code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Forontiers Pipe
# ## Importing code
# +
# %load_ext autoreload
# %autoreload 2
import os
from os import listdir
from os.path import isfile, join, isdir
import shutil
import pygraphviz
import networkx
import plotly
import rpy2
from tkinter import filedialog
from tkinter import *
import itertools
import threading
import subprocess
from functools import partial
import Processing.include_exclude_alignment as in_ex_align
import Processing.genome_3nt as genome_3nt
from Processing.analyze_editing_percent import filter_pileup_by_categories,analyse_multiple_editing_percent_files
from Processing.pileup_sorting import pileup_sort
from parallel_commands import parallel_commands, Command
from Processing.union_set_pileup import union_pileups
from Filtering.filter_pileup_by_multiple_existing_snps import snp_algebra, snp_detect
from Filtering.filter_pileup_by_consensus_site import filter_by_consensus
from Experiments.forontiers_jupyter.site_loss_by_group_plot import site_loss_by_group_plot
from Experiments.forontiers_jupyter.editing_type_count_by_group_plot import editing_type_count_by_group
from pipe_utils import *
from PIL import Image #has to be imported after pipe_utils/tkinter to avoid Image being overwritten
#import Processing.genome_to_transcriptome # Todo - check what is the real name
# -
# ## Aligning the data
# ### Building the allignment graph spec
# Now we are building the allignment graph. this graph allows us to run number of alignments one after the other , which each alignment is on the fastq files which containes the lines the previous alignment didn't align.
#
# Beside the alignment that is being done in each step, we can do an additional action before or\and after the alignment.
#
# Below, we are defining a spec_dict that will contain the structure of the alignments and before and\or after actions.
#
# #### Defining the alignment nodes
# We define the different types of alignments in a python dictionary where:
# * the key is the name we choose for this alignment
# * the value is a 3-tuple of the form (alignment_flags,preprocessing_func,post_processing_func)
#alignment types these are flags that will go to the aligner
norep_align = " -m 2 -n 3"
repetative_align = " -m 100 -n 3 "
# +
# building dict
#no special processing
spec_dict = dict()
spec_dict["norep"]=(norep_align,None,None)
spec_dict["rep"]=(repetative_align,None,None)
#adding non rep and rep 3nt genome
for base_pairs_name,pre,post in genome_3nt_all_combination_spec():
spec_dict["norep_hyper_"+base_pairs_name]=(norep_align,pre,post)
for base_pairs_name,pre,post in genome_3nt_all_combination_spec():
spec_dict["rep_hyper_"+base_pairs_name]=(repetative_align,pre,post)
#adding rep and norep transcriptome for each 3nt combination
#TODO - if you want to remove the transcriptom , delete it here
for base_pairs_name in all_genome_pair_combinations():
spec_dict["norep_transcriptome_"+"_".join(base_pairs_name)]=(norep_align,transcriptom_func,None)
for base_pairs_name in all_genome_pair_combinations():
spec_dict["rep_transcriptome_"+"_".join(base_pairs_name)]=(repetative_align,transcriptom_func,None)
#print(spec_dict)
# -
# #### Defining the graph dependecies
# To define our graph dependencies, we generate a dictionary whos keys are all the names of `spec_dict` and the values of each key, `k`, is a list with the names of "fathers" of `k`.
# +
# generate empty dict
graph_dict={name:[] for name in spec_dict.keys()}
for key,fathers in graph_dict.items():
# non rep is starting node
if key == "norep":
continue
# rep is son of norep
if key == "rep":
fathers.append("norep")
continue
# if we reached here, the name has a base_pair
key_split=key.split('_')
base_pair_name='_'.join(key_split[-2:])
# hyper norep are all sons of rep
if "norep_hyper_" in key:
fathers.append("rep")
continue
# hyper rep is son of hyper norep with same bps
if "rep_hyper_" in key:
fathers.append("norep_hyper_"+base_pair_name)
continue
# hyper norep are all sons of rep
if "norep_transcriptome_" in key:
fathers.append("rep_hyper_"+base_pair_name)
continue
# hyper norep are all sons of rep
if "rep_transcriptome_" in key:
fathers.append("norep_transcriptome_"+base_pair_name)
continue
#print(graph_dict)
# +
dg = networkx.DiGraph(graph_dict)
graphviz_dg = networkx.nx_agraph.to_agraph(dg.reverse())
graphviz_dg.layout(prog='dot')
graphviz_dg.draw('graph.png')
image = Image.open('graph.png')
image.thumbnail((960,960)) #keeps the ratio, but take the longer side in one of the bounderies consideration
display(image)
# -
# #### Defining the plot, groupings
# Like in the case of 3nt genomes, we need to split a single conecpt into multiple alignment nodes even though we would like to aggregate all said nodes for visualization and statistics. Conversly, sometimes there are too many nodes for a metadata plot of all of them to be managable and we would like to cluster the nodes for visualization purposes.
# Here we group our nodes into distinct groups, using a dictionary, they key will be the name of the group and the value will be a set of all node names belonging to our group.
# +
group_dict ={
"norep":{"norep"},
"rep":{"rep"},
"norep_hyper":{"norep_hyper_"+"".join(name) for name in all_genome_pair_combinations()},
"rep_hyper":{"rep_hyper_"+"".join(name) for name in all_genome_pair_combinations()},
"norep_transcriptome":{"norep_transcriptome_"+"".join(name) for name in all_genome_pair_combinations()},
"rep_transcriptome":{"rep_transcriptome_"+"".join(name) for name in all_genome_pair_combinations()}
}
steps_reverse = ["rep_transcriptome","norep_transcriptome","rep_hyper","norep_hyper","rep","norep"]
#print(group_dict)
# -
# ### Selecting your source files
# choose analysis output folder
# +
proj_folder ="/home/lammlab/Documents/frontiers_test/tt"
if proj_folder is None:
proj_folder = folder_selector()
print(proj_folder)
# -
# choose the exprimental rna seq fastqs for the analysis - mark a number of files by pressing "ctrl" in between files choosing
# +
positive_fastqs = ['/home/lammlab/Documents/frontiers_test/files_for_test/test_test_mRNA.fastq']
if positive_fastqs is None:
positive_fastqs = files_selector()
print (positive_fastqs)
# -
# choose the reference fasta file for the analysis
# +
reference_library = "/home/lammlab/Documents/frontiers_test/files_for_test/ws220-genes_expanded.fasta"
if reference_library is None:
reference_library = file_selector()
print(reference_library)
# -
# choose the control fastqs for the analysis (dna reads and ADAR mutant reads)- mark a number of files by pressing "ctrl" in between files choosing
# +
negative_fastqs = ['/home/lammlab/Documents/frontiers_test/files_for_test/test_test_negative.fastq']
if negative_fastqs is None:
negative_fastqs = files_selector()
print(negative_fastqs)
# -
# ### Running alignment
# Make sure that, your `spec,graph,group` dictionaries are configures properly. The next cell will run the inclusion exclusion alignment.
# #### Creating the directories structure
# +
original_output_folder_name = "original"
# creating the folders for the positive fastqs analysis, as subdirs of the proj_folder
fastq_projects_list = []
for fastq_file in positive_fastqs:
new_proj=os.path.join(proj_folder,"results_"+os.path.basename(fastq_file).split(".")[0])
if not os.path.exists(new_proj):
os.mkdir(new_proj)
fastq_projects_list.append(new_proj)
# creating the folders for the negative fastqs analysis, as subdirs of the proj_folder
neg_fastq_project_list = []
for fastq_file in negative_fastqs:
new_proj=os.path.join(proj_folder,"results_"+os.path.basename(fastq_file).split(".")[0])
if not os.path.exists(new_proj):
os.mkdir(new_proj)
neg_fastq_project_list.append(new_proj)
all_projects = [] + fastq_projects_list + neg_fastq_project_list
# creating the output folder of the pileup files from the alignment graph
# and the subdirectories of the rest of the analysis
for project in all_projects:
if project in fastq_projects_list:
# creating the results folder of the editing type plot for the positive fastqs
if not os.path.exists(new_proj):
os.mkdir(os.path.join(project,"editing_type_plot"))
os.mkdir(os.path.join(project,"site_loss_plot"))
for group in group_dict.keys():
dir1=os.path.join(project,group)
if not os.path.exists(new_proj):
os.mkdir(dir1)
# creating the pileup results folders for every group (the folder before filterring),
# for negative and positive fastqs
if not os.path.exists(new_proj):
os.mkdir(os.path.join(dir1, original_output_folder_name))
os.mkdir(os.path.join(dir1, "sorted_pileups"))
if project in fastq_projects_list:
# creating the results filterring folders for the positive fastqs steps.
if not os.path.exists(os.path.join(dir1,"no_change_filterout")):
os.mkdir(os.path.join(dir1,"no_change_filterout"))
os.mkdir(os.path.join(dir1,"reads_threshold_filterout"))
os.mkdir(os.path.join(dir1,"snp_removed"))
os.mkdir(os.path.join(dir1,"editing_sites_filterout"))
os.mkdir(os.path.join(dir1,"consensus_filterout"))
os.mkdir(os.path.join(dir1,"editing_percent_analysis"))
# -
# ##### directories structure overview
# this is how the directories structure looks like:
print_structure(proj_folder)
# #### aligning
# +
#calling the function that run's the analysis (alignments)
#TODO - maybe need to change from threads (threading lib) to multiprocessing - use the example of parallel_commands below
print(reference_library)
threads = []
for project, fastq_file in zip(fastq_projects_list,positive_fastqs):
t = threading.Thread(target=in_ex_align.do_include_exclude_alignment, args=(fastq_file, reference_library, spec_dict, graph_dict, project))
t.start()
threads.append(t)
#in_ex_align.do_include_exclude_alignment(fastq_file, reference_library, spec_dict,graph_dict, outdir=project)
for project, fastq_file in zip(neg_fastq_project_list,negative_fastqs):
t = threading.Thread(target=in_ex_align.do_include_exclude_alignment, args=(fastq_file, reference_library, spec_dict, graph_dict, project))
t.start()
threads.append(t)
for thread in threads:
thread.join()
#for project, fastq_file in zip(neg_fastq_project_list, negative_fastqs):
# in_ex_align.do_include_exclude_alignment(fastq_file, reference_library, spec_dict,graph_dict, project)
# -
# #### moving the files to folders
# +
# moving the
for project in all_projects:
#move each group to a unique group folder
files = [f for f in listdir(project) if isfile(join(project, f))]
print(files)
#group_dict = dict()
for step in group_dict.keys():
for file in files:
for step in steps_reverse:
if "_"+step in file:
shutil.move(os.path.join(project,file),os.path.join(project,step))
break
# on each folder call to create a pileup from the sam files
# -
# ### Pileup creation
# +
#create bam
#TODO - maybe need to change from threads (threading lib) to multiprocessing - use the example of parallel_commands below
threads=[]
for project in all_projects:
for group in group_dict.keys():
dir1=os.path.join(project,group)
files = [f for f in listdir(dir1) if (isfile(os.path.join(dir1, f)) and f.split(".")[-1] == "sam")]
for file in files:
sample_name=".".join(file.split(".")[:-1])
command="samtools view -bS {file_name} > {out_name}.bam && samtools sort {out_name}.bam -o {out_name}.sorted_bam" \
.format(file_name=os.path.join(dir1,file), out_name=os.path.join(dir1,sample_name))
'''print(command)
res = subprocess.call(command,shell=True)
if res:
print("error")'''
print(command)
t = threading.Thread(target=call_process(command))
t.start()
threads.append(t)
for thread in threads:
thread.join()
# +
#create pileup
#TODO - maybe need to change from threads (threading lib) to multiprocessing - use the example of parallel_commands below
for project in all_projects:
for group in group_dict.keys():
dir1=os.path.join(project,group)
files = [f for f in listdir(dir1) if (isfile(os.path.join(dir1, f)) and f.split(".")[-1] == "sorted_bam")]
output_folder = os.path.join(dir1, original_output_folder_name)
#os.mkdir(output_folder)
for file in files:
sample_name=".".join(file.split(".")[:-1])
command="samtools mpileup -f {fasta} {file_name} | tail -n +3 > {out_name}.pileup" \
.format(fasta=reference_library,file_name=os.path.join(dir1,file), out_name=os.path.join(output_folder,sample_name) )
t = threading.Thread(target=call_process(command))
t.start()
threads.append(t)
for thread in threads:
thread.join()
# -
# sorting the pileup files created
# +
commands = []
#TODO - move the raw data - bam,sam,sorted_bam to a new raw folder
# TODO - make sure in the sorting process that the _sorted.pileup files doesnt get
# to the folder otherwise the plot will not come out correctly
for proj in fastq_projects_list:
for group in group_dict.keys():
dir2=os.path.join(proj,group)
indir=os.path.join(dir2,original_output_folder_name)
outdir=os.path.join(dir2,"sorted_pileups")
files = [f for f in listdir(indir) if ( isfile(os.path.join(indir, f)) and (f.split(".")[-1] == "pileup") and ("_sorted.pileup" not in f) ) ]
for file in files:
output=os.path.join(outdir,file)
filename=os.path.join(indir,file)
com=Command(pileup_sort,[filename,output])
commands.append(com)
parallel_commands(commands)
# -
# ### Filter no change pileuplines
# filter out the lines of pileup that shows no editing (the lins with only "." or "," in them)
# +
commands = []
for proj in fastq_projects_list:
for group in group_dict.keys():
dir2=os.path.join(proj,group)
dir1=os.path.join(dir2,"sorted_pileups")
outdir=os.path.join(dir2,"no_change_filterout")
files = [f for f in listdir(dir1) if (isfile(os.path.join(dir1, f)) and f.split(".")[-1] == "pileup") and "sorted.pileup" not in f]
for file in files:
pileup=os.path.join(dir1,file)
output_pileup=os.path.join(outdir,file)
args_array=[pileup,output_pileup,None,True,None,None]
com=Command(filter_pileup_by_categories, args_array)
commands.append(com)
parallel_commands(commands)
# -
# ### Filter by reads threshold
# +
import Processing.analyze_editing_percent as editing
commands = []
threshold=1 #todo - choose threashold - in 0-100 not 0.something
for proj in fastq_projects_list:
for group in group_dict.keys():
dir1=os.path.join(proj,group)
indir=os.path.join(dir1,"no_change_filterout")
outdir=os.path.join(dir1,"reads_threshold_filterout")
files = [f for f in listdir(indir) if (isfile(os.path.join(indir, f)) and f.split(".")[-1] == "pileup")]
for file in files:
pileup=os.path.join(indir,file)
output_pileup=os.path.join(outdir,file)
com=Command(filter_pileup_by_categories,(pileup,output_pileup,threshold,None,None,None))
commands.append(com)
parallel_commands(commands)
# -
# ### SNP removal of mutant and DNA
# todo - need to ask on what fasta (genes\chrome) to select the right file. and if need to do it twice
# +
#union_pileups
##### union the pileups for the negative files ####
# todo - maybe we dont need to union because of the snp removel script ?
commands=[]
for proj in neg_fastq_project_list:
for group in group_dict.keys():
dir1=os.path.join(proj, group)
indir=os.path.join(dir1, original_output_folder_name)
output=os.path.join(indir,group+"_union.pileup")
files = [os.path.join(indir,f) for f in listdir(indir) if (isfile(os.path.join(indir, f)) and f.split(".")[-1] == "pileup")]
com=Command(union_pileups,[files,output])
commands.append(com)
parallel_commands(commands)
# +
#SNP removal for the main fastq
negative_pileup_list = []
commands = []
for proj in neg_fastq_project_list:
for group in group_dict.keys():
dir2=os.path.join(proj,group)
dir1 = os.path.join(dir2,original_output_folder_name)
files = [ os.path.join(dir1,f) for f in listdir(dir1) if ( isfile(os.path.join(dir1, f)) and ("union" in f ) ) ]
negative_pileup_list = negative_pileup_list + files
for proj in fastq_projects_list:
for group in group_dict.keys():
dir1=os.path.join(proj,group)
indir=os.path.join(dir1,"reads_threshold_filterout")
outdir=os.path.join(dir1,"snp_removed")
files = [f for f in listdir(indir) if (isfile(os.path.join(indir, f)) and f.split(".")[-1] == "pileup")]
for file in files:
output=os.path.join(outdir,file)
in_file=os.path.join(indir,file)
com=Command(snp_algebra, [in_file,negative_pileup_list,snp_detect,output,True])
commands.append(com)
parallel_commands(commands)
# -
# ### Filter by editing sites (by editing percent and noise)
# +
commands = []
editing_thresold=10#todo - fill in read threshold
noise_threshold=90#todo - fill in read threshold
for proj in fastq_projects_list:
for group in group_dict.keys():
dir1=os.path.join(proj,group)
indir=os.path.join(dir1,"snp_removed")
outdir=os.path.join(dir1,"editing_sites_filterout")
#os.mkdir(outdir)
files = [f for f in listdir(indir) if (isfile(os.path.join(indir, f)) and f.split(".")[-1] == "pileup")]
for file in files:
pileup=os.path.join(indir,file)
output_pileup=os.path.join(outdir,file)
com=Command(filter_pileup_by_categories,[pileup,output_pileup,None,None,editing_thresold,noise_threshold])
commands.append(com)
parallel_commands(commands)
# -
# ### Consensus
# +
#TODO - CHECK IF OUTPUT MAKE SENSE
is_sorted=True
k=0.3#todo- fill in the k
commands = []
for proj in fastq_projects_list:
for group in group_dict.keys():
dir1=os.path.join(proj,group)
indir=os.path.join(dir1,"editing_sites_filterout")
outdir=os.path.join(dir1,"consensus_filterout")
files = [os.path.join(indir,f) for f in listdir(indir) if (isfile(os.path.join(indir, f)) and f.split(".")[-1] == "pileup")]
com=Command(filter_by_consensus,[files,k,outdir,is_sorted])
commands.append(com)
parallel_commands(commands)
# -
# ### Site loss plot
# +
commands = []
list_order_of_folders=["consensus_filterout","editing_sites_filterout","snp_removed","reads_threshold_filterout","no_change_filterout"]
for proj in fastq_projects_list:
outdir = os.path.join(proj,"site_loss_plot")
output_file_name=os.path.join(outdir,"site_loss_plot.png")
folders = [os.path.join(proj,f) for f in listdir(proj) if isdir(os.path.join(proj, f)) and "site_loss_plot" not in f and "editing_type_plot" not in f]
com=Command(site_loss_by_group_plot,[folders,output_file_name,original_output_folder_name,list_order_of_folders])
commands.append(com)
parallel_commands(commands)
# -
# ### Editing type count plot
# #### Editing percent analysis
# +
commands = []
min_editing=0.0#todo- fill up
max_noise=1.0#todo- fill up
min_reads=1#todo- fill up
for proj in fastq_projects_list:
for group in group_dict.keys():
dir1=os.path.join(proj,group)
indir=os.path.join(dir1,"consensus_filterout") #input : the pileups after the intire thing of filtering
outdir=os.path.join(dir1,"editing_percent_analysis")
files = [os.path.join(indir, f) for f in listdir(indir) if (isfile(os.path.join(indir, f)) and f.split(".")[-1] == "pileup")]
#analyse_multiple_editing_percent_files(files,out_dir=outdir,add_headers=True,
# min_editing=editing_percent_threshold,
# max_noise=noise_percent_threshold,
# min_reads=count_threshold, edit_tag="edited",csv_genes=True)
vars_list=[files,outdir,None,True,False,min_editing,max_noise,min_reads,"edited",False]
com=Command(analyse_multiple_editing_percent_files,vars_list)
commands.append(com)
parallel_commands(commands)
# -
# #### plot
# +
commands = []
#input : the pileups after the intire thing - then do editing percent anaysis
for proj in fastq_projects_list:
outdir=os.path.join(proj,"editing_type_plot")
#os.mkdir(outdir)
output_file_name=os.path.join(outdir,"editing_type_plot.png")
list_of_dirs=[]
for group in group_dict.keys():
indir=os.path.join(os.path.join(proj,group),"editing_percent_analysis")
list_of_dirs.append(indir)
com=Command(editing_type_count_by_group,[list_of_dirs,output_file_name])
commands.append(com)
parallel_commands(commands)
# -
| Experiments/forontiers_jupyter/Forontiers_Pipe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import numpy as np
import music21 as m21
import pandas as pd
import json
import matplotlib.pyplot as plt
from scipy import stats
from scipy import spatial
import time
import math
from IPython.display import display
from collections import Counter
np.random.seed(777)
# -
# ## Functions
# Function to retrieve a list of midi pitch events and its timestamp
def getMelodyDeltaTimes(eventsintrack):
# Initialize array
DeltaTimes = []
# Initialize cumulative sum
cum_sum = 0
# Initialize variable to track the time delta
prev_deltatime = 0
# Traverse the events
for ev in eventsintrack:
# If a note starts
if (ev.isNoteOn()):
# Get the pitch name and save it with the cumulative sum, midi pitch and name
pitch_in_time = m21.pitch.Pitch(ev.pitch)
DeltaTimes.append((cum_sum, prev_deltatime, pitch_in_time.midi, pitch_in_time.spanish, pitch_in_time))
# Restart the delta time
prev_deltatime = 0
# Else if there is a delta time
elif(str(ev.type) == "DeltaTime"):
# We sum the time
cum_sum += ev.time
# We sum it to the current delta time
prev_deltatime += ev.time
# Return the array
return DeltaTimes
def get_MelodyShapeNgram_NOREST(melody_w_times):
ngram_list = []
for m_el in melody_w_times:
# print(m_el)
current_element = [m_el[2], m_el[0], max(m_el[1],1), 0]
# print(current_element)
ngram_list.append(current_element)
return ngram_list
# # Groups for analysis
# * Reference for the axis
# * Groups of dataset
Y_AXIS_MIDI_PATHS = [
"./CalebRascon/CORPUS/MIDI/",
"./CalebRascon/MIDI_Grammar_SOLO_LEN12/",
"./MilesDavis/MIDI_Grammar_SOLO_LEN12/",
"./CharlieParker/MIDI_Grammar_SOLO_LEN12/"
]
X_AXIS_MIDI_PATHS = [
"./CalebRascon/CORPUS/MIDI/",
"./CalebRascon/MIDI_Grammar_SOLO_LEN12/",
"./MilesDavis/MIDI_Grammar_SOLO_LEN12/",
"./CharlieParker/MIDI_Grammar_SOLO_LEN12/",
"./CalebRascon/MIDI_Grammar_TRADE_Caleb/",
"./CalebRascon/MIDI_Grammar_TRADE_Miles/",
"./CalebRascon/MIDI_Grammar_TRADE_CharlieParker/"
]
# %%time
note_representation_bspline = {"fam":{}}
for MIDI_path_query in Y_AXIS_MIDI_PATHS:
for MIDI_path_test in X_AXIS_MIDI_PATHS:
similarities_all_v_all = {}
for root_ref, dirs_ref, files_ref in os.walk(MIDI_path_query):
for name_ref in files_ref:
# print("+++++++++++++++++++++++++++++")
# print(name_ref)
melody_score_A = m21.converter.parseFile(os.path.join(root_ref, name_ref))
midi_tracks_A = m21.midi.translate.streamToMidiFile(melody_score_A)
melody_w_times_A = getMelodyDeltaTimes(midi_tracks_A.tracks[0].events)
similarities_from_reference = []
similarities_all_v_all[name_ref] = {}
for root, dirs, files in os.walk(MIDI_path_test):
for name in files:
# print(name)
melody_score_B = m21.converter.parseFile(os.path.join(root, name))
midi_tracks_B = m21.midi.translate.streamToMidiFile(melody_score_B)
melody_w_times_B = getMelodyDeltaTimes(midi_tracks_B.tracks[0].events)
comparison_id = "MP_A;{0};M_A;{1};MP_B;{2};M_B;{3}".format(MIDI_path_query, name_ref, MIDI_path_test, name)
print(comparison_id)
# We Save the representation for the family
note_representation_bspline["fam"][comparison_id] = {"msa":[], "msb":[]}
note_representation_bspline["fam"][comparison_id]["msa"] = get_MelodyShapeNgram_NOREST(melody_w_times_A)
note_representation_bspline["fam"][comparison_id]["msb"] = get_MelodyShapeNgram_NOREST(melody_w_times_B)
with open('./note_representation_bspline.json', 'w') as outfile:
json.dump(note_representation_bspline, outfile)
| CaseStudy/CCE_DataGroups_RAWNOTES_BSPLINE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ghost
#
# According to [Wikipedia](https://en.wikipedia.org/wiki/Ghost_(game)):
#
# > **Ghost** *is a written or spoken word game in which players take turns adding letters to a growing word fragment, trying not to be the one to complete a valid word. Each fragment must be the beginning of an actual word, and usually some minimum is set on the length of a word that counts, such as three or four letters. The player who completes a word loses.*
#
# I'd like to create a program to allow any two players (human or computer) to play the game, and I'd like to figure out who wins if both players play optimally. The concepts I will need to define, and my implementation choices, are as follows:
#
# - **Words**: I will read a standard online word list, `enable1`, and make a set of all the words of sufficient length.
# - **Fragment**: a fragment is a `str` of letters, such as `'gho'`.
# - **Beginning**: each word has a set of valid beginnings: for `ghost` it is `{'', g, gh, gho, ghos, ghost}`. "Prefix" is a synonym of "beginning".
# - **Vocabulary**: A `Vocabulary` object holds a set of all the `words` in a dictionary, as well as all the valid `fragments` (beginnings) of the words.
# - **Player**: The first player will be called player `0`; the second player `1`.
# - **Play**: A play is a new fragment formed by adding one letter to the end of the existing fragment.
# - **Legal Play**: A play that is a valid prefix of some word. `enable1.legal_plays('gho') = {'ghos, 'ghou'}`.
# - **Strategy**: A strategy is a function with signature `strategy(vocab, fragment) -> play`.
# - **Game**: `play_game(vocab, *strategies)` plays a game between two (or more) player strategies.
#
#
# # Vocabulary: Words, Fragments, Legal Plays, and `enable1`
#
# `Vocabulary(words)` takes a collection of words as input, stores the words as a set, and also stores all the legal fragments of those words. `legal_plays(fragments)` gives a set of all plays that can be formed by adding a letter to create a legal word fragment. I also define the function `words` to split any string into component words.
# +
class Vocabulary:
"Holds a set of legal words and a set of legal prefix fragments of those words."
def __init__(self, words, minlength=3):
self.words = {word for word in words if len(word) >= minlength}
self.fragments = {word[:i] for word in self.words for i in range(len(word) + 1)}
def legal_plays(self, fragment):
return {fragment + L for L in alphabet} & self.fragments
alphabet = 'abcdefghijklmnopqrstuvwxyz'
words = str.split # Function to split a str into words
# -
# Here is a small example with a vocabulary of just three words:
# +
v = Vocabulary(words('game ghost ghoul'))
v.words
# -
v.fragments
# And here is a large vocabulary, from a standard online Scrabble™ word list known as `enable1`:
! [ -e enable1.txt ] || curl -O http://norvig.com/ngrams/enable1.txt
enable1 = Vocabulary(words(open('enable1.txt').read()))
# Let's explore `enable1`:
len(enable1.words), len(enable1.fragments), max(enable1.words, key=len)
enable1.legal_plays('gho')
enable1.legal_plays('ew')
enable1.legal_plays('th')
# # Players and Winners
#
# The first player is `0` and the second player is `1`. These names are convenient because:
# - During the course of the game, the player whose turn it is to play next is always the length of the current fragment mod 2.
# - When the game ends, the winning player is the length of the current fragment mod 2.
#
to_play = winner = lambda fragment: len(fragment) % 2
# # Who Wins?
#
# Who wins a game if both players play rationally? To answer that, consider the general situation at some point
# during the game where
# a player is presented with a fragment. That player can win if either:
# - The fragment is a complete word (meaning the other player just formed the word, and thereby lost).
# - The fragment is not a legal fragment (meaning the other player made a mistake, and thereby lost).
# - At least one of the legal plays in this position puts the opponent in a position from which they *cannot* win.
#
# The function `win(vocab, fragment)` implements this. It returns a winning fragment if there is one, otherwise `False`. In particular, it returns `fragment` if the current player has already won (because `fragment` is
# a word or illegal fragment), and it returns one of the legal plays if that play leads to a position from
# which the opponent *cannot* `win`.
def win(vocab, fragment=''):
"""Does the current player have a forced win?
If so, return a play (or the current fragment) that leads to a win."""
if fragment in vocab.words or fragment not in vocab.fragments:
return fragment
for play in vocab.legal_plays(fragment):
if not win(vocab, play):
return play
return False
# Let's test `win` to gain some confidence that we got it right:
# No winning play because all words have odd number of letters.
win(Vocabulary(words('cat camel gecko')))
# 'g' is a winning play (but 'c' would be a loser)
win(Vocabulary(words('cat camel goat gerbil')))
# No winning play; doomed to 'camel' or 'gar'
win(Vocabulary(words('cat camel goat gecko gerbil gar')))
# 'g' wins because 'ga' can be answered with 'gan' and 'ge' with 'ge'
win(Vocabulary(words('cat camel goat gecko gerbil gar gannet')))
# # TL;DR: The Answer
#
# Can the first player win with the `enable1` vocabulary?
win(enable1)
# **No.** The game is a win for the second player, not the first.
# This agrees with [xkcd](https://xkcd.com/)'s Randall Monroe, who [says](https://blog.xkcd.com/2007/12/31/ghost/) *"I hear if you use the Scrabble wordlist, it’s always a win for the second player."*
#
# But ... Wikipedia says that the minimum word length can be "three or four letters." In `enable1` the limit was three; let's try again with a limit of four:
# +
enable1_4 = Vocabulary(enable1.words, 4)
win(enable1_4)
# -
# **Yes.** The first player can win with this vocabulary, by playing `'n'` first (and there might be other first plays that also force a win). It makes sense that it is easier for the first player to win, because we've eliminated a bunch of three-letter words from the vocabulary, all of which are losers for the first player. So here's a good meta-strategy: Say "*Hey, let's play a game of Ghost. We can use the `enable1` word list. Would you like the limit to be 3 or 4 letters?*" Then if your opponent says three (or four) you can say "*OK, since you decided that, I'll decide to go second (or first).*"
# # Playing the Game: Strategies
#
# We define a *strategy* as a function that is given a vocabulary and a fragment as arguments, and returns a legal play. Below we define `rational`, a strategy that wins whenever it is possible to do so and plays randomly otherwise, and `ask` (a strategy factory that returns a strategy that, when called, will ask the named person to input a fragment).
# +
import random
def rational(vocab, fragment):
"Select a play that makes opponent not win (if possible), otherwise a random play."
return win(vocab, fragment) or random.choice(list(vocab.legal_plays(fragment)))
def ask(name):
"Return a strategy that asks for the next letter."
return lambda _, fragment: input('Player ' + name + ' given "' + fragment + '" plays? ')
# -
# Here is a function to play a game. You give it a vocabulary, two (or possibly more) strategies, and optionally a `verbose` keyword to say if you want a line printed for each play or not.
# +
from itertools import cycle
def play(vocab, *strategies, verbose=False):
"Return (winner, final_fragment) for a game of Ghost between these strategies."
fragment = ''
for (p, strategy) in cycle(enumerate(strategies)): # p is the player number
play = strategy(vocab, fragment)
if verbose:
print('Player {}, given "{}", plays "{}".'.format(p, fragment, play))
if play not in vocab.legal_plays(fragment):
return (winner(fragment + '?'), play) # Player loses for making an illegal play
elif play in vocab.words:
return (winner(play), play) # Player loses for making a word
else:
fragment = play # Keep playing
# -
play(enable1, rational, rational)
# +
# Does player 1 win every time?
[play(enable1, rational, rational, verbose=False) for _ in range(20)]
# -
play(enable1_4, rational, rational)
play(enable1, ask('P'), rational)
# # Minimizing Possible Outcomes
#
# Now we know how to play perfectly, if we have a computer handy to execute the strategy.
# But can we summarize the strategy into a form that is small enough that a human can memorize it? I will define the function `outcomes(vocab, fragment, player)` to return a set of all the words that are possible outcomes of a game, where the opponent can use any strategy whatsoever, but `player` uses a strategy that is:
#
# - *Rational*: plays towards a forced win whenever there is one.
# - *Exploitive*: otherwise tries to give the opponent an opportunity to make a mistake that can be exploited.
# - *Minimizing*: within the above constraints, returns the smallest possible set of words.
# +
import textwrap
def outcomes(vocab, fragment, player):
"The smallest set of outcome words, if player tries to win, and make the set small."
if fragment in vocab.words:
return {fragment}
else:
cases = [outcomes(vocab, play, player) for play in vocab.legal_plays(fragment)]
if to_play(fragment) == player: # Player picks the top priority case
return min(cases, key=lambda words: priority(words, player))
else: # Oher player could pick anything
return set.union(*cases)
def priority(words, player):
"""Return (lossiness, number_of_words, total_number_of_letters),
where lossiness is 0 if no losses, 1 if mixed losses/wins, 2 if all losses.
The idea is to find the list of outcome words that minimizes this triple."""
lossiness = (0 if all(winner(word) == player for word in words) else
1 if any(winner(word) == player for word in words) else
2)
return (lossiness, len(words), sum(map(len, words)))
def report_outcomes(vocab, player):
"Find minimal outcomes for player; print info."
oc = outcomes(vocab, '', player)
winners = {w for w in oc if winner(w) == player}
def fill(label, words):
if words:
text = '{} ({}): {}'.format(label, len(words), ' '.join(sorted(words)))
print(textwrap.fill(text))
print('Outcomes ({}) for player {}:'.format(len(oc), player))
fill('Losers', oc - winners)
fill('Winners', winners)
# -
# ## Minimizing Outcomes for Player 0
#
# Let's see what minimal set of words player 0 can force the game into (with both vocabularies):
report_outcomes(enable1, 0)
# **Interesting!** There are only 6 words; it wouldn't be hard for a human to memorize these. Then, when you are playing as player 0, pick `'q'` first, and then try to steer the game to one of the 5 words with an even number of letters. Unfortunately, one word, `'qursh'` (a monetary unit of Saudi Arabia), has an odd number of letters, which means that if the opponent replies to `'q'` with `'qu'` and to `'qur'` with `'qurs'`, then player 0 will lose. But if the opponent makes any other responses, player 0 will win.
report_outcomes(enable1_4, 0)
# **Neat!** Only 7 words, and the first player can always win by forcing the opponent to one of these words.
#
# ## Minimizing Outcomes for Player 1
#
# Since player 0 can pick any letter, the minimal `outcomes` set for player 1 must be at least 26 words. Let's see how much bigger it turns out to be.
#
# With `enable1` we already know that player 1 can force a win, so all the words in the `outcomes` set will have odd length:
report_outcomes(enable1, 1)
# Memorize this list and you will never lose as player 1.
#
# How about with the other vocabulary?
report_outcomes(enable1_4, 1)
# In this case there are 85 words, four of which are losses for player 1. But the other 81 words are wins, so with this strategy you'd have a good chance against an imperfect opponent.
# # SuperGhost
#
# In the variant *SuperGhost*, players can add a letter to either the beginning or the end of a fragment, as long as this forms a fragment that is part of some word. As Wikipedia says, given the fragment `era`, a player might play `bera` or `erad`. I was thinking of SuperGhost when I made the design decision to encapsulate `legal_plays` as a method of `Vocabulary`, rather than as a separate function. Because I did that, I should be able to use all the existing code if I just make a new class, `SuperVocabulary`, that finds *all* fragments (i.e. infixes) rather than just the beginning fragments (i.e. prefixes), and if I change `legal_plays` to add letters to both ends.
class SuperVocabulary(Vocabulary):
"Holds a set of legal words and a set of legal infix fragments of those words."
def __init__(self, words, minlength=3):
self.words = {word for word in words if len(word) >= minlength}
self.fragments = {word[i:j] for word in self.words
for i in range(len(word))
for j in range(i, len(word) + 1)}
def legal_plays(self, fragment):
"All plays (adding a letter to fragment) that form a valid infix."
return {p for L in alphabet for p in (fragment + L, L + fragment)} & self.fragments
# Now I will create `SuperVocabulary` objects for 3- and 4-letter versions of `enable1`, and check out how many fragments there are in each variant:
enable1_3super = SuperVocabulary(enable1.words, 3)
enable1_4super = SuperVocabulary(enable1.words, 4)
# Example legal plays
enable1_3super.legal_plays('crop')
# +
# Can the first player win in SuperGhost with 3-letter words?
win(enable1_3super)
# +
# How about with a 4-letter limit?
win(enable1_4super)
# -
# The first player can win with or without three-letter words. And unless the first player is perfect, the rational strategy can do pretty well as seond player as well. Here is a sample game:
play(enable1_3super, ask('P'), rational)
# I would like to give a concise summary of the strategy for SuperGhost, but my existing `outcomes` function won't do it. That's because it is not enough to know that a particular word results in a win; we have to know in what order the letters of the word are added. I'll leave it as an exercise to find a good way to summarize SuperGhost strategies.
#
# # SuperDuperGhost
#
# In the variant *SuperDuperGhost*, players have an option to reverse the fragment before adding a letter to the beginning or end. As Wikipedia says, given the fragment `era`, a player might play `bera, erad, nare,` or `aren`.
# Wikipedia is not clear, but I interpret this as meaning that the fragment played must still be a fragment of a word (not a reversed fragment of a word). Again, all we need is a new subclass:
class SuperDuperVocabulary(SuperVocabulary):
"Holds a set of legal words and a set of legal infix fragments of those words."
def legal_plays(self, fragment):
"All plays that form a valid infix; optionally reverse fragment first."
def all4(f, L): return (f + L, f[::-1] + L, L + f, L + f[::-1])
return {p for L in alphabet for p in all4(fragment, L)} & self.fragments
enable1_3duper = SuperDuperVocabulary(enable1.words)
enable1_4duper = SuperDuperVocabulary(enable1.words, 4)
# Example legal plays
enable1_3duper.legal_plays('crop')
# Now we should check who wins. But I'm impateint: I tried `win(enable1_3duper)`, waited a minute, and it still hadn't returned, so I interrupted that computation and threw in a `lru_cache` decorator; which stops `win` from wasting time repeating the same computation over and over. This brings the time down to about 2 seconds.
# +
from functools import lru_cache
@lru_cache(None)
def win(vocab, fragment=''):
"""Does the current player have a forced win?
Return fragment if the player has already won, or return a play that forces a win."""
if fragment in vocab.words or fragment not in vocab.fragments:
return fragment
for play in vocab.legal_plays(fragment):
if not win(vocab, play):
return play
return False
# -
win(enable1_3duper)
win(enable1_4duper)
# The first player can win with either vocabulary. Here's a sample game.
play(enable1_3duper, rational, rational, verbose=True)
# Let's see how many fragments each vocabulary takes:
[len(v.fragments)
for v in [enable1, enable1_4, enable1_3super, enable1_4super, enable1_3duper, enable1_4duper]]
# # Summary
#
# Here's a summary of what we have learned. (*Note:* the bold **qursh** means it is a losing word):
#
# | Variant | Shortest | Winner | First Player Forced Outcomes | 2nd Outcomes | Fragments
# |:----: |:---: |:---: |:---: |:---: |---:
# | Ghost | 3 | Second | qaid qiviut qoph **qursh** qurush qwerty | 55 words | 387,878
# | Ghost | 4 | First | naan nene ngultrum nirvanic nolo null nyctalopia | 85 words | 387,844
# | Super | 3 | First | ? | ? | 1,076,434
# | Super | 4 | First | ? | ? | 1,076,431
# | SuperDuper | 3 | First| ? | ? | 1,076,434
# | SuperDuper | 4 | First| ? | ? | 1,076,431
#
# # Further Work
#
# Here are some additional ideas to play with:
#
# - **Exploitation:** What are some good strategies when you are not guaranteed to win, to exploit an imperfect human opponent? Can you steer the game so that you win if the opponent is unfamiliar with some obscure word(s)? You might need a file of [word frequencies](http://norvig.com/ngrams/count_1w.txt).
# - **Security:** A strategy function could *cheat*, and modify `vocab.words`, inserting or deleting some crucial words to ensure victory. Can you harden `play` (and/or change `Vocabulary`) to protect against that?
# - **Pruning:** Currently `Vocabulary` saves words and fragments that could never be reached in a game. For example, because `'the'` is a word that ends the game, we could never reach `'them'` or `'theme'` or `'thermoluminescences'`. Can you prune away these redundant words/fragments?
# - **Multi-player:** `play(enable1, ask('A'), ask('B'), ask('C'))` will play a three-player game. But `rational` (along with `win` and `winner`) would no longer work, since they assume there are exactly two players. Can you alter them to allow *n* players?
# - **SuperGhost Summary:** Can you summarize a SuperGhost or SuperDuperGhost strategy in a way that a human can memorize?
# - **Xghost:** In *Xghost*, a letter can be added anywhere, so from the fragment `'era'` you could play `'erba'`.
# - **Spook:** In *Spook*, letters can be rearranged before adding one, so from the fragment `'era'` you could play `'bear'`.
| ipynb/Ghost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style='text-align:center'>Simulação de Canal de Comunicação segundo Modelo Erceg</h1>
# +
import numpy as np
import random
import matplotlib.pyplot as plt
from PIL import Image
import math
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
# #%%latex
# -
# O modelo Erceg foi construído e estudado de acordo com o dado experimentais coletados pela AT&T Wireless Service em 95 estações dentro dos EUA operando a 1,9GHz. O modelo enumera três categorias de terrenos que provocam perdas no sinal e eles são divididos da seguinte maneira; a categoria A representa terrenos com morros e colinas e com alta de densidade de obejetos no caminho do sinal, o que representa uma alta perda de percurso. Categoria C representa um terreno plano com baixa densidade de objetos no caminho do sinal, representando uma baixa perda do sinal, já a categoria B é constituída por terrenos montanhososo com baixa densidade de objetos ou terrenos plano com densidade consideravel de objetos no caminho do sinal, basicamente a categoria B representa um meio termo entre as categorias A e C, com perda de sinal mediana quando compara com as outras.
#
# Para todas as três categorias o caminho de perda mediana é representada pela mesma equação com a condição de $d > d_0$, no caso a seguinte equação.$$
# \begin{equation}\label{eq:erceg}
# P_L(dB) = 20\log_{10}(4\pi d_0/\lambda)+10\gamma \log_{10}(d/d_0)+s
# \end{equation}
# $$Onde $\lambda$ é o comprimento de onda do sinal, s é o efeito de sombreamento do sinal, $\gamma$ representa o caminho de menor perda considerando as três categorias do modelo, representado pela seguinte equação
# $$
# \gamma = a - bh_b + c/h_b
# $$
# $h_b$ representa a altura da estação base em metros, normalmente entre 10 e 80m, $d_0=100m$ e os valores nominais de a, b e c variam de acordo com a catgoria do terrenos, seus valores variam de acordo a seguinte tabela:
#
# |Parametro|Categoria A|Categoria B|Categoria C|
# |---------|-----------|-----------|-----------|
# |a|4,6|3|3,6|
# |b|0,0076|0,0065|0,005|
# |c|12,6|17,1|20|
#
#
#
#
# Definiçao das variáveis $d_0$, $h_b$ e $s$
d0 = 100
hb = 50
s = 6
f = 1900000000
# Definição das distâncias a serem simuladas
d = np.arange(1, 1000, 1)
# Construção dos arrays das categorias
a = np.array([4.6, 3, 3.6])
b = np.array([0.0076, 0.0065, 0.005])
c = np.array([12.6, 17.1, 20])
# Array com os valores de gamma
gamma = [a[i]-b[i]*hb+c[i]/hb for i in range(3)]
# Com os valores instânciados podemos analisar a resposta para cada uma das três situações
v1 = 20*np.log10(4*math.pi*d0*f/300000000)+10*gamma[0]*np.log10(d/d0)+s
v2 = 20*np.log10(4*math.pi*d0*f/300000000)+10*gamma[1]*np.log10(d/d0)+s
v3 = 20*np.log10(4*math.pi*d0*f/300000000)+10*gamma[2]*np.log10(d/d0)+s
# Plotando os resultados obtidos
# +
plt.plot(d, v1, label='Categoria A')
plt.plot(d, v2, label='Categoria B')
plt.plot(d, v3, label='Categoria C')
plt.xlabel('Distancia entre estacao movel e a base [m]')
plt.ylabel('Perda de potencia incidente na estacao movel [db]')
plt.legend()
# plt.title('Perda de Potencia em funcao da distancia entre estacao movel e base')
# -
# Analisando as curva obtidas em função da distância entre a estação rádio base e o dispositivo móvel podemos verificar para distâncias $d<d_0$ as curvas apresentam comportamento similar, apresentado perdas de potência próximas, um forte indicativo de que os fatores usados para compor o modelo Erceg entregam praticamente os mesmos valores independente da categoria do terreno por onde sinal se propaga. É interessante notar que as divergência entre as curvas para cada categoria se tornam evidentes quando $d>2d_0$, fazendo as curvas divergirem.
# Um ponto interessante a se simular é fixar a distância entre a estação rádio móvel, no caso para as três situações possíveis, $d<d_0$, $d=d_0$ e $d>d_0$ e verificar o comportamento de uma frequencia variavel, alguns limites para esse caso são importantes pois o modelo Erceg foi projetado para trabalhar com frequências próximas de 2GHz, logo não faz sentido simular algo para todo o espectro, logo a simulação foi centrada em 2GHz com uma faixa de $+/-$300MHz, ou seja, foi simulado o comportamento da perda de potência para uma distância fixa para a faixa de frequência iniciando em 1,7GHz e finalizando em 2,3GHz no passo de 100KHz.
# +
fval = np.arange(1700000000, 2300000000, 100000)
fv1 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[0]*np.log10(100/d0)+s
fv2 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[1]*np.log10(100/d0)+s
fv3 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[2]*np.log10(100/d0)+s
fv4 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[0]*np.log10(50/d0)+s
fv5 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[1]*np.log10(50/d0)+s
fv6 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[2]*np.log10(50/d0)+s
fv7 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[0]*np.log10(200/d0)+s
fv8 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[1]*np.log10(200/d0)+s
fv9 = 20*np.log10(4*math.pi*d0*fval/300000000)+10*gamma[2]*np.log10(200/d0)+s
# +
plt.plot(fval, fv1, label='Categoria A d=d0')
plt.plot(fval, fv2, label='Categoria B d=d0')
plt.plot(fval, fv3, label='Categoria C d=d0')
plt.plot(fval, fv4, label='Categoria A d<d0')
plt.plot(fval, fv5, label='Categoria B d<d0')
plt.plot(fval, fv6, label='Categoria C d<d0')
plt.plot(fval, fv7, label='Categoria A d>d0')
plt.plot(fval, fv8, label='Categoria B d>d0')
plt.plot(fval, fv9, label='Categoria C d>d0')
plt.xlabel('Frequencia do sinal')
plt.ylabel('Perda de potencia incidente na estacao movel [db]')
plt.legend()
# -
# Analisando as curvas geradas, chama atenção que quando $d=d_0$ as curvas são iguais, ou seja, independente da categoria a variação da frequência produz o mesmo resultado, contudo este comportamente era previsto pelo modelo pois com $d=d_0$ o termo que depende das categorias de terreno é anulado do modelo, fazendo com que ele dependesse apenas da frequência.
#
# Para distâncias $d<d_0$ o modelo apresenta baixa perdas quando comparado com os outros casos simulados, configurando este como o caso ideal de uso. Para distâncias $d>d_0$ o modelo já apresenta perdas maiores quando colocado em comparação com as outras curvas geradas.
# Outra simulacao possivel seria variar a altura da torre fixando a uma frequencia e um $d$ especifico assim seria possivel enteder qual a relacao da alura da torre
# http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.210.3876&rep=rep1&type=pdf
# http://morse.colorado.edu/~tlen5510/text/classwebch3.html
# https://www.mathworks.com/matlabcentral/fileexchange/39322-erceg-model
# https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Working%20With%20Markdown%20Cells.html
# <h1 style='text-align:center'>Simulação de Propagacao em Multipercurso</h1>
# $$X(t)=X_c(t)+jX_s(t)$$
# $$X_c(t)=\dfrac{2}{\sqrt M}\sum_{n=1}^{M}\cos (\Psi_n)\cos (\omega_d t \cos a_n + \phi)$$
# $$X_s(t)=\dfrac{2}{\sqrt M}\sum_{n=1}^{M}\sin (\Psi_n)\cos (\omega_d t \cos a_n + \phi)$$
# $$a_n = \dfrac{2\pi n - \pi + \theta}{4M}$$
#
# Onde $\theta$, $\phi$ e $\Psi_n$ sao estatisticamente independente, variando entre $[-\pi,\pi)$ para todo $n$. A simulacao do artigo foi testada com $M = 8$
#
# A expressão pode ser manilpulada de forma a reunir todos os fatores em uma única expressão.
# $$
# X_k(t)= \sqrt{\dfrac{2}{M}} \left\{ \sum_{n=1}^{M}\cos (\Psi_{n,k})\cos \left[\omega_d t \cos \left(\dfrac{2\pi n - \pi + \theta}{4M}\right) + \phi_k \right] \right\} + j\left\{ \sum_{n=1}^{M}\sin (\Psi_{n,k})\cos \left[\omega_d t \cos \left(\dfrac{2\pi n - \pi + \theta}{4M}\right) + \phi_k \right] \right\}
# $$
# Primeiro é preciso instanciar as bibliotecas para gerar um distribuição uniforme para $\theta$, $\phi$ e $\Psi_n$ entre $[-\pi,\pi)$
# +
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import pandas as pd
fig, ax = plt.subplots(figsize=(25, 15))
M = 8
const = np.sqrt(2/M)
ts = np.arange(0, 100, step=0.025)
Nstats = [10, 50, 100]
colors = ['r', 'g', 'b']
trials = {}
for i, Nstat in enumerate(Nstats):
for N in range(Nstat):
Xc_t = []
theta_multi = np.random.uniform(-np.pi, np.pi)
phi_multi = np.random.uniform(-np.pi, np.pi)
psi_multi = np.random.uniform(-np.pi, np.pi, size=M)
for t in ts:
def return_an(n):
return (2*np.pi*n-np.pi+theta_multi)/(4*M)
Xc = np.sum(np.array([np.cos(psi_multi[m-1])*np.cos(2*np.pi*t*np.cos(return_an(m)+phi_multi))
for m in range(1, M+1)]))
Xc_t.append(const*Xc)
Xc_t = np.array(Xc_t)
trials[N] = Xc_t
df = pd.DataFrame(trials)
averages = df.mean(axis=1)
c = np.correlate(averages.values, averages.values, mode='same')
sns.lineplot(ts, c / c.max(), color=colors[i], label='Nstat ' + str(Nstat), ax=ax)
ax.set_xlim(50, 65)
ax.legend()
ax.set_xticklabels(list(range(0, 15, 2)))
ax.set_xlabel('Tempo Normalizado')
ax.set_ylabel('Autocorrelação X_c(t)')
# -
# +
E_0 = np.sqrt(2)
C_n = 1/np.sqrt(4*M+2)
phi_multi = np.random.uniform(-np.pi, np.pi)
gc_t = []
ts = np.arange(0, 30, step=0.025)
M = 8
phi_n = np.random.uniform(-np.pi, np.pi)
a_n = np.random.uniform(-np.pi, np.pi)
for t in ts:
def return_an(n):
return p.random.uniform(-np.pi, np.pi)(2*np.pi*n/(4*M+2))
gc = E_0*np.sum(np.array([C_n*np.cos(2*np.pi*t*np.cos(a_n))
for m in range(1, M+1)]))
gc_t.append(gc)
fig, ax = plt.subplots(figsize=(25, 15))
c = np.correlate(np.array(gc_t), np.array(gc_t), mode='same')
sns.lineplot(ts, c / c.max(), ax=ax)
ax.set_xlim(15, 30)
# -
| Channel-Simulation/Erceg-Model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import keras
from keras.datasets import mnist
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten, Conv2D, MaxPool2D, Activation
import tensorflow as tf
# +
NUM_CLASSES = 10
IMG_ROWS = IMG_COLS = 28
# test ve egitim icin verileri otomatik cikartiliyor
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# egitim icin kullanilacak inputlarin boyutu degistirildi, xtrain=array, 1 tane kanal rayisi var siyahbeyaz. rgbde 3 kanal var
X_train = X_train.reshape(X_train.shape[0], IMG_ROWS, IMG_COLS, 1)
X_test = X_test.reshape(X_test.shape[0], IMG_ROWS, IMG_COLS, 1)
X_train = X_train.astype('float32') / 255.0
X_test = X_test.astype('float32') / 255.0
#y_train i matrixe cevir -> one_hot'in karsiligi
y_train = keras.utils.to_categorical(y_train, NUM_CLASSES)
y_test = keras.utils.to_categorical(y_test, NUM_CLASSES)
# +
model = Sequential()
#resimlere aktivasyon yapmak zorundayiz cunku ozniteliklerin degerleri cok artiyor, degersizler azaliyor.
# aktivasyondan gecirip degersizleri sifirlayarak onlardan kurtulmaya calisiliyor (negatif degerer sifir olsun).
# 32 tane farkli filtre uygula
model.add(Conv2D(32, kernel_size=(5, 5), activation='relu', input_shape=(IMG_ROWS, IMG_COLS,1)))
# 2x2 pizelleri geziyor, en buyugunu aliyor. gereksiz bilgileri atip boyut dusuruluyor
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Conv2D(64, kernel_size=(5, 5), activation='relu'))
model.add(MaxPool2D(pool_size=(2, 2)))
#model.add(Dropout(0.35))
# duz katman
model.add(Flatten())
# dense=hiddenlayer=fullconnected. 1 gizli katmanda 128 noron var. gizli katman eklemek icin tekrar ayni sekilde add
model.add(Dense(128, activation='relu'))
model.add(Dense(NUM_CLASSES))
# model.add(Dense(NUM_CLASSES, activation='softmax')) #exit layer
# normally it is better to use the code line above but because of
# dependency problems in the current environment, I used the code line below
model.add(Activation(tf.nn.softmax))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
# -
model.fit(X_train, y_train, batch_size=128, epochs=10, verbose=1, validation_data=(X_test, y_test))
model.evaluate(X_test, y_test, verbose=1)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import random
def show_digit(pixels):
img = pixels.reshape(28, 28)
plt.axis('off')
plt.imshow(img, cmap='gray_r')
# +
sample = random.choice(X_test)
show_digit(sample)
sample = sample.reshape(1, 28, 28, 1)
predictions = model.predict(sample)[0]
for i, v in enumerate(predictions):
print("Probability of being %d: %.6f%%" % (i, v * 100))
# -
| Keras CNN Basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## The Flexbox layout
#
# The `HBox` and `VBox` classes above are special cases of the `Box` widget.
#
# The `Box` widget enables the entire CSS flexbox spec as well as the Grid layout spec, enabling rich reactive layouts in the Jupyter notebook. It aims at providing an efficient way to lay out, align and distribute space among items in a container.
#
# Again, the whole flexbox spec is exposed via the `layout` attribute of the container widget (`Box`) and the contained items. One may share the same `layout` attribute among all the contained items.
#
# ### Acknowledgement
#
# The following flexbox tutorial on the flexbox layout follows the lines of the article [A Complete Guide to Flexbox](https://css-tricks.com/snippets/css/a-guide-to-flexbox/) by <NAME>, and uses text and various images from the article [with permission](https://css-tricks.com/license/).
#
# ### Basics and terminology
#
# Since flexbox is a whole module and not a single property, it involves a lot of things including its whole set of properties. Some of them are meant to be set on the container (parent element, known as "flex container") whereas the others are meant to be set on the children (known as "flex items").
# If regular layout is based on both block and inline flow directions, the flex layout is based on "flex-flow directions". Please have a look at this figure from the specification, explaining the main idea behind the flex layout.
#
# 
#
# Basically, items will be laid out following either the `main axis` (from `main-start` to `main-end`) or the `cross axis` (from `cross-start` to `cross-end`).
#
# - `main axis` - The main axis of a flex container is the primary axis along which flex items are laid out. Beware, it is not necessarily horizontal; it depends on the flex-direction property (see below).
# - `main-start | main-end` - The flex items are placed within the container starting from main-start and going to main-end.
# - `main size` - A flex item's width or height, whichever is in the main dimension, is the item's main size. The flex item's main size property is either the ‘width’ or ‘height’ property, whichever is in the main dimension.
# cross axis - The axis perpendicular to the main axis is called the cross axis. Its direction depends on the main axis direction.
# - `cross-start | cross-end` - Flex lines are filled with items and placed into the container starting on the cross-start side of the flex container and going toward the cross-end side.
# - `cross size` - The width or height of a flex item, whichever is in the cross dimension, is the item's cross size. The cross size property is whichever of ‘width’ or ‘height’ that is in the cross dimension.
#
# ### Properties of the parent
#
# 
#
#
# #### display
#
# `display` can be `flex` or `inline-flex`. This defines a flex container (block or inline).
#
# #### flex-flow
#
# `flex-flow` is a shorthand for the `flex-direction` and `flex-wrap` properties, which together define the flex container's main and cross axes. Default is `row nowrap`.
#
# - `flex-direction` (column-reverse | column | row | row-reverse )
#
# This establishes the main-axis, thus defining the direction flex items are placed in the flex container. Flexbox is (aside from optional wrapping) a single-direction layout concept. Think of flex items as primarily laying out either in horizontal rows or vertical columns.
# 
#
# - `flex-wrap` (nowrap | wrap | wrap-reverse)
#
# By default, flex items will all try to fit onto one line. You can change that and allow the items to wrap as needed with this property. Direction also plays a role here, determining the direction new lines are stacked in.
# 
#
# #### justify-content
#
# `justify-content` can be one of `flex-start`, `flex-end`, `center`, `space-between`, `space-around`. This defines the alignment along the main axis. It helps distribute extra free space left over when either all the flex items on a line are inflexible, or are flexible but have reached their maximum size. It also exerts some control over the alignment of items when they overflow the line.
# 
#
# #### align-items
#
# `align-items` can be one of `flex-start`, `flex-end`, `center`, `baseline`, `stretch`. This defines the default behaviour for how flex items are laid out along the cross axis on the current line. Think of it as the justify-content version for the cross-axis (perpendicular to the main-axis).
# 
#
# #### align-content
# `align-content` can be one of `flex-start`, `flex-end`, `center`, `baseline`, `stretch`. This aligns a flex container's lines within when there is extra space in the cross-axis, similar to how justify-content aligns individual items within the main-axis.
# 
#
# **Note**: this property has no effect when there is only one line of flex items.
#
# ### Properties of the items
#
# 
#
# The flexbox-related CSS properties of the items have no impact if the parent element is not a flexbox container (i.e. has a `display` attribute equal to `flex` or `inline-flex`).
#
# #### order
# By default, flex items are laid out in the source order. However, the `order` property controls the order in which they appear in the flex container.
# <img src="./images/order-2.svg" alt="Order" style="width: 500px;"/>
#
# #### flex
# `flex` is shorthand for three properties, `flex-grow`, `flex-shrink` and `flex-basis` combined. The second and third parameters (`flex-shrink` and `flex-basis`) are optional. Default is `0 1 auto`.
#
# - `flex-grow`
#
# This defines the ability for a flex item to grow if necessary. It accepts a unitless value that serves as a proportion. It dictates what amount of the available space inside the flex container the item should take up.
#
# If all items have flex-grow set to 1, the remaining space in the container will be distributed equally to all children. If one of the children a value of 2, the remaining space would take up twice as much space as the others (or it will try to, at least).
# 
#
# - `flex-shrink`
#
# This defines the ability for a flex item to shrink if necessary.
#
# - `flex-basis`
#
# This defines the default size of an element before the remaining space is distributed. It can be a length (e.g. `20%`, `5rem`, etc.) or a keyword. The `auto` keyword means *"look at my width or height property"*.
#
# #### align-self
#
# `align-self` allows the default alignment (or the one specified by align-items) to be overridden for individual flex items.
#
# 
#
# ### The VBox and HBox helpers
#
# The `VBox` and `HBox` helper classes provide simple defaults to arrange child widgets in vertical and horizontal boxes. They are roughly equivalent to:
#
# ```Python
# def VBox(*pargs, **kwargs):
# """Displays multiple widgets vertically using the flexible box model."""
# box = Box(*pargs, **kwargs)
# box.layout.display = 'flex'
# box.layout.flex_flow = 'column'
# box.layout.align_items = 'stretch'
# return box
#
# def HBox(*pargs, **kwargs):
# """Displays multiple widgets horizontally using the flexible box model."""
# box = Box(*pargs, **kwargs)
# box.layout.display = 'flex'
# box.layout.align_items = 'stretch'
# return box
# ```
#
#
# ### Examples
# **Four buttons in a VBox. Items stretch to the maximum width, in a vertical box taking `50%` of the available space.**
# +
from ipywidgets import Layout, Button, Box
items_layout = Layout( width='auto') # override the default width of the button to 'auto' to let the button grow
box_layout = Layout(display='flex',
flex_flow='column',
align_items='stretch',
border='solid',
width='50%')
words = ['correct', 'horse', 'battery', 'staple']
items = [Button(description=word, layout=items_layout, button_style='danger') for word in words]
box = Box(children=items, layout=box_layout)
box
# -
# **Three buttons in an HBox. Items flex proportionally to their weight.**
# +
from ipywidgets import Layout, Button, Box, VBox
# Items flex proportionally to the weight and the left over space around the text
items_auto = [
Button(description='weight=1; auto', layout=Layout(flex='1 1 auto', width='auto'), button_style='danger'),
Button(description='weight=3; auto', layout=Layout(flex='3 1 auto', width='auto'), button_style='danger'),
Button(description='weight=1; auto', layout=Layout(flex='1 1 auto', width='auto'), button_style='danger'),
]
# Items flex proportionally to the weight
items_0 = [
Button(description='weight=1; 0%', layout=Layout(flex='1 1 0%', width='auto'), button_style='danger'),
Button(description='weight=3; 0%', layout=Layout(flex='3 1 0%', width='auto'), button_style='danger'),
Button(description='weight=1; 0%', layout=Layout(flex='1 1 0%', width='auto'), button_style='danger'),
]
box_layout = Layout(display='flex',
flex_flow='row',
align_items='stretch',
width='70%')
box_auto = Box(children=items_auto, layout=box_layout)
box_0 = Box(children=items_0, layout=box_layout)
VBox([box_auto, box_0])
# -
# **A more advanced example: a reactive form.**
#
# The form is a `VBox` of width '50%'. Each row in the VBox is an HBox, that justifies the content with space between..
# +
from ipywidgets import Layout, Button, Box, FloatText, Textarea, Dropdown, Label, IntSlider
form_item_layout = Layout(
display='flex',
flex_flow='row',
justify_content='space-between'
)
form_items = [
Box([Label(value='Age of the captain'), IntSlider(min=40, max=60)], layout=form_item_layout),
Box([Label(value='Egg style'),
Dropdown(options=['Scrambled', 'Sunny side up', 'Over easy'])], layout=form_item_layout),
Box([Label(value='Ship size'),
FloatText()], layout=form_item_layout),
Box([Label(value='Information'),
Textarea()], layout=form_item_layout)
]
form = Box(form_items, layout=Layout(
display='flex',
flex_flow='column',
border='solid 2px',
align_items='stretch',
width='50%'
))
form
# -
# **A more advanced example: a carousel.**
# +
from ipywidgets import Layout, Button, Box, Label
item_layout = Layout(height='100px', min_width='40px')
items = [Button(layout=item_layout, description=str(i), button_style='warning') for i in range(40)]
box_layout = Layout(overflow_x='scroll',
border='3px solid black',
width='500px',
height='',
flex_flow='row',
display='flex')
carousel = Box(children=items, layout=box_layout)
VBox([Label('Scroll horizontally:'), carousel])
# -
# #### *Compatibility note*
#
# The `overflow_x` and `overflow_y` options are deprecated in ipywidgets `7.5`. Instead, use the shorthand property `overflow='scroll hidden'`. The first part specificies overflow in `x`, the second the overflow in `y`.
# ## A widget for exploring layout options
#
# The widgets below was written by ipywidgets user [<NAME> (@DougRzz)](https://github.com/DougRzz). If you want to look through the source code to see how it works, take a look at this [notebook he contributed](cssJupyterWidgetStyling-UI.ipynb).
#
# Use the dropdowns and sliders in the widget to change the layout of the box containing the five colored buttons. Many of the CSS layout optoins described above are available, and the Python code to generate a `Layout` object reflecting the settings is in a `TextArea` in the widget.
# +
#from layout_preview import layout
#layout
# -
| jupyter_interactive_widgets/notebooks/reference_guides/.ipynb_checkpoints/guide-flex-box-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.10 (''venv'': venv)'
# language: python
# name: python3
# ---
# # Visualizing Decision Boundaries
#
# In this notebook, we visualize the decision boundaries of the link classifier based on [this scikit-learn example](https://scikit-learn.org/stable/auto_examples/classification/plot_classifier_comparison.html).
# +
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import SGDClassifier
from sklearn.ensemble import HistGradientBoostingClassifier
from imblearn.ensemble import BalancedBaggingClassifier
import json
from tqdm import tqdm
h = 0.02 # step size in the mesh
names = [
"Logistic Regression",
"Balanced Bagging",
]
classifiers = [
SGDClassifier(loss="log", class_weight="balanced"),
BalancedBaggingClassifier(
base_estimator=HistGradientBoostingClassifier(random_state=42),
n_estimators=10,
random_state=42,
n_jobs=2,
)
]
def make_links():
training_data_path = "../Screen2Vec/simplifiedscreen2vec/data/links.json"
with open(training_data_path) as training_data_json:
initial_training_data = json.load(training_data_json)["links"]
X = []
Y = []
for data_point in tqdm(initial_training_data):
is_link = data_point["isLink"]
source_is_clickable_probability = data_point["link"][
"sourceClickableProbability"
]
source_target_text_similarity = data_point["link"]["sourceTargetTextSimilarity"]
X.append(
[source_is_clickable_probability]
+ [source_target_text_similarity]
)
Y.append(int(is_link))
return np.array(X), np.array(Y)
datasets = [
make_links(),
]
figure = plt.figure(figsize=(9, 3))
i = 1
# iterate over datasets
for ds_cnt, ds in enumerate(datasets):
# preprocess dataset, split into training and test part
X, y = ds
X = StandardScaler().fit_transform(X)
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.4, random_state=42
)
x_min, x_max = X[:, 0].min() - 0.5, X[:, 0].max() + 0.5
y_min, y_max = X[:, 1].min() - 0.5, X[:, 1].max() + 0.5
xx, yy = np.meshgrid(np.arange(x_min, x_max, h), np.arange(y_min, y_max, h))
# just plot the dataset first
cm = plt.cm.RdBu
cm_bright = ListedColormap(["#FF0000", "#0000FF"])
ax = plt.subplot(len(datasets), len(classifiers) + 1, i)
if ds_cnt == 0:
ax.set_title("Input data")
# Plot the training points
ax.scatter(X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright, edgecolors="k")
# Plot the testing points
ax.scatter(
X_test[:, 0], X_test[:, 1], c=y_test, cmap=cm_bright, alpha=0.6, edgecolors="k"
)
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xticks(())
ax.set_yticks(())
ax.set_xlabel("clickable probability")
ax.set_ylabel("text similarity")
i += 1
# iterate over classifiers
for name, clf in zip(names, classifiers):
ax = plt.subplot(len(datasets), len(classifiers) + 1, i)
clf.fit(X_train, y_train)
score = clf.score(X_test, y_test)
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min, x_max]x[y_min, y_max].
if hasattr(clf, "decision_function"):
Z = clf.decision_function(np.c_[xx.ravel(), yy.ravel()])
else:
Z = clf.predict_proba(np.c_[xx.ravel(), yy.ravel()])[:, 1]
# Put the result into a color plot
Z = Z.reshape(xx.shape)
ax.contourf(xx, yy, Z, cmap=cm, alpha=0.8)
# Plot the training points
ax.scatter(
X_train[:, 0], X_train[:, 1], c=y_train, cmap=cm_bright, edgecolors="k"
)
# Plot the testing points
ax.scatter(
X_test[:, 0],
X_test[:, 1],
c=y_test,
cmap=cm_bright,
edgecolors="k",
alpha=0.6,
)
ax.set_xlim(xx.min(), xx.max())
ax.set_ylim(yy.min(), yy.max())
ax.set_xticks(())
ax.set_yticks(())
if ds_cnt == 0:
ax.set_title(name)
ax.text(
xx.max() - 0.3,
yy.min() + 0.3,
("%.2f" % score).lstrip("0"),
size=15,
horizontalalignment="right",
)
i += 1
plt.tight_layout()
plt.show()
# +
import pandas as pd
links_data_path = "../Screen2Vec/simplifiedscreen2vec/data/links.json"
with open(links_data_path) as links_data_json:
links = json.load(links_data_json)["links"]
links_df = pd.DataFrame(pd.json_normalize(links))
links_df.head()
# -
links_df.describe()
links_df.groupby("isLink").describe()
rough_total_sample_size = 100
sample = links_df.groupby('isLink').apply(lambda link_class: link_class.sample(int(rough_total_sample_size/2))).reset_index(drop=True)
sample.head()
sample.describe()
sample.groupby("isLink").describe()
# +
import seaborn as sns
sns.displot(data=sample, x="link.sourceClickableProbability", hue="isLink", kind="kde")
# -
sns.boxplot(data=sample, x="isLink", y="link.sourceClickableProbability")
sns.displot(data=sample, x="link.sourceTargetTextSimilarity", hue="isLink", kind="kde")
sns.boxplot(data=sample, x="isLink", y="link.sourceTargetTextSimilarity")
| visualizing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importando as bibliotecas necessárias
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sb
import os
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import classification_report
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
from collections import Counter
# Caminho do dataset
path = 'C:/Users/Thiago/Documents/CSV/felicidade_2017.csv'
# Criando o dataframe
df = pd.read_csv(path)
# Renomeando as colunas
df.columns = 'Pais Ranking Pontuação wh wl PIB Familia Exp_vida Liberdade Generosidade Confiança_governo Distopia_residual'.split()
# Excluindo duas colunas redundantes
df.drop(['wh','wl','Familia'],axis=1,inplace=True)
# Visualizando os 5 primeiros registros
df.head()
# Scatter plot de duas colunas do dataframe
sb.jointplot(x=df['PIB'],y=df['Pontuação'],kind='scatter');
# Heatmap de correlação entre as variáveis
plt.figure(figsize=(12,8))
sb.heatmap(df.corr(),annot=True,cmap='magma',fmt='.2f')
plt.title('Heatmap de correlação')
plt.xticks(rotation=45)
plt.tight_layout()
# Pair plot das variáveis: Score, PIB e expectativa de vida
sb.pairplot(df,
vars = ['Pontuação','Exp_vida','Confiança_governo'],
diag_kind ='kde',
plot_kws = {'alpha': 0.4, 's': 80, 'edgecolor': 'k'},
size=3.5);
plt.savefig('pairplot.png',dpi=70)
# KDE das variáveis: Score e PIB
sb.jointplot(x=df['Score'],y=df['PIB'],kind='kde');
# +
# Função para classificar os países pelo seu score
# Se score >= media(score) = país feliz
# Senão = país triste
def Rotular(df):
lista_target = []
for score in df['Pontuação']:
if score >= df['Pontuação'].mean():
lista_target.append(1)
else:
lista_target.append(0)
return lista_target
# Criando uma nova coluna no dataframe
df['Target'] = Rotular(df)
# -
# Contagem de países felizes e infelizes
sb.countplot(df['Target'])
plt.title('0 - Infeliz | 1 - Feliz')
plt.ylabel('Contagem')
# +
# Divide os dados em características e varíavel preditiva
# onde X são as características (Score, PIB, etc..)
# e y é o Target
def SplitData():
X = np.array([])
y = np.array([])
X = df.drop(['Pais','Ranking','Target'],axis=1)
y = df['Target']
# Não é necessário, pois os dados já estão na mesma escala
#X = MinMaxScaler().fit_transform(X)
return X,y
X,y = SplitData()
# -
# Dividindo os dados em 70% para treinamento e 30% para teste
X_train,X_test,y_train,y_test = train_test_split(X, y, test_size=0.3)
# Print das divisões
print('Total de amostras:', X.shape[0])
print('Amostras p/ treino:', X_train.shape[0])
print('Amostras p/ teste:', X_test.shape[0])
# Print da distribuição dos países
print('Países felizes/infelizes p/ treino:',Counter(y_train))
print('Paiíses infelizes/felizes p/ teste:',Counter(y_test))
# Definindo o modelo. profundidade da árvore = 3
model = DecisionTreeClassifier(max_depth=3)
# Treinando o modelo
model.fit(X_train,y_train);
# Armazenando as predições
y_pred = model.predict(X_test)
# Comparativo dos valores reais contra as predições
for real,predito in zip(y_test,y_pred):
print('Real:{} | Predito:{}'.format(real,predito))
# Acurácia no treinamento e no teste
print('Acurácia treino:',model.score(X_train,y_train))
print('Acurácia teste:',model.score(X_test,y_test))
# Relatório de classificação
print('| Classification Report |\n')
print(classification_report(y_test,y_test))
# Matriz de confusão
print('| Matriz de confusão |\n')
print(pd.crosstab(y_test, y_pred, rownames=['Real'], colnames=['Predito'], margins=True))
# +
# https://www.linkedin.com/in/thiagomunich
| Ranking-Felicidade/.ipynb_checkpoints/Happiness-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
# default_exp models.deepfm
# -
# # DeepFM
# > A pytorch implementation of DeepFM.
#
# DeepFM consists of an FM component and a deep component which are integrated in a parallel structure. The FM component is the same as the 2-way factorization machines which is used to model the low-order feature interactions. The deep component is a multi-layered perceptron that is used to capture high-order feature interactions and nonlinearities. These two components share the same inputs/embeddings and their outputs are summed up as the final prediction. It is worth pointing out that the spirit of DeepFM resembles that of the Wide & Deep architecture which can capture both memorization and generalization. The advantages of DeepFM over the Wide & Deep model is that it reduces the effort of hand-crafted feature engineering by identifying feature combinations automatically.
#
# 
#hide
from nbdev.showdoc import *
from fastcore.nb_imports import *
from fastcore.test import *
# # v1
# +
#export
import torch
from torch import nn
from recohut.models.layers.embedding import EmbeddingLayer
from recohut.models.layers.common import MLP_Layer, LR_Layer, FM_Layer
from recohut.models.bases.ctr import CTRModel
# -
#export
class DeepFM(CTRModel):
def __init__(self,
feature_map,
model_id="DeepFM",
task="binary_classification",
learning_rate=1e-3,
embedding_initializer="torch.nn.init.normal_(std=1e-4)",
embedding_dim=10,
hidden_units=[64, 64, 64],
hidden_activations="ReLU",
net_dropout=0,
batch_norm=False,
**kwargs):
super(DeepFM, self).__init__(feature_map,
model_id=model_id,
**kwargs)
self.embedding_layer = EmbeddingLayer(feature_map, embedding_dim)
self.fm_layer = FM_Layer(feature_map, output_activation=None, use_bias=False)
self.dnn = MLP_Layer(input_dim=embedding_dim * feature_map.num_fields,
output_dim=1,
hidden_units=hidden_units,
hidden_activations=hidden_activations,
output_activation=None,
dropout_rates=net_dropout,
batch_norm=batch_norm,
use_bias=True)
self.output_activation = self.get_final_activation(task)
self.init_weights(embedding_initializer=embedding_initializer)
def forward(self, inputs):
feature_emb = self.embedding_layer(inputs)
y_pred = self.fm_layer(inputs, feature_emb)
y_pred += self.dnn(feature_emb.flatten(start_dim=1))
if self.output_activation is not None:
y_pred = self.output_activation(y_pred)
return y_pred
# Example
params = {'model_id': 'DeepFM',
'data_dir': '/content/data',
'model_root': './checkpoints/',
'learning_rate': 1e-3,
'optimizer': 'adamw',
'task': 'binary_classification',
'loss': 'binary_crossentropy',
'metrics': ['logloss', 'AUC'],
'embedding_dim': 10,
'hidden_units': [300, 300, 300],
'hidden_activations': 'relu',
'net_regularizer': 0,
'embedding_regularizer': 0,
'batch_norm': False,
'net_dropout': 0,
'batch_size': 64,
'epochs': 3,
'shuffle': True,
'seed': 2019,
'use_hdf5': True,
'workers': 1,
'verbose': 0}
model = DeepFM(ds.dataset.feature_map, **params)
pl_trainer(model, ds, max_epochs=5)
# ## v2
# > **References:-**
# - <NAME>, et al. DeepFM: A Factorization-Machine based Neural Network for CTR Prediction, 2017.
# - https://github.com/rixwew/pytorch-fm/blob/master/torchfm/model/dfm.py
# +
#export
import torch
from recohut.models.layers.common import FeaturesEmbedding, FeaturesLinear, MultiLayerPerceptron
# +
#export
class FactorizationMachine(torch.nn.Module):
def __init__(self, reduce_sum=True):
super().__init__()
self.reduce_sum = reduce_sum
def forward(self, x):
"""
:param x: Float tensor of size ``(batch_size, num_fields, embed_dim)``
"""
square_of_sum = torch.sum(x, dim=1) ** 2
sum_of_square = torch.sum(x ** 2, dim=1)
ix = square_of_sum - sum_of_square
if self.reduce_sum:
ix = torch.sum(ix, dim=1, keepdim=True)
return 0.5 * ix
class DeepFM_v2(torch.nn.Module):
"""
A pytorch implementation of DeepFM.
Reference:
<NAME>, et al. DeepFM: A Factorization-Machine based Neural Network for CTR Prediction, 2017.
"""
def __init__(self, field_dims, embed_dim, mlp_dims, dropout):
super().__init__()
self.linear = FeaturesLinear(field_dims)
self.fm = FactorizationMachine(reduce_sum=True)
self.embedding = FeaturesEmbedding(field_dims, embed_dim)
self.embed_output_dim = len(field_dims) * embed_dim
self.mlp = MultiLayerPerceptron(self.embed_output_dim, mlp_dims, dropout)
def forward(self, x):
"""
:param x: Long tensor of size ``(batch_size, num_fields)``
"""
embed_x = self.embedding(x)
x = self.linear(x) + self.fm(embed_x) + self.mlp(embed_x.view(-1, self.embed_output_dim))
return torch.sigmoid(x.squeeze(1))
# -
# ## v3
# > **References:-**
# - https://github.com/huangjunheng/recommendation_model/tree/master/deepFM
# +
#export
from collections import namedtuple, defaultdict
import torch
from torch import nn as nn
# -
#exporti
class FM(nn.Module):
def __init__(self, p, k):
super(FM, self).__init__()
self.p = p
self.k = k
self.linear = nn.Linear(self.p, 1, bias=True)
self.v = nn.Parameter(torch.Tensor(self.p, self.k), requires_grad=True)
self.v.data.uniform_(-0.01, 0.01)
self.drop = nn.Dropout(0.3)
def forward(self, x):
linear_part = self.linear(x)
inter_part1 = torch.pow(torch.mm(x, self.v), 2)
inter_part2 = torch.mm(torch.pow(x, 2), torch.pow(self.v, 2))
pair_interactions = torch.sum(torch.sub(inter_part1, inter_part2), dim=1)
self.drop(pair_interactions)
output = linear_part.transpose(1, 0) + 0.5 * pair_interactions
return output.view(-1, 1)
#export
class DeepFM_v3(nn.Module):
def __init__(self, feat_sizes, sparse_feature_columns, dense_feature_columns,dnn_hidden_units=[400, 400,400], dnn_dropout=0.0, ebedding_size=4,
l2_reg_linear=0.00001, l2_reg_embedding=0.00001, l2_reg_dnn=0, init_std=0.0001, seed=1024,
device='cpu'):
super(DeepFM_v3, self).__init__()
self.feat_sizes = feat_sizes
self.device = device
self.dense_feature_columns = dense_feature_columns
self.sparse_feature_columns = sparse_feature_columns
self.embedding_size = ebedding_size
self.l2_reg_linear = l2_reg_linear
self.bias = nn.Parameter(torch.zeros((1, )))
self.init_std = init_std
self.dnn_dropout = dnn_dropout
self.embedding_dic = nn.ModuleDict({feat:nn.Embedding(self.feat_sizes[feat], self.embedding_size, sparse=False)
for feat in self.sparse_feature_columns})
for tensor in self.embedding_dic.values():
nn.init.normal_(tensor.weight, mean=0, std=self.init_std)
self.embedding_dic.to(self.device)
self.feature_index = defaultdict(int)
start = 0
for feat in self.feat_sizes:
if feat in self.feature_index:
continue
self.feature_index[feat] = start
start += 1
self.input_size = self.embedding_size * len(self.sparse_feature_columns)+len(self.dense_feature_columns)
# fm
self.fm = FM(self.input_size, 10)
# DNN
self.dropout = nn.Dropout(self.dnn_dropout)
self.hidden_units = [self.input_size] + dnn_hidden_units
self.Linears = nn.ModuleList([nn.Linear(self.hidden_units[i], self.hidden_units[i+1]) for i in range(len(self.hidden_units)-1)])
self.relus = nn.ModuleList([nn.ReLU() for i in range(len(self.hidden_units)-1)])
for name, tensor in self.Linears.named_parameters():
if 'weight' in name:
nn.init.normal_(tensor, mean=0, std=self.init_std)
self.dnn_outlayer = nn.Linear(dnn_hidden_units[-1], 1, bias=False).to(self.device)
def forward(self, x):
# x shape 1024*39
sparse_embedding = [self.embedding_dic[feat](x[:, self.feature_index[feat]].long()) for feat in self.sparse_feature_columns]
sparse_embedding = torch.cat(sparse_embedding, dim=-1)
# print(sparse_embedding.shape) # batch * 208
dense_value = [x[:, self.feature_index[feat]] for feat in
self.dense_feature_columns]
dense_value = torch.cat(dense_value, dim=0)
dense_value = torch.reshape(dense_value, (len(self.dense_feature_columns), -1))
dense_value = dense_value.T
# print(dense_value.shape) # batch * 13
input_x = torch.cat((dense_value, sparse_embedding), dim=1)
# print(input_x.shape) # batch * 221
fm_logit = self.fm(input_x)
for i in range(len(self.Linears)):
fc = self.Linears[i](input_x)
fc = self.relus[i](fc)
fc = self.dropout(fc)
input_x = fc
dnn_logit = self.dnn_outlayer(input_x)
y_pre = torch.sigmoid(fm_logit+dnn_logit+self.bias)
return y_pre
# +
import numpy as np
import pandas as pd
import torch
import torch.nn as nn
from torch.utils.data import DataLoader
from sklearn.metrics import log_loss, roc_auc_score
from recohut.datasets.criteo import CriteoSampleDataset
def get_auc(loader, model):
pred, target = [], []
model.eval()
with torch.no_grad():
for x, y in loader:
x, y = x.to(device).float(), y.to(device).float()
y_hat = model(x)
pred += list(y_hat.cpu().numpy())
target += list(y.cpu().numpy())
auc = roc_auc_score(target, pred)
return auc
root = '/content/data'
batch_size = 1024
epochs = 10
seed = 1024
lr = 0.00005
wd = 0.00001
device = 'cpu'
ds = CriteoSampleDataset(root=root)
train_tensor_data, test_tensor_data = ds.load()
train_loader = DataLoader(train_tensor_data, shuffle=True, batch_size=batch_size)
test_loader = DataLoader(test_tensor_data, batch_size=batch_size)
sparse_features = ['C' + str(i) for i in range(1, 27)]
dense_features = ['I' + str(i) for i in range(1, 14)]
# model = NFM(ds.feat_sizes, embedding_size, ds.linear_feature_columns, ds.dnn_feature_columns).to(device)
model = DeepFM_v3(ds.feat_sizes, sparse_feature_columns=sparse_features, dense_feature_columns=dense_features,
dnn_hidden_units=[1000, 500, 250], dnn_dropout=0.9, ebedding_size=16,
l2_reg_linear=1e-3, device=device)
loss_func = nn.BCELoss(reduction='mean')
optimizer = torch.optim.Adam(model.parameters(), lr=lr, weight_decay=wd)
for epoch in range(epochs):
total_loss_epoch = 0.0
total_tmp = 0
model.train()
for index, (x, y) in enumerate(train_loader):
x, y = x.to(device).float(), y.to(device).float()
y_hat = model(x)
optimizer.zero_grad()
loss = loss_func(y_hat, y)
loss.backward()
optimizer.step()
total_loss_epoch += loss.item()
total_tmp += 1
auc = get_auc(test_loader, model)
print('epoch/epoches: {}/{}, train loss: {:.3f}, test auc: {:.3f}'.format(epoch, epochs, total_loss_epoch / total_tmp, auc))
# +
# class DeepFM(PointModel):
# def __init__(self, n_users, n_items, embedding_dim, batch_norm=True, dropout=0.1, num_layers=3, act_function='relu'):
# """
# Args:
# n_users : int, the number of users
# n_items : int, the number of items
# embedding_dim : int, the number of latent factoact_function : str, activation function for hidden layer
# num_layers : int, number of hidden layers
# batch_norm : bool, whether to normalize a batch of data
# dropout : float, dropout rate
# """
# super().__init__()
# self.num_layers = num_layers
# self.user_embedding = nn.Embedding(
# num_embeddings=n_users, embedding_dim=embedding_dim
# )
# self.item_embedding = nn.Embedding(
# num_embeddings=n_items, embedding_dim=embedding_dim
# )
# self.user_bias = nn.Embedding(n_users, 1)
# self.item_bias = nn.Embedding(n_items, 1)
# self.bias_ = nn.Parameter(torch.tensor([0.0]))
# fm_modules = []
# if batch_norm:
# fm_modules.append(nn.BatchNorm1d(embedding_dim))
# fm_modules.append(nn.Dropout(dropout))
# self.fm_layers = nn.Sequential(*fm_modules)
# deep_modules = []
# in_dim = embedding_dim * 2 # user & item
# for _ in range(num_layers): # _ is dim if layers is list
# out_dim = in_dim
# deep_modules.append(nn.Linear(in_dim, out_dim))
# in_dim = out_dim
# if batch_norm:
# deep_modules.append(nn.BatchNorm1d(out_dim))
# if act_function == 'relu':
# deep_modules.append(nn.ReLU())
# elif act_function == 'sigmoid':
# deep_modules.append(nn.Sigmoid())
# elif act_function == 'tanh':
# deep_modules.append(nn.Tanh())
# deep_modules.append(nn.Dropout(dropout))
# self.deep_layers = nn.Sequential(*deep_modules)
# self.deep_out = nn.Linear(in_dim, 1, bias=False)
# self._init_weights()
# def _init_weights(self):
# nn.init.normal_(self.item_embedding.weight, std=0.01)
# nn.init.normal_(self.user_embedding.weight, std=0.01)
# nn.init.constant_(self.user_bias.weight, 0.0)
# nn.init.constant_(self.item_bias.weight, 0.0)
# # for deep layers
# for m in self.deep_layers:
# if isinstance(m, nn.Linear):
# nn.init.xavier_normal_(m.weight)
# nn.init.xavier_normal_(self.deep_out.weight)
# def forward(self, users, items):
# embed_user = self.user_embedding(users)
# embed_item = self.item_embedding(items)
# fm = embed_user * embed_item
# fm = self.fm_layers(fm)
# y_fm = fm.sum(dim=-1)
# y_fm = y_fm + self.user_bias(users) + self.item_bias(items) + self.bias_
# if self.num_layers:
# fm = self.deep_layers(fm)
# y_deep = torch.cat((embed_user, embed_item), dim=-1)
# y_deep = self.deep_layers(y_deep)
# # since BCELoss will automatically transfer pred with sigmoid
# # there is no need to use extra nn.Sigmoid(pred)
# pred = y_fm + y_deep
# return pred.view(-1)
# -
#hide
# %reload_ext watermark
# %watermark -a "Sparsh A." -m -iv -u -t -d -p recohut
| nbs/models/models.deepfm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: demo
# language: python
# name: demo
# ---
# #### Import all the required packages
# +
## basic packages
import numpy as np
import re
import csv
import time
import pandas as pd
from itertools import product
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('darkgrid')
##gensim
import gensim
import gensim.corpora as corpora
from gensim.utils import simple_preprocess
from gensim.models import CoherenceModel
##spacy and nltk
import spacy
from nltk.corpus import stopwords
from spacy.lang.en.stop_words import STOP_WORDS
##vis
# import pyLDAvis
# import pyLDAvis.gensim_models as gensimvis
# pyLDAvis.enable_notebook()
import warnings
warnings.filterwarnings("ignore",category=DeprecationWarning)
# -
# #### load the metadata of podcast transcripts
# +
global df, show_descriptions
meta_data = []
with open("../data/metadata.tsv") as csvfile:
csvreader = csv.reader(csvfile,delimiter="\t")
for row in csvreader:
meta_data.append(row)
df = pd.DataFrame(meta_data[1:],columns=meta_data[0])
show_filename_prefixes = df.show_filename_prefix
episode_filename_prefixes = df.episode_filename_prefix
shows = df.groupby(by=['show_name'])
show_names = shows.apply(lambda x: x.show_name.unique()[0])
# +
genres_topics = ["comedy","news","crime","science","economics","politics","education",\
"sports","lifestyle","health","wellbeing","religion","faith","music",\
"art","fashion","literature","humanities","drama","fitness","drama",\
"fantasy","scifi","gameshow","news quiz","games","game","mental",\
"humor","research","technology","society","social","culture","lifestyle",\
"songs","cooking","culinary","food","travel","films","film","movies","tv",\
"climate","space","planet","digital","artificial intelligence", "ai",\
"cars","car","nutrition","wellness","family","history","geography","physics",\
"mathematics","math","chemistry","biology","documentary","commentary","nfl",\
"mls","nba","mlb","stocks","stock","market","wall","street","wallstreet","business",\
"reality","shows","investing","social media","biography","biographies",\
"data science","medicine","media","books","book","europe","asia","canada",\
"south america","north america","america","usa","netflix","adventure","pets","dogs",\
"cats","dog","cat","nintendo","xbox","playstation","ps4","ps5","theatre","mars"\
"tennis","australia","conspiracy","war","epidemic","pandemic","climate","change"\
"astrology","novel","church","christ","romance","english","kids","astronomy"\
"design","entrepreneurship","marketing","digital","christian","christianity","boardgames",\
"boardgame","videogames","videogame","twitch","currency","cryptocurrency","federal","bank",\
"blockchain","bitcoin","nyse","nft","import","export","capital","money","exchange","boxing",\
"mma","wrestling","excercise","excercises","gym","bodybuilding","body-building","yoga",\
"stamina","strength","calories","meditation","physical","healthy","pope","bible","catholic",\
"catholicism","testament"]
formats = ["monologue","interview","storytelling","repurposed",\
"bite-sized","co-host conversation","debate","narrative",\
"scripted","improvised"]
# -
podcasts_genres_topics = {}
for k,show in enumerate(show_names):
keywords = show.lower().split(" ")
for word in keywords:
if word in genres_topics:
if (k,show) in podcasts_genres_topics:
if word not in podcasts_genres_topics[(k,show)]:
podcasts_genres_topics[(k,show)].append(word)
else:
podcasts_genres_topics[(k,show)] = [word]
podcasts = [item[1] for item in podcasts_genres_topics.keys()]
nlp = spacy.load("en_core_web_sm")
stops_nltk = set(stopwords.words("english"))
stops_spacy = STOP_WORDS.union({'ll', 've', 'pron','okay','oh','like','know','yea','yep','yes','no',\
"like","oh","yeah","okay","wow","podcast","rating","ratings","not",\
"support","anchor","podcasts","episode","http","https","5star","reviews",\
"review","instagram","tiktok","amazon","apple","twitter","goole",\
"facebook","send","voice message","message","voice","subscribe","follow",\
"sponsor","links","easiest","way","fuck","fucking","talk","discuss",\
"world","time","want","join","learn","week","things","stuff","find",\
"enjoy","welcome","share","talk","talking","people","gmail","help","today",\
"listen","best","stories","story","hope","tips","great","journey",\
"topics","email","questions","question","going","life","good","friends",\
"friend","guys","discussing","live","work","student","students","need",\
"hear","think","change","free","better","little","fucking","fuck","shit",\
"bitch","sex","easiest","way","currently","follow","follows","needs",\
"grow","stay","tuned","walk","understand","tell","tells","ask","helps",\
"feel","feels","look","looks","meet","relate","soon","quick","dude","girl",\
"girls","guy","literally","spotify","google","totally","played","young",\
"begin","began","create","month","year","date","day","terms","lose","list",\
"bought","brings","bring","buy","percent","rate","increase","words","value",\
"search","awesome","followers","finn","jake","mark","america","american",\
"speak","funny","hours","hour","honestly","states","united","franklin",\
"patrick","john","build","dave","excited","process","processes","based",\
"focus","star","mary","chris","taylor","gotta","liked","hair","adam","chat",\
"named","died","born","country","mother","father","children","tools",\
"countries","jordan","tommy","listeners","water","jason","lauren","alex",\
"laguna","jessica","kristen","examples","example","heidi","stephen","utiful",\
"everybody","sorry","came","come","meet","whoa","whoaa","yay","whoaw",\
"anybody","somebody","cool","watch","nice","shall"})
stops = stops_nltk.union(stops_spacy)
health_keywords = ["fitness","health","diet","nutrition","healthy","meditation","mental",\
"physical","excercise","calories","gym","bodybuilding","body-building",\
"stamina","strength","excercise","yoga","wellness"]
health_category = [(key,val) for key,val in podcasts_genres_topics.items() if ("fitness" in val)\
or ("health" in val)\
or ("diet" in val)\
or ("nutrition" in val)\
or ("healthy" in val)\
or ("meditation" in val)\
or ("mental" in val)\
or ("physical" in val)\
or ("excercise" in val)\
or ("calories" in val)\
or ("gym" in val)\
or ("bodybuilding" in val)\
or ("body-building" in val)\
or ("stamina" in val)\
or ("strength" in val)\
or ("excercise" in val)\
or ("yoga" in val)\
or ("wellness" in val)\
or ("wellbeing" in val)]
# +
d = {}
for val in podcasts_genres_topics.values():
for word in health_keywords:
if word in val:
if word in d:
d[word] += 1
else:
d[word] = 1
plt.figure(figsize=(10,8))
plt.bar(d.keys(),d.values())
plt.title('Distribution of podcast episodes related to health/fitness',fontsize=16)
plt.xlabel('Keyword',fontsize=16)
plt.ylabel('Keyword frequency',fontsize=16)
plt.xticks(rotation=90,fontsize=14)
plt.yticks(fontsize=14);
# +
number_of_topics = [5,6,7,8,9,10,15]
df_parameters = list(product([2,3,4,5,6,7,8,9,10],[0.3,0.4,0.5,0.6,0.7,0.8,0.9]))
hyperparams = list(product(number_of_topics,df_parameters))
sports_cs = []
with open('/home1/sgmark/capstone-project/model/coherence_scores_health_category.csv','r') as f:
reader = csv.reader(f)
for row in reader:
sports_cs.append([float(x) for x in row])
best_hp_setting = hyperparams[np.argmax([x[4] for x in sports_cs])]
# -
# #### The individual transcript location
def file_location(show,episode):
search_string = local_path + "/spotify-podcasts-2020" + "/podcasts-transcripts" \
+ "/" + show[0] \
+ "/" + show[1] \
+ "/" + "show_" + show \
+ "/"
return search_string
# #### load the transcripts
# +
transcripts = {}
for podcast,genre in health_category:
for i in shows.get_group(podcast[1])[['show_filename_prefix','episode_filename_prefix']].index:
show,episode = shows.get_group(podcast[1])[['show_filename_prefix','episode_filename_prefix']].loc[i]
s = show.split("_")[1]
try:
with open('../podcast_transcripts/'+s[0]+'/'+s[1]+'/'+show+'/'+episode+'.txt','r') as f:
transcripts[(show,episode)] = f.readlines()
f.close()
except Exception as e:
pass
keys = list(transcripts.keys())
# +
# Cleaning & remove urls and links
def remove_stops(text,stops):
final = []
for word in text:
if (word not in stops) and (len(word)>3) and (not word.endswith('ing')) and (not word.endswith('ly')):
final.append(word)
return final
def clean_text(docs):
final = []
for doc in docs:
clean_doc = remove_stops(doc, stops)
final.extend(clean_doc)
return final
def lemmatization(text_data):
nlp = spacy.load("en_core_web_sm")
texts = []
for text in text_data:
doc = nlp(text)
lem_text = []
for token in doc:
if (token.pos_=="VERB") or (token.pos_=="ADV"):
pass
else:
lem_text.append(token.lemma_)
texts.append(lem_text)
return texts
# -
# #### tokenize/convert text into words
def normalize_docs(text_data):
final_texts = []
for text in text_data:
new_text = gensim.utils.simple_preprocess(text,deacc=True)
final_texts.append(new_text)
return final_texts
# +
docs = []
for text in transcripts.values():
docs.append(' '.join(clean_text(normalize_docs(text))))
texts = lemmatization(docs)
texts = [remove_stops(text,stops) for text in texts]
# -
# ### Using bigrams
# +
from gensim.models.phrases import Phrases
bigram = Phrases(texts, min_count=best_hp_setting[1][0])
for i in range(len(texts)):
for token in bigram[texts[i]]:
if '_' in token:
texts[i].append(token)
# -
# #### Construct a corpus of words as a bag of words
# +
dictionary = corpora.Dictionary(texts)
dictionary.filter_extremes(no_below=best_hp_setting[1][0],no_above=best_hp_setting[1][1])
# -
corpus = [dictionary.doc2bow(text) for text in texts]
# #### Hyperparameter tuning
# +
# from itertools import product
# number_of_topics = [5,6,7,8,9,10,15]
# df_parameters = list(product([2,3,4,5,6,7,8,9,10],[0.3,0.4,0.5,0.6,0.7,0.8,0.9]))
# coherence_scores_umass = np.zeros((len(number_of_topics),len(df_parameters)))
# coherence_scores_uci = np.zeros((len(number_of_topics),len(df_parameters)))
# coherence_scores_npmi = np.zeros((len(number_of_topics),len(df_parameters)))
# j = 0
# for num in number_of_topics:
# i = 0
# for n,m in df_parameters:
# dictionary = corpora.Dictionary(texts)
# dictionary.filter_extremes(no_below=n,no_above=m)
# corpus = [dictionary.doc2bow(text) for text in texts]
# num_topics = num
# chunksize = 200
# passes = 20
# iterations = 500
# eval_every = None
# lda_model = gensim.models.ldamodel.LdaModel(corpus,
# id2word=dictionary,
# num_topics=num_topics,
# chunksize=chunksize,
# passes=passes,
# iterations=iterations,
# alpha='auto',
# eta='auto',
# random_state = 123,
# eval_every=eval_every)
# cm = CoherenceModel(lda_model, texts=texts,corpus=corpus, coherence= 'c_uci')
# coherence_scores_uci[j,i] = cm.get_coherence()
# cm = CoherenceModel(lda_model, texts=texts,corpus=corpus, coherence= 'c_npmi')
# coherence_scores_npmi[j,i] = cm.get_coherence()
# cm = CoherenceModel(lda_model, corpus=corpus, coherence= 'u_mass')
# coherence_scores_umass[j,i] = cm.get_coherence()
# with open("coherence_scores_health_category.csv",'a') as f:
# writer = csv.writer(f)
# writer.writerow([num,n,m,coherence_scores_uci[j,i],coherence_scores_npmi[j,i],\
# coherence_scores_umass[j,i]])
# i += 1
# print(i)
# j += 1
# print(j)
# -
# #### Final model
# +
# %%time
import logging
logging.basicConfig(filename='health_topics.log', encoding='utf-8',format='%(asctime)s : %(levelname)s : %(message)s', level=logging.DEBUG)
num_topics = best_hp_setting[0]
chunksize = 200
passes = 50
iterations = 500
eval_every = None
lda_model = gensim.models.ldamodel.LdaModel(corpus,
id2word=dictionary,
num_topics=num_topics,
chunksize=chunksize,
passes=passes,
iterations=iterations,
alpha='auto',
eta='auto',
random_state=123,
eval_every=eval_every)
# +
top_topics = lda_model.top_topics(corpus,texts=texts,coherence='c_npmi') #, num_words=20)
