code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# look at tools/set_up_magics.ipynb
yandex_metrica_allowed = True ; get_ipython().run_cell('# one_liner_str\n\nget_ipython().run_cell_magic(\'javascript\', \'\', \n \'// setup cpp code highlighting\\n\'\n \'IPython.CodeCell.options_default.highlight_modes["text/x-c++src"] = {\\\'reg\\\':[/^%%cpp/]} ;\'\n \'IPython.CodeCell.options_default.highlight_modes["text/x-cmake"] = {\\\'reg\\\':[/^%%cmake/]} ;\'\n)\n\n# creating magics\nfrom IPython.core.magic import register_cell_magic, register_line_magic\nfrom IPython.display import display, Markdown, HTML\nimport argparse\nfrom subprocess import Popen, PIPE, STDOUT, check_output\nimport html\nimport random\nimport sys\nimport os\nimport re\nimport signal\nimport shutil\nimport shlex\nimport glob\nimport time\n\n@register_cell_magic\ndef save_file(args_str, cell, line_comment_start="#"):\n parser = argparse.ArgumentParser()\n parser.add_argument("fname")\n parser.add_argument("--ejudge-style", action="store_true")\n parser.add_argument("--under-spoiler-threshold", type=int, default=None)\n args = parser.parse_args(args_str.split())\n \n cell = cell if cell[-1] == \'\\n\' or args.no_eof_newline else cell + "\\n"\n cmds = []\n with open(args.fname, "w") as f:\n f.write(line_comment_start + " %%cpp " + args_str + "\\n")\n for line in cell.split("\\n"):\n line_to_write = (line if not args.ejudge_style else line.rstrip()) + "\\n"\n if line.startswith("%"):\n run_prefix = "%run "\n if line.startswith(run_prefix):\n cmds.append(line[len(run_prefix):].strip())\n f.write(line_comment_start + " " + line_to_write)\n continue\n comment_prefix = "%" + line_comment_start\n if line.startswith(comment_prefix):\n cmds.append(\'#\' + line[len(comment_prefix):].strip())\n f.write(line_comment_start + " " + line_to_write)\n continue\n raise Exception("Unknown %%save_file subcommand: \'%s\'" % line)\n else:\n f.write(line_to_write)\n f.write("" if not args.ejudge_style else line_comment_start + r" line without \\n")\n for cmd in cmds:\n if cmd.startswith(\'#\'):\n display(Markdown("\\#\\#\\#\\# `%s`" % cmd[1:]))\n else:\n display(Markdown("Run: `%s`" % cmd))\n if args.under_spoiler_threshold:\n out = check_output(cmd, stderr=STDOUT, shell=True, universal_newlines=True)\n out = out[:-1] if out.endswith(\'\\n\') else out\n out = html.escape(out)\n if len(out.split(\'\\n\')) > args.under_spoiler_threshold:\n out = "<details> <summary> output </summary> <pre><code>%s</code></pre></details>" % out\n elif out:\n out = "<pre><code>%s</code></pre>" % out\n if out:\n display(HTML(out))\n else:\n get_ipython().system(cmd)\n\n@register_cell_magic\ndef cpp(fname, cell):\n save_file(fname, cell, "//")\n \n@register_cell_magic\ndef cmake(fname, cell):\n save_file(fname, cell, "#")\n\n@register_cell_magic\ndef asm(fname, cell):\n save_file(fname, cell, "//")\n \n@register_cell_magic\ndef makefile(fname, cell):\n fname = fname or "makefile"\n assert fname.endswith("makefile")\n save_file(fname, cell.replace(" " * 4, "\\t"))\n \n@register_line_magic\ndef p(line):\n line = line.strip() \n if line[0] == \'#\':\n display(Markdown(line[1:].strip()))\n else:\n try:\n expr, comment = line.split(" #")\n display(Markdown("`{} = {}` # {}".format(expr.strip(), eval(expr), comment.strip())))\n except:\n display(Markdown("{} = {}".format(line, eval(line))))\n \n \ndef show_log_file(file, return_html_string=False):\n obj = file.replace(\'.\', \'_\').replace(\'/\', \'_\') + "_obj"\n html_string = \'\'\'\n <!--MD_BEGIN_FILTER-->\n <script type=text/javascript>\n var entrance___OBJ__ = 0;\n var errors___OBJ__ = 0;\n function halt__OBJ__(elem, color)\n {\n elem.setAttribute("style", "font-size: 14px; background: " + color + "; padding: 10px; border: 3px; border-radius: 5px; color: white; "); \n }\n function refresh__OBJ__()\n {\n entrance___OBJ__ -= 1;\n if (entrance___OBJ__ < 0) {\n entrance___OBJ__ = 0;\n }\n var elem = document.getElementById("__OBJ__");\n if (elem) {\n var xmlhttp=new XMLHttpRequest();\n xmlhttp.onreadystatechange=function()\n {\n var elem = document.getElementById("__OBJ__");\n console.log(!!elem, xmlhttp.readyState, xmlhttp.status, entrance___OBJ__);\n if (elem && xmlhttp.readyState==4) {\n if (xmlhttp.status==200)\n {\n errors___OBJ__ = 0;\n if (!entrance___OBJ__) {\n if (elem.innerHTML != xmlhttp.responseText) {\n elem.innerHTML = xmlhttp.responseText;\n }\n if (elem.innerHTML.includes("Process finished.")) {\n halt__OBJ__(elem, "#333333");\n } else {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n }\n }\n return xmlhttp.responseText;\n } else {\n errors___OBJ__ += 1;\n if (!entrance___OBJ__) {\n if (errors___OBJ__ < 6) {\n entrance___OBJ__ += 1;\n console.log("req");\n window.setTimeout("refresh__OBJ__()", 300); \n } else {\n halt__OBJ__(elem, "#994444");\n }\n }\n }\n }\n }\n xmlhttp.open("GET", "__FILE__", true);\n xmlhttp.setRequestHeader("Cache-Control", "no-cache");\n xmlhttp.send(); \n }\n }\n \n if (!entrance___OBJ__) {\n entrance___OBJ__ += 1;\n refresh__OBJ__(); \n }\n </script>\n\n <p id="__OBJ__" style="font-size: 14px; background: #000000; padding: 10px; border: 3px; border-radius: 5px; color: white; ">\n </p>\n \n </font>\n <!--MD_END_FILTER-->\n <!--MD_FROM_FILE __FILE__.md -->\n \'\'\'.replace("__OBJ__", obj).replace("__FILE__", file)\n if return_html_string:\n return html_string\n display(HTML(html_string))\n\n \nclass TInteractiveLauncher:\n tmp_path = "./interactive_launcher_tmp"\n def __init__(self, cmd):\n try:\n os.mkdir(TInteractiveLauncher.tmp_path)\n except:\n pass\n name = str(random.randint(0, 1e18))\n self.inq_path = os.path.join(TInteractiveLauncher.tmp_path, name + ".inq")\n self.log_path = os.path.join(TInteractiveLauncher.tmp_path, name + ".log")\n \n os.mkfifo(self.inq_path)\n open(self.log_path, \'w\').close()\n open(self.log_path + ".md", \'w\').close()\n\n self.pid = os.fork()\n if self.pid == -1:\n print("Error")\n if self.pid == 0:\n exe_cands = glob.glob("../tools/launcher.py") + glob.glob("../../tools/launcher.py")\n assert(len(exe_cands) == 1)\n assert(os.execvp("python3", ["python3", exe_cands[0], "-l", self.log_path, "-i", self.inq_path, "-c", cmd]) == 0)\n self.inq_f = open(self.inq_path, "w")\n interactive_launcher_opened_set.add(self.pid)\n show_log_file(self.log_path)\n\n def write(self, s):\n s = s.encode()\n assert len(s) == os.write(self.inq_f.fileno(), s)\n \n def get_pid(self):\n n = 100\n for i in range(n):\n try:\n return int(re.findall(r"PID = (\\d+)", open(self.log_path).readline())[0])\n except:\n if i + 1 == n:\n raise\n time.sleep(0.1)\n \n def input_queue_path(self):\n return self.inq_path\n \n def wait_stop(self, timeout):\n for i in range(int(timeout * 10)):\n wpid, status = os.waitpid(self.pid, os.WNOHANG)\n if wpid != 0:\n return True\n time.sleep(0.1)\n return False\n \n def close(self, timeout=3):\n self.inq_f.close()\n if not self.wait_stop(timeout):\n os.kill(self.get_pid(), signal.SIGKILL)\n os.waitpid(self.pid, 0)\n os.remove(self.inq_path)\n # os.remove(self.log_path)\n self.inq_path = None\n self.log_path = None \n interactive_launcher_opened_set.remove(self.pid)\n self.pid = None\n \n @staticmethod\n def terminate_all():\n if "interactive_launcher_opened_set" not in globals():\n globals()["interactive_launcher_opened_set"] = set()\n global interactive_launcher_opened_set\n for pid in interactive_launcher_opened_set:\n print("Terminate pid=" + str(pid), file=sys.stderr)\n os.kill(pid, signal.SIGKILL)\n os.waitpid(pid, 0)\n interactive_launcher_opened_set = set()\n if os.path.exists(TInteractiveLauncher.tmp_path):\n shutil.rmtree(TInteractiveLauncher.tmp_path)\n \nTInteractiveLauncher.terminate_all()\n \nyandex_metrica_allowed = bool(globals().get("yandex_metrica_allowed", False))\nif yandex_metrica_allowed:\n display(HTML(\'\'\'<!-- YANDEX_METRICA_BEGIN -->\n <script type="text/javascript" >\n (function(m,e,t,r,i,k,a){m[i]=m[i]||function(){(m[i].a=m[i].a||[]).push(arguments)};\n m[i].l=1*new Date();k=e.createElement(t),a=e.getElementsByTagName(t)[0],k.async=1,k.src=r,a.parentNode.insertBefore(k,a)})\n (window, document, "script", "https://mc.yandex.ru/metrika/tag.js", "ym");\n\n ym(59260609, "init", {\n clickmap:true,\n trackLinks:true,\n accurateTrackBounce:true\n });\n </script>\n <noscript><div><img src="https://mc.yandex.ru/watch/59260609" style="position:absolute; left:-9999px;" alt="" /></div></noscript>\n <!-- YANDEX_METRICA_END -->\'\'\'))\n\ndef make_oneliner():\n html_text = \'("В этот ноутбук встроен код Яндекс Метрики для сбора статистики использований. Если вы не хотите, чтобы по вам собиралась статистика, исправьте: yandex_metrica_allowed = False" if yandex_metrica_allowed else "")\'\n html_text += \' + "<""!-- MAGICS_SETUP_PRINTING_END -->"\'\n return \'\'.join([\n \'# look at tools/set_up_magics.ipynb\\n\',\n \'yandex_metrica_allowed = True ; get_ipython().run_cell(%s);\' % repr(one_liner_str),\n \'display(HTML(%s))\' % html_text,\n \' #\'\'MAGICS_SETUP_END\'\n ])\n \n\n');display(HTML(("В этот ноутбук встроен код Яндекс Метрики для сбора статистики использований. Если вы не хотите, чтобы по вам собиралась статистика, исправьте: yandex_metrica_allowed = False" if yandex_metrica_allowed else "") + "<""!-- MAGICS_SETUP_PRINTING_END -->")) #MAGICS_SETUP_END
# # Аттрибуты файлов и файловых дескрипторов
#
# <p><a href="https://www.youtube.com/watch?v=bMmE7PPA1LQ&list=PLjzMm8llUm4AmU6i_hPU0NobgA4VsBowc&index=12" target="_blank">
# <h3>Видеозапись семинара</h3>
# </a></p>
#
#
# [Р<NAME>](https://github.com/victor-yacovlev/mipt-diht-caos/tree/master/practice/stat_fcntl)
#
#
# Сегодня в программе:
# * <a href="#stat" style="color:#856024"> Атрибуты файлов и разные способы их получения </a>
# * <a href="#time" style="color:#856024"> Извлечем время доступа из атрибутов файла </a>
# * <a href="#username" style="color:#856024"> Определим логин пользователя, изменившего файл </a>
# * <a href="#access" style="color:#856024"> Проверим свои права на файл </a>
# * <a href="#link" style="color:#856024"> Ссылки жесткие и символические </a>
# * <a href="#fds" style="color:#856024"> Атрибуты файловых дескрипторов </a>
# * <a href="#hw" style="color:#856024"> Комментарии к ДЗ </a>
#
# ## <a name="stat"></a> Атрибуты файлов и разные способы их получения
#
# Сигнатуры функций, с помощью которых можно получить аттрибуты
#
# ```c
# #include <sys/types.h>
# #include <sys/stat.h>
# #include <unistd.h>
#
# int stat(const char *pathname, struct stat *buf);
# int fstat(int fd, struct stat *buf);
# int lstat(const char *pathname, struct stat *buf);
#
# #include <fcntl.h> /* Definition of AT_* constants */
# #include <sys/stat.h>
#
# int fstatat(int dirfd, const char *pathname, struct stat *buf,
# int flags);
# ```
#
#
# Описание структуры из `man 2 stat`:
#
# ```c
# struct stat {
# dev_t st_dev; /* ID of device containing file */
# ino_t st_ino; /* inode number */
# mode_t st_mode; /* protection */
# nlink_t st_nlink; /* number of hard links */
# uid_t st_uid; /* user ID of owner */
# gid_t st_gid; /* group ID of owner */
# dev_t st_rdev; /* device ID (if special file) */
# off_t st_size; /* total size, in bytes */
# blksize_t st_blksize; /* blocksize for filesystem I/O */
# blkcnt_t st_blocks; /* number of 512B blocks allocated */
#
# /* Since Linux 2.6, the kernel supports nanosecond
# precision for the following timestamp fields.
# For the details before Linux 2.6, see NOTES. */
#
# struct timespec st_atim; /* time of last access */
# struct timespec st_mtim; /* time of last modification */
# struct timespec st_ctim; /* time of last status change */
#
# #define st_atime st_atim.tv_sec /* Backward compatibility */
# #define st_mtime st_mtim.tv_sec
# #define st_ctime st_ctim.tv_sec
# };
# ```
#
# Особый интерес будет представлять поле `.st_mode`
#
# Биты соответствующие маскам:
# * `0170000` - тип файла.
#
# Эти биты стоит рассматривать как одно число, по значению которого можно определить тип файла. Сравнивая это число с:
# * `S_IFSOCK 0140000 socket`
# * `S_IFLNK 0120000 symbolic link`
# * `S_IFREG 0100000 regular file`
# * `S_IFBLK 0060000 block device`
# * `S_IFDIR 0040000 directory`
# * `S_IFCHR 0020000 character device`
# * `S_IFIFO 0010000 FIFO`
# * `0777` - права на файл.
#
# Эти биты можно рассматривать как независимые биты, каджый из которых отвечает за право (пользователя, его группы, всех остальных) (читать/писать/выполнять) файл.
# **fstat** - смотрит по файловому дескриптору
# +
# %%cpp stat.c
# %run gcc stat.c -o stat.exe
# %run rm -rf tmp && mkdir tmp && touch tmp/a && ln -s ./a tmp/a_link && mkdir tmp/dir
# %run ./stat.exe < tmp/a
# %run ./stat.exe < tmp/dir
# %run ./stat.exe < tmp/a_link
#include <sys/types.h>
#include <sys/stat.h>
#include <fcntl.h>
#include <unistd.h>
#include <stdio.h>
#include <assert.h>
int main(int argc, char *argv[])
{
struct stat s;
fstat(0, &s); // get stat for stdin
printf("is regular: %s ", ((s.st_mode & S_IFMT) == S_IFREG) ? "yes" : "no "); // can use predefined mask
printf("is directory: %s ", S_ISDIR(s.st_mode) ? "yes" : "no "); // or predefined macro
printf("is symbolic link: %s\n", S_ISLNK(s.st_mode) ? "yes" : "no ");
return 0;
}
# -
# Обратите внимание, что для симлинки результат показан как для регулярного файла, на который она ссылается.
# Тут, вероятно, замешан bash, который открывает файл на месте stdin для нашей программы. Видно, что он проходит по симлинкам.
#
# **stat** - смотри по имени файла, следует по симлинкам
# +
# %%cpp stat.c
# %run gcc stat.c -o stat.exe
# %run rm -rf tmp && mkdir tmp && touch tmp/a && ln -s ./a tmp/a_link && mkdir tmp/dir
# %run ./stat.exe tmp/a
# %run ./stat.exe tmp/dir
# %run ./stat.exe tmp/a_link
#include <sys/types.h>
#include <sys/stat.h>
#include <fcntl.h>
#include <unistd.h>
#include <stdio.h>
#include <assert.h>
int main(int argc, char *argv[])
{
assert(argc == 2);
struct stat s;
stat(argv[1], &s); // Следует по симлинкам
printf("is regular: %s ", ((s.st_mode & S_IFMT) == S_IFREG) ? "yes" : "no "); // can use predefined mask
printf("is directory: %s ", S_ISDIR(s.st_mode) ? "yes" : "no "); // or predefined macro
printf("is symbolic link: %s\n", S_ISLNK(s.st_mode) ? "yes" : "no ");
return 0;
}
# -
# Обратите внимание, что для симлинки результат показан как для регулярного файла, на который она ссылается.
#
# **lstat** - смотрит по имени файла, не следует по симлинкам.
# +
# %%cpp stat.c
# %run gcc stat.c -o stat.exe
# %run rm -rf tmp && mkdir tmp && touch tmp/a && ln -s ./a tmp/a_link && mkdir tmp/dir
# %run ./stat.exe tmp/a
# %run ./stat.exe tmp/dir
# %run ./stat.exe tmp/a_link
#include <sys/types.h>
#include <sys/stat.h>
#include <fcntl.h>
#include <unistd.h>
#include <stdio.h>
#include <assert.h>
int main(int argc, char *argv[])
{
assert(argc == 2);
struct stat s;
lstat(argv[1], &s); // Не следует по симлинкам, то есть можно узнать stat самого файла симлинки
printf("is regular: %s ", ((s.st_mode & S_IFMT) == S_IFREG) ? "yes" : "no "); // can use predefined mask
printf("is directory: %s ", S_ISDIR(s.st_mode) ? "yes" : "no "); // or predefined macro
printf("is symbolic link: %s\n", S_ISLNK(s.st_mode) ? "yes" : "no ");
return 0;
}
# -
# Сейчас результат для симлинки показан как для самой симлинки. Так как используем специальную функцию.
#
# **open(...O_NOFOLLOW | O_PATH) + fstat** - открываем файл так, чтобы не следовать по симлинкам и далее смотрим его stat
#
# Кстати, открываем не очень честно. С опцией O_PATH нельзя потом применять read, write и еще некоторые операции.
# +
# %%cpp stat.c
# %run gcc stat.c -o stat.exe
# %run rm -rf tmp && mkdir tmp && touch tmp/a && ln -s ./a tmp/a_link && mkdir tmp/dir
# %run ./stat.exe tmp/a
# %run ./stat.exe tmp/dir
# %run ./stat.exe tmp/a_link
#define _GNU_SOURCE // need for O_PATH
#include <sys/types.h>
#include <sys/stat.h>
#include <fcntl.h>
#include <unistd.h>
#include <stdio.h>
#include <assert.h>
int main(int argc, char *argv[])
{
assert(argc == 2);
struct stat s;
int fd = open(argv[1], O_RDONLY | O_NOFOLLOW | O_PATH);
assert(fd >= 0);
fstat(fd, &s);
printf("is regular: %s ", ((s.st_mode & S_IFMT) == S_IFREG) ? "yes" : "no "); // can use predefined mask
printf("is directory: %s ", S_ISDIR(s.st_mode) ? "yes" : "no "); // or predefined macro
printf("is symbolic link: %s\n", S_ISLNK(s.st_mode) ? "yes" : "no ");
close(fd);
return 0;
}
# -
# Здесь тоже результат показан для самой симлинки, поскольку вызываем open со специальными опциями, чтобы не следовать по симлинкам.
#
# **Поэтому важно понимать, какое поведение вы хотите и использовать stat, fstat или lstat**
# ## <a name="time"></a> Извлечем время доступа из атрибутов файла
# +
# %%cpp stat.c
# %run gcc stat.c -o stat.exe
# %run rm -rf tmp && mkdir tmp
# %run touch tmp/a && sleep 2
# %run ln -s ./a tmp/a_link && sleep 2
# %run mkdir tmp/dir
# %run ./stat.exe < tmp/a
# %run ./stat.exe < tmp/dir
# %run ./stat.exe < tmp/a_link
#include <sys/types.h>
#include <sys/stat.h>
#include <fcntl.h>
#include <unistd.h>
#include <stdio.h>
#include <assert.h>
#include <time.h>
int main(int argc, char *argv[])
{
struct stat s;
fstat(0, &s);
printf("update time: %s", asctime(gmtime(&s.st_mtim.tv_sec))); // '\n' есть в строке генерируемой asctime
printf("access time: %s", asctime(gmtime(&s.st_atim.tv_sec)));
return 0;
}
# -
# ## example from man 2 stat
# +
# %%cpp stat.c
# %run gcc stat.c -o stat.exe
# %run rm -rf tmp && mkdir tmp
# %run touch tmp/a
# %run ln -s ./a tmp/a_link
# %run mkdir tmp/dir
# %run ./stat.exe tmp/a
# %run ./stat.exe tmp/dir
# %run ./stat.exe tmp/a_link
# %run ./stat.exe /bin/sh
#include <sys/types.h>
#include <sys/stat.h>
#include <time.h>
#include <stdio.h>
#include <stdlib.h>
int main(int argc, char *argv[])
{
struct stat sb;
if (argc != 2) {
fprintf(stderr, "Usage: %s <pathname>\n", argv[0]);
exit(EXIT_FAILURE);
}
if (stat(argv[1], &sb) == -1) {
perror("stat");
exit(EXIT_FAILURE);
}
printf("File type: ");
switch (sb.st_mode & S_IFMT) {
case S_IFBLK: printf("block device\n"); break;
case S_IFCHR: printf("character device\n"); break;
case S_IFDIR: printf("directory\n"); break;
case S_IFIFO: printf("FIFO/pipe\n"); break;
case S_IFLNK: printf("symlink\n"); break;
case S_IFREG: printf("regular file\n"); break;
case S_IFSOCK: printf("socket\n"); break;
default: printf("unknown?\n"); break;
}
printf("I-node number: %ld\n", (long) sb.st_ino);
printf("Mode: %lo (octal)\n",
(unsigned long) sb.st_mode);
printf("Link count: %ld\n", (long) sb.st_nlink);
printf("Ownership: UID=%ld GID=%ld\n",
(long) sb.st_uid, (long) sb.st_gid);
printf("Preferred I/O block size: %ld bytes\n",
(long) sb.st_blksize);
printf("File size: %lld bytes\n",
(long long) sb.st_size);
printf("Blocks allocated: %lld\n",
(long long) sb.st_blocks);
printf("Last status change: %s", ctime(&sb.st_ctime));
printf("Last file access: %s", ctime(&sb.st_atime));
printf("Last file modification: %s", ctime(&sb.st_mtime));
exit(EXIT_SUCCESS);
}
# -
# ## <a name="username"></a> get user string name
# +
# %%cpp stat.c
# %run gcc stat.c -o stat.exe
# %run rm -r tmp2 ; mkdir tmp2
# %run touch tmp2/a && echo $PASSWORD | sudo -S touch tmp2/b # create this file with sudo
# %run ./stat.exe < tmp2/a # created by me
# %run ./stat.exe < tmp2/b # created by root (with sudo)
#include <sys/types.h>
#include <sys/stat.h>
#include <stdlib.h>
#include <pwd.h>
#include <stdio.h>
#include <assert.h>
int main(int argc, char *argv[])
{
struct stat s;
fstat(0, &s);
struct passwd *pw = getpwuid(s.st_uid);
assert(pw);
printf("%s\n", pw->pw_name);
return 0;
}
# -
# ## <a name="access"></a> Проверка своих прав
# +
# %%cpp stat.c
# %run gcc stat.c -o stat.exe
# %run rm -rf tmp2 && mkdir tmp2
# %run touch tmp2/a
# %run touch tmp2/b && chmod +x tmp2/b
# %run ./stat.exe tmp2/a # usual
# %run ./stat.exe tmp2/aa # not existent file
# %run ./stat.exe tmp2/b # executable
#include <sys/types.h>
#include <sys/stat.h>
#include <stdlib.h>
#include <unistd.h>
#include <stdio.h>
#include <assert.h>
int main(int argc, char *argv[])
{
assert(argc >= 1);
struct stat s;
int status = access(argv[1], X_OK);
fprintf(stderr, "Can execute: %s\n", (status == 0) ? "yes" : "no");
if (status < 0) {
perror("Can not execute because");
}
return 0;
}
# -
# ## <a name="link"></a> Ссылки жесткие и символические
!(rm x.txt ; rm x_hard.txt ; rm x_sym.txt) 2> /dev/null
# !touch x_ordinary.txt
# !touch x.txt
# !link x.txt x_hard.txt
# !ln -s x.txt ./x_sym.txt
# !ls -la x*
# !echo "Hello" > x_ordinary.txt
# !ls -la x*
# ## <a name="fds"></a> Атрибуты файловых дескрипторов
#
# `fcntl(fd, F_GETFD, 0)`, флаг `FD_CLOEXEC`
# +
# %%cpp fcntl_flags.cpp
# %run gcc fcntl_flags.cpp -o fcntl_flags.exe
# %run ./fcntl_flags.exe
#ifndef _GNU_SOURCE
#define _GNU_SOURCE
#endif
#include <stdio.h>
#include <stdlib.h>
#include <unistd.h>
#include <assert.h>
#include <fcntl.h>
#include <sys/resource.h>
#include <sys/types.h>
#include <sys/wait.h>
void describe_fd(const char* prefix, int fd) {
int ret = fcntl(fd, F_GETFD, 0); // Функция принимает 3 аргумента, поэтому обязаны записать все 3 (даже если третий не будет использоваться)
if (ret & FD_CLOEXEC) {
printf("%s: fd %d has CLOEXEC flag\n", prefix, fd);
} else {
printf("%s: fd %d doesn't have CLOEXEC flag\n", prefix, fd);
}
}
int main() {
int fd[2];
pipe(fd);
describe_fd("pipe", fd[0]);
pipe2(fd, O_CLOEXEC);
describe_fd("pipe2 + O_CLOEXEC", fd[0]);
pipe(fd);
fcntl(fd[0], F_SETFD, fcntl(fd[0], F_GETFD, 0) | FD_CLOEXEC); //руками сделали так что у pipe есть флаг O_CLOEXEC
describe_fd("pipe + manually set flag", fd[0]);
return 0;
}
# -
# `fcntl(fd, F_GETFL, 0)`, флаги `O_RDWR`, `O_RDONLY`, `O_WRONLY`, `O_APPEND`, `O_TMPFILE`, `O_ASYNC`, `O_DIRECT`
#
# На самом деле это только ограниченное подмножество флагов из тех, что указываются при открытии файла.
# +
# %%cpp fcntl_open_flags.cpp
# %run gcc fcntl_open_flags.cpp -o fcntl_open_flags.exe
# %run ./fcntl_open_flags.exe
#ifndef _GNU_SOURCE
#define _GNU_SOURCE
#endif
#include <stdio.h>
#include <stdlib.h>
#include <unistd.h>
#include <assert.h>
#include <fcntl.h>
#include <sys/resource.h>
#include <sys/types.h>
#include <sys/wait.h>
void describe_fd(const char* prefix, int fd) {
int ret = fcntl(fd, F_GETFL, 0);
#define flag_cond_str_expanded(flag, mask, name) ((ret & (mask)) == flag ? name : "")
#define flag_cond_str_mask(flag, mask) flag_cond_str_expanded(flag, mask, #flag)
#define flag_cond_str(flag) flag_cond_str_expanded(flag, flag, #flag)
//printf("%d\n", ret & 3);
printf("%s: %s, %s, %s, %s, %s, %s, %s, %s, %s, %s, %s\n", prefix
, flag_cond_str_mask(O_RDONLY, O_RDONLY | O_WRONLY | O_RDWR)
, flag_cond_str_mask(O_WRONLY, O_RDONLY | O_WRONLY | O_RDWR)
, flag_cond_str_mask(O_RDWR, O_RDONLY | O_WRONLY | O_RDWR)
, flag_cond_str(O_TRUNC)
, flag_cond_str(O_APPEND)
, flag_cond_str(O_NONBLOCK)
, flag_cond_str(O_CREAT)
, flag_cond_str(O_CLOEXEC)
, flag_cond_str(O_TMPFILE)
, flag_cond_str(O_ASYNC)
, flag_cond_str(O_DIRECT)
);
}
void check_fd(int fd) {
if (fd < 0) {
perror("open");
assert(fd >= 0);
}
}
int main() {
describe_fd("0 (stdin)", 0);
describe_fd("1 (stdout)", 1);
describe_fd("2 (stderr)", 2);
int f1 = open("fcntl_open_flags.1", O_CREAT|O_TRUNC|O_WRONLY, 0664); check_fd(f1);
describe_fd("f1 O_CREAT|O_TRUNC|O_WRONLY", f1);
int f2 = open("fcntl_open_flags.2", O_CREAT|O_RDWR, 0664); check_fd(f2);
describe_fd("f2 O_CREAT|O_RDWR", f2);
int f3 = open("fcntl_open_flags.2", O_WRONLY|O_APPEND); check_fd(f3);
describe_fd("f3 O_WRONLY|O_APPEND", f3);
int f4 = open("fcntl_open_flags.2", O_RDONLY|O_NONBLOCK|O_ASYNC|O_DIRECT); check_fd(f4);
describe_fd("f4 O_RDONLY|O_NONBLOCK|O_ASYNC|O_DIRECT", f4);
int f5 = open("./", O_TMPFILE|O_RDWR, 0664); check_fd(f5);
describe_fd("f5 O_TMPFILE|O_RDWR", f5);
int fds[2];
pipe2(fds, O_CLOEXEC);
describe_fd("pipe2(fds, O_CLOEXEC)", fds[0]);
return 0;
}
# -
# +
# %%cpp istty.c
# %run gcc istty.c -o istty.exe
# %run ./istty.exe > a.txt
# %run ./istty.exe
#include <unistd.h>
#include <stdlib.h>
#include <pwd.h>
#include <stdio.h>
#include <assert.h>
int main(int argc, char *argv[])
{
if (isatty(STDOUT_FILENO)) {
fprintf(stderr, "\033[0;31mIt's terminal\033[0m\n");
} else {
fprintf(stderr, "It's NOT terminal\n");
}
return 0;
}
# -
# ## <a name="hw"></a> Комментарии к ДЗ
#
# * `PATH_MAX` - есть такая замечательная константа, обязательно узнайте почему она существует и зачем она нужна, перед тем как делать дз. (Иначе есть большая вероятность получить реджект).
# * Есть классные функции `strncmp`, `memcmp`. Прочитайте про них до того, как сравнивать какие-то байтики.
| sem11-file-attributes/file-attrib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center>Climate Action Tracking Prompt:</center>
# ## <center> PDF Data Extraction using NLP </center>
# <hr>
# ### PDF parsing:
#
# PDF is a document format developed by Adobe, based on PostScript. PDFs are highly favoured by the scientific community. Most of the documents and proposals related to climate action are in the PDF format. For parsing these documents, we can consider these PDFs to be divided as follows:
#
# 1. <b>Text:</b> Information can be extracted using NLP.
# 2. <b>Images:</b> Surprisingly there are no Python libraries for extracting images from PDFs, PDF parsers need to be written to extract images.
# 3. <b>Tables:</b> Although libraries are available for extracting tables such as camelot and tabula, they do not always work properly nor do they give consistent and accurate performance.
# ### 1) Parsing Text in PDFs using Python and NLP:
#
# The methodology used is as follows:
#
# #### Reading PDF using PyPDF2:
# +
import PyPDF2
pdf = open('action_example_dublin.pdf', 'rb')
pdfReader = PyPDF2.PdfFileReader(pdf)
#print(pdfReader.numPages)
# creating a list of page objects
page = [pdfReader.getPage(x) for x in range(pdfReader.numPages)]
# extracting text from page
#page_text = [page[x].extractText() for x in range(len(page))]
# closing the pdf object
#pdf.close()
# -
# #### Extracting data using Spacy:
# +
import spacy
import pandas as pd
nlp = spacy.load('en_core_web_md')
# -
doc = nlp(page[20].extractText().replace('\n', ''))
print(doc)
# #### Entitities in the text:
from spacy import displacy
displacy.render(doc, style='ent')
# #### Analysis:
#
# As we can see from the above visualizations, the Name Entity Recognizer (NER) does a pretty good job of identifying different entities even though it is not trained in the context of the given documents. For eg. the quantity NER fails to identify units of measurement such as CO2 and CO2/per employee as part of Quantity Entity. Datasets and Entity relations need to be created in the context of climate action proposals.
#
# Using the default Spacy models will definitely not yield good results for data extraction. However for the sake of prototyping, let us look at an idealized example text and how we might be able to extract information from it using Spacy into Pandas Dataframes.
text = """Dublin is a great city. Dublin has proposed to reduce emissions in the city by 50% till 2025.
At this rate Dublin will thus have net emissions of minus 5% by 2040. The carbon footprint of The Dublin Finance Department
is 1719 tonnes, with the number emissions expected to rise by 2% next year in France. Apple in Dublin has reduced
emissions by 2000 tonnes this year.
"""
text
doc = nlp(text.replace('\n', ' '))
displacy.render(doc, style='ent')
doc = nlp(text)
displacy.render(doc, style='dep')
# +
Columns = {'Organization or Actor': [], 'Location': [], 'Action': [], 'Quantity': [], 'Date or Timeline': []}
df = pd.DataFrame(Columns)
# -
df
for idx, sent in enumerate(doc.sents):
sentence = nlp(str(sent))
actor_list, location_list, action_list, date_list, quantity_list = [], [], [], [], []
#print(sentence.ents)
for token in sentence:
if token.pos_ == "ADJ":
action_list.append(token.text)
if token.pos_ == "VERB":
action_list.append(token.text)
if token.dep_ == "pobj" and token.ent_type_ == "GPE":
location_list.append(token.text)
elif token.ent_type_ == "GPE":
actor_list.append(token.text)
for ent in sentence.ents:
if ent.label_ == "ORG":
actor_list.append(ent.text)
if ent.label_ == "GPE":
span = doc[ent.start:ent.end]
#for token in sentence:
#if token.dep_ == "pobj" and token.ent_type == "GPE":
#location_list.append(ent.text)
#else:
# actor_list.append(ent.text)
if ent.label_ == "DATE":
date_list.append(ent.text)
if ent.label_ == "QUANTITY":
quantity_list.append(ent.text)
if ent.label_ == "PERCENT":
quantity_list.append(ent.text)
#print(actor_list, location_list, action_list, date_list, quantity_list)
if len(actor_list)!=0 and len(quantity_list)!=0 and len(date_list)!=0:
df = df.append({
'Organization or Actor': ', '.join(actor_list),
'Location': ', '.join(location_list),
'Action': ', '.join(action_list),
'Quantity': ', '.join(quantity_list),
'Date or Timeline': ', '.join(date_list)
}, ignore_index = True)
df
# Thus we get the required dataframe which helps to quantify data in the text. This could be further processed using pandas and some text manipulation to get better results.
# ## References Used:
#
# This Jupyter notebook borrows heavily from the following sources:
# - https://course.spacy.io/
# - https://towardsdatascience.com/python-for-pdf-ef0fac2808b0
# - https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5551970/
# ## Summary:
#
# The above code for information was an oversimplified attempt of the research paper titled [Prescription Extraction Using CRFs and Word Embeddings](https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5551970/). By training context specific datasets and entity relationships much better results could be obtained for the above approach.
#
# In the course of working on this project, it was found that there is still a lack of standardization around the documents used for climate action proposals. A standard structure would go a long way in aiding such information extraction from PDFs, especially due to the lack of availability of proper libraries for working with PDFs.
| PDF parsing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python3
# ---
# # The Battle of Neighborhoods | Business Proposal | Introduction
# ## Introduction:
#
# The purpose of this Project is to help people in exploring better facilities around their neighborhood. It will help people making smart and efficient decision on selecting great neighborhood out of numbers of other neighborhoods in Scarborough, Toranto.
#
# Lots of people are migrating to various states of Canada and needed lots of research for good housing prices and reputated schools for their children. This project is for those people who are looking for better neighborhoods. For ease of accessing to Cafe, School, Super market, medical shops, grocery shops, mall, theatre, hospital, like minded people, etc.
#
# This Project aim to create an analysis of features for a people migrating to Scarborough to search a best neighborhood as a comparative analysis between neighborhoods. The features include median housing price and better school according to ratings, crime rates of that particular area, road connectivity, weather conditions, good management for emergency, water resources both freash and waste water and excrement conveyed in sewers and recreational facilities.
#
# It will help people to get awareness of the area and neighborhood before moving to a new city, state, country or place for their work or to start a new fresh life.
#
#
# ## Problem Which Tried to Solve:
#
# The major purpose of this project, is to suggest a better neighborhood in a new city for the person who are shiffting there. Social presence in society in terms of like minded people. Connectivity to the airport, bus stand, city center, markets and other daily needs things nearby.
#
# 1. Sorted list of house in terms of housing prices in a ascending or descending order
# 2. Sorted list of schools in terms of location, fees, rating and reviews
#
#
# ## The Location:
# Scarborough is a popular destination for new immigrants in Canada to reside. As a result, it is one of the most diverse and multicultural areas in the Greater Toronto Area, being home to various religious groups and places of worship. Although immigration has become a hot topic over the past few years with more governments seeking more restrictions on immigrants and refugees, the general trend of immigration into Canada has been one of on the rise.
#
#
# ## Foursquare API:
# This project would use Four-square API as its prime data gathering source as it has a database of millions of places, especially their places API which provides the ability to perform location search, location sharing and details about a business.
#
#
# ## Work Flow:
# Using credentials of Foursquare API features of near-by places of the neighborhoods would be mined. Due to http request limitations the number of places per neighborhood parameter would reasonably be set to 100 and the radius parameter would be set to 500.
#
#
# ## Clustering Approach:
# To compare the similarities of two cities, we decided to explore neighborhoods, segment them, and group them into clusters to find similar neighborhoods in a big city like New York and Toronto. To be able to do that, we need to cluster data which is a form of unsupervised machine learning: k-means clustering algorithm
#
#
# ## Libraries Which are Used to Develope the Project:
# Pandas: For creating and manipulating dataframes.
#
# Folium: Python visualization library would be used to visualize the neighborhoods cluster distribution of using interactive leaflet map.
#
# Scikit Learn: For importing k-means clustering.
#
# JSON: Library to handle JSON files.
#
# XML: To separate data from presentation and XML stores data in plain text format.
#
# Geocoder: To retrieve Location Data.
#
# Beautiful Soup and Requests: To scrap and library to handle http requests.
#
# Matplotlib: Python Plotting Module.
| Week 4 Business Proposal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Defining Column Roles with NimbusML - Relevance Judgement
#
# In NimbusML, when developing a pipeline, (usually for the last learner) users can specify the column roles, such as feature, label, weight, group (for ranking problem), etc.. With this definition, a full dataset with all thoses columns can be fed to the training function. The learner will extract the useful columns from the dataset automatically.
#
# In this tutorial, we solve a learning to rank problem using Microsoft Bing data. More specifically, it is a Subset Ranking problem with real-valued relevance labels. The [full dataset](https://www.microsoft.com/en-us/research/project/mslr/) is available online. Queries and urls provided by a web search engine (Microsoft Bing) are presented as pairs (see the following figure). Each query-url pair is then evaluated by relevence, which takes 5 values (0 for irrelevenat and 4 for perfectly relevant), serving as the label for this machine learning problem. Features are extracted by the Bing team, and those include term frequencies, length of url, etc. for each query-url pair. The model aims at predicting the scores of each pair that in the same order as the relevance within the same group.
#
# In this notebook, due to license issues, we use a randomly generated ranking data just to demonstrate the usage of the package.
from IPython.display import display,IFrame,Image
display(Image(filename='images/Ranking2.PNG'))
# ## Loading Data
# +
# Getting input file path from package
# Due to licensing, user needs to download data files himself
import os
import numpy as np
import pandas as pd
from nimbusml.datasets import get_dataset
from nimbusml import FileDataStream, Role, Pipeline
from nimbusml.ensemble import LightGbmRanker
from nimbusml.feature_extraction.categorical import OneHotHashVectorizer
train_file = get_dataset('msltrtrain').as_filepath()
test_file = get_dataset('msltrtest').as_filepath()
# -
# A snapshot of the input training data is shown below:
#
# |Relevance|QueryID|CoveredQueryTermNumber-Body|StreamLength-Body|InverseDocumentFrequency-Body|SumOfTermFrequency-Title|QualityScore|
# |-----|----------|--------------|-------|--------|-------------|------|
# |2|1|3|156|6.94|2|1|
# |3|1|3|406|6.94|1|3|
# |2|6|3|146|10.22|6|3|
# |5|6|3|209|10.22|4|23|
#
# Each row corresponds to a url-query pair. The first column is the relevance for the url to the corresponding query. The first and second rows are for the same query, but the second url has higher relavance score than the first one. The second column is the query id. The rest columns include the features for this url-query pair. There are 136 features in total, some of which are not shown here. The ranking model predicts the relevance label for each url-query pair.
# We generate the schema and create DataFileStream.
train_stream = FileDataStream.read_csv(
train_file, sep = ',',
header = False,
collapse=True,
names={0:'Label', 1:'GroupId', (2,137):'Features'},
numeric_dtype=np.float32, dtype={'GroupId':str})
test_stream = FileDataStream.read_csv(
test_file,
sep = ',',
header = False,
collapse=True,
names={0:'Label', 1:'GroupId', (2,137):'Features'},
numeric_dtype=np.float32,
dtype={'GroupId':str})
# ## Training
model = Pipeline([
OneHotHashVectorizer(output_kind='Key', columns='GroupId'), #If we can generate schema setting GroupId:U4, we don't need this
LightGbmRanker(feature="Features", label='Label', group_id='GroupId')
])
model.fit(train_stream, 'Label');
# Notice that we add a `OneHotHashVectorizer` transform for the GroupId column. The original dataset has column GroupId as string. However, for LightGbmRanker, the group_id column must be a 'key' type. Therefore, we add this tranformation. The `ToKey` transform would not work in this situation because it would mark any group id's that it didn't encounter in the training dataset as "missing" and cause the ranker to incorrectly see these samples as a single query.
#
# The above code identifies the feature, label and the group_id for LightGbmRanker(). The loss function is estimated per group, and the group_id is needed for each query-url pair. In this case, the group_id is the same as the query id.
#
# There are three ways to specify those roles for different columns, other than the syntax above, the following syntax is also allowed:
#
# from nimbusml import Role
# model = LightGbmRanker() << {Role.Feature: 'Features', Role.Label: 'Label', Role.GroupId: 'GroupID'}
# model = LightGbmRanker(columns = {Role.Feature: 'Features', Role.Label: 'Label', Role.GroupId: 'GroupID'})
# Not only for ranking problems, the same syntax can also be used for classificaion, regression, etc.. In the fitting function, no "y" will be needed as long as all the columns are included in the training dataset (either a dataframe or a FileDataStream).
# ## Evaluation
# Similar to other examples with pipeline, the model can be evaluated using .test(). A perfect ranking model would predict higher score for url-query pair with high relevance. We can see from the above table that all the test samples belong to the same query. Urls have higher relevance tend to have higher score. Note that the model is only trained with 100 rows of data and should be improved with large sample size.
# +
metrics, scores = model.test(test_stream, 'Label', group_id = 'GroupId', evaltype='ranking', output_scores = True)
import pandas as pd
scores[['Relevance','QueryID']] = pd.read_csv(test_file, header = None)[[0,1]]
# Show the top 10 predicted items for query 5.
print("Prediction scores: ")
scores[scores.QueryID == 5].sort_values('Score', ascending=False).head(10)
# -
# ## Discussion
# The Normalized Discounted Cumulative Gain (NDCG) and Discounted Cumulative Gain (DCG) at a particular rank position (1,2, or 3) are reported by the .test() function if setting evaltype='ranking'. DCG evaluates the quality of a rank with respect to the original relevance scores. It is the sum of the discounted relevance scores for the top N items in the ranking. The discounting factor, which is logarithmically proportional to the position of the item, is meant to penalize highly relevant items that appear lower in the ranking. DCG can be computed as follows:
#
# <img src="images/2-5-1.png" align = "middle"/>
#
# Where N is the threshold of ranking to account for in DCG, and g<sub>i</sub> the relavance score for the i<sup>th</sup> item in the ranking. Since the number of items in a result can vary by query, DCG is not a consistent metric when comparing results for multiple queries. NDCG normalizes DCG by the max(DCG) or the Ideal DCG (IDCG), which is the DCG for the perfect ranking where all items are ranked in decreasing order of relevance. This scales NDCG to a range of [0,1] so that comparisons of rankings across multiple queries are possible.
print("Performance Metrics: ")
metrics
| samples/2.5 [Numeric] Learning-to-Rank with Microsoft Bing Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.10 64-bit
# name: python3
# ---
# +
# This is a sample script to analyze the results of measuring talker_listener.
# Please build the trace_analysis package before using it.
# The source code used for the measurement can be found here:
# https://github.com/hsgwa/trace_samples
# +
import os
import sys
from pathlib import Path
sys.path.append(os.path.join(os.path.dirname(Path().resolve()), '..'))
# -
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
# build and source trace_analysis package
from trace_analysis import Architecture, Application, Lttng
trace_dir_path = './cyclic_pipeline_intra_process/'
lttng = Lttng(trace_dir_path, force_conversion=True)
arch = Architecture()
architecture_path = './cyclic_pipeline_intra_process/architecture.yaml'
arch.import_file(file_path = architecture_path, file_type='yaml', latency_composer=lttng)
app = Application(arch)
# +
node = app.nodes[0]
print('node_name: ' + node.node_name)
callback = node.callbacks[0]
print('callback_type: ' + callback.TYPE_NAME)
print('subscription topic name :' + callback.topic_name)
print('callback_name: ' + callback.callback_name)
print('callback symbol: ' + callback.symbol)
print('callback unique name: ' + callback.unique_name)
# -
start_callback_unique_name = app.callbacks[0].unique_name
end_callback_unique_name = app.callbacks[0].unique_name
print('path search')
print('start_callback: ' + start_callback_unique_name)
print('end_callback: ' + end_callback_unique_name)
paths = app.search_paths(start_callback_unique_name, end_callback_unique_name)
path = paths[0]
t, latency = path.to_timeseries()
plt.plot(t, latency)
path.to_timeseries()[1]
histogram, bins = path.to_histogram(binsize_ns=100000)
plt.step(bins[:-1], histogram, where='post')
df = path.to_dataframe(remove_dropped=True)
df
# +
for i, row in df.iterrows():
x = row.values
y = np.array(range(len(x))) * -1
plt.plot(x, y, marker='o')
break
plt.yticks(y, df.columns)
# -
| trace_analysis/sample/lttng_samples/cyclic_pipeline_intra_process.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # So easy, *voilà*!
#
# In this example notebook, we demonstrate how voila can render custom interactive matplotlib figures using the [ipympl](https://github.com/matplotlib/ipympl) widget.
# +
# %matplotlib widget
import ipympl
import numpy as np
import matplotlib.pyplot as plt
x = np.linspace(0, 2 * np.pi, 500)
y1 = np.sin(x)
y2 = np.sin(3 * x)
fig, ax = plt.subplots()
ax.fill(x, y1, 'b', x, y2, 'r', alpha=0.3)
plt.show()
# +
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('ggplot')
fig, axes = plt.subplots(ncols=2, nrows=2)
ax1, ax2, ax3, ax4 = axes.ravel()
# scatter plot (Note: `plt.scatter` doesn't use default colors)
x, y = np.random.normal(size=(2, 200))
ax1.plot(x, y, 'o')
# sinusoidal lines with colors from default color cycle
L = 2 * np.pi
x = np.linspace(0, L)
ncolors = len(plt.rcParams['axes.prop_cycle'])
shift = np.linspace(0, L, ncolors, endpoint=False)
for s in shift:
ax2.plot(x, np.sin(x + s), '-')
ax2.margins(0)
# bar graphs
x = np.arange(5)
y1, y2 = np.random.randint(1, 25, size=(2, 5))
width = 0.25
ax3.bar(x, y1, width)
ax3.bar(x + width, y2, width,
color=list(plt.rcParams['axes.prop_cycle'])[2]['color'])
ax3.set_xticks(x + width)
ax3.set_xticklabels(['a', 'b', 'c', 'd', 'e'])
# circles with colors from default color cycle
for i, color in enumerate(plt.rcParams['axes.prop_cycle']):
xy = np.random.normal(size=2)
ax4.add_patch(plt.Circle(xy, radius=0.3, color=color['color']))
ax4.axis('equal')
ax4.margins(0)
plt.show()
# -
| notebooks/ipympl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/karencfisher/COVID19/blob/main/notebooks/covidradio_weighted_baseline.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="z-UlU3Kxziv_" outputId="6aa25bec-a36f-4359-9638-569e6cad4ad5"
from google.colab import drive
drive.mount('/content/drive')
BASE_PATH = '/content/drive/MyDrive/COVID-19_Radiography_Dataset'
# !unzip -q /content/drive/MyDrive/COVID-19_Radiography_Dataset/data.zip
# !wget https://raw.githubusercontent.com/karencfisher/COVID19/main/tools/util.py
# + id="a35edfe8"
import os
import random
import shutil
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
import cv2
from sklearn.metrics import confusion_matrix, roc_curve, roc_auc_score, classification_report
from sklearn.utils import class_weight
from tensorflow.keras.models import Sequential, load_model
from tensorflow.keras.layers import Dense, Conv2D, MaxPooling2D, Dropout, Flatten
from tensorflow.keras.preprocessing.image import ImageDataGenerator, load_img, img_to_array
from tensorflow.keras.callbacks import EarlyStopping, ModelCheckpoint, ReduceLROnPlateau
from tensorflow.keras import backend as K
import util
# + [markdown] id="742f7c67"
# #### Get metadata on training set
# + id="ff683d55"
classes = ('normal', 'COVID')
data_path = 'data'
counts = []
filelists = []
for item in classes:
filelist = os.listdir(os.path.join(data_path, 'train', item))
filelists.append(filelist)
counts.append(len(filelist))
# + [markdown] id="5b616950"
# #### Visualize samples of the image data by class
# + colab={"base_uri": "https://localhost:8080/", "height": 309} id="1aae1989" outputId="c8d79427-1332-4d4e-e7fa-c49408a3f0e8"
f, axes = plt.subplots(len(classes), 3, sharey=True)
f.set_figwidth(10)
plt.tight_layout()
for i, item in enumerate(classes):
images = random.sample(filelists[i], 3)
for j, image in enumerate(images):
img_file = os.path.join(data_path, 'train', item, image)
img = plt.imread(img_file)
axes[i][j].imshow(img, cmap='gray')
axes[i][j].set_title(item)
# + [markdown] id="9b6309cb"
# #### Plot prevalence of classes
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="ecf54e07" outputId="cc0c6b94-8c00-466d-c1f3-3cba2c041eca"
plt.bar(classes, counts)
plt.title('Prevalence of Classes');
# + [markdown] id="62848c1f"
# #### Data generators to import images into the classifier
# + colab={"base_uri": "https://localhost:8080/"} id="2f25e256" outputId="bd97486d-4225-4341-ce95-7d5c52024fc7"
datagen = ImageDataGenerator(rescale = 1 / 255.)
train_path = os.path.join(data_path, 'train')
train_gen = datagen.flow_from_directory(directory=train_path,
target_size=(299, 299),
batch_size=32,
shuffle=True,
class_mode='binary',
classes = classes)
valid_path = os.path.join(data_path, 'valid')
valid_gen = datagen.flow_from_directory(directory=valid_path,
target_size=(299, 299),
batch_size=32,
shuffle=False,
class_mode='binary',
classes=classes)
test_path = os.path.join(data_path, 'test')
test_gen = datagen.flow_from_directory(directory=test_path,
target_size=(299, 299),
batch_size=32,
shuffle=True,
class_mode='binary',
classes = classes)
# + [markdown] id="b0998e41"
# #### Build a simple CNN as a baseline
# + colab={"base_uri": "https://localhost:8080/"} id="ecdb2234" outputId="58bdd35d-a2a2-4262-f69f-673adf722dc8"
model = Sequential([Conv2D(32, (3,3), activation='relu', input_shape=(299, 299, 3)),
MaxPooling2D(2, 2),
Conv2D(32, (3,3), activation='relu'),
MaxPooling2D(2, 2),
Conv2D(64, (3,3), activation='relu'),
MaxPooling2D(2, 2),
Conv2D(128, (3,3), activation='relu'),
MaxPooling2D(2, 2),
Conv2D(256, (3,3), activation='relu'),
MaxPooling2D(2, 2),
Flatten(),
Dropout(0.2),
Dense(256, activation='relu'),
Dense(128, activation='relu'),
Dense(1, activation='sigmoid')])
model.summary()
# + [markdown] id="Q8dYbN57JVod"
#
# + [markdown] id="kMYGfQBZIZkA"
# #### Weighted binary crossentropy function
#
# $$\mathcal{L}(y, \hat{y}, \epsilon) = - \frac{1}{N} \sum (w_{p} y \log(\hat{y} + \epsilon) + w_{n}(1-y) \log( 1 - \hat{y} + \epsilon ) ) $$
# + [markdown] id="b94cbc9f"
# #### Compile and train
# + colab={"base_uri": "https://localhost:8080/"} id="d937ac75" outputId="96525811-bba0-4151-db47-81a83bac764b"
model.compile(loss=util.Weighted_Loss(train_gen.classes), optimizer='adam', metrics=['accuracy'])
stop = EarlyStopping(monitor='val_loss', patience=5, restore_best_weights=True)
reduce_lr = ReduceLROnPlateau(monitor='val_loss', factor=0.2,
patience=5, min_lr=0.001)
history = model.fit(train_gen,
epochs=25,
steps_per_epoch=len(train_gen),
validation_data=valid_gen,
validation_steps=len(valid_gen),
verbose=1,
callbacks=[stop, reduce_lr])
# + id="60159680"
model_path = os.path.join(BASE_PATH, 'weighted_model.h5')
model.save(model_path)
# + [markdown] id="624f109a"
# #### Evaluate the model
#
# (Accuracy is not everything.)
# + id="4f773d19" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="6c6d9a32-dc09-4edf-aa83-4d5c6d7f151a"
plt.plot(history.history['accuracy'], 'r', label='Accuracy Training')
plt.plot(history.history['val_accuracy'], 'b', label='Accuracy Validation')
plt.title('Accuracy Training and Validation')
plt.ylabel('Accuracy')
plt.xlabel('Epoch')
plt.legend(loc=0)
plt.show()
# + id="UP5z6fW_u6Dt"
model_path = os.path.join(BASE_PATH, 'weighted_model.h5')
model = load_model(model_path, compile=False)
model.compile(loss=util.Weighted_Loss(train_gen.classes), optimizer='adam',
metrics=['accuracy'])
# + id="77884219" colab={"base_uri": "https://localhost:8080/"} outputId="e9f5e41f-be68-4147-e144-3b9b9a58d504"
score = model.evaluate(valid_gen, steps=len(valid_gen))
print(f'Accuracy Score = {round(score[1], 2) * 100}%')
# + [markdown] id="98f397e8"
# #### Calculate confusion matrix, precision, recall, f1
# + id="e5030343" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="ef8c64f3-7d28-4af0-f643-45a57d33a30b"
y_true = np.array(valid_gen.classes)
y_true = np.expand_dims(y_true, axis=1)
y_raw_pred = model.predict(valid_gen, verbose=1)
y_pred = y_raw_pred > 0.5
y_pred = y_pred.astype(np.uint8)
cf = confusion_matrix(y_true, y_pred)
sns.heatmap(cf, xticklabels=classes, yticklabels=classes, annot=True, fmt='.2f');
# + [markdown] id="4e7583b1"
# #### ROC-AUC curve and AUC score
# + [markdown] id="a686abdb"
# #### Calculate Sensitivity, specificity, and PPV
# Calculate PPV (Positive Predictive Value) using Bayes Theorem
#
#
# $$ PPV = \frac{sensitivity \times prevalence}{sensitivity\times prevalence + (1 - specificty)\times(1 - prevalence)} $$
# + id="fa16c625" colab={"base_uri": "https://localhost:8080/", "height": 80} outputId="9366dd8a-e81f-4e0e-fd80-95bde7b6e63d"
labels = ['covid']
metrics_df, roc_curves = util.model_metrics(y_true, y_pred, labels)
metrics_df
# + id="1c4834ec" colab={"base_uri": "https://localhost:8080/", "height": 295} outputId="05fbf94c-2f89-438c-cab8-622feb4c1ee2"
fpr, tpr, _ = roc_curve(y_true, y_raw_pred)
plt.plot(fpr, tpr)
plt.xlabel("FPR")
plt.ylabel("TPR")
plt.title(f'ROC-AUC curve')
plt.show()
# + [markdown] id="R4ti-AsDY_YA"
# #### Individual prediction from the test set
#
# We'll just pick a random example and run it through the model
# + id="zhwE2DYAWz0l"
counts = []
filelists = []
for item in classes:
filelist = os.listdir(os.path.join(data_path, 'test', item))
filelists.append(filelist)
counts.append(len(filelist))
# + colab={"base_uri": "https://localhost:8080/", "height": 703} id="BoMXv1iG8aey" outputId="a09683bc-2302-4d55-ac05-df171a152e01"
for i, item in enumerate(classes):
f, axes = plt.subplots(1, 3, sharey=True)
f.set_figwidth(15)
plt.tight_layout()
image_file = random.choice(filelists[i])
image_path = os.path.join(data_path, 'test', item, image_file)
image = load_img(image_path)
image = img_to_array(image)
pred_image = np.expand_dims(image, axis=0)
pred_image = pred_image / 255.
pred = model.predict(pred_image)
prediction = int(pred[0][0] >= 0.5)
if prediction:
confidence = round(pred[0][0] * 100, 2)
else:
confidence = round((1 - pred[0][0]) * 100, 2)
img = plt.imread(image_path)
axes[0].imshow(img, cmap='gray')
axes[0].set_title(item)
cam = util.grad_cam(model, pred_image, 0, 'conv2d_4')
img = plt.imread(image_path)
axes[1].imshow(img, cmap='gray')
pos = axes[1].imshow(cam, alpha=0.5)
f.colorbar(pos, ax=axes[1])
axes[1].set_title(item)
cam_gs = cv2.cvtColor(cam, cv2.COLOR_RGB2GRAY)
axes[2].imshow(img, cmap='gray')
axes[2].contour(cam_gs)
axes[2].set_title(item)
plt.show()
print(f'Actual class: {item}')
print(f'Predicted class: {classes[prediction]}')
print(f'Confidence: {confidence}%')
# + id="x285ZHUlc5jV"
| notebooks/covidradio_weighted_baseline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modelos de Lenguaje
# +
# %matplotlib inline
import codecs
import numpy as np
import pandas as pd
# -
with codecs.open('food.txt', 'r') as f:
raw_corpus = []
for line in f:
raw_corpus.append('<s> {} </s>'.format(line.strip()))
raw_corpus = ' '.join(raw_corpus)
raw_corpus
corpus = np.array(raw_corpus.split(' '))
corpus
# ## Modelo de Unigrama
unique, counts = np.unique(corpus, return_counts=True)
repetitions = dict(zip(unique, counts))
# P(A)
prob_A = repetitions['A'] / corpus.shape[0]
prob_A
# P(B)
prob_B = repetitions['B'] / corpus.shape[0]
prob_B
# P(C)
prob_C = repetitions['C'] / corpus.shape[0]
prob_C
# ## Modelo de Bigramas
bigram_matrix = pd.DataFrame(index=unique, columns=unique)
bigram_matrix = bigram_matrix.fillna(0)
bigram_matrix = bigram_matrix.drop(['</s>'], axis=0)
bigram_matrix = bigram_matrix.drop(['<s>'], axis=1)
bigram_matrix
for i, w1 in enumerate(corpus):
if w1 != '</s>':
try:
w2 = corpus[i + 1]
if w2 != '<s>':
bigram_matrix.at[w1, w2] += 1
except IndexError:
break
bigram_matrix
# +
for word in unique:
try:
bigram_matrix.loc[[word]] = (bigram_matrix.loc[[word]] + 1) / (repetitions[word] + unique.shape[0])
except KeyError:
continue
bigram_matrix
# -
bigram_matrix.T.plot()
# P(ACB) = P(A)*P(C|A)*P(B|C)
prob_ACB = prob_A * bigram_matrix.at['A', 'C'] * bigram_matrix.at['C', 'B']
prob_ACB
# P(CAB) =
prob_CAB = prob_C * bigram_matrix.at['C', 'A'] * bigram_matrix.at['A', 'B']
prob_CAB
| Sesion3.ipynb |
(* -*- coding: utf-8 -*-
(* --- *)
(* jupyter: *)
(* jupytext: *)
(* text_representation: *)
(* extension: .ml *)
(* format_name: light *)
(* format_version: '1.5' *)
(* jupytext_version: 1.14.4 *)
(* kernelspec: *)
(* display_name: OCaml 4.07.1 *)
(* language: OCaml *)
(* name: ocaml-jupyter *)
(* --- *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* <center> *)
(* *)
(* <h1 style="text-align:center"> Lambda Calculus : Encodings </h1> *)
(* <h2 style="text-align:center"> CS3100 Fall 2019 </h2> *)
(* </center> *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Review *)
(* *)
(* ### Previously *)
(* *)
(* * Semantics of untyped lambda calculus *)
(* + β-reductions, reduction strategies, normal forms, extensionality *)
(* *)
(* ### Today *)
(* *)
(* * Encodings *)
(* + Booleans, Arithmetic, Pairs, Recursion *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Power of Lambdas *)
(* *)
(* * Despite its simplicity, the lambda calculus is quite expressive: it is **Turing complete**! *)
(* * Means we can encode any computation we want *)
(* + if we are sufficiently clever... *)
(* * Examples *)
(* + Booleans & predicate logic. *)
(* + Pairs *)
(* + Lists *)
(* + Natural numbers & arithmetic. *)
(* *)
(* $\newcommand{\br}{\rightarrow_{\beta}}$ *)
(* + slideshow={"slide_type": "slide"}
#use "init.ml"
let p = Lambda_parse.parse_string
let var x = Var x
let app l =
match l with
| [] -> failwith "ill typed app"
| [x] -> x
| x::y::xs -> List.fold_left (fun expr v -> App (expr, v)) (App(x,y)) xs
let lam x e = Lam(x,e)
let eval ?(log=true) ?(depth=1000) s =
s
|> Eval.eval ~log ~depth Eval.reduce_normal
|> Syntax.string_of_expr
(* -
p "\\x.x";;
var "x";;
app [var "x"; var "y"; var "z"];;
lam "x" (var "y");;
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Booleans *)
(* + slideshow={"slide_type": "fragment"}
let tru = p "\\t.\\f.t"
let fls = p "\\t.\\f.f"
(* + [markdown] slideshow={"slide_type": "fragment"}
(* * Now we can define a `test` function such that *)
(* + `test tru v w` $\br$ `v` *)
(* + `test fls v w` $\br$ `w` *)
(* + slideshow={"slide_type": "fragment"}
let test = p "\\l.\\m.\\n.l m n"
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Booleans *)
(* *)
(* Now *)
(* *)
(* ```ocaml *)
(* test tru v w *)
(* ``` *)
(* *)
(* evaluates to *)
(* + slideshow={"slide_type": "fragment"}
eval @@ app [test; app [lam "x" (var "x"); tru]; var "v"; var "w"]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Booleans *)
(* *)
(* Similarly, *)
(* *)
(* ```ocaml *)
(* test fls v w *)
(* ``` *)
(* *)
(* evaluates to *)
(* -
eval @@ app [test; fls; var "v"; var "w"]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Booleans *)
(* + [markdown] slideshow={"slide_type": "-"}
(* `fls` itself is a function. `test fls v w` is equivalent to `fls v w`. *)
(* + slideshow={"slide_type": "fragment"}
eval @@ app [fls; var "v"; var "w"]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Logical operators *)
(* *)
(* ```ocaml *)
(* and = λb.λc.b c fls *)
(* or = λb.λc.b tru c *)
(* not = λb.b fls tru *)
(* ``` *)
(* + slideshow={"slide_type": "fragment"}
let and_ = lam "b" (lam "c" (app [var "b"; var "c"; fls]))
let or_ = lam "b" (lam "c" (app [var "b"; tru; var "c"]))
let not_ = lam "b" (app [var "b"; fls; tru])
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Logical Operators *)
(* + slideshow={"slide_type": "-"}
eval @@ app [and_; tru; fls]
(* + [markdown] slideshow={"slide_type": "fragment"}
(* The above is a **proof** for `true /\ false = false` *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Logical operators *)
(* *)
(* Encode implies using standard formulation. *)
(* *)
(* \\[ *)
(* \begin{array}{rl} *)
(* & p \implies q \equiv \neg p \vee q \\ *)
(* \mathbf{Theorem 1.} & a \wedge b \implies a *)
(* \end{array} *)
(* \\] *)
(* + slideshow={"slide_type": "fragment"}
let implies = lam "p" (lam "q" (app [or_; app [not_; var "p"]; var "q"]))
let thm1 = lam "a" (lam "b" (app [implies; app [and_; var "a"; var "b"]; var "a"]))
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Logical operators *)
(* -
eval ~log:false (app [thm1; tru; fls])
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz *)
(* *)
(* What is the lambda calculus encoding for `xor x y` *)
(* *)
(* | x | y | xor x y | *)
(* |---|---|:-------:| *)
(* | T | T | F | *)
(* | T | F | T | *)
(* | F | T | T | *)
(* | F | F | F | *)
(* *)
(* 1. x x y *)
(* 2. x (y tru fls) y *)
(* 3. x (y fls tru) y *)
(* 4. y x y *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz *)
(* *)
(* What is the lambda calculus encoding for `xor x y` *)
(* *)
(* | x | y | xor x y | *)
(* |---|---|:-------:| *)
(* | T | T | F | *)
(* | T | F | T | *)
(* | F | T | T | *)
(* | F | F | F | *)
(* *)
(* 1. x x y *)
(* 2. x (y tru fls) y *)
(* 3. x (y fls tru) y ✅ *)
(* 4. y x y *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Pairs *)
(* -
let mk_pair x y = (x,y)
let fst (x,y) = x
let snd (x,y) = y
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Pairs *)
(* *)
(* * Encoding of a pair `(a,b)` *)
(* + Pair Constructor : (a,b) = λf.λs.λb.b f s *)
(* -
let pair = p "λf.λs.λb.b f s"
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Pairs *)
(* -
eval @@ app [pair; var "v"; var "w"]
(* * The pair **value** is a function that takes a **boolean** as an argument and applies the elements of the pair to it. *)
(* * `b` is a boolean is a **convention** that we should follow. *)
(* + No type safety. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Pair accessor functions *)
(* *)
(* * Recall that a pair value is a function `λb.b v w` *)
(* + where `v` and `w` are the first and second elements of the pair. *)
(* * We can define accessors `fst` and `snd` as follows: *)
(* + fst = λp.p tru *)
(* + snd = λp.p fls *)
(* -
let fst = lam "p" (app [var "p"; tru])
let snd = lam "p" (app [var "p"; fls])
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Pair accessor functions *)
(* -
eval ~log:true @@ app [fst; app[pair; var "v"; var "w"]]
eval ~log:false @@ app [snd; app [pair; var "v"; var "w"]]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Pair swap function *)
(* *)
(* In OCaml, *)
(* *)
(* ```ocaml *)
(* let swap (x,y) = (y,x) *)
(* ``` *)
(* *)
(* In lambda calculus, *)
(* *)
(* ```ocaml *)
(* swap = λp.λb.b (snd p) (fst p) *)
(* ``` *)
(* -
let swap = lam "p" (lam "b" (app [var "b"; app [snd;var "p"]; app [fst; var "p"]]))
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Pair swap function *)
(* *)
(* Let's try *)
(* *)
(* ```ocaml *)
(* fst (swap (v,w)) *)
(* ``` *)
(* -
eval ~log:false @@ app [fst; app [swap; app[pair; var "v"; var "w"]]]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Natural numbers *)
(* *)
(* * 0 = λs.λz.z *)
(* * 1 = λs.λz.s z *)
(* * 2 = λs.λz.s (s z) *)
(* * 3 = λs.λz.s (s (s z)) *)
(* *)
(* i.e., n = λs.λz.(apply `s` n times to `z`) *)
(* *)
(* Also known as **Church numerals**. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Natural numbers *)
(* + slideshow={"slide_type": "-"}
let zero = p ("λs.λz.z")
let one = p ("λs.λz.s z")
let two = p ("λs.λz.s (s z)")
let three = p ("λs.λz.s (s (s z))")
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz *)
(* *)
(* What will be the OCaml type of church encoded numeral? *)
(* *)
(* 1. `('a -> 'b) -> 'a -> 'b` *)
(* 2. `('a -> 'a) -> 'a -> 'a` *)
(* 3. `('a -> 'a) -> 'b -> int` *)
(* 4. `(int -> int) -> int -> int` *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz *)
(* *)
(* What will be the OCaml type of church encoded numeral? *)
(* *)
(* 1. `('a -> 'b) -> 'a -> 'b` *)
(* 2. `('a -> 'a) -> 'a -> 'a` ✅ *)
(* 3. `('a -> 'a) -> 'b -> int` *)
(* 4. `(int -> int) -> int -> int` *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Operations on numbers: Successor *)
(* *)
(* Successor function is: *)
(* *)
(* ```ocaml *)
(* scc = λn.λs.λz.s (n s z) *)
(* ``` *)
(* + slideshow={"slide_type": "fragment"}
let scc = p ("λn.λs.λz.s (n s z)")
(* + slideshow={"slide_type": "-"}
eval @@ app [scc; zero]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Operations on numbers : is_zero *)
(* *)
(* Check if the given number is zero: *)
(* *)
(* ```ocaml *)
(* is_zero = λn.n (λy.fls) tru *)
(* ``` *)
(* + slideshow={"slide_type": "fragment"}
let is_zero = lam "n" (app [var "n"; lam "y" fls; tru])
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Operations on numbers : is_zero *)
(* -
eval @@ app [is_zero; zero]
eval @@ app [is_zero; one]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic *)
(* *)
(* ```ocaml *)
(* plus = λm.λn.λs.λz.m s (n s z) *)
(* mult = λm.λn.λs.m (n s) *)
(* ``` *)
(* + slideshow={"slide_type": "fragment"}
let plus = p ("λm.λn.λs.λz.m s (n s z)")
let mult = p ("λm.λn.λs.m (n s)")
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: addition *)
(* -
eval @@ app [plus; one; two]
(* + [markdown] slideshow={"slide_type": "fragment"}
(* Proves 1 + 2 = 3. Can build a theory of arithmetic over lambda calculus. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: multiplication *)
(* -
eval @@ app [mult; two; zero]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: predecessor *)
(* *)
(* It turns out predecessor function is much more tricky compared to successor. *)
(* *)
(* ```ocaml *)
(* zz = pair zero zero *)
(* ss = λp. pair (snd p) (plus one (snd p)) *)
(* ``` *)
(* + [markdown] slideshow={"slide_type": "fragment"}
(* ```ocaml *)
(* zz = (0,0) *)
(* ss zz = (0,1) *)
(* ss (ss zz) = (1,2) *)
(* ss (ss (ss zz)) = (2,3) *)
(* ``` *)
(* etc. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: predecessor *)
(* *)
(* It turns out predecessor function is much more tricky compared to successor. *)
(* *)
(* ```ocaml *)
(* zz = pair zero zero *)
(* ss = λp. pair (snd p) (plus one (snd p)) *)
(* prd = λm. fst (m ss zz) *)
(* ``` *)
(* + slideshow={"slide_type": "fragment"}
let zz = app [pair; zero; zero]
let ss = lam "p" (app [pair; app [snd; var "p"]; app [plus; one; app [snd; var "p"]]])
let prd = lam "m" (app [fst; app [var "m"; ss; zz]])
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: Predecessor *)
(* -
eval ~log:false @@ app [prd; three]
eval ~log:false @@ app [prd; zero]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: Subtraction *)
(* *)
(* `sub` computes `m-n`: *)
(* *)
(* ```ocaml *)
(* sub = λm.λn.n prd m *)
(* ``` *)
(* *)
(* Intuition: apply predecessor `n` times on `m`. *)
(* -
let sub = lam "m" (lam "n" (app [var "n"; prd; var "m"]))
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: Subtraction *)
(* -
eval ~log:false @@ app [sub; three; two]
eval ~log:false @@ app [sub; two; three]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: equal *)
(* *)
(* * `m - n = 0` $\implies$ `m = n`. *)
(* + But we operate on natural numbers. *)
(* + `3 - 4 = 0` $\implies$ `3 = 4`. *)
(* * `m - n = 0 && n - m = 0` $\implies$ `m = n`. *)
(* -
let equal =
let mnz = app [is_zero; app [sub; var "m"; var "n"]] in
let nmz = app [is_zero; app [sub; var "n"; var "m"]] in
lam "m" (lam "n" (app [and_; mnz; nmz]))
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Arithmetic: equal *)
(* -
eval ~log:false @@ app [equal; two; two]
eval ~log:false @@ app [equal; app[sub; three; two]; two]
eval ~log:false @@ app [equal; app[sub; two; three]; zero]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Fixed points *)
(* *)
(* * Given a function $f$, if $x = f(x)$ then $x$ is said to be a fixed point for $f$. *)
(* + $f(x) = x^2$ has two fixed points 0 and 1. *)
(* + $f(x) = x + 1$ has no fixed points. *)
(* * For lambda calculus, $N$ is said to be a fixed point of $F$ if $F ~N =_{\beta} N$ *)
(* + In the untyped lambda calculus, every term F has a fixed point! *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Fixed points *)
(* *)
(* * Let `D = λx.x x`, then *)
(* + `D D = (λx.x x) (λx.x x)` $\rightarrow_{\beta}$ `(λx.x x) (λx.x x) = D D`. *)
(* * So `D D` is an infinite loop *)
(* + In general, self-application is how you get looping *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Fixed points *)
(* *)
(* $ *)
(* \require{color} *)
(* \newcommand{\yb}[1]{\colorbox{yellow}{$#1$}} *)
(* $ *)
(* *)
(* Let $Y = \lambda f.(\lambda x.f ~(x ~x)) ~(\lambda x.f ~(x ~x))$, then *)
(* *)
(* \\[ *)
(* \begin{array}{rl} *)
(* & Y ~F = (\lambda \yb{f}.(\lambda x.f ~(x ~x)) ~(\lambda x.f ~(x ~x))) ~F \\ *)
(* \rightarrow_{\beta} & (\lambda \yb{x}.F ~(x ~x)) ~\yb{(\lambda x.F ~(x ~x))} \\ *)
(* \rightarrow_{\beta} & F \yb{((λx.F (x x)) (λx.F (x x)))} \\ *)
(* \rightarrow_{\beta} & F ~(Y ~F) *)
(* \end{array} *)
(* \\] *)
(* *)
(* * Therefore, `Y F = F(Y F)`. *)
(* + `Y F` is said to be the fixed point of `F`. *)
(* + `Y F = F(Y F) = F(F(Y F)) = ...` *)
(* + `Y` (`y`-combinator) can be used to achieve recursion. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Fixed point: Factorial *)
(* *)
(* ```ocaml *)
(* fact = λf.λn.if n = 0 then 1 else n * f (n-1) *)
(* ``` *)
(* *)
(* * Second argument `n` is the integer. *)
(* * First argument `f` is the function to call for the recursive case. *)
(* * We will use y-combinator to achieve recursion. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Fixed point: Factorial *)
(* *)
(* \\[ *)
(* \begin{array}{rl} *)
(* & (Y ~\text{fact}) ~1 = ~\text{fact} ~(Y ~\text{fact}) 1 \\ *)
(* \rightarrow_{beta} & \text{if } 1 = 0 \text{ then } 1 \text{ else } 1 * ((Y \text{ fact}) ~0) \\ *)
(* \rightarrow_{beta} & 1 * ((Y \text{ fact}) ~0) \\ *)
(* \rightarrow_{beta} & 1 * (\text{fact } (Y \text{ fact}) ~0) \\ *)
(* \rightarrow_{beta} & 1 * \text{if } 0 = 0 \text{ then } 1 \text{ else } 1 * ((Y \text{ fact}) ~0) \\ *)
(* \rightarrow_{beta} & 1 * 1 \\ *)
(* \rightarrow_{beta} & 1 \\ *)
(* \end{array} *)
(* \\] *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Fixed point: Factorial *)
(* *)
(* -
let y = p "λf.(λx.f (x x)) (λx.f (x x))"
let fact =
let tst = app [is_zero; var "n"] in
let fb = app [mult; var "n"; app [var "f"; app [prd; var "n"]]] in
lam "f" (lam "n" (app [tst; one; fb]))
(* + slideshow={"slide_type": "slide"}
eval ~log:false @@ app [y; fact; two]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz *)
(* *)
(* The y-combinator Y = λf.(λx.f (x x)) (λx.f (x x)) is a fixed pointer combinator under which reduction strategy? *)
(* *)
(* 1. Call-by-value *)
(* 2. Call-by-name *)
(* 3. Both *)
(* 4. Neither *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Quiz *)
(* *)
(* The y-combinator Y = λf.(λx.f (x x)) (λx.f (x x)) is a fixed pointer combinator under which reduction strategy? *)
(* *)
(* 1. Call-by-value *)
(* 2. Call-by-name ✅ *)
(* 3. Both *)
(* 4. Neither *)
(* *)
(* Under call-by-value, we will keep indefinitely expanding `Y F = F (Y F) = F (F (Y F)) = ...`. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Fixed point: Z combinator *)
(* *)
(* There is indeed a fixed point combinator for call-by-value called the Z combinator *)
(* *)
(* ```ocaml *)
(* Z = λf. (λx. f (λy. x x y)) (λx. f (λy. x x y)) *)
(* ``` *)
(* *)
(* which is just an $\eta$-expansion of the Y combinator *)
(* *)
(* ```ocaml *)
(* Y = λf. (λx. f (x x)) (λx. f (x x)) *)
(* ``` *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Fixed point: Z combinator *)
(* *)
(* \\[ *)
(* \begin{array}{rl} *)
(* & Z ~F = (λ\yb{f}.(λx.f ~(λy.x ~x ~y)) ~(λx.f ~(λy.x ~x ~y))) ~\yb{F} \\ *)
(* \rightarrow_{\beta V} & (λ\yb{x}.F ~(λy.x ~x ~y)) ~\yb{(λx.F ~(λy.x ~x ~y))} \\ *)
(* \rightarrow_{\beta V} & F ~(λy.\yb{(λx.F ~(λy.x ~x ~y)) ~(λx.F ~(λy.x ~x ~y))} ~y) \\ *)
(* \rightarrow_{\beta V} & F ~(λy. (Z ~F) ~y) *)
(* \end{array} *)
(* \\] *)
(* *)
(* The $\eta$-expansion has prevented further reduction. *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Recursive data structures: Lists *)
(* *)
(* * We have earlier seen simple data strcuture `pair` encoded in lambda calculus. *)
(* * Recursive data structures can also be encoded. *)
(* + List, Trees, etc. *)
(* * Mogensen–Scott encoding *)
(* + Take constructors as arguments *)
(* + slideshow={"slide_type": "fragment"}
let nil = p "\\c.\\n.n"
let cons = p "\\h.\\t.\\c.\\n.c h t"
(* + slideshow={"slide_type": "fragment"}
let l2 = app [cons; two; app [cons; one; nil]] (* [2;1] *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Lists: Head and Tail *)
(* *)
(* Empty list (`nil`) is `λc.λn.n` and non-empty list is `λc.λn.c h t`. *)
(* + slideshow={"slide_type": "-"}
let hd = lam "l" (app [var "l"; tru; nil])
let tl = lam "l" (app [var "l"; fls; nil])
(* + slideshow={"slide_type": "fragment"}
eval ~log:false @@ app [hd; l2];;
eval ~log:false @@ app [tl; l2];;
eval ~log:false @@ app [hd; nil];;
eval ~log:false @@ app [tl; nil]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Lists: is_nil? *)
(* *)
(* Empty list (`nil`) is `λc.λn.n` and non-empty list is `λc.λn.c h t`. *)
(* *)
(* ```ocaml *)
(* is_nil = λl.l (λx.λy.fls) tru *)
(* ``` *)
(* *)
(* Similar idea to `is_zero` function *)
(* *)
(* ```ocaml *)
(* is_zero = λn.n (λy.fls) tru *)
(* ``` *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Lists: is_nil? *)
(* -
let is_nil = lam "l" (app [var "l"; lam "x" (lam "y" fls); tru])
(* + slideshow={"slide_type": "fragment"}
eval ~log:false @@ app [is_nil; l2];;
eval ~log:false @@ app [is_nil; nil]
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Lists: Length *)
(* *)
(* Empty list (`nil`) is `λc.λn.n` and non-empty list is `λc.λn.c h t`. *)
(* *)
(* ```ocaml *)
(* length = λf.λl.if is_nil l then zero else succ (f (tl l)) *)
(* ``` *)
(* *)
(* where `f` is the recursive function. *)
(* + slideshow={"slide_type": "fragment"}
let len =
let cond = app[is_nil; var "l"] in
let flsc = app[scc; app[var "f"; app [tl; var "l"]]] in
app [y; lam "f" (lam "l" (app [cond; zero; flsc]))]
(* + slideshow={"slide_type": "slide"}
eval ~log:false @@ app [len; nil];;
eval ~log:false @@ app [len; l2];;
(* + [markdown] slideshow={"slide_type": "slide"}
(* ## Discussion *)
(* *)
(* * Lambda calculus is Turing-complete *)
(* + Most powerful language possible *)
(* + Can represent pretty much anything in a "real" language *)
(* * Using clever encodings *)
(* * But programs would be *)
(* + Pretty slow (10000 + 1 → thousands of function calls) *)
(* + Pretty large (10000 + 1 → hundreds of lines of code) *)
(* + Pretty hard to understand (recognize 10000 vs. 9999) *)
(* * In practice, *)
(* + We use richer, more expressive languages *)
(* + That include built-in primitives *)
(* + [markdown] slideshow={"slide_type": "slide"}
(* <center> *)
(* *)
(* <h1 style="text-align:center"> Fin. </h1> *)
(* </center> *)
| lectures/lec10/lec10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Ou0PGp_4icRo"
# # Time windows - 2 methods
# + id="gqWabzlJ63nL"
import numpy as np
import tensorflow as tf
from tensorflow.keras.preprocessing.sequence import TimeseriesGenerator
# + id="1tl-0BOKkEtk"
def window_dataset(series, time_steps, batch_size=32,
shuffle_buffer=1000):
dataset = tf.data.Dataset.from_tensor_slices(series)
dataset = dataset.window(time_steps + 1, shift=1, drop_remainder=True)
dataset = dataset.flat_map(lambda window: window.batch(time_steps + 1))
#dataset = dataset.shuffle(shuffle_buffer)
dataset = dataset.map(lambda window: (window[:-1], window[-1]))
dataset = dataset.batch(batch_size).prefetch(1)
return dataset
# -
x = np.arange(10)
print(x)
time_steps = 4
batch_size = 1
train_set = window_dataset(x, time_steps, batch_size)
for x, y in train_set:
print("x :\n", x.numpy())
print("y :\n", y.numpy())
# +
# define dataset
series = np.array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10])
series = np.arange(10)
time_steps = 4 # steps here
generator = TimeseriesGenerator(series, series, length=time_steps, batch_size=1)
# number of samples
print('Samples: %d' % len(generator))
for i in range(len(generator)):
x, y = generator[i]
print('%s => %s' % (x, y))
| 4. Time Series/AZ/Data Sampling/01_sample_window-01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="v-njGXZoIu7l" papermill={"duration": 0.009365, "end_time": "2021-05-12T18:31:16.661237", "exception": false, "start_time": "2021-05-12T18:31:16.651872", "status": "completed"} tags=[]
# <h1> HubMap - Hacking the Kidney </h1>
# <h3> Goal - Mapping the human body at function tissue unit level - detect glomeuli FTUs in kidney </h3>
#
# Implementation of Kaggle Notebook - Accuracy Prize Winner - WhatsGoinOn <br>
# Description - Use 2-5 fold models to predict on test image masks <br>
# Input - models, test images, sample_submission.csv <br>
# Output - submission_whatsgoinon.csv (rle for test images) <br>
#
# <b>How to use?</b><br>
# Change the basepath to where your data lives and you're good to go. <br>
#
# <h6> Step 1 - Import useful libraries </h6>
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" id="pHEON1kvIu7r" papermill={"duration": 3.08492, "end_time": "2021-05-12T18:31:19.754675", "exception": false, "start_time": "2021-05-12T18:31:16.669755", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from PIL import Image
import tifffile as tiff
import cv2
import os
import gc
from tqdm.notebook import tqdm
import rasterio
from rasterio.windows import Window
from fastai.vision.all import *
from torch.utils.data import Dataset, DataLoader
import tensorflow as tf
import warnings
warnings.filterwarnings("ignore")
# + [markdown] id="PsC7s0NBIu7s" papermill={"duration": 0.008339, "end_time": "2021-05-12T18:31:19.811650", "exception": false, "start_time": "2021-05-12T18:31:19.803311", "status": "completed"} tags=[]
# <h6> Step 2 - Helper functions </h6>
# + _kg_hide-input=true id="bjjwgkUnIu7s" papermill={"duration": 0.026863, "end_time": "2021-05-12T18:31:19.847047", "exception": false, "start_time": "2021-05-12T18:31:19.820184", "status": "completed"} tags=[]
#functions to convert encoding to mask and mask to encoding
def enc2mask(encs, shape):
img = np.zeros(shape[0]*shape[1], dtype=np.uint8)
for m,enc in enumerate(encs):
if isinstance(enc,np.float) and np.isnan(enc): continue
s = enc.split()
for i in range(len(s)//2):
start = int(s[2*i]) - 1
length = int(s[2*i+1])
img[start:start+length] = 1 + m
return img.reshape(shape).T
def mask2enc(mask, n=1):
pixels = mask.T.flatten()
encs = []
for i in range(1,n+1):
p = (pixels == i).astype(np.int8)
if p.sum() == 0: encs.append(np.nan)
else:
p = np.concatenate([[0], p, [0]])
runs = np.where(p[1:] != p[:-1])[0] + 1
runs[1::2] -= runs[::2]
encs.append(' '.join(str(x) for x in runs))
return encs
#https://www.kaggle.com/bguberfain/memory-aware-rle-encoding
#with transposed mask
def rle_encode_less_memory(img):
#the image should be transposed
pixels = img.T.flatten()
# This simplified method requires first and last pixel to be zero
pixels[0] = 0
pixels[-1] = 0
runs = np.where(pixels[1:] != pixels[:-1])[0] + 2
runs[1::2] -= runs[::2]
return ' '.join(str(x) for x in runs)
# + id="TR94UeLoIu7t" papermill={"duration": 0.041336, "end_time": "2021-05-12T18:31:19.896992", "exception": false, "start_time": "2021-05-12T18:31:19.855656", "status": "completed"} tags=[]
sz = 2048 #the size of tiles
sz_reduction = 2 #reduce the original images by 4 times
expansion = 512
TH = 0.5 #threshold for positive predictions
# https://www.kaggle.com/iafoss/256x256-images
mean = np.array([0.65459856,0.48386562,0.69428385])
std = np.array([0.15167958,0.23584107,0.13146145])
s_th = 40 #saturation blancking threshold
p_th = 1000*(sz//256)**2 #threshold for the minimum number of pixels
identity = rasterio.Affine(1, 0, 0, 0, 1, 0)
def img2tensor(img,dtype:np.dtype=np.float32):
if img.ndim==2 : img = np.expand_dims(img,2)
img = np.transpose(img,(2,0,1))
return torch.from_numpy(img.astype(dtype, copy=False))
class HuBMAPDataset(Dataset):
def __init__(self, idx, sz=sz, sz_reduction=sz_reduction, expansion=expansion):
self.data = rasterio.open(os.path.join(test,idx+'.tiff'), transform = identity,
num_threads='all_cpus')
# some images have issues with their format
# and must be saved correctly before reading with rasterio
if self.data.count != 3:
subdatasets = self.data.subdatasets
self.layers = []
if len(subdatasets) > 0:
for i, subdataset in enumerate(subdatasets, 0):
self.layers.append(rasterio.open(subdataset))
self.shape = self.data.shape
self.sz_reduction = sz_reduction
self.sz = sz_reduction*sz
self.expansion = sz_reduction*expansion
self.pad0 = (self.sz - self.shape[0]%self.sz)%self.sz
self.pad1 = (self.sz - self.shape[1]%self.sz)%self.sz
self.n0max = (self.shape[0] + self.pad0)//self.sz
self.n1max = (self.shape[1] + self.pad1)//self.sz
def __len__(self):
return self.n0max*self.n1max
def __getitem__(self, idx):
# the code below may be a little bit difficult to understand,
# but the thing it does is mapping the original image to
# tiles created with adding padding, as done in
# https://www.kaggle.com/iafoss/256x256-images ,
# and then the tiles are loaded with rasterio
# n0,n1 - are the x and y index of the tile (idx = n0*self.n1max + n1)
n0,n1 = idx//self.n1max, idx%self.n1max
# x0,y0 - are the coordinates of the lower left corner of the tile in the image
# negative numbers correspond to padding (which must not be loaded)
x0,y0 = -self.pad0//2 + n0*self.sz - self.expansion//2, -self.pad1//2 + n1*self.sz- self.expansion//2
# make sure that the region to read is within the image
p00,p01 = max(0,x0), min(x0+self.sz+self.expansion,self.shape[0])
p10,p11 = max(0,y0), min(y0+self.sz+self.expansion,self.shape[1])
img = np.zeros((self.sz+self.expansion,self.sz+self.expansion,3),np.uint8)
# mapping the loade region to the tile
if self.data.count == 3:
img[(p00-x0):(p01-x0),(p10-y0):(p11-y0)] = np.moveaxis(self.data.read([1,2,3],
window=Window.from_slices((p00,p01),(p10,p11))), 0, -1)
else:
for i,layer in enumerate(self.layers):
img[(p00-x0):(p01-x0),(p10-y0):(p11-y0),i] =\
layer.read(1,window=Window.from_slices((p00,p01),(p10,p11)))
if self.sz_reduction != 1:
img = cv2.resize(img,((self.sz+self.expansion)//self.sz_reduction,(self.sz+self.expansion)//self.sz_reduction),
interpolation = cv2.INTER_AREA)
#check for empty imges
hsv = cv2.cvtColor(img, cv2.COLOR_BGR2HSV)
h,s,v = cv2.split(hsv)
if (s>s_th).sum() <= p_th or img.sum() <= p_th:
#images with -1 will be skipped
return img2tensor((img/255.0 - mean)/std), -1
else: return img2tensor((img/255.0 - mean)/std), idx
# + id="oza-gjKAIu7t" papermill={"duration": 0.027498, "end_time": "2021-05-12T18:31:19.933229", "exception": false, "start_time": "2021-05-12T18:31:19.905731", "status": "completed"} tags=[]
#iterator like wrapper that returns predicted masks
class Model_pred:
def __init__(self, models, dl, tta:bool=True, half:bool=False):
self.models = models
self.dl = dl
self.tta = tta
self.half = half
def __iter__(self):
count=0
with torch.no_grad():
for x,y in iter(self.dl):
if ((y>=0).sum() > 0): #exclude empty images
x = x[y>=0].to(device)
y = y[y>=0]
if self.half: x = x.half()
py = None
for model in self.models:
p = model(x)
p = torch.sigmoid(p).detach()
if py is None: py = p
else: py += p
if self.tta:
#x,y,xy flips as TTA
flips = [[-1],[-2],[-2,-1]]
for f in flips:
xf = torch.flip(x,f)
for model in self.models:
p = model(xf)
p = torch.flip(p,f)
py += torch.sigmoid(p).detach()
py /= (1+len(flips))
py /= len(self.models)
py = F.upsample(py, scale_factor=sz_reduction, mode="bilinear")
py = py.permute(0,2,3,1).float().cpu()
batch_size = len(py)
for i in range(batch_size):
yield py[i,expansion*sz_reduction//2:-expansion*sz_reduction//2,expansion*sz_reduction//2:-expansion*sz_reduction//2],y[i]
count += 1
def __len__(self):
return len(self.dl.dataset)
# + _kg_hide-input=true id="gpVGG-_5Iu7u" papermill={"duration": 0.061964, "end_time": "2021-05-12T18:31:20.021808", "exception": false, "start_time": "2021-05-12T18:31:19.959844", "status": "completed"} tags=[]
import torch.nn as nn
# import torch.tensor as Tensor
import torch
from fastai.vision.all import *
from torchvision.models.resnet import ResNet, Bottleneck
class FPN(nn.Module):
def __init__(self, input_channels:list, output_channels:list):
super().__init__()
self.convs = nn.ModuleList(
[nn.Sequential(nn.Conv2d(in_ch, out_ch*2, kernel_size=3, padding=1),
nn.ReLU(inplace=True), nn.BatchNorm2d(out_ch*2),
nn.Conv2d(out_ch*2, out_ch, kernel_size=3, padding=1))
for in_ch, out_ch in zip(input_channels, output_channels)])
def forward(self, xs:list, last_layer):
hcs = [F.interpolate(c(x),scale_factor=2**(len(self.convs)-i),mode='bilinear')
for i,(c,x) in enumerate(zip(self.convs, xs))]
hcs.append(last_layer)
return torch.cat(hcs, dim=1)
class UnetBlock(nn.Module):
def __init__(self, up_in_c:int, x_in_c:int, nf:int=None, blur:bool=False,
self_attention:bool=False, **kwargs):
super().__init__()
self.shuf = PixelShuffle_ICNR(up_in_c, up_in_c//2, blur=blur, **kwargs)
self.bn = nn.BatchNorm2d(x_in_c)
ni = up_in_c//2 + x_in_c
nf = nf if nf is not None else max(up_in_c//2,32)
self.conv1 = ConvLayer(ni, nf, norm_type=None, **kwargs)
self.conv2 = ConvLayer(nf, nf, norm_type=None,
xtra=SelfAttention(nf) if self_attention else None, **kwargs)
self.relu = nn.ReLU(inplace=True)
def forward(self, up_in:Tensor, left_in:Tensor) -> Tensor:
s = left_in
up_out = self.shuf(up_in)
cat_x = self.relu(torch.cat([up_out, self.bn(s)], dim=1))
return self.conv2(self.conv1(cat_x))
class _ASPPModule(nn.Module):
def __init__(self, inplanes, planes, kernel_size, padding, dilation, groups=1):
super().__init__()
self.atrous_conv = nn.Conv2d(inplanes, planes, kernel_size=kernel_size,
stride=1, padding=padding, dilation=dilation, bias=False, groups=groups)
self.bn = nn.BatchNorm2d(planes)
self.relu = nn.ReLU()
self._init_weight()
def forward(self, x):
x = self.atrous_conv(x)
x = self.bn(x)
return self.relu(x)
def _init_weight(self):
for m in self.modules():
if isinstance(m, nn.Conv2d):
torch.nn.init.kaiming_normal_(m.weight)
elif isinstance(m, nn.BatchNorm2d):
m.weight.data.fill_(1)
m.bias.data.zero_()
class ASPP(nn.Module):
def __init__(self, inplanes=512, mid_c=256, dilations=[6, 12, 18, 24], out_c=None):
super().__init__()
self.aspps = [_ASPPModule(inplanes, mid_c, 1, padding=0, dilation=1)] + \
[_ASPPModule(inplanes, mid_c, 3, padding=d, dilation=d,groups=4) for d in dilations]
self.aspps = nn.ModuleList(self.aspps)
self.global_pool = nn.Sequential(nn.AdaptiveMaxPool2d((1, 1)),
nn.Conv2d(inplanes, mid_c, 1, stride=1, bias=False),
nn.BatchNorm2d(mid_c), nn.ReLU())
out_c = out_c if out_c is not None else mid_c
self.out_conv = nn.Sequential(nn.Conv2d(mid_c*(2+len(dilations)), out_c, 1, bias=False),
nn.BatchNorm2d(out_c), nn.ReLU(inplace=True))
self.conv1 = nn.Conv2d(mid_c*(2+len(dilations)), out_c, 1, bias=False)
self._init_weight()
def forward(self, x):
x0 = self.global_pool(x)
xs = [aspp(x) for aspp in self.aspps]
x0 = F.interpolate(x0, size=xs[0].size()[2:], mode='bilinear', align_corners=True)
x = torch.cat([x0] + xs, dim=1)
return self.out_conv(x)
def _init_weight(self):
for m in self.modules():
if isinstance(m, nn.Conv2d):
torch.nn.init.kaiming_normal_(m.weight)
elif isinstance(m, nn.BatchNorm2d):
m.weight.data.fill_(1)
m.bias.data.zero_()
import torch.hub
# hub_model = torch.hub.load(
# 'moskomule/senet.pytorch',
# 'se_resnet50',
# pretrained=True,)
import torchvision
class UneXt(nn.Module):
def __init__(self, m, stride=1, **kwargs):
super().__init__()
#encoder
# m = torch.hub.load('facebookresearch/semi-supervised-ImageNet1K-models',
# 'resnext101_32x4d_swsl')
# m = torch.hub.load('facebookresearch/semi-supervised-ImageNet1K-models',
# 'resnext50_32x4d_swsl', pretrained=False)
#m = ResNet(Bottleneck, [3, 4, 23, 3], groups=32, width_per_group=4)
#m = torchvision.models.resnext50_32x4d(pretrained=False)
# m = torch.hub.load(
# 'moskomule/senet.pytorch',
# 'se_resnet101',
# pretrained=True,)
#m=torch.hub.load('zhanghang1989/ResNeSt', 'resnest50', pretrained=True)
self.enc0 = nn.Sequential(m.conv1, m.bn1, nn.ReLU(inplace=True))
self.enc1 = nn.Sequential(nn.MaxPool2d(kernel_size=3, stride=2, padding=1, dilation=1),
m.layer1) #256
self.enc2 = m.layer2 #512
self.enc3 = m.layer3 #1024
self.enc4 = m.layer4 #2048
#aspp with customized dilatations
self.aspp = ASPP(2048,256,out_c=512,dilations=[stride*1,stride*2,stride*3,stride*4])
self.drop_aspp = nn.Dropout2d(0.5)
#decoder
self.dec4 = UnetBlock(512,1024,256)
self.dec3 = UnetBlock(256,512,128)
self.dec2 = UnetBlock(128,256,64)
self.dec1 = UnetBlock(64,64,32)
self.fpn = FPN([512,256,128,64],[16]*4)
self.drop = nn.Dropout2d(0.1)
self.final_conv = ConvLayer(32+16*4, 1, ks=1, norm_type=None, act_cls=None)
def forward(self, x):
enc0 = self.enc0(x)
enc1 = self.enc1(enc0)
enc2 = self.enc2(enc1)
enc3 = self.enc3(enc2)
enc4 = self.enc4(enc3)
enc5 = self.aspp(enc4)
dec3 = self.dec4(self.drop_aspp(enc5),enc3)
dec2 = self.dec3(dec3,enc2)
dec1 = self.dec2(dec2,enc1)
dec0 = self.dec1(dec1,enc0)
x = self.fpn([enc5, dec3, dec2, dec1], dec0)
x = self.final_conv(self.drop(x))
x = F.interpolate(x,scale_factor=2,mode='bilinear')
return x
# + [markdown] id="tcJ0WYIuIu7x" papermill={"duration": 0.008819, "end_time": "2021-05-12T18:32:04.820874", "exception": false, "start_time": "2021-05-12T18:32:04.812055", "status": "completed"} tags=[]
# <h6> Step 3 - Set configuration and paths </h6>
# + id="WfY_KuBPIu7s" papermill={"duration": 0.029592, "end_time": "2021-05-12T18:31:19.794350", "exception": false, "start_time": "2021-05-12T18:31:19.764758", "status": "completed"} tags=[]
DATA = Path(r'C:\Users\soodn\Downloads\Naveksha\Kaggle HuBMAP\Data\kidney-data')
MODELS_rsxt50 = [f'model_OG/hubmaptest44/fold{i}.pth' for i in range(5)]
MODELS_rsxt101 = [f'model_OG/hubmaptest45/fold{i}.pth' for i in range(5)]
test = DATA/'private test'
df_sample = pd.read_csv(DATA/'sample_submission_pvt.csv')
df_test = pd.read_csv(DATA/'private_test.csv')
bs = 2
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# + id="4NmLCbtVIu7w" papermill={"duration": 44.771748, "end_time": "2021-05-12T18:32:04.802749", "exception": false, "start_time": "2021-05-12T18:31:20.031001", "status": "completed"} tags=[]
import gc
models = []
for path in MODELS_rsxt50:
state_dict = torch.load(path)
model = UneXt(m=torchvision.models.resnext50_32x4d(pretrained=False)).cuda()
model = nn.DataParallel(model)
model.load_state_dict(state_dict)
model.float()
model.eval()
#model.to(device)
models.append(model)
for path in MODELS_rsxt101:
state_dict = torch.load(path)
model = UneXt(m=ResNet(Bottleneck, [3, 4, 23, 3], groups=32, width_per_group=4)).cuda()
model = nn.DataParallel(model)
model.load_state_dict(state_dict)
model.float()
model.eval()
#model.to(device)
models.append(model)
del state_dict
# -
# <h6> Step 4 - Make Prediction </h6>
# + _kg_hide-output=true colab={"referenced_widgets": ["112d789b9c824b0786d2a71bc3e4f38f"]} id="exgfYD2aIu7y" outputId="ac17da08-7794-4fbe-86f3-b2b308331460" papermill={"duration": 6228.498649, "end_time": "2021-05-12T20:15:53.328441", "exception": false, "start_time": "2021-05-12T18:32:04.829792", "status": "completed"} tags=[]
names,preds = [],[]
for idx,row in tqdm(df_sample.iterrows(),total=len(df_sample)):
idx = row['id']
ds = HuBMAPDataset(idx)
#rasterio cannot be used with multiple workers
dl = DataLoader(ds,bs,num_workers=0,shuffle=False,pin_memory=True)
mp = Model_pred(models,dl)
#generate masks
mask = torch.zeros(len(ds),ds.sz,ds.sz,dtype=torch.int8)
for p,i in iter(mp):
# print(p.shape)
mask[i.item()] = p.squeeze(-1) > TH
#reshape tiled masks into a single mask and crop padding
mask = mask.view(ds.n0max,ds.n1max,ds.sz,ds.sz).\
permute(0,2,1,3).reshape(ds.n0max*ds.sz,ds.n1max*ds.sz)
mask = mask[ds.pad0//2:-(ds.pad0-ds.pad0//2) if ds.pad0 > 0 else ds.n0max*ds.sz,
ds.pad1//2:-(ds.pad1-ds.pad1//2) if ds.pad1 > 0 else ds.n1max*ds.sz]
rle = rle_encode_less_memory(mask.numpy())
names.append(idx)
preds.append(rle)
del mask, ds, dl
gc.collect()
# + id="Vs-ofmkEIu7y" papermill={"duration": 0.513902, "end_time": "2021-05-12T20:15:53.901034", "exception": false, "start_time": "2021-05-12T20:15:53.387132", "status": "completed"} tags=[]
df = pd.DataFrame({'id':names,'predicted':preds})
df.to_csv('submission-wgo.csv',index=False)
| models/3-Whats goin on/WhatsGoinOn-Inference-Kidney.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ZU90KHE5jwQp"
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import zipfile
from PIL import Image,ImageOps
# + id="fg9ksUowsfmX"
zip = zipfile.ZipFile('Train.zip')
zip.extractall()
# + colab={"base_uri": "https://localhost:8080/"} id="b-gK4upEj_mw" outputId="02b676fc-66e3-42ed-afad-a7b3f427c860"
import os
cur_path = os.getcwd()
print(cur_path)
# + id="OtvJvHHvlBaH"
classs={
1:'Negative',
-1:'Positive'
}
# + id="COZTBV0HEZ8b"
data=[]
labels=[]
# + id="ru1nrhinkDNW"
classes = 2
for i in range(classes):
path = os.path.join(cur_path,'Train/',str(i))
images = os.listdir(path)
#print(images)
for a in images:
image = Image.open(path + '//'+ a)
image=ImageOps.grayscale(image)
image = image.resize((64,64))
image = np.array(image)
image=image.flatten()
data.append(image)
labels.append(i)
print("{0} Loaded".format(a))
# + colab={"base_uri": "https://localhost:8080/"} id="8CS_CGmHGUf9" outputId="a93099c6-0668-4922-88ee-823935418341"
labels
# + colab={"base_uri": "https://localhost:8080/"} id="Facn54WHGaJC" outputId="01cd3fe3-ed01-48ba-c340-8beb4937f8b8"
len(labels)
# + colab={"base_uri": "https://localhost:8080/"} id="a13erm6GG0tI" outputId="7f7008e8-7ec5-42df-de8e-2fabe933bbef"
labels[20001]
# + id="bUzg73pHGcUG"
for i in range(len(labels)):
if labels[i]==int('0'):
labels[i]=int('-1')
else:
break
# + colab={"base_uri": "https://localhost:8080/"} id="A3srX8m1GrRm" outputId="02a464f3-994f-4860-88dd-40c6ac6bf7c4"
labels
# + id="uLtpwxS1kbFg"
#Converting lists into numpy arrays
data = np.array(data)
labels = np.array(labels)
# + colab={"base_uri": "https://localhost:8080/"} id="qqd_PDKpHIGI" outputId="4a2891bf-1373-45d5-8948-805f15cc9a98"
len(data)
# + id="FaHW24oilWQ8"
from sklearn.model_selection import train_test_split
# + id="w5t3VFS5lZQL"
X_train, X_test, y_train, y_test = train_test_split(data, labels, test_size=0.25, random_state=42)
# + colab={"base_uri": "https://localhost:8080/"} id="ZXc7earvlbbJ" outputId="3b3c1071-4cde-4a99-c774-e583e5bb3518"
from sklearn.ensemble import IsolationForest
clf = IsolationForest(max_samples=100000, random_state=42)
clf.fit(X_train,y_train)
# + id="rZc4vOEnlv8F"
y_pred = clf.predict(X_test)
y_pred=np.round(y_pred)
np.set_printoptions(precision=2)
# + colab={"base_uri": "https://localhost:8080/"} id="QviCk0oLmAq1" outputId="76d6c512-74d0-44c4-a4cf-b5320fd0ada4"
print(np.concatenate((y_pred.reshape(len(y_pred),1), y_test.reshape(len(y_test),1)),1))
# + colab={"base_uri": "https://localhost:8080/"} id="YzGXl2d1mBNa" outputId="95128dde-f90d-4fea-9215-ddf5225959cf"
from sklearn.metrics import accuracy_score
print("Accuracy Score for the algorithm=>{}%".format(round(accuracy_score(y_test,y_pred)*100),2))
| part-1/SD_IF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mopala/ML-DL/blob/main/Mnist/cnn_classifier.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" colab={"base_uri": "https://localhost:8080/"} id="fl8UX2uYSf0K" outputId="9e7b2401-db86-48ea-bf68-54e2b4637693"
import tensorflow as tf
import numpy as np
from tensorflow import keras
from keras import layers
print(tf.__version__)
# + _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" id="j87zn0zVSf0L"
mnist=tf.keras.datasets.mnist
(X_train,Y_train),(X_test,Y_test)=mnist.load_data()
# + id="8ITUqqJQSf0M"
x_train=np.array(X_train)
x_train=x_train.reshape([-1,28,28,1])
x_train=x_train/255
x_test=np.array(X_test)
x_test=x_test/255
x_test=x_test.reshape([-1,28,28,1])
# + colab={"base_uri": "https://localhost:8080/"} id="TzHQh5niSf0N" outputId="38b1e2d3-5e44-4015-f3f9-59cf6c3b899f"
model=keras.Sequential()
model.add(keras.Input(shape=(28,28,1)))
model.add(layers.Conv2D(16,(7,7)))
model.add(layers.BatchNormalization(axis=3))
model.add(layers.Activation('relu'))
model.add(layers.Dropout(0.2))
model.add(layers.MaxPooling2D((2,2)))
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="u7ZB5cwdSf0N" outputId="15195343-45cd-478d-97cc-21a4c1a02435"
model.add(keras.Input(shape=(28,28,1)))
model.add(layers.Conv2D(128,(5,5)))
model.add(layers.BatchNormalization(axis=3))
model.add(layers.Activation('relu'))
model.add(layers.Dropout(0.2))
model.add(layers.MaxPooling2D((2,2)))
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="UmxKraPKSf0N" outputId="368ceece-4016-4721-8de0-8794d319f99d"
model.add(layers.Flatten())
model.add(layers.Dense(10,activation='softmax'))
model.summary()
# + id="ys-wnqHDSf0N"
model.compile(optimizer='Adam',loss='categorical_crossentropy',metrics='accuracy')
# + id="bFP0KGTHSf0N"
y_train=tf.keras.utils.to_categorical(Y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="1W_OoSyuSf0N" outputId="b3c5dc6c-dd0f-443e-a578-1f2669a6eeff"
model.fit(x_train,y_train,batch_size=32,epochs=10)
# + colab={"base_uri": "https://localhost:8080/"} id="BC3ek3aaSf0O" outputId="61dda48c-0daa-443f-cd3b-79f22c5b882f"
Y_pred=model.predict_classes(x_test)
print("The accuracy on test set is ",np.mean(Y_pred==Y_test))
# + id="6XyKdgGOSf0P"
| Mnist/cnn_classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="oYM61xrTsP5d"
# # EfficientNetV2 with tf-hub
#
# + [markdown] id="MfBg1C5NB3X0"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://github.com/google/automl/blob/master/efficientnetv2/tfhub.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on github
# </a>
# </td><td>
# <a target="_blank" href="https://colab.research.google.com/github/google/automl/blob/master/efficientnetv2/tfhub.ipynb">
# <img width=32px src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td><td>
# <!----<a href="https://tfhub.dev/google/collections/image/1"><img src="https://www.tensorflow.org/images/hub_logo_32px.png" />TF Hub models</a>--->
# </td>
# </table>
# + [markdown] id="L1otmJgmbahf"
# ## 1.Introduction
#
# [EfficientNetV2](https://arxiv.org/abs/2104.00298) is a family of classification models, with better accuracy, smaller size, and faster speed than previous models.
