code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
# +
# Reading Data
df_resale = pd.read_csv('Landed_Housing_sorted.csv')
df_resale['Address'] = df_resale['address']
folder = 'Non-Landed Sorted Data'
df_cc = pd.read_csv('./Landed Sorted Data/CC_NEW.csv')
df_cc = df_cc.rename(columns={'0' : 'Address','1' : 'CC','2' : 'distance_cc'})
df_cc['distance_cc'] = df_cc['distance_cc'].str[:-2].astype(float)
df_cc = df_cc.drop(['CC'], axis=1)
df_hawker = pd.read_csv('./Landed Sorted Data/hawker_NEW.csv')
df_hawker = df_hawker.rename(columns={'0' : 'Address','1' : 'hawker','2' : 'distance_hawker'})
df_hawker['distance_hawker'] = df_hawker['distance_hawker'].str[:-2].astype(float)
df_hawker = df_hawker.drop(['hawker'], axis=1)
df_mrt = pd.read_csv('./Landed Sorted Data/MRT_NEW.csv')
df_mrt = df_mrt.rename(columns={'0' : 'Address','1' : 'MRT','2' : 'distance_mrt'})
df_mrt['distance_mrt'] = df_mrt['distance_mrt'].str[:-2].astype(float)
df_mrt = df_mrt.drop(['MRT'], axis=1)
df_npc = pd.read_csv('./Landed Sorted Data/npc_NEW.csv')
df_npc = df_npc.rename(columns={'0' : 'Address','1' : 'NPC','2' : 'distance_npc'})
df_npc['distance_npc']= df_npc['distance_npc'].str[:-2].astype(float)
df_npc = df_npc.drop(['NPC'], axis=1)
df_ps = pd.read_csv('./Landed Sorted Data/ps_NEW.csv')
df_ps = df_ps.rename(columns={'0' : 'Address','1' : 'PS','2' : 'distance_primary_school'})
df_ps['distance_primary_school'] = df_ps['distance_primary_school'].str[:-2].astype(float)
df_ps = df_ps.drop(['PS'], axis=1)
df_ss = pd.read_csv('./Landed Sorted Data/SS_NEW.csv')
df_ss = df_ss.rename(columns={'0' : 'Address','1' : 'SS','2' : 'distance_secondary_school'})
df_ss['distance_secondary_school'] = df_ss['distance_secondary_school'].str[:-2].astype(float)
df_ss = df_ss.drop(['SS'], axis=1)
df_sm = pd.read_csv('./Landed Sorted Data/supermarket_NEW.csv')
df_sm = df_sm.rename(columns={'0' : 'Address','1' : 'SM','2' : 'distance_supermarket'})
df_sm['distance_supermarket'] = df_sm['distance_supermarket'].str[:-2].astype(float)
df_sm = df_sm.drop(['SM'], axis=1)
df_city = pd.read_csv('./Landed Sorted Data/City_NEW.csv')
df_city = df_city.rename(columns={'0' : 'Address','1' : 'City','2' : 'distance_city'})
df_city['distance_city'] = df_city['distance_city'].str[:-2].astype(float)
df_city = df_city.drop(['City'], axis=1)
# -
df_resale.head()
import re
df_resale['Tenure'].unique()
s = df_resale['Tenure'].str.findall('\d+')
def years_left(df):
a = re.findall('\d+',df)
if len(a)!= 0:
left = int(a[0]) - (2021 - int(a[1]))
else:
left = 999999
return left
df_resale['remaining_lease_yrs'] = df_resale['Tenure'].apply(years_left)
df_resale['remaining_lease_yrs']
# +
#The dictionary for cleaning up the categorical columns
cleanup_nums = {"flat_type_num": {"Detached": 1, "Semi-detached": 2,"Terrace": 3, "Strata Detached": 4
, "Strata Semi-detached": 5, "Strata Terrace": 6},
"Planning Area_num": {"North": 1, "North-East": 2,"East": 3, "West": 4,
"Central": 5},
}
#To convert the columns to numbers using replace:
df_resale['flat_type_num'] = df_resale['Type']
df_resale['Planning Area_num'] = df_resale['Planning Area']
df_resale = df_resale.replace(cleanup_nums)
df_resale.head()
# -
merged = pd.merge(df_resale,df_cc, on=['Address'], how="outer")
merged = pd.merge(merged,df_hawker, on=['Address'], how="outer")
merged = pd.merge(merged,df_mrt, on=['Address'], how="outer")
merged = pd.merge(merged,df_npc, on=['Address'], how="outer")
merged = pd.merge(merged,df_ps, on=['Address'], how="outer")
merged = pd.merge(merged,df_ss, on=['Address'], how="outer")
merged = pd.merge(merged,df_sm, on=['Address'], how="outer")
merged = pd.merge(merged,df_city, on=['Address'], how="outer")
#merged = pd.merge(merged,df_meta, on=['Address'], how="outer")
merged = merged.dropna()
#merged.to_csv('Complete_dataset_landed.csv')
merged = merged.rename(columns={"Area (Sqft)": "floor_area_sqm"})
merged['resale_price'] = merged['Price ($)']/merged['No. of Units']
# +
dataset_features = merged[['resale_price', 'Postal District','flat_type_num' ,'floor_area_sqm', 'Planning Area_num', 'remaining_lease_yrs',
'distance_secondary_school','distance_primary_school', 'distance_mrt', 'distance_supermarket', 'distance_hawker',
'distance_city', 'distance_npc', 'distance_cc','Mature_Estate']]
print(len(dataset_features))
# Only "y varable"
resale_p = dataset_features['resale_price']
# All other indepdendent variables
X = dataset_features[[ 'Postal District','flat_type_num' ,'floor_area_sqm', 'Planning Area_num', 'remaining_lease_yrs',
'distance_secondary_school','distance_primary_school', 'distance_mrt', 'distance_supermarket', 'distance_hawker',
'distance_city', 'distance_npc', 'distance_cc','Mature_Estate']]
dataset_features.head()
# -
dataset_features.dtypes
# +
#Have correlation analysis for resale price with all variables:
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import numpy as np
corrMatrix = dataset_features.corr()
test = corrMatrix.iloc[[0]]
test = test.iloc[: , 1:]
print(test)
"""plt.subplots(figsize=(12,9))
sns.heatmap(corrMatrix, xticklabels=corrMatrix.columns, yticklabels=corrMatrix.columns, cmap='coolwarm', annot=True)"""
# -
plt.subplots(figsize=(13,0.5))
sns.heatmap(test, xticklabels=test.columns, yticklabels=test.index, cmap='coolwarm', annot=True)
plt.yticks(rotation = 'horizontal')
# +
floor_area = merged.sort_values(by=['floor_area_sqm'])
plt.fill_between(floor_area['floor_area_sqm'], floor_area['resale_price'], lw=2)
plt.ylabel('Resale Price')
plt.title('Landed Housing Floor Area')
plt.show()
# +
remaining = merged.sort_values(by=['remaining_lease_yrs'])
remaining = remaining.loc[remaining['remaining_lease_yrs']>30]
remaining = remaining.loc[remaining['resale_price']<10000000]
y_pos = np.arange(len(remaining))
plt.bar(y_pos, remaining['resale_price'], align='center', alpha=0.5,width=20)
plt.ylabel('Resale Price')
plt.xticks(y_pos, remaining['remaining_lease_yrs'])
plt.title('Landed Housing Remaining Lease Years')
plt.show()
# -
remaining = merged.sort_values(by=['remaining_lease_yrs'])
plt.ylabel('Resale Price')
plt.scatter(remaining['remaining_lease_yrs'], remaining['resale_price'],s=1, alpha=0.5)
plt.title('Landed Housing Remaining Lease Years')
plt.show()
# +
city = merged.sort_values(by=['distance_city'])
plt.plot(city['distance_city'], city['resale_price'], lw=2)
plt.ylabel('Resale Price')
plt.title('Landed Housing Distance to City')
plt.xlabel('Distance to City')
plt.show()
# -
postal_district = merged.groupby(by = ['Postal District'])['resale_price'].mean().reset_index()
y_pos = np.arange(len(postal_district))
plt.bar(y_pos, postal_district['resale_price'], align='center', alpha=0.5)
plt.ylabel('Resale Price')
plt.xticks(y_pos, postal_district['Postal District'])
plt.title('Landed Housing Postal District')
plt.show()
# +
from sklearn.model_selection import train_test_split
from sklearn import linear_model
from sklearn.metrics import mean_squared_error, r2_score
import numpy as np
from sklearn import tree
from sklearn.tree import DecisionTreeRegressor
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import AdaBoostRegressor, BaggingRegressor
np.random.seed(100)
X_train, X_test, y_train, y_test = train_test_split(X, resale_p, test_size=.3, random_state=0)
Models = ["OLS", "AdaBoost", "Decision Tree"]
MSE_lst = []
# OLS:
regr = linear_model.LinearRegression() # Create linear regression object
np.random.seed(100)
regr.fit(X_train, y_train) # Train the model using the training sets
y_pred_ols = regr.predict(X_test) # Make predictions using the testing set
MSE_ols = mean_squared_error(y_test, y_pred_ols) # performance statistic
MSE_lst.append(MSE_ols)
# Boosting
adaboosting = AdaBoostRegressor()
adaboosting.fit(X=X_train, y=y_train)
y_pred_boosting = adaboosting.predict(X=X_test)
MSE_adaboost = mean_squared_error(y_test, y_pred_boosting)
MSE_lst.append(MSE_adaboost)
# Bagging
bagging = BaggingRegressor(DecisionTreeRegressor())
bagging.fit(X=X_train, y=y_train)
y_pred_dt = bagging.predict(X=X_test)
MSE_bag = mean_squared_error(y_test, y_pred_dt)
MSE_lst.append(MSE_bag)
# -
import matplotlib.pyplot as plt
print(len(y_test))
fig, ax = plt.subplots(figsize=(20,10))
ax.plot(range(len(y_test[:100])), y_test[:100], '-b',label='Actual')
ax.plot(range(len(y_pred_ols[:100])), y_pred_ols[:100], 'r', label='Predicted')
plt.show()
import matplotlib.pyplot as plt
print(len(y_test))
fig, ax = plt.subplots(figsize=(20,10))
ax.plot(range(len(y_test[:100])), y_test[:100], '-b',label='Actual')
ax.plot(range(len(y_pred_boosting[:100])), y_pred_boosting[:100], 'r', label='Predicted')
plt.show()
import matplotlib.pyplot as plt
print(len(y_test))
fig, ax = plt.subplots(figsize=(20,10))
ax.plot(range(len(y_test[:100])), y_test[:100], '-b',label='Actual')
ax.plot(range(len(y_pred_dt[:100])), y_pred_dt[:100], 'r', label='Predicted')
plt.show()
# +
y_pred_df_ols = pd.DataFrame(y_pred_dt, columns= ['y_pred'])
print(len(y_pred_df_ols))
print(len(X_test))
pred_res1 = pd.concat([X_test,y_test], axis=1)
print(len(pred_res1))
pred_res1 = pred_res1.reset_index(drop=True)
pred_res2 = pd.concat([pred_res1,y_pred_df_ols], axis=1)
pred_res2
# +
import pandas as pd
import numpy as np
from sklearn import datasets, linear_model
from sklearn.linear_model import LinearRegression
import statsmodels.api as sm
from scipy import stats
X2 = sm.add_constant(X_train)
est = sm.OLS(y_train, X2)
est2 = est.fit()
print(est2.summary())
# -
| [CODE] Data Exploration/Data Exploration Landed Housing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # XML Parser and Dataframe Creator
#
# The first step is to parse XML files into documents.
#
#
# Useful Resources:
# * <NAME>, "[Processing XML in Python—ElementTree](https://towardsdatascience.com/processing-xml-in-python-elementtree-c8992941efd2)," Accessed Sept. 22, 2020.
# +
# Import necessary libraries.
import re, glob, csv, sys, os
import pandas as pd
import xml.etree.ElementTree as ET
# Declare directory location to shorten filepaths later.
abs_dir = "/Users/quinn.wi/Documents/"
# Gather all .xml files using glob.
list_of_files = glob.glob(abs_dir + "Data/PSC/JQA/*/*.xml")
# -
# ## Define Functions
# +
'''
Arguments of Functions:
namespace:
ancestor:
xpath_as_string:
attrib_val_str:
'''
# Read in file and get root of XML tree.
def get_root(xml_file):
tree = ET.parse(xml_file)
root = tree.getroot()
return root
# Get namespace of individual file from root element.
def get_namespace(root):
namespace = re.match(r"{(.*)}", str(root.tag))
ns = {"ns":namespace.group(1)}
return ns
# Get document id.
def get_document_id(ancestor, attrib_val_str):
doc_id = ancestor.get(attrib_val_str)
return doc_id
# Get date of document.
def get_date_from_attrValue(ancestor, xpath_as_string, attrib_val_str, namespace):
date = ancestor.find(xpath_as_string, namespace).get(attrib_val_str)
return date
def get_peopleList_from_attrValue(ancestor, xpath_as_string, attrib_val_str, namespace):
people_list = []
for elem in ancestor.findall(xpath_as_string, namespace):
person = elem.get(attrib_val_str)
people_list.append(person)
# Return a string object of 'list' to be written to output file. Can be split later.
return ','.join(people_list)
# Get plain text of every element (designated by first argument).
def get_textContent(ancestor, xpath_as_string, namespace):
text_list = []
for elem in ancestor.findall(xpath_as_string, namespace):
text = ''.join(ET.tostring(elem, encoding='unicode', method='text'))
# Add text (cleaned of additional whitespace) to text_list.
text_list.append(re.sub(r'\s+', ' ', text))
# Return concetanate text list.
return ' '.join(text_list)
# -
# ## Declare Variables
# +
# Declare regex to simplify file paths below
regex = re.compile(r'.*/\d{4}/(.*)')
# Declare document level of file. Requires root starting point ('.').
doc_as_xpath = './/ns:div/[@type="entry"]'
# Declare date element of each document.
date_path = './ns:bibl/ns:date/[@when]'
# Declare person elements in each document.
person_path = './/ns:p/ns:persRef/[@ref]'
# Declare text level within each document.
text_path = './ns:div/[@type="docbody"]/ns:p'
# -
# ## Parse Documents
# +
# %%time
# Open/Create file to write contents.
with open(abs_dir + 'Output/ParsedXML/JQA_dataframe.txt', 'w') as outFile:
# Write headers for table.
outFile.write('file' + '\t' + 'entry' + '\t' + 'date' + '\t' + \
'people' + '\t' + 'text' + '\n')
# Loop through each file within a directory.
for file in list_of_files:
# Call functions to create necessary variables and grab content.
root = get_root(file)
ns = get_namespace(root)
for eachDoc in root.findall(doc_as_xpath, ns):
# Call functions.
entry = get_document_id(eachDoc, '{http://www.w3.org/XML/1998/namespace}id')
date = get_date_from_attrValue(eachDoc, date_path, 'when', ns)
people = get_peopleList_from_attrValue(eachDoc, person_path, 'ref', ns)
text = get_textContent(eachDoc, text_path, ns)
# Write results in tab-separated format.
outFile.write(str(regex.search(file).groups()) + '\t' + entry + \
'\t' + date + '\t' + people + '\t' + text + '\n')
# -
# ## Import Dataframe
# +
dataframe = pd.read_csv(abs_dir + 'Output/ParsedXML/JQA_dataframe.txt', sep = '\t')
dataframe
# -
| Jupyter_Notebooks/Parsers/JQA_XML-Parser.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:c3dev] *
# language: python
# name: conda-env-c3dev-py
# ---
# # Reconstructing ancestral states
#
# This app takes a `model_result` and returns a `tabular_result` consisting of the posterior probabilities of ancestral states for each node of a tree. These probabilities are computed using the marginal reconstruction algorithm.
#
# We first fit a model to the sample data.
# +
from cogent3.app import io, evo
reader = io.load_aligned(format="fasta")
aln = reader("../data/primate_brca1.fasta")
gn = evo.model("GN", tree="../data/primate_brca1.tree")
result = gn(aln)
# -
# ## Define the `ancestral_states` app
reconstuctor = evo.ancestral_states()
states_result = reconstuctor(result)
states_result
# The `tabular_result` is keyed by the node name. Each value is a `DictArray`, with header corresponding to the states and rows corresponding to alignment position.
states_result['edge.0']
# If not included in the newick tree file, the internal node names are automatically generated open loading. You can establish what those are by interrogating the tree bound to the likelihood function object. (If you move your mouse cursor over the nodes, their names will appear as hover text.)
result.tree.get_figure(contemporaneous=True).show(width=500, height=500)
| doc/app/evo-ancestral-states.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] deletable=true editable=true
# # Neural Network Basics
#
# Compiled by <NAME> (@sunilmallya)
# + [markdown] deletable=true editable=true
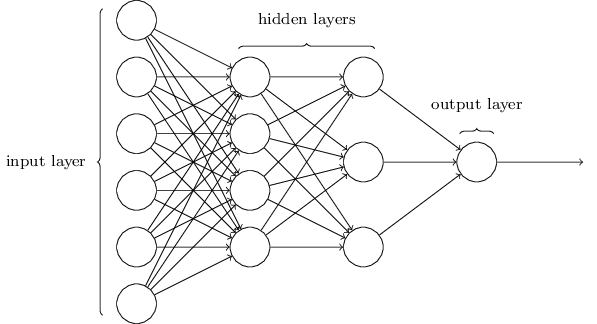
# # Basic Neural Network layout
#
# 
#
#
# + [markdown] deletable=true editable=true
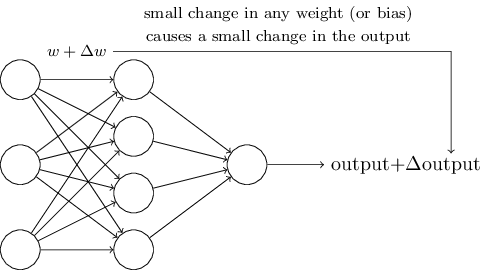
# 
#
#
# images from: http://neuralnetworksanddeeplearning.com/chap1.html
# -
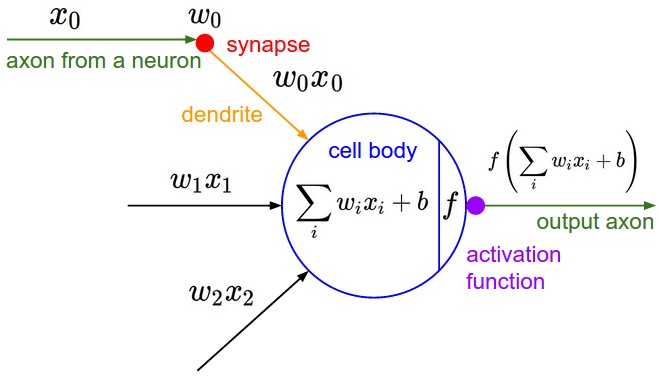
# ## Neural Network Equation
#
# 
#
# img src http://cs231n.github.io/assets/nn1/neuron_model.jpeg
# ## The Learning Process
#
# 
# + [markdown] deletable=true editable=true
# # Loss Functions and Optimizations
#
#
# 
#
# + [markdown] deletable=true editable=true
# # Layers
#
#
# ## Multilayer perceptrons
#
# Here's where things start to get interesting. Before, we mapped our inputs directly onto our outputs through a single linear transformation.
#
# 
#
# This model is perfectly adequate when the underlying relationship between our data points and labels is approximately linear. When our data points and targets are characterized by a more complex relationship, a linear model and produce sucky results. We can model a more general class of functions by incorporating one or more *hidden layers*.
#
# 
#
# Here, each layer will require it's own set of parameters. To make things simple here, we'll assume two hidden layers of computation.
# -
# ### Word Embeddings
#
# Word Embedding turns text into numbers
#
# - ML Algorithms don't understand text, they require input to be vectors of continuous values
#
# - Benefits:
# - Dimensionality Reduction
# - Group similar words together (Contextual Similarity)
#
#
# ##### A common Embedding technique: Word2Vec
#
# https://github.com/saurabh3949/Word2Vec-MXNet/blob/master/Word2vec%2Bwith%2BGluon.ipynb
#
# 
#
# ##### [Embedding Visualization]
# https://ronxin.github.io/wevi/
#
# https://medium.com/towards-data-science/deep-learning-4-embedding-layers-f9a02d55ac12
# + [markdown] deletable=true editable=true
# # MXNet cheat sheet
#
#
# PDF
# https://s3.amazonaws.com/aws-bigdata-blog/artifacts/apache_mxnet/apache-mxnet-cheat.pdf
#
# https://aws.amazon.com/blogs/ai/exploiting-the-unique-features-of-the-apache-mxnet-deep-learning-framework-with-a-cheat-sheet/
# + deletable=true editable=true
| nn_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Confidence and prediction intervals
#
# > <NAME>
# > Laboratory of Biomechanics and Motor Control ([http://demotu.org/](http://demotu.org/))
# > Federal University of ABC, Brazil
# For a finite univariate random variable with a normal probability distribution, the mean $\mu$ (a measure of central tendency) and variance $\sigma^2$ (a measure of dispersion) of a population are the well known formulas:
#
# $$ \mu = \frac{1}{N}\sum_{i=1}^{N} x_i $$
#
# $$ \sigma^2 = \frac{1}{N}\sum_{i=1}^{N} (x_i - \mu)^2 $$
#
# For a more general case, a continuous univariate random variable $x$ with [probability density function](http://en.wikipedia.org/wiki/Probability_density_function) (pdf), $f(x)$, the mean and variance of a population are:
#
# $$ \mu = \int_{\infty}^{\infty} x f(x)\: dx $$
#
# $$ \sigma^2 = \int_{\infty}^{\infty} (x-\mu)^2 f(x)\: dx $$
#
# The pdf is a function that describes the relative likelihood for the random variable to take on a given value.
# Mean and variance are the first and second central moments of a random variable. The standard deviation $\sigma$ of the population is the square root of the variance.
#
# The [normal (or Gaussian) distribution](http://en.wikipedia.org/wiki/Normal_distribution) is a very common and useful distribution, also because of the [central limit theorem](http://en.wikipedia.org/wiki/Central_limit_theorem), which states that for a sufficiently large number of samples (each with many observations) of an independent random variable with an arbitrary probability distribution, the means of the samples will have a normal distribution. That is, even if the underlying probability distribution of a random variable is not normal, if we sample enough this variable, the means of the set of samples will have a normal distribution.
#
# The probability density function of a univariate normal (or Gaussian) distribution is:
#
# $$ f(x) = \frac{1}{\sqrt{2\pi\sigma^2}} \exp\Bigl(-\frac{(x-\mu)^2}{2\sigma^2}\Bigr) $$
#
# The only parameters that define the normal distribution are the mean $\mu$ and the variance $\sigma^2$, because of that a normal distribution is usually described as $N(\mu,\:\sigma^2)$.
#
# Here is a plot of the pdf for the normal distribution:
# import the necessary libraries
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython.display import display, Latex
from scipy import stats
import sys
sys.path.insert(1, r'./../functions') # directory of BMC Python functions
from pdf_norm_plot import pdf_norm_plot
pdf_norm_plot();
# The horizontal axis above is shown in terms of the number of standard deviations in relation to the mean, which is known as standard score or $z$ score:
#
# $$ z = \frac{x - \mu}{\sigma} $$
#
# So, instead of specifying raw values in the distribution, we define the pdf in terms of $z$ scores; this conversion process is called standardizing the distribution (and the result is known as standard normal distribution). Note that because $\mu$ and $\sigma$ are known parameters, $z$ has the same distribution as $x$, in this case, the normal distribution.
#
# The percentage numbers in the plot are the probability (area under the curve) for each interval shown in the horizontal label.
# An interval in terms of z score is specified as: $[\mu-z\sigma,\;\mu+z\sigma]$.
# The interval $[\mu-1\sigma,\;\mu+1\sigma]$ contains 68.3% of the population and the interval $[\mu-2\sigma,\;\mu+2\sigma]$ contains 95.4% of the population.
# These numbers can be calculated using the function `stats.norm.cdf()`, the [cumulative distribution function](http://en.wikipedia.org/wiki/Cumulative_distribution_function) (cdf) of the normal distribution at a given value:
print('Cumulative distribution function (cdf) of the normal distribution:')
for i in range(-3, 4):
display(Latex(r'%d $\sigma:\;$ %.2f' %(i, stats.norm.cdf(i, loc=0, scale=1)*100) + ' %'))
# The parameters `loc` and `scale` are optionals and represent mean and variance of the distribution. The default is `loc=0` and `scale=1`.
# A commonly used proportion is 95%. The value that results is this proportion can be found using the function `stats.norm.ppf()`. If we want to find the $\pm$value for the interval that will result in 95% of the population inside, we have to consider that 2.5% of the population will stay out of the interval in each tail of the distribution. Because of that, the number we have to use with the `stats.norm.ppf()` is 0.975:
print('Percent point function (inverse of cdf) of the normal distribution:')
display(Latex(r'ppf(.975) = %.2f' % stats.norm.ppf(.975, loc=0, scale=1)))
# Or we can use the function `stats.norm.interval` which already gives the interval:
print('Confidence interval around the mean:')
stats.norm.interval(alpha=0.95, loc=0, scale=1)
# So, the interval $[\mu-1.96\sigma,\;\mu+1.96\sigma]$ contains 95% of the population.
#
# Now that we know how the probability density function of a normal distribution looks like, let's demonstrate the central limit theorem for a uniform distribution. For that, we will generate samples of a uniform distribution, calculate the mean across samples, and plot the histogram of the mean across samples:
fig, ax = plt.subplots(1, 4, sharey=True, squeeze=True, figsize=(12, 5))
x = np.linspace(0, 1, 100)
for i, n in enumerate([1, 2, 3, 10]):
f = np.mean(np.random.random((1000, n)), 1)
m, s = np.mean(f), np.std(f, ddof=1)
fn = (1/(s*np.sqrt(2*np.pi)))*np.exp(-(x-m)**2/(2*s**2)) # normal pdf
ax[i].hist(f, 20, normed=True, color=[0, 0.2, .8, .6])
ax[i].set_title('n=%d' %n)
ax[i].plot(x, fn, color=[1, 0, 0, .6], linewidth=5)
plt.suptitle('Demonstration of the central limit theorem for a uniform distribution', y=1.05)
plt.show()
# **Statistics for a sample of the population**
#
# Parameters (such as mean and variance) are characteristics of a population; statistics are the equivalent for a sample. For a population and a sample with normal or Gaussian distribution, mean and variance is everything we need to completely characterize this population or sample.
#
# The difference between sample and population is fundamental for the understanding of probability and statistics.
# In Statistics, a sample is a set of data collected from a population. A population is usually very large and can't be accessed completely; all we have access is a sample (a smaller set) of the population.
#
# If we have only a sample of a finite univariate random variable with a normal distribution, both mean and variance of the population are unknown and they have to be estimated from the sample:
#
# $$ \bar{x} = \frac{1}{N}\sum_{i=1}^{N} x_i $$
#
# $$ s^2 = \frac{1}{N-1}\sum_{i=1}^{N} (x_i - \bar{x})^2 $$
#
# The sample $\bar{x}$ and $s^2$ are only estimations of the unknown true mean and variance of the population, but because of the [law of large numbers](http://en.wikipedia.org/wiki/Law_of_large_numbers), as the size of the sample increases, the sample mean and variance have an increased probability of being close to the population mean and variance.
#
# **Prediction interval around the mean**
#
# For a sample of a univariate random variable, the area in an interval of the probability density function can't be interpreted anymore as the proportion of the sample lying inside the interval. Rather, that area in the interval is a prediction of the probability that a new value from the population added to the sample will be inside the interval. This is called a [prediction interval](http://en.wikipedia.org/wiki/Prediction_interval). However, there is one more thing to correct. We have to adjust the interval limits for the fact that now we have only a sample of the population and the parameters $\mu$ and $\sigma$ are unknown and have to be estimated. This correction will increase the interval for the same probability value of the interval because we are not so certain about the distribution of the population.
# To calculate the interval given a desired probability, we have to determine the distribution of the z-score equivalent for the case of a sample with unknown mean and variance:
#
# $$ \frac{x_{n+i}-\bar{x}}{s\sqrt{1+1/n}} $$
#
# Where $x_{n+i}$ is the new observation for which we want to calculate the prediction interval.
# The distribution of the ratio above is called <a href="http://en.wikipedia.org/wiki/Student's_t-distribution">Student's t-distribution</a> or simply $T$ distribution, with $n-1$ degrees of freedom. A $T$ distribution is symmetric and its pdf tends to that of the
# standard normal as $n$ tends to infinity.
#
# Then, the prediction interval around the sample mean for a new observation is:
#
# $$ \left[\bar{x} - T_{n-1}\:s\:\sqrt{1+1/n},\quad \bar{x} + T_{n-1}\:s\:\sqrt{1+1/n}\right] $$
#
# Where $T_{n-1}$ is the $100((1+p)/2)^{th}$ percentile of the Student's t-distribution with n−1 degrees of freedom.
#
# For instance, the prediction interval with 95% of probability for a sample ($\bar{x}=0,\;s^2=1$) with size equals to 10 is:
np.asarray(stats.t.interval(alpha=0.95, df=25-1, loc=0, scale=1)) * np.sqrt(1+1/10)
# For a large sample (e.g., 10000), the interval approaches the one for a normal distribution (according to the [central limit theorem](http://en.wikipedia.org/wiki/Central_limit_theorem)):
np.asarray(stats.t.interval(alpha=0.95, df=10000-1, loc=0, scale=1)) * np.sqrt(1+1/10000)
# Here is a plot of the pdf for the normal distribution and the pdf for the Student's t-distribution with different number of degrees of freedom (n-1):
fig, ax = plt.subplots(1, 1, figsize=(10, 5))
x = np.linspace(-4, 4, 1000)
f = stats.norm.pdf(x, loc=0, scale=1)
t2 = stats.t.pdf(x, df=2-1)
t10 = stats.t.pdf(x, df=10-1)
t100 = stats.t.pdf(x, df=100-1)
ax.plot(x, f, color='k', linestyle='--', lw=4, label='Normal')
ax.plot(x, t2, color='r', lw=2, label='T (1)')
ax.plot(x, t10, color='g', lw=2, label='T (9)')
ax.plot(x, t100, color='b', lw=2, label='T (99)')
ax.legend(title='Distribution', fontsize=14)
ax.set_title("Normal and Student's t distributions", fontsize=20)
ax.set_xticks(np.linspace(-4, 4, 9))
xtl = [r'%+d$\sigma$' %i for i in range(-4, 5, 1)]
xtl[4] = r'$\mu$'
ax.set_xticklabels(xtl)
ax.set_ylim(-0.01, .41)
plt.grid()
plt.rc('font', size=16)
plt.show()
# It's common to use 1.96 as value for the 95% prediction interval even when dealing with a sample; let's quantify the error of this approximation for different sample sizes:
T = lambda n: stats.t.ppf(0.975, n-1)*np.sqrt(1+1/n) # T distribution
N = stats.norm.ppf(0.975) # Normal distribution
for n in [1000, 100, 10]:
print('\nApproximation error for n = %d' %n)
print('Using Normal distribution: %.1f%%' % (100*(N-T(n))/T(n)))
# For n=1000, the approximation is good, for n=10 it is bad, and it always underestimates.
#
# **Standard error of the mean**
#
# The [standard error of the mean](http://en.wikipedia.org/wiki/Standard_error) (sem) is the standard deviation of the sample-mean estimate of a population mean and is given by:
#
# $$ sem = \frac{s}{\sqrt{n}} $$
#
# **Confidence interval**
#
# In statistics, a [confidence interval](http://en.wikipedia.org/wiki/Confidence_interval) (CI) is a type of interval estimate of a population parameter and is used to indicate the reliability of an estimate ([Wikipedia](http://en.wikipedia.org/wiki/Confidence_interval)). For instance, the 95% confidence interval for the sample-mean estimate of a population mean is:
#
# $$ \left[\bar{x} - T_{n-1}\:s/\sqrt{n},\quad \bar{x} + T_{n-1}\:s/\sqrt{n}\right] $$
#
# Where $T_{n-1}$ is the $100((1+p)/2)^{th}$ percentile of the Student's t-distribution with n−1 degrees of freedom.
# For instance, the confidence interval for the mean with 95% of probability for a sample ($\bar{x}=0,\;s^2=1$) with size equals to 10 is:
stats.t.interval(alpha=0.95, df=10-1, loc=0, scale=1) / np.sqrt(10)
# The 95% CI means that if we randomly obtain 100 samples of a population and calculate the CI of each sample (i.e., we replicate the experiment 99 times in a independent way), 95% of these CIs should contain the population mean (the true mean). This is different from the prediction interval, which is larger, and gives the probability that a new observation is inside this interval. Note that the confidence interval DOES NOT give the probability that the true mean (the mean of the population) is inside this interval. The true mean is a parameter (fixed) and it is either inside the calculated interval or not; it is not a matter of chance (probability).
#
# Let's simulate samples of a population ~ $N(\mu=0, \sigma^2=1) $ and calculate the confidence interval for the samples' mean:
n = 20 # number of observations
x = np.random.randn(n, 100) # 100 samples with n observations
m = np.mean(x, axis=0) # samples' mean
s = np.std(x, axis=0, ddof=1) # samples' standard deviation
T = stats.t.ppf(.975, n-1) # T statistic for 95% and n-1 degrees of freedom
ci = m + np.array([-s*T/np.sqrt(n), s*T/np.sqrt(n)])
out = ci[0, :]*ci[1, :] > 0 # CIs that don't contain the true mean
fig, ax = plt.subplots(1, 1, figsize=(13, 5))
ind = np.arange(1, 101)
ax.axhline(y=0, xmin=0, xmax=n+1, color=[0, 0, 0])
ax.plot([ind, ind], ci, color=[0, 0.2, 0.8, 0.8], marker='_', ms=0, linewidth=3)
ax.plot([ind[out], ind[out]], ci[:, out], color=[1, 0, 0, 0.8], marker='_', ms=0, linewidth=3)
ax.plot(ind, m, color=[0, .8, .2, .8], marker='.', ms=10, linestyle='')
ax.set_xlim(0, 101)
ax.set_ylim(-1.1, 1.1)
ax.set_title("Confidence interval for the samples' mean estimate of a population ~ $N(0, 1)$",
fontsize=18)
ax.set_xlabel('Sample (with %d observations)' %n, fontsize=18)
plt.show()
# Four out of 100 95%-CI's don't contain the population mean, about what we predicted.
#
# And the standard deviation of the samples' mean per definition should be equal to the standard error of the mean:
print("Samples' mean and standard deviation:")
print('m = %.3f s = %.3f' % (np.mean(m), np.mean(s)))
print("Standard deviation of the samples' mean:")
print('%.3f' % np.std(m, ddof=1))
print("Standard error of the mean:")
print('%.3f' % (np.mean(s)/np.sqrt(20)))
# Likewise, it's common to use 1.96 for the 95% confidence interval even when dealing with a sample; let's quantify the error of this approximation for different sample sizes:
T = lambda n: stats.t.ppf(0.975, n-1) # T distribution
N = stats.norm.ppf(0.975) # Normal distribution
for n in [1000, 100, 10]:
print('\nApproximation error for n = %d' %n)
print('Using Normal distribution: %.1f%%' % (100*(N-T(n))/T(n)))
# For n=1000, the approximation is good, for n=10 it is bad, and it always underestimates.
#
# For the case of a multivariate random variable, see [Prediction ellipse and prediction ellipsoid](http://nbviewer.ipython.org/github/demotu/BMC/blob/master/notebooks/PredictionEllipseEllipsoid.ipynb).
# ## References
#
# - <NAME>, <NAME> (1991) [Statistical Intervals: A Guide for Practitioners](http://books.google.com.br/books?id=ADGuRxqt5z4C). <NAME> & Sons.
# - Montgomery (2013) [Applied Statistics and Probability for Engineers](http://books.google.com.br/books?id=_f4KrEcNAfEC). John Wiley & Sons.
| notebooks/ConfidencePredictionIntervals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Summary
# This notebook contains:
# - torch implementations of a few linear algebra techniques:
# - forward- and back-solving
# - LDLt decomposition
# - QR decomposition via Householder reflections
#
# - initial implementations of secure linear regression and <NAME>'s [DASH](https://github.com/jbloom22/DASH/) that leverage PySyft for secure computation.
#
# These implementations linear regression and DASH are not currently strictly secure, in that a few final steps are performed on the local worker for now. That's because our implementations of LDLt decomposition, QR decomposition, etc. don't quite work for the PySyft `AdditiveSharingTensor` just yet. They definitely do in principle (because they're compositions of operations the SPDZ supports), but there are still a few details to hammer out.
#
# ## Contents
# [Ordinary least squares regression and LDLt decomposition](#OLSandLDLt)
# * [LDLt decomposition, forward/back-solving](#LDLt)
# * [Secure linear regression example](#OLS)
#
# [DASH](#dashqr)
# * [QR decomposition via Householder transforms](#qr)
# * [DASH example](#dash)
import numpy as np
import torch as th
import syft as sy
from scipy import stats
sy.create_sandbox(globals())
# # <a id='OLSandLDLt'>Ordinary least squared regression and LDLt decomposition</a>
# ## <a id='LDLt'>LDLt decomposition, forward/back-solving</a>
#
# These are torch implementations of basic linear algebra routines we'll use to perform regression (and also in parts of the next section).
# - Forward/back-solving allows us to solve triangular linear systems efficiently and stably.
# - LDLt decomposition lets us write symmetric matrics as a product LDL^t where L is lower-triangular and D is diagonal (^t denotes transpose). It performs a role similar to Cholesky decomposition (which is normally available as method of a torch tensor), but doesn't require computing square roots. This makes makes LDLt a better fit for the secure setting.
# +
def _eye(n):
"""th.eye doesn't seem to work after hooking torch, so just adding
a workaround for now.
"""
return th.FloatTensor(np.eye(n))
def ldlt_decomposition(x):
"""Decompose the square, symmetric, full-rank matrix X as X = LDL^t, where
- L is upper triangular
- D is diagonal.
"""
n, _ = x.shape
l, diag = _eye(n), th.zeros(n).float()
for j in range(n):
diag[j] = x[j, j] - (th.sum((l[j, :j] ** 2) * diag[:j]))
for i in range(j + 1, n):
l[i, j] = (x[i, j] - th.sum(diag[:j] * l[i, :j] * l[j, :j])) / diag[j]
return l, th.diag(diag), l.transpose(0, 1)
def back_solve(u, y):
"""Solve Ux = y for U a square, upper triangular matrix of full rank"""
n = u.shape[0]
x = th.zeros(n)
for i in range(n - 1, -1, -1):
x[i] = (y[i] - th.sum(u[i, i+1:] * x[i+1:])) / u[i, i]
return x.reshape(-1, 1)
def forward_solve(l, y):
"""Solve Lx = y for L a square, lower triangular matrix of full rank."""
n = l.shape[0]
x = th.zeros(n)
for i in range(0, n):
x[i] = (y[i] - th.sum(l[i, :i] * x[:i])) / l[i, i]
return x.reshape(-1, 1)
def invert_triangular(t, upper=True):
"""
Invert by repeated forward/back-solving.
TODO: -Could be made more efficient with vectorized implementation of forward/backsolve
-detection and validation around triangularity/squareness
"""
solve = back_solve if upper else forward_solve
t_inv = th.zeros_like(t)
n = t.shape[0]
for i in range(n):
e = th.zeros(n, 1)
e[i] = 1.
t_inv[:, [i]] = solve(t, e)
return t_inv
def solve_symmetric(a, y):
"""Solve the linear system Ax = y where A is a symmetric matrix of full rank."""
l, d, lt = ldlt_decomposition(a)
# TODO: more efficient to just extract diagonal of d as 1D vector and scale?
x_ = forward_solve(l.mm(d), y)
return back_solve(lt, x_)
# +
"""
Basic tests for LDLt decomposition.
"""
def _assert_small(x, failure_msg=None, threshold=1E-5):
norm = x.norm()
assert norm < threshold, failure_msg
def test_ldlt_case(a):
l, d, lt = ldlt_decomposition(a)
_assert_small(l - lt.transpose(0, 1))
_assert_small(l.mm(d).mm(lt) - a, 'Decomposition is inaccurate.')
_assert_small(l - th.tril(l), 'L is not lower triangular.')
_assert_small(th.triu(th.tril(d)) - d, 'D is not diagonal.')
print(f'PASSED for {a}')
def test_solve_symmetric_case(a, x):
y = a.mm(x)
_assert_small(solve_symmetric(a, y) - x)
print(f'PASSED for {a}, {x}')
a = th.tensor([[1, 2, 3],
[2, 1, 2],
[3, 2, 1]]).float()
x = th.tensor([1, 2, 3]).float().reshape(-1, 1)
test_ldlt_case(a)
test_solve_symmetric_case(a, x)
# -
# ## <a id='OLS'>Secure linear regression example</a>
# #### Problem
# We're solving
# $$ \min_\beta \|X \beta - y\|_2 $$
# in the situation where the data $(X, y)$ is horizontally partitioned (each worker $w$ owns chunks $X_w, y_w$ of the rows of $X$ and $y$).
#
# #### Goals
# We want to do this
# * securely
# * without network overhead or MPC-related costs that scale with the number of rows of $X$.
#
# #### Plan
#
# 1. (**local plaintext compression**): each worker locally computes $X_w^t X_w$ and $X_w^t y_w$ in plain text. This is the only step that depends on the number of rows of X, and it's performed in plaintext.
# 2. (**secure summing**): securely compute the sums $$\begin{align}X^t X &= \sum_w X^t_w X_w \\ X^t y &= \sum_w X^t_w y_w \end{align}$$ as an AdditiveSharingTensor. Some worker or other party (here the local worker) will have a pointers to those two AdditiveSharingTensors.
# 3. (**secure solve**): We can then solve $X^tX\beta = X^ty$ for $\beta$ by a sequence of operations on those pointers (specifically, we apply `solve_symmetric` defined above).
#
# #### Example data:
# The correct $\beta$ is $[1, 2, -1]$
X = th.tensor(10 * np.random.randn(30000, 3))
y = (X[:, 0] + 2 * X[:, 1] - X[:, 2]).reshape(-1, 1)
# Split the data into chunks and send a chunk to each worker, storing pointers to chunks in two `MultiPointerTensor`s.
# +
workers = [alice, bob, theo]
crypto_provider = jon
chunk_size = int(X.shape[0] / len(workers))
def _get_chunk_pointers(data, chunk_size, workers):
return [
data[(i * chunk_size):((i+1)*chunk_size), :].send(worker)
for i, worker in enumerate(workers)
]
X_ptrs = sy.MultiPointerTensor(
children=_get_chunk_pointers(X, chunk_size, workers))
y_ptrs = sy.MultiPointerTensor(
children=_get_chunk_pointers(y, chunk_size, workers))
# -
# ### local compression
# This is the only step that depends on the number of rows of $X, y$, and it's performed locally on each worker in plain text. The result is two `MultiPointerTensor`s with pointers to each workers' summand of $X^tX$ (or $X^ty$).
# +
Xt_ptrs = X_ptrs.transpose(0, 1)
XtX_summand_ptrs = Xt_ptrs.mm(X_ptrs)
Xty_summand_ptrs = Xt_ptrs.mm(y_ptrs)
# -
# ### secure sum
# We add those summands up in two steps:
# - share each summand among all other workers
# - move the resulting pointers to one place (here just the local worker) and add 'em up.
def _generate_shared_summand_pointers(
summand_ptrs,
workers,
crypto_provider):
for worker_id, summand_pointer in summand_ptrs.child.items():
shared_summand_pointer = summand_pointer.fix_precision().share(
*workers, crypto_provider=crypto_provider)
yield shared_summand_pointer.get()
# +
XtX_shared = sum(
_generate_shared_summand_pointers(
XtX_summand_ptrs, workers, crypto_provider))
Xty_shared = sum(_generate_shared_summand_pointers(
Xty_summand_ptrs, workers, crypto_provider))
# -
# ### secure solve
# The coefficient $\beta$ is the solution to
# $$X^t X \beta = X^t y$$
#
# We solve for $\beta$ using `solve_symmetric`. Critically, this is a composition of linear operations that should be supported by `AdditiveSharingTensor`. Unlike the classic Cholesky decomposition, the $LDL^t$ decomposition in step 1 does not involve taking square roots, which would be challenging.
#
#
# **TODO**: there's still some additional work required to get `solve_symmetric` working for `AdditiveSharingTensor`, so we're performing the final linear solve publicly for now.
beta = solve_symmetric(XtX_shared.get().float_precision(), Xty_shared.get().float_precision())
beta
# # <a id='dashqr'>DASH and QR-decomposition</a>
# ## <a id='qr'>QR decomposition</a>
#
# A $m \times n$ real matrix $A$ with $m \geq n$ can be written as $$A = QR$$ for $Q$ orthogonal and $R$ upper triangular. This is helpful in solving systems of equations, among other things. It is also central to the compression idea of [DASH](https://arxiv.org/pdf/1901.09531.pdf).
# +
"""
Full QR decomposition via Householder transforms,
following Numerical Linear Algebra (Trefethen and Bau).
"""
def _apply_householder_transform(a, v):
return a - 2 * v.mm(v.transpose(0, 1).mm(a))
def _build_householder_matrix(v):
n = v.shape[0]
u = v / v.norm()
return _eye(n) - 2 * u.mm(u.transpose(0, 1))
def _householder_qr_step(a):
x = a[:, 0].reshape(-1, 1)
alpha = x.norm()
u = x.copy()
# note: can get better stability by multiplying by sign(u[0, 0])
# (where sign(0) = 1); is this supported in the secure context?
u[0, 0] += u.norm()
# is there a simple way of getting around computing the norm twice?
u /= u.norm()
a = _apply_householder_transform(a, u)
return a, u
def _recover_q(householder_vectors):
"""
Build the matrix Q from the Householder transforms.
"""
n = len(householder_vectors)
def _apply_transforms(x):
"""Trefethen and Bau, Algorithm 10.3"""
for k in range(n-1, -1, -1):
x[k:, :] = _apply_householder_transform(
x[k:, :],
householder_vectors[k])
return x
m = householder_vectors[0].shape[0]
n = len(householder_vectors)
q = th.zeros(m, m)
# Determine q by evaluating it on a basis
for i in range(m):
e = th.zeros(m, 1)
e[i] = 1.
q[:, [i]] = _apply_transforms(e)
return q
def qr(a, return_q=True):
"""
Args:
a: shape (m, n), m >= n
return_q: bool, whether to reconstruct q
Returns:
orthogonal q of shape (m, m) (None if return_q is False)
upper-triangular of shape (m, n)
"""
m, n = a.shape
assert m >= n, \
f"Passed a of shape {a.shape}, must have a.shape[0] >= a.shape[1]"
r = a.copy()
householder_unit_normal_vectors = []
for k in range(n):
r[k:, k:], u = _householder_qr_step(r[k:, k:])
householder_unit_normal_vectors.append(u)
if return_q:
q = _recover_q(householder_unit_normal_vectors)
else:
q = None
return q, r
# +
"""
Basic tests for QR decomposition
"""
def _test_qr_case(a):
q, r = qr(a)
# actually have QR = A
_assert_small(q.mm(r) - a, "QR = A failed")
# Q is orthogonal
m, _ = a.shape
_assert_small(
q.mm(q.transpose(0, 1)) - _eye(m),
"QQ^t = I failed"
)
# R is upper triangular
lower_triangular_entries = th.tensor([
r[i, j].item() for i in range(r.shape[0])
for j in range(i)])
_assert_small(
lower_triangular_entries,
"R is not upper triangular"
)
print(f"PASSED for \n{a}\n")
def test_qr():
_test_qr_case(
th.tensor([[1, 0, 1],
[1, 1, 0],
[0, 1, 1]]).float()
)
_test_qr_case(
th.tensor([[1, 0, 1],
[1, 1, 0],
[0, 1, 1],
[1, 1, 1],]).float()
)
test_qr()
# -
# ## <a id='dash'>DASH implementation</a>
# We follow https://github.com/jbloom22/DASH/.
#
# The overall structure is roughly analogous to the linear regression example above.
#
# - There's a local compression step that's performed separately on each worker in plaintext.
# - We leverage PySyft's SMCP features to perform secure summation.
# - For now, the last few steps are performed by a single player (the local worker).
# - Again, this could be performed securely, but there are still a few hitches with getting our torch implementation of QR decomposition to work for an `AdditiveSharingTensor`.
# +
def _generate_worker_data_pointers(
n, m, k, worker,
beta_correct, gamma_correct, epsilon=0.01
):
"""
Return pointers to worker-level data.
Args:
n: number of rows
m: number of transient
k: number of covariates
beta_correct: coefficients for transient features (tensor of shape (m, 1))
gamma_correct: coefficients for covariates (tensor of shape (k, 1))
epsilon: scale of noise added to response
Return:
y, X, C: pointers to response, transients, and covariates
"""
X = th.randn(n, m).send(worker)
C = th.randn(n, k).send(worker)
y = (X.mm(beta_correct.copy().send(worker)).reshape(-1, 1) +
C.mm(gamma_correct.copy().send(worker)).reshape(-1, 1))
y += (epsilon * th.randn(n, 1)).send(worker)
return y, X, C
def _dot(x):
return (x * x).sum(dim=0).reshape(-1, 1)
def _secure_sum(worker_level_pointers, workers, crypto_provider):
"""
Securely add up an interable of pointers to (same-sized) tensors.
Args:
worker_level_pointers: iterable of pointer tensors
workers: list of workers
crypto_provider: worker
Returns:
AdditiveSharingTensor shared among workers
"""
return sum([
p.fix_precision(precision_fractional=10).share(*workers, crypto_provider=crypto_provider).get()
for p in worker_level_pointers
])
# -
def dash_example_secure(
workers, crypto_provider,
n_samples_by_worker, m, k,
beta_correct, gamma_correct,
epsilon=0.01
):
"""
Args:
workers: list of workers
crypto_provider: worker
n_samples_by_worker: dict mapping worker ids to ints (number of rows of data)
m: number of transients
k: number of covariates
beta_correct: coefficient for transient features
gamma_correct: coefficient for covariates
epsilon: scale of noise added to response
Returns:
beta, sigma, tstat, pval: coefficient of transients and accompanying statistics
"""
# Generate each worker's data
worker_data_pointers = {
p: _generate_worker_data_pointers(
n, m, k, workers[p],
beta_correct, gamma_correct,
epsilon=epsilon)
for p, n in n_samples_by_worker.items()
}
# to be populated with pointers to results of local, worker-level computations
Ctys, CtXs, yys, Xys, XXs, Rs = {}, {}, {}, {}, {}, {}
def _sum(pointers):
return _secure_sum(pointers, list(players.values()), crypto_provider)
# worker-level compression step
for p, (y, X, C) in worker_data_pointers.items():
# perform worker-level compression step
yys[p] = y.norm()
Xys[p] = X.transpose(0, 1).mm(y)
XXs[p] = _dot(X)
Ctys[p] = C.transpose(0, 1).mm(y)
CtXs[p] = C.transpose(0, 1).mm(X)
_, R_full = qr(C, return_q=False)
Rs[p] = R_full[:k, :]
# Perform secure sum
# - We're returning result to the local worker and computing there for the rest
# of the way, but should be possible to compute via SMPC (on a pointers to AdditiveSharingTensors)
# - still afew minor-looking issues with implementing invert_triangular/qr for
# AdditiveSharingTensor
yy = _sum(yys.values()).get().float_precision()
Xy = _sum(Xys.values()).get().float_precision()
XX = _sum(XXs.values()).get().float_precision()
Cty = _sum(Ctys.values()).get().float_precision()
CtX = _sum(CtXs.values()).get().float_precision()
# Rest is done publicly on the local worker for now
_, R_public = qr(
th.cat([R.get() for R in Rs.values()], dim=0),
return_q=False)
invR_public = invert_triangular(R_public[:k, :])
Qty = invR_public.transpose(0, 1).mm(Cty)
QtX = invR_public.transpose(0, 1).mm(CtX)
QtXQty = QtX.transpose(0, 1).mm(Qty)
QtyQty = _dot(Qty)
QtXQtX = _dot(QtX)
yyq = yy - QtyQty
Xyq = Xy - QtXQty
XXq = XX - QtXQtX
d = sum(n_samples_by_worker.values()) - k - 1
beta = Xyq / XXq
sigma = ((yyq / XXq - (beta ** 2)) / d).abs() ** 0.5
tstat = beta / sigma
pval = 2 * stats.t.cdf(-abs(tstat), d)
return beta, sigma, tstat, pval
# +
players = {
worker.id: worker
for worker in [alice, bob, theo]
}
# de
n_samples_by_player = {
alice.id: 100000,
bob.id: 200000,
theo.id: 100000
}
crypto_provider = jon
m = 100
k = 3
d = sum(n_samples_by_player.values()) - k - 1
beta_correct = th.ones(m, 1)
gamma_correct = th.ones(k, 1)
dash_example_secure(
players, crypto_provider,
n_samples_by_player, m, k,
beta_correct, gamma_correct)
# -
| examples/experimental/plaintext_speed_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## The dataset is from Kaggle where we have 7503 values or rows in training set and 3243 rows in test set and test set does not have target column. Dataset is about Disaster tweets and we will be classifying fake and real tweets
# https://www.kaggle.com/c/nlp-getting-started/data?select=train.csv
import numpy as np
from nltk.tokenize import TweetTokenizer
import pandas as pd
from nltk.corpus import stopwords
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
from nltk.stem.porter import PorterStemmer
from nltk.stem import WordNetLemmatizer
from sklearn.metrics import classification_report, roc_curve, roc_auc_score, plot_confusion_matrix
ps = PorterStemmer()
import re
import seaborn as sns
import nltk
import scipy
import matplotlib.pyplot as plt
import networkx as nx
from gensim.models import word2vec
import spacy
from sklearn.feature_extraction.text import TfidfVectorizer
# %matplotlib inline
from nltk.stem.porter import PorterStemmer
from sklearn.naive_bayes import MultinomialNB
from sklearn.model_selection import train_test_split
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
df = pd.read_csv('train.csv')
df_test = pd.read_csv('test.csv')
print(df.tail(3))
print(df_test.shape)
print(df.shape)
print(df.head(3))
## Checking for missing values
print(df.isnull().sum())
print(df_test.isnull().sum())
# #### We can see that the target variable is not available in df_test dataset and we are going to classify the tweets in df_test dataset
df_test.info()
## Removing missing values
df.drop(['keyword', 'location'], axis = 1, inplace = True)
df_test.drop(['keyword', 'location'], axis = 1, inplace = True)
nltk.download('wordnet')
## Applying Regex to remove punctuation, hyperlinks and numbers
## Converting text to lower
## Applying Lemmatizer to stemming and get the meaninful words
ps = PorterStemmer()
lemmatizer = WordNetLemmatizer()
corpus = []
for i in range(0, len(df)):
#review = re.sub(r'^https?:\/\/.*[\r\n]*', '', df['text'][i], flags=re.MULTILINE)
#review = re.sub(r'(https|http)?:\/\/(\w|\.|\/|\?|\=|\&|\%)*\b', '', df['text'][i], flags=re.MULTILINE)
review = re.sub(r"http\S+", "", df['text'][i])
review = re.sub('[^a-zA-Z\d+]', ' ', review)
review = re.sub('[0-9]', '', review)
review = review.lower()
review = review.split()
#review = [ps.stem(word) for word in review if not word in stopwords.words('english')]
review = [lemmatizer.lemmatize(word, pos = 'v') for word in review if not word in stopwords.words('english')]
review = [lemmatizer.lemmatize(word, pos = 'n') for word in review]
review = [lemmatizer.lemmatize(word, pos = 'a') for word in review]
review = ' '.join(review)
corpus.append(review)
## We have 113,654 words in our dataset
print(df.shape)
df['text'].apply(lambda x: len(x.split(' '))).sum()
## Applying the same for test dataset
ps = PorterStemmer()
lemmatizer = WordNetLemmatizer()
corpus_test = []
for i in range(0, len(df_test)):
review = re.sub(r"http\S+", "", df_test['text'][i])
review = re.sub('[^a-zA-Z\d+]', ' ', review)
review = re.sub('[0-9]', '', review)
review = review.lower()
review = review.split()
review = [lemmatizer.lemmatize(word, pos = 'v') for word in review if not word in stopwords.words('english')]
review = [lemmatizer.lemmatize(word, pos = 'n') for word in review]
review = [lemmatizer.lemmatize(word, pos = 'a') for word in review]
review = ' '.join(review)
corpus_test.append(review)
corpus_test[1:4]
corpus[1:4]
## Before proceeding to model, checking for class imbalance
classes = df['target'].value_counts()
plt.figure(figsize=(4,4))
sns.barplot(classes.index, classes.values, alpha=0.8)
plt.ylabel('Number of Occurrences', fontsize=12)
plt.xlabel('target', fontsize=12)
plt.legend('0', '1')
plt.show()
df['target'].value_counts()
# ### We do not have a class imbalance in our dataset
## Creating a Dictionary to see most frequent words
wordfreq = {}
for sentence in corpus:
tokens = nltk.word_tokenize(sentence)
for token in tokens:
if token not in wordfreq.keys():
wordfreq[token] = 1
else:
wordfreq[token] += 1
## Using heap module in python to see 10 most frequent words
import heapq
most_freq = heapq.nlargest(200, wordfreq, key=wordfreq.get)
most_freq[0:10]
## One way to create features for Bag of words
sentence_vectors = []
for sentence in corpus:
sentence_tokens = nltk.word_tokenize(sentence)
sent_vec = []
for token in most_freq:
if token in sentence_tokens:
sent_vec.append(1)
else:
sent_vec.append(0)
sentence_vectors.append(sent_vec)
sentence_vectors = np.asarray(sentence_vectors)
sentence_vectors
## Importing CountVectorizer to create bag of words and
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features=1000)
X = cv.fit_transform(corpus).toarray()
y = df['target']
## These are the features for Bag of words
X[1:5]
### Splitting data for training and test data and applying Naive Bayes Classification
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.30, random_state = 0)
clf = MultinomialNB().fit(X_train, y_train)
y_pred_clf = clf.predict(X_test)
print("Training set score using Naive Bayes Classifier: {:.2f}".format(clf.score(X_train, y_train)))
print("Testing set score using Naive Bayes Classifier: {:.2f}" .format(clf.score(X_test, y_test)))
lr = LogisticRegression()
print(X_train.shape, y_train.shape)
train = lr.fit(X_train, y_train)
y_pred = lr.predict(X_test)
print('Training set score using Logistic Regression:{:.2f}'.format(train.score(X_train, y_train)))
print('Test set score:{:.2f}'.format(train.score(X_test, y_test)))
plot_confusion_matrix(lr,X_test, y_test)
# +
from sklearn import ensemble
rfc = ensemble.RandomForestClassifier()
train1 = rfc.fit(X_train, y_train)
print('Training set score using Random forest Classifier:{:.2f}'.format(rfc.score(X_train, y_train)))
print('Test set score using Random Forest Classifier:{:.2f}'.format(rfc.score(X_test, y_test)))
# -
print(classification_report(y_test, y_pred))
y_pred_proba = lr.predict_proba(X_test)[:,1]
y_pred_proba
fpr,tpr, thresholds = roc_curve(y_test, y_pred_proba)
plt.plot(fpr,tpr)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title("ROC CURVE of TWEETS", color = 'blue')
plt.xlabel('False Possitive Rate(1-Specificity)')
plt.ylabel('True Possitive Rate(Sensitivity)')
plt.grid(True)
print("The area under ROC CURVE using Logistic Regression with BOW: {:.2f}".format(roc_auc_score(y_test, y_pred_proba)))
# #### Logistic Regression is the best model from the above 3 models but the gap between test and train data is less with Naive Bayes Classifier
# Creating the TF-IDF model
from sklearn.feature_extraction.text import TfidfVectorizer
cv1 = TfidfVectorizer()
X_td = cv1.fit_transform(corpus).toarray()
X_train1, X_test1, y_train1, y_test1 = train_test_split(X_td, y, test_size = 0.20, random_state = 0)
clf1 = MultinomialNB().fit(X_train1, y_train1)
y_pred1 = clf1.predict(X_test1)
confusion_td = confusion_matrix(y_test1, y_pred1)
print(confusion_td)
print("TF-IDF Score for Naive Bayes Training Set is {:.2f}".format(clf1.score(X_train1, y_train1)))
print("TF-IDF Score for Naive Bayes Test Set is: {:.2f}".format(clf1.score(X_test1, y_test1)))
lr1 = LogisticRegression()
train1 = lr1.fit(X_train1, y_train1)
print('TF-IDF score of Training set with Logistic Regression: {:.2f}'.format(lr1.score(X_train1, y_train1)))
print('TF-IDF score for Test set with Logistic Regression: {:.2f}'.format(lr1.score(X_test1, y_test1)))
plot_confusion_matrix(lr1, X_test1, y_test1)
# +
from sklearn import ensemble
rfc2 = ensemble.RandomForestClassifier()
train5 = rfc2.fit(X_train1, y_train1)
print('Training set score using Random forest Classifier:{:.2f}'.format(rfc2.score(X_train1, y_train1)))
print('Test set score using Random Forest Classifier:{:.2f}'.format(rfc2.score(X_test1, y_test1)))
# -
y_pred_tfidf = lr1.predict(X_test1)
print(classification_report(y_test1, y_pred_tfidf))
y_pred_prob1 = lr1.predict_proba(X_test1)[:,1]
fpr,tpr, thresholds = roc_curve(y_test1, y_pred_prob1)
plt.plot(fpr,tpr)
plt.xlim([0.0, 1.0])
plt.ylim([0.0, 1.0])
plt.title("ROC CURVE of TWEETS TFIDF", color = 'blue')
plt.xlabel('False Possitive Rate(1-Specificity)')
plt.ylabel('True Possitive Rate(Sensitivity)')
plt.grid(True)
plt.show()
print('Area under the ROC Curve TFIDF: {:.2f}'.format(roc_auc_score(y_test1, y_pred_prob1)))
text1 = df['text']
text = df['text']
nlp = spacy.load('en_core_web_sm')
text_doc = nlp('text')
tweet_tokenizer = TweetTokenizer()
tokens1 = []
for sent in corpus:
for word in tweet_tokenizer.tokenize(sent):
if len(word) < 2:
continue
tokens1.append(word.lower())
print("The number of tokens we have in our training dataset are {}" .format(len(tokens1)))
## Creating tokens using TweetTokenizer from NLTK library
tweet_tokenizer = TweetTokenizer()
tweet_tokens = []
for sent in corpus:
review2 = tweet_tokenizer.tokenize(sent)
tweet_tokens.append(review2)
tweet_tokens[1]
## Removing punctuation, numbers and hyperlinks from the text
corpus1 = []
for i in range(0, len(df)):
review1 = re.sub(r"http\S+", "", df['text'][i])
review1 = re.sub('[^a-zA-Z\d+]', ' ', review1)
review1 = review1.split()
review1 = ' '.join(review1)
corpus1.append(review1)
## Language Parsing using spacy
nlp = spacy.load('en_core_web_sm')
corpus_spacy = []
for i in corpus1:
text_doc = nlp(i)
corpus_spacy.append(text_doc)
from collections import Counter
# Utility function to calculate how frequently words appear in the text.
def word_frequencies(corpus_spacy, include_stop = False):
# Build a list of words.
# Strip out punctuation and, optionally, stop words.
words = []
for token in corpus_spacy:
for j in token:
if not j.is_punct and (not j.is_stop and not include_stop):
words.append(j.text)
# Build and return a Counter object containing word counts.
return Counter(words)
corpus_freq = word_frequencies(corpus_spacy).most_common(30)
print('corpus_spacy includes stop words:', corpus_freq)
# #### Dividing the data into target1 and target0 inorder to look at freq words in each category
corpus4 = ' '.join(corpus)
# +
import gensim
from gensim.models import word2vec
model = word2vec.Word2Vec(
tweet_tokens,
workers=4, # Number of threads to run in parallel (if your computer does parallel processing).
min_count=50, # Minimum word count threshold.
window=6, # Number of words around target word to consider.
sg=0, # Use CBOW because our corpus is small.
sample=1e-3 , # Penalize frequent words.
size=300, # Word vector length.
hs=1 # Use hierarchical softmax.
)
print('done!')
# -
## Find most similar words to life
print(model.wv.most_similar(positive = ['life']))
print(model.wv.most_similar(negative = ['life']))
# +
## Use t-SNE to represent high-dimensional data in a lower-dimensional space.
from sklearn.manifold import TSNE
def tsne_plot(model):
"Create TSNE model and plot it"
labels = []
tokens = []
for word in model.wv.vocab:
tokens.append(model[word])
labels.append(word)
tsne_model = TSNE(perplexity=40, n_components=2, init='pca', n_iter=2500, random_state=23)
new_values = tsne_model.fit_transform(tokens)
x = []
y = []
for value in new_values:
x.append(value[0])
y.append(value[1])
plt.figure(figsize=(18, 18))
for i in range(len(x)):
plt.scatter(x[i],y[i])
plt.annotate(labels[i],
xy=(x[i], y[i]),
xytext=(5, 2),
textcoords='offset points',
ha='right',
va='bottom')
plt.show()
tsne_plot(model)
# -
# #### We can see from the above plot that words like collide, evacuate, crash, smoke, blow, explode, shoot are closer to each other and they should be in a real tweets.
# +
## Creating wordcloud for visualizing most important words
from PIL import Image
wc_text = corpus4
custom_mask = np.array(Image.open('twitter_mask.png'))
wc = WordCloud(background_color = 'white', max_words = 500, mask = custom_mask, height =
5000, width = 5000)
wc.generate(wc_text)
image_colors = ImageColorGenerator(custom_mask)
plt.figure(figsize=(20,10))
plt.imshow(wc, interpolation = 'bilinear')
plt.axis('off')
plt.show()
# -
## Creating features using bag of words for test data set
cv = CountVectorizer(max_features=1000)
test_features = cv.fit_transform(corpus_test).toarray()
test_features
lr = LogisticRegression()
pred = lr.fit(X,y)
print(test_features.shape)
y_pred1 = lr.predict(test_features)
y_pred1.sum()
# ## We determined that Logistic regression using Bag of words as the best model, using our best model we have classified real and fake tweets. We have 1142 real tweets about disasters and 2121 fake tweets in the predicted test data set.
# During the age of Social Media where we get all the updates and News from social media like Twitter, Facebook, it is very important to differentiate real and fake tweets. With this model we can differentiate real tweets about disasters from fake tweets. This model not only helps in flagging fake tweets, it is also helpful to identify real tweets and assist people who are in need of help. Once a model is deployed into production and providing utility to the business, it is necessary to monitor how well the model is performing to implement something that will continuously update the database as new data is generated. We can use a scalable messaging platform like Kafka to send newly acquired data to a long running Spark Streaming process. The Spark process can then make a new prediction based on the new data and update the operational database.
| Final Capstone.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: odor-states
# language: python
# name: odor-states
# ---
# +
from scipy.linalg import block_diag
import matplotlib as mpl
import matplotlib.pyplot as plt
import networkx as nx
import numpy as np
# %config Completer.use_jedi = False
mpl.rcParams.update({'font.size': 20})
# -
# # Algorithm Schematic (Fig 3)
np.random.seed(39274)
mat = 1-block_diag(np.ones((4,4)),np.ones((3,3)),np.ones((4,4)))
flip = np.random.choice([0,1],p=[0.8,0.2],size=(11,11))
mat2 = np.logical_xor((flip+flip.T>0),mat)
np.random.seed(34715)
inv_G = nx.from_numpy_matrix(1-mat,create_using=nx.Graph)
G = nx.from_numpy_matrix(mat2,create_using=nx.Graph)
G_bar = nx.from_numpy_matrix(1-mat2,create_using=nx.Graph)
pos = nx.layout.fruchterman_reingold_layout(inv_G,k=0.8)
M = G.number_of_edges()
# +
plt.figure(figsize=(6,6))
nodes = nx.draw_networkx_nodes(G, pos, node_size=800, node_color='grey')
edges = nx.draw_networkx_edges(G, pos, node_size=800,
arrowsize=10, width=0.5,edge_color='grey')
ax = plt.gca()
ax.set_axis_off()
plt.savefig(f"Figures/Final.svg")
plt.show()
# +
plt.figure(figsize=(6,6))
nodes = nx.draw_networkx_nodes(G, pos, node_size=800, node_color='grey')
edges = nx.draw_networkx_edges(G_bar, pos, node_size=800,
arrowsize=10, width=0.5,edge_color='grey')
ax = plt.gca()
ax.set_axis_off()
plt.savefig(f"Figures/Final_bar.svg")
plt.show()
# -
np.random.seed(965305)
inv_G = nx.from_numpy_matrix(1-mat,create_using=nx.Graph)
G = nx.from_numpy_matrix(mat2,create_using=nx.Graph)
G_bar = nx.from_numpy_matrix(1-mat2,create_using=nx.Graph)
pos = nx.layout.fruchterman_reingold_layout(inv_G,k=2)
M = G.number_of_edges()
# +
plt.figure(figsize=(6,6))
nodes = nx.draw_networkx_nodes(G, pos, node_size=800, node_color='grey')
edges = nx.draw_networkx_edges(G, pos, node_size=800,
arrowsize=10, width=0.5,edge_color='grey')
ax = plt.gca()
ax.set_axis_off()
plt.savefig(f"Figures/Initial.svg")
plt.show()
# +
plt.figure(figsize=(6,6))
nodes = nx.draw_networkx_nodes(G, pos, node_size=800, node_color='grey')
edges = nx.draw_networkx_edges(G_bar, pos, node_size=800,
arrowsize=10, width=0.5,edge_color='grey')
ax = plt.gca()
ax.set_axis_off()
plt.savefig(f"Figures/Initial_bar.svg")
plt.show()
# -
mat = np.loadtxt(f'../modules/matrix_2.csv',delimiter=",")
module = np.loadtxt(f'../modules/matrix_2_modules.csv')
order = np.argsort(module)
plt.figure(figsize=(7,7))
plt.imshow(mat,aspect='equal',cmap=plt.cm.inferno)
plt.clim(-0.2,1.2)
plt.xticks([0,9,19,29],[1,10,20,30])
plt.xlabel('Neuron Number')
plt.yticks([0,9,19,29],[1,10,20,30],rotation=90)
plt.ylabel('Neuron Number')
plt.savefig("Figures/Initial_mat.svg")
plt.figure(figsize=(7,7))
plt.imshow(1-mat,aspect='equal',cmap=plt.cm.inferno)
plt.clim(-0.2,1.2)
plt.xticks([0,9,19,29],[1,10,20,30])
plt.xlabel('Neuron Number')
plt.yticks([0,9,19,29],[1,10,20,30],rotation=90)
plt.ylabel('Neuron Number')
plt.savefig("Figures/Initial_mat_bar.svg")
plt.figure(figsize=(7,7))
plt.imshow((1-mat)[order,:][:,order],aspect='equal',cmap=plt.cm.inferno)
plt.clim(-0.2,1.2)
plt.xticks(np.arange(30),[f"{x:.0f}" for x in np.sort(module)])
plt.xlabel('Community Number')
plt.yticks(np.arange(30),[f"{x:.0f}" for x in np.sort(module)],rotation=90)
plt.ylabel('Community Number')
plt.savefig("Figures/Final_mat_bar.svg")
plt.figure(figsize=(7,7))
plt.imshow(mat[order,:][:,order],aspect='equal',cmap=plt.cm.inferno)
plt.clim(-0.2,1.2)
plt.xticks(np.arange(30),[f"{x:.0f}" for x in np.sort(module)])
plt.xlabel('Community Number')
plt.yticks(np.arange(30),[f"{x:.0f}" for x in np.sort(module)],rotation=90)
plt.ylabel('Community Number')
plt.savefig("Figures/Final_mat.svg")
| fig3/fig3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from unicode_info.database import generate_data_for_experiment, generate_positive_pairs_consortium, generate_negative_pairs_consortium
def generate_similarities():
num_pairs = 1000
supported_consortium_feature_vectors, supported_consortium_clusters_dict = generate_data_for_experiment()
positive_pairs = generate_positive_pairs_consortium(supported_consortium_clusters_dict, num_pairs)
negative_pairs = generate_negative_pairs_consortium(supported_consortium_clusters_dict, num_pairs)
cos_sim = lambda features_x, features_y: np.dot(features_x, features_y) / (np.linalg.norm(features_x) * np.linalg.norm(features_y))
calc_sim = lambda pair: cos_sim(supported_consortium_feature_vectors[pair[0]], supported_consortium_feature_vectors[pair[1]])
pos_sim = np.array(list(map(calc_sim, positive_pairs)))
neg_sim = np.array(list(map(calc_sim, negative_pairs)))
return pos_sim, neg_sim
def calculate_recall(pos_sim, threshold):
return np.count_nonzero(pos_sim > threshold) / len(pos_sim)
def calculate_fpr(neg_sim, threshold):
return np.count_nonzero(neg_sim > threshold) / len(neg_sim)
# -
pos_sim, neg_sim = generate_similarities()
thres = 0.9
print(calculate_recall(pos_sim, thres))
print(calculate_fpr(neg_sim, thres))
thres = 0.85
print(calculate_recall(pos_sim, thres))
print(calculate_fpr(neg_sim, thres))
thres = 0.80
print(calculate_recall(pos_sim, thres))
print(calculate_fpr(neg_sim, thres))
| threshold_investigation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 第五章 PyTorch常用工具模块
# 在训练神经网络过程中,需要用到很多工具,其中最重要的三部分是:数据、可视化和GPU加速。本章主要介绍Pytorch在这几方面的工具模块,合理使用这些工具能够极大地提高编码效率。
# ### 5.1 数据处理
#
# 在解决深度学习问题的过程中,往往需要花费大量的精力去处理数据,包括图像、文本、语音或其它二进制数据等。数据的处理对训练神经网络来说十分重要,良好的数据处理不仅会加速模型训练,更会提高模型效果。考虑到这点,PyTorch提供了几个高效便捷的工具,以便使用者进行数据处理或增强等操作,同时可通过并行化加速数据加载。
#
# #### 5.1.1 数据加载
#
# 在PyTorch中,数据加载可通过自定义的数据集对象。数据集对象被抽象为`Dataset`类,实现自定义的数据集需要继承Dataset,并实现两个Python魔法方法:
# - `__getitem__`:返回一条数据,或一个样本。`obj[index]`等价于`obj.__getitem__(index)`
# - `__len__`:返回样本的数量。`len(obj)`等价于`obj.__len__()`
# 这里我们以Kaggle经典挑战赛["Dogs vs. Cat"](https://www.kaggle.com/c/dogs-vs-cats/)的数据为例,来详细讲解如何处理数据。"Dogs vs. Cats"是一个分类问题,判断一张图片是狗还是猫,其所有图片都存放在一个文件夹下,根据文件名的前缀判断是狗还是猫。
# %env LS_COLORS = None
# !tree --charset ascii data/dogcat/
import torch as t
from torch.utils import data
# + active=""
# import os
# from PIL import Image
# import numpy as np
#
# class DogCat(data.Dataset):
# def __init__(self, root):
# imgs = os.listdir(root)
# # 所有图片的绝对路径
# # 这里不实际加载图片,只是指定路径,当调用__getitem__时才会真正读图片
# self.imgs = [os.path.join(root, img) for img in imgs]
#
# def __getitem__(self, index):
# img_path = self.imgs[index]
# # dog->1, cat->0
# label = 1 if 'dog' in img_path.split('/')[-1] else 0
# pil_img = Image.open(img_path)
# array = np.asarray(pil_img)
# data = t.from_numpy(array)
# return data, label
#
# def __len__(self):
# return len(self.imgs)
# -
dataset = DogCat('./data/dogcat/')
img, label = dataset[0] # 相当于调用dataset.__getitem__(0)
for img, label in dataset:
print(img.size(), img.float().mean(), label)
# 通过上面的代码,我们学习了如何自定义自己的数据集,并可以依次获取。但这里返回的数据不适合实际使用,因其具有如下两方面问题:
# - 返回样本的形状不一,因每张图片的大小不一样,这对于需要取batch训练的神经网络来说很不友好
# - 返回样本的数值较大,未归一化至[-1, 1]
# 针对上述问题,PyTorch提供了torchvision[^1]。它是一个视觉工具包,提供了很多视觉图像处理的工具,其中`transforms`模块提供了对PIL `Image`对象和`Tensor`对象的常用操作。
#
# 对PIL Image的操作包括:
# - `Scale`:调整图片尺寸,长宽比保持不变
# - `CenterCrop`、`RandomCrop`、`RandomResizedCrop`: 裁剪图片
# - `Pad`:填充
# - `ToTensor`:将PIL Image对象转成Tensor,会自动将[0, 255]归一化至[0, 1]
#
# 对Tensor的操作包括:
# - Normalize:标准化,即减均值,除以标准差
# - ToPILImage:将Tensor转为PIL Image对象
#
# 如果要对图片进行多个操作,可通过`Compose`函数将这些操作拼接起来,类似于`nn.Sequential`。注意,这些操作定义后是以函数的形式存在,真正使用时需调用它的`__call__`方法,这点类似于`nn.Module`。例如要将图片调整为$224\times 224$,首先应构建这个操作`trans = Resize((224, 224))`,然后调用`trans(img)`。下面我们就用transforms的这些操作来优化上面实现的dataset。
# [^1]: https://github.com/pytorch/vision/
# +
import os
from PIL import Image
import numpy as np
from torchvision import transforms as T
transform = T.Compose([
T.Resize(224), # 缩放图片(Image),保持长宽比不变,最短边为224像素
T.CenterCrop(224), # 从图片中间切出224*224的图片
T.ToTensor(), # 将图片(Image)转成Tensor,归一化至[0, 1]
T.Normalize(mean=[.5, .5, .5], std=[.5, .5, .5]) # 标准化至[-1, 1],规定均值和标准差
])
class DogCat(data.Dataset):
def __init__(self, root, transforms=None):
imgs = os.listdir(root)
self.imgs = [os.path.join(root, img) for img in imgs]
self.transforms=transforms
def __getitem__(self, index):
img_path = self.imgs[index]
label = 0 if 'dog' in img_path.split('/')[-1] else 1
data = Image.open(img_path)
if self.transforms:
data = self.transforms(data)
return data, label
def __len__(self):
return len(self.imgs)
dataset = DogCat('./data/dogcat/', transforms=transform)
img, label = dataset[0]
for img, label in dataset:
print(img.size(), label)
# -
# 除了上述操作之外,transforms还可通过`Lambda`封装自定义的转换策略。例如想对PIL Image进行随机旋转,则可写成这样`trans=T.Lambda(lambda img: img.rotate(random()*360))`。
# torchvision已经预先实现了常用的Dataset,包括前面使用过的CIFAR-10,以及ImageNet、COCO、MNIST、LSUN等数据集,可通过诸如`torchvision.datasets.CIFAR10`来调用,具体使用方法请参看官方文档[^1]。在这里介绍一个会经常使用到的Dataset——`ImageFolder`,它的实现和上述的`DogCat`很相似。`ImageFolder`假设所有的文件按文件夹保存,每个文件夹下存储同一个类别的图片,文件夹名为类名,其构造函数如下:
# ```
# ImageFolder(root, transform=None, target_transform=None, loader=default_loader)
# ```
# 它主要有四个参数:
# - `root`:在root指定的路径下寻找图片
# - `transform`:对PIL Image进行的转换操作,transform的输入是使用loader读取图片的返回对象
# - `target_transform`:对label的转换
# - `loader`:给定路径后如何读取图片,默认读取为RGB格式的PIL Image对象
#
# label是按照文件夹名顺序排序后存成字典,即{类名:类序号(从0开始)},一般来说最好直接将文件夹命名为从0开始的数字,这样会和ImageFolder实际的label一致,如果不是这种命名规范,建议看看`self.class_to_idx`属性以了解label和文件夹名的映射关系。
#
# [^1]: http://pytorch.org/docs/master/torchvision/datasets.html
# !tree --charset ASCII data/dogcat_2/
from torchvision.datasets import ImageFolder
dataset = ImageFolder('data/dogcat_2/')
# cat文件夹的图片对应label 0,dog对应1
dataset.class_to_idx
# 所有图片的路径和对应的label
dataset.imgs
# 没有任何的transform,所以返回的还是PIL Image对象
dataset[0][1] # 第一维是第几张图,第二维为1返回label
dataset[0][0] # 为0返回图片数据
# 加上transform
normalize = T.Normalize(mean=[0.4, 0.4, 0.4], std=[0.2, 0.2, 0.2])
transform = T.Compose([
T.RandomResizedCrop(224),
T.RandomHorizontalFlip(),
T.ToTensor(),
normalize,
])
dataset = ImageFolder('data/dogcat_2/', transform=transform)
# 深度学习中图片数据一般保存成CxHxW,即通道数x图片高x图片宽
dataset[0][0].size()
to_img = T.ToPILImage()
# 0.2和0.4是标准差和均值的近似
to_img(dataset[0][0]*0.2+0.4)
# `Dataset`只负责数据的抽象,一次调用`__getitem__`只返回一个样本。前面提到过,在训练神经网络时,最好是对一个batch的数据进行操作,同时还需要对数据进行shuffle和并行加速等。对此,PyTorch提供了`DataLoader`帮助我们实现这些功能。
# DataLoader的函数定义如下:
# `DataLoader(dataset, batch_size=1, shuffle=False, sampler=None, num_workers=0, collate_fn=default_collate, pin_memory=False, drop_last=False)`
#
# - dataset:加载的数据集(Dataset对象)
# - batch_size:batch size
# - shuffle::是否将数据打乱
# - sampler: 样本抽样,后续会详细介绍
# - num_workers:使用多进程加载的进程数,0代表不使用多进程
# - collate_fn: 如何将多个样本数据拼接成一个batch,一般使用默认的拼接方式即可
# - pin_memory:是否将数据保存在pin memory区,pin memory中的数据转到GPU会快一些
# - drop_last:dataset中的数据个数可能不是batch_size的整数倍,drop_last为True会将多出来不足一个batch的数据丢弃
from torch.utils.data import DataLoader
dataloader = DataLoader(dataset, batch_size=3, shuffle=True, num_workers=0, drop_last=False)
dataiter = iter(dataloader)
imgs, labels = next(dataiter)
imgs.size() # batch_size, channel, height, weight
# dataloader是一个可迭代的对象,意味着我们可以像使用迭代器一样使用它,例如:
# ```python
# for batch_datas, batch_labels in dataloader:
# train()
# ```
# 或
# ```
# dataiter = iter(dataloader)
# batch_datas, batch_labesl = next(dataiter)
# ```
# 在数据处理中,有时会出现某个样本无法读取等问题,比如某张图片损坏。这时在`__getitem__`函数中将出现异常,此时最好的解决方案即是将出错的样本剔除。如果实在是遇到这种情况无法处理,则可以返回None对象,然后在`Dataloader`中实现自定义的`collate_fn`,将空对象过滤掉。但要注意,在这种情况下dataloader返回的batch数目会少于batch_size。
# +
class NewDogCat(DogCat): # 继承前面实现的DogCat数据集
def __getitem__(self, index):
try:
# 调用父类的获取函数,即 DogCat.__getitem__(self, index)
return super(NewDogCat,self).__getitem__(index)
except:
return None, None
from torch.utils.data.dataloader import default_collate # 导入默认的拼接方式
def my_collate_fn(batch):
'''
batch中每个元素形如(data, label)
'''
# 过滤为None的数据
batch = list(filter(lambda x:x[0] is not None, batch))
if len(batch) == 0: return t.Tensor()
return default_collate(batch) # 用默认方式拼接过滤后的batch数据
# -
dataset = NewDogCat('data/dogcat_wrong/', transforms=transform)
dataset[5]
dataloader = DataLoader(dataset, 2, collate_fn=my_collate_fn, num_workers=1,shuffle=True)
for batch_datas, batch_labels in dataloader:
print(batch_datas.size(),batch_labels.size())
# 来看一下上述batch_size的大小。其中第2个的batch_size为1,这是因为有一张图片损坏,导致其无法正常返回。而最后1个的batch_size也为1,这是因为共有9张(包括损坏的文件)图片,无法整除2(batch_size),因此最后一个batch的数据会少于batch_szie,可通过指定`drop_last=True`来丢弃最后一个不足batch_size的batch。
# 对于诸如样本损坏或数据集加载异常等情况,还可以通过其它方式解决。例如但凡遇到异常情况,就随机取一张图片代替:
# ```python
# class NewDogCat(DogCat):
# def __getitem__(self, index):
# try:
# return super(NewDogCat, self).__getitem__(index)
# except:
# new_index = random.randint(0, len(self)-1)
# return self[new_index]
# ```
# 相比较丢弃异常图片而言,这种做法会更好一些,因为它能保证每个batch的数目仍是batch_size。但在大多数情况下,最好的方式还是对数据进行彻底清洗。
# DataLoader里面并没有太多的魔法方法,它封装了Python的标准库`multiprocessing`,使其能够实现多进程加速。在此提几点关于Dataset和DataLoader使用方面的建议:
# 1. 高负载的操作放在`__getitem__`中,如加载图片等。
# 2. dataset中应尽量只包含只读对象,避免修改任何可变对象,利用多线程进行操作。
#
# 第一点是因为多进程会并行的调用`__getitem__`函数,将负载高的放在`__getitem__`函数中能够实现并行加速。
# 第二点是因为dataloader使用多进程加载,如果在`Dataset`实现中使用了可变对象,可能会有意想不到的冲突。在多线程/多进程中,修改一个可变对象,需要加锁,但是dataloader的设计使得其很难加锁(在实际使用中也应尽量避免锁的存在),因此最好避免在dataset中修改可变对象。例如下面就是一个不好的例子,在多进程处理中`self.num`可能与预期不符,这种问题不会报错,因此难以发现。如果一定要修改可变对象,建议使用Python标准库`Queue`中的相关数据结构。
#
# ```python
# class BadDataset(Dataset):
# def __init__(self):
# self.datas = range(100)
# self.num = 0 # 取数据的次数
# def __getitem__(self, index):
# self.num += 1
# return self.datas[index]
# ```
# 使用Python `multiprocessing`库的另一个问题是,在使用多进程时,如果主程序异常终止(比如用Ctrl+C强行退出),相应的数据加载进程可能无法正常退出。这时你可能会发现程序已经退出了,但GPU显存和内存依旧被占用着,或通过`top`、`ps aux`依旧能够看到已经退出的程序,这时就需要手动强行杀掉进程。建议使用如下命令:
#
# ```
# ps x | grep <cmdline> | awk '{print $1}' | xargs kill
# ```
#
# - `ps x`:获取当前用户的所有进程
# - `grep <cmdline>`:找到已经停止的PyTorch程序的进程,例如你是通过python train.py启动的,那你就需要写`grep 'python train.py'`
# - `awk '{print $1}'`:获取进程的pid
# - `xargs kill`:杀掉进程,根据需要可能要写成`xargs kill -9`强制杀掉进程
#
# 在执行这句命令之前,建议先打印确认一下是否会误杀其它进程
# ```
# ps x | grep <cmdline> | ps x
# ```
# PyTorch中还单独提供了一个`sampler`模块,用来对数据进行采样。常用的有随机采样器:`RandomSampler`,当dataloader的`shuffle`参数为True时,系统会自动调用这个采样器,实现打乱数据。默认的是采用`SequentialSampler`,它会按顺序一个一个进行采样。这里介绍另外一个很有用的采样方法:
# `WeightedRandomSampler`,它会根据每个样本的权重选取数据,在样本比例不均衡的问题中,可用它来进行重采样。
#
# 构建`WeightedRandomSampler`时需提供两个参数:每个样本的权重`weights`、共选取的样本总数`num_samples`,以及一个可选参数`replacement`。权重越大的样本被选中的概率越大,待选取的样本数目一般小于全部的样本数目。`replacement`用于指定是否可以重复选取某一个样本,默认为True,即允许在一个epoch中重复采样某一个数据。如果设为False,则当某一类的样本被全部选取完,但其样本数目仍未达到num_samples时,sampler将不会再从该类中选择数据,此时可能导致`weights`参数失效。下面举例说明。
# +
dataset = DogCat('data/dogcat/', transforms=transform)
# 狗的图片被取出的概率是猫的概率的两倍
# 两类图片被取出的概率与weights的绝对大小无关,只和比值有关
weights = [2 if label == 1 else 1 for data, label in dataset]
weights
# -
from torch.utils.data.sampler import WeightedRandomSampler
sampler = WeightedRandomSampler(weights,\
num_samples=9,\
replacement=True)
dataloader = DataLoader(dataset,
batch_size=3,
sampler=sampler)
for datas, labels in dataloader:
print(labels.tolist())
# 可见猫狗样本比例约为1:2,另外一共只有8个样本,但是却返回了9个,说明肯定有被重复返回的,这就是replacement参数的作用,下面将replacement设为False试试。
sampler = WeightedRandomSampler(weights, 8, replacement=False)
dataloader = DataLoader(dataset, batch_size=4, sampler=sampler)
for datas, labels in dataloader:
print(labels.tolist())
# 在这种情况下,num_samples等于dataset的样本总数,为了不重复选取,sampler会将每个样本都返回,这样就失去weight参数的意义了。
#
# 从上面的例子可见sampler在样本采样中的作用:如果指定了sampler,shuffle将不再生效,并且sampler.num_samples会覆盖dataset的实际大小,即一个epoch返回的图片总数取决于`sampler.num_samples`。
# ### 5.2 计算机视觉工具包:torchvision
# 计算机视觉是深度学习中最重要的一类应用,为了方便研究者使用,PyTorch团队专门开发了一个视觉工具包`torchvion`,这个包独立于PyTorch,需通过`pip instal torchvision`安装。在之前的例子中我们已经见识到了它的部分功能,这里再做一个系统性的介绍。torchvision主要包含三部分:
#
# - models:提供深度学习中各种经典网络的网络结构以及预训练好的模型,包括`AlexNet`、VGG系列、ResNet系列、Inception系列等。
# - datasets: 提供常用的数据集加载,设计上都是继承`torhc.utils.data.Dataset`,主要包括`MNIST`、`CIFAR10/100`、`ImageNet`、`COCO`等。
# - transforms:提供常用的数据预处理操作,主要包括对Tensor以及PIL Image对象的操作。
# +
from torchvision import models
from torch import nn
# 加载预训练好的模型,如果不存在会进行下载
# 预训练好的模型保存在 ~/.torch/models/下面
resnet34 = models.squeezenet1_1(pretrained=True, num_classes=1000)
# 修改最后的全连接层为10分类问题(默认是ImageNet上的1000分类)
resnet34.fc=nn.Linear(512, 10)
# -
from torchvision import datasets
# 指定数据集路径为data,如果数据集不存在则进行下载
# 通过train=False获取测试集
dataset = datasets.MNIST('data/', download=True, train=False, transform=transform)
# Transforms中涵盖了大部分对Tensor和PIL Image的常用处理,这些已在上文提到,这里就不再详细介绍。需要注意的是转换分为两步,第一步:构建转换操作,例如`transf = transforms.Normalize(mean=x, std=y)`,第二步:执行转换操作,例如`output = transf(input)`。另外还可将多个处理操作用Compose拼接起来,形成一个处理转换流程。
from torchvision import transforms
to_pil = transforms.ToPILImage()
to_pil(t.randn(3, 64, 64))
# torchvision还提供了两个常用的函数。一个是`make_grid`,它能将多张图片拼接成一个网格中;另一个是`save_img`,它能将Tensor保存成图片。
len(dataset)
dataloader = DataLoader(dataset, shuffle=True, batch_size=16)
from torchvision.utils import make_grid, save_image
dataiter = iter(dataloader)
img = make_grid(next(dataiter)[0], 4) # 拼成4*4网格图片,且会转成3通道
to_img(img)
save_image(img, 'a.png')
Image.open('a.png')
# ### 5.3 可视化工具
# 在训练神经网络时,我们希望能更直观地了解训练情况,包括损失曲线、输入图片、输出图片、卷积核的参数分布等信息。这些信息能帮助我们更好地监督网络的训练过程,并为参数优化提供方向和依据。最简单的办法就是打印输出,但其只能打印数值信息,不够直观,同时无法查看分布、图片、声音等。在本节,我们将介绍两个深度学习中常用的可视化工具:Tensorboard和Visdom。
# #### 5.3.1 Tensorboard
#
# Tensorboard最初是作为TensorFlow的可视化工具迅速流行开来。作为和TensorFlow深度集成的工具,Tensorboard能够展现你的TensorFlow网络计算图,绘制图像生成的定量指标图以及附加数据。但同时Tensorboard也是一个相对独立的工具,只要用户保存的数据遵循相应的格式,tensorboard就能读取这些数据并进行可视化。这里我们将主要介绍如何在PyTorch中使用tensorboardX[^1]进行训练损失的可视化。
# TensorboardX是将Tensorboard的功能抽取出来,使得非TensorFlow用户也能使用它进行可视化,几乎支持原生TensorBoard的全部功能。
#
# 
# [^1]:https://github.com/lanpa/tensorboardX
#
# tensorboard的安装主要分为以下两步:
# - 安装TensorFlow:如果电脑中已经安装完TensorFlow可以跳过这一步,如果电脑中尚未安装,建议安装CPU-Only的版本,具体安装教程参见TensorFlow官网[^1],或使用pip直接安装,推荐使用清华的软件源[^2]。
# - 安装tensorboard: `pip install tensorboard`
# - 安装tensorboardX:可通过`pip install tensorboardX`命令直接安装。
#
# tensorboardX的使用非常简单。首先用如下命令启动tensorboard:
# ```bash
# tensorboard --logdir <your/running/dir> --port <your_bind_port>
# ```
#
# 下面举例说明tensorboardX的使用。
#
# [^1]: https://www.tensorflow.org/install/
# [^2]: https://mirrors.tuna.tsinghua.edu.cn/help/tensorflow/
from tensorboardX import SummaryWriter
# 构建logger对象,logdir用来指定log文件的保存路径
# flush_secs用来指定刷新同步间隔
logger = SummaryWriter(log_dir='experimient_cnn', flush_secs=2)
for ii in range(100):
logger.add_scalar('data/loss', 10-ii**0.5)
logger.add_scalar('data/accuracy', ii**0.5/10)
# 打开浏览器输入`http://localhost:6006`(其中6006应改成你的tensorboard所绑定的端口),即可看到如图2所示的结果。
# 
# 左侧的Horizontal Axis下有三个选项,分别是:
# - Step:根据步长来记录,log_value时如果有步长,则将其作为x轴坐标描点画线。
# - Relative:用前后相对顺序描点画线,可认为logger自己维护了一个`step`属性,每调用一次log_value就自动加1。
# - Wall:按时间排序描点画线。
#
# 左侧的Smoothing条可以左右拖动,用来调节平滑的幅度。点击右上角的刷新按钮可立即刷新结果,默认是每30s自动刷新数据。可见tensorboard_logger的使用十分简单,但它只能统计简单的数值信息,不支持其它功能。
#
# 感兴趣的读者可以从github项目主页获取更多信息,本节将把更多的内容留给另一个可视化工具:Visdom。
# [^4]: https://github.com/lanpa/tensorboard-pytorch
# #### 5.3.2 Visdom
# Visdom[^5]是Facebook专门为PyTorch开发的一款可视化工具,其开源于2017年3月。Visdom十分轻量级,但却支持非常丰富的功能,能胜任大多数的科学运算可视化任务。其可视化界面如图3所示。
#
# 
# [^5]: https://github.com/facebookresearch/visdom
# Visdom可以创造、组织和共享多种数据的可视化,包括数值、图像、文本,甚至是视频,其支持PyTorch、Torch及Numpy。用户可通过编程组织可视化空间,或通过用户接口为生动数据打造仪表板,检查实验结果或调试代码。
#
# Visdom中有两个重要概念:
# - env:环境。不同环境的可视化结果相互隔离,互不影响,在使用时如果不指定env,默认使用`main`。不同用户、不同程序一般使用不同的env。
# - pane:窗格。窗格可用于可视化图像、数值或打印文本等,其可以拖动、缩放、保存和关闭。一个程序中可使用同一个env中的不同pane,每个pane可视化或记录某一信息。
#
# 如图4所示,当前env共有两个pane,一个用于打印log,另一个用于记录损失函数的变化。点击clear按钮可以清空当前env的所有pane,点击save按钮可将当前env保存成json文件,保存路径位于`~/.visdom/`目录下。也可修改env的名字后点击fork,保存当前env的状态至更名后的env。
# 
#
# Visdom的安装可通过命令`pip install visdom`。安装完成后,需通过`python -m visdom.server`命令启动visdom服务,或通过`nohup python -m visdom.server &`命令将服务放至后台运行。Visdom服务是一个web server服务,默认绑定8097端口,客户端与服务器间通过tornado进行非阻塞交互。
#
# Visdom的使用有两点需要注意的地方:
# - 需手动指定保存env,可在web界面点击save按钮或在程序中调用save方法,否则visdom服务重启后,env等信息会丢失。
# - 客户端与服务器之间的交互采用tornado异步框架,可视化操作不会阻塞当前程序,网络异常也不会导致程序退出。
#
# Visdom以Plotly为基础,支持丰富的可视化操作,下面举例说明一些最常用的操作。
# + language="sh"
# # 启动visdom服务器
# # nohup python -m visdom.server &
# +
import torch as t
# +
import visdom
# 新建一个连接客户端
# 指定env = u'test1',默认端口为8097,host是‘localhost'
vis = visdom.Visdom(env=u'test1',use_incoming_socket=False)
x = t.arange(1, 30, 0.01)
y = t.sin(x)
vis.line(X=x, Y=y, win='sinx', opts={'title': 'y=sin(x)'})
# -
# 输出的结果如图5所示。
# 
# 下面逐一分析这几行代码:
# - vis = visdom.Visdom(env=u'test1'),用于构建一个客户端,客户端除指定env之外,还可以指定host、port等参数。
# - vis作为一个客户端对象,可以使用常见的画图函数,包括:
#
# - line:类似Matlab中的`plot`操作,用于记录某些标量的变化,如损失、准确率等
# - image:可视化图片,可以是输入的图片,也可以是GAN生成的图片,还可以是卷积核的信息
# - text:用于记录日志等文字信息,支持html格式
# - histgram:可视化分布,主要是查看数据、参数的分布
# - scatter:绘制散点图
# - bar:绘制柱状图
# - pie:绘制饼状图
# - 更多操作可参考visdom的github主页
#
# 这里主要介绍深度学习中常见的line、image和text操作。
#
# Visdom同时支持PyTorch的tensor和Numpy的ndarray两种数据结构,但不支持Python的int、float等类型,因此每次传入时都需先将数据转成ndarray或tensor。上述操作的参数一般不同,但有两个参数是绝大多数操作都具备的:
# - win:用于指定pane的名字,如果不指定,visdom将自动分配一个新的pane。如果两次操作指定的win名字一样,新的操作将覆盖当前pane的内容,因此建议每次操作都重新指定win。
# - opts:选项,接收一个字典,常见的option包括`title`、`xlabel`、`ylabel`、`width`等,主要用于设置pane的显示格式。
#
# 之前提到过,每次操作都会覆盖之前的数值,但往往我们在训练网络的过程中需不断更新数值,如损失值等,这时就需要指定参数`update='append'`来避免覆盖之前的数值。而除了使用update参数以外,还可以使用`vis.updateTrace`方法来更新图,但`updateTrace`不仅能在指定pane上新增一个和已有数据相互独立的Trace,还能像`update='append'`那样在同一条trace上追加数据。
# +
# append 追加数据
for ii in range(0, 10):
# y = x
x = t.Tensor([ii])
y = x
vis.line(X=x, Y=y, win='polynomial', update='append' if ii>0 else None)
# updateTrace 新增一条线
x = t.arange(0, 9, 0.1)
y = (x ** 2) / 9
vis.line(X=x, Y=y, win='polynomial', name='this is a new Trace',update='new')
# -
# 打开浏览器,输入`http://localhost:8097`,可以看到如图6所示的结果。
# 
# image的画图功能可分为如下两类:
# - `image`接收一个二维或三维向量,$H\times W$或$3 \times H\times W$,前者是黑白图像,后者是彩色图像。
# - `images`接收一个四维向量$N\times C\times H\times W$,$C$可以是1或3,分别代表黑白和彩色图像。可实现类似torchvision中make_grid的功能,将多张图片拼接在一起。`images`也可以接收一个二维或三维的向量,此时它所实现的功能与image一致。
# +
# 可视化一个随机的黑白图片
vis.image(t.randn(64, 64).numpy())
# 随机可视化一张彩色图片
vis.image(t.randn(3, 64, 64).numpy(), win='random2')
# 可视化36张随机的彩色图片,每一行6张
vis.images(t.randn(36, 3, 64, 64).numpy(), nrow=6, win='random3', opts={'title':'random_imgs'})
# -
# 其中images的可视化输出如图7所示。
# 
# `vis.text`用于可视化文本,支持所有的html标签,同时也遵循着html的语法标准。例如,换行需使用`<br>`标签,`\r\n`无法实现换行。下面举例说明。
vis.text(u'''<h1>Hello Visdom</h1><br>Visdom是Facebook专门为<b>PyTorch</b>开发的一个可视化工具,
在内部使用了很久,在2017年3月份开源了它。
Visdom十分轻量级,但是却有十分强大的功能,支持几乎所有的科学运算可视化任务''',
win='visdom',
opts={'title': u'visdom简介' }
)
# 
# ### 5.4 使用GPU加速:cuda
# 这部分内容在前面介绍Tensor、Module时大都提到过,这里将做一个总结,并深入介绍相关应用。
#
# 在PyTorch中以下数据结构分为CPU和GPU两个版本:
# - Tensor
# - nn.Module(包括常用的layer、loss function,以及容器Sequential等)
#
# 它们都带有一个`.cuda`方法,调用此方法即可将其转为对应的GPU对象。注意,`tensor.cuda`会返回一个新对象,这个新对象的数据已转移至GPU,而之前的tensor还在原来的设备上(CPU)。而`module.cuda`则会将所有的数据都迁移至GPU,并返回自己。所以`module = module.cuda()`和`module.cuda()`所起的作用一致。
#
# nn.Module在GPU与CPU之间的转换,本质上还是利用了Tensor在GPU和CPU之间的转换。`nn.Module`的cuda方法是将nn.Module下的所有parameter(包括子module的parameter)都转移至GPU,而Parameter本质上也是tensor(Tensor的子类)。
#
# 下面将举例说明,这部分代码需要你具有两块GPU设备。
#
# P.S. 为什么将数据转移至GPU的方法叫做`.cuda`而不是`.gpu`,就像将数据转移至CPU调用的方法是`.cpu`?这是因为GPU的编程接口采用CUDA,而目前并不是所有的GPU都支持CUDA,只有部分Nvidia的GPU才支持。PyTorch未来可能会支持AMD的GPU,而AMD GPU的编程接口采用OpenCL,因此PyTorch还预留着`.cl`方法,用于以后支持AMD等的GPU。
tensor = t.Tensor(3, 4)
# 返回一个新的tensor,保存在第1块GPU上,但原来的tensor并没有改变
tensor.cuda(0)
tensor.is_cuda # False
# 不指定所使用的GPU设备,将默认使用第1块GPU
tensor = tensor.cuda()
tensor.is_cuda # True
module = nn.Linear(3, 4)
module.cuda(device = 1)
module.weight.is_cuda # True
class VeryBigModule(nn.Module):
def __init__(self):
super(VeryBigModule, self).__init__()
self.GiantParameter1 = t.nn.Parameter(t.randn(100000, 20000)).cuda(0)
self.GiantParameter2 = t.nn.Parameter(t.randn(20000, 100000)).cuda(1)
def forward(self, x):
x = self.GiantParameter1.mm(x.cuda(0))
x = self.GiantParameter2.mm(x.cuda(1))
return x
# 上面最后一部分中,两个Parameter所占用的内存空间都非常大,大概是8个G,如果将这两个都同时放在一块GPU上几乎会将显存占满,无法再进行任何其它运算。此时可通过这种方式将不同的计算分布到不同的GPU中。
# 关于使用GPU的一些建议:
# - GPU运算很快,但对于很小的运算量来说,并不能体现出它的优势,因此对于一些简单的操作可直接利用CPU完成
# - 数据在CPU和GPU之间,以及GPU与GPU之间的传递会比较耗时,应当尽量避免
# - 在进行低精度的计算时,可以考虑`HalfTensor`,它相比于`FloatTensor`能节省一半的显存,但需千万注意数值溢出的情况。
# 另外这里需要专门提一下,大部分的损失函数也都属于`nn.Moudle`,但在使用GPU时,很多时候我们都忘记使用它的`.cuda`方法,这在大多数情况下不会报错,因为损失函数本身没有可学习的参数(learnable parameters)。但在某些情况下会出现问题,为了保险起见同时也为了代码更规范,应记得调用`criterion.cuda`。下面举例说明。
# +
# 交叉熵损失函数,带权重
criterion = t.nn.CrossEntropyLoss(weight=t.Tensor([1, 3]))
input = t.randn(4, 2).cuda()
target = t.Tensor([1, 0, 0, 1]).long().cuda()
# 下面这行会报错,因weight未被转移至GPU
# loss = criterion(input, target)
# 这行则不会报错
criterion.cuda()
loss = criterion(input, target)
criterion._buffers
# -
# 而除了调用对象的`.cuda`方法之外,还可以使用`torch.cuda.device`,来指定默认使用哪一块GPU,或使用`torch.set_default_tensor_type`使程序默认使用GPU,不需要手动调用cuda。
# +
# 如果未指定使用哪块GPU,默认使用GPU 0
x = t.cuda.FloatTensor(2, 3)
# x.get_device() == 0
y = t.FloatTensor(2, 3).cuda()
# y.get_device() == 0
# 指定默认使用GPU 1
with t.cuda.device(1):
# 在GPU 1上构建tensor
a = t.cuda.FloatTensor(2, 3)
# 将tensor转移至GPU 1
b = t.FloatTensor(2, 3).cuda()
print(a.get_device() == b.get_device() == 1 )
c = a + b
print(c.get_device() == 1)
z = x + y
print(z.get_device() == 0)
# 手动指定使用GPU 0
d = t.randn(2, 3).cuda(0)
print(d.get_device() == 2)
# -
t.set_default_tensor_type('torch.cuda.FloatTensor') # 指定默认tensor的类型为GPU上的FloatTensor
a = t.ones(2, 3)
a.is_cuda
# 如果服务器具有多个GPU,`tensor.cuda()`方法会将tensor保存到第一块GPU上,等价于`tensor.cuda(0)`。此时如果想使用第二块GPU,需手动指定`tensor.cuda(1)`,而这需要修改大量代码,很是繁琐。这里有两种替代方法:
#
# - 一种是先调用`t.cuda.set_device(1)`指定使用第二块GPU,后续的`.cuda()`都无需更改,切换GPU只需修改这一行代码。
# - 更推荐的方法是设置环境变量`CUDA_VISIBLE_DEVICES`,例如当`export CUDA_VISIBLE_DEVICE=1`(下标是从0开始,1代表第二块GPU),只使用第二块物理GPU,但在程序中这块GPU会被看成是第一块逻辑GPU,因此此时调用`tensor.cuda()`会将Tensor转移至第二块物理GPU。`CUDA_VISIBLE_DEVICES`还可以指定多个GPU,如`export CUDA_VISIBLE_DEVICES=0,2,3`,那么第一、三、四块物理GPU会被映射成第一、二、三块逻辑GPU,`tensor.cuda(1)`会将Tensor转移到第三块物理GPU上。
#
# 设置`CUDA_VISIBLE_DEVICES`有两种方法,一种是在命令行中`CUDA_VISIBLE_DEVICES=0,1 python main.py`,一种是在程序中`import os;os.environ["CUDA_VISIBLE_DEVICES"] = "2"`。如果使用IPython或者Jupyter notebook,还可以使用`%env CUDA_VISIBLE_DEVICES=1,2`来设置环境变量。
# 从 0.4 版本开始,pytorch新增了`tensor.to(device)`方法,能够实现设备透明,便于实现CPU/GPU兼容。这部份内容已经在第三章讲解过了。
# 从PyTorch 0.2版本中,PyTorch新增分布式GPU支持。分布式是指有多个GPU在多台服务器上,而并行一般指的是一台服务器上的多个GPU。分布式涉及到了服务器之间的通信,因此比较复杂,PyTorch封装了相应的接口,可以用几句简单的代码实现分布式训练。分布式对普通用户来说比较遥远,因为搭建一个分布式集群的代价十分大,使用也比较复杂。相比之下一机多卡更加现实。对于分布式训练,这里不做太多的介绍,感兴趣的读者可参考文档[^distributed]。
# [^distributed]: http://pytorch.org/docs/distributed.html
# #### 5.4.1 单机多卡并行
# 要实现模型单机多卡十分容易,直接使用 `new_module = nn.DataParallel(module, device_ids)`, 默认会把模型分布到所有的卡上。多卡并行的机制如下:
# - 将模型(module)复制到每一张卡上
# - 将形状为(N,C,H,W)的输入均等分为 n份(假设有n张卡),每一份形状是(N/n, C,H,W),然后在每张卡前向传播,反向传播,梯度求平均。要求batch-size 大于等于卡的个数(N>=n)
#
# 在绝大多数情况下,new_module的用法和module一致,除了极其特殊的情况下(RNN中的PackedSequence)。另外想要获取原始的单卡模型,需要通过`new_module.module`访问。
# #### 5.4.2 多机分布式
# ### 5.5 持久化
# 在PyTorch中,以下对象可以持久化到硬盘,并能通过相应的方法加载到内存中:
# - Tensor
# - Variable
# - nn.Module
# - Optimizer
#
# 本质上上述这些信息最终都是保存成Tensor。Tensor的保存和加载十分的简单,使用t.save和t.load即可完成相应的功能。在save/load时可指定使用的pickle模块,在load时还可将GPU tensor映射到CPU或其它GPU上。
#
# 我们可以通过`t.save(obj, file_name)`等方法保存任意可序列化的对象,然后通过`obj = t.load(file_name)`方法加载保存的数据。对于Module和Optimizer对象,这里建议保存对应的`state_dict`,而不是直接保存整个Module/Optimizer对象。Optimizer对象保存的主要是参数,以及动量信息,通过加载之前的动量信息,能够有效地减少模型震荡,下面举例说明。
#
a = t.Tensor(3, 4)
if t.cuda.is_available():
a = a.cuda(1) # 把a转为GPU1上的tensor,
t.save(a,'a.pth')
# 加载为b, 存储于GPU1上(因为保存时tensor就在GPU1上)
b = t.load('a.pth')
# 加载为c, 存储于CPU
c = t.load('a.pth', map_location=lambda storage, loc: storage)
# 加载为d, 存储于GPU0上
d = t.load('a.pth', map_location={'cuda:1':'cuda:0'})
t.set_default_tensor_type('torch.FloatTensor')
from torchvision.models import SqueezeNet
model = SqueezeNet()
# module的state_dict是一个字典
model.state_dict().keys()
# Module对象的保存与加载
t.save(model.state_dict(), 'squeezenet.pth')
model.load_state_dict(t.load('squeezenet.pth'))
optimizer = t.optim.Adam(model.parameters(), lr=0.1)
t.save(optimizer.state_dict(), 'optimizer.pth')
optimizer.load_state_dict(t.load('optimizer.pth'))
all_data = dict(
optimizer = optimizer.state_dict(),
model = model.state_dict(),
info = u'模型和优化器的所有参数'
)
t.save(all_data, 'all.pth')
all_data = t.load('all.pth')
all_data.keys()
| chapter05-utilities/chapter5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project Scenario
# You're a data scientist in an epidemiology department. <br>
#
# The government is waging a war on diabetes, and you're at the frontline. Your weapon is your Python skills, and your bullets are data.