# Average topic coherence is the sum of topic coherences of all topics, divided by the number of topics.
avg_topic_coherence = sum([t[1] for t in top_topics])/num_topics
print('Average topic coherence: %.4f.' % avg_topic_coherence)
print(f'topic coherence scores: {[t[1] for t in top_topics]}')
# -
# ### Visualizing data
# +
# vis = pyLDAvis.gensim_models.prepare(lda_model,corpus,dictionary,mds="mmds",R=10)
# vis
# +
# from pprint import pprint
# pprint(top_topics)
# +
import pickle
pickle.dump(lda_model,open('../model/health_episodes_lda_model.pkl','wb'))
pickle.dump(dictionary,open('../model/health_episodes_dictionary.pkl','wb'))
pickle.dump(corpus,open('../model/health_episodes_corpus.pkl','wb'))
pickle.dump(texts,open('../model/health_episodes_texts.pkl','wb'))
# +
# import pickle
# file = open('../model/health_episodes_lda_model.pkl','rb')
# lda_model = pickle.load(file)
# file.close()
# file = open('../model/health_episodes_corpus.pkl','rb')
# corpus = pickle.load(file)
# file.close()
# file = open('../model/health_episodes_dictionary.pkl','rb')
# dictionary = pickle.load(file)
# file.close()
# -
def get_main_topic_df(model, bow, texts):
topic_list = []
percent_list = []
keyword_list = []
podcast_list = []
episode_list = []
duration_list = []
publisher_list = []
for key,wc in zip(keys,bow):
podcast_list.append(df[(df['show_filename_prefix'] == key[0])&(df['episode_filename_prefix'] == key[1])].show_name.iloc[0])
episode_list.append(df[(df['show_filename_prefix'] == key[0])&(df['episode_filename_prefix'] == key[1])].episode_name.iloc[0])
duration_list.append(round(float(df[(df['show_filename_prefix'] == key[0])&(df['episode_filename_prefix'] == key[1])].duration.iloc[0])))
publisher_list.append(df[(df['show_filename_prefix'] == key[0])&(df['episode_filename_prefix'] == key[1])].publisher.iloc[0])
topic, percent = sorted(model.get_document_topics(wc), key=lambda x: x[1], reverse=True)[0]
topic_list.append(topic)
percent_list.append(round(percent, 3))
keyword_list.append(' '.join(sorted([x[0] for x in model.show_topic(topic)])))
result_df = pd.concat([pd.Series(podcast_list, name='Podcast_name'),
pd.Series(episode_list, name='Episode_name'),
pd.Series(topic_list, name='Dominant_topic'),
pd.Series(percent_list, name='Percent'),
pd.Series(texts, name='Processed_text'),
pd.Series(keyword_list, name='Keywords'),
pd.Series(duration_list, name='Duration of the episode'),
pd.Series(publisher_list, name='Publisher of the show')], axis=1)
return result_df
main_topic_df = get_main_topic_df(lda_model,corpus,texts)
main_topic_df.to_pickle('health_topics_main_df.pkl')
main_topic_df.head(5)
plt.figure(figsize=(10,8))
topics_groups = main_topic_df.groupby('Dominant_topic')
plt.bar(range(best_hp_setting[0]),topics_groups.count()['Podcast_name'],width=0.5)
plt.title('Dominant topic frequency in the investment/stocks category of podcast episodes',fontsize=16)
plt.xlabel('Dominant Topic index',fontsize=16)
plt.ylabel('Number of episodes',fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14);
# +
representatives = pd.DataFrame()
for k in topics_groups.groups.keys():
representatives = pd.concat([representatives,
topics_groups.get_group(k).sort_values(['Percent'], ascending=False).head(1)])
for k,words in enumerate(representatives.Keywords):
print(f'topic {k}: {words}')
# +
# print('Document: {} Dominant topic: {}\n'.format(representatives.index[2],
# representatives.loc[representatives.index[2]]['Dominant_topic']))
# print([sentence.strip() for sentence in transcripts[keys[representatives.index[2]]]])
# -
num_topics = best_hp_setting[0]
def word_count_by_topic(topic=0):
d_lens = [len(d) for d in topics_groups.get_group(topic)['Processed_text']]
plt.figure(figsize=(10,8))
plt.hist(d_lens)
large = plt.gca().get_ylim()[1]
d_mean = round(np.mean(d_lens), 1)
d_median = np.median(d_lens)
plt.plot([d_mean, d_mean], [0,large], label='Mean = {}'.format(d_mean))
plt.plot([d_median, d_median], [0,large], label='Median = {}'.format(d_median))
plt.legend()
plt.xlabel('Document word count',fontsize=16)
plt.ylabel('Number of documents',fontsize=16)
plt.xticks(fontsize=14)
plt.yticks(fontsize=14)
# +
from ipywidgets import interact, IntSlider
slider = IntSlider(min=0, max=num_topics-1, step=1, value=0, description='Topic')
interact(word_count_by_topic, topic=slider);
# -
lda_top_words_index = set()
for i in range(lda_model.num_topics):
lda_top_words_index = lda_top_words_index.union([k for (k,v) in lda_model.get_topic_terms(i)])
#print('Indices of top words: \n{}\n'.format(lda_top_words_index))
words_we_care_about = [{dictionary[tup[0]]: tup[1] for tup in lst if tup[0] in list(lda_top_words_index)}
for lst in corpus]
lda_top_words_df = pd.DataFrame(words_we_care_about).fillna(0).astype(int).sort_index(axis=1)
lda_top_words_df['Cluster'] = main_topic_df['Dominant_topic']
k=1
clusterwise_words_dist = lda_top_words_df.groupby('Cluster').get_group(k)
plt.figure(figsize=(30,8))
plt.bar(list(clusterwise_words_dist.sum()[:-1].transpose().index),\
list(clusterwise_words_dist.sum()[:-1].transpose()))
plt.title(f'Term frequencies of keywords of topic: {k}',fontsize=16)
plt.xlabel('Keywords in the topics',fontsize=16)
plt.ylabel('Word frequency',fontsize=16)
plt.xticks(rotation=90,fontsize=14)
plt.yticks(fontsize=14);
word_totals = {k:{y[1]:y[0] for y in x[0]} for k,x in enumerate(top_topics)}
# +
import matplotlib.pyplot as plt
from ipywidgets import interact, IntSlider
from wordcloud import WordCloud
def show_wordcloud(topic=0):
cloud = WordCloud(background_color='white', colormap='viridis')
cloud.generate_from_frequencies(word_totals[topic])
plt.figure(figsize=(10,8))
plt.gca().imshow(cloud)
plt.axis('off')
plt.tight_layout()
slider = IntSlider(min=0, max=best_hp_setting[0]-1, step=1, value=0, description='Topic')
interact(show_wordcloud, topic=slider);
# -
| model/.ipynb_checkpoints/LDA_model-health-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from tqdm.notebook import tqdm
import numpy as np
from collections import Counter
from sklearn.metrics import f1_score, accuracy_score as acc, precision_score as prec, recall_score as rec
import torch
from transformers import BertForSequenceClassification, BertTokenizer, XLNetForSequenceClassification, XLNetTokenizer, RobertaForSequenceClassification, RobertaTokenizer, AdamW, get_linear_schedule_with_warmup
from torch.utils.data import TensorDataset, DataLoader, RandomSampler, SequentialSampler
import random
# Set random seeds for reproducibility on a specific machine
torch.backends.cudnn.benchmark = False
torch.backends.cudnn.deterministic = True
torch.manual_seed(1)
torch.cuda.manual_seed(1)
random.seed(1)
np.random.seed(1)
np.random.RandomState(0)
# +
train = pd.read_csv('../annotated_data/train.tsv', sep='\t', header=0)
train['Sentence'] = train['Sentence'].apply(lambda x: x.lower())
train_sentences = train['Sentence'].tolist()
train_labels_DS = train['DS_Label'].values
train_labels_Maj = train['Majority_label'].values
dev = pd.read_csv('../annotated_data/dev.tsv', sep='\t', header=0)
dev['Sentence'] = dev['Sentence'].apply(lambda x: x.lower())
dev_sentences = dev['Sentence'].tolist()
dev_labels_DS = dev['DS_Label'].values
dev_labels_Maj = dev['Majority_label'].values
test = pd.read_csv('../annotated_data/test.tsv', sep='\t', header=0)
test['Sentence'] = test['Sentence'].apply(lambda x: x.lower())
test_sentences = test['Sentence'].tolist()
test_labels_DS = test['DS_Label'].values
test_labels_Maj = test['Majority_label'].values
# -
# Creating new columns that might help later on with the sequence labelling task:
# 1. *Post.ID* The reddit post from which the replies are scraped -
# 2. *Reply.ID* The id number of the reply, as ordered by reddit's best algorithm (need to check up on this with Ben)
# 3. *Sent.Num* The sentence number, in order, from within the reply
# +
train['Post.ID'] = train['ID'].apply(lambda x: x.split('-')[0])
train['Reply.ID'] = train['ID'].apply(lambda x: x.split('-')[1])
train['Sent.Num'] = train['ID'].apply(lambda x: x.split('-')[2])
dev['Post.ID'] = dev['ID'].apply(lambda x: x.split('-')[0])
dev['Reply.ID'] = dev['ID'].apply(lambda x: x.split('-')[1])
dev['Sent.Num'] = dev['ID'].apply(lambda x: x.split('-')[2])
test['Post.ID'] = test['ID'].apply(lambda x: x.split('-')[0])
test['Reply.ID'] = test['ID'].apply(lambda x: x.split('-')[1])
test['Sent.Num'] = test['ID'].apply(lambda x: x.split('-')[2])
train.set_index('ID',inplace=True)
train.head()
# +
# def BIO_convert(array):
# '''
# Convert a sequence of 1s and 0s to BIO(Beginning-Inside-Outside) format
# '''
# bio = ['O' for i in range(len(array))]
# if 1 not in array:
# return bio
# else:
# bio[array.index(1)] = 'B'
# for k in range(array.index(1)+1, len(array)):
# if array[k] == 1 and bio[k-1] == 'B':
# bio[k] = 'I'
# elif array[k] == 0:
# bio[k] = 'O'
# elif array[k] == 1 and array[k-1] == 0:
# bio[k] = 'B'
# return bio
# +
# MODELS = {'bert': (BertForSequenceClassification, BertTokenizer,
# 'bert-base-cased'),
# 'xlnet': (XLNetForSequenceClassification, XLNetTokenizer,
# 'xlnet-base-cased'),
# 'roberta': (RobertaForSequenceClassification, RobertaTokenizer,
# 'roberta-base')}
# -
# ## Normal Sequence Classification with Fine tuned BERT (DS labels)
# +
# # Set the maximum length of sequence - just the longest length sentence from
# # train, test and dev
# MAX_LEN = max(max([len(a.split()) for a in train_sentences]),
# max([len(a.split()) for a in dev_sentences]),
# max([len(a.split()) for a in test_sentences]))
# # Select a batch size for training. For fine-tuning xlnet on a specific task, the authors recommend a batch size of 16 or 32
# batch_size = 32
# # Choose gpu or cpu
# if torch.cuda.is_available():
# device = torch.device('cuda:0')
# else:
# device = torch.device('cpu')
# # How many labels in your problem?
# num_labels = np.unique(train_labels_DS).shape[0]
# # Load tokenizer XForSequenceClassification model, the pretrained args.model
# # with a single linear classification layer on top
# tokenizer = MODELS['xlnet'][1].from_pretrained(MODELS['xlnet'][2])
# model = MODELS['xlnet'][0].from_pretrained(MODELS['xlnet'][2], num_labels=num_labels).to(device)
# +
# model = XLNetForSequenceClassification.from_pretrained("xlnet-base-cased", num_labels=2,
# hidden_dropout_prob=0.5).to(device)
# +
# Convert our text into tokens that corresponds to BERT library
# train_input_ids = [tokenizer.encode(sent, add_special_tokens=True,max_length=MAX_LEN,pad_to_max_length=True)
# for sent in train_sentences]
# train_input_ids = torch.tensor(train_input_ids)
# # Create a mask of 1 for all input tokens and 0 for all padding tokens
# train_attention_masks = [[float(i>0) for i in seq] for seq in train_input_ids]
# train_attention_masks = torch.tensor(train_attention_masks)
# train_labels_DS = torch.tensor(train_labels_DS)
# # Same for dev
# dev_input_ids = [tokenizer.encode(sent, add_special_tokens=True,max_length=MAX_LEN,pad_to_max_length=True)
# for sent in dev_sentences]
# dev_input_ids = torch.tensor(dev_input_ids)
# dev_attention_masks = [[float(i>0) for i in seq] for seq in dev_input_ids]
# dev_attention_masks = torch.tensor(dev_attention_masks)
# dev_labels_DS = torch.tensor(dev_labels_DS)
# print(train_input_ids.shape, dev_input_ids.shape, train_attention_masks.shape, dev_attention_masks.shape)
# +
# Create an iterator of our data with torch DataLoader. This helps save on memory during training because, unlike a for loop, with an iterator the entire dataset does not need to be loaded into memory
# train_data = TensorDataset(train_input_ids[:32],train_attention_masks[:32],train_labels_DS[:32])
# train_sampler = RandomSampler(train_data)
# train_dataloader = DataLoader(train_data,sampler=train_sampler,batch_size=batch_size)
# dev_data = TensorDataset(dev_input_ids[:32],dev_attention_masks[:32],dev_labels_DS[:32])
# dev_sampler = RandomSampler(dev_data)
# dev_dataloader = DataLoader(dev_data,sampler=dev_sampler,batch_size=batch_size)
# +
# Load BertForSequenceClassification, the pretrained BERT model with a single linear classification layer on top
# model = BertForSequenceClassification.from_pretrained("bert-base-cased", num_labels=2,
# hidden_dropout_prob=0.5).to(device)
# # Parameters:
# lr = 2e-7
# adam_epsilon = 1e-8
# # Number of training epochs (authors recommend between 2 and 4)
# epochs = 6
# num_warmup_steps = 0
# num_training_steps = len(train_dataloader)*epochs
# ### In Transformers, optimizer and schedules are splitted and instantiated like this:
# optimizer = AdamW(model.parameters(), lr=lr,eps=adam_epsilon,correct_bias=False) # To reproduce BertAdam specific behavior set correct_bias=False
# scheduler = get_linear_schedule_with_warmup(optimizer, num_warmup_steps=num_warmup_steps, num_training_steps=num_training_steps) # PyTorch scheduler
# -
# ### Training/Fine-tuning step
# +
# Store our loss and accuracy for plotting
# train_loss_set = []
# learning_rate = []
# validation_acc = [0]
# # Gradients gets accumulated by default
# model.zero_grad()
# for _ in range(1,epochs+1):
# print("<" + "="*22 + F" Epoch {_} "+ "="*22 + ">")
# # Calculate total loss for this epoch
# batch_loss = 0
# for step, batch in enumerate(tqdm(train_dataloader)):
# # Set our model to training mode (as opposed to evaluation mode)
# model.train()
# # Add batch to GPU
# batch = tuple(t.to(device) for t in batch)
# # Unpack the inputs from our dataloader
# b_input_ids, b_input_mask, b_labels = batch
# from IPython.core.debugger import set_trace;set_trace()
# # Forward pass
# outputs = model(b_input_ids, token_type_ids=None, attention_mask=b_input_mask, labels=b_labels)
# loss = outputs[0]
# # Backward pass
# loss.backward()
# # Clip the norm of the gradients to 1.0
# # Gradient clipping is not in AdamW anymore
# torch.nn.utils.clip_grad_norm_(model.parameters(), 1.0)
# # Update parameters and take a step using the computed gradient
# optimizer.step()
# # Update learning rate schedule
# scheduler.step()
# # Clear the previous accumulated gradients
# optimizer.zero_grad()
# # Update tracking variables
# batch_loss += loss.item()
# # Calculate the average loss over the training data.
# avg_train_loss = batch_loss / len(train_dataloader)
# #store the current learning rate
# for param_group in optimizer.param_groups:
# print("\n\tCurrent Learning rate: ",param_group['lr'])
# learning_rate.append(param_group['lr'])
# train_loss_set.append(avg_train_loss)
# print(F'\n\tAverage Training loss: {avg_train_loss}')
# # Put model in evaluation mode to evaluate loss on the validation set
# model.eval()
# # Tracking variables
# eval_accuracy, eval_f1_score, nb_eval_steps = 0, 0, 0
# for batch in dev_dataloader:
# # Add batch to GPU
# batch = tuple(t.to(device) for t in batch)
# # Unpack the inputs from our dataloader
# b_input_ids, b_input_mask, b_labels = batch
# # Telling the model not to compute or store gradients, saving memory and speeding up validation
# with torch.no_grad():
# # Forward pass, calculate logit predictions
# (logits,) = model(b_input_ids, token_type_ids=None, attention_mask=b_input_mask)
# # Move logits and labels to CPU
# logits = logits.to('cpu').numpy()
# label_ids = b_labels.to('cpu').numpy()
# pred_flat = np.argmax(logits, axis=1).flatten()
# labels_flat = label_ids.flatten()
# tmp_eval_accuracy = acc(pred_flat, labels_flat)
# tmp_eval_f1_score = f1_score(labels_flat, pred_flat)
# eval_accuracy += tmp_eval_accuracy
# eval_f1_score += tmp_eval_f1_score
# nb_eval_steps += 1
# validation_acc.append(eval_accuracy/nb_eval_steps)
# print(F'\n\tValidation Accuracy: {eval_accuracy/nb_eval_steps}')
# # print(F'\n\tValidation MCC Accuracy: {eval_mcc_accuracy/nb_eval_steps}')
# print(F'\n\tValidation F1 Score: {eval_f1_score/nb_eval_steps}')
# # if valid_acc[-1] < valid_acc[-2]:
# # break
# -
| Notebooks/Advice_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [Day 14](https://adventofcode.com/2020/day/14): Docking Data
# +
import re
with open("../data/14.txt", "r") as f:
code = re.sub(r"mask = (.*)\n", r"mem.mask = '\1'\n", f.read())
# -
# ## Part 1
# +
class Memory(dict):
"""Memory as a dictionary of address-value pairs."""
def __init__(mem, code):
"""Initialize memory by running (transformed) code."""
exec(code)
def setmask(self, val):
self._mask = self.asint("1" if x == "X" else "0" for x in val)
self._places = [i for i, x in enumerate(reversed(val)) if x == "X"]
self._overwrite = self.asint("0" if x == "X" else x for x in val)
mask = property(fset=setmask)
@staticmethod
def asint(x):
return int("".join(x), 2)
def __setitem__(self, addr, val):
"""Write values to memory."""
for a, v in self.registers(addr, val):
super().__setitem__(a, v)
def registers(self, addr, val):
"""Generate address-value pairs to write; assign dynamically."""
def __iter__(self):
return iter(super().values())
class Part1(Memory):
"""Apply bitmask to memory values."""
def registers(self, addr, val):
yield addr, (val & self._mask) | self._overwrite
assert 7817357407588 == sum(Part1(code))
# -
# ## Part 2
# +
from itertools import product
from operator import lshift
class Part2(Memory):
"""Decode memory addresses."""
def registers(self, addr, val):
addr = (addr | self._overwrite) & (self.BITS36 ^ self._mask)
for bits in product((0, 1), repeat=len(self._places)):
yield addr | sum(map(lshift, bits, self._places)), val
BITS36 = 2**36 - 1
assert 4335927555692 == sum(Part2(code))
| solutions/14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning Pipeline - Feature Engineering
#
# In the following notebooks, we will go through the implementation of each one of the steps in the Machine Learning Pipeline.
#
# We will discuss:
#
# 1. Data Analysis
# 2. **Feature Engineering**
# 3. Feature Selection
# 4. Model Training
# 5. Obtaining Predictions / Scoring
#
#
# We will use the house price dataset available on [Kaggle.com](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data). See below for more details.
#
# ===================================================================================================
#
# ## Predicting Sale Price of Houses
#
# The aim of the project is to build a machine learning model to predict the sale price of homes based on different explanatory variables describing aspects of residential houses.
#
#
# ### Why is this important?
#
# Predicting house prices is useful to identify fruitful investments, or to determine whether the price advertised for a house is over or under-estimated.
#
#
# ### What is the objective of the machine learning model?
#
# We aim to minimise the difference between the real price and the price estimated by our model. We will evaluate model performance with the:
#
# 1. mean squared error (mse)
# 2. root squared of the mean squared error (rmse)
# 3. r-squared (r2).
#
#
# ### How do I download the dataset?
#
# - Visit the [Kaggle Website](https://www.kaggle.com/c/house-prices-advanced-regression-techniques/data).
#
# - Remember to **log in**
#
# - Scroll down to the bottom of the page, and click on the link **'train.csv'**, and then click the 'download' blue button towards the right of the screen, to download the dataset.
#
# - The download the file called **'test.csv'** and save it in the directory with the notebooks.
#
#
# **Note the following:**
#
# - You need to be logged in to Kaggle in order to download the datasets.
# - You need to accept the terms and conditions of the competition to download the dataset
# - If you save the file to the directory with the jupyter notebook, then you can run the code as it is written here.
# # Reproducibility: Setting the seed
#
# With the aim to ensure reproducibility between runs of the same notebook, but also between the research and production environment, for each step that includes some element of randomness, it is extremely important that we **set the seed**.
# +
# to handle datasets
import pandas as pd
import numpy as np
# for plotting
import matplotlib.pyplot as plt
# for the yeo-johnson transformation
import scipy.stats as stats
# to divide train and test set
from sklearn.model_selection import train_test_split
# feature scaling
from sklearn.preprocessing import MinMaxScaler
# to save the trained scaler class
import joblib
# to visualise al the columns in the dataframe
pd.pandas.set_option('display.max_columns', None)
# +
# load dataset
data = pd.read_csv('train.csv')
# rows and columns of the data
print(data.shape)
# visualise the dataset
data.head()
# -
# # Separate dataset into train and test
#
# It is important to separate our data intro training and testing set.
#
# When we engineer features, some techniques learn parameters from data. It is important to learn these parameters only from the train set. This is to avoid over-fitting.
#
# Our feature engineering techniques will learn:
#
# - mean
# - mode
# - exponents for the yeo-johnson
# - category frequency
# - and category to number mappings
#
# from the train set.
#
# **Separating the data into train and test involves randomness, therefore, we need to set the seed.**
# +
# Let's separate into train and test set
# Remember to set the seed (random_state for this sklearn function)
X_train, X_test, y_train, y_test = train_test_split(
data.drop(['Id', 'SalePrice'], axis=1), # predictive variables
data['SalePrice'], # target
test_size=0.1, # portion of dataset to allocate to test set
random_state=0, # we are setting the seed here
)
X_train.shape, X_test.shape
# -
# # Feature Engineering
#
# In the following cells, we will engineer the variables of the House Price Dataset so that we tackle:
#
# 1. Missing values
# 2. Temporal variables
# 3. Non-Gaussian distributed variables
# 4. Categorical variables: remove rare labels
# 5. Categorical variables: convert strings to numbers
# 5. Put the variables in a similar scale
# ## Target
#
# We apply the logarithm
y_train = np.log(y_train)
y_test = np.log(y_test)
# ## Missing values
#
# ### Categorical variables
#
# We will replace missing values with the string "missing" in those variables with a lot of missing data.
#
# Alternatively, we will replace missing data with the most frequent category in those variables that contain fewer observations without values.
#
# This is common practice.
# +
# let's identify the categorical variables
# we will capture those of type object
cat_vars = [var for var in data.columns if data[var].dtype == 'O']
# MSSubClass is also categorical by definition, despite its numeric values
# (you can find the definitions of the variables in the data_description.txt
# file available on Kaggle, in the same website where you downloaded the data)
# lets add MSSubClass to the list of categorical variables
cat_vars = cat_vars + ['MSSubClass']
# cast all variables as categorical
X_train[cat_vars] = X_train[cat_vars].astype('O')
X_test[cat_vars] = X_test[cat_vars].astype('O')
# number of categorical variables
len(cat_vars)
# +
# make a list of the categorical variables that contain missing values
cat_vars_with_na = [
var for var in cat_vars
if X_train[var].isnull().sum() > 0
]
# print percentage of missing values per variable
X_train[cat_vars_with_na ].isnull().mean().sort_values(ascending=False)
# +
# variables to impute with the string missing
with_string_missing = [
var for var in cat_vars_with_na if X_train[var].isnull().mean() > 0.1]
# variables to impute with the most frequent category
with_frequent_category = [
var for var in cat_vars_with_na if X_train[var].isnull().mean() < 0.1]
# -
with_string_missing
# +
# replace missing values with new label: "Missing"
X_train[with_string_missing] = X_train[with_string_missing].fillna('Missing')
X_test[with_string_missing] = X_test[with_string_missing].fillna('Missing')
# -
for var in with_frequent_category:
# there can be more than 1 mode in a variable
# we take the first one with [0]
mode = X_train[var].mode()[0]
print(var, mode)
X_train[var].fillna(mode, inplace=True)
X_test[var].fillna(mode, inplace=True)
# +
# check that we have no missing information in the engineered variables
X_train[cat_vars_with_na].isnull().sum()
# +
# check that test set does not contain null values in the engineered variables
[var for var in cat_vars_with_na if X_test[var].isnull().sum() > 0]
# -
# ### Numerical variables
#
# To engineer missing values in numerical variables, we will:
#
# - add a binary missing indicator variable
# - and then replace the missing values in the original variable with the mean
# +
# now let's identify the numerical variables
num_vars = [
var for var in X_train.columns if var not in cat_vars and var != 'SalePrice'
]
# number of numerical variables
len(num_vars)
# +
# make a list with the numerical variables that contain missing values
vars_with_na = [
var for var in num_vars
if X_train[var].isnull().sum() > 0
]
# print percentage of missing values per variable
X_train[vars_with_na].isnull().mean()
# +
# replace missing values as we described above
for var in vars_with_na:
# calculate the mean using the train set
mean_val = X_train[var].mean()
print(var, mean_val)
# add binary missing indicator (in train and test)
X_train[var + '_na'] = np.where(X_train[var].isnull(), 1, 0)
X_test[var + '_na'] = np.where(X_test[var].isnull(), 1, 0)
# replace missing values by the mean
# (in train and test)
X_train[var].fillna(mean_val, inplace=True)
X_test[var].fillna(mean_val, inplace=True)
# check that we have no more missing values in the engineered variables
X_train[vars_with_na].isnull().sum()
# +
# check that test set does not contain null values in the engineered variables
[var for var in vars_with_na if X_test[var].isnull().sum() > 0]
# +
# check the binary missing indicator variables
X_train[['LotFrontage_na', 'MasVnrArea_na', 'GarageYrBlt_na']].head()
# -
# ## Temporal variables
#
# ### Capture elapsed time
#
# We learned in the previous notebook, that there are 4 variables that refer to the years in which the house or the garage were built or remodeled.
#
# We will capture the time elapsed between those variables and the year in which the house was sold:
def elapsed_years(df, var):
# capture difference between the year variable
# and the year in which the house was sold
df[var] = df['YrSold'] - df[var]
return df
for var in ['YearBuilt', 'YearRemodAdd', 'GarageYrBlt']:
X_train = elapsed_years(X_train, var)
X_test = elapsed_years(X_test, var)
# now we drop YrSold
X_train.drop(['YrSold'], axis=1, inplace=True)
X_test.drop(['YrSold'], axis=1, inplace=True)
# ## Numerical variable transformation
#
# ### Logarithmic transformation
#
# In the previous notebook, we observed that the numerical variables are not normally distributed.
#
# We will transform with the logarightm the positive numerical variables in order to get a more Gaussian-like distribution.
for var in ["LotFrontage", "1stFlrSF", "GrLivArea"]:
X_train[var] = np.log(X_train[var])
X_test[var] = np.log(X_test[var])
# check that test set does not contain null values in the engineered variables
[var for var in ["LotFrontage", "1stFlrSF", "GrLivArea"] if X_test[var].isnull().sum() > 0]
# same for train set
[var for var in ["LotFrontage", "1stFlrSF", "GrLivArea"] if X_train[var].isnull().sum() > 0]
# ### Yeo-Johnson transformation
#
# We will apply the Yeo-Johnson transformation to LotArea.
# +
# the yeo-johnson transformation learns the best exponent to transform the variable
# it needs to learn it from the train set:
X_train['LotArea'], param = stats.yeojohnson(X_train['LotArea'])
# and then apply the transformation to the test set with the same
# parameter: see who this time we pass param as argument to the
# yeo-johnson
X_test['LotArea'] = stats.yeojohnson(X_test['LotArea'], lmbda=param)
print(param)
# -
# check absence of na in the train set
[var for var in X_train.columns if X_train[var].isnull().sum() > 0]
# check absence of na in the test set
[var for var in X_train.columns if X_test[var].isnull().sum() > 0]
# ### Binarize skewed variables
#
# There were a few variables very skewed, we would transform those into binary variables.
# +
skewed = [
'BsmtFinSF2', 'LowQualFinSF', 'EnclosedPorch',
'3SsnPorch', 'ScreenPorch', 'MiscVal'
]
for var in skewed:
# map the variable values into 0 and 1
X_train[var] = np.where(X_train[var]==0, 0, 1)
X_test[var] = np.where(X_test[var]==0, 0, 1)
# -
# ## Categorical variables
#
# ### Apply mappings
#
# These are variables which values have an assigned order, related to quality. For more information, check Kaggle website.
# +
# re-map strings to numbers, which determine quality
qual_mappings = {'Po': 1, 'Fa': 2, 'TA': 3, 'Gd': 4, 'Ex': 5, 'Missing': 0, 'NA': 0}
qual_vars = ['ExterQual', 'ExterCond', 'BsmtQual', 'BsmtCond',
'HeatingQC', 'KitchenQual', 'FireplaceQu',
'GarageQual', 'GarageCond',
]
for var in qual_vars:
X_train[var] = X_train[var].map(qual_mappings)
X_test[var] = X_test[var].map(qual_mappings)
# +
exposure_mappings = {'No': 1, 'Mn': 2, 'Av': 3, 'Gd': 4}
var = 'BsmtExposure'
X_train[var] = X_train[var].map(exposure_mappings)
X_test[var] = X_test[var].map(exposure_mappings)
# +
finish_mappings = {'Missing': 0, 'NA': 0, 'Unf': 1, 'LwQ': 2, 'Rec': 3, 'BLQ': 4, 'ALQ': 5, 'GLQ': 6}
finish_vars = ['BsmtFinType1', 'BsmtFinType2']
for var in finish_vars:
X_train[var] = X_train[var].map(finish_mappings)
X_test[var] = X_test[var].map(finish_mappings)
# +
garage_mappings = {'Missing': 0, 'NA': 0, 'Unf': 1, 'RFn': 2, 'Fin': 3}
var = 'GarageFinish'
X_train[var] = X_train[var].map(garage_mappings)
X_test[var] = X_test[var].map(garage_mappings)
# +
fence_mappings = {'Missing': 0, 'NA': 0, 'MnWw': 1, 'GdWo': 2, 'MnPrv': 3, 'GdPrv': 4}
var = 'Fence'
X_train[var] = X_train[var].map(fence_mappings)
X_test[var] = X_test[var].map(fence_mappings)
# -
# check absence of na in the train set
[var for var in X_train.columns if X_train[var].isnull().sum() > 0]
# ### Removing Rare Labels
#
# For the remaining categorical variables, we will group those categories that are present in less than 1% of the observations. That is, all values of categorical variables that are shared by less than 1% of houses, well be replaced by the string "Rare".
#
# To learn more about how to handle categorical variables visit our course [Feature Engineering for Machine Learning](https://www.udemy.com/course/feature-engineering-for-machine-learning/?referralCode=A855148E05283015CF06) in Udemy.
# +
# capture all quality variables
qual_vars = qual_vars + finish_vars + ['BsmtExposure','GarageFinish','Fence']
# capture the remaining categorical variables
# (those that we did not re-map)
cat_others = [
var for var in cat_vars if var not in qual_vars
]
len(cat_others)
# +
def find_frequent_labels(df, var, rare_perc):
# function finds the labels that are shared by more than
# a certain % of the houses in the dataset
df = df.copy()
tmp = df.groupby(var)[var].count() / len(df)
return tmp[tmp > rare_perc].index
for var in cat_others:
# find the frequent categories
frequent_ls = find_frequent_labels(X_train, var, 0.01)
print(var, frequent_ls)
print()
# replace rare categories by the string "Rare"
X_train[var] = np.where(X_train[var].isin(
frequent_ls), X_train[var], 'Rare')
X_test[var] = np.where(X_test[var].isin(
frequent_ls), X_test[var], 'Rare')
# -
# ### Encoding of categorical variables
#
# Next, we need to transform the strings of the categorical variables into numbers.
#
# We will do it so that we capture the monotonic relationship between the label and the target.
#
# To learn more about how to encode categorical variables visit our course [Feature Engineering for Machine Learning](https://www.udemy.com/course/feature-engineering-for-machine-learning/?referralCode=A855148E05283015CF06) in Udemy.
# +
# this function will assign discrete values to the strings of the variables,
# so that the smaller value corresponds to the category that shows the smaller
# mean house sale price
def replace_categories(train, test, y_train, var, target):
tmp = pd.concat([X_train, y_train], axis=1)
# order the categories in a variable from that with the lowest
# house sale price, to that with the highest
ordered_labels = tmp.groupby([var])[target].mean().sort_values().index
# create a dictionary of ordered categories to integer values
ordinal_label = {k: i for i, k in enumerate(ordered_labels, 0)}
print(var, ordinal_label)
print()
# use the dictionary to replace the categorical strings by integers
train[var] = train[var].map(ordinal_label)
test[var] = test[var].map(ordinal_label)
# -
for var in cat_others:
replace_categories(X_train, X_test, y_train, var, 'SalePrice')
# check absence of na in the train set
[var for var in X_train.columns if X_train[var].isnull().sum() > 0]
# check absence of na in the test set
[var for var in X_test.columns if X_test[var].isnull().sum() > 0]
# +
# let me show you what I mean by monotonic relationship
# between labels and target
def analyse_vars(train, y_train, var):
# function plots median house sale price per encoded
# category
tmp = pd.concat([X_train, np.log(y_train)], axis=1)
tmp.groupby(var)['SalePrice'].median().plot.bar()
plt.title(var)
plt.ylim(2.2, 2.6)
plt.ylabel('SalePrice')
plt.show()
for var in cat_others:
analyse_vars(X_train, y_train, var)
# -
# The monotonic relationship is particularly clear for the variables MSZoning and Neighborhood. Note how, the higher the integer that now represents the category, the higher the mean house sale price.
#
# (remember that the target is log-transformed, that is why the differences seem so small).
# ## Feature Scaling
#
# For use in linear models, features need to be either scaled. We will scale features to the minimum and maximum values:
# +
# create scaler
scaler = MinMaxScaler()
# fit the scaler to the train set
scaler.fit(X_train)
# transform the train and test set
# sklearn returns numpy arrays, so we wrap the
# array with a pandas dataframe
X_train = pd.DataFrame(
scaler.transform(X_train),
columns=X_train.columns
)
X_test = pd.DataFrame(
scaler.transform(X_test),
columns=X_train.columns
)
# -
X_train.head()
# +
# let's now save the train and test sets for the next notebook!
X_train.to_csv('xtrain.csv', index=False)
X_test.to_csv('xtest.csv', index=False)
y_train.to_csv('ytrain.csv', index=False)
y_test.to_csv('ytest.csv', index=False)
# +
# now let's save the scaler
joblib.dump(scaler, 'minmax_scaler.joblib')
# -
# That concludes the feature engineering section.
#
# # Additional Resources
#
# - [Feature Engineering for Machine Learning](https://www.udemy.com/course/feature-engineering-for-machine-learning/?referralCode=A855148E05283015CF06) - Online Course
# - [Packt Feature Engineering Cookbook](https://www.packtpub.com/data/python-feature-engineering-cookbook) - Book
# - [Feature Engineering for Machine Learning: A comprehensive Overview](https://trainindata.medium.com/feature-engineering-for-machine-learning-a-comprehensive-overview-a7ad04c896f8) - Article
# - [Practical Code Implementations of Feature Engineering for Machine Learning with Python](https://towardsdatascience.com/practical-code-implementations-of-feature-engineering-for-machine-learning-with-python-f13b953d4bcd) - Article
| section-04-research-and-development/02-machine-learning-pipeline-feature-engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # 可选scipy加速例程(numpy.dual)
#
# https://numpy.org/doc/stable/reference/routines.dual.html
#
# + pycharm={"name": "#%%\n"}
import numpy as np
a = np.array([[1, 2], [3, 4]])
np.linalg.det(a)
| numpy/reference/17_routines.dual.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Network analysis for coauthors - network through time
#
# This looks at how the network changes through time with network metrics.
# +
# %load_ext autoreload
# %autoreload 2
from src import util as u
import pandas as pd
from functools import reduce
import operator
import networkx as nx
# -
nodes = pd.read_csv(u.fn_nodes)
# Import author data with dates
start = 1758
end = 2018
years = end-start
timepoints = 4
interval = round(years/3, 0)
year1= start + interval*1
year2= start + interval*2
print(start, year1)
print(year1, year2) # postal mail invented
print(year2, end) # airlines invested
# +
auth = u.get_spp_df()
auth = auth.drop(columns=['status'])
auth1 = auth[(auth['date'] >= start) & (auth['date'] < year1)][['idx', 'full_name_of_describer']]
auth2 = auth[(auth['date'] >= year1) & (auth['date'] < year2)][['idx', 'full_name_of_describer']]
auth3 = auth[(auth['date'] >= year2) & (auth['date'] <= end)][['idx', 'full_name_of_describer']]
df_li = [auth1, auth2, auth3]
df_li = [x.groupby('idx')['full_name_of_describer'].apply(lambda x: "%s" % '; '.join(x)) for x in df_li]
# +
# Get pairs
def li_pairs(source):
source = str.split(source, "; ")
result = []
N = len(source)
if (N <= 1):
return [(source[0], None)]
else:
for p1 in range(N):
for p2 in range(p1+1,len(source)):
result.append((source[p1],source[p2]))
return result
# -
print(df_li[2].iloc[2])
print(li_pairs(df_li[2].iloc[2]))
print(li_pairs(df_li[2].iloc[63]))
df_li = [x.apply(lambda x: li_pairs(x)) for x in df_li]
li = [[] for i in range(3)]
for i in range(len(df_li)):
df = df_li[i]
for j, row in df.iteritems():
li[i].append(row[0])
df_li = [pd.DataFrame(x, columns=['p1', 'p2']) for x in li]
df_li = [pd.DataFrame(x).groupby(['p1', 'p2']).size() for x in df_li]
df_li = [x.reset_index() for x in df_li]
def get_edges(df):
edges = []
for i, row in df.iterrows():
edges.append((row.p1, row.p2, row[0]))
return edges
df_li = [get_edges(df) for df in df_li]
def get_node_names(edges):
all_node_names = []
all_node_names = [all_node_names + [e[0], e[1]] for e in edges]
all_node_names = reduce(operator.add, all_node_names)
return set(all_node_names)
node_names = [get_node_names(df) for df in df_li]
# Load into graph
g_li = [nx.Graph() for x in range(3)]
for i in range(len(df_li)):
g_li[i].add_nodes_from(node_names[i])
g_li[i].add_weighted_edges_from(df_li[i])
for i in range(len(df_li)):
print("Network", i)
density = nx.density(g_li[i])
print("Network density:", round(density*100, 1), "%")
triadic_closure = nx.transitivity(g_li[i])
print("Triadic closure:", round(triadic_closure*100, 1), "%")
# Get subgraphs
subgraphs = [c for c in sorted(nx.connected_components(g_li[i]), key=len, reverse=True)]
print("Number of subgraphs:", len(subgraphs))
# Largest component
components = nx.connected_components(g_li[i])
largest_component = max(components, key=len) # max number of nodes
# Create a "subgraph" of just the largest component
# Then calculate the diameter of the subgraph, just like you did with density.
subgraph = g_li[i].subgraph(largest_component)
diameter = nx.diameter(subgraph)
print("Network diameter of largest component:", diameter)
print([v[0] for v in g_li[i].nodes(data=True)])
print("\n")
| 2020-08-31-jsa-type-v2-ch3-coauth/2019-11-11-co-author-networkx-time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import pickle
import re
from joblib import load, dump
from sklearn.neighbors import NearestNeighbors
pd.set_option('display.max_columns', 24)
# -
# joblib our model
def joblib_model():
"""
Opens our .csv file for model training
Trains a nearest neighbor model on our Spotify dataset, and joblib dumps the model
Returns the model, and the knn file
input parameters:
----------------
None
output:
------
model
"""
# import data into a dataframe
df = pd.read_csv('./spotify_rock.csv', index_col=0)
# training dataframe for NN model (drop str columns, 'artists', 'song')
df_train = df[df.columns[3:]]
# make a copy of the truncated dataframe
df_train_trunc = df_train.copy()
# create an instance of the Nearest Neighbors class
model = NearestNeighbors(n_neighbors=5, algorithm='kd_tree')
# fit the model
model.fit(df_train_trunc)
# joblib the model
dump(model, "knn_model.joblib", compress=True)
return model
model_knn = joblib_model()
# +
# create test use query(ies)
# user_req = df.index[(df['song'] == 'Magic Carpet Ride') & (df['artists'] == 'Steppenwolf')]
# user_req = df.index[(df['song'] == 'Know Your Enemy') & (df['artists'] == 'Rage Against The Machine')]
user_req = df.index[(df['song'] == 'Jump') & (df['artists'] == 'Van Halen')]
submit = user_req[0]
# return model inputs based on user artist/song entry
series = df.iloc[user_req, 3:].to_numpy()
# query model based on input
neighbors = model_knn.kneighbors(series, return_distance=False)
# checkout results of NN
neighbors[0]
# -
song_artists = pd.read_csv("./songs_artists.csv", index_col=0)
song_artists.head()
song_artists.loc[song_artists['artists'] == 'The Beatles']
# explore what the model returns as neighbors
for _ in range(len(neighbors[0])):
display(song_artists[song_artists.index == neighbors[0][_]])
# load the model
classifier = load('./knn_model.joblib')
# send through the same query using the joblib model
neighbors = classifier.kneighbors(series, return_distance=False)
# checkout results of NN
neighbors[0]
| notebooks/Nearest_Neighbor_Joblib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from numpy import geomspace
import itertools
import random
from tqdm import tqdm
# # CoinOracle Class
class CoinOracle:
def __init__(self, n_coins, secret_coin, is_heavier):
assert secret_coin <= n_coins
# Secret attributes
self.__secret_coin = secret_coin
self.__is_heavier = is_heavier
self.__coin_weights = list([5 for _ in range(n_coins+1)])
self.__coin_weights[secret_coin] = 5.1 if is_heavier else 4.9
self.__experiment_results = []
self.__check_guess_count = 0
def __repr__(self):
return f"CoinOracle: {self.n_coins} coins and {self.n_tests} tests made"
def __weight(self,coin_list):
return sum([self.__coin_weights[i] for i in coin_list])
# Accessible attributes
@property
def n_coins(self):
return len(self.__coin_weights)-1
@property
def n_tests(self):
return len(self.__experiment_results)
@property
def experiment_results(self):
return self.__experiment_results
# Key methods
def is_left_side_heavier(self, left_side, right_side):
if self.__weight(left_side) > self.__weight(right_side):
result = 1
elif self.__weight(left_side) < self.__weight(right_side):
result = -1
else:
result = 0
self.experiment_results.append((left_side,right_side, result))
return result
def check_guess(self, secret_coin, is_heavier):
self.__check_guess_count += 1
return secret_coin==self.__secret_coin and is_heavier==self.__is_heavier
# # Solutions
# ### Marta
def marta(temp_oracle):
coins = list(range(temp_oracle.n_coins + 1))
last_weigh_diff = ([], None) # left group, weigh in result
while True:
if len(coins) == 2:
break
chunk_size = len(coins) // 3
chunks = [coins[i * chunk_size:(i + 1) * chunk_size] for i in
range((len(coins) + chunk_size - 1) // chunk_size)]
lf_sd_heavier = temp_oracle.is_left_side_heavier(chunks[0], chunks[1])
if lf_sd_heavier == 0:
coins = [0]
for i in range(2, len(chunks)):
coins += chunks[i]
else:
coins = chunks[0] + chunks[1]
last_weigh_diff = (chunks[0], lf_sd_heavier)
if last_weigh_diff[1] is None:
return coins[1], not temp_oracle.is_left_side_heavier([coins[0]], [coins[1]]) == 1
else:
heavier = last_weigh_diff[1] * -1 if coins[1] in last_weigh_diff[0] else last_weigh_diff[1]
return coins[1], not heavier == 1
# ### <NAME>
# +
from math import floor
from typing import List, Tuple
# Let the result of is_left_side_heavier(L, R) be one of (-1, 0, 1) as RESULT
# The Scenarios Metric below is given by the tuple:
# (Sum of RESULT, Sum of abs(RESULT), RESULT(A vs. B))
HEAVY_IN_A = (2, 2, 1)
HEAVY_IN_B = (0, 2, -1)
HEAVY_IN_C = (-2, 2, 0)
LIGHT_IN_A = (-2, 2, -1)
LIGHT_IN_B = (0, 2, 1)
LIGHT_IN_C = (2, 2, 0)
def n_sized_split(
original_list: List[int],
n: int,
) -> Tuple[List[int], List[int], List[int], List[int]]:
"""
Returns a list equally divided in 'n' parts and the leftover, assuming the
given list is large enough.
"""
K = len(original_list)
N = floor(K / n)
split_list = [original_list[i*N:(i+1)*N] for i in range(0, n)]
rest_list = original_list[n*N:]
split_list.append(rest_list)
return split_list
def is_heavier_than_control(temp_oracle, coin: int, control: int = 0) -> bool:
"""
Determines whether some coin is heavier than the control.
"""
left = [coin]
right = [control]
result = temp_oracle.is_left_side_heavier(left, right)
if result == 1:
return True
return False
def L_vs_R(temp_oracle, L: int, R: int) -> Tuple[int, bool]:
"""
Determines which coin is heavier, assuming they do not have the same weight.
"""
coin = 0
is_heavier = None
result_L = temp_oracle.is_left_side_heavier([L], [0])
result_R = temp_oracle.is_left_side_heavier([R], [0])
if result_L == 1:
coin = L
is_heavier = True
elif result_L == -1:
coin = L
is_heavier = False
elif result_R == 1:
coin = R
is_heavier = True
elif result_R == -1:
coin = R
is_heavier = False
return coin, is_heavier
def reduce_list(temp_oracle, coin_list: List[int]) -> Tuple[int, bool]:
"""
Recursive call to reduce a given list of coins until we have the result.
"""
coin = 0
N = len(coin_list)
# The stop conditions for the recursion which we can easily evaluate
if N == 1:
coin = coin_list[0]
return coin, is_heavier_than_control(temp_oracle, coin)
elif N == 2:
left = coin_list[0]
right = coin_list[1]
return L_vs_R(temp_oracle, left, right)
A, B, C, *rest = n_sized_split(coin_list, 3)
D = rest[0] if len(rest) > 0 else []
# We compare the bundles against each other
result_A_B = temp_oracle.is_left_side_heavier(A, B)
result_A_C = temp_oracle.is_left_side_heavier(A, C)
result_B_C = temp_oracle.is_left_side_heavier(B, C)
# And compute some metrics
result_sum = result_A_B + result_A_C + result_B_C
result_abs_sum = abs(result_A_B) + abs(result_A_C) + abs(result_B_C)
metric = (result_sum, result_abs_sum, result_A_B)
if metric == HEAVY_IN_A or metric == LIGHT_IN_A:
coin = A
elif metric == HEAVY_IN_B or metric == LIGHT_IN_B:
coin = B
elif metric == HEAVY_IN_C or metric == LIGHT_IN_C:
coin = C
else:
coin = D
return reduce_list(temp_oracle, coin)
def joao_carmo(temp_oracle) -> Tuple[int, bool]:
# The first coin is a control which we know to be always a good coin
coin = 0
is_heavier = False
N = temp_oracle.n_coins
coin, is_heavier = reduce_list(temp_oracle, list(range(0, N + 1)))
return coin, is_heavier
# -
# ### Miguel
def miguel(temp_oracle):
possible_coins = list(range(1, temp_oracle.n_coins+1))
found_unequal_weights = False
while not found_unequal_weights:
n = len(possible_coins)
left_side = possible_coins[round(n/3): round(2*n/3)]
right_side = possible_coins[round(2*n/3):]
while len(left_side)<len(right_side):
left_side.append(0)
while len(right_side)<len(left_side):
right_side.append(0)
result = temp_oracle.is_left_side_heavier(left_side,right_side)
found_unequal_weights = result != 0
if not found_unequal_weights:
possible_coins = possible_coins[:round(n/3)]
left_side_is_heavier = result == 1
if left_side_is_heavier:
heavier_coins,lighter_coins = left_side, right_side
else:
heavier_coins,lighter_coins = right_side, left_side
unique_coin = len(heavier_coins) + len(lighter_coins)<2
while not unique_coin:
a = len(heavier_coins)
b = len(lighter_coins)
left_side = heavier_coins[:round(a/3)] + lighter_coins[:round(b/3)]
right_side = heavier_coins[round(a/3):round(2*a/3)] + lighter_coins[round(b/3):round(2*b/3)]
while len(left_side)<len(right_side):
left_side.append(0)
while len(right_side)<len(left_side):
right_side.append(0)
result = temp_oracle.is_left_side_heavier(left_side, right_side)
if result == 0:
heavier_coins = heavier_coins[round(2*a/3):]
lighter_coins = lighter_coins[round(2*b/3):]
elif result == 1:
heavier_coins = heavier_coins[:round(a/3)]
lighter_coins = lighter_coins[round(b/3):round(2*b/3)]
else:
heavier_coins = heavier_coins[round(a/3):round(2*a/3)]
lighter_coins = lighter_coins[:round(b/3)]
unique_coin = len(heavier_coins) + len(lighter_coins)<2
if heavier_coins:
return (heavier_coins[0], True)
if lighter_coins:
return (lighter_coins[0], False)
# # Your solution
#
# The solution is similar to a brute force solution, but it stops when the fake coin is found. Can you improve it?
def your_custom_solution(temp_oracle):
i=1
while temp_oracle.is_left_side_heavier([i],[0])==0:
i+=1
coin = i
is_heavier = temp_oracle.experiment_results[-1][2]==1
return (coin, is_heavier)
# # The code below runs games for a given list of solvers
# Runs one game for each size in sizes, using all solvers passed. Filename is used to store the plot in the end as a file.
def run_comparison(sizes, solvers):
df = pd.DataFrame(
max(sizes)+1,
columns=[solver.__name__ for solver in solvers],
index=sizes
)
secret_coins = {size: random.choice(range(size))+1 for size in sizes}
n_pairs = len(sizes)*len(solvers)
for (size, solver) in tqdm(itertools.product(sizes, solvers), total=n_pairs):
oracle = CoinOracle(
n_coins = size,
secret_coin = secret_coins[size],
is_heavier = random.choice([True,False])
)
(guessed_secret_coin, guess_is_heavier) = solver(oracle)
if oracle.check_guess(guessed_secret_coin, guess_is_heavier):
df[solver.__name__][size] = oracle.n_tests
else:
print(f"Solver {solver.__name__} incorrectly guessed for size {size} with secret_coin {secret_cois[size]}")
return df
sizes = sorted(set(map(lambda x: floor(x), geomspace(10,10**7, 1000))), reverse=True)
solvers = [marta, joao_carmo, miguel]
df = run_comparison(sizes, solvers)
df.T
# ### The plot below shows the number of experiments conducted by your custom solution vs the theoretical maximum, for multiple sizes. What is the best score you can get?
#
# Remark: It is possible that you occasionally get a better result than the theoretical optimal, for example if you happen to be lucky and test the fake coin in the very first run. However it is impossible to consistently beat the theoretical optimal
from math import log
df["theoretical_optimum"] = list(map(lambda x: log(2*x, 3), df.index))
df.plot.area(figsize=(16,8), stacked=False, logx=True).figure.savefig('results.png')
| Coins_DoE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import networkx as nx
alice = nx.Graph()
alice.add_edge('Alice','Rabbit')
alice.add_edge('Alice','Cat')
alice.add_edge('Alice','Queen')
alice.add_edge('Alice','Caterpillar')
alice.add_edge('Alice','Hatter')
alice.add_edge('Alice','Hare')
alice.add_edge('Rabbit','Queen')
alice.add_edge('Hatter','Dormouse')
alice.add_edge('Hatter','Hare')
alice.add_edge('Hare','Dormouse')
print(nx.info(alice))
# degree to which any node is linked ot each other (if all linekd to each other then 1, in ours not all are linekd)
nx.average_clustering(alice)
# shortest path typically through Alice at center
nx.shortest_path(alice, 'Caterpillar', 'Queen')
# exploring betweenness centrality - how likely others will pass through you
nx.betweenness_centrality(alice)
# degree centrality is how many links you have compared to how many you could
#alice is only not connected to the dormouse
nx.degree_centrality(alice)
# how many (not just shortest) in total lead to this person/object - useful for larger networks
nx.eigenvector_centrality(alice)
# plenty of lbraries available to make thsi prettier that are not nx - convenient for checking
nx.draw_networkx(alice)
# we can make a directional graph
bb = nx.DiGraph()
bb.add_edge('Walter','Emelio')
bb.add_edge('Walter','Krazy-8')
bb.add_edge('Walter','Jack')
bb.add_edge('Walter','Walter')
bb.add_edge('Tuco','No-Doze')
bb.add_edge('Hank','Tuco')
bb.add_edge('Jesse','Joaquin')
bb.add_edge('Jack','Hank')
# spring layout spaces things out
nx.spring_layout(bb)
nx.draw_networkx(bb)
# +
#Now add weights and relations (how this is related to that)
bb = nx.DiGraph()
bb.add_edge('Walter','Emelio', relation='poisoned')
bb.add_edge('Walter','Krazy-8', relation='strangled')
bb.add_edge('Walter','Jack', relation='shot')
bb.add_edge('Walter','Walter', relation='shot')
bb.add_edge('Tuco','No-Doze', relation='punched')
bb.add_edge('Hank','Tuco', relation='shot')
bb.add_edge('Jesse','Joaquin', relation='shot')
bb.add_edge('Jack','Hank', relation='shot')
nx.spring_layout(bb)
nx.draw_networkx(bb)
# +
import networkx as nx
#networkx has built-in demo graphs
graph = nx.florentine_families_graph()
nx.draw_networkx(graph)
# -
# new lesson 6.3
import tarfile
import urllib.request as urllib
# get a network from SNAP and store it int he same folder as this notebook
url = 'https://snap.stanford.edu/data/facebook.tar.gz'
stream = urllib.urlopen(url)
archive = tarfile.open(fileobj=stream, mode='r|gz')
archive.extractall()
fn = 'facebook/0.edges'
open(fn).readlines()[:15]
# handy function for loading edges
graph = nx.read_edgelist(fn)
print(nx.info(graph))
# note that the ego node like our Alice node above is excluded otherwise everything would link to it.
# In this case, the person whose network this is.
nx.draw_networkx(graph, with_labels=False, node_size=10)
# we can see little groups that do not know any of the bigger group members
# grabbing a zip this time
from zipfile import ZipFile
from io import BytesIO
#Let's get a new file
# +
#this time we are not extracting to our folder, using the files directly via cache
url = 'http://www-personal.umich.edu/~mejn/netdata/football.zip'
with urllib.urlopen(url) as stream:
with ZipFile(BytesIO(stream.read())) as archive:
archive.printdir()
txt = archive.read('football.txt').decode()
gml = archive.read('football.gml').decode()
# -
# gml = Graph Markup Language - this includes nodes and edges
# this one shows games played between teams
print(txt)
graph = nx.parse_gml(gml)
nx.draw_networkx(graph)
# +
# add options to make it readable (and pass options in a clean way)
# also paint all black except ASU
colors = ['black' for node in graph]
colors[8] = 'maroon'
# could normally pass one color as string but highlighting is cool
options = {
'node_color':colors,
'node_size':50,
'linewidths':0,
'width':0.1,
'with_labels':False
}
nx.draw_networkx(graph, **options)
# you can see the football conferences
# -
import geopandas as gpd
import matplotlib.pyplot as plt
# NEW SECTION on Mechanical Turk
| Module 6/Practice Book for Videos.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kool7/Lyft_Motion_Prediction_for_Autonomous_Vehicles_2020_Kaggle/blob/master/colab/multimode/Lyft_L5_V4_448x448.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="hYFrAu2fFfrw"
# # Introduction
#
# This is the 4 version of previous notebooks which were used in [this](https://www.kaggle.com/c/lyft-motion-prediction-autonomous-vehicles/overview) competition Lyft Motion Prediction for Autonomous Vehicles
# . For full EDA please visit [here](https://www.kaggle.com/kool777/lyft-level5-eda-training-inference).
#
# * Objective - Our task is to build motion prediction models. We need to predict the position of cars, cyclists and pedestrians in AV's environment. We need to predict the location of agents in next 50 frames.
#
# * Tasks Performed :-
# * Using new version of l5kit (1.1.0)
# * Training for 60000 iterations
# * Used negative log loss
#
# This notebook is based on discussions [here](https://www.kaggle.com/c/lyft-motion-prediction-autonomous-vehicles/discussion/186492) and [here](https://www.kaggle.com/c/lyft-motion-prediction-autonomous-vehicles/discussion/187825).
# + id="Qo09lAQkE-wz"
# mount google drive
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="-NVbgGgCSdxE"
# # Data Preparation
#
# We are going to download data from kaggle.
# + id="CrYy8Sn3SbVm"
import os
os.makedirs('/content/lyft-motion-prediction-autonomous-vehicles/', exist_ok=True)
os.chdir('/content/lyft-motion-prediction-autonomous-vehicles/')
# %cd '/content/lyft-motion-prediction-autonomous-vehicles/'
# !pwd
# + id="fVwSpM0ASiwN"
os.makedirs('/content/lyft-motion-prediction-autonomous-vehicles/scenes/', exist_ok=True)
os.chdir('/content/lyft-motion-prediction-autonomous-vehicles/scenes/')
# !wget --header="Host: storage.googleapis.com" --header="User-Agent: Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/85.0.4183.102 Safari/537.36" --header="Accept: text/html,application/xhtml+xml,application/xml;q=0.9,image/avif,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9" --header="Accept-Language: en-US,en;q=0.9" --header="Referer: https://www.kaggle.com/" --header="Cookie: _ga=GA1.3.1723755215.1594569279" --header="Connection: keep-alive" "https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/19990/1472735/compressed/scenes.zip?GoogleAccessId=<EMAIL>&Expires=1601530855&Signature=VUL5ViMOqoxt4ZberYrAxJsQTJmUd%2BNtPLKvi%2FYgw%2Fg3V9qzXQB%2Bp9320GKErjcqUmuHMgydxoiMZXDt8TNUCk%2BxpbQMGVPxwhOyVP0NqKMIi74YrKNbzjiLTabNtUSHEx1%2FgWDxtgNhkyNslAtT%2FENUV8zqZrABKX2Lo7VKv0f7s9r%2FjM%2BX4FMavjAgPc5YlHsSNGPwsVhNUraRPFSkeEpwakgLSq6uF4VhwV3bIhcSzsnxSZH9wLYZ5Lwj9rMryHNhmxTrQESENMkdht%2FAdPqCe4t6KOFgwe6r%2B6rWT%2BqcQaFK6fDqQHHw0gIj3VE4HNDkVqaV4ayzhuHIeNbBzQ%3D%3D&response-content-disposition=attachment%3B+filename%3Dscenes.zip" -c -O 'scenes.zip'
# !unzip ./scenes.zip
# + id="REoxKIELStjg"
# %cd '/content/lyft-motion-prediction-autonomous-vehicles/scenes/'
# !pwd
# !ls
# + id="8oCkUO9FSuWz"
# %rm -r validate.zarr/
# %rm -r sample.zarr/
# %rm scenes.zip
# + id="Y6MmoxKwSwjy"
# %cd ..
# !pwd
# !ls
# + id="28jlNtkDSyZt"
os.makedirs('/content/lyft-motion-prediction-autonomous-vehicles/aerial_map/', exist_ok=True)
os.chdir('/content/lyft-motion-prediction-autonomous-vehicles/aerial_map/')
# !wget --header="Host: storage.googleapis.com" --header="User-Agent: Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/85.0.4183.102 Safari/537.36" --header="Accept: text/html,application/xhtml+xml,application/xml;q=0.9,image/avif,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9" --header="Accept-Language: en-US,en;q=0.9" --header="Referer: https://www.kaggle.com/" --header="Cookie: _ga=GA1.3.1723755215.1594569279" --header="Connection: keep-alive" "https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/19990/1472735/compressed/aerial_map.zip?GoogleAccessId=<EMAIL>&Expires=1601531043&Signature=QTkS13tw%2BItCroMzlv4SWj9%2F3e9LjKXgjdBw9ANbGAkBQMRIlRp1xD4eUlUBHwxUsw5l7VEIDjun0ndl%2FlIudvpMBwCHg%2BDCXWRrMslh9LWn7hFMulCoZLEO4oaou3LWs1vjCIk0%2BMSyLsWUBYvoqzTznSVm2mDhVEa%2B4sd%2FzfU%2FkLeJviUhA1%2BODufDNM%2B%2Bcyma%2FeLMF7FeKEoSce0Jslur1bQbhdroQZ%2FQ22PXTuDRd0JasgPkNvLRYH%2BDfCo3hVFE%2Bn176Wv59R3jg0Tymk2mZtZJCSMUnp2YaOR6sjqNKc3Rxk3RrAIaqXhwgkJebEQTiZhXip8wz9flmsV7cw%3D%3D&response-content-disposition=attachment%3B+filename%3Daerial_map.zip" -c -O 'aerial_map.zip'
# !unzip ./aerial_map.zip
# + id="3PL-zLtUS0UX"
# %rm -r aerial_map.zip
# + id="Deg25CmxS2gs"
# %cd ..
# !pwd
# !ls
# + id="9Asb5SpxS4OT"
os.makedirs('/content/lyft-motion-prediction-autonomous-vehicles/semantic_map/', exist_ok=True)
os.chdir('/content/lyft-motion-prediction-autonomous-vehicles/semantic_map/')
# !wget --header="Host: storage.googleapis.com" --header="User-Agent: Mozilla/5.0 (Windows NT 10.0; Win64; x64) AppleWebKit/537.36 (KHTML, like Gecko) Chrome/85.0.4183.102 Safari/537.36" --header="Accept: text/html,application/xhtml+xml,application/xml;q=0.9,image/avif,image/webp,image/apng,*/*;q=0.8,application/signed-exchange;v=b3;q=0.9" --header="Accept-Language: en-US,en;q=0.9" --header="Referer: https://www.kaggle.com/" --header="Cookie: _ga=GA1.3.1723755215.1594569279" --header="Connection: keep-alive" "https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/19990/1472735/upload/semantic_map.zip?GoogleAccessId=<EMAIL>&Expires=1601531085&Signature=KS3o7dPKQgF9ubQPtFYhYRft%2B84qbqczH4S1QdIOgYP2haztAtLTNYJ64xPKuWGH6XkcTVUV6%2F8vCCNNAPeENWi6B8%2Fy1LFaM46Orv%2FXg5b6779KVP4tG%2FI6d5w2sy3rLBSU3clZ7oKffhqxjsadCyEI4JCPz0Ub2l3U1OcDprdn3SYiAmkOOmpn8k%2F9sRgTOOfuMeEPMrXSSY8HdcDtnYwK3YiQqoe2qrWBDLOmvfUV1oYYBBZ1NwCykEJz%2B5U%2FcUq5ydOeBYQhvGW0o4MKFUqZDhGVGlqMufUs6n791UpymD1kzRlQRt7unux9jAoC5JTCR4bK0GIqojXqYiRtng%3D%3D&response-content-disposition=attachment%3B+filename%3Dsemantic_map.zip" -c -O 'semantic_map.zip'
# !unzip ./semantic_map.zip
# + id="URx1otO8S8Yz"
# %rm -r semantic_map.zip
# %cd ..
# !pwd
# !ls
# + [markdown] id="ivPxuLK4S9tK"
# # Installing Dependencies
#
# We need to install some of the dependencies in order to process our data and train model.
# + id="n20EXXkES_tE"
# this script transports l5kit and dependencies
# !pip -q install pymap3d==2.1.0
# !pip -q install protobuf==3.12.2
# !pip -q install transforms3d
# !pip -q install zarr
# !pip -q install ptable
# previous version 1.0.6
# !pip -q install --no-dependencies l5kit
# + [markdown] id="mYLJ5n4zWmmy"
# # Import Packages
# + id="u8doy1hJUUdM" outputId="e866b020-274c-494e-97ee-4b860bd00db9" colab={"base_uri": "https://localhost:8080/", "height": 50}
# import packages
import numpy as np
import gc, os, time
from tqdm import tqdm
import albumentations as A
from google.colab import files
from typing import Dict, Callable
# deeplearning Pytorch
import torch
from torch import Tensor, nn, optim
from torch.utils.data import DataLoader
from torchvision.models.resnet import resnet18, resnet34, resnet50
from torch.optim.lr_scheduler import StepLR
from torch.utils.data.dataset import Dataset
# L5 Toolkit
import l5kit
from l5kit.data import LocalDataManager, ChunkedDataset
from l5kit.dataset import AgentDataset, EgoDataset
from l5kit.rasterization import build_rasterizer
print(f'L5 Toolkit Version : {l5kit.__version__}')
print(f'Pytorch Version : {torch.__version__}')
# + id="73VXrQ3rWz4P"
# root directory
INPUT_DIR = '/content/lyft-motion-prediction-autonomous-vehicles/'
# + [markdown] id="bhR0J7nTX4k-"
# # Configurations
# + id="t5rJvsHJX3lt"
cfg = {
'format_version': 4,
'model_params': {
'model_architecture': 'resnet18',
'history_num_frames': 10,
'history_step_size': 1,
'history_delta_time': 0.1,
'future_num_frames': 50,
'future_step_size': 1,
'future_delta_time': 0.1
},
'raster_params': {
'raster_size': [448, 448],
'pixel_size': [0.5, 0.5],
'ego_center': [0.25, 0.5],
'map_type': 'py_semantic',
'satellite_map_key': 'aerial_map/aerial_map.png',
'semantic_map_key': 'semantic_map/semantic_map.pb',
'dataset_meta_key': 'meta.json',
'filter_agents_threshold': 0.5
},
'train_data_loader': {
'key': 'scenes/train.zarr',
'batch_size': 64,
'shuffle': True,
'num_workers': 0
},
'train_params': {
'max_num_steps': 100000,
'image_coords': True,
'verbose': 100,
}
}
# + id="na9-qZ_5ZUX6"
# set env variable for data
os.environ["L5KIT_DATA_FOLDER"] = INPUT_DIR
dm = LocalDataManager(None)
# get config
print(cfg)
# + [markdown] id="49OapdQhh-az"
# # Augmentations
# + id="bUDF0-xNh949"
class TransformDataset(Dataset):
def __init__(self, dataset: Dataset, transform: Callable):
self.dataset = dataset
self.transform = transform
def __getitem__(self, index):
batch = self.dataset[index]
return self.transform(batch)
def __len__(self):
return len(self.dataset)
def transform(batch):
return batch["image"], batch["target_positions"], batch["target_availabilities"]
train_transform = A.Compose([
A.Cutout(max_h_size=int(224 * 0.125), max_w_size=int(224 * 0.125), num_holes=3, p=0.6),
A.Normalize()
])
# + [markdown] id="KdCQuloqaP-Q"
# # Data Loader
# + id="ZACcgClMaQoC"
def get_data(cfg):
'''
Description
----------------
Prepare image data by
getting muti-channel tensors
and converting them to rgb images.
It iterates over other agents annotations
and passes the AgentDataset into Pytorch
Dataloader.
Arguments:
cfg -- Configuration file
Returns:
train_dataloader -- Pytorch dataloader
'''
# rasterizercfg, dm
rasterizer = build_rasterizer()
# Train dataset/dataloader
train_cfg = cfg["train_data_loader"]
train_zarr = ChunkedDataset(dm.require(train_cfg["key"])).open()
train_dataset = AgentDataset(cfg, train_zarr, rasterizer)
train_dataset = TransformDataset(train_dataset, train_transform)
train_dataloader = DataLoader(train_dataset,
shuffle=train_cfg["shuffle"],
batch_size=train_cfg["batch_size"],
num_workers=train_cfg["num_workers"])
return train_dataloader
# + [markdown] id="66TMlEJND7_r"
# # Loss Function: Negative Log Likelihood
# + id="fatGqNZgD7Af"
def negativeLogLikelihood(gt: Tensor, pred: Tensor, confidences: Tensor, avails: Tensor):
'''
Description
-------------------------------
Compute Negative log-likelihood
Arguments:
ground_truth -- array of shape (time)x(2D coords)
prediction -- array of shape (modes)x(time)x(2D coords)
confidences -- array of shape (modes) with a confidence for each mode in each sample
avails -- array of shape (time) with the availability for each gt timestep
Returns:
error -- negative log-likelihood (floating number)
'''
assert len(pred.shape) == 4, f"expected 3D (MxTxC) array for pred, got {pred.shape}"
batch_size, num_modes, future_len, num_coords = pred.shape
assert gt.shape == (batch_size, future_len, num_coords), f"expected 2D (Time x Coords) array for gt, got {gt.shape}"
assert confidences.shape == (batch_size, num_modes), f"expected 1D (Modes) array for gt, got {confidences.shape}"
assert torch.allclose(torch.sum(confidences, dim=1), confidences.new_ones((batch_size,))), "confidences should sum to 1"
assert avails.shape == (batch_size, future_len), f"expected 1D (Time) array for gt, got {avails.shape}"
# assert all data are valid
assert torch.isfinite(pred).all(), "invalid value found in pred"
assert torch.isfinite(gt).all(), "invalid value found in gt"
assert torch.isfinite(confidences).all(), "invalid value found in confidences"
assert torch.isfinite(avails).all(), "invalid value found in avails"
gt = torch.unsqueeze(gt, 1)
avails = avails[:, None, :, None]
error = torch.sum(((gt - pred) * avails) ** 2, dim=-1)
with np.errstate(divide="ignore"):
error = torch.log(confidences) - 0.5 * torch.sum(error, dim = -1)
max_value, _ = error.max(dim = 1, keepdim = True)
error = -torch.log(torch.sum(torch.exp(error - max_value), dim = -1, keepdim=True)) - max_value
return torch.mean(error)
# + [markdown] id="5DyqS_aQEDDS"
# # Model: resnet 18
# + id="QT_ZyHq8EElN"
class LyftMultiModel(nn.Module):
def __init__(self, cfg: Dict, num_modes=3):
super(LyftMultiModel, self).__init__()
architecture = cfg["model_params"]["model_architecture"]
backbone = eval(architecture)(pretrained=True, progress=True)
self.backbone = backbone
num_history_channels = (cfg["model_params"]["history_num_frames"] + 1) * 2
num_in_channels = 3 + num_history_channels
self.backbone.conv1 = nn.Conv2d(
num_in_channels,
self.backbone.conv1.out_channels,
kernel_size=self.backbone.conv1.kernel_size,
stride=self.backbone.conv1.stride,
padding=self.backbone.conv1.padding,
bias=False,
)
if architecture == "resnet50":
backbone_out_features = 2048
else:
backbone_out_features = 512
# X, Y coords for the future positions (output shape: batch_sizex50x2)
self.future_len = cfg["model_params"]["future_num_frames"]
num_targets = 2 * self.future_len
# You can add more layers here.
self.head = nn.Sequential(
nn.Linear(in_features=backbone_out_features, out_features=4096),
)
self.num_preds = num_targets * num_modes
self.num_modes = num_modes
self.logit = nn.Linear(4096, out_features=self.num_preds + num_modes)
def forward(self, x):
x = self.backbone.conv1(x)
x = self.backbone.bn1(x)
x = self.backbone.relu(x)
x = self.backbone.maxpool(x)
x = self.backbone.layer1(x)
x = self.backbone.layer2(x)
x = self.backbone.layer3(x)
x = self.backbone.layer4(x)
x = self.backbone.avgpool(x)
x = torch.flatten(x, 1)
x = self.head(x)
x = self.logit(x)
# pred (batch_size)x(modes)x(time)x(2D coords)
# confidences (batch_size)x(modes)
bs, _ = x.shape
pred, confidences = torch.split(x, self.num_preds, dim=1)
pred = pred.view(bs, self.num_modes, self.future_len, 2)
assert confidences.shape == (bs, self.num_modes)
confidences = torch.softmax(confidences, dim=1)
return pred, confidences
# + id="yn4rHhF0Egno"
def forward(data, model, device, criterion = negativeLogLikelihood):
# USE GPU FOR TRAINING
inputs = data["image"].to(device)
target_availabilities = data["target_availabilities"].to(device)
targets = data["target_positions"].to(device)
# FORWARD PASS TO GET OUTPUT/LOGITS
preds, confidences = model(inputs)
loss = criterion(targets, preds, confidences, target_availabilities)
return loss, preds
# + id="h2aan49I4jqX"
# Used in V3
def forward_v3(data, model, device, criterion = negativeLogLikelihood):
# USE GPU FOR TRAINIG
inputs = data["image"].to(device)
target_availabilities = data["target_availabilities"].to(device)
targets = data["target_positions"].to(device)
matrix = data["world_to_image"].to(device)
centroid = data["centroid"].to(device)[:,None,:].to(torch.float)
# FORWARD PASS TO GET OUTPUT/LOGITS
outputs = model(inputs)
bs,tl,_ = targets.shape
assert tl == cfg["model_params"]["future_num_frames"]
if cfg['train_params']['image_coords']:
targets = targets + centroid
targets = torch.cat([targets,torch.ones((bs,tl,1)).to(device)], dim=2)
targets = torch.matmul(matrix.to(torch.float), targets.transpose(1,2))
targets = targets.transpose(1,2)[:,:,:2]
bias = torch.tensor([56.25, 112.5])[None,None,:].to(device)
targets = targets - bias
confidences, pred = outputs[:,:3], outputs[:,3:]
pred = pred.view(bs, 3, tl, 2)
assert confidences.shape == (bs, 3)
confidences = torch.softmax(confidences, dim=1)
# CALCULATE LOSS: NEGATIVE LOG LIKELIHOOD
loss = criterion(targets, pred, confidences, target_availabilities)
loss = torch.mean(loss)
if cfg['train_params']['image_coords']:
matrix_inv = torch.inverse(matrix)
pred = pred + bias[:,None,:,:]
pred = torch.cat([pred,torch.ones((bs,3,tl,1)).to(device)], dim=3)
pred = torch.stack([torch.matmul(matrix_inv.to(torch.float), pred[:,i].transpose(1,2))
for i in range(3)], dim=1)
pred = pred.transpose(2,3)[:,:,:,:2]
pred = pred - centroid[:,None,:,:]
return loss, pred, confidences
# + id="segKq6UeadH-"
# INSTANTIATE MODEL CLASS
model = LyftMultiModel(cfg)
# USE GPU FOR TRAINING MODEL
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
# GPU FOR MODEL
model.to(device)
# INSTANTIATE OPTIMIZER CLASS
optimizer = optim.Adam(model.parameters(), lr=1e-3)
# + [markdown] id="ESyAw4KasqNa"
# # Checkpoint
#
# Run this cell when you want to resume training your model after certain number of epochs. Uncomment all the lines if you have saved weights earlier to resume training.
# + id="Vst3EJ-ScrDh"
# WEIGHT_FILE = '/content/drive/My Drive/Kaggle/Lyft L5 Motion Prediction/resnet18_448x448_model_state_v3_1400.pth'
# checkpoint = torch.load(WEIGHT_FILE, map_location=device)
# epoch = checkpoint['epoch']
# model.load_state_dict(checkpoint['model_state_dict'])
# optimizer.load_state_dict(checkpoint['optimizer_state_dict'])
# + [markdown] id="2bs2RpyrtEkv"
# # Train Model
# + id="qUdbWBNWs3rb"
def train(cfg, device, model, optimizer):
losses_train = []
print('getting Dataloader...')
train_dataloader = get_data(cfg)
progress_bar = tqdm(range(epoch, cfg['train_params']['max_num_steps']))
start = time.time()
print('Training...')
for i in progress_bar:
try:
data = next(iter(train_dataloader))
except StopIteration:
data = next(iter(train_dataloader))
model.train()
torch.set_grade_enabled(True)
# CLEAR GRADIENTS W.R.T TO PARAMETERS
optimizer.zero_grad()
# FORWARD PASS TO GET OUTPUT
loss, _ = forward(data, model, device)
# GETTING GRADIENTS W.R.T TO PARAMETERS
loss.backward()
# UPDATING PARAMETERS
optimizer.step()
# Appending losses in a list
losses_train.append(loss.item())
if i % cfg['train_params']['verbose'] == 0:
torch.save({'epoch': i + 1,
'model_state_dict': model.state_dict(),
'optimizer_state_dict': optimizer.state_dict()},
f'/content/drive/My Drive/Kaggle/Lyft L5 Motion Prediction/resnet18_448x448_model_state_v4_{i}.pth')
progress_bar.set_description(f'loss: {loss.item()} loss_avg: {np.mean(losses_train)}')
# + [markdown] id="sF1d1ufiy0WA"
# # Training Model
# + id="B9mYGlBjyo1U"
train(cfg, device, model, optimizer)
| colab/multimode/Lyft_L5_V4_448x448.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: geocomp
# language: python
# name: geocomp
# ---
# # An introduction to Interact
#
# "The interact function (ipywidgets.interact) automatically creates user interface (UI) controls for exploring code and data interactively. It is the easiest way to get started using IPython’s widgets." [Using Interact Documentation](https://ipywidgets.readthedocs.io/en/stable/examples/Using%20Interact.html)
import matplotlib.pyplot as plt
# ## Why would we want to use Interact in the first place?
#
# An _interactive_ figure speaks a thousand words.
# +
# # %load plotter.py
"""
Required for the ipywidgets demo notebook.
"""
import numpy as np
import matplotlib.pyplot as plt
from ipywidgets import interact, interactive, fixed, interact_manual
import ipywidgets as widgets
params = dict(
colormap=['viridis', 'plasma', 'inferno', 'magma', 'Greys', 'Greys_r'],
section=widgets.RadioButtons(options=['inline', 'xline', 'timeslice'],
value='inline',description='slicer',disabled=False),
inline=widgets.IntSlider(value=300,min=0,max=600,step=1,
continuous_update=False,description='<font color="red">inline</>'),
xline=widgets.IntSlider(value=240,min=0,max=480,step=1,
continuous_update=False,description='<font color="green">xline</>'),
timeslice=widgets.IntSlider(value=125,min=0,max=250,step=1,
continuous_update=False,description='<font color="blue">timeslice</>'),
)
@interact(**params)
def seismic_plotter(colormap, section, inline, xline, timeslice):
"""
Plot a chosen seismic iLine, xLine or timeslice with a choice of colormap.
"""
vol = np.load('../../data/Penobscot_0-1000ms.npy')
# Sections dictionary.
sections = {
'inline': {'amp': vol[inline,:,:].T, 'line': inline, 'shrink_val': 0.6,
'axhline_y': timeslice, 'axhline_c': 'b',
'axvline_x': xline, 'axvline_c': 'g',
'axspine_c': 'r'},
'xline': {'amp': vol[:,xline,:].T, 'line': xline, 'shrink_val': 0.5,
'axhline_y': timeslice, 'axhline_c': 'b',
'axvline_x': inline, 'axvline_c': 'r',
'axspine_c': 'g'},
'timeslice': {'amp': vol[:,:,timeslice], 'line': timeslice, 'shrink_val': 0.95,
'axhline_y': xline, 'axhline_c': 'g',
'axvline_x': inline, 'axvline_c': 'r',
'axspine_c': 'b'},
}
# Scale amplitudes.
ma = np.percentile(vol, 98)
# Plot figure.
fig, ax = plt.subplots(figsize=(18, 6), ncols=1)
sec = sections[section]
im = ax.imshow(sec['amp'], aspect=0.5, vmin=-ma, vmax=ma, cmap=colormap)
ax.set_title(f'Penobscot_0-1000ms {section} {sec["line"]}')
plt.colorbar(im, ax=ax, shrink=sec['shrink_val']).set_label(colormap)
# Add projected lines.
ax.axhline(y=sec['axhline_y'], linewidth=2, color=sec['axhline_c'])
ax.axvline(x=sec['axvline_x'], linewidth=2, color=sec['axvline_c'])
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(2)
ax.spines[axis].set_color(sec['axspine_c'])
plt.show()
return
# -
# ## Basic interact
#
# Let's jump in: for a basic function `vp_from_dt` that prints out `Vp` for a given `DT`, we can construct a simple slider like so:
def vp_from_dt(dt):
vp = 10e6 / dt
return print(f'vp = {vp:.2f}')
# +
from ipywidgets import interact
interact(vp_from_dt, dt=3500)
# -
# For a similar function that requires a `boolean`, we can make a toggle, also notice that by saving the `interact()` function to a `name`, we suppress the output `<function __main__.convert_vp_to_dt(vp, convert_to_dt)>`.
#
# We have also created a dropdown list automatically by passing an `iterable` to the `interact()` function.
# +
import numpy as np
vp = np.arange(2400, 2750, 50)
def convert_vp_to_dt(vp, convert_to_dt):
if convert_to_dt:
output = 1e6/vp
else:
output = vp
return print(f'The result is {output:.2f}')
# -
my_boolean = interact(convert_vp_to_dt, vp=vp, convert_to_dt=True)
# If we need to get input from the user, we can use an `input box`, (see [here](https://stackoverflow.com/questions/35361038/using-ipython-ipywidget-to-create-a-variable) for example):
#
# - We first define variables for the `input` and `saved` variables
# - Then we define a `get_input()` function that will assign the input value to the saved variable
# - Next we call the `on_submit()` function on the `input variable`, passing it the `get_input()` function
# + tags=["hide"]
import ipywidgets as widgets
input_string = widgets.Text(placeholder='Please type something in this box')
saved_string = widgets.Text()
def get_input(input_text):
"""bind the input text to a variable"""
saved_string.value = input_string.value
return saved_string
# + tags=["hide"]
input_string.on_submit(get_input)
# -
# We can now use the `input_variable` to see an input box:
input_string
# And the `output_variable` will be updated `on_submit()`:
saved_string
# We now have access to the `output_variable` and can saved it to a string for manipulation:
my_string = saved_string.value
my_string.upper()
# Also note that the `saved_variable` remains bound to the `input_variable`, so if you type something else in the `input_variable` box, the `saved_variable` is updated. The reverse is _not_ true however.
# ### There are many widget types available:
# - [Numeric widgets](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Numeric-widgets)
# - [Boolean widgets](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Boolean-widgets)
# - [Selection widgets](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Selection-widgets)
# - [String widgets](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#String-widgets)
# - [Image](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Image)
# - [Button](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Button)
# - [Output](https://ipywidgets.readthedocs.io/en/latest/examples/Output%20Widget.html)
# - [Play (Animation) widget](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Play-(Animation)-widget)
# - [Date picker](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Date-picker)
# - [Color picker](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Color-picker)
# - [Controller](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Controller)
# - [Container/Layout widgets](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Container/Layout-widgets)
#
# Almost all have different `keyword arguments` that can be set as in the `IntSlider` example below:
widgets.IntSlider(
value=12,
min=0,
max=100,
step=1,
description='Slider:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='d'
)
# + [markdown] tags=["exe"]
# ### Exercise
# Try to replicate the range slider below using a min of 0 and a max of 20.
# Once you've got it working, see what changes you can make to it.
#
# <img src='../data/range-slider.png'/>
#
# If you don't know where to start, you'll find all the widgets [here](https://ipywidgets.readthedocs.io/en/stable/examples/Widget%20List.html#Widget-List).
# +
# Your code here.
# -
# + tags=["hide"]
widgets.IntRangeSlider(
value=[3, 12],
min=0,
max=20,
step=1,
description='Range:',
disabled=False,
continuous_update=False,
orientation='horizontal',
readout=True,
readout_format='d',
)
# -
# ## Interact usage
#
# `interact` can also be used as a `decorator`. Interact [decorators](https://wiki.python.org/moin/PythonDecorators#What_is_a_Decorator) allow you expand the functionality of your function and interact with it in a single shot. As this `square_or_double()` example function shows, interact also works with functions that have multiple arguments. [source](https://ipywidgets.readthedocs.io/en/stable/examples/Using%20Interact.html#Basic-interact)
# +
method = widgets.RadioButtons(options=['squared','doubled'],description='option')
y = widgets.IntSlider(value=5,min=0,max=10,step=1,description='y')
@interact(method=method, y=y)
def square_or_double(method, y):
if method == 'squared':
result = y**2
else:
result = y*2
return print(f'{y} {method} = {result}')
# + [markdown] tags=["exe"]
# ### Exercise
# Write a function that returns `a` to the power of `b` but use the interact decorator to make both `a` and `b` interactive (between 0 and 10 in steps of 1), add a toggle to negate the result.
# +
# Your code here.
# -
# + tags=["hide"]
@interact(a=widgets.IntSlider(value=2,min=0,max=10,step=1,description='a'),
b=widgets.IntSlider(value=4,min=0,max=10,step=1,description='b'),
negate=widgets.Checkbox(value=False,description='negate'))
def pow_a_b(a, b, negate):
"""return a to the power of b or negative a**b"""
if negate:
out = -a**b
else:
out = a**b
return out
# + tags=["hide"]
@interact(a=(0,10,1), b=(0,10,1), negate=False)
def pow_a_b(a, b, negate):
"""return a to the power of b or negative a**b"""
if negate:
out = -a**b
else:
out = a**b
return out
# -
# ### Worked Example
#
# Let's build an example of an interactive wavelet using [Bruges](https://github.com/agile-geoscience/bruges), we'll use:
# - [Ricker](https://github.com/agile-geoscience/bruges/blob/master/bruges/filters/wavelets.py)
# - [Gabor](https://github.com/agile-geoscience/bruges/blob/master/bruges/filters/wavelets.py)
# - [sinc](https://github.com/agile-geoscience/bruges/blob/master/bruges/filters/wavelets.py)
# - [cosine](https://github.com/agile-geoscience/bruges/blob/master/bruges/filters/wavelets.py)
# +
from bruges.filters.wavelets import ricker, gabor, sinc, cosine
w, t = ricker(duration=0.128, dt=0.001, f=25, return_t=True)
fig, ax = plt.subplots(figsize=(15, 6), ncols=1)
ax.plot(t, w)
ax.grid()
ax.set_title(f'ricker wavelet - frequency=25')
plt.show()
# + [markdown] tags=["exe"]
# ### Exercise
# Let's turn this into an interactive function:
# - first define a function
# - copy the code above into that function
# - use an interact decorator and widget to have frequency by a slider (allow a range from 1Hz to 75Hz in steps of 1Hz)
#
# Remember to correct the title.
# +
# Your code here.
# -
# + tags=["hide"]
@interact(frequency=widgets.IntSlider(value=25,min=1,max=75,step=1))
def plot_filter(frequency):
w, t = ricker(duration=0.128, dt=0.001, f=frequency, return_t=True)
fig, ax = plt.subplots(figsize=(15, 6), ncols=1)
ax.plot(t, w)
ax.grid()
ax.set_title(f'ricker wavelet - frequency={frequency}')
plt.show()
return
# + [markdown] tags=["exe"]
# ### Exercise
# Now let's allow the user to pass both duration _and_ dt as interactive arguments, using your code above:
# - add two more arguments to the function
# - define these arguments `duration` and `dt` as `Interact.widgets`
#
# For `duration` use a value 0.256 seconds with a minimum of 0.04 seconds, a maximum of 0.512 seconds and steps of 0.004 seconds.