#
#
# This doc describes some examples with EfficientNetV2 tfhub. For more details, please visit the official code: https://github.com/google/automl/tree/master/efficientnetv2
# + [markdown] id="mmaHHH7Pvmth"
# ## 2.Select the TF2 SavedModel module to use
# + id="FlsEcKVeuCnf" colab={"base_uri": "https://localhost:8080/"} outputId="a81e6d95-2d0c-4625-e25e-e62cbd753558"
import itertools
import os
import matplotlib.pylab as plt
import numpy as np
import tensorflow as tf
import tensorflow_hub as hub
print('TF version:', tf.__version__)
print('Hub version:', hub.__version__)
print('Phsical devices:', tf.config.list_physical_devices())
def get_hub_url_and_isize(model_name, ckpt_type, hub_type):
if ckpt_type == '-1k':
ckpt_type = '' # json doesn't support empty string
hub_url_map = {
'efficientnetv2-b0': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-b0/{hub_type}',
'efficientnetv2-b1': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-b1/{hub_type}',
'efficientnetv2-b2': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-b2/{hub_type}',
'efficientnetv2-b3': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-b3/{hub_type}',
'efficientnetv2-s': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-s/{hub_type}',
'efficientnetv2-m': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-m/{hub_type}',
'efficientnetv2-l': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-l/{hub_type}',
'efficientnetv2-s-21k': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-s-21k/{hub_type}',
'efficientnetv2-m-21k': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-m-21k/{hub_type}',
'efficientnetv2-l-21k': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-l-21k/{hub_type}',
'efficientnetv2-s-21k-ft1k': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-s-21k-ft1k/{hub_type}',
'efficientnetv2-m-21k-ft1k': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-m-21k-ft1k/{hub_type}',
'efficientnetv2-l-21k-ft1k': f'gs://cloud-tpu-checkpoints/efficientnet/v2/hub/efficientnetv2-l-21k-ft1k/{hub_type}',
# efficientnetv1
'efficientnet_b0': f'https://tfhub.dev/tensorflow/efficientnet/b0/{hub_type}/1',
'efficientnet_b1': f'https://tfhub.dev/tensorflow/efficientnet/b1/{hub_type}/1',
'efficientnet_b2': f'https://tfhub.dev/tensorflow/efficientnet/b2/{hub_type}/1',
'efficientnet_b3': f'https://tfhub.dev/tensorflow/efficientnet/b3/{hub_type}/1',
'efficientnet_b4': f'https://tfhub.dev/tensorflow/efficientnet/b4/{hub_type}/1',
'efficientnet_b5': f'https://tfhub.dev/tensorflow/efficientnet/b5/{hub_type}/1',
'efficientnet_b6': f'https://tfhub.dev/tensorflow/efficientnet/b6/{hub_type}/1',
'efficientnet_b7': f'https://tfhub.dev/tensorflow/efficientnet/b7/{hub_type}/1',
}
image_size_map = {
'efficientnetv2-b0': 224,
'efficientnetv2-b1': 240,
'efficientnetv2-b2': 260,
'efficientnetv2-b3': 300,
'efficientnetv2-s': 384,
'efficientnetv2-m': 480,
'efficientnetv2-l': 480,
'efficientnet_b0': 224,
'efficientnet_b1': 240,
'efficientnet_b2': 260,
'efficientnet_b3': 300,
'efficientnet_b4': 380,
'efficientnet_b5': 456,
'efficientnet_b6': 528,
'efficientnet_b7': 600,
}
hub_url = hub_url_map.get(model_name + ckpt_type)
image_size = image_size_map.get(model_name, 224)
return hub_url, image_size
# + [markdown] id="REiLDGq_W5FA"
# ## 3.Inference with Panda image
# + id="E32RGKBEWq76" colab={"base_uri": "https://localhost:8080/", "height": 799} outputId="1d635114-509e-4b52-86c0-befd5573b4c9"
# Build model
import tensorflow_hub as hub
model_name = 'efficientnetv2-s' #@param {type:'string'}
ckpt_type = '-21k-ft1k' # @param ['-21k', '-21k-ft1k', '-1k']
hub_type = 'classification' # @param ['classification', 'feature-vector']
hub_url, image_size = get_hub_url_and_isize(model_name, ckpt_type, hub_type)
tf.keras.backend.clear_session()
m = hub.KerasLayer(hub_url, trainable=False)
m.build([None, 224, 224, 3]) # Batch input shape.
# Download label map file and image
labels_map = '/tmp/labels_map.txt'
image_file = '/tmp/panda.jpg'
tf.keras.utils.get_file(image_file, 'https://upload.wikimedia.org/wikipedia/commons/f/fe/Giant_Panda_in_Beijing_Zoo_1.JPG')
tf.keras.utils.get_file(labels_map, 'https://storage.googleapis.com/cloud-tpu-checkpoints/efficientnet/eval_data/labels_map.txt')
# preprocess image.
image = tf.keras.preprocessing.image.load_img(image_file, target_size=(224, 224))
image = tf.keras.preprocessing.image.img_to_array(image)
image = (image - 128.) / 128.
logits = m(tf.expand_dims(image, 0), False)
# Output classes and probability
pred = tf.keras.layers.Softmax()(logits)
idx = tf.argsort(logits[0])[::-1][:5].numpy()
import ast
classes = ast.literal_eval(open(labels_map, "r").read())
for i, id in enumerate(idx):
print(f'top {i+1} ({pred[0][id]*100:.1f}%): {classes[id]} ')
from IPython import display
display.display(display.Image(image_file))
# + [markdown] id="yTY8qzyYv3vl"
# ## 4.Finetune with Flowers dataset.
# + [markdown] id="c_12xwDuZFOQ"
# Get hub_url and image_size
#
# + id="50FYNIb1dmJH"
# Build model
import tensorflow_hub as hub
model_name = 'efficientnetv2-b0' #@param {type:'string'}
ckpt_type = '-1k' # @param ['-21k', '-21k-ft1k', '-1k']
hub_type = 'feature-vector' # @param ['feature-vector']
batch_size = 32#@param {type:"integer"}
hub_url, image_size = get_hub_url_and_isize(model_name, ckpt_type, hub_type)
# + [markdown] id="lus9bIA-bQgj"
# Get dataset
# + id="umB5tswsfTEQ" colab={"base_uri": "https://localhost:8080/"} outputId="8df0490e-13cd-4de4-fa0a-a6551ecdb6d5"
data_dir = tf.keras.utils.get_file(
'flower_photos',
'https://storage.googleapis.com/download.tensorflow.org/example_images/flower_photos.tgz',
untar=True)
datagen_kwargs = dict(rescale=1./255, validation_split=.20)
dataflow_kwargs = dict(target_size=(image_size, image_size),
batch_size=batch_size,
interpolation="bilinear")
valid_datagen = tf.keras.preprocessing.image.ImageDataGenerator(
**datagen_kwargs)
valid_generator = valid_datagen.flow_from_directory(
data_dir, subset="validation", shuffle=False, **dataflow_kwargs)
do_data_augmentation = False #@param {type:"boolean"}
if do_data_augmentation:
train_datagen = tf.keras.preprocessing.image.ImageDataGenerator(
rotation_range=40,
horizontal_flip=True,
width_shift_range=0.2, height_shift_range=0.2,
shear_range=0.2, zoom_range=0.2,
**datagen_kwargs)
else:
train_datagen = valid_datagen
train_generator = train_datagen.flow_from_directory(
data_dir, subset="training", shuffle=True, **dataflow_kwargs)
# + [markdown] id="u2e5WupIw2N2"
# Training the model
# + id="9f3yBUvkd_VJ" colab={"base_uri": "https://localhost:8080/"} outputId="99b79db6-d9f9-438e-9011-8221e8c5377c"
# whether to finetune the whole model or just the top layer.
do_fine_tuning = True #@param {type:"boolean"}
num_epochs = 2 #@param {type:"integer"}
tf.keras.backend.clear_session()
model = tf.keras.Sequential([
# Explicitly define the input shape so the model can be properly
# loaded by the TFLiteConverter
tf.keras.layers.InputLayer(input_shape=[image_size, image_size, 3]),
hub.KerasLayer(hub_url, trainable=do_fine_tuning),
tf.keras.layers.Dropout(rate=0.2),
tf.keras.layers.Dense(train_generator.num_classes,
kernel_regularizer=tf.keras.regularizers.l2(0.0001))
])
model.build((None, image_size, image_size, 3))
model.summary()
model.compile(
optimizer=tf.keras.optimizers.SGD(learning_rate=0.005, momentum=0.9),
loss=tf.keras.losses.CategoricalCrossentropy(from_logits=True, label_smoothing=0.1),
metrics=['accuracy'])
steps_per_epoch = train_generator.samples // train_generator.batch_size
validation_steps = valid_generator.samples // valid_generator.batch_size
hist = model.fit(
train_generator,
epochs=num_epochs, steps_per_epoch=steps_per_epoch,
validation_data=valid_generator,
validation_steps=validation_steps).history
# + id="oi1iCNB9K1Ai" colab={"base_uri": "https://localhost:8080/", "height": 301} outputId="c8e2fc22-b4bd-4950-bce3-6c4af197f405"
def get_class_string_from_index(index):
for class_string, class_index in valid_generator.class_indices.items():
if class_index == index:
return class_string
x, y = next(valid_generator)
image = x[0, :, :, :]
true_index = np.argmax(y[0])
plt.imshow(image)
plt.axis('off')
plt.show()
# Expand the validation image to (1, 224, 224, 3) before predicting the label
prediction_scores = model.predict(np.expand_dims(image, axis=0))
predicted_index = np.argmax(prediction_scores)
print("True label: " + get_class_string_from_index(true_index))
print("Predicted label: " + get_class_string_from_index(predicted_index))
# + [markdown] id="YCsAsQM1IRvA"
# Finally, the trained model can be saved for deployment to TF Serving or TF Lite (on mobile) as follows.
# + id="LGvTi69oIc2d" colab={"base_uri": "https://localhost:8080/"} outputId="710b8803-3e34-4948-96e7-d905c5c96d64"
saved_model_path = f"/tmp/saved_flowers_model_{model_name}"
tf.saved_model.save(model, saved_model_path)
# + [markdown] id="QzW4oNRjILaq"
# ## Optional: Deployment to TensorFlow Lite
#
# [TensorFlow Lite](https://www.tensorflow.org/lite) for mobile. Here we also runs tflite file in the TF Lite Interpreter to examine the resulting quality.
# + id="Va1Vo92fSyV6" colab={"base_uri": "https://localhost:8080/"} outputId="11956d83-1b22-4330-f57a-bfc893ec2b2a"
optimize_lite_model = True #@param {type:"boolean"}
#@markdown Setting a value greater than zero enables quantization of neural network activations. A few dozen is already a useful amount.
num_calibration_examples = 81 #@param {type:"slider", min:0, max:1000, step:1}
representative_dataset = None
if optimize_lite_model and num_calibration_examples:
# Use a bounded number of training examples without labels for calibration.
# TFLiteConverter expects a list of input tensors, each with batch size 1.
representative_dataset = lambda: itertools.islice(
([image[None, ...]] for batch, _ in train_generator for image in batch),
num_calibration_examples)
converter = tf.lite.TFLiteConverter.from_saved_model(saved_model_path)
if optimize_lite_model:
converter.optimizations = [tf.lite.Optimize.DEFAULT]
if representative_dataset: # This is optional, see above.
converter.representative_dataset = representative_dataset
lite_model_content = converter.convert()
with open(f"/tmp/lite_flowers_model_{model_name}.tflite", "wb") as f:
f.write(lite_model_content)
print("Wrote %sTFLite model of %d bytes." %
("optimized " if optimize_lite_model else "", len(lite_model_content)))
# + id="_wqEmD0xIqeG"
interpreter = tf.lite.Interpreter(model_content=lite_model_content)
# This little helper wraps the TF Lite interpreter as a numpy-to-numpy function.
def lite_model(images):
interpreter.allocate_tensors()
interpreter.set_tensor(interpreter.get_input_details()[0]['index'], images)
interpreter.invoke()
return interpreter.get_tensor(interpreter.get_output_details()[0]['index'])
# + id="JMMK-fZrKrk8" colab={"base_uri": "https://localhost:8080/"} outputId="f5e205ee-42da-4041-b397-26408ad78b88"
#@markdown For rapid experimentation, start with a moderate number of examples.
num_eval_examples = 50 #@param {type:"slider", min:0, max:700}
eval_dataset = ((image, label) # TFLite expects batch size 1.
for batch in train_generator
for (image, label) in zip(*batch))
count = 0
count_lite_tf_agree = 0
count_lite_correct = 0
for image, label in eval_dataset:
probs_lite = lite_model(image[None, ...])[0]
probs_tf = model(image[None, ...]).numpy()[0]
y_lite = np.argmax(probs_lite)
y_tf = np.argmax(probs_tf)
y_true = np.argmax(label)
count +=1
if y_lite == y_tf: count_lite_tf_agree += 1
if y_lite == y_true: count_lite_correct += 1
if count >= num_eval_examples: break
print("TF Lite model agrees with original model on %d of %d examples (%g%%)." %
(count_lite_tf_agree, count, 100.0 * count_lite_tf_agree / count))
print("TF Lite model is accurate on %d of %d examples (%g%%)." %
(count_lite_correct, count, 100.0 * count_lite_correct / count))
| efficientnetv2/tfhub.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Dogs v Cats super-charged!
# Put these at the top of every notebook, to get automatic reloading and inline plotting
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# +
# This file contains all the main external libs we'll use
from fastai.imports import *
from fastai.transforms import *
from fastai.conv_learner import *
from fastai.model import *
from fastai.dataset import *
from fastai.sgdr import *
from fastai.plots import *
PATH = "data/dogscats/"
sz=299
arch=resnext50
bs=28
# -
tfms = tfms_from_model(arch, sz, aug_tfms=transforms_side_on, max_zoom=1.1)
data = ImageClassifierData.from_paths(PATH, tfms=tfms, bs=bs, num_workers=4)
learn = ConvLearner.pretrained(arch, data, precompute=True, ps=0.5)
learn.fit(1e-2, 1)
learn.precompute=False
learn.fit(1e-2, 2, cycle_len=1)
learn.unfreeze()
lr=np.array([1e-4,1e-3,1e-2])
learn.fit(lr, 3, cycle_len=1)
learn.save('224_all_50')
learn.load('224_all_50')
# +
log_preds,y = learn.TTA()
probs = np.mean(np.exp(log_preds),0)
accuracy(probs,y)
# -
# ## Analyzing results
# +
preds = np.argmax(probs, axis=1)
probs = probs[:,1]
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y, preds)
# -
plot_confusion_matrix(cm, data.classes)
# +
def rand_by_mask(mask): return np.random.choice(np.where(mask)[0], 4, replace=False)
def rand_by_correct(is_correct): return rand_by_mask((preds == data.val_y)==is_correct)
def plot_val_with_title(idxs, title):
imgs = np.stack([data.val_ds[x][0] for x in idxs])
title_probs = [probs[x] for x in idxs]
print(title)
return plots(data.val_ds.denorm(imgs), rows=1, titles=title_probs)
def plots(ims, figsize=(12,6), rows=1, titles=None):
f = plt.figure(figsize=figsize)
for i in range(len(ims)):
sp = f.add_subplot(rows, len(ims)//rows, i+1)
sp.axis('Off')
if titles is not None: sp.set_title(titles[i], fontsize=16)
plt.imshow(ims[i])
def load_img_id(ds, idx): return np.array(PIL.Image.open(PATH+ds.fnames[idx]))
def plot_val_with_title(idxs, title):
imgs = [load_img_id(data.val_ds,x) for x in idxs]
title_probs = [probs[x] for x in idxs]
print(title)
return plots(imgs, rows=1, titles=title_probs, figsize=(16,8))
def most_by_mask(mask, mult):
idxs = np.where(mask)[0]
return idxs[np.argsort(mult * probs[idxs])[:4]]
def most_by_correct(y, is_correct):
mult = -1 if (y==1)==is_correct else 1
return most_by_mask((preds == data.val_y)==is_correct & (data.val_y == y), mult)
# -
plot_val_with_title(most_by_correct(0, False), "Most incorrect cats")
plot_val_with_title(most_by_correct(1, False), "Most incorrect dogs")
| courses/dl1/lesson1-rxt50.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # PART FOUR
#
# Take a matplotlib bar chart from any data set we've worked with (including the scrabble or NBA data) and use Illustrator to make it look a little bit nicer. Whatever "nicer" might mean.
# +
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
import matplotlib
matplotlib.rcParams['pdf.fonttype']=42
# -
performance_df = pd.read_excel("NBA stats.xlsx", sheetname="Player performance")
salary_df = pd.read_excel("NBA stats.xlsx", sheetname="2016-2017 Salaries")
df = performance_df.merge(salary_df, left_on='Player', right_on='NAME')
df.head()
salary_df.groupby('TEAM')['SALARY'].mean().sort_values(ascending=False).head(5)
salary_df.groupby('TEAM')['SALARY'].mean().sort_values(ascending=False).head(5).plot(kind='barh')
plt.savefig("nba-team-salary.pdf", transparent=True)
| homework1/homework1_part4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] toc="true"
# # Table of Contents
# <div class="toc" style="margin-top: 1em;"><ul class="toc-item" id="toc-level0"></ul></div>
# +
### Figures for Metro Vancouver Preliminary Report 2016 ###
# +
import datetime
import matplotlib.pyplot as plt
import matplotlib.colors as mpl_colors
import matplotlib.colorbar as mpl_colorbar
import netCDF4 as nc
import numpy as np
from salishsea_tools import geo_tools
# %matplotlib inline
# -
#End of spit
late = 49.205
lone = -123.26
#Further up spit
lats = 49.214
lons = -123.22
bathy = nc.Dataset('../../NEMO-forcing/grid/bathy_meter_SalishSea2.nc')
model_lats = bathy.variables['nav_lat'][:]
model_lons = bathy.variables['nav_lon'][:]
depth = bathy.variables['Bathymetry'][:]
dx = 1.2
lat = late + dx * (late - lats)
lon = lone + dx * (lone - lons)
y, x = geo_tools.find_closest_model_point(lon, lat, model_lons, model_lats)
print(depth[y, x])
print(y, x)
print('And in Fortran for ariane', y + 1, x + 1)
# point I used
outy = y - 1
outx = x - 1
print('Point I used', outx, outy)
ugrid = nc.Dataset('../../../E471/miniproj3/eosc471/uVelocity_2016apr09_01.nc')
darray = ugrid.variables['depth'][:]
print (darray[27-1])
uvel = ugrid.variables['uVelocity'][:]
print (uvel.shape)
#traj = np.loadtxt('../../../E471/miniproj3/eosc471/traj_2016apr09.txt', delimiter=' ')
#traj = np.loadtxt('../../../E471/miniproj3/traj_2016apr17.txt', delimiter=' ')
traj = np.loadtxt('../../../E471/miniproj3/traj_2016oct14.txt', delimiter=' ')
print (traj[0])
print (traj.shape)
deep = 110
shallow = 35
# +
# Make a figure and axes with dimensions as desired.
fig = plt.figure(figsize=(7, 7))
ax1 = fig.add_axes([0.05, 0.00, 0.72, 0.05])
ax = fig.add_axes([0.05, 0.15, 0.9, 0.9])
mesh = ax.pcolormesh(model_lons, model_lats, depth, cmap='winter_r')
ax.set_xlim((-123.5, -123.1))
ax.set_ylim((49.05, 49.35))
for i in range(traj.shape[0]):
scaled_z = 1-(deep + traj[i, 3]) / (deep - shallow)
cmap = plt.cm.plasma_r
color = cmap(scaled_z)
ax.scatter(traj[i, 1], traj[i, 2], c=color)
norm = mpl_colors.Normalize(vmin=deep, vmax=shallow)
cb1 = mpl_colorbar.ColorbarBase(
ax1, cmap=cmap, norm=norm, orientation='horizontal')
cb1.set_label('Particle Depth (m)')
ax.plot(model_lons[outy, outx], model_lats[outy, outx], 'wo')
ax.set_yticks((49.1, 49.2, 49.3))
ax.set_yticklabels(('49.1 N', '49.2 N', '49.3 N'))
ax.set_ylabel('Latitude')
cb = fig.colorbar(mesh, ax=ax)
cb.set_label('Water Depth (m)')
ax.set_xticks((-123.5, -123.3, -123.1))
ax.set_xticklabels(('123.5 W', '123.3 W', '123.1 W'))
ax.set_xlabel('Longitude')
ax.set_title('October 14, 2016');
# -
tidegrid = nc.Dataset('miniproj3/PointAtkinson.nc')
ssh = tidegrid['ssh'][:,0,0]
print (ssh.shape)
plt.plot(ssh)
tidegrid = nc.Dataset('miniproj3/PointAtkinson_whole.nc')
ssh = tidegrid['ssh'][:,0,0]
times = tidegrid['time'][:]
print (ssh.shape)
plt.plot(ssh[24500:25000])
2880/96.
1600/96.
800/96.
print (times[24700])
| IonaOutfall/PreliminaryReport2016.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import syft as sy
import numpy as np
from syft.core.adp.entity import Entity
# ### Loading the dataset
data = pd.read_csv("../datasets/ca - feb 2021.csv")[0:10]
data.head()
# ### Logging into the domain
# Let's login into the domain
ca = sy.login(email="<EMAIL>", password="<PASSWORD>", port=8081)
# ### Upload the dataset to Domain node
# +
# We will upload only the first few rows
# All these three columns are of `int` type
# NOTE: casting this tensor as np.int32 is REALLY IMPORTANT. We need to create flags for this or something
canada_trade = ((np.array(list(data['Trade Value (US$)'])) / 100000)[0:10]).astype(np.int32)
trade_partners = ((list(data['Partner'])))[0:10]
entities = list()
for i in range(len(trade_partners)):
entities.append(Entity(name=trade_partners[i]))
# Upload a private dataset to the Domain object, as the root owner
sampled_canada_dataset = sy.Tensor(canada_trade)
sampled_canada_dataset.public_shape = sampled_canada_dataset.shape
sampled_canada_dataset = sampled_canada_dataset.private(0, 3, entity=entities[0]).tag("trade_flow")
# -
ca.load_dataset(
assets={"Canada Trade": sampled_canada_dataset},
name="Canada Trade Data - First few rows",
description="""A collection of reports from Canada's statistics
bureau about how much it thinks it imports and exports from other countries.""",
)
ca.datasets
# ### Create a Data Scientist User
ca.users.create(
**{
"name": "<NAME>",
"email": "<EMAIL>",
"password": "<PASSWORD>",
"budget":200
}
)
# ### Accept/Deny Requests to the Domain
ca.requests.pandas
ca.requests[-1].accept()
| notebooks/trade_demo/demo/Data Owner - Canada.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SageMath 9.4
# language: sage
# name: sagemath-9.4
# ---
# 
# ---
#
# <NAME> (<EMAIL>)
#
# ---
#
# # Propiedades de la Solución de Schwarzschild
# ---
#
# ## Resumen
#
# Este cuaderno utiliza [SageMath](https://www.sagemath.org) y [SageManifolds](https://sagemanifolds.obspm.fr/index.html) para calcular algunas propiedades geométricas del espacio-tiempo de Schwarzschild incluyendo simbolos de Christoffel, el tensor de Riemann, el tensor de Ricci y el escalar de Kretschmann.
# Nota: Este cuaderno fue creado utilizando SageMath >= 9.4
version()
# Activamos la visualización matemática via LaTeX,
# %display latex
# ## 1. Definición del Espacio-Tiempo
#
# El espaciotiempo se define como una variedad Lorentziana 4-dimensional,
MU = Manifold(4, 'MU', structure='Lorentzian', latex_name='\mathcal{M}')
print(MU)
# ### 1.1. Coordenadas Estandard
#
# Se introducen las coordenadas esfericas (estandard) o **coordenadas de Schwarzschild-Droste** utilizando el método `.chart()` del objeto variedad`MU`. El argumento del método `.chart()` es una cadena de texto que define cada una de las coordenadas y sus rangos, si son diferentes de $(-\infty, +\infty)$, al igual que el sìmbolo LaTeX que las representará.
#
# Note que las variables de `Python` para cada coordenada son declaradas dentro del operador `<...>`en el lado izquierdo. La variable `X` denotará la carta coordenada.
X.<t,r,th,ph> = MU.chart(r"t r:(0,+oo) th:(0,pi):\theta ph:(0,2*pi):\phi")
X
# ## 2. Tensor Metrico
#
# Para definir la métrica de Schwarzschild, es necesario declarar el parámetro de masa $M$ como una variable simbólica utilizando al función `var()`. Adicionalmente asumiremos que este parámetro toma valores positivos.
M = var('M')
assume(M>=0)
a = var('a', domain='real')
Sigma = var('Sigma', latex_name='\\Sigma')
Delta = var('Delta', latex_name='\\Delta')
F = 1 - 2*M/r
Sigma = r**2 + a**2*cos(th)**2
Delta = r**2 - 2*M*r + a**2
#Delta = F*Sigma + a**2*sin(th)**2
# Ahora definimos el tensor métrico
g = MU.metric()
g[0,0] = -(1 - 2*M*r/Sigma)
g[1,1] = Sigma/Delta
g[2,2] = Sigma
g[3,3] = (r**2 + a**2 + 2*M*r*(a*sin(th))**2/Sigma)*sin(th)^2
g[0,3] = -2*M*r*a*(sin(th))**2/Sigma
g.display()
g = MU.metric()
g[0,0] = (Delta-a**2*sin(th)**2)/Sigma
g[1,1] = -Sigma/Delta
g[2,2] = -Sigma
g[3,3] = -((r**2 + a**2)**2 - Delta*a**2*sin(th)**2)*sin(th)^2/Sigma
g[0,3] = (r**2 + a**2 - Delta)*a*sin(th)**2/Sigma
g.display()
g[:]
# ## 3. Simbolos de Christoffel (Conexiones)
#
# Los simbolos de Christoffel (de segunda especie) para la métrica $g$ en las coordenadas de Schwarzschild-Droste se definen como
#
# \begin{equation}
# \Gamma^\alpha_{\mu \nu} = \frac{1}{2} g^{\alpha \sigma} \left[ \partial_\mu g_{\sigma \nu} + \partial_\nu g_{\mu \sigma } - \partial_\sigma g_{\mu \nu} \right]
# \end{equation}
#
#
# Estas componentes se pueden obtener mediante el método `.christoffel_symbols_display()` del objeto métrica, `g`. Por defecto, solo se mostrarán las componentes no-nulas y no-redundantes (i.e. se tiene en cuenta la propiedad de simetría $\Gamma^\alpha_{\mu \nu} = \Gamma^\alpha_{\nu \mu}$).
#
# De igual forma, utilizando el símbolo `?` al final del comando, i.e. `g.christoffel_symbols_display?` se visualiza la información y opciones disponibles para el método.
g.christoffel_symbols_display?
g.christoffel_symbols_display()
# También es posible acceder a cada componente particular,
g.christoffel_symbols()[0,1,0]
# Podemos comprobar que la simetría en los indices inferiores se satisface,
g.christoffel_symbols()[0,0,1] == g.christoffel_symbols()[0,1,0]
# ## 4. Tensor de Curvatura
#
# El tensor de curvatura de Riemann se define como
#
# \begin{equation}
# R^\mu_{\,\, \nu \rho \sigma} = \frac{\partial \Gamma^\mu_{ \nu \sigma}}{\partial x^{\rho}} - \frac{\partial \Gamma^\mu_{ \nu \rho}}{\partial x^{\sigma}} + \Gamma^{\alpha}_{\nu \sigma}\Gamma^\mu_{\rho \alpha} - \Gamma^{\alpha}_{\ \nu \rho}\Gamma^\mu_{\sigma \alpha}
# \end{equation}
#
#
# Este puede calculares mediante el método `.riemann()` de la métrica,
Riemann = g.riemann()
print(Riemann)
# **Es importante notar que este es un tensor de orden (1,3)**.
Riemann
# Para visualizar sus componentes utilizamos el método `.display()`,
Riemann.display()
Riemann[:]
# Cada componente particular se visualiza utilizando los índices correspondientes. Por ejemplo, la componente $R^0_{\ \,101} = R^t_{\ \,trt}$ será
Riemann[0,1,1,0]
Riemann[0,1,0,1]
# Para obtener el tensor de curvatura del tipo (0,4) se utiliza el método `.down()`:
Riemann_dddd = Riemann.down(g)
Riemann_dddd
Riemann_dddd.display()
Riemann_dddd.display_comp()
Riemann_dddd[0,1,0,1]
Riemann_dddd[0,1,1,0]
Riemann_dddd[1,0,0,1]
Riemann_dddd[1,0,0,1]
# ## 5. Tensor de Ricci
#
# El tensor de Ricci se calcula con el método `.ricci()` de la metrica,
Ricci = g.ricci()
Ricci[:]
# Como era de esperar, todas sus componentes son cero en este caso,
Ricci.display()
Ricci[:]
# ### Tensor de Ricci Tensor (Version Alternativa)
#
# Otra forma de calcular este tensor es contrayendo el trensor de Riemann
#
# \begin{equation}
# R_{\mu \nu} = R^\alpha_{\,\, \mu \alpha \nu} = g ^{\alpha \beta} R_{\alpha \mu \beta \nu}
# \end{equation}
Ric = g.up(g)['^{ab}'] * Riemann.down(g)['_{acbd}']
Ricci[:]
# ### 5.1. Escalar de Curvatura de Ricci
#
# El escalar de curvatura puede obtenerse contrayendo la métrica con el tesnor de Ricci,
R = g['_{ab}'] * Ricci.up(g)['^{ab}']
print(R)
R.display()
# La expresión simbólica de un campo escalar se realiza utilizando el método `.expr()`:
R.expr()
# También es posible utilizar el método `.ricci_scalar()`
R = g.ricci_scalar()
R.expr()
# ## 6. Escalar de Kretschmann
#
# El escalar de Kretschmann es el "cuadrado" del tensor de Riemann,
#
# \begin{equation}
# K = R_{\mu \nu \rho \sigma} R^{\mu \nu \rho \sigma}
# \end{equation}
#
# Para calcularlo, se necesita el tensor de curvtura con todos los indices abajo y con todos los indices arriba, i.e. $R_{\mu \nu \rho \sigma}$ y
# $R^{\mu \nu \rho \sigma}$. Estos se evaluan utilizando los métodos`.down()` y `.up()`. Luego se realiza la contracción de los índices en el orden establecido,
K = Riemann.down(g)['_{abcd}'] * Riemann.up(g)['^{abcd}']
K
K.display()
K.expr()
# ### Summary of methods
#
# ```
# Manifold.chart()
# Manifold.metric()
# ```
#
#
# ```
# metric.christoffel_symbols_display()
# metric.christoffel_symbols()[]
# metric.riemann()
# metric.ricci()
# ```
#
#
# ```
# tensor.display()
# tensor[:]
# tensor.expr()
# tensor.up(metric)
# tensor.down(metric)
# ```
g.display?
Einstein = Ricci - R*g/2
Einstein[:]
Einstein_ud = Einstein.up(g,1)
Einstein_ud[:]
Einstein[0,0].simplify()
| 04.Kerr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6
# language: python
# name: python36
# ---
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
# # Creating and Updating a Docker Image before Deployment as a Webservice
#
# This notebook demonstrates how to make changes to an existing docker image, before deploying it as a webservice.
#
# Knowing how to do this can be helpful, for example if you need to debug the execution script of a webservice you're developing, and debugging it involves several iterations of code changes. In this case it is not an option to deploy your application as a webservice at every iteration, because the time it takes to deploy your service will significantly slow you down. In some cases, it may be easier to simply run the execution script on the command line, but this not an option if your script accumulates data across individual calls.
#
# We describe the following process:
#
# 1. Configure your Azure Workspace.
# 2. Create a Docker image, using the Azure ML SDK.
# 3. Test your Application by running the Docker container locally.
# 4. Update the execution script inside your running Docker container.
# 5. Commit the changes in your Docker container to its image
# 6. Update the image in the Azure Container Registry (ACR).
# 7. Deploy your Docker image as an Azure Container Instance ([ACI](https://azure.microsoft.com/en-us/services/container-instances/)) Webservice.
# 8. Test your Webservice.
#
# > Several cells below are completely commented out. This is because they can only be run on Jupyter, but not on Azure Databricks. If you do have access to Jupyter, we recommend to explore these cells, because they give you an insight into how to debug a docker container.
# ### Prerequisites
# - You need to have an [Azure](https://azure.microsoft.com) subscription. You will also need to know a subscription_id and resource_group. One way to discover those is to visit the [azure portal](https://portal.azure.com) and look use the same as those of the DSVM.
#
# **Note:**
# - This code was tested on a Data Science Virtual Machine ([DSVM](https://azure.microsoft.com/en-us/services/virtual-machines/data-science-virtual-machines/)), running Ubuntu Linux 16.04 (Xenial). **Do *not* try to run this notebook on *Databricks***, because you may run into trouble executing some of the shell and docker commands.
# - If you get an error message when trying to import `azureml` in the first cell below, you probably have to switch to using the correct kernel: `Python [conda env:amladpm]`.
# ## Configure your Azure Workspace
#
# We start by selecting your workspace, and make sure we have access to it from here. In order for this to work, make sure you followed the instructions for creating a workspace in your development environment:
#
# - [DSVM](../lab0.0_Setting_Up_Env/configure_environment_DSVM.ipynb)
# - [Aure Databricks](../lab0.0_Setting_Up_Env/configure_environment_ADB.ipynb)
# +
# # %matplotlib inline
import os
from azureml.core import Workspace
import pandas as pd
import urllib
config_path = '..'
# # run this if you are using Jupyter (instead of azure datarbicks)
# config_path = os.path.expanduser('~')
ws = Workspace.from_config(path=os.path.join(config_path, 'aml_config', 'config.json'))
ws.get_details()
# -
# Let's make sure that you have the correction version of the Azure ML SDK installed on your workstation or VM. If you don't have the write version, please follow these [Installation Instructions](https://docs.microsoft.com/en-us/azure/machine-learning/service/quickstart-create-workspace-with-python#install-the-sdk).
# + tags=["check version"]
import azureml
# display the core SDK version number
print("Azure ML SDK Version: ", azureml.core.VERSION)
# -
# ## Create a Docker image using the Azure ML SDK
# ### Create a template execution script for your application
#
# We are going to start with just a barebones execution script for your webservice. This script will calculate the running average of numbers thrown at it.
#
# We recommend that you execute the cells for your `score.py` scripts twice.
#
# 1. With a `#` sign at the beginning of the first line. This way you can detect typos and syntax errors during execution.
# 2. If the script runs OK, you can remove the `#` sign, to write the script to a file instead of executing it.
# +
# # #%%writefile score.py
# import json # we use json in order to interact with the anomaly detection service via a RESTful API
# # The init function is only run once, when the webservice (or Docker container) is started
# def init():
# global running_avg, curr_n
# running_avg = 0.0
# curr_n = 0
# pass
# # the run function is run everytime we interact with the service
# def run(raw_data):
# """
# Calculates rolling average according to Welford's online algorithm.
# https://en.wikipedia.org/wiki/Algorithms_for_calculating_variance#Online
# :param raw_data: raw_data should be a json query containing a dictionary with the key 'value'
# :return: runnin_avg (float, json response)
# """
# global running_avg, curr_n
# value = json.loads(raw_data)['value']
# n_arg = 5 # we calculate the average over the last "n" measures
# curr_n += 1
# n = min(curr_n, n_arg) # in case we don't have "n" measures yet
# running_avg += (value - running_avg) / n
# return json.dumps(running_avg)
# -
# ### Create environment file for your Conda environment
#
# Next, create an environment file (environment.yml) that specifies all the python dependencies of your script. This file is used to ensure that all of those dependencies are installed in the Docker image. Let's assume your Webservice will require ``azureml-sdk``, ``scikit-learn``, and ``pynacl``.
# + tags=["set conda dependencies"]
from azureml.core.conda_dependencies import CondaDependencies
myenv = CondaDependencies()
myenv.add_conda_package("scikit-learn")
myenv.add_pip_package("pynacl==1.2.1")
myenv.add_pip_package("pyculiarity")
myenv.add_pip_package("pandas")
myenv.add_pip_package("numpy")
with open("environment.yml","w") as f:
f.write(myenv.serialize_to_string())
# -
# Review the content of the `environment.yml` file.
with open("environment.yml","r") as f:
print(f.read())
# ### Create the initial Docker image
#
# The next step is to create a docker image.
#
# We start by creating downloading a scoring script that we prepared for this course. You can skip the details of this script, if you are in a hurry. Briefly, it contains our solution to online anomaly detection from the previous lab.
# +
# uncomment the below lines to see the solution
filename = 'AD_score.py'
urllib.request.urlretrieve(
os.path.join('https://raw.githubusercontent.com/Azure/LearnAI-ADPM/master/solutions/', filename),
filename='score.py')
with open('score.py') as f:
print(f.read())
# +
# %%time
from azureml.core.image import ContainerImage
# configure the image
image_config = ContainerImage.image_configuration(execution_script = "score.py",
runtime = "python",
conda_file = "environment.yml")
# create the docker image. this should take less than 5 minutes
image = ContainerImage.create(name = "my-docker-image",
image_config = image_config,
models = [],
workspace = ws)
# we wait until the image has been created
image.wait_for_creation(show_output=True)
# let's save the image location
imageLocation = image.serialize()['imageLocation']
# -
# ## Test your Application by running the Docker container locally
#
# ### Download the created Docker image from the Azure Container Registry ([ACR](https://azure.microsoft.com/en-us/services/container-registry/))
#
# Here we use some [cell magic](https://ipython.readthedocs.io/en/stable/interactive/magics.html) to exchange variables between python and bash.
# +
# # %%bash -s "$imageLocation"
# # get the location of the docker image in ACR
# imageLocation=$1
# # extract the address of the repository within ACR
# repository=$(echo $imageLocation | cut -f 1 -d ".")
# # echo "Attempting to login to repository $repository"
# az acr login --name $repository
# # echo
# # echo "Trying to pull image $imageLocation"
# docker pull $imageLocation
# -
# ### Start the docker container
#
# We use standard Docker commands to start the container locally.
# +
# # %%bash -s "$imageLocation"
# # extract image name and tag from imageLocation
# image_name=$(echo $1 | cut -f 1 -d ":")
# tag=$(echo $1 | cut -f 2 -d ":")
# # echo "Image name: $image_name, tag: $tag"
# # extract image ID from list of downloaded docker images
# image_id=$(docker images $image_name:$tag --format "{{.ID}}")
# # echo "Image ID: $image_id"
# # we forward TCP port 5001 of the docker container to local port 8080 for testing
# # echo "Starting docker container"
# docker run -d -p 8888:5001 $image_id
# -
# ### Test the docker container
#
# We test the docker container, by sending some data to it to see how it responds - just as we would with a Webservice.
#
# > If you get an error message below, you may just have to wait a couple of seconds.
# +
# import json
# import requests
# import numpy as np
# import matplotlib.pyplot as plt
# values = np.random.normal(0,1,100)
# values = np.cumsum(values)
# running_avgs = []
# for value in values:
# raw_data = {"value": value}
# r = requests.post('http://localhost:8888/score', json=raw_data)
# result = json.loads(r.json())
# running_avgs.append(result)
# plt.close()
# plt.plot(values)
# plt.plot(running_avgs)
# display()
# -
# ## Modifying the container
#
# Let's make a change to the the execution script: Copy the changed ``AD_score.py`` into the running docker container and commit the changes to the container image.
# +
# # %%bash -s $imageLocation
# image_location=$1
# # extract image name and tag from imageLocation
# image_name=$(echo $image_location | cut -f 1 -d ":")
# tag=$(echo $image_location | cut -f 2 -d ":")
# # echo "Image name: $image_name, tag: $tag"
# # extract image id
# image_id=$(docker images $image_name:$tag --format "{{.ID}}")
# # echo "Image ID: $image_id"
# # extract container ID
# container_id=$(docker ps | tail -n1 | cut -f 1 -d " ")
# # echo "Container ID: $container_id"
# # # copy modified scoring script again
# docker cp AD_score.py $container_id:/var/azureml-app/score.py
# sleep 1
# # stop the container
# docker restart $container_id
# -
# ## Test the container
# ### Load telemetry data
# +
base_path = 'https://sethmottstore.blob.core.windows.net'
data_dir = os.path.join(base_path, 'predmaint')
print("Reading data ... ", end="")
telemetry = pd.read_csv(os.path.join(data_dir, 'telemetry.csv'))
print("Done.")
print("Parsing datetime...", end="")
telemetry['datetime'] = pd.to_datetime(telemetry['datetime'], format="%m/%d/%Y %I:%M:%S %p")
telemetry.columns = ['timestamp', 'machineID', 'volt', 'rotate', 'pressure', 'vibration']
print("Done.")
# +
# import numpy as np
# import json
# import requests
# def test_docker(telemetry, n=None):
# """
# n is the number of sensor readings we are simulating
# """
# if not n:
# n = telemetry.shape[0]
# machine_ids = [1] # telemetry['machineID'].unique()
# timestamps = telemetry['timestamp'].unique()
# out_df = pd.DataFrame()
# for timestamp in timestamps[:10]:
# np.random.shuffle(machine_ids)
# for machine_id in machine_ids:
# data = telemetry.loc[(telemetry['timestamp'] == timestamp) & (telemetry['machineID'] == machine_id), :]
# json_data = data.to_json()
# input_data = {"data": json_data}
# r = requests.post('http://localhost:8888/score', json=input_data)
# result = pd.read_json(json.loads(r.json()))
# out_df = out_df.append(result, ignore_index=True)
# return out_df
# +
# print("Processing ... ")
# out_df = test_docker(telemetry)
# print("Done.")
# out_df
# -
# ### Push the updated container to ACR
#
# **First**, test your Docker container again (run the json query above), to ensure that the changes are having the expected effect.
#
# **Then** you can push the image into ACR, so that it can be retrieved by the Azure ML SDK when you want to deploy your Webservice.
# +
# # %%bash -s "$imageLocation"
# image_location=$1
# # extract container ID
# container_id=$(docker ps | tail -n1 | cut -f 1 -d " ")
# # echo "Container ID: $container_id"
# # commit changes made in the container to the local copy of the image
# docker commit $container_id $image_location
# docker push $image_location
# -
# Let's try to deploy the container to ACI, just to make sure everything behaves as expected.
# +
# %%time
from azureml.core.webservice import Webservice
from azureml.core.image import ContainerImage
from azureml.core.webservice import AciWebservice
# create configuration for ACI
aciconfig = AciWebservice.deploy_configuration(cpu_cores=1,
memory_gb=1,
tags={"data": "some data", "method" : "machine learning"},
description="Does machine learning on some data")
# pull the image
image = ContainerImage(ws, name='my-docker-image')
# deploy webservice
service_name = 'my-web-service'
service = Webservice.deploy_from_image(deployment_config = aciconfig,
image = image,
name = service_name,
workspace = ws)
service.wait_for_deployment(show_output = True)
print(service.state)
# +
import numpy as np
import json
import requests
def test_webservice(telemetry, n=None):
"""
n is the number of sensor readings we are simulating
"""
if not n:
n = telemetry.shape[0]
machine_ids = [1] # telemetry['machineID'].unique()
timestamps = telemetry['timestamp'].unique()
out_df = pd.DataFrame()
for timestamp in timestamps[:n]:
np.random.shuffle(machine_ids)
for machine_id in machine_ids:
data = telemetry.loc[(telemetry['timestamp'] == timestamp) & (telemetry['machineID'] == machine_id), :]
json_data = data.to_json()
input_data = bytes(json.dumps({"data": json_data}), encoding = 'utf8')
result = pd.read_json(json.loads(service.run(input_data=input_data)))
out_df = out_df.append(result, ignore_index=True)
return out_df
# -
print("Processing ... ")
out_df = test_webservice(telemetry, n=10)
print("Done.")
out_df
service.serialize()
# ## Clean up resources
#
# To keep the resource group and workspace for other tutorials and exploration, you can delete only the ACI deployment using this API call:
service.delete()
# # The end
# Copyright (c) Microsoft Corporation. All rights reserved.
#
# Licensed under the MIT License.
| lab01.5_AD_Model_Deployment/lab01.5_AD_model_deployment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook explains all the steps taken to pre-process the unigram dataset.
# +
import csv
import pandas as pd
import numpy as np
pd.options.mode.chained_assignment = None
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib
matplotlib.style.use('ggplot')
from scipy.interpolate import interp1d
from nltk.stem import WordNetLemmatizer
lemmatizer = WordNetLemmatizer()
def tf(x):
return 1+np.log10(x)
from itertools import groupby
from operator import itemgetter
import pickle as pkl
import re
def patternmaker(x):
x=np.array(x.notnull())
x=x.astype(int)
#print(x)
val = ''.join(map(str, x))
#print(val)
return val
pd.set_option('display.float_format', lambda x: '%.3f' % x)
# -
# # Introduction
#
# The datasets are from the site http://www.ngrams.info/download_coha.asp.
# For my experimentation, I wanted to see if this dataset could be a candidate for novel compound prediction.
#
# The uni-gram dataset has the following fields -
# 1. count - Frequency of the uni-gram
# 2. unigram - The uni-gram
# 3. pos - The POS of the uni-gram, and
# 4. decade -The decade timestamp, where 1 is for 1810s, 2 for 1820s and so on.
#
#
#
# For the initial trials I looked only into noun-noun compounds (I did perform experiments with adjective noun pairs, but many non compounds were returned).
#
unigram=pd.read_csv("1_pos_y_cs_y.txt",quotechar='"',delimiter="\t",quoting=csv.QUOTE_ALL,encoding='cp1252',skiprows=[1])
unigram.columns=['count','unigram','pos','decade']
pos_counts=unigram.pos.value_counts().to_frame()
pos_counts['perc']=round((pos_counts.pos/pos_counts.pos.sum())*100,3)
pos_counts.head(10)
# If we look at the pos tags, we see that there are about 280 of them. The POS tag description is provided on
# http://ucrel.lancs.ac.uk/claws7tags.html. The large value of possible tags is due to the presence of “DITTO
# Tags” (explanation found at the bottom of the webpage).
# # Selection of only noun tags
# From the tag explainations, nn, nn1 and nn2 are tags for common nouns (the '1' and '2' denote if the noun is singular or plural).
noun_tags=["nn","nn1","nn2"]
#adj_tags=["jj"]
unigram=unigram[unigram.pos.isin(noun_tags)]
nouns=dict.fromkeys(noun_tags,'noun')
#adj=dict.fromkeys(adj_tags,'adj')
unigram.head(10)
# # Exploratory data analysis
# As we see from the table above, the large number of tags should not be a concern as most have really small frequencies.
# If we look at the count of words per decade, we see that the counts are increasing.
# This should of course, come at no surprise.
decade_counts=unigram.decade.value_counts().to_frame()
decade_counts['perc']=round((decade_counts.decade/decade_counts.decade.sum())*100,3)
decade_counts
decade_counts.drop('decade',axis=1).plot.bar()
#possibile_tags=["nn","nn1","nn2","jj"]
noun_counts=pos_counts[pos_counts.index.isin(noun_tags)]
print(noun_counts)
print("\n Percentage of words that are nouns is ", round(noun_counts.perc.sum(),3) ,"%")
# # Lemmatization
# Lemmatization is performed on the dataset so that both singular and plural nouns get lemmatized to their common lemma form.
unigram.drop(['pos'],axis=1,inplace=True)
unigram.info()
# +
unigram['unigram']=unigram['unigram'].str.lower()
unigram['word']=unigram['unigram'].apply(lambda x: lemmatizer.lemmatize(str(x)))
unigram.drop(['unigram'],axis=1,inplace=True)
display(unigram.head(10))
unigram.info()
# -
# Following operations were performed -
#
# 1. substitute all noun tags with 'noun'
# 2. merge the unigram and its tag together
# 3. reduce all words to lowercase
# Some rows need to be merged after the tag class being simplified.
#
# Also only words occuring at least 3 times in a decade are chosen.
unigram=unigram.groupby(['word','decade'])['count'].sum().to_frame()
unigram=unigram.reset_index(level=['word','decade'])
#unigram=unigram[unigram['count']>=10]
display(unigram.head(10))
unigram.info()
# # Shallow to dense representation
# We can now finally pivot the dataset (convert it from a shallow to dense represenation).
#
#
# I have also added 3 extra columns which are,
#
# **cf** : Collection frequency, which is the log of the sum of the term across decades, ie log(1+sum(term).
#
# **presence** : Number of decades a term is present in.
#
# **pattern** : A string that is of length 18. A 0 if the word was not present in the particular decade and 1 otherwise. Could be useful later on during the LSTM reprsentation.
#
# Also, the decades are now suitably labeled.
#
# In order to reduce bias against words with extremely high frequencies, all counts are logarithmically reduced.
#
#
# +
unigram=unigram.pivot(index='word',columns='decade',values='count')
unigram.columns=['1810s','1820s','1830s','1840s','1850s','1860s','1870s','1880s','1890s','1900s','1910s','1920s','1930s','1940s','1950s','1960s','1970s','1980s']
#unigram = unigram.fillna(0)
#unigram[unigram.index.str.contains("['-]")==False]
unigram['cf']=unigram.sum(axis=1)
unigram['presence']=unigram.drop('cf',axis=1).count(axis=1)
#unigram['idf']=np.log10(18/unigram['presence'])
unigram['presence']=unigram["presence"].astype('category')
#unigram[unigram.columns[:-2]]=unigram[unigram.columns[:-2]].applymap(tf)
unigram['pattern']=unigram[unigram.columns[:-2]].apply(patternmaker,axis=1)
#unigram['term_weight']=unigram['cf']*unigram['idf']
display(unigram.head(10))
unigram.info()
# -
unigram.cf.sum()
unigram.to_csv('unigrams.csv',sep="\t")
# # End of Report
#
# All the details below are not important for the Novel Compound Generation and were for data analysis.
# The NANs in the above dataset are when words were not present for the particular decade.
# Even after certain pre-processing steps, the dataset appears to be messy due to presence of
# 1. Words such as “a-blowin_noun”,“a'throat_noun” (which might need to be parsed further),
# 2. Words such as “a'mighty_noun”, a'ready_noun (which seem to be missing letters)
# 3. words such as “zwei_noun” (which are not used in english).
# There also exist several uni-grams that are compound nouns already.
#unigram.loc[["a-blowin_noun","a'throat_noun","a'mighty_noun","zwei_noun"]]
unigram.index.get_loc('bed_noun')
# There also exist several uni-grams that are compound nouns already.
unigram.shape
unigram.describe()
# From the description table above, we see how higher counts can be biased towards the importance of a term.
#
# Hence another transformation is performed - Raw frequencies to term frequencies.
#
pattern_counts=unigram.pattern.value_counts().to_frame()
pattern_counts['perc']=round((pattern_counts['pattern']/pattern_counts['pattern'].sum())*100,3)
pattern_counts.sort_index(inplace=True)
pattern_counts
# +
#xlabels=pattern_counts[pattern_counts.pattern>4000].index
#xlabels
# -
pattern_counts.sort_values('perc', ascending=False).head(20).drop('perc',axis=1).plot.bar(figsize=(10,10))
# +
#fig, ax = plt.subplots()
#pattern_counts.pattern.plot(ax=ax,figsize=(15,15),rot=45)
#tick_idx = plt.xticks()[0]
#year_labels = df.years[tick_idx].values
#ax.xaxis.set_ticklabels(xlabels)
#pattern_counts.pattern.plot(figsize=(15,15),xticks=xlabels)
# -
pattern_counts.perc[pattern_counts.perc>=5.0].sum()
# From the table above we see that that log normalization has helped in reducing the variation of the terms across the decades.
# There are still 357971 possible unigrams in the dataset, and that would mean we would theoritically 357971*357971 rows in the bigram dataset.
#
# It then makes sence to figure out ways of eliminating uni-grams that are less "informative".
#
# A unigram could be considered to be "informative" if it -
#
# 1. Occurs consistently across the decades. (Given by the presence column)
# 2. Has a good frequency accoess the decades. (Given by the cf column)
#
#
# Given below are graphs and more statistics that hopefully could help you in deciding the correct amount of unigrams that should be selected.
decade_presence=unigram.presence.value_counts().to_frame()
decade_presence['perc']=round((decade_presence.presence/decade_presence.presence.sum())*100,3)
decade_presence.sort_index(inplace=True)
decade_presence
# The table above and the plot below show how the proportion of words changes w.r.t the Presence column.
fig, ax = plt.subplots()
decade_presence.drop('presence',axis=1).plot.bar(ax=ax,figsize=(10,10))
plt.xlabel('Number of decades the unigram is present in', fontsize=18)
plt.ylabel('Proportion of words', fontsize=18)
plt.show()
# What we could decipher from above is that several words only belong in 4 of fewer decades. Such words could be removed as they might be helpful in discovering future compounds.
#
# You could make a call on how important this statistic is.
# +
fig, ax = plt.subplots()
unigram.drop('presence',axis=1).plot.box(ax=ax,showfliers=True,figsize=(20,15),legend =True,notch=True)
plt.xlabel('Decades', fontsize=18)
plt.ylabel('Frequency', fontsize=18)
plt.show()
# -
# From the boxplot of Decades vs. the Frequency of a unigram we can see the visual represention of the description of the unigram dataset.
#
# The cf (Collection Frequency) column encompasses the information of a unigram for all the decades, and hence should be useful in selecting the possible unigrams.
unigram.hist(figsize=(20,20),alpha=0.5,bins=20,normed=1)
# The histograms do not seem to highlight anything out of the blue.
# The graphs below are maybe where Lonneke could give her perpective.
#
# Each individual graph shows how the Collection Frequency varies w.r.t the Presence column.
unigram.hist(by= 'presence',column ='cf', figsize=(20, 20),bins=50,sharex=True,log=True,sharey=True,normed=True,range=(1,7))
# In both the Histograms and the Boxplots, following are observed -
#
# 1. The distribution of the cf spreads out (becomes more normal as well) as the presence of a unigram increases.
# 2. The median and the IQR (the region represented by the box in the bloxplot) keep shifting higher.
#
#
# +
#fig, ax = plt.subplots()
unigram.boxplot(by= 'presence',column ='cf',figsize=(20, 20),notch=True)
#plt.figure(figsize=(20, 20))
#plt.axhline(y=2, color='r', linestyle='-')
#plt.xlabel('Presence', fontsize=18)
#plt.ylabel('Collection Frequency', fontsize=18)
#plt.show()
# -
# The boxplots could help you in deciding on the cutoff value for the collection frequency.
#
# For example, if you wish to select unigrams that have a collection frequency > 2 and have been seen in 4 or more decades, then you would the graph below.
#
# +
fig, ax = plt.subplots()
unigram[['presence','cf']].boxplot(ax=ax,by= 'presence',figsize=(20,20),notch=True)
#plt.figure(figsize=(20, 20))
plt.axhline(y=2, color='r', linestyle='-')
plt.axvline(x=4.5,color='g',linestyle='-')
plt.xlabel('Presence', fontsize=10)
plt.ylabel('Collection Frequency', fontsize=10)
plt.show()
# -
# The fact that the distributions are changing should not too suprising, as it makes sence that the cf inscreases as the decades go by.
ax=unigram.groupby('presence')['cf'].mean().plot()
ax.set(xlabel="Presence", ylabel="Collection Frequency")
# I hope all the above statistics helped you understand the complexity of the problem.
#
# Now, if we were to select a cut-off for the collection frequency value as we did before, we would get the new reduced dataset as below.
newunigram=unigram[((unigram.cf>2.0)==True) & (unigram.presence.isin([1,2,3,4])==False)]
newunigram
pattern_counts=newunigram.pattern.value_counts().to_frame()
pattern_counts['perc']=round((pattern_counts['pattern']/pattern_counts['pattern'].sum())*100,3)
pattern_counts.sort_index(inplace=True)
pattern_counts
pattern_counts.sort_values('perc', ascending=False).head(20).drop('perc',axis=1).plot.bar()
# As you can see, the dataset is reduced to 1/3rd its original size.
#
# We, however are still left with unsuitable unigrams such as "a-blaze_adj", "a'mighty_noun",etc.
#
# You can access all the unigrams in the dataset using the code below (they are grouped by the first letter of the unigram)
unigram_list=newunigram.index.tolist()
unigram_dict={}
for letter, words in groupby(sorted(unigram_list), key=itemgetter(0)):
#print(letter)
for word in words:
if letter not in unigram_dict:
unigram_dict[letter]=list()
unigram_dict[letter].append(word)
else:
unigram_dict[letter].append(word)
for letter in unigram_dict:
print("There are ",len(unigram_dict[letter]),"terms with the letter ", letter)
# You could now try to look through these terms and if any outliers exist.
#
# For example, if we look at some unigrams that start with a 'b' we see there exist outliers which -
#
# 1. Have symbols in between the words (b'nai_noun,b'y_noun) . These must be errors in reading the documents during OCR step
# 2. Are already compounds (baby-blue_adj,baby-clothes_noun,back-and-forth_adj).
#
#
#
#
# I hope all of the above steps and explained helped you in understanding the task.
#
# So, to reiterate your task would be to remove of unwanted unigrams that are present in the dataset.
# They could be due
# 1. Errors during scanning of the document,
# 2. Unigrams that are not in fact, unigrams
# 3. Unigrams that occur too infrequently and/or in too few decades
unigram_dict['b']
newunigram.to_csv("unigrams.csv")
| src/Notebooks/Constituents.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# This notebook is just used as scratch paper.
# nbi:hide
import ipywidgets as widgets
from ipywidgets import interact, interactive, interact_manual
import nbinteract as nbi
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# +
def x_vals(n):
return np.arange(n)
def y_vals(xs, n):
return xs ** n
nbi.line(x_vals, y_vals,
n=(1, 10))
# -
# nbi:right
@interact(x=(0, 10))
def plot(x):
plt.plot(np.arange(x), np.arange(x) ** 2)
# nbi:left
@interact(x=(0, 10))
def square(x):
return x * x
# This cell should display as a full row.
# nbi:hide_in
@interact(x=(0, 10))
def plot(x):
plt.plot(np.arange(x), np.exp(np.arange(x)))
# nbi:right
@interact(x=(0, 10))
def plot(x):
plt.plot(np.arange(x), np.arange(x) ** 2)
# nbi:left
@interact(x=(0, 10))
def square(x):
return x * x
# nbi:right
@interact(x=(0, 10))
def plot(x):
plt.plot(np.arange(x), np.arange(x) ** 3)
| notebooks/scratch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 1.116612, "end_time": "2021-02-22T16:59:54.263647", "exception": false, "start_time": "2021-02-22T16:59:53.147035", "status": "completed"} tags=[]
import numpy as np
import pandas as pd
import re
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
# + papermill={"duration": 0.120243, "end_time": "2021-02-22T16:59:54.395911", "exception": false, "start_time": "2021-02-22T16:59:54.275668", "status": "completed"} tags=[]
import pandas as pd
df = pd.read_table('SMSSpamCollection.txt', header=None)
df.columns = ['target', 'message']
# + papermill={"duration": 0.022866, "end_time": "2021-02-22T16:59:54.431375", "exception": false, "start_time": "2021-02-22T16:59:54.408509", "status": "completed"} tags=[]
df.shape
# + [markdown] papermill={"duration": 0.011894, "end_time": "2021-02-22T16:59:54.456178", "exception": false, "start_time": "2021-02-22T16:59:54.444284", "status": "completed"} tags=[]
# # Build a simple Bag of Words from Scratch
# + papermill={"duration": 0.027299, "end_time": "2021-02-22T16:59:54.496644", "exception": false, "start_time": "2021-02-22T16:59:54.469345", "status": "completed"} tags=[]
docs = ["Here at the Wall",
"What are the main reasons for.....",
"There are 700 possiblities that Alex will meet Alex Prime",
"Alpha prime is the member of Prime Groups",
"Is that all you got ?"]
# Preprocess the text
def preprocess(text):
text = text.lower()
text = re.sub(r"[^a-zA-Z0-9]", " ", text)
text = text.strip()
text = text.split(" ")
return text
preprocessed_docs = [preprocess(d) for d in docs]
print(preprocessed_docs)
# + papermill={"duration": 0.026337, "end_time": "2021-02-22T16:59:54.536612", "exception": false, "start_time": "2021-02-22T16:59:54.510275", "status": "completed"} tags=[]
def create_bow(docs):
bow = []
for d in docs:
count = dict()
for words in d:
count[words] = count.get(words, 0) + 1
bow.append(count)
return bow
create_bow(preprocessed_docs)
# + [markdown] papermill={"duration": 0.013027, "end_time": "2021-02-22T16:59:54.563849", "exception": false, "start_time": "2021-02-22T16:59:54.550822", "status": "completed"} tags=[]
# # Dealing with the DataSet at Hand
# + papermill={"duration": 0.025982, "end_time": "2021-02-22T16:59:54.603443", "exception": false, "start_time": "2021-02-22T16:59:54.577461", "status": "completed"} tags=[]
def preprocess_text(text):
text = text.lower()
text = re.sub(r"[^a-zA-Z0-9]", " ", text)
text = text.strip()
text = text.split()
text = ' '.join(list(filter(lambda x : x not in ['', ' '], text)))
return text
# + papermill={"duration": 0.12496, "end_time": "2021-02-22T16:59:54.742945", "exception": false, "start_time": "2021-02-22T16:59:54.617985", "status": "completed"} tags=[]
df.message = df.message.apply(preprocess_text)
df
# + papermill={"duration": 0.03924, "end_time": "2021-02-22T16:59:54.797842", "exception": false, "start_time": "2021-02-22T16:59:54.758602", "status": "completed"} tags=[]
# Split the data into Train/Test
X_train, X_test, y_train, y_test = train_test_split(df.message.values, df.target.values, test_size=0.1, stratify=df.target)
# + papermill={"duration": 0.024465, "end_time": "2021-02-22T16:59:54.838041", "exception": false, "start_time": "2021-02-22T16:59:54.813576", "status": "completed"} tags=[]
# In sklearn you can create a BOW using the CountVectorizer() function
bow = CountVectorizer(stop_words='english')
# + papermill={"duration": 0.260527, "end_time": "2021-02-22T16:59:55.113472", "exception": false, "start_time": "2021-02-22T16:59:54.852945", "status": "completed"} tags=[]
# Fit the bag of words on the training docs
bow.fit(X_train)
# + papermill={"duration": 0.141994, "end_time": "2021-02-22T16:59:55.271033", "exception": false, "start_time": "2021-02-22T16:59:55.129039", "status": "completed"} tags=[]
X_train = bow.transform(X_train)
X_test = bow.transform(X_test)
# + papermill={"duration": 0.050196, "end_time": "2021-02-22T16:59:55.336624", "exception": false, "start_time": "2021-02-22T16:59:55.286428", "status": "completed"} tags=[]
naive_bayes = MultinomialNB()
naive_bayes.fit(X_train, y_train)
# + papermill={"duration": 0.045808, "end_time": "2021-02-22T16:59:55.397842", "exception": false, "start_time": "2021-02-22T16:59:55.352034", "status": "completed"} tags=[]
from sklearn.metrics import accuracy_score, precision_score, recall_score, f1_score
print(f'Accuracy : {accuracy_score(y_test, naive_bayes.predict(X_test)):.3f}')
print(f'Precision : {precision_score(y_test, naive_bayes.predict(X_test), pos_label="spam"):.3f}')
print(f'Recall : {recall_score(y_test, naive_bayes.predict(X_test), pos_label="spam"):.3f}')
print(f'F1-Score : {f1_score(y_test, naive_bayes.predict(X_test), pos_label="spam"):.3f}')
# + [markdown] papermill={"duration": 0.015813, "end_time": "2021-02-22T16:59:55.429829", "exception": false, "start_time": "2021-02-22T16:59:55.414016", "status": "completed"} tags=[]
# # Testing the Model
# + papermill={"duration": 0.029701, "end_time": "2021-02-22T16:59:55.475872", "exception": false, "start_time": "2021-02-22T16:59:55.446171", "status": "completed"} tags=[]
# Test sample input
text = "You've Won! Winning an unexpected prize sounds great in theory. ..."
p_text = preprocess_text(text)
print(p_text)
p_text = bow.transform([p_text])
naive_bayes.predict_proba(p_text)[0][1]
# + [markdown] papermill={"duration": 0.016848, "end_time": "2021-02-22T16:59:55.509992", "exception": false, "start_time": "2021-02-22T16:59:55.493144", "status": "completed"} tags=[]
# # Saving model
# +
import pickle
with open('model_spam.pickle', 'wb') as handle:
pickle.dump(naive_bayes, handle)
with open('vectorizer_spam.pickle', 'wb') as handle:
pickle.dump(bow, handle)
| notebooks/spam_filter/naive-bayes-spam-classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KSY1526/myblog/blob/master/_notebooks/2022-01-23-FirstDeep3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="UBi2owFuBMgk"
# # "[처음 시작하는 딥러닝] 3. 밑바닥부터 만들어보는 CNN"
# - author: <NAME>
# - categories: [book, jupyter, Deep Learning, matrix, math, class]
# + [markdown] id="t7Ksob3VUnaE"
# # 기본 세팅
# + id="QuJTJyKDBHko"
import numpy as np
from numpy import ndarray
# + id="elQnyttoUrBw"
def assert_same_shape(output, output_grad):
assert output.shape == output_grad.shape, \
'''
두 ndarray의 모양이 같아야 하는데,
첫 번째 ndarray의 모양은 {0}이고
두 번째 ndarray의 모양은 {1}이다.
'''.format(tuple(output_grad.shape), tuple(output.shape))
return None
def assert_dim(t, dim):
assert len(t.shape) == dim, \
'''
이 텐서는 {0}차원이어야 하는데, {1}차원이다.
'''.format(dim, len(t.shape))
return None
# + [markdown] id="xjHQJSO7U-JJ"
# 필요한 차원을 잘 입력했는지 확인하는 함수를 선언합니다.
# + [markdown] id="VDNvhn1xBMl4"
# # 1차원 합성 곱
# + id="LpZDABFyQ9RA"
input_1d = np.array([1,2,3,4,5])
param_1d = np.array([1,1,1])
# + colab={"base_uri": "https://localhost:8080/"} id="zGDkVwLzSfAU" outputId="f377387a-6dc6-4e03-c5cd-f044cf2f6581"
def _pad_1d(inp, num): # 원래 데이터와 패딩 길이 입력
z = np.array([0])
z = np.repeat(z, num)
return np.concatenate([z, inp, z])
_pad_1d(input_1d, 1)
# + [markdown] id="MiCQOz44Tn8F"
# 간단한 함수로 1차원 데이터를 패딩한 모습입니다.
#
# 입력 데이터와 합성곱 연산을 한 출력 데이터의 크기를 같게하기 위해선 벗어나는 범위에 대해서 0 값을 채워주는 패딩을 하게 됩니다.
#
# 패딩 크기는 필터 크기를 2로 나눈 값에 정수부분이 입력과 출력을 같게하는 패딩의 크기가 됩니다.
# + colab={"base_uri": "https://localhost:8080/"} id="UNW4Xm-ES05q" outputId="820c4c88-ebc9-4d6a-b478-3c252bf7a3e1"
def conv_1d(inp, param): # 입력 값과 필터 값 입력
# 1차원 입력인지 확인합니다
assert_dim(inp, 1)
assert_dim(param, 1)
# 입력 값에 패딩을 덧붙입니다.
param_len = param.shape[0]
param_mid = param_len // 2
input_pad = _pad_1d(inp, param_mid)
# 초기값 부여
out = np.zeros(inp.shape)
# 1차원 합성곱 연산 수행
for o in range(out.shape[0]):
for p in range(param_len):
out[o] += param[p] * input_pad[o+p]
# 출력 모양이 입력과 동일한지 확인
assert_same_shape(inp, out)
return out
conv_1d(input_1d, param_1d)
# + [markdown] id="t9dI88xSWssj"
# 가중치가 [1,1,1] 인 간단한 합성곱 연산을 진행했습니다.
# + colab={"base_uri": "https://localhost:8080/"} id="KVxgyRWlWhkb" outputId="cb0d2f30-a561-45aa-95a7-3c1cac34f4e4"
# 출력값의 합을 구하는 함수
def conv_1d_sum(inp, param):
out = conv_1d(inp, param)
return np.sum(out)
input_1d = np.array([1,2,3,4,5])
input_1d_2 = np.array([1,2,3,4,6])
input_1d_3 = np.array([1,2,3,5,5])
param_1d = np.array([1,1,1])
param_1d_2 = np.array([2,1,1])
print(conv_1d_sum(input_1d, param_1d))
print(conv_1d_sum(input_1d_2, param_1d))
print(conv_1d_sum(input_1d_3, param_1d))
print(conv_1d_sum(input_1d, param_1d_2))
# + [markdown] id="mD3eCUzbY5lW"
# 입력값과 필터값이 달라짐에 따라 출력값의 합이 어떻게 바뀌는지 비교했습니다.
#
# 끝 쪽 입력값이 1 증가할때는 출력값의 합이 2증가, 가운데 쪽 입력값(패딩 영향 안받는)이 1 증가할때는 출력값의 합이 3증가합니다.
#
# 또 필터값이 1 증가할때 출력값의 합이 10 증가합니다.
# + colab={"base_uri": "https://localhost:8080/"} id="TUACXeaqYzPs" outputId="53852bb1-b8b8-44bb-ea39-349d5e81b5cd"
def _param_grad_1d(inp, param, output_grad = None):
# 입력값 패딩 추가
param_len = param.shape[0]
param_mid = param_len // 2
input_pad = _pad_1d(inp, param_mid)
if output_grad is None:
# 출력값의 기울기를 입력하지 않으면 1로 초기화.
# 왜냐하면 출력값의 합의 기울기이기 때문에 기울기를 유지하는 1을 쓰면됨.
output_grad = np.ones_like(inp)
else:
assert_same_shape(inp, output_grad)
# 모든 기울기의 초기값을 0으로 줍니다.
param_grad = np.zeros_like(param)
input_grad = np.zeros_like(inp)
for o in range(inp.shape[0]): # 0~4
for p in range(param.shape[0]): # 0~2
# 필터값의 기울기는 실제 영향을 받는 입력값의 합으로 됨
param_grad[p] += input_pad[o+p] * output_grad[o]
assert_same_shape(param_grad, param)
return param_grad
_param_grad_1d(input_1d, param_1d)
# + [markdown] id="6Cu2zz8pgLax"
# 1차원 합성 곱의 역방향 함수중 먼저 필터(파라미터) 기울기를 구하는 함수 입니다.
#
# 조금 어려운데 결과값을 간단히 해석하면 파라미터가 1 증가했을때 출력값의 합이 각각 10, 15, 14 증가한다는 것 입니다.
# + colab={"base_uri": "https://localhost:8080/"} id="NPQln7w4ch5_" outputId="64728efd-e269-4dc0-ef55-9eee61c72415"
def _input_grad_1d(inp, param, output_grad = None):
# 입력값 패딩 추가
param_len = param.shape[0]
param_mid = param_len // 2
input_pad = _pad_1d(inp, param_mid)
if output_grad is None:
# 출력값의 기울기를 입력하지 않으면 1로 초기화.
# 왜냐하면 출력값의 합의 기울기이기 때문에 기울기를 유지하는 1을 쓰면됨.
output_grad = np.ones_like(inp)
else:
assert_same_shape(inp, output_grad)
# 원할한 연산을 위해 범위 내 값은 1을, 범위를 벗어나는 것들은 0으로함.
# 패딩도 같은 효과를 냄.
output_pad = _pad_1d(output_grad, param_mid)
# 모든 기울기의 초기값을 0으로 줍니다.
param_grad = np.zeros_like(param)
input_grad = np.zeros_like(inp)
for o in range(inp.shape[0]): # 0~4
for f in range(param.shape[0]): # 0~2
# 입력값의 기울기는 실제 영향을 받는 필터값의 합으로 됨
input_grad[o] += output_pad[o + param_len - f - 1] * param[f]
assert_same_shape(param_grad, param)
return input_grad
_input_grad_1d(input_1d, param_1d)
# + [markdown] id="aF8UOdngsSGy"
# 입력값에 따른 출력값의 변동이 얼마나 되는지를 나타내는 함수 입니다.
#
# 첫번째와 마지막은 입력값이 1 증가할때 크기가 1 작은 2만큼 증가하고 나머지 값들은 3만큼 증가합니다.
#
# 패딩한 것에 영향받는 값을 제외하고 필터의 개수(3)만큼 영향력이 있다고 생각하면 됩니다.
# + [markdown] id="4kmH3crCswxY"
# # 배치 입력 적용하기
# + colab={"base_uri": "https://localhost:8080/"} id="XIHaAaNWsPdZ" outputId="062260ef-5c74-4f56-e8b1-0abb5f76dbff"
input_1d_batch = np.array([[0,1,2,3,4,5,6],
[1,2,3,4,5,6,7]])
def _pad_1d_batch(inp, num):
outs = [_pad_1d(obs, num) for obs in inp]
return np.stack(outs)
_pad_1d_batch(input_1d_batch, 1)
# + [markdown] id="zJ-GOQ5WtchM"
# 입력값이 2개 이상인 배치에도 적용하기 위해 기존 구현한 함수를 확장하겠습니다.
#
# 패딩의 경우 기존함수를 반복문을 이용해서 여러번 호출하면 됩니다.
# + colab={"base_uri": "https://localhost:8080/"} id="ZT607kOltVzD" outputId="eee77b48-97cb-4a3f-d7e5-1ad313f831fa"
def conv_1d_batch(inp, param):
outs = [conv_1d(obs, param) for obs in inp]
return np.stack(outs)
conv_1d_batch(input_1d_batch, param_1d)
# + [markdown] id="97VFqdJlt8Ls"
# 순방향 계산에 경우에도 같은 방식으로 확장했습니다.
# + colab={"base_uri": "https://localhost:8080/"} id="C0Oll_xLt5fY" outputId="966ac06b-7f36-4049-f0d5-eb43dfdc001c"
def input_grad_1d_batch(inp, param):
out = conv_1d_batch(inp, param)
out_grad = np.ones_like(out) # 출력기울기 값의 형태가 배치이므로 이에 맞게 조정
batch_size = out_grad.shape[0] # 배치 크기가 나옴
grads = [_input_grad_1d(inp[i], param, out_grad[i]) for i in range(batch_size)]
return np.stack(grads)
input_grad_1d_batch(input_1d_batch, param_1d)
# + [markdown] id="vUR9vE-422Vx"
# 입력값에 따른 출력값이 얼마나 되는지 구하는 함수를 배치로 확장했습니다.
#
# 기울기는 입력값에 영향이 있지 않기 때문에 어느 입력값이 입력되던 그대로 출력됩니다.
# + colab={"base_uri": "https://localhost:8080/"} id="OOYXeGkD21Ry" outputId="3163c82a-329e-4bf0-f5b9-22ad3d90c505"
def param_grad_1d_batch(inp, param):
output_grad = np.ones_like(inp)
# 단순 합의 기울기이기 때문에 모든 값을 1로 둡니다.
inp_pad = _pad_1d_batch(inp, 1)
out_pad = _pad_1d_batch(inp, 1)
param_grad = np.zeros_like(param)
for i in range(inp.shape[0]): # 배치 크기만큼
for o in range(inp.shape[1]): # 인풋 길이만큼
for p in range(param.shape[0]): # 필터 길이만큼
# 전부 합해줍니다.
param_grad[p] += inp_pad[i][o+p] * output_grad[i][o]
return param_grad
param_grad_1d_batch(input_1d_batch, param_1d)
# + [markdown] id="2-wvYSaM4r0i"
# 필터값에 따른 출력값이 얼마나 변하는지를 구하는 함수를 배치로 확장했습니다.
#
# 이때 필터에 대한 기울기는 배치 단위인데 필터는 모든 관찰과 합성곱 연선이 이뤄지므로 모든 값을 다 더해야합니다.
#
# 즉 모든 요소의 합이 필터 값이 바뀜에 따라서 얼마나 바뀌는지를 구하는 것 입니다.
# + [markdown] id="UeC1sRPv5Lrq"
# # 2차원 합성곱
# + colab={"base_uri": "https://localhost:8080/"} id="fWt-wHJw4nmc" outputId="634f2b49-9943-4540-9ec0-23ec0d8573b3"
imgs_2d_batch = np.random.randn(3, 28, 28)
param_2d = np.random.randn(3,3)
def _pad_2d_obs(inp, num):
# 가로 단위로 앞 뒷 값 각각 패딩
inp_pad = _pad_1d_batch(inp, num)
# 가로로 윗 2줄, 아래 2줄 패딩
other = np.zeros((num, inp.shape[0] + num * 2))
return np.concatenate([other, inp_pad, other])
def _pad_2d(inp, num):
# 첫번째 차원은 배치 크기에 해당함.
outs = [_pad_2d_obs(obs, num) for obs in inp]
return np.stack(outs)
_pad_2d(imgs_2d_batch, 1).shape
# + [markdown] id="9-K9wYNL7bpT"
# 2차원 단위에 입력값을 가지고 패딩을 진행했습니다.
#
# _pad_2d_obs 함수는 패딩을 실질적으로 진행하는 함수이고, _pad_2d 함수는 배치 단위로 확장하는 함수 입니다.
# + colab={"base_uri": "https://localhost:8080/"} id="xEofY3516hTU" outputId="7d15faa5-0575-4cbb-b792-86dde37b12b5"
def _compute_output_obs_2d(obs, param):
param_mid = param.shape[0] // 2
obs_pad = _pad_2d_obs(obs, param_mid)
out = np.zeros_like(obs)
# 2차원 필터를 거처 출력값을 만듭니다.
for o_w in range(out.shape[0]): # 출력값 가로길이
for o_h in range(out.shape[1]): # 출력값 세로길이
for p_w in range(param.shape[0]): # 필터 가로길이
for p_h in range(param.shape[1]): # 필터 세로길이
out[o_w][o_h] += param[p_w][p_h] * obs_pad[o_w+p_w][o_h+p_h]
return out
def _compute_output_2d(img_batch, param):
assert_dim(img_batch, 3)
outs = [_compute_output_obs_2d(obs, param) for obs in img_batch]
return np.stack(outs)
_compute_output_2d(imgs_2d_batch, param_2d).shape
# + [markdown] id="oSTOSCTL_87u"
# 2차원 단위에 순방향 계산입니다. 패딩을 먼저 시킨 뒤 2차원 필터를 통과시켜 모든 값을 합친 값을 출력해줍니다.
# + colab={"base_uri": "https://localhost:8080/"} id="8gL9p75x_4cP" outputId="dfc3f7d8-a968-4b24-d5d1-6c813e0dbfec"
def _compute_grads_obs_2d(input_obs, output_grad_obs, param):
# 입력을 나타내는 2차원값, 출력 기울기를 나타내는 2차원값(여기선 모두 1을 사용), 2차원 필터
param_size = param.shape[0] # 2차원 필터의 가로 세로가 같다고 가정합니다.
# 출력 기울기에 패딩을 먼저 덧붙입니다. 원본값은 1로, 나머지 값은 0으로하여 원본값만 유지하게 합니다.
output_obs_pad = _pad_2d_obs(output_grad_obs, param_size // 2)
input_grad = np.zeros_like(input_obs) # 초기 기울기는 0으로 합니다.
for i_w in range(input_obs.shape[0]): # 입력값 가로길이
for i_h in range(input_obs.shape[1]): # 입력값 세로길이
for p_w in range(param_size): # 필터 가로길이
for p_h in range(param_size): # 필터 세로길이
input_grad[i_w][i_h] += output_obs_pad[i_w + param_size - p_w -1][i_h + param_size - p_h -1] * param[p_w][p_h]
return input_grad
def _compute_grads_2d(inp, output_grad, param):
grads = [_compute_grads_obs_2d(inp[i], output_grad[i], param) for i in range(output_grad.shape[0])]
return np.stack(grads)
img_grads = _compute_grads_2d(imgs_2d_batch, np.ones_like(imgs_2d_batch), param_2d)
img_grads.shape
# + [markdown] id="hClsrKHlDEPp"
# 역방향 계산을 2차원으로 구현했습니다. 그 중 입력 기울기를 계산하는 절차인데요.
#
# 출력 기울기에 패딩을 덧붙이고 해당하는 가중치와의 합 연산을 하면 입력 기울기를 계산할 수 있습니다.
# + colab={"base_uri": "https://localhost:8080/"} id="-vhxIQxSCw5F" outputId="05326f58-50dc-4ce0-e0c4-82b9e68e9b3a"
def _param_grad_2d(inp, output_grad, param):
# 입력을 나타내는 3차원값, 출력 기울기를 나타내는 3차원값(여기선 모두 1을 사용), 2차원 필터
param_size = param.shape[0] # 2차원 필터의 가로 세로가 같다고 가정합니다.
inp_pad = _pad_2d(inp, param_size // 2) # 입력 값을 패딩합니다.
param_grad = np.zeros_like(param) # 초기 가중치 기울기를 0으로 합니다.
img_shape = output_grad.shape[1:] # 첫 값은 배치크기이므로 빼고 실행하기 위해.
for i in range(inp.shape[0]): # 배치 크기
for o_w in range(img_shape[0]): # 입력값 가로길이
for o_h in range(img_shape[1]): # 입력값 세로길이
for p_w in range(param_size): # 필터 가로길이
for p_h in range(param_size): # 필터 세로길이
param_grad[p_w][p_h] += inp_pad[i][o_w+p_w][o_h+p_h] * output_grad[i][o_w][o_h]
return param_grad
param_grad = _param_grad_2d(imgs_2d_batch, np.ones_like(imgs_2d_batch), param_2d)
param_grad
# + [markdown] id="RWyw0QlqHczd"
# 역방향 계산 중 필터 기울기를 구하는 부분을 2차원으로 구현했습니다.
#
# 여기서는 입력값에 패딩을 덧붙이고 합 연산을 했는데, 1차원하고 크게 다를게 없습니다.
#
# 배치 연산까지 한번에 진행하는 함수를 구현했는데 모든 배치의 입력값을 순회합니다.
# + [markdown] id="-DKpH9UubJfA"
# # 채널 추가하기
# + id="MiYbTw-9b0xY"
def _compute_output_obs(obs, param):
assert_dim(obs, 3)
assert_dim(param, 4)
param_size = param.shape[2]
param_mid = param_size // 2
obs_pad = _pad_2d_channel(obs, param_mid)
in_channels = param.shape[0]
out_channels = param.shape[1]
img_size = obs.shape[1]
out = np.zeros((out_channels,) + obs.shape[1:])
for c_in in range(in_channels):
for c_out in range(out_channels):
for o_w in range(img_size):
for o_h in range(img_size):
for p_w in range(param_size):
for p_h in range(param_size):
out[c_out][o_w][o_h] += \
param[c_in][c_out][p_w][p_h] * obs_pad[c_in][o_w+p_w][o_h+p_h]
return out
def _output(inp, param):
outs = [_compute_output_obs(obs, param) for obs in inp]
return np.stack(outs)
# + [markdown] id="b1y5WIxBcYcp"
# 합성곱층은 2차원으로 서로 엮인 뉴런 외에도 특징 맵과 같은 수의 채널을 갖습니다.
#
# 이런식으로 입력된 데이터를 다루기 위해 채널이 있는 순방향 연산을 구현했습니다. 역방향 연산은 생략합니다.
# + [markdown] id="LDNmTwtedOy-"
# # 기타 가벼운 이론
# + [markdown] id="jcUvG5VNdQiY"
# 이미지 데이터에서는 서로 가까운 픽셀 간에 유의미한 의미가 있는 조합이 나올 가능성이 높습니다.
#
# 즉 픽셀간 얼마나 공간적으로 가까운지를 나타내기 위해 합성곱 연산을 수행합니다.
#
# 풀링이란 각 특징 맵을 다운샘플링하여 데이터의 크기를 줄이는 방법입니다. 예시로 각 영역 픽셀 값의 최대값을 사용할 수 있습니다.
#
# 계산양 감소라는 이점이 있으나, 정보 손실또한 크기 때문에 이 방법을 쓰는데 다양한 의견이 있습니다.
#
# 스트라이드란 필터가 움직이는 간격이 지금까지는 1이였는데 이 값을 의미하는 용어로 커질수록 다운샘플링 효과가 커집니다.
#
# 최근 제안되는 고급 합성곱 신경망 구조에서는 풀링 대신 스트라이드를 2이상으로 설정해 다운샘플링 효과를 얻습니다.
| _notebooks/2022-01-23-FirstDeep3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from __future__ import print_function
import argparse
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torchvision import datasets, transforms
import numpy as np
import os
class CelebA_Attr(torch.utils.data.Dataset):
def __init__(self, root_dir, train_or_test, restricted_degree=0):
self._root_dir = root_dir
self._train_or_test = train_or_test
info_pak = np.load(os.path.join(self._root_dir, 'celeba_attr.npz'))
train_idxs = info_pak['train_idxs']
val_idxs = info_pak['val_idxs']
test_idxs = info_pak['test_idxs']
attribute_names = info_pak['attribute_names']
attributes = info_pak['attributes']
male_attr_idx = 20
def get_label(idxs, restricted_degree):
def jj(attr):
important_attributes_idx = [0, 1, 4, 9, 16, 18, 22, 24, 29, 30, 34, 36, 37, 38]
x = np.array([0 for i in range(attr.shape[0])])
for i in important_attributes_idx:
x = x + attr[:, i]
return x
label = attributes[idxs]
sig = jj(label) >= restricted_degree
label = label[sig]
data = np.delete(label, [male_attr_idx], 1)
label = label[:, male_attr_idx]
return data.astype('float32'), label
if self._train_or_test=='train':
self._data, self._label = get_label(train_idxs, restricted_degree)
elif self._train_or_test=='test':
self._data, self._label = get_label(test_idxs, restricted_degree)
def __len__(self):
return self._label.shape[0]
def __getitem__(self, index):
return self._data[index], self._label[index]
# +
# Training settings
batch_size = 32
test_batch_size = 1000
epochs = 10
lr = 0.001
eps = 1e-8
save_model = False
# +
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
kwargs = {'num_workers': 0, 'pin_memory': True} if use_cuda else {}
train_loader = torch.utils.data.DataLoader(
CelebA_Attr('./data/toy_celeba', 'train'),
batch_size=batch_size, shuffle=True, **kwargs)
test_loader = torch.utils.data.DataLoader(
CelebA_Attr('./data/toy_celeba', 'test'),
batch_size=test_batch_size, shuffle=True, **kwargs)
model = torch.nn.Sequential(
torch.nn.Linear(39, 200),
torch.nn.ReLU(),
torch.nn.Linear(200, 2),
torch.nn.LogSoftmax(dim=1)
).to(device)
optimizer = optim.Adam(model.parameters(), lr=lr, eps=eps)
# +
def train(model, device, train_loader, optimizer, criterion, epoch):
running_loss = 0.0
running_corrects = 0
model.train()
for batch_idx, (data, target) in enumerate(train_loader):
data, target = data.to(device), target.to(device)
optimizer.zero_grad()
output = model(data)
loss = criterion(output, target)
_, preds = torch.max(output, 1)
# _, labels = torch.max(target, 1)
loss.backward()
optimizer.step()
running_loss += loss.item() * data.size(0)
running_corrects += torch.sum(preds == target)
epoch_loss = running_loss / len(train_loader.dataset)
epoch_acc = running_corrects.double() / len(train_loader.dataset)
print('{} Loss: {:.4f} Acc: {:.4f}'.format(
'train', loss, epoch_acc))
def test(model, device, test_loader, criterion):
model.eval()
test_loss = 0
correct = 0
with torch.no_grad():
for data, target in test_loader:
data, target = data.to(device), target.to(device)
output = model(data)
test_loss += criterion(output, target).item() # sum up batch loss
pred = output.argmax(dim=1, keepdim=True) # get the index of the max log-probability
correct += pred.eq(target.view_as(pred)).sum().item()
test_loss /= len(test_loader.dataset)
print('Test set: Average loss: {:.4f}, Accuracy: {}/{} ({:.2f}%)\n'.format(
test_loss, correct, len(test_loader.dataset),
100. * correct / len(test_loader.dataset)))
# +
criterion = nn.CrossEntropyLoss(reduction='sum')
for epoch in range(1, epochs + 1):
print('Epoch {}'.format(epoch))
train(model, device, train_loader, optimizer, criterion, epoch)
test(model, device, test_loader, criterion)
# -
if (save_model):
torch.save(model.state_dict(),"mnist_cnn.pt")
| gender-classification/Attributes2Label_DNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ciência de dados - Unidade 3
#
# *Por: <NAME>, <NAME>, <NAME> e <NAME>*
#
# ### Objetivos
#
# O objetivo desse projeto é explorar os [datasets da UFRN](http://dados.ufrn.br/group/despesas-e-orcamento) contendo informações sobre requisições de material, requisições de manutenção e empenhos sob o contexto da [diminuição de verba](https://g1.globo.com/educacao/noticia/rio-grande-do-norte-veja-a-evolucao-do-orcamento-repassado-pelo-mec-as-duas-universidades-federais-do-estado.ghtml) que a UFRN recentemente vem sofrendo devido a crise financeira.
#
# De acordo com a pesquisa feita pelo nosso grupo, as fontes dizem que os cortes atingem [principalmente serviços terceirizados](https://g1.globo.com/educacao/noticia/90-das-universidades-federais-tiveram-perda-real-no-orcamento-em-cinco-anos-verba-nacional-encolheu-28.ghtml) como limpeza, manutenção e segurança, além de benefícios para estudantes de baixa renda, dado que estas [não são despesas obrigatórias] (https://g1.globo.com/educacao/noticia/salario-de-professores-das-universidades-federais-e-despesa-obrigatoria-mas-auxilio-estudantil-nao-entenda-a-diferenca.ghtml), ao contrário do pagamento de aposentadorias e pensões e o pagamento de pessoal ativo, no entanto, em [entrevista](http://www.tribunadonorte.com.br/noticia/na-s-vamos-receber-o-ma-nimo-diz-reitora-da-ufrn/399980), a atual reitora disse que o setor mais afetado seria o de obras e sua gestão, o que pode ser uma informação mais confiável, visto que até 2017 todo o orçamento era destinado diretamente as universidades federais, portanto eles decidiam como todos os gastos eram feitos. Isso mudou em 2018, já que o Ministério da Educação adotou uma nova metodologia que restringe ainda mais os gastos à "matriz Andifes" de forma que 50% do orçamento passou a ser gerenciado pelo próprio ministério da educação, logo a comparação do orçamento de 2018 com os anteriores deixa de ser possível.
