code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
from astropy.table import Table, vstack
from astropy.utils.data import download_file
from astropy.time import Time
from bs4 import BeautifulSoup
from urllib.request import HTTPError
base_url = "http://mkwc.ifa.hawaii.edu/archive/wx/ukirt/"
# +
tables = []
for i in range(1991, 2019):
print('Downloading year:', i)
try:
raw_table = open(download_file(base_url + 'ukirt-wx.{0}.dat'.format(i), cache=True)).read()
# Files before 2007 have a header row
if i < 2007:
table = Table.read(raw_table, format='ascii')
# The 2007 table is in a different format from the others, skip it
elif i in [2007, 2008]:
pass
# Files later than 2007 have no header
else:
table = Table.read(path, format='ascii.fixed_width_no_header', delimiter=' ')
# If the file is inaccessible, skip it and move on
except HTTPError:
pass
tables.append(table)
# +
columns = ['col1', 'col2', 'col3', 'col4']
labels = ['year', 'month', 'day', 'hour'] # figure this out...
for col_ind, label in zip(columns, labels):
if label not in tables[13].colnames:
tables[13].rename_column(col_ind, label)
# columns = ['col1', 'col2', 'col3', 'col4']
# labels = ['year', 'month', 'day', 'hour'] # figure this out...
# for col_ind, label in zip(columns, labels):
# if label not in tables[14].colnames:
# tables[14].rename_column(col_ind, label)
# columns = ['col1', 'col2', 'col3', 'col4']
# labels = ['year', 'month', 'day', 'hour'] # figure this out...
# for col_ind, label in zip(columns, labels):
# if label not in tables[15].colnames:
# tables[15].rename_column(col_ind, label)
# -
for table in tables:
if 'year' in table.colnames:
time_strings = ["{0}-{1}-{2} {3}:{4}:{5}"
.format(row['year'], row['month'], row['day'],
row['hour'], row['min'], row['sec'])
for row in table]
times = Time(time_strings, format='iso')
table['times'] = times
else:
time_strings = ["{0}-{1:02d}-{2} {3:02d}:{4:02d}"
.format(row['col1'], row['col2'], row['col3'],
row['col4'], row['col5']).replace(',', '')
for row in table]
times = Time(time_strings, format='iso')
table['times'] = times
# +
fig, ax = plt.subplots(figsize=(8, 4))
for table in tables:
if 'year' in table.colnames:
ax.plot_date(table['times'].plot_date, table['air_temp(C)'], marker='.', color='k', alpha=0.2)
else:
# Need to figure out how to handle this case when there are no column headers...
pass
plt.setp(ax.get_xticklabels(), rotation=45)
ax.set_ylim([-10, 20])
ax.set_xlabel('Time')
ax.set_ylabel('Air Temp [C]')
ax.set_title('UKIRT')
# -
| ukirt.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
from PIL import Image
from numpy import *
from pylab import *
import ncut
ncut = reload(ncut)
from scipy.misc import imresize
im = array(Image.open('C-uniform03.ppm'))
m, n = im.shape[:2]
wid = 50
rim = imresize(im, (wid, wid), interp='bilinear')
rim = array(rim, 'f')
A = ncut.ncut_graph_matrix(rim, sigma_d=1, sigma_g=1e-2)
code, V = ncut.cluster(A, k=3, ndim=3)
codeim = imresize(code.reshape(wid, wid), (m, n), interp='nearest')
figure()
imshow(codeim)
gray()
show()
code, V = ncut.cluster(A, k=4, ndim=3)
figure()
for i in range(4):
subplot(1, 4, i+1)
imshow(imresize(V[i].reshape(wid, wid), (m, n), interp='bilinear'))
axis('off')
show()
prime_img = V[0].reshape(wid, wid)
figure()
gray()
imshow(prime_img)
show()
from scipy.ndimage import filters
# +
sigma1 = 1
sigma2 = 3
im2 = filters.gaussian_filter(prime_img, sigma1)
im3 = filters.gaussian_filter(prime_img, sigma2)
im4 = im2 - im3
im5 = sqrt(im4**2)
figure(figsize=(16, 16))
gray()
subplot(3, 2, 1)
imshow(prime_img)
axis('off')
subplot(3, 2, 2)
imshow(im2)
axis('off')
subplot(3, 2, 3)
imshow(im3)
axis('off')
subplot(3, 2, 4)
imshow(im4)
axis('off')
subplot(3, 2, 5)
imshow(im5)
axis('off')
show()
# -
im6 = im4.copy()
threshold = 0
im6[im6<=threshold] = 0
im6[im6>threshold] = 1
im6 = 1 - im6
figure(0)
gray()
imshow(im6)
axis('off')
show()
| Chapter-9/CV Book Chapter 9 Exercise 7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import the datetime module
import datetime
# +
# initialize a datetime object
atime = datetime.datetime(2018,6,28,18,30)
print(atime)
print(type(atime))
later = atime + datetime.timedelta(hours=6)
print(later)
# +
# What time is it?
today = datetime.datetime.now()
print(today)
print(today - atime)
# +
# initialize a datetime object with an arbitrary formatted string
timestring = "21/11/17 16:30"
anothertime = datetime.datetime.strptime(timestring, "%d/%m/%y %H:%M")
newtimestring = anothertime.strftime('%Y-%m-%d_%H%M')
print(newtimestring)
# +
def timerange(begin, end):
result = []
pivot = begin
while pivot <= end:
result.append(pivot)
pivot += datetime.timedelta(days=1)
return result
timerange(datetime.datetime(2018,6,28), today)
# +
# list the file names in the data folder
import os
filelist = os.listdir('data')
print(filelist)
print(os.getcwd())
os.chdir('data/')
os.chdir('..')
os.mkdir('newfolder')
os.rmdir('newfolder')
os.makedirs('newfolder/newsubfolder/')
os.removedirs('newfolder/newsubfolder/')
#os.rename('A.txt', 'B.txt')
last_changed = os.stat('data/20180624_leipzig_categorize.nc').st_mtime # modification time
dt_last_changed = datetime.datetime.fromtimestamp(last_changed)
print(f'Modification time: {dt_last_changed:%Y%m%d %H:%M:%S}')
# +
# Write a function that returns a datetime object inizialized by a string containting some kind of formatted date
def dt_from_filename(filename):
return datetime.datetime.strptime(filename[:8], '%Y%m%d')
dt_from_filename(filelist[1])
# -
# list comprehension
[dt_from_filename(filename) for filename in filelist]
| notebooks_solution/part2_standardlib.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="RB8OAOdBc6Bz"
import pandas as pd
# + id="K8-gi7QKc7re" colab={"base_uri": "https://localhost:8080/", "height": 359} outputId="e0dc63be-975b-4085-c229-45c89721b23c"
movies = pd.read_csv('data.csv', usecols = ['id', 'name', 'description'])
movies.head(10)
# + colab={"base_uri": "https://localhost:8080/"} id="zZFx7f46uyF4" outputId="dea7a768-6e88-475d-9bee-9ce6826f1beb"
movies.info()
# + [markdown] id="rUdvL8qWdMks"
# ## Analysis
# + id="9gwHEkvtuoFu"
import matplotlib.pyplot as plt
from wordcloud import WordCloud, STOPWORDS
# + id="4TS52HKhdL51"
movies['name'] = movies['name'].astype('str')
name_corpus = ' '.join(movies['name'])
# + colab={"base_uri": "https://localhost:8080/", "height": 466} id="TMVLjqPndROe" outputId="3f96ec6f-f6ed-49f7-b4fe-2f20a35d8317"
name_wordcloud = WordCloud(stopwords = STOPWORDS, background_color = 'white', height = 2000, width = 4000).generate(name_corpus)
plt.figure(figsize = (16,8))
plt.imshow(name_wordcloud)
plt.axis('off')
plt.show()
# + id="BHcNIegLdPsB"
movies['description'] = movies['description'].astype('str')
description_corpus = ' '.join(movies['description'])
# + colab={"base_uri": "https://localhost:8080/", "height": 466} id="rEuryWQui-P3" outputId="7bafcf92-f3b3-41e0-9b98-6beb2eec493e"
description_wordcloud = WordCloud(stopwords = STOPWORDS, background_color = 'white', height = 2000, width = 4000).generate(description_corpus)
plt.figure(figsize = (16,8))
plt.imshow(description_wordcloud)
plt.axis('off')
plt.show()
# + [markdown] id="jXgfqXyhYnMG"
#
# + id="x_2K_0Ldilq0"
movies['content'] = movies[['name', 'description']].astype(str).apply(lambda x: ' // '.join(x), axis = 1)
movies['content'].fillna('Null', inplace = True)
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="FDux4vDoink-" outputId="17344731-89a8-4131-a3ab-43441ddb630b"
movies
# + [markdown] id="8O2V44A4jht5"
# ## Training
# + id="0jK51MWmuhZG"
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.metrics.pairwise import linear_kernel
# + [markdown] id="06v7Uf5ljp9Y"
# ### TF-IDF (Term Frequency - Inverse Document Frequency)
# https://en.wikipedia.org/wiki/Tf%E2%80%93idf
# + id="Zaq56HpkiqYn"
tf = TfidfVectorizer(analyzer = 'word', ngram_range = (1, 2), min_df = 0, stop_words = 'english')
tfidf_matrix = tf.fit_transform(movies['content'])
# + [markdown] id="Fz81YsI7jnIY"
# ### Cosine Similarity
#
# https://en.wikipedia.org/wiki/Cosine_similarity
#
# + id="kdpwpi9riuc8"
cosine_similarities = linear_kernel(tfidf_matrix, tfidf_matrix)
# + id="N8I_bQrUPO80"
# from sklearn.feature_extraction.text import CountVectorizer
# from sklearn.metrics.pairwise import cosine_similarity
# tf = CountVectorizer(stop_words = 'english')
# tfidf_matrix = tf.fit_transform(movies['content'])
# cosine_similarities = cosine_similarity(tfidf_matrix, tfidf_matrix)
# + [markdown] id="ixLjmNq1oA-S"
# store the top 100 similar items
# + id="NwAtDpDLiwFt"
results = {}
for idx, row in movies.iterrows():
similar_indices = cosine_similarities[idx].argsort()[:-100:-1]
similar_items = [(cosine_similarities[idx][i], movies['id'][i]) for i in similar_indices]
results[row['id']] = similar_items[1:]
# + id="Avk8VD4LRwbv" colab={"base_uri": "https://localhost:8080/"} outputId="7207962e-775e-4f42-950e-39021791e572"
type(results)
# + [markdown] id="8tLupLI2oIsl"
# ## Prediction
# + id="kjSSAqpioULf"
def getName(id):
return movies.loc[movies['id'] == id]['name'].tolist()
def getDesc(id):
return movies.loc[movies['id'] == id]['description'].tolist()
# + id="QIeiRaoiixXn"
def recommend(item_id, num):
print('{} movies similar to {}'.format(num, getName(item_id)))
print('---------------------------------------')
recs = results[item_id][:num]
for i, rec in enumerate(recs):
print('Movie Id: {}'.format(rec[1]))
print('score: {}'.format(rec[0]))
print('Name: {}'.format(getName(rec[1])))
print('Description: {}\n'.format(getDesc(rec[1])))
# + colab={"base_uri": "https://localhost:8080/"} id="gucU7jJ8iy1I" outputId="f0567830-9213-4232-e189-6f54c0f81f13"
recommend(item_id = 155, num = 5)
# + colab={"base_uri": "https://localhost:8080/"} id="_QnmpsQl3Coe" outputId="03706f59-e78e-4e1b-cf99-c7a11187ccb2"
recommend(item_id = 120, num = 5)
| notebooks/movie_recommander.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 203 Activation
#
# View more, visit my tutorial page: https://morvanzhou.github.io/tutorials/
# My Youtube Channel: https://www.youtube.com/user/MorvanZhou
#
# Dependencies:
# * torch: 0.5.0a0 (0.4.1+)
# * matplotlib
#
import torch
import torch.nn.functional as F
from torch.autograd import Variable
import matplotlib.pyplot as plt
# ### Firstly generate some fake data
x = torch.linspace(-5, 5, 200) # x data (tensor), shape=(200, 1) # torch.Tensor
x = Variable(x) #torch.Tensor
x_np = x.data.numpy() # numpy array for plotting # numpy.ndarray
# ### Following are popular activation functions
# +
# # deprecated, used in torch: 0.1.11
# y_relu = F.relu(x).data.numpy()
# y_sigmoid = F.sigmoid(x).data.numpy()
# y_tanh = F.tanh(x).data.numpy()
# y_softplus = F.softplus(x).data.numpy()
# # y_softmax = F.softmax(x)
y_relu = torch.relu(x).data.numpy()
y_sigmoid = torch.sigmoid(x).data.numpy()
y_tanh = torch.tanh(x).data.numpy()
y_softplus = F.softplus(x).data.numpy() #there's no softplus in torch
y_softmax = torch.softmax(x, dim=0).data.numpy()
# softmax is a special kind of activation function, it is about probability
# and will make the sum as 1.
# -
import numpy as np
np.sum(y_softmax)
# ### Plot to visualize these activation function
# %matplotlib inline
# +
plt.figure(1, figsize=(16, 6))
plt.subplot(231)
plt.plot(x_np, y_relu, c='red', label='relu')
plt.ylim((-1, 5))
plt.legend(loc='best')
plt.subplot(232)
plt.plot(x_np, y_sigmoid, c='red', label='sigmoid')
plt.ylim((-0.2, 1.2))
plt.legend(loc='best')
plt.subplot(233)
plt.plot(x_np, y_tanh, c='red', label='tanh')
plt.ylim((-1.2, 1.2))
plt.legend(loc='best')
plt.subplot(234)
plt.plot(x_np, y_softplus, c='red', label='softplus')
plt.ylim((-0.2, 6))
plt.legend(loc='best')
plt.subplot(235)
plt.plot(x_np, y_softmax, c='red', label='softmax')
plt.ylim((-0.05, 0.1))
plt.legend(loc='best')
plt.show()
# -
| tutorial-contents-notebooks/203_activation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="5fGL2VYp13Vi"
# # Analyzing interstellar reddening and calculating synthetic photometry
# + [markdown] colab_type="text" id="6UzIN7ds1-G1"
# ## Authors
#
# <NAME>, <NAME>, <NAME>, <NAME>
#
# Input from <NAME>, <NAME>, <NAME>, <NAME>
# + [markdown] colab_type="text" id="Btb9Da5P4nLr"
# ## Learning Goals
# - Investigate extinction curve shapes
# - Deredden spectral energy distributions and spectra
# - Calculate photometric extinction and reddening
# - Calculate synthetic photometry for a dust-reddened star by combining `dust_extinction` and `synphot`
# - Convert from frequency to wavelength with `astropy.unit` equivalencies
# - Unit support for plotting with `astropy.visualization`
#
#
# ## Keywords
# dust extinction, synphot, astroquery, units, photometry, extinction, physics, observational astronomy
#
# ## Companion Content
#
# * [Bessell & Murphy (2012)](https://ui.adsabs.harvard.edu/#abs/2012PASP..124..140B/abstract)
#
#
#
# ## Summary
#
# In this tutorial, we will look at some extinction curves from the literature, use one of those curves to deredden an observed spectrum, and practice invoking a background source flux in order to calculate magnitudes from an extinction model.
#
# The primary libraries we'll be using are [dust_extinction](https://dust-extinction.readthedocs.io/en/latest/) and [synphot](https://synphot.readthedocs.io/en/latest/), which are [Astropy affiliated packages](https://www.astropy.org/affiliated/).
#
# We recommend installing the two packages in this fashion:
# ```
# pip install synphot
# pip install dust_extinction
# ```
# This tutorial requires v0.7 or later of `dust_extinction`. To ensure that all commands work properly, make sure you have the correct version installed. If you have v0.6 or earlier installed, run the following command to upgrade
# ```
# pip install dust_extinction --upgrade
# ```
# + colab={} colab_type="code" id="vFDq1xGXz_t4"
import matplotlib.pyplot as plt
# %matplotlib inline
import numpy as np
import astropy.units as u
from astropy.table import Table
from dust_extinction.parameter_averages import CCM89, F99
from synphot import units, config
from synphot import SourceSpectrum,SpectralElement,Observation,ExtinctionModel1D
from synphot.models import BlackBodyNorm1D
from synphot.spectrum import BaseUnitlessSpectrum
from synphot.reddening import ExtinctionCurve
from astroquery.simbad import Simbad
from astroquery.mast import Observations
import astropy.visualization
# + [markdown] colab_type="text" id="ssir1UjtD_Wn"
# # Introduction
# + [markdown] colab_type="text" id="1rvwpEs946ju"
# Dust in the interstellar medium (ISM) extinguishes background starlight. The wavelength dependence of the extinction is such that short-wavelength light is extinguished more than long-wavelength light, and we call this effect *reddening*.
#
# If you're new to extinction, here is a brief introduction to the types of quantities involved.
# The fractional change to the flux of starlight is
# $$
# \frac{dF_\lambda}{F_\lambda} = -\tau_\lambda
# $$
#
# where $\tau$ is the optical depth and depends on wavelength. Integrating along the line of sight, the resultant flux is an exponential function of optical depth,
# $$
# \tau_\lambda = -\ln\left(\frac{F_\lambda}{F_{\lambda,0}}\right).
# $$
#
# With an eye to how we define magnitudes, we usually change the base from $e$ to 10,
# $$
# \tau_\lambda = -2.303\log\left(\frac{F_\lambda}{F_{\lambda,0}}\right),
# $$
#
# and define an extinction $A_\lambda = 1.086 \,\tau_\lambda$ so that
# $$
# A_\lambda = -2.5\log\left(\frac{F_\lambda}{F_{\lambda,0}}\right).
# $$
#
#
# There are two basic take-home messages from this derivation:
#
# * Extinction introduces a multiplying factor $10^{-0.4 A_\lambda}$ to the flux.
# * Extinction is defined relative to the flux without dust, $F_{\lambda,0}$.
#
# + [markdown] colab_type="text" id="K9gwBSy_R_2E"
# Once astropy and the affiliated packages are installed, we can import from them as needed:
# + [markdown] colab_type="text" id="NYgj1w2X7gXc"
# # Example 1: Investigate Extinction Models
# + [markdown] colab_type="text" id="ifscJWwyEZtS"
# The `dust_extinction` package provides various models for extinction $A_\lambda$ normalized to $A_V$. The shapes of normalized curves are relatively (and perhaps surprisingly) uniform in the Milky Way. The little variation that exists is often parameterized by the ratio of extinction ($A_V$) to reddening in the blue-visual ($E_{B-V}$),
# $$
# R_V \equiv \frac{A_V}{E_{B-V}}
# $$
#
# where $E_{B-V}$ is differential extinction $A_B-A_V$. In this example, we show the $R_V$-parameterization for the Clayton, Cardelli, & Mathis (1989, CCM) and the Fitzpatrick (1999) models. [More model options are available in the `dust_extinction` documentation.](https://dust-extinction.readthedocs.io/en/latest/dust_extinction/model_flavors.html)
# + colab={"base_uri": "https://localhost:8080/", "height": 358} colab_type="code" id="b1uhXbRl79FR" outputId="e8999675-8ac1-4398-deb2-3bc5d36342ae"
# Create wavelengths array.
wav = np.arange(0.1, 3.0, 0.001)*u.micron
for model in [CCM89, F99]:
for R in (2.0,3.0,4.0):
# Initialize the extinction model
ext = model(Rv=R)
plt.plot(1/wav, ext(wav), label=model.name+' R='+str(R))
plt.xlabel('$\lambda^{-1}$ ($\mu$m$^{-1}$)')
plt.ylabel('A($\lambda$) / A(V)')
plt.legend(loc='best')
plt.title('Some Extinction Laws')
plt.show()
# + [markdown] colab_type="text" id="d-1XRyovX028"
# Astronomers studying the ISM often display extinction curves against inverse wavelength (wavenumber) to show the ultraviolet variation, as we do here. Infrared extinction varies much less and approaches zero at long wavelength in the absence of wavelength-independent, or grey, extinction.
# + [markdown] colab_type="text" id="TMEWEKJf-iSL"
# # Example 2: Deredden a Spectrum
# + [markdown] colab_type="text" id="zTeDJ-dI1cQj"
# Here we deredden (unextinguish) the IUE ultraviolet spectrum and optical photometry of the star $\rho$ Oph (HD 147933).
#
# First, we will use astroquery to fetch the archival [IUE spectrum from MAST](https://archive.stsci.edu/iue/):
# + colab={"base_uri": "https://localhost:8080/", "height": 193} colab_type="code" id="Zdwr_mdZcDeh" outputId="d274490e-e6e2-449f-aaed-29dd0908a308"
obsTable = Observations.query_object("HD 147933",radius="1 arcsec")
obsTable_spec=obsTable[obsTable['dataproduct_type']=='spectrum']
obsTable_spec.pprint()
obsids = obsTable_spec[39]['obsid']
dataProductsByID = Observations.get_product_list(obsids)
manifest = Observations.download_products(dataProductsByID)
# + [markdown] colab_type="text" id="MevN670kcI4B"
# We read the downloaded files into an astropy table:
# + colab={"base_uri": "https://localhost:8080/", "height": 124} colab_type="code" id="THilzBfVcDb-" outputId="31470912-d7f6-43fa-8ff5-714a40cc770e"
t_lwr = Table.read('./mastDownload/IUE/lwr05639/lwr05639mxlo_vo.fits')
print(t_lwr)
# + [markdown] colab_type="text" id="IiQ9u8QJcKtC"
# The `.quantity` extension in the next lines will read the Table columns into Quantity vectors. Quantities keep the units of the Table column attached to the numpy array values.
# + colab={} colab_type="code" id="P54jimyWcDZa"
wav_UV = t_lwr['WAVE'][0,].quantity
UVflux = t_lwr['FLUX'][0,].quantity
# + [markdown] colab_type="text" id="WCIgXzhdcUn7"
# Now, we use astroquery again to fetch photometry from Simbad to go with the IUE spectrum:
# + colab={} colab_type="code" id="lYPVhZrXcDWw"
custom_query = Simbad()
custom_query.add_votable_fields('fluxdata(U)','fluxdata(B)','fluxdata(V)')
phot_table=custom_query.query_object('HD 147933')
Umag=phot_table['FLUX_U']
Bmag=phot_table['FLUX_B']
Vmag=phot_table['FLUX_V']
# + [markdown] colab_type="text" id="OAxOt2AwcVfF"
# To convert the photometry to flux, we look up some [properties of the photometric passbands](http://ned.ipac.caltech.edu/help/photoband.lst), including the flux of a magnitude zero star through the each passband, also known as the zero-point of the passband.
# + colab={} colab_type="code" id="cKMXywyCcDUU"
wav_U = 0.3660 * u.micron
zeroflux_U_nu = 1.81E-23 * u.Watt/(u.m*u.m*u.Hz)
wav_B = 0.4400 * u.micron
zeroflux_B_nu = 4.26E-23 * u.Watt/(u.m*u.m*u.Hz)
wav_V = 0.5530 * u.micron
zeroflux_V_nu = 3.64E-23 * u.Watt/(u.m*u.m*u.Hz)
# + [markdown] colab_type="text" id="3cFcAlSOcjht"
# The zero-points that we found for the optical passbands are not in the same units as the IUE fluxes. To make matters worse, the zero-point fluxes are $F_\nu$ and the IUE fluxes are $F_\lambda$. To convert between them, the wavelength is needed. Fortunately, astropy provides an easy way to make the conversion with *equivalencies*:
# + colab={} colab_type="code" id="pOFMtrNAcDRa"
zeroflux_U = zeroflux_U_nu.to(u.erg/u.AA/u.cm/u.cm/u.s,
equivalencies=u.spectral_density(wav_U))
zeroflux_B = zeroflux_B_nu.to(u.erg/u.AA/u.cm/u.cm/u.s,
equivalencies=u.spectral_density(wav_B))
zeroflux_V = zeroflux_V_nu.to(u.erg/u.AA/u.cm/u.cm/u.s,
equivalencies=u.spectral_density(wav_V))
# + [markdown] colab_type="text" id="7XlrfQtjctNG"
# Now we can convert from photometry to flux using the definition of magnitude:
# $$
# F=F_0\ 10^{-0.4\, m}
# $$
# + colab={} colab_type="code" id="p_DaxRzjcDOZ"
Uflux = zeroflux_U * 10.**(-0.4*Umag)
Bflux = zeroflux_B * 10.**(-0.4*Bmag)
Vflux = zeroflux_V * 10.**(-0.4*Vmag)
# + [markdown] colab_type="text" id="ODhSujFjc0hM"
# Using astropy quantities allow us to take advantage of astropy's unit support in plotting. [Calling `astropy.visualization.quantity_support` explicitly turns the feature on.](http://docs.astropy.org/en/stable/units/quantity.html#plotting-quantities) Then, when quantity objects are passed to matplotlib plotting functions, the axis labels are automatically labeled with the unit of the quantity. In addition, quantities are converted automatically into the same units when combining multiple plots on the same axes.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 300} colab_type="code" id="x31EhibTcDFg" outputId="70578733-20e9-4b5b-b65e-c09f1894c0da"
astropy.visualization.quantity_support()
plt.plot(wav_UV,UVflux,'m',label='UV')
plt.plot(wav_V,Vflux,'ko',label='U, B, V')
plt.plot(wav_B,Bflux,'ko')
plt.plot(wav_U,Uflux,'ko')
plt.legend(loc='best')
plt.ylim(0,3E-10)
plt.title('rho Oph')
plt.show()
# + [markdown] colab_type="text" id="vCfk0D87dDE8"
# Finally, we initialize the extinction model, choosing values $R_V = 5$ and $E_{B-V} = 0.5$. This star is famous in the ISM community for having large-$R_V$ dust in the line of sight.
# + colab={} colab_type="code" id="wdp5ERuqcC8_"
Rv = 5.0 # Usually around 3, but about 5 for this star.
Ebv = 0.5
ext = F99(Rv=Rv)
# + [markdown] colab_type="text" id="cpN5n_MIdv26"
# To extinguish (redden) a spectrum, multiply by the `ext.extinguish` function. To unextinguish (deredden), divide by the same `ext.extinguish`, as we do here:
# + colab={"base_uri": "https://localhost:8080/", "height": 355} colab_type="code" id="DyxkYpdxdvfc" outputId="de570750-83c7-4496-f13e-3f1f97eff8ef"
plt.semilogy(wav_UV,UVflux,'m',label='UV')
plt.semilogy(wav_V,Vflux,'ko',label='U, B, V')
plt.semilogy(wav_B,Bflux,'ko')
plt.semilogy(wav_U,Uflux,'ko')
plt.semilogy(wav_UV,UVflux/ext.extinguish(wav_UV,Ebv=Ebv),'b',
label='dereddened: EBV=0.5, RV=5')
plt.semilogy(wav_V,Vflux/ext.extinguish(wav_V,Ebv=Ebv),'ro',
label='dereddened: EBV=0.5, RV=5')
plt.semilogy(wav_B,Bflux/ext.extinguish(wav_B,Ebv=Ebv),'ro')
plt.semilogy(wav_U,Uflux/ext.extinguish(wav_U,Ebv=Ebv),'ro')
plt.legend(loc='best')
plt.title('rho Oph')
plt.show()
# + [markdown] colab_type="text" id="L-yuz7YfFMxm"
# Notice that, by dereddening the spectrum, the absorption feature at 2175 Angstrom is removed. This feature can also be seen as the prominent bump in the extinction curves in Example 1. That we have smoothly removed the 2175 Angstrom feature suggests that the values we chose, $R_V = 5$ and $E_{B-V} = 0.5$, are a reasonable model for the foreground dust.
#
# Those experienced with dereddening should notice that that `dust_extinction` returns $A_\lambda/A_V$, while other routines like the IDL fm_unred procedure often return $A_\lambda/E_{B-V}$ by default and need to be divided by $R_V$ in order to compare directly with `dust_extinction`.
# + [markdown] colab_type="text" id="WqtsQTZbp9nz"
# # Example 3: Calculate Color Excess with `synphot`
# + [markdown] colab_type="text" id="WwsM9_56FVYr"
# Calculating broadband *photometric* extinction is harder than it might look at first. All we have to do is look up $A_\lambda$ for a particular passband, right? Under the right conditions, yes. In general, no.
#
# Remember that we have to integrate over a passband to get synthetic photometry,
# $$
# A = -2.5\log\left(\frac{\int W_\lambda F_{\lambda,0} 10^{-0.4A_\lambda} d\lambda}{\int W_\lambda F_{\lambda,0} d\lambda} \right),
# $$
#
# where $W_\lambda$ is the fraction of incident energy transmitted through a filter. See the detailed appendix in [Bessell & Murphy (2012)](https://ui.adsabs.harvard.edu/#abs/2012PASP..124..140B/abstract)
# for an excellent review of the issues and common misunderstandings in synthetic photometry.
#
# There is an important point to be made here. The expression above does not simplify any further. Strictly speaking, it is impossible to convert spectral extinction $A_\lambda$ into a magnitude system without knowing the wavelength dependence of the source's original flux across the filter in question. As a special case, if we assume that the source flux is constant in the band (i.e. $F_\lambda = F$), then we can cancel these factors out from the integrals, and extinction in magnitudes becomes the weighted average of the extinction factor across the filter in question. In that special case, $A_\lambda$ at $\lambda_{\rm eff}$ is a good approximation for magnitude extinction.
#
# In this example, we will demonstrate the more general calculation of photometric extinction. We use a blackbody curve for the flux before the dust, apply an extinction curve, and perform synthetic photometry to calculate extinction and reddening in a magnitude system.
#
# + [markdown] colab_type="text" id="avGGI5fVX2wV"
# First, let's get the filter transmission curves:
# + colab={"base_uri": "https://localhost:8080/", "height": 155} colab_type="code" id="wO-wwoOnp2B3" outputId="20b76dae-ea0d-4f29-ebfb-28e41f29895c"
# Optional, for when the STScI ftp server is not answering:
config.conf.vega_file='http://ssb.stsci.edu/cdbs/calspec/alpha_lyr_stis_008.fits'
config.conf.johnson_u_file='http://ssb.stsci.edu/cdbs/comp/nonhst/johnson_u_004_syn.fits'
config.conf.johnson_b_file='http://ssb.stsci.edu/cdbs/comp/nonhst/johnson_b_004_syn.fits'
config.conf.johnson_v_file='http://ssb.stsci.edu/cdbs/comp/nonhst/johnson_v_004_syn.fits'
config.conf.johnson_r_file='http://ssb.stsci.edu/cdbs/comp/nonhst/johnson_r_003_syn.fits'
config.conf.johnson_i_file='http://ssb.stsci.edu/cdbs/comp/nonhst/johnson_i_003_syn.fits'
config.conf.bessel_j_file='http://ssb.stsci.edu/cdbs/comp/nonhst/bessell_j_003_syn.fits'
config.conf.bessel_h_file='http://ssb.stsci.edu/cdbs/comp/nonhst/bessell_h_004_syn.fits'
config.conf.bessel_k_file='http://ssb.stsci.edu/cdbs/comp/nonhst/bessell_k_003_syn.fits'
u_band = SpectralElement.from_filter('johnson_u')
b_band = SpectralElement.from_filter('johnson_b')
v_band = SpectralElement.from_filter('johnson_v')
r_band = SpectralElement.from_filter('johnson_r')
i_band = SpectralElement.from_filter('johnson_i')
j_band = SpectralElement.from_filter('bessel_j')
h_band = SpectralElement.from_filter('bessel_h')
k_band = SpectralElement.from_filter('bessel_k')
# + [markdown] colab_type="text" id="hekvrcEGpvYd"
# If you are running this with your own python, see the [synphot documentation](https://synphot.readthedocs.io/en/latest/#installation-and-setup) on how to install your own copy of the necessary files.
# + [markdown] colab_type="text" id="vQTSrYpvJeaY"
# Next, let's make a background flux to which we will apply extinction. Here we make a 10,000 K blackbody using the model mechanism from within `synphot` and normalize it to $V$ = 10 in the Vega-based magnitude system.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="NuPS0Ij2ncC0" outputId="056bfa64-e194-4217-869e-3690206032d5"
# First, create a blackbody at some temperature.
sp = SourceSpectrum(BlackBodyNorm1D, temperature=10000)
# sp.plot(left=1, right=15000, flux_unit='flam', title='Blackbody')
# Get the Vega spectrum as the zero point flux.
vega = SourceSpectrum.from_vega()
# vega.plot(left=1, right=15000)
# Normalize the blackbody to some chosen magnitude, say V = 10.
vmag = 10.
v_band = SpectralElement.from_filter('johnson_v')
sp_norm = sp.normalize(vmag * units.VEGAMAG, v_band, vegaspec=vega)
sp_norm.plot(left=1, right=15000, flux_unit='flam', title='Normed Blackbody')
# + [markdown] colab_type="text" id="H4jYc_F8CiHi"
# Now we initialize the extinction model and choose an extinction of $A_V$ = 2. To get the `dust_extinction` model working with `synphot`, we create a wavelength array and make a spectral element with the extinction model as a lookup table.
# + colab={"base_uri": "https://localhost:8080/", "height": 295} colab_type="code" id="9H4VHd0VCCBG" outputId="7713cbd6-e68f-4a9a-85cc-9723b99ef6f2"
# Initialize the extinction model and choose the extinction, here Av = 2.
ext = CCM89(Rv=3.1)
Av = 2.
# Create a wavelength array.
wav = np.arange(0.1, 3, 0.001)*u.micron
# Make the extinction model in synphot using a lookup table.
ex = ExtinctionCurve(ExtinctionModel1D,
points=wav, lookup_table=ext.extinguish(wav, Av=Av))
sp_ext = sp_norm*ex
sp_ext.plot(left=1, right=15000, flux_unit='flam',
title='Normed Blackbody with Extinction')
# + [markdown] colab_type="text" id="h0dxfniFF-Bf"
# Synthetic photometry refers to modeling an observation of a star by multiplying the theoretical model for the astronomical flux through a certain filter response function, then integrating.
# + colab={} colab_type="code" id="_1m5IFT4E2v_"
# "Observe" the star through the filter and integrate to get photometric mag.
sp_obs = Observation(sp_ext, v_band)
sp_obs_before = Observation(sp_norm, v_band)
# sp_obs.plot(left=1, right=15000, flux_unit='flam',
# title='Normed Blackbody with Extinction through V Filter')
# + [markdown] colab_type="text" id="vFtJs0PhGLzx"
# Next, `synphot` performs the integration and computes magnitudes in the Vega system.
# + colab={"base_uri": "https://localhost:8080/", "height": 69} colab_type="code" id="ycvVHlOGCRur" outputId="c939feab-31fb-4f68-f6d9-c749c020d04e"
sp_stim_before = sp_obs_before.effstim(flux_unit='vegamag', vegaspec=vega)
sp_stim = sp_obs.effstim(flux_unit='vegamag', vegaspec=vega)
print('before dust, V =', np.round(sp_stim_before,1))
print('after dust, V =', np.round(sp_stim,1))
# Calculate extinction and compare to our chosen value.
Av_calc = sp_stim - sp_stim_before
print('$A_V$ = ', np.round(Av_calc,1))
# + [markdown] colab_type="text" id="asE7JUPuHddK"
# This is a good check for us to do. We normalized our spectrum to $V$ = 10 mag and added 2 mag of visual extinction, so the synthetic photometry procedure should reproduce these chosen values, and it does. Now we are ready to find the extinction in other passbands.
# + [markdown] colab_type="text" id="woWiGJKwGI2v"
# We calculate the new photometry for the rest of the Johnson optical and the Bessell infrared filters. We calculate extinction $A = \Delta m$ and plot color excess, $E(\lambda - V) = A_\lambda - A_V$.
#
# Notice that `synphot` calculates the effective wavelength of the observations for us, which is very useful for plotting the results. We show reddening with the model extinction curve for comparison in the plot.
# + colab={"base_uri": "https://localhost:8080/", "height": 436} colab_type="code" id="0PMDsoC68w4b" outputId="c52d74e5-c64e-46ee-e1a3-0282e6c9349a"
bands = [u_band,b_band,v_band,r_band,i_band,j_band,h_band,k_band]
for band in bands:
# Calculate photometry with dust:
sp_obs = Observation(sp_ext, band, force='extrap')
obs_effstim = sp_obs.effstim(flux_unit='vegamag', vegaspec=vega)
# Calculate photometry without dust:
sp_obs_i = Observation(sp_norm, band, force='extrap')
obs_i_effstim = sp_obs_i.effstim(flux_unit='vegamag', vegaspec=vega)
# Extinction = mag with dust - mag without dust
# Color excess = extinction at lambda - extinction at V
color_excess = obs_effstim - obs_i_effstim - Av_calc
plt.plot(sp_obs_i.effective_wavelength(), color_excess,'or')
print(np.round(sp_obs_i.effective_wavelength(),1), ',',
np.round(color_excess,2))
# Plot the model extinction curve for comparison
plt.plot(wav,Av*ext(wav)-Av,'--k')
plt.ylim([-2,2])
plt.xlabel('$\lambda$ (Angstrom)')
plt.ylabel('E($\lambda$-V)')
plt.title('Reddening of T=10,000K Background Source with Av=2')
plt.show()
# + [markdown] colab_type="text" id="9B4gdXzn8LtA"
# ## Exercise
# Try changing the blackbody temperature to something very hot or very cool. Are the color excess values the same? Have the effective wavelengths changed?
#
# Note that the photometric extinction changes because the filter transmission is not uniform. The observed throughput of the filter depends on the shape of the background source flux.
# -
| tutorials/notebooks/color-excess/color-excess.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # Generating C Code to implement Method of Lines Timestepping for Explicit Runge Kutta Methods
#
# ## Authors: <NAME> & <NAME>
#
# ## This tutorial notebook generates three blocks of C Code in order to perform Method of Lines timestepping.
#
# **Notebook Status:** <font color='green'><b> Validated </b></font>
#
# **Validation Notes:** This tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented [below](#code_validation). All Runge-Kutta Butcher tables were validated using truncated Taylor series in [a separate module](Tutorial-RK_Butcher_Table_Validation.ipynb). Finally, C-code implementation of RK4 was validated against a trusted version. C-code implementations of other RK methods seem to work as expected in the context of solving the scalar wave equation in Cartesian coordinates.
#
# ### NRPy+ Source Code for this module:
# * [MoLtimestepping/C_Code_Generation.py](../edit/MoLtimestepping/C_Code_Generation.py)
# * [MoLtimestepping/RK_Butcher_Table_Dictionary.py](../edit/MoLtimestepping/RK_Butcher_Table_Dictionary.py) ([**Tutorial**](Tutorial-RK_Butcher_Table_Dictionary.ipynb)) Stores the Butcher tables for the explicit Runge Kutta methods
#
# ## Introduction:
#
# When numerically solving a partial differential equation initial-value problem, subject to suitable boundary conditions, we implement Method of Lines to "integrate" the solution forward in time.
#
#
# ### The Method of Lines:
#
# Once we have the initial data for a PDE, we "evolve it forward in time", using the [Method of Lines](https://reference.wolfram.com/language/tutorial/NDSolveMethodOfLines.html). In short, the Method of Lines enables us to handle
# 1. the **spatial derivatives** of an initial value problem PDE using **standard finite difference approaches**, and
# 2. the **temporal derivatives** of an initial value problem PDE using **standard strategies for solving ordinary differential equations (ODEs), like Runge Kutta methods** so long as the initial value problem PDE can be written in the first-order-in-time form
# $$\partial_t \vec{f} = \mathbf{M}\ \vec{f},$$
# where $\mathbf{M}$ is an $N\times N$ matrix containing only *spatial* differential operators that act on the $N$-element column vector $\vec{f}$. $\mathbf{M}$ may not contain $t$ or time derivatives explicitly; only *spatial* partial derivatives are allowed to appear inside $\mathbf{M}$.
#
# You may find the next module [Tutorial-ScalarWave](Tutorial-ScalarWave.ipynb) extremely helpful as an example for implementing the Method of Lines for solving the Scalar Wave equation in Cartesian coordinates.
#
# ### Generating the C code:
# This module describes how three C code blocks are written to implement Method of Lines timestepping for a specified RK method. The first block is dedicated to allocating memory for the appropriate number of grid function lists needed for the given RK method. The second block will implement the Runge Kutta numerical scheme based on the corresponding Butcher table. The third block will free up the previously allocated memory after the Method of Lines run is complete. These blocks of code are stored within the following three header files respectively
#
# 1. `MoLtimestepping/RK_Allocate_Memory.h`
# 1. `MoLtimestepping/RK_MoL.h`
# 1. `MoLtimestepping/RK_Free_Memory.h`
#
# The generated code is then included in future Start-to-Finish example tutorial notebooks when solving PDEs numerically.
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# This notebook is organized as follows
#
# 1. [Step 1](#initializenrpy): Initialize needed Python/NRPy+ modules
# 1. [Step 2](#diagonal): Checking if Butcher Table is Diagonal
# 1. [Step 3](#ccode): Generating the C Code
# 1. [Step 3.a](#allocate): Allocating Memory, `MoLtimestepping/RK_Allocate_Memory.h`
# 1. [Step 3.b](#rkmol): Implementing the Runge Kutta Scheme for Method of Lines Timestepping, `MoLtimestepping/RK_MoL.h`
# 1. [Step 3.c](#free): Freeing Allocated Memory, `MoLtimestepping/RK_Free_Memory.h`
# 1. [Step 4](#code_validation): Code Validation against `MoLtimestepping.RK_Butcher_Table_Generating_C_Code` NRPy+ module
# 1. [Step 5](#latex_pdf_output): Output this notebook to $\LaTeX$-formatted PDF file
# <a id='initializenrpy'></a>
#
# # Step 1: Initialize needed Python/NRPy+ modules [Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
# Let's start by importing all the needed modules from Python/NRPy+:
import sympy as sp # Import SymPy, a computer algebra system written entirely in Python
import os, shutil # Standard Python modules for multiplatform OS-level functions
from MoLtimestepping.RK_Butcher_Table_Dictionary import Butcher_dict
# <a id='diagonal'></a>
#
# # Step 2: Checking if a Butcher table is Diagonal [Back to [top](#toc)\]
# $$\label{diagonal}$$
#
# A diagonal Butcher table takes the form
#
# $$\begin{array}{c|cccccc}
# 0 & \\
# a_1 & a_1 & \\
# a_2 & 0 & a_2 & \\
# a_3 & 0 & 0 & a_3 & \\
# \vdots & \vdots & \ddots & \ddots & \ddots \\
# a_s & 0 & 0 & 0 & \cdots & a_s \\ \hline
# & b_1 & b_2 & b_3 & \cdots & b_{s-1} & b_s
# \end{array}$$
#
# where $s$ is the number of required predictor-corrector steps for a given RK method (see [<NAME>. (2008)](https://onlinelibrary.wiley.com/doi/book/10.1002/9780470753767)). One known diagonal RK method is the classic RK4 represented in Butcher table form as:
#
# $$\begin{array}{c|cccc}
# 0 & \\
# 1/2 & 1/2 & \\
# 1/2 & 0 & 1/2 & \\
# 1 & 0 & 0 & 1 & \\ \hline
# & 1/6 & 1/3 & 1/3 & 1/6
# \end{array} $$
#
# Diagonal Butcher tables are nice when it comes to saving required memory space. Each new step for a diagonal RK method, when computing the new $k_i$, does not depend on the previous calculation, and so there are ways to save memory. Significantly so in large three-dimensional spatial grid spaces.
# +
def diagonal(key):
diagonal = True # Start with the Butcher table is diagonal
Butcher = Butcher_dict[key][0]
L = len(Butcher)-1 # Establish the number of rows to check for diagonal trait, all bust last row
row_idx = 0 # Initialize the Butcher table row index
for i in range(L): # Check all the desired rows
for j in range(1,row_idx): # Check each element before the diagonal element in a row
if Butcher[i][j] != sp.sympify(0): # If any element is non-zero, then the table is not diagonal
diagonal = False
break
row_idx += 1 # Update to check the next row
return diagonal
# State whether each Butcher table is diagonal or not
for key, value in Butcher_dict.items():
if diagonal(key) == True:
print("The RK method "+str(key)+" is diagonal!")
else:
print("The RK method "+str(key)+" is NOT diagonal!")
# -
# <a id='ccode'></a>
#
# # Step 3: Generating the C Code [Back to [top](#toc)\]
# $$\label{ccode}$$
#
# The following sections build up the C code for implementing the Method of Lines timestepping algorithm for solving PDEs. To see what the C code looks like for a particular method, simply change the `RK_method` below, otherwise it will default to `"RK4"`.
# Choose a method to see the C code print out for
RK_method = "RK3 Ralston"
# <a id='free'></a>
#
# ## Step 3.a: Freeing Allocated Memory, `MoLtimestepping/RK_Free_Memory.h` [Back to [top](#toc)\]
# $$\label{free}$$
#
# We define the function `RK_free()` which generates the C code for freeing the memory that was being occupied by the grid functions lists that had been allocated. The function writes the C code to the header file `MoLtimestepping/RK_Free_Memory.h`
# Step 3.a: When allocating memory, we populate a list malloced_gridfunctions,
# which is used here to determine which gridfunctions need memory freed,
# via the free() command. Free the mallocs!
def free_allocated_memory(outdir,RK_method,malloced_gridfunctions):
# This step is made extremely easy, as we had to
with open(os.path.join(outdir, "RK_Free_Memory.h"), "w") as file:
file.write("// Code snippet freeing gridfunction memory for \"" + RK_method + "\" method:\n")
for gridfunction in malloced_gridfunctions:
file.write("free(" + gridfunction + ");\n")
# <a id='rkmol'></a>
#
# ## Step 3.b: Implementing the Runge Kutta Scheme for Method of Lines Timestepping, `MoLtimestepping/RK_MoL.h` [Back to [top](#toc)\]
# $$\label{rkmol}$$
#
# We define the function `RK_MoL()` which generates the C code for implementing Method of Lines using a specified Runge Kutta scheme. The function writes the C code to the header file `MoLtimestepping/RK_MoL.h`.
# +
# Step 3.b: Main driver function for outputting all the MoL C Code
def MoL_C_Code_Generation(RK_method = "RK4", RHS_string = "", post_RHS_string = "",outdir="MoLtimestepping/",
MemAllocOnly=False):
####### Step 3.b.i: Allocating Memory
malloc_str = "// Code snippet allocating gridfunction memory for \"" + RK_method + "\" method:\n"
# Loop over grids
malloced_gridfunctions = []
# Set gridfunction type
type_str = "REAL *restrict "
# Define a couple useful functions for outputting the needed C code for allocating memory
def malloc_gfs_str(varname):
malloced_gridfunctions.append(varname)
memory_alloc_str = " = (REAL *)malloc(sizeof(REAL) * NUM_EVOL_GFS * Nxx_plus_2NGHOSTS_tot"+")"
return type_str + varname + memory_alloc_str + ";\n"
def diagnostic_output_gfs_equal_to(gfs):
return type_str + "diagnostic_output_gfs"+" = "+gfs + ";\n"
# No matter the method we define gridfunctions "y_n_gfs" to store the initial data
malloc_str += malloc_gfs_str("y_n_gfs")
if diagonal(RK_method) == True and "RK3" in RK_method:
malloc_str += malloc_gfs_str("k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs")
malloc_str += malloc_gfs_str("k2_or_y_nplus_a32_k2_gfs")
malloc_str += diagnostic_output_gfs_equal_to("k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs")
else:
if diagonal(RK_method) == False: # Allocate memory for non-diagonal Butcher tables
# Determine the number of k_i steps based on length of Butcher Table
num_k = len(Butcher_dict[RK_method][0])-1
# For non-diagonal tables an intermediate gridfunction "next_y_input" is used for rhs evaluations
malloc_str += malloc_gfs_str("next_y_input_gfs")
for i in range(num_k): # Need to allocate all k_i steps for a given method
malloc_str += malloc_gfs_str("k"+str(i+1)+"_gfs")
malloc_str += diagnostic_output_gfs_equal_to("k1_gfs")
else: # Allocate memory for diagonal Butcher tables, which use a "y_nplus1_running_total gridfunction"
malloc_str += malloc_gfs_str("y_nplus1_running_total_gfs")
if RK_method != 'Euler': # Allocate memory for diagonal Butcher tables that aren't Euler
# Need k_odd for k_1,3,5... and k_even for k_2,4,6...
malloc_str += malloc_gfs_str("k_odd_gfs")
malloc_str += malloc_gfs_str("k_even_gfs")
malloc_str += diagnostic_output_gfs_equal_to("y_nplus1_running_total_gfs")
with open(os.path.join(outdir,"RK_Allocate_Memory.h"), "w") as file:
file.write(malloc_str)
if MemAllocOnly:
free_allocated_memory(outdir,RK_method,malloced_gridfunctions)
return
########################################################################################################################
# EXAMPLE
# ODE: y' = f(t,y), y(t_0) = y_0
# Starting at time t_n with solution having value y_n and trying to update to y_nplus1 with timestep dt
# Example of scheme for RK4 with k_1, k_2, k_3, k_4 (Using non-diagonal algorithm) Notice this requires storage of
# y_n, y_nplus1, k_1 through k_4
# k_1 = dt*f(t_n, y_n)
# k_2 = dt*f(t_n + 1/2*dt, y_n + 1/2*k_1)
# k_3 = dt*f(t_n + 1/2*dt, y_n + 1/2*k_2)
# k_4 = dt*f(t_n + dt, y_n + k_3)
# y_nplus1 = y_n + 1/3k_1 + 1/6k_2 + 1/6k_3 + 1/3k_4
# Example of scheme RK4 using only k_odd and k_even (Diagonal algorithm) Notice that this only requires storage
# k_odd = dt*f(t_n, y_n)
# y_nplus1 = 1/3*k_odd
# k_even = dt*f(t_n + 1/2*dt, y_n + 1/2*k_odd)
# y_nplus1 += 1/6*k_even
# k_odd = dt*f(t_n + 1/2*dt, y_n + 1/2*k_even)
# y_nplus1 += 1/6*k_odd
# k_even = dt*f(t_n + dt, y_n + k_odd)
# y_nplus1 += 1/3*k_even
########################################################################################################################
####### Step 3.b.ii: Implementing the Runge Kutta Scheme for Method of Lines Timestepping
Butcher = Butcher_dict[RK_method][0] # Get the desired Butcher table from the dictionary
num_steps = len(Butcher)-1 # Specify the number of required steps to update solution
# Diagonal RK3 only!!!
def single_RK_substep(commentblock, RHS_str, RHS_input_str, RHS_output_str, RK_lhss_list, RK_rhss_list,
post_RHS_list, post_RHS_output_list, indent = " "):
return_str = commentblock + "\n"
if not isinstance(RK_lhss_list,list):
RK_lhss_list = [RK_lhss_list]
if not isinstance(RK_rhss_list,list):
RK_rhss_list = [RK_rhss_list]
if not isinstance(post_RHS_list,list):
post_RHS_list = [post_RHS_list]
if not isinstance(post_RHS_output_list,list):
post_RHS_output_list = [post_RHS_output_list]
# Part 1: RHS evaluation:
return_str += RHS_str.replace("RK_INPUT_GFS", RHS_input_str).\
replace("RK_OUTPUT_GFS",RHS_output_str)+"\n"
# Part 2: RK update
return_str += "LOOP_ALL_GFS_GPS"+"(i) {\n"
for lhs,rhs in zip(RK_lhss_list,RK_rhss_list):
return_str += indent + lhs + "[i] = " + rhs.replace("_gfs","_gfs") + ";\n"
return_str += "}\n"
# Part 3: Call post-RHS functions
for post_RHS,post_RHS_output in zip(post_RHS_list,post_RHS_output_list):
return_str += post_RHS.replace("RK_OUTPUT_GFS",post_RHS_output)+"\n"
return return_str+"\n"
RK_str = "// C code implementation of " + RK_method + " Method of Lines timestepping.\n"
if diagonal(RK_method) == True and "RK3" in RK_method:
# In a diagonal RK3 method, only 3 gridfunctions need be defined. Below implements this approach.
# k_1
RK_str += """
// In a diagonal RK3 method like this one, only 3 gridfunctions need be defined. Below implements this approach.
// Using y_n_gfs as input, k1 and apply boundary conditions\n"""
RK_str += single_RK_substep(
commentblock = """
// ***k1 substep:***
// 1. We will store k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs now as
// ... the update for the next rhs evaluation y_n + a21*k1*dt
// Post-RHS evaluation:
// 1. Apply post-RHS to y_n + a21*k1*dt""",
RHS_str = RHS_string,
RHS_input_str = "y_n_gfs", RHS_output_str = "k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs",
RK_lhss_list = ["k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs"],
RK_rhss_list = ["("+sp.ccode(Butcher[1][1]).replace("L","")+")*k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs[i]*dt + y_n_gfs[i]"],
post_RHS_list = [post_RHS_string], post_RHS_output_list = ["k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs"])
# k_2
RK_str += single_RK_substep(
commentblock="""
// ***k2 substep:***
// 1. Reassign k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs to be the running total y_{n+1}; a32*k2*dt to the running total
// 2. Store k2_or_y_nplus_a32_k2_gfs now as y_n + a32*k2*dt
// Post-RHS evaluation:
// 1. Apply post-RHS to both y_n + a32*k2 (stored in k2_or_y_nplus_a32_k2_gfs)
// ... and the y_{n+1} running total, as they have not been applied yet to k2-related gridfunctions""",
RHS_str=RHS_string,
RHS_input_str="k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs", RHS_output_str="k2_or_y_nplus_a32_k2_gfs",
RK_lhss_list=["k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs","k2_or_y_nplus_a32_k2_gfs"],
RK_rhss_list=["("+sp.ccode(Butcher[3][1]).replace("L","")+")*(k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs[i] - y_n_gfs[i])/("+sp.ccode(Butcher[1][1]).replace("L","")+") + y_n_gfs[i] + ("+sp.ccode(Butcher[3][2]).replace("L","")+")*k2_or_y_nplus_a32_k2_gfs[i]*dt",
"("+sp.ccode(Butcher[2][2]).replace("L","")+")*k2_or_y_nplus_a32_k2_gfs[i]*dt + y_n_gfs[i]"],
post_RHS_list=[post_RHS_string,post_RHS_string],
post_RHS_output_list=["k2_or_y_nplus_a32_k2_gfs","k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs"])
# k_3
RK_str += single_RK_substep(
commentblock="""
// ***k3 substep:***
// 1. Add k3 to the running total and save to y_n
// Post-RHS evaluation:
// 1. Apply post-RHS to y_n""",
RHS_str=RHS_string,
RHS_input_str="k2_or_y_nplus_a32_k2_gfs", RHS_output_str="y_n_gfs",
RK_lhss_list=["y_n_gfs","k2_or_y_nplus_a32_k2_gfs"],
RK_rhss_list=["k1_or_y_nplus_a21_k1_or_y_nplus1_running_total_gfs[i] + ("+sp.ccode(Butcher[3][3]).replace("L","")+")*y_n_gfs[i]*dt"],
post_RHS_list=[post_RHS_string],
post_RHS_output_list=["y_n_gfs"])
else:
y_n = "y_n_gfs"
if diagonal(RK_method) == False:
for s in range(num_steps):
next_y_input = "next_y_input_gfs"
# If we're on the first step (s=0), we use y_n gridfunction as input.
# Otherwise next_y_input is input. Output is just the reverse.
if s==0: # If on first step:
RHS_input = y_n
else: # If on second step or later:
RHS_input = next_y_input
RHS_output = "k" + str(s + 1) + "_gfs"
if s == num_steps-1: # If on final step:
RK_lhs = y_n
RK_rhs = y_n + "[i] + dt*("
else: # If on anything but the final step:
RK_lhs = next_y_input
RK_rhs = y_n + "[i] + dt*("
for m in range(s+1):
if Butcher[s+1][m+1] != 0:
if Butcher[s+1][m+1] != 1:
RK_rhs += " + k"+str(m+1)+"_gfs[i]*("+sp.ccode(Butcher[s+1][m+1]).replace("L","")+")"
else:
RK_rhs += " + k"+str(m+1)+"_gfs[i]"
RK_rhs += " )"
post_RHS = post_RHS_string
if s == num_steps-1: # If on final step:
post_RHS_output = y_n
else: # If on anything but the final step:
post_RHS_output = next_y_input
RK_str += single_RK_substep(
commentblock="// ***k" + str(s + 1) + " substep:***",
RHS_str=RHS_string,
RHS_input_str=RHS_input, RHS_output_str=RHS_output,
RK_lhss_list=[RK_lhs], RK_rhss_list=[RK_rhs],
post_RHS_list=[post_RHS],
post_RHS_output_list=[post_RHS_output])
else:
y_nplus1_running_total = "y_nplus1_running_total_gfs"
if RK_method == 'Euler': # Euler's method doesn't require any k_i, and gets its own unique algorithm
RK_str += single_RK_substep(
commentblock="// ***Euler timestepping only requires one RHS evaluation***",
RHS_str=RHS_string,
RHS_input_str=y_n, RHS_output_str=y_nplus1_running_total,
RK_lhss_list=[y_n], RK_rhss_list=[y_n+"[i] + "+y_nplus1_running_total+"[i]*dt"],
post_RHS_list=[post_RHS_string],
post_RHS_output_list=[y_n])
else:
for s in range(num_steps):
# If we're on the first step (s=0), we use y_n gridfunction as input.
# and k_odd as output.
if s == 0:
RHS_input = "y_n_gfs"
RHS_output = "k_odd_gfs"
# For the remaining steps the inputs and outputs alternate between k_odd and k_even
elif s%2 == 0:
RHS_input = "k_even_gfs"
RHS_output = "k_odd_gfs"
else:
RHS_input = "k_odd_gfs"
RHS_output = "k_even_gfs"
RK_lhs_list = []
RK_rhs_list = []
if s != num_steps-1: # For anything besides the final step
if s == 0: # The first RK step
RK_lhs_list.append(y_nplus1_running_total)
RK_rhs_list.append(RHS_output+"[i]*dt*("+sp.ccode(Butcher[num_steps][s+1]).replace("L","")+")")
RK_lhs_list.append(RHS_output)
RK_rhs_list.append(y_n+"[i] + "+RHS_output+"[i]*dt*("+sp.ccode(Butcher[s+1][s+1]).replace("L","")+")")
else:
if Butcher[num_steps][s+1] !=0:
RK_lhs_list.append(y_nplus1_running_total)
if Butcher[num_steps][s+1] !=1:
RK_rhs_list.append(y_nplus1_running_total+"[i] + "+RHS_output+"[i]*dt*("+sp.ccode(Butcher[num_steps][s+1]).replace("L","")+")")
else:
RK_rhs_list.append(y_nplus1_running_total+"[i] + "+RHS_output+"[i]*dt")
if Butcher[s+1][s+1] !=0:
RK_lhs_list.append(RHS_output)
if Butcher[s+1][s+1] !=1:
RK_rhs_list.append(y_n+"[i] + "+RHS_output+"[i]*dt*("+sp.ccode(Butcher[s+1][s+1]).replace("L","")+")")
else:
RK_rhs_list.append(y_n+"[i] + "+RHS_output+"[i]*dt")
post_RHS_output = RHS_output
if s == num_steps-1: # If on the final step
if Butcher[num_steps][s+1] != 0:
RK_lhs_list.append(y_n)
if Butcher[num_steps][s+1] != 1:
RK_rhs_list.append(y_n+"[i] + "+y_nplus1_running_total+"[i] + "+RHS_output+"[i]*dt*("+sp.ccode(Butcher[num_steps][s+1]).replace("L","")+")")
else:
RK_rhs_list.append(y_n+"[i] + "+y_nplus1_running_total+"[i] + "+RHS_output+"[i]*dt)")
post_RHS_output = y_n
RK_str += single_RK_substep(
commentblock="// ***k" + str(s + 1) + " substep:***",
RHS_str=RHS_string,
RHS_input_str=RHS_input, RHS_output_str=RHS_output,
RK_lhss_list=RK_lhs_list, RK_rhss_list=RK_rhs_list,
post_RHS_list=[post_RHS_string],
post_RHS_output_list=[post_RHS_output])
with open(os.path.join(outdir,"RK_MoL.h"), "w") as file:
file.write(RK_str)
####### Step 3.b.iii: Freeing Allocated Memory
free_allocated_memory(outdir,RK_method,malloced_gridfunctions)
MoL_C_Code_Generation(RK_method,"rhs_eval(Nxx,Nxx_plus_2NGHOSTS,dxx, RK_INPUT_GFS, RK_OUTPUT_GFS);",
"apply_bcs(Nxx,Nxx_plus_2NGHOSTS, RK_OUTPUT_GFS);")
print("This is the MoL timestepping RK scheme C code for the "+str(RK_method)+" method: \n")
with open(os.path.join("MoLtimestepping/","RK_MoL.h"), "r") as file:
print(file.read())
# -
# <a id='code_validation'></a>
#
# # Step 4: Code Validation against `MoLtimestepping.RK_Butcher_Table_Generating_C_Code` NRPy+ module [Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# As a code validation check, we verify agreement in the dictionary of Butcher tables between
#
# 1. this tutorial and
# 2. the NRPy+ [MoLtimestepping.RK_Butcher_Table_Generating_C_Code](../edit/MoLtimestepping/RK_Butcher_Table_Generating_C_Code.py) module.
#
# We generate the header files for each RK method and check for agreement with the NRPY+ module.
# +
import sys
import MoLtimestepping.C_Code_Generation as MoLC
print("\n\n ### BEGIN VALIDATION TESTS ###")
import filecmp
for key, value in Butcher_dict.items():
MoL_C_Code_Generation(key,"rhs_eval(Nxx,Nxx_plus_2NGHOSTS,dxx, RK_INPUT_GFS, RK_OUTPUT_GFS);",
"apply_bcs(Nxx,Nxx_plus_2NGHOSTS, RK_OUTPUT_GFS);")
for filename in ["RK_Allocate_Memory.h","RK_MoL.h","RK_Free_Memory.h"]:
shutil.copy(os.path.join("MoLtimestepping/",filename), os.path.join("MoLtimestepping/",filename+key+".h"))
MoLC.MoL_C_Code_Generation(key, "rhs_eval(Nxx,Nxx_plus_2NGHOSTS,dxx, RK_INPUT_GFS, RK_OUTPUT_GFS);",
"apply_bcs(Nxx,Nxx_plus_2NGHOSTS, RK_OUTPUT_GFS);")
for filename in ["RK_Allocate_Memory.h","RK_MoL.h","RK_Free_Memory.h"]:
if filecmp.cmp(os.path.join("MoLtimestepping/",filename), os.path.join("MoLtimestepping/",filename+key+".h")) == False:
print("VALIDATION TEST FAILED ON files: "+os.path.join("MoLtimestepping/",filename)+" and "+
os.path.join("MoLtimestepping/",filename+key+".h"))
sys.exit(1)
print("VALIDATION TEST PASSED on all files from "+str(key)+" method")
print("### END VALIDATION TESTS ###")
# -
# <a id='latex_pdf_output'></a>
#
# # Step 5: Output this notebook to $\LaTeX$-formatted PDF \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-RK_Butcher_Table_Generating_C_Code.pdf](Tutorial-RK_Butcher_Table_Generating_C_Code.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import cmdline_helper as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("Tutorial-Method_of_Lines-C_Code_Generation")
| Tutorial-Method_of_Lines-C_Code_Generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="7eb26a4698dc8954acb121b489b07b3130c40cd8"
# **This is an interesting dataset for building Deep Learning Neural Networks. here we use tensorflow keras API to form the model.**
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-output=false _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# Import the necessary libraries
# TensorFlow and tf.keras
import tensorflow as tf
from tensorflow import keras
# Helper libraries
import matplotlib.pyplot as plt
import numpy as np
from os import listdir
from os.path import join
import cv2
import pandas
import os
import random
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# Set the path of the input folder
# data = "../input/flowers/flowers/"
data = "flowers/"
# List out the directories inside the main input folder
folders = os.listdir(data)
print(folders)
# + _uuid="18d55fd068cd1ca21ed14a4e566b215a75e9d45e"
# Import the images and resize them to a 128*128 size
# Also generate the corresponding labels
image_names = []
train_labels = []
train_images = []
size = 64,64
for folder in folders:
for file in os.listdir(os.path.join(data,folder)):
if file.endswith("jpg"):
image_names.append(os.path.join(data,folder,file))
train_labels.append(folder)
img = cv2.imread(os.path.join(data,folder,file))
im = cv2.resize(img,size)
train_images.append(im)
else:
continue
# + _uuid="8ea726765c318e3e855d37b262a7b9ac66bf3351"
# Transform the image array to a numpy type
train = np.array(train_images)
train.shape
# + _uuid="7946f5528f1217ef93cd94df26320ed1b5722935"
# Reduce the RGB values between 0 and 1
train = train.astype('float32') / 255.0
# + _uuid="5b0e814039e66a966e8cebe4662b502a788de87b"
# Extract the labels
label_dummies = pandas.get_dummies(train_labels)
labels = label_dummies.values.argmax(1)
# + _uuid="111462613dba646ccad64e2221175ff86b84bc0b"
pandas.unique(train_labels)
# + _uuid="3b2373c20c84544a16dd071395289e85f0222724"
pandas.unique(labels)
# + _uuid="09860dbb816fa04eef09fc2d4fa260403c0af079"
# Shuffle the labels and images randomly for better results
union_list = list(zip(train, labels))
random.shuffle(union_list)
train,labels = zip(*union_list)
# Convert the shuffled list to numpy array type
train = np.array(train)
labels = np.array(labels)
# + _uuid="31958ba17cb150683936e02e3556833bd928bd71"
# Develop a sequential model using tensorflow keras
model = keras.Sequential([
keras.layers.Flatten(input_shape=(64,64,3)),
keras.layers.Dense(128, activation=tf.nn.tanh),
keras.layers.Dense(5, activation=tf.nn.softmax)
])
# + _uuid="644b9f7c5687e36e4f34a8cd49db344acf7c39e4"
# Compute the model parameters
model.compile(optimizer=tf.train.AdamOptimizer(),
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
# + _uuid="f438f031c38fa58cd6c35c679560e3babacccc86"
# Train the model with 5 epochs
model.fit(train,labels, epochs=50)
# -
| Flowers Recognition/code/obsolete/oldest/flower-classification-model-tensorflow.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
A = np.array([[1,2], [3,4]])
A.shape
B = np.array([[5,6], [7,8]])
B.shape
np.dot(A,B)
# ### 3.3.3 ニューラルネットワークの行列の積
X = np.array([1, 2])
X.shape
W = np.array([[1,3,5], [2,4,6]])
print(W)
W.shape
Y = np.dot(X, W)
print(Y)
# ### 各層における信号伝達の実装
def sigmoid(x):
return 1 / (1 + np.exp(-x))
# +
X = np.array([1.0, 0.5])
W1 = np.array([[0.1, 0.3, 0.5], [0.2, 0.4, 0.6]])
B1 = np.array([0.1, 0.2, 0.3])
print(W1.shape)
print(X.shape)
print(B1.shape)
# +
A1 = np.dot(X, W1) + B1
Z1 = sigmoid(A1)
print(A1)
print(Z1)
# +
W2 = np.array([[0.1, 0.4], [0.2, 0.5], [0.3, 0.6]])
B2 = np.array([0.1, 0.2])
print(Z1.shape)
print(W2.shape)
print(B2.shape)
# +
A2 = np.dot(Z1, W2) + B2
Z2 = sigmoid(A2)
print(A2)
print(Z2)
# +
def identity_function(x):
return x
W3 = np.array([[0.1, 0.3], [0.2, 0.4]])
B3 = np.array([0.1, 0.2])
A3 = np.dot(Z2, W3)+ B3
Y = identity_function(A3)
Y
# +
def init_network():
network = {}
network['W1'] = np.array([[0.1, 0.3, 0.5], [0.2, 0.4, 0.6]])
network['b1'] = np.array([0.1, 0.2, 0.3])
network['W2'] = np.array([[0.1, 0.4], [0.2, 0.5], [0.3, 0.6]])
network['b2'] = np.array([0.1, 0.2])
network['W3'] = np.array([[0.1, 0.3], [0.2, 0.4]])
network['b3'] = np.array([0.1, 0.2])
return network
def forward(network, x):
W1, W2, W3 = network['W1'], network['W2'], network['W3']
b1, b2, b3 = network['b1'], network['b2'], network['b3']
a1 = np.dot(x, W1) + b1
z1 = sigmoid(a1)
a2 = np.dot(z1, W2) + b2
z2 = sigmoid(a2)
a3 = np.dot(z2, W3) + b3
y = identity_function(a3)
return y
network = init_network()
x = np.array([1.0, 0.5])
y = forward(network, x)
print(y)
# -
# ### 3.5.1 恒等関数とソフトマックス関数
a = np.array([0.3, 2.9, 4.0])
exp_a = np.exp(a)
print(exp_a)
sum_exp_a = np.sum(exp_a)
print(sum_exp_a)
# ### 3.5.2 ソフトマックス関数の実装上の注意
a = np.array([1010, 1000, 990])
np.exp(a) / np.sum(np.exp(a))
c = np.max(a)
a - c
np.exp(a - c) / np.sum(np.exp(a - c))
def softmax(a):
c = np.max(a)
exp_a = np.exp(a - c)
sum_exp_a = np.sum(exp_a)
y = exp_a / sum_exp_a
return y
a = np.array([0.3, 2.9, 4.0])
y = softmax(a)
print(y)
np.sum(y)
# ### 3.6.1 MNISTデータセット
# +
import sys, os
sys.path.append(os.pardir)
from dataset.mnist import load_mnist
(x_train, t_train), (x_test, t_test) =\
load_mnist(flatten=True, normalize=False)
# それぞれのデータの形状を出力
print(x_train.shape)
print(t_train.shape)
print(x_test.shape)
print(t_test.shape)
# +
import sys, os
sys.path.append(os.pardir)
import numpy as np
from dataset.mnist import load_mnist
from PIL import Image
def img_show(img):
pil_img = Image.fromarray(np.uint8(img))
pil_img.show()
(x_train, t_train), (x_test, t_test) =\
load_mnist(flatten=True, normalize=False)
img = x_train[0]
label = t_train[0]
print(label)
print(img.shape)
img = img.reshape(28, 28)
print(img.shape)
img_show(img)
# +
def get_data():
(x_train, t_train), (x_test, t_test) =\
load_mnist(normalize=True, flatten=True, one_hot_label=False)
return x_test, t_test
def init_network():
with open("sample_weight.pkl", 'rb') as f:
network = pickle.load(f)
return network
def predict(network, x):
W1, W2, W3 = network['W1'], network['W2'], network['W3']
b1, b2, b3 = network['b1'], network['b2'], network['b3']
a1 = np.dot(x, W1) + b1
z1 = sigmoid(a1)
a2 = np.dot(z1, W2) + b2
z2 = sigmoid(a2)
a3 = np.dot(z2, W3) + b3
y = softmax(a3)
return y
| .ipynb_checkpoints/m_ch03-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Dependencies
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _kg_hide-input=false _kg_hide-output=true _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
from utillity_script_cloud_segmentation import *
seed = 0
seed_everything(seed)
warnings.filterwarnings("ignore")
# -
# ### Load data
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _kg_hide-input=true _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
train = pd.read_csv('../input/understanding_cloud_organization/train.csv')
hold_out_set = pd.read_csv('../input/clouds-data-split/hold-out.csv')
X_train = hold_out_set[hold_out_set['set'] == 'train']
X_val = hold_out_set[hold_out_set['set'] == 'validation']
print('Compete set samples:', len(train))
print('Train samples: ', len(X_train))
print('Validation samples: ', len(X_val))
# Preprocecss data
train['image'] = train['Image_Label'].apply(lambda x: x.split('_')[0])
display(X_train.head())
# -
# # Model parameters
# +
BACKBONE = 'efficientnetb0'
BATCH_SIZE = 16
EPOCHS = 30
LEARNING_RATE = 3e-4
HEIGHT = 320
WIDTH = 480
CHANNELS = 3
N_CLASSES = 4
ES_PATIENCE = 7
RLROP_PATIENCE = 2
DECAY_DROP = 0.2
STEP_SIZE_TRAIN = len(X_train)//BATCH_SIZE
STEP_SIZE_VALID = len(X_val)//BATCH_SIZE
model_path = '55-unet_%s_%sx%s.h5' % (BACKBONE, HEIGHT, WIDTH)
train_images_path = '../input/cloud-images-resized-320x480/train_images320x480/train_images/'
# + _kg_hide-input=false
preprocessing = sm.get_preprocessing(BACKBONE)
augmentation = albu.Compose([albu.HorizontalFlip(p=0.5),
albu.VerticalFlip(p=0.5),
albu.GridDistortion(p=0.5),
albu.ShiftScaleRotate(scale_limit=0.5, rotate_limit=0,
shift_limit=0.1, border_mode=0, p=0.5)
])
# -
# ### Data generator
# + _kg_hide-input=false
train_generator = DataGenerator(
directory=train_images_path,
dataframe=X_train,
target_df=train,
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
augmentation=augmentation,
seed=seed)
valid_generator = DataGenerator(
directory=train_images_path,
dataframe=X_val,
target_df=train,
batch_size=BATCH_SIZE,
target_size=(HEIGHT, WIDTH),
n_channels=CHANNELS,
n_classes=N_CLASSES,
preprocessing=preprocessing,
seed=seed)
# -
# # Model
# + _kg_hide-input=false _kg_hide-output=true
model = sm.Unet(backbone_name=BACKBONE,
encoder_weights='imagenet',
classes=N_CLASSES,
activation='sigmoid',
input_shape=(None, None, CHANNELS))
checkpoint = ModelCheckpoint(model_path, monitor='val_loss', mode='min', save_best_only=True)
es = EarlyStopping(monitor='val_loss', mode='min', patience=ES_PATIENCE, restore_best_weights=True, verbose=1)
rlrop = ReduceLROnPlateau(monitor='val_loss', mode='min', patience=RLROP_PATIENCE, factor=DECAY_DROP, verbose=1)
metric_list = [dice_coef, sm.metrics.iou_score, sm.metrics.f1_score]
callback_list = [checkpoint, es, rlrop]
optimizer = RAdam(learning_rate=LEARNING_RATE, warmup_proportion=0.1)
model.compile(optimizer=optimizer, loss=sm.losses.bce_dice_loss, metrics=metric_list)
model.summary()
# + _kg_hide-input=true _kg_hide-output=true
history = model.fit_generator(generator=train_generator,
steps_per_epoch=STEP_SIZE_TRAIN,
validation_data=valid_generator,
validation_steps=STEP_SIZE_VALID,
callbacks=callback_list,
epochs=EPOCHS,
verbose=2).history
# -
# ## Model loss graph
# + _kg_hide-input=true
plot_metrics(history, metric_list=['loss', 'dice_coef', 'iou_score', 'f1-score'])
| Model backlog/Training/Segmentation/Kaggle/55-unet-efficientnetb0.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Representing image using matching pursuit (MP) with Gaussian basis functions
#
# This notebook uses a variant of the matching pusuit algorithm (Elad, 2014) to decompose a frame from a microscopy video into a weighted sum of Gaussian basis functions.
# %load_ext autoreload
# %autoreload
# %matplotlib inline
# +
import h5py
import dpmeans
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as widgets
from skimage.util import img_as_float, img_as_uint
from scipy import ndimage
from scipy.stats import multivariate_normal
from otimage import io
from otimage.imagerep import greedy_mp
# -
# ### Parameters
# +
# Relative threshold applied to image
THRESHOLD_REL = 0.15
# Minimum number of pixels for component to be considered
CELL_MIN_SIZE = 50
# -
# ### Load Zimmer data
# +
# Path to file containing Zimmer data
in_fpath = '/home/mn2822/Desktop/WormOT/data/zimmer/mCherry_v00065-01581.hdf5'
# Index of frame to use
t_frame = 500
with io.ZimmerReader(in_fpath) as reader:
frame = reader.get_frame(t_frame)
# -
# ### Select section of frame for test image
# +
# Section of XY plane where head is found
head_x = (500, 650)
head_y = (250, 525)
head_z = (0, 33)
# Extract section
img = frame[head_x[0]:head_x[1], head_y[0]:head_y[1], head_z[0]:head_z[1]]
img = img_as_float(img)
# Compute image min and max
img_min = np.min(img)
img_max = np.max(img)
# Display max projection
plt.imshow(np.max(img, 2).T);
plt.axis('off');
# +
wx = 15
wy = 15
wz = 5
xg, yg, zg = np.mgrid[-wx:wx+1, -wy:wy+1, -wz:wz+1]
weight_grid = np.stack((xg, yg, zg), axis=-1)
mean = np.array([0, 0, 0])
cov = np.diag([8.0, 8.0, 1.5])
weights_nn = multivariate_normal.pdf(weight_grid, mean=mean, cov=cov)
weights = weights_nn / np.sqrt(np.sum(weights_nn ** 2))
plt.imshow(np.max(weights, 2).T, origin='lower', extent=[-15, 15, -15, 15])
plt.title('Convolution weights')
plt.xlabel('x')
plt.ylabel('y')
plt.colorbar();
# -
# %%time
pts, vals, img_conv = greedy_mp(img, weights, 1)
# +
img_rep = np.zeros((img.shape[0] + 2 * wx, img.shape[1] + 2 * wy, img.shape[2] + 2 * wz))
pt = pts[0]
pt_mod = pt + np.array([wx, wy, wz])
x_sl = slice(pt_mod[0] - wx, pt_mod[0] + wx + 1)
y_sl = slice(pt_mod[1] - wy, pt_mod[1] + wy + 1)
z_sl = slice(pt_mod[2] - wz, pt_mod[2] + wz + 1)
img_rep[x_sl, y_sl, z_sl] = vals[0] * weights
img_rep = img_rep[wx:-wx, wy:-wy, wz:-wz]
plt.figure(figsize=(12, 12))
plt.subplot(131)
plt.imshow(np.max(img, 2).T);
plt.axis('off');
plt.title('original image')
plt.subplot(132)
plt.imshow(np.max(img_rep, 2).T)
plt.axis('off')
plt.title('first MP component');
plt.subplot(133)
plt.imshow(np.max(img_conv, 2).T)
plt.axis('off')
plt.title('convolved residual');
# +
points, vals, img_conv = greedy_mp(img, weights, 300)
plt.imshow(np.max(img_conv, 2).T)
plt.colorbar()
# +
recon_img = np.zeros((img.shape[0] + 2 * wx, img.shape[1] + 2 * wy, img.shape[2] + 2 * wz))
points_mod = points + np.array([wx, wy, wz])
for i in range(points_mod.shape[0]):
pt = points_mod[i]
x_sl = slice(pt[0] - wx, pt[0] + wx + 1)
y_sl = slice(pt[1] - wy, pt[1] + wy + 1)
z_sl = slice(pt[2] - wz, pt[2] + wz + 1)
recon_img[x_sl, y_sl, z_sl] += vals[i] * weights
r_img = recon_img[wx:-wx, wy:-wy, wz:-wz]
# +
plt.figure(figsize=(10, 10))
plt.subplot(131)
plt.imshow(np.max(img, 2).T)
plt.title('original image')
plt.subplot(132)
plt.imshow(np.max(r_img, 2).T)
plt.title('reconstruction')
sq_res_img = (r_img - img) ** 2
plt.subplot(133)
plt.imshow(np.max(sq_res_img, 2).T)
plt.title('residual')
plt.savefig('greedy_mp_1.png')
# -
| python/notebooks/image_representation/04_matching_pursuit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image restoration / segmentation project
# ## Maxflow - mincut example
#
# maxflow is a Python module for max-flow/min-cut computations. It wraps the C++ maxflow library by <NAME>, which implements the algorithm described in
#
# An Experimental Comparison of Min-Cut/Max-Flow Algorithms for Energy Minimization in Vision. <NAME> and <NAME>. TPAMI.
#
# This module aims to simplify the construction of graphs with complex layouts. It provides two Graph classes, Graph[int] and Graph[float], for integer and real data types.
#
# To install, run
#
# pip install PyMaxFlow
#
# for your python. Here we assume python3, but python2 is also supported
#
# +
## Once installed, you can use the maxflow package
import maxflow
# -
# ## Graph example
#
# <img src="graph.png" alt="Graph example" style="width: 200px;"/>
# +
g = maxflow.Graph[int](2, 2)
# Add two (non-terminal) nodes. Get the index to the first one.
nodes = g.add_nodes(2)
# Create two edges (forwards and backwards) with the given capacities.
# The indices of the nodes are always consecutive.
g.add_edge(nodes[0], nodes[1], 1, 2)
# Set the capacities of the terminal edges...
# ...for the first node.
g.add_tedge(nodes[0], 2, 5)
# ...for the second node.
g.add_tedge(nodes[1], 9, 4)
flow = g.maxflow()
print("Maximum flow:", flow)
# -
print("Segment of the node 0:", g.get_segment(nodes[0]))
print("Segment of the node 1:", g.get_segment(nodes[1]))
# ### interpretation
# The result can be interpreted in the following way:
#
# <img src="graph2.png" alt="Graph result" style="width: 200px;"/>
# ### Flow network from an image
#
# Flow network can be directly computed from an image, which is very convenient:
# http://pmneila.github.io/PyMaxflow/maxflow.html
#
# We use as example a noisy image of the letter A:
#
#
# <img src="a2.png" alt="Graph result" style="width: 80px;"/>
# +
import numpy as np
import scipy
from scipy.misc import imread
import maxflow
img = imread("a2.png")
# -
# ### Image restoration
#
# We will restore the image minimizing the energy
#
# $$E(\mathbf{x}) = \sum_i D_i(x_i) + \lambda \sum_{(i,j)\in\mathcal{C}} |x_i - x_j|$$,
#
# where $\mathbf{x} \in \{0,1\}^N$ are the values of the restored image, $N$ is the number of pixels. The unary term $D_i(0)$ (resp $D_i(1)$) is the penalty for assigning the value 0 (resp 1) to the i-th pixel. Each $D_i$ depends on the values of the noisy image, which are denoted as $p_i$:
#
# $D_i(x_i)$ = \begin{cases} p_i & \textrm{if } x_i=0\\
# 255-p_i & \textrm{if } x_i=1 \end{cases}.
#
# Thus, $D_i$ is low when assigning the label 0 to dark pixels or the label 1 to bright pixels, and high otherwise. The value $\lambda$ is the regularization strength. The larger $\lambda$ the smoother the restoration. We set it to 50. This $\lambda$ does not need to be a fixed constant for every pixel, but in this case it is. It may not depend on $\mathbf{x}$.
#
# The maximum flow algorithm is widely used to minimize energy functions of this type. We build a graph which represents the above energy. This graph has as many non-terminal nodes as pixels in the image. The nodes are connected in a grid arrangement, so that the nodes corresponding to neighbor pixels are connected by a forward and a backward edge. The capacities of all non-terminal edges is $\lambda$. The capacities of the edges from the source node are set to $D_i(0)$, and the capacities of the edges to the sink node are $D_i(1)$.