#
# In this project, train a machine learning model to predict whether an individual is at risk of getting diabetes.
#
# # Parts
# - Part I: Data collection and cleaning
# - Part II: Data visualisation and statistics
# - Part III: Machine Learning model training
#
# # Part I: Data collection and cleaning
# Get data from [UCI Machine Learning Repository](https://archive.ics.uci.edu/ml/datasets/Early+stage+diabetes+risk+prediction+dataset.)
import pandas as pd
# read CSV into DataFrame
df = pd.read_csv("../Dataset/diabetes_data_upload.csv")
df
# +
# replace values in DataFrame with .replace
df = df.replace("No", 0)
df = df.replace("Yes", 1)
df = df.replace("Positive", 1)
df = df.replace("Negative", 0)
# consider Gender column as isMale
df = df.replace("Male", 1)
df = df.replace("Female", 0)
df
# -
# check for missing values
df.isnull().sum()
# check dtypes of columns
# as long as int or float, good
df.dtypes
# +
replace = {"Gender": "ismale"}
df = df.rename(columns=replace)
# lowercase everything in columns
df.columns.str.lower()
# -
# lowercase everything in columns
df.columns = df.columns.str.lower()
df
# export DataFrame to CSV
df.to_csv("../Dataset/diabetes_data_clean.csv", index=None)
pd.read_csv("../Dataset/diabetes_data_clean.csv")
# # Summary
# 1. Collect data from UCI Repository
# 2. Replaced strings to 1s and 0s
# 3. Replaced changed column name
# 4. Lowercased everything in columns
# 5. Exported clean DataFrame to new CSV
| Notebooks/Project Early Diabetes Detection Part I.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''base'': conda)'
# name: python395jvsc74a57bd0c480343c6863bdb6a59057b7d5b44ce36b9db870283dc107bcd3b770ee670e9a
# ---
import engine
import feature
import display
import evaluation
# 首先指定圖庫路徑,並且載入想要搜索的圖片,以及圖片標記。
data = engine.data(folder='resource/jpg/data/')
# data.load(path='resource/jpg/quiz/seafood/8.jpg', label='seafood')
# data.load(path='_query/cat_8.jpg', label='cat')
# data.load(path='_query/dog_8.jpg', label='dog')
# data.load(path='_query/soup_7.jpg', label='soup')
data.load(path='_query/meat_8.jpg', label='meat')
data.table.head()
# 第一筆資料代表我想檢索的圖片,後續則是待會會進行比對的圖庫資料。
comparsion = {}
# + active=""
# 使用 rgb 以及 gray 的特徵來進行比對,比對長條圖的方法有 'L2 norm' 、 'Chi-square' 、 'Correlation' 、 'intersection' 。
# -
##
color = feature.color(table=data.table, use='rgb')
comparsion['rgb l2'] = color.compare(method='l2')
comparsion['rgb chisquare'] = color.compare(method='chisquare')
comparsion['rgb correlation'] = color.compare(method='correlation')
comparsion['rgb intersection'] = color.compare(method='intersection')
##
color = feature.color(table=data.table, use='gray')
comparsion['gray l2'] = color.compare(method='l2')
comparsion['gray chisquare'] = color.compare(method='chisquare')
comparsion['gray correlation'] = color.compare(method='correlation')
comparsion['gray intersection'] = color.compare(method='intersection')
# 使用 sobel 以及 canny 的方法來提取 texture/edge 特徵來進行比對,比對長條圖的方法有 'L2 norm' 、 'Chi-square' 、 'Correlation' 、 'intersection' 。
##
texture = feature.texture(table=data.table, use="sobel(xy)")
comparsion['sobel(xy) l2'] = texture.compare(method='l2')
comparsion['sobel(xy) chisquare'] = texture.compare(method='chisquare')
comparsion['sobel(xy) correlation'] = texture.compare(method='correlation')
comparsion['sobel(xy) intersection'] = texture.compare(method='intersection')
##
texture = feature.texture(table=data.table, use="canny")
comparsion['canny l2'] = texture.compare(method='l2')
comparsion['canny chisquare'] = texture.compare(method='chisquare')
comparsion['canny correlation'] = texture.compare(method='correlation')
comparsion['canny intersection'] = texture.compare(method='intersection')
# 以 ROC 曲線來評估每個搜索的方法。
display.auc(comparsion, save="LOG/auc plot.png")
# 設定 threshold ,可以針對每個方法計算對應的 recall 以及 precision 分數。
summary = {}
summary['rgb l2'] = evaluation.summarize(comparsion['rgb l2'], threshold=0.5, method='rgb l2')
summary['rgb chisquare'] = evaluation.summarize(comparsion['rgb chisquare'], threshold=0.5, method='rgb chisquare')
summary['rgb correlation'] = evaluation.summarize(comparsion['rgb correlation'], threshold=0.5, method='rgb correlation')
summary['rgb intersection'] = evaluation.summarize(comparsion['rgb intersection'], threshold=0.5, method='rgb intersection')
summary['gray l2'] = evaluation.summarize(comparsion['gray l2'], threshold=0.5, method='gray l2')
summary['gray chisquare'] = evaluation.summarize(comparsion['gray chisquare'], threshold=0.5, method='gray chisquare')
summary['gray correlation'] = evaluation.summarize(comparsion['gray correlation'], threshold=0.5, method='gray correlation')
summary['gray intersection'] = evaluation.summarize(comparsion['gray intersection'], threshold=0.5, method='gray intersection')
summary['sobel(xy) l2'] = evaluation.summarize(comparsion['sobel(xy) l2'], threshold=0.5, method='sobel(xy) l2')
summary['sobel(xy) chisquare'] = evaluation.summarize(comparsion['sobel(xy) chisquare'], threshold=0.5, method='sobel(xy) chisquare')
summary['sobel(xy) correlation'] = evaluation.summarize(comparsion['sobel(xy) correlation'], threshold=0.5, method='sobel(xy) correlation')
summary['sobel(xy) intersection'] = evaluation.summarize(comparsion['sobel(xy) intersection'], threshold=0.5, method='sobel(xy) intersection')
summary['canny l2'] = evaluation.summarize(comparsion['canny l2'], threshold=0.5, method='canny l2')
summary['canny chisquare'] = evaluation.summarize(comparsion['canny chisquare'], threshold=0.5, method='canny chisquare')
summary['canny correlation'] = evaluation.summarize(comparsion['canny correlation'], threshold=0.5, method='canny correlation')
summary['canny intersection'] = evaluation.summarize(comparsion['canny intersection'], threshold=0.5, method='canny intersection')
report = evaluation.report(summary)
report.to_csv('LOG/report summary.csv', index=False)
report
# 顯示一個搜尋的結果,根據相似程度由高到低進行排序,設定參數 number 可以顯示圖片的數量。
display.search(table=comparsion['rgb l2'], number=6, save='LOG/cache.png')
comparsion['rgb l2'].to_csv("LOG/BetterLog.csv", index=False)
| 1/demonstration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
from upath import UPath
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### local filesystem
#
# If you give a local path, `UPath` defaults to `pathlib.PosixPath` or `pathlib.WindowsPath`
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
local_path = UPath('/tmp')
local_path
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# If you give it a scheme registered with fsspec, it will return a UPath which uses fsspec FileSystem backend
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
local_upath = UPath('file:/tmp')
local_upath
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# ### fsspec filesystems
#
# with `UPath` you can connect to any fsspec FileSystem and interact with it in with it as you would with your local filesystem using pathlib. Connection arguments can be given in a couple of ways:
#
# You can give them as keyword arguments as described for each filesystem in the fsspec docs:
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
ghpath = UPath('github:/', org='fsspec', repo='universal_pathlib', sha='main')
# + [markdown] ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# or define them in the path/url, in which case they will be appropriately parsed:
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
ghpath = UPath('github://fsspec:universal_pathlib@main/')
ghpath
# -
# With a `UPath` object instantiated, you can now interact with the paths with the usual `pathlib.Path` API
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
for p in ghpath.iterdir():
print(p)
# -
# The `glob` method is also available for most filesystems. Note the syntax here is as defined in `fsspec`, rather than that of pathlib.
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
for p in ghpath.glob('**.py'):
print(p)
# -
# All the standard path methods and attributes of `pathlib.Path` are available too:
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
readme_path = ghpath / 'README.md'
readme_path
# -
# To get the full path as a string use:
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
str(readme_path)
# -
# You can also use the path attribute to get just the path:
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
# path attribute added
readme_path.path
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
readme_path.name
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
readme_path.stem
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
readme_path.suffix
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
readme_path.exists()
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
readme_path.read_text()[:19]
# -
# Some filesystems may require extra imports to use.
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
import s3fs
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
s3path = UPath("s3://spacenet-dataset")
# + ein.hycell=false ein.tags="worksheet-0" slideshow={"slide_type": "-"}
for p in s3path.iterdir():
print(p)
# -
# You can chain paths with the `/` operator and read text or binary contents.
(s3path / "LICENSE.md").read_text()
with (s3path / "LICENSE.md").open("rt", encoding="utf-8") as f:
print(f.read(22))
# Globbing also works for many filesystems.
from itertools import islice
for p in islice((s3path / "AOIs" / "AOI_3_Paris").glob("**.TIF"), 5):
print(p)
| notebooks/examples.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="FHoH-r-fUiKq"
from __future__ import absolute_import, division, print_function
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import numpy as np
import matplotlib.pyplot as plt
# + colab={} colab_type="code" id="a1uFgkbrUjZY"
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = fashion_mnist.load_data()
# + colab={} colab_type="code" id="QQ1w29lc0Dak"
# print(np.min(train_images[0]))
# print(np.max(train_images[0]))
# + colab={} colab_type="code" id="BbcsAWVVUsqW"
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# + colab={} colab_type="code" id="YMrRq-yNV6bP"
# train_images.shape
# + colab={} colab_type="code" id="fjubKHOZV9U9"
# len(train_labels)
# + colab={} colab_type="code" id="VBCXhh5AV_kD"
# train_labels
# + colab={} colab_type="code" id="_DoqerPeWFAJ"
# test_images.shape
# + colab={} colab_type="code" id="Hn9EomDdWK_2"
# len(test_labels)
# + colab={} colab_type="code" id="D37qfihCWNLL"
# plt.imshow(test_images[0])
# plt.colorbar()
# + colab={} colab_type="code" id="1VBvJP98WRUV"
train_images = train_images / 255.0
test_images = test_images / 255.0
# + colab={} colab_type="code" id="Nl8y6SaQWZAN"
# plt.figure(figsize=(10,10))
# for i in range(25):
# plt.subplot(5,5,i+1)
# plt.xticks([])
# plt.yticks([])
# plt.grid(False)
# plt.imshow(train_images[i], cmap=plt.cm.binary)
# plt.xlabel(class_names[train_labels[i]])
# plt.show()
# + colab={} colab_type="code" id="_EcnZaDXWb0x"
model = keras.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(128, activation=tf.nn.relu),
keras.layers.Dense(10, activation=tf.nn.softmax)
])
# + colab={} colab_type="code" id="AnBMnmc6Wnqw"
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + colab={"base_uri": "https://localhost:8080/", "height": 208} colab_type="code" id="-cNOp6EnWw5m" outputId="96f7357d-1671-41c2-b5ce-e1970f8715a2"
model.fit(train_images, train_labels, epochs=5)
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="Tohjs7C1WzcJ" outputId="f6461cf7-2937-49aa-c8e9-739f58d651a5"
test_loss, test_acc = model.evaluate(test_images, test_labels)
print('Test accuracy:', test_acc)
# + colab={} colab_type="code" id="Gin1px8IXBHR"
predictions = model.predict(test_images)
# + colab={"base_uri": "https://localhost:8080/", "height": 69} colab_type="code" id="QnkuMzdOXD1e" outputId="7c88eb95-e0e0-406c-d4ad-2438676f1922"
predictions[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="BdXQjlKMXGFV" outputId="81a9c4f2-05d9-4928-8a41-4815b4670820"
np.argmax(predictions[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="demY47eEXPye" outputId="504efce9-09f5-45e8-8a7d-5b4bf6c4d1db"
test_labels[0]
# + colab={} colab_type="code" id="CwVcaNj2XVK3"
## Data Sc
# def plot_image(i, predictions_array, true_label, img):
# predictions_array, true_label, img = predictions_array[i], true_label[i], img[i]
# plt.grid(False)
# plt.xticks([])
# plt.yticks([])
# plt.imshow(img, cmap=plt.cm.binary)
# predicted_label = np.argmax(predictions_array)
# if predicted_label == true_label:
# color = 'blue'
# else:
# color = 'red'
# plt.xlabel("{} {:2.0f}% ({})".format(class_names[predicted_label],
# 100*np.max(predictions_array),
# class_names[true_label]),
# color=color)
# def plot_value_array(i, predictions_array, true_label):
# predictions_array, true_label = predictions_array[i], true_label[i]
# plt.grid(False)
# plt.xticks([])
# plt.yticks([])
# thisplot = plt.bar(range(10), predictions_array, color="#777777")
# plt.ylim([0, 1])
# predicted_label = np.argmax(predictions_array)
# thisplot[predicted_label].set_color('red')
# thisplot[true_label].set_color('blue')
# + colab={"base_uri": "https://localhost:8080/", "height": 203} colab_type="code" id="_bw41D7uXcrj" outputId="e9a53fe2-9a23-4a87-90a8-de9004e3f6d2"
## Data Sc
# i = 0
# plt.figure(figsize=(6,3))
# plt.subplot(1,2,1)
# plot_image(i, predictions, test_labels, test_images)
# plt.subplot(1,2,2)
# plot_value_array(i, predictions, test_labels)
# plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 203} colab_type="code" id="rxef0vb6XkuA" outputId="f4f086d4-6f1c-4c36-ba67-ab3eea27d3b3"
## Data Sc
# i = 12
# plt.figure(figsize=(6,3))
# plt.subplot(1,2,1)
# plot_image(i, predictions, test_labels, test_images)
# plt.subplot(1,2,2)
# plot_value_array(i, predictions, test_labels)
# plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 589} colab_type="code" id="-6fP5pXKXnXC" outputId="52d1a1f7-a709-4d33-e37d-d9472b1185d8"
# # Plot the first X test images, their predicted label, and the true label
# # Color correct predictions in blue, incorrect predictions in red
# ## Data Sc
# num_rows = 5
# num_cols = 3
# num_images = num_rows*num_cols
# plt.figure(figsize=(2*2*num_cols, 2*num_rows))
# for i in range(num_images):
# plt.subplot(num_rows, 2*num_cols, 2*i+1)
# plot_image(i, predictions, test_labels, test_images)
# plt.subplot(num_rows, 2*num_cols, 2*i+2)
# plot_value_array(i, predictions, test_labels)
# plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="FqdZeErgX7Qr" outputId="7ea225c3-88b5-465a-ff97-8754ec7126ef"
# # Grab an image from the test dataset
# img = test_images[0]
# print(img.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="7SlR2TPaXsr6" outputId="3a945ce9-16b8-4424-9651-fa9b7e33577e"
# # Add the image to a batch where it's the only member.
# img = (np.expand_dims(img,0))
# print(img.shape)
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="au5PW3gkX-1Q" outputId="7f274087-36a8-4ff1-9ab1-f667e8be3b34"
# predictions_single = model.predict(img)
# print(predictions_single)
# + colab={} colab_type="code" id="eGgtU0uRYBhc"
# plot_value_array(0, predictions_single, test_labels)
# _ = plt.xticks(range(10), class_names, rotation=45)
# + colab={} colab_type="code" id="vklFgbGdYEZW"
# np.argmax(predictions_single[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="xj9cRqZFYHRh" outputId="8668f36a-d3b7-48ed-c6a6-f32007998803"
import cv2
gray = cv2.imread("coat.jpg", cv2.IMREAD_GRAYSCALE)
gray = cv2.resize(255-gray, (28, 28))
gray = gray/255
pr = model.predict_classes(gray.reshape(1, 28, 28))
print(pr)
print(class_names[pr[0]])
# + colab={} colab_type="code" id="V194K7_uyjtv"
# plt.imshow(gray)
# plt.colorbar()
# + colab={} colab_type="code" id="bu1Mxo7_zvHf"
fn = "mymodel.h5"
model.save_weights(fn)
| Deep Learning/deep/keras_training_savedata.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Strokes Gained Statistics
# “The strokes gained concept was initially developed by Professor <NAME> of Columbia University, utilizing ShotLink data that has been made available to academic institutions for research since 2007. Strokes gained is a better method for measuring performance because it compares a player's performance to the rest of the field and because it can isolate individual aspects of the game. Traditional golf statistics, such as greens in regulation and putts per green, are influenced by a player's performance on shots other than those being measured.”
#
# (*Taken from an informational web-page of the PGA Tour*)
#
# ## Table of contents <a id="toc"></a>
#
# 1. [Data Wrangling](#wrangling)
# 1. [Load Course Data](#loadcourse)
# 1. [Load Game Data](#loadgame)
# 2. [Strokes Gained Categories](#categories)
# 1. [Base Tables](#basetables)
# 1. [Metrics](#metrics)
# 2. [Preparing Summaries](#summaries)
# 1. [Per Category Summaries](#catsummaries)
# 1. [Category Groups](#catgroups)
# 1. [Classic Statistics](#classics)
# ## Data Wrangling <a name="wrangling"></a>
# (<a href="#toc">back to ToC</a>)
#
import os
import re
import json
import datetime
# ### Load Course Data <a name="loadcourse"></a>
# (<a href="#toc">back to ToC</a>)
#
# Course data consists of meta data describing geo-coordinates and other aspects of every hole on the course.
# +
courses_path = "/Users/npi/prg/python/strokesgained/strogain/static"
course_name = "Open.9"
meta_info_file = "course.json"
course_meta_path = os.path.join(courses_path, course_name, meta_info_file)
with open(course_meta_path, 'r') as f:
course_meta = json.load(f)
# -
# For convenience, we define a function to get the level-par score for a hole. We will use this later to attach the par score to every shot of a game.
# +
def par_for_hole(holeno):
return course_meta['holes'][str(holeno)]['par']
par_for_hole(2)
# -
# ### Load Game Data <a name="loadgame"></a>
# (<a href="#toc">back to ToC</a>)
#
# Load a set of games. Games describe all the shots of a round of golf.
#
# Parameters will be provided by an HTML form, but we use constants for now.
games_path = "/Users/npi/prg/python/strokesgained/instance/games" # substitute this with flask-variables
user = "Norbert"
#date_range = [datetime.date.today()-datetime.timedelta(days = 10), datetime.date.today()]
date_range = [datetime.date.today()-datetime.timedelta(days = 12),
datetime.date.today()-datetime.timedelta(days = 10)]
# We will load all the rounds of golf in a given date range. First prepare some helper functions.
# +
# Check if a date is within a certain range
def check_date_range(date_as_string, date_from, date_to):
d = datetime.datetime.strptime(date_as_string, "%Y-%m-%d").date()
return date_from <= d <= date_to
all_games_rx = re.compile('game-(\d\d\d\d-\d\d-\d\d).json')
# Return a list of all the game filenames for a given date range
def games_in_range(uid, date_from, date_to):
dirname = os.path.join(games_path, uid)
files = os.listdir(dirname)
flist = []
for fname in files:
#print(fname)
m = all_games_rx.search(fname)
if m:
dstr = m.group(1)
#print("date = '%s'" % dstr)
if check_date_range(dstr, date_from, date_to):
flist.append(fname)
return flist
#check_date_range("2020-10-07", datetime.date.today(), datetime.date.today()+datetime.timedelta(days = 3))
games_in_range("Norbert", date_range[0], date_range[1])
# -
# We will process game files one by one, loading their JSON contents. *action* will be a function which takes the JSON as a parameter.
# +
def process_game_files(game_filenames, action, initial, *params):
res = initial
for gf in game_filenames:
m = all_games_rx.search(gf)
game_data_path = os.path.join(games_path, user, gf)
with open(game_data_path, 'r') as f:
game = json.load(f)
print("Processing game %s" % gf)
res = action(game, m.group(1), initial, *params)
return res
#game['game']['1']['pin']
# -
# Game data is collected in a Pandas data frame.
import numpy as np
import pandas as pd
# We define an action function which collects all the strokes in a game and appends them as rows to a data frame.
# +
def load_game_action(game, gdate, initial, *params):
holes = [ holeno for holeno in game['game']]
print("Game has holes", holes)
df = initial
for h in holes:
strokes = game['game'][h]['strokes']
for s in strokes:
row = {
'round': gdate,
'hole': h,
'stroke': int(s['stroke']),
'from': s['from'],
'to': s['to'],
'lie': s['lie'],
'result': s['result'],
'pin': game['game'][h]['pin']
}
#print(row)
df = df.append(row, ignore_index=True)
return df
def load_games_as_dataframe(game_filenames):
df = pd.DataFrame(columns=['round', 'hole', 'stroke', 'from', 'to', 'lie', 'result', 'pin'])
df['stroke'] = df['stroke'].astype(int)
df = process_game_files(game_filenames, load_game_action, df)
return df
game_filenames = games_in_range(user, date_range[0], date_range[1])
df = load_games_as_dataframe(game_filenames)
df.head(7)
# -
# We need to calculate geo-distances. *Strokes gained* is all about getting nearer to the hole.
# +
import geopy.distance
def dist_to_pin(row):
return geopy.distance.distance(row['from'], row['pin']).m
def rest_to_pin(row):
return geopy.distance.distance(row['to'], row['pin']).m
# 'm' returns distance in meters
d = geopy.distance.distance([48.27065440030751, 11.750349998474123], [48.269147574488386, 11.750752329826357]).m
d
# -
# For *strokes gained* calculation we do not need absolute geo-coordinates, but rather distances to the pin.
# +
def calculate_shot_distances(shot_dataframe):
shot_dataframe['from2pin'] = shot_dataframe.apply(dist_to_pin, axis=1)
shot_dataframe['to2pin'] = shot_dataframe.apply(rest_to_pin, axis=1)
shot_dataframe['dist'] = shot_dataframe['from2pin']-shot_dataframe['to2pin']
shot_dataframe = shot_dataframe.drop(columns=['from', 'to', 'pin'])
return shot_dataframe
df = calculate_shot_distances(df)
df.head(7)
# -
# Some calculations will rely on the par score for a hole. We extract it from the course meta data.
# +
def set_par_for_hole(row):
return par_for_hole(row['hole'])
df['par'] = df.apply(set_par_for_hole, axis=1)
df.head(7)
# -
# ## Strokes Gained Categories <a name="categories"></a>
# (<a href="#toc">back to ToC</a>)
#
# Statistics about strokes gained are clustered into shot categories.
#
# A shot's cluster depends its lie and distance to the pin, e.g. an approach shot 146m from the rough is in category "*Appr-150_R*". We need shot categories for the start position of a shot and for the resulting position.
# row is a dataframe row, distance_col selects either start position or resulting position.
# lie is one of H (holed), G (on the green), F (fairway), B (bunker), R (rough), T (tee).
def cat_sg(row, distance_col, lie):
d = row[distance_col]
l = row[lie]
cat = "other"
if l == "H":
cat = "holed"
elif l == "G":
if d <= 1.2:
cat = "Putt-1"
elif d <= 3:
cat = "Putt-3"
else:
cat = "Putt"
elif l == "B":
if d <= 30:
cat = "Bunker-Green"
elif d <= 70:
cat = "Bunker-70"
elif d <= 70:
cat = "Bunker-100"
else:
cat = "Bunker-150"
elif l == "F" or l == "R":
if d <= 20:
cat = "Short-20"
elif d <= 40:
cat = "Short-40_" + l
elif d <= 70:
cat = "Appr-70_" + l
elif d <= 110:
cat = "Appr-110_" + l
elif d <= 150:
cat = "Appr-150_" + l
elif d <= 180:
cat = "Appr-180_" + l
else:
cat = "Appr-long"
elif l == "T":
if d <= 200:
cat = "Tee-200"
else:
cat = "Tee-400"
return cat
# Now we're ready to fully categorize each shot.
# +
df['sg_cat_from'] = df.apply(lambda r: cat_sg(r, 'from2pin', 'lie'), axis=1)
df['sg_cat_to'] = df.apply(lambda r: cat_sg(r, 'to2pin', 'result'), axis=1)
df.head(7)
# -
# ### Strokes Gained Base Tables <a name="basetables"></a>
# (<a href="#toc">back to ToC</a>)
#
# Strokes gained calculations are based on empirical data, gathered by the PGA Tour via its *Shot Link* programme. <NAME> published them in his book, where from the tables below have been extracted.
# Strokes gained tables for (male) PGA tour pros
#sg_table_pros_json = """
sg_table_pros = {
"holed": 0,
"Putt-1": 1.04,
"Putt-3": 1.42,
"Putt-10": 1.87,
"Putt": 2.4,
"Short-20": 2.6,
"Short-40_F": 2.65,
"Short-40_R": 2.8,
"Appr-70_F": 2.75,
"Appr-70_R": 2.96,
"Appr-110_F": 2.85,
"Appr-110_R": 3.08,
"Appr-150_F": 2.98,
"Appr-150_R": 3.23,
"Appr-180_F": 3.19,
"Appr-180_R": 3.42,
"Appr-long": 3.6,
"Bunker-Green": 2.82,
"Bunker-70": 3.2,
"Bunker-110": 3.25,
"Bunker-150": 3.3,
"Tee-200": 3.17,
"Tee-400": 4.08
}
#"""
#sg_table_pros = json.loads(sg_table_pros_json)
# +
# Derived strokes gained tables for 18 handicaps
#sg18_table_json = """
sg18_table = {
"holed": 0,
"Putt-1": 1.1,
"Putt-3": 1.8,
"Putt-10": 2.19,
"Putt": 2.65,
"Short-20": 2.75,
"Short-40_F": 2.9,
"Short-40_R": 3,
"Appr-70_F": 3,
"Appr-70_R": 3.17,
"Appr-110_F": 3.15,
"Appr-110_R": 3.31,
"Appr-150_F": 3.34,
"Appr-150_R": 3.44,
"Appr-180_F": 3.58,
"Appr-180_R": 3.71,
"Appr-long": 3.89,
"Bunker-Green": 3.15,
"Bunker-70": 3.41,
"Bunker-110": 3.48,
"Bunker-150": 3.51,
"Tee-200": 3.9,
"Tee-400": 5.2
}
#"""
#sg18_table = json.loads(sg18_table_json)
#def sg(row):
# sg_f = sg18_table[row['sg_cat_from']]
# sg_t = sg18_table[row['sg_cat_to']]
# return sg_f - sg_t - 1
#df['sg'] = df.apply(sg, axis=1)
# -
# For bookkeeping and UI-output we declare a natural order for shot categories.
# +
#sg_target_sort = """
sg_sort = {
"holed": 0,
"Putt-1": 1,
"Putt-3": 2,
"Putt-10": 3,
"Putt": 4,
"Short-20": 5,
"Short-40_F": 6,
"Short-40_R": 7,
"Appr-70_F": 8,
"Appr-110_F": 9,
"Appr-150_F": 10,
"Appr-180_F": 11,
"Appr-70_R": 12,
"Appr-110_R": 13,
"Appr-150_R": 14,
"Appr-180_R": 15,
"Appr-long": 16,
"Bunker-Green": 17,
"Bunker-70": 18,
"Bunker-110": 19,
"Bunker-150": 20,
"Tee-200": 21,
"Tee-400": 22
}
#"""
#sg_sort = json.loads(sg_target_sort)
def sg_cat_key(elem):
return sg_sort[elem[0]]
print(sg_cat_key(('Appr-150_F', 0)))
# -
#sg_official_name = """
sg_official = {
"holed": "",
"Putt-1": "Putt ≤ 1m",
"Putt-3": "Putt ≤ 3m",
"Putt-10": "Putt ≤ 10m",
"Putt": "Distance Putt",
"Short-20": "Around Green ≤ 20m",
"Short-40_F": "Around Green ≤ 40m",
"Short-40_R": "Around Green ≤ 40m from Rough",
"Appr-70_F": "Approach ≤ 70m from Fairway",
"Appr-110_F": "Approach ≤ 110m from Fairway",
"Appr-150_F": "Approach ≤ 150m from Fairway",
"Appr-180_F": "Approach ≤ 180m from Fairway",
"Appr-70_R": "Approach ≤ 70m from Rough",
"Appr-110_R": "Approach ≤ 110m from Rough",
"Appr-150_R": "Approach ≤ 150m from Rough",
"Appr-180_R": "Approach ≤ 180m from Rough",
"Appr-long": "Long Approach",
"Bunker-Green": "Greenside Bunker",
"Bunker-70": "Long Bunker Shot",
"Bunker-110": "Short Fairway Bunker Shot",
"Bunker-150": "Long Fairway Bunker Shot",
"Tee-200": "",
"Tee-400": "Tee Shot"
}
#"""
#sg_official = json.loads(sg_official_name)
# There will be situations where we need an ordered list/tuple of shot categories. This function will prepare one.
# +
#def sg_cat_inx(elem):
# return sg_sort[elem[1]]
# Return a tuple of names and a tuple of sequence numbers
def make_category_index():
order = sorted(sg_sort.items(), key=sg_cat_key)
cat_names_ordered, cat_index = zip(*order)
return cat_names_ordered, cat_index
#make_category_index()[0] # show the names tuple
# -
# ### Strokes Gained Metrics <a name="metrics"></a>
# (<a href="#toc">back to ToC</a>)
#
# For easier handling we load the strokes gained base metrics into a data frame as well.
# +
# Get pro's and am's strokes-gained data ordered
def extract_pro_and_am_strokes_gained():
pro_data_tuples = sorted(sg_table_pros.items(), key=sg_cat_key)
_, pro_strokes_gained_list = zip(*pro_data_tuples) # unpack a list of pairs into two tuples
am_data_tuples = sorted(sg18_table.items(), key=sg_cat_key)
_, am_strokes_gained_list = zip(*am_data_tuples)
return pro_strokes_gained_list, am_strokes_gained_list
#extract_pro_and_am_strokes_gained()
# -
# With ordered data points at hand, now we're able to pull the strokes-gained baseline data as a data frame.
# +
def get_strokes_gained_base_data_frame():
pro_strokes_gained_list, am_strokes_gained_list = extract_pro_and_am_strokes_gained()
pros = pd.Series(pro_strokes_gained_list)
ams = pd.Series(am_strokes_gained_list)
cats, inx = make_category_index()
#cats = pd.Series(pro_x)
sgdf = pd.DataFrame({
'Category': cats,
'Seq': inx,
'Pro': pros,
'Am18': ams
}).set_index('Category')
return sgdf
sgdf = get_strokes_gained_base_data_frame()
sgdf.head()
# -
# Let's take a quick detour and visualize the strokes gained base tables and discuss implications.
# +
cats, inx = make_category_index()
p = sgdf.plot.line(y=['Pro', 'Am18'], grid=True, rot=90, xticks=inx, figsize=(5,3))
p.set_xticklabels(cats);
p
# -
# The gap between the two lines visualizes the proficiency difference between a PGA tour pro and a 18-handicap. The values for the 18-hcp are a result of some educated guesswork and measuring my own performance (as an 18-hcp).
#
# For different handicaps, our task will be to interpolate between (or extrapolate) the two curves. Let's define an interpolations function as a helper.
# +
def interp(f, intv):
return f*(intv[1]-intv[0])+intv[0]
print("1/3[1,2] = %.2f" % interp(1/3, (1, 2)))
# -
# Now we're able to calculate a strokes-gained target for a given category and handicap.
# +
def hcp_sg(hcp, cat):
cat_row = sgdf.loc[cat]
sg = interp(hcp/18, (cat_row['Pro'], cat_row['Am18']))
#print(cat_row)
return sg
print("Hcp 18 target for short game = %.2f" % hcp_sg(18, 'Short-20'))
print("Hcp 10 target for long approaches = %.2f" % hcp_sg(10, 'Appr-long'))
# -
# Tee-shots are only slightly more complicated.
# +
def teeshot_sg(hcp, distance):
t200 = hcp_sg(hcp, 'Tee-200')
t400 = hcp_sg(hcp, 'Tee-400')
#print("%.2f / %.2f" % (t200, t400))
return interp((distance-200)/200, (t200, t400))
#print(teeshot_sg(0, 400))
#print(teeshot_sg(0, 300))
#print(teeshot_sg(18, 400))
#print(teeshot_sg(18, 366))
# -
# Now we include columns for SG-0 and SG-18 in our data frame.
# +
def insert_SG_for_hcps(hcp, df):
def local_sg(row):
cat = row['sg_cat_from']
mysg = hcp_sg(hcp, cat)
if cat == 'tee-200' or cat == 'Tee-400':
#print("interpolating SG for tee shot")
mysg = teeshot_sg(hcp, row['from2pin'])
#print("my sg = %.2f" % mysg)
result_sg = hcp_sg(hcp, row['sg_cat_to'])
#print("-> sg = %.2f" % result_sg)
return mysg - result_sg - 1
df['SG-'+str(hcp)] = df.apply(local_sg, axis=1)
#return local_sg(row)
return df
#print(df.iloc[0])
#print("\nstrokes gained for tee shot = %.2f" % sg_for_hcp(12, df.iloc[0]))
df = insert_SG_for_hcps(0, df)
df = insert_SG_for_hcps(18, df)
df.head()
# -
# Finally, we need to fix one small thing: For analytical purposes tee-shots for par-3s fall into shot categories of approach shots from the fairway.
# +
def fix_teeshot_cat(row):
cat_from = row['sg_cat_from']
if row['lie'] == "T" and row['par'] == 3:
row['lie'] = "F"
cat_from = cat_sg(row, 'from2pin', 'lie')
print(cat_from)
return cat_from
df['sg_category'] = df.apply(fix_teeshot_cat, axis=1)
df.head()
# -
# ## Preparing Summaries <a name="summaries"></a>
# (<a href="#toc">back to ToC</a>)
#
# Taking a look at individual shots is great, but users need summary information on what to work on with their game.
#
#
# ### Per Category Summaries <a name="catsummaries"></a>
# (<a href="#toc">back to ToC</a>)
#
# First we extract a condensed data frame.
dfx = df[['sg_category', 'SG-0', 'SG-18']]
dfx.head()
# The first statistic of interest will be the mean SG-value for every shot category. *pivot_table*( ) uses mean-aggregation by default.
# +
# https://pandas.pydata.org/pandas-docs/stable/user_guide/reshaping.html
#df_sg_means = dfx.pivot(columns='sg_category', values=['SG-0', 'SG-18']).mean()
df_sg_means = pd.pivot_table(dfx, index='sg_category')
df_sg_means.head()
# -
m0 = df_sg_means #['SG-0']
m0 = m0.reindex(cats).drop(['holed', 'Tee-200'])
SG_details_table = pd.DataFrame(m0)
#SG_details_table.columns = ["SG"]
SG_details_table.head()
# +
SG_details_table['label'] = SG_details_table.apply(lambda row: sg_official[row.name], axis=1)
SG_details_table.head()
# -
# For HTML output we need a way to iterate over rows by means of the Jinja template engine.
SG_details_table.to_dict(orient='records')[0:5] # create an ordered list of dicts, iteratable
# ### Category Groups <a name="catgroups"></a>
# (<a href="#toc">back to ToC</a>)
#
# The PGA clusters strokes gained categories into clusters for easier communication.
# +
category_groups = {
"holed": '',
"Putt-1": 'Putting',
"Putt-3": 'Putting',
"Putt-10": 'Putting',
"Putt": 'Putting',
"Short-20": 'Around the Green',
"Short-40_F": 'Around the Green',
"Short-40_R": 'Around the Green',
"Appr-70_F": 'Approach',
"Appr-110_F": 'Approach',
"Appr-150_F": 'Approach',
"Appr-180_F": 'Approach',
"Appr-70_R": 'Approach',
"Appr-110_R": 'Approach',
"Appr-150_R": 'Approach',
"Appr-180_R": 'Approach',
"Appr-long": 'Approach',
"Bunker-Green": 'Around the Green',
"Bunker-70": 'Around the Green',
"Bunker-110": 'Approach',
"Bunker-150": 'Approach',
"Tee-200": '',
"Tee-400": 'Driving'
}
cat_group_names = ["Putting","Around the Green","Approach","Driving"]
# -
# We attach the category group label to each row / shot.
# +
df['cat_group'] = df['sg_category'].apply(lambda x: category_groups[x]).astype("category")
df['cat_group'].cat.set_categories(cat_group_names,inplace=True)
dfy = df[['cat_group', 'SG-0', 'SG-18']]
dfy.head()
# -
# Now we're ready to aggregate SG statistics per category group.
# +
group_SG_stats = pd.pivot_table(dfy, index='cat_group')
group_SG_stats.head()
# -
# As explained above Jinja templates must be able to iterate over the table.
# +
group_SG_stats['group_name'] = group_SG_stats.index #cat_group_names
group_SG_stats.to_dict(orient='records')
# -
# ### Classic Statistics <a name="classics"></a>
# (<a href="#toc">back to ToC</a>)
# **Driving Distance** is defined by <NAME> as the 75th percentile of all non-par-3 tee shots. We will be a bit more optimistic and use the 80th percentile.
# +
drives = df[['sg_category', 'dist']].where(df['sg_category'] == 'Tee-400', axis=0)
driving_distance = drives.quantile(.8, axis=0)
print(type(driving_distance[0]))
print("Driving Distance = %.2f" % driving_distance[0])
# -
# Another classic is **Greens in Regulation**. This is a bit more involved.
#
# First filter for shots to the green or into the hole.
# +
greens = df[['round', 'hole', 'stroke', 'par', 'result']]
ongreen = greens['result']=="G"
inhole = greens['result']=="H" # include hole-in-ones and eagles
greens = greens.where(ongreen | inhole)
greens.loc[greens['stroke'].notna()].head(3)
# -
# We group by (round, hole) and take the shot with the lowest stroke number (= *np.min*). This gives us the first shot per hole to land on the green (or in the hole). In the next step we can compare this shot to par.
# +
gir = pd.pivot_table(greens, index=['round', 'hole'], values=['stroke', 'par'], aggfunc=np.min)
gir['stroke'] = gir['stroke'].astype(int)
gir['par'] = gir['par'].astype(int)
print(gir.dtypes)
gir.head(3)
# -
# Now we introduce the *GiR* metric. It is >= 0 for a "Green in Regulation" and negativ otherwise.
# +
gir['GiR'] = gir['par'] - gir['stroke'] - 2
gir = gir.drop(['stroke'], axis=1)
gir.head()
# -
# As usual we want to be able to export it as a dict for the UI. This is intended for an overview of a round of golf, flagging GiR per hole.
gir['hole'] = gir.index
gir.to_dict(orient='records')
# We're left with one small task: calculate the overall percentage for GiR.
# +
GiR_percentage = len(gir[gir['GiR'] >= 0].index)/(len(gir.index))*100.0
print("Overall GiR = %.2f %%" % GiR_percentage)
# -
# The last classic statistic we'll calulate is **Fairways hit**.
# +
fw = df[['round', 'hole', 'stroke', 'sg_category', 'result']]
onfw = fw['result']=="F"
istee = fw['sg_category']=="Tee-400"
fw = fw.where(istee)
teeshot_count = len(fw.loc[fw['result'].notna()])
fw = fw.where(onfw)
fw_count = len(fw.loc[fw['result'].notna()])
fw_hit = fw_count/teeshot_count*100.0
print("Overall count of long tee-shots = %d" % teeshot_count)
print("Overall FW = %.2f %%" % fw_hit)
fw.loc[fw['result'].notna()].head(3)
# -
# Now let's construct a table showing all the non-par-3 holes where the *1st* tee shot ended up on the fairway. We have to eliminate all tee shots with s shot index > 1. The data could include tee shots e.g. as 3rd shots (onto the fairway) after the first tee shot ended up OB (resulting in 2 tee shots for the same hole). We count these "reloaded" shots for our overall statistics, as they help us asses our driving capabilities, but we won't display them for course summaries.
# +
fw = fw.where(fw['stroke']< 1.5) # eliminate tee-shots after first one has been a miss
fw = fw[['round', 'hole', 'result']]
fw = fw.loc[fw['result'].notna()] # finally drop NaNs
fw.head(3)
# -
# As usual we want to be able to export it as a dict for the UI. This is intended for an overview of a round of golf, flagging *Fairways hit* per hole.
fw.to_dict(orient='records')
# This is a function to calculate all the classic statistics in one go. There is nothing new here, just collecting the code snippets above.
# +
def classic_stats(df):
drives = df[['sg_category', 'dist']].where(df['sg_category'] == 'Tee-400', axis=0)
driving_distance = drives.quantile(.8, axis=0)[0]
greens = df[['round', 'hole', 'stroke', 'par', 'result']]
ongreen = greens['result']=="G"
inhole = greens['result']=="H"
greens = greens.where(ongreen | inhole)
gir = pd.pivot_table(greens, index=['round', 'hole'], values=['stroke', 'par'], aggfunc=np.min)
gir['GiR'] = gir['par'] - gir['stroke'] - 2
gir = gir.drop(['stroke'], axis=1)
gir['hole'] = gir.index
gir_dict = gir.to_dict(orient='records')
GiR_percentage = len(gir[gir['GiR'] >= 0].index)/(len(gir.index))*100.0
fw = df[['round', 'hole', 'stroke', 'sg_category', 'result']]
onfw = fw['result']=="F"
istee = fw['sg_category']=="Tee-400"
fw = fw.where(istee)
teeshot_count = len(fw.loc[fw['result'].notna()])
fw = fw.where(onfw)
fw_count = len(fw.loc[fw['result'].notna()])
fw_hit = fw_count/teeshot_count*100.0
fw = fw.where(fw['stroke']< 1.5)
fw = fw[['round', 'hole', 'result']]
fw = fw.loc[fw['result'].notna()]
fw_dict = fw.to_dict(orient='records')
stats = {
'driving_distance': driving_distance,
'GiR': gir_dict,
'GiR_pcnt': GiR_percentage,
'FW_hit': fw_dict,
'FW_pcnt': fw_hit
}
return stats
classic_stats(df)
# -
h = df.loc[df['result']=="H"][['hole','par','stroke']]
h = pd.pivot_table(h, index=["par"])
h['par'] = h.index
h.to_dict(orient="record")
| notebooks/SG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Intro
#
# This is a demo of the paper Teachable Reinforcement Learning via Advice Distillation showing how humans can use advice to coach agents through new tasks!
#
# For more details, check out our NeurIPS paper and video: https://neurips.cc/Conferences/2021/Schedule?showEvent=27834
# # Setup - do this once
#
# To avoid version conflicts, we recommend running this in a conda env with python 3.7.
#
# conda create --name teachable_rl python=3.7
# conda activate teachable_rl
# pip install notebook
#
# You either need to run this on a device with a display. If you're running on a machine without one, use port forwarding:
#
# ssh -L 9999:localhost:9999 INSERT_SERVER_NAME
# jupyter notebook --no-browser --port 9999
#
#
# We use two environments: [BabyAI](https://github.com/mila-iqia/babyai) and [AntMaze](https://github.com/rail-berkeley/d4rl). If you would like to use AntMaze, please [install Mujoco](https://github.com/openai/mujoco-py).
# + tags=["hide-input"]
# !git clone https://github.com/aliengirlliv/teachable 1> /dev/null
# + tags=["hide-input"]
# cd teachable
# + tags=["hide-input"]
# !pip install -r reqs.txt 1> /dev/null
# -
# cd ..
# # Setup - Do this each time you reload the notebook
# %matplotlib tk
# cd teachable
from final_demo import *
from IPython.display import HTML
import pathlib
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
# # Setup
#
# ## Instructions
# 1. Select the collection mode.
# - "Advice" runs the "Improvement" phase of our method, allowing you to coach an agent using waypoint advice
# - "Demos" lets you collect trajectories providing actions each timestep.
# 2. Select a save name (any string describing this experiment).
# 3. Collect demos below
#
# # Collection
# To collect data, run the block below. A window will open which lets you collect data.
#
# In our human exps, we found you can reach okay performance (> 50% success) on this env with about 30 mins of human collection time.
#
# ## Task
#
# **BabyAI**: The agent's task is to unlock a door by collecting a matching-colored key and using it on the corresponding door. (To speed up training time, we always spawn the agent in the same room as the locked door, and the key is always in the same spot.)
#
# **Ant**: The agent's task is to reach the pink target.
#
# ### Using Advice
#
# Note: Collecting advice requires a mouse.
#
# **BabyAI**: The agent you will be coaching has been pretrained to understand Waypoint advice. It has never seen this particular environment/task before, and has never had to unlock a door. Click on a square to tell the agent to head there and manipulate any item present. Use the scrollwheel to advance.
#
# **Ant**: The agent you will be coaching has been pretrained to understand Waypoint advice. It has never seen an environment this large before. Click on a point to tell the agent to head there. Use the scrollwheel to advance.
#
# ### Providing Demos (BabyAI env only)
# Use the arrow keys to navigate, Page Up/Down to manipulate objects, and Space to open doors.
#
# ## Using Pre-collected data
# We include a buffer of data collected using 30 mins of human time using Advice. You can only load this data if you have CUDA enabled.
env_type = 'Ant' # Options are 'BabyAI', 'Ant'
collect_type = 'Advice' # Options are 'Advice', 'Demos', or 'Precollected'
save_path = 'temp3' # Any string
collector = HumanFeedback(env_type=env_type, collect_type=collect_type,
save_path=save_path, seed=124)
# # Train
#
# Here, we train an advice-free policy on the collected trajectories using the buffer of collected trajectories.
#
#
# It will train for 20 itrs, but feel free to pause it before then if you'd like to see thet trained model.
args = make_args(collector, save_path)
run_experiment(args)
# # Visualize
#
# Play a video of the agent you trained. This agent was trained using the coached rollouts you provided. This agent does **not** receive advice.
display_trained_model(save_path)
# Plot the agent's success rate during training.
plot(save_path)
| FinalDemo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
sys.path.append('../../../GraphGallery/')
sys.path.append('../../../GraphAdv/')
from graphgallery.nn.models import GCN
from graphadv.attack.untargeted import FGSM
import matplotlib.pyplot as plt
# -
from graphgallery.data import NPZDataset
data = NPZDataset('citeseer', root="~/GraphData/datasets/", verbose=False, standardize=True)
adj, x, labels = data.graph.unpack()
idx_train, idx_val, idx_test = data.split(random_state=15)
attacker = FGSM(adj, x, labels, idx_train, device="GPU", seed=None, surrogate_args={'idx_val':idx_val})
attacker.reset()
attacker.attack(0.05, structure_attack=True, feature_attack=False)
# show logs
attacker.show_edge_flips(detail=False)
# # Before Attack
model_before = GCN(adj, x, labels, device='GPU', norm_x=None, seed=42)
model_before.build()
his_before = model_before.train(idx_train, verbose=1, epochs=100)
loss, accuracy = model_before.test(idx_test)
print(f'Test loss {loss:.5}, Test accuracy {accuracy:.2%}')
# # After Attack
model_after = GCN(attacker.A, x, labels, device='GPU', norm_x=None, seed=42)
model_after.build()
his_after = model_after.train(idx_train, verbose=1, epochs=100)
loss, accuracy = model_after.test(idx_test)
print(f'Test loss {loss:.5}, Test accuracy {accuracy:.2%}')
# # Visulation
def plot(his_before, his_after, metric="loss"):
with plt.style.context(['science', 'no-latex']):
plt.plot(his_before.history[metric])
plt.plot(his_after.history[metric])
plt.legend([f'{metric.title()} Before', f'{metric.title()} After'])
plt.ylabel(f'{metric.title()}')
plt.xlabel('Epochs')
plt.show()
plot(his_before, his_after, metric="loss")
plot(his_before, his_after, metric="acc")
| examples/Untargeted Attack/test_FGSM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Observations and Insights
#
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import scipy.stats as sts
import numpy as np
# Study data files
mouse_metadata_path = "data/Mouse_metadata.csv"
study_results_path = "data/Study_results.csv"
# Read the mouse data and the study results
mouse_metadata = pd.read_csv(mouse_metadata_path)
study_results = pd.read_csv(study_results_path)
# Combine the data into a single dataset
merged_df=pd.merge(mouse_metadata, study_results, how='left', on='Mouse ID')
# Display the data table for preview
merged_df.head()
# -
mouse_metadata.head()
study_results.head()
# Checking the number of mice.
len(merged_df['Mouse ID'].unique())
# Getting the duplicate mice by ID number that shows up for Mouse ID and Timepoint.
duplicate_id=merged_df.loc[merged_df.duplicated(subset=['Mouse ID','Timepoint']),'Mouse ID'].unique()
duplicate_id
# Optional: Get all the data for the duplicate mouse ID.
duplicate_id_df = merged_df.loc[merged_df['Mouse ID']=='g989']
duplicate_id_df
# Create a clean DataFrame by dropping the duplicate mouse by its ID.
cleaned_df = merged_df.loc[merged_df['Mouse ID']!='g989']
cleaned_df
# Checking the number of mice in the clean DataFrame.
len(cleaned_df['Mouse ID'].unique())
# ## Summary Statistics
# +
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Use groupby and summary statistical methods to calculate the following properties of each drug regimen:
# mean, median, variance, standard deviation, and SEM of the tumor volume.
# Assemble the resulting series into a single summary dataframe.
mean_mouse = cleaned_df.groupby('Drug Regimen').mean()['Tumor Volume (mm3)']
median_mouse = cleaned_df.groupby('Drug Regimen').median()['Tumor Volume (mm3)']
var_mouse = cleaned_df.groupby('Drug Regimen').var()['Tumor Volume (mm3)']
std_mouse = cleaned_df.groupby('Drug Regimen').std()['Tumor Volume (mm3)']
sem_mouse = cleaned_df.groupby('Drug Regimen').sem()['Tumor Volume (mm3)']
summary_df = pd.DataFrame({'Mean Tumor Volume':mean_mouse,
'Median Tumor Volume':median_mouse,
'Variance Tumor Volume':var_mouse
,'Standard Deviation of Tumor Volume':std_mouse,
'SEM of Tumor Volume':sem_mouse})
summary_df
# -
# Generate a summary statistics table of mean, median, variance, standard deviation, and SEM of the tumor volume for each regimen
# Using the aggregation method, produce the same summary statistics in a single line
#df.groupby('A').agg({'B': ['min', 'max'], 'C': 'sum'})
summary_df2 = cleaned_df.groupby('Drug Regimen').agg({'Tumor Volume (mm3)':['mean', 'median','var','std','sem']})
summary_df2
# ## Bar and Pie Charts
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pandas.
# .valuecounts on drug regimen =mouse_count
color_list = ["green", "red", "blue", "yellow", "purple", "orange", "coral", "black","brown", "gray"]
regimen_summary = cleaned_df['Drug Regimen'].value_counts()
regimen_summary.plot(kind='bar',figsize=(10,5),rot=0,color=color_list,alpha=.65)
# Set a Title for the chart
plt.title('Total Number of Measurements per Regimen')
plt.xlabel('Drug Regimen')
plt.ylabel('Number of Measurements')
plt.ylim(125,250)
plt.show()
# +
# Generate a bar plot showing the total number of measurements taken on each drug regimen using pyplot.
#regimen_summary = cleaned_df['Drug Regimen'].value_counts()
#regimen_summary
drug_id_time_df = cleaned_df[["Drug Regimen","Timepoint","Mouse ID"]]
x = drug_id_time_df['Drug Regimen'].unique().tolist()
y = drug_id_time_df['Drug Regimen'].value_counts().tolist()
plt.figure()
plt.bar(x,y,color=color_list, alpha=.8,width=.4)
plt.title('Total Mice Per Timepoint for Drug Regiment')
plt.xlabel('Drug Regimen')
plt.ylabel('Number of Mice')
plt.ylim(100, 250)
# +
# Generate a pie plot showing the distribution of female versus male mice using pandas
M_vs_F = cleaned_df["Sex"].value_counts()
#print(M_vs_F)
gender = ["Male", "Female",]
explode = (0, .1)
M_vs_F.plot(kind="pie",autopct="%1.1f%%",startangle=140,colors = ['lightsalmon','darkturquoise'],explode = (0, .07),shadow=True)
plt.title('Distribution of Mouse Sexes')
plt.show()
# +
# Generate a pie plot showing the distribution of female versus male mice using pyplot
M_vs_F = cleaned_df["Sex"].value_counts()
#print(M_vs_F)
# Labels for the sections of our pie chart
gender = ["Male", "Female",]
# The colors of each section of the pie chart
color = color_list
# Tells matplotlib to seperate the "Female" section from the others
explode = (0, .07)
# Creates the pie chart based upon the values above
# Automatically finds the percentages of each part of the pie chart
plt.pie(M_vs_F, colors=['orchid','paleturquoise'],autopct="%1.1f%%", shadow=True, startangle=140, labels=gender, explode=explode,)
plt.title('Distribution of Mouse Sexes')
plt.show()
# -
# ## Quartiles, Outliers and Boxplots
# +
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
spec_drug_df = cleaned_df.loc[(cleaned_df["Drug Regimen"] == "Capomulin") |(cleaned_df["Drug Regimen"] == "Ramicane") |
(cleaned_df["Drug Regimen"] == "Infubinol") | (cleaned_df["Drug Regimen"] == "Ceftamin")]
# Start by getting the last (greatest) timepoint for each mouse
gg = spec_drug_df.groupby("Mouse ID")
for each_mouse in gg:
max_timepoint = gg["Timepoint"].max()
max_df = pd.DataFrame({
"Maximum Timepoint" : max_timepoint
})
new_df = pd.merge(spec_drug_df,max_df, on = "Mouse ID")
new_df = new_df.loc[new_df["Timepoint"] == new_df["Maximum Timepoint"]]
final_tumor_volume = new_df[["Mouse ID", "Tumor Volume (mm3)"]]
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
final_volume_merge = pd.merge(cleaned_df, final_tumor_volume, how = "outer", on = "Mouse ID")
rn_fnl_vol_df = final_volume_merge.rename(columns = ({
"Tumor Volume (mm3)_y" : "Final Tumor Volume (mm3)",
"Tumor Volume (mm3)_x" : "Tumor Volume (mm3)"
}))
rn_fnl_vol_df.head(30)
# +
# Calculate the final tumor volume of each mouse across four of the treatment regimens:
# Capomulin, Ramicane, Infubinol, and Ceftamin
# Start by getting the last (greatest) timepoint for each mouse
# Merge this group df with the original dataframe to get the tumor volume at the last timepoint
merged_data = drug_success[['Mouse ID','Timepoint']].merge(cleaned_df,on=['Mouse ID','Timepoint'],how="left")
merged_data.head()
tumor_vol_list = []
treatment_list = ["Capomulin", "Ramicane", "Infubinol", "Ceftamin"]
for drug in treatment_list:
final_tumor_vol = merged_data.loc[merged_data["Drug Regimen"] == drug, 'Tumor Volume (mm3)']
tumor_vol_list.append(final_tumor_vol)
fig1, axl = plt.subplots()
# ax1.set_ylabel('Final Tumor Volume (mm3)')
# axl.boxplot(tumor_vol_list)
# plt.show()
axl.boxplot(tumor_vol_list, labels = treatment_list)
# plt.ylabel('Final Tumor Volume (mm3)')
plt.title("Drug Trial Results baased on Tumor Volume (mm3)")
plt.ylabel("Tumor Volume (mm3)")
plt.xlabel("Drug Adminstered")
plt.grid=(True)
# -
# ## Line and Scatter Plots
# Generate a line plot of tumor volume vs. time point for a mouse treated with Capomulin
capomulin_data = cleaned_df[cleaned_df["Drug Regimen"] == "Capomulin"]
capomulin_mouse_data = capomulin_data[capomulin_data["Mouse ID"] == "s185"]
x_line = capomulin_mouse_data["Timepoint"]
y_line = capomulin_mouse_data["Tumor Volume (mm3)"]
plt.plot(x_line, y_line)
plt.title("Treatment of Mouse 's185' on Capomulin")
plt.xlabel("Time (days)")
plt.ylabel("Tumor Volume (mm3)")
plt.grid(True)
plt.xlim(0,45.5)
plt.ylim(0,50)
plt.show()
# Generate a scatter plot of average tumor volume vs. mouse weight for the Capomulin regimen
grouped_mouse = capomulin_data.groupby(["Mouse ID"])
grouped_weight = grouped_mouse["Weight (g)"].mean()
avg_tumor_size_bymouse = grouped_mouse["Tumor Volume (mm3)"].mean()
plt.scatter(x = grouped_weight, y = avg_tumor_size_bymouse)
plt.title("Average Tumor Size (mm3) vs. Weight of Mouse during Capomulin Drug Trial")
plt.xlabel("Weight of Mouse (g)")
plt.ylabel("Average Tumor Size (mm3)")
plt.grid(True)
plt.xlim(12,28)
plt.ylim(30,50)
plt.show()
# ## Correlation and Regression
# Calculate the correlation coefficient and linear regression model
# for mouse weight and average tumor volume for the Capomulin regimen
corr_coeff = round(sts.pearsonr(grouped_weight,avg_tumor_size_bymouse)[0],2)
plt.scatter(x = grouped_weight, y = avg_tumor_size_bymouse)
plt.title("Average Tumor Size (mm3) vs. Weight of Mouse during Capomulin Drug Trial")
plt.xlabel("Weight of Mouse (g)")
plt.ylabel("Average Tumor Size (mm3)")
plt.grid(True)
plt.xlim(14,26)
plt.ylim(30,50)
linregress = sts.linregress(x = grouped_weight, y = avg_tumor_size_bymouse)
slope = linregress[0]
intercept = linregress[1]
bestfit = slope*grouped_weight + intercept
plt.plot(grouped_weight,bestfit, "--",color = "red")
plt.show()
print(f'The correlation coeffienct is {corr_coeff} for the Mouse Weight against the Tumor volume.')
| Pymaceuticals/pymaceuticals_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Face completion with a multi-output estimators
#
#
# This example shows the use of multi-output estimator to complete images.
# The goal is to predict the lower half of a face given its upper half.
#
# The first column of images shows true faces. The next columns illustrate
# how extremely randomized trees, k nearest neighbors, linear
# regression and ridge regression complete the lower half of those faces.
#
#
#
# +
print(__doc__)
import numpy as np
import matplotlib.pyplot as plt
from sklearn.datasets import fetch_olivetti_faces
from sklearn.utils.validation import check_random_state
from sklearn.ensemble import ExtraTreesRegressor
from sklearn.neighbors import KNeighborsRegressor
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import RidgeCV
# Load the faces datasets
data = fetch_olivetti_faces()
targets = data.target
data = data.images.reshape((len(data.images), -1))
train = data[targets < 30]
test = data[targets >= 30] # Test on independent people
# Test on a subset of people
n_faces = 5
rng = check_random_state(4)
face_ids = rng.randint(test.shape[0], size=(n_faces, ))
test = test[face_ids, :]
n_pixels = data.shape[1]
# Upper half of the faces
X_train = train[:, :(n_pixels + 1) // 2]
# Lower half of the faces
y_train = train[:, n_pixels // 2:]
X_test = test[:, :(n_pixels + 1) // 2]
y_test = test[:, n_pixels // 2:]
# Fit estimators
ESTIMATORS = {
"Extra trees": ExtraTreesRegressor(n_estimators=10, max_features=32,
random_state=0),
"K-nn": KNeighborsRegressor(),
"Linear regression": LinearRegression(),
"Ridge": RidgeCV(),
}
y_test_predict = dict()
for name, estimator in ESTIMATORS.items():
estimator.fit(X_train, y_train)
y_test_predict[name] = estimator.predict(X_test)
# Plot the completed faces
image_shape = (64, 64)
n_cols = 1 + len(ESTIMATORS)
plt.figure(figsize=(2. * n_cols, 2.26 * n_faces))
plt.suptitle("Face completion with multi-output estimators", size=16)
for i in range(n_faces):
true_face = np.hstack((X_test[i], y_test[i]))
if i:
sub = plt.subplot(n_faces, n_cols, i * n_cols + 1)
else:
sub = plt.subplot(n_faces, n_cols, i * n_cols + 1,
title="true faces")
sub.axis("off")
sub.imshow(true_face.reshape(image_shape),
cmap=plt.cm.gray,
interpolation="nearest")
for j, est in enumerate(sorted(ESTIMATORS)):
completed_face = np.hstack((X_test[i], y_test_predict[est][i]))
if i:
sub = plt.subplot(n_faces, n_cols, i * n_cols + 2 + j)
else:
sub = plt.subplot(n_faces, n_cols, i * n_cols + 2 + j,
title=est)
sub.axis("off")
sub.imshow(completed_face.reshape(image_shape),
cmap=plt.cm.gray,
interpolation="nearest")
plt.show()
| 01 Machine Learning/scikit_examples_jupyter/plot_multioutput_face_completion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false deletable=true id="OJkcQuONXhj4" new_sheet=false run_control={"read_only": false}
# # Série 2
#
# Dans cette série on va s'occuper de la cryptographie asymétrique et faire quelques exercices avec. On utilise la librairie libnacl (prononcé "lib-salt", peut-être "lib-sel" en français). Elle est bien connue et permet d'utiliser les algorithmes cryptographiques à différents niveaux.
#
# Exercice 1 va vous montrer comment on signe un message, et dans l'Exercice 2 on va essayer de faire un echange de Diffie-Hellman.
# + [markdown] button=false deletable=true id="xc0a_EOTck4x" new_sheet=false run_control={"read_only": false}
# # Exercice 2
#
# Ici on va faire une petite introduction à Diffie-Hellman. La première partie va donne une introduction dans les opérations cryptographiques nécessaires. La deuxième partie vérifie que l'opération Diffie-Hellman marche. Et la troisième partie combine le Diffie-Hellman avec la signature nécessaire pour pouvoir utiliser une clé à longue durée.
#
# ## 1. Connaissance
#
# Création de clés: libnacl vous donne la possibilité de créer une paire de clés avec la méthode suivante:
#
# priv, pub = libnacl.crypto_keypair()
#
# Multiplication: avec les courbes elliptiques, une multiplications peut se faire entre deux clés privées (vu que ce sont des nombres) ou entre une clé privée et une clé publique. Si vous connaissez le RSA, ceci correspond à peu près à l'exponentiation modulaire. Avec libnacl, la méthode à utiliser est la suivante:
#
# resultat = libnacl.crypto_scalarmult(pub, priv)
#
# Dans l'exemple suivant, on va vérifier que les deux parties du Diffie-Hellman donnent bien le même résultat.
# + button=false colab={"base_uri": "https://localhost:8080/", "height": 368} deletable=true executionInfo={"elapsed": 2479, "status": "error", "timestamp": 1620330981935, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiT9d_hiiZE9FHyx2dphb24xe4f0737HSVlpv54=s64", "userId": "10501199511424753986"}, "user_tz": -120} id="4qOznnPDckVh" jupyter={"outputs_hidden": false} new_sheet=false outputId="5d83ac62-f84e-405a-8990-2414a9d0e570" run_control={"read_only": false}
# Exercice 2 - Partie 1
import libnacl, ctypes
def crypto_scalarmult(priv, pub):
'''
Cette méthode retourne le résultat de la multiplication de la clé privée avec
la clé publique
'''
if len(priv) != libnacl.crypto_box_SECRETKEYBYTES:
raise ValueError('Invalid secret key')
if len(pub) != libnacl.crypto_box_PUBLICKEYBYTES:
raise ValueError('Invalid public key')
result = ctypes.create_string_buffer(libnacl.crypto_box_PUBLICKEYBYTES)
if libnacl.nacl.crypto_scalarmult(result, priv, pub):
raise libnacl.CryptError('Failed to compute scalar product')
return result.raw
pub, priv = libnacl.crypto_box_keypair()
res = crypto_scalarmult(priv, pub)
print("Clé privée:", priv.hex())
print("Clé publique:", pub.hex())
print("Multiplication:", res.hex())
# -
# ## 2. Compréhension
#
# Supposons qu'Alice et Bob veulent créer une clé symétrique en utilisant Diffie-Hellman. Ils vont donc faire la chose suivante:
#
# 1. Alice et Bob vont créer leurs paires de clés:
# ```
# alice_pub, alice_priv pour Alice
# bob_pub, bob_priv pour Bob
# ```
#
#
# 2. Alice envoie sa clé publique (alice_pub) à Bob
# 3. Bob envoie sa clé publique (bob_pub) à Alice
# 3. Bob fait le calcul suivant:
# ```
# bob_symetrique = bob_priv * alice_pub
# ```
#
#
# 4. Alice fait de même:
# ```
# alice_symetrique = alice_priv * bob_pub
# ```
#
#
# A la fin les deux doivent avoir le même résultat!
#
# Utilisez les éléments que vous avez vu dans la 1ère partie pour implémenter cet algorithme! Vous n'avez pas besoin de copier la méthode `crypto_scalarmult` - elle est disponible une fois que vous avez exécuté le block correspondant.
#
# Pour finaliser l'exercice, vous pouvez encore faire le hachage sha256 du résultat de la multiplication. Ceci assure que d'éventuels structures de la clé publique ne sont plus visibles.
# + button=false deletable=true id="9HYfk1lTc7Hp" new_sheet=false run_control={"read_only": false}
# Exercice 1 - Partie 2
# -
# ## 3. Application
#
# Maintenant on suppose qu'Alice et Bob ont une paire de clés long terme, c-à-dire qu'ils utilisent pour plusieurs échanges Diffie-Hellman. Ces clés sont connues d'avance, donc Bob connaît la clé publique longue durée d'Alice, et vice-versa.
#
# Ajouter la signature à l'exercice sur le Diffie-Hellman, en vérifiant de chaque côté. On aura donc:
#
# ### Préparation:
#
# Alice et Bob choisissent leur paires de clés longue durée et échangent la partie publique: alice_pub_long et bob_pub_long
#
# ### Algorithme:
#
# 1. Alice et Bob choisissent une paire de clés pour la session
# 2. Alice envoie sa clé publique, signée par sa clé longue durée
# 3. Bob vérifie la signature avec la clé publique de longue durée d'Alice
# 4. Bob envoie sa clé publique, signée par sa clé longue durée
# 5. Alice vérifie la signature avec la clé publique de longue durée de Bob
# 6. Alice et Bob font le calcule Diffie-Hellman, puis font un sha256 dessus
# + button=false deletable=true id="-Fk-3jvpdEcq" new_sheet=false run_control={"read_only": false} tags=[]
# Exercice 1 - Partie 3
| Jour-3/Serie-2/jour_3_serie_2_exo_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.1 64-bit
# language: python
# name: python_defaultSpec_1595328765133
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Tricks
# -
# First, lets make some variables that we can do things to as a demo:
# + slideshow={"slide_type": "slide"}
pets = ["dog", "goat", "pig", "sheep", "cattle", "zebu", "cat", "chicken",
"guinea pig", "donkey", "duck", "water buffalo",
"western honey bee", "dromedary camel", "horse", "silkmoth",
"pigeon", "goose", "yak", "bactrian camel", "llama", "alpaca",
"guineafowl", "ferret", "muscovy duck", "barbary dove",
"bali cattle", "gayal", "turkey", "goldfish", "rabbit", "koi",
"canary", "society finch", "fancy mouse", "siamese fighting fish",
"fancy rat and lab rat", "mink", "red fox", "hedgehog", "guppy"]
# pets = list(range(40))
numbers = list(range(30))
message_dict = {"name": "Ben",
"Year": 2017,
"Location": "Sydney",
"Greeting": "Yo whatup now and give a brother room",
"Fact": "It would take 1,200,000 mosquitoes, each " +
"sucking once, to completely drain the " +
"average human of blood",
"Alphabet Inc Class A": "847.80USD",
"fruit": ["apple", "apricot", "avocado", "abiu"]}
# + [markdown] slideshow={"slide_type": "slide"}
# # Slicing
#
# This is a powerful Python idea that is now available in a few other languages.
#
# Try and come up with your own way to remember the syntax, and if you do, share it with me!
# + tags=[]
print("[:10]", pets[:10])
# + tags=[]
print("\n[35:]", pets[35:])
# + tags=[]
print("\n[4:20]", pets[4:20])
# + tags=[]
print("\n[:-35]", pets[:-35])
# + tags=[]
print("\n[::-1]", pets[::-1])
# -
some_pets = pets[4:10]
some_pets
# + [markdown] slideshow={"slide_type": "slide"}
# # Append vs Extend
#
# We've seen append a lot, it adds something to a list.
#
# _Extend_ works on lists, and adds the _contents_ of the list to a list. It's rare that you'll need it, but it's really helpful when you do.
# + slideshow={"slide_type": "fragment"} tags=[]
new_list = []
new_list.append(numbers)
new_list.append(some_pets)
print(new_list)
# + slideshow={"slide_type": "fragment"} tags=[]
another_new_list = []
another_new_list.extend(numbers)
another_new_list.extend(some_pets)
print(another_new_list)
# + [markdown] slideshow={"slide_type": "slide"}
# # Truthy values
#
# You don't always need to say `x == True`, if `x` is _truthy_, you can use it directly.
# + slideshow={"slide_type": "fragment"}
thing = False
if thing:
print("You'll never see me")
# + tags=[]
if "":
print("Empty strings are falsy")
if "Hello":
print("Non empty strings are truthy")
if 0:
print("Zero is falsy")
if 6:
print("Non-zero numbers are truthy")
if -7:
print("Non-zero numbers are truthy, even negative ones")
if []:
print("Empty lists are falsy")
if ["stuff"]:
print("Non empty lists are truthy...")
if [False]:
print("...even if they're full of falsy things")
if None:
print("None is falsy")
if False:
print("False is falsy")
if True:
print("True is truthy")
# + slideshow={"slide_type": "fragment"} tags=[]
thing = "hi!"
if thing:
print("Woah, look at me!" )
# + slideshow={"slide_type": "fragment"}
thing = None
if thing:
print("I'm a sneaky snake")
# + [markdown] slideshow={"slide_type": "slide"}
# # List comprehensions
# + slideshow={"slide_type": "fragment"} tags=[]
pet_name_lengths = []
for p in pets:
pet_name_lengths.append(len(p))
print(pet_name_lengths)
# + slideshow={"slide_type": "fragment"} tags=[]
pet_name_lengths = [len(p) for p in pets]
print(pet_name_lengths)
# + tags=[]
short_pets = [p for p in pets if len(p) < 6]
print(short_pets)
# + tags=[]
big_short_pets = [p.upper() for p in pets if len(p) < 6]
print(big_short_pets)
# + [markdown] slideshow={"slide_type": "slide"}
# # λambdas
#
# Lambdas are anonymous functions. They're used a bit as Pandas apply functions, but they're a bit crap these days (in python 3), and comprehensions will usually do the job.
#
# They're massive in javascript, so often if people are converting, they'll reach for them.
# + slideshow={"slide_type": "fragment"} tags=[]
def get_2x_len(my_string):
return len(my_string) * 2
print(list(map(get_2x_len, pets)))
# + tags=[]
print(list(map(lambda x: len(x) * 2, pets)))
# + tags=[]
print([len(p) * 2 for p in pets])
# + slideshow={"slide_type": "fragment"} tags=[]
print(list(map(lambda x: x[1], pets)))
# + tags=[]
print([x[1] for x in pets])
# + slideshow={"slide_type": "fragment"} tags=[]
print(list(map(len, pets)))
# + [markdown] slideshow={"slide_type": "slide"}
# # Built ins
#
# Some functions don't need to be imported, they're always available.
# + slideshow={"slide_type": "-"}
from random import randint
my_odd_list = [randint(0,100) for _ in range(30)]
# + slideshow={"slide_type": "fragment"}
max(my_odd_list)
# + slideshow={"slide_type": "fragment"}
min(my_odd_list)
# + slideshow={"slide_type": "slide"}
list(zip(range(len(pets)), pets))
# + slideshow={"slide_type": "slide"} tags=[]
for p in enumerate(pets):
print(p)
# + [markdown] slideshow={"slide_type": "slide"}
# # Generators
# + slideshow={"slide_type": "slide"}
a = enumerate(pets)
# -
next(a)
# + slideshow={"slide_type": "slide"} tags=[]
def a_generator():
counter = 0
while True:
yield counter
counter += 3
g = a_generator()
print(next(g))
print(next(g))
print(next(g))
# -
# # Dictionary comprehensions
# + tags=[]
# all yours terry!
L = range(10)
d = {k:v for k, v in zip(pets, L)}
print(d)
# + tags=[]
import collections
d = collections.defaultdict(list)
d = collections.Counter()
d = collections.Counter(my_odd_list)
print(d)
print()
print(max(d, key=lambda x: d[x]))
# -
| tricks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sqc
import numpy as np
from exercises import CRz,qft,cqft
import matplotlib.pyplot as plt
import matplotlib.animation as animation
# +
def period(x,L):
if x >= L/2:
return x - L
return x
def w(j,N):
assert(j<N and j>=0)
if j == N-1:
return -2**j
return 2**j
def CexpIaX(c,a,xbits,op):
N=len(xbits)
for j in range(len(xbits)):
op=CRz(c,xbits[j],a*w(j,N),op)
return op
def CexpIaP(c,a,xbits,op):
N=len(xbits)
op=qft(op,mask=xbits,inverse=False)
op=CexpIaX(c,a*2.*np.pi / 2**N,xbits,op)
op=qft(op,mask=xbits,inverse=True)
return op
nxbits=5
nabits=1
nbits=nxbits+nabits
s0=sqc.state(nbits,basis=["|%d>|%d>" % (x//2**nxbits,period(x%2**nxbits,2**nxbits))
for x in range(2**nbits)])
s1=sqc.operator(nbits).X(1).X(2).X(nxbits)*s0
print(s1)
s2=CexpIaP(nxbits,11,range(nxbits),sqc.operator(nbits))*s1
print(s2)
# +
def CexpIaX2(c,a,xbits,op):
N=len(xbits)
l=a/12.*(2.+4.**N)-a/2.
op=CRz(c,xbits[0],l,CRz(c,xbits[0],l,op.X(xbits[0])).X(xbits[0]))
for j in range(len(xbits)):
op=CRz(c,xbits[j],-a*w(j,N),op)
for i in range(len(xbits)):
if i != j:
t=a/4.*w(j,N)*w(i,N)
ga=xbits[i]
gb=xbits[j]
op=op.CNOT(gb,ga)
op=op.X(ga)
op=CRz(c,ga,t,op)
op=op.X(ga)
op=CRz(c,ga,-t,op)
op=op.CNOT(gb,ga)
return op
def CexpIaP2(c,a,xbits,op):
N=len(xbits)
op=qft(op,mask=xbits,inverse=False)
op=CexpIaX2(c,a*(2.*np.pi / 2**N)**2.,xbits,op)
op=qft(op,mask=xbits,inverse=True)
return op
# -
s3=CexpIaX2(nxbits,1,range(nxbits),sqc.operator(nbits))*s1
print(np.log(s3[6+2**nxbits])/1j+12.*np.pi)
def timeEvolutionPlot(dt,steps,s1):
Tdt=CexpIaP2(nxbits,dt,range(nxbits),sqc.operator(nbits))
print(len(Tdt.m))
sn=s1
for tstep in range(steps):
res=sqc.sample(sn, n=100, mask=range(nxbits))
sn=Tdt*sn
xvals=[ period(x,2**nxbits) for x in res.keys() ]
yvals=res.values()
plt.xlabel('x')
plt.xlim(-2**nxbits/2,2**nxbits/2)
plt.ylabel('count')
plt.title('t=%g' % (tstep*dt))
plt.bar(xvals,yvals,width=0.75)
plt.show()
print(s1)
timeEvolutionPlot(0.1,30,s1)
# Time-periodicity: all eigenvalues are multiple of (2pi/L)**2 -> once smallest eval has 2pi-periodicity,
# entire system has 2pi-periodicity -> temporal loop
Tcrit=2*np.pi / (2*np.pi/32)**2.
timeEvolutionPlot(Tcrit/20,21,s1)
# +
# Now discuss circuit to create smeared packet
#def Rot(i,t,op):
# return op.Rz(i,-np.pi/2).H(i).Rz(i,-2*t).H(i).Rz(i,t).X(i).Rz(i,t).X(i).Rz(i,np.pi/2)
#s1=CexpIaP(nxbits,0.5,range(nxbits),sqc.operator(nbits).X(0).X(nxbits))*s0
nxbits=8
nabits=1
nbits=nxbits+nabits
s0=sqc.state(nbits,basis=["|%d>|%d>" % (x//2**nxbits,period(x%2**nxbits,2**nxbits))
for x in range(2**nbits)])
def createSource(xbits,p,op):
n=len(xbits)
for i in range(n):
op=op.H(xbits[i]).Rz(xbits[i],-p*2**i)
return op
s1=createSource(range(0),2*np.pi/2**nxbits*64,sqc.operator(nbits).X(nxbits))*s0
print(s1)
timeEvolutionPlot(5,4,s1)
#s2=expIaP2(0.3,range(nxbits),sqc.operator(nxbits))*s1
#s3=qft(sqc.operator(nxbits),mask=range(nxbits),inverse=False)*s2
#timeEvolutionPlot(0.1,30,s3)
# +
s1=createSource(range(4),2*np.pi/2**nxbits*64,sqc.operator(nbits).X(nxbits))*s0
print(s1)
timeEvolutionPlot(5,4,s1)
| examples/chapter3/one-dim-free-particle.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chapter 7: Sequential Data (Review Questions)
# The questions below assume that you have read the [first <img height="12" style="display: inline-block" src="../static/link/to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/develop/07_sequences/00_content.ipynb), [second <img height="12" style="display: inline-block" src="../static/link/to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/develop/07_sequences/01_content.ipynb), and the [third <img height="12" style="display: inline-block" src="../static/link/to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/develop/07_sequences/03_content.ipynb) part of Chapter 7. Some questions regard the [Appendix <img height="12" style="display: inline-block" src="../static/link/to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/develop/07_sequences/05_appendix.ipynb); that is indicated with a **\***.
#
# Be concise in your answers! Most questions can be answered in *one* sentence.
# ## Essay Questions
# Answer the following questions *briefly*!
# **Q1**: We have seen **containers** and **iterables** before in [Chapter 4 <img height="12" style="display: inline-block" src="../static/link/to_nb.png">](https://nbviewer.jupyter.org/github/webartifex/intro-to-python/blob/develop/04_iteration/02_content.ipynb#Containers-vs.-Iterables). How do they relate to **sequences**?
# < your answer >
# **Q2**: What are **abstract base classes**? How can we make use of the ones from the [collections.abc <img height="12" style="display: inline-block" src="../static/link/to_py.png">](https://docs.python.org/3/library/collections.abc.html) module in the [standard library <img height="12" style="display: inline-block" src="../static/link/to_py.png">](https://docs.python.org/3/library/index.html)?
# < your answer >
# **Q3**: How are the *abstract behaviors* of **reversibility** and **finiteness** essential for *indexing* and *slicing* sequences?
# < your answer >
# **Q4**: Explain the difference between **mutable** and **immutable** objects in Python with the examples of the `list` and `tuple` types!
# < your answer >
# **Q5**: What is the difference between a **shallow** and a **deep** copy of an object? How can one of them become a "problem?"
# < your answer >
# **Q6**: Many **list methods** change `list` objects "**in place**." What do we mean by that?
# < your answer >
# **Q7.1**: `tuple` objects have *two* primary usages. First, they can be used in place of `list` objects where **mutability** is *not* required. Second, we use them to model data **records**.
#
# Describe why `tuple` objects are a suitable replacement for `list` objects in general!
# < your answer >
# **Q7.2\***: What do we mean by a **record**? How are `tuple` objects suitable to model records? How can we integrate a **semantic meaning** when working with records into our code?
# < your answer >
# **Q8**: How is (iterable) **packing** and **unpacking** useful in the context of **function definitions** and **calls**?
# < your answer >
# ## True / False Questions
# Motivate your answer with *one short* sentence!
# **Q9**: `sequence` objects are *not* part of core Python but may be imported from the [standard library <img height="12" style="display: inline-block" src="../static/link/to_py.png">](https://docs.python.org/3/library/index.html).
# < your answer >
# **Q10**: The built-in [.sort() <img height="12" style="display: inline-block" src="../static/link/to_py.png">](https://docs.python.org/3/library/stdtypes.html#list.sort) function takes a *finite* **iterable** as its argument an returns a *new* `list` object. On the contrary, the [sorted() <img height="12" style="display: inline-block" src="../static/link/to_py.png">](https://docs.python.org/3/library/functions.html#sorted) method on `list` objects *mutates* them *in place*.
# < your answer >
# **Q11**: Passing **mutable** objects as arguments to functions is not problematic because functions operate in a **local** scope without affecting the **global** scope.
# < your answer >
| 07_sequences/07_review.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # ML Table Creation 1
#
# First iteration of ML Table creation for a cleaned table ready for ML. The table will be used as reference in ML testing. The table is uploaded to the teams S3 bucket as "MLTable1.csv"
#
# #### Method:
#
# This is code for creating a ML which consist of satellite detections (NASA FIRMS detections) considered as instances which were merged wwith weather and soil data from the USDA SCAN network.
#
# The table table is then merged with WFIGS actual records of wildifres based on initial coordinates and date the fires started.
#
# Certain columns were dropped for multiple resons described in the notebook.
#
# Lastly, the nan values were dropped, giving a table ready for ML reference.
#
# #### Additional Note:
#
# Most FIRMS detections were not associated with a major fire. The ML activities are in attempt to train a model to predict if a FIRMS detection will lead to a fire.
