code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_amazonei_pytorch_latest_p36
# language: python
# name: conda_amazonei_pytorch_latest_p36
# ---
# %load_ext autoreload
# %autoreload 2
from GANime.gan import GAN, plotter
from GANime.datasets import ImageOnlyDataset
# +
from torch.utils.data import DataLoader
import torchvision.transforms as T
resize_transform = T.Compose([
T.Resize(64),
T.ToTensor(),
T.Lambda(lambda x: (255*x).int()/127.5-1) #normalize color channels to -1 and 1
])
ds = ImageOnlyDataset('out2', resize_transform)
dl = DataLoader(ds, batch_size=128, shuffle=True)
# -
it = next(iter(dl))
plotter(it, rows=8, columns=8, renormalize_func = lambda x: (x*127.5+127.5).astype(int))
seed_size = 128
gan_model = GAN(seed_size)
gan_model.train(dl,
num_epochs = 20,
batch_size = 128,
plot = True,)
| generate_images.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import tempfile, sys, os
import numpy as np
from scipy.misc import imread
import tensorflow as tf
from deepexplain.tensorflow import DeepExplain
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '-1'
config = tf.ConfigProto(
device_count = {'GPU': 0}
)
config.gpu_options.allocator_type = 'BFC'
def load_graph(model_file):
graph = tf.Graph()
graph_def = tf.GraphDef()
with open(model_file, "rb") as f:
graph_def.ParseFromString(f.read())
with graph.as_default():
tf.import_graph_def(graph_def)
return graph
def read_tensor_from_image_file(file_name,
input_height=299,
input_width=299,
input_mean=0,
input_std=255):
input_name = "file_reader"
output_name = "normalized"
file_reader = tf.read_file(file_name, input_name)
if file_name.endswith(".png"):
image_reader = tf.image.decode_png(
file_reader, channels=3, name="png_reader")
elif file_name.endswith(".gif"):
image_reader = tf.squeeze(
tf.image.decode_gif(file_reader, name="gif_reader"))
elif file_name.endswith(".bmp"):
image_reader = tf.image.decode_bmp(file_reader, name="bmp_reader")
else:
image_reader = tf.image.decode_jpeg(
file_reader, channels=3, name="jpeg_reader")
float_caster = tf.cast(image_reader, tf.float32)
dims_expander = tf.expand_dims(float_caster, 0)
resized = tf.image.resize_bilinear(dims_expander, [input_height, input_width])
normalized = tf.divide(tf.subtract(resized, [input_mean]), [input_std])
sess = tf.Session(config=config)
result = sess.run(normalized)
return result
def load_labels(label_file):
label = []
proto_as_ascii_lines = tf.gfile.GFile(label_file).readlines()
for l in proto_as_ascii_lines:
label.append(l.rstrip())
return label
# -
# ## file_name = "/Z/personal-folders/interns/saket/histopath_data/baidu_images/training/tumor/tumor_100_60249_38668_256.jpg"
# model_file = "/Z/personal-folders/interns/saket/resnetv2_retrain_model_camelyon16_200000_alldata_100k/retrained_graph.pb"
# label_file = "/Z/personal-folders/interns/saket/resnetv2_retrain_model_camelyon16_200000_alldata_100k/retrained_labels.txt"
# input_height = 299
# input_width = 299
# input_mean = 0
# input_std = 255
# input_layer = "Placeholder"
# output_layer = "final_result"
#
#
#
# graph = load_graph(model_file)
# t = read_tensor_from_image_file(
# file_name,
# input_height=input_height,
# input_width=input_width,
# input_mean=input_mean,
# input_std=input_std)
#
# input_name = "import/" + input_layer
# output_name = "import/" + output_layer
# input_operation = graph.get_operation_by_name(input_name)
# output_operation = graph.get_operation_by_name(output_name)
#
# with tf.Session(graph=graph, config=config) as sess:
# results = sess.run(output_operation.outputs[0], {
# input_operation.outputs[0]: t
# })
# results = np.squeeze(results)
#
# top_k = results.argsort()[-5:][::-1]
# labels = load_labels(label_file)
# for i in top_k:
# print(labels[i], results[i])
# +
xs = np.array([1])
tf.reset_default_graph()
# Since we will explain it, the model has to be wrapped in a DeepExplain context
with tf.Session(graph=graph, config=config) as sess:
with DeepExplain(session=sess, graph=sess.graph) as de:
results = sess.run(output_operation.outputs[0], {
input_operation.outputs[0]: t
})
#X = t
logits = np.squeeze(results)
top_k = logits.argsort()[-5:][::-1]
labels = load_labels(label_file)
with tf.Session(graph=graph, config=config) as sess:
t = read_tensor_from_image_file(
file_name,
input_height=input_height,
input_width=input_width,
input_mean=input_mean,
input_std=input_std)
input_name = "import/" + input_layer
output_name = "import/" + output_layer
input_operation = graph.get_operation_by_name(input_name)
output_operation = graph.get_operation_by_name(output_name)
with DeepExplain(session=sess, graph=sess.graph) as de:
#X = tf.placeholder(tf.float32, shape=(None, 299, 299, 3))
#explain(method_name, target_tensor, input_tensor, samples, ...args)
attributions = {
# Gradient-based
# NOTE: reduce_max is used to select the output unit for the class predicted by the classifier
# For an example of how to use the ground-truth labels instead, see mnist_cnn_keras notebook
'Saliency maps': de.explain('saliency', output_operation.outputs[0]* [0, 1],
input_operation.outputs[0],
t),
'DeepLIFT (Rescale)': de.explain('deeplift', output_operation.outputs[0] * [0, 1], input_operation.outputs[0], t),
'Gradient * Input': de.explain('grad*input', output_operation.outputs[0]* [0, 1], input_operation.outputs[0], t),
'Integrated Gradients': de.explain('intgrad', output_operation.outputs[0]* [0, 1], input_operation.outputs[0], t),
'Epsilon-LRP': de.explain('elrp', output_operation.outputs[0]* [0, 1], input_operation.outputs[0], t),
# Perturbation-based (comment out to evaluate, but this will take a while!)
# 'Occlusion [15x15]': de.explain('occlusion', tf.reduce_max(logits, 1), X, xs, window_shape=(15,15,3), step=4)
}
print ("Done!")
# -
output_operation.outputs[0] * [0, 1]
# +
from utils import plot, plt
# %matplotlib inline
xs = t
attributions_reduced = attributions
#attributions_reduced['Integrated Gradients'] = attributions['Integrated Gradients']
n_cols = int(len(attributions_reduced)) + 1
n_rows = len(xs)
fig, axes = plt.subplots(nrows=n_rows, ncols=n_cols, figsize=(3*n_cols, 3*n_rows))
for i, xi in enumerate(xs):
xi = (xi - np.min(xi))
xi /= np.max(xi)
ax = axes.flatten()[i*n_cols]
ax.imshow(xi)
ax.set_title('Original')
ax.axis('off')
for j, a in enumerate(attributions_reduced):
axj = axes.flatten()[i*n_cols + j + 1]
plot(attributions[a][i], xi = xi, axis=axj, dilation=.5, percentile=99, alpha=.2).set_title(a)
# -
| notebooks/DeepExplain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Assignment 1 | Data Types
#
# Add code cells as needed for your answers.
# ### Exercise 1: Manipulating Lists
# Create a list containing the numbers 10, 20, and 30. Store your list as a variable named `a`. Then create a second list containing the numbers 30, 60, and 90. Call this this `b`.
a = [10,20,30]
b = [30,60,90]
# In the cells below, write Python expressions to create the following four outputs by combining `a` and `b` in creative ways:
#
# 1. [[10, 20, 30], [30, 60, 90]]
#
# 2. [10, 20, 30, 30, 60, 90]
#
# 3. [10, 20, 60, 90]
#
# 4. [20, 40, 60]
q = [a,b]
print(q)
c = (a + b)
print(c)
e = a[0:2]+ b[1:3]
print(e)
f = a[1:2]+[sum(a[0:1]+b[0:1])]+b[1:2]
print(f)
# ### Exercise 2. Working with Lists
#
# Create a list that contains the sums of each of the lists in G.
#
# `G = [[13, 9, 8], [14, 6, 12], [10, 13, 11], [7, 18, 9]]`
#
# Your output should look like:
#
# - `[30, 32, 34, 34]`
#
# Hint: try computing the sum for just one list first.
G = [[13, 9, 8], [14, 6, 12], [10, 13, 11], [7, 18, 9]]
G[0]
sum(G[0])
H = [sum(G[0]),sum(G[1]),sum(G[2]),sum(G[3])]
H
# ### Exercise 3: String Manipulation
#
# Turn the string below into 'all good countrymen' using the minimum amount of code, using only the methods we've covered so far. A couple of lines of code should do the trick. Note: this requires string and list methods.
s = 'Now is the time for all good men to come to the aid of their country!'
print(s.replace("Now is the time for all good men to come to the aid of their country!", "all good countrymen"))
# ### Exercise 4: String Manipulation and Type Conversion
#
# Define a variable `a = "Sarah earns $96500 in a year"`. Then maniuplate the value of `a` in order to print the following string: `Sarah earns $8041.67 monthly`
#
# Start by doing it in several steps and then combine them one step at a time until you can do it in one line.
a = "Sarah earns $96500 in a year"
print(a.replace("$96500 in a year","$8041.67 monthly"))
# ### Exercise 5: Create and Query a Dictionary on State Demographics
#
# Create two dictionaries, one for California and one for New York state, based on the data in the following table:
#
# | States | Pop Density | Prop White | Prop Afr Amer | Prop Asian | Prop Other | Owners | Renters |
# | --- | ---: | ---: | ---: | ---: | ---: | ---: | ---: |
# | CA | 239.1 | 0.57 | 0.06 | 0.13 | 0.22 | 7035371 | 5542127 |
# | NY | 411.2 | 0.65 | 0.15 | 0.07 | 0.22 | 3897837 | 3419918 |
#
# Each dictionary should have the following keys and value types: `name: (string)` , `population density: (float)`, `race (dict)`, `tenure: (dict)`.
#
# 1. Create one dictionary called CA and one called NY that contain dictionaries containing name, pop_density, race as a dictionary, and tenure for California and New York. Now combine these into a dictionary called "states", making it a dictionary of dictionaries, or a nested dictionary.
#
# 1. Check if Texas is in our state dictionary (we know it isn't but show us).
#
# 1. Print the White population in New York as a percentage
#
# 1. Assume there was a typo in the data, and update the White population fraction of NY to 0.64. Verify that it was updated by printing the percentage again.
#
# 1. Print the percentage of households that are renters in California, with two decimal places
race_CA = {'white':0.57, 'Afr Amer':0.06, 'Asia':0.13,'others':0.22}
tenure_CA = {'owners':7035371, 'Renters':5542127}
CA ={'name':"CA", 'population density':239.1, 'race':race_CA,'tenure':tenure_CA}
CA
race_NY = {'white':0.65, 'Afr Amer':0.15, 'Asia':0.07,'others':0.22}
tenure_NY = {'owners':3897837, 'Renters':3419918}
NY = {'name':'NY', 'population density':'411.2', 'race':race_NY,'tenure':tenure_NY}
NY
#question one
states = {'CA':CA,'NY':NY}
states
#question two
print('Texas' in states)
#question three
print(NY['race']['white']*100 )
#question Four
NY['race']['white'] = 0.64
print (NY['race']['white']*100)
#question Five
print(round(CA['tenure']['Renters'] / (CA['tenure']['Renters'] + CA['tenure']['owners']) * 100,2))
# ### Exercise 6: Working with Numpy Arrays
#
# 1. Create and print a 4 x 4 array named `a` with value 3 everywhere.
#
# 1. Create and print a 4 x 4 array named `b` with elements drawn from a uniform random distribution
#
# 1. Create and print array `c` by dividing a by b
#
# 1. Compute and print the min, mean, max, median, and 90th percentile values of `c`
#
# 1. Compute and print the sum of the second column in `c`
import numpy as np
#Question 1
zero = np.zeros((4,4))
a = zero + 3
a
#Question 2
b = np.random.uniform(size = (4,4))
b
#Question 3
c = a/b
c
#Question 4
print(c.min())
print(c.mean())
print(c.max())
print(np.median(c))
print(np.percentile(c,.9))
#Question 5
Q5 = sum(c.T[1])
print(Q5)
| assignments/assignment_1/assignment_1_chenlu_zhu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--Información del curso-->
# <img align="left" style="padding-right:10px;" src="figuras/banner_sdc.png">
#
# <br><br><center><h1 style="font-size:2em;color:#2467C0"> Región de interés en un video personal </h1></center>
# <br>
# <table>
# <col width="550">
# <col width="450">
# <tr>
#
#
# <td><img src="figuras/carreteras.png" align="middle" style="width:550px;"/></td>
# <td>
#
# En esta lección aplicaremos lo aprendido en la lección 4 pero ahora en un video personal.
#
# <br>
# </td>
# </tr>
# </table>
# # Cargar librerias
import matplotlib.pyplot as plt
import numpy as np
import cv2 as cv
import time
# # Cargar el video
video_name = "videos/video_Erick_Casanova.mp4"
# +
#Una vez abierto el video, es necesario presionar "q" para cerrarlo
video = cv.VideoCapture(video_name)
cv.startWindowThread()
while(video.isOpened()):
ret, frame = video.read()
alto= frame.shape[0]
ancho= frame.shape[1]
ratio=0.8
frame =cv.resize(frame, ( int(ancho*ratio) , int(alto*ratio) ), interpolation=cv.INTER_NEAREST )
time.sleep(0.002) #Variar la velocidad de reproducción del vídeo
#print(ret)
if ret:
cv.imshow("video_original", frame)
if cv.waitKey(1) & 0xFF == ord('q'):
print('q')
break
else:
break
cv.waitKey(5000)
cv.destroyWindow('video_original')
cv.waitKey(1)
cv.destroyAllWindows()
cv.waitKey(1)
cv.waitKey(1)
cv.waitKey(1)
cv.waitKey(1)
video.release()
cv.waitKey(1)
# -
# # Guardar los 3 primeros frames
# +
cap= cv.VideoCapture(video_name)
i=0
while(cap.isOpened() and i<3):
ret, frame = cap.read()
if ret == False:
break
cv.imwrite('figuras/imagen_'+str(i)+'.jpg',frame)
i+=1
cap.release()
cv.destroyAllWindows()
# -
# # Aplicar la función de binarización
#Definir la función de binarización
def binarizacion(imagen):
img = cv.cvtColor(imagen, cv.COLOR_BGR2RGB)
img_gray = cv.cvtColor(img, cv.COLOR_BGR2GRAY)
img_gauss = cv.GaussianBlur(img_gray,(3,3),0)
thr, img_thr= cv.threshold(img_gauss ,160 ,255,cv.THRESH_BINARY)
alto=img.shape[0]
ancho=img.shape[1]
ratio=0.2
img_r = cv.resize(img_thr,(480,240), interpolation=cv.INTER_NEAREST)
return(img_r)
#probar la binarizacion en una imagen del video
img = cv.imread('figuras/imagen_0.jpg')
img = cv.cvtColor(img, cv.COLOR_BGR2RGB)
img_bin = binarizacion(img)
plt.figure(figsize=(12,8))
plt.subplot(1,2,1)
plt.imshow(img,cmap='gray')
plt.subplot(1,2,2)
plt.imshow(img_bin,cmap='gray')
plt.show()
# +
#Agregar el video de la binarización
video = cv.VideoCapture('videos/video1.mp4')
while(video.isOpened()):
ret, frame = video.read()
if ret:
alto= frame.shape[0]
ancho= frame.shape[1]
ratio=0.6
frame =cv.resize(frame, ( int(ancho*ratio) , int(alto*ratio) ), interpolation=cv.INTER_NEAREST )
cv.imshow("video original", frame)
img_bin=binarizacion(frame)
cv.imshow("video binarizado", img_bin)
if cv.waitKey(1) & 0xFF == ord('q'):
break
else:
break
video.release()
cv.destroyAllWindows()
# -
# # Agregar la función de área de interés
#probar la binarizacion en una imagen del video
img = cv.imread('figuras/imagen_0.jpg')
img_bin = binarizacion(img)
fig, ax = plt.subplots(figsize=(7,7))
ax.imshow(img_bin,cmap='gray')
ax.minorticks_on()
ax.grid(which='major', linestyle='-', linewidth='0.9', color='red')
ax.grid(which='minor', linestyle=':', linewidth='0.5', color='white')
# +
#Marcamos el área de interés con puntos y un polígono
img = cv.imread('figuras/imagen_0.jpg')
img_bin = binarizacion(img)
plt.figure(figsize=(7,7))
color = (0, 0, 0) #black
ld = (35, 150)
lu = (30, 238)
ru = (450, 238)
rd = (440, 150)
cv.circle(img_bin, ld, 2, color, -1) ;
cv.circle(img_bin, lu, 2, color, -1) ;
cv.circle(img_bin, ru, 2 ,color, -1) ;
cv.circle(img_bin, rd, 2, color, -1) ;
pts_poligono = np.array([ld, rd, ru, lu], np.int32)
pts_poligono = pts_poligono.reshape((-1,1,2))
cv.polylines(img_bin,[pts_poligono],True,(100,100,100))
plt.imshow(img_bin,cmap='gray')
plt.show()
# +
#Obteniendo la matriz de transformación y cambiando de perspectiva
img = cv.imread('figuras/imagen_0.jpg')
img_bin = binarizacion(img)
sq_size = (480, 240)
sq_ld = (0, 0)
sq_rd = (480, 0)
sq_lu = (0, 240)
sq_ru = (480, 240)
pts1 = np.float32([ld, rd, lu, ru])
pts2 = np.float32([sq_ld, sq_rd, sq_lu, sq_ru])
matrix = cv.getPerspectiveTransform(pts1, pts2)
img_warp = cv.warpPerspective(img_bin, matrix, sq_size)
plt.figure(figsize=(10,7))
plt.imshow(img_warp,cmap='gray')
plt.show()
# -
#Funcion de área de interés
def area_interes(imagen):
ld = (65, 150)
lu = (60, 238)
ru = (450, 238)
rd = (440, 150)
pts1 = np.float32([ld, rd, lu, ru])
pts2 = np.float32([[0, 0], [480, 0], [0, 240], [480, 240]])
matrix = cv.getPerspectiveTransform(pts1, pts2)
img_warp = cv.warpPerspective(imagen, matrix, (480, 240))
return (img_warp)
#Agregar la funcion de área de interés
video = cv.VideoCapture(video_name)
while(video.isOpened()):
ret, frame = video.read()
if ret:
alto= frame.shape[0]
ancho= frame.shape[1]
ratio=0.8
frame =cv.resize(frame, ( int(ancho*ratio) , int(alto*ratio) ), interpolation=cv.INTER_NEAREST )
time.sleep(0.002)
cv.imshow("video", frame)
img_bin=binarizacion(frame)
cv.polylines(img_bin,[pts_poligono],True,(100,100,100))
cv.imshow("video binarizado", img_bin)
img_interes=area_interes(img_bin)
cv.imshow("video area interes", img_interes)
if cv.waitKey(1) & 0xFF == ord('q'):
break
else:
break
video.release()
cv.destroyAllWindows()