#
# <hr>
# # 0 - Importando as bibliotecas
#
# Aqui utilizaremos o *pip* para instalar as bibliotecas necessárias para executar o notebook, sendo estas:
#
# - pandas
# - numpy
# - matplotlib
# - wordcloud
# !pip install pandas
# !pip install numpy
# !pip install matplotlib
# !pip install wordcloud
# # 1 - Lendo os datasets
#
# Nessa seção nós iremos importar os datasets contendo informações sobre requisiçoes de manutenção, requisições de material de serviço e empenhos, todos disponíveis no site de dados da UFRN.
#
# Na célula abaixo nós definimos uma lista com os arquivos que iremos precisar, lemos todos eles e os guardamos em um dicionário.
# +
import pandas as pd
from os import path
# Lista com o nome dos arquivos de todos os datasets que iremos utilizar
dataset_names = ['requisicaomanutencao.csv', 'requisicaomaterialservico.csv', 'empenhos.csv']
# Pasta em que os datasets se encontram
dataset_path = 'datasets'
# Dicionário onde eles serão armazenados
data = {}
# Loop que itera sobre todos os nomes definidos e armazena os dados lidos no dicionário
for name in dataset_names:
data[name[:-4]] = pd.read_csv(path.join(dataset_path, name), sep=';', low_memory=False)
# -
# Mostrando 'requisicaomanutencao.csv'
data['requisicaomanutencao']
# Mostrando 'requisicaomaterialservico.csv'
data['requisicaomaterialservico']
# Mostrando 'empenhos.csv'
data['empenhos']
# # 2 - Explorando e limpando os datasets
#
# Nessa seção é feita a análise das diferentes colunas dos datasets para identificar seus significados e suas utilidades para os problemas que iremos analisar. Sendo feita essa análise, nós então limpamos os datasets para que eles se tornem mais legíveis e mais fáceis de manusear.
# ## 2.1 - Requisição de manutenção
#
# Trata-se de um dataset listando todas as requisições de manutenções da UFRN desde 2005. Lembrando que iremos analisar apenas dados de 2008 a 2017, que são os anos em que temos o valor da verba total da UFRN.
maintenance_data = data['requisicaomanutencao']
print(maintenance_data.head())
print(maintenance_data.divisao.unique())
# ### 2.11 - Descrevendo as colunas e valores
#
# Observando o resultado da célula acima, podemos fazer as seguintes conclusões sobre as colunas:
#
# - <span style="color:red"><b>numero</b></span>: ID da requisição, não é relevante para o problema.
#
# - **ano**: Ano em que foi feita requisição de manutenção
#
# - **divisão**: Diz a divisão para qual a manutenção foi requisitada, assume os seguintes valores: 'Serviços Gerais', 'Instalações Elétricas e Telecomunicações', 'Instalações Hidráulicas e Sanitárias', 'Viário', 'Ar condicionado', 'Outros'.
#
# - **id_unidade_requisitante**: ID da unidade que fez a requisição.
#
# - **nome_unidade_requisitante**: Nome da unidade que fez a requisição.
#
# - **id_unidade_custo**: ID da unidade para qual o custo será destinado (pode ser igual a requisitante).
#
# - **nome_unidade_custo**: Nome da unidade para qual o custo será destinado (poder ser igual a requisitante).
#
# - **data_cadastro**: Data em que a requisição foi cadastrada.
#
# - **descricao**: Descrição da requisição, geralmente uma justificativa para aquela manutenção.
#
# - **local**: Local exato em que será feito a manutenção, pode ser uma sala, laboratório etc
#
# - <span style="color:red"><b>usuario</b></span>: Usuário que solicitou a manutenção. Provavelmente não tem utilidade para nosso problema.
#
# - **status**: Diz o status atual da requisição. Pode ajudar na análise de custos, considerando apenas as que já foram aprovadas, comparando a proporção de aprovadas e reprovadas para cada setor.
# ### 2.12 - Removendo colunas desnecessárias
#
# - <span style="color:red"><b>numero</b></span>: É apenas o ID da requisição
# - <span style="color:red"><b>usuario</b></span>: Não precisamos saber o usuário para nossa análise
# +
def remove_cols(df_input, dropped_columns):
'''
This functions receives a dataframe and a list of column names as input. It checks if each column exist,
and if they do, they're removed.
'''
for dropped_column in dropped_columns:
if dropped_column in df_input:
df_input = df_input.drop([dropped_column], axis=1)
return df_input
maintenance_dropped = ['numero', 'usuario']
maintenance_data = remove_cols(maintenance_data, maintenance_dropped)
maintenance_data.head()
# -
# ### 2.13 - Removendo outliers e valores desnecessários
#
# Aqui iremos analisar os valores do nosso dataset e determinar quais podemos remover ou modificar de modo a facilitar a nossa análise.
print(maintenance_data.status.value_counts())
# **Observação:**
#
# Checando os status, podemos perceber que a maioria dos valores ocorrem um número muito pequeno de vezes e não precisamos deles para nossa análise, portanto iremos eliminar os valores com 800 ocorrências ou menos
maintenance_data = maintenance_data.groupby('status').filter(lambda x: len(x) > 800)
maintenance_data.status.value_counts()
# **Observação:**
#
# Sobram portanto 5 valores possíveis para a coluna **status**. Porém, para nossa análise de custos, precisamos saber apenas se a requisição foi negada ou autorizada. Analisando os status restantes, podemos considerar que toda requisição que tiver valor diferente de negada pode ser considerada como autorizada.
# +
def convert_status(status_val):
'''Converts the value of all strings in the status column to AUTORIZADA, unless their value is NEGADA.'''
if status_val == 'NEGADA':
return status_val
else:
return 'AUTORIZADA'
maintenance_data['status'] = maintenance_data['status'].apply(convert_status)
maintenance_data.status.value_counts()
# -
maintenance_data.info()
print(maintenance_data.divisao.value_counts())
print(maintenance_data.nome_unidade_custo.value_counts())
# ### 2.14 - Lidando com valores nulos
#
# Aqui nós utilizaremos o método [pandas.DataFrame.info](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.info.html) para verificar quais os valores nulos das colunas de nosso dataset. A partir disso, dependendo da quantidade de colunas com valores nulos e do tipo de dado, nós iremos decidir o que fazer com esses valores.
maintenance_data.info()
maintenance_data.divisao.value_counts()
# **Observação**
#
# Utilizando o método [pandas.DataFrame.info](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.info.html) percebemos que existem muitos valores **NULL** na coluna *local* e alguns poucos na coluna *divisao*. Para a coluna *local*, iremos preencher as linhas **nulas** com seus valores de *nome_unidade_custo*. Para a coluna *divisao*, iremos preencher com o valor 'Outros', que é um dos mais comuns.
# +
import numpy as np
maintenance_data['local'] = np.where(maintenance_data.local.isnull(), maintenance_data.nome_unidade_custo, maintenance_data.local)
maintenance_data['divisao'] = maintenance_data['divisao'].fillna('Outros')
maintenance_data.info()
# -
# Resultado final da limpeza
maintenance_data.head()
# ## 2.2 - Requisição de material de serviço
#
# Esse dataset lista todas as requisições de materiais e serviços contratados pela UFRN desde 2008.
material_request_data = data['requisicaomaterialservico']
print('===== Primeiras linhas =====')
print(material_request_data.head())
print('===== Contagem de valores de natureza_despesa =====')
print(material_request_data.natureza_despesa.value_counts())
print('===== Contagem de valores de status =====')
print(material_request_data.status.value_counts())
# ### 2.21 - Descrevendo as colunas e valores
#
# Observando o resultado da célula acima, podemos fazer as seguintes conclusões sobre as colunas:
#
# - <span style="color:red"><b>numero</b></span>: ID da requisição, não é relevante.
#
# - **ano**: Ano em que foi feita a requisição.
#
# - **id_unidade_requisitante**: ID da unidade que fez a requisição, toda unidade tem um ID único.
#
# - **nome_unidade_requisitante**: Nome da unidade que fez a requisição.
#
# - **id_unidade_custo**: ID da unidade para qual os custos serão destinados, pode ser diferente da requisitante.
#
# - **nome_unidade_custo**: Nome da unidade para qual os custos serão destinados, pode ser diferente da requisitante.
#
# - **data_envio**: Data em que a requisição foi enviada.
#
# - <span style="color:red"><b>numero_contrato</b></span>: Aparentemente as requisições são feitas por meio de contratos, esse é o número do contrato.
#
# - **contratado**: Empresa contratada para fornecer o material.
#
# - <span style="color:red"><b>natureza_despesa</b></span>: Em todas as linhas analisadas, essa coluna tem o valor 'SERV. PESSOA JURÍDICA'.
#
# - **valor**: Valor pedido pela requisição.
#
# - **observacoes**: Comentário feito pela pessoa que fez a requisição, explicando o motivo desta
#
# - **status**: O status atual da requisição, está diretamente ligada ao empenho e pode assumir os seguintes valores: 'ENVIADA', 'PENDENTE ATENDIMENTO', 'CADASTRADA', 'ESTORNADA', 'LIQUIDADA', 'PENDENTE AUTORIZAÇÃO', 'FINALIZADA', 'EM_LIQUIDACAO', 'NEGADA', 'A_EMPENHAR', 'EMPENHO_ANULADO', 'AUTORIZADA', 'CANCELADA\n'.
# ### 2.22 - Removendo colunas desnecessárias
#
# As seguintes colunas serão dropadas
#
# - <span style="color:red"><b>numero</b></span>: Trata-se apenas do ID da requisição, não é necessário
#
# - <span style="color:red"><b>numero_contrato</b></span>: Informação desnecessária para a análise
#
# - <span style="color:red"><b>natureza_despesa</b></span>: Possui o mesmo valor em todas as linhas
# +
material_dropped = ['numero' ,'natureza_despesa', 'numero_contrato']
material_request_data = remove_cols(material_request_data, material_dropped)
print(material_request_data.head())
# -
# ### 2.23 - Removendo outliers e valores desnecessários
#
# Aqui iremos analisar os dados do nosso dataset e determinar quais podemos remover ou modificar de modo a facilitar a nossa análise.
print(material_request_data.status.value_counts())
# **Observação:**
# Verificando a contagem de valores da coluna *status*, percebemos que grande parte dos valores possíveis tem um número muito pequeno de ocorrências no dataset. Esses valores com poucas ocorrências influenciam pouco na nossa análise, portanto iremos eliminá-los.
# +
allowed_status = ['LIQUIDADA', 'EM_LIQUIDACAO', 'ENVIADA', 'ESTORNADA', 'FINALIZADA', 'CADASTRADA']
material_request_data = material_request_data[material_request_data.status.isin(allowed_status)]
print(material_request_data.status.value_counts())
# -
# ### 2.24 - Lidando com valores nulos
#
# Aqui nós utilizaremos o método [pandas.DataFrame.info](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.info.html) para verificar quais os valores nulos das colunas de nosso dataset. A partir disso, dependendo da quantidade de colunas com valores nulos e do tipo de dado, nós iremos decidir o que fazer com esses valores.
material_request_data.info()
material_request_data[material_request_data.data_envio.isnull()].head(n=20)
# - **data_envio**: Possui vários valores nulos. Como a maioria deles está bem separado um do outro e o dataset está ordenado por data, podemos preenchê-los usando o valor dessa coluna em linhas anteriores.
# - **observacoes**: Algumas observações também tem valores nulos, iremos simplesmente settar esses para uma string vazia.
# +
material_request_data.data_envio = material_request_data.data_envio.fillna(method='ffill')
material_request_data.observacoes = material_request_data.observacoes.fillna('')
material_request_data.info()
# -
# ## 2.3 - Empenhos
#
# Dataset contendo a relação de todos os empenhos efetuados pela UFRN desde 2001.
#
# O empenho da despesa importa em deduzir do saldo de determinada dotação orçamentária a parcela necessária à execução das atividades do órgão. É a forma de comprometimento de recursos orçamentários. Nenhuma despesa poderá ser realizada sem prévio empenho (art. 60 da Lei n° 4.320/64), sendo realizado após autorização do Ordenador de Despesa em cada Unidade Gestora Executora.
# +
empenhos_data = data['empenhos']
print(empenhos_data.head())
print(empenhos_data.data.value_counts())
# -
# ### 2.31 - Descrevendo as colunas e valores
#
# - <span style="color:red"><b>cod_empenho</b></span>: ID do empenho;
#
# - **ano**: Ano em que foi solicitado o empenho;
#
# - **modalidade**: O empenho da despesa pode assumir três tipos diferentes:
# - a) Ordinário – a despesa com valor exato deve ser liquidada e paga de uma única vez;
# - b) Estimativo – O valor total da despesa é estimado, podendo ser liquidado e pago em parcelas mensais;
# - c) Global – a despesa total é conhecida e seu pagamento é parcelado, de acordo com cronograma de execução.
#
# - **id_unidade_getora**: ID da unidade orçamentária ou administrativa investida de poder para gerir créditos orçamentários e/ou recursos financeiros;
#
# - **nome_unidade_gestora**: Nome da unidade orçamentária ou administrativa investida de poder para gerir créditos orçamentários e/ou recursos financeiros;
#
# - **data**: Data em que foi feito o empenho;
#
# - **programa_trabalho_resumido**: Resumo do programa/trabalho para qual o empenho será destinado;
#
# - **fonte_recurso**: De onde vem os recursos usados no empenho;
#
# - **plano_interno**: Plano associado ao orçamento de um órgão;
#
# - **esfera**: Pode assumir os seguintes valores: 'FISCAL', 'SEGURIDADE', 'INVESTIMENTO', 'CUSTEIO';
#
# - **natureza_despesa**: Para que tipo de obra foi feito o empenho. Podemos verificar a despesa para desenvolvimento de software, entre os valores dessas colunas temos: 'MAT. CONSUMO', 'SERV. PESSOA JURÍDICA', 'EQUIP. MATERIAL PERMANENTE', 'OBRAS E INSTALAÇÕES', 'PASSAGENS', 'SERVIÇOS DE TECNOLOGIA DA INFORMAÇÃO E COMUNICAÇÃO', 'DESENVOLVIMENTO DE SOFTWARE', 'DIV.EXERCÍCIOS ANTERIORES', 'SERV. PESSOA FÍSICA' 'LOC. MÃO-DE-OBRA', 'SERVIÇOS / UG-GESTÃO' etc.
#
# - **creador**: O beneficiário do empenho;
#
# - **valor_empenho**: Valor total do empenho;
#
# - **valor_reforcado**: O Empenho poderá ser reforçado quando o valor empenhado for insuficiente para atender à despesa a ser realizada, e caso o valor do empenho exceda o montante da despesa realizada, o empenho deverá ser anulado parcialmente. Será anulado totalmente quando o objeto do contrato não tiver sido cumprido, ou ainda, no caso de ter sido emitido incorretamente. Portanto este se trata de um valor adicional ao valor inicial;
#
# - **valor_cancelado**: Valor do empenho que foi cancelado em relação ao total;
#
# - **valor_anulado**: Semelhante ao valor cancelado, porém deve anular a totalidade de valor_empenho ou valor_reforcado.
#
# - **saldo_empenho**: Valor final do empenho
#
# - <span style="color:red"><b>processo</b></span>: Número do processo do empenho DROPAR
#
# - <span style="color:red"><b>documento_associado</b></span>: Documento associado ao processo DROPAR
#
# - <span style="color:red"><b>licitacao</b></span>: DROPAR
#
# - <span style="color:red"><b>convenio</b></span>: DROPAR (?) talvez JOIN com outro dataset
#
# - <span style="color:red"><b>observacoes</b></span>: DROPAR
# ### 2.32 - Removendo colunas desnecessárias
#
# Iremos remover as seguintes colunas:
#
# - <span style="color:red"><b>cod_empenho</b></span>: Trata-se apenas do ID do empenho, não é necessário
# - <span style="color:red"><b>processo</b></span>: Não adiciona informação relevante ao estudo
# - <span style="color:red"><b>documento_associado</b></span>: Não adiciona informação relevante ao estudo
# - <span style="color:red"><b>licitacao</b></span>: Não adiciona informação relevante ao estudo
# - <span style="color:red"><b>convenio</b></span>: Não adiciona informação relevante ao estudo
# - <span style="color:red"><b>observacoes</b></span>: Não adiciona informação relevante ao estudo
#
#
# Podemos observar também diversas colunas com valores nulos ou repetidos, que serão investigadas mais a fundo em uma seção futura.
# +
empenhos_dropped = ['cod_empenho', 'processo', 'documento_associado', 'licitacao', 'convenio', 'observacoes']
empenhos_data = remove_cols(empenhos_data, empenhos_dropped)
print(empenhos_data.head())
# -
# ### 2.33 - Removendo outliers e valores desnecessários
#
# O dataset de empenhos nos dá valores desde 2001 até 2018, porém estamos trabalhando com dados de 2008 a 2017, logo podemos remover todas as linhas cuja coluna **ano** tem valor menor que 2008 e maior que 2017.
# +
# Defining a vector with the years we'll analyse
years = [2008, 2009, 2010, 2011, 2012, 2013, 2014, 2015, 2016, 2017]
empenhos_data = empenhos_data[empenhos_data.ano.isin(years)]
# -
# ### 2.34 - Lidando com valores nulos
#
# Aqui nós utilizaremos o método [pandas.DataFrame.info](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.info.html) para verificar quais os valores nulos das colunas de nosso dataset. A partir disso, dependendo da quantidade de colunas com valores nulos e do tipo de dado, nós iremos decidir o que fazer com esses valores.
# +
empenhos_data.info()
empenhos_data[empenhos_data.valor_anulado.notnull()].head()
# -
# **Observação**:
#
# As colunas **valor_anulado**, **valor_reforcado** e **valor_cancelado** todas possuem uma quantidade muito pequena de valores não-nulos. Como as colunas **valor_empenho** e **saldo_empenho** possuem todos os valores, nós não precisamos das outras para fazermos nossa análise, logo podemos dropá-las.
# +
valores_drop = ['valor_reforcado', 'valor_anulado', 'valor_cancelado']
empenhos_data = remove_cols(empenhos_data, valores_drop)
empenhos_data.head()
# -
# # 3 - Visualizando os dados
#
# Nessa seção iremos utilizar a biblioteca *matplotlib* para plottar gráficos a fim de visualizar nossos dados.
# ## 3.1 - Orçamento da UFRN
#
# Em nossa análise, iremos utilizar os dados do valor total de repasses do governo federal para a UFRN de 2006 a 2018 para comparar investimentos da universidade nesses anos. Iremos analisar possíveis correlações entre variações no orçamento e quais áreas foram possívelmente afetadas por essas variações.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
years = [2008, 2009, 2010, 2011, 2012, 2013, 2014, 2015, 2016, 2017]
budget = [62010293, 136021308, 203664331, 172999177, 221801098, 246858171, 228864259, 207579799, 230855480, 186863902]
# +
# Plottagem do orçamento da UFRN de 2008 a 2017, podemos perceber que caiu em todos os anos desde 2013, exceto por 2016.
budget_scaled = [value / 1000000 for value in budget]
plt.rcParams['figure.figsize'] = (11,7)
plt.plot(years, budget_scaled, 'r')
plt.scatter(years, budget_scaled, color='green')
plt.xlabel("Ano")
plt.ylabel("Orçamento (em milhões de reais)")
plt.xticks(years)
plt.show()
# -
# ## 3.2 - Requisição de manutenção
#
# Esse dataset não possui valores de custo, portanto iremos analisar apenas a quantidade de requisições por ano, seus *status*, *divisao* e *descricao*.
# +
autorized_count_year = []
denied_count_year = []
for year in years:
status_count = maintenance_data[maintenance_data.ano == year].status.value_counts()
autorized_count_year.append(status_count['AUTORIZADA'])
denied_count_year.append(status_count['NEGADA'])
# +
import datetime
from matplotlib.dates import date2num
bar_width = 0.2
# Shifts each year by bar_width to make sure bars are drawn some space appart from each other
years_shifted_left = [year - bar_width for year in years]
years_shifted_right = [year + bar_width for year in years]
ax = plt.subplot(111)
ax.bar(years_shifted_left, autorized_count_year, width=bar_width, color='g', align='center')
ax.bar(years, denied_count_year, width=bar_width, color='r', align='center')
legends = ['Autorizadas', 'Negadas']
plt.legend(legends)
plt.ylabel("Quantidade")
plt.xlabel("Ano")
plt.xticks(years)
plt.title("Manutenções autorizadas x negadas de 2008 a 2017")
plt.show()
# +
divisao_year_count = []
# Keeps all unique values for 'divisao' column.
divisao_values = maintenance_data.divisao.unique()
for year in years:
maintenance_data_year = maintenance_data[maintenance_data.ano == year]
divisao_year_count.append(maintenance_data_year.divisao.value_counts())
# If a key doesn't exist in the count, we add it.
for possible_value in divisao_values:
for year_count in divisao_year_count:
if possible_value not in year_count.index:
year_count[possible_value] = 0
for year_count in divisao_year_count:
print(year_count)
# +
bar_width = 0.15
# Shifts each year by bar_width to make sure bars are drawn some space appart from each other
years_shifted_left = [year - bar_width for year in years]
years_shifted_right = [year + bar_width for year in years]
ax = plt.subplot(111)
colors = ['red', 'green', 'blue', 'orange', 'grey', 'black']
half_div_len = len(divisao_values) / 2.0
for i, divisao in enumerate(divisao_values):
total_divisao_count = []
for year in years:
total_divisao_count.append(divisao_year_count[i][divisao])
ax.bar(years, total_divisao_count, width=bar_width, color=colors[i], align='center')
plt.legend(divisao_values)
plt.ylabel("Quantidade")
plt.xlabel("Ano")
plt.xticks(years)
plt.title("Proporção dos tipos de manutenção de 2008 a 2017.")
plt.show()
# -
# ## 3.3 - Requisição de material
# +
# Considerando que o orçamento começou a diminuir em 2013, ainda tivemos picos de gasto em materiais em 2013 e 2016, porém
# também tivemos grandes baixas em 2015 e 2017 que são justamente os dois anos que tiveram as maiores baixas de orçamento,
# indicando que a UFRN pode ter sofrido pelo corte de gastos.
material_spending = []
for year in years:
material_spending.append(material_request_data[material_request_data.ano == year].valor.sum() / 1000000)
plt.plot(years, material_spending, 'r')
plt.scatter(years, material_spending, color='green')
plt.xlabel("Ano")
plt.ylabel("Gasto com material (em milhões de reais)")
plt.xticks(years)
plt.title("Valor gasto com material na UFRN de 2008 a 2017.")
plt.show()
# -
# ## 3.4 - Empenhos
# +
valor_year = []
saldo_year = []
for year in years:
valor_year.append(empenhos_data[empenhos_data.ano == year].valor_empenho.sum() / 1000000)
saldo_year.append(empenhos_data[empenhos_data.ano == year].saldo_empenho.sum() / 1000000)
plt.plot(years, valor_year, 'r', label='Valor pedido')
plt.scatter(years, valor_year, color='blue')
plt.title("Valor total pedido pelos empenhos da UFRN de 2006 a 2017.")
plt.xlabel('Ano')
plt.ylabel('Valor total (milhões)')
plt.xticks(years)
plt.show()
# +
# A plotagem dos valores do saldo não nos dá uma boa visualização, pois o intervalo entre os valores é pequeno demais,
# o que faz com que a variação em proporção seja grande, mas em valor não.
plt.plot(years, saldo_year, 'g')
plt.scatter(years, saldo_year, color='blue')
plt.title("Valor total empenhado pela UFRN de 2006 a 2017.")
plt.xlabel('Ano')
plt.ylabel('Saldo (milhões)')
plt.xticks(years)
plt.show()
# +
# O gráfico de barras nos dá uma visualização melhor. Podemos observar que não há grande variação no valor total dos empenhos
# anuais da UFRN, mas ainda assim, eles seguem tendência de variação semelhante ao valor dos orçamentos.
plt.bar(years, saldo_year)
plt.title("Saldo autorizado pelos empenhos da UFRN de 2006 a 2017.")
plt.xlabel("Ano")
plt.ylabel("Gastos (em milhões de reais)")
plt.xticks(years)
plt.show()
# +
bar_width = 0.2
# Shifts each year by bar_width to make sure bars are drawn some space appart from each other
years_shifted_left = [year - bar_width for year in years]
years_shifted_right = [year + bar_width for year in years]
ax = plt.subplot(111)
ax.bar(years_shifted_left, valor_year, width=bar_width, color='g', align='center')
ax.bar(years_shifted_right, saldo_year, width=bar_width, color='b', align='center')
ax.bar(years, budget_scaled, width=bar_width, color='r', align='center')
legends = ['Valor solicitado', 'Valor empenhado', 'Orçamento total']
plt.legend(legends)
plt.ylabel("Valor (milhões)")
plt.xlabel("Ano")
plt.xticks(years)
plt.title("Valor pedido vs. Valor empenhado vs. Orçamento")
plt.show()
# -
| .ipynb_checkpoints/empenhos_requests_ufrn_budget-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import sys
import time
import numpy as np
import cv2
from matplotlib import pyplot as plt
import importlib
import torch
import torchvision
from torchvision import datasets, transforms
from torch import nn, optim
import torch.nn.functional as F
sys.path.append("../")
import config
from model_training import training_data_loader
# -
# device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
cpu_device = torch.device('cpu')
# +
TRAINING_DIRS_1 = [
'31_03_21__318__3or4_people/1/006__11_44_59',
'31_03_21__318__3or4_people/1/007__11_48_59',
'31_03_21__318__3or4_people/1/008__11_52_59',
'31_03_21__318__3or4_people/1/009__11_57_00',
]
TRAINING_DIRS_2 = [
'31_03_21__318__3or4_people/2/000__14_15_19',
'31_03_21__318__3or4_people/2/001__14_19_19',
'31_03_21__318__3or4_people/2/002__14_23_19',
'31_03_21__318__3or4_people/2/003__14_27_20',
'31_03_21__318__3or4_people/2/004__14_31_20',
'31_03_21__318__3or4_people/2/005__14_35_20',
'31_03_21__318__3or4_people/2/006__14_39_20',
'31_03_21__318__3or4_people/2/007__14_43_20',
'31_03_21__318__3or4_people/2/008__14_47_20',
'31_03_21__318__3or4_people/2/009__14_51_20',
'31_03_21__318__3or4_people/2/010__14_55_20',
'31_03_21__318__3or4_people/2/011__14_59_20',
'31_03_21__318__3or4_people/2/012__15_03_21',
]
VALIDATION_DIRS_1 = [
'31_03_21__318__3or4_people/2/013__15_07_21',
'31_03_21__318__3or4_people/2/014__15_11_21',
'31_03_21__318__3or4_people/2/015__15_15_21',
'31_03_21__318__3or4_people/2/016__15_19_21',
]
#_training_data_1 = training_data_loader.load_data_for_labeled_batches(labeled_batch_dirs=TRAINING_DIRS_1)
_training_data_2 = training_data_loader.load_data_for_labeled_batches(labeled_batch_dirs=TRAINING_DIRS_2)
_validation_data_1 = training_data_loader.load_data_for_labeled_batches(labeled_batch_dirs=VALIDATION_DIRS_1)
augmented_data_training = training_data_loader.AugmentedBatchesTrainingData()
#augmented_data_training.add_training_batch(_training_data_1)
augmented_data_training.add_training_batch(_training_data_2)
augmented_data_validation = training_data_loader.AugmentedBatchesTrainingData()
augmented_data_validation.add_training_batch(_validation_data_1)
# -
augmented_data_training.print_stats()
augmented_data_validation.print_stats()
# +
MAX_PEOPLE_TO_COUNT = 4
l1_out_features = 128
conv2_out_channels = 32
l1_in_features = config.IR_CAMERA_RESOLUTION_X * config.IR_CAMERA_RESOLUTION_Y // ((2*2)**2) \
* conv2_out_channels
class ModelSingleFrameCnn(nn.Module):
def __init__(self):
super().__init__()
self.conv1 = nn.Conv2d(in_channels=1, out_channels=16, kernel_size=(3,3), stride=1, padding=1)
self.conv2 = nn.Conv2d(in_channels=16, out_channels=conv2_out_channels, kernel_size=3, stride=1, padding=1)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, kernel_size=2, stride=2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, kernel_size=2, stride=2)
x = x.view(-1, l1_in_features)
return x
class ModelSingleFrame1(nn.Module):
def __init__(self, cnn_model):
super(ModelSingleFrame1, self).__init__()
self.cnn_model = cnn_model
self.l1 = nn.Linear(in_features=l1_in_features, out_features=l1_out_features)
self.l2 = nn.Linear(in_features=l1_out_features, out_features=MAX_PEOPLE_TO_COUNT+1)
def forward(self, x):
x = self.cnn_model(x)
x = F.relu(self.l1(x))
x = F.dropout(x, p=0.2)
x = F.log_softmax(self.l2(x), dim=1)
return x
cnn_model = ModelSingleFrameCnn().double().to(device)
model = ModelSingleFrame1(cnn_model).double().to(device)
# -
l1_in_features
# +
from model_training.training_data_loader import AugmentedBatchesTrainingData
class IrPersonsDataset(torch.utils.data.Dataset):
def __init__(self, augmented_data: AugmentedBatchesTrainingData, transform=None):
self.augmented_data = AugmentedBatchesTrainingData
self.transform = transform
self._index_to_batch_and_subindex_map = {}
i = 0
for batch in augmented_data.batches:
for j in range(len(batch.raw_ir_data)):
self._index_to_batch_and_subindex_map[i] = (batch, j)
i += 1
def __len__(self):
return len (self._index_to_batch_and_subindex_map)
def __getitem__(self, idx):
if torch.is_tensor(idx):
raise Exception("Not supported")
#idx = idx.tolist()
batch, subindex = self._index_to_batch_and_subindex_map[idx]
frame = batch.normalized_ir_data[subindex][np.newaxis, :, :]
return frame, len(batch.centre_points[subindex])
# +
training_dataset = IrPersonsDataset(augmented_data_training)
validation_dataset = IrPersonsDataset(augmented_data_validation)
# it makes no sense to split all data, as most of the frames are almost identical
# training_dataset, validation_dataset = torch.utils.data.random_split(all_data_dataset, [training_data_len, validation_data_len])
trainloader = torch.utils.data.DataLoader(training_dataset, batch_size=64, shuffle=True)
valloader = torch.utils.data.DataLoader(validation_dataset, batch_size=1, shuffle=True)
print(len(trainloader))
print(len(valloader))
# +
dataiter = iter(trainloader)
ir_frmaes, labels = dataiter.next()
print(ir_frmaes.shape)
print(labels.shape)
ir_frame_normalized_0 = ir_frmaes[0].numpy().squeeze()
plt.imshow(ir_frame_normalized_0)
print(f'Persons: {labels[0]}')
#print(ir_frame_normalized_0)
#print(np.min(ir_frame_normalized_0))
# -
# +
criterion = nn.NLLLoss()
images, labels = next(iter(trainloader))
print(images.shape)
logps = model(images) #log probabilities
loss = criterion(logps, labels) #calculate the NLL loss
# -
# +
optimizer = optim.SGD(model.parameters(), lr=0.003, momentum=0.9)
time0 = time.time()
epochs = 10
for e in range(epochs):
running_loss = 0
for images, labels in trainloader:
# Training pass
optimizer.zero_grad()
output = model(images)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
print("Epoch {} - Training loss: {}".format(e, running_loss/len(trainloader)))
print("\nTraining Time (in minutes) =",(time.time()-time0)/60)
# -
# +
images, labels = next(iter(valloader))
with torch.no_grad():
logps = model(images)
ps = torch.exp(logps)
probab = list(ps.numpy()[0])
plt.imshow(images[0].numpy().squeeze())
print("Predicted number of persons =", probab.index(max(probab)))
print(probab)
# +
correct_count = 0
tested_frames = 0
number_of_frames_with_n_persons = {}
number_of_frames_with_n_persons_predicted_correctly = {}
for frame, labels in valloader:
for i in range(len(labels)):
with torch.no_grad():
logps = model(frame)
ps = torch.exp(logps)
probab = list(ps.numpy()[0])
pred_label = probab.index(max(probab))
true_label = labels.numpy()[i]
number_of_frames_with_n_persons[pred_label] = \
number_of_frames_with_n_persons.get(pred_label, 0) + 1
if true_label == pred_label:
correct_count += 1
number_of_frames_with_n_persons_predicted_correctly[pred_label] = \
number_of_frames_with_n_persons_predicted_correctly.get(pred_label, 0) + 1
tested_frames += 1
print(f"Number of tested frames: {tested_frames}")
print(f"Model Accuracy = {correct_count / tested_frames}")
print('Predicted:\n' + '\n'.join([f' {count} frames with {no} persons' for no, count in number_of_frames_with_n_persons.items()]))
print('Predicted correctly:\n' + '\n'.join([f' {count} frames with {no} persons' for no, count in number_of_frames_with_n_persons_predicted_correctly.items()]))
# -
TIME_STEPS = 32
# +
from model_training.training_data_loader import AugmentedBatchesTrainingData
class IrPersonsDatasetSeries(torch.utils.data.Dataset):
def __init__(self, augmented_data: AugmentedBatchesTrainingData, transform=None):
self.augmented_data = AugmentedBatchesTrainingData
self.transform = transform
self._index_to_batch_and_subindex_map = {}
i = 0
for batch in augmented_data.batches:
for j in range(len(batch.raw_ir_data)):
self._index_to_batch_and_subindex_map[i] = (batch, j)
i += 1
def __len__(self):
return (len(self._index_to_batch_and_subindex_map) - TIME_STEPS) // TIME_STEPS -1
def __getitem__(self, idx):
# TIME_STEPS consecutive frames
real_frame_idx = [idx*TIME_STEPS + i for i in range(TIME_STEPS)]
frames = [self.get_absolute_frame_data(i) for i in real_frame_idx]
label = self.get_absolute_frame_label(real_frame_idx[-1]) # ???
frames_4d = np.stack(frames, axis = 0)
#print(frames_4d.shape)
return frames_4d, label
def get_absolute_frame_data(self, idx):
batch, subindex = self._index_to_batch_and_subindex_map[idx]
frame = batch.normalized_ir_data[subindex][np.newaxis, :, :]
#print(frame.shape)
return frame
def get_absolute_frame_label(self, idx):
batch, subindex = self._index_to_batch_and_subindex_map[idx]
frame = batch.normalized_ir_data[subindex][np.newaxis, :, :]
return len(batch.centre_points[subindex])
training_dataset_series = IrPersonsDatasetSeries(augmented_data_training)
trainloader_series = torch.utils.data.DataLoader(training_dataset_series, batch_size=16, shuffle=True)
validation_dataset_series = IrPersonsDatasetSeries(augmented_data_training)
valloader_series = torch.utils.data.DataLoader(validation_dataset_series, batch_size=1, shuffle=True)
# criterion = nn.NLLLoss()
# images, labels = next(iter(trainloader))
# print(images.shape)
# print(labels)
# logps = model(images) #log probabilities
# print(logps)
# loss = criterion(logps, labels) #calculate the NLL loss
# -
# +
#torch.randn([5, 32, 60, 60]).shape
# -
# +
dataiter = iter(trainloader_series)
frames_series, labels = dataiter.next()
plt.imshow(frames_series[0, 0, 0])
frames_series.shape # [batch_size, timesteps, C, H, W]
# +
# cnn_out = cnn_model(frames)
# cnn_out.shape # [batch_size, outputs]
# -
# +
class CnnLstmModel(nn.Module):
def __init__(self, pretrained_cnn_model):
super(CnnLstmModel, self).__init__()
self.pretrained_cnn_model = pretrained_cnn_model
self.lstm = nn.LSTM(l1_in_features, hidden_size=l1_out_features, num_layers=2, batch_first=True)
# move l1 to cnn? Or does lstm behaves as l1? Or add it here
self.l1 = nn.Linear(in_features=l1_out_features, out_features=l1_out_features)
self.l2 = nn.Linear(in_features=l1_out_features, out_features=MAX_PEOPLE_TO_COUNT+1)
def forward(self, x):
batch_size, timesteps, C, H, W = x.size()
c_in = x.view(batch_size * timesteps, C, H, W)
x = self.pretrained_cnn_model(c_in)
lstm_in = x.view(batch_size, timesteps, -1)
lstm_out, hidden = self.lstm(lstm_in)
#print(lstm_out.shape)
lstm_out_last_frame = lstm_out[:,-1,:]
#print(lstm_out_last_frame.shape)
x = F.relu(self.l1(lstm_out_last_frame)) # to enable l1 also
# x = F.relu(lstm_out_last_frame)
x = F.dropout(x, p=0.2) # ???
x = F.log_softmax(self.l2(x), dim=1)
return x
cnn_lstm_model = CnnLstmModel(pretrained_cnn_model=cnn_model).double()
cnn_lstm_model
frames_series, labels = iter(trainloader_series).next()
print(frames_series.shape)
out = cnn_lstm_model(frames_series)
out.shape
out[0, -1]
# -
# +
criterion = nn.NLLLoss()
images_series, labels = next(iter(trainloader_series))
print(images_series.shape)
logps = cnn_lstm_model(images_series) #log probabilities
loss = criterion(logps, labels) #calculate the NLL loss
# -
# +
optimizer = optim.SGD(model.parameters(), lr=0.003, momentum=0.9)
time0 = time.time()
epochs = 10
for e in range(epochs):
running_loss = 0
for frame_series, labels in trainloader_series:
# Training pass
optimizer.zero_grad()
output = cnn_lstm_model(frame_series)
loss = criterion(output, labels)
loss.backward()
optimizer.step()
running_loss += loss.item()
else:
print("Epoch {} - Training loss: {}".format(e, running_loss/len(trainloader)))
print("\nTraining Time (in minutes) =",(time.time()-time0)/60)
# -
# +
correct_count = 0
tested_frames = 0
number_of_frames_with_n_persons = {}
number_of_frames_with_n_persons_predicted_correctly = {}
for frame_series, labels in valloader_series:
for i in range(len(labels)):
with torch.no_grad():
logps = cnn_lstm_model(frame_series)
ps = torch.exp(logps)
probab = list(ps.numpy()[0])
pred_label = probab.index(max(probab))
true_label = labels.numpy()[i]
number_of_frames_with_n_persons[pred_label] = \
number_of_frames_with_n_persons.get(pred_label, 0) + 1
if true_label == pred_label:
correct_count += 1
number_of_frames_with_n_persons_predicted_correctly[pred_label] = \
number_of_frames_with_n_persons_predicted_correctly.get(pred_label, 0) + 1
tested_frames += 1
print(f'{correct_count}/{tested_frames}')
print(f"Number of tested frames: {tested_frames}")
print(f"Model Accuracy = {correct_count / tested_frames}")
print('Predicted:\n' + '\n'.join([f' {count} frames with {no} persons' for no, count in number_of_frames_with_n_persons.items()]))
print('Predicted correctly:\n' + '\n'.join([f' {count} frames with {no} persons' for no, count in number_of_frames_with_n_persons_predicted_correctly.items()]))
| data_processing/notebooks/obsoleted/E_lstm_classification_random_batches.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Linked List Nth to Last Node
#
# ## Problem Statement
# Write a function that takes a head node and an integer value **n** and then returns the nth to last node in the linked list. For example, given:
class Node:
def __init__(self, value):
self.value = value
self.nextnode = None
# **Example Input and Output:**
# +
a = Node(1)
b = Node(2)
c = Node(3)
d = Node(4)
e = Node(5)
a.nextnode = b
b.nextnode = c
c.nextnode = d
d.nextnode = e
# This would return the node d with a value of 4, because its the 2nd to last node.
target_node = nth_to_last_node(2, a)
# -
target_node.value
# ## Solution
# Fill out your solution below:
def nth_to_last_node(n, head):
pass
# # Test Your Solution
# +
"""
RUN THIS CELL TO TEST YOUR SOLUTION AGAINST A TEST CASE
PLEASE NOTE THIS IS JUST ONE CASE
"""
from nose.tools import assert_equal
a = Node(1)
b = Node(2)
c = Node(3)
d = Node(4)
e = Node(5)
a.nextnode = b
b.nextnode = c
c.nextnode = d
d.nextnode = e
####
class TestNLast(object):
def test(self,sol):
assert_equal(sol(2,a),d)
print 'ALL TEST CASES PASSED'
# Run tests
t = TestNLast()
t.test(nth_to_last_node)
# -
# ## Good Job!
| code/algorithms/course_udemy_1/Linked Lists/Problems/.ipynb_checkpoints/Linked List Nth to Last Node -checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
a = tf.constant(2)
b = tf.constant(3)
with tf.Session() as sess:
print('a=2, b=3')
print('Addition with constants: {}'.format(sess.run(a+b)))
print('Multiplication with constants: {}'.format(sess.run(a*b)))
a = tf.placeholder(tf.int16)
b = tf.placeholder(tf.int16)
add = tf.add(a, b)
mul = tf.mul(a, b)
with tf.Session() as sess:
print('Addition with variables: {}'.format(sess.run(add, feed_dict={a: 2, b:3})))
print('Multiplication with variables: {}'.format(sess.run(mul, feed_dict={a: 2, b:3})))
matrix1 = tf.constant([[3., 3.]])
matrix2 = tf.constant([[2.], [2.]])
product = tf.matmul(matrix1, matrix2)
with tf.Session() as sess:
result = sess.run(product)
print(result)
| tensorflow/tensorflow-1b.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="KRtjic_f9hZF"
# !pip install otter-grader==1.1.6
from google.colab import drive
drive.mount('/content/gdrive')
# %cd /content/gdrive/MyDrive/'Colab Notebooks'/Colab-data-8/hw/hw03
# + deletable=false editable=false id="TozrZ5RQ9SUR"
# Initialize Otter
import otter
grader = otter.Notebook()
# + [markdown] id="i3XeozBN9SUU"
# # Homework 3: Table Manipulation and Visualization
# + [markdown] id="vEdLa0GR9SUU"
# **Reading**:
# * [Visualization](https://www.inferentialthinking.com/chapters/07/visualization.html)
# + [markdown] id="OUYAicC99SUV"
# Please complete this notebook by filling in the cells provided.
#
# **Throughout this homework and all future ones, please be sure to not re-assign variables throughout the notebook!** For example, if you use `max_temperature` in your answer to one question, do not reassign it later on. Moreover, please be sure to only put your written answers in the provided cells.
# + id="EcNKp8D-9SUV"
# Don't change this cell; just run it.
import numpy as np
from datascience import *
# These lines do some fancy plotting magic.\n",
import matplotlib
# %matplotlib inline
import matplotlib.pyplot as plots
plots.style.use('fivethirtyeight')
# + [markdown] id="1_rxOzqM9SUW"
# ## 1. Unemployment
#
# + [markdown] id="YrPwk0bD9SUW"
# The Federal Reserve Bank of St. Louis publishes data about jobs in the US. Below, we've loaded data on unemployment in the United States. There are many ways of defining unemployment, and our dataset includes two notions of the unemployment rate:
#
# 1. Among people who are able to work and are looking for a full-time job, the percentage who can't find a job. This is called the Non-Employment Index, or NEI.
# 2. Among people who are able to work and are looking for a full-time job, the percentage who can't find any job *or* are only working at a part-time job. The latter group is called "Part-Time for Economic Reasons", so the acronym for this index is NEI-PTER. (Economists are great at marketing.)
#
# The source of the data is [here](https://fred.stlouisfed.org/categories/33509).
# + [markdown] deletable=false editable=false id="f6TzKxlr9SUX"
# **Question 1.** The data are in a CSV file called `unemployment.csv`. Load that file into a table called `unemployment`.
#
# <!--
# BEGIN QUESTION
# name: q1_1
# -->
# + id="MTCU0yMz9SUX"
unemployment = ...
unemployment
# + deletable=false editable=false id="66XFQl9P9SUX"
grader.check("q1_1")
# + [markdown] deletable=false editable=false id="GweCSP4i9SUX"
# **Question 2.** Sort the data in descending order by NEI, naming the sorted table `by_nei`. Create another table called `by_nei_pter` that's sorted in descending order by NEI-PTER instead.
#
# <!--
# BEGIN QUESTION
# name: q1_2
# -->
# + id="guKI6NWK9SUY"
by_nei = ...
by_nei_pter = ...
# + deletable=false editable=false id="kCnOS_Pb9SUY"
grader.check("q1_2")
# + [markdown] deletable=false editable=false id="_jK_Ijjj9SUY"
# **Question 3.** Use `take` to make a table containing the data for the 10 quarters when NEI was greatest. Call that table `greatest_nei`.
#
# `greatest_nei` should be sorted in descending order of `NEI`. Note that each row of `unemployment` represents a quarter.
#
# <!--
# BEGIN QUESTION
# name: q1_3
# -->
# + id="zj9ZAdeo9SUZ"
greatest_nei = ...
greatest_nei
# + id="tFL2phQg9SUZ"
greatest_nei.take(0).column(0).item(0) == '2009-10-01'
# + [markdown] deletable=false editable=false id="PjczPeYa9SUZ"
# **Question 4.** It's believed that many people became PTER (recall: "Part-Time for Economic Reasons") in the "Great Recession" of 2008-2009. NEI-PTER is the percentage of people who are unemployed (and counted in the NEI) plus the percentage of people who are PTER. Compute an array containing the percentage of people who were PTER in each quarter. (The first element of the array should correspond to the first row of `unemployment`, and so on.)
#
# *Note:* Use the original `unemployment` table for this.
#
# <!--
# BEGIN QUESTION
# name: q1_4
# -->
# + id="DfcEFOb09SUZ"
pter = ...
pter
# + deletable=false editable=false id="PpDdzxhU9SUa"
grader.check("q1_4")
# + [markdown] deletable=false editable=false id="pqKD_8gZ9SUa"
# **Question 5.** Add `pter` as a column to `unemployment` (named "PTER") and sort the resulting table by that column in descending order. Call the table `by_pter`.
#
# Try to do this with a single line of code, if you can.
#
# <!--
# BEGIN QUESTION
# name: q1_5
# -->
# + id="Lbqntl0N9SUa"
by_pter = ...
by_pter
# + deletable=false editable=false id="sR6U7oxN9SUa"
grader.check("q1_5")
# + [markdown] deletable=false editable=false id="Pwag-VtI9SUb"
# **Question 6.**
#
# Create a line plot of the PTER over time.
#
# To do this, create a new table called `pter_over_time` that adds the `year` array and the `pter` array to the `unemployment` table. Label these columns `Year` and `PTER`. Then, generate a line plot using one of the table methods you've learned in class.
#
# <!--
# BEGIN QUESTION
# name: q1_6
# -->
# + id="JGdF7jk59SUb"
year = 1994 + np.arange(by_pter.num_rows)/4
pter_over_time = ...
...
# + deletable=false editable=false id="oMFv4OrZ9SUb"
grader.check("q1_6")
# + [markdown] deletable=false editable=false id="bkIUb9hp9SUb"
# **Question 7.** Were PTER rates high during the Great Recession (that is to say, were PTER rates particularly high in the years 2008 through 2011)? Assign highPTER to `True` if you think PTER rates were high in this period, and `False` if you think they weren't.
#
# <!--
# BEGIN QUESTION
# name: q1_7
# -->
# + id="PDvFyPa29SUc"
highPTER = ...
# + deletable=false editable=false id="W9oGxaPp9SUc"
grader.check("q1_7")
# + [markdown] id="Oe1_3NyC9SUc"
# ## 2. <NAME>
#
# + [markdown] id="SiPdbGbI9SUc"
# The following table gives census-based population estimates for each state on both July 1, 2015 and July 1, 2016. The last four columns describe the components of the estimated change in population during this time interval. **For all questions below, assume that the word "states" refers to all 52 rows including Puerto Rico & the District of Columbia.**
#
# The data was taken from [here](http://www2.census.gov/programs-surveys/popest/datasets/2010-2016/national/totals/nst-est2016-alldata.csv).
#
# If you want to read more about the different column descriptions, click [here](http://www2.census.gov/programs-surveys/popest/datasets/2010-2015/national/totals/nst-est2015-alldata.pdf)!
#
# The raw data is a bit messy - run the cell below to clean the table and make it easier to work with.
# + id="5T9_Trnq9SUd"
# Don't change this cell; just run it.
pop = Table.read_table('nst-est2016-alldata.csv').where('SUMLEV', 40).select([1, 4, 12, 13, 27, 34, 62, 69])
pop = pop.relabeled('POPESTIMATE2015', '2015').relabeled('POPESTIMATE2016', '2016')
pop = pop.relabeled('BIRTHS2016', 'BIRTHS').relabeled('DEATHS2016', 'DEATHS')
pop = pop.relabeled('NETMIG2016', 'MIGRATION').relabeled('RESIDUAL2016', 'OTHER')
pop = pop.with_columns("REGION", np.array([int(region) if region != "X" else 0 for region in pop.column("REGION")]))
pop.set_format([2, 3, 4, 5, 6, 7], NumberFormatter(decimals=0)).show(5)
# + [markdown] deletable=false editable=false id="SxSYWpRp9SUd"
# **Question 1.** Assign `us_birth_rate` to the total US annual birth rate during this time interval. The annual birth rate for a year-long period is the total number of births in that period as a proportion of the population size at the start of the time period.
#
# **Hint:** Which year corresponds to the start of the time period?
#
# <!--
# BEGIN QUESTION
# name: q2_1
# -->
# + id="PB8_azct9SUe"
us_birth_rate = ...
us_birth_rate
# + deletable=false editable=false id="JjZi36UO9SUe"
grader.check("q2_1")
# + [markdown] deletable=false editable=false id="3k7-XoC89SUe"
# **Question 2.** Assign `movers` to the number of states for which the **absolute value** of the **annual rate of migration** was higher than 1%. The annual rate of migration for a year-long period is the net number of migrations (in and out) as a proportion of the population size at the start of the period. The `MIGRATION` column contains estimated annual net migration counts by state.
#
# <!--
# BEGIN QUESTION
# name: q2_2
# -->
# + id="jvoxhSAD9SUe"
migration_rates = ...
movers = ...
movers
# + deletable=false editable=false id="306z6icy9SUf"
grader.check("q2_2")
# + [markdown] deletable=false editable=false id="uN6JZzaY9SUf"
# **Question 3.** Assign `west_births` to the total number of births that occurred in region 4 (the Western US).
#
# **Hint:** Make sure you double check the type of the values in the region column, and appropriately filter (i.e. the types must match!).
#
# <!--
# BEGIN QUESTION
# name: q2_3
# -->
# + id="k-TD1Hoo9SUf"
west_births = ...
west_births
# + deletable=false editable=false id="0y6j3LMg9SUf"
grader.check("q2_3")
# + [markdown] deletable=false editable=false id="RltVQH0l9SUf"
# **Question 4.** Assign `less_than_west_births` to the number of states that had a total population in 2016 that was smaller than the *total number of births in region 4 (the Western US)* during this time interval.
#
# <!--
# BEGIN QUESTION
# name: q2_4
# -->
# + id="3g9eMH789SUg"
less_than_west_births = pop.where('2016', are.below(west_births)).num_rows
less_than_west_births
# + deletable=false editable=false id="XkO3Ldg19SUg"
grader.check("q2_4")
# + [markdown] deletable=false editable=false id="dJue4U-y9SUg"
# **Question 5.**
#
# In the next question, you will be creating a visualization to understand the relationship between birth and death rates. The annual death rate for a year-long period is the total number of deaths in that period as a proportion of the population size at the start of the time period.
#
# What visualization is most appropriate to see if there is an association between birth and death rates during a given time interval?
#
# 1. Line Graph
# <br>
# 2. Scatter Plot
# <br>
# 3. Bar Chart
#
# Assign `visualization` below to the number corresponding to the correct visualization.
#
# <!--
# BEGIN QUESTION
# name: q2_5
# -->
# + id="iIY6mevQ9SUg"
visualization = ...
# + deletable=false editable=false id="UEg6KgMK9SUg"
grader.check("q2_5")
# + [markdown] deletable=false editable=false id="1xE-JZgd9SUg"
# <!-- BEGIN QUESTION -->
#
# **Question 6.**
#
# In the code cell below, create a visualization that will help us determine if there is an association between birth rate and death rate during this time interval. It may be helpful to create an intermediate table here.
#
# <!--
# BEGIN QUESTION
# name: q2_6
# manual: true
# -->
# + export_pdf=true manual_problem_id="birth_death_rates_graph" id="eWQNJTBM9SUh"
# Generate your chart in this cell
...
# + [markdown] deletable=false editable=false id="uaPrlKee9SUh"
# <!-- END QUESTION -->
#
# **Question 7.** `True` or `False`: There is an association between birth rate and death rate during this time interval.
#
# Assign `assoc` to `True` or `False` in the cell below.
#
# <!--
# BEGIN QUESTION
# name: q2_7
# -->
# + id="ynEMOF6f9SUh"
assoc = ...
# + deletable=false editable=false id="EMRw56fm9SUh"
grader.check("q2_7")
# + [markdown] id="67cxLIy39SUh"
# ## 3. Marginal Histograms
#
# + [markdown] id="tHQ7u1Ur9SUh"
# Consider the following scatter plot: 
#
# The axes of the plot represent values of two variables: $x$ and $y$.
# + [markdown] id="tHxv4qTQ9SUi"
# Suppose we have a table called `t` that has two columns in it:
#
# - `x`: a column containing the x-values of the points in the scatter plot
# - `y`: a column containing the y-values of the points in the scatter plot
# + [markdown] id="lnH8dZCt9SUi"
# Below, you are given two histograms, each of which corresponds to either column `x` or column `y`.
# + [markdown] id="T3Ymw6cU9SUi"
# **Histogram A:** 
# **Histogram B:** 
# + [markdown] deletable=false editable=false id="JI0Q2ar29SUi"
# **Question 1.** Suppose we run `t.hist('x')`. Which histogram does this code produce? Assign `histogram_column_x` to either 1 or 2.
#
# 1. Histogram A
# 2. Histogram B
#
# <!--
# BEGIN QUESTION
# name: q3_1
# manual: false
# -->
# + id="LgCj_AYa9SUi"
histogram_column_x = ...
# + deletable=false editable=false id="PQGwDAs39SUj"
grader.check("q3_1")
# + [markdown] deletable=false editable=false id="DbWY8faH9SUj"
# <!-- BEGIN QUESTION -->
#
# **Question 2.** State at least one reason why you chose the histogram from Question 1. Make sure to indicate which histogram you selected (ex: "I chose histogram A because ...").
#
# <!--
# BEGIN QUESTION
# name: q3_2
# manual: true
# -->
# + [markdown] id="QRj57fCO9SUj"
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false id="NaVevlYT9SUj"
# <!-- END QUESTION -->
#
# **Question 3.** Suppose we run `t.hist('y')`. Which histogram does this code produce? `Assign histogram_column_y` to either 1 or 2.
#
# 1. Histogram A
# 2. Histogram B
#
# <!--
# BEGIN QUESTION
# name: q3_3
# manual: false
# -->
# + id="LMtf-8t39SUj"
histogram_column_y = ...
# + deletable=false editable=false id="r1lxi3o39SUk"
grader.check("q3_3")
# + [markdown] deletable=false editable=false id="Zdq1rJWp9SUk"
# <!-- BEGIN QUESTION -->
#
# **Question 4.** State at least one reason why you chose the histogram from Question 3. Make sure to indicate which histogram you selected (ex: "I chose histogram A because ...").
#
# <!--
# BEGIN QUESTION
# name: q3_4
# manual: true
# -->
# + [markdown] id="E_92vkQ99SUk"
# _Type your answer here, replacing this text._
# + [markdown] id="L3LbTCcb9SUk"
# <!-- END QUESTION -->
#
#
#
# ## 4. Uber
#
# + [markdown] id="lqLouvHU9SUk"
# Below we load tables containing 200,000 weekday Uber rides in the Manila, Philippines, and Boston, Massachusetts metropolitan areas from the [Uber Movement](https://movement.uber.com) project. The `sourceid` and `dstid` columns contain codes corresponding to start and end locations of each ride. The `hod` column contains codes corresponding to the hour of the day the ride took place. The `ride time` column contains the length of the ride, in minutes.
# + for_question_type="solution" id="orVJdAV29SUk"
boston = Table.read_table("boston.csv")
manila = Table.read_table("manila.csv")
print("Boston Table")
boston.show(4)
print("Manila Table")
manila.show(4)
# + [markdown] deletable=false editable=false id="bGBA_z_w9SUk"
# <!-- BEGIN QUESTION -->
#
# **Question 1.** Produce histograms of all ride times in Boston using the given bins.
#
# <!--
# BEGIN QUESTION
# name: q4_1
# manual: true
# -->
#
# + export_pdf=true manual_problem_id="uber_1" id="-jt6mYzh9SUl"
equal_bins = np.arange(0, 120, 5)
...
# + [markdown] deletable=false editable=false id="hZGcF82w9SUl"
# <!-- END QUESTION -->
#
# <!-- BEGIN QUESTION -->
#
# **Question 2.** Now, produce histograms of all ride times in Manila using the given bins.
#
# <!--
# BEGIN QUESTION
# name: q4_2
# manual: true
# -->
# + export_pdf=true id="IbdpuiPv9SUl"
...
# Don't delete the following line!
plots.ylim(0, 0.05)
# + [markdown] deletable=false editable=false id="v0MYkEk39SUl"
# <!-- END QUESTION -->
#
# **Question 3.** Assign `boston_under_10` and `manila_under_10` to the percentage of rides that are less than 10 minutes in their respective metropolitan areas. Use the height variables provided below in order to compute the percentages. Your solution should only use height variables, numbers, and mathematical operations. You should not access the tables boston and manila in any way.
#
# <!--
# BEGIN QUESTION
# name: q4_3
# manual: false
# -->
# + manual_problem_id="uber_2" id="32QkZWjr9SUl"
boston_under_5_height = 1.2
manila_under_5_height = 0.6
boston_5_to_under_10_height = 3.2
manila_5_to_under_10_height = 1.4
boston_under_10 = ...
manila_under_10 = ...
# + deletable=false editable=false id="MecWFSqP9SUl"
grader.check("q4_3")
# + [markdown] deletable=false editable=false id="r0fdHWlO9SUm"
# **Question 4.** Let's take a closer look at the distribution of ride times in Manila. Assign `manila_median_bin` to an integer (1, 2, 3, or 4) that corresponds to the bin that contains the median time
#
# 1: 0-15 minutes
# 2: 15-40 minutes
# 3: 40-60 minutes
# 4: 60-80 minutes
#
# *Hint:* The median of a sorted list has half of the list elements to its left, and half to its right
#
# <!--
# BEGIN QUESTION
# name: q4_4
# manual: false
# -->
# + id="xpGs_wxu9SUm"
manila_median_bin = ...
manila_median_bin
# + deletable=false editable=false id="9Ve_WetS9SUm"
grader.check("q4_4")
# + [markdown] deletable=false editable=false id="lrc8NUBm9SUm"
# <!-- BEGIN QUESTION -->
#
# **Question 5.** What is the main difference between the two histograms. What might be causing this?
#
# **Hint:** Try thinking about external factors that may be causing the difference!
#
# <!--
# BEGIN QUESTION
# name: q4_5
# manual: true
# -->
# + [markdown] id="l9SoU9PZ9SUm"
# _Type your answer here, replacing this text._
# + [markdown] deletable=false editable=false id="KMm1Jg9R9SUm"
# ---
#
# To double-check your work, the cell below will rerun all of the autograder tests.
# + deletable=false editable=false id="HJ-lXN_N9SUm"
grader.check_all()
# + [markdown] deletable=false editable=false id="4pZ5l1169SUn"
# ## Submission
#
# Make sure you have run all cells in your notebook in order before running the cell below, so that all images/graphs appear in the output. The cell below will generate a zip file for you to submit. **Please save before exporting!**
# + deletable=false editable=false id="8vZswECa9SUn"
# Save your notebook first, then run this cell to export your submission.
grader.export(pdf=False)
# + [markdown] id="gc5-AH599SUn"
#
| hw/hw03/hw03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Managing software issues
# ### Issues
#
# Code has *bugs*. It also has *features*, things it should do.
#
# A good project has an organised way of managing these. Generally you should use an issue tracker.
# ### Some Issue Trackers
#
# There are lots of good issue trackers.
#
# The most commonly used open source ones are [Trac](http://trac.edgewall.org/) and [Redmine](http://www.redmine.org/).
#
# Cloud based issue trackers include [Lighthouse](http://lighthouseapp.com/) and [GitHub](https://github.com/blog/831-issues-2-0-the-next-generation).
#
# Commercial solutions include [Jira](https://www.atlassian.com/software/jira).
#
# In this course, we'll be using the GitHub issue tracker.
# ### Anatomy of an issue
#
# * Reporter
# * Description
# * Owner
# * Type [Bug, Feature]
# * Component
# * Status
# * Severity
# ### Reporting a Bug
#
# The description should make the bug reproducible:
#
# * Version
# * Steps
#
# If possible, submit a minimal reproducing code fragment.
# ### Owning an issue
#
# * Whoever the issue is assigned to works next.
# * If an issue needs someone else's work, assign it to them.
# ### Status
#
# * Submitted
# * Accepted
# * Underway
# * Blocked
# ### Resolutions
#
# * Resolved
# * Will Not Fix
# * Not reproducible
# * Not a bug (working as intended)
# ### Bug triage
#
# Some organisations use a severity matrix based on:
#
# * Severity [Wrong answer, crash, unusable, workaround, cosmetic...]
# * Frequency [All users, most users, some users...]
# ### The backlog
#
# The list of all the bugs that need to be fixed or
# features that have been requested is called the "backlog".
# ### Development cycles
#
# Development goes in *cycles*.
#
# Cycles range in length from a week to three months.
#
# In a given cycle:
#
# * Decide which features should be implemented
# * Decide which bugs should be fixed
# * Move these issues from the Backlog into the current cycle. (Aka Sprint)
# ### GitHub issues
#
# GitHub doesn't have separate fields for status, component, severity etc.
# Instead, it just has labels, which you can create and delete.
#
# See for example [Jupyter](https://github.com/jupyter/notebook/issues?labels=bug&page=1&state=open)
| ch04packaging/09Issues.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Combining Datasets: Concat and Append
# Some of the most interesting studies of data come from combining different data sources.
# These operations can involve anything from very straightforward concatenation of two different datasets, to more complicated database-style joins and merges that correctly handle any overlaps between the datasets.
# ``Series`` and ``DataFrame``s are built with this type of operation in mind, and Pandas includes functions and methods that make this sort of data wrangling fast and straightforward.
#
# Here we'll take a look at simple concatenation of ``Series`` and ``DataFrame``s with the ``pd.concat`` function; later we'll dive into more sophisticated in-memory merges and joins implemented in Pandas.
#
# We begin with the standard imports:
import pandas as pd
import numpy as np
# For convenience, we'll define this function which creates a ``DataFrame`` of a particular form that will be useful below:
# +
def make_df(cols, ind):
"""Quickly make a DataFrame"""
data = {c: [str(c) + str(i) for i in ind]
for c in cols}
return pd.DataFrame(data, ind)
# example DataFrame
make_df('ABC', range(3))
# -
# ## Recall: Concatenation of NumPy Arrays
#
# Concatenation of ``Series`` and ``DataFrame`` objects is very similar to concatenation of Numpy arrays, which can be done via the ``np.concatenate`` function as discussed in [The Basics of NumPy Arrays](02.02-The-Basics-Of-NumPy-Arrays.ipynb).
# Recall that with it, you can combine the contents of two or more arrays into a single array:
x = [1, 2, 3]
y = [4, 5, 6]
z = [7, 8, 9]
np.concatenate([x, y, z])
# The first argument is a list or tuple of arrays to concatenate.
# Additionally, it takes an ``axis`` keyword that allows you to specify the axis along which the result will be concatenated:
x = [[1, 2],
[3, 4]]
np.concatenate([x, x], axis=1)
# ## Simple Concatenation with ``pd.concat``
# Pandas has a function, ``pd.concat()``, which has a similar syntax to ``np.concatenate`` but contains a number of options that we'll discuss momentarily:
#
# ```python
# # Signature in Pandas v0.18
# pd.concat(objs, axis=0, join='outer', join_axes=None, ignore_index=False,
# keys=None, levels=None, names=None, verify_integrity=False,
# copy=True)
# ```
#
# ``pd.concat()`` can be used for a simple concatenation of ``Series`` or ``DataFrame`` objects, just as ``np.concatenate()`` can be used for simple concatenations of arrays:
ser1 = pd.Series(['A', 'B', 'C'], index=[1, 2, 3])
ser2 = pd.Series(['D', 'E', 'F'], index=[4, 5, 6])
pd.concat([ser1, ser2])
# It also works to concatenate higher-dimensional objects, such as ``DataFrame``s:
df1 = make_df('AB', [1, 2])
df2 = make_df('AB', [3, 4])
df1
df2
pd.concat([df1, df2])
# By default, the concatenation takes place row-wise within the ``DataFrame`` (i.e., ``axis=0``).
# Like ``np.concatenate``, ``pd.concat`` allows specification of an axis along which concatenation will take place.
# Consider the following example:
df3 = make_df('AB', [0, 1])
df4 = make_df('CD', [0, 1])
df3
df4
pd.concat([df3, df4], axis=1)
# ### Duplicate indices
#
# One important difference between ``np.concatenate`` and ``pd.concat`` is that Pandas concatenation *preserves indices*, even if the result will have duplicate indices!
# Consider this simple example:
x = make_df('AB', [0, 1])
y = make_df('AB', [2, 3])
y.index = x.index # make duplicate indices!
x
y
pd.concat([x, y])
# Notice the repeated indices in the result.
# While this is valid within ``DataFrame``s, the outcome is often undesirable.
# ``pd.concat()`` gives us a few ways to handle it.
# #### Catching the repeats as an error
#
# If you'd like to simply verify that the indices in the result of ``pd.concat()`` do not overlap, you can specify the ``verify_integrity`` flag.
# With this set to True, the concatenation will raise an exception if there are duplicate indices.
# Here is an example, where for clarity we'll catch and print the error message:
try:
pd.concat([x, y], verify_integrity=True)
except ValueError as e:
print("ValueError:", e)
# #### Ignoring the index
#
# Sometimes the index itself does not matter, and you would prefer it to simply be ignored.
# This option can be specified using the ``ignore_index`` flag.
# With this set to true, the concatenation will create a new integer index for the resulting ``Series``:
x
y
pd.concat([x, y], ignore_index=True)
# #### Adding MultiIndex keys
#
# Another option is to use the ``keys`` option to specify a label for the data sources; the result will be a hierarchically indexed series containing the data:
pd.concat([x, y], keys=['x', 'y'])
# ### Concatenation with joins
#
# In the simple examples we just looked at, we were mainly concatenating ``DataFrame``s with shared column names.
# In practice, data from different sources might have different sets of column names, and ``pd.concat`` offers several options in this case.
# Consider the concatenation of the following two ``DataFrame``s, which have some (but not all!) columns in common:
df5 = make_df('ABC', [1, 2])
df6 = make_df('BCD', [3, 4])
df5
df6
pd.concat([df5, df6])
# By default, the entries for which no data is available are filled with NA values.
# To change this, we can specify one of several options for the ``join`` and ``join_axes`` parameters of the concatenate function.
# By default, the join is a union of the input columns (``join='outer'``), but we can change this to an intersection of the columns using ``join='inner'``:
pd.concat([df5, df6], join='inner')
# Another option is to directly specify the index of the remaininig colums using the ``join_axes`` argument, which takes a list of index objects.
# Here we'll specify that the returned columns should be the same as those of the first input:
# +
# pd.concat([df5, df6], join_axes=[df5.columns]) # deprecated in version 0.25
df7 = pd.concat([df5, df6], axis=1)
df7 = df7.reindex(df5.index)
df7
# -
# The combination of options of the ``pd.concat`` function allows a wide range of possible behaviors when joining two datasets; keep these in mind as you use these tools for your own data.
# ### The ``append()`` method
#
# Because direct array concatenation is so common, ``Series`` and ``DataFrame`` objects have an ``append`` method that can accomplish the same thing in fewer keystrokes.
# For example, rather than calling ``pd.concat([df1, df2])``, you can simply call ``df1.append(df2)``:
df1
df2
df1.append(df2)
# Keep in mind that unlike the ``append()`` and ``extend()`` methods of Python lists, the ``append()`` method in Pandas does not modify the original object–instead it creates a new object with the combined data.
# It also is not a very efficient method, because it involves creation of a new index *and* data buffer.
# Thus, if you plan to do multiple ``append`` operations, it is generally better to build a list of ``DataFrame``s and pass them all at once to the ``concat()`` function.
#
# In the next section, we'll look at another more powerful approach to combining data from multiple sources, the database-style merges/joins implemented in ``pd.merge``.
# For more information on ``concat()``, ``append()``, and related functionality, see the ["Merge, Join, and Concatenate" section](http://pandas.pydata.org/pandas-docs/stable/merging.html) of the Pandas documentation.
| day3/06. pandas - Concat-And-Append.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# 
# <hr>
# # Research to Production
# ## Long Short Term Memory
# ---------
# ### Introduction
# This notebook explains how to you can use the Research Environment to develop and test a Long Short Term Memory hypothesis, then put the hypothesis in production.
#
# <a href="https://www.quantconnect.com/docs/v2/research-environment/tutorials/research-to-production/long-short-term-memory">Documentation</a>
#
# Recurrent neural networks (RNN) are a powerful tool in deep learning. These models quite accurately mimic how humans process sequencial information and learn. Unlike traditional feedforward neural networks, RNNs have memory. That is, information fed into them persists and the network is able to draw on this to make inferences.
#
# Long Short-term Memory (LSTM) is a type of RNN. Instead of one layer, LSTM cells generally have four, three of which are part of "gates" -- ways to optionally let information through. The three gates are commonly referred to as the forget, input, and output gates. The forget gate layer is where the model decides what information to keep from prior states. At the input gate layer, the model decides which values to update. Finally, the output gate layer is where the final output of the cell state is decided. Essentially, LSTM separately decides what to remember and the rate at which it should update.
#
# <img src="https://cdn.quantconnect.com/i/tu/lstm-cell-2.png">
# <b>An exmaple of a LSTM cell: x is the input data, c is the long-term memory, h is the current state and serve as short-term memory, $\sigma$ and $tanh$ is the non-linear activation function of the gates.</b><br/>
# <i><font font-size="2px">Image source: https://en.wikipedia.org/wiki/Long_short-term_memory#/media/File:LSTM_Cell.svg</font></i>
#
# ### Create Hypothesis
# LSTM models have produced some great results when applied to time-series prediction. One of the central challenges with conventional time-series models is that, despite trying to account for trends or other non-stationary elements, it is almost impossible to truly predict an outlier like a recession, flash crash, liquidity crisis, etc. By having a long memory, LSTM models are better able to capture these difficult trends in the data without suffering from the level of overfitting a conventional model would need in order to capture the same data.
#
# For a very basic application, we're hypothesizing LSTM can offer an accurate prediction in future price.
#
# ### Prerequisites
# If you use Python, you must understand how to work with pandas DataFrames and Series. If you are not familiar with pandas, refer to the <a href="https://pandas.pydata.org/docs/">pandas documentation</a>.
# ### Import Packages
# We'll need to import packages to help with data processing, visualisation and deep learning. Import keras, sklearn, numpy and matplotlib packages by the following:
# +
from keras.layers import LSTM, Dense, Dropout
from keras.models import Sequential
from keras.callbacks import EarlyStopping
from sklearn.preprocessing import MinMaxScaler
import numpy as np
from matplotlib import pyplot as plt
# -
# ### Get Historical Data
# To begin, we retrieve historical data for research.
# +
# Instantiate a QuantBook.
qb = QuantBook()
# Select the desired tickers for research.
assets = "SPY"
# Call the AddEquity method with the tickers, and its corresponding resolution. Then store their Symbols. Resolution.Minute is used by default.
qb.AddEquity(assets, Resolution.Minute)
# Call the History method with qb.Securities.Keys for all tickers, time argument(s), and resolution to request historical data for the symbol.
history = qb.History(qb.Securities.Keys, datetime(2020, 1, 1), datetime(2022, 1, 1), Resolution.Daily)
history
# -
# ### Preparing Data
# We'll have to process our data as well as build the LSTM model before testing the hypothesis. We would scale our data to for better covergence.
# +
# Select the close column and then call the unstack method.
close = history['close'].unstack(level=0)
# Initialize MinMaxScaler to scale the data onto [0,1].
scaler = MinMaxScaler(feature_range = (0, 1))
# Transform our data.
df = pd.DataFrame(scaler.fit_transform(close), index=close.index)
# Select input data
input_ = df.iloc[1:]
# Shift the data for 1-step backward as training output result.
output = df.shift(-1).iloc[:-1]
# Split the data into training and testing sets. In this example, we use the first 80% data for trianing, and the last 20% for testing.
splitter = int(input_.shape[0] * 0.8)
X_train = input_.iloc[:splitter]
X_test = input_.iloc[splitter:]
y_train = output.iloc[:splitter]
y_test = output.iloc[splitter:]
# Build feauture and label sets (using number of steps 60, and feature rank 1).
features_set = []
labels = []
for i in range(60, X_train.shape[0]):
features_set.append(X_train.iloc[i-60:i].values.reshape(-1, 1))
labels.append(y_train.iloc[i])
features_set, labels = np.array(features_set), np.array(labels)
features_set = np.reshape(features_set, (features_set.shape[0], features_set.shape[1], 1))
# -
# ### Build Model
# We construct the LSTM model.
# +
# Build a Sequential keras model.
model = Sequential()
# Create the model infrastructure.
# Add our first LSTM layer - 50 nodes
model.add(LSTM(units = 50, return_sequences=True, input_shape=(features_set.shape[1], 1)))
# Add Dropout layer to avoid overfitting
model.add(Dropout(0.2))
# Add additional layers
model.add(LSTM(units=50, return_sequences=True))
model.add(Dropout(0.2))
model.add(LSTM(units=50))
model.add(Dropout(0.2))
model.add(Dense(units = 5))
model.add(Dense(units = 1))
# Compile the model. We use Adam as optimizer for adpative step size and MSE as loss function since it is continuous data.
model.compile(optimizer = 'adam', loss = 'mean_squared_error', metrics=['mae', 'acc'])
# Set early stopping callback method.
callback = EarlyStopping(monitor='loss', patience=3, verbose=1, restore_best_weights=True)
# Display the model structure.
model.summary()
# -
# Fit the model to our data, running 20 training epochs.
model.fit(features_set, labels, epochs = 20, batch_size = 100, callbacks=[callback])
# Note that different training session's results will not be the same since the batch is randomly selected.
# ### Test the Hypothesis
# We would test the performance of this ML model to see if it could predict 1-step forward price precisely. To do so, we would compare the predicted and actual prices.
# +
# Get testing set features for input.
test_features = []
for i in range(60, X_test.shape[0]):
test_features.append(X_test.iloc[i-60:i].values.reshape(-1, 1))
test_features = np.array(test_features)
test_features = np.reshape(test_features, (test_features.shape[0], test_features.shape[1], 1))
# Make predictions.
predictions = model.predict(test_features)
# Transform predictions back to original data-scale.
predictions = scaler.inverse_transform(predictions)
actual = scaler.inverse_transform(y_test.values)
# Plot the results
plt.figure(figsize=(15, 10))
plt.plot(actual[60:], color='blue', label='Actual')
plt.plot(predictions , color='red', label='Prediction')
plt.title('Price vs Predicted Price ')
plt.legend()
plt.show()
| 03 Research Environment/03 Tutorials/07 Research to Production/08 Long Short-Term Memory/00 Long Short Term Memory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# https://leetcode.com/problems/trim-a-binary-search-tree/
# Given the root of a binary search tree and 2 numbers min and max, trim the tree such that all the numbers in the new tree are between min and max (inclusive). The resulting tree should still be a valid binary search tree. So, if we get this tree as input:
# +
# 8
# / \
#
# -
# When `node.val > high`, we know that the trimmed binary tree must occur to the left of the node. Similarly, when `node.val < low`, the trimmed binary tree occurs to the right of the node. Otherwise, we will trim both sides of the tree.
#
# **Solution (DFS):**
# Definition for a binary tree node.
# class TreeNode(object):
# def __init__(self, val=0, left=None, right=None):
# self.val = val
# self.left = left
# self.right = right
class Solution(object):
def trimBST(self, root, low, high):
"""
:type root: TreeNode
:type low: int
:type high: int
:rtype: TreeNode
"""
def trim(node):
if not node:
return None
elif node.val > high:
return trim(node.left)
elif node.val < low:
return trim(node.right)
else:
node.left = trim(node.left)
node.right = trim(node.right)
return node
return trim(root)
# Time Complexity: $O(N)$, where $N$ is the total number of nodes in the given tree. We visit each node at most once.
#
# Space Complexity: $O(N)$, Even though we don't explicitly use any additional memory, the call stack of our recursion could be as large as the number of nodes in the worst case.
#
#
| notebooks/Trees/6. Trim a Binary Search Tree Interview Problem.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fashion MNIST Classification: 3 Layer CNN vs SVM
# ### Comparing the accuracy of a 3 layer Convolutional Neural Network and a Support Vector Machine
# #### By <NAME>
# This is a dataset of 60,000 28x28 grayscale images of 10 fashion categories, along with a test set of 10,000 images.
# One of the goals of this project is to show the efficiency and effectiveness of neural networks over hyperplane based classification models.
#
# The follow labels indicate the type of fashion item:
# - 0: T-shirt/top
# - 1: Trouser
# - 2: Pullover
# - 3: Dress
# - 4: Coat
# - 5: Sandal
# - 6: Shirt
# - 7: Sneaker
# - 8: Bag
# - 9: Ankle boot
#
# The following resources were used to help build the model and plot graphs:
# 1. https://www.tensorflow.org/tutorials/keras/classification
# 2. CNN Jupyter Notebook by <NAME>
# 3. My previous hw for making plots
# 4. https://towardsdatascience.com/a-comprehensive-guide-to-convolutional-neural-networks-the-eli5-way-3bd2b1164a53
# ## Importing Libraries
#
# I will be utilizing 7 libaries: Pandas for initial dataframes, Numpy for array reshaping, SKlearn for our support vector machine model, Matplotlib plot plotting, and Tensorflow Keras for our primary CNN.
# +
import pandas as pd
import numpy as np
import sklearn as sk
import matplotlib.pyplot as plt
import tensorflow as tf
from sklearn import svm
from mpl_toolkits.axes_grid1 import ImageGrid
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Dense,Flatten,MaxPooling2D
from keras.layers import Conv2D
from tensorflow import keras
from tensorflow.keras import datasets, layers, models
from sklearn import metrics
# -
# ## Loading Data
# Load the data from the keras library, and view the shape of it.
#load data
fash = tf.keras.datasets.fashion_mnist.load_data()
#load training and testing data
(x_train, y_train), (x_test, y_test) = fash
# +
#What's the shape of our data?
print("x_train shape: ", x_train.shape)
print("y_train: ", y_train.shape)
print("x_test: ", x_test.shape)
print("y_test: ", y_test.shape)
m_train = x_train.shape[0]
m_test = x_test.shape[0]
num_px = x_train.shape[1]
print ("Number of training examples: ", m_train)
print ("Number of testing examples: ",m_test)
print ("Height and Width of each image: ", num_px)
# -
# ## Visualizing the Data
#
# Here we will visualize what the images look like with their attached label.
# +
#Function source from <NAME>
fig, axes = plt.subplots(4, 4, figsize=(10,10))
for i in range(4):
for j in range(4):
axes[i,j].imshow(x_train[i*4 + j, :, :], cmap='binary')
axes[i,j].set_title("Label: " + str(y_train[i*4+j]))
axes[i,j].axis('off')
plt.show()
# -
# ## SVM Classification
#
# We will first predict our model accuracy with a Support Vector Machine. When comparing minimal classes in a SVM, the model operates pretty quickly, however when running 9 features for classification on tens of thousands of images, the runtime increases quite a bit.
#shape data to fit svm model
ntrain, nx, ny = x_train.shape
ntest, nx1, ny1 = x_test.shape
x_trainAdj = x_train.reshape((ntrain,nx*ny))
x_testAdj = x_test.reshape((ntest,nx1*ny1))
#fit model
np.random.seed(345)
svmodel=svm.SVC(kernel='poly',degree=2)
svmodel.fit(x_trainAdj,y_train)
#predict and calculate prediction
np.random.seed(345)
svmodel.predict([x_testAdj[0]])
y_pred = svmodel.predict(x_testAdj)
acc = metrics.accuracy_score(y_test, y_pred, normalize=True)
print('SVM accuracy is : ' + str(acc))
# ## Setting up the Convolutional Neural Network
#
# This is a 3 layer convolutional neural network. The benefit of running a CNN is it automatically detects important features with the passing of each layer and filter. Usually when adding more layers to a neural network, the model is able to fit more complex patterns, however this comes at the cost of increasing dimensionality. With this we can store more values in a weight which may cause our model to overfit.
# +
# Normalize pixel values to be between 0 and 1
x_train = x_train.reshape(x_train.shape[0],28,28,1)
x_test = x_test.reshape(x_test.shape[0],28,28,1)
trainX = x_train / 255.0
testX = x_test/ 255.0
#one hot encode y labels
trainY = to_categorical(y_train)
testY = to_categorical(y_test)
# +
#Built off the primary model in Exercise 3
np.random.seed(345)
# define the variable input_s here, which is the size of the images in CIFAR10.(width, height, channel)
input_s = (28,28,1)
#base model
model = keras.Sequential()
#first layer
model.add(Conv2D(32, kernel_size=(3, 3),activation='relu',kernel_initializer='he_normal',input_shape=input_s))
model.add(MaxPooling2D((2,2)))
#second layer
model.add(Conv2D(64, (3, 3), activation='relu'))
#third layer
model.add(Conv2D(128, (3, 3), activation='relu'))
#convert matrix to single array
model.add(Flatten())
#Transform vector
model.add(Dense(64, activation='relu', kernel_initializer='he_uniform'))
model.add(Dense(128, activation='relu', kernel_initializer='he_uniform'))
#add dropout to reduce overfitting
model.add(layers.Dropout(0.2))
#add softmax prediction layer
model.add(Dense(10, activation='softmax'))
#compile
model.compile(optimizer='Adam', loss='categorical_crossentropy', metrics=['accuracy'])
#set epochs
epo = 10
#fit the model
hist = model.fit(trainX, trainY, epochs=epo, batch_size=64, validation_data=(x_test, testY), verbose=1)
# +
acc1 = hist.history['accuracy']
acc_val = hist.history['val_accuracy']
epoch_range = range(len(acc))
plt.plot(epoch_range,acc,'g', label = "Training Accuracy")
plt.plot(epoch_range,acc_val,'b',label = "Validation Accuracy")
plt.title('Training and validation accuracy')
plt.legend()
plt.figure()
plt.show();
# -
np.random.seed(345)
score = model.evaluate(testX, testY, verbose=1)
print("CNN Accuracy: " + str(score[1]))
# ## Conclusion
#
# The difference in accuracies between my svm and my cnn is 0.049. In my tests, it takes about 30 minutes for the svm to run, and about half that time for the cnn to run. There is a definite edge for convolutional neural networks when using a large dataset. If we had less data and features we might see better performance and accuracy for an support vector model.
#
# There were a few problems I faced in coding both of the svm and cnn model. The biggest ones were dealing with the shape of my input data. For the svm I had to reshape the data to fit the svm model. I previously was trying to input 3 dimensional data, when the model required 2 dimensions. Likewise on the cnn I had to reshape the data to be a tensor.
#
# Prior to normalizing the data, I was getting very poor accuracies: 26% and 10%. I thought adding a dropout layer would help as that's a way to reduce overfitting. After I normalized my data, I saw much better prediction values: >89%. I kept in the dropout layer to play with what different values would do. I saw when increasing the dropout value to around 0.25 I saw a decrease in accuracy to about 2%, however when decreasing the dropout value my prediction increased to about 91%.
#
# Overall we see that using a CNN on a large image dataset is much better than using a support vector model. Moving forward I would like to test this model on other image classication datasets to see how well it performs. I would also like to increase the epochs it runs through to see how the validation validation accuracy changes.
| Fashion_MNIST_Project_Hinds.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# name: python2
# ---
# + [markdown] id="JndnmDMp66FL" colab_type="text"
# ##### Copyright 2018 Google LLC.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + id="hMqWDc_m6rUC" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}} cellView="both"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="pNqKk1MmrakH" colab_type="text"
# # Semantic Dictionaries -- Building Blocks of Interpretability
#
# This colab notebook is part of our **Building Blocks of Intepretability** series exploring how intepretability techniques combine together to explain neural networks. If you haven't already, make sure to look at the [**corresponding paper**](https://distill.pub/2018/building-blocks) as well!
#
# This notebook studies **semantic dictionaries**. The basic idea of semantic dictionaries is to marry neuron activations to visualizations of those neurons, transforming them from abstract vectors to something more meaningful to humans. Semantic dictionaries can also be applied to other bases, such as rotated versions of activations space that try to disentangle neurons.
#
# <br>
# <img src="https://storage.googleapis.com/lucid-static/building-blocks/notebook_heroes/semantic-dictionary.jpeg" width="648"></img>
# <br>
#
# This tutorial is based on [**Lucid**](https://github.com/tensorflow/lucid), a network for visualizing neural networks. Lucid is a kind of spiritual successor to DeepDream, but provides flexible abstractions so that it can be used for a wide range of interpretability research.
#
# **Note**: The easiest way to use this tutorial is [as a colab notebook](), which allows you to dive in with no setup. We recommend you enable a free GPU by going:
#
# > **Runtime** → **Change runtime type** → **Hardware Accelerator: GPU**
#
# Thanks for trying Lucid!
#
# + [markdown] id="hOBBuzMaxU37" colab_type="text"
# # Install / Import / Load
# + [markdown] id="UL1yOZtjqkcj" colab_type="text"
# This code depends on [Lucid](https://github.com/tensorflow/lucid) (our visualization library), and [svelte](https://svelte.technology/) (a web framework). The following cell will install both of them, and dependancies such as TensorFlow. And then import them as appropriate.
# + id="AA17rJBLuyYH" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 102} outputId="3acd867e-4fc2-4369-8684-cbdcd3f70c7d" executionInfo={"status": "ok", "timestamp": 1520312194763, "user_tz": 480, "elapsed": 15116, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# !pip install --quiet lucid==0.0.5
# !npm install -g svelte-cli@2.2.0
import numpy as np
import tensorflow as tf
import lucid.modelzoo.vision_models as models
import lucid.optvis.render as render
from lucid.misc.io import show, load
from lucid.misc.io.showing import _image_url
import lucid.scratch.web.svelte as lucid_svelte
# + [markdown] id="HfG1iu5KdmUV" colab_type="text"
# # Semantic Dictionary Code
# + [markdown] id="GABn20jcn4MC" colab_type="text"
# ## **Defining the interface**
#
# First, we define our "semantic dictionary" interface as a [svelte component](https://svelte.technology/). This makes it easy to manage state, like which position we're looking at.
# + id="ZhUtaAmcOIrw" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 102} outputId="20678755-ab0c-43fc-fa6c-24bdc361fc96" executionInfo={"status": "ok", "timestamp": 1520312330316, "user_tz": 480, "elapsed": 542, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
# %%html_define_svelte SemanticDict
<div class="figure">
<div class="input_image">
<div class="image" style="background-image: url({{image_url}}); z-index: -10;"></div>
<svg class="pointer_container" viewBox="0 0 {{N[0]}} {{N[1]}}">
{{#each xs as x}}
{{#each ys as y}}
<rect x={{x}} y={{y}} width=1 height=1
class={{(x == pos[0] && y == pos[1])? "selected" : "unselected"}}
on:mouseover="set({pos: [x,y]})"></rect>
{{/each}}
{{/each}}
</svg>
</div>
<div class="dict" >
{{#each present_acts as act, act_ind}}
<div class="entry">
<div class="sprite" style="background-image: url({{spritemap_url}}); width: {{sprite_size}}px; height: {{sprite_size}}px; background-position: -{{sprite_size*(act.n%sprite_n_wrap)}}px -{{sprite_size*Math.floor(act.n/sprite_n_wrap)}}px; --info: {{act.n}};"></div>
<div class="value" style="height: {{sprite_size*act.v/1000.0}}px;"></div>
</div>
{{/each}}
</div>
</div>
<style>
.figure {
padding: 10px;
width: 1024px;
}
.input_image {
display: inline-block;
width: 224px;
height: 224px;
}
.input_image .image, .input_image .pointer_constainer {
position: absolute;
width: 224px;
height: 224px;
border-radius: 8px;
}
.pointer_container rect {
opacity: 0;
}
.pointer_container .selected {
opacity: 1;
fill: none;
stroke: hsl(24, 100%, 50%);
stroke-width: 0.1px;
}
.dict {
height: 128px;
display: inline-block;
vertical-align: bottom;
padding-bottom: 64px;
margin-left: 64px;
}
.entry {
margin-top: 9px;
margin-right: 32px;
display: inline-block;
}
.value {
display: inline-block;
width: 32px;
border-radius: 8px;
background: #777;
}
.sprite {
display: inline-block;
border-radius: 8px;
}
.dict-text {
display: none;
font-size: 24px;
color: #AAA;
margin-bottom: 20px;
}
</style>
<script>
function range(n){
return Array(n).fill().map((_, i) => i);
}
export default {
data () {
return {
spritemap_url: "",
sprite_size: 64,
sprite_n_wrap: 1e8,
image_url: "",
activations: [[[{n: 0, v: 1}]]],
pos: [0,0]
};
},
computed: {
present_acts: (activations, pos) => activations[pos[1]][pos[0]],
N: activations => [activations.length, activations[0].length],
xs: (N) => range(N[0]),
ys: (N) => range(N[1])
},
helpers: {range}
};
</script>
# + [markdown] id="F4l9Ki-UoVko" colab_type="text"
# ## **Spritemaps**
#
# In order to use the semantic dictionaries, we need "spritemaps" of channel visualizations.
# These visualization spritemaps are large grids of images (such as [this one](https://storage.googleapis.com/lucid-static/building-blocks/sprite_mixed4d_channel.jpeg)) that visualize every channel in a layer.
# We provide spritemaps for GoogLeNet because making them takes a few hours of GPU time, but
# you can make your own channel spritemaps to explore other models. Check out other notebooks on how to
# make your own neuron visualizations.
#
# It's also worth noting that GoogLeNet has unusually semantically meaningful neurons. We don't know why this is -- although it's an active area of research for us. More sophisticated interfaces, such as neuron groups, may work better for networks where meaningful ideas are more entangled or less aligned with the neuron directions.
# + id="BpGLiyEEoPfB" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
layer_spritemap_sizes = {
'mixed3a' : 16,
'mixed3b' : 21,
'mixed4a' : 22,
'mixed4b' : 22,
'mixed4c' : 22,
'mixed4d' : 22,
'mixed4e' : 28,
'mixed5a' : 28,
}
def googlenet_spritemap(layer):
assert layer in layer_spritemap_sizes
size = layer_spritemap_sizes[layer]
url = "https://storage.googleapis.com/lucid-static/building-blocks/googlenet_spritemaps/sprite_%s_channel_alpha.jpeg" % layer
return size, url
# + [markdown] id="eP9jcxiLowkZ" colab_type="text"
# ## **User facing constructor**
#
# Now we'll create a convenient API for creating semantic dictionary visualizations. It will compute the network activations for an image, grab an appropriate spritemap, and render the interface.
# + id="9czK9sf1d1bU" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}}
googlenet = models.InceptionV1()
googlenet.load_graphdef()
def googlenet_semantic_dict(layer, img_url):
img = load(img_url)
# Compute the activations
with tf.Graph().as_default(), tf.Session():
t_input = tf.placeholder(tf.float32, [224, 224, 3])
T = render.import_model(googlenet, t_input, t_input)
acts = T(layer).eval({t_input: img})[0]
# Find the most interesting position for our initial view
max_mag = acts.max(-1)
max_x = np.argmax(max_mag.max(-1))
max_y = np.argmax(max_mag[max_x])
# Find appropriate spritemap
spritemap_n, spritemap_url = googlenet_spritemap(layer)
# Actually construct the semantic dictionary interface
# using our *custom component*
lucid_svelte.SemanticDict({
"spritemap_url": spritemap_url,
"sprite_size": 110,
"sprite_n_wrap": spritemap_n,
"image_url": _image_url(img),
"activations": [[[{"n": n, "v": float(act_vec[n])} for n in np.argsort(-act_vec)[:4]] for act_vec in act_slice] for act_slice in acts],
"pos" : [max_y, max_x]
})
# + [markdown] id="k0-gZUpApZyz" colab_type="text"
# # Now let's make some semantic dictionaries!
# + id="MEWC-UKdqRGC" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 265} outputId="096bc577-d5d9-4c48-a914-411b8e77b03a" executionInfo={"status": "ok", "timestamp": 1520312336706, "user_tz": 480, "elapsed": 2455, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
googlenet_semantic_dict("mixed4d", "https://storage.googleapis.com/lucid-static/building-blocks/examples/dog_cat.png")
# + id="Izf_YqCRe6E7" colab_type="code" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{}], "base_uri": "https://localhost:8080/", "height": 265} outputId="ed2b0826-52b4-4714-e019-773162c2b170" executionInfo={"status": "ok", "timestamp": 1520312338047, "user_tz": 480, "elapsed": 1259, "user": {"displayName": "", "photoUrl": "", "userId": ""}}
googlenet_semantic_dict("mixed4d", "https://storage.googleapis.com/lucid-static/building-blocks/examples/flowers.png")
| notebooks/building-blocks/SemanticDictionary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="pUVMRN-3IJU7"
import pandas as pd
import matplotlib.pyplot as plt
# # !pip install yfinance
import yfinance as yf
# + [markdown] colab_type="text" id="YbpgXGAJIJVE"
# ### 1. Load Financial Data of Company
# + colab={} colab_type="code" id="7uaPKUZHIJVH"
# Set file path for specific company data
firmData = pd.read_csv('./morningstarData/sabr.csv', skiprows=2, index_col='Unnamed: 0')
# + colab={"base_uri": "https://localhost:8080/", "height": 360} colab_type="code" id="V3GgVINrIJVM" outputId="84b8da44-ead2-492a-9e1e-54bce1a42fd5"
firmData.head()
# + [markdown] colab_type="text" id="AYdrkRSRIJVT"
# ### 2. Read Metrics From Financial Data
#
# ### Functions for Metrics
# +
# Return a specific stock metric of at a specific date
# Remove commas if any
def search_value(index_name, date):
# return float(firmData.loc[index_name, date].replace(",", ""))
return float(firmData.loc[index_name, date])
# Return all historical data for a specific stock metric
# Remove commas if any
def historical_value(index_name):
# return firmData.loc[index_name, ].str.replace(",","").astype(float)
return firmData.loc[index_name, ].astype(float)
# TODO: Create function that acts like search_value, however dynamically uses most recent YE
# -
# ##### (a) Shares Outstanding
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="j_vFXw--IJVU" outputId="fc6f0543-f630-4026-d01e-aa40cdb0487b"
# Read 2019 shares outstanding
# Note: This line might not work if the company doesn't have a Sept fiscal year end
# Note: Or, if the company does have Sept fiscal year end, might not have gone public in 2019
print(search_value('Shares Mil', '2019-12'))
# Read all historical data of shares outstanding
print(historical_value('Shares Mil'))
# + [markdown] colab_type="text" id="UqttGWPBIJVv"
# ##### (b) EPS
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="sN3kc5ERIJVw" outputId="aab3de88-f291-41b3-a5d9-e701db0b3a65"
# Read 2019 EPS
print(search_value('Earnings Per Share USD', '2019-12'))
# Read all historical data of shares outstanding
print(historical_value('Earnings Per Share USD'))
# + [markdown] colab_type="text" id="OdZJbvJgIJV3"
# ##### (c) Dividend Rate
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="YGck0iJLIJV_" outputId="07980dfc-a1e0-464f-b44a-aac71e719f6c"
# Read 2019 dividend rate
print(search_value('Dividends USD', '2019-12'))
# Read historical dividend rate
# Note: NaN manipulation will be introduced later
print(historical_value('Dividends USD'))
# + [markdown] colab_type="text" id="6IrS3Vk1IJWG"
# ##### (d) Dividend Yield
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="rBBJbfElIJWI" outputId="73402a6a-589a-4dca-92c8-2fafdb1900ec"
# Read 2019 dividend yield
print(search_value('Payout Ratio % *', '2019-12'))
# Read historical dividend yield
# Note: TTM = Trailing Twelve Months
print(historical_value('Payout Ratio % *'))
# + [markdown] colab_type="text" id="t5MmSkvwIJWO"
# ##### (e) Debt/Equity Ratio
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="GsK8JpzXIJWO" outputId="d6effc84-92da-4a68-a0b2-6b0e459bde39"
# Read 2019 D/E Ratio
print(search_value('Debt/Equity', '2019-12'))
# Read historical D/E ratio
print(historical_value('Debt/Equity'))
# + [markdown] colab_type="text" id="7KazUwjrIJWT"
# ##### (f) Book Value Per Share
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="h6RqEP8WIJWU" outputId="922c45d2-db1d-41a5-e603-c4c50f4c5162"
# Read 2019 book value per share
print(search_value('Book Value Per Share * USD', '2019-12'))
# Read historical book value per share
print(historical_value('Book Value Per Share * USD'))
# + [markdown] colab_type="text" id="JGDD-821IJWZ"
# ##### (g) ROE (Return on Equity)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="a_5z0KgZIJWZ" outputId="db1ad314-2bad-479b-e5fa-82a71651c013"
# Read 2019 ROE
print(search_value('Return on Equity %', '2019-12'))
# Read historical ROE
print(historical_value('Return on Equity %'))
# + [markdown] colab_type="text" id="okzk5BQrIJWd"
# ##### (h) Current Ratio
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="ynvMbpk0IJWd" outputId="6e6c5ef3-8105-4d5d-b25a-57d87d6e6c6a"
# Read 2019 current ratio
print(search_value('Current Ratio', '2019-12'))
# Read historical current ratio
print(historical_value('Current Ratio'))
# + [markdown] colab_type="text" id="miWi8QhJIJWh"
# ### 3. Get Complete Stock Price Dataframe
# + colab={} colab_type="code" id="ohiyR2huIJWh"
# Set ticker symbol
ticker = "SABR"
stock = yf.Ticker(ticker)
# Get complete history of stock prices
stock_price = stock.history(period='max')
stock_price.head()
# -
# Line graph of stock price over time
stock_price['Close'].plot()
plt.xlabel("Date", labelpad=15)
plt.ylabel("Stock Price", labelpad=15)
plt.title(f"{ticker} Stock Price Over Time", y=1.02, fontsize=22);
# + [markdown] colab_type="text" id="7P0lQIxeIJWj"
# ### 4. Calculate Price/Earnings (PE) & Price/Book Value (P/BV)
#
# ##### (a) Latest P/E
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="NO-ekSyoIJWk" outputId="c33fac86-41ba-4b20-fbf8-9038f6fdbdd8"
# Obtain the most recent closing price of the current stock
# TODO: Change based on current date, should just be close price of yesterday
latest_price = stock_price.loc['2020-09-11','Close']
# Obtain the most recent fiscal year end EPS
latest_eps = search_value('Earnings Per Share USD', '2019-12')
# Calculate the P/E ratio
latest_PtoE = latest_price/latest_eps
print(latest_PtoE)
# + [markdown] colab_type="text" id="3sYRzw9OIJWl"
# ##### (b) Latest P/BV
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="UlF_ATanIJWl" outputId="fa22cd43-dcee-47d8-ddf5-49c190d620c3"
# Obtain the most recent closing price of the current stock
# TODO: Change based on current date, should just be close price of yesterday
latest_price = stock_price.loc['2020-09-11','Close']
# Obtain the most recent fiscal year end book value per share
latest_eps = search_value('Book Value Per Share * USD', '2019-12')
# Calculate the P/BV ratio
latest_PtoBV = latest_price/latest_eps
print(latest_PtoBV)
# + [markdown] colab_type="text" id="cf9H7tWeIJWn"
# ### 5. Formatting to One Concise Dataframe
#
# ##### (a) Transpose Company Dataframe
# For the convenience of calculation, we want to transpose the dataframe. Such that the time would be rows and metrics would be columns.