#
# For `dt` use a value 0.001 seconds with a minimum of 0.0001 seconds, a maximum of 0.008 seconds and steps of 0.0001 seconds. You may want to use the `**kwargs` `readout_format='.4f'` for `dt`.
#
# N.B.: you can optionally add `continuous_update=False` to the arguments of your `widgets` in order to avoid 'choppy' display when you move the sliders.
# +
# Your code here.
# -
# + tags=["hide"]
@interact(frequency=widgets.IntSlider(value=25,min=1,max=75,step=1,continuous_update=False),
duration=widgets.FloatSlider(value=0.256,min=0.04,max=0.512,step=0.004,continuous_update=False),
dt=widgets.FloatSlider(value=0.001,min=0.0001,max=0.008,
step=0.0001,continuous_update=False,
readout_format='.4f'))
def plot_filter(frequency,duration,dt):
w, t = ricker(duration=duration, dt=dt, f=frequency, return_t=True)
fig, ax = plt.subplots(figsize=(15, 6), ncols=1)
ax.plot(t, w)
ax.grid()
ax.set_title(f'ricker wavelet - frequency={frequency}')
plt.show()
return
# + [markdown] tags=["exe"]
# ### Exercise
# Now let's see if we can fill the wavelet between zero and positive values of the wavelet, for this you can use the matplotlib function `.fill_between()`, you might need to read the [docs](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.fill_between.html) or look at an [example](https://matplotlib.org/examples/pylab_examples/fill_between_demo.html) to figure out how to use this function.
# +
# your code here
# -
# + tags=["hide"]
@interact(frequency=widgets.IntSlider(value=25,min=1,max=75,step=1,continuous_update=False),
duration=widgets.FloatSlider(value=0.256,min=0.04,max=0.512,step=0.004,continuous_update=False),
dt=widgets.FloatSlider(value=0.001,min=0.0001,max=0.008,
step=0.0001,continuous_update=False,
readout_format='.4f'),
filled=widgets.Checkbox(value=True,description='fill wavelet',disabled=False)
)
def plot_filter(frequency,duration,dt,filled):
w, t = ricker(duration=duration, dt=dt, f=frequency, return_t=True)
fig, ax = plt.subplots(figsize=(15, 6), ncols=1)
ax.plot(t, w)
ax.grid()
ax.set_title(f'ricker wavelet - frequency={frequency}')
# define fill_between() parameters
x_min = -duration / 2
x_max = duration / 2
x = np.arange(x_min, x_max, dt)
if filled:
ax.fill_between(x, 0, w, where=w > 0, color='k')
plt.show()
return
# + [markdown] tags=["exe"]
# ### Exercise
# Finally, let's see if we can add a choice of wavelets to the function, so that the user can choose between 'ricker', 'gabor', 'sinc' and 'cosine' for example (these all have the same input parameters), there are different ways to achieve this for example using a `ToggleButtons` or a `Select` widget.
#
# Once again, remember to correct the title.
# +
# your code here
# -
# + tags=["hide"]
FUNCS={'ricker': ricker,'gabor': gabor,'sinc': sinc,'cosine': cosine}
@interact(wavelet=widgets.ToggleButtons(options=FUNCS,description='wavelet',button_style='success'),
frequency=widgets.IntSlider(value=25,min=1,max=75,step=1,continuous_update=False),
duration=widgets.FloatSlider(value=0.256,min=0.04,max=0.512,step=0.004,continuous_update=False),
dt=widgets.FloatSlider(value=0.001,min=0.0001,max=0.008,
step=0.0001,continuous_update=False,
readout_format='.4f'),
filled=widgets.Checkbox(value=True,description='fill wavelet',disabled=False)
)
def plot_filter(wavelet, frequency, duration, dt, filled):
w, t = wavelet(duration=duration, dt=dt, f=frequency, return_t=True)
fig, ax = plt.subplots(figsize=(15, 6), ncols=1)
ax.plot(t, w)
ax.grid()
ax.set_title(f'{wavelet.__name__} wavelet - frequency={frequency}')
# define fill_between() parameters
x_min = -duration / 2
x_max = duration / 2
x = np.arange(x_min, x_max, dt)
if filled:
ax.fill_between(x, 0, w, where=w > 0, color='k')
plt.show()
return
# + tags=["hide"]
# A final version with all formatting
FUNCS={'ricker': ricker,'gabor': gabor,'sinc': sinc,'cosine': cosine}
@interact(wavelet=widgets.ToggleButtons(options=FUNCS.keys(),description='wavelet',button_style='success'),
duration=widgets.FloatSlider(value=0.256,min=0.04,max=0.512,step=0.004,
description='duration',
continuous_update=False,
readout_format='.3f'),
dt=widgets.FloatSlider(value=0.001,min=0.0001,max=0.008,step=0.0001,
description='dt',
continuous_update=False,
readout_format='.4f'),
frequency=widgets.IntSlider(value=25,min=1,max=75,step=1,
description='frequency',
continuous_update=False,
readout_format='d'),
filled=widgets.Checkbox(value=True,description='fill wavelet',disabled=False)
)
def plot_filter(wavelet, duration, dt, frequency, filled):
"""
Plot a filter:
Args:
function (function): one of ['ricker', 'gabor', 'sinc', 'cosine']
duration (float): The length in seconds of the wavelet.
dt (float): The sample interval in seconds.
frequency (ndarray): Dominant frequency of the wavelet in Hz.
fill (boolean): whether the filter plot is filled between 0 and wavelet.
Returns:
ndarray. {function} wavelet with centre frequency 'frequency' sampled on t.
"""
# call the wavelet function
w, t = FUNCS[wavelet](duration, dt, f=frequency, return_t=True)
# create the plot
fig, ax = plt.subplots(figsize=(15, 6), ncols=1)
ax.plot(t, w, color='black')
ax.grid()
ax.set_title(f'{wavelet} wavelet, frequency={frequency}, duration={duration}, dt={dt}')
# define fill_between() parameters
x_min = -duration / 2
x_max = duration / 2
x = np.arange(x_min, x_max, dt)
# fill wavelet
if filled:
ax.fill_between(x, 0, w, where=w > 0, color='k')
# show the plot
plt.show()
return
# -
# ### Summary
#
# Let's summarise by looking at the initial reason we looked at interact:
@interact(
colormap=['viridis', 'plasma', 'inferno', 'magma', 'Greys', 'Greys_r'],
section=widgets.RadioButtons(options=['inline', 'xline', 'timeslice'],
value='inline',description='slicer',disabled=False),
inline=widgets.IntSlider(value=300,min=0,max=600,step=1,
continuous_update=False,description='<font color="red">inline</>'),
xline=widgets.IntSlider(value=240,min=0,max=480,step=1,
continuous_update=False,description='<font color="green">xline</>'),
timeslice=widgets.IntSlider(value=125,min=0,max=250,step=1,
continuous_update=False,description='<font color="blue">timeslice</>'),
)
def seismic_plotter(colormap, section, inline, xline, timeslice):
"""Plot a chosen seismic ILine, XLine or Timeslice with a choice of colormaps"""
# load a volume
vol = np.load('data/Penobscot_0-1000ms.npy')
# sections dictionary
sections = {
'inline': {'amp': vol[inline,:,:].T, 'line': inline, 'shrink_val': 0.6,
'axhline_y': timeslice, 'axhline_c': 'b',
'axvline_x': xline, 'axvline_c': 'g',
'axspine_c': 'r'},
'xline': {'amp': vol[:,xline,:].T, 'line': xline, 'shrink_val': 0.5,
'axhline_y': timeslice, 'axhline_c': 'b',
'axvline_x': inline, 'axvline_c': 'r',
'axspine_c': 'g'},
'timeslice': {'amp': vol[:,:,timeslice], 'line': timeslice, 'shrink_val': 0.95,
'axhline_y': xline, 'axhline_c': 'g',
'axvline_x': inline, 'axvline_c': 'r',
'axspine_c': 'b'},
}
# scale amplitudes
ma = np.percentile(vol, 98)
# plot figure
fig, ax = plt.subplots(figsize=(18, 6), ncols=1)
sec = sections[section]
im = ax.imshow(sec['amp'], aspect=0.5, vmin=-ma, vmax=ma, cmap=colormap)
ax.set_title(f'Penobscot_0-1000ms {section} {sec["line"]}')
plt.colorbar(im, ax=ax, shrink=sec['shrink_val']).set_label(colormap)
# add projected lines
ax.axhline(y=sec['axhline_y'], linewidth=2, color=sec['axhline_c'])
ax.axvline(x=sec['axvline_x'], linewidth=2, color=sec['axvline_c'])
for axis in ['top','bottom','left','right']:
ax.spines[axis].set_linewidth(2)
ax.spines[axis].set_color(sec['axspine_c'])
plt.show()
return
# <hr />
#
# <div>
# <img src="https://avatars1.githubusercontent.com/u/1692321?s=50"><p style="text-align:center">© Agile Geoscience 2019</p>
# </div>
| Interactive_widgets_in_Jupyter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Widget Example Using Ginga
# See https://astrowidgets.readthedocs.io for additional details about the widget, including installation notes.
from astrowidgets import ImageWidget
# +
from ginga.misc.log import get_logger
logger = get_logger('my viewer', log_stderr=True,
log_file=None, level=30)
# -
w = ImageWidget(logger=logger)
# For this example, we use an image from Astropy data repository and load it as `CCDData`. Feel free to modify `filename` to point to your desired image.
#
# Alternately, for local FITS file, you could load it like this instead:
# ```python
# w.load_fits(filename, numhdu=numhdu)
# ```
# Or if you wish to load a data array natively (without WCS):
# ```python
# from astropy.io import fits
# with fits.open(filename, memmap=False) as pf:
# arr = pf[numhdu].data.copy()
# w.load_array(arr)
# ```
# +
filename = 'http://data.astropy.org/photometry/spitzer_example_image.fits'
numhdu = 0
# Loads NDData
# NOTE: memmap=False is needed for remote data on Windows.
# NOTE: Some file also requires unit to be explicitly set in CCDData.
from astropy.nddata import CCDData
ccd = CCDData.read(filename, hdu=numhdu, format='fits')
w.load_nddata(ccd)
# -
# Ginga key bindings documented at http://ginga.readthedocs.io/en/latest/quickref.html . Note that not all documented bindings would work here. Please use an alternate binding, if available, if the chosen one is not working.
#
# Here are the ones that worked during testing with Firefox 52.8.0 on RHEL7 64-bit:
#
# Key | Action | Notes
# --- | --- | ---
# `+` | Zoom in |
# `-` | Zoom out |
# Number (0-9) | Zoom in to specified level | 0 = 10
# Shift + number | Zoom out to specified level | Numpad does not work
# ` (backtick) | Reset zoom |
# Space > `q` > arrow | Pan |
# ESC | Exit mode (pan, etc) |
# `c` | Center image
# Space > `d` > up/down arrow | Cycle through color distributions
# Space > `d` > Shift + `d` | Go back to linear color distribution
# Space > `s` > Shift + `s` | Set cut level to min/max
# Space > `s` > Shift + `a` | Set cut level to 0/255 (for 8bpp RGB images)
# Space > `s` > up/down arrow | Cycle through cuts algorithms
# Space > `l` | Toggle no/soft/normal lock |
#
# *TODO: Check out Contrast Mode next*
# A viewer will be shown after running the next cell.
# In Jupyter Lab, you can split it out into a separate view by right-clicking on the viewer and then select
# "Create New View for Output". Then, you can drag the new
# "Output View" tab, say, to the right side of the workspace. Both viewers are connected to the same events.
w
# This next cell captures print outputs. You can pop it out like the viewer above. It is very convenient for debugging purpose.
# Capture print outputs from the widget
display(w.print_out)
# The following cell changes the visibility or position of the cursor info bar.
#
w.cursor = 'top' # 'top', 'bottom', None
print(w.cursor)
# The rest of the calls demonstrate how the widget API works. Comment/uncomment as needed. Feel free to experiment.
# Programmatically center to (X, Y) on viewer
w.center_on((1, 1))
# Programmatically offset w.r.t. current center
w.offset_to(10, 10)
# +
from astropy.coordinates import SkyCoord
# Change the values here if you are not using given
# example image.
ra_str = '01h13m23.193s'
dec_str = '+00d12m32.19s'
frame = 'icrs'
# Programmatically center to SkyCoord on viewer
w.center_on(SkyCoord(ra_str, dec_str, frame=frame))
# +
# Change the values if needed.
deg_offset = 0.001
# Programmatically offset (in degrees) w.r.t.
# SkyCoord center
w.offset_to(deg_offset, deg_offset, skycoord_offset=True)
# -
# Show zoom level
print(w.zoom_level)
# Programmatically zoom image on viewer
w.zoom(2)
# Capture what viewer is showing and save RGB image.
# Need https://github.com/ejeschke/ginga/pull/688 to work.
w.save('test.png')
# Get all available image stretch options
print(w.stretch_options)
# Get image stretch algorithm in use
print(w.stretch)
# Change the stretch
w.stretch = 'histeq'
print(w.stretch)
# Get all available image cuts options
print(w.autocut_options)
# Get image cut levels in use
print(w.cuts)
# Change the cuts by providing explicit low/high values
w.cuts = (0, 100)
print(w.cuts)
# Change the cuts with an autocut algorithm
w.cuts = 'zscale'
print(w.cuts)
# This enables click to center.
w.click_center = True
# Now, click on the image to center it.
# Turn it back off so marking (next cell) can be done.
w.click_center = False
# This enables marking mode.
w.start_marking()
print(w.is_marking)
# Now, click on the image to mark a point of interest.
# When done, set back to False.
w.stop_marking()
print(w.is_marking)
# +
# Get table of markers
markers_table = w.get_markers(marker_name='all')
# Default display might be hard to read, so we do this
print('{:^8s} {:^8s} {:^28s}'.format(
'X', 'Y', 'Coordinates'))
for row in markers_table:
c = row['coord'].to_string('hmsdms')
print('{:8.2f} {:8.2f} {}'.format(
row['x'], row['y'], c))
# -
# Erase markers from display
w.reset_markers()
# The following works even when we have set `w.is_marking=False`. This is because `w.is_marking` only controls the interactive marking and does not affect marking programmatically.
# Programmatically re-mark from table using X, Y.
# To be fancy, first 2 points marked as bigger
# and thicker red circles.
w.marker = {'type': 'circle', 'color': 'red', 'radius': 50,
'linewidth': 2}
w.add_markers(markers_table[:2])
# You can also change the type of marker to cross or plus
w.marker = {'type': 'cross', 'color': 'cyan', 'radius': 20}
w.add_markers(markers_table[2:])
# +
# Erase them again
w.reset_markers()
# Programmatically re-mark from table using SkyCoord
w.add_markers(markers_table, use_skycoord=True)
# -
# Start marking again
w.start_marking()
print(w.is_marking)
# Stop marking AND clear markers.
# Note that this deletes ALL of the markers
w.stop_marking(clear_markers=True)
print(w.is_marking)
# The next cell randomly generates some "stars" to mark. In the real world, you would probably detect real stars using `photutils` package.
# +
import numpy as np
from astropy.table import Table
# Maximum umber of "stars" to generate randomly.
max_stars = 1000
# Number of pixels from edge to avoid.
dpix = 20
# Image from the viewer.
img = w._viewer.get_image()
# Random "stars" generated.
bad_locs = np.random.randint(
dpix, high=img.shape[1] - dpix, size=[max_stars, 2])
# Only want those not near the edges.
mask = ((dpix < bad_locs[:, 0]) &
(bad_locs[:, 0] < img.shape[0] - dpix) &
(dpix < bad_locs[:, 1]) &
(bad_locs[:, 1] < img.shape[1] - dpix))
locs = bad_locs[mask]
# Put them in table
t = Table([locs[:, 1], locs[:, 0]], names=('x', 'y'))
print(t)
# -
# Mark those "stars" based on given table with X and Y.
w.add_markers(t)
# The following illustrates how to control number of markers displayed using interactive widget from `ipywidgets`.
# +
# Set the marker properties as you like.
w.marker = {'type': 'circle', 'color': 'red', 'radius': 10,
'linewidth': 2}
# Define a function to control marker display
def show_circles(n):
"""Show and hide circles."""
w.reset_markers()
t2show = t[:n]
w.add_markers(t2show)
with w.print_out:
print('Displaying {} markers...'.format(len(t2show)))
# -
# We redisplay the image widget below above the slider. Note that the slider affects both this view of the image widget and the one near the top of the notebook.
# +
from IPython.display import display
import ipywidgets as ipyw
from ipywidgets import interactive
# Show the slider widget.
slider = interactive(show_circles,
n=ipyw.IntSlider(min=0,max=len(t),step=1,value=0, continuous_update=False))
display(w, slider)
# -
# Now, use the slider. The chosen `n` represents the first `n` "stars" being displayed.
| example_notebooks/ginga_widget.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
def zombies:
"""
SIR Zombie Model aka Zombie Apocalypse Model
s'(t) = h - beta*s*z - delta(s)*s
I'(t) = beta*s*z - ro*I - delta(I)*I
Z'(t) = ro*I - alpha*s*z
D'(t) = delta(s)*s + delta(I)*I + alpha*s*z
s'(t): takes into account those who are suseptible to get infected with the zombie virus
h : number of humans we begin with
beta*s*z : Those infected with zombie virus
delta(s)*s : Those who are dead/killed
I'(t): Takes into account those are infected
ro*I : Number of infected rate
beta*s*z : Those infected with zombie virus
delta(I)*I : When people are killed/dead
Z'(t): Takes into account those are NOW zombies
ro*I : Number of infected rate
alpha*s*z : human and zombie interaction. Humans killing zombies
D'(t): Those who are dead
delta(s)*s : Those who are dead/killed
delta(I)*I : When people are killed/dead
alpha*s*z : human and zombie interaction. Humans killing zombies
"""
| .ipynb_checkpoints/Daisy's changes-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.3.7
# language: julia
# name: julia 0.3
# ---
using DataFrames
using JSON
using Iterators
using taxis
using HDF5, JLD
using Stats
using kNN
using sequenceCompare
#reload("taxis")
#reload("sequenceCompare")
nprocs()
taxi_df, taxi_validation_df = taxis.LoadData("/home/tony/ML/taxi/taxi2_time/train_200k.csv", "/home/tony/ML/taxi/taxi2_time/test.csv")
0
train_coords = taxi_df[:COORDS_TEST][1:4]
#train_lens = [c[:,1:int(ceil(rand(1)[1]*size(c,2)))] for c in train_coords]
# +
println("looking at taxi id information")
head(taxi_df)
taxi_ids = taxi_df[:TAXI_ID]
taxi_ids_dict = Dict{Int64, Int64}()
for id in taxi_ids
taxi_ids_dict[id] = get(taxi_ids_dict, id, 0) + 1
end
taxi_id_counts = [x::Int64 for x in values(taxi_ids_dict)]
describe(taxi_id_counts)
println("number less than 10: ", sum(taxi_id_counts .< 20))
println("number of taxi ids: ", length(keys(taxi_ids_dict)))
#taxis_by_id = [id => taxi_df[taxi_df[:TAXI_ID].==id,:] for id in keys(taxi_ids_dict)]
val_ids = [get(taxi_ids_dict, x, 0)::Int64 for x in taxi_validation_df[:TAXI_ID]]
sum(val_ids .== 0)
# -
println("finding unique number of coords")
all_coords_val = hcat(taxi_validation_df[:COORDS]...)
all_coords = hcat(taxi_df[:COORDS]...)
# # Creating coord dict
small_taxi_df = GetTableOrderedSubset(taxi_df, 190000)
coordsDB = ConstructCoordsDatabase(small_taxi_df, 2)
describe([length(x)::Int64 for x in values(coordsDB)])
taxi_df[:GRID_START] = [round(c,2) for c in taxi_df[:START]]
taxi_validation_df[:GRID_START] = [round(c,2) for c in taxi_validation_df[:START]]
taxi_df[:GRID_END] = [round(c,2) for c in taxi_df[:END]]
taxi_validation_df[:GRID_END] = [round(c,2) for c in taxi_validation_df[:END]]
# +
function getGridDicts(train_df, test_df, grid_delta=.01)
train_df[:GRID_START] = [round(c,2) for c in train_df[:START]]
test_df[:GRID_START] = [round(c,2) for c in test_df[:START] ]
train_df[:GRID_END] = [round(c,2) for c in train_df[:END] ]
test_df[:GRID_END] = [round(c,2) for c in test_df[:END] ]
grids_dict = Dict()
for i in 1:length(train_df[:GRID_START])
coord_pair = train_df[:GRID_START][i]
res = get(grids_dict, coord_pair, Any[])
push!(res, train_df[:COORDS][i])
grids_dict[coord_pair] = res
end
mean_length_dict = Dict()
mean_dest_dict = Dict()
for grid_coord in keys(grids_dict)
paths = grids_dict[grid_coord]
endpoints = [x[:,end] for x in paths]
path_lens = [int(length(x)/2)::Int64 for x in paths]
#println("hey")
mean_dest_dict[grid_coord] = mean(endpoints)
#println("2222")
mean_length_dict[grid_coord] = round(mean(path_lens))
end
return mean_length_dict, mean_dest_dict
end
mean_length_dict, mean_dest_dict = getGridDicts()
# -
values(mean_length_dict)
# +
pairs = collect(keys(grids_dict))
grid_coord = pairs[1]
p = grids_dict[grid_coord]
println("num paths: ", length(p))
endpoints = [x[:,end] for x in p]
paths_lens = [int(length(x)/2)::Int64 for x in p]
println("endpoiktns and paths")
println(mean(paths_lens))
println(length(p[1]))
for x in p
println(length(x))
end
println(mean(endpoints))
# -
sum([haskey(grids_dict, x) for x in taxi_validation_df[:GRID_START]])
mean([c for c in train_df[:END][1:5]])
median(train_df[:NUM_COORDS][1:5])
# #Creating new features
#
# +
function GetDateInfo(df)
if haskey(df, :DAYOFWEEK)
return df
end
function GetDistanceData(df)
if haskey(df, :DISTANCE)
return df
end
# -
# # Training models
# +
function euclideanDist(p1, p2)
return sqrt((p1[1]-p2[1])^2 + (p1[2]-p2[2])^2)
end
function findClosestTrainingExample2(all_train_coords, test_path, w=1)
num_paths = length(all_train_coords)
best_dist = 9999.0
best_path = all_train_coords[1]
for k=1:num_paths
train_path = all_train_coords[k]
if w <= 1
dist = DTWDistance(train_path, test_path)
else
dist = DTWDistance(train_path, test_path, w)
end
#println("k=", k, " dist=", dist)
if dist < best_dist
#println(k, ", old best: ", best_dist, " new best: ", dist)
best_dist = dist
best_path = all_train_coords[k]
end
end
return best_path
end
# Create a DTW/START_DIFF metric
function findClosestTrainingExample3(all_train_coords, test_path, w=1)
num_paths = length(all_train_coords)
best_score = 999999.0
best_path = all_train_coords[1]
for k=1:num_paths
train_path = all_train_coords[k]
test_path = test_path[:,1:min(end, size(train_path,2)+2)]
if size(train_path,2) < size(test_path, 2)
continue
end
if w <= 1
dist = DTWDistance(train_path, test_path)
else
dist = DTWDistance(train_path, test_path, w)
end
start_diff = float(euclideanDist(test_path[:,1], train_path[:,1])+0.000001)
# score is 1/dist * 1/start_diff = 1/(dist*start_diff)
# we'll minimize score = (dist*start_diff)
score = float(dist * start_diff)
if score < best_score
best_score = score
best_path = all_train_coords[k]
end
end
return best_path
end
function findClosestTrainingExamplesDTWstarDiffAvg(all_train_coords, test_path, w=2, num_avg=30)
num_paths = length(all_train_coords)
dists = [float(DTWDistance(train_path, test_path, w)) for train_path in all_train_coords]
start_diffs = [float(euclideanDist(test_path[:,1], train_path[:,1])+0.0000001) for train_path in all_train_coords]
num_coords = [size(train_path,2) for train_path in all_train_coords]
df = DataFrame(DISTS = dists, START_DIFF = start_diffs, NUM_COORDS = num_coords, DTW_START_DIFF = dists .* start_diffs)
sort!(df, cols=[:DTW_START_DIFF])
avg_length = median(df[:NUM_COORDS][1:min(end,num_avg)])
return int(avg_length)
end
function findClosestTrainingExampleForTestSet2(train_df, test_df, max_subset=1000, w=1)
all_train_paths = train_df[:COORDS]
all_test_paths = test_df[:COORDS]
num_train_paths = length(all_train_paths)
num_test_paths = length(all_test_paths)
train_paths_subset = all_train_paths[7000:(7000+max_subset)]
closest_examples = cell(num_test_paths)
for k=1:num_test_paths
if k % 20 == 0
println(k, "/", num_test_paths, " for ", num_train_paths, " train path examples")
end
test_path = all_test_paths[k]
test_path_len = size(test_path, 2)
test_taxi_id = test_df[:TAXI_ID][k]
same_taxi_df = train_df[train_df[:TAXI_ID] .== test_taxi_id,:]
if size(same_taxi_df,1) == 0
println("no taxi ids found for ", test_taxi_id)
closest_training_example = findClosestTrainingExample(train_paths_subset, test_path, w)
closest_examples[k] = closest_training_example
else
#println(size(same_taxi_df,1), " - number of taxi id's routes")
#println("test path length: ", test_path_len)
#println("taxi id: ", test_taxi_id)
same_taxi_paths = same_taxi_df[same_taxi_df[:NUM_COORDS] .>= test_path_len,:][:COORDS][1:min(end,max_subset)]
println(size(same_taxi_df,1), " - number of taxi id's routes with ", length(same_taxi_paths),
" greater than length ", test_path_len,"!")
if length(same_taxi_paths) == 0
println("all paths filtered out! resorting to random search again!")
closest_training_example = findClosestTrainingExample(train_paths_subset, test_path, w)
closest_examples[k] = closest_training_example
else
closest_training_example = findClosestTrainingExample(same_taxi_paths, test_path, w)
closest_examples[k] = closest_training_example
end
end
end
return closest_examples
end
function findClosestTrainingExampleForTestSet3(train_df, test_df, max_subset=4000, w=1)
all_train_paths = train_df[:COORDS]
all_test_paths = test_df[:COORDS]
num_train_paths = length(all_train_paths)
num_test_paths = length(all_test_paths)
train_paths_subset = all_train_paths[7000:(7000+max_subset)]
closest_examples = cell(num_test_paths)
for k=1:num_test_paths
if k % 20 == 0
println(k, "/", num_test_paths, " for ", num_train_paths, " train path examples")
end
test_path = all_test_paths[k]
test_path_len = size(test_path, 2)
test_taxi_id = test_df[:TAXI_ID][k]
same_taxi_df = train_df[train_df[:TAXI_ID] .== test_taxi_id,:]
if size(same_taxi_df,1) == 0
println("no taxi ids found for ", test_taxi_id)
closest_training_example = findClosestTrainingExample3(train_paths_subset, test_path, w)
closest_examples[k] = closest_training_example
else
#println(size(same_taxi_df,1), " - number of taxi id's routes")
#println("test path length: ", test_path_len)
#println("taxi id: ", test_taxi_id)
same_taxi_paths = same_taxi_df[same_taxi_df[:NUM_COORDS] .>= test_path_len,:][:COORDS]
println(size(same_taxi_df,1), " - number of taxi id's routes with ", length(same_taxi_paths),
" greater than length ", test_path_len,"!")
if length(same_taxi_paths) == 0
println("all paths filtered out! resorting to random search again!")
closest_training_example = findClosestTrainingExample3(train_paths_subset, test_path, w)
closest_examples[k] = closest_training_example
else
closest_training_example = findClosestTrainingExample3(same_taxi_paths, test_path, w)
closest_examples[k] = closest_training_example
end
end
end
return closest_examples
end
function avgDTWstartDiffScore(train_df, test_df, max_subset=4000, w=2, num_avg=30)
all_train_paths = train_df[:COORDS]
all_test_paths = test_df[:COORDS]
num_train_paths = length(all_train_paths)
num_test_paths = length(all_test_paths)
train_paths_subset = all_train_paths[7000:(7000+max_subset)]
closest_examples = cell(num_test_paths)
for k=1:num_test_paths
if k % 20 == 0
println(k, "/", num_test_paths, " for ", num_train_paths, " train path examples")
end
test_path = all_test_paths[k]
test_path_len = size(test_path, 2)
test_taxi_id = test_df[:TAXI_ID][k]
same_taxi_df = train_df[train_df[:TAXI_ID] .== test_taxi_id,:]
if size(same_taxi_df,1) == 0
println("no taxi ids found for ", test_taxi_id)
closest_training_example = findClosestTrainingExamplesDTWstarDiffAvg(train_paths_subset, test_path, w, num_avg)
closest_examples[k] = closest_training_example
else
#println(size(same_taxi_df,1), " - number of taxi id's routes")
#println("test path length: ", test_path_len)
#println("taxi id: ", test_taxi_id)
same_taxi_paths = same_taxi_df[same_taxi_df[:NUM_COORDS] .>= test_path_len,:][:COORDS]
println(size(same_taxi_df,1), " - number of taxi id's routes with ", length(same_taxi_paths),
" greater than length ", test_path_len,"!")
if length(same_taxi_paths) == 0
println("all paths filtered out! resorting to random search again!")
closest_training_example = findClosestTrainingExamplesDTWstarDiffAvg(train_paths_subset, test_path, w, num_avg)
closest_examples[k] = closest_training_example
else
closest_training_example = findClosestTrainingExamplesDTWstarDiffAvg(same_taxi_paths, test_path, w, num_avg)
closest_examples[k] = closest_training_example
end
end
end
return closest_examples
end
# +
function startEndPrediction(train_paths, test_paths)
num_test_examples = length(test_paths)
end
# +
println("SUBMISSION PREDICTION")
# 1. Using subset of all_train_coords and scanning it for each test example
#all_train_coords = taxi_df[:COORDS][1:20]
#all_validation_coords = taxi_validation_df[:COORDS]
#test_guess_paths = findClosestTrainingExampleForTestSet(all_train_coords, all_validation_coords)
#taxi_validation_df[:GUESS_PATHS] = test_guess_paths
# 2. Scanning paths of same taxi
all_train_coords = taxi_df[:COORDS]
all_validation_coords = taxi_validation_df[:COORDS]
test_guess_lengths = avgDTWstartDiffScore(taxi_df, taxi_validation_df, 4000, 2, 30)
taxi_validation_df[:GUESS_NUM_COORDS] = test_guess_lengths
# +
println("Local prediction")
#scanning subset of train paths for all test paths
#all_train_coords = taxi_df[:COORDS][10000:10400]
#test_df = tail(taxi_df, 100)
#all_test_coords = test_df[:COORDS_TEST]
#test_guess_paths = findClosestTrainingExampleForTestSet(all_train_coords, all_test_coords, 2)
#test_df[:GUESS_PATHS] = test_guess_paths
#looking at paths for same driver
train_df = taxi_df[1:90000,:]
test_df = tail(taxi_df, 10)
test_coords = test_df[:COORDS]
test_df[:COORDS] = test_df[:COORDS_TEST]
test_guess_lengths = avgDTWstartDiffScore(train_df, test_df, 1000, 2, 140)
test_df[:GUESS_NUM_COORDS] = test_guess_lengths
test_df[:COORDS] = test_coords #change it back for scoring
score_coords_guess(test_df)
# +
function score_path_guess(test_df)
pred_paths = test_df[:GUESS_PATHS]
actual_paths = test_df[:COORDS]
pred_times = [((size(x,2)-1)*15)::Int64 for x in pred_paths]
actual_times = [((size(x,2)-1)*15)::Int64 for x in actual_paths]
score = sqrt(mean((log(max(pred_times, 660)+1)-log(actual_times+1)).^2))
println("time score: ", score)
for k=1:length(pred_times)
println("pred: ", pred_times[k], ", actual: ", actual_times[k], ", delta error: ", pred_times[k]-actual_times[k])
end
end
function score_coords_guess(test_df)
pred_num_coords = test_df[:GUESS_NUM_COORDS]
actual_paths = test_df[:COORDS]
pred_times = [((x-1)*15)::Int64 for x in pred_num_coords]
actual_times = [((size(x,2)-1)*15)::Int64 for x in actual_paths]
score = sqrt(mean((log(max(pred_times, 660)+1)-log(actual_times+1)).^2))
println("time score: ", score)
for k=1:length(pred_times)
println("pred: ", pred_times[k], ", actual: ", actual_times[k], ", delta error: ", pred_times[k]-actual_times[k])
end
end
score_path_guess(test_df)
# -
log([1 2 3])
# +
#taxi_validation_df[:GUESS_PATHS] = test_guess_paths
#guess_times = [length(x)*15 for x in test_guess_paths]
num_test_examples = length(test_guess_paths)
guess_times = Array(Int64, num_test_examples)
dest_coords = cell(num_test_examples)
all_test_paths = taxi_validation_df[:COORDS]
test_guess_paths = taxi_validation_df[:GUESS_PATHS]
for k=1:num_test_examples
test_path = all_test_paths[k]
best_guess_path = test_guess_paths[k]
test_path_time = length(test_path)*15
best_guess_time = length(best_guess_path)*15
if test_path_time > best_guess_time
println(k, ": guessing ", best_guess_time, " but existing time is ", test_path_time)
#best_guess_time = test_path_time + 100
end
guess_times[k] = max(test_path_time, best_guess_time)
end
submission_validation = guess_times
# -
size(taxi_df[:COORDS][1],2)
# #Generating Submission
# +
# beat the benchmark example
#mean_time = mean(times_validation)
#submission_validation = [max(x, mean_time) for x in times_validation]
#submission_validation
df_submission = DataFrame()
df_submission[:TRIP_ID] = taxi_validation_df[:TRIP_ID]
df_submission[:TRAVEL_TIME] = submission_validation
writetable("7th_submission_DTW_START_DIFF.csv", df_submission)
# +
lats = [float(x[2,end])::Float64 for x in taxi_validation_df[:GUESS_PATHS]]
lons = [float(x[1,end])::Float64 for x in taxi_validation_df[:GUESS_PATHS]]
df_submission = DataFrame()
df_submission[:TRIP_ID] = taxi_validation_df[:TRIP_ID]
df_submission[:LATITUDE] = lats
df_submission[:LONGITUDE] = lons
writetable("1th_sub_endpoint_DTW_START_DIFF.csv", df_submission)
# +
immutable Point2{T}
x::T
y::T
end
D = [Point2(1.,2.) => 42]
haskey(D, Point2(1., 2.)) #False!
# -
taxi_validation_df[:COORDS]
| julia_taxi2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import sqlite3 as sql
import pandas as pd
import numpy as np
import matplotlib.pyplot as plot
conn = sql.connect('trvlag.db')
query = '''SELECT * FROM booking'''
df = pd.read_sql_query(query, conn)
df
# +
listCountry0 = df['COUNTRY_NAME'] == df['COUNTRY_NAME'][0]
listCountry1 = df['COUNTRY_NAME'] == df['COUNTRY_NAME'][1]
listCountry2 = df['COUNTRY_NAME'] == df['COUNTRY_NAME'][4]
data = {str(df['COUNTRY_NAME'][0]): len(df[listCountry0]),
str(df['COUNTRY_NAME'][1]): len(df[listCountry1]),
str(df['COUNTRY_NAME'][4]): len(df[listCountry2]) }
courses = list(data.keys())
values = list(data.values())
fig = plot.figure(figsize = (8, 4))
plot.bar(courses, values, color ='blue', width = 0.3)
plot.xlabel("Country")
plot.ylabel("Count")
plot.title("No of entries in the country")
plot.show()
# -
| trvl_agency/trvlag_report.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # GeoEnrichment
# + [markdown] slideshow={"slide_type": "-"}
# GeoEnrichment provides the ability to
# * get facts about a location or area.
# * information about the people, places, and businesses
# * in a specific area or
# * within a certain distance or drive time from a location.
# * large collection of data sets including population, income, housing, consumer behavior, and the natural environment.
# * Site analysis is a popular application
# + [markdown] slideshow={"slide_type": "-"}
# ### Login
# + slideshow={"slide_type": "skip"}
from arcgis.gis import GIS
from arcgis.geoenrichment import *
gis = GIS(profile='agol_profile', verify_cert=False)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## GeoEnrichment coverage
# + slideshow={"slide_type": "fragment"}
countries = get_countries()
print("Number of countries for which GeoEnrichment data is available: " + str(len(countries)))
#print a few countries for a sample
countries[0:10]
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Filtering countries by properties
# + slideshow={"slide_type": "fragment"}
[c.properties.name for c in countries if c.properties.continent == 'Oceania']
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Discovering information for a country
# * Data collections,
# * Sub-geographies and
# * Available reports for a country
# + slideshow={"slide_type": "fragment"}
aus = Country.get('Australia')
# + [markdown] slideshow={"slide_type": "skip"}
# Commonly used properties for the country are accessible using `Country.properties`.
# + slideshow={"slide_type": "fragment"}
aus.properties.name
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Data collections and analysis variables
# + slideshow={"slide_type": "fragment"}
df = aus.data_collections
df.head()
# + slideshow={"slide_type": "-"}
# call the shape property to get the total number of rows and columns
df.shape
# + [markdown] slideshow={"slide_type": "fragment"}
# Query the `EducationalAttainment` data collection and get all the unique `analysisVariable`s under that collection
# + slideshow={"slide_type": "-"}
df.loc['EducationalAttainment']['analysisVariable'].unique()
# + slideshow={"slide_type": "fragment"}
# view a sample of the `Age` data collection
df.loc['EducationalAttainment'].head()
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Enriching an address
# + slideshow={"slide_type": "-"}
sdf = enrich(study_areas=["Parliament Dr, Canberra ACT 2600, Australia"],
data_collections=['EducationalAttainment'])
# -
sdf.spatial.plot()
# + [markdown] slideshow={"slide_type": "subslide"}
# # Reports
# + slideshow={"slide_type": "fragment"}
aus.reports.head(6)
# + slideshow={"slide_type": "-"}
# total number of reports available
aus.reports.shape
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Creating Reports
# + slideshow={"slide_type": "-"}
import tempfile
report = create_report(study_areas=["Parliament Dr, Canberra ACT 2600, Australia"],
report="AustraliaFoodAndBeverageSpendingMDS",
export_format="PDF",
out_folder=tempfile.gettempdir(), out_name="FoodAndBeverageSpending.pdf")
print(report)
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Finding named statistical areas
#
# Each country has several named statistical areas in a hierarchy of geography levels (such as states, counties, zip codes, etc).
# + slideshow={"slide_type": "skip"}
# %config IPCompleter.greedy=True
# + slideshow={"slide_type": "fragment"}
de = Country.get("Germany")
de.subgeographies.states['Hamburg']
# + slideshow={"slide_type": "fragment"}
de.subgeographies.states["Hamburg"].districts['Hamburg,_Freie_und_Hansestadt']
# -
de.subgeographies.postcodes2['Berlin']
# + [markdown] slideshow={"slide_type": "skip"}
# The named areas can also be drawn on a map, as they include a `geometry` property.
# + slideshow={"slide_type": "subslide"}
m1 = gis.map('Hamburg, Germany', zoomlevel=9)
m1
# + slideshow={"slide_type": "-"}
m1.draw(de.subgeographies.states["Hamburg"].districts['Hamburg,_Freie_und_Hansestadt'].geometry)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Different geography levels for different country
# + slideshow={"slide_type": "-"}
india = Country.get('India')
# + slideshow={"slide_type": "fragment"}
india.subgeographies.states['Uttar_Pradesh'].districts['Baghpat'].subdistricts['Baraut']
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Searching for named areas within a country
# + slideshow={"slide_type": "fragment"}
riversides_in_usa = usa.search('Riverside')
print("number of riversides in the US: " + str(len(riversides_in_usa)))
# list a few of them
riversides_in_usa[:10]
# -
# For instance, you can make a map of all the riversides in the US
# + slideshow={"slide_type": "subslide"}
usamap = gis.map('United States', zoomlevel=4)
usamap
# + slideshow={"slide_type": "-"}
for riverside in riversides_in_usa:
usamap.draw(riverside.geometry)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Filtering named areas by geography level
# + slideshow={"slide_type": "fragment"}
[level['id'] for level in usa.levels]
# + slideshow={"slide_type": "fragment"}
usa.search(query='Riverside', layers=['US.Counties'])
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Study Areas
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Accepted forms of study areas
#
# - **Street address locations** - Locations can be passed as strings of input street addresses, points of interest or place names.
# + **Example:** `"380 New York St, Redlands, CA"`
#
# - **Multiple field input addresses** - Locations described as multiple field input addresses, using dictionaries.
# + **Example:**
# {"Address" : "380 New York Street",
# "City" : "Redlands",
# "Region" : "CA",
# "Postal" : 92373}
#
# - **Point and line geometries** - Point and line locations, using `arcgis.geometry` instances.
# + **Example Point Location: **
#
# `arcgis.geometry.Geometry({"x":-122.435,"y":37.785})`
#
# + ** Example Point location obtained using find_businesses() above: **
#
# `arcgis.geometry.Geometry(businesses.iloc[0]['SHAPE'])`
#
# - **Buffered study areas** - `BufferStudyArea` instances to change the ring buffer size or create drive-time service areas around points specified using one of the above methods. BufferStudyArea allows you to buffer point and street address study areas. They can be created using the following parameters:
# * area: the point geometry or street address (string) study area to be buffered
# * radii: list of distances by which to buffer the study area, eg. [1, 2, 3]
# * units: distance unit, eg. Miles, Kilometers, Minutes (when using drive times/travel_mode)
# * overlap: boolean, uses overlapping rings/network service areas when True, or non-overlapping disks when False
# * travel_mode: None or string, one of the supported travel modes when using network service areas
# + **Example Buffered Location: **
#
# `pt = arcgis.geometry.Geometry({"x":-122.435,"y":37.785})
# buffered_area = BufferStudyArea(area=pt, radii=[1,2,3], units="Miles", overlap=False)`
#
# - **Network service areas** - `BufferStudyArea` also allows you to define drive time service areas around points as well as other advanced service areas such as walking and trucking.
# + **Example: **
#
# `pt = arcgis.geometry.Geometry({"x":-122.435,"y":37.785})
# buffered_area = BufferStudyArea(area=pt, radii=[1,2,3], units="Minutes", travel_mode="Driving")`
#
# - **Named statistical areas** -
# + **Example:**
#
# `usa.subgeographies.states['California'].zip5['92373']`
#
# - **Polygon geometries** - Locations can given as polygon geometries.
# + **Example Polygon geometry: **
#
# `arcgis.geometry.Geometry({"rings":[[[-117.185412,34.063170],[-122.81,37.81],[-117.200570,34.057196],[-117.185412,34.063170]]],"spatialReference":{"wkid":4326}})`
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: Enriching a named statistical area
# Enriching zip code 92373 in California using the 'Age' data collection:
# + slideshow={"slide_type": "-"}
redlands = usa.subgeographies.states['California'].zip5['92373']
# + slideshow={"slide_type": "fragment"}
enrich(study_areas=[redlands], data_collections=['Age'] )
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: Enrich all counties in a state
# + slideshow={"slide_type": "-"}
ca_counties = usa.subgeographies.states['California'].counties
# + slideshow={"slide_type": "fragment"}
counties_df = enrich(study_areas=ca_counties, data_collections=['Age'])
counties_df.head(10)
# + slideshow={"slide_type": "subslide"}
m2 = gis.map('California')
m2
# + slideshow={"slide_type": "-"}
item = gis.content.import_data(df=counties_df, title="CA county population")
# + slideshow={"slide_type": "-"}
item
# + slideshow={"slide_type": "-"}
m2.add_layer(item.layers[0], {'renderer': 'ClassedColorRenderer',
'field_name':'FEM0'})
# + slideshow={"slide_type": "-"}
item.delete()
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: Using comparison levels
# + slideshow={"slide_type": "-"}
enrich(study_areas=[redlands], data_collections=['Age'],
comparison_levels=['US.Counties', 'US.States'])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: Buffering locations using non overlapping disks
#
# The example below creates non-overlapping disks of radii 1, 3 and 5 Miles respectively from a street address and enriches these using the 'Age' data collection.
# + slideshow={"slide_type": "-"}
buffered = BufferStudyArea(area='380 New York St Redlands CA 92373',
radii=[1,3,5], units='Miles', overlap=False)
enrich(study_areas=[buffered], data_collections=['Age'])
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example: Using drive times as study areas
#
# The example below creates 5 and 10 minute drive times from a street address and enriches these using the 'Age' data collection.
# + slideshow={"slide_type": "-"}
buffered = BufferStudyArea(area='380 New York St Redlands CA 92373',
radii=[5, 10], units='Minutes',
travel_mode='Driving')
drive_time_df = enrich(study_areas=[buffered], data_collections=['Age'])
# + slideshow={"slide_type": "-"}
drive_time_df
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Visualize results on a map
# + [markdown] slideshow={"slide_type": "skip"}
# The returned spatial dataframe can be visualized on a map as shown below:
# + slideshow={"slide_type": "-"}
redlands_map = gis.map('Redlands, CA')
redlands_map.basemap = 'dark-gray-vector'
redlands_map
# + slideshow={"slide_type": "-"}
drive_time_df.spatial.plot(redlands_map,
renderer_type='c', # for class breaks renderer
method='esriClassifyNaturalBreaks', # classification algorithm
class_count=3, # choose the number of classes
col='bufferRadii', # numeric column to classify
cmap='prism', # color map to pick colors from for each class
alpha=0.7) # specify opacity
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Saving GeoEnrichment Results
# + slideshow={"slide_type": "-"} active=""
# gis.content.import_data(df=drive_time_df, title="Age statistics within 5,10 minutes of drive time from Esri")
| 04. Performing Analysis with the Python API/04 - geoenrichment/05 - Geoenrichment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Singko25/Linear-Algebra-58020/blob/main/SAAVEDRA_VECTOR.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="OII_-vczBEf0"
# Matrix And Its Operations
#
# + colab={"base_uri": "https://localhost:8080/"} id="LXcz2148BK5y" outputId="54a8fd65-4c4f-478b-db33-a7e0e2bf602a"
import numpy as nmp
#Create a 1x3 array
a = nmp.array([1,2,3])
print(a)
# + colab={"base_uri": "https://localhost:8080/"} id="9BXlm8DgDFA1" outputId="58d3806f-3649-4e87-c65b-e74740ed96b5"
b = nmp.array([[1,2,3],[-1,-2,-3]])
print(b)
# + colab={"base_uri": "https://localhost:8080/"} id="hXfnV93jD86b" outputId="39082773-5300-4ec2-c62a-5679fd30ea30"
c = nmp.array([[1,2,3],[-1,-2,-3],[4,5,6]])
print(c)
# + colab={"base_uri": "https://localhost:8080/"} id="tBBwSBAYEhW4" outputId="6f8ed1ad-03e7-4c10-a2fd-99387a92f70f"
d = nmp.array([[1,2,3],[-1,-2,-3],[4,5,6]])
print(d)
e = nmp.diagonal([[1,2,3],[-1,-2,-3],[4,5,6]])
print(e)
#diagonal
# + colab={"base_uri": "https://localhost:8080/"} id="qbnaxR7rFEE4" outputId="e0587f80-150a-49e7-812a-e287505aa684"
f = nmp.eye(3)
print(f)
#identity
# + colab={"base_uri": "https://localhost:8080/"} id="7uKzsN-bFUWJ" outputId="fb5c8aed-d086-4cae-faf1-ed2c10df4020"
g = nmp.zeros((8,3))
print(g)
#zero
# + colab={"base_uri": "https://localhost:8080/"} id="fAAMZtm2FcQx" outputId="a5f4f974-e689-43d3-800b-4297c9558628"
h = nmp.empty((0,5))
print(h)
#empty
# + colab={"base_uri": "https://localhost:8080/"} id="4fBbDIjLFz4F" outputId="d46c2f65-7a6e-40aa-f4d3-fdf5399bb7cd"
#matrix addition
i = nmp.array([[1,2,3],[-1,-2,-3]])
j = nmp.array([[3,4,5],[-3,-4,-5]])
print(j+i)
# + colab={"base_uri": "https://localhost:8080/"} id="rYdJc0SFGYBO" outputId="04cb1fbb-5ec9-4c5d-f23e-ff14874d5aa7"
#substraction
k = nmp.array([[1,2,3],[-1,-2,-3]])
l = nmp.array([[3,4,5],[-3,-4,-5]])
print(k-l)
| SAAVEDRA_VECTOR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 35} executionInfo={"elapsed": 38, "status": "ok", "timestamp": 1638961611402, "user": {"displayName": "Sinatra Lab", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16209430936604316323"}, "user_tz": -60} id="Cj_x8fjYj96X" outputId="25c1aa8e-1963-4b3a-9674-0e1d507baac4"
''' VMP 2022-03-02: parhaps not in final report actually.
Now uses one overall path '''
# + id="DJNs_RZaB3dn"
# overall path to the project
path = "path/to/base"
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 27, "status": "ok", "timestamp": 1638961611404, "user": {"displayName": "Sinatra Lab", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16209430936604316323"}, "user_tz": -60} id="zz1H6BlkkFyG" outputId="92d97c44-bad2-426f-c2d4-cc04cafd9685"
# check RAM
from psutil import virtual_memory
ram_gb = virtual_memory().total / 1e9
print('Your runtime has {:.1f} gigabytes of available RAM\n'.format(ram_gb))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 15374, "status": "ok", "timestamp": 1638961626761, "user": {"displayName": "Sinatra Lab", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16209430936604316323"}, "user_tz": -60} id="56ozGyGgkIt3" outputId="98d4a1a2-f1c4-4e5b-8444-36167c7f89f2"
# basic setup
from google.colab import drive
drive.mount('/content/gdrive')
# + id="4WK_jek3kKzO"
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import math
import re
from tqdm import tqdm
from os import listdir
from os.path import isfile, join
pd.options.mode.chained_assignment = None # default='warn'
# + id="KrqXaiH-JjVW"
# outpath (what changes)
outPath = f"{path}/DATA/collaboration/network/GCC_size/"
# + id="DscH2Mr4Dt9l"
# inpath for the document
inPath = f"{path}/DATA/collaboration/network/metrics/"
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 24, "status": "ok", "timestamp": 1638961626771, "user": {"displayName": "Sinatra Lab", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "16209430936604316323"}, "user_tz": -60} id="c6B-XcxL7NCh" outputId="42b3e4b1-53a5-4511-f94a-61a4d9011c51"
''' check network size (relative) '''
# + id="C9MZWowm7SFd"
# relative network size files
GCC_netsize = sorted([x for x in listdir(f"{inPath}") if x.endswith("netsize.csv")])
# + id="Pd3g-wbAfs2H"
# regex pattern & setup
pattern_date = '\d{4}-\d{2}'
pattern_field = '([a-zA-Z]+)'
# + id="oq7z_2cSeKx_"
# just loop
dfs = []
for filex in GCC_netsize:
d = pd.read_csv(f"{inPath}{filex}")
d['start_date'] = re.findall(pattern_date, filex)[0]
dfs.append(d)
share_df = pd.concat(dfs)
share_df = share_df.assign(
node_fraction = lambda x: x.GCC_nodes / x.G_nodes,
edge_fraction = lambda x: x.GCC_edges / x.G_edges,
edge_w_fraction = lambda x: x.GCC_edges_w / x.G_edges_w,
GCC_edge_div_edgew = lambda x: x.GCC_edges / x.GCC_edges_w
)
share_df.to_csv(f"{outPath}GCC_netsize.csv", index = False)
| Collaboration/network/prepare_data/netsize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/simecek/PseudoDNA_Generator/blob/master/models/3UTR_AWD_LSTM_v0.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="y8NYCcZxvM2P" colab_type="text"
# ## Setup
# + id="NeFnR_pcvEED" colab_type="code" colab={}
# !pip install fastai2>=0.0.11 ipywidgets matplotlib nbdev>=0.2.12 pandas scikit_learn sentencepiece
# + id="ExfZF-bTvSkj" colab_type="code" colab={}
from fastai2.text.all import *
import pandas as pd
import torch
# + id="ERTuSw3U-Yxu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="751b7780-a556-4777-bf7b-a7a52a50f8be"
torch.cuda.is_available(), torch.cuda.device_count(), torch.cuda.get_device_name(0)
# + id="qkgkc8rNQCwY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 128} outputId="51475a65-e4a5-4a63-b9ec-52c2ed344c53"
# Mount to your Google Drive allowing lesson files will be saved to your Drive location
from google.colab import drive
drive.mount('/content/drive')
# + id="uAMnbJuNxBkU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 236} outputId="30c09f99-2135-435e-dfbc-7ea50dcf2b34"
dt = pd.read_csv("/content/drive/My Drive/data/random/random_3utr.csv")
train = dt[(dt.chr!="1") & ~dt.seq.isna()]
test = dt[(dt.chr=="1") & ~dt.seq.isna()]
print(dt.shape, train.shape, test.shape)
dt.head()
# + id="V-WYDE4wvZEA" colab_type="code" colab={}
# !rm -rf split_tok
# !rm -rf split
# !mkdir split
# !mkdir split/train
# !mkdir split/valid
# !mkdir split/train/1/
# !mkdir split/valid/1/
# + id="5KII2R0JvkQT" colab_type="code" colab={}
# splitting the file into training and test part
N = len(dt.seq)
for i, s in enumerate(train.seq):
open("split/train/1/seq"+str(i)+".txt", 'w').writelines([s])
for i, s in enumerate(test.seq):
open("split/valid/1/seq"+str(i)+".txt", 'w').writelines([s])
# + [markdown] id="DsWZp8dnxSqa" colab_type="text"
# ## Tokenizer
# + id="GbRbPRbJv0VB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 217} outputId="352fb1c6-5b3b-4964-cba6-4ff57ce7d272"
# !wget https://raw.githubusercontent.com/simecek/PseudoDNA_Generator/master/models/genomic_tokenizer2.py
# + id="Qm5Dgbicxa_Z" colab_type="code" colab={}
from genomic_tokenizer2 import tkn2
# + [markdown] id="bMNqgj9n1TEw" colab_type="text"
# ## Data Loaders
# + id="zFgyggiNxlz_" colab_type="code" colab={}
dls_lm = TextDataLoaders.from_folder(Path("./split"), bs=128, seed=42,
is_lm=True,
tok_tfm=tkn2, seq_len=50)
# + id="TmftEQ6bx42-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 474} outputId="6bd8319b-2d5f-43c5-8d13-ad0e04d81a43"
dls_lm.show_batch()
# + [markdown] id="7sr_NFaB1uHH" colab_type="text"
# ## Model and Learning
# + id="xcOLVeQk0tJ8" colab_type="code" colab={}
learn = language_model_learner(
dls_lm, AWD_LSTM, drop_mult=0.3, pretrained=False,
metrics=[accuracy, Perplexity()])
# + id="dZOgoDpnPl2E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="89b20b6b-f7a9-43ca-faa6-692abe612db9"
learn.lr_find()
# + id="GdWkfIiG11dd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="cd52e50e-02b3-41e2-cb1a-8ad4f982ad58"
learn.fit_one_cycle(5, 1e-2)
learn.export("/content/drive/My Drive/DNAModels/3UTRs/3UTR_AWD_LSTM_v0.pkl")
| models/3UTR_AWD_LSTM_v0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="054sbUkob2c3" colab_type="text"
# # Google Colab Nasıl Kullanılır?
# + [markdown] id="pmECBCuVdu8N" colab_type="text"
# ## Google Colab Nedir
#
# Google Colab, etkileşimli kodlama için web tabanlı bir iPython Notebook hizmetidir. Google Colab **Nvidia K80 GPU'ya ücretsiz erişim** ve hızlı prototipleme ve görselleştirme için popüler bir araç haline geliyor. Colab'ı öğrenmesi kolaydır ve interaktif kodlama için başka bir popüler araç olan Jupyter Notebook'a çok benzer.
#
# + [markdown] id="ES75IrSxB3Fk" colab_type="text"
# # Colab ile İlgili Temel Bilgiler
# + [markdown] id="TU6draQQrq5M" colab_type="text"
# Bu dosyayı salt okunur olarak görüntülüyorsanız (yani paylaşılan bağlantıyı kullanarak), başlığın hemen altındaki **OPEN IN PLAYGROUND** seçeneğini tıklayın.
#
# Oyun alanı modu, bilgisayara kaydetmeyen geçici bir oturumda salt okunur not defterinin bir kopyasını açar.
#
# Herhangi bir değişikliği kaydetmek istiyorsanız, not defterini kaydetmelisiniz (bu aşağıda açıklanmıştır).
#
# 
# + [markdown] id="48jvUKSArw0k" colab_type="text"
# Bir Colab dosyası, bağımsız olarak çalıştırılabilen kod blokları olan hücrelerden oluşur. Hücreler, sol üst taraftaki **+** düğmelerini tıklayarak eklenebilir.
#
# Üzerine tıklayarak bir hücre seçebilirsiniz. Aşağıda bir kod hücresi örneği verilmiştir.
# + id="WWf-TXIEEDVv" colab_type="code" colab={}
print("Bu bir 'kod' hücresidir.")
a = 4
print("a: %d" % a)
# + [markdown] id="81E69UM6Mgxb" colab_type="text"
# ## Hücreleri Çalıştırma ve Organize Etme
#
# Bir hücreyi veya hücre grubunu çalıştırmak için birkaç seçenek vardır. Bu seçenekler, başlığın **Çalışma Zamanı (Runtime)** menüsünde listelenir.
#
# Alternatif olarak, sol taraftaki 'Run' düğmesiyle tek bir hücre yürütülebilir.
#
# 
#
#
# Yukarıdaki örnek kod hücresini çalıştırmayı deneyin. Ardından, aşağıdaki kod hücresini çalıştırın.
# + id="pNCJOXTum_bm" colab_type="code" colab={}
b = 5 * a
print("a: %d" % b)
print("b: %d" % b)
# + [markdown] id="32do8DDcnhqs" colab_type="text"
# Her bir hücrenin bağımsız olarak çalıştırılabileceğini ve bir değişiklik olması durumunda, yalnızca bu değişiklikle ilgili hücrelerin yeniden çalıştırılması gerektiğini unutmayın.
#
# Bu nedenle, her şeyi tek bir hücreye yazmak yerine kodunuzu hücre bloklarına ayırmak önemlidir. Bu modüler yaklaşım, kodun bazı bölümlerini her şeyi yeniden çalıştırmadan değiştirmeyi kolaylaştırır.
# + [markdown] id="Vwc8IPD7QMNI" colab_type="text"
# ## Colab dosyasını kaydetme ve dışa aktarma
# + [markdown] id="VGJ8HgMhE61D" colab_type="text"
# Yukarıda belirtildiği gibi, Playground yalnızca geçici oturum sağlar, bu nedenle değişiklikleri kaydetmek için not defteri dosyalarının kaydedilmesi gerekir.
#
# Colab not defterini kaydetmenin en kolay yolu, Google Drive'ınıza kaydetmektir. **File > Save a copy in Drive (Dosya> Bir kopyasını Drive'a kaydet)**'i tıklayın. Bu öğreticinin bir kopyasını kaydettikten sonra, Google Drive veya Colab kontrol panelinizden açabilirsiniz.
#
# 
#
# 
#
# Google Drive'ınızda bu kurs için bir klasör (örn. ColabTutorialTR) oluşturmayı deneyin ve dağınıklığı azaltmak için tüm Colab dosyalarını buraya kaydedin.
# + [markdown] id="fRI4jVyo_-zI" colab_type="text"
# Colab, **File > Download .ipynb** veya **File > Download .py** ile bir notebook dosyası veya Python dosyası olarak dışa aktarılabilir.
#
# Dışa aktarılan not defteri dosyaları Google Colab'a yeniden yüklenebilir veya Jupyter Not Defteri'nde kullanılabilir.
# + [markdown] id="DByG4RzaAZtP" colab_type="text"
# Colab dosyasını PDF olarak kaydetmek için **File > Print** ve ardından PDF olarak kaydedin.
#
# Örneğin, Google Chrome'da **Open PDF in Previe (PDF'yi Önizlemede Aç)**'ı tıklayın.
#
# 
# + [markdown] id="hisAbf-8Iz_J" colab_type="text"
# Colab'ı paylaşılabilir bağlantı kullanarak paylaşabilirsiniz. Sağ üst köşedeki **Share (Paylaş)**' düğmesini ve **Get sharable link (Paylaşılabilir bağlantı al)**ı tıklayın.
#
# 
# + [markdown] id="i5Sxrqw7IF-5" colab_type="text"
# # Bash Komutlarını Çalıştırma ve Paketleri Yükleme
#
# ## Bash Komutları
#
# Bash komutları, komutun önüne '' **!** '' eklenerek çalıştırılabilir.
#
# Örneğin, aşağıdaki 'pwd' komutu geçerli çalışma dizinine giden yolu yazdırır.
# + id="vMJkei4GI9kW" colab_type="code" colab={}
# !pwd
# + [markdown] id="UY7u68dsvPjh" colab_type="text"
# 'ls' komutu geçerli dizindeki içerik listesini yazdırır.
# + id="FQL7Bqp_vdDO" colab_type="code" colab={}
# !ls
# + [markdown] id="Y_uVFPv7JjVH" colab_type="text"
# Bash komutu paketleri yüklemek veya dosyaları indirmek için kullanılabilir (örn. Veri kümeleri).
# + [markdown] id="AIV4udfDLwHd" colab_type="text"
# ## Paketleri Yükleme
#
# En yaygın kitaplıklar (PyTorch dahil) Colab'a zaten yüklenmiştir.
#
# Python paketlerini kurmak için şunu kullanın:
# + id="MCAfZtdKLWie" colab_type="code" colab={}
# !pip install [package name]
# + [markdown] id="2dTwYO5dLaBs" colab_type="text"
# Advanced Package Tool : Gelişmiş Paket Aracı (apt) aracılığıyla kullanılabilen diğer paketleri yüklemek için
#
# + id="UTRJlO55LdKf" colab_type="code" colab={}
# !apt-get install [package name]
# + [markdown] id="DNVYbPlnEmZz" colab_type="text"
# # Google Drive'ı kullanma
#
# Yukarıda belirtildiği gibi Google Drive, Colab notebook dosyalarını kaydetmek için kullanılabilir. Ancak bu, Colab oturumunuzun Google Drive'ınıza erişeceği anlamına gelmez. Bazen Colab oturumundan dosya yüklemek veya kaydetmek isteyebilirsiniz ve bunu yapmak için önce Google Drive'ı bağlamanız gerekir.
#
# Google Drive'ı bağlamadan önce, `/content` dizininde `gdrive` dizini bulunmaz;
# + id="QxN9GI2iG0VQ" colab_type="code" colab={}
# !ls /content
# + [markdown] id="Co70w2WwG4_Q" colab_type="text"
# Google Drive'ı bağlamak için aşağıdaki kodu çalıştırın ve yetkilendirme kodunu almak için bağlantıya gidin. Yetkilendirme kodunu, kodu çalıştırarak oluşturulan kutuya yapıştırın ve enter tuşuna basın.
# + id="CXnyoIOFGCYk" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/gdrive')
# + [markdown] id="X7e_EBGeHv5b" colab_type="text"
# Başarıyla bağlandıktan sonra, tüm Google Drive dosyalarınıza `/content/gdrive/My\ Drive/` altından erişilebilir olmalıdır.
# + id="tFyK89OEGkOc" colab_type="code" colab={}
# !ls /content/gdrive/My\ Drive
# + [markdown] id="i3F5Xe8GICzu" colab_type="text"
# Google Drive'ı bağladıktan sonra, veri kümesini Google Drive'a indirip Colab oturumunda kullanabilir veya oturumun çıktılarını Google Drive'ınıza kaydedebilirsiniz.
# + [markdown] id="_F2NZuHRE99n" colab_type="text"
# # Donanım Hızlandırıcısını Kullanma
#
# GPU (veya TPU) kullanmak için**Runtime > Change runtime type** seçeneğine gidin ve **Hardware accelerator** altında **GPU** veya **TPU** seçeneğini belirleyin.
#
# 
#
# GPU'yu etkinleştirdikten sonra, aşağıdaki ifade **print true** olmalıdır.
# + id="ZpeWhDXCGZn_" colab_type="code" colab={}
import torch
torch.cuda.is_available()
| Tutorial/ColabTutorialTR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression Analysis
#
# Regression is a statistical method used in finance, investing, and other disciplines that attempts to determine the strength and character of the relationship between one dependent variable (usually denoted by Y) and a series of other variables (known as independent variables).
#
# Regression helps investment and financial managers to value assets and understand the relationships between variables, such as commodity prices and the stocks of businesses dealing in those commodities.
#
# (Ref: https://www.investopedia.com/terms/r/regression.asp#:~:text=What%20Is%20Regression%3F,(known%20as%20independent%20variables). )
#
# <b>Example:</b>
# - House prices: Larger the house Higher the price
# - Size : Explanatory variable (x)
# - Price : Dependant variable (y)
# - Basically if we know the size of the house, we can expect the price of the house
#
# <b>Regression Types</b>
# 1. Simple Regression : One variable (linear equation)
# $$
# y = \alpha + \beta x
# $$
# - y : dependent
# - x : explanatory (independent)
# - <b>NOTE:</b> Simple regression uses only one variable therefore its estimations will not be accurate, therefore we can present it like :
# $$
# y = \alpha + \beta x + error
# $$
# error is also called 'residuals'
#
#
# 2. Multivariate regression: More than one variable
#
#
#
# +
import numpy as np
import pandas as pd
from scipy import stats
import statsmodels.api as sm
import matplotlib.pyplot as plt
data = pd.read_excel('samplefiles/HousesOnSale.xlsx')
data
# -
data[['House Price', 'House Size']] # get list which will have index, house price, house size columns
# ### Univariate Regression
# explanatory variable
x = data['House Size']
# dependent variable
y = data['House Price']
print(x)
print(y)
plt.scatter(x,y)
plt.show()
plt.scatter(x,y)
plt.axis([0,2500,0,1500000]) # plot axis start from 0,0 to 2500,1500000
plt.title('House Price/Size')
plt.ylabel('Price')
plt.xlabel('Size')
plt.show()
# +
x1 = sm.add_constant(x)
# Ordinary Least Squares : https://financetrain.com/ordinary-least-squares-ols
regression = sm.OLS(y,x1).fit()
regression.summary()
# in the results, R-squared (should be 0.612 based on
# the sample file) means, 61.2% of the results can ve explanaed
# with the explanatory variable we used which is the House Price)
# So, because of 61% is a high number, this is a good model
# -
# #### Measuring how predictable is a regression test:
#
# - Are all regressions created equal ?
# - They are not all equal
# - Some of the explanatory variables are better at predicting other variables
# - for example, for house prices, size is a good pricing indicator. Another example can be location, etc..
#
# - Check this [R-Squared](https://www.investopedia.com/terms/r/r-squared.asp)
# - R-Square varies between 0% and 100%. The higher it is, the more predictive power the model has
#
#
# ## Calculating Alpha, Beta and R-Squared
#
# regression.summary() method used above does provide alpha, beta, and R-squared, alternatively we can use the below approach
#
slope, intercept, r_value, p_value, std_err = stats.linregress(x,y)
# +
print('Slope')
print(slope)
print('Intercept')
print(intercept)
print('r_value')
print(r_value)
print('r_squared')
print(r_value ** 2)
print('p_value')
print(p_value)
print('std_err')
print(std_err)
# -
inte
| pyt-in-finance/03_RegressionAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# TODO: filled in dataset path, e.g. '/demo-mount/datasets/cyclist'
cyclist_dataset_path = None
# cyclist_dataset_path = '/demo-mount/datasets/cyclist'
# TODO: filled in output voc dataset path, e.g. '/demo-mount/datasets/cyclist-voc'
output_dataset_path = None
# output_dataset_path = '/demo-mount/datasets/cyclist-voc'
# TODO: filled cs231n project directory, e.g. '/home/lizhe/cs231n_project'
cs231n_project_path = None
# cs231n_project_path = '/home/lizhe/cs231n_project'
# TODO: filled in the output kitti format eval data, e.g. '/demo-mount/datasets/cyclist-kitti'
# eval_kitti library need a kitti format data as input, so we need to convert the validation and test data
# from tsinghua dataset to kitti format.
output_kitti_format_eval_data_path = None
# output_kitti_format_eval_data_path = '/demo-mount/datasets/cyclist-kitti'
assert cyclist_dataset_path is not None
assert output_dataset_path is not None
assert cs231n_project_path is not None
assert output_kitti_format_eval_data_path is not None
training_subdir = 'leftImg8bit/train/tsinghuaDaimlerDataset/'
validation_subdir = 'leftImg8bit/valid/tsinghuaDaimlerDataset/'
test_subdir = 'leftImg8bit/test/tsinghuaDaimlerDataset/'
# + magic_args="-s \"$cyclist_dataset_path\" \"$training_subdir\" \"$validation_subdir\" \"$test_subdir\"" language="bash"
#
# cd $1
# ls -1 $2 | cut -d. -f1 > train.txt
# ls -1 $3 | cut -d. -f1 > valid.txt
# ls -1 $4 | cut -d. -f1 > test.txt
#
# # head -500 full_train.txt > train.txt
# # head -500 full_valid.txt > valid.txt
# # head -500 full_test.txt > test.txt
# + magic_args="-s \"$cyclist_dataset_path\" \"$output_dataset_path\" \"$cs231n_project_path\"" language="bash"
#
# # This takes around 30 minutes to finish
# cd $3/vod-converter
# python vod_converter/main.py --from tsinghua-daimler --from-path $1 --to voc --to-path $2
# + magic_args="-s \"$cyclist_dataset_path\" \"$output_dataset_path\"" language="bash"
# # Generate train.txt, valid.txt and test.txt
#
# cp $1/train.txt $2/VOC2012/ImageSets/Main/raw_train.txt
# cp $1/valid.txt $2/VOC2012/ImageSets/Main/raw_valid.txt
# cp $1/test.txt $2/VOC2012/ImageSets/Main/raw_test.txt
#
# cd $2/VOC2012/ImageSets/Main/
#
# sed 's/_leftImg8bit//g' raw_train.txt > train.txt
# sed 's/_leftImg8bit//g' raw_valid.txt > valid.txt
# sed 's/_leftImg8bit//g' raw_test.txt > test.txt
#
# rm raw_train.txt
# rm raw_valid.txt
# rm raw_test.txt
#
# head -100 train.txt > small_train.txt
# head -10 valid.txt > small_valid.txt
# head -10 test.txt > small_test.txt
# + magic_args="-s \"$cs231n_project_path\" \"$output_dataset_path\"" language="bash"
# # Create linkage for detectron2 to load the data
# # Supposely the error can be ignored, you can check whether the linkage is correctly created in the printed directory.
#
# mkdir -p $1/detectron2-ResNeSt/datasets/tsinghua_cyclist
# ln -s $2/VOC2012 $1/detectron2-ResNeSt/datasets/tsinghua_cyclist/VOC2012
# echo $1/detectron2-ResNeSt/datasets/tsinghua_cyclist/VOC2012
# + magic_args="-s \"$cyclist_dataset_path\" \"$output_kitti_format_eval_data_path\" \"$cs231n_project_path\" \"$training_subdir\" \"$validation_subdir\" \"$test_subdir\"" language="bash"
#
# cd $1
# ls -1 $4 | cut -d. -f1 > full_train.txt
# ls -1 $5 | cut -d. -f1 > full_valid.txt
# ls -1 $6 | cut -d. -f1 > full_test.txt
#
# # No need to convert training
# head -0 full_train.txt > train.txt
# # Convert all fo the validation data and test data
# cp full_valid.txt valid.txt
# cp full_test.txt test.txt
#
# # head -0 full_train.txt > train.txt
# # head -10 full_valid.txt > valid.txt
# # head -10 full_test.txt > test.txt
#
# cd $3/vod-converter
# mkdir -p /demo-mount/datasets/cyclist-kitti
# python vod_converter/main.py --from tsinghua-daimler --from-path $1 --to kitti --to-path $2
#
# cd $1
# cp full_train.txt train.txt
# cp full_valid.txt valid.txt
# cp full_test.txt test.txt
# + magic_args="-s \"$output_kitti_format_eval_data_path\" \"$cs231n_project_path\"" language="bash"
# mkdir $2/eval_kitti/build/data/object -p
# ln -s $1/training/label_2 $2/eval_kitti/build/data/object/label_2
# + magic_args="-s \"$cs231n_project_path\"" language="bash"
#
# # Rebuild eval_kitti as there's code change
# cd $1/eval_kitti/build
# cmake ..
# make
# -
| convert_cyclist_data_to_voc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction to Binary Search and Complexity Analysis with Python
#
# ### Part 1 of "Data Structures and Algorithms in Python"
#
# [Data Structures and Algorithms in Python](https://pythondsa.com) is beginner-friendly introduction to common data structures (linked lists, stacks, queues, graphs) and algorithms (search, sorting, recursion, dynamic programming) in Python, designed to help you prepare for coding interviews and assessments. Check out the full series here:
#
# 1. [Binary Search and Complexity Analysis](https://jovian.ai/aakashns/python-binary-search)
# 2. [Python Classes and Linked Lists](https://jovian.ai/aakashns/python-classes-and-linked-lists)
# 3. Arrays, Stacks, Queues and Strings (coming soon)
# 4. Binary Search Trees and Hash Tables (coming soon)
# 5. Insertion Sort, Merge Sort and Divide-and-Conquer (coming soon)
# 6. Quicksort, Partitions and Average-case Complexity (coming soon)
# 7. Recursion, Backtracking and Dynamic Programming (coming soon)
# 8. Knapsack, Subsequence and Matrix Problems (coming soon)
# 9. Graphs, Breadth-First Search and Depth-First Search (coming soon)
# 10. Shortest Paths, Spanning Trees & Topological Sorting (coming soon)
# 11. Disjoint Sets and the Union Find Algorithm (coming soon)
# 12. Interview Questions, Tips & Practical Advice (coming soon)
#
#
# Earn a verified certificate of accomplishment for this course by signing up here: http://pythondsa.com .
#
# Ask questions, get help & participate in discussions on the community forum: https://jovian.ai/forum/c/data-structures-and-algorithms-in-python/78
# ### Prerequisites
#
# This course assumes very little background in programming and mathematics, and you can learn the required concepts here:
#
# - Basic programming with Python ([variables](https://jovian.ai/aakashns/first-steps-with-python), [data types](https://jovian.ai/aakashns/python-variables-and-data-types), [loops](https://jovian.ai/aakashns/python-branching-and-loops), [functions](https://jovian.ai/aakashns/python-functions-and-scope) etc.)
# - Some high school mathematics ([polynomials](https://www.youtube.com/watch?v=Vm7H0VTlIco), [vectors, matrices](https://www.youtube.com/watch?v=0oGJTQCy4cQ&list=PLSQl0a2vh4HCs4zPpOEdF2GuydqS90Yb6) and [probability](https://www.youtube.com/watch?v=uzkc-qNVoOk))
# - No prior knowledge of data structures or algorithms is required
#
# We'll cover any additional mathematical and theoretical concepts we need as we go along.
#
#
# ## How to Run the Code
#
# The best way to learn the material is to execute the code and experiment with it yourself. This tutorial is an executable [Jupyter notebook](https://jupyter.org). You can _run_ this tutorial and experiment with the code examples in a couple of ways: *using free online resources* (recommended) or *on your computer*.
#
# #### Option 1: Running using free online resources (1-click, recommended)
#
# The easiest way to start executing the code is to click the **Run** button at the top of this page and select **Run on Binder**. You can also select "Run on Colab" or "Run on Kaggle", but you'll need to create an account on [Google Colab](https://colab.research.google.com) or [Kaggle](https://kaggle.com) to use these platforms.
#
#
# #### Option 2: Running on your computer locally
#
# To run the code on your computer locally, you'll need to set up [Python](https://www.python.org), download the notebook and install the required libraries. We recommend using the [Conda](https://docs.conda.io/projects/conda/en/latest/user-guide/install/) distribution of Python. Click the **Run** button at the top of this page, select the **Run Locally** option, and follow the instructions.
#
# > **Jupyter Notebooks**: This notebook is made of _cells_. Each cell can contain code written in Python or explanations in plain English. You can execute code cells and view the results instantly within the notebook. Jupyter is a powerful platform for experimentation and analysis. Don't be afraid to mess around with the code & break things - you'll learn a lot by encountering and fixing errors. You can use the "Kernel > Restart & Clear Output" menu option to clear all outputs and start again from the top.
#
# Try executing the cells below:
# Import a library module
import math
# Use a function from the library
math.sqrt(49)
# ## Problem
#
# This course takes a coding-focused approach towards learning. In each notebook, we'll focus on solving one problem, and learn the techniques, algorithms, and data structures to devise an *efficient* solution. We will then generalize the technique and apply it to other problems.
#
#
#
# In this notebook, we focus on solving the following problem:
#
# > **QUESTION 1:** Alice has some cards with numbers written on them. She arranges the cards in decreasing order, and lays them out face down in a sequence on a table. She challenges Bob to pick out the card containing a given number by turning over as few cards as possible. Write a function to help Bob locate the card.
#
# <img src="https://i.imgur.com/mazym6s.png" width="480">
#
# This may seem like a simple problem, especially if you're familiar with the concept of _binary search_, but the strategy and technique we learning here will be widely applicable, and we'll soon use it to solve harder problems.
# ## Why You Should Learn Data Structures and Algorithms
#
# Whether you're pursuing a career in software development or data science, it's almost certain that you'll be asked to solve programming problems like *reversing a linked list* or *balancing a binary tree* in a technical interview or coding assessment.
#
# It's well known, however, that you will almost never face these problems in your job as a software developer. So it's reasonable to wonder why such problems are asked in interviews and coding assessments. Solving programming problems demonstrates the following traits:
#
# 1. You can **think about a problem systematically** and solve it systematically step-by-step.
# 2. You can **envision different inputs, outputs, and edge cases** for programs you write.
# 3. You can **communicate your ideas clearly** to co-workers and incorporate their suggestions.
# 4. Most importantly, you can **convert your thoughts and ideas into working code** that's also readable.
#
# It's not your knowledge of specific data structures or algorithms that's tested in an interview, but your approach towards the problem. You may fail to solve the problem and still clear the interview or vice versa. In this course, you will learn the skills to both solve problems and clear interviews successfully.
#
# ## The Method
#
# Upon reading the problem, you may get some ideas on how to solve it and your first instinct might be to start writing code. This is not the optimal strategy and you may end up spending a longer time trying to solve the problem due to coding errors, or may not be able to solve it at all.
#
# Here's a systematic strategy we'll apply for solving problems:
#
# 1. State the problem clearly. Identify the input & output formats.
# 2. Come up with some example inputs & outputs. Try to cover all edge cases.
# 3. Come up with a correct solution for the problem. State it in plain English.
# 4. Implement the solution and test it using example inputs. Fix bugs, if any.
# 5. Analyze the algorithm's complexity and identify inefficiencies, if any.
# 6. Apply the right technique to overcome the inefficiency. Repeat steps 3 to 6.
#
# _"Applying the right technique"_ is where the knowledge of common data structures and algorithms comes in handy.
#
# Use this template for solving problems by applying this method: https://jovian.ai/aakashns/python-problem-solving-template
# ## Solution
#
#
# ### 1. State the problem clearly. Identify the input & output formats.
#
# You will often encounter detailed word problems in coding challenges and interviews. The first step is to state the problem clearly and precisely in abstract terms.
#
# <img src="https://i.imgur.com/mazym6s.png" width="480">
#
# In this case, for instance, we can represent the sequence of cards as a list of numbers. Turning over a specific card is equivalent to accessing the value of the number at the corresponding position the list.
#
# <img src="https://i.imgur.com/G9fBarb.png" width="600">
#
# The problem can now be stated as follows:
#
# #### Problem
#
# > We need to write a program to find the position of a given number in a list of numbers arranged in decreasing order. We also need to minimize the number of times we access elements from the list.
#
# #### Input
#
# 1. `cards`: A list of numbers sorted in decreasing order. E.g. `[13, 11, 10, 7, 4, 3, 1, 0]`
# 2. `query`: A number, whose position in the array is to be determined. E.g. `7`
#
# #### Output
#
# 3. `position`: The position of `query` in the list `cards`. E.g. `3` in the above case (counting from `0`)
#
#
#
# Based on the above, we can now create the signature of our function:
def locate_card(cards, query):
pass
# **Tips**:
#
# * Name your function appropriately and think carefully about the signature
# * Discuss the problem with the interviewer if you are unsure how to frame it in abstract terms
# * Use descriptive variable names, otherwise you may forget what a variable represents
#
#
# ### 2. Come up with some example inputs & outputs. Try to cover all edge cases.
#
# Before we start implementing our function, it would be useful to come up with some example inputs and outputs which we can use later to test out problem. We'll refer to them as *test cases*.
#
# Here's the test case described in the example above.
cards = [13, 11, 10, 7, 4, 3, 1, 0]
query = 7
output = 3
# We can test our function by passing the inputs into function and comparing the result with the expected output.
result = locate_card(cards, query)
print(result)
result == output
# Obviously, the two result does not match the output as we have not yet implemented the function.
#
# We'll represent our test cases as dictionaries to make it easier to test them once we write implement our function. For example, the above test case can be represented as follows:
test = {
'input': {
'cards': [13, 11, 10, 7, 4, 3, 1, 0],
'query': 7
},
'output': 3
}
# The function can now be tested as follows.
locate_card(**test['input']) == test['output']
# Our function should be able to handle any set of valid inputs we pass into it. Here's a list of some possible variations we might encounter:
#
# 1. The number `query` occurs somewhere in the middle of the list `cards`.
# 2. `query` is the first element in `cards`.
# 3. `query` is the last element in `cards`.
# 4. The list `cards` contains just one element, which is `query`.
# 5. The list `cards` does not contain number `query`.
# 6. The list `cards` is empty.
# 7. The list `cards` contains repeating numbers.
# 8. The number `query` occurs at more than one position in `cards`.
# 9. (can you think of any more variations?)
#
# > **Edge Cases**: It's likely that you didn't think of all of the above cases when you read the problem for the first time. Some of these (like the empty array or `query` not occurring in `cards`) are called *edge cases*, as they represent rare or extreme examples.
#
# While edge cases may not occur frequently, your programs should be able to handle all edge cases, otherwise they may fail in unexpected ways. Let's create some more test cases for the variations listed above. We'll store all our test cases in an list for easier testing.
tests = []
# +
# query occurs in the middle
tests.append(test)
tests.append({
'input': {
'cards': [13, 11, 10, 7, 4, 3, 1, 0],
'query': 1
},
'output': 6
})
# -
# query is the first element
tests.append({
'input': {
'cards': [4, 2, 1, -1],
'query': 4
},
'output': 0
})
# query is the last element
tests.append({
'input': {
'cards': [3, -1, -9, -127],
'query': -127
},
'output': 3
})
# cards contains just one element, query
tests.append({
'input': {
'cards': [6],
'query': 6
},
'output': 0
})
# The problem statement does not specify what to do if the list `cards` does not contain the number `query`.
#
# 1. Read the problem statement again, carefully.
# 2. Look through the examples provided with the problem.
# 3. Ask the interviewer/platform for a clarification.
# 4. Make a reasonable assumption, state it and move forward.
#
# We will assume that our function will return `-1` in case `cards` does not contain `query`.
# cards does not contain query
tests.append({
'input': {
'cards': [9, 7, 5, 2, -9],
'query': 4
},
'output': -1
})
# cards is empty
tests.append({
'input': {
'cards': [],
'query': 7
},
'output': -1
})
# numbers can repeat in cards
tests.append({
'input': {
'cards': [8, 8, 6, 6, 6, 6, 6, 3, 2, 2, 2, 0, 0, 0],
'query': 3
},
'output': 7
})
# In the case where `query` occurs multiple times in `cards`, we'll expect our function to return the first occurrence of `query`.
#
# While it may also be acceptable for the function to return any position where `query` occurs within the list, it would be slightly more difficult to test the function, as the output is non-deterministic.
# query occurs multiple times
tests.append({
'input': {
'cards': [8, 8, 6, 6, 6, 6, 6, 6, 3, 2, 2, 2, 0, 0, 0],
'query': 6
},
'output': 2
})
# Let's look at the full set of test cases we have created so far.
tests
# Great, now we have a fairly exhaustive set of test cases to evaluate our function.
#
# Creating test cases beforehand allows you to identify different variations and edge cases in advance so that can make sure to handle them while writing code. Sometimes, you may start out confused, but the solution will reveal itself as you try to come up with interesting test cases.
#
#
# **Tip:** Don't stress it if you can't come up with an exhaustive list of test cases though. You can come back to this section and add more test cases as you discover them. Coming up with good test cases is a skill that takes practice.
#
#
# ### 3. Come up with a correct solution for the problem. State it in plain English.
#
# Our first goal should always be to come up with a _correct_ solution to the problem, which may necessarily be the most _efficient_ solution. The simplest or most obvious solution to a problem, which generally involves checking all possible answers is called the _brute force_ solution.
#
# In this problem, coming up with a correct solution is quite easy: Bob can simply turn over cards in order one by one, till he find a card with the given number on it. Here's how we might implement it:
#
# 1. Create a variable `position` with the value 0.
# 3. Check whether the number at index `position` in `card` equals `query`.
# 4. If it does, `position` is the answer and can be returned from the function
# 5. If not, increment the value of `position` by 1, and repeat steps 2 to 5 till we reach the last position.
# 6. If the number was not found, return `-1`.
#
# > **Linear Search Algorithm**: Congratulations, we've just written our first _algorithm_! An algorithm is simply a list of statements which can be converted into code and executed by a computer on different sets of inputs. This particular algorithm is called linear search, since it involves searching through a list in a linear fashion i.e. element after element.
#
#
# **Tip:** Always try to express (speak or write) the algorithm in your own words before you start coding. It can be as brief or detailed as you require it to be. Writing is a great tool for thinking clearly. It's likely that you will find some parts of the solution difficult to express, which suggests that you are probably unable to think about it clearly. The more clearly you are able to express your thoughts, the easier it will be for you to turn into code.
# ### 4. Implement the solution and test it using example inputs. Fix bugs, if any.
#
# Phew! We are finally ready to implement our solution. All the work we've done so far will definitely come in handy, as we now exactly what we want our function to do, and we have an easy way of testing it on a variety of inputs.
#
# Here's a first attempt at implementing the function.
def locate_card(cards, query):
# Create a variable position with the value 0
position = 0
# Set up a loop for repetition
while True:
# Check if element at the current position matche the query
if cards[position] == query:
# Answer found! Return and exit..
return position
# Increment the position
position += 1
# Check if we have reached the end of the array
if position == len(cards):
# Number not found, return -1
return -1
# Let's test out the function with the first test case
test
result = locate_card(test['input']['cards'], test['input']['query'])
result
result == output
# Yay! The result matches the output.
#
# To help you test your functions easily the `jovian` Python library provides a helper function `evalute_test_case`. Apart from checking whether the function produces the expected result, it also displays the input, expected output, actual output from the function, and the execution time of the function.
# !pip install jovian --upgrade --quiet
from jovian.pythondsa import evaluate_test_case
evaluate_test_case(locate_card, test)
# While it may seem like we have a working solution based on the above test, we can't be sure about it until we test the function with all the test cases.
#
# We can use the `evaluate_test_cases` (plural) function from the `jovian` library to test our function on all the test cases with a single line of code.
from jovian.pythondsa import evaluate_test_cases
evaluate_test_cases(locate_card, tests)
# Oh no! Looks like our function encountered an error in the sixth test case. The error message suggests that we're trying to access an index outside the range of valid indices in the list. Looks like the list `cards` is empty in this case, and may be the root of the problem.
#
# Let's add some `print` statements within our function to print the inputs and the value of the `position` variable in each loop.
def locate_card(cards, query):
position = 0
print('cards:', cards)
print('query:', query)
while True:
print('position:', position)
if cards[position] == query:
return position
position += 1
if position == len(cards):
return -1
# +
cards6 = tests[6]['input']['cards']
query6 = tests[6]['input']['query']
locate_card(cards6, query6)
# -
# Clearly, since `cards` is empty, it's not possible to access the element at index 0. To fix this, we can check whether we've reached the end of the array before trying to access an element from it. In fact, this can be terminating condition for the `while` loop itself.
def locate_card(cards, query):
position = 0
while position < len(cards):
if cards[position] == query:
return position
position += 1
return -1
# Let's test the failing case again.
tests[6]
locate_card(cards6, query6)
# The result now matches the expected output. Do you now see the benefit of listing test cases beforehand? Without a good set of test cases, we may never have discovered this error in our function.
#
# Let's verify that all the other test cases pass too.
evaluate_test_cases(locate_card, tests)
# Our code passes all the test cases. Of course, there might be some other edge cases we haven't thought of which may cause the function to fail. Can you think of any?
#
# **Tip**: In a real interview or coding assessment, you can skip the step of implementing and testing the brute force solution in the interest of time. It's generally quite easy to figure out the complexity of the brute for solution from the plain English description.
# ### 5. Analyze the algorithm's complexity and identify inefficiencies, if any.
#
# Recall this statement from original question: _"Alice challenges Bob to pick out the card containing a given number by **turning over as few cards as possible**."_ We restated this requirement as: _"Minimize the number of times we access elements from the list `cards`"_
#
# <img src="https://i.imgur.com/mazym6s.png" width="480">
#
# Before we can minimize the number, we need a way to measure it. Since we access a list element once in every iteration, for a list of size `N` we access the elements from the list up to `N` times. Thus, Bob may need to overturn up to `N` cards in the worst case, to find the required card.
#
# Suppose he is only allowed to overturn 1 card per minute, it may take him 30 minutes to find the required card if 30 cards are laid out on the table. Is this the best he can do? Is a way for Bob to arrive at the answer by turning over just 5 cards, instead of 30?
#
# The field of study concerned with finding the amount of time, space or other resources required to complete the execution of computer programs is called _the analysis of algorithms_. And the process of figuring out the best algorithm to solve a given problem is called _algorithm design and optimization_.
#
#
# ### Complexity and Big O Notation
#
# > **Complexity** of an algorithm is a measure of the amount of time and/or space required by an algorithm for an input of a given size e.g. `N`. Unless otherwise stated, the term _complexity_ always refers to the worst-case complexity (i.e. the highest possible time/space taken by the program/algorithm to process an input).
#
# In the case of linear search:
#
# 1. The _time complexity_ of the algorithm is `cN` for some fixed constant `c` that depends on the number of operations we perform in each iteration and the time taken to execute a statement. Time complexity is sometimes also called the _running time_ of the algorithm.
#
# 2. The _space complexity_ is some constant `c'` (independent of `N`), since we just need a single variable `position` to iterate through the array, and it occupies a constant space in the computer's memory (RAM).
#
#
# > **Big O Notation**: Worst-case complexity is often expressed using the Big O notation. In the Big O, we drop fixed constants and lower powers of variables to capture the trend of relationship between the size of the input and the complexity of the algorithm i.e. if the complexity of the algorithm is `cN^3 + dN^2 + eN + f`, in the Big O notation it is expressed as **O(N^3)**
#
# Thus, the time complexity of linear search is **O(N)** and its space complexity is **O(1)**.
#
#
#
# ### Save and upload your work to Jovian
#
# Whether you're running this Jupyter notebook online or on your computer, it's essential to save your work from time to time. You can continue working on a saved notebook later or share it with friends and colleagues to let them execute your code. [Jovian](https://jovian.ai/platform-features) offers an easy way of saving and sharing your Jupyter notebooks online.
# !pip install jovian --upgrade --quiet
import jovian
jovian.commit(project='python-binary-search', environment=None)
# ### 6. Apply the right technique to overcome the inefficiency. Repeat steps 3 to 6.
#
# At the moment, we're simply going over cards one by one, and not even utilizing the face that they're sorted. This is called a *brute force* approach.
#
# It would be great if Bob could somehow guess the card at the first attempt, but with all the cards turned over it's simply impossible to guess the right card.