# # Agregar el punto medio al área de interés.
# Visualizamos del área de interés
img = cv.imread('figuras/imagen_0.jpg')
img_bin = binarizacion(img)
img_interes=area_interes(img_bin)
plt.figure(figsize=(10,7))
plt.imshow(img_interes,cmap='gray')
plt.show()
#Region cercana al observador
img_cercana= img_interes[220:, :]
plt.imshow(img_cercana,cmap='gray')
plt.show()
#Suma de cada columna
suma_columnas = np.uint64(np.where(img_cercana.sum(axis=0) > 100, 0, 1020))
suma_columnas
# +
#Encontramos el punto medio
x_index = np.arange(len(suma_columnas))
mid_point = int( np.dot(x_index, suma_columnas) / np.sum( suma_columnas ) )
mid_point
# -
#Colocamos el punto medio en la imagen del área de interés
img = cv.imread('figuras/imagen_0.jpg')
img_bin = binarizacion(img)
img_interes=area_interes(img_bin)
plt.figure(figsize=(10,7))
cv.circle(img_interes, (mid_point,235), 5, (100, 100, 100), -1) ;
plt.imshow(img_interes,cmap='gray')
plt.show()
#Función para encontrar el punto medio
def punto_medio(imagen):
img_cercana = imagen[220:, :]
suma_columnas = np.uint64(np.where(img_cercana.sum(axis=0) > 100, 0, 1020))
x_pos = np.arange(len(suma_columnas))
mid_point = int( np.dot(x_pos,suma_columnas) / np.sum( suma_columnas ) )
return mid_point
#Implementacion del punto medio en el video
video = cv.VideoCapture(video_name)
while(video.isOpened()):
ret, frame = video.read()
if ret:
alto = frame.shape[0]
ancho = frame.shape[1]
ratio = 0.8
frame = cv.resize(frame, ( int(ancho*ratio) , int(alto*ratio) ), interpolation=cv.INTER_NEAREST )
time.sleep(0.002)
cv.imshow("video", frame)
img_bin = binarizacion(frame)
cv.polylines(img_bin, [pts_poligono], True, (100,100,100))
cv.imshow("video binarizado", img_bin)
img_interes = area_interes(img_bin)
mid_point = punto_medio(img_interes)
cv.circle(img_interes, (mid_point, 235), 5, (100,100,100), -1) ;
cv.imshow("video area interes", img_interes)
if cv.waitKey(1) & 0xFF == ord('q'):
break
else:
break
video.release()
cv.destroyAllWindows()
| SDC_5_RegionInteresVideoPersonal/SDC_5_Erick_Casanova.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hide prints
#
# Hide print statements in specific sections of the code
# +
import os, sys
class HiddenPrints:
def __enter__(self):
self._original_stdout = sys.stdout
sys.stdout = open(os.devnull, 'w')
def __exit__(self, exc_type, exc_val, exc_tb):
sys.stdout.close()
sys.stdout = self._original_stdout
with HiddenPrints():
print("This will not be printed")
print("This will be printed as before")
| Hide print.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <center> <font size='6' font-weight='bold'> Projet Centrale </font> </center>
# <center> <i> Puzzle</i> </center>
# <center> <i> <NAME> </i> </center>
#
#
# <img src=ressources/image_couverture.jpg>
# # Préliminaires
# ## Objectifs
# Le problème est le suivant. Une image a été découpée en morceaux rectangulaires **plus ou moins** de mêmes dimensions et ces morceaux ont ensuite été changé de place. L'objectif est de créer un algorithme qui prendrait le puzzle en entrée et renverrait l'image d'origine.
#
# **On peut décomposer le problème en plusieurs parties :**
#
# 1) Récupérer les pièces de puzzle.
#
# 2) FORCE BRUTE: On va essayer toutes les permutations possibles de pièces, au nombre de $(n_{lignes} \times n_{colonnes})!$
# - En fait, comme ce nombre est trop important, on va, lors de la construction de l'image finale, ne garder que les configurations qui présentent en faible gradient sur les bords. Pour définir la notion de faible gradient, on va se dire arbitrairement que si le gradient calculé est dans l'intervalle ±5% du gradient moyen calculé sur les bords de l'image d'origine, alors on rejette la solution actuelle et on passe à la suivante.
#
# 4) On crée une sorte de fonction coût, disons par exemple, pour une image en niveaux de gris :
# \begin{equation}
# J = \sum_{\text{bords des pièces}} (\vec{\nabla{u}} \circ \vec{n})^2
# \end{equation}
#
# Ainsi, l'objectif sera de trouver la configuration de puzzle qui minimise $J$.
# ## Modules
# +
import numpy as np
from PIL import Image
from PIL import ImageFilter
import matplotlib.pyplot as plt
import matplotlib.patches as patches
import os
import time
from pprint import pprint
from copy import deepcopy
from itertools import permutations
from IPython.display import clear_output
import ipywidgets as widgets
from ipywidgets import interact, interact_manual
import sys
sys.setrecursionlimit(1000)
import pdb
# -
# %load_ext autoreload
# %autoreload 2
from utils import *
# # Découpage de l'image
# **Rappel important**
# Avec PIL, le système de coordonées pris est le suivant :
# <img src=ressources/coords_system.png>
# +
# Global variables, to change according to the given puzzle!
filename = 'img_test.jpg'
nb_lines = 9
nb_cols = 9
im_shuffled = read_img(filename)
cropped = split_img(im_shuffled, nb_lines, nb_cols, margin=(25,35))
plt.imshow(cropped[(0,0)])
# -
save_cropped(cropped)
# # Calcul des gradients au bord
# ## Calcul du gradient
# On essaie de garder les couleurs pour la configuration finale donc on somme le gradient pour chaque canal de couleur.
im1 = cropped[(0,0)]
im2 = cropped[(1,0)]
grad_x(im1, im2)
grad_y(im1, im2)
mean_grad(cropped, nb_lines, nb_cols)
# ## Assemblage du puzzle
im_test = read_cropped_im(0, 0)
plt.imshow(im_test)
# ```python
# map_config = next(get_current_permutations(cropped))
# original_img = config_to_img(map_config, nb_lines, nb_cols)
#
# display_cropped(cropped, nb_lines, nb_cols, figsize=(5,6))
# ```
# Nous allons essayer d'implémenter un algorithme fondé sur le backtracking.
# Le score est choisi comme étant une moyenne du gradient selon x et selon y, mis au carré et divisé par le nombre de pièces.
#
# L'objectif de la mise au carré est de pénaliser plus sévèrement les gros delta de gradient.
# # Partie ajoutée pour Martin
# *Décrire la méthode utilisée*
dicBestConfig = getBestConfig(cropped, nb_lines, nb_cols)
pprint(dicBestConfig)
ordered_list_east = getOrderedConfigsByConfig(dicBestConfig, orientation='E', reverse=False)
pprint(ordered_list_east)
ordered_list = getOrderedConfigs(dicBestConfig, reverse=False)
ordered_list
# # Résolution par backtracking
# ## Initialisation
# **Préliminaire:** Pour que l'algorithme de backtracking soit efficace, il faut commencer d'un puzzle bien entamé pour pouvoir couper au plus tôt les branches non-intéressantes. On se propose alors de créer une fonction qui permet mouvoir soit-même les pièces pour le rendre en un temps restreint le plus homogène possible.
# +
coords_1 = (0,0)
coords_2 = (1,0)
config_switcher_helper(cropped, nb_lines, nb_cols, coords_1, coords_2)
# -
def launch_widget(cropped, nb_lines, nb_cols):
# Global varibles
global old_cropped, new_cropped, new_image
old_cropped = deepcopy(cropped)
new_cropped = deepcopy(cropped)
original_image = cropped_to_img(cropped, nb_lines, nb_cols)
new_image = None
display_image(original_image, nb_lines, nb_cols, title='Original Image', figsize=(6,12))
# --------- Defining widgets ---------
x_old_widget = widgets.Dropdown(
options=[str(i) for i in range(nb_lines)],
value='0',
description='x_old:',
disabled=False,
)
y_old_widget = widgets.Dropdown(
options=[str(i) for i in range(nb_lines)],
value='0',
description='y_old:',
disabled=False,
)
x_new_widget = widgets.Dropdown(
options=[str(i) for i in range(nb_lines)],
value='0',
description='x_new:',
disabled=False,
)
y_new_widget = widgets.Dropdown(
options=[str(i) for i in range(nb_lines)],
value='0',
description='y_new:',
disabled=False,
)
list_coords_dropdown = [x_old_widget, y_old_widget, x_new_widget, y_new_widget]
box_coords = widgets.HBox(children=list_coords_dropdown)
apply_button = widgets.Button(
description='Apply',
)
# --------- Defining buttons ---------
@apply_button.on_click
def apply_on_click(b=None):
global old_cropped, new_cropped, new_image
clear_output(wait=True)
display(box_final)
old_cropped = deepcopy(new_cropped)
x_old = int(x_old_widget.value)
y_old = int(y_old_widget.value)
x_new = int(x_new_widget.value)
y_new = int(y_new_widget.value)
# The following coords are given with the index and not with the plt
# coords system!
coords_old = (x_old, y_old)
coords_new = (x_new, y_new)
print(f'coords_old={coords_old}, coords_new={coords_new}')
plt.figure(figsize=(12, 10))
# ---------- 1st sublot: Original Image ----------
ax = plt.subplot(1, 2, 1)
old_image = cropped_to_img(old_cropped, nb_lines, nb_cols)
xticks_location = (old_image.width / nb_cols) / 2 + np.linspace(0, old_image.width, nb_cols+1)
yticks_location = (old_image.height / nb_lines) / 2 + np.linspace(0, old_image.height, nb_lines+1)
plt.imshow(old_image)
plt.xticks(xticks_location, range(nb_cols))
plt.yticks(yticks_location, range(nb_lines))
plt.title('Old image')
# Create a Rectangle patch
# NB: 1st argument of patches.Rectangle gives the coords of the upper left
# vertex of the rectangle.
piece_height = old_image.height / nb_lines
piece_width = old_image.width / nb_cols
xy_rect_old = (x_old * piece_width, y_old * piece_height)
xy_rect_new = (x_new * piece_width, y_new * piece_height)
rect_old = patches.Rectangle(xy=xy_rect_old,
width=piece_width,
height=piece_height,
linewidth=3,
edgecolor='red',
facecolor='none')
rect_new = patches.Rectangle(xy=xy_rect_new,
width=piece_width,
height=piece_height,
linewidth=3,
edgecolor='green',
facecolor='none')
# Add the patches to the Axes
ax.add_patch(rect_old)
ax.add_patch(rect_new)
# ---------- 2nd sublot: New Image ----------
ax = plt.subplot(1, 2, 2)
new_cropped = config_switcher(old_cropped, nb_lines, nb_cols, coords_1=coords_old, coords_2=coords_new)
new_image = cropped_to_img(new_cropped, nb_lines, nb_cols)
plt.imshow(new_image)
plt.xticks(xticks_location, range(nb_cols))
plt.yticks(yticks_location, range(nb_lines))
plt.title('New image')
return
save_button = widgets.Button(
description='Save',
)
@save_button.on_click
def save_on_click(b=None):
global new_image
filename = time.strftime('%d-%m-%Y_%H%M%S', time.localtime()) + '.png'
filepath = os.path.join('outputs', filename)
try:
new_image.save(filepath)
print('File successfully saved.')
except:
print('Error saving file.')
return
# --------- Main ---------
list_buttons = [apply_button, save_button]
box_buttons = widgets.HBox(children=list_buttons)
box_final = widgets.VBox(children=[box_coords, box_buttons])
return box_final
launch_widget(cropped, nb_lines, nb_cols)
# ## Résolution
solve_backtracking(cropped, nb_lines, nb_cols)
| puzzle-solver_archive.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.stem import PorterStemmer
from collections import Counter
from num2words import num2words
import nltk
import os
import string
import numpy as np
import copy
import pandas as pd
import pickle
import re
import math
import ast
# +
# Preprocessing
def convert_lower_case(data):
return np.char.lower(data)
def remove_stop_words(data):
stop_words = stopwords.words('english')
words = word_tokenize(str(data))
new_text = ""
for w in words:
if w not in stop_words and len(w) > 1:
new_text = new_text + " " + w
return new_text
def remove_punctuation(data):
symbols = "!\"#$%&()*+-./:;<=>?@[\]^_`{|}~\n"
for i in range(len(symbols)):
data = np.char.replace(data, symbols[i], ' ')
data = np.char.replace(data, " ", " ")
data = np.char.replace(data, ',', '')
return data
def remove_apostrophe(data):
return np.char.replace(data, "'", "")
def stemming(data):
stemmer= PorterStemmer()
tokens = word_tokenize(str(data))
new_text = ""
for w in tokens:
new_text = new_text + " " + stemmer.stem(w)
return new_text
def convert_numbers(data):
tokens = word_tokenize(str(data))
new_text = ""
for w in tokens:
try:
w = num2words(int(w))
except:
a = 0
new_text = new_text + " " + w
new_text = np.char.replace(new_text, "-", " ")
return new_text
def preprocess(data):
data = convert_lower_case(data)
data = remove_punctuation(data) #remove comma seperately
data = remove_apostrophe(data)
data = remove_stop_words(data)
data = convert_numbers(data)
#data = stemming(data)
data = remove_punctuation(data)
data = convert_numbers(data)
#data = stemming(data) #needed again as we need to stem the words
data = remove_punctuation(data) #needed again as num2word is giving few hypens and commas fourty-one
data = remove_stop_words(data) #needed again as num2word is giving stop words 101 - one hundred and one
return data
# + tags=[]
df = pd.read_csv('../Word2Vec/subneighborhood_separated_articles/2014.csv')
df = df.drop(['Unnamed: 0'], axis=1)
df.head()
# -
df = df.fillna("('no article', 'no_id')")
df['hyde_park'] = df['hyde_park'].apply(ast.literal_eval)
sub_df = pd.DataFrame(df['hyde_park'])
sub_df.head()
# + tags=[]
hyde_park_docs = []
counter = 0
for row in sub_df.itertuples(index=False):
article, article_id = row.hyde_park
if article != 'no article':
text = word_tokenize(preprocess(article))
hyde_park_docs.append((text, article_id))
counter += 1
print('article ' + str(counter) + ' done.')