# + colab={} colab_type="code" id="kRxRWL1ZIJWn"
# Transpose dataframe
firmData = firmData.T
firmData.head()
# + [markdown] colab_type="text" id="Vkp6AyXAIJWr"
# ##### (b) Extract metrics we need
# The financial data contains a lot of metrics, we are not going to use most of them. So, let's create a smaller dataframe with whatever metrics we will use.
# + colab={"base_uri": "https://localhost:8080/", "height": 357} colab_type="code" id="l9qYfm3KIJWr" outputId="d61526f0-f7d3-4615-e5af-7e2ee9d2f8be"
col_names = ['Shares Mil','Earnings Per Share USD','Dividends USD',
'Payout Ratio % *', 'Debt/Equity', 'Book Value Per Share * USD',
'Return on Equity %', 'Current Ratio']
firm_df = firmData[col_names]
firm_df.head()
# + [markdown] colab_type="text" id="Y5Tc4CImIJWt"
# ##### (c) Rename Columns
#
# Now we have a concise dataframe `firm_df`, which is easier to use. Now let's get rid of the column special symbols and unify the column name style by renaming them.
# + colab={"base_uri": "https://localhost:8080/", "height": 309} colab_type="code" id="vOY7_ZngIJWt" outputId="067904c5-2cd1-4ee9-b1af-c3759937e5b1"
new_col = ['shares_outstanding', 'earning_per_share', 'dividend_rate',
'dividend_yield', 'debt_to_equity', 'book_value_per_share',
'return_on_equity', 'current_ratio']
firm_df.columns = new_col
firm_df.head()
# + [markdown] colab_type="text" id="XsMSAdIdIJWv"
# Now the dataframe looks great! Attention, do not get the order of new column (`new_col`) names wrong. It should be in exactly the same order as current column.
| stockAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from bertviz.attention_gpt2 import AttentionGPT2Data, show
from bertviz.pytorch_pretrained_bert import GPT2Model, GPT2Tokenizer
# + language="javascript"
# require.config({
# paths: {
# d3: '//cdnjs.cloudflare.com/ajax/libs/d3/3.4.8/d3.min'
# }
# });
# + language="javascript"
# IPython.OutputArea.auto_scroll_threshold = 9999;
# -
def call_html():
import IPython
display(IPython.core.display.HTML('''
<script src="/static/components/requirejs/require.js"></script>
<script>
requirejs.config({
paths: {
base: '/static/base',
"d3": "https://cdnjs.cloudflare.com/ajax/libs/d3/3.5.8/d3.min",
jquery: '//ajax.googleapis.com/ajax/libs/jquery/2.0.0/jquery.min',
},
});
</script>
'''))
model = GPT2Model.from_pretrained('gpt2')
tokenizer = GPT2Tokenizer.from_pretrained('gpt2')
text = "The dog on the ship ran off, and the dog was found by the crew."
map_data = AttentionGPT2Data(model, tokenizer)
tokens, atts = map_data.get_data(text)
call_html()
show(tokens, atts)
| bertviz/bertviz_summary_gpt2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# #!/usr/bin/env python
# coding: utf-8
from utils import *
"""
######################################################################
1. add metric "siameseEucDist" - Done
2. Do data processing: siamese and classification data togther for multitasking - under procsess
3. Modify the trainer function
######################################################################
"""
# import torch
# from torchvision.datasets import FashionMNIST
# from torchvision import transforms
# mean, std = 0.28604059698879553, 0.35302424451492237
# batch_size = 256
# train_dataset = FashionMNIST('../data/FashionMNIST', train=True, download=True,
# transform=transforms.Compose([
# transforms.ToTensor(),
# transforms.Normalize((mean,), (std,))
# ]))
# test_dataset = FashionMNIST('../data/FashionMNIST', train=False, download=True,
# transform=transforms.Compose([
# transforms.ToTensor(),
# transforms.Normalize((mean,), (std,))
# ]))
# cuda = torch.cuda.is_available()
# kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
# train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size, shuffle=True, **kwargs)
# test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
# n_classes = 10
import torch
from torchvision.datasets import MNIST
from torchvision import transforms
mean, std = 0.1307, 0.3081
train_dataset = MNIST('../data/MNIST', train=True, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((mean,), (std,))
]))
test_dataset = MNIST('../data/MNIST', train=False, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((mean,), (std,))
]))
n_classes = 10
# ## Common setup
# In[3]:
import torch
from torch.optim import lr_scheduler
import torch.optim as optim
from torch.autograd import Variable
from metrics import *
from trainer import fit, fit_siam
import numpy as np
cuda = torch.cuda.is_available()
# get_ipython().run_line_magic('matplotlib', 'inline')
import matplotlib
import matplotlib.pyplot as plt
import os
from utils import *
import utils
import sys
# gpu, seed_offset, MVLW = int(sys.argv[1]), int(sys.argv[2]), int(sys.argv[3])
gpu, seed_offset, MVLW = 3, 50, 0
os.environ["CUDA_VISIBLE_DEVICES"] = "1,2,3,4"
mnist_classes = ['0', '1', '2', '3', '4', '5', '6', '7', '8', '9']
colors = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728',
'#9467bd', '#8c564b', '#e377c2', '#7f7f7f',
'#bcbd22', '#17becf']
def plot_embeddings(embeddings, targets, xlim=None, ylim=None):
plt.figure(figsize=(10,10))
for i in range(10):
inds = np.where(targets==i)[0]
plt.scatter(embeddings[inds,0], embeddings[inds,1], alpha=0.5, color=colors[i])
if xlim:
plt.xlim(xlim[0], xlim[1])
if ylim:
plt.ylim(ylim[0], ylim[1])
plt.legend(mnist_classes)
def extract_embeddings(dataloader, model):
with torch.no_grad():
model.eval()
embeddings = np.zeros((len(dataloader.dataset), 2))
labels = np.zeros(len(dataloader.dataset))
k = 0
for images, target in dataloader:
if cuda:
images = images.cuda(gpu)
embeddings[k:k+len(images)] = model.get_embedding(images).data.cpu().numpy()
labels[k:k+len(images)] = target.numpy()
k += len(images)
return embeddings, labels
# Set up data loaders
from datasets import SiameseMNIST, SiameseMNIST_MT
## Step 1
# siamese_train_dataset = SiameseMNIST(train_dataset) # Returns pairs of images and target same/different
# siamese_test_dataset = SiameseMNIST(test_dataset)
# batch_size = 128
# kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
# siamese_train_loader = torch.utils.data.DataLoader(siamese_train_dataset, batch_size=batch_size, shuffle=True, **kwargs)
# siamese_test_loader = torch.utils.data.DataLoader(siamese_test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
# Step 1 - Multi-task data
# Set up the network and training parameters
from networks import EmbeddingNet, SiameseNet, SiameseNet_ClassNet
from losses import ContrastiveLoss
from losses import ContrastiveLoss_mod, CrossEntropy
# torch.manual_seed(0)
# np.random.seed(0)
log_interval = 5
siamese_MT_train_dataset = SiameseMNIST_MT(train_dataset, seed=0) # Returns pairs of images and target same/different
siamese_MT_test_dataset = SiameseMNIST_MT(test_dataset, seed=0)
interval = 0.025
write_list = []
mw_list = []
# For reproducibility
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
metric_classes=[SimpleSiamDistAcc, AccumulatedAccuracyMetric_mod]
margin = 1.0
if MVLW:
init_mix_weight = 0.5
n_epochs = 20
start, interval, end = 0, 1, 1
seedmax = 1
log_tag = "auto"
else:
# start, interval, end = 0, interval, 1.0 + interval
MW = 0.85
start, interval, end = MW, 0.25, MW + interval
seedmax = 1
n_epochs = 20
log_tag = "gridsearch"
# -
for k in range(0, seedmax):
for mwk, mix_weight in enumerate(np.arange(start, end, interval)):
if MVLW:
mix_weight = init_mix_weight
print("Mix weight count:", mwk, mix_weight)
mix_weight = round(mix_weight, 3)
# for _, _ in enumerate([1]):
# for _, mix_weight in enumerate(np.arange(0, 0.2, interval)):
seed = k + seed_offset
torch.manual_seed(seed)
np.random.seed(seed)
# mix_weight = 0.5 ### initial weight
exp_tag = "margin{}_seedoffset_{}_epoch_{}_intvl{}".format(margin, seed_offset, n_epochs, interval)
batch_size = 2**11
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
# siamese_MT_train_loader = torch.utils.data.DataLoader(siamese_MT_train_dataset, batch_size=batch_size, shuffle=True, **kwargs)
# siamese_MT_test_loader = torch.utils.data.DataLoader(siamese_MT_test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
siamese_train_loader = torch.utils.data.DataLoader(siamese_MT_train_dataset, batch_size=batch_size, shuffle=False, **kwargs)
siamese_test_loader = torch.utils.data.DataLoader(siamese_MT_test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
# Step 2
embedding_net = EmbeddingNet()
# Step 3
model = SiameseNet(embedding_net)
model_mt = SiameseNet_ClassNet(embedding_net, n_classes=n_classes)
if cuda:
model.cuda(gpu)
model_mt.cuda(gpu)
# Step 4
# loss_fn = ContrastiveLoss(margin)
loss_fn = ContrastiveLoss_mod(margin)
loss_fn_ce = CrossEntropy()
lr = 1e-3
optimizer = optim.Adam(model.parameters(), lr=lr)
scheduler = lr_scheduler.StepLR(optimizer, 8, gamma=0.1, last_epoch=-1)
# class AccumulatedAccuracyMetric_mod(Metric):
loss_fn_tup = (loss_fn, loss_fn_ce)
# for Siamese + Classifier net
model_mt = SiameseNet_ClassNet(embedding_net, n_classes=n_classes)
if cuda:
model.cuda(gpu)
model_mt.cuda(gpu)
mix_weight = torch.tensor(mix_weight).cuda(gpu)
write_var, mix_weight, mix_weight_list = fit_siam(gpu, siamese_train_loader, siamese_test_loader, model_mt, loss_fn_tup, optimizer, scheduler, n_epochs, cuda, log_interval, mix_weight, MVLW, metric_classes=metric_classes, seed=seed)
write_list.append(write_var)
mw_list.append(', '.join(mix_weight_list))
write_txt("log_{}/{}.txt".format(log_tag, exp_tag), write_list)
# write_txt("mw_est/{}2.txt".format(exp_tag), mw_list)
print("write var:", write_var)
# +
# train_dataset=siamese_MT_train_dataset
# test_dataset=siamese_MT_test_dataset
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size, shuffle=False, **kwargs)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
train_embeddings_cl, train_labels_cl = extract_embeddings(train_loader, model_mt)
plot_embeddings(train_embeddings_cl, train_labels_cl)
val_embeddings_cl, val_labels_cl = extract_embeddings(test_loader, model_mt)
plot_embeddings(val_embeddings_cl, val_labels_cl)
# -
| siamese_MNIST_multiTask.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Ruby 2.4.1
# language: ruby
# name: ruby
# ---
# +
require 'csv'
require 'redis'
require 'json'
require 'date'
#1 year version (2017)
INPUTDATA = 'input_data_1y.txt'
#10 years version (2008-2017)
#INPUTDATA = 'input_data_10y.txt'
redis = Redis.new(host: "redis", port: 6379, db: 0)
f = File.open(INPUTDATA, 'r')
f.readline
redis.flushdb
key = 0
begin
loop do
if key % 10000 == 0 then puts "#{Time.now} insert-data: #{key} processed" end
redis.set(key,f.readline.strip.split("\t").map{|i| i.strip}.to_s)
key = key + 1
end
rescue StandardError => e
puts e
end
size = key
(0..size - 1).each do |key|
if key % 10000 == 0 then puts "#{Time.now} prr-params: #{key} processed" end
t = redis.get(key).nil? ? nil : eval(redis.get(key))
if t
year, month = Date.parse(t[0]).year, Date.parse(t[0]).month
drug, soc = t[1], t[2]
redis.incr({incremental: false, year: year, month: month, drug: drug, soc: soc})
redis.incr({incremental: false, year: year, month: month, drug: drug})
redis.incr({incremental: false, year: year, month: month, soc: soc})
redis.incr({incremental: false, year: year, month: month})
(year..2017).each do |y|
start_month = year == y ? month : 1
(start_month..12).each do |m|
redis.incr({incremental: true, year: y, month: m, drug: drug, soc: soc})
redis.incr({incremental: true, year: y, month: m, drug: drug})
redis.incr({incremental: true, year: y, month: m, soc: soc})
redis.incr({incremental: true, year: y, month: m})
end
end
end
end
| DockerContainer/TEDAR/sciruby/Init.ipynb |
# ---
# jupyter:
# jupytext:
# formats: ipynb,md:myst
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (XPython)
# language: python
# name: xpython
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Information Theory
# + [markdown] slideshow={"slide_type": "-"} tags=["remove-cell"]
# **CS1302 Introduction to Computer Programming**
# ___
# + [markdown] slideshow={"slide_type": "subslide"}
# As mentioned in previous lectures, the following two lists `coin_flips` and `dice_rolls` simulate the random coin flips and rollings of a dice:
# + nbgrader={"grade": false, "grade_id": "random", "locked": true, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# Do NOT modify any variables defined here because some tests rely on them
import random
random.seed(0) # for reproducible results.
num_trials = 200
coin_flips = ['H' if random.random() <= 1/2 else 'T' for i in range(num_trials)]
dice_rolls = [random.randint(1,6) for i in range(num_trials)]
print('coin flips: ',*coin_flips)
print('dice rolls: ',*dice_rolls)
# + [markdown] slideshow={"slide_type": "subslide"}
# **What is the difference of the two random processes?
# Can we say one process has more information content than the other?**
# + [markdown] slideshow={"slide_type": "-"}
# In this Lab, we will use dictionaries to store their distributions and then compute their information content using information theory, which was introduced by *<NAME>*. It has [numerous applications](https://www.khanacademy.org/computing/computer-science/informationtheory):
# - *compression* (to keep files small)
# - *communications* (to send data mobile phones), and
# - *machine learning* (to identify relevant features to learn from).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Entropy
# + [markdown] slideshow={"slide_type": "fragment"}
# Mathematically, we denote a distribution as $\mathbf{p}=[p_i]_{i\in \mathcal{S}}$, where
# - $\mathcal{S}$ is the set of distinct outcomes, and
# - $p_i$ denotes the probability (chance) of seeing outcome $i$.
# + [markdown] slideshow={"slide_type": "fragment"}
# The following code shown in the lecture uses a dictionary to store the distribution for a sequence efficiently without storing outcomes with zero counts:
# + nbgrader={"grade": false, "grade_id": "dist", "locked": true, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# Do NOT modify any variables defined here because some tests rely on them
def distribute(seq):
'''Returns a dictionary where each value in a key-value pair is
the probability of the associated key occuring in the sequence.
'''
p = {}
for i in seq:
p[i] = p.get(i,0) + 1/len(seq)
return p
# tests
coin_flips_dist = distribute(coin_flips)
dice_rolls_dist = distribute(dice_rolls)
print('Distribution of coin flips:', coin_flips_dist)
print('Distribution of dice rolls:', dice_rolls_dist)
# + [markdown] slideshow={"slide_type": "fragment"}
# For $\mathbf{p}$ to be a valid distribution, the probabilities $p_i$'s have to sum to $1$, i.e.,
#
# $$\sum_{i\in \mathcal{S}} p_i = 1, $$
# which can be verified as follows:
# + slideshow={"slide_type": "-"}
import math
assert math.isclose(sum(coin_flips_dist.values()),1) and math.isclose(sum(dice_rolls_dist.values()),1)
# + [markdown] slideshow={"slide_type": "subslide"}
# **How to measure the information content?**
# + slideshow={"slide_type": "-"} language="html"
# <iframe width="912" height="513" src="https://www.youtube.com/embed/2s3aJfRr9gE" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
# + [markdown] slideshow={"slide_type": "subslide"}
# In information theory, the information content of a distribution is measured by its [*entropy*](https://en.wikipedia.org/wiki/Entropy_(information_theory)) defined as:
#
# $$ \begin{aligned} H(\mathbf{p}) &:= \sum_{i\in \mathcal{S}} p_i \overbrace{\log_2 \tfrac{1}{p_i}}^{\text{called surprise} } \\ &= - \sum_{i\in \mathcal{S}} p_i \log_2 p_i \kern1em \text{(bits)} \end{aligned} $$
# with $p_i \log_2 \frac{1}{p_i} = 0$ if $p_i = 0$ because $\lim_{x\downarrow 0} x \log_2 \frac1x = 0$.
# + [markdown] slideshow={"slide_type": "fragment"}
# For instance, if $\mathbf{p}=(p_{H},p_{T})=(0.5,0.5)$, then
#
# $$\begin{aligned} H(\mathbf{p}) &= 0.5 \log_2 \frac{1}{0.5} + 0.5 \log_2 \frac{1}{0.5} \\ &= 0.5 + 0.5 = 1 \text{ bit,}\end{aligned} $$
# i.e., an outcome of a fair coin flip has one bit of information content, as expected.
# + [markdown] slideshow={"slide_type": "fragment"}
# On the other hand, if $\mathbf{p}=(p_{H},p_{T})=(1,0)$, then
# $$\begin{aligned} H(\mathbf{p}) &= 1 \log_2 \frac{1}{1} + 0 \log_2 \frac{1}{0} \\ &= 0 + 0 = 0 \text{ bits,}\end{aligned} $$
# i.e., an outcome of a biased coin flip that always comes up head has no information content, again as expected.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Define a function `entropy` that
# - takes a distribution $\mathbf{p}$ as its argument, and
# - returns the entropy $H(\mathbf{p})$.
#
# Handle the case when $p_i=0$, e.g., using the short-circuit evaluation of logical `and`.
# + nbgrader={"grade": false, "grade_id": "entropy", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def entropy(dist):
### BEGIN SOLUTION
return sum([p and -p * math.log2(p) for p in dist.values()])
### END SOLUTION
# + nbgrader={"grade": true, "grade_id": "test-entropy", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# tests
assert math.isclose(entropy({'H': 0.5, 'T': 0.5}), 1)
assert math.isclose(entropy({'H': 1, 'T': 0}), 0)
assert math.isclose(entropy(dict.fromkeys(range(1, 7), 1 / 6)), math.log2(6))
### BEGIN HIDDEN TESTS
assert entropy(dict(zip(range(3), [0.5, 0.25, 0.25]))) == 1.5
### END HIDDEN TESTS
# + [markdown] slideshow={"slide_type": "slide"}
# ## Uniform distribution maximizes entropy
# + [markdown] slideshow={"slide_type": "fragment"}
# Intuitively,
# - for large enough numbers of fair coin flips, we should have $\mathcal{S}=\{H,T\}$ and $p_H=p_T=0.5$, i.e., equal chance for head and tail.
# - for large enough numbers of fair dice rolls, we should have $p_i=\frac16$ for all $i\in \mathcal{S}=\{1,2,3,4,5,6\}$.
# + slideshow={"slide_type": "-"}
import matplotlib.pyplot as plt
def plot_distribution(seq):
dist = distribute(seq)
plt.stem(dist.keys(), # set-like view of the keys
dist.values(), # view of the values
use_line_collection=True)
plt.xlabel('Outcomes')
plt.title('Distribution')
plt.ylim(0, 1)
import ipywidgets as widgets
n_widget = widgets.IntSlider(
value=1,
min=1,
max=num_trials,
step=1,
description='n:',
continuous_update=False,
)
widgets.interactive(lambda n: plot_distribution(coin_flips[:n]),n=n_widget)
# + slideshow={"slide_type": "-"}
widgets.interactive(lambda n: plot_distribution(dice_rolls[:n]),n=n_widget)
# + [markdown] slideshow={"slide_type": "fragment"}
# A distribution is called a *uniform distribution* if all its distinct outcomes have the same probability of occuring, i.e.,
#
# $$ p_i = \frac{1}{|\mathcal{S}|}\kern1em \text{for all }i\in \mathcal{S}, $$
# where $|\mathcal{S}|$ is the mathematical notation to denote the size of the set $\mathcal{S}$.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Define a function `uniform` that
# - takes a sequence of possibly duplicate outcomes, and
# - returns a uniform distribution on the distinct outcomes.
# + nbgrader={"grade": false, "grade_id": "uniform", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def uniform(outcomes):
'''Returns the uniform distribution (dict) over distinct items in outcomes.'''
### BEGIN SOLUTION
distinct_outcomes = set(outcomes)
return dict.fromkeys(distinct_outcomes, 1 / len(distinct_outcomes))
### END SOLUTION
# + nbgrader={"grade": true, "grade_id": "test-uniform", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# tests
assert uniform('HT') == {'H': 0.5, 'T': 0.5}
assert uniform('HTH') == {'H': 0.5, 'T': 0.5}
fair_dice_dist = uniform(range(1, 7))
assert all(
math.isclose(fair_dice_dist[k], v) for k, v in {
1: 0.16666666666666666,
2: 0.16666666666666666,
3: 0.16666666666666666,
4: 0.16666666666666666,
5: 0.16666666666666666,
6: 0.16666666666666666
}.items())
### BEGIN HIDDEN TESTS
assert uniform(range(10)) == dict.fromkeys(range(10),1/10)
### END HIDDEN TESTS
# + [markdown] slideshow={"slide_type": "subslide"}
# **What is the entropy for uniform distributions?**
# + [markdown] slideshow={"slide_type": "fragment"}
# By definition,
#
# $$ \begin{aligned} H(\mathbf{p}) &:= \sum_{i\in \mathcal{S}} p_i \log_2 \tfrac{1}{p_i} \\ &= \sum_{i\in \mathcal{S}} \frac{1}{|\mathcal{S}|} \log_2 |\mathcal{S}| = \log_2 |\mathcal{S}| \kern1em \text{(bits)} \end{aligned} $$
#
# This reduces to the formula you learned in Lecture 1 and Lab 1 regarding the number of bits required to represent a set. This is the maximum possible entropy for a given finite set of possible outcomes.
# + [markdown] slideshow={"slide_type": "fragment"}
# You can use this result to test whether you have implemented both `entropy` and `uniform` correctly:
# + slideshow={"slide_type": "-"}
assert all(math.isclose(entropy(uniform(range(n))), math.log2(n)) for n in range(1,100))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Joint distribution and its entropy
# + [markdown] slideshow={"slide_type": "fragment"}
# If we duplicate a sequence of outcomes multiple times, the total information content should remain unchanged, NOT doubled, because the duplicate contain the same information as the original. We will verify this fact by creating a [joint distribution](https://en.wikipedia.org/wiki/Joint_probability_distribution)
#
# $$\mathbf{p}=[p_{ij}]_{i\in \mathcal{S},j\in \mathcal{T}}$$
# - where $\mathcal{S}$ and $\mathcal{T}$ are sets of outcomes; and
# - $p_{ij}$ is the chance we see outcomes $i$ and $j$ simultaneously.
#
# The subscript $ij$ in $p_{ij}$ denotes a tuple $(i,j)$, NOT the multiplication $i\times j$. We also have
#
# $$\sum_{i\in \mathcal{S}} \sum_{j\in \mathcal{T}} p_{ij} = 1.$$
#
#
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Define a function `jointly_distribute` that
# - takes two sequences `seq1` and `seq2` of outcomes with the same length, and
# - returns the joint distribution represented as a dictionary where each key-value pair has the key being a tuple `(i,j)` associated with the probability $p_{ij}$ of seeing `(i,j)` in `zip(seq1,seq2)`.
# + nbgrader={"grade": false, "grade_id": "jointly_distribute", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def jointly_distribute(seq1, seq2):
'''Returns the joint distribution of the tuple (i,j) of outcomes from zip(seq1,seq2).'''
### BEGIN SOLUTION
p, n = {}, min(len(seq1),len(seq2))
for ij in zip(seq1, seq2):
p[ij] = p.get(ij, 0) + 1 / n
return p
### END SOLUTION
# + code_folding=[] nbgrader={"grade": true, "grade_id": "test-jointly_distribute", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# tests
assert jointly_distribute('HT', 'HT') == {('H', 'H'): 0.5, ('T', 'T'): 0.5}
assert jointly_distribute('HHTT', 'HTHT') == {
('H', 'H'): 0.25,
('H', 'T'): 0.25,
('T', 'H'): 0.25,
('T', 'T'): 0.25
}
coin_flips_duplicate_dist = {
('T', 'T'): 0.5350000000000004,
('H', 'H'): 0.4650000000000003
}
coin_flips_duplicate_ans = jointly_distribute(coin_flips, coin_flips)
assert all(
math.isclose(coin_flips_duplicate_ans[i], pi)
for i, pi in coin_flips_duplicate_dist.items())
### BEGIN HIDDEN TESTS
assert jointly_distribute('HT', 'TH') == {('H', 'T'): 0.5, ('T', 'H'): 0.5}
assert jointly_distribute('HTTH', 'HHTT') == {
('H', 'H'): 0.25,
('H', 'T'): 0.25,
('T', 'H'): 0.25,
('T', 'T'): 0.25
}
coin_flips_dice_rolls_dist = {
('T', 4): 0.09500000000000001,
('T', 1): 0.10000000000000002,
('H', 3): 0.09500000000000001,
('H', 6): 0.09000000000000001,
('T', 2): 0.11000000000000003,
('H', 4): 0.075,
('T', 5): 0.06499999999999999,
('H', 5): 0.05499999999999999,
('T', 3): 0.09000000000000001,
('T', 6): 0.075,
('H', 1): 0.11500000000000003,
('H', 2): 0.035
}
coin_flips_dice_rolls_ans = jointly_distribute(coin_flips, dice_rolls)
assert all(
math.isclose(coin_flips_dice_rolls_ans[i], pi)
for i, pi in coin_flips_dice_rolls_dist.items())
### END HIDDEN TESTS
# + [markdown] slideshow={"slide_type": "fragment"}
# If you have implemented `entropy` and `jointly_distribute` correctly, you can verify that duplicating a sequence will give the same entropy.
# + slideshow={"slide_type": "-"}
assert math.isclose(entropy(jointly_distribute(coin_flips,coin_flips)), entropy(distribute(coin_flips)))
assert math.isclose(entropy(jointly_distribute(dice_rolls,dice_rolls)), entropy(distribute(dice_rolls)))
# + [markdown] slideshow={"slide_type": "fragment"}
# However, for two sequences generated independently, the joint entropy is roughly the sum of the individual entropies.
# + slideshow={"slide_type": "-"}
coin_flips_entropy = entropy(distribute(coin_flips))
dice_rolls_entropy = entropy(distribute(dice_rolls))
cf_dr_entropy = entropy(jointly_distribute(coin_flips, dice_rolls))
print(f'''Entropy of coin flip: {coin_flips_entropy}
Entropy of dice roll: {dice_rolls_entropy}
Sum of the above entropies: {coin_flips_entropy + dice_rolls_entropy}
Joint entropy: {cf_dr_entropy}''')
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Conditional distribution and entropy
# + [markdown] slideshow={"slide_type": "fragment"}
# Mathematically, we denote a [conditional distribution](https://en.wikipedia.org/wiki/Conditional_probability_distribution) as $\mathbf{q}:=[q_{j|i}]_{i\in \mathcal{S}, j\in \mathcal{T}}$, where
# - $\mathcal{S}$ and $\mathcal{T}$ are two sets of distinct outcomes, and
# - $q_{j|i}$ denotes the probability (chance) of seeing outcome $j$ given the condition that outcome $i$ is observed.
#
# For $\mathbf{q}$ to be a valid distribution, the probabilities $q_{j|i}$'s have to sum to $1$ for every $i$, i.e.,
#
# $$\sum_{j\in \mathcal{T}} q_{j|i} = 1 \kern1em \text{for all }i\in \mathcal{S} $$
# + [markdown] slideshow={"slide_type": "fragment"}
# For example, suppose we want to compute the distribution of coin flips given dice rolls, then the following assign `q_H_1` and `q_T_1` to the values $q_{H|1}$ and $q_{T|1}$ respectively:
# + slideshow={"slide_type": "-"}
coin_flips_1 = [j for i, j in zip(dice_rolls, coin_flips) if i == 1]
q_H_1 = coin_flips_1.count('H') / len(coin_flips_1)
q_T_1 = coin_flips_1.count('T') / len(coin_flips_1)
print('Coin flips given dice roll is 1:', coin_flips_1)
print('Distribution of coin flip given dice roll is 1: {{ "H": {}, "T": {}}}'.format(q_H_1, q_T_1))
assert math.isclose(q_H_1 + q_T_1, 1)
# + [markdown] slideshow={"slide_type": "fragment"}
# Note that `q_H_1 + q_T_1` is 1 as expected. Similarly, we can assign `q_H_2` and `q_T_2` to the values $q_{H|2}$ and $q_{T|2}$ respectively.
# + slideshow={"slide_type": "-"}
coin_flips_2 = [j for i, j in zip(dice_rolls, coin_flips) if i == 2]
q_H_2 = coin_flips_2.count('H') / len(coin_flips_2)
q_T_2 = coin_flips_2.count('T') / len(coin_flips_2)
print('Coin flips given dice roll is 2:', coin_flips_2)
print('Distribution of coin flip given dice roll is 2: {{ "H": {}, "T": {}}}'.format(q_H_2, q_T_2))
assert math.isclose(q_H_2 + q_T_2, 1)
# + [markdown] slideshow={"slide_type": "fragment"}
# Finally, we want to store the conditional distribution as a nested dictionary so that `q[i]` points to the distribution
#
# $$[q_{j|i}]_{j\in \mathcal{T}} \kern1em \text{for }i\in \mathcal{S}.$$
# + slideshow={"slide_type": "-"}
q = {}
q[1] = dict(zip('HT',(q_H_1, q_T_1)))
q[2] = dict(zip('HT',(q_H_2, q_T_2)))
q
# + [markdown] slideshow={"slide_type": "fragment"}
# Of course, the above dictionary is missing the entries for other possible outcomes of the dice rolls.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Define a function `conditionally_distribute` that
# - takes two sequences `seq1` and `seq2` of outcomes of the same length, and
# - returns the conditional distribution of `seq2` given `seq1` in the form of a nested dictionary efficiently without storing the unobserved outcomes.
#
# In the above example, `seq1` is `dice_rolls` while `seq2` is `coin_flips`.
#
# *Hint:* For an efficient implementation without traversing the input sequences too many times, consider using the following solution template and the `setdefault` method.
# ```Python
# def conditionally_distribute(seq1, seq2):
# q, count = {}, {} # NOT q = count = {}
# for i in seq1:
# count[i] = count.get(i, 0) + 1
# for i, j in zip(seq1, seq2):
# q[i][j] = ____________________________________________________
# return q
# ```
# + nbgrader={"grade": false, "grade_id": "conditionally_distribute", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def conditionally_distribute(seq1, seq2):
'''Returns the conditional distribution q of seq2 given seq1 such that
q[i] is a dictionary for observed outcome i in seq1 and
q[i][j] is the probability of observing j in seq2 given the
corresponding outcome in seq1 is i.'''
### BEGIN SOLUTION
q, count = {}, {} # NOT q = count = {}
for i in seq1:
count[i] = count.get(i, 0) + 1
for i, j in zip(seq1, seq2):
q[i][j] = q.setdefault(i, {}).setdefault(j, 0) + 1 / count[i]
return q
### END SOLUTION
# + nbgrader={"grade": true, "grade_id": "test-conditionally_distribute", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# tests
cf_given_dr_dist = {
4: {
'T': 0.5588235294117647,
'H': 0.4411764705882353
},
1: {
'T': 0.46511627906976744,
'H': 0.5348837209302325
},
3: {
'H': 0.5135135135135135,
'T': 0.4864864864864865
},
6: {
'H': 0.5454545454545454,
'T': 0.45454545454545453
},
2: {
'T': 0.7586206896551724,
'H': 0.2413793103448276
},
5: {
'T': 0.5416666666666666,
'H': 0.4583333333333333
}
}
cf_given_dr_ans = conditionally_distribute(dice_rolls, coin_flips)
assert all(
math.isclose(cf_given_dr_ans[i][j], v)
for i, d in cf_given_dr_dist.items() for j, v in d.items())
### BEGIN HIDDEN TESTS
dr_given_cf_ans = conditionally_distribute(coin_flips, dice_rolls)
dr_given_cf_dist = {
'T': {
4: 0.17757009345794392,
1: 0.18691588785046728,
2: 0.205607476635514,
5: 0.12149532710280374,
3: 0.16822429906542055,
6: 0.14018691588785046
},
'H': {
3: 0.20430107526881722,
6: 0.1935483870967742,
4: 0.16129032258064516,
5: 0.11827956989247312,
1: 0.24731182795698925,
2: 0.07526881720430108
}
}
assert all(
math.isclose(dr_given_cf_ans[i][j], v)
for i, d in dr_given_cf_dist.items() for j, v in d.items())
### END HIDDEN TESTS
# + [markdown] slideshow={"slide_type": "fragment"}
# The [*conditional entropy*](https://en.wikipedia.org/wiki/Conditional_entropy) is defined for a conditional distribution $\mathbf{q}=[q_{j|i}]_{i\in \mathcal{S},j\in\mathcal{T}}$ and a distribution $\mathbf{p}=[p_i]_{i\in \mathcal{S}}$ as follows:
#
# $$ H(\mathbf{q}|\mathbf{p}) = \sum_{i\in \mathcal{S}} p_i \sum_{j\in \mathcal{T}} q_{j|i} \log_2 \frac{1}{q_{j|i}}, $$
# where, by convention,
# - the summand of the outer sum is 0 if $p_i=0$ (regardless of the values of $q_{j|i}$), and
# - the summand of the inner sum is 0 if $q_{j|i}=0$.
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Define a function `conditional_entropy` that
# - takes
# - a distribution p as its first argument,
# - a conditional distribution q as its second argument, and
# - returns the conditional entropy $H(\mathbf{q}|\mathbf{p})$.
#
# Handle the cases when $p_i=0$ and $q_{j|i}=0$ as well.
# + nbgrader={"grade": false, "grade_id": "conditional_entropy", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def conditional_entropy(p, q):
'''Returns the conditional entropy of the conditional distribution q given
the distribution p.'''
result = 0
for i in q:
result+= p[i]*entropy(q[i])
return result
# + nbgrader={"grade": true, "grade_id": "test-conditional_entropy", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# tests
cf_given_dr_dist = {
4: {
'T': 0.5588235294117647,
'H': 0.4411764705882353
},
1: {
'T': 0.46511627906976744,
'H': 0.5348837209302325
},
3: {
'H': 0.5135135135135135,
'T': 0.4864864864864865
},
6: {
'H': 0.5454545454545454,
'T': 0.45454545454545453
},
2: {
'T': 0.7586206896551724,
'H': 0.2413793103448276
},
5: {
'T': 0.5416666666666666,
'H': 0.4583333333333333
}
}
assert conditional_entropy({'H': 0.5, 'T': 0.5},{'H': {'H': 0.5, 'T': 0.5}, 'T': {'H': 0.5, 'T': 0.5}}) == 1
assert conditional_entropy({'H': 0, 'T': 1},{'H': {'H': 0.5, 'T': 0.5}, 'T': {'H': 0.5, 'T': 0.5}}) == 1
assert conditional_entropy({'H': 0.5, 'T': 0.5},{'H': {'H': 1, 'T': 0}, 'T': {'H': 0, 'T': 1}}) == 0
assert conditional_entropy({'H': 0.5, 'T': 0.5},{'H': {'H': 1, 'T': 0}, 'T': {'H': 0.5, 'T': 0.5}}) == 0.5
assert math.isclose(conditional_entropy(dice_rolls_dist, cf_given_dr_dist), 0.9664712793722372)
### BEGIN HIDDEN TESTS
dr_given_cf_dist = {
'T': {
4: 0.17757009345794392,
1: 0.18691588785046728,
2: 0.205607476635514,
5: 0.12149532710280374,
3: 0.16822429906542055,
6: 0.14018691588785046
},
'H': {
3: 0.20430107526881722,
6: 0.1935483870967742,
4: 0.16129032258064516,
5: 0.11827956989247312,
1: 0.24731182795698925,
2: 0.07526881720430108
}
}
assert conditional_entropy({'H': 1, 'T': 0},{'H': {'H': 0.5, 'T': 0.5}, 'T': {'H': 0.5, 'T': 0.5}}) == 1
assert conditional_entropy({'H': 0.5, 'T': 0.5},{'H': {'H': 0, 'T': 1}, 'T': {'H': 1, 'T': 0}}) == 0
assert conditional_entropy({'H': 0.5, 'T': 0.5},{'H': {'H': 0.5, 'T': 0.5}, 'T': {'H': 0, 'T': 1}}) == 0.5
assert math.isclose(conditional_entropy(coin_flips_dist, dr_given_cf_dist), 2.531672712273041)
### END HIDDEN TESTS
# + [markdown] slideshow={"slide_type": "subslide"}
# The joint probability $p_{ij}$ over $i\in \mathcal{S}$ and $j\in \mathcal{T}$ can be calculated as follows
#
# $$p_{ij} = p_{i} q_{j|i}$$
# where $p_i$ is the probability of $i$ and $q_{j|i}$ is the conditional probability of $j$ given $i$.
# -
# **Exercise** Define a function `joint_distribution` that
# - takes the distribution $p$ and conditional distribution $q$ as arguments, and
# - returns their joint distribution.
# + nbgrader={"grade": false, "grade_id": "joint_distribution", "locked": false, "schema_version": 3, "solution": true, "task": false}
def joint_distribution(p,q):
### BEGIN SOLUTION
return {(i,j):p[i]*q[i][j] for i in p for j in q[i]}
### END SOLUTION
# + nbgrader={"grade": true, "grade_id": "test-joint_distribution", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# tests
assert joint_distribution({'H': 0.5, 'T': 0.5},{'H': {'H': 0.5, 'T': 0.5}, 'T': {'H': 0.5, 'T': 0.5}}) == {('H', 'H'): 0.25, ('H', 'T'): 0.25, ('T', 'H'): 0.25, ('T', 'T'): 0.25}
assert joint_distribution({'H': 0, 'T': 1},{'H': {'H': 0.5, 'T': 0.5}, 'T': {'H': 0.5, 'T': 0.5}}) == {('H', 'H'): 0.0, ('H', 'T'): 0.0, ('T', 'H'): 0.5, ('T', 'T'): 0.5}
assert joint_distribution({'H': 0.5, 'T': 0.5},{'H': {'H': 1, 'T': 0}, 'T': {'H': 0, 'T': 1}}) == {('H', 'H'): 0.5, ('H', 'T'): 0.0, ('T', 'H'): 0.0, ('T', 'T'): 0.5}, {'H': 0.5, 'T': 0.5}
### BEGIN HIDDEN TESTS
assert joint_distribution({'H': 1, 'T': 0},{'H': {'H': 0.5, 'T': 0.5}, 'T': {'H': 0.5, 'T': 0.5}}) == {('H', 'H'): 0.5, ('H', 'T'): 0.5, ('T', 'H'): 0.0, ('T', 'T'): 0.0}
assert joint_distribution({'H': 0.5, 'T': 0.5},{'H': {'H': 0, 'T': 1}, 'T': {'H': 1, 'T': 0}}) == {('H', 'H'): 0.0, ('H', 'T'): 0.5, ('T', 'H'): 0.5, ('T', 'T'): 0.0}
### END HIDDEN TESTS
# + [markdown] slideshow={"slide_type": "subslide"}
# Finally, a fundamental information identity, called the [*chain rule*](https://en.wikipedia.org/wiki/Conditional_entropy#Chain_rule), is that the joint entropy is equal to
#
# $$ H(\mathbf{p}) + H(\mathbf{q}|\mathbf{p})$$
# for any distribution $\mathbf{p}$ over outcome $i\in \mathcal{S}$ and conditional distribution $\mathbf{q}$ over outcome $j\in \mathcal{T}$ given outcome $i \in \mathcal{S}$.
# + [markdown] slideshow={"slide_type": "fragment"}
# If you have implemented `jointly_distribute`, `conditionally_distribute`, `entropy`, and `conditional_entropy` correctly, we can verify the identity as follows.
# + slideshow={"slide_type": "-"}
def validate_chain_rule(seq1,seq2):
p = distribute(seq1)
q = conditionally_distribute(seq1,seq2)
pq = jointly_distribute(seq1,seq2)
H_pq = entropy(pq)
H_p = entropy(p)
H_q_p = conditional_entropy(p,q)
print(f'''Entropy of seq1: {H_p}
Conditional entropy of seq2 given seq1: {H_q_p}
Sum of the above entropies: {H_p + H_q_p}
Joint entropy: {H_pq}''')
assert math.isclose(H_pq,H_p + H_q_p)
# + slideshow={"slide_type": "-"}
validate_chain_rule(coin_flips,coin_flips)
# + slideshow={"slide_type": "-"}
validate_chain_rule(dice_rolls,coin_flips)
| source/Lab8/Information Theory.ipynb |
# Transformers installation
# ! pip install transformers datasets
# To install from source instead of the last release, comment the command above and uncomment the following one.
# # ! pip install git+https://github.com/huggingface/transformers.git
# # Perplexity of fixed-length models
# Perplexity (PPL) is one of the most common metrics for evaluating language models. Before diving in, we should note
# that the metric applies specifically to classical language models (sometimes called autoregressive or causal language
# models) and is not well defined for masked language models like BERT (see [summary of the models](https://huggingface.co/docs/transformers/master/en/model_summary)).
#
# Perplexity is defined as the exponentiated average negative log-likelihood of a sequence. If we have a tokenized
# sequence $X = (x_0, x_1, \dots, x_t)$, then the perplexity of $X$ is,
#
# $$\text{PPL}(X) = \exp \left\{ {-\frac{1}{t}\sum_i^t \log p_\theta (x_i|x_{<i}) } \right\}$$
#
# where $\log p_\theta (x_i|x_{<i})$ is the log-likelihood of the ith token conditioned on the preceding tokens $x_{<i}$ according to our model. Intuitively, it can be thought of as an evaluation of the model's ability to predict uniformly among the set of specified tokens in a corpus. Importantly, this means that the tokenization procedure has a direct impact on a model's perplexity which should always be taken into consideration when comparing different models.
#
# This is also equivalent to the exponentiation of the cross-entropy between the data and model predictions. For more
# intuition about perplexity and its relationship to Bits Per Character (BPC) and data compression, check out this
# [fantastic blog post on The Gradient](https://thegradient.pub/understanding-evaluation-metrics-for-language-models/).
# ## Calculating PPL with fixed-length models
# If we weren't limited by a model's context size, we would evaluate the model's perplexity by autoregressively
# factorizing a sequence and conditioning on the entire preceding subsequence at each step, as shown below.
#
# <img width="600" alt="Full decomposition of a sequence with unlimited context length" src="https://huggingface.co/datasets/huggingface/documentation-images/resolve/main/ppl_full.gif"/>
#
# When working with approximate models, however, we typically have a constraint on the number of tokens the model can
# process. The largest version of [GPT-2](https://huggingface.co/docs/transformers/master/en/model_doc/gpt2), for example, has a fixed length of 1024 tokens, so we
# cannot calculate $p_\theta(x_t|x_{<t})$ directly when $t$ is greater than 1024.
#
# Instead, the sequence is typically broken into subsequences equal to the model's maximum input size. If a model's max
# input size is $k$, we then approximate the likelihood of a token $x_t$ by conditioning only on the
# $k-1$ tokens that precede it rather than the entire context. When evaluating the model's perplexity of a
# sequence, a tempting but suboptimal approach is to break the sequence into disjoint chunks and add up the decomposed
# log-likelihoods of each segment independently.
#
# <img width="600" alt="Suboptimal PPL not taking advantage of full available context" src="https://huggingface.co/datasets/huggingface/documentation-images/resolve/main/ppl_chunked.gif"/>
#
# This is quick to compute since the perplexity of each segment can be computed in one forward pass, but serves as a poor
# approximation of the fully-factorized perplexity and will typically yield a higher (worse) PPL because the model will
# have less context at most of the prediction steps.
#
# Instead, the PPL of fixed-length models should be evaluated with a sliding-window strategy. This involves repeatedly
# sliding the context window so that the model has more context when making each prediction.
#
# <img width="600" alt="Sliding window PPL taking advantage of all available context" src="https://huggingface.co/datasets/huggingface/documentation-images/resolve/main/ppl_sliding.gif"/>
#
# This is a closer approximation to the true decomposition of the sequence probability and will typically yield a more
# favorable score. The downside is that it requires a separate forward pass for each token in the corpus. A good
# practical compromise is to employ a strided sliding window, moving the context by larger strides rather than sliding by
# 1 token a time. This allows computation to proceed much faster while still giving the model a large context to make
# predictions at each step.
# ## Example: Calculating perplexity with GPT-2 in 🤗 Transformers
# Let's demonstrate this process with GPT-2.
from transformers import GPT2LMHeadModel, GPT2TokenizerFast
device = 'cuda'
model_id = 'gpt2-large'
model = GPT2LMHeadModel.from_pretrained(model_id).to(device)
tokenizer = GPT2TokenizerFast.from_pretrained(model_id)
# We'll load in the WikiText-2 dataset and evaluate the perplexity using a few different sliding-window strategies. Since
# this dataset is small and we're just doing one forward pass over the set, we can just load and encode the entire
# dataset in memory.
from datasets import load_dataset
test = load_dataset('wikitext', 'wikitext-2-raw-v1', split='test')
encodings = tokenizer('\n\n'.join(test['text']), return_tensors='pt')
# With 🤗 Transformers, we can simply pass the `input_ids` as the `labels` to our model, and the average negative
# log-likelihood for each token is returned as the loss. With our sliding window approach, however, there is overlap in
# the tokens we pass to the model at each iteration. We don't want the log-likelihood for the tokens we're just treating
# as context to be included in our loss, so we can set these targets to `-100` so that they are ignored. The following
# is an example of how we could do this with a stride of `512`. This means that the model will have at least 512 tokens
# for context when calculating the conditional likelihood of any one token (provided there are 512 preceding tokens
# available to condition on).
# +
import torch
from tqdm import tqdm
max_length = model.config.n_positions
stride = 512
nlls = []
for i in tqdm(range(0, encodings.input_ids.size(1), stride)):
begin_loc = max(i + stride - max_length, 0)
end_loc = min(i + stride, encodings.input_ids.size(1))
trg_len = end_loc - i # may be different from stride on last loop
input_ids = encodings.input_ids[:,begin_loc:end_loc].to(device)
target_ids = input_ids.clone()
target_ids[:,:-trg_len] = -100
with torch.no_grad():
outputs = model(input_ids, labels=target_ids)
neg_log_likelihood = outputs[0] * trg_len
nlls.append(neg_log_likelihood)
ppl = torch.exp(torch.stack(nlls).sum() / end_loc)
# -
# Running this with the stride length equal to the max input length is equivalent to the suboptimal, non-sliding-window
# strategy we discussed above. The smaller the stride, the more context the model will have in making each prediction,
# and the better the reported perplexity will typically be.
#
# When we run the above with `stride = 1024`, i.e. no overlap, the resulting PPL is `19.64`, which is about the same
# as the `19.93` reported in the GPT-2 paper. By using `stride = 512` and thereby employing our striding window
# strategy, this jumps down to `16.53`. This is not only a more favorable score, but is calculated in a way that is
# closer to the true autoregressive decomposition of a sequence likelihood.
| notebooks/transformers_doc/tensorflow/perplexity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import random
suits = ('Hearts', 'Diamonds', 'Spades', 'Clubs')
ranks = ('Two', 'Three', 'Four', 'Five', 'Six', 'Sevem', 'Eight', 'Nine', 'Ten', 'Jack', 'Queen', 'King', 'Ace')
values = {'Two': 2, 'Three': 3, 'Four': 4, 'Five': 5, 'Six': 6, 'Sevem': 7, 'Eight': 8, 'Nine': 9, 'Ten': 10, 'Jack': 11, 'Queen': 12, 'King': 13, 'Ace': 14}
playing = True
# -
class Card:
def __init__(self, suit, rank):
self.suit = suit
self.rank = rank
def __str__(self):
return self.rank + ' Of ' + self.suit
class Deck:
def __init__(self):
self.deck = []
for suit in suits:
for rank in ranks:
self.deck.append(Card(suit, rank))
def __str__(self):
deck_comp = ''
for card in self.deck:
deck_comp += '\n' + card.__str__()
return "The deck has: " + deck_comp
def shuffle(self):
random.shuffle(self.deck)
def deal(self):
single_card = self.deck.pop()
return single_card
test_deck = Deck()
print(test_deck)
test_duck = Deck()
test_duck.shuffle()
print(test_duck)
class Hand:
def __init__(self):
self.cards = []
self.value = 0
self.aces = 0
def add_card(self, card):
self.cards.append(card)
self.value += values[card.rank]
if card.rank == 'Ace':
self.aces += 1
def adjust_for_ace(self):
while self.value > 21 and self.aces:
self.value -= 10
self.aces -= 1
# +
test_deck = Deck()
test_deck.shuffle()
test_player = Hand()
pulled_card = test_deck.deal()
print(pulled_card)
test_player.add_card(pulled_card)
print(test_player.value)
# -
class Chips:
def __init__(self):
self.total = 100
self.bet = 0
def win_bet(self):
self.total += self.bet
def lose_bet(self):
self.total -= self.bet
def take_bet(chips):
while True:
try:
chips.bet = int(input('How many chips you would like to bet?: '))
except ValueError:
print('Sorry, Your bet must be integer')
else:
if chips.bet > chips.total:
print('Sorry Pal, You do not have enough chips! You have: {}'.format(chips.total))
else:
break
def hit(deck, hand):
hand.add_card(deck.deal())
hand.adjust_for_ace()
def hit_or_stand(deck, hand):
global playing
while True:
x = input("Would you like to Hit or Stand? Enter 'h' or 's' ")
if x[0].lower() == 'h':
hit(deck, hand)
elif x[0].lower() == 's':
print("Player Stands. Dealer is playing")
playing = False
else:
print("Sorry, Please Try Again !!")
continue
break
# +
def player_busts(player, dealer, chips):
print("Player Busts!!")
chips.lose_bet()
def player_wins(player, dealer, chips):
print("Player Wins")
chips.win_bet()
def dealer_busts(player, dealer, chips):
print('Player Wins! Dealer Busted')
chips.win_bet()
def dealer_wins(player, dealer, chips):
print("Dealer Wins")
chips.lose_bet()
def push(player, dealer):
print("Dealer and Player, Tie!! Push")
# +
def show_some(player, dealer):
print("\nDealer's Hand:")
print(" <card hidden>")
print('', dealer.cards[1])
print("\nPlayer's Hand:", *player.cards, sep = '\n ')
def show_all(player, dealer):
print("\nDealer's Hand:", *dealer.cards, sep = '\n ')
print("Dealer's Hand = ", dealer.value)
print("\nPlayer's Hand:", *player.cards, sep='\n ')
print("\nPlayer's Hand = ", player.value)
# -
while True:
print("Welcome to BlackJack")
deck = Deck()
deck.shuffle()
player_hand = Hand()
player_hand.add_card(deck.deal())
player_hand.add_card(deck.deal())
dealer_hand = Hand()
dealer_hand.add_card(deck.deal())
dealer_hand.add_card(deck.deal())
player_chips = Chips()
take_bet(player_chips)
show_some(player_hand, dealer_hand)
while playing:
hit_or_stand(deck, player_hand)
show_some(player_hand, dealer_hand)
if player_hand.value > 21:
player_busts(player_hand, dealer_hand, player_chips)
break
if player_hand.value <= 21:
while dealer_hand.value < 17:
hit(deck, dealer_hand)
show_all(player_hand, dealer_hand)
if dealer_hand.value > 21:
dealer_busts(player_hand, dealer_hand, player_chips)
elif dealer_hand.value > player_hand.value:
dealer_wins(player_hand, dealer_hand, player_chips)
elif dealer_hand.value < player_hand.value:
player_wins(player_hand, dealer_hand, player_chips)
else:
push(player_hand, dealer_hand)
print('\n Player total chips are: {}'.format(player_chips.total))
new_game = input("Would you like to play another hand? Y/N")
if new_game[0].lower() == 'y':
playing = True
continue
else:
print('Thanks for playing')
break
| Jose Portilla/Python Bootcamp Go from Zero to Hero in Python 3/Python Milestoen Project 2/Black Jack.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PaddlePaddle 2.0.0b0 (Python 3.5)
# language: python
# name: py35-paddle1.2.0
# ---
# # Transform In Transformer 介绍
#
# **TNT(Transform In Transformer)** 是继ViT 和 DeiT之后的优异vision transformer(视觉transformer)。在视觉任务上有较好性能。
#
# 整体上来说,TNT较以前的模型在transformer处理上有更多的细节上的提升。
#
# 提升点:
#
# 1. **patch-level + pixel-level** 两级结合,即利用Attention特性对patch-level上图像的**全局特征**进行高质量提取,同时又利用pixel-level对全局下的**局部特征**进行进一步提取,保证了图片的较为完整空间关系。
#
# 2. **position encode** 进行位置编码,保证图片在split时空间结构被较好地保留下来,这是以前的视觉transformer不具有的。
#
# 3. pixel-level的embedding实现嵌入后还将继续嵌入到patch-level的embedding中。
#
# 4. embedding的结果会与position encode进行结合,保证图片特征提取过程中**完整的空间结构**。
#
# 严谨性:
#
# 1. 利用消融实验,对**head数,position encode,two-level**的必要性进行了实验证明。
#
#
# > 下面就TNT复现代码进行讲解,适当的补充TNT体系结构的说明。
# ## 完整实现代码
#
# -- models
#
# * tnt_layers.py: TNT模型所涉及到的组件网络层实现 + 代码注释
#
# * tnt_model.py: TNT模型实现、与基本small、big模型的配置 + 代码注释
#
# * tset.py: 提供TNT模型的简单使用方法
#
#
# > 下边是一个测试文件,其它信息可前往test.py中查看
# +
# %cd models/
from test import create_tnt_by_basecfg
# on_start_test: True, 表示进行基本测试
model = create_tnt_by_basecfg(num_classes = 10, img_size = 224, in_chans = 3, choice_big=False, on_start_test=True)
# -
# # 一、TNT 模型基本流程解析
#
# (图源:论文)
#
# 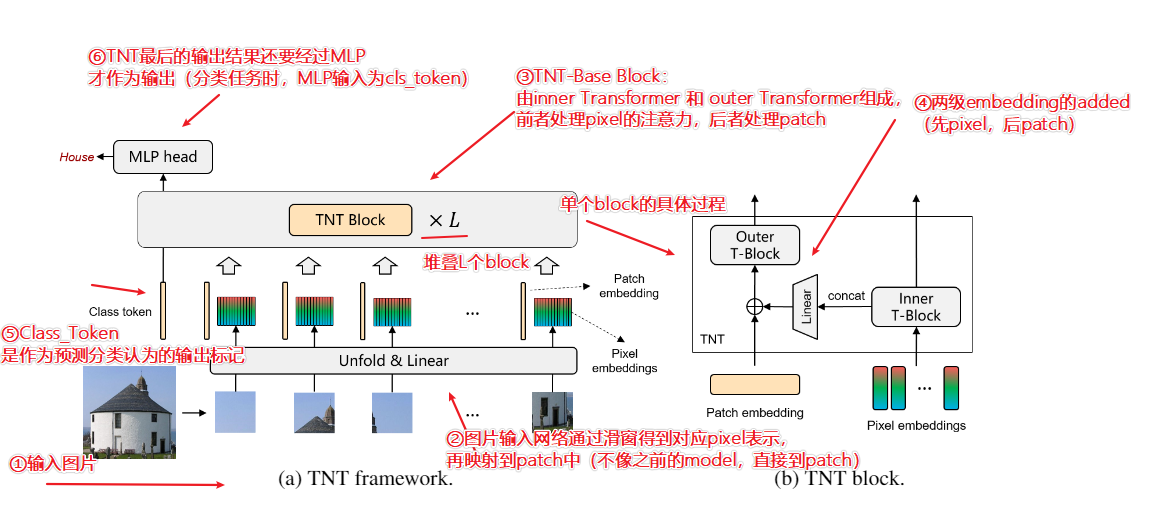
#
# 基本流程如图中所示,基本步骤总结如下:
#
#
# | 步骤 | 组件 | info |
# | -------- | -------- | -------- |
# | 第一步 | Unfold + Conv2D | 将输入图片分割成指定的pixel大小和patch大小 |
# | 第二步 | TNT Block | 堆叠的TNT块将patch与pixel作为输入进行处理 |
# | 第三步 | inner transformer | 在TNT中处理pixel |
# | 第四步 | outer transformer | 在TNT中处理patch |
# | 第五步 | TNT Blocks | 反复第二到第四步,直到第L块运行结束 |
# | 第六步 | MLP head | 将TNT Blocks输出中的class_token部分作为输入,从而得到分类任务的输出结果 |
#
# # 二、TNT代码解析
#
# 就上边所说的流程进行代码复现,为了描述完整性,按照以下顺序介绍:
#
# 1. Attention部分
#
# 2. MLP部分
#
# 3. DropPath部分(添加的丢弃策略)
#
# 4. TNT Block部分
#
# 5. Pixel Embed部分
#
# 以上顺序与表格中略有不同,增添部分为论文代码实现的一些策略。
#
# > 具体内容,将在代码中去介绍,每一个部分的代码前边,会有对该部分代码的主要解析,与参数说明
# ## 0. 基本的依赖库
# +
import paddle # 提供数据操作方法
from paddle import nn # 网络层API
from paddle.nn import functional as F # 常见方法
import math
import os
# -
# ## 1. Attention部分
#
# 注意力部分,主要是通过对输入数据进行注意力编码,然后将注意力结果叠加到原输入数据上。
#
# Attention前后数据形状不发生改变,仅发生数据值变化。
#
# 主要实现思路:
#
# 1. 将输入进行指定维度的线性映射(Linear)
#
# 2. 第一次映射,同时获取questions 和 keys的映射数据
#
# 3. 然后分离两者,并将两者进行矩阵乘积,然后通过softmax,计算question在key上的注意力结果
#
# 4. 第二次映射,获取values的映射数据
#
# 5. question与key的注意力结果与value进行矩阵乘积,将注意力作用到value上
#
# 6. 最后将value映射回输入大小,适当丢弃,最后输出
#
#
# > 下边是代码实现 + 代码注释
class Attention(nn.Layer):
'''
Multi-Head Attention
'''
def __init__(self, in_dims, hidden_dims, num_heads=8, qkv_bias=False, attn_drop=0., proj_drop=0., logging=False):
super(Attention, self).__init__()
''' Attention
params list:
in_dims: 输入维度大小
hidden_dim: 隐藏层维度大小
num_heads: 注意力头数量
qkv_bias: 是否对question、keys、values开启映射
attn_drop: 注意力丢弃率
proj_drop: 映射丢弃率
logging: 是否输出Attention的init参数日志
'''
# 确保输入层、隐藏层维度为偶数,且不为零,否则在头划分映射时会发生大小错误
assert in_dims % 2 == 0 and in_dims != 0, \
'please make sure the input_dims(now: {0}) is an even number.(%2==0 and !=0)'.format(in_dims)
assert hidden_dims % 2 == 0 and hidden_dims != 0, \
'please make sure the hidden_dims(now: {0}) is an even number.(%2==0 and !=0)'.format(hidden_dims)
self.in_dims = in_dims # ATT输入大小
self.hidden_dims = hidden_dims # ATT隐藏层大小
self.num_heads = num_heads # ATT的头数目
self.head_dims = hidden_dims // num_heads # 将ATT隐藏层大小按照头数平分,作为ATT-head的维度大小
self.scale = self.head_dims ** -0.5 # 缩放比例按照头的唯独大小进行开(-0.5次幂)
self.qkv_bias = qkv_bias
self.attn_drop = attn_drop
self.proj_drop = proj_drop
# 输出日志信息
if logging:
print('\n—— Attention Init-Logging ——')
print('{0:20}'.format(list(dict(in_dims=self.in_dims).keys())[0]), ': {0:12}'.format(self.in_dims))
print('{0:20}'.format(list(dict(hidden_dims=self.hidden_dims).keys())[0]), ': {0:12}'.format(self.hidden_dims))
print('{0:20}'.format(list(dict(num_heads=self.num_heads).keys())[0]), ': {0:12}'.format(self.num_heads))
print('{0:20}'.format(list(dict(head_dims=self.head_dims).keys())[0]), ': {0:12}'.format(self.head_dims))
print('{0:20}'.format(list(dict(scale=self.scale).keys())[0]), ': {0:12}'.format(self.scale))
print('{0:20}'.format(list(dict(qkv_bias=self.qkv_bias).keys())[0]), ': {0:12}'.format(self.qkv_bias))
print('{0:20}'.format(list(dict(attn_drop=self.attn_drop).keys())[0]), ': {0:12}'.format(self.attn_drop))
print('{0:20}'.format(list(dict(proj_drop=self.proj_drop).keys())[0]), ': {0:12}'.format(self.proj_drop))
'''
questions + keys | values
project layers
'''
# questions + keys 的映射层: *2 就是在一次操作下将两层一同映射
# qkv_bias:是否开启bias --> 开启默认全零初始化
self.qk = nn.Linear(self.in_dims, self.hidden_dims*2, bias_attr=qkv_bias)
# values 的映射层
# qkv_bias:是否开启bias --> 开启默认全零初始化
self.v = nn.Linear(self.in_dims, self.in_dims, bias_attr=qkv_bias)
# ATT的丢弃层
self.attn_drop = nn.Dropout(attn_drop)
'''
注意力结果映射层
'''
self.proj = nn.Linear(self.in_dims, self.in_dims)
self.proj_drop = nn.Dropout(proj_drop)
@paddle.jit.to_static
def forward(self, inputs):
x = inputs
B, N, C= x.shape # B:batch_size, N:patch_number, C:input_channel
# print('\n—— Attention Forward-Logging ——')
# print('{0:20}'.format(list(dict(B=B).keys())[0]), ': {0:12}'.format(B))
# print('{0:20}'.format(list(dict(N=N).keys())[0]), ': {0:12}'.format(N))
# print('{0:20}'.format(list(dict(C=C).keys())[0]), ': {0:12}'.format(C))
# 利用输入映射question 和 keys维度的特征
# print('input(x): ', x.numpy().shape)
qk = self.qk(x) # 将输入映射到question + keys上
# print('qk_project: ', qk.numpy().shape)
qk = paddle.reshape(qk, shape=(B, N, 2, self.num_heads, self.head_dims)) # 将question + keys分离
# print('qk_reshape: ', qk.numpy().shape)
qk = paddle.transpose(qk, perm=[2, 0, 3, 1, 4]) # 重新排列question和keys的数据排布
# print('qk_transpose: ', qk.numpy().shape)
'''
①上面实现的划分,正好对应: head_dims = hidden_dims // num_heads
②排布更新为: 映射类别(question+keys),batch_size, head_number, patch_number, head_dims
'''
q, k = qk[0], qk[1] # 分离question 和 keys
# print('q: ', q.numpy().shape)
# print('k: ', k.numpy().shape)
# 利用输入映射 values 维度的特征
v = self.v(x).reshape(shape=(B, N, self.num_heads, -1)).transpose(perm=(0, 2, 1, 3))
# print('v: ', v.numpy().shape)
# 通过question 与 keys矩阵积,计算patch的注意力结果
attn = paddle.matmul(q, k.transpose(perm=(0, 1, 3, 2))) * self.scale
# print('attn_matrix*: ', attn.numpy().shape)
'''
k.transpose(perm=(0, 1, 3, 2)) : 最后两维发生转置 --> 用于矩阵乘法,实现注意力大小计算(question 对 keys)
* self.scale : 针对注意力头数进行一定的缩放,稳定值
'''
attn = F.softmax(attn, axis=-1) # 通过softmax整体估算注意力 -- 对每一个patch上的hidden_dim进行注意力计算
# print('attn_softmax: ', attn.numpy().shape)
attn = self.attn_drop(attn) # 丢弃部分注意力结果
# 将注意力结果与value进行矩阵乘法结合
x = paddle.matmul(attn, v).transpose(perm=(0, 2, 1, 3)).reshape(shape=(B, N, -1))
# print('x_matrix*: ', x.numpy().shape)
'''
attn 与 v 矩阵乘: 实现注意力叠加
transpose(perm=(0, 2, 1, 3)): 将patch与head维度互换(转置) -- 保证reshape不发生错误合并
reshape(shape=(B, N, -1)): 转换回:batch_size, patch_num, out_dims形式 -- out_dims = num_head * head_dims
'''
x = self.proj(x) # 将注意力叠加完成的结果进行再映射,将其映射回输入大小
# print('x_proj: ', x.numpy().shape)
x = self.proj_drop(x) # 丢弃部分结果
return x
# ## 2. MLP部分
#
# 多层感知机部分是整个过程中最简单的部分,只需要两层线性层(Linear)即可
#
# 主要实现思路:
#
# 1. 确定输入、输出、隐藏层大小
#
# 2. 构建一层输入层,实现输入大小到隐藏大小的映射
#
# 3. 构建一层输出层,实现隐藏大小到输出大小的映射
#
# 4. 构建前向时,输入层后先跟GELU激活函数,再跟丢弃层
#
# 5. 然后经过输出层 + 丢弃层,得到MLP的输出
#
# > 下边是代码实现 + 代码注释
class MLP(nn.Layer):
'''
两层fc的感知机
'''
def __init__(self, in_features, hidden_features=None, out_features=None, act_layer=nn.GELU, drop=0.):
super(MLP, self).__init__()
''' MLP
params list:
in_features: 输入大小
hidden_features: 隐藏层大小
out_features: 输出大小
act_layer: 激活层
drop: 丢弃率
'''
# 如果前项为None,则返回后向作为赋值内容
out_features = out_features or in_features
hidden_features = hidden_features or in_features
# 第一层输入
self.fc1 = nn.Linear(in_features, hidden_features)
self.act = act_layer()
# 第二层输出
self.fc2 = nn.Linear(hidden_features, out_features)
self.drop = nn.Dropout()
@paddle.jit.to_static
def forward(self, inputs):
x = inputs
x = self.fc1(x)
x = self.act(x)
x = self.drop(x)
x = self.fc2(x)
x = self.drop(x)
return x
# ## 3. DropPath部分
#
# 路径丢弃,与普通的Dropout相比,路径丢弃使得丢弃的数据更多。
#
# 所谓路径,就是沿着shape中的第一个维度进行一整个的丢弃(全部置为0);
#
# 而普通的丢弃,仅仅对全部参数进行随机的丢弃,而DropPath则是将实现从逐个元素跨越到了整个轴上。
#
# > drop_path: 考虑到大面积丢弃,所以对未丢弃的数据的值进行了一定的扩增
#
# 主要实现思路:
#
# 1. 首先在运行前判断是否进行丢弃 —— drop为0,不丢弃;在训练中,不丢弃
#
# 2. 计算保持率
#
# 3. 得到路径丢弃的随机丢弃分布 -- random_tensor
#
# 4. 扩增数据,并实现丢弃
#
#
# > 下边是代码实现 + 代码注释
class DropPath(nn.Layer):
'''删除路径数据
延一个路径进行丢弃(沿数据第一个维度进行丢弃)
丢弃的对应path下,所有数据置为0
'''
def __init__(self, drop_prob=None):
super(DropPath, self).__init__()
''' DropPath
params list:
drop_prob: 丢弃率
'''
self.drop_prob = drop_prob
@paddle.jit.to_static
def forward(self, inputs):
x = inputs
return self.drop_path(x) # self.training是否在训练模型下
def drop_path(self, x):
'''
具体的path丢弃操作: 改变对应的值,不改变数据形状
'''
if self.drop_prob == 0. or not self.training:
return x
keep_prob = paddle.to_tensor([1 - self.drop_prob])
# 作batch_size维度大小的shape结构--(batch_size, 1, 1, ...)
shape = (x.shape[0], ) + (1, ) * (x.ndim - 1) # batch_size, 1*原ndim减去batch_size维度的大小
random_tensor = keep_prob + paddle.rand(shape=shape, dtype=x.dtype) # 按照划分的shape创建一个[0, 1)均匀分布随机tensor
# 利用[0,1)均匀分布产生的值 + 保持率,就可以实现等比例的保留和丢弃
# 由于随机性,可以保证丢弃的随机性
# 由于值总是在[0,1)间,所以只要得到的值 + keep_prob大于一个阈值,就保留
# 但是因为值时均匀分布的,虽然每一个位置上值时随机取到的,但是确实均匀划分的,
# 因此这样相加后可以实现对应丢弃概率下的丢弃path,并非一定会执行丢弃
# print(keep_prob + paddle.rand(shape=[2,]))
# print(paddle.floor(keep_prob + paddle.rand(shape=[2,])))
random_tensor = paddle.floor(random_tensor) # 将1作为阈值,从而floor向下取整筛选满足的数据
# 仅仅留下 0, 1
# print(random_tensor)
# print(x[0, 0, 0])
# print(keep_prob)
# print(paddle.divide(x, keep_prob)[0, 0, 0])
# print(random_tensor.shape)
# print('x: ', x.numpy())
output = paddle.divide(x, keep_prob) * random_tensor
# print('output: ', output.numpy())
return output
# ## 4. TNT Block部分
#
# TNT Block是TNT模型中的特征提取结构。
#
# TNT Block由Tin 与 Tout组成,利用Tin 对 pixel进行特征提取;利用Tout 对 patch进行特征提取。
#
# 具体流程:先Tin,然后再Tout
#
# 主要实现思路:
#
# 1. 先构建Tin,利用Attention + MLP + Linear实现
#
# 2. 输入的pixel_embed嵌入数据先通过Attention提取,作为当前层的pixel_embed结果,
#
# 3. 再通过MLP进行映射提取,又进行叠加,得到当前block最终的pixel数据
#
# 4. 然后利用Linear 对 pixel进行映射,实现从pixel到patch的映射,并将其加到patch_embed嵌入数据中
#
# 5. 再构建Tout, 只需Attention + MLP
#
# 6. 将上边加过pixel_embed的patch_embed依次通过Attention和MLP,得到Block的输出
#
#
# > 下边是代码实现 + 代码注释
class TNT_Block(nn.Layer):
'''
实现inner transfromer 和 outer transformer, 从pixel-level 和 patch-level进行数据特征提取
特性:
输入输出前后,tensor不发生shape变化(中间过程存在部分映射有shape变化)
'''
def __init__(self, patch_embeb_dim, in_dim, num_pixel, out_num_heads=12,
in_num_head=4, mlp_ratio=4.,qkv_bias=False, drop=0.,
attn_drop=0., drop_path=0., act_layer=nn.GELU, norm_layer=nn.LayerNorm):
'''TNT_Block
params list:
patch_embeb_dim : patch的嵌入维度大小(也是实际输入数据的映射空间大小)
in_dim : 单个patch的维度大小(不包含pixel-level维度)
num_pixel : patch下的(in_dim维度)元素对应pixel的比例 1:num_pixel,即pixel个数(也是论文中指的patch2pixel分辨率)
out_num_heads : 输出(outer attn)的注意力头
in_num_head : 输入(inner attn)的注意力头
mlp_ratio : outer transformer中感知机隐藏层的维度缩放率
qkv_bias : question 、 keys 、values对应的(线性)映射层bias启用标记
drop : MLP部分、proj部分的丢弃率
attn_drop : attn部分的丢弃率
drop_path : 路径丢弃的丢弃率
act_layer : 激活层
norm_layer : 归一化层
'''
super(TNT_Block, self).__init__()
# Inner transformer
# 输入-注意力计算 -- pixel level
self.in_attn_norm = norm_layer(in_dim) # 层归一化--attention的归一化层
self.in_attn = Attention(in_dims=in_dim, hidden_dims=in_dim,
num_heads=in_num_head, qkv_bias=qkv_bias,
attn_drop=attn_drop, proj_drop=drop) # attention输出,tensor的shape不变
# 输入-多层感知机进行维度映射
self.in_mlp_norm = norm_layer(in_dim) # 层归一化--mlp的归一化层
self.in_mlp = MLP(in_features=in_dim, hidden_features=int(in_dim*4),
out_features=in_dim, act_layer=act_layer, drop=drop) # mlp输出,tensor的shape不变
# 输入-线性映射输出
self.in_proj_norm = norm_layer(in_dim) # 层归一化--proj的归一化层
self.in_proj = nn.Linear(in_dim * num_pixel, patch_embeb_dim, bias_attr=True) # proj输出,tensor的shape发生改变
# outer transformer
# 输出-注意力计算 -- patch level
self.out_attn_norm = norm_layer(patch_embeb_dim)
self.out_attn = Attention(in_dims=patch_embeb_dim, hidden_dims=patch_embeb_dim,
num_heads=out_num_heads, qkv_bias=qkv_bias,
attn_drop=attn_drop, proj_drop=drop)
self.out_mlp_norm = norm_layer(patch_embeb_dim)
self.out_mlp = MLP(in_features=patch_embeb_dim, hidden_features=int(patch_embeb_dim * mlp_ratio),
out_features=patch_embeb_dim, act_layer=act_layer, drop=drop)
# 公用方法
# 路径丢弃
self.drop_path = DropPath(drop_prob=drop_path) if drop_path > 0. else self.Identity # self.Identity()占位方法,不对数据做任何处理
@paddle.jit.to_static
def forward(self, pixel_embeb, patch_embeb):
'''
params list:
pixel_embeb: 上一个block输出的pixel-level out tensor
patch_embeb: 上一个block输出的patch-level out tensor
'''
# inner work
# 1. 注意力嵌入 added
pixel_embeb = pixel_embeb + self.drop_path(self.in_attn(self.in_attn_norm(pixel_embeb)))
# 2. mlp嵌入 added
pixel_embeb = pixel_embeb + self.drop_path(self.in_mlp(self.in_mlp_norm(pixel_embeb)))
# pixel嵌入的pathc叠加,在outer中完成
# outer work
B, N, C = patch_embeb.shape # B:batch_size N:Patch_Number C:Feature_map_channel
# 线性映射pixel到patch维度,N-1 means;映射前后不包括class_token
# 映射是需要完整映射,不需要路径丢弃
pixel_embeb_proj2patch = self.in_proj(self.in_proj_norm(pixel_embeb).reshape(shape=(B, N-1, -1)))
# patch叠加上pixel的embeb数据,从patch1 --> patchn
# 不在这里操作class_token
patch_embeb[:, 1:] = patch_embeb[:, 1:] + pixel_embeb_proj2patch
# 1. 注意力嵌入 added
patch_embeb = patch_embeb + self.drop_path(self.out_attn(self.out_attn_norm(patch_embeb)))
# print(patch_embeb.shape)
# 2. mlp嵌入 added
patch_embeb = patch_embeb + self.drop_path(self.out_mlp(self.out_mlp_norm(patch_embeb)))
return pixel_embeb, patch_embeb
def Identity(self, x):
'''
do nothing, only return input
'''
return x
# ## 5. Pixel Embed部分
#
# Pixel Embed的像素嵌入部分是整个TNT模型的入口,实现对图像进行合理提取,根据预置的patch大小等参数生成pixel的嵌入数据。
#
# 【这里得到的pixel将可以在后边模型组网中实现从pixel2patch的直接映射——这就是TNT中提出的two-level extract(提取) features】
#
# 主要实现思路:
#
# 1. 根据给定的参数信息,进行一层卷积得到指定通道的特征图 -- 这是的通道数实际上就是单个pathc的维度(在不包含pixel级是的大小)【个人理解,如果有错误,可以在评论区指导一下,谢谢!】
#
# 2. 根据stride,进行卷积提取——实际上卷积核大小与padding刚好可以保证卷积前后图像大小不变,但由于stride,导致特征图缩小了
#
# 3. 再经过一个滑窗函数,进行滑窗展开,得到完整的pixel表示(pixel嵌入结果)
#
# 4. 输出pixel嵌入结果
#
# > 下边是代码实现 + 代码注释
class Pixel_Embed(nn.Layer):
def __init__(self, img_size=224, patch_size=16, in_chans=3, in_dim=48, stride=4):
super(Pixel_Embed, self).__init__()
'''Pixel_Embed: 像素嵌入--完成后才有patch嵌入
params_list:
img_size: 输入图片大小
patch_size: 当前一个patch的预置大小
in_chans: 输入的图像通道数
in_dim: 设定的输入维度 -- 即预定的patch的个数
stride: 分块时,使用卷积、滑窗的步长,决定着patch向下划分pixel时的分辨率(不是patch的分辨率)
'''
self.img_size = img_size
self.num_patches = (img_size // patch_size) ** 2
# 平方解释:img_size是宽时,//patch_size得到一行可划分多少个,而同样的列就有多少个
# 这里考虑完整划分patch的个数
self.in_dim = in_dim # 每一个patch对应的分辨率 -- 即patch-level的分辨率
self.new_patch_size = math.ceil(patch_size / stride) # 向上取整 -- 确定向下划分pixel的分辨率
self.stride = stride # 卷积 + 滑窗的步长
'''
两步实现图像到patch的映射,与patch到pixel的分割
'''
self.proj = nn.Conv2D(in_channels=in_chans, out_channels=self.in_dim,
kernel_size=7, padding=3, stride=self.stride)
# 7 // 2 == 3, padding = 3, conv后会保持原图大小 -- 在stride=1时
self.unfold = F.unfold
# 对输入提取滑动块
@paddle.jit.to_static
def forward(self, inputs, pixel_pos):
x = inputs
B, C, H, W = x.shape
# 验证是否与所需大小一致
assert H == self.img_size and W == self.img_size, \
f"Input image size ({H}*{W}) doesn't match model ({self.img_size}*{self.img_size})."
x = self.proj(x)
x = self.unfold(x, kernel_sizes=self.new_patch_size, strides=self.stride) # 提取滑块,获得对应的滑块结果
# unfold将img转换为(B, Cout, Lout)
# Cout = Channel * kernel_sizes[0] * kernel_sizes[1] , 即每一次滑窗在图片上得到参数个数
# Lout = hout * wout —— 滑动block的个数
# hout,wout 类似卷积在图片对应h,w上的滑动次数
x = paddle.transpose(x, perm=[0, 2, 1]) # to be shape: (B, Lout, Cout)
x = paddle.reshape(x, shape=(B * self.num_patches, self.in_dim, self.new_patch_size, self.new_patch_size))
# 再分解为需要的编码形式: (Batch_size * patch_number, in_dim, new_patch_size, new_patch_size)
# Batch_size * patch_number:将每个batch得到的patch数乘以batch_size得到总的patch数量
# in_dim: 当前设定的输入维度大小
# new_patch_size: 由stride确定的 patch 的分辨率 -- 对应patch下feature map的w,h
x = x + pixel_pos # 加上位置编码
x = paddle.reshape(x, shape=(B * self.num_patches, self.in_dim, -1)) # 拼接pixel-level的元素
x = paddle.transpose(x, perm=[0, 2, 1])
# 转换为(B * self.num_patches, patch_dim2pixel_size, self.in_dim)
# patch_dim2pixel_size: 即原in_dim下所有序列元素的拼接大小
# 原来是,in_dim对应dim下的pixels
# 现在是,每一个pixel对应in_dim的情况
return x
# # 三、TNT模型构建
#
# (图源:论文)
#
# 
#
# 接下来就上面实现的组件代码进行模型构建。
# 主要构建流程:
#
# 1. 首先构建 Pixel Embed, 并搭建从pixel到patch的映射网络, 完成图上第一步和第二步的操作
#
# 2. 创建class_token标记,为分类任务创建标记 -- 初始化采用随机截断正态分布
#
# 3. 创建pixel 与 patch的position encoder,实现论文中的位置编码 -- 初始化采用随机截断正态分布
#
# 4. 构建TNT Blocks,搭建特征提取主要网络 -- 完成图中第三和第四步
#
# 5. 构建head,对TNT Blocks的输出的class_token进行指定任务的输出 -- 实现分类结果的输出 -- 完成后几步
#
# 6. 预测结果并非限定在0-1.之间,实际使用预测,还需添加softmax函数进行预测
class TNT(nn.Layer):
def __init__(self, img_size=224, patch_size=16, in_chans=3,
num_classes=10, embed_dim=768, in_dim=48, depth=12,
out_num_heads=12, in_num_head=4, mlp_ratio=4., qkv_bias=False,
drop_rate=0., attn_drop_rate=0.,drop_path_rate=0.,
norm_layer=nn.LayerNorm, first_stride=4):
super(TNT, self).__init__()
'''TNT
params_list:
img_size: 输入图片大小(提前明确)
patch_size: patch的大小(提前明确)
in_chans: 输入图片通道数
num_classes: 分类类别
embed_dim: 嵌入维度大小 -- 也是总的feature大小
in_dim: 每个patch的维度大小 --
但不是每个patch对应的实际全部元素,
全部元素还要加上in_dim[x]*每一个大小对应划分pixel的数量
depth: 深度(block数量)
out_num_heads: outer transformer的header数
in_num_head: inner transformer的header数
mlp_ratio: outer中mlp的隐藏层缩放比例
qkv_bias: question、keys、values是否启用bias
drop_rate: mlp、一般层(比如:映射输出、位置编码输出时)丢弃率
attn_drop_rate: 注意力丢弃率
drop_path_rate: 路径丢弃率
norm_layer: 归一化层
first_stride: 图片输入提取分块的步长--img --> patch
'''
self.num_classes = num_classes # 分类数
self.embed_dim = embed_dim # 嵌入维度 == 特征数
self.num_features = self.embed_dim # 嵌入维度 == 特征数
# 先完成pixel-level的嵌入
self.pixel_embeb = Pixel_Embed(img_size=img_size, patch_size=patch_size,
in_chans=in_chans, in_dim=in_dim, stride=first_stride)
self.num_patches = self.pixel_embeb.num_patches # 当前pixel等效的实际的patch个数
self.new_patch_size = self.pixel_embeb.new_patch_size # 当前等效的每一个patch对应的pixel的分辨率(w == h)
self.num_pixel = self.new_patch_size ** 2 # 当前每个patch实际划分的分辨率,w*h = w**2 得到patch2pixel的序列大小
# 在进行patch-level嵌入
# 从pixel映射到patch上,要对每一个patch展开为pixel下的数据通过层归一化
self.first_proj_norm_start = norm_layer(self.num_pixel * in_dim) # self.nwe_pixel * in_dim, 即每一个patch对应的全部元素
# 然后映射到指定嵌入维度上
self.first_proj = nn.Linear(self.num_pixel * in_dim, self.embed_dim) # 将全部每一个patch对应的pixel都映射到指定嵌入维度大小的空间
# 在经过一次归一化,输出
self.first_proj_norm_end = norm_layer(self.embed_dim)
# 分类标记
# 截断正态分布来填充初始化cls_token、patch_pos、pixel_pos
self.cls_token = paddle.create_parameter(shape=(1, 1, self.embed_dim), dtype='float32', attr=nn.initializer.TruncatedNormal(std=0.02))
# 位置编码
# patch_position_encode: self.num_patches + 1对应实际patch数目+上边的分类标记
self.patch_pos = paddle.create_parameter(shape=(1, self.num_patches + 1, self.embed_dim), dtype='float32', attr=nn.initializer.TruncatedNormal(std=0.02))
# pixel_position_encode: in_dim对应每一个patch的大小, self.new_patch_size对应patch划分为pixel的分辨率
self.pixel_pos = paddle.create_parameter(shape=(1, in_dim, self.new_patch_size, self.new_patch_size), dtype='float32', attr=nn.initializer.TruncatedNormal(std=0.02))
# 位置编码的丢弃
self.pos_drop = nn.Dropout(p=drop_rate)
# 在TNT中使用了path_drop, 进行路径丢弃
# 为了丢弃更具随机性,提高鲁棒性,进行随机丢弃率的制作 -- 根据深度生成对应数量的丢弃率
drop_path_random_rates = [r.numpy().item() for r in paddle.linspace(0, drop_path_rate, depth)]
# 相同块,采用迭代生成
tnt_blocks = []
for i in range(depth): # 根据深度添加TNT块
tnt_blocks.append(
TNT_Block(patch_embeb_dim=self.embed_dim, in_dim=in_dim, num_pixel=self.num_pixel, # 嵌入大小, patch-level大小,patch2pixel大小
out_num_heads=out_num_heads, in_num_head=in_num_head, mlp_ratio=mlp_ratio, # outer transformer头数,inner transformer头数,感知机隐藏层缩放比
qkv_bias=qkv_bias, drop=drop_rate, attn_drop=attn_drop_rate, # transormer中bias启动情况,映射层的丢弃率, 注意力层的丢弃率
drop_path=drop_path_random_rates[i], norm_layer=norm_layer) # 路径丢弃的丢弃率,归一化层
)
# 放入顺序层结构中
self.tnt_blocks = nn.Sequential(*tnt_blocks) # 输入前后不发生shape变化
# tnt_blocks最后的输出还要经过一层归一化
self.tnt_block_end_norm = norm_layer(self.embed_dim) # 沿用前边最初归一化层的嵌入大小进行归一化设置
# 输出任务结果 -- 这里是利用cls_token进行分类,embed_dim是整个模型的嵌入维度大小,也是cls_token的最后1维度的大小
self.head = nn.Linear(self.embed_dim, self.num_classes) if self.num_classes > 0 else TNT_Block.Identity()
# 初始化网络参数
self._init_weights()
def _init_weights(self):
'''
完成整个网络的初始化工作
'''
for l in self.sublayers():
if isinstance(l, nn.Linear):
# 随即截断正态分布填充初始化
l.weight = paddle.create_parameter(shape = (l.weight.shape), dtype=l.weight.dtype,
name=l.weight.name, attr=nn.initializer.TruncatedNormal(std=0.02))
# bias 默认开启就初始化为0 -- 不指定其它初始化方式时
elif isinstance(l, nn.LayerNorm):
# bias 默认开启就初始化为0 -- 不指定其它初始化方式时
# 常量1.0初始化
l.weight = paddle.create_parameter(shape = (l.weight.shape), dtype=l.weight.dtype,
name=l.weight.name, attr=nn.initializer.Constant(value=1.0))
def get_classfier(self):
'''
用于获取分类任务头进行相应的任务预测、输出等
[在当前任务不是必要的]
'''
return self.head
def reset_classfier(self, num_classes):
'''
用于修改分类任务头的分类数目
'''
self.num_classes = num_classes # 修改模型分类参数
self.head = nn.Linear(self.embeb_dim, self.num_classes) if self.num_classes > 0 else TNT_Block.Identity()
def upstream_forward(self, inputs):
'''
上有任务前向运算:TNT特征提取部分的网络运算
'''
x = inputs
B = x.shape[0] # batch_size
# 1. 先进行pixel-level的嵌入
pixel_embeb = self.pixel_embeb(x, self.pixel_pos)
# 2. 再从pixel-level上升到patch-level的嵌入(即向上映射)
# 依次通过: pixel2patch的layer_norm, 然后进行映射,最后再通过一层layer_norm完成整个映射过程
# 其中输入的pixel_embeb要经过shape变换,将散布在不同in_dim下的参数进行拼接到对应patch下
patch_embeb = self.first_proj_norm_end(self.first_proj(self.first_proj_norm_start(pixel_embeb.reshape(shape=(B, self.num_patches, -1)))))
patch_embeb = paddle.concat([self.cls_token, patch_embeb], axis=1) # 将分类任务标记拼接到patch的嵌入空间中
patch_embeb = patch_embeb + self.patch_pos # 加上位置编码
patch_embeb = self.pos_drop(patch_embeb) # 丢弃一部分编码结果
for tnt_block in self.tnt_blocks: # 将前期处理好的嵌入信息,进行迭代,进行信息地进一步提取
pixel_embeb, patch_embeb = tnt_block(pixel_embeb, patch_embeb)
patch_embeb = self.tnt_block_end_norm(patch_embeb)
return patch_embeb[:, 0] # 0号位置为cls_token对应的位置
@paddle.jit.to_static
def forward(self, inputs):
x = inputs
# 主体前向传播
x = self.upstream_forward(x) # 返回cls_token,用于分类用
# 分类任务
x = self.head(x) # 执行分类
return x
# # 四、简单总结
#
# 复现TNT整体上来说没那么多复杂的设计,但是其Conv+滑窗、position encoder、two-level的设计思想却有一定的吸收难度。
#
# 通过two-level,对全局特征和局部特征进行融合,改善以前的transformer在视觉上的不足。
#
# 利用position encoder巩固图像的空间结构。
#
# 参数调整建议:
#
# 1.in_num_head 尽量不动;
#
# 2.改embeding_size可以按照64的倍数增加;
#
# 3.改动size,dim等参数均要被2整除才可保证模型运行
#
# 4.mlp_ratio 在过拟合时,可以适当缩小
#
# > 具体的代码都在models中,test.py为测试代码
# > 有问题欢迎评论区讨论
#
# > 姓名:蔡敬辉
#
# > 学历:大三(在读)
#
# > 爱好:喜欢参加一些大大小小的比赛,不限于计算机视觉——有共同爱好的小伙伴可以关注一下哦~后期会持续更新一些自制的竞赛baseline和一些竞赛经验分享
#
# > 主要方向:目标检测、图像分割与图像识别--在学习NLP, 正在捣鼓FPGA
#
# > 联系方式:qq:3020889729 微信:cjh3020889729
#
# > 学校:西南科技大学
| tnt_rep.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''anaconda3'': virtualenv)'
# name: python_defaultSpec_1596073614085
# ---
import numpy as np
import pandas as pd
import seaborn as sns
df = pd.read_csv('../../data/book/House-Price.csv', header=0)
# + tags=[]
df.head()
# -
sns.jointplot(df.n_hot_rooms,df.price)
sns.scatterplot(df.n_hot_rooms,df.price)
df[df.n_hot_rooms>80]
| notebooks/lin-reg/PreProcess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lustraka/puzzles/blob/main/AoC2021/AoC_21.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="dDZWBB4Ll10C"
# # Advent of Code Puzzles
# [Advent of Code 2021](https://adventofcode.com/2021) | [reddit/adventofcode](https://www.reddit.com/r/adventofcode/)
# + id="ri3O_1i3l2mo"
import requests
import pandas as pd
import numpy as np
path = 'https://raw.githubusercontent.com/lustraka/puzzles/main/AoC2021/data/'
# + [markdown] id="TnFAd8h1l8NI"
# ## [Day 21](https://adventofcode.com/2021/day/21): <NAME>
# ### Part I
# - **Unknown**: The score of the losing player times the number of times the die was rolled during the game.
# - **Data**: Starting positions of two players.
# - **Condition**:
# - A game board has a circular track containing 10 spaces clockwise.
# - The 100-sided deterministic dice rolls a sequence of numbers from 1 to 100.
# - **Plan**:
# - Define the class Player which has a score, board and a method for moving accross the board while counting score.
# - Define a class Dice with a roll counter and method for rolling thrice.
#
# + id="bCYFkYumjpe7"
class Player:
def __init__(self, starting_position):
self.score = 0
self.board = np.roll(np.arange(1,11), -starting_position+1)
def move(self, steps):
self.board = np.roll(self.board, -steps)
self.score += self.board[0]
def __repr__(self):
return f"Player(space={self.board[0]}, score={self.score})"
class DiracDice:
def __init__(self):
self.dice = np.arange(1,101)
self.roll_counter = 0
def roll_thrice(self):
value = 0
for i in range(3):
value += self.dice[0]
self.roll_counter += 1
self.dice = np.roll(self.dice, -1)
return value
def __repr__(self):
return f"Dice(last_value={self.dice[0]}, roll_counter={self.roll_counter})"
# + id="p1AZWha2tqrg" outputId="93daa575-7586-4ee8-9d27-464d892920ca" colab={"base_uri": "https://localhost:8080/"}
p1, p2 = Player(4), Player(8)
dice = DiracDice()
for i in range(4):
p1.move(dice.roll_thrice()), p2.move(dice.roll_thrice())
p1, p2, dice
# + [markdown] id="uKOUED9LzE6T"
# - **Plan** cont.
# - Players roll, move and score
# - If player 1 or player 2 score reaches at least 1000 the game ends
# + id="JNj0DKPcjpkN" outputId="6a9de20a-c4e2-4642-9418-d995b015bc24" colab={"base_uri": "https://localhost:8080/"}
def solve_part1(sp1, sp2):
# Instantiate players and a dice
p1, p2 = Player(sp1), Player(sp2)
dice = DiracDice()
while True:
p1.move(dice.roll_thrice())
if p1.score > 999:
return p2.score * dice.roll_counter
p2.move(dice.roll_thrice())
if p2.score > 999:
return p1.score * dice.roll_counter
solve_part1(4,8)
# + id="CHiYaZpzhLmP" outputId="bd65a426-34b3-4193-9fd1-c0f3e7b931e0" colab={"base_uri": "https://localhost:8080/"}
solve_part1(7,3)
# + [markdown] id="LaGY7AU0funo"
#
# ### Part II
# - **Unknown**:
# - **Data**:
# - **Condition**:
# -
# -
# - **Plan**:
# -
#
# + id="4NBFFtvGhLfO"
# + id="apnNzI6ehLW3"
# + id="bRRErS4jsoqR"
| AoC2021/AoC_21.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1A.soft - Notions de SQL - correction
#
# Correction des exercices du premier notebooks relié au SQL.
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# %matplotlib inline
# ## Recupérer les données
import os
if not os.path.exists("td8_velib.db3"):
from pyensae import download_data
download_data("td8_velib.zip", website = 'xd')
from pyensae.sql import import_flatfile_into_database
dbf = "td8_velib.db3"
import_flatfile_into_database(dbf, "td8_velib.txt") # 2 secondes
import_flatfile_into_database(dbf, "stations.txt", table="stations") # 2 minutes
# %load_ext pyensae
# %SQL_connect td8_velib.db3
# ## Exercice 1
# %%SQL
SELECT COUNT(*) FROM (
SELECT DISTINCT last_update FROM td8_velib
) ;
# %%SQL
SELECT MIN(last_update), MAX(last_update) FROM td8_velib ;
# ## Exercice 2
# %%SQL
SELECT number, COUNT(*) AS nb
FROM td8_velib
WHERE available_bikes==0 AND last_update >= '2013-09-10 11:30:19'
GROUP BY number
ORDER BY nb DESC
# ## Exercice 3 : plage horaires de cinq minutes où il n'y a aucun vélo disponible
# %%SQL
SELECT nb, COUNT(*) AS nb_station
FROM (
-- requête de l'exercice précédent
SELECT number, COUNT(*) AS nb
FROM td8_velib
WHERE available_bikes==0 AND last_update >= '2013-09-10 11:30:19'
GROUP BY number
)
GROUP BY nb
# ## Exercice 4 : distribution horaire par station et par tranche de 5 minutes
# %%SQL
SELECT A.number, A.heure, A.minute, 1.0 * A.nb_velo / B.nb_velo_tot AS distribution_temporelle
FROM (
SELECT number, heure, minute, SUM(available_bikes) AS nb_velo
FROM td8_velib
WHERE last_update >= '2013-09-10 11:30:19'
GROUP BY heure, minute, number
) AS A
JOIN (
SELECT number, heure, minute, SUM(available_bikes) AS nb_velo_tot
FROM td8_velib
WHERE last_update >= '2013-09-10 11:30:19'
GROUP BY number
) AS B
ON A.number == B.number
--WHERE A.number in (8001, 8003, 15024, 15031) -- pour n'afficher que quelques stations
ORDER BY A.number, A.heure, A.minute
# ## Zones de travail et zones de résidence
# %%SQL --df=df
SELECT number, SUM(distribution_temporelle) AS velo_jour
FROM (
-- requête de l'exercice 4
SELECT A.number, A.heure, A.minute, 1.0 * A.nb_velo / B.nb_velo_tot AS distribution_temporelle
FROM (
SELECT number, heure, minute, SUM(available_bikes) AS nb_velo
FROM td8_velib
WHERE last_update >= '2013-09-10 11:30:19'
GROUP BY heure, minute, number
) AS A
JOIN (
SELECT number, heure, minute, SUM(available_bikes) AS nb_velo_tot
FROM td8_velib
WHERE last_update >= '2013-09-10 11:30:19'
GROUP BY number
) AS B
ON A.number == B.number
)
WHERE heure >= 10 AND heure <= 16
GROUP BY number
df = df.sort_values("velo_jour").reset_index()
df["index"] = range(0, df.shape[0])
df.head()
df.plot(x="index", y="velo_jour")
# ## JOIN avec la table stations et les stations "travail"
#
# On trouve les arrondissements où les stations de vélib sont les plus remplies en journée au centre de Paris.
# %%SQL
SELECT C.number, name, lat, lng, velo_jour FROM
(
-- requête de la partie précédente
SELECT number, SUM(distribution_temporelle) AS velo_jour
FROM (
-- requête de l'exercice 4
SELECT A.number, A.heure, A.minute, 1.0 * A.nb_velo / B.nb_velo_tot AS distribution_temporelle
FROM (
SELECT number, heure, minute, SUM(available_bikes) AS nb_velo
FROM td8_velib
WHERE last_update >= '2013-09-10 11:30:19'
GROUP BY heure, minute, number
) AS A
JOIN (
SELECT number, heure, minute, SUM(available_bikes) AS nb_velo_tot
FROM td8_velib
WHERE last_update >= '2013-09-10 11:30:19'
GROUP BY number
) AS B
ON A.number == B.number
)
WHERE heure >= 10 AND heure <= 16
GROUP BY number
) AS C
INNER JOIN stations
ON C.number == stations.number
| _doc/notebooks/td1a_soft/td1a_sql_correction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import networks data to the 3DCityDB
#
# This notebook shows how the **IntegrCiTy Data Access Layer** (DAL) can be used to import data to the 3DCityDB.
# In this specific example, **geometric data** (2D line diagram) and of a gas network is **extracted** from separate sources (shapefiles), **consolidated** and **stored** in the database.
#
# ## Extracting the data
#
# In this example, we extract and consolidate data from various files.
# In this case, all the date is tored in 2 separate shapefiles:
# * **Gas network pipes**: The positions of all pipes are stored as 2D line diagrams, where each pipe is defined by 2 points (start and end point of a straight pipe).
# * **Gas network nodes**: Nodes are points in space where either a pipe is connected to a network boundary (substation, building) or where two or more pipes are joined.
#
# In this example, both data sets use a consistent set of IDs to refer to pipes and nodes, which makes it possible to easily link the data from both sets.
# For instance, there is a pipe called *gas_pipe83-224*, whose starting and ending points coincide with *gas_node83* and *gas_node224*, respectively.
# By checking the data of these nodes, it is possible to see that this pipe is connected to a building on one side (*gas_node83* is connected to *building126*) and that two other pipes are joined to it (*gas_pipe223-224* and *gas_pipe224-225* via *gas_node224*).
# In real-life applications it might take another pre-processing step to link the available data.
#
# The image below visualizes part of the network data from the shapefile (with the help of [QGIS](https://www.qgis.org/)).
#
# <img src="./img/network.png" style="height:14cm">
#
# Before we start, let's define a few tuples and dicts that will be helpful to collect and process the data from the shapefiles:
# +
from collections import namedtuple
# This tuple and dict are used to collect data common to all nodes.
NodeData = namedtuple( 'NodeData', [ 'level', 'x' ,'y' ] )
nodes = {}
# This tuple and dict are used to collect data common to nodes that represent networks sinks (i.e., connections to building).
SinkData = namedtuple( 'SinkData', [ 'level', 'building_name' ] )
sinks = {}
# This tuple and dict are used to collect data common to nodes that represent networks sources (i.e., connections to substations).