#
# We could build this graph as in the first example. First, we would add all the nodes. Then, we would iterate over the nodes adding the edges properly. However, this is extremely slow in Python, especially when dealing with large images or stacks of images. PyMaxflow provides methods for building some complex graphs with a few calls. In this example we review add_grid_nodes, add_grid_edges, which add edges with a fixed capacity to the grid, and add_grid_tedges, which sets the capacities of the terminal edges for multiple nodes:
# Create the graph.
g = maxflow.Graph[int]()
# Add the nodes. nodeids has the identifiers of the nodes in the grid.
nodeids = g.add_grid_nodes(img.shape)
# Add non-terminal edges with the same capacity.
g.add_grid_edges(nodeids, 50)
# Add the terminal edges. The image pixels are the capacities
# of the edges from the source node. The inverted image pixels
# are the capacities of the edges to the sink node.
g.add_grid_tedges(nodeids, img, 255-img)
# Finally, we perform the maxflow computation and get the results:
# Find the maximum flow.
g.maxflow()
# Get the segments of the nodes in the grid.
sgm = g.get_grid_segments(nodeids)
# The method get_grid_segments returns an array with the same shape than nodeids. It is almost equivalent to calling get_segment once for each node in nodeids, but much faster. For the i-th cell, the array stores False if the i-th node belongs to the source segment (i.e., the corresponding pixel has the label 1) and True if the node belongs to the sink segment (i.e., the corresponding pixel has the label 0). We now get the labels for each pixel:
# +
# %matplotlib notebook
# The labels should be 1 where sgm is False and 0 otherwise.
img2 = np.int_(np.logical_not(sgm))
# Show the result.
from matplotlib import pyplot as plt
plt.figure(figsize=(2,2))
plt.imshow(img)
plt.figure(figsize=(2,2))
plt.imshow(img2)
plt.show()
# -
# ### More complex layouts
#
# The method add_grid_edges is a powerful tool to create complex layouts. The first argument, nodeids is an array of node identifiers with the shape of the grid of nodes where the edges will be added. The edges to add and their final capacities are computed using the arguments weights and structure.
#
# weights is an array and its shape must be broadcastable to the shape of nodeids. Thus every node will have a associated weight. structure is an array with the same dimensions as nodeids and with an odd shape. It defines the local neighborhood of every node.
#
# Given a node, the structure array is centered on it. Edges are created from that node to the nodes of its neighborhood corresponding to nonzero entries of structure. The capacity of the new edge will be the product of the weight of the initial node and the corresponding value in structure. Additionally, a reverse edge with the same capacity will be added if the argument symmetric is True (by default).
#
# Therefore, the weights argument allows to define an inhomogeneous graph, with different capacities in different areas of the grid. On the other hand, besides defining the local neighborhood of each node, structure enables anisotropic edges, with different capacities depending on their orientation.
#
# ### Extension example on github
#
# The github of the PyMaxFlow package is here: https://github.com/pmneila/PyMaxflow
#
# The file examples/layout_examples.py and the documentation of maxflow.GraphInt.add_grid_edges() contain several different layouts than can be created with add_grid_edges. A more involved example is in examples/layout_example2.py, where a complex graph is created using several calls to maxflow.GraphInt.add_grid_edges() and maxflow.GraphInt.add_grid_tedges().
| files/maxflow_segmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/hhk54250/20MA573-HHK/blob/master/hw11/Untitled6.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="Vrt6miKWHcww" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9a9a18db-2cfd-442e-f3a9-ac04290d7971"
import scipy.stats as ss
import numpy as np
c = ss.norm(0,1).cdf(-2)
print("the exact price is", c)
# + id="Sex1jsuqHg6k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f108e009-a72d-4139-a49a-37dda71bad31"
def OMC(N,alpha):
s=0
for i in range(N):
if np.random.normal(0,1)<-2:
s+=1
return s/N
OMC(1000,2)
# + id="ur88UmdlHjmR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d4a1d0ad-b673-46d3-ad44-10f428981a3f"
def IS(N,alpha):
s=0
for i in range(N):
A= np.random.normal(-alpha,1)
if A<-2:
s+= np.exp(alpha**2/2 + alpha*A)
return s/N
IS(1000,2)
# + id="7r9q_Ti_Hsa0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="14e5d6be-3009-4d68-d2b8-e2c22620b1fa"
var_IS=0
var_omc=0
for i in range(10000):
var_omc+= (c - OMC(1000,2))**2
var_IS += (c - IS(1000,2))**2
mse_omc = var_omc/10000
mse_IS = var_IS/10000
print(mse_omc)
print(mse_IS)
# + id="z1Gh-wHQHwhI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="f4048710-f836-4ec4-b3d2-7b0532e9f1a8"
def MSE_OMC(alpha_hat,N):
for i in range (N):
x=ss.norm(0,1).cdf(-2)
MSE_OMC = (1/N)*(x-x**2)
return MSE_OMC
MSE_OMC(3,10000)
# + id="vwmWwkknH5Nl" colab_type="code" colab={}
def MSE_IS(alpha_hat,N):
for i in range (N):
x=ss.norm(0,1).cdf(-2)
y=ss.norm(0,1).cdf(-2-alpha_hat)
mse_IS = (1/N)*np.exp(alpha_hat**2)*y-(x**2)
return mse_IS
MSE_IS(2,10000)
# + id="WaU3T_v9IArX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 182} outputId="abd0bf30-aca9-49bd-c64f-692d3c8c3bcc"
diff = MSE_OMC(3,10000)-MSE_IS(3,10000)
diff
| hw11/Untitled6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import itertools
from keras.preprocessing import text, sequence
from keras import utils
from sklearn.preprocessing import LabelBinarizer, LabelEncoder
from keras.layers import Dense, Activation, Dropout
from keras.models import Sequential
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.metrics import confusion_matrix
df=pd.read_json("Team12_Chevron.json")
df.head(5)
df['sentiment'].value_counts()
train_size = int(len(df) * .8)
print ("Train size: %d" % train_size)
print ("Test size: %d" % (len(df) - train_size))
# +
train_posts = df['text'][:train_size]
train_tags = df['sentiment'][:train_size]
test_posts = df['text'][train_size:]
test_tags = df['sentiment'][train_size:]
# -
max_words = 1000
tokenize = text.Tokenizer(num_words=max_words, char_level=False)
tokenize.fit_on_texts(train_posts) # only fit on train
x_train = tokenize.texts_to_matrix(train_posts)
x_test = tokenize.texts_to_matrix(test_posts)
# Use sklearn utility to convert label strings to numbered index
encoder = LabelEncoder()
encoder.fit(train_tags)
y_train = encoder.transform(train_tags)
y_test = encoder.transform(test_tags)
# Converts the labels to a one-hot representation
num_classes = np.max(y_train) + 1
y_train = utils.to_categorical(y_train, num_classes)
y_test = utils.to_categorical(y_test, num_classes)
# Inspect the dimenstions of our training and test data (this is helpful to debug)
print('x_train shape:', x_train.shape)
print('x_test shape:', x_test.shape)
print('y_train shape:', y_train.shape)
print('y_test shape:', y_test.shape)
# This model trains very quickly and 2 epochs are already more than enough
# Training for more epochs will likely lead to overfitting on this dataset
# You can try tweaking these hyperparamaters when using this model with your own data
batch_size = 32
epochs = 2
# +
# Build the model
model = Sequential()
model.add(Dense(512, input_shape=(max_words,)))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(num_classes))
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# -
history = model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_split=0.1)
score = model.evaluate(x_test, y_test,
batch_size=batch_size, verbose=1)
print('Test score:', score[0])
print('Test accuracy:', score[1])
# +
text_labels = encoder.classes_
for i in range(10):
prediction = model.predict(np.array([x_test[i]]))
predicted_label = text_labels[np.argmax(prediction)]
print(test_posts.iloc[i][:50], "...")
print('Actual label:' + test_tags.iloc[i])
print("Predicted label: " + predicted_label + "\n")
# +
y_softmax = model.predict(x_test)
y_test_1d = []
y_pred_1d = []
for i in range(len(y_test)):
probs = y_test[i]
index_arr = np.nonzero(probs)
one_hot_index = index_arr[0].item(0)
y_test_1d.append(one_hot_index)
for i in range(0, len(y_softmax)):
probs = y_softmax[i]
predicted_index = np.argmax(probs)
y_pred_1d.append(predicted_index)
# -
def plot_confusion_matrix(cm, classes,
title='Confusion matrix',
cmap=plt.cm.Blues):
"""
This function prints and plots the confusion matrix.
Normalization can be applied by setting `normalize=True`.
"""
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title, fontsize=30)
plt.colorbar()
tick_marks = np.arange(len(classes))
plt.xticks(tick_marks, classes, rotation=45, fontsize=22)
plt.yticks(tick_marks, classes, fontsize=22)
fmt = '.2f'
thresh = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
plt.text(j, i, format(cm[i, j], fmt),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.ylabel('True label', fontsize=25)
plt.xlabel('Predicted label', fontsize=25)
cnf_matrix = confusion_matrix(y_test_1d, y_pred_1d)
plt.figure(figsize=(24,20))
plot_confusion_matrix(cnf_matrix, classes=text_labels, title="Confusion matrix")
plt.show()
| Uncovering Sentiments using EDGAR Datasets/BOW model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from multiprocessing import cpu_count
import numpy as np
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
from pycebox.ice import ice, ice_plot, pdp
from pdpbox.info_plots import target_plot, actual_plot
from pdpbox.pdp import pdp_isolate, pdp_plot
import seaborn as sns
import sklearn
from sklearn.model_selection import train_test_split
from sklearn.neural_network import MLPClassifier
from sklearn.ensemble import GradientBoostingClassifier, GradientBoostingRegressor
from sklearn.metrics import accuracy_score, classification_report, confusion_matrix, r2_score
# -
notes = pd.read_csv('../data/banknote.csv')
notes
X = notes.drop(columns=['Class'])
y = notes['Class']
# +
test_size=0.8
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size, random_state=42)
gbc = GradientBoostingClassifier()
gbc.fit(X_train.values, y_train)
y_pred = gbc.predict(X_test.values)
print("Accuracy Score: %f" %(accuracy_score(y_pred, y_test)))
print(confusion_matrix(y_pred, y_test))
print(classification_report(y_pred, y_test))
df = X_train
features = list(range(len(df.columns)))
feature_names = df.columns
plt.figure(1, figsize=(15, 15))
f, ax = plot_partial_dependence(gbc, X_train, features=features, feature_names=feature_names, n_cols=4, n_jobs=cpu_count())
# +
test_size=0.015
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=test_size, random_state=42)
mlp = MLPClassifier(hidden_layer_sizes=(25,), alpha=0.001)
mlp.fit(X_train.values, y_train)
y_pred = mlp.predict(X_test.values)
y_prob = mlp.predict_proba(X_test)[:, 0]
print("Accuracy: %f" %(accuracy_score(y_pred, y_test)))
print(confusion_matrix(y_pred, y_test))
print(classification_report(y_pred, y_test))
df = X_train
features = list(range(len(df.columns)))
feature_names = df.columns
# for i in range(len(df.columns)):
# plt.figure(i, figsize=(10, 10))
# out = pdp_isolate(mlp, X, model_features=feature_names, feature=feature_names[i], n_jobs=cpu_count())
# plot_params = {
# 'title':'Partial dependence plot (PDP) for %s' %(feature_names[i]),
# 'xlabel':'%s' %(feature_names[i]),
# 'ylabel':'Authenticity of Banknote'
# }
# pdp_plot(out, feature_names[i], figsize=(10, 10), ncols=1, plot_params=plot_params)
# plt.savefig('../Plots/PDPPlotWhole{}.png'.format(feature_names[i]))
# plt.savefig('../Plots/PDPPlotWhole{}.pdf'.format(feature_names[i]))
# for i in range(len(df.columns)):
# plt.figure(i, figsize=(10, 10))
# actual_plot(mlp, X_train)
def callable(X_test):
return mlp.predict_proba(X_test)[:, 0]
for i in range(len(df.columns)):
plt.figure(i, figsize=(10, 10))
ax = plt.gca()
ice_data = ice(X_test, feature_names[i], callable)
ice_plot(ice_data=ice_data, c='k', plot_points=True,plot_pdp=True, cmap='RdBu', ax=ax)
ax.grid()
ax.set_xlabel('%s' % (feature_names[i]))
ax.set_ylabel('Authenticity of Banknote')
ax.set_title('ICE Plots for %s' %(feature_names[i]))
# plt.savefig('../Plots/ICEPlot{}Centered.png'.format(feature_names[i]))
# plt.savefig('../Plots/ICEPlot{}Centered.pdf'.format(feature_names[i]))
# -
| src/ICEPlots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import scipy.stats as stats
from sklearn.preprocessing import MinMaxScaler
maxrapsodo = pd.read_csv('maxrapsodo.csv')
allgeyerrap = pd.read_csv('rapsodonew.csv')
allgeyertm = pd.read_csv('allgeyertm.csv')
allgeyerrap = allgeyerrap.round(1)
allgeyerrap = allgeyerrap.sort_values(by=['Date'])
allgeyerrap = allgeyerrap.rename(columns={"Pitch Type": "PitchType1"})
allgeyerrap
allgeyertm = allgeyertm.sort_values(by=['Pitch Time'])
allgeyertm = allgeyertm.drop([8,2])
allgeyertm
combine = allgeyertm.join(allgeyerrap)
combine = combine.drop(columns=['Pitcher Name','Pitch Time','Pitch ID','Horizontal Approach Angle (°)','Zone Location','No','Strike Zone Side','Is Strike','Strike Zone Height','tilt_normed_vert','tilt_normed_horz','Gyro Degree (deg)',
'Spin Efficiency (SZ)','Spin Rate (rpm)','Release Height','Release Side'])
#combine = combine.drop(columns=['Pitcher Name','Pitch Time','Pitch ID','Horizontal Approach Angle (°)',
# 'Zone Location','No','Strike Zone Side','Is Strike','Strike Zone Height','tilt_normed_vert',
# 'tilt_normed_horz','Gyro Degree (deg)'])
combine.columns
#combine['Tilt'] = pd.to_datetime(combine['Tilt'], format='%H:%M:%S').dt.strftime('%M:%S')
combine["Tilt"] = combine["Tilt"].str.replace(":","")
combine["Spin Axis"] = combine["Spin Axis"].str.replace(":","")
combine["Spin Axis"] = combine["Spin Axis"].astype(float)
combine["Tilt"] = combine["Tilt"].astype(float)
combine = combine.sort_values(["Date", "Release Speed (mph)",'Speed','VB','Induced Vertical Break (in)'],
ascending = (True, True, True, True, True))
combine
combine['axisdev'] = np.subtract(combine['Spin Axis'], combine['Tilt'])
df = pd.read_csv('sample.csv')
df
df["Spin Axis"] = df["Spin Axis"].str.replace(":","")
df["Tilt"] = df["Tilt"].str.replace(":","")
df["Spin Axis"] = df["Spin Axis"].astype(int)
df["Tilt"] = df["Tilt"].astype(int)
grouped = df.groupby(df['Pitch Type'])
sinker = grouped.get_group("Sinker")
changeup = grouped.get_group("ChangeUp")
slider = grouped.get_group("Slider")
sinker['axisdev'] = np.subtract(sinker['Tilt'], sinker['Spin Axis'])
slider['axisdev'] = np.subtract(slider['Tilt'], slider['Spin Axis'])
changeup['axisdev'] = np.subtract(changeup['Tilt'], changeup['Spin Axis'])
sinker['hbdiff'] = np.subtract(sinker['Horizontal Break'], sinker['HB'])
slider['hbdiff'] = np.subtract(slider['Horizontal Break'], slider['HB'])
changeup['hbdiff'] = np.subtract(changeup['Horizontal Break'], changeup['HB'])
sinker['vbdiff'] = np.subtract(sinker['Induced Vertical Break'], sinker['VB'])
slider['vbdiff'] = np.subtract(slider['Induced Vertical Break'], slider['VB'])
changeup['vbdiff'] = np.subtract(changeup['Induced Vertical Break'], changeup['VB'])
sinker = sinker.drop(columns=['Pitch Type'])
changeup = changeup.drop(columns=['Pitch Type'])
slider = slider.drop(columns=['Pitch Type'])
sinkerz = sinker.apply(stats.zscore)
changeupz = changeup.apply(stats.zscore)
sliderz = slider.apply(stats.zscore)
sinkerheight = sinkerz["Release Height"][1]
sinkerside = sinkerz['Release Side'][1]
sinkerdev = sinkerz['axisdev'][1]
sinkerhb = sinkerz['hbdiff'][1]
sinkervb = sinkerz['vbdiff'][1]
sinkervelo = sinker['Speed'].mean()
value = (sinkervelo*.37) +(sinkervb*.05) + (sinkerhb*.25) + (sinkerdev*.1) + (sinkerside*.11) + (sinkerheight*.12)
value
def stuff(df):
for i in range(len(df)):
value = (df['Speed'][i]*.37) +(df['vbdiff'][i]*.05) + (df['hbdiff'][i]*.25) + (df['axisdev'][i]*.1) + (df['Release Side'][i]*.11) + (df['Release Height'][i]*.12)
#df['Value'] = ''
df['Value'][i] = value
stuff(sinkerz)
# +
#sinkerz['Value'] = MinMaxScaler().fit_transform(np.array(sinkerz['Value']).reshape(-1,1))
# -
sinkerz
full = pd.read_csv('full1.csv')
pitches = ['Sinker']
full = full[full['Pitch Type'].isin(pitches)]
full = full.dropna()
full = full.reset_index(drop=True)
full = full.drop(columns = ['tilt_normed_vert','tilt_normed_horz','Pitch Time','Zone Location','Location Height (ft)', 'Location Side (ft)',
'Pitch Type','Pitcher Name'])
full["Spin Axis"] = full["Spin Axis"].str.replace(":","")
full["Tilt"] = full["Tilt"].str.replace(":","")
full["Spin Axis"] = full["Spin Axis"].astype(int)
full["Tilt"] = full["Tilt"].astype(int)
full['axisdev'] = np.subtract(full['Tilt'], full['Spin Axis'])
full['hbdiff'] = np.subtract(full['Horizontal Break'], full['HB'])
full['vbdiff'] = np.subtract(full['Induced Vertical Break'], full['VB'])
nfull = full.apply(stats.zscore)
nfull['Value'] = ''
stuff(nfull)
full
scaler = MinMaxScaler(feature_range=(20,80))
nfull['Value'] = scaler.fit_transform(np.array(nfull['Value']).reshape(-1,1))
display(nfull)
import matplotlib.tri as tri
import matplotlib.pyplot as plt
x = full['Horizontal Break']
y = full['Induced Vertical Break']
z = nfull['Value']
plt.tricontour(x, y, z, 15, linewidths=1.5, cmap='coolwarm')
plt.tricontourf(x, y, z, 15)
plt.ylabel('IVB')
plt.xlabel('HB')
plt.colorbar()
plt.grid()
sinkerzn = sinkerz.copy()
column = 'Value'
sinkerzn[column] = scaler.fit_transform(np.array(sinkerzn[column]).reshape(-1,1))
display(sinkerzn)
X = sinker['Horizontal Break']
Y = sinker['Induced Vertical Break']
Z = sinkerzn['Value']
plt.tricontour(X, Y, Z, 15, linewidths=1.5, cmap='coolwarm')
plt.tricontourf(X, Y, Z, 15)
plt.ylabel('IVB')
plt.xlabel('HB')
plt.colorbar()
plt.grid()
velo = input("Enter your velo: ")
velo = float(velo)
vbrap = input('Enter your vertical break on rapsodo: ')
vbrap = float(vbrap)
hbrap = input('Enter your horizontal break on rapsodo: ')
hbrap = float(hbrap)
vbtm = input('Enter your vertical break on trackman: ')
vbtm = float(vbtm)
hbtm = input('Enter your horizontal break on trackman: ')
hbtm = float(hbtm)
tiltrap = input('Enter your tilt on rapsodo without the colon: ')
tiltrap = float(tiltrap)
tilttm = input('Enter your tilt on trackman without the colon: ')
tilttm = float(tiltrap)
relh = input('Enter your release height: ')
rels = input('Enter your release side: ')
relh = float(relh)
rels = float(rels)
vbdiff = vbrap - vbtm
hbdiff = hbrap - hbtm
tiltdiff = tilttm - tiltrap
newrow = {'Speed':velo,'Induced Vertical Break':vbtm,'Horizontal Break':hbtm,'HB':hbrap,'VB':vbrap,
'Release Height':relh,'Release Side':rels,'vbdiff':vbdiff,'hbdiff':hbdiff,'axisdev':tiltdiff}
sinkerrow = sinker.append(newrow,ignore_index=True)
sinkerrowz = sinkerrow.apply(stats.zscore)
sinkerrowz['Value']= ''
stuff(sinkerrowz)
sinkerrowz['Value'] = scaler.fit_transform(np.array(sinkerrowz['Value']).reshape(-1,1))
display(sinkerrowz)
X = sinkerrow['Horizontal Break']
Y = sinkerrow['Induced Vertical Break']
Z = sinkerrowz['Value']
plt.tricontour(X, Y, Z, 15, linewidths=1.5, cmap='coolwarm')
plt.tricontourf(X, Y, Z, 15)
plt.ylabel('IVB')
plt.xlabel('HB')
plt.colorbar()
plt.grid()
snkvalue = (velo*.37) +(vbdiff*.05) + (hbdiff*.25) + (tiltdiff*.1) + (rels*.11) + (relh*.12)
snkvalue
snkz = (snkvalue - sinkerz.mean())/sinkerz.std()
snkz['Value']
def sinkerstuff():
velo = input("Enter your velo: ")
velo = float(velo)
vbrap = input('Enter your vertical break on rapsodo: ')
vbrap = float(vbrap)
hbrap = input('Enter your horizontal break on rapsodo: ')
hbrap = float(hbrap)
vbtm = input('Enter your vertical break on trackman: ')
vbtm = float(vbtm)
hbtm = input('Enter your horizontal break on trackman: ')
hbtm = float(hbtm)
tiltrap = input('Enter your tilt on rapsodo without the colon: ')
tiltrap = float(tiltrap)
tilttm = input('Enter your tilt on trackman without the colon: ')
tilttm = float(tiltrap)
relh = input('Enter your release height: ')
rels = input('Enter your release side: ')
relh = float(relh)
rels = float(rels)
vbdiff = vbrap - vbtm
hbdiff = hbrap - hbtm
tiltdiff = tilttm - tiltrap
newrow = {'Speed':velo,'Induced Vertical Break':vbtm,'Horizontal Break':hbtm,'HB':hbrap,'VB':vbrap,
'Release Height':relh,'Release Side':rels,'vbdiff':vbdiff,'hbdiff':hbdiff,'axisdev':tiltdiff}
global full
full = full.append(newrow,ignore_index=True)
fullz = full.apply(stats.zscore)
fullz['Value'] = ''
stuff(fullz)
fullz['Value'] = scaler.fit_transform(np.array(fullz['Value']).reshape(-1,1))
X = full['Horizontal Break']
Y = full['Induced Vertical Break']
Z = fullz['Value']
plt.tricontour(X, Y, Z, 15, linewidths=1.5, cmap='coolwarm')
plt.tricontourf(X, Y, Z, 15, cmap = 'coolwarm')
plt.ylabel('IVB')
plt.xlabel('HB')
plt.colorbar()
plt.grid()
print(fullz['Value'].iloc[-1])
sinkerstuff()
full
| stuff.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from collections import OrderedDict
from collections import namedtuple
import numpy as np
from scipy import stats
# R precision
def r_precision(targets, predictions, max_n_predictions=500):
# Assumes predictions are sorted by relevance
# First, cap the number of predictions
predictions = predictions[:max_n_predictions]
# Calculate metric
target_set = set(targets)
target_count = len(target_set)
return float(len(set(predictions[:target_count]).intersection(target_set))) / target_count
def dcg(relevant_elements, retrieved_elements, k, *args, **kwargs):
"""Compute the Discounted Cumulative Gain.
Rewards elements being retrieved in descending order of relevance.
\[ DCG = rel_1 + \sum_{i=2}^{|R|} \frac{rel_i}{\log_2(i + 1)} \]
Args:
retrieved_elements (list): List of retrieved elements
relevant_elements (list): List of relevant elements
k (int): 1-based index of the maximum element in retrieved_elements
taken in the computation
Note: The vector `retrieved_elements` is truncated at first, THEN
deduplication is done, keeping only the first occurence of each element.
Returns:
DCG value
"""
retrieved_elements = __get_unique(retrieved_elements[:k])
relevant_elements = __get_unique(relevant_elements)
if len(retrieved_elements) == 0 or len(relevant_elements) == 0:
return 0.0
# Computes an ordered vector of 1.0 and 0.0
score = [float(el in relevant_elements) for el in retrieved_elements]
# return score[0] + np.sum(score[1:] / np.log2(
# 1 + np.arange(2, len(score) + 1)))
return np.sum(score / np.log2(1 + np.arange(1, len(score) + 1)))
def ndcg(relevant_elements, retrieved_elements, k, *args, **kwargs):
"""Compute the Normalized Discounted Cumulative Gain.
Rewards elements being retrieved in descending order of relevance.
The metric is determined by calculating the DCG and dividing it by the
ideal or optimal DCG in the case that all recommended tracks are relevant.
Note:
The ideal DCG or IDCG is on our case equal to:
\[ IDCG = 1+\sum_{i=2}^{min(\left| G \right|, k)}\frac{1}{\log_2(i +1)}\]
If the size of the set intersection of \( G \) and \( R \), is empty, then
the IDCG is equal to 0. The NDCG metric is now calculated as:
\[ NDCG = \frac{DCG}{IDCG + \delta} \]
with \( \delta \) a (very) small constant.
The vector `retrieved_elements` is truncated at first, THEN
deduplication is done, keeping only the first occurence of each element.
Args:
retrieved_elements (list): List of retrieved elements
relevant_elements (list): List of relevant elements
k (int): 1-based index of the maximum element in retrieved_elements
taken in the computation
Returns:
NDCG value
"""
# TODO: When https://github.com/scikit-learn/scikit-learn/pull/9951 is
# merged...
idcg = dcg(
relevant_elements, relevant_elements, min(k, len(relevant_elements)))
if idcg == 0:
raise ValueError("relevent_elements is empty, the metric is"
"not defined")
true_dcg = dcg(relevant_elements, retrieved_elements, k)
return true_dcg / idcg
def __get_unique(original_list):
"""Get only unique values of a list but keep the order of the first
occurence of each element
"""
return list(OrderedDict.fromkeys(original_list))
Metrics = namedtuple('Metrics', ['r_precision', 'ndcg', 'plex_clicks'])
# playlist extender clicks
def playlist_extender_clicks(targets, predictions, max_n_predictions=500):
# Assumes predictions are sorted by relevance
# First, cap the number of predictions
predictions = predictions[:max_n_predictions]
# Calculate metric
i = set(predictions).intersection(set(targets))
for index, t in enumerate(predictions):
for track in i:
if t == track:
return float(int(index / 10))
return float(max_n_predictions / 10.0 + 1)
# def compute all metrics
def get_all_metrics(targets, predictions, k):
return Metrics(r_precision(targets, predictions, k),
ndcg(targets, predictions, k),
playlist_extender_clicks(targets, predictions, k))
MetricsSummary = namedtuple('MetricsSummary', ['mean_r_precision',
'mean_ndcg',
'mean_plex_clicks',
'coverage'])
# +
###skip
import os
Meta1Resultspath='/home/ubuntu/SpotifyChallenge/notebooks/Reranking/TestingQueryResults/Meta1/'
Meta2Resultspath='/home/ubuntu/SpotifyChallenge/notebooks/Reranking/TestingQueryResults/Meta2/'
QEPRFResultspath='/home/ubuntu/SpotifyChallenge/notebooks/Reranking/TestingQueryResults/QEPRF750/'
Meta1Files=[Meta1Resultspath+x for x in os.listdir(Meta1Resultspath)]
Meta2Files=[Meta2Resultspath+x for x in os.listdir(Meta2Resultspath)]
QEPRFFiles=[QEPRFResultspath+x for x in os.listdir(QEPRFResultspath)]
###skip
import codecs
def parseMetaFiles(path):
playlistId=path.split('/')[-1].split('.op')[0]
with codecs.open(path, 'r', encoding='utf-8') as f:
lines = f.read().splitlines()
rank=0
resultSet=[]
for result in lines[1:]:
try:
rank=rank+1
splits=result.split('\t')
score = splits[0]
trackid= splits[1]
resultSet.append((rank,trackid,score))
except:
print result
return "QueryError"
return(playlistId,resultSet)
####skip
Meta1Op=[]
err1=[]
Meta2Op=[]
err2=[]
for f in Meta1Files:
res=parseMetaFiles(f)
if res !="QueryError":
Meta1Op.append(res)
else:
err1.append(f)
for f in Meta2Files:
res=parseMetaFiles(f)
if res !="QueryError":
Meta2Op.append(res)
else:
err2.append(f)
####skip
import codecs
def QEPRFParse(path):
playlistId=path.split('/')[-1].split('.op')[0]
with codecs.open(path, 'r', encoding='utf-8') as f:
lines = f.read().splitlines()
inputQueries=lines[0].split('# query: ')[1].split()
resultSet=[]
pairResults= lines[1].split(' #weight(')[2].split(') )')[0].split('" ')
rank=0
for result in pairResults[:-1]:
try:
rank=rank+1
splits=result.split('"')
score = splits[0].strip()
trackid= splits[1].strip()
resultSet.append((rank,trackid,score))
except:
print result
return "QueryError"
return(playlistId,inputQueries,resultSet)
###skip
QEPRFOp=[]
err3=[]
for f in QEPRFFiles:
res=QEPRFParse(f)
if res !="QueryError":
QEPRFOp.append(res)
else:
err3.append(f)
###skip
import pickle
pidTrackMapping=pickle.load(open('./BiPartites/AllDataPidTrackListBipartite.pkl','rb'))
####skip
import pickle
import os
import codecs
from random import shuffle
pkl = os.listdir('./SplitsInformation/')
count=0
DS={}
for fpkl in pkl:
if fpkl in ['testing25RandPid.pkl', 'testing25Pid.pkl', 'testing1Pid.pkl', 'testing100Pid.pkl', 'testing10Pid.pkl', 'testing5Pid.pkl', 'testing100RandPid.pkl']:
testType=fpkl.replace('.pkl','')
if 'Rand' in fpkl:
listLen=int(fpkl.split('testing')[1].split('Rand')[0])
qtype='Rand'
else :
listLen=int(fpkl.split('testing')[1].split('Pid')[0])
qtype='Normal'
testingPids=pickle.load(open('./SplitsInformation/'+fpkl,'rb'))
for pid in testingPids:
pid=str(pid)
referenceSet=[x.replace('spotify:track:','') for x in pidTrackMapping[pid]]
DS[pid]=(testType,qtype,listLen,referenceSet)
####skip
import pickle
import os
import codecs
from random import shuffle
pkl = os.listdir('./SplitsInformation/')
testingTitleonlyPids=[]
for fpkl in pkl:
if fpkl =='testingOnlyTitlePid.pkl':
testType=fpkl.replace('.pkl','')
listLen=0
qtype='Normal'
testingPids=pickle.load(open('./SplitsInformation/'+fpkl,'rb'))
for pid in testingPids:
pid=str(pid)
referenceSet=[x.replace('spotify:track:','') for x in pidTrackMapping[pid]]
DS[pid]=(testType,qtype,listLen,referenceSet)
testingTitleonlyPids=[str(x) for x in testingPids]
# +
TestFile='./Training/ExternalAndW2VAsFeatures-BigRecall-TestingFile750-2080.txt'
with open(TestFile) as f:
test = f.readlines()
PidTestTracks={}
for l in test:
pid=l.split()[1].split(':')[1].strip()
track=l.split('#')[1].strip()
PidTestTracks.setdefault(pid,[]).append(track)
# -
len(PidTestTracks)
# +
scoresfile='./Training/External2080Model-500Trees-NDCG20-tc-1-lr01-leaf50.txt'
with open(scoresfile) as f:
scores = f.readlines()
from collections import defaultdict
from random import shuffle
PidTracksScores={}
for l in scores:
pid=l.split()[0].strip()
trackScore=l.split()[2].strip()
PidTracksScores.setdefault(pid,[]).append(float(trackScore))
rerankedCandidates={}
for pid,tracksList in PidTestTracks.items():
scoresList=PidTracksScores[pid]
zippedPairs=zip(tracksList,scoresList)
shuffle(zippedPairs)
rerankedCandidates[pid]=[x[0] for x in sorted(zippedPairs, key=lambda x: x[1], reverse=True)]
####continue here
evalSets=[]
for pl in QEPRFOp:
plId=pl[0]
if plId in rerankedCandidates:
exposed=pl[1]
candidates=rerankedCandidates[plId]
candidates=[x for x in candidates if x not in exposed]
refVals= DS[plId]
testtype=refVals[0]
orderType=refVals[1]
exposedLen=refVals[2]
playlist=refVals[3]
if orderType=='Normal':
groundTruth=playlist[exposedLen:]
else:
groundTruth=[x for x in playlist if x not in exposed]
evalSets.append((groundTruth, candidates[:500], testtype, exposedLen))
for pl in Meta2Op:
plId=pl[0]
if plId in testingTitleonlyPids and plId in rerankedCandidates:
exposed=[]
candidates=rerankedCandidates[plId]
refVals= DS[plId]
testtype=refVals[0]
orderType=refVals[1]
exposedLen=refVals[2]
playlist=refVals[3]
groundTruth=playlist[exposedLen:]
evalSets.append((groundTruth, candidates[:500], testtype, exposedLen))
####continue here
'''
r_precision(targets, predictions, k),
ndcg(targets, predictions, k),
playlist_extender_clicks(targets, predictions, k)
'''
indivSumsCounts= defaultdict(int)
indivSumsRecall = defaultdict(int)
indivSumsNdcg = defaultdict(int)
indivSumsRprec = defaultdict(int)
indivSumsClicks = defaultdict(int)
globalNdcg=0
globalRprec=0
globalClicks=0
globalRecall=0
count=0
for evalTuple in evalSets:
targets=evalTuple[0]
predictions=evalTuple[1]
testType=evalTuple[2]
tupNdcg=ndcg(targets,predictions,500)
tuprprec=r_precision(targets,predictions,500)
tupClicks=playlist_extender_clicks(targets,predictions,500)
globalNdcg+=tupNdcg
indivSumsNdcg[testType]+=tupNdcg
globalRprec+=tuprprec
indivSumsRprec[testType]+=tuprprec
globalClicks+=tupClicks
indivSumsClicks[testType]+=tupClicks
indivSumsCounts[testType]+=1
recallSetSize= len(set(predictions)&set(targets))
refSetSize=len(targets)
recall=recallSetSize*1.0/refSetSize
globalRecall+=recall
indivSumsRecall[testType]+=recall
count+=1
for k, v in indivSumsCounts.items():
indivSumsRecall[k]=indivSumsRecall[k]/v
indivSumsNdcg[k]=indivSumsNdcg[k]/v
indivSumsRprec[k]=indivSumsRprec[k]/v
indivSumsClicks[k]=indivSumsClicks[k]/v
print scoresfile , 'Recall:' , globalRecall/count,'NDCG:', globalNdcg/count, 'RPrec:', globalRprec/count,'Clicks:', globalClicks/count
| notebooks/FixedNewEvalReranking.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (tensorflow-2.0)
# language: python
# name: tensorflow-2.0
# ---
# # T81-558: Applications of Deep Neural Networks
# **Module 11: Natural Language Processing and Speech Recognition**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Module 11 Material
#
# * Part 11.1: Getting Started with Spacy in Python [[Video]](https://www.youtube.com/watch?v=bv_iVVrlfbU) [[Notebook]](t81_558_class_11_01_spacy.ipynb)
# * Part 11.2: Word2Vec and Text Classification [[Video]](https://www.youtube.com/watch?v=qN9hHlZKIL4) [[Notebook]](t81_558_class_11_02_word2vec.ipynb)
# * Part 11.3: What are Embedding Layers in Keras [[Video]](https://www.youtube.com/watch?v=Ae3GVw5nTYU) [[Notebook]](t81_558_class_11_03_embedding.ipynb)
# * **Part 11.4: Natural Language Processing with Spacy and Keras** [[Video]](https://www.youtube.com/watch?v=Ae3GVw5nTYU) [[Notebook]](t81_558_class_11_04_text_nlp.ipynb)
# * Part 11.5: Learning English from Scratch with Keras and TensorFlow [[Video]](https://www.youtube.com/watch?v=Ae3GVw5nTYU) [[Notebook]](t81_558_class_11_05_english_scratch.ipynb)
# # Part 11.4: Natural Language Processing with Spacy and Keras
#
# In this part we will see how to use Spacy and Keras together.
#
# ### Word-Level Text Generation
#
# There are a number of different approaches to teaching a neural network to output free-form text. The most basic question is if you wish the neural network to learn at the word or character level. In many ways, learning at the character level is the more interesting of the two. The LSTM is learning construct its own words without even being shown what a word is. We will begin with character-level text generation. In the next module, we will see how we can use nearly the same technique to operate at the word level. The automatic captioning that will be implemented in the next module is at the word level.
#
# We begin by importing the needed Python packages and defining the sequence length, named **maxlen**. Time-series neural networks always accept their input as a fixed length array. Not all of the sequence might be used, it is common to fill extra elements with zeros. The text will be divided into sequences of this length and the neural network will be trained to predict what comes after this sequence.
from tensorflow.keras.callbacks import LambdaCallback
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
from tensorflow.keras.layers import LSTM
from tensorflow.keras.optimizers import RMSprop
import numpy as np
import random
import sys
import io
import requests
import re
# +
import requests
r = requests.get("https://data.heatonresearch.com/data/t81-558/text/treasure_island.txt")
raw_text = r.text.lower()
print(raw_text[0:1000])
# +
import spacy
nlp = spacy.load("en_core_web_sm")
doc = nlp(raw_text)
vocab = set()
tokenized_text = []
for token in doc:
word = ''.join([i if ord(i) < 128 else ' ' for i in token.text])
word = word.strip()
if not token.is_digit \
and not token.like_url \
and not token.like_email:
vocab.add(word)
tokenized_text.append(word)
print(f"Vocab size: {len(vocab)}")
# -
print(list(vocab)[:20])
word2idx = dict((n, v) for v, n in enumerate(vocab))
idx2word = dict((n, v) for n, v in enumerate(vocab))
tokenized_text = [word2idx[word] for word in tokenized_text]
tokenized_text
# cut the text in semi-redundant sequences of maxlen words
maxlen = 6
step = 3
sentences = []
next_words = []
for i in range(0, len(tokenized_text) - maxlen, step):
sentences.append(tokenized_text[i: i + maxlen])
next_words.append(tokenized_text[i + maxlen])
print('nb sequences:', len(sentences))
sentences[0:5]
# +
import numpy as np
print('Vectorization...')
x = np.zeros((len(sentences), maxlen, len(vocab)), dtype=np.bool)
y = np.zeros((len(sentences), len(vocab)), dtype=np.bool)
for i, sentence in enumerate(sentences):
for t, word in enumerate(sentence):
x[i, t, word] = 1
y[i, next_words[i]] = 1
# -
x.shape
y.shape
y[0:5]
# +
# build the model: a single LSTM
print('Build model...')
model = Sequential()
model.add(LSTM(128, input_shape=(maxlen, len(vocab))))
model.add(Dense(len(vocab), activation='softmax'))
optimizer = RMSprop(lr=0.01)
model.compile(loss='categorical_crossentropy', optimizer=optimizer)
# -
model.summary()
def sample(preds, temperature=1.0):
# helper function to sample an index from a probability array
preds = np.asarray(preds).astype('float64')
preds = np.log(preds) / temperature
exp_preds = np.exp(preds)
preds = exp_preds / np.sum(exp_preds)
probas = np.random.multinomial(1, preds, 1)
return np.argmax(probas)
def on_epoch_end(epoch, _):
# Function invoked at end of each epoch. Prints generated text.
print("****************************************************************************")
print('----- Generating text after Epoch: %d' % epoch)
start_index = random.randint(0, len(tokenized_text) - maxlen - 1)
for temperature in [0.2, 0.5, 1.0, 1.2]:
print('----- temperature:', temperature)
#generated = ''
sentence = tokenized_text[start_index: start_index + maxlen]
#generated += sentence
o = ' '.join([idx2word[idx] for idx in sentence])
print(f'----- Generating with seed: "{o}"')
#sys.stdout.write(generated)
for i in range(100):
x_pred = np.zeros((1, maxlen, len(vocab)))
for t, word in enumerate(sentence):
x_pred[0, t, word] = 1.
preds = model.predict(x_pred, verbose=0)[0]
next_index = sample(preds, temperature)
next_word = idx2word[next_index]
#generated += next_char
sentence = sentence[1:]
sentence.append(next_index)
sys.stdout.write(next_word)
sys.stdout.write(' ')
sys.stdout.flush()
print()
# +
print_callback = LambdaCallback(on_epoch_end=on_epoch_end)
model.fit(x, y,
batch_size=128,
epochs=60,
callbacks=[print_callback])
# -
| t81_558_class_11_04_text_nlp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Decision Tree Hyperparameters in sklearn
# +
# #
# Classification and Regreession Tree
# 在每个节点上根据每一个维度d 和每一个阈值 v 进行二分得到的二叉树
# slearn 默认使用的是CART
# 当建立决策树时。在每个节点上都要进行二次划分。 预测O(logm) , 训练复杂度O(n*m*logm ) same with KNN 容易出现过拟合问题
# 非参数学习方法 - KNN and DT
# 剪枝 可以减低模型复杂度和防止过拟合
# 超参数:
# 1. max_depth - 最大树深度
# 2. min_samples_leaf - 每个叶子节点上最少多少个样本才继续往下划分
# 3. min_samples_split - 每个节点上最少多少个样本才继续往下划分
# 4. max_leaf_nodes - 最大的叶子节点数
# -
# ### Hyperparameters: max_depth/min_samples_leaf/min_samples_split/max_leaf_nodes
import numpy as np
import matplotlib.pyplot as plt
# +
from sklearn import datasets
X, y = datasets.make_moons(noise=0.20, random_state=200) # noise 比较大的情况下
# -
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
plt.show()
# +
from sklearn.tree import DecisionTreeClassifier
# max_depth 不做限制
dt_clf = DecisionTreeClassifier() #默认的是jini系数,max_step不做限制。这样会划分到每个决策点的jini系数为零为止。
dt_clf.fit(X, y)
# -
def plot_decision_boundary(model, axis):
x0, x1 = np.meshgrid(
np.linspace(axis[0], axis[1], int((axis[1]-axis[0])*100)).reshape(-1, 1),
np.linspace(axis[2], axis[3], int((axis[3]-axis[2])*100)).reshape(-1, 1),
)
X_new = np.c_[x0.ravel(), x1.ravel()]
y_predict = model.predict(X_new)
zz = y_predict.reshape(x0.shape)
from matplotlib.colors import ListedColormap
custom_cmap = ListedColormap(['#EF9A9A','#FFF59D','#90CAF9'])
plt.contourf(x0, x1, zz, cmap=custom_cmap)
plot_decision_boundary(dt_clf, axis=[-1.5, 2.5, -1.0, 1.5])
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
plt.show()
# +
# 传入max_depth 超参数来限制过拟合情况-
dt_clf2 = DecisionTreeClassifier(max_depth=2)
dt_clf2.fit(X, y)
plot_decision_boundary(dt_clf2, axis=[-1.5, 2.5, -1.0, 1.5])
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
plt.show()
# +
# 对于一个节点来数,要多少样本才继续对这几点拆分下去 - min_samples_split = 10
# 么有过拟合
dt_clf3 = DecisionTreeClassifier(min_samples_split=10)
dt_clf3.fit(X, y)
plot_decision_boundary(dt_clf3, axis=[-1.5, 2.5, -1.0, 1.5])
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
plt.show()
# +
# min_samples_leaf 对于一个叶子节点说,至少要几个样本才能继续拆分
dt_clf4 = DecisionTreeClassifier(min_samples_leaf=6)
dt_clf4.fit(X, y)
plot_decision_boundary(dt_clf4, axis=[-1.5, 2.5, -1.0, 1.5])
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
plt.show()
# +
# 最多有几个叶子节点 max_leaf_nodes
dt_clf5 = DecisionTreeClassifier(max_leaf_nodes=5)
dt_clf5.fit(X, y)
plot_decision_boundary(dt_clf5, axis=[-1.5, 2.5, -1.0, 1.5])
plt.scatter(X[y==0,0], X[y==0,1])
plt.scatter(X[y==1,0], X[y==1,1])
plt.show()
# -
| 5_Decision_Tree_Hyperparameters_max_depth_min_samples_leaf_min_samples_split_max_leaf_nodes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import torch
import sys
sys.path.extend(['..'])
from mvn import MVNIso, create_mog
from distributions import log_prob_banana as log_p
from util import acf, ess, estimate_log_z
from math import log, pi, sqrt, exp
import matplotlib.pyplot as plt
import time
from tqdm.notebook import trange
import numpy as np
from scipy.integrate import quad
from samplers.mcmc import HMC
from pathlib import Path
# %matplotlib inline
# +
distrib = 'banana'
component = 'MVNIso'
lam = 4.487
chain = 0
file = Path('results') / f"hmc_{distrib}_{component}_{lam:.3f}_{chain:03d}.dat"
hmc_results = torch.load(file)
# -
plt.plot(hmc_results['samples'])
full_mixture = create_mog([MVNIso(theta=th) for th in hmc_results['samples']])
x_grid = torch.linspace(-4,4,301)
xx, yy = torch.meshgrid(x_grid, x_grid)
xy = torch.stack([xx.flatten(), yy.flatten()], dim=1)
plt.contourf(xx, yy, full_mixture.log_prob(xy).exp().reshape(xx.size()))
# Estimate $MI[x;\theta]$ using the 'full' mixture
# +
def mixture_entropy(mog, n):
sample_x = mog.sample((n,))
log_m_x = mog.log_prob(sample_x)
entropy_m = -log_m_x.mean().item()
entropy_m_mcse = log_m_x.std().item() / sqrt(n)
return entropy_m, entropy_m_mcse
def mixture_kl(mog_q, mog_p, n):
sample_x = mog_q.sample((n,))
log_q_x = mog_q.log_prob(sample_x)
log_p_x = mog_p.log_prob(sample_x)
log_diff = (log_q_x - log_p_x)
return log_diff.mean().item(), log_diff.std().item() / sqrt(n)
# +
n_entropy_samples = 5000
entropy_m, entropy_m_mcse = mixture_entropy(full_mixture, n_entropy_samples)
print("H[m] is ", entropy_m, "±", entropy_m_mcse)
each_entropy_q = torch.tensor([MVNIso(theta=th).entropy() for th in hmc_results['samples']])
avg_entropy_q = each_entropy_q.mean().item()
avg_entropy_q_mcse = each_entropy_q.std().item() / sqrt(ess(each_entropy_q.view(-1,1)).item())
print("E[H[q]] is ", avg_entropy_q, "±", avg_entropy_q_mcse)
true_mi = entropy_m - avg_entropy_q
true_mi_mcse = sqrt(entropy_m_mcse**2 + avg_entropy_q_mcse**2)
print("MI[x;θ] is", true_mi, "±", true_mi_mcse)
# -
t_values = torch.arange(20, dtype=torch.int)+1
runs = 100
kl_mt_m = torch.zeros(runs, len(t_values))
kl_mt_m_mcse = torch.zeros(runs, len(t_values))
for j, t in enumerate(t_values):
for i in range(runs):
idx = torch.randint(hmc_results['samples'].size(0), size=(t,))
mixture_t = create_mog([MVNIso(theta=th) for th in hmc_results['samples'][idx, :]])
# Compute KL(m_T || m)
kl_mt_m[i,j], kl_mt_m_mcse[i,j] = mixture_kl(mixture_t, full_mixture, n_entropy_samples)
avg_kl_mt_m = kl_mt_m.mean(dim=0)
avg_kl_mt_m_mcse = (kl_mt_m.std(dim=0)**2/runs + (kl_mt_m_mcse**2).mean(dim=0)).sqrt()
plt.figure(figsize=(3,2))
plt.errorbar(t_values, y=avg_kl_mt_m, yerr=avg_kl_mt_m_mcse, color='k')
plt.errorbar(t_values, y=true_mi/t_values, yerr=true_mi_mcse/t_values, color='r')
plt.legend([r'$E[KL(m_T||m)]$', r'$\frac{1}{T}MI[x;\theta]$'])
plt.xticks(t_values[::2])
plt.xlabel('T')
plt.title('Numerical check of "variance" approximation')
plt.savefig('numerical_variance_approx.svg')
| notebooks/numerical_variance_approx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Illuminating the Latent Space of an MNIST GAN
#
# One of the most popular applications of [Generative Adversarial Networks](https://en.wikipedia.org/wiki/Generative_adversarial_network) is generating fake images. In particular, websites like [this person does not exist](https://thispersondoesnotexist.com) serve a GAN that generates fake images of people ([this x does not exist](https://thisxdoesnotexist.com) provides a comprehensive list of such websites). Such websites are entertaining, especially when one is asked to figure out [which face is real](https://www.whichfaceisreal.com/index.php).
#
# Usually, these websites extract fake images by sampling the GAN’s latent space. For those unfamiliar with GANs, this means that each image is associated with a real valued vector of $n$ components. But since these vectors are typically generated randomly, the usefulness of these websites breaks down when we wish to search for a specific image.
#
# For instance, suppose that instead of fake faces, we want to generate fake handwriting, specifically the digit eight (8). We could train a GAN on the MNIST dataset and produce a generator network that generates fake digits. Now, we can repeatedly sample the latent space until an eight appears. However, if we want to _find_ an eight, we could optimize latent space directly with CMA-ES. To ensure that we generate eights, we could use the output classification prediction of a LeNet-5 classifier as the objective (see [Bontrager 2018](https://arxiv.org/abs/1705.07386)).<sup>1</sup> But notice that the latent space likely contains many examples of the digit eight, and they might vary in the weight of the pen stroke or the lightness of the ink color. If we make these properties our behavior characteristics, we could search latent space for many different examples of eight in a single run!
#
# [Fontaine 2021](https://arxiv.org/abs/2007.05674) takes exactly this approach when generating new levels for the classic video game [Super Mario Bros](https://en.wikipedia.org/wiki/Super_Mario_Bros). They term this approach “Latent Space Illumination”, as they explore quality diversity (QD) algorithms (including [CMA-ME](https://arxiv.org/pdf/1912.02400.pdf)) as a method to search the latent space of a video game level GAN and illuminate the behavior space of possible level mechanics. In this tutorial, we illuminate the latent space of the aforementioned MNIST GAN by mimicking the approach taken in [Fontaine 2021](https://arxiv.org/abs/2007.05674).
#
# **(1)** Since the discriminator of the GAN is only trained to evaluate how realistic an image is, it cannot detect specific digits. Hence, we need the LeNet-5 to check that the digit is an 8.
# ## Setup
#
# First, we install pyribs, PyTorch, and several utilities.
# %pip install ribs torch torchvision numpy matplotlib
# +
import time
import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
import torchvision
# -
# Below, we check what device is available for PyTorch. On Colab, activate the GPU by clicking "Runtime" in the toolbar at the top. Then, click "Change Runtime Type", and select "GPU".
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
print(device)
# ## Loading the GAN and Classifier
#
# For this tutorial, we pretrained a GAN that generates MNIST digits using the code from [a beginner GAN tutorial](https://debuggercafe.com/generating-mnist-digit-images-using-vanilla-gan-with-pytorch/). We also pretrained a [LeNet-5](https://en.wikipedia.org/wiki/LeNet) classifier for the MNIST dataset using the code in our [Fooling MNIST tutorial](https://docs.pyribs.org/en/latest/tutorials/fooling_mnist.html). Below, we define the network structures.
# +
class Generator(nn.Module):
"""Generator network for the GAN."""
def __init__(self, nz):
super(Generator, self).__init__()
# Size of the latent space (number of dimensions).
self.nz = nz
self.main = nn.Sequential(
nn.Linear(self.nz, 256),
nn.LeakyReLU(0.2),
nn.Linear(256, 512),
nn.LeakyReLU(0.2),
nn.Linear(512, 1024),
nn.LeakyReLU(0.2),
nn.Linear(1024, 784),
nn.Tanh(),
)
def forward(self, x):
return self.main(x).view(-1, 1, 28, 28)
class Discriminator(nn.Module):
"""Discriminator network for the GAN."""
def __init__(self):
super(Discriminator, self).__init__()
self.n_input = 784
self.main = nn.Sequential(
nn.Linear(self.n_input, 1024),
nn.LeakyReLU(0.2),
nn.Dropout(0.3),
nn.Linear(1024, 512),
nn.LeakyReLU(0.2),
nn.Dropout(0.3),
nn.Linear(512, 256),
nn.LeakyReLU(0.2),
nn.Dropout(0.3),
nn.Linear(256, 1),
nn.Sigmoid(),
)
def forward(self, x):
x = x.view(-1, 784)
return self.main(x)
LENET5 = nn.Sequential(
nn.Conv2d(1, 6, (5, 5), stride=1, padding=0), # (1,28,28) -> (6,24,24)
nn.MaxPool2d(2), # (6,24,24) -> (6,12,12)
nn.ReLU(),
nn.Conv2d(6, 16, (5, 5), stride=1, padding=0), # (6,12,12) -> (16,8,8)
nn.MaxPool2d(2), # (16,8,8) -> (16,4,4)
nn.ReLU(),
nn.Flatten(), # (16,4,4) -> (256,)
nn.Linear(256, 120), # (256,) -> (120,)
nn.ReLU(),
nn.Linear(120, 84), # (120,) -> (84,)
nn.ReLU(),
nn.Linear(84, 10), # (84,) -> (10,)
nn.LogSoftmax(dim=1), # (10,) log probabilities
).to(device)
LENET5_MEAN_TRANSFORM = 0.1307
LENET5_STD_DEV_TRANSFORM = 0.3081
# -
# Next, we load the pretrained weights for each network.
# +
import os
from urllib.request import urlretrieve
from pathlib import Path
LOCAL_DIR = Path("lsi_mnist_weights")
LOCAL_DIR.mkdir(exist_ok=True)
WEB_DIR = "https://raw.githubusercontent.com/icaros-usc/pyribs/master/examples/tutorials/_static/"
# Download the model files to LOCAL_DIR.
for filename in [
"mnist_generator.pth",
"mnist_discriminator.pth",
"mnist_classifier.pth",
]:
model_path = LOCAL_DIR / filename
if not model_path.is_file():
urlretrieve(WEB_DIR + filename, str(model_path))
# Load the weights of each network from its file.
g_state_dict = torch.load(
str(LOCAL_DIR / "mnist_generator.pth"),
map_location=device,
)
d_state_dict = torch.load(
str(LOCAL_DIR / "mnist_discriminator.pth"),
map_location=device,
)
c_state_dict = torch.load(
str(LOCAL_DIR / "mnist_classifier.pth"),
map_location=device,
)
# Instantiate networks and insert the weights.
generator = Generator(nz=128).to(device)
discriminator = Discriminator().to(device)
generator.load_state_dict(g_state_dict)
discriminator.load_state_dict(d_state_dict)
LENET5.load_state_dict(c_state_dict)
# -
# ## LSI with CMA-ME on MNIST GAN
#
# After loading the GAN and the classifier, we can begin exploring the latent space of the GAN with the pyribs implementation of CMA-ME. Thus, we import and initialize the `GridArchive`, `ImprovementEmitter`, and `Optimizer` from pyribs.
#
# For the `GridArchive`, we choose a 2D behavior space with "boldness" and "lightness" as the behavior characteristics. We approximate "boldness" of a digit by counting the number of white pixels in the image, and we approximate "lightness" by averaging the values of the white pixels in the image. We define a "white" pixel as a pixel with value at least 0.5 (pixels are bounded to the range $[0,1]$). Since there are 784 pixels in an image, boldness is bounded to the range $[0, 784]$. Meanwhile, lightness is bounded to the range $[0.5, 1]$, as that is the range of a white pixel.
# +
from ribs.archives import GridArchive
archive = GridArchive(
[200, 200], # 200 bins in each dimension.
[(0, 784), (0.5, 1)], # Boldness range, lightness range.
)
# -
# Next, we use 5 instances of `ImprovementEmitter`, each with batch size of 30. Each emitter begins with a zero vector of the same dimensionality as the latent space and an initial step size $\sigma=0.2$.
# +
from ribs.emitters import ImprovementEmitter
emitters = [
ImprovementEmitter(
archive,
np.zeros(generator.nz),
0.2,
batch_size=30,
) for _ in range(5)
]
# -
# Finally, we construct the optimizer to connect the archive and emitters together.
# +
from ribs.optimizers import Optimizer
optimizer = Optimizer(archive, emitters)
# -
# With the components created, we now generate latent vectors. As we use 5 emitters with batch size of 30 and run 30,000 iterations, we evaluate 30,000 * 30 * 5 = 4,500,000 latent vectors in total. This loop should take 15-30 min to run.
# + tags=[]
total_itrs = 30_000
flat_img_size = 784 # 28 * 28
start_time = time.time()
for itr in range(1, total_itrs + 1):
sols = optimizer.ask()
with torch.no_grad():
tensor_sols = torch.tensor(
sols,
dtype=torch.float32,
device=device,
)
# Shape: len(sols) x 1 x 28 x 28
generated_imgs = generator(tensor_sols)
# Normalize the images from [-1,1] to [0,1].
normalized_imgs = (generated_imgs + 1.0) / 2.0
# We optimize the score of the digit being 8. Other digits may also be
# used.
lenet5_normalized = ((normalized_imgs - LENET5_MEAN_TRANSFORM) /
LENET5_STD_DEV_TRANSFORM)
objs = torch.exp(LENET5(lenet5_normalized)[:, 8]).cpu().numpy()
# Shape: len(sols) x 784
flattened_imgs = normalized_imgs.cpu().numpy().reshape(
(-1, flat_img_size))
# The first bc is the "boldness" of the digit (i.e. number of white
# pixels). We consider pixels with values larger than or equal to 0.5
# to be "white".
# Shape: len(sols) x 1
boldness = np.count_nonzero(flattened_imgs >= 0.5,
axis=1,
keepdims=True)
# The second bc is the "lightness" of the digit (i.e. the mean value of
# the white pixels).
# Shape: len(sols) x 1
flattened_imgs[flattened_imgs < 0.5] = 0 # Set non-white pixels to 0.
# Add 1 to avoid dividing by zero.
lightness = (np.sum(flattened_imgs, axis=1, keepdims=True) /
(boldness + 1))
# Each BC entry is [boldness, lightness].
bcs = np.concatenate([boldness, lightness], axis=1)
optimizer.tell(objs, bcs)
if itr % 1000 == 0:
print(
f"Iteration {itr} complete after {time.time() - start_time}s - "
f"Archive size: {len(archive.as_pandas(include_solutions=False))}")
# -
# ## Visualization
#
# Below, we visualize the archive after all evaluations. The x-axis is the boldness and the y-axis is the lightness. The color indicates the objective value. We can see that we found many images that the classifier strongly believed to be an eight.
# +
from ribs.visualize import grid_archive_heatmap
plt.figure(figsize=(8, 6))
grid_archive_heatmap(archive)
plt.title("LSI MNIST")
plt.xlabel("Boldness")
plt.ylabel("Lightness")
plt.show()
# -
# Next, we display a grid of digits generated from a selected set of latent vectors in the archive.
# + tags=[]
from torchvision.utils import make_grid
def show_grid_img(x_start,
x_num,
x_step_size,
y_start,
y_num,
y_step_size,
archive,
figsize=(8, 6)):
"""Displays a grid of images from the archive.
Args:
x_start (int): Starting index along x-axis.
x_num (int): Number of images to generate along x-axis.
x_step_size (int): Index step size along x-axis.
y_start (int): Starting index along y-axis.
y_num (int): Number of images to generate along y-axis.
y_step_size (int): Index step size along y-axis.
archive (GridArchive): Archive with results from CMA-ME.
figsize ((int, int)): Size of the figure for the image.
"""
elites = archive.as_pandas()
x_range = np.arange(x_start, x_start + x_step_size * x_num, x_step_size)
y_range = np.arange(y_start, y_start + y_step_size * y_num, y_step_size)
grid_indexes = [(x, y) for y in np.flip(y_range) for x in x_range]
imgs = []
img_size = (28, 28)
for index in grid_indexes:
x, y = index
sol_row = elites[(elites["index_0"] == x) & (elites["index_1"] == y)]
if sol_row.empty:
print(
f"Index ({x}, {y}) solution does not exist at the specified indexes."
)
return
latent_vec = sol_row.iloc[0]["solution_0":].to_numpy()
with torch.no_grad():
img = generator(
torch.tensor(latent_vec.reshape(1, generator.nz),
dtype=torch.float32,
device=device))
# Normalize images to [0,1].
normalized = (img.reshape(1, *img_size) + 1) / 2
imgs.append(normalized)
plt.figure(figsize=figsize)
img_grid = make_grid(imgs, nrow=x_num, padding=0)
plt.imshow(np.transpose(img_grid.cpu().numpy(), (1, 2, 0)),
interpolation='nearest',
cmap='gray')
# Change labels to be BC values.
plt.xlabel("Boldness")
plt.ylabel("Lightness")
x_ticklabels = [
round(archive.boundaries[0][i])
for i in [x_start + x_step_size * k for k in range(x_num + 1)]
]
y_ticklabels = [
round(archive.boundaries[1][i], 2) for i in [
y_start + y_step_size * y_num - y_step_size * k
for k in range(y_num + 1)
]
]
plt.xticks([img_size[0] * x for x in range(x_num + 1)], x_ticklabels)
plt.yticks([img_size[0] * x for x in range(y_num + 1)], y_ticklabels)
# -
# As we can see below, digits get bolder as we go along the x-axis. Meanwhile, as we go along the y-axis, the digits get brighter. For instance, the image in the bottom right corner is grey and bold, while the image in the top left corner is white and thin.
show_grid_img(10, 8, 7, 105, 6, 15, archive)
# Here we display images from a wider range of the archive. Note that in order to generate images with high boldness values, CMA-ME generated images that do not look realistic (see the bottom right corner in particular).
show_grid_img(10, 8, 15, 90, 6, 15, archive)
# To determine how realistic all of the images in the archive are, we can evaluate them with the discriminator network of the GAN. Below, we create a new archive where the objective value of each solution is the discriminator score. BCs remain the same.
# +
df = archive.as_pandas()
discriminator_archive = GridArchive(
[200, 200], # 200 bins in each dimension.
[(0, 784), (0.5, 1)], # Boldness range, lightness range.
)
discriminator_archive.initialize(generator.nz)
# Evaluate each solution in the archive and insert it into the new archive.
for _, row in df.iterrows():
latent = np.array(row.loc["solution_0":])
bcs = row.loc[["behavior_0", "behavior_1"]]
# No need to normalize to [0, 1] since the discriminator takes in images in
# the range [-1, 1].
img = generator(
torch.tensor(latent.reshape(1, generator.nz),
dtype=torch.float32,
device=device))
obj = discriminator(img).item()
discriminator_archive.add(latent, obj, bcs)
# -
# Now, we can plot a heatmap of the archive with the discriminator score. The large regions of low score (in black) show that many images in the archive are not realistic, even though LeNet-5 had high confidence that these images showed the digit eight.
plt.figure(figsize=(8, 6))
grid_archive_heatmap(discriminator_archive)
plt.title("Discriminator Evaluation")
plt.xlabel("Boldness")
plt.ylabel("Lightness")
plt.show()
# ## Conclusion
#
# By searching the latent space of an MNIST GAN, CMA-ME found images of the digit eight that varied in boldness and lightness. Even though the LeNet-5 network had high confidence that these images were eights, it turned out that many of these images were highly unrealistic --- when we evaluated them with the GAN's discriminator network, the images mostly received low scores.
#
# _In short, we found that large portions of the GAN's latent space are unrealistic_. This is not surprising because during training, the GAN generates fake images by randomly sampling the latent space from a fixed Gaussian distribution, and some portions of the distribution are less likely to be sampled. Thus, we have the following questions, which we leave open for future exploration:
#
# - How can we ensure that CMA-ME searches for realistic eights?
# - While searching for realistic eights, can we also search for other digits at the same time?
| examples/tutorials/lsi_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: feos
# language: python
# name: feos
# ---
# +
from feos_pcsaft.si import *
from feos_pcsaft.dft import *
from feos_pcsaft import PcSaftParameters
import matplotlib.pyplot as plt
import numpy as np
# -
params = PcSaftParameters.from_json(["methanol"], "adsorption_params.json")
func = PcSaftFunctional(params)
T = 350*KELVIN
state = State(func, T, pressure=BAR)
potential = ExternalPotential.LJ93(3.0, 100.0, 0.08)
#potential = ExternalPotential.Steele(3.0, 100.0, 0.08)
# +
f, ax = plt.subplots(1,2,figsize=(15,5))
pore_sizes = [5,10,20]
slit_pore = Pore1D(Geometry.Cartesian, 100*ANGSTROM, potential, 4096).initialize(state)
cylindrical_pores = [Pore1D(Geometry.Cylindrical, r*ANGSTROM, potential).initialize(state) for r in pore_sizes]
spherical_pores = [Pore1D(Geometry.Spherical, r*ANGSTROM, potential).initialize(state) for r in pore_sizes]
for cpore, spore, size in zip(cylindrical_pores, spherical_pores, pore_sizes):
ax[0].plot(cpore.r/ANGSTROM - size, cpore.external_potential.T, label=f'${size}$')
ax[1].plot(spore.r/ANGSTROM - size, spore.external_potential.T, label=f'${size}$')
for a in ax:
a.plot(slit_pore.z/ANGSTROM-50, slit_pore.external_potential.T, label='$\infty$')
a.axis([-6,0,-20,55])
a.legend()
# -
pmin = 0.2*BAR
pmax = 2.5*BAR
cyl10 = Adsorption1D.equilibrium_isotherm(func, T, (pmin, pmax, 151), Pore1D(Geometry.Cylindrical, 10*ANGSTROM, potential))
cyl20 = Adsorption1D.equilibrium_isotherm(func, T, (pmin, pmax, 151), Pore1D(Geometry.Cylindrical, 20*ANGSTROM, potential))
sph10 = Adsorption1D.equilibrium_isotherm(func, T, (pmin, pmax, 151), Pore1D(Geometry.Spherical, 10*ANGSTROM, potential))
sph20 = Adsorption1D.equilibrium_isotherm(func, T, (pmin, pmax, 151), Pore1D(Geometry.Spherical, 20*ANGSTROM, potential))
car10 = Adsorption1D.equilibrium_isotherm(func, T, (pmin, pmax, 151), Pore1D(Geometry.Cartesian, 20*ANGSTROM, potential))
car20 = Adsorption1D.equilibrium_isotherm(func, T, (pmin, pmax, 151), Pore1D(Geometry.Cartesian, 40*ANGSTROM, potential))
plt.plot(cyl10.pressure/BAR, cyl10.total_adsorption/(np.pi*100*ANGSTROM**2)/(KILO*MOL/METER**3), 'r', label='cyl10')
plt.plot(cyl20.pressure/BAR, cyl20.total_adsorption/(np.pi*400*ANGSTROM**2)/(KILO*MOL/METER**3), 'r--', label='cyl20')
plt.plot(sph10.pressure/BAR, sph10.total_adsorption/(4/3*np.pi*1000*ANGSTROM**3)/(KILO*MOL/METER**3), 'b', label='sph10')
plt.plot(sph20.pressure/BAR, sph20.total_adsorption/(4/3*np.pi*8000*ANGSTROM**3)/(KILO*MOL/METER**3), 'b--', label='sph20')
plt.plot(car10.pressure/BAR, car10.total_adsorption/(10*ANGSTROM)/(KILO*MOL/METER**3), 'g', label='car10')
plt.plot(car20.pressure/BAR, car20.total_adsorption/(20*ANGSTROM)/(KILO*MOL/METER**3), 'g--', label='car20')
plt.legend()
# +
f, ax = plt.subplots(1,2,figsize=(15,5))
ax[0].plot(cyl10.profiles[-1].r/ANGSTROM, (cyl10.profiles[-1].density/(KILO*MOL/METER**3)).T, label='cyl10')
ax[1].plot(cyl20.profiles[-1].r/ANGSTROM, (cyl20.profiles[-1].density/(KILO*MOL/METER**3)).T, label='cyl20')
ax[0].plot(sph10.profiles[-1].r/ANGSTROM, (sph10.profiles[-1].density/(KILO*MOL/METER**3)).T, label='sph10')
ax[1].plot(sph20.profiles[-1].r/ANGSTROM, (sph20.profiles[-1].density/(KILO*MOL/METER**3)).T, label='sph20')
ax[0].plot(car10.profiles[-1].r/ANGSTROM, (car10.profiles[-1].density/(KILO*MOL/METER**3)).T, label='car10')
ax[1].plot(car20.profiles[-1].r/ANGSTROM, (car20.profiles[-1].density/(KILO*MOL/METER**3)).T, label='car20')
ax[0].set_xlim(0,10)
ax[1].set_xlim(0,20)
for a in ax:
a.set_ylim(0,50)
a.legend()
# -
| examples/pore_geometry.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Lecture
class CreditCard:
"""A consumer credit card"""
def __init__(self, customer, bank, acnt, limit):
"""Create a new credit card instance.
The initial balance is zero.
:param customer: the name of the customer
:param bank: the name of the bank
:param acnt: the account identifier
:param limit: credit limit
"""
self._customer = customer
self._bank = bank
self._account = acnt
self._limit = limit
self._balance = 0
def get_customer(self):
"""Return name of the customer"""
return self._customer
def get_bank(self):
"""Return the bank's name."""
return self._bank
def get_account(self):
"""Return the card identifying number (typically stored as a string."""
return self._account
def get_limit(self):
"""Return current credit limit."""
return self._limit
def get_balance(self):
"""Return current balance."""
return self._balance
def charge(self, price):
"""Charge given price to the card, assuming sufficient credit limit.
Return True if charge was processed; False if charge was denied.
"""
if price + self._balance > self._limit:
return False
else:
self._balance += price
return True
def make_payment(self, amount):
"""Process customer payment that reduces balance."""
self._balance -= amount
# +
# Inheritance of the credit card class
class PredatoryCreditCard(CreditCard):
"""An extension to CreditCard that compounds interest and fees."""
def __init__(self, customer, bank, acnt, limit, apr):
"""Create a new predatory credit card instance.
The initial balance is zero.
customer the name of the customer
bank the name of the bank
acnt the acount identifier
limit credit limit
apr annual percentage rate
"""
super().__init__(customer, bank, acnt, limit)
self._apr = apr
def charge(self, price):
"""Charge given price to the card, assuming sufficient credit limit.
Return True if charge was processed.
Return False and assess $5 fee if charge is denied.
"""
success = super().charge(price)
if not success:
self._balance += 5
return success
def process_month(self):
"""Assess monthly interest on outstanding balance."""
if self._balance > 0:
# if positive balance, convert APR to monthly multiplicative factor
monthly_factor = pow(1 + self._apr, 1/12)
self._balance *= monthly_factor
# -
predatory = PredatoryCreditCard('John', 'AA Bank', '1234 5678', 100000000, 0.03)
predatory.charge(1000)
predatory.get_balance()
predatory.process_month()
predatory.get_balance()
# +
## Abstract Base Classes
from abc import ABCMeta, abstractmethod
class Sequence(metaclass=ABCMeta):
"""Our own version of collections.Sequence abstract base class."""
@abstractmethod
def __len__(self):
"""Return the length of the sequence."""
@abstractmethod
def __getitem__(self, item):
"""Return the element at index item of the sequence."""
def __contains__(self, val):
"""Retrun True if val found in the sequence; False otherwise."""
for j in range(len(self)):
if self[j] == val:
return True
return False
def index(self, val):
"""REturn leftmost index at which val is found (or raise ValueError)."""
for j in range(len(self)):
if self[j] == val:
return j
raise ValueError('value not in sequence')
def count(self, val):
"""Return the number of elements equal to given value."""
k = 0
for j in range(len(self)):
if self[j] == val:
k += 1
return k
# -
Sequence()
# +
#Reinforcement
# -
# R24
class Flower:
def __init__(self, name, petal_number,price):
self.n = name
self.pn = petal_number
self.p = price
# +
# R25
class CreditCard:
"""A consumer credit card"""
def __init__(self, customer, bank, acnt, limit):
"""Create a new credit card instance.
The initial balance is zero.
:param customer: the name of the customer
:param bank: the name of the bank
:param acnt: the account identifier
:param limit: credit limit
"""
self._customer = customer
self._bank = bank
self._account = acnt
self._limit = limit
self._balance = 0
def get_customer(self):
"""Return name of the customer"""
return self._customer
def get_bank(self):
"""Return the bank's name."""
return self._bank
def get_account(self):
"""Return the card identifying number (typically stored as a string."""
return self._account
def get_limit(self):
"""Return current credit limit."""
return self._limit
def get_balance(self):
"""Return current balance."""
return self._balance
def charge(self, price):
"""Charge given price to the card, assuming sufficient credit limit.
Return True if charge was processed; False if charge was denied.