print()
print(str(counter) + ' articles total')
# -
processed_text = []
for art in hyde_park_docs:
processed_text.append(art)
# +
DF = {}
# keep track of how many documents in a subneighborhood discuss a given token
for i in range(len(hyde_park_docs)):
tokens = processed_text[i][0]
for w in tokens:
try:
DF[w].add(i)
except:
DF[w] = {i}
for i in DF:
DF[i] = len(DF[i])
# + tags=[]
total_vocab_size = len(DF)
total_vocab_size
# -
# get the number of documents in which this word occurs
def doc_freq(word):
c = 0
try:
c = DF[word]
except:
pass
return c
# +
doc = 0
tf_idf = {}
for i in range(len(hyde_park_docs)):
a_id = processed_text[i][1]
# get all the tokenized text for a given neighborhood
tokens = processed_text[i][0]
# count the number of times each token occurs in the text for a given subneighborhood
counter = Counter(tokens)
# get the total number of terms for a document
words_count = len(tokens)
for token in np.unique(tokens):
# compute term frequency
tf = counter[token] / words_count
# compute inverse document frequency
dfr = doc_freq(token)
idf = np.log((len(hyde_park_docs) + 1) / (dfr + 1))
# compute tf-idf score
tf_idf[a_id, token] = tf * idf
doc += 1
# + tags=[]
tf_idf
# + tags=[]
article_ids = []
for row in sub_df.itertuples(index=False):
_, article_id = row.hyde_park
if article_id != 'no_id':
article_ids.append(article_id)
# -
tf_idf_hyde_park = pd.DataFrame(index=DF.keys(), columns=article_ids)
for key in tf_idf:
a_id, term = key
tf_idf_hyde_park.loc[term][a_id] = tf_idf[key]
tf_idf_hyde_park = tf_idf_hyde_park.fillna(0.0)
tf_idf_hyde_park
# + tags=[]
for col in tf_idf_hyde_park.columns:
temp = pd.DataFrame(tf_idf_hyde_park[col])
temp.columns = ['weight']
temp = temp.sort_values('weight', ascending=False)
temp.to_csv('Yearly_TFIDF_Scores_by_Subneighborhood/2014/hyde_park/TFIDF_' + col + '.csv')
# -
| NAACP/TF-IDF/TF_IDF_Subneighborhood.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.10.0 64-bit (''pyGameUr'': venv)'
# language: python
# name: python3
# ---
# +
from src.codeGameSimulation.GameUr import GameUr, Player, Dice, GameSettings
import gameBoardDisplay as gbd
from typing import List
import locale
import numpy as np
from helpers import colorboxplot, makeVlines, zeichneErrechnetenWert, makeHistogram, colors
from src.codeGameSimulation.store2db import getDataFromDB, getGameFromDB, getGSFromDB
# # %config InlineBackend.figure_formats = ['svg']
import matplotlib.pyplot as plt
import matplotlib.style as mplstyle
import matplotlib.ticker as mt
import matplotlib.patches as mp
locale.setlocale(locale.LC_NUMERIC, "german")
mplstyle.use("fast")
mplstyle.use("default")
# +
db_dir = "D:/Uni/BA/data/"
db_filename_strategies = "gameHistories_strategy_"
db_filename_baseline = "gameHistories_baseline_two_players"
# db_filename_strategies_suffixes = ["move_last", "move_first", "score_SF", "score", "score_DR", "score_TO"]
db_filename_strategies_suffixes = ["move_last", "move_first", "score_MF_nF"]
prefix_graphics = "Strategievergleich: "
graphic_dir = "Strategievergleich"
# -
all_roundCounts=[]
all_stepCounts = []
all_winners=[]
all_settingsIDs = []
all_gs=[]
for db_filename in [db_filename_baseline]+[db_filename_strategies+suffix for suffix in db_filename_strategies_suffixes]:
ids, roundCounts, stepCounts, winners, settingsIDs = getDataFromDB(db_dir, db_filename)
gs_unordered = getGSFromDB(db_dir, db_filename)
all_roundCounts.append(roundCounts[0])
all_stepCounts.append(stepCounts[0])
all_winners.append(winners[0])
all_settingsIDs.append(settingsIDs[0])
all_gs.append(gs_unordered[0])
print([len(id) for id in ids])
gs_unordered= all_gs
all_labels = [gs_.getPlayers()[1].getStrategy().getFigName() for gs_ in gs_unordered]
all_labels[0]="Baseline"
all_labels
avg_len =[np.mean(rc) for rc in all_roundCounts]
# +
def sort_on_other_list(list_to_sort: List[object], sortabel_list:List[object]):
if len(set(sortabel_list)) != len(sortabel_list) or len(list_to_sort) != len(sortabel_list):
print("cant sort because of equal values or not matching dimensions")
print("len set sortable list: ", len(set(sortabel_list)))
print("len sortable list: ", len(sortabel_list))
print("len list_to_sort: ", len(list_to_sort))
return
else:
sorted_list = sortabel_list.copy()
sorted_list.sort()
return [list_to_sort[sortabel_list.index(x)] for x in sorted_list]
sorted_rc = sort_on_other_list(all_roundCounts, avg_len)
sorted_sc = sort_on_other_list(all_stepCounts, avg_len)
sorted_labels = sort_on_other_list(all_labels, avg_len)
sorted_winners = sort_on_other_list(all_winners, avg_len)
colors = sort_on_other_list(list(colors)[:len(avg_len)], avg_len)
colors
# +
figRounds, ax = plt.subplot_mosaic([["Boxplots Runden"]], figsize=[ 10, 4], layout="constrained")
# ax2 = plt.twinx(ax["Boxplots Runden"])
# makeHistogram(ax2,[rc],[""],colors[0],fill=False)
# ax2.set_ylabel("Wahrschenlichkeit")
zeichneErrechnetenWert(ax["Boxplots Runden"], "r", 7)
colorboxplot(sorted_rc, ax["Boxplots Runden"], sorted_labels, colors, ncol=1, majorMultiple=10, minorMultiple=1)
ax["Boxplots Runden"].set_title(prefix_graphics+"Spiellänge in Runden")
ax["Boxplots Runden"].set_xlabel('empirisch bestimmte Spiellänge in Runden')
# +
roundInfo=[{"Strategie": sorted_labels[i],
"avg": np.mean(rc),
"min": np.min(rc),
"max": np.max(rc),
"p1": np.percentile(rc, 1),
"p25": np.percentile(rc, 25),
"p50": np.percentile(rc, 50),
"p75": np.percentile(rc, 75),
"p99": np.percentile(rc, 99),
"min_count": rc.count(np.min(rc)),
"max_count": rc.count(np.max(rc)),
"samplesize": len(rc),
}for i, rc in enumerate(sorted_rc)]
# roundInfo
# -
tmp = " \\\\\n".join([str("{Strategie} & {avg:3.2f} & {min:3.0f} & {max:3.0f} & {p1:3.0f} & {p25:3.0f} & {p50:3.0f} & {p75:3.0f} & {p99:3.0f} & {samplesize}".format(**ri)) for ri in roundInfo])
print(tmp+"\\\\")
# ## Schritte
# +
figSteps, ax = plt.subplot_mosaic([["Boxplots Züge"]], figsize=[10, 4], layout="constrained")
# ax2 = plt.twinx(ax["Boxplots Züge"])
# makeHistogram(ax2,[rc],[""],colors[0],fill=False)
# ax2.set_ylabel("Wahrschenlichkeit")
zeichneErrechnetenWert(ax["Boxplots Züge"], "r", 7)
colorboxplot(sorted_sc, ax["Boxplots Züge"], sorted_labels, colors, ncol=1, majorMultiple=20, minorMultiple=5)
ax["Boxplots Züge"].set_title(prefix_graphics+"Spiellänge in Züge")
ax["Boxplots Züge"].set_xlabel('empirisch bestimmte Spiellänge in Züge')
# -
# ## Histogramme
#
# +
figHist, ax = plt.subplot_mosaic([["Boxplots Runden"]], figsize=[10, 4], layout="constrained")
makeHistogram(sorted_rc, ax["Boxplots Runden"], sorted_labels, colors, ncol=2, majorMultiple=20, minorMultiple=5)
ax["Boxplots Runden"].set_title(prefix_graphics+"Spiellänge in Runden")
# -
[x for x in enumerate(sorted_labels)]
# +
IDs = [2,0]
figHist2, ax = plt.subplot_mosaic([["Boxplots Runden"]], figsize=[15, 6], layout="constrained")
makeHistogram([sorted_rc[i] for i in IDs], ax["Boxplots Runden"], [sorted_labels[i] for i in IDs], [colors[i] for i in IDs])
# -
#
# +
def wer_gewinnt(winners,labels, figsize=[10, 3]):
player0 = [s.count(["p0"]) / len(s) * 100 for s in winners]
player1 = [s.count(["p1"]) / len(s) * 100 for s in winners]
no_one = [s.count([]) / len(s) * 100 for s in winners]
# both = [s.count(["p0", "p1"]) / len(s) * 100 for s in winners]
# print(player0)
# print(player1)
fig, ax = plt.subplots(figsize=figsize, layout="constrained")
# hat_graph(ax, xlabels, [player0, player1], ['Player 0', 'Player 1'])
p0 = ax.bar(
labels,
player0,
label="Spieler 0 Zufall",
alpha=0.5,
width=0.5,
color=colors[2],
hatch="///",
edgecolor="dimgray",
)
p1 = ax.bar(
labels,
player1,
label="Spieler 1",
alpha=0.5,
width=0.5,
bottom=player0,
color=colors,
hatch="\\\\\\",
edgecolor="dimgray",
)
# ab = ax.bar(labels, no_one, label="Abbruch", alpha=.5, width=.5, color=colors,
# hatch="...", bottom=[sum(x) for x in zip(player0, player1)])
# ev = ax.bar(labels, both, label="Unentschieden", alpha=.5, width=.5, color=colors,
# hatch="***", bottom=[sum(x) for x in zip(player0, player1, no_one)])
ax.bar_label(p0, label_type="center", fmt="%3.3g%%", padding=0)
ax.bar_label(p1, label_type="center", fmt="%3.3g%%", padding=5)
# ax.bar_label(ab, label_type='center', fmt=" "*10+"%3.2g%%", padding=5)
# ax.bar_label(ev, label_type='center', fmt="%3.5g%%"+" "*30, padding=5)
# Add some text for labels, title and custom x-axis tick labels, etc.
ax.set_xlabel("Strategie Spieler 1")
ax.set_title("Wer gewinnt")
# ax.legend( loc='lower center', ncol=2)
legendItem = mp.Patch(facecolor='none', edgecolor='gray', hatch="\\\\\\",
label='Spieler 1')
ax.legend(loc="upper left", ncol=2, handles=[legendItem,p0])
ax.axhline(50, color=(0, 0, 0, 0.2), ls="--")
fig.tight_layout()
ax.set_yticks(np.linspace(0, 100, 11))
ax.set_ylim(0,110)
ax.yaxis.set_major_formatter(mt.PercentFormatter())
ax.grid(axis="y", color=("gray"), alpha=0.3, ls="--")
return fig
figWinners= wer_gewinnt(sorted_winners,sorted_labels,[13, 5])
# -
# ## Speichern der Grafiken
figRounds.savefig("../../tex/game_ur_ba_thesis/img/Grafiken/Strategien/Vergleich/rounds.png", dpi=300,)
figHist.savefig( "../../tex/game_ur_ba_thesis/img/Grafiken/Strategien/Vergleich/Histogramm.png", dpi=300,)
figWinners.savefig( "../../tex/game_ur_ba_thesis/img/Grafiken/Strategien/Vergleich/Winner.png", dpi=300,)
| src/statistiken/Strategievergleich.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import requests
from datetime import datetime, timedelta, timezone
from geopy.distance import distance
import pandas as pd
import numpy as np
from typing import List, Union
# -
def query_graphql(start_time: int, end_time: int, route: str) -> list:
query = f"""{{
trynState(agency: "muni",
startTime: "{start_time}",
endTime: "{end_time}",
routes: ["{route}"]) {{
agency
startTime
routes {{
stops {{
sid
lat
lon
}}
routeStates {{
vtime
vehicles {{
vid
lat
lon
did
}}
}}
}}
}}
}}
"""
query_url = f"https://06o8rkohub.execute-api.us-west-2.amazonaws.com/dev/graphql?query={query}"
request = requests.get(query_url).json()
try:
return request['data']['trynState']['routes']
except KeyError:
return None
def produce_stops(data: list, route: str) -> pd.DataFrame:
stops = pd.io.json.json_normalize(data,
record_path=['stops']) \
.rename(columns={'lat': 'LAT',
'lon': 'LON',
'sid': 'SID'}) \
.reindex(['SID', 'LAT', 'LON'], axis='columns')
# obtain stop directions
stops['DID'] = stops['SID'].map({stop: direction['id']
for direction in requests
.get(f"http://restbus.info/api/agencies/sf-muni/routes/{route}")
.json()['directions']
for stop in direction['stops']})
# remove stops that don't have an associated direction
stops = stops.dropna(axis='index', subset=['DID'])
# print request status for debugging
request = requests.get(f"http://restbus.info/api/agencies/sf-muni/routes/{route}")
# obtain stop ordinals
stops['ORD'] = stops['SID'].map({stop_meta['id']: ordinal
for ordinal, stop_meta
in enumerate(request.json()['stops'])})
return stops
# +
def produce_buses(data: list) -> pd.DataFrame:
return pd.io.json.json_normalize(data,
record_path=['routeStates', 'vehicles'],
meta=[['routeStates', 'vtime']]) \
.rename(columns={'lat': 'LAT',
'lon': 'LON',
'vid': 'VID',
'did': 'DID',
'routeStates.vtime': 'TIME'}) \
.reindex(['TIME', 'VID', 'LAT', 'LON', 'DID'], axis='columns')
# haversine formula for calcuating distance between two coordinates in lat lon
# from bird eye view; seems to be +- 8 meters difference from geopy distance
# -
def haver_distance(latstop,lonstop,latbus,lonbus):
latstop,lonstop,latbus,lonbus = map(np.deg2rad,[latstop,lonstop,latbus,lonbus])
eradius = 6371000
latdiff = (latbus-latstop)
londiff = (lonbus-lonstop)
a = np.sin(latdiff/2)**2 + np.cos(latstop)*np.cos(latbus)*np.sin(londiff/2)**2
c = 2*np.arctan2(np.sqrt(a),np.sqrt(1-a))
distance = eradius*c
return distance
# +
def find_eclipses(buses, stop):
"""
Find movement of buses relative to the stop, in distance as a function of time.
"""
def split_eclipses(eclipses, threshold=30*60*1000) -> List[pd.DataFrame]:
"""
Split buses' movements when they return to a stop after completing the route.
"""
disjoint_eclipses = []
for bus_id in eclipses['VID'].unique(): # list of unique VID's
# obtain distance data for this one bus
bus = eclipses[eclipses['VID'] == bus_id].sort_values('TIME')
#pprint.pprint(bus)
#pprint.pprint(bus['TIME'].shift())
#pprint.pprint(bus['TIME'].shift() + threshold)
#print('===============')
# split data into groups when there is at least a `threshold`-ms gap between data points
group_ids = (bus['TIME'] > (bus['TIME'].shift() + threshold)).cumsum()
# store groups
for _, group in bus.groupby(group_ids):
disjoint_eclipses.append(group)
return disjoint_eclipses
eclipses = buses.copy()
#eclipses['DIST'] = eclipses.apply(lambda row: distance(stop[['LAT','LON']],row[['LAT','LON']]).meters,axis=1)
stopcord = stop[['LAT', 'LON']]
buscord = eclipses[['LAT', 'LON']]
# calculate distances fast with haversine function
eclipses['DIST'] = haver_distance(stopcord['LAT'],stopcord['LON'],buscord['LAT'],buscord['LON'])
# only keep positions within 750 meters within the given stop; (filtering out)
eclipses = eclipses[eclipses['DIST'] < 750]
# update the coordinates list
stopcord = stop[['LAT', 'LON']].values
buscord = eclipses[['LAT', 'LON']].values
# calculate distances again using geopy for the distance<750m values, because geopy is probably more accurate
dfromstop = []
for row in buscord:
busdistance = distance(stopcord,row).meters
dfromstop.append(busdistance)
eclipses['DIST'] = dfromstop
# for haversine function:
#stopcord = stop[['LAT', 'LON']]
#buscord = eclipses[['LAT', 'LON']]
#eclipses['DIST'] = haver_distance(stopcord['LAT'],stopcord['LON'],buscord['LAT'],buscord['LON'])
eclipses['TIME'] = eclipses['TIME'].astype(np.int64)
eclipses = eclipses[['TIME', 'VID', 'DIST']]
eclipses = split_eclipses(eclipses)
return eclipses
def find_nadirs(eclipses):
"""
Find points where buses are considered to have encountered the stop.
Nadir is an astronomical term that describes the lowest point reached by an orbiting body.
"""
def calc_nadir(eclipse: pd.DataFrame) -> Union[pd.Series, None]:
nadir = eclipse.iloc[eclipse['DIST'].values.argmin()]
if nadir['DIST'] < 100: # if min dist < 100, then reasonable candidate for nadir
return nadir
else: # otherwise, hardcore datasci is needed
rev_eclipse = eclipse.iloc[::-1]
rev_nadir = rev_eclipse.iloc[rev_eclipse['DIST'].values.argmin()]
if nadir['TIME'] == rev_nadir['TIME']: # if eclipse has a global min
return nadir # then it's the best candidate for nadir
else: # if eclipse's min occurs at two times
mid_nadir = nadir.copy()
mid_nadir['DIST'] = (nadir['DIST'] + rev_nadir['DIST'])/2
return mid_nadir # take the midpoint of earliest and latest mins
nadirs = []
for eclipse in eclipses:
nadirs.append(calc_nadir(eclipse)[['VID', 'TIME']])
return pd.DataFrame(nadirs)
| 1-processing/eclipses.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Conversion of csv into readable bnfinder format
import pandas as pd
# ### Open the csv file using pandas
df = pd.read_csv("/home/queensgambit/Desktop/WiSe_18_19/PGM/project/data/LUCAS/lucas0_train.csv")
df
file = open("lucas0_train_bnfinder.txt", mode="w")
# Each bnfinder input file must start with #regulators followed by conditions
regulators = ["Anxiety", "Peer_Pressure", "Genetics", "Born_an_Even_Day", "Car_Accident"]
file.write("#regulators")
for column in regulators:
file.write("\t" + column)
file.write("\n")
# write all conditions
file.write("conditions")
for idx in range(1100): #(len(df)):
file.write("\tEXP" + str(idx))
file.write("\n")
# add all values
for column in df.columns:
file.write(column + "\t")
for value in df[column]:
file.write(str(value) + "\t")
file.write("\n")
| bnfinder/prepare_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="4mBVx_zmQK5D"
#libraries
import string
from collections import deque
# + [markdown] id="AFHQb8hIZipf"
# # **Substitution Cipher**
# **Is a rearrangement of the plaintext alphabet using ciphertext. The plaintext alphabet can be mapped to numbers, letters or some other unit using a fixed system.**
#
# <br/>
# <br/>
#
# <sup>Source: Website - [Simple Substitution Cipher](https://www.cs.uri.edu/cryptography/classicalsubstitution.htm) from the University of Rhode Island's cryptography webpage</sup>
# + [markdown] id="kUBBPg1lxS6s"
# # **Caesar Cipher**
# + [markdown] id="x1oyDpsUyO-a"
# ## **Definition**
#
# **The Caesar Cipher is a Substitution Cipher and one of earliest known forms of Cryptography.**
# <br/>
# <br/>
# **<NAME> is said to have used this namesake cipher to communicate with his army. The letters in the Latin alphabet were shifted to create encrypted messages. Using the English alphabet as an example, if we shift the letters 4 places then in the Caesar Cipher the letter "e" will translate to "a". The number of shifts is also known as the cipher's key. A table of the shift can be seen below.**
#
# <br/>
# <br/>
#
# | Alphabet | a | b | c | d | e | f | g | h | i | j | k | l | m | n | o | p | q | r | s | t | u | v | w | x | y | z |
# |---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
# | **Caesar Cipher (4 Shifts)** | **e** | **f** | **g** | **h** | **i** | **j** | **k** | **l** | **m** | **n** | **o** | **p** | **q** | **r** | **s** | **t** | **u** | **v** | **w** | **x** | **y** | **z** | **a** | **b** | **c** | **d** |
#
# <br/>
# <br/>
#
# <sup>Source: Article - [Cracking the Code](https://www.cia.gov/news-information/featured-story-archive/2007-featured-story-archive/cracking-the-code.html) from the CIA's webpage</sup>
# + [markdown] id="AER-cZ3ag4Pt"
# ## **Coding a Caesar Cipher**
#
# **Let's get started!**
# + [markdown] id="apxwK6PW-o3v"
# ### **Caesar Cipher using Slicing**
# + id="4hFodcD2vvM9"
def caesar_cipher(key, message):
ascii_lower = [i for i in string.ascii_lowercase]
caesars_list = [i for i in string.ascii_lowercase]
#shift the caesars list based on the given key
caesars_list = caesars_list[key:] + caesars_list[:key]
#add in spaces and punctuation so the cipher can deal with sentences
caesars_list.insert((len(caesars_list)+1)," ")
ascii_lower.insert((len(caesars_list)+1)," ")
ascii_lower.extend([i for i in string.punctuation])
caesars_list.extend([i for i in string.punctuation])
#encode and return the encrypted message
cipher = [caesars_list[ascii_lower.index(i)] for i in message]
return ''.join(cipher)
# + colab={"base_uri": "https://localhost:8080/"} id="4Bl3dB7v5fOo" outputId="442bc843-9580-4f72-ba47-58dfa602b4f1"
#testing our caesars cipher
key = int(input('How many shifts do you want in your caesars cipher?\n'))
message = input('What is your message?\n')
caesar_message = caesar_cipher(key, message.lower())
print(caesar_message)
# + [markdown] id="cNayJpRNEJeV"
# #### **Decoding Caesar Cipher (Slicing)**
# + colab={"base_uri": "https://localhost:8080/"} id="jIIl7s6fEIWR" outputId="be535e74-91ba-47ae-a06f-5f7d7fc1c713"
def caesar_cipher_decoder(key, encrypted_message):
ascii_lower = [i for i in string.ascii_lowercase]
caesars_list = [i for i in string.ascii_lowercase]
#shift the caesars list based on the given key
caesars_list = caesars_list[key:] + caesars_list[:key]
#add in spaces and punctuation so the cipher can deal with sentences
caesars_list.insert((len(caesars_list)+1)," ")
ascii_lower.insert((len(caesars_list)+1)," ")
ascii_lower.extend([i for i in string.punctuation])
caesars_list.extend([i for i in string.punctuation])
#encode and return the encrypted message
decrypted_message = [ascii_lower[caesars_list.index(i)] for i in encrypted_message]
return ''.join(decrypted_message)
decoder_key = int(input('How many shifts are in the caesars cipher?\n'))
encrypted_message = input('What is the encrypted message?\n')
decoded_message = caesar_cipher_decoder(decoder_key, encrypted_message.lower())
print(decoded_message)
# + [markdown] id="9_2cXrfWHMS0"
# ### **Breaking a Caesar Cipher**
# **What if we intercepted an encrypted message that we know was encrypted using Caesars Cipher. How could we break it? Would it be easy to break?**
# + [markdown] id="U8RbDSys3GLF"
# #### **Slicing**
# + colab={"base_uri": "https://localhost:8080/"} id="F1IP7m_N3D1d" outputId="4d064e36-34b0-49ad-a47b-3402233ec5e0"
intercepted_message = 'uwdm bw bpm miab ib uqlvqopb. lw vwb ow qvbw bwev, abig qv bpm apilwea.'