#
#
# <img src="https://i.imgur.com/mazym6s.png" width="480">
#
# The next best idea would be to pick a random card, and use the fact that the list is sorted, to determine whether the target card lies to the left or right of it. In fact, if we pick the middle card, we can reduce the number of additional cards to be tested to half the size of the list. Then, we can simply repeat the process with each half. This technique is called binary search. Here's a visual explanation of the technique:
#
#
#
# <img src="https://miro.medium.com/max/494/1*3eOrsoF9idyOp-0Ll9I9PA.png" width="480">
#
#
#
# ### 7. Come up with a correct solution for the problem. State it in plain English.
#
# Here's how binary search can be applied to our problem:
#
# 1. Find the middle element of the list.
# 2. If it matches queried number, return the middle position as the answer.
# 3. If it is less than the queried number, then search the first half of the list
# 3. If it is greater than the queried number, then search the second half of the list
# 4. If no more elements remain, return -1.
#
#
jovian.commit()
# ### 8. Implement the solution and test it using example inputs. Fix bugs, if any.
# Here's an implementation of binary search for solving our problem. We also print the relevant variables in each iteration of the `while` loop.
def locate_card(cards, query):
lo, hi = 0, len(cards) - 1
while lo <= hi:
mid = (lo + hi) // 2
mid_number = cards[mid]
print("lo:", lo, ", hi:", hi, ", mid:", mid, ", mid_number:", mid_number)
if mid_number == query:
return mid
elif mid_number < query:
hi = mid - 1
elif mid_number > query:
lo = mid + 1
return -1
# Let's test it out using the test cases.
evaluate_test_cases(locate_card, tests)
# Looks like it passed 8 out of 9 tests! Let's look at the failed test.
evaluate_test_case(locate_card, tests[8])
# Seems like our function returned the position `7`. Let's check what lies at this position in the input list.
cards8 = tests[8]['input']['cards']
query8 = tests[8]['input']['cards']
query8[7]
# Seems like we did locate a 6 in the array, it's just that it wasn't the first 6. As you can guess, this is because in binary search, we don't go over indices in a linear order.
#
# So how do we fix it?
#
# When we find that `cards[mid]` is equal to `query`, we need to check whether it is the first occurrence of `query` in the list i.e the number that comes before it.
#
# `[8, 8, 6, 6, 6, 6, 6, 6, 3, 2, 2, 2, 0, 0, 0]`
#
# To make it easier, we'll define a helper function called `test_location`, which will take the list `cards`, the `query` and `mid` as inputs.
#
# +
def test_location(cards, query, mid):
mid_number = cards[mid]
print("mid:", mid, ", mid_number:", mid_number)
if mid_number == query:
if mid-1 >= 0 and cards[mid-1] == query:
return 'left'
else:
return 'found'
elif mid_number < query:
return 'left'
else:
return 'right'
def locate_card(cards, query):
lo, hi = 0, len(cards) - 1
while lo <= hi:
print("lo:", lo, ", hi:", hi)
mid = (lo + hi) // 2
result = test_location(cards, query, mid)
if result == 'found':
return mid
elif result == 'left':
hi = mid - 1
elif result == 'right':
lo = mid + 1
return -1
# -
evaluate_test_case(locate_card, tests[8])
evaluate_test_cases(locate_card, tests)
# In fact, once we have written out the algorithm, we may want to add a few more test cases:
#
# 1. The number lies in first half of the array.
# 2. The number lies in the second half of the array.
# Here is the final code for the algorithm (without the `print` statements):
# +
def test_location(cards, query, mid):
if cards[mid] == query:
if mid-1 >= 0 and cards[mid-1] == query:
return 'left'
else:
return 'found'
elif cards[mid] < query:
return 'left'
else:
return 'right'
def locate_card(cards, query):
lo, hi = 0, len(cards) - 1
while lo <= hi:
mid = (lo + hi) // 2
result = test_location(cards, query, mid)
if result == 'found':
return mid
elif result == 'left':
hi = mid - 1
elif result == 'right':
lo = mid + 1
return -1
# -
# Try creating a few more test cases to test the algorithm more extensively.
#
# Let's save our work before continuing.
jovian.commit()
# ### 9. Analyze the algorithm's complexity and identify inefficiencies, if any.
#
# Once again, let's try to count the number of iterations in the algorithm. If we start out with an array of N elements, then each time the size of the array reduces to half for the next iteration, until we are left with just 1 element.
#
# Initial length - `N`
#
# Iteration 1 - `N/2`
#
# Iteration 2 - `N/4` i.e. `N/2^2`
#
# Iteration 3 - `N/8` i.e. `N/2^3`
#
# ...
#
# Iteration k - `N/2^k`
#
#
# Since the final length of the array is 1, we can find the
#
# `N/2^k = 1`
#
# Rearranging the terms, we get
#
# `N = 2^k`
#
# Taking the logarithm
#
# `k = log N`
#
# Where `log` refers to log to the base 2. Therefore, our algorithm has the time complexity **O(log N)**. This fact is often stated as: binary search _runs_ in logarithmic time. You can verify that the space complexity of binary search is **O(1)**.
#
#
#
#
#
# ### Binary Search vs. Linear Search
#
# Su
#
def locate_card_linear(cards, query):
position = 0
while position < len(cards):
if cards[position] == query:
return position
position += 1
return -1
# +
large_test = {
'input': {
'cards': list(range(10000000, 0, -1)),
'query': 2
},
'output': 9999998
}
# +
result, passed, runtime = evaluate_test_case(locate_card_linear, large_test, display=False)
print("Result: {}\nPassed: {}\nExecution Time: {} ms".format(result, passed, runtime))
# +
result, passed, runtime = evaluate_test_case(locate_card, large_test, display=False)
print("Result: {}\nPassed: {}\nExecution Time: {} ms".format(result, passed, runtime))
# -
# The binary search version is over 55,000 times faster than the linear search version.
#
# Furthermore, as the size of the input grows larger, the difference only gets bigger. For a list 10 times, the size, linear search would run for 10 times longer, whereas binary search would only require 3 additional operations! (can you verify this?) That's the real difference between the complexities **O(N)** and **O(log N)**.
#
# Another way to look at it is that binary search runs `c * N / log N` times faster than linear search, for some fixed constant `c`. Since `log N` grows very slowly compared to `N`, the difference gets larger with the size of the input. Here's a graph showing how the comparing common functions for running time of algorithms ([source](https://dev.to/b0nbon1/understanding-big-o-notation-with-javascript-25mc)):
#
# <img src="https://res.cloudinary.com/practicaldev/image/fetch/s--NR3M1nw8--/c_limit%2Cf_auto%2Cfl_progressive%2Cq_auto%2Cw_880/https://thepracticaldev.s3.amazonaws.com/i/z4bbf8o1ly77wmkjdgge.png" width="480">
#
# Do you see now why we ignore constants and lower order terms while expressing the complexity using the Big O notation?
# ## Generic Binary Search
#
# Here is the general strategy behind binary search, which is applicable to a variety of problems:
#
# 1. Come up with a condition to determine whether the answer lies before, after or at a given position
# 1. Retrieve the midpoint and the middle element of the list.
# 2. If it is the answer, return the middle position as the answer.
# 3. If answer lies before it, repeat the search with the first half of the list
# 4. If the answer lies after it, repeat the search with the second half of the list.
#
# Here is the generic algorithm for binary search, implemented in Python:
def binary_search(lo, hi, condition):
"""TODO - add docs"""
while lo <= hi:
mid = (lo + hi) // 2
result = condition(mid)
if result == 'found':
return mid
elif result == 'left':
hi = mid - 1
else:
lo = mid + 1
return -1
# The worst-case complexity or running time of binary search is **O(log N)**, provided the complexity of the condition used to determine whether the answer lies before, after or at a given position is **O(1)**.
#
# Note that `binary_search` accepts a function `condition` as an argument. Python allows passing functions as arguments to other functions, unlike C++ and Java.
#
# We can now rewrite the `locate_card` function more succinctly using the `binary_search` function.
def locate_card(cards, query):
def condition(mid):
if cards[mid] == query:
if mid > 0 and cards[mid-1] == query:
return 'left'
else:
return 'found'
elif cards[mid] < query:
return 'left'
else:
return 'right'
return binary_search(0, len(cards) - 1, condition)
# Note here that we have defined a function within a function, another handy feature in Python. And the inner function can access the variables within the outer function.
evaluate_test_cases(locate_card, tests)
# The `binary_search` function can now be used to solve other problems too. It is a tested piece of logic.
#
#
# > **Question**: Given an array of integers nums sorted in ascending order, find the starting and ending position of a given number.
#
# This differs from the problem in only two significant ways:
#
# 1. The numbers are sorted in increasing order.
# 2. We are looking for both the increasing order and the decreasing order.
#
# Here's the full code for solving the question, obtained by making minor modifications to our previous function:
# +
def first_position(nums, target):
def condition(mid):
if nums[mid] == target:
if mid > 0 and nums[mid-1] == target:
return 'left'
return 'found'
elif nums[mid] < target:
return 'right'
else:
return 'left'
return binary_search(0, len(nums)-1, condition)
def last_position(nums, target):
def condition(mid):
if nums[mid] == target:
if mid < len(nums)-1 and nums[mid+1] == target:
return 'right'
return 'found'
elif nums[mid] < target:
return 'right'
else:
return 'left'
return binary_search(0, len(nums)-1, condition)
def first_and_last_position(nums, target):
return first_position(nums, target), last_position(nums, target)
# -
# We can test our solution by making a submission here: https://leetcode.com/problems/find-first-and-last-position-of-element-in-sorted-array/
# ## The Method - Revisited
#
# Here's a systematic strategy we've applied for solving the problem:
#
# 1. State the problem clearly. Identify the input & output formats.
# 2. Come up with some example inputs & outputs. Try to cover all edge cases.
# 3. Come up with a correct solution for the problem. State it in plain English.
# 4. Implement the solution and test it using example inputs. Fix bugs, if any.
# 5. Analyze the algorithm's complexity and identify inefficiencies, if any.
# 6. Apply the right technique to overcome the inefficiency. Repeat steps 3 to 6.
#
# Use this template for solving problems using this method: https://jovian.ai/aakashns/python-problem-solving-template
#
# This seemingly obvious strategy will help you solve almost any programming problem you will face in an interview or coding assessment.
#
# The objective of this course is to rewire your brain to think using this method, by applying it over and over to different types of problems. This is a course about thinking about problems systematically and turning those thoughts into code.
# ## Problems for Practice
#
#
# Here are some resources to learn more and find problems to practice.
#
# * Assignment on Binary Search: https://jovian.ai/aakashns/python-binary-search-assignment
# * Binary Search Problems on LeetCode: https://leetcode.com/problems/binary-search/
# * Binary Search Problems on GeeksForGeeks: https://www.geeksforgeeks.org/binary-search/
# * Binary Search Problems on Codeforces: https://codeforces.com/problemset?tags=binary+search
#
# Use this template for solving problems: https://jovian.ai/aakashns/python-problem-solving-template
#
# Start a discussion on the forum: https://jovian.ai/forum/c/data-structures-and-algorithms-in-python/lesson-1-binary-search-linked-lists-and-complex/81
#
# Try to solve at least 5-10 problems over the week to master binary search.
jovian.commit()
| Data_Structures_and_Algorithms_in_Python/Lesson_1-Binary_Search,_Linked_Lists_and_Complexity/.ipynb_checkpoints/python-binary-search-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # vizzu
#
# [Vizzu](https://lib.vizzuhq.com/) is a free, open-source Javascript/C++ library utilizing a generic dataviz engine that generates many types of charts and seamlessly animates between them. It can be used to create static charts but more importantly it is designed for building animated data stories and interactive explorers as Vizzu enables showing different perspectives of the data that the viewers can easily follow due to the animation.
#
#
#
# # ipyvizzu
#
# ipyvizzu is a [jupyter notebook](https://jupyter.org/) integration for the vizzu project. ipyvizzu works only in the jupyter notebook environment.
# +
from ipyvizzu import Chart, Data, Config
data = Data()
data.add_series("Foo", ['Alice', 'Bob', 'Ted'])
data.add_series("Bar", [15, 32, 12])
data.add_series("Baz", [5, 2, 2])
chart = Chart()
chart.animate(data)
chart.animate(Config({"x": "Foo", "y": "Bar", "color": "Foo"}))
chart.animate(Config({"geometry": "circle"}))
chart.animate(Config({"x": "Foo", "y": "Baz", "color": "Foo"}))
chart.animate(Config({"geometry": "rectangle"}))
# -
# **Note:** `Chart` only generates a javascript code. The vizzu calls are evaulated by the browser. Therefore if the vizzu figure is blank you should check the console of your browser where the javascript reports its errors.
#
# * [ipyvizzu options](options.html)
#
# The examples bellow are copied from the [vizzu tutorial](https://lib.vizzuhq.com/0.4/). You can read more information from there.
# * [Data](examples/data.html)
# * [Axes, title, tooltip](examples/axes.html)
# * [Geometry](examples/geometry.html)
# * [Channels & legend](examples/channels.html)
# * [Group/stack](examples/group.html)
# * [Sorting](examples/sorting.html)
# * [Align & range](examples/align.html)
# * [Aggregate/drill-down](examples/aggregate.html)
# * [Orientation, split & polar](examples/orientation.html)
# * [Without coordinates & noop channel](examples/without_coordinates.html)
# * [Color palette & fonts](examples/palette_font.html)
# * [Chart layout](examples/layout.html)
# * [Animation options](examples/animation_options.html)
| docs/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import cv2 as cv
import numpy as np
import pandas as pd
import matplotlib.image as mpimg
import argparse
import random as rng
rng.seed(12345)
import matplotlib.pyplot as plt
# %matplotlib inline
man_data = pd.read_csv('MAnzaNaDaTa.csv')
e = 5000
x = 0
y = 0
z = 0
w = 0
out_f = {"Nombre de la imagen":"Train (%d)" %e, "x": x , "y": y , "l": z,"a": w}
sesult = man_data.append(out_f,ignore_index=True)
for i in range (-1,3200):
##Gala_d
try:
img = cv.imread('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Gala_d/Gala-d (%d).png' %(i))
if img is None:
pass
else:
lo= Otsuu(img)
x,y,z,w, boun = untitled(lo)
out_f = {"Nombre de la imagen":"Gala_d (%d)" %i, "x": float(x)/800 , "y": float(y)/600 , "l": float(x+z)/800,"a": float(y+w)/600}
sesult = sesult.append(out_f,ignore_index=True)
cv.imwrite('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Gala_d/Gala_d_mask (%d).png' %(i), lo)
crop_img = lo[y:y+w, x:x+z]
cv.imwrite('/home/skunkllr/Documentos/TAREAS/REcortE_Mz/train/Gala_d/Gala_d_rec (%d).png' %(i), crop_img)
except:
pass
##Gala_s
try:
img = cv.imread('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Gala_s/Gala-s (%d).png' %(i))
if img is None:
pass
else:
lo= Otsuu(img)
x,y,z,w, boun = untitled(lo)
out_f = {"Nombre de la imagen":"Gala_s (%d)" %i, "x": float(x)/800 , "y": float(y)/600 , "l": float(x+z)/800,"a": float(y+w)/600}
sesult = sesult.append(out_f,ignore_index=True)
cv.imwrite('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Gala_s/Gala_s_mask (%d).png' %(i), lo)
crop_img = lo[y:y+w, x:x+z]
cv.imwrite('/home/skunkllr/Documentos/TAREAS/REcortE_Mz/train/Gala_s/Gala_s_rec (%d).png' %(i), crop_img)
except:
pass
##Golden_d
try:
img = cv.imread('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Golden_d/Golden-d (%d).png' %(i))
if img is None:
pass
else:
lo= Otsuu(img)
x,y,z,w, boun = untitled(lo)
out_f = {"Nombre de la imagen":"Golden_d (%d)" %i, "x": float(x)/800 , "y": float(y)/600 , "l": float(x+z)/800,"a": float(y+w)/600}
sesult = sesult.append(out_f,ignore_index=True)
cv.imwrite('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Golden_d/Golden_d_mask (%d).png' %(i), lo)
crop_img = lo[y:y+w, x:x+z]
cv.imwrite('/home/skunkllr/Documentos/TAREAS/REcortE_Mz/train/Golden_d/Golden_d_rec (%d).png' %(i), crop_img)
except:
pass
##Golden_s
try:
img = cv.imread('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Golden_s/Golden-s (%d).png' %(i))
if img is None:
pass
else:
lo= Otsuu(img)
x,y,z,w, boun = untitled(lo)
out_f = {"Nombre de la imagen":"Golden_s (%d)" %i, "x": float(x)/800 , "y": float(y)/600 , "l": float(x+z)/800,"a": float(y+w)/600}
sesult = sesult.append(out_f,ignore_index=True)
cv.imwrite('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Golden_s/Golden_s_mask (%d).png' %(i), lo)
crop_img = lo[y:y+w, x:x+z]
cv.imwrite('/home/skunkllr/Documentos/TAREAS/REcortE_Mz/train/Golden_s/Golden_s_rec (%d).png' %(i), crop_img)
except:
pass
##Granny_d
try:
img = cv.imread('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Granny_d/Granny_d (%d).png' %(i))
if img is None:
pass
else:
lo= Otsuu(img)
x,y,z,w, boun = untitled(lo)
out_f = {"Nombre de la imagen":"Granny_d (%d)" %i, "x": float(x)/800 , "y": float(y)/600 , "l": float(x+z)/800,"a": float(y+w)/600}
sesult = sesult.append(out_f,ignore_index=True)
cv.imwrite('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Granny_d/Granny_d_mask (%d).png' %(i), lo)
crop_img = lo[y:y+w, x:x+z]
cv.imwrite('/home/skunkllr/Documentos/TAREAS/REcortE_Mz/train/Granny_d/Granny_d_rec (%d).png' %(i), crop_img)
except:
pass
##Granny_s
try:
img = cv.imread('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Granny_s/Granny_s (%d).png' %(i))
if img is None:
pass
else:
lo= Otsuu(img)
x,y,z,w, boun = untitled(lo)
out_f = {"Nombre de la imagen":"Granny_s (%d)" %i, "x": float(x)/800 , "y": float(y)/600 , "l": float(x+z)/800,"a": float(y+w)/600}
sesult = sesult.append(out_f,ignore_index=True)
cv.imwrite('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Granny_s/Granny_s_mask (%d).png' %(i), lo)
crop_img = lo[y:y+w, x:x+z]
cv.imwrite('/home/skunkllr/Documentos/TAREAS/REcortE_Mz/train/Granny_s/Granny_s_rec (%d).png' %(i), crop_img)
except:
pass
##Red_d
try:
img = cv.imread('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Red_d/Red_d (%d).png' %(i))
if img is None:
pass
else:
lo= Otsuu(img)
x,y,z,w, boun = untitled(lo)
out_f = {"Nombre de la imagen":"Red_d (%d)" %i, "x": float(x)/800 , "y": float(y)/600 , "l": float(x+z)/800,"a": float(y+w)/600}
sesult = sesult.append(out_f,ignore_index=True)
cv.imwrite('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Red_d/Red_d_mask (%d).png' %(i), lo)
crop_img = lo[y:y+w, x:x+z]
cv.imwrite('/home/skunkllr/Documentos/TAREAS/REcortE_Mz/train/Red_d/Red_d_rec (%d).png' %(i), crop_img)
except:
pass
##Red_s
try:
img = cv.imread('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Red_s/Red_s (%d).png' %(i))
if img is None:
pass
else:
lo= Otsuu(img)
x,y,z,w, boun = untitled(lo)
out_f = {"Nombre de la imagen":"Red_s (%d)" %i, "x": float(x)/800 , "y": float(y)/600 , "l": float(x+z)/800,"a": float(y+w)/600}
sesult = sesult.append(out_f,ignore_index=True)
cv.imwrite('/home/skunkllr/Documentos/TAREAS/IMAGENES_MANZANAS_SD/train/Red_s/Red_s_mask (%d).png' %(i), lo)
crop_img = lo[y:y+w, x:x+z]
cv.imwrite('/home/skunkllr/Documentos/TAREAS/REcortE_Mz/train/Red_s/Red_s_rec (%d).png' %(i), crop_img)
except:
pass
sesult.to_csv('MAnzaNaDaTa.csv')
# -
def Otsuu(image):
b,g,r = cv.split(image)
img_grises = b
kernel = np.ones((5,5), np.uint8)
blur = cv.GaussianBlur(b,(5,5),0)
ret3, inverse_mask = cv.threshold(blur,0,255,cv.THRESH_BINARY+cv.THRESH_OTSU)
mask = cv.bitwise_not(inverse_mask)
kernel = np.ones((7,7),np.uint8)
mask_morph = cv.morphologyEx(mask, cv.MORPH_OPEN, kernel)
retval, labels, stats, centroids = cv.connectedComponentsWithStats(mask_morph)
# Map component labels to hue val
label_hue = np.uint8(179*labels/np.max(labels))
blank_ch = 255*np.ones_like(label_hue)
labeled_img = cv.merge([label_hue, blank_ch, blank_ch])
# cvt to BGR for display
labeled_img = cv.cvtColor(labeled_img, cv.COLOR_HSV2BGR)
# set bg label to black
labeled_img[label_hue==0] = 0
#print('Cantidad de componentes conectados: ', retval)
if retval > 2:
area_min = 200
for i in range(0, retval):
area = stats[i,cv.CC_STAT_AREA]
if area < area_min:
#print('Area: {}'.format(area))
indices = np.where(labels == i)
mask[indices] = 0
labeled_img = mask
#plt.imshow(labeled_img)
img_erosion = cv.erode(labeled_img, kernel, iterations=1)
labeled_img = cv.dilate(img_erosion,kernel, iterations=3)
return labeled_img
else :
#plt.imshow(labeled_img)
img_erosion = cv.erode(labeled_img, kernel, iterations=1)
labeled_img = cv.dilate(img_erosion,kernel, iterations=3)
return labeled_img
def untitled(tc4):
img = tc4
gray = cv.cvtColor(img, cv.COLOR_BGR2GRAY)
gray = cv.blur(gray, (3,3))
canny_output = cv.Canny(gray, 60, 60 * 2)
_, contours, _ = cv.findContours(canny_output,cv.RETR_TREE,cv.CHAIN_APPROX_NONE)
contours_poly = [None]*len(contours)
boundRect = [None]*len(contours)
for i, c in enumerate(contours):
contours_poly[i] = cv.approxPolyDP(c, 3, True)
boundRect[i] = cv.boundingRect(contours_poly[i])
drawing = np.zeros((canny_output.shape[0], canny_output.shape[1], 3), dtype=np.uint8)
for i in range(len(contours)):
color = (rng.randint(0,256), rng.randint(0,256), rng.randint(0,256))
cv.rectangle(drawing, (int(boundRect[i][0]), int(boundRect[i][1])), \
(int(boundRect[i][0]+boundRect[i][2]), int(boundRect[i][1]+boundRect[i][3])), color, 2)
#plt.imshow(drawing)
a,b,c,d = boundRect [0]
return a,b,c,d, boundRect
| Segmentacion/SegMenTacION)_Otsu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="be9d36d21776f47dcc5f8458ecfca473718c13bf"
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# This Python 3 environment comes with many helpful analytics libraries installed
# It is defined by the kaggle/python docker image: https://github.com/kaggle/docker-python
# For example, here's several helpful packages to load in
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
# Input data files are available in the "../input/" directory.
# For example, running this (by clicking run or pressing Shift+Enter) will list the files in the input directory
import os
import pathlib
import shutil
print(os.listdir("../input"))
# Any results you write to the current directory are saved as output.
from sklearn import metrics
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
from fastai.imports import *
from fastai.torch_imports import *
from fastai.transforms import *
from fastai.conv_learner import *
from fastai.model import *
from fastai.dataset import *
from fastai.sgdr import *
from fastai.plots import *
# + _uuid="bc768c376526fd9dd58000eb23c01a40adc1a90b"
PATH="../input/dogs-vs-cats-redux-kernels-edition"
# + _uuid="9d0c746478074f7e58698fa404abf5f56f0c4285"
os.makedirs(f'../working/train/dog', exist_ok=True)
os.makedirs(f'../working/train/cat', exist_ok=True)
os.makedirs(f'../working/valid/dog', exist_ok=True)
os.makedirs(f'../working/valid/cat', exist_ok=True)
os.makedirs(f'../working/test', exist_ok=True)
# + _uuid="5fbfb38ac6a780395d44fc9faaf3666c9c9296e9"
filenames = os.listdir(f'{PATH}/train')
# + _uuid="1c04ef114b9c8d41fb51ad1dae06d26153259051"
dog_indexes = []
cat_indexes = []
for fn in filenames:
if 'dog' in fn:
dog_indexes.append(fn.split('.')[1])
else:
cat_indexes.append(fn.split('.')[1])
print(len(dog_indexes), len(cat_indexes))
# + [markdown] _uuid="dfc23e20a5e5a5757305d01a7570e7fdadb85579"
# Get Validation set
# + _uuid="74a065e806196f5f106baeaec4b3b24d78ba3aa0"
val_dog_indexes = random.sample(dog_indexes, int(len(dog_indexes) * 0.2))
val_cat_indexes = random.sample(cat_indexes, int(len(cat_indexes) * 0.2))
print(len(val_dog_indexes), len(val_cat_indexes))
# + _uuid="e820ecf9f7a9e10d11c678cd1e279c9a3fd7dfd2"
for fn in filenames:
if 'dog' in fn:
dog_id = fn.split('.')[1]
if dog_id in val_dog_indexes:
shutil.copy(f'{PATH}/train/{fn}', f'../working/valid/dog')
else:
shutil.copy(f'{PATH}/train/{fn}', f'../working/train/dog')
if 'cat' in fn:
cat_id = fn.split('.')[1]
if cat_id in val_cat_indexes:
shutil.copy(f'{PATH}/train/{fn}', f'../working/valid/cat')
else:
shutil.copy(f'{PATH}/train/{fn}', f'../working/train/cat')
# + _uuid="792c4a5d2a66dbf6e5fb88be3da598342729dfbd"
for fn in os.listdir(f'{PATH}/test'):
shutil.copy(f'{PATH}/test/{fn}', f'../working/test')
# + _uuid="133d885556162679d9bbec4b44be1bfb6153c90c"
PATH = '../working'
print(len(os.listdir(f'{PATH}/train/cat')))
print(len(os.listdir(f'{PATH}/train/dog')))
print(len(os.listdir(f'{PATH}/valid/cat')))
print(len(os.listdir(f'{PATH}/valid/dog')))
# + _uuid="e788ef7baf06ccaeccbfb9069c52c99536d589cc"
cat_filenames = os.listdir(f'{PATH}/train/cat')
sample_image = plt.imread(f'{PATH}/train/cat/{cat_filenames[0]}')
plt.imshow(sample_image)
# + _uuid="e340c157767b673d4041318fb71a3d6132f97217"
sample_image.shape
# + _uuid="1fd7af917de875a9ae68694b6f7e87bf06133aa1"
filename_size_map = {fn: PIL.Image.open(f'{PATH}/train/cat/{fn}').size for fn in cat_filenames}
# + _uuid="4fc43f09e060bd443e3d5a5b0f6f17e2089b0b48"
rows, cols = zip(*filename_size_map.values())
# + _uuid="d057ff0eb5fbab0db6aa98f63208340bf161205e"
plt.hist(rows)
# + _uuid="5adf292474d05f9d74a4d344e34310340e0fab00"
plt.hist(cols)
# + _uuid="6ef038d169bdebb3d1909341ce3382b1fb28b0a9"
arch = resnet152
# + _uuid="4e52313a126bbe0361fd89e73bcabc974951da99"
def get_data(img_size, batch_size):
tfms = tfms_from_model(arch, img_size, transforms_side_on, 1.1)
return ImageClassifierData.from_paths(f'{PATH}/', batch_size, tfms, test_name='test')
# + _uuid="9d160b0d8ae063533baac0dbfb3589e3505362a2"
data = get_data(224, 64)
data.path = pathlib.Path('.')
learn = ConvLearner.pretrained(arch, data, precompute=False)
# + _uuid="d13afefc24702a0ac016e9374a630c7ddff2c916"
learn.lr_find()
# + _uuid="d066bb46bf97b6c9ff46ea61ed565c8bbf37ba1f"
learn.sched.plot()
# + _uuid="7610ddbd4e31e82724172816fb9bee005775878c"
learn.fit(1e-2, 3, cycle_len=1, cycle_mult=2)
# + _uuid="eec53b281c9d543598da6e0fe6386c3d049c5280"
learn.save('last_layer_226')
# + _uuid="dde3afb8265f12859b4957b55ff32e8a9b076ccd"
learn.load('last_layer_226')
# + _uuid="a2e9c1919c8a62fd221ac877ebcec84e95d37dc6"
log_prob, y = learn.TTA(is_test=True)
prob = np.mean(np.exp(log_prob), 0)
# + _uuid="9937af34d22425386a916b978afe38c8a11cabab"
df = pd.DataFrame(prob[:, 1])
submit_df = df.rename(columns = {0:'label'})
submit_df.head()
submit_df.size
# + _uuid="119b228b1c7a75e611705c831a1a88f4f1fce919"
submit_df.insert(0, 'id', [fn[:-4] for fn in os.listdir(f'{PATH}/test')])
submit_df.head()
# + _uuid="1baf042546b637ab13d44618ad621efba4d4f53f"
SUBMIT_PATH = './submit'
os.makedirs(SUBMIT_PATH, exist_ok=True)
submit_df.to_csv(f'{SUBMIT_PATH}/submit.gz', compression='gzip', index=False)
# + _uuid="8fa31cd43539adcecd81958ce5e17fd1e737be89"
FileLink(f'{SUBMIT_PATH}/submit.gz')
# + _uuid="b0d0a54a17951ffae434bad54e2621353a60bf45"
| dog_cat_redux/dog_cat_redux.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .ps1
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .NET (PowerShell)
# language: PowerShell
# name: .net-powershell
# ---
# # T1055.001 - Dynamic-link Library Injection
# Adversaries may inject dynamic-link libraries (DLLs) into processes in order to evade process-based defenses as well as possibly elevate privileges. DLL injection is a method of executing arbitrary code in the address space of a separate live process.
#
# DLL injection is commonly performed by writing the path to a DLL in the virtual address space of the target process before loading the DLL by invoking a new thread. The write can be performed with native Windows API calls such as <code>VirtualAllocEx</code> and <code>WriteProcessMemory</code>, then invoked with <code>CreateRemoteThread</code> (which calls the <code>LoadLibrary</code> API responsible for loading the DLL). (Citation: Endgame Process Injection July 2017)
#
# Variations of this method such as reflective DLL injection (writing a self-mapping DLL into a process) and memory module (map DLL when writing into process) overcome the address relocation issue as well as the additional APIs to invoke execution (since these methods load and execute the files in memory by manually preforming the function of <code>LoadLibrary</code>).(Citation: Endgame HuntingNMemory June 2017)(Citation: Endgame Process Injection July 2017)
#
# Running code in the context of another process may allow access to the process's memory, system/network resources, and possibly elevated privileges. Execution via DLL injection may also evade detection from security products since the execution is masked under a legitimate process.
# ## Atomic Tests:
# Currently, no tests are available for this technique.
# ## Detection
# Monitoring Windows API calls indicative of the various types of code injection may generate a significant amount of data and may not be directly useful for defense unless collected under specific circumstances for known bad sequences of calls, since benign use of API functions may be common and difficult to distinguish from malicious behavior. Windows API calls such as <code>CreateRemoteThread</code> and those that can be used to modify memory within another process, such as <code>VirtualAllocEx</code>/<code>WriteProcessMemory</code>, may be used for this technique.(Citation: Endgame Process Injection July 2017)
#
# Monitor DLL/PE file events, specifically creation of these binary files as well as the loading of DLLs into processes. Look for DLLs that are not recognized or not normally loaded into a process.
#
# Analyze process behavior to determine if a process is performing actions it usually does not, such as opening network connections, reading files, or other suspicious actions that could relate to post-compromise behavior.
| playbook/tactics/privilege-escalation/T1055.001.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # AGDCv2 Landsat analytics example using USGS Surface Reflectance
# ### Import the required libraries
# %matplotlib inline
from matplotlib import pyplot as plt
import datacube
from datacube.model import Range
from datetime import datetime
dc = datacube.Datacube(app='dc-example')
from datacube.storage import masking
from datacube.storage.masking import mask_valid_data as mask_invalid_data
import pandas
import xarray
import numpy
import json
import vega
from datacube.utils import geometry
numpy.seterr(divide='ignore', invalid='ignore')
import folium
from IPython.display import display
import geopandas
from shapely.geometry import mapping
from shapely.geometry import MultiPolygon
import rasterio
import shapely.geometry
import shapely.ops
from functools import partial
import pyproj
from datacube.model import CRS
from datacube.utils import geometry
## From http://scikit-image.org/docs/dev/auto_examples/plot_equalize.html
from skimage import data, img_as_float
from skimage import exposure
datacube.__version__
# ### Include some helpful functions
def datasets_union(dss):
thing = geometry.unary_union(ds.extent for ds in dss)
return thing.to_crs(geometry.CRS('EPSG:4326'))
import random
def plot_folium(shapes):
mapa = folium.Map(location=[17.38,78.48], zoom_start=8)
colors=['#00ff00', '#ff0000', '#00ffff', '#ffffff', '#000000', '#ff00ff']
for shape in shapes:
style_function = lambda x: {'fillColor': '#000000' if x['type'] == 'Polygon' else '#00ff00',
'color' : random.choice(colors)}
poly = folium.features.GeoJson(mapping(shape), style_function=style_function)
mapa.add_children(poly)
display(mapa)
# determine the clip parameters for a target clear (cloud free image) - identified through the index provided
def get_p2_p98(rgb, red, green, blue, index):
r = numpy.nan_to_num(numpy.array(rgb.data_vars[red][index]))
g = numpy.nan_to_num(numpy.array(rgb.data_vars[green][index]))
b = numpy.nan_to_num(numpy.array(rgb.data_vars[blue][index]))
rp2, rp98 = numpy.percentile(r, (2, 99))
gp2, gp98 = numpy.percentile(g, (2, 99))
bp2, bp98 = numpy.percentile(b, (2, 99))
return(rp2, rp98, gp2, gp98, bp2, bp98)
def plot_rgb(rgb, rp2, rp98, gp2, gp98, bp2, bp98, red, green, blue, index):
r = numpy.nan_to_num(numpy.array(rgb.data_vars[red][index]))
g = numpy.nan_to_num(numpy.array(rgb.data_vars[green][index]))
b = numpy.nan_to_num(numpy.array(rgb.data_vars[blue][index]))
r_rescale = exposure.rescale_intensity(r, in_range=(rp2, rp98))
g_rescale = exposure.rescale_intensity(g, in_range=(gp2, gp98))
b_rescale = exposure.rescale_intensity(b, in_range=(bp2, bp98))
rgb_stack = numpy.dstack((r_rescale,g_rescale,b_rescale))
img = img_as_float(rgb_stack)
return(img)
def plot_water_pixel_drill(water_drill):
vega_data = [{'x': str(ts), 'y': str(v)} for ts, v in zip(water_drill.time.values, water_drill.values)]
vega_spec = """{"width":720,"height":90,"padding":{"top":10,"left":80,"bottom":60,"right":30},"data":[{"name":"wofs","values":[{"code":0,"class":"dry","display":"Dry","color":"#D99694","y_top":30,"y_bottom":50},{"code":1,"class":"nodata","display":"No Data","color":"#A0A0A0","y_top":60,"y_bottom":80},{"code":2,"class":"shadow","display":"Shadow","color":"#A0A0A0","y_top":60,"y_bottom":80},{"code":4,"class":"cloud","display":"Cloud","color":"#A0A0A0","y_top":60,"y_bottom":80},{"code":1,"class":"wet","display":"Wet","color":"#4F81BD","y_top":0,"y_bottom":20},{"code":3,"class":"snow","display":"Snow","color":"#4F81BD","y_top":0,"y_bottom":20},{"code":255,"class":"fill","display":"Fill","color":"#4F81BD","y_top":0,"y_bottom":20}]},{"name":"table","format":{"type":"json","parse":{"x":"date"}},"values":[],"transform":[{"type":"lookup","on":"wofs","onKey":"code","keys":["y"],"as":["class"],"default":null},{"type":"filter","test":"datum.y != 255"}]}],"scales":[{"name":"x","type":"time","range":"width","domain":{"data":"table","field":"x"},"round":true},{"name":"y","type":"ordinal","range":"height","domain":["water","not water","not observed"],"nice":true}],"axes":[{"type":"x","scale":"x","formatType":"time"},{"type":"y","scale":"y","tickSize":0}],"marks":[{"description":"data plot","type":"rect","from":{"data":"table"},"properties":{"enter":{"xc":{"scale":"x","field":"x"},"width":{"value":"1"},"y":{"field":"class.y_top"},"y2":{"field":"class.y_bottom"},"fill":{"field":"class.color"},"strokeOpacity":{"value":"0"}}}}]}"""
spec_obj = json.loads(vega_spec)
spec_obj['data'][1]['values'] = vega_data
return vega.Vega(spec_obj)
# ## Plot the spatial extent of our data for each product
plot_folium([datasets_union(dc.index.datasets.search_eager(product='ls5_ledaps_scene')),\
datasets_union(dc.index.datasets.search_eager(product='ls7_ledaps_scene')),\
datasets_union(dc.index.datasets.search_eager(product='ls8_ledaps_scene'))])
# ## Inspect the available measurements for each product
dc.list_measurements()
# ## Specify the Area of Interest for our analysis
# +
# Hyderbad
# 'lon': (78.40, 78.57),
# 'lat': (17.36, 17.52),
# Lake Singur
# 'lat': (17.67, 17.84),
# 'lon': (77.83, 78.0),
# Lake Singur Dam
query = {
'lat': (17.72, 17.79),
'lon': (77.88, 77.95),
}
# -
# ## Load Landsat Surface Reflectance for our Area of Interest
# +
products = ['ls5_ledaps_scene','ls7_ledaps_scene','ls8_ledaps_scene']
datasets = []
for product in products:
ds = dc.load(product=product, measurements=['nir','red', 'green','blue'], output_crs='EPSG:32644',resolution=(-30,30), **query)
ds['product'] = ('time', numpy.repeat(product, ds.time.size))
datasets.append(ds)
sr = xarray.concat(datasets, dim='time')
sr = sr.isel(time=sr.time.argsort()) # sort along time dim
sr = sr.where(sr != -9999)
# +
##### include an index here for the timeslice with representative data for best stretch of time series
# don't run this to keep the same limits as the previous sensor
#rp2, rp98, gp2, gp98, bp2, bp98 = get_p2_p98(sr,'red','green','blue', 0)
rp2, rp98, gp2, gp98, bp2, bp98 = (300.0, 2000.0, 300.0, 2000.0, 300.0, 2000.0)
print(rp2, rp98, gp2, gp98, bp2, bp98)
# -
plt.imshow(plot_rgb(sr,rp2, rp98, gp2, gp98, bp2, bp98,'red',
'green', 'blue', 0),interpolation='nearest')
# ## Load Landsat Pixel Quality for our area of interest
# +
datasets = []
for product in products:
ds = dc.load(product=product, measurements=['cfmask'], output_crs='EPSG:32644',resolution=(-30,30), **query).cfmask
ds['product'] = ('time', numpy.repeat(product, ds.time.size))
datasets.append(ds)
pq = xarray.concat(datasets, dim='time')
pq = pq.isel(time=pq.time.argsort()) # sort along time dim
del(datasets)
# -
# ## Visualise pixel quality information from our selected spatiotemporal subset
pq.attrs['flags_definition'] = {'cfmask': {'values': {'255': 'fill', '1': 'water', '2': 'shadow', '3': 'snow', '4': 'cloud', '0': 'clear'}, 'description': 'CFmask', 'bits': [0, 1, 2, 3, 4, 5, 6, 7]}}
pandas.DataFrame.from_dict(masking.get_flags_def(pq), orient='index')
# ### Plot the frequency of water classified in pixel quality
water = masking.make_mask(pq, cfmask ='water')
water.sum('time').plot(cmap='nipy_spectral')
# ### Plot the timeseries at the center point of the image
plot_water_pixel_drill(pq.isel(y=int(water.shape[1] / 2), x=int(water.shape[2] / 2)))
del(water)
# ## Remove the cloud and shadow pixels from the surface reflectance
mask = masking.make_mask(pq, cfmask ='cloud')
mask = abs(mask*-1+1)
sr = sr.where(mask)
mask = masking.make_mask(pq, cfmask ='shadow')
mask = abs(mask*-1+1)
sr = sr.where(mask)
del(mask)
del(pq)
sr.attrs['crs'] = CRS('EPSG:32644')
# ## Spatiotemporal summary NDVI median
ndvi_median = ((sr.nir-sr.red)/(sr.nir+sr.red)).median(dim='time')
ndvi_median.attrs['crs'] = CRS('EPSG:32644')
ndvi_median.plot(cmap='YlGn', robust='True')
# ## NDVI trend over time in cropping area Point Of Interest
poi_latitude = 17.749343
poi_longitude = 77.935634
p = geometry.point(x=poi_longitude, y=poi_latitude, crs=geometry.CRS('EPSG:4326')).to_crs(sr.crs)
# ### Create a subset around our point of interest
subset = sr.sel(x=((sr.x > p.points[0][0]-1000)), y=((sr.y < p.points[0][1]+1000)))
subset = subset.sel(x=((subset.x < p.points[0][0]+1000)), y=((subset.y > p.points[0][1]-1000)))
# ### Plot subset image with POI at centre
plt.imshow(plot_rgb(subset,rp2, rp98, gp2, gp98, bp2, bp98,'red',
'green', 'blue',0),interpolation='nearest' )
# ### NDVI timeseries plot
((sr.nir-sr.red)/(sr.nir+sr.red)).sel(x=p.points[0][0], y=p.points[0][1], method='nearest').plot(marker='o')
| notebooks_ledaps/hyderabad_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES']='0'
from deoldify.visualize import *
plt.style.use('dark_background')
torch.backends.cudnn.benchmark=True
colorizer = get_video_colorizer()
# # Instructions
#
# ### source_url
# Type in a url hosting a video from YouTube, Imgur, Twitter, Reddit, Vimeo, etc. Many sources work! GIFs also work. Full list here: https://ytdl-org.github.io/youtube-dl/supportedsites.html NOTE: If you want to use your own video, you can set source_url to None and just upload the file to video/source/ in Jupyter. Just make sure that the file_name parameter matches the file you uploaded.
#
#
# ### file_name
# Name this whatever sensible file name you want (minus extension)! It should actually exist in video/source if source_url=None
#
#
# ### render_factor
# The default value of 21 has been carefully chosen and should work -ok- for most scenarios (but probably won't be the -best-). This determines resolution at which the color portion of the video is rendered. Lower resolution will render faster, and colors also tend to look more vibrant. Older and lower quality film in particular will generally benefit by lowering the render factor. Higher render factors are often better for higher quality videos and inconsistencies (flashy render) will generally be reduced, but the colors may get slightly washed out.
#
#
# ### file_name_ext
# There's no reason to changes this.
#
#
# ### result_path
# Ditto- don't change.
#
#
# ### How to Download a Copy
# Simply shift+right click on the displayed video and click "Save video as..."!
#
#
# ## Pro Tips
# 1. If a video takes a long time to render and you're wondering how well the frames will actually be colorized, you can preview how well the frames will be rendered at each render_factor by using the code at the bottom. Just stop the video rendering by hitting the stop button on the cell, then run that bottom cell under "See how well render_factor values perform on a frame here". It's not perfect and you may still need to experiment a bit especially when it comes to figuring out how to reduce frame inconsistency. But it'll go a long way in narrowing down what actually works.
#
#
# ## Troubleshooting
# The video player may wind up not showing up, in which case- make sure to wait for the Jupyter cell to complete processing first (the play button will stop spinning). Then follow these alternative download instructions
#
# 1. In the menu to the left, click Home icon.
# 2. By default, rendered video will be in /video/result/
#
# If a video you downloaded doesn't play, it's probably because the cell didn't complete processing and the video is in a half-finished state.
# If you get a 'CUDA out of memory' error, you probably have the render_factor too high. The max is 44 on 11GB video cards.
# ## Colorize!!
# +
#NOTE: Max is 44 with 11GB video cards. 21 is a good default
render_factor=21
#NOTE: Make source_url None to just read from file at ./video/source/[file_name] directly without modification
source_url='https://twitter.com/silentmoviegifs/status/1116751583386034176'
file_name = 'DogShy1926'
file_name_ext = file_name + '.mp4'
result_path = None
if source_url is not None:
result_path = colorizer.colorize_from_url(source_url, file_name_ext, render_factor=render_factor)
else:
result_path = colorizer.colorize_from_file_name(file_name_ext)
show_video_in_notebook(result_path)
# -
# ## See how well render_factor values perform on a frame here
for i in range(10,45,2):
colorizer.vis.plot_transformed_image('video/bwframes/' + file_name + '/00001.jpg', render_factor=i, display_render_factor=True, figsize=(8,8))
| VideoColorizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Thermal Conduction
# This examples shows how OpenPNM can be used to simulate thermal conduction on a generic grid of nodes. The result obtained from OpenPNM is compared to the analytical result.
# As usual, start by importing OpenPNM, and the SciPy library.
# %matplotlib inline
import numpy as np
import scipy as sp
import openpnm as op
# %config InlineBackend.figure_formats = ['svg']
np.random.seed(10)
ws = op.Workspace()
ws.settings["loglevel"] = 40
np.set_printoptions(precision=5)
# ## Generating the Network object
#
# Next, 2D a **Network** is generated with dimensions of 10x50 elements. The lattice spacing is given by *Lc*. Boundaries are added all around the edges of **Network** object using the ``add_boundariy_pores`` method.
divs = [10, 50]
Lc = 0.1 # cm
pn = op.network.Cubic(shape=divs, spacing=Lc)
pn.add_boundary_pores(['left', 'right', 'front', 'back'])
# ## Creating a Phase object
# All simulations require a phase object which possess the thermosphysical properties of the system. In this case, we'll create a generic phase object, call it copper, though it has no properties; we'll add these by hand later.
# Create Phase object and associate with a Physics object
Cu = op.phases.GenericPhase(network=pn)
# ## Assigning Thermal Conductance to Copper
# In a proper OpenPNM model we would create a Geometry object to manage all the geometrical properties, and a Physics object to calculate the thermal conductance based on the geometric information and the thermophysical properties of copper. In the present case, however, we'll just calculate the conductance manually and assign it to ```Cu```.
# Add a unit conductance to all connections
Cu['throat.thermal_conductance'] = 1
# Overwrite boundary conductances since those connections are half as long
Ps = pn.pores('*boundary')
Ts = pn.find_neighbor_throats(pores=Ps)
Cu['throat.thermal_conductance'][Ts] = 2
# ## Generating the algorithm objects and running the simulation
# The last step in the OpenPNM simulation involves the generation of a **Algorithm** object and running the simulation.
# Setup Algorithm object
alg = op.algorithms.FourierConduction(network=pn)
alg.setup(phase=Cu)
inlets = pn.pores('right_boundary')
outlets = pn.pores(['front_boundary', 'back_boundary', 'right_boundary'])
T_in = 30*np.sin(np.pi*pn['pore.coords'][inlets, 1]/5)+50
alg.set_value_BC(values=T_in, pores=inlets)
alg.set_value_BC(values=50, pores=outlets)
alg.run()
# This is the last step usually required in a OpenPNM simulation. The algorithm was run, and now the simulation data obtained can be analyzed. For illustrative purposes, the results obtained using OpenPNM shall be compared to an analytical solution of the problem in the following.
# First let's rehape the 'pore.temperature' array into the shape of the network while also extracting only the internal pores to avoid showing the boundaries.
import matplotlib.pyplot as plt
sim = alg['pore.temperature'][pn.pores('internal')]
temp_map = np.reshape(a=sim, newshape=divs)
plt.subplots(1, 1, figsize=(10, 5))
plt.imshow(temp_map, cmap=plt.cm.plasma);
plt.colorbar();
# Also, let's take a look at the average temperature:
print(f"T_average (numerical): {alg['pore.temperature'][pn.pores('internal')].mean():.5f}")
# The analytical solution is computed as well, and the result is the same shape as the network (including the boundary pores).
# Calculate analytical solution over the same domain spacing
X = pn['pore.coords'][:, 0]
Y = pn['pore.coords'][:, 1]
soln = 30*np.sinh(np.pi*X/5)/np.sinh(np.pi/5)*np.sin(np.pi*Y/5) + 50
soln = soln[pn.pores('internal')]
soln = np.reshape(soln, (divs[0], divs[1]))
plt.subplots(1, 1, figsize=(10, 5))
plt.imshow(soln, cmap=plt.cm.plasma);
plt.colorbar();
# Also, let's take a look at the average temperature:
print(f"T_average (analytical): {soln.mean():.5f}")
# Both the analytical solution and OpenPNM simulation can be subtracted from each other to yield the difference in both values.
diff = soln - temp_map
plt.subplots(1, 1, figsize=(10, 5))
plt.imshow(diff, cmap=plt.cm.plasma);
plt.colorbar();
print(f"Minimum error: {diff.min():.5f}, maximum error: {diff.max():.5f}")
# The maximum error is 0.01 degrees on a 50 degree profile, which is quite good and thus demonstrates that the OpenPNM finite difference approach is versatile despite being simple.
| examples/notebooks/algorithms/single_phase/continuum_heat_transfer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Approximation
# ## Outline
#
# 1. Setup
# 2. Polynomial interpolation
# 3. Resources
# ## Setup
# coefficientscoefficientscoefficientsIn many computational economics applications, we need to replace an analytically intractable function $f : R^n \rightarrow R$ with a numerically tractable approximation $\hat{f}$. In some applications, f can be evaluated at any point of its domain, but with difficulty, and we wish to replace it with an approximation $\hat{f}$ that is easier to work with.
#
# We study interpolation, a general strategy for forming a tractable approximation to a function that can be evaluated at any point of its domain. Consider a real-valued function $f$ defined on an interval of the real line that can be evaluated at any point of its domain.
#
# Generally, we will approximate $f$ using a function $\hat{f}$ that is a finite linear combination of n known basis functions $\phi_1, \phi_2, ..., \phi_n$ of our choosing:
#
# \begin{align*}
# f(x) \approx \hat{f}(x) \equiv \sum_{j=1}^n c_j \phi_j(x).
# \end{align*}
#
# We will fix the n basis coefficients $c_1 , c_2 , ... , c_n$ by requiring $\hat{f}$ to interpolate, that is, agree with $f$ , at $n$ interpolation nodes $x_1 , x_2 , ... , x_n$ of our choosing.
#
# The most readily recognizable basis is the monomial basis:
#
# \begin{align*}
# \phi_0(x) &= 1 \\
# \phi_1(x) &= x \\
# \phi_2(x) &= x^2 \\
# . \\
# \phi_n(x)&= x^n.
# \end{align*}
# This can be used to construct the polynomial approximations:
#
# \begin{align*}
# f(x) \approx \hat{f}(x) \equiv c_0 + c_1 x + c_2 x^2 + ... c_n x^n
# \end{align*}
#
# There are other basis functions with more desirable properties and there are many different ways to choose the interpolation nodes.
#
# Regardless of how the $n$ basis functions and nodes are chosen, computing the basis coefficients reduces to solving a linear equation.
#
# \begin{align*}
# \sum_{j=1}^n c_j \phi_j(x) = f(x), \qquad i = 1, ..., n
# \end{align*}
#
# Interpolation schemes differ only in how the basis functions $\phi_j$ and interpolation nodes $x_j$ are chosen.
#
#
#
# +
from functools import partial
from temfpy.interpolation import runge
import warnings
from scipy.interpolate import interp1d
import matplotlib.pyplot as plt
import numpy as np
from approximation_algorithms import get_interpolator_flexible_basis_flexible_nodes
from approximation_algorithms import get_interpolator_monomial_flexible_nodes
from approximation_algorithms import get_interpolator_monomial_uniform
from approximation_algorithms import get_interpolator_runge_baseline
from approximation_auxiliary import compute_interpolation_error
from approximation_auxiliary import get_chebyshev_nodes
from approximation_auxiliary import get_uniform_nodes
from approximation_plots import plot_two_dimensional_problem
from approximation_plots import plot_reciprocal_exponential
from approximation_plots import plot_runge_different_nodes
from approximation_plots import plot_runge_function_cubic
from approximation_plots import plot_two_dimensional_grid
from approximation_plots import plot_approximation_nodes
from approximation_plots import plot_basis_functions
from approximation_plots import plot_runge_multiple
from approximation_plots import plot_runge
from approximation_problems import problem_reciprocal_exponential
from approximation_problems import problem_two_dimensions
# -
# ## Polynomial interpolation
# A polynomial is an expression consisting of variables and coefficients, that involves only the operations of addition, subtraction, multiplication, and non-negative integer exponentiation of variables.
#
# The Weierstrass Theorem asserts that any continuous real-valued function can be approximated to an arbitrary degree of accuracy over a bounded interval by a polynomial.
#
# Specifically, if $f$ is continuous on $[a, b]$ and $\epsilon > 0$, then there exists a polynomial $p$ such that
#
# \begin{align*}
# \max_{x\in[a, b]} |f(x) - p(x)| < \epsilon
# \end{align*}
#
# * How to find a polynomial that provides a desired degree of accuracy?
#
# * What degree of the polynomial is required?
#
# ### Naive polynomial interpolation
#
# Let's start with a basic setup, where we use a uniform grid and monomial basis functions.
#
# \begin{align*}
# \hat{f}(x) \equiv \sum_{j=0}^n c_j x^j
# \end{align*}
??get_uniform_nodes
plot_approximation_nodes([5, 10, 15, 20], nodes="uniform")
# Now we can get a look at the interpolation nodes.
plot_basis_functions("monomial")
# Let's look at the performance of this approach for the Runge function for $x\in[0, 1]$.
#
# \begin{align*}
# f(x) = \frac{1}{(1 + 25 x^2)}
# \end{align*}
plot_runge()
# Due to its frequent use, `numpy` does offer a convenience class to work with polynomials. See [here](https://numpy.org/devdocs/reference/routines.polynomials.html) for its documentation.
from numpy.polynomial import Polynomial as P # noqa E402
from numpy.polynomial import Chebyshev as C # noqa E402
# We will use the attached methods to develop a flexible interpolation set in an iterative fashion.'
??get_interpolator_runge_baseline
with warnings.catch_warnings():
warnings.simplefilter("ignore")
interpolant = get_interpolator_runge_baseline(runge)
xvalues = np.linspace(-1, 1, 10000)
yfit = interpolant(xvalues)
# ### _Question_
#
# * Why the warnings?
#
# Since we have a good understanding what is causing the warning, we can simply turn it of going forward. A documentation that shows how to deal with more fine-grained filters is available [here](https://pymotw.com/3/warnings/).
warnings.simplefilter("ignore")
# Now we are read to plot it against the true function.
fig, ax = plt.subplots()
ax.plot(xvalues, runge(xvalues), label="True")
ax.plot(xvalues, yfit, label="Approximation")
ax.legend()
# We evaluate the error in our approximation by the the following statistic.
??compute_interpolation_error
compute_interpolation_error(yfit - runge(xvalues))
# ### _Exercises_
#
# 1. Generalize the function to allow to approximate the function with a polynomial of generic degree.
# 2. How does the quality of the approximation change as we increase the number of interpolation points?
with warnings.catch_warnings():
warnings.simplefilter("ignore")
plot_runge_multiple()
# What can be done? First we explore a different way to choose the the nodes.
#
# Theory asserts that the best way to approximate a continuous function with a polynomial over a bounded interval $[a, b]$ is to interpolate it at so called Chebychev nodes:
#
# \begin{align*}
# x_i = \frac{a + b}{2} + \frac{b - a}{2}\cos\left(\frac{n - i + 0.5}{n}\pi\right)
# \end{align*}
#
#
??get_chebyshev_nodes
# Let's look at a visual representation.
plot_approximation_nodes([5, 10, 15, 20], nodes="chebychev")
# The Chebychev nodes are not evenly spaced and do not include the endpoints of the approximation interval. They are more closely spaced near the endpoints of the approximation interval and less so near the center.
#
#
# If $f$ is continuous ...
#
# * Rivlin’s Theorem asserts that Chebychev-node polynomial interpolation is nearly optimal, that is, it affords an approximation error that is very close to the lowest error attainable with another polynomial of the same degree.
#
# * Jackson’s Theorem asserts that Chebychev-node polynomial interpolation is consistent, that is, the approximation error vanishes as the degree of the polynomial increases.
??get_interpolator_monomial_flexible_nodes
intertp = get_interpolator_monomial_flexible_nodes(runge, 11, nodes="chebychev")
intertp(np.linspace(-1, 1, 10))
# Let's compare the performance of the two approaches.
plot_runge_different_nodes()
# However, merely interpolating at the Chebychev nodes does not eliminate ill-conditioning. Ill-conditioning stems from the choice of basis functions,
# not the choice of interpolation nodes. Fortunately, there is alternative to the monomial basis that is ideal for expressing Chebychev-node polynomial interpolants. The optimal basis for expressing Chebychev-node
# polynomial interpolants is called the Chebychev polynomial basis.
plot_basis_functions("chebychev")
# Combining the Chebychev basis polynomials and Chebychev interpolation nodes yields an extremely well-conditioned interpolation equation and allows to approximate any continuous function to high precision. Let's put it all together now.
??get_interpolator_flexible_basis_flexible_nodes
# How well can we actually do now?
for degree in [5, 10, 15]:
interp = get_interpolator_flexible_basis_flexible_nodes(
runge, degree, nodes="uniform", basis="monomial"
)
xvalues = np.linspace(-1, 1, 10000)
yfit = interp(xvalues)
fig, ax = plt.subplots()
ax.plot(xvalues, runge(xvalues), label="True")
ax.plot(xvalues, yfit, label="Approximation")
ax.legend()
ax.set_title(f"Degree {degree}")
# ### Spline interpolation
#
# Piecewise polynomial splines, or simply splines for short, are a rich, flexible class of functions that may be used instead of high degree polynomials to approximate a real-valued function over a bounded interval. Generally, an order $k$ spline consists of a series of $k^{th}$
# degree polynomial segments spliced together so as to preserve continuity of derivatives of order $k - 1$ or less
#
# * A first-order or **linear spline** is a series of line segments spliced together to form a continuous function.
#
# * A third-order or **cubic spline** is a series of cubic polynomials segments spliced together to form a twice continuously differentiable function.
#
#
# <img src="material/fig-spline-two.png" width=500 height=500 />
# <img src="material/fig-spline-four.png" width=500 height=500 />
# <img src="material/fig-spline-eight.png" width=500 height=500 />
#
# A linear spline with n + 1 evenly-spaced interpolation nodes $x_0 , x_1 , ... , x_n$ on the interval $[a, b]$ may be written as a linear combination of the $n + 1$ basis functions:
#
# \begin{align*}
# \phi_j(x) = \begin{cases}
# 1 - \frac{|x - x_j|}{h} & \qquad |x - x_j| \leq h \\
# 0 & \\
# \end{cases}
# \end{align*}
#
# where $h = (b - a)/n$ is the distance between the nodes.
#
#
# The linear spline approximant of $f$ takes thus the form:
#
# \begin{align*}
# \hat{f}(x) = \sum_{j=1}^{n} f(x_j)\phi_j(x)
# \end{align*}
plot_basis_functions("linear")
# This kind of interpolation procedure is frequently used in practice and readily available in `scipy`. The `interp1` function is documented [here](https://docs.scipy.org/doc/scipy/reference/generated/scipy.interpolate.interp1d.html).
x_fit = get_uniform_nodes(10, -1, 1)
f_inter = interp1d(x_fit, runge(x_fit))
f_inter(0.5)
# Let's get a feel for this approach using our earlier test function.
# +
x_eval = get_uniform_nodes(10000, -1, 1)
for degree in [3, 5, 10, 15]:
x_fit = get_uniform_nodes(degree, -1, 1)
interp = interp1d(x_fit, runge(x_fit))
yfit = interp(xvalues)
fig, ax = plt.subplots()
ax.plot(xvalues, runge(xvalues), label="True")
ax.plot(xvalues, yfit, label="Approximation")
ax.legend()
ax.set_title(f"Degree {degree}")
# -
# ### _Question_
#
# * How about other ways to place the interpolation nodes?
# Another widely-used specification relies on cubic splines. Here are the corresponding basis functions.
#
# <img src="material/fig-cubic-spline-basis.png" width=500 height=500 />
#
#
# It is directly integrated into the `interp1` function.
x_fit = get_uniform_nodes(10, -1, 1)
f_inter = interp1d(x_fit, runge(x_fit), kind="cubic")
f_inter(0.5)
# How about approximating Runge's function.
plot_runge_function_cubic()
# Let's take stock of our interpolation toolkit by running a final benchmarking exercise and then try to extract some
#
#
# ### _Exercises_
#
# Let's consider two test functions: problem_reciprocal_exponential, problem_kinked.
#
# 1. Visualize both over the range from -1 to 1. What is the key differences in their properties?
# 2. Set up a function that allows you to flexibly interpolate using either Chebychev polynomials (monomial basis, Chebychev nodes) or linear and cubic splines.
# 3. Compare the performance for the following degrees: 10, 20, 30.
# We collect some rules-of-thumb:
#
# * Chebychev-node polynomial interpolation dominates spline function interpolation whenever the function is smooth.
#
# * Spline interpolation may perform better than polynomial interpolation if the underlying function exhibits a high degree of curvature or a derivative discontinuity.
# ### Multidimensional interpolation
#
# Univariate interpolation methods can be extended to higher dimensions by applying tensor product principles. We consider the problem of interpolating a bivariate real-valued function $f$ over an interval:
#
# \begin{align*}
# I = \{(x, y) | a_x \leq x \leq b_x, a_y \leq y \leq b_y\}
# \end{align*}
#
#
# Let $\phi_{x_1} , \phi_{x_2} , ... , \phi_{x_n}$ x and $x_1 , x_2 , . . . , x_{n_x}$ be $n_x$ univariate basis functions and $n_x$ interpolation nodes for the interval $[a_x , b_x]$ and let $\phi_{y_1} , \phi_{y_2} , ... , \phi_{y_n}$ x and $y_1 , y_2 , . . . , y_{n_y}$ be $n_x$ univariate basis functions and $n_y$ interpolation nodes for the interval $[a_y , b_y]$.
#
# Then an $n = n_x n_y$ bivariate function basis defined on $I$ may be obtained by forming the tensor product of the univariate basis functions: $\phi_{ij} (x, y) = \phi^x_i (x) \phi^y_j(y)$ for $i = 1, 2, ... , n_x$ and $j = 1, 2, ... , n_y$. Similarly, a grid of $n = n_x n_y$ interpolation nodes for $I$ may be obtained by forming the Cartesian product of the univariate interpolation nodes
#
# \begin{align*}
# \{ (x_i , y_j ) | i = 1, 2, . . . , n_x ; j = 1, 2, . . . , n_y \}.
# \end{align*}
#
# Typically, multivariate tensor product interpolation schemes inherit the favorable qualities of their univariate parents. An approximant for $f$ then takes the form:
#
# \begin{align*}
# \hat{f}(x_1, x_2) = \sum_{i=1}^{n_x} \sum_{j=1}^{n_y} c_{ij}\phi_{ij}(x_i, y_j)
# \end{align*}
#
# However, this straightforward extension to the multivariate setting suffers from the **curse of dimensionality**. For example, the number of interpolation nodes increases exponentially in the number of dimensions.
#
#
# As an aside, we now move to the multidimensional setting where we often have to apply the same operation across multidimensional arrays and `numpy` provides some suitable capabilities to do this very fast if one makes an effort in understanding its [broadcasting rules](https://numpy.org/doc/stable/user/theory.broadcasting.html#array-broadcasting-in-numpy).
plot_two_dimensional_grid("uniform")
plot_two_dimensional_grid("chebychev")
# Let's see how we can transfer the ideas to polynomial interpolation to the two-dimensional setting.
#
# \begin{align*}
# f(x, y) = \frac{\cos(x)}{\sin(y)}
# \end{align*}
??plot_two_dimensional_problem
plot_two_dimensional_problem()
# Now, let's fit a two-dimensional polynomial approximation. We will have to rely on the `scikit-learn` library.
from sklearn.preprocessing import PolynomialFeatures # noqa: E402
from sklearn.linear_model import LinearRegression # noqa: E402
import sklearn # noqa: E402
# We first need to set up an approximating model using some of its provided functionality. One of the functions at the core of this workflow is `np.meshgrid` which takes a bit of getting used to. Let's check out its [documentation](https://numpy.org/doc/stable/reference/generated/numpy.meshgrid.html) first and so some explorations.
x_fit, y_fit = get_chebyshev_nodes(100), get_chebyshev_nodes(100)
# We now combine the univariate interpolation nodes into a two-dimensional grid, adjust it to meet the structure expected by `scikit-learn`, expand it to contain all polynomials (including interactions), and fit a linear regression model.
# +
X_fit, Y_fit = np.meshgrid(x_fit, y_fit)
grid_fit = np.array(np.meshgrid(x_fit, y_fit)).T.reshape(-1, 2)
y = [problem_two_dimensions(*point) for point in grid_fit]
poly = PolynomialFeatures(degree=6)
X_poly = poly.fit_transform(grid_fit)
clf = LinearRegression().fit(X_poly, y)
# -
# How well are we doing? As usual, we will simply compare the true and approximated values of the function over a fine grid.
# +
x_eval = get_uniform_nodes(100)
y_eval = get_uniform_nodes(100)
Z_eval = np.tile(np.nan, (100, 100))
Z_true = np.tile(np.nan, (100, 100))
for i, x in enumerate(x_eval):
for j, y in enumerate(y_eval):
point = [x, y]
Z_eval[i, j] = clf.predict(poly.fit_transform([point]))[0]
Z_true[i, j] = problem_two_dimensions(*point)
fig = plt.figure()
ax = fig.gca(projection="3d")
ax.plot_surface(*np.meshgrid(x_eval, y_eval), Z_eval - Z_true)
# -
#
# ## Resources
#
# * https://relate.cs.illinois.edu/course/cs450-f18/file-version/a7a1965adf0479d36f1a34889afe55e2ec61a532/demos/upload/07-interpolation/Chebyshev%20interpolation.html
#
# * https://www.unioviedo.es/compnum/labs/PYTHON/Interpolation.html
#
# * https://www.johndcook.com/blog/2017/11/06/chebyshev-interpolation/
#
# * https://numpy.org/devdocs/reference/routines.polynomials.html
| labs/approximation/notebook.nbconvert.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Code Notebook 3: Feature Selection and Engineering #
# In this notebook, I will be engineering categorical features into a numeric scale when possible. I will examine correlations between features identified in Part 2 as promising predictors and use this information, combined with the observations from Part 2, to choose the best combinations of features for regression. Any final pre-processing of those features, such as dropping or filling NaN values appropriately, dropping extreme outliers, or combining features in feature interactions, will complete the feature engineering process in this notebook.
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import graphtools as gt
# -
train = pd.read_csv('../datasets/train_1.csv', keep_default_na=False, na_values='null')
test = pd.read_csv('../datasets/test_1.csv', keep_default_na=False, na_values='null')
# ## VIII. Converting Hierarchical Categorical Data to Numeric Scale ##
# In this section, all categorical variables that have a clear heirarchy will be converted to a numeric scale.
train['exter_qual'].value_counts()
train['exter_cond'].value_counts()
train['bsmt_qual'].value_counts()
train['bsmt_cond'].value_counts()
train['bsmt_exposure'].value_counts()
train['heating_qc'].value_counts()
train['central_air'].value_counts()
train['kitchen_qual'].value_counts()
train['functional'].value_counts()
train['fireplace_qu'].value_counts()
train['bsmtfin_type_1'].value_counts()
train['bsmtfin_type_2'].value_counts()
train['electrical'].value_counts()
train['garage_finish'].value_counts()
train['garage_qual'].value_counts()
train['garage_cond'].value_counts()
train['paved_drive'].value_counts()
train['pool_qc'].value_counts()
# +
# Assigning categorical variables to be converted as appropriate.
rating_to_scaled = ['exter_qual', 'exter_cond', 'bsmt_qual', 'bsmt_cond', 'heating_qc', 'kitchen_qual', 'fireplace_qu',
'garage_qual', 'garage_cond', 'pool_qc']
exposure_to_scaled = ['bsmt_exposure']
yes_no_to_binary = ['central_air']
functional_to_scaled = ['functional']
finish_to_scaled = ['bsmtfin_type_1', 'bsmtfin_type_2']
elec_to_scaled = ['electrical']
garfin_to_scaled = ['garage_finish']
paved_to_scaled = ['paved_drive']
# -
def rating_to_scale(df, rating_cols, exposure_cols, yes_no_cols, fun_cols, fin_cols, elec_cols, garfin_cols, paved_cols):
# Dictionaries for converting from categorical rating scales to numerical scales.
rating_to_scale = {
'NA' : 0,
'Po' : 1,
'Fa' : 2,
'TA' : 3,
'Gd' : 4,
'Ex' : 5,
'' : np.nan
}
exposure_to_scale = {
'NA' : np.nan,
'No' : 0,
'Mn' : 1,
'Av' : 2,
'Gd' : 3
}
yes_no_scale = {
'Y' : 1,
'N' : 0
}
fun_scale = {
'Sal' : 0,
'Sev' : 1,
'Maj2' : 2,
'Maj1' : 3,
'Mod' : 4,
'Min2' : 5,
'Min1' : 6,
'Typ' : 7
}
fin_scale = {
'' : np.nan,
'NA' : 0,
'Unf' : 1,
'LwQ' : 2,
'Rec' : 3,
'BLQ' : 4,
'ALQ' : 5,
'GLQ' : 6
}
elec_scale = {
'Mix' : 1,
'FuseP' : 2,
'FuseF' : 3,
'FuseA' : 4,
'SBrkr' : 5
}
garfin_scale = {
'' : np.nan,
'NA' : 0,
'Unf' : 1,
'RFn' : 2,
'Fin' : 3
}
paved_scale = {
'N' : 0,
'P' : 1,
'Y' : 2
}
# These loops apply the above dictionaries to convert each categorical column as previously assigned.
for col in rating_cols:
df[col] = df[col].map(rating_to_scale)
for col in exposure_cols:
df[col] = df[col].map(exposure_to_scale)
for col in yes_no_cols:
df[col] = df[col].map(yes_no_scale)
for col in fun_cols:
df[col] = df[col].map(fun_scale)
for col in fin_cols:
df[col] = df[col].map(fin_scale)
for col in elec_cols:
df[col] = df[col].map(elec_scale)
for col in garfin_cols:
df[col] = df[col].map(garfin_scale)
for col in paved_cols:
df[col] = df[col].map(paved_scale)
return df
# +
# Apply the maps designed above to the train and test splits.
train = rating_to_scale(train, rating_to_scaled, exposure_to_scaled, yes_no_to_binary, functional_to_scaled,
finish_to_scaled, elec_to_scaled, garfin_to_scaled, paved_to_scaled)
test = rating_to_scale(test, rating_to_scaled, exposure_to_scaled, yes_no_to_binary, functional_to_scaled,
finish_to_scaled, elec_to_scaled, garfin_to_scaled, paved_to_scaled)
# -
# ## IX. Correlated Features and Feature Interactions ##
corr = train.corr()
plt.figure(figsize=(2,20))
sns.heatmap(data=corr[['saleprice']].sort_values(by='saleprice', ascending=False), annot=True, vmin=-1, vmax=1, cmap='viridis', linewidths=1);
# Recreating the heatmap now that more features are encoded numerically.
bsmt_features = [col for col in train.columns if 'bsmt' in col]
# +
corr = train[bsmt_features].corr()
plt.figure(figsize=(20,10))
sns.heatmap(corr, annot=True, vmin=-1, vmax=1, cmap='viridis', linewidths=1);
# -
# There are a very large number of basement features, but the most promising tend to be correlated with each other.
gt.quick_scatter(train['total_bsmt_sf']*train['bsmt_full_bath'],
train['saleprice'])
garage_features = [col for col in train.columns if 'gar' in col]
corr = train[garage_features].corr()
plt.figure(figsize=(20,10))
sns.heatmap(corr, annot=True, vmin=-1, vmax=1, cmap='viridis', linewidths=1);
# Nearly all garage features are significantly correlated with each other. I should only use a few, ideally in a feature interaction.
gt.quick_scatter(train['garage_yr_blt'].fillna(0)*train['garage_area'],
train['saleprice'])
# This looks like an excellent combination of garage features. It is also a nice solution of how to handle the garage_yr_blt feature when there is no garage. It can now be encoded as 0.
# See the documentation for .multi_bar_target_mean() in graphtools.py for references to resources I used for this graph.
price_by_neighborhood = train['saleprice'].groupby(train['neighborhood']).mean()
plt.figure(figsize=(20,5))
plot = plt.bar(price_by_neighborhood.index, price_by_neighborhood)
plt.xlabel('Neighborhood', fontsize='x-large')
plt.ylabel('Saleprice', fontsize='x-large')
plt.title(f'Mean Saleprice by Neighborhood', fontsize='x-large')
plt.xticks(rotation=70)
plt.bar_label(plot, train['neighborhood'].groupby(train['neighborhood']).size());
# Select neighborhoods with average saleprice at least one standard deviation above or below the mean average saleprice.
high_neighborhood_cutoff = price_by_neighborhood.mean() + price_by_neighborhood.std()
low_neighborhood_cutoff = price_by_neighborhood.mean() - price_by_neighborhood.std()
expensive_neighborhoods = price_by_neighborhood[price_by_neighborhood > high_neighborhood_cutoff]
cheap_neighborhoods = price_by_neighborhood[price_by_neighborhood < low_neighborhood_cutoff]
print(expensive_neighborhoods)
print(cheap_neighborhoods)
# Neighborhood looks like a very strong detmining factor for price with a nice distribution of houses across neighborhoods. I previously tried OneHotEncoding this feature, but it was detrimental to my model. I am now going to create 3 bins for neighborhoods with low, tyical, or high mean saleprices.
# A list of the most promising overall size-related features from EDA.
size_features = ['1st_flr_sf', 'gr_liv_area', 'lot_area', 'totrms_abvgrd', 'full_bath', 'half_bath']
corr = train[size_features].corr()
plt.figure(figsize=(10,5))
sns.heatmap(corr, annot=True, vmin=-1, vmax=1, cmap='viridis', linewidths=1);
# gr_liv_area, half_bath, and lot_area seems like a good combination.
# A list of the most promising age-related features from EDA.
size_features = ['year_built', 'year_remod/add']
corr = train[size_features].corr()
plt.figure(figsize=(10,5))
sns.heatmap(corr, annot=True, vmin=-1, vmax=1, cmap='viridis', linewidths=1);
# Probably best to stick with one of these.
# A licolumnshe most promising quality-related features from EDA.
quality_features = ['overall_qual', 'exter_qual', 'kitchen_qual', 'fireplace_qu', 'heating_qc']
corr = train[quality_features].corr()
plt.figure(figsize=(10,5))
sns.heatmap(corr, annot=True, vmin=-1, vmax=1, cmap='viridis', linewidths=1);
# Many of these are correalated. I'll try overall_qual, fireplace_qu, and heating_qc
keep_features = ['saleprice', 'id', # REQUIRED
'total_bsmt_sf', 'bsmt_full_bath', # BASEMENT
'garage_yr_blt', 'garage_area', # GARAGE
'half_bath', 'gr_liv_area', 'lot_area', # OVERALL SIZE
'neighborhood', # LOCATION
'overall_qual', 'fireplace_qu', 'heating_qc', # QUALITY
'wood_deck_sf', 'open_porch_sf', 'enclosed_porch',
'3ssn_porch', 'screen_porch', # PORCH
'mas_vnr_area', # MASONRY VENEER
'year_built' # AGE
]
# +
# List of features in keep_features that need to be dummified.
# dummy_features = ['mas_vnr_type']
# -
# Every time I OneHotEncoded something it went poorly, so I finally didn't use anything I could convert to a numerical scale.
# ## X. Removing Outliers ##
# This custom function graphs scatterplots of all numeric columns in a dataframe vs. one selected column.
gt.multi_scatter(train[keep_features], 'saleprice', 3, 20)
# Collecting a list of indices of rows with extreme outliers based on the scatterplots above. This is also where I will drop any rows with NaN values that I can't otherwise account for.
outlier_rows = list(train.index[ (train['lot_area'] > 80_000) |
(train['1st_flr_sf'] > 3000) |
(train['gr_liv_area'] > 4000) |
(train['bsmtfin_sf_1'] > 3000) |
(train['total_bsmt_sf'] > 4000) |
(train['bsmt_qual'].isna()) |
(train['garage_yr_blt'] > 2010)]
)
outlier_rows
# + tags=[]
# # List to hold indices of rows with outliers.
# outlier_rows = []
# # Iterate through all rows being sent to the regression except saleprice.
# for col in keep_features[1:]:
# # Ignore the row if it isn't numeric.
# if (train[col].dtype == int) | (train[col].dtype == float):
# # Identify positive outliers and add them to the list if they are not already in it.
# pos_outliers = list(train.index[train[col] > train[col].mean() + 3 * train[col].std()])
# new_pos_outliers = [row for row in pos_outliers if row not in outlier_rows]
# # Identify negative outliers and add them to the list if they are not already in it.
# neg_outliers = list(train.index[train[col] < train[col].mean() - 3 * train[col].std()])
# new_neg_outliers = [row for row in neg_outliers if row not in outlier_rows]
# # Add the new outliers to outlier_rows
# outlier_rows += new_pos_outliers
# outlier_rows += new_neg_outliers
# # check how many rows are being dropped.
# len(outlier_rows)
# +
# Reminder of the shape of the dataframe being sent to regression, for comparison with number of outlier rows being dropped.
# train[keep_features].shape
# -
# The commented out code above made my results worse. It seems that removing all any data more than 3 standard deviations from the mean before sending it to the model is too agressive.
# + [markdown] tags=[]
# ## XI. Feature Engineering and Outlier Pipelines ##
# -
def pipe_select(df, keep):
return df[keep]
train = pipe_select(train, keep_features)
# saleprice will always be the first feature in keep_features. It must be excluded for the test data because it isn't provided.
test = pipe_select(test, keep_features[1:])
def group_neighborhoods(df):
neighborhood_bins = {}
for neighborhood in df['neighborhood'].unique():
if neighborhood in expensive_neighborhoods:
neighborhood_bins[neighborhood] = 3
elif neighborhood in cheap_neighborhoods:
neighborhood_bins[neighborhood] = 1
else:
neighborhood_bins[neighborhood] = 2
df['neighborhood'] = df['neighborhood'].map(neighborhood_bins)
return df
train = group_neighborhoods(train)
test = group_neighborhoods(test)
# The commented out cells below are from my attempt to OneHotEncode categorical features. This was not good for my model, and I instead chose to break neighborhood into three bins and avoid other features that would have required OneHotEncoding.
# +
# def pipe_engineer(df_train, df_test, dummify):
# df_train = pd.get_dummies(df_train, columns=dummify)
# df_test = pd.get_dummies(df_test, columns=dummify)
# for col in df_train.columns:
# if (col not in df_test.columns) & (col != 'saleprice'):
# df_test[col] = 0
# return df_train, df_test
# +
# train, test = pipe_engineer(train, test, dummy_features)
# -
# Year features are a special case. We might be inclined to convert them to age, but this would be a problem with feature interactions. For example, if I multiple garage age by garage size, then they work against each other. A small age and large size would result in a small total, rather than large. Due to this, I will use "newness," which will be the number of years after 1890 than the garage was built. This was chosen because the oldest house provided was built in 1895, so this should avoid any 0's. If a house has no garage, then year built can be 0, since it will multiply a 0 size in a feature interaction and have no effect.
train['garage_yr_blt'].min()
def pipe_interact(df):
# Adjust garage_yr_blt to be years since 1890 and fill NaN with 0.
df['garage_yr_blt'] = df['garage_yr_blt'] - 1890
df['garage_yr_blt'].fillna(0, inplace=True)
# Also adjust year_built to have the same scale.
df['year_built'] = df['year_built'] - 1890
df['mas_vnr_area'].fillna(0, inplace=True)
# Adding deck/porch features into one column that will represent total deck area.
df['porch_space'] = df['wood_deck_sf'] + df['open_porch_sf'] + df['enclosed_porch'] + df['3ssn_porch'] + df['screen_porch']
df.drop(columns=['wood_deck_sf', 'open_porch_sf', 'enclosed_porch', '3ssn_porch', 'screen_porch'], inplace=True)
# Feature interactions.
df['garage_total'] = df['garage_area'] * df['garage_yr_blt']
df.drop(columns=['garage_area', 'garage_yr_blt'], inplace=True)
df['bsmt_total'] = df['total_bsmt_sf'] * df['bsmt_full_bath']
df.drop(columns=['total_bsmt_sf', 'bsmt_full_bath'], inplace=True)
return df
train = pipe_interact(train)
test = pipe_interact(test)
def pipe_outlier(df, outlier_list):
df.drop(outlier_list, inplace=True)
return df
train = pipe_outlier(train, outlier_rows)
train.to_csv('../datasets/train_2.csv', index=False, na_rep='null')
test.to_csv('../datasets/test_2.csv', index=False, na_rep='null')
train.shape
test.shape
# All features that are being passed to the model have been properly encoded. NaN values and outliers have been addressed, and interactions between related features have been established. All features have been selected to avoid excessively strong correlations, so these features are representative of broader categories. For example, total size above ground is a good stand-in for other size-related features like numbers of bathrooms. This completes the process of preparing features for modelling.
#
# Below are the features included in the final version of the regression model:
#
# |Feature|Type|Description|
# |---|---|---|
# |saleprice|int|Sale price of the house in dollar|
# |id|int|House identifier necessary for Kaggle submission, exculded from model|
# |half_bath|int|Number of half-bathrooms above grade|
# |gr_liv_area|int|Total above ground living area|
# |lot_area|int|Lot size in sq feet|
# |neighborhood|int|1 if neighborhood is 1 std below the mean for average sale price, 3 if 1 std above the mean, 2 otherwise|
# |overall_qual|int|Overall quality rating of the house on an increasing scale|
# |fireplace_qu|float|Fireplace quality rating on increasing numeric scale with 0 being no fireplace|
# |heating_qc|float|Heating quality on an increasing numeric scale|
# |mas_vnr_area|float|Total masonry veneer area|
# |year_built|int|Year built minus 1890 to allow for a more natural scale when using scaling algorithms|
# |porch_space|int|Sum of the area of all porch types in sq feet|
# |garage_total|float|Product of the original features garage area and year built, with year built subtracted by 1890 and 0 if no garage|
# |bsmt_total|float|Product of the original features total basement surface area (sq ft) and basement full bathrooms|
#
#
| code/part3-feature-selection-engineering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Entrada de datos por teclado
# En python es posible introducir datos por teclado, ya sea para introducir parámetros en nuestro flujo de trabajo, o bien para evaluar "funciones" simple. Veamos esto a través de ejemplos.
#
# #### Ejemplo 1 (Caso de uso).
# A un TAAS trabajando en Arauco Forestal le piden construir una función que calcule el área de una zona de sus terrenos de cultivo, para marcar las zonas que se talarán para la extracción de celulosa. Como datos de entrada ofrecen el ancho y largo. La herramienta quieren escribirla en Python para disponibilizarla (a futuro) como una cloud function. ¿Cómo hacer esto?
# +
# usamos la instrucción input para
# tomar los valores de largo y ancho.
ancho = float(input("Ancho del terreno [m]: "))
largo = float(input("Largo del terreno [m]: "))
print('El área del terreno evaluado es ',ancho*largo, '[m^2]')
# -
# #### Problema 1.
# Un emprendedor que quiere construir una app para ofrecer un servicio de encomiendas desea poder calcular cuanto debe cobrar cada uno de los usuarios de su app, considerando aspectos como ancho, largo, alto y peso.
#
# Este deduce la siguiente fórmula como calculadora de costos:
#
# Costo=(largo x alto x ancho + 2 x peso) x 1000
#
# Calcule el costo de envío si:
# - largo = 1
# - ancho = 2
# - alto = 3
# - peso = 4
# # Funciones
# Una función puede imaginarse como una maquina en la que introducimos objetos, esta los procesa, y devuelve otro objeto. Imaginen cualquier maquina ...
# Veamos una de las maquinas/funciones mas simples ... toma dos números y los suma.
# sintaxis
def suma1(numero1, numero2):
return numero1+numero2
# hay otra forma de definir funciones
suma2=lambda num1,num2: num1+num2
# +
# Buenas prácticas:
# Sintaxis
def nombre_de_la_funcion(argumento1,argumento2):
'''
Documentación de la Función
-------------------------------------
Aquí explicamos que hace la función.
Parámetros
----------
argumento1: explicamos el significado de cada uno de los argumentos
de la función, así como sus restricciones.
argumento2: idem.
Resultado/Retorno
-------------------
Lo que la función devuelve como resultado del proceso que realiza.
Ejemplo
--------
>>> nombre_de_la_funcion('hola','amigo')
hola amigo 123
Referencias
----------
.. [1] Alvarez-Socorro, A.J. Buenas prácticas a la hora de escribir
funciones en Python, Laboratorio de I+D+I Zenta, 2019.
'''
return str(argumento1)+' '+str(argumento2)+' 123'
nombre_de_la_funcion('hola','amigo')
# +
# Construya funciones para los ejemplo 1 y para el problema 1.
# +
def costo_del_envio(ancho,alto,largo,peso):
'''
Documentación:
---------------
Esta función calcula el costo del envio
----
ancho: float
alto: float
largo: float
peso: float
salida
------
precio: float
'''
precio=(largo * alto * ancho + 2 * peso) * 1000
return precio
print(costo_del_envio(1,2,3,4))
# -
# Las funciones pueden ser tan generales y complejas como el caso lo requiera. Inicialmente, no tienen dependencia explicita con el tipo de dato, sin embargo, si efectuamos alguna operación no permitida, esta nos dará un error.
# +
# Definimos una función que calcule el promedio de una
# serie de números (i.e. en una lista)
L1=[2,3,46,23,51,15]
# 16.666
def promedio1(L):
acum=0
for i in L:
acum=acum+i
pro=acum/len(L)
return pro
def promedio2(L):
return sum(L)/len(L)
promedio3=lambda L: sum(L)/len(L)
# -
promedio3(L1)
# +
# construyamos una función que calcule la desviación estándar
# la fórmula de desviación estándar es
# raiz(sum(x-mu)^2/N)
def desviacion_estandar(L1):
mu=promedio2(L1)
s=0
for i in L1:
s=s+(i-mu)**2
var=((1/len(L1))*s)**0.5
return var
desviacion_estandar(L1)
# +
L1=[2,3,46,23,51,15]
def desviacion_estandar2(L1):
mu=promedio2(L1)
s=0
LAUX=[(i-mu)**2 for i in L1]
s=sum(LAUX)
var=((1/len(L1))*s)**0.5
return var
desviacion_estandar2(L1)
# -
# Comparamos con numpy
import numpy as np
np.mean(L1)
np.std(L1)
# +
# considere el siguiente código
anchura = int(input("Anchura del rectángulo: "))
altura = int(input("Altura del rectángulo: "))
for i in range(altura):
for j in range(anchura):
print("* ", end="")
print()
# construya una función que use anchura y altura como argumentos
# y grafique un rectangulo de 10 x 6
# -
# # problemas de funciones
#
# 1. Escriba una función en Python que encuentre el máximo entre tres números.
#
#
# +
def maxi(a,b,c):
return max(a,b,c)
print(maxi(-1,10,2))
def suma_l(L):
acum=0
for i in L:
acum=acum+i
return acum
def suma_l(L):
acum=1
for i in L:
acum=acum*i
return acum
# -
#
# 2. Usando un for, escriba una función que sume todo los numeros de una lista.
# Lista Ejemplo : [9, 1, 3, 1, 6]
# Output : 20
#
# 3. Escriba una función en Python que multiplique todos los números de una lista.
# Lista Ejemplo: (1, 2, 3,5, -1)
# Output : -30
#
# 4. Escriba una función en Python que escriba una cadena al revés.
# Cadena de ejemplo: "ZentaGroup123"
# Output : "321puorGatneZ"
#
# 5. Escriba una función que calcule el factorial de un número.
# $$n!=n*(n-1)*(n-2)\ldots 2* 1$$
#
# 6. Escriba una función que verifique si un número está en una lista. (For/Else :o)
#
# 7. Escriba una función que cuente el número de mayúsculas y minusculas en una cadena de caracteres.
# Cadena de ejemplo : 'ZentaGroupNuNoA'
# Output :
# No. de caracteres en mayúscula: 5
# No. de caracteres en minúscula: 11
#
# 8. Escriba una función que tome una lista y devuelva una nueva lista sin que ninguno de sus elementos esté repetido.
# Lista de prueba : [1,2,3,3,3,3,4,5]
# Lista resultante : [1, 2, 3, 4, 5]
#
# 9. Escriba una función que chequee si una palabra o frase es palindroma y regrese True o False según sea el caso.
# Nota: Una palabra o frase es palindroma si se lee de la misma forma hacia delante y hacia atrás. Por ejemplo, "yo soy", "somos o no somos".
#
# 10. Escriba una función que tenga como argumento una secuencia de palabras separadas por guiones y su salida sea la misma secuencia pero ordenada alfabéticamente.