"""
if price + self._balance > self._limit:
return False
else:
try:
self._balance += price
return True
except:
raise ValueError
def make_payment(self, amount):
"""Process customer payment that reduces balance."""
if amount < 0:
raise ValueError
else:
self._balance -= amount
# -
class Vector:
"""Represent a vector in a multidimensional space"""
def __init__(self, value):
"""Create d-dimensional vector of zeros."""
if isinstance(value, int):
self._coords = [0] * value
elif isinstance(value, list):
self._coords = [i for i in value]
else:
raise ValueError('Vector accepts int or list only')
def __len__(self):
"""Return the dimension of the vector."""
return len(self._coords)
def __getitem__(self, j):
"""Return jth coordinate of vector."""
return self._coords[j]
def __setitem__(self, j, val):
"""Set jth coordinate of vector to given value."""
self._coords[j] = val
def __add__(self, other):
"""Return sum of two vectors."""
if len(self) != len(other):
raise ValueError('dimensions must agree')
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] + other[j]
return result
def __radd__(self, other):
"""Return sum of two vectors."""
if len(self) != len(other):
raise ValueError('dimensions must agree')
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] + other[j]
return result
def __sub__(self, other):
"""Return subtraction of two vectors."""
if len(self)!= len(other):
raise ValueError('dimensions must agree')
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] - other[j]
return result
def __mul__(self, other):
"""Return multiplication of a scalar or a vector"""
if isinstance(other, (int, float)):
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] * other
elif len(self) == len(other):
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] * other[j]
else:
raise ValueError('multiplicand should be a scalar or a vector with same dimension as multiplier')
return result
def __rmul__(self, other):
"""Return multiplication of a scalar or a vector"""
if isinstance(other, (int, float)):
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] * other
elif len(self) == len(other):
result = Vector(len(self))
for j in range(len(self)):
result[j] = self[j] * other[j]
else:
raise ValueError('multiplicand should be a scalar or a vector with same dimension as multiplier')
return result
def __neg__(self):
result = Vector(len(self))
for j in range(len(self)):
result[j] = -self[j]
return result
def __eq__(self, other):
"""Return True if vectgor has same coordinates as other."""
return self._coords == other._coords
def __ne__(self, other):
"""Return True if vector differs from other."""
return not self == other # This rely on existing __eq__ definition
def __str__(self):
"""Produce string representation of vector."""
return '<' + str(self._coords)[1:-1] + '>'
def __repr__(self):
"""For representation."""
return 'Vector(%r)' % self._coords
| Data Structure and Algorithms/Section2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
a=pd.read_csv("/home/technosoft/Answers.csv")
a
b=pd.read_csv("/home/technosoft/Questions.csv")
b
del a
del a1
b=pd.read_csv("/home/technosoft/Questions.csv")
b1=b["Body"]
len(b)
len(b)
b11=[]
for i in b1:
i = i.replace('<p>', '').replace('</p>', '').replace('<ul>','').replace('</ul>','').replace('"','')
b11.append(i)
b11
del a
del a11
len(b11)
len(a11)
| datasetup.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# name: python385jvsc74a57bd0402135d49870b961f74d796f924ad426c34ba2a9a18d44bdd08100ee6884e7b2
# ---
# # Data Visualization
# ## Import Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from plotly.subplots import make_subplots
import plotly.graph_objects as go
from plotly.offline import download_plotlyjs, init_notebook_mode, plot, iplot
pd.set_option("display.max_columns", 120)
import plotly.offline as py
# %matplotlib inline
# ## Import Datasets
dataset = pd.read_csv('data/cleaned_train_v2.csv')
# ## Data Preparation
# Converting POSIX data from visiStartTime column and replace it in date column
dataset['date'] = pd.to_datetime(dataset['visitStartTime'], unit='s').dt.strftime('%Y-%m-%d-%H')
# +
dataset = dataset.assign(
Date = lambda x: pd.to_datetime(x['date']).dt.date,
Year = lambda x: pd.to_datetime(x['date']).dt.year,
Month = lambda x: pd.to_datetime(x['date']).dt.month,
Day = lambda x: pd.to_datetime(x['date']).dt.day,
hour = lambda x: pd.to_datetime(x['date']).dt.hour
)
print(f'Start of year: {dataset.Year.min()}')
print(f'Start of year: {dataset.Year.max()}')
dataset["year_month"] = pd.to_datetime(dataset['visitStartTime'], unit='s').dt.strftime('%Y-%m')
# -
# # Hypotheses:
# 1. Users who come from "Origanic search" are doing more transactions.
# 2. Users who visited the Store more than 3 times made also a transaction.
# 3. Most of the Sessions are from mobile users (change count to sum for amount of transactions).
# 4. Most visits are from USA.
# 5. Most revenues are generate from the USA.
# 6. Users made less transactions in February.
# 7. Sessions through Operating System MacOS tend to have more Revenue.
# ## 1. Users who come from "Origanic search" are doing more transactions
df_nonz = dataset[dataset['totals.transactionRevenue']>0].sort_values(by ="totals.transactionRevenue", ascending=False)
df_z = dataset[dataset['totals.transactionRevenue']==0].sort_values(by ="totals.transactionRevenue", ascending=False)
sns.set(font_scale=1.2)
fig, ax = plt.subplots(figsize=(15,8))
ax = sns.histplot(data=dataset, x="channelGrouping")
sns.set(font_scale=1.2)
fig, ax = plt.subplots(figsize=(15,8))
ax = sns.histplot(data=df_nonz, x="channelGrouping")
# ### Value of transactions per channel
obj = df_nonz.groupby('channelGrouping')["totals.transactionRevenue"].sum().sort_values(ascending =False).reset_index()
fig, ax = plt.subplots(figsize=(10,6))
ax = sns.barplot(data=obj, x=obj['channelGrouping'].head(10), y=obj['totals.transactionRevenue'].head(10))
ax.set(xlabel="Channel Grouping", ylabel = "Revenue in $")
# ## 2. Users who visited the Store more than 3 times made also a transaction
obj = dataset.groupby('visitNumber')["totals.transactionRevenue"].mean().sort_values(ascending =False).reset_index()
fig, ax = plt.subplots(figsize=(15,8))
ax = sns.barplot(data=obj, x=obj['visitNumber'].head(10), y=obj['totals.transactionRevenue'].head(10))
# ## 3. Most of the Sessions are from mobile users
# (change count to sum for amount of transactions)
obj = dataset.groupby(['channelGrouping','device.isMobile'])["totals.transactionRevenue"].count().sort_values(ascending =False).reset_index()
ax = sns.catplot(y="totals.transactionRevenue",x="channelGrouping",kind="bar", hue="device.isMobile",height=8.27, aspect=11.7/8.27,
edgecolor=".6", data=obj)
ax.set(xlabel="Channel Grouping", ylabel = "No. of Sessions")
# ## 4. Most visits are from USA
tmp = dataset["geoNetwork.country"].value_counts()
country_visits = pd.DataFrame(data={'geoNetwork.country': tmp.values}, index=tmp.index).reset_index()
country_visits.columns = ['Country', 'Visits']
def plot_country_map(data, location, z, legend, title, colormap='Viridis'):
data = dict(type = 'choropleth',
colorscale = colormap,
autocolorscale = False,
reversescale = False,
locations = data[location],
locationmode = 'country names',
z = data[z],
text = data[z],
colorbar = {'title':legend})
layout = dict(title = title,
geo = dict(showframe = False,
projection = {'type': 'natural earth'}))
choromap = go.Figure(data = [data], layout=layout)
iplot(choromap)
plot_country_map(country_visits, 'Country', 'Visits', 'Visits', 'Visits per country')
# ## 5. Most revenues are generate from the USA
# Select the visits with non-zero transaction revenue and calculate the sums
tmp = df_nonz.groupby(['geoNetwork.country'])['totals.transactionRevenue'].sum()
country_total = pd.DataFrame(data={'total': tmp.values}, index=tmp.index).reset_index()
country_total.columns = ['Country', 'Total']
country_total['Total'] = np.log1p(country_total['Total'])
plot_country_map(country_total, 'Country', 'Total', 'Total(log)', 'Total revenues per country (log scale)')
# ## 6. Users made less transactions in February
tmp = dataset.groupby('Date')['totals.transactionRevenue'].agg(['size'])
tmp.columns = ["Total"]
tmp = tmp.sort_index()
# +
def plot_scatter_data(data, xtitle, ytitle, title, color='blue'):
trace = go.Scatter(
x = data.index,
y = data.values,
name=ytitle,
marker=dict(
color=color
),
mode='lines'
)
data = [trace]
layout = dict(title = title,
xaxis = dict(title = xtitle), yaxis = dict(title = ytitle),
)
fig = dict(data=data, layout=layout)
iplot(fig, filename='lines')
# -
plot_scatter_data(tmp['Total'],'Date','No. of sessions','Sessions including zero transactions','green')
tmp_nonz = df_nonz.groupby('Date')['totals.transactionRevenue'].agg(['size'])
tmp_nonz.columns = ["Total"]
tmp_nonz = tmp_nonz.sort_index()
plot_scatter_data(tmp_nonz['Total'],'Date','No. of sessions','Sessions with revenue only','red')
# ### Predicted Transactions
dataset_pred = pd.read_csv('models/dataset_pred.csv')
tmp0 = dataset.groupby('Date')['totals.transactionRevenue'].agg(['sum'])
tmp0.columns = ["Total"]
tmp0 = tmp0.sort_index()
plot_scatter_data(tmp0['Total'],'Date', 'Amountin $','Total Revenue for year 2016-2018','orange')
def plot_scatter_data_join(data, data2, xtitle, ytitle, title):
trace = go.Scatter(
x = data.index,
y = data.values,
name=ytitle,
marker=dict(
color='maroon'
),
mode='lines'
)
trace2 = go.Scatter(
x = data2.index,
y = data2.values,
name='Actual Rev. in $',
marker=dict(
color='rgb(94,163,192)'
),
mode='lines'
)
# data = [trace, trace2]
layout = dict(title = title,
xaxis = dict(title = xtitle), yaxis = dict(title = ytitle)
)
#fig = dict(data=data, layout=layout)
fig = make_subplots(specs=[[{"secondary_y": True}]])
fig.add_trace(trace)
fig.add_trace(trace2,secondary_y=True)
fig['layout'].update(height = 400, width = 1150, title = title,xaxis=dict(
tickangle=0
))
iplot(fig, filename='lines')
tmp1 = dataset_pred.groupby('date')['Label'].agg(['sum'])
tmp1.columns = ["Label"]
tmp1 = tmp1.sort_index()
tmp2 = dataset_pred.groupby('date')['Target'].agg(['sum'])
tmp2.columns = ["Target"]
tmp2 = tmp2.sort_index()
plot_scatter_data_join(tmp1['Label'],tmp2['Target'],'date', 'Predicted Rev. in $','Result: Revenue generating sessions from May - Oct 2018')
### Unlog
dataset_pred['Label'] = np.expm1(dataset_pred['Label'])
tmp1 = dataset_pred.groupby('date')['Label'].agg(['sum'])
tmp1.columns = ["Label"]
tmp1 = tmp1.sort_index()
plot_scatter_data_join(tmp1['Label'],tmp0['Total'],'date', 'Predicted Rev. in $','Revenue generating sessions from Aug 2016 - Oct 2018')
# ### Total hits binned (Change to visitnumber instead of hits)
bins = [0, 50, 100, 150, 200, 250, 300, 350, 400]
labels = ["0-50","51-100","101-150","151-200","201-250","251-300","301-350","351-400"]
dataset['binned'] = pd.cut(dataset['visitNumber'], bins=bins, labels=labels)
obj = dataset.groupby('binned')["totals.transactionRevenue"].mean().sort_values(ascending =False).reset_index()
#obj.head(20)
fig, ax = plt.subplots(figsize=(10,6))
ax = sns.barplot(data=obj, x=obj['binned'].head(20), y=obj['totals.transactionRevenue'])
ax.set(xlabel="Visits grouped", ylabel = "Revenue in $")
gdf = dataset.groupby("fullVisitorId")["totals.transactionRevenue"].sum().reset_index()
nrc = gdf[gdf['totals.transactionRevenue']==0]
rc = gdf[gdf['totals.transactionRevenue']>0]
print("The number of nonrevenue customers are ", len(nrc))
print("The number of revenue generating customers are ", len(rc))
print("the ratio of revenue generating customers are {0:0.4}%".format(len(rc)/len(gdf)*100))
labels = ['Non revenue generating customers','revenue generating customers']
values = [1307589,16141]
plt.axis("equal")
plt.pie(values, labels=labels, radius=1.5, autopct="%0.2f%%",shadow=True, explode=[0,0.8], colors=['lightskyblue','lightcoral'])
plt.show()
# ## Country Distribution
# +
country_series = dataset["geoNetwork.country"].value_counts().head(25)
country_count = country_series.shape[0]
print("Total No. Of Countries: ", country_count)
country_series = country_series.head(25)
trace = go.Bar(
x=country_series.index,
y=country_series.values,
marker=dict(
color=country_series.values,
showscale=True
),
)
layout = go.Layout(title="Countrywise Observation Count")
data = [trace]
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename="country")
# -
# - 50% of the observerations are registered from Americas
# - 360K Observations are from USA alone.
# - Note, China is not there. Google is baned in China
# - Following USA, its India, Is it because of the population
# ## Plotly/Dash
from plotly import tools
import plotly.offline as py
py.init_notebook_mode(connected=True)
import plotly.graph_objs as go
import plotly.figure_factory as ff
# ### Sessions from revenue sessions and non-revenue sessions wrt Time
# +
trace = [
go.Histogram(x=dataset['hour'],
opacity = 0.7,
name="Total Sessions",
hoverinfo="y",
marker=dict(line=dict(width=1.6),
color='grey')
),
go.Histogram(x=df_nonz[df_nonz['totals.transactionRevenue'].notnull()]['hour'],
visible=False,
opacity = 0.7,
name = "Non-zero revenue Sessions",
hoverinfo="y",
marker=dict(line=dict(width=1.6),
color='green')
),
go.Histogram(x=df_z[df_z['totals.transactionRevenue'].notnull()]['hour'],
visible=False,
opacity = 0.7,
name = "Zero revenue Sessions",
hoverinfo="y",
marker=dict(line=dict(width=1.6),
color='orange')
)
]
layout = go.Layout(title='Sessioning hours',
paper_bgcolor = 'rgb(240, 240, 240)',
plot_bgcolor = 'rgb(240, 240, 240)',
autosize=True, xaxis=dict(tickmode="linear", title="Hour of the Day for the year 2017-2018"),
yaxis=dict(title="No. of Sessions",
titlefont=dict(size=17)),
)
updatemenus = list([
dict(
buttons=list([
dict(
args = [{'visible': [True, False, False]}],
label="Total Sessions",
method='update',
),
dict(
args = [{'visible': [False, True, False]}],
label="Non-zero revenue Sessions",
method='update',
),
dict(
args = [{'visible': [False, False, True]}],
label="Zero revenue Sessions",
method='update',
),
]),
direction="down",
pad = {'r':10, "t":10},
x=0.1,
y=1.25,
yanchor='top',
)
])
layout['updatemenus'] = updatemenus
fig = dict(data=trace, layout=layout)
fig = py.iplot(fig)
fig
# -
# ## 7. Sessions through Operating System MacOS tend to have more Revenue.
# +
df_nonz['day_frame'] = 0
df_nonz['day_frame'] = np.where((df_nonz["hour"]>=0) & (df_nonz["hour"]<4), 'overnight',
df_nonz['day_frame'])
df_nonz['day_frame'] = np.where((df_nonz["hour"]>=4) & (df_nonz["hour"]<8), 'dawn',
df_nonz['day_frame'])
df_nonz['day_frame'] = np.where((df_nonz["hour"]>=8) & (df_nonz["hour"]<12), 'morning',
df_nonz['day_frame'])
df_nonz['day_frame'] = np.where((df_nonz["hour"]>=12) & (df_nonz["hour"]<14), 'lunch',
df_nonz['day_frame'])
df_nonz['day_frame'] = np.where((df_nonz["hour"]>=14) & (df_nonz["hour"]<18), 'afternoon',
df_nonz['day_frame'])
df_nonz['day_frame'] = np.where((df_nonz["hour"]>=18) & (df_nonz["hour"]<21), 'evening',
df_nonz['day_frame'])
df_nonz['day_frame'] = np.where((df_nonz["hour"]>=21) & (df_nonz["hour"]<24), 'night',
df_nonz['day_frame'])
# +
fv = df_nonz.pivot_table(index="device.operatingSystem",columns="day_frame",
values="totals.transactionRevenue",aggfunc=lambda x:x.sum())
fv = fv[['morning', 'lunch', 'afternoon', 'evening','night','overnight', 'dawn']]
fv = fv.sort_values(by='morning', ascending=False)[:6]
trace = go.Heatmap(z=[fv.values[0],fv.values[1],fv.values[2],fv.values[3],
fv.values[4],fv.values[5]],
x=['morning', 'lunch', 'afternoon', 'evening', 'night',
'overnight','dawn'],
y=fv.index.values, colorscale='Purples', reversescale = False
)
data=[trace]
layout = go.Layout(
title='Total Revenue by Device OS<br>(parts of the day)')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig)
# -
# ## Total sessions, non-zero revenue count and Revenue counts from operating systems
# +
color = ['tomato', 'bisque','lightgreen', 'gold', 'tan', 'lightgrey', 'cyan']
def PieChart(column, title, limit):
revenue = "totals.transactionRevenue"
count_trace = dataset.groupby(column)[revenue].size().nlargest(limit).reset_index()
non_zero_trace = df_nonz.groupby(column)[revenue].count().nlargest(limit).reset_index()
rev_trace = df_nonz.groupby(column)[revenue].sum().nlargest(limit).reset_index()
trace1 = go.Pie(labels=count_trace[column],
values=count_trace[revenue],
name= "Sessions",
hole= .5, textfont=dict(size=10),
domain= {'x': [0, .32]},
marker=dict(colors=color))
trace2 = go.Pie(labels=non_zero_trace[column],
values=non_zero_trace[revenue],
name="Revenue",
hole= .5, textfont=dict(size=10),
domain= {'x': [.34, .66]})
trace3 = go.Pie(labels=rev_trace[column],
values=rev_trace[revenue],
name="Revenue",
hole= .5, textfont=dict(size=10),
domain= {'x': [.68, 1]})
layout = dict(title= title, font=dict(size=15), legend=dict(orientation="h"),
annotations = [
dict(
x=.10, y=.5,
text='<b>Total <br>Sessions',
showarrow=False,
font=dict(size=12)
),
dict(
x=.50, y=.5,
text='<b>Non-zero <br>Sessions<br>',
showarrow=False,
font=dict(size=12)
),
dict(
x=.88, y=.5,
text='<b>Total<br>Revenue',
showarrow=False,
font=dict(size=12)
)
])
fig = dict(data=[trace1, trace2,trace3], layout=layout)
py.iplot(fig)
# -
PieChart("device.operatingSystem", "Operating System", 4)
| 7_data_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# disclaimer: To ensure that the notebook can be run from (more or less) any point, I try to load the relevant functions or modules whenever I use them in a cell. This is generally not good practice as it adds unneccesary overhead
# # 0. Image representation as numerical arrays
# ### We start by importing numpy and creating and printing a simple 9x9 checkerboard array
#
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload
import numpy as np
#make a 9x9 checkerboard
checkBoard = #
print(checkBoard)
# -
# ### Then, we import [pyplot](https://matplotlib.org/api/pyplot_api.html) and [image](https://matplotlib.org/api/image_api.html) modules from the ploting library [matplotlib](https://matplotlib.org/3.1.1/api/index.html). Using it, we can display our checkerboard array in image form:
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
plt.imshow(checkBoard, cmap='gray', interpolation='nearest')
plt.show()
# ### As another simple example, we will import the [data](https://scikit-image.org/docs/dev/api/skimage.data.html) module image processing library [scikit-image](https://scikit-image.org/) and load a small image of a bush.
#
# #### First, we want to print the pixel values:
# +
from skimage import data
image_of_a_bush = data.lfw_subset()
image_of_a_bush = image_of_a_bush[0,:,:]
#print the #of dimentions, the shape, and the pixel values of the image
print("The number of dimensions of the image is: ", image_of_a_bush.ndim)
print("The size of the image is: ", image_of_a_bush.shape)
print(image_of_a_bush)
# -
# ### Can you see the bush?
#
# #### Next, show the image:
# +
plt.figure(figsize=(1,1))
# display the image
plt.#
# -
# # 1. Pixel-level operations
# ### Now that we have a sense of what a digital image is, let's start manipulating it. We'll begin with simple pixel-level operations
#
# ## 1.1 Basic pixel-level operations
# ### Let's look at a more interesting image. From scikit-image data we'll open a example IHC image, and plot it using pyplot.
# +
from skimage import data
import matplotlib.pyplot as plt
import numpy as np
image_hist = data.immunohistochemistry()
#check the size of the image
print("The number of dimensions of the image is: ", image_hist.ndim)
print("The size of the image is: ", image_hist.shape)
plt.imshow(image_hist, cmap=plt.cm.gray)
# -
# ### Seems like we have an RGB image. Let's look at every channel independently.
# +
plt.figure(figsize=(15,5))
plt.subplot(131)
plt.gca().set_title('Red channel')
plt.imshow(, cmap='Reds', interpolation='nearest')
plt.subplot(132)
plt.gca().set_title('Green channel')
plt.imshow(, cmap='Greens', interpolation='nearest')
plt.subplot(133)
plt.gca().set_title('Blue channel')
plt.imshow(, cmap='Blues', interpolation='nearest')
plt.show()
# -
#for the moment let's look at only the first color channel
image_hist = image_hist[:,:,0]
plt.gca().set_title('First channel')
plt.imshow(image_hist, cmap=plt.cm.gray)
# ### We can invert the image using the *invert* function from [scikit-images utilities module](https://scikit-image.org/docs/dev/api/skimage.util.html):
# +
from skimage.util import invert
inverted_image = #
plt.figure(figsize=(15,5))
plt.subplot(121)
plt.gca().set_title('original image')
plt.imshow(image_hist, cmap=plt.cm.gray)
plt.subplot(122)
plt.gca().set_title('inverted image')
plt.imshow(inverted_image, cmap=plt.cm.gray)
# -
# ### Let's try some other pixel-level operations. We'll use the [Exposure module](https://scikit-image.org/docs/dev/api/skimage.exposure.html#skimage.exposure.adjust_sigmoid) from scikit image.
#
# 1. A gamma correction applies the nonlinear transform $V_{out} = V_{in}^\gamma$.
#
# 2. A log transform applies $V_{out} = log(V_{in}+1)$.
#
# 3. A sigmoid transform applies $V_{out} = \frac{1}{1+e^{gain\cdot(\text{cutoff}-V_{in})}}$.
#
# 4. Equalization transforms the intensity histogram of an image to a uniform distribution. It often enhances the contrast of the image
#
# 5. Contrast Limited Adaptive Histogram Equalization (CLAHE) works similarly to equalization thats applied separately to different regions of the image.
#
# Try to apply these by calling the relevant function from skimage.exposure, or by direct calculation.
#
# Play with the different parameters and see how they change the output.
# +
from skimage import exposure
# apply gamma scaling with gamma=2
gamma=
gamma_corrected =
# apply logarithmic scaling
logarithmic_corrected =
# apply sigmoidal scaling with cutoff=0.4
cutoff =
sigmoid_corrected =
# equalize
equalize_corrected =
# apply Contrast Limited Adaptive Histogram Equalization (CLAHE)
CLHA_corrected =
plt.figure(figsize=(15,10))
plt.subplot(231)
plt.gca().set_title('original')
plt.imshow(image_hist, cmap=plt.cm.gray)
plt.subplot(232)
plt.gca().set_title('gamma corrected')
plt.imshow(gamma_corrected, cmap=plt.cm.gray)
plt.subplot(233)
plt.gca().set_title('log corrected')
plt.imshow(logarithmic_corrected, cmap=plt.cm.gray)
plt.subplot(234)
plt.gca().set_title('sigmoid')
plt.imshow(sigmoid_corrected, cmap=plt.cm.gray)
plt.subplot(235)
plt.gca().set_title('equalized')
plt.imshow(equalize_corrected, cmap=plt.cm.gray)
plt.subplot(236)
plt.gca().set_title('CLHA corrected')
plt.imshow(CLHA_corrected, cmap=plt.cm.gray)
# -
# ## 1.2 Image filtering
#
# ### Spatial filtering is an image processing technique for changing the intensities of a pixel according to the intensities of some neighborhood of pixels.
# <img src="./images/same_padding_no_strides.gif" width="400" height="200" >
#
#
# ### The *Kernel* of the filter defines the neighborhood and the weights asigned to each pixel in the neighborhood:
#
# <img src="./images/spatialFilter.jpg" width="400" height="200" >
# This procedure is formally a convolution and is marked by an asterisk: $I_o = I_i\ast f$.
#
# *side note: since a convolution in the spatial domain is equivalent to multiplication in the frequency domain. Sometimes it is more computationally reasonable to calculate these in fourier space.*
#
# *side side note: filtering can also be performed in the frequency domain by directly removing a set of frequencies from an image.*
#
#
# ### The kernel can be of any shape/size, it is applied to each pixel in the image, and the output is a new, filtered, image. The output image is often called the *response* to the given filter.
# Example, local average: <img src="./images/spatialFilterExample.jpg" width="300" height="150">
#
#
# #### Filtering is an incredibly versatile tool with which you can emphasize certain features or remove other features.
# #### Image processing operations implemented with filtering include smoothing, sharpening, and edge enhancement.
#
#
# ### To implement different image filters, we will use the [filters module from scikit-image](https://scikit-image.org/docs/dev/api/skimage.filters.html)
# ### 1.2.1 Smoothing
#
# #### Smoothing, aka low-pass filtering, is used for removing high-frequency noise from images. Most commonly, a gaussian kernel is used, but others (e.g. local mean/median) work too. We'll see the effect of gaussian filtering.
#
# Try to change the value of sigma (width of the gaussian) to see how the output changes.
# +
import matplotlib.pyplot as plt
import numpy as np
from skimage import filters
image_hist = data.immunohistochemistry()
sigma = 2
gauss_filtered_img =
plt.figure(figsize=(15,8))
plt.subplot(121)
plt.gca().set_title('original image')
plt.imshow(image_hist, cmap=plt.cm.gray)
plt.subplot(122)
plt.gca().set_title('response, gaussian smoothing')
plt.imshow(gauss_filtered_img, cmap=plt.cm.gray)
# -
# ### 1.2.2 Sharpening
#
# #### sharpening is sometimes used to enhance a blurry (i.e. crappy) image.
#
#
# 1. Start with input image
# 2. Apply gaussian filter with very narrow kernel
# 3. Subtract filtered image from input image to get only high frequency components
# 3. Amplify (alpha) and add high frequency components to original input image
#
# +
filter_blurred_f = filters.gaussian(gauss_filtered_img, sigma=0.5, multichannel=False)
alpha = 3
sharpened = gauss_filtered_img + alpha * (gauss_filtered_img - filter_blurred_f)
plt.figure(figsize=(15,8))
plt.subplot(121)
plt.gca().set_title('input - blury image')
plt.imshow(gauss_filtered_img, cmap=plt.cm.gray)
plt.subplot(122)
plt.gca().set_title('sharpened')
plt.imshow(sharpened, cmap=plt.cm.gray)
# -
# ### 1.2.3 Edge enhancement
#
# #### Edge detecting filters work by measuring the local spatial gradient of an image. Common types are the Sobel, Prewitt and Roberts.
#
# #### The filters are usually applied to each direction individually and then the total magnitude of the gradient is calculated.
# $G = \sqrt{G_x^2+G_y^2}$
#
# #### Sobel:
# <img src="./images/sobmasks.gif" width="200" height="100" align="left">
#
# <br><br><br><br>
#
# #### Prewitt:
# <img src="./images/premasks.png" width="200" height="100" align="left">
#
# <br><br><br><br>
#
# #### Roberts:
# <img src="./images/robmasks.gif" width="200" height="100" align="left">
#
#
# +
# sobel magnitude
filtered_img =
# sobel horizontal
filtered_img_h =
# sobel vertical
filtered_img_v =
plt.figure(figsize=(15,16))
plt.subplot(221)
plt.gca().set_title('input image')
plt.imshow(image_hist[:,:,2], cmap=plt.cm.gray)
plt.subplot(222)
plt.gca().set_title('sobel filter response - magnitude')
plt.imshow(filtered_img, cmap=plt.cm.gray)
plt.subplot(223)
plt.gca().set_title('sobel filter response - horizontal edges')
plt.imshow(np.abs(filtered_img_h), cmap=plt.cm.gray)
plt.subplot(224)
plt.gca().set_title('sobel filter response - vertical edges')
plt.imshow(np.abs(filtered_img_v), cmap=plt.cm.gray)
# -
# ## 1.3 Masking
#
# ### A mask is a binary image (0s and 1s) that typically separates a given input image into Foreground (interesting) and Background (boring) regions, or for picking a region-of-interest (ROI). A mask is *applied* to an image by element-wise multiplication. The size of a mask must be *identical* to the size of the image it's applied to.
#
#
# ### Let's begin by creating a simple circular mask. We'll create an array where the value at each point is it's distance from the center of the image, and display it as an image:
# +
import matplotlib.pyplot as plt
import numpy as np
#dimensions in x and y
y = 512
x = 512
#position of center
centY = np.ceil(y/2)
centX = np.ceil(x/2)
#create the grid
yy,xx = np.indices((y,x))
#create radial distance map
radialDist =
#display
plt.gca().set_title('Radial distance')
plt.imshow(radialDist, cmap='gray', interpolation='nearest')
plt.show()
# -
# ### Of these points, we'll pick a circle of radius 100 and display it as an image:
# +
circ1 =
plt.show()
plt.gca().set_title('Circle with radius 100')
plt.imshow(circ1, cmap='inferno', interpolation='nearest')
# -
# #### This object is a **mask**. If you multiply this matrix of 0s and 1s with an image of the same size, only the parts that are ==1 will remain
# ### Let's apply this mask to our histology image.
plt.gca().set_title('Masked first channel')
plt.imshow(, cmap=plt.cm.gray)
# ### What happens if we invert the mask?
# +
inverted_mask =
plt.figure(figsize=(15,5))
plt.subplot(121)
plt.gca().set_title('inverted mask')
plt.imshow(inverted_mask, cmap=plt.cm.gray)
plt.subplot(122)
plt.gca().set_title('inverted masked image')
plt.imshow(image_hist[:,:,2]*, cmap=plt.cm.gray)
# -
# **Just for closure, let's see what happens when we look at the full RGB image and try to apply the mask**
# +
image = data.immunohistochemistry()
masked_image = image*circ1
plt.imshow(masked_image, cmap=plt.cm.gray)
# -
# **Whoops. Seems like something is wrong. Our problem is that numpy didn't know how to multiply a 512x512x3 with a 512x512 mask. Numpy makes solving this very easy by adding a singleton dimension (look up broadcasting in your spare time).**
image = data.immunohistochemistry()
plt.gca().set_title('Masked image')
masked_image = mage*np.expand_dims(circ1,2)
plt.imshow(masked_image, cmap=plt.cm.gray)
# ## 1.4 Thresholding
# ## 1.4.1 Simple thresholding
# ### Thresholding an image is the process of setting an intensity (or intensities) for separating the different components of an image.
#
# <img src="./images/Thresholding.png" width="600" height="600" >
#
#
# #### In simplest case, the foreground and background have very different intensities. In that case thresholding is just clustering pixels by their intensity levels.
#
# +
#this function from skimage converts images of integer types into floats, which are easier to work with.
import matplotlib.pyplot as plt
import numpy as np
from skimage import img_as_float
from skimage import data
# First, let's create a noisy image of blobs
image_blobs = img_as_float(data.binary_blobs(length=512, seed=1))
sigma = 0.22
image_blobs += np.random.normal(loc=0, scale=sigma, size=image_blobs.shape)
print("The number of dimensions of the image is: ", image_blobs.ndim)
print("The size of the image is: ", image_blobs.shape)
plt.imshow(image_blobs, cmap=plt.cm.gray)
# -
# ### To find the right threshold, let's examine a histogram of pixel intensity values
plt.hist(image_blobs.flatten(),bins=250)
plt.show()
# Pick an appropriate threshold, by eye, and see if you can remove the background.
# What happens when you increase or decrease the threshold?
# +
thresh =
mask =
masked_image =
plt.figure(figsize=(15,5))
plt.subplot(131)
plt.gca().set_title('original')
plt.imshow(image_blobs, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(132)
plt.gca().set_title('mask')
plt.imshow(mask, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(133)
plt.gca().set_title('masked image')
plt.imshow(masked_image, interpolation='nearest', cmap=plt.cm.gray)
# -
# ### Our mask looks ok, but it has a lot of salt & pepper speckle noise. Why is that?
# We can try and use what we learned before about filtering to clean up our results. What filter should we use?
# +
from skimage import filters
thresh = 0.5
mask =
masked_image =
plt.figure(figsize=(15,5))
plt.subplot(131)
plt.gca().set_title('original')
plt.imshow(image_blobs, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(132)
plt.gca().set_title('mask')
plt.imshow(mask, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(133)
plt.gca().set_title('masked image')
plt.imshow(masked_image, interpolation='nearest', cmap=plt.cm.gray)
# -
# It's usually a good idea before creating a mask to despeckle an image using a narrow gaussian filter!
# ## 1.4.2 Morphological operations
# Morphology is a broad set of image processing operations that process images based on shapes. In a morphological operation, each pixel in the image is adjusted based on the value of other pixels in its neighborhood. By choosing the size and shape of the neighborhood, you can construct a morphological operation that is sensitive to specific shapes in the input image. (explanation from Mathworks)
#
# Morphological operations are based around a *structuring element*, which is a small binary image, often of a disk or a square. The structuring element is positioned at all possible locations in the image and it is compared with the corresponding neighbourhood of pixels. Some operations test whether the element "fits" within the neighbourhood, while others test whether it "hits" or intersects the neighbourhood.
#
# Common operations for image processing
#
# Erosion - output image =1 wherever the structuring element **fits** (erodes the mask)
#
# Dilation - output image =1 wherever the structuring element **hits** (expands the mask)
#
# Opening - Erosion followed by dilation (opens gaps in spots where the mask is weakly connected)
#
# Closing - Dilation followed by erosion (closes holes in the mask)
#
#
# A very thorough explanation of morphological operationscould be found [here](https://www.cs.auckland.ac.nz/courses/compsci773s1c/lectures/ImageProcessing-html/topic4.htm)
# +
from skimage.morphology import erosion, dilation, opening, closing
from skimage.morphology import disk
#define a "disk" structuring element with radius 10
selem =
#apply erosion, dilation, opening, and closing
erosion_mask =
dilation_mask =
opening_mask =
closing_mask =
plt.figure(figsize=(15,10))
plt.subplot(231)
plt.gca().set_title('mask')
plt.imshow(mask, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(232)
plt.gca().set_title('erosion')
plt.imshow(erosion_mask, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(233)
plt.gca().set_title('dilation')
plt.imshow(dilation_mask, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(235)
plt.gca().set_title('opening')
plt.imshow(opening_mask, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(236)
plt.gca().set_title('closing')
plt.imshow(closing_mask, interpolation='nearest', cmap=plt.cm.gray)
# -
# ## 1.4.3 Masking actual data
#
# ### We'll repeat the thresholding procedure using an actual microscopy image of fluorescent nuclei
# +
import matplotlib.pyplot as plt
import numpy as np
from skimage import img_as_float
import matplotlib.image as mpimg
#this is how we load an image from the hard drive
image_nuclei = img_as_float(mpimg.imread("../Data/xy040-1.png"))
fig = plt.figure(num=None, figsize=(7.1, 4.6), dpi=80, facecolor='w', edgecolor='k')
print("The number of dimensions of the image is: ", image_nuclei.ndim)
print("The size of the image is: ", image_nuclei.shape)
plt.imshow(image_nuclei, cmap=plt.cm.gray, vmin=0, vmax=0.01)
# -
# ### Again, let's plot a histogram of intensity values
plt.
plt.xlim((0, 0.02))
plt.show()
# ### And again we'll pick a value by eye:
# +
thresh =
#remember to despeckle before creating a mask!
mask =
plt.figure(figsize=(8,15))
plt.subplot(311)
plt.gca().set_title('original')
plt.imshow(image_nuclei, interpolation='nearest', cmap=plt.cm.gray, vmin=0, vmax=0.01)
plt.subplot(312)
plt.gca().set_title('mask')
plt.imshow(mask, interpolation='nearest', cmap=plt.cm.gray)
plt.subplot(313)
plt.gca().set_title('masked image')
plt.imshow(image_nuclei*mask, interpolation='nearest', cmap=plt.cm.gray, vmin=0, vmax=0.01)
# -
# ### Not bad! But also not very scalable. If we have 100s of images we can't look at them one-by-one and find thresholds by eye. Next, we'll look at some methods for automatically finding the thresholds.
# ## 1.5 Automated threshold calculation
#
# ### There is a very large list of algorithms for threshold calculation out there that are optimized for different situations. We will briefly review a few of the most common ones.
#
# ### 1.5.1 Iterative mean thresholding
# Algorithm:
# 1. Start with some threshold $T_i$
# 2. Compute the means $m_0$ and $m_1$ of the BG and FG
# 3. Update $T_{i+1} = \frac{m_0+m_1}{2}$
# 4. Repeat until it converges
#
# <img src="./images/meanThresh.gif" width="400" height="400" >
#
# ### 1.5.2 Otsu thresholding
# The algorithm exhaustively searches for the threshold that minimizes the intra-class variance, defined for a given threshold $T$ as a weighted sum of variances of the two classes:
# $\sigma^2_w(T)=\omega_0(T)\sigma^2_0(T)+\omega_1(T)\sigma^2_1(T)$
#
# For 2 classes, minimizing the intra-class variance is equivalent to maximizing inter-class variance, which is much easier to calculate:
# \begin{align}
# \sigma^2_b(T) & =\sigma^2-\sigma^2_w(T)=\omega_0(\mu_0-\mu_T)^2+\omega_1(\mu_1-\mu_T)^2 \\
# & =\omega_0(T) \omega_1(T) \left[\mu_0(T)-\mu_1(T)\right]^2
# \end{align}
#
# <img src="./images/Otsu's_Method_Visualization.gif" width="400" height="400" >
#
# ### 1.5.3 Triangle thresholding
# Algorithm:
# 1. Draw a straight line between the histogram peak and the brightest value.
# 2. From every point on that line, draw the shortest connecting line to the histogram.
# 3. Find longest of these connecting lines.
# 4. Threshold is set at the intersection of that line and the curve.
# <img src="./images/triThresh.png" width="400" height="400" >
#
# *note: Triangle thresholding is good for situations where the image is mostly background, and there is no clear "peak" of bright pixels.*
#
#
# ### [scikit-image's filters module](https://scikit-image.org/docs/dev/api/skimage.filters.html) implements a large variety of thresholding algorithms. Let's apply the ones we just learned about.
#
# +
from skimage import filters
#calculate iterative mean threshold
meanThresh =
print(meanThresh)
#calculate otsu threshold
OtsuThresh =
print(OtsuThresh)
#calculate triangle
TriThresh =
print(TriThresh)
# -
# ### Let's look at the resulting masks we get with each of these thresholds
# +
fig = plt.figure(num=None, figsize=(12, 8), dpi=80)
ax1 = fig.add_axes([0.1,0.6,0.4,0.4])
ax1.hist(image_nuclei.flatten(),bins=250)
ax1.axvline(meanThresh, color='g', linestyle='--')
ax1.axvline(OtsuThresh, color='r', linestyle='--')
ax1.axvline(TriThresh, color='k', linestyle='--')
ax1.legend(['mean' ,'otsu', 'triangle'])
ax1.set_title('histogram')
ax2 = fig.add_axes([0.6,0.6,0.4,0.4])
#get iterative mean mask (remember to despeckle)
mask_mean =
ax2.imshow(mask_mean)
ax2.set_title('Iterative mean')
ax2.set_axis_off()
ax2 = fig.add_axes([0.1,0.1,0.4,0.4])
#get otsu mask
mask_otsu =
ax2.imshow(mask_otsu)
ax2.set_title('Otsu')
ax2.set_axis_off()
ax2 = fig.add_axes([0.6,0.1,0.4,0.4])
#get triangle mask
mask_tri =
ax2.imshow(mask_tri)
ax2.set_title('Triangle')
ax2.set_axis_off()
# -
# ### 1.5.4 Local thresholding
# #### All of the methods we saw so far are *global* in the sense that the same threshold is applied to the whole picture. Sometimes we can have an image with vastly different intensity distributions at different locations. Using local thresholding, we can overcome such cases.
#
# Let's compare the results from a global (Otsu) and a local threshold.
# +
from skimage import data
image = data.page()
fig = plt.figure(num=None, figsize=(12, 8), dpi=80)
#global thresholding
threshGlobal = filters.threshold_otsu(image)
ax1 = fig.add_axes([0.1,0.6,0.4,0.4])
ax1.set_title('mask - Otsu threshold')
plt.imshow(image ,cmap='gray')
ax2 = fig.add_axes([0.6,0.6,0.4,0.4])
ax2.set_title('mask - Otsu threshold')
plt.imshow(image>threshGlobal,cmap='gray')
#local thresholding
#Try and change this number and see what happens
block_size = 81
#calculate local threshold map
threshLocal =
ax1 = fig.add_axes([0.1,0.2,0.4,0.4])
ax1.imshow(threshLocal,cmap='gray')
ax1.set_title('local threshold map')
ax2 = fig.add_axes([0.6,0.2,0.4,0.4])
ax2.set_title('mask - Local threshold')
plt.imshow(image>threshLocal,cmap='gray')
# -
# # 2. Image segmentation
# ### Image segmentation is the process of partitioning a digital image into multiple segments. The goal of segmentation is to simplify and/or change the representation of an image into something that is more meaningful and easier to analyze.
# <img src="./images/imageSegmentation.png" width="400" height="400" >
#
# ## 2.1 Connected components
# ### After we generate a mask, the simplest segmentation is achieved by taking regions in the mask that are connected and labeling each one as a separate object.
#
# #### We begin by generating a simple mask using the triangle threshold method:
# +
import matplotlib.pyplot as plt
import numpy as np
from skimage import img_as_float
import matplotlib.image as mpimg
from skimage import filters
image_nuclei = img_as_float(mpimg.imread("../Data/xy040-1.png"))
TriThresh = filters.threshold_triangle(image_nuclei)
#despeckle
mask = filters.gaussian(image_nuclei, sigma=1)>TriThresh
# -
# ### [scikit-image's measure module](https://scikit-image.org/docs/dev/api/skimage.measure.html) implements a variety of useful methods for segmentation. the *label* function returns a *labeled image* of connected components (CCs). Each CC is uniquely numbered by an integer.
#
# $\begin{bmatrix}
# 1 & 1 & 0 & 0 & 2\\
# 1 & 1 & 0 & 2 & 2\\
# 0 & 0 & 0 & 0 & 0\\
# 0 & 3 & 0 & 4 & 4\\
# 0 & 0 & 0 & 4 & 4\\
# \end{bmatrix}$
#
# +
from skimage import measure
#generate a labeled matrix of connected components
labels =
plt.figure(figsize=(12,5))
plt.subplot(121)
plt.imshow(mask, cmap='gray')
plt.subplot(122)
plt.imshow(labels, cmap='nipy_spectral')
# -
# #### We can easily generate a *mask* for a **specific** CC using the binary operation *labels==i*
# +
i=43
mask_of_CC_i =
plt.figure(figsize=(10,5))
plt.imshow(mask_of_CC_i, cmap='gray')
# -
# ## Problem with simple CC segmentation : overlapping objects
#
# ### We often really care about having only a single object per label. Using CC, any overlapping objects will merge into one blob:
i=87
mask_of_CC_i = labels==i
plt.imshow(mask_of_CC_i, cmap='gray')
# ### These problems can be partially resolved using morphological operations, but there's no silver bullet
# +
from skimage.morphology import erosion, dilation, opening, closing
from skimage.morphology import disk
#define a "disk" structuring element
selem1 = disk(10)
selem2 = disk(7)
plt.figure(figsize=(15,10))
plt.subplot(121)
plt.gca().set_title('original')
plt.imshow(mask, cmap='nipy_spectral')
plt.subplot(122)
plt.gca().set_title('opening')
plt.imshow(dilation(erosion(mask, selem1),selem2), interpolation='nearest', cmap='nipy_spectral')
# -
# ## 2.2 Watershed based segmentation
#
# ### 2.2.1 The watershed algorithm
# The watershed transformation treats the image it operates upon like a topographic map, with the brightness of each point representing its height, and finds the lines that run along the tops of ridges.
#
# <img src="./images/Diagram-of-watershed-algorithm.png" width="400" height="400" >
#
# More precisely, the algorithm goes as follows:
# 1. *Label* local minima (i.e. $S_1$, $S_2$)
# 2. Move to next higher intensity level
# 3. Assign to each point the label of it's closest label set.
# **<font color='red'>By passing the argument *watershed_line = 1* Points equidistant to multiple sets are labeled as boundaries and intensities set to 0</font>**
# 4. Repeat until all points are labeled
#
# <img src="./images/watershed1.png" width="400" height="400" >
#
# Let's start with a very naive application. We will invert the image, and then simply apply the *watershed* function from the [scikit-image morphology module](https://scikit-image.org/docs/dev/api/skimage.morphology). The function returns a labeled image. We'll plot the edges of that image using a Sobel filter.
# +
import matplotlib.pyplot as plt
import numpy as np
from skimage import img_as_float
import matplotlib.image as mpimg
from skimage import filters
from skimage.util import invert
from skimage.morphology import watershed
image_nuclei = img_as_float(mpimg.imread("../Data/xy040-1.png"))
#invert image
image_to_watershed =
#Calculate watershed transform, remember to pass the watershed_line = 1 argument
labels_naive =
plt.figure(figsize=(15,10))
#let's look at all the boundaries
plt.figure(figsize=(12,20))
plt.subplot(211)
plt.gca().set_title('image fed to watershed')
plt.imshow(image_to_watershed, cmap='gray')
plt.subplot(212)
plt.gca().set_title('watershed result')
plt.imshow(filters.sobel(labels_naive), cmap='nipy_spectral')
# -
# ### So this clearly didn't work. Why? How do we fix it?
#
# Noise generates a ton of local minima. Each gets its own basin. This leads to massive oversegmentation.
#
# #### Watershed segmentation is only a *part* of a segmentation pipeline. Preprocessing (denoising, smoothing, seeding minima) of the image is CRUCIAL for it to work well.
#
# <img src="./images/PreprocessingApproaches.png" width="600" height="400" >
#
# The first thing we'll do is to apply the mask that we found before. This is simply done by adding a *mask* argument to the watershed function.
# +
import matplotlib.pyplot as plt
import numpy as np
from skimage import img_as_float
import matplotlib.image as mpimg
from skimage import filters
from skimage.util import invert
from skimage.morphology import watershed
image_nuclei = img_as_float(mpimg.imread("../Data/xy040-1.png"))
#calculate mask
mask = filters.gaussian(image_nuclei, sigma=1)>TriThresh
#apply mask and invert image
masked image =
inverted_masked_image =
image_to_watershed = inverted_masked_image
#Calculate watershed transform
#Now, also pass the mask to the watershed function so it avoids segmenting the BG
labels_masked =
#let's look at all the boundaries
plt.figure(figsize=(12,20))
plt.subplot(211)
plt.gca().set_title('image fed to watershed')
plt.imshow(image_to_watershed, cmap='gray')
plt.subplot(212)
plt.gca().set_title('watershed result')
plt.imshow(filters.sobel(labels_masked), cmap='nipy_spectral')
# -
# #### So we got rid of all the BG regions, but we are still oversegmenting. Why?
#
# Let's try to smoothen the image and get rid of the many local minima. How wide should the gaussian kernel be?
# +
import matplotlib.pyplot as plt
import numpy as np
from skimage import img_as_float
import matplotlib.image as mpimg
from skimage import filters
from skimage.util import invert
from skimage.morphology import watershed
image_nuclei = img_as_float(mpimg.imread("../Data/xy040-1.png"))
#Calculate mask
mask = filters.gaussian(image_nuclei, sigma=1)>TriThresh
#mask, smooth, and invert the image
masked_image = image_nuclei*mask
sigma_for_smoothing =
smoothed_masked_image =
inverted_smoothed_masked_image = invert(smoothed_masked_image)
image_to_watershed = inverted_smoothed_masked_image
#Calculate watershed transform
#pass the mask to the watershed function so it avoids segmenting the BG
labels_masked_smooth = watershed(image_to_watershed, watershed_line = 1, mask=mask)
#let's look at all the boundaries
plt.figure(figsize=(12,20))
plt.subplot(211)
plt.gca().set_title('image fed to watershed')
plt.imshow(image_to_watershed, cmap='gray')
plt.subplot(212)
plt.gca().set_title('watershed result')
plt.imshow(filters.sobel(labels_masked_smooth), cmap='nipy_spectral')
# -
# ### We're starting to get somewhere!! Can we do better?
#
# #### We can do more to help the algorithm by providing local markers (seeds) from which to start the process
#
# #### We will find seeds by calculating local maxima over areas that are larger than 30x30 pixels using the footprint argument for the function peak_local_max
# +
import matplotlib.pyplot as plt
import numpy as np
from skimage import img_as_float
import matplotlib.image as mpimg
from skimage import filters
from skimage import measure
from skimage.util import invert
from skimage.morphology import watershed
from skimage.feature import peak_local_max
image_nuclei = img_as_float(mpimg.imread("../Data/xy040-1.png"))
mask = filters.gaussian(image_nuclei, sigma=1)>TriThresh
#mask, smooth, and invert the image
masked_image = image_nuclei*mask
sigma_for_smoothing = 4
smoothed_masked_image = filters.gaussian(masked_image, sigma=sigma_for_smoothing)
inverted_smoothed_masked_image = invert(smoothed_masked_image)
image_to_watershed = inverted_smoothed_masked_image
#find local peaks to use as seeds
#focus on this function. Look at the effect of different arguments!
#Specifically. look at the footprint argument
MaskedImagePeaks =
#This is for presentation of our markers
#create disk structuring element of radius 5
selem = disk(5)
#dilate local peaks so that close ones merge
peakMask = dilation(MaskedImagePeaks,selem)
# label local peak regions to find initial markers
markers = measure.label(peakMask)
#pass the *markers* argument to the watershed function
labels_localmax_markers = watershed(image_to_watershed,markers, watershed_line = 1, mask=mask)
#let's look at all the boundaries
plt.figure(figsize=(12,20))
plt.subplot(211)
plt.gca().set_title('image fed to watershed')
plt.imshow(image_to_watershed-peakMask, cmap='gray')
plt.clim((0.95, 1))
plt.subplot(212)
plt.gca().set_title('watershed result')
plt.imshow(filters.sobel(labels_localmax_markers), cmap='nipy_spectral')
# -
# #### This is pretty good! We're still getting a few errors here and there, but there's no big systematic over- or under- segmentation. This is a typical good result when dealing with real data.
# # 3. Feature extraction
# ### Feature extraction is a process of dimensionality reduction by which an initial raw image is reduced to a list of objects and attributes
#
# <img src="./images/feat_ext.png" width="600" height="400" >
#
# ## 3.1 Extracting region properties
#
# ### [scikit-image's measure module](https://scikit-image.org/docs/dev/api/skimage.measure.html) implements a method called *regionprops* that accepts a labeled mask of connected components, and, optionally, a corresponding image, and returns a list. Each object on the list contains useful data about the size, shape, position, and intensity ([see the full list here](https://scikit-image.org/docs/dev/api/skimage.measure.html#skimage.measure.regionprops)) of a specific component.
#
# The length of the list is equal to the total number of objects detected.
#
#
# #### We'll start by extracting the number of CC we found and the area of each CC
# +
from skimage import measure
#We use regionprops to extract properties on all the CCs
props = measure.regionprops(labels_localmax_markers,image_nuclei)
#how many total connected components did we get?
print(len(props))
props[1].perimeter
# +
#This is how we make a list of a specific property for each CC
areas = [r.area for r in props]
#Do the same for the "mean_intensity" property
intensities = [r.mean_intensity for r in props]
#let's look at all the boundaries
plt.figure(figsize=(12,6))
plt.subplot(121)
plt.gca().set_title('areas')
plt.hist(areas)
plt.subplot(122)
plt.gca().set_title('intensities')
plt.hist(intensities)
# -
# ## 3.2 Some options for data presentation
#
# **We can look at individual objects we found**
i=2
plt.imshow(props[i].intensity_image)
plt.gca().set_title('Single cell')
# **Let's use a scatter plot to compare our results to the image**
# +
intensities = np.array([r.mean_intensity for r in props])
centroids = np.array([r.centroid for r in props])
fig = plt.figure(figsize=(12,15))
fig.add_axes([0.1,0.6,0.4,0.25])
plt.gca().set_title('original')
plt.imshow(image_nuclei, interpolation='nearest', cmap=plt.cm.gray, vmin=0, vmax=0.02)
fig.add_axes([0.6,0.6,0.4,0.25])
plt.scatter(centroids[:,1],centroids[:,0], c=intensities)
plt.axis('equal')
plt.gca().invert_yaxis()
# -
# **Or even nicer scatter plots!**
# +
intensities = np.array([r.mean_intensity for r in props])
areas = np.array([r.area for r in props])
centroids = np.array([r.centroid for r in props])
fig = plt.figure(figsize=(12,15))
fig.add_axes([0.1,0.6,0.4,0.25])
plt.gca().set_title('original')
plt.imshow(image_nuclei, interpolation='nearest', cmap=plt.cm.gray, vmin=0, vmax=0.02)
fig.add_axes([0.6,0.6,0.4,0.25])
plt.scatter(centroids[:,1],centroids[:,0], c=intensities, s=areas/20)
plt.axis('equal')
plt.gca().invert_yaxis()
plt.text(centroids[10,1],centroids[10,0],props[10].label)
# -
# **You can even draw your points directly on the image!**
# +
intensities = np.array([r.mean_intensity for r in props])
areas = np.array([r.area for r in props])
centroids = np.array([r.centroid for r in props])
fig = plt.figure(figsize=(12,15))
fig.add_axes([0.1,0.6,0.8,0.5])
plt.gca().set_title('original')
plt.imshow(image_nuclei, interpolation='nearest', cmap=plt.cm.gray, vmin=0, vmax=0.02)
plt.gca().patch.set_alpha(0.5)
plt.scatter(centroids[:,1],centroids[:,0], c=intensities, alpha=1)
# -
# ## 3.3 Converting regionprops to a table (Dataframe)
#
# #### Let's define some useful functions for converting a list of props into a pandas dataframe. These should become obsolete soon since the new version of scikit-image will have this functionality #### These are some useful functions for converting a list of props into a pandas dataframe
# +
import pandas as pd
def scalar_attributes_list(im_props):
"""
Makes list of all scalar, non-dunder, non-hidden
attributes of skimage.measure.regionprops object
"""
attributes_list = []
for i, test_attribute in enumerate(dir(im_props[0])):
#Attribute should not start with _ and cannot return an array
#does not yet return tuples
if test_attribute[:1] != '_' and not\
isinstance(getattr(im_props[0], test_attribute), np.ndarray):
attributes_list += [test_attribute]
return attributes_list
def regionprops_to_df(im_props):
"""
Read content of all attributes for every item in a list
output by skimage.measure.regionprops
"""
attributes_list = scalar_attributes_list(im_props)
# Initialise list of lists for parsed data
parsed_data = []
# Put data from im_props into list of lists
for i, _ in enumerate(im_props):
parsed_data += [[]]
for j in range(len(attributes_list)):
parsed_data[i] += [getattr(im_props[i], attributes_list[j])]
# Return as a Pandas DataFrame
return pd.DataFrame(parsed_data, columns=attributes_list)
# -
# ### Now, to get all the properties in table form we simply run:
props_df = regionprops_to_df(props)
props_df
# ### Finally, if we imaged our cells in multiple channels, we would want to use the same segmented nuclei and measure intensities of other channels.
# +
from skimage import measure
from skimage import img_as_float
import matplotlib.image as mpimg
image_2ndChannel = img_as_float(mpimg.imread("../Data/xy040-2.png"))
# extract regionprops using labels_localmax_markers mask from image_2ndChannel
props_other_channel = measure.regionprops(labels_localmax_markers,image_2ndChannel)
plt.figure(figsize=(12,6))
plt.subplot(121)
plt.gca().set_title('Nuclei')
plt.imshow(image_nuclei, cmap='gray')
plt.subplot(122)
plt.gca().set_title('other channel')
plt.imshow(image_2ndChannel, cmap='gray')
# -
# ### Extract only the intensity related features
# +
mean_2nd_channel = [r.mean_intensity for r in props_other_channel]
max_2nd_channel = [r.max_intensity for r in props_other_channel]
min_2nd_channel = [r.min_intensity for r in props_other_channel]
plt.gca().set_title('intensities of 2nd channel')
plt.hist(mean_2nd_channel)
# -
# ### Add these new features to the pandas dataframe
# +
props_df['mean_intensity_ch2'] = mean_2nd_channel
props_df['max_intensity_ch2'] = max_2nd_channel
props_df['min_intensity_ch2'] = min_2nd_channel
props_df
# -
# **Sometimes it's easier to see a bimodal distribution in log scale**
# +
fig = plt.figure(figsize=(12,6))
fig.add_axes([0.1,0.1,0.4,0.4])
plt.gca().set_title('Histogram of intensities')
plt.hist(props_df.mean_intensity_ch2,20)
fig.add_axes([0.6,0.1,0.4,0.4])
plt.gca().set_title('Histogram of log of intensities')
plt.hist(np.log(props_df.mean_intensity_ch2),20)
# -
# **We can compare distributions of different channels**
# +
plt.figure(figsize=(12,6))
plt.gca().set_title('scatter plot of intensities')
plt.scatter(np.log(props_df['max_intensity_ch2']), np.log(props_df['max_intensity']))
plt.xlabel('Ch2')
plt.ylabel('Ch1')
# -
# ### And so, we've successfully implemented a simple image segmentation pipeline for multicolor microscopy data.
#
#
# #### Fin.
#
#
#
#
| 20190511/JupyterNotebooks/DIP_AOY_Student.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Medidas de riesgo
#
# <img style="float: right; margin: 0px 0px 15px 15px;" src="http://www.thebluediamondgallery.com/wooden-tile/images/risk.jpg" width="450px" height="150px" />
#
# > Existen $10^{11}$ estrellas en la galaxia. Ese solía ser un número grandísimo. Sin embargo son solo cien billones. Es menos que la deuda nacional! (de EUA) Solíamos llamarlos números astronómicos. Ahora, deberíamos llamarlos números económicos.
#
# **<NAME>**
#
# Referencias:
# - www.risklab.es/es/seminarios/pasados/septiembre2005.pdf
# - www.emagister.com/uploads_user_home/Comunidad_Emagister_5840_valor.pdf
# - repositorio.uchile.cl/bitstream/handle/2250/127411/149%20Medidas_de_Riesgo_Financiero_Rafael_Romero_M.pdf
#importar los paquetes que se van a usar
import pandas as pd
import pandas_datareader.data as web
import numpy as np
from sklearn.neighbors import KernelDensity
import datetime
from datetime import datetime
import scipy.stats as st
import scipy as sp
import scipy.optimize as scopt
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
#algunas opciones para Python
pd.set_option('display.notebook_repr_html', True)
pd.set_option('display.max_columns', 6)
pd.set_option('display.max_rows', 10)
pd.set_option('display.width', 78)
pd.set_option('precision', 3)
# ## 0. Introducción y apuntes históricos
#
# Existen en el mundo factores fundamentales que han contribuido al desarrollo de la gestión, medición, y el control del riesgo financiero:
#
# - Alto nivel de inestabilidad económica en los mercados.
# - Fundamentalmente la volatilidad se ve reflejada, en los siguientes factores: La volatilidad de los mercados accionarios, la volatilidad de la tasa de cambio, la volatilidad de la tasa de interés y la volatilidad de los precios en el mercado de los commodities.
# - Crecimiento de las actividades de negociación.
# - Durante los últimos años, tanto el número de activos negociados como el volumen de éstos, ha tenido un incremento considerable a nivel mundial. Se ha evolucionado considerablemente en el desarrollo de numerosos tipos de instrumentos, los cuales han facilitado las transacciones sobre activos previamente ilíquidos. Los mercados de derivados financieros como las opciones, futuros, forwards y swaps, han tenido un crecimiento notable a partir de los años 70, cuando hubo un importante desarrollo de sus aspectos teóricos, liderado por autores como <NAME>, <NAME>, Myron Scholes.
# - Avances en la tecnología.
# - Mejor poder computacional, mejores técnicas computacionales. Se ha avanzado en la cultura de la información, ya que las empresas han comenzado a tomar conciencia acerca de la importancia de tener bases de datos, esenciales para un posterior análisis del riesgo. Estos avances tecnológicos permiten obtener, de forma rápida, información fundamental para la toma de decisiones de inversión.
# ___
# ### Conceptos básicos relativos al riesgo
#
# - Etimológicamente la palabra riesgo deriva del italiano risico o rischio, y éste a la vez del árabe risq, que significa ”lo que depara la providencia”.
#
# - La palabra riesgo puede entenderse como la contingencia o proximidad a un daño; es decir, tiene que **ver con el futuro**, con la necesidad de prever y anticiparnos antes de cierta contingencia.
#
# - En el contexto de las finanzas, cuando hablamos de riesgo nos estamos refiriendo a la **posibilidad de pérdida o ganancia** debido a los cambios sobre los factores que afectan el valor de un activo. Por esa razón, es importante que se identifiquen, se midan, se controlen, y se haga un **monitoreo** de los diversos tipos de riesgo a los que están expuestos los inversores en los mercados de capitales.
#
# - Por lo general, el riesgo se percibe como una exposición a eventos negativos; no obstante, el riesgo bien manejado representa una gran oportunidad de obtener rentabilidades significativas en los portafolios financieros. En este sentido, y con una visión positiva, el riesgo financiero es más considerado como una _“oportunidad de ganar”_ que como una “posibilidad de perder”.
# ___
# ### Tipos de riesgos financieros
#
# 1.**Riesgo de mercado**, asociado a las fluctuaciones de los mercados financieros, y en el que se distinguen:
# - Riesgo de cambio, consecuencia de la volatilidad del mercado de divisas.
# - Riesgo de tipo de interés, consecuencia de la volatilidad de los tipos de interés.
# - Riesgo de mercado (en acepción restringida), que se refiere específicamente a la volatilidad de los mercados de instrumentos financieros tales como acciones, deuda, derivados, etc.
#
# 2.**Riesgo de crédito**, consecuencia de la posibilidad de que una de las partes de un contrato financiero no asuma sus obligaciones.
#
# 3.**Riesgo de liquidez o de financiación**, y que se refiere al hecho de que una de las partes de un contrato financiero no pueda obtener la liquidez necesaria para asumir sus obligaciones a pesar de disponer de los activos —que no puede vender con la suficiente rapidez y al precio adecuado— y la voluntad de hacerlo.
#
# 4.**Riesgo operativo**, que es entendido como la posibilidad de ocurrencia de pérdidas financieras, originadas por fallas o insuficiencias de procesos, personas, sistemas internos, tecnología, y en la presencia de eventos externos imprevistos.
#
# 5.**Riesgo país o riesgo soberano**, es todo riesgo inherente a operaciones transnacionales y, en particular, a las financiaciones desde un país a otro.
#
# 6.**Riesgo sistémico**, puede ser interpretado como "inestabilidad del sistema financiero, potencialmente catastrófico, causado por eventos idiosincráticos o condiciones en los intermediarios financieros".
#
# > Referencia:https://es.wikipedia.org/wiki/Riesgo_financiero
# ### Apuntes históricos
#
# Históricamente se pueden identificar tres periodos relevantes en cuanto al desarrollo de las finanzas modernas:
#
# 1. **Modelo de media-varianza (Markowitz, 1952-1956)**
# - Antes de esto el riesgo financiero era considerado como un *factor de corrección* entre el retorno esperado de una inversión y el retorno real. De modo que no se podía definir el "riesgo" sino hasta que se tenía el resultado de la decisión de inversión. Markowitz propuso como medidas de riesgo la varianza (para inversiones individuales) y la covarianza (para portafolios). Con esta forma de medir el riesgo se pudo optimizar...
# $$P \ \text{(Portafolio->Retornos normales)}\\
# Var(P) = \sigma^2=x_1^2\sigma_1^2 +2x_1x_2 \sigma_{12}+ x_2^2 \sigma_2^2$$
#
#
# 2. **Modelos en tiempo continuo (Merton, Black, Scholes, 1969-1973)**
# - Estos modelos fueron un gran paso teórico que se evidenciaron prácticamente con la introducción de los instrumentos derivados, puesto que permitieron su valuación.
#
# 3. **Medidas de riesgo (Artzner, 1997-1999)**
# - Desarrollos que pretenden modelar situaciones más reales como sesgo, colas anchas, etcétera.
# ___
# Supongamos que tenemos un portafolio de acciones. ¿Qué tan riesgosa es esa inversión?
#
# **Objetivo:** producir un único numero para resumir la exposición de la inversión al riesgo de mercado.
#
# - Pregunta inicial: ¿Cuánto podría perder en el peor escenario?
# - Mala pregunta: lo podrías perder todo.
#
# Bueno, pero
#
# - ¿Cuál es la pérdida que estamos $100\alpha \%$ confiados que no excederemos en $T$ tiempo?
#
# Ejemplo:
#
# $VaR_{0.9}=1000.000$ en 10 días:
# - Estoy 90% seguro que no perderé más de $1000.000$ en mi inversión en los siguientes 10 días.
# - Hay un 90% de probabilidad que mi pérdida sea menor a $1000.000$ en los siguientes 10 días.
# - Hay un 10% de probabilidad que mi pérdida sea mayor a $1000.000$ en los siguientes 10 días.
# ## 1. Value-at-Risk (VaR)
#
# Dado un horizonte temporal $T$ y un nivel de confianza $100 \alpha \%$, el **VaR** es la pérdida máxima respecto al rendimiento esperado sobre el horizonte temporal $T$ que sólo es superada con una probabilidad $1 − \alpha$.
#
# $$ VaR = \mu - \alpha$$
#
# donde $\mu$ es el retorno medio, $\alpha$ es el menor valor tal que a la izquierda de ese valor la probabilidad este en cierto nivel, por ejemplo 1%.
# 
# ### Ejemplo:
#
# Suponga que el retorno de cada uno de los dos activos que forman el portafolio se distribuye normal. Un activo tiene un retorno esperado de 20% y el otro de 15%. La varianza del primer activo es 0,08, la del segundo es 0,05 y la covarianza es 0,02. La ponderación de cada activo es la misma.
#
# El retorno esperado del portafolio es:
# $$\mu = {1 \over 2} 0.20+{1 \over 2}0.15 =0.175$$
#
# La varianza del portafolio es:
# $$\sigma^2 = {1 \over 2}^2 0.08 +{1 \over 2}^2 0.05 + 2{1 \over 2}{1 \over 2} 0.02 = 0.043$$
# $$ \sigma = \sqrt 0.043 = 0.206$$
#
# ¿Cómo definir $\alpha$?
#
# Se debe de tomar un significancia que oscila tipicamente entre el 1%,5% o 10%. Luego se debe de encontrar el valor de la variable aleatoria tal que a la izquierda de ese valor, quede el nivel de significancia tomado. Por ejemplo si se toma un nivel de significancia del 1%, $\alpha$ se encuentra como:
alpha = st.norm.ppf(0.05,loc=0.175,scale=0.206)
alpha
# Finalmente se encuentra que:
# $$VaR = 0.175 +0.305=0.48$$
#
# Que se interpreta como que hay un 1% de probabilidad de perder más que 48% en el próximo período.
# ## Parte práctica
# Descargamos precios de cierre de activos correspondientes a American Airlines y a Coca-Cola, entre 2010 y 2017
# +
def get_historical_closes(ticker, start_date, end_date=None):
# # closes = web.YahooDailyReader(ticker, start_date, end_date).read().sort_index('major_axis')
# closes = web.YahooDailyReader(symbols=ticker, start=start_date, end=end_date).read()
# index = pd.to_datetime(closes.loc['date',:,tickers[0]].values)
# closes.set_axis(index, axis=1, inplace=True)
# closes = closes.loc['adjclose']#.sort_index().dropna()
# closes = pd.DataFrame(np.array(closes.as_matrix(), dtype=np.float64), columns=ticker, index=closes.index)
# closes.index.name = 'Date'
# return closes#.loc['Adj Close'].dropna()
closes = web.DataReader(ticker, 'iex', start)
index = pd.to_datetime(closes[ticker[0]].index)
closes = np.asarray([closes[i]['close'] for i in ticker]).T
closes = pd.DataFrame(closes, columns=ticker,index= index)
return closes
####### Calculation of log-returns
def calc_daily_ret(closes):
return np.log(closes/closes.shift(1)).dropna()
# +
start = '2013-01-01'
# end = '2018-04-30'
ticker = ['AAPL','KO','AAL']#,'SPY']
closes = get_historical_closes(ticker, start)
closes.plot(figsize=(8,6));
# +
# ticker = ['AA','KO']
# start_date = '2010-01-01'
# end_date = '2017-12-31'
# closes = get_historical_closes(ticker, start_date, end_date)
# closes.plot(figsize=(8,6));
# -
# Calculamos rendimientos...
daily_ret = calc_daily_ret(closes)
daily_ret.plot(figsize=(8,6));
# Definimos un portafolio con las anteriores acciones. Tomemos igual participación en ambas...
def def_portfolio(tickers, participation=None):
if (participation is None):
participation = np.ones(len(tickers))/len(tickers)
portfolio = pd.DataFrame({'Tickers': tickers, 'Participation': participation}, index=tickers)
return portfolio
portfolio = def_portfolio(ticker, participation=np.array([1,1,1])/3)
portfolio
# Con la información podemos calcular rendimientos anuales, o mensuales
# +
def calc_annual_ret(daily_ret):
return np.exp(daily_ret.groupby(lambda date: date.year).sum())-1
def calc_monthly_ret(daily_ret):
return np.exp(daily_ret.groupby(lambda date: '%4d-%02d'%(date.year,date.month)).sum())-1
# Ejemplificar el funcionamiento de la función
# daily_ret.groupby(lambda date: date.year).sum()
# daily_ret.index.year
# -
annual_ret = calc_annual_ret(daily_ret)
monthly_ret = calc_monthly_ret(daily_ret)
# Graficar...
monthly_ret.plot(figsize=(8,6));
annual_ret.plot(figsize=(8,6));
# Calculamos el rendimiento ponderado del portafolio (mensual)
# +
def calc_portfolio_ret(portfolio, returns, name='Value'):
total_sum = portfolio.Participation.sum()
weight = portfolio.Participation/total_sum
weighted_ret = returns*weight
return pd.DataFrame({name: weighted_ret.sum(axis=1)})
# Explicación de los comandos
# p = portfolio.Participation/1
# w=(monthly_ret*p).sum(axis=1)
# monthly_ret,w
# -
portfolio_ret_m = calc_portfolio_ret(portfolio, monthly_ret)
portfolio_ret_m.plot(figsize=(8,6));
# ### Métodos para estimar el VaR
#
# **Con datos históricos**
#
# Calculamos cuantiles empíricos de un histograma de rendimientos (diarios, semanales, en periodos de N días, mensuales, anuales).
#
# Como ejemplo, tomamos los rendimientos mensuales del portafolio.
portfolio_ret_m.hist(bins=30, normed=True, histtype='stepfilled', alpha=0.5);
# El cuantil empírico 0.01 de los retornos mensuales del portafolio, menos el retorno mensual esperado ($VaR_{0.99}$ a un mes) es:
portfolio_ret_m.mean().Value-portfolio_ret_m.quantile(0.01).Value
# Esto quiere decir que con un $99\%$ de confianza, la peor pérdida mensual del portafolio no será mayor a $9.92\%$.
# > ## **Ejercicio**
# > Calcular el $VaR_{0.95}$ a un año.
portfolio_ret_a = calc_portfolio_ret(portfolio, annual_ret)
portfolio_ret_a.plot(figsize=(8,6));
portfolio_ret_a.mean().Value-portfolio_ret_a.quantile(0.05).Value
# En un año, la máxima pérdida podría ser de $24\%$ con una confianza del $95\%$.
# **Usando métodos Monte Carlo**
#
# Simulamos varios escenarios con condiciones aleatorias de mercado. Calculamos la pérdida para cada escenario. Usamos los datos de los escenarios para establecer el riesgo del portafolio.
# Ajustamos una función de densidad empírica con Kernel Gaussianos
kde = KernelDensity(kernel='gaussian', bandwidth=0.001).fit(portfolio_ret_m)
# Interpretación de `bandwidth`
# 
# Simulamos varios escenarios de rendimientos...
nscen = 100000
ret_sim = kde.sample(n_samples=nscen, random_state=None)
plt.hist(ret_sim, bins=30);
# Obtenemos el cuantil 0.01 de los retornos simulados del portafolio, menos el retorno mensual esperado ($VaR_{0.99}$ a un mes) es:
np.mean(ret_sim)-np.percentile(ret_sim, 1)
# Esto quiere decir que con un $99\%$ de confianza, la peor pérdida mensual del portafolio no será mayor a $14.19\%$.
# **Ejercicio** Calcular el $VaR_{0.95}$ a un año.
kde_a = KernelDensity(kernel='gaussian', bandwidth=0.001).fit(portfolio_ret_a)
nscen = 100000
ret_sim = kde_a.sample(n_samples=nscen, random_state=None)
plt.hist(ret_sim, bins=30);
np.mean(ret_sim)-np.percentile(ret_sim, 5)
# En un año, la máxima pérdida podría ser de $24.8\%$ con una confianza del $95\%$.
# ## 2. Deficiencia del VaR como medida de riesgo
#
# Matemáticamente:
#
# - Una medida de riesgo es una función que cuantifica riesgo (como el VaR)
#
# - Una medida de riesgo determina la cantidad de un capital que debe ser reservada. El objetivo de esta reserva es garantizar la presencia de capital que pueda cubrir la manifestación de un evento riesgoso que produzca una pérdida (crédito).
# - Desde un punto de vista matemático, una medida de riesgo es una función $\rho:\mathcal{L}\to\mathbb{R}\cup\lbrace\infty\rbrace$, donde $\mathcal{L}$ es el espacio lineal de pérdidas.
# ### Coherencia
#
# Una medida de riesgo se dice coherente si:
#
# 1. Es **monótona**: $Z_1,Z_2\in\mathcal{L}$, $Z_1\leq Z_2$ entonces $\rho(Z_1)\leq\rho(Z_2)$. Implica que hay posibilidad de un ordenamiento (riesgo mayor asociado a mayor pérdida o beneficio).
# 2. Es **subaditiva**: $Z_1,Z_2\in\mathcal{L}$ entonces $\rho(Z_1+Z_2)\leq\rho(Z_1)+\rho(Z_2)$. Implica que hay un incentivo a la diversificación.
# 3. Es **positivamente homogenea**: $a\geq 0$, $Z\in\mathcal{L}$ entonces $\rho(aZ)=a\rho(Z)$. Implica proporcionalidad del riesgo.
# 4. Es **translacionalmente invariante**: $b\in\mathbb{R}$, $Z\in\mathcal{L}$ entonces $\rho(Z+b)=\rho(Z)-b$. Implica que por el hecho de agregar retornos por valor seguro $b$ a un retorno aleatorio $Z$ el riesgo $\rho$ decrece por un factor de $b$.
# A parte de la implicación financiera de la subaditividad, esta condición está fuertemente relacionada con la convexidad (propiedad de prioritaria en optimización).
# **El VaR falla, en general, en la subaditividad.**
# ## 3. (Co)Varianza como medida de riesgo.
#
# El riesgo no son solo eventos de pérdida. De hecho, como dijimos antes, el riesgo es una oportunidad para ganar. El riesgo finalmente esta asociado con *no saber*.
#
# Dado que los portafolios son funciones lineales de activos, la covarianza es convexa (en particular subaditiva), lo cual es esencial en la teoría de Markowitz.
#
# Veamos un ejemplo con los datos que tenemos...
# Covarianza y correlacion de los activos
monthly_ret.cov()
monthly_ret.corr()
# Concatenamos con portafolio y calculamos varianzas individuales...
total_ret = pd.concat([monthly_ret, portfolio_ret_m], axis=1)
total_ret
total_ret.std()**2
# Verificación subaditividad
(total_ret.std()**2).Value<=(total_ret.std()**2).AAL+(total_ret.std()**2).KO \
+(total_ret.std()**2).AAPL
# Gráfica
total_ret.plot(figsize=(8,6));
sns.jointplot('AAL', 'KO', data=monthly_ret, color="k").plot_joint(sns.kdeplot, zorder=0, n_levels=60);
sns.jointplot('AAPL','AAL', data=monthly_ret, color="k").plot_joint(sns.kdeplot, zorder=0, n_levels=30);
# Beneficia mucho: correlación negativa o en su defecto cercana a cero.
# **Ejemplo**: considere los activos con los siguientes rendimientos y un portafolio de igual participación.
rendimientos = pd.DataFrame({'Acción A': [0.1, 0.24, 0.05, -0.02, 0.2],
'Acción B': [-0.15, -0.2, -0.01, 0.04, -0.15]})
rendimientos
portafolio = def_portfolio(['Acción A', 'Acción B'])
portafolio
rendimientos_port = calc_portfolio_ret(portafolio, rendimientos)
rendimientos_port
rendimientos_totales = pd.concat([rendimientos, rendimientos_port], axis=1)
rendimientos.cov()
rendimientos.corr()
rendimientos_totales.std()**2
rendimientos_totales.plot(figsize=(8,6));
# ## Tarea
#
# Consultar la medida CVaR (VaR condicional). Dar una breve descripción usando LaTeX en celdas Markdown.
#
# Además, implementar en python, tomar como ejemplo un par de activos y verificar subaditividad.
# ## Trabajo
# 1. Se cuenta con tres portafolios, compuesto por los siguientes activos:
# - Alcoa Corporation (AA), American Airlines (AAL), Moneygram International Inc. (MGI) , Apple (AAPL),American Eagle (AEO), SL Green Realty Corporation (SLG), Westwater Resources Inc. (WWR), eBay Inc. (EBAY), Facebook Inc. (FB), Twitter Inc. (TWTR).
# - Starbucks Corporation (SBUX), Microsoft Corporation (MSFT), Coca-Cola Company (The) (KO), Nike Inc. (NKE), Texas Instruments Incorporated (TXN), Bank of America Corporation (BAC), McDonald's Corporation (MCD), The Walt Disney Company (DIS), Walmart Inc. (WMT), Best Buy Co. Inc. (BBY), Amazon.com Inc. (AMZN).
# - MGM Resorts International (MGM), Universal Corporation (UVV), Pandora Media Inc. (P), Spotify Technology S.A. (SPOT), Netflix Inc. (NFLX), Motorola Solutions Inc. (MSI), Twenty-First Century Fox Inc. (FOX), Paramount Group Inc. (PGRE), Sony Corporation (SNE), Viacom Inc. (VIA), Time Warner Inc. (TWX).
#
# El objetivo es obtener el VaR de cada uno de los portafolios, bajo los siguientes supuestos:
# - Suponer que la participación de cada uno de los activos es la misma.
#
# > Página de donse se extraen los datos: https://iextrading.com/apps/stocks/#/BWA
#
# ## Puntos extra
# - Encontrar la participación óptima de cada portafolio usando Marcowitz.
# <script>
# $(document).ready(function(){
# $('div.prompt').hide();
# $('div.back-to-top').hide();
# $('nav#menubar').hide();
# $('.breadcrumb').hide();
# $('.hidden-print').hide();
# });
# </script>
#
# <footer id="attribution" style="float:right; color:#808080; background:#fff;">
# Created with Jupyter by <NAME>. and modified by <NAME>.
# </footer>
| TEMA-3/Clase22_MedidasRiesgo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Обучение с учителем
# ## Применение линейной регрессии
# В этом задании вам предлагается изучить и применить на практике модели линейной регресии,
# доступные в библиотеке `scikit-learn`.
# Модель линейной регрессии имеет ряд проблем, связанных с переобучением. Другими словами, модель слишком точно приближает зависимость между признаками и целевой переменной на тренировочной выборке, и, как результат, делает достаточно грубые или неточные предсказания на новых данных. Регуляризация является эффективным решением данной проблемы. Введем понятие Гребневой Регрессии (Ridge Regression).
#
# Сформулируем задачу минимизации функционала $Q(\beta)$ для нахождения линейной зависимости между целевой переменной $Y$ и признаками.
#
# $$
# \begin{equation*}
# Q(\beta) = \| Y - X \beta \|^2 + \lambda \| \beta \|^2 \rightarrow \min\limits_{\beta}.
# \end{equation*}
# $$
#
# В данном случае, $X$ - матрица признаков, $\beta$ - вектор параметров, который требуется найти. Таким образом, мы добавляем к функции потерь регуляризатор с параметром $\lambda$, штрафующий большие значения вектора весов $\beta$.
#
# Настройка коэффициента регуляризации $\lambda$ - это важная задача в построении модели Гребневой регрессии.
# ## Загрузка и разбиение данных
# Мы будем решать задачу линейной регресии на наборе данных `Boston House Prices`. Он представляет из себя набор различных факторов, которые некоторым образов влияют на среднюю рыночную цену дома в разных районах Бостона. Например, уровень преступности или среднее количество комнат. Наша задача - предсказать цену нового объекта, опираясь на значения этих признаков.
#
# Начнем с загрузки этого набора данных и его разбиения. `Boston House Prices` является одним из втроенных наборов данных, доступных в `scikit-learn`. Для его загрузки достаточно вызвать функцию `load_boston` из `sklearn.datasets`. Далее, предлагается случайно разбить выборку на тренировочную и тестовую выборку, используя функцию `train_test_split` из `sklearn.model_selection`, вызвав ее с параметром `random_state=54` и `test_size=0.33`.
# +
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_boston
boston = load_boston()
X_train, X_test, y_train, y_test = train_test_split(boston.data, boston.target, random_state=54, test_size=0.33)
# -
# ## Настройка параметра регуляризации в Гребневой регрессии
# Модель Гребневой регрессии представлена классом `Ridge` из `sklearn.linear_model`. Конструктор этого класса содержит аргумент `alpha`, обратное значение которого соответствует параметру регуляризации $\lambda$. При использовании данной модели установите значение аргумента `random_state=42`. Найдите оптимальное значение коэффициента `alpha` в диапазоне `{10, 20, 30, ..., 90}` начиная со значения 10 до 90 с шагом 10, соответствующее минимальному значению среднеквадратической ошибки на тестовой выборке. Так же укажите это минимальное значение, округленное до трех знаков после запятой. Эти два значения будут являться первым `answer1` и вторым `answer2` ответом в этом практическом задание. Постройте график зависимости среднеквадратической ошибки от значения параметра `alpha`.
# ### *РЕШЕНИЕ*
# +
import numpy as np
from sklearn.linear_model import Ridge
from sklearn.metrics import mean_squared_error
scores = []
coef = []
for alpha in range(10, 100, 10):
model = Ridge(alpha=alpha, random_state=42)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
mse_score = mean_squared_error(y_test, y_pred)
scores.append(mse_score)
coef.append(alpha)
index = np.array(scores).argmin()
answer1 = coef[index]
answer2 = round(scores[index], 3)
# +
from matplotlib import pyplot as plt
plt.plot(coef, scores, '--o')
plt.title("MSE(alpha)")
plt.xlabel("alpha")
plt.ylabel("mean squared error, 1000$'s")
plt.show()
# -
# В результате, мы нашли оптимальное значение коэффициента `alpha` из предложенных, которое соответствует наиболее точной модели на новых данных. Таким образом, Гребневая регрессия собирает максимум информации из предложенных тренировочных данных.
# ## Метод регрессии LASSO
# Еще одним методом регуляризации, похожем на Гребневую регрессию, является LASSO (least absolute shrinkage and selection operator). В данном случае, задача минимизации функции потерь формулируется следующим образом.
#
# $$
# \begin{equation*}
# \left\lbrace
# \begin{array}{c}
# Q(\beta) = \| Y - X \beta \|^2, \\
# \sum\limits_{i=0}^N |\beta_i| < \varkappa.
# \end{array}
# \right.
# \end{equation*}
# $$
#
# При уменьшении значения $\varkappa$ все больше коэффициентов $\beta_i$ обнуляются. Таким образом, LASSO является своеобразным методом отбора признаков и понижения размерности, подробнее о которых вы узнаете на 4-й неделе нашего курса. Этот метод регуляризации выбирает самые информативные признаки, отбрасывая остальные.
# Загрузите набор данных `Diabetes`, используя функцию `load_diabetes` из `sklearn.datasets`. `Diabetes` содержит в себе информацию о различных признаках, таких как возраст, пол, среднее кровяное давление человека и прочие. В качестве целевой переменной выступает количественный показатель прогрессирования заболевания диабет через год после определенного периода (baseline).
# +
from sklearn.datasets import load_diabetes
diabetes = load_diabetes()
# -
# Метод LASSO представлен классом `Lasso` из `sklearn.linear_model`. Обучите эту модель на всем наборе данных `Diabetes`. Выберете в качестве параметра конструктора класса `random_state=42`. Какое количество признаков было отсеяно данной моделью? Для этого проанализируйте вектор весов `coef_`, являющийся атрибутом данной модели. В качестве ответа `answer3` на это задание приведите отношение числа выбранных моделью признаков к их общему числу. Ответ округлите до одного знака после запятой.
# ### *РЕШЕНИЕ*
# +
from sklearn.linear_model import Lasso
model = Lasso(random_state=42)
model.fit(diabetes.data, diabetes.target)
numFeatures = len(diabetes.feature_names)
answer3 = float(numFeatures - list(model.coef_).count(0)) / numFeatures
answer3 = round(answer3, 1)
# -
# Далее обучите модель `Lasso` с параметром конструктора `random_state=42` на тренировочной выборке датасета `Boston House Prices` и посчитайте значение среднеквадратической ошибки на тестовой выборке. Приведите это значение в качестве ответа `answer4`, округленного до трех знаков после запятой. Сравните это значение с результатом, полученным Гребневой регрессией.