#
# This will support the team's goal **“Given that a FIRMS (Fire Information for Resource Management System) detection is found, how might we predict if the detection will turn into a wildfire by referencing historical WFIGS (Wildland Fire
# Interagency Geospatial Services) data?”.**
#general imports to start
#general imports
import pandas as pd
import boto3
import numpy as np
pd.set_option('display.max_columns', 200)
# ## Load and Merge the Data
# +
#load in the csvs
#TODO For Team: enter the credentails below to run
S3_Key_id=''
S3_Secret_key=''
def pull_data(Key_id, Secret_key, file):
"""
Function which CJ wrote to pull data from S3
"""
BUCKET_NAME = "gtown-wildfire-ds"
OBJECT_KEY = file
client = boto3.client(
's3',
aws_access_key_id= Key_id,
aws_secret_access_key= Secret_key)
obj = client.get_object(Bucket= BUCKET_NAME, Key= OBJECT_KEY)
file_df = pd.read_csv(obj['Body'])
return (file_df)
#Pull in the firms and scan df
file = 'FIRMSandSCANFull2018toApr2022_V2.csv'
df_FirmsAndScan = pull_data(S3_Key_id, S3_Secret_key, file)
df_FirmsAndScan.head()
# -
#pull in the big WFIGS dataframe
file = 'WFIGS_big_Pulled5-8-2022.csv'
df_wfigsbig = pull_data(S3_Key_id, S3_Secret_key, file)
df_wfigsbig.head()
# now starting to transform the fire db
latlonground = 1 #Sets the number of decimal places to round latlong
#getting the fire discovered date into the same format as the same date
df_wfigsbig['disc_date'] = df_wfigsbig['FireDiscoveryDateTime'].astype(str).str[0:10]
df_wfigsbig['disc_date'] = df_wfigsbig['disc_date'].astype(str).str.replace('/', '-')
# #convert the init long and lat frame to 2 decimals like i did in the satellite df
df_wfigsbig['init_lat_rounded'] = round(df_wfigsbig['Y'], latlonground) #rounds lat long into new column, using latlonground # of decimal pts
df_wfigsbig['init_long_rounded'] = round(df_wfigsbig['X'], latlonground) #rounds lat long into new column, using latlonground # of decimal pts
#make the date loc column
df_wfigsbig['disc_date_loc'] = df_wfigsbig['disc_date'] + ',' + df_wfigsbig['init_lat_rounded'].astype(str) + ',' + df_wfigsbig['init_long_rounded'].astype(str)
df_wfigsbig.head()
#Merging on the date + location key.
dfDiscovered = df_FirmsAndScan.merge(df_wfigsbig, left_on='date_loc', right_on='disc_date_loc', how='inner') #creates df of only dsetected areas
dfMerged = df_FirmsAndScan.merge(df_wfigsbig, left_on='date_loc', right_on='disc_date_loc', how='left') #creates df of both
dfMerged['FIRE_DETECTED'] = dfMerged['OBJECTID'].isnull() #if there was no fire detectection set to true
dfMerged['FIRE_DETECTED'] = ~dfMerged['FIRE_DETECTED'].astype(bool) #flip so that if detection merged set to true
print(dfMerged.shape)
dfMerged.head()
#check the number of merges
dfDiscovered.shape
# ## Clean and Prep the Data
#beign cleaning the data by removing unnecessary columns:
#drop the columns we VERY likely do not need for this approach
coltodrop = ['Unnamed: 0',
'Unnamed: 0.1',
'acq_date',
'acq_time',
'version',
'lat',
'long',
'hour',
'date_loc',
'date_hour_loc',
'newdate',
'nearestStation',
'Date',
'Station Id',
'State Code',
'Network Code',
'Station Name',
'Elevation (ft)',
'Latitude',
'Longitude',
'County Name',
'HUC6 Name',
'HUC8 Name',
'Wind Speed Average 324in (mph)',
'Wind Speed Average 128in (mph)',
'date_new',
'X',
'Y',
'OBJECTID',
'ABCDMisc',
'ADSPermissionState',
'CalculatedAcres',
'ContainmentDateTime',
'ControlDateTime',
'DailyAcres',
'DiscoveryAcres',
'DispatchCenterID',
'EstimatedCostToDate',
'FinalFireReportApprovedByTitle',
'FinalFireReportApprovedByUnit',
'FinalFireReportApprovedDate',
'FireBehaviorGeneral',
'FireBehaviorGeneral1',
'FireBehaviorGeneral2',
'FireBehaviorGeneral3',
'FireCause',
'FireCauseGeneral',
'FireCauseSpecific',
'FireCode',
'FireDepartmentID',
'FireDiscoveryDateTime',
'FireMgmtComplexity',
'FireOutDateTime',
'FireStrategyConfinePercent',
'FireStrategyFullSuppPercent',
'FireStrategyMonitorPercent',
'FireStrategyPointZonePercent',
'FSJobCode',
'FSOverrideCode',
'GACC',
'ICS209ReportDateTime',
'ICS209ReportForTimePeriodFrom',
'ICS209ReportForTimePeriodTo',
'ICS209ReportStatus',
'IncidentManagementOrganization',
'IncidentName',
'IncidentShortDescription',
'IncidentTypeCategory',
'IncidentTypeKind',
'InitialLatitude',
'InitialLongitude',
'InitialResponseAcres',
'InitialResponseDateTime',
'IrwinID',
'IsFireCauseInvestigated',
'IsFireCodeRequested',
'IsFSAssisted',
'IsMultiJurisdictional',
'IsReimbursable',
'IsTrespass',
'IsUnifiedCommand',
'LocalIncidentIdentifier',
'PercentContained',
'PercentPerimeterToBeContained',
'POOCity',
'POOCounty',
'POODispatchCenterID',
'POOFips',
'POOJurisdictionalAgency',
'POOJurisdictionalUnit',
'POOJurisdictionalUnitParentUnit',
'POOLandownerCategory',
'POOLandownerKind',
'POOLegalDescPrincipalMeridian',
'POOLegalDescQtr',
'POOLegalDescQtrQtr',
'POOLegalDescRange',
'POOLegalDescSection',
'POOLegalDescTownship',
'POOPredictiveServiceAreaID',
'POOProtectingAgency',
'POOProtectingUnit',
'POOState',
'PredominantFuelGroup',
'PredominantFuelModel',
'PrimaryFuelModel',
'SecondaryFuelModel',
'TotalIncidentPersonnel',
'UniqueFireIdentifier',
'WFDSSDecisionStatus',
'CreatedBySystem',
'ModifiedBySystem',
'IsDispatchComplete',
'OrganizationalAssessment',
'StrategicDecisionPublishDate',
'CreatedOnDateTime_dt',
'ModifiedOnDateTime_dt',
'Source',
'GlobalID',
'IsCpxChild',
'CpxName',
'CpxID',
'disc_date',
'init_lat_rounded',
'init_long_rounded',
'disc_date_loc',
'time',
'newdatetime'
]
dfMerged = dfMerged.drop(coltodrop, axis = 1) #drops unwanted columns for ML process
dfMerged.head()
#remove columns we (likely) dont need for this approach
coltodrop = ['latitude',
'longitude',
'daynight',
'Soil Moisture Percent -4in (pct) Start of Day Values',
'HUC4 Name']
dfMerged = dfMerged.drop(coltodrop, axis = 1) #drops unwanted columns for ML process
dfMerged.head()
#checking some countsd of values on missing columns
collist = list(dfMerged.columns) #get the list of columns in the df
numbrows = dfMerged.shape[0] #saves number of riows in the df
for col in collist:
missingcount = dfMerged[col].isnull().sum() #count missing vals
missingpercent = round((missingcount/numbrows)*100, 2) #gets the percent compared to the entire dataframe
print('number of nan for '+col+':'+str(missingcount)+' ; '+str(missingpercent)+'%') #prints the result
#there appears to be a lot of missing data for 'type' and 'precipitation month to date'. drop those for this approach
dfMerged = dfMerged.drop('type', axis = 1)
dfMerged = dfMerged.drop('Precipitation Month-to-date (in) Start of Day Values', axis = 1)
#the station distance is not relevant for a fire prediction so dropping
#we may want to drop rows where the nearest station distance is too far in the future
dfMerged = dfMerged.drop('StationDist', axis = 1)
dfMerged.head()
#'confidence' appears to be starnge. based off the link below: n=nominal l = low and h = high
# https://cdn.earthdata.nasa.gov/conduit/upload/10575/MODIS_C6_Fire_User_Guide_B.pdf
#another helpful link:
# https://earthdata.nasa.gov/faq/firms-faq#ed-confidence
# lets approximate low 20, nominal 50, and high 80 for this table
# this can be adjusted later, if needed
mapping = {'l': 20, 'n': 50, 'h': 100}
dfMerged = dfMerged.replace({'confidence': mapping})
dfMerged['confidence'] = pd.to_numeric(dfMerged['confidence']) #convert confidence from string to integer
#now drop the nan rows
print('df shape before dropping all nan: '+str(dfMerged.shape))
dfMerged = dfMerged.dropna() #drop all nan values
print('df shape after dropping all nan: '+str(dfMerged.shape))
#check how many trues we have
dfMerged['FIRE_DETECTED'].value_counts()
#now begin to one hot encode the 'satellite' column
#first check what kind of values we have for the satellite column
dfMerged['satellite'].value_counts()
#weird how '1' appears twice, convert to list to check why and if different datatype
dfMerged['satellite'].unique().tolist()
#looks like some 1's are strings and some are ints. converting everything to string
dfMerged['satellite'] = dfMerged['satellite'].astype(str)
#check to make sure we are good now
dfMerged['satellite'].value_counts()
#add the one hot encoded df to the merged dataframe and drop the original satellite column
sat_dummies = pd.get_dummies(dfMerged.satellite)
dfMerged = pd.concat([dfMerged, sat_dummies], axis=1) #adds in the one hot encoded staellite df
dfMerged = dfMerged.drop('satellite', axis = 1) #drop the satellite column
dfMerged.head()
#now one hot encode instrument
ohe = pd.get_dummies(dfMerged.instrument) #creates the one hot encoded df of instrument
dfMerged = pd.concat([dfMerged, ohe], axis=1) #merges the dataframe with the one hot encoded df
dfMerged = dfMerged.drop('instrument', axis = 1) #drop the instrument column
dfMerged.head()
#because there is only 2 instruments (MODIS and VIIRS) we can drop one of the columns
#we will keep the MODIS column, so 1 = MODIS and 0 = VIIRS instrument
dfMerged = dfMerged.drop('VIIRS', axis = 1) #drop the instrument column
#the column name 'HUC2 Name' will not work for the necessary syntax because it has a space in it.
dfMerged = dfMerged.rename(columns={"HUC2 Name": "HUC2Name"}) #renames the HUC2 Name column to remove spaces
#now one hot encode the region by HUC2 name:
ohe = pd.get_dummies(dfMerged.HUC2Name) #creates the one hot encoded df of instrument
dfMerged = pd.concat([dfMerged, ohe], axis=1) #merges the dataframe with the one hot encoded df
dfMerged = dfMerged.drop('HUC2Name', axis = 1) #drop the instrument column
dfMerged.head()
#check the data types to make sure they are all values and not strings
dfMerged.dtypes
# ### Save the data
dfMerged.to_csv('MLTable1.csv')
| Ingestion and Wrangling/MLTableCreation/ML Table Creation 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yoyoyo-yo/DeepLearningMugenKnock/blob/master/notes_pytorch/AE/AE_MNIST_pytorch.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="xa9STwNFx962"
# # AutoEncoder
#
# + [markdown] id="3u9ehbSkx963"
# ## Import and Config
# + id="tDVr-eTxrXhe"
# !pip install -q --upgrade albumentations==0.5.1
# + id="fx5MA7lRm4tz"
import os
import time
from tqdm.notebook import tqdm
import numpy as np
import cv2
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
from sklearn.model_selection import KFold, GroupKFold, StratifiedKFold
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.utils.data import Dataset, DataLoader
import albumentations as A
import albumentations.pytorch as Ap
import torchvision
import torchvision.models as models
# + id="ZxI9fnf1nRS5"
IN_HEIGHT, IN_WIDTH = 28, 28
RESOURCE = "GPU"
FOLD = "StratifiedKFold" # KFold, GroupKFold, StratifiedKFold, StratifiedGroupXX
FOLD_N = 2 # fold num
RANDOM_SEED = 42
if RESOURCE == "CPU":
device = torch.device("cpu")
elif RESOURCE == "GPU":
device = torch.device("cuda")
# + [markdown] id="UL99hDa-x967"
# ## Define Model
# + id="GxdY2y83x967"
class Flatten(nn.Module):
def forward(self, x):
return x.view(x.size()[0], -1)
class Reshape(nn.Module):
def __init__(self, c, h, w):
super(Reshape, self).__init__()
self.c = c
self.h = h
self.w = w
def forward(self, x):
x = x.view(x.size()[0], self.c, self.h, self.w)
return x
class AutoEncoder(nn.Module):
def __init__(self, dim=32, in_channel=1):
super(AutoEncoder, self).__init__()
# Encoder
self.module_encoder = nn.Sequential(
Flatten(),
nn.Linear(IN_HEIGHT * IN_WIDTH * in_channel, dim * 16),
nn.ReLU(),
nn.Linear(dim * 16, dim * 8),
nn.ReLU(),
nn.Linear(dim * 8, dim),
nn.ReLU(),
)
# Decoder
self.module_decoder = nn.Sequential(
nn.Linear(dim, dim * 8),
nn.ReLU(),
nn.Linear(dim * 8, dim * 16),
nn.ReLU(),
nn.Linear(dim * 16, IN_WIDTH * IN_HEIGHT * in_channel),
nn.Tanh(),
Reshape(in_channel, IN_HEIGHT, IN_WIDTH)
)
def forward(self, x):
x = self.module_encoder(x)
return self.module_decoder(x)
def forward_encoder(self, x):
return self.module_encoder(x)
# + [markdown] id="hm-tUIRkppbl"
# # Dataset
# + id="Hl6fXlMTpqij"
train_ds = torchvision.datasets.MNIST(root="./", train=True, download=True, transform=None)
train_Xs = train_ds.data.numpy()[..., None].astype(np.float32)
train_ys = np.array(train_ds.targets)
class MnistDataset(Dataset):
def __init__(self, xs, ys, transforms=None):
self.xs = xs
self.ys = ys
self.transforms=transforms
self.data_num = len(xs)
def __len__(self):
return self.data_num
def __getitem__(self, idx):
x = self.xs[idx] / 127.5 - 1
y = self.ys[idx]
if self.transforms:
transformed = self.transforms(image=x)
x = transformed["image"]
return x, y
# + id="B2l9tCkerR7v"
transforms_train = A.Compose([
A.HorizontalFlip(p=0.5),
A.VerticalFlip(p=0.5),
A.ShiftScaleRotate(p=0.5),
A.RandomRotate90(p=0.5),
# A.Resize(IN_HEIGHT, IN_WIDTH),
# A.Normalize(max_pixel_value=255.0, p=1.0),
Ap.ToTensorV2(p=1.0),
])
transforms_val = A.Compose([
# A.Resize(IN_HEIGHT, IN_WIDTH),
# A.Normalize(max_pixel_value=255.0, p=1.0),
Ap.ToTensorV2(p=1.0),
])
# + [markdown] id="DsNa1zJ1pRfC"
# # Train
# + id="pzKwyS264N18"
def show_sample(Xs, ys, show_num=8):
Xs = Xs.detach().cpu().numpy()[:, 0]
Xs = (Xs * 127.5 + 127.5).astype(np.uint8)
ys = ys.detach().cpu().numpy()[:, 0]
ys = (ys * 127.5 + 127.5).astype(np.uint8)
plt.figure(figsize=(12, 2))
for i in range(show_num):
# show input
x = Xs[i]
plt.subplot(2, show_num, i + 1)
plt.imshow(x, cmap="gray")
plt.axis('off')
plt.title('Input{}'.format(i + 1))
# show output
y = ys[i]
plt.subplot(2, show_num, i + 1 + show_num)
plt.imshow(y, cmap="gray")
plt.axis('off')
plt.title('Output{}'.format(i + 1))
plt.show()
# + id="pKRGTQIMpS5r"
def train():
# fold
if FOLD == "KFold":
kf = KFold(n_splits=FOLD_N, shuffle=True, random_state=RANDOM_SEED)
spl = kf.split(train_ds)
elif FOLD == "GroupKFold":
kf = GroupKFold(n_splits=FOLD_N)
spl = kf.split(train_ds.data, train_ds.targets, train_ds.targets)
elif FOLD == "StratifiedKFold":
kf = StratifiedKFold(n_splits=FOLD_N, shuffle=True, random_state=RANDOM_SEED)
spl = kf.split(train_ds.data, train_ds.targets)
else:
print("invalid fold")
return None
train_models = []
train_model_paths = []
EPOCH = 10
for fold_i, (train_idx, val_idx) in enumerate(spl):
train_losses = []
val_losses = []
print(f"{FOLD} fold:{fold_i + 1}/{FOLD_N}")
print(f"train_N={len(train_idx)}, val_N={len(val_idx)}")
#---
# datasert
#---
X_train = train_Xs[train_idx]
y_train = train_ys[train_idx]
X_val = train_Xs[val_idx]
y_val = train_ys[val_idx]
dataset_train = MnistDataset(X_train, y_train, transforms=transforms_train)
dataset_val = MnistDataset(X_val, y_val, transforms=transforms_val)
dataloader_train = DataLoader(dataset_train, batch_size=512, num_workers=4, shuffle=True, pin_memory=True)
dataloader_val = DataLoader(dataset_val, batch_size=512, num_workers=4, shuffle=False, pin_memory=True)
train_n = len(X_train)
val_n = len(X_val)
target_n = 1
#---
# model
#---
model = AutoEncoder()
model = model.to(device)
criterion = nn.MSELoss()
# optimizer = optim.SGD(model.parameters(), lr=0.01)
optimizer = optim.Adam(model.parameters(), lr=0.001)
#---
# epoch
#---
for epoch in range(EPOCH):
model.train()
tr_loss = 0
correct = 0
total = 0
#---
# train
#---
train_time_start = time.time()
for step, batch in enumerate(dataloader_train):
optimizer.zero_grad()
xs = batch[0].to(device) # image
ys = batch[1].to(device) # target
outputs = model(xs)
loss = criterion(outputs, xs)
loss = loss# / train_n
loss.backward()
loss = loss.item()
tr_loss += loss
_, predicted = torch.max(outputs.data, 1)
total += ys.size(0)
correct += 0#(predicted == ys).sum().item()
optimizer.step()
train_losses.append(tr_loss)
train_time_end = time.time()
#---
# val
#---
model.eval()
val_loss = 0
val_correct = 0
val_total = 0
val_time_start = time.time()
val_labels = []
val_preds = []
with torch.no_grad():
for step, batch in enumerate(dataloader_val):
xs = batch[0].to(device) # image
ys = batch[1].to(device) # target
outputs = model(xs)
loss = criterion(outputs, xs)
val_loss += loss.item()# / val_n
_, predicted = torch.max(outputs.data, 1)
val_total += ys.size(0)
val_correct += 0#(predicted == ys).sum().item()
val_labels.extend(ys.detach().cpu().numpy().tolist())
val_preds.extend(predicted.detach().cpu().numpy().tolist())
val_time_end = time.time()
train_time_total = train_time_end - train_time_start
val_time_total = val_time_end - val_time_start
total_time = train_time_total + val_time_total
val_losses.append(val_loss)
print(f"fold:{fold_i + 1} epoch:{epoch + 1}/{EPOCH} [tra]loss:{tr_loss:.4f} [val]loss:{val_loss:.4f} [time]total:{total_time:.2f}sec tra:{train_time_total:.2f}sec val:{val_time_total:.2f}sec")
if (epoch + 1) % 100 == 0:
savename = f"model_epoch{epoch + 1}_{EPOCH}_{FOLD}_{fold_i + 1}_{FOLD_N}.pth"
torch.save(model.state_dict(), savename)
print(f"model saved to >> {savename}")
show_sample(xs, outputs)
#---
# save model
#---
savename = f"model_epoch{EPOCH}_{FOLD}_{fold_i + 1}_{FOLD_N}.pth"
torch.save(model.state_dict(), savename)
print(f"model saved to >> {savename}")
print()
train_models.append(model)
train_model_paths.append(savename)
fig, ax1 = plt.subplots()
ax2 = ax1.twinx()
ax1.grid()
ax1.plot(train_losses, marker=".", markersize=6, color="red", label="train loss")
ax1.plot(val_losses, marker=".", markersize=6, color="blue", label="val loss")
h1, l1 = ax1.get_legend_handles_labels()
h2, l2 = ax2.get_legend_handles_labels()
ax1.legend(h1+h2, l1+l2, loc="upper right")
ax1.set(xlabel="Epoch", ylabel="Loss")
ax2.set(ylabel="Accuracy")
plt.show()
return train_models, train_model_paths
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="kWJlRDdHrxNG" outputId="ca32bab6-05e2-4067-b8c0-ba6f70837273"
train_models, train_model_paths = train()
# + [markdown] id="ODVD2zTZDYAZ"
# # Test
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="0MvosjsLDXvQ" outputId="8bcfa397-8f2d-4e62-de0f-af376e1de3ed"
# test
def test(train_models):
for model in train_models:
model.eval()
model_num = len(train_models)
test_ds = torchvision.datasets.MNIST(root="./", train=False, download=True, transform=None)
test_Xs = test_ds.data.numpy().astype(np.float32)[..., None]
test_ys = np.array(test_ds.targets)
dataset_test = MnistDataset(test_Xs, test_ys, transforms=transforms_val)
dataloader_test = DataLoader(dataset_test, batch_size=512, num_workers=4, shuffle=False, pin_memory=True)
preds = []
correct = 0
with torch.no_grad():
for step, batch in enumerate(dataloader_test):
Xs = batch[0].to(device) # image
ys = batch[1]
_preds = np.zeros([len(ys), 10])
for model in train_models:
model_preds = model(Xs)
show_sample(Xs, model_preds)
break
test(train_models)
# + id="fzuAc0vYOUZ0"
| notes_pytorch/AE/AE_MNIST_pytorch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Applying IntelligentElement to relational data
#
# Relational data uses keys to access nested structures and retrieve them across database tables.
#
# For example: a client record from a table may have `client_id`, that is used to index and retrieve multiple rows from another datatable.
#
# ## Imports
#
# Let us import the packages that will be used.
# +
#misc
# %matplotlib inline
import matplotlib.pyplot as plt
import pandas as pd
import zipfile
#learning
from keras import Model
from keras import layers as L
from keras import backend as K
import numpy as np
#add two parent levels to path
import os,sys,inspect
currentdir = os.path.dirname(os.path.abspath(inspect.getfile(inspect.currentframe())))
parentdir = os.path.dirname(os.path.dirname(currentdir))
sys.path.insert(0,parentdir)
# %load_ext autoreload
# %autoreload 2
import IntelligentElement as IE
# -
# # Load Data
#
# ## Open .csv files from zip
zip_data = zipfile.ZipFile('elodata/elo_data.zip', 'r')
allfiles = [f for f in zip_data.namelist()]
allfiles
train_data = pd.read_csv(zip_data.open('train.csv'), parse_dates=["first_active_month"]).fillna(0)
test_data = pd.read_csv(zip_data.open('test.csv'), parse_dates=["first_active_month"]).fillna(0)
train_data.head()
hist_trans = pd.read_csv(zip_data.open('historical_transactions.csv'), index_col='card_id', parse_dates=["purchase_date"]).fillna(0)
hist_trans = hist_trans.sort_index()
hist_trans.head()
new_trans = pd.read_csv(zip_data.open('new_merchant_transactions.csv'), index_col='card_id', parse_dates=["purchase_date"]).fillna(0)
new_trans = new_trans.sort_index()
new_trans.head()
mercs = pd.read_csv(zip_data.open('merchants.csv'), index_col='merchant_id').fillna(0)
mercs = mercs.sort_index()
mercs.head()
train_data.loc[1].card_id in hist_trans.index
hist_trans.loc[train_data.loc[2].card_id]
# ## Data Preprocessing
#
# ### train.csv / test.csv
# +
#get labels
y_train = train_data['target'].tolist()
y_train = np.exp(y_train) - np.exp(min(y_train))
y_train = np.log(1+y_train)
plt.figure(figsize=(15,5))
plt.hist(y_train,bins=100)
# +
from sklearn.preprocessing import LabelEncoder
def preproc_data(df):
ans = df[ ['card_id', 'feature_3'] ]
dum1 = pd.get_dummies(train_data['feature_1'],prefix='f1')
ans = pd.concat([ans,dum1],axis=1)
dum2 = pd.get_dummies(train_data['feature_2'],prefix='f2')
ans = pd.concat([ans,dum2],axis=1)
return ans
train_data_prep = preproc_data(train_data)
train_data_prep.head()
# -
# ### history.csv
# +
#hist_trans = hist_trans.head(1000)
# +
pd.options.mode.chained_assignment = None # default='warn'
#label_encoders
history_categoricals = ['category_3', 'category_2', 'category_1', 'merchant_category_id',
'subsector_id','city_id','state_id']
def preproc_history(df, label_encoders = None):
if label_encoders is None:
lbl_encs = {}
else:
lbl_encs = label_encoders
ans = df[ ['merchant_id', 'month_lag', 'purchase_amount'] ]
ans['month_lag'] = -ans['month_lag']/13
ans['authorized_flag'] = df.authorized_flag.apply(lambda x : int(x=='Y'))
ans['purchase_dayofweek'] = df.purchase_date.apply(lambda x : x.dayofweek)
ans['purchase_dayofmonth'] = df.purchase_date.apply(lambda x : x.day)
ans['purchase_month'] = df.purchase_date.apply(lambda x : x.month)
for col in history_categoricals:
if label_encoders is None:
lblenc=LabelEncoder()
ans[col] = lblenc.fit_transform(df[col].tolist())
lbl_encs[col]=lblenc
else:
ans[col] = label_encoders[col].transform(df[col].tolist())
return ans, lbl_encs
# -
hist_trans_prep, hist_trans_lbl_encs=preproc_history(hist_trans)
hist_trans_prep.head()
# ### new_merchant_transactions.csv
new_trans_prep, new_trans_lbl_encs = preproc_history(new_trans)
new_trans_prep.head()
# ### merchants.csv
merc_categoricals = ['category_4', 'category_2', 'category_1', 'city_id','state_id', 'merchant_group_id',
'merchant_category_id', 'subsector_id', 'most_recent_sales_range', 'most_recent_purchases_range']
def preproc_mercs(df):
lbl_encs = {}
ans = df[ ['numerical_1', 'numerical_2',
'avg_sales_lag3', 'avg_purchases_lag3', 'active_months_lag3',
'avg_sales_lag6', 'avg_purchases_lag6', 'active_months_lag6',
'avg_sales_lag12','avg_purchases_lag12','active_months_lag12'] ]
ans['avg_sales_lag3'] = np.log(1+ans['avg_sales_lag3'])
ans['avg_sales_lag6'] = np.log(1+ans['avg_sales_lag6'])
ans['avg_sales_lag12'] = np.log(1+ans['avg_sales_lag12'])
ans['avg_purchases_lag3'] = np.log(1+np.clip(ans['avg_purchases_lag3'],0,1e5))
ans['avg_purchases_lag12'] = np.log(1+np.clip(ans['avg_purchases_lag12'],0,1e5))
ans['avg_purchases_lag6'] = np.log(1+np.clip(ans['avg_purchases_lag6'],0,1e5))
ans['numerical_1'] = ans['numerical_1']/25.
ans['numerical_2'] = ans['numerical_2']/25.
ans['active_months_lag3'] = ans['active_months_lag3']/12.
ans['active_months_lag12'] = ans['active_months_lag12']/12.
ans['active_months_lag6'] = ans['active_months_lag6']/12.
for col in merc_categoricals:
lblenc=LabelEncoder()
ans[col] = lblenc.fit_transform(df[col].tolist())
lbl_encs[col]=lblenc
return ans, lbl_encs
mercs_prep, mercs_lblencs = preproc_mercs(mercs)
mercs_prep.head()
# # Build IntelligentElement classes
x_train = train_data.index.tolist()
# +
n_val=int( 0.1*len(y_train) )
x_val = x_train[0:n_val]
y_val = y_train[0:n_val]
x_train = x_train[n_val:]
y_train = y_train[n_val:]
# -
# ## history
def get_hist_info(x):
row = train_data_prep.iloc[x]
card_id = row.card_id
#card_id = 'C_ID_0001238066' #train_data_prep.iloc[x].card_id
#print(card_id)
idx0 = hist_trans_prep.index.searchsorted(card_id, side='left')
idxf = hist_trans_prep.index.searchsorted(card_id, side='right')
#print('{} {}'.format(idx0,idxf))
df_slice = hist_trans_prep.iloc[slice(idx0, idxf)].sort_values(by=['month_lag'])
df_slice = df_slice.drop(columns = ['merchant_id'])
df_slice['pad'] = 1 #identifies that this info exists, ie, was not padded
if idx0==idxf:
return np.zeros( (1,len(df_slice.columns)) )
else:
return np.array(df_slice.values)
# +
temp = get_hist_info(0)
inp_shape = (None, temp.shape[1])
########
# Model
########
inp = L.Input(inp_shape)
#numericals
x = L.Lambda(lambda q: q[:, :, 0:3])(inp)
#deal with categoricals
vals=[x]
curval = L.Lambda(lambda q: q[:, :, 3:4])(inp) #dayofweek
curval = L.Embedding(7,4)(curval)
curval = L.Lambda(lambda q: K.squeeze(q,axis=2) )(curval)
vals.append(curval)
curval = L.Lambda(lambda q: q[:, :, 4:5])(inp) #dayofmonth
curval = L.Embedding(31,4)(curval)
curval = L.Lambda(lambda q: K.squeeze(q,axis=2) )(curval)
vals.append(curval)
curval = L.Lambda(lambda q: q[:, :, 5:6])(inp) #month
curval = L.Embedding(12,4)(curval)
curval = L.Lambda(lambda q: K.squeeze(q,axis=2) )(curval)
vals.append(curval)
for k in range(len(history_categoricals)):
dict_size = len(set(hist_trans_lbl_encs[history_categoricals[k]].classes_))
embdim = int(np.sqrt(dict_size))+2
#print(history_categoricals[k])
#print( embdim )
pos=k+6
curval = L.Lambda(lambda q: q[:, :, pos:pos+1])(inp)
curval = L.Embedding(dict_size, embdim)(curval)
curval = L.Lambda(lambda q: K.squeeze(q,axis=2) )(curval)
vals.append(curval)
x = L.Concatenate()(vals)
x = L.Bidirectional(L.CuDNNLSTM(32))(x)
hist_model = Model(inputs=inp, outputs=x)
#hist_model.summary()
#########
# IE
#########
hist_ie = IE.IntelligentElement(x_train, hist_model, inp_shape, preprocess_function=get_hist_info, val_data=x_val, name='hist_ie')
m, ii, oo = hist_ie.retrieve_model_inputs_outputs()
print('\n\nRetrieved model')
m.summary()
# -
hist_ie.get_batch([2,12,46])[0].shape
hist_ie.get_batch([2,12,46], from_set='val')[0].shape
# ## root
# +
def get_train_info(x):
row = train_data_prep.iloc[x]
card_id = row.card_id
row = row.drop('card_id')
return np.array(row.tolist()).astype(int)
root_feats = hist_ie.model.output_shape[-1]+get_train_info(0).shape[0]
root_shape = (get_train_info(0).shape[0],)
inp = L.Input( (root_feats,) )
x=inp
for k in range(4):
x = L.Dense(4*root_feats, activation='relu')(x)
x = L.Dense(1)(x)
root_model = Model(inputs=inp, outputs=x)
root_model.summary()
#########
# IE
#########
root_ie = IE.IntelligentElement(x_train, root_model, root_shape, preprocess_function=get_train_info, val_data=x_val,
children_ie=[hist_ie], name='root_ie')
m, ii, oo = root_ie.retrieve_model_inputs_outputs()
print('\n\nRetrieved model')
m.summary()
# -
len(root_ie.get_batch([1,10,20]))
# # Data Generation and Model Training
train_gen = IE.IEDataGenerator(root_ie, y_train, labeltype=float)
val_gen = IE.IEDataGenerator(root_ie, y_val, labeltype=float, from_set = 'val')
xx,yy = train_gen.__getitem__(0)
print('{} {} {}'.format(xx[0].shape, xx[1].shape, yy.shape))
# +
from keras.optimizers import Adam
#m.compile(optimizer=Adam(lr=4e-4, clipvalue=0.5), loss='mean_squared_error') #, metrics=[mean_iou, 'categorical_accuracy'])
m.compile(optimizer=Adam(lr=4e-4, clipvalue=0.5), loss='logcosh') #, metrics=[mean_iou, 'categorical_accuracy'])
# +
from keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau
earlystopper = EarlyStopping(patience=10, verbose=1, monitor='val_loss')
checkpointer = ModelCheckpoint('model-reldb.h5', verbose=1, save_best_only=True, monitor='val_loss')
reduce_lr = ReduceLROnPlateau(factor=0.5, patience=5, min_lr=1e-5, verbose=1, monitor='loss')
# -
results = m.fit_generator(train_gen, epochs=100,
#use_multiprocessing = False, workers=4,
validation_data=val_gen,
callbacks=[earlystopper, checkpointer, reduce_lr])
# # Tests and Verifications
new_trans_prep, temp = preproc_history(new_trans)
new_trans_prep.head()
idx = mercs_prep.index.searchsorted('M_ID_0000699140')
mercs_prep.iloc[idx]
new_trans.index.searchsorted('C_ID_0001238066', side='left')
new_trans.index.searchsorted('C_ID_0001238066', side='right')
sl = slice(2,28)
new_trans.iloc[sl]
slist = s.tolist()
slist[0:3]
train_data.iloc[150]
| examples/relationaldb/Apply IntelligentElement to relational data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="7ip_xEFS5F5A"
#description: This is a 'smart' chat bot program
# + colab={"base_uri": "https://localhost:8080/"} id="_V4nxoMp7J0k" outputId="8e42f4e0-ab2c-48e0-ad32-e972a3b51ee8"
pip install nltk
# + colab={"base_uri": "https://localhost:8080/"} id="g-JHMKoF7T-O" outputId="ce09d791-b90f-4de1-cb9a-d2778f2ffa7d"
pip install newspaper3k
# + id="yf1zxMky7eET"
#import the libraries
from newspaper import Article
import random
import string
import nltk
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics.pairwise import cosine_similarity
import numpy as np
import warnings
warnings.filterwarnings('ignore')
# + colab={"base_uri": "https://localhost:8080/"} id="L7hoHwCy-HZL" outputId="49644baa-2e19-4879-d6ec-664e9a2162ef"
#Download the punkt package
nltk.download('punkt', quiet=True)
# + id="KS6UOHaB-k-l"
#Get the article
article = Article('https://www.mayoclinic.org/diseases-conditions/chronic-kidney-disease/symptoms-causes/syc-20354521')
article.download()
article.parse()
article.nlp
corpus = article.text
# + colab={"base_uri": "https://localhost:8080/"} id="9WgtqhOE_lIG" outputId="35b41023-b1da-4629-967f-c79385f2b257"
#Print the articles text
print(corpus)
# + id="bcNNqY-Q_6i8"
#Tokenization
text = corpus
sentense_list = nltk.sent_tokenize(text) #A list of sentences
# + colab={"base_uri": "https://localhost:8080/"} id="Xgt8va2JAm9F" outputId="65946bdb-338c-45e0-b10a-3dd3537a92b8"
#Print the list of sentences
print(sentense_list)
# + id="0d1ajGE4Bzm-"
# A function to return a random greeting response to a users greeting
def greeting_response(text):
text = text.lower
#Bots greeting response
bot_greeting = [ "howdy" ,'hi', 'hello', 'hola']
#users greeting
user_greeting = ['hi', 'hey', 'hello','hola', 'greetings', 'wassup']
for word in text.split():
if word in user_greeting:
return random.choice(bot_greeting)
# + id="ckBib9CZEsFZ"
def index_sort(list_var):
length = len(list_var)
list_index = list(range(0), length)
x = list_var
for i in range(length):
for j in range(length):
if x[list_index[i]] > x[list_index[j]]:
#swap
temp = list_index[i]
list_index[i] = list_index[j]
list_index[j] = temp
return list_index
# + id="nSRBv7WkDSiX"
#Create the bots response
def bot_response(user_input):
user_input = user_input.lower()
sentense_list.append(user_input)
bot_response = ''
cm = CountVectorizer().fit_transform(sentence_list)
similarity_scores = cosine_similarity(cm[-1], cm)
similarity_scores_list = similarity_scores.flatten()
index = index_sort(similarity_scores_list)
index = index[1:]
response_flag = 0
j = 0
for i in range(len(index)):
if similarity_scores_list[index[i]] > 0.0:
bot_response = bot_response+' '+sentense_list[index[i]]
response_flag = 1
j = j+1
if j > 2:
break
if response_flag ==0:
bot_response = bot_response+ ' '+"I apologize, I don't understand"
sentense_list.remove(user_input)
return bot_response
# + colab={"base_uri": "https://localhost:8080/"} id="YAbgvTWbC60b" outputId="c757a55b-46e6-4ebb-8292-cca31a8ce010"
#Start the chat
print('Doc bot: i am a doctor bot or Doc bot for short. i will answer your quiries about heart disease.If you want to exit, type bye.')
exit_list = ['exit', 'see you later', 'bye', 'quite','break']
while(True):
user_input = input()
if user_input.lower() in exit_list:
print('Doc bot: chat with you later !')
break
else:
if greeting_response(user_input) != None:
print('Doc bot: '+greeting_response(user_input))
else:
print('Doc bot: '+bot_response(user_input))
| A_Smart_AI_Chat_Bot_Using_Python_&_Machine_Learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finetuning of the pretrained Japanese BERT model
#
# Finetune the pretrained model to solve multi-class classification problems.
# This notebook requires the following objects:
# - trained sentencepiece model (model and vocab files)
# - pretraiend Japanese BERT model
#
# We make test:dev:train = 2:2:6 datasets.
# +
import configparser
import glob
import os
import pandas as pd
import subprocess
import sys
import tarfile
import pandas as pd
from urllib.request import urlretrieve
CURDIR = os.getcwd()
CONFIGPATH = os.path.join(CURDIR, os.pardir, 'config.ini')
config = configparser.ConfigParser()
config.read(CONFIGPATH)
# -
# ## Data preparing
#
# You need execute the following cells just once.
FILEURL = config['HIS-DATA']['DATADIR']
EXTRACTDIR = config['HIS-DATA']['TEXTDIR']
df = pd.read_csv(FILEURL, delimiter='\t')
df.head()
# Save data as tsv files.
# test:dev:train = 2:2:6. To check the usability of finetuning, we also prepare sampled training data (1/5 of full training data).
# +
df[:len(df) // 10].to_csv( os.path.join(EXTRACTDIR, "test.tsv"), sep='\t', index=False)
df[len(df) // 10:len(df)*2 // 10].to_csv( os.path.join(EXTRACTDIR, "dev.tsv"), sep='\t', index=False)
df[len(df)*2 // 10:].to_csv( os.path.join(EXTRACTDIR, "train.tsv"), sep='\t', index=False)
### 1/5 of full training data.
# df[:len(df) // 5].to_csv( os.path.join(EXTRACTDIR, "test.tsv"), sep='\t', index=False)
# df[len(df) // 5:len(df)*2 // 5].to_csv( os.path.join(EXTRACTDIR, "dev.tsv"), sep='\t', index=False)
# df[len(df)*2 // 5:].sample(frac=0.2, random_state=23).to_csv( os.path.join(EXTRACTDIR, "train.tsv"), sep='\t', index=False)
# -
# ## Finetune pre-trained model
#
# It will take a lot of hours to execute the following cells on CPU environment.
# You can also use colab to recieve the power of TPU. You need to uplode the created data onto your GCS bucket.
#
# [](https://colab.research.google.com/drive/1zZH2GWe0U-7GjJ2w2duodFfEUptvHjcx)
PRETRAINED_MODEL_PATH = '../model/model.ckpt-1000000'
FINETUNE_OUTPUT_DIR = '../model/his_output'
output_ckpts = glob.glob("{}/model.ckpt*data*".format(FINETUNE_OUTPUT_DIR))
latest_ckpt = sorted(output_ckpts)[-1]
PRETRAINED_MODEL_PATH = latest_ckpt.split('.data-00000-of-00001')[0]
PRETRAINED_MODEL_PATH
# +
# %%time
# It will take many hours on CPU environment.
# !python3 ../src/run_classifier.py \
# --task_name=his \
# --do_train=true \
# --do_eval=true \
# --data_dir=../data/his \
# --model_file=../model/wiki-ja.model \
# --vocab_file=../model/wiki-ja.vocab \
# --init_checkpoint={PRETRAINED_MODEL_PATH} \
# --max_seq_length=32 \
# --train_batch_size=16 \
# --learning_rate=5e-5 \
# --num_train_epochs=20 \
# --output_dir={FINETUNE_OUTPUT_DIR}
# -
# ## Predict using the finetuned model
#
# Let's predict test data using the finetuned model.
# +
import sys
sys.path.append("../src")
import tokenization_sentencepiece as tokenization
from run_classifier import HISProcessor
from run_classifier import model_fn_builder
from run_classifier import file_based_input_fn_builder
from run_classifier import file_based_convert_examples_to_features
from utils import str_to_value
# +
sys.path.append("../bert")
import modeling
import optimization
import tensorflow as tf
# +
import configparser
import json
import glob
import os
import pandas as pd
import tempfile
bert_config_file = tempfile.NamedTemporaryFile(mode='w+t', encoding='utf-8', suffix='.json')
bert_config_file.write(json.dumps({k:str_to_value(v) for k,v in config['BERT-CONFIG'].items()}))
bert_config_file.seek(0)
bert_config = modeling.BertConfig.from_json_file(bert_config_file.name)
# -
output_ckpts = glob.glob("{}/model.ckpt*data*".format(FINETUNE_OUTPUT_DIR))
latest_ckpt = sorted(output_ckpts)[-1]
FINETUNED_MODEL_PATH = latest_ckpt.split('.data-00000-of-00001')[0]
# FINETUNED_MODEL_PATH = '../model/his_output/model.ckpt-33000'
FINETUNED_MODEL_PATH
class FLAGS(object):
'''Parameters.'''
def __init__(self):
self.model_file = "../model/wiki-ja.model"
self.vocab_file = "../model/wiki-ja.vocab"
self.do_lower_case = True
self.use_tpu = False
self.output_dir = "/dummy"
self.data_dir = "../data/his"
self.max_seq_length = 64
self.init_checkpoint = FINETUNED_MODEL_PATH
self.predict_batch_size = 4
# The following parameters are not used in predictions.
# Just use to create RunConfig.
self.master = None
self.save_checkpoints_steps = 1
self.iterations_per_loop = 1
self.num_tpu_cores = 1
self.learning_rate = 0
self.num_warmup_steps = 0
self.num_train_steps = 0
self.train_batch_size = 0
self.eval_batch_size = 0
FLAGS = FLAGS()
processor = HISProcessor()
label_list = processor.get_labels()
# +
tokenizer = tokenization.FullTokenizer(
model_file=FLAGS.model_file, vocab_file=FLAGS.vocab_file,
do_lower_case=FLAGS.do_lower_case)
tpu_cluster_resolver = None
is_per_host = tf.contrib.tpu.InputPipelineConfig.PER_HOST_V2
run_config = tf.contrib.tpu.RunConfig(
cluster=tpu_cluster_resolver,
master=FLAGS.master,
model_dir=FLAGS.output_dir,
save_checkpoints_steps=FLAGS.save_checkpoints_steps,
tpu_config=tf.contrib.tpu.TPUConfig(
iterations_per_loop=FLAGS.iterations_per_loop,
num_shards=FLAGS.num_tpu_cores,
per_host_input_for_training=is_per_host))
# +
model_fn = model_fn_builder(
bert_config=bert_config,
num_labels=len(label_list),
init_checkpoint=FLAGS.init_checkpoint,
learning_rate=FLAGS.learning_rate,
num_train_steps=FLAGS.num_train_steps,
num_warmup_steps=FLAGS.num_warmup_steps,
use_tpu=FLAGS.use_tpu,
use_one_hot_embeddings=FLAGS.use_tpu)
estimator = tf.contrib.tpu.TPUEstimator(
use_tpu=FLAGS.use_tpu,
model_fn=model_fn,
config=run_config,
train_batch_size=FLAGS.train_batch_size,
eval_batch_size=FLAGS.eval_batch_size,
predict_batch_size=FLAGS.predict_batch_size)
# +
predict_examples = processor.get_test_examples(FLAGS.data_dir)
predict_file = tempfile.NamedTemporaryFile(mode='w+t', encoding='utf-8', suffix='.tf_record')
file_based_convert_examples_to_features(predict_examples, label_list,
FLAGS.max_seq_length, tokenizer,
predict_file.name)
predict_drop_remainder = True if FLAGS.use_tpu else False
predict_input_fn = file_based_input_fn_builder(
input_file=predict_file.name,
seq_length=FLAGS.max_seq_length,
is_training=False,
drop_remainder=predict_drop_remainder)
# -
result = estimator.predict(input_fn=predict_input_fn)
# +
# %%time
# It will take a few hours on CPU environment.
result = list(result)
# -
result[:2]
# Read test data set and add prediction results.
import pandas as pd
test_df = pd.read_csv("../data/his/test.tsv", sep='\t')
test_df['predict'] = [ label_list[elem['probabilities'].argmax()] for elem in result ]
test_df.head(10)
sum( test_df['label'] == test_df['predict'] ) / len(test_df)
# A littel more detailed check using `sklearn.metrics`.
# !pip install scikit-learn
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
print(classification_report(test_df['label'], test_df['predict']))
print(confusion_matrix(test_df['label'], test_df['predict']))
# ### Simple baseline model.
import pandas as pd
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
train_df = pd.read_csv("../data/his/train.tsv", sep='\t')
dev_df = pd.read_csv("../data/his/dev.tsv", sep='\t')
test_df = pd.read_csv("../data/his/test.tsv", sep='\t')
# !sudo apt-get install -q -y mecab libmecab-dev mecab-ipadic mecab-ipadic-utf8
# !pip install mecab-python3==0.7
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.ensemble import GradientBoostingClassifier
import MeCab
m = MeCab.Tagger("-Owakati")
train_dev_df = pd.concat([train_df, dev_df])
# +
train_dev_xs = train_dev_df['text'].apply(lambda x: m.parse(x))
train_dev_ys = train_dev_df['label']
test_xs = test_df['text'].apply(lambda x: m.parse(x))
test_ys = test_df['label']
# -
vectorizer = TfidfVectorizer(max_features=750)
train_dev_xs_ = vectorizer.fit_transform(train_dev_xs)
test_xs_ = vectorizer.transform(test_xs)
# The following set up is not exactly identical to that of BERT because inside Classifier it uses `train_test_split` with shuffle.
# In addition, parameters are not well tuned, however, we think it's enough to check the power of BERT.
# +
# %%time
# model = GradientBoostingClassifier(n_estimators=200,
# validation_fraction=len(train_df)/len(dev_df),
# n_iter_no_change=5,
# tol=0.01,
# random_state=23)
### 1/5 of full training data.
# model = GradientBoostingClassifier(n_estimators=200,
# validation_fraction=len(dev_df)/len(train_df),
# n_iter_no_change=5,
# tol=0.01,
# random_state=23)
from sklearn.svm import LinearSVC
# from sklearn.linear_model import LogisticRegression
# from sklearn.ensemble import RandomForestClassifier
model = LinearSVC()
model.fit(train_dev_xs_, train_dev_ys)
# -
print(classification_report(test_ys, model.predict(test_xs_)))
print(confusion_matrix(test_ys, model.predict(test_xs_)))
| notebook/finetune-to-his-corpus.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finding out the impact of the news
# Here we are calculating the Mean and standard deviation of the volume before the news broke out and mean of the volume after the news broke out. If the news came on a Saturday or Sunday (when the market is closed), we ignore the news.
#
# If the mean volume value was higher, we know that it created some impact on the market.
# +
import numpy as np
import pandas as pd
import csv
time1=[]
time2=[]
def check(time):
try:
s = int(time[11:13]+time[14:16])
if(s>=945 and s<=1459):
return True
else:
return False
except:
print ("except")
pass
return time
index = 0
arr = np.ones(30)
file = open(r'BANK OF BARODA Quant.csv','r',encoding='ISO-8859-1')
file1= open(r'Bank of Baroda News.csv','r',encoding='ISO-8859-1')
new = open(r'Impact.csv','a',newline='',encoding='ISO-8859-1')
writ=csv.writer(new)
writ.writerow(['Time','Title','Day','MeanVol-30','StddevVol-30','MeanVol+30'])
read1=csv.reader(file1)
next(read1, None)
for j in read1:
index = 0
count =0
sum=0
flag=0
arr = np.ones(30)
file = open(r'BANK OF BARODA Quant.csv','r',encoding='ISO-8859-1')
read=csv.reader(file)
next(read, None)
temp=pd.to_datetime(j[3])
day = temp.strftime("%A")
if day == 'Sunday' or day =='Saturday':
print ('Ignored')
continue
if(not check(j[3])):
continue
for i in read:
arr[(index)%30] = float(i[6])
index+=1
if flag==1:
sum+=float(i[6])
count +=1
if count ==30:
writ.writerow([j[3],j[5],day,mean,stddev,(sum/30)])
break
if j[3] in i[1]:
flag = 1
print(i[1])
print(arr)
stddev = np.std(arr,axis=0)
mean = np.mean(arr,axis=0)
file.close()
# -
new.close()
| Merging quant and news/Finding news impact/NewsImpact.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import xarray as xr
from mlmicrophysics.models import DenseNeuralNetwork
from sklearn.ensemble import RandomForestClassifier
from mlmicrophysics.explain import partial_dependence_2d
from mlmicrophysics.data import subset_data_files_by_date, log10_transform, neg_log10_transform, assemble_data_files
from sklearn.preprocessing import StandardScaler, RobustScaler, MaxAbsScaler, MinMaxScaler
from sklearn.metrics import r2_score, confusion_matrix, mean_squared_error
from keras.models import save_model
from keras.layers import Input, Dense, Dropout, GaussianNoise, Activation, Concatenate, BatchNormalization
from keras.models import Model
from keras.regularizers import l2
from keras.optimizers import Adam, SGD
import keras.backend as K
import yaml
from matplotlib.colors import LogNorm
scalers = {"MinMaxScaler": MinMaxScaler,
"MaxAbsScaler": MaxAbsScaler,
"StandardScaler": StandardScaler,
"RobustScaler": RobustScaler}
transforms = {"log10_transform": log10_transform,
"neg_log10_transform": neg_log10_transform}
import call_collect
# ls /glade/p/cisl/aiml/dgagne/cam_mp_files_run2_csv/
# # Data Loading
path = "/glade/p/cisl/aiml/dgagne/cam_mp_files_run2_csv/"
#path = "/Users/dgagne/data/cam_mp_files_run2_csv/"
train_files, val_files, test_files = subset_data_files_by_date(path, "_qc_*.csv", train_date_start=0, train_date_end=8000,
test_date_start=8100, test_date_end=16000)
train_files_qr, val_files_qr, test_files_qr = subset_data_files_by_date(path, "_qr_*.csv", train_date_start=0, train_date_end=8000,
test_date_start=8100, test_date_end=16000)
input_transforms_yaml = """
RHO_CLUBB_lev: log10_transform
QC_TAU_in: log10_transform
NC_TAU_in: log10_transform
QR_TAU_in: log10_transform
NR_TAU_in: log10_transform
"""
output_transforms_yaml = """
qctend_TAU:
0: [">=", -1e-18, "zero_transform", "None"]
1: ["<", -1e-18, "neg_log10_transform", "StandardScaler"]
qrtend_TAU:
0: ["<=", 1e-18, "zero_transform", "None"]
1: [">", 1e-18, "log10_transform", "StandardScaler"]
nctend_TAU:
0: [">=", -1e-18, "zero_transform", "None"]
1: ["<", -1e-18, "neg_log10_transform", "StandardScaler"]
nrtend_TAU:
-1: ["<", 0, "neg_log10_transform", "StandardScaler"]
0: ["==", 0, "zero_transform", "None"]
1: [">", 0, "log10_transform", "StandardScaler"]
qctend_MG2:
0: [">=", -1e-18, "zero_transform", "None"]
1: ["<", -1e-18, "neg_log10_transform", "StandardScaler"]
qrtend_MG2:
0: ["<=", 1e-18, "zero_transform", "None"]
1: [">", 1e-18, "log10_transform", "StandardScaler"]
nctend_MG2:
0: [">=", -1e-18, "zero_transform", "None"]
1: ["<", -1e-18, "neg_log10_transform", "StandardScaler"]
nrtend_MG2:
-1: ["<", 0, "neg_log10_transform", "StandardScaler"]
0: ["==", 0, "zero_transform", "None"]
1: [">", 0, "log10_transform", "StandardScaler"]
"""
input_transforms = yaml.load(input_transforms_yaml)
output_transforms = yaml.load(output_transforms_yaml)
input_transforms
# +
input_scaler = StandardScaler()
input_cols = ["QC_TAU_in", "QR_TAU_in", "NC_TAU_in", "NR_TAU_in", "RHO_CLUBB_lev"]
output_cols = ["qctend_TAU", "qrtend_TAU", "nctend_TAU", "nrtend_TAU",
"qctend_MG2", "qrtend_MG2", "nctend_MG2", "nrtend_MG2"]
scaled_input_train,\
labels_train,\
transformed_out_train,\
scaled_out_train,\
output_scalers = assemble_data_files(train_files, input_cols, output_cols, input_transforms,
output_transforms, input_scaler, subsample=0.1)
scaled_input_train_qr,\
labels_train_qr,\
transformed_out_train_qr,\
scaled_out_train_qr,\
output_scalers_qr = assemble_data_files(train_files_qr, input_cols, output_cols, input_transforms,
output_transforms, input_scaler, subsample=0.1)
# -
original_out_train_nc = np.zeros(scaled_out_train_qr.shape[0])
original_out_train_nc[labels_train_qr["nctend_TAU"] == 1] = -10 ** output_scalers_qr["nctend_TAU"][1].inverse_transform(
scaled_out_train_qr.loc[labels_train_qr["nctend_TAU"] == 1, ["nctend_TAU"]]).ravel()
plt.hist(np.log10(-original_out_train_nc[original_out_train_nc < 0]), bins=50)
plt.gca().set_yscale("log")
scaled_input_train_qr.hist()
scaled_input_train.hist()
input_scaler_df = pd.DataFrame({"mean":input_scaler.mean_, "scale": input_scaler.scale_},
index=input_cols)
#input_scaler_df.to_csv("/glade/p/cisl/aiml/dgagne/cam_run2_models/input_scale_values.csv",
# index_label="input")
input_scaler_df
input_scaler_df2 = pd.read_csv("/glade/p/cisl/aiml/dgagne/cam_run2_models/input_scale_values.csv", index_col="input")
input_scaler_df2
out_scales_list = []
for var in output_scalers.keys():
for out_class in output_scalers[var].keys():
print(var, out_class)
if output_scalers[var][out_class] is not None:
out_scales_list.append(pd.DataFrame({"mean": output_scalers[var][out_class].mean_,
"scale": output_scalers[var][out_class].scale_},
index=[var + "_" + str(out_class)]))
out_scales_df = pd.concat(out_scales_list)
print(out_scales_df)
#out_scales_df.to_csv("/glade/p/cisl/aiml/dgagne/cam_run2_models/output_scale_values.csv",
# index_label="output")
out_scales_list = []
for var in output_scalers.keys():
for out_class in output_scalers[var].keys():
print(var, out_class)
if output_scalers[var][out_class] is not None:
out_scales_list.append(pd.DataFrame({"mean": output_scalers_qr[var][out_class].mean_,
"scale": output_scalers_qr[var][out_class].scale_},
index=[var + "_" + str(out_class)]))
out_scales_df = pd.concat(out_scales_list)
print(out_scales_df)
#out_scales_df.to_csv("/glade/p/cisl/aiml/dgagne/cam_run2_models/output_scale_values.csv",
# index_label="output")
out_scales_df2 = pd.read_csv("/glade/p/cisl/aiml/dgagne/cam_run2_models/output_scale_values.csv", index_col="output")
out_scales_df2
scaled_input_test,\
labels_test,\
transformed_out_test,\
scaled_out_test,\
output_scalers_test = assemble_data_files(test_files, input_cols, output_cols, input_transforms,
output_transforms, input_scaler, output_scalers=output_scalers,
subsample=0.1)
original_input_train = pd.DataFrame(10 ** input_scaler.inverse_transform(scaled_input_train),
columns=scaled_input_train.columns)
# +
pd_bins = 20
pd_vals = np.zeros((len(input_cols), pd_bins))
for v, var in enumerate(input_cols):
pd_vals[v] = np.logspace(np.log10(original_input_train[var].min()), np.log10(original_input_train[var].max()), pd_bins)
# -
pd_bins = 20
pd_vals = np.zeros((len(input_cols), pd_bins))
pd_grid_tau = np.zeros((len(input_cols), pd_bins, 5))
for v, var in enumerate(input_cols):
print(var)
pd_vals[v] = np.logspace(np.log10(original_input_train[var].min()), np.log10(original_input_train[var].max()), pd_bins)
train_copy = original_input_train.loc[labels_train["qctend_TAU"] == 1].copy()
for p, pd_val in enumerate(pd_vals[v]):
print(p, pd_val)
train_copy.loc[:, var] = pd_val
out_vals = np.array(call_collect.call_collect(1800.0,
np.ones(train_copy.shape[0]) * 12,
train_copy["RHO_CLUBB_lev"].values,
train_copy["QC_TAU_in"].values,
train_copy["NC_TAU_in"].values,
train_copy["QR_TAU_in"].values,
train_copy["NR_TAU_in"].values))
out_vals[0][out_vals[0] > -1e-15] = 0
print(out_vals.shape)
pd_grid_tau[v, p, 0] = np.log10(-out_vals[0][out_vals[0] < 0]).mean()
pd_grid_tau[v, p, 1] = np.log10(-out_vals[1][out_vals[1] < 0]).mean()
pd_grid_tau[v, p, 2] = np.log10(out_vals[2][out_vals[2] > 0]).mean()
pd_grid_tau[v, p, 3] = np.log10(out_vals[3][out_vals[3] > 0]).mean()
pd_grid_tau[v, p, 4] = np.log10(-out_vals[3][out_vals[3]< 0]).mean()
out_var_names = ["QC", "NC", "QR", "NR"]
for o, out_var_name in enumerate(out_var_names):
fig, axes = plt.subplots(3, 2, figsize=(6, 9))
plt.subplots_adjust(hspace=0.3, wspace=0.3)
for a, ax in enumerate(axes.ravel()):
ax.plot(np.log10(pd_vals[a]), pd_grid_tau[a, :, o], '-')
ax.set_xlabel(input_cols[a].replace("_", " "))
fig.suptitle(f"TAU Microphysics Partial Dependence {out_var_name}", fontsize=14, y=0.93)
plt.savefig(f"tau_partial_dependence_{out_var_name.lower()}.png", bbox_inches="tight", dpi=200)
plt.close()
print(call_collect.call_collect.__doc__)
qctend_tau, nctend_tau, qrtend_tau, nrtend_tau = call_collect.call_collect(1800.0,
np.ones(original_input_train["RHO_CLUBB_lev"].shape[0]) * 273.0,
original_input_train["RHO_CLUBB_lev"].values,
original_input_train["QC_TAU_in"].values,
original_input_train["NC_TAU_in"].values,
original_input_train["QR_TAU_in"].values,
original_input_train["NR_TAU_in"].values)
fig, axes = plt.subplots(3,2, figsize=(10, 10))
transformed_input_train = pd.DataFrame(input_scaler.inverse_transform(scaled_input_train), columns=input_cols)
for a, ax in enumerate(axes.ravel()):
if a < len(input_cols):
ax.set_yscale("log")
ax.hist(transformed_input_train[input_cols[a]], bins=20)
ax.set_title(input_cols[a])
class DenseNeuralNetwork(object):
"""
A Dense Neural Network Model that can support arbitrary numbers of hidden layers.
Attributes:
hidden_layers: Number of hidden layers
hidden_neurons: Number of neurons in each hidden layer
inputs: Number of input values
outputs: Number of output values
activation: Type of activation function
output_activation: Activation function applied to the output layer
optimizer: Name of optimizer or optimizer object.
loss: Name of loss function or loss object
use_noise: Whether or not additive Gaussian noise layers are included in the network
noise_sd: The standard deviation of the Gaussian noise layers
use_dropout: Whether or not Dropout layers are added to the network
dropout_alpha: proportion of neurons randomly set to 0.
batch_size: Number of examples per batch
epochs: Number of epochs to train
verbose: Level of detail to provide during training
model: Keras Model object
"""
def __init__(self, hidden_layers=1, hidden_neurons=4, activation="relu",
output_activation="linear", optimizer="adam", loss="mse", use_noise=False, noise_sd=0.01,
lr=0.001, use_dropout=False, dropout_alpha=0.1, batch_size=128, epochs=2,
l2_weight=0.01, sgd_momentum=0.9, adam_beta_1=0.9, adam_beta_2=0.999, decay=0, verbose=0,
classifier=False):
self.hidden_layers = hidden_layers
self.hidden_neurons = hidden_neurons
self.activation = activation
self.output_activation = output_activation
self.optimizer = optimizer
self.optimizer_obj = None
self.sgd_momentum = sgd_momentum
self.adam_beta_1 = adam_beta_1
self.adam_beta_2 = adam_beta_2
self.loss = loss
self.lr = lr
self.l2_weight = l2_weight
self.batch_size = batch_size
self.use_noise = use_noise
self.noise_sd = noise_sd
self.use_dropout = use_dropout
self.dropout_alpha = dropout_alpha
self.epochs = epochs
self.decay = decay
self.verbose = verbose
self.classifier = classifier
self.y_labels = None
self.model = None
self.optimizer_obj = None
def build_neural_network(self, inputs, outputs):
"""
Create Keras neural network model and compile it.
Args:
inputs (int): Number of input predictor variables
outputs (int): Number of output predictor variables
"""
nn_input = Input(shape=(inputs,), name="input")
nn_model = nn_input
for h in range(self.hidden_layers):
nn_model = Dense(self.hidden_neurons, activation=self.activation,
kernel_regularizer=l2(self.l2_weight), name=f"dense_{h:02d}")(nn_model)
if self.use_dropout:
nn_model = Dropout(self.dropout_alpha, name=f"dropout_h_{h:02d}")(nn_model)
if self.use_noise:
nn_model = GaussianNoise(self.noise_sd, name=f"ganoise_h_{h:02d}")(nn_model)
nn_model = Dense(outputs,
activation=self.output_activation, name=f"dense_{self.hidden_layers:02d}")(nn_model)
self.model = Model(nn_input, nn_model)
if self.optimizer == "adam":
self.optimizer_obj = Adam(lr=self.lr, beta_1=self.adam_beta_1, beta_2=self.adam_beta_2, decay=self.decay)
elif self.optimizer == "sgd":
self.optimizer_obj = SGD(lr=self.lr, momentum=self.sgd_momentum, decay=self.decay)
self.model.compile(optimizer=self.optimizer, loss=self.loss)
def fit(self, x, y):
inputs = x.shape[1]
if len(y.shape) == 1:
outputs = 1
else:
outputs = y.shape[1]
if self.classifier:
outputs = np.unique(y).size
self.build_neural_network(inputs, outputs)
if self.classifier:
self.y_labels = np.unique(y)
y_class = np.zeros((y.shape[0], self.y_labels.size), dtype=np.int32)
for l, label in enumerate(self.y_labels):
y_class[y == label, l] = 1
self.model.fit(x, y_class, batch_size=self.batch_size, epochs=self.epochs, verbose=self.verbose)
else:
self.model.fit(x, y, batch_size=self.batch_size, epochs=self.epochs, verbose=self.verbose)
return
def save_fortran_model(self, filename):
nn_ds = xr.Dataset()
num_dense = 0
layer_names = []
for layer in self.model.layers:
if "dense" in layer.name:
layer_names.append(layer.name)
dense_weights = layer.get_weights()
nn_ds[layer.name + "_weights"] = ((layer.name + "_in", layer.name + "_out"), dense_weights[0])
nn_ds[layer.name + "_bias"] = ((layer.name + "_out",), dense_weights[1])
nn_ds[layer.name + "_weights"].attrs["name"] = layer.name
nn_ds[layer.name + "_weights"].attrs["activation"] = layer.get_config()["activation"]
num_dense += 1
nn_ds["layer_names"] = (("num_layers",), np.array(layer_names))
nn_ds.attrs["num_layers"] = num_dense
nn_ds.to_netcdf(filename, encoding={'layer_names':{'dtype': 'S1'}})
return
def predict(self, x):
if self.classifier:
y_prob = self.model.predict(x, batch_size=self.batch_size)
y_out = self.y_labels[np.argmax(y_prob, axis=1)].ravel()
else:
y_out = self.model.predict(x, batch_size=self.batch_size).ravel()
return y_out
def predict_proba(self, x):
y_prob = self.model.predict(x, batch_size=self.batch_size)
return y_prob
# # Model Training
# +
dnn_qr_class = DenseNeuralNetwork(hidden_layers=6, hidden_neurons=30,
loss="categorical_crossentropy",
output_activation="softmax",
activation="relu", epochs=10,
batch_size=2048, verbose=1, lr=0.0001, l2_weight=0.0001, classifier=True)
dnn_nc_class = DenseNeuralNetwork(hidden_layers=6, hidden_neurons=10,
loss="categorical_crossentropy",
output_activation="softmax",
activation="relu", epochs=10,
batch_size=2048, verbose=1, lr=0.0001, l2_weight=0.0001, classifier=True)
dnn_qr = DenseNeuralNetwork(hidden_layers=6, hidden_neurons=30,
loss="mse", activation="relu", epochs=10,
batch_size=2048, verbose=1, lr=0.0001, l2_weight=0.0001)
dnn_nc = DenseNeuralNetwork(hidden_layers=6, hidden_neurons=30,
loss="mse", activation="relu", epochs=10,
batch_size=2048, verbose=1, l2_weight=0.0001, lr=0.0001)
# -
dnn_qr_class.fit(scaled_input_train, labels_train["qrtend_TAU"])
dnn_qr.fit(scaled_input_train.loc[labels_train["qrtend_TAU"] == 1],
scaled_out_train.loc[labels_train["qrtend_TAU"] == 1, "qrtend_TAU"].values)
dnn_nc.fit(scaled_input_train, scaled_out_train.loc[:, "nctend_TAU"])
dnn_nc_class.fit(scaled_input_train, labels_train["nctend_TAU"])
save_model(dnn_nc.model, "/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nc.h5")
save_model(dnn_qr.model, "/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_qr.h5")
save_model(dnn_nc_class.model, "/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nc_class.h5")
save_model(dnn_qr_class.model, "/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_qr_class.h5")
dnn_qr_class.save_fortran_model("/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_qr_class_fortran.nc")
dnn_qr.save_fortran_model("/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_qr_fortran.nc")
dnn_nc_class.save_fortran_model("/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nc_class_fortran.nc")
dnn_nc.save_fortran_model("/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nc_fortran.nc")
dnn_nr_class = DenseNeuralNetwork(hidden_layers=6, hidden_neurons=30,
loss="categorical_crossentropy",
output_activation="softmax",
activation="relu", epochs=10,
batch_size=2048, verbose=1, l2_weight=0.0001, lr=0.0001, classifier=True)
dnn_nr_class.fit(scaled_input_train, labels_train["nrtend_TAU"])
save_model(dnn_nr_class.model, "/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nr_class.h5")
dnn_nr_class.save_fortran_model("/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nr_class_fortran.nc")
test_nr_sign = dnn_nr_class.predict(scaled_input_test)
# +
cf = confusion_matrix(labels_test["nrtend_TAU"].values, test_nr_sign)
print(cf / cf.sum())
# -
cm_qr = confusion_matrix(labels_test["qrtend_TAU"].values, dnn_qr_class.predict(scaled_input_test))
cm_qr / cm_qr.sum()
nr_pos_net = DenseNeuralNetwork(hidden_layers=6, hidden_neurons=30,
loss="mse", activation="relu", epochs=10,
batch_size=2048, verbose=1, l2_weight=0.0001, lr=0.0001)
nr_neg_net = DenseNeuralNetwork(hidden_layers=6, hidden_neurons=30,
loss="mse", activation="relu", epochs=10,
batch_size=2048, verbose=1, l2_weight=0.0001, lr=0.0001)
pos_sub = labels_train["nrtend_TAU"] == 1
neg_sub = labels_train["nrtend_TAU"] == -1
nr_pos_net.fit(scaled_input_train.loc[pos_sub], scaled_out_train.loc[pos_sub, "nrtend_TAU"])
nr_neg_net.fit(scaled_input_train.loc[neg_sub], scaled_out_train.loc[neg_sub, "nrtend_TAU"])
save_model(nr_pos_net.model, "/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nr_pos.h5")
save_model(nr_neg_net.model, "/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nr_neg.h5")
nr_pos_net.save_fortran_model("/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nr_pos_fortran.nc")
nr_neg_net.save_fortran_model("/glade/p/cisl/aiml/dgagne/cam_run2_models/dnn_nr_neg_fortran.nc")
# ls -ltr /glade/p/cisl/aiml/dgagne/cam_run2_models
nr_pred_values = np.zeros(scaled_input_test.shape[0])
nr_pred_values[test_nr_sign > 0] = 10 ** output_scalers["nrtend_TAU"][1].inverse_transform(nr_pos_net.predict(scaled_input_test[test_nr_sign > 0]).ravel())
nr_pred_values[test_nr_sign < 0] = -10 ** output_scalers["nrtend_TAU"][-1].inverse_transform(nr_neg_net.predict(scaled_input_test[test_nr_sign < 0]).ravel())
# ## Verification
pred_tendencies = pd.DataFrame(0, index=scaled_out_test.index, columns=output_cols[1:4], dtype=float)
qr_class_preds = dnn_qr_class.predict(scaled_input_test)
pred_tendencies.loc[:, "qrtend_TAU"] = output_scalers["qrtend_TAU"][1].inverse_transform(dnn_qr.predict(scaled_input_test))
pred_tendencies.loc[:, "nctend_TAU"] = output_scalers["nctend_TAU"][1].inverse_transform(dnn_nc.predict(scaled_input_test))
pred_tendencies.loc[:, "nrtend_TAU"] = nr_pred_values
confusion_matrix(np.where(labels_test["nrtend_TAU"] != 0, 1, 0), labels_test["nctend_TAU"])
confusion_matrix(labels_test["nrtend_MG2"], labels_test["qrtend_MG2"])
confusion_matrix(np.where(labels_test["nrtend_TAU"] != 0, 1, 0), labels_test["qrtend_MG2"])
cm = confusion_matrix(labels_test["nrtend_TAU"], labels_test["nrtend_MG2"])
print(cm)
cm = confusion_matrix(labels_test["nctend_TAU"], labels_test["nctend_MG2"])
print(cm)
pred_tendencies.to_csv("/glade/p/cisl/aiml/dgagne/cam_run2_nn_predictions.csv")
# +
fig, axes = plt.subplots(3, 3, figsize=(10, 10))
plt.subplots_adjust(0.03, 0.03, 0.96, 0.95, wspace=0.2)
all_bins = [np.linspace(-16, -4, 50), np.linspace(-10,6, 50), np.linspace(-200, 300, 50)]
original_out_test = np.zeros(transformed_out_test["nrtend_TAU"].size)
original_out_test[labels_test["nrtend_TAU"] > 0] = 10 ** transformed_out_test.loc[labels_test["nrtend_TAU"] > 0,
"nrtend_TAU"]
original_out_test[labels_test["nrtend_TAU"] < 0] = -10 ** transformed_out_test.loc[labels_test["nrtend_TAU"] < 0,
"nrtend_TAU"]
axes[0, 0].hist(pred_tendencies.loc[labels_test["qrtend_TAU"] == 1, "qrtend_TAU"],
bins=all_bins[0], color='skyblue')
axes[0, 0].hist(transformed_out_test["qrtend_TAU"],
bins=all_bins[0], histtype="step", color="navy", lw=3)
axes[0, 0].set_ylabel("TAU Neural Network", fontsize=18)
axes[0, 0].set_yscale("log")
axes[0, 1].hist(pred_tendencies.loc[:, "nctend_TAU"],
bins=all_bins[1], color='skyblue')
axes[0, 1].hist(transformed_out_test["nctend_TAU"],
bins=all_bins[1], histtype="step", color="navy", lw=3)
axes[0, 1].set_yscale("log")
#axes[0, 1].set_xlabel("Emulated NC", fontsize=14)
axes[0, 2].hist(pred_tendencies.loc[:, "nrtend_TAU"],
bins=all_bins[2], color='skyblue')
axes[0, 2].hist(original_out_test,
bins=all_bins[2], histtype="step", color="navy", lw=3)
axes[0, 2].set_yscale("log")
#axes[0, 2].set_xlabel("Emulated NR", fontsize=14)
axes[1, 0].hist(transformed_out_test["qrtend_TAU"],
bins=all_bins[0], color='navy')
axes[1, 0].set_ylabel("TAU Bin", fontsize=18)
axes[1, 0].set_yscale("log")
#axes[1, 0].set_xlabel("Bin QR", fontsize=14)
axes[1, 1].hist(transformed_out_test["nctend_TAU"],
bins=all_bins[1], color='navy')
axes[1, 1].set_yscale("log")
#axes[1, 1].set_xlabel("Bin NC", fontsize=14)
axes[1, 2].hist(original_out_test,
bins=all_bins[2], color='navy')
axes[1, 2].set_yscale("log")
#axes[1, 2].set_xlabel("Bin NR", fontsize=14)
axes[2, 0].hist(transformed_out_test["qrtend_MG2"],
bins=all_bins[0], color='purple')
axes[2, 0].hist(transformed_out_test["qrtend_TAU"],
bins=all_bins[0], histtype="step", color="navy", lw=3)
axes[2, 0].set_yscale("log")
axes[2, 0].set_ylabel("MG2 Bulk", fontsize=18)
axes[2, 0].set_xlabel("$\log_{10}(\Delta$Rain Mass)", fontsize=14)
axes[2, 1].hist(transformed_out_test["nctend_MG2"],
bins=all_bins[1], color='purple')
axes[2, 1].hist(transformed_out_test["nctend_TAU"],
bins=all_bins[1], histtype="step", color="navy", lw=3)
axes[2, 1].set_yscale("log")
axes[2, 1].set_xlabel("$\log_{10}(\Delta$Number Cloud Droplets)", fontsize=14)
original_out_test_mg = np.zeros(transformed_out_test["nrtend_MG2"].size)
original_out_test_mg[labels_test["nrtend_MG2"] > 0] = 10 ** transformed_out_test.loc[labels_test["nrtend_MG2"] > 0,
"nrtend_MG2"]
original_out_test_mg[labels_test["nrtend_MG2"] < 0] = -10 ** transformed_out_test.loc[labels_test["nrtend_MG2"] < 0,
"nrtend_MG2"]
axes[2, 2].hist(original_out_test_mg,
bins=all_bins[2], color='purple')
axes[2, 2].hist(original_out_test,
bins=all_bins[2], histtype="step", color="navy", lw=3)
axes[2, 2].set_yscale("log")
axes[2, 2].set_xticks(np.arange(-200, 350, 100))
axes[1, 2].set_xticks(np.arange(-200, 350, 100))
axes[0, 2].set_xticks(np.arange(-200, 350, 100))
axes[2, 2].set_xlabel("$\Delta$ Number of Rain Drops", fontsize=14)
#axes[0, 0].set_ylabel("Frequency", fontsize=14)
#axes[1, 0].set_ylabel("Frequency", fontsize=14)
#axes[2, 0].set_ylabel("Frequency", fontsize=14)
fig.suptitle("Bin Microphysics Tendency Emulation Distributions", y=0.98, fontsize=22, fontweight="bold")
plt.savefig("nn_bin_hist.png", dpi=200, bbox_inches="tight")
plt.savefig("nn_bin_hist.pdf", dpi=200, bbox_inches="tight")
# -
plt.hist(original_out_test_mg[original_out_test_mg > 0], bins=100)
#plt.hist(original_out_test, bins=100)
plt.gca().set_yscale("log")
# +
plt.hist(np.log10(original_out_test_mg[original_out_test_mg > 0]), bins=np.arange(-12, 6, 0.1), histtype="step", lw=3, label="MG NR")
plt.hist(np.log10(original_out_test[original_out_test > 0]), bins=np.arange(-12, 6, 0.1), histtype="step", lw=3, label="TAU NR")
plt.hist(np.log10(pred_tendencies["nrtend_TAU"].values[pred_tendencies["nrtend_TAU"] > 0]), bins=np.arange(-12, 6, 0.1), histtype="step", lw=3, label="NN NR")
plt.gca().set_yscale("log")
plt.xlabel("Positive NR Tendency")
plt.legend(loc=2)
plt.savefig("mg_tau_nr_pos_hist.png", dpi=200, bbox_inches="tight")
# +
plt.hist(np.log10(-original_out_test_mg[original_out_test_mg < 0]), bins=np.arange(-12, 6, 0.1), histtype="step", lw=3, label="MG NR")
plt.hist(np.log10(-original_out_test[original_out_test < 0]), bins=np.arange(-12, 6, 0.1), histtype="step", lw=3, label="TAU NR")
plt.hist(np.log10(-pred_tendencies["nrtend_TAU"].values[pred_tendencies["nrtend_TAU"] < 0]), bins=np.arange(-12, 6, 0.1), histtype="step", lw=3, label="NN NR")
plt.gca().set_yscale("log")
plt.xlabel("Negative NR Tendency")
plt.legend(loc=2)
plt.savefig("mg_tau_nr_neg_hist.png", dpi=200, bbox_inches="tight")
# -
plt.hist(original_out_test_mg[original_out_test_mg < 0], bins=100)
plt.hist(original_out_test[original_out_test < 0], bins=100)
plt.gca().set_yscale("log")
transformed_out_test.loc[transformed_out_test["qrtend_TAU"] < 0, "qrtend_TAU"]
rmses = np.zeros(5)
rmses[0] = np.sqrt(mean_squared_error(transformed_out_test.loc[transformed_out_test["qrtend_TAU"] < 0, "qrtend_TAU"],
pred_tendencies.loc[transformed_out_test["qrtend_TAU"] < 0, "qrtend_TAU"]))
rmses[1] = np.sqrt(mean_squared_error(transformed_out_test["nctend_TAU"],
pred_tendencies.loc[:, "nctend_TAU"]))
original_out_test = np.zeros(transformed_out_test["nrtend_TAU"].size)
original_out_test[labels_test["nrtend_TAU"] > 0] = 10 ** transformed_out_test.loc[labels_test["nrtend_TAU"] > 0,
"nrtend_TAU"]
original_out_test[labels_test["nrtend_TAU"] < 0] = -10 ** transformed_out_test.loc[labels_test["nrtend_TAU"] < 0,
"nrtend_TAU"]
rmses[2] = np.sqrt(mean_squared_error(original_out_test,
pred_tendencies.loc[:, "nrtend_TAU"]))
rmses[3] = np.sqrt(mean_squared_error(np.log10(-pred_tendencies["nrtend_TAU"][pred_tendencies["nrtend_TAU"] < 0]),
np.log10(np.maximum(-original_out_test[pred_tendencies["nrtend_TAU"] < 0], 1e-20))))
rmses[4] = np.sqrt(mean_squared_error(np.log10(pred_tendencies["nrtend_TAU"][pred_tendencies["nrtend_TAU"] > 0]),
np.log10(np.maximum(original_out_test[pred_tendencies["nrtend_TAU"] > 0], 1e-20))))
r2s = np.zeros(5)
r2s[0] = np.corrcoef(transformed_out_test.loc[transformed_out_test["qrtend_TAU"] < 0, "qrtend_TAU"],
pred_tendencies.loc[transformed_out_test["qrtend_TAU"] < 0, "qrtend_TAU"])[0, 1] ** 2
r2s[1] = np.corrcoef(transformed_out_test["nctend_TAU"],
pred_tendencies.loc[:, "nctend_TAU"])[0, 1] ** 2
r2s[2] = np.corrcoef(original_out_test,
pred_tendencies.loc[:, "nrtend_TAU"])[0, 1] ** 2
r2s[3] = np.corrcoef(np.log10(-pred_tendencies["nrtend_TAU"][pred_tendencies["nrtend_TAU"] < 0]),
np.log10(np.maximum(-original_out_test[pred_tendencies["nrtend_TAU"] < 0], 1e-20)))[0, 1] ** 2
r2s[4] = np.corrcoef(np.log10(pred_tendencies["nrtend_TAU"][pred_tendencies["nrtend_TAU"] > 0]),
np.log10(np.maximum(original_out_test[pred_tendencies["nrtend_TAU"] > 0], 1e-20)))[0, 1] ** 2
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
plt.subplots_adjust(wspace=0.3)
axes[0].hist2d(pred_tendencies["qrtend_TAU"],
transformed_out_test["qrtend_TAU"],
cmin=1, bins=all_bins[0], norm=LogNorm())
axes[0].set_xlabel("Emulated dQR/dt", fontsize=18)
axes[0].set_ylabel("Bin dQR/dt", fontsize=18)
axes[1].set_xticks(np.arange(-16, 7))
axes[1].set_yticks(np.arange(-16, 7))
axes[0].plot(all_bins[0], all_bins[0], 'k--')
axes[0].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses[0], r2s[0]), fontsize=16)
axes[1].hist2d(pred_tendencies["nctend_TAU"],
transformed_out_test["nctend_TAU"],
cmin=1, bins=all_bins[1], norm=LogNorm())
axes[1].set_xlabel("Emulated dNC/dt", fontsize=18)
axes[1].set_ylabel("Bin dNC/dt", fontsize=18)
axes[1].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses[1], r2s[1]), fontsize=16)
axes[1].set_xticks(np.arange(-10, 7))
axes[1].set_yticks(np.arange(-10, 7))
axes[1].plot(all_bins[1], all_bins[1], 'k--')
axes[2].hist2d(pred_tendencies["nrtend_TAU"],
original_out_test,
cmin=1, bins=all_bins[2], norm=LogNorm())
axes[2].set_xlabel("Emulated dNR/dt", fontsize=18)
axes[2].set_ylabel("Bin dNR/dt", fontsize=18)
axes[2].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses[2], r2s[2]), fontsize=16)
#axes[2].set_xticks(np.arange(-400, 500, 100))
#axes[2].set_yticks(np.arange(-400, 500, 100))
axes[2].plot(all_bins[2], all_bins[2], 'k--')
plt.savefig("nn_bin_hist2d.png", dpi=300, bbox_inches="tight")
# +
fig, axes = plt.subplots(1, 4, figsize=(17, 4))
plt.subplots_adjust(wspace=0.3)
axes[0].hist2d(pred_tendencies["qrtend_TAU"],
transformed_out_test["qrtend_TAU"],
cmin=1, bins=all_bins[0], norm=LogNorm())
axes[0].set_xlabel("Emulated dQR/dt", fontsize=18)
axes[0].set_ylabel("Bin dQR/dt", fontsize=18)
axes[1].set_xticks(np.arange(-16, 7))
axes[1].set_yticks(np.arange(-16, 7))
axes[0].plot(all_bins[0], all_bins[0], 'k--')
axes[0].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses[0], r2s[0]), fontsize=16)
axes[1].hist2d(pred_tendencies["nctend_TAU"],
transformed_out_test["nctend_TAU"],
cmin=1, bins=all_bins[1], norm=LogNorm())
axes[1].set_xlabel("Emulated dNC/dt", fontsize=18)
axes[1].set_ylabel("Bin dNC/dt", fontsize=18)
axes[1].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses[1], r2s[1]), fontsize=16)
axes[1].set_xticks(np.arange(-10, 7))
axes[1].set_yticks(np.arange(-10, 7))
axes[1].plot(all_bins[1], all_bins[1], 'k--')
axes[2].hist2d(np.log10(-pred_tendencies["nrtend_TAU"][pred_tendencies["nrtend_TAU"] < 0]),
np.log10(np.maximum(-original_out_test[pred_tendencies["nrtend_TAU"] < 0], 1e-20)),
cmin=1, bins=np.linspace(-10, 5, 50), norm=LogNorm())
axes[2].set_xlabel("Emulated dNR/dt", fontsize=18)
axes[2].set_ylabel("Bin dNR/dt", fontsize=18)
axes[2].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses[3], r2s[3]), fontsize=16)
axes[3].hist2d(np.log10(pred_tendencies["nrtend_TAU"][pred_tendencies["nrtend_TAU"] > 0]),
np.log10(np.maximum(original_out_test[pred_tendencies["nrtend_TAU"] > 0], 1e-20)),
cmin=1, bins=np.linspace(-10, 5, 50), norm=LogNorm())
axes[3].set_xlabel("Emulated dNR/dt", fontsize=18)
axes[3].set_ylabel("Bin dNR/dt", fontsize=18)
axes[3].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses[4], r2s[4]), fontsize=16)
#axes[2].set_xticks(np.arange(-400, 500, 100))
#axes[2].set_yticks(np.arange(-400, 500, 100))
axes[2].plot(all_bins[2], all_bins[2], 'k--')
axes[3].plot(all_bins[2], all_bins[2], 'k--')
plt.savefig("nn_bin_hist2d.png", dpi=300, bbox_inches="tight")
# -
def inverse_nr(nr_labels, nr_vals):
inverse_vals = np.zeros(nr_vals.size)
inverse_vals[nr_labels > 0] = 10 ** nr_vals[nr_labels > 0]
inverse_vals[nr_labels < 0] = -10 ** nr_vals[nr_labels < 0]
return inverse_vals
rmses_bulk = np.zeros(3)
rmses_bulk[0] = np.sqrt(mean_squared_error(transformed_out_test["qrtend_TAU"],
transformed_out_test["qrtend_MG2"]))
rmses_bulk[1] = np.sqrt(mean_squared_error(transformed_out_test["nctend_TAU"],
transformed_out_test["nctend_MG2"]))
rmses_bulk[2] = np.sqrt(mean_squared_error(inverse_nr(labels_test["nrtend_TAU"], transformed_out_test["nrtend_TAU"]),
inverse_nr(labels_test["nrtend_MG2"], transformed_out_test["nrtend_MG2"])))
r2s_bulk = np.zeros(3)
r2s_bulk[0] = np.corrcoef(transformed_out_test["qrtend_TAU"],
transformed_out_test["qrtend_MG2"])[0, 1] ** 2
r2s_bulk[1] = np.corrcoef(transformed_out_test["nctend_TAU"],
transformed_out_test["nctend_MG2"])[0, 1] ** 2
r2s_bulk[2] = np.corrcoef(inverse_nr(labels_test["nrtend_TAU"], transformed_out_test["nrtend_TAU"]),
inverse_nr(labels_test["nrtend_MG2"], transformed_out_test["nrtend_MG2"]))[0, 1] ** 2
fig, axes = plt.subplots(1, 3, figsize=(13, 4))
plt.subplots_adjust(wspace=0.3)
axes[0].hist2d(transformed_out_test["qrtend_MG2"],
transformed_out_test["qrtend_TAU"],
cmin=1, bins=all_bins[0], norm=LogNorm())
axes[0].set_xlabel("Bulk dQR/dt", fontsize=18)
axes[0].set_ylabel("Bin dQR/dt", fontsize=18)
axes[1].set_xticks(np.arange(-16, 7))
axes[1].set_yticks(np.arange(-16, 7))
axes[0].plot(np.arange(-16, 7),np.arange(-16, 7), 'k--')
axes[0].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses_bulk[0], r2s_bulk[0]), fontsize=16)
axes[1].hist2d(transformed_out_test["nctend_MG2"],
transformed_out_test["nctend_TAU"],
cmin=1, bins=all_bins[1], norm=LogNorm())
axes[1].set_xlabel("Bulk dNC/dt", fontsize=18)
axes[1].set_ylabel("Bin dNC/dt", fontsize=18)
axes[1].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses_bulk[1], r2s_bulk[1]), fontsize=16)
axes[1].set_xticks(np.arange(-10, 7))
axes[1].set_yticks(np.arange(-10, 7))
axes[1].plot(np.arange(-3, 7),np.arange(-3, 7), 'k--')
axes[2].hist2d(inverse_nr(labels_test["nrtend_MG2"], transformed_out_test["nrtend_MG2"]),
inverse_nr(labels_test["nrtend_TAU"], transformed_out_test["nrtend_TAU"]),
cmin=1, bins=all_bins[2], norm=LogNorm())
axes[2].set_xlabel("Bulk dNR/dt", fontsize=18)
axes[2].set_ylabel("Bin dNR/dt", fontsize=18)
axes[2].set_title("RMSE: {0:0.3f}; $R^2$: {1:0.3f}".format(rmses_bulk[2], r2s_bulk[2]), fontsize=16)
#axes[2].set_xticks(np.arange(-400, 500, 100))
#axes[2].set_yticks(np.arange(-400, 500, 100))
#axes[2].plot(np.arange(-400, 500, 100),np.arange(-400, 500, 100), 'k--')
plt.savefig("nn_bulk_hist2d.png", dpi=300, bbox_inches="tight")
inverse_nr(labels_test["nrtend_TAU"], transformed_out_test["nrtend_TAU"]).max()
inverse_nr(labels_test["nrtend_MG2"], transformed_out_test["nrtend_MG2"]).max()
# # Partial Dependence Plots
def partial_dependence_1d(x, model, var_index, var_vals):
"""
Calculate how the mean prediction of an ML model varies if one variable's value is fixed across all input
examples.
Args:
x: array of input variables
model: scikit-learn style model object
var_index: column index of the variable being investigated
var_vals: values of the input variable that are fixed.
Returns:
Array of partial dependence values.
"""
partial_dependence = np.zeros(var_vals.shape)
x_copy = np.copy(x)
for v, var_val in enumerate(var_vals):
x_copy[:, var_index] = var_val
partial_dependence[v] = model.predict(x_copy).mean()
return partial_dependence
pd_bins = 20
pd_vals = np.zeros((len(input_cols), pd_bins))
for v, var in enumerate(input_cols):
print(var, scaled_input_train[var].min(), scaled_input_train[var].max())
pd_vals[v] = np.linspace( scaled_input_train[var].min(), scaled_input_train[var].max(), pd_bins)
pd_vals_scaled = input_scaler.transform(log10_transform(pd_vals).T).T
pd_output_qr = np.zeros((len(input_cols), pd_vals.shape[1]))
rand_sub = np.random.permutation(np.arange(np.sum(labels_train["qrtend_TAU"] == 1)))[:50000]
for v, var in enumerate(input_cols):
print(v, var)
pd_output_qr[v] = partial_dependence_1d(scaled_input_train.loc[labels_train["qrtend_TAU"] == 1].iloc[rand_sub], dnn_qr, v,
pd_vals_scaled[v])
pd_output_nc = np.zeros((len(input_cols), pd_vals.shape[1]))
for v, var in enumerate(input_cols):
print(v, var)
pd_output_nc[v] = partial_dependence_1d(scaled_input_train, dnn_nc, v, pd_vals_scaled[v])
pd_output_nr_pos = np.zeros((len(input_cols), pd_vals.shape[1]))
for v, var in enumerate(input_cols):
print(v, var)
pd_output_nr_pos[v] = partial_dependence_1d(scaled_input_train.loc[labels_train["nrtend_TAU"] == 1], nr_pos_net, v, pd_vals_scaled[v])
pd_output_nr_neg = np.zeros((len(input_cols), pd_vals.shape[1]))
for v, var in enumerate(input_cols):
print(v, var)
pd_output_nr_neg[v] = partial_dependence_1d(scaled_input_train.loc[labels_train["nrtend_TAU"] == -1], nr_neg_net, v, pd_vals_scaled[v])
pd_grid_tau.shape
#pd_unscaled = input_scaler.inverse_transform(pd_vals.T)
pd_unscaled_qr = output_scalers["qrtend_TAU"][1].inverse_transform(pd_output_qr)
fig, axes = plt.subplots(3, 2, figsize=(9, 6), sharey=False)
plt.subplots_adjust(hspace=0.5, wspace=0.3)
for a, ax in enumerate(axes.ravel()):
ax.plot(log10_transform(pd_vals[a]), pd_unscaled_qr[a], color='b', label="Emulator")
#ax.plot(log10_transform(pd_vals[a]), pd_grid_tau[a, :, 2], 'k-', label="TAU")
ax.set_xlabel(input_cols[a].replace("_TAU_in", "").replace("_", " "), fontsize=12)
#ax.set_yticks(np.round(np.linspace(np.round(pd_unscaled_qr[a].min(), 1),
# np.round(pd_unscaled_qr[a].max(), 2), 4), 2))
if a == 0:
ax.legend()
fig.suptitle("Partial Dependence for QR Tendencies", fontsize=16, y=0.94)
plt.savefig("pdp_qr_20190420.png", dpi=200, bbox_inches="tight")
# +
pd_unscaled = input_scaler.inverse_transform(pd_vals.T)
pd_unscaled_nc = output_scalers["nctend_TAU"][1].inverse_transform(pd_output_nc)
fig, axes = plt.subplots(3, 2, figsize=(10, 6), sharey=False)
plt.subplots_adjust(hspace=0.5, wspace=0.3)
for a, ax in enumerate(axes.ravel()):
ax.plot(log10_transform(pd_vals[a]), pd_unscaled_nc[a], 'b-', label="Emulator")
ax.plot(log10_transform(pd_vals[a]), pd_grid_tau[a, :, 1], 'k-', label="TAU")
ax.set_xlabel(input_cols[a].replace("_TAU_in", "").replace("_", " "), fontsize=12)
#ax.set_yticks(np.round(np.linspace(np.round(pd_unscaled_nc[a].min(), 1),
# np.round(pd_unscaled_nc[a].max(), 2), 4), 2))
if a == 0:
ax.legend()
fig.suptitle("Partial Dependence for NC Tendencies", fontsize=16, y=0.94)
plt.savefig("pdp_nc.png", dpi=200, bbox_inches="tight")
# +
pd_unscaled = input_scaler.inverse_transform(pd_vals.T)
pd_unscaled_nr_pos = output_scalers["nrtend_TAU"][1].inverse_transform(pd_output_nr_pos)
pd_unscaled_nr_neg = output_scalers["nrtend_TAU"][-1].inverse_transform(pd_output_nr_neg)
fig, axes = plt.subplots(3, 2, figsize=(10, 6), sharey=False)
plt.subplots_adjust(hspace=0.5, wspace=0.3)
for a, ax in enumerate(axes.ravel()):
ax.plot(log10_transform(pd_vals[a]), pd_unscaled_nr_pos[a], 'b-', label="Emulator")
ax.plot(log10_transform(pd_vals[a]), pd_grid_tau[a, :, 3], 'k-', label="TAU")
ax.set_xlabel(input_cols[a].replace("_TAU_in", "").replace("_", " "), fontsize=12)
ax.set_yticks(np.round(np.linspace(np.round(pd_unscaled_nr_pos[a].min(), 1),
np.round(pd_unscaled_nr_pos[a].max(), 2), 4), 2))
if a == 0:
ax.legend()
fig.suptitle("Partial Dependence for NR+ Tendencies", fontsize=16, y=0.94)
plt.savefig("pdp_nr_pos.png", dpi=200, bbox_inches="tight")
# +
pd_unscaled = input_scaler.inverse_transform(pd_vals.T)
pd_unscaled_nr_pos = output_scalers["nrtend_TAU"][1].inverse_transform(pd_output_nr_pos)
pd_unscaled_nr_neg = output_scalers["nrtend_TAU"][-1].inverse_transform(pd_output_nr_neg)
fig, axes = plt.subplots(3, 2, figsize=(10, 6), sharey=False)
plt.subplots_adjust(hspace=0.5, wspace=0.3)
for a, ax in enumerate(axes.ravel()):
ax.plot(log10_transform(pd_vals[a]), pd_unscaled_nr_neg[a], 'b-', label="Emulator")
ax.plot(log10_transform(pd_vals[a]), pd_grid_tau[a, :, 4], 'k-', label="TAU")
ax.set_xlabel(input_cols[a].replace("_TAU_in", "").replace("_", " "), fontsize=12)
#ax.set_yticks(np.round(np.linspace(np.round(pd_unscaled_nr_neg[a].min(), 1),
# np.round(pd_unscaled_nr_neg[a].max(), 2), 4), 2))
if a == 0:
ax.legend()
fig.suptitle("Partial Dependence for NR- Tendencies", fontsize=16, y=0.94)
plt.savefig("pdp_nr_neg.png", dpi=200, bbox_inches="tight")
# -
all_pd_out = [pd_output_qr, pd_output_nc, pd_output_nr_pos, pd_output_nr_neg]
pd_names = ["QR", "NC", "NR+", "NR-"]
colors = ['red', 'green', 'blue', 'purple']
fig, axes = plt.subplots(3,2, figsize=(10, 6), sharey=False)
plt.subplots_adjust(hspace=0.5, wspace=0.3)
for a, ax in enumerate(axes.ravel()):
for o in range(len(all_pd_out)):
ax.plot(log10_transform(pd_vals[ a]),
(all_pd_out[o][a] - all_pd_out[o][a].min()) / (all_pd_out[o][a].max() - all_pd_out[o][a].min()),
color=colors[o])
ax.set_xlabel(input_cols[a].replace("_TAU_in", "").replace("_", " "), fontsize=12)
plt.figlegend(pd_names, loc='lower center', ncol=4)
plt.suptitle("Partial Dependence for Bin Neural Network Emulators", y=0.95, fontsize=16)
plt.savefig("pdp_all.png", dpi=200, bbox_inches="tight")
pd_grid_nc = partial_dependence_2d(scaled_input_train, dnn_nc, 5, pd_vals[5], 6, pd_vals[6])
plt.contourf(pd_unscaled[:, 6], pd_unscaled[:, 5], pd_grid_nc, np.arange(-3, 3.5, .5), cmap="RdBu_r")
plt.xlabel(input_cols[6])
plt.ylabel(input_cols[5])
plt.colorbar()
| notebooks/bin_mp_nn_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Capstone Final Report
# ## Amazon Video Game Reviews NLP
# <NAME>
# #### Problem - Text Data!
#
# Text data is collected for many purposes. This could include chats where a customer/client has a concern and reviews of products/services are some of the most common. Other times, text that has been published needs to be analyed to create summaries or extract key topics. Natural Langauge Processing (NLP) is a processes that provides methods for cleaning text data, breaking it down into manageable chunks, and extracting meaningful insights.
#
# This analysis focuses on reviews collected by Amazon for video game purchases between 1996-2004. The data include a star rating and text written by the customer describing how they feel about the game. It is very useful to be able to predict a customer or client's positive or negative opinion from text. This information can be used to help make decisions for certain products, create recommendations, or provide areas for improvement on products.
#
# The primary goal is to analyze a subset of the text reviews and use the star rating (converted to positive or negative) to create a model capable of predicting whether a video game review is positive or negative. The first approach for this is to use a pre-trained model, that has already analyzed text data for positive or negative sentiment. If this model is successful, then it can be deployed without having to create a custom model. This would save time and resources from a business perspective.
#
# If the pre-trained model isn't accurate or fails to predict the correct label, a model will be trained on the data of interest. There are several reasons why a pre-trained model could be unsuccessful. One ver yimportant reason is that no model is trained on every possible combination of features. In other words, a pre-trained model may use data for training that is very dissimilar to the data for which predictions are desired. Training of a model in this case would be supervised learning, where pre-labeled data are used to train a model. The effectiveness of the model will be assessed on hold-out test data that the model was not exposed to during training.
# #### The Data
# - The data for this analysis were obtained from the following source:
# > Justifying recommendations using distantly-labeled reviews and fined-grained aspects
# <NAME>, <NAME>, <NAME>
# Empirical Methods in Natural Language Processing (EMNLP), 2019
# [https://nijianmo.github.io/amazon/index.html#files](https://nijianmo.github.io/amazon/index.html#files)
# - The specific file used was the Video Games data with at least 5 reviews per game.
# #### Exploratory Data Analysis
# Several steps were taken to explore the data:
# 1. Parse the JSON file using pandas built-in JSON reader
# 2. Use basic pandas functions to explore the data structure
# 3. Isolate the text data (features) and the ratings (target) and remove any missing values from the data
# 4. Explore the text data for interesting features
# - Word frequencies (most common words)
# - Anomalous words
# - Number of reviews in which a word appears
# - Generate summary plots and a word cloud to visualize the data
# More details on the exploratory analysis can be found in `1_Video_Games_Exploratory_NLP.ipynb` file. The summary plots and descriptions are displayed here.
#
# The comments in each cell explain the purpose of the code.
# import packages
import numpy as np
import pandas as pd
# read in data and clean the dataframe for missing values
# extract only the relevant columns
vg = pd.read_json('../Amazon_Data/Video_Games_5.json.gz', lines=True, compression='gzip')
vg = vg.loc[:,['overall', 'reviewText']]
vg = vg.dropna(how='any')
vg.loc[:,'overall'] = vg.overall.astype('int16')
# import stop words from spacy and add some additional words identified during data exploration
from spacy.lang.en.stop_words import STOP_WORDS
stops = list(STOP_WORDS)
issue_words = ['\ufeff1', '\x16', '...', '\x10once', 's', '1', '2', '3', '4', '5']
stops.extend(issue_words)
# +
# import additional nlp tools
import string
import spacy
from spacy.lang.en import English
# generate puntuations string
punctuations = string.punctuation
# +
# define remove chars function to remove special characters
import re
def rmv_spec_chars(sentence):
# completely remove most punctuation
sentence = re.sub("[\\\[\]@_!#$%^&*()<>?/\|}{~:']", '', sentence)
# replace hypens with spaces to split those words
sentence = re.sub('-', ' ', sentence)
return sentence
# -
# create tokenizer to parse text into tokens with special characters removed
parser = English()
def my_tokenizer(sentence):
sentence = rmv_spec_chars(sentence)
mytokens = parser(sentence)
mytokens = [word.lemma_.lower().strip() if word.lemma_ != "-PRON-"
else word.lower_ for word in mytokens]
mytokens = [word for word in mytokens if word not in stops and word not in punctuations]
return mytokens
# create word_list function to generate a list of words in the text
def get_word_list(text, word_list):
for sentence in text:
for word in my_tokenizer(sentence):
word_list.append(word)
# get a list of all words each time they occur in the reviews
# using tokenizer and the get_word_list function
word_list = []
get_word_list(vg.reviewText, word_list)
# #### Generating summary stats for the words and appearances
# +
from collections import Counter, defaultdict
# get a count of every word
token_counts = Counter(word_list)
# -
# find the number of unique words in the reviews
# stop words excluded
print('The total number of unique words is {}'.format(len(token_counts)))
# #### Generate some basic visualizations for the number of word appearances
# +
import matplotlib.pyplot as plt
# create histogram of word appearances
# most words occur infrequently, so threshold for num influences this plot greatly
values = []
for tup in token_counts.most_common():
word, num = tup
if num > 10000:
values.append(num)
_ = plt.hist(values, bins=100)
_ = plt.title('Histogram for Number of Appearances')
_ = plt.xlabel('Number of Times a Word Appears')
_ = plt.ylabel('Number of Words')
# -
# This histogram shows that most words appear infrequently, but there are a few words that appear way more than the others.
# +
import seaborn as sns
# plot the 20 most common words
words = []
values = []
for tup in token_counts.most_common(20):
word, num = tup
words.append(word)
values.append(num)
_ = sns.barplot(words, values, palette='muted')
_ = plt.xticks(rotation=90)
_ = plt.title('20 Most Common Words')
_ = plt.xlabel('Word')
_ = plt.ylabel('Occurrences')
# -
# Unsurprisingly, the word that appears the most is 'game'. The other most common words are also displayed here with their number of total appearances. Also of interest is the number of reviews in which a word appears. If a word only appears once, then it may make sense to exclude it from analysis.
# +
# get a count of the number of reviews where a word appears
def get_num_docs(text_series):
"""Take a text series and and return a default dict."""
# initialize default dict
num_docs = defaultdict(int)
# iterate through and populate the default dict
for text in text_series:
ls = []
for word in my_tokenizer(text):
if word not in ls:
ls.append(word)
for x in ls:
num_docs[x] += 1
# return the default dict
return num_docs
# -
# get the number of docs in which each word appears
num_docs = get_num_docs(vg.reviewText)
# store a list of the number of appearances for each word
apps = []
for key, val in num_docs.items():
apps.append(val)
# plot the cdf for the number of reviews where each word appears
_ = plt.hist(apps, cumulative=True, histtype='step', density=True, bins=np.arange(100))
_ = plt.xlim(0,25)
_ = plt.xlabel('Number of Docs')
_ = plt.ylabel('Cumulative Density')
_ = plt.title('Density of Words Appearing \nin at Least X Reviews')
# So about 60% of the words in the vocabulary appear in only one review! When creating the sentiment analysis model, it could be useful to set a minimum appearance of 2.
# The next visualization for the words will be a wordcloud using the WordCloud class.
# import wordcloud
from wordcloud import WordCloud
# initialize wordcloud object
wc = WordCloud(background_color='white', stopwords=stops, max_words=200,
max_font_size=40, scale=3, random_state=42)
# generate the wordcloud
wc.generate(' '.join(word_list))
# +
# show the wordcloud
fig = plt.figure(1, figsize=(12, 12))
plt.axis('off')
plt.imshow(wc)
plt.show()
# -
# ## Summary of Exploratory Analysis
# The text data had some very interesting features. It was surprising to find many special characters and emojis in the data. For this analysis, these were removed, and the analysis was performed on the clean text. I think this is a very important result, given that this was not much of an issue when I was originally working with book reviews.
#
# This shows the importance of understanding the particular dataset for the problem. While the book reviews came from the same source (Amazon for the years 1996-2014), the content was different enough to require different preprocessing techniques.
#
# Plotting word counts showed that over half of the words only appeared in one review. The most common words were plotted, and unsurprisingly, the most common word was game. If using standard models to predict sentiment of reviews, it would be recommended to include only words that appear in two or more reviews.
#
# The next step was to use the Flair package for sentiment analysis.
# ## Sentiment Analysis of Video Game Reviews
# The primary goal here is to obtain a model capable of predicting positive or negative sentiment by analyzing the text of a video game review. The first approach is to use a pre-trained model from the Flair package. It should be noted that this model was trained on IMDB reviews, so the model must be assessed for usefulness with a different data set.
#
# The pre-trained model did fail, so a model was trained. To see the results of the pre-trained model, see `2_Video_Games_Flair_Model.ipynb`. Only the training of the model is displayed here.
# ### Important
# This recurrent neural network employed with the Flair model uses a substantial amount of resources. It tooks a couple of weeks of trial and error before the model finally completed using a GPU and high RAM instance on Google Colab. Only the output is copied here.
# + colab={"base_uri": "https://localhost:8080/", "height": 80} colab_type="code" id="e21SkqIDWQCY" outputId="492d1ee7-3386-450f-85df-ff9d63bae72d"
# import flair
from flair.models import TextClassifier
from flair.data import Sentence
# + colab={} colab_type="code" id="JZB1HRYWXrnv"
# map sentiment for two-class model
vg.loc[:,'pt_sentiment'] = vg.overall.map({1: 0, 2: 0, 3: 1, 4: 1, 5: 1}).astype('int16')
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="lFqtjP64X87p" outputId="8538ae68-bae7-48a4-a2f6-cdf8a6243617"
# get sentiment counts
class1_counts = vg.pt_sentiment.value_counts()[1]
class0_counts = vg.pt_sentiment.value_counts()[0]
print('Positive counts {}'.format(class1_counts))
print('Negative counts {}'.format(class0_counts))
# + [markdown] colab_type="text" id="BRUqFQgmcTu5"
# Because the samples were imbalanced, the majority class was downsampled to even out the classes and improve performance of model training.