# Ejemplo : verde-rojo-amarillo-negro-blanco
# Resultado : amarillo-blanco-...
def inversa(A):
CR=''
for i in range(len(A)-1,-1,-1):
CR=CR+A[i]
return CR
print(inversa('ZentaGroup123'))
def factorial(n):
if n==1: return 1
else: return n*factorial(n-1)
print(factorial(10))
| Clase_3.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// # magic commands
// #!lsmagic
// #!about
var x = 10;
var q = 0;
// #!who
// #!whos
// # display
Console.WriteLine("Hello world");
Console.WriteLine(h2("hello world"));
display("hello world");
display(h2("hello world"))
h2[style: "color: red"]("hello world")
h2[style: "color: red"]("hello world")
// +
var pocketViewTagMethods = typeof(PocketViewTags).GetProperties().Select(m => m.Name);
string.Join(",", pocketViewTagMethods)
// -
select(option("first"), option("second"))
// # #load
#load "C:\Users\dcost\source\repos\JupyterNotebooks\start.csx"
Add(100, 200)
// # object formatter
class Student { public string FirstName { get; set; } public string LastName { get; set; }}
var student = new Student { FirstName = "John", LastName = "Smith" };
display(student)
Formatter.Register<Student>((student, writer) =>
{
writer.Write(b[style: "color: green"]($"First name: {student.FirstName}, last name: {student.LastName}"));
}, "text/html");
display(student)
// +
//Formatter.ResetToDefault(); // this resets all object formatters (inclusing charts objects!), not recommended
// -
// # charts
using XPlot.Plotly;
var numbers = Enumerable.Range(1, 100).ToList();
numbers
Random rnd = new Random();
var randomNumbersA = numbers.Select(_ => rnd.Next(100));
var randomNumbersB = numbers.Select(_ => rnd.Next(50));
randomNumbersA.ToList()
// #### object formatter for list of ints
// +
static string BuildHideRowsScript(long uniqueId)
{
var script = $"var allRows = document.querySelectorAll('#table_{uniqueId} tbody tr:nth-child(n)'); ";
script += "for (let i = 0; i < allRows.length; i++) { allRows[i].style.display='none'; } ";
return script;
}
static string BuildPageScript(long uniqueId, int size)
{
var script = $"var page = parseInt(document.querySelector('#page_{uniqueId}').innerHTML) - 1; ";
script += $"var pageRows = document.querySelectorAll(`#table_{uniqueId} tbody tr:nth-child(n + ${{page * {size} + 1 }})`); ";
script += $"for (let j = 0; j < {size}; j++) {{ pageRows[j].style.display='table-row'; }} ";
return script;
}
static string GotoPageIndex(long uniqueId, long page)
{
var script = $"document.querySelector('#page_{uniqueId}').innerHTML = {page + 1}; ";
return script;
}
static string UpdatePageIndex(long uniqueId, int step, long maxPage)
{
var script = $"var page = parseInt(document.querySelector('#page_{uniqueId}').innerHTML) - 1; ";
script += $"page = parseInt(page) + parseInt({step}); ";
script += $"page = page < 0 ? 0 : page; ";
script += $"page = page > {maxPage} ? {maxPage} : page; ";
script += $"document.querySelector('#page_{uniqueId}').innerHTML = page + 1; ";
return script;
}
// +
using Microsoft.AspNetCore.Html;
Formatter.Register<List<int>>((ints, writer) =>
{
const int MAX = 10000;
const int SIZE = 10;
var uniqueId = DateTime.Now.Ticks;
if (ints.Count > SIZE)
{
var maxMessage = ints.Count > MAX ? $" (showing a max of {MAX} rows)" : string.Empty;
var title = h3[style: "text-align: center;"]($"List of ints - {ints.Count} rows {maxMessage}");
//var header = new List<IHtmlContent>
//{
// th(i("index"))
//};
//header.AddRange(df.Columns.Select(c => (IHtmlContent)th(c.Name)));
// table body
var maxRows = Math.Min(MAX, ints.Count);
var rows = new List<List<IHtmlContent>>();
for (var index = 0; index < Math.Min(MAX, ints.Count); index++)
{
var cells = new List<IHtmlContent>
{
td(i((index)))
};
//foreach (var obj in ints[index])
//{
// cells.Add(td(obj));
//}
cells.Add(td(ints[index]));
rows.Add(cells);
}
//navigator
var footer = new List<IHtmlContent>();
BuildHideRowsScript(uniqueId);
if (ints.Count > SIZE)
{
var paginateScriptFirst = BuildHideRowsScript(uniqueId) + GotoPageIndex(uniqueId, 0) + BuildPageScript(uniqueId, SIZE);
footer.Add(button[style: "margin: 2px;", onclick: paginateScriptFirst]("⏮"));
var paginateScriptPrevTen = BuildHideRowsScript(uniqueId) + UpdatePageIndex(uniqueId, -10, (maxRows - 1) / SIZE) + BuildPageScript(uniqueId, SIZE);
footer.Add(button[style: "margin: 2px;", onclick: paginateScriptPrevTen]("⏪"));
var paginateScriptPrev = BuildHideRowsScript(uniqueId) + UpdatePageIndex(uniqueId, -1, (maxRows - 1) / SIZE) + BuildPageScript(uniqueId, SIZE);
footer.Add(button[style: "margin: 2px;", onclick: paginateScriptPrev]("◀️"));
footer.Add(b[style: "margin: 2px;"]("Page"));
footer.Add(b[id: $"page_{uniqueId}", style: "margin: 2px;"]("1"));
var paginateScriptNext = BuildHideRowsScript(uniqueId) + UpdatePageIndex(uniqueId, 1, (maxRows - 1) / SIZE) + BuildPageScript(uniqueId, SIZE);
footer.Add(button[style: "margin: 2px;", onclick: paginateScriptNext]("▶️"));
var paginateScriptNextTen = BuildHideRowsScript(uniqueId) + UpdatePageIndex(uniqueId, 10, (maxRows - 1) / SIZE) + BuildPageScript(uniqueId, SIZE);
footer.Add(button[style: "margin: 2px;", onclick: paginateScriptNextTen]("⏩"));
var paginateScriptLast = BuildHideRowsScript(uniqueId) + GotoPageIndex(uniqueId, (maxRows - 1) / SIZE) + BuildPageScript(uniqueId, SIZE);
footer.Add(button[style: "margin: 2px;", onclick: paginateScriptLast]("⏭️"));
}
else
{
BuildHideRowsScript(uniqueId);
footer.Add(b[style: "margin: 2px;"]("Page"));
footer.Add(b[id: $"page_{uniqueId}", style: "margin: 2px;"]("0"));
}
//table
var t = table[id: $"table_{uniqueId}"](
caption(title),
thead(tr(header)),
tbody(rows.Select(r => tr[style: "display: none"](r))),
tfoot(tr(td[style: "text-align: center;"](footer)))
);
writer.Write(t);
}
else
{
var rows = new List<List<IHtmlContent>>();
for (var index = 0; index < ints.Count; index++)
{
var cells = new List<IHtmlContent>
{
td(i((index)))
};
//foreach (var obj in ints[index])
//{
//cells.Add(td(obj));
//}
cells.Add(td(ints[index]));
rows.Add(cells);
}
//table
var t = table[id: $"table_{uniqueId}"](
thead(tr(header)),
tbody(rows.Select(r => tr(r)))
);
writer.Write(t);
}
//show first page
writer.Write($"<script>{BuildPageScript(uniqueId, SIZE)}</script>");
}, "text/html");
// -
randomNumbersA.ToList()
// #### box plot segmentation
// +
var segmentationDiagram = Chart.Plot(new[] {
new Graph.Box { y = randomNumbersA, name = "First set of numbers" },
new Graph.Box { y = randomNumbersB, name = "Second set of numbers" }
});
var layout = new Layout.Layout()
{
title = "Box plot segmentation"
};
segmentationDiagram.WithLayout(layout);
display(segmentationDiagram);
// -
| SmartFireAlarm/Jupyter/Jupyter Start.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas_datareader as pdr
import datetime
t = pdr.get_data_yahoo('000333.SZ', start=datetime.datetime(2016,10,1), end=datetime.datetime.now())
print(t)
t.head()
t.describe()
import pandas as pd
type(t)
t.index
monthly_t = t.resample('M').mean()
print monthly_t
import matplotlib.pyplot as plt
import matplotlib.pyplot as plt
t['Close'].plot(grid=True)
plt.show()
import numpy as np
daily_close = t[['Adj Close']]
daily_pct_change = daily_close.pct_change()
daily_pct_change.fillna(0, inplace=True)
print(daily_pct_change)
daily_log_returns = np.log(daily_close.pct_change()+1)
print(daily_log_returns)
daily_pct_change.hist(bins=50)
plt.show()
cum_daily_return = (1 + daily_pct_change).cumprod()
print(cum_daily_return)
cum_daily_return.plot(figsize=(12,8))
plt.show()
cum_monthly_return = cum_daily_return.resample("M").mean()
print(cum_monthly_return)
cum_monthly_return.plot()
daily_volume = t[['Volume']]
daily_volume.plot(kind='bar')
print(daily_volume)
t.head()
daily_volume = t[['Volume']]
daily_volume.head()
daily_volume.plot(kind='bar')
plt.show()
monthly_volume = daily_volume.resample('W').mean()
monthly_volume.head()
monthly_volume.plot(kind='bar')
plt.show()
monthly_volume = daily_volume.resample('BM').mean()
monthly_volume.plot(kind='bar')
plt.show()
monthly_volume.hist()
plt.show()
monthly_volume.hist(bins=20, sharex=True, figsize=(12,8))
plt.show()
daily_volume.hist(bins=20, sharex=True, figsize=(12,8))
plt.show()
daily_volume.head()
moving_avg_vol = daily_volume.rolling(window=10).mean()
moving_avg_vol.head()
print(moving_avg_vol)
moving_avg_vol.plot(figsize=(12,8))
plt.show()
daily_close_vol = t[['Adj Close', 'Volume']]
daily_close_vol.head()
min_periods = 75
vol = daily_pct_change.rolling(min_periods).std() * np.sqrt(min_periods)
vol.plot(figsize=(10, 8))
plt.show()
t = pdr.get_data_yahoo('000333.SZ', start=datetime.datetime(2015,10,1), end=datetime.datetime.now())
short_window = 40
long_window = 100
signals = pd.DataFrame(index=t.index)
signals['signal'] = 0.0
signals['short_mavg'] = t['Close'].rolling(window=short_window, min_periods=1, center=False).mean()
signals['long_mavg'] = t['Close'].rolling(window=long_window, min_periods=1, center=False).mean()
signals['signal'][short_window:] = np.where(signals['short_mavg'][short_window:]
> signals['long_mavg'][short_window:], 1.0, 0.0)
signals.head(50)
signals['positions'] = signals['signal'].diff()
print(signals['positions'][:50])
fig, ax1 = None, None
fig = plt.figure(figsize=(15,8))
ax1 = fig.add_subplot(111, ylabel='Price in $')
t['Close'].plot(ax=ax1, color='r', lw=2.)
signals[['short_mavg', 'long_mavg']].plot(ax=ax1, lw=2.)
ax1.plot(signals.loc[signals.positions == 1.0].index,
signals.short_mavg[signals.positions == 1.0],
'^', markersize=10, color='m')
ax1.plot(signals.loc[signals.positions == -1.0].index,
signals.short_mavg[signals.positions == -1.0],
'v', markersize=10, color='k')
plt.show()
print(signals.short_mavg[signals.positions == 1.0])
print(signals.loc[signals.positions == 1.0].index)
| demo/stock_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Helper Methods
# +
from IPython.core.magic import register_cell_magic
@register_cell_magic
def write_and_run(line, cell):
"""Write command to file before executing on notebook"""
args = line.split()
append = ''
if '-a' in args:
append = args.pop()
for arg in args:
file = arg
mode = 'w' if append != '-a' else 'a'
with open(file, mode) as f:
f.write(f"{cell}\n")
get_ipython().run_cell(cell)
# -
# # Dependencies Installation
# !pip install -r requirements.txt
# ## Connect to our DB server
# First thing we should do is to connect to our DB(test)
#
# **PS**: Do not do this in production
SCRIPT_FILENAME = 'scripts/simple_etl.py'
# %store SCRIPT_FILENAME
# +
# %%write_and_run $SCRIPT_FILENAME
import os
import pandas as pd
from dotenv import dotenv_values
from sqlalchemy import create_engine, inspect
# -
# Read environmental variables defined in the `.env` file, if not available read from environments
# +
# %%write_and_run $SCRIPT_FILENAME -a
CONFIG = dotenv_values('.env')
if not CONFIG:
CONFIG = os.environ
connection_uri = "postgresql+psycopg2://{}:{}@{}:{}".format(
CONFIG["POSTGRES_USER"],
CONFIG["POSTGRES_PASSWORD"],
CONFIG['POSTGRES_HOST'],
CONFIG["POSTGRES_PORT"],
)
# -
# Create a connection to the DB.
# +
# %%write_and_run $SCRIPT_FILENAME -a
engine = create_engine(connection_uri, pool_pre_ping=True)
engine.connect()
# -
# # Extract the data from the hosting service
# %%write_and_run $SCRIPT_FILENAME -a
# Extract
dataset = "https://gist.githubusercontent.com/mmphego/5b6fc4d6dc3c8fba4fce9d994a2fe16b/raw/ab5df0e76812e13df5b31e466a5fb787fac0599a/wine_quality.csv"
# Load in the 'Wine Quality' dataset and creare a dataframe
# +
# %%write_and_run $SCRIPT_FILENAME -a
df = pd.read_csv(dataset)
# -
# Always good to make sure your data look correct
df.head()
# Simple table summary containing basic statistical metrics (count, mean, std, min, max, and percentiles)
df.describe()
# Notice that index 12 contains an object (string), our future ML model will not understand this.
# Therefore we will need to transform it to numeric
df.info()
# ## Load the raw data into a production system
#
# Once we have our data in a dataframe, we can then save it into our DB for future use
# %%write_and_run $SCRIPT_FILENAME -a
# load to DB
table_name = 'wine_quality_raw_dataset'
df.to_sql(table_name, engine, if_exists='replace')
# Check if table was created!
f"{table_name!r} exists!" if table_name in inspect(engine).get_table_names() else f"{table_name} does not exist!"
# # Transform it into a usable format
# Now we impute the color from `str` to `int`
# %%write_and_run $SCRIPT_FILENAME -a
# transformation
df_transform = df.copy()
winecolor_encoded = pd.get_dummies(df_transform['winecolor'], prefix='winecolor')
df_transform[winecolor_encoded.columns.to_list()] = winecolor_encoded
df_transform.drop('winecolor', axis=1, inplace=True)
# apply normalization techniques
for column in df_transform.columns:
df_transform[column] = (df_transform[column] - df_transform[column].mean()) / df_transform[column].std()
df_transform.describe()
df_transform.head()
# # Load transformed data into a production system
# Once our data transformation is complete, we can save our dataframe in an SQL table
# %%write_and_run $SCRIPT_FILENAME -a
# load
table_name = table_name.replace('raw', 'clean')
# %%write_and_run $SCRIPT_FILENAME -a
df.to_sql(table_name, engine, if_exists='replace')
# Check if table was created!
f"{table_name!r} exists!" if table_name in inspect(engine).get_table_names() else f"{table_name} does not exist!"
# # Read table from SQL
# for sanity!
# ## Raw Dataset
pd.read_sql("SELECT * FROM wine_quality_raw_dataset", engine)
# ## Cleaned Dataset
pd.read_sql("SELECT * FROM wine_quality_clean_dataset", engine)
engine
# # Done!
# +
# %%writefile dags/simple_etl_dag.py
import os
from functools import wraps
import pandas as pd
from airflow.models import DAG
from airflow.utils.dates import days_ago
from airflow.operators.python import PythonOperator
from dotenv import dotenv_values
from sqlalchemy import create_engine, inspect
args = {"owner": "me myself and I", "start_date": days_ago(1)}
dag = DAG(dag_id="simple_etl_dag", default_args=args, schedule_interval=None)
DATASET_URL = "https://gist.githubusercontent.com/mmphego/5b6fc4d6dc3c8fba4fce9d994a2fe16b/raw/ab5df0e76812e13df5b31e466a5fb787fac0599a/wine_quality.csv"
CONFIG = dotenv_values(".env")
if not CONFIG:
CONFIG = os.environ
def logger(fn):
from datetime import datetime, timezone
@wraps(fn)
def inner(*args, **kwargs):
called_at = datetime.now(timezone.utc)
print(f">>> Running {fn.__name__!r} function. Logged at {called_at}")
to_execute = fn(*args, **kwargs)
print(f">>> Function: {fn.__name__!r} executed. Logged at {called_at}")
return to_execute
return inner
@logger
def connect_db():
print("Connecting to DB")
connection_uri = "postgresql+psycopg2://{}:{}@{}:{}".format(
CONFIG["POSTGRES_USER"],
CONFIG["POSTGRES_PASSWORD"],
CONFIG["POSTGRES_HOST"],
CONFIG["POSTGRES_PORT"],
)
engine = create_engine(connection_uri, pool_pre_ping=True)
engine.connect()
return engine
@logger
def extract(dataset_url):
print(f"Reading dataset from {dataset_url}")
df = pd.read_csv(dataset_url)
return df
@logger
def transform(df):
# transformation
print("Transforming data")
def impute_color(x):
return 0 if x == "white" else 1
df["winecolor"] = df["winecolor"].apply(impute_color)
return df
@logger
def check_table_exists(table_name, engine):
if table_name in inspect(engine).get_table_names():
print(f"{table_name!r} exists in the DB!")
else:
print(f"{table_name} does not exist in the DB!")
@logger
def load_to_db(df, table_name, engine):
print(f"Loading dataframe to DB on table: {table_name}")
df.to_sql(table_name, engine, if_exists="replace")
check_table_exists(table_name, engine)
@logger
def etl():
db_engine = connect_db()
raw_df = extract(DATASET_URL)
raw_table_name = "wine_quality_raw_dataset"
clean_df = transform(raw_df)
clean_table_name = "wine_quality_clean_dataset"
load_to_db(raw_df, raw_table_name, db_engine)
load_to_db(clean_df, clean_table_name, db_engine)
with dag:
run_etl_task = PythonOperator(task_id="run_etl_task", python_callable=etl)
# -
| Simple ETL.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 ('base')
# language: python
# name: python3
# ---
import sqlite3
import re #Expressoes regulares
import time #Tempo/hora
import pycountry #Leitura de países em siglas formato ISO
import numpy as np #Manipulação de dados
import pandas as pd # ^^
import matplotlib.pyplot as plt #Criação de gráficos
import seaborn as sns # ^^
from matplotlib import cm
from sklearn.feature_extraction.text import CountVectorizer #Vetor para calculos
import warnings
warnings.filterwarnings('ignore')
sns.set_theme(style = "whitegrid")
# !imdb-sqlite
conexao = sqlite3.connect("imdb.db")
tabelas = pd.read_sql_query("SELECT NAME AS 'Table_name' FROM sqlite_master WHERE type ='table'", conexao)
# #### Quais são os tipos mais comuns de obras no IMDB?
#
# Se a obra é um longa metragem, um curta, um especial de TV, por exemplo
# +
#Consulta SQL
consultaTipo = '''SELECT type,COUNT(*) AS COUNT FROM titles GROUP BY type'''
#Extração do resultado
resultadoTipo = pd.read_sql_query(consultaTipo, conexao)
#Cálculo percentual de cada gênero
resultadoTipo['percentual'] = (resultadoTipo['COUNT'] / resultadoTipo['COUNT'].sum()) * 100
#Exibição do resultado
display(resultadoTipo)
# +
#Ordenação e filtragem de dados
#Criação de um dicionário vazio
others = {}
#Filtro de no máximo 5% (categorias que possuem abaixo de 5% serão consideradas "others")
others['COUNT'] = resultadoTipo[resultadoTipo['percentual'] < 5]['COUNT'].sum()
#Gravação do percentual
others['percentual'] = resultadoTipo[resultadoTipo['percentual'] < 5]['percentual'].sum()
#Ajuste do nome
others['type'] = 'Others'
#Filtro dos demais tipos de obras (apenas 3 categorias pontuam mais de 5%)
resultadoTipo = resultadoTipo[resultadoTipo['percentual'] > 5]
#Concatenação com o dataframe das demais categorias
resultadoTipo = resultadoTipo.append(others, ignore_index = True)
#Ordenação do resultado
resultadoTipo = resultadoTipo.sort_values(by = 'COUNT', ascending = False)
# +
#Criação de gráficos
#Ajuste das labels
labels = [str(resultadoTipo['type'][i]) + ' ' + '[' + str(round(resultadoTipo['percentual'][i], 2)) + '%' + ']' for i in resultadoTipo.index]
#Escolha das cores
cs = cm.Set3(np.arange(100))
#Criação da figura
figura = plt.figure
#Gráfico de rosca
plt.pie(resultadoTipo['COUNT'], labeldistance = 1, radius = 3, colors = cs, wedgeprops = dict(width = 0.8))
plt.legend(labels = labels, loc = 'center', prop = {'size': 12})
plt.title("Distribuição de Títulos", loc = 'Center', fontdict = {'fontsize':20, 'fontweight':20})
plt.show
# -
# #### Quais são os gêneros mais comuns de filmes no IMDB?
#
# +
#"Selecione gêneros, contando todos a partir de títulos, onde o tipo é "filme", ordene por gênero"
consultaGenero = '''SELECT genres, COUNT(*) FROM titles WHERE type = 'movie' GROUP BY genres'''
resultadoGenero = pd.read_sql_query(consultaGenero, conexao)
# -
#Remoção de valores vazios (\N)
removeVazio = resultadoGenero['genres'].dropna()
# +
display(resultadoGenero)
# +
#Criação de um count vectorizer
#Expressão regular para filtrar as strings
regex = '(?u)\\b[\\w-]+\\b'
#Construção do vetor
vetor = CountVectorizer(token_pattern = regex, analyzer = 'word').fit(removeVazio)
#Vetorização do dataset
bag_generos = vetor.transform(removeVazio)
#Retorno de gêneros únicos
generos_unicos = vetor.get_feature_names()
#Dataframe de gêneros
generos = pd.DataFrame(bag_generos.todense(), columns = generos_unicos, index = removeVazio.index)
# -
#Visualização parcial dos dados
generos.info()
#Remoção da coluna N, criada sem razão
generos = generos.drop(columns = 'n', axis = 0)
# +
#Lapidação dos dados e criação do gráfico
#Percentual de gêneros
generos_percentual = 100 * pd.Series(generos.sum()).sort_values(ascending = False) / generos.shape[0]
#Criação do plot
plt.Figure(figsize = (16, 8))
sns.barplot(x = generos_percentual.values, y = generos_percentual.index, orient = "h", palette = "terrain" )
plt.ylabel("Gênero")
plt.xlabel("\nPercentual de Filmes (%)")
plt.title('\nNúmero de Títulos por Gênero\n')
plt.show
# -
# #### Quais países produzem mais filmes?
#
consultaPaises = ''' SELECT region, COUNT(*) Number_of_movies FROM
akas JOIN titles ON
akas.title_id = titles.title_id
WHERE region != 'None'
AND type = \'movie\'
GROUP BY region
'''
# Conta-se o número de filmes de cada regiao (akas), eliminando regiões vazias, filtrando para apenas filmes e agrupando por região
# Visualização apenas por siglas atrapalha a interpretação
resultadoPaises = pd.read_sql_query(consultaPaises, conexao)
display(resultadoPaises)
# +
# Listas auxiliares
nomes_paises = []
contagem = []
# Loop para obter o país de acordo com a região
for i in range (resultadoPaises.shape[0]):
try:
coun = resultadoPaises['region'].values[i]
nomes_paises.append(pycountry.countries.get(alpha_2 = coun).name) # Usa-se o pycountry para converter as siglas em nomes de países
contagem.append(resultadoPaises['Number_of_movies'].values[i])
except:
continue
# +
# Preparação do dataframe
df_filmes_paises = pd.DataFrame()
df_filmes_paises['country'] = nomes_paises
df_filmes_paises['Movie_Count'] = contagem
# Ordenação dos resultados
df_filmes_paises = df_filmes_paises.sort_values(by = 'Movie_Count', ascending = False)
df_filmes_paises.head(100)
# +
# Criação do gráfico
plt.figure(figsize = (20,8))
sns.barplot(y = df_filmes_paises[:20].country, x = df_filmes_paises[:20].Movie_Count, orient = 'h')
for i in range(0,20):
plt.text(df_filmes_paises.Movie_Count[df_filmes_paises.index[i]]-1,
i + 0.30, round(df_filmes_paises["Movie_Count"][df_filmes_paises.index[i]],2))
plt.ylabel('País')
plt.xlabel('\nNúmero de Filmes')
plt.title('\nNúmero de Filmes por País\n')
plt.show
| analise_IMDB/analise_imdb.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 股指对冲
# ### 1.股指期货概述
# 股指期货,用于对冲股票市场风险的金融衍生工具,恰当的应用可以有效的规避投资风险,股指期货T0交易制度,双向交易,且支持做多也支持做空的交易方式,是对T 1交易制度、单向交易的股票市场是极大的扩展,同时也兼具期货本身的套利、投机等功能属性。
# 在量化交易策略中,有不少追求alpha收益的策略,如果能够找到稳定赚钱的alpha,就可以通过指数期货的对冲,规避掉市场风险部分,获取超额收益。
# 和其他的期货品种一样,期货市场同样也存在着期保值者、投机者、套利者,股指期货市场的表现同样也表达了投资者对股票市场未来走势看法,如在2015年熊市挡住,股指期货就曾被认为是股市下跌的元凶。
# 股指期货到期进行交割时就是股票指数了,受政策变化、市场流动性缩减、手续费提升的影响,指数期货与股票指数的实际走势也发生着变化,本篇文章就是带大家看一下股指期货与股指的具体表现。
# 基差率(期货价格/指数价格 -1)
# ### 2.股指期货的几个重要时间点
# 目前,市场上已经开放的股指期货有三个,沪深300股指期货(交易代码IF)、中证500股指期货(交易代码IC),上证50股指期货(交易代码IH),首先,这里我们先整体看一下三个指数期货发布以来到现在的成交量、基差率(期货价格/指数价格 -1)的变化情况
# * 沪深300股指期货:沪深300指数自2005年4月8日正式发布,指数期货在2010年4月16日发布
# * 上证50股指期货:上证50指数自2004年1月2日起正式发布,指数期货在2015年4月16日发布
# * 中证500股指期货:中证500指数在2007年1月15日正式发布,指数期货在2015年4月16日发布
# 其次,列出几次重要的制度变化时间点
# * 金融期货限制:2015年9月7日开始,限制内容,,,,,,
# (将期指非套保持仓保证金提高至40%,平仓手续费提高至万分之二十三,单个产品单日开仓交易量超过10手认定为异常交易行为,旨在进一步抑制市场过度投机)
# * 金融期货第一次松绑:2017年2月17日,,,
# (自2月17日起,股指期货日内过度交易行为的监管标准从原先的10手调整为20手,套期保值交易开仓数量不受此限;自2月17日结算时起,沪深300、上证50股指期货非套期保值交易保证金调整为20%,中证500股指期货非套期保值交易保证金调整为30%(三个产品套保持仓交易保证金维持20%不变);自2月17日起,沪深300、上证50、中证500股指期货平今仓交易手续费调整为成交金额的万分之九点二。)
# * 金融期货第三次松绑:2018年12月3日,,,
# (中金所发布公告称,经中国证监会同意,自2018年12月3日结算时起,将沪深300、上证50股指期货交易保证金标准统一调整为10%,中证500股指期货交易保证金标准统一调整为15%;二是自2018年12月3日起,将股指期货日内过度交易行为的监管标准调整为单个合约50手,套期保值交易开仓数量不受此限;三是自2018年12月3日起,将股指期货平今仓交易手续费标准调整为成交金额的万分之四点六。)
# ### 3.不同时间段的表现
# 沪深300股指期货分阶段的成交量、基差的均值数据
# 
# 中证500股指期货分阶段的成交量、基差数据统计
# 
# 上证50股指期货分阶段的成交量、基差数据统计
# 
# ### 4.分阶段描述各个时期的特点。
# 
# 这张图统计了自2015年限制之后,三个股指期货的成交量变化,成交量均在稳步提升,我们看到在政策不断松绑的
# 成交量回升显著,上证50从近5000手上升至超过2万手,沪深300从1.25万手上升智4万手以上
# 中证500中间出现了缩量的情况,但当前月的均量也已经是成倍的增长
# (上证50、沪深300集中了大盘蓝筹股、中证500则反映着中小股的走势,从2017年年初之后,市场就是大盘股主导,估计与这行情有着一定的关系)
# 
# 再来看基差率
#
# 1.基差逐步减小,深度贴水的状况逐渐改善
# 2.在2015年4月到2015年9月期间,即没有做限制之前,基差均值非常大,说明过度投机的市场,也会导致股指期货与股指偏离过大.
# 3.自2015年股灾之后基差均值首次出现正值
# 4.中证500基差率近几年均值一直维持在0.5%左右
| .ipynb_checkpoints/6.1 股指对冲-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class Queue_using_two_stack:
def __init__(self):
self.__Array1 = []
self.__Array2 = []
def Enqueue(self, data):
self.__Array1.append(data)
def Dequeue(self):
if self.IsEmpty_of_Queue():
print("Hey Queue is Empty")
return None
while not self.IsEmpty_of_Queue():
data = self.__Array1.pop()
self.__Array2.append(data)
popped_element = self.__Array2.pop()
while not self.IsEmpty_2():
data = self.__Array2.pop()
self.__Array1.append(data)
return popped_element
def Front(self):
if self.IsEmpty_of_Queue():
print("Hey Queue is Empty")
return None
while not self.IsEmpty_of_Queue():
data = self.__Array1.pop()
self.__Array2.append(data)
front_element = self.__Array2[len(self.__Array2) - len(self.__Array2)]
while not self.IsEmpty_2():
data = self.__Array2.pop()
self.__Array1.append(data)
return front_element
def Size_of_Queue(self):
return len(self.__Array1)
def size_2(self):
return len(self.__Array2)
def IsEmpty_of_Queue(self):
return self.Size_of_Queue() == 0
def IsEmpty_2(self):
return self.size_2() == 0
# + pycharm={"name": "#%%\n"}
q = Queue_using_two_stack()
q.Enqueue(1)
q.Enqueue(2)
q.Dequeue()
q.Dequeue()
q.Size_of_Queue()
q.IsEmpty_of_Queue()
q.Dequeue()
q.Front()
q.Enqueue(11)
q.Front()
| 14. Queues/Queue_using_two_Stack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.5 64-bit
# name: python3
# ---
with open('/media/julian/Datasets1/french_corpus/wiki_news.txt', 'r') as f:
corpus = f.readlines()
len(corpus)
vocab = set(corpus[0][:-1].split(' '))
for sentence in corpus:
for word in sentence[:-1].split(' '):
vocab.add(word)
vocab = sorted(list(vocab))
def generate_line(word):
return word + '\t' + ''.join(list(map(lambda a: a + ' ', list(word)))) + '|\n'
with open('./lexicon.lst', 'a') as f:
for word in vocab:
f.writelines([generate_line(word)])
| utils/generate-vocab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: jobflow
# language: python
# name: jobflow
# ---
# # Five-minute quickstart
#
# In this quickstart, you will:
#
# - Create some jobs.
# - Use the jobs in a Flow.
# - Run the Flow.
# - Examine the outputs of the Flow.
# - Get a flavor of the Python API
#
# This tutorial will emphasize "hands-on" usage of jobflow and not explain things in detail.
#
# ## Define jobs
#
# The atomic building block of jobflows are jobs. Creating a job is as easy as writing a python function. All you need to do is use the `@job` decorator.
# + nbsphinx="hidden"
import warnings
warnings.filterwarnings("ignore", "Using `tqdm.autonotebook.tqdm`")
# +
from jobflow import job
@job
def add(a, b):
return a + b
# -
# Any call to the `add` function will return a `Job` object. This is essentially a function call that will be executed later.
add_first = add(1, 5)
# Jobs have outputs that can be accessed using the `output` attribute. As the job has not yet been executed, the output is currently a reference to the future output.
add_first.output
# The output of a job can be used as the input to another job.
add_second = add(add_first.output, 3)
# ## Create a Flow
#
# A Flow is a collection of Jobs or other Flow objects. Let's create a Flow from the `add_first` and `add_second` jobs we just made:
# +
from jobflow import Flow
flow = Flow([add_first, add_second])
# -
# The order of the jobs in the input array does not matter. Their execution order will be determined by their connectivity. Because `add_second` takes the output of `add_first` as an input, the `add_first` will always run before `add_second`.
#
# The connectivity of the jobs in a flow can be visualized using:
flow.draw_graph(figsize=(3, 3)).show()
# ## Run the Flow
# Jobflow supports running Flows locally or on remote clusters. Below we run the Flow locally using the `run_locally` function.
#
# <div class="alert alert-info">
# **Note**
# <p>Running Flows on remote clusters can be achieved using the FireWorks package and is covered in the [Running Flows with FireWorks tutorial](https://hackingmaterials.lbl.gov/jobflow/tutorials/6-fireworks.html).</p>
# </div>
# +
from jobflow.managers.local import run_locally
responses = run_locally(flow)
# -
# The numbers in brackets after the job function name give the job unique identifier (UUID).
# ## Examine Flow outputs
# The `run_locally` function returns the output of all jobs. The the format of the output is:
#
# ```python
# {
# job_uuid: {
# job_index: {
# Response()
# }
# }
# }
# ```
#
# The `job_index` and `Response()` objects are introduced in later tutorials. The main thing to know is that the Response contains the output of the Job and any other commands for controlling the Flow execution.
responses
# We can check the output of specific jobs using:
responses[add_first.uuid][1].output
# ## Next steps
#
# Now that you’ve successfully run your first Flow, we encourage you to learn about all the different options jobflow provides for designing and running workflows. A good next step is the [Introductory tutorial](https://materialsproject.github.io/jobflow/tutorials/2-introduction.html), which covers things more slowly than this quickstart.
| tutorials/1-quickstart.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.neighbors import NearestNeighbors
import statsmodels.formula.api as smf
from scipy.stats import ttest_ind
import matplotlib.pyplot as plt
import pandas as pd
pd.options.display.float_format = "{:,.2f}".format
# -
# # Regression and matching estimators in causal effects
# In this problem set we are going to compare the consistency of regression and matching estimators of causal effects based on Dehejia & Wahba (1999). For that we employ the experimental study from LaLonde (1986), which provides an opportunity to estimate true treatment effects. We then use these results to evaluate the performance of (treatment effect) estimators one can usuallly obtain in observational studies.
#
# LaLonde (1986) implements the data from the National Supported Work program (NSW) -- temporary employment program designed to help disadvantaged workers lacking basic job skills move into the labor market by giving them work experience and counseling in sheltered environment. Unlike other federally sponsored employment programs, the NSW
# program assigned qualiffed applications randomly. Those assigned to the treatment group received all the bene
# ts of the NSW program, while those assigned to the control group were left to fend for themselves.
#
# To produce the observational study, we select the sample from the Current Population Survey (CPS) as the comparison group and merge it with the treatment group. We do this to obtain a data set which resembles the data which is commonly used in scientific practice. The two data sets are explained below:*
#
# - **nsw_dehejia.csv** is field-experiment data from the NSW. It contains variables as education, age, ethnicity, marital status, preintervention (1975) and postintervention (1978) earnings of the eligible male applicants. Dehejia & Wahba (1999) also transform the LaLonde (1986) data set to have observations on preintervention 1974 earnings; motivation is explained in their paper.
#
# - **cps.csv** is a non-experimental sample from the CPS which selects all males under age 55 and contains the same range of variables.
# ## Task A
# *Create the table with the sample means of characteristics by age, education, preintervention earnings, etc. for treated and control groups of NSW sample (you can use the Table 1 from Dehejia and Wahba (1999) as a benchmark). Is the distribution of preintervention variables similar across the treatment and control groups? Check the differences on significance. Add to the table the CPS sample means. Is the comparison group different from the treatment group in terms of age, marital status, ethnicity, and preintervention earnings?*
# +
demographics = ["age", "ed", "black", "hisp", "married", "nodeg", "age2"]
dtypes = dict()
for column in ["treat"] + demographics:
dtypes[column] = int
df_nsw = pd.read_csv("data/nsw_dehejia.csv", dtype=dtypes)
df_nsw.index.name = "individual"
df_nsw.head()
# -
# How does a summary of the data look like?
df_nsw.describe()
# Let's look at the mean differences by treatment status.
df_nsw.groupby("treat").mean()
df_nsw.groupby("treat").mean().diff()
# Are these differences statistically significant?
for column in demographics:
treated = df_nsw.query("treat == 1")[column]
control = df_nsw.query("treat == 0")[column]
stat = ttest_ind(treated, control)[1]
print(f"{column:<7} {stat:7.3f}")
df_cps = pd.read_csv("data/cps.csv", dtype=dtypes)
df_cps.index.name = "individual"
df_cps.head()
# How does a summary of the data look like?
df_cps.describe()
# Let's compare mean differences between the synthetic control group and the treatment group.
for column in demographics:
treated = df_nsw.query("treat == 1")[column]
control = df_cps[column]
stat = ttest_ind(treated, control)[1]
print(f"{column:<7} {stat:7.3f}")
# ## Task B. Regression Adjustment
#
# *In this section we compare the results of regression estimates with selection on observables as discussed in the lecture 6.*
# ### Task B.1
# *Merge the treatment group data from the NSW sample with the comparison group data from the CPS sample to imitate an observational study.*
# +
df_nsw["sample"] = "NSW"
df_cps["sample"] = "CPS"
df_obs = pd.concat([df_nsw.query("treat == 1"), df_cps])
df_obs.set_index(["sample"], append=True, inplace=True)
df_obs.sort_index(inplace=True)
df_obs.loc[(slice(1, 5), "NSW"), :]
# -
# ### Task B.2
# *Which assumption need to hold such that conditioning on observables can help in obtaining an unbiased estimate of the true treatment effect?*
# $$E[Y^1|D = 1, S] = E[Y^1|D = 0, S]$$
# $$E[Y^0|D = 1, S] = E[Y^0|D = 0, S]$$
# ### Task B.3
# *Run a regression on both experimental and non-experimental data using the specification: RE78 on a constant, a treatment indicator, age, age2, education, marital status, no degree, black, hispanic, RE74, and RE75. We recommend using statsmodels, but you are free to use any other software. Is the treatment effect estimate of the observational study consistent with the true estimate?*
# We first construct the regression equation.
# +
indep_vars = df_obs.columns.tolist()
indep_vars.remove("re78")
formula = "re78 ~ " + " " " + ".join(indep_vars)
formula
# -
# Now we can run the model on both datasets.
for label, data in [("observational", df_obs), ("experimental", df_nsw)]:
stat = smf.ols(formula=formula, data=data).fit().params["treat"]
print(f"Estimate based on {label} data: {stat:7.3f}")
# ## Task C. Matching on Propensity Score
#
# Recall that the propensity score p(Si) is the probability of unit i having been assigned to treatment. Most commonly this function is modeled to be dependent on various covariates. We write $p(S_i) := Pr(D_i = 1|S_i) = E(D_i|S_i).$ One assumption that makes estimation strategies feasible is $S_i \perp D_i|p(S_i)$ which means that, conditional on the propensity score, the covariates are independent of assignment to treatment. Therefore, conditioning on the propensity score, each individual has the same probability of assignment to treatment,
# as in a randomized experiment.*
#
# Estimation is done in two steps. First, we estimate the propensity score using a logistic regression model. Secondly, we match the observations on propensity score employing nearest-neighbor algorithm discussed in the lecture 5. That is, each treatment unit is matched to the comparison unit with the closest propensity score -- the unmatched comparison units are discarded.
# ### Task C.1
# *Before we start with matching on propensity score, let's come back to another matching strategy which was discussed in Lecture 5 - matching on stratification. Looking at the data could you name at least two potential reasons why matching on stratification might be impossible to use here?*
# Data contains continuous variables; formed stratas might not have treated and control units available at the same time.
# ### Task C.2
# *Employing our imitated observational data run a logistic regression on the following specification: treatment indicator on age, education, marital status, no degree, black, hispanic, RE74, and RE75. Use, for example, [statsmodels](https://www.statsmodels.org/stable/index.html) for this task. Then extract a propensity score for every individual as a probability to be assigned into treatment.*
formula = "treat ~ age + ed + black + hisp + married + nodeg + re74 + re75"
df_obs["pscore"] = smf.logit(formula=formula, data=df_obs).fit().predict()
# ### Task C.3
# *Before proceeding further we have to be sure that propensity scores of treatment units overlap with the propensity scores of control units. Draw a figure showing the distribution of propensity score across treatment and control units (we use the packages matplotlib and seaborn). Do we observe common support?*
# +
fig, ax = plt.subplots()
df_control = df_obs.query("treat == 0")["pscore"]
df_treated = df_obs.query("treat == 1")["pscore"]
ax.hist([df_control, df_treated], density=True, label=["Control", "Treated"])
ax.set_ylim(0, 5)
ax.set_xlim(0, 1)
ax.set_ylabel("Density")
ax.set_xlabel("Propensity scores")
ax.legend()
# -
# ### Task C.4
# *Match each treatment unit with control unit one-to-one with replacement. We use the package sklearn.neighbors: apply the algorithm NearestNeighbors to the propensity score of treated and control units and extract the indices of matched control units.*
def get_matched_dataset(df):
training_data = df.query("treat == 0")["pscore"].to_numpy().reshape(-1, 1)
eval_point = df.query("treat == 1")["pscore"].to_numpy().reshape(-1, 1)
neigh = NearestNeighbors(n_neighbors=1)
neigh.fit(training_data)
matched = neigh.kneighbors(eval_point, return_distance=False)[:, 0]
df_treated = df.query("treat == 1")
df_matched = df.query("treat == 0").iloc[matched]
df_sample = pd.concat([df_treated, df_matched])
return df_sample
# ### Task C.5
# *Construct new data set with matched observations. Run the regression to obtain matching on propensity score estimate. Is it more or less consistent estimate of the true effect comparing to the regression estimate with selection on observables? How could you explain this result?*
df_sample = get_matched_dataset(df_obs)
stat = smf.ols(formula="re78 ~ treat", data=df_sample).fit().params["treat"]
print(f"Estimate based on matched for re78 data: {stat:7.3f}")
# Regression model neglects important nonlinear terms and interactions (Rubin 1973). The benefit of matching over regression is that it is non-parametric (but you do have to assume that you have the right propensity score specification in case of matching).
# Let's further explore two selected issues in matching, i.e. the use of placebo testing and trimming.
stat = smf.ols(formula="re75 ~ treat", data=df_sample).fit().params["treat"]
print(f"Estimate based on matched for re75 data: {stat:7.3f}")
# What happens if we trim our dataset?
for value in [0.025, 0.05, 0.1, 0.15]:
lower, upper = value, 1 - value
df_trimmed = df_obs.loc[df_obs["pscore"].between(lower, upper), :]
df_sample = get_matched_dataset(df_trimmed)
stat = smf.ols(formula="re78 ~ treat", data=df_sample).fit().params["treat"]
print(f"{value:5.3f}: {stat:7.3f}")
# ## References
#
# * **Bureau of Labor Statistics. (1974, 1975, 1978)**. [Current Population Survey](https://www.census.gov/programs-surveys/cps.html).
#
#
# * **<NAME>., and <NAME>. (1999)**. [Causal effects in nonexperimental studies: Reevaluating the evaluation of training programs](https://www.jstor.org/stable/2669919?seq=1). *Journal of the American Statistical Association*, 94(448), 1053-1062.
#
#
# * **<NAME>. (1986)**. [Evaluating the econometric evaluation of training programs with experimental data](https://www.jstor.org/stable/1806062?seq=1). *American Economic Review*, 76(4), 604-620.
| problem-sets/matching-estimators/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fast Python3 For Beginners
# ___
# ```
# try...
#
# except...
#
# finally...
# ```
# ### 1.try
try:
print('try...')
r = 10/0
print('result:', r)
except ZeroDivisionError as e:
print('except:', e)
finally:
print('finally')
print('END')
# **logging error**
# > Python内置的`logging`模块可以非常容易地**记录错误信息**:
#
# > Python's built-in `logging` module makes it very easy to record error messages:
# +
# err_logging.py
import logging
def foo(s):
return 10 / int(s)
def bar(s):
return foo(s) * 2
def main():
try:
bar('0')
except Exception as e:
logging.exception(e)
main()
print('END')
# -
# **raising error**
try:
a = input("type a number")
r = 10 / int(a)
if int(a) < 0:
raise ValueError
print('r:', r)
except ZeroDivisionError as e:
logging.exception(e)
finally:
print("finally")
print("END")
# ### 2.Debug
# **Method_1: `print()`**
# > 用print()把可能有问题的变量打印出来看看:
#
# > Use `print()` to print out variables that may be problematic.
# **Method_2: `assert`**
# > 凡是用print()来辅助查看的地方,都可以用断言(assert)来替代:
# assert的意思是,表达式n != 0应该是True,否则,根据程序运行的逻辑,后面的代码肯定会出错。
# 如果断言失败,assert语句本身就会抛出AssertionError
#
# > `print()` can be replaced with `assert`:
# Assert means that the judgement expression like `n != 0` should be True, otherwise, according to the logic of the program, the following code will definitely make mistakes.
# If the assertion fails, the assert statement itself throws an `Assertion Error`
# +
def foo(s):
n = int(s)
assert n != 0, 'n is zero!'
return 10 / n
def main():
foo('0')
main()
# -
# **Method_3: `logging`**
# > 和assert比,logging不会抛出错误,而且可以输出到文件:
#
# > Compared with `assert`, `logging` won't through `errors`, and can write error messages to file.
# +
import logging
logging.basicConfig(level=logging.INFO)
s = '0'
n = int(s)
logging.info('n = %d' % n)
print(10 / n)
# -
'Done!\N{Cat}'
| 03 Exception Handling/00_Exception_Handling_A.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.3 64-bit (''complete3.8'': conda)'
# language: python
# name: python38364bitcomplete38conda6b4851e4606c45b69c6094ba1f069d7d
# ---
# This notebook simulates data for a hypothetical farmland appraisal modeling task. We look in particular at the different model fits during k-fold cross-validation, the estimate of generalization error produced by cross-validation, and the confidence interval for the generalization error.
#
# This notebook produces figures published in The Crosstab Kite's article [Research Digest: What does cross-validation really estimate?](https://crosstab.io/articles/bates-cross-validation), which digests the research paper [Cross-validation: what does it estimate and how well does it do it?](https://arxiv.org/abs/2104.00673) by Bates, Hastie, and Tibshirani.
#
# The plot styling is intended for the figures as they appear in the article, so they look really bad in this notebook. That's known and ok.
# # 0. Setup
# +
import numpy as np
import pandas as pd
import plotly.offline as pyo
import plotly.graph_objects as go
from sklearn.model_selection import KFold
from sklearn.linear_model import LinearRegression
import scipy.stats as stats
pyo.init_notebook_mode()
# +
## Generic plot style
baseline_style = dict(
font=dict(family="Arial", size=36),
template="simple_white",
)
marker_size = 26
# -
# # 1. Generate data
# The true regression function of sale price is quadratic in property acreage. The distribution of acreage and sale prices is intended to very loosely mimic agricultural property values in the Hill Country of Texas, based on [data from Texas A&M](https://www.recenter.tamu.edu/data/rural-land/).
# +
np.random.seed(18)
n = 100
acreage_mean = 120
acreage_sd = 30
price_sd = 350000
target = "price"
# +
df = pd.DataFrame({"acres": np.random.normal(acreage_mean, acreage_sd, n)})
noise = np.random.normal(loc=0, scale=price_sd, size=n)
df["sq_acres"] = df["acres"] ** 2
df[target] = 2000 * df["acres"] + 50 * df["sq_acres"] + noise
df.sample(5)
# +
fig = go.Figure()
fig.add_trace(
go.Scatter(
x=df["acres"],
y=df["price"],
mode="markers",
marker=dict(
symbol="circle",
color="rgba(100, 149, 237, 0.35)",
size=marker_size,
line=dict(width=2, color="#15388d"),
),
showlegend=False,
)
)
fig.update_layout(baseline_style)
fig.update_layout(xaxis_title="Acres", yaxis_title="Sale price ($)")
fig.write_image("sim_farm_sales.png", height=1400, width=1400)
fig.show()
# -
# Make a grid of values for the `acres` features, for plotting quadratic model fits.
xgrid = pd.DataFrame(
{"acres": np.linspace(df["acres"].min() - 5, df["acres"].max() + 5, 100)}
)
xgrid["sq_acres"] = xgrid["acres"] ** 2
# # 2. Select the best model form with 5-fold cross-validation
# Scikit-learn has a convenience function [`cross_val_score`](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.cross_val_score.html#sklearn.model_selection.cross_val_score) that makes this a lot less verbose. Here we use the `KFold` iterator to show the steps more carefully and to more closely match the Bates, et al. paper. Specifically, the Bates, et al. paper computes the cross-validation error a little differently than most. Most sources say to take the average of the per-fold test errors, but Bates et al. record then error for each point when it's in the test, then take the average of all points at the end.
# +
linear_errors = np.array([])
quad_errors = np.array([])
kfold = KFold(n_splits=5)
for ix_train, ix_test in kfold.split(df):
# Split data
df_train = df.loc[ix_train]
df_test = df.loc[ix_test]
# Fit the linear model and get test RMSE
linear_model = LinearRegression()
linear_model.fit(df_train[["acres"]], df_train[[target]])
linear_ystar = linear_model.predict(df_test[["acres"]]).flatten()
linear_errors = np.append(linear_errors, (df_test[target] - linear_ystar))
# Draw the trained linear model on the plot.
fig.add_trace(
go.Scatter(
x=xgrid["acres"],
y=linear_model.predict(xgrid[["acres"]]).flatten(),
mode="lines",
line=dict(width=3, dash="dash", color="orange"),
showlegend=False,
)
)
# Fit the quadratic model and get test RMSE
quad_model = LinearRegression()
quad_model.fit(df_train[["acres", "sq_acres"]], df_train[target])
quad_ystar = quad_model.predict(df_test[["acres", "sq_acres"]]).flatten()
quad_errors = np.append(quad_errors, (df_test[target] - quad_ystar))
# Draw the trained quadratic model on the plot.
fig.add_trace(
go.Scatter(
x=xgrid["acres"],
y=quad_model.predict(xgrid[["acres", "sq_acres"]]).flatten(),
mode="lines",
line=dict(width=3, dash="dash", color="purple"),
showlegend=False,
)
)
linear_cv_rmse = (linear_errors ** 2).mean() ** 0.5
quad_cv_rmse = (quad_errors ** 2).mean() ** 0.5
print(f"{linear_cv_rmse=}")
print(f"{quad_cv_rmse=}")
# -
# As expected, given that the true regression function is quadratic, the quadratic form has lower cross-validation error.
# +
fig.add_annotation(
x=205,
y=1.65e6,
text=f"Linear model<br>5-fold CV fits<br>CV RMSE: ${linear_cv_rmse:,.2f}",
showarrow=False,
font=dict(color="orange"),
)
fig.add_annotation(
x=150,
y=2.8e6,
text=f"Quadratic model<br>5-fold CV fits<br>CV RMSE: ${quad_cv_rmse:,.2f}",
showarrow=False,
font=dict(color="purple"),
)
fig.write_image("cv_model_fits.png", height=1400, width=1400)
fig.show()
# -
# # 3. Re-fit best predictive model to the full dataset
final_model = LinearRegression()
final_model.fit(df[["acres", "sq_acres"]], df[target])
# # 4. Illustrate generalization
# What we really care about is the model's generalization error, which is the average model prediction error (measured by our squared error loss function) on new data points from the same distribution. Here we just manually create two new data points for the purpose of illustration on our schematic plot.
df_new = pd.DataFrame({"acres": [90, 170]})
df_new["sq_acres"] = df_new["acres"] ** 2
df_new["ystar"] = final_model.predict(df_new[["acres", "sq_acres"]])
df_new["price"] = [5.8e5, 1.1e6]
df_new
# Plot the final model with the new points and the model's predictions for those points.
# +
fig = go.Figure()
fig.add_trace(
go.Scatter(
x=df["acres"],
y=df["price"],
mode="markers",
marker=dict(
symbol="circle",
color="rgba(100, 149, 237, 0.35)",
size=marker_size,
line=dict(width=2, color="#15388d"),
),
showlegend=False,
)
)
fig.add_trace(
go.Scatter(
name="Final model",
x=xgrid["acres"],
y=final_model.predict(xgrid[["acres", "sq_acres"]]).flatten(),
mode="lines",
line=dict(width=6, color="purple"),
)
)
fig.add_trace(
go.Scatter(
name="New point true values (unknown)",
x=df_new["acres"],
y=df_new["price"],
mode="markers",
marker=dict(
symbol="circle-open", color="red", size=marker_size + 4, line_width=4
),
)
)
fig.add_trace(
go.Scatter(
name="New point predictions",
x=df_new["acres"],
y=df_new["ystar"],
mode="markers",
marker=dict(symbol="x", color="red", size=marker_size + 4),
)
)
fig.add_annotation(
x=200,
y=6.8e5,
text="Averge RMSE<br>for new points: ?",
showarrow=False,
font=dict(color="red"),
)
fig.update_layout(baseline_style)
fig.update_layout(
xaxis_title="Acres", yaxis_title="Sale price ($)", legend=dict(x=0.1, y=0.9)
)
fig.write_image("final_model.png", height=1400, width=1400)
fig.show()
# -
# # Naïve standard error and confidence interval of generalization error
# This is what Bates, et al. call the *naïve cross-validation interval*. As they show, this is not a good idea - the interval is too narrow to cover the true generalization error with the intended frequency.
#
# Note that even though our loss function is squared error, we take the square root here to get RMSE for interpretability.
significance = 0.1
tail_prob = 1 - significance / 2
z_quantile = stats.norm.ppf(tail_prob)
print(f"{z_quantile=}") # just write the value explicitly in the article
# +
std_err = (quad_errors ** 2).std(ddof=1) / np.sqrt(n)
avg_loss = quad_cv_rmse ** 2
rmse_ci_lower = (avg_loss - z_quantile * std_err) ** 0.5
rmse_ci_upper = (avg_loss + z_quantile * std_err) ** 0.5
print(f"{quad_cv_rmse=}")
print(f"{rmse_ci_lower=}")
print(f"{rmse_ci_upper=}")
# -
# This is a suprisingly high 90% confidence interval for generalization error. Crazy to think that it's not even wide enough to actually cover with 90% frequency.
| farmland_value_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <a href="https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv" target="_blank"><img align="left" src="data/cover.jpg" style="width: 76px; height: 100px; background: white; padding: 1px; border: 1px solid black; margin-right:10px;"></a>
# *This notebook contains an excerpt from the book [Machine Learning for OpenCV](https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv) by <NAME>.
# The code is released under the [MIT license](https://opensource.org/licenses/MIT),
# and is available on [GitHub](https://github.com/mbeyeler/opencv-machine-learning).*
#
# *Note that this excerpt contains only the raw code - the book is rich with additional explanations and illustrations.
# If you find this content useful, please consider supporting the work by
# [buying the book](https://www.packtpub.com/big-data-and-business-intelligence/machine-learning-opencv)!*
# <!--NAVIGATION-->
# < [Implementing Agglomerative Hierarchical Clustering](08.04-Implementing-Agglomerative-Hierarchical-Clustering.ipynb) | [Contents](../README.md) | [Understanding Perceptrons](09.01-Understanding-perceptrons.ipynb) >
# # Using Deep Learning to Classify Handwritten Digits
#
# In this chapter, we want to wrap our heads around some simple versions of artificial neural
# nets, such as the McCulloch-Pitts neuron, the [perceptron](09.01-Understanding-Perceptrons.ipynb),
# and the [multi-layer perceptron](09.02-Implementing-a-Multi-Layer-Perceptron-in-OpenCV.ipynb).
# Once we have familiarized ourselves with the basics, we will be ready to implement a more
# [sophisticated deep neural net](09.03-Getting-Acquainted-with-Deep-Learning.ipynb)
# in order to classify handwritten digits from the popular
# [MNIST database](09.04-Classifying-Handwritten-Digits.ipynb)
# (short for Mixed National Institute of Standards and Technology
# database). For this, we will be making use of [Keras](09.05-Training-a-Deep-Neural-Net-Using-Keras.ipynb),
# a high-level neural network library,
# which is also frequently used by researchers and tech companies.
#
# Along the way, we want to get answers to the following questions:
# - How do I implement perceptrons and multilayer perceptrons in OpenCV?
# - What is the difference between stochastic and batch gradient descent, and how does it fit in with backpropagation?
# - How do I know what size my neural net should be?
# - How can I use Keras to build sophisticated deep neural networks?
#
#
# ## Outline
#
# - [Understanding Perceptrons](09.01-Understanding-Perceptrons.ipynb)
# - [Implementing a Multi-Layer Perceptron in OpenCV](09.02-Implementing-a-Multi-Layer-Perceptron-in-OpenCV.ipynb)
# - [Getting Acquainted with Deep Learning](09.03-Getting-Acquainted-with-Deep-Learning.ipynb)
# - [Training an MLP in OpenCV to Classify Handwritten Digits](09.04-Training-an-MLP-in-OpenCV-to-Classify-Handwritten-Digits)
# - [Training a Deep Neural Net to Classify Handwritten Digits Using Keras](09.05-Training-a-Deep-Neural-Net-to-Classify-Handwritten-Digits-Using-Keras)
#
# > The book offers a detailed treatment of the McCulloch-Pitts neuron, the perceptron, multi-layer perceptrons, and backrpopagation. For more information on the same, please refer to the book.
#
#
# Excited? Then let's go!
# <!--NAVIGATION-->
# < [Implementing Agglomerative Hierarchical Clustering](08.04-Implementing-Agglomerative-Hierarchical-Clustering.ipynb) | [Contents](../README.md) | [Understanding Perceptrons](09.01-Understanding-perceptrons.ipynb) >
| notebooks/09.00-Using-Deep-Learning-to-Classify-Handwritten-Digits.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using PyMC3 samplers on PyMC4 models
# %load_ext autoreload
# %autoreload 2
import pymc4 as pm
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
# +
# Create simple pymc4 model
@pm.model(auto_name=True)
def t_test():
mu = pm.Normal(0, 1)
model = t_test.configure()
model._forward_context.vars
func = model.make_log_prob_function()
# -
# Create function to evaluate logp and dlogp over array of inputs
@tf.function
def logp_array(array):
#mu = array[0]
with tf.GradientTape() as tape:
tape.watch(array)
logp = func(array)
grad = tape.gradient(logp, array)
return logp, grad
# As the above function expects TF inputs and outputs, wrap it as PyMC3's samplers want numpy
def logp_wrapper(array):
logp, grad = logp_array(tf.convert_to_tensor(array))
return logp.numpy(), grad.numpy()
from pymc4.hmc import HamiltonianMC
size = 1
n_samples = 500
tf.random.set_seed(123)
np.random.seed(123)
hmc = HamiltonianMC(logp_dlogp_func=logp_wrapper, size=size, adapt_step_size=False)
curr = np.ones(size, dtype='float32') * .05
posterior_samples = []
stats = []
# +
# # %%time # NB: uncommenting cell magic %%time will prevent variable from escaping local cell scope
for i in range(n_samples):
curr, stat = hmc.step(curr)
posterior_samples.append(curr)
stats.append(stat)
if i % 10 == 0:
print(i)
print(hmc.step_size)
trace = np.array(posterior_samples)
# -
# Compare with `PyMC3`
import pymc3 as pm3
with pm3.Model() as model3:
pm3.Normal('x', 0, 1)
# +
np.random.seed(123)
with model3:
hmc3 = pm3.HamiltonianMC(adapt_step_size=True)
point = {'x': np.array(.05)}
trace3 = []
# +
# %%time
for i in range(n_samples):
point, _ = hmc3.step(point)
trace3.append(point['x'])
if i % 10 == 0:
print(i)
print(hmc3.step_size)
# -
import seaborn as sns
sns.distplot(trace)
sns.distplot(trace3)
# There still seems to be a problem here where in the PyMC4 implementation, the step_size keeps getting smaller and smaller, causing the sampler to take very long. Haven't figured it out yet.
hmc.step_size
hmc3.step_size
hmc.potential._stds
hmc3.potential._stds
| notebooks/context_design/pymc3_samplers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="--OoGccFbvaj"
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/AbdelMahm/FSR/blob/master/IDDLO-29-20/Notebooks/PCA.ipynb"><img src="https://colab.research.google.com/img/colab_favicon_256px.png" />Run in Google Colab</a>
# </td>
# </table>
# + [markdown] id="LUs6MdhLbva3"
# # Dimensionality Reduction
# + id="T7G_YnfCbva6"
import sys
import sklearn
import os
# %matplotlib inline
import matplotlib as mpl
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import sklearn.linear_model
import urllib.request
from sklearn import preprocessing
from sklearn import pipeline
# + [markdown] id="c1-Q3xmNbva8"
# ### Load Data
# + id="tySv0M0qbva8" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="1f2f2c05-9cc6-4d16-ae06-96f93bd7f356"
url = "https://archive.ics.uci.edu/ml/machine-learning-databases/iris/iris.data"
# load dataset into Pandas DataFrame
df = pd.read_csv(url, names=['sepal length','sepal width','petal length','petal width','target'])
df.head()
# + id="KXnV2_ELbva9"
from sklearn.preprocessing import StandardScaler
features = ['sepal length', 'sepal width', 'petal length', 'petal width']
# Separating out the features
X = df.loc[:, features].values
# Separating out the target
y = df.loc[:,['target']].values
# Standardizing the features
X = StandardScaler().fit_transform(X)
# + [markdown] id="k1jNGAiqbva-"
# ### PCA components
# + id="kpKcKvebbva-" colab={"base_uri": "https://localhost:8080/"} outputId="5cc8a9e1-a55b-4ec7-d1b0-21a54606edf6"
from sklearn.decomposition import PCA
#pca = PCA(n_components=2)
pca = PCA(n_components=3)
principalComponents = pca.fit_transform(X)
#principalDf = pd.DataFrame(data = principalComponents, columns = ['PC1', 'PC2'])
principalDf = pd.DataFrame(data = principalComponents, columns = ['PC1', 'PC2','PC3'])
pca.explained_variance_ratio_
# + [markdown] id="FnhCpDB32Prh"
# on remarque que meme si on projete nos données sur 3 axes , l'axe PC1 concentre toujours 72% de l'information
# + id="fwvmUkIBbvbA" colab={"base_uri": "https://localhost:8080/", "height": 206} outputId="73bb909d-eb3f-4563-dcf3-89c343cc858d"
finalDf = pd.concat([principalDf, df[['target']]], axis = 1)
finalDf.head()
# + [markdown] id="bDLAxB4PbvbB"
# ### Visualization
# + id="tZHVdveWbvbC" colab={"base_uri": "https://localhost:8080/", "height": 523} outputId="1386072d-042b-4464-d182-c088a0e22e37"
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('PC1', fontsize = 15)
ax.set_ylabel('PC2', fontsize = 15)
ax.set_title('2 component PCA', fontsize = 20)
targets = ['Iris-setosa', 'Iris-versicolor', 'Iris-virginica']
colors = ['r', 'g', 'b']
for target, color in zip(targets,colors):
indicesToKeep = finalDf['target'] == target
ax.scatter(finalDf.loc[indicesToKeep, 'PC1']
, finalDf.loc[indicesToKeep, 'PC2']
, c = color
, s = 50)
ax.legend(targets)
ax.grid()
# + id="zbX64QDebvbC" colab={"base_uri": "https://localhost:8080/"} outputId="beb89c41-58b5-4b99-8668-5302468a3368"
pca.explained_variance_ratio_
# + [markdown] id="gNgeFfGgfUIa"
# L'axe PC1 concentre 72.7% de l'information
# + [markdown] id="uNCzTnSYbvbD"
# ## PCA to speed up ML model
# + id="SjDkqruxbvbE"
from sklearn.datasets import fetch_openml
X, y = fetch_openml('mnist_784', version=1, return_X_y=True)
# + id="bDXbJy2GbvbF"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, train_size=6/7, test_size=10000)
# + [markdown] id="tm-9Kul9bvbF"
# ### Apply Logistic Regression without PCA
# + id="fC_rDJyvbvbG"
# %%time
from sklearn.linear_model import LogisticRegression
logisticRegr = LogisticRegression(solver = 'lbfgs')
logisticRegr.fit(X_train, y_train)
score = logisticRegr.score(X_test, y_test)
print(score)
# + [markdown] id="eQqzBDaYbvbG"
# ### Apply Logistic Regression after PCA
# + id="zuIkKt8NbvbG"
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
# Fit on training set only.
scaler.fit(X_train)
# Apply transform to both the training set and the test set.
X_train_scale = scaler.transform(X_train)
X_test_scale = scaler.transform(X_test)
# + id="N914X0ElbvbH"
from sklearn.decomposition import PCA
# Make an instance of the Model (retain 95% of the variance)
pca = PCA(.95)
pca.fit(X_train)
pca.n_components_
# + id="uWHMzZCMbvbI"
X_train_scale_pca = pca.transform(X_train_scale)
X_test_scale_pca = pca.transform(X_test_scale)
# + id="NFIpjh9tbvbJ"
# %%time
from sklearn.linear_model import LogisticRegression
logisticRegr = LogisticRegression(solver = 'lbfgs')
logisticRegr.fit(X_train_scale_pca, y_train)
score_pca = logisticRegr.score(X_test_scale_pca, y_test)
print(score_pca)
# + [markdown] id="VRpl1VXnbvbM"
# ## Return to Original Space (5% missed)
# + id="aH7FGWyMbvbR"
lower_dimensional_data = pca.fit_transform(X_train) # fit and transform in the same time
# + id="cbuGqFdAbvbS"
approximation = pca.inverse_transform(lower_dimensional_data)
# + id="QMHot9F6bvbT"
plt.figure(figsize=(8,4));
# Original Image
plt.subplot(1, 2, 1);
plt.imshow(X_train[2].reshape(28,28),
cmap = plt.cm.gray, interpolation='nearest',
clim=(0, 255));
plt.xlabel('784 components', fontsize = 14)
plt.title('Original Image', fontsize = 20);
# 154 principal components
plt.subplot(1, 2, 2);
plt.imshow(approximation[2].reshape(28, 28),
cmap = plt.cm.gray, interpolation='nearest',
clim=(0, 255));
plt.xlabel('154 components', fontsize = 14)
plt.title('95% of Explained Variance', fontsize = 20);
# + id="9oJ1rdgUbvbT"
| Copie_de_PCA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # LSTM-FCN for time series classification
#
# Project on [GitHub](https://github.com/titu1994/LSTM-FCN), [paper](https://ieeexplore.ieee.org/document/8141873/). Models work OK for univariate time series.
#
# 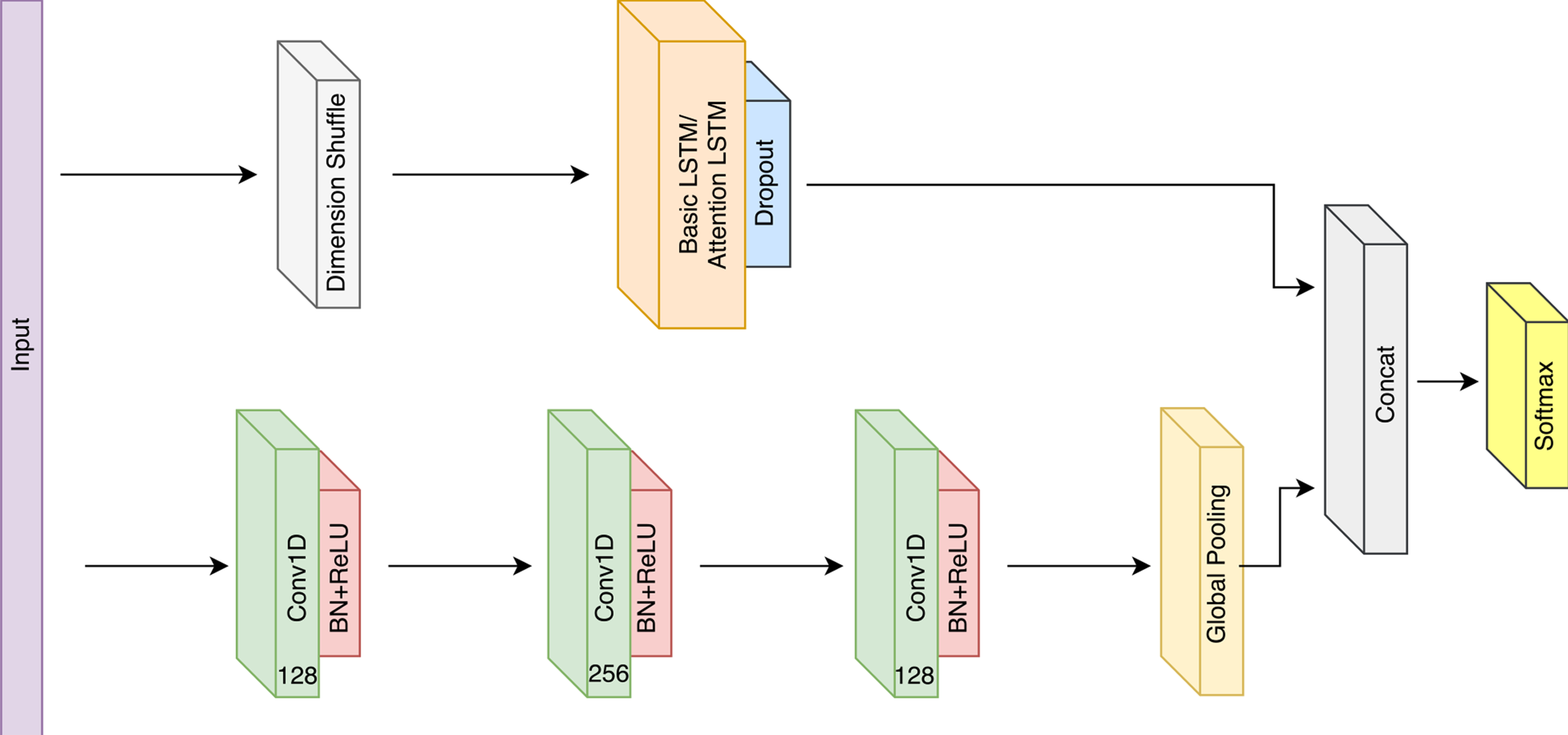
from nns.nns import (estimate, printable_dataframe)
from nns.models.ts_classification import LSTM_FCN
# # LSTM FCN
perf = []
estimate(LSTM_FCN(), perf, perf)
printable_dataframe(perf, ignore_phase=False)
| notebooks/lstm_fcn.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .js
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Javascript (Node.js)
// language: javascript
// name: javascript
// ---
// # Lab Four
// ---
//
// Ok for this lab we're going to complete our Iteration 0.
//
// Our Goals are:
// - Introduce the team
// - Define team meeting cadence, means of communication, and scheduled activities
// - Discuss means of documentation, and development
// - Create Design Specifications
// - Create Initial Backlog
// - Assign Roles (These will change since we want every one to be in every role at some point)
//
// ## First things first, what's your initial idea for the project!?
// This is allowed to change and be altered but the alterations have to be approved by the shareholder! You can't just show up with a different product than you promised. That's the quickest way to not get paid!
//
// **When you have this also put it in the Slack channel that you'll make in the next step! I may not take the first idea. The channel with me will serve as a means to communicate with each other if you so choose as well as me!**
// Our project idea is called Digital Passport. This application allows people to post tourist pictures and the location in which the pictures were taken. Thus, other users can look up a desired travel location and see and comment on other people's pictures from that place. It is a great way for users to do vacation research and see the places they can visit and things they can do in a travel destination.
// ## Once you have that come up with a team/company name! Make a slack channel with all the members of your team and <NAME>!
//
//
//
// Phoenix Foundation.
// ## How often are we meeting for Stand ups?
//
//
// Every Monday, Wednesday, and Friday.
// ## What is our iteration length? We're all locked into 2 week iterations (This ones a gimmie 😂)
//
//
// 2 week iteration length.
// ## What is our preferred means of communication? (Slack, Email, Text, Zoom call?)
//
//
// We prefferably use Discord to communicate.
// ## What means of communication do we want for Stand Ups?
//
//
// Discord.
// ## What means of communication do we want for Iteration Planning?
//
//
// A Discord voice channel.
// ## What means of communication do we want for Retrospectives?
//
//
// Discord voice channels.
// # You need to have your project idea settled before you answer these!
// ## Come up with your design specifications! What is your 1.0 release supposed to look like?
//
//
// Version 1.0 will be focused around a map that the user will be able to zoom in and out of and pan around to find a location they are looking for. They can click in a general metropolitan area, where they will be directed to a page for them to scroll through the posts from that location. Users will be able to make an account, which will let them comment on others' posts and save posts to a personal collection for vacation planning purposes.
// ## Come up with a way to manage and create an initial backlog of things to do. Put the link in your Slack Channel's description as well as here!
//
//
// Trello will be used to manage our backlog of things to do. Here is the link: https://trello.com/invite/b/VubnuuNs/d1ec23b84e048a4892acef27bc7e7b03/sd2-final-project-development-flow
// ## What's everyone's starting role (Choose an IM and PO everyone else is Dev)
// IM: <NAME>;
// PO: <NAME>;
// Dev: <NAME> & <NAME>
| Labs/Lab 4/Lab 4 SUBMISSION.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neural regression
# The point of this experiment is to see if the numbers randomly generated by a human being (kept in the file numbers.py) can be predicted using mathematics and various regression technique
import numpy as np
from matplotlib import pyplot
from datetime import datetime as dt
import json
# ### Visualizing the numbers
from data.numbers import numbers
pyplot.plot(numbers)
pyplot.title('Number of entries: {0}. Range: {1}-{2}'.format(*[len(numbers), min(numbers), max(numbers)]))
# ### Specify the parameters
# +
train_endpoint = int(len(numbers)*0.7) + 1 # Using 70% for training
x_train = np.array(range(1, train_endpoint+1))
y_train = np.array(numbers[:train_endpoint])
x_test = np.array(range(train_endpoint, len(numbers)))
y_test = np.array(numbers[train_endpoint:])
tested_degrees = range(1, 101) # Use polynomials with degrees from 2 to 100
output_dir = f'data/{dt.now()}.txt'
polydata_dir = 'data/polydata.json'
min_effectiveness = 0.5 # Any polynomial that performs equally or better than this value will be saved
soft_performance_bound = 1
print('Length of training dataset - {0}\nLength of testing dataset - {1}'.format(*[len(y_train), len(y_test)]))
# -
# ### Define functions to periodically save progress
def log(string, use_separators=True):
with open(output_dir%dt.now(), 'a+') as file:
separators = '=' * 70 if use_separators else ''
file.write(string + '\n' + separators + '\n')
def save_polydata(degree, coefficients):
with open(polydata_dir, 'w+') as file:
jsn = dict(json.load(file))
if degree in list(jsn.keys()):
print(f'There is already an entry for the degree {degree}. Overwriting...')
jsn[degree] = coefficients
json.save(jsn, file)
def clear_file():
open(output_dir, 'w+').close()
print('File cleared\n')
# +
clear_file()
total_start_time = dt.now()
performed_poorly = dict()
performed_sufficiently = dict()
print(f'Starting time - {total_start_time}\n')
log(f'Starting time - {total_start_time}', False)
for degree in tested_degrees:
start_time = dt.now()
print('='*5,'Using polynomial with degree ',degree,'='*5)
log(f'\t[{dt.now()}] Using polynomial with degree {degree}')
model = np.polynomial.polynomial.Polynomial.fit(x_train, y_train, deg=degree)
poly = np.poly1d(model.convert().coef)
strict_matches = 0 # Strict performance measures the number of cases when the model predicted the number exactly (1=1, 2=2)
soft_matches = 0 # Soft performance measures the number of cases when the model predicted the number NOT EXACTLY, but within allowed bounds
total_cases = 0
for (x, y) in zip(x_test, y_test):
total_cases += 1
result = poly(x)
if result == y:
strict_matches += 1
elif abs(result - y) <= soft_performance_bound:
soft_matches += 1
strict_performance = ( strict_matches / total_cases ) * 100
soft_performance = ( soft_matches / total_cases ) * 100
if any([strict_performance >= min_effectiveness, soft_performance >= min_effectiveness]):
performed_sufficiently[degree] = {'strict_performance': strict_performance, 'soft_performance': soft_performance}
print(f'\tModel with degree {degree} performes well enough:\n\t\tStrict performance: {strict_performance}%\n\t\tSoft performance: {soft_performance}%')
save_polydata(degree, poly)
log(f'\tModel with degree {degree} performes well enough:\n\t\tStrict performance: {strict_performance}%\n\t\tSoft performance: {soft_performance}%')
else:
print('\tModel performes poorly\n')
end_time = dt.now()
elapsed_time = end_time - start_time
print(f'Elapsed: {elapsed_time}\n')
| Experiment.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.1
# language: julia
# name: julia-1.5
# ---
# # Simulating the Atomic Bomb
# ## Packages
# +
#using Pkg
#pkg"add SimJulia"
#pkg"add Distributions"
#pkg"add StatsPlots"
using SimJulia
using Distributions
using Plots
using StatsPlots
using CSV
using Logging
# -
# ## Constants
const Nₐ = 6.02214086e23 # atoms / mole
const ρᵤ = 19.1 # g / cm3
const mᵤ = 235.0439299 # g / mole
const nᵤ = ρᵤ * Nₐ / mᵤ # atoms / cm3
const mₙ = 1.008664916 # g / mole
const Mₙ = mₙ / Nₐ * 1e-3 # kg
const k = 1.38064852e-23 # J / K
const q = 1.60217662e-19 # C
const A = mᵤ / mₙ
const α = (A - 1)^2 / (A + 1) ^2
const numberofspontaneousfis = 0.0003; # / g / s
ρᵤ * 4/3 * π * 9^3 * numberofspontaneousfis
# ## Distributions
# +
const cosΘdistr = Uniform(-1, 1)
const cosϕdistr = Uniform(-1, 1)
const energy = 1e-3:1e-3:15
function wattspectrum(energy) # MeV
0.453 * sinh(sqrt(2.29*energy))*exp(-1.036*energy)
end
const spectrum = wattspectrum.(energy)
const wattdistr = Categorical(spectrum ./ sum(spectrum))
const numberofneutronsdistr = Categorical([0,0.6,0.36,0.04])
const numberofneutronsspontaneousdistr = Categorical([0.2,0.74,0.06]);
# -
# ## Data
# +
σt = CSV.read("sigma_total.txt")
σf = CSV.read("sigma_fission.txt")
σa = CSV.read("sigma_absorption.txt")
σi = CSV.read("sigma_inelastic.txt")
function Σ(energy::Float64) # 1 / cm
i = findfirst(e -> e > energy, σt[:, 1])
σ = σt[i, 2] + (energy - σt[i, 1]) / (σt[i-1, 1] - σt[i, 1]) * (σt[i-1, 2] - σt[i, 2])
nᵤ * σ * 1e-24
end
function ΔtΔl(energy::Float64)
Δl = -log(rand()) / Σ(energy)
v = sqrt(2 * energy * q / Mₙ) * 100
Δl / v, Δl
end;
# -
# ## Types
# +
struct Bomb
radius :: Float64 # cm
generated :: Vector{Int64}
neutrons :: Vector{Int64}
times :: Vector{Float64} # s
function Bomb(radius::Real)
new(radius, Float64[], Int64[], Float64[])
end
end
mutable struct Neutron
r :: Float64 # cm
cosθ :: Float64
energy :: Float64 # eV
function Neutron(r::Float64, energy::Float64, cosθ::Float64 = rand(cosΘdistr))
new(r, cosθ, energy)
end
end
function Neutron(sim::Simulation, bomb::Bomb, r::Float64, energy::Float64=energy[rand(wattdistr)] * 1e6)
neutron = Neutron(r, energy)
time = now(sim)
@info("$time: create neutron at position $r with cosθ = $(neutron.cosθ) and energy = $(neutron.energy) eV")
push!(bomb.times, time)
push!(bomb.neutrons, 1)
Δt, Δl = ΔtΔl(neutron.energy)
@callback collision(timeout(sim, Δt), bomb, neutron, Δl)
end;
# -
# ## Callback
# +
function spontaneousfission(ev::AbstractEvent, bomb::Bomb)
sim = environment(ev)
for _ in rand(numberofneutronsspontaneousdistr)
Neutron(sim, bomb, rand() * bomb.radius)
end
rate = ρᵤ * 4/3 * π * bomb.radius^3 * numberofspontaneousfis
@callback spontaneousfission(timeout(sim, -log(rand()) / rate), bomb)
end
function collision(ev::AbstractEvent, bomb::Bomb, neutron::Neutron, Δl::Float64)
sim = environment(ev)
time = now(ev)
r′ = sqrt(neutron.r^2 + Δl^2 + 2*neutron.r*Δl*neutron.cosθ)
if r′ > bomb.radius
@info("$(now(sim)): neutron has left the bomb")
push!(bomb.times, time)
push!(bomb.neutrons, -1)
push!(bomb.generated, 0)
else
i = findfirst(e -> e > neutron.energy, σt[:, 1])
σtot = σt[i, 2] + (neutron.energy - σt[i, 1]) / (σt[i-1, 1] - σt[i, 1]) * (σt[i-1, 2] - σt[i, 2])
i = findfirst(e -> e > neutron.energy, σf[:, 1])
σfis = σf[i, 2] + (neutron.energy - σf[i, 1]) / (σf[i-1, 1] - σf[i, 1]) * (σf[i-1, 2] - σf[i, 2])
i = findfirst(e -> e > neutron.energy, σa[:, 1])
σabs = σa[i, 2] + (neutron.energy - σa[i, 1]) / (σa[i-1, 1] - σa[i, 1]) * (σa[i-1, 2] - σa[i, 2])
i = findfirst(e -> e > neutron.energy, σi[:, 1])
i = i == 1 ? 2 : i
σin = σi[i, 2] + (neutron.energy - σi[i, 1]) / (σi[i-1, 1] - σi[i, 1]) * (σi[i-1, 2] - σi[i, 2])
rnd = rand()
if rnd < σfis / σtot
n = rand(numberofneutronsdistr)
@info("$(now(sim)): fission with creation of $n neutrons")
for _ in 1:n
Neutron(sim, bomb, r′)
end
push!(bomb.times, time)
push!(bomb.neutrons, -1)
push!(bomb.generated, n)
elseif rnd < (σabs + σfis) / σtot
@info("$(now(sim)): neutron absorbed")
push!(bomb.times, time)
push!(bomb.neutrons, -1)
push!(bomb.generated, 0)
elseif rnd < (σin + σabs + σfis) / σtot
@info("$(now(sim)): inelastic scattering")
n = 1
Neutron(sim, bomb, r′)
push!(bomb.times, time)
push!(bomb.neutrons, -1)
else
cosϕ = rand(cosϕdistr)
cosψ = (A * cosϕ + 1) / sqrt(A^2 + 2 * A * cosϕ +1)
neutron.r = r′
neutron.energy *= 0.5 * (1 + α + (1 - α) * cosϕ)
θ = acos(neutron.cosθ)
ψ = acos(cosψ)
θplusψ = θ + ψ
θminψ = ψ < π / 2 ? θ - ψ : θ - ψ + 2π
neutron.cosθ = cos(θplusψ + rand() * (θminψ - θplusψ))
@info("$(now(sim)): elastic scattering at position $r′ with cosθ = $(neutron.cosθ) and energy = $(neutron.energy) eV")
Δt, Δl = ΔtΔl(neutron.energy)
@callback collision(timeout(sim, Δt), bomb, neutron, Δl)
end
end
((sum(bomb.generated) > 500 && sum(bomb.neutrons) == 0) || (time > 1 && sum(bomb.neutrons) == 0) || sum(bomb.generated) > 1000) && throw(StopSimulation())
end
# -
# ## Simulation
sim = Simulation()
bomb = Bomb(8.0)
@callback spontaneousfission(timeout(sim, 0.0), bomb)
run(sim)
mean(bomb.generated)
# ## Plot
i = findlast(x->x==0, cumsum(bomb.neutrons))
i = i === nothing ? 1 : i
plot(bomb.times[i+1:end], cumsum(bomb.neutrons)[i+1:end], seriestype=:scatter, ylabel="N", xlabel="time [s]")
#plot(bomb.times, cumsum(bomb.neutrons), seriestype=:scatter, ylabel="N", xlabel="time [s]")
# ## Monte Carlo
const RUNS = 100
const RADII = 5:12;
Logging.disable_logging(LogLevel(1000));
ks = zeros(Float64, RUNS, length(RADII))
for (i, r) in enumerate(RADII)
for j in 1:RUNS
sim = Simulation()
bomb = Bomb(r)
@callback spontaneousfission(timeout(sim, 0.0), bomb)
run(sim)
ks[j, i] = mean(bomb.generated)
end
end
boxplot(reshape(collect(RADII), 1, length(RADII)), ks, label=reshape(collect(RADII), 1, length(RADII)), legend=:bottomright, xlabel="R [cm]", ylabel="k")
mean(ks, dims=1)
plot(RADII, [mean(ks, dims=1) ...], seriestype=:scatter, xlabel="R [cm]", ylabel="k")
| Lectures_old/Lecture 12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Choroplethmapbox animation
# We define an animated Choroplethmapbox, where each frame consists in plotting only two counties from Switzerland.
import numpy as np
import json
import plotly.graph_objs as go
# +
import urllib.request
def read_geojson(url):
with urllib.request.urlopen(url) as url:
jdata = json.loads(url.read().decode())
return jdata
# -
swiss_url = 'https://raw.githubusercontent.com/empet/Datasets/master/swiss-cantons.geojson'
jdata = read_geojson(swiss_url)
# +
import pandas as pd
data_url = "https://raw.githubusercontent.com/empet/Datasets/master/Swiss-synthetic-data.csv"
df = pd.read_csv(data_url)
df.head()
# -
len(jdata['features']), len(df)
# Define a list of geojson type dicts (its elements would correspond to geojson files for different countries in a continent, when
# the animation is intended for that case):
geojs =[]
for k in range(0, 26, 2):
geojs.append(dict(type='FeatureCollection',
features = jdata['features'][k:k+2]))
geojs[5].keys()
# +
import plotly.express as px
fig = px.choropleth_mapbox(df[:2],
geojson=geojs[0],
featureidkey='properties.id',
locations='canton-id',
color='2018',
color_continuous_scale ='matter_r',
zoom=6.25,
center={'lat': 46.8181877 , 'lon':8.2275124 },
mapbox_style='carto-positron')
fig.update(frames = fig.frames, layout = fig.layout)
fig.update_layout(title_text='Choroplethmapbox animation',
title_x=0.5,
width=900, height=500,
coloraxis_cmin=min(df['2018']),
coloraxis_cmax=max(df['2018']),
coloraxis_colorbar_thickness=25);
# -
# Below we are selecting from the df a new dataframe consisting in only two rows and one column from the general dataframe, df:
k =3
df.iloc[k:k+2, [1]]
df.iloc[k:k+2, [1]]['canton-id']
frames = [go.Frame(
data=[go.Choroplethmapbox(geojson=geojs[k//2],
locations=df.iloc[k:k+2, [1]]['canton-id'], # dataframe consisting in rows k:k+2 from df['canton-id'],
z=df.iloc[k:k+2, [2]]['2018'], # rows k:k+2 from df['2018']
)],
name=f'fr{k//2}') for k in range(0, len(df), 2)]
fig.update(frames=frames);
f_duration =350 #frame duration
fig.update_layout(updatemenus=[dict(type='buttons',
y=0,
x=1.25,
active=0,
buttons=[dict(label='Play',
method='animate',
args=[None,
dict(frame=dict(duration=f_duration,
redraw=True),
transition=dict(duration=0),
fromcurrent=True,
mode='immediate')])])],
sliders = [dict(steps= [dict(method= 'animate',
args= [[ f'fr{k}'],
dict(mode= 'immediate',
frame= dict( duration=f_duration, redraw= True ),
fromcurrent=True,
transition=dict( duration= 0))],
label=f"fr{k}") for k in range(len(frames))],
minorticklen=0,
x=0,
len=1)]);
fig.show()
import chart_studio.plotly as py
py.plot(fig, filename='choroplethmbx-anim')
# **Remarks**:
# **1.** This method of decomposing the initial geojson file into many geojson type dicts has the advantage that requires less
# data to be involved in the animation.
# Another method could be to set `geojson=jdata` in the definition of `fig= px.choropleth_mapbox()`,
# and pass to each frame data only `locations` and `z`. In this case each frame inherits
# the other attributes set in `fig.data[0]`, i.e.
# for each frame is used the entire jdata file. Obviously this method is not recommended for large geojson files read as jdata.
#
# **2.** If `fig.data[0]` is defined directly as an instance of the class `go.Choroplethmapbox`, not via `px.choroplethmapbox()`, then to ensure the correct mapping of z-values in each frame, to the associated colorscale, the attributes `zmin`, `zmax` must be set, as can be seen below (note that for `go.Choroplethmapbox` we have `zmin`, `zmax` as attributes setting the least and the biggest value to be mapped to a colorscale, while using `px.choropleth_mapbox()', the same effect has setting
# coloraxis['cmin'], coloraxis['cmax'], and these differences of names can be a bit confusing.
# Alternative definition of fig, defined above by `px.choropleth_mapbox()`:
# +
fig = go.Figure(go.Choroplethmapbox(geojson=geojs[0],
locations=df.iloc[0:2 , [1]]['canton-id'],
z=df.iloc[0:2 , [2]]['2018'],
featureidkey='properties.id',
colorscale='matter_r',
colorbar_thickness=25,
zmin=min(df['2018']),
zmax=max(df['2018']),
marker_line_width=1))
fig.update_layout(title_text = 'Choroplethmapbox animation',
title_x=0.5,
width=800, height=500,
mapbox=dict(style='carto-positron',
zoom=6.25,
center = {"lat": 46.8181877 , "lon":8.2275124 },
))
# -
| Choroplethmapbox-animation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MDN Hands On Tutorial
#
# This notebook demonstrates the construction of a simple MDN, and compares it to a regular neural network.
#
# Read about MDNs on the [original paper](https://publications.aston.ac.uk/373/1/NCRG_94_004.pdf) by <NAME>.
# > This revision has been adapted to work with Tensorflow 2. Note that `tensorflow_probability` has been moved from the main code to it's own package (`pip install tensorflow_probability`)
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import seaborn as sns
import tensorflow as tf
import tensorflow_probability as tfp
import matplotlib.pyplot as plt
from sklearn.utils import shuffle
from tensorflow import keras
from keras import optimizers
from keras.models import Sequential, Model
from keras.layers import Dense, Activation, Layer, Input
# The network we'll construct will try to learn the following relation between $x$ and $f(x)$:
#
# $$f(x) = x^2-6x+9$$
#
#
# Note that this simply $y = x^2$ shifted three steps to the left (global minimum is at $x=3$).
def f(x):
return x**2-6*x+9
# In order to make the data a little bit more realistic, we'll add a normally-distributed noise, which will be location-dependent - the larger $x$ is, the larger the noisier the data will be. So, our data generator will obey the following relation:
#
# $$g(x) = f(x) + \epsilon(x) $$
#
# $$ \text{where}: \epsilon(x) = N(0,\sigma_0 x)$$
#
# Where $N(\mu,\sigma)$ is the normal distribution with mean $\mu$ and STD of $\sigma$.
#
# The `data_generator` below function creates $n$ nosiy data samples for a given `x`, where $n$ is defined by `samples`. Notice that technically, `data_generator` yields $g(x) = N(f(x),\sigma_0 x)$, as mathematically that's the same thing.
def data_generator(x,sigma_0,samples):
return np.random.normal(f(x),sigma_0*x,samples)
# We'll now generate our dataset for $1<x<5$.
#
# The purple line in the plot presents the "clean" function $f(x)$ for this range.
# +
sigma_0 = 0.1
x_vals = np.arange(1,5.2,0.2)
x_arr = np.array([])
y_arr = np.array([])
samples = 50
for x in x_vals:
x_arr = np.append(x_arr, np.full(samples,x))
y_arr = np.append(y_arr, data_generator(x,sigma_0,samples))
x_arr, y_arr = shuffle(x_arr, y_arr)
x_test = np.arange(1.1,5.1,0.2)
fig, ax = plt.subplots(figsize=(10,10))
plt.grid(True)
plt.xlabel('x')
plt.ylabel('g(x)')
ax.scatter(x_arr,y_arr,label='sampled data')
ax.plot(x_vals,list(map(f,x_vals)),c='m',label='f(x)')
ax.legend(loc='upper center',fontsize='large',shadow=True)
plt.show()
# -
# ## Regular neural network
# We'll now train a neural network which will receive $x$ as input and our noisy $g(x)$ but will have to learn the relation $x \rightarrow f(x)$.
#
# The network is constructed of two hidden layers, each with 12 nodes and the $\tanh(x)$ activation function (note we use a linear activation on the last output layer which is the same as no activation function at all, since keras defaults to sigmoid if none is given for Dense layers).
#
# We set the learning rate $\alpha=0.0003$, 50 examples per mini-batch and a total of 500 epoches.
epochs = 500
batch_size = 50
learning_rate = 0.0003
model = Sequential()
model.add(Dense(12,input_shape=(1,),activation="tanh"))
model.add(Dense(12,activation="tanh"))
model.add(Dense(1,activation="linear"))
adamOptimizer = optimizers.Adam(learning_rate=learning_rate)
model.compile(loss='mse',optimizer=adamOptimizer,metrics=['mse'])
history_cache = model.fit(x_arr,
y_arr,
verbose=0, # write =1 if you wish to see the progress for each epoch
epochs=epochs,
batch_size=batch_size)
y_pred = model.predict(x_test)
fig, ax = plt.subplots(figsize=(10,10))
plt.grid(True)
plt.xlabel('x')
plt.ylabel('y')
ax.scatter(x_arr,y_arr,c='b',label='sampled data')
ax.scatter(x_test,y_pred,c='r',label='predicted values')
ax.plot(x_vals,list(map(f,x_vals)),c='m',label='f(x)')
ax.legend(loc='upper center',fontsize='large',shadow=True)
plt.show()
print('Final cost: {0:.4f}'.format(history_cache.history['mse'][-1]))
# It seems to be doing quite good in predicting $f(x)$, but we can clearly see that the network learnt nothing about the size of the noise.