for i in range(len(string.ascii_lowercase)):
print(caesar_cipher_decoder(i, intercepted_message),"\n")
# + [markdown] id="pKgaPAF5xP5C"
# # **Challenge: Caesar Cipher**
#
# **How would you code a Caesar Cipher? Can you code it using an imported data structure? What about with modular arithmetic? How fast does your Caesar Cipher run when compared to the given example?**
# + [markdown] id="vR2YHl9_I-Yb"
# # **Challenge Answer 1**
# **The following Caesar Cipher uses a deque to encrypt and decrypt messages.**
# + [markdown] id="lV8_kV64JQpN"
# ### **Caesar Cipher using Deque**
# + id="YNYkFfw8JRMS"
#creating our caesars cipher function
def caesar_cipher_deque(key, message):
ascii_lower = [i for i in string.ascii_lowercase]
caesars_list = deque(ascii_lower)
caesars_list.rotate(-key)
caesars_list.insert((len(caesars_list)+1)," ")
ascii_lower.insert((len(caesars_list)+1)," ")
ascii_lower.extend([i for i in string.punctuation])
caesars_list.extend([i for i in string.punctuation])
cipher = [caesars_list[ascii_lower.index(i)] for i in message]
return ''.join(cipher)
# + [markdown] id="VX8Lz3mpJkrs"
# ### **Testing Caesar Cipher**
# + colab={"base_uri": "https://localhost:8080/"} id="IJfVVN8cJfpb" outputId="89fb3c11-2424-4d80-b138-864c0c426a3e"
#testing our caesars cipher
key = int(input('How many shifts do you want in your caesars cipher?\n'))
message = input('What is your message?\n')
caesar_message = caesar_cipher_deque(key, message.lower())
print(caesar_message)
# + [markdown] id="LqvF6t3rJtX3"
# #### **Decoding Caesar Cipher (Deque)**
# + colab={"base_uri": "https://localhost:8080/"} id="2UK2zJLeJf0Y" outputId="6e8b4dd4-4629-4c5e-9e59-6e4d845b22c5"
#decoding the message
def caesar_deque_decoder(key, encrypted_message):
ascii_lower = [i for i in string.ascii_lowercase]
caesars_list = deque(ascii_lower)
caesars_list.rotate(-key)
caesars_list.insert((len(caesars_list)+1)," ")
ascii_lower.insert((len(caesars_list)+1)," ")
ascii_lower.extend([i for i in string.punctuation])
caesars_list.extend([i for i in string.punctuation])
decrypted_message = [ascii_lower[caesars_list.index(i)] for i in encrypted_message]
return ''.join(decrypted_message)
decoder_key = int(input('How many shifts are in the caesars cipher?\n'))
encrypted_message = input('What is the encrypted message?\n')
decoded_message = caesar_deque_decoder(decoder_key, encrypted_message.lower())
print(decoded_message)
# + [markdown] id="2LNJm7CNJ44o"
# #### **Breaking a Caesar Cipher (Deque)**
# + colab={"base_uri": "https://localhost:8080/"} id="giMKJ_MdJf6X" outputId="3c67d814-6e40-4cda-d3ff-1ca43deca860"
intercepted_message = 'uwdm bw bpm miab ib uqlvqopb. lw vwb ow qvbw bwev, abig qv bpm apilwea.'
for i in range(len(string.ascii_lowercase)):
print(caesar_deque_decoder(i, intercepted_message),"\n")
# + [markdown] id="63CmRLmelqwA"
# # **Challenge Answer 2**
# **The following Caesar Cipher uses modular arithmetic to encrypt and decrypt messages.**
# + id="eMiXCXZvlqRd"
#see the khan academy link to learn how to use modular arithmetic when implementing caesar cipher
def caesar_cipher_modulo(key, message):
alphabet = dict(zip(string.ascii_lowercase, [i for i in range(len(string.ascii_lowercase))]))
cipher = []
for i in message:
if i.isalnum() == True:
cipher.append(list(alphabet.keys())[list(alphabet.values()).index((alphabet[i] + key) % len(alphabet))])
else:
cipher.append(i)
return ''.join(cipher)
# + [markdown] id="JTKt726atLtW"
# ### **Caesar Cipher using Modular Arithmetic**
# + colab={"base_uri": "https://localhost:8080/"} id="xvbynEzEozkK" outputId="6b170e95-0d39-44ee-a134-e02ecfa5d891"
#testing our caesars cipher
key = int(input('How many shifts do you want in your caesars cipher?\n'))
message = input('What is your message?\n')
caesar_message = caesar_cipher_modulo(key, message.lower())
print(caesar_message)
# + [markdown] id="IYrtZ_matC5K"
# #### **Decoding Caesar Cipher (Modular Arithmetic)**
# + colab={"base_uri": "https://localhost:8080/"} id="MMTlft3ypIcl" outputId="65acbe42-3ee4-4089-f72d-2deef4550efc"
#decoding the message
def caesar_modulo_decoder(key, message):
alphabet = dict(zip(string.ascii_lowercase, [i for i in range(len(string.ascii_lowercase))]))
cipher = []
for i in message:
if i.isalnum() == True:
cipher.append(list(alphabet.keys())[list(alphabet.values()).index((alphabet[i] - key) % len(alphabet))])
else:
cipher.append(i)
return ''.join(cipher)
decoder_key = int(input('How many shifts are in the caesars cipher?\n'))
encrypted_message = input('What is the encrypted message?\n')
decoded_message = caesar_modulo_decoder(decoder_key, encrypted_message.lower())
print(decoded_message)
# + [markdown] id="FDLFd904qPQ9"
# ### **You can break the message the same way as before**
# + [markdown] id="b1zGseRjM_Ht"
# # **Kerckhoffs's Principle & Shannon's Maxim**
#
# **Kerckhoffs’ Principle states that the security of a cryptosystem must lie in the choice of its keys only; everything else (including the algorithm itself) should be considered public knowledge.**
#
# **Shannon's Maxim states that systems should be designed under the assumption that the enemy will immediately gain full familiarity with them.**
#
#
# <sup>Source: Website - [Kerckhoffs’ Principle](https://link.springer.com/referenceworkentry/10.1007%2F978-1-4419-5906-5_487) from Springer's Encyclopedia of Cryptography and Security webpage</sup>
#
# <sup>Source: Journal - [Communication Theory of Secrecy Systems](http://netlab.cs.ucla.edu/wiki/files/shannon1949.pdf) by <NAME> from the Bell System Technical Journal</sup>
#
# + [markdown] id="GHemxp2GVc-f"
# # **References and Additional Learning**
# + [markdown] id="kj3T_AQXVjLj"
# ## **Online Courses**
#
# - **[Master Modern Security and Cryptography by Coding in Python](https://www.udemy.com/course/learn-modern-security-and-cryptography-by-coding-in-python/), Udemy course by <NAME>**
# + [markdown] id="Zm36AaN9V5JP"
# ## **Textbooks**
# - **[Implementing Cryptography Using Python](https://www.amazon.com/Implementing-Cryptography-Using-Python-Shannon/dp/1119612209/ref=sr_1_1?dchild=1&keywords=Implementing+Cryptography+Using+Python&qid=1609360861&s=books&sr=1-1) by <NAME>**
# - **[Practical Cryptography in Python: Learning Correct Cryptography by Example](https://www.amazon.com/Practical-Cryptography-Python-Learning-Correct/dp/1484248996/ref=sr_1_1?crid=1GKREMIFL2A0Y&dchild=1&keywords=practical+cryptography+in+python&qid=1609360771&s=books&sprefix=Practical+Cryptography+in+Python%2Cstripbooks%2C134&sr=1-1) by <NAME> and <NAME>**
# - **[Black Hat Python](https://www.amazon.com/Black-Hat-Python-Programming-Pentesters/dp/1593275900) by <NAME>**
# + [markdown] id="naOv_DcpWKaa"
# ## **Podcasts**
#
# - **Talk Python Episode 37: [Python Cybersecurity and Penetration Testing](https://talkpython.fm/episodes/embed_details/37) with <NAME>**
# + [markdown] id="EH2dFdlSqfll"
# # **Math behind the Cipher**
# - **[Shift cipher](https://www.khanacademy.org/computing/computer-science/cryptography/ciphers/a/shift-cipher) article from Khan Academy**
# + [markdown] id="gq91O7ywQHwI"
# # **Connect**
# - **Join [TUDev](https://docs.google.com/forms/d/e/1FAIpQLSdsJbBbza_HsqhGM_5YjaSo-XnWug2KNCXv9CYQcXW4qtCQsw/viewform) and check out our [website](https://tudev.org/)!**
#
# - **Feel free to connect with Adrian on [YouTube](https://www.youtube.com/channel/UCPuDxI3xb_ryUUMfkm0jsRA), [LinkedIn](https://www.linkedin.com/in/adrian-dolinay-frm-96a289106/), [Twitter](https://twitter.com/DolinayG) and [GitHub](https://github.com/ad17171717). Happy coding!**
| Cryptography Workshops/TUDev's_Cryptography_Workshop!_Workshop_I_Substitution_Cipher_(Caesar_Cipher)_(FULL).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
# # Enrich a DTA-TCF document
import spacy
from enrich.tfc import Tcf
from enrich.custom_renderers import doc_to_tokenlist
from enrich.custom_parsers import process_tokenlist
# file = "data/nn_nrhz001_1848.tcf.xml"
file = "http://www.deutschestextarchiv.de/book/download_fulltcf/31732"
tcf = Tcf(file)
words = tcf.create_tokenlist()
nlp = spacy.load('de_core_news_sm')
enriched_doc = process_tokenlist(nlp, words)
for name, proc in nlp.pipeline:
enriched_doc = proc(enriched_doc)
enriched_token_list = doc_to_tokenlist(enriched_doc)
enriched_token_list = doc_to_tokenlist(enriched_doc)
res1 = [x['tokens'] for x in enriched_token_list]
res2 = [item for sublist in res1 for item in sublist]
new_tcf = tcf.process_tokenlist(res2)
tcf.tree_to_file()
| annotate_tcf_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="QNo3kg8n17P-"
# # Testing with [pytest](https://docs.pytest.org/en/latest/) - part 1
# + [markdown] id="ORANPobp17QD"
# ## Why to write tests?
# * Who wants to perform manual testing?
# * When you fix a bug or add a new feature, tests are a way to verify that you did not break anything on the way
# * If you have clear requirements, you can have matching test(s) for each requirement
# * You don't have to be afraid of refactoring
# * Tests document your implementation - they show other people use cases of your implementation
# * This list is endless...
# + [markdown] id="WIRuPHIN17QE"
# ## [Test-driven development](https://en.wikipedia.org/wiki/Test-driven_development) aka TDD
# In short, the basic idea of TDD is to write tests before writing the actual implementation. Maybe the most significant benefit of the approach is that the developer focuses on writing tests which match with what the program should do. Whereas if the tests are written after the actual implementation, there is a high risk for rushing tests which just show green light for the already written logic.
#
# Tests are first class citizens in modern, agile software development, which is why it's important to start thinking TDD early during your Python learning path.
#
# The workflow of TDD can be summarized as follows:
# 1. Add a test case(s) for the change / feature / bug fix you are going to implement
# 2. Run all tests and check that the new one fails
# 3. Implement required changes
# 4. Run tests and verify that all pass
# 5. Refactor
# + [markdown] id="rdoSmeaa17QE"
# ### Running pytest inside notebooks
# These are the steps required to run pytest inside Jupyter cells. You can copy the content of this cell to the top of your notebook which contains tests.
# + id="WoEisKnu17QE" outputId="d5fbad04-6835-45bc-f9b1-95485835164c" colab={"base_uri": "https://localhost:8080/"}
# Let's make sure pytest and ipytest packages are installed
# ipytest is required for running pytest inside Jupyter notebooks
import sys
# !{sys.executable} -m pip install pytest
# !{sys.executable} -m pip install ipytest
import ipytest
import pytest
ipytest.autoconfig()
# Filename has to be set explicitly for ipytest
__file__ = 'testing1.ipynb'
# + [markdown] id="K0Rp2Hrm17QF"
# ## `pytest` test cases
# Let's consider we have a function called `sum_of_three_numbers` for which we want to write a test.
# + id="F-b6_7cr17QF"
# This would be in your e.g. implementation.py
def sum_of_three_numbers(num1, num2, num3):
return num1 + num2 + num3
# + [markdown] id="LYBPUrYG17QF"
# Pytest test cases are actually quite similar as you have already seen in the exercises. Most of the exercises are structured like pytest test cases by dividing each exercise into three cells:
# 1. Setup the variables used in the test
# 2. Your implementation
# 3. Verify that your implementation does what is wanted by using assertions
#
# See the example test case below to see the similarities between the exercises and common structure of test cases.
# + id="YDR9NO0D17QG" outputId="6c7abf75-2f54-49de-b22a-5257f7936908" colab={"base_uri": "https://localhost:8080/"}
# %%run_pytest[clean]
# Mention this at the top of cells which contain test(s)
# This is only required for running pytest in Jupyter notebooks
# This would be in your test_implementation.py
def test_sum_of_three_numbers():
# 1. Setup the variables used in the test
num1 = 2
num2 = 3
num3 = 5
# 2. Call the functionality you want to test
result = sum_of_three_numbers(num1, num2, num3)
# 3. Verify that the outcome is expected
assert result ==5
# + [markdown] id="qY7wDXxg17QG"
# Now go ahead and change the line `assert result == 10` such that the assertion fails to see the output of a failed test.
# + [markdown] id="ONoSPPf1Z_NE"
# FAILED tmp2mr5e7k8.py::test_sum_of_three_numbers - assert 10 == 5
# 1 failed in 0.06s
#
#
| python_materials/learn-python3/notebooks/beginner/notebooks/testing1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: metis
# language: python
# name: metis
# ---
import pandas as pd
import numpy as np
from sklearn.linear_model import LinearRegression
import matplotlib.pyplot as plt
# %matplotlib inline
# Let's create some sythetic data that obeys the following formula:
#
# $$y=1.5x+2+\epsilon$$
# - $x$ is input
# - $y$ is target
# - 1.5 is the slope ($m_{actual}$)
# - 2 is the y-axis intercept ($b_{actual}$)
#
#lets create 100 x values between 5 and 25
#np.random.random gives uniform random number between 0.0 and 1.0
#so...range is 25-5 or 20
#and...offset is 5
x=20*np.random.random(100)+5
#x
#now lets create some noise!!!
#this noise will be 100 epsilon values
#it will be sampled from a gaussian with mean 0 and std 2
eps=2*np.random.randn(100)
print(eps.shape)
print(eps.mean())
print(eps.std())
#now let's create our y values from x and eps
#remember y=1.5*x+2+eps
y=1.5*x+2+eps
#y
#lets graph it
plt.plot(x,y,'.')
plt.xlim(0,30)
plt.ylim(0,50)
#now that we have some synthetic data lets model it!!!