SourceData = namedtuple( 'SourceData', [ 'level', 'p_lim_kw', 'p_pa' ] )
sources = {}
# This tuple and dict are used to collect data about pipes.
PipeData = namedtuple( 'PipeData', [ 'length', 'diameter_m', 'from_node', 'to_node' ] )
pipes = {}
# -
# Like in the previous example, the [Python Shapefile Library (PyShp)](https://pypi.org/project/pyshp/) is used to exatract the shapefile data for the nodes:
# +
import shapefile
import os, os.path
gas_nodes_data = shapefile.Reader( os.path.join( os.getcwd(), '..', '1_data', 'shapefiles', 'gas_network_nodes' ) )
assert( gas_nodes_data.shapeType == shapefile.POINT )
# -
# Let's go through and extract the nodes data.
# We use the tuples and dicts defined above to extract data common to all the nodes as well as data specific to sources and sinks:
for data in gas_nodes_data:
# Retrieve 2D coordinates.
x = data.shape.points[0][0]
y = data.shape.points[0][1]
# Extract data common to all nodes.
nodes[data.record['name']] = NodeData( data.record['level'], x, y )
# Extract data only relevant for sinks and sources.
if data.record['type'] == 'SINK':
sinks[data.record['name']] = SinkData( data.record['level'], data.record['build_id'] )
elif data.record['type'] == 'SRCE':
sources[data.record['name']] = SourceData( data.record['level'], data.record['p_lim_kw'], data.record['p_lim_kw'] )
# Now load the pipes data:
gas_pipes_data = shapefile.Reader( os.path.join( os.getcwd(), '..', '1_data', 'shapefiles', 'gas_network_pipes' ) )
assert( gas_pipes_data.shapeType == shapefile.POLYLINE )
# Now go through and extract the pipes data, using the dict and tuple defined above:
# +
from math import *
for data in gas_pipes_data:
# Retrieve the pipe's start and end point.
p1 = data.shape.points[0]
p2 = data.shape.points[1]
# Calculate the pipe's length.
length = sqrt( pow( p1[0] - p2[0], 2 ) + pow( p1[1] - p2[1], 2) )
# Extract the relevant data.
pipes[data.record['name']] = \
PipeData( length, data.record['diameter_m'], data.record['from_node'], data.record['to_node'] )
# -
# ## Accessing the 3DCityDB through the IntegrCiTy DAL
#
# Like in the previous examples, load [package dblayer](https://github.com/IntegrCiTy/dblayer).
# The following lines import the core of the package (*dblayer*) for accessing the database:
# +
from dblayer import *
connect = PostgreSQLConnectionInfo(
user = 'postgres',
pwd = '<PASSWORD>',
host = 'localhost',
port = '5432',
dbname = 'citydb'
)
db_access = DBAccess()
db_access.connect_to_citydb( connect )
# -
# Specify the spatial reference identifier (SRID) used by the 3DCityDB instance. If you have used the setup scripts for installing the extended 3DCityDB provided as part of package dblayer, then the default SRID is 4326.
srid = 4326
# ## Consolidating the network data and adding it to the database
#
# Networks can be represented in the 3DCityDB with the help of the [Utility Network ADE](https://github.com/TatjanaKutzner/CityGML-UtilityNetwork-ADE) (UNADE).
# This CityGML domain extension provides a very flexible framework to store both **topographical data** (e.g., coordinates and shapes) and **topological data** (e.g., functional connections between network features) of various types of networks.
# However, with flexibility also comes complexity, which makes the usage of the UNADE a non-trivial task for new users.
# To this end, [package dblayer](https://github.com/IntegrCiTy/dblayer) provides helper functions (*dblayer.helpers.utn*) that will be used further down in this notebook.
#
# Start by adding a new network object to the 3DCityDB called *gas_network*. The returned values are the IDs of the associated (but still empty) *Network* and *NetworkGraph* objects, which represent the topographical and topological aspects of the network, respectively.
# +
from dblayer.helpers.utn.gas_network import *
( ntw_id, ntw_graph_id ) = write_network_to_db(
db_access,
name = 'gas_network',
id = 3000
)
# -
# We will start building up the network contents in the 3DCityDB by adding the nodes using function *write_network_node_to_db*.
# For every new node added to the network, this functions returns an instance of class *GasNetworkNodeData*, which holds the most relevant information of the associated database object (see [here](https://github.com/IntegrCiTy/dblayer/blob/master/dblayer/helpers/utn/gas_network.py) for details).
# For further processing, we also store this information in a separate dict:
# +
# Dict for storing information returned from database.
nodes_db_data = {}
for name, data in nodes.items():
nodes_db_data[name] = write_network_node_to_db(
db_access,
name,
data.level,
Point2D( data.x, data.y ),
srid,
ntw_id,
ntw_graph_id )
# -
# Next, we add the pipes to the database with the help of function *write_round_pipe_to_db*.
# For this, we can already make good use of the information returned in the previous step:
for name, data in pipes.items():
pipe_id = write_round_pipe_to_db(
db_access,
name,
# Provide information about database object associated with the starting node:
nodes_db_data[data.from_node],
# Provide information about database object associated with the end node:
nodes_db_data[data.to_node],
srid,
ntw_id,
ntw_graph_id,
int_diameter = data.diameter_m,
int_diameter_unit = 'm',
)
# We also want to link the network sinks to the buildings already stored in the database.
# For this, we retrieve all buildings data:
# +
buildings_db_data = db_access.get_citydb_objects(
'Building',
table_name='building',
schema='citydb_view'
)
buildings_db_id = { b.name: b.id for b in buildings_db_data}
# -
# Network sinks can be represented by objects of type *TerminalElement*, which are associated both to a network node and another CityGML object (a building in this case).
# To make this association, we can use function *write_gas_sink_to_db*:
for name, data in sinks.items():
write_gas_sink_to_db(
db_access,
name,
# Provide information about database object associated with the network node:
nodes_db_data[name],
# Provide default value for consumption (static) of this terminal element:
10.,
'kW',
srid,
ntw_id,
ntw_graph_id,
# Provide ID of city object that should be linked:
buildings_db_id[data.building_name]
)
# Similarly, we can represent sources (e.g., substations) by objects of type *TerminalElement*.
# To make this association, we can use function *write_feeder_to_db*:
for name, data in sources.items():
write_feeder_to_db(
db_access,
name,
# Provide information about database object associated with the network node:
nodes_db_data[name],
# Provide default value for generation (rating, static) of this terminal element:
data.p_lim_kw,
data.p_pa,
srid,
ntw_id,
ntw_graph_id
)
# ## Storing the data to the 3DCityDB
#
# Above, the data was *added* to the database session. In order to make it persistent, i.e., to store it permanently in the database, it has to be *committed* to the 3DCityDB.
# This is done via *commit_citydb_session*:
db_access.commit_citydb_session()
# Finally, delete the instance of class DBAccess to close the session.
del db_access
# Next up is notebook [3a_sim_setup.ipynb](../3_simulate/3a_sim_setup.ipynb), which demonstrates how to use the IntegrCiTy DAL to create a simulation setup for analyzing this gas network.
| 2_citydb/2b_gas_network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Robot Sensors
#
# A robot senses the world through cameras and other sensors, but these sensors are not perfectly accurate. In the video, you saw an example of a robot in a 1D world made of colored grid cells; all cells were either green or red. The robot then sensed that it was in a red grid cell.
#
# The probability that this reading was accurate, which we'll call the prbability that the sensor has hit its target, `pHit`, was `0.6` and the probability that this reading was inaccurate (the sensor has missed its target) and the robot was *actually* in a green cell was `pMiss` equal to `0.2`.
#
# In this notebook, let's go through how this works step by step.
# ### Uniform Distribution
#
# The robot starts with a map with a length of 5 cells. Since the robot does not know where it is at first, the probability of being in any space is the same; a uniform distribution!
#
# importing resources
import matplotlib.pyplot as plt
import numpy as np
# ex. initialize_robot(5) = [0.2, 0.2, 0.2, 0.2, 0.2]
def initialize_robot(grid_length):
''' Takes in a grid length and returns
a uniform distribution of location probabilities'''
p = [1.0/grid_length] * grid_length
return p
# I'll also include a helper function for visualizing this distribution. The below function, `display_map` will output a bar chart showing the probability that a robot is in each grid space. The y-axis has a range of 0 to 1 for the range of probabilities. For a uniform distribution, this will look like a flat line. You can choose the width of each bar to be <= 1 should you want to space these out.
def display_map(grid, bar_width=1):
if(len(grid) > 0):
x_labels = range(len(grid))
plt.bar(x_labels, height=grid, width=bar_width, color='b')
plt.xlabel('Grid Cell')
plt.ylabel('Probability')
plt.ylim(0, 1) # range of 0-1 for probability values
plt.title('Probability of the robot being at each cell in the grid')
plt.xticks(np.arange(min(x_labels), max(x_labels)+1, 1))
plt.show()
else:
print('Grid is empty')
# initialize a 5 cell, 1D world
p = initialize_robot(5)
print(p)
display_map(p)
# ### Probability After Sense
#
# Then the robot senses that it is in a red cell, and updates its probabilities. As per our example:
#
# * The probability that it is sensing the correct color is `pHit = 0.6`.
# * The probability that it is sensing the incorrect color (in this case: seeing red but *actually* in a green cell) is `pMiss = 0.2`
#
# <img src='images/robot_sensing.png' width=50% height=50% />
#
# #### Next, we write code that outputs a new grid, `p`, after multiplying each entry by pHit or pMiss at the appropriate places.
#
# Remember that the red cells (cell 1 and 2) are "hits" and the other green cells are "misses."
#
# Note that you may see values that are not exact due to how machines imperfectly represent floating points.
# +
# given initial variables
p = initialize_robot(5)
pHit = 0.6
pMiss = 0.2
# Creates a new grid, with modified probabilities, after sensing
# All values are calculated by a product of 1. the sensing probability for a color (pHit for red)
# and 2. the current probability of a robot being in that location p[i]; all equal to 0.2 at first.
p[0] = p[0]*pMiss
p[1] = p[1]*pHit
p[2] = p[2]*pHit
p[3] = p[3]*pMiss
p[4] = p[4]*pMiss
print(p)
display_map(p)
# -
# You should see that the red grid cells (1 and 2) have a higher probability than the green cells. One thing that may look strange is how low these probability bars are, and you may have noticed that these don't accurately represent a probability distribution because the components of this list do not add up to 1!
#
# ### QUIZ: Compute the sum of all of these probabilities.
#
# What do these values add up to and how do you think we can turn this into a probability distribution whose components do add up to 1?
#
# In the next code cell, write code to sum up the values in the new world, `p`.
# +
# What is the sum of all the values in p?
print("current p sum", sum(p))
## TODO: add up all the values in the list of location probabilities to determine the answer
current_total = sum(p)
p = [i/current_total for i in p]
print("normalized p list", p)
print("new p sum", sum(p))
display_map(p)
| 3-object-tracking-and-localization/activities/2-robot-location/2. Probability After Sense.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas as pd
df=pd.read_excel('Price.xlsx')
df.head(4)
df
plt.plot(df.area,df.price, 'r', alpha=0.8)
plt.xlabel("Area in sq. feet--->")
plt.ylabel('Price in US $--->')
| plotting.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="text-align: center;">
# <h2>INFSCI 2915 Foundations- Machine Learning - Spring 2018 </h2>
# <h1 style="font-size: 250%;">Assignment #3</h1>
# <h3>Issued Monday, 3/26/2018; Due Monday, 11:59pm, 4/09/2018</h3>
# <h3>Total points: 100 </h3>
# </div>
# Type in your information in the double quotes
firstName = ""
lastName = ""
pittID = ""
# <h3> Problem #1.Linear Discriminant Analysis (LDA)and Quadratic Discriminant Analysis(QDA) <br>[30 points]</h3>
# Writing a code is not required for this problem.
# <h4> Problem #1-1</h4> <br>
# Assume we have K classes to be classified with one feature $(x)$. The prior probability of
# class $k$ is $𝜋_{k} = 𝑃(𝑌 = 𝑘)$. Assume that the feature in class k has Gaussian distribution of
# mean $μ_{k}$ and variance $σ^2 (𝒩(μ,𝜎^{2}))$.The variance is the same for all classes.
# Prove that the Bayes’ classifier (that chooses class k with largest $𝑃(𝑌 = 𝑘|𝑥))$ is equivalent to assigning an observation to the class for which the discriminant function $𝛿_{k}(x)$ is
# maximized, where
# \begin{array} \\
# 𝛿_{k}(x) = x\frac{\mu _{k}}{\sigma ^{2}}- \frac{\mu_{2}^{k}}{2\sigma ^{2}}+ log(\pi _{k})
# \end{array}
# <br> What is the name of this classifier?
#
# +
#Write your narrative answer here (no code is needed for this part)
# -
# <h4> Problem #1-2</h4> <br>
# Extend **Problem #1-1** to include **p** features. With features from each class drawn from a
# Gaussian distribution with mean vector $μ_{k}$ and covariance matrix $Σ_{k}$ (which is now
# different for each class). Find the discriminant function that maximizes **𝑃(𝑌 = 𝑘|𝑥)**. Is
# the relationship with the feature vector **x** linear?<br> What is this classifier?
#
# +
#Write your narrative answer here (no code is needed for this part)
# -
# <h4> Problem #1-3</h4> <br>
# - Explain Bias-variance trade-off in choosing between LDA and QDA
# +
#Write your narrative answer here (no code is needed for this part)
# -
# <h3> Problem #2. Support Vector Machines [20 points]</h3>
# Writing a code is not required for this problem.
# <h4> Problem #2-1 Answer the following quesionts </h4>
#
# - Describe limitations of using Maximal Margin Classifier
# - What is the difference between Support Vector Classifier and Maximal Margin Classifier
# - Explain Bias-variance trade-off that occurs when we choose large and small values for the Slack Variables
#
# +
#Write your narrative answer here (no code is needed for this part)
# -
# <h4> Problem #2-2</h4><br>
# How does the Radial Basis Function Kernel in SVM measure the similarity between a test point and a training example? Discuss the impact of choosing a large RBF parameter **𝛾**
# on the learning algorithm.
# +
#Write your narrative answer here (no code is needed for this part)
# -
# <h3> Problem #3 Classification Performance Evaluation and Cross validation [30 points]</h3>
#
# <h4> Problem #3-1</h4> <br>
# In a fraud detection system, a QDA classifier’s confusion matrix is found to be:
# | |Predicted Class - Not fraud| Predicted Class - fraud|
# |:--:|:-------------------------------:|
# |Actual class – Not fraud|1000|20|
# |Actual class – fraud|30|5|
# - Evaluate the overall error rate and the accuracy<br>
# - Evaluate the precision and the recall <br>
# - Is dataset balanced?
# - Comment on the results.
# +
#Write your narrative answer here (no code is needed for this part)
# -
# <h4> Problem #3-2</h4> <br>
# Assuming we want to lower the misclassification of fraud. How can you modify the classifier to do better classification using information from confusion matrix?
# +
#Write your narrative answer here (no code is needed for this part)
# -
# <h4> Problem #3-3 Cross validation, SVM </h4> <br>
# In this exercise, we will use SVM for breast cancer classification (malignant or benign).<br>
# Use a code below to download the dataset:
#
from sklearn.datasets import load_breast_cancer
dataset = load_breast_cancer()
print(dataset.keys())
# Follow steps to answer questions.
# - Scale the features with MinMaxScaler
# - Split breast cancer dataset into two, test dataset and train dataset
# - Find best SVM classifier model. Try values of C=[0.001, 0.1, 1, 10, 1000] and Gamma = [0.001, 0.1, 1, 10, 1000].
# - Use 3-fold cross validation to find the best parameters (using all possible combinations of these values for C and gamma).
# - Report your parameters
# - Report accuracy, confusion matrix, precision, and recall (use Test dataset, SVM classifier model with best parameters)
# - Comment your results
# +
#Write your answer here
# -
# <h4> Problem #3-4 Cross validation, Logistic regression </h4>
#
# - Repeat **Problem #3-3** but instead of SVM find best Logistic regression classifier model. Try values of C= [0.001,0.01,1,10,100, 1000] and Penalty = ["l1","l2"]
#
# +
#Write your answer here
# -
# <h3> Problem #4 Neural Networks [20 points]</h3>
# In this exercise we will clasify Iris species (Setosa, Versicolor, Virginica) using Neural Networks.<br>
# Use a code below to download the dataset:
from sklearn.datasets import load_iris
dataset=load_iris()
print(dataset.keys())
# Follow steps to answer questions.
# - Scale the feautures with MinMaxScaler, then use sklearn MLPClassifier.
# - Build a model that has two hidden layers, the first layer has 10 neurons and second layer has 5 neurons.
# - Use 'relu' activation function, and set the regularization parameter alpha=0.5.
# - Set the random_state=0 for splitting and building all models.
#
# <h4> Problem #4-1 </h4>
# - Use gradient descent to solve the optimization problem (i.e. get the weights), and choose random_state=0 (which corresponds to a particular initializationo of weight values).
# - Report accuracy, confusion matrix, precision, and recall
# +
#Write your answer here
# -
# <h4> Problem #4-2 </h4>
# - Repeat **Problem #4-1** but with a model that use random_state=10 to initialize the weights.
# - Report accuracy, confusion matrix, precision, and recall
# +
#Write your answer here
# -
# <h4> Problem #4-3 </h4>
# - Repeat **Problem #4-2** but with model that use L-BFGS (a numerical quasi-Newton method) instead of stochastic gradient descent to find the weights.
# - Report accuracy, confusion matrix, precision, and recall
# - Comment on results
# +
#Write your answer here
| assignment3/Assignment 3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3.6 - AzureML
# language: python
# name: python3-azureml
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# # online nets
#
# Deep learning is powerful but computationally expensive, frequently requiring massive compute budgets. In persuit of cost-effective-yet-powerful AI, this work explores and evaluates a heuristic which should lend to more-efficient use of data through online learning.
#
# Goal: evaluate a deep learning alternative capable of true online learning. Solution requirements:
#
# 1. catastrophic forgetting should be impossible;
# 2. all data is integrated into sufficient statistics of fixed dimension;
# 3. and our solution should have predictive power comparable to deep learning.
#
# ## modeling strategy
#
# We will not attempt to derive sufficient statistics for an entire deep net, but instead leverage well-known sufficient statistics for least squares models,
# so will have sufficient statistics per deep net layer. If this can be empirically shown effective, we'll build-out the theory afterwards.
#
# Recognizing a deep net as a series of compositions, as follows.
#
# $ Y + \varepsilon \approx \mathbb{E}Y = \sigma_3 \circ \beta_3^T \circ \sigma_2 \circ \beta_2^T \circ \sigma_1 \circ \beta_1^T X $
#
# So, we can isolate invidivdual $\beta_j$ matrices using (psuedo-)inverses $\beta_j^{-1}$ like so.
#
# $ \sigma_2^{-1} \circ \beta_3^{-1} \circ \sigma_3^{-1} (Y) \approx \beta_2^T \circ \sigma_1 \circ \beta_1^T X $
#
# In this example, if we freeze all $\beta_j$'s except $\beta_2$, we are free to update $\hat \beta_2$ using $\tilde Y = \sigma_2^{-1} \circ \beta_3^{-1} \circ \sigma_3^{-1} (Y) $
# and $\tilde X = \sigma_1 \circ \beta_1^T X $.
#
# Using a least squares formulation for fitting to $\left( \tilde X, \tilde Y \right)$, we get sufficient statistics per layer.
# + [markdown] nteract={"transient": {"deleting": false}}
# # model code definitions
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1640111005277}
import torch
TORCH_TENSOR_TYPE = type(torch.tensor(1))
## NOT USED
def padded_diagonal(diag_value, n_rows, n_cols):
## construct diagonal matrix
n_diag = min(n_rows, n_cols)
diag = torch.diag(torch.tensor([diag_value]*n_diag))
if n_rows > n_cols:
## pad rows
pad = n_rows - n_cols
return torch.cat([diag, torch.zeros((pad, n_cols))], 0)
if n_cols > n_rows:
## pad cols
pad = n_cols - n_rows
return torch.cat([diag, torch.zeros((n_rows, pad))], 1)
## no padding
return diag
def padded_row(value, n_rows, n_cols):
row = torch.tensor([value]*n_cols).reshape((1, -1))
return torch.cat([row, torch.zeros((n_rows-1, n_cols))], 0)
def iterated_diagonals(diag_value, n_rows, n_cols):
## construct diagonal matrix
n_diag = min(n_rows, n_cols)
diag = torch.diag(torch.tensor([diag_value]*n_diag))
if n_rows > n_cols:
## pad rows
pad = n_rows//n_cols + 1
return torch.cat([diag]*pad, 0)[:n_rows, :n_cols]
if n_cols > n_rows:
## pad cols
pad = n_cols//n_rows + 1
return torch.cat([diag]*pad, 1)[:n_rows, :n_cols]
## no padding
return diag
class OnlineDenseLayer:
'''
A single dense net, formulated as a least squares model.
'''
def __init__(self, p, q, activation=lambda x:x, activation_inverse=lambda x:x, lam=1., clip=10000., dlamdn=0.):
'''
inputs:
- p: input dimension
- q: output dimension
- activation: non-linear function, from R^p to R^q. Default is identity.
- activation_inverse: inverse of the activation function. Default is identity.
- lam: regularization term
- clip: predicted value clipping limit
- dlamdn: rate of lambda growth relative to n for online regularization
'''
lam = float(lam)
clip = float(clip)
dlamdn = float(dlamdn)
self.__validate_inputs(p=p, q=q, lam=lam, clip=clip, dlamdn=dlamdn)
self.p = p
self.q = q
self.clip = clip
self.activation = activation
self.activation_inverse = activation_inverse
self.batch_norm_forward_mean = None
self.batch_norm_forward_std = None
self.batch_norm_forward_n = 0
self.batch_norm_backward_mean = None
self.batch_norm_backward_std = None
self.batch_norm_backward_n = 0
self.lam = lam
self.xTy = iterated_diagonals(lam, p+1,q) # +1 for intercept
self.yTx = iterated_diagonals(lam, q+1,p)
self.xTx_inv = torch.diag(torch.tensor([1./lam]*(p+1)))
self.yTy_inv = torch.diag(torch.tensor([1./lam]*(q+1)))
self.betaT_forward = torch.matmul(self.xTx_inv, self.xTy)
self.betaT_forward = torch.transpose(self.betaT_forward, 0, 1)
self.betaT_backward = torch.matmul(self.yTy_inv, self.yTx)
self.betaT_backward = torch.transpose(self.betaT_backward, 0, 1)
self.x_forward = None
self.y_forward = None
self.x_backward = None
self.y_backward = None
pass
def forward(self, x):
'creates and stores x_forward and y_forward, then returns activation(y_forward)'
self.__validate_inputs(x=x, p=self.p)
x = self.batch_norm(x, forward=True) ## TODO use fitting or not
self.x_forward = x
x = torch.cat((torch.tensor([[1.]]), x), dim=0) # intercept
self.y_forward = torch.matmul(self.betaT_forward, x) # predict
self.y_forward = torch.clip(self.y_forward, -self.clip, self.clip)
return self.activation(self.y_forward)
def backward(self, y):
'creates and stores x_backward and y_backward, then returns y_backward'
y = self.activation_inverse(y)
self.__validate_inputs(y=y, q=self.q)
y = self.batch_norm(y, forward=False) ## TODO use fitting or not
self.y_backward = y
y = torch.cat((torch.tensor([[1.]]), y), dim=0)
self.x_backward = torch.matmul(self.betaT_backward, y)
self.x_forward = torch.clip(self.x_forward, -self.clip, self.clip)
return self.x_backward
def forward_fit(self):
'uses x_forward and y_backward to update forward model, then returns <NAME> denominator'
self.__validate_inputs(x=self.x_forward, y=self.y_backward, p=self.p, q=self.q)
x = torch.cat((torch.tensor([[1.]]), self.x_forward), dim=0)
self.xTx_inv, sm_denom = self.sherman_morrison(self.xTx_inv, x, x)
self.xTy += torch.matmul(x, torch.transpose(self.y_backward, 0, 1))
self.betaT_forward = torch.matmul(self.xTx_inv, self.xTy)
self.betaT_forward = torch.transpose(self.betaT_forward, 0, 1)
return sm_denom
def backward_fit(self):
'uses x_backward and y_forward to update backward model, then returns Sherman Morrison denominator'
self.__validate_inputs(x=self.x_forward, y=self.y_backward, p=self.p, q=self.q)
y = torch.cat((torch.tensor([[1.]]), self.y_backward), dim=0)
self.yTy_inv, sm_denom = self.sherman_morrison(self.yTy_inv, y, y)
self.yTx += torch.matmul(y, torch.transpose(self.x_backward, 0, 1))
self.betaT_backward = torch.matmul(self.yTy_inv, self.yTx)
self.betaT_backward = torch.transpose(self.betaT_backward, 0, 1)
return sm_denom
def batch_norm(self, x, forward, fitting=True):
'''
batch normalize tensor
inputs:
- x: (tensor) to be normalized
- forward: (boolean) indicates prediction is forward, instead of backward
- fitting: (boolean) if in model fitting, update values
'''
## retrieve
if forward:
m = self.batch_norm_forward_mean
s = self.batch_norm_forward_std
n = self.batch_norm_forward_n
else:
m = self.batch_norm_backward_mean
s = self.batch_norm_backward_std
n = self.batch_norm_backward_n
## caculate
if n == 0:
n = 1
m = x.mean()
s = x.std()
else:
n += 1
m = x.mean()/n + m*(n-1)/n
s = x.std()/n + s*(n-1)/n
## store
if fitting:
if forward:
self.batch_norm_forward_mean = m
self.batch_norm_forward_std = s
self.batch_norm_forward_n = n
else:
self.batch_norm_backward_mean = m
self.batch_norm_backward_std = s
self.batch_norm_backward_n = n
## no dividing by zero
if s < 1e-3:
s = 1e-3
return (x - m)/s
@staticmethod
def sherman_morrison(inv_mat, vec1, vec2):
'''
applies Sherman Morrison updates, (mat + vec1 vec2^T)^{-1}
inputs:
- inv_mat: an inverted matrix
- vec1: a column vector
- vec2: a column vector
returns:
- updated matrix
- the Sherman Morrison denominator, for tracking numerical stability
'''
v2t = torch.transpose(vec2, 0, 1)
denominator = 1. + torch.matmul(torch.matmul(v2t, inv_mat), vec1)
numerator = torch.matmul(torch.matmul(inv_mat, vec1), torch.matmul(v2t, inv_mat))
updated_inv_mat = inv_mat - numerator / denominator
return updated_inv_mat, float(denominator)
def __validate_inputs(self, p=None, q=None, lam=None, x=None, y=None, clip=None, dlamdn=None):
'raises value exceptions if provided parameters are invalid'
if q is not None:
if not isinstance(q, int):
raise ValueError('`q` must be int!')
if q <= 0:
raise ValueError('`q` must be greater than zero!')
if p is not None:
if not isinstance(p, int):
raise ValueError('`p` must be int!')
if p <= 0:
raise ValueError('`p` must be greater than zero!')
if lam is not None:
if not (isinstance(lam, float) or isinstance(lam, int)):
raise ValueError('`lam` must be float or int!')
if lam < 0:
raise ValueError('`lam` must be non-negative!')
if x is not None and p is not None:
if type(x) != TORCH_TENSOR_TYPE:
raise ValueError('`x` must be of type `torch.tensor`!')
if list(x.shape) != [p,1]:
raise ValueError('`x.shape` must be `[p,1]`!')
if torch.isnan(x).any():
raise ValueError('`x` contains `nan`!')
pass
if y is not None and q is not None:
if type(y) != TORCH_TENSOR_TYPE:
raise ValueError('`y` must be of type `torch.tensor`!')
if list(y.shape) != [q,1]:
raise ValueError('`y.shape` must be `[q,1]`')
if torch.isnan(y).any():
raise ValueError('`y` contains `nan`!')
pass
if clip is not None:
if type(clip) != float:
raise ValueError('`clip` my be of type `float`!')
if clip <= 0.:
raise ValueError('`clip` must be positive!')
pass
if dlamdn is not None:
if type(dlamdn) != float:
raise ValueError('`dlamdn` my be of type `float`!')
if dlamdn < 0.:
raise ValueError('`dlamdn` must be non-negative!')
pass
pass
pass
class OnlineNet:
'online, sequential dense net'
def __init__(self, layer_list):
## validate inputs
if type(layer_list) != list:
raise ValueError('`layer_list` must be of type list!')
for layer in layer_list:
if not issubclass(type(layer), OnlineDenseLayer):
raise ValueError('each item in `layer_list` must be an instance of a subclass of `OnlineDenseLayer`!')
## assign
self.layer_list = layer_list
pass
def forward(self, x):
'predict forward'
for layer in self.layer_list:
x = layer.forward(x)
return x
def backward(self, y):
'predict backward'
for layer in reversed(self.layer_list):
y = layer.backward(y)
return y
def fit(self):
'assumes layers x & y targets have already been set. Returns Sherman Morrison denominators per layer in (forward, backward) pairs in a list'
sherman_morrison_denominator_list = []
for layer in self.layer_list:
forward_smd = layer.forward_fit()
backward_smd = layer.backward_fit()
sherman_morrison_denominator_list.append((forward_smd, backward_smd))
return sherman_morrison_denominator_list
def __reduce_sherman_morrison_denominator_list(self, smd_pair_list):
'returns the value closest to zero'
if type(smd_pair_list) != list:
raise ValueError('`smd_pair_list` must be of type `list`!')
if len(smd_pair_list) == 0:
return None
smallest_smd = None
for smd_pair in smd_pair_list:
if type(smd_pair) != tuple:
raise ValueError('`smd_pair_list` must be list of tuples!')
if smallest_smd is None:
smallest_smd = smd_pair[0]
if abs(smallest_smd) > abs(smd_pair[0]):
smallest_smd = smd_pair[0]
if abs(smallest_smd) > abs(smd_pair[1]):
smallest_smd = smd_pair[1]
return float(smallest_smd)
def __call__(self, x, y=None):
'''
If only x is given, a prediction is made and returned.
If x and y are given, then the model is updated, and returns
- the prediction
- the sherman morrison denominator closest to zero, for tracking numerical stability
'''
y_hat = self.forward(x)
if y is None:
return y_hat
self.backward(y)
self.layer_list[0].x_forward = x
self.layer_list[0].x_backward = x
self.layer_list[-1].y_forward = y
self.layer_list[-1].y_backward = y
smd_pair_list = self.fit()
smallest_smd = self.__reduce_sherman_morrison_denominator_list(smd_pair_list)
return y_hat, smallest_smd
## tests
## test 1: <NAME>
a = torch.tensor([[2., 1.], [1., 2.]])
b = torch.tensor([[.1],[.2]])
sm_inv, _ = OnlineDenseLayer.sherman_morrison(torch.inverse(a),b,b)
num_inv = torch.inverse(a+torch.matmul(b, torch.transpose(b,0,1)))
err = float(torch.abs(sm_inv - num_inv).sum())
assert(err < 1e-5)
# + [markdown] nteract={"transient": {"deleting": false}}
# # first experiment: mnist classification
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1640112291633}
from tqdm import tqdm
from torchvision import datasets, transforms
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.1307,), (0.3081,))
])
dataset1 = datasets.MNIST('../../data', train=True, download=True, transform=transform)
dataset2 = datasets.MNIST('../../data', train=False, transform=transform)
train_loader = torch.utils.data.DataLoader(dataset1)
test_loader = torch.utils.data.DataLoader(dataset2)
n_labels = 10
lam = 1000.
clip=100.
## activation functions
## torch.sigmoid
inv_sigmoid = lambda x: -torch.log((1/(x+1e-8))-1)
leaky_relu_alpha = .1
leaky_relu = lambda x: (x > 0)*x + (x <= 0)*x*leaky_relu_alpha
inv_leaky_relu = lambda x: (x > 0)*x + (x <= 0)*x/leaky_relu_alpha
model = OnlineNet(
[
OnlineDenseLayer(p=1*1*28*28, q=100, activation=leaky_relu, activation_inverse=inv_leaky_relu, lam=lam, clip=clip),
OnlineDenseLayer(p=100, q=100, activation=leaky_relu, activation_inverse=inv_leaky_relu, lam=lam, clip=clip),
OnlineDenseLayer(p=100, q=100, activation=leaky_relu, activation_inverse=inv_leaky_relu, lam=lam, clip=clip),
OnlineDenseLayer(p=100, q=n_labels, activation=torch.sigmoid, activation_inverse=inv_sigmoid, lam=lam, clip=clip)
]
)
def build_data(image, label):
'format data from iterator for model'
y = torch.tensor([1. if int(label[0]) == idx else 0. for idx in range(n_labels)]) ## one-hot representation
x = image.reshape([-1]) ## flatten
## shrink so sigmoid inverse is well-defined
y = y*.90 + .05
## reshape to column vectors
x = x.reshape([-1,1])
y = y.reshape([-1,1])
return x, y
def match(y, y_hat):
y = y.reshape(-1)
y_hat = y_hat.reshape(-1)
if y.argmax() == y_hat.argmax():
return 1.
return 0.
errs = []
stab = []
y_std = []
acc = [0.]
def fit():
pbar = tqdm(train_loader)
for [image, label] in pbar:
x, y = build_data(image, label)
## fit
y_hat, stability = model(x, y)
## stats
err = float((y - y_hat).abs().sum())
errs.append(err)
stab.append(stability)
std = float(y_hat.std())
y_std.append(std)
acc_n = max(len(acc), 1000)
acc.append( match(y,y_hat)/acc_n + acc[-1]*(acc_n-1)/acc_n)
pbar.set_description(f'acc: {acc[-1]}, err: {err}, y_std: {std}, stab: {stability}')
## train error
## TODO
pass
fit()
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1640112588981}
import matplotlib.pyplot as plt
print('acc')
plt.plot(acc[100:])
plt.show()
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1640112592148}
print('errs')
plt.plot(errs)
plt.show()
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1640112594525}
print('y_std')
plt.plot(y_std)
plt.show()
# + jupyter={"source_hidden": false, "outputs_hidden": false} nteract={"transient": {"deleting": false}} gather={"logged": 1640112596460}
print('stab')
plt.plot(stab)
plt.show()
| online-nets/.ipynb_aml_checkpoints/online-nets-checkpoint2021-11-21-19-19-14Z.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Chapter6.2 学習とテスト
# +
# testdatasetというデータセットを作成、既にあれば新たに作る必要はない
# # !bq mk testdataset
# wdbc テープルを作成
# # !bq load --autodetect testdataset.wdbc ../datasets/wdbc.csv
# -
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# pandas DataFrameに読み込みます
# <font color="red">**PROJECTID**を書き換えて実行してください</font>
# +
# BigQueryクエリ結果をDataFrameに読み込む
query = 'SELECT * FROM testdataset.wdbc ORDER BY index'
dataset = pd.read_gbq(project_id='PROJECTID', query=query)
# データの先頭5行を表示
dataset.head()
# -
# データの整形
# +
# 'M'を0, 'B'を1に変換
dataset['diagnostic'] = dataset['diagnostic'].apply(
lambda x: 0 if x == 'M' else 1)
# 'index'カラムを削除
dataset.drop('index', axis=1, inplace=True)
# -
# DataFrameからarrayに変換
X_dataset = dataset.drop('diagnostic', axis=1).as_matrix()
y_dataset = dataset.diagnostic.as_matrix()
# 学習とテストデータの分割
# +
from sklearn.model_selection import train_test_split
# 学習用とテスト用にデータセットを分ける
X_train, X_test, y_train, y_test = train_test_split(
X_dataset, y_dataset, test_size=0.2, random_state=42)
# -
# ランダムフォレストで識別
# +
from sklearn.ensemble import RandomForestClassifier
# 識別器のインスタンスを生成
classifier = RandomForestClassifier(random_state=42)
# 学習用データを学習
classifier.fit(X_train, y_train)
# テスト用データで推論し、正解率を算出
classifier.score(X_test, y_test)
# -
# SVMで識別
# +
from sklearn.svm import SVC
# 識別器のインスタンスを生成
classifier = SVC()
# 学習用データを学習
classifier.fit(X_train, y_train)
# テスト用データで推論し、正解率を算出
classifier.score(X_test, y_test)
# -
# データのレンジを合わせる
# +
from sklearn.preprocessing import MinMaxScaler
# 最大値が1、最小値が0になるよう各特徴量をスケールする
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# 再度、学習と推論を行う
classifier.fit(X_train, y_train)
classifier.score(X_test, y_test)
# -
# K-Fold クロスバリデーション
# +
from sklearn.model_selection import KFold
# データを4分割するモード
kf = KFold(n_splits=4, random_state=42, shuffle=False)
test_data = np.zeros(32)
# 学習用とテスト用に分割された配列のインデックスを取得
for train_index, test_index in kf.split(test_data):
# test_indexのインデックスだけ値を1にし、他は0の配列を作る
dat = np.zeros(32)
dat[test_index] = 1
# データを可視化
plt.gray()
plt.matshow(dat.reshape(1, 32), extent=[0, 32, 0, 1])
plt.gca().set_yticks([])
plt.gca().set_xticks(range(32), minor='true')
plt.grid(which='minor')
plt.show()
# -
# KFoldクロスバリデーションでSVMを学習
# +
# 識別器のインスタンスを作成
classifier = SVC()
kf = KFold(n_splits=4, random_state=42, shuffle=True)
for train_index, test_index in kf.split(X_dataset):
# データセットを学習とテストに分ける
X_train, X_test = X_dataset[train_index], X_dataset[test_index]
y_train, y_test = y_dataset[train_index], y_dataset[test_index]
# スケール
scaler = MinMaxScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# 学習と正解率の算出
classifier.fit(X_train, y_train)
print(classifier.score(X_test, y_test))
| Part2/Chapter6.2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="of3HGFCW2ii7"
# <a id='Q0'></a>
# <center><a target="_blank" href="http://www.propulsion.academy"><img src="https://drive.google.com/uc?id=1McNxpNrSwfqu1w-QtlOmPSmfULvkkMQV" width="200" style="background:none; border:none; box-shadow:none;" /></a> </center>
# <center> <h4 style="color:#303030"> Python for Data Science, Homework, template: </h4> </center>
# <center> <h1 style="color:#303030">Breast Cancer Selection</h1> </center>
# <p style="margin-bottom:1cm;"></p>
# <center style="color:#303030"><h4>Propulsion Academy, 2021</h4></center>
# <p style="margin-bottom:1cm;"></p>
#
# <div style="background:#EEEDF5;border-top:0.1cm solid #EF475B;border-bottom:0.1cm solid #EF475B;">
# <div style="margin-left: 0.5cm;margin-top: 0.5cm;margin-bottom: 0.5cm">
# <p><strong>Goal:</strong> Binary classification on Breast Cancer data</p>
# <strong> Sections:</strong>
# <a id="P0" name="P0"></a>
# <ol>
# <li> <a style="color:#303030" href="#SU">Set Up </a> </li>
# <li> <a style="color:#303030" href="#P1">Modeling</a></li>
# <li> <a style="color:#303030" href="#P2">Evaluation</a></li>
# </ol>
# <strong>Topics Trained:</strong> Binary Classification.
# </div>
# </div>
#
#
# <nav style="text-align:right"><strong>
# <a style="color:#00BAE5" href="https://monolith.propulsion-home.ch/backend/api/momentum/materials/intro-2-ds-materials/" title="momentum"> SIT Introduction to Data Science</a>|
# <a style="color:#00BAE5" href="https://monolith.propulsion-home.ch/backend/api/momentum/materials/intro-2-ds-materials/weeks/week2/day1/index.html" title="momentum">Week 2 Day 1, Applied Machine Learning</a>|
# <a style="color:#00BAE5" href="https://colab.research.google.com/drive/1DK68oHRR2-5IiZ2SG7OTS2cCFSe-RpeE?usp=sharing" title="momentum"> Assignment, Classification of the success of pirate attacks</a>
# </strong></nav>
# + [markdown] id="ckLGGhLpmYD8"
# <a id='SU' name="SU"></a>
# ## [Set up](#P0)
# + id="sdNiUvJ9sTLA" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619033928493, "user_tz": -330, "elapsed": 183775, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14834829200815078196"}} outputId="4e3e406c-4bd7-4685-af0d-5dcabf3aad6f"
# !sudo apt-get install build-essential swig
# !curl https://raw.githubusercontent.com/automl/auto-sklearn/master/requirements.txt | xargs -n 1 -L 1 pip install
# !pip install auto-sklearn
# !pip install pipelineprofiler # visualize the pipelines created by auto-sklearn
# !pip install shap
# !pip install --upgrade plotly
# !pip3 install -U scikit-learn
# + [markdown] id="RspLUVmbsTLB"
# ### Package imports
# + id="3c7D7zi94i5b" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1619036963049, "user_tz": -330, "elapsed": 11783, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14834829200815078196"}} outputId="ae3d3f08-0af4-4262-d186-35a3d24a2690"
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.preprocessing import StandardScaler
from sklearn.linear_model import LinearRegression
from sklearn import metrics
from sklearn import set_config
from sklearn.pipeline import Pipeline
from pandas_profiling import ProfileReport
from sklearn.impute import SimpleImputer
from sklearn.metrics import mean_squared_error, accuracy_score, f1_score
from sklearn.base import BaseEstimator, TransformerMixin
import autosklearn.classification
import PipelineProfiler
import plotly.express as px
import plotly.graph_objects as go
from joblib import dump
import shap
import datetime
import logging
import matplotlib.pyplot as plt
# + id="mbtwn7358_Xc"
from sklearn.model_selection import train_test_split, cross_val_score
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.preprocessing import StandardScaler, OrdinalEncoder, OneHotEncoder
from sklearn import preprocessing
from sklearn.metrics import silhouette_score
from sklearn import set_config
from sklearn.pipeline import Pipeline
from sklearn.compose import ColumnTransformer
from sklearn.linear_model import LogisticRegression
from sklearn.impute import SimpleImputer
# + [markdown] id="7BGAPlne9D_N"
# Connect to Google Drive
# + colab={"base_uri": "https://localhost:8080/"} id="LUmqb8oH4m3P" executionInfo={"status": "ok", "timestamp": 1618733289353, "user_tz": -330, "elapsed": 1933, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14834829200815078196"}} outputId="80746f8e-e25b-4844-a24f-881af1c4e889"
from google.colab import drive
drive.mount('/content/drive', force_remount=True)
# + id="utIL2E025xId"
data_path="/content/drive/MyDrive/Introduction2DataScience/W2D2_Assignment/w2d2/data/raw/"
model_path = "/content/drive/MyDrive/Introduction2DataScience/W2D2_Assignment/w2d2/models/"
timesstr = str(datetime.datetime.now()).replace(' ', '_')
logging.basicConfig(filename=f"{model_path}explog_{timesstr}.log", level=logging.INFO)
# + id="tkxjKyn15opR"
pd.set_option('display.max_rows', 25)
# + id="8VJBC3wf6Azw"
set_config(display='diagram')
# + [markdown] id="tiBsfiFvstdj"
# Please Download the data from [this source](https://drive.google.com/file/d/1uMM8qdQSiHHjIiYPd45EPzXH7sqIiQ9t/view?usp=sharing), and upload it on your introduction2DS/data google drive folder.
# + [markdown] id="g6UK6Rj4ozP4"
# <a id='P1' name="P1"></a>
# ## [Loading Data and Train-Test Split](#P0)
#
# + [markdown] id="G92XL9EOqqLG"
# **Load the csv file as a DataFrame using Pandas**
# + id="vhLQe4H-qqLH"
# your code here
df = pd.read_csv(f'{data_path}data-breast-cancer.csv')
df['diagnosis']=df['diagnosis'].map({'M':1, 'B':0})
# + id="ZUXoCNfwdpG9"
df = df.drop(["Unnamed: 32", "id"], axis=1)
# + id="_y1VzaWvB5mE"
test_size = 0.2
random_state = 42
# + id="kRY7IjIl51tu"
Y = df['diagnosis']
X = df.drop('diagnosis', axis=1)
x_train, x_test, y_train, y_test = train_test_split(X, Y, test_size=test_size, random_state=random_state, stratify=Y)
# + id="LEsPw6aAesMB"
logging.info(f'train test split with test_size={test_size} and random state={random_state}')
# + id="Y3QSCMrTewGV"
total_time = 60
per_run_time_limit = 30
# + colab={"base_uri": "https://localhost:8080/", "height": 88} id="-wZsvtfIezHT" executionInfo={"status": "ok", "timestamp": 1618733390855, "user_tz": -330, "elapsed": 57602, "user": {"displayName": "akshat jhalani", "photoUrl": "", "userId": "14834829200815078196"}} outputId="9db39fc2-6e8d-464a-a612-c1f7b0af5dc2"
automl = autosklearn.classification.AutoSklearnClassifier(
time_left_for_this_task=total_time,
per_run_time_limit=per_run_time_limit,
)
automl.fit(x_train, y_train)
# + id="EZoZEXEkWHup"
# profiler_data= PipelineProfiler.import_autosklearn(automl)
# PipelineProfiler.plot_pipeline_matrix(profiler_data)
# + [markdown] id="-zX0WtORr_41"
# _Your Comments here_
# + id="06byp_jjfixk"
logging.info(f'Ran autosklearn regressor for a total time of {total_time} seconds, with a maximum of {per_run_time_limit} seconds per model run')
# + colab={"base_uri": "https://localhost:8080/"} id="K3kkwMrifjVX" executionInfo={"status": "ok", "timestamp": 1618733409617, "user_tz": -330, "elapsed": 8080, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14834829200815078196"}} outputId="bfdf20ed-6450-4b02-b4f3-2c9044751f5f"
dump(automl, f'{model_path}model{timesstr}.pkl')
# + id="47v4-o50flb2"
logging.info(f'Saved regressor model at {model_path}model{timesstr}.pkl ')
# + id="dpBy9Cksgc9Z"
logging.info(f'autosklearn model statistics:')
logging.info(automl.sprint_statistics())
# + [markdown] id="c91uTIghgmix"
# <a id='P2' name="P2"></a>
# ## [Model Evaluation and Explainability](#P0)
# + id="MtXkGJIP5el0"
y_pred = automl.predict(x_test)
# + id="YH60TOMQfe0o"
logging.info(f"Accuracy is {accuracy_score(y_test, y_pred)}, \n F1 score is {f1_score(y_test, y_pred)}")
# + [markdown] id="BbtABjHRhdoN"
# #### Model Explainability
# + id="9EmfLmYrKSm0"
explainer = shap.KernelExplainer(model = automl.predict, data = x_test.iloc[:50, :], link = "identity")
# + colab={"base_uri": "https://localhost:8080/", "height": 92, "referenced_widgets": ["a1a60015c93b4de78b6e43e38d87ec39", "b2e65808f33b491a9ebb86eb41c33afa", "0db667967a6b451e8649284631803285", "9fdaa9d3b9a2444393968d7758f12f73", "16c5da3bd7d34d19948d2ed00403ab84", "fcc657f2f7d0468aacfe44181ecaf6c5", "a26010f7a62c4f62aa5afea20b0bd623", "a46349db9b434db183c4cb9e6c25fd33"]} id="qLLcd_mRhiEv" executionInfo={"status": "ok", "timestamp": 1618733427577, "user_tz": -330, "elapsed": 5839, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14834829200815078196"}} outputId="a0b58a86-37ca-4b57-d28f-d4ceff72fe6c"
# Set the index of the specific example to explain
x_idx = 0
shap_value_single = explainer.shap_values(X = x_test.iloc[x_idx:x_idx+1,:], nsamples = 100)
x_test.iloc[x_idx:x_idx+1,:]
# print the JS visualization code to the notebook
shap.initjs()
shap.force_plot(base_value = explainer.expected_value,
shap_values = shap_value_single,
features = x_test.iloc[x_idx:x_idx+1,:],
show=False,
matplotlib=True
)
plt.savefig(f"{model_path}shap_example_{timesstr}.png")
logging.info(f"Shapley example saved as {model_path}shap_example_{timesstr}.png")
# + colab={"base_uri": "https://localhost:8080/", "height": 66, "referenced_widgets": ["07d4d7b0e2a84ec7a85914b695ca94c3", "<KEY>", "b6fb133cca0748bca6ce539fc4b850bd", "523b739bb9de435da490a04db58815a3", "0ab1ef1882d6449d8f66b2e39005df74", "6ce8818aa8fa48fb8d481999fb24e15f", "af70da7deacc435281f59df94a718cc2", "ea54ed61392e4c809cd323d65d5710b6"]} id="NJnC-fjkh3ZS" executionInfo={"status": "ok", "timestamp": 1618733579924, "user_tz": -330, "elapsed": 154567, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14834829200815078196"}} outputId="a1e20c05-601d-4ada-cf27-99d364822469"
shap_values = explainer.shap_values(X = x_test.iloc[0:50,:], nsamples = 100)
# + colab={"base_uri": "https://localhost:8080/", "height": 43} id="2-qFguLnh32h" executionInfo={"status": "ok", "timestamp": 1618733580644, "user_tz": -330, "elapsed": 154494, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "14834829200815078196"}} outputId="deb5d013-964a-4b66-8d00-a6f3189b5c47"
# print the JS visualization code to the notebook
shap.initjs()
fig = shap.summary_plot(shap_values = shap_values,
features = x_test.iloc[0:50,:],
show=False)
plt.savefig(f"{model_path}shap_summary_{timesstr}.png")
logging.info(f"Shapley summary saved as {model_path}shap_summary_{timesstr}.png")
# + [markdown] id="I6tLyrF3MCho"
# --------------
# # End of This Notebook
| notebooks/SIT_W2D1_Breast_Cancer_Model_Deployment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''hra'': conda)'
# name: python3
# ---
# # Random Forest Classifier
# Load the packages
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import AdaBoostClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import classification_report
# Load the data
train_df = pd.read_csv('./../../../../data/train/train.csv')
test_df = pd.read_csv('./../../../../data/test/test.csv')
# Load the feature selection result
feature_selector = pd.read_csv('./../../../../data/feature_ranking.csv')
feature_selector.set_index('Unnamed: 0', inplace=True)
# Separate feature space from target variable
y_train = train_df['Attrition']
X_train = train_df.drop('Attrition', axis=1)
y_test = test_df['Attrition']
X_test = test_df.drop('Attrition', axis=1)
# We will be running models for different set of features and evaluate their performances. We start with complete dataset and then start with meaximum feature score of 8 to 5.
# Declare the model paramters for searching
param_grid = dict(
n_estimators = [25, 50, 100, 200],
learning_rate = [1, 0.1, 0.01, 0.001]
)
# Declare and train the model
ada_clf = AdaBoostClassifier(DecisionTreeClassifier(class_weight="balanced", max_features=None, criterion='entropy', max_depth=40))
ada = GridSearchCV(estimator=ada_clf, param_grid=param_grid, scoring='f1', n_jobs=-1)
# ## Complete data
# Train the model
dt.fit(X_train, y_train)
# Get the parameters for the best model
dt.best_estimator_
# Predict using model
y_pred = dt.predict(X_test)
# Make the classification report
print(classification_report(y_test, y_pred))
# The results not better than that of logistic regression. The precision, recall and f1 of attrition is not at all good as that of random forest.
# Train the model
ada.fit(X_train, y_train)
# Get the parameters for the best model
ada.best_estimator_
# Predict using model
y_pred = ada.predict(X_test)
# Make the classification report
print(classification_report(y_test, y_pred))
# The adaboost classifier is performing as good as the decision tree classifier only. The results for attrition does not seem to be good.
# ## Feature score of 8
# +
# Create the new dataset
# Get features with feature score of 8
features = feature_selector[feature_selector['Total']==8].index.tolist()
X_train_8 = X_train.loc[:, features]
X_test_8 = X_test.loc[:, features]
# -
# Train the model
ada.fit(X_train_8, y_train)
# Predict with model
y_pred_8 = ada.predict(X_test_8)
# Make the report
print(classification_report(y_test, y_pred))
# There is no improvement in the result. But since this model uses less number of features, it better to use it in production in order to improve the retraining and inferencing with huge load of data.
# Since the least number of features that could be used gave the same performance as all the features, it is better to skip the other scores since the chance of improvement in result is quite less.
| hrbook/files/models/adaboost.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [[source]](../api/alibi.explainers.shap_wrappers.rst)
# # Kernel SHAP
# ## Overview
# The Kernel SHAP (**SH**apley **A**dditive ex**P**lanations) algorithm is based on the paper [A Unified Approach to Interpreting Model Predictions](https://papers.nips.cc/paper/7062-a-unified-approach-to-interpreting-model-predictions) by Lundberg et al. and builds on the open source [shap library](https://github.com/slundberg/shap) from the paper's first author.
#
# The algorithm provides model-agnostic (*black box*), human interpretable explanations suitable for regression and classification models applied to tabular data. This method is a member of the *additive feature attribution methods* class; feature attribution refers to the fact that the change of an outcome to be explained (e.g., a class probability in a classification problem) with respect to a *baseline* (e.g., average prediction probability for that class in the training set) can be attributed in different proportions to the model input features.
#
# A simple illustration of the explanation process is shown in Figure 1. Here we see depicted a model which takes as an input features such as `Age`, `BMI` or `Sex` and outputs a continuous value. We know that the average value of that output in a dataset of interest is `0.1`. Using the Kernel SHAP algorithm, we attribute the `0.3` difference to the input features. Because the sum of the attribute values equals `output - base rate`, this method is _additive_. We can see for example that the `Sex` feature contributes negatively to this prediction whereas the remainder of the features have a positive contribution. For explaining this particular data point, the `Age` feature seems to be the most important. See our examples on how to perform explanations with this algorithm and visualise the results using the `shap` library visualisations [here](../examples/kernel_shap_wine_intro.ipynb), [here](../examples/kernel_shap_wine_lr.ipynb) and [here](../examples/kernel_shap_adult_lr.ipynb).
# 
# Figure 1: Cartoon illustration of black-box explanation models with Kernel SHAP
#
# Image Credit: <NAME> (see source [here](https://github.com/slundberg/shap))
# ## Usage
# In order to compute the shap values , the following hyperparameters can be set when calling the `explain` method:
#
# * `nsamples`: Determines the number of subsets used for the estimation of the shap values. A default of `2*M + 2**11` is provided where `M` is the number of features. One is encouraged to experiment with the number of samples in order to determine a value that balances explanation accuracy and runtime.
#
# * `l1_reg`: can take values `0`, `False` to disable, `auto` for automatic regularisation selection, `bic` or `aic` to use $\ell_1$ regularised regression with the Bayes/Akaike information criteria for regularisation parameter selection, `num_features(10)` to specify the number of feature effects to be returned or a float value that is used as the regularisation coefficient for the $\ell_1$ penalised regression. The default option `auto` uses the least angle regression algorithm with the Akaike Information Criterion if a fraction smaller than `0.2` of the total number of subsets is enumerated.
#
# If the dataset to be explained contains categorical variables, then the following options can be specified _unless_ the categorical variables have been grouped (see example below):
#
# * `summarise_result`: if True, the shap values estimated for dimensions of an encoded categorical variable are summed and a single shap value is returned for the categorical variable. This requires that both arguments below are specified:
# * `cat_var_start_idx`: a list containing the column indices where categorical variables start. For example if the feature matrix contains a categorical feature starting at index `0` and one at index `10`, then `cat_var_start_idx=[0, 10]`.
# * `cat_vars_enc_dim`: a list containing the dimension of the encoded categorical variables. The number of columns specified in this list is summed for each categorical variable starting with the corresponding index in `cat_var_start_idx`. So if `cat_var_start_idx=[0, 10]` and `cat_vars_enc_dim=[3, 5]`, then the columns with indices `0, 1` and `2` and `10, 11, 12, 13` and `14` will be combined to return one shap value for each categorical variable, as opposed to `3` and `5`.
# ### Explaining continuous datasets
# #### Initialisation and fit
# The explainer is initialised by specifying:
#
# * a predict function.
# * optionally, setting `link='logit'` if the the model to be explained is a classifier that outputs probabilities. This will apply the logit function to convert outputs to margin space.
# * optionally, providing a list of `feature_names`
#
# Hence assuming the classifier takes in 4 inputs and returns probabilities of 3 classes, we initialise its explainer as:
#
# ```python
# from alibi.explainers import KernelShap
#
# predict_fn = lambda x: clf.predict_proba(x)
# explainer = KernelShap(predict_fn, link='logit', feature_names=['a','b','c','d'])
# ```
# To fit our classifier, we simply pass our background or 'reference' dataset to the explainer:
#
# ```python
# explainer.fit(X_reference)
# ```
#
# Note that `X_reference` is expected to have a `samples x features` layout.
# #### Explanation
# To explain an instance `X`, we simply pass it to the explain method:
#
# ```python
# explanation = explainer.explain(X)
# ```
# The returned explanation object has the following fields:
#
# * `explanation.meta`:
#
# ```python
# {'name': 'KernelShap',
# 'type': ['blackbox'],
# 'explanations': ['local', 'global'],
# 'params': {'groups': None,
# 'group_names': None,
# 'weights': None,
# 'summarise_background': False
# }
# }
# ```
#
# This field contains metadata such as the explainer name and type as well as the type of explanations this method can generate. In this case, the `params` attribute shows that none of the `fit` method optional parameters have been set.
#
# * `explanation.data`:
#
# ```python
# {'shap_values': [array([ 0.8340445 , 0.12000589, -0.07984099, 0.61758141]),
# array([-0.71522546, 0.31749045, 0.3146705 , -0.13365639]),
# array([-0.12984616, -0.47194649, -0.23036243, -0.52314911])],
# 'expected_value': array([0.74456904, 1.05058744, 1.15837362]),
# 'link': 'logit',
# 'feature_names': ['a', 'b', 'c', 'd'],
# 'categorical_names': {},
# 'raw': {
# 'raw_prediction': array([ 2.23635984, 0.83386654, -0.19693058]),
# 'prediction': array([0]),
# 'instances': array([ 0.93884707, -0.63216607, -0.4350103 , -0.91969562]),
# 'importances': {
# '0': {'ranked_effect': array([0.8340445 , 0.61758141, 0.12000589, 0.07984099]),
# 'names': ['a', 'd', 'b', 'c']},
# '1': {'ranked_effect': array([0.71522546, 0.31749045, 0.3146705 , 0.13365639]),
# 'names': ['a', 'b', 'c', 'd']},
# '2': {'ranked_effect': array([0.52314911, 0.47194649, 0.23036243, 0.12984616]),
# 'names': ['d', 'b', 'c', 'a']},
# 'aggregated': {'ranked_effect': array([1.67911611, 1.27438691, 0.90944283, 0.62487392]),
# 'names': ['a', 'd', 'b', 'c']}
# }
# }
# }
# ```
#
# This field contains:
#
# * `shap_values`: a list of length equal to the number of model outputs, where each entry is an array of dimension `samples x features` of shap values. For the example above , only one instance with 4 features has been explained so the shap values for each class are of dimension `1 x 4`
# * `expected_value`: an array of the expected value for each model output across `X_reference`
# * `link`: which function has been applied to the model output prior to computing the `expected_value` and estimation of the `shap_values`
# * `feature_names`: a list with the feature names, if provided. Defaults to a list containing strings of with the format `feature_{}` if no names are passed
# * `categorical_names`: a mapping of the categorical variables (represented by indices in the `shap_values` columns) to the description of the category
# * `raw`: this field contains:
# * `raw_prediction`: a `samples x n_outputs` array of predictions for each instance to be explained. Note that this is calculated by applying the link function specified in `link` to the output of `pred_fn`
# * `prediction`: a `samples` array containing the index of the maximum value in the `raw_prediction` array
# * `instances`: a `samples x n_features` array of instances which have been explained
# * `importances`: a dictionary where each entry is a dictionary containing the sorted average magnitude of the shap value (`ranked_effect`) along with a list of feature names corresponding to the re-ordered shap values (`names`). There are `n_outputs + 1` keys, corresponding to `n_outputs` and to the aggregated output (obtained by summing all the arrays in `shap_values`)
#
# Please see our examples on how to visualise these outputs using the `shap` library visualisations [here](../examples/kernel_shap_wine_intro.ipynb), [here](../examples/kernel_shap_wine_lr.ipynb) and [here](../examples/kernel_shap_adult_lr.ipynb).
# ### Explaining heterogeneous (continuous and categorical) datasets
# When the dataset contains both continuous and categorical variables, `categorical_names`, an optional mapping from the encoded categorical features to a description of the category can be passed in addition to the `feature_names` list. This mapping is currently used for determining what type of summarisation should be applied if `X_reference` is large and the `fit` argument `summarise_background='auto'` or `summarise_background=True` but in the future it might be used for annotating visualisations. The definition of the map depends on what method is used to handle the categorical variables.
# #### By grouping categorical data
# By grouping categorical data we estimate a single shap value for each categorical variable.
# ##### Initialisation and fit
# Assume that we have a dataset with features such as `Marital Status` (first column), `Age` (2nd column), `Income` (3rd column) and `Education` (4th column). The 2nd and 3rd column are continuous variables, whereas the 1st and 4th are categorical ones.
#
# The mapping of categorical variables could be generated from a Pandas dataframe using the utility `gen_category_map`, imported from `alibi.utils.data`. For this example the output could look like:
#
# ```python
# category_map = {
# 0: ["married", "divorced"],
# 3: ["high school diploma", "master's degree"],
# }
# ```
# Hence, using the same predict function as before, we initialise the explainer as:
#
# ```python
# explainer = KernelShap(
# predict_fn,
# link='logit',
# feature_names=["Marital Status", "Age", "Income", "Education"],
# categorical_names=category_map,
# )
# ```
#
# To group our data, we have to provide the `groups` list, which contains lists with indices that are grouped together. In our case this would be:
# ```python
# groups = [[0, 1], [2], [3], [4, 5]]
# ```
# Similarly, the group_names are the same as the feature names
# ```python
# group_names = ["Marital Status", "Age", "Income", "Education"]
# ```
# Note that, in this case, the keys of the `category_map` are indices into `groups`. To fit our explainer we pass _one-hot encoded_ data to the explainer along with the grouping information.
#
# ```python
# explainer.fit(
# X_reference,
# group_names=group_names,
# groups=groups,
# )
# ```
# ##### Explanation
# To perform an explanation, we pass _one hot encoded_ instances `X` to the `explain` method:
#
# ```python
# explanation = explainer.explain(X)
# ```
# The explanation returned will contain the grouping information in its `meta` attribute
#
# ```python
# {'name': 'KernelShap',
# 'type': ['blackbox'],
# 'explanations': ['local', 'global'],
# 'params': {'groups': [[0, 1], [2], [3], [4, 5]],
# 'group_names': ["Marital Status", "Age", "Income", "Education"] ,
# 'weights': None,
# 'summarise_background': False
# }
# }
# ```
# whereas inspecting the `data` attribute shows that one shap value is estimated for each of the four groups:
#
# ```python
# {'shap_values': [array([ 0.8340445 , 0.12000589, -0.07984099, 0.61758141]),
# array([-0.71522546, 0.31749045, 0.3146705 , -0.13365639]),
# array([-0.12984616, -0.47194649, -0.23036243, -0.52314911])],
# 'expected_value': array([0.74456904, 1.05058744, 1.15837362]),
# 'link': 'logit',
# 'feature_names': ["Marital Status", "Age", "Income", "Education"],
# 'categorical_names': {},
# 'raw': {
# 'raw_prediction': array([ 2.23635984, 0.83386654, -0.19693058]),
# 'prediction': array([0]),
# 'instances': array([ 0.93884707, -0.63216607, -0.4350103 , -0.91969562]),
# 'importances': {
# '0': {'ranked_effect': array([0.8340445 , 0.61758141, 0.12000589, 0.07984099]),
# 'names': ['a', 'd', 'b', 'c']},
# '1': {'ranked_effect': array([0.71522546, 0.31749045, 0.3146705 , 0.13365639]),
# 'names': ['a', 'b', 'c', 'd']},
# '2': {'ranked_effect': array([0.52314911, 0.47194649, 0.23036243, 0.12984616]),
# 'names': ['d', 'b', 'c', 'a']},
# 'aggregated': {'ranked_effect': array([1.67911611, 1.27438691, 0.90944283, 0.62487392]),
# 'names': ['a', 'd', 'b', 'c']}
# }
# }
# }
# ```
# #### By summing output
# An alternative to grouping, with a higher runtime cost, is to estimate one shap value for each dimension of the one-hot encoded data and sum the shap values of the encoded dimensions to obtain only one shap value per categorical variable.
# ##### Initialisation and fit
# The initialisation step is as before:
#
# ```python
# explainer = KernelShap(
# predict_fn,
# link='logit',
# feature_names=["Marital Status", "Age", "Income", "Education"],
# categorical_names=category_map,
# )
# ```
#
# However, note that the keys of the `category_map` have to correspond to the locations of the categorical variables after the effects for the encoded dimensions have been summed up (see details below).
#
# The fit step requires _one hot encoded_ data and simply takes the reference dataset:
# ```python
# explainer.fit(X_reference)
# ```
# ##### Explanation
# To obtain a single shap value per categorical result, we have to specify the following arguments to the `explain` method:
#
# * `summarise_result`: indicates that some shap values will be summed
# * `cat_vars_start_idx`: the column indices where the first encoded dimension is for each categorical variable
# * `cat_vars_enc_dim`: the length of the encoding dimensions for each categorical variable
#
# ```python
# explanation = explainer.explain(
# X,
# summarise_result=True,
# cat_vars_start_idx=[0, 4],
# cat_vars_enc_dim=[2, 2],
# )
# ```
#
#
# In our case `Marital Status` starts at column `0` and occupies 2 columns, `Age` and `Income` occupy columns `2` and `3` and `Education` occupies columns `4` and `5`.
# #### By combining preprocessor and predictor
# Finally, an alternative is to combine the preprocessor and the predictor together in the same object, and fit the explainer on data _before preprocessing_.
# ##### Initialisation and fit
# To do so, we first redefine our predict function as
# ```python
# predict_fn = lambda x: clf.predict(preprocessor.transform(x))
# ```
# The explainer can be initialised as:
# ```python
#
# explainer = KernelShap(
# predict_fn,
# link='logit',
# feature_names=["Marital Status", "Age", "Income", "Education"],
# categorical_names=category_map,
# )
# ```
# Then, the explainer should be fitted on _unprocessed_ data:
#
# ```python
# explainer.fit(X_referennce_unprocessed)
# ```
# ##### Explanation
# We can explain _unprocessed records_ simply by calling `explain`:
#
# ```python
# explanation = explainer.explain(X_unprocessed)
# ```
# ### Running batches of explanations in parallel
# Increases in the size of the background dataset, the number of samples used to estimate the shap values or simply explaining a large number of instances dramatically increase the cost of running Kernel SHAP.
#
# To explain batches of instances in parallel, first run ``pip install alibi[ray]`` to install required dependencies and then simply initialise `KernelShap` specifying the number of physical cores available as follows:
#
# ```python
# distrib_kernel_shap = KernelShap(predict_fn, distributed_opts={'n_cpus': 10}
# ```
#
# To explain, simply call the `explain` as before - no other changes are required.
# ### Miscellaneous
# #### Runtime considerations
# For a given instance, the runtime of the algorithm depends on:
#
# * the size of the reference dataset
# * the dimensionality of the data
# * the number of samples used to estimate the shap values
# ##### Adjusting the size of the reference dataset
# The algorithm automatically warns the user if a background dataset size of more than `300` samples is passed. If the runtime of an explanation with the original dataset is too large, then the algorithm can automatically subsample the background dataset during the `fit` step. This can be achieve by specifying the fit step as
#
# ```python
# explainer.fit(
# X_reference,
# summarise_background=True,
# n_background_samples=150,
# )
# ```
#
# or
# ```python
# explainer.fit(
# X_reference,
# summarise_background='auto'
# )
# ```
#
# The `auto` option will select `300` examples, whereas using the boolean argument allows the user to directly control the size of the reference set. If categorical variables or grouping options are specified, the algorithm uses subsampling of the data. Otherwise, a kmeans clustering algorithm is used to select the background dataset and the samples are weighted according to the frequency of occurrence of the cluster they are assigned to, which is reflected in the `expected_value` attribute of the explainer.
#
# As described above, the explanations are performed with respect to the expected (or weighted-average) output over this dataset so the shap values will be affected by the dataset selection. We recommend experimenting with various ways to choose the background dataset before deploying explanations.
# ##### The dimensionality of the data and the number of samples used in shap value estimation
# The dimensionality of the data has a slight impact on the runtime, since by default the number of samples used for estimation is `2*n_features + 2**11`. In our experiments, we found that either grouping the data or fitting the explainer on unprocessed data resulted in run time savings (but did not run rigorous comparison experiments). If grouping/fitting on unprocessed data alone does not give enough runtime savings, the background dataset could be adjusted. Additionally (or alternatively), the number of samples could be reduced as follows:
#
# ```python
# explanation = explainer.explain(X, nsamples=500)
# ```
# We recommend experimenting with this setting to understand the variance in the shap values before deploying such configurations.
# #### Imbalanced datasets
# In some situations, the reference datasets might be imbalanced so one might wish to perform an explanation of the model behaviour around $x$ with respect to $\sum_{i} w_i f(y_i)$ as opposed to $\mathbb{E}_{\mathcal{D}}[f(y)]$. This can be achieved by passing a list or an 1-D numpy array containing a weight for each data point in `X_reference` as the `weights` argument of the `fit` method.
# ## Theoretical overview
# Consider a model $f$ that takes as an input $M$ features. Assume that we want to explain the output of the model $f$ when applied to an input $x$. Since the model output scale does not have an origin (it is an [affine space](https://en.wikipedia.org/wiki/Affine_space)), one can only explain the difference of the observed model output with respect to a chosen origin point. This point can be taken to be the function output value for an arbitrary record or the average output over a set of records, $\mathcal{D}$. Assuming the latter case, for the explanation to be accurate, one requires
#
# $$
# f(x) - \mathbb{E}_{y \sim \mathcal{D}}[f(y)] = \sum_{i=1}^M \phi_i
# $$
#
# where $\mathcal{D}$ is also known as a _background dataset_ and $\phi_i$ is the portion of the change attributed to the $i$th feature. This portion is sometimes referred to as feature importance, effect or simply shap value.
#
# One can conceptually imagine the estimation process for the shap value of the $i^{th}$ feature $x_i$ as consisting of the following steps:
#
# - enumerate all subsets $S$ of the set $F = \{1, ..., M\} \setminus \{i\}$
# - for each $S \subseteq F \setminus \{i\}$, compute the contribution of feature $i$ as $C(i|S) = f(S \cup \{i\}) - f(S)$
# - compute the shap value according to
# $$
# \phi_i := \frac{1}{M} \sum \limits_{{S \subseteq F \setminus \{i\}}} \frac{1}{ M - 1 \choose |S|} C(i|S).
# $$
#
# The semantics of $f(S)$ in the above is to compute $f$ by treating $\bar{S}$ as missing inputs. Thus, we can imagine the process of computing the SHAP explanation as starting with $S$ that does not contain our feature, adding feature $i$ and then observing the difference in the function value. For a nonlinear function the value obtained will depend on which features are already in $S$, so we average the contribution over all possible ways to choose a subset of size $|S|$ and over all subset sizes. The issue with this method is that:
#
# - the summation contains $2^M$ terms, so the algorithm complexity is $O(M2^M)$
# - since most models cannot accept an arbitrary pattern of missing inputs at inference time, calculating $f(S)$ would involve model retraining the model an exponential number of times
#
# To overcome this issue, the following approximations are made:
#
# - the missing features are simulated by replacing them with values from the background dataset
# - the feature attributions are estimated instead by solving
#
# $$
# \min \limits_{\phi_i, ..., \phi_M} \left\{ \sum \limits_{S \subseteq F} \left[ f(S) - \sum \limits_{j \in S} \phi_j \right]^2 \pi_x(S) \right\}
# $$
#
# where
#
# $$
# \pi_x(S) = \frac{M-1}{{M \choose |S|} |S|(M - |S|)}
# $$
#
# is the Shapley kernel (Figure 2).
# 
#
# Figure 2: Shapley kernel
# Note that the optimisation objective implies above an exponential number of terms. In practice, one considers a finite number of samples `n`, selecting `n` subsets $S_1, ..., S_n$ according to the probability distribution induced by the kernel weights. We can see that the kernel favours either small or large subset sizes, since most of the information about the effect of a particular feature for an outcome change can be obtained by excluding that feature or excluding all the features except for it from the input set.
#
# Therefore, Kernel SHAP returns an approximation of the true Shapley values, whose variability depends on factors such as the size of the structure of the background dataset used to estimate the feature attributions and the number of subsets of missing features sampled. Whenever possible, algorithms specialised for specific model structures (e.g., Tree SHAP, Linear SHAP, integrated gradients) should be used since they are faster and more accurate.
# ### Comparison to other methods
# Like [LIME](https://arxiv.org/abs/1602.04938), this method provides *local explanations*, in the sense that the attributions are estimated to explain the change from a baseline *for a given data point*, $x$. LIME computes the feature attributions by optimising the following objective in order to obtain a locally accurate explanation model (i.e., one that approximates the model to explained well around an instance $x$):
#
# $$
# \zeta = \text{arg}\min_{g \in \mathcal{G}} L(f, g, \pi_{x}) + \Omega(g).
# $$
#
# Here $f$ is the model to be explained, $g$ is the explanation model (assumed linear), $\pi$ is a local kernel around instance $x$ (usually cosine or $\ell_2$ kernel) and $\Omega(g)$ penalises explanation model complexity. The choices for $L, \pi$ and $\Omega$ in LIME are heuristic, which can lead to unintuitive behaviour (see [Section 5](https://papers.nips.cc/paper/7062-a-unified-approach-to-interpreting-model-predictions) of Lundberg et al. for a study). Instead, by computing the shap values according to the weighted regression in the previous section,
# the feature attributions estimated by Kernel SHAP have desirable properties such as *local accuracy* , _consistency_ and *missingness*, detailed in [Section 3](https://papers.nips.cc/paper/7062-a-unified-approach-to-interpreting-model-predictions) of Lundberg et al..
# Although, in general, local explanations are limited in that it is not clear to what a given explanation applies *around* and instance $x$ (see anchors algorithm overview [here](Anchors.ipynb) for a discussion), insights into global model behaviour can be drawn by aggregating the results from local explanations (see the work of Lundberg et al. [here](https://www.nature.com/articles/s42256-019-0138-9)). In the future, a distributed version of the Kernel SHAP algorithm will be available in order to reduce the runtime requirements necessary for explaining large datasets.
# ## Examples
# ### Continuous Data
# [Introductory example: Kernel SHAP on Wine dataset](../examples/kernel_shap_wine_intro.nblink)
# [Comparison with interpretable models](../examples/kernel_shap_wine_lr.nblink)
# ### Mixed Data
# [Handling categorical variables with Kernel SHAP: an income prediction application](../examples/kernel_shap_adult_lr.nblink)
# [Handlling categorical variables with Kernel SHAP: fitting explainers on data before pre-processing](../examples/kernel_shap_adult_categorical_preproc.nblink)
# [Distributed Kernel SHAP: paralelizing explanations on multiple cores](../examples/distributed_kernel_shap_adult_lr.nblink)
| doc/source/methods/KernelSHAP.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib
import numpy as np
import os
import yaml
import shutil
from collections import defaultdict
matplotlib.style.use('seaborn-notebook')
matplotlib.pyplot.rcParams['figure.figsize'] = (16, 10)
matplotlib.pyplot.rcParams['font.family'] = 'sans-serif'
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
# -
# # Task1
# +
# %time
exp_dir = "../exps/task1/two_headed_best/"
dev_outputs = defaultdict(list)
def load_exps(exp_dir):
experiments = []
for subdir in os.scandir(exp_dir):
dev_fn = os.path.join(subdir.path, "dev.out")
if not os.path.exists(dev_fn):
print("No dev.out: {}".format(subdir.path))
continue
config_fn = os.path.join(subdir.path, "config.yaml")
with open(config_fn) as f:
config = yaml.load(f)
config['config_language'] = config['language']
config['language'] = '-'.join(config['dev_file'].split('/')[-1].split('-')[:-1])
config['train_size'] = config['train_file'].split('/')[-1].split('-')[-1]
exp_d = config
dev_output = pd.read_table(dev_fn, names=["lemma", "inflected", "tags"])
if len(dev_output['inflected']) == 0:
print(subdir.path)
dev_outputs[(config['language'], config['train_size'])].append(dev_output)
dev_acc_fn = os.path.join(subdir.path, "dev.word_accuracy")
if not os.path.exists(dev_acc_fn):
print("Dev accuracy file does not exist in dir: {}".format(subdir.path))
else:
with open(dev_acc_fn) as f:
exp_d['dev_acc'] = float(f.read())
experiments.append(exp_d)
experiments = pd.DataFrame(experiments)
return experiments
experiments = load_exps(exp_dir)
sum(len(v) for v in dev_outputs.values())
# +
gold = {}
for fn in os.scandir("../data/conll2018/task1/all"):
if not fn.path.endswith('-dev'):
continue
language = '-'.join(fn.name.split('-')[:-1])
with open(fn.path) as f:
inflected = [l.strip().split('\t')[1] for l in f]
gold[language] = pd.Series(inflected)
for fn in os.scandir("../data/conll2018/task1/surprise"):
if not fn.path.endswith('-dev'):
continue
language = '-'.join(fn.name.split('-')[:-1])
with open(fn.path) as f:
inflected = [l.strip().split('\t')[1] for l in f]
gold[language] = pd.Series(inflected)
# -
experiments.groupby('train_size').dev_acc.describe()
# +
merged = {}
for (lang, size), outputs in dev_outputs.items():
output = pd.concat(outputs, axis=1)
merged[(lang, size)] = pd.concat(outputs, axis=1)['inflected']
merged[('latin', 'high')].head()
# +
results = []
for (lang, size), outputs in merged.items():
d = {'language': lang, 'train_size': size}
for i in range(outputs.shape[1]):
acc = (outputs.iloc[:, i] == gold[lang]).mean()
d[i] = acc
d['majority'] = (outputs.mode(axis=1).iloc[:, 0] == gold[lang]).mean()
results.append(d)
# -
dev_results = pd.DataFrame(results)
dev_results['majority_wins'] = dev_results[dev_results.columns[2:-1]].max(axis=1) < dev_results.majority
dev_results[dev_results['train_size']=='low'].majority_wins.value_counts()
dev_results.majority_wins.value_counts()
# ### Save best configs
# +
outdir = "../config/task1/highest_dev_acc"
keep = ['batch_size', 'commit_hash', 'dataset_class', 'dropout', 'early_stopping_window', 'epochs',
'inflected_embedding_size', 'inflected_hidden_size', 'inflected_num_layers',
'lemma_embedding_size', 'lemma_hidden_size', 'lemma_num_layers', 'model',
'numpy_random_seed', 'optimizer', 'share_vocab',
'tag_embedding_size', 'tag_hidden_size', 'tag_num_layers', 'train_file', 'dev_file',
'save_min_epoch', 'torch_random_seed']
for (lang, train_size), idx in experiments.groupby(['language', 'train_size']).dev_acc.idxmax().iteritems():
out_fn = os.path.join(outdir, "{}_{}.yaml".format(lang, train_size))
in_config = experiments.loc[idx].experiment_dir + "/config.yaml"
with open(in_config) as f:
cfg = yaml.load(f)
cfg = {k: v for k, v in cfg.items() if k in keep}
cfg['train_file'] = 'data/conll2018' + cfg['train_file'].split('conll2018')[1]
cfg['dev_file'] = 'data/conll2018' + cfg['dev_file'].split('conll2018')[1]
with open(out_fn, 'w') as f:
yaml.dump(cfg, f, default_flow_style=False)
# -
# # Submission 1
#
# Model with the highest dev acc, no majority voting
output_dir = "../submissions/task1/01/task1"
for (lang, train_size), idx in experiments.groupby(['language', 'train_size']).dev_acc.idxmax().iteritems():
exp = experiments.loc[idx]
test_output = os.path.join(exp.experiment_dir, "test.out")
submission_path = os.path.join(output_dir, "{}-{}-out".format(lang, train_size))
shutil.copy2(test_output, submission_path)
# # Submission 2
#
# Simple majority vote
output_dir = "../submissions/task1/02/task1"
for (lang, train_size), group in experiments.groupby(['language', 'train_size']).groups.items():
all_outputs = None
for i, idx in enumerate(group):
exp = experiments.loc[idx]
test_fn = os.path.join(exp.experiment_dir, "test.out")
test_output = pd.read_table(test_fn, names=['lemma', i, 'tags'])
if all_outputs is None:
all_outputs = test_output[[i]]
else:
all_outputs = pd.concat((all_outputs, test_output[[i]]), axis=1)
majority = pd.concat((test_output[['lemma']], all_outputs.mode(axis=1)[0], test_output[['tags']]), axis=1)
output_fn = os.path.join(output_dir, "{}-{}-out".format(lang, train_size))
majority.to_csv(output_fn, index=False, header=False, sep="\t", na_rep='nan')
# # Submission 3
#
# batch size = 16
# +
# %time
exp_dir = "../exps/task1/two_headed_best/"
def load_exps(exp_dir):
experiments = []
for subdir in os.scandir(exp_dir):
dev_fn = os.path.join(subdir.path, "dev.batch16.out")
if not os.path.exists(dev_fn):
print("No dev.out: {}".format(subdir.path))
continue
config_fn = os.path.join(subdir.path, "config.yaml")
with open(config_fn) as f:
config = yaml.load(f)
config['config_language'] = config['language']
config['language'] = '-'.join(config['dev_file'].split('/')[-1].split('-')[:-1])
config['train_size'] = config['train_file'].split('/')[-1].split('-')[-1]
exp_d = config
dev_acc_fn = os.path.join(subdir.path, "dev.batch16.word_accuracy")
if not os.path.exists(dev_acc_fn):
print("Dev accuracy file does not exist in dir: {}".format(subdir.path))
else:
with open(dev_acc_fn) as f:
exp_d['dev_acc'] = float(f.read())
experiments.append(exp_d)
experiments = pd.DataFrame(experiments)
return experiments
experiments = load_exps(exp_dir)
sum(len(v) for v in dev_outputs.values())
# -
output_dir = "../submissions/task1/03/task1"
for (lang, train_size), idx in experiments.groupby(['language', 'train_size']).dev_acc.idxmax().iteritems():
exp = experiments.loc[idx]
test_output = os.path.join(exp.experiment_dir, "test.out")
submission_path = os.path.join(output_dir, "{}-{}-out".format(lang, train_size))
shutil.copy2(test_output, submission_path)
# # Task2 - Track 1
# +
exp_dirs = ["../exps/task2/track1/final", "../exps/task2/track1/default"]
task2_track1 = []
for exp_dir in exp_dirs:
for subdir in os.scandir(exp_dir):
config_fn = os.path.join(subdir.path, "config.yaml")
with open(config_fn) as f:
config = yaml.load(f)
exp_d = config
exp_d['language'] = config['train_file'].split('/')[-1].split('-')[0]
exp_d['train_size'] = config['train_file'].split('/')[-1].split('-')[2]
dev_acc_fn = os.path.join(subdir.path, "dev.word_accuracy")
if not os.path.exists(dev_acc_fn):
print("Dev accuracy file does not exist in {}".format(subdir.path))
else:
with open(dev_acc_fn) as f:
exp_d['dev_acc'] = float(f.read())
test_out_fn = os.path.join(subdir.path, "test.out")
if not os.path.exists(test_out_fn):
print("Test output file does not exist in {}".format(subdir.path))
else:
exp_d['test_output_path'] = test_out_fn
task2_track1.append(exp_d)
task2_track1 = pd.DataFrame(task2_track1)
# -
task2_track1.groupby(['language', 'train_size']).dev_acc.agg(['mean', 'max', 'std', 'size'])
t = task2_track1
t[(t.language=='sv') & (t.train_size=='high')]
# ## Save best configs
# +
outdir = "../config/task2/track1/highest_dev_acc"
keep = ['batch_size', 'char_embedding_size', 'commit_hash', 'context_hidden_size', 'context_num_layers',
'dataset_class', 'decoder_hidden_size', 'decoder_num_layers',
'dev_file', 'dropout', 'early_stopping_window', 'epochs',
'include_same_forms_ratio', 'lemma_hidden_size', 'lemma_num_layers',
'min_epochs', 'model', 'numpy_random_seed', 'optimizer',
'share_context_encoder', 'share_embedding', 'share_vocab',
'tag_embedding_size', 'tag_hidden_size', 'tag_num_layers',
'save_min_epoch',
'torch_random_seed', 'train_file', 'word_hidden_size', 'word_num_layers']
for (language, train_size), idx in task2_track1.groupby(['language', 'train_size']).dev_acc.idxmax().iteritems():
out_fn = os.path.join(outdir, "{}_{}.yaml".format(language, train_size))
in_config = task2_track1.loc[idx].experiment_dir + "/config.yaml"
with open(in_config) as f:
cfg = yaml.load(f)
cfg = {k: v for k, v in cfg.items() if k in keep}
cfg['train_file'] = 'data/conll2018' + cfg['train_file'].split('conll2018')[1]
cfg['dev_file'] = 'data/conll2018' + cfg['dev_file'].split('conll2018')[1]
with open(out_fn, 'w') as f:
yaml.dump(cfg, f, default_flow_style=False)
# -
# ## Submission - there is only one
#
# choose the one with the highest dev acc
submission_dir = "../submissions/task2/01/task2"
for (language, train_size), idx in task2_track1.groupby(['language', 'train_size']).dev_acc.idxmax().iteritems():
experiment = task2_track1.loc[idx]
target_name = "{}-1-{}-out".format(language, train_size)
target_fn = os.path.join(submission_dir, target_name)
shutil.copy2(experiment.test_output_path, target_fn)
# +
exp_dirs = ["../exps/task2/track2/default/"]
task2_track2 = []
for exp_dir in exp_dirs:
for subdir in os.scandir(exp_dir):
config_fn = os.path.join(subdir.path, "config.yaml")
with open(config_fn) as f:
config = yaml.load(f)
exp_d = config
exp_d['language'] = config['train_file'].split('/')[-1].split('-')[0]
exp_d['train_size'] = config['train_file'].split('/')[-1].split('-')[2]
dev_acc_fn = os.path.join(subdir.path, "dev.word_accuracy")
if not os.path.exists(dev_acc_fn):
print("Dev accuracy file does not exist in {}".format(subdir.path))
else:
with open(dev_acc_fn) as f:
exp_d['dev_acc'] = float(f.read())
test_out_fn = os.path.join(subdir.path, "test.out")
if not os.path.exists(test_out_fn):
print("Test output file does not exist in {}".format(subdir.path))
else:
exp_d['test_output_path'] = test_out_fn
task2_track2.append(exp_d)
task2_track2 = pd.DataFrame(task2_track2)
# -
task2_track2.groupby(['language', 'train_size']).dev_acc.max().to_frame()
# ### Save best configs
# +
outdir = "../config/task2/track2/highest_dev_acc"
keep = ['batch_size', 'char_embedding_size', 'commit_hash', 'context_hidden_size', 'context_num_layers',
'dataset_class', 'decoder_hidden_size', 'decoder_num_layers',
'dev_file', 'dropout', 'early_stopping_window', 'epochs',
'include_same_forms_ratio', 'lemma_hidden_size', 'lemma_num_layers',
'min_epochs', 'model', 'numpy_random_seed', 'optimizer',
'share_context_encoder', 'share_embedding', 'share_vocab',
'tag_embedding_size', 'tag_hidden_size', 'tag_num_layers',
'save_min_epoch',
'torch_random_seed', 'train_file', 'word_hidden_size', 'word_num_layers']
for (language, train_size), idx in task2_track2.groupby(['language', 'train_size']).dev_acc.idxmax().iteritems():
out_fn = os.path.join(outdir, "{}_{}.yaml".format(language, train_size))
in_config = task2_track2.loc[idx].experiment_dir + "/config.yaml"
with open(in_config) as f:
cfg = yaml.load(f)
cfg = {k: v for k, v in cfg.items() if k in keep}
cfg['train_file'] = 'data/conll2018' + cfg['train_file'].split('conll2018')[1]
cfg['dev_file'] = 'data/conll2018' + cfg['dev_file'].split('conll2018')[1]
with open(out_fn, 'w') as f:
yaml.dump(cfg, f, default_flow_style=False)
# -
submission_dir = "../submissions/task2/01/task2"
for (language, train_size), idx in task2_track2.groupby(['language', 'train_size']).dev_acc.idxmax().iteritems():
experiment = task2_track2.loc[idx]
target_name = "{}-2-{}-out".format(language, train_size)
target_fn = os.path.join(submission_dir, target_name)
shutil.copy2(experiment.test_output_path, target_fn)
# # Sanity checks
#
# ## Task1
#
# 1. Is every submission file the same length as the input?
# 2. Does every line have 3 fields?
# 3. Do the inputs match?
# +
test_dir = "../data/conll2018/task1/all"
submission_dir = "../submissions/task1"
for submission_no in os.listdir(submission_dir):
for subm_file in os.scandir(os.path.join(submission_dir, submission_no, 'task1')):
language = '-'.join(subm_file.name.split('-')[:-2])
test_fn = os.path.join(test_dir, '{}-covered-test'.format(language))
if not os.path.exists(test_fn):
test_fn = os.path.join(test_dir, "..", "surprise", '{}-covered-test'.format(language))
with open(test_fn) as input_f, open(subm_file) as output_f:
for inp_line in input_f:
outp_line = next(output_f)
infd = inp_line.strip().split('\t')
outfd = outp_line.strip().split('\t')
try:
assert len(outfd) == 3
assert infd[0] == outfd[0]
assert infd[-1] == outfd[-1]
except AssertionError:
print(submission_no, subm_file.path, infd, outfd)
# -
# ## Task2
# +
test_dir = "../data/conll2018/task2/testsets/"
submission_dir = "../submissions/task2"
for submission_no in os.listdir(submission_dir):
for subm_file in os.scandir(os.path.join(submission_dir, submission_no, 'task2')):
language = subm_file.name.split('-')[0]
track = subm_file.name.split('-')[1]
test_fn = os.path.join(test_dir, '{}-track{}-covered'.format(language, track))
with open(test_fn) as input_f, open(subm_file) as output_f:
for inp_line in input_f:
outp_line = next(output_f)
if not inp_line.strip():
assert not outp_line.strip()
continue
infd = inp_line.strip().split('\t')
outfd = outp_line.strip().split('\t')
if track == '1':
try:
assert len(outfd) == 3
assert infd[2] == outfd[2]
except AssertionError:
print(inp_line)
print(outp_line)
print(submission_no, subm_file.path, infd, outfd)
elif track == '2':
pass
else:
raise ValueError("unknown track: {}".format(track))
# -
# # Create submission archives
#
# ## Task1
# + language="bash"
#
# old_pwd=$(pwd)
#
# for sub in $( ls ../submissions/task1); do
# tar_name="BME-HAS-$sub-1.tgz"
# echo $sub
# cd ../submissions/task1/$sub
# mkdir -p arabic_slovene_fixed
# mkdir -p arabic_slovene_fixed/task1
# cp task1/arabic* arabic_slovene_fixed/task1
# cp task1/slovene* arabic_slovene_fixed/task1
# cd arabic_slovene_fixed
# tar czf $tar_name task1
# mv $tar_name ../../../tgz/arabic_slovene_fixed/
# cd $old_pwd
# done
# + language="bash"
#
# for sub in $( ls ../submissions/task1); do
# tar_name="BME-HAS-$sub-1.tgz"
# cd ../submissions/task1/$sub
# tar czf $tar_name task1
# mv $tar_name ../../tgz
# cd -
# done
# + language="bash"
#
# for sub in $( ls ../submissions/task2); do
# tar_name="BME-HAS-$sub-2.tgz"
# cd ../submissions/task2/$sub
# tar czf $tar_name task2
# mv $tar_name ../../tgz
# cd -
# done
# -
| notebooks/create_submissions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="iws1EZcbaquY"
# # Text Classification using XGBoost and CountVectorizer
# + [markdown] id="3sYA5-5xayzK"
# This Code Template is for Text Classification using XGBoost along with Text Feature technique CountVectorizer from Scikit-learn in python.
#
# <img src="https://cdn.blobcity.com/assets/gpu_recommended.png" height="25" style="margin-bottom:-15px" />
# + [markdown] id="DxclZ9Jka773"
# ### Required Packages
# -
# !pip install nltk
# !pip install xgboost
# + colab={"base_uri": "https://localhost:8080/"} id="DVQ8QbNMa9SM" outputId="c17a2354-e5ca-4626-f5d9-8270beb87842"
import warnings
warnings.filterwarnings('ignore')
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as se
import re, string
import nltk
from nltk.tokenize import word_tokenize
from nltk.corpus import stopwords, wordnet
from nltk.stem import SnowballStemmer, WordNetLemmatizer
nltk.download('stopwords')
nltk.download('punkt')
nltk.download('averaged_perceptron_tagger')
nltk.download('wordnet')
from imblearn.over_sampling import RandomOverSampler
from sklearn.feature_extraction.text import CountVectorizer
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder
from sklearn.metrics import plot_confusion_matrix,classification_report
from xgboost import XGBClassifier
# + [markdown] id="vcKNbYf-cFXE"
# ### Initialization
# + [markdown] id="gMC9nA6dcFl2"
# Filepath of CSV file
# + id="XYms4d_4cLcc"
#filepath
file_path= ''
# + [markdown] id="tYMsqgb2gezr"
# **Target** variable for prediction.
# + id="8dqaqzN4ghl9"
target=''
# + [markdown] id="thVb7na9glcy"
# Text column containing all the text data
#
# + id="FWgps0zmgpB9"
text=""
# + [markdown] id="-wZ37AFSdTPW"
# ## Data Fetching
# + [markdown] id="u_bvM8SadTpO"
# Pandas is an open-source, BSD-licensed library providing high-performance, easy-to-use data manipulation and data analysis tools.
#
# We will use panda's library to read the CSV file using its storage path.And we use the head function to display the initial row or entry.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="vbgp9mOggxtI" outputId="7744a9b5-cef0-4382-9c79-8e6fd931153f"
df=pd.read_csv(file_path)
df.head()
# + [markdown] id="7FKVbIjqj-0_"
# ### Data Preprocessing
# + [markdown] id="renAqXpkj-35"
# Since the majority of the machine learning models in the Sklearn library doesn't handle string category data and Null value, we have to explicitly remove or replace null values. The below snippet have functions, which removes the null value if any exists. And convert the string classes data in the datasets by encoding them to integer classes.
# + id="gy_OFA53kD_J"
#convert to lowercase, strip and remove punctuations
def preprocess(text):
text = text.lower()
text=text.strip()
text=re.compile('<.*?>').sub('', text)
text = re.compile('[%s]' % re.escape(string.punctuation)).sub(' ', text)
text = re.sub('\s+', ' ', text)
text = re.sub(r'\[[0-9]*\]',' ',text)
text=re.sub(r'[^\w\s]', '', str(text).lower().strip())
text = re.sub(r'\d',' ',text)
text = re.sub(r'\s+',' ',text)
return text
# STOPWORD REMOVAL
def stopword(string):
a= [i for i in string.split() if i not in stopwords.words('english')]
return ' '.join(a)
#LEMMATIZATION
# Initialize the lemmatizer
wl = WordNetLemmatizer()
# This is a helper function to map NTLK position tags
def get_wordnet_pos(tag):
if tag.startswith('J'):
return wordnet.ADJ
elif tag.startswith('V'):
return wordnet.VERB
elif tag.startswith('N'):
return wordnet.NOUN
elif tag.startswith('R'):
return wordnet.ADV
else:
return wordnet.NOUN
# Tokenize the sentence
def lemmatizer(string):
word_pos_tags = nltk.pos_tag(word_tokenize(string)) # Get position tags
a=[wl.lemmatize(tag[0], get_wordnet_pos(tag[1])) for idx, tag in enumerate(word_pos_tags)] # Map the position tag and lemmatize the word/token
return " ".join(a)
def textfinalpreprocess(string):
return lemmatizer(stopword(preprocess(string)))
def data_preprocess(df, target):
df = df.dropna(axis=0, how = 'any')
df[target] = LabelEncoder().fit_transform(df[target])
return df
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="4ajKAt-1v220" outputId="ca5eebe4-7f11-4b60-d3c3-3499993809bf"
df = data_preprocess(df, target)
df[text] = df[text].apply(lambda x: textfinalpreprocess(x))
df.head()
# + [markdown] id="_pAHqMokjI0Q"
# ### Feature Selections
# + [markdown] id="mtsrehs6jJDK"
# It is the process of reducing the number of input variables when developing a predictive model. Used to reduce the number of input variables to both reduce the computational cost of modelling and, in some cases, to improve the performance of the model.
#
# We will assign all the required input features to X and target/outcome to Y.
# + id="_MNVdxu6jMjD"
X=df[text]
Y=df[target]
# + [markdown] id="RwwCPRS3Du30"
# #### Distribution Of Target Variable
# + colab={"base_uri": "https://localhost:8080/", "height": 405} id="dobw-AfYDy3g" outputId="d25a4d8b-5d1f-423b-ea12-5bf2d442fcfc"
plt.figure(figsize = (10,6))
se.countplot(Y)
# + [markdown] id="u3iVx_6v_-De"
# ### Data Splitting
# Since we are using a univariate dataset, we can directly split our data into training and testing subsets. The first subset is utilized to fit/train the model. The second subset is used for prediction. The main motive is to estimate the performance of the model on new data.
# + id="jKoGwzU3AAkW"
x_train,x_test,y_train,y_test=train_test_split(X,Y,test_size=0.2,random_state=123)
# + [markdown] id="ufHKpTiN90UC"
# ### Feature Transformation
# + [markdown] id="R_PpHDfQ-BJU"
# **CountVectorizer** is a text transformation technique provided by the scikit-learn library that transforms a given text into a vector on the basis of the frequency (count) of each word that occurs in the entire text.
#
# It converts a collection of text documents to a matrix of token counts
#
# ***For more information on CountVectorizer [click here](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html)***
# + id="WU9H2MXk-b70"
vectorizer = CountVectorizer()
vectorizer.fit(x_train)
x_train = vectorizer.transform(x_train)
x_test = vectorizer.transform(x_test)
# -
x_train,y_train = RandomOverSampler(random_state=123).fit_resample(x_train, y_train)
# + [markdown] id="d4ByFdUKCQ7k"
# ## Model
# + [markdown] id="DOhgjmbaCQwC"
# XGBoost is an optimized distributed gradient boosting library designed to be highly efficient, flexible and portable. It implements machine learning algorithms under the Gradient Boosting framework. XGBoost provides a parallel tree boosting (also known as GBDT, GBM) that solve many data science problems in a fast and accurate way.
# + [markdown] id="r1YQa7jLCXUt"
#
# #### Model Tuning Parameters
# > - booster -> Which booster to use. Can be gbtree, gblinear or dart; gbtree and dart use tree based models while gblinear uses linear functions.
#
# > - validate_parameters -> When set to True, XGBoost will perform validation of input parameters to check whether a parameter is used or not. The feature is still experimental. It’s expected to have some false positives.
#
# > - num_pbuffer -> Size of prediction buffer, normally set to number of training instances. The buffers are used to save the prediction results of last boosting step.
#
# > - num_feature -> Feature dimension used in boosting, set to maximum dimension of the feature
#
# Read more at [xgboost.readthedocs.io](https://xgboost.readthedocs.io/en/latest/parameter.html)
# + colab={"base_uri": "https://localhost:8080/"} id="uQ1b_UYhAoB4" outputId="ead85146-b112-4297-9986-c59e33dc318c"
model=XGBClassifier()
model.fit(x_train,y_train)
# + [markdown] id="GYatcCDDC2JP"
# #### Model Accuracy
# score() method return the mean accuracy on the given test data and labels.
#
# In multi-label classification, this is the subset accuracy which is a harsh metric since you require for each sample that each label set be correctly predicted.
# + colab={"base_uri": "https://localhost:8080/"} id="C8b5oAH7C3Ya" outputId="03064f1b-1706-403a-bd46-aa723fcd8a53"
print("Accuracy score {:.2f} %\n".format(model.score(x_test,y_test)*100))
# + [markdown] id="yas-d-CAC9bz"
# #### Confusion Matrix
# A confusion matrix is utilized to understand the performance of the classification model or algorithm in machine learning for a given test set where results are known.
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="MXz0euBYC-95" outputId="6e7b964e-0d72-4182-b497-9c049ef6ba7c"
plot_confusion_matrix(model,x_test,y_test,cmap=plt.cm.Blues)
# + [markdown] id="gny6G4OsDBTT"
# #### Classification Report
# A Classification report is used to measure the quality of predictions from a classification algorithm. How many predictions are True, how many are False.