#
# ## Mixture density network (MDN)
# Let's try an MDN now. We'll use the same network as in the previous section, with one important change:
# the output layer now has two nodes (which are constructed as two layers of 1 node for technical simplicity), which we named `mu` and `sigma`
#
# Note the new cost function: we create a normal distribution out of the predicted `mu` and `sigma`, and then minimize the negative log-likelihood of this distribution yielding the target value `y`. Mathematically, our cost function is the negative logarithm of the normal distribution's probability density function (PDF):
#
# $$Cost = -\log (PDF) = -\log\left(\frac{1}{\sqrt{2\pi}\sigma}\cdot\exp{\left[-\frac{(y-\mu)^{2}}{2\sigma^{2}}\right]}\right)$$
def mdn_cost(mu, sigma, y):
dist = tfp.distributions.Normal(loc=mu, scale=sigma)
return tf.reduce_mean(-dist.log_prob(y))
# We'll use `elu + 1` as the activation function for `sigma`, as it must always be non-negative. The Exponential Linear Unit (ELU) is defined as:
#
# $$ ELU(x) = \begin{cases} x & x\ge0 \\ \exp{(x)}-1 & x < 0 \end{cases} $$
epochs = 500
batch_size = 50
learning_rate = 0.0003
InputLayer = Input(shape=(1,))
Layer_1 = Dense(12,activation="tanh")(InputLayer)
Layer_2 = Dense(12,activation="tanh")(Layer_1)
mu = Dense(1, activation="linear")(Layer_2)
sigma = Dense(1, activation=lambda x: tf.nn.elu(x) + 1)(Layer_2)
y_real = Input(shape=(1,))
lossF = mdn_cost(mu,sigma,y_real)
model = Model(inputs=[InputLayer, y_real], outputs=[mu, sigma])
model.add_loss(lossF)
adamOptimizer = optimizers.Adam(learning_rate=learning_rate)
model.compile(optimizer=adamOptimizer,metrics=['mse'])
history_cache = model.fit([x_arr, y_arr], #notice we are using an input to pass the real values due to the inner workings of keras
verbose=0, # write =1 if you wish to see the progress for each epoch
epochs=epochs,
batch_size=batch_size)
print('Final cost: {0:.4f}'.format(history_cache.history['loss'][-1]))
mu_pred, sigma_pred = model.predict(list((x_test,x_test))) # the model expects a list of arrays as it has 2 inputs
fig, ax = plt.subplots(figsize=(10,10))
plt.grid(True)
plt.xlabel('x')
plt.ylabel('y')
ax.errorbar(x_test,mu_pred,yerr=np.absolute(sigma_pred),c='r',ls='None',marker='.',ms=10,label='predicted distributions')
ax.scatter(x_arr,y_arr,c='b',alpha=0.05,label='sampled data')
ax.errorbar(x_vals,list(map(f,x_vals)),yerr=list(map(lambda x: sigma_0*x,x_vals)),c='b',lw=2,ls='None',marker='.',ms=10,label='true distributions')
ax.plot(x_vals,list(map(f,x_vals)),c='m',label='f(x)')
ax.legend(loc='upper center',fontsize='large',shadow=True)
plt.show()
# The plot above shows the results learnt by the network. In red are the networks predictions for 𝜇
# and 𝜎, and in blue are the actual 𝜇 and 𝜎 used for the training set. The actual data can be seen faded in the background. We can clearly see the network has learnt not just 𝑥→𝑓(𝑥), but also the noise creating 𝑥→𝑔(𝑥).
| mdn-tf2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + code_folding=[]
# just imports
# %load_ext autoreload
# %autoreload 2
import sys
import time
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
from torch.utils import data
from torch import utils
from torch.optim import Adam
from torchvision import transforms
from torchvision import datasets
import numpy as np
from matplotlib import pyplot as plt
from pandas import read_fwf, DataFrame
from tqdm import tqdm_notebook as tqdm
import matplotlib.gridspec as gridspec
from scipy.ndimage.interpolation import rotate
from sklearn.model_selection import train_test_split
# + code_folding=[]
# local imports
sys.path.append('../')
from VAE.rg_dataset import LRG, BalancedDataSetLRG
from VAE.loss_funcs import VAE_Loss
from VAE import vae_models
import pickle
# + code_folding=[]
# %%time
data_path = '../data/'
aug=1
lrg_datasets = pickle.load( open( "lrg.p", "rb" ) )
# +
normal_data_loader = utils.data.DataLoader(lrg_datasets['full'], batch_size=128, shuffle=False)
data_loader_lrg = utils.data.DataLoader(lrg_datasets['train'], batch_size=128, shuffle=True)
# -
lbs = np.array(lrg_data_set.labels)
lbs = lbs[lbs > 0]
lbs = lbs[lbs>2]
np.sum(lbs > 3)
# len(lrg_data_set.labels)
# set(lbs)
# + code_folding=[1, 54, 59, 66, 71]
class VAE(nn.Module):
def __init__(self, lt_dim=4, k=None, batch_norm=True):
super(VAE, self).__init__()
self.k = k
n_layers = len(self.k)
encoder_layers = []
decoder_layers = []
for i in range( n_layers -1) :
in_c, out_c = self.k[i], self.k[i + 1]
if(in_c == 'M'): continue
stride = 1
if out_c == 'M':
stride = 2
i += 1
out_c = self.k[i + 1]
layer = nn.Conv2d(in_c, out_c, kernel_size=3, padding=1, stride=stride)
encoder_layers.append(layer)
if batch_norm:
encoder_layers.append(nn.BatchNorm2d(out_c))
encoder_layers.append(nn.ReLU(inplace=True))
self.encoder = nn.Sequential(*encoder_layers)
for i in range(n_layers - 1, 0, -1):
in_c, out_c = self.k[i], self.k[i - 1]
if(in_c == 'M'): continue
stride = 1
output_padding=0
if out_c == 'M':
stride = 2
i -= 1
out_c = self.k[i - 1]
output_padding=1
layer = nn.ConvTranspose2d(in_c, out_c, kernel_size=3, padding=1,
output_padding=output_padding, stride=stride)
decoder_layers.append(layer)
if batch_norm:
decoder_layers.append(nn.BatchNorm2d(out_c))
decoder_layers.append(nn.ReLU(inplace=True))
self.decoder = nn.Sequential(*decoder_layers[:-1])
self.fc_mu = nn.Sequential(
nn.Linear(self.k[-1]*2*2, lt_dim*2),
nn.Linear(lt_dim*2, lt_dim)
)
self.fc_ep = nn.Sequential(
nn.Linear(self.k[-1]*2*2, lt_dim*2),
nn.Linear(lt_dim*2, lt_dim)
)
self.fc_dc = nn.Linear(lt_dim, self.k[-1]*2*2)
def encode(self, x):
encoded = self.encoder(x)
encoded = encoded.view(-1, self.k[-1]*2*2)
return self.fc_mu(encoded), self.fc_ep(encoded)
def reparameterize(self, mu, logvar):
std = torch.exp(0.5*logvar)
eps = torch.randn_like(std)
if self.training :
return mu + eps*std
return mu
def decode(self, x):
x = F.relu(self.fc_dc(x))
x = x.view(-1, self.k[-1], 2, 2) #reshape
return torch.sigmoid(self.decoder(x))
def forward(self, x):
mu, var = self.encode(x)
z = self.reparameterize(mu, var)
d = self.decode(z)
return d, mu, var
# -
model = torch.load('SimpleBVAE_Class_all')
sample = iter(normal_data_loader).next()
s = sample[0][1:2]
with torch.no_grad():
e = model.encode(s.to('cuda'))[0]
d = model.decode(e).to('cpu')
f, ax = plt.subplots(1, 2, figsize=(10, 5))
ax[0].set_title('Original')
ax[1].set_title('Reconstruida')
ax[0].imshow(s[0][0], cmap='gray')
ax[1].imshow(d[0][0], cmap='gray')
ax[0].axis('off')
ax[1].axis('off')
plt.show()
# +
# [1.5, 2, 2, 1, 8, 2, 1, 3]
j = 7
m = 1.5
ar = np.arange(-2, 2.5, .5)
f, ax = plt.subplots(1, len(ar), figsize=(20, 9))
for k, i in enumerate(ar):
b = torch.tensor(e)
b[0][j] = e[0][j] + m*i
with torch.no_grad():
d = model.decode(b).cpu()[0][0]
ax[k].imshow(d, cmap='gray')
ax[k].set_aspect('equal')
ax[k].axis('off')
# +
fig = plt.figure(figsize=(14, 13))
gs1 = gridspec.GridSpec(8, 9, figure=fig)
gs1.update(wspace=0.02, hspace=0.02) # set the spacing between axes.
m_vecotr = [.75, 1, 1, .5, 4, 1, .5, 1.5]
for j in range(8):
for k, i in enumerate(np.arange(-2,2.5,.5)):
ax1 = plt.subplot(gs1[j*9+k])
plt.axis('off')
b = torch.tensor(e)
b[0][j] = e[0][j] + m_vecotr[j]*i
with torch.no_grad():
d = model.decode(b).cpu()[0][0]
ax1.imshow(d, cmap='gray')
ax1.set_aspect('equal')
plt.show()
# -
model.eval()
enc_vals = []
labels = []
j = 12
with torch.no_grad():
for i, (data, target) in enumerate(data_loader_lrg):
e = model.encode(data.to('cuda'))[0]
enc_vals += (e.to('cpu').tolist())
labels += target.tolist()
j -= 1
# if j == 0:
# break
enc_vals = np.array(enc_vals)
print(len(labels))
Y = np.array(labels)
X = enc_vals
Y = np.array(labels)
from sklearn.manifold import TSNE
X_embedded = TSNE(n_components=2).fit_transform(X)
plt.figure(figsize=(15,10))
plt.scatter(X_embedded[:,0], X_embedded[:,1], c=Y,alpha=0.8)
plt.show()
Y2 = Y[Y > 0]
X_embedded2 = X_embedded[Y > 0]
plt.figure(figsize=(15,10))
plt.scatter(X_embedded2[:,0], X_embedded2[:,1], c = Y2 < 3)
plt.show()
Y3 = Y2[Y2 < 3]
X_embedded3 = X_embedded2[Y2 < 3]
plt.figure(figsize=(15,10))
plt.scatter(X_embedded3[:,0], X_embedded3[:,1], c = Y3 == 1)
plt.show()
np.arange(-2,2.5,.5)
| notebooks/TestBVAE_Class_TSNE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Working with snapshots
#
# Here, the following topics are going to be covered:
#
# - What is a snapshot
# - How to create it
# - How it is saved next to the measurement data
# - How to extract snapshot from the dataset
# ### Useful imports
# +
from pprint import pprint # for pretty-printing python variables like 'dict'
import json # for converting JSON data into python 'dict'
import qcodes
from qcodes import Parameter, Station, \
initialise_database, \
new_experiment, Measurement
from qcodes.tests.instrument_mocks import DummyInstrument
# -
# ## What is a snapshot
#
# Often times experiments comprise a complex network of interconnected instruments. Their numerous settings define the overall behavior of the experimental setup. Obviously, the experimental setup has a direct impact on the measured data which is of prime interest for researchers. In order to capture this link, the measured data should have metadata associated with it. An important part of that metadata is a captured state of the experimental setup. In QCoDeS terms, this is called snapshot.
# ## How to create a snapshot
#
# All QCoDeS instruments and parameters (and some other objects too, like `InstrumentChannel`s) support snapshotting, which means that they provide a method to retrieve their state.
#
# Let's look at snapshots of various objects.
# ### Snapshot example for Parameter object
# Let's create a `Parameter`, call its `snapshot` method, and then inspect the output of that method.
#
# The returned snapshot is a python dictionary that reflects all the important properties of that parameter (check the name, label, unit, and even value).
# +
p = Parameter('p', label='Parameter P', unit='kg', set_cmd=None, get_cmd=None)
p.set(123)
snapshot_of_p = p.snapshot()
pprint(snapshot_of_p)
# -
# In case you want to use the snapshot object in your code, you can refer to its contents in the same way as you work with python dictionaries, for example:
print(f"Value of {snapshot_of_p['label']} was {snapshot_of_p['value']} (when it was snapshotted).")
# Note that the implementation of a particular QCoDeS object defines which attributes are snapshotted and how. For example, `Parameter` implements a keyword argument `snapshot_value` which allows to choose if the value of the parameter is snapshotted (the reasons for this are out of scope of this article). (Another interesting keyword argument of `Parameter` that is realated to snapshotting is `snapshot_get` - refer to `Parameters`'s docstring for more information.)
# Below is a demonstration of the `snapshot_value` keyword argument, notice that the value of the parameter is not part of the snapshot.
# +
q = Parameter('q', label='Parameter Q', unit='A', snapshot_value=False, set_cmd=None, get_cmd=None)
p.set(456)
snapshot_of_q = q.snapshot()
pprint(snapshot_of_q)
# -
# ### Snapshot of an Instrument
#
# Now let's have a brief look at snapshots of instruments. For the sake of exercise, we are going to use a "dummy" instrument.
# A dummy instrument with two parameters, "input" and "output", plus a third one we'll use later.
instr = DummyInstrument('instr', gates=['input', 'output', 'gain'])
instr.gain(11)
# +
snapshot_of_instr = instr.snapshot()
pprint(snapshot_of_instr, indent=4)
# -
# ### Station and its snapshot
#
# Experimental setups are large, and instruments tend to be quite complex in that they comprise many parameters and other stateful parts. It would be very time-consuming for the user to manually go through every instrument and parameter, and collect the snapshot data.
#
# Here is where the concept of station comes into play. Instruments, parameters, and other submodules can be added to a [Station object](../Station.ipynb) ([nbviewer.jupyter.org link](https://nbviewer.jupyter.org/github/QCoDeS/Qcodes/tree/master/docs/examples/Station.ipynb)). In turn, the station has its `snapshot` method that allows to create a collective, single snapshot of all the instruments, parameters, and submodules.
#
# Note that in this article the focus is on the snapshot feature of the QCoDeS `Station`, while it has some other features (also some legacy once).
# Let's create a station, and add a parameter, instrument, and submodule to it. Then we will print the snapshot. Notice that the station is aware of insturments and stand-alone parameters, and classifies them into dedicated lists within the snapshot.
# +
station = Station()
station.add_component(p)
station.add_component(instr)
# Note that it is also possible to add components
# to a station via arguments of its constructor, like this:
# station = Station(p, instr)
# +
snapshot_of_station = station.snapshot()
pprint(snapshot_of_station)
# -
# ## Saving snapshot next to the measurement data
#
# With the power of the station object, it is now possible to conveniently associate the snapshot information with the measured data.
#
# In order to do so, a station needs to be created, and then that station needs to be provided to the `Measurement` object. If no station is explicitly provided, the `Measurement` object will use the default station, `Station.default` (refer to `Measurement` and `Station` objects docstrings for more information). At the moment the new measurement run is started, a snapshot of the whole station will be taken, and added next to the measured data.
#
# Note that the snapshot gets stored in a JSON format (an automatic convertion from python dictionary to JSON takes place). This is done in order to ensure that the snapshot can be read in environments other than python. JSON is an extemely popular data format, and all platforms/environments/languages/frameworks have means to read JSON-formatted data.
# Here is how it looks in the code. We will create a new experiment. Then we are going to reuse the station object created above, and create a new measurement object. Then we will perform a dummy measurement. After that we are going to extract the snapshot from the resulting dataset, and print it.
# +
# Let's initialize a database to ensure that it exists
initialise_database()
# Let's create a new experiment
experiment = new_experiment('snapshot_experiment', 'no_sample_yet')
# +
measurement = Measurement(experiment, station)
measurement.register_parameter(instr.input)
measurement.register_parameter(instr.output, setpoints=[instr.input])
# +
with measurement.run() as data_saver:
input_value = 111
instr.input.set(input_value)
instr.output.set(222) # assuming that the instrument measured this value on the output
data_saver.add_result((instr.input, input_value),
(instr.output, instr.output()))
# For convenience, let's work with the dataset object directly
dataset = data_saver.dataset
# -
# ## Extracting snapshot from dataset
# Now we have a dataset that contains data from the measurement run. It also contains the snapshot.
#
# In order to access the snapshot, use the `DataSet`'s properties called `snapshot` and `snapshot_raw`. As their docstrings declare, the former returns the snapshot of the run as a python dictionary, while the latter returns it as JSON string (in other words, in exactly the same format as it is stored in the experiments database).
snapshot_of_run = dataset.snapshot
snapshot_of_run_in_json_format = dataset.snapshot_raw
# To prove that these snapshots are the same, use `json.loads` or `json.dumps` to assert the values of the variables:
assert json.loads(snapshot_of_run_in_json_format) == snapshot_of_run
assert json.dumps(snapshot_of_run) == snapshot_of_run_in_json_format
# Finally, let's pretty-print the snapshot. Notice that the values of the `input` and `output` parameters of the `instr` instrument have `0`s as values, and not `111` and `222` that were set during the measurement run.
pprint(snapshot_of_run)
# Note that the snapshot that we have just loaded from the dataset is almost the same as the snapshot that we directly obtained from the station above. The only difference is that the snapshot loaded from the dataset has a top-level `station` field. If you do not trust me, have a look at the following `assert` statement for the proof.
assert {'station': snapshot_of_station} == snapshot_of_run
# ## Comparing how two DataSets were taken
# Suppose something went wrong in an experiment, and you'd like to compare what changed since a known-good run.
# QCoDeS lets you do this by taking a *diff* between the snapshots for two `DataSet` instances.
# +
measurement = Measurement(experiment, station)
measurement.register_parameter(instr.input)
measurement.register_parameter(instr.output, setpoints=[instr.input])
# +
instr.gain(400) # Oops!
with measurement.run() as data_saver:
input_value = 111
instr.input.set(input_value)
instr.output.set(222) # assuming that the instrument measured this value on the output
data_saver.add_result((instr.input, input_value),
(instr.output, instr.output()))
# For convenience, let's work with the dataset object directly
bad_dataset = data_saver.dataset
# -
# The `diff_param_values` function tells us about the parameters that changed betw
from qcodes.utils.metadata import diff_param_values
diff_param_values(dataset.snapshot, bad_dataset.snapshot).changed
| docs/examples/DataSet/Working with snapshots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sqlalchemy import create_engine
from matplotlib import pyplot as plt
from config import pg_password
# Creating connection string using f string to keep my postgres password hidden
connection_str = f"postgres://postgres:{pg_password}@192.168.127.12:5432/Employees"
engine = create_engine(connection_str)
# Setting variable to hold SQL query that will pull all rows from the "salaries" table
cmd = "SELECT * FROM salaries"
# Running query and converting it to a DataFrame
salaries_df = pd.read_sql(cmd, con=engine)
salaries_df
# Creating histogram to visualize the the frequency salaries are paid.
salaries_df["salary"].hist()
plt.title("Histogram of Salaries")
plt.xlabel("Salary")
plt.ylabel("Frequency")
plt.show()
# Creating variable to define the SQL query that will pull all of the data from the "titles" table.
cmd = "Select * From titles"
titles_df = pd.read_sql(cmd, con=engine)
titles_df
# Creating variable to define the SQL query that will pull all of the data from the "employees" table.
cmd = "SELECT * FROM employees"
employees_df = pd.read_sql(cmd, con=engine)
employees_df
# Merging salaries_df and employees_df so that salaies can be displayed with employees.
employee_salary_df = pd.merge(salaries_df, employees_df, on = "emp_no")
employee_salary_df
# Merging the titles_df with the employee_salary_df so that I can use the emp_title_id to include each employees title.
salaries_title_df=pd.merge(employee_salary_df, titles_df, left_on="emp_title_id", right_on="title_id")
salaries_title_df
# Grouping by title so that I can get the average salary per title.
grouped_salaries = salaries_title_df.groupby("title", as_index=False).agg({"salary": "mean"})
avg_salaries_df = grouped_salaries.round(decimals=2)
avg_salaries_df
# Creating table to visualize the average salary per job title.
plt.title("Average Salary per Job Title")
plt.xlabel("Job Title")
plt.ylabel("Average Salary")
plt.bar(
avg_salaries_df["title"], avg_salaries_df["salary"], color="orange"
)
plt.xticks(rotation=65)
plt.show()
# Here I am learning that I do not have a job after all. It was all just an April Fool's joke. I'm so disappointed.
# It's ok, though. I wouldn't take a job that only paid $40,000 a year
my_id = salaries_title_df.loc[salaries_title_df["emp_no"] == 499942]
my_id
| Employee_SQL/SQL_Challenge_Employees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.13 ('py38')
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="cz3i7oBg8pKw" outputId="f03b31c0-5fd5-4d6a-e8bd-95bb235fb9ef"
# from google.colab import drive
# drive.mount('/content/gdrive')
# + colab={"base_uri": "https://localhost:8080/"} id="4VOVI054suq2" outputId="efb88694-28ba-4243-a296-94f10561c880"
# !nvidia-smi
# + colab={"base_uri": "https://localhost:8080/"} id="CrCji_SWtMLL" outputId="d97e02eb-798b-440a-ba36-6f800a431ca1"
# %cd /home/agent/Documents/Project/Knowledge-Distillation
import torch, timm
import torch.nn as nn
import torchvision.models as models
from prepare_dataloader import loaders, dataset_sizes, num_classes# , class_to_idx, idx_to_class
from distiller.teacher_train import training
#TODO create teacher model
teacher = models.resnet34(pretrained=True, progress=True)
teacher.fc = nn.Linear(in_features=teacher.fc.in_features,
out_features=num_classes, bias=True)
# teacher.load_state_dict(
# torch.load("weights/teacher.pth"))
#NOTE train teacher model
epoch_warmup = 1
epoch = 1
path_save_weight = "weights/teacher.pth"
# -
teacher = training(loaders, dataset_sizes, epoch_warmup, epoch, teacher, path_save_weight)
# + colab={"base_uri": "https://localhost:8080/"} id="g3FbQd9Vv829" outputId="e1e25e41-9bb0-424c-e63f-666d76ea232f"
# %cd /content/gdrive/MyDrive/Knowledge-Distillation
from distiller.student_train import training_kd
student = models.resnet18(pretrained=True, progress=True)
student.fc = nn.Linear(in_features=student.fc.in_features,
out_features=num_classes, bias=True)
# student.load_state_dict(
# torch.load("weights/student.pth"))
#NOTE train teacher model
epoch_warmup = 1
epoch = 1
path_save_weight = "/content/gdrive/MyDrive/Classify_pytorch/Weights/student.pth"
student = training_kd(student, teacher, epoch_warmup, epoch, path_save_weight)
# + id="PFbW07ISMTlT"
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "watson21"
# !git add student_train.py
# !git add teacher_train.py
# !git add data.py
# !git add model.py
# !git commit -m "fix grad"
# !git push
# + colab={"base_uri": "https://localhost:8080/"} id="gR3Y435V6Agx" outputId="7ef45ce1-9967-461f-e76d-d070d8cfae08"
import torchvision.models.resnet as resnet
def get_size_model(model):
# model = resnet.resnet18()
param_size = 0
for param in model.parameters():
param_size += param.nelement() * param.element_size()
buffer_size = 0
for buffer in model.buffers():
buffer_size += buffer.nelement() * buffer.element_size()
size_all_mb = (param_size + buffer_size) / 1024**2
print('{} : {:.3f}MB'.format(model.__class__.__name__, size_all_mb))
return size_all_mb
models = [resnet.resnet18(),
resnet.resnet34(),
resnet.resnet50(),
resnet.resnet101(),
resnet.resnet152(),
resnet.resnext50_32x4d(),
resnet.resnext101_32x8d(),
resnet.wide_resnet50_2(),
resnet.wide_resnet101_2()]
for model in models:
get_size_model(model)
del models
# + id="gYw6w2FD9ZV5"
| Knowledge_Distillation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Finding Lane Lines on the Road**
#
#
# This project helps to detect lane lines in images using Python and OpenCV and then applying it for videos as an end goal.
#
# ---
#
# ### Table of Contents
# 1. [Libraries and Packages](#1-libraries-and-packages)
# 2. [Finding Lane Lines](#2-finding-lane-lines)
# - [Modifying `draw_lines()` Function](#modifying-draw_lines-function)
# 3. [Potential Shortcomings with Current Pipeline](#3-potential-shortcomings-with-current-pipeline)
# 4. [Possible Improvements to Pipeline](#4-possible-improvements-to-pipeline)
# 5. [Conclusion](#5-conclusion)
# 6. [License](#6-license)
#
# ---
#
# ### 1. Libraries and Packages
# The following libraries and packages have been used in this project: **`scipy`**, **`matplotlib.pyplot`**, **`matplotlib.image`**, **`numpy`**, **`cv2`**, **`sklearn.linear_model`**, **`sklearn.preprocessing`**, **`sklearn.pipeline`**, **`moviepy.editor`**, and **`IPython.display`**.
#
# ### 2. Finding Lane Lines
#
# First of all I started using the simple methods for making a pipeline, as described in the course. The steps were as following:
#
# 1. Converting the RGB image to a Grayscale image using `cv2.cvtColor()`.
# 2. Applying `cv2.GaussianBlur()` to make the image smoother.
# 3. Applying the OpenCV Canny edge detection function, `cv2.Canny()`, to find edges in the image
# 4. Creating and applying a mask to find the region of interest. The region of interest includs the lane line which the car is moving on.
# - It helps to remove other objects on the image that are redundant and make it easier to make pipelines.
# 5. Applying _Hough Transform_ on the edge detected image to find lines.
# 6. Modifying the `draw_lines()` function to make pipelines on the image.
# - I will describe how to modify `draw_lines()` function later.
#
# You can change/adjust the parameters of _Canny_ or _Hough Transform_ functions to find pipelines for the test images. But when it comes to the videos, especially the challange video, it's not able to find pipelines for the all frames accurately!
#
# Before explaining _the steps to improve lane line detection_ I will describe `draw_lines()` function modification steps.
#
#
# #### Modifying `draw_lines()` Function
# In order to draw a single line on the left and right lanes, I modified the `draw_lines(img, lines, color=[255, 0, 0], thickness=8)` function by calculating the left line and the right lane slopes. Then I drew a solid line with the calculated slope. To accomplish that the steps are as following:
# 1. **Calculate the average of positive slopes and negative slopes separately.** In most cases the left line has a negative (-) slope and the right line has a positive (+) slope. So, this step gives us a good estimate of the final line slopes.
# ```python
# slopes = [((y2-y1)/(x2-x1)) for line in lines for x1,y1,x2,y2 in line]
# lane_slopes_right = np.average(np.array([m for m in slopes if m > 0]))
# lane_slopes_left = np.average(np.array([m for m in slopes if m < 0]))
# ```
# 2. **Find the left and right lines.** To define a line uniquely, only **line slope** and **a point on the line** are needed. I created a function called `get_line(slope, x0, y0, y_min, y_max)` to define a line with its parameters and make it reusable.
# ```python
# def get_line(slope, x0, y0, y_min, y_max):
# x1 = int(x0 + (y_max - y0) / slope)
# x2 = int(x0 + (y_min - y0) / slope)
# return x1, y_max, x2, y_min
# ```
# This function gets the line parameters (`slope`, `x0`, `y0`) and range of line (`y_min`, `y_max`) and returns the _start point_ (`x1, y_max`) and the _end point_ (`x2, y_min`) of the line.
# 3. **Finding line parameters `slope, x0, y0, y_min, y_max`.** We know the line `slope` from first step. So, it's time to find the rest!
#
# i. Categorize the left and right lines.
# ```python
# lines_left = [line for line in lines for x1,y1,x2,y2 in line if ((y2-y1)/(x2-x1)) < 0]
# lines_right = [line for line in lines for x1,y1,x2,y2 in line if ((y2-y1)/(x2-x1)) > 0]
# ```
# ii. Find the upper (a) and lower (b) points of each side. It helps to find `x0, y0, y_min, y_max`.
# ```python
# # Left side
# l_a_dict = {y1:x1 for line in lines_left for x1, y1, x2, y2 in line}
# l_a_y = min(l_a_dict, key=l_a_dict.get)
# l_a = [l_a_dict[l_a_y], l_a_y]
# l_b_dict = {y2:x2 for line in lines_left for x1, y1, x2, y2 in line}
# l_b_y = max(l_b_dict, key=l_b_dict.get)
# l_b = [l_b_dict[l_b_y], l_b_y]
#
# # Right side
# r_a_dict = {y1:x1 for line in lines_right for x1, y1, x2, y2 in line}
# r_a_y = min(r_a_dict, key=r_a_dict.get)
# r_a = [r_a_dict[r_a_y], r_a_y]
# r_b_dict = {y2:x2 for line in lines_right for x1, y1, x2, y2 in line}
# r_b_y = max(r_b_dict, key=r_b_dict.get)
# r_b = [r_b_dict[r_b_y], r_b_y]
# ```
# I used `l_b` and `r_b` as `x0, y0` for the left and the right lines respectively.
#
# iii. Finding vertical range of lines `y_min, y_max`.
# ```python
# ymin = min(l_b_y, r_a_y)
# ymax = max(l_a_y, r_b_y)
# ```
# 4. **Add a single line to the image as the left pipeline.** I used `cv2.line()` function for adding a line to the image.
# ```python
# l_x1, l_y1, l_x2, l_y2 = get_line(lane_slopes_left, l_b[0], l_b[1], ymin, ymax)
# cv2.line(img, (l_x1, l_y1), (l_x2, l_y2), color, thickness)
# ```
# Do the same thing for the right side:
# ```python
# r_x1, r_y1, r_x2, r_y2 = get_line(lane_slopes_right, r_a[0], r_a[1], ymin, ymax)
# cv2.line(img, (r_x1, r_y1), (r_x2, r_y2), color, thickness)
# ```
# `weighted_img()` also could be used instead of `cv2.line()` for adding translucent pipelines to the original image.
#
# ### 3. Potential Shortcomings with Current Pipeline
# Before improving the lane line detection code, potential shortcomings need to be identified. There are two important shortcoming here:
# 1. **Color selection:** The algorithm for color selection is quite simple and it's useless, for instance, when the brightness of the image is high or the contrast is low or there is a light-color object on the road close to the lane line sides, etc.
# 2. **Finding the solid left and right lines:** The road isn't always straight and it turns to the left an right. So, _striaght pipeline_ is a wrong assumption. The truth is that a pipeline is actually a curve!
#
# ### 4. Possible Improvements to Pipeline
# First step is to _improve the color selection algorithm_ and second step is _estimating the best possible curve line for the pipeline_. For the rest of this section I use `test_images/whiteCarLaneSwitch.jpg` to show result of each step.
#
# <br/>
# <figure>
# <img src="article/whiteCarLaneSwitch_source.jpg" width="460" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> The image which is used to show result of each step. </i> </p>
# </figcaption>
# </figure>
#
# #### Color Selection
# A possible improvement would be using a different color space like HLS instead of RGB for selecting white and yellow colors. Another one is that do this separately for each color!
#
# ##### Different Color Space
# To start, I created `copy_hls()` function for converting color space from `RGB` to `HLS`.
# ```python
# def copy_hls(image):
# img = np.copy(image)
# # Convert BGR to HLS
# hls = cv2.cvtColor(img, cv2.COLOR_RGB2HLS)
# return hls
# ```
#
# > **Note:** It is a good practice to initially create a copy of the source image and then make changes to the copied version instead of the main version.
#
# ##### White and Yellow Mask
# The `get_white_yellow_mask(img)` function generates a mask to filter out almost every objects that doesn't have white or yellow colors. Using the lower and upper range of color and `cv2.inRange()` the mask for the a given color could be found. `mask_y` for the yellow objects:
# ```python
# # Convert BGR to HLS
# img_hls = copy_hls(img)
#
# # Yellow mask
# # define range of yellow color in HLS color space
# lower_range = np.array(y_lower, dtype=np.uint8)
# upper_range = np.array(y_upper, dtype=np.uint8)
#
# # Apply the range values to the HLS image to get only yellow colors
# mask_y = cv2.inRange(img_hls, lower_range, upper_range)
# ```
# and `mask_w` for the white color objects:
# ```python
# # White mask
# # define range of white color in HLS color space
# lower_range = np.array(w_lower, dtype=np.uint8)
# upper_range = np.array(w_upper, dtype=np.uint8)
#
# # Apply the range values to the HLS image to get only white colors
# mask_w = cv2.inRange(img_hls, lower_range, upper_range)
# ```
# > At the end of section 3, I will explain about lower and upper ranges of yellow and white colors.
#
# Then mix `mask_y` and `mask_w` to a single mask for both colors using `cv2.bitwise_or()`.
# ```python
# # Mix the generated masks
# mask_wy = cv2.bitwise_or(mask_y, mask_w)
# idx = mask_wy != 0
# mask_wy[idx] = 255
# ```
#
# The result is as following:
#
# <table style="width:100%; text-align: center;">
# <tr>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_mask_y.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> `mask_y`: Yellow color mask </i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_mask_w.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> `mask_w`: White color mask </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# <tr>
# <td colspan="2">
# <figure>
# <img src="article/whiteCarLaneSwitch_mask_wy.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> `mask_wy`: Mixed mask for both yellow and white </i> </p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# #### Canny Edge Detection
# The `get_canny(img, mask)` function creates a convenient canny image. By using `mask` we can extract the regions in the image that includes white and/or yellow objects. I call it `region_of_interest`.
# ```python
# region_of_interest = cv2.bitwise_and(img, mask)
# ```
# Then I made it a little bit blur to reduce the noise of the image which also helps gaining a better canny image.
# ```python
# region_of_interest_blur = gaussian_blur(region_of_interest, kernel_size)
# ```
# At the end, I converted it to a canny image using `canny` function. It's one of helper functions and it uses
# ```python
# region_of_interest_canny = canny(region_of_interest_blur, canny_low_threshold, canny_high_threshold)
# ```
#
# You can see the results below:
#
# <table style="width:100%; text-align: center;">
# <tr>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_region_of_interest.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> `region_of_interest` : The region of interest</i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_region_of_interest_blur.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> `region_of_interest_blur`: The blurred region of interest to reduce the noise</i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# <tr>
# <td colspan="2">
# <figure>
# <img src="article/whiteCarLaneSwitch_region_of_interest_canny.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> `region_of_interest_canny`: Canny edge detected image </i> </p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# #### Transforming the Lane Area to a Rectangle
# To simplify, the lane area in canny image is transformed to a rectangle. For this purpose `transform_to_rectangle()` has been used. In order to implement this function `cv2.getPerspectiveTransform()` and `cv2.warpPerspective()` of OpenCV are used.
# <br/>
# <table style="width:100%; text-align: center;">
# <tr>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_region_of_interest_canny.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> Canny edge detected image</i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_img_trans_org2rec.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> Transforming the lane line area to the rectangle </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# #### Separating the Left and Right Sides
# Before finding the left and right curved pipelines, I separated the left and right lines using `separate_to_left_right()` as following:
#
# <table style="width:100%; text-align: center;">
# <tr>
# <td colspan="2">
# <figure>
# <img src="article/whiteCarLaneSwitch_img_trans_org2rec.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> Transforming the lane line area to the rectangle </i> </p>
# </figcaption>
# </figure>
# </td>
# </tr>
# <tr>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_img_left.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> Used for finding the left curved line</i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_img_right.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> Used for finding the right curved line </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# #### Hough Transform
# Next step is applying _Hough Transform_ by using `find_hough_lines()` function to the both left and right side images. The results for each side is a set of lines and each line contains a couple of points that indicates the starting and ending points of it.
# You can see these points below:
# <table style="width:100%; text-align: center;">
# <tr>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_img_left_plus_hough_lines.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> The starting and ending points of Hough Transform lines for the left side </i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_img_right_plus_hough_lines.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> The starting and ending points of Hough Transform lines for the right side </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# #### Find the Best Line with Polynomial Interpolation
# `scikit-learn` has some pakcages for Polynomial Interpolation (see an example [here](http://scikit-learn.org/stable/auto_examples/linear_model/plot_polynomial_interpolation.html)). In fact, I used linear regression with polynomial features to approximate a nonlinear function that fits the points which were found from the previous step. I used the following packages:
# ```python
# from sklearn.linear_model import Ridge
# from sklearn.preprocessing import PolynomialFeatures
# from sklearn.pipeline import make_pipeline
# ```
# The function that is responsible to fit the best line is `find_pipeline`. The polynomial with a degree equal to 2 gives the best results. If you increase or decrease the degree of polynomial, the underfitting and overfitting problems occur. For more information see [here](http://scikit-learn.org/stable/auto_examples/model_selection/plot_underfitting_overfitting.html#sphx-glr-auto-examples-model-selection-plot-underfitting-overfitting-py). Finally, the calculated pipeline formula would be used to draw pipelines using `draw_pred_lines()` function (image below).
# <br/>
# <table style="width:100%; text-align: center;">
# <tr>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_calculated_curved_pipelines.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> The calculated pipelines </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# > I created a `buffer_model` for the calculated pipeline models. In each step after calculating the model I stored them (left and right models) in this buffer. And they will be used to a more accurate estimation of the pipelines for the next frame. To clarify, assume we have **two first frames** of a video: `F1` and `F2`. The steps are as following:
# > 1. Calculate the pipeline predict model for `F1` (`pred_model_1`).
# > 2. Use this `pred_model_1` to add an estimated pipelines to `F1`.
# > 3. Add it to the buffer.
# > 4. Next frame (`F2`) comes for estimation.
# > 5. Calculate the pipeline predict model for `F2`(`pred_model_2`).
# > 6. Use `pred_model_1` and `pred_model_2` to add an estimated pipelines to `F2`.
# > - If deviation of a point on the pipe line 2 with respect to the same point on the line 1 exceeds a specified threshold, `th_0`, replace it with a point which has a deviation equal to the `th_0` otherwise do nothing.
# >
# > The step 6 really makes the changes of curvature smooth.
#
# #### Adding The Green Zone and Retransform it to the Original Shape
# The safe driving area for a car is the area between the pipelines that I call it "Green Zone." Let's fill the area between red pipelines with the green color using `add_green_zone()` function. For implementing this function I used `cv2.fillPoly()` from OpenCV (figure below). Then retransform it to the original perspective using `transform_back_to_origin()`. The results are as following:
# <br/>
# <table style="width:100%; text-align: center;">
# <tr>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_curved_pipelines_with_green_zone.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> The calculated pipelines with green zone</i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_img_trans_rec2org.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> Transformed result </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# #### Adding the Pipelines with Green Zone to the Original Image
# This is a time for adding the pipeline with green zone to the original image. `weighted_img()` function adds a translucent version to the original image (figure below).
# <br/>
# <table style="width:100%; text-align: center;">
# <tr>
# <td>
# <figure>
# <img src="article/whiteCarLaneSwitch_final.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> The final result </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# #### `find_pipelines_and_green_zone()` Function
# `find_pipelines_and_green_zone()` function (including all the mentioned steps) adds pipe lines and green zone to a given road image. You can see this function below:
# ```python
# def find_pipelines_and_green_zone(img):
# mask = get_white_yellow_mask(img)
#
# canny = get_canny(img, mask)
#
# rows,cols,ch = img.shape
# img_trans = transform_to_rectangle(canny, rows, cols)
# img_left, img_right = separate_to_left_right(img_trans)
#
# lines_left, lines_right = find_hough_lines(img_left, img_right)
#
# len_x_history = 40
# len_x_plot = 10
# x_history = np.int16(np.linspace(0, rows - 1, len_x_history))
# x_plot = np.int16(np.linspace(0, rows - 1, len_x_plot))
# y_plot_left, y_plot_right = find_pipeline(rows, cols, lines_left, lines_right, x_history, x_plot)
#
# # Draw lines
# line_width = 40
# img_lines = draw_pred_lines(rows, cols, y_plot_left, y_plot_right, x_plot, line_width)
# img_green_zone = add_green_zone(img_lines, y_plot_left, y_plot_right, x_plot)
#
# # Transform region of interest
# img_trans_back = transform_back_to_origin(img_green_zone, rows, cols)
#
# # Add it to the original image
# final = weighted_img(img, img_trans_back)
# return final
# ```
#
# #### Test Images
# The result of the other test images are as following:
#
# <table style="width:100%; text-align: center;">
# <tr>
# <td>
# <figure>
# <img src="article/solidWhiteCurve_final.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> solidWhiteCurve.jpg </i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/solidWhiteRight_final.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> solidWhiteRight.jpg </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# <tr>
# <td>
# <figure>
# <img src="article/solidYellowCurve_final.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> solidYellowCurve.jpg </i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/solidYellowCurve2_final.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> solidYellowCurve2.jpg </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# <tr>
# <td>
# <figure>
# <img src="article/solidYellowCurveChal_final.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> solidYellowCurveChal.jpg </i></p>
# </figcaption>
# </figure>
# </td>
# <td>
# <figure>
# <img src="article/solidYellowLeft_final.jpg" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> <i> solidYellowLeft.jpg </i></p>
# </figcaption>
# </figure>
# </td>
# </tr>
# </table>
#
# #### Tuning the Parameters
# I put the parameters for different parts of this code in one place to tune them conveniently. The final result for these parameters are as following:
# ```python
# # Color in range
# y_lower = [10, 0, 120]
# y_upper = [40, 255, 255]
# w_lower = [16, 182, 0]
# w_upper = [255, 255, 255]
#
# # Transform
# coef_w_top = 0.15
# coef_w_dwn = 1.00
# offset_v_top = 0.63
# offset_v_dwn = 0.02
#
# # Blur
# kernel_size = 9
#
# # Canny
# canny_low_threshold = 0
# canny_high_threshold = 255
#
# # Make a blank the same size as our image to draw on
# rho = 1 # distance resolution in pixels of the Hough grid
# theta = np.pi/180 * 0.5 # angular resolution in radians of the Hough grid
# threshold = 10 # minimum number of votes (intersections in Hough grid cell)
# min_line_len = 15 # minimum number of pixels making up a line
# max_line_gap = 10 # maximum gap in pixels between connectable line segments
#
# # Interpolation
# degree = 2
# ```
#
# These values has been tuned for all videos including the challenge video.
#
# ##### You can find the output videos [here](test_videos_output) or watch them online [here](https://www.youtube.com/watch?v=S0_758-sbnc&index=2&list=PLgjxKJEo-VjELoTEKXEjA8Vi7s5xZCzI9).
#
#
# ### 5. Conclusion
# Using better approaches for "color selection" and "pipeline estimation" results to a more accurate output. Also using the buffer makes the changes of the pipeline curvature smoother.
#
# ### 6. License
# [MIT License](LICENSE).
| README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Exercise 04: Plotting Stock Price Data in Tabs
# We will revisit the basic widgets and build a simple plot that will display the first 25 data points for the selected stock. The displayed stock can be changed with a drop-down menu.
# The dataset of this exercise contains temporal stock price data.
# This means we'll be looking at data over a range of time.
# > The dataset for this exercise is quite big and has to be downloaded and added into the data folder.
# The dataset is available in the data folder of the GitHub repository; here is the link to it: https://bit.ly/2UaLtSV
# #### Loading our dataset
# importing the necessary dependencies
import pandas as pd
# +
# make bokeh display figures inside the notebook
from bokeh.io import output_notebook
output_notebook()
# -
# loading the Dataset with geoplotlib
dataset = pd.read_csv('../../Datasets/stock_prices.csv')
# looking at the dataset
dataset.head()
# Since the date column has no information about the hour, minute, and second, we want to avoid displaying them in the visualization later on and simply display the year, month, and day.
#
# Therefore we'll create a new column that holds the formatted short version of the date value.
# +
# mapping the date of each row to only the year-month-day format
from datetime import datetime
def shorten_time_stamp(timestamp):
shortened = timestamp[0]
if len(shortened) > 10:
parsed_date=datetime.strptime(shortened, '%Y-%m-%d %H:%M:%S')
shortened=datetime.strftime(parsed_date, '%Y-%m-%d')
return shortened
dataset['short_date'] = dataset.apply(lambda x: shorten_time_stamp(x), axis=1)
dataset.head()
# -
# **Note:**
# The exectuion of the cell will take a moment since it's a fairly large dataset.
# Please be patient.
#
# The last, newly added, column now holds the timestamp without the hour, minute, and second information.
# ---
# #### Creating a basic plot and adding a widget
# In this task, we will create a basic visualization with the stock price dataset. This will be your first interactive visualization in which you can dynamically change the stock that is displayed in the graph.
#
# We will get used to one of the above described interactive widgets, the dropdown menu. It will be the main point of interaction for our visualization.
# +
# importing the necessary dependencies
from ipywidgets import interact
from bokeh.models.widgets import Panel, Tabs
from bokeh.plotting import figure, show
# -
# In order to structure our notebook better, we want to write a adaptable method that gets a subsection of stock data as argument and builds a two tab `Pane` object, that lets us switch between the two views in our visualization.
# The first tab will contain a line plot of the given data, the second one a circle-based representation of the same data.
# A legend will display the name of the currently viewed at stock.
# method to build the tab-based plot
def get_plot(stock):
stock_name=stock['symbol'].unique()[0]
line_plot=figure(title='Stock prices',
x_axis_label='Date', x_range=stock['short_date'],
y_axis_label='Price in $USD')
line_plot.line(stock['short_date'], stock['high'], legend_label=stock_name)
line_plot.xaxis.major_label_orientation = 1
circle_plot=figure(title='Stock prices',
x_axis_label='Date', x_range=stock['short_date'],
y_axis_label='Price in $USD')
circle_plot.circle(stock['short_date'], stock['high'], legend_label=stock_name)
circle_plot.xaxis.major_label_orientation = 1
line_tab=Panel(child=line_plot, title='Line')
circle_tab=Panel(child=circle_plot, title='Circles')
tabs = Tabs(tabs=[ line_tab, circle_tab ])
return tabs
# Before we can build our interaction, we have to get a list of all the stock names that are present in the dataset.
# Once we have done that, we can then use this list as an input for the interact element.
# With each interaction of the dropdown, our displayed data will then be updated.
#
# To keep it simple, we only want to display the first 25 entries of each stock in this task.
# By default, the stock of Apple should be displayed, it's symbol in the dataset is 'AAPL'.
# extracing all the stock names
stock_names=dataset['symbol'].unique()
# +
# creating the dropdown interaction and building the plot
# based on selection
@interact(Stock=stock_names)
def get_stock_for(Stock='AAPL'):
stock = dataset[dataset['symbol'] == Stock][:25]
show(get_plot(stock))
# -
# **Note:**
# We can already see that each date is displayed on the x-axis. If we want to display a bigger time range, we have to customize the ticks on our x-axis. This can be done using `Ticker` objects.
| Exercise04/Exercise04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Evaluation of a Pipeline and its Components
#
# [](https://colab.research.google.com/github/deepset-ai/haystack/blob/master/tutorials/Tutorial5_Evaluation.ipynb)
#
# To be able to make a statement about the quality of results a question-answering pipeline or any other pipeline in haystack produces, it is important to evaluate it. Furthermore, evaluation allows determining which components of the pipeline can be improved.
# The results of the evaluation can be saved as CSV files, which contain all the information to calculate additional metrics later on or inspect individual predictions.
# + [markdown] id="lEKOjCS5U7so" pycharm={"is_executing": true}
# ### Prepare environment
#
# #### Colab: Enable the GPU runtime
# Make sure you enable the GPU runtime to experience decent speed in this tutorial.
# **Runtime -> Change Runtime type -> Hardware accelerator -> GPU**
#
# <img src="https://raw.githubusercontent.com/deepset-ai/haystack/master/docs/img/colab_gpu_runtime.jpg">
# + colab={"base_uri": "https://localhost:8080/"} id="xhFIMX_7U7ss" outputId="285b2491-01e5-4bfd-cba9-c2279d4417c4" pycharm={"name": "#%%\n"}
# Make sure you have a GPU running
# !nvidia-smi
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="vgmFOp82Oht_" outputId="5bbcbb42-3a90-43a9-ebfd-598a98fa7143" pycharm={"name": "#%%\n"}
# Install the latest release of Haystack in your own environment
# #! pip install farm-haystack
# Install the latest master of Haystack
# !pip install --upgrade pip
# !pip install git+https://github.com/deepset-ai/haystack.git#egg=farm-haystack[colab]
# -
# ## Start an Elasticsearch server
# You can start Elasticsearch on your local machine instance using Docker. If Docker is not readily available in your environment (eg., in Colab notebooks), then you can manually download and execute Elasticsearch from source.
# + id="tNoaWcDKOhuL" pycharm={"name": "#%%\n"}
# If Docker is available: Start Elasticsearch as docker container
# from haystack.utils import launch_es
# launch_es()
# Alternative in Colab / No Docker environments: Start Elasticsearch from source
# ! wget https://artifacts.elastic.co/downloads/elasticsearch/elasticsearch-7.9.2-linux-x86_64.tar.gz -q
# ! tar -xzf elasticsearch-7.9.2-linux-x86_64.tar.gz
# ! chown -R daemon:daemon elasticsearch-7.9.2
import os
from subprocess import Popen, PIPE, STDOUT
es_server = Popen(
["elasticsearch-7.9.2/bin/elasticsearch"], stdout=PIPE, stderr=STDOUT, preexec_fn=lambda: os.setuid(1) # as daemon
)
# wait until ES has started
# ! sleep 30
# -
# ## Fetch, Store And Preprocess the Evaluation Dataset
# + colab={"base_uri": "https://localhost:8080/"} id="tTXxr6TAOhuz" outputId="586d4775-4354-4ed9-a72c-c30bedcdfbee" pycharm={"is_executing": true, "name": "#%%\n"}
from haystack.utils import fetch_archive_from_http
# Download evaluation data, which is a subset of Natural Questions development set containing 50 documents with one question per document and multiple annotated answers
doc_dir = "data/tutorial5"
s3_url = "https://s3.eu-central-1.amazonaws.com/deepset.ai-farm-qa/datasets/nq_dev_subset_v2.json.zip"
fetch_archive_from_http(url=s3_url, output_dir=doc_dir)
# + id="T-G7Ay2jU7s_"
# make sure these indices do not collide with existing ones, the indices will be wiped clean before data is inserted
doc_index = "tutorial5_docs"
label_index = "tutorial5_labels"
# + id="B_NEtezLOhu5" pycharm={"name": "#%%\n"}
# Connect to Elasticsearch
from haystack.document_stores import ElasticsearchDocumentStore
# Connect to Elasticsearch
document_store = ElasticsearchDocumentStore(
host="localhost",
username="",
password="",
index=doc_index,
label_index=label_index,
embedding_field="emb",
embedding_dim=768,
excluded_meta_data=["emb"],
)
# + colab={"base_uri": "https://localhost:8080/"} id="bRFsQUAJOhu_" outputId="477031b9-5c2c-4128-ef5f-54db86259734" pycharm={"name": "#%%\n"}
from haystack.nodes import PreProcessor
# Add evaluation data to Elasticsearch Document Store
# We first delete the custom tutorial indices to not have duplicate elements
# and also split our documents into shorter passages using the PreProcessor
preprocessor = PreProcessor(
split_length=200,
split_overlap=0,
split_respect_sentence_boundary=False,
clean_empty_lines=False,
clean_whitespace=False,
)
document_store.delete_documents(index=doc_index)
document_store.delete_documents(index=label_index)
# The add_eval_data() method converts the given dataset in json format into Haystack document and label objects. Those objects are then indexed in their respective document and label index in the document store. The method can be used with any dataset in SQuAD format.
document_store.add_eval_data(
filename="data/tutorial5/nq_dev_subset_v2.json",
doc_index=doc_index,
label_index=label_index,
preprocessor=preprocessor,
)
# + [markdown] id="gy8YwmSYOhvE" pycharm={"name": "#%% md\n"}
# ## Initialize the Two Components of an ExtractiveQAPipeline: Retriever and Reader
# + id="JkhaPMIJOhvF" pycharm={"name": "#%%\n"}
# Initialize Retriever
from haystack.nodes import ElasticsearchRetriever
retriever = ElasticsearchRetriever(document_store=document_store)
# Alternative: Evaluate dense retrievers (EmbeddingRetriever or DensePassageRetriever)
# The EmbeddingRetriever uses a single transformer based encoder model for query and document.
# In contrast, DensePassageRetriever uses two separate encoders for both.
# Please make sure the "embedding_dim" parameter in the DocumentStore above matches the output dimension of your models!
# Please also take care that the PreProcessor splits your files into chunks that can be completely converted with
# the max_seq_len limitations of Transformers
# The SentenceTransformer model "sentence-transformers/multi-qa-mpnet-base-dot-v1" generally works well with the EmbeddingRetriever on any kind of English text.
# For more information and suggestions on different models check out the documentation at: https://www.sbert.net/docs/pretrained_models.html
# from haystack.retriever import EmbeddingRetriever, DensePassageRetriever
# retriever = EmbeddingRetriever(document_store=document_store, model_format="sentence_transformers",
# embedding_model="sentence-transformers/multi-qa-mpnet-base-dot-v1")
# retriever = DensePassageRetriever(document_store=document_store,
# query_embedding_model="facebook/dpr-question_encoder-single-nq-base",
# passage_embedding_model="facebook/dpr-ctx_encoder-single-nq-base",
# use_gpu=True,
# max_seq_len_passage=256,
# embed_title=True)
# document_store.update_embeddings(retriever, index=doc_index)
# + colab={"base_uri": "https://localhost:8080/", "height": 313, "referenced_widgets": ["118aedffeecd4f109ae04a4561baeb08", "59efb57b419242a1aba4d20084e29d38", "ddaf59cedca143c8b4fe005a51077323", "25d7818d6f7b4b628ab3f83f2c2fa6a3", "<KEY>", "<KEY>", "3d211e6614f4451e9d14866cb3a8332d", "<KEY>", "<KEY>", "<KEY>", "8e0958b5dc27412e9f0332da3457ffdb", "<KEY>", "b480c97c1d9944b9a8dd09ed6e1e9bd3", "ed683929822b4084ba33e89b23936b16", "94c88d0fc3f949fbacfb6b4fcd99cc63", "b262e92ff6484405a0e9364f6ecafb6a", "bacf1704dbaf4176afbe2cbcc8e036ef", "fe56b1d86ab84675b82781a1f8edd40a", "d7e3c8e1e0424cec9dc1b97090b5af87", "98da94a8d7b94fb4a08adcebea15e114", "<KEY>", "2bd3bd51ae644c1894a2ddca09d14e85", "<KEY>", "<KEY>", "2c028a3f096344d68071d78387efa117", "<KEY>", "<KEY>", "76f4af76b42f460fa34d5f00a9656dc5", "73d7fdd3f38349b4882124d8351eace5", "ea439e2251ed467fb3a775f0c8e0c3bb", "<KEY>", "fc011913e8464d439a97fe75ef5f9fa1", "8a9f9b7bab8e40278430a35720066a61", "5db857b352964db3a617568ff1dce86d", "7752437041f745a4af4b9130df3fefa7", "5f94d400ea884c1cadfc966e44849b3a", "<KEY>", "994ae85181664e2e87a2ee18a7a237ba", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "6f028c7e888e4ae5ab5c1e42ff142b5f", "c8ba8c2a210b45f6a9b5257589babac3", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "eb11ea5785284bf6a15cc31ad643ed88", "<KEY>", "<KEY>", "<KEY>", "d4dbfa5e89e7432dbed34606a786fd6f", "7e058076836e438daf5399428eabac5e", "<KEY>", "69750fea2e7149eab8928282ba9bae29", "<KEY>", "<KEY>", "<KEY>", "4dad7e58cc47436aafe38230514325a1", "<KEY>", "2a9ef1f2f43d47b28cd0ff7ef4a21ade", "caa374f7dc5045218c6f71f322d8e6be", "e567fab4446544f795be2eb0a6705f9c", "<KEY>", "3096cae7388e4b988df306be9cc58afd"]} id="cW3Ypn_gOhvK" outputId="4b5feff7-ae9f-4cd8-de1e-944f0eb66f66" pycharm={"name": "#%%\n"}
# Initialize Reader
from haystack.nodes import FARMReader
reader = FARMReader("deepset/roberta-base-squad2", top_k=4, return_no_answer=True)
# Define a pipeline consisting of the initialized retriever and reader
from haystack.pipelines import ExtractiveQAPipeline
pipeline = ExtractiveQAPipeline(reader=reader, retriever=retriever)
# The evaluation also works with any other pipeline.
# For example you could use a DocumentSearchPipeline as an alternative:
# from haystack.pipelines import DocumentSearchPipeline
# pipeline = DocumentSearchPipeline(retriever=retriever)
# + [markdown] id="7i84KXONOhvc" pycharm={"name": "#%% md\n"}
# ## Evaluation of an ExtractiveQAPipeline
# Here we evaluate retriever and reader in open domain fashion on the full corpus of documents i.e. a document is considered
# correctly retrieved if it contains the gold answer string within it. The reader is evaluated based purely on the
# predicted answer string, regardless of which document this came from and the position of the extracted span.
#
# The generation of predictions is seperated from the calculation of metrics. This allows you to run the computation-heavy model predictions only once and then iterate flexibly on the metrics or reports you want to generate.
#
# + pycharm={"name": "#%%\n"}
from haystack.schema import EvaluationResult, MultiLabel
# We can load evaluation labels from the document store
# We are also opting to filter out no_answer samples
eval_labels = document_store.get_all_labels_aggregated(drop_negative_labels=True, drop_no_answers=True)
## Alternative: Define queries and labels directly
# eval_labels = [
# MultiLabel(
# labels=[
# Label(
# query="who is written in the book of life",
# answer=Answer(
# answer="every person who is destined for Heaven or the World to Come",
# offsets_in_context=[Span(374, 434)]
# ),
# document=Document(
# id='1b090aec7dbd1af6739c4c80f8995877-0',
# content_type="text",
# content='Book of Life - wikipedia Book of Life Jump to: navigation, search This article is
# about the book mentioned in Christian and Jewish religious teachings...'
# ),
# is_correct_answer=True,
# is_correct_document=True,
# origin="gold-label"
# )
# ]
# )
# ]
# Similar to pipeline.run() we can execute pipeline.eval()
eval_result = pipeline.eval(labels=eval_labels, params={"Retriever": {"top_k": 5}})
# + pycharm={"name": "#%%\n"}
# The EvaluationResult contains a pandas dataframe for each pipeline node.
# That's why there are two dataframes in the EvaluationResult of an ExtractiveQAPipeline.
retriever_result = eval_result["Retriever"]
retriever_result.head()
# + pycharm={"name": "#%%\n"}
reader_result = eval_result["Reader"]
reader_result.head()
# + pycharm={"name": "#%%\n"}
# We can filter for all documents retrieved for a given query
query = "who is written in the book of life"
retriever_book_of_life = retriever_result[retriever_result["query"] == query]
# + pycharm={"name": "#%%\n"}
# We can also filter for all answers predicted for a given query
reader_book_of_life = reader_result[reader_result["query"] == query]
# + pycharm={"name": "#%%\n"}
# Save the evaluation result so that we can reload it later and calculate evaluation metrics without running the pipeline again.
eval_result.save("../")
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Calculating Evaluation Metrics
# Load an EvaluationResult to quickly calculate standard evaluation metrics for all predictions,
# such as F1-score of each individual prediction of the Reader node or recall of the retriever.
# To learn more about the metrics, see [Evaluation Metrics](https://haystack.deepset.ai/guides/evaluation#metrics-retrieval)
# + pycharm={"name": "#%%\n"}
saved_eval_result = EvaluationResult.load("../")
metrics = saved_eval_result.calculate_metrics()
print(f'Retriever - Recall (single relevant document): {metrics["Retriever"]["recall_single_hit"]}')
print(f'Retriever - Recall (multiple relevant documents): {metrics["Retriever"]["recall_multi_hit"]}')
print(f'Retriever - Mean Reciprocal Rank: {metrics["Retriever"]["mrr"]}')
print(f'Retriever - Precision: {metrics["Retriever"]["precision"]}')
print(f'Retriever - Mean Average Precision: {metrics["Retriever"]["map"]}')
print(f'Reader - F1-Score: {metrics["Reader"]["f1"]}')
print(f'Reader - Exact Match: {metrics["Reader"]["exact_match"]}')
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Generating an Evaluation Report
# A summary of the evaluation results can be printed to get a quick overview. It includes some aggregated metrics and also shows a few wrongly predicted examples.
# + pycharm={"name": "#%%\n"}
pipeline.print_eval_report(saved_eval_result)
# -
# ## Advanced Evaluation Metrics
# As an advanced evaluation metric, semantic answer similarity (SAS) can be calculated. This metric takes into account whether the meaning of a predicted answer is similar to the annotated gold answer rather than just doing string comparison.
# To this end SAS relies on pre-trained models. For English, we recommend "cross-encoder/stsb-roberta-large", whereas for German we recommend "deepset/gbert-large-sts". A good multilingual model is "sentence-transformers/paraphrase-multilingual-mpnet-base-v2".
# More info on this metric can be found in our [paper](https://arxiv.org/abs/2108.06130) or in our [blog post](https://www.deepset.ai/blog/semantic-answer-similarity-to-evaluate-qa).
# + pycharm={"name": "#%%\n"}
advanced_eval_result = pipeline.eval(
labels=eval_labels, params={"Retriever": {"top_k": 1}}, sas_model_name_or_path="cross-encoder/stsb-roberta-large"
)
metrics = advanced_eval_result.calculate_metrics()
print(metrics["Reader"]["sas"])
# -
# ## Isolated Evaluation Mode
# The isolated node evaluation uses labels as input to the Reader node instead of the output of the preceeding Retriever node.
# Thereby, we can additionally calculate the upper bounds of the evaluation metrics of the Reader. Note that even with isolated evaluation enabled, integrated evaluation will still be running.
#
# + pycharm={"name": "#%%\n"}
eval_result_with_upper_bounds = pipeline.eval(
labels=eval_labels, params={"Retriever": {"top_k": 5}, "Reader": {"top_k": 5}}, add_isolated_node_eval=True
)
# + pycharm={"name": "#%%\n"}
pipeline.print_eval_report(eval_result_with_upper_bounds)
# -
# ## Evaluation of Individual Components: Retriever
# Sometimes you might want to evaluate individual components, for example, if you don't have a pipeline but only a retriever or a reader with a model that you trained yourself.
# Here we evaluate only the retriever, based on whether the gold_label document is retrieved.
# + pycharm={"name": "#%%\n"}
## Evaluate Retriever on its own
# Note that no_answer samples are omitted when evaluation is performed with this method
retriever_eval_results = retriever.eval(top_k=5, label_index=label_index, doc_index=doc_index)
# Retriever Recall is the proportion of questions for which the correct document containing the answer is
# among the correct documents
print("Retriever Recall:", retriever_eval_results["recall"])
# Retriever Mean Avg Precision rewards retrievers that give relevant documents a higher rank
print("Retriever Mean Avg Precision:", retriever_eval_results["map"])
# -
# Just as a sanity check, we can compare the recall from `retriever.eval()` with the multi hit recall from `pipeline.eval(add_isolated_node_eval=True)`.
# These two recall metrics are only comparable since we chose to filter out no_answer samples when generating eval_labels and setting doc_relevance_col to `"gold_id_match"`. Per default `calculate_metrics()` has doc_relevance_col set to `"gold_id_or_answer_match"` which interprets documents as relevant if they either match the gold_id or contain the answer.
metrics = eval_result_with_upper_bounds.calculate_metrics(doc_relevance_col="gold_id_match")
print(metrics["Retriever"]["recall_multi_hit"])
# ## Evaluation of Individual Components: Reader
# Here we evaluate only the reader in a closed domain fashion i.e. the reader is given one query
# and its corresponding relevant document and metrics are calculated on whether the right position in this text is selected by
# the model as the answer span (i.e. SQuAD style)
# + pycharm={"name": "#%%\n"}
# Evaluate Reader on its own
reader_eval_results = reader.eval(document_store=document_store, label_index=label_index, doc_index=doc_index)
top_n = reader_eval_results["top_n"]
# Evaluation of Reader can also be done directly on a SQuAD-formatted file without passing the data to Elasticsearch
# reader_eval_results = reader.eval_on_file("../data/nq", "nq_dev_subset_v2.json", device=device)
# Reader Top-N-Accuracy is the proportion of predicted answers that match with their corresponding correct answer including no_answers
print(f"Reader Top-{top_n}-Accuracy:", reader_eval_results["top_n_accuracy"])
# Reader Top-1-Exact Match is the proportion of questions where the first predicted answer is exactly the same as the correct answer including no_answers
print("Reader Top-1-Exact Match:", reader_eval_results["EM"])
# Reader Top-1-F1-Score is the average overlap between the first predicted answers and the correct answers including no_answers
print("Reader Top-1-F1-Score:", reader_eval_results["f1"])
# Reader Top-N-Accuracy is the proportion of predicted answers that match with their corresponding correct answer excluding no_answers
print(f"Reader Top-{top_n}-Accuracy (without no_answers):", reader_eval_results["top_n_accuracy_text_answer"])
# Reader Top-N-Exact Match is the proportion of questions where the predicted answer within the first n results is exactly the same as the correct answer excluding no_answers (no_answers are always present within top n).
print(f"Reader Top-{top_n}-Exact Match (without no_answers):", reader_eval_results["top_n_EM_text_answer"])
# Reader Top-N-F1-Score is the average overlap between the top n predicted answers and the correct answers excluding no_answers (no_answers are always present within top n).
print(f"Reader Top-{top_n}-F1-Score (without no_answers):", reader_eval_results["top_n_f1_text_answer"])
# -
# Just as a sanity check, we can compare the top-n exact_match and f1 metrics from `reader.eval()` with the exact_match and f1 from `pipeline.eval(add_isolated_node_eval=True)`.
# These two approaches return the same values because pipeline.eval() calculates top-n metrics per default. Small discrepancies might occur due to string normalization in pipeline.eval()'s answer-to-label comparison. reader.eval() does not use string normalization.
metrics = eval_result_with_upper_bounds.calculate_metrics(eval_mode="isolated")
print(metrics["Reader"]["exact_match"])
print(metrics["Reader"]["f1"])
# + [markdown] id="8QJ68G12U7tb" pycharm={"name": "#%% md\n"}
# ## About us
#
# This [Haystack](https://github.com/deepset-ai/haystack/) notebook was made with love by [deepset](https://deepset.ai/) in Berlin, Germany
#
# We bring NLP to the industry via open source!
# Our focus: Industry specific language models & large scale QA systems.
#
# Some of our other work:
# - [German BERT](https://deepset.ai/german-bert)
# - [GermanQuAD and GermanDPR](https://deepset.ai/germanquad)
# - [FARM](https://github.com/deepset-ai/FARM)
#
# Get in touch:
# [Twitter](https://twitter.com/deepset_ai) | [LinkedIn](https://www.linkedin.com/company/deepset-ai/) | [Slack](https://haystack.deepset.ai/community/join) | [GitHub Discussions](https://github.com/deepset-ai/haystack/discussions) | [Website](https://deepset.ai)
#
# By the way: [we're hiring!](https://www.deepset.ai/jobs)
| tutorials/Tutorial5_Evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="ojm_6E9f9Kcf"
# # Refactor
#
# Start with notebook 306 and refactor the code.
# Code improvement had been deferred during SOARS project.
# + colab={"base_uri": "https://localhost:8080/"} id="RmPF4h_YI_sT" outputId="a1edd650-41ca-4e7e-e543-a8e36a0b40c9"
import time
def show_time():
t = time.time()
print(time.strftime('%Y-%m-%d %H:%M:%S %Z', time.localtime(t)))
show_time()
# + id="VQY7aTj29Kch"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.model_selection import KFold
from sklearn.model_selection import cross_val_score
from sklearn.metrics import roc_curve
from sklearn.metrics import roc_auc_score
from keras.models import Sequential
from keras.layers import Dense,Embedding,Dropout
from keras.layers import Flatten,TimeDistributed
from keras.losses import BinaryCrossentropy
from keras.callbacks import ModelCheckpoint
from keras.models import load_model
# + colab={"base_uri": "https://localhost:8080/"} id="xUxEB53HI_sk" outputId="50c1004f-ef2f-4f51-d2d6-358a70a9a4fb"
import sys
IN_COLAB = False
try:
from google.colab import drive
IN_COLAB = True
except:
pass
if IN_COLAB:
print("On Google CoLab, mount cloud-local file, get our code from GitHub.")
PATH='/content/drive/'
#drive.mount(PATH,force_remount=True) # hardly ever need this
drive.mount(PATH) # Google will require login credentials
DATAPATH=PATH+'My Drive/data/' # must end in "/"
import requests
r = requests.get('https://raw.githubusercontent.com/ShepherdCode/Soars2021/master/SimTools/RNA_describe.py')
with open('RNA_describe.py', 'w') as f:
f.write(r.text)
from RNA_describe import ORF_counter
r = requests.get('https://raw.githubusercontent.com/ShepherdCode/Soars2021/master/SimTools/GenCodeTools.py')
with open('GenCodeTools.py', 'w') as f:
f.write(r.text)
from GenCodeTools import GenCodeLoader
r = requests.get('https://raw.githubusercontent.com/ShepherdCode/Soars2021/master/SimTools/KmerTools.py')
with open('KmerTools.py', 'w') as f:
f.write(r.text)
from KmerTools import KmerTools
r = requests.get('https://raw.githubusercontent.com/ShepherdCode/Soars2021/master/SimTools/DataPrep.py')
with open('DataPrep.py', 'w') as f:
f.write(r.text)
from DataPrep import DataPrep
else:
print("CoLab not working. On my PC, use relative paths.")
DATAPATH='data/' # must end in "/"
sys.path.append("..") # append parent dir in order to use sibling dirs
from SimTools.RNA_describe import ORF_counter
from SimTools.GenCodeTools import GenCodeLoader
from SimTools.KmerTools import KmerTools
from SimTools.DataPrep import DataPrep
BESTMODELPATH=DATAPATH+"BestModel"
LASTMODELPATH=DATAPATH+"LastModel"
# + [markdown] id="8buAhZRfI_sp"
# ## Data Load
# + colab={"base_uri": "https://localhost:8080/"} id="h94xptH1tI82" outputId="1768facb-1428-432e-bdb0-e16ae90109f4"
PC_TRAINS=1000
NC_TRAINS=1000
PC_TESTS=1000
NC_TESTS=1000
PC_LENS=(200,4000)
NC_LENS=(200,4000) # Wen used 3500 for hyperparameter, 3000 for train
PC_FILENAME='gencode.v38.pc_transcripts.fa.gz'
NC_FILENAME='gencode.v38.lncRNA_transcripts.fa.gz'
PC_FULLPATH=DATAPATH+PC_FILENAME
NC_FULLPATH=DATAPATH+NC_FILENAME
MAX_K = 3
INPUT_SHAPE=(None,84) # 4^3 + 4^2 + 4^1
NEURONS=32
DROP_RATE=0.30
EPOCHS=200
SPLITS=3
FOLDS=3
show_time()
# + colab={"base_uri": "https://localhost:8080/"} id="VNnPagXjtI85" outputId="5f0a0e19-625d-4348-ae2c-959322fc1628"
loader=GenCodeLoader()
loader.set_label(1)
loader.set_check_utr(True) # ORF-restricted
loader.set_check_size(*PC_LENS) # length-restricted
pcdf=loader.load_file(PC_FULLPATH)
print("PC seqs loaded:",len(pcdf))
loader.set_label(0)
loader.set_check_utr(False) # length-restricted
loader.set_check_size(*NC_LENS)
ncdf=loader.load_file(NC_FULLPATH)
print("NC seqs loaded:",len(ncdf))
show_time()
# + colab={"base_uri": "https://localhost:8080/"} id="ShtPw_fGtI9E" outputId="7e2cef66-c962-4fc3-c907-eb2170fa5901"
def dataframe_extract_sequence(df):
return df['sequence'].tolist()
pc_all = dataframe_extract_sequence(pcdf)
nc_all = dataframe_extract_sequence(ncdf)
pcdf=None
ncdf=None
show_time()
print("PC seqs pass filter:",len(pc_all),type(pc_all))
print("NC seqs pass filter:",len(nc_all),type(nc_all))
# Garbage collection to reduce RAM footprint
#PC seqs pass filter: 55381
#NC seqs pass filter: 46919
# -
print("Simulated sequence characteristics:")
oc = ORF_counter()
print("PC seqs")
oc.describe_sequences(pc_all)
print("NC seqs")
oc.describe_sequences(nc_all)
oc=None
show_time()
# + [markdown] id="CCNh_FZaI_sv"
# ## Data Prep
# -
dp = DataPrep()
Xseq,y=dp.combine_pos_and_neg(pc_all,nc_all)
nc_all=None
pc_all=None
nc_all=None
print("The first few shuffled labels:")
print(y[:30])
show_time()
Xfrq=KmerTools.seqs_to_kmer_freqs(Xseq,MAX_K)
Xseq = None
y=np.asarray(y)
show_time()
# Assume X and y were shuffled.
train_size=PC_TRAINS+NC_TRAINS
X_train=Xfrq[:train_size]
X_test=Xfrq[train_size:]
y_train=y[:train_size]
y_test=y[train_size:]
print("Training set size=",len(X_train),"=",len(y_train))
print("Reserved test set size=",len(X_test),"=",len(y_test))
Xfrq=None
y=None
show_time()
# + [markdown] id="dJ4XhrzGI_s-"
# ## Load a trained neural network
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="BM6UjBzrrEAV" outputId="4d7ffa1e-f349-40ef-ee05-a3a8b038c3c7"
show_time()
model = load_model(BESTMODELPATH)
print(model.summary())
# + [markdown] id="OsytC9VUrEAX"
# ## Test the neural network
# + id="8hIqe1r1rEAa"
def show_test_AUC(model,X,y):
ns_probs = [0 for _ in range(len(y))]
bm_probs = model.predict(X)
ns_auc = roc_auc_score(y, ns_probs)
bm_auc = roc_auc_score(y, bm_probs)
ns_fpr, ns_tpr, _ = roc_curve(y, ns_probs)
bm_fpr, bm_tpr, _ = roc_curve(y, bm_probs)
plt.plot(ns_fpr, ns_tpr, linestyle='--', label='Guess, auc=%.4f'%ns_auc)
plt.plot(bm_fpr, bm_tpr, marker='.', label='Model, auc=%.4f'%bm_auc)
plt.title('ROC')
plt.xlabel('False Positive Rate')
plt.ylabel('True Positive Rate')
plt.legend()
plt.show()
print("%s: %.2f%%" %('AUC',bm_auc*100.0))
def show_test_accuracy(model,X,y):
scores = model.evaluate(X, y, verbose=0)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# -
print(type(X_test))
print(type(X_test[0]))
print(type(X_test[0][0]))
print(X_test[0])
#<class 'numpy.ndarray'>
#<class 'numpy.ndarray'>
#<class 'numpy.float64'>
print()
print(type(y_test))
print(type(y_test[0]))
# + colab={"base_uri": "https://localhost:8080/", "height": 469} id="tGf2PcxRC8jT" outputId="489f5c26-0ff9-4f02-f078-39e3abcc51ba"
print("Accuracy on test data.")
show_time()
show_test_AUC(model,X_test,y_test)
show_test_accuracy(model,X_test,y_test)
show_time()
# + id="-cSLTNfzrEAo"
| Notebooks/Jas_309_Refactor.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from recap_argument_graph import Graph, Node, Edge, NodeCategory
from pathlib import Path
import glob, json
import pandas as pd
def toDataFrame(graph:Graph):
data = {'child_text':[], 'parent_text':[], 'stance':[]}
for child, link in graph._outgoing_nodes.items():
if child.category.name != "I":
continue
else:
# if node does not exist
if len(link) == 0:
continue
else:
# per definition there is only one s_node between two i_nodes
s_node = [n for n in link][0]
link2 = graph._outgoing_nodes[s_node]
parent = [n for n in link2][0]
data['child_text'].append(child.text)
data['stance'].append(s_node.category.name)
data["parent_text"].append(parent.text)
df = pd.DataFrame(data=data)
df.reset_index(inplace=True)
return df
# +
english = glob.glob("kialo_graphs/english/*.json")
dataframes = []
for discussion in english:
graph = Graph.open(Path(discussion))
dataframe = toDataFrame(graph)
dataframes.append(dataframe)
df_english = pd.concat(dataframes)
df_english
# -
df_english.to_csv("english_stances.csv")
# +
deutsch = glob.glob("kialo_graphs/deutsch/*.json")
dataframes = []
for discussion in deutsch:
graph = Graph.open(Path(discussion))
dataframe = toDataFrame(graph)
dataframes.append(dataframe)
df_deutsch = pd.concat(dataframes)
df_deutsch
# -
df_deutsch.to_csv("deutsch_stances.csv")
| recap_am/preprocessing/graph_to_dataframe.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.1
# language: julia
# name: julia-0.4
# ---
push!(LOAD_PATH, pwd());
using TSSM
V(x) = x^2
m = SchroedingerReal1D(1024, -30, 30, potential=V, cubic_coupling=390)
psi=wave_function(m)
f(x)=exp(-x^2)
set!(psi, f)
include("groundstate.jl")
groundstate!(psi, extrapolation_order=2)
using PyPlot
x=get_nodes(m);
u=get_data(psi,true)
to_real_space!(psi)
plot(x, u[1:end-2].^2)
| examples/GroundstateDemo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''base'': conda)'
# name: python3
# ---
# # Project Introduction
# # Ayiti Analytics Probability and Statistics Project
# Use all the files to have a final datasets to have the following columns
# <ul>
# <li>questid</li>
# <li>gender</li>
# <li>age (age of applicant)</li>
# <li>communeName (use data prep in commune file)</li>
# <li>application_date (created at in quest file)</li>
# <li>enroll_date (created_at in enroll file)</li>
# <li>is_enroll (Yes/No) (use data prep in enroll file)</li>
# <li>Education Level</li>
# <li>Communication channels(hear_AA1)</li>
# <li>Bootcamp Insterest (after_AA)</li>
# <li>Payement Date (use ord and transaction files)</li>
# <li>Payed (Yes/No)</li>
# <li>list Technologies as columns based (use get_dummies)</li>
# <li>list Study domains (use get_dummies)</li>
# <li>Job is formal</li>
# <li>Have computer at home</li>
# <li>Have internet at home</li>
#
# </ul>
#
# ### Data Wrangling
# #### Importing Python Packages
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from datetime import date
from scipy import stats
from scipy.stats import chi2_contingency
from scipy.stats import f_oneway
from scipy.stats import wilcoxon
# -
# #### Loading the datasets.
commune=pd.read_excel("commune.xlsx")
enroll = pd.read_csv("enroll.csv")
quest = pd.read_csv("quest.csv")
industry = pd.read_csv("industry.csv")
ord = pd.read_csv("ord.csv")
study_domain = pd.read_csv("study_domain.csv")
transaction = pd.read_csv("transaction.csv")
technology = pd.read_csv("technology.csv")
# #### - Displaying informations about the dataset
quest.info()
# #### - Merging the questionnaire dataset and the Commune dataset
# +
quest['department'] = quest['department'].apply(lambda x : str(x))
quest['department']= quest['department'].apply(lambda x : x.upper())
quest['commune']= quest['commune'].apply(lambda x : x.upper())
quest
first_tab=pd.merge(quest,commune, how = 'left', left_on=['department','commune'], right_on=['ADM1_PCODE','Commune_Id'])
first_tab.head()
# -
# #### -Creating a column for the applicants that paid to access the course .
# +
transaction['Paid'] = 'Yes'
ord['Paid'] = 'Yes'
x = transaction.loc[:,['Paid','user_id','created_at']]
y = ord.loc[:,['Paid','user_id','created_at']]
trans_ord= pd.concat([x,y],axis=0)
trans_ord= trans_ord.rename(columns={'created_at': 'Payment_Date'})
trans_ord.head()
# -
# #### - Merging the enroll dataset and transactions datasets
# +
enroll.head()
enroll= enroll.rename(columns={'created_at': 'enroll_date'})
enroll['is_enroll'] = 'Yes'
scnd_tab=pd.merge(enroll,trans_ord, how = 'left',on = ['user_id'] )
scnd_tab= scnd_tab.loc[:,['user_id','quest_id','enroll_date','Paid','Payment_Date','is_enroll']]
scnd_tab
# -
# #### -Merging The modified quest dataset and the modified enroll dataset
third_tab=pd.merge(left=first_tab,right=scnd_tab, how = 'left', on=['quest_id'])
third_tab['is_enroll']= third_tab['is_enroll'].fillna('no')
third_tab['Paid']= third_tab['Paid'].fillna('no')
third_tab.head()
# #### - Converting Quantitative variables into dummies. Using study domain and technology Dataset
# ##### A-
study_domain1 = pd.get_dummies(data=study_domain[["quest_id", "values"]], columns=['values'], prefix="", prefix_sep="")
study_domain1=study_domain1.groupby("quest_id").sum()
study_domain1
fourth_tab=pd.merge(left=third_tab,right=study_domain1, how = 'left', on=['quest_id'])
fourth_tab
# ##### B-
# +
technologyy = pd.get_dummies(data=technology[["quest_id", "values"]], columns=['values'], prefix="", prefix_sep="")
technologyyy=technologyy.groupby("quest_id").sum()
fifth_tab=pd.merge(left=fourth_tab,right=technologyyy, how = 'left', on=['quest_id'])
fifth_tab
# -
# #### - Dropping unecessary columns
# +
columns_drop= ['Unnamed: 0','modified_at','current_employed','Commune_Id','Commune_FR','ADM1_PCODE','study_domain','department','university','commune']
# -
# #### - Creating our final dataset
final_tab= fifth_tab.drop(labels=None,columns=columns_drop, level=None, inplace=False)
y = pd.DataFrame(final_tab.created_at.str.split('T',1).tolist(),columns = ['Date_created_at','To_drop'])
final_tab['created_at'] = y.iloc[:,0]
# ### Transforming Data
# #### A- Creating the Age columns
# +
final_tab['dob'] = final_tab['dob'].astype(str)
final_tab['dob'].replace({'3 aout 1977':'03/08/1977'},inplace = True)
final_tab['dob'] = pd.to_datetime(final_tab['dob'])
final_tab['created_at'] = pd.to_datetime(final_tab['created_at'])
final_tab['Age'] =final_tab['created_at'] - final_tab['dob']
final_tab['Age'] = final_tab['Age']//np.timedelta64(1,"Y")
# -
drop_data = final_tab[(final_tab['Age'] == 1.0) | (final_tab['Age'] == 0) |(final_tab['Age'] == -1)]
final_tab.drop(index= drop_data.index, inplace= True)
final_tab[(final_tab['Age'] ==1) | (final_tab['Age'] ==0) |(final_tab['Age'] ==-1)]
drop_data['Age'].replace({(1,0,-1):final_tab['Age'].mean()},inplace = True)
drop_data = drop_data.reset_index()
drop_data = drop_data.drop(columns= 'index')
final_tab = pd.concat([final_tab,drop_data], axis = 0)
final_tab = final_tab.reset_index()
final_tab =final_tab.drop(columns= 'index')
move = final_tab.pop('Age')
final_tab.insert(2,'Age',move)
final_tab['Age'] = final_tab['Age'].fillna(final_tab['Age'].mean())
final_tab['Age'] = final_tab['Age'].astype(float)
#final_tab = final_tab.drop(columns=['user_id','dob'],axis=1)
final_tab.Age.value_counts()
# #### - Removing the timestamp
# +
date_list=['created_at','enroll_date','Payment_Date']
for i in lisst:
final_tab[i] =final_tab[i].apply(lambda x : str(x).split("T")[0])
final_tab[i] =final_tab[i].apply(lambda x : pd.to_datetime(x))
# -
cols=final_tab.iloc[:,16:].columns
for i in cols:
final_tab[i]= final_tab[i].fillna(0.0)
final_tab.info()
# ### 1. How many observations and variables are there in the dataset
#
# here your codes
final_tab.shape
print('The number of observations in the dataset is:',final_tab.shape[0])
print('The number of variables in the dataset is:',final_tab.shape[1])
# ### 2.A Calculate the average age of the observations
# * On the whole dataset
# * On the whole male dataset
# * On all the female dataset
# ### 2.B Calculate
# * variance of the whole datatset,male and female
# * kurtosis of the whole datatset,male and female
# * skewness of the whole datatset,male and female
# ### 2.C Does the age distribution follow a normal distribution
#
# ### 2.A Calculate the average age of the observations
# +
# here your codes
avg=final_tab['Age'].mean()
print('The average age of the observations in the whole dataset is', np.round(avg,2), 'years old')
avg_male=final_tab[final_tab['gender']== 'male'].Age.mean()
print('The average age of the observations for the male dataset is', np.round(avg_male,2), 'years old')
avg_female=final_tab[final_tab['gender']== 'female'].Age.mean()
print('The average age of the observations for the female dataset is', np.round(avg_female,2), 'years old','\n')
# -
# ### 2.B Calculate variance of the whole datatset,male and female
# +
var=final_tab['Age'].var()
print('The age variance of the observations in the whole dataset is', np.round(var,2))
var_male=final_tab[final_tab['gender']== 'male'].Age.var()
print('The age variance of the observations for the male is', np.round(var_male,2))
var_female=final_tab[final_tab['gender']== 'female'].Age.var()
print('The age variance of the observations for the female is', np.round(var_female,2),'\n')
# -
# ### 2.B Calculate the kurtosis of the whole datatset,male and female
# +
kurtosis=final_tab['Age'].kurtosis()
print('The kurtosis of the age distribution the whole dataset is', np.round(kurtosis,2))
kurtosis_male=final_tab[final_tab['gender']== 'male'].Age.kurtosis()
print('The kurtosis of the age distribution for the male is', np.round(kurtosis_male,2))
kurtosis_female=final_tab[final_tab['gender']== 'female'].Age.kurtosis()
print('The kurtosis of the age distributionfor the female is', np.round(kurtosis_female,2),'\n')
# -
# ### 2.B Calculate the skewness of the whole datatset,male and female
skewness=final_tab['Age'].skew()
print('The skewness age of the distribution in the whole dataset is', np.round(skewness,2))
skewness_male=final_tab[final_tab['gender']== 'male'].Age.skew()
print('The skewness age of the distribution for the male is', np.round(skewness_male,2))
skewness_female=final_tab[final_tab['gender']== 'female'].Age.skew()
print('The skewness age of the distribution for the female is', np.round(skewness_female,2),'\n')
# ### 2.C Does the age distribution follow a normal distribution
# +
#Testing the distribution for normality
from scipy.stats import shapiro
statistic,pvalue=shapiro(final_tab['Age'])
alpha = 0.05
print('The Shapiro Wilk Test for the whole dataset')
print(pvalue)
if pvalue > alpha:
print('ho Distribution follows a gaussian distribution (fail to reject H0)')
else:
print('h1 Distribution doesnt follows a gaussian distribution (reject H0)')
# -
statistic,pvalue =shapiro(final_tab[final_tab['gender']== 'male'].Age)
print('The Shapiro Wilk Test for the male dataset')
print(pvalue)
if pvalue > alpha:
print('ho Distribution follows a gaussian distribution (fail to reject H0)')
else:
print('h1 Distribution doesnt follows a gaussian distribution (reject H0)')
statistic,pvalue =shapiro(final_tab[final_tab['gender']== 'female'].Age)
print('The Shapiro Wilk Test for the female dataset')
print(pvalue)
if pvalue > alpha:
print('ho Distribution follows a gaussian distribution (fail to reject H0)')
else:
print('h1 Distribution doesnt follows a gaussian distribution (reject H0)')
# ### 3.Display the age frequency disbrution
# * On the whole dataset
# * On the whole male dataset
# * On all the female dataset
bins = 50
sns.displot(final_tab['Age'],bins = bins,kde = True)
sns.displot(final_tab[final_tab['gender']== 'male'].Age, bins = bins,kde = True)
sns.displot(final_tab[final_tab['gender']== 'female'].Age,bins= bins,kde = True)
# ### 4. Can we say that the average age (24 years old) of the observations can be considered as the average age of the population likely to participate in this bootcamp. Justify your answer
# +
#here your codes
# H0 : Mu = 24
# H1 : Mu != 24
stat, p = wilcoxon(final_tab.Age - 24)
print('Statistics=%.2f, p=%.2f' % (stat, p))
# interpretation
alpha = 0.05
if p > alpha:
print('ho mu=24 (fail to reject H0)')
else:
print('h1 mu!= 24 (reject H0)')
# -
# #### No, We cannot say that 24 years can be considered as the average of the population likely to participate in this bootcamp. By realizing a Wilcoxon test , the p-value obtained (0.00) is lower than the significance level 0.05. The age difference of the population is different than 24 years old.
# ### 5. Calculate the average age of participants for each communication channel
#here your codes
display( final_tab['Age'].groupby(by= final_tab['hear_AA_1']).mean())
# ### 6. Display an age boxplot for each communication channel
#here your codes
plt.figure(figsize=(15,6))
sns.boxplot(data=final_tab,x="hear_AA_1",y="Age")
plt.title('Boxplot of applicants age by communication channels')
# ### 7 .Is there a significant age difference between these groups
# +
Facebook_list_age = final_tab[final_tab['hear_AA_1'] == 'Facebook']['Age'].values
Unknown_list_age = final_tab[final_tab['hear_AA_1'] == 'unknown']['Age'].values
Friend_list_age = final_tab[final_tab['hear_AA_1'] == 'Friend']['Age'].values
ESIH_list_age = final_tab[final_tab['hear_AA_1'] == 'ESIH']['Age'].values
WhatsApp_list_age = final_tab[final_tab['hear_AA_1'] == 'WhatsApp']['Age'].values
Bootcamp_Alumni_list_age = final_tab[final_tab['hear_AA_1'] == 'Bootcamp Alumni']['Age'].values
LinkedIn_list_age = final_tab[final_tab['hear_AA_1'] == 'LinkedIn']['Age'].values
Ayiti_Analytics_Website_list_age = final_tab[final_tab['hear_AA_1'] == 'Ayiti Analytics Website']['Age'].values
other_list_age = final_tab[final_tab['hear_AA_1'] == 'other']['Age'].values
Instagram_list_age = final_tab[final_tab['hear_AA_1'] == 'Instagram']['Age'].values
# +
#here your codes
# Ho : There is no significant age difference between these groups
# h1 : There is a significant age difference between the mean of these groups
stat, p = f_oneway(Facebook_list_age, Unknown_list_age, Friend_list_age, ESIH_list_age, WhatsApp_list_age, Bootcamp_Alumni_list_age, LinkedIn_list_age, Ayiti_Analytics_Website_list_age, other_list_age, Instagram_list_age)
print(p)
if p < 0.05:
print("Hypothesis Rejected : There is a significant age difference between the mean of these groups")
else:
print("Hypothesis Accepted :There is no significant age difference between these groups ")
# -
# ### 8.Plan a BA strategy for each communication channel regarding the age group
# * Do content marketing
#
# Content marketing is a marketing program that centers on creating, publishing, and distributing content for your target audience -- usually online -- the goal of which is to attract new customers.
#
#
# The application process for the bootcamp is 100% online. The announcements were made on the social networks of Ayiti Analytics. 56.4% of the applicants say they heard about AA via social networks and 26.8% of the applicants hear about AA from their friends who might also have heard about it on social media, but we don't have enough data to confirm this. AA has the team and the technology, the next step is to create more content that can attract applicants like a blog post, videos, infographics,newsletters. And also use email marketing to remain in touch with subscribers
#
#
#
# ### 9.According to the observations what is the probability of being in each channel of communication knowing your are a woman
# #### Displaying the proportion of male and female in the dataset
res =final_tab.gender.value_counts(normalize=True).to_frame()
display(res)
# #### Displaying the probability of being one of the genders and being in one of the communications channels
# +
#here your codes
gender_table = pd.pivot_table(final_tab,'Age',index='hear_AA_1',columns=['gender'],aggfunc = ['count'],fill_value=0)
my_pivot = gender_table.sort_values(by= ('count','female'),ascending = False)
my_pivot['Probabilty of being female and being in one of the communications channels'] =gender_table[('count','female')]/final_tab.shape[0]
my_pivot['Probabilty of being male and being in one of the communications channels'] = gender_table[('count','male')]/final_tab.shape[0]
my_pivot=my_pivot.iloc[:,[2,3]]
display(my_pivot)
# -
my_pivot['female'] =my_pivot['Probabilty of being female and being in one of the communications channels']/ res.loc["female","gender"]
my_pivot.loc[:,['female']]
# ### 10.According to the observations what is the probability of being in each channel of communication knowing your are a man
#here your codes
my_pivot['male'] =my_pivot['Probabilty of being male and being in one of the communications channels']/ res.loc["male","gender"]
my_pivot
# ### 11. Deduce the probability of being a woman knowing each communication channel
#here your codes
communication_table = pd.pivot_table(final_tab,'Age',index='hear_AA_1',columns='gender',aggfunc = ['count'],fill_value=0,margins = True)
communication_table= communication_table.iloc[:-1,-1].to_frame()
communication_table[('count','All')] = communication_table[('count','All')]/final_tab.shape[0]
#Using Bayes Theorem to deduce the probabilty
communication_table['Probability of being in each channel of communication knowing your are a woman']= my_pivot.loc[:,['female']]
communication_table['Probability of being in each channel of communication knowing your are a man']= my_pivot.loc[:,['male']]
communication_table['Probability of being a female knowing each communication channels']= (communication_table['Probability of being in each channel of communication knowing your are a woman'] * (0.188))/communication_table[('count','All')]
communication_table['Probability of being a male knowing each communication channels']= (communication_table['Probability of being in each channel of communication knowing your are a man'] * (0.188))/communication_table[('count','All')]
communication_table.iloc[:,3]
# ### 12. Deduce the probability of being a man knowing each communication channel
#here your codes
communication_table.iloc[:,4]
# ### 13 Display a plot to see Gender vs Communication Channels .Is there any dependency between communication channels and gender?
#here
def my_chisqure_test(data=final_tab,columns=None,significance_level=0.05,figsize=(5,5),heatmap=False):
result1 = pd.pivot_table(data=data,index=columns[0],columns=columns[1],values="Age",aggfunc="count",fill_value=0,margins = True)
display(result1)
stat, p, dof,s= chi2_contingency(result1)
print("p value: " + str(p))
if p <= significance_level:
print('Reject NULL HYPOTHESIS')
else:
print('ACCEPT NULL HYPOTHESIS : The variables are independent')
# Force the aspect ratio of the blocks to be equal using "square" parameter
plt.figure(figsize=figsize)
if heatmap:
ax = sns.heatmap(result1,cmap="YlGnBu", linewidths=.5,
annot=True ,annot_kws={'size':14} ,fmt=".1f" , cbar=True ,square = True)
plt.yticks(rotation=0)
else:
values = result1.iloc[:-1,:].sort_values(by =('male'),ascending=True)
ax =values.sort_values(by =('male'),ascending=True)[[('male'),('female')]].plot(kind="barh",figsize=(15,10),stacked= True,alpha =0.7)
ylab = ax.set_ylabel('Number of Applicants')
xlab = ax.set_xlabel('Communications Channels')
title = ax.set_title('Which communications channels did the applicants hear of AA ?')
index =0
for male, female ,total in zip(values[('male')],values[('female')], values[('All')]):
if male != total and female != total :
ax.text(male/2 ,
index,
str(np.round((male/total)*100,1)) + "%",
fontdict=dict(color='white',fontsize=10,horizontalalignment="center",verticalalignment="center")
)
ax.text(male + female/2 ,
index,
str(np.round((female/total)*100,1)) + "%",
fontdict=dict(color='blue',fontsize=10,horizontalalignment="center",verticalalignment="center")
)
elif female == total:
ax.text(female/2 ,
index,
str(np.round((female/total)*100,1)) + "%",
fontdict=dict(color='blue',fontsize=10,horizontalalignment="center",verticalalignment="center")
)
else:
ax.text(male/2 ,
index,
str(np.round((male/total)*100,1)) + "%",
fontdict=dict(color='white',fontsize=10,horizontalalignment="center",verticalalignment="center")
)
index+=1
plt.show()
return result1
my_chisqure_test(final_tab,columns=['hear_AA_1','gender'],significance_level=0.05,figsize=(5,15),heatmap=False)
# ### 16 Use the same method to display plot and know if is there any dependency between communication channels and Bootcamp Insterest?