# + colab={} colab_type="code" id="1gkLoV3pOTqR"
# import resample
from sklearn.utils import resample
# down-sample to balance classes
vg_class1 = vg[vg.pt_sentiment == 1]
vg_class0 = vg[vg.pt_sentiment == 0]
# downsample majority class
vg_class1_down = resample(vg_class1, replace=False, n_samples=vg_class0.shape[0], random_state=42)
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="dygdNkWLRKoj" outputId="fb02c2d4-4470-4d12-e62e-95f8f00c1651"
# concat the dfs back together
vg_down = pd.concat([vg_class1_down, vg_class0])
vg_down.pt_sentiment.value_counts()
# + [markdown] colab_type="text" id="MYtY2iuLceJF"
# Previous exploratory analysis revealed the presence of many special characters. These will be removed from the text prior to training.
# + colab={} colab_type="code" id="8FCZaeCAp1g4"
# clean the text
vg_down.loc[:,'clean_text'] = vg_down.reviewText.apply(rmv_spec_chars)
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="m1V4Jy0lqKol" outputId="68100d71-f45e-486c-9c70-a6e2434501bb"
# indicate which text fields are now invalid due to entries that only contain
# white space after removal of special characters
vg_down.loc[:,'invalid'] = vg_down.clean_text.apply(lambda x: bool(re.match('^\s+$', x)))
vg_down.invalid.value_counts()
# + colab={} colab_type="code" id="IG17vF6MlxBw"
# remove these from the dataset
vg_down = vg_down[vg_down.invalid == False]
# + colab={} colab_type="code" id="CFBtUFOWtwYR"
# identify additional invalid entries that have had every character removed
# by the cleaning function (entry was entirely special characters)
vg_down.loc[:,'invalid'] = vg_down.clean_text.apply(lambda x: len(x) == 0)
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="5kXb9pAKuMF1" outputId="6c763de7-01cc-400f-c9b7-08900d69c61f"
# inspect the number of invalid entries that are empty
vg_down.invalid.value_counts()
# + colab={} colab_type="code" id="zaqK7A9JuUvF"
# remove additional invalid text entries
vg_down = vg_down[vg_down.invalid == False]
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="LLiwIhAIlxGw" outputId="b50eebc7-bc04-498f-eecf-d2f40998a442"
# classes are still approximately balanced
vg_down.pt_sentiment.value_counts()
# + colab={} colab_type="code" id="7oHi_MVyYEUG"
# split the data into train/validation
train_text, test_text, train_labels, test_labels = train_test_split(vg_down.clean_text,
vg_down.pt_sentiment,
test_size=0.20,
random_state=42,
stratify=vg_down.pt_sentiment)
# + colab={} colab_type="code" id="RuYATGB-nUrL"
# split the test data into test/valid
valid_text, test_text, valid_labels, test_labels = train_test_split(test_text, test_labels,
test_size=0.50,
random_state=42,
stratify=test_labels)
# + [markdown] colab_type="text" id="T2fKmBvpgb1N"
# ### Manual model training
#
# A model was trained using the video games data to try and increase model effectiveness.
# + colab={} colab_type="code" id="uQPZBUb6dQ9T"
# manual model training
# + colab={} colab_type="code" id="V179PtJZdQ64"
# write trianing data to file
# format for flair is below
# '__label__X Here is the text for the review'
# where 'X' is the label for the review (0 is negative, 1 is positive)
import os.path
idxs = train_text.index.values
max_idx = max(idxs)
if os.path.isfile('train.txt'):
print ("File already created.")
else:
with open('train.txt', 'w') as file:
for idx in idxs:
line = '__label__'
line += str(train_labels[idx]) + ' '
line += str.replace(train_text[idx], '\n', ' ')
line += '\n'
file.write(line)
# + colab={} colab_type="code" id="G4AOGzDnempD"
# write dev.txt
max_idx = max(valid_text.index.values)
if os.path.isfile('dev.txt'):
print ("File already created.")
else:
with open('dev.txt', 'w') as file:
for idx in valid_text.index.values:
line = '__label__'
line += str(valid_labels[idx]) + ' '
line += str.replace(valid_text[idx], '\n', ' ')
line += '\n'
file.write(line)
# + colab={} colab_type="code" id="zzYWvHfPduhj"
# write test.txt
import os.path
max_idx = max(test_text.index.values)
if os.path.isfile('test.txt'):
print ("File already created.")
else:
with open('test.txt', 'w') as file:
for idx in test_text.index.values:
line = '__label__'
line += str(test_labels[idx]) + ' '
line += str.replace(test_text[idx], '\n', ' ')
line += '\n'
file.write(line)
# + colab={} colab_type="code" id="8zs1wk8GkbOb"
# import necessary packages
from flair.data import Corpus
from flair.datasets import ClassificationCorpus
from flair.trainers import ModelTrainer
from pathlib import Path
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="EJoxUVFTQTtP" outputId="79747810-7ac7-490f-9438-5afe3d9609e6"
# check for GPU availability
import torch
torch.cuda.is_available()
# + colab={} colab_type="code" id="Yx_H5-TWQAR0"
# set storage and GPU usage
import flair
device = torch.device('cuda:0')
map_location=lambda storage, loc: storage.cuda()
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="1kFH0vcdkiSZ" outputId="e2c800e9-cd15-4153-fb1c-3c13f240ba7b"
# create corpus containing training, test and dev data
corpus: Corpus = ClassificationCorpus('./',
test_file='test.txt',
dev_file='dev.txt',
train_file='train.txt')
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="JbH84dyYoaf4" outputId="f36f8a28-415a-41fa-e540-4d61139d61fa"
type(corpus)
# + colab={} colab_type="code" id="B58JDz5LodEa"
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="ouF7bUX3oieE" outputId="55738b51-7b26-4910-fc85-70a5f79474af"
# generate label dictionary
# fails if there are text entries that are invalid
# (see removal of invalid entries above)
label_dictionary=corpus.make_label_dictionary()
# + colab={} colab_type="code" id="MAp1sAmSkxB3"
# import embeddings
from flair.embeddings import TokenEmbeddings, WordEmbeddings, StackedEmbeddings
from flair.embeddings import FlairEmbeddings, DocumentRNNEmbeddings
from typing import List
# + colab={"base_uri": "https://localhost:8080/", "height": 343} colab_type="code" id="41HWFz33k85h" outputId="e5c2b8d5-8050-443c-e877-3c66953adc29"
# create word embeddings
word_embeddings = [WordEmbeddings('glove'), FlairEmbeddings('news-forward-fast'),
FlairEmbeddings('news-backward-fast')]
# + colab={} colab_type="code" id="08e0-1TglAoi"
# build the RNN document embeddings
document_embeddings = DocumentRNNEmbeddings(word_embeddings, hidden_size=512,
reproject_words=True, reproject_words_dimension=256,
rnn_type='LSTM')
# + colab={} colab_type="code" id="hrMGol1-lHBz"
# create the classifier
classifier = TextClassifier(document_embeddings,
label_dictionary=label_dictionary,
multi_label=False)
# + colab={} colab_type="code" id="GLKUNdNSlLBQ"
# create a model trainer
trainer = ModelTrainer(classifier, corpus)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="AkaWGTe7lO9w" outputId="d4af82b9-fc03-40c9-b2a5-4be57587ab1c"
# train the model
# takes approx 12.5 hours to train on Google Colab using Pro GPU
trainer.train('./', max_epochs=10, monitor_test=True, embeddings_storage_mode='gpu')
# + [markdown] colab_type="text" id="2WP2ZAnYh4M9"
# ### Results of model training
#
# There are several files created during model training, which will need to be saved/downloaded. The most important is 'best-model.pt' which actually contained the trained model.
#
# The final model f1 scores were around 86% for both classes on the dev and test data, which is much better than using the pretrained model.
# -
# create a plot comparing the two models' performances
df = pd.DataFrame({'Model': ['Pre-Trained', 'Custom', 'Pre-Trained', 'Custom',
'Pre-Trained', 'Custom', 'Pre-Trained', 'Custom',
'Pre-Trained', 'Custom', 'Pre-Trained', 'Custom'],
'Metric': ['Negative Precision', 'Negative Precision',
'Positive Precision', 'Positive Precision',
'Negative Recall', 'Negative Recall',
'Positive Recall', 'Positive Recall',
'Negative F1', 'Negative F1',
'Positive F1', 'Positive F1'],
'Value': [0.28, 0.86, 0.96, 0.86, 0.75, 0.86, 0.76, 0.86, 0.41, 0.86, 0.85, 0.86]})
ax = sns.barplot(x='Metric', y='Value', hue='Model', data=df)
_ = plt.xlabel('Performance Metric')
_ = plt.xticks(rotation=45)
_ = plt.ylabel('Score')
_ = plt.title('Pre-Trained vs. Custom Model Comparison')
_ = ax.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# ## Final Report Summary
#
# This analysis found some interesting features in the data. One of the most interesting is that this data showed it was very different than the Amazon books data that was used in the very early stages of this project. The failure of the pre-trained model to perform well also showed that this data was very different than the IMDB data that model used for training. The lesson here, is that it is a bad assumption that a model trained on text data from one source are automatically going to generalize to other types of text data.
#
# This analysis provides examples of several tools used to work with text data. These include parsing the data into tokens or individual words, getting summary statistics on the data including total word counts, number of word appearances, and the number of documents in which a word appears. Visualizations were used to help communicate this information in a clear way.
#
# The process to create a sentiment classifier for text data using Flair was also displayed, and some challenges in doing so were also documented. This was a challenging project, but the applications for many different businesses make these tools very valuable.
#
# Running text data through sentiment analysis can help to identify pleased customers/clients and those that are angry or disappointed. This can help to prioritize responses or generate actions that can lead to better outcomes for the business. Additional NLP tools like providing text summaries, can also be very useful for classifying text input to help direct customers to the right department/agent or to sort articles or other text data into categories.
#
# Future analysis on this data set could try to identify categories of reviews or use the items, customers, and their ratings to create a recommendation model for suggesting video games to customers using similarities in customer choices or suggestions based on similar items.
| 3_Final_Report_VG_NLP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gagan3012/zeroshot-learning/blob/master/Zero_Shot_learning_experiment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="m9Xl0NnDx0NI"
import pandas as pd
df = pd.read_csv("/content/bquxjob_3bc46ddf_17ce26263bb.csv")
df.head()
# + id="C35EOzq63UoA"
# !pip install newspaper3k==0.2.8
# + id="tYTt3_Fm03u3"
from bs4 import BeautifulSoup
import newspaper
def get_article_metadata_newspaper(df):
"""
Get/extract the metadata of news article, including, article heading and main content, from it's URL.
Arguments:
url: (str) link of news article (starting with http:// or https://)
Returns:
dict:
title: (str) extracted heading of news article
text: (str) extracted main content of news article
"""
df = df.drop_duplicates(subset=['SOURCEURL'],ignore_index = True)
df["text"] = 0
df["title"]= 0
for i in range(len(df)):
url = df['SOURCEURL'][i]
article = newspaper.Article(url)
try:
article.download()
article.parse()
except:
continue
df["text"][i] = article.text
df["title"][i] = article.title
return df
# + id="ek79OLEw3Lbq"
df_new = get_article_metadata_newspaper(df)
# + id="7kUvF0d06XmV"
df_new.head()
# + colab={"base_uri": "https://localhost:8080/"} id="4ESG4tOpDzjg" outputId="98f43163-3b65-4792-e229-be5078af0215"
# !pip install transformers
# + colab={"base_uri": "https://localhost:8080/"} id="4U_x523DIb8n" outputId="3e56fe5b-c384-4731-e535-1eb3be858537"
len(df_new["text"][0])
# + id="-2PLWQqQkNVA"
import re
def stringupdate(i):
strList = df_new["Themes"][i].split(";")
strList = list(map(lambda x: str(x).replace('WB_', ''), strList))
strList = list(map(lambda x: re.sub('^\d+', '', x), strList))
strList = list(map(lambda x: str(x).replace('_', '', 1), strList))
strList = list(filter(None, strList))
return strList
# + [markdown] id="Tai__nZ0tHlN"
# ##### ZSL
# This is the checkpoint for bart-large after being trained on the MultiNLI (MNLI) dataset.
#
# Additional information about this model:
#
# The bart-large model page
# BART: Denoising Sequence-to-Sequence Pre-training for Natural Language Generation, Translation, and Comprehension
# BART fairseq implementation
# NLI-based Zero Shot Text Classification
# Yin et al. proposed a method for using pre-trained NLI models as a ready-made zero-shot sequence classifiers. The method works by posing the sequence to be classified as the NLI premise and to construct a hypothesis from each candidate label. For example, if we want to evaluate whether a sequence belongs to the class "politics", we could construct a hypothesis of This text is about politics.. The probabilities for entailment and contradiction are then converted to label probabilities.
#
# This method is surprisingly effective in many cases, particularly when used with larger pre-trained models like BART and Roberta. See this blog post for a more expansive introduction to this and other zero shot methods, and see the code snippets below for examples of using this model for zero-shot classification both with Hugging Face's built-in pipeline and with native Transformers/PyTorch code.
# + id="sZJosrMtB88E"
from transformers import pipeline
classifier = pipeline("zero-shot-classification",
model="facebook/bart-large-mnli")
# + id="Xg2w8OnEDy0m" colab={"base_uri": "https://localhost:8080/"} outputId="803e3cdf-f50a-4db6-98ca-dc66cbb8a6ea"
sequence_to_classify =df_new["title"][3]
candidate_labels = stringupdate(3)
classifier(sequence_to_classify, candidate_labels, multi_label=True)
# + id="mCGijDkYSoWO"
| Zero_Shot_learning_experiment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="zX6H255Agj68"
# <a href="https://colab.research.google.com/github/google-research/text-to-text-transfer-transfrormer/blob/main/notebooks/t5-deploy.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="Yo_HOomXe1f2"
# ##### Copyright 2020 The T5 Authors
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + colab={} colab_type="code" id="Rz9fAJ8PexKB"
# Copyright 2020 The T5 Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
# + [markdown] colab_type="text" id="geoZEiaLdGfR"
# # T5 SavedModel Export and Inference
#
# This notebook guides you through the process of exporting a [T5](https://github.com/google-research/text-to-text-transformer) `SavedModel` for inference. It uses the fine-tuned checkpoints in the [T5 Closed Book QA](https://github.com/google-research/google-research/tree/main/t5_closed_book_qa) repository for the [Natural Questions](https://ai.google.com/research/NaturalQuestions/) task as an example, but the same process will work for any model trained with the `t5` library.
#
# For more general usage of the `t5` library, please see the main [github repo](https://github.com/google-research/text-to-text-transfer-transformer) and fine-tuning [colab notebook](https://goo.gle/t5-colab).
#
#
# + [markdown] colab_type="text" id="WtS5hODBKtR_"
# ## Install T5 Library
# + colab={} colab_type="code" id="UHCx-R4M-D0a"
# !pip install -q t5
# + [markdown] colab_type="text" id="IFuyCiHpLCh7"
# ## Export `SavedModel` to local storage
#
# NOTE: This will take a while for XL and XXL.
# + cellView="both" colab={} colab_type="code" id="-Y7QSepo9a8H"
MODEL = "small_ssm_nq" #@param["small_ssm_nq", "t5.1.1.xl_ssm_nq", "t5.1.1.xxl_ssm_nq"]
import os
saved_model_dir = f"/content/{MODEL}"
# !t5_mesh_transformer \
# --model_dir="gs://t5-data/pretrained_models/cbqa/{MODEL}" \
# --use_model_api \
# --mode="export_predict" \
# --export_dir="{saved_model_dir}"
saved_model_path = os.path.join(saved_model_dir, max(os.listdir(saved_model_dir)))
# + [markdown] colab_type="text" id="JUH5BkcYK3At"
# ## Load `SavedModel` and create helper functions for inference
#
# NOTE: This will take a while for XL and XXL.
# + colab={} colab_type="code" id="xiBSnGuu-em0"
import tensorflow as tf
import tensorflow_text # Required to run exported model.
model = tf.saved_model.load(saved_model_path, ["serve"])
def predict_fn(x):
return model.signatures['serving_default'](tf.constant(x))['outputs'].numpy()
def answer(question):
return predict_fn([question])[0].decode('utf-8')
# + [markdown] colab_type="text" id="HSlRnhz7VpTu"
# ## Ask some questions
#
# We must prefix each question with the `nq question:` prompt since T5 is a multitask model.
# + colab={} colab_type="code" id="CE1bO4hw--Zh"
for question in ["nq question: where is google's headquarters",
"nq question: what is the most populous country in the world",
"nq question: name a member of the beatles",
"nq question: how many teeth do humans have"]:
print(answer(question))
# + [markdown] colab_type="text" id="BlTOrC7iaCZD"
# ## Package in Docker image
# + [markdown] colab_type="text" id="ndK6zIryaKTX"
# ```bash
# MODEL_NAME=model-name
# SAVED_MODEL_PATH=/path/to/export/dir
#
# # Download the TensorFlow Serving Docker image and repo:
# docker pull tensorflow/serving:nightly
#
# # First, run a serving image as a daemon:
# docker run -d --name serving_base tensorflow/serving:nightly
#
# # Next, copy your `SavedModel` to the container's model folder:
# docker cp $SAVED_MODEL_PATH serving_base:/models/$MODEL_NAME
#
# # Now, commit the container that's serving your model:
# docker commit --change "ENV MODEL_NAME ${MODEL_NAME}" serving_base $MODEL_NAME
#
# # Finally, save the image to a tar file:
# docker save $MODEL_NAME -o $MODEL_NAME.tar
#
# # You can now stop `serving_base`:
# docker kill serving_base
# ```
# + [markdown] colab_type="text" id="WYQiZke3nXD_"
# ```bash
# docker run -t --rm -p 8501:8501 --name $MODEL_NAME-server $MODEL_NAME &
#
# curl -d '{"inputs": ["nq question: what is the most populous country?"]}' \
# -X POST http://localhost:8501/v1/models/$MODEL_NAME:predict
#
# docker stop $MODEL_NAME-server
# ```
| notebooks/t5-deploy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="Tce3stUlHN0L"
# ##### Copyright 2018 The TensorFlow Authors.
#
#
# + colab={} colab_type="code" id="tuOe1ymfHZPu"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="MfBg1C5NB3X0"
# # Title
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/not_a_real_link"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/tools/templates/notebook.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/tools/templates/notebook.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="r6P32iYYV27b"
# {**Fix these links**}
# + [markdown] colab_type="text" id="xHxb-dlhMIzW"
# ## Overview
#
# {Include a paragraph or two explaining what this example demonstrates, who should be interested in it, and what you need to know before you get started.}
#
#
# + [markdown] colab_type="text" id="MUXex9ctTuDB"
# ## Setup
# + colab={} colab_type="code" id="IqR2PQG4ZaZ0"
from __future__ import absolute_import, division, print_function, unicode_literals
import tensorflow.compat.v2 as tf
tf.enable_v2_behavior()
import numpy as np
# + [markdown] colab_type="text" id="1Eh-iCRVBm0p"
# {Put all your imports and installs up into a setup section.}
# + [markdown] colab_type="text" id="UhNtHfuxCGVy"
# ## Notes
# + [markdown] colab_type="text" id="kKhmFeraTdEI"
# For general instructions on how to write docs for Tensorflow see [Writing TensorFlow Documentation](https://www.tensorflow.org/community/documentation).
#
# The tips below are specific to notebooks for tensorflow.
# + [markdown] colab_type="text" id="2V22fKegUtF9"
# ### General
#
# * Include the collapsed license at the top (this uses Colab's "Form" mode to hide the cells).
# * Only include a single `H1` title.
# * Include the button-bar immediately under the `H1`.
# * Include an overview section before any code.
# * Put all your installs and imports in a setup section.
# * Always include the three `__future__` imports.
# * Save the notebook with the Table of Contents open.
# * Write python3 compatible code.
# * Keep cells small (~max 20 lines).
#
# + [markdown] colab_type="text" id="YrsKXcPRUvK9"
# ### Working in GitHub
#
# * Be consistent about how you save your notebooks, otherwise the JSON-diffs will be a mess.
#
# * This notebook has the "Omit code cell output when saving this notebook" option set. GitHub refuses to diff notebooks with large diffs (inline images).
#
# * [reviewnb.com](http://reviewnb.com) may help. You can access it using this bookmarklet:
#
# ```
# javascript:(function(){ window.open(window.location.toString().replace(/github\.com/, 'app.reviewnb.com').replace(/files$/,"")); })()
# ```
#
# * To open a GitHub notebook in Colab use the [Open in Colab](https://chrome.google.com/webstore/detail/open-in-colab/iogfkhleblhcpcekbiedikdehleodpjo) extension (or make a bookmarklet).
#
# * The easiest way to edit a notebook in GitHub is to open it with Colab from the branch you want to edit. Then use File --> Save a copy in GitHub, which will save it back to the branch you opened it from.
#
# * For PRs it's helpful to post a direct Colab link to the PR head: https://colab.research.google.com/github/{user}/{repo}/blob/{branch}/{path}.ipynb
#
# + [markdown] colab_type="text" id="QKp40qS-DGEZ"
# ### Code Style
#
#
# * Notebooks are for people. Write code optimized for clarity.
#
# * Demonstrate small parts before combining them into something more complex. Like below:
# + colab={} colab_type="code" id="KtylpxOmceaC"
#Build the model
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, activation='relu', input_shape=(None, 5)),
tf.keras.layers.Dense(3)
])
# + colab={} colab_type="code" id="mMOeXVmbdilM"
# Run the model on a single batch of data, and inspect the output.
result = model(tf.constant(np.random.randn(10,5), dtype = tf.float32)).numpy()
print("min:", result.min())
print("max:", result.max())
print("mean:", result.mean())
print("shape:", result.shape)
# + colab={} colab_type="code" id="U82B_tH2d294"
# Compile the model for training
model.compile(optimizer=tf.keras.optimizers.Adam(),
loss=tf.keras.losses.categorical_crossentropy)
# + [markdown] colab_type="text" id="g3-lzxbCZi-H"
# * Publishing to tensorflow.org doesn't yet support interactive plots like altair, or the [graph embedding trick](https://colab.research.google.com/github/tensorflow/tensorflow/blob/master/tensorflow/examples/tutorials/deepdream/deepdream.ipynb#scrollTo=LrucdvgyQgks).
#
# * Keep examples quick. Use small datasets, or small slices of datasets. Don't train to convergence, train until it's obvious it's making progress.
#
# * For a large example, don't try to fit all the code in the notebook. Host the code somewhere. Use the notebook to [download the example code](https://www.tensorflow.org/tutorials/estimators/linear), and demonstrate how it works.
# + [markdown] colab_type="text" id="TJdqBNBbS78n"
# ### Code content
#
# Use the highest level API that gets the job done (unless the goal is to demonstrate the low level API).
#
# Use `keras.Sequential` > keras functional api > keras model subclassing > ...
#
# Use `model.fit` > `model.train_on_batch` > manual `GradientTapes`.
#
# Use eager-style code.
#
# Use `tensorflow_datasets` and `tf.data` where possible.
#
# Avoid `compat.v1`.
#
#
# + [markdown] colab_type="text" id="78HBT9cQXJko"
# ### Text
#
# * Use an imperative style. "Run a batch of images through the model."
#
# * Use sentence case in titles/headings.
#
# * Use short titles/headings: "Download the data", "Build the Model", "Train the model".
#
#
| tools/templates/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import gc
import os
import warnings
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from tqdm import tqdm
import multiprocessing
from keras import backend as K
warnings.filterwarnings(action='ignore')
K.image_data_format()
# +
# 더 자세한 내용은
# 케라스 코리아 김태영 님의 자료 https://tykimos.github.io/2017/06/10/CNN_Data_Augmentation/
# 를 추천합니다.
params = {
# Generator Parameter
'random_state': 2,
# 회전하는 최대 각도
'rotation_range': 10,
# 좌우로 이동할 최대 비율
'width_shift_range': 0.20,
# 상하로 이동할 최대 비율
'height_shift_range': 0.20,
# 회전 및 밀림 값의 최대 라디안
'shear_range': 0.50,
# 축소/확대 할 최대 비율
'zoom_range': 0.20,
'horizontal_flip': True,
'brightness_range': (0.7, 1.5),
# Model Parameter
# inception_resnet_v2 의 경우 (299, 299)를 많이 사용합니다.
'img_size': (299, 299),
'input_shape': (299, 299, 3),
'batch_size': 16,
# 한 번 split하여 generate 한 데이터 셋을 학습할 횟수
'epochs_per_generator': 5,
# Batch를 불러올 때 Multiprocessing을 사용합니다.
# 라이젠 등 코어가 많은 CPU 환경에서 많은 성능 향상을 가져올 수 있습니다.
'nb_workers': multiprocessing.cpu_count() // 2
}
# -
# 혹 다른 데이터 셋 추가(Pretrained Model Weights)로 인해 PATH가 변경된다면 아래 PATH를 수정
DATA_PATH = 'E:\\data\\2019-3rd-ml-month-with-kakr'
os.listdir(DATA_PATH)
# +
# 이미지 폴더 경로
TRAIN_IMG_PATH = os.path.join(DATA_PATH, 'cropped_train')
TEST_IMG_PATH = os.path.join(DATA_PATH, 'cropped_test')
# CSV 파일 경로
df_train = pd.read_csv(os.path.join(DATA_PATH, 'train.csv'))
df_test = pd.read_csv(os.path.join(DATA_PATH, 'test.csv'))
df_class = pd.read_csv(os.path.join(DATA_PATH, 'class.csv'))
# +
# class를 정수 값에서 문자열로 치환합니다.
df_train['class'] = df_train['class'].astype('str')
df_train = df_train[['img_file', 'class']]
df_test = df_test[['img_file']]
# +
from sklearn.model_selection import train_test_split
def train_val_split(params, df_train):
# training, validation set을
# 8:2비율로 Random한 index들로 split합니다.
its = np.arange(df_train.shape[0])
train_idx, val_idx = train_test_split(its, train_size=0.8, random_state=params['random_state'])
X_train = df_train.iloc[train_idx, :]
X_val = df_train.iloc[val_idx, :]
# 다음 번 split을 위해 random state 값을 변경합니다.
# random state 값을 지정한 이유는, 지정한 random state를 통해 index를 매번 갖게 split할 수 있어
# 디버깅에 용이하기 때문입니다.
params['random_state'] += 10
return X_train, X_val
def train_val_split_fixed(params, df_train, i, n):
# training, validation set을
# 8:2비율로 고정된 index들로 split합니다.
# 50 epochs 기준으로
# 모든 데이터를 반드시 4번씩 training 하게 되고,
# 모든 데이터는 반드시 1번씩 validaton 하게 됩니다.
its = list(range(df_train.shape[0]))
chunks = [its[j::n] for j in range(n)]
i = i % n
X_val = df_train.iloc[chunks[i], :]
chunks.pop(i)
X_train = df_train.iloc[sum(chunks, []), :]
return X_train, X_val
# +
from keras.applications.inception_resnet_v2 import InceptionResNetV2, preprocess_input
from keras.preprocessing.image import ImageDataGenerator
#
def make_generator(params, X_train, X_val):
# Define Generator config
train_datagen = ImageDataGenerator(
rotation_range=params['rotation_range'],
width_shift_range=params['width_shift_range'],
height_shift_range=params['height_shift_range'],
shear_range=params['shear_range'],
zoom_range=params['zoom_range'],
horizontal_flip=params['horizontal_flip'],
brightness_range=params['brightness_range'],
preprocessing_function=preprocess_input)
val_datagen = ImageDataGenerator(preprocessing_function=preprocess_input)
# Make Generator
train_generator = train_datagen.flow_from_dataframe(
dataframe=X_train,
directory=TRAIN_IMG_PATH,
x_col='img_file',
y_col='class',
target_size=params['img_size'],
color_mode='rgb',
class_mode='categorical',
batch_size=params['batch_size'],
seed=params['random_state']
)
validation_generator = val_datagen.flow_from_dataframe(
dataframe=X_val,
directory=TRAIN_IMG_PATH,
x_col='img_file',
y_col='class',
target_size=params['img_size'],
color_mode='rgb',
class_mode='categorical',
batch_size=params['batch_size'],
shuffle=False
)
return train_generator, validation_generator
# -
def get_steps(num_samples, batch_size):
if (num_samples % batch_size) > 0 :
return (num_samples // batch_size) + 1
else:
return num_samples // batch_size
# +
from keras.models import Sequential
from keras.layers import Dense, GlobalAveragePooling2D
cnn_model = InceptionResNetV2(include_top=False, input_shape=params['input_shape'])
model = Sequential()
model.add(cnn_model)
model.add(GlobalAveragePooling2D())
model.add(Dense(196, activation='softmax', kernel_initializer='he_normal'))
model.summary()
# -
model.compile(optimizer='adam', loss='categorical_crossentropy', metrics=['acc'])
# +
# %%time
from keras.callbacks import ModelCheckpoint, EarlyStopping
# checkpoint에서 모델을 저장할 path
filepath = os.path.join(DATA_PATH, 'inception_resnet_v2_ep{epoch:03d}_vloss-{val_loss:.4f}_vacc-{val_acc:.4f}.h5')
# 학습을 이어서 할 경우, model filename만 지정해주면 됩니다.
# from keras.models import load_model
# model_filename = 'inception_resnet_v2_ep050_vloss-0.2162_vacc-0.9361.h5'
# model = load_model(os.path.join(DATA_PATH, model_filename))
checkpoint = ModelCheckpoint(filepath, monitor='val_acc', verbose=1, save_best_only=True)
# earlystop = EarlyStopping(monitor='val_acc', min_delta=0, patience=5, verbose=1, mode='auto')
# callbacks = [checkpoint, earlystop]
callbacks = [checkpoint]
histories = []
# 최대 학습 횟수
epochs = 100
for i in range(epochs // params['epochs_per_generator']):
print(params)
X_train, X_val = train_val_split(params, df_train)
train_generator, validation_generator = make_generator(params, X_train, X_val)
params.update({
'nb_train_samples': len(X_train),
'nb_validation_samples': len(X_val)
})
histories.append(
model.fit_generator(
train_generator,
steps_per_epoch=get_steps(params['nb_train_samples'], params['batch_size']),
# 한번 generate 된 데이터를 학습할 횟수
epochs=params['epochs_per_generator'] * (i + 1),
validation_data=validation_generator,
validation_steps=get_steps(params['nb_validation_samples'], params['batch_size']),
callbacks=callbacks,
workers=params['nb_workers'],
initial_epoch=params['epochs_per_generator'] * i
)
)
save_model_filename = 'inception_resnet_v2_' + str(epochs) + '_vloss-' +\
str(round(histories[-1].history['val_loss'][-1], 4)) + '_vacc-' + str(round(histories[-1].history['val_acc'][-1], 4)) + '.h5'
model.save(os.path.join(DATA_PATH, save_model_filename))
gc.collect()
# -
# Plot training & validation loss values
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('Model loss')
plt.ylabel('Loss')
plt.xlabel('Epoch')
plt.legend(['Train', 'Val'], loc='upper left')
plt.show()
# +
# %%time
from keras.models import load_model
params.update({
'nb_test_samples': len(df_test)
})
model_filename = 'inception_resnet_v2_ep061_vloss-0.0776_vacc-0.9780.h5'
test_datagen = ImageDataGenerator(preprocessing_function=preprocess_input)
test_generator = test_datagen.flow_from_dataframe(
dataframe=df_test,
directory=TEST_IMG_PATH,
x_col='img_file',
y_col=None,
target_size=params['img_size'],
color_mode='rgb',
class_mode=None,
batch_size=params['batch_size'],
shuffle=False)
model = load_model(os.path.join(DATA_PATH, model_filename))
prediction = model.predict_generator(
generator = test_generator,
steps = get_steps(params['nb_test_samples'], params['batch_size']),
verbose=1,
workers=params['nb_workers']
)
# +
import os
predicted_class_indices=np.argmax(prediction, axis=1)
X_train, X_val = train_val_split(params, df_train)
train_generator, _ = make_generator(params, X_train, X_val)
# Generator class dictionary mapping
labels = (train_generator.class_indices)
labels = dict((v, k) for k,v in labels.items())
predictions = [labels[k] for k in predicted_class_indices]
submission = pd.read_csv(os.path.join(DATA_PATH, 'sample_submission.csv'))
submission['class'] = predictions
submission.to_csv(os.path.join(DATA_PATH, os.path.splitext(model_filename)[0] + '.csv'), index=False)
submission.head()
# -
# #### **Reference:**
# https://medium.com/@vijayabhaskar96/tutorial-on-keras-flow-from-dataframe-1fd4493d237c
# https://keras.io/
# http://www.arxiv.org/abs/1512.03385
# https://pillow.readthedocs.io/en/stable/
# https://www.kaggle.com/guglielmocamporese/macro-f1-score-keras
| inception-resnet-v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # An Introduction to Ethical Supervised Learning - Solution Manual
# <hr/>
# ## Overview:
# - Measuring performance of supervised learning predictors that are:
# - classification problems
# - binary predictors
# - Introduction to non-discriminatory ML predictor models
# - Charting performance of a TransRisk score case study to determine if it passes as non-discriminatory
# <hr/>
# ## Part 1: A Brief Introduction to Non-Discriminatory Machine Learning Predictors
# For companies that use classification based predictors, sometimes the predicted outcome of individuals within a group will fully influence the decision that is made for that individual. This needs to be treated particularly carefully when the decision being made is an <i>Important Benefit</i> - ie) health care, loan approval, or college admission. What if the data that is being used to train the model is inherently discriminatory? What if factors that created the data we use was inherently discriminatory and we didn't even know? Then the outcome predicted would also be discriminatory.<br/><br/>
# This is what non-discriminatory predictors seek to solve. For example, <b>The Equal Opportunity Model</b> requires that the true positive rate for all groups in a dataset to be the same in order to achieve fairness. What does this mean in terms of performance for binary classifiers? (Write in terms of 1's an 0's below)
# ** Write Answer Here: **
# Among all of the actual 1's, it's the percentage you predicted were 1.
# <hr/>
# ## Part 2: Introducing the TransRisk Dataset
# For this part of the tutorial, we will be working with a dataset that represents the distribution of TransRisk scores for non-defaulters (the 'Good' - people who have previously paid off their loans on time) and defaulters (the 'Bad' - people who previously haven't paid their loans on time) against four main demographic groups: Asian, Hispanic, Black, and White. Go ahead and import this data to take a look. What collected information to create TransRisk scores could be inherently discriminatory?
import pickle
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib.lines as mlines
# %matplotlib inline
totalData = pd.read_csv("ShortenedData.csv")
totalData.head()
# For loan approval, usually a bank will set a <b>threshold TransRisk score</b> that determines who is approved and who is denied. For example, if the threshold was 60: everyone with a TransRisk score below 60 would be denied the loan, and everyone with a TransRisk score above 60 would be approved a loan.
# <br/><br/>
# How should a predictor go about deciding who should get a loan and who should not? It makes sense to say all of the people who <i>deserve</i> a loan should receive one. In the case of the TransRisk score, the group of people who <i>deserve</i> a loan would be the non-defaulters.
# <br/><br/>
# Following this logic, in theory the probability of a non-defaulter getting a loan ($\hat Y$ = 1) at any threshold TransRisk score should be the same amongst all four groups. Finish the function below to plot the distribution of non-defaulters from one group getting ($\hat Y$ = 1) based on a threshold value of TransRisk scores. Then, get the probabilities for all four demographic groups and plot them on top of eachother.
def getGraphData(dataset, graphType):
dataset = dataset.set_index("TransRisk Score")
x = []
y = []
for index, row in dataset.iterrows():
curr_race_non_default = dataset.loc[index:]["Good"].sum()
total_race_non_default = dataset["Good"].sum()
yVal = curr_race_non_default / total_race_non_default
x.append(index)
y.append(yVal)
plt.plot(x, y, graphType)
white_non_default = totalData[["TransRisk Score", "Demographic", "Good"]][totalData["Demographic"] == "white"]
asian_non_default = totalData[["TransRisk Score", "Demographic", "Good"]][totalData["Demographic"] == "asian"]
black_non_default = totalData[["TransRisk Score", "Demographic", "Good"]][totalData["Demographic"] == "black"]
hispanic_non_default = totalData[["TransRisk Score", "Demographic", "Good"]][totalData["Demographic"] == "hispanic"]
# ** Plot Graph Below **
# +
getGraphData(asian_non_default, 'b-')
getGraphData(white_non_default, 'g-')
getGraphData(black_non_default, 'c-')
getGraphData(hispanic_non_default, 'm-')
plt.title("Probability of Non-Defaulters Getting $\hat Y$ = 1 (Beneficial Outcome)" )
blue_line = mlines.Line2D([], [], color='blue', marker='.',
markersize=15, label='Asian')
green_line = mlines.Line2D([], [], color='green', marker='.',
markersize=15, label='White')
cyan_line = mlines.Line2D([], [], color='cyan', marker='.',
markersize=15, label='Black')
purple_line = mlines.Line2D([], [], color='purple', marker='.',
markersize=15, label='Hispanic')
plt.legend(handles=[blue_line, green_line, cyan_line, purple_line])
# -
# <hr/>
# ## Part 3: Calculating Performance
# Now that we've seen the likelihood of non-defaulting individuals from each of the four demographic groups to be approved a loan based on threshold value, let's check the performance of this model. Take a look at the original data again. This part will be easier if you can separate the data into four different dataframes, one for each demographic. <i>Note: it is also helpful if you set the index to be the TransRisk score.</i>
whites = totalData[["TransRisk Score", "Demographic", "Good", "Bad"]][totalData["Demographic"] == "white"]
asians = totalData[["TransRisk Score", "Demographic", "Good", "Bad"]][totalData["Demographic"] == "asian"]
blacks = totalData[["TransRisk Score", "Demographic", "Good", "Bad"]][totalData["Demographic"] == "black"]
hispanics = totalData[["TransRisk Score", "Demographic", "Good", "Bad"]][totalData["Demographic"] == "hispanic"]
whites.set_index("TransRisk Score", inplace=True)
asians.set_index("TransRisk Score", inplace=True)
blacks.set_index("TransRisk Score", inplace=True)
hispanics.set_index("TransRisk Score", inplace=True)
# ** Complete : Calculate the following problem for both the White and Black demographic groups with a threshold TransRisk score of 60**
#
# For all of the individuals that <i>deserve</i> a loan, how many will receive one?
totalGood = whites["Good"].sum()
tp = whites["Good"].loc[60:].sum()
white_sensitivity = tp / totalGood
totalGood = blacks["Good"].sum()
tp = blacks["Good"].loc[60:].sum()
black_sensitivity = tp / totalGood
white_sensitivity, black_sensitivity
# <hr/>
# ### Analysis:
#
# What you just calculated is the <b>sensitivity</b> of the White and Black demographic groups for a single threshold value (60). If you recall from the beginning of this tutorial, for the Equalized Opportunity fairness model the main requirement for achieving fairness is to ensure that the true positive rates (also known as the sensitivity) are the same for all groups. As we saw in our plot from Part 2, Equalized Opportunity was definitely not being satisfied. So, how might we go about finding an easy solution to this problem? The answer lies in utilizing this performance metric. To satisfy the requirement, we need to find a point where all demographic groups have the same sensitivity and the same defaulting rate. <br/><br/>
# It makes sense that sensitivity must be the same for all demographic groups (since that is what we are trying to accomplish with the <b>Equalized Opportunity Model</b>). But why do we need to have the same defaulting rate? This is because there is going to be a separate sensitivity value for each threshold we choose. It would be easy to choose a threshold score that gives us a sensitivity of 1 (the ideal sensitivity), but then what if choosing that score creates an unfair percentage of defaulters who are allowed loans for one demographic over another? That is why it is important to make sure that the percentage of defaulters is about the same for each demographic group as well, to ensure fairness.<br/><br/>
# *** How do you choose which percent default is the best? *** <br/><br/>
# This question depends on the company that is giving out the loans. Many banks will stick to an 18% loss function. Meaning that for all of the loans they give out, there's an 18% chance that the people granted a loan will default. Based on this example, you will try to find four different threshold scores (one for each demographic) that provide the same sensitivity for all, and the same 18% probability of defaulting for all.
#
# <hr/>
# ### Step 1: Visualize the Sensitivity
# <i>Create a plot representing the sensitivity and TransRisk scores to visualize the sensitivity versus the threshold score for all demographic groups</i>
import numpy as np
def getSensitivityDF(demographic, data):
scores = []
percent_default = []
totalGood = data["Good"].sum()
sensitivity = []
for index, row in data.iterrows():
currTP = data["Good"].loc[index:].sum()
sensitivity.append(currTP / totalGood)
scores.append(index)
percent_default.append(1 - (currTP / totalGood))
df = pd.DataFrame({ 'Score' : scores,
'Demographic' : np.full(len(scores), demographic),
'Sensitivity' : sensitivity,
'Percent_Default': percent_default})
return df
white_sensitivity = getSensitivityDF("white", whites)
asian_sensitivity = getSensitivityDF("asian", asians)
black_sensitivity = getSensitivityDF("black", blacks)
hispanic_sensitivity = getSensitivityDF("hispanic", hispanics)
sensitivities = pd.concat([white_sensitivity,
asian_sensitivity,
black_sensitivity,
hispanic_sensitivity], axis=0)
plt.plot(white_sensitivity["Score"], white_sensitivity["Sensitivity"], 'b-')
plt.plot(asian_sensitivity["Score"], asian_sensitivity["Sensitivity"], 'g-')
plt.plot(black_sensitivity["Score"], black_sensitivity["Sensitivity"], 'c-')
plt.plot(hispanic_sensitivity["Score"], hispanic_sensitivity["Sensitivity"], 'm-')
# <hr/>
# ### Step 2: Utilize 18% Loss
# <i>Now that you have all of your sensitivity for each demographic, find the percent default for every calculation of sensitivity. </i> <br/><br/>
# *** Important Notes: ***
# - The sensitivity should be cumulative (because a threshold score means everyone at a score and above will recieve the loan)
# - The percent_default should also be cumulative
# - Is there an easy way to calculate percent default based off the calculations you've already made?
# +
## Solution notes:
## The cumulative percent default is just 1 - sensitivity
## Which means a percent default of 18% is a sensitivity value of ~ 82%
## So, find the sensitivity of ~ 82% for each demographic, and the score at that sensitivity
## is the score you should choose!
## Below, we calculate what the scores look like closest to 18% default for each demographic for visual clarity.
# -
min_white = min(white_sensitivity["Percent_Default"], key=lambda x:abs(x-0.18))
white_sensitivity[white_sensitivity["Percent_Default"] == min_white]
min_asian = min(asian_sensitivity["Percent_Default"], key=lambda x:abs(x-0.18))
asian_sensitivity[asian_sensitivity["Percent_Default"] == min_asian]
min_black = min(black_sensitivity["Percent_Default"], key=lambda x:abs(x-0.18))
black_sensitivity[black_sensitivity["Percent_Default"] == min_black]
min_hispanic = min(hispanic_sensitivity["Percent_Default"], key=lambda x:abs(x-0.18))
hispanic_sensitivity[hispanic_sensitivity["Percent_Default"] == min_hispanic]
# <hr/>
# ### Step 3: Find the final 'fair' scores for each demographic
# <i>Now that we have found scores for each demographic that have the same percent default of 18% and the same sensitivity, write them below. These are the four threshold values that you would need to use to determine loan approval with this dataset in order to satisfy the <b>Equal Opportunity Model.</b></i> <br/><br/>
#
# *** Write Scores Here: ***
# +
white_final = white_sensitivity[white_sensitivity["Percent_Default"] == min_white].Score
asian_final = asian_sensitivity[asian_sensitivity["Percent_Default"] == min_asian].Score
black_final = black_sensitivity[black_sensitivity["Percent_Default"] == min_black].Score
hispanic_final = hispanic_sensitivity[hispanic_sensitivity["Percent_Default"] == min_hispanic].Score
white_final, asian_final, black_final, hispanic_final
# -
# <hr/>
# ## Conclusion
# Congratulations! You have successfully taken a discriminatory set of data and utilized a machine learning fairness model to make it into a fair predictor. As our research showed, it was very obvious that the data involved in creating the supervised learning predictors for loan approval from TransRisk scores was inherently discriminatory.<br/><br/> What are some other possible solutions for optimizing performance of these models to ensure non-discriminatory decision making?
# <br/><br/>
# TransRisk data and non-discriminatory analysis courtesy of https://arxiv.org/pdf/1610.02413.pdf
| .ipynb_checkpoints/Tutorial-Solution-Condensed-Data-Correct-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plotly.com/python/getting-started/) by downloading the client and [reading the primer](https://plotly.com/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plotly.com/python/getting-started/#initialization-for-online-plotting) or [offline](https://plotly.com/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plotly.com/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# #### Imports
# The tutorial below imports [NumPy](http://www.numpy.org/), [Pandas](https://plotly.com/pandas/intro-to-pandas-tutorial/), [SciPy](https://www.scipy.org/) and [PeakUtils](http://pythonhosted.org/PeakUtils/).
# +
import plotly.plotly as py
import plotly.graph_objs as go
import plotly.figure_factory as ff
import numpy as np
import pandas as pd
import scipy
import peakutils
# -
# #### Tips
# Our method for finding the area under any peak is to find the area from the `data values` to the x-axis, the area from the `baseline` to the x-axis, and then take the difference between them. In particular, we want to find the areas of these functions defined on the x-axis interval $I$ under the peak.
#
# Let $T(x)$ be the function of the data, $B(x)$ the function of the baseline, and $Area$ the peak integration area between the baseline and the first peak. Since $T(x) \geq B(x)$ for all $x$, then we know that
#
# $$
# \begin{align}
# A = \int_{I} T(x)dx - \int_{I} B(x)dx
# \end{align}
# $$
# #### Import Data
# For our example below we will import some data on milk production by month:
# +
milk_data = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/monthly-milk-production-pounds.csv')
time_series = milk_data['Monthly milk production (pounds per cow)']
time_series = np.asarray(time_series)
df = milk_data[0:15]
table = ff.create_table(df)
py.iplot(table, filename='milk-production-dataframe')
# -
# #### Area Under One Peak
# +
baseline_values = peakutils.baseline(time_series)
x = [j for j in range(len(time_series))]
time_series = time_series.tolist()
baseline_values = baseline_values.tolist()
rev_baseline_values = baseline_values[:11]
rev_baseline_values.reverse()
area_x = [0,1,2,3,4,5,6,7,8,9,10,11,10,9,8,7,6,5,4,3,2,1]
area_y = time_series[:11] + rev_baseline_values
trace = go.Scatter(
x=x,
y=time_series,
mode='lines',
marker=dict(
color='#B292EA',
),
name='Original Plot'
)
trace2 = go.Scatter(
x=x,
y=baseline_values,
mode='markers',
marker=dict(
size=3,
color='#EB55BF',
),
name='Bassline'
)
trace3 = go.Scatter(
x=area_x,
y=area_y,
mode='lines+markers',
marker=dict(
size=4,
color='rgb(255,0,0)',
),
name='1st Peak Outline'
)
first_peak_x = [j for j in range(11)]
area_under_first_peak = np.trapz(time_series[:11], first_peak_x) - np.trapz(baseline_values[:11], first_peak_x)
area_under_first_peak
annotation = go.Annotation(
x=80,
y=1000,
text='The peak integration for the first peak is approximately %s' % (area_under_first_peak),
showarrow=False
)
layout = go.Layout(
annotations=[annotation]
)
trace_data = [trace, trace2, trace3]
fig = go.Figure(data=trace_data, layout=layout)
py.iplot(fig, filename='milk-production-peak-integration')
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'python-Peak-Integration.ipynb', 'python/peak-integration/', 'Peak Integration | plotly',
'Learn how to integrate the area between peaks and bassline in Python.',
title='Peak Integration in Python | plotly',
name='Peak Integration',
language='python',
page_type='example_index', has_thumbnail='false', display_as='peak-analysis', order=4,
ipynb= '~notebook_demo/121')
# -
| _posts/python-v3/peak-analysis/peak-integration/python-Peak-Integration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# qgrid - An interactive grid for viewing and editing pandas DataFrames
# =======================================================
# Qgrid is an Jupyter notebook widget which uses a javascript library called SlickGrid to render pandas DataFrames within a Jupyter notebook. It was developed for use in [Quantopian's hosted research environment]( https://www.quantopian.com/research?utm_source=github&utm_medium=web&utm_campaign=qgrid-nbviewer).
#
# The purpose of this notebook is to give an overview of what qgrid is capable of. Execute the cells below to generate some qgrids using a diverse set of DataFrames.
# ## Overview
# * [SlickGrid](https://github.com/mleibman/SlickGrid) is a javascript grid which allows users to scroll, sort,
# and filter hundreds of thousands of rows with extreme responsiveness.
# * [Pandas](https://github.com/pydata/pandas) is a powerful data analysis / manipulation library for Python, and DataFrames are the primary way of storing and manipulating two-dimensional data in pandas.
#
# [Qgrid](https://github.com/quantopian/qgrid) renders pandas DataFrames as SlickGrids, which enables users to explore the entire contents of a DataFrame using intuitive sorting and filtering controls. It's built on the ipywidget framework and is designed to be used in Jupyter notebook, Jupyterhub, or Jupyterlab
# ## What's new in 1.0.3
# * Added the ability to listen for events on all QgridWidget instances (using `qgrid.on`) as well as on individual instances (using `QgridWidget.on`).
# * **Breaking API Change:** Previously the recommended (but not officially documented) way of attaching event handlers to a QgridWidget instance was to listen for changes to the ``_df`` attribute using the ``observe`` method (i.e.``qgrid_widget.observe(handle_df_changed, names=['_df'])``). This method will no longer work for most events (scrolling, sorting, filtering, etc) so the new ``QgridWidget.on`` method should be used instead.
# ## API & Usage
# API documentation is hosted on [readthedocs](http://qgrid.readthedocs.io/en/widget-guidelines/).
#
# The API documentation can also be accessed via the "?" operator in IPython. To use the "?" operator, type the name of the function followed by "?" to see the documentation for that function, like this:
# ```
# qgrid.show_grid?
# qgrid.set_defaults?
# qgrid.set_grid_options?
# qgrid.enable?
# qgrid.disable?
#
# ```
# ## Example 1 - Render a DataFrame with many different types of columns
import numpy as np
import pandas as pd
import qgrid
randn = np.random.randn
df_types = pd.DataFrame({
'A' : pd.Series(['2013-01-01', '2013-01-02', '2013-01-03', '2013-01-04',
'2013-01-05', '2013-01-06', '2013-01-07', '2013-01-08', '2013-01-09'],index=list(range(9)),dtype='datetime64[ns]'),
'B' : pd.Series(randn(9),index=list(range(9)),dtype='float32'),
'C' : pd.Categorical(["washington", "adams", "washington", "madison", "lincoln","jefferson", "hamilton", "roosevelt", "kennedy"]),
'D' : ["foo", "bar", "buzz", "bippity","boppity", "foo", "foo", "bar", "zoo"] })
df_types['E'] = df_types['D'] == 'foo'
qgrid_widget = qgrid.QgridWidget(df=df_types, show_toolbar=True)
qgrid_widget
# If you make any sorting/filtering changes, or edit the grid by double clicking, you can retrieve a copy of your DataFrame which reflects these changes by calling `get_changed_df` on the `QgridWidget` instance returned by `show_grid`.
qgrid_widget.get_changed_df()
# ## Example 2 - Render a DataFrame with 1 million rows
# *Note: The reason for the redundant "import" statements in the next cell (and many subsequent cells) is because it allows us to run the cells in any order.*
# +
import pandas as pd
import numpy as np
import qgrid
# set the default max number of rows to 10 so the larger DataFrame we render don't take up to much space
qgrid.set_grid_option('maxVisibleRows', 10)
df_scale = pd.DataFrame(np.random.randn(1000000, 4), columns=list('ABCD'))
# duplicate column B as a string column, to test scalability for text column filters
df_scale['B (as str)'] = df_scale['B'].map(lambda x: str(x))
q_scale = qgrid.show_grid(df_scale, show_toolbar=True, grid_options={'forceFitColumns': False, 'defaultColumnWidth': 200})
q_scale
# -
q_scale.get_changed_df()
# ## Example 3 - Render a DataFrame returned by Yahoo Finance by enabling automatic qgrids
# +
import pandas as pd
import numpy as np
import qgrid
randn = np.random.randn
# Get a pandas DataFrame containing the daily prices for the S&P 500 from 1/1/2014 - 1/1/2017
from pandas_datareader.data import DataReader
spy = DataReader(
'SPY',
'yahoo',
pd.Timestamp('2014-01-01'),
pd.Timestamp('2017-01-01'),
)
# Tell qgrid to automatically render all DataFrames and Series as qgrids.
qgrid.enable()
# Render the DataFrame as a qgrid automatically
spy
# -
# Disable automatic display so we can display DataFrames in the normal way
qgrid.disable()
# ## Example 4 - Render a DataFrame with a multi-index
#
# Create a sample DataFrame using the `wb.download` function and render it without using qgrid
import qgrid
import pandas as pd
from pandas_datareader import wb
df_countries = wb.download(indicator='NY.GDP.PCAP.KD', country=['all'], start=2005, end=2008)
df_countries.columns = ['GDP per capita (constant 2005 US$)']
qgrid.show_grid(df_countries)
df_countries
# ## Example 5 - Render a DataFrame with an interval column
# Create a sample DataFrame using the `wb.download` function and render it without using qgrid
# +
import numpy as np
import pandas as pd
import qgrid
td = np.cumsum(np.random.randint(1, 15*60, 1000))
start = pd.Timestamp('2017-04-17')
df_interval = pd.DataFrame(
[(start + pd.Timedelta(seconds=d)) for d in td],
columns=['time'])
freq = '15Min'
start = df_interval['time'].min().floor(freq)
end = df_interval['time'].max().ceil(freq)
bins = pd.date_range(start, end, freq=freq)
df_interval['time_bin'] = pd.cut(df_interval['time'], bins)
qgrid.QgridWidget(df=df_interval, show_toolbar=True)
# -
df_interval
# ## Example 6 - Render a DataFrame with unnamed columns
# Create a sample DataFrame using the `wb.download` function and render it without using qgrid
# +
import numpy as np
import pandas as pd
import qgrid
arrays = [['bar', 'bar', 'baz', 'baz', 'foo', 'foo', 'qux', 'qux'],
['one', 'two', 'one', 'two', 'one', 'two', 'one', 'two']]
df_multi = pd.DataFrame(np.random.randn(8, 4), index=arrays)
qgrid.show_grid(df_multi, show_toolbar=True)
# -
df_multi
# ## Example 7 - Render a narrow DataFrame inside a Layout widget
# Create a sample DataFrame with only two columns using `randint`, and render it in a Layout widget that's 20% of the width of the output area.
import numpy as np
import pandas as pd
import qgrid
import ipywidgets as ipyw
randn = np.random.randn
df_types = pd.DataFrame(np.random.randint(1,14,14))
qgrid_widget = qgrid.QgridWidget(df=df_types, show_toolbar=False)
qgrid_widget.layout = ipyw.Layout(width='20%')
qgrid_widget
# ## Example 8 - Render a DataFrame with an index and column that contain multiple types
import pandas as pd
import qgrid
df = pd.DataFrame({'A': [1.2, 'xy', 4], 'B': [3, 4, 5]})
df = df.set_index(pd.Index(['yz', 7, 3.2]))
view = qgrid.QgridWidget(df=df)
view
# ## Example 9 - Render a DataFrame with a Period index and Period column
import pandas as pd
import qgrid
range_index = pd.period_range(start='2000', periods=10, freq='B')
df = pd.DataFrame({'a': 5, 'b': range_index}, index=range_index)
view = qgrid.QgridWidget(df=df)
view
# ## Example 10 - Render a DataFrame with NaN and None
import pandas as pd
import numpy as np
import qgrid
df = pd.DataFrame([(pd.Timestamp('2017-02-02'), None, 3.4), (np.nan, 2, 4.7), (pd.Timestamp('2017-02-03'), 3, None)])
qgrid.show_grid(df)
| notebooks/qgrid - basic example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
import numpy.linalg as LA
import csv
import cvxpy as cp
import math
import sys
ros_path = '/opt/ros/kinetic/lib/python2.7/dist-packages'
if ros_path in sys.path:
sys.path.remove(ros_path)
import cv2 as cv
sys.path.append('/opt/ros/kinetic/lib/python2.7/dist-packages')
import glob
# -
# termination criteria
criteria = (cv.TERM_CRITERIA_EPS + cv.TERM_CRITERIA_MAX_ITER, 30, 0.001)
# prepare object points, like (0,0,0), (1,0,0), (2,0,0) ....,(6,5,0)
objp = np.zeros((6*7,3), np.float32)
objp[:,:2] = np.mgrid[0:7,0:6].T.reshape(-1,2)
# Arrays to store object points and image points from all the images.
objpoints = [] # 3d point in real world space
imgpoints = [] # 2d points in image plane.
images = glob.glob('/home/jonathan/mav0/cali_tong/cam0/*.png')
for fname in images:
img = cv.imread(fname)
gray = cv.cvtColor(img, cv.COLOR_BGR2GRAY)
# Find the chess board corners
ret, corners = cv.findChessboardCorners(gray, (7,6), None)
# If found, add object points, image points (after refining them)
if ret == True:
objpoints.append(objp)
corners2 = cv.cornerSubPix(gray,corners, (11,11), (-1,-1), criteria)
imgpoints.append(corners)
# Draw and display the corners
cv.drawChessboardCorners(img, (7,6), corners2, ret)
cv.imshow('img', img)
cv.waitKey(250)
cv.destroyAllWindows()
ret, mtx, dist, rvecs, tvecs = cv.calibrateCamera(objpoints, imgpoints, gray.shape[::-1], None, None)
img = cv.imread('/home/jonathan/mav0/cali_tong/cam0/1403709072787836928.png')
h, w = img.shape[:2]
newcameramtx, roi = cv.getOptimalNewCameraMatrix(mtx, dist, (w,h), 1, (w,h))
# undistort
dst = cv.undistort(img, mtx, dist, None, newcameramtx)
# crop the image
x, y, w, h = roi
dst = dst[y:y+h, x:x+w]
cv.imwrite('~/calibresult.png', dst)
| ch13/.ipynb_checkpoints/cam_calibration-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # LOWER TRIANGULAR CORRELATION MATRIX
#
# A seaborn example to plot a lower triangular correlation matrix.
# +
# Import packages
from string import ascii_letters
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
sns.set(style="white")
# Generate a large random dataset
rs = np.random.RandomState(33)
d = pd.DataFrame(data=rs.normal(size=(100, 26)),
columns=list(ascii_letters[26:]))
# Compute the correlation matrix
corr = d.corr()
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(11, 9))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
plt.show()
| Seaborn - Correlation Matrix.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python
# language: python3
# name: python3
# ---
# Notebook created: 2021-08-29 11:57:45
# Generated from: source/01_comparison/includes/groupby.rst
# pandas provides a flexible `groupby` mechanism that allows similar aggregations. See the
# groupby documentation for more details and examples.
# + hide-output=false
tips_summed = tips.groupby(["sex", "smoker"])[["total_bill", "tip"]].sum()
tips_summed
| 01_comparison/includes/groupby.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import warnings
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from statsmodels.stats.outliers_influence import variance_inflation_factor
# # %matplotlib inline
warnings.filterwarnings('ignore')
# -
# # set image building to True/False
# * if build_images == False: faster to run, but images won't be generated
build_images = True
# # define functions
# +
def visualize_correlations(file_folder, df, file, annotation=True):
# constroi matriz de correlação de Pearson com valores arredondados (2 decimais)
corrmatrix = df.corr().apply(lambda x: round(x, 2))
#constroi heatmap
f, ax = plt.subplots(figsize=(10, 10))
sns.heatmap(corrmatrix, cmap='RdBu', center=0, square=True, annot=annotation, annot_kws={'fontsize':9}, cbar_kws={'shrink':.8 },
yticklabels=1, ax=ax, linewidths=0.1) #
plt.title('Variable correlations', pad=20, size=15)
plt.savefig(os.path.join(file_folder, file), bbox_inches = "tight")
def calculate_vif(df, threshold=5):
collinear_vars = []
variables = list(range(df.shape[1]))
dropped = True
while dropped:
dropped = False
vif = [variance_inflation_factor(df.iloc[:, variables].values, ix)
for ix in range(df.iloc[:, variables].shape[1])]
maxloc = vif.index(max(vif))
if max(vif) > threshold:
var = df.iloc[:, variables].columns[maxloc]
del variables[maxloc]
collinear_vars.append(var)
dropped = True
return collinear_vars
# def save_csv(sufix, label=False):
# file = sufix + '.csv'
# if label==False:
# df = eval(sufix)
# else:
# df = pd.DataFrame(eval(sufix), columns=[target_var])
# df.to_csv(os.path.join(outputs, file))
# -
def get_numerical_mask(df):
type_mask = []
for i in df.dtypes:
if str(i).startswith('float') or str(i).startswith('int'): # or str(i).startswith('bool')
type_mask.append(True)
else: type_mask.append(False)
num_cols = list(np.array(df.columns)[type_mask])
other_cols = list(np.array(df.columns)[[not elem for elem in type_mask]])
return num_cols, other_cols
# # Data capture
# +
inputs = os.path.join('..', 'data', '03_processed')
outputs = os.path.join('..', 'data', '03_processed')
reports = os.path.join('..', 'data', '06_reporting')
data = pd.read_csv(os.path.join(inputs, 'X_train_onehot.csv'), index_col='id')
data_test = pd.read_csv(os.path.join(inputs, 'X_test_onehot.csv'), index_col='id')
y_train = pd.read_csv(os.path.join(inputs, 'y_train.csv'), index_col='id')
# -
# # Visualize correlations
# ### visualize only continuous data
# +
numerical_cols, _ = get_numerical_mask(data)
# remove dummies as I already checked that they are uncorrelated
dummy_cols = [c for c in numerical_cols if c.startswith('dummy')]
numerical_cols = [c for c in numerical_cols if not c.startswith('dummy')]
# -
if build_images:
data_vis = data[numerical_cols]
data_vis['y'] = y_train['y']
visualize_correlations(reports, data_vis, '04correlations.jpg')
# # visualize dummies
if build_images:
data_vis = data[dummy_cols]
data_vis['y'] = y_train['y']
visualize_correlations(reports, data_vis, '04correlations_dummy.jpg')
# # visualize correlations with response variable
if build_images:
data_vis = data[numerical_cols]
data_vis['y'] = y_train['y']
data_vis.corr()['y'].sort_values(ascending = False).plot(kind='bar')
plt.savefig(os.path.join(reports, '04targetcorrelations.jpg'), bbox_inches = "tight")
# # Solve multicollinearity
# From the perspective of machine learning models, multicolinearity might bring some problems:
# * Coefficients become more sensitive to small changes in the model.
# * Reduction on the precision when estimating coefficients.
# One way to measure collinearity is through VIF (variance inflation factor), which measures the rise of variance of the estimation of a parameter given the adition of a new variable. To accomplish this, we can use statsmodel, a Python package. More information at:
# * https://www.statsmodels.org/stable/generated/statsmodels.stats.outliers_influence.variance_inflation_factor.html
#
# The rule of thumb is to consider the variable collinear when VIF>5. So we are going to use 5 as the threshold for the removal ot variables. An alternative would be to apply PCA on those variables, but on that case, we would lose explainability.
collinear_vars = calculate_vif(data)
collinear_vars
# # report collinear vars
import csv
with open(os.path.join(reports, 'collinear_vars.csv'),'w', newline='') as file:
writer = csv.writer(file, delimiter=',')
writer.writerow((['feature']))
for val in collinear_vars:
writer.writerow([val])
# + active=""
# # save data with collinearity treatment
# -
| notebooks/06_check_correlations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/JHORIZ-RODEL-AQUINO/OOP-1-2/blob/main/OOP_Concepts.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="zrREriDmSAHv"
# #Python Classes and Objects
# + [markdown] id="qkVLMbT_SmIE"
# Create A Class
# + id="f5Qx2h87Sc6V"
class MyClass:
pass
# + colab={"base_uri": "https://localhost:8080/"} id="stjAvmSISwMu" outputId="2737c7ee-32a0-4fad-e4e4-286f41efd1dd"
class OOP1_2:
X = 5
print(X)
# + [markdown] id="03aG-arGUHg6"
# Create Objects
# + colab={"base_uri": "https://localhost:8080/"} id="yMdySit7TGzM" outputId="d2e4797a-79dd-4660-b6fa-c8e7af7c0e69"
class OOP1_2:
def __init__(self, name, age):
self.name = name # attributes
self.age = age
def identity(self):
print(self.name, self.age)
person = OOP1_2("Jhoriz", 19) # create objects
print(person.name)
print(person.age)
print(person.identity)
# + colab={"base_uri": "https://localhost:8080/"} id="qms8riERWMMZ" outputId="fa272a26-bf55-4469-cc99-a5700f9dd362"
#Modify the Object Name
person.name = "Rodel"
print(person.name)
print(person.age)
# + colab={"base_uri": "https://localhost:8080/"} id="Zlyy-VlEW5HB" outputId="bf84690d-3596-483f-c546-037a399b021c"
person.age = 38
print(person.name)
print(person.age)
# + colab={"base_uri": "https://localhost:8080/", "height": 200} id="Ujn7e1BnXC8K" outputId="d114fd7d-cc85-491f-cf1a-4ca3b74a792c"
# Delete the Object
del person.name
# + colab={"base_uri": "https://localhost:8080/", "height": 165} id="3aQy0V6gXOBE" outputId="62cfbc65-a91c-4b72-dc92-c66c20905b91"
print(person.name)
# + colab={"base_uri": "https://localhost:8080/"} id="jY55E5fYXjDu" outputId="451abb30-5c01-4ced-9afd-d2d13fbd944f"
print(person.age)
# + [markdown] id="KwwEi_agXoYG"
# Application 1 - Write a Python program that computes the area of a square, and name its class as Square, side as attribute.
# + colab={"base_uri": "https://localhost:8080/"} id="JD5Xyr0lZLhl" outputId="e4312a16-5f1a-4d89-ca02-ef3b26788d04"
class Square:
def __init__(self, side):
self.side = side
def area(self):
return self.side * self.side
def display(self):
print("the area of the square is", self.area())
square = Square(4)
print(square.side)
square.display()
# + [markdown] id="Nm6wEG9abY-r"
# Application 2 - Write a Python program that display your full name, age, course, school.
# Create a calss named MyClass, and name, age, course and school as attributes.
# + colab={"base_uri": "https://localhost:8080/"} id="ihX76GDdfuKl" outputId="64cd0361-e76d-4bc5-b0c1-ebd364390a7f"
class MyClass:
def __init__(self, name, age, course, school):
self.name = name
self.age = age
self.course = course
self.school = school
def display_info(self):
print("Name:", self.name)
print("Age:", self.age)
print("Course:", self.course)
print("School:", self.school)
def introduce_myself(self):
print("\n\nHi guys!...")
print("..My name is " + self.name + ". I am", self.age,
"years old...")
print("...Just like y'all, I am also a first year student under " +
"the course " + self.course + " here in " + self.school + "..")
print("....I chose this course because I like the idea of engineering " +
"combined with programming.....That's all !!!")
student_1 = MyClass("<NAME>", 19,
"BS in Computer Engineering", "Cavite State University")
student_1.display_info()
student_1.introduce_myself()
| OOP_Concepts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="gBvgSF6c_rk9"
#
#
#
# Experimental code to produce a abc synthesizable netlist from a sci-kit learn random forest.
# Untested. Proceed with caution.
# author: <EMAIL>
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="V68GP6x8yf9E" outputId="e66c913b-29db-46a9-c366-7cee945723bd"
# #%tensorflow_version 1.x
import os
import re
import numpy as np
# TF is used only to read MNIST data
import tensorflow as tf
import sklearn
from sklearn.ensemble import RandomForestClassifier
print('The scikit-learn version is {}.'.format(sklearn.__version__))
print('The TF version is {}.'.format(tf.__version__))
# + colab={} colab_type="code" id="_m9iOo25y9xv"
def tree_predict(tree, node, x):
assert node >= 0
left = tree.children_left
right = tree.children_right # -1 is sentinel for none
feats = tree.feature # -2 is sentinel for none
thresh = tree.threshold
values = tree.value
if feats[node] == -2: # leaf node
assert left[node] == -1
assert right[node] == -1
return values[node] / values[node].sum()
else:
assert left[node] != -1
assert right[node] != -1
# note: we are int'ing the threshold since we don't think it matters
# as the features are all ints anyway
if x[feats[node]] <= int(thresh[node]):
return tree_predict(tree, left[node], x)
else:
return tree_predict(tree, right[node], x)
def forest_predict(model, x, debug=False):
res = tree_predict(model.estimators_[0].tree_, 0, x)
for estimator in model.estimators_[1:]:
res += tree_predict(estimator.tree_, 0, x)
if debug:
print(res.reshape(-1).astype(np.int32))
return res.reshape(-1).argmax()
def accuracy(model, examples):
return np.array([forest_predict(model, example) for example in examples])
# + colab={"base_uri": "https://localhost:8080/", "height": 323} colab_type="code" id="_zaLSBNO0Izi" outputId="aad1f57a-eedb-4c27-ccf6-c15329143f5b"
def generate(name, randomize_labels, nverify=1000):
(tx, ty), (vx, vy) = tf.keras.datasets.mnist.load_data()
if randomize_labels:
ty = np.roll(ty, 127) # np.random.permutation(ty)
tx = tx.reshape(60000, -1)
vx = vx.reshape(10000, -1)
# note we turn off bootstrap so that samples are taken without resampling
# and as a result sample weights are always 1 and so inference is simpler
# m = RandomForestClassifier(n_estimators=10, bootstrap=False, random_state=0)
# TODO: tiny tree
m = RandomForestClassifier(n_estimators=2, max_depth=3, bootstrap=False, random_state=0)
m.fit(tx, ty)
print(m)
print("name = {}, ta = {}, va = {}".format(name, m.score(tx, ty),
m.score(vx, vy)))
nverify = min(60000, nverify)
mine = accuracy(m, tx[:nverify])
golden = m.predict(tx[:nverify])
assert (mine == golden).all()
# print(np.arange(nverify)[mine != golden])
print("verified")
# write_model(m, name)
# print("done writing {}".format(name))
return m
mreal = generate('real', randomize_labels=False)
mrand = generate('random', randomize_labels=True)
# + colab={} colab_type="code" id="5yDyMjua_rlJ" outputId="a6b3ac06-c625-4321-ebd4-3908b27d02c7"
def dump_tree(tree, node, tree_id, n_classes_y, file):
assert node >= 0
left = tree.children_left
right = tree.children_right # -1 is sentinel for none
feats = tree.feature # -2 is sentinel for none
thresh = tree.threshold
values = tree.value
for i in range(n_classes_y):
print(' wire [7:0] n_{}_{}_{};'.format(tree_id, node, i), file=file)
if feats[node] == -2: # leaf node
assert left[node] == -1
assert right[node] == -1
#print(' wire [7:0] n{};'.format(node), file=file)
# for some reason (multi output classes?) tree.value has an extra dimension
assert values[node].shape == (1, n_classes_y)
class_probabilities = (values[node] / values[node].sum())[0]
for i in range(n_classes_y):
p_float = class_probabilities[i]
p_fixed = int(p_float * 255. + 0.5)
print(' assign n_{}_{}_{} = 8\'h{:x}; // {}'.format(tree_id, node, i, p_fixed, p_float), file=file)
return
else:
assert left[node] != -1
assert right[node] != -1
# note: we are int'ing the threshold since we don't think it matters
# as the features are all ints anyway
dump_tree(tree, left[node], tree_id, n_classes_y, file=file)
dump_tree(tree, right[node], tree_id, n_classes_y, file=file)
#for i in range(n_classes_y):
# print(' wire [7:0] n{}_{};'.format(node, i), file=file)
print(' wire c_{}_{};'.format(tree_id, node), file=file)
assert 0. <= thresh[node]
assert thresh[node] < 255.
threshold = int(thresh[node])
print(' assign c_{}_{} = x{} <= 8\'h{:x};'.format(tree_id, node, feats[node], threshold), file=file)
for i in range(n_classes_y):
print(' assign n_{}_{}_{} = c_{}_{} ? n_{}_{}_{} : n_{}_{}_{};'.format(
tree_id, node, i,
tree_id, node,
tree_id, left[node], i,
tree_id, right[node], i),
file=file)
def dump_verilog(model, width_x, n_classes_y):
with open('output.v', 'w') as f:
print("module forest(", file=f)
for i in range(width_x):
print(" input wire [7:0] x{}{}".format(i, ','), file=f)
for i in range(n_classes_y):
print(" output wire [15:0] y{}{}".format(i, ',' if i < n_classes_y - 1 else ''), file=f)
print(" );", file=f)
for i, estimator in enumerate(model.estimators_):
print(' // dumping tree {}'.format(i), file=f)
dump_tree(estimator.tree_, node=0, tree_id=i, n_classes_y=n_classes_y, file=f)
for c in range(n_classes_y):
print(' wire [15:0] s_{}_{};'.format(i, c), file=f)
print(' wire [15:0] e_{}_{};'.format(i, c), file=f)
print(' assign e_{}_{} = {} 8\'h0, n_{}_0_{} {};'.format(i, c, '{', i, c, '}'), file=f)
if i > 0:
print(' assign s_{}_{} = s_{}_{} + e_{}_{};'.format(i, c, i - 1, c, i, c), file=f)
else:
print(' assign s_{}_{} = e_{}_{};'.format(i, c, i, c), file=f)
for c in range(n_classes_y):
print(' assign y{} = s_{}_{};'.format(c, len(model.estimators_) - 1, c), file=f)
print("endmodule", file=f)
dump_verilog(mreal, width_x=784, n_classes_y=10)
# !head output.v
# verilator can take 3 mins to lint the resulting Verilog file if 10 trees and unlimited depth is used!
# # !verilator output.v --lint-only
# # !abc/abc -c "%read output.v; %blast; &ps; &put; write test_syn.v"
# #!cat test_syn.v
# + colab={} colab_type="code" id="RccyGmmK_rlM"
# ABC limitations:
# read silently fails whereas %read works
# if a PO is not driven an assertion fails in blast
# verilator limitations:
# sometimes when the input is bad verilator may get stuck!
| qkeras-master/experimental/forest_gen.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
x = np.linspace(0,6*np.pi,101)
y = np.sin(x)
# Not visible in "Dark Theme"
plt.plot(x,y)
plt.show()
# +
# Extra work -> worth it!
plt.figure()
ax = plt.subplot(111)
ax.plot(x,y)
# Edges...
ax.spines['bottom'].set_color('grey')
ax.spines['top'].set_color('grey')
ax.spines['right'].set_color('grey')
ax.spines['left'].set_color('grey')
# Ticks
ax.tick_params(axis='x', colors='grey')
ax.tick_params(axis='y', colors='grey')
# Labels
ax.yaxis.label.set_color('grey')
ax.xaxis.label.set_color('grey')
# Title
ax.title.set_color('grey')
plt.show()
# -
| Plots/JupyterLab Dark Theme Adjustments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="l_LulNCC8z96"
# ## Upper Confidence Bound (UCB) Algorithm
#
# 1. Apply the Upper Confidence Bound algorithm to optimize the best ad to convert the best customers' click rate
#
# 2. **Input** = Ads_CTR_Optimisation.csv
#
#
#
#
#
# + [markdown] id="xpXdowrE9DxW"
# ## Importing the libraries
# + id="nhKd4hWx9GFt"
import numpy as np
import matplotlib.pyplot as plt #graphs
import pandas as pd
# + [markdown] id="E--Vc-xXF-Si"
# # Importing the dataset
# + id="XCX22pc2F8Cz"
dataset = pd.read_csv('Ads_CTR_Optimisation.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="d9FQJ8W_ETyF" outputId="7e757ff9-1ecd-46f2-d654-9e1b90860c65"
print(len(dataset))
# + [markdown] id="9FS_CMupdWvR"
# #Implementing UCB
# + id="K_UKH0_9GSrT"
import math
# + id="ui3FyLGRCbzj"
N = 525
d = 10
ads_selected = []
numbers_of_selections = [0] * d
sums_of_rewards = [0] * d
total_reward = 0
# + id="8akq12NEFCK4"
for n in range(0,N):
ad = 0
max_upper_bound = 0
for i in range(0,d):
if numbers_of_selections[i] > 0:
average_reward = sums_of_rewards[i] / numbers_of_selections[i]
delta_i = math.sqrt(1.5*math.log(n+1)/numbers_of_selections[i])
upper_bound = average_reward + delta_i
else:
upper_bound = 1e400
if upper_bound > max_upper_bound:
max_upper_bound = upper_bound
ad = i
ads_selected.append(ad)
numbers_of_selections[ad] += 1
reward = dataset.values[n, ad]
sums_of_rewards[ad] += reward
total_reward += reward
# + id="CuVcZ6xLKuD1"
print(ads_selected)
# + [markdown] id="YsgM4SdJi-XS"
# ##Visualising the results
#
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="QzIp_DvRCKhG" outputId="4b6931d7-01c4-44db-ef2f-3e33aa5c59e3"
plt.hist(ads_selected)
plt.title('Histogram of ads selections')
plt.xlabel('Ads')
plt.ylabel('Number of times each ad was selected')
plt.show()
# + id="93o9HHSyLNQi"
| Part 6 - Reinforce Learning/Sec 32 - Upper Confidence Bound/ML_AZ_Sec32_UpperConfidenceBound.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Quantum-Classical Hybrid Machine Learning
# Quantum-classical hybrid machine learning is a kind of NISQ algorithm for machine learning using a quantum computer.
# As the name suggests, it uses a quantum computer and a classical computer hybridly.
# The basic idea is the same as VQE and QAOA.
# The quantum computer and the classical computer are run alternately, and the parameter $\theta$ that the quantum circuit has is repeatedly updated to minimize (maximize) the target quantity.
# The calculation to find the updated values of parameters is done by the classical computer.
#
# Classical computer performs the calculations that are possible with classical calculations, and by keeping the parameters as classical data, the quantum circuit can be initialized and executed repeatedly.
# This reduces the size of the quantum circuit and makes it suitable for execution on NISQ devices.
#
# VQE and QAOA defined a Hamiltonian for the problem, and updated the parameters of the quantum circuit so that the expected value of the Hamiltonian is minimum (maximum). In contrast, in quantum-classical hybrid machine learning, a loss function that takes the measurement results of the quantum circuit as its argument is defined in the classical computer side.
# As a post-processing step, the classical computer calculates the output of the loss function and updates the parameters to minimize the loss function.
# This eliminates the need to define a Hamiltonian and allows us to directly apply the loss function, which is widely used in machine learning.
# The quantum circuitry that runs in a quantum computer can be divided into three major parts.
#
# First, in the "Data encoding" layer, the input data for training is embedded into the quantum circuit.
# Next, the "Parametric quantum circuit" layer consists of a combination of two qubit gates to entangle and a rotational gate to update the parameters. It is in this part that the probability distribution of the model is learned by updating parameters.
# Finally, the "Measurement" layer measures the quantum state and passes the measurement results to the classical computer.
# Learning the parameters $\theta$ of a quantum circuit is similar to learning the weight parameters in a conventional neural network.
# Therefore, the elements necessary for successful learning are similar to those of neural networks, for example,
#
# 1. How to embed data
# 2. Loss function design
# 3. Parameter update method
# 4. Parametric quantum circuit (learning layer) design
#
# A variety of methods have been proposed and used for each of these.
# In the following tutorials, we will show some simple learning examples using concrete models.
# ### References
# Benedetti, Marcello, et al. "Parameterized quantum circuits as machine learning models." Quantum Science and Technology 4.4 (2019): 043001.
# ## Numerical differentiation
# From the function $f(x)$ and the small change $h$, the derivative can be obtained numerically.
#
# $$
# \frac{df(x)}{dx} = \lim_{h \rightarrow 0}\frac{f(x+h) - f(x)}{h}
# $$
#
# For example, with $f(x) = x^2$, $h=0.1$, the derivative at $x=1$ is as follows
x = 1
h = 0.1
((x+h)**2 - x**2) / h
# ## Partial differentiation
# In the parameterized quantum circuits often used in NISQ, many angles appear in the circuit as variable parameters.
# Partial differentiation allows you to find the differential coefficient at a particular parameter.
#
# Partial derivative of $f(x, y)$ with respect to $x$.
#
# $$
# \frac{\partial f(x,y)}{\partial x}
# $$
#
# Partial derivative of $f(x, y)$ with respect to $y$.
#
# $$
# \frac{\partial f(x,y)}{\partial y}
# $$
# ## Gradient
# The gradient is the partial derivative for each parameter written in the form of a vector.
#
# $$
# (\frac{\partial f(x,y)}{\partial x},\frac{\partial f(x,y)}{\partial y})
# $$
#
# The gradient is useful for finding the point where the function takes its minimum value (minimum point, minima).
# The derivative becomes zero at the minima, and the points around them (neighborhoods) have positive or negative slopes.
#
# - If the coordinates of the current location is greater than the minimum point, the slope is positive. Therefore, subtracting the slope from the current location coordinates will bring you closer to the minimum point.
# - If the coordinate of the current location is less than the minimum point, the slope is negative. So again, subtracting the slope from the current location coordinates will bring you closer to the minimum point.
#
# From the above, we can find the minimum point by updating the current location using the gradient. The $e$ denotes the learning rate here.
#
# $$
# x' = x -e\cdot\frac{\partial f(x,y)}{\partial x}\\
# y' = y -e\cdot\frac{\partial f(x,y)}{\partial y}
# $$
#
# Repeat this process.
# ## Gradient descent
# Let's try VQE with a circuit using RY gates to find the minimum expected value of a measurement.
# +
from blueqat import Circuit
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
def abs_sq(k):
return np.square(np.abs(k))
# Z expectation value
def exptZ(a):
res = Circuit().ry(a)[0].run()
return abs_sq(res[0]) - abs_sq(res[1])
# Initail setting
ainit = [np.random.rand()*2*np.pi]
a = ainit.copy()
h = 0.001
e = 0.01
# Update and logging of gradient
for i in range(1000):
a.append(a[i] - e*(exptZ(a[i]+h) - exptZ(a[i]))/h)
# Last value
print(a[-1])
# Plot expectation value
plt.plot(list(map(exptZ, a)))
plt.show()
# -
# The $Z$ measurement expectation value of one qubit is minimized when the state is $\lvert 1\rangle$.
# After minimizing the expectation value, the $RY$ gate rotation angle is approximately $\pi = 3.14... $, which changes the initial state $\lvert 0\rangle$ to $\lvert 1\rangle$.
# From the above, we succeeded VQE.
# ## Parameter shift rule
#
# Reference: arXiv:1803.00745 [quant-ph] (https://arxiv.org/abs/1803.00745)
# Numerical differentiation with a small numerical value $h$ has a problem in running on a real quantum computer.
# The expected value computed by a quantum computer has variance due to sampling and noise. If the small numerical value $h$ is too small, it will be buried in those variance, and if it is too large, the numerical differentiation will be inaccurate. Therefore, it is necessary to adjust $h$ as a hyperparameter.
#
# Against this background, research is also being conducted on methods for calculating the gradient in quantum computers.
# Here we explain a typical method, "parameter shift rule".
#
# Think about $\langle \hat{B}\rangle$, expected value of an observable $\hat{B}$.
# For the parameter $\theta$ in the Pauli rotation gate $U(\theta) = \exp{(-i\theta P)}\ (P\in \{X, Y, Z, I\})$, the gradient of $\langle \hat{B}\rangle$ is obtained as follows
#
#
# $$\frac{\partial \langle \hat{B}\rangle}{\partial \theta} = \frac{1}{2} \bigl(\langle \hat{B}\rangle_+ - \langle \hat{B}\rangle_- \bigr)$$
#
# $$U(\theta + \frac{\pi}{2}) = \langle \hat{B}\rangle_+$$
# $$U(\theta - \frac{\pi}{2}) = \langle \hat{B}\rangle_-$$
#
# In other words, for each parameter for which you want to find the gradient, it's sufficinet to run the $+\frac{\pi}{2}$ and $-\frac{\pi}{2}$ shifted quantum circuits and calculate the expected value, respectively.
# Since this method uses a fixed and sufficiently large shift $\pm\frac{\pi}{2}$ instead of a small change $h$, the above issue can be solved.
#
# This can be applied to parameters $\theta$ of, such as, $RX(\theta)$ gate, $RY(\theta)$ gate, and $RZ(\theta)$ gate. By combining this with a two-qubit entangle gate, it is possible to construct a general quantum-classical hybrid machine learning circuit.
#
# The following is an example of the gradient descent method calculation from earlier again using the parameter shift rule.
# +
# Initail setting
ainit = [np.random.rand()*2*np.pi]
a = ainit.copy()
e = 0.01
# Update and logging of gradient
for i in range(1000):
grad = (exptZ(a[i]+np.pi/2) - exptZ(a[i]-np.pi/2)) / 2
a.append(a[i] - e*grad)
# Last value
print(a[-1])
# Plot expectation value
plt.plot(list(map(exptZ, a)))
plt.show()
# -
# ## Loss function
# VQE minimizes the expected value of measurement, but in the above approach, the object of minimization does not necessarily have to be the expected value of measurement.