# ### *РЕШЕНИЕ*
model = Lasso(random_state=42)
model.fit(X_train, y_train)
y_pred = model.predict(X_test)
mse_score = mean_squared_error(y_test, y_pred)
answer4 = round(mse_score, 3)
print("mse error(ridge) < mse error(lasso) ? {}".format(answer2 < answer4))
# # Строка с ответами
output = "alpha: {0}\nmse error(ridge): {1:.3f}\nratio: {2:.1f}\nmse error(lasso): {3:.3f}"
print(output.format(answer1, answer2, answer3, answer4))
| W3/reguralization_test/Teacher_solve.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import datetime
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.dates as mdates
import pandas as pd
from pandas_datareader import data, wb
import pprint
import statsmodels.tsa.stattools as ts
def plot_price_series(df, ts1, ts2):
months = mdates.MonthLocator() # every month
fig, ax = plt.subplots()
ax.plot(df.index, df[ts1], label=ts1)
ax.plot(df.index, df[ts2], label=ts2)
ax.xaxis.set_major_locator(months)
ax.xaxis.set_major_formatter(mdates.DateFormatter('%b %Y'))
ax.set_xlim(datetime.datetime(2012, 1, 1), datetime.datetime(2013, 1, 1))
ax.grid(True)
fig.autofmt_xdate()
plt.xlabel('Month/Year')
plt.ylabel('Price ($)')
plt.title('%s and %s Daily Prices' % (ts1, ts2))
plt.legend()
plt.show()
def plot_scatter_series(df, ts1, ts2):
plt.xlabel('%s Price ($)' % ts1)
plt.ylabel('%s Price ($)' % ts2)
plt.title('%s and %s Price Scatterplot' % (ts1, ts2))
plt.scatter(df[ts1], df[ts2])
plt.show()
def plot_residuals(df):
months = mdates.MonthLocator() # every month
fig, ax = plt.subplots()
ax.plot(df.index, df["res"], label="Residuals")
ax.xaxis.set_major_locator(months)
ax.xaxis.set_major_formatter(mdates.DateFormatter('%b %Y'))
ax.set_xlim(datetime.datetime(2012, 1, 1), datetime.datetime(2013, 1, 1))
ax.grid(True)
fig.autofmt_xdate()
plt.xlabel('Month/Year')
plt.ylabel('Price ($)')
plt.title('Residual Plot')
plt.legend()
plt.plot(df["res"])
plt.show()
# +
start = datetime.datetime(2012, 1, 1)
end = datetime.datetime(2013, 1, 1)
arex = data.DataReader("AREX", "yahoo", start, end)
wll = data.DataReader("WLL", "yahoo", start, end)
df = pd.DataFrame(index=arex.index)
df["AREX"] = arex["Adj Close"]
df["WLL"] = wll["Adj Close"]
# Plot the two time series
plot_price_series(df, "AREX", "WLL")
# Display a scatter plot of the two time series
plot_scatter_series(df, "AREX", "WLL")
# Calculate optimal hedge ratio "beta"
res = ols(y=df['WLL'], x=df["AREX"])
beta_hr = res.beta.x
# Calculate the residuals of the linear combination
df["res"] = df["WLL"] - beta_hr*df["AREX"]
# Plot the residuals
plot_residuals(df)
# Calculate and output the CADF test on the residuals
cadf = ts.adfuller(df["res"])
pprint.pprint(cadf)
# -
| quant/plot_yahoo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/KshitijBhat/ML_Enthusiasts_Ishana/blob/main/Main.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="bj1ySrITL5d-"
# Requires GPU (CUDA). Please connect to a GPU runtime.
#
# Be sure to mount/upload the drive folder `ML_enthsiasts_Ishana`
# + id="2Nq2UCSa5G0J"
import os
import cv2
import numpy as np
import matplotlib.pyplot as plt
import networkx as nx
import warnings
warnings.filterwarnings("ignore")
import json
# !git clone https://github.com/zlckanata/DeepGlobe-Road-Extraction-Challenge.git
# !mv DeepGlobe-Road-Extraction-Challenge DeepGlobe
# !git clone https://github.com/KshitijBhat/ML_Enthusiasts_Ishana.git
from ML_Enthusiasts_Ishana.Helpers import *
def get_path(image,t, image_path):
kernel = np.array([[0, -1, 0],
[-1, 5,-1],
[0, -1, 0]])
img_sharp = cv2.filter2D(image, ddepth=-1, kernel=kernel)
out = cv2.addWeighted( image, 1.1, img_sharp, 0.1, -5)
squares = split_stitch(out)
masks = []
for cut_img in squares:
mask = solver.test_one_img_from_path(cut_img)
mask[mask>0.7] = 255
mask[mask<=0.7] = 0
mask = np.concatenate([mask[:,:,None],mask[:,:,None],mask[:,:,None]],axis=2)
masks.append(mask.astype(np.uint8))
prediction = split_stitch(image,masks=masks)
points_path = image_path[0:len(image_path)-4] + ".json"
points = open(points_path)
coords = json.load(points)
start = np.array(coords["Start"])
end = np.array(coords["End"])
out = prediction
out = cv2.cvtColor(out,cv2.COLOR_BGR2GRAY)
grid = np.ceil((255-out)/255).astype('int')
edges = (cv2.Canny(out, threshold1=100, threshold2=200))/255
ngrid = narrowize(grid, edges,t)
G = initgraph(ngrid)
start = tuple(start[::-1])
end = tuple(end[::-1])
astar_path = generate_path(G,start,end,euclidean)
return astar_path
def submit():
ans = {}
for id in image_ids:
image = cv2.imread(source+id)
paths = get_path(image,4,source+id)
if paths == None:
paths = get_path(image,0,source+id)
path = []
for point in paths:
(x,y) = point
path.append([int(y),int(x)])
ans[id] = path
print(id,': Done')
json_object = json.dumps(ans, indent = 4)
with open("Submission.json", "w") as outfile:
outfile.write(json_object)
# + [markdown] id="QAvyOv-R8gOr"
# Upload the 60% images data and JSON files to a folder and fill in the path in the following cell . If it is in a github repo, clone it here and fill in the corresponding path. Example: `source = 'drive/MyDrive/Ishana/Data/'`
#
# JSON file name should be same as of the image.
#
# 60% data Folder structure:
# ```
# Data
# |_____Image1.json
# |_____Image1.png
# |_____Image2.json
# |_____Image2.png
# : ...
# : ...
#
# ```
#
#
#
# + id="IAeWN-ZF_Xbb"
source = '<insert path here>' #Like 'drive/MyDrive/Ishana/Data/'
val = os.listdir(source)
image_ids = list()
for filename in val:
if filename.endswith('.png'):
image_ids.append(filename)
print(image_ids)
# + [markdown] id="R67Kmwd_LWNL"
# Unzip the `weights.zip` attached in the folder named `ML_enthusiasts_Ishana` by running the following cell.
#
# Insert the path of the `weights.zip` file.
#
# Example: `model=drive/Shareddrives/ML_enthusiasts_Ishana/weights.zip`
# + id="TabWFhcyH7Di"
# !model=<insert path here> ; unzip $model
solver = TTAFrame(DinkNet34)
solver.load('log01_dink34.th')
# + [markdown] id="0WHzg_jTd-ER"
# To generate the JSON file for the final submission, run the following cell. The JSON file will be generated as `Submission.json` in the Files section. Download it for evaluation
# + id="WYJMqLRn9Hhy"
submit()
| Main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src= 'https://github.com/data-scince-explorer/data_vs_wild/blob/main/data_vs_wild.jpg?raw=true' width=500/>
# ### 1. Reading Data - with Pandas ###
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
sns.set() #import Seaborn functionalities
mushroom_df = pd.read_csv('data_vs_wild.csv')
mushroom_df.info()
mushroom_df.columns
# Let’s have a look at data dimensionality, features names, and feature types.
# Numbers of rows and columns
mushroom_df.shape
# Number of rows
len(mushroom_df)
# First 5 records of data
mushroom_df.head()
# Describe basic/statistical of data
mushroom_df.describe()
mushroom_df.describe().transpose() # change the rows and columns
mushroom_df.columns = [col.strip().replace('-', '_').lower() for col in mushroom_df.columns]
# Print the new columns
mushroom_df.columns
mushroom_df.head()
# +
# Open the mushroom properties details data
mushroom_prop_df=pd.read_csv('mushroom_features.csv')
mushroom_prop_df.head()
# -
mushroom_prop_df.info
mushroom_prop_df.shape
# replace "_" from mush_feature column of mushroom_prop_df dataframe
mushroom_prop_df=mushroom_prop_df.replace(to_replace ='-', value = '_', regex = True)
mushroom_prop_df.info
mushroom_prop_df.to_csv('new_mushroom_feature.csv')
new_mushroom_prop_df=pd.read_csv('new_mushroom_feature.csv')
new_mushroom_prop_df.info
mushroom_df['cap_color']
# ### Replace the "-" with "_" in the all columns to recognise in th code
# +
mushroom_df = mushroom_df.set_index('cap_shape')
mushroom_prop_df = mushroom_prop_df.set_index('m_code').rename( columns={'m_property':'cap_shape'} )
mushroom_prop_df.combine_first(mushroom_df)
# +
df = pd.DataFrame({'id1': [1001,1002,1001,1003,1004,1005,1002,1006],
'value1': ["a","b","c","d","e","f","g","h"],
'value3': ["yes","no","yes","no","no","no","yes","no"]})
dfReplace = pd.DataFrame({'id2': [1001,1002],
'value2': ["rep1","rep2"]})
print(df)
print()
print(dfReplace)
# +
df = df.set_index('id1')
#dfReplace = dfReplace.set_index('id2').rename( columns={'value2':'value1'} )
dfReplace = dfReplace.set_index('id2').rename( columns={'value2':'id1'} )
dfReplace.combine_first(df)
# +
# i=0
# for i in range(len(mushroom_df)):
# idx_prop=0
# for idx_prop in range(len(mushroom_prop_df)):
# if mushroom_df[i]
# -
# to see statistics on non-numerical features,
mushroom_df.describe(include=['object'])
# For categorical (type `object`) and boolean (type `bool`) features we can use the `value_counts` method. Let’s have a look at the distribution of `Churn`:
mushroom_df['safe'].value_counts() # to see how many mushrooms are not posioneous
# 753 users out of 247 are *loyal*; their `Fraud` value is `0`. To calculate fractions, pass `normalize=True` to the `value_counts` function.
mushroom_df['safe'].value_counts(normalize=True) # To see the fractions of mushrooms
(mushroom_df['safe'].value_counts().plot(
kind='bar',
figsize=(5, 20),
title='Distribution of Target Variable',
color='brown'
)
);
plt.show()
mushroom_df['safe'].mean()
mushroom_df.columns
#mushroom_df.iloc[0][1] #, 'cap_surface':'bruises']
mushroom_df['stalk_color_below_ring']
mushroom_df.iloc[0:5, 0:3]
import seaborn as sns
# sns.set_theme(style="whitegrid")
# tips = sns.load_dataset("tips")
# ax = sns.boxplot(x=tips["total_bill"])
ax = sns.boxplot(x="cap_surface", y="safe", data=mushroom_df)
ax = sns.boxplot(x="cap_surface", y="safe", hue="stalk_color_below_ring",
data=mushroom_df, palette="Set3")
# ax = sns.boxplot(x="cap_surface", y="safe", data=mushroom_df, order=["stalk_color_below_ring", "gill_spacing"])
ax = sns.stripplot(x="cap_surface", y="safe", data=mushroom_df)
# +
_, axes = plt.subplots(nrows=3, ncols=3, figsize=(15, 7))
sns.countplot(x='cap_surface', data=mushroom_df, ax=axes[0][0]);
sns.countplot(x='cap_color', data=mushroom_df, ax=axes[0][1]);
sns.countplot(x='bruises', data=mushroom_df, ax=axes[0][2]);
sns.countplot(x='odor', data=mushroom_df, ax=axes[1][0]);
sns.countplot(x='gill_attachment', data=mushroom_df, ax=axes[1][1]);
sns.countplot(x='gill_spacing', data=mushroom_df, ax=axes[1][2]);
sns.countplot(x='gill_size', data=mushroom_df, ax=axes[2][0]);
sns.countplot(x='gill_color', data=mushroom_df, ax=axes[2][1]);
sns.countplot(x='stalk_shape', data=mushroom_df, ax=axes[2][2]);
# -
mushroom_df.columns
# +
_, axes = plt.subplots(nrows=3, ncols=3, figsize=(15, 7))
sns.countplot(x='stalk_root', data=mushroom_df, ax=axes[0][0]);
sns.countplot(x='stalk_surface_above_ring', data=mushroom_df, ax=axes[0][1]);
sns.countplot(x='stalk_color_below_ring', data=mushroom_df, ax=axes[0][2]);
sns.countplot(x='veil_type', data=mushroom_df, ax=axes[1][0]);
sns.countplot(x='veil_color', data=mushroom_df, ax=axes[1][1]);
sns.countplot(x='ring_number', data=mushroom_df, ax=axes[1][2]);
sns.countplot(x='ring_type', data=mushroom_df, ax=axes[2][0]);
sns.countplot(x='spore_print_color', data=mushroom_df, ax=axes[2][1]);
sns.countplot(x='population', data=mushroom_df, ax=axes[2][2]);
# +
_, axes = plt.subplots(nrows=1, ncols=1, figsize=(15, 7))
sns.countplot(x='stalk_root', data=mushroom_df);
# -
# #### 1.2.3. Distributions of categorical features
mushroom_df.columns
# +
# Distributions of categorical features
plt.rcParams['figure.figsize'] = 8,6
sns.countplot(y='cap_surface', data=mushroom_df)
plt.show()
sns.countplot(y='cap_color', data=mushroom_df)
plt.show()
# -
# <h4>Pandas scatter matrix function helps visualize the relationship between features</h4>
# Use with care though, because it is processor intensive
# +
from pandas.plotting import scatter_matrix
p=scatter_matrix(mushroom_df, alpha=0.2, figsize=(30, 20), diagonal='kde')
# -
# Box and Whisker Plots
import matplotlib.pyplot as plt
plt.rcParams['figure.figsize'] = 20,20 # control plot size
mushroom_df.plot(kind='box', subplots=True, layout=(6,6), sharex=False, sharey=False)
plt.show()
| data_vs_wild.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Converting from OneMap Singapore's [X Y coordinates](https://www.onemap.gov.sg/docs/#3414-svy21-to-3857) in 3414(SVY21) format to latitude and longitude in 4326(WGS84) format
#
# Information loads very quickly when the url is opened but the information took very long to be retrieved using the requests.get function. Hence instead of using the OneMap API to convert the coordinates, here's a much quicker alternative.
import webbrowser
url = "https://developers.onemap.sg/commonapi/convert/3414to4326?X=28983.788791079794&Y=33554.5098132845"
webbrowser.open(url)
# ## Using requests.get
from urllib.request import Request, urlopen
import requests
from bs4 import BeautifulSoup
import time
hdr = {'User-Agent': 'Mozilla/5.0'}
x_list = [28983.78879107979]
y_list = [33554.5098132845]
for i in range(0,1):
url = "https://developers.onemap.sg/commonapi/convert/3414to4326?X={}4&Y={}".format(x_list[i],y_list[i])
start = time.time()
print('retrieving...')
bookpage = requests.get(url, headers=hdr)
end = time.time()
soup = BeautifulSoup(bookpage.text, "html.parser")
print(end - start)
soup
list_latlong = []
list_latlong.append((x,y,soup))
list_latlong
# ## Using pyproj - better alternative
import pyproj
from pyproj import Transformer
import pyproj
p = pyproj.CRS("epsg:3414")
p_to = pyproj.CRS("epsg:4326")
# ### Applying to a list
def xy_to_lonlat(x, y):
transformer = pyproj.Transformer.from_crs(p, p_to, always_xy=True)
lonlat = transformer.transform(x, y)
return lonlat[0], lonlat[1]
x_list = [28983.78879107979]
y_list = [33554.5098132845]
for i in range(0,1):
lon, lat = xy_to_lonlat(x_list[i],y_list[i])
print(lat,lon)
# ### Applying to a pandas dataframe
def xy_to_lonlat(x, y):
transformer = pyproj.Transformer.from_crs(p, p_to, always_xy=True)
lonlat = transformer.transform(x, y)
return lonlat#lonlat[0], lonlat[1]
import warnings
warnings.filterwarnings("ignore")
df["lonlat"] = df.apply(lambda x: xy_to_lonlat(x.x_coord, x.y_coord), axis=1)
df['lonlat']= df['lonlat'].astype(str)
df[['longitude','latitude']] = df['lonlat'].str.split(',',expand=True)
df['longitude'] = df['longitude'].str.replace('(','')
df['latitude'] = df['latitude'].str.replace(')','')
# References:
# https://github.com/pyproj4/pyproj/issues/537
| notebooks/OneMapSG_XY_LatLon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="TzlyU5nrvpWp"
# ## **Arrow Detection with Direction**
# + [markdown] colab_type="text" id="YLjAfhqxxOPq"
# Import Libraries
# + colab={} colab_type="code" id="mXfHdocv7pq7"
import numpy as np
import matplotlib.pyplot as plt
from PIL import Image
from keras import Sequential
from keras.layers import Dense
from keras.models import Model
from keras.layers import Conv2D, MaxPool2D, \
Dropout, Dense, Input, concatenate, \
GlobalAveragePooling2D, AveragePooling2D,\
Flatten, Conv2DTranspose
import cv2
import numpy as np
import math
print(tf.__version__)
# + [markdown] colab_type="text" id="eObs1ZkFyjUF"
# Define Network
# + colab={} colab_type="code" id="SlLUP1wf-bPH"
in_size = 16
x = keras.layers.Input(shape=(160, 160, 3))
gen1 = keras.layers.Conv2D(8, (4, 4), padding = 'SAME', activation = 'relu')(x)
P1 = keras.layers.MaxPooling2D(2, padding = 'SAME')(gen1)
gen2 = keras.layers.Conv2D(16, (4, 4), padding = 'SAME', activation = 'relu')(P1)
P2 = keras.layers.MaxPooling2D(4, padding = 'SAME')(gen2)
gen3 = keras.layers.Conv2D(32, (4, 4), padding = 'SAME', activation = 'relu')(P2)
P3 = keras.layers.MaxPooling2D(2, padding = 'SAME')(gen3)
gen4 = keras.layers.Conv2D(16, (4, 4), padding = 'SAME', activation = 'relu')(P3)
P4 = keras.layers.MaxPooling2D(4, padding = 'SAME')(gen4)
gen4 = keras.layers.Conv2D(8, (4, 4), padding = 'SAME', activation = 'sigmoid')(P4)
P4 = keras.layers.MaxPooling2D(2, padding = 'SAME')(gen4)
x4 = keras.layers.Flatten()(P4)
x5 = keras.layers.Dense(3, activation = 'softmax')(x4)
model = keras.models.Model([x], x5)
model.summary()
model.compile(optimizer='adam', loss='sparse_categorical_crossentropy', metrics=['accuracy'])
# + [markdown] colab_type="text" id="XMMWheM3yo_i"
# Load pretrained model
# -
model = keras.models.load_model('/home/aashish/Downloads/image_upscaling/edge_detection/Arrow224/Arrow224/arrow.h5')
# + [markdown] colab_type="text" id="7jzFIS9Oyr2p"
# Test Webcam footage
# + colab={} colab_type="code" id="KiEe3OqQ7prj"
#img = Image.open('/home/aashish/Downloads/image_upscaling/edge_detection/Arrow224/Arrow224/test/left/L8.jpg')
video_capture = cv2.VideoCapture(0)
while True:
ret, frame = video_capture.read()
#img.load
#plt.imshow(img)
#plt.show()
img = frame
#img = img.resize((160, 160), Image.ANTIALIAS)
npimg = np.asarray(img, dtype="uint8" )
print('shape', np.shape(npimg))
test_image = np.resize(npimg, (1, 160, 160, 3))
print('shape', np.shape(test_image))
predictions = model.predict(test_image)
print(predictions)
classes = ['L', 'R', 'N']
print(classes[np.argmax(predictions)])
font = cv2.FONT_HERSHEY_SIMPLEX
org = (50, 50)
fontScale = 1
color = (255, 0, 0)
thickness = 2
cv2.putText(frame,classes[np.argmax(predictions)], org, font, fontScale, color, thickness, cv2.LINE_AA)
cv2.imshow('Frame', frame)
if cv2.waitKey(1) & 0xFF == ord('q'):
break
video_capture.release()
cv2.destroyAllWindows()
# -
| Arrow-Detection/Arrow_detect_run.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: C#
# language: csharp
# name: csharp
# ---
# 
# ## Welcome to The QuantConnect Research Page
# #### Refer to this page for documentation https://www.quantconnect.com/docs#Introduction-to-Jupyter
# #### Contribute to this template file https://github.com/QuantConnect/Lean/blob/master/Jupyter/BasicCSharpQuantBookTemplate.ipynb
# ## QuantBook Basics
# The following example is ready to be used in our Docker container, reference the ReadMe for more details on setting this up.
#
#
#
# In order to use this notebook locally you will need to make a few small changes:
#
# 1. Either create the notebook in your build folder (`bin/debug`) **or** set working directory of the notebook to it like so in the first cell:
#
# ```Directory.SetCurrentDirectory("PathToLean/Lean/Launcher/bin/Debug/");```
#
# 2. Load "QuantConnect.csx" instead of "../QuantConnect.csx", this is again because of the Notebook position relative to the build files.
#
# ### Start QuantBook
# - Load "QuantConnect.csx" with all the basic imports
# - Create a QuantBook instance
#load "../QuantConnect.csx"
var qb = new QuantBook();
# ### Using the Web API
# Our script `QuantConnect.csx` automatically loads an instance of the web API for you to use.**
#
# Look at Lean's [Api](https://github.com/QuantConnect/Lean/tree/master/Api) class for more functions to interact with the cloud
#
#
# ##### **Note: This will only connect if you have your User ID and Api token in `config.json`
// Show that our api object is connected to the Web Api
Console.WriteLine(api.Connected);
// Get our list of projects from the cloud and print their names
var projectResponse = api.ListProjects();
foreach (var project in projectResponse.Projects) {
Console.WriteLine(project.Name);
}
# ### Selecting Asset Data
# Checkout the QuantConnect [docs](https://www.quantconnect.com/docs#Initializing-Algorithms-Selecting-Asset-Data) to learn how to select asset data.
var spy = qb.AddEquity("SPY");
var eur = qb.AddForex("EURUSD");
var btc = qb.AddCrypto("BTCUSD");
var fxv = qb.AddData<FxcmVolume>("EURUSD_Vol", Resolution.Hour);
# ### Historical Data Requests
#
# We can use the QuantConnect API to make Historical Data Requests. The data will be presented as multi-index pandas.DataFrame where the first index is the Symbol.
#
# For more information, please follow the [link](https://www.quantconnect.com/docs#Historical-Data-Historical-Data-Requests).
// Gets historical data from the subscribed assets, the last 360 datapoints with daily resolution
var h1 = qb.History(qb.Securities.Keys, 360, Resolution.Daily);
// Gets historical data from the subscribed assets, from the last 30 days with daily resolution
var h2 = qb.History(qb.Securities.Keys, TimeSpan.FromDays(360), Resolution.Daily);
// Gets historical data from the subscribed assets, between two dates with daily resolution
var h3 = qb.History(btc.Symbol, new DateTime(2014,1,1), DateTime.Now, Resolution.Daily);
// Only fetchs historical data from a desired symbol
var h4 = qb.History(spy.Symbol, 360, Resolution.Daily);
// Only fetchs historical data from a desired symbol
var h5 = qb.History<QuoteBar>(eur.Symbol, TimeSpan.FromDays(360), Resolution.Daily);
// Fetchs custom data
var h6 = qb.History<FxcmVolume>(fxv.Symbol, TimeSpan.FromDays(360));
| Research/KitchenSinkCSharpQuantBookTemplate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: eclipse
# language: python
# name: eclipse
# ---
# Import all libs
import matplotlib.pyplot as plt
plt.style.use(["science", "ieee"])
import datetime as dt
import sys
sys.path.extend(["code/", "code/rt/", "code/sd/"])
from models import plotting_electron_density, extract_modeled_parameters
from plots import *
dat, H, T = extract_modeled_parameters(dstart=dt.datetime(2017,8,21,15), dend=dt.datetime(2017,8,21,21,5),
latlon = [40.015,-105.2705], p_time = dt.datetime(2017,8,21,17,46),
params=["WI"], base="dataset/parsed/%s_parsed_wi.pickle")
np.max(dat["WI"]), np.min(dat["WI"])
plot_params(H, T, dat["WI"], dt.datetime(2017,8,21,17,46), fname="data/%s/Boulder_v.density_{d}D.png",
location="Boulder", pname="WI", vlim=[0, 31], dlim=3)
# Import all libs
import matplotlib.pyplot as plt
plt.style.use(["science", "ieee"])
import datetime as dt
import sys
sys.path.extend(["code/", "code/rt/", "code/sd/"])
from models import plotting_electron_density, extract_modeled_parameters
from plots import *
dat, H, T = extract_modeled_parameters(dstart=dt.datetime(2017,8,21,15), dend=dt.datetime(2017,8,21,21,5),
latlon = [40.015,-105.2705], p_time = dt.datetime(2017,8,21,17,46),
params=["U", "V"], base="dataset/parsed/%s_parsed_uv.pickle")
np.max(dat["U"]), np.min(dat["U"]), np.max(dat["V"]), np.min(dat["V"])
plot_params(H, T, dat["U"], dt.datetime(2017,8,21,17,46), fname="data/%s/Boulder_v.density_{d}D.png",
location="Boulder", pname="U", vlim=[10, 100], dlim=10)
plot_params(H, T, dat["V"], dt.datetime(2017,8,21,17,46), fname="data/%s/Boulder_v.density_{d}D.png",
location="Boulder", pname="V", vlim=[0, 200], dlim=3)
| Analysis.uvw.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Basic Operations
#
# This lecture will cover some basic operations with Spark DataFrames.
#
# We will play around with some stock data from Apple.
from pyspark.sql import SparkSession
# + jupyter={"outputs_hidden": true}
# May take awhile locally
spark = SparkSession.builder.appName("Operations").getOrCreate()
# + jupyter={"outputs_hidden": false}
# Let Spark know about the header and infer the Schema types!
df = spark.read.csv('appl_stock.csv',inferSchema=True,header=True)
# + jupyter={"outputs_hidden": false}
df.printSchema()
# -
# ## Filtering Data
#
# A large part of working with DataFrames is the ability to quickly filter out data based on conditions. Spark DataFrames are built on top of the Spark SQL platform, which means that is you already know SQL, you can quickly and easily grab that data using SQL commands, or using the DataFram methods (which is what we focus on in this course).
# + jupyter={"outputs_hidden": false}
# Using SQL
df.filter("Close<500").show()
# + jupyter={"outputs_hidden": false}
# Using SQL with .select()
df.filter("Close<500").select('Open').show()
# + jupyter={"outputs_hidden": false}
# Using SQL with .select()
df.filter("Close<500").select(['Open','Close']).show()
# -
# Using normal python comparison operators is another way to do this, they will look very similar to SQL operators, except you need to make sure you are calling the entire column within the dataframe, using the format: df["column name"]
#
# Let's see some examples:
# + jupyter={"outputs_hidden": false}
df.filter(df["Close"] < 200).show()
# + jupyter={"outputs_hidden": false}
# Will produce an error, make sure to read the error!
df.filter(df["Close"] < 200 and df['Open'] > 200).show()
# + jupyter={"outputs_hidden": false}
# Make sure to add in the parenthesis separating the statements!
df.filter( (df["Close"] < 200) & (df['Open'] > 200) ).show()
# + jupyter={"outputs_hidden": false}
# Make sure to add in the parenthesis separating the statements!
df.filter( (df["Close"] < 200) | (df['Open'] > 200) ).show()
# + jupyter={"outputs_hidden": false}
# Make sure to add in the parenthesis separating the statements!
df.filter( (df["Close"] < 200) & ~(df['Open'] < 200) ).show()
# + jupyter={"outputs_hidden": false}
df.filter(df["Low"] == 197.16).show()
# + jupyter={"outputs_hidden": false}
# Collecting results as Python objects
df.filter(df["Low"] == 197.16).collect()
# + jupyter={"outputs_hidden": true}
result = df.filter(df["Low"] == 197.16).collect()
# + jupyter={"outputs_hidden": false}
# Note the nested structure returns a nested row object
type(result[0])
# + jupyter={"outputs_hidden": true}
row = result[0]
# -
# Rows can be called to turn into dictionaries
# + jupyter={"outputs_hidden": false}
row.asDict()
# + jupyter={"outputs_hidden": false}
for item in result[0]:
print(item)
# -
# That is all for now Great Job!
| Python-and-Spark-for-Big-Data-master/Spark_DataFrames/DataFrame_Basic_Operations.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
# This line will include DEBUG level log messages from the start of the app,
# as well as include a "Local server" as provider (at http://localhost:5000/optimade/v<MAJOR>),
# where <MAJOR> is the major version number of the currently supported OPTIMADE spec version.
#os.environ["OPTIMADE_CLIENT_DEBUG"] = "True"
# +
from optimade_client import (
HeaderDescription,
OptimadeClientFAQ,
OptimadeLog,
OptimadeQueryProviderWidget,
OptimadeQueryFilterWidget,
OptimadeSummaryWidget,
)
from ipywidgets import dlink, HTML
from IPython.display import display
from tools_optimade_client import QEInputButton
disable_providers = [
"cod",
"tcod",
"nmd",
"oqmd",
"aflow",
"matcloud",
"mpds",
"necro",
"jarvis",
]
skip_databases = {"Materials Cloud": ["optimade-sample", "li-ion-conductors"]}
database_grouping = {
"Materials Cloud": {
"General": ["curated-cofs"],
"Projects": [
"2dstructures",
"2dtopo",
"pyrene-mofs",
"scdm",
"sssp",
"stoceriaitf",
"tc-applicability",
"threedd",
]}
}
selector = OptimadeQueryProviderWidget(
width_ratio=(38, 51),
database_limit=50,
disable_providers=disable_providers,
skip_databases=skip_databases,
provider_database_groupings=database_grouping,
)
filters = OptimadeQueryFilterWidget(
button_style='primary',
result_limit=25,
subparts_order=[
"filter_header",
"filters",
"query_button",
"structures_header",
"sort_selector",
"structure_page_chooser",
"structure_drop",
"error_or_status_messages",
],
)
summary = OptimadeSummaryWidget(direction='horizontal', button_style='info')
qe_input_generator_button = QEInputButton(button_style="info")
_ = dlink((selector, 'database'), (filters, 'database'))
_ = dlink((filters, 'structure'), (summary, 'entity'))
_ = dlink((filters, 'structure'), (qe_input_generator_button, 'structure'))
HeaderDescription(button_style='info')
# -
OptimadeClientFAQ()
OptimadeLog()
# +
display(HTML('<h2 style="margin-below:0px;padding-below:0px;">Query a provider\'s database</h2>'))
display(selector, filters, summary)
display(qe_input_generator_button)
| OPTIMADE-Client.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Understanding lists and manipulating lines
#
# By [<NAME>](http://www.decontextualize.com/)
#
# In this tutorial, I explain how the list data structure works in Python. After going over the basics, I'll show you how to use list comprehensions as a powerful and succinct method to poetically manipulate lines and words from a text.
#
# ## Lists: the basics
#
# A list is a type of value in Python that represents a sequence of values. The list is a very common and versatile data structure in Python and is used frequently to represent (among other things) tabular data and words in a text. Here's how you write one out in Python:
[5, 10, 15, 20, 25, 30]
# That is: a left square bracket, followed by a series of comma-separated expressions, followed by a right square bracket. Items in a list don't have to be values; they can be more complex expressions as well. Python will evaluate those expressions and put them in the list.
[5, 2*5, 3*5, 4*5, 5*5, 6*5]
# Lists can have an arbitrary number of values. Here's a list with only one value in it:
[5]
# And here's a list with no values in it:
[]
# Here's what happens when we ask Python what type of value a list is:
type([1, 2, 3])
# It's a value of type `list`.
#
# Like any other kind of Python value, you can assign a list to a variable:
my_numbers = [5, 10, 15, 20, 25, 30]
# ### Getting values out of lists
#
# Once we have a list, we might want to get values *out* of the list. You can write a Python expression that evaluates to a particular value in a list using square brackets to the right of your list, with a number representing which value you want, numbered from the beginning (the left-hand side) of the list. Here's an example:
[5, 10, 15, 20][2]
# If we were to say this expression out loud, it might read, "I have a list of four things: 5, 10, 15, 20. Give me back the second item in the list." Python evaluates that expression to `15`, the second item in the list.
#
# Here's what it looks like to use this indexing notation on a list stored in a variable:
my_numbers[2]
# #### The second item? Am I seeing things. 15 is clearly the third item in the list.
#
# You're right---good catch. But for reasons too complicated to go into here, Python (along with many other programming languages!) starts list indexes at 0, instead of 1. So what looks like the third element of the list to human eyes is actually the second element to Python. The first element of the list is accessed using index 0, like so:
[5, 10, 15, 20][0]
# The way I like to conceptualize this is to think of list indexes not as specifying the number of the item you want, but instead specifying how "far away" from the beginning of the list to look for that value.
#
# If you attempt to use a value for the index of a list that is beyond the end of the list (i.e., the value you use is higher than the last index in the list), Python gives you an error:
my_numbers[47]
# Note that while the type of a list is `list`, the type of an expression using index brackets to get an item out of the list is the type of whatever was in the list to begin with. To illustrate:
type(my_numbers)
type(my_numbers[0])
# #### Indexes can be expressions too
#
# The thing that goes inside of the index brackets doesn't have to be a number that you've just typed in there. Any Python expression that evaluates to an integer can go in there.
my_numbers[2 * 2]
x = 3
[5, 10, 15, 20][x]
# ### Other operations on lists
#
# Because lists are so central to Python programming, Python includes a number of built-in functions that allow us to write expressions that evaluate to interesting facts about lists. For example, try putting a list between the parentheses of the `len()` function. It will evaluate to the number of items in the list:
len(my_numbers)
len([20])
len([])
# The `in` operator checks to see if the value on the left-hand side is in the list on the right-hand side.
3 in my_numbers
15 in my_numbers
# The `max()` function will evaluate to the highest value in the list:
readings = [9, 8, 42, 3, -17, 2]
max(readings)
# ... and the `min()` function will evaluate to the lowest value in the list:
min(readings)
# The `sum()` function evaluates to the sum of all values in the list.
sum([2, 4, 6, 8, 80])
# Finally, the `sorted()` function evaluates to a copy of the list, sorted from smallest value to largest value:
sorted(readings)
# ### Negative indexes
#
# If you use `-1` as the value inside of the brackets, something interesting happens:
fib = [1, 1, 2, 3, 5]
fib[-1]
# The expression evaluates to the *last* item in the list. This is essentially the same thing as the following code:
fib[len(fib) - 1]
# ... except easier to write. In fact, you can use any negative integer in the index brackets, and Python will count that many items from the end of the list, and evaluate the expression to that item.
fib[-3]
# If the value in the brackets would "go past" the beginning of the list, Python will raise an error:
fib[-14]
# ### Generating lists with `range()`
# The expression `list(range(n))` returns a list from 0 up to (but not including) `n`. This is helpful when you just want numbers in a sequence:
list(range(10))
# You can specify where the list should start and end by supplying two parameters to the call to `range`:
list(range(-10, 10))
# ## List slices
#
# The index bracket syntax explained above allows you to write an expression that evaluates to a particular item in a list, based on its position in the list. Python also has a powerful way for you to write expressions that return a *section* of a list, starting from a particular index and ending with another index. In Python parlance we'll call this section a *slice*.
#
# Writing an expression to get a slice of a list looks a lot like writing an expression to get a single value. The difference is that instead of putting one number between square brackets, we put *two* numbers, separated by a colon. The first number tells Python where to begin the slice, and the second number tells Python where to end it.
[4, 5, 6, 10, 12, 15][1:4]
# Note that the value after the colon specifies at which index the slice should end, but the slice does *not* include the value at that index. (You can tell how long the slice will be by subtracting the value before the colon from the value after it.)
#
# Also note that---as always!---any expression that evaluates to an integer can be used for either value in the brackets. For example:
x = 3
[4, 5, 6, 10, 12, 15][x:x+2]
# Finally, note that the type of a slice is `list`:
type(my_numbers)
type(my_numbers[1:4])
# ### Omitting slice values
#
# Because it's so common to use the slice syntax to get a list that is either a slice starting at the beginning of the list or a slice ending at the end of the list, Python has a special shortcut. Instead of writing:
my_numbers[0:3]
# You can leave out the `0` and write this instead:
my_numbers[:3]
# Likewise, if you wanted a slice that starts at index 4 and goes to the end of the list, you might write:
my_numbers[4:]
# Getting the last two items in `my_numbers`:
my_numbers[:2]
# ### Negative index values in slices
#
# Now for some tricky stuff: You can use negative index values in slice brackets as well! For example, to get a slice of a list from the fourth-to-last element of the list up to (but not including) the second-to-last element of the list:
my_numbers[-4:-2]
# To get the last three elements of the list:
my_numbers[:-3]
# All items from `my_numbers` from the third item from the end of the list upto the end of the list:
my_numbers[-3:]
# ### Strings and lists
#
# Strings and lists share a lot of similarities! The same square bracket slice and index syntax works on strings the same way it works on lists:
message = "importantly"
message[1]
message[-2]
message[-5:-2]
# Weirdly, `max()` and `min()` also work on strings... they just evaluate to the letter that comes latest and earliest in alphabetical order (respectively):
max(message)
min(message)
# You can turn a string into a list of its component characters by passing it to `list()`:
list(message)
list("我爱猫!😻")
# The letters in a string in alphabetical order:
sorted(list(message))
# ## List comprehensions: Applying transformations to lists
#
# A very common task in both data analysis and computer programming is applying some operation to every item in a list (e.g., scaling the numbers in a list by a fixed factor), or to create a copy of a list with only those items that match a particular criterion (e.g., eliminating values that fall below a certain threshold). Python has a succinct syntax, called a *list comprehension*, which allows you to easily write expressions that transform and filter lists.
#
# A list comprehension has a few parts:
#
# - a *source list*, or the list whose values will be transformed or filtered;
# - a *predicate expression*, to be evaluated for every item in the list;
# - (optionally) a *membership expression* that determines whether or not an item in the source list will be included in the result of evaluating the list comprehension, based on whether the expression evaluates to `True` or `False`; and
# - a *temporary variable name* by which each value from the source list will be known in the predicate expression and membership expression.
#
# These parts are arranged like so:
#
# > `[` *predicate expression* `for` *temporary variable name* `in` *source list* `if` *membership expression* `]`
#
# The words `for`, `in`, and `if` are a part of the syntax of the expression. They don't mean anything in particular (and in fact, they do completely different things in other parts of the Python language). You just have to spell them right and put them in the right place in order for the list comprehension to work.
#
# Here's an example, returning the squares of integers zero up to ten. First, we'll create a list `to_ten` that contains the range of numbers:
to_ten = list(range(10))
to_ten
[x * x for x in to_ten]
# In the example above, `x*x` is the predicate expression; `x` is the temporary variable name; and `[0, 1, 2, 3, 4, 5, 6, 7, 8, 9]` is the source list. There's no membership expression in this example, so we omit it (and the word `if`).
#
# There's nothing special about the variable `x`; it's just a name that we chose. We could easily choose any other temporary variable name, as long as we use it in the predicate expression as well. Below, I use the name of one of my cats as the temporary variable name, and the expression evaluates the same way it did with `x`:
[shumai * shumai for shumai in to_ten]
# Notice that the type of the value that a list comprehension evaluates to is itself type `list`:
type([x * x for x in to_ten])
# The `source` doesn't have to be a variable that contains a list. It can be *any expression that evaluates to a list*, or in fact any expression that evaluates to an [iterable](https://docs.python.org/3/glossary.html#term-iterable). For example:
[x * x for x in [0, -1, 1, -2, 2, -3, 3]]
# ... or:
[x * x for x in range(10)]
# We've used the expression `x * x` as the predicate expression in the examples above, but you can use any expression you want. For example, to scale the values of a list by 0.5:
[x * 0.5 for x in range(10)]
# In fact, the expression in the list comprehension can just be the temporary variable itself, in which case the list comprehension will simply evaluate to a copy of the original list:
[x for x in range(10)]
# You don't technically even need to use the temporary variable in the predicate expression:
[42 for x in range(10)]
# > Bonus exercise: Write a list comprehension for the list `range(5)` that evaluates to a list where every value has been multiplied by two (i.e., the expression would evaluate to `[0, 2, 4, 6, 8]`).
# ### The membership expression
#
# As indicated above, you can include an expression at the end of the list comprehension to determine whether or not the item in the source list will be evaluated and included in the resulting list. One way, for example, of including only those values from the source list that are greater than or equal to five:
[x*x for x in range(10) if x >= 5]
# ## Splitting strings into lists
#
# The `split()` method is a funny thing you can do with a string to transform it into a list. If you have an expression that evaluates to a string, you can put `.split()` right after it, and Python will evaluate the whole expression to mean "take this string, and 'split' it on white space, giving me a list of strings with the remaining parts." For example:
"this is a test".split()
# Notably, while the `type` of a string is `str`, the type of the result of `split()` is `list`:
type("this is a test".split())
# If the string in question has some delimiter in it other than whitespace that we want to use to separate the fields in the resulting list, we can put a string with that delimiter inside the parentheses of the `split()` method. Maybe you can tell where I'm going with this at this point!
#
# ### From string to list of numbers: an example
#
# For example, I happen to have here a string that represents the total points scored by <NAME> in each of his NBA games in the 2013-2014 regular season.
#
# > 17,25,26,25,35,18,25,33,39,30,13,21,22,35,28,27,26,23,21,21,24,17,25,30,24,18,38,19,33,26,26,15,30,32,32,36,25,21,34,30,29,27,18,34,30,24,31,13,37,36,42,33,31,20,61,22,19,17,23,19,21,24,43,15,25,32,38,17,13,32,17,34,38,29,37,36,27
#
# You can either cut-and-paste this string from the notes, or see a file on github with these values [here](https://gist.githubusercontent.com/aparrish/56ea528159c97b085a34/raw/8406bdd866101cf64347349558d9a806c82aceb7/scores.txt).
#
# Now if I just cut-and-pasted this string into a variable and tried to call list functions on it, I wouldn't get very helpful responses:
raw_str = "17,25,26,25,35,18,25,33,39,30,13,21,22,35,28,27,26,23,21,21,24,17,25,30,24,18,38,19,33,26,26,15,30,32,32,36,25,21,34,30,29,27,18,34,30,24,31,13,37,36,42,33,31,20,61,22,19,17,23,19,21,24,43,15,25,32,38,17,13,32,17,34,38,29,37,36,27"
max(raw_str)
# This is wrong—we know that <NAME> scored more than nine points in his highest scoring game. The `max()` function clearly does strange things when we give it a string instead of a list. The reason for this is that all Python knows about a string is that it's a *series of characters*. It's easy for a human to look at this string and think, "Hey, that's a list of numbers!" But Python doesn't know that. We have to explicitly "translate" that string into the kind of data we want Python to treat it as.
#
# > Bonus advanced exercise: Take a guess as to why, specifically, Python evaluates `max(raw_str)` to `9`. Hint: what's the result of `type(max(raw_str))`?
#
# What we want to do, then, is find some way to convert this string that *represents* integer values into an actual Python list of integer values. We'll start by splitting this string into a list, using the `split()` method, passing `","` as a parameter so it splits on commas instead of on whitespace:
str_list = raw_str.split(",")
str_list
# Looks good so far. What does `max()` have to say about it?
max(str_list)
# This.. works. (But only by accident—see below.) But what if we wanted to find the total number of points scored by LBJ? We should be able to do something like this:
sum(str_list)
# ... but we get an error. Why this error? The reason lies in what kind of data is in our list. We can check the data type of an element of the list with the `type()` function:
type(str_list[0])
# A-ha! The type is `str`. So the error message we got before (`unsupported operand type(s) for +: 'int' and 'str'`) is Python's way of telling us, "You gave me a list of strings and then asked me to add them all together. I'm not sure what I can do for you."
#
# So there's one step left in our process of "converting" our "raw" string, consisting of comma-separated numbers, into a list of numbers. What we have is a list of strings; what we want is a list of numbers. Fortunately, we know how to write an expression to transform one list into another list, applying an expression to each member of the list along the way—it's called a list comprehension. Equally fortunately, we know how to write an expression that converts a string representing an integer into an actual integer (`int()`). Here's how to write that expression:
[int(x) for x in str_list]
# Let's double-check that the values in this list are, in fact, integers, by spot-checking the first item in the list:
type([int(x) for x in str_list][0])
# Hey, voila! Now we'll assign that list to a variable, for the sake of convenience, and then check to see if `sum()` works how we expect it to.
int_list = [int(x) for x in str_list]
sum(int_list)
# Wow! 2089 points in one season! Good work, <NAME>.
# ## Join: Making strings from lists
#
# Once we've created a list of words, it's a common task to want to take that
# list and "glue" it back together, so it's a single string again, instead of
# a list. So, for example:
element_list = ["hydrogen", "helium", "lithium", "beryllium", "boron"]
glue = ", and "
glue.join(element_list)
# The `.join()` method needs a "glue" string to the left of it---this is the
# string that will be placed in between the list elements. In the parentheses
# to the right, you need to put an expression that evaluates to a list. Very
# frequently with `.join()`, programmers don't bother to assign the "glue"
# string to a variable first, so you end up with code that looks like this:
words = ["this", "is", "a", "test"]
" ".join(words)
# When we're working with `.split()` and `.join()`, our workflow usually looks
# something like this:
#
# 1. Split a string to get a list of units (usually words).
# 2. Use some of the list operations discussed above to modify or slice the list.
# 3. Join that list back together into a string.
# 4. Do something with that string (e.g., print it out).
#
# With this in mind, here's a program that splits a string into words, randomizes the order of the words, then prints out the results:
text = "it was a dark and stormy night"
words = text.split()
random.shuffle(words)
' '.join(words)
# ## Lists and randomness
#
# Python's `random` library provides several helpful functions for performing
# chance operations on lists. The first is `shuffle`, which takes a list and
# randomly shuffles its contents:
import random
ingredients = ["flour", "milk", "eggs", "sugar"]
random.shuffle(ingredients)
ingredients
# The second is `choice`, which returns a single random element from list.
import random
ingredients = ["flour", "milk", "eggs", "sugar"]
random.choice(ingredients)
# Finally, the `sample` function returns a list of values, selected at random,
# from a list. The `sample` function takes two parameters: the first is a list,
# and the second is how many items should be in the resulting list of randomly
# selected values:
import random
ingredients = ["flour", "milk", "eggs", "sugar"]
random.sample(ingredients, 2)
# ## Text files and lists of lines
#
# The `open()` function allows you to read text from a file. When used as the source in a list comprehension, the predicate expression will be evaluated for *each line* of text in the file. For example:
[line for line in open("sea_rose.txt")]
# What we have here is a *list of strings*. Each element in the list corresponds to a single line from the text file.
#
# Notice that you see the `\n` character at the end of each string; this is Python letting you know that there's a character in the file that indicates where the linebreaks should occur. By default, Python doesn't strip this character out. To make our data a little bit cleaner, let's use the `.strip()` method in the predicate expression to get rid of the newline character (and any other accompanying whitespace characters):
[line.strip() for line in open("sea_rose.txt")]
# Assigning this to a variable, we can do things like get the lines of the poem from index 3 up to index 9:
poem = [line.strip() for line in open("sea_rose.txt")]
poem[3:9]
# Or we can get a few lines at random:
random.sample(poem, 3)
# Or sort the poem in alphabetical order:
sorted(poem)
# The resulting list variable can itself be used as the source expression in other list comprehensions:
[line[:5] for line in poem]
# ### Transforming lines of text
#
# Wait, how did I do that thing with that poem, where the letters are all weird? It's like this weird pomo l=a=n=g=u=a=g=e poetry now. Completely unpublishable, I'll get kicked right out of the Iowa Writer's Workshop. Very cool.
#
# It turns out you can make changes to the predicate expression in order to make changes to the way the text looks in the output. We're modifying the text by transforming the strings. There are a handful of really easy things you can do to strings of characters in Python to make the text do weird and interesting things. I'm going to show you a few.
#
# First, I'm going to make a variable called `poem` and assign to it the result of reading in that Robert Frost poem. The road one where he decides to take a road, but not another road, and it is very momentous.
poem = [line.strip() for line in open("frost.txt")]
# What we want to do now is write a list comprehension that transforms the lines in the poem somehow. The key to this is to change the predicate expression in a list comprehension. The simplest possible text transformation is nothing at all: just make a new list that looks like the old list in every way.
[line for line in poem]
# That list comprehension basically translates to the following: "Hey python, hey yes you, python! Take a look at the list of strings in the variable poem that I defined earlier. I want you to make a new list, and here's how that new list should look: for every item of that list—let's call the item line—put an item with whatever that line is into the new list." Python: "So, uh, make a copy of the list?" You: "Yeah I guess basically."
#
# Another simple transformation is to not do anything with the data in the line at all, and have Python put another string altogether into the new list:
["I'm <NAME>, howdy howdy howdy" for line in poem]
# Neither of these are interesting from anything other than a theoretical perspective, which means if you're a humanities scholar or something you can just stop here and start typing up your monograph on conceptions of identity and iteration in algorithmic media. But as for me and my students, we are ARTISTS and ENGINEERS and it's important that we CHANGE THE WORLD and SHOW RESULTS.
# ### String methods in the predicate expression
#
# Recall from the tutorial on strings that string expressions in Python have a number of methods that can be called on them that return a copy of the string with some transformation applied, like `.lower()` (converts the string to lower case) or `.replace()` (replaces matching substrings with some other string). We can use these *in the predicate expression* to effect that transformation on every item in the list. To make every string in the upper case, for example, call the .upper() method on the temporary variable line. This makes Frost look really mad:
[line.upper() for line in poem]
# And `.replace()` is a fun one. You need to put two comma-separated strings between the parentheses. Python will replace every occurrence of the first string with the second string. To make our poem more colloquial, for example:
[line.replace("I", "my dude") for line in poem]
# If you have ever wondered, "What would this roady poem sound like if you tickled <NAME> while he read it aloud," then you're in luck because Python has answered that question.
[line.replace("a", "aheHEEhaHAha") for line in poem]
# The `.strip()` method is helpful! We used it above to strip off whitespace, but if you give it a string as a parameter (inside the parentheses), it will remove all of the characters inside that string from the beginning and end of every line. This is a convenient way to, e.g., remove punctuation from the ends of lines:
[line.strip(",;.!:—") for line in poem]
# You can use the `+` operator to build up strings from each line as well:
["☛ " + line + " ☚" for line in poem]
# Using string slices, we can create some abstract poetry from parts of each line. Here we smoosh the first five characters of each line up against the last five characters:
[line[:5] + line[-5:] for line in poem]
# You may find discover a desire deep inside of you to use more than one of these transformations on the predicate expression. "Impossible," says a nearby moustachioed man, monocle popping from his orbital socket. But it can be done! In two ways. First, you can perform the transformation by assigning the result of one list comprehension to a variable, and then using that result in a second list comprehension. For example, to turn this poem into a telegram, we'll first convert it to upper case:
upper_frost = [line.upper() for line in poem]
# And then we'll get rid of punctuation at the end of the line:
upper_frost_no_punct = [line.strip(",;.!:—") for line in upper_frost]
# And then append the string STOP to the end of each line:
[line + " STOP" for line in upper_frost_no_punct]
# Not bad, but sort of inconvenient! You can actually write that whole thing using one expression. Any of those weird methods (`.lower()`, `.upper()`, etc.) mentioned above can be chained: you can attach them not just to line but to any other expression you made with line. Likewise, the `+` operator can be used with line but also any expression that results from performing a transformation on line. For example, you can rewrite the three list comprehensions above using one list comprehension with chained operators:
[line.upper().strip(",;.!:—") + " STOP" for line in poem]
# This is especially useful for multiple replacements. Here's the Swedish Chef version:
[line.replace("i", "ö").replace("o", "ö").replace("a", "ö").replace("e", "ur") for line in poem]
# ### Filtering lines
#
# Using the membership expression of a list comprehension, we can make a list of only the lines from the text file that match particular criteria. Any of the various expressions that answer questions about strings can be used in this spot. For example, to find all of the lines of a particular length:
[line for line in poem if len(line) == 33]
# Lines that have the string `travel`:
[line for line in poem if "travel" in line]
# Lines that start with `And`:
[line for line in poem if line.startswith("And")]
# ## Text files and lists of words
#
# Lines are an interesting unit to work with, especially with poetic source texts, as they give us an easy handle on large (but not too large) syntactic units. A more traditional unit of analysis is the word. Fortunately for us, getting words from a text file is (relatively) easy.
#
# Calling `open(filename).read()` will read the file `filename` into a Python string. We can then use the `.split()` method to split that string into a list of words. Without any parameters, the `.split()` method just breaks the string up into units delimited by any kind of whitespace (whether that's a space character, a tab, a newline, etc.). So, for example, to get all of the words from our Frost poem:
frost_txt = open("frost.txt").read() # evaluates to a string
words = frost_txt.split() # split evaluates to a list of strings
words
# Or, more succinctly:
words = open("frost.txt").read().split()
# Now we can ask simple questions about this poem, like how many words does it have?
len(words)
# We can create a new weird poem by randomly sampling words from the original:
random.sample(words, 20)
# Or sort the words in alphabetical order:
sorted(words)
# Using a list comprehension, we can get all of the words that meet particular criteria, like the words that have more than seven characters:
[item for item in words if len(item) > 7]
# Or all of the words that start with the letter `a`:
[item for item in words if item.startswith("a")]
# ## Formatting lists
#
# You've doubtlessly noticed with the previous examples that whenever we evaluate a list, Jupyter Notebook displays *the actual syntax you'd need to reproduce the list*, i.e., with the square brackets, quotes, commas and everything. That's the default way Python shows values when you evaluate it—which is usually helpful, because it means you can just take the text in the cell output and paste it into another notebook and you've got exactly the same value, without needing to re-type it. However, when we're creating poetic output, the extra Python punctuation is undesirable.
#
# To make the output prettier, we need to do the following steps:
#
# * Create a string from the list
# * *Print* the string (don't just evaluate it)
#
# To create a string from the list, we use the `.join()` method, as outlined above! For example, here's Smooshed Frost from earlier in the notebook, assigned to a variable:
smooshed = [line[:5] + line[-5:] for line in poem]
type(smooshed)
# This is a *list of strings*. To create a version of this where all of the strings are joined together, we'll use `.join()`. In this case, let's say that we want each string in the list to be a single line of text in the output, so we'll use `\n` (the newline character) as the "glue."
"\n".join(smooshed)
# Ugh, that's still not right though! It's one string now, but when we evaluate the expression Python is *showing* us the escape characters instead of *interpreting* them. To get Python to interpret them, we have to send the whole thing to the print function, like so:
print("\n".join(smooshed))
# There we go! Ready to submit to our favorite poetry journal. Of course, you don't have to join with a newline character. Let's say you want to make a prose poem (i.e., no line breaks) by randomly sampling fifty words in the Frost poem. We'll use the space character as the glue:
print(" ".join(random.sample(words, 50)))
# Nice!
# ## Making changes to lists
#
# Often we'll want to make changes to a list after we've created it---for
# example, we might want to append elements to the list, remove elements from
# the list, or change the order of elements in the list. Python has a number
# of methods for facilitating these operations.
#
# The first method we'll talk about is `.append()`, which adds an item on to
# the end of an existing list.
ingredients = ["flour", "milk", "eggs"]
ingredients.append("sugar")
ingredients
# Notice that invoking the `.append()` method doesn't itself evaluate to
# anything! (Technically, it evaluates to a special value of type `None`.)
# Unlike many of the methods and syntactic constructions we've looked at so far,
# the `.append()` method changes the underlying value---it doesn't return a
# new value that is a copy with changes applied.
#
# There are two methods to facilitate removing values from a list: `.pop()` and
# `.remove()`. The `.remove()` method removes from the list the first value that
# matches the value in the parentheses:
ingredients = ["flour", "milk", "eggs", "sugar"]
ingredients.remove("flour")
ingredients
# (Note that `.remove()`, like `.append()` doesn't evaluate to anything---it
# changes the list itself.)
#
# The `.pop()` method works slightly differently: give it an expression that
# evaluates to an integer, and it evaluates to the expression at the index
# named by the integer. But it also has a side effect: it *removes* that item
# from the list:
ingredients = ["flour", "milk", "eggs", "sugar"]
ingredients.pop(1)
ingredients
# > EXERCISE: What happens when you try to `.pop()` a value from a list at an index that doesn't exist in the list? What happens you try to `.remove()` an item from a list if that item isn't in that list to begin with?
#
# > ANOTHER EXERCISE: Write an expression that `.pop()`s the second-to-last item from a list. SPOILER: <span style="background: black;">(Did you guess that you could use negative indexing with `.pop()`?</span>
#
# The `.sort()` and `.reverse()` methods do exactly the same thing as their
# function counterparts `sorted()` and `reversed()`, with the only difference
# being that the methods don't evaluate to anything, instead opting to change
# the list in-place.
ingredients = ["flour", "milk", "eggs", "sugar"]
ingredients.sort()
ingredients
ingredients = ["flour", "milk", "eggs", "sugar"]
ingredients.reverse()
ingredients
# > EXERCISE: Write a Python command-line program that prints out the lines of a text file in random order.
# ## Iterating over lists with `for`
#
# The list comprehension syntax discussed earlier is very powerful: it allows you to succinctly transform one list into another list by thinking in terms of filtering and modification. But sometimes your primary goal isn't to make a new list, but simply to perform a set of operations on an existing list.
#
# Let's say that you want to print every string in a list. Here's a short text:
text = "it was the best of times, it was the worst of times"
# We can make a list of all the words in the text by splitting on whitespace:
words = text.split()
# Of course, we can see what's in the list simply by evaluating the variable:
words
# But let's say that we want to print out each word on a separate line, without any of Python's weird punctuation. In other words, I want the output to look like:
#
# it
# was
# the
# best
# of
# times,
# it
# was
# the
# worst
# of
# times
#
# But how can this be accomplished? We know that the `print()` function can display an individual string in this manner:
print("hello")
# So what we need, clearly, is a way to call the `print()` function with every item of the list. We could do this by writing a series of `print()` statements, one for every item in the list:
print(words[0])
print(words[1])
print(words[2])
print(words[3])
print(words[4])
print(words[5])
print(words[6])
print(words[7])
print(words[8])
print(words[9])
print(words[10])
print(words[11])
# Nice, but there are some problems with this approach:
#
# 1. It's kind of verbose---we're doing exactly the same thing multiple times, only with slightly different expressions. Surely there's an easier way to tell the computer to do this?
# 2. It doesn't scale. What if we wrote a program that we want to produce hundreds or thousands of lines. Would we really need to write a `print` statement for each of those expressions?
# 3. It requires us to know how many items are going to end up in the list to begin with.
# Things are looking grim! But there's hope. Performing the same operation on all items of a list is an extremely common task in computer programming. So common,
# that Python has some built-in syntax to make the task easy: the `for` loop.
#
# Here's how a `for` loop looks:
#
# for tempvar in sourcelist:
# statements
#
# The words `for` and `in` just have to be there---that's how Python knows it's
# a `for` loop. Here's what each of those parts mean.
#
# * *tempvar*: A name for a variable. Inside of the for loop, this variable will contain the current item of the list.
# * *sourcelist*: This can be any Python expression that evaluates to a list---a variable that contains a list, or a list slice, or even a list literal that you just type right in!
# * *statements*: One or more Python statements. Everything tabbed over underneath the `for` will be executed once for each item in the list. The statements tabbed over underneath the `for` line are called the *body* of the loop.
#
# Here's what the `for` loop for printing out every item in a list might look like:
for item in words:
print(item)
# The variable name `item` is arbitrary. You can pick whatever variable name you like, as long as you're consistent about using the same variable name in the body of the loop. If you wrote out this loop in a long-hand fashion, it might look like this:
item = words[0]
print(item)
item = words[1]
print(item)
item = words[2]
print(item)
item = words[3]
print(item)
# etc.
# Of course, the body of the loop can have more than one statement, and you can assign values to variables inside the loop:
for item in words:
yelling = item.upper()
print(yelling)
# You can also include other kinds of nested statements inside the `for` loop, like `if/else`:
for item in words:
if len(item) == 2:
print(item.upper())
elif len(item) == 3:
print(" " + item)
else:
print(item)
# This structure is called a "loop" because when Python reaches the end of the statements in the body, it "loops" back to the beginning of the body, and executes the same statements again (this time with the next item in the list).
#
# Python programmers tend to use `for` loops most often when the problem would otherwise be too tricky or complicated to solve using a list comprehension. It's easy to paraphrase any list comprehension in `for` loop syntax. For example, this list comprehension, which evaluates to a list of the squares of even integers from 1 to 25:
[x * x for x in range(1, 26) if x % 2 == 0]
# You can rewrite this list comprehesion as a `for` loop by starting out with an empty list, then appending an item to the list inside the loop. The source list remains the same:
result = []
for x in range(1, 26):
if x % 2 == 0:
result.append(x * x)
result
# ## Conclusion
#
# We've put down the foundation today for you to become fluent in Python's very powerful and super-convenient syntax for lists. We've also done a bit of data parsing and analysis! Pretty good for day one.
#
# Further resources:
#
# * [Lists](http://openbookproject.net/thinkcs/python/english3e/lists.html), from [How To Think Like A Computer Scientist: Learning with Python](http://openbookproject.net/thinkcs/python/english3e/index.html)
# * [Lists](https://docs.python.org/3/tutorial/introduction.html#lists) and [More on Lists](https://docs.python.org/3/tutorial/datastructures.html#more-on-lists) from the [Official Tutorial](https://docs.python.org/3/tutorial/index.html)
| understanding-lists-manipulating-lines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import time
import string
# **1**. (25 points)
#
# The following iterative sequence is defined for the set of positive integers:
#
# - n → n/2 (n is even)
# - n → 3n + 1 (n is odd)
#
# Using the rule above and starting with 13, we generate the following sequence:
#
# 13 → 40 → 20 → 10 → 5 → 16 → 8 → 4 → 2 → 1
#
# It can be seen that this sequence (starting at 13 and finishing at 1) contains 10 terms. Although it has not been proved yet (Collatz Problem), it is thought that all starting numbers finish at 1.
#
# Which starting number, under one million, produces the longest chain?
#
# NOTE: Once the chain starts the terms are allowed to go above one million.
# **Straightforward Solution**: Loop through 1 - 1,000,000, and keep track of the maximum length of chain and corresponding starting number.
# +
# %%time
def length(starting_num):
l = 1
while starting_num != 1:
starting_num = starting_num // 2 if starting_num % 2 == 0 else starting_num * 3 + 1
l += 1
return l
maximum_len, maximum_start = None, None
for num in range(1, 1000001):
cur_len = length(num)
if maximum_len is None or cur_len > maximum_len: # cannot change order: short circuit evaluation of `or`
maximum_len, maximum_start = cur_len, num
print(maximum_start)
# -
# **Recursion:** The recursion can be formulated as $$\texttt{chain-len}(i) = 1 + \left\{\begin{aligned}\texttt{chain-len}(i // 2)\quad i\text{ is even};\\\texttt{chain-len}(i * 3 + 1)\quad i\text{ is odd}.
# \end{aligned}\right.$$
# +
# %%time
# num_function_calls = 0
def length(starting_num):
# global num_function_calls
# num_function_calls += 1
if starting_num == 1:
return 1
elif starting_num % 2 == 0:
return 1 + length(starting_num // 2)
else:
return 1 + length(starting_num * 3 + 1)
maximum_len, maximum_start = None, None
for num in range(1, 1000001):
cur_len = length(num)
if maximum_len is None or cur_len > maximum_len: # cannot change order: short circuit evaluation of `or`
maximum_len, maximum_start = cur_len, num
print(maximum_start)
# print(f'Total function calls: {num_function_calls}')
# -
# Recursion is slower as a result of many function calls. However, recursion often helps in simplifying complex problems.
# **Faster Solution:** We can actually make use of the information we obtained in previous computation.
# Take the following example: Suppose we have calculated `chain-len(10) = 7`: 10 → 5 → 16 → 8 → 4 → 2 → 1. We can speed up the calculation in two ways:
#
# 1) We should not need to calculate `chain-len(16)`;
#
# 2) We should not need to calculate `chain-len(13)` to the last step (13 → 40 → 20 → 10 → 5 → 16 → 8 → 4 → 2 → 1).
# +
# %%time
# num_function_calls = 0
chain_len_dict = {1: 1}
def length(starting_num):
# global num_function_calls
num_function_calls += 1
# already calculated: just return the value
if starting_num in chain_len_dict.keys():
return chain_len_dict[starting_num]
new_num = starting_num // 2 if starting_num % 2 == 0 else starting_num * 3 + 1
# if the length of chain from new_num is l, then the length from starting_num is l+1.
result = 1 + length(new_num)
# cache the result
chain_len_dict[starting_num] = result
return result
maximum_len, maximum_start = None, None
for num in range(1, 1000001):
cur_len = length(num)
if maximum_len is None or cur_len > maximum_len: # cannot change order: short circuit evaluation of `or`
maximum_len, maximum_start = cur_len, num
print(maximum_start)
# print(f'Total function calls: {num_function_calls}')
# -
# **Optional: only for your interest**
#
# - List: `a = [..., ..., ..., ..., ..., ...]`. Use `a[i]` to denote the value of `chain-len(i)`.
#
# - Dict: `a = {...:..., ...:..., ...:..., ...:...}`. When you visit `i`, put `i:chain-len(i)` into the dictionary.
#
# In Python, searching whether an element is in a dictionary (based on hashmap) and fetching the key takes $O(1)$ to $O(n)$ time (depending on how many elements are in the set), while searching whether an element is in a list takes $O(n)$ time. Therefore, we prefer using a dictionary to store the visited elements.
#
# However, taking an element out from a list using index only takes $O(1)$ time. Therefore, for the list implementation, we do not simply put visited elements in a list; instead, we create a big list where each element corresponds to an index. This is called as "exchanging space for time".
# **Fact 1**: We will sooner or later have calculated all `chain-len(i)` for `1 <= i <= 1000000`;
#
# **Fact 2**: We may encounter some `chain-len(i)` values where `i` is larger than `1000000`, and we do not yet know how much the number can be.
#
# Due to fact 2, we cannot pre-assign a list and use `list[i]` to store the value of `chain-len(i)`. (We even don't know how large the list should be. Even if we know an upper bound, this may also be a waste of memory space.)
#
# Due to fact 1, if we use a dictionary to store `i` and corresponding `chain-len(i)`, the dictionary will at least have 1000000 elements, which might make searching slower.
#
# A compromise is to use a list for `1 <= i <= 1000000` and use a dictionary for other values encountered.
# +
# %%time
chain_len_list = [None] * 1000001
chain_len_list[1] = 1 # Why we need this?
chain_len_dict = {}
def length(starting_num):
# already calculated: just return the value
if starting_num <= 1000000 and chain_len_list[starting_num]:
return chain_len_list[starting_num]
if starting_num > 1000000 and starting_num in chain_len_dict.keys():
return chain_len_dict[starting_num]
new_num = starting_num // 2 if starting_num % 2 == 0 else starting_num * 3 + 1
# if the length of chain from new_num is l, then the length from starting_num is l+1.
result = 1 + length(new_num)
# cache the result
if starting_num <= 1000000:
chain_len_list[starting_num] = result
else:
chain_len_dict[starting_num] = result
return result
maximum_len, maximum_start = None, None
for num in range(1, 1000001):
cur_len = length(num)
if maximum_len is None or cur_len > maximum_len: # cannot change order: short circuit evaluation of `or`
maximum_len, maximum_start = cur_len, num
print(maximum_start)
# -
# **2** (25 points)
#
#
# - Perform the median polish to calculate just the *residuals* for this [example](https://mgimond.github.io/ES218/Week11a.html) in Python.
# - Use the matrix `xs` provided
# - Display the final result after 3 iterations to 1 decimal place and check if it agrees with
#
# 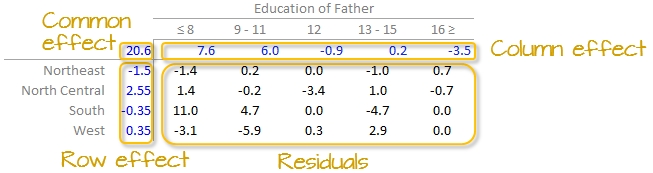
xs = np.array([
(25.3,32.1,38.8,25.4),
(25.3,29,31,21.1),
(18.2,18.8,19.3,20.3),
(18.3,24.3,15.7,24),
(16.3,19,16.8,17.5)
]).T
# +
def loop(array, nloop):
nrow, ncol = array.shape
# Step 1: Compute overall median and residual table
overall_median = np.median(array)
residual_table = array - overall_median # broadcast
row_effect = np.zeros(nrow)
col_effect = np.zeros(ncol)
for i in range(nloop):
# Step 2: Compute the row medians
row_medians = np.median(residual_table, axis = 1)
med_col_effect = np.median(col_effect)
# Step 3: Create a new residual table from the row medians
row_effect += row_medians
overall_median += med_col_effect
col_effect -= med_col_effect
residual_table -= row_medians[:, None]
# Step 4: Compute the column medians
col_medians = np.median(residual_table, axis = 0)
med_row_effect = np.median(row_effect)
# Step 5: Create a new residual table from the column medians
col_effect += col_medians
overall_median += med_row_effect
row_effect -= med_row_effect
residual_table -= col_medians
return np.round(residual_table, 1)
loop(xs, 3)
# +
def loop(array, nloop):
overall_median = np.median(array)
residual_table = array - overall_median
for i in range(3):
row_medians = np.median(residual_table, axis = 1)
residual_table -= row_medians[:, None]
col_medians = np.median(residual_table, axis = 0)
residual_table -= col_medians
return np.round(residual_table, 1)
loop(xs, 3)
# -
# **3**. (50 points)
#
# A Caesar cipher is a very simple method of encoding and decoding data. The cipher simply replaces characters with the character offset by $k$ places. For example, if the offset is 3, we replace `a` with `d`, `b` with `e` etc. The cipher wraps around so we replace `y` with `b`, `z` with `c` and so on. Punctuation, spaces and numbers are left unchanged.
#
# - Write a function `encode` that takes as arguments a string and an integer offset and returns the encoded cipher.
# - Write a function `decode` that takes as arguments a cipher and an integer offset and returns the decoded string.
# - Write a function `auto_decode` that takes as argument a cipher and uses a statistical method to guess the optimal offset to decode the cipher, assuming the original string is in English which has the following letter frequency:
#
# ```python
# freq = {
# 'a': 0.08167,
# 'b': 0.01492,
# 'c': 0.02782,
# 'd': 0.04253,
# 'e': 0.12702,
# 'f': 0.02228,
# 'g': 0.02015,
# 'h': 0.06094,
# 'i': 0.06966,
# 'j': 0.00153,
# 'k': 0.00772,
# 'l': 0.04025,
# 'm': 0.02406,
# 'n': 0.06749,
# 'o': 0.07507,
# 'p': 0.01929,
# 'q': 0.00095,
# 'r': 0.05987,
# 's': 0.06327,
# 't': 0.09056,
# 'u': 0.02758,
# 'v': 0.00978,
# 'w': 0.0236,
# 'x': 0.0015,
# 'y': 0.01974,
# 'z': 0.00074
# }
# ```
#
# - Encode the following nursery rhyme using a random offset from 10 to 20, then recover the original using `auto_decode`:
#
# ```text
# Baa, baa, black sheep,
# Have you any wool?
# Yes, sir, yes, sir,
# Three bags full;
# One for the master,
# And one for the dame,
# And one for the little boy
# Who lives down the lane.
# ```
def encode(s, offset):
offset = offset % 26 # let offset be within 0 and 25 (mod 26)
shift = lambda letters: letters[offset:] + letters[:offset]
# offset is defined outside shift, so it can be used in the lambda expression.
# offset is similar to global variable for shift
all_letters = string.ascii_lowercase + string.ascii_uppercase
shifted_letters = shift(string.ascii_lowercase) + shift(string.ascii_uppercase)
return s.translate(str.maketrans(all_letters, shifted_letters))
# $-30 = -2 \times 26 + 22$ (python, R) - round down the quotient
#
# $-30 = -1 \times 26 + (-4)$ (C, C++) - round the quotient towards zero
#
# In both cases, you can write `offset = (offset % 26 + 26) % 26` to get non-negative remainders.
def decode(s, offset):
return encode(s, -offset)
# How can we determine what offset the rhyme is encoded? We can do this based on the frequencies of each letter.
#
# An easy way is to identify the most common letter as `'e'`, since it has the highest frequency of appearance.
# If you suspect that only using one letter might be subject to randomness,
# - Theoretical frequency: $f = (f_1, f_2, \cdots, f_{26})$.
#
# - Empirical frequency under offset $i$: $f^{(i)} = (f_1^{(i)}, f_2^{(i)},\cdots, f_{26}^{(i)})$.
#
# Find $i$ such that $f^{(i)}$ is the closest to $f$.
# $f$ and $f^{(i)}$ are two points in $\mathbb{R}^{26}$, or the 25-simplex $\Delta^{25}$.
#
# Metrics:
#
# - inner product: $d_i = \sum f_jf_j^{(i)}$;
#
# - $L_2$-norm: $d_i^2 = \sum (f_j - f_j^{(i)})^2$;
#
# Kullback-Leibler divergence: $D_{KL}(f || f^{(i)})$.
freq = {
'a': 0.08167,
'b': 0.01492,
'c': 0.02782,
'd': 0.04253,
'e': 0.12702,
'f': 0.02228,
'g': 0.02015,
'h': 0.06094,
'i': 0.06966,
'j': 0.00153,
'k': 0.00772,
'l': 0.04025,
'm': 0.02406,
'n': 0.06749,
'o': 0.07507,
'p': 0.01929,
'q': 0.00095,
'r': 0.05987,
's': 0.06327,
't': 0.09056,
'u': 0.02758,
'v': 0.00978,
'w': 0.0236,
'x': 0.0015,
'y': 0.01974,
'z': 0.00074
}
rhyme = '''
Baa, baa, black sheep,
Have you any wool?
Yes, sir, yes, sir,
Three bags full;
One for the master,
And one for the dame,
And one for the little boy
Who lives down the lane.
'''
encoded_rhyme = encode(rhyme, np.random.randint(10, 21))
print(encoded_rhyme)
from collections import Counter
import re
l = Counter(re.sub(r'[^a-zA-Z]', '', encoded_rhyme.lower())).most_common(1)[0][0] # find the most common letter
offset = ord(l) - ord('e')
decoded_rhyme = decode(encoded_rhyme, offset)
print(decoded_rhyme)
| homework/solutions/Homework01 - Sample Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 第15讲 编写方法绘制线段
# ### Problem 问题描述
# 数轴上有6个点,对应的数字分别为`[-4.5, -2.0, 0, 1.5, 3.5, 5.0]`。连接其中任意的两个点可以形成一条线段。请问这6个点一共可形成多少条线段?参照下图,在宽高分别为600和400的绘图区绘制一条数轴,在数轴上标记这些点,并用不同颜色绘制出这6个点能形成的所有的线段并给每条线段标记一个序号。为避免绘制出的线段相互重合,每条线段间隔10个像素。图中已经绘制出了从`-4.5`开始到所有其它点的线段。
#
# 下面的代码已经完成了数轴绘制和对每一个点的标记。
#
# <img src="figures/L015_draw_all_lines.png" width="600px"/>
# +
from turtle import setup, reset, pu, pd, bye, left, right, fd, bk, screensize
from turtle import goto, seth, write, ht, st, home, pen, dot
nums = [-4.5, -2.0, 0, 1.5, 3.5, 5.0]
colors = ["orange", "black", "red", "green", "blue"]
SCALE = 50
width, height = 600, 400 # 窗口的宽度和高度(单位为:像素)
setup(width, height, 0, 0)
# -
from qianglib import draw_axis, mark
reset()
draw_axis(width, height, SCALE)
nums = [-4.5, -2.0, 0, 1.5, 3.5, 5.0]
colors = ["orange", "black", "red", "green", "blue"]
i = 0
n_colors = len(colors)
while i < len(nums):
mark(nums[i], size=5, color=colors[i % n_colors])
i += 1
# ### Math Background 数学背景
#
# 1. 简单的排列组合
# + [markdown] heading_collapsed=true
# ### Prerequisites 预备知识
#
# #### 1. 排列和组合
#
# - 4个小朋友Jason, Sophie, Tony, Yunzi按从前到后的次序排成一个纵队,可以排成多少个不一样的队伍?
# -
classmates = ["Jason", "Sophie", "Tony", "Yunzi"]
# #### 2. 充分理解`goto(x, y)`的意义
st() # show turtle
pu()
goto(100, 0) # turtle moved to 2.0 on x Axis.
dot(5, "red")
goto(100, 50) # turtle moved to 50 pixels above 2.0 on x Axis
dot(5, "blue")
goto(100, -50) # turtle moved to 50 pixles below 2.0 on x Axis
dot(5, "black")
ht()
# #### 3. 声明绘制一条线段的方法
# - 考虑绘制一条线段需要知道哪些数据:线段的两个端点的位置,线的粗细以及颜色
# - 给方法以及这些数据进行命名,方法名可定为`line`,两个端点的位置可以分别为`start`,`end`,线的粗细设为`linewidth`,颜色为`color`
# - 为了避免绘出的线发生重叠,考虑将线段整体往上平移一定的距离,这个距离可以用变量`offset`来表示,
# - 不要忘记了还有一个比例尺,用变量`scale`表示。
#
# 这样我们可以如下声明绘制线条的方法:
# ```python
# def line(start, end, linewidth, color, offset, scale):
# #TODO: need implementation 需要实现该方法
# return
# ```
# 其中`def`是一个关键词,它表明将要声明一个方法`line`是方法名,上述所有在方法名后的括号`()`内声明的变量名又叫方法接受的**参数**。改行以冒号`:`结束,随后每一行代码相对于`def`都有缩进,直到最后执行`return`语句,完成这个方法或者缩进结束。冒号以后开始的直至`return`语句(或者缩进结束)构成了这个方法的**方法体**。
# #### 4. 实现先前声明的方法
#
# 实现一个方法指的是在方法体内部编写代码以完成方法设定的功能的过程。
#
# 在`line`方法体内,我们需要依次完成下面5个步骤:
# 1. 提起画笔
# 2. 移动画笔至要绘制线段的一个端点(`start`),端点的位置与参数`start`、`offset`和`scale`的值有关。
# 3. 设定好画笔的颜色和粗细
# 4. 落下画笔准备绘制
# 5. 移动画笔至要绘制线段的另一个端点(`end`),同样位置需要计算得到。
#
# 在上面的几个步骤中,设置画笔颜色和粗细这一步的次序可以相对随意些。
#
# 这样我们可以如下声明绘制线条的方法:
# ```python
# def line(start, end, linewidth, color, offset, scale):
# pu() # 提起画笔
# pen(pensize=linewidth, pencolor=color) # 设定画笔粗细和颜色
# goto(start * scale, offset) # 移动画笔至某端点
# pd() # 落下画笔准备绘图
# goto(end * scale, offset) # 移动画笔值另端点完成线条绘制
# return # 离开方法,返回。
# ```
# 有了这样的实现,我们就可以在代码中调用这个方法来完成一条线段的绘制,下面的代码使用黑色的画笔,3个像素的粗细来绘制一条从(`-4.5*SCALE`, `15`)表示的点到(`-2.0*SCALE`, `15`)表示的点之间的线段。这条线段位于x轴上方5个像素。如下图所示:
# ```python
# line(-4.5, -2.0, 3, "black", 15, SCALE)
# ```
#
# <img src="figures/L015_one_line.png" width="600px"/>
# 在调用方法时,给方法提供的参数的次序需要与声明方法时的次序一致,否则会出错或预期以外的功能。
#
# 如果需要给绘制的线段进行标号,那么需要额外完成一些诸如:提起、移动画笔和绘制文字的步骤。同时我们还应该有标号使用的数字(或文字),需要新设定一个参数来表示它,可以用`label`来表示,下面的代码请读者自己添加这几行代码给这个方法添加线段标号的功能。
# ```python
# def line(start, end, linewidth, color, offset, scale, label):
# # ..... # 省略了之前的一些代码
# pu() # 提起画笔
# goto(start * scale - 10, offset) # 移动画笔至要标号的位置
# write(str(label), align="right") # 书写标号使用默认字体
# return # 离开方法,返回
# ```
# #### 5. 练习:
# 1. 补全line方法使得其可以对所绘制的线段进行标号
# 2. 调用你刚刚完成的具有标号功能的`line`方法,绘制一条线段。线段的起点对应数轴上-2.0表示的位置,终点对应数轴上5.0表示的位置,线宽选择5个像素,颜色从提供的颜色中任意选择一种,绘制的线段距离x轴的约50像素远,比例尺选择默认的比例尺,标号用你自己的名字字符串(例如"Celine")。
# ### 3. Solution 编程求解
#
# <img src="figures/L015_solution_figure.png" width="600px"/>
def line(start, end, linewidth, color, offset, scale, label):
pu() # 提起画笔
pen(pensize=linewidth, pencolor=color) # 设定画笔粗细和颜色
goto(start * scale, offset) # 移动画笔至某端点
pd() # 落下画笔准备绘图
goto(end * scale, offset) # 移动画笔值另端点完成线条绘制
pu() # 提起画笔
goto(start * scale - 10, offset) # 移动画笔至要标号的位置
write(str(label), align="right") # 书写标号使用默认字体
return # 离开方法,返回
# + tags=[]
nums = [-4.5, -2.0, 0, 1.5, 3.5, 5.0]
colors = ["orange", "black", "red", "green", "blue"]
i, j = 0, 1 # index i, j: start, end in nums
height = 10
n_drawn = 0
while i < len(nums) - 1:
j = i + 1
while j < len(nums):
line(nums[i], nums[j], 3, colors[n_drawn % n_colors], # 可以换行
height * (n_drawn + 1), SCALE, label=n_drawn+1)
n_drawn += 1
j += 1
i += 1
# -
# ### Summary 知识点小结
# 1. 如何用循环嵌套循环来列举各种可能的排列组合
# 2. 充分理解`goto`方法及其接受的参数的意义
# 3. 学习编写接受参数的方法,理解参数的次序的重要性
# 4. 学习调用自己编写的方法
# 5. 复习`dot`方法,复习比例尺概念
# ### 计算机小知识
# 暂缺
# + [markdown] heading_collapsed=true
# ### Assignments 作业
# **注意**:本讲所有的作业涉及到绘图时均使用本讲示例所使用的数轴和比例尺。
#
# **Attention**: All the following assignments concerning drawing are supposed to use the axis and scale that are used in this lecture.
# + [markdown] hidden=true
# 1. 创建(声明和实现)两个不同的方法计算两个不同的正整数之间所有正整数的和,其中一个方法得到的和包括给出的这两个正整数参数;另一个方法得到的和不包括给出的这两个正整数参数。在两个方法体内部输出得到的和。观察比较结果有什么不一样。并用下面的一系列数据来测试你编写的方法。<br>
# Create(declare and implement) two defferent methods to calculate the sum between two different positive integers. In one of the methods, the sum includes the two integers while in another method, the sumd doesn't. Test your methods by using the data provided in below table and print out the result following the format in the table as well.<br>
#
# | num1 | num2 | sum(include num1,2) | sum(exclude num1,2) |
# | ---------:|-----------:|:---------------------|:--------------------|
# | 12 | 12 | sum(inclu) = 24 | sum(exclu) = 0 |
# | 1 | 10 | sum(inclu) = 55 | sum(exclu) = 44 |
# | 10 | 1 | sum(inclu) = 55 | sum(exclu) = 44 |
# | 100 | 1000 | sum(inclu) = 495550 | sum(exclu) = 444450 |
#
# +
# use this cell to create your methods
1 10
sum1 = 1 + 2 +3 + 。。。 +10
sum2 = 2 + 3 + 。。+ 9
# +
# run your method(4 times for each method) with the test data in this cell.
# -
# 2. 仅编写一个方法来完成前一题。<br>
# Complete the previous question by only creating(declaring and implementing) one method.
#
# **提示**:在创建这个新方法时,新增加一个参数,这个参数可以取两个不同的值,在方法体内不根据这个参数的值使用`if`条件语句来动态的决定计算的和是否包括前两个正整数参数。<br>
# **Hint**: Add another parameter to indicate whether the sum include the two positive integers or not when declaring this new method. and In the method body, adjust your codes to handl this two scenarios by using `if` statement.
# +
# use this cell to create your method
# +
# run your method 8 times with the test data in this cell.
# -
# 3. 声明并且实现一个方法,该方法接受数轴上的一个点表示的数字作为参数1,另一个数作为参数2,完成以参数1为表示的位置为中心、参数2表示的数据为边长其中两条边平行于x轴的正方形的绘制,同时要求你实现的这个方法在绘制正方形时可以在调用方法时确定正方形边的颜色和线宽。<br>
# Declare and implement a method which accepts a number denoting a point in x axis as paramter1 and another number as paramter2 to draw a square with the parameter1 as the square's center and parameter2 as its side length. Two sides of the drawn square should be parallel to X axis. Besides, your method should also be able to draw squares with different colors and line widths.<br>
# 实现这个方法后,用下面的数据来绘制3个正方形。结果应如图所示:<br>
# Use the following test data to draw three squares after you complete implementing the method. The result should be like the following figure:<br>
#
# | center | side length | color | line width |
# | ---------:|-----------:|:---------:|:-----------:|
# | -2.0 | 2.0 | red | 3 |
# | 1.5 | 3.0 | green | 4 |
# | 0.0 | 1.0 | blue | 3 |
#
#
# **要求**:使用`goto`方法来移动画笔,禁止使用`left`,`right`,`fd`,`bk`等方法。 <br>
#
# **Requirement**: Only use `goto` to move pen, you are not allowed to use methods like `left`,`right`,`fd`,`bk`;<br>
#
# **提示**:先计算出正方形每一个顶点的位置。<br>
#
# **Hint**: Calculate the position of each vertex of the square bfore you draw it.
# <br>
# <img src="figures/L015_exercise_3.png" width="600px"/>
# + [markdown] hidden=true
# 4. 声明并且实现一个方法,该方法接受数轴上的两个不同的点表示的数字作为参数,完成以这两个点为端点的位于数轴上方的一个正三角形的绘制,同时要求你实现的这个方法在绘制正三角形时可以在调用方法时确定三角形边的颜色和线宽。<br>
# Declare and implement a method which accepts two numbers denoting two points in x axis as paramters to draw a regular triangle, one side of which is located on X axis and the other vertex is above the axis.Besides, your method should also be able to draw the triangles with different colors and line widths.<br>
# 实现这个方法后,用下面的数据来绘制3个三角形。结果应如图所示:<br>
# Use the following test data to draw three regular triangles after you complete implementing the method. The result should be like the following figure:<br>
#
# | num1 | num2 | color | line width |
# | ---------:|-----------:|:---------:|:-----------:|
# | -4.5 | 0.0 | red | 3 |
# | -2.0 | 1.5 | green | 4 |
# | 1.5 | 3.5 | blue | 5 |
#
#
# **要求**:尽可能不使用`left`,`right`,`fd`,`bk`等方法,而使用`goto`方法来移动画笔。<br>
#
# **Requirement**: Try to only use `goto` to move pen instead of using `left`,`right`,`fd`,`bk`.<br>
#
# **提示**:使用goto方法时,需要用到一个数值`1.732`来帮助定位三角形的端点。请在你实现方法的代码里使用这个数值。<br>
#
# **Hint**: You may need `1.732` to help locate the position of a vertex. Please use this value when implementing your method.
#
# <img src="figures/L015_exercise_4.png" width="600px"/>
# -
# 5. [**难,选做**]用数字"1,2,3,4"这4个数字来组成一个5位数,其中每一个数至少用到1次,一共可以组成多少个5位数,其中最小的和最大的数分别是什么数?<br>
#
# [**hard, optional**]Comose a 5-digit positive integer using the number 1,2,3, and 4 on each bit. Each number should at least be used once. How many 5-digit positive number can be composed? what is the minimal and maximal integer?<br>
#
# **提示1 Hint1**: `123 = 1*100 + 2*10 + 3*1` <br>
#
# **提示2**: 可以使用一个`list`类型的变量,把所有找到的5位数都存放在这个变量中,这个列表型变量的长度就是所有5位数的个数,列表中最大和最小的元素就分别是能形成的最大的和最小的5位数<br>
#
# **Hint2**: You can use a `list` variable, put all the integers to it, then the length of this variable is the number of all integers. and minimal and maximal elements are the minimal and maximal value of all composed 5-digit integers.
| source/2021/100Beginner/content/015_how_many_lines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1D Linear Regression Using Gradient Descent
#
# Here we will introduce 1D Linear Regression and compute the weights using Gradient Descent.
#
# This notebook will give you an algorithm that will work for n number of input features but first we'll test it on 1D data.