# here your codes
def my_chisqure_test(data=final_tab,columns=None,significance_level=0.05,figsize=(5,5),heatmap=False):
result1 = pd.pivot_table(data=data,index=columns[0],columns=columns[1],values="Age",aggfunc="count",fill_value=0)
stat, p, dof,s= chi2_contingency(result1)
print("p value: " + str(p))
if p <= significance_level:
print('There is not independecy between the variables :Reject NULL HYPOTHESIS')
else:
print('There is independency ACCEPT NULL HYPOTHESIS')
# Force the aspect ratio of the blocks to be equal using "square" parameter
plt.figure(figsize=figsize)
if heatmap:
ax = sns.heatmap(result1,cmap="YlGnBu", linewidths=.5,
annot=True ,annot_kws={'size':14} ,fmt=".1f" , cbar=True ,square = True)
plt.yticks(rotation=0)
else:
ax =result1.sort_values(by='Improving my data analysis skills').plot(kind="barh",stacked=True,figsize=(10,4))
plt.show()
return result1
my_chisqure_test(final_tab,columns=['hear_AA_1','after_AA'],significance_level=0.05,figsize=(5,15),heatmap=False)
# ### 17.Plan a BA strategy for each communication channel, Bootcamp Insterest regarding the gender
# Find the right approach for each gender :
#
# In order to determine the ideal target group, the interest of the target should be taken into account with the insight provided by the dataset,we can assert that there is dependency between the two variables of Bootcamp interest and communication channels .But it is also important to include other factors for a successful campaign. In the case of gender-specific advertising messages, it is important to avoid clichés.
#
#
# This is especially advisable for gender-related campaigns, such as Women’s Day. Women are particularly well-received if the content is conveyed by emotions, people or family. We can assert it with friends and bootcamp alumni being the top communication channels by the women In contrast, male users prefer campaigns with action and competitive situations. The campaign content should also be transferred to the visual level: clear, bright newsletters with an emotional design are appealing to female customers, while men find a rational approach with technical details and comparisons appealing.Whatsapp and friend are the top communicationg channels for men
#
# ### 19. Calculate
# * P(Bootcamp Insterest,Communication channels,Gender/Payed=yes)
# * P(Bootcamp Insterest,Communication channels,Gender/Payed=no)
result12 = pd.pivot_table(data=final_tab,columns='after_AA',index=['Paid','hear_AA_1','gender'],values="quest_id",aggfunc="count",fill_value=0,margins = True)
result12 = result12 /final_tab[final_tab['Paid'] == 'Yes'].shape[0]
result12
result123 = pd.pivot_table(data=final_tab[final_tab['Paid'] == 'no'],columns='after_AA',index=['Paid','hear_AA_1','gender'],values="quest_id",aggfunc="count",fill_value=0,margins = True)
result123 = result123 /final_tab[final_tab['Paid'] == 'no'].shape[0]
result123
# ### 20 reduce
# * P(Payed="yes"/Bootcamp Insterest="Increase Skill",Communication channels="Friend",Gender="male")
# * P(Payed="no"/Bootcamp Insterest="Increase Skill",Communication channels="Friend",Gender="male")
# here your codes
m = pd.pivot_table(data=final_tab,columns='after_AA',index=['hear_AA_1','gender'],values="quest_id",aggfunc="count",fill_value=0,margins = True)
m=final_tab[final_tab['Paid'] == 'Yes'].shape[0]/m
g = result12*m
g = g.fillna("-")
display(g)
g.loc[pd.IndexSlice['Friend','male'],'Improving my data analysis skills']
# ### Based on these findings, propose strategies to increase our sales?
#
# * 1 - Recognize the customer's problems or needs
# It all starts with understanding your customers.With the insights obtained,we can see where the bootcamp interest can play a big part with attracting customers. Knowing that the bootcamp could upscale their skills or open doors on the professional world.
#
#
# * 2- People Buy Benefits
#
# People don’t buy products, they buy the results that product will give. AA have to start the process of identifying their ideal customer by making a list of all of the benefits that their customers will enjoy by using their products or services.After participating in the bootcamp, the applicants want to Improving their data skills , Finding an internship or start their own company after learning all the job-ready skills
#
#
#
# * 3- Develop Your Competitive Advantage
#
# You define your competitive advantage, the reason for buying your products or services, in terms of the benefits, results or outcomes that your customer will enjoy from purchasing your product or service that they would not fully enjoy from purchasing the product or service of your competitor. AA is for now the only company providing this services in Haiti, Extending their market should be the next move for them to be able to reach out more young people that loves tech in haiti.(refers to first Python Project)
#
# Focus on the benefits of what makes your product better than others.
#
#
| data (2).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.12 ('ds_study')
# language: python
# name: python3
# ---
# +
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LogisticRegression
import statsmodels.api as sm
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
COVID = pd.read_excel('./owid-covid-data.xlsx')
COVID = COVID[COVID['location']=='South Korea']
COVID = COVID.fillna(0)
COVID.head()
# -
COVID.tail()
COVID = COVID[['total_cases', 'new_cases', 'new_cases_smoothed', 'total_deaths', 'new_deaths', 'new_deaths_smoothed']]
COVID.head()
COVID.info()
# +
X = COVID.drop(['new_deaths'], axis=1)
y = COVID['new_deaths']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=13)
# +
from sklearn.pipeline import Pipeline
from sklearn.tree import DecisionTreeClassifier
from sklearn.preprocessing import StandardScaler
estimators = [
('scalor', StandardScaler()),
('clf', DecisionTreeClassifier())
]
pipe = Pipeline(estimators)
# -
pipe.set_params(clf__max_depth=2)
pipe.set_params(clf__random_state=13)
pipe.fit(X_train, y_train)
# +
from sklearn.metrics import accuracy_score
y_pred_tr = pipe.predict(X_train)
y_pred_test = pipe.predict(X_test)
print('Train acc :', accuracy_score(y_train, y_pred_tr))
print('Test acc :', accuracy_score(y_test, y_pred_test))
# +
from sklearn.model_selection import GridSearchCV
from sklearn.tree import DecisionTreeClassifier
params = {'max_depth': [i for i in range(4, 14, 2)]}
be_tree = DecisionTreeClassifier(max_depth=2, random_state=13)
# -
GridSearch = GridSearchCV(estimator=be_tree, param_grid=params, cv=5)
GridSearch.fit(X, y)
# +
import pprint
pp = pprint.PrettyPrinter(indent=4)
pp.pprint(GridSearch.cv_results_)
# -
COVIDdr = COVID.drop(['new_deaths', 'new_deaths_smoothed'], axis=1)
# +
import statsmodels.formula.api as smf
lm_model = smf.ols(formula='y ~ X', data=COVIDdr).fit()
# -
COVID.info()
X_train
lm_model.params
COVID
COVID.drop(['iso_code','continent','location','date'], axis=1,inplace=True)
COVID.drop(['tests_units'], axis=1, inplace=True)
COVID.info()
X = COVID.drop(['new_deaths','total_deaths','new_deaths_per_million', 'new_deaths_smoothed', 'new_deaths_smoothed_per_million'], axis=1)
y = COVID['new_deaths']
lm = sm.OLS(y, X).fit()
lm.rsquared
lm.summary()
death = lm.predict(X)
COVIDNew = COVID[['total_cases', 'new_cases', 'new_cases_smoothed','reproduction_rate' , 'hosp_patients','positive_rate' , 'total_vaccinations', 'people_vaccinated', 'people_fully_vaccinated', 'total_boosters', 'stringency_index','human_development_index' , 'excess_mortality_cumulative_per_million', 'new_people_vaccinated_smoothed']]
X = COVIDNew
y = COVID['new_deaths']
lm = sm.OLS(y, X).fit()
lm.rsquared
lm.summary()
dead= lm.predict(X)
COVIDNew['Predict Death'] = dead
COVIDNew['Predict with all'] = death
COVIDNew['Real death'] = COVID['new_deaths']
COVIDNew
lm.aic
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=13)
# +
from sklearn.linear_model import LinearRegression
reg = LinearRegression()
reg.fit(X_train, y_train)
# +
from sklearn.metrics import mean_squared_error
pred_tr = reg.predict(X_train)
pred_test = reg.predict(X_test)
rmse_tr = np.sqrt(mean_squared_error(y_train, pred_tr))
rmse_test = np.sqrt(mean_squared_error(y_test, pred_test))
# -
print('RMSE of Train Data: ', rmse_tr)
print('RMSE of Test Data: ', rmse_test)
# +
def stepwise_feature_selection(X_train, y_train, variables=X_train.columns.tolist() ):
import statsmodels.api as sm
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
y = y_train ## 반응 변수
selected_variables = [] ## 선택된 변수들
sl_enter = 0.05
sl_remove = 0.05
sv_per_step = [] ## 각 스텝별로 선택된 변수들
adjusted_r_squared = [] ## 각 스텝별 수정된 결정계수
steps = [] ## 스텝
step = 0
while len(variables) > 0:
remainder = list(set(variables) - set(selected_variables))
pval = pd.Series(index=remainder) ## 변수의 p-value ## 기존에 포함된 변수와 새로운 변수 하나씩 돌아가면서 ## 선형 모형을 적합한다.
for col in remainder:
X = X_train[selected_variables+[col]]
X = sm.add_constant(X)
model = sm.OLS(y,X).fit(disp=0)
pval[col] = model.pvalues[col]
min_pval = pval.min()
if min_pval < sl_enter: ## 최소 p-value 값이 기준 값보다 작으면 포함
selected_variables.append(pval.idxmin()) ## 선택된 변수들에대해서 ## 어떤 변수를 제거할지 고른다.
while len(selected_variables) > 0:
selected_X = X_train[selected_variables]
selected_X = sm.add_constant(selected_X)
selected_pval = sm.OLS(y,selected_X).fit(disp=0).pvalues[1:] ## 절편항의 p-value는 뺀다
max_pval = selected_pval.max()
if max_pval >= sl_remove: ## 최대 p-value값이 기준값보다 크거나 같으면 제외
remove_variable = selected_pval.idxmax()
selected_variables.remove(remove_variable)
else:
break
step += 1
steps.append(step)
adj_r_squared = sm.OLS(y,sm.add_constant(X_train[selected_variables])).fit(disp=0).rsquared_adj
adjusted_r_squared.append(adj_r_squared)
sv_per_step.append(selected_variables.copy())
else:
break
fig = plt.figure(figsize=(100,10))
fig.set_facecolor('white')
font_size = 15
plt.xticks(steps,[f'step {s}\n'+'\n'.join(sv_per_step[i]) for i,s in enumerate(steps)], fontsize=12)
plt.plot(steps,adjusted_r_squared, marker='o')
plt.ylabel('Adjusted R Squared',fontsize=font_size)
plt.grid(True)
plt.show()
return selected_variables
selected_variables = stepwise_feature_selection(X_train, y_train)
# -
model = sm.OLS(y_train, sm.add_constant(pd.DataFrame(X_train[selected_variables]))).fit(disp=0)
print(model.summary())
COVID
COVID
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=13)
corona_tree = DecisionTreeClassifier(max_depth=4, random_state=13)
corona_tree.fit(X_train, y_train)
y_pred_tr = corona_tree.predict(X_train)
y_pred_test = corona_tree.predict(X_test)
print('Train Acc: ', accuracy_score(y_train, y_pred_tr))
print('Test Acc: ',accuracy_score(y_test, y_pred_test))
# -
dict(zip(X_train.columns, corona_tree.feature_importances_))
X = COVID.drop(['new_deaths','total_deaths','new_deaths_per_million', 'new_deaths_smoothed', 'new_deaths_smoothed_per_million', 'icu_patients_per_million'], axis=1)
y = COVID['new_deaths']
# +
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=13)
corona_tree = DecisionTreeClassifier(max_depth=4, random_state=13)
corona_tree.fit(X_train, y_train)
y_pred_tr = corona_tree.predict(X_train)
y_pred_test = corona_tree.predict(X_test)
print('Train Acc: ', accuracy_score(y_train, y_pred_tr))
print('Test Acc: ',accuracy_score(y_test, y_pred_test))
# -
dict(zip(X_train.columns, corona_tree.feature_importances_))
| ML_Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # About
#
# Run `tcpdump` to see if we can read the raw bytes off the wire and verify endianess of the TCP header.
# ! sudo tcpdump -nn -c1 -X tcp dst port 80
# In the above capture we pass the following options to the `tcpdump` command:
#
# * `-nn` Do not resolve hostnames, don't convert protocol and port numbers etc. to names either.
# * `-c` Capture _x_ packets and then stop (in our case we just capture 1).
# * `tcp` Capture only TCP packets
# * `dst port` Filter only packets going to port 80
# * `-X` Print data of the packet in hex.
# Note that the output of `tcpdump` will change everytime we run it. So for reference here is the ouput that we will use for the remainder of the text:
#
# ```
# tcpdump: verbose output suppressed, use -v or -vv for full protocol decode
# listening on eno1, link-type EN10MB (Ethernet), capture size 262144 bytes
# 14:17:35.632129 IP 10.10.138.125.49166 > 109.238.50.21.80: Flags [S], seq 3608769402, win 29200, options [mss 1460,sackOK,TS val 4978207 ecr 0,nop,wscale 7], length 0
# 0x0000: 4500 003c 8565 4000 4006 80cc 0a0a 8a7d E..<.e@.@......}
# 0x0010: 6dee 3215 c00e 0050 d719 737a 0000 0000 m.2....P..sz....
# 0x0020: a002 7210 34b9 0000 0204 05b4 0402 080a ..r.4...........
# 0x0030: 004b f61f 0000 0000 0103 0307 .K..........
# 1 packet captured
# 2 packets received by filter
# 0 packets dropped by kernel
# 29 packets dropped by interface
# ```
#
# The quick glance we see that the packet captured was going from:
#
# * 10.10.138.125.49166, where 10.10.138.125 is the source IP and 49166 is the source port.
#
# To:
# * 192.168.3.11.80, where 192.168.3.11 is the destination IP and 80 is the destination port.
#
# Good with that in place we can look at the bytes of the packet, it was a bit hard to find but it seems that `tcpdump` prints also the IP header. See format [here](https://en.wikipedia.org/wiki/IPv4).
#
# The first four bits are the IP version used, which is 0x4, so we are using IP version 4. The length of the IP header in 32 bit words is the next four bits. Which is 0x5, so we konw that our TCP header will start $5\cdot32$ bits into the data. This is equivalent to skipping over 40 hex digits.
#
# This means that the first 32 bit word of the TCP header must be `c00e 0050`.
#
# Important note is that memory wise this is stored as (where memory is some pointer to our data):
#
# * `memory[0] = c0;`
# * `memory[1] = 0e;`
# * `memory[2] = 00;`
# * `memory[3] = 50;`
#
# When we interpret the memory we have to be a bit careful, both the source an destination port stored in this 32 bit memory are stored using _network byte order_ or _big endian_ which means that the most significant byte comes first.
#
# * The first 16 bits are the source port `0xc00e` which equals 49166 in decimal.
# * The second 16 bits are the destination port `0x0050` which equals 80 in decimal.
#
# This matches our inital expectation.
# Lets try to parse this with bitter:
# ! git clone <EMAIL>:steinwurf/bitter.git
# We also need to get be able to read the big endian numbers and convert them to the host manchines endianess. This can be done with our endian library:
# ! git clone <EMAIL>:steinwurf/endian.git
# +
# %%file parse_tcp_header.cpp
#include <iostream>
#include <bitter/reader.hpp>
#include <endian/big_endian.hpp>
int main()
{
uint8_t data[] = {0xc0,0x0e,0x00,0x50};
std::cout << "src port = " << endian::big_endian::get<uint16_t>(data) << std::endl;
std::cout << "dest port = " << endian::big_endian::get<uint16_t>(data+2) << std::endl;
return 0;
}
# -
# ! g++ -std=c++11 -Ibitter/src -Iendian/src parse_tcp_header.cpp
# Lets run it:
# ! ./a.out
| examples/tcpdump/tcpdump.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="A0T8R4yTdALH" colab_type="code" outputId="099108b8-3523-4f6a-a1be-86b90c62388c" colab={"base_uri": "https://localhost:8080/", "height": 34}
from keras.datasets import cifar10
import numpy as np
# + id="PlTnKyW5dSEs" colab_type="code" outputId="8ca7b619-acc9-4c19-f37a-d3cfb19b282e" colab={"base_uri": "https://localhost:8080/", "height": 51}
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
# + id="PLBZoTN2dbVO" colab_type="code" outputId="fe42637d-b825-49ad-ef4f-aa5b5b770825" colab={"base_uri": "https://localhost:8080/", "height": 347}
import matplotlib.pyplot as plt
plt.imshow(x_train[30])
plt.show()
# + id="rBfb0Ydid40g" colab_type="code" colab={}
from keras.utils import to_categorical
y_train=to_categorical(y_train)
y_test=to_categorical(y_test)
# + id="so3lT3BXfK2z" colab_type="code" outputId="87ac1ece-c269-4b02-daee-90dbdbf2dc9a" colab={"base_uri": "https://localhost:8080/", "height": 34}
x_train=x_train.reshape(-1,32,32,3)
x_test=x_test.reshape(-1,32,32,3)
x_train.shape,x_test.shape,y_train.shape,y_test.shape
# + id="Co-TNqfefsCg" colab_type="code" colab={}
x_train=x_train/255
x_test=x_test/255
# + id="spRqHud9gT8o" colab_type="code" colab={}
from keras.models import Model
from keras.layers import Input,Conv2D,MaxPooling2D,Flatten,Dense,Dropout
# + id="kid2wZAMgiBH" colab_type="code" colab={}
inputs=Input(shape=(32,32,3))
c1=Conv2D(64,(3,3),padding="same",activation="relu")(inputs)
m1=MaxPooling2D(padding="same")(c1)
drop1=Dropout(0.3)(m1)
c2=Conv2D(64,(3,3),padding="same",activation="relu")(drop1)
m2=MaxPooling2D(padding="same")(c2)
drop2=Dropout(0.3)(m2)
c3=Conv2D(64,(5,5),padding="same",activation="relu")(drop2)
m3=MaxPooling2D(padding="same")(c3)
drop2=Dropout(0.3)(m3)
conv_out=Flatten()(drop2)
d1=Dense(512,activation="relu")(conv_out)
out=Dense(10,activation="softmax")(d1)
# + id="h_b51t4dhpl0" colab_type="code" colab={}
model=Model(inputs=inputs,outputs=out)
model.compile(optimizer="adam",loss="categorical_crossentropy",metrics=["accuracy"])
# + id="elOeZiweewAl" colab_type="code" outputId="5a47d34d-0838-4401-d86e-8b54712ded5a" colab={"base_uri": "https://localhost:8080/", "height": 578}
model.summary()
# + id="ZJkGIXJ6jH8M" colab_type="code" outputId="16a245c2-24fd-4228-c974-e3e1328b9cb9" colab={"base_uri": "https://localhost:8080/", "height": 731}
model.fit(x_train,y_train,batch_size=64,epochs=20,validation_data=(x_test,y_test))
# + id="tDpEIcR-jn3N" colab_type="code" outputId="ce28a036-1626-4935-8c0e-80e7a9a3847f" colab={"base_uri": "https://localhost:8080/", "height": 68}
model.evaluate(x_train,y_train),model.evaluate(x_test,y_test)
| cifar10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Boston house prices prediction using 'scikit-learn':
# The goal of this notebook is to familiarise machine learning concepts using Boston House prices dataset from scikit learn library.
#
# **Introduction:**
# <br>Scikit-learn (formerly scikits.learn and also known as sklearn) is a free software machine learning library for the Python programming language. It features various classification, regression and clustering algorithms including support vector machines, random forests, gradient boosting, k-means and DBSCAN, and is designed to interoperate with the Python numerical and scientific libraries NumPy and SciPy. (Ref: Wikipedia)
#
# **Dependencies:**
# 1. Python 3.7+
# 2. numpy, pandas, scikit-learn libraries installed
# 3. IDE that opens .ipnyb files eg: Jupyter
#
#
# **Supervised Machine Learning:**
# It is a type of Machine Learning where a target variable is predicted from input variables by estimating the underlying function f(x) through various techniques.
# Here, we work with various supervised machine learning algorithm to predict the price of a house in boston.
# **Dataset:** The dataset is taken from scikit-learn library as it already comes as a builtin. So we manipulate the data by cleaning and preparing for our prediction.
# **Data Science Pipeline:** A typical data science process or pipelines has a set of iterative operations involved and we flip back and forth in the pipeline, as a result we achieve a model serving the business purpose to it's best. Below is a popular data science pipeline called CRISP-DM.
# <img src=https://cdn-images-1.medium.com/max/800/0*yt4EAnw2EfixSEJW.png>
# <center>CRISP-DM Process (Image credit: KDnuggets)
# So, we will start my importing necessary libraries.
import numpy as np
import pandas as pd
from sklearn.datasets import load_boston
import seaborn as sns
import matplotlib.pyplot as plt
# The dataset is available as load_boston() in scikit-learn datasets. When viewed, it is a dictionary with data, target variable, column names are present. We would clean it to create a tidy pandas dataframe.
# How it is stored in the library
load_boston()
# assigning it to 'boston'
boston = load_boston()
# What we do is, create a DataFrame with data as the data array stored in the above dictionary, column names as feature_names and a new column called target or price. So let me explain what each column actually mean in the dataset.
boston_df = pd.DataFrame(data = boston.data, columns=boston.feature_names)
boston_df['target'] = boston.target
boston_df.head()
# **Dataset information:**
#
# Number of instances: 506
#
# Number of attributes or columns: 14
#
# Attribute information:
#
# 1. CRIM per capita crime rate by town
#
# 2. ZN proportion of residential land zoned for lots over 25,000 sq.ft.
#
# 3. INDUS proportion of non-retail business acres per town
#
# 4. CHAS Charles River dummy variable (= 1 if tract bounds river; 0 otherwise)
#
# 5. NOX nitric oxides concentration (parts per 10 million)
#
# 6. RM average number of rooms per dwelling
#
# 7. AGE proportion of owner-occupied units built prior to 1940
#
# 8. DIS weighted distances to five Boston employment centres
#
# 9. RAD index of accessibility to radial highways
#
# 10. TAX full-value property-tax rate per $10,000
#
# 11. PTRATIO pupil-teacher ratio by town
#
# 12. B 1000(Bk - 0.63)^2 where Bk is the proportion of blacks by town
#
# 13. LSTAT % lower status of the population
#
# 14. MEDV Median value of owner-occupied homes in $1000’s
boston_df
# +
f, ax = plt.subplots(figsize = (15,20))
sns.heatmap(boston_df.corr(),
annot=True,
square=True,
cmap='coolwarm',
cbar_kws={'shrink': 0.3,
'ticks': [-1, -0.5, 0, 0.5, 1]},
linewidths=0.5,
vmin=-1,
vmax=1,
annot_kws={'size': 10})
ax.set_yticklabels(boston_df.corr().columns, rotation = 0)
ax.set_xticklabels(boston_df.corr().columns)
sns.set_style({'xtick.bottom': True}, {'ytick.left': True})
# -
| .ipynb_checkpoints/boston-house-prices-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stop Detection
#
# <img align="right" src="https://anitagraser.github.io/movingpandas/pics/movingpandas.png">
#
# There are no definitive answers when it comes to detecting / extracting stops from movement trajectories. Due to tracking inaccuracies, movement speed rarely goes to true zero. GPS tracks, for example, tend to keep moving around the object's stop location.
#
# Suitable stop definitions are also highly application dependent. For example, an application may be interested in analyzing trip purposes. To do so, analysts would be interested in stops that are longer than, for example, 5 minutes and may try to infer the purpose of the stop from the stop location and time. Shorter stops, such as delays at traffic lights, however would not be relevant for this appication.
#
# In the MovingPandas **TrajectoryStopDetector** implementation, a stop is detected if the movement stays within an area of specified size for at least the specified duration.
# ## Setup
# %matplotlib inline
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
# +
import pandas as pd
import geopandas as gpd
from datetime import datetime, timedelta
import sys
sys.path.append("..")
import movingpandas as mpd
print(mpd.__version__)
import warnings
warnings.simplefilter("ignore")
# -
# ## Loading Geolife Sample
# %%time
df = gpd.read_file('data/demodata_geolife.gpkg')
df['t'] = pd.to_datetime(df['t'])
df = df.set_index('t').tz_localize(None)
print("Finished reading {} rows".format(len(df)))
traj_collection = mpd.TrajectoryCollection(df, 'trajectory_id')
print(traj_collection)
# ## Stop Detection with a SingleTrajectory
# %%time
my_traj = traj_collection.trajectories[0]
stops = mpd.TrajectoryStopDetector(my_traj).get_stop_segments(min_duration=timedelta(seconds=60), max_diameter=100)
len(stops)
for x in mpd.TrajectoryStopDetector(my_traj).get_stop_time_ranges(min_duration=timedelta(seconds=60), max_diameter=100): print(x)
( my_traj.hvplot(title='Stops in Trajectory {}'.format(my_traj.id), line_width=7.0, tiles='CartoLight', color='slategray', width=700, height=700) *
stops.hvplot( size=200, line_width=7.0, tiles=None, color='deeppink') *
stops.get_start_locations().hvplot(geo=True, size=200, color='deeppink') )
# %%time
split = mpd.StopSplitter(my_traj).split(min_duration=timedelta(seconds=60), max_diameter=100)
( my_traj.hvplot(title='Stops in Trajectory {}'.format(my_traj.id), line_width=7.0, tiles='CartoLight', color='slategray', width=400, height=700) *
stops.get_start_locations().hvplot(geo=True, size=200, color='deeppink') +
split.hvplot(title='Trajectory {} split at stops'.format(my_traj.id), line_width=7.0, tiles='CartoLight', width=400, height=700)
)
for segment in split:
print(segment)
# ## Stop Detection for TrajectoryCollections
#
# The process is the same as for individual trajectories.
# %%time
stops = mpd.TrajectoryStopDetector(traj_collection).get_stop_segments(min_duration=timedelta(seconds=60), max_diameter=100)
len(stops)
( traj_collection.hvplot(width=700, height=700, line_width=7.0, tiles='CartoLight', color='slategray') *
stops.hvplot( size=200, line_width=7.0, tiles=None, color='deeppink') *
stops.get_start_locations().hvplot(geo=True, size=200, color='deeppink') )
# ## Continue exploring MovingPandas
#
# 1. [Getting started](1-getting-started.ipynb)
# 1. [Trajectory aggregation (flow maps)](2-generalization-and-aggregation.ipynb)
# 1. [Stop detection](3-stop-detection.ipynb)
| tutorials/3-stop-detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="79715e0d-3fc3-4e90-b515-342a9679c654" _uuid="4c87f41b97eaf6f8d864b9cebe4a2ede1cd281b0"
#Auxílio do Tutorial: https://matheusfacure.github.io/2017/05/12/tensorflow-essencial/
import tensorflow as tf
import gzip
import pickle
import numpy as np
import pandas as pd
from sklearn import preprocessing
from sklearn.preprocessing import MinMaxScaler
from sklearn.model_selection import train_test_split
import os # para criar pastas
from sklearn.metrics import r2_score, accuracy_score
# + _cell_guid="8fd468d4-60fd-45d1-b3e5-6004952f9c0a" _uuid="4cba27eb5ec04f67677b6de8c5a560dfe5536670"
train = pd.read_csv('../input/train.csv')
test = pd.read_csv('../input/test.csv')
train.head()
# + _cell_guid="144bf270-f5a0-4812-aa0c-1252ba5a9f7f" _uuid="b730f88ea29aeff5b5fbaafe6711c0b8209ac1d5"
def process_age(df,cut_points,label_names):
df["Age"] = df["Age"].fillna(-0.5)
df["Age_categories"] = pd.cut(df["Age"],cut_points,labels=label_names)
return df
cut_points = [-1,0,5,12,18,35,60,100]
label_names = ["Missing","Infant","Child","Teenager","Young Adult","Adult","Senior"]
train = process_age(train,cut_points,label_names)
test = process_age(test,cut_points,label_names)
def create_dummies(df,column_name):
dummies = pd.get_dummies(df[column_name],prefix=column_name)
df = pd.concat([df,dummies],axis=1)
return df
train = create_dummies(train,"Pclass")
test = create_dummies(test,"Pclass")
train = create_dummies(train,"Sex")
test = create_dummies(test,"Sex")
train = create_dummies(train,"Age_categories")
test = create_dummies(test,"Age_categories")
columns = ['Pclass_1', 'Pclass_2', 'Pclass_3', 'Sex_female', 'Sex_male',
'Age_categories_Missing','Age_categories_Infant',
'Age_categories_Child', 'Age_categories_Teenager',
'Age_categories_Young Adult', 'Age_categories_Adult',
'Age_categories_Senior','Survived','SibSp','Parch','Fare']
columns_test = ['Pclass_1', 'Pclass_2', 'Pclass_3', 'Sex_female', 'Sex_male',
'Age_categories_Missing','Age_categories_Infant',
'Age_categories_Child', 'Age_categories_Teenager',
'Age_categories_Young Adult', 'Age_categories_Adult',
'Age_categories_Senior','SibSp','Parch','Fare']
df = train[columns]
df.head()
# + _cell_guid="14a5b84c-0d0d-4c2d-8a11-d53ce1cb3a51" _uuid="ca5470f5d6212edca62572b312897bf6e05dbc47"
le = preprocessing.LabelEncoder()
df_encoded = df.apply(le.fit_transform)
#list(le.classes_)
#list(le.inverse_transform([2, 2, 1]))
df_encoded.astype(float)
scaler = MinMaxScaler()
df_encoded[df_encoded.columns] = scaler.fit_transform(df_encoded[df_encoded.columns])
df_encoded.head()
X = df_encoded.drop(['Survived'], axis=1)
y = df_encoded['Survived']
X.head()
# + _cell_guid="c460bef9-8bc3-43f1-b379-0c6f0e10772c" _uuid="e7fb9e81b68ae15858a51a3454baa715b44b4f39"
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
print('Formato dos dados:', X_train.shape, y_train.shape)
# + _cell_guid="713f5b25-fd3d-43cf-95d4-3e454f88d0d4" _uuid="3ba133c60da91729b84ef52f58b5676689f8527c"
# definindo constantes
lr = 1e-2 # taxa de aprendizado
n_iter = 2501 # número de iterações de treino
n_inputs = X_train.shape[1] # número de variáveis independentes
n_outputs = 1 # número de variáveis dependentes
graph = tf.Graph() # isso cria um grafo
with graph.as_default(): # isso abre o grafo para que possamos colocar operações e variáveis dentro dele.
tf.set_random_seed(1)
# adiciona as variáveis ao grafo
W = tf.Variable(tf.truncated_normal([n_inputs, n_outputs], stddev=.1), name='Weight')
b = tf.Variable(tf.zeros([n_outputs]), name='bias')
######################################
# Monta o modelo de regressão linear #
######################################
# Camadas de Inputs
x_input = tf.placeholder(tf.float32, [None, n_inputs], name='X_input')
y_input = tf.placeholder(tf.float32, [None, n_outputs], name='y_input')
# Camada Linear
y_pred = tf.add(tf.matmul(x_input, W), b, name='y_pred')
# Camada de custo ou função objetivo
EQM = tf.reduce_mean(tf.square(y_pred - y_input), name="EQM")
# otimizador
optimizer = tf.train.AdamOptimizer(learning_rate=lr).minimize(EQM)
# inicializador
init = tf.global_variables_initializer()
# para salvar o modelo treinado
saver = tf.train.Saver()
# + _cell_guid="698eb24e-3488-42a9-88b3-87931070387f" _uuid="3ac0e1a557aff0ce440b14cc246dac2fec0677a9"
# criamos uma pasta para salvar o modelo
if not os.path.exists('tmp'):
os.makedirs('tmp')
# abrimos a sessão tf
with tf.Session(graph=graph) as sess:
sess.run(init) # iniciamos as variáveis
# cria um feed_dict
feed_dict = {x_input: X_train, y_input: y_train.values.reshape(-1,1)}
# realizamos as iterações de treino
for step in range(n_iter + 1):
# executa algumas operações do grafo
_, l = sess.run([optimizer, EQM], feed_dict=feed_dict)
if (step % 500) == 0:
print('Custo na iteração %d: %.2f \r' % (step, l), end='')
saver.save(sess, "./tmp/my_model.ckpt")
# + _cell_guid="a8f83cfa-f158-40ef-ac24-d7d1fe45686d" _uuid="84a33012563a3a1c0fcdcc3ccbd84971d36ac83c"
# novamente, abrimos a sessão tf
with tf.Session(graph=graph) as sess:
# restauramos o valor das variáveis
saver.restore(sess, "./tmp/my_model.ckpt", )
# rodamos o nó de previsão no grafo
y_hat = sess.run(y_pred, feed_dict={x_input: X_test})
print('\nR2: %.3f' % r2_score(y_pred=y_hat, y_true=y_test))
print('\nAccuracy %.3f' % accuracy_score(y_test, y_hat.round(), normalize=True))
| v2/Titanic/titanic-linear-regression-tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# ----------
# User Instructions:
#
# Create a function compute_value which returns
# a grid of values. The value of a cell is the minimum
# number of moves required to get from the cell to the goal.
#
# If a cell is a wall or it is impossible to reach the goal from a cell,
# assign that cell a value of 99.
# ----------
import copy
grid = [[0, 1, 0, 0, 0, 0],
[0, 1, 0, 0, 0, 0],
[0, 1, 1, 1, 1, 0],
[0, 1, 0, 0, 1, 0],
[0, 0, 0, 0, 1, 0]]
goal = [len(grid)-1, len(grid[0])-1]
cost = 1 # the cost associated with moving from a cell to an adjacent one
delta = [[-1, 0 ], # go up
[ 0, -1], # go left
[ 1, 0 ], # go down
[ 0, 1 ]] # go right
delta_name = ['^', '<', 'v', '>']
def compute_value(grid,goal,cost):
# ----------------------------------------
# insert code below
# ----------------------------------------
value_map = copy.deepcopy(grid) #
for row in range(len(value_map)): # 所有元素乘-1
for ii in range(len(value_map[0])):
value_map[row][ii] = grid[row][ii]*99
close_set = [goal]
stage_cost = 0
hist = []
hist.append(goal)
#display(hist)
while len(close_set) != 0: #如果考察集不是空集
next_set = []
stage_cost = stage_cost+ cost
#display(stage_cost)
for i in range(len(close_set)):#取任一个当前考察集里的元素
x = close_set[i][0] # 第几行
y = close_set[i][1] # 第几列
for ii in range(len(delta)):
#display(ii)
x2 = x + delta[ii][0] #行数变化
y2 = y + delta[ii][1] #列数变化
if x2 >= 0 and x2 < len(grid) and y2 >=0 and y2 < len(grid[0]): # 在范围内
#display('in range')
#display(hist)
#display([[x2,y2]])
#display([[x2,y2]] not in hist )
#display(grid[x2][y2] == 0)
if ([x2,y2] not in hist) and grid[x2][y2] == 0: #没有被考察过,而且可行 [X2,Y2] instead of [[x2,y2]]
#display('find a next step')
value_map[x2][y2] = stage_cost
next_set.append([x2,y2])
hist.append([x2,y2]) #加入已查找
close_set = []
close_set = copy.deepcopy(next_set) #更新当前考察集
#display(close_set)
#display(hist)
#display(value_map)
#Final Process for the un-reach area to set to 99
for row in range(len(value_map)):
for col in range(len(value_map[0])):
if value_map[row][col] == 0:
value_map[row][col] = 99
value_map[goal[0]][goal[1]] =0
# make sure your function returns a grid of values as
# demonstrated in the previous video.
return value_map
# -
compute_value(grid,goal,cost)
| Path_Generation/Search_exercise/.ipynb_checkpoints/DynamicPrograming-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Berkeley 311 calls - Lecture notes
# ## Modules
#
# A **module** is a way for Python to store functions and variables so they can be reused. You `import` modules to use those functions and variables. Typically, all the `import` statements are the top of a file or notebook.
import pandas as pd
import altair as alt
import requests
# The `requests` module is already built into Python. We installed the pandas and altair modules when we installed `requirements-3.8.5.txt` earlier.
#
# A **package** is a kind of module that uses "dotted module names." That just means you can use methods with the name of the package followed by a period and the method, e.g. `requests.get()`.
#
# When you `import` a module, you can give it an alias using `as`. Above, we gave the pandas module the alias of `pd`. That means we can call the library’s methods using `pd.read_csv()` instead of `pandas.read_csv()`. We always `import pandas as pd` because that's what the pandas-using community has decided as a convention; likewise, we always `import altair as alt`.
#
# Sometimes we call modules like pandas and altair "libraries." That's a general software term; there's no special Python thing called "library."
# ## What is pandas?
#
# It's a Python Data Analysis library.
#
# [Pandas](https://pandas.pydata.org/) is a library that allows you to view table data and perform lots of different kinds of operations on that table. In pandas, a table is called a **dataframe**. If you’ve used Excel or Google Sheets, a dataframe should look familiar to you. There are rows and columns. You have column headers. You have discrete rows.
#
# ## What is Altair?
#
# [Altair](https://altair-viz.github.io/) is a data visualization library for Python. `matplotlib` is usually the first data viz module Python programmers learn, but Altair is easier to use.
# ## Download data
#
# We're going to download [311 call data](https://data.cityofberkeley.info/311/311-Cases-COB/bscu-qpbu) from the City of Berkeley's Open Data Portal. You can run the cell below. It will download the data into a file called `berkeley_311.csv`.
#
# If you want to download something straight from the Internet again, you can copy this code but swap out the url and the file name. (But don't forget to `import requests` at the top of your notebook.)
# +
url = 'https://data.cityofberkeley.info/api/views/bscu-qpbu/rows.csv?accessType=DOWNLOAD'
r = requests.get(url, allow_redirects=False)
# write the content of the request into a file called `berkeley_311.csv`
open('berkeley_311.csv', 'wb').write(r.content)
# -
# ## Import the csv into a `pandas` dataframe
#
# We use a method called `pd.read_csv()` to import a csv file into a dataframe.
#
# Make sure you assign the dataframe into a variable. Below, we're calling the dataframe `berkeley_311_original`.
berkeley_311_original = pd.read_csv('berkeley_311.csv')
# In these notes, I'm going to use the term `df` to stand for 'dataframe' — you will see `df` when you're searching the Internet for answers. In your actual work, you'll replace `df` with whatever you called your dataframe; in this case, that's `berkeley_311_original`.
# ## View data
#
# ### `df.head()` and `df.tail()`
#
# Use `df.head()` to view the first 5 rows and the last 5 rows of the dataframe.
berkeley_311_original.head()
# You'll notice that there are numbers `0`, `1`, `2`, `3`, and `4` added to the dataframe to the left. That's called the **index** of the dataframe. An index is basically a row id.
#
# You can also use `df.head(20)` to view the first 20 rows.
# How would you view the last 5 rows? Use `df.tail()`. This might look similar to the command-line lecture we had a few weeks ago.
berkeley_311_original.tail()
# If you call the dataframe on its own, you'll get both the first 5 rows and the last 5 rows.
berkeley_311_original
# ### `df.info()`
#
# Use `df.info()` to get more information on the dataframe. In particular, this method is useful in that it shows us the column names and what `dtype` the column is. (I'll explain `dtype` below.)
berkeley_311_original.info()
# ### What is dtype?
#
# You remember when we talked about Python **types**, like `int`, `float`, and `string`? Above, we have `int64` instead of `int`, and `float64` instead of `float`. (`object` is pretty close to `string`, but not exactly.)
#
# Here, **dtype** stands for **data type** and comes from a module called `numpy`. Even though we did not `import numpy`, the pandas module imported numpy within its own code.
#
# Side note on dtypes: Sometimes, data doesn't import correctly, and you have to `df.read_csv()` again while simultaneously specifying the dtype. We're not going to do that today because it looks like most of the data imported OK. But we will convert the dtype of two columns so we can perform certain calculations.
#
# Let's take a look at a bit of the dataframe again.
berkeley_311_original.head()
# There are 2 columns that we want to convert. **Date_Opened** and **Date_Closed** are both `object` dtype, but we want to change them to a `datetime64` dtype. That allows us to do some math operations, like sort by the earliest date in the dataframe.
berkeley_311_original['Date_Opened'].min()
berkeley_311_original['Date_Opened'].max()
# The operations above are wrong — they are sorted by string, not by actual date! We saw 2021 data when we called the df.tail() method earlier.
# ## Properly type the data
#
# ### Copy the original dataframe
#
# Before we start convert the 2 columns, let's copy the original dataframe into a new dataframe. Below, I'm going to use `df.copy()` to create a copy of the original dataframe. We're not going to alter the original dataframe at all. That way, if we run into any problems later, we can compare our edited dataframe with the original dataframe.
berkeley_311 = berkeley_311_original.copy()
# ### Convert columns to `datetime`
#
# Let's convert **Date_Opened** first so we can contrast the two columns. The syntax for this conversion is:
#
# ```python
# df['column_name'] = pd.to_datetime(df['column_name'])
# ```
berkeley_311['Date_Opened'] = pd.to_datetime(berkeley_311['Date_Opened'])
# By the way, `berkeley_311['Date_Opened']` is a pandas **series**. We don't have to worry too much about that right now, but I want you to have to the right terminology. We're converting a series to a version of itself that passed through the `pd.to_datetime()` method.
# Look at the dataframe now and compare **Date_Opened** and **Date_Closed**.
berkeley_311.head()
# See how they look different? You can also call `df.info()` again.
berkeley_311.info()
# Let's now convert the 2nd column `Date_Closed`:
berkeley_311['Date_Closed'] = pd.to_datetime(berkeley_311['Date_Closed'])
# And let's take a peek at the change:
berkeley_311.head()
# By the way, you might notice that we call `df.head()` and `df.tail()` a lot to check our work. That's OK. Later on, you will be able to run more code at once before calling one of those methods, but for now, it's good to check your work often.
# ### .min() and .max()
#
# Now we can find the earliest date and the latest date of both columns, by performing `series.min()` and `series.max()` on these series. (There's no equivalent of `df`/dataframe for **series**, unfortunately.)
berkeley_311['Date_Opened'].min()
berkeley_311['Date_Opened'].max()
# The min and max tell us that the year 2010 is not complete (and for that matter, neither is the current year, although we wouldn't need pandas to tell us that). If we analyze the data by year later, we might want to exclude 2010 and 2021 data.
# ### Get the difference of the two date columns
#
# Pandas allows you to get the difference of two dates by literally subtracting one datetime column from another. We'll create a new column called **Close_Time** that shows us how long it took for a case to be closed.
berkeley_311['Close_Time'] = berkeley_311['Date_Closed'] - berkeley_311['Date_Opened']
# The resulting column will not be a `datetime` dtype. It will be a `timedelta` dtype. The term "delta" is often used to mean "change." Observe the last column:
berkeley_311.head()
# ## A brief detour into data analysis
#
# Now that we've converted the columns, we can do some interesting operations on them.
#
# ### Mean
#
# Get the mean of a column by calling `series.mean()`.
berkeley_311['Close_Time'].mean()
# The average time the city took to close a case was around 61 days. This is for the whole dataset, from early 2010 to now.
# ### Median
berkeley_311['Close_Time'].median()
# But the median time was around 5 days.
# + [markdown] tags=[]
# ### Min
# -
berkeley_311['Close_Time'].min()
# The shortest amount of time was 0 days and 0 seconds. That might be a public comment that didn't require follow-up. We can check on that later.
# ### Max
berkeley_311['Close_Time'].max()
# One case seems to have taken 3,373 days! That's almost 10 years. That seems like way too long. There might be an error here.
#
# Let's take a quick detour into _subsetting_ the data. That means to take a smaller set of the data based on some conditions. I'll explain how to subset more later, but for now, check out the following code:
berkeley_311[berkeley_311['Close_Time'] >= '3373 days']
# It's hard to know why it took so long to close this case without asking the city for more information.
# ### Sort data
#
# We can even look at the top 10 cases that took the longest time to resolve. You'll use the `df.sort_values()` method.
#
# Let's break down the below code before we run it. You can see there are 2 options within the parentheses for `.sort_values()`:
# ```python
# by=['Close_Time'], ascending=False
# ```
#
# The `by` argument tells us which column we will sort the dataframe by. You always need to include this argument. You can sort by multiple columns, too.
#
# The optional `ascending` argument tells us if we want the dataframe to sort from smallest to largest or earliest to latest. By default, `ascending` is set to `True`, so we're going to change it here so it's `False`.
#
# Next, I don't want to see the entire dataframe, just the first 10 rows. So I'm going to call `df.head(10)`.
berkeley_311.sort_values(by=['Close_Time'], ascending=False).head(10)
# So that's a preview of an interesting analysis we can do. I showed you one fun part before we moved onto the harder part, checking and vetting the data. Usually, we need to do that first. But we did convert the columns to datetime, which is part of making sure the data was valid.
# ## Clean data, part 2: Check and vet the data
#
# ### Unique identifier for every row?
#
# First, I want to see if `Case_ID` has a unique ID for every row. Why? When you're doing a data analysis, every row should have its own unique ID. Hopefully, the agency that gave you the data has provided a unique ID. Sometimes, though, they don't. In those cases, you want to create a unique ID for every row.
#
# I get a count of unique values by calling `.nunique()` on a column.
berkeley_311['Case_ID'].nunique()
# How many rows do we have again? We can use `df.info()` to get the number of rows, or we can scroll up to see again. I'm feeling lazy, so let's just call `len(df)`. (Do you remember that we learned `len()` for both strings and data structures earlier?)
len(berkeley_311)
# So there are 588,022 unique case IDs, but 588,028 rows in this dataframe. There might be duplicates or there could be missing data. Let's check for both.
#
# We're going to check by **subsetting** the data.
# ### Subsetting
#
# This is the general structure of how you subset data in pandas.
#
# ```python
# df[ expression ]
# ```
#
# That's not very descriptive. What's the _expression_? There are lots of different ways we write these expressions in pandas. I'm going to show you a handful of different kinds today, but know there are a bunch more!
#
# Ultimately, what I want is a list of the duplicate `Case_ID`s. I'll then subset the dataframe to show any row that has a Case_ID that is on that list.
#
#
# We'll first check to see which rows in `berkeley_311` are exact duplicates.
berkeley_311[berkeley_311.duplicated()]
# There are no exact duplicates. Now let's check specifically to see rows in which `Case_ID`s are duplicated.
berkeley_311[berkeley_311['Case_ID'].duplicated()]
# What's annoying about this is that it only shows ONE instance of the Case_ID. What I want is a list of those Case IDs. How do I make a list of the values in one column?
#
# First, I'm going to create a new dataframe that has the duplicated IDs. We're going to create a copy.
dupe_cases = berkeley_311[berkeley_311['Case_ID'].duplicated()].copy()
dupe_cases
# Now we create a `list` or `array` (the numpy version of a list) of those IDs:
# +
dupe_case_ids = dupe_cases['Case_ID'].to_list()
# to create an array, you can use this instead:
# dupe_case_ids = dupe_cases['Case_ID'].unique()
# -
# Now we'll call `dupe_case_ids`, so we can see what's in it.
dupe_case_ids
# Now we'll subset the data by finding cases in our edited dataframe `berkeley_311`.
berkeley_311[berkeley_311['Case_ID'].isin(dupe_case_ids)]
# OK, this is weird, it looks like all the cases are for the same address. Still, it's hard to tell what's going on, so I'm going to sort that dataframe by **Case_ID**.
berkeley_311[berkeley_311['Case_ID'].isin(dupe_case_ids)].sort_values(by=['Case_ID'])
# The thing that looks different is the geocoding between those cases.
#
# FYI, all the code we used above didn't change the original dataframe. We were subsetting, but we did not subset with a new variable.
#
# Now, we're going to drop the duplicated cases, and reset the variable `berkeley_311`.
berkeley_311 = berkeley_311.drop_duplicates(subset=['Case_ID'])
berkeley_311
# By the way, you might have noticed there's something called `NaT` in one of the rows above. `NaT` stands for _not a time_ and is kind of like `None` or an empty cell in Google Sheets. For non-time related blank cells, you'll see `NaN` (not a number) instead of `NaT`.
#
# The difference between `None` and `NaN`/`NaT` is that the latter allows you to perform calcuations and skip any blank cells. That means, you probably need to check for how many `NaN`/`NaT` cells exist in your dataframe. If there are a lot of them, your analysis might not be valid. You can quickly check for that with the `df.info()` method we learned earlier. There's a column called `Non-Null Count`.
berkeley_311.info()
# Which columns have a lot of null values?
# ### Assert
# The keyword `assert` is a good way for us to check if the length of the dataframe now matches the number of unique IDs.
assert len(berkeley_311) == berkeley_311['Case_ID'].nunique()
# If the assertion is `True`, nothing happens. But if the assertion is `False`, you'll get an error. You might want to use these kinds of assertions when you have to re-run your notebooks or have to import updated datasets.
# ## Export a clean version of the data to a csv
#
# We can use the `df.to_csv()` method to export a clean copy of the csv. That way, you can instantly import the clean data in a new notebook instead of rerunning the code in this notebook.
#
# Before we run the code below, let's take a closer look:
#
# ```python
# berkeley_311.to_csv('berkeley_311_clean.csv', index=False)
# ```
#
# The first argument in `df.to_csv()` is the name of the file we're going to export our dataframe into. In this case, that's `berkeley_311_clean.csv`.
#
# The second argument is `index=False`. This means that I don't want pandas to export those row ids (0, 1, 2, 3, etc.) that show up at the very lefthand side of the dataframe.
berkeley_311.to_csv('berkeley_311_clean.csv', index=False)
# Try removing `index=False` and see what happens.
| berkeley_311_calls_lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 02: Working with numbers
# **1**. (10 points)
#
# Normalize the $3 \times 4$ diagonal matrix with diagonal (1, ,2, 3) so all rows have mean 0 and standard deviation 1. The matrix has 0 everywhere not on the diagonal.
# **2**. (10 points)
#
# A fixed point of a funciton is a value that remains the same when the funciton is applied to it, that is $f(x) = x$. Write a function that finds the fixed poitn of another function $f$ given an intiial value $x_0$. For example, if
#
# $$f(x) \rightarrow \sqrt{x}$$
#
# and
#
# $x_0$ is any positive real number, then the function should return 1 since
#
# $$\sqrt{1} = 1$$
#
# Not all funcitons have a fixed point - if it taakes over 1,000 iterations, the fucntion shold return None.
#
# - Use the function signature `fixed_point(f, x0, max_iter=1000)`.
# - Test with `fixed_point(np.sqrt, 10)`.
# **3**. (10 points)
#
# Use `np.fromfunction` to construc the following matrix
#
# ```python
# array([[5, 0, 0, 0, 5],
# [0, 4, 0, 4, 0],
# [0, 0, 3, 0, 0],
# [0, 2, 0, 2, 0],
# [1, 0, 0, 0, 1]])
# ```
# **4**. (15 points)
#
# Simulate $n$ coin toss experiments, in which you toss a coin $k$ times for each experiment. Find the maximum run length of heads (e.g. the sequence `T,T,H,H,H,T,H,H` has a maximum run length of 3 heads in each experiment. What is the most common maximum run length?
#
# Let $n$ = 10,000 and $k=100$.
# **5**. (15 points)
#
# Wikipedia gives this algorithm for finding prime numbers
#
# To find all the prime numbers less than or equal to a given integer n by Eratosthenes' method:
#
# - Create a list of consecutive integers from 2 through n: (2, 3, 4, ..., n).
# - Initially, let p equal 2, the smallest prime number.
# - Enumerate the multiples of p by counting to n from 2p in increments of p, and mark them in the list (these will be 2p, 3p, 4p, ...; the p itself should not be marked).
# - Find the first number greater than p in the list that is not marked. If there was no such number, stop. Otherwise, let p now equal this new number (which is the next prime), and repeat from step 3.
# - When the algorithm terminates, the numbers remaining not marked in the list are all the primes below n.
#
# Find all primes less than 1,000 using this method.
#
# - You may use `numpy` and do not have to follow the algorithm exactly if you can achieve the same results.
# **6**. (40 points)
#
# Write code to generate a plot similar to those shown below using the explanation for generation of 1D Cellular Automata found [here](http://mathworld.wolfram.com/ElementaryCellularAutomaton.html). You should only need to use standard Python, `numpy` and `matplotllib`.
#
# 
#
#
#
# The input to the function making the plots should be a simple list of rules
#
# ```python
# rules = [30, 54, 60, 62, 90, 94, 102, 110, 122, 126,
# 150, 158, 182, 188, 190, 220, 222, 250]
# make_plots(rules, niter, ncols)
# ```
#
# You may, of course, write other helper functions to keep your code modular.
| homework/HW02_numbers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="yTDqVBbh36V_" colab_type="text"
# # Assignment 2
# ## Sentiment Classification
#
# This assignment approaches the sentiment classification problem by using deep learning techniques and also exposes some ethical problem related to these methods.
#
# This is what we’re going to do:
#
# # + Acquire some pre-computed word embeddings to represent the meanings of words
# # + Acquire training and test data, with gold-standard examples of positive and negative words
# # + Train a simple classifier to recognize other positive and negative words based on their word embeddings
# # + Compute sentiment scores for sentences of text using this classifier
# # + Analyze the results to look for unwanted bias.
# + [markdown] id="litP4Bg836WB" colab_type="text"
# ## Word embeddings
#
# There are several datasets of pre-trained English word embeddings such as `word2vec`, pretrained on Google News data, and `GloVe`, pretrained on the Common Crawl of web pages. We will use `GloVe`.
#
# GloVe comes in three sizes: 6B, 42B, and 840B. The 42B version is pretty good and is also neatly trimmed to a vocabulary of 1 million words. We will just use the 42B version.
#
# > **GloVe.42B** data: 42B tokens, 1.9M vocab, uncased, 300d vectors, 1.75 GB download
# + id="SE7Br8Ui36WC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 351} outputId="ec52738b-6a65-4917-bd13-2b5ef1e217da"
# !wget http://nlp.stanford.edu/data/glove.42B.300d.zip
# + id="gIly4FM436WF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="a3f8df53-8a9d-494b-e98b-ed2a564e8732"
# !unzip glove.42B.300d.zip
# + id="xGy_qF5t36WH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="b3343b12-c072-43ee-ad44-a75076b85e88"
import numpy as np
import pandas as pd
from tqdm import tqdm
def load_embeddings(filename):
labels = []
rows = []
with open(filename, encoding='utf-8') as infile:
for i, line in tqdm(enumerate(infile)):
items = line.rstrip().split(' ')
if len(items) == 2:
# This is a header row giving the shape of the matrix
continue
labels.append(items[0])
values = np.array([float(x) for x in items[1:]], 'f')
rows.append(values)
arr = np.vstack(rows)
return pd.DataFrame(arr, index=labels, dtype='f')
embeddings = load_embeddings('glove.42B.300d.txt')
embeddings.shape
# + [markdown] id="mJ2HHZpc36WJ" colab_type="text"
# ## Positive and Negative Words
#
# We need some input about which words are positive and which words are negative. There are many sentiment lexicons you could use, but we’re going to go with a very straightforward lexicon from https://www.cs.uic.edu/~liub/FBS/sentiment-analysis.html
#
# There is a copy of these files in the GitHub repository of the course.
# + id="rWyxpDkX36WJ" colab_type="code" colab={}
def load_lexicon(filename):
lexicon = []
with open(filename, encoding='latin-1') as infile:
for line in infile:
line = line.rstrip()
if line and not line.startswith(';'):
lexicon.append(line)
return lexicon
pos_words = load_lexicon('positive-words.txt')
neg_words = load_lexicon('negative-words.txt')
# + [markdown] id="r71VdqOc36WL" colab_type="text"
# Some of these words are not in the GloVe vocabulary. Those words end up with rows full of NaN to indicate their missing embeddings, so we will use Pandas to clean the dataset.
# + id="zm6frm7X36WM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 251} outputId="81510a68-12d9-413a-e6c2-a872b36f2f51"
pos_vectors = embeddings.loc[pos_words].dropna()
neg_vectors = embeddings.loc[neg_words].dropna()
# + [markdown] id="ie0PrSrx36WO" colab_type="text"
# Now we make arrays of the desired inputs and outputs. The inputs are the embeddings, and the outputs are `1` for positive words and `-1` for negative words. We also make sure to keep track of the words they’re labeled with, so we can interpret the results.
# + id="m621qcuJ36WP" colab_type="code" colab={}
vectors = pd.concat([pos_vectors, neg_vectors])
targets = np.array([1 for entry in pos_vectors.index] + [-1 for entry in neg_vectors.index])
labels = list(pos_vectors.index) + list(neg_vectors.index)
# + id="Gr0h65tU36WQ" colab_type="code" colab={}
from sklearn.model_selection import train_test_split
train_vectors, test_vectors, train_targets, test_targets, train_labels, test_labels = \
train_test_split(vectors, targets, labels, test_size=0.1, random_state=0)
# + [markdown] id="WmvRa6EU36WS" colab_type="text"
# Now it is time to make your classifier, and train it by running the training vectors through it for 100 iterations. You can use a logistic function as the loss, so that the resulting classifier can output the probability that a word is positive or negative.
# + id="JYYhRoD9EvOV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="ebed526f-c346-40bc-f9d3-fb3be3aa9e4f"
train_vectors.shape, train_targets.shape
# + id="c6rKVM8yJKk4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="069cd8ba-a00e-4c3c-b489-95e2b501e07f"
for a, b in {'a': 1, 'b': 2}.items():
print(a,b)
# + id="qhUhDc73G1aN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 338} outputId="e66f45df-79cf-4f53-c5e2-f28759a58103"
# write your classifier here
from sklearn.tree import DecisionTreeClassifier
from sklearn.metrics import accuracy_score
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import LinearSVC
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import SGDClassifier
X = train_vectors; y = train_targets
for name, model in {
'DecisionTreeClassifier': DecisionTreeClassifier(),
'MLPClassifier': MLPClassifier(hidden_layer_sizes=(100,), max_iter=100, random_state=1),
'RandomForestClassifier': RandomForestClassifier(),
'KNeighborsClassifier': KNeighborsClassifier(),
'LinearSVC': LinearSVC(),
'GaussianNB': GaussianNB(),
'SGDClassifier': SGDClassifier(loss='log', max_iter=100)
}.items():
model.fit(X, y)
print(name + " accuracy on train: ", accuracy_score(y.flatten(), model.predict(X)))
print(name + " accuracy on train: ", accuracy_score(test_targets.flatten(), model.predict(test_vectors)))
# + id="039sm98qEUhG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 104} outputId="ae3b2e24-66c7-4c25-ace3-89519b9dfa88"
from sklearn.linear_model import LogisticRegression
model = LogisticRegression()
model.fit(X, y)
print("LogisticRegression accuracy on train: ", accuracy_score(y.flatten(), model.predict(X)))
print("LogisticRegression accuracy on train: ", accuracy_score(test_targets.flatten(), model.predict(test_vectors)))
# + [markdown] id="d7nkqCLT36WU" colab_type="text"
# Evaluate your classifier on the test vectors. Which is the classifier performace (in terms of accuracy)?
# + id="hC0jVNW836WV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="52316d99-1af4-4fd4-ada6-1011904c2ced"
# Compute your classifier accuracy
# It should predict the correct sentiment for sentiment words outside of its training data 95% of the time.
print("accuracy on train: ", accuracy_score(test_targets.flatten(), model.predict(test_vectors)))
# + [markdown] id="uK1i_QdU36WW" colab_type="text"
# Let’s define a function that we can use to see the sentiment that this classifier predicts for particular words, then use it to see some examples of its predictions on the test data.
# + id="RHjbLz4436WX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 761} outputId="16812b48-84bd-44cb-8c93-f64007e39c61"
def vecs_to_sentiment(vecs):
# predict_log_proba gives the log probability for each class
predictions = model.predict_log_proba(vecs)
# To see an overall positive vs. negative classification in one number,
# we take the log probability of positive sentiment minus the log
# probability of negative sentiment.
return predictions[:, 1] - predictions[:, 0]
def words_to_sentiment(words):
vecs = embeddings.loc[words].dropna()
log_odds = vecs_to_sentiment(vecs)
return pd.DataFrame({'sentiment': log_odds}, index=vecs.index)
# Show 20 examples from the test set
words_to_sentiment(test_labels).ix[:20]
# + [markdown] id="J4526kr336WY" colab_type="text"
# ## Sentiment score for text
#
# There are many ways to combine sentiments for word vectors into an overall sentiment score. The simplest way is to average them.
# + id="A3dEGQHR36WZ" colab_type="code" colab={}
import re
TOKEN_RE = re.compile(r"\w.*?\b")
# The regex above finds tokens that start with a word-like character (\w), and continues
# matching characters (.+?) until the next word break (\b). It's a relatively simple
# expression that manages to extract something very much like words from text.
def text_to_sentiment(text):
tokens = [token.casefold() for token in TOKEN_RE.findall(text)]
sentiments = words_to_sentiment(tokens)
return sentiments['sentiment'].mean()
# + id="hQTzNCdS36Wa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="1ab9c390-081c-48f1-d993-a15bef8cbd33"
text_to_sentiment("this example is pretty cool")
# + id="qTzmNPsS36Wc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 33} outputId="44862985-6fd6-49a1-ae5c-34820e6877f1"
text_to_sentiment("meh, this example sucks")
# + [markdown] id="yEOgMznp36We" colab_type="text"
# Let’s see what it does with a few variations on a neutral sentence:
# + id="JJGhHOQV36Wf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="1e5e3327-05fd-4ef7-afeb-2d4de140293b"
print(text_to_sentiment("Let's go get Italian food"))
print(text_to_sentiment("Let's go get Chinese food"))
print(text_to_sentiment("Let's go get Mexican food"))
# + id="0ireRAwn36Wh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 84} outputId="82a1966f-5c7a-46b4-935f-7c1308412634"
print(text_to_sentiment("My name is Emily"))
print(text_to_sentiment("My name is Heather"))
print(text_to_sentiment("My name is Yvette"))
print(text_to_sentiment("My name is Shaniqua"))
# + [markdown] id="wRGzWes336Wi" colab_type="text"
# The system has associated wildly different sentiments with people’s names. You can look at these examples and many others and see that the sentiment is generally more positive for stereotypically-white names, and more negative for stereotypically-black names.
# + [markdown] id="x_E9QiMm36Wj" colab_type="text"
# ## Ethical problem
#
# We want to learn how to not make something like this again. So let’s put more data through it, and statistically measure how bad its bias is.
#
# Here we have four lists of names that tend to reflect different ethnic backgrounds, mostly from a United States perspective. The first two are lists of predominantly “white” and “black” names adapted from Caliskan et al.’s article. I also added typically Hispanic names, as well as Muslim names that come from Arabic or Urdu; these are two more distinct groupings of given names that tend to represent your background.
# + id="b5_7Pgtl36Wj" colab_type="code" colab={}
NAMES_BY_ETHNICITY = {
# The first two lists are from the Caliskan et al. appendix describing the
# Word Embedding Association Test.
'White': [
'Adam', 'Chip', 'Harry', 'Josh', 'Roger', 'Alan', 'Frank', 'Ian', 'Justin',
'Ryan', 'Andrew', 'Fred', 'Jack', 'Matthew', 'Stephen', 'Brad', 'Greg', 'Jed',
'Paul', 'Todd', 'Brandon', 'Hank', 'Jonathan', 'Peter', 'Wilbur', 'Amanda',
'Courtney', 'Heather', 'Melanie', 'Sara', 'Amber', 'Crystal', 'Katie',
'Meredith', 'Shannon', 'Betsy', 'Donna', 'Kristin', 'Nancy', 'Stephanie',
'Bobbie-Sue', 'Ellen', 'Lauren', 'Peggy', 'Sue-Ellen', 'Colleen', 'Emily',
'Megan', 'Rachel', 'Wendy'
],
'Black': [
'Alonzo', 'Jamel', 'Lerone', 'Percell', 'Theo', 'Alphonse', 'Jerome',
'Leroy', 'Rasaan', 'Torrance', 'Darnell', 'Lamar', 'Lionel', 'Rashaun',
'Tyree', 'Deion', 'Lamont', 'Malik', 'Terrence', 'Tyrone', 'Everol',
'Lavon', 'Marcellus', 'Terryl', 'Wardell', 'Aiesha', 'Lashelle', 'Nichelle',
'Shereen', 'Temeka', 'Ebony', 'Latisha', 'Shaniqua', 'Tameisha', 'Teretha',
'Jasmine', 'Latonya', 'Shanise', 'Tanisha', 'Tia', 'Lakisha', 'Latoya',
'Sharise', 'Tashika', 'Yolanda', 'Lashandra', 'Malika', 'Shavonn',
'Tawanda', 'Yvette'
],
# This list comes from statistics about common Hispanic-origin names in the US.
'Hispanic': [
'Juan', 'José', 'Miguel', 'Luís', 'Jorge', 'Santiago', 'Matías', 'Sebastián',
'Mateo', 'Nicolás', 'Alejandro', 'Samuel', 'Diego', 'Daniel', 'Tomás',
'Juana', 'Ana', 'Luisa', 'María', 'Elena', 'Sofía', 'Isabella', 'Valentina',
'Camila', 'Valeria', 'Ximena', 'Luciana', 'Mariana', 'Victoria', 'Martina'
],
# The following list conflates religion and ethnicity, I'm aware. So do given names.
#
# This list was cobbled together from searching baby-name sites for common Muslim names,
# as spelled in English. I did not ultimately distinguish whether the origin of the name
# is Arabic or Urdu or another language.
#
# I'd be happy to replace it with something more authoritative, given a source.
'Arab/Muslim': [
'Mohammed', 'Omar', 'Ahmed', 'Ali', 'Youssef', 'Abdullah', 'Yasin', 'Hamza',
'Ayaan', 'Syed', 'Rishaan', 'Samar', 'Ahmad', 'Zikri', 'Rayyan', 'Mariam',
'Jana', 'Malak', 'Salma', 'Nour', 'Lian', 'Fatima', 'Ayesha', 'Zahra', 'Sana',
'Zara', 'Alya', 'Shaista', 'Zoya', 'Yasmin'
]
}
# + [markdown] id="bW8d-UTG36Wl" colab_type="text"
# Now we’ll use Pandas to make a table of these names, their predominant ethnic background, and the sentiment score we get for them:
# + id="7LrHlgku36Wl" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 741} outputId="135e2efd-1899-4a0c-b3c5-0d2ff0ce3248"
def name_sentiment_table():
frames = []
for group, name_list in sorted(NAMES_BY_ETHNICITY.items()):
lower_names = [name.lower() for name in name_list]
sentiments = words_to_sentiment(lower_names)
sentiments['group'] = group
frames.append(sentiments)
# Put together the data we got from each ethnic group into one big table
return pd.concat(frames)
name_sentiments = name_sentiment_table()
name_sentiments.ix[::25]
# + id="qV-_PiWH36Wp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 300} outputId="acec494f-7d07-4cf9-c06f-ae44a28f8182"
import seaborn
plot = seaborn.swarmplot(x='group', y='sentiment', data=name_sentiments)
plot.set_ylim([-10, 10])
# + id="AEMEogIQ36Wr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 280} outputId="d70bdb53-2e5d-491d-b4a6-c740fdeab63f"
plot = seaborn.barplot(x='group', y='sentiment', data=name_sentiments, capsize=.1)
# + id="o6QOaT7kCIx_" colab_type="code" colab={}
| Assignment 2. Ethics and Sentiment Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''.venv'': venv)'
# name: python_defaultSpec_1599791128016
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/kangsuek/image-background-remove-tool/blob/master/docs/other/try.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="sqwsUfoI3SnG" colab_type="text"
# # Install image-background-remove-tool
# + id="7C4rC_HQi1gq" colab_type="code" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="4398e711-6fa5-4412-b124-92028cf14a2f"
#@title Choose branch
branch = 'master' #@param {type: "string"}
# !git clone https://github.com/kangsuek/image-background-remove-tool.git -b $branch
# %cd /content/image-background-remove-tool
# !pip install -r requirements.txt
# !cd ./tools && echo "all" | python setup.py
# + tags=[]
# 서버의 GPU availability 상태확인
# !nvidia-smi # Jupyter Notebook에서 명령어를 입력할 경우
# + tags=[]
# 사용가능한 GPU 디바이스의 갯수, 현재 셋업 되어있는 GPU 넘버, 그리고 GPU 디바이스의 이름을 출력
import torch
# 현재 Setup 되어있는 device 확인
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print ('Available devices ', torch.cuda.device_count())
print ('Current cuda device ', torch.cuda.current_device())
print(torch.cuda.get_device_name(device))
# + [markdown] id="pF-4SVcB3gjK" colab_type="text"
# # 배경이미지 제거 테스트
# + id="B6XlZr9pBB8b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} cellView="form" outputId="9a10fd4b-11a4-4a69-b143-9375c74a944f" tags=[]
#@markdown ### *img_url의 sample image를 1.jpg부터 31.jpg까지 변경해보며 테스트하면 됩니다.
img_url = 'https://github.com/kangsuek/image-background-remove-tool/raw/master/docs/imgs/input/1.jpg' #@param {type: "string"}
preprocessing = "bbd-fastrcnn" #@param ["bbd-fastrcnn", "bbmd-maskrcnn", "None"] {allow-input: false}
model_name = "u2net" #@param ["u2net", "basnet", "u2netp", "mobile_net_model", "xception_model"] {allow-input: false}
postprocessing = "rtb-bnb" #@param ["rtb-bnb", "rtb-bnb2", "No"] {allow-input: false}
# !wget -q -O ./1.jpg "$img_url" > /dev/null
from IPython.display import display
from PIL import Image
import os
# 입력 이미지
display(Image.open("./1.jpg"))
if os.path.exists("./1.jpg"):
f = "1.jpg"
# !python main.py -i ./1.jpg -o ./1.png -m $model_name -prep $preprocessing -postp $postprocessing
image = Image.open("./1.png")
new = Image.new("RGB", image.size, (255, 255, 255))
new.paste(image, (0,0), image)
# 결과이미지
display(new)
| docs/other/try.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
quantitative_data = pd.read_csv("../../resulting data/quantitative_data.csv", sep="\t", index_col=0)
quantitative_data.head()
quantitative_data.describe()
quantitative_data.plot.box(rot=90, figsize=(15,5))
quantitative_data.plot(kind="bar", x="code", y="verses", color="green",figsize=(10,5))
relative_quantitative_data = quantitative_data.copy().plot.bar(rot=90, figsize=(15,5), x="code")
# +
relative_quantitative_data = quantitative_data.copy()
for column in relative_quantitative_data.columns:
if(relative_quantitative_data[column].dtype == np.float64 or relative_quantitative_data[column].dtype == np.int64):
relative_quantitative_data[column] = relative_quantitative_data[column]/quantitative_data["verses"]
else:
relative_quantitative_data[column]
# -
relative_quantitative_data.head()
relative_quantitative_data.plot.box(rot=90, figsize=(15,5))
relative_quantitative_data.plot(kind="bar", x="code", y=["diff pers","diff orgs", "diff plas"], figsize=(18,5), title="Amount of different entities in each book of the Bible (relative to amount of verses)")
relative_quantitative_data.plot(kind="bar", x="code", y=["pers","orgs", "plas"], figsize=(20,5))
relative_quantitative_data.plot(kind="bar", x="code", y=["qs-oral", "qs-written","qs-prayer"], figsize=(20,5))
relative_quantitative_data.plot(kind="bar", x="code", y=["qs-written", "qs-soCalled","qs-prayer","qs-song","qs-idea",], figsize=(20,5))
# +
relative_quantitative_data.plot(kind="bar", x="code", y=["qs-dream","qs-oath"], figsize=(20,5))
# -
qs_columns = ["qs-dream","qs-oath","qs-written", "qs-soCalled","qs-song","qs-idea","qs-oral", "qs-prayer"]
sorted(qs_columns)
relative_quantitative_data.plot(kind="bar", x="code", y=["qs-oral", "qs-written","qs-prayer"], figsize=(20,5))
relative_quantitative_data.plot(kind="bar", x="code", y=["qs-oral", "qs-written","qs-prayer"], figsize=(20,5))
quantitative_data.plot.scatter(x="verses",y="pericopes")
quantitative_data.plot.scatter(x="diff orgs",y="diff plas")
| code/python/.ipynb_checkpoints/testing visualizations of books-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # An Introduction to Graph Neural Networks
# By <NAME>, on 20 June 2020
# <br> <br>
# Graphs in the real world, such as social networks, chemical molecules and biological knowledge graphs, are rich with information that cannot be found in individual entities. A method for learning graph representations or node classification would be extremely valuable. Unfortunately, the modern deep learning toolbox is designed for grids (ie. images) and simple sequences (ie. text). CNNs and RNNs cannot generalize to graphs that have arbitrary size, complex topological structures and no fixed node ordering. Graph neural networks (GNN) provide a powerful tool to learn representations from any arbitrary graph structure by leveraging local network neighborhoods. This tutorial aims to (1) introduce the concept of graph neural networks, (2) discuss the quatitative motivation behind different GNN architectures and (3) implement these architectures using the PyTorch Geometric library.
# <br> <br>
# +
from IPython.lib.display import YouTubeVideo
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
from torch_geometric.data import Data, GraphSAINTRandomWalkSampler
from torch_geometric.datasets import Planetoid, Entities
from torch_geometric.nn import GCNConv, RGCNConv, GATConv, SAGEConv, JumpingKnowledge, GINConv, DeepGraphInfomax
torch.manual_seed(200620)
np.random.seed(200620)
# -
# ## What is a Graph Neural Network?
# Graph neural networks have a somewhat universal architecture where forward propagation uses a 'neighborhood aggregation' technique. The model iteratively produces new feature representations for a given node by aggregating the current feature representations of its adjacent nodes’ (ie. neighbors) and the node itself. Here, an iteration is parametrized by a layer of the neural network. This means that the computational graph of the neural network is defined by the neighborhood of each node. And, each layer in the graph neural network can be thought of as a step where each node aggregates messages from its neighboring nodes, as visualized by Microsoft in this [video](https://www.youtube.com/watch?v=cWIeTMklzNg):
YouTubeVideo('cWIeTMklzNg', height=500, width=890, start=70, end=460)
# We have a graph $\mathcal{G} = (\mathcal{V}, \mathcal{E})$ described by an adjacency matrix $A$. The goal is to learn a function of node input features for node classification. The model takes as input a feature descriptions summarized by matrix $X$ which is $N \times D$ ($N$: number of nodes, $D$: number of input features) and produces a node-level output $Z$ (a $N \times C$ feature matrix, where $C$ is the number of classes per node). At the node level, each layer of the GNN is doing neighborhood aggregation to transform the node representation. At the graph level, every neural network layer can then be written as a non-linear function $f$:
# <br><br>$$H^{(l+1)}=f^{(l+1)}(H^{(l)},A)$$<br>
# with $H^{(0)}=X$ and $H^{(L)}=Z$, where L is the number of layers. Notice that each node started with an input vector of length $D$ and ended at a classification vector of length $C$, where $D$ need not equal $C$. This is possible because after doing neighborhood aggregation, $f$ applies a non-linear transformation to node representations using its parameters, a weight matrix $W$ and an activation $\sigma$. The dimension of node representations will change from one layer to another when $W$ is not a square matrix. Note that each layer learns it's own parameters. The specific GNN architechtures then differ only in how $f(⋅,⋅)$ is chosen and parameterized.
# ## GNN Architectures
#
# In recent years, many architectures of graph neural networks have been introduced. I will explore the quantitative motivations behind some of the most influential architectures in the field. I will also implement simple models using these architectures in PyTorch and evaluate them on the benchmark Cora dataset.
#
#
# ### Dataset
print('For this tutorial, we will use a standard citation dataset that is commonly used to benchmark GNN performance.')
print('The Cora dataset is a homogeneous, undirected graph where nodes are publications linked by citations.')
dataset = Planetoid(root='/tmp/Cora', name='Cora', split='full')
print('It contains:')
graph = dataset[0]
print('\t- {:d} labels'.format(dataset.num_classes))
print('\t- {:d} nodes'.format(graph.num_nodes))
print('\t- {:d} training'.format(graph.train_mask.sum().item()))
print('\t- {:d} validation'.format(graph.val_mask.sum().item()))
print('\t- {:d} testing'.format(graph.test_mask.sum().item()))
# ### Tools for Learning
# #### Hyperparameters
# +
class Hyperparameters():
def __init__(self):
self.num_node_features = None
self.num_classes = None
self.lr = 0.005
self.w_decay = 5e-4
self.dropout = 0.3
self.epochs = 200
self.cuda = True
self.device = None
args = Hyperparameters()
args.num_node_features = graph.num_node_features
args.num_classes = dataset.num_classes
args.cuda = args.cuda and torch.cuda.is_available()
if args.cuda:
args.device = torch.device('cuda:2')
else:
args.device = torch.device('cpu')
# -
# #### Supervised learning
# The `LearnGraph`class can train an arbitrary GNN on an arbitrary graph for node classification. By default, binary cross entropy loss and adam optimizer are used.
# + code_folding=[]
class LearnGraph():
def __init__(self, graph, model, args, criterion=None):
self.args = args
self.graph = graph.to(self.args.device)
self.model = model.to(self.args.device)
if not criterion:
criterion = nn.CrossEntropyLoss()
self.criterion = criterion
self.optim = torch.optim.Adam(self.model.parameters(), lr=self.args.lr, weight_decay=self.args.w_decay)
self.train_loss = []
self.val_loss = []
self.train_complete = False
def learn(self) -> None:
# tracks training and validation loss over epochs
# can add early stopping mechanism by comparing losses
for epoch in range(self.args.epochs):
if self.train_complete: return
tl = self.train_epoch()
self.train_loss.append(tl)
vl = self.val()
self.val_loss.append(vl)
self.train_complete = True
def train_epoch(self) -> float:
# trains a single epoch (ie. one pass over the full graph) and updates the models parameters
# returns the loss
self.model.train()
labels = self.graph.y[self.graph.train_mask]
self.optim.zero_grad()
output = self.model.forward(self.graph)
loss = self.criterion(output[self.graph.train_mask], labels)
loss.backward()
self.optim.step()
return loss.data.item()
def val(self) -> float:
# returns the validation loss
self.model.eval()
labels = self.graph.y[self.graph.val_mask]
output = self.model.forward(self.graph)
loss = self.criterion(output[self.graph.val_mask], labels)
return loss.data.item()
def test(self) -> float:
# returns the test accuracy
if not self.train_complete:
self.learn()
self.model.eval()
labels = self.graph.y[self.graph.test_mask]
_, pred = self.model.forward(self.graph).max(dim=1)
correct = float ( pred[self.graph.test_mask].eq(labels).sum().item() )
acc = correct / self.graph.test_mask.sum().item()
return acc
# -
# ### Graph Convolutional Networks
# **[GCNConv](https://pytorch-geometric.readthedocs.io/en/latest/modules/nn.html#torch_geometric.nn.conv.GCNConv)** from Kipf and Welling: [Semi-Supervised Classification with Graph Convolutional Networks](https://arxiv.org/abs/1609.02907) (ICLR 2017)
# Also see [this blog by Kipf](http://tkipf.github.io/graph-convolutional-networks/)
# <br> <br>
# The basic propagation rule is $$f: \sigma\left(A H^{(l)} W^{(l)}\right)$$
# According to the adjacency matrix, we sum the feature vectors of all neighboring nodes but not the node itself. We fix this by enforcing self-loops: $\hat{A} = A + I$
# Another limitation is that $\hat{A}$ is not normalized, so multiplication with $\hat{A}$ will completely change the scale of the feature vectors. Let $\hat{D}$ be the diagonal node degree matrix. Simple normalization involves using $\hat{D}^{-1} \hat{A} \text{ instead of } \hat{A}$. In practice, we do symmetric normalization such that the final propagation rule is <br><br>
# $$f: \sigma\left(\hat{D}^{-\frac{1}{2}} \hat{A} \hat{D}^{-\frac{1}{2}} H^{(l)} W^{(l)}\right)$$<br>
# For a given node $i$ feature vector, where $ N_{i}$ is its neighborhood and $c_{i j}$ is a normalization constant for the edge $(i, j)$, the update is: <br> <br>
# $$h_{i}^{(l+1)}=\sigma\ \left(\ \sum_{j \ \in \ N_{i}} \frac{1}{c_{i j}} \ h_{j}^{(l)} \ W^{(l)}\ \right)$$<br>
#
#
# +
class GNN(torch.nn.Module):
def __init__(self):
super(GNN, self).__init__()
self.conv1 = GCNConv(args.num_node_features, 64)
self.conv2 = GCNConv(64, args.num_classes)
self.transition = nn.Sequential(
nn.ReLU(),
nn.Dropout(p=args.dropout)
)
def forward(self, graph):
x, edge_index = graph.x, graph.edge_index
x = self.conv1(x, edge_index)
x = self.transition(x)
x = self.conv2(x, edge_index)
return x
learner = LearnGraph(model=GNN(), graph=graph, args=args)
acc = learner.test()
print('Accuracy: {:.1%}'.format(acc))
# -
# ### Relational Graph Convolutional Networks
# **[RGCNConv](https://pytorch-geometric.readthedocs.io/en/latest/modules/nn.html#torch_geometric.nn.conv.RGCNConv)** from Schlichtkrull *et al.*: [Modeling Relational Data with Graph Convolutional Networks](https://arxiv.org/abs/1703.06103) (ESWC 2018)
# <br> <br>
# 'Knowledge' graphs have nodes connected by many different relationships. To capture many types of links in graphs, we can apply relation-specific transformations on incoming messages. For a specific node $i$, $\mathcal{N}_{i}^{r}$ contains its neighbors connected by link $r$. $c_{i, r}$ is a normalization constant (such as $|\mathcal{N}_{i}^{r}|$).
# <br> <br> $$h_{i}^{(l+1)}=\sigma\left(\ \sum_{r \ \in \ \mathcal{R}} \ \sum_{j \ \in \ \mathcal{N}_{i}^{r}} \frac{1}{c_{i, r}} \ W_{r}^{(l)} h_{j}^{(l)}+W_{0}^{(l)} h_{i}^{(l)}\right)$$<br>
# The parameters of the network grow rapidly with the number of relations in the graph. We need regularlization to prevent overfitting on rare relations. Using basis decompostion, each $W_{r}^{(l)}$ is defined as a linear combination of basis vectors $V_{b}^{(l)} \in \mathbb{R}^{d^{(l+1)} \times d^{(l)}}$, the space of $W_{r}^{(l)}$. Only the coefficients $a_{r b}^{(l)}$ depend on $r$. This method alleviates overfitting by creating weight sharing across frequent and rare relations.
# <br> <br> $$W_{r}^{(l)}=\sum_{b=1}^{B} a_{r b}^{(l)} V_{b}^{(l)}$$<br>
# Since R-GCN is applied to heterogenous graphs, we will use the MUTAG graph instead of Cora. RGCN auto-generates unique embeddings as input features for nodes if no features are provided, as in MUTAG. Minor changes made to `LearnGraph`to accomodate MUTAG.
# + code_folding=[]
mutag_dataset = Entities(root='/tmp/MUTAG', name='MUTAG')
mutag_graph = mutag_dataset[0]
class LearnMUTAG():
def __init__(self, graph, model, args, criterion=None):
self.args = args
self.graph = graph.to(self.args.device)
self.model = model.to(self.args.device)
if not criterion:
criterion = nn.CrossEntropyLoss()
self.criterion = criterion
self.optim = torch.optim.Adam(self.model.parameters(), lr=self.args.lr, weight_decay=self.args.w_decay)
self.train_loss = []
self.train_complete = False
def learn(self) -> None:
for epoch in range(self.args.epochs):
if self.train_complete: return
tl = self.train_epoch()
self.train_loss.append(tl)
self.train_complete = True
def train_epoch(self) -> float:
self.model.train()
labels = self.graph.train_y
self.optim.zero_grad()
output = self.model.forward(self.graph)
loss = self.criterion(output[self.graph.train_idx], labels)
loss.backward()
self.optim.step()
return loss.data.item()
def test(self) -> float:
# returns the test accuracy
if not self.train_complete:
self.learn()
self.model.eval()
labels = self.graph.test_y
_, pred = self.model.forward(self.graph).max(dim=1)
correct = float ( pred[self.graph.test_idx].eq(labels).sum().item() )
acc = correct / len(self.graph.test_idx)
return acc
class GNN(torch.nn.Module):
def __init__(self):
super(GNN, self).__init__()
self.conv1 = RGCNConv(mutag_graph.num_nodes, 16, mutag_dataset.num_relations, num_bases=30)
self.conv2 = RGCNConv(16, mutag_dataset.num_classes, mutag_dataset.num_relations, num_bases=30)
self.transition = nn.Sequential(
nn.ReLU(),
nn.Dropout(p=args.dropout)
)
def forward(self, graph):
x, edge_index, edge_type = graph.x, graph.edge_index, graph.edge_type
x = self.conv1(x, edge_index, edge_type)
x = self.transition(x)
x = self.conv2(x, edge_index, edge_type)
return x
learner = LearnMUTAG(model=GNN(), graph=mutag_graph, args=args)
acc = learner.test()
print('Accuracy: {:.1%}'.format(acc))
# -
# ### Graph Attention Networks
# **[GATConv](https://pytorch-geometric.readthedocs.io/en/latest/modules/nn.html#torch_geometric.nn.conv.GATConv)** from Veličković *et al.*: [Graph Attention Networks](https://arxiv.org/abs/1710.10903) (ICLR 2018)
# <br> <br>
# GATConv is inspired by the use of self-attention (Google transformer) which has achieved SOTA performance in NLP. The idea is that nodes can 'attend' over their neighbors and select the direction from which they recieve information. For a single layer, where nodes go from $F$ to $F^{\prime}$ features, the following steps are applied:
#
# 1. Linear transformation of input features using a weight matrix, $\mathbf{W} \in \mathbb{R}^{F^{\prime} \times F}$
#
# 2. Compute attention coefficients $e_{i j} \in \mathbb{R}$ for each node pair $(i, j)$ using a shared attention mechanism $a$. Here this mechanism is a single-layer feedforward network network with a the parameter vector, $\overrightarrow{\mathbf{a}} \in \mathbb{R}^{2 F^{\prime}}$
# <br> <br>$$e_{i j}=a\left(\mathbf{W} \vec{h}_{i}, \mathbf{W} \vec{h}_{j}\right) = \text{LeakyReLU}\left(\overrightarrow{\mathbf{a}}^{T}\left[\mathbf{W} \vec{h}_{i} \| \mathbf{W} \vec{h}_{j}\right]\right)$$ <br>
#
# 3. Normalize attention coefficients across nodes: $\alpha_{i j}=\operatorname{softmax}_{j}\left(e_{i j}\right)$
#
# 4. Compute output features as a linear combination of input features corresponding to their normalized attention coefficients
# <br> <br> $$\vec{h}_{i}^{\prime}=\sigma\left(\sum_{j \in \mathcal{N}_{i}} \alpha_{i j} \mathbf{W} \vec{h}_{j}\right)$$ <br>
# 5. This process can be stabilized using multi-head attention, which concatenates/averages multiple indepedently computed output features from independent attention layers <br> <br>
#
#
# +
class GNN(torch.nn.Module):
def __init__(self):
super(GNN, self).__init__()
self.conv1 = GATConv(args.num_node_features, 8, heads=8)
self.conv2 = GATConv(64, args.num_classes, heads=1)
self.transition = nn.Sequential(
nn.ReLU(),
nn.Dropout(p=args.dropout)
)
def forward(self, graph):
x, edge_index = graph.x, graph.edge_index
x = self.conv1(x, edge_index)
x = self.transition(x)
x = self.conv2(x, edge_index)
return x
learner = LearnGraph(model=GNN(), graph=graph, args=args)
acc = learner.test()
print('Accuracy: {:.1%}'.format(acc))
# -
# ### GraphSAGE
# **[SAGEConv](https://pytorch-geometric.readthedocs.io/en/latest/modules/nn.html#torch_geometric.nn.conv.SAGEConv)** from Hamilton *et al.*: [Inductive Representation Learning on Large Graphs](https://arxiv.org/abs/1706.02216) (NIPS 2017)
# <br> <br>
# GraphSAGE generalizes GCN to use trainable aggregation functions beyond simple convolutions. For a given layer $(l+1)$, at each node $i$, the embeddings $h_{j}^{(l)}$ for all nodes $j$ in its neighborhood $\mathcal{N}(i)$ are combined into a single vector $h^{(l+1)}_{\mathcal{N}(i)}$ using an aggregation function. Aggregating the neighborhood separately from the node itself implictly adds skip connections across layers.
# <br> <br> $$\mathbf{h}_{\mathcal{N}(i)}^{(l+1)} = \operatorname{AGGREGATE}_{(l+1)}\left(\left\{\mathbf{h}_{j}^{(l)}, \forall j \in \mathcal{N}(i)\right\}\right)$$ <br>
# Aggregation functions can include mean (~traditional GCN), LSTM and Pooling. In max pooling, each neighbor vector is fed through a single-layer neural network and then an elementwise max-pooling operation is applied. Max pooling implicitly selects the important nodes, much like Graph Attention Networks.
# <br> <br> $$\mathrm{AGGREGATE}_{(l+1)}^{\mathrm{pool}}=\max \left(\left\{\sigma\left(\mathbf{W}_{\mathrm{pool}} \ \mathbf{h}_{j}^{(l+1)}+\mathbf{b}\right), \forall j \in \mathcal{N}(i)\right\}\right)$$ <br>
# The final step is to concatenate the node's current representation $h^{(l)}_{i}$ with the aggregated neighborhood vector $h^{(l+1)}_{\mathcal{N}(i)}$. This concatenated vector is fed through a single-layer neural network to calculate the output representation $h^{(l+1)}_{i}$
# <br> <br> $$\mathbf{h}_{i}^{(l+1)} = \sigma \left(W_{(l+1)} \left[\ h^{(l)}_{i} \ \| \ h^{(l+1)}_{\mathcal{N}(i)}\ \right] \right)$$ <br>
#
# +
class GNN(torch.nn.Module):
def __init__(self):
super(GNN, self).__init__()
self.conv1 = SAGEConv(args.num_node_features, 64, normalize=True)
self.conv1.aggr = 'max'
self.conv2 = SAGEConv(64, args.num_classes, normalize=True)
self.conv2.aggr = 'max'
self.transition = nn.Sequential(
nn.ReLU(),
nn.Dropout(p=args.dropout)
)
print()
def forward(self, graph):
x, edge_index = graph.x, graph.edge_index
x = self.conv1(x, edge_index)
x = self.transition(x)
x = self.conv2(x, edge_index)
return x
learner = LearnGraph(model=GNN(), graph=graph, args=args)
acc = learner.test()
print('Accuracy: {:.1%}'.format(acc))
# -
# ### Jumping Knowledge Networks
# **[Jumping Knowledge](https://pytorch-geometric.readthedocs.io/en/latest/modules/nn.html#torch_geometric.nn.models.JumpingKnowledge)** from Xu *et al.*: [Representation Learning on Graphs with Jumping Knowledge Networks](https://arxiv.org/abs/1806.03536) (ICML 2018)
# <br> <br>
# Many aggregation based models achieve best performance with 2 layer networks. After that, performance degrades despite theoretically greater access to information and even after adding residual connections. In biological networks, the majority of the nodes have few connections, whereas some nodes are hubs. In the same graph, the same number of GNN layers an lead to very different effects for different nodes.
# <br> <br>
# Unlike GAT or GraphSAGE which select the direction of expansion, Jumping Knowledge operates on the locality of expansion. This model proposes two architectural changes – jump connections and a subsequent selective but adaptive aggregation mechanism. As in common neighborhood aggregation networks, each layer increases the size of the influence distribution by aggregating neighborhoods from the previous layer. In the last JK layer, for each node, we select from all of those intermediate representations. If this is done independently for each node, then the model can adapt the effective neighborhood size for each node as needed, resulting in exactly the desired adaptivity. The layer aggregation mechanisms can include concatenation, max pooling, and lstm attention.
# <br> <br>
# +
class GNN(torch.nn.Module):
def __init__(self):
super(GNN, self).__init__()
self.conv1 = GCNConv(args.num_node_features, 64)
self.convx= GCNConv(64, 64)
self.jk = JumpingKnowledge(mode='max')
self.final = nn.Linear(64, args.num_classes)
self.transition = nn.Sequential(
nn.ReLU(),
nn.Dropout(p=args.dropout)
)
def forward(self, graph):
x, edge_index = graph.x, graph.edge_index
xs = []
x = self.conv1(x, edge_index)
x = self.transition(x)
xs.append(x)
for _ in range(5):
x = self.convx(x, edge_index)
x = self.transition(x)
xs.append(x)
x = self.jk(xs)
x = self.final(x)
return x
learner = LearnGraph(model=GNN(), graph=graph, args=args)
acc = learner.test()
print('Accuracy: {:.1%}'.format(acc))
# -
# ### Graph Isomorphism Network
# **[GINConv](https://pytorch-geometric.readthedocs.io/en/latest/modules/nn.html#torch_geometric.nn.conv.GINConv)** from Xu *et al.*: [How Powerful are Graph Neural Networks?](https://arxiv.org/abs/1810.00826) (ICLR 2019)
# <br> <br>
# This work aimed to increase the expressive power of GNNs. The Weisfeiler-Lehman (WL) test for graph isomorphism uses injective node aggregation to distinguish graphs from each other. They show that GNNs can be as powerful as the WL test in distinguishing graph structures if the GNN’s aggregation scheme is highly expressive and can model injective (one-one) functions. A nodes neighbors are defined as a multiset, i.e., a set with possibly repeating elements. Sum aggregators can represent injective (and universal) functions over multisets. On the other hand, mean or max aggregators (used in GraphSAGE or GCN) are not injective multiset functions. The Graph Isomorphism Network uses a multi-layer perceptron to do neighborhood aggregation. Note that this argument holds under their key assumption that "node input features are from a countable universe" which is a very simplistic view of input features.