# * where:
# - Precision:- Accuracy of positive predictions.
# - Recall:- Fraction of positives that were correctly identified.
# - f1-score:- percent of positive predictions were correct
# - support:- Support is the number of actual occurrences of the class in the specified dataset.
# + colab={"base_uri": "https://localhost:8080/"} id="AowdAdt4DaPh" outputId="9e53e419-38e2-4f51-8f4e-440260a854cc"
print(classification_report(y_test,model.predict(x_test)))
# + [markdown] id="R2tcbGJGDhK4"
#
# #### Creator: <NAME> , Github: [Profile](https://github.com/Sbhawal)
| Natural Language Processing/NLP/TextClassification_XGBoost_CountVectorizer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
arr = np.array([[1,2],[3,4]])
arr
np.diag(arr)
np.arange(10)
a = np.random.rand(4,3)
a
a = np.random.randn(4,3)
a
a = np.random.randint(100)
a < 35
import re
l = "a,b c,d e f-9887;2BaB"
re.split(",| |-|;",l)
l = "EBOLA _VIRUS22"
l
l[4:2:-1]
arr = np.arange(45).reshape(3,3,5)
arr
arr2 = arr.ravel()
arr2
arr2 = arr2.reshape(5,9)
arr2
arr2.shape
arr2.T
arr.resize(5,9)
arr
arr.shape
arr[-1]
arr[1:4,2:-1]
arr[1:4,[1,2,6,7]]
arr = arr.reshape(3,-1)
arr
# +
a = np.array([[1,2],[3,4]])
b = np.array([[5,6],[7,8]])
c = np.vstack((a,b))
c
# -
d = np.hstack((a,b))
d
np.concatenate((a,b),axis=0)
np.concatenate((a,b),axis=1)
np.column_stack((a,b))
np.row_stack((a,b))
# +
a = np.array([1,2])
b = np.array([3,4])
np.column_stack((a,b))
# -
np.row_stack((a,b))
| 1/Part 1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # FINN Preprocessor: Process MODIS Burned Area into polygon Shapefile
# ## 1. Setting Envoronments
# ### Systems settings
# Most likely no need to be edited.
# +
# python libraries
import sys
import os
import re
import glob
import datetime
import subprocess
import shlex
from urllib.parse import urlparse
from importlib import reload
import gdal
import matplotlib.pylab as plt
# finn preproc codes
sys.path = sys.path + ['../code_anaconda']
import downloader
import af_import
import rst_import
import polygon_import
import run_step1
import run_step2
import export_shp
import plotter
# -
# database settings
os.environ['PGDATABASE'] = 'gis'
os.environ['PGUSER'] = 'finn'
os.environ['PGPASSWORD'] = '<PASSWORD>'
os.environ['PGHOST'] = 'localhost'
os.environ['PGPORT'] = '5432'
# Make sure that the PostGIS database is ready.
# show info for the database
# !psql postgres -c 'SELECT version();'
# !pg_lsclusters
# TODO i want to move this to Dockerfile somehow
# create plpython, needed only once for the database
try:
p = subprocess.run(shlex.split("psql -d %s -c 'CREATE LANGUAGE plpython3u;'" % os.environ['PGDATABASE']),
check=True, stderr=subprocess.PIPE)
except subprocess.CalledProcessError as e:
if 'already exists' in e.stderr.decode():
print(e.stderr.decode().replace('ERROR','OK').strip())
pass
# ### Settings for Burnt Area Datasets
# MODIS burned area will be downloaded, if needed, for the region specified by the rectangle
# 1. `year_rst`, `month_rst`: MODIS burned area year/month to be processed
# 2. either
# `four_corners`: (LowerLeft_lon, LowerLeft_Lat, UpperRight_lon, UpperRight_lat) or
# `extent_shp`: shape file (could be polygon of area of interest, points of fires)
# +
# MODIS raster datasets' year
#first_year_rst = 2017
#first_month_rst = 1
first_year_rst = 2018
first_month_rst = 1
last_year_rst = 2018
last_month_rst = 11
# tag to identify dataset
tag_bdt = 'modbdt_2018_tx'
# +
# Geographic extent of download
# specify either one of below (comment out one line with #)
four_corners = (-107, 25, -93, 37) # LL corner lon, LL corner LAT, UR corner Lon, UR corner Lat)
#extent_shp = './north_central_america.shp' # shape file of North and Central America (i can create this from AllRegion polygon)
# +
# get year/month series
yrmo0 = datetime.date(first_year_rst, first_month_rst, 1)
yrmo1 = datetime.date(last_year_rst, last_month_rst, 1)
yrmos = [yrmo0 + datetime.timedelta(days=_) for _ in range((yrmo1-yrmo0).days)]
yrmos = [_ for _ in yrmos if _.day == 1]
yrmos.append(yrmo1)
# tags to identify datasets, automatically set to be modlct_YYYY, modvcf_YYYY
#tag_bdt = 'modbdt_%d%02d' % (year_rst, month_rst)
tags_bdt = ['modbdt_%d%02d' % (yrmo.year, yrmo.month) for yrmo in yrmos]
tags_bdt
# -
# ---
# ## 2. Download raster datasets
# Raster files URL and directories to save data
# all downloads are stored in following dir
download_rootdir = '../downloads'
# +
# earthdata's URL for BA
#url_bdt = 'https://e4ftl01.cr.usgs.gov/MOTA/MCD64A1.006/%d.%02d.01/' % (year_rst, month_rst)
urls_bdt = ['https://e4ftl01.cr.usgs.gov/MOTA/MCD64A1.006/%d.%02d.01/' % (yrmo.year, yrmo.month) for yrmo in yrmos]
#ddir_bdt = download_rootdir +'/'+ ''.join(urlparse(url_bdt)[1:3])
ddirs_bdt = [download_rootdir +'/'+ ''.join(urlparse(url_bdt)[1:3]) for url_bdt in urls_bdt]
print('BDT downloads goes to %s' % ddirs_bdt)
# -
# Download burned area raster, <b>only for the tiles needed for the active fire file</b>
if 'four_corners' in locals() and not four_corners is None:
# use four corner
poly = "POLYGON((%f %f, %f %f, %f %f, %f %f, %f %f))" % (
four_corners[0], four_corners[1],
four_corners[0], four_corners[3],
four_corners[2], four_corners[3],
four_corners[2], four_corners[1],
four_corners[0], four_corners[1],
)
elif 'extent_shp' in locals() and not extent_shp is None:
# use shape file
poly = extent_shp
else:
raise RuntimeError('Specify region of interest!')
reload(downloader)
for url_bdt in urls_bdt:
downloader.download_only_needed(url = url_bdt, droot = download_rootdir, pnts=poly)
# Verify BDT files' checksum. If a file is corrupted, the file is downloaded again.
for ddir_bdt, url_bdt in zip(ddirs_bdt, urls_bdt):
downloader.purge_corrupted(ddir = ddir_bdt, url=url_bdt)
# ## 3. Import raster datasets
# Downloaded files need preprocessing, which is to extract the only raster band needed, and also make coordinate system to be WGS84. Intermediate files are created in following directories.
# +
workdir_bdt = '../proc_rst_%s' % tag_bdt
print('BDT preprocessing occurs in %s' % workdir_bdt)
# -
# ### Import Buned Area
# First grab hdf file names from the download directory
fnames_bdt = {}
n = 0
for ddir_bdt, yrmo in zip(ddirs_bdt, yrmos):
search_string = "%(ddir_bdt)s/MCD64A1.A%(year_rst)s???.h??v??.006.*.hdf" % dict(
ddir_bdt = ddir_bdt, year_rst=yrmo.year)
fnames_bdt[yrmo] = sorted(glob.glob(search_string))
n += len(fnames_bdt[yrmo])
print('found %d hdf files' % n )
if n == 0:
raise RuntimeError("check if downloads are successful and search string to be correct: %s" % search_string)
# The next command performs three tasks, "merge", "resample" and "import". First two task creates intermediate GeoTiff files in <i>work_dir</i>. Last task actually import the data into database's <i>raster</i> schema.
#
# You can run only selected tasks with run_XXX flags to `False`, when you know that processing failed in the middle and you resolved the issue.
reload(rst_import)
for yrmo in yrmos:
print(yrmo)
rst_import.main(tag_bdt, fnames=fnames_bdt[yrmo], workdir = workdir_bdt,
run_merge=True, run_resample=True, run_import=True)
# merge shapes into one
shpfiles = glob.glob(os.path.join(workdir_bdt, 'rsp', 'MCD64A1.A???????.shp'))
oname = os.path.join(workdir_bdt, '.'.join(('MCD64A1', tag_bdt, 'shp')))
cmd = ['ogr2ogr', '-update', '-append', oname, ]
for shp in shpfiles:
cmdx = cmd + [shp]
subprocess.run(cmdx, check=True)
| proc_rst_modbdt_2018_tx/main_proc_rst_modbdt_2018_tx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import sklearn.datasets
import sklearn.cluster
import sklearn.preprocessing
import sklearn.neural_network
import sklearn.linear_model
import sklearn.model_selection
import torch
import pandas as pd
datos = pd.read_csv('data.csv')
datos = np.array(datos,dtype = float)[:-1,1:]
X = datos[:,:-1]
Y = datos[:,-1]
Y[Y != 1] = 0
X_train, X_test, Y_train, Y_test = sklearn.model_selection.train_test_split(X, Y, test_size=0.7)
scaler = sklearn.preprocessing.StandardScaler()
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
# +
model = torch.nn.Sequential(
torch.nn.Conv1d(1, 15, kernel_size=15, stride=1),
torch.nn.MaxPool1d(kernel_size=6),
torch.nn.Conv1d(15, 1, kernel_size=2, stride=1),
torch.nn.Linear(25, 2)
)
criterion = torch.nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=0.2) #lr: learning rate
epochs = 60
loss_values = np.zeros(epochs)
F1_values_train = np.zeros(epochs)
F1_values_test = np.zeros(epochs)
for epoch in range(epochs):
X_new = np.expand_dims(X_train, 1)
inputs = torch.autograd.Variable(torch.Tensor(X_new).float())
targets = torch.autograd.Variable(torch.Tensor(Y_train).long())
optimizer.zero_grad()
out = model(inputs)
out = out.squeeze(dim=1) # necesario para quitar la dimension intermedia de channel
loss = criterion(out, targets)
loss.backward()
optimizer.step()
values, Y_predicted = torch.max(out.data, 1)
loss_values[epoch] = loss.item()
F1_values_train[epoch] = sklearn.metrics.f1_score(Y_train, Y_predicted, average='macro')
X_new = np.expand_dims(X_test, 1)
inputs_test = torch.autograd.Variable(torch.Tensor(X_new).float())
out_test = model(inputs_test)
out_test = out_test.squeeze(dim=1)
values, Y_predicted_test = torch.max(out_test.data, 1)
F1_values_test[epoch] = sklearn.metrics.f1_score(Y_test, Y_predicted_test, average='macro')
# -
plt.plot(np.arange(epochs), loss_values)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.plot(np.arange(epochs), F1_values_train, label='train')
plt.plot(np.arange(epochs), F1_values_test, label='test')
plt.xlabel('epoch')
plt.ylabel('F1')
plt.legend()
| deep_eeg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rlalastjd782/rlalastjd782.github.io/blob/main/chatbotproject/chatbot_project.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="AqhU02jVwkG-" outputId="f9149eaa-d835-4fce-a10c-70ff41f773b2"
import re
import requests
from bs4 import BeautifulSoup
import pandas as pd
# url 따오기
# 판례
# 뒷자리
numlist2= []
for num in range(3,600):
num = format(num,'05')
numlist2.append(num)
# 앞자리
numlist = []
for num in range(1,25):
num = format(num,'03')
numlist.append(num)
# url 코드
urls = []
for i in numlist:
for j in numlist2:
url = "https://www.klac.or.kr/legalinfo/counselView.do?folderId=000&scdFolderId=&pageIndex=1&searchCnd=0&searchWrd=&caseId=case-{0}-{1}".format(i,j)
urls.append(url)
#print(urls[0:10])
data = []
for url in urls[0:15] :
print(url)
webpage = requests.get(url)
#print(webpage.text)
soup = BeautifulSoup(webpage.content, "html.parser")
dt = soup.select_one("#print_page").text
dt = dt.replace("\n", "")
data.append(dt)
#print(data)
#data.strip()
#print(data[1])
df = pd.DataFrame({
'data' : data,
'url' : url
})
print(df)
| chatbotproject/chatbot_project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/bhukyavamshirathod/Cpp-Primer/blob/master/Image_Classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="_rFZDWlhGoJ8"
# **Step 1 : Gather Data**
#
# 1. Laptop
# 2. Bikes
# 3. Red Apple
# 4. Foot Ball
# + id="dvWWLwiBx0Wd" colab={"base_uri": "https://localhost:8080/"} outputId="db0e2dba-94f3-4d26-f58b-36ba1fd71171"
# !pip install ipython-autotime
# %load_ext autotime
# + colab={"base_uri": "https://localhost:8080/"} id="8YtHDCFCplgH" outputId="154aa5ed-3352-40d5-e1d2-fa3df12bda70"
# !pip install bing-image-downloader
# + colab={"base_uri": "https://localhost:8080/"} id="TMvLBubj-I4t" outputId="3d42a332-94a4-4ebc-9fda-67d712d63a3d"
from bing_image_downloader import downloader
downloader.download("Red Apple", limit=30, output_dir='Images_datasets', adult_filter_off=True, force_replace=False, timeout=60)
# + colab={"base_uri": "https://localhost:8080/"} id="DWBrm591dgTo" outputId="7c88ab19-c53b-4042-9d6f-91639a319e40"
downloader.download("laptop", limit=30, output_dir='Images_datasets', adult_filter_off=True, force_replace=False, timeout=60)
# + colab={"base_uri": "https://localhost:8080/"} id="P3re2-8RdjM7" outputId="1a682317-0aaa-4c4e-9f8c-d71f83e66598"
downloader.download("foot ball", limit=30, output_dir='Images_datasets', adult_filter_off=True, force_replace=False, timeout=60)
# + colab={"base_uri": "https://localhost:8080/"} id="jk64irRNcart" outputId="05fda794-e7e1-4eca-bedc-b9800ca6ad67"
downloader.download("Bikes", limit=30, output_dir='Images_datasets', adult_filter_off=True, force_replace=False, timeout=60)
# + colab={"base_uri": "https://localhost:8080/"} id="biKHOEKVGEVi" outputId="d165e835-335f-4300-86cc-42a5755739b2"
# %ls /content/Images_datasets
# + id="uWmr3r3TLb5v" colab={"base_uri": "https://localhost:8080/"} outputId="9f5013ac-d7dd-45b7-91b6-a0599bb75dc5"
# Importing Libraries
import numpy as np
import pandas as pd
import os
import matplotlib.pyplot as plt
from skimage.transform import resize
from skimage.io import imread
from sklearn.model_selection import train_test_split
from sklearn.naive_bayes import MultinomialNB
from sklearn.metrics import accuracy_score,confusion_matrix,classification_report
# + [markdown] id="yonh_DofyVR_"
# **Step 2 : Data Preprocessing**
# + id="PmpZ-FfCLxyb" colab={"base_uri": "https://localhost:8080/"} outputId="7fe30777-db39-462b-e0a1-2e6740876668"
# Flattening Images
Target = []
Images = []
Flated_Data = []
Data_Dir = '/content/Images_datasets'
CATEGORIES = ['laptop','Red Apple','foot ball','Bikes']
for i in CATEGORIES:
class_target = CATEGORIES.index(i)
path = os.path.join(Data_Dir,i)
for img in os.listdir(path):
img_array = imread(os.path.join(path,img))
img_resized = resize(img_array,(150,150,3))
Flated_Data.append(img_resized.flatten())
Images.append(img_resized)
Target.append(class_target)
Flated_Data = np.array(Flated_Data)
Target = np.array(Target)
Images = np.array(Images)
# + id="qwxaT9eaMiVX" colab={"base_uri": "https://localhost:8080/", "height": 456} outputId="32ffefc2-131f-4f35-8434-de43ef444b99"
# Creating DataFrame...
Data_Frame = pd.DataFrame(Flated_Data)
Data_Frame['Target'] = Target
Data_Frame
# + id="y6-FI15kTL3_" colab={"base_uri": "https://localhost:8080/"} outputId="be2a171d-2657-4c02-f637-2714ee1dde6b"
x = Flated_Data
y = Target
y
# + id="UiVfbQ1OQZk8" colab={"base_uri": "https://localhost:8080/"} outputId="61b41332-e9c0-4b97-eea8-04ff0e48fa4b"
# Splitting Data into Training and Testing sets...
x_train,x_test,y_train,y_test = train_test_split(x,y,random_state=0,stratify=y)
print(x_train.shape)
print(x_test.shape)
# + id="y_2VDSyBVt0L" colab={"base_uri": "https://localhost:8080/"} outputId="84ce72fe-9b2a-4667-ff2a-6743939f03fb"
np.unique(y_train,return_counts=True)
# + id="ykbzNC5LV6FS" colab={"base_uri": "https://localhost:8080/"} outputId="161a29fd-5611-4e20-9ebc-b18d4e1cdbca"
np.unique(y_test,return_counts=True)
# + [markdown] id="qybaHfu76V5P"
# **Step 3 : Algorithms**
# + id="kLXFlm90WAQS" colab={"base_uri": "https://localhost:8080/"} outputId="25d49561-032a-4047-9ffb-324cacf07ad2"
#Building Model and Traing the Model...
# from sklearn.svm import SVC # 79%
# model = SVC(C=1)
# model.fit(x_train,y_train)
from sklearn.linear_model import LogisticRegression # 93%
model = LogisticRegression(max_iter = 1000)
model.fit(x_train,y_train)
# from sklearn.multiclass import OneVsRestClassifier # 85%
# from sklearn.svm import LinearSVC
# model = OneVsRestClassifier(LinearSVC(random_state=0)).fit(x_train, y_train)
# from sklearn.naive_bayes import CategoricalNB
# model = CategoricalNB()
# model.fit(x_train,y_train)
# model = MultinomialNB() # 83%
# model.fit(x_train,y_train)
# + id="jl9ha4VTWDJZ" colab={"base_uri": "https://localhost:8080/"} outputId="376515cc-df3d-448e-e08c-0e4d0e8c1e71"
# Predicting and Testing the Model...
y_pred = model.predict(x_test)
y_pred
# + id="hsKqCXD63o3r" colab={"base_uri": "https://localhost:8080/"} outputId="475686b2-6552-4c1b-ce9b-d0bd1845f5ee"
print("Number of mislabeled points out of a total %d points : %d"
... % (x_test.shape[0], (y_test != y_pred).sum()))
# + [markdown] id="RDSMyXsE6deQ"
# **Step 4 : Evaluation**
# + id="SnfF9Eq6WRQR" colab={"base_uri": "https://localhost:8080/"} outputId="a4a6a2a5-5559-43fa-b8c8-1840ef593f1c"
# Evaluation of the Model...
confusion_matrix(y_pred,y_test)
# + id="cClQVvjjWYJT" colab={"base_uri": "https://localhost:8080/"} outputId="ea7a0b72-57d7-42ef-8a53-b4d0e76f3b52"
accuracy_score(y_pred,y_test)
# + id="FY3apU4UWaui" colab={"base_uri": "https://localhost:8080/"} outputId="bd782eb7-af6b-46d2-de93-2e8f3bd58a09"
print(classification_report(y_pred,y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="3AXyR83oqe7S" outputId="0cacc2be-256d-4bf3-8944-238c0d093d9b"
# Finding the Best Parameters while using SVM Algorithm...
# from sklearn.model_selection import GridSearchCV
# from sklearn import svm
# para_grid = [
# {'C':[1,10,100,1000],'kernel':['linear']},
# {'C':[1,10,100,1000],'gamma':[0.001,0.0001],'kernel':['rbf']}
# ]
# svc = svm.SVC(probability = True)
# clf = GridSearchCV(svc,para_grid)
# clf.fit(x_train,y_train)
# + [markdown] id="kp4gwmx0Qo0q"
# **Step 5 : Predicting the given Image**
# + id="Ekz3u8qa3BsX" colab={"base_uri": "https://localhost:8080/"} outputId="1129303d-c458-4e3e-8224-05795dde450a"
#Saving the Model
import pickle
pickle.dump(model,open('img_model.p','wb'))
# + id="35JzEwd63mZ8" colab={"base_uri": "https://localhost:8080/"} outputId="14409679-8828-493d-e0b7-9cd7047e7613"
model1 = pickle.load(open('img_model.p','rb'))
# + colab={"base_uri": "https://localhost:8080/", "height": 337} id="ur0zwoQ63vFM" outputId="776202b7-ff9f-478c-e1b7-91da390858ec"
# Testing a New Image
Flat_data = []
url = input('Enter Your URL : ')
img = imread(url)
img_resized = resize(img,(150,150,3))
Flat_data.append(img_resized.flatten())
Flat_data = np.array(Flat_data)
print(img.shape)
plt.imshow(img_resized)
y_out = model1.predict(Flat_data)
y_out = CATEGORIES[y_out[0]]
print(f' Predicted Output :{y_out}')
| Image_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# +
from random import choice
import tensorflow as tf
import pickle
import numpy as np
import pandas as pd
from annsa.template_sampling import *
# -
from annsa.load_dataset import load_full
# #### Load template dataset
# +
background_dataset = pd.read_csv('../.data/background_template_dataset.csv')
source_dataset = pd.read_csv('../data/shielded_templates_200kev_dataset.csv')
source_dataset, spectra_dataset, all_keys = load_full(source_dataset, background_dataset)
# -
# #### Create Dataset
# +
total_spectra = 1e1
dataset = {'sources': [],
'backgrounds': [],
'keys': []}
# simulate sources
isotopes_processed = 0
for isotope in set(source_dataset['isotope'].values):
for _ in np.arange(total_spectra):
background_cps = np.random.poisson(200)
integration_time = 10 ** np.random.uniform(np.log10(10), np.log10(3600))
signal_to_background = np.random.uniform(0.1, 3)
calibration = [np.random.uniform(0, 10),
np.random.uniform(0.8, 1.2),
0]
fwhm = choice([7.0, 7.5, 8.0])
source_spectrum, background_spectrum = make_random_spectrum(source_dataset,
background_dataset,
background_cps=background_cps,
integration_time=integration_time,
signal_to_background=signal_to_background,
calibration=calibration,
isotope=isotope,
fwhm=fwhm,)
dataset['sources'].append(source_spectrum)
dataset['backgrounds'].append(background_spectrum)
dataset['keys'].append(isotope)
isotopes_processed+=1
print(isotopes_processed)
np.save('spectra_dataset_complete_'+str(int(total_spectra)),dataset)
# -
| examples/source-interdiction/dataset-generation/Dataset-Generation-Complete.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="RdhQd8heU5jV" colab_type="code" colab={}
# !wget -c https://repo.continuum.io/archive/Anaconda3-5.2.0-Linux-x86_64.sh
# !chmod +x Anaconda3-5.2.0-Linux-x86_64.sh
# !bash ./Anaconda3-5.2.0-Linux-x86_64.sh -b -f -p /usr/local
# + id="uwjvlEbioC9I" colab_type="code" colab={}
# %env PYTHONPATH=
# + [markdown] id="PKh9n6J2zhVi" colab_type="text"
#
# + id="VWhJDRYYpgdJ" colab_type="code" colab={}
import sys
_ = (sys.path
.append("/usr/local/lib/python3.6/site-packages"))
# + id="d2rsmdP0WJp4" colab_type="code" colab={}
# !source activate -n pvnet python=3.7 --yes
# + id="6Tmgcvp8q3q_" colab_type="code" colab={}
# !source activate pvnet
# + id="I4q59w1PrcwT" colab_type="code" colab={}
# !conda init
# + id="Tm_oBruuNO4a" colab_type="code" colab={}
# %cd /content
# + id="FC-SUD5caa48" colab_type="code" colab={}
# !apt-get --purge remove cuda nvidia* libnvidia-*
# !dpkg -l | grep cuda- | awk '{print $2}' | xargs -n1 dpkg --purge
# !apt-get remove cuda-*
# !apt autoremove
# !apt-get update
# !wget https://developer.nvidia.com/compute/cuda/9.2/Prod/local_installers/cuda-repo-ubuntu1604-9-2-local_9.2.88-1_amd64 -O cuda-repo-ubuntu1604-9-2-local_9.2.88-1_amd64.deb
# !dpkg -i cuda-repo-ubuntu1604-9-2-local_9.2.88-1_amd64.deb
# !apt-key add /var/cuda-repo-9-2-local/7fa2af80.pub
# !apt-get update
# !apt-get install cuda-9.2
# !pip install http://download.pytorch.org/whl/cu92/torch-0.4.1-cp36-cp36m-linux_x86_64.whl
# !pip install torchvision
# !pip install http://download.pytorch.org/whl/cu92/torch-0.4.1-cp36-cp36m-linux_x86_64.whl
# + id="daaVbdFexSa8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="b52d3be5-1006-4817-a14e-f9259ba384b7"
# # !pip install torchvision
# !pip install http://download.pytorch.org/whl/cu92/torch-0.4.1-cp36-cp36m-linux_x86_64.whl
# + id="nEwtbfQCLRTE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="b366c833-6ee9-4f45-c7cc-06a1f2f83e33"
# !sudo apt-get install gcc-6
# !sudo apt-get install g++-6
# !sudo update-alternatives --install /usr/bin/g++ g++ /usr/bin/g++-6 10
# !sudo update-alternatives --install /usr/bin/gcc gcc /usr/bin/gcc-6 10
# + id="6WJhNGXIsVyx" colab_type="code" colab={}
# !sudo apt-get install libglfw3-dev libglfw3
# + id="-NbswY-zsiON" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="1028f05a-b654-4277-f1be-ad05bbdeb6a3"
# %cd /content/drive/'My Drive'/clean-pvnet-master
# + id="qS4xe9Nwsy2U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="6e0e2921-896c-45ce-a47f-be5dd92db5fb"
# # !pip install cyglfw3
# !pip install -r requirements.txt
# + id="n3SYLErWBV80" colab_type="code" colab={}
# !nvcc --version
# + id="SM66y23EtIE9" colab_type="code" colab={}
# %env ROOT=drive/'My Drive'/clean-pvnet-master
# + id="pudQhWijtpHH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="fd1d004e-59b7-47ff-d163-f2c1b03bd861"
# %env CUDA_HOME="/usr/local/cuda-9.0"
# %env LD_LIBRARY_PATH='$LD_LIBRARY_PATH:/usr/local/cuda-9.0/lib64'
# # %env CUDA_HOME=
# + id="Ll7S-8Bjts0X" colab_type="code" colab={}
# cd lib/csrc/dcn_v2/
# + id="t19tgkQMt60k" colab_type="code" colab={}
# !python setup.py build_ext --inplace
# + id="uLAZYFsZuree" colab_type="code" colab={}
# cd ../ransac_voting
# + id="JNQH41bWuv1S" colab_type="code" colab={}
# # !sudo apt install -lcudart-dev
# # !sudo ln -s /usr/local/cuda/lib64/libcudart.so.9.2.88 /usr/lib/libcudart.so
# !python setup.py build_ext --inplace
# + id="FZMZIl0pu58E" colab_type="code" colab={}
# cd ../nn
# + id="vB3aviR8u7Wm" colab_type="code" colab={}
# !python setup.py build_ext --inplace
# + id="Euu6-Gf3vCSU" colab_type="code" colab={}
# cd ../fps
# + id="1wG4buSXL3PS" colab_type="code" colab={}
# %cd data/custom
# !unzip /content/drive/My\ Drive/HybridPose/data/custom/masks/masks.zip -d mask
# + [markdown] id="y_oeS1xdM6QD" colab_type="text"
#
# + id="TvoKayj7M6qL" colab_type="code" colab={}
import os
for count, filename in enumerate(filter(lambda x: x.endswith('png'), os.listdir(os.path.join('mask/ship')))):
dst =filename.split('.png')[0].split('mask')[1]+'0001.jpg'
src ='mask/ship/'+ filename
dst ='mask/'+ dst
print(src)
print(dst)
# rename() function will
# rename all the files
os.rename(src, dst)
# + id="2zWkNfLdvFYl" colab_type="code" colab={}
# !python setup.py build_ext --inplace
# + id="cK8J_HGHvMnC" colab_type="code" colab={}
# cd ../uncertainty_pnp
# + id="0GhoItb7vPlR" colab_type="code" colab={}
# !sudo apt-get install libgoogle-glog-dev
# + id="R5C_oenUvUb9" colab_type="code" colab={}
# !sudo apt-get install libsuitesparse-dev
# + id="Vm3em_0xvY_l" colab_type="code" colab={}
# !sudo apt-get install libatlas-base-dev
# + id="XfvZ1bDsvc_m" colab_type="code" colab={}
# !python setup.py build_ext --inplace
# + id="9AyqaJBy0JPs" colab_type="code" colab={}
# !sudo dpkg --configure -a
# + id="cTbwt7FRwV9V" colab_type="code" colab={}
# cd data
# + id="x5GmyLVCwifD" colab_type="code" colab={}
# !sudo ln -s /content/drive/mydrive/clean-pvnet-master/ship_dataset /custom
# + id="pz7rf74gEcR5" colab_type="code" colab={}
# cd ..
# + id="CV3B2zH5qNdN" colab_type="code" colab={}
# !python train_net.py --cfg_file configs/custom.yaml train.batch_size 4 resume True
# + id="-YUPOUdlNxEw" colab_type="code" colab={}
# # !jupyter tensorboard enable
# %tensorboard --logdir data/record/pvnet
# + id="lK3QOf8vzQ6U" colab_type="code" colab={}
# !pip install tensorboard
# + id="xIfZqi9py41i" colab_type="code" colab={}
# %load_ext tensorboard
# + id="p4yTKE_fx9d7" colab_type="code" colab={}
# !pip install -q tf-nightly-2.0-preview
# Load the TensorBoard notebook extension
# %load_ext tensorboard
# + id="xe6xO673kw3f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="5d0bb835-0555-4164-bb89-d888ae704427"
# !pip install open3d-python
# !pip install open3d
# + id="AAoOe8nPOE1H" colab_type="code" colab={}
# !pip install tensorboard
# + id="-waZKQXfvbK6" colab_type="code" colab={}
# !pip install jupyter-tensorboard
# + id="fywSk41qfaUS" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="36e1fa55-3a58-4131-ef8b-253098447946"
# # !python run.py --type custom
# %cd /content/drive/'My Drive'/clean-pvnet-master
# !python run.py --type evaluate --cfg_file configs/custom.yaml
# + id="ED5WYYP09ln0" colab_type="code" colab={}
# %matplotlib --list
# + id="aM67hnHrMVDC" colab_type="code" colab={}
# !export LD_LIBRARY_PATH=/usr/local/cuda/lib64 && sudo ldconfig
# + id="le78eCSOSff7" colab_type="code" colab={}
from google.colab import drive
drive.mount('/content/drive')
# + id="OeSUx9_23PsZ" colab_type="code" colab={}
# %cd /content/drive/'My Drive'/pvnet-rendering
# !sudo python run.py --type rendering
# + id="kY0ofq18CSGz" colab_type="code" colab={}
# !pip install opencv-python
# + id="NkEgCGcerWrk" colab_type="code" colab={}
import cv2
import numpy as np
import glob
print('hi')
img_array = []
size=(640,480)
dirFiles=glob.glob('/content/drive/My Drive/clean-pvnet-master/test_video/*.jpg')
# dirFiles=glob.glob('/content/drive/My Drive/clean-pvnet-master/data/video/*.jpg')
dirFiles=sorted(dirFiles, key=lambda x: int(x.split(".")[0].split("video/")[1]))
for filename in dirFiles:
# if filename.contains('0001'):
# pass
print(filename)
img = cv2.imread(filename)
height, width, layers = img.shape
size = (width,height)
img_array.append(img)
# img_array.sort()
out = cv2.VideoWriter('/content/drive/My Drive/clean-pvnet-master/test_video/proj5.avi',cv2.VideoWriter_fourcc(*'DIVX'), 17, size)
for i in range(len(img_array)):
out.write(img_array[i])
out.release()
# + id="zaQ9y__UBUl_" colab_type="code" colab={}
import numpy
numpy.version.version
# + id="BIcIDQLiNaXv" colab_type="code" colab={}
# cd ../clean-pvnet-master/
# + id="27xncg6SWJya" colab_type="code" colab={}
cd ../pvnet-rendering/
# + id="xQkxG0XKNO-6" colab_type="code" colab={}
# !ln -s '/content/drive/My Drive/clean-pvnet-master/data/oceandatasety/MASATI-v2' MASATI
# + id="NVybZSt2kQOg" colab_type="code" colab={}
# !wget http://groups.csail.mit.edu/vision/SUN/releases/SUN2012pascalformat.tar.gz
# + id="VEd7DH0lopcz" colab_type="code" colab={}
# # !wget -O /usr/local/blender-2.79a-linux-glibc219-x86_64.tar.bz2 https://download.blender.org/release/Blender2.79/blender-2.79a-linux-glibc219-x86_64.tar.bz2
# !chmod +x blender-2.79a-linux-glibc219-x86_64/blender
# !chmod +x blender-2.79a-linux-glibc219-x86_64/2.79/python/bin/pip
# !chmod +x blender-2.79a-linux-glibc219-x86_64/2.79/python/bin/pip3
# !chmod +x blender-2.79a-linux-glibc219-x86_64/2.79/python/bin/python3.5m
# + id="VuQCwrMmTxzw" colab_type="code" colab={}
# !pip install msgpack
# !pip install easydict
# !pip install lmdb
# !pip install OpenEXR
# !pip install transforms3d
# # !sudo ./blender-2.79a-linux-glibc219-x86_64/2.79/python/bin/pip3 install opencv-python
# # !sudo ./blender-2.79a-linux-glibc219-x86_64/2.79/python/bin/pip3 --no-cache-dir install numpy
# # !sudo ./blender-2.79a-linux-glibc219-x86_64/2.79/python/bin/pip3 install numpy
# # !sudo ./blender-2.79a-linux-glibc219-x86_64/2.79/python/bin/pip3 install Pillow
# # !blender --version
# + id="I92UbItPozEQ" colab_type="code" colab={}
# + id="XNiCQJ1tuCvh" colab_type="code" colab={}
import os
os.environ['SDL_AUDIODRIVER'] = 'dsp'
# + id="PaSKwVL8ma6L" colab_type="code" colab={}
# !sudo python run.py --type fuse
# + id="Li7uwQ272gxa" colab_type="code" colab={}
import sys
_ = (sys.path
.append("/content/drive/'My Drive'/pvnet-rendering/blender/Automold--Road-Augmentation-Library"))
# + id="7Kl5hnSluu3S" colab_type="code" colab={}
# cp -r /usr/local/lib/python3.6/site-packages/transforms3d* /content/drive/'My Drive'/pvnet-rendering/blender-2.79a-linux-glibc219-x86_64/2.79/python/bin/site-packages
# + id="Y7KZYgIf_o0L" colab_type="code" colab={}
# !ln -s /content/drive/'My Drive'/pvnet-rendering/SUN2012 SUN
# + id="6z2FcYn_CLzi" colab_type="code" colab={}
# !nvcc --version
# + id="F1QwspiNIbNy" colab_type="code" colab={}
# !chmod +x ./blender-2.79a-linux-glibc219-x86_64/blender
# + id="5bTVt6edT-GZ" colab_type="code" colab={}
# !wget https://developer.download.nvidia.com/compute/cuda/repos/ubuntu1804/x86_64/cuda-repo-ubuntu1804_10.1.243-1_amd64.deb
# !sudo apt-key adv --fetch-keys https://developer.download.nvidia.com/compute/cuda/repos/ubuntu1804/x86_64/7fa2af80.pub
# !sudo dpkg -i cuda-repo-ubuntu1804_10.1.243-1_amd64.deb
# !sudo apt-get update
# !wget http://developer.download.nvidia.com/compute/machine-learning/repos/ubuntu1804/x86_64/nvidia-machine-learning-repo-ubuntu1804_1.0.0-1_amd64.deb
# !sudo apt install ./nvidia-machine-learning-repo-ubuntu1804_1.0.0-1_amd64.deb
# !sudo apt-get update
# + id="ldQBa9aqWeG8" colab_type="code" colab={}
# !sudo update-alternatives --remove-all gcc
# !sudo update-alternatives --remove-all g++
# + id="fKVWN6X_UQDu" colab_type="code" colab={}
# !sudo apt-get install --no-install-recommends nvidia-driver-430
# + id="NDmIGqp7UY2l" colab_type="code" colab={}
# !sudo apt-get install --no-install-recommends \
# cuda-10-1 \
# libcudnn7=7.6.4.38-1+cuda10.1 \
# libcudnn7-dev=7.6.4.38-1+cuda10.1
# + id="nDqteJAQVOLJ" colab_type="code" colab={}
# !sudo apt install g++-5
# !sudo apt install gcc-5
# + id="mCu7ZaAyg5Cp" colab_type="code" colab={}
# !sudo update-alternatives --install /usr/bin/gcc gcc /usr/bin/gcc-5 10
# !sudo update-alternatives --install /usr/bin/gcc gcc /usr/bin/gcc-5 20
# !sudo update-alternatives --install /usr/bin/g++ g++ /usr/bin/g++-5 10
# !sudo update-alternatives --install /usr/bin/g++ g++ /usr/bin/g++-5 20
# !sudo update-alternatives --install /usr/bin/cc cc /usr/bin/gcc 30
# !sudo update-alternatives --set cc /usr/bin/gcc
# !sudo update-alternatives --install /usr/bin/c++ c++ /usr/bin/g++ 30
# !sudo update-alternatives --set c++ /usr/bin/g++
# + id="bDH0P32JkjcX" colab_type="code" colab={}
# !pip install torchvision==0.2.1 --no-deps --no-cache-dir
# + id="OyayUvx067yr" colab_type="code" colab={}
# !conda install pytorch==0.4.1 torchvision==0.2.1 -c pytorch
# + id="sS1aZHXKPuko" colab_type="code" colab={}
# !conda install torchvision -c pytorch -c conda-forge
# + id="tFHIYmFxh0hk" colab_type="code" colab={}
# !conda list
| pvnet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <p class='cooltitle' style="font-size:35px; text-align:center;" >Enzyme Kinetics</p>
#
# <br><br>
#
# In this notebook, we're going to implement some basic enzyme kinetics notions in Python, enzyme kinetics play a major role in Neuroscience as they dictate how ion channels and molecular interaction networks in intracellular signaling are regulated. We're going to use differential equations to express those kinetics because they are considered as the mathematical objects of scientific modeling.
# + [markdown] toc=true
# <h1>Table of contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Zero-and-First-Order-Reactions" data-toc-modified-id="Zero-and-First-Order-Reactions-1"><span class="toc-item-num">1 </span>Zero and First Order Reactions</a></span></li><li><span><a href="#Enzymatic-Equilibrium" data-toc-modified-id="Enzymatic-Equilibrium-2"><span class="toc-item-num">2 </span>Enzymatic Equilibrium</a></span></li><li><span><a href="#Michaelis-Menten-Henri-Equation" data-toc-modified-id="Michaelis-Menten-Henri-Equation-3"><span class="toc-item-num">3 </span>Michaelis-Menten-Henri Equation</a></span></li></ul></div>
# -
# Let's start by importing some libraries that we'll use mainly here and in the rest of the Notebooks.
# + hide_input=false
import numpy as np # A Python package for scientific computing
import sympy as sp # A Python library for symbolic mathematics
import matplotlib.pyplot as plt # A data visualization Library in Python
# -
# # Zero and First Order Reactions
# <br>
#
#
# - Considering a chemical reaction catalyzed by an enzyme for a reactant $A$ that transforms into a product $B$ :
#
# $$A \xrightarrow{enzyme} B$$
#
# > If the concentration of the enzyme is far less than the concentration of $A$, we can say that the enzyme is saturated and the depletion of the reactant $A$ or the appearance of the product $B$ is constant and is independant from the concentration of the reactant $A$.
# $$\frac{d[B]}{dt} = - \frac{d[A]}{dt} = k $$
# $k$ being the rate constant of the reaction
#
# - We can see that this expression is equivalent to :
# $$\boxed{- \frac{d[A]}{dt} = k[A]^0 = k}$$
# >The rate of the reaction $\frac{d[A]}{dt}$ does not depend on the concentration of reactants,but only on the rate constant $k$, in what we call a zero-order chemical reaction. This is a differential equation, solving it consists of finding the function that gives the conentration of the reactant in function of time $A(t)$, it is easy to solve at hand but sometimes differential equations get complicated really fast, this is why we use computers.
#
# - If the rate depends linearly upon the concentration of the reactant $A$, the equation becomes :
# $$\boxed{- \frac{d[A]}{dt} = k[A]}$$
# > The first order reaction, this too is considered as a differential equation and can be solved the same way as the previous one.
#
#
# - In order to solve differential equations using python, we can use two different approaches : <br>
# >1 - Analytical methods; by using Python's Sympy library for symbolic mathematics.<br>
# 2 - Numerical Methods; by implementing Euler's method which is used for solving ordinary differential equations (ODEs) and then we can use Python's NumPy Library and its useful N-dimensional array objects that are the cosidered the basis of scientific computing.
# <font size="+2"><b>Zero-order</b></font> <br> <br>
# **Analytical Method**
#
# > Let's start first by initialising Sympy's symbol objects which we will be able to manipulate
# + cell_style="center"
k, t, C1 = sp.symbols(
'k t C1') # Rate constant k , time t and arbitrary integration constant C1
# -
# > The reactant concentration $A$ should be initialised as Sympy Function Object and it should be in the differentiated form with respect to time $t$
# + cell_style="center"
A = sp.Function('A')
dAdt = A(t).diff(t)
# -
# > Now we can write the zero order equation
# + cell_style="center"
zero_order = sp.Eq(-dAdt, k)
zero_order
# -
# > The next step is to solve this differential equation, this can be easily done by using SymPy's function dsolve that takes the equation and the variable to be solved to as parameters.
# + cell_style="center"
analytic_gen_sol_zero = sp.dsolve(zero_order, A(t))
analytic_gen_sol_zero
# -
# > This is the general solution to our differential equation, in order to calculate the arbitrary constant C1 we can initialize $t=0$ in our general solution to find a particular one.
# + cell_style="center"
C1_zero = analytic_gen_sol_zero.subs(
t, 0) # This substitutes t with the value of 0 in our general solution
C1_zero
# -
# > The arbitrary constant $C_1$ is the concentration of the reactant at time 0 (the initial condition)
# + cell_style="center"
analytic_gen_sol_zero = analytic_gen_sol_zero.subs(C1, A(0))
analytic_gen_sol_zero
# -
# > This is the formula for the general solution of the differential equation, in order to find a particular solution let's consider that the rate constant $k=2$ and the initial concentration of the reactant $A(0) = 10$
analytic_par_sol_zero = analytic_gen_sol_zero.subs({A(0): 10, k: 2})
analytic_par_sol_zero
# > Since this is an algebraic SymPy expression, we cannot numerically evaluate it, we have to transform it into a lambda function, we can use SymPy's lambdify function to achieve that.
analytic_sol_zero = sp.lambdify(
t, analytic_par_sol_zero.rhs,
"numpy") # We took the right hand side with .rhs method from our solution
# **Numerical Method**
#
# > In order to numerically solve differential equations, we have to implement the Euler's method, let's proceed by writing a python function that will work with every differential equation. *For a detailed explanation of Euler's Method, check out Steven Strogatz's Nonlinear dynamics and Chaos*
def euler(init_cond, *constants, equation, dt=0.01, Tmax=10):
"""Euler's Method for solving ODEs.
init_cond : The initial condition to start solving from.
constants : a variable length argument with constants in the ODE
equation : the differential equation to be solved.
dt : the time interval between every step.
Tmax : the maximal amount of time
"""
for step in range(int(Tmax / dt)): # How many steps to take
init_cond += dt * equation(init_cond, *constants) # Euler's method
yield init_cond # A generator python expression
# > This Euler function with only 4 lines will take any kind of differential equation and will spit out a generator object that is the solution for a certain array of time. Now we can write the zero-order differential equation as a Lambda function in Python and pass it to our Euler function.
zero_order = lambda c,k : -k
# > Likewise, Let's take $A(0) = 10$ as our initial used concentration (starting condition) and $k=2$
Ao, const = 10, 2
# > So the numerical solution is :
numeric_sol_zero = list(
euler(Ao, const, equation=zero_order
)) # Transform generator into list to visualize with matplotlib
# **Plotting the solutions** <br>
# Now that we have both the solution, let's visualize them.
def plot_solutions(t, numeric, analytic, title):
"""This function plots the numerical and the analytical solutions
to differential equation for a given time vector t
"""
fig, ax = plt.subplots(1, 2, figsize=(10, 4), dpi=150)
plt.subplot(121) # Numerical Solution Plot
plt.plot(t, numeric, 'b', label='$A(t)$')
plt.title('Numerical Solution')
plt.legend()
plt.subplot(122) # Analytical Solution Plot
plt.plot(t, analytic(time), 'g', label='$A(t)$')
plt.xlabel('Time', position=(-0.1, 0))
plt.title('Analytical Solution')
plt.suptitle(title)
plt.legend()
time = np.arange(0, 10, 0.01) # Same vector used for euler's method
plot_solutions(time, numeric_sol_zero, analytic_sol_zero, "Zero-order reaction $A(0)=10$")
# <font size="+2"><b>First-order</b></font> <br> <br>
# **Analytical Method**<br>
# > We're going to use the same steps taken for zero order
# + hide_input=false
first_order = sp.Eq(-dAdt, k*A(t))
first_order
# -
# > After the first order equation has been initialised, let's find the general solution with SymPy
analytic_gen_sol_first = sp.dsolve(first_order, A(t))
analytic_gen_sol_first
# > As always, we'll take $t=0$ to find the arbitrary constant
C1_1 = analytic_gen_sol_first.subs(t,0)
C1_1
# > So the solution is :
analytic_gen_sol_first = analytic_gen_sol_first.subs(C1, A(0))
analytic_gen_sol_first
# > Now to find a particular solution, let's take $A(0) = 10$ and $k=2$
analytic_par_sol_first = analytic_gen_sol_first.subs({A(0) : 10, k : 2})
analytic_par_sol_first
# > And finally we transform it into a lambda function :
analytic_sol_first = sp.lambdify(t, analytic_par_sol_first.rhs, "numpy")
# **Numerical Method**
# > Let's initialize a Python function for the First order reaction
first_order = lambda c,k : -k*c
# > And the solution, using Euler's Method and with the same starting condition and rate constant, will be :
numeric_sol_first = list(euler(Ao, const, equation=first_order))
# **Plotting the solutions** <br>
# And now let's visulaize the solutions
plot_solutions(
time, numeric_sol_first, analytic_sol_first,
'First-order reaction $A(0)=10$') #Same time vector used earlier
# <hr class="sep">
# # Enzymatic Equilibrium
#
# - Let's consider now a molecule that passes from a chemical conformation to another one in a reversible manner, like an ion channel that opens and closes, this reaction will be :
#
# $$A \overset{\alpha}{\underset{\beta}\rightleftharpoons} B$$
#
# - $[A] + [B] = c_0$ is always constant, and taking into account the law of conservation of mass we can see that :
# $$\frac{d[B]}{dt} = \alpha[A] - \beta[B]$$
#
# <br>
#
# - If we take into account the fraction of each conformation where $f_\alpha + f_\beta = 1$ : <br>
# $$\frac{df_\beta}{dt} = \alpha f_\alpha - \beta f_\beta$$
#
# - Considering $f_\alpha = 1 - f_\beta$, we get :
# $$\frac{df}{dt} = \alpha(1 - f) - \beta f = \alpha - (\alpha + \beta)f$$
#
# <br>
#
#
# - Finally, consider $\tau = \frac{1}{\alpha+\beta}$ and $f_\infty = \frac{\alpha}{\alpha+\beta}$ :
#
# $$\boxed{\tau\frac{df}{dt} = f_\infty - f(t)}$$
#
# > This too is a differential equation, now we're going to solve it similairly to zero and first order reactions.
# **Analytical Method**
# > We're going to use SymPy's algebraic notation, let's initialize our constants and functions.
tau, f_infty= sp.symbols('tau f_\infty')
f = sp.Function('f')
dfdt = f(t).diff(t)
# >Let's see our equation :
equilibrium = sp.Eq(tau*dfdt, f_infty - f(t))
equilibrium
# > Now it's time to see the general solution :
analytic_gen_sol_eq = sp.dsolve(equilibrium, f(t))
analytic_gen_sol_eq
# > Let's find the arbitrary constant $C_1$
eq_0 = analytic_gen_sol_eq.subs(t,0) # Considering that t=0
C1_eq = sp.Eq(sp.solve(eq_0,C1)[0], C1) # Solving the equation t=0 to find C1
C1_eq
# > And the solution is :
analytic_gen_sol_eq = analytic_gen_sol_eq.subs(C1,C1_eq.lhs)
analytic_gen_sol_eq
# > It would be interesting to visualize the solution $f(t)$, so let's assign some values for our constants
analytic_par_sol_eq = analytic_gen_sol_eq.subs({f_infty : 6, tau : 2, f(0) : 0})
analytic_par_sol_eq
# > And the final step is always to transform the SymPy expression into a lambda function :
analytic_sol_eq = sp.lambdify(t, analytic_par_sol_eq.rhs, "numpy")
# **Numerical Method**
#
# > We should first create a python function (anonymous function) for our equilibrium reaction :
eq = lambda f,f_oo,tau : (f_oo - f)/ tau
# > Let's assign some values for our constants
f_const, finfty_const, tau_const = 0, 6, 2
# > And the numerical solution using Euler's method will be :
numeric_sol_eq = list(euler(f_const, finfty_const, tau_const, equation=eq))
# **Plotting the solutions**
plot_solutions(time, numeric_sol_eq, analytic_sol_eq,
'Chemical equilibrium $f(0) = 0$')
# <hr class="sep">
# # Michaelis-Menten-Henri Equation
#
#
#
# - Considering the following reaction :
# $$ S + E \overset{k_1}{\underset{k_{-1}}\rightleftharpoons} ES \overset{k_2}{\rightarrow} P + E$$
#
# > Whereas S : Substrate, E : Enzyme, ES : Enzyme-Substrate Complex, P : Product <br>
# **The Michaelis-Menten equation** is :
#
# $$\boxed{ v = \frac{d[P]}{dt} = \frac{V_{max}[S]}{K_m + [S]}}$$
# > Where $V_{max} = k_2[E]$ and $K_m$ is the Michaelis constant, it is the concentration of the substrate $[S]$ when the initial rate $v$ is equal to $\frac{V_{max}}{2}$
#
#
# - If we wanted to graphically determine the constants $K_m$ and $V_{max}$, we're going to use **the Lineweaver-Burk representation** of this equation which we obtain by inversing the earlier equation :
#
# $$ \boxed{\frac{1}{v} = \frac{K_m}{V_{max}}\frac{1}{[S]} + \frac{1}{V_{max}}}$$
#
# > Let's first start by implementing our two equations :
# +
def mich_ment(substrate, vmax, km) :
"""This function returns the rate of the reaction v
from the Michaelis-Menten equation.
Substrate : an array of substrate concentrations
vmax, km : constants of the MM equation, type int"""
return (vmax*substrate)/(km + substrate)
def line_burk(substrate, vmax, km) :
"""The Lineweaver-Burk representation, it sends back 1/v
Substrate : an array of substrate concentrations
vmax, km : constants of the MM equation, type int"""
return (km/(vmax*substrate)) + (1/vmax)
# -
# > Now let's take a look at how they are grapically represented :
# +
s = np.arange(0.1,10, 0.1) # Substrate concentrations between 0.1 and 10
fig, ax = plt.subplots(1, 2, figsize = (12,5), dpi = 150)
plt.subplot(121) # Michaelis-Menten plot
plt.plot(s, mich_ment(s,vmax = 6, km= 1)) # v-s curve
plt.axhline(y = 6, linestyle = '--', color = 'y', label = '$V_{max}$') # Vmax
plt.axvline(x = 1, linestyle = '--', color = 'r')
plt.plot(1,3, 'og' , label = '$K_m = 50 \% V{max}$') # Km
plt.title('The Michaelis-Menten equation')
plt.xlabel(r'$[S]$')
plt.ylabel(r'$v$', rotation = 0)
plt.legend()
plt.subplot(122) # Lineweaver-Burk plot
plt.plot(1/s, line_burk(s,vmax = 6, km= 1)) # 1/v - 1/s curve
plt.title('The Lineweaver-Burk representation')
plt.axvline(x = 1, linestyle = '--', color = 'r')
plt.plot(1,1/3, 'og' , label = r'$\frac{1}{K_m}$') # 1/Km, 1/Vmax = 1/3
plt.xlabel(r'$\frac{1}{[S]}$')
plt.ylabel(r'$\frac{1}{v}$', rotation = 0)
plt.legend()
# -
# <hr class="sep">
| Notebooks/1_Enzyme_Kinetics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Correlation between Detected Breeding Sites and Larval Survey V2
# +
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
from plotly import tools
from plotly.graph_objs import *
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
init_notebook_mode(connected=True)
import plotly.graph_objs as go
import matplotlib as mpl
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
import os, json, itertools
from tqdm import tqdm
from sklearn import model_selection
from sklearn import metrics
from sklearn import preprocessing
from scipy.stats import pearsonr
from sklearn import linear_model
from copy import deepcopy
from scipy.stats.stats import pearsonr, spearmanr
from shapely.geometry import Polygon
from collections import Counter
sns.set(color_codes=True)
month = ['Jan','Feb','Mar','Apr','May','Jun','Jul','Aug','Sep','Oct','Nov','Dec']
categories = np.array(['bin','bowl','bucket','cup','jar','pottedplant','tire','vase']).reshape(-1,1)
data_dir = os.path.join('..','..','data')
# -
# ## 0. Helper Functions
# ### 0.2 Select from Data
# +
def get_detect(df_detect, addrcode, columns=None):
if columns is None:
detect = round(df_detect.loc[df_detect['addrcode'] == addrcode].mean()['total'], 2)
# cup =round(df_detect.loc[df_detect['addrcode'] == addrcode].mean()['cup'], 2)
# vase = round(df_detect.loc[df_detect['addrcode'] == addrcode].mean()['vase'], 2)
# detect = detect-cup-vase
else:
# Breeding Site Feature
detect = df_detect.loc[df_detect['addrcode'] == addrcode][columns].copy()
return detect
def get_survey(df_survey, dengue_season, addrcode):
if dengue_season:
months = [6,7,8,9,10,11]
title = 'Dengue Season'
else:
months = [1,2,3,4,5,6,7,8,9,10,11,12]
title = 'Entire Year'
survey = round(df_survey.loc[
(df_survey['addrcode'] == addrcode) &
(df_survey.index.month.isin(months))
]['bi'].mean(), 2)
return survey, title
def get_cases(df_cases, dengue_season, addrcode):
if dengue_season:
months = [6,7,8,9,10,11]
title = 'Dengue Season'
else:
months = [1,2,3,4,5,6,7,8,9,10,11,12]
title = 'Entire Year'
cases = round(df_cases.loc[
(df_cases['addrcode'] == addrcode) &
(df_cases.index.month.isin(months))
]['cases'].sum(), 2)
return cases, title
def get_area(df_area, addrcode):
area = round(df_area.loc[df_area['addrcode'] == addrcode]['area'].mean(), 2)
return area
def get_population(df_population, addrcode):
population = round(df_population.loc[df_population['addrcode'] == addrcode]['population'].mean(), 2)
return population
def get_gsv_month(df_detect, addrcode):
month = df_detect.loc[df_detect['addrcode'] == addrcode].index.month[0]
return month
def get_gsv_coverage(df_gsv_coverage, addrcode):
coverage = df_gsv_coverage.loc[df_gsv_coverage['addrcode'] == addrcode]['image_area'].mean()
return coverage
def get_dict_info(df_dictionary, addrcode):
dict_info = df_dictionary.loc[df_dictionary['addrcode'] == addrcode]
return dict_info
# -
# ## 1. Load Data
# +
# Load Surveys Data
def filter_survey(df_survey, index='bi'):
df_filtered = []
for addrcode in df_survey['addrcode'].unique():
tmp = df_survey.loc[df_survey['addrcode'] == addrcode].copy()
if len(tmp) == 1 and tmp[index].mean() < 100:
df_filtered.append(tmp.copy())
df_filtered.append(tmp[np.abs(tmp[index]-tmp[index].mean()) <= (1*tmp[index].std())].copy())
df_filtered = pd.concat(df_filtered, axis=0)
return df_filtered
df_survey = pd.read_csv(os.path.join(data_dir,'breeding-sites','csv','addrcode-index','larval-survey.csv'))
df_survey = df_survey.replace(0, np.nan)
df_survey = df_survey.dropna(axis=0, how='any')
df_survey = df_survey.reset_index(drop=True)
df_survey['addrcode'] = df_survey['addrcode'].astype('int')
df_survey['date'] = pd.to_datetime(df_survey['date'], format='%Y-%m')
df_survey = df_survey.set_index('date')
df_survey = df_survey.sort_index()
df_survey = df_survey['2015':'2017']
df_survey = filter_survey(df_survey, index='bi')
df_survey.head(1)
# print('Total data points:',len(df_survey))
# Load Detected Breeding Sites
df_detect = pd.read_csv(os.path.join(data_dir,'breeding-sites','csv','addrcode-index','detection.csv'))
df_detect['date'] = pd.to_datetime(df_detect['date'], format='%Y-%m')
df_detect['addrcode'] = df_detect['addrcode'].astype('int')
df_detect = df_detect.set_index('date')
df_detect = df_detect.sort_index()
df_detect.head(1)
# print('Total data points:',len(df_detect))
# Load Dengue Cases
df_cases = pd.read_csv(os.path.join(data_dir,'dengue-cases','dengue-cases.csv'))
df_cases['date'] = pd.to_datetime(df_cases['date'], format='%Y-%m')
df_cases['addrcode'] = df_cases['addrcode'].astype('int')
df_cases = df_cases.set_index('date')
df_cases = df_cases.sort_index()
df_cases = df_cases['2015':'2017']
df_cases.head(1)
# Area
df_area = pd.read_csv(os.path.join(data_dir,'shapefiles','csv','subdistricts-area.csv'))
df_area['addrcode'] = df_area['addrcode'].astype('int')
df_area.head(1)
# print('Total data points:',len(df_area))
# Population
df_population = pd.read_csv(os.path.join(data_dir,'population','addrcode-index','population.csv'))
df_population['addrcode'] = df_population['addrcode'].astype('int')
df_population.head(1)
# print('Total data points:',len(df_population))
# Dictionary File
df_dictionary = pd.read_csv(os.path.join(data_dir,'shapefiles','csv','addrcode-dictionary.csv'))
df_dictionary['addrcode'] = df_dictionary['addrcode'].astype('int')
df_dictionary.head(1)
# GSV Coverage
df_gsv_coverage = pd.read_csv(os.path.join(data_dir,'shapefiles','csv','gsv-coverage.csv'))
df_gsv_coverage['addrcode'] = df_gsv_coverage['addrcode'].astype('int')
df_gsv_coverage.head(1)
# -
# ## 2. Correlation between Total Breeeding Site Counts and Cases
def correlation_total_bs(df_survey, df_detect, df_area, df_cases, df_population, column='total'):
titles = []
traces = []
for dengue_season, province in list(itertools.product([False, True], [['10',],['80'],['81']])):
X, y = [], []
names = []
for addrcode in df_detect['addrcode'].unique():
province_id = str(addrcode)[:2]
if province_id not in province: continue
detect = get_detect(df_detect, addrcode)
cases, title = get_cases(df_cases, dengue_season, addrcode)
area = get_area(df_area, addrcode)
population = get_population(df_population, addrcode)
if np.isnan(detect) or np.isnan(cases) or np.isnan(population): continue
X.append(cases)
y.append(population)
names.append(addrcode)
province = ', '.join(province)
pearson_val = pearsonr(X, y)
spearman_val = spearmanr(X, y)
trace_1 = go.Scatter(
x=X,
y=y,
mode='markers',
name=province,
text=names,
marker=dict(size=14, opacity=0.5)
)
X, y = np.array(X), np.array(y)
regr = linear_model.LinearRegression()
regr.fit(X.reshape(-1, 1), y.reshape(-1, 1))
y_pred = np.squeeze(regr.predict(X.reshape(-1, 1)))
trace_2 = go.Scatter(
x = X,
y = y_pred,
mode = 'lines',
line = dict(width = 4),
name=province
)
titles.append(
title+' ('+str(len(X))+' data points)'+', province: '+province+ \
'<br>Pearson: '+str(round(pearson_val[0],4))+ ', p-value: '+str(round(pearson_val[1],4))+ \
'<br>Spearman: '+str(round(spearman_val[0],4))+', p-value: '+str(round(spearman_val[1],4))
)
traces.append([trace_1, trace_2])
fig = tools.make_subplots(rows=2, cols=3, subplot_titles=tuple(titles), horizontal_spacing = 0.05, vertical_spacing=0.15)
k = 0
for i in range(2):
for j in range(3):
fig.append_trace(traces[k][0], i+1, j+1)
fig.append_trace(traces[k][1], i+1, j+1)
fig['layout']['xaxis'+str(k+1)].update(title='Dengue Cases')
fig['layout']['yaxis'+str(k+1)].update(title='Population')
k+=1
fig['layout'].update(height=1200, hovermode='closest')
iplot(fig)
return X, y
X, y = correlation_total_bs(df_survey, df_detect, df_area, df_cases, df_population)
# ## 2.
# +
def correlation_matrix_plot(dengue_season=False, province=['80'], norm='image_area'):
brd_sites=['bin','bowl','bucket','jar','pottedplant','tire','vase','cup']
X, y, names = [], [], []
for addrcode in df_detect['addrcode'].unique():
province_id = str(addrcode)[:2]
if province_id not in province: continue
cases, title = get_cases(df_cases, dengue_season, addrcode)
population = get_population(df_population, addrcode)
area = get_area(df_area, addrcode)
image_area = get_gsv_coverage(df_gsv_coverage, addrcode)
detect = np.squeeze(get_detect(df_detect, addrcode, brd_sites).values)
detect = norm_detection(norm, detect, area, image_area, population)
if np.isnan(cases) or np.isnan(population): continue
# Combine Features
X.append(list(detect) + [cases])
names.append(addrcode)
X = np.array(X)
df_features = pd.DataFrame.from_records(X, columns=['bin','bowl','bucket','jar','pottedplant',
'tire','vase','cup','cases'])
# plt.style.use('seaborn-dark-palette')
# plt.figure(figsize=(16,11))
# sns.set(font_scale=1.3)
# sns.heatmap(df_features.corr(), annot=True, fmt=".2f", cmap="YlGnBu")
# plt.title('Dengue season: '+str(dengue_season) + ', Norm: '+norm + \
# ',\n Province: '+''.join(province)+', Data shape: '+str(X.shape))
# plt.show()
df_features['addrcode'] = names
df_features.set_index('addrcode', inplace=True)
return df_features
def norm_detection(norm, detect, area, image_area, population):
if norm == 'image_area':
detect = detect/image_area
elif norm == 'land_area':
detect = detect/area
elif norm == 'population/image_area':
detect = (detect*population)/(image_area)
elif norm == 'population/land_area':
detect = (population*detect)/(area)
elif norm == 'population':
detect = detect/population
return detect
# -
# ## 3. Correlation between *Predicted* Breeding Site Counts and Breteau Index
# +
df = pd.read_csv('/home/poom/Desktop/finaldata.csv')
for x in df.addrcode.values:
if x not in df_features.index:
print(x)
for x in df_features.index:
if x not in df.addrcode.values:
print(x)
# +
df_features = correlation_matrix_plot(
dengue_season=False,
province=['10','80','81'],
norm=''
)
X = df_features.drop('cases', axis=1).values
y = df_features['cases'].values
X = df_features.loc[df.addrcode].drop('cases', axis=1).values
y = df_features.loc[df.addrcode]['cases'].values
X.shape, y.shape
predicted = model_selection.cross_val_predict(linear_model.LinearRegression(), X, y, cv=10)
_=plt.title('R-squared:' + str(round(metrics.r2_score(y, predicted),4)) + ', Pearson: ' + str(np.round(pearsonr(y, predicted),4)[0]))
_=sns.set()
_=sns.regplot(y, predicted)
# -
df_features.loc[df.addrcode].to_csv('/home/poom/Desktop/norm_features.csv')
| src/correlation/correlation-dengue-case-v2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pRH(3.6)
# language: python
# name: prh
# ---
# # Camera Calibration Demo
# > A little demonstration of camera calibration process.
#
# - toc: true
# - branch: master
# - badges: true
# - comments: true
# - categories: [jupyter, camera calibration, python, computer vision]
# - hide: false
# - search_exclude: false
# ## Brief Introduction
# In camera calibration problem, we are given a calibration pattern, which is an object with features of known size or geometry, and we want to find the intrinsic (and or extrinsic) parameters of a camera. A calibration pattern could be a chessboard pattern or rounded dots.
#
# ### Intrinsic Parameters
# $$\begin{bmatrix} x\\ y\\ 1 \end{bmatrix} = \begin{bmatrix} \frac{f}{p_x}&(tan\,\alpha)\frac{f}{p_y}&c_x\\ &\frac{f}{p_y}&c_y\\ &&1 \end{bmatrix} \begin{bmatrix} x_{\mathcal R}\\ y_{\mathcal R}\\ 1 \end{bmatrix}$$
# * $\begin{bmatrix} x\\ y\\ 1 \end{bmatrix}$: image coordinates
# * $\begin{bmatrix} x_{\mathcal R}\\ y_{\mathcal R}\\ 1 \end{bmatrix}$: real world coordinates
# * $f$: focal length
# * $p_x, p_y$: width and height of pixels
# * $\begin{bmatrix} c_x\\ c_y \end{bmatrix}$: principal point
# * $\alpha$: skew angle
#
# The equation can be simplified as below
#
# $$\begin{bmatrix} x\\ y\\ 1 \end{bmatrix} = \begin{bmatrix} f_x&s&c_x\\ &f_y&c_y\\ &&1 \end{bmatrix} \begin{bmatrix} x_{\mathcal R}\\ y_{\mathcal R}\\ 1 \end{bmatrix}$$
#
# This is a common camera matrix. It can be further simplified to
#
# $$m = K\,m_{\mathcal R}$$
#
# ### Extrinsic Parameters
# $$M^\prime = \begin{bmatrix} R&t\\ 0_3^T&1 \end{bmatrix}\,M$$
# * $R$: Rotation matrix ($3 \times 3$ matrix)
# * $t$: translation vector($\begin{bmatrix}t_x\\t_y\\t_z \end{bmatrix}$)
# * 3 for $R$ and 3 for $t \to$ total 6 degree of freedom
#
# ### Combined Projection Matrix
# $$\begin{bmatrix} x\\y\\1 \end{bmatrix} \sim \begin{bmatrix} f_x&s&c_x\\ 0&f_y&c_y\\ 0&0&1 \end{bmatrix} \, \begin{bmatrix} 1&0&0&0\\ 0&1&0&0\\ 0&0&1&0 \end{bmatrix} \, \begin{bmatrix} R&t\\ 0_3^T&1 \end{bmatrix} \, \begin{bmatrix} X\\Y\\Z\\1\end{bmatrix}$$
# * [image] = [Intrinsic K] [projection] [extrinsic] [real world]
# * $m \sim K \, [R|t] \, M$
# * $m \sim P \; M$
# * $P$: $3 \times 4$ matrix, camera projection matrix
#
# ### Camera Calibration
# Camera calibration is composed of two major steps as below.
# 1. Compute the matrix $P$ from a set of corresponding points of image and real world coordinates.
# 2. Decompose $P$ into $K, R, t$ with QR decomposition
import os
import cv2
import numpy as np
import matplotlib.pyplot as plt
def imageshow(img, cmap='bgr', savefile=False, filename='output'):
"""
Print image or Save it.
img: Image to show or to save.
cmap: Color map of img. Default to 'bgr'
savefile: To save image or not. Default to 'False'.
filename: The filename you want the output image has.
Default to 'output.jpg'
"""
cwd = os.getcwd() + '/'
des = cwd + filename + '.jpg'
if savefile:
cv2.imwrite(des, img)
else:
plt.figure(figsize=(8,8))
if(cmap == 'bgr'):
# OpenCV considers float only when values range
# from 0-1.
img = img.astype('uint8')
# Change order from bgr to rgb for plt to show
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
plt.imshow(img)
else:
plt.imshow(img, cmap=cmap)
plt.title(filename)
plt.show()
# ## Load Image
# Open the image
img_name_1 = 'data/chessboard_1.jpg'
img_name_2 = 'data/chessboard_2.jpg'
img_1 = cv2.imread(img_name_1)
img_2 = cv2.imread(img_name_2)
imageshow(img_1, filename="chessboard_1")
imageshow(img_2, filename="chessboard_2")
# ## Load Image and Real World Coordinate
# Image coordinates can be obtained by manual labelling or corner detection. However, coordinates from solely corner detection are not so accurate, we can use two straight lines to help labelling the coordinates.
#
# In this demo, we label the image coordinates manually.
# +
#collapse-output
# Open the corner label file
corner_label_1 = np.load('Point2D_1.npy')
corner_label_2 = np.load('Point2D_2.npy')
print(f"Coordinates for chessboard_1:({len(corner_label_1)} points in total)\n", corner_label_1)
print(f"Coordinates for chessboard_2:({len(corner_label_2)} points in total)\n", corner_label_2)
# Open the 3D points file
threeD_point = np.loadtxt('data/Point3D.txt')
print(f"Coordinates for chessboard in real world:({len(threeD_point)} points in total)\n", threeD_point)
# -
# ## Homogeneous Coordinates
# We borrow a coordinates for manually labelled coordinates and real world coordinates.
# +
#collapse-output
# Make homogeneous coordinates of corner label and 3D points
tmp = np.ones((corner_label_1.shape[0], 1))
corner_label_1_homo = np.concatenate((corner_label_1, tmp), axis = 1)
tmp = np.ones((corner_label_2.shape[0], 1))
corner_label_2_homo = np.concatenate((corner_label_2, tmp), axis = 1)
tmp = np.ones((threeD_point.shape[0], 1))
threeD_point_homo = np.concatenate((threeD_point, tmp), axis = 1)
print(f"Coordinates for chessboard_1:({len(corner_label_1_homo)} points in total)\n", corner_label_1_homo)
print(f"Coordinates for chessboard_2:({len(corner_label_2_homo)} points in total)\n", corner_label_2_homo)
print(f"Coordinates for chessboard in real world:({len(threeD_point_homo)} points in total)\n", threeD_point_homo)
# -
# ## Step 1. Compute Matrix $P$
# We can use corresponding points between image and real world coordinates to get compute matrix $P$, since each correspondence generates two equations as below.
# $$x_i = \frac{p_{11}X_i+p_{12}Y_i+p_{13}Z_i+p_{14}}{p_{31}X_i+p_{32}Y_i+p_{33}Z_i+p_{34}}, \, y_i = \frac{p_{21}X_i+p_{22}Y_i+p_{23}Z_i+p_{24}}{p_{31}X_i+p_{32}Y_i+p_{33}Z_i+p_{34}}$$
#
# They can be written as
# $$\begin{aligned}x_i(p_{31}X_i+p_{32}Y_i+p_{33}Z_i+p_{34}) &= p_{11}X_i+p_{12}Y_i+p_{13}Z_i+p_{14}\\
# y_i(p_{31}X_i+p_{32}Y_i+p_{33}Z_i+p_{34}) &= p_{21}X_i+p_{22}Y_i+p_{23}Z_i+p_{24}
# \end{aligned}$$
#
# They can also be written as
# $$\begin{bmatrix} X_i&Y_i&Z_i&1&0&0&0&0&-x_iX_i&-x_iY_i&-x_iZ_i&-x_i\\ 0&0&0&0&X_i&Y_i&Z_i&1&-y_iX_i&-y_iY_i&-y_iZ_i&-y_i\end{bmatrix}\,P = 0$$
# * $P$ is a 12-vector
#
# We can compute matrix $P$ by concatenate $n$ correspondences to generate matrix $A$, which is a $2n \times 12$ matrix as below
# $$A = \begin{bmatrix} X_1&Y_1&Z_1&1&0&0&0&0&-x_1X_1&-x_1Y_1&-x_1Z_1&-x_1\\ 0&0&0&0&X_1&Y_1&Z_1&1&-y_1X_1&-y_1Y_1&-y_1Z_1&-y_1\\ &&&&&&&\vdots \\ X_n&Y_n&Z_n&1&0&0&0&0&-x_nX_n&-x_nY_n&-x_nZ_n&-x_n\\ 0&0&0&0&X_n&Y_n&Z_n&1&-y_nX_n&-y_nY_n&-y_nZ_n&-y_n\\\end{bmatrix}$$
# * Theoretically, $n \geq 6$ is enough, but we find $n >> 6$ in practice.
# * In this demo, we use $36$ corresponding points to compute matrix $P$, therefore, we have matrix $A$ in the shape of $72 \times 12$.
#
# ### Numerical Solution
# We can use the eigenvector with the smallest eigenvalue of $A^TA$ as a solution of matrix $P$, which minimizes $||AP||$ and makes $||P|| = 1$.
# Here we return matrix $P$ in the shape of $3 \times 4$.
def getProjectionMatrix(corner_label, threeD_point):
"""
Get the projection matrix from given 2D and 3D points.
Parameters:
* corner_label: 2D points in the image.
* threeD_points: 3D points from the real world.
Return:
* P: Projection matrix.
"""
# Concatenate the equations
# Ai = [[X1, Y1, Z1, 1, 0, 0, 0, 0, −x1X1, −x1Y1, −x1Z1, −x1],
# [0, 0, 0, 0, X1, Y1, Z1, 1, −y1X1, −y1Y1, −y1Z1, −y1],...]
# A is a (2N, 12) matrix, N is the point number
A = np.zeros((2*threeD_point.shape[0], 12))
tmp = [12]
for i in range(threeD_point.shape[0]):
tmp[0:4] = threeD_point[i]
tmp[4:8] = [0.0, 0.0, 0.0, 0.0]
tmp[8:] = -(corner_label[i][0]*threeD_point[i])
A[2*i] = tmp
tmp[0:4] = [0.0, 0.0, 0.0, 0.0]
tmp[4:8] = threeD_point[i]
tmp[8:] = -(corner_label[i][1]*threeD_point[i])
A[2*i+1] = tmp
print(f"matrix A is in the shape of {A.shape[0]} X {A.shape[1]}")
AT_A = np.dot(A.T, A)
eigenValues, eigenVectors = np.linalg.eig(AT_A)
# print(eigenValues)
# print(eigenVectors)
# Find P, which is the eigenvector with the smallest eigenvalue
idx = eigenValues.argsort()[0]
eigenValues = eigenValues[idx]
# The column of the returned eigenvector is real eigen vector.
P = eigenVectors[:, idx]
P = P.reshape((3, 4))
print(f"matrix P is in the shape of {P.shape[0]} X {P.shape[1]}")
print(f"Test ||P|| (sqrt(sum(P^2))): {np.sqrt(np.sum(P**2))}")
return P
### Calculate projection matrix, P ###
P_1 = getProjectionMatrix(corner_label_1_homo, threeD_point_homo)
print("-"*50)
P_2 = getProjectionMatrix(corner_label_2_homo, threeD_point_homo)
# ## Step 2. Decomposing $P$
# In this step, we want to get intrinsic and extrinsic parameters out of matrix $P$.
#
# We first extract a $3 \times 3$ submatrix $M$ out of matrix $P$.
# $$M = K\,R$$
# Matrix $M$ is the product of an upper triangular ($K = \begin{bmatrix} f_x&s&c_x\\ &f_y&c_y\\ &&1 \end{bmatrix}\,$) and rotation ($R \to 3 \times 3$) matrix.
#
# Then we decompose matrix $M$ into $K, R$ by QR decomposition.
#
# After that, we obtain $t$ by equation
# $$t = K^{-1}\,\begin{bmatrix}p_{14}\\p_{24}\\p_{34}\end{bmatrix}$$
#
# One should be noticed that the matrix $K$ contains a skew angle parameter $s$, and $s = \tan \theta$. $\theta$ is the angle between the image axes.
def getKRT(P):
"""
Get the intrinsic parameters K, extrinsic parameters R,
and translation vector t.
Parameters:
* P: Projection Matrix
Return:
* K: 3X3 upper triangular matrix.
* R: 3X3 orthogonal matrix.
* t: 3X1 matrix.
"""
# Normalize P with ||P31,P32,P33||
P = P/np.sqrt(P[2][0]**2 + P[2][1]**2 + P[2][2]**2)
# Do RQ decomposition to get intrinsic parameter K,
# and extrinsic parameters R by QR decomposition
M = P[:3, :3]
Q, R = np.linalg.qr(np.linalg.inv(M))
K = np.linalg.inv(R)
R = np.linalg.inv(Q)
# Compute t, translation vector
P_T = P[:3, 3].reshape(3, 1)
t = np.dot(np.linalg.inv(K), P_T)
print("K: ")
print(K)
print("R: ")
print(R)
# print(np.dot(R, R.T))
print("t: ")
print(t)
return K, R, t
### Calculate parameters K, R, and t.###
K_1, R_1, t_1 = getKRT(P_1)
print("-"*60)
K_2, R_2, t_2 = getKRT(P_2)
# ## Reprojecting
# Finally, we can reproject the dots from real world coordinates with the $K, R, t$ matrices we have.
# $$\begin{bmatrix} x\\y\\1 \end{bmatrix} \sim K \, \begin{bmatrix} 1&0&0&0\\ 0&1&0&0\\ 0&0&1&0 \end{bmatrix} \, \begin{bmatrix} R&t\\ 0_3^T&1 \end{bmatrix} \, \begin{bmatrix} X\\Y\\Z\\1\end{bmatrix}$$
#
# We can evaluate the reprojection effect by calculating Root Mean Square Error (RMSE) between reprojected points from real world coordinates (marked as pink) and manually labelled image points (marked as yellow).
def ReProject(img, img_name, K, R, t, twoD, threeD):
"""
Reproject and display the 2d points on image by
given K, R, and t.
Parameters:
* img: Image to reproject on.
* img_name: Filename of the image to save.
* K: 3X3 intrinsic parameters.
* R: 3X3 extrinsic parameters rotation matrix.
* t: 3X1 extrinsic parameters translation vector.
* twoD: 2D points that user labeled.
* threeD: 3D points of the 2D points in real world.
"""
# Concatenate the extrinsic matrix
extrinsic = R
extrinsic = np.append(extrinsic, t, axis=1)
a = np.array([0.0, 0.0, 0.0, 1.0]).reshape((1, 4))
extrinsic = np.append(extrinsic, a, axis=0)
projection = np.array([ [1, 0, 0, 0],
[0, 1, 0, 0],
[0, 0, 1, 0]])
P_new = np.dot(np.dot(K, projection), extrinsic)
# print(P_new)
for i in twoD:
x = i[0]
y = i[1]
# circle(image, coordinate, radius, color, thickness)
cv2.circle(img, (x, y), 5, (0, 255, 255), -1) # Yellow
tgs = []
for j in threeD:
tg = np.dot(P_new, j)
tg = tg/tg[2]
x = int(round(tg[0]))
y = int(round(tg[1]))
cv2.circle(img, (x, y), 5, (255, 0, 255), 2) # Pink
tgs += [[x, y]]
tgs = np.asarray(tgs)
rmse = np.sqrt(np.mean((tgs - twoD)**2))
print("Root Mean Square Error(RMSE): %f" % rmse)
# img_name = img_name + '.jpg'
# cv2.imwrite(img_name, img)
# cv2.imshow(img_name, img)
# cv2.waitKey(0)
imageshow(img, filename=img_name)
### Reproject 2D points by K, R, T ###
ReProject(img_1, 'chessboard_1r', K_1, R_1, t_1,
corner_label_1, threeD_point_homo)
ReProject(img_2, 'chessboard_2r', K_2, R_2, t_2,
corner_label_2, threeD_point_homo)
# ## Visualize Camera Position
# The visualizing functions are provided by course TA.
# +
import math
from mpl_toolkits.mplot3d import Axes3D
from mpl_toolkits.mplot3d.art3d import Poly3DCollection
def draw_camera(ax, R, pos, color):
leftTopCorner = [ -0.2, 0.2, -1 ] @ R
leftTopCorner = leftTopCorner / np.linalg.norm(leftTopCorner, 2)*4 + pos
leftBotCorner = [ -0.2, -0.2, -1 ] @ R
leftBotCorner = leftBotCorner / np.linalg.norm(leftBotCorner, 2)*4 + pos
rightTopCorner = [ 0.2, 0.2, -1 ] @ R
rightTopCorner = rightTopCorner / np.linalg.norm(rightTopCorner, 2)*4 + pos
rightBotCorner = [ 0.2, -0.2, -1 ] @ R
rightBotCorner = rightBotCorner / np.linalg.norm(rightBotCorner, 2)*4 + pos
leftTriangle = np.concatenate([pos, leftTopCorner, leftBotCorner], axis=0)
topTriangle = np.concatenate([pos, leftTopCorner, rightTopCorner], axis=0)
rightTriangle = np.concatenate([pos, rightTopCorner, rightBotCorner], axis=0)
botTriangle = np.concatenate([pos, leftBotCorner, rightBotCorner], axis=0)
verts = [list(leftTriangle)]
ax.add_collection3d(Poly3DCollection(verts, facecolors=color))
verts = [list(topTriangle)]
ax.add_collection3d(Poly3DCollection(verts, facecolors=color))
verts = [list(rightTriangle)]
ax.add_collection3d(Poly3DCollection(verts, facecolors=color))
verts = [list(botTriangle)]
ax.add_collection3d(Poly3DCollection(verts, facecolors=color))
def visualize(pts, R1, T1, R2, T2):
'''
Input:
pts: 36x3 3D points
R1: 3x3 rotation matrix of image 1
T1: 3x1 translation vector of image 1
R2: 3x3 roatation matrix of image 2
T2: 3x1 translation vector of image 2
This function will display a chessboard and two cameras.
'''
fig = plt.figure()
ax = Axes3D(fig)
ax.set_xlim3d(-10, 5)
ax.set_ylim3d(10, -5)
ax.set_zlim3d(-5, 10)
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
# pose vector of camera
cameraPoseVector1 = R1.T @ [0, 0, 1]
cameraPoseVector1 /= np.linalg.norm(cameraPoseVector1)
cameraPoseVector2 = R2.T @ [0, 0, 1]
cameraPoseVector2 /= np.linalg.norm(cameraPoseVector2)
angle = math.degrees(math.acos(np.clip(np.dot(cameraPoseVector1, cameraPoseVector2), -1, 1)))
print('Angle between two cameras: ', angle)
# position of camera
cameraPos1 = -T1.T @ R1
cameraPos2 = -T2.T @ R2
# draw
for r in range(1, 9):
for c in range(1, 4):
fourCorner = np.concatenate((pts[r*4+c-5, :], pts[r*4+c-4, :], pts[r*4+c, :], pts[r*4+c-1, :]))
fourCorner.resize((4,3))
if r%2 == c%2:
color = 'black'
else:
color = 'white'
verts = [list(fourCorner)]
ax.add_collection3d(Poly3DCollection(verts, facecolors=color))
# draw cameras
draw_camera(ax, R1, cameraPos1, 'blue')
draw_camera(ax, R2, cameraPos2, 'red')
plt.show()
# -
# %matplotlib notebook
import matplotlib.pyplot as plt
visualize(threeD_point, R_1, t_1, R_2, t_2)
# ## References
# 1. Slides from NTHU Computer Vision lecture (CS6550).
| _notebooks/2022-01-05-Camera-Calibration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## This notebook reads in local xml files in bulk and assembles a dataset which it cleans and reduces and exports as a csv
# +
# imports
import xml.etree.ElementTree as et
import pandas as pd
pd.set_option('max_columns', 100)
import os
import time
import warnings
warnings.simplefilter(action = 'ignore', category = Warning)
# +
# an empty list where we'll put just the dicts we want
selected = []
# keep a count of skipped empty files (0KB) and of ignored DSSTORE or jupyter files
skipped_ctr = 0
ignored_ctr = 0
# create a list of all the local files, the path to them and a file counter to increase
list_of_files = os.listdir('/Users/aidanair/Documents/DATA/V_ENV/virtual_env_tri/SPANISH_ARCHIVE_FILES')
file_ctr = 0
path = '/Users/aidanair/Documents/DATA/V_ENV/virtual_env_tri/SPANISH_ARCHIVE_FILES/'
# read through all the files
for x in range (len(list_of_files)):
# define the file location and name
local_file = path + list_of_files[file_ctr]
# find the file size
filesize = os.path.getsize(local_file)
# if file is empty, skip it
if filesize == 0:
file_ctr += 1
skipped_ctr += 1
continue
# if it's a DSSTORE or NOTEBOOK CHECKPOINT, skip it
other_files = ['/Users/aidanair/Documents/DATA/V_ENV/virtual_env_tri/SPANISH_ARCHIVE_FILES/.DS_Store', '/Users/aidanair/Documents/DATA/V_ENV/virtual_env_tri/SPANISH_ARCHIVE_FILES/.ipynb_checkpoints']
if local_file in other_files:
file_ctr += 1
ignored_ctr += 1
continue
# otherwise:
else:
# create a tree and root to parse the xml
xtree = et.parse(local_file)
xroot = xtree.getroot()
# create empty dict
empty = {}
# define the list of k tags / keys we want in our shorter df
key_list = ['recordid', 'otherrecordid', 'titleproper', 'agencycode', 'agencyname', 'fromdate', 'todate', 'url']
# find all elements then collect the tags as keys, and the text as values
for c in xroot.findall(".//"):
k = c.tag.split('}')[1]
v = c.text
# if the key is in the selected list, add it to our empty dictionary
if k in key_list:
empty[k] = v
# define complete url (for the html page) to which this xml file corresponds and add to dict
ending = local_file.split('_')[-1].split('.')[0]
url = f'http://pares.mcu.es/ParesBusquedas20/catalogo/description/{ending}'
empty['url'] = url
# add our dictionary to our overall list of dicts
selected.append(empty)
# print a counter marker every 5000
if file_ctr % 5000 == 0:
print("file counter currently at:", file_ctr)
# increase counter
file_ctr += 1
# when finished, show the number of dictionaries now in the list (and how many 0KB files or unwanted files were skipped):
print(len(selected))
print(skipped_ctr)
print(ignored_ctr)
# +
# make the df
df = pd.DataFrame(selected)
print(df.shape)
df[:4]
# -
# ## clean the df
# +
# remove 'Ficha de ' from in front of each name
def remove_ficha(titleproper):
splitup = titleproper.split('Ficha de ')
return splitup[1]
df['name'] = df.titleproper.apply(remove_ficha)
# +
# move the id col to the end in order to split it easily
df = df[['name', 'otherrecordid', 'titleproper', 'agencycode', 'agencyname', 'fromdate', 'todate', 'url', 'recordid']]
# +
# check
df[:5]
# +
# split the first part of record id (keeping original) using /
df[['recordid_a', 'recordid_b', 'recordid_x', 'recordid_c']] = df.recordid.str.split("/", expand=True)
# +
# split remainder of record id using ,
df[['recordid_d', 'recordid_e', 'recordid_f', 'recordid_g']] = df.recordid.str.split(",", expand=True)
# +
# check
df[:3]
# +
# reorder the cols, also removing any duplicate or unnecessary cols
df = df[['name', 'recordid_a', 'recordid_b', 'recordid_e', 'recordid_f', 'recordid_g', 'fromdate', 'todate', 'agencycode', 'agencyname', 'recordid', 'otherrecordid', 'url']]
# +
# rename the cols
df.columns = ['name', 'recordid_a', 'recordid_b', 'recordid_c', 'recordid_d', 'recordid_e', 'fromdate', 'todate', 'agencycode', 'agencyname', 'recordid', 'otherrecordid', 'url']
# +
# finished
df[:3]
# +
## check for duplicate rows in the urls column. No url should appear more than once
df.url.value_counts()
# +
# export as a csv, with no index - with first and last file number and date and total length of df
df.to_csv('/Users/aidanair/Desktop/pares_archive_10_074_724_10_117_072_03042021_LEN41442.csv', index = False)
# -
| PARES/PARSE_xml_files.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matplotlib OO API from callbacks
#
# Based on [this example](https://matplotlib.org/gallery/api/agg_oo_sgskip.html). Some notes:
#
# * If you don't use the `pyplot` API, you have to manually attach a canvas. Even then, `fig.show()` still doesn't work because no manager is set.
# * ~~Toggling the image widget's visibility doesn't currently work in voila, so you'll see a broken image icon. You could fix that by starting with a blank image.~~ Looks like hiding the image widget works now!
# * If you use VBox instead of HBox, the image gets stretched horizontally.
import ipywidgets
from matplotlib.backends.backend_agg import FigureCanvasAgg as FigureCanvas
from matplotlib.figure import Figure
# +
def callback(w):
fig = Figure()
FigureCanvas(fig)
ax = fig.add_subplot(111)
ax.plot([1, 2, 3])
fig.savefig('chart.png')
image.value = open('chart.png', 'rb').read()
image.layout = ipywidgets.Layout(visibility='visible')
button = ipywidgets.Button(description='Run')
button.on_click(callback)
image = ipywidgets.Image(layout=ipywidgets.Layout(visibility='hidden'))
ipywidgets.HBox(children=[button, image])
# -
| voila/callback_matplotlib_oo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# +
import numpy as np
import os
import cv2
from tqdm import tqdm
import random
import shutil
import sys
sys.path.append('../inference/')
from face_detector import FaceDetector
# +
IMAGES_DIR = '/home/gpu2/hdd/dan/FDDB/originalPics/'
ANNOTATIONS_PATH = '/home/gpu2/hdd/dan/FDDB/FDDB-folds/'
MODEL_PATH = '../model.pb'
# you will need the content of this folder
# for the official evaluation script
RESULT_DIR = 'result_fddb/'
# -
# # Collect annotated images
annotations = [s for s in os.listdir(ANNOTATIONS_PATH) if s.endswith('ellipseList.txt')]
image_lists = [s for s in os.listdir(ANNOTATIONS_PATH) if not s.endswith('ellipseList.txt')]
annotations = sorted(annotations)
image_lists = sorted(image_lists)
images_to_use = []
for n in image_lists:
with open(os.path.join(ANNOTATIONS_PATH, n)) as f:
images_to_use.extend(f.readlines())
# +
shutil.rmtree(RESULT_DIR, ignore_errors=True)
os.mkdir(RESULT_DIR)
images_to_use = [s.strip() for s in images_to_use]
with open(os.path.join(RESULT_DIR, 'faceList.txt'), 'w') as f:
for p in images_to_use:
f.write(p + '\n')
# -
ellipses = []
for n in annotations:
with open(os.path.join(ANNOTATIONS_PATH, n)) as f:
ellipses.extend(f.readlines())
i = 0
with open(os.path.join(RESULT_DIR, 'ellipseList.txt'), 'w') as f:
for p in ellipses:
# check image order
if 'big/img' in p:
assert images_to_use[i] in p
i += 1
f.write(p)
# # Predict using the trained detector
face_detector = FaceDetector(MODEL_PATH, gpu_memory_fraction=0.5, visible_device_list='1')
predictions = []
for n in tqdm(images_to_use):
image_array = cv2.imread(os.path.join(IMAGES_DIR, n) + '.jpg')
image_array = cv2.cvtColor(image_array, cv2.COLOR_BGR2RGB)
# threshold is important to set low
boxes, scores = face_detector(image_array, score_threshold=0.01)
predictions.append((n, boxes, scores))
with open(os.path.join(RESULT_DIR, 'detections.txt'), 'w') as f:
for n, boxes, scores in predictions:
f.write(n + '\n')
f.write(str(len(boxes)) + '\n')
for b, s in zip(boxes, scores):
ymin, xmin, ymax, xmax = b
h, w = int(ymax - ymin), int(xmax - xmin)
f.write('{0} {1} {2} {3} {4:.4f}\n'.format(int(xmin), int(ymin), w, h, s))
# # Copy images
# for some reason the official evaluation script needs this
for n in tqdm(images_to_use):
p = os.path.join(RESULT_DIR, 'images', n + '.jpg')
os.makedirs(os.path.dirname(p), exist_ok=True)
shutil.copy(os.path.join(IMAGES_DIR, n) + '.jpg', p)
| evaluation/predict_for_FDDB.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/willianrocha/bootcamp-datascience-alura/blob/main/module_1/ds_mod1_lecture5.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="e5M-p5H4qpGO" outputId="d8018d4f-dc48-4eaf-bedd-1f6d5eab7351"
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
import numpy as np
import pandas as pd
from google.colab import drive
drive.mount('/content/drive')
# Mind the path for the csv file in your gdrive
# !cp "/content/drive/MyDrive/alura/bootcamp-data-science/A160324189_28_143_208.csv" .
pd.options.display.float_format = "{:,.2f}".format
dados = pd.read_csv("/content/A160324189_28_143_208.csv", encoding="ISO-8859-1",
skiprows=3, skipfooter=12, sep=";", thousands=".", decimal=",",
engine='python')
# + [markdown] id="0mxUhaXbwA4s"
# Olhar ultimo mês do ordenado por total
# * Slicing
# * Ultimo mês pode ser incompleto, talvez seja melhor usar o penultimo
# * Dados são por data de atendimento, não de processamento
# * Ultimos meses tem uma queda natural em relação ao processamento tardio de internações - Represamento de dados.
# * Encontrar um grafico que represente bem
#
# + id="WebH_nYl5YQ_"
colunas_usaveis = dados.mean().index.tolist()
colunas_usaveis.insert(0, "Unidade da Federação")
usaveis = dados[colunas_usaveis]
usaveis = usaveis.set_index("Unidade da Federação")
# usaveis = usaveis.drop("Total", axis=1)
# usaveis["Total"] = usaveis.sum(axis=1)
ordenado_por_total = usaveis.sort_values("Total", ascending=False)
ordenado_por_total = ordenado_por_total.drop("Total", axis=1)
colunas_interessadas = ordenado_por_total.columns[6:]
ordenado_por_total = ordenado_por_total[colunas_interessadas]
ordenado_por_total = ordenado_por_total / 10**6
# + colab={"base_uri": "https://localhost:8080/"} id="EEaeCxTa5Yof" outputId="d91f4fe0-559b-487f-b71c-0e83e8bb40d1"
mes_mais_recente = ordenado_por_total.columns[-1]
gastos_do_mais_recente = ordenado_por_total[mes_mais_recente]
gastos_do_mais_recente.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="cF47qPhP5cDg" outputId="d42eae3c-d874-4cc6-b01c-15f7a8010ab1"
gastos_do_mais_recente.plot()
# + [markdown] id="BCVIjxCB7Hgu"
# Plot padrão inviável.
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="IGPkJSl26mi7" outputId="4ccb455a-be28-480f-9139-6f63ce4d9c7a"
gastos_do_mais_recente.plot(kind='pie')
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="hR6jFr0x6p_9" outputId="242f947a-ec7f-45e6-9d08-fe1e89b4d34b"
gastos_do_mais_recente.sample(frac=1).plot(kind='pie')
# + [markdown] id="GIzk35re7JzL"
# Pie plot horrivel.
# - Legenda muito pequena
# - Número de fatias inacreditavel
# - Se não estiver ordenado, complicado de tirar conclusões não obvias
# + colab={"base_uri": "https://localhost:8080/"} id="58QOuMY267dA" outputId="5999203e-7336-4014-80ed-1ce256a5c931"
gastos_do_mais_recente / gastos_do_mais_recente.loc["33 Rio de Janeiro"]
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="jNI9eUcc74qX" outputId="8ea72fe7-43fb-4d3a-917d-babca88152e9"
gastos_do_mais_recente.plot(kind='bar')
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="qyVTqaCR9C2g" outputId="dd740720-9adb-4299-b0c3-2d5bdd72329d"
gastos_do_mais_recente = gastos_do_mais_recente.sort_values(ascending=False)
gastos_do_mais_recente.plot(kind='bar')
# + [markdown] id="Rhi1P2j9-_fb"
# # Desafios
# + [markdown] id="j-S40XkR_Dbl"
# Desafio 01: Buscar na documentação do Matplotlib como colocar um grid nos gráficos e adicionar nos gráficos de barra.
#
# Desafio 02: Fazer um gráfico e uma tabela do gasto dos outros estados em função do seu estado, ou qualquer outro de interesse.
#
# Desafio 03: Fazer o cálculo proporcional a população do seu estado e mais um a sua escolha.
#
# Desafio 04: Faça uma análise dos dados analisados, levante hipóteses e compartilhe com a gente no Discord.
#
# Desafio 05: Reproduza as análises feitas em aulas, refletindo e levantando hipóteses.
# + [markdown] id="WDxsQqqv_IV6"
# ## Desafio 01: Buscar na documentação do Matplotlib como colocar um grid nos gráficos e adicionar nos gráficos de barra.
# + colab={"base_uri": "https://localhost:8080/", "height": 387} id="67FnC8YJ9dpd" outputId="2f1f015a-caa1-47bd-a935-d12507147793"
gastos_do_mais_recente.plot(kind='bar', grid=True)
plt.show()
# + [markdown] id="EulP6Rae_YYS"
# ## Desafio 02: Fazer um gráfico e uma tabela do gasto dos outros estados em função do seu estado, ou qualquer outro de interesse.
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="AoAIadek_Otq" outputId="75420668-a5ba-42ad-c4e0-4b550db23502"
label_estado_ref_02 = "15 Pará"
desafio_02 = gastos_do_mais_recente / gastos_do_mais_recente.loc[label_estado_ref_02]
estado_ref = gastos_do_mais_recente.index.get_loc(label_estado_ref_02)
label_colors_02 = ['b']*27
label_colors_02[estado_ref] = 'r'
desafio_02.plot(kind='bar', grid=True, color=label_colors_02, figsize=(10,6))
# + [markdown] id="kl6v826aBuj3"
# ## Desafio 03: Fazer o cálculo proporcional a população do seu estado e mais um a sua escolha.
# + [markdown] id="m0MiJB-rClWW"
# Fonte: https://pt.wikipedia.org/wiki/Lista_de_unidades_federativas_do_Brasil_por_popula%C3%A7%C3%A3o
# + colab={"base_uri": "https://localhost:8080/", "height": 235} id="KiGdDI_TEUc9" outputId="02ffee39-8ee5-4e78-a162-69a31aad4a1a"
# !cp "/content/drive/MyDrive/alura/bootcamp-data-science/pop_por_estado_jul_2020.csv" .
pop_dados = pd.read_csv("/content/pop_por_estado_jul_2020.csv")
desafio_03_gastos_do_mais_recente = gastos_do_mais_recente.copy()
novos_indices = [" ".join(idx.split()[1:]) for idx in desafio_03_gastos_do_mais_recente.index]
desafio_03_gastos_do_mais_recente.index = novos_indices
pop_dados = pop_dados.set_index("Unidade federativa")
pop_dados["Gastos (mi)"] = desafio_03_gastos_do_mais_recente
pop_dados["Gastos per capta"] = pop_dados["Gastos (mi)"]*10**6 / pop_dados["População"]
pop_dados.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 480} id="ui2pXiWPHQWq" outputId="da7ccc20-d145-426c-9645-aa4c9ad2565a"
label_estado_ref_03 = "Pará"
desafio_03 = pop_dados["Gastos per capta"] / pop_dados["Gastos per capta"][label_estado_ref_03]
desafio_03 = desafio_03.sort_values(ascending=False)
estado_ref = desafio_03.index.get_loc(label_estado_ref_03)
label_colors_03 = ['b']*27
label_colors_03[estado_ref] = 'r'
ax = desafio_03.plot(kind='bar', grid=True, color=label_colors_03, figsize=(10,6))
# ax.yaxis.set_major_formatter(ticker.StrMethodFormatter("R$ {x:,.0f}"))
# + [markdown] id="SiN6wXH_NE49"
# ## Desafio 04: Faça uma análise dos dados analisados, levante hipóteses e compartilhe com a gente no Discord.
# + colab={"base_uri": "https://localhost:8080/", "height": 529} id="HpJNQjxXMaNG" outputId="9b888f39-fa22-4996-d061-3412681e33e2"
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(20,6))
desafio_02.plot(kind='bar', grid=True, color=label_colors_02, ax=axes[0],
title="Gastos proporcionais aos recursos destinados ao Pará")
desafio_03.plot(kind='bar', grid=True, color=label_colors_03, ax=axes[1],
title="Gastos proporcionais aos recursos destinados ao Pará por habitante")
# + [markdown] id="w1ZF0hmAPKaP"
# Embora o estado do Pará seja o 10<sup>o</sup> estado em valores absolutos de recursos, que o destaca muito dos outros estados da região norte. Proporcionalmente à sua população, é o 21<sup>o</sup> em valores per capita.