# Here we will use the more general loss function. This way we can recognize it as machine learning.
# +
from blueqat import Circuit
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
def abs_sq(k):
return np.square(np.abs(k))
# expectation value
def expt(a):
res = Circuit().ry(a)[0].run()
return abs_sq(res[0])-abs_sq(res[1])
# loss function
def loss(res,tar):
return np.square(res-tar)
# derivative
def dfx(a,h,tgt=-1):
return (loss(expt(a+h),tgt) - loss(expt(a),tgt))/h
# return (expt(a+h)-expt(a))/h
# gradient
def grad(a,h,e):
for i in range(1000):
a.append(a[i] - e*dfx(a[i],h))
return a
# initialize
a = [np.random.rand()*2*np.pi]
h = 0.001
e = 0.01
ra = grad(a,h,e)
print(ra[-1])
arr = [loss(expt(i),-1) for i in ra]
plt.plot(arr)
plt.show()
print(arr[-1])
# -
# ## momentum SGD
# +
#momentum sgd
def msgd(a,h,e,tgt,alpha):
p_delta = 0
for i in range(100):
update = -e*dfx(a[i],h,tgt) + alpha*p_delta
a.append(a[i] + update)
p_delta = update
return a
#initialization
a = ainit.copy()
h = 0.001
e = 0.01
alpha = 0.9
tgt = -1
result = msgd(a,h,e,tgt,alpha)
arr = [loss(expt(i),tgt) for i in result]
plt.plot(arr)
plt.show()
# -
# ## Adagrad
# +
def adagrad(a,h,e,tgt,epsilon):
G = epsilon
for i in range(100):
g = dfx(a[i],h,tgt)
G += g*g
update = -e/np.sqrt(G)*g
a.append(a[i] + update)
return a
#initialization
a = ainit.copy()
h = 0.001
e = 0.1
epsilon = 1e-08
tgt = -1
result = adagrad(a,h,e,tgt,epsilon)
arr = [loss(expt(i),tgt) for i in result]
plt.plot(arr)
plt.show()
# -
| tutorial/250_quantum_classical_hybrid.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from scipy.special import gamma as gamma_function # Check if correct gamma function
from scipy.stats import norm
from scipy.stats import gamma
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# +
def q_tau(tau, a, b):
return gamma.pdf(tau, a, loc=0, scale=1/b)
def q_mu(x, mu, tau, lam=1.0):
return norm.pdf(x, mu, lam*tau**-.5)
# +
lam_0 = 1.0
a_0 = 10
b_0 = 3.5
mu_0 = 0
mus = np.linspace(-3, 3, 100)
taus = np.linspace(0, 6, 100)
M, T = np.meshgrid(mus, taus, indexing="ij")
Z = np.zeros_like(M)
for i in range(Z.shape[0]):
for j in range(Z.shape[1]):
Z[i][j] = q_mu(mus[i], mu_0, taus[j], lam_0) * q_tau(taus[j], a_0, b_0)
plt.contour(M, T, Z)
plt.xlabel(r'$\mu$')
plt.ylabel(r'$\tau$')
# +
mu = 0
lam = 1.0
a = 2
b =1
def q_mu(x):
return norm.pdf(x, mu, np.sqrt(1 / lam))
def q_tau(tau):
return gamma.pdf(tau, a, loc = 0, scale = 1 / b)
# +
mus = np.linspace(-3, 3, 100)
taus = np.linspace(0, 6, 100)
M, T = np.meshgrid(mus, taus, indexing="ij")
Z = np.zeros_like(M)
for i in range(Z.shape[0]):
for j in range(Z.shape[1]):
Z[i][j] = q_mu(mus[i]) * q_tau(taus[j])
plt.contour(M, T, Z)
plt.xlabel(r'$\mu$')
plt.ylabel(r'$\tau$')
# -
| notebooks/randomsample/variational_guassian_gamma.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from tqdm import tqdm
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import normalize
train = pd.read_csv('data/train_texts.csv')
test = pd.read_csv('data/test_texts.csv')
# В обязательной части поработаем только с текстами. Пока просто любопытно посмотреть, что будет и есть ли смысл усложнять; более того, в BOW / tf-idf репрезентациях получается громадная размерность данных.
train.head()
X_text = train['text']
y = train['class']
# ### BOW
from sklearn.feature_extraction.text import CountVectorizer
vectorizer = CountVectorizer()
vectorizer.fit(X_text)
X_bow = normalize(vectorizer.transform(X_text))
X_bow.shape
X_train, X_val, y_train, y_val = train_test_split(X_bow, y, test_size=0.05, random_state=42)
X_train.shape, X_val.shape
# С мешком слов попробуем логистическую регрессию.
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression(max_iter=10000, random_state=42).fit(X_train, y_train)
clf.score(X_train, y_train), clf.score(X_val, y_val)
# 70%, уже неплохо!
clf = LogisticRegression(max_iter=10000, random_state=42).fit(X_bow, y)
clf.score(X_train, y_train), clf.score(X_val, y_val)
X_test, X_id = normalize(vectorizer.transform(test['text'])), test['id']
def classifier_out(test_data, model, filename):
y_test = model.predict(test_data)
out_rows=list(zip(X_id, y_test))
out_rows = [('Id','Predicted')] + out_rows
out_rows = [f'{t[0]},{t[1]}' for t in out_rows]
with open(f'submissions/{filename}.csv', 'w') as a:
a.write('\n'.join(out_rows))
a.close()
classifier_out(X_test, clf, '1.bow_lr')
# В Kaggle это позволило достичь резов в 0.72. Неплохо для элементарного классификатора!
# ### TF-IDF
from sklearn.feature_extraction.text import TfidfTransformer
X_train_counts = vectorizer.transform(X_text)
X_test_counts = vectorizer.transform(test['text'])
tfidf = TfidfTransformer().fit(X_train_counts)
X = tfidf.transform(X_train_counts)
X_test = tfidf.transform(X_test_counts)
X_train, X_val, y_train, y_val = train_test_split(X, y, test_size=0.05, random_state=42)
X_train.shape, X_val.shape
# С tf-idf попробуем наивный байесовский классификатор.
from sklearn.naive_bayes import MultinomialNB
nb_split = MultinomialNB().fit(X_train, y_train)
nb_split.score(X_train, y_train), nb_split.score(X_val, y_val)
# Результаты чуть хуже, чем у lr + bow
nb_full = MultinomialNB().fit(X, y)
nb_full.score(X_train, y_train), nb_split.score(X_val, y_val)
classifier_out(X_test, nb_full, '2.tfidf_nb')
# 0.68. Хуже! Попробуем логистическую регрессию с tf-idf же.
lr_tfidf_split = LogisticRegression(max_iter=10000, random_state=42).fit(X_train, y_train)
lr_tfidf_split.score(X_train, y_train), lr_tfidf_split.score(X_val, y_val)
# Выглядит как теоретическое улучшение относительно обычного мешка слов!
lr_tfidf_full = LogisticRegression(max_iter=10000, random_state=42).fit(X, y)
lr_tfidf_full.score(X_train, y_train), lr_tfidf_full.score(X_val, y_val)
classifier_out(X_test, lr_tfidf_full, '3.tfidf_lr')
# Улучшили на 1%. Ура, товарищи! Дальше попробуем tensorflow. (прошу обратиться к 3. YABAI.ipynb)
| 2. Sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
word=str(input('word'))
if word.endswith('ch'):
print(word, 'es')
elif word.endswith('sh'):
print(word, 'es')
else:
print(word, 's')
# +
n=input('请输入你的名字')
d=float(input('请输入你的生日'))
if 1.19<d<2.19:
print(n,'你是水瓶座')
elif 2.18<d<3.21:
print(n,'你是双鱼座')
elif 3.20<d<4.20:
print(n,'你是白羊座')
elif 4.19<d<5.21:
print(n,'你是金牛座')
elif 5.20<d<6.22:
print(n,'你是双子座')
elif 6.21<d<7.23:
print(n,'你是巨蟹座')
elif 7.22<d<8.23:
print(n,'你是狮子座')
elif 8.22<d<9.23:
print(n,'你是处女座')
elif 9.22<d<10.23:
print(n,'你是天秤座')
elif 10.22<d<11.23:
print(n,'你是天蝎座')
elif 11.22<d<12.32 or 1.00<d<1.20:
print(n,'你是射手座')
# +
m=int(input('请输入一个整数'))
n=int(input('请输入一个非零整数'))
print('你想干嘛,回车输入;1为求和;2为求积;3为求商的余数')
p=int(input('输入1或2或3'))
total=m+n
y=m*n
s=n%m
q=m%n
if p==1:
if m<n:
while m<n:
m+=1
total+=m
else:
print(total-m)
if n<m:
while n<m:
n+=1
total+=n
else:
print(total-n)
if p==2:
if m<n:
while m<n:
m=m+1
y=y*m
else:
print(y/m)
if n<m:
while n<m:
n=n+1
y=y*n
else:
print(y/n)
if p==3:
if m<n:
print(s)
if n<m:
print(q)
# -
p=float(input('空气污染pm2.5指数'))
if p>500:
print('空气重度污染,请打开空气净化器,戴口罩')
elif 200<p<500:
print('当前污染指数较高')
elif p<200:
print('当前空气污染较小放心出行')
| chapter1/homework/localization/3-22/201611680228(task3).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="PVO3ASMSm8Bx" outputId="3668dd9c-c5e6-4f5c-edd3-544bc1e6d077"
from google.colab import drive
drive.mount('/content/drive')
# + id="uLrd5bwInAd1"
path="/content/drive/MyDrive/Research/cods_comad_plots/SDC/dataset_4/"
# + colab={"base_uri": "https://localhost:8080/"} id="N2_J4Rw2r0SQ" outputId="e66b4c3b-d8f9-44e0-df4f-59dbc7269c57"
import numpy as np
import pandas as pd
from matplotlib import pyplot as plt
from tqdm import tqdm
# %matplotlib inline
from torch.utils.data import Dataset, DataLoader
import torch
import torchvision
import torch.nn as nn
import torch.optim as optim
from torch.nn import functional as F
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
print(device)
# + [markdown] id="F6fjud_Fr0Sa"
# # Generate dataset
# + colab={"base_uri": "https://localhost:8080/"} id="CqdXHO0Cr0Sd" outputId="be8bf807-b022-4712-898c-f80c569fdda7"
np.random.seed(12)
y = np.random.randint(0,10,5000)
idx= []
for i in range(10):
print(i,sum(y==i))
idx.append(y==i)
# + id="ddhXyODwr0Sk"
x = np.zeros((5000,2))
# + id="DyV3N2DIr0Sp"
np.random.seed(12)
x[idx[0],:] = np.random.multivariate_normal(mean = [4,6.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[0]))
x[idx[1],:] = np.random.multivariate_normal(mean = [5.5,6],cov=[[0.01,0],[0,0.01]],size=sum(idx[1]))
x[idx[2],:] = np.random.multivariate_normal(mean = [4.5,4.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[2]))
x[idx[3],:] = np.random.multivariate_normal(mean = [3,3.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[3]))
x[idx[4],:] = np.random.multivariate_normal(mean = [2.5,5.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[4]))
x[idx[5],:] = np.random.multivariate_normal(mean = [3.5,8],cov=[[0.01,0],[0,0.01]],size=sum(idx[5]))
x[idx[6],:] = np.random.multivariate_normal(mean = [5.5,8],cov=[[0.01,0],[0,0.01]],size=sum(idx[6]))
x[idx[7],:] = np.random.multivariate_normal(mean = [7,6.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[7]))
x[idx[8],:] = np.random.multivariate_normal(mean = [6.5,4.5],cov=[[0.01,0],[0,0.01]],size=sum(idx[8]))
x[idx[9],:] = np.random.multivariate_normal(mean = [5,3],cov=[[0.01,0],[0,0.01]],size=sum(idx[9]))
# + colab={"base_uri": "https://localhost:8080/"} id="qh1mDScsU07I" outputId="7fb99a8e-c2a9-4ef3-a608-9ba56ba8e7b4"
x[idx[0]][0], x[idx[5]][5]
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="9Vr5ErQ_wSrV" outputId="d4161888-d2f7-4833-89e4-55ffc792a8de"
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="NG-3RpffwU_i" colab={"base_uri": "https://localhost:8080/"} outputId="58419c8c-df2b-41ac-8877-64d2ca27259c"
bg_idx = [ np.where(idx[3] == True)[0],
np.where(idx[4] == True)[0],
np.where(idx[5] == True)[0],
np.where(idx[6] == True)[0],
np.where(idx[7] == True)[0],
np.where(idx[8] == True)[0],
np.where(idx[9] == True)[0]]
bg_idx = np.concatenate(bg_idx, axis = 0)
bg_idx.shape
# + colab={"base_uri": "https://localhost:8080/"} id="_0JKKUdwoo9b" outputId="e9dcaaf0-659a-4df0-b033-583c6303127f"
np.unique(bg_idx).shape
# + id="6LA5i3Droo60"
x = x - np.mean(x[bg_idx], axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/"} id="s5tfCDfHoo3-" outputId="28f6bd02-6982-4868-d27a-f52829f48d8b"
np.mean(x[bg_idx], axis = 0, keepdims = True), np.mean(x, axis = 0, keepdims = True)
# + id="TpSAZtqzoo1F"
x = x/np.std(x[bg_idx], axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/"} id="NsrCmT8IovQv" outputId="1e8bac86-7b3e-4f5a-ed6e-7b8a7d8a7427"
np.std(x[bg_idx], axis = 0, keepdims = True), np.std(x, axis = 0, keepdims = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="hJ8Jm7YUr0St" outputId="a50918c8-a59a-493e-fa04-d8b9b2590f03"
for i in range(10):
plt.scatter(x[idx[i],0],x[idx[i],1],label="class_"+str(i))
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
# + id="3lMBZEHNBlF2"
foreground_classes = {'class_0','class_1', 'class_2'}
background_classes = {'class_3','class_4', 'class_5', 'class_6','class_7', 'class_8', 'class_9'}
# + colab={"base_uri": "https://localhost:8080/"} id="blRbGZHeCwXU" outputId="531cda2e-798c-4884-add6-79371aaaf1bd"
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,9)
a = []
for i in range(9):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
print(a.shape)
print(fg_class , fg_idx)
# + id="Y43sWeX7C15F" colab={"base_uri": "https://localhost:8080/"} outputId="7971202d-8e99-4926-ef4b-8625418716f2"
a.shape
# + colab={"base_uri": "https://localhost:8080/"} id="ooII7N6UDWe0" outputId="115db9f4-b81a-4a36-8301-02642dc16b85"
np.reshape(a,(18,1)) #not required
# + id="g21bvPRYDL9k" colab={"base_uri": "https://localhost:8080/", "height": 284} outputId="bf6c25c9-a229-4dcf-e184-7dd9cb30147b"
a=np.reshape(a,(9,2))
plt.imshow(a)
# + id="OplNpNQVr0S2"
desired_num = 2000
mosaic_list_of_images =[]
mosaic_label = []
fore_idx=[]
for j in range(desired_num):
np.random.seed(j)
fg_class = np.random.randint(0,3)
fg_idx = np.random.randint(0,9)
a = []
for i in range(9):
if i == fg_idx:
b = np.random.choice(np.where(idx[fg_class]==True)[0],size=1)
a.append(x[b])
# print("foreground "+str(fg_class)+" present at " + str(fg_idx))
else:
bg_class = np.random.randint(3,10)
b = np.random.choice(np.where(idx[bg_class]==True)[0],size=1)
a.append(x[b])
# print("background "+str(bg_class)+" present at " + str(i))
a = np.concatenate(a,axis=0)
mosaic_list_of_images.append(a)
mosaic_label.append(fg_class)
fore_idx.append(fg_idx)
# + id="dwZVmmRBr0S8"
# mosaic_list_of_images = np.concatenate(mosaic_list_of_images,axis=1).T
# + colab={"base_uri": "https://localhost:8080/"} id="OoxzYI-ur0S_" outputId="5f9224ab-3d44-4868-85be-f098a12107ab"
len(mosaic_list_of_images), mosaic_list_of_images[0]
# + colab={"base_uri": "https://localhost:8080/"} id="Nbefu9LEnMu1" outputId="7b599e36-c65f-424e-cf5f-e9bb060100e8"
mosaic_list_of_images_reshaped = np.reshape(mosaic_list_of_images, (2000,9,2))
mean_train = np.mean(mosaic_list_of_images_reshaped[0:1000], axis=0, keepdims= True)
print(mean_train.shape, mean_train)
std_train = np.std(mosaic_list_of_images_reshaped[0:1000], axis=0, keepdims= True)
print(std_train.shape, std_train)
# + id="yldL3sUvnMsE"
mosaic_list_of_images = ( mosaic_list_of_images_reshaped - mean_train ) / std_train
# + colab={"base_uri": "https://localhost:8080/"} id="qWboEmnwnX_r" outputId="4103872b-da24-435e-bfc4-7e4854c3071a"
print(np.mean(mosaic_list_of_images[0:1000], axis=0, keepdims= True))
print(np.std(mosaic_list_of_images[0:1000], axis=0, keepdims= True))
# + colab={"base_uri": "https://localhost:8080/"} id="88zCElC_naib" outputId="a1644fe9-d5ac-4976-86b8-929d6675c62d"
print(np.mean(mosaic_list_of_images[1000:2000], axis=0, keepdims= True))
print(np.std(mosaic_list_of_images[1000:2000], axis=0, keepdims= True))
# + colab={"base_uri": "https://localhost:8080/"} id="3DE8AXVAngEj" outputId="2f0f1b34-2103-4e7d-c9dd-b144d22028e4"
mosaic_list_of_images.shape
# + id="iPoIwbMHx44n"
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label, fore_idx):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx], self.fore_idx[idx]
# + id="fOPAJQJeW8Ah"
batch = 250
msd1 = MosaicDataset(mosaic_list_of_images[0:1000], mosaic_label[0:1000] , fore_idx[0:1000])
train_loader = DataLoader( msd1 ,batch_size= batch ,shuffle=True)
# + id="qjNiQgxZW8bA"
batch = 250
msd2 = MosaicDataset(mosaic_list_of_images[1000:2000], mosaic_label[1000:2000] , fore_idx[1000:2000])
test_loader = DataLoader( msd2 ,batch_size= batch ,shuffle=True)
# + id="30ZAjix3x8CM"
class Focus(nn.Module):
def __init__(self):
super(Focus, self).__init__()
self.fc1 = nn.Linear(2, 1, bias=False)
torch.nn.init.zeros_(self.fc1.weight)
# self.fc2 = nn.Linear(50, 10)
# self.fc3 = nn.Linear(10, 1)
def forward(self,z): #y is avg image #z batch of list of 9 images
y = torch.zeros([batch,2], dtype=torch.float64)
x = torch.zeros([batch,9],dtype=torch.float64)
y = y.to("cuda")
x = x.to("cuda")
# print(x.shape, z.shape)
for i in range(9):
# print(z[:,i].shape)
# print(self.helper(z[:,i])[:,0].shape)
x[:,i] = self.helper(z[:,i])[:,0]
# print(x.shape, z.shape)
x = F.softmax(x,dim=1)
# print(x.shape, z.shape)
# x1 = x[:,0]
# print(torch.mul(x[:,0],z[:,0]).shape)
for i in range(9):
# x1 = x[:,i]
y = y + torch.mul(x[:,i,None],z[:,i])
# print(x.shape, y.shape)
return x, y
def helper(self, x):
x = x.view(-1, 2)
# x = F.relu(self.fc1(x))
# x = F.relu(self.fc2(x))
x = (self.fc1(x))
return x
# + id="0dYXnywAD-4l"
class Classification(nn.Module):
def __init__(self):
super(Classification, self).__init__()
self.fc1 = nn.Linear(2, 3)
torch.nn.init.xavier_normal_(self.fc1.weight)
torch.nn.init.zeros_(self.fc1.bias)
def forward(self, x):
x = x.view(-1, 2)
x = self.fc1(x)
# print(x.shape)
return x
# + id="lSa6O9f6XNf4"
torch.manual_seed(12)
focus_net = Focus().double()
focus_net = focus_net.to("cuda")
# + id="36k3H2G-XO9A"
torch.manual_seed(12)
classify = Classification().double()
classify = classify.to("cuda")
# + colab={"base_uri": "https://localhost:8080/"} id="hOJ8NP14n6Eb" outputId="ec015359-73f8-45fb-a9da-9ae7ab4964f5"
focus_net.helper( torch.randn((1,9,2)).double().to("cuda") )
# + colab={"base_uri": "https://localhost:8080/"} id="estYkNt8oKhL" outputId="7ab9ec9d-d5a6-4c93-e8e2-f7cf70deacc4"
focus_net.fc1.weight
# + colab={"base_uri": "https://localhost:8080/"} id="TNA0bvAeoEpU" outputId="74128812-8c09-4c98-de00-c5df42d7ce60"
classify.fc1.weight, classify.fc1.bias, classify.fc1.weight.shape
# + id="7MHBbL7FXRQk"
import torch.optim as optim
criterion = nn.CrossEntropyLoss()
optimizer_classify = optim.Adam(classify.parameters(), lr=0.01 ) #, momentum=0.9)
optimizer_focus = optim.Adam(focus_net.parameters(), lr=0.01 ) #, momentum=0.9)
# + id="pjD2VZuV9Ed4"
col1=[]
col2=[]
col3=[]
col4=[]
col5=[]
col6=[]
col7=[]
col8=[]
col9=[]
col10=[]
col11=[]
col12=[]
col13=[]
# + id="uALi25pmzQHV" colab={"base_uri": "https://localhost:8080/"} outputId="b00cfbb4-1d76-4126-c6b4-498eb59aba8e"
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
# print(outputs.shape)
_, predicted = torch.max(outputs.data, 1)
# print(predicted.shape)
for j in range(labels.size(0)):
count += 1
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
# print(focus, fore_idx[j], predicted[j])
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
print(count)
print("="*100)
col1.append(0)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
# + id="4vmNprlPzTjP" colab={"base_uri": "https://localhost:8080/"} outputId="9869bfef-040d-4266-bade-dfd1452ef818"
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
# + id="Yl41sE8vFERk" colab={"base_uri": "https://localhost:8080/"} outputId="0cfb7565-15a9-4ff7-cdc6-f047eae4a8dc"
nos_epochs = 1000
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for epoch in range(nos_epochs): # loop over the dataset multiple times
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
running_loss = 0.0
epoch_loss = []
cnt=0
iteration = desired_num // batch
#training data set
for i, data in enumerate(train_loader):
inputs , labels , fore_idx = data
inputs, labels = inputs.to("cuda"), labels.to("cuda")
inputs = inputs.double()
# zero the parameter gradients
optimizer_focus.zero_grad()
optimizer_classify.zero_grad()
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
# print(outputs)
# print(outputs.shape,labels.shape , torch.argmax(outputs, dim=1))
loss = criterion(outputs, labels)
loss.backward()
optimizer_focus.step()
optimizer_classify.step()
running_loss += loss.item()
mini = 3
if cnt % mini == mini-1: # print every 40 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, cnt + 1, running_loss / mini))
epoch_loss.append(running_loss/mini)
running_loss = 0.0
cnt=cnt+1
if epoch % 5 == 0:
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
if(np.mean(epoch_loss) <= 0.001):
break;
if epoch % 5 == 0:
col1.append(epoch + 1)
col2.append(argmax_more_than_half)
col3.append(argmax_less_than_half)
col4.append(focus_true_pred_true)
col5.append(focus_false_pred_true)
col6.append(focus_true_pred_false)
col7.append(focus_false_pred_false)
# print("="*20)
# print("Train FTPT : ", col4)
# print("Train FFPT : ", col5)
#************************************************************************
#testing data set
# focus_net.eval()
with torch.no_grad():
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range (batch):
focus = torch.argmax(alphas[j])
if(alphas[j][focus] >= 0.5):
argmax_more_than_half +=1
else:
argmax_less_than_half +=1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true +=1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false +=1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false +=1
col8.append(argmax_more_than_half)
col9.append(argmax_less_than_half)
col10.append(focus_true_pred_true)
col11.append(focus_false_pred_true)
col12.append(focus_true_pred_false)
col13.append(focus_false_pred_false)
# print("Test FTPT : ", col10)
# print("Test FFPT : ", col11)
# print("="*20)
print('Finished Training')
# + id="5gQoPST5zW2t"
df_train = pd.DataFrame()
df_test = pd.DataFrame()
# + id="In76SYH_zZHV"
columns = ["epochs", "argmax > 0.5" ,"argmax < 0.5", "focus_true_pred_true", "focus_false_pred_true", "focus_true_pred_false", "focus_false_pred_false" ]
# + id="BS4HtOHEzZ0E"
df_train[columns[0]] = col1
df_train[columns[1]] = col2
df_train[columns[2]] = col3
df_train[columns[3]] = col4
df_train[columns[4]] = col5
df_train[columns[5]] = col6
df_train[columns[6]] = col7
df_test[columns[0]] = col1
df_test[columns[1]] = col8
df_test[columns[2]] = col9
df_test[columns[3]] = col10
df_test[columns[4]] = col11
df_test[columns[5]] = col12
df_test[columns[6]] = col13
# + id="1UbTkfLUINTI" colab={"base_uri": "https://localhost:8080/", "height": 424} outputId="74d279fa-dba8-41a3-c5e6-a67a62c2f3ff"
df_train
# + colab={"base_uri": "https://localhost:8080/", "height": 573} id="xyna3USAeKZ9" outputId="1460ba45-627a-4111-ede6-0d73e6a4e962"
# plt.figure(12,12)
plt.plot(col1,col2, label='argmax > 0.5')
plt.plot(col1,col3, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.title("On Training set")
plt.show()
plt.plot(col1,col4, label ="focus_true_pred_true ")
plt.plot(col1,col5, label ="focus_false_pred_true ")
plt.plot(col1,col6, label ="focus_true_pred_false ")
plt.plot(col1,col7, label ="focus_false_pred_false ")
plt.title("On Training set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("training data")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="5lhODjqGoSxL" outputId="30d81ec0-00e3-4883-bf09-28bd2229147c"
plt.figure(figsize=(6,5))
plt.plot(col1,np.array(col4)/10, label ="FTPT")
plt.plot(col1,np.array(col5)/10, label ="FFPT")
plt.plot(col1,np.array(col6)/10, label ="FTPF")
plt.plot(col1,np.array(col7)/10, label ="FFPF")
plt.title("Dataset4 - SDC On Train set")
plt.grid()
# plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.legend() #loc='upper right')
plt.xlabel("epochs", fontsize=14, fontweight = 'bold')
plt.ylabel("percentage train data", fontsize=14, fontweight = 'bold')
plt.savefig(path+"ds4_train.png", bbox_inches="tight")
plt.savefig(path+"ds4_train.pdf", bbox_inches="tight")
plt.savefig("ds4_train.png", bbox_inches="tight")
plt.savefig("ds4_train.pdf", bbox_inches="tight")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="hrb68lvueMA-" outputId="e7e65a50-1c87-42f9-ad49-fa94eb0810ab"
df_test
# + colab={"base_uri": "https://localhost:8080/", "height": 573} id="qR5SnitGeNMN" outputId="6f005526-cde3-4ab1-980a-388ea8af229b"
# plt.figure(12,12)
plt.plot(col1,col8, label='argmax > 0.5')
plt.plot(col1,col9, label='argmax < 0.5')
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.title("On Testing set")
plt.show()
plt.plot(col1,col10, label ="focus_true_pred_true ")
plt.plot(col1,col11, label ="focus_false_pred_true ")
plt.plot(col1,col12, label ="focus_true_pred_false ")
plt.plot(col1,col13, label ="focus_false_pred_false ")
plt.title("On Testing set")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.xlabel("epochs")
plt.ylabel("Testing data")
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="vwP4uAKyoaq7" outputId="6a069b58-1604-436d-df53-6b173dfb964d"
plt.figure(figsize=(6,5))
plt.plot(col1,np.array(col10)/10, label ="FTPT")
plt.plot(col1,np.array(col11)/10, label ="FFPT")
plt.plot(col1,np.array(col12)/10, label ="FTPF")
plt.plot(col1,np.array(col13)/10, label ="FFPF")
plt.title("Dataset4 - SDC On Test set")
plt.grid()
# plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.legend() #loc='upper right')
plt.xlabel("epochs", fontsize=14, fontweight = 'bold')
plt.ylabel("percentage test data", fontsize=14, fontweight = 'bold')
plt.savefig(path+"ds4_test.png", bbox_inches="tight")
plt.savefig(path+"ds4_test.pdf", bbox_inches="tight")
plt.savefig("ds4_test.png", bbox_inches="tight")
plt.savefig("ds4_test.pdf", bbox_inches="tight")
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="TGrzYDWxeO6m" outputId="ad718dd3-c239-4287-c775-c5b79af342e1"
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 train images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
# + colab={"base_uri": "https://localhost:8080/"} id="67_H9pkCeQNs" outputId="4bd4ac52-935a-448d-a94b-c2b5c4c87a10"
correct = 0
total = 0
count = 0
flag = 1
focus_true_pred_true =0
focus_false_pred_true =0
focus_true_pred_false =0
focus_false_pred_false =0
argmax_more_than_half = 0
argmax_less_than_half =0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels , fore_idx = inputs.to("cuda"),labels.to("cuda"), fore_idx.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
for j in range(labels.size(0)):
focus = torch.argmax(alphas[j])
if alphas[j][focus] >= 0.5 :
argmax_more_than_half += 1
else:
argmax_less_than_half += 1
if(focus == fore_idx[j] and predicted[j] == labels[j]):
focus_true_pred_true += 1
elif(focus != fore_idx[j] and predicted[j] == labels[j]):
focus_false_pred_true += 1
elif(focus == fore_idx[j] and predicted[j] != labels[j]):
focus_true_pred_false += 1
elif(focus != fore_idx[j] and predicted[j] != labels[j]):
focus_false_pred_false += 1
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 test images: %d %%' % (
100 * correct / total))
print("total correct", correct)
print("total train set images", total)
print("focus_true_pred_true %d =============> FTPT : %d %%" % (focus_true_pred_true , (100 * focus_true_pred_true / total) ) )
print("focus_false_pred_true %d =============> FFPT : %d %%" % (focus_false_pred_true, (100 * focus_false_pred_true / total) ) )
print("focus_true_pred_false %d =============> FTPF : %d %%" %( focus_true_pred_false , ( 100 * focus_true_pred_false / total) ) )
print("focus_false_pred_false %d =============> FFPF : %d %%" % (focus_false_pred_false, ( 100 * focus_false_pred_false / total) ) )
print("argmax_more_than_half ==================> ",argmax_more_than_half)
print("argmax_less_than_half ==================> ",argmax_less_than_half)
# + colab={"base_uri": "https://localhost:8080/"} id="jhvhkEAyeRpt" outputId="085e5709-dd26-463a-fae4-972996dbcaed"
correct = 0
total = 0
with torch.no_grad():
for data in train_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 train images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + colab={"base_uri": "https://localhost:8080/"} id="OKcmpKwGeS8M" outputId="0dcb52e6-a049-48e4-9b73-bfd8669170fc"
correct = 0
total = 0
with torch.no_grad():
for data in test_loader:
inputs, labels , fore_idx = data
inputs = inputs.double()
inputs, labels = inputs.to("cuda"), labels.to("cuda")
alphas, avg_images = focus_net(inputs)
outputs = classify(avg_images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 1000 test images: %d %%' % ( 100 * correct / total))
print("total correct", correct)
print("total train set images", total)
# + id="xlalQug_egcl"
| CODS_COMAD/SDC on all datasets/type4_focus_linear_classify_linear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: desc-python
# language: python
# name: desc-python
# ---
# Notebook contributed by <NAME> (@rmandelb)
#
# Last Verified to Run: 2019-03-14 (by @rmandelb)
#
# This notebook illustrates some basic analysis of the galaxy populations for Run 1.2i/p. Because of some known issues with these test runs, the analysis is a bit more complicated than it will be for future runs. In particular, the fact that Run 1.2p has a bug that results in no extinction being applied (galaxy internal extinction or MW extinction) while Run 1.2i does include extinction means that even with the same input catalogs, the galaxy populations are not expected to look the same. The implications of this are explored in various places below.
#
# Work on this notebook also contributed to validation of DM outputs for Run 1.2i and to better understanding of data products needed for Run 1.2x:
#
# 1. Some issues were found with an earlier version of the object catalogs in Run 1.2i, c.f. https://github.com/LSSTDESC/DC2-production/issues/308; the current version of this notebook uses a reprocessed tract for which this problem was fixed. The older version mentioned in that issue is `dc2_object_run1.2i_alpha`.
#
# 2. It became clear that the truth catalogs for Run 1.2 as of January 2019 could only be used to analyze Run 1.2p data, since they do not include extinction effects. As a result, a new truth catalog including extinction was produced, to enable object vs. truth comparisons for Run 1.2i
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
import GCRCatalogs
from GCR import GCRQuery
import matplotlib
matplotlib.rcParams.update({'font.size': 16})
# Establish some basic quantities up top.
#
# What tract will we use everywhere for the Run 1.2x?
# The tract below was chosen as an example; it's fine to use others.
tract_num = 4850
# Magnitude cut in i-band, determined such that galaxies in the coadd should be well-detected
# (a reasonably complete sample far from the flux limit).
max_mag_i = 25.0
# Load the object catalogs for Run 1.2x and for the HSC XMM field. We want to check the reference band information, which is not available
# in the smaller 'dc2_object_run1.2x' catalogs, so that's why we use the larger versions with all columns.
catalog_i = GCRCatalogs.load_catalog('dc2_object_run1.2i_all_columns')
catalog_p = GCRCatalogs.load_catalog('dc2_object_run1.2p_all_columns')
catalog_h = GCRCatalogs.load_catalog('hsc-pdr1-xmm')
# Establish some basic cuts intended to yield a galaxy sample with reasonable flux measurements.
basic_cuts = [
GCRQuery('extendedness > 0'), # Extended objects (primarily galaxies)
GCRQuery((np.isfinite, 'mag_i_cModel')), # Select objects that have i-band cmodel magnitudes
GCRQuery('clean'), # The source has no flagged pixels (interpolated, saturated, edge, clipped...)
# and was not skipped by the deblender
GCRQuery('xy_flag == 0'), # Flag for bad centroid measurement
]
# Define cuts on galaxy properties: currently we simply make a sample limited at i<25.
properties_cuts = [
GCRQuery('mag_i_cModel < %f'%max_mag_i),
]
# We can now extract our sample with basic and property cuts. Should decide first what quantities we want.
quantities_sim = ['mag_i_cModel', 'snr_i_cModel', 'ra', 'dec', 'mag_g_cModel', 'mag_r_cModel',
'magerr_i_cModel', 'psf_fwhm_i', 'Ixx_i', 'Ixy_i', 'Iyy_i',
'psFlux_i', 'psFluxErr_i',
'merge_measurement_u', 'merge_measurement_g', 'merge_measurement_r', 'merge_measurement_i', 'merge_measurement_z', 'merge_measurement_y']
quantities = ['mag_i_cModel', 'snr_i_cModel', 'ra', 'dec', 'mag_g_cModel', 'mag_r_cModel',
'magerr_i_cModel', 'psf_fwhm_i', 'Ixx_i', 'Ixy_i', 'Iyy_i']
data_basic_i = catalog_i.get_quantities(quantities_sim,
filters=basic_cuts+properties_cuts,
native_filters=['tract == %d'%tract_num])
data_basic_p = catalog_p.get_quantities(quantities_sim,
filters=basic_cuts+properties_cuts,
native_filters=['tract == %d'%tract_num])
# For HSC, there is no tract selection.
data_basic_h = catalog_h.get_quantities(quantities,
filters=basic_cuts+properties_cuts)
# Some basic diagnostics here.
print('Number of objects passing cuts in Run 1.2i, Run1.2p:')
print(len(data_basic_i['ra']), len(data_basic_p['ra']))
print('Which reference band for the objects in Run 1.2i? Numbers are ugrizy:')
print(np.sum(data_basic_i['merge_measurement_u'].astype(int)),
np.sum(data_basic_i['merge_measurement_g'].astype(int)),
np.sum(data_basic_i['merge_measurement_r'].astype(int)),
np.sum(data_basic_i['merge_measurement_i'].astype(int)),
np.sum(data_basic_i['merge_measurement_z'].astype(int)),
np.sum(data_basic_i['merge_measurement_y'].astype(int)))
print('Which reference band for the objects in Run 1.2p? Numbers are ugrizy:')
print(np.sum(data_basic_p['merge_measurement_u'].astype(int)),
np.sum(data_basic_p['merge_measurement_g'].astype(int)),
np.sum(data_basic_p['merge_measurement_r'].astype(int)),
np.sum(data_basic_p['merge_measurement_i'].astype(int)),
np.sum(data_basic_p['merge_measurement_z'].astype(int)),
np.sum(data_basic_p['merge_measurement_y'].astype(int)))
# The numbers above illustrate basic object counts and choices of reference band. The signature of https://github.com/LSSTDESC/DC2-production/issues/308 was that only u and g were reference bands for Run 1.2i in the original processing. Now it's mostly i-band, as for Run 1.2p.
# Take a basic look at the tract, make sure its geometry is roughly rectangular on the sky (no major holes etc.).
plt.figure(figsize=(10,10))
plt.subplot(111)
plt.hexbin(data_basic_i['ra'], data_basic_i['dec'])
plt.xlabel('RA')
plt.ylabel('Dec')
plt.colorbar()
# The geometry of the tract is very simple and rectangular, with only tiny holes, so we are going to use naive area estimates.
# Approximate area estimate for this tract assuming rectangle geometry.
ra_min = np.min(data_basic_i['ra'])
ra_max = np.max(data_basic_i['ra'])
dec_min = np.min(data_basic_i['dec'])
dec_max = np.max(data_basic_i['dec'])
delta_ra = ra_max-ra_min
delta_dec = dec_max-dec_min
cosdec = np.cos(np.median(data_basic_i['dec']*np.pi/180))
area_est = (delta_ra*cosdec)*delta_dec
print('Tract %d area is approximately %.2f square degrees'%(tract_num,area_est))
coord_filters = [
'ra >= {}'.format(ra_min),
'ra < {}'.format(ra_max),
'dec >= {}'.format(dec_min),
'dec < {}'.format(dec_max),
]
# Number density estimate in square arcmin
num_den_i = len(data_basic_i['ra']) / area_est / 60**2
num_den_p = len(data_basic_p['ra']) / area_est / 60**2
print('Number density for Run 1.2i, Run1.2p with consistent cuts in tract %d: %.1f, %.1f (per sq arcmin)'%(tract_num, num_den_i, num_den_p))
# Now we want to read in the truth catalog roughly within the area of this tract. We select only non-sprinkled galaxies (not stars).
# Deal with different labeling of magnitude columns as well. Important note: truth catalogs for Run 1.2p and 1.2i differ because of the
# Run 1.2p extinction bug.
mag_filters = [
(np.isfinite, 'mag_i'),
'mag_i < %f'%max_mag_i,
]
truth_cat_p = GCRCatalogs.load_catalog('dc2_truth_run1.2_static')
truth_cat_i = GCRCatalogs.load_catalog('dc2_truth_run1.2_static_galaxies')
truth_cat_p.add_quantity_modifier('mag_i', truth_cat_p.get_quantity_modifier('mag_true_i'))
truth_cat_p.add_quantity_modifier('mag_g', truth_cat_p.get_quantity_modifier('mag_true_g'))
truth_cat_p.add_quantity_modifier('mag_r', truth_cat_p.get_quantity_modifier('mag_true_r'))
truth_cat_p.add_quantity_modifier('mag_z', truth_cat_p.get_quantity_modifier('mag_true_z'))
truth_cat_i.add_quantity_modifier('mag_i', truth_cat_i.get_quantity_modifier('mag_true_i'))
truth_cat_i.add_quantity_modifier('mag_g', truth_cat_i.get_quantity_modifier('mag_true_g'))
truth_cat_i.add_quantity_modifier('mag_r', truth_cat_i.get_quantity_modifier('mag_true_r'))
truth_cat_i.add_quantity_modifier('mag_z', truth_cat_i.get_quantity_modifier('mag_true_z'))
truth_native_filters = (coord_filters + ['i < %f'%max_mag_i, 'star==0', 'sprinkled==0'])
truth_data_p = truth_cat_p.get_quantities(['ra', 'dec', 'object_id', 'star', 'sprinkled', 'mag_i', 'mag_g', 'mag_r', 'mag_z'],
filters=mag_filters, native_filters=truth_native_filters)
truth_data_i = truth_cat_i.get_quantities(['ra', 'dec', 'object_id', 'star', 'sprinkled', 'mag_i', 'mag_g', 'mag_r', 'mag_z'],
filters=mag_filters, native_filters=truth_native_filters)
truth_num_den_p = len(truth_data_p['ra']) / area_est / 60**2
truth_num_den_i = len(truth_data_i['ra']) / area_est / 60**2
# Look at HSC coverage. Just a few holes.
plt.figure(figsize=(10,10))
plt.subplot(111)
plt.hexbin(data_basic_h['ra'], data_basic_h['dec'])
plt.xlabel('RA')
plt.ylabel('Dec')
plt.colorbar()
# Approximate area estimate for HSC XMM catalog (turns out it is similar in area to the simulation tract we are inspecting)
ra_min = np.min(data_basic_h['ra'])
ra_max = np.max(data_basic_h['ra'])
dec_min = np.min(data_basic_h['dec'])
dec_max = np.max(data_basic_h['dec'])
delta_ra = ra_max-ra_min
delta_dec = dec_max-dec_min
cosdec = np.cos(np.median(data_basic_h['dec']*np.pi/180))
area_est_h = (delta_ra*cosdec)*delta_dec
print('HSC XMM approximate catalog area in sq deg: %.1f'%area_est_h)
num_den_h = len(data_basic_h['ra']) / area_est_h / 60**2
# Now we plot the *normalized* i-band magnitude distributions in Runs 2.1p, 2.1i, the truth catalogs, and HSC.
# They are normalized so we can focus on the shape of the distribution.
# However, the legend indicates the total number density of galaxies selected with our magnitude cut,
# which lets us find issues with the overall number density matching (or not).
# Both Run 1.2i and Run 1.2p have colors that do not reflect the extragalactic catalogs, for reasons explained
# in https://github.com/LSSTDESC/DC2-production/issues/291.
plt.figure(figsize=(10,10))
plt.subplot(111)
nbins = 50
myrange = [20,max_mag_i]
plt.title('Tract %d'%tract_num)
data_to_plot = [
data_basic_i['mag_i_cModel'],
truth_data_i['mag_i'],
data_basic_p['mag_i_cModel'],
truth_data_p['mag_i'],
data_basic_h['mag_i_cModel']
]
labels_to_plot = [
r'Run 1.2i object catalog: %.1f/arcmin$^2$'%num_den_i,
r'Run 1.2i truth catalog: %.1f/arcmin$^2$'%truth_num_den_i,
r'Run 1.2p object catalog: %.1f/arcmin$^2$'%num_den_p,
r'Run 1.2p truth catalog: %.1f/arcmin$^2$'%truth_num_den_p,
r'HSC XMM field: %.1f/arcmin$^2$'%num_den_h
]
for ind in range(len(data_to_plot)):
plt.hist(data_to_plot[ind], nbins, range=myrange, histtype='step',
label=labels_to_plot[ind], linewidth=2.0, density=True)
plt.legend(loc='upper left')
plt.xlabel('i-band magnitude')
plt.ylabel('normalized distribution')
plt.yscale('log')
# Some commentary on the results shown above:
#
# - The HSC XMM field shows the number counts of galaxies in real data. It follows a fairly typical power-law shape. None of the simulations has as high of a normalization because they are based on an early version of protoDC2 that had a smaller density of galaxies for i<25 (this is a known offset).
# - For Run 1.2p, the truth catalog and the object catalogs show quite consistent number densities and shapes of the distributions. This is a nice sanity check of the results.
# - The same is true for Run 1.2i truth vs. object catalogs.
# - For Run 1.2i, the number counts and shape of the distribution differ from Run 1.2p for reasons that are understood: Run 1.2p image simulations did not include extinction and Run 1.2i did, so there should be fewer detections in Run 1.2i.
# Now g-band magnitude (but don't cut at the same place as in i-band).
plt.figure(figsize=(10,10))
plt.subplot(111)
myrange = [20,max_mag_i+1]
data_to_plot = [
data_basic_i['mag_g_cModel'],
truth_data_i['mag_g'],
data_basic_p['mag_g_cModel'],
truth_data_p['mag_g'],
data_basic_h['mag_g_cModel'],
]
plt.title('Tract %d'%tract_num)
for ind in range(len(data_to_plot)):
plt.hist(data_to_plot[ind], nbins, range=myrange, histtype='step',
label=labels_to_plot[ind], linewidth=2.0, density=True)
plt.legend(loc='upper left')
plt.xlabel('g-band magnitude')
plt.yscale('log')
# Look at the image simulations and HSC (no truth): what is the distribution of PSF FWHM in i-band?
# These are not expected to match in any way, it's just useful information for interpreting the
# results for other quantities.
data_to_plot = [
data_basic_i['psf_fwhm_i'],
data_basic_p['psf_fwhm_i'],
data_basic_h['psf_fwhm_i']
]
labels_to_plot = [
'Run 1.2i object catalog',
'Run 1.2p object catalog',
'HSC XMM field'
]
plt.figure(figsize=(10,10))
plt.subplot(111)
myrange = [0.5,0.9]
plt.title('Tract %d'%tract_num)
for ind in range(len(data_to_plot)):
plt.hist(data_to_plot[ind], nbins, range=myrange, histtype='step',
label=labels_to_plot[ind], linewidth=2.0, density=True)
plt.legend(loc='upper left')
plt.xlabel('i-band PSF FWHM [arcsec]')
plt.yscale('log')
# In this tract, it happens to be the case that the seeing is better in Run 1.2i than in HSC or Run 1.2p. This is relevant to interpreting the PSF-convolved object sizes that are plotted later in this notebook.
# Look at the observed r-i color versus i-band magnitude. (2D histograms)
data_x_to_plot = [
data_basic_i['mag_r_cModel']-data_basic_i['mag_i_cModel'],
truth_data_i['mag_r']-truth_data_i['mag_i'],
data_basic_p['mag_r_cModel']-data_basic_p['mag_i_cModel'],
truth_data_p['mag_r']-truth_data_p['mag_i'],
data_basic_h['mag_r_cModel']-data_basic_h['mag_i_cModel']
]
data_y_to_plot = [
data_basic_i['mag_i_cModel'],
truth_data_i['mag_i'],
data_basic_p['mag_i_cModel'],
truth_data_p['mag_i'],
data_basic_h['mag_i_cModel']
]
labels_to_plot = [
'Run 1.2i object catalog',
'Run 1.2i truth catalog',
'Run 1.2p object catalog',
'Run 1.2p truth catalog',
'HSC XMM field'
]
nbins_2d = 25
myrange = [[-1, 2], [20, 25]]
fig = plt.figure(figsize=(10,10))
for ind in range(len(data_x_to_plot)):
ax = plt.subplot(3,2,ind+1)
ax.hist2d(data_x_to_plot[ind], data_y_to_plot[ind], nbins_2d, range=myrange, normed=True)
plt.title(labels_to_plot[ind])
# Overall axis label
ax = fig.add_subplot(111, frameon=False)
plt.tick_params(labelcolor='none', top='off', bottom='off', left='off', right='off')
plt.grid(False)
plt.xlabel('r-i color')
plt.ylabel('i-band magnitude')
# The above 5-panel plot shows the 2D distribution of the apparent r-i color (horizontal axis) and the apparent i-band magnitude (vertical axis). The two top panels are Run 1.2i object (left) and truth (right); the two middle panels are the comparable quantities for Run 1.2p. The bottom is real HSC data. Run 1.2i, 1.2p, and HSC cannot be compared with each other because of known issues with the inputs, but for a given run, comparing truth vs. object catalogs is fair.
# For the image simulations and HSC (on truth), look at the 2D distribution of i-band magnitude vs.
# i-band magnitude uncertainty. We expect fainter galaxies to have larger magnitude uncertainty.
data_x_to_plot = [
data_basic_i['mag_i_cModel'],
data_basic_p['mag_i_cModel'],
data_basic_h['mag_i_cModel'],
]
data_y_to_plot = [
data_basic_i['magerr_i_cModel'],
data_basic_p['magerr_i_cModel'],
data_basic_h['magerr_i_cModel'],
]
labels_to_plot = [
'Run 1.2i object catalog',
'Run 1.2p object catalog',
'HSC XMM field'
]
myrange = [[20,25], [0,0.25]]
fig = plt.figure(figsize=(10,10))
for ind in range(len(data_x_to_plot)):
ax = plt.subplot(2,2,ind+1)
ax.hist2d(data_x_to_plot[ind], data_y_to_plot[ind], nbins_2d, range=myrange, normed=True)
plt.title(labels_to_plot[ind])
# Overall axis label
ax = fig.add_subplot(111, frameon=False)
plt.tick_params(labelcolor='none', top='off', bottom='off', left='off', right='off')
plt.grid(False)
plt.ylabel('i-band magnitude error')
plt.xlabel('i-band magnitude')
# Calculate the determinant radius for the observed (PSF-convolved) galaxies.
i_det_size = (data_basic_i['Ixx_i']*data_basic_i['Iyy_i']-data_basic_i['Ixy_i']**2)**0.25
p_det_size = (data_basic_p['Ixx_i']*data_basic_p['Iyy_i']-data_basic_p['Ixy_i']**2)**0.25
h_det_size = (data_basic_h['Ixx_i']*data_basic_h['Iyy_i']-data_basic_h['Ixy_i']**2)**0.25
# Look at the apparent magnitude vs. apparent size trend line (includes PSF convolution).
# Units are pixels, so the sims vs. HSC comparison is not quite fair.
data_x_to_plot = [
data_basic_i['mag_i_cModel'][np.isfinite(i_det_size)],
data_basic_p['mag_i_cModel'][np.isfinite(p_det_size)],
data_basic_h['mag_i_cModel'][np.isfinite(h_det_size)],
]
data_y_to_plot = [
i_det_size[np.isfinite(i_det_size)],
p_det_size[np.isfinite(p_det_size)],
h_det_size[np.isfinite(h_det_size)],
]
labels_to_plot = [
'Run 1.2i object catalog',
'Run 1.2p object catalog',
'HSC XMM field'
]
myrange = [[20,25], [0,4]]
fig = plt.figure(figsize=(10,10))
for ind in range(len(data_x_to_plot)):
ax = plt.subplot(2,2,ind+1)
ax.hist2d(data_x_to_plot[ind], data_y_to_plot[ind], nbins_2d, range=myrange, normed=True)
plt.title(labels_to_plot[ind])
# Overall axis label
ax = fig.add_subplot(111, frameon=False)
plt.tick_params(labelcolor='none', top='off', bottom='off', left='off', right='off')
plt.grid(False)
plt.xlabel('i-band magnitude')
plt.ylabel('Apparent radius [pixels]')
# In this final plot, we have the i-band magnitude (horizontal axis) versus the apparent radius in units of pixels, calculated as the determinant of the moment matrix of the PSF-convolved galaxies. There are a few obvious features in this plot:
#
# - In each panel, there is a lower cutoff in the values of the apparent radius. The lower cutoff arises because of the PSF size. For Run 1.2i, the PSF is smaller than in HSC or Run 1.2p, and so the cutoff is lower.
#
# - For HSC and Run 1.2i, it's clear that there is a mild anti-correlation between magnitude and galaxy size, i.e., fainter galaxies are smaller. In Run 1.2p this anticorrelation is not really apparent, but it could be because we're looking at a tract where the seeing is worse.
| contributed/Run_1.2_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/manjulamishra/DS-Code-Pandas_Useful_Functions/blob/master/Simple_and_Complex_Filtering_pandas.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="yCCJNTbLl39z" colab_type="text"
# # Data Filtering
#
# https://www.ritchieng.com/pandas-multi-criteria-filtering/
#
# After data cleaning, this is an import step to take a peak into data.
# + id="zY7pmRAlliPg" colab_type="code" colab={}
import pandas as pd
import numpy as np
# + id="sqU3zK4rmRAI" colab_type="code" colab={}
# let's get a dataset to do some filtering on
url = 'http://bit.ly/imdbratings'
movies = pd.read_csv(url)
# + id="Bc3MDnDmmco6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 198} outputId="1084e9bf-616b-4ca7-90a5-5b36e527b35c"
movies.head()
# + id="FcY6q0nvnAZ6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="efe667d5-28c6-42bf-d155-2869273f41f1"
movies.shape
# + id="kN39STbnmtFg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 145} outputId="590c0606-d6a4-4a5a-a39a-1e880b0ef048"
movies.isna().sum()
# + [markdown] id="Xn0d9IC2oBlU" colab_type="text"
# ## Filtering Rows of pandas DF by Column values
# + id="aFzBRjRCm3Ks" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 126} outputId="a6a8e7c3-5b14-406d-ee7d-31523e8a1d63"
# create a Series/list of True/False using a conditional
# create a filter
is_long = movies.duration >= 200
is_long.head()
# + id="jTQwRJVhpVbu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 615} outputId="f4e7a40e-2e40-415e-8f40-54eb1d105bf0"
# use that filter to filter the df based on that filter
# the result is only the movies which are longer than 200 minutes
movies[is_long]
# + [markdown] id="0c3RSmOerVkE" colab_type="text"
# ### A better way to filter the same rows as above
# + id="Z-6a-ixypX3N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 615} outputId="ceb9a4d2-6e84-4b17-e326-2ad3e8b322af"
# an even better way
movies[movies['duration'] >= 200]
# + [markdown] id="NTBF0IH-ruQ7" colab_type="text"
# ### We want to filter df based on duration and only the genre instead of all the columns
# + id="x5wwb5rYrmhq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 254} outputId="94555088-5430-4820-df58-c43e3eed0db9"
# it outputs the index number and the genre for the movies which either or longer than 300 minutes
# movies[movies['duration']>= 200].genre
movies[movies.duration >= 200]['genre']
# + [markdown] id="JnotKFQVsreG" colab_type="text"
# ### best practice is to use .loc to achieve the same result as above
# + id="Y1xCszPesTJO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 408} outputId="8a2392bc-48a0-449e-a7fd-bcdf2f4c9864"
movies.loc[movies['duration']>= 200, ['genre']]
# + id="orH1LDl9D0HE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b4e43acf-07b0-4684-c762-fd1487face86"
type(movies['duration']>= 200)
# + id="1YPSO3JpxxYa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="9f788909-18ac-4b70-b3d5-6a8fa05b9086"
movies.loc[:, 'genre']
# + [markdown] id="ubHZ3n8dGTjk" colab_type="text"
# **Logic Behind the Above Code**
#
# How the above logic works is by creating a boolean series based on the conditional.
#
# movies[movies['duration] >= 200]
#
# creates a conditional inside which is stored as True/Flase series in a temp variable. It;s same as creating a True/False series explicitily and then passing it inside movies like
#
# is_a_long_move = movies['duration'] >= 200
#
# movies[is_a_long_movie]
#
# Now this will spit out a resulting dataframe where the conditons are True with all column.
#
#
# + [markdown] id="F5BDeqOYHXDX" colab_type="text"
# ### Manually Creating a list og booleans
# + id="Vx8PHU4SyYXa" colab_type="code" colab={}
# create a list
booleans = []
# loop
for minutes in movies['duration']:
if minutes >= 200:
booleans.append(True)
else:
booleans.append(False)
# + id="WrkGvrXy-8np" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="da6e9925-2c05-4ca4-8065-e3fddb6019bf"
len(booleans)
# + id="bQNUmyr7Hz5J" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="eac7e56f-fa29-4e98-9dbd-547787f484cf"
booleans[0:5]
# + [markdown] id="Dh3yiZvbH_nR" colab_type="text"
# **Convert the booleans list into pd.Series**
#
# To make the list usable in our dataframe, we have to convert it into pandas Series object.
# + id="BaRqTSf1H9Uc" colab_type="code" colab={}
long_movies = pd.Series(booleans)
# + [markdown] id="P7z8vcqpIbiR" colab_type="text"
# **pandas operations work on Series e.g. head()**
#
# Since we convereted python list into pandas Series, we can not access the elements using head() method which doesn't work on Python list. Additionally, we can also use the same indexing as on python list on a Series.
# + id="ZjMM2FWVIWy5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 126} outputId="b03c47f0-3bcc-4b7a-e41b-9dc6fbcbbdb9"
# both will produce the same result
# long_movies[0:5]
long_movies.head()
# + id="FBIZeAOaIZWj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 615} outputId="cd5df361-01d2-4705-9af3-3dda4e1fead1"
movies[long_movies]
# + [markdown] id="keUjQ_nTYTZS" colab_type="text"
# ## Multiple Criteria Filerting
# + [markdown] id="TJmHUJ_DYZjx" colab_type="text"
# When the conditionals are wrapped in paranthesis, this is an ordered operation that the brackets are executed first then 'and'
#
# **and (&)**
# + id="gX5KCULzLN29" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 138} outputId="dc92b17d-9780-4249-a0cd-a67987a44f4f"
movies.loc[(movies['duration'] >= 200) & (movies['genre']=='Drama')]
# + [markdown] id="hSM27CTrZSUr" colab_type="text"
# **or**
#
# The movies whihc are not Drama
# + id="PWDMiYZqLRuF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 580} outputId="a19d9528-b84b-4121-acc3-9411598e9f33"
movies[movies['duration'] >= 200 | (movies['genre']=='Drama')]
# + [markdown] id="ldjHKiLya5G1" colab_type="text"
# **Creating a boolean based on two Condtionals**
#
# It creates True false list of the same length as the DF. True if those two conditions are met, else False.
# + id="F690_kmjZce7" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="e612116b-81a6-4b8f-932d-d5dddaf332e6"
# (movie['duration] >= 200) & (movie['genre] == 'Drama') # just the oppostive conditional
(movies['duration'] >= 200) | (movies['genre'] == 'Drama')
# + [markdown] id="NXxSLj13bzwk" colab_type="text"
# **Using OR**
#
# Either the movie is more than 200 minutes long or the genre is Drama or Crime or Action.
# + id="UckFjzXzbJb-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 580} outputId="7c827580-b821-4dea-b747-8357293a9054"
movies[(movies['duration'] >= 200) | (movies['genre']=='Drama') | (movies['genre']=='Crime') | (movies['genre']=='Action')]
# + [markdown] id="9c5umvO5eBtt" colab_type="text"
# ### Fast filtering method
# + id="E7Oho6mVcUTg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 580} outputId="16e8776f-349e-4aba-8e39-3ac8ab2f714d"
filter_list = ['Drama', 'Crime', 'Action']
movies[movies['genre'].isin(filter_list)]
# + id="V4guMQCMeV38" colab_type="code" colab={}
# + [markdown] id="oqQRqjSKcOXx" colab_type="text"
# **More on Complex Queries**
#
# temp2 = df[~df["Def"] & (df["days since"] > 7) & (df["bin"] == 3)]
#
# Or
#
# cond1 = df["bin"] == 3
# cond2 = df["days since"] > 7
# cond3 = ~df["Def"]
#
# temp2 = df[cond1 & cond2 & cond3]
# + id="yD6rhHyocWBv" colab_type="code" colab={}
| Simple_and_Complex_Filtering_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Using `ChemicalEnvironments`
#
# Chemical Environments were created as a way to parse SMIRKS strings and make changes in chemical perception space.
# In this workbook, we will show you have chemical environments are initiated and used to make changes to a SMIRKS pattern.
#
# **Authors**
# * <NAME> from Mobley Group at University of California, Irvine
#
# **Basic Structure of `ChemicalEnvironments`**
#
# `ChemicalEnvironments` are initiated with the following input variables:
# * smirks = any SMIRKS string (if None an empty environment is created)
# * label = this could be anything, a number/str/int, stored at ChemicalEnvironment.label
# * replacements = This is a list of two tuples in the form (short, smirks) to substitute short hand in your SMIRKS strings. This is used to check if your input SMIRKS string or created Chemical Environment are Valid.
#
# **SMIRKS Strings**
#
# Here we use the word SMIRKS to mean SMARTS patterns with indexed atoms, we are not using Chemical Environments to parse SMIRKS strings that describe reactions.
# That means these SMIRKS patterns should not contain multiple molecules (`'.'`) or reaction arrows (`'>>'`).
# Here we will try to explain the SMIRKS patterns used here, but SMARTS and SMIRKS are a complex language.
# SMARTS/SMIRKS strings are similar to SMILES strings with increased complexity.
# For more details about this language see the Daylight tutorials:
# * [SMILES](http://www.daylight.com/dayhtml_tutorials/languages/smiles/index.html)
# * [SMARTS](http://www.daylight.com/dayhtml/doc/theory/theory.smarts.html)
# * [SMIRKS](http://www.daylight.com/dayhtml_tutorials/languages/smirks/index.html)
# import necessary functions
from openforcefield.typing.chemistry import environment as env
from openeye import oechem
# ## Default Chemical Environments
#
# All Chemical Environments can be initated using SMIRKS strings.
# If a `ChemicalEnvironment` is initiated with no SMIRKS pattern, it is an empty structure.
# However, there are 5 subtypes of `ChemicalEnvironments` that match the types of parameters found in the SMIRNOFF format.
# If they are initiated with no SMIRKS pattern, their structure matches a generic for that parameter type, for example `[*:1]~[*:2]` for a bond (that is any atom connected to any other atom by any bond).
# The 5 subtypes are listed below with their expected number of indexed atoms and the corresponding SMIRKS structure:
#
# * `AtomChemicalEnvironment`
# - expects 1 indexed atom
# - default/generic SMIRKS `"[*:1]"`
# * `BondChemicalEnvironment`
# - expects 2 indexed atoms
# - default/generic SMIRKS: `"[*:1]~[*:2]"`
# * `AngleChemicalEnvironment`
# - expects 3 indexed atoms
# - default/generic SMIRKS: `"[*:1]~[*:2]~[*:3]"`
# * `TorsionChemicalEnvironment`
# - expects 4 indexed atoms in a proper dihedral angle
# - default/generic SMIRKS: `"[*:1]~[*:2]~[*:3]~[*:4]"`
# * `ImproperChemicalEnvironment`
# - expects 4 indexed atoms in an improper dihedral angle
# - default/generic SMIRKS: `"[*:1]~[*:2](~[*:3])~[*:4]"`
#
# Here we show how these are initiated. Note that the generic environment is blank, it has the potential to become a SMIRKS pattern, but currently nothing is stored in it. While the subtypes have the shape described above, but wildcards (`'*'` for atoms and `'~'` for bonds).
# +
# NBVAL_SKIP
Env = env.ChemicalEnvironment()
atomEnv = env.AtomChemicalEnvironment()
bondEnv = env.BondChemicalEnvironment()
angleEnv = env.AngleChemicalEnvironment()
torsionEnv = env.TorsionChemicalEnvironment()
impropEnv = env.ImproperChemicalEnvironment()
EnvList = [Env, atomEnv, bondEnv, angleEnv, torsionEnv, impropEnv]
names = ['generic', 'Atom','Bond','Angle','Torsion','(Improper']
for idx, Env in enumerate(EnvList):
print("%10s: %s" % (names[idx], Env.asSMIRKS()))
# -
# ## Initiating `ChemicalEnvironments` from SMIRKS Strings
#
# `ChemicalEnvironments` can be initialized by SMIRKS strings. Here we attempt to show the robustness of this parsing. These patterns are intentionally complicated and therefore can be hard to read by humans. Here are some of the key features we would like to test:
#
# * SMILES strings are SMIRKS strings (i.e. 'CCC' should be stored as 3 atoms bonded in a row).
# * Replacement strings, such as `"$ewg1"` to mean `"[#7!-1,#8!-1,#16!-1,#9,#17,#35,#53]"`
# * Complex recursive SMIRKS such as `"[#6$(*([#6]=[#8])-,=$ewg2))]"`
# * Ring indexing, as in SMILES, SMARTS and SMIKRS use a number after an atom to describe the atoms in a ring, such as "[#6:1]1(-;!@[#1,#6])=;@[#6]-;@[#6]1" to show a cyclopropene ring where atom 1 is in the double bond and is bound to a hydrogen or carbon outside the ring.
# * Hybrid SMIRKS with atomic symbols for the atoms. These do not have to use the square brackets, for example "C(O-[#7,#8])C[C+0]=[*]"
#
# In this set-up we will show that these SMIRKS patterns are parseable with the OpenEye Toolkits, then create a `ChemicalEnvironment` from the SMIRKS string and then print the `ChemicalEnvironment` as a SMIRKS string. Note that due to the nature of SMIRKS patterns the `ChemicalEnvironment` smirks may not identically match the input SMIRKS. A key difference is that every atom in a `ChemicalEnvironment` SMIRKS will be inside square brackets. Also, "blank" bonds, for example in "CCC" will be converted to their literal meaning, single or aromatic.
# +
# NBVAL_SKIP
# define the two replacements strings
replacements = [ ('ewg1', '[#7!-1,#8!-1,#16!-1,#9,#17,#35,#53]'),
('ewg2', '[#7!-1,#8,#16]')]
# define complicated SMIRKS patterns
SMIRKS = ['[#6$(*~[#6]=[#8])$(*-,=$ewg2)]', # complex recursive SMIRKS
'CCC', # SMILES
"[#1:1]-CCC", # simple hybrid
'[#6:1]1(-;!@[#1,#6])=;@[#6]-;@[#6]1', # Complicated ring
'C(O-[#7,#8])CC=[*]', # Hybrid SMIRKS
"[#6$([#6X4](~[$ewg1])(~[#8]~[#1])):1]-[#6X2H2;+0:2]-,=,:;!@;!#[$ewg2:3]-[#4:4]", # its just long
"[#6$([#6X4](~[$ewg1])(~[#8]~[#1])):1]1=CCCC1", # another ring
]
for smirk in SMIRKS:
qmol = oechem.OEQMol()
tmp_smirks = oechem.OESmartsLexReplace(smirk, replacements)
parseable = env.OEParseSmarts(qmol, tmp_smirks)
print("Input SMIRKS: %s" % smirk)
print("\t parseable by OpenEye Tools: %s" % parseable)
Env = env.ChemicalEnvironment(smirks = smirk, replacements = replacements)
print("\t Chemical Environment SMIRKS: %s\n" % Env.asSMIRKS())
# -
# # Structure of `ChemicalEnvironments`
#
# Up until now, we have discussed only how to initiate `ChemicalEnvironment`s. Now we will explain how they are structured and how to use them to make changes to your SMIRKS pattern (and therefor the fragment you are describing).
# To begin with, the overall structure of `ChemicalEnvironment`s is similar to how a chemist might think about a fragment.
# We use NetworkX graphs to store information about the pieces.
# Nodes store information about Atoms and edges (connecting nodes) store information about Bonds.
# Both of these sub-structures, Atoms and Bonds store information about the input SMIRKS pattern in a broken down way so it can be easily editted. The words Atoms and Bonds are capitalized as they are classes in and of themselves.
#
# Both Atoms and Bonds have two types of information
# * ORtypes
# - things that are OR'd together in the SMIRKS string using a comma (',')
# - These have two subtypes:
# - ORbases - typically an atomic number
# - ORdecorators - typically information that might be true for 1 possible atomic number, but not others
# * ANDtypes
# - things that are AND'd together in the SMIRKS string using a semi-colon (';')
#
# This starts to sound complicated, so to try to illustrate how this works, we will use an actual Angle found in the [SMIRNOFF99Frosst](https://github.com/openforcefield/smirnoff99Frosst) forcefield.
#
# Here is the SMIRKS String:
#
# # `"[#6X3,#7:1]~;@[#8;r:2]~;@[#6X3,#7:3]"`
#
# * atom 1 and atom 3
# - ORtypes
# - '#6X3' - a trivalent carbon
# - ORbase = '#6'
# - ORdecorators = ['X3']
# - '#7' is a nitrogen
# - ORbase = '#7'
# - ORdecorators []
# - ANDtypes
# - [] (None)
# * atom 2
# - ORtypes
# - '#8'
# - ORbase = '#8'
# - ORdecorators = []
# - ANDtypes
# - ['r'] it is in a ring
# * bond 1 and 2 are identical
# - ORtypes = None (generic bond ~)
# - ANDtypes = ['@']
# - it is in a ring
#
# ### Selecting Atoms and Bonds
#
# Here we will use the selectAtom and selectBond functions to get a specific atom or bond and then print its information. The 'select' methods ( selectAtom() or selectBond() ) takes an argument descriptor which can be used to select a certain atom or type of atom.
#
# Descriptor input option:
# * None - returns a random atom
# * int - returns that atom or bond by index
# * 'Indexed' - returns a random indexed atom
# * 'Unindexed' - returns a random non-indexed atom
# * 'Alpha' - returns a random atom alpha to an indexed atom
# * 'Beta' - returns a random atom beta to an indexed atom
#
# +
smirks = "[#6X3,#7:1]~;@[#8;r:2]~;@[#6X3,#7:3]"
angle = env.ChemicalEnvironment(smirks = smirks)
# get atom1 and print information
atom1 = angle.selectAtom(1)
print("Atom 1: '%s'" % atom1.asSMIRKS())
print("ORTypes")
for (base, decs) in atom1.getORtypes():
print("\tBase: %s" % base)
str_decs = ["'%s'" % d for d in decs]
str_decs = ','.join(str_decs)
print("\tDecorators: [%s]" % str_decs)
print("ANDTypes:", atom1.getANDtypes())
print()
# get bond1 and print information
bond1 = angle.selectBond(1)
print("Bond 1: '%s'" % bond1.asSMIRKS())
print("ORTypes: ", bond1.getORtypes())
print("ANDTypes: ", bond1.getANDtypes())
# -
# ## Changing ORtypes and ANDtypes
#
# For both ORtypes and ANDtypes for Atoms and Bonds there are "get" and "set" methods.
# The set methods completely rewrite that type.
# There are also methods for add ORtypes and ANDtypes where you add a single entry to the existing list.
#
# Here we will use the set ORtypes to change atom1 to be a trivalent carbon or a divalent nitrogen.
#
# Then we will also add an ORType and ANDType to atom2 so that it could refer to an oxygen ('#8') or trivalent and neutral nitrogen ('#7X3+0') and in one ring ('R1').
#
#
# ### Final SMIRKS string: `"[#6X3,#7X2:1]~;@[#8,#7X3+0;r;R1:2]~;@[#6X3,#7:3]"`
# +
# Change atom1's ORtypes with the setORtype method
new_ORtypes = [ ('#6', ['X3']), ('#7', ['X2']) ]
atom1.setORtypes(new_ORtypes)
print("New Atom 1: %s " % atom1.asSMIRKS())
# Change atom2's AND and OR types with the add*type methods
atom2 = angle.selectAtom(2)
atom2.addANDtype('R1')
atom2.addORtype('#7', ['X3', '+0'])
print("New Atom 2: %s" % atom2.asSMIRKS())
print("\nNew SMIRKS: %s" % angle.asSMIRKS())
# -
# ## Adding new Atoms
#
# The addAtom method is used to introduce atoms bound to existing atoms.
# You can add an empty atom or specify information about the new bond and new atom.
# Here are the parameters for the addAtom method:
# ```
# Parameters
# -----------
# bondToAtom: atom object, required
# atom the new atom will be bound to
# bondORtypes: list of tuples, optional
# strings that will be used for the ORtypes for the new bond
# bondANDtypes: list of strings, optional
# strings that will be used for the ANDtypes for the new bond
# newORtypes: list of strings, optional
# strings that will be used for the ORtypes for the new atom
# newANDtypes: list of strings, optional
# strings that will be used for the ANDtypes for the new atom
# newAtomIndex: int, optional
# integer label that could be used to index the atom in a SMIRKS string
# beyondBeta: boolean, optional
# if True, allows bonding beyond beta position
# ```
#
# The `addAtom` method returns the created atom.
#
# Here we will add an alpha atom (oxygen) to atom 3 that is not in a ring and then a beta atom (hydrogen) bound to the alpha atom.
#
# ### New SMIRKS pattern: `"[#6X3,#7X2:1]~;@[#8,#7+0X3;R1:2]~;@[#6X3,#7:3]~;!@[#8X2H1;R0]~[#1]"`
#
# +
atom3 = angle.selectAtom(3)
alpha_ORtypes = [('#8', ['X2', 'H1'])]
alpha_ANDtypes = ['R0']
alpha_bondANDtypes = ['!@']
beta_ORtypes = [('#1', [])]
alpha = angle.addAtom(atom3, bondANDtypes = alpha_bondANDtypes,
newORtypes = alpha_ORtypes, newANDtypes = alpha_ANDtypes)
beta = angle.addAtom(alpha, newORtypes = beta_ORtypes)
print("Alpha Atom SMIRKS: %s" % alpha.asSMIRKS())
print("Beta Atom SMIRKS: %s" % beta.asSMIRKS())
print()
print("New overall SMIRKS: %s" % angle.asSMIRKS())
# -
# ## Removing Atoms
#
# The removeAtom method works how you would expect. It removes the specified atom and the bond connecting it to the fragment.
# You cannot remove indexed atoms (if you want to remove their OR and AND decorators you can set them to empty lists).
# The other option with the `removeAtom` method is to say only remove it if the atom is undecorated. This is done by setting the input variable `isEmpty` to True (default is False). When `isEmpty` is True, the atom is only removed if it has 1 ORtype and no ANDtypes.
#
# The `removeAtom` method returns True if the atom was removed and False if it was not.
#
# As an example, we will remove the hydrogen in the beta position to atom3 that was added above.
#
# ### New SMIRKS pattern: `"New overall SMIRKS: [#6X3,#7X2:1]~;@[#8,#7+0X3;R1:2]~;@[#6X3,#7:3]~;!@[#8X2H1;R0]"`
removed = angle.removeAtom(beta)
print("The hydrogen beta to atom3 was remove: ", removed)
print("Updated SMIRKS string: %s" % angle.asSMIRKS())
# ## Other `ChemicalEnvironment` Methods
#
# There are a variety of other methods that let you get information about the stored fragment. This includes:
#
# 1. Getting information about an atom or bond in an environment (i.e. `isAlpha` returns a boolean)
# * Get atoms or bonds in each type of position:
# - `getAtoms` or `getBonds`
# - returns all atoms or bonds
# - `getIndexedAtoms` or `getIndexedBonds`
# - `getAlphaAtoms` or `getAlphaBonds`
# - `getBetaAtoms` or `getBetaBonds`
# - `getUnindexedAtoms` or `getUnindexedBonds`
# * Report the minimum order of a bond with `Bond.getOrder`
# - Note this is the minimum so a bond that is single or double (`'-,='`) will report the order as 1
# * Report the valence and bond order around an atom can be reported with `getValence` and `getBondORder`
# * Get a bond between two atoms (or determine if the atoms are bonded) with `getBond(atom1, atom2)`
# * Get atoms bound to a specified atom with `getNeighbors`
#
# Here we will show how each of these method types is used:
# 1. Getting information about an atom or bond in an environment (i.e. isAlpha returns a boolean)
# Check if the alpha atom above is any of the following
print("Above a carbon atom ('%s') was added in the alpha position to atom 3. This atom is ..." % alpha.asSMIRKS())
print("\t Indexed: ", angle.isIndexed(alpha))
print("\t Unindexed: ", angle.isUnindexed(alpha))
print("\t Alpha: ", angle.isAlpha(alpha))
print("\t Beta: ", angle.isBeta(alpha))
# NOTE - These methods can take an atom or a bond as an argument
# +
# 2. Get atoms or bonds in each type of position, for example getIndexedAtoms or getAlphaBonds
# We will print the SMIRKS for each indexed atom:
indexed = angle.getIndexedAtoms()
print("Here are the SMIRKS strings for the Indexed atoms in the example angle:")
for a in indexed:
print("\tAtom %i: '%s'" % (a.index, a.asSMIRKS()))
print()
bonds = angle.getBonds()
print("Here are the SMIRKS strings for ALL bonds in the example angle:")
for b in bonds:
print("\t'%s'" % b.asSMIRKS())
# -
# 3. Report the minimum order of a bond with Bond.getOrder
bond1 = angle.selectBond(1)
print("Bond 1 (between atoms 1 and 2) has a minimum order of %i" % bond1.getOrder())
# 4. Report the valence and bond order around an atom can be reported with getValence and getBondORder
atom3 = angle.selectAtom(3)
print("Atom 3 has a valency of %i" % angle.getValence(atom3))
print("Atom 3 has a minimum bond order of %i" % angle.getBondOrder(atom3))
# 5. Get a bond between two atoms (or determine if the atoms are bonded) with getBond(atom1, atom2)
# Check for bonds between each pair of indexed atoms
atom_pairs = [ (1,2), (2,3), (1,3) ]
for (A,B) in atom_pairs:
atomA = angle.selectAtom(A)
atomB = angle.selectAtom(B)
# check if there is a bond between the two atoms
bond = angle.getBond(atomA, atomB)
if bond is None:
print("There is no bond between Atom %i and Atom %i" % (A, B))
else:
print("The bond between Atom %i and Atom %i has the pattern '%s'" % (A, B, bond.asSMIRKS()))
# 6. Get atoms bound to a specified atom with getNeighbors
# get the neighbors for each indexed atom
for A in [1,2,3]:
atomA = angle.selectAtom(A)
print("Atom %i has the following neighbors" % A)
for a in angle.getNeighbors(atomA):
print("\t '%s' " % a.asSMIRKS())
print()
| examples/deprecated/chemicalEnvironments/using_environments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Coding Exercise #0307
# ### 1. Correlation:
import pandas as pd
import numpy as np
import scipy.stats as st
import os
# !wget --no-clobber https://raw.githubusercontent.com/stefannae/SIC-Artificial-Intelligence/main/SIC_AI_Coding_Exercises/SIC_AI_Chapter_04_Coding_Exercises/data_iris.csv
# Read in the data.
df = pd.read_csv('data_iris.csv', header='infer')
df.shape
df.head(3)
# Set aside two variables.
x = df['Petal.Length']
y = df['Sepal.Length']
# #### 1.1. Pearson:
# Using the SciPy function.
# Correlation and p-value.
np.round(st.pearsonr(x,y),3)
# Using the Pandas function.
x.corr(y)
# Correlation array.
np.round(df.corr(),3)
# #### 1.2. Spearman:
# Using the SciPy function.
# Correlation and p-value.
np.round(st.spearmanr(x,y),3)
# #### 1.3. Kendall:
# Using the SciPy function.
# Correlation and p-value.
np.round(st.kendalltau(x,y),3) # 상관계수와 p-값.
# ### 2. Interval Estimation of the Correlation:
# !wget --no-clobber https://raw.githubusercontent.com/stefannae/SIC-Artificial-Intelligence/main/SIC_AI_Coding_Exercises/SIC_AI_Chapter_04_Coding_Exercises/data_studentlist.csv
# Read in another data set.
df = pd.read_csv('data_studentlist.csv', header='infer')
df.head(3)
# Set aside two variables.
x = df.height
y = df.weight
# #### 2.2. Confidence Interval of the Pearson Correlation:
# Apply the Fisher's z-transformation.
# See the lecture.
n = len(x)
r = x.corr(y)
z = np.arctanh(r)
std_error_z = 1/np.sqrt(n-3)
# 95% confidence interval.
# Expressed as a dictionary object.
{'low':np.tanh(z-st.norm.ppf(0.975)*std_error_z), 'high': np.tanh(z+st.norm.ppf(0.975)*std_error_z)}
# 99% confidence interval.
# Expressed as a dictionary object.
{'low':np.tanh(z-st.norm.ppf(0.995)*std_error_z), 'high': np.tanh(z+st.norm.ppf(0.995)*std_error_z)}
| SIC_AI_Coding_Exercises/SIC_AI_Chapter_04_Coding_Exercises/ex_0307.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
"""
Copyright (c) 2016, <NAME>
All rights reserved.
Redistribution and use in source and binary forms, with or without
modification, are permitted provided that the following conditions are met:
1. Redistributions of source code must retain the above copyright notice, this
list of conditions and the following disclaimer.
2. Redistributions in binary form must reproduce the above copyright notice,
this list of conditions and the following disclaimer in the documentation
and/or other materials provided with the distribution.
3. Neither the name of the copyright holder nor the names of its contributors
may be used to endorse or promote products derived from this software
without specific prior written permission.
THIS SOFTWARE IS PROVIDED BY THE COPYRIGHT HOLDERS AND CONTRIBUTORS "AS IS"
AND ANY EXPRESS OR IMPLIED WARRANTIES, INCLUDING, BUT NOT LIMITED TO, THE
IMPLIED WARRANTIES OF MERCHANTABILITY AND FITNESS FOR A PARTICULAR PURPOSE
ARE DISCLAIMED. IN NO EVENT SHALL THE COPYRIGHT HOLDER OR CONTRIBUTORS BE
LIABLE FOR ANY DIRECT, INDIRECT, INCIDENTAL, SPECIAL, EXEMPLARY, OR
CONSEQUENTIAL DAMAGES (INCLUDING, BUT NOT LIMITED TO, PROCUREMENT OF
SUBSTITUTE GOODS OR SERVICES; LOSS OF USE, DATA, OR PROFITS; OR BUSINESS
INTERRUPTION) HOWEVER CAUSED AND ON ANY THEORY OF LIABILITY, WHETHER IN
CONTRACT, STRICT LIABILITY, OR TORT (INCLUDING NEGLIGENCE OR OTHERWISE)
ARISING IN ANY WAY OUT OF THE USE OF THIS SOFTWARE, EVEN IF ADVISED OF THE
POSSIBILITY OF SUCH DAMAGE.
"""
import os
import numpy as np
import matplotlib.pyplot as plt
plt.rcdefaults()
from matplotlib.lines import Line2D
from matplotlib.patches import Rectangle
from matplotlib.collections import PatchCollection
NumConvMax = 8
NumFcMax = 20
White = 1.
Light = 0.7
Medium = 0.5
Dark = 0.3
Black = 0.
def add_layer(patches, colors, size=24, num=5,
top_left=[0, 0],
loc_diff=[3, -3],
):
# add a rectangle
top_left = np.array(top_left)
loc_diff = np.array(loc_diff)
loc_start = top_left - np.array([0, size])
for ind in range(num):
patches.append(Rectangle(loc_start + ind * loc_diff, size, size))
if ind % 2:
colors.append(Medium)
else:
colors.append(Light)
def add_mapping(patches, colors, start_ratio, patch_size, ind_bgn,
top_left_list, loc_diff_list, num_show_list, size_list):
start_loc = top_left_list[ind_bgn] \
+ (num_show_list[ind_bgn] - 1) * np.array(loc_diff_list[ind_bgn]) \
+ np.array([start_ratio[0] * size_list[ind_bgn],
-start_ratio[1] * size_list[ind_bgn]])
end_loc = top_left_list[ind_bgn + 1] \
+ (num_show_list[ind_bgn + 1] - 1) \
* np.array(loc_diff_list[ind_bgn + 1]) \
+ np.array([(start_ratio[0] + .5 * patch_size / size_list[ind_bgn]) *
size_list[ind_bgn + 1],
-(start_ratio[1] - .5 * patch_size / size_list[ind_bgn]) *
size_list[ind_bgn + 1]])
patches.append(Rectangle(start_loc, patch_size, patch_size))
colors.append(Dark)
patches.append(Line2D([start_loc[0], end_loc[0]],
[start_loc[1], end_loc[1]]))
colors.append(Black)
patches.append(Line2D([start_loc[0] + patch_size, end_loc[0]],
[start_loc[1], end_loc[1]]))
colors.append(Black)
patches.append(Line2D([start_loc[0], end_loc[0]],
[start_loc[1] + patch_size, end_loc[1]]))
colors.append(Black)
patches.append(Line2D([start_loc[0] + patch_size, end_loc[0]],
[start_loc[1] + patch_size, end_loc[1]]))
colors.append(Black)
def label(xy, text, xy_off=[0, 4]):
plt.text(xy[0] + xy_off[0], xy[1] + xy_off[1], text,
family='sans-serif', size=8)
if __name__ == '__main__':
fc_unit_size = 2
layer_width = 40
patches = []
colors = []
fig, ax = plt.subplots()
############################
# conv layers
size_list = [32, 18, 10, 6, 4]
num_list = [3, 32, 32, 48, 48]
x_diff_list = [0, layer_width, layer_width, layer_width, layer_width]
text_list = ['Inputs'] + ['Feature\nmaps'] * (len(size_list) - 1)
loc_diff_list = [[3, -3]] * len(size_list)
num_show_list = map(min, num_list, [NumConvMax] * len(num_list))
top_left_list = np.c_[np.cumsum(x_diff_list), np.zeros(len(x_diff_list))]
for ind in range(len(size_list)):
add_layer(patches, colors, size=size_list[ind],
num=num_show_list[ind],
top_left=top_left_list[ind], loc_diff=loc_diff_list[ind])
label(top_left_list[ind], text_list[ind] + '\n{}@{}x{}'.format(
num_list[ind], size_list[ind], size_list[ind]))
############################
# in between layers
start_ratio_list = [[0.4, 0.5], [0.4, 0.8], [0.4, 0.5], [0.4, 0.8]]
patch_size_list = [5, 2, 5, 2]
ind_bgn_list = range(len(patch_size_list))
text_list = ['Convolution', 'Max-pooling', 'Convolution', 'Max-pooling']
for ind in range(len(patch_size_list)):
add_mapping(patches, colors, start_ratio_list[ind],
patch_size_list[ind], ind,
top_left_list, loc_diff_list, num_show_list, size_list)
label(top_left_list[ind], text_list[ind] + '\n{}x{} kernel'.format(
patch_size_list[ind], patch_size_list[ind]), xy_off=[26, -65])
############################
# fully connected layers
size_list = [fc_unit_size, fc_unit_size, fc_unit_size]
num_list = [768, 500, 2]
num_show_list = map(min, num_list, [NumFcMax] * len(num_list))
x_diff_list = [sum(x_diff_list) + layer_width, layer_width, layer_width]
top_left_list = np.c_[np.cumsum(x_diff_list), np.zeros(len(x_diff_list))]
loc_diff_list = [[fc_unit_size, -fc_unit_size]] * len(top_left_list)
text_list = ['Hidden\nunits'] * (len(size_list) - 1) + ['Outputs']
for ind in range(len(size_list)):
add_layer(patches, colors, size=size_list[ind], num=num_show_list[ind],
top_left=top_left_list[ind], loc_diff=loc_diff_list[ind])
label(top_left_list[ind], text_list[ind] + '\n{}'.format(
num_list[ind]))
text_list = ['Flatten\n', 'Fully\nconnected', 'Fully\nconnected']
for ind in range(len(size_list)):
label(top_left_list[ind], text_list[ind], xy_off=[-10, -65])
############################
colors += [0, 1]
collection = PatchCollection(patches, cmap=plt.cm.gray)
collection.set_array(np.array(colors))
ax.add_collection(collection)
plt.tight_layout()
plt.axis('equal')
plt.axis('off')
plt.show()
fig.set_size_inches(8, 2.5)
fig_dir = './'
fig_ext = '.png'
fig.savefig(os.path.join(fig_dir, 'convnet_fig' + fig_ext),
bbox_inches='tight', pad_inches=0)
# -
| Data Preprocessing/.ipynb_checkpoints/plot-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# INF6804 Vision par ordinateur
#
# Polytechnique Montréal
#
# Exemple du calcul d'un axe par analyse en composantes principales (ACP)
from skimage import data
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.lines as mlines
# Lecture ou création d'une image binaire.
#Image 1
#image = np.invert(data.horse())
#Image 2
image = np.zeros((100,100))
image[35:85,20:40] = 1
#Affichage
plt.figure(figsize=(4,4))
plt.imshow(image, plt.get_cmap('binary'))
plt.show()
# Listes contenant les coordonnées en X et en Y des pixels à True.
LesX = []
LesY = []
for i in range(image.shape[0]):
for j in range(image.shape[1]):
if image[i,j] == True:
LesX = LesX + [j]
LesY = LesY + [i]
# Moyennes et covariance des coordonnées pour calculer les vecteurs propres, valeurs propres.
MoyX = np.mean(LesX)
MoyY = np.mean(LesY)
LesX = LesX - MoyX
LesY = LesY - MoyY
matcov = np.cov(LesX,LesY)
# Calcul des vecteurs propres/valeurs propres pour trouver les axes principaux. Trouve le système de coordonnées optimal pour représenter les pixels à True.
val, vec = np.linalg.eig(matcov)
maxval = np.argmax(val)
# Affichage de l'axe ayant le plus d'énergie.
# +
plt.figure(figsize=(4,4))
plt.imshow(image, plt.get_cmap('binary'))
plt.plot(MoyX,MoyY, marker='o', color = 'r')
ax = plt.gca()
xmin = MoyX - 100*vec[maxval,0]
xmax = MoyX + 100*vec[maxval,0]
ymin = MoyY - 100*vec[maxval,1]
ymax = MoyY + 100*vec[maxval,1]
l = mlines.Line2D([xmin,xmax], [ymin,ymax])
ax.add_line(l)
plt.show()
# -
| PCAAxis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import csv
import pandas as pd
# +
# import graph data
def load_data(csvname):
# load in data
data = np.asarray(pd.read_csv(csvname))
# import data and reshape appropriately
X = data[:,0:-1]
y = data[:,-1]
y.shape = (len(y),1)
#print('Test MAE :',np.amin(X, axis=0))
print('Eval MAE :',np.amin(y, axis=0))
return X,y
def make_graph(variable):
train, evaluation = load_data(variable)
plt.plot(train, linestyle='solid',label='training set')
plt.plot(evaluation, linestyle='solid',label='evaluation set')
plt.xlabel("Epoch")
plt.ylabel("MAE")
plt.legend();
plt.show()
# -
make_graph('resultplain17_1.csv')
make_graph('resultsrnet17_1.csv')
make_graph('resultirnet17_1.csv')
make_graph('resultplain24_1.csv')
make_graph('resultsrnet24_1.csv')
make_graph('resultirnet24_1.csv')
make_graph('resultplain48_2.csv')
make_graph('resultsrnet48_1.csv')
make_graph('resultirnet48_1.csv')
| Graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 (''.rssv_model'': venv)'
# language: python
# name: python3
# ---
# The behavior of the risky asset, with a constant interest rate $r$, is described by
# $$dS_t = rS_tdt + \sigma(X_t)dWt$$
# where $X$ is a continuous time markov chain and $W$ is a standard brownian motion.
#
# To simulate this, we first instanciate a continuous time markov with, for example, 3 states and provide 3 values for sigma.
# +
from stochastic_process.markov_chain import Sigma, MarkovChain
sigma = Sigma(0.2, 0.5, 0.8)
X = MarkovChain(5, 3, 7, 2, 4, 9)
# -
# The combination fo this two allow us to compute $\sigma$'s paths.
# +
import numpy as np
import matplotlib.pyplot as plt
# Compute X_paths. Here we just compute one
delta_t = 0.01
random_sample_for_X = np.random.rand(1, 100)
X_paths = X.paths(random_sample_for_X, delta_t)
X_path = X_paths[0]
# Map X state's values to sigma's state values
sigma_paths = sigma.paths(X_paths)
sigma_path = sigma_paths[0]
plt.plot(sigma_path)
# -
# We can now create a risky asset built on this two objects and a specified interest rate $r$.
# +
from stochastic_process.risky_asset import RiskyAsset
r = 0.05
S_0 = 1
S = RiskyAsset(current_price=S_0)
S_paths = S.paths(delta_t, sigma_paths, r)
S_path = S_paths[0]
plt.plot(S_path)
# -
# Considering the two process $\sigma_t$ and $S_t$, one can see how $\sigma$'s values impact the behavior of S:
# +
fig=plt.figure(figsize=(16, 8))
S_ax = fig.add_subplot(121)
sigma_ax = fig.add_subplot(122)
S_ax.plot(S_path)
sigma_ax.plot(sigma_path)
| risky_asset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import xarray as xr
from tqdm import tqdm
from RPLB_acc_LC import RPLB_acc_LC
# set pure spectral properties
lambda_0 = 800e-9 # central wavelength
tau_0 = 10e-15 # FL pulse duration 1/e2 intensity half-width (same def as beam waist)
# set initial beam waist
w_0 = 4e-6
# calculate Rayleigh range
z_R = (np.pi*w_0**2)/(lambda_0)
# beam power
P = 100e12
# spectral phase
GDD = 0*(1e-15)**2
TOD = 0*(1e-15)**3
# intial electron velocity
beta_0 = 0
# set tuning parameters
z_0 = np.linspace(-0.2*z_R, +0.2*z_R, 9, endpoint=True)
Psi_0 = np.linspace(0, 2*np.pi, 14, endpoint=False)
tau_p = np.linspace(-3, +1, 9)*(1e-15)
# create result variable
KE_final = np.zeros(shape=(len(z_0), len(Psi_0), len(tau_p)))
#loop over inital position of test particle
for i in range(0, len(z_0)):
# loop over CEO phase of laser
for j in range(0, len(Psi_0)):
# loop over tau_p
for k in range(0, len(tau_p)):
KE_final[i, j, k] = RPLB_acc_LC(lambda_0, tau_0, w_0, P, Psi_0[j], GDD, TOD, z_0[i], beta_0, tau_p[k])
# +
KE_final = xr.DataArray(KE_final, coords=[z_0/z_R, Psi_0, tau_p], dims=['z_0', 'CEO', 'tau_p'])
plt.figure(figsize=(5, 5))
(KE_final.max(dim='z_0').max(dim='CEO')/1e6).plot()
plt.tight_layout()
# -
| LC_OnAxis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
N = int(input())
if N%2 == 0 :
print("even")
N = int(input())
if N%2 == 0 :
print("even")
else :
print("odd")
N = int(input())
if N > 0 :
print("positive")
elif N == 0 :
print("zero")
else :
print("negative")
a = int(input())
if a > 0 :
print("positive")
print("%d is positive number." %a)
print("Indentation is very important in Python.")
a = int(input())
if a > 0 :
print("positive")
print("%d is positive number." %a)
print("Indentation is very important in Python.")
a = int(input())
if a > 0 :
if a%2 == 0 :
print(a, "Positive even number")
else :
print(a, "Positive odd number")
elif a == 0 :
print(a, "Zero")
else :
if a%2 == 0 : print(a, "Negative even number")
else : print(a, "Negative odd number")
| 04 if~else (stu).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Object Detection
#
# **Authors:**
#
# - [<NAME>](https://github.com/AngusTheMack) ([1106817](mailto:<EMAIL>))
# - [<NAME>](https://github.com/nmichlo) ([1386161](mailto:<EMAIL>))
#
# **Achievement** Detecting the bounding box of snakes within images to better perform classification later on.
#
# ## Introduction
# This notebook is based off of techniques from [this article](https://medium.com/@Stormblessed/2460292bcfb) and [this repo](https://github.com/GokulEpiphany/contests-final-code/tree/master/aicrowd-snake-species) by .
#
# ----------------------
# +
# Utilities
import sys
import os
from tqdm.notebook import tqdm
import imageio
import matplotlib.pyplot as plt
from fastai.vision.data import ImageList
import json
from pprint import pprint
# Add root of project to PYTHON_PATH so we can import correctly
if os.path.abspath('../') not in {os.path.abspath(path) for path in sys.path}:
sys.path.insert(0, os.path.abspath('../'))
# Import SSIC common stuffs
from ssic.ssic import SSIC
from ssic.util import set_random_seed, cache_data
# if you dont have a .env file set it here
os.environ.setdefault('DATASET_DIR', '~/downloads/datasets/ssic')
# Initialise SSIC paths, data and other stuffs, searches for a .env file in the project with these variables specified, also checkpoints os.environ and sys.path
SSIC.init()
# -
# ## User Setup
#
# **DATASETS FROM**: https://medium.com/@Stormblessed/2460292bcfb
#
# **INSTRUCTIONS**:
# Download both of the following into the `DATASET_DIR` above, then extract the dataset into that same directory **take care** all the images are not inside a folder within the zip.
# - labeled dataset: https://drive.google.com/file/d/1q14CtkQ9r7rlxwLuksWAOduhDjUb-bBE/view
# - drive link: https://drive.google.com/file/d/18dx_5Ngmc56fDRZ6YZA_elX-0ehtV5U6/view
# ## Code
# +
# LOAD IMAGES:
IMAGES_DIR = os.path.join(SSIC.DATASET_DIR, 'train-object-detect')
assert os.path.isdir(IMAGES_DIR)
imagelist = ImageList.from_folder(IMAGES_DIR)
# LOAD ANNOTATIONS:
ANNOTATIONS_PATH = os.path.join(SSIC.DATASET_DIR, 'annotations.json')
assert os.path.isfile(ANNOTATIONS_PATH)
with open(ANNOTATIONS_PATH, 'r') as file:
ANNOTATIONS = json.load(file)
# -
# Show One Example
pprint(ANNOTATIONS[0])
plt.imshow(imageio.imread(SSIC.get_train_image_info()[ANNOTATIONS[0]['filename']]['path']))
# +
class SnakeDetector(nn.Module):
def __init__(self, arch=models.resnet18):
super().__init__()
self.cnn = create_body(arch)
self.head = create_head(num_features_model(self.cnn) * 2, 4)
def forward(self, im):
x = self.cnn(im)
x = self.head(x)
return 2 * (x.sigmoid_() - 0.5)
def loss_fn(preds, targs, class_idxs):
return L1Loss()(preds, targs.squeeze())
# -
| notebooks/object_detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
import torchvision
import math
# import model
from model import CenterNet,focal_loss
# -
use_cuda = True
device = torch.device('cuda:4' if use_cuda else 'cpu')
x = torch.Tensor([0]).cuda(device)
centerNet = CenterNet(3)
centerNet.to(device)
centerNet.mode
x = torch.randn(5,3,512,512)
target = []
for i in range(x.shape[0]):
target.append(
dict(
classes = torch.Tensor([1,2,1,0,1]),
bboxes = torch.FloatTensor([[10,70,250,345],
[100,45,124,145],
[178,54,230,310],
[187,49,400,310],
[145,214,340,440]])
)
)
# + tags=[]
x = x.cuda(device)
# + tags=[]
# %%time
for i in range(100):
result,losses = centerNet(x,target)
# -
x.is_cuda
fl = focal_loss(2,4,'mean',5)
fl(losses['heatmap'],losses['heatmap'])
torch.stack()
| Object Detection/CenterNet/train_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="DlJAbvFqOedG"
# # Mount Drive
# + colab={"base_uri": "https://localhost:8080/"} id="4-7JU3Q7q_XT" outputId="6cc74863-3650-45b5-a8a7-a16e40f9f1b7"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/"} id="DqqXI7Izk65p" outputId="be7f4527-ee34-4eba-e492-e567fc4c6e3e"
# !pip install -U -q PyDrive
# !pip install httplib2==0.15.0
import os
from pydrive.auth import GoogleAuth
from pydrive.drive import GoogleDrive
from pydrive.files import GoogleDriveFileList
from google.colab import auth
from oauth2client.client import GoogleCredentials
from getpass import getpass
import urllib
# 1. Authenticate and create the PyDrive client.
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
# Cloning PAL_2021 to access modules.
# Need password to access private repo.
if 'CLIPPER' not in os.listdir():
cmd_string = 'git clone https://github.com/PAL-ML/CLIPPER.git'
os.system(cmd_string)
# + [markdown] id="RGlVnbugxFK2"
# # Installation
# + [markdown] id="gmgIrfT8hDNE"
# ## Install multi label metrics dependencies
# + colab={"base_uri": "https://localhost:8080/"} id="b6xXPAFbe6Gp" outputId="2338673d-52e6-4a0a-8050-3c5d4b533406"
# ! pip install scikit-learn==0.24
# + [markdown] id="3rKe3HqM523g"
# ## Install CLIP dependencies
# + colab={"base_uri": "https://localhost:8080/"} id="poS-WNDixIhY" outputId="83414f05-96a0-423c-b7fb-c4e1cf2b9a5c"
import subprocess
CUDA_version = [s for s in subprocess.check_output(["nvcc", "--version"]).decode("UTF-8").split(", ") if s.startswith("release")][0].split(" ")[-1]
print("CUDA version:", CUDA_version)
if CUDA_version == "10.0":
torch_version_suffix = "+cu100"
elif CUDA_version == "10.1":
torch_version_suffix = "+cu101"
elif CUDA_version == "10.2":
torch_version_suffix = ""
else:
torch_version_suffix = "+cu110"
# + colab={"base_uri": "https://localhost:8080/"} id="uA-69W8M59nA" outputId="cd8bb8c6-2d82-4443-c686-d470d637d0bf"
# ! pip install torch==1.7.1{torch_version_suffix} torchvision==0.8.2{torch_version_suffix} -f https://download.pytorch.org/whl/torch_stable.html ftfy regex
# + colab={"base_uri": "https://localhost:8080/"} id="sYwBZS1N6A3d" outputId="4634e20a-4a58-4ff3-bb86-98883dbc65c9"
# ! pip install ftfy regex
# ! wget https://openaipublic.azureedge.net/clip/bpe_simple_vocab_16e6.txt.gz -O bpe_simple_vocab_16e6.txt.gz
# + colab={"base_uri": "https://localhost:8080/"} id="9oIcNBYB8lz3" outputId="41b99918-abb5-44bf-af4c-fcad3a83b504"
# !pip install git+https://github.com/Sri-vatsa/CLIP # using this fork because of visualization capabilities
# + [markdown] id="OLU-gp7n8__E"
# ## Install clustering dependencies
# + id="6TLg9ozo9Hvc"
# !pip -q install umap-learn>=0.3.7
# + [markdown] id="9z1WQnXdLHy2"
# ## Install dataset manager dependencies
# + colab={"base_uri": "https://localhost:8080/"} id="J1vvMx7_LLSp" outputId="25d70286-fd26-45a8-c3c1-f329e5b7a24b"
# !pip install wget
# + [markdown] id="NzsubsEm72rr"
# # Imports
# + id="KZI62a6G74kw"
# ML Libraries
import tensorflow as tf
import tensorflow_hub as hub
import torch
import torch.nn as nn
import torchvision.models as models
import torchvision.transforms as transforms
import keras
# Data processing
import PIL
import base64
import imageio
import pandas as pd
import numpy as np
import json
from PIL import Image
import cv2
from sklearn.feature_extraction.image import extract_patches_2d
# Plotting
import seaborn as sns
import matplotlib.pyplot as plt
import matplotlib.patches as patches
from IPython.core.display import display, HTML
from matplotlib import cm
import matplotlib.image as mpimg
# Models
import clip
# Datasets
import tensorflow_datasets as tfds
# Clustering
# import umap
from sklearn import metrics
from sklearn.cluster import KMeans
#from yellowbrick.cluster import KElbowVisualizer
# Misc
import progressbar
import logging
from abc import ABC, abstractmethod
import time
import urllib.request
import os
from sklearn.metrics import jaccard_score, hamming_loss, accuracy_score, f1_score
from sklearn.preprocessing import MultiLabelBinarizer
# Modules
from CLIPPER.code.ExperimentModules import embedding_models
from CLIPPER.code.ExperimentModules.dataset_manager import DatasetManager
from CLIPPER.code.ExperimentModules.weight_imprinting_classifier import WeightImprintingClassifier
from CLIPPER.code.ExperimentModules import simclr_data_augmentations
from CLIPPER.code.ExperimentModules.utils import (save_npy, load_npy,
get_folder_id,
create_expt_dir,
save_to_drive,
load_all_from_drive_folder,
download_file_by_name,
delete_file_by_name)
logging.getLogger('googleapicliet.discovery_cache').setLevel(logging.ERROR)
# + [markdown] id="zU_gxQ0KbwMh"
# # Initialization & Constants
# + [markdown] id="txBvu3lU-EVO"
# ## Dataset details
# + colab={"base_uri": "https://localhost:8080/"} id="bih5tBPdbx3u" outputId="fca1c741-bb65-476d-aad2-39935cd90fd3"
dataset_name = 'CelebAAttributes'
folder_name = "CelebAAttributes-Embeddings-22-03-21"
# Change parentid to match that of experiments root folder in gdrive
parentid = '1bK72W-Um20EQDEyChNhNJthUNbmoSEjD'
# Filepaths
train_labels_filename = "train_labels.npz"
val_labels_filename = "val_labels.npz"
test_labels_filename = "test_labels.npz"
train_embeddings_filename_suffix = "_embeddings_train.npz"
val_embeddings_filename_suffix = "_embeddings_val.npz"
test_embeddings_filename_suffix = "_embeddings_test.npz"
# Initialize sepcific experiment folder in drive
folderid = create_expt_dir(drive, parentid, folder_name)
# + [markdown] id="7adeWKpB-Gdk"
# ## Threshold range
#
#
# + id="alpQcgdL-BTd"
min_threshold = 0.44
max_threshold = 0.72
threshold_stepsize = 0.01
# + [markdown] id="dTMseBlA-xyO"
# ## Few shot learning parameters
# + id="ks-9cfhX-wvz"
num_ways = 5 # [5, 20]
num_shot = 5 # [5, 1]
num_eval = 15 # [5, 10, 15, 19]
num_episodes = 100
shuffle = False
# + [markdown] id="A2EubiVL_gRG"
# ## Image embedding and augmentations
# + id="wgkPidFD_frf"
embedding_model = embedding_models.CLIPEmbeddingWrapper()
num_augmentations = 0 # [0, 5, 10]
trivial=False # [True, False]
# + [markdown] id="DpVmoi5pAW4K"
# ## Training parameters
# + id="2oki_jMyAXg-"
# List of number of epochs to train over, e.g. [5, 10, 15, 20]. [0] indicates no training.
train_epochs_arr = [0]
# + [markdown] id="UxTa8MVsvQCN"
# # Load data
# + colab={"base_uri": "https://localhost:8080/"} id="n6S124Jfwuu5" outputId="84170e6f-d514-44ac-f38e-ebc55448c898"
def get_ndarray_from_drive(drive, folderid, filename):
download_file_by_name(drive, folderid, filename)
return np.load(filename)['data']
test_labels = get_ndarray_from_drive(drive, folderid, test_labels_filename)
# + colab={"base_uri": "https://localhost:8080/"} id="4p2JpaGHmqbl" outputId="6f419b8e-c78b-478e-fcc3-811ef75ebf5a"
dm = DatasetManager()
test_data_generator = dm.load_dataset('celeba_faces', split='test')
class_names = dm.get_class_names()
print(class_names)
# + [markdown] id="iLbRqaYxzbr7"
# # Create label dictionary
# + colab={"base_uri": "https://localhost:8080/"} id="Emz85fNX0Vif" outputId="1735562d-9032-4f49-b5e6-2b4b1511bab8"
unique_labels = np.arange(test_labels.shape[1])
print(len(unique_labels))
# + id="0TubS-RLzeVM"
label_dictionary = {la:[] for la in unique_labels}
for i in range(len(test_labels)):
la = np.where(test_labels[i] == 1)
for l in la[0]:
label_dictionary[l].append(i)
# + [markdown] id="xP6ftkDCvKul"
# # Weight Imprinting models on train data embeddings
# + [markdown] id="fJIGo_R86GQi"
# ## Function definitions
# + id="bXj6BCFkNp3R"
def get_item_from_dataset(idx):
dm = DatasetManager()
test_data_generator = dm.load_dataset('celeba_faces', split='test')
item = next(itertools.islice(test_data_generator, idx, None))
return item
# + id="pv68q8TH6tvG"
def start_progress_bar(bar_len):
widgets = [
' [',
progressbar.Timer(format= 'elapsed time: %(elapsed)s'),
'] ',
progressbar.Bar('*'),' (',
progressbar.ETA(), ') ',
]
pbar = progressbar.ProgressBar(
max_value=bar_len, widgets=widgets
).start()
return pbar
# + id="DtoopzLRStTO"
def prepare_indices(
num_ways,
num_shot,
num_eval,
num_episodes,
label_dictionary,
labels,
shuffle=False
):
eval_indices = []
train_indices = []
wi_y = []
eval_y = []
label_dictionary = {la:label_dictionary[la] for la in label_dictionary if len(label_dictionary[la]) >= (num_shot+num_eval)}
unique_labels = list(label_dictionary.keys())
pbar = start_progress_bar(num_episodes)
for s in range(num_episodes):
# Setting random seed for replicability
np.random.seed(s)
_train_indices = []
_eval_indices = []
selected_labels = np.random.choice(unique_labels, size=num_ways, replace=False)
for la in selected_labels:
la_indices = label_dictionary[la]
# select = np.random.choice(la_indices, size = num_shot+num_eval, replace=False)
# tr_idx = list(select[:num_shot])
# ev_idx = list(select[num_shot:])
tr_idx = []
ev_idx = []
while len(tr_idx) < num_shot:
idx = np.random.choice(la_indices)
if idx not in _train_indices and idx not in _eval_indices and idx not in tr_idx:
tr_idx.append(idx)
while len(ev_idx) < num_eval:
idx = np.random.choice(la_indices)
if idx not in _train_indices and idx not in _eval_indices and idx not in tr_idx and idx not in ev_idx:
ev_idx.append(idx)
_train_indices = _train_indices + tr_idx
_eval_indices = _eval_indices + ev_idx
if shuffle:
np.random.shuffle(_train_indices)
np.random.shuffle(_eval_indices)
train_indices.append(_train_indices)
eval_indices.append(_eval_indices)
_wi_y = []
for idx in _train_indices:
la = np.where(labels[idx] == 1)
_wi_y.append(list([l for l in la[0] if l in selected_labels]))
_eval_y = []
for idx in _eval_indices:
la = np.where(labels[idx] == 1)
_eval_y.append(list([l for l in la[0] if l in selected_labels]))
wi_y.append(_wi_y)
eval_y.append(_eval_y)
pbar.update(s+1)
return train_indices, eval_indices, wi_y, eval_y
# + id="4M5aHABX7k8a"
def embed_images(
embedding_model,
train_indices,
num_augmentations,
trivial=False
):
def augment_image(image, num_augmentations, trivial):
""" Perform SimCLR augmentations on the image
"""
if np.max(image) > 1:
image = image/255
augmented_images = [image]
def _run_filters(image):
width = image.shape[1]
height = image.shape[0]
image_aug = simclr_data_augmentations.random_crop_with_resize(
image,
height,
width
)
image_aug = tf.image.random_flip_left_right(image_aug)
image_aug = simclr_data_augmentations.random_color_jitter(image_aug)
image_aug = simclr_data_augmentations.random_blur(
image_aug,
height,
width
)
image_aug = tf.reshape(image_aug, [image.shape[0], image.shape[1], 3])
image_aug = tf.clip_by_value(image_aug, 0., 1.)