# <br> <br>
# +
class GNN(torch.nn.Module):
def __init__(self):
super(GNN, self).__init__()
self.transition = nn.Sequential(
nn.ReLU(),
nn.Dropout(p=args.dropout)
)
self.mlp1 = nn.Sequential(
nn.Linear(args.num_node_features, 256),
nn.ReLU(),
nn.Linear(256, 64),
)
self.conv1 = GINConv(self.mlp1)
self.mlp2 = nn.Sequential(
nn.Linear(64, 16),
nn.ReLU(),
nn.Linear(16, args.num_classes),
)
self.conv2= GINConv(self.mlp2)
def forward(self, graph):
x, edge_index = graph.x, graph.edge_index
x = self.conv1(x, edge_index)
x = self.transition(x)
x = self.conv2(x, edge_index)
return x
learner = LearnGraph(model=GNN(), graph=graph, args=args)
acc = learner.test()
print('Accuracy: {:.1%}'.format(acc))
# -
# ### Deep Graph Infomax
# **[Deep Graph Infomax](https://pytorch-geometric.readthedocs.io/en/latest/modules/nn.html#torch_geometric.nn.models.DeepGraphInfomax)** from Veličković *et al.*: [Deep Graph Infomax](https://arxiv.org/abs/1809.10341) (ICLR 2019)
# <br> <br>
# Unsupervised embeddings have traditionally been trained with random-walk objectives, which can over-emphasize proximity information at the expense of structural information. Since encoders already enforce an inductive bias that neighboring nodes have similar representations, it is unclear whether random-walk objectives actually provide any useful signal. Deep graph infomax is an alternative objective for unsupervised graph learning that is based upon mutual information.
# <br> <br>
# The DGI objective seeks to maximize local mutual information by obtaining obtain node (i.e., local) representations that capture the global information content of the entire graph. As all of the derived patch representations are driven to preserve mutual information with the global graph summary, this allows for discovering and preserving similarities on the patch-level. This is useful because distant nodes with similar structural roles are known to be a strong predictor for many node classification tasks.
# <br> <br>
# The DGI model is defined by an encoder $\mathcal{E}$, discriminator $\mathcal{D}$, readout function $\mathcal{R}$ and corruption function $\mathcal{C}$. The GNN encoder outputs node representations $h_{i}$. The readout function gives a summary vector, $\vec{s}$, which is the global graph representation. The discriminator takes the graph summary $s$ and a node $h_{i}$ and assigns a co-occurence probablity $\mathcal{D}(h_{i}, s)$ to the pair. For negative samples, the graph structured is randomly changed using $\mathcal{C}$, and the same process is repeated. The objective is structured in binary cross entropy form:
# <br> <br>$$\mathcal{L}=\frac{1}{N+M}\left(\sum_{i=1}^{N} \mathbb{E}_{(\mathbf{X}, \mathbf{A})}\left[\log \mathcal{D}\left(\vec{h}_{i}, \vec{s}\right)\right]+\sum_{j=1}^{M} \mathbb{E}_{(\widetilde{\mathbf{X}}, \widetilde{\mathbf{A}})}\left[\log \left(1-\mathcal{D}\left(\overrightarrow{\widetilde{h}_{j}}, \vec{s}\right)\right)\right]\right)$$ <br>
# For supervised learning, we can incorporate the DGI objective in an intermediate, hidden layer in conjunction with the traditional BCE loss at the final layer.<br> <br>
# + code_folding=[]
class LearnDeepGraphInfomax():
def __init__(self, graph, enc_dgi, enc_cls, args, criterion=None):
self.args = args
self.graph = graph.to(self.args.device)
self.dgi_model = DeepGraphInfomax(enc_dgi.hidden_ch, enc_dgi, enc_dgi.summary, enc_dgi.corruption)
self.dgi_model = self.dgi_model.to(self.args.device)
self.cls_model = enc_cls.to(self.args.device)
if not criterion:
criterion = nn.CrossEntropyLoss()
self.criterion = criterion
parameters = [*self.dgi_model.parameters()] + [*self.cls_model.parameters()]
self.optim = torch.optim.Adam(parameters, lr=self.args.lr, weight_decay=self.args.w_decay)
self.train_loss = []
self.val_loss = []
self.train_complete = False
def learn(self) -> None:
for epoch in range(self.args.epochs):
if self.train_complete: return
tl = self.train_epoch()
self.train_loss.append(tl)
vl = self.val()
self.val_loss.append(vl)
self.train_complete = True
def train_epoch(self) -> float:
self.dgi_model.train()
self.cls_model.train()
labels = self.graph.y[self.graph.train_mask]
self.optim.zero_grad()
pos_z, neg_z, summary = self.dgi_model.forward(x=self.graph.x, edge_index=self.graph.edge_index)
output = self.cls_model.forward(pos_z, self.graph.edge_index)
loss = self.dgi_model.loss(pos_z, neg_z, summary) + self.criterion(output[self.graph.train_mask], labels)
loss.backward()
self.optim.step()
return loss.data.item()
def val(self) -> float:
self.dgi_model.eval()
self.cls_model.eval()
labels = self.graph.y[self.graph.val_mask]
pos_z, neg_z, summary = self.dgi_model.forward(self.graph.x, self.graph.edge_index)
output = self.cls_model.forward(pos_z, self.graph.edge_index)
loss = self.dgi_model.loss(pos_z, neg_z, summary) + self.criterion(output[self.graph.val_mask], labels)
return loss.data.item()
def test(self) -> float:
if not self.train_complete:
self.learn()
self.dgi_model.eval()
self.cls_model.eval()
labels = self.graph.y[self.graph.test_mask]
pos_z, neg_z, summary = self.dgi_model.forward(self.graph.x, self.graph.edge_index)
_, pred = self.cls_model.forward(pos_z, self.graph.edge_index).max(dim=1)
correct = float ( pred[self.graph.test_mask].eq(labels).sum().item() )
acc = correct / self.graph.test_mask.sum().item()
return acc
class Encoder_DGI(torch.nn.Module):
def __init__(self, hidden_ch=64):
super(Encoder_DGI, self).__init__()
self.hidden_ch = hidden_ch
self.conv = GCNConv(args.num_node_features, hidden_ch)
self.activation = nn.PReLU()
def corruption(self, x, edge_index):
# corrupted features are obtained by row-wise shuffling of the original features
# corrupted graph consists of the same nodes but located in different places
return x[torch.randperm(x.size(0))], edge_index
def summary(self, z, *args, **kwargs):
return torch.sigmoid(z.mean(dim=0))
def forward(self, x, edge_index):
x = self.conv(x, edge_index)
x = self.activation(x)
return x
class Encoder_CLS(torch.nn.Module):
def __init__(self, hidden_ch=64):
super(Encoder_CLS, self).__init__()
self.conv = GCNConv(hidden_ch, args.num_classes)
def forward(self, x, edge_index):
return self.conv(x, edge_index)
learner = LearnDeepGraphInfomax(enc_dgi=Encoder_DGI(), enc_cls=Encoder_CLS(), graph=graph, args=args)
acc = learner.test()
print('Accuracy: {:.1%}'.format(acc))
# -
# ### GraphSAINT
# **[GraphSAINT](https://pytorch-geometric.readthedocs.io/en/latest/modules/data.html#torch_geometric.data.GraphSAINTSampler)** from Zeng *et al.*: [GraphSAINT: Graph Sampling Based Inductive Learning Method](https://arxiv.org/abs/1907.04931) (ICLR 2020)
# <br> <br>
# As the GNN becomes deeper, training time can grow exponentially due to "neighbor explosion". GraphSAINT samples the training graph first and then builds a full GNN on the subgraph. Intuitively, nodes of higher influence on each other should have higher probability to form a subgraph. This enables the sampled nodes to “support” each other without going outside the minibatch. Unfortunately, such strategy results in non-identical node sampling probability, and introduces bias in the minibatch estimator. GraphSAINT employs normalization techniques so that the feature learning does not give preference to nodes more frequently sampled. The GraphSAINT sampler can be applied to any graph irrespective of the GNN being trained on that graph.
# <br> <br>
# +
class LearnGraphSAINT():
def __init__(self, graph, model, args, criterion=None):
self.args = args
self.graph = graph.to(self.args.device)
self.model = model.to(self.args.device)
self.loader = GraphSAINTRandomWalkSampler(self.graph, batch_size=100, walk_length=2, num_steps=self.args.epochs)
if not criterion:
criterion = nn.CrossEntropyLoss()
self.criterion = criterion
self.optim = torch.optim.Adam(self.model.parameters(), lr=self.args.lr, weight_decay=self.args.w_decay)
self.train_loss = []
self.val_loss = []
self.train_complete = False
def learn(self) -> None:
for epoch, batch in enumerate(self.loader):
if self.train_complete: return
tl = self.train_batch(batch)
self.train_loss.append(tl)
vl = self.val()
self.val_loss.append(vl)
self.train_complete = True
def train_batch(self, batch) -> float:
self.model.train()
labels = batch.y[batch.train_mask]
self.optim.zero_grad()
output = self.model.forward(batch)
loss = self.criterion(output[batch.train_mask], labels)
loss.backward()
self.optim.step()
return loss.data.item()
def val(self) -> float:
self.model.eval()
labels = self.graph.y[self.graph.val_mask]
output = self.model.forward(self.graph)
loss = self.criterion(output[self.graph.val_mask], labels)
return loss.data.item()
def test(self) -> float:
if not self.train_complete:
self.learn()
self.model.eval()
labels = self.graph.y[self.graph.test_mask]
_, pred = self.model.forward(self.graph).max(dim=1)
correct = float ( pred[self.graph.test_mask].eq(labels).sum().item() )
acc = correct / self.graph.test_mask.sum().item()
return acc
class GNN(torch.nn.Module):
def __init__(self):
super(GNN, self).__init__()
self.conv1 = GCNConv(args.num_node_features, 64)
self.conv2 = GCNConv(64, args.num_classes)
self.transition = nn.Sequential(
nn.ReLU(),
nn.Dropout(p=args.dropout)
)
def forward(self, graph):
x, edge_index = graph.x, graph.edge_index
x = self.conv1(x, edge_index)
x = self.transition(x)
x = self.conv2(x, edge_index)
return x
learner = LearnGraphSAINT(model=GNN(), graph=graph, args=args)
acc = learner.test()
print('Accuracy: {:.1%}'.format(acc))
# -
# ## More...
# * PyTorch Geometric [GitHub](https://github.com/rusty1s/pytorch_geometric) – even more architectures and examples
# * Customizing GNNs – using the [Message Passing base class](https://pytorch-geometric.readthedocs.io/en/latest/notes/create_gnn.html) in PyTorch Geometric
# * [Deep Graph Library](https://www.dgl.ai) – an alternative to PyTorch Geometric
| 01-intro/gnn-intro.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="LcTzETSHkQVV"
# # Deep learning with fast.ai cookbook - MNIST "hello world" example
# Minimal example using the MNIST dataset to demonstrate a minimimal, "hello world" type fast.ai application.
#
# Here are the fastai curated dataset choices for MNIST:
#
# **MNIST_SAMPLE**: 3s and 7s only
# ~~~
# |-- train
# | |-- 3
# | `-- 7
# `-- valid
# |-- 3
# `-- 7
# ~~~
#
# **MNIST_TINY**: 3s and 7s only
# ~~~
# |-- models
# |-- test
# |-- train
# | |-- 3
# | `-- 7
# `-- valid
# |-- 3
# `-- 7
# ~~~
#
# **MNIST_VAR_SIZE_TINY**: 3s and 7s only
# ~~~
# |-- models
# |-- test
# |-- train
# | |-- 3
# | `-- 7
# `-- valid
# |-- 3
# `-- 7
# ~~~
#
# **MNIST**: complete dataset
# ~~~
# |-- testing
# | |-- 0
# | |-- 1
# | |-- 2
# | |-- 3
# | |-- 4
# | |-- 5
# | |-- 6
# | |-- 7
# | |-- 8
# | `-- 9
# `-- training
# |-- 0
# |-- 1
# |-- 2
# |-- 3
# |-- 4
# |-- 5
# |-- 6
# |-- 7
# |-- 8
# `-- 9
# ~~~
#
# + colab={"base_uri": "https://localhost:8080/"} id="7abffi5ckQVZ" outputId="259860bc-69c5-41b6-e583-92fc51be77c1"
# imports for notebook boilerplate
# !pip install -Uqq fastbook
import fastbook
from fastbook import *
from fastai.vision.all import *
# + colab={"base_uri": "https://localhost:8080/"} id="W30zzDOHkQVb" outputId="6f75dc0d-0eba-4b0f-8386-80057294da11"
# set up the notebook for fast.ai
fastbook.setup_book()
# + colab={"base_uri": "https://localhost:8080/", "height": 37} id="QdA_eLdikQVb" outputId="ddcc80a6-bb94-4b77-83dd-c0d065e29746"
# In Gradient, datasets get saved in /storage/data when untar_data is called
# if the dataset has not been copied there already
path = untar_data(URLs.MNIST)
# + colab={"base_uri": "https://localhost:8080/"} id="buwpbGBxkQVc" outputId="8d71ebc6-4d75-485e-85ac-f15b47bd9a56"
# examine the directory structure of the dataset
path.ls()
# + colab={"base_uri": "https://localhost:8080/", "height": 116} id="SmIsJL1LkQVc" outputId="d762c475-0c1b-4c04-8388-aaa39d58e4f1"
# %%time
# create an image dataloaders object using the path
# note that because of the directory structure of the dataset
# the train and valid sets have to be explicitly specified
# details here: https://github.com/fastai/fastai/issues/1129
dls = ImageDataLoaders.from_folder(path, train='training', valid='testing')
# create a learner object using the dataloaders that was just defined
# architecture is resnet18; see https://pytorch.org/hub/pytorch_vision_resnet/
# loss function is selected for multi class classification
# accuracy is the metric used to optimize
learn = cnn_learner(dls, resnet18, pretrained=False,
loss_func=LabelSmoothingCrossEntropy(), metrics=accuracy)
# fit the model for one epoch using 1cycle policy
# see https://docs.fast.ai/callback.schedule.html#Learner.fit_one_cycle
learn.fit_one_cycle(1, 0.1)
# + colab={"base_uri": "https://localhost:8080/", "height": 192} id="PBkUx2JekQVd" outputId="3b7d3725-0c4a-4064-a6c5-2e5f4221b557"
# show a batch of training data
dls.train.show_batch(max_n=4, nrows=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 192} id="zz-dmspLkQVd" outputId="4722672c-f7ae-4115-994e-1ac236fdfe08"
# show a batch of validation data
dls.valid.show_batch(max_n=4, nrows=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 45} id="IDCbDmRVkQVe" outputId="fd350bd5-f9a0-46e8-9d76-c6caff11227a"
# take a look at one of the test images
img_files = get_image_files(path/"testing")
img = PILImage.create(img_files[7000])
img
# + colab={"base_uri": "https://localhost:8080/", "height": 233} id="pUjS4G7VkQVf" outputId="317e77e9-4e8a-4908-d011-5d6fc765a624"
# show the images with the highest loss
interp = ClassificationInterpretation.from_learner(learn)
interp.plot_top_losses(4, nrows=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="OwgibxldkQVf" outputId="58ed7367-3eed-498d-85c3-cbcbcdc7dbd1"
# examine the model, including its layers
learn.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 45} id="sxxjZZfGkQVg" outputId="7bdbf7c7-23ab-4bd0-b45d-85befedee2ab"
# select an image from the test set
img = PILImage.create(img_files[0])
img
# + colab={"base_uri": "https://localhost:8080/", "height": 70} id="8zK47Ta3kQVg" outputId="d3273ff3-0e1b-4943-c5dd-7f3e393dfea6"
# apply the trained model to the image
learn.predict(img)
# + colab={"base_uri": "https://localhost:8080/", "height": 45} id="O41AhTUCkQVg" outputId="faa54fa6-1890-4116-921b-132ffef4f201"
# select a different test set image and apply the model to it to get a prediction
img = PILImage.create(img_files[2030])
img
# + colab={"base_uri": "https://localhost:8080/", "height": 70} id="DDZYZ6UQkQVh" outputId="ae3f334a-382f-46b6-f2ae-9fdcaec0155e"
learn.predict(img)
# + colab={"base_uri": "https://localhost:8080/", "height": 45} id="v-VJpctskQVh" outputId="a09740ef-6cac-4cbe-ab55-048a32364efe"
# select a different image and apply the model to it to get a prediction
img = PILImage.create(img_files[5800])
img
# + colab={"base_uri": "https://localhost:8080/", "height": 70} id="n1DMQd8ZkQVh" outputId="3508c065-9ddf-4bd7-8912-8eaff704b7f8"
learn.predict(img)
# + id="B92WVU7akQVh"
| _notebooks/2022-03-23-mnist_hello_world.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neural art check-point viewer
# Simple notebook to load the images saved during an nstyle training run and view them together.
import glob
dir_name = 'output/'
prefix = dir_name + 'e_'
files = glob.glob(prefix+'*.jpg')
# Sort by numeric part
files.sort(key=lambda x: int(x[len(prefix):-4]))
files.append(dir_name+'final.jpg')
files[:5]
# Set up some code to display the images in a notebook
import IPython.display
def show_img(fn):
h = IPython.display.HTML("<img src='"+fn+"'>")
IPython.display.display(h)
# ## Animated display
# Flip through the frames fairly quickly for a movie-like experience.
# NOTE: this tends to **work much better the second time** through after
# the browser has cached all the images.
# Animated Display
import time
for img in files:
IPython.display.clear_output()
print(img)
show_img(img)
time.sleep(.3)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from skimage import io
for img_fn in files:
print(img_fn)
im = io.imread(img_fn)
plt.figure(figsize=(3,2))
plt.axis('off')
plt.imshow(im)
plt.show()
# -
| checkpoint-viewer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Dual-tone multi-frequency (DTMF) signaling
#
# DTMF signaling is the way analog phones send the number dialed by a user over to the central phone office. This was in the day before all-digital networks and cell phones were the norm, but the method is still used for in-call option selection ("press 4 to talk to customer service"...).
#
# The mechanism is rather clever: the phone's keypad is arranged in a $4\times 3$ grid and each button is associated to *two* frequencies according to this table:
#
#
# | | **1209 Hz** | **1336 Hz** | **1477 Hz** |
# |------------|:-----------:|:-----------:|:-----------:|
# | **697 Hz** | 1 | 2 | 3 |
# | **770 Hz** | 4 | 5 | 6 |
# | **852 Hz** | 7 | 8 | 9 |
# | **941 Hz** | * | 0 | # |
#
#
# The frequencies in the table have been chosen so that they are "coprime"; in other words, no frequency is a multiple of any other, which reduces the probability of erroneously detecting the received signals due to interference. When a button is pressed, the two corresponding frequencies are generated simultaneously and sent over the line. For instance, if the digit '1' is pressed, the generated signal will be:
#
# $$
# x(t) = \sin(2\pi\cdot 1209\cdot t) + \sin(2\pi\cdot697\cdot t)
# $$
#
#
# The official specifications for the DTMF standard further stipulate that:
#
# * each tone should be at least 65ms long
# * tones corresponding to successive digits should be separated by a silent gap of at least 65ms
#
#
# In this notebook we will build a DTMF decoder based on the Discrete Fourier Transform.
# Of course here we will use discrete-time signals exclusively so, if the clock of the system is $F_s$, each DTMF tone will be of the form:
# $$
# x[n] = \sin(2\pi\,(f_l/F_s)\, n) + \sin(2\pi\,(f_h/F_s)\,n)
# $$
#
# The first thing to do is to write a DTMF encoder.
# $f_l$ the lower frequency and $f_h$ the higher one.
#
# **(?1)** What is the clock of the system $F_s$?
# +
# first our usual bookkeeping
# %pylab inline
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
import IPython
# the "clock" of the system
FS = 24000
# -
# **Note** that the
# > `Populating the interactive namespace from numpy and matplotlib`
#
# was the by-product of `%pylab inline`
def dtmf_dial(number):
DTMF = {
'1': (697, 1209), '2': (697, 1336), '3': (697, 1477),
'4': (770, 1209), '5': (770, 1336), '6': (770, 1477),
'7': (852, 1209), '8': (852, 1336), '9': (852, 1477),
'*': (941, 1209), '0': (941, 1336), '#': (941, 1477),
}
MARK = 0.1
SPACE = 0.1
n = np.arange(0, int(MARK * FS))
x = np.array([])
for d in number:
s = np.sin(2*np.pi * DTMF[d][0] / FS * n) + np.sin(2*np.pi * DTMF[d][1] / FS * n)
x = np.concatenate((x, s, np.zeros(int(SPACE * FS))))
return x
# OK, that was easy. Let's test it and evaluate it "by ear":
# +
x=dtmf_dial('123##45')
IPython.display.Audio(x, rate=FS)
# -
# #### Stopped here (2020/10/28 (水) 16h00)
# - Next time remember that you should make an effort to understand the inner working of `dtmf_dial()`
# Now let's start thinking about the decoder. We will use the following strategy:
#
# * split the signal into individual digit tones by looking at the position of the gaps
# * perform a DFT on the digit tones
# * look at the peaks of the Fourier transform and recover the dialed number
#
# Here we assume whe have the whole signal in memory, i.e. we will perform *batch* processing; clearly a more practical system would decode the incoming signal as it arrives sample by sample (real-time processing); you are more than encouraged to try and implement such an algorithm.
#
# To split the signal the idea is to look at the local energy over small windows: when the signal is silence, we will cut it.
#
# Let's see how we can do that; let's look at the raw data first
plt.plot(x);
# OK so, clearly, we should be able to find the high and low energy sections of the signal. Let's say that we use an analysis window of 240 samples which, at our $F_s$ corresponds to an interval of 10ms. We can easily find the local energy like so:
# +
# split the signal in 240-sample chunks and arrange them as rows in a matrix
# (truncate the data vector to a length multiple of 240 to avoid errors)
w = np.reshape(x[:(len(x)/240)*240], (-1, 240))
# compute the energy of each chunk by summing the squares of the elements of each row
we = np.sum(w * w, axis=1)
plt.plot(we);
# -
# From the plot, it appears clearly that we can set a threshold of about 200 to separate tone sections from silence sections. Let's write a function that returns the start and stop indices of the tone sections in an input signal
def dtmf_split(x, win=240, th=200):
edges = []
w = np.reshape(x[:(len(x)/win)*win], (-1, win))
we = np.sum(w * w, axis=1)
L = len(we)
ix = 0
while ix < L:
while ix < L and we[ix] < th:
ix = ix+1
if ix >= L:
break # ending on silence
iy = ix
while iy < L and we[iy] > th:
iy = iy+1
edges.append((ix * win, iy * win))
ix = iy
return edges
print dtmf_split(x)
# Looks good. Now that we have a splitter, let's run a DFT over the tone sections and find the DTMF frequencies that are closest to the peaks of the DFT magnitude. The "low" DTMF frequencies are in the 697 Hz to 941 Hz range, while the high frequencies in the 1209 Hz to 1477 Hz range, so we will look for a DFT peak in each of those intervals. For instance, let's look at the first tone, and let's look at the peaks in the DFT:
X = abs(np.fft.fft(x[0:2400]))
plt.plot(X[0:500]);
# We clearly have identifiable peaks. The only thing we need to pay attention to is making sure that we map real-world frequencies to the DFT plot correctly (and vice versa).
def dtmf_decode(x, edges = None):
# the DTMF frequencies
LO_FREQS = np.array([697.0, 770.0, 852.0, 941.0])
HI_FREQS = np.array([1209.0, 1336.0, 1477.0])
KEYS = [['1', '2', '3'], ['4', '5', '6'], ['7', '8', '9'], ['*', '0', '#']]
# frequency ranges to search for low and high DTMF tones
LO_RANGE = (680.0, 960.0)
HI_RANGE = (1180.0, 1500.0)
number = []
# now examine each tone in turn. the freqency mapping on the DFT
# axis will be dependent on the length of the data vector
if edges is None:
edges = dtmf_split(x)
for g in edges:
# compute the DFT of the tone segment
X = abs(np.fft.fft(x[g[0]:g[1]]))
N = len(X)
# compute the resolution in Hz of a DFT bin
res = float(FS) / N
# find the peak location within the low freq range
a = int(LO_RANGE[0] / res)
b = int(LO_RANGE[1] / res)
lo = a + np.argmax(X[a:b])
# find the peak location within the high freq range
a = int(HI_RANGE[0] / res)
b = int(HI_RANGE[1] / res)
hi = a + np.argmax(X[a:b])
# now match the results to the DTMF frequencies
row = np.argmin(abs(LO_FREQS - lo * res))
col = np.argmin(abs(HI_FREQS - hi * res))
# and finally convert that to the pressed key
number.append(KEYS[row][col])
return number
dtmf_decode(x)
# Yay! It works! As always, in communication systems, the receiver is much more complicated than the receiver.
#
# Of course this is a very simplified setup and we have glossed over a lot of practical details. For instance, in the splitting function, the thresholds are not determined dynamically and this may create problems in the presence of noise. Similarly, we just detect a frequency peak in the spectrum, but noise may make things more complicated.
#
# For instance, listen to the following noise-corrupted version of the original signal. Although the tones are still detectable by ear, the segmentation algorithm fails and returns a single digit.
# +
noisy = x + np.random.uniform(-2, 2, len(x))
IPython.display.Audio(noisy, rate=FS)
# -
dtmf_decode(noisy)
# If we **carefully** change the segmentation threshold, we can still decode
dtmf_decode(x, dtmf_split(x, th=220))
# but if we're not careful...
dtmf_decode(x, dtmf_split(x, th=250))
# The sensitivity to the segmentation threshold confirms the fact that segmentation should be performed using more sophisticated techniques, which what happens in practical systems.
| epfl/2020/hw-ipynb/DTMF/DTMF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Sort and Clean conference data.
# It writes to `sorted_data.yml` and `cleaned_data.yml`, copy those to the conference.yml after screening.
import yaml
import datetime
import sys
from shutil import copyfile
from builtins import input
import pytz
# +
try:
# for python newer than 2.7
from collections import OrderedDict
except ImportError:
# use backport from pypi
from ordereddict import OrderedDict
try:
from yaml import CLoader as Loader, CDumper as Dumper
except ImportError:
from yaml import Loader, Dumper
from yaml.representer import SafeRepresenter
_mapping_tag = yaml.resolver.BaseResolver.DEFAULT_MAPPING_TAG
def dict_representer(dumper, data):
return dumper.represent_dict(data.iteritems())
def dict_constructor(loader, node):
return OrderedDict(loader.construct_pairs(node))
Dumper.add_representer(OrderedDict, dict_representer)
Loader.add_constructor(_mapping_tag, dict_constructor)
Dumper.add_representer(str,
SafeRepresenter.represent_str)
def ordered_dump(data, stream=None, Dumper=yaml.Dumper, **kwds):
class OrderedDumper(Dumper):
pass
def _dict_representer(dumper, data):
return dumper.represent_mapping(
yaml.resolver.BaseResolver.DEFAULT_MAPPING_TAG,
data.items())
OrderedDumper.add_representer(OrderedDict, _dict_representer)
return yaml.dump(data, stream, OrderedDumper, **kwds)
dateformat = '%Y-%m-%d %H:%M:%S'
tba_words = ["tba","tbd"]
right_now = datetime.datetime.utcnow().replace(microsecond=0).strftime(dateformat)
# -
def query_yes_no(question, default="no"):
"""Ask a yes/no question via input() and return their answer.
"question" is a string that is presented to the user.
"default" is the presumed answer if the user just hits <Enter>.
It must be "yes" (the default), "no" or None (meaning
an answer is required of the user).
The "answer" return value is True for "yes" or False for "no".
"""
valid = {"yes": True, "y": True, "ye": True,
"no": False, "n": False}
if default is None:
prompt = " [y/n] "
elif default == "yes":
prompt = " [Y/n] "
elif default == "no":
prompt = " [y/N] "
else:
raise ValueError("invalid default answer: '%s'" % default)
while True:
sys.stdout.write(question + prompt)
choice = input().lower()
if default is not None and choice == '':
return valid[default]
elif choice in valid:
return valid[choice]
else:
sys.stdout.write("Please respond with 'yes' or 'no' "
"(or 'y' or 'n').\n")
with open("../_data/conferences.yml", 'r') as stream:
try:
data = yaml.load(stream, Loader=Loader)
print("Initial Sorting:")
for q in data:
print(q["deadline"]," - ",q["name"])
print("\n\n")
conf = [x for x in data if x['deadline'].lower() not in tba_words]
tba = [x for x in data if x['deadline'].lower() in tba_words]
# just sort:
conf.sort(key=lambda x: pytz.utc.normalize(datetime.datetime.strptime(x['deadline'], dateformat).replace(tzinfo=pytz.timezone(x['timezone']))))
print("Date Sorting:")
for q in conf+tba:
print(q["deadline"]," - ",q["name"])
print("\n\n")
conf.sort(key=lambda x: pytz.utc.normalize(datetime.datetime.strptime(x['deadline'], dateformat).replace(tzinfo=pytz.timezone(x['timezone']))).strftime(dateformat) < right_now)
print("Date and Passed Deadline Sorting with tba:")
for q in conf+tba:
print(q["deadline"]," - ",q["name"])
print("\n\n")
with open('sorted_data.yml', 'w') as outfile:
for line in ordered_dump(conf+tba, Dumper=yaml.SafeDumper, default_flow_style=False, explicit_start=True).replace('\'', '"').splitlines():
outfile.write('\n')
outfile.write(line.replace('- name:', '\n- name:'))
except yaml.YAMLError as exc:
print(exc)
if query_yes_no("Did you check the sorted data and would like to replace the original data?"):
copyfile('sorted_data.yml','../_data/conferences.yml')
with open('sorted_data.yml', 'r') as stream:
try:
conf = yaml.load(stream, Loader=Loader)
print("Initial Data:")
for q in conf:
print(q["deadline"]," - ",q["name"])
print("\n\n")
clean_conf = []
for q in conf:
dates,year=q["date"].split(",")
start_date = dates.strip().split(" ")[0].strip()+" "+dates.split("-")[1].strip()+" "+year.strip()
try:
datetime.datetime.strptime(start_date, "%B %d %Y").strftime(dateformat)
except ValueError:
start_date = dates.split("-")[1].strip()+" "+year.strip()
if datetime.datetime.strptime(start_date, "%B %d %Y").strftime(dateformat) >= right_now:
clean_conf.append(q)
else:
print("Passed: "+q["deadline"]," - ",q["name"])
print("\n\n")
print("Cleaned Data:")
for q in clean_conf:
print(q["deadline"]," - ",q["name"])
with open('cleaned_data.yml', 'w') as outfile:
for line in ordered_dump(clean_conf, Dumper=yaml.SafeDumper, default_flow_style=False, explicit_start=True).replace('\'', '"').splitlines():
outfile.write('\n')
outfile.write(line.replace('- name:', '\n- name:'))
except yaml.YAMLError as exc:
print(exc)
if query_yes_no("Did you check the cleaned data and would like to replace the original data?"):
copyfile('cleaned_data.yml','../_data/conferences.yml')
# Thanks to https://gist.github.com/oglops/c70fb69eef42d40bed06
| utils/Data_Handling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="wJcYs_ERTnnI"
# ##### Copyright 2021 The TensorFlow Authors.
# + cellView="form" id="HMUDt0CiUJk9"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="77z2OchJTk0l"
# # Migrate the fault tolerance mechanism
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/guide/migrate/fault_tolerance">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/guide/migrate/fault_tolerance.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/guide/migrate/fault_tolerance.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/guide/migrate/fault_tolerance.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="n4O6fPyYTxZv"
# Fault tolerance refers to a mechanism of periodically saving the states of trackable objects, such as parameters and models. This enables you to recover them in the event of a program/machine failure during training.
#
# This guide first demonstrates how to add fault tolerance to training with `tf.estimator.Estimator` in TensorFlow 1 by specifying metric saving with `tf.estimator.RunConfig`. Then, you will learn how to implement fault tolerance for training in Tensorflow 2 in two ways:
#
# - If you use the Keras `Model.fit` API, you can pass the `tf.keras.callbacks.BackupAndRestore` callback to it.
# - If you use a custom training loop (with `tf.GradientTape`), you can arbitrarily save checkpoints using the `tf.train.Checkpoint` and `tf.train.CheckpointManager` APIs.
#
# Both of these methods will back up and restore the training states in [checkpoint](../../guide/checkpoint.ipynb) files.
#
# + [markdown] id="pHJfmkCFUhQf"
# ## Setup
# + id="VXnPvQi8Ui1F"
import tensorflow.compat.v1 as tf1
import tensorflow as tf
import numpy as np
import tempfile
import time
# + id="Tww-uIoiUlsT"
mnist = tf.keras.datasets.mnist
(x_train, y_train),(x_test, y_test) = mnist.load_data()
x_train, x_test = x_train / 255.0, x_test / 255.0
# + [markdown] id="TtlucRG_Uro_"
# ## TensorFlow 1: Save checkpoints with tf.estimator.RunConfig
#
# In TensorFlow 1, you can configure a `tf.estimator` to save checkpoints every step by configuring `tf.estimator.RunConfig`.
#
# In this example, start by writing a hook that artificially throws an error during the fifth checkpoint:
# + id="Q8shCkV2jKcc"
class InterruptHook(tf1.train.SessionRunHook):
# A hook for artificially interrupting training.
def begin(self):
self._step = -1
def before_run(self, run_context):
self._step += 1
def after_run(self, run_context, run_values):
if self._step == 5:
raise RuntimeError('Interruption')
# + [markdown] id="ZXbQ6cFlkoIM"
# Next, configure `tf.estimator.Estimator` to save every checkpoint and use the MNIST dataset:
# + id="1EKXzi4Qj2Eb"
feature_columns = [tf1.feature_column.numeric_column("x", shape=[28, 28])]
config = tf1.estimator.RunConfig(save_summary_steps=1,
save_checkpoints_steps=1)
path = tempfile.mkdtemp()
classifier = tf1.estimator.DNNClassifier(
feature_columns=feature_columns,
hidden_units=[256, 32],
optimizer=tf1.train.AdamOptimizer(0.001),
n_classes=10,
dropout=0.2,
model_dir=path,
config = config
)
train_input_fn = tf1.estimator.inputs.numpy_input_fn(
x={"x": x_train},
y=y_train.astype(np.int32),
num_epochs=10,
batch_size=50,
shuffle=True,
)
# + [markdown] id="sGP7Nyenk1gr"
# Begin training the model. An artificial exception will be raised by the hook you defined earlier.
# + id="xWKMsmt6jYSL"
try:
classifier.train(input_fn=train_input_fn,
hooks=[InterruptHook()],
max_steps=10)
except Exception as e:
print(f'{type(e).__name__}:{e}')
# + [markdown] id="DekxJkgWk-4N"
# Rebuild the `tf.estimator.Estimator` using the last saved checkpoint and continue training:
# + id="vqMVTiJMjcH7"
classifier = tf1.estimator.DNNClassifier(
feature_columns=feature_columns,
hidden_units=[256, 32],
optimizer=tf1.train.AdamOptimizer(0.001),
n_classes=10,
dropout=0.2,
model_dir=path,
config = config
)
classifier.train(input_fn=train_input_fn,
max_steps = 10)
# + [markdown] id="T5LtVtmvYx7J"
# ## TensorFlow 2: Back up and restore with a callback and Model.fit
#
# In TensorFlow 2, if you use the Keras `Model.fit` API for training, you can provide the `tf.keras.callbacks.BackupAndRestore` callback to add the fault tolerance functionality.
#
# To help demonstrate this, let's first start by defining a callback class that artificially throws an error during the fifth checkpoint:
#
# + id="Ci3yB6A5lwJu"
class InterruptingCallback(tf.keras.callbacks.Callback):
# A callback for artificially interrupting training.
def on_epoch_end(self, epoch, log=None):
if epoch == 4:
raise RuntimeError('Interruption')
# + [markdown] id="AhU3VTYZoDh-"
# Then, define and instantiate a simple Keras model, define the loss function, call `Model.compile`, and set up a `tf.keras.callbacks.BackupAndRestore` callback that will save the checkpoints in a temporary directory:
# + id="1VOQLDNkl2bl"
def create_model():
return tf.keras.models.Sequential([
tf.keras.layers.Flatten(input_shape=(28, 28)),
tf.keras.layers.Dense(512, activation='relu'),
tf.keras.layers.Dropout(0.2),
tf.keras.layers.Dense(10)
])
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
model = create_model()
model.compile(optimizer='adam',
loss=loss,
metrics=['accuracy'],
steps_per_execution=10)
log_dir = tempfile.mkdtemp()
backup_restore_callback = tf.keras.callbacks.BackupAndRestore(
backup_dir = log_dir
)
# + [markdown] id="LRRWmZqsvMrq"
# Now, start training the model with `Model.fit`. During training, checkpoints will be saved thanks to the `backup_restore_callback` defined above, while the `InterruptingCallback` will raise an artificial exception to simulate a failure.
# + id="8bVO79qWl4Uv"
try:
model.fit(x=x_train,
y=y_train,
epochs=10,
validation_data=(x_test, y_test),
callbacks=[backup_restore_callback, InterruptingCallback()])
except Exception as e:
print(f'{type(e).__name__}:{e}')
# + [markdown] id="EWidh234vcRf"
# Next, instantiate the Keras model, call `Model.compile`, and continue training the model with `Model.fit` from a previously saved checkpoint:
# + id="3IWPH0Cmn2wi"
model = create_model()
model.compile(optimizer='adam',
loss=loss,
metrics=['accuracy'],
steps_per_execution=10)
model.fit(x=x_train,
y=y_train,
epochs=10,
validation_data=(x_test, y_test),
callbacks=[backup_restore_callback])
# + [markdown] id="OdWexHUUaEB6"
# ## TensorFlow 2: Write manual checkpoints with a custom training loop
#
# If you use a custom training loop in TensorFlow 2, you can implement a fault tolerance mechanism with the `tf.train.Checkpoint` and `tf.train.CheckpointManager` APIs.
#
# This example demonstrates how to:
#
# - Use a `tf.train.Checkpoint` object to manually create a checkpoint, where the trackable objects you want to save are set as attributes.
# - Use a `tf.train.CheckpointManager` to manage multiple checkpoints.
#
# Start by defining and instantiating the Keras model, the optimizer, and the loss function. Then, create a `Checkpoint` that manages two objects with trackable states (the model and the optimizer), as well as a `CheckpointManager` for logging and keeping several checkpoints in a temporary directory.
# + id="hPnIRKC8aDwE"
model = create_model()
optimizer = tf.keras.optimizers.SGD(learning_rate=0.001)
loss_fn = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
log_dir = tempfile.mkdtemp()
epochs = 5
steps_per_epoch = 5
checkpoint = tf.train.Checkpoint(model=model, optimizer=optimizer)
checkpoint_manager = tf.train.CheckpointManager(
checkpoint, log_dir, max_to_keep=2)
# + [markdown] id="L2tK4fm6xNkJ"
# Now, implement a custom training loop where after the first epoch every time a new epoch starts the last checkpoint is loaded:
# + id="GhQphF5jxPWU"
for epoch in range(epochs):
if epoch > 0:
tf.train.load_checkpoint(save_path)
print(f"\nStart of epoch {epoch}")
for step in range(steps_per_epoch):
with tf.GradientTape() as tape:
logits = model(x_train, training=True)
loss_value = loss_fn(y_train, logits)
grads = tape.gradient(loss_value, model.trainable_weights)
optimizer.apply_gradients(zip(grads, model.trainable_weights))
save_path = checkpoint_manager.save()
print(f"Checkpoint saved to {save_path}")
print(f"Training loss at step {step}: {loss_value}")
# + [markdown] id="rQUS8nO9FZlH"
# ## Next steps
#
# To learn more about fault tolerance and checkpointing in TensorFlow 2, consider the following documentation:
#
# - The `tf.keras.callbacks.BackupAndRestore` callback API docs.
# - The `tf.train.Checkpoint` and `tf.train.CheckpointManager` API docs.
# - The [Training checkpoints](../../guide/checkpoint.ipynb) guide, including the _Writing checkpoints_ section.
#
# You may also find the following material related to [distributed training](../..guide/distributed_training.ipynb) useful:
#
# - The _Fault tolerance_ section in the [Multi-worker training with Keras](../../tutorials/distribute/multi_worker_with_keras.ipynb) tutorial.
# - The _Handing task failure_ section in the [Parameter server training](../../tutorials/distribute/parameter_server_training.ipynb) tutorial.
| site/en-snapshot/guide/migrate/fault_tolerance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# #%%appyter init
# NOTE: The following line is only for debugging, in regular appyters using it from pip, it would be omitted
import os, sys; sys.path.insert(0, os.path.realpath('..'))
from appyter import magic
magic.init(lambda _=globals: _())
# # My Title
import numpy as np
import plotly.graph_objects as go
for n in range(2):
fig = go.Figure()
X = np.random.normal(loc=0, scale=1, size=(10000, 2))
fig.add_trace(go.Scattergl(
mode='markers',
x=X[:, 0],
y=X[:, 1],
text=[f"{x}, {y}" for x, y in X],
))
fig.show()
for n in range(2):
fig = go.Figure()
X = np.random.normal(loc=0, scale=1, size=(500000, 2))
fig.add_trace(go.Scattergl(
mode='markers',
x=X[:, 0],
y=X[:, 1],
text=[f"{x}, {y}" for x, y in X],
))
fig.show()
for n in range(2):
fig = go.Figure()
X = np.random.normal(loc=0, scale=1, size=(10000, 2))
fig.add_trace(go.Scattergl(
mode='markers',
x=X[:, 0],
y=X[:, 1],
text=[f"{x}, {y}" for x, y in X],
))
fig.show()
| example/droptest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import requests
import os
import json
# +
def connect_to_twitter():
bearer_token = os.environ.get("BEARER_TOKEN")
return {"Authorization" : "Bearer {}".format(bearer_token)}
headers = connect_to_twitter()
# +
def make_request(headers):
url = "https://api.twitter.com/2/tweets/search/recent?query=ExtremeWeather"
return requests.request("GET", url , headers=headers).jsonon()
response = make_request(headers)
print(response)
| Extraccion/TwitterExtract/TwitterAPI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: myDL
# language: python
# name: mydl
# ---
# +
# Date: 15-03-2020
# Author: <NAME>
# -
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import random
from scipy.optimize import curve_fit
from scipy.special import erf
# %matplotlib inline
df_confirmed = pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv')
df_deaths = pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv')
df_recovered = pd.read_csv('https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv')
df_confirmed.head()
# # Morocco
df_confirmed[df_confirmed['Country/Region']=='Morocco'].loc[:,'3/2/20':] #'3/2/20'
# #### draw the curves
plt.figure(figsize=(12, 8))
plt.xticks(rotation=50,size=8)
plt.plot(list(df_confirmed)[44:],df_confirmed[df_confirmed['Country/Region']=='Morocco'].loc[:, '3/2/20':].values[0,:], '*-',color='blue')
plt.plot(list(df_recovered)[44:],df_recovered[df_recovered['Country/Region']=='Morocco'].loc[:, '3/2/20':].values[0,:], '*-',color='green')
plt.plot(list(df_deaths)[44:],df_deaths[df_deaths['Country/Region']=='Morocco'].loc[:, '3/2/20':].values[0,:], '*-', color='red')
plt.legend(['confimed','Recovered','death'],fontsize=12)
plt.title('COVID-19 in Morocco')
plt.xlabel('Date',size=12)
plt.ylabel('Cases',size=12)
#plt.style.use('dark_background')
plt.show()
# # 7-day forecast of COVID-19 infections in Morocco
# At this time, data is still scarce and not suitable for accurate modeling, these results are then highly uncertain for now.
# #### Fit an exponential curve (worst case, not realistic)
# population is finite thus an exponential model is not realistic but gives insights about the beggining of an infection
# #### function to extend dates
from datetime import timedelta, datetime
## https://github.com/Lewuathe/COVID19-SIR/blob/master/solver.py
def extend_index(index, new_size):
values = index.values
current = datetime.strptime(index[-1], '%m/%d/%y')
while len(values) < new_size:
current = current + timedelta(days=1)
values = np.append(values, datetime.strftime(current, '%m/%d/%y'))
return values
dataConfirmedMorocco = df_confirmed[df_confirmed['Country/Region']=='Morocco'].loc[:, '3/2/20':].iloc[0]
dataRecoveredMorocco = df_recovered[df_recovered['Country/Region']=='Morocco'].loc[:, '3/2/20':].iloc[0]
dataDeathMorocco = df_deaths[df_deaths['Country/Region']=='Morocco'].loc[:, '3/2/20':].iloc[0]
nbdays = (datetime.today() - datetime.strptime('3/2/20', '%m/%d/%y')).days + 1 #nbdays of available data from 3/2/20
# #### curve fitting
# +
U,V = curve_fit(lambda t,a,b,c: a*np.exp(b*t)+c,
np.arange(1,nbdays),
dataConfirmedMorocco)
# -
U, V
plt.figure(figsize=(12, 8))
plt.xticks(rotation=50,size=8)
plt.plot(list(df_confirmed)[44:],df_confirmed[df_confirmed['Country/Region']=='Morocco'].loc[:, '3/2/20':].values[0,:], '*-',color='blue')
plt.plot(extend_index(dataConfirmedMorocco.index,nbdays+7),U[0]*np.exp(U[1]*np.arange(1,nbdays+8))+U[2], '--',color='magenta')
plt.legend(['actual confirmed','exponential fit'],fontsize=12)
plt.title('7 days forecast - Exponential fit for confirmed cases')
plt.xlabel('Date',size=12)
plt.ylabel('Cases',size=12)
#plt.style.use('dark_background')
plt.show()
# #### prediction for the next 7 days
np.floor(U[0]*np.exp(U[1]*np.arange(nbdays,nbdays+7))+U[2])
# ### Fit a logisitic curve (also not very realistic)
# This model is widely used for estimating the growth of a population and also to simply model infections
def logistic_model(x,a,b,c):
return c/(1+np.exp(-(a*x+b)))
# +
nbdays = (datetime.today() - datetime.strptime('3/2/20', '%m/%d/%y')).days + 1 #nbdays of available data from 3/2/20
U,V = curve_fit(logistic_model,
np.arange(1,nbdays),
dataConfirmedMorocco,
p0=[2,-60,10000])#p0=[2,-60,1000]
# -
U,V
# #### !!!!!!!!!!!!!! data is not sufficient to find a good logistic fit...
plt.figure(figsize=(12, 8))
plt.xticks(rotation=50,size=8)
plt.plot(list(df_confirmed)[44:],df_confirmed[df_confirmed['Country/Region']=='Morocco'].loc[:, '3/2/20':].values[0,:], '*-',color='blue')
plt.plot(extend_index(dataConfirmedMorocco.index,nbdays+17),U[2]/(1+np.exp(-(U[0]*np.arange(1,nbdays+18)+U[1]))), '--',color='magenta')
plt.legend(['actual confirmed','Logistic fit'],fontsize=12)
plt.title('7 days forecast - Logistic fit for confirmed cases')
plt.xlabel('Date',size=12)
plt.ylabel('Cases',size=12)
#plt.style.use('dark_background')
plt.show()
np.floor(U[2]/(1+np.exp(-(U[0]*np.arange(nbdays,nbdays+8)+U[1]))))
# somewhat similar to the exp curve for now (it's just the beginning in Morocco...)
# # !!!! SIR model !!!!
# #### Realistic model known to epidemiologists, check the excellent video by 3blue1brown https://www.youtube.com/watch?v=gxAaO2rsdIs
# ### Now let's find the best $\beta$ and $\gamma$ for Morocco data
# +
N=3e7
def loss(point, data, recovered, N, s_0, i_0, r_0):
#https://github.com/Lewuathe/COVID19-SIR/blob/master/solver.py
size = len(data)
beta, gamma = point
def SIR(t, y):
S = y[0]
I = y[1]
R = y[2]
return [-beta*S*I/N, beta*S*I/N-gamma*I, gamma*I]
solution = solve_ivp(SIR, [0, size], [s_0,i_0,r_0], t_eval=np.arange(0, size, 1), vectorized=True)
l1 = np.sqrt(np.mean((solution.y[1] - data)**2))
l2 = np.sqrt(np.mean((solution.y[2] - recovered)**2))
alpha = 0.1
return alpha * l1 + (1 - alpha) * l2
# +
from scipy.integrate import solve_ivp
from scipy.optimize import minimize
data = dataConfirmedMorocco - dataRecoveredMorocco - dataDeathMorocco
###
s_0 = 3e7
i_0 = 1
r_0 = 0
###
result = minimize(loss, [0.001, 0.001], args=(data.astype('float64'), dataRecoveredMorocco.astype('float64'), N, s_0, i_0, r_0), method='L-BFGS-B', bounds=[(0., 1), (0., 1)], options={'disp':10})#, bounds=[(0.00000001, 0.4), (0.00000001, 0.4)]
beta, gamma = result.x
print(beta,gamma,beta/gamma)
# -
def SIR(t, y):
S = y[0]
I = y[1]
R = y[2]
return [-beta*S*I/N, beta*S*I/N-gamma*I, gamma*I]
# +
predict_range = nbdays + 150
new_index = extend_index(dataConfirmedMorocco.index, predict_range)
size = len(new_index)
predictMorocco = solve_ivp(SIR, [0, size], [s_0,i_0,r_0], t_eval=np.arange(0, size, 1))
# +
infected_actual = np.concatenate((data.values, [None] * (size - len(data.values))))
recovered_actual = np.concatenate((dataRecoveredMorocco.values, [None] * (size - len(data.values))))
plt.figure(figsize=(20,10))
plt.plot(new_index[::2], infected_actual[::2], '*-',color='black', label='actual infected')
plt.plot(new_index[::2], recovered_actual[::2], '*-',color='magenta', label='actual recovered')
plt.plot(new_index[::2], predictMorocco.y[0,::2], label='Suspected')
plt.plot(new_index[::2], predictMorocco.y[1,::2], label='Infected')
plt.plot(new_index[::2], predictMorocco.y[2,::2], label='Recovered')
plt.xticks(rotation=90)
plt.legend()
# -
# #### same but log-scale
plt.figure(figsize=(20,10))
plt.semilogy(new_index[::2], infected_actual[::2], '*-',color='black', label='actual infected')
plt.semilogy(new_index[::2], recovered_actual[::2], '*-',color='magenta', label='actual recovered')
plt.semilogy(new_index[::2], predictMorocco.y[0,::2], label='Suspected')
plt.semilogy(new_index[::2], predictMorocco.y[1,::2], label='Infected')
plt.semilogy(new_index[::2], predictMorocco.y[2,::2], label='Recovered')
plt.xticks(rotation=90)
plt.legend()
# # !!!! SEIR model !!!!
def lossSEIR(point, data, recovered, N, s_0, e_0, i_0, r_0):
#https://github.com/Lewuathe/COVID19-SIR/blob/master/solver.py
size = len(data)
alpha, beta, gamma = point
def SEIR(t, y):
S = y[0]
E = y[1]
I = y[2]
R = y[3]
return [-beta*S*I/N, beta*S*I/N - alpha*E, alpha*E - gamma*I, gamma*I]
solution = solve_ivp(SEIR, [0, size], [s_0,e_0,i_0,r_0],
t_eval=np.arange(0, size, 1), vectorized=True)
l1 = np.sqrt(np.mean((solution.y[2] - data)**2))
l2 = np.sqrt(np.mean((solution.y[3] - recovered)**2))
alpha = 0.5
#print(l1,l2, alpha * l1 + (1 - alpha) * l2)
return alpha * l1 + (1 - alpha) * l2
# +
from scipy.integrate import solve_ivp
from scipy.optimize import minimize
data = dataConfirmedMorocco - dataRecoveredMorocco - dataDeathMorocco
###
s_0 = 3e7-1
e_0 = 0
i_0 = 1
r_0 = 0
###
result = minimize(lossSEIR,
[0.1,0.1,0.1],
args=(data.astype('float64'), dataRecoveredMorocco.astype('float64'), N, s_0, e_0, i_0, r_0),
method='L-BFGS-B',
bounds=[(0., 5.), (0., 10.), (0., 5.)])#, bounds=[(0.00000001, 0.4), (0.00000001, 0.4)]
alpha, beta, gamma = result.x
print(alpha, beta, gamma, beta/gamma)
# -
def SEIR(t, y):
S = y[0]
E = y[1]
I = y[2]
R = y[3]
return [- beta*S*I/N, beta*S*I/N - alpha*E, alpha*E - gamma*I, gamma*I]
# +
predict_range = nbdays + 120
new_index = extend_index(dataConfirmedMorocco.index, predict_range)
size = len(new_index)
predictMorocco = solve_ivp(SEIR,
[0, size],
[s_0,e_0,i_0,r_0],
t_eval=np.arange(0, size, 1))
# +
infected_actual = np.concatenate((data.values, [None] * (size - len(data.values))))
recovered_actual = np.concatenate((dataRecoveredMorocco.values, [None] * (size - len(data.values))))
plt.figure(figsize=(20,10))
plt.plot(new_index[::2], infected_actual[::2], '*-',color='black', label='actual infected')
plt.plot(new_index[::2], recovered_actual[::2], '*-',color='magenta', label='actual recovered')
plt.plot(new_index[::2], predictMorocco.y[0,::2], label='Suspected')
plt.plot(new_index[::2], predictMorocco.y[1,::2], label='Exposed')
plt.plot(new_index[::2], predictMorocco.y[2,::2], label='Infected', color='red')
plt.plot(new_index[::2], predictMorocco.y[3,::2], label='Recovered', color='green')
plt.xticks(rotation=90)
plt.legend()
# -
# log scale
plt.figure(figsize=(20,10))
plt.semilogy(new_index[::2], infected_actual[::2], '*-',color='black', label='actual infected')
plt.semilogy(new_index[::2], recovered_actual[::2], '*-',color='magenta', label='actual recovered')
plt.semilogy(new_index[::2], predictMorocco.y[0,::2], label='Suspected')
plt.semilogy(new_index[::2], predictMorocco.y[1,::2], label='Exposed')
plt.semilogy(new_index[::2], predictMorocco.y[2,::2], label='Infected', color='red')
plt.semilogy(new_index[::2], predictMorocco.y[3,::2], label='Recovered', color='green')
plt.xticks(rotation=90)
plt.legend()
np.max(predictMorocco.y[2,:])
| notebooks/COVID-19-Morocco.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # xarray compatibility
#
# **scmdata** allows datat to be exported to xarray. This makes it easy to use xarray's many helpful features, most of which are not natively provided in scmdata.
# NBVAL_IGNORE_OUTPUT
import numpy as np
from scmdata import ScmRun
def get_data(years, n_ensemble_members, end_val, rand_pct):
return (np.arange(years.shape[0]) / years.shape[0] * end_val)[
:, np.newaxis
] * (rand_pct * np.random.random((years.shape[0], n_ensemble_members)) + 1)
# +
# NBVAL_IGNORE_OUTPUT
years = np.arange(1750, 2500 + 1)
variables = ["gsat", "gmst"]
n_variables = len(variables)
n_ensemble_members = 100
start = ScmRun(
np.hstack(
[
get_data(years, n_ensemble_members, 5.5, 0.1),
get_data(years, n_ensemble_members, 6.0, 0.05),
]
),
index=years,
columns={
"model": "a_model",
"scenario": "a_scenario",
"variable": [v for v in variables for i in range(n_ensemble_members)],
"region": "World",
"unit": "K",
"ensemble_member": [
i for v in variables for i in range(n_ensemble_members)
],
},
)
start
# -
# The usual scmdata methods are of course available.
# NBVAL_IGNORE_OUTPUT
start.plumeplot(
quantile_over="ensemble_member", hue_var="variable", hue_label="Variable"
)
# However, we can cast to an xarray DataSet and then all the xarray methods become available too.
# NBVAL_IGNORE_OUTPUT
xr_ds = start.to_xarray(dimensions=("ensemble_member",))
xr_ds
# For example, calculating statistics.
# NBVAL_IGNORE_OUTPUT
xr_ds.median(dim="ensemble_member")
# Plotting timeseries.
# NBVAL_IGNORE_OUTPUT
xr_ds["gsat"].plot.line(hue="ensemble_member", add_legend=False);
# Selecting and plotting timeseries.
# NBVAL_IGNORE_OUTPUT
xr_ds["gsat"].sel(ensemble_member=range(10)).plot.line(
hue="ensemble_member", add_legend=False
);
# Scatter plots.
# NBVAL_IGNORE_OUTPUT
xr_ds.plot.scatter(x="gsat", y="gmst", hue="ensemble_member", alpha=0.3)
# Or combinations of calculations and plots.
# NBVAL_IGNORE_OUTPUT
xr_ds.median(dim="ensemble_member").plot.scatter(x="gsat", y="gmst")
| notebooks/xarray-compatibility.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:svecachesim_ipykernel_conda_env]
# language: python
# name: conda-env-svecachesim_ipykernel_conda_env-py
# ---
# + language="markdown"
#
# # Sequential SuperLU Traces
#
# The traces share the following properties:
#
# * CFD Problems
# * 100% Pattern symmetry
# * <100% Numeric symmetry
# * Full rank
#
# This trace is from `EXAMPLE/ditersol`, which ships with SuperLU.
# We have slightly modified it to accept matrices as command line arguments,
# as opposed to reading them off of `stdin`.
#
#
# -
import sveCacheSim as sim
import CacheModels
import matplotlib.pyplot as plt
import numpy as nppace
import pandas as pd
import pickle
from tqdm import tqdm
import importlib
importlib.reload(sim)
pass
import os
# +
os.chdir('/storage/home/hhive1/plavin3/data/ModelSwapping')
slu = {}
## Choose a group of runs
#GROUP='slu'
GROUP='parsec'
traces_slu = ['steam1', 'steam2', 'orsirr_1', 'orsirr_2', 'orsreg_1']
traces_parsec = ['blackscholes', 'bodytrack', 'ferret', 'fluidanimate', 'freqmine']
models = ['BASE', 'FR', 'M4', 'M8', 'ALL']
if GROUP =='slu':
traces = traces_slu
elif GROUP == 'parsec':
traces = traces_parsec
else:
raise ValueError()
for tr in traces:
slu[tr] = {}
for mo in models:
if GROUP == 'slu':
filename = 'DataV4/slu_{}-{}-data.pkl'.format(tr, mo)
elif GROUP=='parsec':
filename = 'DataV4/{}-{}-data.pkl'.format(tr, mo)
try:
slu[tr][mo] = sim.load_object(filename)
except:
print('Failed to load {}'.format(filename))
# + language="markdown"
#
# ### Phase Trace Object
# We first need to grab the phase trace object from the stats of our simulation. It is stored in `stats_*.phase_trace`. This object
# has type `PhaseTrace` which is defined in `PhaseDetector.py` and includes 4 fields.
#
# * `trace`: The phase assigned to each interval. -1 represents an interval not identified as part of a phase.
# * `nphases`: The number of unique phases identified.
# * `phase_count`: A list of length nphases where phase_count[i] is the number of intervals assigned to phase i
# * `phase_unique`: A list of length nphases where phase_unique[i] is the number of times phase i was entered
#
# Let's take a look at these for the 'large' trace.
# +
def phase_summary(name, trace):
tr = trace.phase_trace
print(name, ':')
print(' Identified', tr.nphases, 'phases')
print(' Number of intervals assigned to each phase', tr.phase_count)
#print(' Number of times each phase was entered', tr.phase_unique)
for tr in slu.keys():
if 'BASE' in slu[tr].keys():
phase_summary(tr, slu[tr]['BASE'])
# +
def plot_trace(ax, label, trace, axis_labels):
ax.scatter([*range(len(trace))], trace,s=2, color='black')
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.set_title(label)
if axis_labels:
ax.set_ylabel('Phase Number (-1 is uncategorized)')
ax.set_xlabel('Interval number (Interval = 10k inst)')
fig, ax = plt.subplots(1, len(slu.keys()), figsize=(16,4))
for idx, tr in enumerate(slu.keys()):
if 'BASE' in slu[tr].keys():
plot_trace(ax[idx], tr, slu[tr]['BASE'].phase_trace.trace, idx == 0)
#plt.savefig('plots/phase-traces.svg')
plt.show()
# +
def get_acc(st1, st2, phase):
# t1 is assumed to be from the base cache
t1 = st1.cache_trace[0] # cache trace from st1
t2 = st2.cache_trace[0] # cache trace from st2
# Just get accesses where were are in phase `phase` and in state is Swapped (2)
pick_state = t2['state'] == 2
pick_phase = t2['phase'] == phase
pick = np.logical_and(pick_state, pick_phase)
# Get the hit/miss from each
t1_hits = t1['isHit'][pick]
t2_hits = t2['isHit'][pick]
num_correct = np.sum(np.equal(t1_hits, t2_hits))
acc = num_correct / len(t1_hits)
return acc
def get_stats(stats):
nphases = stats['BASE'].phase_trace.nphases
name_map={'FR':'Fixed Rate', 'M4': 'Markov 4', 'M8':'Markov 8', 'ALL':'All'}
data = {}
for name in stats:
if name is 'BASE':
continue
acc = []
for i in range(nphases):
acc.append(get_acc(stats['BASE'], stats[name], i))
data[name_map[name]] = acc
return pd.DataFrame(data)
acc = {}
for tr in slu.keys():
if 'BASE' in slu[tr].keys():
print('{} Accuracy'.format(tr))
acc[tr] = get_stats(slu[tr])
print(acc[tr])
print()
# +
nn=1
def acc_plot(ax, acc, slu, tr):
df = acc[tr]
ny = slu[tr]['BASE'].phase_trace.nphases
#ax=plt.gca()
df.plot(xticks=[*range(0,ny,1)],
yticks=[i/nn for i in range(0,nn+1)],
title='{} per-phase accuracy'.format(tr),ax=ax,
marker='o', linestyle=(0, (2,4)))
ax.set_ylabel('Accuracy')
ax.set_xlabel('Phase')
#ax.get_legend().remove()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
cmap = plt.get_cmap("tab10")
#ax.text( x=ny-1+.1, y=df['Fixed Rate'][ny-1], s='Fixed Rate', color=cmap(0))
#ax.text( x=4.1, y=df['Markov 4'][ny-1], s='Markov 4', color=cmap(1))
#ax.text( x=4.1, y=df['Markov 8'][ny-1], s='Markov 8', color=cmap(2))
plt.tight_layout()
fig, ax = plt.subplots(len(slu.keys()), 1, figsize=(8.5,10))
for idx, tr in enumerate(slu.keys()):
if 'BASE' in slu[tr].keys():
acc_plot(ax[idx], acc, slu, tr)
#acc_plot(acc, slu, 'steam2')
#plt.savefig('plots/per-phase-accuracy.svg')
# +
nn=1
maxidx=len(slu.keys())
barwidth=.2
colors = plt.cm.get_cmap('tab10').colors
colormap={'All': colors[0], 'Markov 4':colors[1], 'Markov 8':colors[2], 'Fixed Rate':colors[3]}
def acc_plot(ax, acc, slu, tr, idx):
df = acc[tr]
ny = slu[tr]['BASE'].phase_trace.nphases
#ax=plt.gca()
print(tr)
for idx,k in enumerate(acc[tr]):
bars=acc[tr][k]
r = np.arange(len(bars)) + barwidth*idx
ax.bar(r, bars, color=colormap[k], width=barwidth, edgecolor='white', label='var1')
#ax.xticks([r + barwidth for r in range(len(bars))], ['A', 'B', 'C', 'D'])
ax.set_yticks([0,1])
ax.set_xticks([*range(0,22,1)])
#df.plot(xticks=[*range(0,ny,1)],
# yticks=[i/nn for i in range(0,nn+1)],
# title='{}'.format(tr),ax=ax,
# marker='o', linestyle=(0, (2,4)))
#if idx == maxidx - 1:
# ax.set_ylabel('Accuracy')
# ax.set_xlabel('Phase')
#ax.get_legend().remove()
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
#cmap = plt.get_cmap("tab10")
#plt.tight_layout()
fig, ax = plt.subplots(len(slu.keys()), 1, figsize=(10,9))
for idx, tr in enumerate(slu.keys()):
if 'BASE' in slu[tr].keys():
acc_plot(ax[idx], acc, slu, tr, idx)
#acc_plot(acc, slu, 'steam2')
#plt.savefig('plots/per-phase-accuracy.svg')
# + language="markdown"
#
# ## Accuracy Over Time
# Let's now take a look at how accuracy changes over the course of a simulation. Does it get worse over time?
#
# As we're using the small trace, we won't see much here. Please re-run the notebook to get a better large chart.
#
# The reason a legend isn't generated is because it is hard to place it well. To get a legend, re-run the plot and uncomment the line labeled as such.
# Then grab the legend and paste it onto the plot in a good area.
# +
interval_len = 10000
def acc_over_time(st1, st2, phase):
t1 = st1.cache_trace[0] # cache trace from st1
t2 = st2.cache_trace[0] # cache trace from st2
# Just get accesses where were are in phase `phase` and in state is Swapped (2)
pick_state = t2['state'] == 2
pick_phase = t2['phase'] == phase
pick = np.logical_and(pick_state, pick_phase)
# Get the hit/miss from each
t1_hits = t1['isHit'][pick]
t2_hits = t2['isHit'][pick]
nintervals = int(len(t1_hits) // interval_len)-1
acc = []
for i in range(nintervals):
start = i*interval_len
end = (i+1)*interval_len
t1_hits_rest = t1_hits[start:end]
t2_hits_rest = t2_hits[start:end]
num_correct = np.sum(np.equal(t1_hits_rest, t2_hits_rest))
acc.append(num_correct / interval_len)
return acc
nphases = stats['base'].phase_trace.nphases
name_map={'fr':'Fixed Rate', 'm4': 'Markov 4', 'm8':'Markov 8', 'all':'All'}
all_df = {}
for i in range(nphases):
data = {}
for name in stats:
if name is 'base':
continue
data[name_map[name]] = acc_over_time(stats['base'], stats[name], i)
data_df = pd.DataFrame(data)
all_df[i] = data_df
# + jupyter={"source_hidden": true}
pretty=['Initialization', 'Marker', 'High Locality', 'Vector Add', 'Random']
def acc_over_time_plot(ax, approx, m4, m8, phase):
ax.plot(approx, label='Fixed Rate')
ax.plot(m4, label='Markov 4')
ax.plot(m8, label='Markov 8')
ax.set_ylim((0,1))
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.get_yaxis().set_ticks([])
ax.set_title('Phase {}'.format(phase))
ax.text(x=0, y=.05, s=pretty[phase], color='grey')
if phase == 0:
ax.set_ylabel('Accuracy')
ax.get_yaxis().set_ticks([.2, .4, .6, .8, 1.])
ax.set_xlabel('Interval number')
if phase == 4:
pass
#ax.legend(loc=10) ##UNCOMMENT ME FOR LEGEND
fig, ax = plt.subplots(1, 5, figsize=(12,3), squeeze=False)
for phase in range(5):
acc_over_time_plot(ax[0,phase],
all_df[phase]['Fixed Rate'],
all_df[phase]['Markov 4'],
all_df[phase]['Markov 8'],
phase)
#plt.savefig('plots/acc-over-time.svg')
plt.show()
# + language="markdown"
#
# ## Accuracy as a function of Model Size
#
# This is just a fun little plot.
# +
bs=8192
sz = {'base':1, 'fr':24/bs, 'm4':(3 * (4 * 4) * 8)/bs, 'm8':(3 * (8 * 8) * 8)/bs}
fig, ax = plt.subplots()
cmap = plt.get_cmap("tab10")
for i in range(5):
ax.scatter([sz['fr']], accuracy_df['Fixed Rate'][i], marker='${}$'.format(i), color=cmap(0), s=60)
for i in range(5):
ax.scatter([sz['m4']], accuracy_df['Markov 4'][i], marker='${}$'.format(i), color=cmap(1), s=60)
for i in range(5):
ax.scatter([sz['m8']], accuracy_df['Markov 8'][i], marker='${}$'.format(i), color=cmap(2), s=60)
ax.scatter(1,1, color=cmap(3))
ax.set_xscale('log')
labs=np.float64([24/bs, (3 * (4 * 4) * 8)/bs, (3 * (8 * 8) * 8)/bs, 1])*100
ax.get_xaxis().set_ticks([24/bs, (3 * (4 * 4) * 8)/bs, (3 * (8 * 8) * 8)/bs, 1])
ax.set_xticklabels(['{:.2f}%'.format(labs[i]) for i in range(4)])
ax.get_yaxis().set_ticks([.4,.6, .8, 1])
ax.set_xlabel('Model Size (% of Base Cache)')
ax.set_ylabel('Accuracy')
ax.set_ylim(0,1.05)
ax.set_xlim(0,1.5)
ax.text(x=labs[0]/100, y=.05, s='Fixed Rate', rotation=90, color='grey')
ax.text(x=labs[1]/100, y=.05, s='Markov 4', rotation=90, color='grey')
ax.text(x=labs[2]/100, y=.05, s='Markov 8', rotation=90, color='grey')
ax.text(x=labs[3]/100, y=.05, s='Base', rotation=90, color='grey')
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.set_title('Points are Phase Numbers', size=10)
fig.suptitle('Accuracy as a function of Model Size')
#plt.savefig('plots/acc_vs_modelsize.svg')
plt.show()
# + language="markdown"
#
# ## Locality Analysis
#
# Finally, we get to the hardest plot to generate, the locality plot.
# +
def index(array, item):
for idx, val in enumerate(array):
if val == item:
return idx
return -1
def rightshift(val: np.uint64, shift: np.uint64) -> np.uint64:
return val >> shift
def reuse(trace, shift, outfile=None):
stack = np.array([], dtype=np.int64)
out = []
for i in tqdm(range(len(trace))):
addr = rightshift(trace[i], np.uint64(shift)) # Cache line
idx = index(stack, addr)
if idx == -1:
stack = np.insert(stack, 0, addr)
else:
out.append(idx)
stack = np.delete(stack, idx)
stack = np.insert(stack, 0, addr)
if outfile is not None:
save_object(out, outfile)
print('Wrote reuse trace to: {}'.format(outfile))
return out
# +
reuse_perphase_base = {}
reuse_perphase_approx = {}
reuse_perphase_markov4 = {}
reuse_perphase_markov8 = {}
gran=6
def get_addrs(data, phase):
df = pd.DataFrame(data)
return np.array(df[df['phase']==phase]['addr'], dtype=np.uint64)
for phase in range(0,5):
reuse_perphase_base[phase] = reuse(get_addrs(stats['base'].cache_trace[1], phase), gran)
reuse_perphase_approx[phase] = reuse(get_addrs(stats['fr'].cache_trace[1], phase), gran)
reuse_perphase_markov4[phase] = reuse(get_addrs(stats['m4'].cache_trace[1], phase), gran)
reuse_perphase_markov8[phase] = reuse(get_addrs(stats['m8'].cache_trace[1], phase), gran)
# +
binwidth=16
#maxbin={1:50, 2:850, 3:225, 4:50}
maxbin={0:200,1:200, 2:200, 3:200, 4:200}
yl=45000
ylim={0:yl,1:yl, 2:yl, 3:yl, 4:yl}
def bins(data, binwidth, maxbin):
#return range(min(data), min(maxbin, max(data) + binwidth), binwidth)
# print(data)
return range(min(data), maxbin, binwidth)
def plot(ax, data, title, phase):
if not data:
data = [1]
ax.hist(data, bins=bins(data, binwidth, maxbin=maxbin[phase]), color='black')
ax.set_ylim((0,ylim[phase]))
#ax.suptitle(title)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.get_yaxis().set_ticks([])
if phase == 0:
ax.get_yaxis().set_ticks([30000])
ax.set_yticklabels(['30k'])
#plot(reuse_perphase_base[1], 'Base Cache', maxbin=maxbin[1])
#plot(reuse_perphase_approx[1], 'Fixed Rate Cache', maxbin=maxbin[1])
#plot(reuse_perphase_markov4[1], '4-State Markov Cache', maxbin=maxbin[1])
#plot(reuse_perphase_markov8[1], '8-State Markov Cache', maxbin=maxbin[1])
#i = 1
fig, ax = plt.subplots(4, 5, figsize=(8,8))
for i in range(5):
#phase = i+1
phase=i
plot(ax[0,i], reuse_perphase_base[phase], 'Base Cache', phase)
plot(ax[1,i], reuse_perphase_approx[phase], 'Fixed Rate Cache', phase)
plot(ax[2,i], reuse_perphase_markov4[phase], '4-State Markov Cache', phase)
plot(ax[3,i], reuse_perphase_markov8[phase], '8-State Markov Cache', phase)
#plt.savefig('plots/locality_per_phase.svg')
plt.show()
# -
| SuperLU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="https://rhyme.com/assets/img/logo-dark.png" align="center"> <h2 align="center">Logistic Regression: A Sentiment Analysis Case Study</h2>
#
#
# ### Introduction
# ___
# - IMDB movie reviews dataset
# - http://ai.stanford.edu/~amaas/data/sentiment
# - Contains 25000 positive and 25000 negative reviews
# <img src="https://i.imgur.com/lQNnqgi.png" align="center">
# - Contains at most reviews per movie
# - At least 7 stars out of 10 $\rightarrow$ positive (label = 1)
# - At most 4 stars out of 10 $\rightarrow$ negative (label = 0)
# - 50/50 train/test split
# - Evaluation accuracy
# <b>Features: bag of 1-grams with TF-IDF values</b>:
# - Extremely sparse feature matrix - close to 97% are zeros
# <b>Model: Logistic regression</b>
# - $p(y = 1|x) = \sigma(w^{T}x)$
# - Linear classification model
# - Can handle sparse data
# - Fast to train
# - Weights can be interpreted
# <img src="https://i.imgur.com/VieM41f.png" align="center" width=500 height=500>
# ### Task 1: Loading the dataset
# ---
# +
import pandas as pd
df = pd.read_csv('./movie_data.csv')
df.head(5)
# -
# ## <h2 align="center">Bag of words / Bag of N-grams model</h2>
# ### Task 2: Transforming documents into feature vectors
# ___
# Note: If you are starting the notebook from this task, you can run cells from all previous tasks in the kernel by going to the top menu and Kernel > Restart and Run All
# ___
# Below, we will call the fit_transform method on CountVectorizer. This will construct the vocabulary of the bag-of-words model and transform the following three sentences into sparse feature vectors:
# 1. The sun is shining
# 2. The weather is sweet
# 3. The sun is shining, the weather is sweet, and one and one is two
#
# Raw term frequencies: *tf (t,d)*—the number of times a term t occurs in a document *d*
# ### Task 3: Word relevancy using term frequency-inverse document frequency
# ___
# Note: If you are starting the notebook from this task, you can run cells from all previous tasks in the kernel by going to the top menu and Kernel > Restart and Run All
# ___
# $$\text{tf-idf}(t,d)=\text{tf (t,d)}\times \text{idf}(t,d)$$
# $$\text{idf}(t,d) = \text{log}\frac{n_d}{1+\text{df}(d, t)},$$
# where $n_d$ is the total number of documents, and df(d, t) is the number of documents d that contain the term t.
# The equations for the idf and tf-idf that are implemented in scikit-learn are:
#
# $$\text{idf} (t,d) = log\frac{1 + n_d}{1 + \text{df}(d, t)}$$
# The tf-idf equation that is implemented in scikit-learn is as follows:
#
# $$\text{tf-idf}(t,d) = \text{tf}(t,d) \times (\text{idf}(t,d)+1)$$
# $$v_{\text{norm}} = \frac{v}{||v||_2} = \frac{v}{\sqrt{v_{1}^{2} + v_{2}^{2} + \dots + v_{n}^{2}}} = \frac{v}{\big (\sum_{i=1}^{n} v_{i}^{2}\big)^\frac{1}{2}}$$
# ### Example:
# $$\text{idf}("is", d3) = log \frac{1+3}{1+3} = 0$$
# Now in order to calculate the tf-idf, we simply need to add 1 to the inverse document frequency and multiply it by the term frequency:
#
# $$\text{tf-idf}("is",d3)= 3 \times (0+1) = 3$$
# ### Task 4: Calculate tf-idf of the term *is*:
# ___
# Note: If you are starting the notebook from this task, you can run cells from all previous tasks in the kernel by going to the top menu and Kernel > Restart and Run All
# ___
#
# $$\text{tfi-df}_{norm} = \frac{[3.39, 3.0, 3.39, 1.29, 1.29, 1.29, 2.0 , 1.69, 1.29]}{\sqrt{[3.39^2, 3.0^2, 3.39^2, 1.29^2, 1.29^2, 1.29^2, 2.0^2 , 1.69^2, 1.29^2]}}$$$$=[0.5, 0.45, 0.5, 0.19, 0.19, 0.19, 0.3, 0.25, 0.19]$$$$\Rightarrow \text{tfi-df}_{norm}("is", d3) = 0.45$$
# ### Task 5:Data Preparation
# ___
# Note: If you are starting the notebook from this task, you can run cells from all previous tasks in the kernel by going to the top menu and Kernel > Restart and Run All
# ___
'''import re
def preprocessor(text):
text = re.sub('<[^>]*>', '', text)
emoticons = re.findall('(?::|;|=)(?:-)?(?:\)|\(|D|P)', text)
text = re.sub('[\W]+', ' ', text.lower()) +\
' '.join(emoticons).replace('-', '')
return text'''
#
#
# ### Task 6: Tokenization of documents
# ___
# Note: If you are starting the notebook from this task, you can run cells from all previous tasks in the kernel by going to the top menu and Kernel > Restart and Run All
# ___
#
#
# ### Task 7: Document classification via a logistic regression model
# ___
# Note: If you are starting the notebook from this task, you can run cells from all previous tasks in the kernel by going to the top menu and Kernel > Restart and Run All
# ___
# ### Task 8: Load saved model from disk
# ___
# Note: If you are starting the notebook from this task, you can run cells from all previous tasks in the kernel by going to the top menu and Kernel > Restart and Run All
# ___
#
# ### Task 9: Model accuracy
# ___
# Note: If you are starting the notebook from this task, you can run cells from all previous tasks in the kernel by going to the top menu and Kernel > Restart and Run All
# ___
| Logistic regression/task2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#pip install pycoingecko
# -
from pycoingecko import CoinGeckoAPI
import plotly.graph_objects as go
import pandas as pd
coinGecko=CoinGeckoAPI()
bitcoin_data=coinGecko.get_coin_market_chart_by_id(id='bitcoin',vs_currency='usd', days=60)
bitcoin_price_data=bitcoin_data['prices']
data=pd.DataFrame(bitcoin_price_data, columns=['TimeStamp','Price'])
data
data['Date'] = pd.to_datetime(data['TimeStamp'], unit='ms')
data
candlestick_data=data.groupby(data.Date.dt.date).agg({'Price':['min','max','first','last']})
fig=go.Figure(data=[go.Candlestick(x=candlestick_data.index,
open=candlestick_data['Price']['first'],
high=candlestick_data['Price']['max'],
low=candlestick_data['Price']['min'],
close=candlestick_data['Price']['last'],)])
fig.update_layout(xaxis_rangeslider_visible=False, xaxis_title='Date',
yaxis_title='Price (USD $)', title='Bitcoin Candlestick Chart over past 60 days')
| Cryptocurrency Price.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="VbWZg69t_IAy" colab_type="code" colab={}
# Execute this command to check GPU model
# In Colab the best one is Tesla P100
# !nvidia-smi
# + id="nCNglsAsvhpL" colab_type="code" colab={}
# Execute this to mount drive (if needed)
from google.colab import drive
drive.mount('/content/drive')
# + id="xrRXmGLEfOm_" colab_type="code" colab={}
import sys
sys.path.append('/content/drive/My Drive/DeepNILM')
import data_loading
import data_understanding
import data_model
import model_training
import model_testing
import data_ingestion
import metrics
# + id="Z7O772OS074C" colab_type="code" colab={}
# To Reload modules after updating them
import importlib
importlib.reload(data_loading)
importlib.reload(data_understanding)
importlib.reload(data_model)
importlib.reload(model_training)
importlib.reload(model_testing)
importlib.reload(data_ingestion)
importlib.reload(metrics)
# + [markdown] id="PWOskQg5fmTl" colab_type="text"
# The purpose of this notebook is to train a deep neural network to solve the following Energy Disaggregation task: infer the power consumption of two appliances (dishwasher and fridge) given the overall consumption (main) of an house.
#
# The followed approach is mainly inspired by the Sequence-to-point one described by Zhang et al. (2017) (https://arxiv.org/abs/1612.09106).
# + [markdown] id="KB-ATEzxgbjT" colab_type="text"
# ## Data Loading
# + id="CFyw2MZZg0VN" colab_type="code" colab={}
dishwasher_power = data_loading.read_csv_data("drive/My Drive/Progetto/dishwasher_train.csv")
fridge_power = data_loading.read_csv_data('drive/My Drive/Progetto/fridge_train.csv')
main_power = data_loading.read_csv_data('drive/My Drive/Progetto/main_train.csv')
# + [markdown] id="z9CiXHZKhEgk" colab_type="text"
# ## Data Understanding
#
# We make first an explorative analysis of the time series.
# + id="lgEY8i7wtpai" colab_type="code" colab={}
print('###################################################')
print('Data Understanding for dishwasher power consumption')
print('###################################################')
data_understanding.data_explore(dishwasher_power)
print('###################################################')
print('Data Understanding for fridge power consumption')
print('###################################################')
data_understanding.data_explore(fridge_power)
print('###################################################')
print('Data Understanding for main power consumption')
print('###################################################')
data_understanding.data_explore(main_power)
# + [markdown] id="1yq9F_6P1vjv" colab_type="text"
# Then, we visualize the time series in a given day.
# + id="XYJ8UXy14-AH" colab_type="code" colab={}
start_time_visualization = '2019-02-06 00:00:00'
end_time_visualization = '2019-02-07 00:00:00'
data_understanding.plot_interval(main_power, start_time_visualization,
end_time_visualization, title='Random Daily Overall Power Consumption')
data_understanding.plot_interval(dishwasher_power, start_time_visualization,
end_time_visualization, title='Random Daily Dishwasher Power Consumption')
data_understanding.plot_interval(fridge_power, start_time_visualization,
end_time_visualization, title='Fridge Daily Power Consumption')
# + [markdown] id="w38GtrFoSa6V" colab_type="text"
# ## Dishwasher's consumption inference
#
# Let's train the model for dishwasher with early stopping enabled.
# + id="88Hn0ZNHLGso" colab_type="code" colab={}
model_training.train_model(appliance_name='dishwasher',
main_path='drive/My Drive/Progetto/main_train.csv',
appliance_path='drive/My Drive/Progetto/dishwasher_train.csv',
train_end_timestamp='2019-03-01 00:00:00',
window_size=600,
batch_size=512,
build_model_func=data_model.build_dishwasher_cnn,
epochs=20,
patience=4,
early_stopping=True,
rescaling='normalize',
split=True,
plot_model=True)
# + [markdown] id="QHJ74yOG0dZw" colab_type="text"
# Now we train the model for a fixed number of epochs.
# + id="3f-30C-BXpxT" colab_type="code" colab={}
model = model_training.train_model(appliance_name='dishwasher',
main_path='drive/My Drive/Progetto/main_train.csv',
appliance_path='drive/My Drive/Progetto/dishwasher_train.csv',
window_size=600,
batch_size=512,
build_model_func=data_model.build_dishwasher_cnn,
epochs=8,
rescaling='normalize')
# + [markdown] id="wC7sQ0qa0iDl" colab_type="text"
# The model is saved for future reuse.
# + id="DqrR2lyHkzqQ" colab_type="code" colab={}
model.save('drive/My Drive/DeepNILM/dishwasher_model')
# + [markdown] id="gMrIz0SghmPz" colab_type="text"
# Let's try the test module.
# + id="f6h40UL02VIH" colab_type="code" colab={}
ground_truth, predicted_values = model_testing.test_model(appliance_name='dishwasher',
main_path='drive/My Drive/Progetto/main_train.csv',
appliance_path='drive/My Drive/Progetto/dishwasher_train.csv',
model_path='drive/My Drive/DeepNILM/dishwasher_model',
window_size=600,
batch_size=512,
rescaling='normalize',
appliance_min_power=0.0,
appliance_max_power=2570.6,
main_min_power=73.48100000000002,
main_max_power=6048.699999999999)
# + id="SxZvZeLgjlHj" colab_type="code" colab={}
f1 = metrics.compute_F1_score(predicted_values, ground_truth)
print('Energy based F1 score on test set: {}'.format(f1))
# + [markdown] id="6Ea996Hyxr_a" colab_type="text"
# ## Fridge's Consumption inference
#
# Let's train the model for fridge with early stopping enabled.
#
#
# + id="fzLy2z80Kqd_" colab_type="code" colab={}
model_training.train_model(appliance_name='fridge',
main_path='drive/My Drive/Progetto/main_train.csv',
appliance_path='drive/My Drive/Progetto/fridge_train.csv',
train_end_timestamp='2019-03-01 00:00:00',
window_size=600,
batch_size=512,
build_model_func=data_model.build_fridge_cnn,
epochs=50,
patience=10,
early_stopping=True,
rescaling='standardize',
split=True,
plot_model=True)
# + [markdown] id="RmB26DUNMFLU" colab_type="text"
# Now we train the model for a fixed number of epochs.
# + id="VwSHRGvzMMns" colab_type="code" colab={}
model = model_training.train_model(appliance_name='fridge',
main_path='drive/My Drive/Progetto/main_train.csv',
appliance_path='drive/My Drive/Progetto/fridge_train.csv',
window_size=600,
batch_size=512,
build_model_func=data_model.build_fridge_cnn,
epochs=7,
rescaling='standardize')
# + [markdown] id="a3HQlFh8MM3C" colab_type="text"
# The model is saved for future reuse.
# + id="DmxfAm6RMQFk" colab_type="code" colab={}
model.save('drive/My Drive/DeepNILM/fridge_model')
# + [markdown] id="j-I_YHK1hq6k" colab_type="text"
# Let's try the test module.
# + id="1pIrVjZ8YcOo" colab_type="code" colab={}
ground_truth, predicted_values = model_testing.test_model(appliance_name='fridge',
main_path='drive/My Drive/Progetto/main_train.csv',
appliance_path='drive/My Drive/Progetto/fridge_train.csv',
model_path='drive/My Drive/DeepNILM/fridge_model',
window_size=600,
batch_size=512,
rescaling='standardize',
appliance_mean_power=37.23710644724372,
appliance_std_power=46.9886959530205,
main_mean_power=370.91555422946004,
main_std_power=549.1880538356259)
# + id="eiEoEXpz4PdJ" colab_type="code" colab={}
f1 = metrics.compute_F1_score(predicted_values, ground_truth)
print('Energy based F1 score on test set: {}'.format(f1))
| TrainingNotebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Extraindo uma imagem PNG do GEE com o Pillow
#
# Neste notebook, utilizando o exemplo anterior de extração de máscaras, iremos exibir as imagens diretamente no notebook utilizando a biblioteca Pillow e o Request e, posteriormente, savá-la em disco.
#
# Primeiramente, vamos importar as bibliotecas e inicializar o GEE:
# +
# importação da bibliotecas
import ee
import PIL
import requests
from PIL import Image
from io import BytesIO
# inicialização do GEE
ee.Initialize()
# -
# Funções principais utilizadas por esse notebook (comentadas no notebook anterior):
# +
# Função para aplicar à imagem vinda da coleção a máscara de água
def mascara_agua(imagem):
qa = imagem.select('pixel_qa')
return qa.bitwiseAnd(1 << 2).eq(0)
# Função para aplicar à imagem vinda da coléção a máscara de nuvem/sombra de nuvem
def mascara_nuvem(imagem):
qa = imagem.select('pixel_qa')
return qa.bitwiseAnd(1 << 3).eq(0) and (qa.bitwiseAnd(1 << 5).eq(0)) and (qa.bitwiseAnd(1 << 6).eq(0)) and (qa.bitwiseAnd(1 << 7).eq(0))
# função para aplicar as máscaras
def aplicar_mascaras(imagem):
# criar uma imagem em branco/vazio para evitar problemas no fundo ao gerar um PNG
# usamos valores dummies (neste caso, branco)
vazio = ee.Image(99999)
# máscara de água
agua = vazio.updateMask(mascara_agua(imagem).Not()).rename('agua')
# máscara de nuvem (criará uma imagem com apenas nuvens)
# caso a imagem não tenha nuvens, ela ficará toda branca
nuvem = vazio.updateMask(mascara_nuvem(imagem).Not()).rename('nuvem')
# podemos ainda, ao contrário da linha anterior, REMOVER as nuvens
# notem que retiramos a função .Not (negação)
sem_nuvem = vazio.updateMask(mascara_nuvem(imagem)).rename('sem_nuvem')
# aplicar o indice NDVI
ndvi = imagem.expression('(nir - red) / (nir + red)',{'nir':imagem.select('B5'),'red':imagem.select('B4')}).rename('ndvi')
# assim como fizemos para o NDVI, retornamos uma imagem com as novas bandas
return imagem.addBands([ndvi,agua,nuvem,sem_nuvem])
# função para aplicar uma máscara em uma banda específica
# A mascará a ser aplicada
def aplicar_mascara_banda(imagem, banda_mascara, banda_origem, band_destino):
# Primeiramente, temos que aplicar a máscara desejada na banda de origem, que será nomeada para a banda de destino
# Podemos, inclusive, sobscrever a banda de origem, sem problemas
imagem_mascara = imagem.select(banda_origem).updateMask(imagem.select(banda_mascara)).rename(band_destino)
# Depois, temos que criar uma imagem em branco que receberá a máscara, renomeando também para banda de destino
imagem_mascara = ee.Image(99999).blend(imagem_mascara).rename(band_destino)
# Retornar a imagem com a nova banda nomeada com a string da banda_destino
return imagem.addBands([imagem_mascara])
# -
# Agora, vamos definir a geometria e as datas (baseada na Latitude e Longitude) da nossa área de estudo e consultá-la no GEE (mesmo do notebook anterior):
# +
# Notem que foi criada uma coordenada (Latitude e Longitude) através de uma string, posteriormente repartida pelas virgulas
# Essa abordagem é importante para quando utilizarmos a linha da comando
coordenadas = "-48.53801472648439,-22.503806214013736,-48.270222978437516,-22.7281869567509"
# Aqui, usamos uma ferramenta do Python chamada de unpacking
x1,y1,x2,y2 = coordenadas.split(",")
# Criamos a geometria com base nas coordenadas 'quebradas' acima
geometria = geometry = ee.Geometry.Polygon(
[[[float(x1),float(y2)],
[float(x2),float(y2)],
[float(x2),float(y1)],
[float(x1),float(y1)],
[float(x1),float(y2)]]])
# String de datas
datas = "2014-10-13,2014-10-14"
# Divisão das duas datas pela vírgula, novamente usando a técnica de unpacking
inicio,fim = datas.split(",")
# Consultando a coleção com base na área de estudo e datas selecionadas
colecao = ee.ImageCollection('LANDSAT/LC08/C01/T1_SR').filterBounds(geometria).filterDate(inicio,fim).filterMetadata('CLOUD_COVER','less_than', 30)
# aplicar a função 'aplicar_mascaras' em todas as imagens (irá adicionar as bandas 'agua', 'nuvem', 'sem_nuvem' nas imagens):
colecao = colecao.map(aplicar_mascaras)
# extraindo a imagem mediana da coleção
imagem = colecao.median()
# -
# Agora, vamos aplicar as máscaras individualmente na banda NDVI:
# +
# Aplicamos as três máscaras individualmente na banda NDVI
# A função irá adicionar as já mencionadas bandas de origem a medida que for sendo executada, linha a linha
imagem = aplicar_mascara_banda(imagem, 'agua', 'ndvi', 'ndvi_agua')
imagem = aplicar_mascara_banda(imagem, 'nuvem', 'ndvi', 'ndvi_nuvem')
imagem = aplicar_mascara_banda(imagem, 'sem_nuvem', 'ndvi', 'ndvi_sem_nuvem')
imagem = aplicar_mascara_banda(imagem, 'agua', 'ndvi_sem_nuvem', 'ndvi_agua_sem_nuvem')
# Depois, cortamos a imagem
# scale = escala do sensor. No caso do Landsat-8/OLI são 30 metros
imagem_corte = imagem.clipToBoundsAndScale(geometry=geometria,scale=30)
# -
# Utilizando o Pillow e o Request, iremos exibir extrair uma imagem da banda 'ndvi_agua_sem_nuvem' através da plataforma GEE:
# Exibindo a imagem com o Pillow, Requests e BytesIO diretamente no notebook
PIL.Image.open(BytesIO(requests.get(imagem_corte.select(['ndvi_agua_sem_nuvem']).getThumbUrl({'min':-1, 'max':1})).content))
# Essa imagem pode ser, inclusive, salva em um arquivo com o comando:
# Salvar imagem em um arquivo
imagem_pillow = PIL.Image.open(BytesIO(requests.get(imagem_corte.select(['ndvi_agua_sem_nuvem']).getThumbUrl({'min':-1, 'max':1})).content))
imagem_pillow.save('images/7-ndvi_bbhr.png')
# ### Verifiquem a imagem '7-ndvi_bbhr.png' na pasta 'images'
| .ipynb_checkpoints/7-Extraindo uma imagem PNG do GEE com o Pillow-checkpoint.ipynb |