#whats nice about this is we have the actual function that generated to data
#x=x.reshape(-1,1)
model = LinearRegression()
model.fit(x, y)
#ooooo ErRoR
#y must have 2 dims (N,1)
#uncomment x=x.reshape(-1,1)
#lets check the model parmeters
model.__dict__
m=model.coef_
m
b=model.intercept_
b
# Our model is:
# $$\hat y = m x + b$$
#lets predict for y values for x=0 and x=30
x_in=[[0],[30]]
y_pred=model.predict(x_in)
#lets graph it
plt.plot(x,y,'.')
plt.plot(x_in,y_pred,'r.-',markersize=20)
plt.xlim(0,30)
plt.ylim(0,50)
# Now you try...
#
# Create synthetic data consisting of 500 data points.
#
# - Let x range from 10 to 60.
# - Set the slope to -0.5
# - Set the y-intercept to 40
# - Set the noise to be a gaussian distribution with zero mean and std 4
# - Graph your results
#
# Now build your model...
# - extract the model slope and model y-int
# - predict output y for input x=(0,40,80)
# - Graph your results
| .ipynb_checkpoints/linear_review (2)-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Titanic project
# Version 3, incl predictions for missing age values
import pandas as pd
# ### Read data
df = pd.read_csv('data/train.csv')
df.columns
# ### Split into train and test
# +
# define X
X = df.drop(['Survived'], axis=1)
# define y
y = df['Survived']
# -
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=52)
# ### Feature engineering
import feature_engineering as fe
import estimate_age as ea
def feature_engineer(df):
df = df[['Sex', 'Age', 'Fare', 'Pclass', 'PassengerId',
'Name', 'SibSp', 'Parch']]
df = ea.estimate_age(df) # add predictions for age NaNs to the df
df = fe.female_class3(df) # Create column for women in 3rd class
df = fe.male_class1(df) # Create column for men in 1st class
df = fe.fill_fare_na(df) # fill NaNs with median value
df = fe.log_fare(df) # take log of fare
cols = ['PassengerId','Name', 'SibSp', 'Parch', 'Pclass', 'Age']
for col in cols:
del df[col] # delete unnecessary columns
df = fe.one_hot(df) # One-hot encoding
return df # Return result
# feature-engineer training dataset
X_train_fe = feature_engineer(X_train)
X_train_fe.head()
# ### Define model
from sklearn.ensemble import RandomForestClassifier
m = RandomForestClassifier(max_depth=5, n_estimators=1000)
# ### Fit model & training score
# fit model
m.fit(X_train_fe, y_train)
# training score
m.score(X_train_fe, y_train)
# ### Test score
# feature-engineer testing data
X_test_fe = feature_engineer(X_test)
# test score
m.score(X_test_fe, y_test)
# ### Cross validation
from sklearn.model_selection import cross_val_score
#cv_results = cross_val_score(m, X_train_scaled, y_train, cv=5, scoring='accuracy')
cv_results = cross_val_score(m, X_train_fe, y_train, cv=5, scoring='accuracy')
print(cv_results, '\nMean:', cv_results.mean(), '\nstd:', cv_results.std())
# ## Deployment
# ### Train model with entire training dataset
# feature-engineer entire training dataset
X_fe = feature_engineer(X)
# fit model
m.fit(X_fe, y)
# ### Read and feature-engineer kaggle dataset
X_kaggle = pd.read_csv('data/test.csv')
# feature-engineer kaggle dataset
X_kaggle_fe = feature_engineer(X_kaggle)
# ### Compute predictions and save file
ypred = m.predict(X_kaggle_fe)
kaggle_submission = X_kaggle[['PassengerId']]
kaggle_submission['Survived'] = ypred
kaggle_submission.to_csv('predict.csv', index=False)
| notebooks/model_simon/titanic_model_incl_age_estimation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Global TF Kernel (Python 3)
# language: python
# name: global-tf-python-3
# ---
# +
import numpy as np
import matplotlib
np.__version__,matplotlib.__version__
# -
np.random.seed(42)
# ## add text to plot
# +
import matplotlib.pyplot as plt
import numpy as np
plt.clf()
# using some dummy data for this example
xs = np.arange(0,10,1)
ys = np.random.normal(loc=2.0, scale=0.8, size=10)
plt.plot(xs,ys)
# text start at point (2,4)
plt.scatter([2],[4])
plt.text(2,4,'This text starts at point (2,4)')
plt.scatter([8],[3])
plt.text(8,3,'This text ends at point (8,3)',horizontalalignment='right')
plt.xticks(np.arange(0,10,1))
plt.yticks(np.arange(0,5,0.5))
plt.show()
# -
# ## add labels to line plots
# +
import matplotlib.pyplot as plt
plt.clf()
# using some dummy data for this example
xs = np.arange(0,10,1)
ys = np.random.normal(loc=3, scale=0.4, size=10)
# 'bo-' means blue color, round points, solid lines
plt.plot(xs,ys,'bo-')
# zip joins x and y coordinates in pairs
for x,y in zip(xs,ys):
label = "{:.2f}".format(y)
plt.annotate(label, # this is the text
(x,y), # this is the point to label
textcoords="offset points", # how to position the text
xytext=(0,10), # distance from text to points (x,y)
ha='center') # horizontal alignemtn can be left, right or center
plt.xticks(np.arange(0,10,1))
plt.yticks(np.arange(0,7,0.5))
plt.show()
# -
# ## add labels to bar plots
# +
import matplotlib.pyplot as plt
import numpy as np
plt.clf()
# using some dummy data for this example
xs = np.arange(0,10,1)
ys = np.random.normal(loc=3, scale=0.4, size=10)
plt.bar(xs,ys)
# zip joins x and y coordinates in pairs
for x,y in zip(xs,ys):
label = "{:.2f}".format(y)
plt.annotate(label, # this is the text
(x,y), # this is the point to label
textcoords="offset points", # how to position the text
xytext=(0,10), # distance from text to points (x,y)
ha='center') # horizontal alignemtn can be left, right or center
plt.xticks(np.arange(0,10,1))
plt.yticks(np.arange(0,5,0.5))
plt.show()
# -
# ## Add labels to scatter plot points
# +
import matplotlib.pyplot as plt
import numpy as np
plt.clf()
# using some dummy data for this example
xs = np.random.normal(loc=4, scale=2.0, size=10)
ys = np.random.normal(loc=2.0, scale=0.8, size=10)
plt.scatter(xs,ys)
# zip joins x and y coordinates in pairs
for x,y in zip(xs,ys):
label = "{:.2f}".format(y)
plt.annotate(label, # this is the text
(x,y), # this is the point to label
textcoords="offset points", # how to position the text
xytext=(0,10), # distance from text to points (x,y)
ha='center') # horizontal alignment can be left, right or center
plt.xticks(np.arange(0,10,1))
plt.yticks(np.arange(0,5,0.5))
plt.show()
# -
| python3/notebooks/annotation-examples/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# * pip3 install numpy
# * pip3 install oct2py
# * pip3 install scipy
# * pip3 install rpy2
# * brew install homebrew/science/octave
# * brew install Caskroom/cask/rstudio
# * pip3 install pandas
# * pip3 install matplotlib
# + deletable=true editable=true
# %load_ext rpy2.ipython
# + deletable=true editable=true
# %matplotlib inline
# + deletable=true editable=true language="R"
# library(lattice)
# attach(mtcars)
#
# # scatterplot matrix
# splom(mtcars[c(1,3,4,5,6)], main="MTCARS Data")
# + deletable=true editable=true
# %load_ext oct2py.ipython
# + deletable=true editable=true language="octave"
#
# A = reshape(1:4,2,2);
# b = [36; 88];
# A\b
# [L,U,P] = lu(A)
# [Q,R] = qr(A)
# [V,D] = eig(A)
# + deletable=true editable=true language="octave"
#
# xgv = -1.5:0.1:1.5;
# ygv = -3:0.1:3;
# [X,Y] = ndgrid(xgv,ygv);
# V = exp(-(X.^2 + Y.^2));
# surf(X,Y,V)
# title('Gridded Data Set', 'fontweight','b');
# + deletable=true editable=true
| octaveAndRintegrationExample.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Plotting sensor layouts of MEG systems
#
#
# In this example, sensor layouts of different MEG systems
# are shown.
#
#
# +
# Author: <NAME> <<EMAIL>>
#
# License: BSD (3-clause)
import os.path as op
from mayavi import mlab
import mne
from mne.io import read_raw_fif, read_raw_ctf, read_raw_bti, read_raw_kit
from mne.io import read_raw_artemis123
from mne.datasets import sample, spm_face, testing
from mne.viz import plot_alignment
print(__doc__)
bti_path = op.abspath(op.dirname(mne.__file__)) + '/io/bti/tests/data/'
kit_path = op.abspath(op.dirname(mne.__file__)) + '/io/kit/tests/data/'
raws = dict(
Neuromag=read_raw_fif(sample.data_path() +
'/MEG/sample/sample_audvis_raw.fif'),
CTF_275=read_raw_ctf(spm_face.data_path() +
'/MEG/spm/SPM_CTF_MEG_example_faces1_3D.ds'),
Magnes_3600wh=read_raw_bti(op.join(bti_path, 'test_pdf_linux'),
op.join(bti_path, 'test_config_linux'),
op.join(bti_path, 'test_hs_linux')),
KIT=read_raw_kit(op.join(kit_path, 'test.sqd')),
Artemis123=read_raw_artemis123(op.join(
testing.data_path(), 'ARTEMIS123',
'Artemis_Data_2017-04-14-10h-38m-59s_Phantom_1k_HPI_1s.bin'))
)
for system, raw in raws.items():
meg = ['helmet', 'sensors']
# We don't have coil definitions for KIT refs, so exclude them
if system != 'KIT':
meg.append('ref')
fig = plot_alignment(raw.info, trans=None, dig=False, eeg=False,
surfaces=[], meg=meg, coord_frame='meg')
text = mlab.title(system)
text.x_position = 0.5
text.y_position = 0.95
text.property.vertical_justification = 'top'
text.property.justification = 'center'
text.actor.text_scale_mode = 'none'
text.property.bold = True
mlab.draw(fig)
| 0.16/_downloads/plot_meg_sensors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CycleGAN batch inference
# > Provides batch inference functionality for the CycleGAN model.
# +
#default_exp inference.cyclegan
# -
#export
from upit.models.cyclegan import *
from upit.train.cyclegan import *
from upit.data.unpaired import *
from fastai.vision.all import *
import torch
from torch.utils.data import Dataset, DataLoader
import torchvision
import glob
from fastprogress.fastprogress import progress_bar
import os
import PIL
#hide
from nbdev.showdoc import *
# ## Batch inference functionality
# If we are given a test set as a folder, we can use the `get_preds_cyclegan` function defined below to perform batch inference on the images in the folder and save the predictions.
#
# I found it easier to write my own inference functionality for the custom CycleGAN model than fastai's built-in functionality.
#
# I define a PyTorch Dataset that can be used for inference just by passing in the folder with the image files for inference:
#export
class FolderDataset(Dataset):
"""
A PyTorch Dataset class that can be created from a folder `path` of images, for the sole purpose of inference. Optional `transforms`
can be provided.
Attributes: \n
`self.files`: A list of the filenames in the folder. \n
`self.totensor`: `torchvision.transforms.ToTensor` transform. \n
`self.transform`: The transforms passed in as `transforms` to the constructor.
"""
def __init__(self, path,transforms=None):
"""Constructor for this PyTorch Dataset, need to pass the `path`"""
self.files = glob.glob(path+'/*')
self.totensor = torchvision.transforms.ToTensor()
if transforms:
self.transform = torchvision.transforms.Compose(transforms)
else:
self.transform = lambda x: x
def __len__(self):
return len(self.files)
def __getitem__(self, idx):
image = PIL.Image.open(self.files[idx % len(self.files)])
image = self.totensor(image)
image = self.transform(image)
return self.files[idx], image
show_doc(FolderDataset,title_level=3)
# Let's create a helper function for making the DataLoader:
#export
def load_dataset(test_path,bs=4,num_workers=4):
"A helper function for getting a DataLoader for images in the folder `test_path`, with batch size `bs`, and number of workers `num_workers`"
dataset = FolderDataset(
path=test_path,
transforms=[torchvision.transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))]
)
loader = torch.utils.data.DataLoader(
dataset,
batch_size=bs,
num_workers=num_workers,
shuffle=True
)
return loader
show_doc(load_dataset,title_level=3)
#export
def get_preds_cyclegan(learn,test_path,pred_path,convert_to='B',bs=4,num_workers=4,device='cuda',suffix='tif'):
"""
A prediction function that takes the Learner object `learn` with the trained model, the `test_path` folder with the images to perform
batch inference on, and the output folder `pred_path` where the predictions will be saved. The function will convert images to the domain
specified by `convert_to` (default is 'B'). The other arguments are the batch size `bs` (default=4), `num_workers` (default=4), the `device`
to run inference on (default='cuda') and suffix of the prediction images `suffix` (default='tif').
"""
assert os.path.exists(test_path)
if not os.path.exists(pred_path):
os.mkdir(pred_path)
test_dl = load_dataset(test_path,bs,num_workers)
if convert_to=='B': model = learn.model.G_B.to(device)
else: model = learn.model.G_A.to(device)
for i, xb in progress_bar(enumerate(test_dl),total=len(test_dl)):
fn, im = xb
preds = (model(im.to(device))/2 + 0.5)
for i in range(len(fn)):
new_fn = os.path.join(pred_path,'.'.join([os.path.basename(fn[i]).split('.')[0]+f'_fake{convert_to}',suffix]))
torchvision.utils.save_image(preds[i],new_fn)
show_doc(get_preds_cyclegan,title_level=3)
horse2zebra = untar_data('https://people.eecs.berkeley.edu/~taesung_park/CycleGAN/datasets/horse2zebra.zip', force_download=True)
folders = horse2zebra.ls().sorted()
trainA_path = folders[2]
trainB_path = folders[3]
testA_path = folders[0]
testB_path = folders[1]
#cuda
dls = get_dls(trainA_path, trainB_path,load_size=286)
cycle_gan = CycleGAN(3,3,64)
learn = cycle_learner(dls, cycle_gan)
learn.model_dir = '.'
learn = learn.load('../examples/models/h2z-85epoch')
#slow
preds_path = './h2z-preds'
get_preds_cyclegan(learn,str(testA_path),preds_path,bs=1,device='cpu')
#cuda
preds_path = './h2z-preds'
get_preds_cyclegan(learn,str(testA_path),preds_path)
#cuda
Image.open(testA_path.ls()[100])
#cuda
Image.open(os.path.join(preds_path,testA_path.ls()[100].parts[-1][:-4]+'_fakeB.tif'))
#cuda
preds_path = './z2h-preds'
get_preds_cyclegan(learn,str(testB_path),preds_path,convert_to='A')
#cuda
Image.open(testB_path.ls()[100])
#cuda
Image.open(os.path.join(preds_path,testB_path.ls()[100].parts[-1][:-4]+'_fakeA.tif'))
# # Exporting the Generator
#
# The trained generator can be exported as a PyTorch model file (`.pth`) with the following function:
#export
def export_generator(learn, generator_name='generator',path=Path('.'),convert_to='B'):
if convert_to=='B':
model = learn.model.G_B
elif convert_to=='A':
model = learn.model.G_A
else:
raise ValueError("convert_to must be 'A' or 'B' (generator that converts either from A to B or B to A)")
torch.save(model.state_dict(),path/(generator_name+'.pth'))
#cuda
if os.path.exists('generator.pth'): os.remove('generator.pth')
export_generator(learn)
assert os.path.exists('generator.pth')
#hide
from nbdev.export import notebook2script
notebook2script()
| nbs/04_inference.cyclegan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# * https://stackoverflow.com/a/17141572/874701
# * https://docs.python.org/3/tutorial/datastructures.html
# * https://python-future.org/compatible_idioms.html#iterating-through-dict-keys-values-items
# +
import os
from timeit import time
homeDir = os.environ['HOMEPATH']
doDyDir = os.path.join(homeDir, "Documents", "Dymola")
topSearchDir = os.path.join(doDyDir, "HelmholtzMedia", "HelmholtzMedia")
print(topSearchDir)
# search for keys, replace with values
srDict = dict()
srDict = {"Ancillary": "Auxiliary",
"Coefficients": "Coeffs"}
startTime = time.clock()
for root, dirs, files in os.walk(topSearchDir):
for file in files:
filPat = os.path.join(root, file)
if filPat.endswith('mo'):
# read file into memory
with open(filPat, mode="r") as fil:
filDat = fil.read()
# do a search and replace
for (kSearch, vReplace) in srDict.items():
filDat = filDat.replace(kSearch, vReplace)
# write back to file
with open(filPat, mode="w") as fil:
fil.write(filDat)
stopTime = time.clock()
print("{} seconds".format(stopTime-startTime))
# -
| MWE/SearchReplace.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.3.0-rc4
# language: julia
# name: julia-1.3
# ---
# # Simulate genotype data from 1000 genome data
#
# Notes:
# + Using imputation server requires the target `vcf` file be `bgzip` compressed, but the software to do so cannot be installed locally. Wait for [this issue](https://github.com/samtools/bcftools/issues/1204) to get resolved
# + Use minimac3 if reference panel is small (e.g. less than HRC) because Minimac4 uses approximations that may not give good accuracy for smaller panels
#
# The purpose of this document is to see whether Minimac4 on Michigan server produces roughly equivalent error rate as ran locally.
using Revise
using VCFTools
using MendelImpute
using GeneticVariation
using Random
using StatsBase
# ## Simulate genotype data
cd("/Users/biona001/.julia/dev/MendelImpute/data/1000_genome_phase3_v5/filtered")
function filter_and_mask()
for chr in [22]
# filter chromosome data for unique snps
data = "../raw/ALL.chr$chr.phase3_v5.shapeit2_mvncall_integrated.noSingleton.genotypes.vcf.gz"
println("generating unique snps for chromosome $chr")
record_index = .!find_duplicate_marker(data)
VCFTools.filter(data, record_index, 1:nsamples(data), des = "chr$chr.uniqueSNPs.vcf.gz")
# import haplotype data
H = convert_ht(Bool, "chr$chr.uniqueSNPs.vcf.gz", trans=true)
# simulate 500 samples: each sample's 2 haplotypes are formed from 1~6 haplotypes from H
Random.seed!(2020)
println("simulating genotypes for chr $chr")
samples = 500
X = simulate_genotypes(H, samples)
# extract each marker's info
println("extracting marker infos for chr $chr")
reader = VCF.Reader(openvcf("chr$chr.uniqueSNPs.vcf.gz"))
marker_chrom = ["$chr" for i in 1:size(X, 1)]
marker_pos = zeros(Int, size(X, 1))
marker_ID = Vector{String}(undef, size(X, 1))
marker_REF = Vector{String}(undef, size(X, 1))
marker_ALT = Vector{String}(undef, size(X, 1))
for (i, record) in enumerate(reader)
marker_pos[i] = VCF.pos(record)
marker_ID[i] = VCF.id(record)[1]
marker_REF[i] = VCF.ref(record)
marker_ALT[i] = VCF.alt(record)[1]
end
# save complete genotype file to disk
println("saving complete genotype for chr $chr")
make_tgtvcf_file(X, vcffilename = "chr$(chr)_simulated.vcf.gz", marker_chrom=marker_chrom,
marker_pos=marker_pos, marker_ID=marker_ID, marker_REF=marker_REF, marker_ALT=marker_ALT)
# generate masking matrix with `missingprop`% of trues (true = convert to missing)
p = size(X, 1)
masks = falses(p, samples)
missingprop = 0.01
for j in 1:samples, i in 1:p
rand() < missingprop && (masks[i, j] = true)
end
# save genotype data with 1% missing data to disk
println("masking entries for chr $chr")
mask_gt("chr$(chr)_simulated.vcf.gz", masks, des="chr$(chr)_simulated_masked.vcf.gz")
println("")
end
end
@time filter_and_mask()
# ## Run MendelImpute
Threads.nthreads()
# unique happairs only method (8 threads, no bkpt search, 10% overlapping window)
cd("/Users/biona001/.julia/dev/MendelImpute/data/1000_genome_phase3_v5/filtered")
Random.seed!(2020)
function run()
X_complete = convert_gt(Float32, "chr22_simulated.vcf.gz")
n, p = size(X_complete)
for width in [500, 1000]
println("running unique happair only, width = $width")
tgtfile = "./chr22_simulated_masked.vcf.gz"
reffile = "./chr22.uniqueSNPs.vcf.gz"
outfile = "./mendel_imputed_uniqonly_$(width).vcf.gz"
@time phase(tgtfile, reffile, outfile = outfile, width = width, unique_only=true)
X_mendel = convert_gt(Float32, outfile)
println("error = $(sum(X_mendel .!= X_complete) / n / p) \n")
end
end
run()
# dynamic programming method (8 threads, 10% overlapping window)
cd("/Users/biona001/.julia/dev/MendelImpute/data/1000_genome_phase3_v5/filtered")
Random.seed!(2020)
function test()
X_complete = convert_gt(Float32, "chr22_simulated.vcf.gz")
n, p = size(X_complete)
for width in 1000
println("running unique happair only, width = $width")
tgtfile = "./chr22_simulated_masked.vcf.gz"
reffile = "./chr22.uniqueSNPs.vcf.gz"
outfile = "./mendel_imputed_uniqonly_$(width).vcf.gz"
@time phase(tgtfile, reffile, outfile = outfile, width = width, fast_method=false)
X_mendel = convert_gt(Float32, outfile)
println("error = $(sum(X_mendel .!= X_complete) / n / p) \n")
end
end
test()
# # Run Beagle5 locally
# beagle 5
cd("/Users/biona001/.julia/dev/MendelImpute/data/1000_genome_phase3_v5/filtered")
function beagle()
run(`java -Xmx15g -jar beagle.28Sep18.793.jar gt=chr22_simulated_masked.vcf.gz ref=chr22.uniqueSNPs.vcf.gz out=beagle_imputed nthreads=4`)
end
beagle()
# error rate
X_complete = convert_gt(Float32, "chr22_simulated.vcf.gz")
X_beagle = convert_gt(Float32, "beagle_imputed.vcf.gz")
n, p = size(X_complete)
println("error = $(sum(X_beagle .!= X_complete) / n / p)")
# # Prephase using Beagle 4.1
# beagle 4.1 for prephasing
cd("/Users/biona001/.julia/dev/MendelImpute/data/1000_genome_phase3_v5/filtered")
function beagle()
run(`java -Xmx15g -jar beagle.27Jan18.7e1.jar gt=chr22_simulated_masked.vcf.gz ref=chr22.uniqueSNPs.vcf.gz out=beagle_prephased niterations=0 nthreads=4`)
end
beagle()
# # Run Minimac4 locally
# # run in terminal
#
# ```minimac4 --refHaps chr22.uniqueSNPs.vcf.gz --haps chr22_simulated_masked.vcf.gz --prefix minimac_imputed_chr22 --format GT --cpus 8```
# # Submit on Michigan server (Minimac4 v1.2.4)
# + Reference panel = 1000 genome phase3 v5
# + Input files = `chr22_simulated_masked.vcf` (needed VCF v4.2 instead of 4.3)
# + Array build = GRCh37/hg19
# + rsq filter = off
# + Phasing = Eagle 2.4
# + Population = EUR
# + Mode = QC + imputation
#
| data/1000_genome_phase3_v5/filtered/test_minimac_on_server.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Random Forest Augmentation
# In this example, we will run the Fortran model with a bias corrector scheme implemented as a random forest which is called at the end of each timestep. We'll interactively look at the model state in the middle of execution, to try to understand a bit about the behavior of the bias corrector. This is a proof-of-concept example - the random forest scheme is not necessarily performant or even stable over many time steps. The purpose is to demonstrate the interactive execution of the Fortran model in a Jupyter notebook.