#
# ## Process
#
# * Loading the data
# * Exploring the data
# * Adding bias to our input
# * Computing the cost function $J(w)$
# * Implementing Gradient Descent
# * Computing the hypothesis
# * Plotting our line of best fit
# * Making a Prediction
# * Visualizing $J(w)$
# * Determine how well the model performed
#
# ## Equations
# The objective of linear regression is to minimize the cost function
#
# $$ J(w) = \frac{1}{2m} \sum_{i=1}^m \left( h_{w}(x^{(i)}) - y^{(i)}\right)^2$$
#
# where $m$ is the number of training examples,
#
# $y^{(i)}$ is the label or output value from the dataset for a specific row which is determined by the index $i$,
#
# $x^{(i)}$ is a specific row of the feature matrix which is determined once again by the index $i$,
#
# $h_w(x)$ is the hypothesis which is given by the linear model
#
# $$ h_w(x) = w^T x = w_0x_0 + w_1 x_1$$
#
# where $w$ is a column vector of weights, i.e.,
# $$ w = \begin{bmatrix} w_0 \\ w_1 \\ w_2 \\ \vdots \\ w_n \end{bmatrix} $$
#
# These weights are estimated using the gradient descent algorithm specifically the batch gradient descent algorithm which is a procedure of simultaneously updating the values of $\theta_j$, i.e.,
#
# $$ \theta_j := \theta_j - \alpha \frac{1}{m} \sum_{i=1}^m \left( h_\theta(x^{(i)}) - y^{(i)}\right)x_j^{(i)} $$
#
# where $j = 0, 1, ..., n$ represents the feature index number with $n$ representing the number of features,
#
# $\alpha$ represents the learning rate,
#
# $x_j^{(i)}$ represents a value in the feature matrix for a specific row which is determined by the index $i$ and the value in the specified row is determined by the feature index $j$, and
#
# $:=$ means to assign the value computed on the right-hand side to the variable on the left-hand side.
#
# Note (1): As long as the learning rate $\alpha$ is not too large with each step of gradient descent, the weights $\theta_j$ will become closer to the optimal values that will achieve the lowest cost $J(\theta)$.
#
# If $\alpha$ is too large this can cause gradient descent to diverge which means a local minimum or global minimum will not be found, and the smaller $\alpha$ is the longer it will take gradient descent to find the minimum.
#
# So, you can think of alpha as the size of the step that is being taken with each iteration.
#
# Note (2): Here we will be performing 1D Linear Regression which means $n = 1$, and our training data has 97 rows which means $m = 97$
# +
import os
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
# %matplotlib inline
# -
# ## Loading the data
data = np.loadtxt('data/population_vs_profit.txt', delimiter=',')
m = len(data)
n = len(data[0]) - 1
print("Shape of data", data.shape)
print("Number of samples", m)
print("Number of features", n)
X, y = data[:, n-1], data[:, n]
# ## Exploring the data
#
# ### Plotting the data
#
# #### Is our data linear?
#
# We can show a simple 2D plot to get a general idea if our feature has a linear relationship with our output.
plt.scatter(X, y)
plt.ylabel('Profit in $10,000')
plt.xlabel('Population in 10,000s');
# Our data seems to be pretty linear so Linear Regression seems like a good option.
# ### Visualizing the shape of our data
#
# $X$ is a feature matrix that is $m$ $x$ $(n + 1)$ where:
#
# * $m =$ number of training examples
# * $n =$ number of features
#
# $\vec{y}$ is our ouput vector that is $m$ $x$ $1$
print("The shape of X is", X.shape)
print("The shape of y is", y.shape)
# ## Adding bias to our input
#
# This is the process of adding the feature $x_0$ to
#
# $$ h_\theta(x) = \theta_0 + \theta_1 x_1 $$
#
# To do this we add a column of 1's to our feature matrix and call it $x_0$
#
# Note: If we have more than one feature and our features differ by orders of magnitude, then we would normalize the features before adding the bias to our input. Feature normalization is utilized because it can cause gradient descent to converge at a faster rate. However, here we have only one feature, so we will not be utilizing feature normalization in this example.
# +
# here we add a column of 1's for our bias we'll use stack for numpy arrays with shape (m,)
# the numpy function stack joins arrays along a given axis
# the first axis (axis=0) refers to rows (training examples) and the second axis (axis=1) refers to columns (features).
# Note: Running this cell more than once will result in an error since all the input arrays will no longer be the same shape
X = np.stack([np.ones(m), X], axis=1)
print("Shape of X is", X.shape)
print("Example feature vector", X[0])
# -
# ## Computing the cost function $J(\theta)$
# function to compute cost
def computeCost(X, y, theta):
# get number of training examples
m = y.shape[0]
# compute hypothesis
h = X@theta
# compute cost function
J = 1/(2*m)*sum((h-y)**2)
return J
# ## Implementing Gradient Descent
# function for implementing gradient descent
def gradientDescent(X, y, theta, alpha, num_iters):
# get number of training examples
m = y.shape[0]
# Use a list to store the values of the cost function for every iteration
J_history = []
for i in range(num_iters):
# Note: h can also be calculated as h = theta@X.T, h = theta.T@X.T, or h = X@theta.T because the shape of theta is (2,) in numpy which means it can act as a row vector or a column vector and it will return h with the same shape of (97,)
h = X@theta
theta = theta - alpha * (1/m) * (h-y)@X
# save the cost on every iteration
J_history.append(computeCost(X, y, theta))
return theta, J_history
# +
# initialize the weigths to 0
theta = np.zeros(2)
# some gradient descent settings
num_iters = 1500
alpha = 0.01
theta, J_history = gradientDescent(X, y, theta, alpha, num_iters)
print('Theta found by gradient descent: {:.4f}, {:.4f}'.format(*theta))
print('Expected theta values (approximately): [-3.6303, 1.1664]')
# -
# ## Computing the hypothesis
#
# We'll take our feature matrix $X$ and pretend we dont know our output vector $\vec{y}$.
#
# Now using the weights we found we'll try and predict what $\vec{y}$ should be at any point.
# +
# Note: h can also be calculated as h = theta@X.T, h = theta.T@X.T, or h = X@theta.T because the shape of theta is (2,) in numpy which means it can act as a row vector or a column vector and it will return h with the same shape of (97,)
h = X@theta
# allows us to make sure h has the expected shape
print("Shape of h is", h.shape)
# -
# ## Plotting our line of best fit
# X[:, 1] returns all of the rows of X in its second column remember the first column of X consists of 1's
plt.scatter(X[:, 1], y)
plt.scatter(X[:, 1], h, c='green')
plt.ylabel('Profit in $10,000s')
plt.xlabel('Population in 10,000s')
plt.plot(X[:, 1], h, c='red');
# # Making a prediction
#
# Now we can create an input vector and get an estimated result.
# +
# Note: 4.5 corresponds to a of population of 15,000 since the each value in the dataset is divided by 10,000
input_vec = [4.5]
# add a bias since all inputs must begin with 1
input_vec_bias = np.append(1, input_vec)
# make a prediction
profit = input_vec_bias@theta
print("Input vector", input_vec)
print("Input vector with bias", input_vec_bias)
print("theta", theta)
print("Profit", profit*10000)
# -
# ## Visualizing $J(\theta)$
# +
# grid over which we will calculate J
theta0_vals = np.linspace(-10, 10, 100)
theta1_vals = np.linspace(-1, 4, 100)
# initialize J_vals to a matrix of 0's
J_vals = np.zeros((theta0_vals.shape[0], theta1_vals.shape[0]))
# Fill out J_vals
for i, theta0 in enumerate(theta0_vals):
for j, theta1 in enumerate(theta1_vals):
J_vals[i, j] = computeCost(X, y, [theta0, theta1])
# Because of the way meshgrids work in the surf command, we need to
# transpose J_vals before calling surf, or else the axes will be flipped
J_vals = J_vals.T
# surface plot
fig = plt.figure(figsize=(12, 5))
ax = fig.add_subplot(121, projection='3d')
ax.plot_surface(theta0_vals, theta1_vals, J_vals, cmap='viridis')
ax.xaxis.set_tick_params(labelsize=8)
plt.xlabel('theta0')
plt.ylabel('theta1')
plt.title('Surface')
# contour plot
# Plot J_vals as 15 contours spaced logarithmically between 0.01 and 100
ax = plt.subplot(122)
plt.contour(theta0_vals, theta1_vals, J_vals, linewidths=2, cmap='viridis', levels=np.logspace(-2, 3, 20))
plt.xlabel('theta0')
plt.ylabel('theta1')
plt.plot(theta[0], theta[1], 'ro', ms=10, lw=2)
plt.title('Contour, showing minimum')
pass
# -
# ## Determine how well the model performed
#
# We need some numerical measure to see how well our model performed
#
# For this we can use $R^2$ (R-Squared)
#
# We ususally use this for any regression not just Linear Regression
#
# The definition for R-Squared is the following:
#
# $$R^2 = 1 - \frac{SS_{res}}{SS_{tot}}$$
#
# where:
#
# $SS_{res}$ is the sum of squared residual
#
# and
#
# $SS_{tot}$ is the sum of squared total
#
# These are defined as:
#
# $$SS_{res} = \sum^m_{i=1}(y^{(i)} - h_\theta(X))^2$$
#
# <br />
#
# $$SS_{tot} = \sum^m_{i=1}(y^{(i)} - \bar{y}^{(i)})^2$$
#
# $R^2 = 1$ is a perfect model
#
# $R^2 = 0$ is basically the average (50%)
#
# $R^2 = -$ is worse than just computing the average
# +
SSres = sum((y-h)**2)
SStot = sum((y-y.mean())**2)
R2 = 1 - SSres/SStot
print('SSres is: ', SSres)
print('SStot is:', SStot)
print('R-squared is: ', R2)
# -
| LinearRegression/03-1D-LR-GD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Science User Case - Inspecting a Candidate List
# Ogle et al. (2016) mined the NASA/IPAC Extragalactic Database (NED) to identify a new type of galaxy: Superluminous Spiral Galaxies. Here's the paper:
#
# Here's the paper: https://ui.adsabs.harvard.edu//#abs/2016ApJ...817..109O/abstract
#
# Table 1 lists the positions of these Super Spirals. Based on those positions, let's create multiwavelength cutouts for each super spiral to see what is unique about this new class of objects.
# Import some python modules so we don't get inundated with unimportant warnings when we run our notebook cells.
# +
import warnings
warnings.filterwarnings("ignore")
import sys
sys.path.append('workshop-dev-master')
from navo_utils.image import Image, ImageColumn
from navo_utils.spectra import Spectra, SpectraColumn
from navo_utils.cone import Cone
from navo_utils.tap import Tap
from navo_utils.utils import astropy_table_from_votable_response
from astropy.table import Table
# Utility functions used in the workshop.
import navo_utils.utils
# -
# ## Search NED for objects in this paper.
from astroquery.ned import Ned
objects_in_paper = Ned.query_refcode('2016ApJ...817..109O')
objects_in_paper.show_in_notebook()
# ## The NED query returns all objects in the paper, not just the galaxies in Table 1. Let's filter these results to only include the galaxies.
# Hint: There is a difference between strings and byte strings
#Let's see whether we are dealing with strings or byte strings
objects_in_paper['Type'].data[0]
# +
#Byte strings it is!
galaxies = objects_in_paper[objects_in_paper['Type'] == b'G']
galaxies.show_in_notebook()
# -
# # Search the NAVO Registry for WISE, GALEX, and SDSS image services
# +
from navo_utils.registry import Registry
wise_services = Registry.query(keyword='allwise', service_type='image')
print(f'{len(wise_services)} result(s) found.')
wise_services
# +
galex_services = Registry.query(keyword='galex', service_type='image')
print(f'{len(galex_services)} result(s) found.')
galex_services
# +
sdss_services = Registry.query(keyword='sdss', service_type='image')
print(f'{len(sdss_services)} result(s) found.')
sdss_services
# -
# # For each position, show the corresponding WISE, GALEX, and SDSS images.
# +
from astropy.coordinates import SkyCoord # High-level coordinates
from navo_utils.image import Image
import astropy.units as u #is this needed?
missions = ['GALEX', '2MASS', 'AllWISE']
base_urls = ['http://mast.stsci.edu/portal_vo/Mashup/VoQuery.asmx/SiaV1?MISSION=GALEX&',
'http://irsa.ipac.caltech.edu/ibe/sia/twomass/allsky/allsky?',
'https://irsa.ipac.caltech.edu/ibe/sia/wise/allwise/p3am_cdd?']
#Create an astropy table with this info.
search_list = Table([missions, base_urls], names = ('mission', 'access_url'))
size = '0' #diameter in degrees
for object in galaxies:
pos = SkyCoord(object['RA(deg)'], object['DEC(deg)'], frame="icrs", unit=(u.deg, u.deg))
for mission in missions:
wise_results = Image.query(coords=pos, radius=size, service=wise_services[0])
table = wise_results[0]
table.show_in_notebook()
galex_results = Image.query(coords=pos, radius=size, service=galex_services[0])
sdss_results = Image.query(coords=pos, radius=size, service=sdss_services[0])
#xid = SDSS.query_region(pos, spectro=True) #requires matches to have spectroscopy, not just photometry:
#im = SDSS.get_images(matches=xid, band='g')
# -
# ## Grab the SDSS images for each super spiral
# +
from astroquery.sdss import SDSS
from astropy.coordinates import SkyCoord # High-level coordinates
import astropy.units as u
for object in galaxies:
pos = SkyCoord(object['RA(deg)'], object['DEC(deg)'], frame="icrs", unit=(u.deg, u.deg))
xid = SDSS.query_region(pos, spectro=True) #requires matches to have spectroscopy, not just photometry:
im = SDSS.get_images(matches=xid, band='g')
| Science Use Case - Inspecting a Candidate List.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sos
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SoS
# language: sos
# name: sos
# ---
# + [markdown] kernel="SoS"
# ## On the Generation of HTML Report from Jupyter Notebook
#
# This example demonstrates how to use the `%preview` magic to preview results and how to generate a report in HTML format from a Jupyter notebook with SoS kernel. The report has the following features:
#
# 1. The report contains all details of the analysis but only displays output or both input and output of selected cells by default.
# 2. A control panel (hidden at the top left corner of the screen) can be used to display hidden contents.
# 3. The report can contain interactive tables and plots generated from the `%preview` magic of SoS. The tables are sortable and filterable, and is displayed in a scrollable window so you can include complete tables with thousand of records in a compact format. You can hover over data points of scatter plots to get details of each data point.
# -
# ### Description of data
#
# We have got an excel file with a list of differentially expressed genes for certain experiment. The analysis was done with ensembl genes so it is difficult to figure out what they are. The goal of the analysis is to annotate the gene IDs with HGNC names.
#
# Let us first read the data from the excel file and have a look:
# + kernel="SoS" tags=["report_output"]
# %preview -n data -l 2000
depends: Py_Module('xlrd')
import pandas as pd
data = pd.read_excel('DEG_list.xlsx')
# -
# ### Data analysis
#
# The [biomaRt](https://bioconductor.org/packages/release/bioc/html/biomaRt.html) bioconductor package provides required datasets to annotate the ensembl IDs so we transfer the gene list from Python to R, and use the biomaRt package to annotate the IDs.
#
# As we can see from the following list, the genes are correctly annotated
# + kernel="R"
# %get data
# + kernel="R" tags=["report_output"]
# %preview -n hgnc[1:5,]
library(biomaRt)
ensembl <- useEnsembl(biomart='ensembl')
#listDatasets(ensembl)
ensembl <- useEnsembl(biomart="ensembl", dataset="mmusculus_gene_ensembl")
hgnc <- getBM(attributes=c('ensembl_gene_id', 'external_gene_name'),
filters = 'ensembl_gene_id', values = data['ensembl_gene_id'], mart = ensembl)
annotated <- merge(data, hgnc, by='ensembl_gene_id', all.x=TRUE)
# -
# ### Result
#
# We transfer the annotated result back to Python and write it to another excel file. The content of the file is previewed and list here:
#
# + kernel="SoS" tags=["report_output"]
# %get annotated --from R
# %preview -n annotated_DEG_list.xlsx -l 2000
annotated = annotated.set_index('external_gene_name')
annotated.sort_values(by='padj', inplace=True)
annotated.to_excel('annotated_DEG_list.xlsx')
# + [markdown] kernel="SoS"
# Just to demonstrate the scatterplot style of `%preview` magic and the `report_cell` tag, here is the command to display a scatter plot of `log2FoldChange` vs `baseMean`. With the tooltip, you can easily figure out which gene has the highest `baseMean`.
# + kernel="SoS" tags=["report_cell"]
# %preview annotated -n -s scatterplot log2FoldChange baseMean \
--tooltip ensembl_gene_id pvalue padj --log y
# -
# ### Report Generation
#
# Now that we have completed our analysis, you can mark the cells with tags to control how they appear in the report. If you are not familiar with Jupyter, a `tag` is any string that can be attached to a cell. You can view, add, and remove tags from the tag toolbar, which can be turned on using `View` -> `Cell Toolbar` -> `Tags`.
#
# You can
# 1. Mark code cells that will be outputed with shortcut `Ctrl-Shift-O`. This will add a `report_output` tag to the cell and mark the cell with a gray bar to the right of the output area.
# 2. Using the same shortcut, mark markdown cell that will be hidden with tag `hide_output`.
# 3. Exclude cells from the report by tagging them with tag `scratch`. There is no shortcut so you have to do this manually.
#
# Now you have a notebook with the following types of cells, they will appear in the generated report as follows:
#
# |Cell type | Tag | Default status in report |
# |---|---|---|---|
# |Markdown cell| None | Displayed|
# |code cell| None | Hidden |
# |Markdown cell|`hide_output` | Hidden|
# |Code cell|`report_cell`| Displayed with hidden input and messages| |Code cell|`report_output`| Displayed with hidden messages |
# |Code cell|`scratch`| Excluded, not in report |
# |Markdown cell|`scratch`| Excluded, not in report |
#
#
# To generate a report, you can execute cell magic
# ```
# # %sossave --to html --force
# ```
# to convert `file.ipynb` to `file.html` under the same directory. The `--force` option allows SoS to overwrite an existing report. This command is available from the dropdown list of the panel cell input box so you do not have to enter it manually.
#
# If you would like to save the report in another name or to another directory, you can use magic
# ```
# # %sossave myreport.html --force
# ```
# or use convert the notebook from the command line using command
# ```
# $ sos convert myanalysis.ipynb myanalysis.html --template sos-report
# ```
# + [markdown] tags=["scratch"]
# Just for demonstration purposes, this paragraph will not be displayed in the report because of the inserted `scratch` tag.
# + [markdown] tags=["hide_output"]
# And this paragraph will be hidden from the report because of the `hide_output` tag.
| src/examples/Preview_and_Report_Generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="gTXsykMOc1xg"
# Importing libraries
import os
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + colab={"base_uri": "https://localhost:8080/"} id="08wQyTMz_Wqn" outputId="6bdb5d9b-002b-4b43-94f5-402b20c1acbd"
# !rm modified_public.zip
# !rm 4aaa1591-b5b2-42ce-a6f5-a2031a2439a1
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="LsX72k7Ci8EO" outputId="2447ab93-c77b-49a6-ea23-d7554c7798af"
# !wget -nc https://competitions.codalab.org/my/datasets/download/4aaa1591-b5b2-42ce-a6f5-a2031a2439a1 -O modified_public.zip
# !unzip -o modified_public.zip
# !ls
# + colab={"base_uri": "https://localhost:8080/"} id="IudSeJ_-jFwo" outputId="f0cf6d80-a361-4f1c-f765-c844c449fbb3"
# !mv modified_public/* .
# !ls
# + id="OlB6Gm9MV_yJ"
class_names = ['airplane', 'car', 'bird', 'cat', 'deer', 'dog', 'horse', 'ship', 'truck']
class_names_label = {class_name:i for i, class_name in enumerate(class_names)}
nb_classes = len(class_names)
IMAGE_SIZE = (32, 32)
# + [markdown] id="waWjT1jwV_yJ"
# ## Loading the Data
# We have to write a load_data function that load the images and the labels from the folder.
# + id="ISS7wzLkV_yJ"
def load_data():
"""
Load the data:
- 14,034 images to train the network.
- 3,000 images to evaluate how accurately the network learned to classify images.
"""
datasets = ['./xs.npy', './ys.npy']
output = []
images = []
labels = []
print("Loading {}".format(datasets[0]))
images = np.load(datasets[0])
print("Loading {}".format(datasets[1]))
labels = np.load(datasets[1])
output.append((images, labels))
return output
# + colab={"base_uri": "https://localhost:8080/"} id="SwUawFEqV_yJ" outputId="afef174b-5d0b-4be6-a161-392b47d44c7e"
(X, y) = load_data()[0]
print("Loading ./xt_modified.npy")
X_test = np.load('./xt_modified.npy')
# + [markdown] id="l00r1EVRV_yK"
# ## Let's explore the dataset
# ### We can ask ourselves:
#
# - How many training and testing examples do we have ?
# - What is the size of the images ?
# - What is the proportion of each observed category ?
# + colab={"base_uri": "https://localhost:8080/"} id="nn2lybi1QETE" outputId="b6efcd7c-20e0-459d-b379-fd881558a712"
n_labelled = X.shape[0]
n_unlabelled = X_test.shape[0]
print ("Number of labelled examples: {}".format(n_labelled))
print ("Number of unlabelled examples: {}".format(n_unlabelled))
print ("Each image is of size: {}".format(IMAGE_SIZE))
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="fGJ1-oCyhpIh" outputId="f50c3bed-f177-4b7d-84e2-5c9f6dc40e54"
_, labelled_counts = np.unique(y, return_counts=True)
pd.DataFrame({'train': labelled_counts},
index=class_names
).plot.bar()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 267} id="wgoYy7mDiXta" outputId="9bdd384b-eeab-4372-e5af-2ec2f5f9911c"
plt.pie(labelled_counts,
explode=([0]*nb_classes) ,
labels=class_names,
autopct='%1.1f%%')
plt.axis('equal')
plt.title('Proportion of each observed category')
plt.show()
# + id="r6OgzwXEisCC"
def display_random_image(class_names, images, labels):
"""
Display a random image from the images array and its correspond label from the labels array.
"""
index = np.random.randint(images.shape[0])
plt.figure()
plt.imshow(images[index])
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.title('Image #{} : '.format(index) + class_names[labels[index]])
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 268} id="Ssft_9zri0Qf" outputId="30f28528-5d35-407c-af17-11c832d9b2ea"
display_random_image(class_names, X, y)
# + id="5k4T_IuMjKWa"
def display_random_unlabelled_image(images):
"""
Display a random image from the images array and its correspond label from the labels array.
"""
index = np.random.randint(images.shape[0])
plt.figure()
plt.imshow(images[index])
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="wE6M2EoIjaEp" outputId="53d8da34-8292-481c-c799-99bece541a19"
display_random_unlabelled_image(X_test)
# + [markdown] id="idz2CYG7i_Ey"
# We can also display the first 25 images from the training set directly with a loop to get a better view
# + id="h3zYhbPJi0T0"
def display_examples(class_names, images, labels):
"""
Display 25 images from the images array with its corresponding labels
"""
fig = plt.figure(figsize=(10,10))
fig.suptitle("Some examples of images of the dataset", fontsize=16)
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(images[i], cmap=plt.cm.binary)
plt.xlabel(class_names[labels[i]])
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 661} id="zAWyohSXV_yL" outputId="6684be71-a4c6-416e-a536-5d5d050642ea"
display_examples(class_names, X, y)
# + id="Py238Sx5jkeS"
def display_unlabelled_examples(images):
"""
Display 25 images from the images array
"""
fig = plt.figure(figsize=(10,10))
fig.suptitle("Some examples of images of the dataset", fontsize=16)
for i in range(25):
plt.subplot(5,5,i+1)
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(images[i], cmap=plt.cm.binary)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 650} id="ap9DOifUjtqh" outputId="1c2c4742-bad6-43f3-8037-88e7b42c5e60"
display_unlabelled_examples(X_test)
# + id="ZaE4hySd9Rwd"
| notebooks/eda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Import the required libraries and files
# +
import pandas as pd
import numpy as np
import fasttext
from scipy.stats import uniform
import pickle
from tqdm import tqdm
import re
import os
from prettytable import PrettyTable
# scikit-learn
from sklearn.model_selection import StratifiedKFold, cross_val_score, cross_val_predict, cross_validate
from sklearn.model_selection import RandomizedSearchCV, GridSearchCV, train_test_split
from sklearn.linear_model import LogisticRegression
from sklearn.svm import SVC
# Tensorflow
import kerastuner as kt
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers, Sequential
from tensorflow.keras.layers import Dense
# Local Packages
import hats.config
import hats.utility as ut
from hats.data_preprocessing import Preprocessing
import hats.ml_model as ml
from hats.config import CONFIG
# Plotting
import matplotlib.pyplot as plt
import seaborn as sns
# %reload_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings('ignore')
# +
home_data = pd.read_csv('../dataset/dataset.csv', sep=';')
translate_data = pd.read_csv('../dataset/translations_data.csv', sep=';')
sms_translations_data = pd.read_csv('../dataset/sms_translations.csv', sep=',')
stop_words = []
with open('../dataset/stop_words.txt') as f:
stop_words = f.readlines()
stop_words = [word.replace('\n', '') for word in stop_words]
# +
import random
ind = [random.randint(0, home_data.shape[0]) for _ in range(10)]
home_data.iloc[ind]
# +
plt.figure(figsize=(7, 5))
ax = sns.countplot('label', data=home_data, color='lightblue')
plt.xlabel('Label', fontsize=15)
plt.ylabel('count', fontsize=15)
plt.xticks(rotation=90)
for p in ax.patches:
h = np.round(p.get_height(), decimals=2)
ax.annotate(str(h), (p.get_x() + 0.25, p.get_height() *1.005))
diff = p.get_width() - 0.35
p.set_width(0.35)
p.set_x(p.get_x() + diff * .5)
plt.show()
# -
# # Fasttext Model Training
# + tags=[]
data_preprocess: Preprocessing = Preprocessing(stop_words, sms_translations_data)
home_data_preprocessed = data_preprocess.preprocessing(home_data.copy())
data_preprocess.saveToCsv(home_data_preprocessed)
# Process the output file to remove double quotes ("")
# !sed -i 's/"//g' ../output/comm_preprocessed.txt
# -
# Train a fasttext model in supervised fashion
ft_model = ml.createFasttextModel(CONFIG.OUTPUT_DATASET_FILE)
command = data_preprocess.strpreprocessing('mai chahta hu ki tum geyser band kr do')
print(command)
ft_model.predict(command)
ft_model.get_sentence_vector('mai chahta hu ki tum geyser band kr do').shape
# # Create additional columns to preprocessed dataset
# ## 1. Create <i><b>sent_vec</b></i> column in main dataset for sentence vectors
home_data_vectorized = data_preprocess.convertCommandToVector(home_data_preprocessed, ft_model)
home_data_vectorized.head(5)
# ## 2. Add a column for each class using OVR scheme
#
# After adding the columns, create a single layer perceptron model with 150 inputs and 1 output with sigmoid activation.
#
# Total number of such models will be equal to the number of classes in the dataset. This is to train multiple models using the OVR technique and while predicting, we will use all the models to predict the final class label of the test command.
home_data_ovr = ut.add_class_ovr_cols(home_data_vectorized.copy())
# # Tensorflow Model
# +
# ml.nn_tune_train(data=home_data_ovr, model_names=home_data_ovr['label'].unique())
# -
# Create the models for each class
models = ml.createPerceptronModels(home_data_ovr['label'].unique())
# +
# Compile each model
# for m_name in models.keys():
# models[m_name]['model'].compile(optimizer=keras.optimizers.Adam(), loss='binary_crossentropy', metrics=['accuracy', ml._f1_score])
models = ml.compileModels(models)
# -
models = ml.nn_modelTrain(data=home_data_ovr, models=models)
# #### Plotting the models loss and accuracy
fig = ut.plot(models)
fig.savefig('fig.png', quality=95, bbox_inches='tight', pad_inches=0.5)
ml.print_nn_results(models)
# +
test_command = 'panka on mat kar'
# prediction = ml.nn_modelPredict(test_command, ft_model, data_preprocess, models)
prediction = ml.nn_modelPredict(test_command, ft_model, data_preprocess, model_names=list(models.keys()))
# -
# # Scikit-learn models
# Grid search using SVC
train_df, test_df = ut.data_split(home_data_ovr, test_size=0.3)
X_train, y_train = train_df['sent_vec'].tolist(), train_df['y']
X_train
classifiers = ml.train(train_df)
results = ml.test(classifiers, test_df)
# +
accuracy_scores = {}
for clf_name in classifiers.keys():
accuracy_scores[clf_name] = np.asarray([classifiers[clf_name]['train_accuracy'], \
results[clf_name]['test_accuracy']])
accuracy_score_df = pd.DataFrame.from_dict(data=accuracy_scores, orient='index', columns=['train', 'test'])
ax = accuracy_score_df.plot(kind='bar', colormap='Paired', figsize=(10, 5))
plt.xticks(rotation=45)
plt.title(f"Model Comparison (Accuracy) dims = {CONFIG.FT_DIMS}")
plt.xlabel("ML Model")
plt.ylabel("Accuracy")
plt.show()
# -
scores['SVC'].keys()
# +
scores = ml.cross_val(classifiers, home_data_ovr, test_df)
train_table = PrettyTable()
train_table.field_names = ['Model Name', 'Precision', 'Recall', 'F1', 'Accuracy']
for name, val in scores.items():
train_table.add_row([name, np.mean(val['train_precision_macro']), np.mean(val['train_recall_macro']),
np.mean(val['train_f1_macro']), np.mean(val['train_accuracy'])])
print('Training Results'.center(105, ' '))
print(train_table)
test_table = PrettyTable()
test_table.field_names = ['Model Name', 'Precision', 'Recall', 'F1', 'Accuracy']
for name, val in scores.items():
test_table.add_row([name, np.mean(val['test_precision_macro']), np.mean(val['test_recall_macro']),
np.mean(val['test_f1_macro']), np.mean(val['test_accuracy'])])
print('\n\n')
print('Testing Results'.center(105, ' '))
print(test_table)
clf, train_precision, train_recall, train_f1, train_accuracy = [], [], [], [], []
test_precision, test_recall, test_f1, test_accuracy = [], [], [], []
for name, val in scores.items():
clf.append(name)
train_precision.append(np.mean(val['train_precision_macro']))
train_recall.append(np.mean(val['train_recall_macro']))
train_f1.append(np.mean(val['train_f1_macro']))
train_accuracy.append(np.mean(val['train_accuracy']))
test_precision.append(np.mean(val['test_precision_macro']))
test_recall.append(np.mean(val['test_recall_macro']))
test_f1.append(np.mean(val['test_f1_macro']))
test_accuracy.append(np.mean(val['test_accuracy']))
score_dict = {'Classifier': clf, 'Train Precision': train_precision, 'Train Recall': train_recall,
'Train F1': train_f1, 'Train Accuracy': train_accuracy, 'Test Precision': test_precision,
'Test Recall': test_recall, 'Test F1': test_f1, 'Test Accuracy': test_accuracy}
score_df = pd.DataFrame.from_dict(score_dict)
# sns.barplot(x='Classifier', y=['Train Precision', 'Train Recall', 'Train F1', 'Train Accuracy'], data=score_df, hue='Classifier')
sns.barplot(x='Classifier', y='Train Precision', data=score_df)
# -
score_df.to_csv('../dataset/scores.csv', index=False)
scores = ml.cross_val(classifiers, train_df, test_df)
scores
# ### Show best estimators for all the models
for clf_name in classifiers:
print(classifiers[clf_name]['best_estimators'])
ml.cross_validate(classifiers, X_train, X_test, y_train, y_test)
test_command = 'gayser on karo ho'
test_command_preprocessed = data_proprocess.strpreprocessing(test_command)
print(f"Test Command: {test_command_preprocessed}")
clf = pickle.load(open(classifiers['SVC']['filename'], 'rb'))
for clf_name in classifiers:
prediction: str = ml.predict(test_command_preprocessed, ft_model, classifiers[clf_name]['filename'])
print(f"{clf_name} - predicted '{prediction}'")
ft_model.get_nearest_neighbors('giser', k=4)
ft_model.predict(test_command)
| app/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 ('base')
# language: python
# name: python3
# ---
# # Exercises for pricing derivatives on a Binomial Tree FINA60211(A)
#
# Pour la version française de ce fichier, [cliquez ici](bino_exercises_fr.ipynb).
#
# This interactive notebook and the associated `pedagogical_binomial_model` python module provide you with tools to generate your own exercises in pricing options on a binomial tree. You can use this notebook to:
#
# - learn to construct forward binomial trees
# - learn to calculate probabilities of states on a binomial tree
# - learn to price options by recursive single-period risk-neutral pricing and discounting
# - learn to price European options by calculating risk-neutral expectations of the terminal value
#
# The module `pedagogical_binomial_model` follows the conventions and notation of our course slides.
# ## Working with the module
#
# In order to learn how to use the module, you can simply read the documentation in the `pedagogical_binomial_model.py` file or try the following `help` call:
import pedagogical_binomial_model as pbm
help(pbm.binomial_tree)
# ## Examples
#
# ### Create a binomial tree for the stock price
#
# Create a tree to describe the evolution of the stock price:
#
# - for three months
# - in one-month steps
#
# Use the following parameters for the tree:
#
# - $S_0$ = 100
# - $r_f$ = 2%
# - $\delta$ = 1%
# - $\sigma$ = 20%
# +
stock_price = 100
number_of_model_steps = 3 # three months
length_of_model_step = 1/12 # a month is 1/12 years
risk_free_rate = 0.02
dividend_yield = 0.01
stock_volatility = 0.2 # annualized
stock_price_tree = pbm.binomial_tree(stock_price, number_of_model_steps, length_of_model_step, risk_free_rate, dividend_yield, stock_volatility)
# -
# To inspect the tree you can use the associated `print` method. The periods of the tree are indexed from $0$ to `number_of_model_steps`. Let's have a look at the output at time $1$.
#
# Note that the periods to be printed have to be given as a `list`: for example, `[0, 1, 2]`.
#
# In the output below you'll see that
#
# - Periods are numbered and separated by headers
# - For each period, all possible stock price states are printed
# - All states are named following the convention that says how many `U` and `D` moves are required to reach a given state, for example `U1-D0` for the "up" state at period $1$ and `U0-D1` for the "down" state at period 1.
#
# There is extra information in the output:
#
# - The **"Up" Transition Probability**: $p^{\star}$ from our slides
# - The **Probability of State** : the probability of achieving a given state via all possible trajectories
# - The **Multi-Period Discount Factor**: the discount factor **from** the period in the header **to** the terminal period; $e^{-r_f (K-k) h}$ where the terminal period is $K$ and the current period is $k$.
# - The **Single-Period Discount Factor**: $e^{-r_f h}$
# - The "Up" and "Down" factors are $u$ and $d$.
stock_price_tree.print([1])
# ### Exercise:
#
# Go back to your slides and use the stock price parameters from the preceding section to build a forward binomial tree by hand. Then use this module to check your results.
# ### Price a European put option
#
# You can use the model to price a European put option. The idea is to first construct the tree that describes the evolution of the stock price and then price the option *on that tree*.
#
# This means that one of the option's key parameters, **time to maturity**, depends on the tree!
#
# Let's price an option with the following parameters:
#
# - Expiration in six months
# - Strike price of \$95
#
# #### Define the stock price tree
#
# Our existing binomial tree, `stock_price_tree` describes the evolution of the price over three months. Therefore, we **cannot** use it to price the option, and we have to create a new tree.
#
# We will stick to a three-period tree. This means that we need two-month periods, i.e., $h = 2/12$ on the tree.
# +
stock_price = 100
number_of_model_steps = 3 # three steps
length_of_model_step = 2/12 # a month is 1/12 years
risk_free_rate = 0.02
dividend_yield = 0.01
stock_volatility = 0.2 # annualized
stock_price_tree = pbm.binomial_tree(stock_price, number_of_model_steps, length_of_model_step, risk_free_rate, dividend_yield, stock_volatility)
# -
# #### Define the option
#
# To define a European put option, we will use the `european_put` class from the `pbm` module. Here's the call to `help(pbm.european_put)`.
#
# An `european_put` is a `derivative`, so if you're interested, you might want to follow up by reading `help(pbm.derivative)` and the code.
help(pbm.european_put)
# The above help listing tells us how to define an `european_put`:
# ```
# | european_put(strike_price)
# ...
# | Methods defined here:
# |
# | __init__(self, strike_price)
# | Define an European Put
# |
# | Parameters
# | ----------
# | strike_price : float
# | strike price of the option
# ```
#
# Simply, set a `strike_price` and write `my_option = pbm.european_put(strike_price)`.
strike_price = 95
my_euro_put = pbm.european_put(strike_price)
# #### Price the option
#
# Having defined the stock price tree and the option, we can price the option.
#
# We do it by using the `pricing()` method of the `stock_price_tree` on the option.
my_euro_put = stock_price_tree.pricing(my_euro_put)
# Seemingly, nothing happened.
#
# However, `my_euro_put` now contains a full binomial tree with all incremental pricing information!
#
# The `derivative` class and all related classes (i.e., European and American calls and puts in `pbm`) have a `print` method which is identical to the `print` method for the binomial tree that we examined above.
#
# To learn about the put's value at time $0$, you can simply type `my_euro_put.print([0])` and see in the field `"Derivative"` that the option is worth $3.1966.
my_euro_put.print([0])
# #### Exercise: Price the option using the expected terminal payoff
#
# For European derivatives we can use the alternative approach to pricing. With $K$ periods of length $h$ years, the price of a derivative $G(S_K)$ is:
# $$
# P = e^{-r_f \times h \times K}E^{\star}\left[ G(S_K) \right]\, ,
# $$
# the risk-neutral expectation of the payoff, discounted at the risk-free rate.
#
# To see the probabilities of the terminal states together with the stock prices in these states, type `my_euro_put.print([3])`.
my_euro_put.print([3])
# In the `"Derivative"` field you can see the values of the payoff $G(S_K)$ which here is $\max(95 - S_K, 0)$. Then use the `Probability of State` and appropriate `Multi-Period Discount Factor` from period $0$ to period $K$ to calculate the price of the European Put with pen and paper.
# #### Extracurricular (for the nerds)
#
# You can recover all the information that is printed on the tree to use it as data in a computer program. The following code snippets extract the risk-neutral probabilities of the states at the final node of the tree, and the corresponding payoffs.
#
# In the end, the `.trunk` attribute of the option's `pricing_tree` is a `dictionary`.
type(my_euro_put.pricing_tree.trunk)
# +
import numpy as np
rn_probs = [my_euro_put.pricing_tree.trunk["Period 3"][state]["Probability of State"] for state in my_euro_put.pricing_tree.trunk["Period 3"].keys()]
rn_probs = np.array(rn_probs)
print("These are the risk-neutral probabilities converted to a numpy array")
print(rn_probs)
# +
payoffs = [my_euro_put.pricing_tree.trunk["Period 3"][state]["Derivative"] for state in my_euro_put.pricing_tree.trunk["Period 3"].keys()]
payoffs = np.array(payoffs)
print("These are the put's payoffs")
print(payoffs)
# -
# Now you can calculate the risk-neutral expectation and discount it to period $0$:
# +
put_price = np.sum(rn_probs * payoffs)
put_price = my_euro_put.pricing_tree.trunk["Period 0"]["State 0"]["Multi-Period Discount Factor"] * put_price
print("The price of the put option is ${0:1.4f}.".format(put_price))
# -
# ### Price an American put option and compare with the European option
#
# The American option is more *optional* than the European option, because it can be exercised any time before expiration. If it's more optional... then its value should be **higher** than the European option's.
#
# The difference between the prices of otherwise identical American and European options is called the **early exercise premium**.
#
# We can use our binomial model to price the American option estimate the premium.
#
# An american put option can be instantiated from the class `american_put`. We will use the same tree and strike price as before.
my_amer_put = pbm.american_put(95)
my_amer_put = stock_price_tree.pricing(my_amer_put)
my_amer_put.print([0])
my_euro_put.print([0])
# The price of the American put is \$3.2435 and the price of the European put is \$3.1966.
#
# The **early exercise premium** is equal to \$0.0469!
# ## Further exercises
#
# Play around with this module. Use it to answer the following questions:
#
# ### Exercise 1
#
# What is the early exercise premium for call options if $\delta = 0 $?
#
# ### Exercise 2
#
# What is the early exercise premium for put options if $r = 0$?
#
# ### Exercise 3
#
# What happens with the price of an European option if the binomial model's time grid gets *denser"?
#
# Through this we mean that we could keep the model's horizon constant but describe it in a larger number of steps. Above, we used three steps for a six-month horizon. What if we used six (i.e., monthly steps)? What if we used 180 (i.e., daily steps)?
#
# First, define a tree where `length_of_model_step = 1/365` and the `number_of_model_steps = 180`.
#
# Then, write a function which evaluates the Black-Scholes option pricing formula and compare the prices from the formula to the prices from a binomial model with very many steps (say, 1 step per day).
#
# Price a European option (call or put) with both methods.
#
# ### Exercise 4
#
# The prices of European put and call options **with the same strike and maturity** are strongly related to each other! This relation is called the **put-call parity**. It can be stated as follows:
# $$
# C - P = S e^{-(r_f - \delta)T} - K \times e^{-r_f T}\, ,
# $$
# where $C$ and $P$ are, respectively, the call and put option prices, $S$ is the stock price, $K$ is the strike of both options, $r_f$ is the risk-free rate, $\delta$ is the dividend yield, and $T$ is the maturity of both options.
#
# Use the binomial model to:
#
# 1. Demonstrate that the put-call parity relation indeed holds,
# 2. Examine how the discrepancy from the relation depends on $\delta$ for American options.
#
| bino_exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Project : Whale identification using its fluke's image
#
# The dataset has been taken from HappyWhale.com, which strives to identify whales using image agorithms. In order to attempt the same, a convolutional neural network model is described to solve this problem of fluke identification.
#
# To aid whale conservation efforts, scientists use photo surveillance systems to monitor ocean activity. They use the shape of whales’ tails and unique markings found in footage to identify what species of whale they’re analyzing and meticulously log whale pod dynamics and movements. For the past 40 years, most of this work has been done manually by individual scientists, leaving a huge trove of data untapped and underutilized.
#
# <img src="files/humpback-whale-fluke.jpg">
# +
#This cell imports all the necessary libraries needed for completing the project
import numpy as np
import pandas as pd
from PIL import Image
from glob import glob
from keras.preprocessing.image import ImageDataGenerator
from keras.applications.inception_v3 import InceptionV3
from keras.layers import Dense,MaxPooling2D,Conv2D,Activation,Flatten
from keras.models import Sequential
from keras import backend as K
from sklearn.preprocessing import OneHotEncoder,LabelEncoder
import matplotlib.pyplot as plt
# +
train_images = glob("train/*jpg")
test_images = glob("test/*jpg")
data = pd.read_csv("train.csv")
data["Image"] =data["Image"].map( lambda x : "train/"+x)
ImageToLabelDict = dict(zip(data["Image"],data["Id"]))
# +
#Image are imported with a resizing
def ImportImage(file):
img = Image.open(file).convert("RGB").resize((139,139))
return np.array(img)
train_img = np.array([ImportImage(img) for img in train_images])
X = train_img
# +
#Displaying image of desired index
plt.imshow(X[9])
# -
print("The number of unique whale ID's : ", len(data['Id'].unique()))
data['Id'].value_counts()[:100]
#Label encoding and One Hot Encoding for the training labels
lenc=LabelEncoder()
oenc=OneHotEncoder()
ly=lenc.fit_transform(data['Id']).reshape((-1,1))
y=pd.get_dummies(ly[:,0])
# +
#Since the training data is pretty less, we will use data augmentation to generate more data
print("The shape corresponds to (m,nh,nw,nc) : ", X.shape)
print("Number of training examples = ",X.shape[0])
X = X.reshape((-1,139,139,3))
input_shape = X[0].shape
X_train = X.astype("float32")
y_train = np.array(y)
Gen = ImageDataGenerator(
rescale=1./255,
rotation_range=15,
width_shift_range=.15,
height_shift_range=.15,
horizontal_flip=True)
#Fitting the ImageDataGenerater
Gen.fit(X_train, augment=True)
# -
# ### Now, we will create the model to be fitted on the training set. We will be using some Convolutional, Maxpooling and Inception blocks for this
#
# <img src="files/basic_inception_model.png">
#
# The above image shows a basic inception block which follows the "do all" approach. Different filter sizes are used for Convolution and a MaxPooling layer is also used. The outputs from al these are stacked to portray as a single layer result
# +
def InceptionModel():
model=Sequential()
# model.add(InceptionV3(weights='imagenet', include_top=True,input_shape=input_shape))
model.add(Conv2D(50,kernel_size=(3,3),strides=1,activation='relu',input_shape=input_shape))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Conv2D(50,kernel_size=(3,3),strides=1,activation='relu'))
model.add(MaxPooling2D(pool_size=(6,6)))
model.add(Conv2D(50,kernel_size=(3,3),strides=1,activation='relu'))
model.add(MaxPooling2D(pool_size=(6,6)))
model.add(Flatten())
model.add(Dense(4251,activation='softmax'))
return model
# +
inception=InceptionModel()
inception.compile(loss='categorical_crossentropy',
optimizer='Adam',
metrics=['accuracy'])
inception.summary()
# +
batch_size=15
epochs=5
inception.fit_generator(Gen.flow(X_train, y_train, batch_size=batch_size),
steps_per_epoch= X_train.shape[0]//batch_size,
epochs=epochs,
verbose=1)
| Whale identification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/utkarshpaliwal9/Anomaly-Detection/blob/master/Anomaly%20Detection.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="BIt15D3m3cun" colab_type="code" outputId="d7fdcd15-6817-410b-c26b-3b40fcb089e3" colab={"base_uri": "https://localhost:8080/", "height": 121}
from google.colab import drive
drive.mount('/content/drive')
# + id="Ug7YMTZ7VELX" colab_type="code" outputId="5ba2086f-9225-4921-a784-0ac96bb05fa0" colab={"base_uri": "https://localhost:8080/", "height": 222}
# # !wget "https://uc56f5099bb273efb6ae5f378ae8.dl.dropboxusercontent.com/cd/0/get/AwMS2jll2ZsP5gXt8WYNIgJbnEJbN2IDPlt7i8s3RE2wvZ-MLknoXSxdI4SY68PjsyuZeJ8hS4_Umu0YG5A6t8qnf7FDiE5NbOwLTKR9m-Op5A/file?_download_id=26842697584487140544307203506444460731291696667291107310867088734785&_notify_domain=www.dropbox.com&dl=1"
# + id="DbfYsvAHTiVn" colab_type="code" outputId="6cd76249-fcce-40f4-c652-d60c3066f012" colab={"base_uri": "https://localhost:8080/", "height": 390}
# # !wget --no-check-certificate "https://uc50213ee5ba4f5b53184877b462.dl.dropboxusercontent.com/cd/0/get/AwOMV7HYB99EP3qJfiNjSPSS6XXB-FMWrLF4m3MJdBo_0-XUlCCxZTgjA82ko__EvSyFv5CdudBGGSibF9XRYWQfe-D1VnlNTda562xwH1IHww/file?_download_id=506935986253414526729073992478660161518623505098054952831654550397&_notify_domain=www.dropbox.com&dl=1"
# + id="HMejaA1PwYGY" colab_type="code" outputId="3e48cbbc-6479-46c7-c579-db51a0ccfadf" colab={"base_uri": "https://localhost:8080/", "height": 222}
# # !wget --no-check-certificate "https://uc35d8e1ab645d362f5e62baef07.dl.dropboxusercontent.com/cd/0/get/AwP1ZZq4MZP3D4_Qmd0bKaDgOnn1E_08Q59OciHYoavjk1pBc8cGbrlcZrbEkasy0ZNlY9pjMBoIe60x-SYEhPiQnCnP6yvc1AMW1ixcSjWiyA/file?_download_id=227332736045446681711471496235872421701305790600816212777490604019&_notify_domain=www.dropbox.com&dl=1"
# + id="3ZhdieoseHXj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 726} outputId="4483cf22-dadd-4f90-af47-bb051e509ace"
# !wget "https://uc2a897141eb02365a41bcc45e32.dl.dropboxusercontent.com/cd/0/get/AwOK07zn2vhdSuuP42CCOHMn4Xrd4DWjeoD13ul1CI6BMfeqh3UhrSlUfQoimfJx44F4xlBhhAOGS2VrJNbUN9qI1p1c3N9R0qzcvpSmcb2KHA/file?_download_id=41246243229563876909263183277312253646512224200655360698624314548&_notify_domain=www.dropbox.com&dl=1"
# + id="V4AXZW4wUKSd" colab_type="code" colab={}
import torch
from torch.autograd import variable
import cv2
import data
import imageio
# + colab_type="code" id="bGnpDranOCND" colab={}
# !cp '/content/drive/My Drive/Colab Notebooks/anomaly_videos2' anomaly_videos2
# + id="HVwnjvFNODNb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="0c78b58f-623a-4864-fd40-c07df40a09ab"
with open ('/content/drive/My Drive/Colab Notebooks/anomaly_videos2', 'w') as anomaly_videos2:
# !unzip anomaly_videos2
# + id="dsrv1BmNLqfr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="37cf7665-1b5f-42f5-d5f3-2aac7c8408eb"
with open('/content/drive/My Drive/Project/Anomaly-Videos-Part-1/Abuse/Abuse001_x264.mp4') as video:
print(type(video))
# + id="kiCI_mmda2V6" colab_type="code" colab={}
| Anomaly Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Code to run the analysis where we test whether moral opportunists switch between guilt- and inequity-averse brain patterns
import warnings, os, sys, glob, matplotlib, scipy
warnings.filterwarnings("ignore", message="numpy.dtype size changed")
matplotlib.rcParams['pdf.fonttype'] = 42 #To make text readable by Illustrator
import numpy as np
import pandas as pd
import scipy.stats as ss
from matplotlib import pyplot as plt
import seaborn as sns
from nltools.data import Brain_Data, Adjacency
from nltools.mask import expand_mask, collapse_mask
from nltools.stats import threshold, one_sample_permutation, two_sample_permutation, correlation_permutation
from nilearn.plotting import plot_roi
base_dir = '/project/3014018.02/analysis_mri/DataSharingCollection/'
sys.path.append('%sCode/Tools/'%base_dir)
import FigureTools;
# ## Load behavioral clusters
fittedParams = pd.read_csv(os.path.join(base_dir,
'Results/2.Behavior-ClusterModel/ParticipantClustering.csv'),index_col=0)
print fittedParams.head()
subNums = np.array(fittedParams['sub'].unique())
print subNums
print len(subNums)
fittedParams['ClustName'].value_counts()
# ## Load parcellation
nparcel = 200
roi = Brain_Data(os.path.join(base_dir,'Data/6.Parcellation/whole_brain_cluster_labels_PCA=100_k=%s.nii.gz' % nparcel))
roi.plot()
roi_x = expand_mask(roi)
# ## Load GA/IA clustering results
stats_all = pd.read_csv(os.path.join(base_dir,
'Results/4.fMRI-PatternClustering/PatternConjunctionStats.csv'),index_col=0)
stats_all.head()
# +
# Select active parcels
parcels = np.unique(stats_all.loc[(stats_all['clust']!='MO'),'parcel'])
parcelsGA = np.unique(stats_all.loc[(stats_all['clust']=='GA'),'parcel'])
parcelsIA = np.unique(stats_all.loc[(stats_all['clust']=='IA'),'parcel'])
print parcels
print parcelsGA, parcelsIA
totalROI = collapse_mask(roi_x[parcels],auto_label=False)
totalROI.plot();
parcelsCommon = np.intersect1d(parcelsGA,parcelsIA)
parcelsGAonly = np.setdiff1d(parcelsGA,parcelsIA)
parcelsIAonly = np.setdiff1d(parcelsIA,parcelsGA)
# -
# ## Load MO and GAIA beta maps
GAnums = np.array(fittedParams.loc[fittedParams['ClustName']=='GA','sub'])
IAnums = np.array(fittedParams.loc[fittedParams['ClustName']=='IA','sub'])
MOnums = np.array(fittedParams.loc[fittedParams['ClustName']=='MO','sub'])
print len(GAnums),len(IAnums),len(MOnums)
# Load beta maps
dat_GA_x2 = Brain_Data()
dat_GA_x4 = Brain_Data()
dat_GA_x6 = Brain_Data()
dat_IA_x2 = Brain_Data()
dat_IA_x4 = Brain_Data()
dat_IA_x6 = Brain_Data()
dat_MO_x2 = Brain_Data()
dat_MO_x4 = Brain_Data()
dat_MO_x6 = Brain_Data()
screen = 'Mult'
for sub in GAnums:
dat_GA_x2 = dat_GA_x2.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X2')))).mean())
dat_GA_x4 = dat_GA_x4.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X4')))).mean())
dat_GA_x6 = dat_GA_x6.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X6')))).mean())
for sub in IAnums:
dat_IA_x2 = dat_IA_x2.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X2')))).mean())
dat_IA_x4 = dat_IA_x4.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X4')))).mean())
dat_IA_x6 = dat_IA_x6.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X6')))).mean())
for sub in MOnums:
dat_MO_x2 = dat_MO_x2.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X2')))).mean())
dat_MO_x4 = dat_MO_x4.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X4')))).mean())
dat_MO_x6 = dat_MO_x6.append(Brain_Data(glob.glob(os.path.join(base_dir,
'Data/5.fMRI-BetaMaps/p%s_%s%s*.nii'%(sub,screen,'X6')))).mean())
# Standardize per parcel per subject
dat_GA_x2_zscore = dat_GA_x2.copy()
dat_GA_x4_zscore = dat_GA_x4.copy()
dat_GA_x6_zscore = dat_GA_x6.copy()
dat_IA_x2_zscore = dat_IA_x2.copy()
dat_IA_x4_zscore = dat_IA_x4.copy()
dat_IA_x6_zscore = dat_IA_x6.copy()
dat_MO_x2_zscore = dat_MO_x2.copy()
dat_MO_x4_zscore = dat_MO_x4.copy()
dat_MO_x6_zscore = dat_MO_x6.copy()
parcelLocs = []
for parcel in range(200):
print parcel,
parcelLocs.append(np.where(roi_x[parcel].data==1))
print ''
for subInd,sub in enumerate(GAnums):
print sub,
subMapX2 = dat_GA_x2_zscore[subInd].copy()
subMapX4 = dat_GA_x4_zscore[subInd].copy()
subMapX6 = dat_GA_x6_zscore[subInd].copy()
subDatX2 = subMapX2.data
subDatX4 = subMapX4.data
subDatX6 = subMapX6.data
for parcel in range(200):
parcelLoc = parcelLocs[parcel][0]
subDatX2[parcelLoc] = ss.zscore(subDatX2[parcelLoc])
subDatX4[parcelLoc] = ss.zscore(subDatX4[parcelLoc])
subDatX6[parcelLoc] = ss.zscore(subDatX6[parcelLoc])
subMapX2.data = subDatX2
subMapX4.data = subDatX4
subMapX6.data = subDatX6
dat_GA_x2_zscore[subInd] = subMapX2
dat_GA_x4_zscore[subInd] = subMapX4
dat_GA_x6_zscore[subInd] = subMapX6
for subInd,sub in enumerate(IAnums):
print sub,
subMapX2 = dat_IA_x2_zscore[subInd].copy()
subMapX4 = dat_IA_x4_zscore[subInd].copy()
subMapX6 = dat_IA_x6_zscore[subInd].copy()
subDatX2 = subMapX2.data
subDatX4 = subMapX4.data
subDatX6 = subMapX6.data
for parcel in range(200):
parcelLoc = parcelLocs[parcel][0]
subDatX2[parcelLoc] = ss.zscore(subDatX2[parcelLoc])
subDatX4[parcelLoc] = ss.zscore(subDatX4[parcelLoc])
subDatX6[parcelLoc] = ss.zscore(subDatX6[parcelLoc])
subMapX2.data = subDatX2
subMapX4.data = subDatX4
subMapX6.data = subDatX6
dat_IA_x2_zscore[subInd] = subMapX2
dat_IA_x4_zscore[subInd] = subMapX4
dat_IA_x6_zscore[subInd] = subMapX6
for subInd,sub in enumerate(MOnums):
print sub,
subMapX2 = dat_MO_x2_zscore[subInd].copy()
subMapX4 = dat_MO_x4_zscore[subInd].copy()
subMapX6 = dat_MO_x6_zscore[subInd].copy()
subDatX2 = subMapX2.data
subDatX4 = subMapX4.data
subDatX6 = subMapX6.data
for parcel in range(200):
parcelLoc = parcelLocs[parcel][0]
subDatX2[parcelLoc] = ss.zscore(subDatX2[parcelLoc])
subDatX4[parcelLoc] = ss.zscore(subDatX4[parcelLoc])
subDatX6[parcelLoc] = ss.zscore(subDatX6[parcelLoc])
subMapX2.data = subDatX2
subMapX4.data = subDatX4
subMapX6.data = subDatX6
dat_MO_x2_zscore[subInd] = subMapX2
dat_MO_x4_zscore[subInd] = subMapX4
dat_MO_x6_zscore[subInd] = subMapX6
for dat in [dat_GA_x2,dat_GA_x4,dat_GA_x6,
dat_GA_x2_zscore,dat_GA_x4_zscore,dat_GA_x6_zscore,
dat_IA_x2,dat_IA_x4,dat_IA_x6,
dat_IA_x2_zscore,dat_IA_x4_zscore,dat_IA_x6_zscore,
dat_MO_x2,dat_MO_x4,dat_MO_x6,
dat_MO_x2_zscore,dat_MO_x4_zscore,dat_MO_x6_zscore,
]:
print dat.shape()
# Should be nearly 0:
print dat_IA_x2_zscore[np.random.randint(0,len(dat_IA_x2_zscore),size=1)].data[roi_x[np.random.randint(0,199,1)].data==1].mean()
print dat_GA_x6_zscore[np.random.randint(0,len(dat_GA_x6_zscore),size=1)].data[roi_x[np.random.randint(0,199,1)].data==1].mean()
print dat_MO_x2_zscore[np.random.randint(0,len(dat_MO_x2_zscore),size=1)].data[roi_x[np.random.randint(0,199,1)].data==1].mean()
# Should be non 0:
print dat_IA_x2[np.random.randint(0,len(dat_IA_x2),size=1)].data[roi_x[np.random.randint(0,199,1)].data==1].mean()
print dat_GA_x6[np.random.randint(0,len(dat_GA_x6),size=1)].data[roi_x[np.random.randint(0,199,1)].data==1].mean()
print dat_MO_x2[np.random.randint(0,len(dat_MO_x2),size=1)].data[roi_x[np.random.randint(0,199,1)].data==1].mean()
# ## Per parcel, compute similarity of MO z maps to GA-IA z-diff map
# Distance
simMethod = 'correlation'
includeX4 = False
sim_zdiffs_all = pd.DataFrame()
for parcel in parcels:
parcelDat_GA_x2_zscore = dat_GA_x2_zscore.apply_mask(roi_x[parcel])
parcelDat_GA_x4_zscore = dat_GA_x4_zscore.apply_mask(roi_x[parcel])
parcelDat_GA_x6_zscore = dat_GA_x6_zscore.apply_mask(roi_x[parcel])
parcelDat_IA_x2_zscore = dat_IA_x2_zscore.apply_mask(roi_x[parcel])
parcelDat_IA_x4_zscore = dat_IA_x4_zscore.apply_mask(roi_x[parcel])
parcelDat_IA_x6_zscore = dat_IA_x6_zscore.apply_mask(roi_x[parcel])
# Mean
mean_GA_x2 = parcelDat_GA_x2_zscore.mean()
mean_GA_x4 = parcelDat_GA_x4_zscore.mean()
mean_GA_x6 = parcelDat_GA_x6_zscore.mean()
mean_IA_x2 = parcelDat_IA_x2_zscore.mean()
mean_IA_x4 = parcelDat_IA_x4_zscore.mean()
mean_IA_x6 = parcelDat_IA_x6_zscore.mean()
# Difference
mean_zdiff_x2 = mean_GA_x2 - mean_IA_x2
mean_zdiff_x4 = mean_GA_x4 - mean_IA_x4
mean_zdiff_x6 = mean_GA_x6 - mean_IA_x6
sim_zdiff_x2 = dat_MO_x2_zscore.apply_mask(roi_x[parcel]).similarity(
mean_zdiff_x2,method=simMethod)
sim_zdiff_x4 = dat_MO_x4_zscore.apply_mask(roi_x[parcel]).similarity(
mean_zdiff_x4,method=simMethod)
sim_zdiff_x6 = dat_MO_x6_zscore.apply_mask(roi_x[parcel]).similarity(
mean_zdiff_x6,method=simMethod)
if includeX4:
sim_zdiffs = pd.DataFrame(np.transpose(np.vstack([sim_zdiff_x2,sim_zdiff_x4,sim_zdiff_x6])),
columns=['x2','x4','x6'])
else:
sim_zdiffs = pd.DataFrame(np.transpose(np.vstack([sim_zdiff_x2,sim_zdiff_x6])),
columns=['x2','x6'])
sim_zdiffs = sim_zdiffs.melt(var_name = 'Condition', value_name='GA > IA similarity')
sim_zdiffs['parcel'] = parcel
sim_zdiffs_all = sim_zdiffs_all.append(sim_zdiffs)
nMO = len(dat_MO_x2)
sim_zdiffs_all['sub'] = np.tile(range(nMO),[1,len(parcels)*(2+includeX4)]).T
sim_zdiffs_all.to_csv(os.path.join(base_dir,'Results/5.fMRI-MOswitching',
'MO_sim_to_GAIA.csv'))
# ## Load data
sim_zdiffs_all = pd.read_csv(os.path.join(base_dir,'Results/5.fMRI-MOswitching',
'MO_sim_to_GAIA.csv'),index_col=0)
sim_zdiffs_all.head()
sim_zdiffs_all.parcel.unique()
# ## Plot / test
# Plot:
sns.set_context('talk')
fig,ax = plt.subplots(1,1,figsize=[np.min([len(parcels)*4,16]),4])
sns.barplot(data=sim_zdiffs_all,x='parcel',y='GA > IA similarity',hue='Condition',ax=ax)
stats_all = pd.DataFrame(columns=['parcel','t','p','p_corr'])
for parcel in parcels:
stats = ss.ttest_rel(sim_zdiffs_all.loc[(sim_zdiffs_all['parcel']==parcel) &
(sim_zdiffs_all['Condition']=='x6'),'GA > IA similarity'],
sim_zdiffs_all.loc[(sim_zdiffs_all['parcel']==parcel) &
(sim_zdiffs_all['Condition']=='x2'),'GA > IA similarity'])
t_val = stats[0]
p_corr = stats[1]*len(parcels)
if (p_corr < 0.05) & (t_val > 0):
sigstars = '*'
else:
sigstars = ''
print 'Parcel %i, t = %.2f, p = %.4f, p-Bonferroni-corrected = %.4f%s'%(parcel,t_val,stats[1],p_corr,sigstars)
stats_all = stats_all.append(pd.DataFrame([[parcel,t_val,stats[1],p_corr]],columns=stats_all.columns))
plt.show()
# Split by GA/IA map
fig,ax = plt.subplots(1,3,figsize=[15,5],sharey=True,
gridspec_kw={'width_ratios':[float(len(parcelsGAonly))/float(len(parcels)),
float(len(parcelsCommon))/float(len(parcels)),
float(len(parcelsIAonly))/float(len(parcels))]})
sns.barplot(data=sim_zdiffs_all.loc[sim_zdiffs_all['parcel'].isin(parcelsGAonly)],
x='parcel',hue='Condition',y='GA > IA similarity',errwidth=1,capsize=.1,ax=ax[0])
ax[0].set_title('Unique GA parcels')
sns.barplot(data=sim_zdiffs_all.loc[sim_zdiffs_all['parcel'].isin(parcelsCommon)],
x='parcel',hue='Condition',y='GA > IA similarity',errwidth=1,capsize=.1,ax=ax[1])
ax[1].set_title('Common parcels')
sns.barplot(data=sim_zdiffs_all.loc[sim_zdiffs_all['parcel'].isin(parcelsIAonly)],
x='parcel',hue='Condition',y='GA > IA similarity',errwidth=1,capsize=.1,ax=ax[2])
ax[2].set_title('Unique IA parcels')
plt.tight_layout()
# ## Summarize over parcels: subject mean over parcel-wise similarities
nMO = len(dat_MO_x2)
meanSimilarities_allROIs = pd.DataFrame(columns=['sub','cond','GAsim'])
includeX4 = False
if includeX4:
conds = ['x2','x4','x6']
else:
conds = ['x2','x6']
for subInd in range(nMO):
subDat = sim_zdiffs_all.loc[(sim_zdiffs_all['sub']==subInd) & (
sim_zdiffs_all['parcel'].isin(parcels))].copy()
for cond in conds:
subSim = np.mean(subDat.loc[subDat['Condition']==cond,'GA > IA similarity'])
meanSimilarities_allROIs = meanSimilarities_allROIs.append(
pd.DataFrame([[subInd,cond,subSim]],columns=meanSimilarities_allROIs.columns))
meanSimilarities_allROIs.head()
conditionMeans = meanSimilarities_allROIs.groupby('cond').mean()
conditionMeans
# Mean difference in pattern correlation
print conditionMeans.loc['x6','GAsim']-conditionMeans.loc['x2','GAsim']
meanSimilarities_allROIs_piv = meanSimilarities_allROIs.pivot(
index='sub',columns='cond',values='GAsim').reset_index()
meanSimilarities_allROIs_piv['diff'] = meanSimilarities_allROIs_piv['x6']-meanSimilarities_allROIs_piv['x2']
fig,ax = plt.subplots(1,2,figsize=[6,5])
sns.barplot(data=meanSimilarities_allROIs,y='GAsim',x='cond',ax=ax[0])
# for sub in range(20):
# ax[0].plot([0,1],sumSimilarities.loc[sumSimilarities['sub']==sub,'GAsim'],
# color='k',lineWidth=1,lineStyle=':')
sns.barplot(data=meanSimilarities_allROIs_piv,y='diff',alpha=.5,errwidth=1,capsize=.1,zorder=1,ax=ax[1])
sns.swarmplot(data=meanSimilarities_allROIs_piv,y='diff',zorder=2,s=8,alpha=1,ax=ax[1])
plt.tight_layout()
print scipy.stats.ttest_rel(meanSimilarities_allROIs.loc[meanSimilarities_allROIs['cond']=='x2','GAsim'],
meanSimilarities_allROIs.loc[meanSimilarities_allROIs['cond']=='x6','GAsim'])
print scipy.stats.wilcoxon(meanSimilarities_allROIs.loc[meanSimilarities_allROIs['cond']=='x2','GAsim'],
meanSimilarities_allROIs.loc[meanSimilarities_allROIs['cond']=='x6','GAsim'],)
print one_sample_permutation(meanSimilarities_allROIs_piv['diff'])
# #### Simple classification as test (forced choice)
# +
pred = [np.argmax(np.array(meanSimilarities_allROIs.loc[meanSimilarities_allROIs['sub']==i,'GAsim']))
for i in meanSimilarities_allROIs['sub'].unique()]
acc = np.mean(pred)
print acc
print ss.ttest_1samp(pred,.5)
print one_sample_permutation(np.array(pred)-.5)
# -
acc*nMO
# ## Summarize over parcels: split by parcel set unique to GA/IA
nMO = len(dat_MO_x2)
meanSimilarities_perStrategy = pd.DataFrame(columns=['sub','map','cond','GAsim'])
for subInd in range(nMO):
for parcelSetName, parcelSet in zip(['GA','IA'],[parcelsGAonly,parcelsIAonly]):
subDat = sim_zdiffs_all.loc[(sim_zdiffs_all['sub']==subInd) & (
sim_zdiffs_all['parcel'].isin(parcelSet))].copy()
for cond in ['x2','x6']:
subSim = np.mean(subDat.loc[subDat['Condition']==cond,'GA > IA similarity'])
meanSimilarities_perStrategy = meanSimilarities_perStrategy.append(
pd.DataFrame([[subInd,parcelSetName,cond,subSim]],columns=['sub','map','cond','GAsim']))
meanSimilarities_perStrategy.head()
meanSimilarities_perStrategy_piv = meanSimilarities_perStrategy.pivot_table(
index=['sub','map'],columns='cond',values='GAsim').reset_index()
meanSimilarities_perStrategy_piv['diff'] = meanSimilarities_perStrategy_piv['x6']-meanSimilarities_perStrategy_piv['x2']
meanSimilarities_perStrategy_piv.head()
x2All = scipy.stats.ttest_1samp(meanSimilarities_allROIs.loc[meanSimilarities_allROIs['cond']=='x2','GAsim'],0)
x6All = scipy.stats.ttest_1samp(meanSimilarities_allROIs.loc[meanSimilarities_allROIs['cond']=='x6','GAsim'],0)
x2GAdat = meanSimilarities_perStrategy.loc[(meanSimilarities_perStrategy['cond']=='x2')
& (meanSimilarities_perStrategy['map']=='GA'),'GAsim']
x2GA = scipy.stats.ttest_1samp(x2GAdat,0)
x6GAdat = meanSimilarities_perStrategy.loc[(meanSimilarities_perStrategy['cond']=='x6')
& (meanSimilarities_perStrategy['map']=='GA'),'GAsim']
x6GA = scipy.stats.ttest_1samp(x6GAdat,0)
x2IAdat = meanSimilarities_perStrategy.loc[(meanSimilarities_perStrategy['cond']=='x2')
& (meanSimilarities_perStrategy['map']=='IA'),'GAsim']
x2IA = scipy.stats.ttest_1samp(x2IAdat,0)
x6IAdat = meanSimilarities_perStrategy.loc[(meanSimilarities_perStrategy['cond']=='x6')
& (meanSimilarities_perStrategy['map']=='IA'),'GAsim']
x6IA = scipy.stats.ttest_1samp(x6IAdat,0)
print x2All
print x6All
print np.mean(x2GAdat),x2GA
print np.mean(x6GAdat),x6GA
print np.mean(x2IAdat),x2IA
print np.mean(x6IAdat),x6IA
diffAll = scipy.stats.ttest_rel(meanSimilarities_allROIs.loc[meanSimilarities_allROIs['cond']=='x6','GAsim'],
meanSimilarities_allROIs.loc[meanSimilarities_allROIs['cond']=='x2','GAsim'])
diffGA = scipy.stats.ttest_rel(meanSimilarities_perStrategy.loc[(meanSimilarities_perStrategy['cond']=='x6')
& (meanSimilarities_perStrategy['map']=='GA'),'GAsim'],meanSimilarities_perStrategy.loc[
(meanSimilarities_perStrategy['cond']=='x2') & (meanSimilarities_perStrategy['map']=='GA'),'GAsim'])
diffIA = scipy.stats.ttest_rel(meanSimilarities_perStrategy.loc[(meanSimilarities_perStrategy['cond']=='x6')
& (meanSimilarities_perStrategy['map']=='IA'),'GAsim'],meanSimilarities_perStrategy.loc[
(meanSimilarities_perStrategy['cond']=='x2') & (meanSimilarities_perStrategy['map']=='IA'),'GAsim'])
print diffAll
print (np.mean(x6GAdat)-np.mean(x2GAdat)),diffGA
print (np.mean(x6IAdat)-np.mean(x2IAdat)),diffIA
# ##### Figure 5
# +
import matplotlib
matplotlib.rcParams['pdf.fonttype'] = 42
sns.set_context('talk')
sns.set_style('white')
sns.set_palette([sns.color_palette('tab10',3)[i] for i in [2,1]])
fig,ax = plt.subplots(1,2,figsize=[10,6],gridspec_kw={'width_ratios':[1,2]},sharey=False)
sns.barplot(data=meanSimilarities_allROIs,y='GAsim',x='cond',ax=ax[0],
errcolor='k',errwidth=1,capsize=.1)
for sub in range(nMO):
ax[0].plot([0,1],meanSimilarities_allROIs.loc[meanSimilarities_allROIs['sub']==sub,'GAsim'],
color='k',lineWidth=1,lineStyle=':')
ax[0].set(xlabel='Condition',ylabel='Mean similarity to GA > IA difference map',
title='Mean over all ROIs')
sns.barplot(data=meanSimilarities_perStrategy,y='GAsim',x='map',hue='cond',ax=ax[1],
errcolor='k',errwidth=1,capsize=.1)
for sub in range(nMO):
for x,parcelSetName in zip([0,1],['GA','IA']):
ax[1].plot([x-.2,x+.2],meanSimilarities_perStrategy.loc[(meanSimilarities_perStrategy['sub']==sub) & (
meanSimilarities_perStrategy['map']==parcelSetName),'GAsim'],color='k',lineWidth=1,lineStyle=':')
ax[1].set(xlabel='Strategy map',ylabel='Mean similarity to GA > IA difference map',
title='Mean over strategy-specific ROIs')
FigureTools.add_sig_markers(ax[0],relationships=[[0,0,x2All[1]]],ystart = .3)
FigureTools.add_sig_markers(ax[0],relationships=[[1,1,x6All[1]]],ystart = .3)
FigureTools.add_sig_markers(ax[0],relationships=[[0,1,diffAll[1]]],linewidth=1,ystart=.4)
FigureTools.add_sig_markers(ax[1],relationships=[[-.2,-.2,x2GA[1]]],ystart = .5)
FigureTools.add_sig_markers(ax[1],relationships=[[.2,.2,x6GA[1]]],ystart = .5)
FigureTools.add_sig_markers(ax[1],relationships=[[.8,.8,x2IA[1]]],ystart = .4)
FigureTools.add_sig_markers(ax[1],relationships=[[1.2,1.2,x6IA[1]]],ystart = .4)
FigureTools.add_sig_markers(ax[1],relationships=[[-.2,.2,diffGA[1]]],linewidth=1,ystart=.6)
FigureTools.add_sig_markers(ax[1],relationships=[[.8,1.2,diffIA[1]]],linewidth=1,ystart=.5)
#
# Equalize ylim
ylim0 = ax[0].get_ylim()
ylim1 = ax[1].get_ylim()
ylim = [np.min([ylim0[0],ylim1[0]]),np.max([ylim0[1],ylim1[1]])]
ax[0].set_ylim(ylim); ax[1].set_ylim(ylim);
plt.suptitle('Pattern similarity of MO participants to GA versus IA',y=1.02)
plt.tight_layout();
plt.savefig(os.path.join(base_dir,'Results/5.fMRI-MOswitching,
'MO-similarity_AllROIs_ByStrategy.pdf'),
transparent=True, bbox_inches='tight')
# -
| 7.fMRI-MOswitching/1.MOswitching.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework Part 1
# ---
# - Build and train a MLP Model to classify Mnist dataset
#
# 1. MLP Network accepts 1D data. So we should flatten our 2D image, then print the dimension of the result arrays.
#
# 2. Normalize data by rescaling them to (0,1)
#
# 3. Convert label arrays to 1-hot representation (keras.utils.to_categorical)
#
# 4. Define Model
#
# - Hidden Layer 1: Fully Conncted + Relu Activition (e.g. 512 Nuerons)
# - Hidden Layer 2: Fully Connected + Relu Activition (e.g. 512 Neurons)
# - Output Layer: Fully Connected + Softmax Activition
#
# - Also build another model with BatchNormalization and Dropout. Compare these two CNN + MLP models performance for test data
# # Importing the Packages
# ---
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt # for plotting the digit image
# %matplotlib inline
from keras.datasets import mnist
from keras.utils import to_categorical
from keras.models import Sequential
from keras.layers import Dense, Input, Conv2D, MaxPooling2D, Flatten
from keras.optimizers import SGD
from keras.optimizers import RMSprop
from keras.initializers import RandomNormal
from keras.callbacks import TensorBoard
# # Loading the Data
# ---
(X_train, y_train), (X_test, y_test) = mnist.load_data()
# ### Check the random image and its label
# +
rand_num = np.random.randint(60000)
plt.imshow(X_train[rand_num], cmap="gray")
plt.show()
# print its label
print('label:', y_train[rand_num])
# -
# ### The Network accept 1D data. So we need to flatten our 2D image, then print the dimension of the result arrays.
X_train.shape
# ### Reshaping, Normalizing, one-hot-coding
# +
# reshape the data
# NOTE: when data is big it is better to do reshaping and normalinzing inplace, bc copying the opject takes up a lot
# of memory space
NUM_CLASSES = 10
X_train = np.reshape(X_train, [-1, 28*28]).astype('float32')
X_test = np.reshape(X_test, [-1, 28*28]).astype('float32')
# Normalize data by rescaling them to (0,1)
X_train /= 255
X_test /= 255
# Convert label arrays to 1-hot representation
y_train = to_categorical(y_train, NUM_CLASSES)
y_test = to_categorical(y_test, NUM_CLASSES)
# -
print('train shape: ', X_train.shape)
print(X_train.shape[0], 'train samples')
print(X_test.shape[0], 'test samples')
# ## Define the Model
# ---
# ### Add the following layers to the network:
#
# - Hidden Layer 1: Fully Conncted + Relu Activition (e.g. 512 Nuerons)
# - Hidden Layer 2: Fully Connected + Relu Activition (e.g. 512 Neurons)
# - Outout Layer: Fully Connected + Softmax Activition
# initialize the model
model = Sequential()
# Add the layers to model here.
model.add(Dense(512, activation='relu', input_shape=(784,), kernel_initializer=RandomNormal(0,0.01)))
model.add(Dense(512, activation='relu', kernel_initializer=RandomNormal(0,0.01)))
# Output Layer: Fully Connected + Softmax Activition
model.add(Dense(10, activation='softmax', kernel_initializer=RandomNormal(0,0.01)))
# ### Determine loss function, optimizer and metrics for the model
model.compile(loss='categorical_crossentropy',
optimizer=RMSprop(),
metrics=['accuracy'])
batch_size = 128
epochs = 5
model.fit(X_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(X_test, y_test))
# ### Print the review of the model
model.summary()
| Homework/HW-2/mlp_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: torch
# language: python
# name: torch
# ---
# +
# %load_ext autoreload
# %autoreload 2
# +
import os
import numpy as np
from torchvision import transforms
import matplotlib.pyplot as plt
import torch
import torch.nn as nn
from torch.utils.data import DataLoader
from net import BusterNet
from dataset import USCISIDataset
# +
def viz(imgs_np, simi_out_np, mani_out_np, mask_out_np, index):
fig = plt.figure(figsize=(20, 80))
img = imgs_np[index]
img = imgs_np[index] * [0.229, 0.224, 0.225] + [0.485, 0.456, 0.406]
ax = fig.add_subplot(1, 4, 1)
plt.imshow(img)
plt.title('Original Image')
simi_pred = (simi_out_np[index] * 2).astype(np.uint8)
ax = fig.add_subplot(1, 4, 2)
plt.imshow(simi_pred[:,:, 0])
plt.title('Similarity mask')
mani_pred = (mani_out_np[index] * 2).astype(np.uint8)
ax = fig.add_subplot(1, 4, 3)
plt.imshow(mani_pred[:,:, 0])
plt.title('Manipulation mask')
mask_pred = (mask_out_np[index] * 2).astype(np.uint8) * 255
ax = fig.add_subplot(1, 4, 4)
plt.imshow(mask_pred)
plt.title('Output mask')
plt.show()
# +
# Download dataset to ./datasets/
lmdb_dir = './datasets/USCISI-CMFD'
test_file = 'test.keys'
input_size = 256
transform = transforms.Compose([
transforms.ToPILImage(),
transforms.Resize((input_size, input_size)),
transforms.ToTensor(),
transforms.Normalize(mean=[0.485, 0.456, 0.406],
std=[0.229, 0.224, 0.225]),
])
target_transform = transforms.Compose([
transforms.ToPILImage(),
transforms.Resize((input_size, input_size)),
transforms.ToTensor(),
])
test_set = USCISIDataset(lmdb_dir, test_file, transform, target_transform)
test_params = {'batch_size': 16,
'shuffle': False,
'drop_last': True,
# 'collate_fn': collater,
'num_workers': 1}
test_generator = DataLoader(test_set, **test_params)
# +
data = next(iter(test_generator))
imgs, gts, _= data
# +
model = BusterNet(256)
model.load_state_dict(torch.load('trained_model/buster_epoch_13.pth'))
model.eval()
with torch.no_grad():
preds = model(imgs)
mask_out, mani_output, simi_output = preds
mask_out_np = mask_out.permute(0, 2, 3, 1).numpy()
mani_out_np = mani_output.permute(0, 2, 3, 1).numpy()
simi_out_np = simi_output.permute(0, 2, 3, 1).numpy()
# +
index = 11
imgs_np = imgs.permute(0, 2, 3, 1).numpy()
viz(imgs_np, simi_out_np, mani_out_np, mask_out_np, index)
# -
| demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5"
# Loads in numpy, pandas, and os packages
import numpy as np
import pandas as pd
import os
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a"
# loads in train data into pandas dataframe
train_data = pd.read_csv("train.csv")
# -
train_data.head()
# Creates dataframe for the test set
test_data = pd.read_csv('test.csv')
# Shows header forn training set
train_data.head()
# Changes male and female to binary values
train_data['Sex'] = train_data.Sex.map({'male': 0, 'female': 1})
# Redisplays the new dataset
train_data.head()
train_data.describe(include=['O'])
#Creates new datafram with the numeric columns
train2 = train_data[['Pclass', 'SibSp', 'Parch', 'Sex', 'Embarked']]
# shows new dataframe with the binary columns
train2.head()
train2.dropna(inplace=True)
train2.Embarked.unique()
train2['Embarked'] = train2.Embarked.map({'S':0, 'C':1, 'Q':3})
train2.head()
#Imports matplotlib and seaborn modules
import matplotlib.pyplot as plt
# %matplotlib inline
import seaborn as sns
# Creates bar chart with a trendline
sns.distplot(train_data.Age.dropna())
#displayes genders that survived
sns.scatterplot(x=train_data.Age, y=train_data.Survived, hue=train_data.Sex)
# This shows that almost all the surviors were female and very few female died
g= sns.FacetGrid(data = train_data, row = 'Embarked', col='Pclass')
g.map(plt.hist, "Survived")
#imports skleanr to create the module
from sklearn.linear_model import LogisticRegression
Regres1 = LogisticRegression()
train_data.head()
train_data.describe()
#adds Pclass, sib, and Parch to the x axis of the training dataset
X_train = train_data[['Pclass', 'SibSp', 'Parch']]
# +
#adds Pclass, sib, and Parch to the x axis of the testing dataset
X_test = test_data[['Pclass', 'SibSp', 'Parch']]
# -
# Adds survived to the train y axis
Y_train = train_data.Survived
Regres1.fit(X_train, Y_train)
#Tests the regression with the test data sets
prediction1 = Regres1.predict(X_test)
prediction1
# Creates testing dataset
test_data = pd.read_csv("test.csv")
# Shows first 5 rows
test_data.head()
test_data.shape
passId = test_data.PassengerId
submissions = pd.DataFrame({'PassengerId': passId, 'Survived':prediction1})
submissions.head()
# Prints testing to submission dataset
submissions.to_csv("titanic_prediction.csv", index=None)
| titanic/titanic_prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import seaborn as sns
data = pd.read_csv("glass.csv")
data.head()
data['Type'].unique()
sns.countplot(data["Type"])
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split, cross_val_score, KFold, ShuffleSplit, GridSearchCV
from sklearn import metrics
X = data.iloc[:,0:9]
print(X.head())
Y = data["Type"]
print(Y.head())
train_x, test_x, train_y, test_y = train_test_split(X, Y, random_state = 5, test_size = 0.30)
dt_model = DecisionTreeClassifier()
dt_model.fit(train_x, train_y)
predicted = dt_model.predict(test_x)
rms = metrics.mean_squared_error(predicted, test_y)
print(rms)
print(metrics.accuracy_score(predicted, test_y))
kf = KFold(n_splits=3)
for train_index, test_index in kf.split(X):
#rint("TRAIN:", train_x, "TEST:", test_x)
x_train, x_test = X.loc[train_index], X.loc[test_index]
y_train, y_test = Y[train_index], Y[test_index]
dt_model.fit(x_train, y_train)
predicted = dt_model.predict(x_test)
print("Accuracy Score : " + str(metrics.accuracy_score(predicted[0:], y_test.values)))
print(cross_val_score(dt_model, X, Y, cv = 3, scoring="accuracy").mean())
# +
import numpy as np
from sklearn.ensemble import RandomForestClassifier
rf_model = RandomForestClassifier(n_jobs=-1,max_features= 'sqrt' ,n_estimators=50, oob_score = True)
parameter_candidates = [{1},{2}]
param_grid = {
'n_estimators': [100, 200],
'max_features': ['auto', 'sqrt', 'log2']
}
CV_rfc = GridSearchCV(estimator=rf_model, param_grid=param_grid, cv= 5)
CV_rfc.fit(X, Y)
print(CV_rfc.best_params_)
# +
rf_model = RandomForestClassifier(n_jobs=-1,max_features= 'auto' ,n_estimators=200, oob_score = True)
cross_val_score(rf_model, X, Y, cv=10, scoring='accuracy')
# -
| 23rd June Assignments/case study 2/Case Study 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas
import seaborn as sns
from sklearn import linear_model
from sklearn.metrics import r2_score
import numpy as np
#reading the csv file
internet_dataframe = pandas.read_csv("internet.csv")
# +
#print(internet_dataframe)
# -
#save independent values in X and dependent in y
X = internet_dataframe[['Hour','Users']]
y = internet_dataframe['Speed']
# +
#print(y)
# -
sns.scatterplot(x=internet_dataframe['Hour'], y=internet_dataframe['Users'], hue=internet_dataframe['Speed'] )
# +
#create the regression model object and fit the data into it
reg_obj = linear_model.LinearRegression()
reg_obj.fit(X,y)
# -
#predict the speed at 20:00 hours in the evening and with 750 users online
predicted_speed = reg_obj.predict([[20,750]])
print("Predicted Speed : ")
print(predicted_speed)
print(reg_obj.coef_)
| multiple_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import h5py
import numpy as np
from matplotlib import pyplot as plt
from matplotlib.animation import FuncAnimation, ArtistAnimation, PillowWriter
import pandas as pd
from pandas import HDFStore,DataFrame
# ## Object Quantification
# - Example Notebook of how you would quantify segmentation images.
#Path to exported features as a csv, can also export as hdf5.
path_722 = "Finished Examples/722_features-0_table.csv"
nuc_722 = pd.read_csv(path_722)
#Only keeping Label 1 objects corresponding to the segmented nucleolus
nuc_722_f = nuc_722[nuc_722["Predicted Class"]=="Label 1"]
density = nuc_722_f["Mean Intensity_0"]
density_n = nuc_722_f["Mean Intensity in neighborhood_0"]
#Rescale intensity values to physical optical path length values [0.5,2]
nuc_722_f["Mean Intensity_0"] = np.interp(density, (density.min(), density.max()), (-0.5, +2))
nuc_722_f["Mean Intensity in neighborhood_0"] = np.interp(density_n, (density_n.min(), density_n.max()), (-0.5, +2))
#Calculate Aspect Ratio (Width/Height)
nuc_722_f["Aspect Ratio"] = nuc_722_f["Radii of the object_0"]/nuc_722_f["Radii of the object_1"]
#Only keeping columns of interest
nuc_722_f = nuc_722_f[['timestep', 'Size in pixels', 'Aspect Ratio',
'Mean Intensity_0', 'Mean Intensity in neighborhood_0']]
#Converting size to microns
nuc_722_f["Size in pixels"] = nuc_722_f["Size in pixels"]*(6.45/63)
nuc_722_f.head()
nuc_722_f_g = nuc_722_f.groupby("timestep")
#Extracting data from pandas dataframe to a dictionary.
datamatrix = {}
for time in np.arange(60):
data = nuc_722_f_g.get_group(time).to_numpy()[:,1:]
datamatrix[time] = data
#Automatically finding good bins.