# Essa diferença de posição se dá pela sua quantidade de habitantes, que é elevada em comparação com os demais estados da região norte.
# + [markdown] id="FX0E7treOofH"
#
| module_1/ds_mod1_lecture5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Task Structured Tabular Data:
#
# #### Dataset Link:
# Dataset can be found at " /data/structured_data/data.csv " in the respective challenge's repo.
#
# #### Description:
# Tabular data is usually given in csv format (comma-separated-value). CSV files can be read and manipulated using pandas and numpy library in python. Most common datatypes in structured data are 'numerical' and 'categorical' data. Data processing is required to handle missing values, inconsistent string formats, missing commas, categorical variables and other different kinds of data inadequacies that you will get to experience in this course.
#
# #### Objective:
# How to process and manipulate basic structured data for machine learning (Check out helpful links section to get hints)
#
# #### Tasks:
# - Load the csv file (pandas.read_csv function)
# - Classify columns into two groups - numerical and categorical. Print column names for each group.
# - Print first 10 rows after handling missing values
# - One-Hot encode the categorical data
# - Standarize or normalize the numerical columns
#
# #### Ask yourself:
#
# - Why do we need feature encoding and scaling techniques?
# - What is ordinal data and should we one-hot encode ordinal data? Are any better ways to encode it?
# - What's the difference between normalization and standardization? Which technique is most suitable for this sample dataset?
# - Can you solve the level-up challenge: Complete all the above tasks without using scikit-learn library ?
#
# #### Helpful Links:
# - Nice introduction to handle missing values: https://analyticsindiamag.com/5-ways-handle-missing-values-machine-learning-datasets/
# - Scikit-learn documentation for one hot encoding: https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.OneHotEncoder.html
# - Difference between normalization and standardization: https://medium.com/towards-artificial-intelligence/how-when-and-why-should-you-normalize-standardize-rescale-your-data-3f083def38ff
# Import the required libraries
# Use terminal commands like "pip install numpy" to install packages
import numpy as np
import pandas as pd
# import sklearn if and when required
# ## Reading the CSV
df= pd.read_csv("data/structured_data/data.csv")
df.head(10)
# ## Various types of columns :
# # Categorical - Country, Purchased, Price Category of Purchase
# # Numerical - Age, Salary
# ## Filling Missing Values
df['Age'].replace(np.NaN, df['Age'].mean(),inplace=True)
df
df['Salary'].replace(np.NaN, df['Salary'].mean(),inplace=True)
df
# ## Using Label Encoder to convert categorical columns to suitable format
from sklearn.preprocessing import LabelEncoder
le= LabelEncoder()
le= le.fit(df['Country'])
df['Country']= le.transform(df['Country'])
# +
#df['Country']= le.fit_transform(df['Country'])
# -
df
# ## Using get dummies from pandas so that the Machine learning algorithm can interpret these categories
dummies = pd.get_dummies(df.Country)
df=pd.concat([df, dummies], axis=1)
df
df.drop("Country",axis=1,inplace=True)
df
le=LabelEncoder()
df['Purchased']= le.fit_transform(df['Purchased'])
df
df2= pd.DataFrame(columns=['Age','Salary'])
df2['Age']=df['Age']
df2['Salary']= df['Salary']
# ## Using Min-Max scaler to normalize the values
# +
from sklearn import preprocessing
x = df2.values #returns a numpy array
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
df2 = pd.DataFrame(x_scaled)
# -
# ## (X - min(col))/ (max(col)-min(col))
# ## Now values are between 0 and 1
df['Age']=df2[0]
df['Salary']= df2[1]
df
| Solutions + Numpy Basics/task_1_tabular.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sqlite3
import pandas as pd
connection = sqlite3.connect('database.sqlite')
print("Database opened successfully")
tables = pd.read_sql("SELECT * FROM sqlite_master WHERE type='table';", connection)
print(tables)
countries = pd.read_sql("SELECT * FROM Country;", connection)
countries.head()
players = pd.read_sql_query("SELECT * FROM Player", connection)
players.head()
selected_players = pd.read_sql_query("SELECT * FROM Player WHERE height >= 180 AND weight >= 170 ", connection)
print(selected_players)
# ! pip install mysql-connector-python
import mysql.connector
connection = mysql.connector.connect(host="127.0.0.1", # your host
user="root", # username
password="<PASSWORD>" ) # password
print(connection)
mycursor = connection.cursor()
mycursor.execute("SHOW DATABASES")
for x in mycursor:
print(x)
connection = mysql.connector.connect(host="127.0.0.1", # your host
user="root", # username
password="<PASSWORD>" ,
database = 'mysql')
mycursor = connection.cursor()
mycursor.execute("SHOW TABLES")
for x in mycursor:
print(x)
| Chapter02/SQL with Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <h1> Семинар 6. Семантическая сегментация, детекция.
# + [markdown] id="UmJo7IfF_58i"
# Скачаем и рахархивируем заранее все данные, которые понадобятся на семинаре + установим недостающий модуль
# + colab={"base_uri": "https://localhost:8080/"} id="MY2kFBuI_uK_" outputId="d7e73f24-db48-438e-d684-b07b34cb3865"
# !wget https://www.dropbox.com/s/tc1qo73rrm3gt3m/CARVANA.zip # Carvana dataset
# !wget https://www.dropbox.com/s/k886cusbuc1afnq/imagenet-mini.zip # mini image-net dataset
# !wget https://gist.githubusercontent.com/yrevar/942d3a0ac09ec9e5eb3a/raw/238f720ff059c1f82f368259d1ca4ffa5dd8f9f5/imagenet1000_clsidx_to_labels.txt # классы имаджнета
# !unzip -q CARVANA.zip
# !unzip -q imagenet-mini.zip
# !rm -rf ./train/.DS_Store
# !rm -rf ./train_masks/.DS_Store
# !pip install colour
# + colab={"base_uri": "https://localhost:8080/"} id="7upRqguxAo90" outputId="169b4058-db96-443a-db2f-56a9cb92964a"
# !ls -l
# + [markdown] id="ubc0wbOl_iUQ"
# <h2>Часть 1. Семантическая сегментация
# + id="jq_ppf6t_iUQ"
import os
import warnings
from os.path import isfile, join
from typing import Dict, List, Tuple, Union
import cv2
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import torch
import torch.nn.functional as F
import torchvision
import torch.optim as optim
from colour import Color
from PIL import Image
from torch import nn
from torch.utils.data import DataLoader
from torch.utils.data.dataset import Dataset
from torchvision import datasets, models, transforms
from tqdm.notebook import trange, tqdm
warnings.filterwarnings("ignore")
# + [markdown] id="_ff_VyjsEaym"
# <h> В этой части семинара мы напишем свой U-net с нуля, без мам, пап и кредитов,
# обучим и попробуем его на задачу распознавания точной маски автомобиля
# + id="b_jCFjB2_iUT"
class Carvana(Dataset):
def __init__(
self,
root: str,
transform: transforms.Compose = None,
) -> None:
"""
:param root: путь к папке с данными
:param transform: transforms of the images and labels
"""
self.root = os.path.expanduser(root)
self.transform = transform
(self.data_path, self.labels_path) = ([], [])
def load_images(path: str) -> List[str]:
"""
Возвращает список с путями до всех изображений
:param path: путь к папке с данными
:return: лист с путями до всех изображений
"""
images_dir = [join(path, f) for f in os.listdir(path)
if isfile(join(path, f))]
images_dir.sort()
return images_dir
self.data_path = load_images(self.root + "/train")
self.labels_path = load_images(self.root + "/train_masks")
def __getitem__(self, index: int) -> Tuple[torch.Tensor, torch.Tensor]:
"""
:param index: sample index
:return: tuple (img, target) with the input data and its label
"""
img = Image.open(self.data_path[index])
target = Image.open(self.labels_path[index])
if self.transform is not None:
img = self.transform(img)
target = self.transform(target)
target = (target > 0).float()
return (img, target)
def __len__(self):
return len(self.data_path)
def im_show(img_list: List[Tuple[str, str]]) -> None:
"""
Plots images with correspinding segmentation mask
:param img_list: list of paths to images
"""
to_PIL = transforms.ToPILImage()
if len(img_list) > 9:
raise Exception("len(img_list) must be smaller than 10")
fig, axes = plt.subplots(len(img_list), 2, figsize=(16, 16))
fig.tight_layout()
for (idx, sample) in enumerate(img_list):
a = axes[idx][0].imshow(np.array(to_PIL(sample[0])))
b = axes[idx][1].imshow(np.array(to_PIL(sample[1])))
for ax in axes[idx]:
ax.get_xaxis().set_visible(False)
ax.get_yaxis().set_visible(False)
plt.show()
# + [markdown] id="HLiA3F2SMQiA"
# Загрузим датасет при помощи определенного выше класса
# + id="eVBdhAWMLlzy"
train_dataset = Carvana(
root=".",
transform=transforms.Compose([
transforms.Resize((256, 256)),
transforms.ToTensor()
])
)
train_loader = torch.utils.data.DataLoader(
dataset=train_dataset,
batch_size=128,
shuffle=True,
pin_memory=True,
num_workers=4
)
# + [markdown] id="FKUcQZpLMTy_"
# Посмотрим несколько примеров изображений и масок автомобиля
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="2VIj0iLOLou-" outputId="87886f75-29c0-4bee-8902-a6001cfab05b"
img_list = []
for i in range(4):
img, label = train_dataset[np.random.randint(0, 100)]
img_list.append((img, label))
im_show(img_list)
# + [markdown] id="LKLjoUgsLqr1"
# Дополните недостающий код таким образом, чтобы получилась архитектура U-net сети (https://arxiv.org/pdf/1505.04597.pdf).
# Обратите внимание, что при проходе "вниз" количество каналов каждого блока __увеличивается в два раза__.
# Ситуация с проходом "вверх" противоположна, количество каналлов __уменьшается вдвое__.
# Также, начинаем мы __не с 64 каналов__, как на схеме ниже, __а с 16 каналов__.
#
# При проходе вниз в нашей реализации предлагается __дойти до 128 каналов__, чтобы сэкономить время обучения, в оригинальной статье было до 1024. (2 блока, вместо 4х)
# В целом, при отличии количества каналлов от указанных выше чисел, модель все равно будет работать, но для удобства проведения семинара лучше всем договориться об одних числах.
#
# <img src="https://lmb.informatik.uni-freiburg.de/people/ronneber/u-net/u-net-architecture.png">
#
# Подсказка: каждый блок по пути "вниз" представляет из себя двойную свертку с батч нормом и активацией (при создании `Relu` слоя не забывайте `inplace=True`).
# + id="j-8ltVJe_iUV"
class ConvBlock(nn.Module):
def __init__(self, in_size, out_size, kernel_size=3, padding=1, stride=1):
super(ConvBlock, self).__init__()
self.conv = # <your code here>
self.bn = nn.BatchNorm2d(out_size)
self.relu = # <your code here>
def forward(self, x):
return self.relu(self.bn(self.conv(x)))
class Unet(nn.Module):
def __init__(self):
super().__init__()
self.down_1 = nn.Sequential(
ConvBlock(3, 16),
ConvBlock(16, 32, stride=2, padding=1)
)
self.down_2 = # <your code here>
self.middle = ConvBlock(128, 128, kernel_size=1, padding=0)
self.up_2 = # <your code here>
self.up_1 = # <your code here>
self.output = nn.Sequential(
ConvBlock(32, 16),
ConvBlock(16, 1, kernel_size=1, padding=0)
)
def forward(self, x):
down1 = # <your code here>
out = F.max_pool2d(down1, kernel_size=2, stride=2)
down2 = # <your code here>
out = F.max_pool2d(down2, kernel_size=2, stride=2)
out = self.middle(out)
out = nn.functional.interpolate(out, scale_factor=2) # интерполяцией увеличиваем размер фильтра вдвое
out = torch.cat([down2, out], 1)
out = self.up_2(out)
out = # <your code here>
out = # <your code here>
out = # <your code here>
out = nn.functional.interpolate(out, scale_factor=2)
return self.output(out)
# + [markdown] id="tE58rn_ZMohS"
# Определяем функцию для обучения одной эпохи
# + id="b5ZF0LeS_iUb"
def train(train_loader, model, criterion, epoch, num_epochs, device):
model.train()
pbar = tqdm(enumerate(train_loader), total=len(train_loader))
for i, (images, labels) in pbar:
images = images.to(device)
labels = labels.to(device)
optimizer.zero_grad()
outputs = torch.sigmoid(model(images))
loss = criterion(outputs, labels)
accuracy = ((outputs > 0.5) == labels).float().mean()
pbar.set_description(
f"Loss: {round(loss.item(), 4)} "
f"Accuracy: {round(accuracy.item() * 100, 4)}"
)
loss.backward()
optimizer.step()
# + colab={"base_uri": "https://localhost:8080/"} id="4RnYf1lS_iUd" outputId="4d929f03-4843-4634-95d4-c29a105d0119"
device = "cuda"
model = Unet().to(device)
criterion = torch.nn.BCELoss() # https://pytorch.org/docs/stable/generated/torch.nn.BCELoss.html
optimizer = torch.optim.RMSprop(
model.parameters(),
weight_decay=1e-4,
lr=1e-4,
momentum=0.9
)
num_epochs = 1
for epoch in range(0, num_epochs):
train(train_loader, model, criterion, epoch, num_epochs, device=device)
# + [markdown] id="G07YWsb2FDff"
# Посмотрим на результаты
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="5aytNIaaDa4S" outputId="757aa449-0c67-4f8e-ba69-c460902f374d"
img_list = []
for i in range(4):
img, label = train_dataset[np.random.randint(0, 100)]
pred = model(img.unsqueeze(0).cuda()).detach().squeeze(0).cpu()
img_list.append((img, pred))
im_show(img_list)
# + [markdown] id="kVvff-qgkdvT"
# Еще несколько эпох и никакие гринскрины нужны не будут...
# + id="DjnzaZ9vNqX7"
# освободим память
torch.cuda.empty_cache()
# + [markdown] id="sr09Rv6F_iUf"
# ---
# + [markdown] id="BQv54rfdELFv"
# <h2>Часть 2. mean Average Precision
# + [markdown] id="eBJgNh0WELFv"
# Для определения метрики mean Average Precision понадобятся проделать небольшой путь и вспомнить пару понятий.
# + [markdown] id="dHuMyUHKELFw"
# Вспоминаем второй класс ИАДа:
# <br>
# $$ Precision = {TP \over TP + FP} $$
# <br>
# $$ Recall = {TP \over TP + FN} $$
# <br>
# <br>TP - True Positive
# <br>FP - False Positive
# <br>FN - False Negative
# <br> <br> __Precision__ - доля объектов, названных классификатором положительными и при этом действительно являющимися положительными, среди всех объектов, названных классификатором положительными.
#
# __Recall__ - доля объектов, названных классификатором положительными и при этом действительно являющимися положительными, среди всех истинно положительных объектов.
# + [markdown] id="MLB8y0--ELFx"
# <h4>Вспоминаем лекцию 6:
# Intersection over Union</h4>
# <img src=https://www.pyimagesearch.com/wp-content/uploads/2016/09/iou_equation.png>
#
# + [markdown] id="pI5Y3JDlELFx"
# В задаче детекции метки TP, FP, FN (чаще всего) выдаются по следующей логике:
#
# - метка TP выдается в случае если IoU > 0.5 и класс определен правильно
# - метка FP выдается в случае если IoU <= 0.5 и/или если Bounding Box'ы дублируются
# - метка FN выдается если если IoU > 0.5, но неправильно определен класс и/или если бокса нет совсем
# + [markdown] id="gQqzHZmwTTT3"
# <h3> Чтобы было нагляднее: </h3>
#
# * зеленый цвет - истинный бокс и класс
# * синий - наши предсказания
#
# <h4>True Positive </h4> У нас два волка, оба определены своим классом и боксы очевидно имеют IoU больше 0.5.
# <img src="TruePositiveVolks.jpg">
#
# <h4>False Positive </h4> Несмотря на то, что класс определен правильно и бокс в целом выглядит логично на своем месте, IoU слишком мал, поэтому такая детекция получает метку FP.
# <img src="FalsePositiveVolk.jpg">
#
# <h4>False Negative </h4> Потому что несмотря на хорошее пересечение предсказанного бокса с целевым, класс с высокой уверенностью определен неправильно.
# <img src="FalseNegativeVolk.jpg">
# + [markdown] id="v8TbTIshELFx"
# <h3> PR-кривая </h3>
#
# Далее, для подсчёта mAP, нужно построить PR-кривую.
# Напомним, что это кривая, у которой по оси Y - значение Precision, а по оси X - значение recall.
# Эти значения считаются при переборе пороговых вероятностей, начиная с которых объект помечается положительным классом.
#
# Для задачи бинарной классификации мы когда-то строили такую кривую, теперь рассмотрим чуть более сложный случай, где у нас три класса: Волк, Лев и Тигр + решается задача детекции, а не классификации.
#
#
# Внизу представлена таблица с игрушечными данными по предсказаниям модели.
# Допустим из 7 объектов, в датасете у нас только 3 волка.
# В данном случае мы называли объект "действительно волком" если он имел правильный класс и IoU не менее 0.5.
# Таким образом имеем задачу вида one vs all, где интересующим нас классом будет являться именно "волк".
# + [markdown] id="WwawmoyeELFy"
# |Номер строки| Уверенность в том что волк (истинный класс) |IoU не менее 0.5? | Precision | Recall |
# |------------|------------------------|----------------------|-------------|---------|
# |1 |0.92 (Волк) | True | 1.0 | 0.33 |
# |2 |0.83 (Волк) | True | 1.0 | 0.67 |
# |3 |0.77 (Волк) | False | 0.67 | 0.67 |
# |4 |0.71 (Лев) | False | 0.50 | ... |
# |5 |0.67 (Тигр) | False | 0.40 | .... |
# |6 |0.54 (Волк) | True | 0.50 | .... |
# |7 |0.47 (Тигр) | False | 0.50 | 1.0 |
# + [markdown] id="ehDXMLbjELFy"
# <h6>Посчитаем Precision и Recall для порога в 0.9:</h6>
# Здесь все легко, взяли порог в 0.9 и называем волками всех, у кого уверенность в классе "Волк" больше 0.9. Один TP, отсутсвуют FP и два FN (2 и 6 строчки).
# $$Precision ={ 1 \over 1 + 0} = 1.0$$
#
# $$Recall = {1 \over 1 + 2} = 0.33$$
#
# <h6>для порога в 0.8:</h6>
# Здесь тоже без дополнительных сложностей. С таким порогом, во второй строчке у нас нашелся еще один TP, соотоветственно убавился один FN. Остальное осталось так же.
# $$Precision ={ 2 \over 2 + 0} = 1.0$$
#
# $$Recall = {2 \over 2 + 1} = 0.67$$
#
# <h6>для порога в 0.75:</h6>
# А вот при пороге в 0.75 в третьей строчке замечаем, что несмотря на то, что истинный класс действительно "Волк", чем мы и называем данный объект - IoU c истинным боксом меньше 0.5, поэтому присваиваем метку FP.
# $$Precision ={ 2 \over 2 + 1} = 0.67$$
#
# $$Recall = {2 \over 2 + 1} = 0.67$$
#
# <h6>Дальше сами :)</h6>
# <h6>Задание: Посчитать недостающие в таблице значения precision и recall и сравнить с изображенной PR кривой</h6>
#
# PR кривая будет вылядеть следуюшим образом:
# <img src="pr_uno.png">
# + [markdown] id="62fZzJWCELF4"
# <h3> Монотонная PR-кривая </h3>
#
# **Average Precision (AP)** стандартно **определяется как AUC-PR**, то есть как площадь по PR кривой. Из-за того, что Precision и Recall находятся в отрезке от 0 до 1, AP так же определена на этом отрезке. Чем ближе к 1, тем качественнее модель.
#
# Для удобства вычислений, и чуть большей устойчивости к перестановке - вместо того, чтобы терпеть возникшую немонотонность, для всех совпадающих значений recall'a берется максимальный справа от текущей точки precision, то есть график изменится следующим образом:
#
# <img src="pr_dos.png">
#
# Технология та же - для вычисления AP считается AUC под красной кривой:
#
# $$AP = 1 * 0.67 + (1 - 0.67) * 0.5 = 0.835 $$
# + [markdown] id="ZlQ8t0jMELF8"
# <h3>k-point interpolation</h3>
#
# В какой-то момент люди решили, что просто площади теперь не круто и в популярном соревновании PASCAL VOC2008 для вычисления Average Precision использовалась 11-point interpolation.
#
# По-простому: брались 11 значений монотонной PR функции, в точках 0, 0.1, 0.2, ..., 0.9, 1.0 и усреднялись. <br>Для любителей формул:
#
# $$AP = {1 \over11} * \sum P(r), r \in [0.1, 0.2, ..., 0.9, 1.0]$$
# $P(r)$ - значение Precision при определенном Recall.
#
# Графически это все выглядит следующим образом:
#
# <img src="pr_tres.png">
#
# Посчитаем АР данным способом:
# $$AP = {1 \over {11}} (1 * 7 + 3 * 0.5) = 0.77$$
# + [markdown] id="0dN1KII-ELF_"
# <h3>Подсчёт mAP</h3>
#
# Для более <font color="green">fresh </font> PASCAl VOC соревнований (2011 - 2012) было принято решение считать **Average Precision** как обычную __площадь под монотонной PR кривой__. Этого определения мы и будем придерживаться далее.
#
# Все предыдущие графики, наша первая табличка и значения AP считались для одного класса "Волк". Понятно, что подобные значения можно посчитать для каждого класса в выборке. И каждый раз будет принцип one vs all, где различается нужный класс и "все остальные"
#
#
# Метрика __mean Average Precison__ считается как среднее между __AP__ каждого класса, те:
#
# $$mAP = \sum_{c \in C} AP(c)$$
# где С - множество классов.
#
# <br>Ниже - пример того как считать среднее :))
#
# *для Average Precision льва и тигра значения взяты случайно
#
# |AP(Волк)| AP(Лев) | AP(Тигр)|mAp |
# |--------|---------|---------|-------|
# |0.835 |0.83 | 0.77 | 0.81 |
# + [markdown] id="qQAG0kBGELF_"
# ---
# + [markdown] id="JiIqDhk_ELGA"
# <h2>Часть 3. Детекция объектов на изображениях</h2>
# -
# Сегодня на семинаре мы попробуем обучить модель детекции.
# Мы не будем полностью реализовывать архитектуру сети, а возьмем предобученную Faster R-CNN из pytorch и сделаем fine-tuning.
# + [markdown] id="icjhclUNIgCk"
# Для начала загрузим данные. Мы будем пользоваться датасетом [Penn-Fudan](https://www.cis.upenn.edu/~jshi/ped_html/). Он содержит 170 изображений с разметкой сегментации и детекции. Мы воспользуемся последней
# + colab={"base_uri": "https://localhost:8080/"} id="NaGQea5hIgCn" outputId="2b25078d-617d-4ccc-cec2-cd4ac140d703"
# ! curl https://www.cis.upenn.edu/~jshi/ped_html/PennFudanPed.zip > PennFudanPed.zip
# ! unzip -q PennFudanPed.zip
# + id="ZtNWxqoMESqw"
from PIL import ImageDraw
from sklearn.metrics import auc
from torchvision.models.detection import fasterrcnn_resnet50_fpn
from torchvision.models.detection.faster_rcnn import FastRCNNPredictor
# + id="U1g32XYFFYTb"
class PennFudanDataset(torch.utils.data.Dataset):
def __init__(self, root: str, transforms_list: transforms = None):
self.root = root
self.transforms = transforms_list
self.imgs = list(sorted(os.listdir(os.path.join(root, "PNGImages"))))
self.masks = list(sorted(os.listdir(os.path.join(root, "PedMasks"))))
def __getitem__(self, idx):
img_path = os.path.join(self.root, "PNGImages", self.imgs[idx])
mask_path = os.path.join(self.root, "PedMasks", self.masks[idx])
img = Image.open(img_path).convert("RGB")
# note that we haven't converted the mask to RGB,
# because each color corresponds to a different instance
# with 0 being background
mask = Image.open(mask_path)
mask = np.array(mask)
# instances are encoded as different colors
obj_ids = np.unique(mask)
# first id is the background, so remove it
obj_ids = obj_ids[1:]
# split the color-encoded mask into a set
# of binary masks
masks = mask == obj_ids[:, None, None]
# get bounding box coordinates for each mask
num_objs = len(obj_ids)
boxes = []
for i in range(num_objs):
pos = np.where(masks[i])
xmin = np.min(pos[1])
xmax = np.max(pos[1])
ymin = np.min(pos[0])
ymax = np.max(pos[0])
boxes.append([xmin, ymin, xmax, ymax])
# convert everything into a torch.Tensor
boxes = torch.as_tensor(boxes, dtype=torch.float32)
# there is only one class
labels = torch.ones((num_objs,), dtype=torch.int64)
image_id = torch.tensor([idx])
target = {"boxes": boxes, "labels": labels, "image_id": image_id}
if self.transforms is not None:
img = self.transforms(img)
return img, target
def __len__(self):
return len(self.imgs)
def get_transform(train=False):
transforms_list = []
if train:
transforms.append(transforms.RandomHorizontalFlip(0.5))
transforms_list.append(transforms.ToTensor())
return transforms.Compose(transforms_list)
# + id="aYRhV4hfGpq9"
dataset = PennFudanDataset("PennFudanPed/", get_transform())
# + colab={"base_uri": "https://localhost:8080/", "height": 553} id="2PJGAZYs6mQE" outputId="1e91df1b-127b-43ca-c387-4639ac092220"
image, labels = next(iter(dataset))
image = transforms.ToPILImage()(image)
draw = ImageDraw.Draw(image)
for box in labels["boxes"]:
draw.rectangle([(box[0], box[1]), (box[2], box[3])])
image
# + [markdown] id="VABmgMQ3IgDI"
# Теперь подгрузим модель. Среди моделей детекции в pytorch есть только `Faster R-CNN`. Так как мы детектим только 2 класса - пешеходов и фон, нужно изменить выходной слой, предсказывающий классы изображений. Для этого в torchvision есть блок `FastRCNNPredictor`.
#
# <h4>Схема модели Fast R-CNN</h4>
# <img src=https://pytorch.org/tutorials/_static/img/tv_tutorial/tv_image03.png>
# + id="kSUjeN0M7EWk"
def get_detection_model(num_classes=2):
model = fasterrcnn_resnet50_fpn(pretrained=True)
# get the number of input features for the classifier
in_features = model.roi_heads.box_predictor.cls_score.in_features
# replace the pre-trained head with a new one
model.roi_heads.box_predictor = FastRCNNPredictor(in_features, num_classes)
return model
# + [markdown] id="MBvGLhxOIgDP"
# Теперь давайте напишем подсчет IoU, которая пригодится нам для отбора кандидатов.
#
# <h4>Intersection over Union </h4>
# <img src=https://www.pyimagesearch.com/wp-content/uploads/2016/09/iou_equation.png>
# + id="Fx3qsM_fIgDP"
def intersection_over_union(dt_bbox: np.array, gt_bbox: np.array) -> float:
"""
Intersection over Union between two bboxes
(x0, y0) - coordinates of the upper left bbox corner
(x1, y1) - coordinates of the down right bbox corner
:param dt_bbox: list or numpy array of size (4,) [x0, y0, x1, y1]
:param gt_bbox: list or numpy array of size (4,) [x0, y0, x1, y1]
:return : intersection over union
"""
# <your code here>
# + colab={"base_uri": "https://localhost:8080/"} id="mV4wTZpMNotf" outputId="01e21e5b-c601-4bae-aaf8-b7b69d8faf91"
dt_bbox = [0, 0, 2, 2]
gt_bbox = [1, 1, 3, 3]
intersection_over_union(dt_bbox, gt_bbox)
# + id="5Pkva3QFDDed"
def evaluate_sample(target_pred: torch.Tensor, target_true, iou_threshold=0.5):
# правильные прямоугольники
gt_bboxes = target_true["boxes"].numpy()
gt_labels = target_true["labels"].numpy()
# предсказания модели
dt_bboxes = target_pred["boxes"].numpy()
dt_labels = target_pred["labels"].numpy()
dt_scores = target_pred["scores"].numpy()
results = []
# для каждого прямоугольника из предсказания находим максимально близкий прямоугольник среди ответов
for detection_id in range(len(dt_labels)):
dt_bbox = dt_bboxes[detection_id, :]
dt_label = dt_labels[detection_id]
dt_score = dt_scores[detection_id]
detection_result_dict = {"score": dt_score}
max_IoU = 0
max_gt_id = -1
for gt_id in range(len(gt_labels)):
gt_bbox = gt_bboxes[gt_id, :]
gt_label = gt_labels[gt_id]
if gt_label != dt_label:
continue
if intersection_over_union(dt_bbox, gt_bbox) > max_IoU:
max_IoU = intersection_over_union(dt_bbox, gt_bbox)
max_gt_id = gt_id
if max_gt_id >= 0 and max_IoU >= iou_threshold:
# для прямоугольника detection_id нашли правильный ответ, который имеет IoU больше 0.5
detection_result_dict["TP"] = 1
# удаляем эти прямоугольники из данных, чтобы больше не матчить с ними
gt_labels = np.delete(gt_labels, max_gt_id, axis=0)
gt_bboxes = np.delete(gt_bboxes, max_gt_id, axis=0)
else:
detection_result_dict["TP"] = 0
results.append(detection_result_dict)
# возвращаем результат, для кажого прямоугольника говорим, смогли ли сматчить его с чем то из ответов
return results
# + id="roHlrGtVIgDV"
def evaluate(model, test_loader, device):
results = []
model.eval()
nbr_boxes = 0
with torch.no_grad():
for batch, (images, targets_true) in enumerate(test_loader):
images = list(image.to(device).float() for image in images)
targets_pred = model(images)
targets_true = [{k: v.cpu().float() for k, v in t.items()} for t in targets_true]
targets_pred = [{k: v.cpu().float() for k, v in t.items()} for t in targets_pred]
for i in range(len(targets_true)):
target_true = targets_true[i]
target_pred = targets_pred[i]
nbr_boxes += target_true["labels"].shape[0]
# матчим ответы с правильными боксами
results.extend(evaluate_sample(target_pred, target_true))
results = sorted(results, key=lambda k: k["score"], reverse=True)
# считаем точность и полноту, чтобы потом посчитать mAP как auc
acc_TP = np.zeros(len(results))
acc_FP = np.zeros(len(results))
recall = np.zeros(len(results))
precision = np.zeros(len(results))
if results[0]["TP"] == 1:
acc_TP[0] = 1
else:
acc_FP[0] = 1
for i in range(1, len(results)):
acc_TP[i] = results[i]["TP"] + acc_TP[i - 1]
acc_FP[i] = (1 - results[i]["TP"]) + acc_FP[i - 1]
precision[i] = acc_TP[i] / (acc_TP[i] + acc_FP[i])
recall[i] = acc_TP[i] / nbr_boxes
return auc(recall, precision)
# + id="fsqKh0gaIgDa"
def train_one_epoch(model, optimizer, data_loader, device):
model.train()
global_loss = 0
for images, targets in tqdm(data_loader, leave=False, desc="Batch number"):
images = list(image.to(device).float() for image in images)
targets = [{k: v.to(device) for k, v in t.items()} for t in targets]
dict_loss = model(images, targets)
losses = sum(loss for loss in dict_loss.values())
optimizer.zero_grad()
losses.backward()
optimizer.step()
global_loss += float(losses.cpu().detach().numpy())
return global_loss
def train(model, train_loader, test_loader, optimizer, device, num_epochs=5):
for epoch in trange(num_epochs, leave=True, desc=f"Epoch number"):
train_one_epoch(model, optimizer, train_loader, device)
mAP = evaluate(model, test_loader, device=device)
print(f"mAP after epoch {epoch + 1} is {mAP:.3f}")
# + id="PEzKc4eR9BHn"
def collate_fn(batch):
return tuple(zip(*batch))
dataset = PennFudanDataset("PennFudanPed", get_transform(train=False))
indices = torch.randperm(len(dataset)).tolist()
dataset_train = torch.utils.data.Subset(dataset, indices[:-50])
data_loader_train = torch.utils.data.DataLoader(
dataset_train,
batch_size=4,
shuffle=True,
pin_memory=True,
num_workers=4,
collate_fn=collate_fn
)
dataset_test = torch.utils.data.Subset(dataset, indices[-50:])
data_loader_test = torch.utils.data.DataLoader(
dataset_test,
batch_size=1,
shuffle=False,
num_workers=4,
collate_fn=collate_fn
)
# + colab={"base_uri": "https://localhost:8080/", "height": 868, "referenced_widgets": ["0dff8103f53b4b38adb088af2f2d8956", "29bb1547ec9545ce85f339693c2af46b", "4f90da335c9a4cbf8d10fe1dc3c78813", "13277a01bbb94b6ca6c24d60bf29fcfd", "1788c9caf91b4c2bb8d2ef0d291f4b14", "7b7e261e948d4bfbb25df72521f25331", "12f7d44ba70b4b8fb9113164bb597fd9", "88b26b728294483681449ffba56cac49"]} id="0vpF4zUbIgDt" outputId="ebbf2160-74bb-4d7e-dc56-7d8ad3d332aa"
num_classes = 2
device = torch.device("cuda") if torch.cuda.is_available() else torch.device("cpu")
model = get_detection_model(num_classes)
model.to(device)
params = [p for p in model.parameters() if p.requires_grad]
optimizer = torch.optim.SGD(params, lr=0.005, momentum=0.9, weight_decay=0.0005)
train(model, data_loader_train, data_loader_test, optimizer, device)
# + [markdown] id="K0DlzqfsIgDv"
# Давайте посмотрим, как наша модель научилась выделять людей
# + colab={"base_uri": "https://localhost:8080/", "height": 341} id="lZ8_iGQzIgDx" outputId="c7ec4729-846f-441b-9d45-8fa5eab2b4fd"
image, labels = next(iter(dataset_test))
model.eval()
pred = model(image.unsqueeze(0).to(device))[0]
image = transforms.ToPILImage()(image)
draw = ImageDraw.Draw(image)
for box in labels["boxes"]:
draw.rectangle([(box[0], box[1]), (box[2], box[3])])
for box in pred["boxes"]:
draw.rectangle([(box[0], box[1]), (box[2], box[3])], outline="red")
image
# + colab={"base_uri": "https://localhost:8080/"} id="Qrj1cdioj3I3" outputId="82d982e3-5452-4633-a19e-24ac25f3f3ec"
pred
# + colab={"base_uri": "https://localhost:8080/"} id="02VoP-2aMggf" outputId="c5ed9d3c-f23a-4c78-85dd-16c50ecae0b8"
for l_box in labels["boxes"]:
print(intersection_over_union(pred["boxes"][0].cpu().detach().numpy(), l_box.numpy()))
| Courses/IadMl/IntroToDeepLearning/seminars/sem06/sem_06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="5Cr1bQVn_WG7"
# # fastai to PyTorch to Lightning - Part 1
# > A Transfer Learning Comparison
#
# - toc: false
# - branch: master
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [fastai, jupyter, pytorch, pytorch-lightning]
# - image: images/fastai-pytorch-pl/thumbnail.png
# -
# Having been a user of fastai for a while, I figured it was time to try to learn PyTorch properly. As a learning exercise, I decided to try to recreate a fastai image classifier system using transfer learning within Pytorch and, having also recently read about PyTorch Lightning, then take the PyTorch model and recreate it within PyTorch Lightning. This blog series will therefore be in two parts, of which this is Part 1:
#
# 1. fastai to PyTorch
# 2. [PyTorch to PyTorch Lightning](https://mikful.github.io/blog/fastai/jupyter/pytorch/pytorch-lightning/2020/09/21/fastai-pytorch-lightning-part2.html)
#
#
# I found this exercise to be very beneficial for a number of reasons:
#
# 1. I came to understand training a PyTorch model in far greater detail than I had before
# 2. I learned about the trickiness of training a pure PyTorch model to high accuracy vs the by-default implemented best-practices of fastai for image classification
# 3. Having seen how pure PyTorch model is implemented, I came to appreciate the benefits of organising my code into PyTorch Lightning modules
#
# For the comparison we will use the Imagenette dataset, a subset of 10 classes from the original ImageNet dataset devised by fastai to aid the implementation of new algorithms, which is simple enough for the exercise.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="f-O8fRfZ_nYa" outputId="86a7d8b3-1150-459b-da07-fb8ad9b909e1"
#hide
# !pip install -U fastai -q
# and restart runtime
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="fjIDi8ReBD3z" outputId="8c24af51-9c3c-43cd-d5a9-5d422173196d"
#hide
# !pip install pytorch-lightning -q
# + colab={} colab_type="code" id="Ve-W4JQy_WG8"
#hide
import torch
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
import numpy as np
import torchvision
from torchvision import datasets, models, transforms
import matplotlib.pyplot as plt
import time
import os
import copy
from pathlib import Path
import random
from PIL import Image
import fastai
from fastai.vision.all import *
import copy
# reproducibility
torch.manual_seed(0)
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
np.random.seed(0)
cuda_available = torch.cuda.is_available()
# + [markdown] colab_type="text" id="Zu45QUnY_WKN"
# # fastai
#
# Let's first of all set a benchmark performance using fastai's high level API `ImageDataLoaders` and `cnn_learner` functions.
# + [markdown] colab_type="text" id="B-jOOvJv_WHC"
# ## Download Dataset and Define Data Path
# + colab={} colab_type="code" id="F1No3sVz_WHD"
data_dir = untar_data(URLs.IMAGENETTE_160)
# + colab={} colab_type="code" id="D5LPKcPF_WKO"
from fastai.vision.all import *
# -
# The main benefit of the fastai API is that best-practices are instantiated by default, allowing you to get SOTA results extremely quickly on the main task domains the API is designed for (vision, text and tabular) and with a little extra work elsewhere.
#
# A great example of this is the image transformation pipeline, that's well described in [part 5](https://github.com/fastai/fastbook/blob/master/05_pet_breeds.ipynb) of the fastai fastbook. Namely, this incorporates the following benefits which to lead to excellent results:
#
# 1. "Presizing" - i.e. take a larger section of the image on the item transforms (which creates uniform image sizes before they are passed to the GPU for batch transformations)
# 2. the `*aug_transforms` which are a mixture of random flips, warps, brightness and colour changes and random resizing and cropping that are undertaken on the GPU for speed and also in one combined augmentation step, rather than many different steps which leads to interpolation artifacts.
# 3. Normalization by the ImageNet statistics as we are using a pretrained Resnet50, with transfer learning.
#
# On the validation set, by default a centre crop of the images will be taken.
# + colab={} colab_type="code" id="EBufwVVG_WKQ"
dls = ImageDataLoaders.from_folder(data_dir,
valid='val',
item_tfms=RandomResizedCrop(224, min_scale=0.75),
batch_tfms=[*aug_transforms(),Normalize.from_stats(*imagenet_stats)])
# + colab={"base_uri": "https://localhost:8080/", "height": 536} colab_type="code" id="8KYhPpv0_WKS" outputId="2a4b937c-f0f6-40aa-e596-5a7164d39094"
dls.show_batch()
# -
# Another main benefit of fastai is that we can create a transfer learning CNN with a pretrained body (a pretrained resnet18 from the PyTorch Model zoo here), custom head and automatically defined loss function (based on the dataset) and optimizer extremely easily (`CrossEntropyLossFlat` and `Adam`, respectively, in this case). Within this custom head, fastai also implements an `AdaptiveConcatPool` layer, which essentially concatenates `MaxPool` and `AveragePool` layers, such that we can extract more information from the dataset. We'll see the PyTorch implementation of this later. The pretrained base model is automatically frozen other than the `BatchNorm` layers, such that it acts as a fixed-feature extractor for the custom head in the initial training stage before fine-tuning both the head and body's weights.
# + colab={"base_uri": "https://localhost:8080/", "height": 83, "referenced_widgets": ["7ba6b7960d4a4f438471b91e86cab25c", "8e28400e477c4cec95074ea63f85e927", "509d394dd079442387dd7900cf7dd788", "cabf400dbfa045b1a3b09d84a6fcf3c2", "bfba5983ad5f43ad8f3927e13ae2496d", "a35ad9ddb2ac4d4e92f2d8ce8947b6d6", "8bdab047ff314d6180a7f16e9ee7fd72", "b327c4a68a2141899c3a554e9fe41d6d"]} colab_type="code" id="d-RpRJ_x_WKU" outputId="775fc317-4c3e-4d8f-fdec-b3f59886a4a8"
learn = cnn_learner(dls, resnet18, metrics=accuracy)
# -
# Once more, we see a benefit of fastai - a very handy learning rate finder, which is critical to defining the correct maximum learning rate during training.
# + colab={"base_uri": "https://localhost:8080/", "height": 300} colab_type="code" id="k4c61mDE_WKW" outputId="0489dcbd-95f0-49c0-dcec-823d089c79c9"
learn.lr_find()
# -
# Another of fastai's useful functions is the one-cycle policy training loop, which automatically implements Leslie Smith's cosine annealing one-cycle learning rate schedule over the full training duration, while inversely varying the Adam optimizer momentum in another schedule.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="4QTNBPD2_WKY" outputId="7dbdc39f-d6a0-4805-ebb7-081c40a7e55e"
learn.fit_one_cycle(5, lr=3e-3)
# -
# We can see that we've acheived a very high accuracy very quickly. So let's now unfreeze the base model, find a new learning rate and fine-tune the model.
# + colab={} colab_type="code" id="E-Xuv1eu_WKZ"
learn.unfreeze()
# + colab={"base_uri": "https://localhost:8080/", "height": 300} colab_type="code" id="BOFrV8oS_WKb" outputId="87fcaec8-6ab3-4dce-b292-f8d93c79c35c"
learn.lr_find()
# -
# We see once more a nice easy implementation here - discriminative learning rates for the pretrained resnet and the custom head of the model, by the `slice` method, such that we don't damage the pretrained weights by using a too-high learning rate.
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="ZvBIJoCn_WKd" outputId="356965ea-eaf0-4174-bb9a-87f8afb7292a"
learn.fit_one_cycle(5, lr_max=slice(3e-6, 3e-4))
# + [markdown] colab_type="text" id="GwUx8fYdFq7o"
# So, we acheived a final accuracy of 98.5% in 10 epochs, taking less then 9 minutes training overall using fastai.
# + [markdown] colab_type="text" id="tDYMFXXU_WHH"
# # PyTorch
#
# Now we have our fastai benchmark, we'll run through how I tried to recreate this performance (or as close as we can!) within pure PyTorch in the same amount of epochs. The following code is partially based on the following tutorials:
#
# https://pytorch.org/tutorials/beginner/transfer_learning_tutorial.html
#
# https://github.com/Paperspace/PyTorch-101-Tutorial-Series
# + [markdown] colab_type="text" id="WNgkdmS9_WHI"
# ## Define Transforms
#
# Trying to recreate the transforms within the fastai vision learner is extremely hard to do, as the `batch_tfms` take place on the GPU within fastai, but take place on the CPU in Pytorch and in sequential steps.
#
# >note: Having tried many different options for the image augmentations, I settled on those detailed in the code below. It was interesting to note the huge slowdown in Google Colab created by using the `transforms.Resize` function to try to recreate the "presizing" of the fastai transforms mentioned above. The following code took over 14 minutes per 5 epochs and produced no benefit over the final transforms with it:
#
#
# ```python
# # 0.980 accuracy and slow training (14mins / 5 epochs)
# data_transforms = {
# 'train': transforms.Compose([
# transforms.Resize(460),
# transforms.RandomPerspective(0.2),
# transforms.RandomResizedCrop(224),
# transforms.ColorJitter(brightness=0.2, contrast=0.2),
# transforms.ToTensor(),
# transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225]) # imagenet stats
# ]),
# 'val': transforms.Compose([
# transforms.Resize(256),
# transforms.CenterCrop(224),
# transforms.ToTensor(),
# transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
# ]),
# }
# ```
#
# + colab={} colab_type="code" id="ri5qL1te_WHI"
## Final transforms: 98.0% in 5mins / 5 epochs
data_transforms = {
'train': transforms.Compose([
transforms.RandomHorizontalFlip(),
transforms.RandomPerspective(0.2),
transforms.RandomResizedCrop(224, scale=(0.08, 0.75)),
transforms.ColorJitter(brightness=0.2, contrast=0.2),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225]) # imagenet stats
]),
'val': transforms.Compose([
transforms.Resize(256),
transforms.CenterCrop(224),
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225])
]),
}
# + [markdown] colab_type="text" id="-6VfoRiB_WHM"
# ## Define Datasets and DataLoaders
# + [markdown] colab_type="text" id="yC_uemoJ_WHN"
# ### Custom dataset method
#
# We could define our own custom dataset as shown for comparison, however, for convenience we'll use the built in `ImageFolder` method within PyTorch's `datasets` class (further below)
# + colab={} colab_type="code" id="QxuHM6mf_WHO"
class imagenetteDataset(torch.utils.data.Dataset):
def __init__(self, data_dir, data_size = 0, transforms = None):
files = data_dir.glob('**/*')
files = [x for x in files if x.is_file()]
if data_size < 0 or data_size > len(files):
assert("Data size should be between 0 to number of files in the dataset")
if data_size == 0:
data_size = len(files)
self.data_size = data_size
self.files = random.sample(files, self.data_size)
self.transforms = transforms
def __len__(self):
return self.data_size
def __getitem__(self, idx):
image_address = self.files[idx]
image = Image.open(image_address)
if self.transforms==None:
print("No transforms defined, please define and re-run.")
elif self.transforms:
image = self.transforms(image)
label = image_address.parent.name
return image, label
# + [markdown] colab_type="text" id="CWr_p0Gq_WHT"
# ### dataset.ImageFolder method
#
# https://pytorch.org/docs/stable/torchvision/datasets.html#imagefolder
#
# This is a much more straightforward pipeline to implement, but with extra convenience attributes and methods over the above custom dataset e.g. `.classes`, for datasets in the following format:
#
# ```python
# root/dog/xxx.png
# root/dog/xxy.png
# root/dog/xxz.png
# ```
# + colab={} colab_type="code" id="JL5kofrM_WHU"
image_datasets = {x: datasets.ImageFolder(os.path.join(data_dir, x),
data_transforms[x])
for x in ['train', 'val']}
dataloaders = {x: torch.utils.data.DataLoader(image_datasets[x],
batch_size=64,
shuffle=True, num_workers=4)
for x in ['train', 'val']}
dataset_sizes = {x: len(image_datasets[x]) for x in ['train', 'val']}
class_names = image_datasets['train'].classes
num_classes = len(image_datasets['train'].classes)
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# + [markdown] colab_type="text" id="J75D31fH_WHX"
# We can now view information about our dataset:
# + colab={"base_uri": "https://localhost:8080/", "height": 221} colab_type="code" id="02ghkSEb_WHY" outputId="36d4e900-f4f3-452d-b0b0-a495012c6f79"
# See information about our train dataset
image_datasets['train']
# + colab={"base_uri": "https://localhost:8080/", "height": 187} colab_type="code" id="qMUKZGEm_WHc" outputId="05bc7da7-f4d2-4364-f727-1db7ec48afa8"
# See information about our validation dataset
image_datasets['val']
# + colab={"base_uri": "https://localhost:8080/", "height": 408} colab_type="code" id="TWHUx4eS_WHh" outputId="ebe08a29-41d7-44b8-8ffa-895984d1c50d"
# view an example data/label pair
image_datasets['train'][0]
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="tDZi2Hl4_WHl" outputId="e36150aa-8f29-435a-b354-41ccee6cd60c"
image_datasets['train'][0][0].shape
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="0di8anYN_WHo" outputId="7ffd43f9-c583-4b86-dac1-7263a2bf1536"
# Get the number of classes
class_names, num_classes
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="V08hORjC_WHs" outputId="0497b6de-2359-423d-9804-eeaeaa36cb54"
# see total num training batches
len(dataloaders['train'])
# + [markdown] colab_type="text" id="o18ieEDb_WHx"
# Check a a batch of our target values:
# + colab={} colab_type="code" id="s50NtluT_WHy"
inputs, targets = next(iter(dataloaders['train']))
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="hFbOmjaY_WH1" outputId="ef4c374d-3c84-4682-ba48-9103deb0ef99"
targets
# + [markdown] colab_type="text" id="AIcJKs9K_WH5"
# ## Visualize a batch of images
#
# We can see below that there were some interpolation artifacts from the default PyTorch transforms:
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="ofbU5viO_WH5" outputId="5f6484cd-b21b-4718-d426-d3bcdaacc119"
def imshow(inp, title=None):
"""Imshow for Tensor."""
inp = inp.numpy().transpose((1, 2, 0))
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
plt.figure(figsize = (10,6))
plt.imshow(inp)
if title is not None:
plt.title(title)
plt.pause(0.001) # pause a bit so that plots are updated
# Get a batch of training data
inputs, classes = next(iter(dataloaders['train']))
# Choose number items to display
n_items = 4
# Make a grid from batch
out = torchvision.utils.make_grid(inputs[:n_items])
imshow(out, title=[image_datasets['train'].classes[x] for x in classes[:n_items]])
# + [markdown] colab_type="text" id="BzBtw07y_WH9"
# ## Define Model
#
# As per the fastai, we'll use a pretrained model as a feature extractor, by freezing it, adding a custom head and training that first. Then, we'll fine-tune the main pretrained network as well as the custom head.
#
# As mentioned before, fastai implements an `AdaptiveConcatPool2d` layer, so to try to be as faithful as I could, I used the implementation below:
# + colab={} colab_type="code" id="azM4bQj4_WH9"
# fastai adaptive concat pool layer
class AdaptiveConcatPool2d(nn.Module):
"Layer that concats `AdaptiveAvgPool2d` and `AdaptiveMaxPool2d`"
def __init__(self, size=None):
super(AdaptiveConcatPool2d, self).__init__()
self.size = size or 1
self.ap = nn.AdaptiveAvgPool2d(self.size)
self.mp = nn.AdaptiveMaxPool2d(self.size)
def forward(self, x):
return torch.cat([self.mp(x), self.ap(x)], 1)
# -
# I then combined the pretrained model, new custom head with `AdaptiveConcatPool2d` layer in the `nn.Module`.
#
# In addition, I defined some freezing and unfreezing functions to allow a recreation for the staged-training method of fastai training then fine-tuning. The `freeze` function also avoids freezing the `BatchNorm` layers as per fastai.
#
# We can see below the module in the `create_model_body` function how the pretrained network is cut at the final layers to allow for the placement of the new custom head, in the same way as fastai does.
#
# The finally, we instantiate the complete model.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="S6_8y7oN_WIC" outputId="fd24342e-c3e4-4ebc-ec66-d68f04795b94"
class transferNet(nn.Module):
def __init__(self, model=None, num_classes=0):
super(transferNet, self).__init__()
# fastai equivalent pretrained body
self.model_body = model
# fastai equivalent head
self.model_head = nn.Sequential(AdaptiveConcatPool2d(),
nn.Flatten(),
nn.BatchNorm1d(1024, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True),
nn.Dropout(p=0.25, inplace=False),
nn.Linear(in_features=1024, out_features=512, bias=False),
nn.ReLU(inplace=True),
nn.BatchNorm1d(512, eps=1e-05, momentum=0.1, affine=True, track_running_stats=True),
nn.Dropout(p=0.5, inplace=False),
nn.Linear(in_features=512, out_features=num_classes, bias=False),
)
def freeze(self):
for name, param in self.model_body.named_parameters():
if("bn" not in name):
param.requires_grad = False
def unfreeze(self):
for param in self.parameters():
param.requires_grad = True
def forward(self, x):
out = self.model_body(x)
out = self.model_head(out)
return out
# create our the body of our model by cutting pretrained network
def create_model_body(model, cut=-2):
model_body_modules = list(model.children())[:cut]
return nn.Sequential(*model_body_modules)
model_body = create_model_body(model=torchvision.models.resnet18(pretrained=True))
# Now create our network
net = transferNet(model=model_body,
num_classes=num_classes)
net
# + [markdown] colab_type="text" id="hhRazyZX_WIG"
# ## Training
# + [markdown] colab_type="text" id="mKw9FrlHG1jX"
#
# ### Stage 1 - Base Model Frozen
#
# Now that we've defined our dataset, dataloader and transforms to be applied, we can define the training loop.
#
# Now let's freeze our base network so that we are only training the head:
# + colab={} colab_type="code" id="u__dauNr_WIH"
net.freeze()
# + [markdown] colab_type="text" id="s9T8Vbnf_WIK"
# We finally train for X epochs, ensuring we pass the same amount of epochs to the `OneCycleLR` scheduler.
# We evaluate classification accuracy every epoch.
# + colab={} colab_type="code" id="bnr9NggD_WIK"
def train_model(model, criterion, optimizer, scheduler, num_epochs=25):
since = time.time()
best_model_wts = copy.deepcopy(model.state_dict())
best_acc = 0.0
for epoch in range(num_epochs):
print(f'Epoch {epoch + 1}/{num_epochs}')
print('-' * 10)
# Each epoch has a training and validation phase
for phase in ['train', 'val']:
if phase == 'train':
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
running_loss = 0.0
running_corrects = 0
# Iterate over data.
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
# zero the parameter gradients
optimizer.zero_grad()
# forward
# track history if only in train
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
_, preds = torch.max(outputs, 1)
loss = criterion(outputs, labels)
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
running_loss += loss.item() * inputs.size(0)
running_corrects += torch.sum(preds == labels.data)
if phase == 'train':
scheduler.step()
epoch_loss = running_loss / dataset_sizes[phase]
epoch_acc = running_corrects.double() / dataset_sizes[phase]
print(f'{phase} Loss: {epoch_loss:.4f} Acc: {epoch_acc:.4f}')
# deep copy the model
if phase == 'val' and epoch_acc > best_acc:
best_acc = epoch_acc
best_model_wts = copy.deepcopy(model.state_dict())
print()
time_elapsed = time.time() - since
print(f'Training complete in {time_elapsed // 60:.0f}m {time_elapsed % 60:.0f}s')
print(f'Best val Acc: {best_acc:4f}')
# load best model weights
model.load_state_dict(best_model_wts)
return model
# + [markdown] colab_type="text" id="nPuKywFp_WIP"
# Now let's define the following elements before we train:
#
# * the Loss Function - the model's method of determining how well it is predicting and what it will try to minimise (`CrossEntropyLoss` as per fastai's `CrossEntropyLossFlat`)
# * the Optimizer - what will optimise the loss function by gradient descent (Adam as per fastai)
# * A scheduler to step the learning rate as we train - in this case the PyTorch implementation of the one-cycle learning rate policy, with the same maximum learning rate as that defined in our fastai model.
# + colab={} colab_type="code" id="TIe5vOGT_WIQ"
# First let's ensure the model is put onto the GPU
if cuda_available:
net = net.cuda()
# Now we define our loss function criterion
criterion = nn.CrossEntropyLoss()
# Define our optimizer by passing the network's parameters
optimizer = optim.Adam(net.parameters())
# Define our scheduler and training num epochs for one-cycle policy
num_epochs = 5
lr_scheduler = torch.optim.lr_scheduler.OneCycleLR(optimizer,
max_lr=3e-3,
steps_per_epoch=len(dataloaders['train']),
epochs=num_epochs)
# + [markdown] colab_type="text" id="XocOm6UD_WIT"
# And now we can train the head of our model:
# + colab={"base_uri": "https://localhost:8080/", "height": 476} colab_type="code" id="kn0kaeY2_WIU" outputId="e26f9646-6ce7-4149-8a89-ce55f0b864bd"
model_conv = train_model(net, criterion, optimizer,
lr_scheduler, num_epochs=num_epochs)
# + [markdown] colab_type="text" id="WRwDqdPL_WIW"
# ### Stage 2 - Unfreeze and train body
#
# Now that we've trained the head, we can unfreeze our pretrained base network and train that too. However, as per fastai, we'll try to use discriminative learning rates.
#
# First, we'll unfreeze the network:
# + colab={} colab_type="code" id="6CnHFXDh_WIX"
net.unfreeze()
# + [markdown] colab_type="text" id="xpsqIcJ-_WIa"
# Now we can set the discriminative learning rates. We'll need to define two separate optimizers and lr schedulers, for the parameters of the network we want to train with different rates - i.e. the pretrained body and the custom head:
# + colab={} colab_type="code" id="XXFLFuBE_WIb"
# Define two optimizers by passing the network's body and head parameters separately
optimizer = optim.Adam([{"params": net.model_body.parameters()},
{"params": net.model_head.parameters()}])
# Define our scheduler max_lr for one-cycle policy for the head and body separately in a list
num_epochs = 5
lr_scheduler = torch.optim.lr_scheduler.OneCycleLR(optimizer,
max_lr=[3e-6, 3e-4],
steps_per_epoch=len(dataloaders['train']),
epochs=num_epochs)
# + colab={"base_uri": "https://localhost:8080/", "height": 476} colab_type="code" id="dZHltoHt_WIe" outputId="e290d8b5-4533-44aa-a815-f4eac257630d"
model_conv = train_model(net, criterion, optimizer,
lr_scheduler, num_epochs=num_epochs)
# + [markdown] colab_type="text" id="X9sRjdKg_WIh"
#
# ## Results
#
# Using the straight PyTorch transfer learning method we acheived a final validation accuracy of: 97.9% that took around 11.5 minutes. Not bad, but not quite fastai standard! This 0.6% shortfall is likely due to the difference in the image transforms, which is definitely one of the main highlights of using of the fastai library.
#
# In [Part 2](https://mikful.github.io/blog/fastai/jupyter/pytorch/pytorch-lightning/2020/09/21/fastai-pytorch-lightning-part2.html) of this blog series we'll simplify the PyTorch pipeline into Pytorch Lighting modules.
| _notebooks/2020-09-21-fastai-pytorch-lightning-part1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="Y0eq2YR2BU2h"
from tkinter import *
# + id="zFP7g60XIm1X"
class Calculator:
def __init__(self, master):
self.master = master
master.title("Python Calculator")
# + [markdown] id="_lODJEPgLXu-"
# ## We are going to create our screen in this step.
# + id="sYWixCSSIqUG"
# create screen widget
self.screen = Text(master, state='disabled', width=30, height=3,background="white", foreground="blue")
# position screen in window
self.screen.grid(row=0,column=0,columnspan=4,padx=5,pady=5)
self.screen.configure(state='normal')
# initialize screen value as empty
self.equation = ''
# + [markdown] id="ItQ1lFIJLuwu"
# I am use the Text class from the Tkinter module as our screen. It takes height and width and background. These properties determine the appearance of our screen. bcoz of experiment with different dimensions and background colors to get familiar with it.
# + id="GARVOsMYKKY4"
# create buttons using method createButton
b1 = self.createButton(7)
b2 = self.createButton(8)
b3 = self.createButton(9)
b4 = self.createButton(u"\u232B",None)
b5 = self.createButton(4)
b6 = self.createButton(5)
b7 = self.createButton(6)
b8 = self.createButton(u"\u00F7")
b9 = self.createButton(1)
b10 = self.createButton(2)
b11 = self.createButton(3)
b12 = self.createButton('*')
b13 = self.createButton('.')
b14 = self.createButton(0)
b15 = self.createButton('+')
b16 = self.createButton('-')
b17 = self.createButton('=',None,34)
# + [markdown] id="XkfvOR-OMCzO"
# # to add buttons to the calculator board, so that can I , calculate numbers.
# createButton makes use of the Tkinter Button widget
# + id="FoZk5_rtKO7_"
# buttons stored in list
buttons = [b1,b2,b3,b4,b5,b6,b7,b8,b9,b10,b11,b12,b13,b14,b15,b16,b17]
# + [markdown] id="xJZGUJbJMsZd"
# after creating all our buttons using createButton, we store it in a list so i can iterate through it
# + [markdown] id="5Xfs6r_6M_Lf"
# range 1,5 for rows because we are trying to keep only 4 buttons on a row, apart from the = button.
# + id="lSFuWXEdKRdg"
# intialize counter
count = 0
# arrange buttons with grid manager
for row in range(1,5):
for column in range(4):
buttons[count].grid(row=row,column=column)
count += 1
# arrange last button '=' at the bottom
buttons[16].grid(row=5,column=0,columnspan=4)
# + id="Quk6h2kZKc5X"
def createButton(self,val,write=True,width=7):
# this function creates a button, and takes one compulsory argument, the value that should be on the button
return Button(self.master, text=val,command = lambda: self.click(val,write), width=width)
# + id="eSF0ZP7_KhW3"
def click(self,text,write):
# this function handles what happens when you click a button
# 'write' argument if True means the value 'val' should be written on screen, if None, should not be written on screen
if write == None:
# + id="OLna03hMKsTn"
#only evaluate code when there is an equation to be evaluated
if text == '=' and self.equation:
# replace the unicode value of division ./.with python division symbol / using regex
self.equation= re.sub(u"\u00F7", '/', self.equation)
print(self.equation)
answer = str(eval(self.equation))
self.clear_screen()
self.insert_screen(answer,newline=True)
elif text == u"\u232B":
self.clear_screen()
else:
# add text to screen
self.insert_screen(text)
# + id="UldHIJ_0Kz-f"
def clear_screen(self):
#to clear screen
#set equation to empty before deleting screen
self.equation = ''
self.screen.configure(state='normal')
self.screen.delete('1.0', END)
def insert_screen(self, value,newline=False):
self.screen.configure(state='normal')
self.screen.insert(END,value)
# record every value inserted in screen
self.equation += str(value)
self.screen.configure(state ='disabled')
root = Tk()
my_gui = Calculator(root)
root.mainloop()
# + id="_9VyhyiDRyuL"
| calculator (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="6XvCUmCEd4Dm"
# # TensorFlow Datasets
#
# TensorFlow Datasets provides a collection of datasets ready to use with TensorFlow. It handles downloading and preparing the data and constructing a [`tf.data.Dataset`](https://www.tensorflow.org/api_docs/python/tf/data/Dataset).
# + [markdown] colab_type="text" id="J8y9ZkLXmAZc"
# Copyright 2018 The TensorFlow Datasets Authors, Licensed under the Apache License, Version 2.0
# + [markdown] colab_type="text" id="OGw9EgE0tC0C"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/datasets/overview"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/datasets/blob/master/docs/overview.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/datasets/blob/master/docs/overview.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="_7hshda5eaGL"
# ## Installation
#
# `pip install tensorflow-datasets`
#
# Note that `tensorflow-datasets` expects you to have TensorFlow already installed, and currently depends on `tensorflow` (or `tensorflow-gpu`) >= `1.14.0`.
# + cellView="both" colab_type="code" id="boeZp0sYbO41" colab={}
# !pip install -q tensorflow tensorflow-datasets matplotlib
# + colab_type="code" id="TTBSvHcSLBzc" colab={}
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import matplotlib.pyplot as plt
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
# + [markdown] colab_type="text" id="8-ZBEd6Ie28N"
# ## Eager execution
#
# TensorFlow Datasets is compatible with both TensorFlow [Eager mode](https://www.tensorflow.org/guide/eager) and Graph mode. For this colab, we'll run in Eager mode.
# + colab_type="code" id="o9H2EiXzfNgO" colab={}
tf.enable_eager_execution()
# + [markdown] colab_type="text" id="VZZyuO13fPvk"
# ## List the available datasets
#
# Each dataset is implemented as a [`tfds.core.DatasetBuilder`](https://www.tensorflow.org/datasets/api_docs/python/tfds/core/DatasetBuilder) and you can list all available builders with `tfds.list_builders()`.
#
# You can see all the datasets with additional documentation on the [datasets documentation page](https://github.com/tensorflow/datasets/blob/master/docs/datasets.md).
# + colab_type="code" id="FAvbSVzjLCIb" outputId="79f354d8-746a-4fac-ab34-86225904ee46" colab={"base_uri": "https://localhost:8080/", "height": 181}
tfds.list_builders()
# + [markdown] colab_type="text" id="VjI6VgOBf0v0"
# ## `tfds.load`: A dataset in one line
#
# [`tfds.load`](https://www.tensorflow.org/datasets/api_docs/python/tfds/load) is a convenience method that's the simplest way to build and load and `tf.data.Dataset`.
#
# Below, we load the MNIST training data. Setting `download=True` will download and prepare the data. Note that it's safe to call `load` multiple times with `download=True` as long as the builder `name` and `data_dir` remain the same. The prepared data will be reused.
# + colab_type="code" id="dCou80mnLLPV" outputId="0ee326c1-ea25-4c4b-b3dc-a7c29e9b2156" colab={"base_uri": "https://localhost:8080/", "height": 108}
mnist_train = tfds.load(name="mnist", split=tfds.Split.TRAIN)
assert isinstance(mnist_train, tf.data.Dataset)
mnist_train
# + [markdown] id="D4A6yp_Ii3G-" colab_type="text"
# When loading a dataset, the canonical default version is used. It is however recommended to specify the major version of the dataset to use, and to advertise which version of the dataset was used in your results. See the
# [documentation on datasets versioning](https://github.com/tensorflow/datasets/blob/master/docs/) for more details.
# + id="OCXCz-vhj0kE" colab_type="code" colab={}
mnist = tfds.load("mnist:1.*.*")
# + [markdown] colab_type="text" id="u-GAxR79hGTr"
# ## Feature dictionaries
#
# All `tfds` datasets contain feature dictionaries mapping feature names to Tensor values. A typical dataset, like MNIST, will have 2 keys: `"image"` and `"label"`. Below we inspect a single example.
# + colab_type="code" id="YHE21nkHLrER" outputId="2cae50b0-4f36-44b3-d858-71c36403826a" colab={"base_uri": "https://localhost:8080/", "height": 365}
mnist_example, = mnist_train.take(1)
image, label = mnist_example["image"], mnist_example["label"]
plt.imshow(image.numpy()[:, :, 0].astype(np.float32), cmap=plt.get_cmap("gray"))
print("Label: %d" % label.numpy())
# + [markdown] colab_type="text" id="EW-kEK_mhbhy"
# ## `DatasetBuilder`
#
# `tfds.load` is really a thin conveninence wrapper around `DatasetBuilder`. We can accomplish the same as above directly with the MNIST `DatasetBuilder`.
# + colab_type="code" id="9FDDJXmBhpQ4" outputId="2be38dd6-1aa3-4b63-fe8a-f4668147fb14" colab={"base_uri": "https://localhost:8080/", "height": 54}
mnist_builder = tfds.builder("mnist")
mnist_builder.download_and_prepare()
mnist_train = mnist_builder.as_dataset(split=tfds.Split.TRAIN)
mnist_train
# + [markdown] colab_type="text" id="7tlVOAzLlKqc"
# ## Input pipelines
#
# Once you have a `tf.data.Dataset` object, it's simple to define the rest of an input pipeline suitable for model training by using the [`tf.data` API](https://www.tensorflow.org/guide/datasets).
#
# Here we'll repeat the dataset so that we have an infinite stream of examples, shuffle, and create batches of 32.
# + colab_type="code" id="9OQZqGZMlSE8" colab={}
mnist_train = mnist_train.repeat().shuffle(1024).batch(32)
# prefetch will enable the input pipeline to asynchronously fetch batches while
# your model is training.
mnist_train = mnist_train.prefetch(tf.data.experimental.AUTOTUNE)
# Now you could loop over batches of the dataset and train
# for batch in mnist_train:
# ...
# + [markdown] colab_type="text" id="uczpNuc_A7wE"
# ## DatasetInfo
#
# After generation, the builder contains useful information on the dataset:
# + colab_type="code" id="mSamfFznA9Ph" outputId="a3203f4b-a1ca-4290-c7da-d4a148096807" colab={"base_uri": "https://localhost:8080/", "height": 272}
info = mnist_builder.info
print(info)
# + [markdown] id="cspsneov2VbC" colab_type="text"
# `DatasetInfo` also contains useful information about the features:
# + id="u1wL14QH2TW1" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 72} outputId="262f4857-7253-4a16-af00-1ff404b69963"
print(info.features)
print(info.features["label"].num_classes)
print(info.features["label"].names)
# + [markdown] colab_type="text" id="xbrm0WBmBLEI"
# You can also load the `DatasetInfo` directly with `tfds.load` using `with_info=True`.
# + colab_type="code" id="tvZYujQwBL7B" outputId="46fbcb43-697d-460b-da80-02e58bed8b3e" colab={"base_uri": "https://localhost:8080/", "height": 290}
dataset, info = tfds.load("mnist", split="test", with_info=True)
print(info)
| docs/overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Simulation Data
#
# Orbit provide the functions to generate the simulation data including:
#
# 1. Generate the data with time-series trend:
# - random walk
# - arima
# 2. Generate the data with seasonality
# - discrete
# - fourier series
# 3. Generate regression data
# +
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from orbit.utils.simulation import make_trend, make_seasonality, make_regression
plt.style.use('fivethirtyeight')
plt.rcParams['figure.figsize'] = [8, 8]
# %matplotlib inline
# -
# ## Trend
# ### Random Walk
rw = make_trend(200, rw_loc=0.02, rw_scale=0.1, seed=2020)
plt.plot(rw);
# ### ARMA
#
# reference for the ARMA process: https://www.statsmodels.org/stable/generated/statsmodels.tsa.arima_process.ArmaProcess.html
arma_trend = make_trend(200, method='arma', arma=[.8, -.1], seed=2020)
plt.plot(arma_trend);
# ## Seasonality
# ### Discrete
# generating a weekly seasonality(=7) where seasonality wihtin a day is constant(duration=24) on an hourly time-series
ds = make_seasonality(500, seasonality=7, duration=24, method='discrete', seed=2020)
plt.plot(ds);
# ### Fourier
# generating a sine-cosine wave seasonality for a annual seasonality(=365) using fourier series
fs = make_seasonality(365 * 3, seasonality=365, method='fourier', order=5, seed=2020)
plt.plot(fs);
fs
# ## Regression
# generating multiplicative time-series with trend, seasonality and regression components
# define the regression coefficients
coefs = [0.1, -.33, 0.8]
x, y, coefs = make_regression(200, coefs, scale=2.0, seed=2020)
plt.plot(y);
# check if get the coefficients as set up
reg = LinearRegression().fit(x, y)
print(reg.coef_)
| examples/utilities_simulation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + hideCode=true hideOutput=false
import os, sys
try:
from synapse.lib.jupyter import *
import synapse.common as s_common
import synapse.lib.parser as s_parser
import synapse.lib.output as s_output
import synapse.tools.genpkg as s_t_genpkg
except ImportError as e:
# Insert the root path of the repository to sys.path.
# This assumes the notebook is located three directories away
# From the root synapse directory. It may need to be varied
synroot = os.path.abspath('../../../')
sys.path.insert(0, synroot)
from synapse.lib.jupyter import *
import synapse.common as s_common
import synapse.lib.parser as s_parser
import synapse.lib.output as s_output
import synapse.tools.genpkg as s_t_genpkg
# + hideCode=true
# Get a Cortex we can use for testing.
core = await getTempCoreCmdr()
# get the base dir for the demopkg
dpkg_fp = getDocPath('foopkg/foopkg.yml')
# + active=""
# .. highlight:: none
#
# .. _syn-tools-genpkg:
#
# Synapse Tools - genpkg
# ========================
#
# The Synapse ``genpkg`` tool can be used to generate a Storm :ref:`gloss-package` containing new Storm commands and Storm modules from a YAML definition and optionally push it to a Cortex or PkgRepo.
#
# Syntax
# ------
#
# ``genpkg`` is executed from an operating system command shell. The command usage is as follows:
#
# ::
#
# usage: synapse.tools.genpkg [-h] [--push <url>] [--save <path>] [--optic <path>] <pkgfile>
#
# Where:
#
# - ``pkgfile`` is the path to the Storm Package YAML file.
#
# - ``--save`` takes a file name to save the completed package JSON as.
#
# - ``--push`` takes an optional Telepath URL to a Cortex or PkgRepo for the package to be pushed to.
#
# - See :ref:`cortex-connect` for the format to specify a path to a Cortex.
# - Cortex paths can also be specified using aliases defined in the user's ``.syn/aliases.yaml`` file.
#
# - ``--optic`` takes an optional path to a directory containing Optic module files.
#
#
# Package Layout
# --------------
#
# The expected filesystem layout for a Storm package is:
#
# ::
#
# foopkg.yml
# storm/
# ├── commands/
# │ └── foocmd
# ├── modules/
# │ └── foomod
# └── optic/
# └── index.html
#
# Commands and modules defined in the package YAML file are expected to have corresponding files containing the Storm code for their implementation. It is not required to have both commands and modules in a Storm package; you may have a package with only commands, or only modules.
#
#
# Package YAML
# ------------
#
# A Storm package YAML may contain the following definitions:
#
# - ``name``: Name of the Storm package.
# - ``version``: Version of the Storm package. A Cortex may contain multiple versions of the same package.
# - ``synapse_minversion``: Optional minimum required Synapse version a Cortex must be running to load the package.
# - ``onload``: Optional Storm code to run in a Cortex when the package is loaded.
# - ``modules``: Storm module definitions.
# - ``commands``: Storm command definitions.
#
# The example below shows the YAML included in the ``foopkg.yml`` file.
#
# ``foopkg.yml``
# ++++++++++++++
# + hideCode=true
# Ensure the yaml content is valid
_contents = s_common.yamlload(dpkg_fp)
assert _contents.get('name') == 'foopkg'
with s_common.genfile(dpkg_fp) as fd:
buf = fd.read()
print(buf.decode())
# + active=""
# Modules
# -------
#
# Modules can be used to expose reusable Storm functions. Each module defines a ``name``, which is used for importing elsewhere via ``$lib.import()``, and optionally a ``modconf`` dictionary containing additional configuration values which will be accessible in the module's Storm via ``$modconf``.
#
# The example below shows the Storm code included in the ``foomod`` file.
#
#
# ``foomod``
# ++++++++++
#
# + hideCode=true
dpkg_foomod = getDocPath('foopkg/storm/modules/foomod')
with s_common.genfile(dpkg_foomod) as fd:
buf = fd.read()
storm = buf.decode()
# Ensure the package is valid storm
assert len(storm) > 1
_ = s_parser.parseQuery(storm)
print(storm)
# + active=""
# Commands
# --------
#
# Multiple Storm commands can be added to a Storm service package, with each defining the following attributes:
#
# - ``name``: Name of the Storm command to expose in the Cortex.
# - ``descr``: Description of the command which will be available in ``help`` displays.
# - ``asroot``: Whether the command should be run with root permissions. This allows users to be granted access to run the command without requiring them to have all the permissions needed by the Storm command. An example ``asroot`` permission for ``foocmd`` would be ``('storm', 'asroot', 'cmd', 'asroot', 'foocmd')``.
# - ``cmdargs``: An optional list of arguments for the command.
# - ``cmdconf``: An optional dictionary of additional configuration variables to provide to the command Storm execution.
# - ``forms``: List of input and output forms for the command, as well as a list of nodedata keys and the corresponding form on which they may be set by the service.
#
#
# The example below shows the Storm code included in the ``foocmd`` file.
#
# ``foocmd``
# ++++++++++
# + hideCode=true
dpkg_foocmd = getDocPath('foopkg/storm/commands/foocmd')
with s_common.genfile(dpkg_foocmd) as fd:
buf = fd.read()
storm = buf.decode()
# Ensure the package is valid storm
assert len(storm) > 1
_ = s_parser.parseQuery(storm)
print(storm)
# + active=""
#
# Building the Example Package
# ----------------------------
#
# To build the package and push it directly to a Cortex:
#
# ::
#
# python -m synapse.tools.genpkg --push tcp://user:pass@127.0.0.1:27492 foopkg.yml
#
#
# .. NOTE::
# Users must have the ``pkg.add`` permission to add a package to a Cortex.
#
# Once the package has been successfully pushed to the Cortex, the additional Storm Commands will be listed in the output of ``storm help`` under the package they were loaded from:
#
# ::
#
# package: foopkg
# foocmd : One line description on the first line.
#
#
# The new commands may now be used like any other Storm command:
#
# + hideCode=true
# Load the package into the Cortex
lurl = core.core._core.getLocalUrl()
args = ['--push', lurl, dpkg_fp]
outp = s_output.OutPutStr()
retn = await s_t_genpkg.main(args, outp)
assert retn == 0
_ = await core.eval('syn:cmd:package=foopkg', num=1)
_ = await core.eval('[inet:ipv4=192.168.0.113 :asn=20]')
podes = await core.eval('inet:ipv4=192.168.0.113 | foocmd', cmdr=True)
assert podes[0][1]['props'].get('asn') == 40
# + active=""
# If immediately pushing the package to a Cortex is not desired, it can instead be built and saved to ``foo.json`` to load later:
#
# ::
#
# python -m synapse.tools.genpkg --save foo.json foopkg.yml
# + hideCode=true hideOutput=true
# Setup the directory we'll pave the storm package contents into
coredirn = core.core._core.dirn
save_fp = os.path.join(coredirn, 'save.json')
args = ['--save', save_fp, dpkg_fp]
outp = s_output.OutPutStr()
retn = await s_t_genpkg.main(args, outp)
assert retn == 0
data = s_common.jsload(save_fp)
assert data.get('name') == 'foopkg'
assert 'commands' in data
assert 'modules' in data
# + hideCode=true hideOutput=true
await core.fini()
| docs/synapse/userguides/syn_tools_genpkg.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import Stemmer
stemmer = Stemmer.Stemmer('english')
import numpy as np
train=np.load('./datasets/processed/train-restaurant.npy',allow_pickle=True)
test=np.load('./datasets/processed/test-restaurant.npy',allow_pickle=True)
train.shape
test.shape
alldata=np.concatenate((train,test))
alldata.shape
target_list=[]
stem_list=[]
for data in alldata:
target_list.append(data['text_aspect'].strip().split()[-1])
s=stemmer.stemWord(data['text_aspect'].strip().split()[-1])
stem_list.append(s)
stem_set=set(stem_list)
stem_set
# import hash
hash_dict=dict()
hash_dict2=dict()
for i in range(len(alldata)):
text_raw=alldata[i]['text_raw']
target_stem=alldata[i]['text_aspect'].strip().split()[-1]
text_hash=hash(text_raw)
key=text_hash
if key not in hash_dict.keys():
hash_dict[key]=[i]
# print(type(target_stem))
hash_dict2[key]=set([target_stem])
else:
l=hash_dict[key]
l.append(i)
hash_dict[key]=l
s=hash_dict2[key]
print(target_stem)
s.add(str(target_stem))
hash_dict2[key]=s
# +
import random
ratio=0.6
train_target_set=set()
test_target_set=set()
processed=set()
train_val_idx=[]
test_idx=[]
for i in range(len(alldata)):
text_raw=alldata[i]['text_raw']
key=hash(text_raw)
if i in processed:
continue
target_stem=alldata[i]['text_aspect'].strip().split()[-1]
if target_stem in train_target_set:
# add to train
all_idx=hash_dict[key]
all_target=hash_dict2[key]
for idx in all_idx:
train_val_idx.append(idx)
processed.add(idx)
for target in all_target:
train_target_set.add(target)
elif target_stem in test_target_set:
all_idx=hash_dict[key]
all_target=hash_dict2[key]
for idx in all_idx:
test_idx.append(idx)
processed.add(idx)
for target in all_target:
test_target_set.add(target)
else:
# flip
ret=random.random()
if ret<ratio:
# add to train
all_idx=hash_dict[key]
all_target=hash_dict2[key]
for idx in all_idx:
train_val_idx.append(idx)
processed.add(idx)
for target in all_target:
train_target_set.add(target)
else:
# # add to test
all_idx=hash_dict[key]
all_target=hash_dict2[key]
for idx in all_idx:
test_idx.append(idx)
processed.add(idx)
for target in all_target:
test_target_set.add(target)
print(len(train_val_idx),len(test_idx),len(processed),(len(train_val_idx)+len(test_idx))==len(processed))
# -
np.save('./datasets/resplit/train-val-restaurant.npy',alldata[train_val_idx])
np.save('./datasets/resplit/test-restaurant.npy',alldata[test_idx])
train_val=np.load('./datasets/resplit/train-val-restaurant.npy',allow_pickle=True)
target_list=[]
stem_list=[]
for data in alldata:
target_list.append(data['text_aspect'].strip().split()[-1])
s=stemmer.stemWord(data['text_aspect'].strip().split()[-1])
stem_list.append(s)
stem_set=set(stem_list)
stem_set
# import hash
hash_dict=dict()
hash_dict2=dict()
for i in range(len(train_val)):
text_raw=alldata[i]['text_raw']
target_stem=alldata[i]['text_aspect'].strip().split()[-1]
text_hash=hash(text_raw)
key=text_hash
if key not in hash_dict.keys():
hash_dict[key]=[i]
# print(type(target_stem))
hash_dict2[key]=set([target_stem])
else:
l=hash_dict[key]
l.append(i)
hash_dict[key]=l
s=hash_dict2[key]
print(target_stem)
s.add(str(target_stem))
hash_dict2[key]=s
# +
import random
ratio=0.6
train_target_set=set()
val_target_set=set()
processed=set()
train_idx=[]
val_idx=[]
for i in range(len(train_val)):
text_raw=alldata[i]['text_raw']
key=hash(text_raw)
if i in processed:
continue
target_stem=alldata[i]['text_aspect'].strip().split()[-1]
if target_stem in train_target_set:
# add to train
all_idx=hash_dict[key]
all_target=hash_dict2[key]
for idx in all_idx:
train_idx.append(idx)
processed.add(idx)
for target in all_target:
train_target_set.add(target)
elif target_stem in val_target_set:
all_idx=hash_dict[key]
all_target=hash_dict2[key]
for idx in all_idx:
val_idx.append(idx)
processed.add(idx)
for target in all_target:
val_target_set.add(target)
else:
# flip
ret=random.random()
if ret<ratio:
# add to train
all_idx=hash_dict[key]
all_target=hash_dict2[key]
for idx in all_idx:
train_idx.append(idx)
processed.add(idx)
for target in all_target:
train_target_set.add(target)
else:
# # add to val
all_idx=hash_dict[key]
all_target=hash_dict2[key]
for idx in all_idx:
val_idx.append(idx)
processed.add(idx)
for target in all_target:
val_target_set.add(target)
print(len(train_idx),len(val_idx),len(processed),(len(train_idx)+len(val_idx))==len(processed))
# -
np.save('./datasets/resplit/train-restaurant.npy',train_val[train_idx])
np.save('./datasets/resplit/valid-restaurant.npy',train_val[val_idx])
alldata[0].keys()
import numpy as np
train=np.load('./datasets/resplit/train-restaurant.npy',allow_pickle=True)
test=np.load('./datasets/resplit/test-restaurant.npy',allow_pickle=True)
val=np.load('./datasets/resplit/valid-restaurant.npy',allow_pickle=True)
len(train)
len(test)
len(val)
| notebooks/resplit_restaurant.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import requests
import json
# 导入文件并转化为pandas DataFrame对象
excel_path = 'myrecords.xlsx'
d = pd.read_excel(excel_path, sheet_name='Sheet1') #文件中只有sheet1有数据,直接导入。
print(d.head(10)) #打印前10条记录
# -
MYKEY = '<KEY>'
base_url = 'https://apis.map.qq.com/ws/geocoder/v1/?address='
placetext = d['申请人地址']
url = base_url + placetext + '&key=' + MYKEY
url
def get_json(url):
try:
r = requests.get(url)
r.raise_for_status()
r.encoding = 'utf-8'
return r.text
except:
print('Get json Error.')
return "ERROR"
# +
urlist = list(url)
geolist = []
for li in urlist:
jdata = get_json(li)
jdict = json.loads(jdata)
geolist.append(jdict)
geolist
# -
len(geolist)
# +
resultlist = []
addresslist = []
locationlist = []
for geo in geolist:
try:
result = geo['result']
address = geo['result']['address_components']
location = geo['result']['location']
except:
result = None
address = {'city': None,
'district': None,
'province': None,
'street': None,
'street_number': None}
location = {'lat': None,'lng': None}
resultlist.append(result)
addresslist.append(address)
locationlist.append(location)
addressdf = pd.DataFrame(addresslist)
locationdf = pd.DataFrame(locationlist)
geodf = addressdf.join(locationdf)
geodf.fillna(value=np.nan, inplace=True)
geodf
# -
d = d.join(geodf)
d
d.to_excel('recordswithloc.xls',sheet_name='sheet1')
countrypd = pd.read_csv('country.csv')
countryli = list(countrypd['value'])
countryli
import jieba
placetext
# +
matched_placelist = []
for place in list(placetext):#地址匹配
status = 0
seg_place = list(jieba.cut(place,cut_all=True))
seg_place
for seg_word in seg_place:#分词匹配
if status == 0 and len(seg_word)>1:#判断是否已经匹配
for country in countryli:
if seg_word in country and status==0:
matched_placelist.append(country)
status = 1 #匹配成功
if status == 0:
matched_placelist.append(np.nan)
matched_placelist
# -
countrydf = pd.DataFrame(matched_placelist,columns = ['country'])
countrydf
d = d.join(countrydf)
d
d.to_excel('recordswithloc.xls',sheet_name='sheet2')
| myrecords_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Importing libraries and datasets
import pandas as pd
cl_cat = pd.read_csv('client_categories.csv', sep=';')
cl_lst = pd.read_csv('clients_last_2_fixed.csv', sep=';')
transac = pd.read_csv('transac_137k.csv', index_col = 0)
mcc = pd.read_excel('MCC_last.xlsx')
transac.head()
cl_lst.head()
# Посмотрим на категории товаров
mcc['category'].value_counts()
# # Обработка данных
cats = {}
for cat in mcc['category'].value_counts().index:
cats[cat] = list(mcc.loc[mcc.category == cat]['mcc'].values)
mcc_list = transac['mcc'].value_counts().index
for cat_num in mcc_list:
for cat in mcc['category'].value_counts().index:
if cat_num in cats[cat]:
transac.loc[transac.mcc == cat_num, 'mcc'] = cat
col = transac['mcc'].value_counts().index
for name in col:
transac[name] = 0
transac[name] = transac.loc[transac.mcc == name]['amount']
transac.fillna(0, inplace = True)
transac['Money'] = transac['Money']/1000
# Получим датасет, где видны общие затраты на категорию товаров для каждого клиента
transac = transac.groupby(['cnum']).sum().iloc[:,4:]
transac.head()
# Обработаем датасет с клиентами
cl = cl_lst.join(pd.get_dummies(cl_lst['gender']))
cl = cl.join(pd.get_dummies(cl_lst['married_']))
cl = cl.join(pd.get_dummies(cl_lst['residenttype']))
count = ['gender','married_','residenttype']
cl.drop(count, axis=1, inplace=True)
cl.set_index('cnum_', inplace=True)
cl.head()
transac['max_group'] = [transac.columns[i][0] for i in transac.values == transac.max(axis=1)[:,None]]
transac.head()
cl = cl.join(transac['max_group']).dropna()
cats = mcc['category'].value_counts().index
cl['mcc'] = 0
for i in range(len(cats)):
cl.loc[cl.max_group == cats[i], 'mcc'] = i
cl.drop('max_group', axis=1, inplace=True)
cl.head()
X = cl.iloc[:,:-1]
y = cl.iloc[:,-1:]
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2, random_state = 42)
# -
# # RandomForest
# +
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(max_depth = 15)
rfc.fit(X_train, y_train)
# -
rfc_pred = rfc.predict(X_test)
pd.DataFrame(rfc_pred)[0].value_counts()
# +
from sklearn.metrics import accuracy_score
accuracy_score(y_test, rfc_pred)
# -
| ml/Raif.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # I'm just gonna see if I can write a quick and dirty kepler-planet function thing
import matplotlib.pyplot as plt
# %matplotlib inline
filename = 'planets_2019.07.30_19.22.04.csv'
import pandas as pd
import numpy as np
planets = pd.read_csv(filename,
sep=",", comment="#")
# orig function:
def read_kepler_data(fileLocation):
names = ('RowID', 'SysName', 'planetLetter', 'NumberOfPlanets', 'Porb',
'ePorbU', 'ePorbL', 'a', 'ea', 'ecc',
'eeccU', 'eeccL', 'Incl', 'eInclU', 'eInclL',
'pMass', 'epMassU', 'epMassL', 'pMassType', 'sMass',
'esMass', 'sRadius', 'esRadiusU', 'esRadiusL', 'tTime',
'etTimeU', 'etTimeL')
formats = ('f4', 'S12', 'S2', 'f4', 'f4',
'f4', 'f4', 'f4', 'f4', 'f4',
'f4', 'f4', 'f4', 'f4', 'f4',
'f4', 'f4', 'f4', 'S8', 'f4',
'f4', 'f4', 'f4', 'f4', 'f4',
'f4', 'f4')
kepler_data = np.genfromtxt(fileLocation,
comments='#',
delimiter=',',
dtype={'names':names,
'formats':formats})
return kepler_data
planets['pl_hostname'].unique()
# +
# convert based on name:
name = 'ups And'
# only select entries with this name
mask = planets['pl_hostname'] == name
ecc = planets['pl_orbeccen'][mask]
a = planets['pl_orbsmax'][mask]
Porb = planets['pl_orbper'][mask]
Incl = planets['pl_orbincl'][mask]
sMass = planets['st_mass'][mask]
pMass = planets['pl_bmassj'][mask]
#tmax=np.array(tTime).max()
#tTime = []
#for t in planets['pl_orbtper'][mask]:
# tTime.append([tmax,t])
tTime = planets['pl_orbtper'][mask]
tTime
Porb
#ecc,a
# -
tTime
planets.keys()
# +
#planets['pl_orbtper']
# +
# # dict?
kepler_data = {}
#kepler_data['RowID'] = -1
kepler_data['SysName'] = name
#kepler_data['planetLetter'] = 'X'
kepler_data['NumberOfPlanets'] = len(a)
kepler_data['Porb'] = Porb.values
#kepler_data['ePorbU']=0
#kepler_data['ePorbL']=0
kepler_data['a']=a.values
#kepler_data['ea']=0
kepler_data['ecc']=ecc.values
#kepler_data['eeccU']=0
#kepler_data['eeccL']=0
kepler_data['Incl']=Incl.values
#kepler_data['eInclU']=0
kepler_data['pMass']=pMass.values
kepler_data['sMass']=sMass.values
kepler_data['tTime']=tTime.values
#'eInclL',
# 'pMass', 'epMassU', 'epMassL', 'pMassType', 'sMass',
# 'esMass', 'sRadius', 'esRadiusU', 'esRadiusL', 'tTime',
# 'etTimeU', 'etTimeL'
# -
kepler_data
from convert_kepler_data import convert_kepler_data
star_mass, planet_masses, planet_initial_position, planet_initial_velocity, ecc = convert_kepler_data(kepler_data)
star_mass
planet_masses
planet_initial_position
planet_initial_velocity
from kepler_data_table_to_new_planet_system import read_convert_new_system
filename = 'planets_2019.07.30_19.22.04.csv'
# What are the possible names?
planets['pl_hostname'].unique()
# We want one with a few planets. We can play with the table under the exoplanet arxiv: https://exoplanetarchive.ipac.caltech.edu/cgi-bin/TblView/nph-tblView?app=ExoTbls&config=planets
#
# Or lets try looking at some to see what they have in their entries:
# +
name = '11 Com' # try first one
mask = planets['pl_hostname'] == name
# how many planets? lets see how long the eccentricity list is:
planets['pl_orbeccen'][mask]
# -
# Well this one only has 1 planet, so let's try another name:
# +
name = 'ups And' # try first one
mask = planets['pl_hostname'] == name
# how many planets? lets see how long the eccentricity list is:
planets['pl_orbeccen'][mask]
# -
# Ok, this one has a few planets, so let's check it out.
new_kepler_data = read_convert_new_system(filename, 'ups And')
new_kepler_data
# We can convert this now just like we did before:
from convert_kepler_data import convert_kepler_data
star_mass, planet_mass, planet_initial_position, planet_initial_velocity, ecc = convert_kepler_data(kepler_data)
# Then we can use these again as inputs into our hermite solver.
from hermite_library import do_hermite
r_h, v_h, t_h, E_h = do_hermite(star_mass,
planet_mass,
planet_initial_position,
planet_initial_velocity,
tfinal=1e4*5000,
Nsteps=5000, threeDee=True)
# +
# we are doing 1 row of plots with 2 columns of plots
# and making sure that our plot is 2X as wide as long
fig, ax = plt.subplots(1,2, figsize=(6*2, 6))
for i in range(r_h.shape[0]): # loop over number of planets+star
ax[0].plot(r_h[i, 0, :], r_h[i, 1, :])
# making extra fancy with labels
ax[0].set_xlabel('x in AU')
ax[0].set_ylabel('y in AU')
# also set a different-than-default size
ax[0].set_xlim(-5, 5)
ax[0].set_ylim(-5, 5)
# now I'll make energy as a function of time
ax[1].plot(t_h, E_h)
ax[1].set_xlabel('Time in Seconds')
ax[1].set_ylabel('Energy, Normalized')
plt.show()
# -
| lesson08/_Trial_newKeplerPlanets_testFunction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Random Forest
# +
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.datasets import load_iris
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report, confusion_matrix
from sklearn.model_selection import train_test_split
# -
iris = load_iris()
# print(digits.DESCR)
dir(iris)
iris.target_names
df = pd.DataFrame(iris.data, columns=iris.feature_names)
df['target'] = iris.target
df['target_names'] = df['target'].apply(lambda i: iris.target_names[i])
df.head()
X_train, X_test, y_train, y_test = train_test_split(df.drop(['target', 'target_names'], axis=1), df.target, test_size=0.2)
model = RandomForestClassifier(n_estimators=40)
model.fit(X_train, y_train)
model.score(X_test, y_test)
model.predict(iris.data[0:5])
y_pred = model.predict(X_test)
print(classification_report(y_test, y_pred))
def print_cm(y_test, y_pred, figsize=None):
plt.figure(figsize=figsize)
cm = confusion_matrix(y_test, y_pred)
sns.heatmap(cm, annot=True)
plt.xlabel('Predicted')
plt.ylabel('Truth')
print_cm(y_test, y_pred)
| 7_random_forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import importlib
from tqdm import tqdm
import json
import pandas as pd
calculate_doc_term_recall = importlib.import_module('train-deep-ct').calculate_doc_term_recall
calculate_qtokens = importlib.import_module('train-deep-ct').calculate_qtokens
# !nosetests
from util import clean_qtokens
# +
DIR = '/mnt/ceph/storage/data-in-progress/data-teaching/theses/wstud-thesis-probst/deep-ct/'
anchorText = {}
try:
anchorText = clean_qtokens(json.load(open(DIR + 'ms-marco-trainig-set-deepct-qtokens.json', 'r')))
except Exception as e:
print(e)
print('Comment the following lines to create ms-marco-trainig-set-deepct-qtokens.json.')