return image_aug.numpy()
for _ in range(num_augmentations):
if trivial:
aug_image = image
else:
aug_image = _run_filters(image)
augmented_images.append(aug_image)
augmented_images = np.stack(augmented_images)
return augmented_images
embedding_model.load_model()
unique_indices = np.unique(np.array(train_indices))
ds = dm.load_dataset('celeba_faces', split='test')
embeddings = []
IMAGE_IDX = 0
pbar = start_progress_bar(unique_indices.size+1)
num_done=0
for idx, item in enumerate(ds):
if idx in unique_indices:
image = item[IMAGE_IDX]
if num_augmentations > 0:
aug_images = augment_image(image, num_augmentations, trivial)
else:
aug_images = image
processed_images = embedding_model.preprocess_data(aug_images)
embedding = embedding_model.embed_images(processed_images)
embeddings.append(embedding)
num_done += 1
pbar.update(num_done+1)
if idx == unique_indices[-1]:
break
embeddings = np.stack(embeddings)
return unique_indices, embeddings
# + id="GZYucgmROjkm"
def train_model_for_episode(
indices_and_embeddings,
train_indices,
wi_y,
num_augmentations,
train_epochs=None,
train_batch_size=5,
multi_label=True
):
train_embeddings = []
train_labels = []
ind = indices_and_embeddings[0]
emb = indices_and_embeddings[1]
for idx, tr_idx in enumerate(train_indices):
train_embeddings.append(emb[np.argwhere(ind==tr_idx)[0][0]])
train_labels += [wi_y[idx] for _ in range(num_augmentations+1)]
train_embeddings = np.concatenate(train_embeddings)
train_embeddings = WeightImprintingClassifier.preprocess_input(train_embeddings)
wi_weights, label_mapping = WeightImprintingClassifier.get_imprinting_weights(
train_embeddings, train_labels, False, multi_label
)
wi_parameters = {
"num_classes": num_ways,
"input_dims": train_embeddings.shape[-1],
"scale": False,
"dense_layer_weights": wi_weights,
"multi_label": multi_label
}
wi_cls = WeightImprintingClassifier(wi_parameters)
if train_epochs:
# ep_y = train_labels
rev_label_mapping = {label_mapping[val]:val for val in label_mapping}
train_y = np.zeros((len(train_labels), num_ways))
for idx_y, _y in enumerate(train_labels):
for l in _y:
train_y[idx_y, rev_label_mapping[l]] = 1
wi_cls.train(train_embeddings, train_y, train_epochs, train_batch_size)
return wi_cls, label_mapping
# + id="MNkdJHdaOjkn"
def evaluate_model_for_episode(
model,
eval_x,
eval_y,
label_mapping,
metrics=['hamming', 'jaccard', 'subset_accuracy', 'ap', 'map', 'c_f1', 'o_f1', 'c_precision', 'o_precision', 'c_recall', 'o_recall', 'classwise_accuracy', 'c_accuracy'],
threshold=0.7,
multi_label=True
):
eval_x = WeightImprintingClassifier.preprocess_input(eval_x)
logits = model.predict_scores(eval_x).tolist()
if multi_label:
pred_y = model.predict_multi_label(eval_x, threshold)
pred_y = [[label_mapping[v] for v in l] for l in pred_y]
met = model.evaluate_multi_label_metrics(
eval_x, eval_y, label_mapping, threshold, metrics
)
else:
pred_y = model.predict_single_label(eval_x)
pred_y = [label_mapping[l] for l in pred_y]
met = model.evaluate_single_label_metrics(
eval_x, eval_y, label_mapping, metrics
)
return pred_y, met, logits
# + id="_u2vlkKxOjkn"
def run_episode_through_model(
indices_and_embeddings,
train_indices,
eval_indices,
wi_y,
eval_y,
thresholds=None,
num_augmentations=0,
train_epochs=None,
train_batch_size=5,
metrics=['hamming', 'jaccard', 'subset_accuracy', 'ap', 'map', 'c_f1', 'o_f1', 'c_precision', 'o_precision', 'c_recall', 'o_recall', 'classwise_accuracy', 'c_accuracy'],
embeddings=None,
multi_label=True
):
metrics_values = {m:[] for m in metrics}
wi_cls, label_mapping = train_model_for_episode(
indices_and_embeddings,
train_indices,
wi_y,
num_augmentations,
train_epochs,
train_batch_size,
multi_label=multi_label
)
eval_x = embeddings[eval_indices]
ep_logits = []
if multi_label:
for t in thresholds:
pred_labels, met, logits = evaluate_model_for_episode(
wi_cls,
eval_x,
eval_y,
label_mapping,
threshold=t,
metrics=metrics,
multi_label=True
)
ep_logits.append(logits)
for m in metrics:
metrics_values[m].append(met[m])
else:
pred_labels, metrics_values, logits = evaluate_model_for_episode(
wi_cls,
eval_x,
eval_y,
label_mapping,
metrics=['accuracy', 'f1_score'],
multi_label=False
)
ep_logits = logits
return metrics_values, ep_logits
# + id="tz9O1nAUOjkp"
def run_evaluations(
indices_and_embeddings,
train_indices,
eval_indices,
wi_y,
eval_y,
num_episodes,
num_ways,
thresholds,
verbose=True,
normalize=True,
train_epochs=None,
train_batch_size=5,
metrics=['hamming', 'jaccard', 'subset_accuracy', 'ap', 'map', 'c_f1', 'o_f1', 'c_precision', 'o_precision', 'c_recall', 'o_recall', 'classwise_accuracy', 'c_accuracy'],
embeddings=None,
num_augmentations=0,
multi_label=True
):
metrics_values = {m:[] for m in metrics}
all_logits = []
if verbose:
pbar = start_progress_bar(num_episodes)
for idx_ep in range(num_episodes):
_train_indices = train_indices[idx_ep]
_eval_indices = eval_indices[idx_ep]
_wi_y = wi_y[idx_ep]
_eval_y = eval_y[idx_ep]
met, ep_logits = run_episode_through_model(
indices_and_embeddings,
_train_indices,
_eval_indices,
_wi_y,
_eval_y,
num_augmentations=num_augmentations,
train_epochs=train_epochs,
train_batch_size=train_batch_size,
embeddings=embeddings,
thresholds=thresholds,
metrics=metrics,
multi_label=multi_label
)
all_logits.append(ep_logits)
for m in metrics:
metrics_values[m].append(met[m])
if verbose:
pbar.update(idx_ep+1)
return metrics_values, all_logits
# + id="MzmivM_CtSbA"
def get_max_mean_jaccard_index_by_threshold(metrics_thresholds):
max_mean_jaccard = np.max([np.mean(mt['jaccard']) for mt in metrics_thresholds])
return max_mean_jaccard
# + id="UtLfeX4w8tEZ"
def get_max_mean_jaccard_index_with_threshold(metrics_thresholds):
max_mean_jaccard = np.max([np.mean(mt['jaccard']) for mt in metrics_thresholds])
threshold = np.argmax([np.mean(mt['jaccard']) for mt in metrics_thresholds])
return max_mean_jaccard, threshold
# + id="wsKTx1h6tSbL"
def get_mean_max_jaccard_index_by_episode(metrics_thresholds):
mean_max_jaccard = np.mean(np.max(np.array([mt['jaccard'] for mt in metrics_thresholds]), axis=0))
return mean_max_jaccard
# + id="Y1mgLWl5N0hY"
def plot_metrics_by_threshold(
metrics_thresholds,
thresholds,
metrics=['hamming', 'jaccard', 'subset_accuracy', 'ap', 'map', 'c_f1', 'o_f1', 'c_precision', 'o_precision', 'c_recall', 'o_recall', 'classwise_accuracy', 'c_accuracy'],
title_suffix=""
):
legend = []
fig = plt.figure(figsize=(10,10))
if 'jaccard' in metrics:
mean_jaccard_threshold = np.mean(np.array(metrics_thresholds['jaccard']), axis=0)
opt_threshold_jaccard = thresholds[np.argmax(mean_jaccard_threshold)]
plt.plot(thresholds, mean_jaccard_threshold, c='blue')
plt.axvline(opt_threshold_jaccard, ls="--", c='blue')
legend.append("Jaccard Index")
legend.append(opt_threshold_jaccard)
if 'hamming' in metrics:
mean_hamming_threshold = np.mean(np.array(metrics_thresholds['hamming']), axis=0)
opt_threshold_hamming = thresholds[np.argmin(mean_hamming_threshold)]
plt.plot(thresholds, mean_hamming_threshold, c='green')
plt.axvline(opt_threshold_hamming, ls="--", c='green')
legend.append("Hamming Score")
legend.append(opt_threshold_hamming)
if 'map' in metrics:
mean_f1_score_threshold = np.mean(np.array(metrics_thresholds['map']), axis=0)
opt_threshold_f1_score = thresholds[np.argmax(mean_f1_score_threshold)]
plt.plot(thresholds, mean_f1_score_threshold, c='red')
plt.axvline(opt_threshold_f1_score, ls="--", c='red')
legend.append("mAP")
legend.append(opt_threshold_f1_score)
if 'o_f1' in metrics:
mean_f1_score_threshold = np.mean(np.array(metrics_thresholds['o_f1']), axis=0)
opt_threshold_f1_score = thresholds[np.argmax(mean_f1_score_threshold)]
plt.plot(thresholds, mean_f1_score_threshold, c='yellow')
plt.axvline(opt_threshold_f1_score, ls="--", c='yellow')
legend.append("OF1")
legend.append(opt_threshold_f1_score)
if 'c_f1' in metrics:
mean_f1_score_threshold = np.mean(np.array(metrics_thresholds['c_f1']), axis=0)
opt_threshold_f1_score = thresholds[np.argmax(mean_f1_score_threshold)]
plt.plot(thresholds, mean_f1_score_threshold, c='orange')
plt.axvline(opt_threshold_f1_score, ls="--", c='orange')
legend.append("CF1")
legend.append(opt_threshold_f1_score)
if 'o_precision' in metrics:
mean_f1_score_threshold = np.mean(np.array(metrics_thresholds['o_precision']), axis=0)
opt_threshold_f1_score = thresholds[np.argmax(mean_f1_score_threshold)]
plt.plot(thresholds, mean_f1_score_threshold, c='purple')
plt.axvline(opt_threshold_f1_score, ls="--", c='purple')
legend.append("OP")
legend.append(opt_threshold_f1_score)
if 'c_precision' in metrics:
mean_f1_score_threshold = np.mean(np.array(metrics_thresholds['c_precision']), axis=0)
opt_threshold_f1_score = thresholds[np.argmax(mean_f1_score_threshold)]
plt.plot(thresholds, mean_f1_score_threshold, c='cyan')
plt.axvline(opt_threshold_f1_score, ls="--", c='cyan')
legend.append("CP")
legend.append(opt_threshold_f1_score)
if 'o_recall' in metrics:
mean_f1_score_threshold = np.mean(np.array(metrics_thresholds['o_recall']), axis=0)
opt_threshold_f1_score = thresholds[np.argmax(mean_f1_score_threshold)]
plt.plot(thresholds, mean_f1_score_threshold, c='brown')
plt.axvline(opt_threshold_f1_score, ls="--", c='brown')
legend.append("OR")
legend.append(opt_threshold_f1_score)
if 'c_recall' in metrics:
mean_f1_score_threshold = np.mean(np.array(metrics_thresholds['c_recall']), axis=0)
opt_threshold_f1_score = thresholds[np.argmax(mean_f1_score_threshold)]
plt.plot(thresholds, mean_f1_score_threshold, c='pink')
plt.axvline(opt_threshold_f1_score, ls="--", c='pink')
legend.append("CR")
legend.append(opt_threshold_f1_score)
if 'c_accuracy' in metrics:
mean_f1_score_threshold = np.mean(np.array(metrics_thresholds['c_accuracy']), axis=0)
opt_threshold_f1_score = thresholds[np.argmax(mean_f1_score_threshold)]
plt.plot(thresholds, mean_f1_score_threshold, c='maroon')
plt.axvline(opt_threshold_f1_score, ls="--", c='maroon')
legend.append("CACC")
legend.append(opt_threshold_f1_score)
plt.xlabel('Threshold')
plt.ylabel('Value')
plt.legend(legend)
title = title_suffix+"\nMulti label metrics by threshold"
plt.title(title)
plt.grid()
fname = os.path.join(PLOT_DIR, title_suffix)
plt.savefig(fname)
plt.show()
# + [markdown] id="hHs46yaOItTW"
# ## Setting multiple thresholds
# + colab={"base_uri": "https://localhost:8080/"} id="vlo2bBCAIyQj" outputId="594699d3-6a14-4a28-945c-9213125092f8"
thresholds = np.arange(min_threshold, max_threshold, threshold_stepsize)
thresholds
# + [markdown] id="g8CNr6I4rU5d"
# # Main
# + [markdown] id="g73bz0lqrU5e"
# ## Picking indices
# + colab={"base_uri": "https://localhost:8080/"} id="Fm6j-e0brU5f" outputId="8f206163-1f37-4f0d-d184-8512ba8c3a94"
train_indices, eval_indices, wi_y, eval_y = prepare_indices(
num_ways, num_shot, num_eval, num_episodes, label_dictionary, test_labels, shuffle
)
# + colab={"base_uri": "https://localhost:8080/"} id="UEg7SCafPpmz" outputId="11c1ddde-6f41-401f-b8c5-b3407f93a4fa"
indices, embeddings = embed_images(
embedding_model,
train_indices,
num_augmentations,
trivial=trivial
)
# + [markdown] id="lLW-8vrbrU5f"
# ## CLIP
# + id="uK5qZA3mrU5g" colab={"base_uri": "https://localhost:8080/"} outputId="33a27c07-fe07-473a-a402-e20509800c00"
clip_embeddings_test_fn = "clip" + test_embeddings_filename_suffix
clip_embeddings_test = get_ndarray_from_drive(drive, folderid, clip_embeddings_test_fn)
# + id="XGTJsU6yN0S0" colab={"base_uri": "https://localhost:8080/"} outputId="cd4b9f61-08e5-4d30-d448-ac09c5663937"
if train_epochs_arr == [0]:
if trivial:
results_filename = "new_metrics"+dataset_name+"_0t"+str(num_ways)+"w"+str(num_shot)+"s"+str(num_augmentations)+"a_trivial_metrics_with_logits.json"
else:
results_filename = "new_metrics"+dataset_name+"_0t"+str(num_ways)+"w"+str(num_shot)+"s"+str(num_augmentations)+"a_metrics_with_logits.json"
else:
if trivial:
results_filename = "new_metrics"+dataset_name+"_"+str(num_ways)+"w"+str(num_shot)+"s"+str(num_augmentations)+"a_trivial_metrics_with_logits.json"
else:
results_filename = "new_metrics"+dataset_name+"_"+str(num_ways)+"w"+str(num_shot)+"s"+str(num_augmentations)+"a_metrics_with_logits.json"
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
download_file_by_name(drive, folderid, results_filename)
if results_filename in os.listdir():
with open(results_filename, 'r') as f:
json_loaded = json.load(f)
clip_metrics_over_train_epochs = json_loaded['metrics']
logits_over_train_epochs = json_loaded["logits"]
else:
clip_metrics_over_train_epochs = []
logits_over_train_epochs = []
for idx, train_epochs in enumerate (train_epochs_arr):
if idx < len(clip_metrics_over_train_epochs):
continue
print(idx)
clip_metrics_thresholds, all_logits = run_evaluations(
(indices, embeddings),
train_indices,
eval_indices,
wi_y,
eval_y,
num_episodes,
num_ways,
thresholds,
train_epochs=train_epochs,
num_augmentations=num_augmentations,
embeddings=clip_embeddings_test
)
clip_metrics_over_train_epochs.append(clip_metrics_thresholds)
logits_over_train_epochs.append(all_logits)
fin_list = []
for a1 in wi_y:
fin_a1_list = []
for a2 in a1:
fin_a2_list = []
for a3 in a2:
new_val = str(a3)
fin_a2_list.append(new_val)
fin_a1_list.append(fin_a2_list)
fin_list.append(fin_a1_list)
with open(results_filename, 'w') as f:
results = {'metrics': clip_metrics_over_train_epochs,
"logits": logits_over_train_epochs,
"true_labels": fin_list}
json.dump(results, f)
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
delete_file_by_name(drive, folderid, results_filename)
save_to_drive(drive, folderid, results_filename)
# + id="7EkbqgHn4ubD"
def get_best_metric_and_threshold(mt, metric_name, thresholds, optimal='max'):
if optimal=='max':
opt_value = np.max(np.mean(np.array(mt[metric_name]), axis=0))
opt_threshold = thresholds[np.argmax(np.mean(np.array(mt[metric_name]), axis=0))]
if optimal=='min':
opt_value = np.min(np.mean(np.array(mt[metric_name]), axis=0))
opt_threshold = thresholds[np.argmin(np.mean(np.array(mt[metric_name]), axis=0))]
return opt_value, opt_threshold
# + id="gdQCn9JgQ6it" colab={"base_uri": "https://localhost:8080/"} outputId="ed37f895-04ea-4fc8-cc7a-e1c3ebdf9737"
all_metrics = ['hamming', 'jaccard', 'subset_accuracy', 'map', 'c_f1', 'o_f1', 'c_precision', 'o_precision', 'c_recall', 'o_recall', 'c_accuracy']
f1_vals = []
f1_t_vals = []
jaccard_vals = []
jaccard_t_vals = []
final_dict = {}
for ind_metric in all_metrics:
vals = []
t_vals = []
final_array = []
for mt in clip_metrics_over_train_epochs:
if ind_metric == "hamming":
ret_val, ret_t_val = get_best_metric_and_threshold(mt,ind_metric,thresholds,"min")
else:
ret_val, ret_t_val = get_best_metric_and_threshold(mt,ind_metric,thresholds,"max")
vals.append(ret_val)
t_vals.append(ret_t_val)
final_array.append(vals)
final_array.append(t_vals)
final_dict[ind_metric] = final_array
if train_epochs_arr == [0]:
if trivial:
graph_filename = "new_metrics"+dataset_name+"_0t"+str(num_ways)+"w"+str(num_shot)+"s"+str(num_augmentations)+"a_trivial_metrics_graphs.json"
else:
graph_filename = "new_metrics"+dataset_name+"_0t"+str(num_ways)+"w"+str(num_shot)+"s"+str(num_augmentations)+"a_metrics_graphs.json"
else:
if trivial:
graph_filename = "new_metrics"+dataset_name+"_"+str(num_ways)+"w"+str(num_shot)+"s"+str(num_augmentations)+"a_trivial_metrics_graphs.json"
else:
graph_filename = "new_metrics"+dataset_name+"_"+str(num_ways)+"w"+str(num_shot)+"s"+str(num_augmentations)+"a_metrics_graphs.json"
with open(graph_filename, 'w') as f:
json.dump(final_dict, f)
auth.authenticate_user()
gauth = GoogleAuth()
gauth.credentials = GoogleCredentials.get_application_default()
drive = GoogleDrive(gauth)
delete_file_by_name(drive, folderid, graph_filename)
save_to_drive(drive, folderid, graph_filename)
# + id="TPASq7usIXzn"
if trivial:
PLOT_DIR = "NewMetrics_WeightImprintingSigmoid_MultiPred_Train_Augmentations_CelebAAttributes" + str(num_ways) + "w" + str(num_shot) + "s" + str(num_augmentations) + "a_trivial_plots"
else:
PLOT_DIR = "NewMetrics_WeightImprintingSigmoid_MultiPred_Train_Augmentations_CelebAAttributes" + str(num_ways) + "w" + str(num_shot) + "s" + str(num_augmentations) + "a_plots"
os.mkdir(PLOT_DIR)
# + colab={"base_uri": "https://localhost:8080/", "height": 637} id="Sah_ZklUCZ9v" outputId="26a9964c-a243-43d8-f99f-b464e812b0e4"
for mt in clip_metrics_over_train_epochs:
plot_metrics_by_threshold(mt, thresholds, metrics=all_metrics)
| notebooks/WeightImprintingSigmoid/WeightImprintingSigmoid_CelebAAttributes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "4"}
# ## 1. The Statcast revolution
# <p><img style="float: left;margin:5px 20px 5px 1px" src="https://s3.amazonaws.com/assets.datacamp.com/production/project_250/img/judge_wide.jpg"></p>
# <p>This is <NAME>. Judge is one of the physically largest players in Major League Baseball standing 6 feet 7 inches (2.01 m) tall and weighing 282 pounds (128 kg). He also hit the <a href="https://www.mlb.com/news/aaron-judge-sets-statcast-exit-velocity-record/c-235640846">hardest home run</a> ever recorded. How do we know this? <strong>Statcast</strong>.</p>
# <p>Statcast is a state-of-the-art tracking system that uses high-resolution cameras and radar equipment to measure the precise location and movement of baseballs and baseball players. Introduced in 2015 to all 30 major league ballparks, Statcast data is revolutionizing the game. Teams are engaging in an "arms race" of data analysis, hiring analysts left and right in an attempt to gain an edge over their competition. This <a href="https://www.youtube.com/watch?v=9rOKGKhQe8U">video</a> describing the system is incredible.</p>
# <p><strong>In this notebook</strong>, we're going to wrangle, analyze, and visualize Statcast data to compare Mr. Judge and another (extremely large) teammate of his. Let's start by loading the data into our Notebook. There are two CSV files, <code>judge.csv</code> and <code>stanton.csv</code>, both of which contain Statcast data for 2015-2017. We'll use pandas DataFrames to store this data. Let's also load our data visualization libraries, matplotlib and seaborn.</p>
# + tags=["sample_code"] dc={"key": "4"}
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
# Load <NAME>'s Statcast data
judge = pd.read_csv('datasets/judge.csv')
# Load <NAME>'s Statcast data
stanton = pd.read_csv('datasets/stanton.csv')
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "11"}
# ## 2. What can Statcast measure?
# <p>The better question might be, what can't Statcast measure?</p>
# <blockquote>
# <p>Starting with the pitcher, Statcast can measure simple data points such as velocity. At the same time, Statcast digs a whole lot deeper, also measuring the release point and spin rate of every pitch.</p>
# <p>Moving on to hitters, Statcast is capable of measuring the exit velocity, launch angle and vector of the ball as it comes off the bat. From there, Statcast can also track the hang time and projected distance that a ball travels.</p>
# </blockquote>
# <p>Let's inspect the last five rows of the <code>judge</code> DataFrame. You'll see that each row represents one pitch thrown to a batter. You'll also see that some columns have esoteric names. If these don't make sense now, don't worry. The relevant ones will be explained as necessary.</p>
# + tags=["sample_code"] dc={"key": "11"}
# Display all columns (pandas will collapse some columns if we don't set this option)
pd.set_option('display.max_columns', None)
# Display the last five rows of the Aaron Judge file
# ... YOUR CODE FOR TASK 2 ...
judge.tail()
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "18"}
# ## 3. <NAME> and <NAME>, prolific sluggers
# <p><img style="float: left;margin:5px 20px 5px 1px" src="https://s3.amazonaws.com/assets.datacamp.com/production/project_250/img/stanton_wide.jpg"></p>
# <p>This is <NAME>. He is also a very large human being, standing 6 feet 6 inches tall and weighing 245 pounds. Despite not wearing the same jersey as Judge in the pictures provided, in 2018 they will be teammates on the New York Yankees. They are similar in a lot of ways, one being that they hit a lot of home runs. Stanton and Judge led baseball in home runs in 2017, with <a href="https://www.youtube.com/watch?v=tJ6Bz5P6dg4">59</a> and <a href="https://www.youtube.com/watch?v=Gw3pFHMM9fk">52</a>, respectively. These are exceptional totals - the player in third "only" had 45 home runs.</p>
# <p>Stanton and Judge are also different in many ways. One is <a href="http://m.mlb.com/glossary/statcast/batted-ball-event">batted ball events</a>, which is any batted ball that produces a result. This includes outs, hits, and errors. Next, you'll find the counts of batted ball events for each player in 2017. The frequencies of other events are quite different.</p>
# + tags=["sample_code"] dc={"key": "18"}
# All of Aaron Judge's batted ball events in 2017
judge_events_2017 = judge[judge['game_date']>"2016-12-31"].events
print("<NAME> batted ball event totals, 2017:")
print(judge_events_2017.value_counts())
# All of Giancarlo Stanton's batted ball events in 2017
stanton_events_2017 = stanton[stanton['game_date']>"2016-12-31"].events
print("\nGiancarlo Stanton batted ball event totals, 2017:")
print(stanton_events_2017.value_counts())
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "25"}
# ## 4. Analyzing home runs with Statcast data
# <p>So Judge walks and strikes out more than Stanton. Stanton flies out more than Judge. But let's get into their hitting profiles in more detail. Two of the most groundbreaking Statcast metrics are launch angle and exit velocity:</p>
# <ul>
# <li><a href="http://m.mlb.com/glossary/statcast/launch-angle">Launch angle</a>: the vertical angle at which the ball leaves a player's bat</li>
# <li><a href="http://m.mlb.com/glossary/statcast/exit-velocity">Exit velocity</a>: the speed of the baseball as it comes off the bat</li>
# </ul>
# <p>This new data has changed the way teams value both hitters and pitchers. Why? As per the <a href="https://www.washingtonpost.com/graphics/sports/mlb-launch-angles-story/?utm_term=.8d088d31f098">Washington Post</a>:</p>
# <blockquote>
# <p>Balls hit with a high launch angle are more likely to result in a hit. Hit fast enough and at the right angle, they become home runs.</p>
# </blockquote>
# <p>Let's look at exit velocity vs. launch angle and let's focus on home runs only (2015-2017). The first two plots show data points. The second two show smoothed contours to represent density.</p>
# + tags=["sample_code"] dc={"key": "25"}
# Filter to include home runs only
judge_hr = judge[judge['events']=='home_run']
stanton_hr = stanton[stanton['events'] == 'home_run']
# Create a figure with two scatter plots of launch speed vs. launch angle, one for each player's home runs
fig1, axs1 = plt.subplots(ncols=2, sharex=True, sharey=True)
sns.regplot(x='launch_speed', y='launch_angle', fit_reg=False, color='tab:blue', data=judge_hr, ax=axs1[0]).set_title('Aaron Judge\nHome Runs, 2015-2017')
sns.regplot(x='launch_speed', y='launch_angle', fit_reg=False, color='tab:blue', data=stanton_hr, ax=axs1[1]).set_title('Giancarlo Stanton\nHome Runs, 2015-2017')
# Create a figure with two KDE plots of launch speed vs. launch angle, one for each player's home runs
fig2, axs2 = plt.subplots(ncols=2, sharex=True, sharey=True)
sns.kdeplot(judge_hr.launch_speed, judge_hr.launch_angle, cmap="Blues", shade=True, shade_lowest=False, ax=axs2[0]).set_title('Aaron Judge\nHome Runs, 2015-2017')
sns.kdeplot(stanton_hr.launch_speed, stanton_hr.launch_angle, cmap="Blues", shade=True, shade_lowest=False, ax=axs2[1]).set_title('Giancarlo Stanton\nHome Runs, 2015-2017')
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "32"}
# ## 5. Home runs by pitch velocity
# <p>It appears that Stanton hits his home runs slightly lower and slightly harder than Judge, though this needs to be taken with a grain of salt given the small sample size of home runs.</p>
# <p>Not only does Statcast measure the velocity of the ball coming off of the bat, it measures the velocity of the ball coming out of the pitcher's hand and begins its journey towards the plate. We can use this data to compare Stanton and Judge's home runs in terms of pitch velocity. Next you'll find box plots displaying the five-number summaries for each player: minimum, first quartile, median, third quartile, and maximum.</p>
# + tags=["sample_code"] dc={"key": "32"}
# Combine the Judge and Stanton home run DataFrames for easy boxplot plotting
judge_stanton_hr = pd.concat([judge_hr, stanton_hr])
# Create a boxplot that describes the pitch velocity of each player's home runs
sns.boxplot(x = 'release_speed', data = judge_stanton_hr, color = 'tab:blue').set_title('Home Runs, 2015-2017')
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "39"}
# ## 6. Home runs by pitch location (I)
# <p>So Judge appears to hit his home runs off of faster pitches than Stanton. We might call Judge a fastball hitter. Stanton appears agnostic to pitch speed and likely pitch movement since slower pitches (e.g. curveballs, sliders, and changeups) tend to have more break. Statcast <em>does</em> track pitch movement and type but let's move on to something else: <strong>pitch location</strong>. Statcast tracks the zone the pitch is in when it crosses the plate. The zone numbering looks like this (from the catcher's point of view):</p>
# <p><img style="margin:5px 20px 5px 1px; width:20%;" src="https://s3.amazonaws.com/assets.datacamp.com/production/project_250/img/zone.png"></p>
# <p>We can plot this using a 2D histogram. For simplicity, let's only look at strikes, which gives us a 9x9 grid. We can view each zone as coordinates on a 2D plot, the bottom left corner being (1,1) and the top right corner being (3,3). Let's set up a function to assign x-coordinates to each pitch.</p>
# + tags=["sample_code"] dc={"key": "39"}
def assign_x_coord(row):
"""
Assigns an x-coordinate to Statcast's strike zone numbers. Zones 11, 12, 13,
and 14 are ignored for plotting simplicity.
"""
# Left third of strike zone
if row.zone in [1, 4, 7]:
# ... YOUR CODE FOR TASK 6 ...
return(1)
# Middle third of strike zone
if row.zone in [2, 5, 8]:
# ... YOUR CODE FOR TASK 6 ...
return(2)
# Right third of strike zone
if row.zone in [3, 6, 9]:
# ... YOUR CODE FOR TASK 6 ...
return(3)
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "46"}
# ## 7. Home runs by pitch location (II)
# <p>And let's do the same but for y-coordinates.</p>
# + tags=["sample_code"] dc={"key": "46"}
def assign_y_coord(row):
"""
Assigns a y-coordinate to Statcast's strike zone numbers. Zones 11, 12, 13,
and 14 are ignored for plotting simplicity.
"""
# Upper third of strike zone
# ... YOUR CODE FOR TASK 7 ...
if row.zone in [1,2,3]:
return 3
# Middle third of strike zone
# ... YOUR CODE FOR TASK 7 ...
if row.zone in [4,5,6]:
return 2
# Lower third of strike zone
# ... YOUR CODE FOR TASK 7 ...
if row.zone in [7,8,9]:
return 1
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "53"}
# ## 8. <NAME>'s home run zone
# <p>Now we can apply the functions we've created then construct our 2D histograms. First, for <NAME> (again, for pitches in the strike zone that resulted in home runs).</p>
# + tags=["sample_code"] dc={"key": "53"}
# Zones 11, 12, 13, and 14 are to be ignored for plotting simplicity
judge_strike_hr = judge_hr.copy().loc[judge_hr.zone <= 9]
# Assign Cartesian coordinates to pitches in the strike zone for Judge home runs
judge_strike_hr['zone_x'] = judge_strike_hr.apply(assign_x_coord, axis = 1)
judge_strike_hr['zone_y'] = judge_strike_hr.apply(assign_y_coord, axis = 1)
# Plot Judge's home run zone as a 2D histogram with a colorbar
plt.hist2d(judge_strike_hr.zone_x, judge_strike_hr.zone_x, bins = 3, cmap='Blues')
plt.title('Aaron Judge Home Runs on\n Pitches in the Strike Zone, 2015-2017')
plt.gca().get_xaxis().set_visible(False)
plt.gca().get_yaxis().set_visible(False)
cb = plt.colorbar()
cb.set_label('Counts in Bin')
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "60"}
# ## 9. <NAME>on's home run zone
# <p>And now for <NAME>anton.</p>
# + tags=["sample_code"] dc={"key": "60"}
# Zones 11, 12, 13, and 14 are to be ignored for plotting simplicity
stanton_strike_hr = stanton_hr.copy().loc[stanton_hr.zone <= 9]
# Assign Cartesian coordinates to pitches in the strike zone for Stanton home runs
stanton_strike_hr['zone_x'] = stanton_strike_hr.apply(assign_x_coord, axis=1)
stanton_strike_hr['zone_y'] = stanton_strike_hr.apply(assign_y_coord, axis=1)
# Plot Stanton's home run zone as a 2D histogram with a colorbar
plt.hist2d(stanton_strike_hr['zone_x'], stanton_strike_hr['zone_y'], bins = 3, cmap='Blues')
plt.title('Giancarlo Stanton Home Runs on\n Pitches in the Strike Zone, 2015-2017')
plt.gca().get_xaxis().set_visible(False)
plt.gca().get_yaxis().set_visible(False)
cb = plt.colorbar()
cb.set_label('Counts in Bin')
# + tags=["context"] deletable=false editable=false run_control={"frozen": true} dc={"key": "67"}
# ## 10. Should opposing pitchers be scared?
# <p>A few takeaways:</p>
# <ul>
# <li>Stanton does not hit many home runs on pitches in the upper third of the strike zone.</li>
# <li>Like pretty much every hitter ever, both players love pitches in the horizontal and vertical middle of the plate.</li>
# <li>Judge's least favorite home run pitch appears to be high-away while Stanton's appears to be low-away.</li>
# <li>If we were to describe Stanton's home run zone, it'd be middle-inside. Judge's home run zone is much more spread out.</li>
# </ul>
# <p>The grand takeaway from this whole exercise: <NAME> and <NAME> are not identical despite their superficial similarities. In terms of home runs, their launch profiles, as well as their pitch speed and location preferences, are different.</p>
# <p>Should opposing pitchers still be scared?</p>
# + tags=["sample_code"] dc={"key": "67"}
# Should opposing pitchers be wary of <NAME> and <NAME>
should_pitchers_be_scared = True
| A New Era of Data Analysis in Baseball/notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# -*- coding: utf-8 -*-
from __future__ import unicode_literals
# text in Western (Windows 1252)
import pickle
import numpy as np
# import StringIO
import math
from keras import optimizers, metrics
from keras.models import Model
from keras.layers import Dense, Dropout, Input
from keras.layers.merge import concatenate
from keras import regularizers
from keras.layers.convolutional import Conv1D
from keras.layers.convolutional import MaxPooling1D
from keras.constraints import maxnorm
from keras.layers import Flatten
from keras.optimizers import SGD
from keras.models import load_model
# from keras import backend as Input
np.random.seed(7)
# +
# # %run ../../../prepare_data.py
import sys
sys.path.insert(0, '../../../')
from prepare_data import *
# -
# X_train, X_other_features_train, y_train, X_validate, X_other_features_validate, y_validate = generate_full_matrix_inputs()
# save_inputs('../../internal_representations/inputs/shuffeled_matrix_train_inputs_other_features_multilabel.h5', X_train, y_train, other_features = X_other_features_train)
# save_inputs('../../internal_representations/inputs/shuffeled_matrix_validate_inputs_other_features_multilabel.h5', X_validate, y_validate, other_features = X_other_features_validate)
X_train, X_other_features_train, y_train = load_inputs('../../internal_representations/inputs/shuffeled_matrix_train_inputs_other_features_multilabel.h5', other_features=True)
X_validate, X_other_features_validate, y_validate = load_inputs('../../internal_representations/inputs/shuffeled_matrix_validate_inputs_other_features_multilabel.h5', other_features=True)
# X_other_features = create_X_features(content)
# print (X_other_features[178200])
decode_position = 30
print (decode_input(X_train[decode_position], dictionary))
# print (X_other_features_train[0])
print (decode_X_features(feature_dictionary, [X_other_features_train[decode_position]]))
# print (len(X_other_features_train[0]))
# +
num_examples = len(X_train) # training set size
# nn_input_dim = max_word * len(dictionary) # input layer dimensionality
# nn_output_dim = max_num_vowels * max_num_vowels # output layer dimensionality
nn_output_dim = 11
nn_hdim = 516
batch_size = 16
actual_epoch = 10
num_fake_epoch = 20
# Gradient descent parameters (I picked these by hand)
# epsilon = 1 # learning rate for gradient descent
# reg_lambda = 1 # regularization strength
# +
# word_processor = Sequential()
# word_processor.add(Conv1D(43, (3), input_shape=(23, 43), padding='same', activation='relu'))
# word_processor.add(Conv1D(43, (2), padding='same', activation='relu'))
# word_processor.add(Conv1D(43, (2), padding='same', activation='relu'))
# word_processor.add(MaxPooling1D(pool_size=2))
# word_processor.add(Flatten())
# word_processor.add(Dense(516, activation='relu', kernel_constraint=maxnorm(3)))
# metadata_processor = Sequential()
# metadata_processor.add(Dense(256, input_dim=167, activation='relu'))
# model = Sequential()
# model.add(Merge([word_processor, metadata_processor], mode='concat')) # Merge is your sensor fusion buddy
# model.add(Dense(1024, input_dim=(516 + 256), activation='relu'))
# model.add(Dropout(0.3))
# model.add(Dense(1024, activation='relu'))
# model.add(Dropout(0.2))
# model.add(Dense(nn_output_dim, activation='sigmoid'))
# -------------------------------------------------------
conv_input_shape=(23, 43)
othr_input = (167, )
conv_input = Input(shape=conv_input_shape, name='conv_input')
x_conv = Conv1D(43, (3), padding='same', activation='relu')(conv_input)
x_conv = Conv1D(43, (3), padding='same', activation='relu')(x_conv)
x_conv = MaxPooling1D(pool_size=2)(x_conv)
x_conv = Flatten()(x_conv)
# x_conv = Dense(516, activation='relu', kernel_constraint=maxnorm(3))(x_conv)
othr_input = Input(shape=othr_input, name='othr_input')
# x_othr = Dense(256, input_dim=167, activation='relu')(othr_input)
# x_othr = Dropout(0.3)(x_othr)
# x_othr = Dense(256, activation='relu')(othr_input)
x = concatenate([x_conv, othr_input])
# x = Dense(1024, input_dim=(516 + 256), activation='relu')(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.3)(x)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.3)(x)
x = Dense(2048, activation='relu')(x)
x = Dropout(0.3)(x)
x = Dense(1024, activation='relu')(x)
x = Dropout(0.3)(x)
x = Dense(512, activation='relu')(x)
x = Dropout(0.2)(x)
x = Dense(nn_output_dim, activation='sigmoid')(x)
# model = Model(inputs=[conv_input, othr_input], output=x)
# +
model = Model(inputs=[conv_input, othr_input], outputs=x)
# epochs = 5
# lrate = 0.1
# decay = lrate/epochs
# sgd = SGD(lr=lrate, momentum=0.9, decay=decay, nesterov=False)
# model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
# Compile model
# keras.optimizers.Adam(lr=0.001, beta_1=0.9, beta_2=0.999, epsilon=1e-08, decay=0.0)
opt = optimizers.Adam(lr=1E-4, beta_1=0.9, beta_2=0.999, epsilon=1e-08)
model.compile(loss='binary_crossentropy', optimizer=opt, metrics=[actual_accuracy,])
# model.compile(loss='mean_squared_error', optimizer='adam', metrics=['accuracy'])
# +
history = model.fit_generator(generate_fake_epoch(X_train, X_other_features_train, y_train, batch_size), X_train.shape[0]/(batch_size * num_fake_epoch), epochs=actual_epoch*num_fake_epoch, validation_data=([X_validate, X_other_features_validate], y_validate))
# model.fit([X_train, X_other_features_train], y_train, validation_data=([X_validate, X_other_features_validate], y_validate), epochs=1, batch_size=16)
# -
name = '10_epoch'
model.save(name + '.h5')
output = open(name + '_history.pkl', 'wb')
pickle.dump(history.history, output)
output.close()
import matplotlib.pyplot as plt
plt.plot(history.history['actual_accuracy'])
plt.plot(history.history['val_actual_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize history for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
model = load_model('test.h5')
h5f.close()
# evaluate the model
scores = model.evaluate([X_validate, X_other_features_validate], y_validate)
print("\n%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
# +
pos = 0
print(decode_input(X_validate[pos], dictionary))
# predictions = model.predict([X_validate, X_other_features_validate])
predictions2 = model.predict([X_validate[pos:pos+1], X_other_features_validate[pos:pos+1]])
# print(predictions2)
print(np.round(predictions2)[0])
# predictions = np.round(predictions)
print(y_validate[pos])
# accuracy = sum([1 if np.all(y_validate[i] == predictions[i]) else 0 for i in range(X_validate.shape[0])])/float(X_validate.shape[0])
# print(accuracy)
import keras.backend as K
def act_accuracy(y_true, y_pred):
return K.mean(K.equal(K.mean(K.equal(K.round(y_true), K.round(y_pred)), axis=-1), 1.0))
def mean_pred2(y_true, y_pred):
return K.mean(K.equal(K.argmax(y_true, axis=-1), K.argmax(y_pred, axis=-1)))
# return K.mean(K.all(K.equal(y_true, y_pred), axis=-1))
# return K.equal(K.round(y_true), K.round(y_pred))
print(mean_pred(y_validate[pos], predictions[pos]).eval())
print(mean_pred(np.array([[ 0., 1., 0., 1., 0., 0., 0., 0., 0., 0., 0.],
[ 0., 1., 0., 1., 0., 0., 0., 0., 0., 0., 0.]]),
np.array([[ 0., 0.51, 0., 0.51, 0., 0., 0., 0., 0., 0., 0.],
[ 0., 0.92, 0., 0.51, 0., 0., 0., 0., 0., 0., 0.]])).eval())
# print(mean_pred(y_validate[pos], predictions[pos]))
# print(mean_pred2(y_validate[pos], predictions[pos]))
print(metrics.categorical_accuracy(y_validate[pos], predictions[pos]))
# -
predictions = model.predict(X[429145:])
decode_position(predictions[0], max_num_vowels)
# +
def test_accuracy(predictions, y):
dictionary, max_word, max_num_vowels, content, vowels, accetuated_vowels = create_dict()
num_of_pred = len(predictions)
num_of_correct_pred = 0
for i in range(predictions.shape[0]):
if decode_position(predictions[i], max_num_vowels) == decode_position(y[i], max_num_vowels):
num_of_correct_pred += 1
return (num_of_correct_pred/float(num_of_pred)) * 100
print(test_accuracy(predictions, y[429145:]))
# -
predictions.shape
print max_num_vowels
# +
dictionary, max_word, max_num_vowels, content, vowels, accetuated_vowels = create_dict()
feature_dictionary = create_feature_dictionary(content)
def generate_input_from_word(word, max_word, dictionary):
x = np.zeros((max_word, len(dictionary)))
j = 0
for c in list(word):
index = 0
for d in dictionary:
if c == d:
x[j, index] = 1
break
index += 1
j += 1
return x
# model = load_model()
# prediction = predict(model, generate_input_from_word('hidrija'))
# print decode_position(prediction[0])
# -
# %run ../../../prepare_data.py
# generate_X_and_y(dictionary, max_word, max_num_vowels, content, vowels, accetuated_vowels, feature_dictionary)
| cnn/word_accetuation/cnn_dictionary/v1_11/cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .venv
# language: python
# name: .venv
# ---
"""Walking throught the "mini-course" from MLMaster: https://machinelearningmastery.com/time-series-forecasting-python-mini-course/
Lesson 01: Time Series as Supervised Learning.
Lesson 02: Load Time Series Data.
Lesson 03: Data Visualization.
Lesson 04: Persistence Forecast Model.
Lesson 05: Autoregressive Forecast Model.
Lesson 06: ARIMA Forecast Model.
Lesson 07: Hello World End-to-End Project.
"""
#Lesson 01: Time Series as Supervised Learning.
"""Time series problems can be framed as a supervised learning problems. We will reoder data like such:
time, value -> value, value
1, 100 -> ?, 100
2, 110 -> 100, 110
3, 108 -> 110, 108
4, 115 -> 108, 115
5, 120 -> 115, 120
-> 120, ?
This is called the (sliding) window method. This converts time series data to linear data.
"""
# +
#Lesson 02: Load Time Series Data.
"""We will load a dataset and preview it"""
from pandas import read_csv
from matplotlib import pyplot
series = read_csv('https://raw.githubusercontent.com/jbrownlee/Datasets/master/daily-total-female-births.csv', header=0, index_col=0)
print(series.describe())
# -
#Lesson 03: Data Visualization.
"""Simply Plot this to preview it"""
pyplot.plot(series)
pyplot.show()
# +
#Lesson 04: Persistence Forecast Model.
"""Establishing a baseline is essential on any time series forecasting problem.
Three properties of a good technique for making a baseline forecast are:
Simple: A method that requires little or no training or intelligence.
Fast: A method that is fast to implement and computationally trivial to make a prediction.
Repeatable: A method that is deterministic, meaning that it produces an expected output given the same input.
"""
from sklearn.metrics import mean_squared_error
from math import sqrt
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.linear_model import LogisticRegression
def model_persistence(x):
"""https://machinelearningmastery.com/persistence-time-series-forecasting-with-python/"""
return x
predictions = []
actual = series.values[1:]
rmse = sqrt(mean_squared_error(actual, predictions))
| Time Series/time_series_course.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# # Imports and Simulation Parameters
# +
import numpy as np
import math
import cmath
import scipy
import scipy.integrate
import sys
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
hbar = 1.0 / (2.0 * np.pi)
ZERO_TOLERANCE = 10**-6
# +
MAX_VIBRATIONAL_STATES = 1500
STARTING_GROUND_STATES = 9
STARTING_EXCITED_STATES = 9
time_scale_set = 100 #will divide the highest energy to give us the time step
low_frequency_cycles = 20.0 #will multiply the frequency of the lowest frequency mode to get the max time
# -
#See if a factorial_Storage dictionary exists already and if not, create one
try:
a = factorial_storage
except:
factorial_storage = {}
# # Defining Parameters of the System
# +
energy_g = 0
energy_gamma = .1
energy_e = 5
energy_epsilon = .31
Huang_Rhys_Parameter = .80
S = Huang_Rhys_Parameter
#bookkeeping
overlap_storage = {}
electronic_energy_gap = energy_e + .5*energy_epsilon - (energy_g + .5 * energy_gamma)
min_energy = energy_g + energy_gamma * .5
# -
mu_0 = 1.0
# If we set the central frequency of a pulse at the 0->0 transition, and we decide we care about the ratio of the 0->1 transition to the 0->0 transition and set that to be $\tau$ then the desired pulse width will be
# \begin{align}
# \sigma &= \sqrt{-\frac{2 \ln (\tau)}{\omega_{\epsilon}^2}}
# \end{align}
# +
def blank_wavefunction(number_ground_states, number_excited_states):
return np.zeros((number_ground_states + number_excited_states))
def perturbing_function(time):
# stdev = 30000.0 * dt #very specific to 0->0 transition!
stdev = 3000.0 * dt #clearly has a small amount of amplitude on 0->1 transition
center = 6 * stdev
return np.cos(electronic_energy_gap*(time - center) / hbar)*np.exp( - (time - center)**2 / (2 * stdev**2)) / stdev
def time_function_handle_from_tau(tau_proportion):
stdev = np.sqrt( -2.0 * np.log(tau_proportion) / (energy_epsilon/hbar)**2)
center = 6 * stdev
return center, stdev, lambda t: np.cos(electronic_energy_gap*(t - center) / hbar)*np.exp( - (t - center)**2 / (2 * stdev**2)) / stdev
def perturbing_function_define_tau(time, tau_proportion):
center, stdev, f = time_function_handle_from_tau(tau_proportion)
return f(time)
# -
#
# # Defining Useful functions
# $ O_{m}^{n} = \left(-1\right)^{n} \sqrt{\frac{e^{-S}S^{m+n}}{m!n!}} \sum_{j=0}^{\min \left( m,n \right)} \frac{m!n!}{j!(m-j)!(n-j)!}(-1)^j S^{-j} $
# +
def factorial(i):
if i in factorial_storage:
return factorial_storage[i]
if i <= 1:
return 1.0
else:
out = factorial(i - 1) * i
factorial_storage[i] = out
return out
def ndarray_factorial(i_array):
return np.array([factorial(i) for i in i_array])
# -
def overlap_function(ground_quantum_number, excited_quantum_number):
m = ground_quantum_number
n = excited_quantum_number
if (m,n) in overlap_storage:
return overlap_storage[(m,n)]
output = (-1)**n
output *= math.sqrt(math.exp(-S) * S**(m + n) /(factorial(m) * factorial(n)) )
j_indeces = np.array(range(0, min(m,n) + 1))
j_summation = factorial(m) * factorial(n) * np.power(-1.0, j_indeces) * np.power(S, -j_indeces)
j_summation = j_summation / (ndarray_factorial(j_indeces) * ndarray_factorial( m - j_indeces) * ndarray_factorial(n - j_indeces) )
output *= np.sum(j_summation)
overlap_storage[(m,n)] = output
return output
# # Solving the Differential Equation
# \begin{align*}
# \left(\frac{d G_a(t)}{dt} + \frac{i}{\hbar}\Omega_{(a)}\right) &=-E(t)\frac{i}{\hbar} \sum_{b} E_b(t) \mu_{a}^{b}\\
# \left(\frac{d E_b(t)}{dt} + \frac{i}{\hbar} \Omega^{(b)} \right) &=-E(t)\frac{i}{\hbar} \sum_{a} G_a(t) \mu_{a}^{b}
# \end{align*}
# Or in a more compact form:
# \begin{align*}
# \frac{d}{dt}\begin{bmatrix}
# G_a(t) \\
# E_b(t)
# \end{bmatrix}
# = -\frac{i}{\hbar}
# \begin{bmatrix}
# \Omega_{(a)} & E(t) \mu_{a}^{b} \\
# E(t) \mu_{a}^{b} & \Omega^{b}
# \end{bmatrix}
# \cdot
# \begin{bmatrix}
# G_a(t) \\
# E_b(t)
# \end{bmatrix}
# \end{align*}
# +
def ode_diagonal_matrix(number_ground_states, number_excited_states):
#Define the Matrix on the RHS of the above equation
ODE_DIAGONAL_MATRIX = np.zeros((number_ground_states + number_excited_states, number_ground_states + number_excited_states), dtype=np.complex)
#set the diagonals
for ground_i in range(number_ground_states):
ODE_DIAGONAL_MATRIX[ground_i, ground_i] = -1.0j * (energy_g + energy_gamma * (ground_i + .5)) / hbar
for excited_i in range(number_excited_states):
excited_index = excited_i + number_ground_states #the offset since the excited states comes later
ODE_DIAGONAL_MATRIX[excited_index, excited_index] = -1.0j * (energy_e + energy_epsilon * (excited_i + .5)) / hbar
return ODE_DIAGONAL_MATRIX
#now for the off-diagonals
def mu_matrix(c, number_ground_states, number_excited_states):
MU_MATRIX = np.zeros((number_ground_states, number_excited_states), dtype = np.complex)
for ground_a in range(number_ground_states):
for excited_b in range(number_excited_states):
new_mu_entry = overlap_function(ground_a, excited_b)
if ground_a >0:
new_mu_entry += c * math.sqrt(ground_a) * overlap_function(ground_a - 1, excited_b)
new_mu_entry += c * math.sqrt(ground_a+1) * overlap_function(ground_a + 1, excited_b)
MU_MATRIX[ground_a, excited_b] = new_mu_entry
return MU_MATRIX
def ode_off_diagonal_matrix(c_value, number_ground_states, number_excited_states):
output = np.zeros((number_ground_states + number_excited_states, number_ground_states + number_excited_states), dtype=np.complex)
MU_MATRIX = mu_matrix(c_value, number_ground_states, number_excited_states)
output[0:number_ground_states, number_ground_states:] = -1.0j * mu_0 * MU_MATRIX / hbar
output[number_ground_states:, 0:number_ground_states] = -1.0j * mu_0 * MU_MATRIX.T / hbar
return output
def IR_transition_dipoles(number_ground_states, number_excited_states):
"outputs matrices to calculate ground and excited state IR emission spectra. Can be combined for total"
output_g = np.zeros((number_ground_states, number_excited_states), dtype = np.complex)
output_e = np.zeros((number_ground_states, number_excited_states), dtype = np.complex)
for ground_a in range(number_ground_states):
try:
output_g[ground_a, ground_a + 1] = 1.0
output_g[ground_a + 1, ground_a] = 1.0
except:
pass
try:
output_g[ground_a, ground_a - 1] = 1.0
output_g[ground_a - 1, ground_a] = 1.0
except:
pass
return output
# -
# \begin{align*}
# \mu(x) &= \mu_0 \left(1 + \lambda x \right) \\
# &= \mu_0 \left(1 + c\left(a + a^{\dagger} \right) \right) \\
# \mu_{a}^{b} &= \mu_0\left(O_{a}^{b} + c\left(\sqrt{a}O_{a-1}^{b} + \sqrt{a+1}O_{a+1}^{b}\right) \right)
# \end{align*}
class VibrationalStateOverFlowException(Exception):
def __init__(self):
pass
def propagate_amplitude_to_end_of_perturbation(c_value, ratio_01_00, starting_ground_states=STARTING_GROUND_STATES, starting_excited_states=STARTING_EXCITED_STATES):
center_time, stdev, time_function = time_function_handle_from_tau(ratio_01_00)
ending_time = center_time + 8.0 * stdev
number_ground_states = starting_ground_states
number_excited_states = starting_excited_states
while number_excited_states + number_ground_states < MAX_VIBRATIONAL_STATES:
#define time scales
max_energy = energy_e + energy_epsilon * (.5 + number_excited_states)
dt = 1.0 / (time_scale_set * max_energy)
ODE_DIAGONAL = ode_diagonal_matrix(number_ground_states, number_excited_states)
ODE_OFF_DIAGONAL = ode_off_diagonal_matrix(c_value, number_ground_states, number_excited_states)
def ODE_integrable_function(time, coefficient_vector):
ODE_TOTAL_MATRIX = ODE_OFF_DIAGONAL * time_function(time) + ODE_DIAGONAL
return np.dot(ODE_TOTAL_MATRIX, coefficient_vector)
#define the starting wavefuntion
initial_conditions = blank_wavefunction(number_ground_states, number_excited_states)
initial_conditions[0] = 1
#create ode solver
current_time = 0.0
ode_solver = scipy.integrate.complex_ode(ODE_integrable_function)
ode_solver.set_initial_value(initial_conditions, current_time)
#Run it
results = []
try: #this block catches an overflow into the highest ground or excited vibrational state
while current_time < ending_time:
# print(current_time, ZERO_TOLERANCE)
#update time, perform solution
current_time = ode_solver.t+dt
new_result = ode_solver.integrate(current_time)
results.append(new_result)
#make sure solver was successful
if not ode_solver.successful():
raise Exception("ODE Solve Failed!")
#make sure that there hasn't been substantial leakage to the highest excited states
re_start_calculation = False
if abs(new_result[number_ground_states - 1])**2 >= ZERO_TOLERANCE:
number_ground_states +=1
print("Increasing Number of Ground vibrational states to %i " % number_ground_states)
# print("check this:", np.abs(new_result)**2, number_ground_states, abs(new_result[number_ground_states])**2)
# raise Exception()
re_start_calculation = True
if abs(new_result[-1])**2 >= ZERO_TOLERANCE:
number_excited_states +=1
print("Increasing Number of excited vibrational states to %i " % number_excited_states)
re_start_calculation = True
if re_start_calculation:
raise VibrationalStateOverFlowException()
except VibrationalStateOverFlowException:
#Move on and re-start the calculation
continue
#Finish calculating
results = np.array(results)
return results, number_ground_states, number_excited_states
raise Exception("NEEDED TOO MANY VIBRATIONAL STATES! RE-RUN WITH DIFFERENT PARAMETERS!")
def get_average_quantum_number_time_series(c_value, ratio_01_00, starting_ground_states=STARTING_GROUND_STATES, starting_excited_states=STARTING_EXCITED_STATES):
results, number_ground_states, number_excited_states = propagate_amplitude_to_end_of_perturbation(c_value, ratio_01_00, starting_ground_states, starting_excited_states)
probabilities = np.abs(results)**2
#calculate the average_vibrational_quantum_number series
average_ground_quantum_number = probabilities[:,0:number_ground_states].dot(np.array(range(number_ground_states)) )
average_excited_quantum_number = probabilities[:,number_ground_states:].dot(np.array(range(number_excited_states)))
return average_ground_quantum_number, average_excited_quantum_number, results, number_ground_states, number_excited_states
# +
def IR_emission_spectrum_after_excitation(c_value, ratio_01_00, starting_ground_states=STARTING_GROUND_STATES, starting_excited_states=STARTING_EXCITED_STATES):
center_time, stdev, time_function = time_function_handle_from_tau(ratio_01_00)
ending_time = center_time + 8.0 * stdev
ending_time += low_frequency_cycles * hbar/min_energy
number_ground_states = starting_ground_states
number_excited_states = starting_excited_states
while number_excited_states + number_ground_states < MAX_VIBRATIONAL_STATES:
#define time scales
max_energy = energy_e + energy_epsilon * (.5 + number_excited_states)
dt = 1.0 / (time_scale_set * max_energy)
ODE_DIAGONAL = ode_diagonal_matrix(number_ground_states, number_excited_states)
ODE_OFF_DIAGONAL = ode_off_diagonal_matrix(c_value, number_ground_states, number_excited_states)
def ODE_integrable_function(time, coefficient_vector):
ODE_TOTAL_MATRIX = ODE_OFF_DIAGONAL * time_function(time) + ODE_DIAGONAL
return np.dot(ODE_TOTAL_MATRIX, coefficient_vector)
#define the starting wavefuntion
initial_conditions = blank_wavefunction(number_ground_states, number_excited_states)
initial_conditions[0] = 1
#create ode solver
current_time = 0.0
ode_solver = scipy.integrate.complex_ode(ODE_integrable_function)
ode_solver.set_initial_value(initial_conditions, current_time)
#Run it
results = []
try: #this block catches an overflow into the highest ground or excited vibrational state
while current_time < ending_time:
# print(current_time, ZERO_TOLERANCE)
#update time, perform solution
current_time = ode_solver.t+dt
new_result = ode_solver.integrate(current_time)
results.append(new_result)
#make sure solver was successful
if not ode_solver.successful():
raise Exception("ODE Solve Failed!")
#make sure that there hasn't been substantial leakage to the highest excited states
re_start_calculation = False
if abs(new_result[number_ground_states - 1])**2 >= ZERO_TOLERANCE:
number_ground_states +=1
print("Increasing Number of Ground vibrational states to %i " % number_ground_states)
# print("check this:", np.abs(new_result)**2, number_ground_states, abs(new_result[number_ground_states])**2)
# raise Exception()
re_start_calculation = True
if abs(new_result[-1])**2 >= ZERO_TOLERANCE:
number_excited_states +=1
print("Increasing Number of excited vibrational states to %i " % number_excited_states)
re_start_calculation = True
if re_start_calculation:
raise VibrationalStateOverFlowException()
except VibrationalStateOverFlowException:
#Move on and re-start the calculation
continue
#Finish calculating
results = np.array(results)
return results, number_ground_states, number_excited_states
raise Exception("NEEDED TOO MANY VIBRATIONAL STATES! RE-RUN WITH DIFFERENT PARAMETERS!"))
low_frequency_cycles
# +
c_values = np.logspace(-3, np.log10(.9), 31)
tau_values = np.logspace(-4, np.log10(.9), 30)
heating_results_ground = np.zeros((c_values.shape[0], tau_values.shape[0]))
heating_results_excited = np.zeros((c_values.shape[0], tau_values.shape[0]))
n_g = STARTING_GROUND_STATES
n_e = STARTING_EXCITED_STATES
dict_cTau_mapsTo_NgNe = {}
for i_c, c in enumerate(c_values):
# as we increase in both tau and
for i_tau, tau in enumerate(tau_values):
print(c, tau)
#make a good guess for the new needed number of simulated states
try:
last_c = c_values[i_c - 1]
except:
last_c = None
try:
last_tau = tau_values[i_tau - 1]
except:
last_tau = None
if last_tau is not None and (c, last_tau) in dict_cTau_mapsTo_NgNe:
n_g_candidate1, n_e_candidate1 = dict_cTau_mapsTo_NgNe[(c, last_tau)]
# n_g_candidate1 += 1
# n_e_candidate1 += 1
else:
n_g_candidate1, n_e_candidate1 = n_g, n_e
if last_c is not None and (last_c, tau) in dict_cTau_mapsTo_NgNe:
n_g_candidate2, n_e_candidate2 = dict_cTau_mapsTo_NgNe[(last_c, tau)]
# n_g_candidate2 += 1
# n_e_candidate2 += 1
else:
n_g_candidate2, n_e_candidate2 = n_g, n_e
n_g = max([n_g_candidate1, n_g_candidate2])
n_e = max([n_e_candidate1, n_e_candidate2])
sys.stdout.flush()
sys.stdout.write("Calculating c=%f, tau=%f at n_g = %i and n_e=%i..." %(c, tau, n_g, n_e))
n_bar_g, n_bar_e, results, num_g, num_e = get_average_quantum_number_time_series(c,
tau,
starting_ground_states = n_g,
starting_excited_states = n_e)
dict_cTau_mapsTo_NgNe[(c, tau)] = (num_g, num_e)
heating_results_ground[i_c, i_tau] = n_bar_g[-1]
heating_results_excited[i_c, i_tau] = n_bar_e[-1]
n_g = num_g
n_e = num_e
# -
n_g, n_e
# +
plt.figure()
plt.title("Ground State Heating")
plt.contourf(np.log10(tau_values),np.log10(c_values), heating_results_ground, 100)
plt.colorbar()
plt.ylabel(r"$c$ log scale")
plt.xlabel(r"$\tau$ log scale")
plt.savefig("ground_state_heating.png")
plt.figure()
plt.title("Excited State Heating")
plt.contourf(np.log10(tau_values),np.log10(c_values), heating_results_excited, 100)
plt.colorbar()
plt.ylabel(r"$c$ log scale")
plt.xlabel(r"$\tau$ log scale")
plt.savefig("excited_state_heating.png")
# -
n_bar_g, n_bar_e, results, number_ground_states = get_average_quantum_number_time_series(.3, .8)
# +
plt.figure()
plt.plot(n_bar_g, label="g")
plt.plot(n_bar_e, label="e")
plt.legend(loc=0)
plt.figure("ground")
plt.title("Ground State Populations")
plt.figure("excited")
plt.title("Excited State Populations")
# plt.semilogy(np.abs(time_function) /np.max(np.abs(time_function)))
for index in range(results.shape[1]):
if index < number_ground_states:
plt.figure("ground")
plt.semilogy(abs(results[:, index])**2, label=index)
else:
plt.figure("excited")
plt.semilogy(abs(results[:, index])**2, label=index - number_ground_states)
plt.figure("ground")
plt.legend(loc=0)
plt.figure("excited")
plt.legend(loc=0)
# -
| MolmerSorenson/.ipynb_checkpoints/HeatingCalculations-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# https://arrow.apache.org/docs/python/parquet.html
# https://arrow.apache.org/docs/python/install.html
# conda install -c conda-forge pyarrow
# Athena - schema update support:
# https://docs.aws.amazon.com/athena/latest/ug/handling-schema-updates-chapter.html
# https://docs.aws.amazon.com/athena/latest/ug/glue-best-practices.html
import pandas as pd
import pyarrow as pa
import numpy as np
import pyarrow.parquet as pq
# -
file_names = ['iris_missing_column','iris_new_column_at_end','iris_new_column_in_middle','iris_new_plant_type', 'iris_setosa','iris_versicolor','iris_virginica']
file_names
for f in file_names:
df = pd.read_csv(f + '.csv')
table = pa.Table.from_pandas(df)
pq.write_table(table, f + '.parquet')
# +
# Read back one of the files
# -
table2 = pq.read_table('iris_new_column_in_middle.parquet')
table2.to_pandas()
| DataLake/Iris/iris_csv_to_parquet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pranjalchaubey/60-days-of-udacity-sixty-ai/blob/master/03%20Sixty%20AI%20Training/Sixty_AI_Training.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="H7LoMj4GA4n_" colab_type="text"
# # Sixty AI - Train a GPT-2 on Text Corpus
#
# <br/> We are now going to use our processed text corpus to train OpenAI's GPT-2 Model.
# <br/> Of course, we are not going to use the _original_ GPT-2 from OpenAI. We are instead going to use _GPT-2 Simple_ from _**Max Woolf**_.
# <br/> GPT-2 Simple is a pretrained GPT-2 model from OpenAI, but with the added functionality of _Finetuning_. We will use our tiny text corpus to finetune a full blown GPT-2 (small _117M_ model) , so that it starts generating some creative text content on its own. This is _NLP Transfer Learning_ live in action!
#
# <br/> For more information about `gpt-2-simple`, you can visit [this GitHub repository](https://github.com/minimaxir/gpt-2-simple).
#
#
# + id="KBkpRgBCBS2_" colab_type="code" colab={}
# Install the GPT-2 Simple library
# !pip install -q gpt-2-simple
# Import Export business
import gpt_2_simple as gpt2
from datetime import datetime
from google.colab import files
# + [markdown] id="Bj2IJLHP3KwE" colab_type="text"
# ## GPU
#
# Colaboratory now uses an Nvidia T4 GPU, which is slightly faster than the old Nvidia K80 GPU for training GPT-2, and has more memory allowing us to train the larger GPT-2 models and generate more text.
#
# Let's verify which GPU is active.
# + id="sUmTooTW3osf" colab_type="code" outputId="7cf845c6-55e7-4378-8f06-f860dae270c9" colab={"base_uri": "https://localhost:8080/", "height": 295}
# !nvidia-smi
# + [markdown] id="0wXB05bPDYxS" colab_type="text"
# ## Downloading GPT-2
#
# We need to download the GPT-2 model first.
#
# There are two sizes of GPT-2:
#
# * `117M` (default): the "small" model, 500MB on disk.
# * `345M`: the "medium" model, 1.5GB on disk.
#
# Larger models have more knowledge, but take longer to finetune and longer to generate text.
# <br/>We will use the smaller 117M model to start things off.
# <br/>The next cell downloads it from Google Cloud Storage and saves it in the Colaboratory VM at `/models/<model_name>`.
#
# + id="P8wSlgXoDPCR" colab_type="code" outputId="9ccd02f0-4e1a-41ab-e1c7-89cb3a7023c3" colab={"base_uri": "https://localhost:8080/", "height": 139}
# Download the smaller 117M model
gpt2.download_gpt2(model_name="117M")
# + [markdown] id="N8KXuKWzQSsN" colab_type="text"
# ## Mounting Google Drive
#
# The best way to get input text to-be-trained into the Colaboratory VM, and to get the trained model *out* of Colaboratory, is to route it through Google Drive *first*.
# <br/>In case you're a little skeptical about putting down the Google Drive auth code (you should be!), I suggest you check out what is going on under the hood in the `gpt-2-simple` library. Simply [click this link](https://github.com/minimaxir/gpt-2-simple/blob/master/gpt_2_simple/gpt_2.py "click this link").
#
# <br/>TL;DR: _It's Safe!_
# + id="puq4iC6vUAHc" colab_type="code" outputId="748f08ad-1e90-4981-aa35-cd6568041846" colab={"base_uri": "https://localhost:8080/", "height": 35}
gpt2.mount_gdrive()
# + [markdown] id="BT__brhBCvJu" colab_type="text"
# ## Uploading our Text Corpus to be Trained
#
# Let's upload our text corpus in the _'Files'_ section (this has to be done manually).
# + id="6OFnPCLADfll" colab_type="code" colab={}
file_name = "final_text_corpus.csv"
model_name = 'run1' # Default Name
# + [markdown] id="LdpZQXknFNY3" colab_type="text"
# ## Finetune GPT-2
#
# Finally, it's time to _finetune_ our Simple GPT-2 Model on our extracted corpus of text.
# <br/>The next cell will start the actual finetuning of GPT-2. It creates a persistent TensorFlow session which stores the training config, then runs the training for the specified number of `steps`. (setting `steps = -1` will run the finetuning indefinitely)
#
# The model checkpoints will be saved in `/checkpoint/run1` by default. The checkpoints are saved every 500 steps and when the cell is stopped.
#
#
#
#
#
# + id="aeXshJM-Cuaf" colab_type="code" outputId="79acffb5-6db9-4bd2-de3f-f07c0e3804ec" colab={"base_uri": "https://localhost:8080/", "height": 1000}
"""
Parameters for gpt2.finetune():
restore_from: Set to 'fresh' to start training from the base GPT-2,
or set to 'latest' to restart training from an existing checkpoint.
sample_every: Number of steps to print example output
print_every: Number of steps to print training progress.
learning_rate: Learning rate for the training.
(default '1e-4', can lower to '1e-5' if you have <1MB input data)
run_name: subfolder within 'checkpoint' to save the model.
This is useful if you want to work with multiple models
(will also need to specify 'run_name' when loading the model)
overwrite: Set to 'True' if you want to continue finetuning an existing
model (w/ restore_from='latest') without creating duplicate copies.
"""
# Start the tf session
# LOL.....why they have a 'session' in tf?! :D
sess = gpt2.start_tf_sess()
# We will train for 1000 epochs, as I have noticed that beyond 1000
# epochs the model starts to overfit on the data
gpt2.finetune(sess,
dataset=file_name,
model_name='117M',
steps=1000,
restore_from='fresh',
run_name=model_name,
print_every=10,
sample_every=200,
save_every=500
)
# + [markdown] id="IXSuTNERaw6K" colab_type="text"
# Copy the trained model to the Google Drive. The checkpoint folder is copied as a `.rar` compressed file.
# + id="VHdTL8NDbAh3" colab_type="code" colab={}
gpt2.copy_checkpoint_to_gdrive(run_name=model_name)
# + [markdown] id="ClJwpF_ACONp" colab_type="text"
# ## Generate Text From The Trained Model
#
# Now that we have trained and/or loaded our finetuned model, its time to generate text!
# + id="8DKMc0fiej4N" colab_type="code" outputId="c3b63fb2-9b9b-4128-e896-6e4b071a04ac" colab={"base_uri": "https://localhost:8080/", "height": 1000}
'''
prefix: force the text to start with a given character sequence and generate
text from there
nsamples: generate multiple texts at a time
batch_size: generate multiple samples in parallel, giving a massive speedup
(in Colaboratory, set a maximum of 20 for batch_size)
length: Number of tokens to generate (default 1023, the maximum)
temperature: The higher the temperature, the crazier the text
(default 0.7, recommended to keep between 0.7 and 1.0)
top_k: Limits the generated guesses to the top k guesses
(default 0 which disables the behavior; if the generated output is
super crazy, you may want to set top_k=40)
top_p: Nucleus sampling: limits the generated guesses to a
cumulative probability. (gets good results on a dataset with top_p=0.9)
truncate: Truncates the input text until a given sequence, excluding that
sequence (e.g. if truncate='<|endoftext|>', the returned text will include
everything before the first <|endoftext|>). It may be useful to combine this
with a smaller length if the input texts are short.
include_prefix: If using truncate and include_prefix=False, the specified
prefix will not be included in the returned text.
'''
gpt2.generate(sess,
length=512,
temperature=0.8,
prefix="Day: ",
nsamples=10,
batch_size=10
)
# + [markdown] id="ig-KVgkCDCKD" colab_type="text"
# # Troubleshoot
#
# If the notebook has errors (e.g. GPU Sync Fail or out-of-memory/OOM), force-kill the Colaboratory virtual machine and restart it with the command below:
# + id="rIHiVP53FnsX" colab_type="code" colab={}
# !kill -9 -1
| Pranjal Chaubey/60-days-of-udacity-sixty-ai/03 Sixty AI Training/Sixty_AI_Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
import cv2
data = '../../data/raw'
# %matplotlib inline
import random
import string
train = pd.read_csv('../../data/raw/train.csv')
depths = pd.read_csv('../../data/raw/depths.csv')
x=1157
img = cv2.imread(data+'/train/images/'+train.iloc[x][0]+'.png')
out = cv2.imread(data+'/train/masks/'+train.iloc[x][0]+'.png')
img = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
rows,cols = img.shape
M = cv2.getRotationMatrix2D((cols/2,rows/2),90,1)
dst = cv2.warpAffine(img,M,(cols,rows))
for i in range(4000):
img = cv2.imread(data+'/train/images/'+train.iloc[i][0]+'.png')
out = cv2.imread(data+'/train/masks/'+train.iloc[i][0]+'.png')
img = cv2.cvtColor(img,cv2.COLOR_BGR2GRAY)
out = cv2.cvtColor(out,cv2.COLOR_BGR2GRAY)
rows,cols = img.shape
M1 = cv2.getRotationMatrix2D((cols/2,rows/2),90,1)
M2 = cv2.getRotationMatrix2D((cols/2,rows/2),180,1)
M3 = cv2.getRotationMatrix2D((cols/2,rows/2),270,1)
M4 = cv2.getRotationMatrix2D((cols/2,rows/2),360,1)
in_1 = cv2.warpAffine(img,M1,(cols,rows))
in_2 = cv2.warpAffine(img,M2,(cols,rows))
in_3 = cv2.warpAffine(img,M3,(cols,rows))
in_4 = cv2.warpAffine(img,M4,(cols,rows))
out_1 = cv2.warpAffine(out,M1, (cols,rows))
out_2 = cv2.warpAffine(out,M2, (cols,rows))
out_3 = cv2.warpAffine(out,M3, (cols,rows))
out_4 = cv2.warpAffine(out,M4, (cols,rows))
cv2.imwrite('../../data/processed/train/images/'+train.iloc[i][0]+'_00.png',in_1)
cv2.imwrite('../../data/processed/train/images/'+train.iloc[i][0]+'_01.png',in_2)
cv2.imwrite('../../data/processed/train/images/'+train.iloc[i][0]+'_02.png',in_3)
cv2.imwrite('../../data/processed/train/images/'+train.iloc[i][0]+'_03.png',in_4)
cv2.imwrite('../../data/processed/train/masks/'+train.iloc[i][0]+'_00.png',out_1)
cv2.imwrite('../../data/processed/train/masks/'+train.iloc[i][0]+'_01.png',out_2)
cv2.imwrite('../../data/processed/train/masks/'+train.iloc[i][0]+'_02.png',out_3)
cv2.imwrite('../../data/processed/train/masks/'+train.iloc[i][0]+'_03.png',out_4)
plt.imshow(img,cmap='gray')
plt.imshow(dst)
cv2.imwrite('../../data/dfds.png',img)
| notebooks/preprocessing/DataAugmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="ojm_6E9f9Kcf"
# # K-mer
# Basic K-mer counting.
# -
import time
def show_time():
t = time.time()
print(time.strftime('%Y-%m-%d %H:%M:%S %Z', time.localtime(t)))
show_time()
PC_SEQUENCES=32000
NC_SEQUENCES=32000
RNA_LEN=32
CDS_LEN=16
# + id="VQY7aTj29Kch"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# -
import sys
IN_COLAB = False
try:
from google.colab import drive
IN_COLAB = True
except:
pass
if IN_COLAB:
print("On Google CoLab, mount cloud-local file, get our code from GitHub.")
PATH='/content/drive/'
#drive.mount(PATH,force_remount=True) # hardly ever need this
#drive.mount(PATH) # Google will require login credentials
DATAPATH=PATH+'My Drive/data/' # must end in "/"
import requests
r = requests.get('https://raw.githubusercontent.com/ShepherdCode/Soars2021/master/SimTools/RNA_describe.py')
with open('RNA_describe.py', 'w') as f:
f.write(r.text)
from RNA_describe import Random_Base_Oracle
else:
print("CoLab not working. On my PC, use relative paths.")
DATAPATH='data/' # must end in "/"
sys.path.append("..") # append parent dir in order to use sibling dirs
from SimTools.RNA_describe import Random_Base_Oracle
MODELPATH="BestModel" # saved on cloud instance and lost after logout
#MODELPATH=DATAPATH+MODELPATH # saved on Google Drive but requires login
# ## K-mer counting
# ### Functions to create the dict of {kmer:count}
def make_kmer_keys(K):
shorter_kmers=['']
for i in range(K):
longer_kmers=[]
for mer in shorter_kmers:
# No support for N or any non-ACGT bases.
longer_kmers.append(mer+'A')
longer_kmers.append(mer+'C')
longer_kmers.append(mer+'G')
longer_kmers.append(mer+'T')
shorter_kmers = longer_kmers
return shorter_kmers
def make_kmer_dict(keys,init=0):
return dict.fromkeys(keys,init)
def make_dict_upto_K(max_K):
keys=make_kmer_keys(1)
for k in range(2,max_K+1):
keys.extend(make_kmer_keys(k))
counts = make_kmer_dict(keys)
return counts
# ### Naive K-mer counting algorithm
# Algorithm:
# 1. for every string
# 1. for every K
# 1. for every position
# 1. kmer=substring
# 2. count{kmer}++
def update_count_one_K(counts,K,rna,tail=False):
L = len(rna)
padding=" "*(K-1)
padded=rna+padding
for i in range(0,L-K+1):
kmer=padded[i:i+K]
counts[kmer] += 1
if tail and K>1:
# for Harvester algorithm, count last letters as special case
for start_pos in range(L-K+1,L):
for end_pos in range(start_pos+1,L+1):
kmer=rna[start_pos:end_pos]
counts[kmer] += 1
return counts
def update_count_upto_K(counts,max_K,sample,tail=False):
for i in range(1,max_K+1):
update_count_one_K(counts,i,sample,tail)
return counts
# ### Harvester K-mer counting algorithm
# Algorithm:
# 1. Count K-mers for max K only
# 2. For each K-mer in counts table:
# 1. For every prefix of the K-mer:
# 1. count{prefix} += count{kmer}
# 3. Handle last K-1 letters of each string as special case
def harvest_counts_from_K(counts,max_K):
for kmer in counts.keys():
klen = len(kmer)
kcnt = counts[kmer]
if klen==max_K and kcnt>0:
for i in range(1,klen):
prefix = kmer[:i]
counts[prefix] += kcnt
return counts
def count_to_frequency(counts,max_K):
freqs = dict.fromkeys(counts.keys(),0.0)
for k in range(1,max_K+1):
tot = 0
for kmer in counts.keys():
if len(kmer)==k:
tot += counts[kmer]
for kmer in counts.keys():
if len(kmer)==k:
freqs[kmer] = 1.0*counts[kmer]/tot
return freqs
# ## Demo
# ### Demo: Naive algorithm
# +
MAX_K = 3
counts1 = make_dict_upto_K(MAX_K)
print("Initial counts:\n",counts1)
sample = "ACCGGGTTTTACGTACGT"
update_count_upto_K(counts1,MAX_K,sample)
print("Final counts:\n",counts1)
# -
# ### Demo: Harvester algorithm
# +
MAX_K = 3
counts2 = make_dict_upto_K(MAX_K)
print("Initial counts:\n",counts2)
sample = "ACCGGGTTTTACGTACGT"
update_count_one_K(counts2,MAX_K,sample,True)
print("Partial counts (just max K and special case letters)\n:",counts2)
harvest_counts_from_K(counts2,MAX_K)
print("Final counts (includes smaller values of K):\n",counts2)
# -
if counts1==counts2:
print("Success. Harvester output matches naive results!")
else:
print("Fail. Harvester output differs from naive results!")
freqs = count_to_frequency(counts2,MAX_K)
print ("Frequency:\n",freqs)
# ## Demo on large dataset
rbo=Random_Base_Oracle(RNA_LEN,True)
pc_all,nc_all = rbo.get_partitioned_sequences(CDS_LEN,10) # just testing
pc_all,nc_all = rbo.get_partitioned_sequences(CDS_LEN,PC_SEQUENCES)
print("Use",len(pc_all),"PC seqs")
print("Use",len(nc_all),"NC seqs")
MAX_K = 3
pc_counts = make_dict_upto_K(MAX_K)
for sample in pc_all:
update_count_one_K(pc_counts,MAX_K,sample,True)
harvest_counts_from_K(pc_counts,MAX_K)
print("PC counts:\n",pc_counts)
pc_freqs = count_to_frequency(pc_counts,MAX_K)
print ("Frequency:\n",pc_freqs)
nc_counts = make_dict_upto_K(MAX_K)
for sample in nc_all:
update_count_one_K(nc_counts,MAX_K,sample,True)
harvest_counts_from_K(nc_counts,MAX_K)
print("NC counts:\n",nc_counts)
nc_freqs = count_to_frequency(nc_counts,MAX_K)
print ("Frequency:\n",nc_freqs)
| Notebooks/Kmer_100.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: machinelearningclass
# language: python
# name: machinelearningclass
# ---
# The following script consists of a Python version of Andrew Ng Stanford Course 'Machine Learning' taught on the Coursera Platform
# Note: All exercise data and structure are credited to Stanford University
#
# **Caveat:** Contrary to the modularity presented in Octave scripts and as I'm using Jupyter Notebooks for educational purposes we will implement the functions on the same notebook where we will call them
# # Exercise 2 - Principle Component Analysis
# +
# Import numpy libraries to deal with matrixes and vectors
import numpy as np
# Import pandas do read data files
import pandas as pd
# Import matplotlib to plot data
import matplotlib.pyplot as plt
# Import Image
from PIL import Image
# Import math
import math
# Import scipy optimization function
from scipy import optimize, io
# Scipy rotate function
from scipy.ndimage import rotate
# Matplotlib notebook property
# %matplotlib inline
# -
# **Let's now look at another famous technique used to reduce dimensionality - Principal Component Analysis:**
# Read the dataset
X = io.loadmat('ex7data1.mat')['X']
# Let's take a look at the points loaded in matrix 'data1'
plt.scatter(X[:,0], X[:,1], color='white', edgecolors='blue')
# PCA is nothing more than a projection of the original data on a new 'axis'.
# <br>
# From the plot above we can 'bring down' the points to the X-axis and only use one variable as a proxy. This, of course, makes you lose information.
# <br>
# <br>
# An important step to develop PCA is to normalize features:
# Normalize Features - Applying standardization of the variables
def featureNormalize(
features: np.array
) -> [np.array, np.array, np.array]:
'''
Computes feature normalization
by subtracting the mean and dividing
by the standard deviation.
Args:
features(np.array): Original data.
Returns:
X_norm(np.array): Normalized data.
mean(float): mean of each column -
array of size n.
sigma(float): standard deviation of
each column - array of size n.
'''
mean = X.mean(axis=0)
# To get the same result as Octave std function you have to set degrees of freedom to 1 with the ddof parameter
sigma = X.std(axis=0,ddof=1)
X_norm = ((X-mean)/sigma)
return X_norm, mean, sigma
X_norm, mu, sigma = featureNormalize(X)
# +
# Executing Principal Component Analysis
def PCA(
X: np.array
) -> [np.array, np.array, np.array]:
'''
Use singular value decomposition
to extract the factorization matrixes
that will be able to construct the original
matrix.
Args:
X(np.array): Original features.
Returns:
U(np.array): First matrix of decomposition.
S(np.array): Second matrix of decomposition.
V(np.array): Third matrix of decomposition.
'''
m, n = X.shape
U = np.zeros(n)
S = np.zeros(n)
# Compute covariance matrix
cov = np.dot(X.T,X)/m
# Use numpy singular value decomposition to return EigenValues and EigenVectors
U, S, V = np.linalg.svd(cov)
return U,S,V
# -
U,S,V = PCA(X_norm)
# Let's take a look at normalized data
plt.scatter(X_norm[:,0], X_norm[:,1], color='white', edgecolors='blue')
# **How can we project this data to 1-dimension?**
K = 1
def projectData(
X: np.array,
U: np.array,
K: int
):
'''
Projects data based on
singular value decomposition in
k dimensions.
Args:
X(np.array): Original variables.
U(np.array): First factor of decomposition.
K(int): Number of dimensions.
Returns:
Z(np.array): Projected data.
'''
Z = np.zeros([X.shape[0],K])
Z = np.dot(X, U[:, :K])
return Z
Z = projectData(X_norm, U, K)
def recoverData(
Z: np.array,
U: np.array,
K: int
)-> np.array:
'''
Recovers data from original projection.
Args:
Z(np.array): Projected data.
U(np.array): First factor of decomposition.
K(int): Number of dimensions.
Returns:
X_rec(np.array): Recovered data.
'''
X_rec = np.zeros([Z.shape[1],U.shape[1]])
X_rec = Z.dot(U[:, :K].T)
return X_rec
X_rec = recoverData(Z, U, K)
# By the figure below, you should understand how the real projection works. Basically, the points are projected into this single line, which can be handy for a lot of machine learning algorithms.
# +
fig, ax = plt.subplots(figsize=(5, 5))
# Plotting normalized data
plt.scatter(X_norm[:,0], X_norm[:,1], color='white', edgecolors='blue')
# Project the new points
plt.scatter(X_rec, X_rec,color='white', edgecolors='red')
# Plot lines connecting
ax.plot(X_rec[:, 0], X_rec[:, 1], 'ro', mec='r', mew=2, mfc='none')
for xnorm, xrec in zip(X_norm, X_rec):
ax.plot([xnorm[0], xrec[0]], [xnorm[1], xrec[1]], '--k', lw=1)
# -
# Above we can see, in Red, the new projected data into a k=1 dimensions. Basically, every point of our bi-dimensional dataser as been projected into a single dimension - be aware that by doing so we lose some information about the original variables.
# Read the Face dataset
X = io.loadmat('ex7faces.mat')['X']
def displayData(
images: np.array,
figsize1: int,
figsize2: int
) -> None:
'''
Plot 10 by 10 grid of images.
Args:
images(np.array): Array of original images.
figsize1(int): Size of grid - horizontal.
figsize2(int): Size of grid - vertical.
Returns:
None
'''
fig=plt.figure(figsize=(figsize1, figsize2))
columns = figsize1
rows = figsize2
for i in range(0, images.shape[0]-1):
img = images[i-1].reshape(32,32)
if i > images.shape[0]:
pass
else:
fig.add_subplot(rows, columns, i+1)
plt.imshow(rotate(img[::-1],270), cmap='gray')
plt.axis('off')
plt.show()
displayData(X[0:101], 10,10)
# **Can you guess what happens if we run PCA on image data?**
X_norm, mu, sigma = featureNormalize(X)
U,S,V = PCA(X_norm)
# Below we are goijg to project our face images into 100 dimensions, less than the original data. We will certainly lose some information about them but we expect that some of the face features will remain intact - such as contours of the faces.
# +
K = 100
Z = projectData(X_norm, U, K)
# -
X_rec = recoverData(Z, U, K)
displayData(X_rec[0:101], 10,10)
# Some high level contours of the faces have been lost. But notice how features such as nose and eyes shape still remain. What happens if we just retain 20 eigenvectors?
K = 20
Z = projectData(X_norm, U, K)
X_rec = recoverData(Z, U, K)
displayData(X_rec[0:101], 10,10)
# Even more details of the faces are stripped out and you start to get a more blurred image. As you project the data to less and less dimension you start to have less detail and information about those images.
# Principal Component Analysis is a pretty good algorithm to compress the size of your data (for example, it is used as a way to stream information across networks with less bandwith) and also to prevent some overfitting of the data. <br>In some cases, the information that you lose by reducing dimensions might also be the data that is preventing the algorithm to generalize for examples different from the training data.
| Programming Assignment 8/.ipynb_checkpoints/Exercise 7 - Principle Component Analysis-checkpoint.ipynb |
# # Estimating the correlation between two variables with a contingency table and a chi-squared test
import numpy as np
import pandas as pd
import scipy.stats as st
import matplotlib.pyplot as plt
# %matplotlib inline
player = '<NAME>'
df = pd.read_csv('https://github.com/ipython-books/'
'cookbook-2nd-data/blob/master/'
'federer.csv?raw=true',
parse_dates=['start date'],
dayfirst=True)
# + podoc={"output_text": "Some tournaments"}
print(f"Number of columns: {len(df.columns)}")
df[df.columns[:4]].tail()
# -
npoints = df['player1 total points total']
points = df['player1 total points won'] / npoints
aces = df['player1 aces'] / npoints
# + podoc={"output_text": "Aces and won points"}
fig, ax = plt.subplots(1, 1)
ax.plot(points, aces, '.')
ax.set_xlabel('% of points won')
ax.set_ylabel('% of aces')
ax.set_xlim(0., 1.)
ax.set_ylim(0.)
# + podoc={"output_text": "Aces and won points"}
df_bis = pd.DataFrame({'points': points,
'aces': aces}).dropna()
df_bis.tail()
# + podoc={"output_text": "Pearson correlation coefficient"}
df_bis.corr()
# -
df_bis['result'] = (df_bis['points'] >
df_bis['points'].median())
df_bis['manyaces'] = (df_bis['aces'] >
df_bis['aces'].median())
# + podoc={"output_text": "Contingency table"}
pd.crosstab(df_bis['result'], df_bis['manyaces'])
# -
st.chi2_contingency(_)
| Section01/05_correlation(1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import csv
import sys
import re
import scipy
import numpy as np
from sklearn import svm
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.model_selection import RandomizedSearchCV
from scipy.stats import randint as sp_randint
from time import time
csv.field_size_limit(sys.maxsize)
# -
df = pd.read_pickle('../data/instances.pkl')
labels = list(set(df['target'].values))
# +
X = []
Y = []
print("Preparing lists...")
for index, row in df.iterrows():
X.append(row["source_code"])
Y.append(row["target"])
# -
print("Extracting features...")
cv = CountVectorizer(binary=True)
cv.fit(X)
instances = cv.transform(X)
X_train, X_test, y_train, y_test = train_test_split(instances, Y, train_size = 0.75, random_state=42)
# # Default parameters
svm_classifier = svm.SVC(random_state=42, verbose=1)
svm_classifier.fit(X_train, y_train)
print("============ EVALUATION on test set:")
print(accuracy_score(y_test, svm_classifier.predict(X_test)))
# # Hyperparametrization
# +
def report(results, n_top=3):
for i in range(1, n_top + 1):
candidates = np.flatnonzero(results['rank_test_score'] == i)
for candidate in candidates:
print("Model with rank: {0}".format(i))
print("Mean validation score: {0:.3f} (std: {1:.3f})".format(
results['mean_test_score'][candidate],
results['std_test_score'][candidate]))
print("Parameters: {0}".format(results['params'][candidate]))
print("")
param_dist = {'C': scipy.stats.expon(scale=100),
'gamma': scipy.stats.expon(scale=.1),
'kernel': ['rbf', 'linear', 'poly'],
'class_weight':['balanced', None]}
svm_classifier2 = svm.SVC(random_state=42)
n_iter_search = 20
random_search = RandomizedSearchCV(svm_classifier2,
param_distributions=param_dist,
n_iter=n_iter_search,
cv=5,
n_jobs=-1)
start = time()
print("Hyperparameter tuning...")
random_search.fit(X_train, y_train)
print("RandomizedSearchCV took %.2f seconds for %d candidates"
" parameter settings." % ((time() - start), n_iter_search))
report(random_search.cv_results_)
print("============ EVALUATION on test set:")
print(accuracy_score(y_test, random_search.best_estimator_.predict(X_test)))
# -
| notebooks/SVM_SourceCode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import boto3
import os, random, string
from tqdm.notebook import tqdm
def random_id():
return ''.join(random.choice(string.ascii_uppercase + string.digits) for _ in range(10))
sqs = boto3.client('sqs')
queue_url = sqs.get_queue_url(QueueName='SomeRegularQueue')['QueueUrl']
fifo_queue_url = sqs.get_queue_url(QueueName='SomeFifoQueue.fifo')['QueueUrl']
# ## Create 5000 messages on each queue
for i in tqdm(range(0, 500)):
sqs.send_message_batch(QueueUrl=queue_url, Entries=[
{
'Id': f'Regular{i}_{j}',
'MessageBody': f'Hi regular queue {i}/{j}'
} for j in range(0, 10)
])
# +
# Send messages to the FIFO queue with random groups so we can have more than one in flight at a time
# -
for i in tqdm(range(0, 500)):
sqs.send_message_batch(QueueUrl=fifo_queue_url, Entries=[
{
'Id': f'Regular{i}_{j}',
'MessageBody': f'Hi regular queue {i}/{j}',
'MessageGroupId': random_id(),
'MessageDeduplicationId': f'{i}_{j}'
} for j in range(0, 10)
])
# ## Receive Messages
# Use a visibility timeout to ensure messages are regarded as 'in-flight', triggering the threshold for
# inflight messages percent for the test project.
sqs.receive_message(QueueUrl=queue_url, VisibilityTimeout=10)
message_ids_regular = []
sqs.receive_message(QueueUrl=queue_url, VisibilityTimeout=10)
for _ in tqdm(range(0, 1201)):
msg_id = sqs.receive_message(QueueUrl=queue_url, VisibilityTimeout=1000)['Messages'][0]['MessageId']
message_ids_regular.append(msg_id)
sqs.receive_message(QueueUrl=fifo_queue_url, VisibilityTimeout=10)
for _ in tqdm(range(0, 181)):
resp = sqs.receive_message(QueueUrl=fifo_queue_url, VisibilityTimeout=1000)
msg_id = resp['Messages'][0]['MessageId']
message_ids_fifo.append(msg_id)
# ## Clean up queues
for url in tqdm([queue_url, fifo_queue_url]):
sqs.purge_queue(QueueUrl=url)
| testing/trigger_sqs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as numpy
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
train = pd.read_csv('titanic_train.csv')
train.head()
# ### Missing Data
train.isnull()
sns.heatmap(train.isnull(),yticklabels=False,cbar=False,cmap='viridis')
# Roughly 20 percent of the Age data is missing. The proportion of Age missing is likely small enough for reasonable replacement with some form of imputation. Looking at the Cabin column, it looks like we are just missing too much of that data to do something useful with at a basic level. We'll probably drop this later, or change it to another feature like "Cabin Known: 1 or 0"
sns.set_style('whitegrid')
sns.countplot(x='Survived',data=train)
sns.set_style('whitegrid')
sns.countplot(x='Survived',hue='Sex',data=train,palette='RdBu_r')
sns.set_style('whitegrid')
sns.countplot(x='Survived',hue='Pclass',data=train,palette='rainbow')
sns.distplot(train['Age'].dropna(),kde=False,color='darkred',bins=40)
train['Age'].hist(bins=30,color='darkred',alpha=0.3)
sns.countplot(x='SibSp',data=train)
train['Fare'].hist(color='green',bins=40,figsize=(8,4))
# ### Cufflinks for plots
import cufflinks as cf
cf.go_offline()
train['Fare'].iplot(kind='hist',bins=30,color='green')
# ### Data Cleaning
# We want to fill in missing age data instead of just dropping the missing age data rows. One way to do this is by filling in the mean age of all the passengers (imputation). However we can be smarter about this and check the average age by passenger class.
plt.figure(figsize=(12, 7))
sns.boxplot(x='Pclass',y='Age',data=train,palette='winter')
# We can see the wealthier passengers in the higher classes tend to be older, which makes sense. We'll use these average age values to impute based on Pclass for Age.
def impute_age(cols):
Age = cols[0]
Pclass = cols[1]
if pd.isnull(Age):
if Pclass == 1:
return 37
elif Pclass == 2:
return 29
else:
return 24
else:
return Age
# Now apply that function!
train['Age'] = train[['Age','Pclass']].apply(impute_age,axis=1)
# Now let's check that heat map again!
sns.heatmap(train.isnull(),yticklabels=False,cbar=False,cmap='viridis')
# Great! Let's go ahead and drop the Cabin column and the row in Embarked that is NaN.
train.drop('Cabin',axis=1,inplace=True)
train.head()
train.dropna(inplace=True)
# ### Converting Categorical Features¶
# We'll need to convert categorical features to dummy variables using pandas! Otherwise our machine learning algorithm won't be able to directly take in those features as inputs.
train.info()
pd.get_dummies(train['Embarked'],drop_first=True).head()
sex = pd.get_dummies(train['Sex'],drop_first=True)
embark = pd.get_dummies(train['Embarked'],drop_first=True)
train.drop(['Sex','Embarked','Name','Ticket'],axis=1,inplace=True)
train.head()
train = pd.concat([train,sex,embark],axis=1)
train.head()
| Basics/#7 Exploratory Data Analysis(EDA).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: cugraph_dev
# language: python
# name: cugraph_dev
# ---
# # Louvain Performance Benchmarking
#
# This notebook benchmarks performance improvement of running the Louvain clustering algorithm within cuGraph against NetworkX. The test is run over eight test networks (graphs) and then results plotted.
# <p><p>
#
#
# #### Notebook Credits
#
# Original Authors: <NAME>
# Last Edit: 06/10/2020
#
#
# #### Test Environment
#
# RAPIDS Versions: 0.15
#
# Test Hardware:
# GV100 32G, CUDA 10,0
# Intel(R) Core(TM) CPU i7-7800X @ 3.50GHz
# 32GB system memory
#
#
#
# #### Updates
# - moved loading ploting libraries to front so that dependencies can be checked before running algorithms
# - added edge values
# - changed timing to including Graph creation for both cuGraph and NetworkX. This will better represent end-to-end times
#
#
#
# #### Dependencies
# - RAPIDS cuDF and cuGraph version 0.6.0
# - NetworkX
# - Matplotlib
# - Scipy
# - data prep script run
#
#
#
# #### Note: Comparison against published results
#
#
# The cuGraph blog post included performance numbers that were collected over a year ago. For the test graphs, int32 values are now used. That improves GPUs performance. Additionally, the initial benchamrks were measured on a P100 GPU.
#
# This test only comparse the modularity scores and a success is if the scores are within 15% of each other. That comparison is done by adjusting the NetworkX modularity score and then verifying that the cuGraph score is higher.
#
# cuGraph did a full validation of NetworkX results against cuGraph results. That included cross-validation of every cluster. That test is very slow and not included here
# ## Load the required libraries
# Import needed libraries
import time
import cugraph
import cudf
import os
# NetworkX libraries
import networkx as nx
from scipy.io import mmread
# NetworkX libraries
try:
import community
except ModuleNotFoundError:
os.system('pip install python-louvain')
import community
try:
import matplotlib
except ModuleNotFoundError:
os.system('pip install matplotlib')
# Loading plotting libraries
import matplotlib.pyplot as plt; plt.rcdefaults()
import numpy as np
# ### Define the test data
# Test File
data = {
'preferentialAttachment' : './data/preferentialAttachment.mtx',
'caidaRouterLevel' : './data/caidaRouterLevel.mtx',
'coAuthorsDBLP' : './data/coAuthorsDBLP.mtx',
'dblp' : './data/dblp-2010.mtx',
'citationCiteseer' : './data/citationCiteseer.mtx',
'coPapersDBLP' : './data/coPapersDBLP.mtx',
'coPapersCiteseer' : './data/coPapersCiteseer.mtx',
'as-Skitter' : './data/as-Skitter.mtx'
}
# ### Define the testing functions
# Read in a dataset in MTX format
def read_mtx_file(mm_file):
print('Reading ' + str(mm_file) + '...')