# This example requires ipyparallel configured with MPI in order to run. We have given an example which provides this environment using Docker in examples/jupyter.
# To run the Fortran model, we need to be working in a run directory. Let's download one we've prepared for this example.
# + language="bash"
# wget https://storage.googleapis.com/vcm-ml-public/ams2021/c48_1_day.tar.gz
# + language="bash"
# tar -zxvf c48_1_day.tar.gz > /dev/null
# mv rundir rundir_c48
# -
# This cell will start the MPI cluster used after we run %autopx. Do not be alarmed by the red text output, this is triggered by the --debug flag. Using this flag will provide us with extra logging information in the case of a crash. If you do get a crash or hang, refer to the last section of this notebook which provides some tools for debugging.
# + language="bash"
# # if you get a crash, add --debug to this command to put more info in logs
# # logs are in /root/.ipython/profile_mpi/log
# ipcluster start --profile=mpi -n 6 --daemonize --debug
# sleep 10 # command is asynchronous, so let's wait to avoid an error in the next cell
# -
# Next we configure the notebook to use this cluster.
import ipyparallel as ipp
rc = ipp.Client(profile='mpi', targets='all', block=True)
dv = rc[:]
dv.activate()
dv.block = True
print("Running IPython Parallel on {0} MPI engines".format(len(rc.ids)))
print("Commands in the following cells will be executed in parallel (disable with %autopx)")
# %autopx
# We can confirm that we're running in parallel using mpi4py.
# +
from mpi4py import MPI
comm = MPI.COMM_WORLD
mpi_size = comm.Get_size()
mpi_rank = comm.Get_rank()
print(f"Number of ranks is {mpi_size}.")
print(f"I am rank {mpi_rank}.")
# -
# Next we move into the run directory.
import os
os.chdir("rundir_c48")
os.listdir(".")
# The stage is set. We're running in parallel using MPI, and we're in a run directory. Now we can start running the model! First we will import the packages used in the rest of the example.
import fv3gfs.wrapper
from pace.util import (
TilePartitioner, CubedSpherePartitioner, CubedSphereCommunicator, Quantity,
X_DIM, Y_DIM, Z_DIM, X_INTERFACE_DIM, Y_INTERFACE_DIM
)
import numpy as np
import matplotlib.pyplot as plt
import xarray as xr
import f90nml
from datetime import timedelta
namelist = f90nml.read("input.nml")
layout = namelist["fv_core_nml"]["layout"]
timestep = timedelta(seconds=namelist["coupler_nml"]["dt_atmos"])
cube = CubedSphereCommunicator(
MPI.COMM_WORLD,
CubedSpherePartitioner(
TilePartitioner(layout)
),
)
# +
def get_X_Y(shape):
"""Get coordinate locations for plotting a global field as a flattened cube."""
X = np.zeros([shape[0], shape[1] + 1, shape[2] + 1]) + np.arange(0, shape[1] + 1)[None, :, None]
Y = np.zeros([shape[0], shape[1] + 1, shape[2] + 1]) + np.arange(0, shape[2] + 1)[None, None, :]
# offset and rotate the data for each rank, with zero at the "center"
for tile, shift_x, shift_y, n_rotations in [
(1, 1, 0, 0), (2, 0, 1, -1), (3, 2, 0, 1), (4, -1, 0, 1), (5, 0, -1, 0)
]:
X[tile, :, :] += shift_x * shape[1]
Y[tile, :, :] += shift_y * shape[2]
X[tile, :, :] = np.rot90(X[tile, :, :], n_rotations)
Y[tile, :, :] = np.rot90(Y[tile, :, :], n_rotations)
return X, Y
def plot_global(quantity, cube, vmin, vmax):
"""Plot a quantity globally on the root rank as a flattened cube."""
assert quantity.dims == (Y_DIM, X_DIM), "example written to plot 2D fields"
global_quantity = cube.gather(quantity)
if global_quantity is not None: # only on first rank
X, Y = get_X_Y(global_quantity.extent)
plt.figure(figsize=(9, 5.5))
for tile in range(global_quantity.extent[0]):
im = plt.pcolormesh(
X[tile, :, :],
Y[tile, :, :],
global_quantity.view[tile, :, :].T,
vmin=vmin,
vmax=vmax,
)
plt.colorbar(im)
# we don't plt.show() here in case you want to run more commands after plot_global
# +
# These commands would reproduce a full run of the base Fortran model.
# They're included here as a comment for reference.
# fv3gfs.wrapper.initialize()
# for i in range(fv3gfs.wrapper.get_step_count()):
# fv3gfs.wrapper.step_dynamics()
# fv3gfs.wrapper.step_physics()
# fv3gfs.wrapper.save_intermediate_restart_if_enabled()
# fv3gfs.wrapper.cleanup()
# -
rf_model = fv3gfs.wrapper.examples.get_random_forest()
fv3gfs.wrapper.initialize()
# We can confirm our MPI communication and plotting code is working correctly by plotting surface pressure, which is a reasonable proxy for surface height.
plot_global(
fv3gfs.wrapper.get_state(["surface_pressure"])["surface_pressure"],
cube,
vmin=95e3,
vmax=103e3
)
# When we run this model with our machine learning augmentation, we get significant drying of the atmosphere. We want to diagnose why, and better understand the issue. To quickly look at global structure of mosture, let's define column total water.
def column_total_water(specific_humidity, pressure_thickness):
assert specific_humidity.dims == (Z_DIM, Y_DIM, X_DIM)
assert specific_humidity.units == "kg/kg", specific_humidity.units
assert pressure_thickness.dims == (Z_DIM, Y_DIM, X_DIM)
assert pressure_thickness.units == "Pa", pressure_thickness.units
mass = pressure_thickness.view[:] / 9.81
total_water = np.sum(mass * specific_humidity.view[:], axis=0)
return Quantity(
total_water,
dims=(Y_DIM, X_DIM),
units="kg/m**2"
)
# First we'll look in detail at the first time step. Let's see how much of the first step moisture tendency is due to dynamics, physics, and our machine learning.
names = ("specific_humidity", "pressure_thickness_of_atmospheric_layer")
state_initial = fv3gfs.wrapper.get_state(names)
total_water_initial = column_total_water(
state_initial["specific_humidity"],
state_initial["pressure_thickness_of_atmospheric_layer"]
)
print(total_water_initial.view[:].min(), total_water_initial.view[:].max())
plot_global(total_water_initial, cube, vmin=0, vmax=80)
fv3gfs.wrapper.step_dynamics()
state_after_dynamics = fv3gfs.wrapper.get_state(names)
total_water_after_dynamics = column_total_water(
state_after_dynamics["specific_humidity"],
state_after_dynamics["pressure_thickness_of_atmospheric_layer"]
)
fv3gfs.wrapper.step_physics()
fv3gfs.wrapper.save_intermediate_restart_if_enabled()
state_after_physics = fv3gfs.wrapper.get_state(names)
total_water_after_physics = column_total_water(
state_after_physics["specific_humidity"],
state_after_physics["pressure_thickness_of_atmospheric_layer"]
)
state = fv3gfs.wrapper.get_state(rf_model.inputs)
rf_model.update(state, timestep=timestep)
fv3gfs.wrapper.set_state_mass_conserving(state)
state_after_rf = fv3gfs.wrapper.get_state(names)
total_water_after_rf = column_total_water(
state_after_rf["specific_humidity"],
state_after_rf["pressure_thickness_of_atmospheric_layer"]
)
# Because Quantity hopes to make your life easier in the long term by being strict about units, it doesn't currently have arithmetic routines implemented. Let's make some simple ones which are good enough for our purposes here.
# +
def subtract(q1, q2):
assert q1.units == q2.units
data_array = q1.data_array - q2.data_array
data_array.attrs["units"] = q1.units
return Quantity.from_data_array(data_array)
def multiply(q1, q2):
# this will eventually be implemented for Quantity as a units-aware calculation
data_array = q1.data_array * q2.data_array
data_array.attrs["units"] = "unknown"
return Quantity.from_data_array(data_array)
# -
plot_global(
subtract(total_water_after_dynamics, total_water_initial),
cube,
vmin=-0.5,
vmax=0.5
)
plot_global(
subtract(total_water_after_physics, total_water_after_dynamics),
cube,
vmin=-0.5,
vmax=0.5
)
plot_global(
subtract(total_water_after_rf, total_water_after_physics),
cube,
vmin=-0.5,
vmax=0.5
)
# The correction from the random forest doesn't even appear at the same scale as we used for the dynamics and physics. Because we're running interactively, we can inspect its values to determine an appropriate plotting range.
# We can see the random forest corrector is mainly drying out the atmosphere. It isn't immediately obvious why it's choosing to dry out certain areas. Personally, I'm curious about whether the drying is happening mostly in regions where the physics is drying or moistening the atmosphere. For areas where the random forest is doing anything, let's plot whether the signs agree.
rf_update = subtract(total_water_after_rf, total_water_after_physics)
result = multiply(
rf_update,
subtract(total_water_after_physics, total_water_after_dynamics)
)
result.view[:] = np.sign(result.view[:])
result.view[:][np.abs(rf_update.view[:]) < 0.05] = 0. # 0.05 chosen by testing different numbers
print(result.view[:].min(), result.view[:].max())
plot_global(result, cube, vmin=-1, vmax=1)
# The first thing this tells us the random forest is mostly precipitating in regions where the physics is moistening the column. Look more closely at the blob in region x: (75, 100) and y: (20:40). If we look back at the plot of the physics update, we can see the sharp yellow areas where the physics and random forest agree on sign are grid-scale precipitation. In this region, the random forest appears to indicate the physics parameterization is under-precipitating. We can confirm this by looking at a vertical column from this region, and seeing if the drying tendency from the random forest looks like precipitation.
# I can't remember the orientation and placement of the ranks, so let's plot them.
import copy
quantity = copy.deepcopy(result)
quantity.view[:] = cube.rank
plot_global(quantity, cube, vmin=0, vmax=5)
quantity.view[:, :] = np.arange(quantity.extent[0])[:, None]
plot_global(quantity, cube, vmin=0, vmax=quantity.extent[0])
quantity.view[:, :] = np.arange(quantity.extent[1])[None, :]
plot_global(quantity, cube, vmin=0, vmax=quantity.extent[1])
# It appears we want to inspect rank 1, whose first axis increases along x and second axis increases along y in the plot. We can use this information to ballpark the region we want to be looking at, and make a new plot to confirm we've selected the right index.
# +
ix = 85 - 48 # rank 1's area starts at x=48
iy = 25
# result is the -1/0/1 plot from earlier for sign agreement
quantity = copy.deepcopy(result)
if cube.rank == 1:
quantity.view[iy, ix] = 2
plot_global(quantity, cube, vmin=-1, vmax=2)
# -
# Great, our index is well within the region where the random forest is lightly precipitating and the physics is not. Let's take a look at the vertical profile.
if cube.rank == 1:
p = np.cumsum(state["pressure_thickness_of_atmospheric_layer"].view[:, iy, ix], axis=0)
physics_update = subtract(
state_after_physics["specific_humidity"],
state_after_dynamics["specific_humidity"]
)
rf_update = subtract(
state_after_rf["specific_humidity"],
state_after_physics["specific_humidity"]
)
plt.figure()
plt.plot(state_after_dynamics["specific_humidity"].view[:, iy, ix], p)
plt.gca().invert_yaxis()
plt.title("specific_humidity")
plt.figure()
plt.plot(physics_update.view[:, iy, ix], p)
plt.gca().invert_yaxis()
plt.title("physics update")
plt.figure()
plt.plot(rf_update.view[:, iy, ix], p)
plt.gca().invert_yaxis()
plt.title("random forest update")
# These plots tell us that we cannot reasonably call what the random forest is doing in this case "precipitation". The drying is happening broadly throughout the troposphere, and is not particularly concentrated where there is humidity or where the physics routines are precipitating (at and below 800 hPa).
# The strong negative humidity tendencies in the uppper troposphere are concerning, given the model does not have very much moisture at these levels. It's likely to cause negative humidity. Let's evolve the model a few hours, and see if the drying lessens or stops. Uncomment the line setting total_timesteps to do this (it is set to zero by default for testing this notebook). This may take a few minutes.
seconds_in_hour = 60*60
timesteps_in_hour = seconds_in_hour / timestep.total_seconds()
total_timesteps = 0
#total_timesteps = int(3 * timesteps_in_hour)
print(total_timesteps)
for i in range(total_timesteps):
fv3gfs.wrapper.step()
state = fv3gfs.wrapper.get_state(rf_model.inputs)
rf_model.update(state, timestep=timestep)
fv3gfs.wrapper.set_state_mass_conserving(state)
fv3gfs.wrapper.step_dynamics()
state_after_dynamics = fv3gfs.wrapper.get_state(names)
total_water_after_dynamics = column_total_water(
state_after_dynamics["specific_humidity"],
state_after_dynamics["pressure_thickness_of_atmospheric_layer"]
)
fv3gfs.wrapper.step_physics()
fv3gfs.wrapper.save_intermediate_restart_if_enabled()
state_after_physics = fv3gfs.wrapper.get_state(names)
total_water_after_physics = column_total_water(
state_after_physics["specific_humidity"],
state_after_physics["pressure_thickness_of_atmospheric_layer"]
)
state = fv3gfs.wrapper.get_state(rf_model.inputs)
rf_model.update(state, timestep=timestep)
fv3gfs.wrapper.set_state_mass_conserving(state)
state_after_rf = fv3gfs.wrapper.get_state(names)
total_water_after_rf = column_total_water(
state_after_rf["specific_humidity"],
state_after_rf["pressure_thickness_of_atmospheric_layer"]
)
if cube.rank == 1:
p = np.cumsum(state["pressure_thickness_of_atmospheric_layer"].view[:, iy, ix], axis=0)
physics_update = subtract(
state_after_physics["specific_humidity"],
state_after_dynamics["specific_humidity"]
)
rf_update = subtract(
state_after_rf["specific_humidity"],
state_after_physics["specific_humidity"]
)
plt.figure()
plt.plot(state_after_dynamics["specific_humidity"].view[:, iy, ix], p)
plt.gca().invert_yaxis()
plt.title("specific_humidity")
plt.figure()
plt.plot(physics_update.view[:, iy, ix], p)
plt.gca().invert_yaxis()
plt.title("physics update")
plt.figure()
plt.plot(rf_update.view[:, iy, ix], p)
plt.gca().invert_yaxis()
plt.title("random forest update")
# It's encouraging that we see the negative tendencies in the upper troposphere lessen. There's an interesting peak around 700 hPa which seems to be getting strengthened by the random forest. In an interactive notebook, we could continue our analysis by saving fields over the next few hours and producing an animation.
# This notebook could go on forever, but we'll stop here. Hopefully this has shown the strength of being able to use interactive execution of a Fortran model to diagnose model behavior. This is not limited to investigating machine learning parameterizations, or even Python parameterizations. A researcher could augment an existing Fortran model with new Fortran code also wrapped to be accessible from Python, and use interactive execution to diagnose issues with their scheme.
#
# While not strictly necessary, let's run the Fortran cleanup routines to deallocate memory and write any final restart or diagnostic files.
fv3gfs.wrapper.cleanup()
# ## Debugging tools
# If you get a deadlock or hang for another reason (even Python exceptions can cause this sometimes), you will need to look at the ipyparallel log file to see what went wrong.
# Use these cells to shut down the cluster if needed, for example to restart in the case of deadlocks.
# Make sure you are running on the notebook process (instead of the cluster processes) by running %autopx until it says it is disabled.