_, bins0 = np.histogram(datamatrix[0][:,0])
_, bins1 = np.histogram(datamatrix[0][:,1])
_, bins2 = np.histogram(datamatrix[0][:,2])
_, bins3 = np.histogram(datamatrix[0][:,3])
# ## Making Animated Histograms of Object measurements over time
# - Requirements: ffmpeg
# - `brew install ffmpeg`
# - check that `which ffmpeg` returns '/usr/local/bin/ffmpeg' if not replace below with the path to ffmpeg on your computer.
plt.rcParams['animation.ffmpeg_path'] = '/usr/local/bin/ffmpeg'
# +
fig, ax = plt.subplots(nrows=2,ncols=2, figsize = (10, 10))
plt.close(fig)
def updateData(i):
#if curr <=2: return
ax[0,0].clear()
ax[0,1].clear()
ax[1,0].clear()
ax[1,1].clear()
ax[0,0].set_title("Size")
ax[0,1].set_title("Aspect Ratio")
ax[1,0].set_title("Mean Density")
ax[1,1].set_title("Mean Density in Neighborhood")
ax[0,0].hist(datamatrix[i][:,0], normed = True, bins = bins0, alpha = 0.5)
ax[0,1].hist(datamatrix[i][:,1], normed = True, bins = bins1, alpha = 0.5)
ax[1,0].hist(datamatrix[i][:,2], normed = True, bins = bins2, alpha = 0.5)
ax[1,1].hist(datamatrix[i][:,3], normed = True, bins = bins3, alpha = 0.5)
simulation = FuncAnimation(fig, updateData, 60, interval=200, repeat=False)
# -
name = "Animated_Histogram.mp4"
simulation.save(name, writer = 'ffmpeg')
| 2019/ilastik/PostProcessing_Object Quantification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from src.features.build_features import get_roast_classification_dataset
from sklearn.model_selection import train_test_split, RandomizedSearchCV, learning_curve
from gensim.utils import simple_preprocess
from gensim.models.doc2vec import TaggedDocument, Doc2Vec
from sklearn.ensemble import RandomForestClassifier
import matplotlib.pyplot as plt
import numpy as np
X, y = get_roast_classification_dataset()
X_tokenized = []
for i, doc in enumerate(X):
tokens = simple_preprocess(doc)
tagged_doc = TaggedDocument(tokens, [i])
X_tokenized.append(tagged_doc)
X_tokenized[1]
# Split into test and train datasets
X_train, X_test, y_train, y_test = train_test_split(X_tokenized, y, test_size=0.2, random_state=25)
# create model
model = Doc2Vec(vector_size=50, min_count=2, epochs=40)
# build a vocabulary
model.build_vocab(X_train)
# view the vocab
model.wv.index_to_key
# vocab size
model.corpus_count
# train model on corpus
model.train(X_train, total_examples=model.corpus_count, epochs=model.epochs)
# look at document vector for example sentence
model.infer_vector(X_tokenized[1].words)
# ## Sanity Check the Model
# It looks like almost all documents are most similar to themselves, which is good.
X_train[1]
inferred_vector = model.infer_vector(X_train[1].words)
model.dv.most_similar([inferred_vector], topn=len(model.dv))
ranks = []
second_ranks = []
for idx in range(len(X_train)):
doc_id = X_train[idx].tags[0]
inferred_vector = model.infer_vector(X_train[idx].words)
sims = model.dv.most_similar([inferred_vector], topn=len(model.dv))
rank = [docid for docid, sim in sims].index(doc_id)
ranks.append(rank)
second_ranks.append(sims[1])
# +
import collections
counter = collections.Counter(ranks)
print(counter)
# -
# ## Classifier
X_train_doc2vec = []
for idx in range(len(X_train)):
inferred_vector = model.infer_vector(X_train[idx].words)
X_train_doc2vec.append(inferred_vector)
X_train_doc2vec[0].shape
# Fit model, and make predictions
rnd_clf = RandomForestClassifier(n_estimators=500, max_depth=32, max_features=50, n_jobs=-1, oob_score=True, random_state=23)
rnd_clf.fit(X_train_doc2vec, y_train)
rnd_clf.oob_score_
rnd_clf = RandomForestClassifier(n_jobs=-1, oob_score=True, random_state=21)
param_dist = {"max_depth": [5, 10, 25, 50, 100, 200],
"n_estimators": [500],
"max_features": [10, 20, 30, 40, 50],
"min_samples_split": [2, 3, 5, 7, 9],
"bootstrap": [True]}
n_iter_search = 10
random_search = RandomizedSearchCV(rnd_clf, param_distributions=param_dist,
n_iter=n_iter_search, random_state=25,
return_train_score=True)
random_search.fit(X_train_doc2vec, y_train)
# +
# Utility function to report best scores
def report(results, n_top=3):
for i in range(1, n_top + 1):
candidates = np.flatnonzero(results['rank_test_score'] == i)
for candidate in candidates:
print("Model with rank: {0}".format(i))
print("Mean validation score: {0:.3f} (std: {1:.3f})"
.format(results['mean_test_score'][candidate],
results['std_test_score'][candidate]))
print("Parameters: {0}".format(results['params'][candidate]))
print("")
report(random_search.cv_results_)
# -
# ## Bigger Doc2Vec Vector
# +
# create model
model = Doc2Vec(vector_size=100, min_count=2, epochs=40)
# build a vocabulary
model.build_vocab(X_train)
# train model on corpus
model.train(X_train, total_examples=model.corpus_count, epochs=model.epochs)
# classifier
X_train_doc2vec = []
for idx in range(len(X_train)):
inferred_vector = model.infer_vector(X_train[idx].words)
X_train_doc2vec.append(inferred_vector)
# Fit model, and make predictions
rnd_clf = RandomForestClassifier(n_estimators=500, max_depth=32, max_features=50, n_jobs=-1, oob_score=True, random_state=23)
rnd_clf.fit(X_train_doc2vec, y_train)
rnd_clf.oob_score_
# +
# create model
model = Doc2Vec(vector_size=25, min_count=2, epochs=40)
# build a vocabulary
model.build_vocab(X_train)
# train model on corpus
model.train(X_train, total_examples=model.corpus_count, epochs=model.epochs)
# classifier
X_train_doc2vec = []
for idx in range(len(X_train)):
inferred_vector = model.infer_vector(X_train[idx].words)
X_train_doc2vec.append(inferred_vector)
# Fit model, and make predictions
rnd_clf = RandomForestClassifier(n_estimators=500, max_depth=32, max_features=25, n_jobs=-1, oob_score=True, random_state=23)
rnd_clf.fit(X_train_doc2vec, y_train)
rnd_clf.oob_score_
# -
# ## Learning Curves
# +
training_set_size = [1, 5, 10, 20, 30, 40, 50, 60, 70, 80, 90, 100, 150, 200, 250, 300, 400, 500, 600, 700, 800, 900, 1000, 1200, 1300, 1400, 1600, 1800, 2000, 2500, 3000, 3500]
train_sizes, train_scores, test_scores, fit_times, _ = learning_curve(
estimator=RandomForestClassifier(n_estimators=500, max_depth=10, max_features=25, n_jobs=-1, oob_score=True, random_state=21),
X=X_train_doc2vec,
y=y_train,
train_sizes=training_set_size,
cv=5,
shuffle=True,
n_jobs=-1,
random_state=21,
return_times=True)
# -
train_scores_mean = np.mean(train_scores, axis=1)
train_scores_std = np.std(train_scores, axis=1)
test_scores_mean = np.mean(test_scores, axis=1)
test_scores_std = np.std(test_scores, axis=1)
fit_times_mean = np.mean(fit_times, axis=1)
fit_times_std = np.std(fit_times, axis=1)
# +
# Plot learning curve
_, axes = plt.subplots(1, 3, figsize=(20, 20))
if axes is None:
_, axes = plt.subplots(1, 3, figsize=(20, 20))
axes[0].set_title("Learning Curves Random Forest")
axes[0].set_ylim(*(0.20, 1.01))
axes[0].set_xlabel("Training examples")
axes[0].set_ylabel("Score")
axes[0].grid()
axes[0].fill_between(train_sizes, train_scores_mean - train_scores_std,
train_scores_mean + train_scores_std, alpha=0.1,
color="r")
axes[0].fill_between(train_sizes, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1,
color="g")
axes[0].plot(train_sizes, train_scores_mean, 'o-', color="r",
label="Training score")
axes[0].plot(train_sizes, test_scores_mean, 'o-', color="g",
label="Cross-validation score")
axes[0].legend(loc="best")
# Plot n_samples vs fit_times
axes[1].grid()
axes[1].plot(train_sizes, fit_times_mean, 'o-')
axes[1].fill_between(train_sizes, fit_times_mean - fit_times_std,
fit_times_mean + fit_times_std, alpha=0.1)
axes[1].set_xlabel("Training examples")
axes[1].set_ylabel("fit_times")
axes[1].set_title("Scalability of the model")
# Plot fit_time vs score
axes[2].grid()
axes[2].plot(fit_times_mean, test_scores_mean, 'o-')
axes[2].fill_between(fit_times_mean, test_scores_mean - test_scores_std,
test_scores_mean + test_scores_std, alpha=0.1)
axes[2].set_xlabel("fit_times")
axes[2].set_ylabel("Score")
axes[2].set_title("Performance of the model")
| notebooks/exploratory/2021-04-24 Exploratory Model Development - Doc2Vec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# + [markdown] deletable=true editable=true
# scheme:
#
# * 1) for data transfer, pick 1st sleep api (h2d) fo stream-0, current cc = 1 (concurrency),
# * 2) check whether there is overalp with stream-
# * 2) if there is overlap, finish cc=1, start from cc++ (cc=2), predit the future ending time
# * 3) during the predicted ending time, check whether there is overlap with stream-2
# * 4) if there is overalap, finish cc=2, start from cc++ (cc=3), predict the future ending time
# * 5) go to step 3) , search through all the cuda streams
# * 6) for each time range, we need to find out how many apis have overlap and which-pair have conflicts or not
# + deletable=true editable=true run_control={"marked": false}
# %load_ext autoreload
# %autoreload 2
import warnings
import pandas as pd
import numpy as np
import os
import sys # error msg, add the modules
import operator # sorting
from math import *
import matplotlib.pyplot as plt
sys.path.append('../../')
import cuda_timeline
import read_trace
import avgblk
import cke
from model_param import *
warnings.filterwarnings("ignore", category=np.VisibleDeprecationWarning)
# + [markdown] deletable=true editable=true run_control={"marked": false}
# # gpu info
# + deletable=true editable=true run_control={"marked": false}
gtx950 = DeviceInfo()
gtx950.sm_num = 6
gtx950.sharedmem_per_sm = 49152
gtx950.reg_per_sm = 65536
gtx950.maxthreads_per_sm = 2048
# + [markdown] deletable=true editable=true
# ### sm resource list and sm trace list
# + deletable=true editable=true
# init SM resources
SM_resList, SM_traceList = init_gpu(gtx950)
# + deletable=true editable=true
SM_resList[0]
# + deletable=true editable=true
SM_traceList[0]
# + [markdown] deletable=true editable=true
# # Understand the input
# + deletable=true editable=true
trace_s1 = 'trace_s1_5m.csv'
df_trace_s1 = read_trace.Trace2dataframe(trace_s1)
trace_s3 = 'trace_s3_5m.csv'
df_trace_s3 = read_trace.Trace2dataframe(trace_s3)
# + deletable=true editable=true
#df_trace_s1
# + deletable=true editable=true
cuda_timeline.plot_trace(df_trace_s1)
# + deletable=true editable=true
#cuda_timeline.plot_trace(df_trace_s2)
# + deletable=true editable=true
cuda_timeline.plot_trace(df_trace_s3)
# + deletable=true editable=true
df_3stream = read_trace.Get_timing_from_trace(df_trace_s3)
tot_runtime = read_trace.GetTotalRuntime(df_3stream)
print tot_runtime
# + [markdown] deletable=true editable=true
# # Kernel Info from the single stream
# + deletable=true editable=true
# extract kernel info from trace
# warning: currently lmted to one kernel
kernel = read_trace.GetKernelInfo(df_trace_s1, gtx950)
Dump_kernel_info(kernel)
# + [markdown] deletable=true editable=true
# # set up cuda stream kernels
# + deletable=true editable=true
# for each stream, have a dd for each kernel
stream_kernel_list = []
stream_num = 3
for sid in range(stream_num):
#print sid
# key will be the kernel order
# value will be the kernel info
kern_dd = {}
kern_dd[0] = Copy_kernel_info(kernel)
stream_kernel_list.append(kern_dd)
Dump_kernel_info(stream_kernel_list[0][0])
# + [markdown] deletable=true editable=true
# ### start kernel from beginning
# + deletable=true editable=true
df_s1_trace_timing = read_trace.Get_timing_from_trace(df_trace_s1)
df_s1 = read_trace.Reset_starting(df_s1_trace_timing)
# + deletable=true editable=true
df_s1
# + [markdown] deletable=true editable=true
# ### set the h2d start for all the cuda streams
# + deletable=true editable=true
# find when to start the stream and update the starting pos for the trace
H2D_H2D_OVLP_TH = 3.158431
df_cke_list = cke.init_trace_list(df_s1, stream_num = stream_num, h2d_ovlp_th = H2D_H2D_OVLP_TH)
# + deletable=true editable=true
#df_cke_list[0]
# + deletable=true editable=true
#df_cke_list[1]
# + deletable=true editable=true
#df_cke_list[2]
# + [markdown] deletable=true editable=true
# ### merge all the cuda stream trace together
# + deletable=true editable=true
df_all_api = cke.init_sort_api_with_extra_cols(df_cke_list)
# + deletable=true editable=true
df_all_api
# + [markdown] deletable=true editable=true
# ### start algorithm
# + deletable=true editable=true
simPos = 0.0
more_streams = stream_num - 1
active_stream_dd = {}
for s in range(stream_num):
active_stream_dd[s] = None
Dump_dd(active_stream_dd)
round_count = 1
while not cke.AllDone(df_all_api):
#
# check whether there is any call in sleep
# if not, which means all wake(or done), finish them all and break the while loop
if NoMoreSleepCalls(df_all_api):
print('no more sleep calls')
df_all_api = FinishRestWakeCalls(df_all_api)
break
#if round_count == 11: break
df_all_api, r1, r1_stream = cke.pick_base_call(df_all_api)
#
# wake it up if r1 is in sleep
if GetInfo(df_all_api, r1, 'status') == 'sleep': df_all_api = SetWake(df_all_api, r1)
#
# if r1 no in the pool add it
if active_stream_dd[r1_stream] == None: active_stream_dd[r1_stream] = r1
print('\n------------\n\n new round ({}) => row {}, stream-id {}'.format(round_count, r1, r1_stream))
Dump_dd(active_stream_dd)
print('simPos {}'.format(simPos))
#if round_count == 11: break
#
# simulation position: simPos should be ahead of r1_start for wake call
#r1_start = GetInfo(df_all_api, r1, 'start')
#if r1_start >= simPos: simPos = r1_start
#print('simulation position : {}'.format(simPos))
#if round_count == 2: break
prev_row = r1
for i in range(0, more_streams):
#if round_count == 4 and i == 0: break
df_all_api, r2, r2_stream = cke.start_next_call(df_all_api, prev_row)
print('=> pick row {}, stream-id {}'.format(r2, r2_stream))
#if round_count == 10 and i == 1: break
#
# check r2 is already running, if yes,continue to pick next
if active_stream_dd[r2_stream] == r2:
print('=> picked row {} is already running. select next'.format(r2))
prev_row = r2
#if round_count == 8 and i == 0: break
continue
#if round_count == 3 and i == 0: break
#if round_count == 2: break
# current position
curPos = GetInfo(df_all_api, r2, 'start')
print('current position : {}'.format(curPos))
Dump_dd(active_stream_dd)
#if round_count == 10 and i == 1: break
#if round == 2 and i == 0: break
if active_stream_dd[r2_stream] == None:
active_stream_dd[r2_stream] = r2
# update trace during the range
print('simPos {} curPos {}'.format(simPos, curPos))
df_all_api,SM_resList, SM_traceList = cke.update_by_range(df_all_api, simPos, curPos,
gtx950,
SM_resList, SM_traceList,
stream_kernel_list)
#
# find the kernel execution time from the sm trace table
#result_kernel_runtime_dd = avgblk.Get_KernTime(SM_traceList)
#print result_kernel_runtime_dd
Dump_dd(active_stream_dd)
#print GetInfo(df_all_api, 2, 'pred_end')
#print GetInfo(df_all_api, 6, 'pred_end')
#if round_count == 8 and i == 1: break
else:
#
# there is stream api ahead, find out which call and terminate it
row_2nd = Find_prevapi_samestream(df_all_api, r2, r2_stream)
print('end prev api call at row {}'.format(row_2nd))
#if round_count == 10 and i == 1: break
#
# end the target row, update the bytes for other call
df_all_api = cke.end_target_row(df_all_api, row_2nd, simPos, curPos)
#if round_count == 9 and i == 1: break
#
# update curPos, since r2 start has been shifted right
curPos = GetInfo(df_all_api, r2, 'start')
df_all_api = UpdateCell(df_all_api, r2, 'current_pos', curPos)
print('row {}, Updated current position : {}'.format(r2, curPos))
#if round_count == 9 and i == 1: break
#
# assume there is no ovlp between row_2nd end and r2_start
row2nd_end = GetInfo(df_all_api, row_2nd, 'end')
df_all_api = cke.move_wake_for_coming_call(df_all_api, row2nd_end, curPos)
#
# update the count for active stream pool: remove row2nd, add r2 to the pool
active_stream_dd[r2_stream] = r2
#Dump_dd(active_stream_dd)
#if round_count == 9 and i == 1: break
#
# shift right
simPos = curPos
#
# update prev_row
prev_row = r2
#
# dump active dd
#Dump_dd(active_stream_dd)
#if i == 0: break
#if round_count == 6 and i == 1: break
#
# end of For loop
# dump active dd
Dump_dd(active_stream_dd)
print('simPos {} curPos {}'.format(simPos, curPos))
#if round_count == 11: break
#
#
# check whether the active pool is full, if yes, terminate the api that ends soon
df_all_api, active_stream_dd, simPos = cke.check_activestream_and_update(df_all_api, active_stream_dd, simPos)
print active_stream_dd
#if round_count == 11: break
round_count += 1
# + deletable=true editable=true
df_all_api
# + deletable=true editable=true
#
# run above
#
| mem_mem/tests/maxCC2_3cke/cc2_3cke-v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import random
import multiprocessing
import numpy as np
from deap import creator, base, tools, algorithms
from vnpy.app.cta_strategy.backtesting import BacktestingEngine,OptimizationSetting
from vnpy.app.cta_strategy.strategies.boll_channel_strategy import BollChannelStrategy
from vnpy.app.cta_strategy.strategies.atr_rsi_strategy import AtrRsiStrategy
from datetime import datetime
import multiprocessing #多进程
from functools import lru_cache
# +
setting = OptimizationSetting()
#setting.add_parameter('atr_length', 10, 50, 2)
#setting.add_parameter('atr_ma_length', 10, 50, 2)
#setting.add_parameter('rsi_length', 4, 50, 2)
#setting.add_parameter('rsi_entry', 4, 30, 1)
setting.add_parameter('boll_window', 4, 50, 2)
#setting.add_parameter('boll_dev', 4, 50, 2)
setting.add_parameter('cci_window', 4, 50, 2)
setting.add_parameter('atr_window', 4, 50, 2)
local_setting = setting.generate_setting()
total_sample = len(local_setting)
print("数据总体:",total_sample)
# -
setting_names = random.choice(local_setting).keys()
setting_names
def parameter_generate():
setting_param = list(random.choice(local_setting).values())
return setting_param
parameter_generate()
setting=dict(zip(setting_names,parameter_generate()))
setting
# +
def object_func(strategy_avg):
""""""
return run_backtesting(tuple(strategy_avg))
#return run_backtesting(strategy_avg)
@lru_cache(maxsize=1000000)
def run_backtesting(strategy_avg):
# 创建回测引擎对象
engine = BacktestingEngine()
engine.set_parameters(
vt_symbol="IF88.CFFEX",
interval="1m",
start=datetime(2016, 1, 1),
end=datetime(2019, 1,1),
rate=0.3/10000,
slippage=0.2,
size=300,
pricetick=0.2,
capital=1_000_000,
)
setting=dict(zip(setting_names,strategy_avg))
#加载策略
#engine.initStrategy(TurtleTradingStrategy, setting)
engine.add_strategy(BollChannelStrategy, setting)
engine.load_data()
engine.run_backtesting()
engine.calculate_result()
result = engine.calculate_statistics(output=False)
return_drawdown_ratio = round(result['return_drawdown_ratio'],2) #收益回撤比
sharpe_ratio= round(result['sharpe_ratio'],2) #夏普比率
return return_drawdown_ratio , sharpe_ratio
# -
object_func(parameter_generate())
target_names = ["return_drawdown_ratio" , "sharpe_ratio"]
def show_result(hof):
for i in range(len(hof)):
solution = hof[i]
parameter=dict(zip(setting_names,solution))
result=dict(zip(target_names,list(object_func(solution))))
print({**parameter, **result})
# +
from time import time
#设置优化方向:最大化收益回撤比,最大化夏普比率
creator.create("FitnessMax", base.Fitness, weights=(1.0, 1.0)) # 1.0 求最大值;-1.0 求最小值
creator.create("Individual", list, fitness=creator.FitnessMax)
def optimize(population=None):
""""""
start = time()
toolbox = base.Toolbox()
# 初始化
toolbox.register("individual", tools.initIterate, creator.Individual,parameter_generate)
toolbox.register("population", tools.initRepeat, list, toolbox.individual)
toolbox.register("mate", tools.cxTwoPoint)
toolbox.register("mutate", tools.mutUniformInt,low = 4,up = 40,indpb=1)
toolbox.register("evaluate", object_func)
toolbox.register("select", tools.selNSGA2)
pool = multiprocessing.Pool(multiprocessing.cpu_count())
toolbox.register("map", pool.map)
#toolbox.register("map", futures.map)
#遗传算法参数设置
MU = 80 #设置每一代选择的个体数
LAMBDA = 100 #设置每一代产生的子女数
POP=100
CXPB, MUTPB, NGEN = 0.95, 0.05,30 #分别为种群内部个体的交叉概率、变异概率、产生种群代数
if population==None:
LAMBDA = POP = int(pow(total_sample, 1/2.7))
MU = int(0.8*POP)
pop = toolbox.population(POP) #设置族群里面的个体数量
hof = tools.ParetoFront() #解的集合:帕累托前沿(非占优最优集)
stats = tools.Statistics(lambda ind: ind.fitness.values)
np.set_printoptions(suppress=True) #对numpy默认输出的科学计数法转换
stats.register("mean", np.mean, axis=0) #统计目标优化函数结果的平均值
stats.register("std", np.std, axis=0) #统计目标优化函数结果的标准差
stats.register("min", np.min, axis=0) #统计目标优化函数结果的最小值
stats.register("max", np.max, axis=0) #统计目标优化函数结果的最大值
print("开始运行遗传算法,每代族群总数:%s, 优良品种筛选个数:%s,迭代次数:%s,交叉概率:%s,突变概率:%s" %(POP,MU,NGEN,CXPB,MUTPB))
#运行算法
algorithms.eaMuPlusLambda(pop, toolbox, MU, LAMBDA, CXPB, MUTPB, NGEN, stats,
halloffame=hof) #esMuPlusLambda是一种基于(μ+λ)选择策略的多目标优化分段遗传算法
end = time()
cost = int((end - start))
print("遗传算法优化完成,耗时%s秒"% (cost))
print("输出帕累托前沿解集:")
show_result(hof)
# -
optimize()
MU = 80 #设置每一代选择的个体数
POP = 100 #设置每一代产生的子女数
CXPB, MUTPB, NGEN = 0.95, 0.05,20
print("开始运行遗传算法,每代族群总数:%s, 优良品种筛选个数:%s,迭代次数:%s,交叉概率:%s,突变概率:%s" %(POP,MU,NGEN,CXPB,MUTPB))
| tests/backtesting/GA_Pre_Final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv('data/src/titanic_train.csv')
df.drop(labels=['Name', 'Ticket', 'Cabin'], axis=1, inplace=True)
df_single = df.set_index('PassengerId')
print(df_single.head())
df_multi = df.set_index(['Sex', 'Pclass', 'Embarked', 'PassengerId']).sort_index()
print(df_multi.head())
print(df_multi.tail())
print(df_multi.mean())
print(df_single.mean())
print(df_multi.max())
print(df_single.max())
print(df_multi.mean(level='Sex'))
print(df_multi.mean(level=0))
print(df_multi.mean(level=1))
print(df_multi.mean(level=2))
print(df_multi.mean(level=['Sex', 'Pclass']))
print(df_multi.mean(level=[0, 1, 2]))
print(df_single.groupby(by='Sex').mean())
print(df_single.groupby(by=['Sex', 'Pclass', 'Embarked']).mean())
print(df_multi.groupby(level='Sex').size())
print(df_multi.groupby(level=2).size())
print(df_multi.groupby(level=[0, 1, 2]).size())
print(df_single.groupby(by=['Sex', 'Pclass', 'Embarked']).size())
| notebook/pandas_multiindex_level_stats.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import multiprocessing
import os
from src.data_extract_utils import *
# -
def get_all_files(dir):
filelist = []
for root, dirs, files in os.walk(dir):
for file in files:
filelist.append(os.path.join(root,file))
return filelist
def process_file(fname, onlyME=True):
print("Worker %d is processing file %s\n" % (os.getpid(), fname))
trial_name = fname[len(database_dir):].split('/')
if onlyME and trial_name[0][-2:] == "MI":
print("Worker %d is SKIPPING file %s\n" % (os.getpid(), fname))
return
run_idx = trial_name[1].split('.')
run_idx = run_idx[0].split('_')
trial_name = trial_name[0] + "_" + run_idx[-1]
pwd = os.getcwd()
processed_file_dir = pwd + "/processed_data"
try:
if not os.path.isdir(processed_file_dir):
os.mkdir(processed_file_dir)
except OSError as error:
print("Cannot create directory. Exiting...")
print(error)
seq_v_class_fname = processed_file_dir + "/" + trial_name + ".pickle"
reject_trials_fname = processed_file_dir + "/" + trial_name + "_reject_trials.pickle"
noneeg_seqs_v_class_fname = processed_file_dir + "/" + trial_name + "_noneeg.pickle"
t1 = time.time()
HDR, data = read_data(fname)
seqs_v_class_map = segregate_data_into_classes(HDR, data)
noneeg_seqs_v_class_map = segregate_noneeg_data_into_classes(HDR, data)
rejected_trials = reject_trials_from_map(seqs_v_class_map)
rejected_trials_map = {}
for key in seqs_v_class_map.keys():
rejected_trials_map[key] = np.zeros(len(seqs_v_class_map[key]), dtype='uint8')
for l in rejected_trials:
rejected_trials_map[l[0]][l[1]] = 1
# CLM = channel_loc_map()
# seqs_v_class_map = data_1D_to_2D(seqs_v_class_map, 9, 9, CLM)
pickle_data(seqs_v_class_map, seq_v_class_fname)
pickle_data(rejected_trials_map, reject_trials_fname)
pickle_data(noneeg_seqs_v_class_map, noneeg_seqs_v_class_fname)
print("Worker %d is done processing file in %f s\n" %
(os.getpid(), time.time() - t1))
# + pycharm={"name": "#%%\n"}
database_dir = "/home/sweet/1-workdir/eeg001-2017/"
filelist = get_all_files(database_dir)
MAX_NPROCESS = multiprocessing.cpu_count()//2
print("There are a total of %d files in %s\n" % (len(filelist), database_dir))
print("**********START PROCESSING ALL FILES**********")
print("MAX NUMBER OF PROCESS = %d" % (MAX_NPROCESS))
p = multiprocessing.Pool(MAX_NPROCESS)
p.map(process_file, filelist)
| data_extraction/src/notebooks/test_data_extract.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
#hide
#|skip
! [ -e /content ] && pip install -Uqq fastai # upgrade fastai on colab
# +
#|all_slow
# -
# # Using the Hugging Face Hub to share and load models
#
# > Integration with the [Hugging Face Hub](https://huggingface.co/docs/huggingface_hub/package_reference/mixins#huggingface_hub.from_pretrained_fastai)
# ## Why share to the Hugging Face Hub
# The Hub is a central platform where anyone can share and explore models, datasets, and ML demos. It aims to build the most extensive collection of Open Source models, datasets, and demos.
#
# Sharing to the Hub could amplify the impact of a fastai `Learner` by making it available for others to download and explore.
#
# Anyone can access all the fastai models in the Hub by filtering the [huggingface.co/models](https://huggingface.co/models) webpage by the fastai library, as in the image below.
#
#
# <img src="images/hf_hub_fastai.png" alt="hf_hub_fastai" width="800" />
# The Hub has built-in [version control based on git](https://huggingface.co/docs/transformers/model_sharing#repository-features) (git-lfs, for large files), discussions, [pull requests](https://huggingface.co/blog/community-update), and [model cards](https://huggingface.co/docs/hub/model-repos#what-are-model-cards-and-why-are-they-useful) for discoverability and reproducibility. For more information on navigating the Hub, see [this introduction](https://github.com/huggingface/education-toolkit/blob/main/01_huggingface-hub-tour.md).
# ## Installation
# Install `huggingface_hub`. Additionally, the integration functions require the following packages:
# - toml,
# - fastai>=2.4,
# - fastcore>=1.3.27
#
# You can install these packages manually or specify `["fastai"]` when installing `huggingface_hub`, and your environment will be ready:
#
# ```
# pip install huggingface_hub["fastai"]
# ```
#
# To share models in the Hub, you will need to have a user. Create it on the [Hugging Face website](https://huggingface.co/join).
# ## Sharing a `Learner` to the Hub
#
# First, log in to the Hugging Face Hub. You will need to create a `write` token in your [Account Settings](http://hf.co/settings/tokens). Then there are three options to log in:
#
# 1. Type `huggingface-cli login` in your terminal and enter your token.
#
# 2. If in a python notebook, you can use `notebook_login`.
#
# ```
# from huggingface_hub import notebook_login
#
# notebook_login()
# ```
#
# 3. Use the `token` argument of the `push_to_hub_fastai` function.
#
#
# Input `push_to_hub_fastai` with the `Learner` you want to upload and the repository id for the Hub in the format of "namespace/repo_name". The namespace can be an individual account or an organization you have write access to (for example, 'fastai/stanza-de'). For more details, refer to the [Hub Client documentation](https://huggingface.co/docs/huggingface_hub/main/en/package_reference/mixins#huggingface_hub.push_to_hub_fastai).
#
# ```py
# from huggingface_hub import push_to_hub_fastai
#
# # repo_id = "YOUR_USERNAME/YOUR_LEARNER_NAME"
# repo_id = "espejelomar/identify-my-cat"
#
# push_to_hub_fastai(learner=learn, repo_id=repo_id)
# ```
#
# The `Learner` is now in the Hub in the repo named [`espejelomar/identify-my-cat`](https://huggingface.co/espejelomar/identify-my-cat). An automatic model card is created with some links and next steps. When uploading a fastai `Learner` (or any other model) to the Hub, it is helpful to edit its model card (image below) so that others better understand your work (refer to the [Hugging Face documentation](https://huggingface.co/docs/hub/model-repos#what-are-model-cards-and-why-are-they-useful)).
#
# <img src="images/hf_model_card.png" alt="hf_model_card" width="800" />
#
# `push_to_hub_fastai` has additional arguments that could be of interest; refer to the [Hub Client Documentation](https://huggingface.co/docs/huggingface_hub/main/en/package_reference/mixins#huggingface_hub.from_pretrained_fastai). The model is a [Git repository](https://huggingface.co/docs/transformers/model_sharing#repository-features) with all the advantages that this entails: version control, commits, branches, [discussions and pull requests](https://huggingface.co/blog/community-update).
#
# ## Loading a Learner from Hub
#
# Load the `Learner` we just shared in the Hub.
#
# ```py
# from huggingface_hub import from_pretrained_fastai
#
# # repo_id = "YOUR_USERNAME/YOUR_LEARNER_NAME"
# repo_id = "espejelomar/identify-my-cat"
#
# learner = from_pretrained_fastai(repo_id)
# ```
#
# The [Hub Client documentation](https://huggingface.co/docs/huggingface_hub/main/en/package_reference/mixins#huggingface_hub.from_pretrained_fastai) includes addtional details on `from_pretrained_fastai`.
#
| nbs/74_huggingface.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # VacationPy
# ----
#
# #### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import gmaps
import os
# Import API key
from config import g_key
# -
# ### Store Part I results into DataFrame
# * Load the csv exported in Part I to a DataFrame
# +
# Study data files
file_loc = "../WeatherPy/output_data/City_Data.csv"
# Read the weather data and the study results
weather_data = pd.read_csv(file_loc)
weather_data.head()
# -
# ### Humidity Heatmap
# * Configure gmaps.
# * Use the Lat and Lng as locations and Humidity as the weight.
# * Add Heatmap layer to map.
# Access maps with unique API key
gmaps.configure(api_key=g_key)
# +
# Store latitude and longitude in locations
location = weather_data[["Latitude", "Longitude"]].astype(float)
# Fill NaN values and convert to float
humidity = weather_data["Humidity"].astype(float)
# +
# Plot Heatmap
fig = gmaps.figure()
# Create heat layer
heat_layer = gmaps.heatmap_layer(location,
weights=humidity,
dissipating=False,
max_intensity=100,
point_radius=1)
# Add layer
fig.add_layer(heat_layer)
#save figure
plt.savefig("output_data/vacationheatmap.png")
fig
# -
# ### Create new DataFrame fitting weather criteria
# * Narrow down the cities to fit weather conditions.
# * Drop any rows will null values.
# +
#Narrow down the DataFrame to find your ideal weather condition.
#A max temperature lower than 80 degrees but higher than 70.
criteria_1 = weather_data.loc[(weather_data["Max Temp"]>70) & (weather_data["Max Temp"]<80),:]
# #Wind speed less than 10 mph.
criteria_2 = criteria_1.loc[criteria_1["Wind Speed"]<10,:]
# #Zero cloudiness.
criteria_3 = criteria_2.loc[criteria_2["Cloudiness"]==0,:]
criteria_3.head()
# -
criteria_3.dropna(inplace=True)
# ### Hotel Map
# * Store into variable named `hotel_df`.
# * Add a "Hotel Name" column to the DataFrame.
# * Set parameters to search for hotels with 5000 meters.
# * Hit the Google Places API for each city's coordinates.
# * Store the first Hotel result into the DataFrame.
# * Plot markers on top of the heatmap.
# +
# from pprint import pprint
# base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
# params = {
# "location": "-33.93, 18.42",
# "radius": "5000",
# "type": "lodging",
# "key": g_key,
# }
# response = requests.get(base_url, params=params).json()
# pprint(response)
# result[0]["name"]
# +
hotel_df = criteria_3.copy()
# set up additional columns to hold information
hotel_df['Hotel Name'] = ""
hotel_df.head()
# +
# find the closest restaurant of each type to coordinates
base_url = "https://maps.googleapis.com/maps/api/place/nearbysearch/json"
params = {
"radius": "5000",
"type": "lodging",
"key": g_key,
}
response = requests.get(base_url, params=params).json()
# use iterrows to iterate through pandas dataframe
for index, row in hotel_df.iterrows():
# get locations of the city
loc_lat=row["Latitude"]
loc_lon=row["Longitude"]
locat=f"{loc_lat},{loc_lon}"
cit = row["City"]
# add keyword to params dict
params['location'] = locat
# assemble url and make API request
print(f"Retrieving Results for Index {index}: {cit}.")
response = requests.get(base_url, params=params).json()
# extract results
results = response['results']
try:
hotel = results[0]["name"]
print(f"Closest hotel to {cit} at {loc_lat}, {loc_lon} is {hotel}.")
hotel_df.loc[index, 'Hotel Name'] = hotel
except (KeyError, IndexError):
print("Missing field/result... skipping.")
print("------------")
# -
hotel_df
# +
# NOTE: Do not change any of the code in this cell
# Using the template add the hotel marks to the heatmap
info_box_template = """
<dl>
<dt>Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
</dl>
"""
# Store the DataFrame Row
# NOTE: be sure to update with your DataFrame name
hotel_info = [info_box_template.format(**row) for index, row in hotel_df.iterrows()]
locations = hotel_df[["Latitude", "Longitude"]]
# +
# Add marker layer ontop of heat map
heat_layer = gmaps.heatmap_layer(location,
weights=humidity,
dissipating=False,
max_intensity=100,
point_radius = 1,
opacity=1)
marker_layer = gmaps.marker_layer(locations,
hover_text='',
label='',
info_box_content=hotel_info)
fig.add_layer(heat_layer)
fig.add_layer(marker_layer)
plt.savefig("output_data/vacation_with_hotels.png")
# Display figure
fig
# -
| VacationPy/VacationPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# name: python392jvsc74a57bd0aee8b7b246df8f9039afb4144a1f6fd8d2ca17a180786b69acc140d282b71a49
# ---
# Generate a histrogram of BFDR for a SAINT file
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from pathlib import Path
saint_file = 'data/mito.txt'
# +
def read_saint(saint_file):
columns = ['Bait', 'Prey', 'PreyGene', 'BFDR']
df = pd.read_csv(saint_file, sep='\t', usecols=columns)
return df
df = read_saint(saint_file)
# +
bins = np.arange(0, 1, 0.05)
df.plot.hist(by='BFDR', bins=bins)
plt.title('BFDR distribution')
plt.xlabel('BFDR')
plt.ylabel('Interactions')
stem = Path(saint_file).stem
outfile = f'output/{stem}.pdf'
plt.savefig(outfile)
# -
| analysis/histogram-saint/histogram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
# %matplotlib notebook
# -
import pysh.samplings as samplings
import pysh.plot as shp
from pysh.samplings import sph2cart
# +
r, theta, phi, weights = samplings.equiangular(4)
x,y,z = sph2cart(r,theta,phi)
shp.scatter(x,y,z)
# +
r, theta, phi, weights = samplings.gaussian(4)
x,y,z = sph2cart(r,theta,phi)
shp.scatter(x,y,z)
# +
r, theta, phi = samplings.eigenmike_em32()
x,y,z = sph2cart(r,theta,phi)
shp.scatter(x,y,z)
# +
r, theta, phi = samplings.icosahedron()
x,y,z = sph2cart(r,theta,phi)
shp.scatter(x,y,z)
# +
n_max = 4
r,theta,phi,w = samplings.hyperinterpolation(n_max)
x,y,z = sph2cart(r,theta,phi)
shp.scatter(x,y,z)
# +
n_max = 3
rad, theta, phi = samplings.healpix(n_max)
x,y,z = sph2cart(rad,theta,phi)
shp.scatter(x,y,z)
| tests/test_samplings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: myconda
# language: python
# name: python3
# ---
# # Sparse Representation Classification (no block validate)
#
# +
import torch
from matplotlib import pyplot as plt
import dataset
from src import src_eval
# -
name = "src_validate"
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
print(device)
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
train_ds = dataset.train()
train_ds = torch.tensor(train_ds, device=device)
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
test_ds = dataset.test()
test_ds = torch.tensor(test_ds, device=device)
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
accu = src_eval(train_ds, train_ds, k=100, device=device)
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
plt.bar(range(len(accu)), accu)
plt.title(f"{name}")
plt.savefig(f"{name}.png")
plt.show()
# + jupyter={"outputs_hidden": false} pycharm={"name": "#%%\n"}
accu.mean()
# -
accu
| src_validate.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="lC45WUtANxZ8"
# # StyleGAN3 Reactive Audio
#
# By <NAME> for the StyleGAN2 Deep Dive class.
#
# This notebook shows one basic example of how to alter your StyleGAN2 vectors with audio. There are lots of different techniques to explore in this, but this is one simple way.
#
# Big thanks to <NAME> who provided the basis for a lot of this code with [this gist](https://gist.github.com/rolux/48f1da6cf2bc6ca5833dbacbf852b348).
# + [markdown] id="m788lAb7OtJt"
# ## Installation
#
# First let’s install the repos and dependencies needed.
# + colab={"base_uri": "https://localhost:8080/"} id="U0QlWt43Lg5j" outputId="39105ad4-2546-4873-f1ae-fc163b64ba17"
# 시작하기 전에 GPU 부터 확인하시오! K80 받으면 차라리 꺼라!
# !nvidia-smi
# + colab={"base_uri": "https://localhost:8080/"} id="dSVmbuBZbRyp" outputId="79bd58c3-d71b-472b-fd73-cf863c24bfe4"
# !wget https://github.com/ninja-build/ninja/releases/download/v1.8.2/ninja-linux.zip # !wget : Web GET의 약어로 웹 상의 파일을 다운로드할 때 사용하는 명령어 -> ninja-linux.zip 다운로드
# !sudo unzip ninja-linux.zip -d /usr/local/bin/ # !sudo 현재 계정에서 root 권한을 이용하여 명령어를 실행할 때 사용 -> ninja-linux.zip 을 unzip
# !sudo update-alternatives --install /usr/bin/ninja ninja /usr/local/bin/ninja 1 --force # 위에서 설치한 특정 패키지를 설치한 후 이 코드를 통해 명령어로 등록을 할 수 있습니다. (참고링크 : https://www.whatwant.com/entry/update-alternatives-%EC%97%AC%EB%9F%AC-%EB%B2%84%EC%A0%84%EC%9D%98-%ED%8C%A8%ED%82%A4%EC%A7%80-%EA%B4%80%EB%A6%AC%ED%95%98%EA%B8%B0)
# + id="5n6wTwbFOofN" outputId="cf7bde8e-618c-4747-af56-169800af4c66" colab={"base_uri": "https://localhost:8080/"}
# !git clone https://github.com/dvschultz/stylegan3.git # use this fork to get interpolation functions
# !pip install opensimplex # needed for noise interpolation
# %cd stylegan3
# + id="KUI3IRUIkJZD"
# StyleGAN3 gen_video.py 분석
"""Generate lerp videos using pretrained network pickle."""
import copy
import os
import re
from typing import List, Optional, Tuple, Union
import click
import dnnlib
import imageio
import numpy as np
import scipy.interpolate
import torch
from tqdm import tqdm
import legacy
# + id="EW1LNTgCkiJs"
def layout_grid(img, grid_w=None, grid_h=1, float_to_uint8=True, chw_to_hwc=True, to_numpy=True):
batch_size, channels, img_h, img_w = img.shape
if grid_w is None:
grid_w = batch_size // grid_h
assert batch_size == grid_w * grid_h # 위의 식을 변환한 것
if float_to_uint8:
img = (img * 127.5 + 128).clamp(0, 255).to(torch.uint8)
img = img.reshape(grid_h, grid_w, channels, img_h, img_w)
img = img.permute(2, 0, 3, 1, 4)
img = img.reshape(channels, grid_h * img_h, grid_w * img_w)
if chw_to_hwc:
img = img.permute(1, 2, 0)
if to_numpy:
img = img.cpu().numpy()
return img
# + id="eTRDQmjpkk8V"
def gen_interp_video(G, mp4: str, seeds, shuffle_seed=None, w_frames=60*4, kind='cubic', grid_dims=(1,1), num_keyframes=None, wraps=2, psi=1, device=torch.device('cuda'), **video_kwargs):
grid_w = grid_dims[0]
grid_h = grid_dims[1]
if num_keyframes is None:
if len(seeds) % (grid_w*grid_h) != 0:
raise ValueError('Number of input seeds must be divisible by grid W*H')
num_keyframes = len(seeds) // (grid_w*grid_h) # 위 식의 값은 0이 나오게 해야합니다!
all_seeds = np.zeros(num_keyframes*grid_h*grid_w, dtype=np.int64)
for idx in range(num_keyframes*grid_h*grid_w):
all_seeds[idx] = seeds[idx % len(seeds)]
if shuffle_seed is not None:
rng = np.random.RandomState(seed=shuffle_seed)
rng.shuffle(all_seeds)
zs = torch.from_numpy(np.stack([np.random.RandomState(seed).randn(G.z_dim) for seed in all_seeds])).to(device)
ws = G.mapping(z=zs, c=None, truncation_psi=psi)
_ = G.synthesis(ws[:1]) # warm up
ws = ws.reshape(grid_h, grid_w, num_keyframes, *ws.shape[1:])
# Interpolation.
grid = []
for yi in range(grid_h):
row = []
for xi in range(grid_w):
x = np.arange(-num_keyframes * wraps, num_keyframes * (wraps + 1))
y = np.tile(ws[yi][xi].cpu().numpy(), [wraps * 2 + 1, 1, 1])
interp = scipy.interpolate.interp1d(x, y, kind=kind, axis=0)
row.append(interp)
grid.append(row)
# Render video.
video_out = imageio.get_writer(mp4, mode='I', fps=60, codec='libx264', **video_kwargs)
for frame_idx in tqdm(range(num_keyframes * w_frames)):
imgs = []
for yi in range(grid_h):
for xi in range(grid_w):
interp = grid[yi][xi]
w = torch.from_numpy(interp(frame_idx / w_frames)).to(device)
img = G.synthesis(ws=w.unsqueeze(0), noise_mode='const')[0]
imgs.append(img)
video_out.append_data(layout_grid(torch.stack(imgs), grid_w=grid_w, grid_h=grid_h))
video_out.close()
# + colab={"base_uri": "https://localhost:8080/"} id="0hQzeHb6JIT5" outputId="e065b5a0-0ec5-4834-8e47-c6e8ba003e49"
# !wget https://raw.github.com/circulosmeos/gdown.pl/master/gdown.pl # gdown 을 사용할 수 있는 함수
# !chmod u+x gdown.pl # chmod : 기존 파일 또는 디렉토리에 대한 접근 권한(파일 모드) 을 변경할 때 사용하는 명령어
# + colab={"base_uri": "https://localhost:8080/"} id="PsKJfBDni-RH" outputId="de65d2d1-49e4-4ac0-83aa-e9f6a184f01f"
# !pip install gdown==4.3
# + colab={"base_uri": "https://localhost:8080/"} id="uUw8pFx9jHBX" outputId="79fd471c-8b31-4b4e-c426-d26a24de9a43"
# awesome_beach.pkl, forest10s.wav download
# !gdown --fuzzy https://drive.google.com/file/d/1_Cneq6wuh2f8_rKES1rbuFT5wYTqpXwD/view?usp=sharing
# !gdown --fuzzy https://drive.google.com/file/d/1wHjX4oFzwbvWYsKzeC0GsVd3jrFnnpfA/view?usp=sharing
# + [markdown] id="4DWRv9BWTr4A"
# ## Upload an audio file
#
# I recommend uploading something simple to start with (think a single instrument or track with silence in it). The file should be in .mp3 or .wav format.
# + colab={"base_uri": "https://localhost:8080/"} id="hnZkq5i0-O_j" outputId="ee2d96ae-3aa8-4799-9964-2484fd5d250d"
# google.colab에 gdrive 넣는 코드
from google.colab import drive
drive.mount('/gdrive',force_remount=True)
# + id="DPIf-xU-TxdN" outputId="cdf358fe-1ea1-447b-9d13-e55f881b6a60" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY>", "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 447}
# google.colab에 파일을 업로드 하는 코드 -> 여기에서 wav 파일을 업로드 하신다고 생각하시면 됩니다.
from google.colab import files
uploaded = files.upload()
# for fn in uploaded.keys():
# print('User uploaded file "{name}" with length {length} bytes'.format(
# name=fn, length=len(uploaded[fn])))
# + [markdown] id="1JXB3HnLjP-j"
# ##Process audio
# The next step process our audio files. Edit the first line of the next cell to point to your audio file. The run the cell after that to process the audio and generate a graph of the volume data.
# + id="KW4dQcTUB1q2"
# stylegan3 모듈을 불러오는 방법
import sys # 파이썬을 설치할 때 함께 설치 되는 라이브러리 모듈
sys.path.append('/content/stylegan3') # sys.path.append를 이용해서 /content/stylegan3 라는 디렉토리를 sys.path에 추가하여 모듈을 불러와서 사용할 수 있도록 하는 코드
# + id="tMKDAPieT7XE" colab={"base_uri": "https://localhost:8080/"} outputId="222f415d-1802-4d38-f91e-73b43a837d96"
import os
import numpy as np
from scipy.interpolate import interp1d
from scipy.io import wavfile
from scipy.signal import savgol_filter # Savitzky–Golay 필터는 데이터를 평활화, 즉 신호 경향을 왜곡하지 않고 데이터의 정밀도를 높이기 위해 디지털 데이터 포인트 세트에 적용 할 수있는 디지털 필터입니다.
import matplotlib.pyplot as plt
import PIL.Image
import moviepy.editor
import dnnlib
import tensorflow as tf
import torch
import pickle
import math
import random
# + id="GbsE9p-2Vl3B"
device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')
# + id="i8TWpZNjTwWa"
# wav_filename 경로 설정 코드
wav_filename = "/content/stylegan3/forest10s.wav"
# + colab={"base_uri": "https://localhost:8080/", "height": 259} id="gopdtgoA0dfg" outputId="0977d592-1f31-409c-a7d7-6c297e1987bb"
audio = {}
fps = 24
# 파형 민감도 설정 / polyorder must be smaller than window_length
window_length = 111 # FFT (FFT란 Fast Fourier Transform의 약자이며, 쉽게 말해서 빠르게 Fourier를 변환하는 방식을 할 때 수행할 시간 간격을 의미합니다.
polyorder = 5 # 정확하게 무슨 의미인지는 모르겠지만, 7 이상의 값에서 wave에서 flat 한 영역이 보입니다.
activeness = 1/2
if not os.path.exists(wav_filename):
audio_clip = moviepy.editor.AudioFileClip(wav_filename)
audio_clip.write_audiofile(wav_filename, fps=44100, nbytes=2, codec='pcm_s16le')
track_name = os.path.basename(wav_filename)[:-4]
rate, signal = wavfile.read(wav_filename)
signal = np.mean(signal, axis=1) # to mono
signal = np.abs(signal)
# seed = signal.shape[0]
duration = signal.shape[0] / rate
frames = int(np.ceil(duration * fps))
samples_per_frame = signal.shape[0] / frames
audio[track_name] = np.zeros(frames, dtype=signal.dtype)
for frame in range(frames):
start = int(round(frame * samples_per_frame))
stop = int(round((frame + 1) * samples_per_frame))
audio[track_name][frame] = np.mean(signal[start:stop], axis=0)
audio[track_name] = savgol_filter(audio[track_name], window_length, polyorder)
audio[track_name] = audio[track_name] / max(audio[track_name])
audio[track_name] = audio[track_name] ** activeness
print("Total frames : ", frames)
for track in sorted(audio.keys()):
plt.figure(figsize=(8, 3))
plt.title(track)
plt.plot(audio[track])
plt.xlabel('Time')
plt.ylabel('Amplitude')
plt.savefig(f'../{track}.png')
# + [markdown] id="ONTlyw6ZUNQl"
# Run the next cell to define some functions we’ll need to use to generate our inference images.
# + id="KyCHTNpzPuWL"
def generate_zs_from_seeds(seeds,Gs):
zs = []
for seed_idx, seed in enumerate(seeds): # seed 인덱스, 값
rnd = np.random.RandomState(seed) # Random 생성값
z = rnd.randn(1, Gs.mapping.z_dim) # [minibatch, component] 가우시안 정규 분포 난수 생성
zs.append(z)
return zs
def convertZtoW(latent, truncation_psi=0.7, truncation_cutoff=9):
latent = torch.from_numpy(latent).to(device)
dlatent = Gs.mapping(latent, 0) # [seed, layer, component]
dlatent_avg = Gs.mapping.w_avg # [component]
for i in range(truncation_cutoff):
dlatent[0][i] = (dlatent[0][i]-dlatent_avg)*truncation_psi + dlatent_avg
return dlatent
def generate_images_in_w_space(dlatents, truncation_psi,folder='random'):
# Gs_kwargs = dnnlib.EasyDict()
# Gs_kwargs.output_transform = dict(func=convert_images_to_uint8, nchw_to_nhwc=True)
# Gs_kwargs.randomize_noise = False
# Gs_kwargs.truncation_psi = truncation_psi
dlatent_avg = Gs.mapping.w_avg # [component]
if folder == 'random':
temp_dir = 'frames%06d'%int(1000000*random.random())
else:
temp_dir = folder
os.system('mkdir %s'%temp_dir)
for row, dlatent in enumerate(dlatents):
print('Generating image for step %d/%d ...' % (row, len(dlatents)))
#row_dlatents = (dlatent[np.newaxis] - dlatent_avg) * np.reshape(truncation_psi, [-1, 1, 1]) + dlatent_avg
dl = (dlatent-dlatent_avg)*truncation_psi + dlatent_avg # dl이 왜 들어가나요?? 내부 네트워크에서 karg를 줬을 때 저렇게 수행이 됩니다. <- truncation에 대해서
row_images = Gs.synthesis(dlatent) # dlatent -> dl 로 변경 시 truncation_psi이 적용 된 상태 -> mapping 에 넣을 때 저 kwargs 값을 주면 됩니다.
row_image = (row_images.permute(0,2,3,1)*127.5+128).clamp(0,255).to(torch.uint8)
row_image = row_image.squeeze(0).cpu().numpy()
PIL.Image.fromarray(row_image, 'RGB').save('%s/frame%05d.png' % (temp_dir, row))
def load_networks(path):
with open(path, 'rb') as stream:
Gs = pickle.load(stream)['G_ema'].to(device)
Gs.eval()
return Gs
# + [markdown] id="kByt3G4yUp02"
# ## Generate Images
#
# ### Use Volume to interpolate between two seeds
# The next cell will take two seed values and do a linear interpolation of them using the volume from your audio. When the audio is silent, it will be the first seed you list. When it is at its loudest it will be the second. Everything in between will be an interpolated value.
# + id="VSPddCx9iVSR" colab={"base_uri": "https://localhost:8080/"} outputId="6b8fe827-1008-49be-8fe7-0d4af2c7470d"
# z noise blend
network_pkl = '/content/stylegan3/awesome_beach.pkl'
seeds = [10, 40, 160, 640, 2560]
seeds_t = [20, 80, 320, 1280, 5120]
truncation_value = 0.7 # 크면 클 수록 변화 되는 정도가 큰 거 같은 느낌??
truncation_psi = 0.7 # 작으면 작을 수록 파도가 잔잔해집니다.
truncation_cutoff = 5 # 아직은 value를 변경함에 따라 어떤 상관관계를 보이는 지 찾지 못 했습니다.
flow_speed = 1.5
flow_energy = 1
# seeds10~5120,value0.7,psi0.7,cutoff7,speed1.5,flow_energy1
# you probably won't need to edit anything below this
Gs = load_networks(network_pkl)
# Gs_kwargs = dnnlib.EasyDict()
# Gs_kwargs.output_transform = dict(func=convert_images_to_uint8, nchw_to_nhwc=True)
# Gs_kwargs.randomize_noise = False
# Gs_syn_kwargs = dnnlib.EasyDict()
# Gs_syn_kwargs.output_transform = dict(func=convert_images_to_uint8, nchw_to_nhwc=True)
# Gs_syn_kwargs.randomize_noise = False
# Gs_syn_kwargs.minibatch_size = 4
w_avg = Gs.mapping.w_avg
def get_ws(n, frames, seed):
filename = f'../ws_{n}_{frames}_{seed}.npy'
if not os.path.exists(filename):
src_ws = np.random.RandomState(seed).randn(n, 512)
ws = np.empty((frames, 512))
for i in range(512):
x = np.linspace(0, 3*frames, 3*len(src_ws), endpoint=False)
y = np.tile(src_ws[:, i], 3)
x_ = np.linspace(0, 3*frames, 3*frames, endpoint=False)
y_ = interp1d(x, y, kind='quadratic', fill_value='extrapolate')(x_)
ws[:, i] = y_[frames:2*frames]
np.save(filename, ws)
else:
ws = np.load(filename)
return ws
def lerp_t(ends_t):
ends_t_lerp = []
for f in range(frames):
y_list = []
for i in range(len(seeds_t)):
y = 1 - abs(1/frames*(f-i/len(seeds_t)*frames))
y_list.append(y)
y_list = np.divide(y_list, sum(y_list))
temp = y_list[i]*ends_t[i]
ends_t_lerp.append(temp)
return ends_t_lerp
def lerp(v0, v1, f, t): # 시드에 따라 움직임이 생성
# return (v0*(1.0-f)+v1*f)
return v0*(1.0-(abs(math.sin(flow_speed * ((f+1)**flow_energy) * math.pi * t / 360)))) + v1*(abs(math.sin(flow_speed * ((f+1)**flow_energy) * math.pi * t / 360)))
ends = generate_zs_from_seeds(seeds,Gs) # 가우시안 정규 분포 난수
ends_t = generate_zs_from_seeds(seeds_t,Gs) # 시간값에 따른 난수
ends_b = []
ends_lerp = lerp_t(ends)
ends_t_lerp = lerp_t(ends_t)
for f in range(frames):
ends_b.append(lerp(ends_t_lerp[f],ends_lerp[f],audio[track_name][f],f))
ends_w = []
for e in range(len(ends_b)):
ends_w.append(convertZtoW(ends_b[e],1))
vectors = ends_w
generate_images_in_w_space(vectors,truncation_value,'frames_test03')
# for e in range(len(ends)):
# ends_w.append(convertZtoW(ends[e],truncation_psi,truncation_cutoff)) # latent Z -> W 로 매핑, truncation 을 사용하지 않기 위해서 0.5 -> 1,0 으로 변경, 노이즈에 따라서 얼마나 바뀌는가?
# for e in range(len(ends_t)):
# ends_w_t.append(convertZtoW(ends_t[e],truncation_psi,truncation_cutoff))
# vectors = [] # 시드별로 프레임 보간
# vectors_blending_A = []
# vectors_blending_B = []
# for i in range((len(seeds)-1)):
# count = 0
# for f in range((frames//(len(seeds)-1))+1):
# vectors.append(lerp(ends_w[i],ends_w[i+1],ends_w_t[i],ends_w_t[i+1],audio[track_name][f],f))
# if f <= ((frames//(len(seeds)-1))+1)*0.1:
# vectors_blending_A.append(lerp(ends_w[i],ends_w[i+1],ends_w_t[i],ends_w_t[i+1],audio[track_name][f],f))
# if f >= ((frames//(len(seeds)-1))+1)*0.9:
# vectors_blending_B.append(lerp(ends_w[i],ends_w[i+1],ends_w_t[i],ends_w_t[i+1],audio[track_name][f],f))
# generate_images_in_w_space(vectors,truncation_value,'frames_test11')
# + [markdown] id="tOkh2DZpV-9W"
# ### Combine the frames into a video and add the audio track back to it
#
# There’s probably a cleaner way to do this all in moviepy but I’m being lazy.
# + id="dPClSNx_Atn-" colab={"base_uri": "https://localhost:8080/"} outputId="2c90a24a-1378-4418-800a-2b121da38971"
# !ffmpeg -r 24 -i /content/stylegan3/frames_test03/frame%05d.png -vcodec libx264 -pix_fmt yuv420p /content/sound-react-volume-test-awesome1.mp4
# + id="R7TUwqrTi4y-" colab={"base_uri": "https://localhost:8080/"} outputId="97f7d981-9e82-43e8-943f-e371c7468b76"
# output file name
mp4_filename = '/content/audio_reactive_wave_sample_test-awesome.mp4' #
# video_clip = moviepy.editor.VideoClip(render_frame, duration=duration)
video_clip = moviepy.editor.VideoFileClip('/content/sound-react-volume-test-awesome1.mp4')
audio_clip_i = moviepy.editor.AudioFileClip('/content/stylegan3/forest10s.wav')
video_clip = video_clip.set_audio(audio_clip_i)
video_clip.write_videofile(mp4_filename, fps=fps, codec='libx264', audio_codec='aac', bitrate='8M')
# + [markdown] id="ibDI8hGWWPD0"
# ### Use Volume to control truncation
# In this example, we’ll use almost the same technique but use volume to change the truncation value.
#
# It’s helpful that both interpolation and truncation are essentially 0.0 to 1.0. This matches the volume signal’s output, but what if we wanted to alter it?
# + id="ifbLsbOcXsgy"
seeds=[135]
seed_z = generate_zs_from_seeds(seeds,Gs)
#Gs_kwargs = dnnlib.EasyDict()
#Gs_kwargs.output_transform = dict(func=convert_images_to_uint8, nchw_to_nhwc=True)
#Gs_kwargs.randomize_noise = False
rnd = np.random.RandomState(seeds[0])
#temp_dir = '%s-trunc_frames%06d'%(track_name,int(1000000*random.random()))
temp_dir = 's-trunc_frames2'
os.system('mkdir %s'%temp_dir)
for f in range(frames):
print('Rendering frame %d/%d ...' % (f,frames))
Gs_kwargs.truncation_psi = audio[track_name][f]
#set_vars({var: rnd.randn(*var.shape.as_list()) for var in noise_vars}) # [height, width]
images = Gs(torch.from_numpy(seed_z[0]).to(device), 0) # [minibatch, height, width, channel]
image = (images.permute(0,2,3,1)*127.5+128).clamp(0,255).to(torch.uint8)
image = image.squeeze(0).cpu().numpy()
PIL.Image.fromarray(image, 'RGB').save('%s/frame%05d.png' % (temp_dir,f))
# + id="gw3negi0e7ll"
# !ffmpeg -r 24 -i /content/s-trunc_frames2/frame%05d.png -vcodec libx264 -pix_fmt yuv420p /content/sound-truncation-volume2.mp4
# + id="g1AC2mF1bwKP"
mp4_filename = '/content/audio_reactive_truncation_wave2.mp4'
# video_clip = moviepy.editor.VideoClip(render_frame, duration=duration)
video_clip = moviepy.editor.VideoFileClip('/content/sound-truncation-volume2.mp4')
audio_clip_i = moviepy.editor.AudioFileClip('/content/ocean-waves-1.wav')
video_clip = video_clip.set_audio(audio_clip_i)
video_clip.write_videofile(mp4_filename, fps=fps, codec='libx264', audio_codec='aac', bitrate='8M')
# + id="nR6VU8QJb4cB"
seeds=[10]
seed_z = generate_zs_from_seeds(seeds,Gs)
Gs_kwargs = dnnlib.EasyDict()
Gs_kwargs.output_transform = dict(func=convert_images_to_uint8, nchw_to_nhwc=True)
Gs_kwargs.randomize_noise = False
rnd = np.random.RandomState(seeds[0])
temp_dir = '%s-trunc_frames%06d'%(track_name,int(1000000*random.random()))
os.system('mkdir %s'%temp_dir)
for f in range(frames):
print('Rendering frame %d/%d ...' % (f,frames))
#edit the next line to alter the volume signal
# new_truncation_value = audio[track_name][f]*2 #multiply by 2 (0.0 to 2.0 for volume signal/truncation value now)
new_truncation_value = (audio[track_name][f]-0.5)*2 #(-1.0 to 1.0 for volume signal/truncation value now)
Gs_kwargs.truncation_psi = new_truncation_value
set_vars({var: rnd.randn(*var.shape.as_list()) for var in noise_vars}) # [height, width]
images = Gs.run(seed_z[0], None, **Gs_kwargs) # [minibatch, height, width, channel]
PIL.Image.fromarray(images[0], 'RGB').save('%s/frame%05d.png' % (temp_dir,f))
# + id="SRdc-crzdkDm"
# !ffmpeg -r 60 -i /content/stylegan2/pleasureisallmine_01-stereo-trunc_frames623374/frame%05d.png -vcodec libx264 -pix_fmt yuv420p /content/sound-truncation-volume.mp4
# + id="hgezjUH3flxa"
mp4_filename = '../volume-trunc-test-v3.mp4'
# video_clip = moviepy.editor.VideoClip(render_frame, duration=duration)
video_clip = moviepy.editor.VideoFileClip('/content/sound-truncation-volume.mp4')
audio_clip_i = moviepy.editor.AudioFileClip('/content/stylegan2/pleasureisallmine_01-stereo.wav')
video_clip = video_clip.set_audio(audio_clip_i)
video_clip.write_videofile(mp4_filename, fps=fps, codec='libx264', audio_codec='aac', bitrate='8M')
# + [markdown] id="N_78NFCdqL68"
# ## Using feature vectors
#
# Let’s look at an example using a feature vector. In this case we’ll just use the straight audio signal.
#
# Upload your feature vector to Colab and then reference it’s location with a variable.
# + id="Kl5w7TaLxldW"
network_pkl = "/content/ffhq.pkl"
_G, _D, Gs = pretrained_networks.load_networks(network_pkl)
Gs_kwargs = dnnlib.EasyDict()
Gs_kwargs.output_transform = dict(func=convert_images_to_uint8, nchw_to_nhwc=True)
Gs_kwargs.randomize_noise = False
Gs_syn_kwargs = dnnlib.EasyDict()
Gs_syn_kwargs.output_transform = dict(func=convert_images_to_uint8, nchw_to_nhwc=True)
Gs_syn_kwargs.randomize_noise = False
Gs_syn_kwargs.minibatch_size = 4
noise_vars = [var for name, var in Gs.components.synthesis.vars.items() if name.startswith('noise')]
w_avg = Gs.get_var('dlatent_avg')
def generate_mov(seed, truncation, direction_vec, scale, n_frames, out_name = 'out', noise_spec = None, loop=True):
"""Generates a mov moving back and forth along the chosen direction vector"""
# Example of reading a generated set of images, and storing as MP4.
# %mkdir out
movieName = f'out/{out_name}.mp4'
offset = -10
step = 20 / n_frames
imgs = []
for i in range(n_frames):
print(f'{i} / {n_frames}')
batch_size = 1
all_seeds = [seed] * batch_size
all_z = np.stack([np.random.RandomState(seed).randn(*Gs.input_shape[1:]) for seed in all_seeds]) # [minibatch, component]
all_w = Gs.components.mapping.run(all_z, None) # [minibatch, layer, component]
if truncation != 1:
w_avg = Gs.get_var('dlatent_avg')
all_w = w_avg + (all_w - w_avg) * truncation # [minibatch, layer, component]
all_w += direction_vec * offset * scale
all_images = Gs.components.synthesis.run(all_w, **Gs_syn_kwargs)
#save image and display
final_im = PIL.Image.fromarray(np.median(all_images, axis=0).astype(np.uint8))
imgs.append(final_im)
#increase offset
offset += step
if loop:
imgs += imgs[::-1]
with imageio.get_writer(movieName, mode='I') as writer:
for image in log_progress(list(imgs), name = "Creating animation"):
writer.append_data(np.array(image))
# + id="PA40ehfqy2S2"
seed = 10 # starting seed (will appear at 0.5)
truncation = 0.7
feature = '/content/profile-c2.npy'
feature_range = 2 # feature_range maps the range of change in features
scale = 1 # scale multiples the strength of the feature (1 is prob fine)
#-------------------
Gs_kwargs = dnnlib.EasyDict()
Gs_kwargs.output_transform = dict(func=convert_images_to_uint8, nchw_to_nhwc=True)
Gs_kwargs.randomize_noise = False
if truncation is not None:
Gs_kwargs.truncation_psi = truncation
set_vars({var: rnd.randn(*var.shape.as_list()) for var in noise_vars}) # [height, width]
w_avg = Gs.get_var('dlatent_avg')
# get starting z and w
rnd = np.random.RandomState(seed)
z = rnd.randn(1, *Gs.input_shape[1:])
w = Gs.components.mapping.run(z, None)
# make dir
feature_name=os.path.basename(feature)[:-4]
dir = '%s_%s_range%02d_seed%05d'%(track_name,feature_name,feature_range,seed)
os.system('mkdir %s'%dir)
# setup
feature_vec = np.load(feature)
min_range = -feature_range
max_range = feature_range
offset = min_range #start value
#generate frames
for f in range(frames):
print('Rendering frame %d/%d ...' % (f,frames))
if truncation != 1:
w = w_avg + (w - w_avg) * truncation # [minibatch, layer, component]
w += feature_vec * offset * scale
#save image and display
image = Gs.components.synthesis.run(w, **Gs_syn_kwargs)
PIL.Image.fromarray(image[0],'RGB').save('%s/frame%05d.png' % (dir,f))
#increase offset
offset = lerp( min_range,max_range,audio[track_name][f] )
# + id="Hbk-mwtQ2oWj"
# !ffmpeg -r 60 -i /content/stylegan2/pleasureisallmine_01-stereo_profile-c2_range02_seed00010/frame%05d.png -vcodec libx264 -pix_fmt yuv420p /content/sound-feature-volume-range2.mp4
# + id="JnoVg2o0AE3_"
mp4_filename = '../volume-feature-test-range2.mp4'
# video_clip = moviepy.editor.VideoClip(render_frame, duration=duration)
video_clip = moviepy.editor.VideoFileClip('/content/sound-feature-volume-range2.mp4')
audio_clip_i = moviepy.editor.AudioFileClip('/content/stylegan2/pleasureisallmine_01-stereo.wav')
video_clip = video_clip.set_audio(audio_clip_i)
video_clip.write_videofile(mp4_filename, fps=fps, codec='libx264', audio_codec='aac', bitrate='8M')
# + id="ZMyuqpcDBqQ3"
| Release/HojinLee/20220218_StyleGAN3_Reactive_Audio_Ho.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Salvar un DataFrame en Google Sheets
# > "ฅ^•ﻌ•^ฅ Para poder comunicarlo al público en general"
#
# - toc: false
# - branch: master
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [pandas]
# Este es uno de los mejores trucos que aprendí trabajando en [Mutt Data](https://muttdata.ai). Del lado del desarrollo, en el día a día, trabajar con Pandas es de lo más común, tanto para análisis exploratorio de datos como manipulación de datos en general. Cuando existen otras partes interesadas en un proyecto, sobre todo personas de negocio, la comunicación es esencial y en Mutt nos iba bastante bien sacando a la luz `DataFrame`s por medio de Google Sheets — ambos están hechos para trabajar con datos tabulares y lo bueno de los documentos de Google es que son fáciles de compartir y que las personas de negocio están acostumbradas a interactuar con hojas de cálculo.
# El paquete que vamos a usar se llama [gspread](https://gspread.readthedocs.io) y es una API en Python para Google Sheets.
#
# pip install gspread
# ## Autenticación
#
# Vamos a necesitar una cuenta de servicio —que es un archivo con credenciales— para habilitar a nuestro programa a escribir en Google Sheets. Estas son las instrucciones.
#
# 1. Ir a la [consola](https://console.cloud.google.com) de Google Cloud Platform (GCP) y crear un proyecto nuevo o seleccionar uno existente (yo creé `mi-proyecto`). Si nunca usaste GCP, vas a tener algunos pasos adicionales.
# 2. En la barra de *Buscar productos y recursos* buscar **Google Drive API** y habilitarla.
# 3. En la barra de *Buscar productos y recursos* buscar **Google Sheets API** y habilitarla.
# 4. En la barra de *Buscar productos y recursos* buscar **cuentas de servicio**, en esa página:
# 5. **+ Crear cuenta de servicio**, completar el formulario; con el nombre de la cuenta de servicio es suficiente (yo elegí `google-sheets`).
# 6. Una vez creada, seleccionarla para entrar en los detalles de la cuenta.
# 7. `Claves > Agregar clave > Crear clave nueva > JSON > Crear`. Aceptar la descarga de la cuenta de servicio. En mi caso, de `mi-proyecto-80a030363d28.json`.
# 8. Mover el archivo a la carpeta de trabajo. Debe estar en lugar seguro.
#
# La cuenta de servicio servirá para todas las planillas de cálculo que necesitemos dentro de un mismo proyecto de GCP.
# ## Acceso a la hoja de cálculo
#
# Este paso es mucho muy importante, **debe realizarse cada vez que utilicemos una hoja de cálculo nueva**.
#
# Ir a la hoja de cálculo y compartirla con el correo electrónico de la cuenta de servicio (es el que figura en el detalle de la cuenta) de la misma manera que haríamos para compartírsela a otra persona mediante su cuenta de correo.
#
# En mi caso, tengo que compartir las hojas con `<EMAIL>`.
#
# ## Pandas
#
# Esta es la función que utilizo para escribir un `DataFrame` en Google Sheets. Hay algunas conversiones de tipos de datos, ya que Pandas y Google Sheets no manejan los mismos tipos.
# +
import gspread
GSHEETS_CREDENTIALS = 'mi-proyecto-80a030363d28.json'
def save_to_gsheets(df, sheet_name, worksheet_name='Sheet1'):
client = gspread.service_account(GSHEETS_CREDENTIALS)
sheet = client.open(sheet_name)
worksheet = sheet.worksheet(worksheet_name)
# convertimos el tipo de las columnas que sean datetime a string
for column in df.columns[df.dtypes == 'datetime64[ns]']:
df[column] = df[column].astype(str)
# reemplazamos valores NaN por strings vacíos
worksheet.update([df.columns.values.tolist()] + df.fillna('').values.tolist())
print(f'DataFrame escrito en la hoja {sheet_name} / {worksheet_name}.')
# -
# Vamos con un ejemplo:
# +
import numpy as np
import pandas as pd
df = pd.DataFrame(np.random.randint(0, 100, size=(7, 4)), columns=list('ABCD'))
df
# -
# Previamente tuve que crear la hoja `Ejemplo Pandas` y darle acceso a la cuenta de servicio.
save_to_gsheets(df, 'Ejemplo Pandas', worksheet_name='Sheet1')
# Este es el resultado 📝.
#
# 
| _notebooks/2020-10-13-Salvar-un-DataFrame-en-Google-Sheets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# **Data preparation** is one of the essential processes in machine learning projects workflow: with well-prepared input even simple algorithm can achieve great result, and without it --- it’s hard to get something meaningful even using the most sophisticated models (remember concept of "[garbage in — garbage out](https://en.wikipedia.org/wiki/Garbage_in,_garbage_out)").
#
# Usually, specific preparation of data for ML modeling can be considered as part of [ETL](https://en.wikipedia.org/wiki/Extract,_transform,_load) process and consists of following steps:
#
# * **feature engineering**: transformation of raw data into proper features, that can be useful for modeling; sometimes, when original data is complex enough (e. g. text, images) this process is also called *feature extraction, feature preparation*.
# * **feature selection**: removing unnecessary features (usually it can help to improve model quality/performance/etc).
#
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler, MinMaxScaler, OrdinalEncoder, OneHotEncoder
from sklearn.decomposition import PCA
from sklearn.feature_selection import VarianceThreshold, SelectFromModel, RFECV, SequentialFeatureSelector
from sklearn.ensemble import RandomForestClassifier, RandomForestRegressor
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import KFold, cross_val_score, train_test_split
from sklearn.pipeline import Pipeline, make_pipeline
from sklearn.metrics import accuracy_score
from sklearn.datasets import make_classification, load_wine, load_breast_cancer, load_diabetes, load_digits
# -
plt.style.use('seaborn-darkgrid')
# +
def plot_scatter(x, y, auto_scaled=True, title=None, clusters=None):
plt.figure(figsize=(4, 4))
plt.scatter(x, y, c=clusters, cmap='bwr')
if not auto_scaled:
plt.axis('square')
plt.grid(True)
plt.title(title)
plt.show()
def return_X_y(data, target_column):
return data.drop(target_column, axis=1), data[target_column]
# -
# # Feature Engineering
# ## Missing Values Preprocessing
housing_data = pd.read_csv('Melbourne_housing_FULL.csv')
# prepare dataset for price regression
housing_data = housing_data[~housing_data['Price'].isnull()]
# Missing values are one of the most common problems you can encounter when you try to prepare your data for machine learning. The reason for the missing values might be human errors, interruptions in the data flow, privacy concerns, and so on. Whatever is the reason, missing values affect the performance of the machine learning models (most of the algorithms do not even accept datasets with missing values).
# First let's check the amount of missing value in each column of our dataset:
housing_data.isnull().mean() # housing_data.isnull().sum() to get absolute numbers
# The most simple strategy is to drop entire rows and/or columns containing missing values based on some threshold (for example, if column contains more than *30%* --- drop it, then drop all rows that still contains some NaN's).
threshold = 0.3
housing_data_dropped = housing_data[housing_data.columns[housing_data.isnull().mean() < threshold]]
housing_data_dropped = housing_data_dropped.dropna(axis=0, how='any') # params is optinal here (matching defaults)
print(f'Original dataset shape (rows, cols): {housing_data.shape}')
print(f'Dataset shape (rows, cols) after dropna: {housing_data_dropped.shape}')
# In general dropping data without additional investigation is not a good approach in most cases since you lose a lot of potentially useful information. For this particular dataset we've fully dropped `Landsize`, `BuildingArea` columns (which actually seem like strong features from common sense).
#
# Usually a better strategy is to impute the missing values, i.e., to infer them from the known part of the data. However, there is an important selection of what you impute to the missing values. You can use default value of missing values in the column. For example, if you have a column that only has `1` and `N\A`, then it is likely that the `N\A` rows may be considered as `0`.