M = mmread(mm_file).asfptype()
return M
# Run the cuGraph Louvain analytic (using nvGRAPH function)
def cugraph_call(M):
t1 = time.time()
# data
gdf = cudf.DataFrame()
gdf['src'] = M.row
gdf['dst'] = M.col
# create graph
G = cugraph.Graph()
G.from_cudf_edgelist(gdf, source='src', destination='dst', renumber=False)
# cugraph Louvain Call
print(' cuGraph Solving... ')
df, mod = cugraph.louvain(G)
t2 = time.time() - t1
return t2, mod
# Run the NetworkX Louvain analytic. THis is done in two parts since the modularity score is not returned
def networkx_call(M):
nnz_per_row = {r: 0 for r in range(M.get_shape()[0])}
for nnz in range(M.getnnz()):
nnz_per_row[M.row[nnz]] = 1 + nnz_per_row[M.row[nnz]]
for nnz in range(M.getnnz()):
M.data[nnz] = 1.0/float(nnz_per_row[M.row[nnz]])
M = M.tocsr()
if M is None:
raise TypeError('Could not read the input graph')
if M.shape[0] != M.shape[1]:
raise TypeError('Shape is not square')
t1 = time.time()
# Directed NetworkX graph
Gnx = nx.Graph(M)
# Networkx
print(' NetworkX Solving... ')
parts = community.best_partition(Gnx)
# Calculating modularity scores for comparison
mod = community.modularity(parts, Gnx)
t2 = time.time() - t1
return t2, mod
# ### Run the benchmarks
# +
# Loop through each test file and compute the speedup
perf = []
names = []
time_cu = []
time_nx = []
#init libraries by doing quick pass
v = './data/preferentialAttachment.mtx'
M = read_mtx_file(v)
trapids = cugraph_call(M)
del M
for k,v in data.items():
M = read_mtx_file(v)
tr, modc = cugraph_call(M)
tn, modx = networkx_call(M)
speedUp = (tn / tr)
names.append(k)
perf.append(speedUp)
time_cu.append(tr)
time_nx.append(tn)
# mod_delta = (0.85 * modx)
print(str(speedUp) + "x faster => cugraph " + str(tr) + " vs " + str(tn))
# -
# ### plot the output
# +
# %matplotlib inline
y_pos = np.arange(len(names))
plt.bar(y_pos, perf, align='center', alpha=0.5)
plt.xticks(y_pos, names)
plt.ylabel('Speed Up')
plt.title('Performance Speedup: cuGraph vs NetworkX')
plt.xticks(rotation=90)
plt.show()
# -
# # Dump the raw stats
perf
time_cu
time_nx
# ___
# Copyright (c) 2020, <NAME>.
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
# ___
| notebooks/cugraph_benchmarks/louvain_benchmark.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''11-film-gex'': conda)'
# name: python3
# ---
# +
import numpy as np
import pandas as pd
from train import read
from pathlib import Path
import joblib
import pyarrow.dataset as ds
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
# CUSTOM
from eval import get_tx, combine_features
# VIZ
size=15
params = {'legend.fontsize': 'large',
'figure.figsize': (4,4),
'axes.labelsize': size,
'axes.titlesize': size,
'xtick.labelsize': size*0.75,
'ytick.labelsize': size*0.75,
'axes.titlepad': 25}
plt.rcParams.update(params)
from matplotlib_inline.backend_inline import set_matplotlib_formats
set_matplotlib_formats('retina')
# +
from scipy.optimize import curve_fit, OptimizeWarning
from scipy.integrate import quad
from eval import *
def plot_drc(params, raw_data):
cpd_order = params['broad_cpd_id'].unique()
results = []
for group,data in params.groupby(['stripped_cell_line_name', 'broad_cpd_id']):
raw_data_sample = raw_data[(raw_data['stripped_cell_line_name']==group[0]) &
(raw_data['broad_cpd_id']==group[1])].copy()
raw_data_sample.loc[:,'IC50'] = data['IC50'].unique().item()
popt = tuple(data[['H', 'EC50', 'Emin', 'Emax']].values.reshape(-1))
raw_data_sample['drc'] = [ll4(x, *popt) for x in raw_data_sample['cpd_conc_umol'].values]
results.append(raw_data_sample)
results = pd.concat(results, ignore_index=True)
results['stripped_cell_line_name'] = results['stripped_cell_line_name'] + "\n IC50=" + results['IC50'].round(2).astype(str)
# order cpds as categories to output plots by decreasing IC50 separation
results = results.set_index('broad_cpd_id').loc[cpd_order].reset_index()
category_cpd = pd.api.types.CategoricalDtype(categories=cpd_order, ordered=True)
results['broad_cpd_id'] = results['broad_cpd_id'].astype(category_cpd)
for group,data in results.groupby(['broad_cpd_id']):
# Annotations
cpd_id = group
cpd_name = data['cpd_name'].unique()[0]
cpd_mech = data['target_or_activity_of_compound'].unique()[0]
# Viz
ax = sns.lineplot(data=data, x='log_cpd_conc_mol', y='drc', hue='stripped_cell_line_name', alpha=0.8, legend=True)
sns.lineplot(data=data, x='log_cpd_conc_mol', y='cpd_pred_pv', hue='stripped_cell_line_name', marker='o', linestyle='', err_style='bars', alpha=0.3, legend=False)
plt.xticks(np.arange(-9, -3, 1))
plt.legend(bbox_to_anchor=(1.07, 1), loc=2, borderaxespad=0., frameon=False, prop={'size':10})
plt.xlabel('Log Concentration (Mol)')
plt.ylabel('Predicted Percent Viability')
plt.subplots_adjust(top=0.8)
plt.suptitle(f'{cpd_id}\n{cpd_name}\n{cpd_mech}', fontsize=12)
plt.show()
def heatmap_top_cpds(df, lines, k, n=20, plot=True):
cols = []
for c in lines:
rows = []
for r in lines:
r_cpds = set(df.loc[r].sort_values(by='IC50')['broad_cpd_id'].unique()[:n])
c_cpds = set(df.loc[c].sort_values(by='IC50')['broad_cpd_id'].unique()[:n])
# jaccard score
rows.append(len(r_cpds.intersection(c_cpds)) / len(r_cpds.union(c_cpds)))
cols.append(rows)
names = [f"{a} (k={b})" for a,b in zip(lines,k)]
inter_df = pd.DataFrame(cols,
index=names,
columns=names)
if plot:
ax = sns.heatmap(inter_df, vmin=0, annot=True, cmap="YlGnBu", annot_kws={"size": 6})
plt.title(f"Top {n} compound intersection by IC50")
return inter_df
def rank_proportion(drc, targets, query_line, k):
rank = drc[drc['stripped_cell_line_name']==query_line].sort_values(by='IC50', ascending=True)
rank = pd.DataFrame(rank['broad_cpd_id'].unique(), columns=['broad_cpd_id'])
#rank = rank.reset_index().rename(columns={'index':'Rank'})
rank['Normalized Rank'] = rank.index / len(rank)
rank['isTarget'] = rank['broad_cpd_id'].isin(targets)
rank['Proportion'] = [rank['isTarget'][:i].sum()/len(targets) for i in range(1,len(rank)+1)]
rank['source'] = f"{query_line} (k={k})"
return rank
def compute_diff(params, base_cl, test_cl):
base = params[params['stripped_cell_line_name']==base_cl]
test = params[params['stripped_cell_line_name']==test_cl]
comb = base.merge(test, on='broad_cpd_id', suffixes=(f"-{base_cl.split('-')[-1]}", f"-{test_cl.split('-')[-1]}"))
comb['IC50_diff'] = comb[f'IC50-Par'] - comb[f"IC50-{test_cl.split('-')[-1]}"]
comb['EC50_diff'] = comb[f'EC50-Par'] - comb[f"EC50-{test_cl.split('-')[-1]}"]
comb['AUC_diff'] = comb[f'AUC-Par'] - comb[f"AUC-{test_cl.split('-')[-1]}"]
# comb = comb[(comb['IC50_diff'] < 10) & (comb['IC50_diff'] > -10)]
# comb = comb[(comb['EC50_diff'] < 100) & (comb['EC50_diff'] > -100)]
return comb.sort_values(by='AUC_diff', ascending=False)
# -
# # Read Data
HCCb_lines = ('HCC1806-Par', 'HCC1806-LM2b')
HCCc_lines = ('HCC1806-Par', 'HCC1806-LM2c')
MDA_lines = ('MDA-Par', 'MDA-LM2')
SW_lines = ('SW480-Par', 'SW480-LvM2')
eval_lines = ('HCC1806-Par', 'HCC1806-LM2b', 'HCC1806-LM2c', 'MDA-Par', 'MDA-LM2', 'SW480-Par', 'SW480-LvM2')
# +
# Get training & testing dose/response metrics
preds = pd.read_csv(f"../data/hani-metastatic/processed/predictions.tsv", sep="\t")
all_df = combine(Path("../data/processed"), preds)
# filter negativ PV
all_df = all_df[all_df['cpd_pred_pv'] >= 0]
# add small eps to reduce log transform errors
all_df['cpd_pred_pv'] = all_df['cpd_pred_pv'] + 1e-32
all_df['log_cpd_conc_mol'] = np.log10(all_df['cpd_conc_umol'] / 1e6)
# Add cpd annotations
ctrp = pd.read_csv("../data/drug_screens/CTRP/v20.meta.per_compound.txt", sep="\t", usecols=['broad_cpd_id', 'cpd_name', 'cpd_status', 'target_or_activity_of_compound', 'gene_symbol_of_protein_target'])
all_df['cpd_name'] = all_df['broad_cpd_id'].map(ctrp.set_index('broad_cpd_id')['cpd_name'])
all_df['target_or_activity_of_compound'] = all_df['broad_cpd_id'].map(ctrp.set_index('broad_cpd_id')['target_or_activity_of_compound'], na_action="unknown")
# -
# # Batch Analysis
# +
data_path = Path("../data/processed")
gene_cols = joblib.load(data_path.joinpath("gene_cols.pkl"))
data_ds = ds.dataset(data_path.joinpath('data.feather'), format='feather')
cols = list(gene_cols) + ['stripped_cell_line_name']
data_df = data_ds.to_table(columns=cols).to_pandas()
data_df = data_df.drop_duplicates(subset=['stripped_cell_line_name']).set_index('stripped_cell_line_name')
# Warning must standardize the original data as is done with model training
data_df = pd.DataFrame(StandardScaler().fit_transform(data_df), index=data_df.index, columns=data_df.columns)
data_df.shape
# +
eval_path = Path("../data/hani-metastatic/processed")
eval_df = pd.read_csv(eval_path.joinpath("eval_data.tsv"), sep="\t")
eval_df = eval_df[cols]
eval_df = eval_df.drop_duplicates(subset=['stripped_cell_line_name']).set_index('stripped_cell_line_name')
eval_df.shape
# +
if not np.array_equal(data_df.columns, eval_df.columns):
raise Exception("Check data alignment")
comb_df = pd.concat([data_df, eval_df])
comb_gene_pca = pd.DataFrame(PCA(n_components=2).fit_transform(comb_df),
index=np.concatenate([data_df.index, eval_df.index]),
columns=["PC_1", "PC_2"])
comb_gene_pca['Source'] = np.concatenate([np.repeat("Train", len(data_df)), eval_df.index])
# viz
ax = sns.scatterplot(data=comb_gene_pca[comb_gene_pca['Source']=='Train'], x='PC_1', y='PC_2', hue='Source', palette='pastel', alpha=0.7)
sns.scatterplot(data=comb_gene_pca[comb_gene_pca['Source']!='Train'], x='PC_1', y='PC_2', hue='Source', ax=ax)
sns.despine()
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0., frameon=False)
plt.title("L1000 Gene Features\nBatch Effect Analysis", size=14)
# -
# # Dendrogram Test Plots
ccle_meta = pd.read_csv("../data/cellular_models/sample_info.csv", sep=",")
ccle_meta.head()
eval_primary = {'HCC1806-Par': 'Colon/Colorectal Cancer',
'HCC1806-LM2b': 'Colon/Colorectal Cancer',
'HCC1806-LM2c': 'Colon/Colorectal Cancer',
'MDA-Par': 'Breast Cancer',
'MDA-LM2': 'Breast Cancer',
'SW480-Par': 'Breast Cancer',
'SW480-LvM2': 'Breast Cancer'}
ccle_primary = ccle_meta.set_index('stripped_cell_line_name')['primary_disease'].to_dict()
all_primary = {**ccle_primary, **eval_primary}
tab20b_cmap = sns.color_palette("tab20b", as_cmap=True)
tab20c_cmap = sns.color_palette("tab20c", as_cmap=True)
from matplotlib.colors import ListedColormap
from matplotlib.patches import Rectangle
large_cmap = ListedColormap(list(tab20b_cmap.colors) + list(tab20c_cmap.colors), name="large_cmap")
large_cmap
# +
row_cl = comb_df.index.to_series()
row_primary = row_cl.map(all_primary)
lut = dict(zip(row_primary.unique(), list(large_cmap.colors)[:row_primary.nunique()]))
row_colors = row_primary.map(lut)
row_colors.name = "" # remove the name of the row_colors
g = sns.clustermap(comb_df, row_colors=row_colors, col_cluster=False, dendrogram_ratio=(0.95, 0.03), colors_ratio=0.04, figsize=(6, 8))
g.ax_heatmap.remove() # remove the heatmap
g.cax.remove() # remove the color bar
g.fig.legend(handles=[Rectangle((0, 0), 0, 0, color=val, label=key) for key, val in lut.items()],
title='Primary Disease', loc='lower left', bbox_to_anchor=[0.8, 0.1])
plt.show()
# +
hcc_primary = all_primary.copy()
HCC_lines = ['HCC1806', 'HCC1806-Par', 'HCC1806-LM2b', 'HCC1806-LM2c']
for line in hcc_primary:
if line in HCC_lines:
hcc_primary[line] = line
elif hcc_primary[line] != 'Colon/Colorectal Cancer':
hcc_primary[line] = 'Other'
set2 = sns.color_palette("Set2", as_cmap=True)
row_cl = comb_df.index.to_series()
row_primary = row_cl.map(hcc_primary)
lut = dict(zip(row_primary.unique(), [list(set2.colors)[-1],] + list(set2.colors)[1:row_primary.nunique()-1] ))
row_colors = row_primary.map(lut)
row_colors.name = "" # remove the name of the row_colors
g = sns.clustermap(comb_df, row_colors=row_colors, col_cluster=False, dendrogram_ratio=(0.95, 0.03), colors_ratio=0.04, figsize=(6, 8))
g.ax_heatmap.remove() # remove the heatmap
g.cax.remove() # remove the color bar
g.fig.legend(handles=[Rectangle((0, 0), 0, 0, color=val, label=key) for key, val in lut.items()],
title='Primary Disease', loc='lower left', bbox_to_anchor=[0.8, 0.3])
plt.show()
# -
# # IC50 Analysis
# +
drc = pd.read_csv("../data/hani-metastatic/processed/drc_parameters.tsv", sep="\t")
# IC50 log transform
drc['IC50'] = np.log10(drc['IC50'] / 1e6)
# Pre-filterting
drc[~drc['stripped_cell_line_name'].isin(eval_lines)].groupby('broad_cpd_id').size().hist(bins=30, alpha=0.5, label='Pre-filtering')
# Fraction of failed DRC models
drc.shape
drc['IC50'].isna().sum() / len(drc)
drc['FUNC'].value_counts()
# drc = drc.dropna()
# drc['FUNC'].value_counts()
# # Filter by Hill coefficient & EC50
# eps = 0.8
# full_drc_shape = drc.shape[0]
# drc = drc[(drc['H']>1-eps) & (drc['H']<1+eps)]
# Filter by EC50
full_drc_shape = drc.shape[0]
drc = drc[drc['EC50']>=1e-3]
drc = drc[drc['EC50']<=300]
print(f"Fraction of DRC remaining after filtering: {drc.shape[0] / full_drc_shape:.3f}")
# Post filtering
drc[~drc['stripped_cell_line_name'].isin(eval_lines)].groupby('broad_cpd_id').size().hist(bins=30, alpha=0.5, label='Post-filtering')
plt.xlabel("# cell lines")
plt.ylabel('Frequency')
plt.title("Number of cell lines tested per compound")
plt.legend()
# +
from sklearn.neighbors import NearestNeighbors
nbrs = NearestNeighbors(n_neighbors=len(comb_df), algorithm='ball_tree').fit(comb_df)
distances, indices = nbrs.kneighbors(comb_df)
indices_df = pd.DataFrame(indices, index=comb_df.index)
distances_df = pd.DataFrame(distances, index=comb_df.index)
# +
cl = 'HCC1806-Par'
query_lines = eval_lines
n = 20
k = ['NA']*len(query_lines)
# Heatmap
heatmap_top_cpds(drc.set_index('stripped_cell_line_name'), lines=query_lines, k=k, n=n)
plt.show()
# rank top cpd recovery
targets = drc[drc['stripped_cell_line_name']==cl].sort_values(by='IC50')['broad_cpd_id'][:n].values
ranks = pd.concat([rank_proportion(drc, targets, ql, i) for ql,i in zip(query_lines,k)], ignore_index=True)
sns.lineplot(data=ranks, x='Normalized Rank', y='Proportion', hue='source', alpha=0.7)
# Put the legend out of the figure
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title(f"Cumulative Proportion of {cl} Ranked Compounds")
plt.show()
# -
top20_cpd_set = (set(drc[(drc['stripped_cell_line_name']==cl)].sort_values(by='IC50').head(20)['broad_cpd_id'].values) for cl in eval_lines)
top20_cpd_set = set.union(*top20_cpd_set)
len(top20_cpd_set)
drc[(~drc['stripped_cell_line_name'].isin(eval_lines))]['IC50'].hist(bins=50, alpha=0.5)
df = drc[(~drc['stripped_cell_line_name'].isin(eval_lines)) & (drc['broad_cpd_id'].isin(top20_cpd_set))].groupby('broad_cpd_id')['IC50'].mean()
df.head()
plt.vlines(df, ymin=0, ymax=20000, color='r', linewidth=0.2)
plt.xlabel("Compound Avg IC50")
plt.ylabel("Frequency")
plt.title("Top Compound Set \nAverage IC50 in Training Data")
# +
cl = 'HCC1806-Par'
n = 20
k = np.arange(0,5,1)
k = list(k) + list(np.geomspace(5, len(indices)-1, num=5, endpoint=True, dtype=np.int))
# Heatmap
query_idx = indices_df.loc[cl][k]
query_lines = indices_df.iloc[query_idx.values].index
query_distances = distances_df.loc[cl][k]
print(dict(zip(query_lines, query_distances)))
heatmap_top_cpds(drc.set_index('stripped_cell_line_name'), lines=query_lines, k=k, n=n)
plt.show()
# rank top cpd recovery
targets = drc[drc['stripped_cell_line_name']==cl].sort_values(by='IC50')['broad_cpd_id'][:n].values
ranks = pd.concat([rank_proportion(drc, targets, ql, i) for ql,i in zip(query_lines,k)], ignore_index=True)
sns.lineplot(data=ranks, x='Normalized Rank', y='Proportion', hue='source', alpha=0.7)
# Put the legend out of the figure
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title(f"Cumulative Proportion of {cl} Ranked Compounds")
plt.show()
# +
cl = 'MDA-Par'
n = 20
k = np.arange(0,5,1)
k = list(k) + list(np.geomspace(5, len(indices)-1, num=5, endpoint=True, dtype=np.int))
# Heatmap
query_idx = indices_df.loc[cl][k]
query_lines = indices_df.iloc[query_idx.values].index
query_distances = distances_df.loc[cl][k]
print(dict(zip(query_lines, query_distances)))
heatmap_top_cpds(drc.set_index('stripped_cell_line_name'), lines=query_lines, k=k, n=n)
plt.show()
# rank top cpd recovery
targets = drc[drc['stripped_cell_line_name']==cl].sort_values(by='IC50')['broad_cpd_id'][:n].values
ranks = pd.concat([rank_proportion(drc, targets, ql, i) for ql,i in zip(query_lines,k)], ignore_index=True)
sns.lineplot(data=ranks, x='Normalized Rank', y='Proportion', hue='source', alpha=0.7)
# Put the legend out of the figure
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title(f"Cumulative Proportion of {cl} Ranked Compounds")
plt.show()
# +
cl = 'SW480-Par'
n = 20
k = np.arange(0,5,1)
k = list(k) + list(np.geomspace(5, len(indices)-1, num=5, endpoint=True, dtype=np.int))
# Heatmap
query_idx = indices_df.loc[cl][k]
query_lines = indices_df.iloc[query_idx.values].index
query_distances = distances_df.loc[cl][k]
print(dict(zip(query_lines, query_distances)))
heatmap_top_cpds(drc.set_index('stripped_cell_line_name'), lines=query_lines, k=k, n=n)
plt.show()
# rank top cpd recovery
targets = drc[drc['stripped_cell_line_name']==cl].sort_values(by='IC50')['broad_cpd_id'][:n].values
ranks = pd.concat([rank_proportion(drc, targets, ql, i) for ql,i in zip(query_lines,k)], ignore_index=True)
sns.lineplot(data=ranks, x='Normalized Rank', y='Proportion', hue='source', alpha=0.7)
# Put the legend out of the figure
plt.legend(bbox_to_anchor=(1.05, 1), loc=2, borderaxespad=0.)
plt.title(f"Cumulative Proportion of {cl} Ranked Compounds")
plt.show()
# +
# def cooks_distance(func, x, y):
# p = func.__code__.co_argcount - len(func.__defaults__) - 1
# popt, pcov = curve_fit(func, x, y)
# primary_yhat = func(x, *popt)
# cooks_dist = []
# for i in range(len(x)):
# popt, pcov = curve_fit(func, np.delete(x, i), np.delete(y, i))
# yhat = func(x, *popt)
# diff = primary_yhat - yhat
# mse = np.mean(diff**2)
# cd = np.sum(diff**2) / (p * mse)
# cooks_dist.append(cd)
# return np.array(cooks_dist)
# -
# # Differenctial Efficacy
from rdkit import Chem
from rdkit.Chem import Draw
from rdkit.Chem.Draw import IPythonConsole
smiles = 'C1=CC(=C(C=C1C2=C(C(=O)C3=C(C=C(C=C3O2)O)O)O)O)O'
m = Chem.MolFromSmiles(smiles)
Draw.MolToMPL(m)
# example
plot_drc(drc[(drc['stripped_cell_line_name'].isin(eval_lines)) & (drc['broad_cpd_id']=='BRD-K55696337')], all_df)
plot_drc(drc[drc['stripped_cell_line_name'].isin(HCCb_lines)].sort_values(by='IC50').head(10), all_df)
for lines in [HCCb_lines, HCCc_lines, MDA_lines, SW_lines]:
print(f"{lines}")
df = compute_diff(drc, *lines).sort_values(by='IC50_diff', ascending=False)
cpds = df.head(5)['broad_cpd_id'].values
df = drc[(drc['stripped_cell_line_name'].isin(lines)) & (drc['broad_cpd_id'].isin(cpds))].set_index('broad_cpd_id').loc[cpds].reset_index()
plot_drc(df, all_df)
df = compute_diff(drc, *MDA_lines).sort_values(by='IC50_diff', ascending=False)
cpds = df.head(20)['broad_cpd_id'].values
df = drc[(drc['stripped_cell_line_name'].isin(lines)) & (drc['broad_cpd_id'].isin(cpds))].set_index('broad_cpd_id').loc[cpds].reset_index()
plot_drc(df, all_df)
| notebooks/2021.09.08_eval.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
using Pkg,Statistics,Random,LinearAlgebra,Printf,GZip,Knet,Plots,Images
L=128;
N_train=2;
p_train=rand(N_train,1);
x_real = rand(N_train,L);
train_real = [x_real p_train];
function threshold_train(x)
y = copy(x)
for i=1:length(x[:,1])
for j = 1:length(x[i,:])-1
y[i,j] = x[i,j] > x[i,end] ? 0 : 1
end
end
return y
end
train_data = threshold_train(train_real)
for i=1:length(train_data[:,1])
push!(lossarr,train_data[i,end]-dot(w[1],train_data[i,1:end-1])+w[2]);
end
w = [0.1*rand(1,L), 0.0]
function loss(w,train_data)
lossarr=zeros(0);
for i=1:length(train_data[:,1])
push!(lossarr,train_data[i,end]-dot(w[1],train_data[i,1:end-1])+w[2]);
end
return mean(abs2,lossarr)
end
lossgradient = grad(loss)
loss(w,train_data)
lossgradient(w,train_data)
function train!(w;lr=0.1)
dw = lossgradient(w,train_data);
for i = 1:length(w)
w[i] -= lr*dw[i];
end
return w
end
| archive/20180913-p1-guess_p.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (phathom)
# language: python
# name: phathom
# ---
# +
import neuroglancer
ip_address = '172.16.31.10'
port = 80
# Use this in IPython to allow external viewing
# neuroglancer.set_server_bind_address(bind_address='192.168.158.128',
# bind_port=80)
if ip_address is not None:
neuroglancer.set_server_bind_address(bind_address=ip_address,
bind_port=port)
viewer = neuroglancer.Viewer()
viewer
# +
from nuggt.utils import ngutils
import numpy as np
def plot_image(img, viewer, layer, shader):
with viewer.txn() as txn:
source = neuroglancer.LocalVolume(img.astype(np.float32))
txn.layers[layer] = neuroglancer.ImageLayer(source=source, shader=shader)
def plot_fixed(fixed_img, viewer, scaling):
fixed_shader = ngutils.red_shader % scaling
plot_image(fixed_img, viewer, 'fixed', fixed_shader)
def plot_moving(moving_img, viewer, scaling):
moving_shader = ngutils.green_shader % scaling
plot_image(moving_img, viewer, 'moving', moving_shader)
def plot_both(fixed_img, moving_img, viewer, scaling):
plot_fixed(fixed_img, viewer, scaling)
plot_moving(moving_img, viewer, scaling)
# +
import zarr
fixed_path = 'fixed.zarr'
nonrigid_path = 'nonrigid.zarr'
fixed_img = zarr.open(fixed_path, mode='r')
nonrigid_img = zarr.open(nonrigid_path, mode='r')
# -
plot_both(fixed_img, nonrigid_img, viewer, 0.001)
| notebooks/.ipynb_checkpoints/neuroglancer_viz-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: vae
# language: python
# name: vae
# ---
# +
import os
import torch
import umap.plot
import numpy as np
import pandas as pd
from mord import OrdinalRidge
from sklearn.linear_model import Ridge, RidgeClassifier
from sklearn.linear_model import Ridge
from sklearn.metrics import accuracy_score, mean_squared_error, r2_score, precision_recall_fscore_support, matthews_corrcoef, confusion_matrix
from sklearn.neural_network import MLPClassifier
from sklearn.svm import LinearSVC
# -
ge = np.load("/home/bram/jointomicscomp/data/GE.npy")
cancerTypes = np.load("/home/bram/jointomicscomp/data/cancerTypes.npy")
cancerType = np.load("/home/bram/jointomicscomp/data/cancerType.npy")
cancer_type_labels = cancerTypes[cancerType.astype(int)]
# +
# From baseline
def evaluate_classification(y_true, y_pred):
# returns accuracy, precision, recall, f1, mcc, confusion_matrix
print(y_true.dtype, y_pred.dtype)
acc = accuracy_score(y_true, y_pred)
pr, rc, f1, _ = precision_recall_fscore_support(y_true, y_pred)
mcc = matthews_corrcoef(y_true, y_pred)
confMat = confusion_matrix(y_true, y_pred)
return [acc, pr, rc, f1, mcc, confMat]
# +
print("RAW DATA: Baseline predictions")
alphas = np.array([1e-4, 1e-3, 1e-2, 1e-1, 0.5, 1.0, 2.0, 5.0, 10., 20.])
# if criterion == 'acc':
# ind = 0
# elif criterion == 'pr':
# ind = 1
# elif criterion == 'rc':
# ind = 2
# elif criterion == 'f1':
# ind = 3
# else:
# assert criterion == 'mcc'
# ind = 4
latent_train = np.load("/home/bram/jointomicscomp/data/BRCA/BR_GE_train.npy")
latent_val = np.load("/home/bram/jointomicscomp/data/BRCA/BRCA_GE_valid.npy")
latent_test = np.load("/home/bram/jointomicscomp/data/BRCA/BRCA_GE_test.npy")
y_train = np.load("/home/bram/jointomicscomp/data/{}/{}_train_stageType.npy".format("BRCA", "BRCA"))
y_val = np.load("/home/bram/jointomicscomp/data/{}/{}_valid_stageType.npy".format("BRCA", "BRCA"))
y_test = np.load("/home/bram/jointomicscomp/data/{}/{}_test_stageType.npy".format("BRCA", "BRCA"))
validationPerformance = np.zeros(alphas.shape[0])
models = []
for i, a in enumerate(alphas):
model = model = RidgeClassifier(alpha=a, fit_intercept=True, normalize=False, random_state=1)
# model = Ridge(alpha=a, fit_intercept=True, normalize=False, random_state=1)
# train
model.fit(latent_train, y_train)
# save so that we don't have to re-train
models.append(model)
# evaluate using user-specified criterion
validationPerformance[i] = evaluate_classification(y_val, model.predict(latent_val))[0]
bestModel = models[np.argmax(validationPerformance)]
predictions = bestModel.predict(latent_test).astype(int)
classifications1 = evaluate_classification(y_test, predictions)
print("Accuracy : ", classifications1[0])
print("Confusion matrix : \n", classifications1[5])
# -
test = bestModel.coef_
z = latent_test
for row in z:
row = np.multiply(row, test)
title = "LinearSVC Gene Expression Test set"
save_dir = "/home/bram/jointomicscomp/umaps"
save_file = "{}/UMAP {} Raw Data.png"\
.format(save_dir, "GE")
background = "white"
color_key_cmap = "Spectral"
# +
mapper = umap.UMAP(
n_neighbors=15,
min_dist=0.1,
n_components=2,
metric='euclidean'
).fit(z)
# -
p = umap.plot.points(mapper, labels=y_test, color_key_cmap=color_key_cmap, background=background)
umap.plot.plt.title(title)
# umap.plot.plt.legend()
umap.plot.plt.savefig(save_file, dpi=1600)
umap.plot.plt.show()
| src/util/Untitled-Copy3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Programming and Database Fundamentals for Data Scientists - EAS503
# ## Basic introduction to `Scikit-Learn`
# `scikit-learn` or `sklearn` is a `Python` machine learning library.
# %matplotlib inline
from matplotlib import pyplot as plt
from helper import *
import numpy as np
import warnings
warnings.filterwarnings("ignore")
# ### Outline
#
# * Classification
# * Model evaluation and selection
# * Transformers, pipelines and feature unions
# * Beyond building classifiers
#
# ## Classification
# Data comes as a finite learning set ${\cal L} = (X, y)$ where
# * Input samples are given as an array $X$ of shape `n_samples` $\times$ `n_features`, taking their values in ${\cal X}$;
# * Output values are given as an array $y$, taking _symbolic_ values in ${\cal Y}$.
#
# The goal of supervised classification is to build an estimator $\varphi: {\cal X} \mapsto {\cal Y}$ minimizing
#
# $$
# Err(\varphi) = \mathbb{E}_{X,Y}\{ \ell(Y, \varphi(X)) \}
# $$
#
# where $\ell$ is a loss function, e.g., the zero-one loss for classification $\ell_{01}(Y,\hat{Y}) = 1(Y \neq \hat{Y})$.
# ## Data
#
# - Input data = Numpy arrays or Scipy sparse matrices ;
# - Algorithms are expressed using high-level operations defined on matrices or vectors (similar to MATLAB) ;
# - Leverage efficient low-leverage implementations ;
# - Keep code short and readable.
# Generate data
from sklearn.datasets import make_blobs
X, y = make_blobs(n_samples=1000, centers=20, random_state=123)
labels = ["b", "r"]
y = np.take(labels, (y < 10))
print(X)
print(y[:5])
# Plot
plt.figure()
for label in labels:
mask = (y == label)
plt.scatter(X[mask, 0], X[mask, 1], c=label)
plt.xlim(-10, 10)
plt.ylim(-10, 10)
plt.show()
# ## A simple and unified API
#
# All learning algorithms in scikit-learn share a uniform and limited API consisting of complementary interfaces:
#
# - a `learn` interface for building and fitting models;
# - a `predict` interface for making predictions;
# - a `transform` interface for converting data.
#
# Goal: enforce a simple and consistent API to __make it trivial to swap or plug algorithms__.
# #### Creating the model
# Import the nearest neighbor class
from sklearn.neighbors import KNeighborsClassifier # Change this to try
# something else
# Set hyper-parameters, for controlling algorithm
clf = KNeighborsClassifier(n_neighbors=5)
# #### Learn a model from training data
clf.fit(X, y)
clf._tree
# #### Predicting
print(clf.predict(X[:5]))
print(clf.predict_proba(X[:5]))
plot_surface(clf,X,y)
# ### Exploring classifiers
from sklearn.tree import DecisionTreeClassifier
clf = DecisionTreeClassifier()
clf.fit(X, y)
plot_surface(clf, X, y)
from sklearn.ensemble import RandomForestClassifier
clf = RandomForestClassifier(n_estimators=500)
clf.fit(X, y)
plot_surface(clf, X, y)
from sklearn.linear_model import LogisticRegression
clf = LogisticRegression()
clf.fit(X, y)
plot_surface(clf, X, y)
from sklearn.svm import SVC
clf = SVC(kernel="rbf",probability=True)
clf.fit(X, y)
plot_surface(clf, X, y)
from sklearn.gaussian_process import GaussianProcessClassifier
clf = GaussianProcessClassifier()
clf.fit(X, y)
plot_surface(clf, X, y)
# ## Model evaluation and selection
# - Recall that we want to learn an estimator $\varphi$ minimizing the generalization error $Err(\varphi) = \mathbb{E}_{X,Y}\{ \ell(Y, \varphi(X)) \}$.
#
# - Problem: Since $P_{X,Y}$ is unknown, the generalization error $Err(\varphi)$ cannot be evaluated.
#
# - Solution: Use a proxy to approximate $Err(\varphi)$.
# ### Train Error
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import zero_one_loss
clf = KNeighborsClassifier(n_neighbors=1)
clf.fit(X, y)
print("Training error =", zero_one_loss(y, clf.predict(X)))
# ### Test Error
from sklearn.neighbors import KNeighborsClassifier
from sklearn.metrics import zero_one_loss
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y)
clf = KNeighborsClassifier(n_neighbors=5)
clf.fit(X_train, y_train)
print("Training error =", zero_one_loss(y_train, clf.predict(X_train)))
print("Test error =", zero_one_loss(y_test, clf.predict(X_test)))
# ### Cross-validation
# Issue:
# - When ${\cal L}$ is small, training on 70% of the data may lead to a model that is significantly different from a model that would have been learned on the entire set ${\cal L}$.
# - Yet, increasing the size of the training set (resp. decreasing the size of the test set), might lead to an inaccurate estimate of the generalization error.
#
# Solution: K-Fold cross-validation.
# - Split ${\cal L}$ into K small disjoint folds.
# - Train on K-1 folds, evaluate the test error one the held-out fold.
# - Repeat for all combinations and average the K estimates of the generalization error.
#
# +
from sklearn.model_selection import KFold
scores = []
for train, test in KFold(n_splits=5, random_state=42).split(X):
X_train, y_train = X[train], y[train]
X_test, y_test = X[test], y[test]
clf = KNeighborsClassifier(n_neighbors=5).fit(X_train, y_train)
scores.append(zero_one_loss(y_test, clf.predict(X_test)))
print("CV error = %f +-%f" % (np.mean(scores), np.std(scores)))
# -
# Shortcut
from sklearn.model_selection import cross_val_score
scores = cross_val_score(KNeighborsClassifier(n_neighbors=5), X, y,
cv=KFold(n_splits=5, random_state=42),
scoring="accuracy")
print("CV error = %f +-%f" % (1. - np.mean(scores), np.std(scores)))
# ## Metrics
# ### Default score
#
# Estimators come with a built-in default evaluation score
# * Accuracy for classification
# * R2 score for regression
y_train = (y_train == "r")
y_test = (y_test == "r")
clf = KNeighborsClassifier(n_neighbors=5)
clf.fit(X_train, y_train)
print("Default score =", clf.score(X_test, y_test))
# ### Accuracy
#
# Definition: The accuracy is the proportion of correct predictions.
from sklearn.metrics import accuracy_score
print("Accuracy =", accuracy_score(y_test, clf.predict(X_test)))
# ### Precision, recall and F-measure
#
# $$Precision = \frac{TP}{TP + FP}$$
# $$Recall = \frac{TP}{TP + FN}$$
# $$F = \frac{2 * Precision * Recall}{Precision + Recall}$$
from sklearn.metrics import precision_score
from sklearn.metrics import recall_score
from sklearn.metrics import fbeta_score
print("Precision =", precision_score(y_test, clf.predict(X_test)))
print("Recall =", recall_score(y_test, clf.predict(X_test)))
print("F =", fbeta_score(y_test, clf.predict(X_test), beta=1))
# ### ROC AUC
#
# Definition: Area under the curve of the false positive rate (FPR) against the true positive rate (TPR) as the decision threshold of the classifier is varied.
# +
from sklearn.metrics import get_scorer
roc_auc_scorer = get_scorer("roc_auc")
print("ROC AUC =", roc_auc_scorer(clf, X_test, y_test))
from sklearn.metrics import roc_curve
fpr, tpr, thresholds = roc_curve(y_test, clf.predict_proba(X_test)[:, 1])
plt.plot(fpr, tpr)
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.show()
# -
# ### Confusion matrix
#
# Definition: number of samples of class $i$ predicted as class $j$.
from sklearn.metrics import confusion_matrix
confusion_matrix(y_test, clf.predict(X_test))
# ### Model selection
#
# - Finding good hyper-parameters is crucial to control under- and over-fitting, hence achieving better performance.
# - The estimated generalization error can be used to select the best model.
#
# ### Under- and over-fitting
#
# - Under-fitting: the model is too simple and does not capture the true relation between X and Y.
# - Over-fitting: the model is too specific to the training set and does not generalize.
# +
from sklearn.model_selection import validation_curve
# Evaluate parameter range in CV
param_range = range(2, 200)
param_name = "max_leaf_nodes"
train_scores, test_scores = validation_curve(
DecisionTreeClassifier(), X, y,
param_name=param_name,
param_range=param_range, cv=5, n_jobs=-1)
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
# Plot parameter VS estimated error
plt.xlabel(param_name)
plt.ylabel("score")
plt.xlim(min(param_range), max(param_range))
plt.plot(param_range, 1. - train_scores_mean, color="red", label="Training error")
plt.fill_between(param_range,
1. - train_scores_mean + train_scores_std,
1. - train_scores_mean - train_scores_std,
alpha=0.2, color="red")
plt.plot(param_range, 1. - test_scores_mean, color="blue", label="CV error")
plt.fill_between(param_range,
1. - test_scores_mean + test_scores_std,
1. - test_scores_mean - test_scores_std,
alpha=0.2, color="blue")
plt.legend(loc="best")
# -
# Best trade-off
print("%s = %d, CV error = %f" % (param_name,
param_range[np.argmax(test_scores_mean)],
1. - np.max(test_scores_mean)))
# ### Hyper-parameter Search
# +
from sklearn.model_selection import GridSearchCV
grid = GridSearchCV(KNeighborsClassifier(),
param_grid={"n_neighbors": list(range(1, 100))},
scoring="accuracy",
cv=5, n_jobs=-1)
grid.fit(X, y) # Note that GridSearchCV is itself an estimator
print("Best score = %f, Best parameters = %s" % (1. - grid.best_score_,
grid.best_params_))
# -
# ## Transformers, pipelines and feature unions
#
# ### Transformers
#
# - Classification (or regression) is often only one or the last step of a long and complicated process;
# - In most cases, input data needs to be cleaned, massaged or extended before being fed to a learning algorithm;
# - For this purpose, Scikit-Learn provides the ``transformer`` API.
# +
# Load digits data
from sklearn.datasets import load_digits
from sklearn.model_selection import train_test_split
digits = load_digits()
X, y = digits.data, digits.target
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=0)
# Plot
sample_id = 42
plt.imshow(X[sample_id].reshape((8, 8)), interpolation="nearest", cmap=plt.cm.Blues)
plt.title("y = %d" % y[sample_id])
plt.show()
# -
# #### Scalers and other normalizers
#
# +
from sklearn.preprocessing import StandardScaler
tf = StandardScaler()
tf.fit(X_train, y_train)
Xt_train = tf.transform(X_train)
print("Mean (before scaling) =", np.mean(X_train))
print("Mean (after scaling) =", np.mean(Xt_train))
# Shortcut: Xt = tf.fit_transform(X)
# See also Binarizer, MinMaxScaler, Normalizer, ...
# -
# Scaling is critical for some algorithms
from sklearn.svm import SVC
clf = SVC()
print("Without scaling =", clf.fit(X_train, y_train).score(X_test, y_test))
print("With scaling =", clf.fit(tf.transform(X_train), y_train).score(tf.transform(X_test), y_test))
# #### Feature selection
# +
# Select the 10 top features, as ranked using ANOVA F-score
from sklearn.feature_selection import SelectKBest, f_classif
tf = SelectKBest(score_func=f_classif, k=10)
Xt = tf.fit_transform(X_train, y_train)
print("Shape =", Xt.shape)
# Plot support
plt.imshow(tf.get_support().reshape((8, 8)), interpolation="nearest", cmap=plt.cm.Blues)
plt.show()
# +
# Feature selection using backward elimination
from sklearn.feature_selection import RFE
from sklearn.ensemble import RandomForestClassifier
tf = RFE(RandomForestClassifier(), n_features_to_select=10, verbose=1)
Xt = tf.fit_transform(X_train, y_train)
print("Shape =", Xt.shape)
# Plot support
plt.imshow(tf.get_support().reshape((8, 8)), interpolation="nearest", cmap=plt.cm.Blues)
plt.show()
# -
# #### Decomposition, factorization or embeddings
# +
# Compute decomposition
from sklearn.decomposition import PCA
# from sklearn.manifold import TSNE
tf = PCA(n_components=2)
Xt_train = tf.fit_transform(X_train)
# Plot
plt.scatter(Xt_train[:, 0], Xt_train[:, 1], c=y_train)
plt.show()
# See also: KernelPCA, NMF, FastICA, Kernel approximations,
# manifold learning, etc
# -
# ### Pipelines
#
# Transformers can be chained in sequence to form a pipeline.
# +
from sklearn.pipeline import make_pipeline
from sklearn.preprocessing import StandardScaler
from sklearn.feature_selection import SelectKBest, f_classif
# Chain transformers to build a new transformer
tf = make_pipeline(StandardScaler(),
SelectKBest(score_func=f_classif, k=10))
tf.fit(X_train, y_train)
# -
Xt_train = tf.transform(X_train)
print("Mean =", np.mean(Xt_train))
print("Shape =", Xt_train.shape)
# Chain transformers + a classifier to build a new classifier
clf = make_pipeline(StandardScaler(),
SelectKBest(score_func=f_classif, k=10),
RandomForestClassifier())
clf.fit(X_train, y_train)
print(clf.predict_proba(X_test)[:5])
# Hyper-parameters can be accessed using step names
print("K =", clf.get_params()["selectkbest__k"])
clf.named_steps
# +
from sklearn.model_selection import GridSearchCV
grid = GridSearchCV(clf,
param_grid={"selectkbest__k": [1, 10, 20, 30, 40, 50],
"randomforestclassifier__max_features": [0.1, 0.25, 0.5]})
grid.fit(X_train, y_train)
print("Best params =", grid.best_params_)
# -
# ### Feature unions
#
# Similarly, transformers can be applied in parallel to transform data in union.
#
# ### Nested composition
#
# Since pipelines and unions are themselves estimators, they can be composed into nested structures.
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.neural_network import MLPClassifier
from sklearn.pipeline import make_union
from sklearn.decomposition import PCA
clf = make_pipeline(
# Build features
make_union(
PCA(n_components=3), # Identity
PCA(),
),
# Select the best features
RFE(RandomForestClassifier(), n_features_to_select=10),
# Train
MLPClassifier()
)
clf.fit(X_train, y_train)
| notebooks/.ipynb_checkpoints/scikit-learn-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="images/logo.jpg" style="display: block; margin-left: auto; margin-right: auto;" alt="לוגו של מיזם לימוד הפייתון. נחש מצויר בצבעי צהוב וכחול, הנע בין האותיות של שם הקורס: לומדים פייתון. הסלוגן המופיע מעל לשם הקורס הוא מיזם חינמי ללימוד תכנות בעברית.">
# # <span style="text-align: right; direction: rtl; float: right;">קבצים</span>
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">הגדרה</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# <dfn>קובץ</dfn> הוא מבנה שמאגד בתוכו נתונים השמורים במחשב שלכם.<br>
# לכל קובץ יש שם וכתובת (<dfn>נתיב</dfn>, או באנגלית <dfn>path</dfn>), והוא מכיל כמות מסוימת של מידע שמשפיעה על גודלו.<br>
# תוכן הקובץ הוא מה שמגדיר את סוגו. סוגי קבצים שונים משמשים למטרות שונות, לדוגמה:
# </p>
# <table style="text-align: right; direction: rtl; clear: both; font-size: 1.3rem">
# <thead>
# <tr>
# <th>סוג הקובץ</th>
# <th>תכלית הקובץ</th>
# <th>דוגמאות לסיומות</th>
# </tr>
# </thead>
# <tbody>
# <tr>
# <td>טקסט</td>
# <td>מכיל מלל בלבד, ללא עיצוב כלשהו (הדגשות, גדלים וכדומה)</td>
# <td>txt</td>
# </tr>
# <tr>
# <td>תמונה</td>
# <td>ייצוג של תמונה דיגיטלית למטרת הצגה חזותית שלה</td>
# <td>png, jpg, gif, bmp</td>
# </tr>
# <tr>
# <td>וידיאו</td>
# <td>ייצוג של סרט או כל וידיאו אחר</td>
# <td>mp4, avi, flv</td>
# </tr>
# <tr>
# <td>פייתון</td>
# <td>מכיל קוד שהתוכנה של פייתון יודעת לקרוא ולהפעיל</td>
# <td>py, pyc, pyd</td>
# </tr>
# <tr>
# <td>הרצה</td>
# <td>מכיל סדרת הוראות המיועדות לקריאה ולהרצה על־ידי המחשב</td>
# <td>exe, dmg</td>
# </tr>
# </tbody>
# </table>
# ## <span style="text-align: right; direction: rtl; float: right; clear: both;">הקדמה</span>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">מהברזל ועד הקובץ</span>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# קבצים הם דרך נהדרת לקבל הצצה חטופה לעומק ולמורכבות של המכונה המדהימה שנקראת מחשב.<br>
# ננסה לצייר לכם תמונה מלאה – נתחיל מהשכבה הנמוכה ביותר, הכונן הקשיח שלכם, ולבסוף נגיע לקבצים.
# </p>
# #### <span style="text-align: right; direction: rtl; float: right; clear: both;">הכונן הקשיח</span>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# <dfn>כונן קשיח</dfn> הוא אחד מהרכיבים הנמצאים בקופסת המחשב שעליו אתם עובדים כרגע.<br>
# זהו הרכיב שמאחסן לטווח ארוך את המידע במחשב שלכם – קבצים, תוכנות, מערכת הפעלה והגדרות, כמו העדפות השפה שלכם.<br>
# היתרון המובהק של כוננים קשיחים הוא שהם יודעים לשמור על המידע שלכם לאורך זמן, גם כאשר הם לא מוזנים בחשמל.
# </p>
# #### <span style="text-align: right; direction: rtl; float: right; clear: both;">ביט</span>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כוננים קשיחים מכילים הרבה יחידות קטנטנות שכל אחת מהן נקראת "ביט".<br>
# <dfn>ביט</dfn> הוא ערך שיכול להיות 0 או 1, כלומר כבוי או דולק.<br>
# בכל כונן קשיח פשוט שנמכר כיום יש מקום ל<em>מאות מיליארדי</em>(!) ביטים כאלו.<br>
# כך נשמר כל המידע שדיברנו עליו בפסקה הקודמת.
# </p>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם נסתכל בעיניים אנושיות על רצפי התאים הקטנטנים, הביטים, שנמצאים בכונן הקשיח, קרוב לוודאי שלא נבין מהם כלום.<br>
# הנה דוגמה לרצף שכזה: 010000100011001100101111010100110011001000110011.<br>
# איך יוצקים משמעות לתוך דבר כזה? מה רצף המספרים הזה אומר בכלל?
# </p>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נדמיין לרגע שאנחנו הממציאים של המחשב, ושהמטרה שלנו היא לייצג טקסט בצורה קלה לשמירה ולקריאה לאחר מכן.<br>
# האם תוכלו לחשוב על דרך לעשות זאת רק באמצעות רצפים של 0 ו־1?
# </p>
#
# <div style="clear: both;"></div>
# #### <span style="text-align: right; direction: rtl; float: right; clear: both;">ייצוג תווים</span>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בעיה דומה של ייצוג טקסט באמצעות אפסים ואחדים בלבד, עלתה אי שם ב־1960.<br>
# אחסוך לכם הרבה מעשיות בדרך, אבל מפה לשם הוחלט על תקן בשם <dfn id="ASCII"><abbr title="American Standard Code for Information Interchange">ASCII</abbr></dfn>, שקובע שכל סידור אפשרי של 8 ביטים שכאלו, שנקרא "<dfn>בייט</dfn>", ייצג תו.<br>
# האות A קיבלה את הייצוג 01000001, האות Z, למשל, קיבלה את הייצוג 01011010, הספרה 7 את הייצוג 00110111 והתו רווח את הייצוג 00100000.<br>
# כך אפשר לקרוא מסר ארוך מאוד, ולהמיר כל 8 ביטים רצופים לתו. אם ננסה לקרוא בשיטה הזו 80 ביטים, נקבל 10 תווים קריאים.
# </p>
# #### <span style="text-align: right; direction: rtl; float: right; clear: both;">ייצוג קבצים אחרים</span>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אבל רגע! מה קורה אם אני רוצה לייצג תמונה של חתול? או סרטון של חתול? או סאונד של... ובכן, חתול?<br>
# כמו שאנשים חכמים חשבו על דרך לייצג ASCII, חכמים אחרים חשבו, עבור כל סוג של קובץ – מתמונה ועד תוכנה שרצה על המחשב, איך מייצגים אותם באמצעות ביטים.<br>
# גם בימים אלו, אנשים מוכשרים רבים יושבים וחושבים על דרכים טובות יותר לייצג מידע בעזרת ביטים, ויוצרים עבורנו סוגי קבצים חדשים.<br>
# </p>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כשאנחנו רוצים לפתוח תמונה של חתול, למשל, אנחנו מפעילים תוכנה ייעודית שיודעת לקרוא ייצוג של תמונות.<br>
# מה שקורה באותו רגע מאחורי הקלעים הוא שהתוכנה קוראת רצפים של 0 ו־1 מהדיסק הקשיח, אותם ביטים שדיברנו עליהם.<br>
# מי שצילם את התמונה השתמש בתוכנה שיודעת להמיר את מה שקרה על מסך המצלמה לביטים שנשמרו על כרטיס הזיכרון שלו.<br>
# מי שתכנת את התוכנה שמציגה לנו כרגע את החתול ידע להורות לה מה לעשות כדי לתרגם את אותם ביטים לתמונה שמוצגת לכם על המסך.<br>
# שניהם פעלו לפי תקן מסוים (יש כמה כאלו, אולי אתם מכירים חלק מהם: JPG, PNG, GIF ועוד), שקובע דרך אחידה לייצג תמונה בעזרת ביטים.<br>
# בסופו של דבר, עבור התוכנה החתול שלכם הוא בסך הכול משבצות קטנטנות בצבעים שונים שמצוירות זו על יד זו.
# </p>
#
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# ייצוג מסוים של קובץ נקרא "<dfn>פורמט</dfn>", או בעברית "<dfn>תַּסְדִיר</dfn>".
# </p>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">קבצים טקסטואליים וקבצים בינאריים</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# חלק מהקבצים, כמו מסמכי טקסט או תוכניות פייתון, יכולים להיות קבצים טקסטואליים.<br>
# ישנם סוגי קבצים טקסטואליים נוספים, כמו <abbr title="Comma Separated Values">CSV</abbr> שמייצג מידע טבלאי באמצעות טקסט, או <abbr title="Hypertext Markup Language">HTML</abbr> שמייצג קוד שבעזרתו המחשב יודע להציג דף אינטרנט.<br>
# <mark>המשותף לקבצים האלו הוא שאם ממירים את הביטים שמהם הם מורכבים לתווים, מתקבלת תוצאה שנוח לבני אדם לקרוא, ולא רק למחשב.</mark>
# </p>
# <figure>
# <img src="images/textual-csv-representation.png" style="display: block; text-align: center; margin-left: auto; margin-right: auto;" width="auto" alt="שני חלונות פתוחים זה ליד זה. בחלון הימני יש תוכנת Microsoft Excel פתוחה, בה נראים כ־15 שורות של מידע אודות יין (נראה שיש עוד מידע רב אם נגלול למטה). שורת הכותרת מכילה את שמות העמודות, ביניהן ארץ מוצא, תיאור, ניקוד, מחיר ועוד, ומתחת לשורת הכותרת ישנם שורות רבות כאשר כל שורה מייצגת יין. בחלון השמאלי ניתן לראות תוכנה פשוטה לעריכת טקסט, ובה מופיעה אותה טבלה בטקסט פשוט. כל שורת יין מופרדת אחת מהשנייה באנטר (שורה חדשה), והתאים בכל שורה מופרדים זה מזה בפסיק.">
# <figcaption style="text-align: center; direction: rtl;">מימין ניתן לראות קובץ CSV פתוח באקסל, ומשמאל את הייצוג הטקסטואלי שלו.</figcaption>
# </figure>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# ישנם גם סוגי קבצים אחרים שלא נועדו לקריאה על־ידי עין אנושית.<br>
# קבצים כאלו נקראים "קבצים בינאריים", ונלמד לטפל בחלק מהם בשלב מתקדם יותר בקורס.<br>
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לפניכם דוגמה לכמה סוגי קבצים בינאריים:
# </p>
# <ul style="text-align: right; direction: rtl; float: right; clear: both;">
# <li>MP3 – תסדיר המיועד עבור קובצי שמע.</li>
# <li>PNG – תסדיר לייצוג תמונות.</li>
# <li>PDF – תסדיר המיועד עבור תצוגה מדויקת של מסמכים.</li>
# <li>XLSX – תסדיר המיועד לאחסון מידע בגיליונות אלקטרוניים (לדוגמה, בעזרת Excel).</li>
# <li>EXE – תסדיר המיועד למערכת ההפעלה חלונות, ומפרט אילו פקודות יש לבצע כדי שתוכנה תרוץ.</li>
# </ul>
# <figure>
# <img src="images/binary-png-representation.png" style="display: block; text-align: center; margin-left: auto; margin-right: auto;" width="auto" alt="שני חלונות פתוחים זה ליד זה. בחלון הימני יש תמונה צבעונית של לוגו הקורס פתוח. בחלון השמאלי ניתן לראות תוכנה פשוטה לעריכת טקסט, ובה מופיעים תווים בלתי דפיסים שלא ניתן לקרוא בעין אנושית.">
# <figcaption style="text-align: center; direction: rtl;">מימין ניתן לראות קובץ PNG פתוח בתוכנה להצגת תמונות, ומשמאל את הייצוג הבינארי שלו כשמנסים להמיר אותו לטקסט.<br>
# קל לראות ש־PNG אינו תסדיר טקסטואלי.</figcaption>
# </figure>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בהמשך החוברת הזו נלמד לטפל בקבצים טקסטואליים.
# </p>
# ## <span style="text-align: right; direction: rtl; float: right;">טיפול בקבצים</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם בעבר השתמשנו בפנקס כדי לשמור מלל, הרי שכיום שימוש בקבצים ממוחשבים עושה הכול לקל ומהיר.<br>
# על קבצים ממוחשבים אפשר לבצע חישובים מסובכים בתוך חלקיק שנייה, ויתרון זה מעניק לנו יכולות שלא היו קיימות בעבר.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# קבצים הם מקור מצוין לקלט ולפלט עבור התוכניות שאנחנו כותבים.<br>
# כמקור קלט, הם יכולים לכלול שורות רבות או מידע מורכב מהרגיל.<br>
# כפלט, הם מאפשרים לנו לשמור מידע בין הרצה להרצה, להעביר את המידע ממקום למקום בקלות ולייצג מידע מורכב בפשטות.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בקובץ <em>passwords.txt</em> שנמצא בתוך תיקיית resources, אספנו לכם את 25 הסיסמאות הנפוצות ביותר בעולם.<br>
# בתור התחלה, ננסה להציץ במה שכתוב בתוך הקובץ בעזרת פייתון.
# </p>
# ### <span style="text-align: right; direction: rtl; float: right;">פתיחת קובץ</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הפונקציה <code>open</code> מאפשרת לנו לפתוח קובץ בעזרת פייתון, כדי להשתמש בו בהמשך התוכנית.<br>
# היא מקבלת 2 פרמטרים: הראשון הוא הנתיב לקובץ, והשני הוא צורת הגישה לקובץ, שעליה מייד נסביר.<br>
# הפונקציה מחזירה ערך שנקרא <dfn>File handler</dfn>, מעין מצביע על הקובץ שעליו נוכל לבצע פעולות.<br>
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# צורת הגישה לקובץ תיבחר לפי המטרה שלשמה אנחנו פותחים אותו:
# </p>
# <ul style="text-align: right; direction: rtl; float: right; clear: both;">
# <li>אם אנחנו מעוניינים לקרוא את הקובץ, צורת הגישה שנבחר תהיה <em>'r'</em> – קריאה, read.</li>
# <li>אם אנחנו מעוניינים לכתוב אל הקובץ ולדרוס את מה שקיים בו, צורת הגישה שנבחר תהיה <em>'w'</em> – כתיבה, write.</li>
# <li>אם אנחנו מעוניינים להוסיף אל הקובץ, צורת הגישה שנבחר תהיה <em>'a'</em> – הוספה, append.</li>
# </ul>
# <div class="align-center" style="display: flex; text-align: right; direction: rtl;">
# <div style="display: flex; width: 10%; float: right; ">
# <img src="images/warning.png" style="height: 50px !important;" alt="אזהרה!">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl;">
# הפרמטר השני, צורת הגישה לקובץ, הוא מחרוזת.<br>
# טעות נפוצה היא לשים שם r, w או a בלי גרשיים סביב.
# </p>
# </div>
# </div>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נתחיל בפתיחת הקובץ, ובהשמה של ה־file handler למשתנה:
# </p>
common_passwords_file = open('resources/passwords.txt', 'r')
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כך נראה הסוג של file handler המצביע לקובץ טקסטואלי:
# </p>
type(common_passwords_file)
# ### <span style="text-align: right; direction: rtl; float: right;">קריאת קובץ</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# עכשיו, כשהקובץ פתוח, נוכל להשתמש בו.<br>
# נבקש מפייתון לקרוא את תוכן הקובץ באמצעות הפעולה <code>read</code>, ונבצע השמה של התוכן שחזר מהפעולה, למשתנה:
# </p>
common_passwords = common_passwords_file.read()
print(common_passwords)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נשים לב שהערך שחזר לנו מפעולת הקריאה הוא מחרוזת לכל דבר:
# </p>
type(common_passwords)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# זיכרו ששורות חדשות מיוצגות על ידי התו <em style="direction: ltr" dir="ltr">\n</em>, וכך גם ב־common_passwords:
# </p>
common_passwords
# <div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
# <div style="display: flex; width: 10%; float: right; clear: both;">
# <img src="images/exercise.svg" style="height: 50px !important;" alt="תרגול">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# פתחו את הקובץ וקראו אותו בעצמכם, קבלו מהמשתמש את הסיסמה שלו, והדפיסו אם היא בין 25 הסיסמאות הנפוצות ביותר.<br>
# <strong>בונוס</strong>: אם היא בין 25 הסיסמאות הנפוצות, החזירו את המיקום שלה ברשימה.
# </p>
# </div>
# </div>
# ### <span style="text-align: right; direction: rtl; float: right;">הסמן</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם ננסה לקרוא שוב את הקובץ, נגלה תופעה מוזרה מעט:
# </p>
common_passwords_again = common_passwords_file.read()
print(common_passwords_again)
# +
# # ???
common_passwords_again == ''
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# קריאה נוספת של הקובץ החזירה לנו הפעם שהוא ריק!<br>
# ודאי תשמחו לגלות שזו התנהגות צפויה – הכל באשמתו של הסמן.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# תופתעו לדעת שאתם כבר מכירים את הרעיון של סמן!<br>
# דמיינו שאתם פותחים מסמך לעריכה, או אפילו פותרים תרגיל במחברת.<br>
# נסו להיזכר בקו המהבהב שמסמן לכם את המיקום של התו הבא שתכתבו.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כאשר אנחנו פותחים קובץ לקריאה בעזרת פייתון, ה<dfn>סמן</dfn> (באנגלית: <dfn>cursor</dfn>) מצביע לתחילת הקובץ.<br>
# ברגע שאתם מבקשים מפייתון לקרוא את הקובץ בעזרת הפעולה <code>read</code>, היא קוראת מהמקום שבו נמצא הסמן ועד סוף הקובץ.<br>
# בזמן הקריאה הסמן יעבור לסוף הקובץ, ולכן כשתנסו לקרוא אותו שוב – תקבלו מחרוזת ריקה.
# </p>
# #### <span style="text-align: right; direction: rtl; float: right;">seek</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כדי להחזיר את הסמן לתחילת הקובץ, נשתמש בפעולה <code>seek</code> ונבקש ממנה להחזיר את הסמן כך שיצביע למקום 0 – לפני התו הראשון:
# </p>
common_passwords_file.seek(0)
print(common_passwords_file.read())
# #### <span style="text-align: right; direction: rtl; float: right;">tell</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# ניתן לראות איפה הסמן נמצא באמצעות הפעולה <code>tell</code>:
# </p>
common_passwords_file.tell()
common_passwords_file.read()
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# מכאן ניתן להסיק, שאם נעשה <code>seek</code> ונספק כארגומנט את המספר שחזר בתא האחרון, נעביר את הסמן כך שיצביע לסוף הקובץ.
# </p>
# ### <span style="text-align: right; direction: rtl; float: right;">קריאה בצורות נוספות</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לעיתים, בייחוד כאשר מדובר בקובץ גדול, נעדיף לקרוא בכל פעם רק חלק מהקובץ.<br>
# נוכל לבחור לקרוא מספר מסוים של תווים באמצעות הפעולה <code>read</code> שאנחנו כבר מכירים,<br>
# אלא שהפעם נעביר לה ארגומנט שיורה לה כמה תווים לקרוא:
# </p>
common_passwords_file.seek(0)
common_passwords_file.read(10)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כדאי לזכור שהסמן יצביע עכשיו על מיקום 10, והפעלה נוספת של פעולת הקריאה תמשיך מהמקום הזה:
# </p>
common_passwords_file.read(5)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# פייתון גם מאפשרת לנו לקרוא עד סוף השורה הנוכחית, שעליה נמצא הסמן, באמצעות הפעולה <code>readline</code>.<br>
# שימו לב שהשורה תסתיים בתווים המייצגים שורה חדשה:
# </p>
common_passwords_file.seek(0)
common_passwords_file.readline()
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לקריאת כל השורות בקובץ, ניתן להשתמש בפעולה <code>readlines</code>, שתחזיר לנו רשימת מחרוזות.<br>
# כל מחרוזת ברשימה מייצגת שורה אחת בקובץ:
# </p>
common_passwords_file.seek(0)
passwords = common_passwords_file.readlines()
print("The passwords variable looks like: " + str(passwords))
print("The type of 'passwords' is: " + str(type(passwords)))
# אם נרצה להיפטר מתו השורה החדשה באחת השורות, נוכל להשתמש בפעולה שלמדנו על מחרוזות:
# strip
print("The most common password in the list is: " + passwords[0].strip())
print("The least common password in the list is: " + passwords[-1].strip())
# <div class="align-center" style="display: flex; text-align: right; direction: rtl; clear: both;">
# <div style="display: flex; width: 10%; float: right; clear: both;">
# <img src="images/exercise.svg" style="height: 50px !important;" alt="תרגול">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כתבו פונקציה שמקבלת נתיב לקובץ, ומחזירה רשימה שבה כל איבר הוא שורה בקובץ.<br>
# בניגוד לפעולה <code>readlines</code>, דאגו שהמחרוזות ברשימה לא יסתיימו בתו שמייצג ירידת שורה.
# </p>
# </div>
# </div>
# ### <span style="text-align: right; direction: rtl; float: right;">סגירת קובץ</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# חשבתם שהשארת דלת פתוחה נחשבת גסות רוח? נראה שעדיין לא שמעתם מה מתכנתים חושבים על קבצים שנשארים פתוחים.<br>
# כיוון שחשוב לנו להיות מנומסים, אנחנו נסגור קבצים לאחר שסיימנו להשתמש בהם.<br>
# קובץ פתוח תופס משאבי מערכת (כמו זיכרון), ולעיתים יגרום לכך שתוכנות אחרות לא יוכלו לגשת אליו.<br>
# השארת קבצים פתוחים היא מנהג מגונה שיגרום להאטה בביצועים ואפילו לקריסות בלתי צפויות, אם יותר מדי file handlers פתוחים.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לא מדובר בפעולה מסובכת מדי. כל מה שתצטרכו לעשות הוא להשתמש בפעולה <code>close</code>:
# </p>
common_passwords_file.close()
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שימו לב שכל ניסיון לעשות שימוש בקובץ אחרי סגירתו, ייכשל:
# </p>
common_passwords_file.read()
# ### <span style="text-align: right; direction: rtl; float: right;">כתיבה לקובץ</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נסתכל שוב על המידע שראינו בתרגיל על הפוקימונים:
# </p>
pokemons = """
#,Name,Type 1,Type 2,Total,HP,Attack,Defense,Sp. Atk,Sp. Def,Speed,Generation,Legendary
1,Bulbasaur,Grass,Poison,318,45,49,49,65,65,45,1,False
2,Ivysaur,Grass,Poison,405,60,62,63,80,80,60,1,False
3,Venusaur,Grass,Poison,525,80,82,83,100,100,80,1,False
4,Charmander,Fire,,309,39,52,43,60,50,65,1,False
5,Charmeleon,Fire,,405,58,64,58,80,65,80,1,False
6,Charizard,Fire,Flying,534,78,84,78,109,85,100,1,False
7,Squirtle,Water,,314,44,48,65,50,64,43,1,False
8,Wartortle,Water,,405,59,63,80,65,80,58,1,False
9,Blastoise,Water,,530,79,83,100,85,105,78,1,False
10,Caterpie,Bug,,195,45,30,35,20,20,45,1,False
11,Metapod,Bug,,205,50,20,55,25,25,30,1,False
12,Butterfree,Bug,Flying,395,60,45,50,90,80,70,1,False
13,Weedle,Bug,Poison,195,40,35,30,20,20,50,1,False
14,Kakuna,Bug,Poison,205,45,25,50,25,25,35,1,False
15,Beedrill,Bug,Poison,395,65,90,40,45,80,75,1,False
16,Pidgey,Normal,Flying,251,40,45,40,35,35,56,1,False
17,Pidgeotto,Normal,Flying,349,63,60,55,50,50,71,1,False
18,Pidgeot,Normal,Flying,479,83,80,75,70,70,101,1,False
19,Rattata,Normal,,253,30,56,35,25,35,72,1,False
"""
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הפעם נכתוב אותו לתוך קובץ בתסדיר CSV.<br>
# כדי לכתוב לקובץ נשתמש בפעולה <code>write</code>, לאחר שנפתח את הקובץ במצב כתיבה (<em>w</em>).
# </p>
# <div class="align-center" style="display: flex; text-align: right; direction: rtl;">
# <div style="display: flex; width: 10%; float: right; ">
# <img src="images/warning.png" style="height: 50px !important;" alt="אזהרה!">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl;">
# <strong>זהירות!</strong> פתיחת קובץ קיים במצב <em>w</em> תמחק את התוכן שלו מיידית.
# </p>
# </div>
# </div>
# <div class="align-center" style="display: flex; text-align: right; direction: rtl;">
# <div style="display: flex; width: 10%; float: right; ">
# <img src="images/tip.png" style="height: 50px !important;" alt="טיפ">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl;">
# אם פתחנו קובץ לכתיבה והנתיב אליו לא קיים במערכת שלנו, פייתון תבדוק אם התיקייה שמעליו קיימת.<br>
# אם כן – פייתון תיצור את הקובץ בשבילנו.
# </p>
# </div>
# </div>
pokemons_file = open('pokemon.csv', 'w')
# נסיר את השורות הריקות והרווחים מהצדדים
clear_pokemon = pokemons.strip()
# ונכתוב לקובץ
pokemons_file.write(clear_pokemon)
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# המספר שפייתון מחזירה הוא כמות התווים שייכתבו לקובץ.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# אם תחפשו את הקובץ במחשב ותפתחו אותו, תגלו שפייתון עדיין לא כתבה אליו את הנתונים.<br>
# פייתון שומרת את הנתונים שביקשתם לכתוב בצד במנגנון זיכרון זמני שנקרא buffer ("מִכְלָא" בעברית, תודה ששאלתם), ותכתוב אותם לקובץ כשתסגרו אותו.<br>
# תוכלו להכריח את פייתון לכתוב לקובץ עוד לפני שסגרתם אותו באמצעות הפעולה <code>flush</code>:
# </p>
pokemons_file.flush()
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# לסיום, לא נשכח לסגור את הקובץ כשאנחנו יודעים שכבר לא נשתמש בו:
# </p>
pokemons_file.close()
# <div class="align-center" style="display: flex; text-align: right; direction: rtl;">
# <div style="display: flex; width: 10%; float: right; ">
# <img src="images/tip.png" style="height: 50px !important;" alt="טיפ">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl;">
# כשאתם בתוך המחברת, ניתן להשתמש בסימן קריאה כדי להריץ פקודות או תוכניות שלא קשורות לפייתון.<br>
# לדוגמה, הפקודה: <code dir="ltr" style="direction: ltr;">!pokemon.csv</code> תריץ את הקובץ שכתבנו.<br>
# שימו לב שלא תוכלו להריץ שום דבר אחר במחברת עד שתסגרו את הקובץ.
# </p>
# </div>
# </div>
# !pokemon.csv
# ### <span style="text-align: right; direction: rtl; float: right;">הוספה לקובץ</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# ניתן לפתוח קובץ במצב '<em>w</em>' אם נרצה לכתוב לקובץ חדש, או לדרוס קובץ קיים.<br>
# אם נרצה להוסיף שורות לקובץ קיים, נפתח את הקובץ במצב ההוספה '<em>a</em>', שמסמן <em>append</em>.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# נוסיף את פוקימון מספר 20, רטיקייט, לקובץ הפוקימונים:
# </p>
new_line = "\n20,Raticate,Normal,,413,55,81,60,50,70,97,1,False"
pokemons_table = open('pokemon.csv', 'a')
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# הוספת תוכן לקובץ במצב הוספה תתבצע באמצעות הפעולה <code>write</code>, בדיוק כמו בכתיבת קובץ חדש:
# </p>
pokemons_table.write(new_line)
pokemons_table.close()
# ## <span style="text-align: right; direction: rtl; float: right;">with</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# מילת המפתח <code>with</code> מאפשרת לנו לאגד כמה שורות באותו הקשר.<br>
# בהקשר של קבצים, היא מאפשרת לנו לאגד תחתיה שורות שמטרתן טיפול בקובץ מסוים.<br>
# לדוגמה, השורות הבאות:
# </p>
# +
passwords_file = open('resources/passwords.txt', 'r')
most_used_password = passwords_file.readline()
other_common_passwords = passwords_file.read()
passwords_file.close()
print(most_used_password.strip())
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# יכולה להיכתב אחרת בעזרת מילת המפתח <code>with</code>:
# </p>
# +
with open('resources/passwords.txt', 'r') as passwords_file:
most_used_password = passwords_file.readline()
other_common_passwords = passwords_file.read()
print(most_used_password.strip())
# -
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שימו לב לצורת השימוש ב־<code>with</code>:
# </p>
# <ol style="text-align: right; direction: rtl; float: right; clear: both;">
# <li>כתבנו את מילת המפתח <code>with</code>.</li>
# <li>מייד לאחר מכן השתמשנו בפונקציה <code>open</code> כדי לפתוח את הקובץ עם הארגומנטים שרצינו.</li>
# <li>השתמשנו במילת המפתח <code>as</code>.</li>
# <li>הכנסנו את שם המשתנה שרצינו שבו ישמר ה־file handler.</li>
# <li>כתבנו נקודתיים כדי לסיים את השורה.</li>
# <li>השתמשנו בהזחה, ואז כתבנו את הפעולות שאנחנו רוצים לעשות בהקשר לקובץ.</li>
# </ol>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בתור מתכנתים, אתם בוחרים איך לכתוב את הקוד שלכם.<br>
# ובכל זאת, קל לראות את היתרונות של שימוש ב־<code>with</code>: <mark>הקוד נעשה קריא ומסודר יותר, והקובץ נסגר לבד כשמסתיימת ההזחה.</mark>
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# כתיבת קוד עם <code>with</code> היא פשוטה יותר ממה שנראה בהתחלה –<br>
# 5 הסעיפים הראשונים התייחסו לשורה הראשונה, ששקולה לשורה הראשונה והרביעית בקוד בלי ה־<code>with</code>.
# </p>
# ## <span style="align: right; direction: rtl; float: right; clear: both;">מונחים</span>
# <dl style="text-align: right; direction: rtl; float: right; clear: both;">
# <dt>קובץ טקסטואלי</dt><dd>קובץ שאפשר להמיר את הבייטים שבו לתווים, ומקבלים קובץ שקריא עבור בני אדם.</dd>
# <dt>קובץ בינארי</dt><dd>קובץ שהתוכן שלו לא נועד לקריאה על־ידי עין אנושית.</dd>
# <dt>סמן</dt><dd>מצביע על המיקום הנוכחי בקובץ, שממנו יתבצעו הפעולות (קריאה, כתיבה וכדומה).</dd>
# <dt>ASCII</dt><dd>שיטה מוסכמת לייצוג תווים.</dd>
# <dt>תסדיר, פורמט</dt><dd>מוסכמה על הדרך שבה ייוצג קובץ מסוים.</dd>
# <dt>File handler</dt><dd>ערך שבאמצעותו ניתן לבצע פעולות המשפיעות על קובץ שפתחנו.</dd>
# </dl>
# ## <span style="text-align: right; direction: rtl; float: right;">תרגולים</span>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">מי דגנים בריאים?</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בקובץ <i>cereal.csv</i> שנמצא בתיקיית resources, ישנו מידע תזונתי על מִדְגַּנִּים שונים (יענו, דגני בוקר).<br>
# ככל שהמדגנים שמופיעים בשורה מסוימת בריאים יותר, כך המספר שמופיע לידם בעמודה rating גבוה יותר.<br>
# מצאו את המדגנים הבריאים ביותר והדפיסו את שמם לצד הציון שקיבלו.<br>
# קרדיט: את הקובץ הבאנו <a href="https://perso.telecom-paristech.fr/eagan/class/igr204/datasets">מכאן</a>.
# </p>
# ### <span style="text-align: right; direction: rtl; float: right; clear: both;">תקווה מארחת</span>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# בקובץ hope.txt שנמצא בתיקיית resources, נמצאת אחת הפואמות המהממות של אמילי דיקנסון, תִּקְוָה הִיא בַּעֲלַת-הַנוֹצָה.<br>
# אך אבוי! הפואמה התבלגנה, וכעת סדר המילים בכל שורה הוא הפוך.
# </p>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# במקום:
# </p>
# <blockquote style="border-left:none !important; position: relative; padding-left: 0.5em; padding: 1.5em; line-height: 1.5em; min-height: 2em; border-right: 3px solid #a93226; background-color: #fbe7e6; font-size: 17px; direction: rtl; text-align: right; clear: both;">
# תִּקְוָה הִיא בַּעֲלַת-הַנוֹצָה<br>
# זוּ בַּנְּשָׁמָה תִשְׁכֹּן –
# </blockquote>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# מופיע:
# </p>
# <blockquote style="border-left:none !important; position: relative; padding-left: 0.5em; padding: 1.5em; line-height: 1.5em; min-height: 2em; border-right: 3px solid #a93226; background-color: #fbe7e6; font-size: 17px; direction: rtl; text-align: right; clear: both;">
# בַּעֲלַת-הַנוֹצָה הִיא תִּקְוָה<br>
# – תִשְׁכֹּן בַּנְּשָׁמָה זוּ
# </blockquote>
# <p style="text-align: right; direction: rtl; float: right; clear: both;">
# שמרו גיבוי של הקובץ, וכתבו קוד שמסדר את הפואמה המבולגנת.<br>
# שמרו את הפואמה המסודרת בקובץ חדש ששמו <i>hope2.txt</i>.
# </p>
# <div class="align-center" style="display: flex; text-align: right; direction: rtl;">
# <div style="display: flex; width: 10%; float: right; ">
# <img src="images/warning.png" style="height: 50px !important;" alt="אזהרה!">
# </div>
# <div style="width: 90%">
# <p style="text-align: right; direction: rtl;">
# <strong>זהירות!</strong> יש פה בעיה שלא למדנו איך לתקן.<br>
# התרגיל בודק גם מה הבנתם מהשיעור על משאבים ברשת בשבוע שעבר :)
# </p>
# </div>
# </div>
| week3/3_Files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Lab 1: Data analysis with numpy
# +
# Name: <NAME>
# CIS D009 Winter
# Assignment 1 Solutions
# -
# Write code to investigate population data of countries of the world.
# <br>
# <br>Note:
# <br>- Do not use pandas for this lab
# <br>- Take advantage of numpy's functions instead of writing loops to access data
# <br>If your code doesn't contain any loop to access data in the population numpy array, you earn 1pt EC
#
# 1. From `countryNames.csv` file, read and store all the country names.
# <br>From `years.txt` file, read and store all the years.
# <br>Print the number of countries and the number of years, with text explanation
# <br>
# <br>Example of print out, including text explanation:
# <br> Countries: 215
# <br> Years: 60
# +
import csv
import random
import numpy as np
data = np.loadtxt("countryNames.csv", delimiter =',', skiprows = 1, usecols=(0,), dtype="str")
print(f"Countries: {len(data)}")
#print(data)
data1 = np.loadtxt("years.txt", delimiter=",", dtype="int")
print(f"Years: {len(data1)}")
#print(data1)
print()
# SECOND METHOD to print the count of Countires.
# This is just to test that we are getting the correct name of the Countries.
# Ex: Line 21 in countryNames.csv file is as below line. In this country name is having comma in the name itself.
# Bahamas, The",BHS,Latin America & Caribbean
with open('countryNames.csv') as csv_file:
csv_reader = csv.reader(csv_file, delimiter=',')
next(csv_reader)
d = [name[0] for name in csv_reader]
print(f"Countries: {len(d)}")
### this is a better choice, given that you only want to store the country names
# -
# 2. From `population.csv` file, create a 2D numpy array of the population.
# <br>Each line of the file corresponds with one country in the `countryNames.csv` file.
# <br>Each column of the file corresponds with one year in the `years.txt` file.
# <br>Print the tuple of the number of rows and number of columns of the numpy array to show that the numbers match the number of countries and years above.
# <br>
# <br>Since the data values are large (in billions) make sure you use the numpy data type that can store large values
with open("population.csv", "r") as file:
numpy_array = np.loadtxt(file, delimiter=",", dtype="uint64")
print(numpy_array.shape)
# 3. Print the total world population in 1960 (first year) and in 2019 (last year), with text explanation.
# <br>Then print the population growth (the difference) between 1960 and 2019, with text explanation.
# <br>To print a large number with commas, use: `f'{largeNum:,}'` where largeNum is the large value
# <br>
# <br>Example of output with large number and text explanation: 1960 population: 3,014,118,076
with open("population.csv", "r") as file:
numpy_array = np.loadtxt(file, delimiter=",", dtype="uint64")
### should not have to run this again, you can use numpy_array from the cell above
# print(numpy_array.shape)
column_sums = numpy_array.sum(axis=0)
print(f"1960 population: {column_sums[0]:,}")
print(f"2019 population: {column_sums[-1]:,}")
print(f"Population growth from 1960 to 2019 is: {column_sums[-1] - column_sums[0]:,}")
# 4. Determine in which quartile the 2019 United States population belongs. Print whether it's in the 25, 50, 75, or higher percentile.
# +
# Calculating 2019 Popuation
numpy_array = np.loadtxt("population.csv", delimiter=",", dtype="uint64")
### don't neeed to do this again
pop_in_2019 = numpy_array[:, -1]
# print(numpy_array[:, -1])
# Calculating Country Names
data = np.loadtxt("countryNames.csv", delimiter =',', skiprows = 1, usecols=(0,), dtype="str")
### don't need to do this again, either
# Calculating country wise population for year 2019
country_wise_data = np.column_stack((data, pop_in_2019))
# print(country_wise_data)
# Cal 25, 50 and 75 percentile values
per_25 = np.percentile(pop_in_2019, 25)
per_50 = np.percentile(pop_in_2019, 50)
per_75 = np.percentile(pop_in_2019, 75)
print(per_25, per_50, per_75)
# Getting US population
us_pop = country_wise_data[np.where(country_wise_data[:,0] == "United States")].flat[1]
# Getting Int value
us_pop_int = int(us_pop)
#Checking and printing US percentile values
if us_pop_int <= per_25:
print("US popuation is less than 25 percentile")
elif us_pop_int <=50:
print("US popuation is less than 50 percentile")
elif us_pop_int <= 75:
print("US popuation is less than 75 percentile")
else:
print("US popuation is higher than 75 percentile")
# arr[arr[:, 1].argsort()]
# -
# 5. Print the median population of 2019. Then prove that it's the correct value by printing the number of countries with population below the median, and the number of countries with population above the median.
with open("population.csv", "r") as file:
numpy_array = np.loadtxt(file, delimiter=",", dtype="uint64")
#print(numpy_array)
# print(type(numpy_array[:, -1]))
print(f"{int(np.median(numpy_array[:, -1]))}")
print(f"Number of countires with less than median population: {len(numpy_array[:, -1][numpy_array[:, -1]<np.median(numpy_array[:, -1])])}")
print(f"Number of countires with more than median population: {len(numpy_array[:, -1][numpy_array[:, -1]>np.median(numpy_array[:, -1])])}")
# 6. Find the top 10 countries with the largest population in 2019. Print the country names and population, sorted by highest to lowest population, and formatted as 2 columns.
# <br>You can use a loop if it's only to print and format the output in 2 columns. The loop won't count toward the 1pt EC of no loops.
# +
# Calculating 2019 Popuation
numpy_array = np.loadtxt("population.csv", delimiter=",", dtype="uint64")
pop_in_2019 = numpy_array[:, -1]
# print(numpy_array[:, -1])
# Calculating Country Names
data = np.loadtxt("countryNames.csv", delimiter =',', skiprows = 1, usecols=(0,), dtype="str")
# Calculating country wise population for year 2019
country_wise_data = np.column_stack((data, pop_in_2019))
len(country_wise_data[country_wise_data[:, 1].argsort()])
# Method 1 is using argsort
top10_country_data = country_wise_data[country_wise_data[:, 1].argsort()][::-1][:10,:]
# print(top10_country_data)
for row in top10_country_data:
print(f"{row[0]:<30} {row[1]:20}")
print()
# Method 2 is using sorted method
top10_country_data_sorted = sorted(country_wise_data, key=lambda row: row[1], reverse=True)[:10]
for row in top10_country_data_sorted:
print(f"{row[0]:<30} {row[1]:20}")
### method 2 will be slower than method 1 for large arrays. Use numpy when possible.
### Either way, these are not the top 10 population numbers. The population become strings
### when you combine the 2 arrays, because numpy arrays can only store one type of data.
### Therefore, the sort is a string sort, not a numeric sort -1.5pts
# -
# 7. A few countries do not have the population data for all years between 1960 and 2019. The missing data is indicated by a 0 in the population count.
# <br>Print the count of the rows (countries) that have missing data.
# <br>Print the index of the rows that have missing data.
numpy_array = np.loadtxt("population.csv", delimiter=",", dtype="uint64")
indices = np.argwhere(np.any(numpy_array == 0, axis=1))
print(f"Count of the rows (countries) that have missing data is: {len(indices):>2}")
print()
print(f"Index of the rows that have missing data: \n{indices}")
# 8. Print the country with the smallest population growth between 1960 and 2019,
# <br>and print the country with the largest population growth between 1960 and 2019.
# <br>The result should not show countries where the population count is 0 in 2019, resulting in a false smallest growth (a negative value), or where the population count is 0 in 1960, resulting in a false largest growth.
# +
#Getting the 2019 and 1960 population data
numpy_array = np.loadtxt("population.csv", delimiter=",", dtype="uint64")
pop_in_2019 = numpy_array[:, -1]
pop_in_1960 = numpy_array[:, 0]
# Getting Country Names
data = np.loadtxt("countryNames.csv", delimiter =',', skiprows = 1, usecols=(0,), dtype="str")
# Creating a 2D string array with country name, 2019 population and 1960 populatio
country_wise_data = np.column_stack((data, pop_in_2019, pop_in_1960 ))
# print(len(country_wise_data))
# Creating the 2D array with 2019 and 1960 population
country_wise_data_new = np.column_stack((pop_in_2019, pop_in_1960 ))
# Finding the row indexes if any column is 0
d1 = country_wise_data_new[np.any(country_wise_data_new == 0, axis=1)]
rows, cols = np.where(country_wise_data_new == 0)
#Deleting the countries who has either 0 population in 1960 or 2019 to satisfy below condition
## The result should not show countries where the population count is 0 in 2019, resulting in a false smallest
## growth (a negative value), or where the population count is 0 in 1960, resulting in a false largest growth.
new_country_wise_data = np.delete(country_wise_data, rows, axis=0)
# Sorting the data based on the population growth from 1960 t0 2020
sorted_country_po_growth_data = sorted(new_country_wise_data, key=lambda row: int(row[1])-int(row[2]), reverse=True)
### use numpy, not Python sorted -1/2pt
### when there are lots of data, a list is slower than a numpy array
# print the country with the largest population growth between 1960 and 2019.
print(f"{sorted_country_po_growth_data[0][0]} has larget population growth of {int(sorted_country_po_growth_data[0][1]) - int(sorted_country_po_growth_data[0][2])}")
# Print the country with the smallest population growth between 1960 and 2019
# print(sorted_country_po_growth_data[-1])
print(f"{sorted_country_po_growth_data[-1][0]} has smallest population growth of {int(sorted_country_po_growth_data[-1][1]) - int(sorted_country_po_growth_data[-1][2])}")
# +
# # +1 EC
| CIS009/2_DataAnalysisWithPandas_DataVisualizationWithMatplotlib/lab1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hashPhoeNiX/ColabNotebooks/blob/master/Marine_Invertebrates_Indentification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="BXPUJWFiDcUO" colab_type="text"
# ## Splitting data into train and validation set
# + id="JA5derPdDbeA" colab_type="code" colab={}
# # !pip install split-folders tqdm
# + id="mxmGFfQgJ9nF" colab_type="code" colab={}
# import split_folders
# split_folders.ratio('train_small', output='data', seed=1337, ratio=(.8, .1, .1))
# + [markdown] id="4xq9BX4GDi7P" colab_type="text"
# ## Import Libraries
# + id="--KkOLoB44X8" colab_type="code" colab={}
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
import matplotlib.pyplot as plt
from PIL import Image
import numpy as np
import torch
from torch import nn
from torch import optim
import torch.nn.functional as F
from torchvision import datasets, transforms, models
from torch.utils.data import random_split
# + [markdown] id="296IZ2Rp7qhO" colab_type="text"
# ## Data Augmentation
# + id="mOGHMGgU5CQA" colab_type="code" colab={}
train_dir = 'data/train/'
valid_dir = 'data/val'
test_dir = 'data/test'
# + [markdown] id="PfnuDLHsOY9P" colab_type="text"
# ## An idea
#
# + id="E8fq22NuOXbz" colab_type="code" colab={}
# doing data augmentation
train_transform = transforms.Compose([transforms.Resize((224,224)),
transforms.RandomRotation(45),
transforms.RandomHorizontalFlip(),
transforms.RandomVerticalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
val_transform = transforms.Compose([transforms.Resize((224,224)),
transforms.RandomRotation(45),
transforms.RandomHorizontalFlip(),
transforms.RandomVerticalFlip(),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
test_transform = transforms.Compose([transforms.Resize((224,224)),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406],
[0.229, 0.224, 0.225])])
# + id="PAfAAaJ-OX82" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="1a38288d-5686-403a-c188-6dbe39a73e86"
# splitting the data into train/validation/test sets
data = datasets.ImageFolder('train_small')
train_size = int(len(data)*0.9)
val_size = int((len(data)-train_size)*0.7)
test_size = int(len(data)-train_size-val_size)
train_data,val_data,test_data = random_split(data,[train_size,val_size,test_size])
print(f'train size: {len(train_data)}\nval size: {len(val_data)}\ntest size: {len(test_data)}')
# + id="AOHkoihmOWV-" colab_type="code" colab={}
train_data.dataset.transform = train_transform
val_data.dataset.transform = val_transform
test_data.dataset.transform = test_transform
batch_size = 16
trainloader = torch.utils.data.DataLoader(train_data,batch_size=batch_size,shuffle=True)
validloader = torch.utils.data.DataLoader(val_data,batch_size=batch_size,shuffle=True)
testloader = torch.utils.data.DataLoader(test_data,batch_size=batch_size,shuffle=False)
# + id="fxY4T6MPOWCp" colab_type="code" colab={}
# + [markdown] id="RnnxOcZ_OdBw" colab_type="text"
# ## Original idea
#
# + id="KvMsRQgS5JWH" colab_type="code" colab={}
# train_transform = transforms.Compose([transforms.RandomRotation(30),
# transforms.RandomResizedCrop(299),
# transforms.RandomHorizontalFlip(),
# transforms.ToTensor()
# ])
# test_transform = transforms.Compose([transforms.RandomResizedCrop(299),
# transforms.ToTensor()
# ])
# trainset = datasets.ImageFolder(train_dir, transform=train_transform)
# validset = datasets.ImageFolder(valid_dir, transform=test_transform)
# testset = datasets.ImageFolder(test_dir, transform=test_transform)
# + id="fCa-5jaw7JPa" colab_type="code" colab={}
# trainloader = torch.utils.data.DataLoader(trainset, shuffle=True, batch_size=32)
# validloader = torch.utils.data.DataLoader(validset, shuffle=True, batch_size=32)
# testloader = torch.utils.data.DataLoader(testset, shuffle=False, batch_size=32)
# + id="qHI1Lgt-R3Kf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="736a0449-e797-472a-f283-6948ed3517ee"
# len(trainset) + len(testset) + len(validset)
# + [markdown] id="ugBXhYLdAcLv" colab_type="text"
# ## Transfer Learning
# + id="n8XJ_xZJNC3Q" colab_type="code" colab={}
def imshow_original(image, ax=None, title=None, normalize=True):
"""Imshow for Tensor."""
if ax is None:
fig, ax = plt.subplots()
image = image.transpose((1, 2, 0))
if normalize:
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
image = std * image + mean
image = np.clip(image, 0, 1)
ax.imshow(image)
ax.spines['top'].set_visible(False)
ax.spines['right'].set_visible(False)
ax.spines['left'].set_visible(False)
ax.spines['bottom'].set_visible(False)
ax.tick_params(axis='both', length=0)
ax.set_xticklabels('')
ax.set_yticklabels('')
return ax
# + id="XGyq4XaR_zbm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="69dbd0c8-29b2-427e-b655-91cdad1fd498"
dataiter = iter(trainloader)
images, labels = dataiter.next()
print(type(images))
print(images.shape)
print(labels.shape)
# imshow_original(images)
# + id="DZkXucDL8XMV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8e3b5887-4aba-443a-86ec-3525ac1ee009"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
device
# + [markdown] id="qS-8TATZASWH" colab_type="text"
# ## Model Training using VGG Pretrained Network
#
# + id="IdSfKC68fN_f" colab_type="code" colab={}
# defining model evaluation function
def evaluation(dataloader, model):
total, correct = 0, 0
for data in dataloader:
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
outputs = model(inputs)
_, pred = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (pred == labels).sum().item()
return (100 * correct / total)
# + id="vL7FrknjfJUS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="2bb20cab-f5f3-4d79-9594-0dfb7e9b45e0"
# Loading Pretrained Network from torchvision.models
vgg = models.vgg16_bn(pretrained=False)
# Freezing the model parameters
for param in vgg.parameters():
param.requires_grad = False
# Building a new vgg model classifier
final_in_features = vgg.classifier[6].in_features
vgg.classifier[6] = nn.Linear(final_in_features, 137)
for param in vgg.parameters():
if param.requires_grad:
print(param.shape)
# define loss and model optimizer
vgg = vgg.to(device)
loss_fn = nn.CrossEntropyLoss()
opt = optim.Adam(vgg.parameters(),lr=0.003)
# + id="1y65Fm5kfF36" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 694} outputId="3533c446-8199-499a-bcdf-bc858cb413d3"
# # %%time
# import copy
# loss_epoch_arr = []
# max_epochs = 15
# batch_size = 32
# min_loss = 10000000
# n_iters = np.ceil(5300/batch_size)
# for epoch in range(max_epochs):
# for i, data in enumerate(trainloader, 0):
# inputs, labels = data
# inputs, labels = inputs.to(device), labels.to(device)
# opt.zero_grad()
# outputs = vgg(inputs)
# loss = loss_fn(outputs, labels)
# loss.backward()
# opt.step()
# if min_loss > loss.item():
# min_loss = loss.item()
# best_model = copy.deepcopy(vgg.state_dict())
# print('Min loss %0.2f' % min_loss)
# if i % 100 == 0:
# print('Iteration: %d/%d, Loss: %0.2f' % (i, n_iters, loss.item()))
# del inputs, labels, outputs
# torch.cuda.empty_cache()
# loss_epoch_arr.append(loss.item())
# vgg.load_state_dict(best_model)
# print(evaluation(trainloader, vgg), evaluation(validloader, vgg))
# + id="AFHCrxk7TBgI" colab_type="code" colab={}
import time
def fit(model, criterion, optimizer, num_epochs=10):
start = time.time()
best_model = model.state_dict()
best_acc = 0.0
train_loss_over_time = []
val_loss_over_time = []
train_acc_over_time = []
val_acc_over_time = []
# each epoch has a training and validation phase
for epoch in range(num_epochs):
print(f'{epoch+1}/{num_epochs} epoch')
for phase in ['train','val']:
if phase == 'train':
data_loader = trainloader
model.train() # set the model to train mode
else:
data_loader = validloader
model.eval() # set the model to evaluate mode
running_loss = 0.0
running_corrects = 0.0
# iterate over the data
for inputs,labels in data_loader:
inputs = inputs.to(device)
labels = labels.to(device)
# zero the parameter gradients
optimizer.zero_grad()
# forward
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
_,pred = torch.max(outputs,dim=1)
loss = criterion(outputs,labels)
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# calculating the loss and accuracy
running_loss += loss.item() * inputs.size(0)
running_corrects += torch.sum(pred == labels.data)
if phase == 'train':
epoch_loss = running_loss/len(train_data)
train_loss_over_time.append(epoch_loss)
epoch_acc = running_corrects.double()/len(train_data)
train_acc_over_time.append(epoch_acc)
else:
epoch_loss = running_loss/len(val_data)
val_loss_over_time.append(epoch_loss)
epoch_acc = running_corrects.double()/len(val_data)
val_acc_over_time.append(epoch_acc)
print(f'{phase} loss: {epoch_loss:.3f}, acc: {epoch_acc:.3f}')
if phase == 'val' and epoch_acc > best_acc:
best_acc = epoch_acc
best_model = model.state_dict()
print('-'*60)
total_time = (time.time() - start)/60
print(f'training complete in: {total_time:.3f} min\nbest accuracy: {best_acc:.3f}')
# load best model weights
model.load_state_dict(best_model)
loss = {'train':train_loss_over_time, 'val':val_loss_over_time}
acc = {'train':train_acc_over_time, 'val':val_acc_over_time}
return model,loss,acc
# + id="XGucnlsTTQtO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="26994051-40f7-4121-abd9-a1f52f8f10e6"
# %time
epochs = 50
history, loss, acc = fit(vgg,loss_fn,opt,num_epochs=epochs)
# + id="TQO9IJLwahKx" colab_type="code" colab={}
# loss, acc
# + id="ZxWnsZcodQpi" colab_type="code" colab={}
def evaluate(model,criterion):
model.eval() # setting the model to evaluate mode
test_loss = 0.0
test_acc = 0.0
preds = []
labels_list = []
for inputs,labels in testloader:
inputs = inputs.to(device)
labels = labels.to(device)
# predicting
with torch.no_grad():
outputs = model(inputs)
loss = criterion(outputs,labels)
_,pred = torch.max(outputs,dim=1)
preds.append(pred)
labels_list.append(labels)
# calculating the loss and accuracy
test_loss += loss.item()*inputs.size(0)
correct = pred.eq(labels.data.view_as(pred))
accuracy = torch.mean(correct.type(torch.FloatTensor))
test_acc += accuracy.item() * inputs.size(0)
# avreging the loss and accuracy
test_loss = test_loss/len(testloader.dataset)
test_acc = test_acc / len(testloader.dataset)
print("test loss: {:.4f} test acc: {:.4f}".format(test_loss,test_acc))
return preds,labels_list
# + id="oz9dbhVjdddU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ec79a950-d9c2-4620-ddb2-2cb522fd64bd"
# testing the model
predictions,labels = evaluate(vgg,loss_fn)
# + id="wkeZ_Yviklla" colab_type="code" colab={}
# the model's name
model_name = 'model_vgg16.pt'
# saving the best trained model
torch.save(vgg.state_dict(), model_name)
# + id="5HgJbrxWgIUN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 54} outputId="58333b73-7644-4946-bd23-a7198bba711c"
print("Acc on training set is {} and validation set is {}".format(evaluation(trainloader, vgg),evaluation(validloader, vgg)))
# + id="0ILDp8akHynP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 294} outputId="044b8bf8-1a30-4051-cdff-656eeed13aa6"
plt.plot(loss_epoch_arr)
plt.xlabel("epochs")
plt.ylabel("loss")
plt.title("loss vs epochs")
plt.show()
# + id="-gNPQiQJhIg2" colab_type="code" colab={}
| Marine_Invertebrates_Indentification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={} tags=[]
# <img width="10%" alt="Naas" src="https://landen.imgix.net/jtci2pxwjczr/assets/5ice39g4.png?w=160"/>
# + [markdown] papermill={} tags=[]
# # WSR - Get daily Covid19 active cases worldmap JHU
# <a href="https://app.naas.ai/user-redirect/naas/downloader?url=https://raw.githubusercontent.com/jupyter-naas/awesome-notebooks/master/WSR/WSR_Get_daily_Covid19_active_cases_worldmap_JHU.ipynb" target="_parent"><img src="https://naasai-public.s3.eu-west-3.amazonaws.com/open_in_naas.svg"/></a>
# + [markdown] papermill={} tags=[]
# **Tags:** #wsr #covid #active-cases #analytics #plotly #automation #naas
# + [markdown] papermill={} tags=[]
# **Author:** [<NAME>](https://www.linkedin.com/in/ACoAABCNSioBW3YZHc2lBHVG0E_TXYWitQkmwog/)
# + [markdown] papermill={} tags=[]
# ## Input
# + [markdown] papermill={} tags=[]
# ### Import libraries
# + papermill={} tags=[]
import pandas as pd
from datetime import datetime
try:
from dataprep.clean import clean_country
except:
# !pip install dataprep --user
from dataprep.clean import clean_country
import plotly.graph_objects as go
import naas
# + [markdown] papermill={} tags=[]
# ### Setup chart title
# + papermill={} tags=[]
title = "COVID 19 - Active cases (in milions)"
# + [markdown] papermill={} tags=[]
# ### Variables
# + papermill={} tags=[]
# Input URLs of the raw csv dataset
urls = [
'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_confirmed_global.csv',
'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_deaths_global.csv',
'https://raw.githubusercontent.com/CSSEGISandData/COVID-19/master/csse_covid_19_data/csse_covid_19_time_series/time_series_covid19_recovered_global.csv'
]
# Output paths
output_image = f"{title}.png"
output_html = f"{title}.html"
# + [markdown] papermill={} tags=[]
# ### Schedule your automation
# + papermill={} tags=[]
# Schedule your job everyday at 8:00 AM (NB: you can choose the time of your scheduling bot)
naas.scheduler.add(cron="0 8 * * *")
#-> Uncomment the line below (by removing the hashtag) to remove your scheduler
# naas.scheduler.delete()
# + [markdown] papermill={} tags=[]
# ## Model
# + [markdown] papermill={} tags=[]
# ### Get data from JHU
# + papermill={} tags=[]
def get_data_url(urls):
df = pd.DataFrame()
for url in urls:
tmp_df = pd.read_csv(url)
tmp_df["Indicator"] = url.split("/time_series_covid19_")[-1].split("_global.csv")[0].capitalize()
df = pd.concat([df, tmp_df])
return df
df_init = get_data_url(urls)
df_init
# + [markdown] papermill={} tags=[]
# ### Get all data from JHU
# + papermill={} tags=[]
def get_all_data(df_init):
df = df_init.copy()
# Cleaning
df = df.drop("Province/State", axis=1)
# Melt data
df = pd.melt(df,
id_vars=["Country/Region", "Lat", "Long", "Indicator"],
var_name="Date",
value_name="Value").fillna(0)
df["Date"] = pd.to_datetime(df["Date"])
# Calc active cases
df_active = df.copy()
df_active.loc[df_active["Indicator"].isin(["Deaths", "Recovered"]), "Value"] = df_active["Value"] * (-1)
df_active["Indicator"] = "Active cases"
# Concat data
df = pd.concat([df, df_active])
# Group by country/region
to_group = ["Country/Region", "Lat", "Long", "Indicator", "Date"]
df = df.groupby(to_group, as_index=False).agg({"Value": "sum"})
# Cleaning
df = df.rename(columns={"Country/Region": "COUNTRY"})
df.columns = df.columns.str.upper()
return df.reset_index(drop=True)
df_clean = get_all_data(df_init)
df_clean
# + [markdown] papermill={} tags=[]
# ### Prep data for worldmap
# + papermill={} tags=[]
def prep_data(df_init):
df = df_init.copy()
# Filter
date_max = df["DATE"].max()
df = df[
(df["INDICATOR"] == "Active cases") &
(df["DATE"] == date_max)
].reset_index(drop=True)
# Clean country
df = clean_country(df, 'COUNTRY', output_format='alpha-3').dropna()
df = df.rename(columns={'COUNTRY_clean': 'COUNTRY_ISO'})
return df.reset_index(drop=True)
df_worldmap = prep_data(df_clean)
df_worldmap
# + [markdown] papermill={} tags=[]
# ### Create worldmap
# + papermill={} tags=[]
def create_worldmap(df):
fig = go.Figure()
fig = go.Figure(data=go.Choropleth(
locations=df['COUNTRY_ISO'],
z=df['VALUE'],
text=df["COUNTRY"] + ": " + df['VALUE'].map("{:,.0f}".format).str.replace(",", " ") + " active cases",
hoverinfo="text",
colorscale='Blues',
autocolorscale=False,
reversescale=False,
marker_line_color='darkgray',
marker_line_width=0.5,
colorbar_tickprefix='',
colorbar_title='Active cases',
))
fig.update_layout(
title=title,
plot_bgcolor="#ffffff",
legend_x=1,
geo=dict(
showframe=False,
showcoastlines=False,
#projection_type='equirectangular'
),
dragmode= False,
width=1200,
height=800,
)
config = {'displayModeBar': False}
fig.show(config=config)
return fig
fig = create_worldmap(df_worldmap)
# + [markdown] papermill={} tags=[]
# ## Output
# + [markdown] papermill={} tags=[]
# ### Export in PNG and HTML
# + papermill={} tags=[]
fig.write_image(output_image, width=1200)
fig.write_html(output_html)
# + [markdown] papermill={} tags=[]
# ### Generate shareable assets
# + papermill={} tags=[]
link_image = naas.asset.add(output_image)
link_html = naas.asset.add(output_html, {"inline":True})
#-> Uncomment the line below to remove your assets
# naas.asset.delete(output_image)
# naas.asset.delete(output_html)
| WSR/WSR_Get_daily_Covid19_active_cases_worldmap_JHU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <NAME>
#
# Fantasy Football Data Visualization
# +
import pandas as pd
import numpy as np
import fantasy_football.src.data.ff as ffdb
import matplotlib.pyplot as plt
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
plt.style.use('ggplot')
# -
# Loading the data. The load function is a wrapper function for several data cleaning functions. Some work still needs to be done to get the future fantasy points and cformatting the name.