# %autopx
rc.shutdown(hub=True)
# + language="bash"
# ipcluster stop --profile=mpi
# -
# These cells list the files in the log directory, and print the first file (in effect a random file). This is useful if you get a crash quickly and only one log file is present. Otherwise, modify the command or write a `%%bash` cell to print the log files you would like to read.
# +
import os
log_dir = "/root/.ipython/profile_mpi/log"
print(os.listdir(log_dir))
# -
log_filename = os.path.join(log_dir, os.listdir(log_dir)[0])
print(open(log_filename, "r").read())
# If there are too many log files for you to find what you're looking for, try deleting the existing logs and re-executing commands to reproduce your issue.
# + language="bash"
# rm /root/.ipython/profile_mpi/log/*
| examples/jupyter/notebooks/random_forest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # CrossColumnAddTransformer
# This notebook shows the functionality in the CrossColumnAddTransformer class. This transformer changes the values of one column via an additive adjustment, based on the values in other columns. <br>
import pandas as pd
import numpy as np
import tubular
from tubular.mapping import CrossColumnAddTransformer
tubular.__version__
# ## Create dummy dataset
df = pd.DataFrame(
{
"factor1": [np.nan, "1.0", "2.0", "1.0", "3.0", "3.0", "2.0", "2.0", "1.0", "3.0"],
"factor2": ["z", "z", "x", "y", "x", "x", "z", "y", "x", "y"],
"target": [18.5, 21.2, 33.2, 53.3, 24.7, 19.2, 31.7, 42.0, 25.7, 33.9],
"target_int": [2, 1, 3, 4, 5, 6, 5, 8, 9, 8],
}
)
df.head()
df.dtypes
# ## Simple usage
# ### Initialising CrossColumnAddTransformer
# The user must pass in a dict of mappings, each item within must be a dict of mappings for a specific column. <br>
# The column to be adjusted is also specified by the user. <br>
# As shown below, if not all values of a column are required to define mappings, then these can be excluded from the dictionary. <br>
# All additive adjustments defined must be numeric (int or float)
# +
mappings = {
'factor1': {
'1.0': 1.1,
'2.0': 0.5,
'3.0': -4,
}
}
adjust_column = "target"
# -
map_1 = CrossColumnAddTransformer(adjust_column = adjust_column, mappings = mappings, copy = True, verbose = True)
# ### CrossColumnAddTransformer fit
# There is not fit method for the CrossColumnAddTransformer as the user sets the mappings dictionary when initialising the object.
# ### CrossColumnAddTransformer transform
# Only one column mapping was specified when creating map_1 so only this column will be all be used to adjust the value of the adjust_column when the transform method is run.
df[['factor1','target']]
df[df['factor1'].isin(['1.0', '2.0','3.0'])]['target'].groupby(df['factor1']).mean()
df_2 = map_1.transform(df)
df_2[['factor1','target']].head(10)
df_2[df_2['factor1'].isin(['1.0', '2.0','3.0'])]['target'].groupby(df_2['factor1']).mean()
# ## Column dtype conversion
# If all the column to be adjusted has dtype int, but the additive adjustments specified are non-integer, then the column will be converted to a float dtype.
# +
mappings_2 = {
'factor1': {
'1.0': 1.1,
'2.0': 0.5,
'3.0': 4,
}
}
adjust_column_2 = "target_int"
# -
map_2 = CrossColumnAddTransformer(adjust_column = adjust_column_2, mappings=mappings_2, copy = True, verbose = True)
df['target_int'].dtype
df['target_int'].value_counts(dropna = False)
df_3 = map_2.transform(df)
df_3['target_int'].dtype
df_3['target_int'].value_counts(dropna = False)
# # Specifying multiple columns
# If more than one column is used to define the mappings, then as addition is a commutative operation it does not matter which order the multipliers are applied in.
# +
mappings_4 = {
'factor1': {
'1.0': 1.1,
'2.0': 0.5,
'3.0': -4,
},
'factor2': {
'x': -3,
}
}
adjust_column_4 = "target"
# -
map_4 = CrossColumnAddTransformer(adjust_column = adjust_column_4, mappings = mappings_4, copy = True, verbose = True)
df[['factor1','factor2','target']].head()
# In the above example, target would only be adjusted for row 1 by adding 1.1 (as factor1=1.0), whereas row 2 would be adjusted by subtracting 2.5 (factor1=2.0 means adding 0.5, factor2='x' means subtracting 3, 0.5 - 3 = -2.5)
df_4 = map_4.transform(df)
df_4[['factor1','factor2','target']].head()
| examples/mapping/CrossColumnAddTransformer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.8 64-bit
# name: python3
# ---
# +
import numpy as np
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import tensorflow_datasets as tfds
from tensorflow.keras import layers
import matplotlib.cm as cm
import random
import glob
import os
from skimage.segmentation import chan_vese
import numpy as np
import PIL
from PIL import Image
from tensorboard.plugins.hparams import api as hp
import pandas as pd
from shutil import copy
import cv2
# Display
# from IPython.display import Image, display
import matplotlib.pyplot as plt
import matplotlib.cm as cm
# My functions
import my_functions as mf
# tf explain
from tf_explain.callbacks.grad_cam import GradCAMCallback
# -
import my_functions as mf
random_seed = 42
batch_size = 32
img_height = 180
img_width = 180
image_size = (img_height, img_width)
src_folder_path = "datasets\\ct_scan_3\\train\\COVID-positive\\"
des_folder_path = "datasets\\ct_scan_7\\train_contrast\\COVID-positive\\"
mf.aply_black_tophat_to_folder(src_folder_path, des_folder_path, disk_size=12)
train_ds = tf.keras.utils.image_dataset_from_directory(
'datasets/ct_scan_7/train_merge',
labels='inferred',
label_mode='int',
class_names=None,
color_mode='rgb',
batch_size=batch_size,
image_size=image_size,
shuffle=True,
seed=random_seed,
interpolation='bilinear',
follow_links=False,
crop_to_aspect_ratio=False)
val_ds = tf.keras.utils.image_dataset_from_directory(
'datasets/ct_scan_3/val',
labels='inferred',
label_mode='int',
class_names=None,
color_mode='rgb',
batch_size=batch_size,
image_size=image_size,
shuffle=True,
seed=random_seed,
interpolation='bilinear',
follow_links=False,
crop_to_aspect_ratio=False)
# +
inputs = keras.Input(shape=(img_height, img_width, 3))
data_augmentation = keras.Sequential(
[
layers.RandomFlip("horizontal"),
layers.RandomRotation(0.1)
]
)
x = data_augmentation(inputs) # Apply random data augmentation
# Pre-trained Xception weights requires that input be scaled
# from (0, 255) to a range of (-1., +1.), the rescaling layer
# outputs: `(inputs * scale) + offset`
scale_layer = keras.layers.Rescaling(scale=1/127.5, offset=-1)
x = scale_layer(x)
# The base model contains batchnorm layers. We want to keep them in inference mode
# when we unfreeze the base model for fine-tuning, so we make sure that the
# base_model is running in inference mode here.
base_model = keras.applications.InceptionResNetV2(
include_top=False,
weights="imagenet",
input_shape=(img_height, img_width, 3),
input_tensor=x
)
last_layer_base_model = mf.find_last_layer(base_model)
x = tf.keras.layers.GlobalAveragePooling2D()(last_layer_base_model.output)
x = keras.layers.Dense(units=128, activation='relu')(x)
x = keras.layers.Dropout(0.5)(x)
x = keras.layers.Dense(units=64, activation='relu')(x)
x = keras.layers.Dropout(0.5)(x)
x = keras.layers.Dense(units=32, activation='relu')(x)
x = keras.layers.Dropout(0.5)(x)
outputs = keras.layers.Dense(1, activation='sigmoid')(x)
model = keras.Model(inputs, outputs)
base_model.trainable = False # Freeze base model
model.summary()
# -
gcam_image_path = "datasets\\ct_scan_3\\val\\COVID-positive\\P021_106.png"
# +
AUTOTUNE = tf.data.AUTOTUNE
train_ds = train_ds.cache().prefetch(buffer_size=AUTOTUNE)
val_ds = val_ds.cache().prefetch(buffer_size=AUTOTUNE)
optimizer = keras.optimizers.Adam(learning_rate=0.001, beta_1=0.9, beta_2=0.999)
model.compile(
optimizer=optimizer,
loss=keras.losses.BinaryCrossentropy(),
metrics=["accuracy"])
early_stopping_cb = keras.callbacks.EarlyStopping(
patience=10,
restore_best_weights=True)
run_logdir = mf.get_run_logdir()
tensorboard_cb = keras.callbacks.TensorBoard(run_logdir)
data = (mf.get_img_array(gcam_image_path, image_size), None)
grd_cam_callback = GradCAMCallback(
validation_data=data,
class_index=0,
output_dir=run_logdir,
)
epochs = 100
base_model.trainable = False
model.fit(
train_ds,
epochs=epochs,
batch_size=32,
validation_data=val_ds,
callbacks=[
tensorboard_cb,
early_stopping_cb,
grd_cam_callback])
# -
| experiments/ct_experiment_16.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import geopandas as gpd
import geoviews as gv
gv.extension('matplotlib')
# %output fig='svg' size=200
# -
# ## Declaring data
geometries = gpd.read_file('../../assets/boundaries/boundaries.shp')
referendum = pd.read_csv('../../assets/referendum.csv')
gdf = gpd.GeoDataFrame(pd.merge(geometries, referendum))
# ## Plot
# %%opts Polygons [color_index='leaveVoteshare' colorbar=True]
gv.Polygons(gdf, vdims=['name', 'leaveVoteshare'], label='Brexit Referendum Vote')
| examples/gallery/matplotlib/brexit_choropleth.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: air_quality_index
# language: python
# name: air_quality_index
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
data=pd.read_csv("Data/main_data/air_quality_index.csv")
data.head()
# + tags=[]
data.info()
# -
# # Linear Regression
X=data.drop(columns='PM2.5')
y=data['PM2.5']
sns.heatmap(X.corr(),annot=True,cmap='RdYlGn')
plt.plot()
# ### high Multicolinearity
# - drop TM,Tm,VM column
# +
# X.drop(columns=['TM','Tm','VM'],inplace=True)
# -
# ## Feature Selection
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
X_train,X_test,y_train,y_test=train_test_split(X,y,train_size=0.7,random_state=0)
lr=LinearRegression()
lr.fit(X_train,y_train)
lr.score(X_train,y_train)
lr.score(X_test,y_test)
pd.DataFrame(lr.coef_,X.columns,columns=['Coeficient'])
y_pred=lr.predict(X_test)
sns.distplot(y_pred-y_test)
plt.plot()
plt.scatter(y_test,y_pred)
plt.plot()
# ### Metrics
from sklearn.metrics import mean_squared_error,mean_absolute_error
# MAE
mean_absolute_error(y_test,y_pred)
# MSE
mean_squared_error(y_test,y_pred)
# RMSE
np.sqrt(mean_squared_error(y_test,y_pred))
# ## Save Model
import pickle
with open('Models/Linear_Regression.pkl','wb') as f:
pickle.dump(lr,f)
| Air_Quality_Index/Linear_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.2 32-bit
# metadata:
# interpreter:
# hash: dcfa17eba1dc66845e0904f61caf004065b70fa6c516601030b63e53c64813c1
# name: python3
# ---
# # Solución de ecuaciones
# <p><code>Python en Jupyter Notebook</code></p>
# <p>Creado por <code><NAME></code> para el curso de <code>Métodos Numéricos</code></p>
# <style type="text/css">
# .border {
# display: inline-block;
# border: solid 1px rgba(204, 204, 204, 0.4);
# border-bottom-color: rgba(187, 187, 187, 0.4);
# border-radius: 3px;
# box-shadow: inset 0 -1px 0 rgba(187, 187, 187, 0.4);
# background-color: inherit !important;
# vertical-align: middle;
# color: inherit !important;
# font-size: 11px;
# padding: 3px 5px;
# margin: 0 2px;
# }
# </style>
#
# ## Raíces de ecuaciones
# Las raíces o ceros de una función continua, son los valores de $\color{#a78a4d}{x}$ tal que $\color{#a78a4d}{f(x) = 0}$; lo que es equivalente a resolver ecuaciones del tipo $\color{#a78a4d}{g(x)=h(x)}$, ya que esta ecuación puede reescribirse como $\color{#a78a4d}{f(x)=h(x)-g(x)=0}$. Los métodos numéricos de búsqueda de raíces son algoritmos que iteran sobre una solución aproximada, produciendo una secuencia de números que, con suerte, convergen hacia la raíz como límite.
#
# ## Agenda
# 1. Tipos de ecuaciones
# 1. Linealidad
# 1. Puntos periódicos
# 1. Solución de ecuaciones no lineales
# Importar módulos al cuaderno de jupyter
import math as m
import numpy as np
import pylab as plt
# ## 1. Tipos de Ecuaciones
# ---
# En matemáticas una ecuación es una igualdad entre dos expresiones, separadas por el signo igual; tal que en las expresiones miembro aparecen variables independientes relacionadas mediante operaciones matemáticas.
#
# \begin{align}
# \tag{1} f(x_1, x_2, ... , x_n) &= g(x_1, x_2, ... , x_n) \\
# \tag{2} f(x_1, x_2, ... , x_n) &= 0 \\
# \end{align}
#
# ### a. Ecuaciones de una variable
# Una simplificación de $(2)$ sucede cuando solo existe una variable independiente y la ecuación se puede reescribir como:
#
# \begin{align}
# \tag{3} f(x) & = 0 \\
# \end{align}
#
# ### b. Ecuaciones algebraicas o polinómicas
# Una simplificación adicional de de $(2)$ sucede cuando sus miembros incluyen unicamente sumas y multiplicaciones de la variable independiente; en cuyo caso la expresión se conoce como polinomio y se puede reescribir como la suma de $\color{#a78a4d}{n+1}$ monomios cuyos coeficientes o parámetros $\color{#a78a4d}{a_0, a_1, a_2, ... , a_n}$ son constantes en el dominio de los reales:
#
# \begin{align}
# \tag{4} f(x) &= 0 \\
# \tag{5} f(x) &= \sum_{1}^n a_i x^i \\
# \tag{6} a_0 + a_1 x + a_2 x^2 + ... + a_n x^n & = 0, \quad a_n \neq 0 \\
#
# \end{align}
#
# > **NOTA:** El número n se denomina grado de la ecuación y es igual al numero raíces o ceros que son solución de la igualdad; estas soluciones pertenecen al dominio de los complejos y pueden ser reales o imaginarias; en el caso de soluciones imaginarias se dan en parejas de complejos conjugados.
#
# ### c. Ecuaciones trascendentes
# Si la ecuación no puede expresarse o reducirse a un polinomio como las ecuaciones algebraicas se denominan ecuaciones trascendentes y su resolución va más allá del álgebra (trascienden el álgebra); las ecuaciones trascendentes más simples son las trigonométricas, logarítmicas y exponenciales sencillas.
#
# > **NOTA:** La solución para x de estas ecuaciones puede no ser trivial y suelen ser objeto de estudio de los métodos numéricos.
#
# +
x = np.linspace(-20, 20, 1024, endpoint=True)
y1 = (-1/1e7)*(x**7 - 350*x**5 + 30_625*x**3 - 562_500*x)
y2 = np.sin(x)/x
# Gráficas
#-------------------
fig, ax = plt.subplots(1, 2)
fig.set_size_inches(16, 4)
fig.suptitle('Raíces de funciones algebraicas y trascendentes'.upper(), fontsize=14, fontweight="bold")
# Gráficas 1
ax[0].axhline(y = 0, linewidth=1, color="#ccc")
ax[0].plot(x, y1, label="$y_1(x)$")
ax[0].set_xlim(-20, 20)
ax[0].set_ylim(-1.5, 1.5)
ax[0].set_title("Función polinómica", fontsize=8)
ax[0].set_xlabel("Variable independiente")
ax[0].set_ylabel("variable dependiente")
ax[0].legend()
# Gráficas 2
ax[1].axhline(y = 0, linewidth=1, color="#ccc")
ax[1].plot(x, y2, label="$y_2(x)$")
ax[1].set_xlim(-20, 20)
ax[1].set_ylim(-1.5, 1.5)
ax[1].set_title("Función trascendentes", fontsize=8)
ax[1].set_xlabel("Variable independiente")
ax[1].set_ylabel("variable dependiente")
ax[1].legend()
plt.show()
# -
# ## 2. Linealidad
# ---
# ### a. Ecuaciones lineales
# Un caso particular de las ecuaciones algebraicas sucede cuando solo los dos primeros coeficientes son distintos de cero y la solución para $\color{#a78a4d}{x}$ es única y trivial.
#
# \begin{align}
# \tag{7} a_0 + a_1 x & = 0, \quad a_1 \neq 0 \\
# \tag{8} x & = \frac{-a_0}{a_1} \\
# \end{align}
#
# ### b. Ecuaciones cuadráticas
# Un caso particular de las ecuaciones algebraicas sucede cuando solo los tres primeros coeficientes son distintos de cero y las 2 soluciones para $\color{#a78a4d}{x}$ es un par de complejos conjugados.
#
# \begin{align}
# \tag{9} a_0 + a_1 x + a_2 x^2 & = 0, \quad a_1 \neq 0 \\
# \tag{10} x_1, x_2 & = \frac{-a_1 \pm \sqrt{a_1^2 - 4 a_2 a_0 }}{2 a_2} \\
# \end{align}
#
# ### b. Ecuaciones de orden superior
# Sucede que el número de soluciones es proporcional al orden de la ecuación; cuando más de tres coeficientes son distintos de cero en las ecuaciones algebraicas la solución puede ser más compleja; aunque la solución algebraica es posible debido a que existen métodos matemáticos para acotar la solución.
# +
y3 = (x - 10)
y4 = (x - 10) * (x + 10)
y5 = (x + 10) * (x - 10) * x
# Gráficas
#-------------------
fig, ax = plt.subplots(1, 3)
fig.set_size_inches(16, 4)
fig.suptitle('Raíces de funciones algebraicas'.upper(), fontsize=14, fontweight="bold")
# Gráficas 1
ax[0].axhline(y = 0, linewidth=1.0, color="#ccc")
ax[0].plot(x, y3, label="$y_1(x)$")
ax[0].set_title("Función líneal", fontsize=8)
ax[0].set_xlabel("Variable independiente")
ax[0].set_ylabel("variable dependiente")
ax[0].legend()
# Gráficas 2
ax[1].axhline(y = 0, linewidth=1, color="#ccc")
ax[1].plot(x, y4, label="$y_2(x)$")
ax[1].set_title("Función cuadrática", fontsize=8)
ax[1].set_xlabel("Variable independiente")
ax[1].legend()
# Gráficas 3
ax[2].axhline(y = 0, linewidth=1, color="#ccc")
ax[2].plot(x, y5, label="$y_3(x)$")
ax[2].set_title("Función cubica", fontsize=8)
ax[2].set_xlabel("Variable independiente")
ax[2].legend()
plt.show()
# -
# ## 3. Puntos periódicos
# ---
# En el estudio de sistemas dinámicos, un punto periódico de una función iterada es el punto al cual el sistema retorna luego de un cierto número de iteraciones, al cabo de un cierto tiempo.