# Another way is to use basic statistics (like *mean* and *medians* of the columns) for imputation.
# +
# const imputing
housing_data_const = housing_data.fillna(value=0)
# mean imputing
housing_data_mean = housing_data.fillna(housing_data.mean(numeric_only=True))
# -
# There are also some advanced technics [KNN Imputation](https://scikit-learn.org/stable/modules/impute.html#nearest-neighbors-imputation), [Multivariate imputation](https://scikit-learn.org/stable/modules/impute.html#multivariate-feature-imputation).
#
# But commonly the most beneficial way is to dig deeper in available data, understand root cases of the problem and develop mixed strategy (for separate features based on investigation results). **Subject matter expertise rules!**
#
# For example, one of the questions you may ask yourself to help figure this out is this:
#
# `Is this value missing because it wasn't recorded or because it doesn’t exist?`
#
# If the value is missing because it doesn’t exist (like the height of the oldest child of someone who doesn't have any children) then it doesn't make sense to try and guess what it might be. These values you probably do want to mark this value using some special tag (or create separate bool feature). On the other hand, if a value is missing because it wasn't recorded, then you may probably use some of the imputation technics mentioned above or even more sophisticated ones.
#
# ## Feature scaling
wine_sklearn = load_wine(as_frame=True)
wine_data, wine_labels = wine_sklearn['data'], wine_sklearn['target']
wine_data
# In real world datasets you can often see multiple features spanning varying degrees of magnitude, range, and units. This is a significant obstacle as a lot of machine learning algorithms are highly sensitive to such things.
#
# To make it simple: algorithm just sees number and does not know what that number represents --- if there is a vast difference in the range say few ranging in thousands and few ranging in dozens, it makes the underlying assumption that higher ranging numbers have superiority of some sort. So, these more significant number starts playing a more decisive role while training the model.
#
# For example, you might be looking at the prices of some products in both Yen and US Dollars. One US Dollar is worth about 100 Yen, but if you don't scale your prices methods like SVM or KNN will consider a difference in price of 1 Yen as important as a difference of 1 US Dollar! This clearly doesn't fit with our intuitions of the world. With currency, you can convert between currencies. But what about if you're looking at something like height and weight? It's not entirely clear how many pounds should equal one inch (or how many kilograms should equal one meter).
#
# By scaling your variables, you can help compare different variables on equal footing (scale).
# ### Standartization
# **Standardization** of datasets is a common requirement for many machine learning models. The idea is to transform the data to the center it by removing the mean value of each feature, then scale it by dividing non-constant features by their standard deviation.
#
# $$scaled\_X = \frac{X - mean(X)}{std(X)}$$, where $X$ is **feature column** (not dataset itself!)
#
# A common approach is to use `StandardScaler` from `sklearn`:
#
scaler = StandardScaler()
wine_data_scaled = scaler.fit_transform(wine_data)
wine_data_scaled
# Let's illustrate the influence of scaling on [PCA](https://en.wikipedia.org/wiki/Principal_component_analysis):
# +
pca = PCA(n_components=2)
wine_data_pca = pca.fit_transform(wine_data)
wine_data_scaled_pca = pca.fit_transform(wine_data_scaled)
fig, (ax1, ax2) = plt.subplots(ncols=2, figsize=(18, 10))
for l, c, m in zip(range(0, 3), ('blue', 'red', 'green'), ('^', 's', 'o')):
ax1.scatter(wine_data_pca[wine_labels == l, 0], wine_data_pca[wine_labels == l, 1],
color=c, label=f'class {l}', alpha=0.5, marker=m)
for l, c, m in zip(range(0, 3), ('blue', 'red', 'green'), ('^', 's', 'o')):
ax2.scatter(wine_data_scaled_pca[wine_labels == l, 0], wine_data_scaled_pca[wine_labels == l, 1],
color=c, label=f'class {l}', alpha=0.5, marker=m)
ax1.set_title('Dataset after PCA')
ax2.set_title('Standardized dataset after PCA')
for ax in (ax1, ax2):
ax.set_xlabel('1st principal component')
ax.set_ylabel('2nd principal component')
ax.legend(loc='upper right')
# -
# ### Normalization
# An alternative standardization is scaling features to lie between a given minimum and maximum value, often between zero and one, or so that the maximum absolute value of each feature is scaled to unit size (also known as **Normalization**. This can be achieved using `MinMaxScaler` or `MaxAbsScaler` from `sklearn`, respectively.
#
# The motivation to use this scaling include robustness to very small standard deviations of features and preserving zero entries in sparse data.
# $$normalised\_X = \frac{X - min(X)}{max(X) - min(X)}$$, where $X$ is **feature column** (not dataset itself!)
# +
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler()
scaler.fit_transform(wine_data)
# -
# ## Log/Power Transform
# Log transformation is a data transformation method in which it replaces each variable $x$ with a $log(x)$. The choice of the logarithm base is usually left up to the analyst and it would depend on the purposes of statistical modeling.
#
# When our original continuous data do not follow the bell curve, we can log transform this data to make it as “normal” as possible so that the statistical analysis results from this data become more valid. In other words, the log transformation reduces or removes the skewness of our original data. The important caveat here is that the original data has to approximately follow a *log-normal distribution*. Otherwise, you can't expect any guarantees that result distribution will be close to normal (but even in such cases log transform can help to improve you scores).
mu, sigma = 5, 1
lognorm_data = np.random.lognormal(mu, sigma, 1000)
plt.figure(figsize=(16,8))
sns.histplot(lognorm_data, stat='probability')
plt.show()
plt.figure(figsize=(16,8))
sns.histplot(np.log(lognorm_data), stat='probability')
plt.show()
# This may sound a bit odd: is it even possible to meet something specific like "log-normal distribution" in real life?
#
# Well, let's plot the price column from Melbourne housing dataset, that we used previously:
plt.figure(figsize=(16,8))
sns.histplot(housing_data['Price'], stat='probability')
plt.show()
# Seems familiar!
#
# Eventually, lognormal distribution of some value in real world is quite common (just like normal distribution). It is suitable for describing length of comments, posted in the internet; the salaries amount; the population of cities and many other things. You may find some [more](https://en.wikipedia.org/wiki/Log-normal_distribution#Occurrence_and_applications) examples just on the wikipedia page.
#
# However, to get some profit from this transformation, the distribution does not necessarily have to be *exactly* lognormal; you can try to apply it to any distribution with a heavy right tail. Furthermore, one can try to use other similar transformations, formulating their own hypotheses on how to approximate the available distribution to a normal. Examples of such transformations are Box-Cox transformation (log is a special case of the Box-Cox transformation) or Yeo-Johnson transformation (extends the range of applicability to negative numbers). Some information about these transformations and their implementations in `sklean` can be found [here](https://scikit-learn.org/stable/modules/preprocessing.html#non-linear-transformation).
# ## Categorical Features Encoding
# Quite often features are not given as continuous values but categorical. For example a person could have features `["male", "female"], ["from Europe", "from US", "from Asia"], ["uses Firefox", "uses Chrome", "uses Safari", "uses Internet Explorer"]`. Such features can be efficiently coded as integers, for instance `["male", "from US", "uses Internet Explorer"]` could be expressed as `[0, 1, 3]` while `["female", "from Asia", "uses Chrome"]` would be `[1, 2, 1]`.
#
# To convert categorical features to such integer codes, we can use the *ordinal encoding*. It transforms each categorical feature to a range of integers (0 to number of categories - 1).
X = [['male', 'US', 'Safari'], ['female', 'Europe', 'Firefox'], ['male', 'Europe', 'Opera']]
pd.DataFrame(X, columns=['gender', 'place', 'browser'])
encoder = OrdinalEncoder()
ordinal_encoded_X = encoder.fit_transform(X)
# Such integer representation can, however, can be unsuitable, for a lot of models: these expect continuous input, and would interpret the categories as being ordered, which is often not desired.
#
# Another possibility to convert categorical features to features that can be used with scikit-learn estimators is to use *one-hot* encoding. The idea is to transforms each categorical feature, that has $n$ different possible categories, into $n$ separate binary features (whether the object belongs to specific category or not).
encoder = OneHotEncoder()
ohe_encoded_X = encoder.fit_transform(X).toarray()
pd.DataFrame(ohe_encoded_X, columns=encoder.get_feature_names())
# It is also possible to encode each column into $n - 1$ columns instead of $n$ columns by using the drop parameter (also called *dummy encoding*). This is useful to avoid co-linearity in the input matrix in some classifiers. Such functionality is useful, for example, when using non-regularized regression, since co-linearity would cause the covariance matrix to be non-invertible.
# You can read about some advanced technics [here](https://www.analyticsvidhya.com/blog/2020/08/types-of-categorical-data-encoding). However, most of them equivalent to one hot encoding to some degree.
#
# # Feature Selection
# Why is it sometimes necessary to select just subset of features and not all at once? The idea of removing features may seem a little counterintuitive, but there is some import motivation here:
#
# 1) First is more connected to engeneering side: the more data, the higher the computational complexity. Removing some unimportant and noisy features can help a lot here.
# 2) The second reason is related to algorithms side: some models can be unstable when data have highly correlated features ([multicolinearity](https://datascience.stackexchange.com/questions/24452/in-supervised-learning-why-is-it-bad-to-have-correlated-features)), some --- when data is noisy.
cancer_sklearn = load_breast_cancer(as_frame=True)
cancer_data, cancer_labels = cancer_sklearn['data'], cancer_sklearn['target']
cancer_data_scaled = StandardScaler().fit_transform(cancer_data)
cancer_data
# ## Statistical Approaches
# The most obvious candidate for removal is a feature whose value remains unchanged, i.e., it contains no information at all. If we build on this thought, it is reasonable to say that features with low variance are worse than those with high variance. So, one can consider cutting features with variance below a certain threshold.
X_generated, y_generated = make_classification(n_samples=1000, n_features=25, n_informative=3,
n_redundant=2, n_repeated=0)
X_generated.shape
print(VarianceThreshold(0.9).fit_transform(X_generated).shape)
print(VarianceThreshold(1).fit_transform(X_generated).shape)
print(VarianceThreshold(1.1).fit_transform(X_generated).shape)
# Keep in mind that we are using absolute value as threshold, so in real world scenario it is necessary to bring all the features to same scale (perform scaling before thresholding).
#
# Personally, I won't recommend using `VarianceTreshold` unless you are completely sure that it's needed and won't make things worse: the low variance does not necessarily mean that feature is not informative. You can also try [other](https://scikit-learn.org/stable/modules/feature_selection.html#univariate-feature-selection) a little bit more advanced statistical approaches.
# ## Selection From Modeling
#
# Basically, the idea is to use some model as an feature importance estimator: for example, we can use linear model with `Lasso` regularization (and feature weights from it) or some tree based models (which have natural ability to compute feature importance). Then, based on received importance/weights we can choose some threshold and take features, that have importance above this value.
selection_model = RandomForestClassifier(random_state=42)
selector = SelectFromModel(selection_model).fit(cancer_data, cancer_labels)
cancer_data_pruned = selector.transform(cancer_data)
print(cancer_data.columns[selector.get_support()])
print(f'Original shape: {cancer_data.shape}')
print(f'Shape after selection: {cancer_data_pruned.shape}')
# +
main_model = LogisticRegression(solver='liblinear', penalty='l1')
pipe_baseline = make_pipeline(StandardScaler(), main_model)
pipe_selection = make_pipeline(StandardScaler(), SelectFromModel(selection_model), main_model) # fix to select only once
print('Result on original data: {:f}'.format(cross_val_score(pipe_baseline, cancer_data, cancer_labels,
scoring='accuracy', cv=5).mean()))
print('Result after selection {:f}'.format(cross_val_score(pipe_selection, cancer_data, cancer_labels,
scoring='accuracy', cv=5).mean()))
# -
# We were able to reduce the number of features significantly, but, as you can see, stable performance is not guaranteed.
# It's also possible to use same model as an importance estimator and actual classifier (regressor).
# As a development of this approach we can consider recursive feature elimination: first, the model is trained on the initial set of features and the importance of each feature is obtained. Then, the least important features are pruned from current set of features. That procedure is recursively repeated on the pruned set until the desired number of features to select is eventually reached.
# +
min_features_to_select = 1
rfecv = RFECV(estimator=main_model, step=1, cv=KFold(3),
scoring='accuracy', min_features_to_select=min_features_to_select)
rfecv.fit(cancer_data_scaled, cancer_labels)
print("Optimal number of features : %d" % rfecv.n_features_)
# -
plt.figure(figsize=(16,8))
plt.plot(range(min_features_to_select,
len(rfecv.grid_scores_) + min_features_to_select),
rfecv.grid_scores_)
plt.show()
# ## Greedy (Sequential) Feature Selection
# Finally, we get to the most reliable method --- trivial brute force: just test all possible subsets of features (train a model on a subset of features, store results, repeat for different subsets, and compare the quality of models to identify the best feature set). This approach is called [Exhaustive Feature Selection](http://rasbt.github.io/mlxtend/user_guide/feature_selection/ExhaustiveFeatureSelector).
#
# However, usually this method is too computationally complex to use for some real word dataset (it's even not available in scikit-learn). To reduce complexity one can the following *greedy* heuristic: tart with zero feature and find the one feature that maximizes a cross-validated score when the model is trained on this single feature. Once that first feature is selected, we repeat the procedure by adding a new feature to the set of selected features. It is possible to iterate until we hit (preselected) maximum number of features or until the quality of the model ceases to increase significantly between iterations.
#
# This algorithm can work in the opposite direction: instead of starting with no feature and greedily adding features, we start with all the features and greedily remove features from the set.
# +
selector = SequentialFeatureSelector(main_model, scoring='accuracy', n_jobs=-1).fit(cancer_data_scaled, cancer_labels)
cancer_data_scaled_pruned = selector.transform(cancer_data_scaled)
print(cancer_data.columns[selector.get_support()])
print(f'Original shape: {cancer_data.shape}')
print(f'Shape after selection: {cancer_data_pruned.shape}\n')
print('Result on original data: {:f}'.format(cross_val_score(main_model, cancer_data_scaled,
cancer_labels, scoring='accuracy', cv=5).mean()))
print('Result after selection {:f}'.format(cross_val_score(main_model, cancer_data_scaled_pruned,
cancer_labels, scoring='accuracy', cv=5).mean()))
# -
# ## Boruta (optional)
#
# Boruta is an algorithm for finding a subset of relevant features. The paper defines a variable as being relevant if there is a subset of attributes in the dataset among which the variable is not redundant when used for prediction.
#
# Easy to read article explaining both algoritm itself and python package, can be found here: https://towardsdatascience.com/boruta-explained-the-way-i-wish-someone-explained-it-to-me-4489d70e154a
#
# A paper can be found here: https://www.researchgate.net/publication/220443685_Boruta_-_A_System_for_Feature_Selection
# # Materials & References
# 1. General article about feature engineering and selection (main reference):
# https://github.com/Yorko/mlcourse.ai/blob/master/jupyter_english/topic06_features_regression/topic6_feature_engineering_feature_selection.ipynb
#
# 2. Feature engineering/preprocessing, using scikit-learn API (great code examples, but really brief explanation):
# https://scikit-learn.org/stable/modules/preprocessing
#
# 3. Feature scaling/normalization:
# https://towardsdatascience.com/all-about-feature-scaling-bcc0ad75cb35
#
# 4. Log Transform/power transform:
# https://medium.com/@kyawsawhtoon/log-transformation-purpose-and-interpretation-9444b4b049c9
#
# 6. Missing values preprocessing using scikit-learn API (great code examples, great explanation):
# https://scikit-learn.org/stable/modules/impute.html
#
# 7. Feature selection scikit-learn API (great code examples, great explanation):
# https://scikit-learn.org/stable/modules/feature_selection.html
#
# 8. Melbourne housing dataset source:
# https://www.kaggle.com/anthonypino/melbourne-housing-market
| 6_feature_engineering_selection/feature_engineering_selection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="dk_hGQEvfuRP"
# ## Dependencies
# + colab={"base_uri": "https://localhost:8080/"} id="iq8x9YBKa2nt" outputId="8a7258bc-8f1b-4a9b-c235-1a774429d5d5"
if 'google.colab' in str(get_ipython()):
print('Running on CoLab')
# !pip install fuzzywuzzy[speedup]
else:
print("No running in colab")
# + id="vMjLOhNq-aFa"
import pandas as pd
import ast
import itertools
import numpy as np
from sklearn.neighbors import NearestNeighbors
from sklearn.preprocessing import MinMaxScaler
from fuzzywuzzy import process
from time import process_time
import pickle
print('all imported')
# + [markdown] id="mGLLfbvHbxtd"
# ##DataFrame
#
# + id="g_JmThSwN_0t"
df_track = pd.read_csv('https://raw.githubusercontent.com/vjmiyagi/DS_BuildWeek_Dec_2020/main/csv/data.csv')
# + colab={"base_uri": "https://localhost:8080/"} id="Au6xjkKQ9M4Z" outputId="8d31abe3-6937-4cee-8327-307c72ed517a"
df_track.info()
# + colab={"base_uri": "https://localhost:8080/"} id="WO92wJioEe2G" outputId="63aaee25-85b3-4ff3-c094-9c9a2bee49cd"
df_track['artists']
# + [markdown] id="3SHIqo8QD6UH"
# ##Spotify df
#
#
# + id="XcR2T-Suz2Jy"
#converting string to a list using ast.literal_eval
# + id="NvvrHlZpOO9d" colab={"base_uri": "https://localhost:8080/"} outputId="e28a8321-ab67-4bcd-a90b-bbd5662d6435"
df_track.artists = df_track.artists.apply(ast.literal_eval)
# returning a list store in a string, str('[a,g,c]') --> list [a,g,c]
type(df_track.artists[0])
# + colab={"base_uri": "https://localhost:8080/"} id="MaD0rMjuzZgN" outputId="2c26a6e8-c632-4841-9c6a-fa8359c71019"
df_track.artists[0]
# + id="eiSt6KW1UU90"
# " ,".join(df_track.artists[0])
#testing joining a list to a single string separated by delimiter
# + id="dxKT-ohzSKzO"
def lst_join(list_artists):
"""transform from list to string
to get rid of the [] of the list"""
Jointed_list = ", ".join(list_artists)
return Jointed_list
df_track.artists = df_track.artists.apply(lst_join)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="RHc0Mdb-Tq_A" outputId="8a8fcd4b-e585-4bec-8f0f-aaa67c144820"
df_track[['artists','id']].head()
# + colab={"base_uri": "https://localhost:8080/"} id="T3qjbGj32trY" outputId="4cb6846e-c87f-4249-d4be-7c538ed5fbaa"
list(df_track.columns.values)
# + [markdown] id="sLVY_Is4fLJ4"
# ## Normalize, Standarize, Model
#
# + id="qAEXOPwZ2poc"
f_labels = ['valence', 'year', 'acousticness', 'danceability',
'duration_ms', 'energy', 'explicit', 'instrumentalness',
'key', 'liveness', 'loudness', 'mode','popularity', 'speechiness', 'tempo']
#selected columns to pass to normalize and pass as a Matrix
# + id="20bbRX0l3sz7"
scaler = MinMaxScaler()
# + id="dUWXIERl3yZH"
spotify = df_track.copy()
# + id="1gigkgPP3_bv"
spotify[f_labels] = scaler.fit_transform(spotify[f_labels])
# + id="lOlbeXAU02II"
model_knn = NearestNeighbors(metric='cosine', algorithm='brute', n_neighbors=5)
# + id="TFxG4FGxMPMp"
spotify_matrix = spotify[f_labels].to_numpy()
# + colab={"base_uri": "https://localhost:8080/"} id="cbAzCwspMOPo" outputId="b5f0e39b-6c06-4eab-92cd-0119b06224dc"
model_knn.fit(spotify_matrix)
# + colab={"base_uri": "https://localhost:8080/"} id="kkDdqchB_iBX" outputId="94d58233-4cfa-4b95-eb24-86b1c036dee7"
spotify_matrix[0]
# + colab={"base_uri": "https://localhost:8080/"} id="-vgZjXi45Lb0" outputId="e4a9ca40-44c4-421b-a6a1-12241b1b212a"
t1_start = process_time()
track_search = process.extractOne('Clancy Lowered the Boom', spotify['name'])
t1_stop = process_time()
print("Elapsed time:", t1_stop, t1_start)
print("Elapsed time during the whole program in seconds:",
t1_stop-t1_start)
# + colab={"base_uri": "https://localhost:8080/"} id="P7ldbrxz7OKF" outputId="2f192100-8f3d-4094-fe60-21ed3acbd95e"
track_search
# + id="HWZLZbePMFyN"
def song_recomender(name_song, n_songs):
"""returns indices from the model,
pass them to df.loc[[]]"""
id_track = process.extractOne(name_song,spotify['name'])[2]
print('Song selected', spotify['name'][id_track], 'id:', id_track)
print('searching for recomendation....')
distances, ids = model_knn.kneighbors(spotify_matrix[[id_track]], n_neighbors=n_songs)
# for i in ids:
# print(spotify[['name','artists','id']].loc[i])
return ids
# + colab={"base_uri": "https://localhost:8080/"} id="uxSkzVauXZxa" outputId="75834c98-4ff6-4784-a9df-28f11318dab1"
song = input('type name of the song and press Enter in your keyboard:')
print("will take a few seconds")
recomendations_20 = list(song_recomender(song,20)[0])
print(recomendations_20, type(recomendations_20))
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="Y17nFycGNeQN" outputId="f581ca99-c1f9-4eb2-b576-1437c75aad8d"
spotify.loc[recomendations_20]
# + [markdown] id="NZLc5LmjClgU"
# ##Save and pickle
# + id="2FVj6tcAhBOx"
if 'google.colab' in str(get_ipython()):
print('Running on CoLab')
from google.colab import files
else:
print("No running in colab")
# + id="Wf_6tH8I8NmI"
filename = './csv/spotify_track_neighbors.urenaj'
pickle.dump(model_knn, open(filename, 'wb'))
#pickle sklearn model
# + id="T9rV7Czo90tM"
#How to un-pickle
# knn_2nd = pickle.load(open(filename, 'rb'))
# distances, x_ids = knn_2nd.kneighbors(spotify_matrix[[0]],3,return_distance=True)
# + id="DK1IwhfyCsr0"
X = spotify_matrix
np.savetxt("./csv/spotify_matrix.csv", X, delimiter=",")
#how to save or pickle np.array
# + id="umYSrvs-GTAQ"
#load np.array
# np.loadtxt("spotify_matrix.csv",delimiter=",")
# + id="dN-rWeNydc5I"
spotify.to_csv('./csv/spotify_tracks.csv',index=False)
# -
| spotify_exploration_wrangling_JU.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Array
#
# Array is a container which can hold a fix number of items and these items should be of the same type.
#
# ```
# arrayName = array(typecode, [Initializers])
# ```
# Typecode are the codes that are used to define the type of value the array will hold. Some common typecodes used are as follows
#
# | Typecode | Value |
# |---|---|
# | b | Represents signed integer of size 1 byte |
# | B | Represents unsigned integer of size 1 byte |
# | c | Represents character of size 1 byte |
# | i | Represents signed integer of size 2 bytes |
# | I | Represents unsigned integer of size 2 bytes |
# | f | Represents floating point of size 4 bytes |
# | d | Represents floating point of size 8 bytes |
# creates an array named array1
from array import *
array1 = array('i', [10,20,30,40,50])
for x in array1:
print(x)
## Accessing Array Element
print(f"The first element: {array1[0]}")
print(f"The first two element: {array1[:2]}")
print(f"The middle two element: {array1[1:4]}")
print(f"The last element: {array1[-1]}")
print(f"The last three element: {array1[-4:]}")
# +
# Insertion Operation
# Insert operation is to insert one or more data elements into an array. Based on the requirement,
# a new element can be added at the beginning, end, or any given index of array.
print(f"Original array: {array1}")
array1.insert(1,60)
print(f"revised array: {array1}")
# -
# ## Deletion Operation
#
# Deletion refers to removing first one existing element from the array and re-organizing all elements of an array.
#
# print(f"Original array: {array1}")
# array1.remove(60)
# print(f"revised array: {array1}")
#
# Search Operation
# You can perform a search for an array element based on its value or its index.
print (array1.index(50))
# +
# Update Operation
# Update operation refers to updating an existing element from the array at a given index.
print(f"Original array: {array1}")
array1[2] = 80
print(f"revised array: {array1}")
# -
# # List
#
# The list is a most versatile datatype available in Python which can be written as a list of comma-separated values (items) between square brackets.
#
# **Important thing about a list is that items in a list need not be of the same type, while for a array they should be same type**
#
# +
# create lists
list1 = ['physics', 'chemistry', 1997, 2000]
list2 = [1, 2, 3, 4, 5 ]
list3 = ["a", "b", "c", "d"]
print(f"list1: {list1}")
print(f"list2: {list2}")
print(f"list3: {list3}")
# -
# Accessing Values
# To access values in lists, use the square brackets for slicing along with the index
# or indices to obtain value available at that index.
print("list1[0]: ", list1[0])
print("list2[1:5]: ", list2[1:5])
# Updating Lists
# You can update single or multiple elements of lists by giving the slice on the left-hand side of the assignment operator
print("Value available at index 2 : ", list1[2])
list1[2] = 2001
print("New value available at index 2 : ", list1[2])
# # Tuple
#
# A tuple is a sequence of immutable Python objects. Tuples are sequences, just like lists. The differences between tuples and lists:
# - the tuples cannot be changed unlike lists
# - tuples use parentheses, (a, b, c), whereas lists use square brackets, [a, b, c].
# +
# Updating Tuples
# Tuples are immutable which means you cannot update or change the values of tuple elements.
# You are able to take portions of existing tuples to create new tuples as the following example demonstrates
tup1 = (12, 34.56)
tup2 = ('abc', 'xyz')
# So let's create a new tuple as follows
tup3 = tup1 + tup2
print(f"New tuple: {tup3}")
# Following action is not valid for tuples
tup1[0] = 100
# +
# Delete Tuple Elements
# Removing individual tuple elements is not possible.
# There is, of course, nothing wrong with putting together
# another tuple with the undesired elements discarded.
# To explicitly remove an entire tuple, just use the del statement.
# -
# # Dictionary
#
# - In Dictionary each key is separated from its value by a colon (:)
# - the items are separated by commas
# - the whole thing is enclosed in curly braces.
# - An empty dictionary without any items is written with just two curly braces, like this {}.
dict = {'Name': 'Zara', 'Age': 7, 'Class': 'First'}
print("dict['Name']: ", dict['Name'])
print("dict['Age']: ", dict['Age'])
# # ChainMap
#
# It is a type of data structure to manage multiple dictionaries together as one unit.
# - Removing duplicate keys. If there are duplicate keys, then only the value from the first key is preserved.
# - The best use of ChainMap is to search through multiple dictionaries at a time and get the proper key-value pair mapping
# - We also see that these ChainMaps behave as stack data structure.
# +
# Examples
import collections
dict1 = {'day1': 'Mon', 'day2': 'Tue'}
dict2 = {'day3': 'Wed', 'day1': 'Thu'}
res = collections.ChainMap(dict1, dict2)
print(res)
# +
# Creating a single dictionary
print(res.maps,'\n')
print('Keys = {}'.format(list(res.keys())))
print('Values = {}'.format(list(res.values())))
print()
# -
# Print all the elements from the result
print('elements:')
for key, val in res.items():
print('{} = {}'.format(key, val))
print()
# Find a specific value in the result
print('day3 in res: {}'.format(('day1' in res)))
print('day4 in res: {}'.format(('day4' in res)))
# +
# Updating Map
# When the element of the dictionary is updated, the result is instantly updated in
# the result of the ChainMap. In the below example we see that the new updated value
# reflects in the result without explicitly applying the ChainMap method again.
print(res.maps,'\n')
dict2['day4'] = 'Fri'
print(res.maps,'\n')
# -
# # 链表
#
# 1) 含义:链表(Linked list)是一种常见的基础数据结构,是一种线性表,但是并不会按线性的顺序存储数据,而是在每一个节点里存到下一个节点的指针(Pointer)。由于不必须按顺序存储,链表在插入的时候可以达到O(1)的复杂度,比另一种线性表顺序表快得多,但是查找一个节点或者访问特定编号的节点则需要O(n)的时间,而顺序表相应的时间复杂度分别是O(logn)和O(1)
#
# 2) 特点:使用链表结构可以克服数组链表需要预先知道数据大小的缺点,链表结构可以充分利用计算机内存空间,实现灵活的内存动态管理。但是链表失去了数组随机读取的优点,同时链表由于增加了结点的指针域,空间开销比较大
#
# 3) 操作:
# 1. is_empty() 链表是否为空
# 3. length() 链表长度
# 3. travel() 遍历链表
# 4. add(item) 链表头部添加
# 5. append(item) 链表尾部添加
# 6. insert(pos, item) 指定位置添加
# 7. remove(item) 删除节点
# 8. search(item) 查找节点是否存在
class LinkNode(object):
def __init(self, item, prev=None, next=None):
self.item = item
self.prev = prev
self.next = next
class DLinkList(object):
def __init__(self):
self.head = None
self.count = 0
def add(self, item):
node = LinkNode(item)
if self.head == None:
self.head = node
node.prev = node
node.next = node
else:
node.next = self.head
self.head.prev = node
self.head = node
self.count += 1
def append(self, item):
pass
def is_empty(self):
return self._head == None
def length(self):
return self.count
# # 堆栈
#
# 1. 含义:堆栈(英语:stack),也可直接称栈,在计算机科学中,是一种特殊的串列形式的数据结构,它的特殊之处在于只能允许在链接串列或阵列的一端(称为堆叠顶端指标,英语:top)进行加入资料(英语:push)和输出资料(英语:pop)的运算。另外堆叠也可以用一维阵列或连结串列的形式来完成。堆叠的另外一个相对的操作方式称为伫列;由于堆叠数据结构只允许在一端进行操作,因而按照后进先出(LIFO, Last In First Out)的原理运作
#
# 2. 特点:先入后出,后入先出;除头尾节点之外,每个元素有一个前驱,一个后继
class Stack(object):
def __init__(self):
self.data = []
def length(self):
return len(self.data)
def is_empty(self):
return self.length == 0
def push(self, item):
self.data.append(item)
def pop(self, item):
return self.data.pop()
# # 队列
#
# 1) 含义:和堆栈类似,唯一的区别是队列只能在队头进行出队操作,所以队列是是先进先出(FIFO, First-In-First-Out)的线性表
#
# 2) 特点:先入先出,后入后出;除尾节点外,每个节点有一个后继;(可选)除头节点外,每个节点有一个前驱
class Queue(object):
def __init__(self):
self.data = []
def dequeue(self):
return self.data.pop(0) if self.data != [] else None
def inqueue(self, item):
self.data.append(item)
# # Dequeue
#
# A double-ended queue, or deque, supports adding and removing elements from either end. he more commonly used stacks and queues are degenerate forms of deques, where the inputs and outputs are restricted to a single end.
# +
import collections
dq = collections.deque(["Monday","Tuesday", "Wesday", "Thursday"])
print(f"Double ended queue: {dq}")
dq.append('Friday'); print(f"Appended at right: {dq}")
dq.appendleft("Sunday"); print(f"Appended at left: {dq}")
dq.pop(); print(f"Deleting from right: {dq}")
dq.popleft(); print(f"Deleting from left: {dq}")
# -
# # 二叉树
#
# 1)定义:二叉树是每个结点最多有两个子树的树结构。它有五种基本形态:二叉树可以是空集;根可以有空的左子树或右子树;或者左、右子树皆为空
# 2)特点:
# 1. 性质1:二叉树第i层上的结点数目最多为$2^{i-1}$(i>=1);
# 2. 性质2:深度为k的二叉树至多有$2^{k-1}$个结点(k>=1);
# 3. 性质3:包含n个结点的二叉树的高度至少为$log_{2} n+1$;
# 4. 性质4:在任意一棵二叉树中,若终端结点的个数为$n_0$,度为2的结点数为$n_2$,则$n_0=n_2+1$
class TreeNode(object):
def __init__(self, item):
self.item = item
self.left_child = None
self.right_child = None
class Tree(object):
def __init__(self):
self.root = None
def add(self, item):
node = TreeNode(item)
if self.root is None:
self.root = node
else:
q = [self.root]
while True:
pop_node = q.pop(0)
if pop_node.left_child is None:
pop_node.left_child = node
break
elif pop_node.right_child is None:
pop_node.right_child = node
break
else:
q.append(pop_node.left_child)
q.append(pop_node.right_child)
def traverse(self):
if self.root is None:
return None
q = [self.root]
res = [self.root.item]
while q != []:
pop_node = q.pop(0)
if pop_node.left_child is not None:
q.append(pop_node.left_child)
res.append(pop_node.left_child.item)
if pop_node.right_child is not None:
q.append(pop_node.right_child)
res.append(pop_node.right_child.item)
return res
def traverse_preorder(self, root, res=None): # 先序遍历
if res is None: res = []
def preorder(root, res):
if root is None: return None
res.append(root.item)
preorder(root.left_child, res)
preorder(root.right_child, res)
preorder(root, res)
return res
def traverse_inorder(self, root, res=None): # 中序序遍历
if res is None: res = []
def inorder(root, res):
if root is None: return None
inorder(root.left_child, res)
res.append(root.item)
inorder(root.right_child, res)
inorder(root, res)
return res
def traverse_postorder(self, root, res=None): # 中序序遍历
if res is None: res = []
def postorder(root, res):
if root is None: return None
postorder(root.left_child, res)
postorder(root.right_child, res)
res.append(root.item)
postorder(root, res)
return res
t = Tree()
for i in range(10):
t.add(i)
# 层序遍历
print(t.traverse())
print(t.traverse_preorder(t.root))
print(t.traverse_inorder(t.root))
print(t.traverse_postorder(t.root))
# # Container datatypes --- collections
# ## Counter
#
# A Counter is a dict subclass for counting hashable objects. It is a collection where elements are stored as dictionary keys and their counts are stored as dictionary values. Counts are allowed to be any integer value including zero or negative counts. The Counter class is similar to bags or multisets in other languages.
# +
from collections import Counter
# 1. initial to take a count in two ways:
cnt = Counter()
for word in ['red', 'blue', 'red', 'green', 'blue', 'blue']:
cnt[word] += 1
print(f"after count: {cnt}")
print(f"Counting directly from array: {Counter(['red', 'blue', 'red', 'green', 'blue', 'blue'])}")
# -
# More examples from
print(f"a new, empty counter {Counter()}")
print(f"a new counter from an iterable {Counter('gallahad') }")
print(f"a new counter from a mapping {Counter({'red': 4, 'blue': 2})}")
print(f"a new counter from keyword args {Counter(cats=4, dogs=8) }")
# Useful APIs
c = Counter(a=4, b=2, c=0, d=-2)
print(c)
print(f"elements: {list(c.elements())}")
# +
#
c = Counter('abracadabra')
top3 = c.most_common(3)
n = 2
least_n = c.most_common()[:-n-1:-1] # n least common elements
print(f"Original Count: {c}")
print(f"Top 3 most common: {top3}")
print(f"N least common elements: {least_n}")
# -
# # Heap
#
# A heap is a binary tree inside an array. A heap is sorted based on the "heap property" that determines the order of the nodes in the tree.
#
# \begin{definition}[Heap]
# Suppose that $arr = [k_0, k_1, k_2, \ldots, k_{n-1}]$, $\forall i \in [0, \frac{n - 2}{2}]$
# \begin{itemize}
# \item \textbf{Max-Heap}
# \begin{equation}
# \begin{cases}
# k_i \geq k_{2i + 1} \\
# k_i \geq k_{2i + 2}
# \end{cases}
# \end{equation}
# \item \textbf{Min-Heap}
# \begin{equation}
# \begin{cases}
# k_i \leq k_{2i + 1} \\
# k_i \leq k_{2i + 2}
# \end{cases}
# \end{equation}
# \end{itemize}
# \end{definition}
#
# If $i$ is the index of a node, then the following formulas give the array indices of its parent and child nodes:
# \begin{aligned}
# parent(i) &= floor(\frac{i - 1}{2}) \\
# left(i) &= 2i + 1 \\
# right(i) &= left(i) + 1 = 2i + 2 \\
# \end{aligned}
# The left and right nodes are always stored right next to each other.
# ## Heap Property
# +
# 1. Using Max- or Min-heap property to verify that if an array is a heap.
arr = [ 10, 7, 2, 5, 1 ]
import math
def parent(index):
return math.floor((index - 1) / 2)
def left(index):
return 2*index + 1
def right(index):
return 2*index + 2
hold_props = [arr[parent(i)] >= arr[i] for i in range(1, len(arr))]
print(hold_props)
# -
# - In Max-heap, all parent nodes is bigger or equal than their children nodes, so the largest item at the root of the tree.
# - In Min-heap, all parent nodes is smaller or equal than their children nodes, so the smallest item at the root of the tree.
# - The root of the heap has the maximum or minimum element, but the sort order of other elements are not predictable.
# ## Heap vs Regular Tree (Binary search tree)
#
# - **Order of the nodes.** In a Binary search tree (BST), the left child must be smaller than its parent, and the right child must be greater. This is not true for a heap. In a max-heap both children must be smaller than the parent, while in a min-heap they both must be greater.
# - **Memory.** Traditional trees take up more memory than just the data they store. You need to allocate additional storage for the node objects and pointers to the left/right child nodes. A heap only uses a plain array for storage and uses no pointers.
# - **Balancing.** A binary search tree must be "balanced" so that most operations have O(log n) performance. You can either insert and delete your data in a random order or use something like an AVL tree or red-black tree, but with heaps we don't actually need the entire tree to be sorted. We just want the heap property to be fulfilled, so balancing isn't an issue. Because of the way the heap is structured, heaps can guarantee O(log n) performance.
# - **Searching.** Whereas searching is fast in a binary tree, it is slow in a heap. Searching isn't a top priority in a heap since the purpose of a heap is to put the largest (or smallest) node at the front and to allow relatively fast inserts and deletes.
# +
import math
class Heap:
def __init__(self):
self.data = []
@property
def size(self):
return len(self.data)
def insert(self, value):
"""
Adds the new element to the end of the heap
and then uses shiftUp() to fix the heap.
"""
self.data.append(value)
self.shiftUp(self.size - 1)
def remove(self):
"""
Removes and returns the maximum value (max-heap) or
the minimum value (min-heap). To fill up the hole left
by removing the element, the very last element is moved
to the root position and then shiftDown() fixes up the heap.
(This is sometimes called "extract min" or "extract max".)
"""
pass
def removeAtIndex(self, index):
"""
Just like remove() with the exception that it allows you to
remove any item from the heap, not just the root. This calls
both shiftDown(), in case the new element is out-of-order with
its children, and shiftUp(), in case the element is out-of-order
with its parents.
"""
pass
def replace(self, index, value):
"""
Assigns a smaller (min-heap) or larger (max-heap) value to a node.
Because this invalidates the heap property, it uses shiftUp() to
patch things up. (Also called "decrease key" and "increase key".)
"""
pass
def search(self, value):
"""
Heaps are not built for efficient searches, but the replace() and
removeAtIndex() operations require the array index of the node,
so you need to find that index. Time: O(n).
"""
pass
def buildHeap(self, array):
"""
Converts an (unsorted) array into a heap by repeatedly calling insert().
If you are smart about this, it can be done in O(n) time.
"""
pass
def peek(self):
"""
The heap also has a peek() function that returns the maximum (max-heap)
or minimum (min-heap) element, without removing it from the heap.
Time: O(1).
"""
return self.data[0] if self.size() > 0 else None
def shiftUp(self, index):
"""
If the element is greater (max-heap) or smaller (min-heap)
than its parent, it needs to be swapped with the parent.
This makes it move up the tree.
Shifting up or down is a recursive procedure that takes O(log n) time.
"""
parent_index = self.parent(index)
while index > 1 and self.data[parent_index] > self.data[index]:
self.data[parent_index], self.data[index] = self.data[index], self.data[parent_index]
index = parent_index
parent_index = self.parent(index)
def shiftDown(self, index):
"""
If the element is smaller (max-heap) or greater (min-heap)
than its children, it needs to move down the tree.
This operation is also called "heapify".
Shifting up or down is a recursive procedure that takes O(log n) time.
"""
pass
def heapify(self):
pass
@staticmethod
def parent(index):
return math.floor((index - 1) / 2)
@staticmethod
def left(index):
return 2 * index + 1
@staticmethod
def right(index):
return 2 * index + 2
if __name__ == "__main__":
pass
# -
# ## Heap queue algorithm in Python
#
# This module provides an implementation of the heap queue algorithm, also known as the priority queue algorithm.
# +
import heapq
# 1. Creat a heap
h1 = []
data = [1, 3, 5, 7, 9, 2, 4, 6, 8, 0]
heapq.heapify(data) # Transform list x into a heap, in-place, in linear time.
print(data)
# -
# Push the value item onto the heap, maintaining the heap invariant.
heapq.heappush(data, 3)
print(data)
# +
# Pop and return the smallest item from the heap, maintaining the heap
# invariant. If the heap is empty, IndexError is raised. To access the
# smallest item without popping it, use heap[0].
result = heapq.heappop(data)
print(data)
print(result)
# -
def heapsort(iterable):
h = []
# build a min-heap
for value in iterable:
heapq.heappush(h, value)
return [heapq.heappop(h) for _ in range(len(iterable))]
heapsort([1, 3, 2, 6, 3, 5, 4, 7, 8, 9])
# # Reference
#
# 1. [Python 中常见的数据结构](https://zhuanlan.zhihu.com/p/69487899)
# 2. [Python对数据结构的实现](https://blog.csdn.net/mxz19901102/article/details/80071864?utm_medium=distribute.pc_relevant.none-task-blog-BlogCommendFromMachineLearnPai2-2.control&depth_1-utm_source=distribute.pc_relevant.none-task-blog-BlogCommendFromMachineLearnPai2-2.control)
# 3. [常见的数据结构](https://zhuanlan.zhihu.com/p/93928546)
# 4. [Heap](https://github.com/raywenderlich/swift-algorithm-club/tree/master/Heap)
| Python/Common Data Struture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # End to End ML with Metaflow and Tempo
#
# We will train two models and deploy them with tempo within a Metaflow pipeline. To understand the core example see [here](https://tempo.readthedocs.io/en/latest/examples/multi-model/README.html)
#
# 
#
# ## MetaFlow Prequisites
#
#
# ### Install metaflow locally
#
# ```
# pip install metaflow
# ```
#
# ### Setup Conda-Forge Support
#
# The flow will use conda-forge so you need to add that channel to conda.
#
# ```
# conda config --add channels conda-forge
# ```
#
#
# ## Iris Flow Summary
# !python src/irisflow.py --environment=conda show
# ## Run Flow locally to deploy to Docker
#
# To run the workflow with a local Docker deployment use the flag:
#
# ```
# --tempo-on-docker true
# ```
#
# !python src/irisflow.py --environment=conda run
# ## Make Predictions with Metaflow Tempo Artifact
from metaflow import Flow
import numpy as np
run = Flow('IrisFlow').latest_run
client = run.data.client_model
client.predict(np.array([[1, 2, 3, 4]]))
# ## Run Flow on AWS and Deploy to Remote Kubernetes
#
# We will now run our flow on AWS Batch and will launch Tempo artifacts onto a remote Kubernetes cluster.
#
# ### Setup AWS Metaflow Support
#
# Note at present this is required even for a local run as artifacts are stored on S3.
#
# [Install Metaflow with remote AWS support](https://docs.metaflow.org/metaflow-on-aws/metaflow-on-aws).
#
# ### Seldon Requirements
#
# For deploying to a remote Kubernetes cluster with Seldon Core installed do the following steps:
#
# #### Install Seldon Core on your Kubernetes Cluster
#
# Create a GKE cluster and install Seldon Core on it using [Ansible to install Seldon Core on a Kubernetes cluster](https://github.com/SeldonIO/ansible-k8s-collection).
#
#
# ### K8S Auth from Metaflow
#
# To deploy services to our Kubernetes cluster with Seldon Core installed, Metaflow steps that run on AWS Batch and use tempo will need to be able to access K8S API. This step will depend on whether you're using GKE or AWS EKS to run
# your cluster.
#
# #### Option 1. K8S cluster runs on GKE
#
# We will need to create two files in the flow src folder:
#
# ```bash
# kubeconfig.yaml
# gsa-key.json
# ```
#
# Follow the steps outlined in [GKE server authentication](https://cloud.google.com/kubernetes-engine/docs/how-to/api-server-authentication#environments-without-gcloud).
#
#
#
#
# #### Option 2. K8S cluster runs on AWS EKS
#
# Make note of two AWS IAM role names, for example find them in the IAM console. The names depend on how you deployed Metaflow and EKS in the first place:
#
# 1. The role used by Metaflow tasks executed on AWS Batch. If you used the default CloudFormation template to deploy Metaflow, it is the role that has `*BatchS3TaskRole*` in its name.
#
# 2. The role used by EKS nodes. If you used `eksctl` to create your EKS cluster, it is the role that starts with `eksctl-<your-cluster-name>-NodeInstanceRole-*`
#
# Now, we need to make sure that AWS Batch role has permissions to access the K8S cluster. For this, add a policy to the AWS Batch task role(1) that has `eks:*` permissions on your EKS cluster (TODO: narrow this down).
#
# You'll also need to add a mapping for that role to `aws-auth` ConfigMap in `kube-system` namespace. For more details, see [AWS docs](https://docs.aws.amazon.com/eks/latest/userguide/add-user-role.html) (under "To add an IAM user or role to an Amazon EKS cluster"). In short, you'd need to add this to `mapRoles` section in the aws-auth ConfigMap:
# ```
# - rolearn: <batch task role ARN>
# username: cluster-admin
# groups:
# - system:masters
# ```
#
# We also need to make sure that the code running in K8S can access S3. For this, add a policy to the EKS node role (2) to allow it to read and write Metaflow S3 buckets.
#
# ### S3 Authentication
# Services deployed to Seldon will need to access Metaflow S3 bucket to download trained models. The exact configuration will depend on whether you're using GKE or AWS EKS to run your cluster.
#
# From the base templates provided below, create your `k8s/s3_secret.yaml`.
#
# ```yaml
# apiVersion: v1
# kind: Secret
# metadata:
# name: s3-secret
# type: Opaque
# stringData:
# RCLONE_CONFIG_S3_TYPE: s3
# RCLONE_CONFIG_S3_PROVIDER: aws
# RCLONE_CONFIG_S3_BUCKET_REGION: <region>
# <...cloud-dependent s3 auth settings (see below)>
# ```
#
# For GKE, to access S3 we'll need to add the following variables to use key/secret auth:
# ```yaml
# RCLONE_CONFIG_S3_ENV_AUTH: "false"
# RCLONE_CONFIG_S3_ACCESS_KEY_ID: <key>
# RCLONE_CONFIG_S3_SECRET_ACCESS_KEY: <secret>
# ```
#
# For AWS EKS, we'll use the instance role assigned to the node, we'll only need to set one env variable:
# ```yaml
# RCLONE_CONFIG_S3_ENV_AUTH: "true"
# ```
#
# We provide two templates to use in the `k8s` folder:
#
# ```
# s3_secret.yaml.tmpl.aws
# s3_secret.yaml.tmpl.gke
# ```
#
# Use one to create the file `s3_secret.yaml` in the same folder
#
# ## Setup RBAC and Secret on Kubernetes Cluster
#
# These steps assume you have authenticated to your cluster with kubectl configuration
# !kubectl create ns production
# !kubectl create -f k8s/tempo-pipeline-rbac.yaml -n production
# Create a Secret from the `k8s/s3_secret.yaml.tmpl` file by adding your AWS Key that can read from S3 and saving as `k8s/s3_secret.yaml`
# !kubectl create -f k8s/s3_secret.yaml -n production
# ## Run Metaflow on AWS Batch
# !python src/irisflow.py \
# --environment=conda \
# --with batch:image=seldonio/seldon-core-s2i-python37-ubi8:1.10.0-dev \
# run
# ## Make Predictions with Metaflow Tempo Artifact
from metaflow import Flow
run = Flow('IrisFlow').latest_run
client = run.data.client_model
import numpy as np
client.predict(np.array([[1, 2, 3, 4]]))
| docs/examples/metaflow/README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from math import pi
import pandas as pd
from bokeh.sampledata.stocks import MSFT
from bokeh.plotting import figure, show, output_notebook
# +
df = pd.DataFrame(MSFT)[:50]
df['date'] = pd.to_datetime(df['date'])
mids = (df.open + df.close)/2
spans = abs(df.close-df.open)
inc = df.close > df.open
dec = df.open > df.close
w = 12*60*60*1000 # half day in ms
# -
output_notebook()
# +
p = figure(x_axis_type="datetime", plot_width=1000)
p.segment(df.date, df.high, df.date, df.low, color='black')
p.rect(df.date[inc], mids[inc], w, spans[inc], fill_color="#D5E1DD", line_color="black")
p.rect(df.date[dec], mids[dec], w, spans[dec], fill_color="#F2583E", line_color="black")
# -
p.title = "MSFT Candlestick"
p.xaxis.major_label_orientation = pi/4
p.grid.grid_line_alpha=0.3
show(p)
| examples/plotting/notebook/candlestick.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In statistics, polynomial regression is a form of regression analysis in which the relationship between the independent variable x and the dependent variable y is modelled as an nth degree polynomial in x. Polynomial regression fits a nonlinear relationship between the value of x and the corresponding conditional mean of y, denoted E(y |x), and has been used to describe nonlinear phenomena such as the growth rate of tissues,the distribution of carbon isotopes in lake sediments, and the progression of disease epidemics Although polynomial regression fits a nonlinear model to the data, as a statistical estimation problem it is linear, in the sense that the regression function E(y | x) is linear in the unknown parameters that are estimated from the data. For this reason, polynomial regression is considered to be a special case of multiple linear regression.
import numpy as np
# create arrays of fake points
x = np.array([0.0, 1.0, 2.0, 3.0, 4.0, 5.0])
y = np.array([0.0, 0.8, 0.9, 0.1, -0.8, -1.0])
# fit up to deg=3
z = np.polyfit(x, y, 3)
z
np.roots(z)
# **Using scikit-learn's PolynomialFeatures**
#
# Generate polynomial and interaction features
# Generate a new feature matrix consisting of all polynomial combinations of the features with degree less than or equal to the specified degree
# +
# Importing the libraries
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# Importing the dataset
datas = pd.read_csv('data.csv')
datas
# -
X = datas.iloc[:, 1:2].values
y = datas.iloc[:, 2].values
# Fitting Linear Regression to the dataset
from sklearn.linear_model import LinearRegression
lin = LinearRegression()
lin.fit(X, y)
# Visualising the Linear Regression results
plt.scatter(X, y, color = 'blue')
plt.plot(X, lin.predict(X), color = 'gray')
plt.title('Linear Regression')
plt.xlabel('Temperature')
plt.ylabel('Pressure')
plt.show()
# +
# Fitting Polynomial Regression to the dataset
from sklearn.preprocessing import PolynomialFeatures
poly = PolynomialFeatures(degree = 4)
X_poly = poly.fit_transform(X)
poly.fit(X_poly, y)
lin2 = LinearRegression()
lin2.fit(X_poly, y)
# -
# Visualising the Polynomial Regression results
plt.scatter(X, y, color = 'blue')
plt.plot(X, lin2.predict(poly.fit_transform(X)), color = 'red')
plt.title('Polynomial Regression')
plt.xlabel('Temperature')
plt.ylabel('Pressure')
plt.show()
| Regression/Polynomial_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import sys
sys.path.append('..')
import keras_multilabel_embedding as kml
import tensorflow as tf
# ## Multi-label embedding (fixed number of labels)
# +
x_ids = [[1, 2, 4], [0, 1, 2], [2, 1, 4], [3, 2, 1]]
x_ids = tf.constant(x_ids)
layer1a = kml.MultiLabelEmbedding(
vocab_size=500000, embed_size=300, random_state=42)
# %timeit y1a = layer1a(x_ids)
# -
# %%time
for _ in range(100):
y1a = layer1a(x_ids)
print(y1a.shape)
# ## Multi-label embedding (variable number of labels)
# +
x_ids = [[1, 2, 4], [0, 1, 2], [2, 1], [3]]
layer1b = kml.MultiLabelEmbedding2(
vocab_size=500000, embed_size=300, random_state=42)
# %timeit y1b = layer1b(x_ids)
# -
# %%time
for _ in range(100):
y1b = layer1b(x_ids)
print(y1b.shape)
# ## Simple dense embedding
# +
# x_ids = [[1, 2, 4], [0, 1, 2], [2, 1, 4], [3, 2, 1]]
x_mask = [[0., 1, 1, 0, 1], [1, 1, 1, 0, 0], [0, 1, 1, 0, 1], [0, 1, 1, 1, 0]]
x_mask = [ex + [0]*(500000-5) for ex in x_mask]
x_mask = tf.constant(x_mask)
layer2 = tf.keras.layers.Dense(300, use_bias=False)
tf.keras.utils.set_random_seed(42)
layer2.build(input_shape=(None, 500000))
layer2.set_weights(layer1a.get_weights())
# %timeit y2 = layer2(x_mask)
# -
# %%time
for _ in range(10):
y2 = layer2(x_mask)
print(y2.shape)
| demo/Runtime Speed Tests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Author: <NAME>
# ### Date Created: 5 Oct 2018
# ## This example jupyer notebook uses clmm code to generate mock data with a set of configuration parameters, producing a catalog of ideal and noisy example data.
import matplotlib.pyplot as plt
# %matplotlib inline
# ### Import mock data module and setup the configuration
import clmm.mock_data as mock
# ### Mock data generation requires a config dictionary
# +
config={}
config['cluster_m'] = 1.e15
config['cluster_z'] = 0.3
config['src_z'] = 0.8
config['concentration'] = 4
config['cosmo'] = 'WMAP7-ML'
config['ngals'] = 10000
config['mdef'] = '200c'
ideal_data = mock.MockData(config=config)
noisy_data = mock.MockData(config=config)
noisy_data_z = mock.MockData(config=config)
# -
# ### Generate the mock catalog from the configuration. Consider 3 examples of increasing complexity:
# * Ideal data, no noise, all galaxies at the same redshift
# * Noisy data, including shape noise and redshift error, all galaxies at the same redshift
# * Noisy data, galaxies following redshift distribution, redshift error, shape noise
ideal_data.generate()
noisy_data.generate(is_shapenoise=True, shapenoise=0.005, is_zerr=True)
noisy_data_z.generate(is_shapenoise=True, is_zdistribution=True, is_zerr=True)
# #### Ideal catalog first entries: no noise on the shape measurement, all galaxies at z=0.3
ideal_data.catalog[0:5]
# #### More realistic catalog: noise has been added on the shape measurement, galaxies follow some photoz distribution, gaussian pdf for each photoz is also provided (along with corresponding bins)
noisy_data_z.catalog[0:2]
# Histogram of the redshift distribution of bkg galaxies (starting at z_cluster + 0.1)
hist = plt.hist(noisy_data_z.catalog['z'], bins=50)
# pdz for the first galaxy in the catalog
plt.plot(noisy_data_z.catalog['z_bins'][0],noisy_data_z.catalog['z_pdf'][0])
# ### Compute and plot shear profile
from clmm import ShearAzimuthalAverager
# +
cl_dict = {'z':config['cluster_z'], 'ra':0.0, 'dec': 0.0}
saa_ideal = ShearAzimuthalAverager(cl_dict,ideal_data.catalog)
saa_noisy = ShearAzimuthalAverager(cl_dict,noisy_data.catalog)
saa_noisy_z = ShearAzimuthalAverager(cl_dict,noisy_data_z.catalog)
saa_ideal.compute_shear()
saa_noisy.compute_shear()
saa_noisy_z.compute_shear()
saa_ideal.make_shear_profile()
saa_noisy.make_shear_profile()
saa_noisy_z.make_shear_profile()
# -
# Ideal data
saa_ideal.plot_profile()
# Noisy data, all galaxies at the same redshift
saa_noisy.plot_profile()
# Noisy data, galaxies following redshift distribution
saa_noisy_z.plot_profile()
| examples/generate_mock_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
#Current version of seaborn generates a bunch of warnings that we will ignore
warnings.filterwarnings('ignore')
sns.set_style('whitegrid')
import gc
import datetime
# %matplotlib inline
color = sns.color_palette()
# +
df = pd.read_csv('C:\\Users\\TOPTAS\\Desktop\\data.csv',encoding = 'ISO-8859-1')
#We must mention the following encoding to avoid errors
df.head()
# -
df.shape
#Rename headers for making our work easier
df.rename(index=str, columns={'InvoiceNo': 'invoice_num',
'StockCode' : 'stock_code',
'Description' : 'description',
'Quantity' : 'quantity',
'InvoiceDate' : 'invoice_date',
'UnitPrice' : 'unit_price',
'CustomerID' : 'cust_id',
'Country' : 'country'}, inplace=True)
df.head()
#Also changing datetime for easy work
df['invoice_date'] = pd.to_datetime(df.invoice_date, format='%m/%d/%Y %H:%M')
#lowercase descriptions are more efficient
df['description'] = df.description.str.lower()
# df_new without missing values
df_new = df.dropna()
# change columns tyoe - String to Int type
df_new['cust_id'] = df_new['cust_id'].astype('int64')
df_new.head()
#Eliminate products that is not in stocks
df_new = df_new[df_new.quantity > 0]
#Get statictics & Using numbers with 2 decimal points is more efficient
df_new.describe().round(2)
#For to get amount that's spent, we multiply quantities with unit prices
df_new['amount_spent'] = df_new['quantity'] * df_new['unit_price']
df_new = df_new[['invoice_num','invoice_date','stock_code','description','quantity','unit_price','amount_spent','cust_id','country']]
#Creating our new table with new features
df_new.insert(loc=2, column='year_month', value=df_new['invoice_date'].map(lambda x: 100*x.year + x.month))
df_new.insert(loc=3, column='month', value=df_new.invoice_date.dt.month)
# # +1 to make Monday=1.....until Sunday=7
df_new.insert(loc=4, column='day', value=(df_new.invoice_date.dt.dayofweek)+1)
df_new.insert(loc=5, column='hour', value=df_new.invoice_date.dt.hour)
df_new.head()
#We categorize countries and customer IDs by counting the quantiy of invoices that each country have
df_new.groupby(by=['cust_id','country'], as_index=False)['invoice_num'].count().head()
# +
orders = df_new.groupby(by=['cust_id','country'], as_index=False)['invoice_num'].count()
#To show customers and their number of orders on graph
plt.subplots(figsize=(16,8))
plt.plot(orders.cust_id, orders.invoice_num)
plt.xlabel('Customers ID')
plt.ylabel('Number of Orders')
plt.title('Number of Orders for different customers')
plt.show()
# -
#Brits are dominating statistics here when we show TOP 5 customers who has invoice the most :)
print('The TOP 5 customers with most number of orders...')
orders.sort_values(by='invoice_num', ascending=False).head()
# +
#We sorted customers according to invoices they have, now it is time to find how much they spent totally
money_spent = df_new.groupby(by=['cust_id','country'], as_index=False)['amount_spent'].sum()
plt.subplots(figsize=(16,8))
plt.plot(money_spent.cust_id, money_spent.amount_spent)
plt.xlabel('Customers ID')
plt.ylabel('Money spent ($)')
plt.title('Money spent for different customers')
plt.show()
# -
#Dutchs are slightly passing Brits in TOP 5 money spent table :)
print('The TOP 5 customers with highest money spent...')
money_spent.sort_values(by='amount_spent', ascending=False).head()
#palplot is plotting the values in a color palette as a horizontal array
myColor = sns.color_palette("bright")
sns.palplot(myColor,2)
#It is time to analyze which months were better than others. Let's see that better on a bar graph
ax = df_new.groupby('invoice_num')['year_month'].unique().value_counts().sort_index().plot(kind='bar',color=color[0],figsize=(16,8))
ax.set_xlabel('Month',fontsize=15)
ax.set_ylabel('Number of orders',fontsize=15)
ax.set_title('Number of orders for different months (1st Dec 2010 - 9th Dec 2011)',fontsize=15)
ax.set_xticklabels(('Dec_10','Jan_11','Feb_11','Mar_11','Apr_11','May_11','Jun_11','July_11','Aug_11','Sep_11','Oct_11','Nov_11','Dec_11'), rotation='horizontal', fontsize=13)
plt.show()
#November has a significant increase compared to other months!
#Let's see now which days the customers ordered products more
#On Saturday the shop is closed so 6th day is excluded
df_new.groupby('invoice_num')['day'].unique().value_counts().sort_index()
#Visualize the results
ax = df_new.groupby('invoice_num')['day'].unique().value_counts().sort_index().plot(kind='bar',color=color[0],figsize=(16,8))
ax.set_xlabel('Day',fontsize=15)
ax.set_ylabel('Number of orders',fontsize=15)
ax.set_title('Number of orders for different days',fontsize=15)
ax.set_xticklabels(('Mon','Tue','Wed','Thur','Fri','Sun'), rotation='horizontal', fontsize=15)
plt.show()
#Let's go into details more. Check which hours are more efficient than others to sell products
df_new.groupby('invoice_num')['hour'].unique().value_counts().iloc[:-1].sort_index()
#Visualize the results for in working hours
ax = df_new.groupby('invoice_num')['hour'].unique().value_counts().iloc[:-1].sort_index().plot(kind = 'bar',color=color[0],figsize=(16,8))
ax.set_xlabel('Hour',fontsize=15)
ax.set_ylabel('Number of orders',fontsize=15)
ax.set_title('Number of orders for different hours',fontsize=15)
ax.set_xticklabels(range(6,21), rotation='horizontal', fontsize=15)
plt.show()
#Statistics for unit price details
df_new.unit_price.describe()
#Let's use a different graph to visualize unit price statistics
plt.subplots(figsize=(16,8))
sns.boxplot(df_new.unit_price)
plt.show()
#To find frequency properly, we need to make unit prices 0
df_free = df_new[df_new.unit_price == 0]
df_free.head()
#Visualize frequency of each months
ax = df_free.year_month.value_counts().sort_index().plot(kind = 'bar',figsize=(16,8), color=color[0])
ax.set_xlabel('Month',fontsize=15)
ax.set_ylabel('Frequency',fontsize=15)
ax.set_title('Frequency for different months (Dec 2010 - Dec 2011)',fontsize=15)
ax.set_xticklabels(('Dec_10','Jan_11','Feb_11','Mar_11','Apr_11','May_11','July_11','Aug_11','Sep_11','Oct_11','Nov_11'), rotation='horizontal', fontsize=13)
plt.show()
# +
group_country_orders = df_new.groupby('country')['invoice_num'].count().sort_values()
# plot number of unique customers in each country
plt.subplots(figsize=(16,8))
group_country_orders.plot(kind = 'barh', fontsize=12, color=color[0])
plt.xlabel('Number of Orders', fontsize=12)
plt.ylabel('Country', fontsize=12)
plt.title('Number of Orders for different Countries', fontsize=12)
plt.show()
# +
#We can see a UK dominance on the previous graph! I wonder about how the graph would be without the UK. Let's see!
group_country_orders = df_new.groupby('country')['invoice_num'].count().sort_values()
del group_country_orders['United Kingdom']
# plot number of unique customers in each country (without UK)
plt.subplots(figsize=(15,8))
group_country_orders.plot(kind = 'barh', fontsize=12, color=color[0])
plt.xlabel('Number of Orders', fontsize=12)
plt.ylabel('Country', fontsize=12)
plt.title('Number of Orders for different Countries', fontsize=12)
plt.show()
# +
group_country_amount_spent = df_new.groupby('country')['amount_spent'].sum().sort_values()
# plot total money spent by each country
plt.subplots(figsize=(15,8))
group_country_amount_spent.plot(kind = 'barh', fontsize=12, color=color[0])
plt.xlabel('Money Spent (Dollar)', fontsize=12)
plt.ylabel('Country', fontsize=12)
plt.title('Money Spent by different Countries', fontsize=12)
plt.show()
# +
group_country_amount_spent = df_new.groupby('country')['amount_spent'].sum().sort_values()
del group_country_amount_spent['United Kingdom']
# Another UK dominance... So let's see the graph without the UK
plt.subplots(figsize=(15,8))
group_country_amount_spent.plot(kind = 'barh', fontsize=12, color=color[0])
plt.xlabel('Money Spent (Dollar)', fontsize=12)
plt.ylabel('Country', fontsize=12)
plt.title('Money Spent by different Countries', fontsize=12)
plt.show()
# -
| btoptas_ecommercedataanalysis-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [anaconda2]
# language: python
# name: Python [anaconda2]
# ---
# <H2>MADATORY PYTHON LIBRARIES</H2>
import ftputil
import numpy as np
import pandas as pd
from shapely.geometry import box
import seaborn as sns
# %matplotlib inline
# <H2>AUXILIARY FUNCTIONS</H2>
def itemize_ftplink(indexfile):
"""returns the keys for the items resulting from splitting by '/' the file_name colummn in the index files"""
if indexfile == 'index_history.txt':
return ['access_protocol','blank', 'host', 'core', 'product_name', 'directory', 'platform_category', 'netCDF_file']
elif indexfile == 'index_monthly.txt':
return ['access_protocol','blank', 'host', 'core', 'product_name', 'directory', 'platform_category', 'timestamp', 'netCDF_file']
else:
return ['access_protocol','blank', 'host', 'core', 'product_name', 'directory', 'timestamp', 'netCDF_file']
def itemize_netCDFname(indexfile):
"""returns the keys for the items resulting from splitting by '_' the netCDF filename"""
if indexfile == 'index_history.txt':
return ['region', 'data', 'data_source', 'code', 'subsettingcode'] #RR_XX_YY_CODE<_ZZZ>.nc
elif indexfile == 'index_monthly.txt':
return ['region', 'timestamp', 'data', 'data_source', 'code'] #RR_YYYYMM_XX_YY_CODE.nc
else:
return ['region', 'fixedname', 'data', 'data_source', 'code', 'timestamp'] #RR_LATEST_XX_YY_CODE_YYYYMMDD.nc
# <h2> SET YOUR CREDENTIALS</h2>
user = '' #type CMEMS user name
password = '' #type CMEMS password
# <h2> TARGET A PRODUCT, HOST AND INDEX FILE</h2>
product_name = 'INSITU_MED_NRT_OBSERVATIONS_013_035' #type aimed In Situ product
host = 'nrt.cmems-du.eu' #type aimed host (nrt.cmems-du.eu or my.cmems-du)
index_file = 'index_latest.txt' #type aimed index file
# <h2>INDEX FILE RAW INFORMATION</h2>
#connect to CMEMS FTP
with ftputil.FTPHost(host, user, password) as ftp_host:
#open the index file to read
with ftp_host.open("Core"+'/'+product_name+'/'+index_file, "r") as indexfile:
raw_index_info = pd.read_csv(indexfile, skiprows=5) #load it as pandas dataframe
raw_index_info.head()
# <h2>INDEX FILE ADVANCED INFORMATION</h2>
raw_index_info[itemize_ftplink(index_file)] = raw_index_info['file_name'].str.split('/',expand=True) #split the ftplink in items
raw_index_info[['netCDF_file_name','netCDF_file_extension']] = raw_index_info['netCDF_file'].str.split('.',expand=True) #split the netCDF name in bare name and extension
raw_index_info[itemize_netCDFname(index_file)]= raw_index_info['netCDF_file_name'].str.split('_',expand=True) #split the netCDF name in items
raw_index_info.head()
# <h2>PARAMETERS BREAKDOWN </h2>
data = {'parameters':[]}
for parameters in raw_index_info['parameters'].tolist():
for item in parameters.split(' '):
if item not in data['parameters'] and item != '':
data['parameters'].append(item)
parameter_dataframe = pd.DataFrame(data=data)
print('There are %s parameters available in this In Situ product; examples:'%(len(parameter_dataframe)))
parameter_dataframe.head() #remove .head() if you wanna display full parameter list
# <h2>DATA SOURCES REPORTING A CERTAIN PARAMETER</h2>
parameter = 'TEMP'
parameter_subset = raw_index_info[raw_index_info['parameters'].str.contains(parameter)]
print('There are a total of %s data sources reporting %s parameter in the %s directory (%s)'%(len(parameter_subset['code'].unique().tolist()), parameter, index_file.split('.')[0].split('_')[1], product_name))
# <h2>PARAMETER'S DATA SOURCES BREAKDOWN</h2>
#create empty dictionary for available platform_types
data = {}
for data_source in parameter_subset['data_source'].unique().tolist(): data[data_source] = 0
print(data)
#count unique platform_code-plaform_types combos (same platform with code XXXX can works as several platform type: i.e vessels with TS (thermosalinometer) and CT (CTD))
combo = parameter_subset['code']+'_'+parameter_subset['data_source']
for combo in combo.unique():
data[combo.split('_')[1]] = data[combo.split('_')[1]] + 1
print(data)
parameter_data_source_breakdown = pd.DataFrame(data=data.values(), index = data.keys(), columns = ['number'])
parameter_data_source_breakdown
parameter_data_source_breakdown.plot.pie(figsize=(10,10), fontsize=15, subplots=True,autopct='%1.0f%%', colors = sns.color_palette())
parameter_data_source_breakdown.plot.bar(figsize=(10,10), fontsize=15, color = sns.color_palette())
# <h2>PARAMETER'S DATA SOURCES & DATA TYPES BREAKDOWN</h2>
#create empty dictionary for available data_types
data = {}
for dtype in parameter_subset['data'].unique().tolist(): data[dtype] = 0
print(data)
#count unique platform_code-plaform's data sources combos (same platform with code XXXX can works as several data sources: i.e vessels with TS (thermosalinometer) and CT (CTD))
for dtype in parameter_subset['data'].unique().tolist():
subset = parameter_subset[parameter_subset['data']== dtype]
temp = {}
for data_source in subset['data_source'].unique().tolist():
temp[data_source] = len(subset[subset['data_source']==data_source])
data[dtype] = temp
print(data)
parameter_dsource_datat_breakdown = pd.DataFrame(data=data)
parameter_dsource_datat_breakdown
parameter_dsource_datat_breakdown.plot.bar(figsize=(10,10), fontsize=15, color = sns.color_palette())
| PythonNotebooks/In_Situ_parameters_breakdown.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Nearest Points Exercise
#
# <NAME> (<EMAIL>)
#
# The latest version of this notebook is available at [https://github.com/Asterics2020-Obelics](https://github.com/Asterics2020-Obelics/School2019/tree/master/numpy)
# + slideshow={"slide_type": "slide"}
import numpy as np
import sys
print("Python version: {0}\n"
"NumPy version: {1}"
.format(sys.version, np.__version__))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Given an array of points (in 3D), find the nearest point for each one.
# + slideshow={"slide_type": "fragment"}
N = 500
n_dims = 3
points = np.random.random((N, n_dims))
points
# + [markdown] slideshow={"slide_type": "fragment"}
# Depending on the data size, our implementation may be faster!
| numpy/3. Nearest Points Exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] nbgrader={}
# # Numpy Exercise 1
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
import numpy as np
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
# + nbgrader={}
import antipackage
import github.ellisonbg.misc.vizarray as va
# + [markdown] nbgrader={}
# ## Checkerboard
# + [markdown] nbgrader={}
# Write a Python function that creates a square `(size,size)` 2d Numpy array with the values `0.0` and `1.0`:
#
# * Your function should work for both odd and even `size`.
# * The `0,0` element should be `1.0`.
# * The `dtype` should be `float`.
# + nbgrader={"checksum": "752eb79a174f018a88efdcf68249ecf6", "solution": true}
def checkerboard(size):
"""Return a 2d checkboard of 0.0 and 1.0 as a NumPy array"""
a = np.zeros((size,size))
x=0
while x < size:
a[x,0] = 1
x += 2
for x in range(0,size):
for y in range(1,size):
a[x,y]= (-1)**(x+y) + a[x,y-1]
return a
# -
checkerboard(4)
# + deletable=false nbgrader={"checksum": "a9259a1539798dd06c53d4699e5b89b7", "grade": true, "grade_id": "numpyex01a", "points": 6}
a = checkerboard(4)
assert a[0,0]==1.0
assert a.sum()==8.0
assert a.dtype==np.dtype(float)
assert np.all(a[0,0:5:2]==1.0)
assert np.all(a[1,0:5:2]==0.0)
b = checkerboard(5)
assert b[0,0]==1.0
assert b.sum()==13.0
assert np.all(b.ravel()[0:26:2]==1.0)
assert np.all(b.ravel()[1:25:2]==0.0)
# + [markdown] nbgrader={}
# Use `vizarray` to visualize a checkerboard of `size=20` with a block size of `10px`.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
# YOUR CODE HERE
va.enable()
va.set_block_size(10)
checkerboard(20)
# + deletable=false nbgrader={"checksum": "e2b72320a52fdce32f4c88bdaa78a92a", "grade": true, "grade_id": "numpyex01b", "points": 2}
assert True
# + [markdown] nbgrader={}
# Use `vizarray` to visualize a checkerboard of `size=27` with a block size of `5px`.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
# YOUR CODE HERE
va.set_block_size(5)
checkerboard(27)
# + deletable=false nbgrader={"checksum": "1f1c7ef9e0b2d19d4290ed461d43ff2c", "grade": true, "grade_id": "numpyex01c", "points": 2}
assert True
| assignments/assignment03/NumpyEx01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import os
import argparse
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torchvision.utils as utils
import pytorch_ssim
import time
from torch.autograd import Variable
from torch.utils.data import DataLoader
from torch.nn.modules.loss import _Loss
from Trans_unet_Gan import *
#from dataset import prepare_data, Dataset
from utils import *
import cv2
import matplotlib.pyplot as plt
from utility import plots as plots, ptcolor as ptcolor, ptutils as ptutils, data as data
from LAB import *
from LCH import *
from torchvision.utils import save_image
# -
os.environ["KMP_DUPLICATE_LIB_OK"]="TRUE"
def split(img):
output=[]
output.append(F.interpolate(img, scale_factor=0.125))
output.append(F.interpolate(img, scale_factor=0.25))
output.append(F.interpolate(img, scale_factor=0.5))
output.append(img)
return output
dtype = 'float32'
os.environ["CUDA_VISIBLE_DEVICES"] = '0'
torch.set_default_tensor_type(torch.FloatTensor)
# Initialize generator
generator = Generator().cuda()
generator.load_state_dict(torch.load("./saved_models/G/generator_795.pth"))
generator.eval()
path='./TestU_90/input/'#要改
path_list = os.listdir(path)
path_list.sort(key=lambda x:int(x.split('.')[0]))
i=1
for item in path_list:
impath=path+item
imgx= cv2.imread(path+item)
imgx=cv2.resize(imgx,(256,256))
imgx = cv2.cvtColor(imgx, cv2.COLOR_BGR2RGB)
imgx = np.array(imgx).astype(dtype)
imgx= torch.from_numpy(imgx)
imgx=imgx.permute(2,0,1).unsqueeze(0)
imgx=imgx/255.0
#plt.imshow(imgx[0,:,:,:])
#plt.show()
imgx = Variable(imgx).cuda()
#print(imgx.shape)
output=generator(imgx)
out=output[3].data
save_image(out, "./output_U90/%d.jpg" % (i), nrow=5, normalize=True)
i=i+1
| .ipynb_checkpoints/test90-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
#
BDA=VINODH
BDA
BDA="VINODH"
BDA
type(BDA)
VINODH=20
type(VINODH)
VIGNESH=80
VINODH-VIGNESH
VINODH+VIGNESH
VIGNESH*VINODH
RAJINI=2.0
type(RAJINI)
SMALLRAJINI=3.0
SMALLRAJINI-RAJINI
BDA[3]
BDA[:3]
BDA[0:4]
BDA[0:2]
sunday="businessdataanalytics"
sunday
sunday[::-1]
sunday[0:22
]
sunday[::-3]
sunday[::]
sunday[::2]
sunday[5]
week=['sun','mon','tue','wed','thu','fri','sat',1,2,3,4,5,6,7,['leave','work','work','leave'],True]
week
week.count('mon')
week.insert(1,5)
week
week.pop()
week
anydicts={"NABH":"national accrediation board of hospital","ASCII":"american standard code for information interchange","IDE":"integrated development environment","PPO":"pre placement offer"}
type(anydicts)
anydicts.values()
anydicts.keys()
anydicts.get("NABH")
anydicts.update({"PPO":"pension payment order"})
anydicts.get("PPO")
anyTuple=("meals","briyani","anwar",12,False,56,9)
anyTuple.count("anwar")
anyTuple.index(12)
anySet={1,8,8,2,3,4,"python",False}
anySet
anySet.add(75)
anySet
True
2<5
2>7
2>0
9+10>100
10==4
10==11-1
1!=2
3<=9
40*2<=800
# # two operands and one operator is statement
#
(40>100) or (50>25)
(40>50)and(100>50)
# # if,elif,else
#
if 40>30:
print("First statement was True")
elif 50>90:
print("Second statement was True")
else:
print("Nothing left hence printing else block")
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense, Conv2D, Dropout, Flatten, MaxPooling2D
import matplotlib.pyplot as plt
# %matplotlib inline
# +
# Downloading the dataset
(x_train, y_train), (x_test, y_test) = tf.keras.datasets.mnist.load_data()
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1)
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
input_shape = (28, 28, 1)
# Making sure that the values are float so that we can get decimal points after division
x_train = x_train.astype('float32')
x_test = x_test.astype('float32')
# Normalizing the RGB codes by dividing it to the max RGB value.
x_train /= 255
x_test /= 255
print('x_train shape:', x_train.shape)
print('Number of images in x_train', x_train.shape[0])
print('Number of images in x_test', x_test.shape[0])
# -
# Creating a Sequential Model and adding the layers
model = Sequential()
model.add(Conv2D(28, kernel_size=(3,3), input_shape=input_shape))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Flatten()) # Flattening the 2D arrays for fully connected layers
model.add(Dense(128, activation=tf.nn.relu))
model.add(Dropout(0.2))
model.add(Dense(10,activation=tf.nn.softmax))
# Compiling and training the model
model.compile(optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
model.fit(x=x_train,y=y_train, epochs=10)
# Evaluate the performance of the model on the test set
model.evaluate(x_test, y_test)
# Save the evaluation model
model.save('evaluation_model')
| evaluation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="agVkpKy_suJb"
# Lambda School Data Science
#
# *Unit 2, Sprint 2, Module 2*
#
# ---
# + id="3FIszjo3suJf"
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/main/data/'
# !pip install category_encoders==2.*
# !pip install pandas-profiling==2.*
# If you're working locally:
else:
DATA_PATH = '../data/'
# + [markdown] id="WCFSl3fDsuJg"
# # Module Project: Random Forests
# + [markdown] id="MVUliXejsuJk"
# This week, the module projects will focus on creating and improving a model for the Tanazania Water Pump datset. Your goal is to create a model to predict whether a water pump is functional, non-functional, or needs repair.
#
# Dataset source: [DrivenData.org](https://www.drivendata.org/competitions/7/pump-it-up-data-mining-the-water-table/).
#
# ## Directions
#
# The tasks for this project are as follows:
#
# - **Task 1:** Sign up for a [Kaggle](https://www.kaggle.com/) account.
# - **Task 2:** Use `wrangle` function to import training and test data.
# - **Task 3:** Split training data into feature matrix `X` and target vector `y`.
# - **Task 4:** Split feature matrix `X` and target vector `y` into training and test sets.
# - **Task 5:** Establish the baseline accuracy score for your dataset.
# - **Task 6:** Build and train `model_dt`.
# - **Task 7:** Calculate the training and validation accuracy score for your model.
# - **Task 8:** Adjust model's `max_depth` to reduce overfitting.
# - **Task 9 `stretch goal`:** Create a horizontal bar chart showing the 10 most important features for your model.
#
# You should limit yourself to the following libraries for this project:
#
# - `category_encoders`
# - `matplotlib`
# - `pandas`
# - `pandas-profiling`
# - `sklearn`
#
# # I. Wrangle Data
# + id="niGk2U8Avfnz"
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
#sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# Split train into train & val
# + id="9MjrrLfRsuJm"
def wrangle(X):
"""Wrangle train, validate, and test sets in the same way"""
# Prevent SettingWithCopyWarning
X = X.copy()
# About 3% of the time, latitude has small values near zero,
# outside Tanzania, so we'll treat these values like zero.
X['latitude'] = X['latitude'].replace(-2e-08, 0)
# When columns have zeros and shouldn't, they are like null values.
# So we will replace the zeros with nulls, and impute missing values later.
# Also create a "missing indicator" column, because the fact that
# values are missing may be a predictive signal.
cols_with_zeros = ['longitude', 'latitude', 'construction_year',
'gps_height', 'population']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
X[col+'_MISSING'] = X[col].isnull()
# Drop duplicate columns
duplicates = ['quantity_group', 'payment_type']
X = X.drop(columns=duplicates)
# Drop recorded_by (never varies) and id (always varies, random)
unusable_variance = ['recorded_by', 'id']
X = X.drop(columns=unusable_variance)
# Convert date_recorded to datetime
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.day
X = X.drop(columns='date_recorded')
# Engineer feature: how many years from construction_year to date_recorded
X['years'] = X['year_recorded'] - X['construction_year']
X['years_MISSING'] = X['years'].isnull()
# return the wrangled dataframe
return X
# + [markdown] id="-ZJDL585suJm"
# **Task 1:** Sign up for a [Kaggle](https://www.kaggle.com/) account. Choose a username that's based on your real name. Like GitHub, Kaggle is part of your public profile as a data scientist.
#
# **Task 2:** Modify the `wrangle` function to engineer a `'pump_age'` feature. Then use the function to read `train_features.csv` and `train_labels.csv` into the DataFrame `df`, and `test_features.csv` into the DataFrame `X_test`.
# + id="edhZDYJmvQoH"
# + id="PiMkZkS2suJo"
df = wrangle(train)
X_test = wrangle(test)
# + [markdown] id="y5-w4xn5suJq"
# # II. Split Data
#
# **Task 3:** Split your DataFrame `df` into a feature matrix `X` and the target vector `y`. You want to predict `'status_group'`.
# + id="LiKERuTnsuJs"
target = 'status_group'
y = df[target]
X = df.drop(columns=target)
# + [markdown] id="Apc1Xe5isuJu"
# **Task 4:** Using a randomized split, divide `X` and `y` into a training set (`X_train`, `y_train`) and a validation set (`X_val`, `y_val`).
# + id="w5Glf3rVsuJu"
from sklearn.model_selection import train_test_split
X_train, X_val, y_train, y_val = train_test_split(X, y, train_size=0.80, random_state=42)
# + [markdown] id="HuOAjaQlsuJv"
# # III. Establish Baseline
#
# **Task 5:** Since this is a **classification** problem, you should establish a baseline accuracy score. Figure out what is the majority class in `y_train` and what percentage of your training observations it represents.