#with open(DIR + 'msmarco-document-train.jsonl') as f:
# for l in tqdm(f):
# l = json.loads(l)
# for target_doc in l['target_document']:
# if target_doc not in anchorText:
# anchorText[target_doc] = []
# anchorText[target_doc] += [l['contents']]
#
#for target_doc in tqdm([i for i in anchorText.keys()]):
# anchorText[target_doc] = calculate_qtokens(anchorText[target_doc])
#json.dump(anchorText, open(DIR + 'ms-marco-trainig-set-deepct-qtokens.json', 'w'))
# -
len(anchorText)
[i for i in anchorText.keys()][:10]
anchorText['D59219']
with open(DIR + 'msmarco-document-passages.jsonl') as docs, open(DIR + 'deep-ct-training-data-ms-marco-training-set-test-overlap-removed.jsonl', 'w') as out_file:
for doc in tqdm(docs):
doc = json.loads(doc)
doc_id = doc['id']
if doc_id not in anchorText:
continue
for passage_id, passage in doc['passages']:
for i in calculate_doc_term_recall([{"doc":{"title": passage}}], anchorText[doc_id]):
i['doc']['position'] = passage_id
i['doc']['id'] = doc_id + '___' + str(passage_id)
out_file.write(json.dumps(i) + '\n')
# +
saved_docs = []
examples = 0
with open(DIR + 'deep-ct-training-data-ms-marco-training-set-test-overlap-removed.jsonl', 'r') as docs:
for l in docs:
l = json.loads(l)
examples += 1
saved_docs += [l['doc']['id'].split('___')[0]]
print('Examples: ', examples)
print('Covered Documents: ', len(set(saved_docs)))
# -
# # Correlations
# +
def correlation(a, b, correlation):
import pandas as pd
import json
# TODO: Add all tokens with weight of 0?
tokens = set([i for i in a.keys()] + [i for i in b.keys()])
df_corr = []
for token in tokens:
df_corr += [{'word': token, 'weight_a': a.get(token, 0), 'weight_b': b.get(token, 0)}]
ret = pd.DataFrame(df_corr).corr(correlation)
if len(ret) != 2:
raise ValueError('Could not handle \n' + json.dumps(a) + '\n' + json.dumps(b))
ret = ret.iloc[0]
if ret.name != 'weight_a':
raise ValueError('Could not handle \n' + json.dumps(a) + '\n' + json.dumps(b))
return float(ret['weight_b'])
def jaccard_similarity(list1, list2):
intersection = len(list(set(list1).intersection(list2)))
union = (len(set(list1)) + len(set(list2))) - intersection
return float(intersection) / union
# -
correlation({'project': 0.3, 'manhatten': 0.7, 'success': 1.0}, {'project': 1.0, 'manhatten': 0.5, 'success': 0.1})
correlation({'project': 0.3, 'manhatten': 0.7, 'success': 1.0}, {'project': .5, 'manhatten': 2, 'success': 0.1})
jaccard_similarity(['a', 'b', 'c'], ['a', 'b', 'c'])
jaccard_similarity(['a', 'b', 'c'], ['a', 'b'])
def calculate_correlation(method_a, method_b):
import json
from tqdm import tqdm
doc_terms_method_a = json.load(open(DIR + method_a))
doc_terms_method_b = json.load(open(DIR + method_b))
ret = []
for doc in tqdm([i for i in doc_terms_method_a.keys()]):
if doc not in doc_terms_method_b or len(doc_terms_method_a[doc].keys()) == 0 or len(doc_terms_method_b[doc].keys()) == 0:
continue
ret += [
{
'doc': doc,
'kendall': correlation(doc_terms_method_a[doc], doc_terms_method_b[doc] ,'kendall'),
'pearson': correlation(doc_terms_method_a[doc], doc_terms_method_b[doc] ,'pearson'),
'jaccard': jaccard_similarity(doc_terms_method_a[doc].keys(), doc_terms_method_b[doc].keys())
}
]
return ret
df_corr_orcas_anchor = pd.DataFrame(calculate_correlation('ms-marco-orcas-deepct-qtokens.json', 'cc-2019-47-deepct-qtokens.json'))
df_corr_orcas_anchor
df_corr_orcas_anchor.describe()
df_corr_orcas_marco_training = pd.DataFrame(calculate_correlation('ms-marco-trainig-set-deepct-qtokens.json', 'ms-marco-orcas-deepct-qtokens.json'))
df_corr_orcas_marco_training
df_corr_orcas_marco_training.describe()
df_corr_anchor_marco_training = pd.DataFrame(calculate_correlation('ms-marco-trainig-set-deepct-qtokens.json', 'cc-2019-47-deepct-qtokens.json'))
df_corr_anchor_marco_training
df_corr_anchor_marco_training.describe()
| src/deepct/train-deep-ct-ms-marco.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from gensim import corpora
import gensim
from gensim.models.doc2vec import TaggedDocument
import nltk
from nltk.corpus import stopwords
import csv
import numpy as np
import string
import re
from sklearn.decomposition import PCA
from sklearn.cluster import KMeans
from sklearn.mixture import GaussianMixture
import matplotlib.pyplot as plt
# +
def get_documents(r):
i = 0
for d in r:
dt = get_words(d)
doc = TaggedDocument([w for w in dt if w not in stopwords.words('english')], [i])
i += 1
yield doc
def get_words(r):
r = re.sub('[^a-zA-Z ]', '', r.lower())
words = nltk.word_tokenize(r)
return [w for w in words if w not in stopwords.words('english')]
with open('../resumes/processed_resumes_work.csv') as f:
r = csv.reader(f)
i = 0
jobs = []
titles = []
docs = []
for row in r:
jobs.append(row[7])
titles.append(row[3])
i += 1
if i >= 5000:
break
sents_gen = get_documents(jobs)
for s in sents_gen:
docs.append(s)
titles = np.array(titles)
# -
doc2vecModel =gensim.models.Doc2Vec(docs, size=20, iter=100)
vects = doc2vecModel.docvecs.doctag_syn0
# reduce dimention for plotting
pca = PCA(n_components = 2)
pca.fit(vects)
reduced= pca.transform(vects)
# k-means cluster
num_cluster = 50
m = KMeans(n_clusters=num_cluster, random_state=0).fit(vects)
p = m.predict(vects)
x, y = zip(*reduced)
plt.scatter(x,y,c=p, cmap='coolwarm')
plt.show()
npjobs = np.array(jobs)
for i in range(num_cluster):
print('-'*100)
wordcounts = {}
cluster_jobs = npjobs[np.where(p == i)]
print(np.random.choice(titles[np.where(p == i)], 5, replace=False))
for j in cluster_jobs:
words = get_words(j)
for w in words:
if w not in wordcounts:
wordcounts[w] = 0
wordcounts[w] += 1
wctuples = wordcounts.items()
revtuples = [(b, a) for a, b in wctuples]
print([s for c, s in sorted(revtuples, reverse=True)][:10])
# gaussian cluster
gm = GaussianMixture(n_components=num_cluster, random_state=0).fit(vects)
pm = gm.predict(vects)
plt.scatter(x,y,c=pm, cmap='coolwarm')
plt.show()
for i in range(num_cluster):
print('-'*100)
wordcounts = {}
cluster_jobs = npjobs[np.where(pm == i)]
print(np.random.choice(titles[np.where(pm == i)], 5, replace=False))
for j in cluster_jobs:
words = get_words(j)
for w in words:
if w not in wordcounts:
wordcounts[w] = 0
wordcounts[w] += 1
wctuples = wordcounts.items()
revtuples = [(b, a) for a, b in wctuples]
print([s for c, s in sorted(revtuples, reverse=True)][:10])
| parsing_scripts/cluster_resume_jobs_by_description.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp evaluation.causal
# -
# hide
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
import tensorflow as tf
physical_devices = tf.config.list_physical_devices('GPU')
tf.config.experimental.set_memory_growth(physical_devices[0], enable=True)
# # Causal Evaluation
#
# > causal
# +
#export
import numpy as np
import pandas as pd
import dowhy
from dowhy import CausalModel
import dowhy.datasets
# -
#export
from enum import Enum, auto
from abc import ABC,abstractmethod
import logging
#export
# Avoid printing dataconversion warnings from sklearn
import warnings
from sklearn.exceptions import DataConversionWarning
warnings.filterwarnings(action='ignore', category=DataConversionWarning)
# +
# Config dict to set the logging level
import logging.config
DEFAULT_LOGGING = {
'version': 1,
'disable_existing_loggers': False,
'loggers': {
'': {
'level': 'WARN',
},
}
}
logging.config.dictConfig(DEFAULT_LOGGING)
# -
# # 0. Utility Classes
# +
#This data is for testing purpuses
# +
#export
class MethodName(Enum):
propensity_score_stratification = 'backdoor.propensity_score_stratification' #<--- This is only for binary treatments
linear_regression = 'backdoor.linear_regression'
propensity_score_weighting = 'backdoor.propensity_score_weighting' #<--- This is only for binary treatments
class TargetUnit(Enum):
atc = 'atc' #Causal effect on the control group
ate = 'ate' #Average Treatment Effect
att = 'att'
class RefuteEstimate(Enum):
random_common_cause = 'random_common_cause' #Add Random Common Cause:
add_unobserved_common_cause = 'add_unobserved_common_cause' #Add Unobserved Common Causes
placebo_treatment_refuter = 'placebo_treatment_refuter' #Placebo Treatment
data_subset_refuter = 'data_subset_refuter' #Removing a random subset of the data
# -
MethodName.propensity_score_stratification.name
TargetUnit.ate.value
#export
class PropensityWeighting(Enum):
ips_weight = {'weighting_scheme':'ips_weight'} #1.Vanilla Inverse Propensity Score weighting (IPS)
ips_normalized_weight = {'weighting_scheme':'ips_normalized_weight'} #2.Self-normalized IPS weighting (also known as the Hajek estimator)
ips_stabilized_weight = {'weighting_scheme':'ips_stabilized_weight'} #3.Stabilized IPS weighting
#export
#create the abstract class for causal methods
class CausalCodeGen(ABC):
#1. Identification
@abstractmethod
def identification(self):
'identify a target estimand under the model'
pass
#2. Estimation
@abstractmethod
def estimation(self):
'Estimate causal effect based on the identified estimand'
pass
#3. Refuting
@abstractmethod
def refuting(self):
'Refute the obtained estimate'
pass
#4. Display
@abstractmethod
def display(self):
pass
# # 1. CausalCodeGen without a directed graph
# Here we specify common causes (confounders) and instruments (if any)
#export
class CausalCodeGenNoGraph(CausalCodeGen):
def __init__(self,
df_data,
treatment = ['treatment'],
outcome = 'outcome',
common_causes = ['confounders']
):
#0. Creating the causal model
self.model= CausalModel(
data = df_data,
treatment = treatment,
outcome = outcome,
common_causes = common_causes,
#effect_modifiers=data["effect_modifier_names"]
)
#1. Identification
def identification(self,proceed_when_unidentifiable=True):
self.identified_estimand = self.model.identify_effect(proceed_when_unidentifiable=proceed_when_unidentifiable)
#print(self.identified_estimand)
print('Identification Done')
pass
#2. Estimation
def estimation(self, method_name, target_units = TargetUnit.ate.value):
self.estimate = self.model.estimate_effect(
self.identified_estimand,
method_name = method_name,
target_units = target_units
)
#print(self.estimate)
#print("Causal Estimate is " + str(self.estimate.value))
print('Estimation Done')
pass
#3. Refuting
def refuting(self, method):
default = "Incorrect method"
return getattr(self, 'refuting_' + str(method), lambda: default)()
def refuting_1(self, random_seed = 1):
'''Adding a random common cause variable
Does the estimation method change its estimate after
we add an independent random variable as a common
cause to the dataset? (Hint: It should not)'''
res_random=self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name = RefuteEstimate.random_common_cause.value,
random_seed = random_seed
)
return res_random
def refuting_2(self, random_seed = 1):
'''Adding an unobserved common cause variable
How sensitive is the effect estimate when we add an
additional common cause (confounder) to the dataset that is correlated with the treatment and the outcome?
(Hint: It should not be too sensitive)
'''
res_unobserved = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name=RefuteEstimate.add_unobserved_common_cause.value,
confounders_effect_on_treatment="binary_flip",
confounders_effect_on_outcome="linear",
effect_strength_on_treatment=0.01,
effect_strength_on_outcome=0.02,
random_seed = random_seed
)
return res_unobserved
def refuting_3(self, random_seed = 1):
'''Replacing treatment with a random (placebo) variable
What happens to the estimated causal effect when we replace
the true treatment variable with an independent random variable?
(Hint: the effect should go to zero)
'''
res_placebo = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name=RefuteEstimate.placebo_treatment_refuter.value,
placebo_type="permute",
random_seed = random_seed
)
return res_placebo
def refuting_4(self, random_seed = 1):
'''Removing a random subset of the data
Does the estimated effect change significantly
when we replace the given dataset with a randomly selected subset?
(Hint: It should not)
'''
res_subset = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name = RefuteEstimate.data_subset_refuter.value,
subset_fraction = 0.9,
random_seed = random_seed
)
return res_subset
#4. Display
def display(self):
self.model.view_model()
# ### Testing CausalCodeGenNoGraph
#tst
data = dowhy.datasets.linear_dataset(
beta=10,
num_common_causes=5,
num_instruments = 0,
num_effect_modifiers=0,
num_samples=50000,
treatment_is_binary=True,
num_discrete_common_causes=1
)
df = data["df"]
print(df.head())
#tst
obj_causalCodeGenNoGraph = CausalCodeGenNoGraph(
df_data = df,
treatment = data["treatment_name"],
outcome = data["outcome_name"],
common_causes = data["common_causes_names"],
)
#tst display
obj_causalCodeGenNoGraph.display()
#tst identification
obj_causalCodeGenNoGraph.identification()
#tst estimation binary
obj_causalCodeGenNoGraph.estimation(
method_name=MethodName.propensity_score_stratification.value #<------- HyperParameter
)
# tst estimation non-binary
obj_causalCodeGenNoGraph.estimation(
#method_name=MethodName.linear_regression.value #<------- HyperParameter
method_name=MethodName.propensity_score_weighting.value #<------- HyperParameter
)
#tst refuting
method = [1,2,3,4] #<----- Hyperparameter
res_method = obj_causalCodeGenNoGraph.refuting(1)
print(res_method)
res_method = [obj_causalCodeGenNoGraph.refuting(i) for i in method]
print(res_method[0])
print(res_method[1])
print(res_method[2])
print(res_method[3])
# # 1.1 Causal CodeGen for Instrumental Variables
#export
class CausalCodeGenIV(CausalCodeGen):
def __init__(self,
df_data,
treatment = ['treatment'],
outcome = 'outcome',
common_causes = ['confounders'],
instruments = ['instruments']
):
#0. Creating the causal model
self.model= CausalModel(
data = df_data,
treatment = treatment,
outcome = outcome,
common_causes = common_causes,
instruments = instruments
#effect_modifiers=data["effect_modifier_names"]
)
#1. Identification
def identification(self,proceed_when_unidentifiable=True):
self.identified_estimand = self.model.identify_effect(proceed_when_unidentifiable=proceed_when_unidentifiable)
#print(self.identified_estimand)
print('Identification Done')
pass
#2. Estimation
def estimation(self, method_name='iv.instrumental_variable', target_units = TargetUnit.ate.value):
self.estimate = self.model.estimate_effect(
self.identified_estimand,
method_name = method_name,
target_units = target_units
)
#print(self.estimate)
#print("Causal Estimate is " + str(self.estimate.value))
print('Estimation Done')
pass
#3. Refuting
def refuting(self, method):
default = "Incorrect method"
return getattr(self, 'refuting_' + str(method), lambda: default)()
def refuting_1(self, random_seed = 1):
'''Adding a random common cause variable
Does the estimation method change its estimate after
we add an independent random variable as a common
cause to the dataset? (Hint: It should not)'''
res_random=self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name = RefuteEstimate.random_common_cause.value,
random_seed = random_seed
)
return res_random
def refuting_2(self, random_seed = 1):
'''Adding an unobserved common cause variable
How sensitive is the effect estimate when we add an
additional common cause (confounder) to the dataset that is correlated with the treatment and the outcome?
(Hint: It should not be too sensitive)
'''
res_unobserved = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name=RefuteEstimate.add_unobserved_common_cause.value,
confounders_effect_on_treatment="binary_flip",
confounders_effect_on_outcome="linear",
effect_strength_on_treatment=0.01,
effect_strength_on_outcome=0.02,
random_seed = random_seed
)
return res_unobserved
def refuting_3(self, random_seed = 1):
'''Replacing treatment with a random (placebo) variable
What happens to the estimated causal effect when we replace
the true treatment variable with an independent random variable?
(Hint: the effect should go to zero)
'''
res_placebo = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name=RefuteEstimate.placebo_treatment_refuter.value,
placebo_type="permute",
random_seed = random_seed
)
return res_placebo
def refuting_4(self, random_seed = 1):
'''Removing a random subset of the data
Does the estimated effect change significantly
when we replace the given dataset with a randomly selected subset?
(Hint: It should not)
'''
res_subset = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name = RefuteEstimate.data_subset_refuter.value,
subset_fraction = 0.9,
random_seed = random_seed
)
return res_subset
#4. Display
def display(self):
self.model.view_model()
# #### Testing IV
#tst
data = dowhy.datasets.linear_dataset(
beta=1,
num_common_causes=10,
num_instruments = 1,
num_effect_modifiers=0,
num_samples=50000,
treatment_is_binary=False,
num_discrete_common_causes=1
)
data.keys()
df = data["df"]
print(df.head())
#tst
obj_causalCodeGenIV = CausalCodeGenIV(
df_data = df,
treatment = data["treatment_name"],
outcome = data["outcome_name"],
common_causes = data["common_causes_names"],
instruments = data['instrument_names']
)
#tst display
obj_causalCodeGenIV.display()
#tst identification
obj_causalCodeGenIV.identification()
#tst estimation IV
obj_causalCodeGenIV.estimation(
#method_name=MethodName.propensity_score_stratification.value #<------- HyperParameter
)
# estimate
obj_causalCodeGenIV.estimate.value
[i for i in range(1,5)]
#Refuting
res_method = [obj_causalCodeGenIV.refuting(i) for i in range(1,5)]
print(res_method[0])
print(res_method[1])
print(res_method[2])
print(res_method[3])
# # 2. CausalCodeGen Estimating effect of multiple treatments
#export
class CausalCodeGenMultiple(CausalCodeGen):
def __init__(self,
df_data,
treatment = ['treatment'],
outcome = 'outcome',
common_causes = ['confounders']
):
#0. Creating the causal model
self.model= CausalModel(
data = df_data,
treatment = treatment,
outcome = outcome,
common_causes = common_causes,
#effect_modifiers=data["effect_modifier_names"]
)
#1. Identification
def identification(self,proceed_when_unidentifiable=True):
self.identified_estimand = self.model.identify_effect(proceed_when_unidentifiable=proceed_when_unidentifiable)
print(self.identified_estimand)
print('Identification Done')
pass
#2. Estimation
def estimation(self, target_units = TargetUnit.ate.value):
self.estimate = self.model.estimate_effect(
self.identified_estimand,
method_name = MethodName.linear_regression.value,
target_units = target_units,
method_params={'need_conditional_estimates': False} #No Effect Modifiers Proposed
)
print(self.estimate)
print("Causal Estimate is " + str(self.estimate.value))
print('Estimation Done')
pass
#3. Refuting
def refuting(self, method):
default = "Incorrect method"
return getattr(self, 'refuting_' + str(method), lambda: default)()
def refuting_1(self, random_seed = 1):
'''Adding a random common cause variable
Does the estimation method change its estimate after
we add an independent random variable as a common
cause to the dataset? (Hint: It should not)'''
res_random=self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name = RefuteEstimate.random_common_cause.value,
random_seed = random_seed
)
return res_random
def refuting_2(self, random_seed = 1):
'''Adding an unobserved common cause variable
How sensitive is the effect estimate when we add an
additional common cause (confounder) to the dataset that is correlated with the treatment and the outcome?
(Hint: It should not be too sensitive)
'''
res_unobserved = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name=RefuteEstimate.add_unobserved_common_cause.value,
confounders_effect_on_treatment="binary_flip",
confounders_effect_on_outcome="linear",
effect_strength_on_treatment=0.01,
effect_strength_on_outcome=0.02,
random_seed = random_seed
)
return res_unobserved
def refuting_3(self, random_seed = 1):
'''Replacing treatment with a random (placebo) variable
What happens to the estimated causal effect when we replace
the true treatment variable with an independent random variable?
(Hint: the effect should go to zero)
'''
res_placebo = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name=RefuteEstimate.placebo_treatment_refuter.value,
#placebo_type="permute",
random_seed = random_seed
)
return res_placebo
def refuting_4(self, random_seed = 1):
'''Removing a random subset of the data
Does the estimated effect change significantly
when we replace the given dataset with a randomly selected subset?
(Hint: It should not)
'''
res_subset = self.model.refute_estimate(
self.identified_estimand,
self.estimate,
method_name = RefuteEstimate.data_subset_refuter.value,
subset_fraction = 0.9,
random_seed = random_seed
)
return res_subset
#4. Display
def display(self):
self.model.view_model()
# ### Testing CausalCodeGenMultiple
#tst
data = dowhy.datasets.linear_dataset(
beta=1,
num_common_causes=5,
num_instruments = 0,
num_effect_modifiers=0,
num_treatments=2, #<---- Multiple Treatments
num_samples=50000,
treatment_is_binary=False,
num_discrete_common_causes=1,
one_hot_encode=False
)
df = data["df"]
print(df.head())
#tst
obj_causalCodeGenMultiple = CausalCodeGenMultiple(
df_data = df,
treatment = data["treatment_name"],
outcome = data["outcome_name"],
common_causes = data["common_causes_names"],
)
#tst display
obj_causalCodeGenMultiple.display()
#tst identification
obj_causalCodeGenMultiple.identification()
#tst estimation binary
obj_causalCodeGenMultiple.estimation()
res_method = obj_causalCodeGenMultiple.refuting(1)
print(res_method)
res_method = obj_causalCodeGenMultiple.refuting(2)
print(res_method)
res_method = obj_causalCodeGenMultiple.refuting(3)
print(res_method)
res_method = obj_causalCodeGenMultiple.refuting(4)
print(res_method)
| nbs/04_evaluation.causal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/RajeevAtla/HTCS-ML-Team-B/blob/master/sentiment_analysis.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="oEph_rmi9lCm" colab_type="text"
# # Building a Sentiment Classifier using Scikit-Learn
#
# Acknowledgement: This is derived from https://towardsdatascience.com/building-a-sentiment-classifier-using-scikit-learn-54c8e7c5d2f0.
# + [markdown] id="blTQmk0V9lCn" colab_type="text"
# <center><img src="https://raw.githubusercontent.com/lazuxd/simple-imdb-sentiment-analysis/master/smiley.jpg"/></center>
# <center><i>Image by AbsolutVision @ <a href="https://pixabay.com/ro/photos/smiley-emoticon-furie-sup%C4%83rat-2979107/">pixabay.com</a></i></center>
#
# > **Sentiment analysis**, an important area in Natural Language Processing, is the process of automatically detecting affective states of text. Sentiment analysis is widely applied to voice-of-customer materials such as product reviews in online shopping websites like Amazon, movie reviews or social media. It can be just a basic task of classifying the polarity of a text as being positive/negative or it can go beyond polarity, looking at emotional states such as "happy", "angry", etc.
#
# Here we will build a classifier that is able to distinguish movie reviews as being either positive or negative. For that, we will use [Large Movie Review Dataset v1.0](http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz)<sup>(2)</sup> of IMDB movie reviews.
# This dataset contains 50,000 movie reviews divided evenly into 25k train and 25k test. The labels are balanced between the two classes (positive and negative). Reviews with a score <= 4 out of 10 are labeled negative and those with score >= 7 out of 10 are labeled positive. Neutral reviews are not included in the labeled data. This dataset also contains unlabeled reviews for unsupervised learning; we will not use them here. There are no more than 30 reviews for a particular movie because the ratings of the same movie tend to be correlated. All reviews for a given movie are either in train or test set but not in both, in order to avoid test accuracy gain by memorizing movie-specific terms.
#
#
# + [markdown] id="7CL4d-HH9lCn" colab_type="text"
# ## Data preprocessing
# + [markdown] id="evm9tjsW9lCo" colab_type="text"
# After the dataset has been downloaded and extracted from archive we have to transform it into a more suitable form for feeding it into a machine learning model for training. We will start by combining all review data into 2 pandas Data Frames representing the train and test datasets, and then saving them as csv files: *imdb_train.csv* and *imdb_test.csv*.
#
# The Data Frames will have the following form:
#
# |text |label |
# |:---------:|:---------:|
# |review1 |0 |
# |review2 |1 |
# |review3 |1 |
# |....... |... |
# |reviewN |0 |
#
# where:
# - review1, review2, ... = the actual text of movie review
# - 0 = negative review
# - 1 = positive review
# + [markdown] id="EckG-g0E9lCo" colab_type="text"
# But machine learnng algorithms work only with numerical values. We can't just input the text itself into a machine learning model and have it learn from that. We have to, somehow, represent the text by numbers or vectors of numbers. One way of doing this is by using the **Bag-of-words** model<sup>(3)</sup>, in which a piece of text(often called a **document**) is represented by a vector of the counts of words from a vocabulary in that document. This model doesn't take into account grammar rules or word ordering; all it considers is the frequency of words. If we use the counts of each word independently we name this representation a **unigram**. In general, in a **n-gram** we take into account the counts of each combination of n words from the vocabulary that appears in a given document.
#
# For example, consider these two documents:
# <br>
# <div style="font-family: monospace;"><center><b>d1: "I am learning" </b></center></div>
# <div style="font-family: monospace;"><center><b>d2: "Machine learning is cool"</b></center></div>
# <br>
# The vocabulary of all words encountered in these two sentences is:
#
# <br/>
# <div style="font-family: monospace;"><center><b>v: [ I, am, learning, machine, is, cool ]</b></center></div>
# <br>
# The unigram representations of d1 and d2:
# <br>
#
# |unigram(d1)|I |am |learning|machine |is |cool |
# |:---------:|:------:|:------:|:------:|:------:|:------:|:------:|
# | |1 |1 |1 |0 |0 |0 |
#
# |unigram(d2)|I |am |learning|machine |is |cool |
# |:---------:|:------:|:------:|:------:|:------:|:------:|:------:|
# | |0 |0 |1 |1 |1 |1 |
#
# And, the bigrams of d1 and d2 are:
#
# |bigram(d1) |I I |I am |I learning|...|machine am|machine learning|...|cool is|cool cool|
# |:---------:|:------:|:------:|:--------:|:-:|:--------:|:--------------:|:-:|:-----:|:-------:|
# | |0 |1 |0 |...|0 |0 |...|0 |0 |
#
# |bigram(d2) |I I |I am |I learning|...|machine am|machine learning|...|cool is|cool cool|
# |:---------:|:------:|:------:|:--------:|:-:|:--------:|:--------------:|:-:|:-----:|:-------:|
# | |0 |0 |0 |...|0 |1 |...|0 |0 |
# + [markdown] id="rcg3cjFI9lCo" colab_type="text"
# Often, we can achieve slightly better results if instead of counts of words we use something called **term frequency times inverse document frequency** (or **tf-idf**). Maybe it sounds complicated, but it is not. Bear with me, I will explain this. The intuition behind this is the following. So, what's the problem of using just the frequency of terms inside a document? Although some terms may have a high frequency inside documents they may not be so relevant for describing a given document in which they appear. That's because those terms may also have a high frequency across the collection of all documents. For example, a collection of movie reviews may have terms specific to movies/cinematography that are present in almost all documents(they have a high **document frequency**). So, when we encounter those terms in a document this doesn't tell much about whether it is a positive or negative review. We need a way of relating **term frequency** (how frequent a term is inside a document) to **document frequency** (how frequent a term is across the whole collection of documents). That is:
#
# $$\begin{align}\frac{\text{term frequency}}{\text{document frequency}} &= \text{term frequency} \cdot \frac{1}{\text{document frequency}} \\ &= \text{term frequency} \cdot \text{inverse document frequency} \\ &= \text{tf} \cdot \text{idf}\end{align}$$
#
# Now, there are more ways used to describe both term frequency and inverse document frequency. But the most common way is by putting them on a logarithmic scale:
#
# $$tf(t, d) = log(1+f_{t,d})$$
# $$idf(t) = log(\frac{1+N}{1+n_t})$$
#
# where:
# $$\begin{align}f_{t,d} &= \text{count of term } \textbf{t} \text{ in document } \textbf{d} \\
# N &= \text{total number of documents} \\
# n_t &= \text{number of documents that contain term } \textbf{t}\end{align}$$
#
# We added 1 in the first logarithm to avoid getting $-\infty$ when $f_{t,d}$ is 0. In the second logarithm we added one fake document to avoid division by zero.
# + [markdown] id="Kt7bdF819lCp" colab_type="text"
# Before we transform our data into vectors of counts or tf-idf values we should remove English **stopwords**<sup>(6)(7)</sup>. Stopwords are words that are very common in a language and are usually removed in the preprocessing stage of natural text-related tasks like sentiment analysis or search.
# + [markdown] id="p5_MOC6Z9lCp" colab_type="text"
# Note that we should construct our vocabulary only based on the training set. When we will process the test data in order to make predictions we should use only the vocabulary constructed in the training phase, the rest of the words will be ignored.
# + [markdown] id="DicmPJO09lCq" colab_type="text"
# Now, let's create the data frames from the supplied csv files:
# + id="Hw_fmTEX9lCq" colab_type="code" colab={}
import pandas as pd
# + id="GE1zgTOi9lCs" colab_type="code" colab={}
# Read in the training and test datasets from previously created csv files
imdb_train = pd.read_csv('csv/imdb_train.csv')
imdb_test = pd.read_csv('csv/imdb_test.csv')
# + id="yPYggSnd9lCu" colab_type="code" colab={} outputId="580e2d8e-c864-413b-a2ce-6b5737254b6a"
# Display information and first few entries from the training and test datasets
pd.set_option('display.max_colwidth', None)
print ("----- Training dataset Info:")
imdb_train.info(verbose=None, buf=None, max_cols=None, memory_usage=None, null_counts=None)
print ("Training dataset Content:")
print(imdb_train.iloc[:5])
print ("\n----- Test dataset Info:")
imdb_test.info(verbose=None, buf=None, max_cols=None, memory_usage=None, null_counts=None)
print ("Test dataset Content:")
print(imdb_test.iloc[:5])
# + [markdown] id="-COMQ8dY9lCw" colab_type="text"
# ### Text vectorization
# + [markdown] id="AtrcxAUp9lCx" colab_type="text"
# Fortunately, for the text vectorization part all the hard work is already done in the Scikit-Learn classes `CountVectorizer`<sup>(8)</sup> and `TfidfTransformer`<sup>(5)</sup>. We will use these classes to transform our csv files into unigram and bigram matrices(using both counts and tf-idf values). (It turns out that if we only use a n-gram for a large n we don't get a good accuracy, we usually use all n-grams up to some n. So, when we say here bigrams we actually refer to uni+bigrams and when we say unigrams it's just unigrams.) Each row in those matrices will represent a document (review) in our dataset, and each column will represent values associated with each word in the vocabulary (in the case of unigrams) or values associated with each combination of maximum 2 words in the vocabulary (bigrams).
#
# `CountVectorizer` has a parameter `ngram_range` which expects a tuple of size 2 that controls what n-grams to include. After we constructed a `CountVectorizer` object we should call `.fit()` method with the actual text as a parameter, in order for it to learn the required statistics of our collection of documents. Then, by calling `.transform()` method with our collection of documents it returns the matrix for the n-gram range specified. As the class name suggests, this matrix will contain just the counts. To obtain the tf-idf values, the class `TfidfTransformer` should be used. It has the `.fit()` and `.transform()` methods that are used in a similar way with those of `CountVectorizer`, but they take as input the counts matrix obtained in the previous step and `.transform()` will return a matrix with tf-idf values. We should use `.fit()` only on training data and then store these objects. When we want to evaluate the test score or whenever we want to make a prediction we should use these objects to transform the data before feeding it into our classifier.
#
# Note that the matrices generated for our train or test data will be huge, and if we store them as normal numpy arrays they will not even fit into RAM. But most of the entries in these matrices will be zero. So, these Scikit-Learn classes are using Scipy sparse matrices<sup>(9)</sup> (`csr_matrix`<sup>(10)</sup> to be more exactly), which store just the non-zero entries and save a LOT of space.
#
# We will use a linear classifier with stochastic gradient descent, `sklearn.linear_model.SGDClassifier`<sup>(11)</sup>, as our model. First we will generate and save our data in 4 forms: unigram and bigram matrix (with both counts and tf-idf values for each). Then we will train and evaluate our model for each these 4 data representations using `SGDClassifier` with the default parameters. After that, we choose the data representation which led to the best score and we will tune the hyper-parameters of our model with this data form using cross-validation in order to obtain the best results.
# + id="xmfPejcN9lCx" colab_type="code" colab={}
from sklearn.feature_extraction.text import CountVectorizer, TfidfTransformer
# + [markdown] id="s6b7Oqp-9lCz" colab_type="text"
# #### Unigram Counts
# + id="1dArWvRR9lCz" colab_type="code" colab={} outputId="304f63a1-3c5a-48ec-f708-8236e0003862"
# Create a unigram vectorizer and process the training set to generate a list of words.
# Note that unigram processing is set via the ngram_range parameter
unigram_vectorizer = CountVectorizer(ngram_range=(1, 1))
unigram_vectorizer.fit(imdb_train['text'].values)
# + id="nHGOebuq9lC2" colab_type="code" colab={} outputId="86a3ee95-00e0-4b5d-d0c3-5bfc765e9725"
# Display the length and a few samples of the unigram vectorizer to show the words that have been extracted
print("Number of words found:", len(unigram_vectorizer.get_feature_names()))
print(unigram_vectorizer.get_feature_names()[10000:10100])
print(unigram_vectorizer.get_feature_names()[50000:50100])
# + id="Xgehvpkr9lC3" colab_type="code" colab={}
# Now process the training dataset to get a count of the words extracted earlier
X_train_unigram = unigram_vectorizer.transform(imdb_train['text'].values)
# + id="eqEqnSZZ9lC5" colab_type="code" colab={} outputId="b3d9065b-f02b-4ed4-e20f-f465e11ab9db"
# Display the attributes the word count matrix; notice it is huge with 25000 rows since we have 25000 entries
# in the training dataset and 74849 columns since we saw above that we have a vocabulary of 74849 words
print(repr(X_train_unigram))
# + [markdown] id="uwUZxydJ9lC7" colab_type="text"
# #### Unigram Tf-Idf
# + id="xx0FsrgL9lC7" colab_type="code" colab={} outputId="d13a68ed-8292-4a9b-faa9-8265b85a7b0e"
# Create a unigram tf-idf vectorizer and load the training set using the word count matrix from earlier
unigram_tf_idf_transformer = TfidfTransformer()
unigram_tf_idf_transformer.fit(X_train_unigram)
# + id="pREGYWEg9lC9" colab_type="code" colab={}
# Now calculate the unigram tf-idf statistics
X_train_unigram_tf_idf = unigram_tf_idf_transformer.transform(X_train_unigram)
# + id="kfUMU4DW9lC_" colab_type="code" colab={} outputId="21653fb8-4354-4322-e4c8-adc9ed39e636"
# Display the attributes the unigram tf-idf matrix; it should be the same size as the unigram matrix above
print(repr(X_train_unigram_tf_idf))
# + [markdown] id="9GIwBJKM9lDB" colab_type="text"
# #### Bigram Counts
# + id="l00sYvzX9lDB" colab_type="code" colab={} outputId="52748c55-184b-409b-c8ec-4c2728e93b79"
# Create a bigram vectorizer and process the training set to generate a list of bigrams.
# Note that bigram processing is set via the ngram_range parameter and so includes unigrams and bigrams
bigram_vectorizer = CountVectorizer(ngram_range=(1, 2))
bigram_vectorizer.fit(imdb_train['text'].values)
# + id="zlRED15i9lDD" colab_type="code" colab={} outputId="c0fba0ba-d30c-4308-a8c2-dfa0c3527d53"
# Display the length and a few samples of the bigram vectorizer to show the bigrams that have been extracted
print("Number of bigrams found:", len(bigram_vectorizer.get_feature_names()))
print(bigram_vectorizer.get_feature_names()[10000:10100])
print(bigram_vectorizer.get_feature_names()[50000:50100])
# + id="W2iPN3dv9lDF" colab_type="code" colab={}
# Now generate bigram statistics on the training set
X_train_bigram = bigram_vectorizer.transform(imdb_train['text'].values)
# + id="dxQaYkVg9lDG" colab_type="code" colab={} outputId="03cd3b38-0b4d-4e17-fcd2-cff8b2b3db89"
# Display the attributes the bigram count matrix; notice it is really huge with 25000 rows since we have 25000 entries
# in the training dataset and 1520266 columns since we saw above that we have 1520266 bigrams
print(repr(X_train_bigram))
# + [markdown] id="Qb745OZ79lDJ" colab_type="text"
# #### Bigram Tf-Idf
# + id="y_o-dnsd9lDJ" colab_type="code" colab={} outputId="5e7a69bc-6cdf-479b-abfe-2a669d57be10"
# Create a bigram tf-idf vectorizer and load the training set using the bigram count matrix from earlier
bigram_tf_idf_transformer = TfidfTransformer()
bigram_tf_idf_transformer.fit(X_train_bigram)
# + id="knxsmxUu9lDL" colab_type="code" colab={}
# Now calculate the bigram tf-idf statistics
X_train_bigram_tf_idf = bigram_tf_idf_transformer.transform(X_train_bigram)
# + id="HOVm5loL9lDN" colab_type="code" colab={} outputId="2068dd54-f136-450a-8820-6da40a83142d"
# Display the attributes the bigram tf-idf matrix; it should be the same size as the bigram matrix above
print(repr(X_train_bigram_tf_idf))
# + [markdown] id="YjConCNN9lDO" colab_type="text"
# ### Try the four different data formats (unigram, bigram with and without tf_idf) on the training set and pick the best
# + [markdown] id="a2Eno3PI9lDO" colab_type="text"
# Now, for each data form we split it into train & validation sets, train a `SGDClassifier` and output the score.
# + id="OL738RU39lDP" colab_type="code" colab={}
from sklearn.linear_model import SGDClassifier
from sklearn.model_selection import train_test_split
from scipy.sparse import csr_matrix
from sklearn.metrics import confusion_matrix, precision_score, recall_score, f1_score
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from typing import Tuple
# + id="9OOLANzU9lDQ" colab_type="code" colab={}
# Helper function to display confusion matrix
def display_confusion_matrix(y_true, y_pred) -> None:
cf_matrix = confusion_matrix(y_true, y_pred)
group_names = ['True Neg','False Pos','False Neg','True Pos']
group_counts = ["{0:0.0f}".format(value) for value in cf_matrix.flatten()]
group_percentages = ["{0:.2%}".format(value) for value in cf_matrix.flatten()/np.sum(cf_matrix)]
labels = [f"{v1}\n{v2}\n{v3}" for v1, v2, v3 in zip(group_names,group_counts,group_percentages)]
labels = np.asarray(labels).reshape(2,2)
sns.heatmap(cf_matrix, annot=labels, fmt='', cmap='coolwarm')
plt.show()
# + id="BCMyDFTa9lDS" colab_type="code" colab={}
def train_and_show_scores(X: csr_matrix, y: np.array, title: str) -> Tuple[float, float]:
X_train, X_valid, y_train, y_valid = train_test_split(
X, y, train_size=0.75, stratify=y
)
clf = SGDClassifier()
clf.fit(X_train, y_train)
train_score = clf.score(X_train, y_train)
valid_score = clf.score(X_valid, y_valid)
print(f'{title}\nTrain score: {round(train_score, 2)} ; Validation score: {round(valid_score, 2)}')
train_pred = clf.predict(X_train)
valid_pred = clf.predict(X_valid)
print(f'Train precision: {round(precision_score(y_train, train_pred), 2)} ; Validation precision: {round(precision_score(y_valid, valid_pred), 2)}')
print(f'Train recall: {round(recall_score(y_train, train_pred), 2)} ; Validation recall: {round(recall_score(y_valid, valid_pred), 2)}')
print(f'Train F1: {round(f1_score(y_train, train_pred), 2)} ; Validation F1: {round(f1_score(y_valid, valid_pred), 2)}')
print("Train Confusion Matrix: ")
print(confusion_matrix(y_train, train_pred))
display_confusion_matrix(y_train, train_pred)
print("Validation Confusion Matrix: ")
print(confusion_matrix(y_valid, valid_pred))
display_confusion_matrix(y_valid, valid_pred)
print("\n")
return train_score, valid_score, clf
# + id="oz2vhlp49lDU" colab_type="code" colab={}
y_train = imdb_train['label'].values
# + id="jAfpF9iX9lDW" colab_type="code" colab={} outputId="f51dcfa9-6d2f-46d1-df5e-a92689d240fc"
uc_train_score, uc_valid_score, uc_clf = train_and_show_scores(X_train_unigram, y_train, '----- Unigram Counts -----')
utfidf_train_score, utfidf_valid_score, utfidf_clf = train_and_show_scores(X_train_unigram_tf_idf, y_train, '----- Unigram Tf-Idf -----')
bc_train_score, bc_valid_score, bc_clf = train_and_show_scores(X_train_bigram, y_train, '----- Bigram Counts -----')
btfidf_train_score, btfidf_valid_score, btfidf_clf = train_and_show_scores(X_train_bigram_tf_idf, y_train, '----- Bigram Tf-Idf -----')
# + id="McGxPaXb9lDX" colab_type="code" colab={} outputId="8ac7fdcf-dcdc-467a-dcdb-fbf3ba1d3da2"
# Display the previously derived scores for the four scenarios
sns.set_style("whitegrid", {'grid.linestyle': '--'})
print ("Training score for the four approaches:")
ax1 = sns.barplot(
x= ['Unigram Count', 'Unigram tf-idf', 'Bigram Count', 'Bigram tf-idf'],
y= [uc_train_score, utfidf_train_score, bc_train_score, btfidf_train_score])
ax1.set(ylim=(0.8, 1.0))
plt.show()
print ("Validation score for the four approaches:")
ax2 = sns.barplot(
x= ['Unigram Count', 'Unigram tf-idf', 'Bigram Count', 'Bigram tf-idf'],
y= [uc_valid_score, utfidf_valid_score, bc_valid_score, btfidf_valid_score])
ax2.set(ylim=(0.8, 1.0))
plt.show()
# + [markdown] id="zE0LbIvB9lDa" colab_type="text"
# The best data form seems to be **bigram with tf-idf** as it gets the highest validation accuracy: **0.9**; so we will choose it as our preferred approach.
# + [markdown] id="gD4vT4Uc9lDb" colab_type="text"
# ### Testing model
# + id="4DFFtigA9lDb" colab_type="code" colab={}
# Transform the test data set into the bigram tf-idf format
X_test = bigram_vectorizer.transform(imdb_test['text'].values)
X_test = bigram_tf_idf_transformer.transform(X_test)
y_test = imdb_test['label'].values
# + id="xJYdvnUH9lDc" colab_type="code" colab={} outputId="1f2143cf-4c04-483a-c115-8d1ef4f17b18"
# Now evaluate the test data using the previously trained bigram tf-idf classifier
clf = btfidf_clf
score = clf.score(X_test, y_test)
print(f'Score: {round(score, 4)}')
test_pred = clf.predict(X_test)
print(f'Test precision: {round(precision_score(y_test, test_pred), 4)}')
print(f'Test recall: {round(recall_score(y_test, test_pred), 4)}')
print(f'Test F1: {round(f1_score(y_test, test_pred), 4)}')
print("Test Confusion Matrix: ")
print(confusion_matrix(y_test, test_pred))
display_confusion_matrix(y_test, test_pred)
print("\n")
# + [markdown] id="AeOTS2Qo9lDe" colab_type="text"
# And we got almost 90% test accuracy. That's not bad for our simple linear model. There are more advanced methods that give better results. The current state-of-the-art on this dataset is **97.42%** <sup>(13)</sup>
# + [markdown] id="5_rNamQ09lDe" colab_type="text"
# ## References
#
# <sup>(1)</sup> [Sentiment Analysis - Wikipedia](https://en.wikipedia.org/wiki/Sentiment_analysis)
# <sup>(2)</sup> [Learning Word Vectors for Sentiment Analysis](http://ai.stanford.edu/~amaas/papers/wvSent_acl2011.pdf)
# <sup>(3)</sup> [Bag-of-words model - Wikipedia](https://en.wikipedia.org/wiki/Bag-of-words_model)
# <sup>(4)</sup> [Tf-idf - Wikipedia](https://en.wikipedia.org/wiki/Tf%E2%80%93idf)
# <sup>(5)</sup> [TfidfTransformer - Scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.TfidfTransformer.html)
# <sup>(6)</sup> [Stop words - Wikipedia](https://en.wikipedia.org/wiki/Stop_words)
# <sup>(7)</sup> [A list of English stopwords](https://gist.github.com/sebleier/554280)
# <sup>(8)</sup> [CountVectorizer - Scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.feature_extraction.text.CountVectorizer.html)
# <sup>(9)</sup> [Scipy sparse matrices](https://docs.scipy.org/doc/scipy/reference/sparse.html)
# <sup>(10)</sup> [Compressed Sparse Row matrix](https://docs.scipy.org/doc/scipy/reference/generated/scipy.sparse.csr_matrix.html#scipy.sparse.csr_matrix)
# <sup>(11)</sup> [SGDClassifier - Scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html)
# <sup>(12)</sup> [RandomizedSearchCV - Scikit-learn documentation](https://scikit-learn.org/stable/modules/generated/sklearn.model_selection.RandomizedSearchCV.html)
# <sup>(13)</sup> [Sentiment Classification using Document Embeddings trained with
# Cosine Similarity](https://www.aclweb.org/anthology/P19-2057.pdf)
| sentiment_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rick1270/DS-Sprint-01-Dealing-With-Data/blob/master/DS_Unit_1_Sprint_Challenge_1_Clayton.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="NooAiTdnafkz" colab_type="text"
# # Data Science Unit 1 Sprint Challenge 1
#
# ## Loading, cleaning, visualizing, and analyzing data
#
# In this sprint challenge you will look at a dataset of the survival of patients who underwent surgery for breast cancer.
#
# http://archive.ics.uci.edu/ml/datasets/Haberman%27s+Survival
#
# Data Set Information:
# The dataset contains cases from a study that was conducted between 1958 and 1970 at the University of Chicago's Billings Hospital on the survival of patients who had undergone surgery for breast cancer.
#
# Attribute Information:
# 1. Age of patient at time of operation (numerical)
# 2. Patient's year of operation (year - 1900, numerical)
# 3. Number of positive axillary nodes detected (numerical)
# 4. Survival status (class attribute)
# -- 1 = the patient survived 5 years or longer
# -- 2 = the patient died within 5 year
#
# Sprint challenges are evaluated based on satisfactory completion of each part. It is suggested you work through it in order, getting each aspect reasonably working, before trying to deeply explore, iterate, or refine any given step. Once you get to the end, if you want to go back and improve things, go for it!
# + [markdown] id="5wch6ksCbJtZ" colab_type="text"
# ## Part 1 - Load and validate the data
#
# - Load the data as a `pandas` data frame.
# - Validate that it has the appropriate number of observations (you can check the raw file, and also read the dataset description from UCI).
# - Validate that you have no missing values.
# - Add informative names to the features.
# - The survival variable is encoded as 1 for surviving >5 years and 2 for not - change this to be 0 for not surviving and 1 for surviving >5 years (0/1 is a more traditional encoding of binary variables)
#
# At the end, print the first five rows of the dataset to demonstrate the above.
# + [markdown] id="cawmnkdQQno-" colab_type="text"
# Attribute Information:
# 1. Age of patient at time of operation (numerical)
# 2. Patient's year of operation (year - 1900, numerical)
# 3. Number of positive axillary nodes detected (numerical)
# 4. Survival status (class attribute)
# 1 = the patient survived 5 years or longer
# 2 = the patient died within 5 year
# + id="287TpoGKFRVK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 297} outputId="b1377779-0b3e-4598-af3b-7eaff62abe95"
# TODO
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
survival_URL = 'http://archive.ics.uci.edu/ml/machine-learning-databases/haberman/haberman.data'
survival_data = pd.read_csv(survival_URL, header = None)
survival_data.describe()
# + id="LMKfaodARrok" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 204} outputId="bfd9bdea-370f-4817-8c68-e048b28a06e1"
survival_data.head()
# + [markdown] id="G7rLytbrO38L" colab_type="text"
# ## Part 2 - Examine the distribution and relationships of the features
#
# Explore the data - create at least *2* tables (can be summary statistics or crosstabulations) and *2* plots illustrating the nature of the data.
#
# This is open-ended, so to remind - first *complete* this task as a baseline, then go on to the remaining sections, and *then* as time allows revisit and explore further.
#
# Hint - you may need to bin some variables depending on your chosen tables/plots.
# + id="n1z1bw07QCo9" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="87ddeac6-b910-42bb-e833-58b0b91285d1"
age_bins = pd.cut(survival_data[0], 10)
age = pd.crosstab(age_bins, survival_data[3], normalize = 'index')
age
# + id="_rWqB-VUZdvN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 413} outputId="f126aca4-497c-43a5-a973-ec02eb824b78"
age.plot.bar(stacked = True)
plt.legend(title='Survival')
plt.show()
# + id="IAkllgCIFVj0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="9799eec1-8ccc-4b2c-bb94-fdbb0e67d3b7"
# TODO
year_bins = pd.cut(survival_data[1], 10)
year = pd.crosstab(year_bins, survival_data[3], normalize='index')
year
# + id="cZ4I8iLYj4go" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 413} outputId="da573ba3-f059-49e9-a3be-0d4cbb5e014f"
year.plot.bar(stacked=True)
plt.legend(title='Survival')
plt.show()
# + id="cPNef3IwoA_4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 390} outputId="58a78074-bde8-4a34-e48b-273f634229f5"
node_bins = pd.cut(survival_data[2], 10)
node = pd.crosstab(node_bins, survival_data[3], normalize='index')
node
# + id="PBHcf2fToBEO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 405} outputId="3274a97a-987b-4bca-e8c7-f6e77db37b7a"
node.plot.bar(stacked=True)
plt.legend(title='Survival')
plt.show()
# + [markdown] id="ZM8JckA2bgnp" colab_type="text"
# ## Part 3 - Analysis and Interpretation
#
# Now that you've looked at the data, answer the following questions:
#
# - What is at least one feature that looks to have a positive correlation with survival?
# Age - Being 30 - 45.5 yo have the highest survival rate
# - What is at least one feature that looks to have a negative correlation with survival?
# Age - Patients over 77 have the lowest rate of survival
# - How are those two features related with each other, and what might that mean?
# While not linear, the younger end of the sample has a much higher rate of survival than the older end
#
# Answer with text, but feel free to intersperse example code/results or refer to it from earlier.
| DS_Unit_1_Sprint_Challenge_1_Clayton.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Introduction
#
# Phase diagram and entry inquiry.
#
# Written using:
# - pymatgen==2017.9.23
# - python==2.7
#
# Note that a API key of materials project may be required.
from pymatgen.analysis.phase_diagram import PDPlotter
plotter = PDPlotter(pd, show_unstable=True)
plotter.show()
# +
# # %matplotlib inline
import matplotlib
import pymatgen as mg
import os
from pymatgen.ext.matproj import MPRester
from pymatgen.analysis.phase_diagram import PhaseDiagram, PDPlotter
rester = mg.MPRester(os.environ['PMG_MAPI_KEY'])
# in your shell .profile/.bashrc as: export PMG_MAPI_KEY=" "
entries = rester.get_entries_in_chemsys(['Ca', 'C', 'O'])
phasediagram = PhaseDiagram(entries)
# -
#Let's show all phases, including unstable ones
plotter_1 = PDPlotter(phasediagram, show_unstable=True)
plotter_2 = PDPlotter(phasediagram, show_unstable=False)
plotter_1.show()
plotter_2.show()
# ## Calculating energy above hull and other phase equilibria properties
#
# To perform more sophisticated analyses, use the PDAnalyzer object.
# +
import collections
data = collections.defaultdict(list)
for e in entries:
decomp, ehull = phasediagram.get_decomp_and_e_above_hull(e)
data["Materials ID"].append(e.entry_id)
data["Composition"].append(e.composition.reduced_formula)
data["Ehull"].append(ehull)
data["Decomposition"].append(" + ".join(["%.2f %s" % (v, k.composition.formula) for k, v in decomp.items()]))
from pandas import DataFrame
df = DataFrame(data, columns=["Materials ID", "Composition", "Ehull", "Decomposition"])
print(df)
# -
# +
pmg_structure = rester.get_structure_by_material_id(structure_id)
pmg_band = rester.get_bandstructure_by_material_id(structure_id)
material_name = pmg_structure.formula.replace('1','').replace(' ','')
spa = SpacegroupAnalyzer(pmg_structure)
# -
# +
# # %matplotlib inline
import matplotlib
import pymatgen as mg
import os
from pymatgen.ext.matproj import MPRester
from pymatgen.analysis.phase_diagram import PhaseDiagram, PDPlotter
rester = mg.MPRester(os.environ['PMG_MAPI_KEY'])
# in your shell .profile/.bashrc as: export PMG_MAPI_KEY=" "
entries = rester.get_entries_in_chemsys(['Bi', 'Pb', 'Ti', 'O'])
phasediagram = PhaseDiagram(entries)
# -
#Let's show all phases, including unstable ones
plotter_1 = PDPlotter(phasediagram, show_unstable=True)
plotter_2 = PDPlotter(phasediagram, show_unstable=False)
plotter_1.show()
plotter_2.show()
# # References
#
# - Convex hull and compositional phase diagram by QZhu: https://github.com/qzhu2017/CMS/wiki/Convex-hull-and-compositional-phase-diagram and his code: https://github.com/qzhu2017/CMS
#
# - Phase Diagram App Manual of Materials Project: https://wiki.materialsproject.org/Phase_Diagram_App_Manual
#
# - Academic Paper: The Quickhull Algorithm for Convex Hull https://www.cise.ufl.edu/~ungor/courses/fall06/papers/QuickHull.pdf
#
| notebooks/20180427-phase-diagram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: dspy3
# language: python
# name: dspy3
# ---
# +
# %matplotlib inline
from matplotlib.collections import LineCollection
import matplotlib.pyplot as plt
from scipy import interpolate
import numpy as np
from numpy.random import rand
from ipywidgets import FloatSlider, interactive, IntSlider
# -
def simple_example(amplitude=2.0, phase=0.0):
plt.rcParams['figure.figsize'] = 8,6
plt.figure()
x = np.linspace(-2*np.pi, 2*np.pi, 1000)
y = amplitude * np.sin(x + phase)
plt.plot(x, y)
plt.xlim(-3, 3)
plt.ylim(-2*np.pi, 2*np.pi)
plt.show()
return
# +
amplitude_slider = FloatSlider(value=2.0, min=0, max=6.0, step=.1)
phase_slider = FloatSlider(value=0.0, min=-np.pi, max=np.pi, step=.10)
interactive(simple_example,
amplitude=amplitude_slider,
phase=phase_slider
)
# +
def spline_demo(num=14, smooth=0, seed=10, brush_strokes=30, alpha=0.5):
a = np.random.RandomState(seed=seed)
x = a.rand(num)
y = a.rand(num)
t = np.arange(0, 1.1, .1)
plt.rcParams['figure.figsize'] = 8, 8
plt.figure()
for brush_stroke in range(brush_strokes):
tck, u = interpolate.splprep(
[x + a.rand(num) / 10.0, y + a.rand(num) / 10.0], s=smooth)
unew = np.arange(0, 1.01, 0.001)
out = interpolate.splev(unew, tck)
plt.plot(out[0], out[1], alpha=alpha, c='black', linewidth=3.0)
plt.xlim(-1.5, 2.)
plt.ylim(-1.5, 2.)
plt.axis('off')
plt.show()
smooth_slider = FloatSlider(value=0, min=0, max=20.0, step=.1)
num_points_slider = IntSlider(value=8, min=4, max=20)
seed_slider = IntSlider(value=4, min=4, max=20)
brush_slider = IntSlider(value=1, min=1, max=20)
alpha_slider = FloatSlider(value=.5, min=0, max=1.0, step=.05)
w = interactive(
spline_demo,
smooth=smooth_slider,
num=num_points_slider,
seed=seed_slider,
brush_strokes=brush_slider,
alpha=alpha_slider)
w
# -
| notebooks/03-Interactive-Splines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import json
import ast
import numpy as np
labelled_data = pd.read_csv("/home/mannu/Downloads/f1265970.csv")
labelled_agg_data = pd.read_csv("/home/mannu/Downloads/a1265970.csv")
print labelled_data.shape
print labelled_agg_data.shape
labelled_data.head()
print len(labelled_data.title.unique()), labelled_data.shape
a = pd.DataFrame({'count' : labelled_data.groupby( [ "url"] ).size()}).reset_index()
a.head()
dic = {}
for n in a["count"]:
if n in dic:
dic[n] = dic[n] + 1
else:
dic[n] = 1
dic
a.url[0]
labelled_data[labelled_data.url == "http://www.asiaone.com/asia/debate-ensues-after-alleged-suicide-attempt-singapore-airlines-passenger-incheon-airport"]
labelled_agg_data
a = pd.read_csv("/home/mannu/Downloads/f1265970.csv")
b = pd.read_csv("/home/mannu/Downloads/a1265970.csv")
a.shape, b.shape
a
b
len(a["url"].unique()), len(b["url"].unique())
_ = pd.DataFrame({'count' : b.groupby( [ "url"] ).size()}).reset_index()
_.sort_values(by="count",ascending=False).head()
annotated_data = pd.read_csv("../../data/annotator_data_dump_with_text")
annotated_data = annotated_data.drop(["Unnamed: 0", "Unnamed: 0.1"],axis=1)
annotated_data.shape
_ = annotated_data.iloc[9]
_.url, _.title, _.summary, _.category
# ### Labelled data
gd = pd.read_csv("../../data/annotator_data_dump_with_text").drop(["Unnamed: 0", "Unnamed: 0.1"],axis=1)
gd.head()
classes = {91: 'health', 92: 'safetysecurity', 93 : 'environment',
94 : 'social_relations', 95 : 'meaning_in_life', 96 : 'achievement',
97 : 'economics', 98 : 'politics', 99 : 'not_applicable', 0 : 'skip' }
s = 0
count=0
for index, row in labelled_data.iterrows():
d = {}
for cat in row["check_one_or_more"].split("\n"):
d[cat] = 0
for cat in ast.literal_eval(gd[gd.url == row["url"]].category.values[0]):
cat = classes[int(cat)]
if cat in d:
d[cat] = 1
else:
d[cat] = 0
temp = 0
for ky in d.keys():
if d[ky] == 1:
temp = temp+1
s = s + float(temp)/len(d.keys())
#print s,d
s/labelled_data.shape[0]
row
labelled_data.check_one_or_more.unique()
"health\nsafetysecurity".split("")
_d = {"1":2}
for ky in _d.keys():
if _d[ky] == 2:
print "很 好"
# ## Find kappa
a = pd.DataFrame({'counts' : labelled_data.groupby( [ "url"] ).size()}).reset_index()
count = 0
d = {}
for url in a[a.counts == 3].url:
d[url] = count
count=count+1
mat = np.zeros((len(d.keys()),9))
_classes = {'health' : 0, 'safetysecurity' : 1, 'environment' : 2,
'social_relations' : 3, 'meaning_in_life' : 4, 'achievement' : 5,
'economics' : 6 , 'politics' : 7, 'not_applicable' : 8, 0 : 'skip' }
for url in d.keys():
for index, row in labelled_data[labelled_data.url == url].iterrows():
q = d[row["url"]]
categories = row["check_one_or_more"].split("\n")
for category in categories:
mat[q][_classes[category]] = mat[q][_classes[category]] + 1
mat[5]
labelled_data[labelled_data.url == d.keys()[2]]
d
labelled_data[labelled_data.url == "http://www.asiaone.com/health/cooler-weather-could-cause-risk-weaker-blood-flow-doctors"]
computeKappa(mat)
# +
DEBUG = False
def checkEachLineCount(mat):
""" Assert that each line has a constant number of ratings
@param mat The matrix checked
@return The number of ratings
@throws AssertionError If lines contain different number of ratings """
n = sum(mat[0])
assert all(sum(line) == n for line in mat[1:]), "Line count != %d (n value)." % n
return n
def computeKappa(mat):
""" Computes the Kappa value
@param n Number of rating per subjects (number of human raters)
@param mat Matrix[subjects][categories]
@return The Kappa value """
n = checkEachLineCount(mat) # PRE : every line count must be equal to n
N = len(mat)
k = len(mat[0])
if DEBUG:
print n, "raters."
print N, "subjects."
print k, "categories."
# Computing p[]
p = [0.0] * k
for j in xrange(k):
p[j] = 0.0
for i in xrange(N):
p[j] += mat[i][j]
p[j] /= N * n
if DEBUG:
print "p =", p
# Computing P[]
P = [0.0] * N
for i in xrange(N):
P[i] = 0.0
for j in xrange(k):
P[i] += mat[i][j] * mat[i][j]
P[i] = (P[i] - n) / (n * (n - 1))
if DEBUG:
print "P =", P
# Computing Pbar
Pbar = sum(P) / N
if DEBUG:
print "Pbar =", Pbar
# Computing PbarE
PbarE = 0.0
for pj in p:
PbarE += pj * pj
if DEBUG:
print "PbarE =", PbarE
if PbarE == 1:
kappa = 1
else:
kappa = (Pbar - PbarE) / (1 - PbarE)
if DEBUG:
print "kappa =", kappa
return kappa
# -
| micromort/analysis/figure 8 - crowd sourcing analysis.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.1
# language: julia
# name: julia-1.4
# ---
using Images,ImageView, Plots,LinearAlgebra,Statistics, FFTW,TestImages
img = rand(Float32,1000,1000)
# img = Gray.(testimage("mandrill"));
# kernel = ones(Float32,15,15);
kernel = rand(Float32,15,15);
size(img)
# # Constant
function pad_constant(img,kernel,constant=0)
kernel_h, kernel_w = size(kernel)
img_h, img_w = size(img)
padded_kernel= ones(img_h,img_w).*(1/(1+exp(-constant)));
pad_h, pad_w = size(padded_kernel)
center_x,center_y = pad_w ÷2, pad_h ÷2
tmp_x = center_x-(kernel_w÷2)
tmp_y = center_y-(kernel_h÷2)
padded_kernel[collect(tmp_x:tmp_x+kernel_w-1),collect(tmp_y:tmp_y+kernel_h-1)] = kernel;
return padded_kernel
end
Gray.(pad_constant(img,kernel))
ker_pad = pad_constant(img, kernel, .3);
x = Gray.(real.(fft(ker_pad)))
y = Gray.(real.(fft(channelview(img))))
img = rand(Float32,1000,1000);
kernel = rand(Float32,15,15);
@time ifft(fft(channelview(img)).*fft(ker_pad))
0.536841/ 0.063075
# # GPU
using CUDA.CUFFT,CUDA,Images
function pad_constant(img,kernel,constant=0)
kernel_h, kernel_w = size(kernel)
img_h, img_w = size(img)
padded_kernel= ones(img_h,img_w)*(1/(1+exp(-constant)));
pad_h, pad_w = size(padded_kernel)
center_x,center_y = pad_w ÷2, pad_h ÷2
tmp_x = center_x-(kernel_w÷2)
tmp_y = center_y-(kernel_h÷2)
padded_kernel[collect(tmp_x:tmp_x+kernel_w-1),collect(tmp_y:tmp_y+kernel_h-1)] = kernel;
return padded_kernel
end
img = rand(9000,9000)
kernel = rand(Float32,15,15);
ker_pad = pad_constant(img, rand(Float32,15,15), .3);
CUDA.allowscalar(false)
@time ifft(fft(CuArray(img))*fft(CuArray(ker_pad)))
0.536841/ 0.002228
CUBLAS.dot
using Images,ImageView, Plots,LinearAlgebra,Statistics,TestImages
img = rand(Float32,50,50)
# img = Gray.(testimage("mandrill"));
kernel = ones(Float32,15,15);
# kernel = rand(Float32,15,15);
# # Max
Gray.(pad_constant(img,kernel,maximum(kernel)))
# # Mean
pad_constant(img,kernel,mean(kernel))
# # Median
Gray.(pad_constant(img,kernel,median(kernel)))
# # Min
pad_constant(img,kernel,minimum(kernel))
# # Edge
# # Linear ramp
# # Reflect
# # Symmetric
# # Wrap
| padding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from multitcn_components import TCNStack, DownsampleLayerWithAttention, LearningRateLogger
import tensorflow as tf
from tensorflow.keras.callbacks import ReduceLROnPlateau, LearningRateScheduler, EarlyStopping, ModelCheckpoint, CSVLogger
from sklearn import preprocessing
import numpy as np
import pandas as pd
from datetime import datetime
import uuid
import sys
import tensorflow_addons as tfa
### Set experiment seed
print("Enter a seed for the experiment:")
seed = input()
if len(seed)!=0 and seed.isdigit():
seed = int(seed)
else:
seed = 192
np.random.seed(seed)
tf.random.set_seed(seed)
# +
def log_experiment_details(just_print=False, complete=False):
""" Log experiment details """
if just_print:
o = sys.stdout
else:
o = open(date_time_string+"--"+experiment_id+".txt","w")
o.write("Date and time: %s \n" % date_time_string)
o.write("Experiment complete : %s\n" % str(experiment_complete))
if complete:
now = datetime.now()
completion_time = now.strftime("%d-%m-%Y-%H-%M-%S")
o.write("Experiment complete at: %s\n" % completion_time)
o.write("Duration in seconds: %f \n"% duration)
o.write("Filename: %s\n" % str(sys.argv[0]))
o.write("\n\n")
o.write(" Training parameters ".center(100,"="))
o.write("\n\n")
o.write("Batch size: %d\n" % batch_size)
o.write("Epochs : %d\n" % epochs)
o.write("Device used: %s\n" % device)
o.write("Random seed used: %s\n" % seed)
o.write("Loss : %s\n" % loss)
o.write("Optimizer config: %s" % str(optimizer.get_config()))
o.write("\n\n")
o.write(" Dataset parameters ".center(100,"="))
o.write("\n\n")
o.write("Dataset description: %s\n" % dataset_description)
o.write("Number of input time series: %d\n" % num_input_time_series)
o.write("Window length: %d\n" % window_length)
o.write("Total sample size: %d \n"% total_samples)
o.write("Training samples: %d\n" % int(train_x.shape[0]*training_percentage))
o.write("Training start date: %s\n" % training_start_date)
o.write("Validation samples: %d\n" % int(train_x.shape[0]*(1-training_percentage)))
o.write("Test samples: %d\n" % test_x.shape[0])
o.write("Test start date: %s\n" % holdout_set_start_date)
o.write("Test end date: %s\n" % holdout_set_end_date)
o.write("Experiment target: %s\n" % experiment_target)
o.write("Dataset preprocessing: %s\n" % dataset_preprocessing)
o.write("Shuffled training and val set: %s\n" % shuffle_train_set)
o.write("Scaled output: %s\n" % scale_output)
o.write("Input preprocessor details: %s, %s\n" %(preprocessor.__class__, preprocessor.get_params()))
o.write("Output scaler details: %s, %s\n" %(out_preprocessor.__class__, out_preprocessor.get_params()))
o.write("\n\n")
o.write(" Specific model parameters ".center(100,"="))
o.write("\n\n")
o.write("tcn_kernel_size : %d\n" % tcn_kernel_size)
o.write("tcn_filter_num : %d\n" % tcn_filter_num)
o.write("tcn_layer_num : %d\n" % tcn_layer_num)
o.write("tcn_use_bias : %s\n" % str(tcn_use_bias))
o.write("tcn_kernel_initializer : %s\n" % tcn_kernel_initializer)
o.write("tcn_dropout_rate : %0.2f\n" % tcn_dropout_rate)
o.write("tcn_dropout_format : %s\n" % tcn_dropout_format)
o.write("tcn_activation : %s\n" % tcn_activation)
o.write("tcn_final_activation : %s\n" % tcn_final_activation)
o.write("tcn_final_stack_activation : %s\n" % tcn_final_stack_activation)
o.write("\n\n")
o.write(" Additional useful notes ".center(100,"="))
o.write("\n\n")
o.write(additional_experiment_notes)
o.write("\n\n")
if not just_print:
o.close()
else:
o.flush()
def windowed_dataset(series, time_series_number, window_size):
"""
Returns a windowed dataset from a Pandas dataframe
"""
available_examples= series.shape[0]-window_size + 1
time_series_number = series.shape[1]
inputs = np.zeros((available_examples,window_size,time_series_number))
for i in range(available_examples):
inputs[i,:,:] = series[i:i+window_size,:]
return inputs
def windowed_forecast(series, forecast_horizon):
available_outputs = series.shape[0]- forecast_horizon + 1
output_series_num = series.shape[1]
output = np.zeros((available_outputs,forecast_horizon, output_series_num))
for i in range(available_outputs):
output[i,:]= series[i:i+forecast_horizon,:]
return output
def shuffle_arrays_together(a,b):
p = np.random.permutation(a.shape[0])
return a[p],b[p]
# +
####### Set up experiment parameters ###############
#Logging parameters
experiment_id = str(uuid.uuid4())
now = datetime.now()
date_time_string = now.strftime("%d-%m-%Y-%H-%M-%S")
# For potential tensorboard use
log_dir="logs/profile/" + date_time_string
tensorboard_callback = tf.keras.callbacks.TensorBoard(log_dir=log_dir, histogram_freq=1, profile_batch = 3)
#Training parameters
epochs = 120
batch_size = 64
starting_lr = 1e-3
optimizer = tfa.optimizers.AdamW(learning_rate=starting_lr, weight_decay=1e-4)
min_lr = 2e-5
loss = 'mse'
## Callbacks
log_name="logs/"+ F"{experiment_id}-{date_time_string}_train_history"
log_history_callback = CSVLogger(log_name)
filepath= experiment_id+"-weights.{epoch:02d}-{val_loss:.4f}.h5"
save_model_callback = ModelCheckpoint(filepath, monitor='val_loss', verbose=0, save_best_only=True, save_weights_only=False, mode='auto', save_freq='epoch')
# Unused callbacks after all
# lr_log_name = "logs/"+ F"{experiment_id}-{date_time_string}_learning_rate_history"
# lr_log_callback = LearningRateLogger(lr_log_name)
#lr_reducer_callback = ReduceLROnPlateau(factor=0.5,cooldown=0,patience=25, min_lr=min_lr, verbose=1, min_delta=0.001)
#lr_schedule = LearningRateScheduler(lambda epoch: 1e-8 * 10**(epoch / 20))
# early_stopping_callback = EarlyStopping(monitor='val_loss', patience=40, min_delta=0.001)
callbacks_list = [log_history_callback, save_model_callback]#, lr_reducer_callback]#, lr_log_callback, early_stopping_callback]
#Dataset parameters
window_length = 96
forecast_horizon = 24
preprocessor = preprocessing.MinMaxScaler()
out_preprocessor = preprocessing.MinMaxScaler()
shuffle_train_set = True
scale_output = True
training_percentage = 0.75
experiment_target = F"Forecasting,{forecast_horizon} steps ahead"
experiment_complete = False
# -
############## Set up model ##########################
class MTCNAModel(tf.keras.Model):
def __init__(self, tcn_layer_num,tcn_kernel_size,tcn_filter_num,window_size,forecast_horizon,num_output_time_series, use_bias, kernel_initializer, tcn_dropout_rate,tcn_dropout_format,tcn_activation, tcn_final_activation, tcn_final_stack_activation):
super(MTCNAModel, self).__init__()
self.num_output_time_series = num_output_time_series
#Create stack of TCN layers
self.lower_tcn = TCNStack(tcn_layer_num,tcn_filter_num,tcn_kernel_size,window_size,use_bias,kernel_initializer,tcn_dropout_rate,tcn_dropout_format,tcn_activation,tcn_final_activation, tcn_final_stack_activation)
self.downsample_att = DownsampleLayerWithAttention(num_output_time_series,window_size, tcn_kernel_size, forecast_horizon, kernel_initializer, None)
def call(self, input_tensor):
x = self.lower_tcn(input_tensor)
x, distribution = self.downsample_att([x,input_tensor[:,:,:self.num_output_time_series]])
return [x[:,i,:] for i in range(self.num_output_time_series)]#, distribution
# +
################ Prepare dataset ###########################
### Note details for logging purposes
dataset_description = "Italian air quality data"
dataset_preprocessing = """Drop time information, Remove NAN rows at end, Replace missing values with 0"""
data = pd.read_csv("Datasets/AirQualityUCI.csv",sep=';',decimal=',')
## Remove NaN rows
data = data.drop(np.arange(9357,9471,1))
# Remove emtpy columns
data = data.drop(['Unnamed: 15','Unnamed: 16'],axis=1)
#Create date object for easy splitting according to dates
dateobj = pd.to_datetime(data["Date"],dayfirst=True) + pd.to_timedelta(data["Time"].str.replace(".00.00",":00:00"))
### For now remove timestamp and output values
data = data.drop(columns=["Date","Time"],axis=1)
#Drop column due to high number of missing values
data = data.drop(['NMHC(GT)'],axis=1)
# Replace missing values with 0
data = data.replace(-200,0)
# Reorganize columns in preparation for second stage (first columns are in order of outputs)
columns = ['CO(GT)','C6H6(GT)','NOx(GT)','NO2(GT)','PT08.S1(CO)','PT08.S2(NMHC)','PT08.S3(NOx)','PT08.S4(NO2)','PT08.S5(O3)','T','RH','AH']
data = data[columns]
## Add date object for splitting
data['DateObj'] = dateobj
# +
#Split data based on dates
training_start_date = pd.Timestamp(year=2004,month=3,day=10)
# Preceding values used only for creating final graph and predicting first values of test set
holdout_preceding_date = pd.Timestamp(year=2004, month=11, day=11)
holdout_set_start_date = pd.Timestamp(year=2004, month=12, day=11)
holdout_set_end_date = pd.Timestamp(year=2005, month=4, day=5)
training_data = data.loc[(data['DateObj']>=training_start_date) & (data['DateObj'] < holdout_set_start_date)]
test_data = data.loc[(data['DateObj'] >= holdout_set_start_date) & (data['DateObj'] < holdout_set_end_date)]
pre_evaluation_period = data.loc[(data['DateObj'] >= holdout_preceding_date) & (data['DateObj'] < holdout_set_start_date)]
input_variables = list(training_data.columns)
training_data = training_data.drop(['DateObj'],axis=1)
test_data = test_data.drop(['DateObj'],axis=1)
# +
##Select prediction target
targets = ['CO(GT)','C6H6(GT)','NOx(GT)','NO2(GT)']
labels = np.array(training_data[targets])
if scale_output:
out_preprocessor.fit(labels)
if "Normalizer" in str(out_preprocessor.__class__):
## Save norm so in case of normalizer we can scale the predictions correctly
out_norm = np.linalg.norm(labels)
labels = preprocessing.normalize(labels,axis=0)
else:
labels= out_preprocessor.transform(labels)
num_input_time_series = training_data.shape[1]
### Make sure data are np arrays in case we skip preprocessing
training_data = np.array(training_data)
### Fit preprocessor to training data
preprocessor.fit(training_data)
if "Normalizer" in str(preprocessor.__class__):
## Save norm so in case of normalizer we can scale the test_data correctly
in_norm = np.linalg.norm(training_data,axis=0)
training_data = preprocessing.normalize(training_data,axis=0)
else:
training_data = preprocessor.transform(training_data)
# +
### Create windows for all data
data_windows = windowed_dataset(training_data[:-forecast_horizon],num_input_time_series,window_length)
label_windows = windowed_forecast(labels[window_length:],forecast_horizon)
### Transpose outputs to agree with model output
label_windows = np.transpose(label_windows,[0,2,1])
samples = data_windows.shape[0]
## Shuffle windows
if shuffle_train_set:
data_windows, label_windows = shuffle_arrays_together(data_windows,label_windows)
### Create train and validation sets
train_x = data_windows
train_y = [label_windows[:,i,:] for i in range(len(targets))]
## In order to use all days of test set for prediction, append training window from preceding period
pre_test_train = pre_evaluation_period[test_data.columns][-window_length:]
test_data = pd.concat([pre_test_train,test_data])
## Create windowed test set with same process
test_labels = np.array(test_data[targets])
#### Preprocess data
test_data = np.array(test_data)
if "Normalizer" in str(preprocessor.__class__):
test_data = test_data/in_norm
else:
test_data = preprocessor.transform(test_data)
test_x = windowed_dataset(test_data[:-forecast_horizon],num_input_time_series,window_length)
test_y = np.transpose(windowed_forecast(test_labels[window_length:],forecast_horizon),[0,2,1])
## Create pre test period for visualization
pre_test_target = np.append(np.array(pre_evaluation_period[targets]),test_labels[:window_length])
total_samples = train_x.shape[0] + test_x.shape[0]
# -
##################### Initialize model parameters ########################
## For simplicity all time series TCNs have the same parameters, though it is relatively easy to change this
tcn_kernel_size = 3
tcn_layer_num = 5
tcn_use_bias = True
tcn_filter_num = 128
tcn_kernel_initializer = 'random_normal'
tcn_dropout_rate = 0.3
tcn_dropout_format = "channel"
tcn_activation = 'relu'
tcn_final_activation = 'linear'
tcn_final_stack_activation = 'relu'
loss = [loss]*len(targets)
# +
# ### Check for GPU
## Make only given GPU visible
gpus = tf.config.experimental.list_physical_devices('GPU')
mirrored_strategy = None
print("GPUs Available: ", gpus)
if len(gpus)==0:
device = "CPU:0"
else:
print("Enter number of gpus to use:")
gpu_num = input()
if len(gpu_num)!=0 and gpu_num.isdigit():
gpu_num = int(gpu_num)
if gpu_num==1:
print("Enter index of GPU to use:")
gpu_idx = input()
if len(gpu_idx)!=0 and gpu_idx.isdigit():
gpu_idx = int(gpu_idx)
tf.config.experimental.set_visible_devices(gpus[gpu_idx], 'GPU')
device = "GPU:0"
else:
mirrored_strategy = tf.distribute.MirroredStrategy(devices=[F"GPU:{i}" for i in range(gpu_num)])
device = " ".join([F"GPU:{i}" for i in range(gpu_num)])
# -
# ##### Initialize model
if mirrored_strategy:
with mirrored_strategy.scope():
model = MTCNAModel(tcn_layer_num,tcn_kernel_size,tcn_filter_num,window_length,forecast_horizon,len(targets), tcn_use_bias, tcn_kernel_initializer, tcn_dropout_rate, tcn_dropout_format, tcn_activation, tcn_final_activation, tcn_final_stack_activation)
model.compile(optimizer,loss,metrics=[tf.keras.metrics.RootMeanSquaredError()])
else:
with tf.device(device):
model = MTCNAModel(tcn_layer_num,tcn_kernel_size,tcn_filter_num,window_length,forecast_horizon,len(targets), tcn_use_bias, tcn_kernel_initializer, tcn_dropout_rate, tcn_dropout_format, tcn_activation, tcn_final_activation, tcn_final_stack_activation)
model.compile(optimizer,loss,metrics=[tf.keras.metrics.RootMeanSquaredError()])
additional_experiment_notes = ""
log_experiment_details(just_print=True)
############## Note here any special notes about this experiment
print("Do you have any special notes for this experiment ? If yes, please write them here (or press Enter):")
additional_experiment_notes = input()
if len(additional_experiment_notes)==0:
additional_experiment_notes = """No special notes"""
# +
#### Create log and start training
log_experiment_details()
start_time = tf.timestamp()
history = model.fit(x=train_x, y= train_y,validation_split=(1-training_percentage), batch_size=batch_size, epochs=epochs, callbacks=callbacks_list)
end_time = tf.timestamp()
duration = end_time - start_time
experiment_complete = True
log_experiment_details(complete=True)
# -
| Suggested-AirQ-Training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# catalogue_local - catalog imagery on a local directory
# +
import datetime
import os
# dt = datetime.date.fromtimestamp(int(f.split('.')[0]))
# print(dt.strftime("%A, %d %B %Y"))
cams = ['c1', 'c2']
types = ['snap','var','timex','bright','dark','rundark']
imagedir = r"C:\\crs\\proj\\2020_PEP_SSF\\practice_folder"
for root, dirs, files in os.walk(imagedir):
print("{0} has {1} files".format(root, len(files)))
# -
| catalogue_local.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # One variable continuous data in `matplotlib` and `seaborn` <img src="images/JHI_STRAP_Web.png" style="width: 150px; float: right;">
# <div class="alert alert-success">
# <h2>Introduction</h2>
# </div>
# This notebook exercise will describe visualisation of one-dimensional continuous data in `Python`, using the `matplotlib` and `seaborn` libraries.
#
# For these examples, we will work with randomly-sampled data from statistical distributions that you will generate using this notebook.
# #### Python imports
#
# To view images in the notebook, we run a cell containing the line:
#
# ```python
# # %matplotlib inline
# ```
#
# This will make `matplotlib` and `NumPy` libraries available to us in the notebook.
# `Seaborn` can be imported with the following code:
#
# ```python
# import seaborn as sns
# ```
# +
# Run the pylab magic in this cell
# %matplotlib inline
import matplotlib.pyplot as plt
# Import seaborn
import seaborn as sns
# Random number generation
import random
# -
# <div class="alert alert-warning">
# <b>We will turn off distracting warning messages</b>
# </div>
#
# ```python
# import warnings
# warnings.filterwarnings('ignore')
# ```
# Turn off warning messages
import warnings
warnings.filterwarnings('ignore')
# <div class="alert alert-warning">
# <b>NOTE:</b> Even if you do not explicitly plot using <b>seaborn</b> as <b>sns</b>, importing it causes style changes from the <b>matplotlib</b> defaults.
# </div>
#
# 
# ### Learning Outcomes
#
# * Generating randomly-distributed example data
# * Representing one-dimensional continuous-valued data with histograms, KDE plots, and rug plots
# * Using `matplotlib` and `seaborn` libraries
# * Presenting arrays of images
# * Use of `figure()` and `subplots()`
# <div class="alert alert-success">
# <h2>Exercise</h2>
# </div>
# <div class="alert alert-success">
# <h3>1. Creating a randomly-distributed 1D dataset</h3>
# </div>
# #### Creating the dataset
#
# The `Numpy` library contains a module called `random` which provides a number of utility functions for generating (pseudo-)random value data. This is imported when you use the `%pylab inline` magic, and you can find out more about this module with `help(random)`.
#
# We will use the `random.random()` function to generate 100 datapoints, which we will assign to the variable `data`. We will explore the distribution of this data graphically using `matplotlib`
#
# ```python
# data = [random.random() for _ in range(100)]
# ```
#
# Other random samplings are available, such as:
#
# * `random.randint(10, size=10)`: integers in a range
# * `random.randn(10)`: standard normal distribution
# * `random.negative_binomial(1, 0.1, 10)`: negative binomial distribution
#
# and so on. You should feel free to experiment with them using the code below
# Create the variable data, containing 100 random values, here
# <div class="alert alert-success">
# <h3>2. Histogram</h3>
# </div>
# #### Base histogram
#
# [Histograms](http://www.datavizcatalogue.com/methods/histogram.html) show the distribution of data over a continuous interval. The total area of the histogram is considered to represent the number of datapoints, where the area of any bar is proportional to the frequency of datapoints in that interval, or *bin*.
#
# You can draw a basic histogram of your dataset using the `hist()` function, as in the code below:
#
# ```python
# n, bins, patches = plt.hist(data)
# ```
# Draw basic histogram in this cell
# You will probably find this basic histogram to be pretty uninspiring. The colour choice is flat, and the whole appearance is quite blocky and uninformative. It also lacks a title or any other detail.
#
# The objects returned by the `hist()` function in the code above are:
#
# * `n`: an *array* of values for each bar in the histogram, reflecting its height on the *y*-axis
# * `bins`: an *array* of the breakpoints for each bin (where the edges of each bin lie on the *x*-axis)
# * `patches`: objects representing the graphical elements for each bar; these can be manipulated to modify the plot's appearance
#
# You can explore the contents of these variables in the cell below
# Explore the contents of n, bins, and patches in this cell
# #### Normalised histograms
#
# The base histogram above reports absolute counts of the data in each bin. A normalised histogram reports *frequencies*, essentially modifying bin heights so that the integral of the histogram (summed area of all the bars) is equal to unity (1). This makes the histogram equivalent to a (stepped) probability density curve.
#
# To generate a normalised histogram, you can set `normed=1` in the call to `hist()`:
#
# ```python
# n, bins, patches = plt.hist(data, normed=1)
# ```
# Create a normalised histogram in this cell
# You will notice that the values on the *y*-axis have changed.
#
# <div class="alert alert-warning">
# <b>NOTE:</b> For some distributions you may notice that, perhaps counter-intuitively, some of the bars may extend past the value 1.0 on the <i>y</i>-axis. This can occur when the width of the bar is less than 1, as the area of each bar is the width of the bin multiplied by its height.
# </div>
# #### Modifying histogram appearance, and `subplots`
#
# We can modify several aspects of the histogram's appearance (see, e.g. `help(hist)` for documentation) in the call to `hist()`:
#
# * `bins`: the number of bins into which the data will be divided
# * `facecolor`: the colour of each rectangle in the histogram
# * `alpha`: the transparency value for the rectangles histogram (useful when layering plots)
#
# We'll try this with the code below:
#
# ```python
# n, bins, patches = plt.hist(data, normed=1, bins=20, facecolor='green', alpha=0.6)
# ```
# Modify the histogram appearance in this cell
# #### Stepfilling
#
# So far, our histograms have filled each of the bars with a border line, which makes the plot look more like a density plot. Other representations are available, such as: `histtype='step'`, that renders the bar chart as a boundary line:
#
# ```python
# n, bins, patches = plt.hist(data, normed=1, bins=20, facecolor='green',
# alpha=0.6, histtype='step')
# ```
# Create a step style histogram
# <div class="alert alert-success">
# <h3>3. Subplots and labels</h3>
# </div>
# We often wish to place multiple sets of axes on the same overall figure, to enable several related visualisations to be placed alongside each other. To do this, we place histograms - or any other plot - into *subplots*.
#
# The general approach for `matplotlib` is as follows:
#
# 1. Create a `Figure()` object
# 2. Add as many subplots (`Axes()`) to the `Figure()` as you require, specifying their location in a grid structure.
# 3. Add your visualisations to the subplot (`Axes()`) objects
#
# <div class="alert alert-warning">
# <b>NOTE:</b> An alternative approach to producing ordered arrangements of plots called <i>faceting</i> is available in other graphics libraries, and can be much simpler to use than this method. <i>Faceting</i> is dealt with in some of the other exercises in this set.
# </div>
#
# In the example below, we will create a 1x3 grid, and revisualise the three histograms from the examples above, with the code below:
#
# ```python
# # Create figure
# fig = plt.figure()
#
# # Create subplot axes
# ax1 = fig.add_subplot(1, 3, 1) # 1x3 grid, position 1
# ax2 = fig.add_subplot(1, 3, 2) # 1x3 grid, position 1
# ax3 = fig.add_subplot(1, 3, 3) # 1x3 grid, position 1
#
# # Add histograms
# ax1.hist(data)
# ax2.hist(data, normed=1)
# ax3.hist(data, normed=1, bins=20, facecolor='green', alpha=0.6)
# ```
# Create 1x3 subplot of histograms in this cell
# The first run of this code will probably be a bit disappointing, as the default aspect ratio will likely make the output image appear cramped and untidy. We can change the overall aspect ratio (and make the figure clearer) by setting a larger overall figure size.
#
# The overall output size of the figure can be set with `figsize=(width, height)`, where `width` and `height` are the output image value in inches. We can set this with the following code:
#
# ```python
# fig = plt.figure(figsize=(10, 3))
# ```
# Replot the subplots in this cell, with a larger figure size
# Modifications, such as addition of labels and titles, can be made to each individual axis in the figure, by calling methods for that plot axis specifically.
#
# Here, the first histogram (`ax1`) has a *y*-axis of absolute count, but the other two plots (`ax2`, `ax3`) have a *y*-axis of normalised frequency. All axes have `data` values along the *x*-axis.
#
# These axis objects are like any other Python object and we can treat them like any other object, so loop over them as in the code below:
#
# ```python
# # Set first axis y-label
# ax1.set_ylabel('count')
#
# # Set second, third axes y-labels
# for axis in (ax2, ax3):
# axis.set_ylabel('frequency')
#
# # Set all axes x-labels
# for axis in (ax1, ax2, ax3):
# axis.set_xlabel('data')
#
# # Set axis titles
# ax1.set_title('basic')
# ax2.set_title('normalised')
# ax3.set_title('modified')
# ```
#
# These changes will modify the figure *in memory*, but won't change any already-rendered images. To see the changes, you have to show the contents of `fig` in an output cell again. First though, apply the `tight_layout()` method so that the labels do not overlap the graphs:
#
# ```python
# fig.tight_layout()
# fig
# ```
#
# * `matplotlib` tight layout guide: [http://matplotlib.org/users/tight_layout_guide.html](http://matplotlib.org/users/tight_layout_guide.html)
# Add axis labels in this cell, and rerender the image
# <div class="alert alert-success">
# <h3>4. Density Plots</h3>
# </div>
# Histograms plot counts or frequencies of one-dimensional data. This gives a (necessarily) blocky representation of the raw data. An alternative representation is the *kernel density estimate* (KDE) plot which smooths the data (usually with a Gaussian basis function).
#
# * Kernel density estimation: [Wikipedia](https://en.wikipedia.org/wiki/Kernel_density_estimation)
#
# There is no KDE/density plot function in `matplotlib`, but the `kdeplot()` function in `seaborn` fits a KDE to a given dataset:
#
# ```python
# sns.kdeplot(data)
# ```
# Draw a seaborn kdeplot()
# `sns.kdeplot` offers quite a bit of control over the choice of kernel and bandwidth reference method for determination of kernel size. For more information see `help(sns.kdeplot)` or the documentation.
#
# * `sns.kdeplot()`: [documentation](https://stanford.edu/~mwaskom/software/seaborn/generated/seaborn.kdeplot.html)
#
# There are several options for kernel choice, e.g.:
#
# ```python
# sns.kdeplot(data, kernel='cos')
# sns.kdeplot(data, kernel='epa')
# ```
#
# and bandwidth can be varied, too:
#
# ```python
# sns.kdeplot(data, kernel='epa', bw=2)
# sns.kdeplot(data, kernel='epa', bw='silverman')
# ```
# Experiment with bandwidth and kernel choice for sns.kdeplot()
# #### Distribution plots with `seaborn`
#
# `seaborn` makes it easy to draw distribution plots combining three representations: histogram, KDE plot and *rug plot*, with the `sns.distplot()` function. By default, only the histogram and KDE plot are shown, but all three types can be controlled by specifying `hist=True`, `kde=True`, `rug=True` (or `False` in each case):
#
# ```python
# sns.distplot(data, rug=True)
# ```
#
# *Rug plots* draw small vertical ticks at each observation point, giving an alternative representation of data density.
#
# <div class="alert alert-warning">
# <b>NOTE:</b> if only the histogram is drawn, the <i>y</i>-axis reverts to count data. Including the KDE sets the <i>y</i> axis to frequency, and essentially forces normalisation of the histogram.
# </div>
#
# Use sns.distplot() to render the data with a rug plot
# #### Subplots with `seaborn`
#
# Although `seaborn` graphs cannot be added to figure subplots on creation in *exactly* the same way as `matplotlib` graphs, some (including `sns.distplot`) can still be added by specifying the `ax` argument, as follows:
#
# ```python
# sns.distplot(data, rug=True, ax=ax3, bins=20)
# ```
#
# So code to generate three renderings of the same data, side-by-side, could be:
#
# ```python
# # Create figure
# fig = plt.figure(figsize=(16, 6))
#
# # Create subplot axes
# ax1 = fig.add_subplot(1, 3, 1) # 1x3 grid, position 1
# ax2 = fig.add_subplot(1, 3, 2) # 1x3 grid, position 1
# ax3 = fig.add_subplot(1, 3, 3) # 1x3 grid, position 1
#
# # Set first axis y-label
# ax1.set_ylabel('count')
#
# # Set second, third axes y-labels
# for axis in (ax2, ax3):
# axis.set_ylabel('frequency')
#
# # Set all axes x-labels
# for axis in (ax1, ax2, ax3):
# axis.set_xlabel('data')
#
# # Set axis titles
# ax1.set_title('histogram')
# ax2.set_title('KDE')
# ax3.set_title('distribution')
#
# # Plot histogram, KDE and all histogram/KDE/rug on three axes
# sns.distplot(data, kde=False, ax=ax1)
# sns.distplot(data, hist=False, ax=ax2)
# sns.distplot(data, rug=True, ax=ax3, bins=20)
# ```
# Render three types of distribution plot with seaborn
| exercises/one_variable_continuous/one_variable_continuous.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from string import ascii_uppercase as uper, ascii_lowercase as lower, digits
import random
check = [959592808, 959852599, 960049253, 926430775, 892811314, 946419251, 929576502, 946419765, 909391664, 925972535, 892613940, 912864564, 12391]
# -
def unsage(check):
x=""
for item in check:
k = format(item, 'x')
x=x+bytes.fromhex(k).decode('utf-8')
return x
def unpush(a):
x=""
n=[]
for j in a:
n.append(j)
for i in n:
if i.isupper() is True:
x=x+upper[(upper.index(i)-3)%26]
elif i.islower() is True:
x=x+lower[(lower.index(i)-3)%26]
elif i.isdigit() is True:
x=x+digits[(digits.index(i)-3)%10]
return x
bytes.fromhex(unpush(unsage(check))).decode('utf-8')
| inctf19-quals/re/sol_patched.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: pyQDQ
# language: python
# name: pyqdq
# ---
from pyQuantumDevice.designer import *
import numpy as np
import matplotlib.pyplot as plt
planView = PlanView(120, 100)
planView.add_finger_gate("Barrier1", np.array([20, 50]), 10)
planView.add_finger_gate("Plunger1", np.array([40, 70]), 10)
planView.add_finger_gate("Barrier2", np.array([60, 50]), 10)
planView.add_finger_gate("Plunger2", np.array([80, 70]), 10)
planView.add_finger_gate("Barrier3", np.array([100, 50]), 10)
planView.add_back_bone(20, 10, {"back": [(20, 5), (60, 5), (100, 5)]})
planView.show()
print(planView.gates[-1].gatewire)
planView.gates[0].location
planView.depth
planView.gates[0].body_length
gate = FingerGate("test1", np.array([0, 0]), 1, 5)
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111)
gate.show(ax)
gate = FingerGate("test2", np.array([0, 0]), 1, 5, "back")
fig = plt.figure(figsize=(10, 10))
ax = fig.add_subplot(111)
gate.show(ax)
| examples/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.0 64-bit ('3.7')
# name: python3
# ---
# # <NAME>: Reinforcement Learning Primer
#
# In this tutorial, we will learn how to apply Reinforcement learning to a problem of path finding. The setting is inspired by [Peter and the Wolf](https://en.wikipedia.org/wiki/Peter_and_the_Wolf) musical fairy tale by Russian composer [<NAME>](https://en.wikipedia.org/wiki/Sergei_Prokofiev). It is a story about young pioneer Peter, who bravely goes out of his house to the forest clearing to chase the wolf. We will train machine learning algorithms that will help Peter to explore the surroinding area and build an optimal navigation map.
#
# First, let's import a bunch of userful libraries:
import matplotlib.pyplot as plt
import numpy as np
import random
import math
# ## Overview of Reinforcement Learning
#
# **Reinforcement Learning** (RL) is a learning technique that allows us to learn an optimal behaviour of an **agent** in some **environment** by running many experiments. An agent in this environment should have some **goal**, defined by a **reward function**.
#
# ## The Environment
#
# For simplicity, let's consider Peter's world to be a square board of size `width` x `height`. Each cell in this board can either be:
# * **ground**, on which Peter and other creatures can walk
# * **water**, on which you obviously cannot walk
# * **a tree** or **grass** - a place where you cat take some rest
# * **an apple**, which represents something Peter would be glad to find in order to feed himself
# * **a wolf**, which is dangerous and should be avoided
#
# To work with the environment, we will define a class called `Board`. In order not to clutter this notebook too much, we have moved all code to work with the board into separate `rlboard` module, which we will now import. You may look inside this module to get more details about the internals of the implementation.
from rlboard import *
# Let's now create a random board and see how it looks:
width, height = 8,8
m = Board(width,height)
m.randomize(seed=13)
m.plot()
# ## Actions and Policy
#
# In our example, Peter's goal would be to find an apple, while avoiding the wolf and other obstacles. To do this, he can essentially walk around until he finds and apple. Therefore, at any position he can chose between one of the following actions: up, down, left and right. We will define those actions as a dictionary, and map them to pairs of corresponding coordinate changes. For example, moving right (`R`) would correspond to a pair `(1,0)`.
actions = { "U" : (0,-1), "D" : (0,1), "L" : (-1,0), "R" : (1,0) }
action_idx = { a : i for i,a in enumerate(actions.keys()) }
# The strategy of our agent (Peter) is defined by a so-called **policy**. Let's consider the simplest policy called **random walk**.
#
# ## Random walk
#
# Let's first solve our problem by implementing a random walk strategy.
# + tags=[]
def random_policy(m):
return random.choice(list(actions))
def walk(m,policy,start_position=None):
n = 0 # number of steps
# set initial position
if start_position:
m.human = start_position
else:
m.random_start()
while True:
if m.at() == Board.Cell.apple:
return n # success!
if m.at() in [Board.Cell.wolf, Board.Cell.water]:
return -1 # eaten by wolf or drowned
while True:
a = actions[policy(m)]
new_pos = m.move_pos(m.human,a)
if m.is_valid(new_pos) and m.at(new_pos)!=Board.Cell.water:
m.move(a) # do the actual move
break
n+=1
walk(m,random_policy)
# -
# Let's run random walk experiment several times and see the average number of steps taken:
# +
def print_statistics(policy):
s,w,n = 0,0,0
for _ in range(100):
z = walk(m,policy)
if z<0:
w+=1
else:
s += z
n += 1
print(f"Average path length = {s/n}, eaten by wolf: {w} times")
print_statistics(random_policy)
# -
# ## Reward Function
#
# To make our policy more intelligent, we need to understand which moves are "better" than others.
#
#
# +
move_reward = -0.1
goal_reward = 10
end_reward = -10
def reward(m,pos=None):
pos = pos or m.human
if not m.is_valid(pos):
return end_reward
x = m.at(pos)
if x==Board.Cell.water or x == Board.Cell.wolf:
return end_reward
if x==Board.Cell.apple:
return goal_reward
return move_reward
# -
# ## Q-Learning
#
# Build a Q-Table, or multi-dimensional array. Since our board has dimensions `width` x `height`, we can represent Q-Table by a numpy array with shape `width` x `height` x `len(actions)`:
Q = np.ones((width,height,len(actions)),dtype=np.float)*1.0/len(actions)
# Pass the Q-Table to the plot function in order to visualize the table on the board:
m.plot(Q)
# ## Essence of Q-Learning: Bellman Equation and Learning Algorithm
#
# Write a pseudo-code for our leaning algorithm:
#
# * Initialize Q-Table Q with equal numbers for all states and actions
# * Set learning rate $\alpha\leftarrow 1$
# * Repeat simulation many times
# 1. Start at random position
# 1. Repeat
# 1. Select an action $a$ at state $s$
# 2. Exectute action by moving to a new state $s'$
# 3. If we encounter end-of-game condition, or total reward is too small - exit simulation
# 4. Compute reward $r$ at the new state
# 5. Update Q-Function according to Bellman equation: $Q(s,a)\leftarrow (1-\alpha)Q(s,a)+\alpha(r+\gamma\max_{a'}Q(s',a'))$
# 6. $s\leftarrow s'$
# 7. Update total reward and decrease $\alpha$.
#
# ## Exploit vs. Explore
#
# The best approach is to balance between exploration and exploitation. As we learn more about our environment, we would be more likely to follow the optimal route, however, choosing the unexplored path once in a while.
#
# ## Python Implementation
#
# Now we are ready to implement the learning algorithm. Before that, we also need some function that will convert arbitrary numbers in the Q-Table into a vector of probabilities for corresponding actions:
def probs(v,eps=1e-4):
v = v-v.min()+eps
v = v/v.sum()
return v
# We add a small amount of `eps` to the original vector in order to avoid division by 0 in the initial case, when all components of the vector are identical.
#
# The actual learning algorithm we will run for 5000 experiments, also called **epochs**:
# +
from IPython.display import clear_output
lpath = []
for epoch in range(10000):
clear_output(wait=True)
print(f"Epoch = {epoch}",end='')
# Pick initial point
m.random_start()
# Start travelling
n=0
cum_reward = 0
while True:
x,y = m.human
v = probs(Q[x,y])
a = random.choices(list(actions),weights=v)[0]
dpos = actions[a]
m.move(dpos,check_correctness=False) # we allow player to move outside the board, which terminates episode
r = reward(m)
cum_reward += r
if r==end_reward or cum_reward < -1000:
print(f" {n} steps",end='\r')
lpath.append(n)
break
alpha = np.exp(-n / 3000)
gamma = 0.5
ai = action_idx[a]
Q[x,y,ai] = (1 - alpha) * Q[x,y,ai] + alpha * (r + gamma * Q[x+dpos[0], y+dpos[1]].max())
n+=1
# -
# After executing this algorithm, the Q-Table should be updated with values that define the attractiveness of different actions at each step. Visualize the table here:
m.plot(Q)
# ## Checking the Policy
#
# Since Q-Table lists the "attractiveness" of each action at each state, it is quite easy to use it to define the efficient navigation in our world. In the simplest case, we can just select the action corresponding to the highest Q-Table value:
# +
def qpolicy_strict(m):
x,y = m.human
v = probs(Q[x,y])
a = list(actions)[np.argmax(v)]
return a
walk(m,qpolicy_strict)
# -
# If you try the code above several times, you may notice that sometimes it just "hangs", and you need to press the STOP button in the notebook to interrupt it.
#
# > **Task 1:** Modify the `walk` function to limit the maximum length of path by a certain number of steps (say, 100), and watch the code above return this value from time to time.
#
# > **Task 2:** Modify the `walk` function so that it does not go back to the places where is has already been previously. This will prevent `walk` from looping, however, the agent can still end up being "trapped" in a location from which it is unable to escape.
# +
def qpolicy(m):
x,y = m.human
v = probs(Q[x,y])
a = random.choices(list(actions),weights=v)[0]
return a
print_statistics(qpolicy)
# -
# ## Investigating the Learning Process
plt.plot(lpath)
# What we see here is that at first the average path length increased. This is probably due to the fact that when we know nothing about the environment - we are likely to get trapped into bad states, water or wolf. As we learn more and start using this knowledge, we can explore the environment for longer, but we still do not know well where apples are.
#
# Once we learn enough, it becomes easier for the agent to achieve the goal, and the path length starts to decrease. However, we are still open to exploration, so we often diverge away from the best path, and explore new options, making the path longer than optimal.
#
# What we also observe on this graph, is that at some point the length increased abruptly. This indicates stochastic nature of the process, and that we can at some point "sploil" the Q-Table coefficients, by overwriting them with new values. This ideally should be minimized by decreasing learning rate (i.e. towards the end of training we only adjust Q-Table values by a small value).
#
# Overall, it is important to remember that the success and quality of the learning process significantly depends on parameters, such as leaning rate, learning rate decay and discount factor. Those are often called **hyperparameters**, to distinguish them from **parameters** which we optimize during training (eg. Q-Table coefficients). The process of finding best hyperparameter values is called **hyperparameter optimization**, and it deserves a separate topic.
# ## Exercise
# #### A More Realistic Peter and the Wolf World
#
# In our situation, Peter was able to move around almost without getting tired or hungry. In a more realistic world, he has to sit down and rest from time to time, and also to feed himself. Let's make our world more realistic by implementing the following rules:
#
# 1. By moving from one place to another, Peter loses **energy** and gains some **fatigue**.
# 2. Peter can gain more energy by eating apples.
# 3. Peter can get rid of fatigue by resting under the tree or on the grass (i.e. walking into a board location with a tree or grass - green field)
# 4. Peter needs to find and kill the wolf
# 5. In order to kill the wolf, Peter needs to have certain levels of energy and fatigue, otherwise he loses the battle.
#
# Modify the reward function above according to the rules of the game, run the reinforcement learning algorithm to learn the best strategy for winning the game, and compare the results of random walk with your algorithm in terms of number of games won and lost.
#
#
# > **Note**: You may need to adjust hyperparameters to make it work, especially the number of epochs. Because the success of the game (fighting the wolf) is a rare event, you can expect much longer training time.
#
#
| 8-Reinforcement/1-QLearning/solution/notebook.ipynb |