# +
fantasy2013 = ffdb.load_data("2013_Fantasy")
fantasy2014 = ffdb.load_data("2014_Fantasy")
fantasy2015 = ffdb.load_data("2015_Fantasy")
fantasy2013df = pd.DataFrame(fantasy2013)
fantasy2014df = pd.DataFrame(fantasy2014)
fantasy2015df = pd.DataFrame(fantasy2015)
# -
# Showing the first ten players in the 2013 NFL Fantasy Football dataset.
print(fantasy2013df.head(10))
# Getting summary statistics for each of the variables in the dataset. PPR is NaN for all players so it should be remved. OvRank has a limited number of values, so it may not be very useful.
print(fantasy2015df.describe())
#PPR is empty for all three years and will be removed
#OvRank has limited values
# Generating a histogram for the yardage by position. A significant portion of the data is within the first few bins, which makes sense because QBs do not frequently catch of rush the ball. It may make sense to create subsets of the data by position for model creation because the positions do different things.
ffdb.hist_yards_by_pos(fantasy2013df, pos = 'QB')
ffdb.hist_yards_by_pos(fantasy2014df, pos = 'RB')
ffdb.hist_yards_by_pos(fantasy2015df, pos = 'WR')
# Following up on the histogram, the next plots are barcharts for yardage by position. TE and WR are somewhat similiar by only having RecYds. It seems that three different models and combining results may be the best route.
ffdb.plot_yards_by_position(fantasy2013df)
ffdb.plot_yards_by_position(fantasy2014df)
ffdb.plot_yards_by_position(fantasy2015df)
# The next plot is a barchart of total fantasy points by team. Some teams have better players than others so if a team has a better QB, all of the other positions should see some benefit.
ffdb.total_fantasy_points_by_team(fantasy2013df)
# The next plot is a scatter plot of fantasy points vs age. The data seems to be fairly random, but it may be worthwile to look further based on position.
plt.scatter(fantasy2015df['Age'], fantasy2015df['FantPt'])
plt.xlabel('Age')
plt.ylabel('Fantasy Points')
plt.title('Fantasy Points vs. Age')
| notebooks/visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function, unicode_literals, absolute_import, division
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
from glob import glob
from tifffile import imread
from csbdeep.utils import Path, normalize
from skimage.segmentation import find_boundaries
from stardist import dist_to_coord, non_maximum_suppression, polygons_to_label
from stardist import random_label_cmap, draw_polygons, sample_points
from stardist import Config, StarDist
np.random.seed(6)
lbl_cmap = random_label_cmap()
# -
# # Data
#
# We assume that data has already been downloaded in via notebook [1_data.ipynb](1_data.ipynb).
# We now load images from the sub-folder `test` that have not been used during training.
# +
X = sorted(glob('data/dsb2018/test/images/*.tif'))
X = list(map(imread,X))
n_channel = 1 if X[0].ndim == 2 else X[0].shape[-1]
axis_norm = (0,1) # normalize channels independently
# axis_norm = (0,1,2) # normalize channels jointly
if n_channel > 1:
print("Normalizing image channels %s." % ('jointly' if axis_norm is None or 2 in axis_norm else 'independently'))
# -
# show all test images
if False:
fig, ax = plt.subplots(7,8, figsize=(16,16))
for i,(a,x) in enumerate(zip(ax.flat, X)):
a.imshow(x,cmap='gray')
a.set_title(i)
[a.axis('off') for a in ax.flat]
plt.tight_layout()
None;
# # Load StarDist models
#
# We assume that two StarDist models have already been trained via notebook [2_training.ipynb](2_training.ipynb).
# Without shape completion
model_no_sc = StarDist(None, name='stardist_no_shape_completion', basedir='models')
# With shape completion
model_sc = StarDist(None, name='stardist_shape_completion', basedir='models')
# ## Prediction
#
# 1. Load image and normalize
# 2. Predict object `prob`abilities and star-convex polygon `dist`ances
# 3. Convert `dist`ances to polygon vertex `coord`inates
# 4. Perform non-maximum suppression for polygons above object probability threshold
# 5. Convert final polygons to label instance image
img = normalize(X[16],1,99.8,axis=axis_norm)
prob, dist = model_sc.predict(img)
coord = dist_to_coord(dist)
points = non_maximum_suppression(coord,prob,prob_thresh=0.4)
labels = polygons_to_label(coord,prob,points)
img_show = img if img.ndim==2 else img[...,0]
fig, ax = plt.subplots(2,2, figsize=(12,12))
for a,d,cm,s in zip(ax.flat, [img_show,prob,dist[...,0],labels], ['gray','magma','viridis',lbl_cmap],
['Input image','Predicted object probability','Predicted distance (0°)','Predicted label instances']):
a.imshow(d,cmap=cm)
a.set_title(s)
a.axis('off')
plt.tight_layout()
None;
# +
from foolbox.models import KerasModel
import keras.backend as K
predictions = model_sc.keras_model.output
print(K.int_shape(predictions[-1]))
model = KerasModel(model_sc.keras_model,bounds=(0, 255))
# -
plt.figure(figsize=(13,12))
points_rnd = sample_points(200,prob>0.2)
plt.subplot(121); plt.imshow(img_show,cmap='gray'); draw_polygons(coord,prob,points_rnd,cmap=lbl_cmap)
plt.axis('off'); plt.title('Polygons randomly sampled')
plt.subplot(122); plt.imshow(img_show,cmap='gray'); draw_polygons(coord,prob,points,cmap=lbl_cmap)
plt.axis('off'); plt.title('Polygons after non-maximum suppression')
plt.tight_layout()
None;
# # Comparing results with and without shape completion
def example(model,i):
img = normalize(X[i],1,99.8,axis=axis_norm)
prob, dist = model.predict(img)
coord = dist_to_coord(dist)
points = non_maximum_suppression(coord,prob,prob_thresh=0.4)
labels = polygons_to_label(coord,prob,points)
img_show = img if img.ndim==2 else img[...,0]
plt.figure(figsize=(13,10))
plt.subplot(121); plt.imshow(img_show,cmap='gray'); plt.axis('off')
draw_polygons(coord,prob,points,show_dist=True)
if model in (model_no_sc,model_sc):
plt.title(('With' if model==model_sc else 'Without') + ' shape completion')
plt.subplot(122); plt.imshow(img_show,cmap='gray'); plt.axis('off')
plt.imshow(labels,cmap=lbl_cmap,alpha=0.5)
plt.tight_layout()
plt.show()
example(model_no_sc,42)
example(model_sc,42)
example(model_no_sc,1)
example(model_sc,1)
example(model_no_sc,15)
example(model_sc,15)
# # Model from paper
model_paper = StarDist(None, name='dsb2018', basedir='../models')
model_paper.load_weights('weights_last.h5')
example(model_paper,29)
| resources/stardist/3_prediction-attack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NumPy Practice
#
# This notebook offers a set of excercises for different tasks with NumPy.
#
# It should be noted there may be more than one different way to answer a question or complete an exercise.
#
# Exercises are based off (and directly taken from) the quick introduction to NumPy notebook.
#
# Different tasks will be detailed by comments or text.
#
# For further reference and resources, it's advised to check out the [NumPy documentation](https://numpy.org/devdocs/user/index.html).
#
# And if you get stuck, try searching for a question in the following format: "how to do XYZ with numpy", where XYZ is the function you want to leverage from NumPy.
# Import NumPy as its abbreviation 'np'
import numpy as np
# +
# Create a 1-dimensional NumPy array using np.array()
a1 = np.array([1,2,3])
# Create a 2-dimensional NumPy array using np.array()
a2 = np.array([[1,2,3], [4,5,6]])
# Create a 3-dimensional Numpy array using np.array()
a3 = np.array([[[1,2,3], [4,5,6]], [[7,8,9],[10,11,13]]])
# -
# Now we've you've created 3 different arrays, let's find details about them.
#
# Find the shape, number of dimensions, data type, size and type of each array.
# Attributes of 1-dimensional array (shape,
# number of dimensions, data type, size and type)
a1.shape, a1.ndim, a1.dtype, type(a1)
# Attributes of 2-dimensional array
a2.shape, a2.ndim, a2.dtype, type(a2)
# Attributes of 3-dimensional array
a3.shape, a3.ndim, a3.dtype, type(a3)
# Import pandas and create a DataFrame out of one
# of the arrays you've created
import pandas as pd
df_2 = pd.DataFrame(a2)
# Create an array of shape (10, 2) with only ones
ones = np.ones([10,2])
ones
# Create an array of shape (7, 2, 3) of only zeros
zeros = np.zeros([7,2,3])
zeros
zeros.shape
# Create an array within a range of 0 and 100 with step 3
my_array = np.arange(0, 100, 3)
my_array
# Create a random array with numbers between 0 and 10 of size (7, 2)
random_array = np.random.randint(10, size=(7,2))
random_array
# Create a random array of floats between 0 & 1 of shape (3, 5)
random_array = np.random.random((3,5))
random_array
# +
# Set the random seed to 42
np.random.seed(42)
# Create a random array of numbers between 0 & 10 of size (4, 6)
random_array = np.random.randint(0, 10, size=(4,6))
random_array
# -
# Run the cell above again, what happens?
#
# Are the numbers in the array different or the same? Why do think this is?
# +
# Create an array of random numbers between 1 & 10 of size (3, 7)
# and save it to a variable
random_array = np.random.randint(0, 10, size=(3,7))
random_array
# Find the unique numbers in the array you just created
np.unique(random_array)
# -
# Find the 0'th index of the latest array you created
random_array[0]
# Get the first 2 rows of latest array you created
random_array[:2, :]
# Get the first 2 values of the first 2 rows of the latest array
random_array[:2, :2]
# Create a random array of numbers between 0 & 10 and an array of ones
# both of size (3, 5), save them both to variables
random_array = np.random.randint(0, 10, size = (3,5))
ones = np.ones([3,5])
random_array
ones
# Add the two arrays together
random_array + ones
# Create another array of ones of shape (5, 3)
ones = np.ones([5,3])
# Try add the array of ones and the other most recent array together
random_array + ones
# When you try the last cell, it produces an error. Why do think this is?
#
# How would you fix it?
random_array.shape
ones.shape
random_array + ones.reshape([3,5])
# ---------
# Create another array of ones of shape (3, 5)
ones = np.ones([3,5])
random_array
ones
# Subtract the new array of ones from the other most recent array
random_array - ones
# Multiply the ones array with the latest array
random_array * ones
# Take the latest array to the power of 2 using '**'
random_array ** 2
# Do the same thing with np.square()
np.square(random_array)
# Find the mean of the latest array using np.mean()
np.mean(random_array)
# Find the maximum of the latest array using np.max()
np.max(random_array)
# Find the minimum of the latest array using np.min()
np.min(random_array)
# Find the standard deviation of the latest array
np.std(random_array)
# Find the variance of the latest array
np.var(random_array)
random_array
# Reshape the latest array to (3, 5, 1)
random_array.reshape([3,5,1])
# Transpose the latest array
random_array.T
# What does the transpose do?
# Create two arrays of random integers between 0 to 10
# one of size (3, 3) the other of size (3, 2)
a1 = np.random.randint(10, size=(3,3))
a2 = np.random.randint(10, size=(3,2))
a1
a2
# Perform a dot product on the two newest arrays you created
a1.dot(a2)
# Create two arrays of random integers between 0 to 10
# both of size (4, 3)
a1 = np.random.randint(10, size=(4,3))
a2 = np.random.randint(10, size=(4,3))
# Perform a dot product on the two newest arrays you created
a1.dot(a2)
# It doesn't work. How would you fix it?
# Take the latest two arrays, perform a transpose on one of them and then perform
# a dot product on them both
a2 = a2.T
a1.dot(a2)
# Notice how performing a transpose allows the dot product to happen.
#
# Why is this?
#
# Checking out the documentation on [`np.dot()`](https://docs.scipy.org/doc/numpy/reference/generated/numpy.dot.html) may help, as well as reading [Math is Fun's guide on the dot product](https://www.mathsisfun.com/algebra/vectors-dot-product.html).
#
# Let's now compare arrays.
# Create two arrays of random integers between 0 & 10 of the same shape
# and save them to variables
a1 = np.random.randint(10, size=(2,3))
a2 = np.random.randint(10, size=(2,3))
a1
a2
# Compare the two arrays with '>'
a1 > a2
# What happens when you compare the arrays with `>`?
# Compare the two arrays with '>='
a1 >= a2
# Find which elements of the first array are greater than 7
a1 > 7
# Which parts of each array are equal? (try using '==')
a1 == a2
# Sort one of the arrays you just created in ascending order
np.sort(a1)
a1.sort()
a1
a1
# Sort the indexes of one of the arrays you just created
a1.argsort()
# Find the index with the maximum value in one of the arrays you've created
a1.argmax()
# Find the index with the minimum value in one of the arrays you've created
a1.argmin()
a1
# Find the indexes with the maximum values down the 1st axis (axis=1)
# of one of the arrays you created
a1.argmax(axis=1)
# Find the indexes with the minimum values across the 0th axis (axis=0)
# of one of the arrays you created
a1.argmin(axis=0)
# Create an array of normally distributed random numbers
np.random.randn(3,5)
# Create an array with 10 evenly spaced numbers between 1 and 100
np.arange(1,100, 10)
# ## Extensions
#
# For more exercises, check out the [NumPy quickstart tutorial](https://docs.scipy.org/doc/numpy-1.15.0/user/quickstart.html). A good practice would be to read through it and for the parts you find interesting, add them into the end of this notebook.
#
# Pay particular attention to the section on broadcasting. And most importantly, get hands-on with the code as much as possible. If in dobut, run the code, see what it does.
#
# The next place you could go is the [Stack Overflow page for the top questions and answers for NumPy](https://stackoverflow.com/questions/tagged/numpy?sort=MostVotes&edited=true). Often, you'll find some of the most common and useful NumPy functions here. Don't forget to play around with the filters! You'll likely find something helpful here.
#
# Finally, as always, remember, the best way to learn something new is to try it. And try it relentlessly. If you get interested in some kind of NumPy function, asking yourself, "I wonder if NumPy could do that?", go and find out.
| Complete Machine Learning and Data Science - Zero to Mastery - AN/07.NumPy/numpy-exercises-MySolution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/probml/probml-notebooks/blob/main/notebooks/hbayes_binom_rats_pymc3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="JzJY-hRWTwYg"
# We fit a hierarchical beta-binomial model to some count data derived from rat survival. (In the book, we motivate this in terms of covid incidence rates.)
# Based on https://docs.pymc.io/notebooks/GLM-hierarchical-binominal-model.html
#
# + id="Kq6ayLpdTtKZ"
import sklearn
import scipy.stats as stats
import scipy.optimize
import matplotlib.pyplot as plt
import seaborn as sns
import time
import numpy as np
import os
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/"} id="pU2KC_mp_5c0" outputId="04dedabf-d32b-4300-9d8a-60ddc34c830a"
try:
import pymc3 as pm
except ModuleNotFoundError:
# %pip install -qq pymc3
import pymc3 as pm
print(pm.__version__)
try:
import arviz as az
except ModuleNotFoundError:
# %pip install -qq arviz
import arviz as az
print(az.__version__)
# + id="NRAKXCqDULZf"
import matplotlib.pyplot as plt
import scipy.stats as stats
import numpy as np
import pandas as pd
#import seaborn as sns
try:
import pymc3 as pm
except ModuleNotFoundError:
# %pip install -qq pymc3
import pymc3 as pm
try:
import arviz as az
except ModuleNotFoundError:
# %pip install -qq arviz
import arviz as az
try:
import theano.tensor as tt
except ModuleNotFoundError:
# %pip install -qq theano
import theano.tensor as tt
# + id="GbnW1XTcUcvu"
np.random.seed(123)
# rat data (BDA3, p. 102)
y = np.array(
[
0,
0,
0,
0,
0,
0,
0,
0,
0,
0,
0,
0,
0,
0,
1,
1,
1,
1,
1,
1,
1,
1,
2,
2,
2,
2,
2,
2,
2,
2,
2,
1,
5,
2,
5,
3,
2,
7,
7,
3,
3,
2,
9,
10,
4,
4,
4,
4,
4,
4,
4,
10,
4,
4,
4,
5,
11,
12,
5,
5,
6,
5,
6,
6,
6,
6,
16,
15,
15,
9,
4,
]
)
n = np.array(
[
20,
20,
20,
20,
20,
20,
20,
19,
19,
19,
19,
18,
18,
17,
20,
20,
20,
20,
19,
19,
18,
18,
25,
24,
23,
20,
20,
20,
20,
20,
20,
10,
49,
19,
46,
27,
17,
49,
47,
20,
20,
13,
48,
50,
20,
20,
20,
20,
20,
20,
20,
48,
19,
19,
19,
22,
46,
49,
20,
20,
23,
19,
22,
20,
20,
20,
52,
46,
47,
24,
14,
]
)
N = len(n)
# + colab={"base_uri": "https://localhost:8080/", "height": 236} id="QDnhJtbDUhZA" outputId="3a11b362-7101-491d-c014-b08e844d6da6"
def logp_ab(value):
"""prior density"""
return tt.log(tt.pow(tt.sum(value), -5 / 2))
with pm.Model() as model:
# Uninformative prior for alpha and beta
ab = pm.HalfFlat("ab", shape=2, testval=np.asarray([1.0, 1.0]))
pm.Potential("p(a, b)", logp_ab(ab))
alpha = pm.Deterministic("alpha", ab[0])
beta = pm.Deterministic("beta", ab[1])
X = pm.Deterministic("X", tt.log(ab[0] / ab[1]))
Z = pm.Deterministic("Z", tt.log(tt.sum(ab)))
theta = pm.Beta("theta", alpha=ab[0], beta=ab[1], shape=N)
p = pm.Binomial("y", p=theta, observed=y, n=n)
# trace = pm.sample(1000, tune=2000, target_accept=0.95)
trace = pm.sample(1000, tune=500)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="euqGyJN8Lhqo" outputId="e0185832-1cea-461a-ae0b-164a4802a9e1"
az.plot_trace(trace)
plt.savefig("hbayes_binom_rats_trace.png", dpi=300)
print(az.summary(trace))
# + colab={"base_uri": "https://localhost:8080/"} id="jT8KC06JMRJX" outputId="bda8534c-9387-41f2-9184-b132e182790c"
# !ls
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="4dgSQ2vAUlc-" outputId="8b5384d9-d2ae-444d-bb1d-988e3a32513d"
J = len(n)
post_mean = np.zeros(J)
samples = trace[theta]
post_mean = np.mean(samples, axis=0)
print("post mean")
print(post_mean)
alphas = trace["alpha"]
betas = trace["beta"]
alpha_mean = np.mean(alphas)
beta_mean = np.mean(betas)
hyper_mean = alpha_mean / (alpha_mean + beta_mean)
print("hyper mean")
print(hyper_mean)
mle = y / n
pooled_mle = np.sum(y) / np.sum(n)
print("pooled mle")
print(pooled_mle)
# axes = az.plot_forest(
# trace, var_names='theta', credible_interval=0.95, combined=True, colors='cycle')
axes = az.plot_forest(trace, var_names="theta", hdi_prob=0.95, combined=True, colors="cycle")
y_lims = axes[0].get_ylim()
axes[0].vlines(hyper_mean, *y_lims)
plt.savefig("hbayes_binom_rats_forest95.pdf", dpi=300)
J = len(n)
fig, axs = plt.subplots(4, 1, figsize=(10, 10))
plt.subplots_adjust(hspace=0.3)
axs = np.reshape(axs, 4)
xs = np.arange(J)
ax = axs[0]
ax.bar(xs, y)
ax.set_title("Number of postives")
ax = axs[1]
ax.bar(xs, n)
ax.set_title("Group size")
ax = axs[2]
ax.bar(xs, mle)
ax.set_ylim(0, 0.5)
ax.hlines(pooled_mle, 0, J, "r", lw=3)
ax.set_title("MLE (red line = pooled)")
ax = axs[3]
ax.bar(xs, post_mean)
ax.hlines(hyper_mean, 0, J, "r", lw=3)
ax.set_ylim(0, 0.5)
ax.set_title("Posterior mean (red line = hparam)")
plt.savefig("hbayes_binom_rats_barplot.pdf", dpi=300)
| notebooks/book2/03/hbayes_binom_rats_pymc3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (flow-master)
# language: python
# name: flow-maser
# ---
# # Tutorial 03: Running RLlib Experiments
#
# This tutorial walks you through the process of running traffic simulations in Flow with trainable RLlib-powered agents. Autonomous agents will learn to maximize a certain reward over the rollouts, using the [**RLlib**](https://ray.readthedocs.io/en/latest/rllib.html) library ([citation](https://arxiv.org/abs/1712.09381)) ([installation instructions](https://flow.readthedocs.io/en/latest/flow_setup.html#optional-install-ray-rllib)). Simulations of this form will depict the propensity of RL agents to influence the traffic of a human fleet in order to make the whole fleet more efficient (for some given metrics).
#
# In this exercise, we simulate an initially perturbed single lane ring road, where we introduce a single autonomous vehicle. We witness that, after some training, that the autonomous vehicle learns to dissipate the formation and propagation of "phantom jams" which form when only human driver dynamics are involved.
#
# ## 1. Components of a Simulation
# All simulations, both in the presence and absence of RL, require two components: a *scenario*, and an *environment*. Scenarios describe the features of the transportation network used in simulation. This includes the positions and properties of nodes and edges constituting the lanes and junctions, as well as properties of the vehicles, traffic lights, inflows, etc... in the network. Environments, on the other hand, initialize, reset, and advance simulations, and act as the primary interface between the reinforcement learning algorithm and the scenario. Moreover, custom environments may be used to modify the dynamical features of an scenario. Finally, in the RL case, it is in the *environment* that the state/action spaces and the reward function are defined.
#
# ## 2. Setting up a Scenario
# Flow contains a plethora of pre-designed scenarios used to replicate highways, intersections, and merges in both closed and open settings. All these scenarios are located in flow/scenarios. For this exercise, which involves a single lane ring road, we will use the scenario `LoopScenario`.
#
# ### 2.1 Setting up Scenario Parameters
#
# The scenario mentioned at the start of this section, as well as all other scenarios in Flow, are parameterized by the following arguments:
# * name
# * vehicles
# * net_params
# * initial_config
#
# These parameters are explained in detail in exercise 1. Moreover, all parameters excluding vehicles (covered in section 2.2) do not change from the previous exercise. Accordingly, we specify them nearly as we have before, and leave further explanations of the parameters to exercise 1.
#
# One important difference between SUMO and RLlib experiments is that, in RLlib experiments, the scenario classes are not imported, but rather called via their *string* names which (for serializtion and execution purposes) must be located within `flow/scenarios/__init__.py`. To check which scenarios are currently available, we execute the below command.
# +
import flow.scenarios as scenarios
print(scenarios.__all__)
# -
# Accordingly, to use the ring road scenario for this tutorial, we specify its (string) names as follows:
# ring road scenario class
scenario_name = "LoopScenario"
# Another difference between SUMO and RLlib experiments is that, in RLlib experiments, the scenario classes do not need to be defined; instead users should simply name the scenario class they wish to use. Later on, an environment setup module will import the correct scenario class based on the provided names.
# +
# input parameter classes to the scenario class
from flow.core.params import NetParams, InitialConfig
# name of the scenario
name = "training_example"
# network-specific parameters
from flow.scenarios.loop import ADDITIONAL_NET_PARAMS
net_params = NetParams(additional_params=ADDITIONAL_NET_PARAMS)
# initial configuration to vehicles
initial_config = InitialConfig(spacing="uniform", perturbation=1)
# -
# ### 2.2 Adding Trainable Autonomous Vehicles
# The `Vehicles` class stores state information on all vehicles in the network. This class is used to identify the dynamical features of a vehicle and whether it is controlled by a reinforcement learning agent. Morover, information pertaining to the observations and reward function can be collected from various `get` methods within this class.
#
# The dynamics of vehicles in the `Vehicles` class can either be depicted by sumo or by the dynamical methods located in flow/controllers. For human-driven vehicles, we use the IDM model for acceleration behavior, with exogenous gaussian acceleration noise with std 0.2 m/s2 to induce perturbations that produce stop-and-go behavior. In addition, we use the `ContinousRouter` routing controller so that the vehicles may maintain their routes closed networks.
#
# As we have done in exercise 1, human-driven vehicles are defined in the `Vehicles` class as follows:
# +
# vehicles class
from flow.core.params import VehicleParams
# vehicles dynamics models
from flow.controllers import IDMController, ContinuousRouter
vehicles = VehicleParams()
vehicles.add("human",
acceleration_controller=(IDMController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=21)
# -
# The above addition to the `Vehicles` class only accounts for 21 of the 22 vehicles that are placed in the network. We now add an additional trainable autuonomous vehicle whose actions are dictated by an RL agent. This is done by specifying an `RLController` as the acceleraton controller to the vehicle.
from flow.controllers import RLController
# Note that this controller serves primarirly as a placeholder that marks the vehicle as a component of the RL agent, meaning that lane changing and routing actions can also be specified by the RL agent to this vehicle.
#
# We finally add the vehicle as follows, while again using the `ContinuousRouter` to perpetually maintain the vehicle within the network.
vehicles.add(veh_id="rl",
acceleration_controller=(RLController, {}),
routing_controller=(ContinuousRouter, {}),
num_vehicles=1)
# ## 3. Setting up an Environment
#
# Several environments in Flow exist to train RL agents of different forms (e.g. autonomous vehicles, traffic lights) to perform a variety of different tasks. The use of an environment allows us to view the cumulative reward simulation rollouts receive, along with to specify the state/action spaces.
#
# Envrionments in Flow are parametrized by three components:
# * env_params
# * sumo_params
# * scenario
#
# ### 3.1 SumoParams
# `SumoParams` specifies simulation-specific variables. These variables include the length of any simulation step and whether to render the GUI when running the experiment. For this example, we consider a simulation step length of 0.1s and activate the GUI.
#
# **Note** For training purposes, it is highly recommanded to deactivate the GUI in order to avoid global slow down. In such case, one just needs to specify the following: `render=False`
# +
from flow.core.params import SumoParams
sumo_params = SumoParams(sim_step=0.1, render=False)
# -
# ### 3.2 EnvParams
#
# `EnvParams` specifies environment and experiment-specific parameters that either affect the training process or the dynamics of various components within the scenario. For the environment `WaveAttenuationPOEnv`, these parameters are used to dictate bounds on the accelerations of the autonomous vehicles, as well as the range of ring lengths (and accordingly network densities) the agent is trained on.
#
# Finally, it is important to specify here the *horizon* of the experiment, which is the duration of one episode (during which the RL-agent acquire data).
# +
from flow.core.params import EnvParams
# Define horizon as a variable to ensure consistent use across notebook
HORIZON=100
env_params = EnvParams(
# length of one rollout
horizon=HORIZON,
additional_params={
# maximum acceleration of autonomous vehicles
"max_accel": 1,
# maximum deceleration of autonomous vehicles
"max_decel": 1,
# bounds on the ranges of ring road lengths the autonomous vehicle
# is trained on
"ring_length": [220, 270],
},
)
# -
# ### 3.3 Initializing a Gym Environment
#
# Now, we have to specify our Gym Environment and the algorithm that our RL agents will use. To specify the environment, one has to use the environment's name (a simple string). A list of all environment names is located in `flow/envs/__init__.py`. The names of available environments can be seen below.
# +
import flow.envs as flowenvs
print(flowenvs.__all__)
# -
# We will use the environment "WaveAttenuationPOEnv", which is used to train autonomous vehicles to attenuate the formation and propagation of waves in a partially observable variable density ring road. To create the Gym Environment, the only necessary parameters are the environment name plus the previously defined variables. These are defined as follows:
env_name = "WaveAttenuationPOEnv"
# ### 3.4 Setting up Flow Parameters
#
# RLlib and rllab experiments both generate a `params.json` file for each experiment run. For RLlib experiments, the parameters defining the Flow scenario and environment must be stored as well. As such, in this section we define the dictionary `flow_params`, which contains the variables required by the utility function `make_create_env`. `make_create_env` is a higher-order function which returns a function `create_env` that initializes a Gym environment corresponding to the Flow scenario specified.
# Creating flow_params. Make sure the dictionary keys are as specified.
flow_params = dict(
# name of the experiment
exp_tag=name,
# name of the flow environment the experiment is running on
env_name=env_name,
# name of the scenario class the experiment uses
scenario=scenario_name,
# simulator that is used by the experiment
simulator='traci',
# sumo-related parameters (see flow.core.params.SumoParams)
sim=sumo_params,
# environment related parameters (see flow.core.params.EnvParams)
env=env_params,
# network-related parameters (see flow.core.params.NetParams and
# the scenario's documentation or ADDITIONAL_NET_PARAMS component)
net=net_params,
# vehicles to be placed in the network at the start of a rollout
# (see flow.core.vehicles.Vehicles)
veh=vehicles,
# (optional) parameters affecting the positioning of vehicles upon
# initialization/reset (see flow.core.params.InitialConfig)
initial=initial_config
)
# ## 4 Running RL experiments in Ray
#
# ### 4.1 Import
#
# First, we must import modules required to run experiments in Ray. The `json` package is required to store the Flow experiment parameters in the `params.json` file, as is `FlowParamsEncoder`. Ray-related imports are required: the PPO algorithm agent, `ray.tune`'s experiment runner, and environment helper methods `register_env` and `make_create_env`.
# +
import json
import ray
try:
from ray.rllib.agents.agent import get_agent_class
except ImportError:
from ray.rllib.agents.registry import get_agent_class
from ray.tune import run_experiments
from ray.tune.registry import register_env
from flow.utils.registry import make_create_env
from flow.utils.rllib import FlowParamsEncoder
# -
# ### 4.2 Initializing Ray
# Here, we initialize Ray and experiment-based constant variables specifying parallelism in the experiment as well as experiment batch size in terms of number of rollouts. `redirect_output` sends stdout and stderr for non-worker processes to files if True.
# +
# number of parallel workers
N_CPUS = 2
# number of rollouts per training iteration
N_ROLLOUTS = 1
ray.init(redirect_output=True, num_cpus=N_CPUS)
# -
# ### 4.3 Configuration and Setup
# Here, we copy and modify the default configuration for the [PPO algorithm](https://arxiv.org/abs/1707.06347). The agent has the number of parallel workers specified, a batch size corresponding to `N_ROLLOUTS` rollouts (each of which has length `HORIZON` steps), a discount rate $\gamma$ of 0.999, two hidden layers of size 16, uses Generalized Advantage Estimation, $\lambda$ of 0.97, and other parameters as set below.
#
# Once `config` contains the desired parameters, a JSON string corresponding to the `flow_params` specified in section 3 is generated. The `FlowParamsEncoder` maps objects to string representations so that the experiment can be reproduced later. That string representation is stored within the `env_config` section of the `config` dictionary. Later, `config` is written out to the file `params.json`.
#
# Next, we call `make_create_env` and pass in the `flow_params` to return a function we can use to register our Flow environment with Gym.
# +
# The algorithm or model to train. This may refer to "
# "the name of a built-on algorithm (e.g. RLLib's DQN "
# "or PPO), or a user-defined trainable function or "
# "class registered in the tune registry.")
alg_run = "PPO"
agent_cls = get_agent_class(alg_run)
config = agent_cls._default_config.copy()
config["num_workers"] = N_CPUS - 1 # number of parallel workers
config["train_batch_size"] = HORIZON * N_ROLLOUTS # batch size
config["gamma"] = 0.999 # discount rate
config["model"].update({"fcnet_hiddens": [16, 16]}) # size of hidden layers in network
config["use_gae"] = True # using generalized advantage estimation
config["lambda"] = 0.97
config["sgd_minibatch_size"] = min(16 * 1024, config["train_batch_size"]) # stochastic gradient descent
config["kl_target"] = 0.02 # target KL divergence
config["num_sgd_iter"] = 10 # number of SGD iterations
config["horizon"] = HORIZON # rollout horizon
# save the flow params for replay
flow_json = json.dumps(flow_params, cls=FlowParamsEncoder, sort_keys=True,
indent=4) # generating a string version of flow_params
config['env_config']['flow_params'] = flow_json # adding the flow_params to config dict
config['env_config']['run'] = alg_run
# Call the utility function make_create_env to be able to
# register the Flow env for this experiment
create_env, gym_name = make_create_env(params=flow_params, version=0)
# Register as rllib env with Gym
register_env(gym_name, create_env)
# -
# ### 4.4 Running Experiments
#
# Here, we use the `run_experiments` function from `ray.tune`. The function takes a dictionary with one key, a name corresponding to the experiment, and one value, itself a dictionary containing parameters for training.
trials = run_experiments({
flow_params["exp_tag"]: {
"run": alg_run,
"env": gym_name,
"config": {
**config
},
"checkpoint_freq": 1, # number of iterations between checkpoints
"checkpoint_at_end": True, # generate a checkpoint at the end
"max_failures": 999,
"stop": { # stopping conditions
"training_iteration": 1, # number of iterations to stop after
},
},
})
# ### 4.5 Visualizing the results
#
# The simulation results are saved within the `ray_results/training_example` directory (we defined `training_example` at the start of this tutorial). The `ray_results` folder is by default located at your root `~/ray_results`.
#
# You can run `tensorboard --logdir=~/ray_results/training_example` (install it with `pip install tensorboard`) to visualize the different data outputted by your simulation.
#
# For more instructions about visualizing, please see `tutorial05_visualize.ipynb`.
# ### 4.6 Restart from a checkpoint / Transfer learning
#
# If you wish to do transfer learning, or to resume a previous training, you will need to start the simulation from a previous checkpoint. To do that, you can add a `restore` parameter in the `run_experiments` argument, as follows:
#
# ```python
# trials = run_experiments({
# flow_params["exp_tag"]: {
# "run": alg_run,
# "env": gym_name,
# "config": {
# **config
# },
# "restore": "/ray_results/experiment/dir/checkpoint_50/checkpoint-50"
# "checkpoint_freq": 1,
# "checkpoint_at_end": True,
# "max_failures": 999,
# "stop": {
# "training_iteration": 1,
# },
# },
# })
# ```
#
# The `"restore"` path should be such that the `[restore]/.tune_metadata` file exists.
#
# There is also a `"resume"` parameter that you can set to `True` if you just wish to continue the training from a previously saved checkpoint, in case you are still training on the same experiment.
| tutorials/tutorial03_rllib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/VinayNooji/FourthBrain/blob/main/Copy_of_1_4_Multivariate_Linear_Regression_Python.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="ON4WbEBvk9Z9"
# # Multivariate Linear Regression
# ## Predicting House Price from Size and Number of Bedrooms
#
# This exercise is modified from the source: https://github.com/kaustubholpadkar/Predicting-House-Price-using-Multivariate-Linear-Regression
# + [markdown] id="RKxoZ5lknWll"
# ### Import Libraries
# + id="Lq6qDjhHnbak"
# %matplotlib inline
from mpl_toolkits.mplot3d import Axes3D
import numpy as np
import matplotlib.pyplot as plt
from sklearn.preprocessing import normalize
# + [markdown] id="W5b_-H-zn3_O"
# ### Read the House Data
#
# NumPy provides the `genfromtxt()` function to load data from a text file into an array. As usual, each row in that file represents a different data point, observation, or example. Its components are separated by commas, so we set the `delimiter` parameter in `genfromtxt()`'s argument list to `','`.
# + id="T1a23JYWn5oM"
data = np.genfromtxt('https://raw.githubusercontent.com/Tan-Moy/medium_articles/master/art2_multivariate_linear_regression/home.txt', delimiter=',')
# + [markdown] id="AK27bq1TnkbT"
# Examine the data's shape with the convenient `.shape` attribute. NumPy arrays and structures based on them all have it.
# + id="Ip2EQ_PGoQJO" colab={"base_uri": "https://localhost:8080/"} outputId="7e4026f0-370c-4c59-a3f3-d4dd2d4709d1"
data
# + [markdown] id="vFbjwHfjoWpT"
# By convention, the last element in an array's `.shape` tuple refers to its number of columns. Our dataset thus has 47 rows and 3 columns.
# + [markdown] id="_cWMna08pDIL"
# Let's visually inspect the first 5 rows of the data
# + id="otHHo0yCpJqj" colab={"base_uri": "https://localhost:8080/"} outputId="9d0e2215-6de2-4170-80f0-5182e94c6d60"
data[:5, :]
# + [markdown] id="IzyXYXMEpYVv"
# The columns from left to right indicate each house's:
#
# 0. Floor area in square feet
# 1. Number of bedrooms
# 2. Price in dollars
#
# The first 2 are our features, i.e. the attributes of the data which we feed into our model to make predictions. The last is our target, the value which we're attempting to predict.
# + [markdown] id="ydBEcRFeiDtS"
# ### Normalize Data
#
# Normalization of feature (and for regression problems, target) columns can mitigate problems that can arise from loss function gradients becoming too large or small, and thus help a model converge faster. There are several ways to normalize data. Here, we're going to rescale each column so that its L2 norm is 1, i.e. the sum of the squares of each element in a given column will be 1.
#
# We'll use the `normalize()` function from scikit-learn's preprocessing module. We'll need to specify the array axis along which we'll be calculating our norms. For aggregating functions (functions like mean and standard deviation) in NumPy and related libraries, the `axis` (or in the case of PyTorch, `dim`) parameter refers to the axis along which the aggregation is performed. Our data is a 2D array. We want to normalize each column, which means that within a given column, we aggregate the numbers from each row. For a 2D array, the 0 axis corresponds to the rows and the 1 axis corresponds to the columns. Thus, we pass `axis=0` into our call to the `normalize()` function.
# + id="y1BwuHufiHJ_"
data = normalize(data, axis=0)
# + [markdown] id="evCdby_Ryh_d"
# Let's inspect the first 5 rows of our data again.
# + id="6mWobVOAymvR" colab={"base_uri": "https://localhost:8080/"} outputId="6c4fd45d-802f-4040-e5ef-e8352be71877"
data[:5, :]
# + [markdown] id="7IYZy8_oywl6"
# The numbers look quite different from before. The floor areas, which were in square feet and on the order of $10^{3}$, are now represented by normalized figures on the order of $10^{-1}$ or just below. Similarly, the prices have been reduced from dollar figures on the order of $10^{5}$.
# + [markdown] id="-BYea-NKzhMb"
# Let's verify that the `normalize()` function did its job properly
# + id="LUjFamH_zsaN" colab={"base_uri": "https://localhost:8080/"} outputId="29d2a014-50a0-402d-eb6b-0ec89594f20c"
# Floor area column
np.sum(np.square(data[:,0]))
# + id="_6QJv4k60Fxl" colab={"base_uri": "https://localhost:8080/"} outputId="e891f291-27c7-4d65-9281-627b8e95975b"
# Bedroom count column
np.sum(np.square(data[:,1]))
# + id="gTczjYHN0Fhu" colab={"base_uri": "https://localhost:8080/"} outputId="e7255d26-8d08-494d-cdba-f469ce43595a"
# Price column
np.sum(np.square(data[:,2]))
# + [markdown] id="O5K5K8pG0VEr"
# To within a roundoff error, our data has, indeed, been normalized properly.
# + [markdown] id="8rMv-L53odN5"
# ### Separate Data into X and Y
#
# I.e., separate the target from the features
# + id="uop_AFQSolHl" colab={"base_uri": "https://localhost:8080/"} outputId="6f2f0e41-29b8-460d-bf60-e8da057ab5e4"
X = data[:, 0:2]
Y = data[:, 2:]
print(Y.shape)
print(X.shape)
# + [markdown] id="wPSTKDAXpLAN"
# ### Visualize the Data
# + id="eqSVKtKBWv9-" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="635ebb31-8ca4-44f0-a416-84c171ebc108"
# Fixing random state for reproducibility
np.random.seed(19680801)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
xs = X[:, 0]
ys = X[:, 1]
zs = Y
ax.scatter(xs, ys, zs)
ax.set_xlabel('size')
ax.set_ylabel('bedroom')
ax.set_zlabel('price')
plt.show()
# + [markdown] id="1qEiGCpL1iae"
# It appears that our data points are distributed closely to a 2D plane. Multivariate linear regression should give us reasonable price predictions.
# + [markdown] id="pf38YSVrIMaQ"
# Now that we've explored our data, let's prepare the model.
# + [markdown] id="Yrjucdifq_mt"
# ### Hyperparameters
#
# First, let's choose the hyperparameters (which the model won't learn, but which will influence the model's outcome and performance). In most machine learning problems, the objective is to learn the values of parameters (which interact with the features of any data point/observation/example fed into the model) such that a loss function (a penalty for predictions in the training data not matching their ground truth values) is minimized. One typically uses numerical approximations of gradients of this loss function to minimize it.
#
# The three most common methods for updating the loss function gradients and learned parameters are:
#
# 1. Batch gradient descent - Each update uses the entire dataset
# 2. Stochastic gradient descent - The gradients and parameters are updated after every training example
# 3. Mini-batch gradient descent - Something in between batch and stochastic gradient descent: The model reads in a subset of the full dataset before updating the gradients and parameters. Mini-batch sizes in powers of 2 are preferable.
# + id="OwVfkk3RrC_I"
# Batch gradient descent
learning_rate = 0.09
max_iteration = 500
# + id="RSXPA9L25UGl"
# Stochastic gradient descent
s_learning_rate = 0.06
s_max_iteration = 500
# + id="IeY-_RqF5V5v"
# Mini-Batch gradient descent
mb_learning_rate = 0.09
mb_max_iteration = 500
batch_size = 16
# + [markdown] id="EzXaG70OrZtn"
# ### Exercise: Parameters
#
# Initialize the arrays of theta parameters (one for each of the gradient descent methods above) which 2D arrays of zeros. Each row should correspond to a feature (column of `X`), with a bias term added (which will allow us to move our plane up or down the price, i.e. z axis). There should only be one element per row.
#
# You may find this function useful: [np.zeros()](https://numpy.org/doc/stable/reference/generated/numpy.zeros.html)
# + id="2INJM9AIrcBu"
### START CODE HERE ###
theta = np.zeros((X.shape[1]+1, 1))
s_theta = np.zeros((X.shape[1]+1, 1))
mb_theta = np.zeros((X.shape[1]+1, 1))
#print(theta.shape, s_theta.shape, mb_theta.shape)
### END CODE HERE ###
# + [markdown] id="FPt2eKjBr1Lx"
# ### Exercise: Hypothesis
#
# Define a function which adds a column of ones (to complement the offset/bias parameter) to the features `X` of the dataset and computes a matrix multiplication between it and an array of learned parameters `theta`. Pay attention to the shapes of `X` and `theta`; matrix multiplication is not commutative.
#
# More generally, a hypothesis function `h` takes in features `X` and learned parameters `theta` to make a prediction, commonly denoted $\hat{Y}$. The loss function (which you'll implement in the code cell immediately following the one for the hypothesis function) is a measure of the difference between your model's predicted target values and actual, ground truth target values.
#
# You may find these functions useful: [np.ones()](https://numpy.org/doc/stable/reference/generated/numpy.ones.html), [np.dot()](https://numpy.org/doc/stable/reference/generated/numpy.dot.html), [np.matmul()](https://numpy.org/doc/stable/reference/generated/numpy.matmul.html)
# + id="enVE4tHWsBBF"
def h (theta, X) :
### START CODE HERE ###
# Add a 0th column of ones to X
tempX = np.ones((X.shape[0], X.shape[1]+1))
tempX[:, 1:] = X
# Return a matrix multiplication of your augmented X and theta
return np.matmul(tempX, theta)
### END CODE HERE ###
# + [markdown] id="IA9I6vaXs7nb"
# ### Exercise: Loss Function
#
# We'll use **half of** the Mean Squared Error (MSE) for our loss function. The MSE is defined as
#
# $
# \textrm{MSE} = \frac{1}{m} \sum_{i=1}^m (Y_i - \hat{Y_i})^2,
# $
#
# where $i$ is a given training example and $m$ is the number of training examples, i.e. rows in our dataset. Recall from above that $\hat{Y}$ is a function of `theta` and `X`.
#
# You may find these functions useful: [np.average()](https://numpy.org/doc/stable/reference/generated/numpy.average.html) [np.mean()](https://numpy.org/doc/stable/reference/generated/numpy.mean.html), [np.square()](https://numpy.org/doc/stable/reference/generated/numpy.square.html)
# + id="3KOQVUjBs-Ub"
def loss (theta, X, Y) :
### START CODE HERE ###
# Return half of the mean squared error between the actual Y and the predicted Y
# Recall that the predicted Y is a function of theta and X
return np.average(np.square(Y - h(theta, X)))/2.0
### END CODE HERE ###
# + colab={"base_uri": "https://localhost:8080/"} id="jxV8aZ5UCiHV" outputId="ebb66a15-5b4c-4f4d-c908-99ab57c3ec99"
print('Mean = ', np.mean(np.square(Y - h(theta, X))), 'Average = ', np.average(np.square(Y - h(theta, X))))
# + [markdown] id="7TE79NG-tyLG"
# ### Exercise: Calculate Gradients
#
# One can verify through straightforward (though somewhat tedious) multivariable calculus that gradient of the loss function $J$ with respect to the array of parameters $\theta$ is
#
# $
# \frac{\partial J}{\partial \theta} = - \frac{1}{m} X^T \cdot (Y - \hat{Y}).
# $
#
# For clarity, $m$ is the number of training examples, the superscript $T$ denotes a matrix transposition, and $\cdot$ denotes a matrix inner product (dot product).
#
# In code, $\frac{\partial J}{\partial \theta}$ is typically denoted `d_theta` or something similar, with the same logic applying to other intermediate parameters for the more complicated models you'll learn about later in this program.
# + id="yy9yxye4t0BJ"
def gradient (theta, X, Y) :
### START CODE HERE ###
# Make a copy of X with an extra initial column of ones to complement the bias parameter
tempX = np.ones((X.shape[0], X.shape[1] + 1))
tempX[:,1:] = X
# Compute d_theta according to the formula in the text cell above
# Make sure it retains the shape (number of features + 1, 1)
d_theta = - np.dot(tempX.T, (Y - h(theta, X)))/X.shape[0]
### END CODE HERE ###
return d_theta
# + [markdown] id="zLd-RcZ0qh0N"
# ### Now we define our gradient descent methods
#
# In general, we update our array of parameters $\theta$ with the formula
#
# $
# \theta := \theta - \alpha \frac{\partial J}{\partial \theta},
# $
#
# or in code,
#
# `theta = theta - learning_rate * d_theta`.
#
# The question is how often `d_theta` is calculated and thus how often `theta` is updated.
# + [markdown] id="jVo5TbKyu9KL"
# ### Exercise: Batch Gradient Descent
#
# Use the full dataset to update `d_theta`
# + id="fprg4-tcu-4v"
def gradient_descent (theta, X, Y, learning_rate, max_iteration, gap) :
cost = np.zeros(max_iteration)
for i in range(max_iteration) :
### START CODE HERE ###
# Use the full dataset to update d_theta
d_theta = gradient(theta, X, Y)
### END CODE HERE ###
theta = theta - learning_rate * d_theta
cost[i] = loss(theta, X, Y)
if i % gap == 0 :
print ('iteration : ', i, ' loss : ', loss(theta, X, Y), ' shape: ', d_theta.shape)
return theta, cost
# + [markdown] id="Si5f2m-9-krz"
# ### Exercise: Mini-Batch Gradient Descent
#
# Use a subset of the data of size `batch_size` to update `d_theta`
# + id="kCGUklWXyov-"
def minibatch_gradient_descent (theta, X, Y, learning_rate, max_iteration, batch_size, gap) :
cost = np.zeros(max_iteration)
for i in range(max_iteration) :
for j in range(0, X.shape[0], batch_size):
### START CODE HERE ###
# Use a subset of the data of size batch_size to update d_theta
d_theta = gradient(theta, X[j:batch_size+j, :], Y[j:batch_size+j, :])
### END CODE HERE ###
theta = theta - learning_rate * d_theta
cost[i] = loss(theta, X, Y)
if i % gap == 0 :
print ('iteration : ', i, ' loss : ', loss(theta, X, Y))
return theta, cost
# + [markdown] id="epWfjT9R-sk3"
# ### Stochastic Gradient Descent
#
# Update `d_theta` at every training example
# + id="cWDNv4BkvpvQ"
def stochastic_gradient_descent (theta, X, Y, learning_rate, max_iteration, gap) :
cost = np.zeros(max_iteration)
for i in range(max_iteration) :
for j in range(X.shape[0]):
### START CODE HERE ###
# Update d_theta at every training example
d_theta = gradient(theta, X[np.newaxis, j, :], Y[np.newaxis, j, :])
### END CODE HERE ###
theta = theta - learning_rate * d_theta
cost[i] = loss(theta, X, Y)
if i % gap == 0 :
print ('iteration : ', i, ' loss : ', loss(theta, X, Y))
return theta, cost
# + [markdown] id="uMIZFxUhwQZO"
# ### Train Model
# + id="-0okDGSCwUC7" colab={"base_uri": "https://localhost:8080/", "height": 381} outputId="cd400351-1b6b-4247-fa88-fad4607b5f9a"
theta, cost = gradient_descent (theta, X, Y, learning_rate, max_iteration, 100)
plt.xlabel("iteration")
plt.ylabel("cost")
plt.plot(range(0, max_iteration), cost)
# + id="YBJreRDKwtvI" colab={"base_uri": "https://localhost:8080/", "height": 381} outputId="bdb268ae-af1b-4b75-d736-38544a21ce0a"
s_theta, s_cost = stochastic_gradient_descent (s_theta, X, Y, s_learning_rate, s_max_iteration, 100)
plt.xlabel("iteration")
plt.ylabel("cost")
plt.plot(range(0, s_max_iteration), s_cost)
# + id="viSuvODWza-1" colab={"base_uri": "https://localhost:8080/", "height": 381} outputId="8c63c53e-a04b-4407-c700-5bc571326bc2"
mb_theta, mb_cost = minibatch_gradient_descent (mb_theta, X, Y, mb_learning_rate, mb_max_iteration, batch_size, 100)
plt.xlabel("iteration")
plt.ylabel("cost")
plt.plot(range(0, mb_max_iteration), mb_cost)
# + [markdown] id="MFp2MUlexVd5"
# ### Optimal values of Parameters using Trained Model
# + id="tcMQWkuvwYu7" colab={"base_uri": "https://localhost:8080/"} outputId="fc062602-4cad-434c-c6d2-ef66e3144e25"
theta
# + id="Orub1Wgdx0yc" colab={"base_uri": "https://localhost:8080/"} outputId="39d14437-306f-4a26-833f-7e16f9d08e89"
s_theta
# + id="TBrs3crW55Ka" colab={"base_uri": "https://localhost:8080/"} outputId="5a521e26-5894-4871-d567-563917e5605d"
mb_theta
# + [markdown] id="Pi99RMws736c"
# ### Cost vs Iteration Plots
# + id="SvzRaqIQ78Pl" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="d60ba5d2-d3f2-4a30-a5af-acfad58dc6f8"
#plot the cost
fig, ax = plt.subplots()
ax.plot(np.arange(max_iteration), cost, 'r')
ax.plot(np.arange(max_iteration), s_cost, 'b')
ax.plot(np.arange(max_iteration), mb_cost, 'g')
ax.legend(loc='upper right', labels=['batch gradient descent', 'stochastic gradient descent', 'mini-batch gradient descent'])
ax.set_xlabel('Iterations')
ax.set_ylabel('Cost')
ax.set_title('Error vs. Training Epoch')
plt.show()
# + [markdown] id="Dh6PmQOwxcKk"
# ### Visualize the Data
# + id="B0h7MJE0xcKo" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="e6d3b7ee-db87-423b-c7ef-1f23c1741c30"
# Fixing random state for reproducibility
np.random.seed(19680801)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
xs = X[:, 0]
ys = X[:, 1]
zs = Y
ax.scatter(xs, ys, zs)
ax.set_xlabel('size')
ax.set_ylabel('bedroom')
ax.set_zlabel('price')
x = y = np.arange(0, 0.3, 0.05)
xp, yp = np.meshgrid(x, y)
z = np.array([h(theta, np.array([[x,y]]))[0, 0] for x,y in zip(np.ravel(xp), np.ravel(yp))])
zp = z.reshape(xp.shape)
ax.plot_surface(xp, yp, zp, alpha=0.7)
plt.show()
# + id="GoZITHpln8TK" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="1147ba78-acee-4141-a7f8-24036bc316c0"
# Fixing random state for reproducibility
np.random.seed(19680801)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
xs = X[:, 0]
ys = X[:, 1]
zs = Y
ax.scatter(xs, ys, zs)
ax.set_xlabel('size')
ax.set_ylabel('bedroom')
ax.set_zlabel('price')
x = y = np.arange(0, 0.3, 0.05)
xp, yp = np.meshgrid(x, y)
z = np.array([h(s_theta, np.array([[x,y]]))[0, 0] for x,y in zip(np.ravel(xp), np.ravel(yp))])
zp = z.reshape(xp.shape)
ax.plot_surface(xp, yp, zp, alpha=0.7)
plt.show()
# + id="KUTo6lsyx_Uv" colab={"base_uri": "https://localhost:8080/", "height": 248} outputId="44cf6a5d-aee8-4235-df60-32bfbb410a13"
# Fixing random state for reproducibility
np.random.seed(19680801)
fig = plt.figure()
ax = fig.add_subplot(111, projection='3d')
x = y = np.arange(0, 0.3, 0.05)
xp, yp = np.meshgrid(x, y)
z = np.array([h(mb_theta, np.array([[x,y]]))[0, 0] for x,y in zip(np.ravel(xp), np.ravel(yp))])
zp = z.reshape(xp.shape)
ax.plot_surface(xp, yp, zp, alpha=0.7)
xs = X[:, 0]
ys = X[:, 1]
zs = Y
ax.scatter(xs, ys, zs)
ax.set_xlabel('size')
ax.set_ylabel('bedroom')
ax.set_zlabel('price')
plt.show()
# + [markdown] id="0pAaVPUztMZV"
# Based on the losses and plots after 500 iterations through the dataset, which gradient descent method appears to yield the best predictions for this particular problem?
# + [markdown] id="-U_2Yg_oTsG4"
# After a lot of trial and error, it appears schotastic gradient fits the data better.
| Copy_of_1_4_Multivariate_Linear_Regression_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from matplotlib import pyplot as plt
import matplotlib
import pandas as pd
import sys
print("Python Version: " + '{0[0]}.{0[1]}'.format(sys.version_info))
print("Matplotlib Version: " + matplotlib.__version__)
out1_roc_data_AvPb_micro_auc_f_path = "./out1_roc_data_AvPb_micro_auc_0.7249_CIs_0.7226_0.7274.txt"
out1_roc_data_AvPb_micro_auc_ = pd.read_csv(out1_roc_data_AvPb_micro_auc_f_path, delimiter = '\t')
out1_roc_data_AvPb_micro_auc_
plt.plot(out1_roc_data_AvPb_micro_auc_["0.000000"], out1_roc_data_AvPb_micro_auc_["0.000000.1"])
plt.title("out1_roc_data_AvPb_micro_auc")
out2_roc_data_AvPb_micro_auc_f_path = "./out2_roc_data_AvPb_micro_auc_0.7804_CIs_0.6952_0.8550.txt"
out2_roc_data_AvPb_micro_auc_ = pd.read_csv(out2_roc_data_AvPb_micro_auc_f_path, delimiter = '\t')
out2_roc_data_AvPb_micro_auc_
plt.plot(out2_roc_data_AvPb_micro_auc_["0.000000"], out2_roc_data_AvPb_micro_auc_["0.000000.1"])
plt.title("out2_roc_data_AvPb_micro_auc")
| Stage 2 Transfer Learning with DeepPATH/test_7483k/.ipynb_checkpoints/Untitled1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # K-means Clustering
# +
# import the library
# %matplotlib inline
import pandas as pd
import numpy as np
import collections
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.preprocessing import StandardScaler
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics.pairwise import euclidean_distances
# convert scientific notation to decimals
pd.set_option('display.float_format', lambda x: '%.2f' % x)
sns.set_style('whitegrid')
# -
# # Problem definition
# Clustering flights according to delay reasons.
# ____________
# # Load the Data
# ### Cleaned, Merged dataset can be downloaded from here: https://www.kaggle.com/arwasheraky/cleaned-flight-delays-2015
df_flights = pd.read_csv('../../Data/flightsmerged.csv', low_memory=False)
print(df_flights.columns)
df_flights.head()
# _________________
# # Feature Engineering
# +
# Keep just the delayed flights
df = df_flights[df_flights['CLASS'] == 'Delayed']
df = df.drop(columns = ['CANCELLATION_REASON','CANCELLED'])
# -
df = df.dropna()
df.shape
# ## Add a column
# +
# Average delay column
df['AVG_DELAY'] = (df['DEPARTURE_DELAY'] + df['ARRIVAL_DELAY']) / 2.0
df['AVG_DELAY'].head(10)
# -
# ## Filter the flights
# +
# Remove flights with avg. delay between [-1,1]
flights_to_remove = list(df[(df['AVG_DELAY'] > -1) & (df['AVG_DELAY'] < 1)].index)
new_df = df[~df.index.isin(flights_to_remove)]
new_df.shape
# -
plt.figure(figsize=(10,6))
plt.hist(new_df['AVG_DELAY'], bins=50, color='purple')
plt.xlabel("Average Delay")
plt.ylabel("Frequency")
plt.show()
# ## Select Model Columns
# +
# Select delay reasons columns
delay_reason = new_df[['AIR_SYSTEM_DELAY','SECURITY_DELAY', 'AIRLINE_DELAY', 'LATE_AIRCRAFT_DELAY','WEATHER_DELAY']]
delay_reason.mean()
# -
X_columns = ['AVG_DELAY'] + list(delay_reason.columns)
# +
# normalize the columns
df_norm = new_df.copy()
df_norm = df_norm[X_columns]
for col in X_columns:
df_norm[col] = new_df[col].astype(float)
df_norm[col] = StandardScaler().fit_transform(df_norm[col].values.reshape(-1, 1))
df_norm.head()
# -
# _______
# # Model Training
# +
k = 5
kmeans = KMeans(n_clusters=k).fit(df_norm.values)
print(set(kmeans.labels_))
print(collections.Counter(kmeans.labels_))
# -
df_results = new_df.copy()
df_norm['cluster'] = kmeans.labels_
df_results['cluster'] = kmeans.labels_
df_results = df_results.reset_index()
# ____________
# # Analyze the Results
# +
# print the most common flights in each cluster
for cluster in sorted(set(kmeans.labels_)):
print("Cluster",cluster," :",
collections.Counter(df_results[df_results['cluster']==cluster]['FLIGHT_NUMBER']).most_common(3))
colors = ['red','navy','orange','purple','green']
n_clusters = len(set(kmeans.labels_))
for col in X_columns:
print(col)
i = 1
plt.figure(figsize=(19,5))
for cluster in sorted(set(kmeans.labels_)):
plt.subplot(1, n_clusters, i)
plt.xlim([df_results[col].min(),df_results[col].max()])
plt.hist(df_results[df_results['cluster']==cluster][col], bins=10, label=str(cluster), alpha=0.7, color=colors[i-1])
plt.title("Cluster "+str(cluster))
i += 1
plt.tight_layout()
plt.show()
# -
# #### Conclusions:
# - Cluster 0: Flights have the highest delay avg, without specific delay reasons. The most contributing factor in the delay could be `Air System Problems`.
# - Cluster 1: Flights delayed mainly due to `Air System Problems`, and secondly `certain conditions in the Airline`
# - Cluster 2: Flights delayed mainly due to `Late Aircraft`
# - Cluster 3: Flights delayed mainly due to `Weather Conditions`, and secondly `Air System Problems`.
# - Cluster 4: Flights delayed mainly due to `Security Issues`.
# +
# for col in X_columns:
# print(col)
# j = 0
# for cluster in set(kmeans.labels_):
# plt.hist(df_results[df_results['cluster']==cluster][col], label="Cluster"+str(cluster), alpha=0.5, bins=20, color=colors[j])
# j+=1
# plt.legend()
# plt.show()
# -
# ___________
# # Model Evaluation
# +
# Inter-Cluster
centroids = []
for cluster in sorted(set(kmeans.labels_)):
centroids.append(df_norm[df_norm['cluster']==cluster][X_columns].mean().values)
distances = []
for c1 in centroids:
for c2 in centroids:
distances.append(euclidean_distances(c1.reshape(-1, 1), c2.reshape(-1, 1))[0][0])
print('Inter Cluster distance = ', np.mean(distances))
# +
## Takes a lot of time ...
# Intra-Cluster
distances = []
for cluster in sorted(set(kmeans.labels_)):
df_filter = df_norm[df_norm['cluster']==cluster]
centroid = df_filter[X_columns].mean().values
for k, v in df_filter[X_columns].iterrows():
distances.append(euclidean_distances(centroid.reshape(-1, 1), v.values.reshape(-1, 1))[0][0])
print('Intra Cluster distance = ', np.mean(distances))
# +
## Couldn't run it...
# Inertia
distances = []
for cluster in sorted(set(kmeans.labels_)):
df_filter = df_norm[df_norm['cluster']==cluster]
centroid = df_filter[X_columns].mean().values
for k, v in df_filter[X_columns].iterrows():
distances.append(euclidean_distances(centroid.reshape(1, -1), v.values.reshape(1, -1), squared=True)[0][0])
print('Inertia = ', np.sum(distances))
| Code/Unsupervised_Learning/flights-clustering-kmeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''data_process'': conda)'
# name: python3
# ---
# +
# step 3
from mne.time_frequency import tfr_morlet
from alive_progress import alive_bar
from hm_tools import *
from tqdm import tqdm
import nibabel as nib
import pandas as pd
import numpy as np
# import scipy.io
import h5py
import mne
import sys
import os
# 计算varibility的代 码要基于mne的epoch文件才可以实现。
subject_num = 34
mne_erp_loc = 'E:/workspace/Trust_Data_and_Results/Haoming/varibility/data/mne_erp_exp2/'
result_loc = 'E:/workspace/Trust_Data_and_Results/Haoming/varibility/result/varibilit_corr_20_exp2/'
circle = 5 # For exp2, circle=5. For Exp 3, circle=2
file_name = ['alpha'] # only calculate the alpha band
varibility_method = 'corr' # 'corr' or 'var'
# 时间窗的长度, 单位
time_window = 0.02
face_per_event = 10
# 需要分析的数据起止时间 第一位是开始的时间,第二位是结束的时间
involve_time = [-0.200, 1.0]
# the number of face in each event(condition)
face_num_per_condition = 10
# +
# 去除数据残缺的被试
bad_subject = np.array([3, 5, 15]) -1
all_subject = np.arange(0,34,1)
# 删掉特定元素
good_subject = np.setdiff1d(all_subject, bad_subject)
# +
def corr_matrix_insert_nan(input_data, nan_position, values, axis):
output_data = input_data
for iter_nan_position in nan_position:# insert nan in every nan position
# insert nan in two direction
output_data = np.insert(arr=output_data, obj=iter_nan_position, values=values, axis=axis[0])
output_data = np.insert(arr=output_data, obj=iter_nan_position, values=values, axis=axis[1])
return output_data
def varibility_corr(eeg_data, time_series, involve_time, window_length):# eeg_data is 2d (trials * time_series)
# 通过对比involve_time和epoch的时间序列来找到是第几位数字开始计算相关性
start_time = np.where(time_series == involve_time[0])[0][0]
end_time = np.where(time_series == involve_time[1])[0][0]
# 生成时间序列结果数组
time_series_var = []
# 对每个时间窗计算一次变异性
for iter_window in range(end_time - start_time):# 步长为1
# 获取每个时间窗的数据 trail
iter_data = eeg_data[:,(start_time + iter_window - round(window_length/2)) : (start_time + iter_window + round(window_length/2))]
# 对该矩阵计算相关系数
corr = np.corrcoef(iter_data)
# 获取对角线以下(k=-1)的下三角
low_triangle = np.tril(corr, k=-1)
# 对下三角求和
all_corr = np.sum(low_triangle)
# 计算下三角的数据点个数
corr_num = (eeg_data.shape[0]**2 - eeg_data.shape[0])/2
# 计算平均相关性
average_corr = all_corr/corr_num
# 通过1-计算差异性
average_var = 1 - average_corr
# 得到每个时间序列的结果
time_series_var.append(average_var)
return time_series_var #输出结果矩阵 time
# +
# preload to check information
# read the epoch data
eeg_epochs = mne.read_epochs(mne_erp_loc + '/subject_1' +'/' + file_name[0] + '_epo.fif', preload = True)
time_series = np.array(eeg_epochs.times)
# 检查event的数量
event_name = eeg_epochs.event_id.keys()
event_name = list(event_name)
event_num = len(event_name)
# check the number of eeg electrode
eeg_data = eeg_epochs.get_data()
electrode_num = eeg_data.shape[1]
# calculate how many power band
power_band_num = len(file_name)
# calculate how many good subject
good_subject_num = len(good_subject)
# check how many time point in the result
result_time_length = int(eeg_epochs.info['sfreq'] * (involve_time[1] - involve_time[0]))
# calculate the time point of each window
if varibility_method == 'corr':
window_length = eeg_epochs.info['sfreq'] * time_window
print('point of the window is:', window_length)
# + tags=["outputPrepend"]
# 平均变异性
all_rsa_result=[]
all_result = np.full((subject_num, power_band_num, event_num, circle, electrode_num, result_time_length),np.nan)
for iter_subject in good_subject:
# 生成每个被试的erp数据存放地址
subject_erp_loc = mne_erp_loc + '/subject_' + str(iter_subject + 1)
iter_subject_result = np.full((power_band_num, event_num, circle, electrode_num, result_time_length),np.nan)
# 包括3个波段,各循环一次
for iter_file in range(power_band_num):
# read the epoch data
eeg_epochs = mne.read_epochs(subject_erp_loc +'/' + file_name[iter_file] + '_epo.fif', preload = True)
iter_power_result = np.full((event_num, circle, electrode_num, result_time_length),np.nan)
# 对每一个event分别进行一次计算(可以大于2个) 两种计算方法
for iter_event in range(event_num):
# 提取每一个event的脑电数据
iter_event_eeg_data = eeg_epochs[event_name[iter_event]].get_data()
# Generate nan matrix for each event (circle * channel * time * trials * trials)
iter_event_result = np.full((circle, electrode_num, result_time_length), np.nan)
for iter_influence_step in range(circle):
# 提取每一个 influence step 的脑电数据
iter_influence_step_eeg_data = iter_event_eeg_data[10*iter_influence_step:10*iter_influence_step+10]
# 检查其中有没有nan,如果有就记录下nan的位置并且把nan位置的数据删除
nan_position, not_nan_position = find_1d_array_nan(iter_influence_step_eeg_data[:,0,0])
if len(nan_position) > 0:
iter_influence_step_eeg_data = iter_influence_step_eeg_data[not_nan_position]
# Generate nan matrix for each influence step (channel * time * trials * trials)
iter_influence_step_result=np.full((electrode_num, result_time_length), np.nan)
for iter_electrode in range(iter_influence_step_eeg_data.shape[1]):
if varibility_method == 'corr':
time_series_corr_matrix = varibility_corr(iter_influence_step_eeg_data[:,iter_electrode,:], time_series, involve_time, window_length)
time_series_corr_matrix = time_series_corr_matrix / np.mean(time_series_corr_matrix[0:int(np.abs(involve_time[0])*eeg_epochs.info['sfreq'])])
# time_series_corr_matrix = rsa_2_variability(time_series_corr_matrix)
else:
print('ERROR: wrong varibility_method name!' )
# save the result per event
iter_influence_step_result[iter_electrode] = time_series_corr_matrix # channel * time * trials * trials
# save the iter influence_step data to previously generated nan iter_event matrix
iter_event_result[iter_influence_step] = iter_influence_step_result
# save the iter event data to previously generated nan iter_power matrix
iter_power_result[iter_event] = iter_event_result
# load the iter power data to previously generated nan iter_subject matrix
iter_subject_result[iter_file] = iter_power_result
# load the iter subject data to previously generated nan all result matrix
all_result[iter_subject] = iter_subject_result
print('')
print("**************************************************************************")
print("******************** subject number:", iter_subject + 1, "/", subject_num,"finished *********************")
print("**************************************************************************")
print('')
all_rsa_result['data'] = all_result
np.save(result_loc +'/' + 'face_alpha_var_nor_result.npy', all_rsa_result)
# -
# # Data analysis part
# +
# import the variation of Exp2 and Exp3
result3_loc = 'E:/workspace/Trust_Data_and_Results/Haoming/varibility/result/varibilit_corr_20_exp3/'
all_var_result2 = np.load(result_loc +'/' + 'face_alpha_var_nor_result.npy', allow_pickle=True).item()
all_var_result3 = np.load(result3_loc +'/' + 'face_alpha_var_nor_result.npy', allow_pickle=True).item()
print(all_var_result2['data'].shape)
# -
# concatenate the data of Exp2 and Exp 3
all_var_result2_data = np.squeeze( all_var_result2['data'])
all_var_result3_data = np.squeeze(all_var_result3['data'])
print(all_var_result3_data.shape)
all_var_result23_data = np.concatenate((all_var_result2_data,all_var_result3_data),axis=2)
print(all_var_result23_data.shape)
# +
# Devide 63 electrode to 6 regions
region_ch_list={}
region_ch_list[0] = ['FT9','Fp1', 'Fpz', 'AF7','F7','FT7','AF3','F5','F3','F1','Fz','FC1','FC3','FC5']
region_ch_list[1] = ['FT10', 'Fpz', 'Fz', 'FC2', 'FC4', 'FC6', 'FT8', 'F2', 'F4', 'F6', 'F8', 'AF4', 'AF8', 'Fp2']
region_ch_list[2] = ['TP9','FT9','FC1','FC3','FC5', 'FT7', 'Cz', 'CPz', 'CP1', 'CP3', 'CP5', 'TP7', 'T7', 'C5', 'C3', 'C1']
region_ch_list[3] = ['TP10','FT10', 'FC2', 'FC4', 'FC6', 'FT8', 'Cz', 'CPz', 'CP2', 'CP4', 'CP6', 'TP8', 'T8', 'C6', 'C4', 'C2']
region_ch_list[4] = ['TP9','Oz', 'POz', 'Pz', 'CPz', 'CP1', 'CP3', 'CP5', 'TP7', 'P1', 'P3', 'P5', 'P7', 'PO3', 'PO7', 'O1']
region_ch_list[5] = ['TP10','Oz', 'POz', 'Pz', 'CPz', 'CP2', 'CP4', 'CP6', 'TP8', 'P2', 'P4', 'P6', 'P8', 'PO4', 'PO8', 'O2']
ch_names = all_var_result2['ch_names']
region_ch_location={}
for iter_region in range(6):
region_ch_location[iter_region]=[]
for iter_channel in range(len(region_ch_list[iter_region])):
iter_channel_position = np.where(np.array(ch_names) == region_ch_list[iter_region][iter_channel])[0][0]
region_ch_location[iter_region].append(iter_channel_position)
# Map the data of 63 eectrodes to 6 regions
region_all_var_result23_data = np.full((34,4,7,6,1200), np.nan)
for iter_region in range(6):
region_all_var_result23_data[:,:,:,iter_region,:] = np.mean(all_var_result23_data[:,:,:,region_ch_location[iter_region],:],axis=3)
print(region_all_var_result23_data.shape)
# +
# Calculate the 1_samp permutation test
n_permutations=10000
permutation_cluster_result = {}
# calculate std
permu_result_loc = 'E:/workspace/Trust_Data_and_Results/Haoming/varibility/result/varibilit_corr_20_exp2/perm_result/'
cond0_region_all_var_result23_data = region_all_var_result23_data[:,0,:,:,:]
print(region_all_var_result23_data.shape)
# tfr_ROI_epoch_data_std={}
# tfr_ROI_epoch_data_std['all_event_std'] = np.std(np.squeeze(all_data_nor['data'][:, :, 0, :, :]), axis=0)
# tfr_ROI_epoch_data_std['event1_std'] = np.std(np.squeeze(all_data_nor['data'][:, :, 1, :, :]), axis=0)
# tfr_ROI_epoch_data_std['event2_std'] = np.std(np.squeeze(all_data_nor['data'][:, :, 2, :, :]), axis=0)
# permutation_cluster_result['std_error'] = tfr_ROI_epoch_data_std
for iter_channel in range(6):
#ROI_num = 10
# compute the cluster test for event 1
iter_channel_result = {}
for iter_event in range(7):
T_obs, clusters, cluster_p_values, H0 = mne.stats.permutation_cluster_1samp_test(cond0_region_all_var_result23_data[good_subject, iter_event, iter_channel, :]-1, out_type='mask',n_permutations=n_permutations, tail=0, verbose=None)
iter_event_result = {'T_obs':T_obs, 'clusters':clusters, 'cluster_p_values':cluster_p_values, 'H0':H0}
iter_channel_result[iter_event] = iter_event_result
print('')
print("**************************************************************************")
print("********************* total number:", iter_channel*7+iter_event + 1, "/", 42,"finished ***********************")
print("**************************************************************************")
print('')
permutation_cluster_result[iter_channel] = iter_channel_result
if not os.path.exists(permu_result_loc):
os.makedirs(permu_result_loc)
np.save(permu_result_loc + 'exp23_permu_result' + '.npy', permutation_cluster_result)
# +
permu_result_loc = 'E:/workspace/Trust_Data_and_Results/Haoming/varibility/result/varibilit_corr_20_exp2/perm_result/'
permutation_cluster_result = np.load(permu_result_loc + 'exp23_permu_result' + '.npy', allow_pickle=True).item()
# -
# function of plot the permutation result
import matplotlib.pyplot as plt
import numpy as np
import os
def plot_erp(permutation_cluster_result, epoch_mean, epoch_data_std, times, event_name, line_color=['orangered','limegreen'],
figsize=(14,6), title_size=25, legend_size=15, labelsize=15, ticksize=20, subplots_adjust=[0.15, 0.15, 0.85, 0.85]):
# keys of the dict permutation_cluster_result
#'event1_result':event1_result, 'event2_result':event2_result, 'compare_result':compare_result}
#'T_obs':T_obs, 'clusters':clusters, 'cluster_p_values':cluster_p_values, 'H0':H0
# keys of the dict epoch_data_mean
# 'event_0' (34, 1300)
# plot for each ROI
event_0_line_color = line_color[0]
event_1_line_color = line_color[1]
event_2_line_color = line_color[2]
plt.close('all')
plt.rcParams['figure.figsize'] = figsize # 设置figure_size尺寸
event_num_plot = [0,2,5]
# (34, 6, 3, 63, 1300)
# epoch_mean={}
# epoch_mean[0] = np.squeeze(np.average(epoch_data['event_0'], axis=0))
# epoch_mean[1] = np.squeeze(np.average(epoch_data['event_1'], axis=0))
for iter_event_num in range(len(event_num_plot)):
iter_color = line_color[iter_event_num]
iter_event = event_num_plot[iter_event_num]
plt.plot(times, epoch_mean[iter_event], color=iter_color, linestyle='--', alpha=0.4)
plt.fill_between(times, epoch_mean[iter_event] - epoch_data_std[iter_event], epoch_mean[iter_event] + epoch_data_std[iter_event], color=iter_color, alpha=0.1)
# event 0 的显著性
for i_c, c in enumerate(permutation_cluster_result[iter_event]['clusters']):
c = c[0]
if permutation_cluster_result[iter_event]['cluster_p_values'][i_c] <= 0.05:
plt.plot(times[c.start : c.stop - 1], epoch_mean[iter_event][c.start : c.stop-1], color=iter_color, alpha=0.9)
#hf = plt.plot(times, T_obs, 'g')
#plt.legend((h, ), ('cluster p-value < 0.05', ))
plt.subplots_adjust(left=subplots_adjust[0], bottom=subplots_adjust[1], right=subplots_adjust[2], top=subplots_adjust[3], hspace=0.1,wspace=0.1)
plt.xlim([times[0]-0.02, times[-1]+0.02])
plt.yticks(size=ticksize, family='Arial')
plt.xticks(size=ticksize, family='Arial')
return plt
# +
# plot varibility erps and permuation result of the variation of Exp2 and Exp3
import matplotlib.pyplot as plt
from matplotlib import font_manager
plot_result_loc = result_loc + '/Exp2_condition0_plot_result/'
# permutation_cluster_result = np.load(permu_result_loc + 'all_event_0_1_alpha' + '.npy', allow_pickle=True).item()
figsize=(12,6)
title_size = 20
labelsize = 15
ticksize=25
fontProperties = font_manager.FontProperties(fname='C:/Windows/Fonts/arial.ttf')
times = np.arange(-0.2, 1, 0.001)
event_name = ['Exp 2 step 1', 'Exp 2 step 3', 'Exp 3 step 6']
chan_name = ['Region 1', 'Region 2', 'Region 3', 'Region 4', 'Region 5', 'Region 6']
# for iter_file in range(len(file_name)):
cond0_region_all_var_result23_data = region_all_var_result23_data[:,0,:,:,:]
if not os.path.exists(plot_result_loc):
os.makedirs(plot_result_loc)
for iter_chan in range(6):
iter_region_all_var_result23_data = cond0_region_all_var_result23_data[good_subject,:,iter_chan,:]
iter_region_all_var_result23_data_std = np.std(iter_region_all_var_result23_data, axis=0)
iter_region_all_var_result23_data_mean = np.average(iter_region_all_var_result23_data, axis=0)
print(iter_region_all_var_result23_data_mean.shape)
plt = plot_erp(permutation_cluster_result[iter_chan], iter_region_all_var_result23_data_mean,
iter_region_all_var_result23_data_std, times, event_name, ticksize = ticksize,
line_color=['#70AD47','#0070C0', '#DF4058'], figsize = figsize)
plt.title(chan_name[iter_chan], family='Arial', fontdict= {'fontsize':title_size})
plt.xlabel("time (s)", fontsize=labelsize, family='Arial')
plt.ylabel("varibility change", fontsize=labelsize, family='Arial')
plt.axvline(times[201], c="gray", ls = "dashed")
plt.plot(times, np.ones(len(times)), color="gray", linestyle="dashed")
plt.savefig(plot_result_loc + '/' + chan_name[iter_chan] + ".png")
# -
# calculate the temporal mean eeg variability 0-300ms
mean_region_all_var_result23_data = np.mean(region_all_var_result23_data[:,:,:,:,200:500],axis=-1)
mean_region_all_var_result23_data.shape
# step 1 and step 2 (pre-influence)
mean_region_all_var_result23_data_123 = np.mean(mean_region_all_var_result23_data[:,:,0:2],axis=2)
mean_region_all_var_result23_data_123.shape
all_corr_result = mean_region_all_var_result23_data_123
# Load the behavior data of Exp2 & 3
behaviorVar_loc = 'E:/workspace/Trust_Data_and_Results/Haoming/varibility/behavior_result/behavior_varibility.npy'
behavior_varibility = np.load(behaviorVar_loc)
behavior_varibility.shape
# +
# Plot:Use the 1-3 mean EEG variability to predict the Exp2-3 behavior change. # mean_all_rsa_result23_0_300
import seaborn as sns
import scipy
import scipy.stats
import matplotlib.pyplot as plt
figsize=(10,10)
title_size=25
legend_size=15
labelsize=23
ticksize=32
plot_loc = 'E:/workspace/Trust_Data_and_Results/Haoming/varibility/result/varibilit_corr_20_exp2/each_influence_varibility/var1_2_mean_predict_behavior_6region_0_300_plot/'
event_name = ['trust0', 'trust1', 'trust2', 'trust3']
all_corr_r, all_corr_p = [], []
for iter_event in range(len(event_name)):
# Generate the correlation image storage address of each condition (Experiment 2 and experiment 3)
iter_event_loc = plot_loc + event_name[iter_event] + '/'
if not os.path.exists(iter_event_loc):
os.makedirs(iter_event_loc)
for iter_channel in range(all_corr_result.shape[-1]):
# Extract behavioral data to X and eeg variability to Y
x = behavior_varibility[good_subject, iter_event]
y = all_corr_result[good_subject, iter_event, iter_channel]
# Calculate correlations and statistical significance, and store results
scipy_corr = scipy.stats.pearsonr(x, y)
# Plot the results for each regions
plt.close('all')
plt.rcParams['figure.figsize'] = figsize # set figure_size
# Significant plots are shown in red and non-significant plots in green
if scipy_corr[1] <= 0.05:
sns.regplot(x=x, y=y, color="r")
else:
sns.regplot(x=x, y=y, color="g")
small2large_position = np.argsort(x)
plt.scatter(x[small2large_position[0:15]],y[small2large_position[0:15]],marker = 'o', color='#ED7D31', s=400)
plt.scatter(x[small2large_position[15:]],y[small2large_position[15:]],marker = 'o', color='#0070C0', s=400)
plt.title('r = '+ str(round(scipy_corr[0], 3)) + ', p-value = '+ str(round(scipy_corr[1], 3)), fontsize=title_size)
plt.xlabel("hehavior varibility", fontsize=labelsize, family='Arial')
plt.ylabel("EXP2-EXP3 EEG variability change rage ", fontsize=labelsize, family='Arial')
plt.yticks(size=ticksize, family='Arial')
plt.xticks(size=ticksize, family='Arial')
plt.xlim([-0.53, 0.23]) #event0: -0.53, 0.23 #event1: -0.33, 0.38
iter_plot_loc = iter_event_loc + '/' + str(iter_channel+1) + '_corr.png'
plt.savefig(iter_plot_loc)
| code/6_exp2_3_variation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (myenv)
# language: python
# name: myenv
# ---
# +
from pprint import pprint
import sklearn
from sklearn import preprocessing
from sklearn.feature_extraction.text import TfidfVectorizer
#import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import numpy as np
from pprint import pprint
import matplotlib.pyplot as plt
#import pyLDAvis
#from sklearn.feature_extraction.text import CountVectorizer
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.metrics import confusion_matrix, precision_score, precision_recall_curve, recall_score, f1_score
from sklearn.naive_bayes import MultinomialNB
#from sklearn.metrics.pairwise import cosine_similarity
#from sklearn.base import BaseEstimator, TransformerMixin
#from sklearn.pipeline import FeatureUnion
#from sklearn.feature_extraction import DictVectorizer
#from sklearn.metrics import accuracy_score
#from sklearn.model_selection import GridSearchCV
#from sklearn.decomposition import LatentDirichletAllocation
#from imblearn.over_sampling import SMOTE
from os import path
from wordcloud import WordCloud, STOPWORDS
#from PIL import Image
from wordcloud import ImageColorGenerator
import re
#import pickle
#import joblib
import spacy
import nltk
from nltk.corpus import stopwords
from nltk.stem import WordNetLemmatizer
import string
from nltk import word_tokenize,sent_tokenize
from nltk.stem.porter import PorterStemmer
#from textblob import TextBlob
#from vaderSentiment.vaderSentiment import SentimentIntensityAnalyzer
import datetime
import warnings
# +
raw_df= pd.read_csv(r'G:/Symbi/Dataset/TelecomPOC New.csv',encoding='utf-8', parse_dates=[1])
#raw_df= pd.read_csv(r'C:/Users/<NAME>/Vi_tweets_test.csv',encoding='utf-8')
raw_df.head()
#neg = raw_df[raw_df.Sentiment1=='Negative']
#neg = neg.reset_index(drop=True)
text = raw_df.Tweet.dropna()
print(text)
#raw_df= raw_df[pd.notnull(raw_df['Tweet'])] #Delete rows where Tweet is null.
# -
def clean_tweet(RawSentence):
"""
removing all hashtags , punctuations, stop_words and links, also stemming words
"""
txt = RawSentence.lower()
word_tokens = word_tokenize(txt)
#word_tokens = word_tokenize(RawSentence)
lemma = WordNetLemmatizer()
nlp = spacy.load('en_core_web_sm')
stop_words = set(stopwords.words('english'))
stop_words.update(["amp", "rt", "cc"]) # adding twitter specific stop words
#stop_words = stop_words - set(['no', 'not','nil']) #removing No, not, nil from stop words
stop_words.update(["worst","airtelpresence","sunday","monday","tuesday","wednesday","thursday","friday","saturday",
"weekend","apparently","extremely","worse","consolation","shouldnt","wont","arent","maybe"
"better", "other","instead","look","yesterday","unacceptable","miss","couldnt","depress","let",
"thank","actually","frustrate","ridiculous","expect","pathetic","terrible","hathway",
"impossible","ask","earlier","literally","years","surprise","horrible","airtel","current","face",
"upset","pls","one","two","three","four","empty","please","good","finally","proper","development",
"situation","company","believe","question","anymore","consolation","mess","shock","longer","twitter",
"completely","finally", "waste","shock","annoy","strand",
"important","break","reason", "first","second","third","attempt","officially","absolutely","hello","yet","know","jio",
"january","february","march","april","may","june","july","august","september","october","november","december","without",
"vodafone","get","vodafoneidea","vicustomercare","vowifi","since","voda","reliancejio","jiocare","time","number"])
txt = re.sub(r"(@\S+)", "", txt) # remove @
txt = re.sub(r'\W', ' ', str(txt)) # remove all special characters including apastrophie
txt = txt.translate(str.maketrans(' ', ' ', string.punctuation)) # remove punctuations
txt = re.sub(r'\s+[a-zA-Z]\s+', ' ', txt) # remove all single characters from within the text (it's -> it s then we need to remove s)
txt = re.sub(r'\^[a-zA-Z]\s+', ' ', txt) # remove all single characters from beginning of the text
txt = re.sub(r'\s+', ' ', txt, flags=re.I) # Substituting multiple spaces with single space
txt = re.sub(r"(http\S+|http)", "", txt) # remove links
#txt = re.sub(r'^b\s+', '', txt) # generally used when text is scrapped online. 'b' is prefixed often indicating binary text. This is not reqd for us
txt = [re.sub('\s+', ' ', t) for t in txt] # Remove new line characters
txt = ''.join([i for i in txt if not i.isdigit()]).strip() # remove digits ()
txt = ' '.join([i for i in txt.split(" ") if i not in stop_words]) # split() is native tokenizer in Python
#txt = ' '.join(lemma.lemmatize(word) for word in txt.split()) # Used lemmatizer later
#txt = ' '.join([PorterStemmer().stem(word=word) for word in txt.split(" ") if word not in stop_words ]) # stem & remove stop words
#soup = BeautifulSoup(txt) #to scrap information from web pages....not needed here
#txt = soup.get_text
def deEmojify(inputString):
return inputString.encode('ascii', 'ignore').decode('ascii')
def stem_tokens(tokens, lemmatize):
lemmatized = []
for item in tokens:
lemmatized.append(lemma.lemmatize(item,'v'))
return lemmatized
txt = deEmojify(txt)
tokens =[]
excluded_tags = {"ADJ", "ADV", "ADP", "PROPN","CCONJ","AUX","DET","PRON","VERB"}
for token in nlp(txt):
if token.pos_ not in excluded_tags:
tokens.append(token.text)
lemm = stem_tokens(tokens, lemma)
#tokens = nltk.word_tokenize(txt)
tokens = [ch for ch in lemm if len(ch)>2] #remove words with character length below 3
#tokens = [ch for ch in tokens if len(ch)<=15] #remove words with character length above 15
joined_text = ' '.join(tokens)
return joined_text
raw_df['clean_tweet'] = raw_df.Tweet.apply(clean_tweet)
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
X=raw_df['clean_tweet']
X_train,X_test =train_test_split(X,test_size=0.3)
#print(X_train)
# calculate number of tokens for each tweet
raw_df['tokens'] = raw_df['clean_tweet'].str.split().str.len()# plot average word count by user
raw_df.groupby('Vertex2')['tokens'].mean().plot(kind='barh',figsize=(10,5))
plt.xlabel('Average Words')
plt.ylabel('Vertex')
plt.show()
# !git clone https://github.com/rwalk/gsdmm.git gsdmm
# !pip install gsdmm
python3 -m pip install google-cloud-dataproc
import gsdmm
from gsdmm import MovieGroupProcess
from tqdm import tqdm# convert string of tokens into tokens list
raw_df['tokens'] = X.apply(lambda x: re.split('\s', x))# create list of token lists
docs = raw_df['tokens'].tolist()
# +
# vectorize the corpus. the vectorizer object will convert text to vector form on the basis of frequency (count) of each word in text
#Vectorization is a process of converting the text data into a machine-readable form. Each word forms the index.
#min_df - will ignore words that are present in min_df percent or less no of documents
#max_df - will take words that are present in max df percent of all the documents
#vectorizer = CountVectorizer(max_df=0.9, min_df=10, ngram_range=(1,2), token_pattern='\w+|\$[\d\.]+|\S+')
#tfidf_vectorizer = TfidfVectorizer(max_df=0.9, min_df=20, ngram_range=(1,2), token_pattern='\w+|\$[\d\.]+|\S+')
#vectorizer = CountVectorizer(max_df=.95, min_df=0.05, ngram_range=(1,2),token_pattern='\w+|\$[\d\.]+|\S+')
#vectorizer = CountVectorizer(max_df=0.9, min_df=0.2, ngram_range=(1,2))
tfidf_vectorizer = TfidfVectorizer(max_df=0.9, min_df=10, ngram_range=(1,2))
# -
#vectors = tfidf_vectorizer.fit_transform(raw_df['clean_tweet']).toarray()
vectors = tfidf_vectorizer.fit_transform(X_train)
vectors.shape
#vectors.nnz /float(vectors.shape[0])
features = tfidf_vectorizer.get_feature_names()
feature_order = np.argsort(tfidf_vectorizer.idf_)[::-1]
top_n = 10
top_n_features = [features[i] for i in feature_order[:top_n]]
print(top_n_features)
Test_Vectors=tfidf_vectorizer.transform(X_test).toarray()
Test_Vectors.shape
from sklearn import metrics
clf = MultinomialNB(alpha=.01)
clf.fit(vectors, X_train )
pred = clf.predict(Test_Vectors)
metrics.f1_score(X_test, pred, average='macro')
| gsdmm/STTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ResNet Model Latency Testing
#
# Testing ResNet model with the default Seldon Tensor and Tensorflow Tensor.
#
# <img src="dog.jpeg"/>
# !cp ../../../proto/prediction.proto ./proto
# !cp -vr ../../../proto/tensorflow/tensorflow .
# !python -m grpc.tools.protoc -I./ --python_out=./ --grpc_python_out=./ ./proto/prediction.proto
# # Download model
#
# !mkdir -p model
# !wget -O model/saved_model.pb https://storage.googleapis.com/inference-eu/models_zoo/resnet_V1_50/saved_model/saved_model.pb
# # Wrap inference
# !s2i build -E environment_grpc . seldonio/seldon-core-s2i-python36:0.13 seldon-resnet2.4
# !docker run --name "resnet" -d --rm -p 5000:5000 -v ${PWD}/model:/model seldon-resnet2.4
# # Test
import json
import requests
import base64
from proto import prediction_pb2
from proto import prediction_pb2_grpc
import grpc
import numpy as np
import pickle
import tensorflow as tf
import cv2
import datetime
import tensorflow as tf
# +
def image_2_vector(input_file):
nparr = np.fromfile(input_file, dtype=np.float32)
print("nparr",nparr.dtype,nparr.shape)
img = cv2.imdecode(nparr, cv2.IMREAD_ANYCOLOR)
print("img",img.dtype,img.shape)
print("Initial size",img.shape)
image = cv2.resize(img, (w, h))
print("image",image.dtype)
print("Converted size",image.shape)
vector = image.reshape((w * h * 3))
print("vector shape",vector.shape, "vector type", vector.dtype )
return vector
def image_2_bytes(input_file):
with open(input_file, "rb") as binary_file:
# Read the whole file at once
data = binary_file.read()
#data = data.tobytes()
#print(data)
print("binary data size:", len(data), type(data))
return data
def run(function,image_path,iterations=1):
w = 224
h = 224
# NOTE(gRPC Python Team): .close() is possible on a channel and should be
# used in circumstances in which the with statement does not fit the needs
# of the code.
with grpc.insecure_channel('localhost:5000') as channel:
stub = prediction_pb2_grpc.ModelStub(channel)
print("seldon stub", stub)
start_time = datetime.datetime.now()
processing_times = np.zeros((0),int)
img = cv2.imread(image_path)
print("img type", type(img))
print("img",img.shape)
print("Initial size",img.shape)
image = cv2.resize(img, (w, h))
image = image.reshape(1, w, h, 3)
print("image",image.dtype)
print("Converted size",image.shape)
if function == "tensor":
datadef = prediction_pb2.DefaultData(
names = 'x',
tensor = prediction_pb2.Tensor(
shape = image.shape,
values = image.ravel().tolist()
)
)
elif function == "tftensor":
print("Create tftensor")
datadef = prediction_pb2.DefaultData(
names = 'x',
tftensor = tf.make_tensor_proto(image)
)
GRPC_request = prediction_pb2.SeldonMessage(
data = datadef
)
for I in range(iterations):
start_time = datetime.datetime.now()
response = stub.Predict(request=GRPC_request)
end_time = datetime.datetime.now()
duration = (end_time - start_time).total_seconds() * 1000
processing_times = np.append(processing_times,np.array([int(duration)]))
print('processing time for all iterations')
for x in processing_times:
print(x,"ms")
print('processing_statistics')
print('average time:',round(np.average(processing_times),1), 'ms; average speed:', round(1000/np.average(processing_times),1),'fps')
print('median time:',round(np.median(processing_times),1), 'ms; median speed:',round(1000/np.median(processing_times),1),'fps')
print('max time:',round(np.max(processing_times),1), 'ms; max speed:',round(1000/np.max(processing_times),1),'fps')
print('min time:',round(np.min(processing_times),1),'ms; min speed:',round(1000/np.min(processing_times),1),'fps')
print('time percentile 90:',round(np.percentile(processing_times,90),1),'ms; speed percentile 90:',round(1000/np.percentile(processing_times,90),1),'fps')
print('time percentile 50:',round(np.percentile(processing_times,50),1),'ms; speed percentile 50:',round(1000/np.percentile(processing_times,50),1),'fps')
print('time standard deviation:',round(np.std(processing_times)))
print('time variance:',round(np.var(processing_times)))
# -
run("tensor","./dog.jpeg",iterations=100)
run("tftensor","./dog.jpeg",iterations=100)
# The stats illustrate that the tftensor payload which is the only difference improves on the latency performance.
# !docker rm -f resnet
| examples/models/resnet/reset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Okfr_uhwhS1X" colab_type="text"
# # Lambda School Data Science - A First Look at Data
#
#
# + [markdown] id="9dtJETFRhnOG" colab_type="text"
# ## Lecture - let's explore Python DS libraries and examples!
#
# The Python Data Science ecosystem is huge. You've seen some of the big pieces - pandas, scikit-learn, matplotlib. What parts do you want to see more of?
# + id="WiBkgmPJhmhE" colab_type="code" colab={}
# TODO - we'll be doing this live, taking requests
# and reproducing what it is to look up and learn things
myList = [187, 69, 420, 117]
# + [markdown] id="pepX6lsqSiNY" colab_type="text"
# # this is new text
# + [markdown] id="lOqaPds9huME" colab_type="text"
# ## Assignment - now it's your turn
#
# Pick at least one Python DS library, and using documentation/examples reproduce in this notebook something cool. It's OK if you don't fully understand it or get it 100% working, but do put in effort and look things up.
# + id="TGUS79cOhPWj" colab_type="code" colab={}
# The essentials, the iconic Trio, the triumvirate
import pandas as pd
import numpy as numpy
import matplotlib.pyplot as plt
# + id="AwbTaWrRnlaE" colab_type="code" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "Ly8gQ29weXJpZ<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 71} outputId="bf71ad1d-43ce-4231-b42a-60adec4d8683"
# getting dataset from kaggle and converting to usable csv file
from google.colab import files # shoutout patricia for saving my life here
uploaded = files.upload()
# be weary of running this cell, took like 10 minutes to load the data
# + id="ByUyBOIdnMgU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 739} outputId="b831ee50-0c48-4696-9ea9-602a34136992"
# importing the data set now that its in usable csv form
df = pd.read_csv('renfe.csv')
# taking a glance at the data set
df.head(10)
# + id="SVLvBF3mpKJq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="cce814e2-2785-46c2-b858-4f4d035a1b8a"
# checking out the shape to see what we're working with
df.shape
# + [markdown] id="sWa_eLvnzdys" colab_type="text"
# oh wow, thats quite a lot of data lol
# + id="yHxqxtmKpX5z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 294} outputId="442886fa-41de-4bf2-f077-472e620c0d1e"
# checking out the various stats on quantifiable data
df.describe()
# + id="Hlnr2ackqAPF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="4df35020-c4d5-4ae6-e127-ac5eb8f6e33e"
# scatter plot of some features I chose that seemed relevant
plt.scatter(df['train_type'], df['price'])
plt.xlabel('train type')
plt.ylabel('price')
plt.title('Some Data From Spanish High Speed railway ')
plt.show()
# + id="ceukZNLZun6b" colab_type="code" colab={}
# going to try some stuff I saw on the example where i found the data
# will need seaborn
import seaborn as sns
# + id="fNWDUd34usnL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 459} outputId="8da55df0-37ef-401f-ee63-cbf39efaebe4"
# saw this on a example I found, these came out quite ugly.
# i will continue to try other methodologies to create something where
# meaningful conclusions can be drawn
sns.pairplot(df);
# + id="p5DyB-E6xYS4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 413} outputId="40f8401e-3bbb-4f24-e293-ddc1ab2b2c50"
plt.figure(figsize= (10,6))
sns.countplot(df.train_class);
# this comes from example as well, came out much prettier than previous cell
# and this actually provides an easy to visualize chart that
# one can easily draw conclusions from
# the conclusion here obviously being:
# On this Spanish railway, Turista is clearly the most used one
# + id="IfX3CiQeyTvs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 413} outputId="db595546-4fe5-4400-fa37-75f43079d6d2"
# same as above, but using a different feature, this time its "Train Type"
plt.figure(figsize= (15,6))
sns.countplot(df.train_type);
# this again, providing a very easy to see conlusion to draw
# on this Spanish railway, "Ave" is the most frequently riden Train Type
# + id="PdOx4gw_2aSK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 269} outputId="3e17d469-177b-44e4-dd21-88af5735b83a"
df['price'].hist();
# one last bit of charting here, we can visualize the distribution of
# price that passengers paid aboard this Spanish railway
# + [markdown] id="BT9gdS7viJZa" colab_type="text"
# ### Assignment questions
#
# After you've worked on some code, answer the following questions in this text block:
#
# 1. Describe in a paragraph of text what you did and why, as if you were writing an email to somebody interested but nontechnical.
#
# 2. What was the most challenging part of what you did?
#
# 3. What was the most interesting thing you learned?
#
# 4. What area would you like to explore with more time?
#
#
# Answers:
# 1. Initially, I had to find a way to get a set of data that wasn't neccessarily usable to start with, and convert it into a format that I could work with using the tools we have here on Python(was quite a headache for a while lol). From there, I just did some of our basic analysis, primarily examining the dimensions of the data, as well as the core statisitcs such as mean, median, standard deviation etc. Then lastly, I charted what I thought to be relevant features from the data that one could use to intuitively draw conclusions.
#
# 2. The most challenging part was taking a data set that wasn't in a nice csv file to begin with, and then getting into a nice easy to work with csv file. In reality, a pretty simple problem to solve, just took a while to find the actual notation to do so.
#
# 3. The most interesting thing I learned would easily have to be the ability to now grab any set of data I find off the internet, save it as a csv file to my computer, and from there be able to import it on to Google colabs and work with the data using Python.
#
# 4. From here, using more of the tools that were touched on in the precourse such as the tools in seaborn and sklearn, and being able to draw meaningful and relevant conclusions about the data I'm working with.
#
#
# + [markdown] id="_XXg2crAipwP" colab_type="text"
# ## Stretch goals and resources
#
# Following are *optional* things for you to take a look at. Focus on the above assignment first, and make sure to commit and push your changes to GitHub (and since this is the first assignment of the sprint, open a PR as well).
#
# - [pandas documentation](https://pandas.pydata.org/pandas-docs/stable/)
# - [scikit-learn documentation](http://scikit-learn.org/stable/documentation.html)
# - [matplotlib documentation](https://matplotlib.org/contents.html)
# - [Awesome Data Science](https://github.com/bulutyazilim/awesome-datascience) - a list of many types of DS resources
#
# Stretch goals:
#
# - Find and read blogs, walkthroughs, and other examples of people working through cool things with data science - and share with your classmates!
# - Write a blog post (Medium is a popular place to publish) introducing yourself as somebody learning data science, and talking about what you've learned already and what you're excited to learn more about.
| module1-afirstlookatdata/LS_DS_111_A_First_Look_at_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import glob
glob.glob("*")
import tokenization
# +
import h5py
filename = 'news_sample.hdf5'
with h5py.File(filename, 'r') as f:
# List all groups
print("Keys: %s" % f.keys())
a_group_key = list(f.keys())[0]
# f.visit(printname)
print(list(f[a_group_key]['10']))
# Get the data
data = list(f[a_group_key])
# -
def printname(name):
print(name)
data
| data/.ipynb_checkpoints/Untitled-checkpoint.ipynb |