# + colab={"base_uri": "https://localhost:8080/"} id="cMtDRR1NsuJv" outputId="22299c0a-eae7-468a-845d-06729ad6eb65"
baseline_acc = y_train.value_counts(normalize=True)
print('Baseline Accuracy Score:', baseline_acc)
# + colab={"base_uri": "https://localhost:8080/", "height": 298} id="u2JSTvPrpz0A" outputId="c4056e93-b816-42e2-9744-8269d449555a"
# %matplotlib
import matplotlib.pyplot as plt
y_train.value_counts(normalize=True).plot(kind = 'barh')
# + [markdown] id="Bn7k5gDKsuJw"
# # IV. Build Model
#
# **Task 6:** Build a `Pipeline` named `model_rf`, and fit it to your training data. Your `Pipeline` should include:
#
# - an `OrdinalEncoder` transformer for categorical features.
# - a `SimpleImputer` transformer fot missing values.
# - a `RandomForestClassifier` predictor.
#
# **Note:** Don't forget to set the `random_state` parameter for your `RandomForestClassifier`. Also, to decrease training time, set `n_jobs` to `-1`.
# + id="KBKjUhA7suJw"
from sklearn.pipeline import make_pipeline
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.ensemble import RandomForestClassifier
model_rf = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
RandomForestClassifier(n_estimators = 120, max_depth= 20, random_state = 0, n_jobs = -1)
)
model_rf.fit(X_train, y_train);
# + [markdown] id="KvvQtkqPsuJx"
# # V. Check Metrics
#
# **Task 7:** Calculate the training and validation accuracy scores for `model_rf`.
# + colab={"base_uri": "https://localhost:8080/"} id="iMj7uhJesuJx" outputId="2736bdbc-e708-4a38-82e5-e045dfb9bad8"
training_acc = model_rf.score(X_train, y_train)
val_acc = model_rf.score(X_val, y_val)
print('Training Accuracy Score:', training_acc)
print('Validation Accuracy Score:', val_acc)
# + [markdown] id="Inf6vlfesuJy"
# # VI. Tune Model
#
# **Task 8:** Tune `n_estimators` and `max_depth` hyperparameters for your `RandomForestClassifier` to get the best validation accuracy score for `model_rf`.
# + colab={"base_uri": "https://localhost:8080/"} id="VP-6jW1usuJ2" outputId="eabd610b-7b67-4725-ee50-c5abdb3122c3"
# Use this cell to experiment and then change
# your model hyperparameters in Task 6
model_rf2 = make_pipeline(
ce.OrdinalEncoder(),
SimpleImputer(),
RandomForestClassifier(n_estimators = 120, max_depth= 20, random_state = 0, n_jobs = -1)
)
model_rf2.fit(X_train, y_train);
training_acc2 = model_rf2.score(X_train, y_train)
val_acc2 = model_rf2.score(X_val, y_val)
print('Training Accuracy Score:', training_acc)
print('Validation Accuracy Score:', val_acc2)
# + colab={"base_uri": "https://localhost:8080/"} id="qqnwCF-U0KxZ" outputId="8c6c4fc7-8cf8-4103-f2a0-476141286312"
print('X_train shape before encoding:', X_train.shape)
encoder = model_rf.named_steps['ordinalencoder']
X_encoded = encoder.transform(X_train)
print('X_train shape after encoding:', X_encoded.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="w7pa0sSP0b0-" outputId="19d8f2d2-040b-4197-dec4-56e2fd49d34b"
print('X_train shape before encoding:', X_test.shape)
encoder = model_rf.named_steps['ordinalencoder']
X_encoded = encoder.transform(X_test)
print('X_train shape after encoding:', X_encoded.shape)
# + [markdown] id="kL8xaCTXsuJ4"
# # VII. Communicate Results
#
# **Task 9:** Generate a list of predictions for `X_test`. The list should be named `y_pred`.
# + colab={"base_uri": "https://localhost:8080/"} id="QOo_ngugsuJ4" outputId="6434cd6a-4e6f-43e2-e767-03a99f1e28e4"
y_pred = []
y_pred = model_rf.predict(X_test)
assert len(y_pred) == len(X_test), f'Your list of predictions should have {len(X_test)} items in it. '
y_pred.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 388} id="DpN84fivz6W_" outputId="d7e65ebe-b35f-4bef-d48c-11e2b0f64f0b"
X_test.head()
# + [markdown] id="dRbVKyHnsuJ4"
# **Task 11 `stretch goal`:** Create a DataFrame `submission` whose index is the same as `X_test` and that has one column `'status_group'` with your predictions. Next, save this DataFrame as a CSV file and upload your submissions to our competition site.
#
# **Note:** Check the `sample_submission.csv` file on the competition website to make sure your submissions follows the same formatting.
# + colab={"base_uri": "https://localhost:8080/", "height": 202} id="XY1UTFhJsuJ5" outputId="30fc7f85-21d3-4f91-a157-fb0577106bf0"
submission = pd.DataFrame(y_pred, columns=['status_group'], index = X_test.index)
submission.head()
# + id="tLg67K376XeN"
#download submission
submission.to_csv('odu_nkiru_water_pump_rf_sub.csv')
| module2-random-forests/LS_DS_222_assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.4.0-dev
# language: julia
# name: julia-0.4
# ---
# # quant-econ Solutions: Estimation of Spectra
#
# Solutions for http://quant-econ.net/jl/estspec.html
# +
using QuantEcon
using PyPlot
srand(42) # reproducible results
# -
# ## Exercise 1
# +
n = 400
phi = 0.5
theta = [0, -0.8]
lp = ARMA(phi, theta)
X = simulation(lp, ts_length=n)
fig, ax = subplots(3, 1)
for (i, wl) in enumerate([15, 55, 175]) # window lengths
x, y = periodogram(X)
ax[i][:plot](x, y, "b-", lw=2, alpha=0.5, label="periodogram")
x_sd, y_sd = spectral_density(lp, two_pi=false, res=120)
ax[i][:plot](x_sd, y_sd, "r-", lw=2, alpha=0.8, label="spectral density")
x, y_smoothed = periodogram(X, "hamming", wl)
ax[i][:plot](x, y_smoothed, "k-", lw=2, label="smoothed periodogram")
ax[i][:legend]()
ax[i][:set_title]("window length = $wl")
end
# -
# ## Exercise 2
| solutions/estspec_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Analysis of French ACC/UAC control sectors
# #### Analysis of Air Traffic Control en-route sectors of Metropolitan France
# ##### *Refer to tables_building.ipynb to generate sectors and volumes files (sectors.csv and volumes.geojson)*
# #### <br> Load tables
import pandas as pd
df = pd.read_csv('sectors.csv')
df.sample(4)
import geopandas as gpd
import numpy as np
gdf = gpd.read_file('volumes.geojson')
gdf.sample(4)
gdf.query('volume=="KD 3"').plot()
# #### <br> Number of Area Control Centers (ACC), sectors and airspace volumes
print('Number of ACCs over Metropolitan France: {}'.format(gdf['acc'].nunique()))
print('Number of control sectors over Metropolitan France: {}'.format(df['control_sector'].nunique()))
print('Number of elementary sectors over Metropolitan France: {}'.format(gdf['elementary_sector'].nunique()))
print('Number of airspace volumes over Metropolitan France: {}'.format(gdf['volume'].count()))
# #### <br> Number of volumes and sectors per ACC
s_cs = df['acc'].value_counts()
s_vol = gdf['acc'].value_counts()
s_es = gdf.groupby('acc')['elementary_sector'].nunique()
df_s = pd.concat([s_vol.rename('nb_volumes'), s_es.rename('nb_elementary_sectors'), s_cs.rename('nb_control_sectors')], axis=1)
df_s['total_sectors'] = df_s['nb_elementary_sectors'] + df_s['nb_control_sectors']
df_s.sort_values(['total_sectors'])
# #### <br> Smallest and largest airspace volumes (horizontal surface) and elementary sectors
# ##### Largest and smallest volumes
gdf["surface"] = gdf['geometry'].area
gdf.sort_values(['surface'], ascending=False, inplace=True)
gdf.head(4)
gdf.tail(4)
# ##### Largest and smallest elementary sectors
gdf.groupby('elementary_sector').max().nlargest(4, 'surface')
gdf.groupby('elementary_sector').max().nsmallest(4, 'surface')
# ##### Display largest and smallest elementary sectors
from shapely.geometry import Polygon
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
france = world[world.name == "France"].copy()
france.geometry = france.geometry.intersection(Polygon([(-10,41),(-10,52),(10,52),(10,41)]))
ax = france.plot(color='white', edgecolor='black', linewidth=1.0)
smalls = list(gdf.groupby('elementary_sector').max().nsmallest(4, 'surface').index)
bigs = list(gdf.groupby('elementary_sector').max().nlargest(4, 'surface').index)
gdf.query('elementary_sector == @smalls').plot(ax=ax, color='green', alpha=0.6)
gdf.query('elementary_sector == @bigs').plot(ax=ax, color='orange', alpha=0.6)
# #### <br> Flight Levels
# ##### All flight levels used as level max in Metropolitan France
gdf.level_max.unique()
# ##### Flight levels used per ACC
for key, group in gdf.groupby('acc'):
print(key, ': ', sorted(group['level_max'].unique()))
# #### <br> Identify complex elementary sectors composed of most different volumes
gdf['elementary_sector'].value_counts().nlargest(3)
ax = france.plot(color='white', edgecolor='black', linewidth=1.0)
complex = list(gdf['elementary_sector'].value_counts().nlargest(3).index)
gdf.query('elementary_sector == @complex').plot(ax=ax, color='red', alpha=0.6)
# #### <br> Display airspace volumes, ACCs and sectors
gdf_low = gdf.query('level_min <= 200 <= level_max')
gdf_high = gdf.query('level_min <= 350 <= level_max')
print('{} volumes at FL200, {} volumes at FL350'.format(gdf_low['volume'].count(),
gdf_high['volume'].count()))
# ##### Display all volumes at FL200
from shapely.geometry import Polygon
world = gpd.read_file(gpd.datasets.get_path('naturalearth_lowres'))
france = world[world.name == "France"].copy()
france.geometry = france.geometry.intersection(Polygon([(-10,41),(-10,52),(10,52),(10,41)]))
ax = france.plot(color='white', edgecolor='black', linewidth=1.0)
custom_colors = [np.random.random(3) for i in range(gdf_low['volume'].count())]
gdf_low.plot(ax=ax, color=custom_colors, alpha=0.8)
# ##### Display all volumes at FL350
# %matplotlib inline
from matplotlib import pyplot as plt
ax = france.plot(color='white', edgecolor='black', linewidth=1.0)
custom_colors = [np.random.random(3) for i in range(gdf_high['volume'].count())]
gdf_high.plot(ax=ax, color=custom_colors, alpha=0.8)
plt.savefig('fl350_volumes.svg', format="svg")
# ##### Display the ACCs at level FL350
pd.set_option('mode.chained_assignment', None)
gdf_high['geometry'] = gdf_high.buffer(0.01)
ax = france.plot(color='white', edgecolor='black', linewidth=1.0)
custom_colors = [np.random.random(3) for i in range(5)]
gdf_high.dissolve(by='acc').plot(ax=ax, color=custom_colors, alpha=0.8)
# ##### Display elementary sectors' volumes of LFBB ACC at level FL190 (FIR Bordeaux)
gdf_vlow = gdf.query('level_min <= 190 <= level_max')
ax = france.plot(color='white', edgecolor='black', linewidth=1.0)
custom_colors = [np.random.random(3) for i in range(gdf_vlow['volume'].count())]
gdf_vlow.query('acc == "LFBB"').dissolve(by='elementary_sector').plot(ax=ax, color=custom_colors, alpha=0.8)
# #### <br> Sectors' centroids and cities
ax = france.plot(color='white', edgecolor='black', linewidth=1.0)
cities = gpd.read_file(gpd.datasets.get_path('naturalearth_cities'))
fr_cities = cities.cx[-10:10, 41:52]
gdf['geometry'].centroid.plot(ax=ax)
fr_cities.plot(ax=ax, color='red')
# ##### Test which volumes/sectors are above a given city (ex: Toulouse)
from shapely.geometry import Point
toulouse = Point(1.433, 43,60)
gdf['above_toulouse'] = gdf['geometry'].map(lambda x: True if x.contains(toulouse) else False)
gdf_a = gdf.query('above_toulouse == True')
gdf_a
ax = france.plot(color='white', edgecolor='black', linewidth=1.0)
custom_colors = [np.random.random(3) for i in range(gdf_a['volume'].count())]
gdf_a.query('above_toulouse == True').plot(ax=ax, color=custom_colors, alpha=0.3)
df_t = pd.DataFrame({'geometry':[toulouse]})
gdf_t = gpd.GeoDataFrame(df_t, geometry='geometry')
gdf_t.plot(ax=ax, color='red')
| atc_sectors_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Scraping with Pandas
# + slideshow={"slide_type": "slide"}
import pandas as pd
# + [markdown] slideshow={"slide_type": "subslide"}
# We can use the `read_html` function in Pandas to automatically scrape any tabular data from a page.
# + slideshow={"slide_type": "fragment"}
data_path = "Resources/cities.csv"
# + slideshow={"slide_type": "fragment"}
data_table_df = pd.read_csv("Resources/cities.csv")
data_table_df.head()
# -
data_table_df[["City", "Country","Date","Cloudiness","Humidity","Lat","Lng","Max Temp", "Wind Speed"]].head()
# + [markdown] slideshow={"slide_type": "slide"}
# ## DataFrames as HTML
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Pandas also had a `to_html` method that we can use to generate HTML tables from DataFrames.
# + slideshow={"slide_type": "fragment"}
data_table_html = data_table_df.to_html()
data_table_html
# + [markdown] slideshow={"slide_type": "subslide"}
# #### You may have to strip unwanted newlines to clean up the table.
# + slideshow={"slide_type": "fragment"}
data_table_html.replace('\n', '')
# + [markdown] slideshow={"slide_type": "subslide"}
# You can also save the table directly to a file.
# + slideshow={"slide_type": "fragment"}
data_table_df.to_html('data_table.html')
# + slideshow={"slide_type": "fragment"}
# OSX Users can run this to open the file in a browser,
# or you can manually find the file and open it in the browser
# !open table.html
# -
| datatohtml.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
# ### 2.3.4 共起行列
include("../common/util.jl")
text = "You say goodbye and I say hello."
corpus, word_to_id, id_to_word = preprocess(text)
println(corpus)
println(id_to_word)
C = [0 1 0 0 0 0 0;
1 0 1 0 1 1 0;
0 1 0 1 0 0 0;
0 0 1 0 1 0 0;
0 1 0 1 0 0 0;
0 1 0 0 0 0 1;
0 0 0 0 0 1 0]
println(C[1, :])
println(C[5, :])
println(C[word_to_id["goodbye"], :])
| ch02/ch02_2_3_4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div align="right"><a href="http://norvig.com"><NAME></a><br><a href="https://github.com/norvig/pytudes">pytudes</a><br>March 2019</div>
#
# # Dice Baseball
#
# The [538 Riddler for March 22, 2019](https://fivethirtyeight.com/features/can-you-turn-americas-pastime-into-a-game-of-yahtzee/) asks us to simulate baseball using probabilities from a 19th century dice game called *Our National Ball Game*:
#
# 1,1: double 2,2: strike 3,3: out at 1st 4,4: fly out
# 1,2: single 2,3: strike 3,4: out at 1st 4,5: fly out
# 1,3: single 2,4: strike 3,5: out at 1st 4,6: fly out
# 1,4: single 2,5: strike 3,6: out at 1st 5,5: double play
# 1,5: base on error 2,6: foul out 5,6: triple
# 1,6: base on balls 6,6: home run
#
#
# The rules left some things unspecified; the following are my current choices (in an early version I made different choices that resulted in slightly more runs):
#
# * On a* b*-base hit, runners advance* b* bases, except that a runner on second scores on a 1-base hit.
# * On an "out at first", all runners advance one base.
# * A double play only applies if there is a runner on first; in that case other runners advance.
# * On a fly out, a runner on third scores; other runners do not advance.
# * On an error all runners advance one base.
# * On a base on balls, only forced runners advance.
#
# I also made some choices about the implementation:
#
# - Exactly one outcome happens to each batter. We call that an *event*.
# - I'll represent events with the following one letter codes:
# - `K`, `O`, `o`, `f`, `D`: strikeout, foul out, out at first, fly out, double play
# - `1`, `2`, `3`, `4`: single, double, triple, home run
# - `E`, `B`: error, base on balls
# - Note the "strike" dice roll is not an event; it is only part of an event. From the probability of a "strike" dice roll, I compute the probability of three strikes in a row, and call that a strikeout event. Sice there are 7 dice rolls giving "strike", the probability of a strike is 7/36, and the probability of a strikeout is (7/36)**3.
# - Note that a die roll such as `1,1` is a 1/36 event, whereas `1,2` is a 2/36 event, because it also represents (2, 1).
# - I'll keep track of runners with a list of occupied bases; `runners = [1, 2]` means runners on first and second.
# - A runner who advances to base 4 or higher has scored a run (unless there are already 3 outs).
# - The function `inning` simulates a half inning and returns the number of runs scored.
# - I want to be able to test `inning` by feeding it specific events, and I also want to generate random innings. So I'll make the interface be that I pass in an *iterable* of events. The function `event_stream` generates an endless stream of randomly sampled events.
# - Note that it is consider good Pythonic style to automatically convert Booleans to integers, so for a runner on second (`r = 2`) when the event is a single (`e = '1'`), the expression `r + int(e) + (r == 2)` evaluates to `2 + 1 + 1` or `4`, meaning the runner on second scores.
# - I'll play 1 million innings and store the resulting scores in `innings`.
# - To simulate a game I just sample 9 elements of `innings` and sum them.
#
# # The Code
# %matplotlib inline
import matplotlib.pyplot as plt
import random
# +
def event_stream(events='2111111EEBBOOooooooofffffD334', strike=7/36):
"An iterator of random events. Defaults from `Our National Ball Game`."
while True:
yield 'K' if (random.random() < strike ** 3) else random.choice(events)
def inning(events=event_stream(), verbose=False) -> int:
"Simulate a half inning based on events, and return number of runs scored."
outs = runs = 0 # Inning starts with no outs and no runs,
runners = [] # ... and with nobody on base
for e in events:
if verbose: print(f'{outs} outs, {runs} runs, event: {e}, runners: {runners}')
# What happens to the batter?
if e in 'KOofD': outs += 1 # Batter is out
elif e in '1234EB': runners.append(0) # Batter becomes a runner
# What happens to the runners?
if e == 'D' and 1 in runners: # double play: runner on 1st out, others advance
outs += 1
runners = [r + 1 for r in runners if r != 1]
elif e in 'oE': # out at first or error: runners advance
runners = [r + 1 for r in runners]
elif e == 'f' and 3 in runners and outs < 3: # fly out: runner on 3rd scores
runners.remove(3)
runs += 1
elif e in '1234': # single, double, triple, homer
runners = [r + int(e) + (r == 2) for r in runners]
elif e == 'B': # base on balls: forced runners advance
runners = [r + forced(runners, r) for r in runners]
# See if inning is over, and if not, whether anyone scored
if outs >= 3:
return runs
runs += sum(r >= 4 for r in runners)
runners = [r for r in runners if r < 4]
def forced(runners, r) -> bool: return all(b in runners for b in range(r))
# -
# # Testing
#
# Let's peek at some random innings:
inning(verbose=True)
inning(verbose=True)
# And we can feed in any events we want to test the code:
inning('2EBB1DB12f', verbose=True)
# That looks good.
#
# # Simulating
#
# Now, simulate a million innings, and then sample from them to simulate a million nine-inning games (for one team):
N = 1000000
innings = [inning() for _ in range(N)]
games = [sum(random.sample(innings, 9)) for _ in range(N)]
# Let's see histograms:
# +
def hist(nums, title):
"Plot a histogram."
plt.hist(nums, ec='black', bins=max(nums)-min(nums)+1, align='left')
plt.title(f'{title} Mean: {sum(nums)/len(nums):.3f}, Min: {min(nums)}, Max: {max(nums)}')
hist(innings, 'Runs per inning:')
# -
hist(games, 'Runs per game:')
| ipynb/Dice Baseball.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Analysis with Python
#
#
# ---
# + [markdown] tags=["solution"]
# ## Instructor notes
#
# *Estimated teaching time:* 30 min
#
# *Estimated challenge time:* 30 min
#
# *Key questions:*
#
# - "How can I import data in Python ?"
# - "What is Pandas ?"
# - "Why should I use Pandas to work with data ?"
#
# *Learning objectives:*
#
# - "Navigate the workshop directory and download a dataset."
# - "Explain what a library is and what libraries are used for."
# - "Describe what the Python Data Analysis Library (Pandas) is."
# - "Load the Python Data Analysis Library (Pandas)."
# - "Use `read_csv` to read tabular data into Python."
# - "Describe what a DataFrame is in Python."
# - "Access and summarize data stored in a DataFrame."
# - "Define indexing as it relates to data structures."
# - "Perform basic mathematical operations and summary statistics on data in a Pandas DataFrame."
# - "Create simple plots."
# -
# ## Automating data analysis tasks in Python
#
# We can automate the process of performing data manipulations in Python. It's efficient to spend time
# building the code to perform these tasks because once it's built, we can use it
# over and over on different datasets that use a similar format. This makes our
# methods easily reproducible. We can also easily share our code with colleagues
# and they can replicate the same analysis.
#
# ### The Dataset
#
# For this lesson, we will be using the Portal Teaching data, a subset of the data
# from Ernst et al
# [Long-term monitoring and experimental manipulation of a Chihuahuan Desert ecosystem near Portal, Arizona, USA](http://www.esapubs.org/archive/ecol/E090/118/default.htm)
#
# We will be using this dataset, which can be downloaded here: [surveys.csv](data/surveys.csv) ... but **don't click** to download it in your browser - **we are going to use Python !**
import urllib.request
# You can also get this URL value by right-clicking the `surveys.csv` link above and selecting "Copy Link Address"
url = 'https://monashdatafluency.github.io/python-workshop-base/modules/data/surveys.csv'
# url = 'https://goo.gl/9ZxqBg' # or a shortened version to save typing
urllib.request.urlretrieve(url, 'surveys.csv')
# If Jupyter is running locally on your computer, you'll now have a file `surveys.csv` in the current working directory.
# You can check by clicking on `File` tab on the top left of the notebook to see if the file exists. If you are running Jupyter on a remote server or cloud service (eg Colaboratory or Azure Notebooks), the file will be there instead.
# We are studying the species and weight of animals caught in plots in our study
# area. The dataset is stored as a `.csv` file: each row holds information for a
# single animal, and the columns represent:
#
# | Column | Description |
# |------------------|------------------------------------|
# | record_id | Unique id for the observation |
# | month | month of observation |
# | day | day of observation |
# | year | year of observation |
# | site_id | ID of a particular plot |
# | species_id | 2-letter code |
# | sex | sex of animal ("M", "F") |
# | hindfoot_length | length of the hindfoot in mm |
# | weight | weight of the animal in grams |
#
#
# The first few rows of our file look like this:
#
# ```
# record_id,month,day,year,site_id,species_id,sex,hindfoot_length,weight
# 1,7,16,1977,2,NL,M,32,
# 2,7,16,1977,3,NL,M,33,
# 3,7,16,1977,2,DM,F,37,
# 4,7,16,1977,7,DM,M,36,
# 5,7,16,1977,3,DM,M,35,
# 6,7,16,1977,1,PF,M,14,
# 7,7,16,1977,2,PE,F,,
# 8,7,16,1977,1,DM,M,37,
# 9,7,16,1977,1,DM,F,34,
# ```
#
# ---
# ## About Libraries
#
# A library in Python contains a set of tools (called functions) that perform
# tasks on our data. Importing a library is like getting a piece of lab equipment
# out of a storage locker and setting it up on the bench for use in a project.
# Once a library is set up, it can be used or called to perform many tasks.
#
# If you have noticed in the previous code `import urllib.request`, we are calling
# a **request** function from library **urllib** to download our dataset from web.
#
#
# ## Pandas in Python
# The dataset we have, is in table format. One of the best options for working with tabular data in Python is to use the
# [Python Data Analysis Library](http://pandas.pydata.org/) (a.k.a. Pandas). The
# Pandas library provides data structures, produces high quality plots with
# [matplotlib](http://matplotlib.org/) and integrates nicely with other libraries
# that use [NumPy](http://www.numpy.org/) (which is another Python library) arrays.
#
# First, lets make sure the Pandas and matplotlib packages are **installed**.
# !pip install pandas matplotlib
# Python doesn't load all of the libraries available to it by default. We have to
# add an `import` statement to our code in order to use library functions. To import
# a library, we use the syntax `import libraryName`. If we want to give the
# library a nickname to shorten the command, we can add `as nickNameHere`. An
# example of importing the pandas library using the common nickname `pd` is below.
import pandas as pd
#
# Each time we call a function that's in a library, we use the syntax
# `LibraryName.FunctionName`. Adding the library name with a `.` before the
# function name tells Python where to find the function. In the example above, we
# have imported Pandas as `pd`. This means we don't have to type out `pandas` each
# time we call a Pandas function.
#
#
# # Reading CSV Data Using Pandas
#
# We will begin by locating and reading our survey data which are in CSV format. CSV stands for Comma-Separated Values and is a common way store formatted data. Other symbols my also be used, so you might see tab-separated, colon-separated or space separated files. It is quite easy to replace one separator with another, to match your application. The first line in the file often has headers to explain what is in each column. CSV (and other separators) make it easy to share data, and can be imported and exported from many applications, including Microsoft Excel.
#
# We can use Pandas' `read_csv` function to pull the file directly into a
# [DataFrame](http://pandas.pydata.org/pandas-docs/stable/dsintro.html#dataframe).
#
#
# ## So What's a DataFrame?
#
# A DataFrame is a 2-dimensional data structure that can store data of different
# types (including characters, integers, floating point values, factors and more)
# in columns. It is similar to a spreadsheet or an SQL table or the `data.frame` in
# R. A DataFrame always has an index (0-based). An index refers to the position of
# an element in the data structure.
#
#
# Note that pd.read_csv is used because we imported pandas as pd
pd.read_csv("surveys.csv")
# The above command outputs a `DateFrame` object, which Jupyter displays as a table (snipped in the middle since there are many rows).
#
# We can see that there were 33,549 rows parsed. Each row has 9
# columns. The first column is the index of the DataFrame. The index is used to
# identify the position of the data, but it is not an actual column of the DataFrame.
# It looks like the `read_csv` function in Pandas read our file properly. However,
# we haven't saved any data to memory so we can work with it.We need to assign the
# DataFrame to a variable. Remember that a variable is a name for a value, such as `x`,
# or `data`. We can create a new object with a variable name by assigning a value to it using `=`.
#
# Let's call the imported survey data `surveys_df`:
surveys_df = pd.read_csv("surveys.csv")
# Notice when you assign the imported DataFrame to a variable, Python does not
# produce any output on the screen. We can view the value of the `surveys_df`
# object by typing its name into the cell.
surveys_df
#
# which prints contents like above.
# You can also select just a few rows, so it is easier to fit on one window, you can see that pandas has neatly formatted the data to fit our screen.
#
# Here, we will be using a function called **head**.
#
# The `head()` function displays the first several lines of a file. It is discussed below.
#
surveys_df.head()
# ## Exploring Our Species Survey Data
#
# Again, we can use the `type` function to see what kind of thing `surveys_df` is:
#
#
type(surveys_df)
#
# As expected, it's a DataFrame (or, to use the full name that Python uses to refer
# to it internally, a `pandas.core.frame.DataFrame`).
#
# What kind of things does `surveys_df` contain? DataFrames have an attribute
# called `dtypes` that answers this:
#
#
surveys_df.dtypes
# All the values in a single column have the same type. For example, months have type
# `int64`, which is a kind of integer. Cells in the month column cannot have
# fractional values, but the weight and hindfoot_length columns can, because they
# have type `float64`. The `object` type doesn't have a very helpful name, but in
# this case it represents strings (such as 'M' and 'F' in the case of sex).
#
# ### Useful Ways to View DataFrame objects in Python
#
# There are many ways to summarize and access the data stored in DataFrames,
# using attributes and methods provided by the DataFrame object.
#
# To access an attribute, use the DataFrame object name followed by the attribute
# name `df_object.attribute`. Using the DataFrame `surveys_df` and attribute
# `columns`, an index of all the column names in the DataFrame can be accessed
# with `surveys_df.columns`.
#
# Methods are called in a similar fashion using the syntax `df_object.method()`.
# As an example, `surveys_df.head()` gets the first few rows in the DataFrame
# `surveys_df` using **the `head()` method**. With a method, we can supply extra
# information in the parens to control behaviour.
#
# Let's look at the data using these.
# + [markdown] tags=["challenge"]
# ## Challenge - DataFrames
#
# Using our DataFrame `surveys_df`, try out the attributes & methods below to see
# what they return.
#
# 1. `surveys_df.columns`
# 2. `surveys_df.shape` Take note of the output of `shape` - what format does it
# return the shape of the DataFrame in? HINT: [More on tuples, here](https://docs.python.org/3/tutorial/datastructures.html#tuples-and-sequences).
# 3. `surveys_df.head()` Also, what does `surveys_df.head(15)` do?
# 4. `surveys_df.tail()`
#
#
#
# + [markdown] tags=["solution"]
# ## Solution - DataFrames
#
# ... try it yourself !
# -
# # Calculating Statistics From Data
#
# We've read our data into Python. Next, let's perform some quick summary
# statistics to learn more about the data that we're working with. We might want
# to know how many animals were collected in each plot, or how many of each
# species were caught. We can perform summary stats quickly using groups. But
# first we need to figure out what we want to group by.
#
# Let's begin by exploring our data:
#
#
# Look at the column names
surveys_df.columns
# Let's get a list of all the species. The `pd.unique` function tells us all of
# the unique values in the `species_id` column.
pd.unique(surveys_df['species_id'])
# + [markdown] tags=["challenge"]
# ## Challenge - Statistics
#
# 1. Create a list of unique site ID's found in the surveys data. Call it
# `site_names`. How many unique sites are there in the data? How many unique
# species are in the data?
#
# 2. What is the difference between `len(site_names)` and `surveys_df['site_id'].nunique()`?
# + [markdown] tags=["solution"]
# ## Solution - Statistics
# + tags=["solution"]
site_names = pd.unique(surveys_df['site_id'])
print(len(site_names), surveys_df['site_id'].nunique())
# -
# # Groups in Pandas
#
# We often want to calculate summary statistics grouped by subsets or attributes
# within fields of our data. For example, we might want to calculate the average
# weight of all individuals per site.
#
# We can calculate basic statistics for all records in a single column using the
# syntax below:
surveys_df['weight'].describe()
#
# We can also extract one specific metric if we wish:
#
#
surveys_df['weight'].min()
surveys_df['weight'].max()
surveys_df['weight'].mean()
surveys_df['weight'].std()
# only the last command shows output below - you can try the others above in new cells
surveys_df['weight'].count()
#
# But if we want to summarize by one or more variables, for example sex, we can
# use **Pandas' `.groupby` method**. Once we've created a groupby DataFrame, we
# can quickly calculate summary statistics by a group of our choice.
#
#
# Group data by sex
grouped_data = surveys_df.groupby('sex')
#
# The **pandas function `describe`** will return descriptive stats including: mean,
# median, max, min, std and count for a particular column in the data. **Note** Pandas'
# `describe` function will only return summary values for columns containing
# numeric data.
#
#
# +
# Summary statistics for all numeric columns by sex
grouped_data.describe()
# Provide the mean for each numeric column by sex
# As above, only the last command shows output below - you can try the others above in new cells
grouped_data.mean()
# -
#
# The `groupby` command is powerful in that it allows us to quickly generate
# summary stats.
#
#
# + [markdown] tags=["challenge"]
# ## Challenge - Summary Data
#
# 1. How many recorded individuals are female `F` and how many male `M`
# - A) 17348 and 15690
# - B) 14894 and 16476
# - C) 15303 and 16879
# - D) 15690 and 17348
#
#
# 2. What happens when you group by two columns using the following syntax and
# then grab mean values:
# - `grouped_data2 = surveys_df.groupby(['site_id','sex'])`
# - `grouped_data2.mean()`
#
#
# 3. Summarize weight values for each site in your data. HINT: you can use the
# following syntax to only create summary statistics for one column in your data
# `by_site['weight'].describe()`
#
#
# + [markdown] tags=["solution"]
# ## Solution- Summary Data
# + tags=["solution"]
## Solution Challenge 1
grouped_data.count()
# + [markdown] tags=["solution"]
# ### Solution - Challenge 2
#
# The mean value for each combination of site and sex is calculated. Remark that the
# mean does not make sense for each variable, so you can specify this column-wise:
# e.g. I want to know the last survey year, median foot-length and mean weight for each site/sex combination:
# + tags=["solution"]
# Solution- Challenge 3
surveys_df.groupby(['site_id'])['weight'].describe()
# + [markdown] tags=["solution"]
# ## Did you get #3 right?
# **A Snippet of the Output from part 3 of the challenge looks like:**
#
# ```
# site_id
# 1 count 1903.000000
# mean 51.822911
# std 38.176670
# min 4.000000
# 25% 30.000000
# 50% 44.000000
# 75% 53.000000
# max 231.000000
# ...
# ```
#
#
# -
# ## Quickly Creating Summary Counts in Pandas
#
# Let's next count the number of samples for each species. We can do this in a few
# ways, but we'll use `groupby` combined with **a `count()` method**.
#
#
#
# Count the number of samples by species
species_counts = surveys_df.groupby('species_id')['record_id'].count()
print(species_counts)
#
# Or, we can also count just the rows that have the species "DO":
#
#
surveys_df.groupby('species_id')['record_id'].count()['DO']
# ## Basic Math Functions
#
# If we wanted to, we could perform math on an entire column of our data. For
# example let's multiply all weight values by 2. A more practical use of this might
# be to normalize the data according to a mean, area, or some other value
# calculated from our data.
#
#
# Multiply all weight values by 2 but does not change the original weight data
surveys_df['weight']*2
# ## Quick & Easy Plotting Data Using Pandas
#
# We can plot our summary stats using Pandas, too.
#
#
# +
## To make sure figures appear inside Jupyter Notebook
# %matplotlib inline
# Create a quick bar chart
species_counts.plot(kind='bar')
# -
# #### Animals per site plot
#
# We can also look at how many animals were captured in each site.
total_count = surveys_df.groupby('site_id')['record_id'].nunique()
# Let's plot that too
total_count.plot(kind='bar')
# + [markdown] tags=["challenge"]
# ## _Extra Plotting Challenge_
#
# 1. Create a plot of average weight across all species per plot.
#
# 2. Create a plot of total males versus total females for the entire dataset.
#
# 3. Create a stacked bar plot, with weight on the Y axis, and the stacked variable being sex. The plot should show total weight by sex for each plot. Some tips are below to help you solve this challenge:
# [For more on Pandas plots, visit this link.](http://pandas.pydata.org/pandas-docs/stable/visualization.html#basic-plotting-plot)
#
#
#
#
# + [markdown] tags=["solution"]
# ### _Solution to Extra Plotting Challenge 1_
# + tags=["solution"]
## Solution Plotting Challenge 1
surveys_df.groupby('site_id').mean()["weight"].plot(kind='bar')
# + [markdown] tags=["solution"]
# ### _Solution to Extra Plotting Challenge 2_
# + tags=["solution"]
# Solution Plotting Challenge 2
## Create plot of total males versus total females for the entire dataset.
surveys_df.groupby('sex').count()["record_id"].plot(kind='bar')
# + [markdown] tags=["solution"]
# ### _Solution to Extra Plotting Challenge 3_
#
# First we group data by site and by sex, and then calculate a total for each site.
# + tags=["solution"]
by_site_sex = surveys_df.groupby(['site_id','sex'])
site_sex_count = by_site_sex['weight'].sum()
# + [markdown] tags=["solution"]
#
# This calculates the sums of weights for each sex within each plot as a table
#
# ```
# site sex
# site_id sex
# 1 F 38253
# M 59979
# 2 F 50144
# M 57250
# 3 F 27251
# M 28253
# 4 F 39796
# M 49377
# <other sites removed for brevity>
# ```
#
# Below we'll use `.unstack()` on our grouped data to figure out the total weight that each sex contributed to each plot.
#
#
# + tags=["solution"]
by_site_sex = surveys_df.groupby(['site_id','sex'])
site_sex_count = by_site_sex['weight'].sum()
site_sex_count.unstack()
# + [markdown] tags=["solution"]
# Now, create a stacked bar plot with that data where the weights for each sex are stacked by plot.
#
# Rather than display it as a table, we can plot the above data by stacking the values of each sex as follows:
# + tags=["solution"]
by_site_sex = surveys_df.groupby(['site_id', 'sex'])
site_sex_count = by_site_sex['weight'].sum()
spc = site_sex_count.unstack()
s_plot = spc.plot(kind='bar', stacked=True, title="Total weight by site and sex")
s_plot.set_ylabel("Weight")
s_plot.set_xlabel("Site")
# -
| workshops/docs/modules/notebooks/working_with_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.9 64-bit (''TrajDetec'': conda)'
# name: python_defaultSpec_1600737096552
# ---
# + tags=[]
import numpy as np
import pandas as pd
import tensorflow as tf
from tensorflow.keras import layers,models
tf.keras.backend.clear_session()
model = models.Sequential()
model.add(layers.Dense(32,input_shape = (None,16),activation = tf.nn.relu)) #通过activation参数指定
model.add(layers.Dense(10))
model.add(layers.Activation(tf.nn.softmax)) # 显式添加layers.Activation激活层
model.summary()
# -
| Computer Science/eat_tensorflow2_in_30_days/practice/5_mid level API/3_activation function/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from multiobject.pytorch import MultiObjectDataLoader, MultiObjectDataset
import numpy as np
import matplotlib.pyplot as plt
import torchvision
import os
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:80% !important; }</style>"))
# -
# # Utils
def visualize_dataset(path_to_dataset):
batch_size = 32
dataset_path = path_to_dataset
train_set = MultiObjectDataset(dataset_path, train=True, split = 1.)
print("Num of Images ", len(train_set))
train_loader = MultiObjectDataLoader(train_set, batch_size=batch_size, shuffle=True)
samples, _ = iter(train_loader).next()
fig=plt.figure(figsize=(12, 12))
N_images = 21
for i in range(1,N_images):
fig.add_subplot(int(np.sqrt(N_images)) +1 , int(np.sqrt(N_images))+1, i)
plt.imshow(np.transpose(samples[i], (1,2,0)))
plt.show()
return None
# # dSprites base
# +
#visualize_dataset('generated/dsprites/multi_dsprites_210407_144010.npz')
# -
# # Custom
# ## Test 1
#
# Add customization of :
# - output folder through --folder
# - file name through --file
# - number of images generated through -n
visualize_dataset('generated/custom/multi_dsprites_test_1.npz')
# # Test 2
#
# Add customization of :
# - patch size through --patch_size
# - frame size through --frame_size
# - allow overlap through --overlap
#
# Parameters : patch_size = 64, frame_size = 128
visualize_dataset('generated/custom/multi_dsprites_test_2.npz')
# ## Test 3
#
# Add customization of :
# - placement of objects through -p --placement ('random', 'center', 'xalign', 'yalign')
#
#
# Parameters : patch_size = 32, frame_size = 128, placement = yalign
visualize_dataset('generated/custom/multi_dsprites_test_3.npz')
# Parameters : patch_size = 32, frame_size = 128, placement = xalign
visualize_dataset('generated/custom/multi_dsprites_test_3_1.npz')
# !python generate_dataset.py --noise 0
data_path = 'generated/custom/multi_dsprites_test_4.npz'
data = np.load(data_path, allow_pickle=True)
x = np.asarray(data['x'], dtype=np.float32) / 255
labels = data['labels']
fig=plt.figure(figsize=(25, 25))
N_images = 24
axs = [None]*N_images
for i in range(1,N_images):
axs[i] = fig.add_subplot(int(np.sqrt(N_images)) +1 , int(np.sqrt(N_images))+1, i)
axs[i].set_title(labels[i][1], fontdict = {'fontsize': 13})
axs[i].set_xticks([])
axs[i].set_yticks([])
plt.imshow(x[i])
plt.show()
# +
#labels
# -
bool(0)
| Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# homework
# -
# # Learning Objectives
# - we learn what are Convolutional Neural Network (CNN)
# - Will talk about CNN components such as stride, max or average pooling
# - Discuss how we can obtain the parameters for CNN
#
# # Activity: Obtain the number of parameters for the following CNN
# ---
# - By default, the strides = (1, 1)
# +
from __future__ import print_function
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3),
activation='relu',
input_shape=(28, 28, 1)))
# it inherited from the previous step so we dont have to define '32'
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
# model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
# model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax'))
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
print(model.summary())
# -
| Notebooks/Convolutional Neural Network (CNN).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Activity 4.04: Visualizing the Top 30 Music YouTube Channels Using Seaborn's FacetGrid
# In this activity, we will visualize the total number of subscribers and the total number of views for the top 30 YouTube channels (as of January 2020) in the category music by using the FacetGrid() function that's provided by the Seaborn library. Visualize the given data using a FacetGrid with two columns. The first column should show the number of subscribers for each YouTube channel, whereas the second column should show the number of views. The goal of this activity is to get some practice working with FacetGrids. Following are the steps to implement this activity:
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# Use pandas to read the youtube.csv data located in the Dataset folder.
mydata = pd.read_csv("../../Datasets/YouTube.csv")
mydata.head()
# Access the data of each group in the column, convert them into a list, and assign this list to variables of each respective group.
channels = mydata[mydata.columns[0]].tolist()
print(channels)
subs = mydata[mydata.columns[1]].tolist()
views = mydata[mydata.columns[2]].tolist()
print(subs)
print(views)
# Create a pandas DataFrame with the preceding data, using the data of each respective group.
data = pd.DataFrame({'YouTube Channels': channels + channels, 'Subscribers/Views in millions': subs + views,
'Type': ['Subscribers'] * len(subs) + ['Views'] * len(views)})
data
# Visualize the given data using a FacetGrid with two columns. The first column should show the number of subscribers for each YouTube channel, whereas the second column should show the number of views.
sns.set()
g = sns.FacetGrid(data, col='Type', hue='Type', sharex=False, height=8)
g.map(sns.barplot, 'Subscribers/Views in millions', 'YouTube Channels')
plt.show()
| Chapter04/Activity4.04/Activity4.04.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="m87CBfRPGiea" executionInfo={"status": "ok", "timestamp": 1618669490842, "user_tz": -330, "elapsed": 1128, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11897582063931328693"}}
import numpy as np
import os
import pickle
import cv2
import numpy as np
import argparse
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.model_selection import GridSearchCV
from sklearn.feature_selection import SelectKBest, f_classif, chi2
from sklearn.metrics import *
import random
# + id="CG8hIfPAFHeI"
# !tar -xvzf /content/drive/MyDrive/LFW/zipped/lfw.tgz
# + colab={"base_uri": "https://localhost:8080/"} id="2DvEjCvYF_Zf" executionInfo={"status": "ok", "timestamp": 1618669494044, "user_tz": -330, "elapsed": 946, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11897582063931328693"}} outputId="5736cf09-c2ac-49c4-82e5-8d1e147e9f77"
persons = os.listdir("/content/lfw")
train_data = []
train_labels = []
print(len(persons))
# + id="N_PWG0aQGk-Q" executionInfo={"status": "ok", "timestamp": 1618669495900, "user_tz": -330, "elapsed": 1077, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11897582063931328693"}}
for i in range(3000):
p1,p2 = random.sample(range(0, 5749),2)
p1imgs = os.listdir(os.path.join("/content/lfw",persons[p1]))
p2imgs = os.listdir(os.path.join("/content/lfw",persons[p2]))
p1name = p1imgs[random.randint(0,len(p1imgs) - 1)]
p2name = p2imgs[random.randint(0,len(p2imgs) - 1)]
train_data.append([os.path.join("lfw/",persons[p1],p1name),os.path.join("lfw/",persons[p2],p2name),0])
# print(p1name,p2name,[os.path.join("lfw/",persons[p1],p1name),os.path.join("lfw/",persons[p2],p2name),0])
# + colab={"base_uri": "https://localhost:8080/"} id="jkLQmNBgOCwn" executionInfo={"status": "ok", "timestamp": 1618669499170, "user_tz": -330, "elapsed": 654, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11897582063931328693"}} outputId="5002c7ce-cbba-4a70-c7b8-1ca79efeffd8"
len(train_data)
# + colab={"base_uri": "https://localhost:8080/"} id="QJbCYMmgJsK5" executionInfo={"status": "ok", "timestamp": 1618669529261, "user_tz": -330, "elapsed": 994, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11897582063931328693"}} outputId="1e91b8dc-d066-497d-9b8e-aebb0c327d16"
cnt = 0
# train_data = []
for i in range(5749):
p1 = i
p1imgs = os.listdir(os.path.join("/content/lfw",persons[p1]))
if len(p1imgs) == 2 or len(p1imgs) == 3:
p1idx,p2idx = random.sample(range(0, len(p1imgs)),2)
p1name = p1imgs[p1idx]
p2name = p1imgs[p2idx]
train_data.append([os.path.join("lfw/",persons[p1],p1name),os.path.join("lfw/",persons[p1],p2name),1])
elif len(p1imgs) >= 4:
iter = random.randint(2,len(p1imgs) // 2)
for _ in range(iter):
p1idx,p2idx = random.sample(range(0, len(p1imgs)),2)
p1name = p1imgs[p1idx]
p2name = p1imgs[p2idx]
train_data.append([os.path.join("lfw/",persons[p1],p1name),os.path.join("lfw/",persons[p1],p2name),1])
# cnt += 1
# i -= 1
# print([os.path.join("lfw/",persons[p1],p1name),os.path.join("lfw/",persons[p1],p2name),1])
print(cnt)
# + id="OLgt4lMlL2fd" executionInfo={"status": "ok", "timestamp": 1618669793903, "user_tz": -330, "elapsed": 1040, "user": {"displayName": "sepnu paus", "photoUrl": "", "userId": "11897582063931328693"}}
# len(train_data)
# print(train_data[:][2])
import pickle
filehandler = open("/content/drive/MyDrive/project-sepnu/src/verification /dataset.pkl","wb")
pickle.dump(train_data,filehandler)
| src/verification classifier/verification_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GRASP+PR
#
# We can take the concepts of GRASP, elite sets and path relinking and combine them into a **hybrid** algorithm.
#
#
# ```python
# def grasp_plus_pr(max_iterations, time_limit):
# best = []
# elite_set = []
#
# do
# s = greedy_construction()
# s = local_search(s)
#
# if cost(s) > cost(best):
# best = s
#
# if len(elite_set) > 0:
# guiding = random(elite_set)
# s = path_relinking(s, guiding)
#
# update(s, elite_set)
#
# until max_iterations or time_limit
#
# return best
# ```
# ## Imports
import numpy as np
import sys
import time
# ## `metapy` imports
# install metapy if running in Google Colab
if 'google.colab' in sys.modules:
# !pip install meta-py
# +
from metapy.tsp import tsp_io as io
from metapy.tsp.euclidean import gen_matrix, plot_tour
from metapy.tsp.objective import OptimisedSimpleTSPObjective
from metapy.local_search.hill_climbing import (HillClimber, TweakTwoOpt)
from metapy.tsp.grasp import (SemiGreedyConstructor,
FixedRCLSizer,
RandomRCLSizer,
MonitoredLocalSearch,
GRASP,
EliteSet)
# -
# ## Load problem
# +
#load file
file_path = 'https://raw.githubusercontent.com/TomMonks/meta-py/main/data/st70.tsp'
#number of rows in the file that are meta_data
md_rows = 6
#read the coordinates
cities = io.read_coordinates(file_path, md_rows)
matrix = gen_matrix(cities, as_integer=True)
# -
# ## Implementation
# * `TSPPathRelinker` implements forward or backward path relinking logic with the option to truncate the relinking after a fixed number of moves.
# * `GRASPPlusPathRelinking` is an modification of the basic `GRASP` and adds path relinking plus an update of the elite set after each iteration.
class TSPPathRelinker:
'''
Path relinking for basic symmetric TSP.
Performs forward or backward relinking with a random
elite solution from the elite set.
After PR is complete a local search is used to take the solution
to a local optimum.
'''
def __init__(self, local_search, tracker, objective,
trunc=None, random_seed=None):
'''
Constructor
local_search: object
local search algorithm
tracker: EliteSet
The elite set tracker object
objective: object
TSP Objective
trunc: int, optional (default=None)
Truncate the relinking to a fixed number of steps
random_seed: int, optional (default=None)
used to control sampling from elite set
'''
self.local_search = local_search
self.tracker = tracker
self.obj = objective
self.trunc = trunc
# used for selecting from the elite set at random
self.rng = np.random.default_rng(random_seed)
self.calls = 0
self.made_it = 0
self.already_elite = 0
def evaluate_neighbour(self, from_city, current, guiding):
'''
Performs a two city swap and returns the cost.
Params:
------
from_city: int
The city to swap
current: np.ndarray
The current solution
guiding: np.ndarray
The guiding solution
obj: Object
The TSP objective function
Returns:
-------
float
'''
from_idx = np.where(current==from_city)[0][0]
to_idx = np.where(guiding==from_city)[0][0]
# swap and evaluate
current[from_idx], current[to_idx] = current[to_idx], current[from_idx]
cost= self.obj.evaluate(current)
#swap back
current[from_idx], current[to_idx] = current[to_idx], current[from_idx]
return cost
def next_restricted_neighbour_move(self, current, guiding):
'''
Iteratively search through a restricted neighbourhood based
on the guiding solution and greedily selects the best move.
Params:
-------
current: np.ndarray
The current solution
guiding: np.ndarray
The guiding solution
obj: Object
The TSP objective function
Returns:
-------
(np.ndarray, float)
(best_solution, best_cost)
'''
# cities in the restricted neighbourhood
swaps = current[current != guiding]
costs = np.full(len(swaps), -np.inf)
i = 0
# is there a way to eliminate the python loop?
for from_city in swaps:
# evaluate all swaps in current restricted neighbourhood
costs[i] = self.evaluate_neighbour(from_city, current, guiding)
i += 1
best_index = np.argmax(costs)
from_city = swaps[best_index]
# get index of cities in current and guiding solutions.
from_idx = np.where(current==from_city)[0][0]
to_idx = np.where(guiding==from_city)[0][0]
# swap and evaluate
current[from_idx], current[to_idx] = current[to_idx], current[from_idx]
return current, costs[best_index]
def relink(self, solution, elite_solution=None, direction='backward'):
'''
Relink between a current and guiding solution
Params:
-------
solution: np.ndarray
a none elite set TSP solution
elite_solution: np.ndarray.
guiding TSP solution
direction: str
forward: move from current towards guiding
backward: swap current and guiding and relink.
'''
# catch local optimum solutions that are already elite
if self.tracker.is_elite(solution):
self.already_elite += 1
return solution, self.obj.evaluate(solution)
# random elite solution if none specified
if elite_solution is None:
elite_solution = self.rng.choice(self.tracker.solutions)
# if 'forward' selected.
if direction == 'forward':
current, guiding = solution, elite_solution
elif direction == 'backward':
current, guiding = elite_solution, solution
else:
raise ValueError('Please select backward of forward relinking')
# moves to relink n / 2
n_moves = len(current[current != guiding]) // 2
# used to truncate path relinking
if self.trunc is not None:
if self.trunc > n_moves:
raise ValueError(f'@trunc must be <= moves to relink {n_moves}')
else:
n_moves = self.trunc
# path relinking
costs = [self.obj.evaluate(solution)]
solutions = [solution]
for i in range(n_moves):
current, cost = self.next_restricted_neighbour_move(current.copy(),
guiding)
solutions.append(current)
costs.append(cost)
# best solution and cost
# error here if the relinking has found no improvements.
best_idx = np.array(costs).argmax()
pr_solution, pr_cost = solutions[best_idx], costs[best_idx]
# local search on relinked solution...
self.local_search.set_init_solution(pr_solution)
self.local_search.solve()
# return local optimum
return self.local_search.best_solutions[0], self.local_search.best_cost
class GRASPPlusPathRelinking:
'''
Greedy Randomised Adaptive Search Procedure algorithm
hybridised with Path RelinkiGRASPPlusPathRelinkingng for the Travelling Salesman Problem
'''
def __init__(self, constructor, local_search, relinker,
tracker, max_iter=1000, time_limit=np.inf):
# semi greedy tour construction method
self.constructor = constructor
# local search procedure
self.local_search = local_search
# path relinker
self.path_relinker = relinker
# elite solution tracker
self.elite_tracker = tracker
# max runtime budget for GRASP
self.max_iter = max_iter
self.time_limit = time_limit
# init solution
self.best_solution = None
self.best = None
def solve(self):
'''
Run GRASP
Returns:
-------
None
'''
self.best_solution = None
self.best = -np.inf
i = 0
start = time.time()
while i < self.max_iter and ((time.time() - start) < self.time_limit):
i += 1
# construction phase
solution = self.constructor.build()
# Improve solution via local search
self.local_search.set_init_solution(solution)
self.local_search.solve()
current_solution = self.local_search.best_solutions[0]
current_cost = self.local_search.best_cost
if not self.elite_tracker.is_empty:
# backwards path relinking with elite solution selected at random
current_solution, current_cost = self.path_relinker.relink(current_solution)
# added because in theory best solution might not go into Elite set due to diversity
if current_cost > self.best:
self.best = current_cost
self.best_solution = current_solution
# update elite solutions
self.elite_tracker.update(current_solution, current_cost)
def compose_grasp(tour, maxtrix, max_iter=50, elite_set_size=10, rcl_size=5,
trunc=15, seeds=(None, None)):
# objective function
obj = OptimisedSimpleTSPObjective(-matrix)
# Two-opt tweaks
tweaker = TweakTwoOpt()
# local search for main GRASP = first improvement hill climbing
ls1 = HillClimber(obj, tour, tweaker)
# local search for path relinking = first improvement hill climbing
ls2 = HillClimber(obj, tour, tweaker)
# semi-greedy constructor and RCL sizer
sizer = FixedRCLSizer(rcl_size)
constructor = SemiGreedyConstructor(sizer, tour, -matrix,
random_seed=seeds[0])
# elite set tracker
tracker = EliteSet(min_delta=1, max_size=elite_set_size)
# path relinking logic
relinker = TSPPathRelinker(ls2, tracker, obj, trunc=trunc, random_seed=seeds[1])
# GRASP + PR framework
solver = GRASPPlusPathRelinking(constructor, ls1, relinker, tracker,
max_iter=max_iter)
return solver
# +
tour = np.arange(len(cities))
solver = compose_grasp(tour, matrix, max_iter=50, seeds=(42, 1966),
rcl_size=10)
print("\nRunning GRASP+PR")
solver.solve()
print("\n** GRASP OUTPUT ***")
print(f"best cost:\t{solver.best}")
print("best solutions:")
print(solver.best_solution)
fig, ax = plot_tour(solver.best_solution, cities, figsize=(12,9))
# -
| content/11_grasp_pr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="tXd5CerxmxGr"
# Make sure the tf version is 2.5.0-dev20201111 or later (for model saving)
# -
pwd
from datetime import datetime;now = datetime.now;t00 = now()
print(t00)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="yj5mv_ZvNl7m" outputId="3281fa15-4cc7-4cde-cd42-1785581a4f8c"
import tensorflow as tf
tf.test.gpu_device_name()
# -
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/"} id="Qq8i5RmyNZq-" outputId="f917cae9-d5c4-468a-bf96-d9770c9aa942"
# #%tensorflow_version 2.x
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
raise SystemError('GPU device not found')
print('Found GPU at: {}'.format(device_name))
# + colab={"base_uri": "https://localhost:8080/"} id="toHPNRS6Nbnc" outputId="09489db9-7851-4f35-da41-694915309b17"
# !nvidia-smi -L
# +
from importlib import reload
import helper_funcs;reload(helper_funcs);from helper_funcs import *
import make_models
reload(make_models)
from make_models import *
# Make tqdm work for colab
from functools import partial
from tqdm import tqdm
tqdm = partial(tqdm, position=0, leave=True)
# + colab={"base_uri": "https://localhost:8080/"} id="pVT3ZT60JTPF" outputId="cecd1920-04b9-4516-a13a-891d78fd8b68"
# #%tensorflow_version 2.x
import tensorflow as tf
import timeit,pickle
device_name = tf.test.gpu_device_name()
if device_name != '/device:GPU:0':
print(
'\n\nThis error most likely means that this notebook is not '
'configured to use a GPU. Change this in Notebook Settings via the '
'command palette (cmd/ctrl-shift-P) or the Edit menu.\n\n')
raise SystemError('GPU device not found')
def cpu():
with tf.device('/cpu:0'):
random_image_cpu = tf.random.normal((100, 100, 100, 3))
net_cpu = tf.keras.layers.Conv2D(32, 7)(random_image_cpu)
return tf.math.reduce_sum(net_cpu)
def gpu():
with tf.device('/device:GPU:0'):
random_image_gpu = tf.random.normal((100, 100, 100, 3))
net_gpu = tf.keras.layers.Conv2D(32, 7)(random_image_gpu)
return tf.math.reduce_sum(net_gpu)
# We run each op once to warm up; see: https://stackoverflow.com/a/45067900
cpu()
gpu()
# Run the op several times.
print('Time (s) to convolve 32x7x7x3 filter over random 100x100x100x3 images '
'(batch x height x width x channel). Sum of ten runs.')
print('CPU (s):')
cpu_time = timeit.timeit('cpu()', number=10, setup="from __main__ import cpu")
print(cpu_time)
print('GPU (s):')
gpu_time = timeit.timeit('gpu()', number=10, setup="from __main__ import gpu")
print(gpu_time)
print('GPU speedup over CPU: {}x'.format(int(cpu_time/gpu_time)))
# + colab={"base_uri": "https://localhost:8080/"} id="wrvyODvPJaOD" outputId="99239232-5868-4d63-d169-2946fc953b2b"
# from google.colab import drive
# drive.mount("/content/drive")
# print('done, mounted')
# + colab={"base_uri": "https://localhost:8080/"} id="RAduH185JatO" outputId="0ae69a86-c9a6-4999-b4e1-e817abe837cf"
# cd '/mmfs1/data/aglinska/BC-MRI-AE/Colab Notebooks'
# + colab={"base_uri": "https://localhost:8080/"} id="bjXikoR3dceV" outputId="850f352e-59c8-4b36-d080-034a4bc6ac8d"
# %matplotlib inline
import os
from matplotlib import pyplot as plt
import seaborn as sns
import numpy as np
import pandas as pd
from tqdm import tqdm
import pandas as pd
from sklearn.metrics import silhouette_score
#from mri_utils import get_MRI_CVAE_3D,get_MRI_CCVAE_3D
print(now()-t00)
# + colab={"base_uri": "https://localhost:8080/"} id="UmLCoBjXmF8M" outputId="cf65bd65-a145-4cb8-929a-683e17c757b1"
# cd ../
# + [markdown] id="t5xW313gmrFJ"
# SHAPE THE DATA
# + colab={"base_uri": "https://localhost:8080/"} id="STcZnREmuLkX" outputId="0ad489fa-c5ef-4fe2-ead8-d6cd94f3eb00"
arr = np.load('./Data/ABIDE-Anat-64iso.npz')
ABIDE_data = arr['data']
ABIDE_subs = arr['subs']
nsubs = ABIDE_data.shape[0]
[arr.shape for arr in [ABIDE_subs,ABIDE_data]]
# + colab={"base_uri": "https://localhost:8080/"} id="71-GquGIuLka" outputId="a3d3dfa4-01b0-412e-89ea-e506fd31b41f"
aa = np.array([ABIDE_data[s,:,:,:].sum() for s in range(ABIDE_data.shape[0])])<5000
ABIDE_data = ABIDE_data[~aa,:,:,:]
ABIDE_subs = ABIDE_subs[~aa]
[arr.shape for arr in [ABIDE_subs,ABIDE_data]]
# + colab={"base_uri": "https://localhost:8080/", "height": 284} id="JANBc_Nm-M2N" outputId="544e4bed-69dd-4064-b910-8e2d87b1dd1b"
df = pd.read_csv('./CSVs/ABIDE-legend.csv',header=0)
df = df.iloc[np.array([df['BIDS_ID'].values[s] in ABIDE_subs for s in range(len(df))])]
df.reset_index(inplace=True)
assert len(df)==len(ABIDE_subs),'dif lenghts'
assert all([df['BIDS_ID'][s]==ABIDE_subs[s] for s in range(len(df))]),'mismatch between df and goodsubs'
df.head(5)
# + colab={"base_uri": "https://localhost:8080/"} id="ZoqlzdzD-U6g" outputId="7f45f299-e6f7-43e2-93fc-a2a8b89af6d8"
#ABIDE_data = np.load('/mmfs1/data/aglinska/3d_grassy_mnist/fake_abide.npy')
TD_subs = ABIDE_data[np.array(df['Subject Type']=='CONTROL'),:,:,:]
DX_subs = ABIDE_data[np.array(df['Subject Type']=='PATIENT'),:,:,:]
print(TD_subs.shape)
print(DX_subs.shape)
# + id="xOdwBNsmF6Ee"
dxArr = np.ones(len(df))
dxArr[np.array(df['Subject Type']=='PATIENT')]=2
# +
# Get Scanner Types and Scanning Site
#df2 = pd.read_csv('/Users/aidasaglinskas/Desktop/BC-MRI-AE/CSVs/ABIDE_BIDS_PATHS.csv',index_col=0)
df2 = pd.read_csv('/mmfs1/data/aglinska/BC-MRI-AE/CSVs/ABIDE_BIDS_PATHS.csv',index_col=0)
df2.index=np.arange(len(df2))
df2 = df2.iloc[np.array([df2['subID'].values[s] in df['BIDS_ID'].values for s in np.arange(len(df2))])]
df2.index=np.arange(len(df2))
df2 = df2.sort_values('subID')
df2.index=np.arange(len(df2))
print(f'{len(df2)}/{len(df)}')
n = len(df2)
df2.head()
assert len(df2)==len(df), 'different lenghts of CSVs'
assert all(np.array([df2['subID'].values[s]==df['BIDS_ID'].values[s] for s in np.arange(len(df))])),'mismatch between CSV orders'
# -
df['ScanSite'] = [val.split('/')[6] for val in df2['sub_path'].values]
df['ScannerType'] = [val.split('/')[8] for val in df2['sub_path'].values]
# +
df['ScannerID'] = str_to_ordinal(df['ScannerType'].values)
df['ScanSiteID'] = str_to_ordinal(df['ScanSite'].values)
patients = df['DxGroup'].values==1
controls = df['DxGroup'].values==2
# -
print(df.shape)
print(ABIDE_data.shape)
dataFnOut = '/mmfs1/data/aglinska/BC-MRI-AE/Data/ABIDE-Anat-64iso-S982.npz'
dfFnOut = '/mmfs1/data/aglinska/BC-MRI-AE/Data/ABIDE_legend.csv'
#np.savez_compressed(dataFnOut,data=ABIDE_data)
df.to_csv(dfFnOut)
# + [markdown] id="BTN1pbr7mnxq"
# # TRAIN THE VAE
# + id="RotozUR54jNt"
train_vae = False
# + id="ny0tO8kropHG"
if train_vae:
from scipy.spatial.distance import pdist
from scipy.spatial.distance import squareform
def plot_recon_vae():
plt.figure(figsize=(5,5));
plt.subplot(2,2,1)
plt.imshow(DX_batch[0,:,:,40]);plt.xticks([]);plt.yticks([]);plt.title('input');
plt.subplot(2,2,2)
plt.imshow(cvae.predict([DX_batch,TD_batch])[0][0,:,:,40,0]);plt.xticks([]);plt.yticks([]);plt.title('reconstruction');
plt.subplot(2,2,3)
plt.imshow(TD_batch[0,32,:,:]);plt.xticks([]);plt.yticks([]);
plt.subplot(2,2,4)
plt.imshow(cvae.predict([DX_batch,TD_batch])[1][0,32,:,:,0]);plt.xticks([]);plt.yticks([]);
# + id="qllddMarnEr2"
import pickle
if train_vae:
latent_dim=32
batch_size=64
disentangle=False
gamma=1
encoder, decoder, vae = get_MRI_CVAE_3D(input_shape=(64, 64, 64, 1), latent_dim=latent_dim, batch_size=batch_size, disentangle=disentangle, gamma=gamma)
loss = list()
print('ready')
#fn = '/mmfs1/data/aglinska/tf_outputs/VAE/Jan4th'
fn = '/mmfs1/data/aglinska/tf_outputs/VAE/Jan12th_16d'
#fn = '/mmfs1/data/aglinska/tf_outputs/VAE/VAE_2D_b64_NoDis_g1'
if os.path.exists(fn+'_loss.pickle'):
print('loading weights')
vae.load_weights(fn)
loss = pickle.load(open(fn+'_loss.pickle','rb'))
else:
print('no weights found - initializing new')
print(fn)
# -
if train_vae:
im1 = ABIDE_data[0:5,:,:,:][0,32,:,:];
im = vae.predict(ABIDE_data[0:5,:,:,:])[0,32,:,:,0];
plot_trainProgress(loss,im,im1);
if train_vae:
batch_size = 64
for i in tqdm(range(1,10000)):
history = vae.train_on_batch(ABIDE_data[np.random.randint(low=0,high=ABIDE_data.shape[0],size=batch_size),:,:,:]);
loss.append(history);
if np.mod(i,25)==0:
im1 = ABIDE_data[0:5,:,:,:][0,32,:,:];
im = vae.predict(ABIDE_data[0:5,:,:,:])[0,32,:,:,0];
plot_trainProgress(loss,im,im1);
if np.mod(i,100)==0:
pickle.dump(loss,open(fn+'_loss.pickle','wb'))
vae.save_weights(fn)
# + [markdown] id="LAbH8lm-nXUt"
# # TRAIN THE CONTRASTIVE VAE
# -
train_cvae=True
items = get_weights(fdir=None)
if train_cvae:
latent_dim = 16
batch_size = 32
#batch_size = 64
beta = 1;gamma = 100
disentangle = True
cvae, z_encoder, s_encoder, cvae_decoder = get_MRI_CCVAE_3D(latent_dim=latent_dim,beta=beta, disentangle=disentangle, gamma=gamma, bias=True, batch_size = batch_size)
loss = list()
fdir = '/mmfs1/data/aglinska/tf_outputs/CVAE/'
#fn = f'{now().strftime("%b%d_%H%M")}_D{latent_dim}B{beta}G{gamma}_D{disentangle}'
fn = 'Feb04_0524_D16B1G100_DTrue'
fn = os.path.join(fdir,fn)
loss = pickle.load(open(fn+'_loss.pickle','rb'))
cvae.load_weights(fn)
fn
# +
# Initial Check
import helper_funcs;reload(helper_funcs);from helper_funcs import *
DX_batch = DX_subs[np.random.randint(low=0,high=DX_subs.shape[0],size=batch_size),:,:,:];
TD_batch = TD_subs[np.random.randint(low=0,high=TD_subs.shape[0],size=batch_size),:,:,:];
if len(loss)==0:
loss.append(np.nan)
im,im1,ss = cvae_query(ABIDE_data,s_encoder,z_encoder,cvae_decoder);
plot_trainProgress(loss,im,im1);
loss = list()
else:
im,im1,ss = cvae_query(ABIDE_data,s_encoder,z_encoder,cvae_decoder);
plot_trainProgress(loss,im,im1);
# + colab={"base_uri": "https://localhost:8080/", "height": 903} id="f3G3VigW9erL" outputId="5da10070-dde3-4d30-cabc-b795a2827082"
import helper_funcs;reload(helper_funcs);from helper_funcs import *
for i in tqdm(range(1,int(1e6))):
#asd_idx,td_idx = get_batch_idx(df,batch_size = batch_size)
#DX_batch = np.take(ABIDE_data, indices=asd_idx, axis=0)
#TD_batch = np.take(ABIDE_data, indices=td_idx, axis=0)
DX_batch = DX_subs[np.random.randint(low=0,high=DX_subs.shape[0],size=batch_size),:,:,:];
TD_batch = TD_subs[np.random.randint(low=0,high=TD_subs.shape[0],size=batch_size),:,:,:];
#print(TD_batch.shape)
hist = cvae.train_on_batch([DX_batch,TD_batch]); # Proper
#hist = cvae.train_on_batch([TD_batch,DX_batch]); # Flipped
assert not np.isnan(hist),'loss is NaN - you f**cked up'
im,im1,ss = net_query();
loss.append(hist);
if np.mod(i,100)==0:
plot_trainProgress(loss,im,im1);
pickle.dump(loss,open(fn+'_loss.pickle','wb'))
plot_four(DX_batch, TD_batch, z_encoder, s_encoder,cvae_decoder,cvae,idx=0)
plot_four(DX_batch, TD_batch, z_encoder, s_encoder,cvae_decoder,cvae,idx=1)
bg_space,sl_space = get_spaces(ABIDE_data, z_encoder, s_encoder, w=2)
cscatter([bg_space,sl_space],c=df['DxGroup'].values)
plt.figure(figsize=(5,5))
plot_sweep(ABIDE_data, z_encoder, s_encoder,cvae_decoder, wspace='z', l=5)
plt.show()
plt.figure(figsize=(5,5))
plot_sweep(ABIDE_data, z_encoder, s_encoder,cvae_decoder,wspace='s', l=5)
plt.show()
plot_cvae_silhouettes(ABIDE_data,z_encoder,s_encoder,patients,keys=None,l=8)
plot_cvae_dif_mat(ABIDE_data,z_encoder,s_encoder,patients)
plot_cvae_dif_mat(ABIDE_data,z_encoder,s_encoder,controls,keys = ['AgeAtScan','ScannerID','ScanSiteID','FIQ'])
if np.mod(i,101)==0:
cvae.save_weights(fn)
# -
| Code/tut-MRI_CCAE_3D_TRAIN_CCAE_andromeda_fx.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.2.0
# language: julia
# name: julia-1.2
# ---
# Imagine we are measuring voltages of a sine wave, and we want to constrain the amplitude and phase.
#
# Our measurements have Gaussian noise of known standard deviation, and occur at known times (assume we can measure time so well that there is practically no uncertainty).
# +
# "y" are our measurements
y = Array([-1.64, 4.63, 6.09, 5.05, 2.66, -0.90, -6.09, -7.16, -2.54, -1.84])
# "noise_std" is the standard deviation of the Gaussian noise on our measurements
noise_std = 1.0
nsamples = length(y)
# "times" are when the measurements were taken (evenly spaced)
times = collect(LinRange(0.0, 2.0*pi, nsamples));
# -
using Plots
plot(times, y)
# Now, we have to write our *model* function. Our model is that this is a sine wave with parameters of *phase offset* and *amplitude*.
function sine_model(t, phase, amplitude)
return amplitude * sin.(t .- phase)
end;
tplot = collect(LinRange(0.0, 2.0*pi, 200));
# I'm going to guess at parameters: phase = 0, amplitude = 6
plot(tplot, sine_model(tplot, 0., 6))
plot!(times, y)
# Now, in order to use MCMC, we must write down our *likelihood*. Given model parameters, how probable are the measurements we made?
function sine_log_likelihood(t, y, phase, amplitude)
ymodel = sine_model(t, phase, amplitude)
log_likelihood = -0.5 * sum((y - ymodel).^2 / noise_std.^2)
return log_likelihood
end;
sine_log_likelihood(times, y, 0., 6.)
sine_log_likelihood(times, y, 0.05, 6.)
# Now, since this is a low-dimensional problem, we can just grid up the parameter space and compute the likelihood at each position! In real problems, you can't do this....
plo,phi = 0., 0.5
alo,ahi = 5., 8.
pp = range(plo, stop=phi, length=100)
aa = range(alo, stop=ahi, length=100)
LL = zeros((length(pp), length(aa)))
for i in 1:length(pp)
for j in 1:length(aa)
LL[i, j] = sine_log_likelihood(times, y, pp[j], aa[i])
end
end
heatmap(pp, aa, LL, xlabel="Phase", ylabel="Amplitude")
heatmap(pp, aa, exp.(LL), xlabel="Phase", ylabel="Amplitude")
# Now, we're going to code up MCMC to see if we can sample this parameter space and get the same answer!
# First, we need to go from a log-likelihood to a log-posterior. This *requires* specifying a *prior* on the parameters.
#
# A common thing to do is to choose a "flat" prior: the prior is constant over the entire range. This is technically wrong ("improper prior").
function sine_log_posterior(t, y, phase, amplitude)
log_like = sine_log_likelihood(t, y, phase, amplitude)
# Flat prior! No preference for phase, amplitude values.
log_prior = 0.
return log_like + log_prior
end;
# Now the actual MCMC algorithm! We need to choose a "proposal distribution" -- how to select the next candidate parameters to jump to. We will make these Gaussians with fixed variances.
# +
chain = []
# initial parameter guess
param_phase = 0.
param_amp = 6.
# proposal jump sizes
jump_phase = 0.01
jump_amp = 0.1
# initial log-posterior.
logprob = sine_log_posterior(times, y, param_phase, param_amp)
# how many steps to take
nsteps = 1_000
for i in 1:nsteps
# propose new parameter values
param_phase_new = param_phase + randn() * jump_phase
param_amp_new = param_amp + randn() * jump_amp
# compute log-posterior at new parameters
logprob_new = sine_log_posterior(times, y, param_phase_new, param_amp_new)
if (exp(logprob_new - logprob) >= rand(Float64))
logprob = logprob_new
param_phase = param_phase_new
param_amp = param_amp_new
end
append!(chain, (param_phase, param_amp))
end
# append! makes "chain" a 1-d vector; reshape to a matrix
chain = reshape(chain, (2,Int64(length(chain)/2)))';
# -
chain
plot(chain[:,1], chain[:,2], xlabel="Phase", ylabel="Amplitude")
histogram2d(chain[:,1], chain[:,2], xlabel="Phase", ylabel="Amplitude")
histogram(chain[:,1], xlabel="Phase", ylabel="Number of MCMC samples")
histogram(chain[:,2], xlabel="Amplitude", ylabel="Number of MCMC samples")
| .ipynb_checkpoints/mcmc-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # VacationPy
# ----
#
# #### Note
# * Keep an eye on your API usage. Use https://developers.google.com/maps/reporting/gmp-reporting as reference for how to monitor your usage and billing.
#
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import requests
import gmaps
import os
import json
# Import API key
from config import g_key
# Must enable gmaps extension to display interactive gmaps in jupyter notebook
# jupyter nbextension enable --py gmaps
# -
# ### Store Part I results into DataFrame
# * Load the csv exported in Part I to a DataFrame
# +
# Load cities.csv from part I
filepath = os.path.join('..', 'WeatherPy', 'Data', 'cities.csv')
cities_data = pd.read_csv(filepath)
# Display DataFrame
cities_data.head()
# +
# Count data points
print(cities_data.count())
# Lat and long already stored as float
cities_data.dtypes
# -
# ### Humidity Heatmap
# * Configure gmaps.
# * Use the Lat and Lng as locations and Humidity as the weight.
# * Add Heatmap layer to map.
# Configure gmaps with API key
gmaps.configure(api_key=g_key)
# +
# Store latitude and longitude data
locations = cities_data[['Latitude', 'Longitude']]
# locations
# Store humidity data
humidity = cities_data['Humidity (%)'].astype(float)
# humidity
# Find max humidity for gmaps max intensity
max_intensity = humidity.max()
# max_intensity
# +
# Create map
fig = gmaps.figure()
# Add heatmap layer displaying humidity
heat_layer = gmaps.heatmap_layer(locations, weights=humidity, dissipating=False,
max_intensity=max_intensity, point_radius=3)
fig.add_layer(heat_layer)
fig
# -
# ### Create new DataFrame fitting weather criteria
# * Narrow down the cities to fit weather conditions.
# * Drop any rows will null values.
# Drop rows with null values
no_null_cities = cities_data.dropna()
no_null_cities
# +
# Narrow down to cities with (1) max temp less than 80*F, (2) max temp greater than 70*F,
# (3) wind speed less than 10 MPH, (4) 0% cloudiness, and (5) humidity less than 80%
ideal_conditions = no_null_cities.loc[(no_null_cities['Max Temperature (*F)'] < 80) &
(no_null_cities['Max Temperature (*F)'] > 70) &
(no_null_cities['Wind Speed (MPH)'] < 10) &
(no_null_cities['Cloudiness (%)'] == 0) &
(no_null_cities['Humidity (%)'] < 80)]
narrow_cities_df = ideal_conditions.reset_index(drop=True)
narrow_cities_df.count()
# -
# ### Hotel Map
# * Store into variable named `hotel_df`.
# * Add a "Hotel Name" column to the DataFrame.
# * Set parameters to search for hotels with 5000 meters.
# * Hit the Google Places API for each city's coordinates.
# * Store the first Hotel result into the DataFrame.
# * Plot markers on top of the heatmap.
# Create lists to hold hotel API response data
h_names = []
h_cities = []
h_countries = []
h_lats = []
h_lngs = []
# +
# Base url for API queries
base_url = 'https://maps.googleapis.com/maps/api/place/nearbysearch/json'
count = 1
# Iterate through each row in hotel_df
for index, row in narrow_cities_df.iterrows():
# Iteration latitude and longitude values
i_lat = narrow_cities_df.iloc[index, 2]
i_lng = narrow_cities_df.iloc[index, 3]
# Save variables to use in params
i_location = f'{i_lat}, {i_lng}'
i_radius = 5000
i_type = 'lodging'
i_keyword = 'hotel'
# Gmaps parameters dictionary
params = {
'location': i_location,
'radius': i_radius,
'types': i_type,
'keyword': i_keyword,
'key': g_key
}
# Call API request with response in .json format
hotel_response = requests.get(base_url, params).json()
# TEST ONLY
# print(hotel_response.url)
# hotel_response = hotel_response.json()
# Print count and city
print(f'Search {count} of {len(narrow_cities_df)}: Hotels near "{narrow_cities_df.iloc[index, 0]}" ')
# Incorporate try/except to skip missing responses and continue iteration
try:
print(f"First Result: {hotel_response['results'][0]['name']}")
h_names.append(hotel_response['results'][0]['name'])
h_cities.append(narrow_cities_df.iloc[index, 0])
h_countries.append(narrow_cities_df.iloc[index, 1])
h_lats.append(hotel_response['results'][0]['geometry']['location']['lat'])
h_lngs.append(hotel_response['results'][0]['geometry']['location']['lng'])
except (IndexError):
print(f'Sorry! No hotels matched search parameters for this city.')
pass
print(f'----------------------------------------------------------')
count = count + 1
# +
# Tests
# print(h_names)
# print(h_cities)
# print(h_countries)
# print(h_lats)
# print(h_lngs)
# +
hotel_dict = {
'City': h_cities,
'Country': h_countries,
'Latitude': h_lats,
'Longitude': h_lngs,
'Hotel Name': h_names
}
hotel_df = pd.DataFrame(hotel_dict)
hotel_df
# +
# NOTE: Do not change any of the code in this cell
# Using the template add the hotel marks to the heatmap
info_box_template = """
<dl>
<dt>Name</dt><dd>{Hotel Name}</dd>
<dt>City</dt><dd>{City}</dd>
<dt>Country</dt><dd>{Country}</dd>
</dl>
"""
# Store the DataFrame Row
# NOTE: be sure to update with your DataFrame name
hotel_info = [info_box_template.format(**row) for index, row in hotel_df.iterrows()]
locations = hotel_df[["Latitude", "Longitude"]]
# +
# Add marker layer ontop of heat map
marker_layer = gmaps.marker_layer(locations, info_box_content=hotel_info)
fig.add_layer(marker_layer)
# Display figure
fig
# -
| VacationPy/VacationPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
X = np.load("souvik/xtrain.npy")
y = np.load("souvik/ytrain.npy")
X1 = np.load('souvik/xtrain_e.npy')
X = X[:,100:200,100:200,:]
X1 = X1[:,100:200,100:200]
X.shape, X1.shape
X = X.astype('float32')
X1 = np.reshape(X1,(-1,100,100,1))
Xtrain = np.concatenate([X,X1],axis=3)
xtrain.shape
def giveSet(a,b):
xs = []
ys = []
bs = 59
for i in range(8):
xs.append(Xtrain[(i*59):((i+1)*59)])
ys.append(y[(i*59):((i+1)*59)])
xtest = np.concatenate([xs[a],xs[b]])
ytest = np.concatenate([ys[a],ys[b]])
xs.pop(a)
xs.pop(b)
ys.pop(a)
ys.pop(b)
xtrain = np.concatenate([xs[0],xs[1],xs[2],xs[3],xs[4],xs[5]])
ytrain = np.concatenate([ys[0],ys[1],ys[2],ys[3],ys[4],ys[5]])
return xtrain,ytrain,xtest,ytest
xtrain,ytrain,xval,yval = giveSet(0,4)
(b==1).sum(),(b==0).sum(), (d==0).sum(), (d==1).sum()
07 16 25 34 04 15 26 37
| fold_sets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Marc-Serenio/OOP-1-1/blob/main/Classes_and_Objects.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="fT3EihYQFV0a"
# Application 2 - Write a python program that displays the Full name of a student, student number, age,course, and school. Create a class name OOP_1-1and create Info()method to display the information of a student with fullname, student_no, age, course, school as attributes
# + colab={"base_uri": "https://localhost:8080/"} id="Fy9imc45FW5Y" outputId="bc5c8877-2778-4e0c-b25c-baac7497581e"
class OOP_1_1:
def __init__(self,fullname,student_no,age,course,school):
self.fullname = fullname
self.student_no = student_no
self.age = age
self.course = course
self.school = school
def info(self):
print(self.fullname,self.student_no,self.age,self.course,self.school)
student = OOP_1_1("<NAME>",202102057,19,"BSCpE","CVSU")
student.info()
# + colab={"base_uri": "https://localhost:8080/"} id="BkE7-NFjGpuQ" outputId="f5c358c5-aa72-42ff-ae9b-d01ce332c674"
class OOP_1_1:
def __init__(self,fullname,student_no,age,course,school):
self.fullname = fullname
self.student_no = student_no
self.age = age
self.course = course
self.school = school
def info(self):
print("My Name is", self.fullname,self.student_no,self.age,self.course,self.school)
#print("My Name is",self.fullname)
#print("My Name is",self.student_no)
#print("My Name is",self.age)
#print("My Name is",self.course)
#print("My Name is",self.school)
student = OOP_1_1("<NAME>",202102057,19,"BSCpE","CVSU")
student.info()
# + colab={"base_uri": "https://localhost:8080/"} id="QSnnbF8cJUd7" outputId="1bb5f8f3-5f12-44cd-f8b8-a7516c207b61"
class Circle:
def __init__(self,r):
self.radius = r
def area(self):
return self.radius**2*3.14
def perimeter(self):
return 2*self.radius*3.14
NewCircle = Circle(8)
print(NewCircle.area())
print(NewCircle.perimeter())
| Classes_and_Objects.ipynb |