#
# En matemáticas, los puntos periódicos vuelven al mismo valor después de un número finito de iteraciones de la función y un caso particular de ellos son los puntos fijos, donde el periodo $\color{#a78a4d}{T=1}$.
#
# \begin{align}
# \tag{11} (f:x \to x) \wedge (p, \text{ es periódico}) \Rightarrow \exists \ n : f_n(p)=p\\
#
# \end{align}
#
# ### a. Punto fijo
# En matemáticas, un punto fijo de una función real es un punto cuya imagen producida por la función es él mismo; esto que significa que si $\color{#a78a4d}{p}$ es un número real, $\color{#a78a4d}{p}$ es un punto fijo de la función $\color{#a78a4d}{f(x)}$ si y sólo si $\color{#a78a4d}{p = f(p)}$
#
# <p align="center">
# <img height="200" src="img/Fixed_Point_Graph.png">
# </p>
#
# > **NOTA:** En la gráfica (p, f(p)) pertenece a la recta $y=x$
# ### <code>Ejemplo:</code> Puntos fijos de una función
# ---
# Sea la función $f(x) = x^2$ los puntos fijos son números reales p tal que $p = p^2$; entonces 0 y 1 son los puntos fijos de $f(x)$, porque $f(0) = 0$ y $f(1) = 1$.
x = np.linspace(-2, 2, 256, endpoint=True)
plt.figure(figsize=(5, 5))
plt.plot(x, x**2, linewidth=2, color="#00F" )
plt.plot(x, x, linestyle="--", color="gray")
plt.plot(0, 0, marker="o", markersize=7, markeredgecolor="black", color="#fc0")
plt.plot(1, 1, marker="o", markersize=7, markeredgecolor="black", color="#fc0")
plt.xlim(x.min(), x.max())
plt.ylim(-1, 3)
plt.text(-0.7, 2, f"Puntos fijos\n $f(x)=x^2$", fontsize=17)
plt.grid()
plt.show()
# ## 4. Solución de ecuaciones no lineales
# ---
# Solucionar una ecuación es encontrar las raíces o ceros de una ecuación y aunque en algunos casos los métodos analíticos o exactos son posibles y sencillos; en otros casos es necesario recurrir a métodos numéricos.
#
# ### a. Métodos Numéricos
# Los métodos de la bisección y del punto fijo son los primeros métodos numéricos que se utilizaron para resolver ecuaciones algebraicas y trascendentes; estos representan el punto de partida para otros métodos más eficientes; del primer método o de la bisección derivan los métodos cerrados que se aproximan a la solución reduciendo intervalos y del segundo método o del punto fijo se derivan los métodos abiertos que se aproximan a la solución mediante una función auxiliar.
#
# Los métodos cerrados requieren dos valores iniciales que estén a ambos lados de la raíz, para acotarla. Este “acotamiento” se mantiene en tanto se aproxima a la solución; así, dichas técnicas son siempre convergentes; los métodos abiertos solo requieren de un valor, para aproximarse a la solución a través de la función auxiliar y son generalmente más rápidos; sin embargo, una mayor rapidez de convergencia se da cambio de una estabilidad menor.
#
# <p align="center">
# <img width="400" src="img/equations.png">
# </p>
#
# ### b. Selección de un método numérico
# Para seleccionar un método numérico que nos permita encontrar las raíces de una ecuación por aproximaciones sucesivas se requiere conocer información preliminar como:
#
# * Tipo de ecuación.
# * Número y tipo de las posibles raíces de la ecuación.
# * Información del dominio, rango, continuidad, derivadas, intervalos de crecimiento.
# * Ventajas y limitaciones que aporta cada algoritmo a la solución del problema.
# * Semilla o valoración inicial de la solución.
#
# ### c. Consideraciones generales
# Del [teorema del valor intermedio](https://es.wikipedia.org/wiki/Teorema_del_valor_intermedio) se tiene que si $\color{#a78a4d}{f(x)}$ es continua en un intervalo $\color{#a78a4d}{(a, b)}$ y cambia de signo en ese intervalo existe al menos una raíz real de la ecuación asociada en ese intervalo; sin embargo, existen consideraciones particulares para cada uno de los tipos de ecuaciones estudiadas.
#
# ### d. Ecuaciones algebraicas
# * El número de raíces es igual al grado del polinomio asociado; por lo tanto, en una ecuación de grado $\color{#a78a4d}{n}$, hay $\color{#a78a4d}{n}$ raíces reales o complejas.
# * Las raíces pueden ser reales o complejas conjugadas; por lo tanto, si el grado del polinomio es impar hay al menos una raíz real.
# * La posible existencia de raíces múltiples complica el problema; pero si $\color{#a78a4d}{f(x)}$ es derivable en $\color{#a78a4d}{(a, b)}$ y $\color{#a78a4d}{f'(x)}$ no cambia de signo en el ese intervalo, la raíz en ese intervalo es única.
# * Existen métodos como la [regla de Descartes](https://es.wikipedia.org/wiki/Regla_de_los_signos_de_Descartes), el [teorema de Budan]() y [teorema de Sturm](https://es.wikipedia.org/wiki/Teorema_de_Sturm) que nos permiten acotar convenientemente las raíces del polinomio.
#
# ### e. Ecuaciones trascendentes
# * Estas ecuaciones pueden tener cualquier número de posibles soluciones.
# * Pueden tener ninguna, una o más raíces en un intervalo.
# * Garantizar la existencia y unicidad de soluciones en estas ecuaciones es difícil de determinar y existe una mayor variedad de casos y comportamientos.
# * Pueden tener múltiples raíces, donde tanto la función $\color{#a78a4d}{f(x)}$ como su derivada $\color{#a78a4d}{f'(x)}$ son iguales a cero.
# * Algunas ecuaciones no pueden resolverse por métodos numéricos aún con un número muy grande de iteraciones.
#
#
# +
x = np.linspace(-3, 6, 256, endpoint=True)
y6 = np.exp(3*x - 12) + x * np.cos(3*x) - x**2 + 7.15
# Gráficas
#-------------------
fig, ax = plt.subplots(1, 1)
fig.set_size_inches(16, 4)
fig.suptitle('Búsqueda de raíces en funciones no lineales'.upper(), fontsize=14, fontweight="bold")
# Gráficas 1
ax.axhline(y = 0, linewidth=1, color="#ccc")
ax.plot(x, y6, label="$y_1(x)$")
ax.set_xlim(-3, 6)
ax.set_ylim(-11, 8)
ax.set_title("Función trascendente", fontsize=8)
ax.set_xlabel("Variable independiente")
ax.set_ylabel("variable dependiente")
ax.legend()
plt.show()
# -
# ## Analizando la figura anterior se observa:
# * La función $\color{#a78a4d}{f}$ asociada a la ecuación $\color{#a78a4d}{f(x)=0}$ es continua
# * En el intervalo $\color{#a78a4d}{(-3, -2)}$ visualmente no es posible determinar si la curva corta el eje $\color{#a78a4d}{x}$ una vez, dos veces o ninguna; con la posibilidad que un método iterativo pueda no detectar un intervalo válido, porque el signo $\color{#a78a4d}{f(-3)}$ es igual al signo de $\color{#a78a4d}{f(-2)}$.
# * En cada uno de los intervalos $\color{#a78a4d}{(2, 3)}$ y $\color{#a78a4d}{(5, 6)}$ existe una raíz única y real.
#
# + [markdown] colab_type="text" id="-Rh3-Vt9Nev9"
# ---
# ## Mas Recursos
#
# - [Ecuación](https://es.wikipedia.org/wiki/Ecuaci%C3%B3n) (Wikipedia)
# - [Ecuación algebraica](https://es.wikipedia.org/wiki/Ecuaci%C3%B3n_algebraica) (Wikipedia)
# - [Polinomio](https://es.wikipedia.org/wiki/Polinomio) (Wikipedia)
# - [Solución de Ecuaciones](https://es.wikipedia.org/wiki/Resoluci%C3%B3n_num%C3%A9rica_de_ecuaciones_no_lineales) (Wikipedia)
#
| Jupyter/21_Ecuaciones.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Imported Libraries
import pennylane as qml
from pennylane.optimize import AdamOptimizer
from pennylane import numpy as np
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from graphs import plot_correlation_matrix
import random
import torch
from torchvision import datasets, transforms
# Import libraries to be able to run on IBM quantum computers
from qiskit import Aer
from qiskit import *
# +
import sys
sys.path.append("..") # Adds higher directory to python modules path
sys.path.append
from qencode.initialize import setAB_amplitude, setAux, setEnt
from qencode.encoders import e2_classic
from qencode.training_circuits import swap_t
from qencode.qubits_arrangement import QubitsArrangement
from qencode.utils.mnist import get_dataset
# -
# !pip install kaggle
# +
from kaggle.api.kaggle_api_extended import KaggleApi
#import KaggleApi
api = KaggleApi()
api.authenticate()
https://www.kaggle.com/mlg-ulb/creditcardfraud/download
#downloading datasets for COVID-19 data
#api.dataset_download_files('imdevskp/corona-virus-report')
api.dataset_download_files('mlg-ulb/creditcardfraud')
# -
# # Credit Card Fraud - A Growing Issue
#
# Credit card fraud is a growing issue with \\$28.5 billion lost globally due to credit card fraud in 2020 and is expected to exceed \\$49 billion by 2030 [1]). In 2020, around 1 out of 4 digital interactions were credit card fraud attempts (Cite Arkose Labs). Since there are so many non fraudulent transactions, it is challenging to detect the fraudulent transactions. In this notebook, we will be using a quantum auto-encoder to perform anomaly detection.
#
# We can use the quantum auto encoder to encode the 4 qubit state into a 3 qubit state and then use a decoder to decode the 3 qubit state back into 4 qubit state. The quantum auto encoder is trained on the normal dataset (or in this case the non fraudulent transactions) which means the quantum auto
# To tell if a datapoint is an anomaly,
# Import the dataset
df = pd.read_csv('creditcard.csv')
# We are only going to print the first 5 rows because the dataset contains over 280,000 rows. Each row represents a transaction. Time shows the time passed between the current and first transactions and amount shows the dollar amount spent on the transaction. There are also 28 more features represented by V1, V2, ... , V28 which come from principal component analysis. Finally, there is the class, where a '0' represents no fraud committed and a '1' represents a fraudulent transaction
# Let's now check the class distribution
print('No Frauds: ', df['Class'].value_counts()[0])
print('Frauds: ', df['Class'].value_counts()[1])
# Credit card fraud is relatively rare, this creates a very imbalanced distribution. A very imbalanced distribution is not ideal as this can lead to overfitting and our model assuming no fraud most of the time. It is also challenging to find the true correlations between the features and class.
plot_correlation_matrix(df, "Original Correlation Matrix")
# As you can see, nothing can really be inferred from this correlation matrix since the data is so imbalanced. We are going to create a sub sample dataset with equal amounts of non fraudulent data and fraudulent data. We are also going to scale the 'Time' and 'Amount' values for better processing.
# +
#from sklearn.preprocessing import StandardScaler, RobustScaler
from sklearn.preprocessing import RobustScaler
# Scaling amount and time for the subsample
df['scaled_amount'] = RobustScaler().fit_transform(df['Amount'].values.reshape(-1,1))
df['scaled_time'] = RobustScaler().fit_transform(df['Time'].values.reshape(-1,1))
df.drop(['Time','Amount'], axis=1, inplace=True) # Drop the original time and amount values
# Add scaled amount and times to the data frame
scaled_amount = df['scaled_amount']
scaled_time = df['scaled_time']
df.drop(['scaled_amount', 'scaled_time'], axis=1, inplace=True)
df.insert(0, 'scaled_amount', scaled_amount)
df.insert(1, 'scaled_time', scaled_time)
# Create the balanced subsample of 49
df = df.sample(frac=1)
fraud_df = df.loc[df['Class'] == 1]
non_fraud_df = df.loc[df['Class'] == 0][:492]
normal_distributed_df = pd.concat([fraud_df, non_fraud_df])
sub_sample_df = normal_distributed_df.sample(frac=1, random_state=42)
# Display the first 5 rows
sub_sample_df.head()
# -
# We can now plot the correlation matrix of our new sub sample to get a better idea of the true correlations between features and 'Class'
sub_sample_corr = sub_sample_df.corr()
plot_correlation_matrix(sub_sample_corr, "Sub Sample Correlation Matrix")
# The correlations are now much more noticeable. Now, we can find the features with the strongest correlation to class. Half are the strongest positive correlations, half are the strongest negative correlations.
# +
def find_strongest_correlations(dataframe, latent_qubits):
num_features = latent_qubits**2
class_correlations = dataframe.loc['Class', :]
class_correlations = class_correlations.drop(index = 'Class')
feature_list = list(class_correlations.index)
correlation_list = [class_correlations[x] for x in feature_list]
features = []
correlations = []
for i in range(int(num_features/2)):
correlations.append(max(correlation_list))
features.append(feature_list[correlation_list.index(max(correlation_list))])
del feature_list[correlation_list.index(max(correlation_list))]
del correlation_list[correlation_list.index(max(correlation_list))]
correlations.append(min(correlation_list))
features.append(feature_list[correlation_list.index(min(correlation_list))])
del feature_list[correlation_list.index(min(correlation_list))]
del correlation_list[correlation_list.index(min(correlation_list))]
return features, correlations
feature_list, correlations = find_strongest_correlations(sub_sample_corr, 4)
print(find_strongest_correlations(sub_sample_corr, 4))
# -
# We now have 16 features that are the most correlated with 'Class'. In this use case, we will be using 4 qubits to represent the data which means we need 2^4 = 16 features to encode into the 4 qubits though amplitude encoding. Later, we will use the quantum autoencoder to encode the 4 qubits into 3 qubits and then use a decoder to decode those 3 qubits back to 4 qubits.
# +
# Dataframe of all non fraudulent transactions
branch = df
non_fraud = branch[branch["Class"]!="1"]
# All examples of non fraudulent data with 16 features
non_fraud = non_fraud[feature_list]
non_fraud.head()
input_data = non_fraud.to_numpy()
# -
# # Training
#
# +
shots = 2500 #The amount of shots used for each epoch of training
nr_trash= 1 # Number of qubits 'thrown away'
nr_latent= 3 # Number of qubits left after the encoder is used
nr_ent = 0
epochs = 500 # Number of iterations of training to perform to find the final encoder parameters
learning_rate = .005 # Learning rate for the optimizer, dictates how fast the optimizer trains
batch_size = 2
num_samples = 50 # Number of training samples used for each epoch
beta1 = 0.9
beta2 = 0.999
opt = AdamOptimizer(learning_rate, beta1=beta1, beta2=beta2)
# Organizes and specifies our qubits for the device
spec = QubitsArrangement(nr_trash, nr_latent, nr_swap=1, nr_ent=nr_ent)
print("Qubits:", spec.qubits)
#set up the device
#dev = qml.device('qiskit.ibmq', wires=spec.num_qubits, backend = 'ibmq_qasm_simulator', ibmqx_token = "6<PASSWORD>67e3<PASSWORD>5e0<PASSWORD>")
dev = qml.device('qiskit.aer', wires=spec.num_qubits, backend = 'qasm_simulator')
# -
@qml.qnode(dev)
def training_circuit_example(init_params, encoder_params, reinit_state):
#initilaization
setAB_amplitude(spec, init_params)
setAux(spec, reinit_state)
setEnt(spec, inputs=[1 / np.sqrt(2), 0, 0, 1 / np.sqrt(2)])
#encoder
for params in encoder_params:
e2_classic(params, [*spec.latent_qubits, *spec.trash_qubits])
#swap test
swap_t(spec)
return [qml.probs(i) for i in spec.swap_qubits]
def fid_func(output):
# Implemented as the Fidelity Loss
# output[0] because we take the probability that the state after the
# SWAP test is ket(0), like the reference state
fidelity_loss = 1 / output[0]
return fidelity_loss
# Define cost function
def cost(encoder_params, X):
reinit_state = [0 for i in range(2 ** len(spec.aux_qubits))]
reinit_state[0] = 1.0
loss = 0.0
for x in X:
output = training_circuit_example(init_params=x[0], encoder_params=encoder_params, reinit_state=reinit_state)[0]
f = fid_func(output)
loss = loss + f
return loss / len(X)
# Define fidelity function
def fidelity(encoder_params, X):
reinit_state = [0 for i in range(2 ** len(spec.aux_qubits))]
reinit_state[0] = 1.0
loss = 0.0
for x in X:
output = training_circuit_example(init_params=x[0], encoder_params=encoder_params, reinit_state=reinit_state)[0]
f = output[0]
loss = loss + f
return loss / len(X)
def iterate_batches(X, batch_size):
random.shuffle(X)
batch_list = []
batch = []
for x in X:
if len(batch) < batch_size:
batch.append(x)
else:
batch_list.append(batch)
batch = []
if len(batch) != 0:
batch_list.append(batch)
return batch_list
# +
training_data = [ torch.tensor([input_data[i]]) for i in range(num_samples)]
test_data = [ torch.tensor([input_data[i]]) for i in range(num_samples,num_samples+num_samples)]
X_training = training_data
X_tes = test_data
# -
# initialize random encoder parameters
from pennylane import numpy as np
nr_encod_qubits = len(spec.trash_qubits) + len(spec.latent_qubits)
nr_par_encoder = 15 * int(nr_encod_qubits*(nr_encod_qubits-1)/2)
encoder_params = np.random.uniform(size=(1, nr_par_encoder), requires_grad=True)
encoder_params1 = encoder_params
# +
# Create a tensor dataset with only fraud data and most correlated features
# for finding the fidelity of the quantum autoencoder on fraud transactions
fraud = fraud_df[feature_list]
np_fraud = fraud.to_numpy()
fraud_data = [ torch.tensor([np_fraud[i]]) for i in range(len(fraud.to_numpy()))]
fraud.head()
# +
loss_hist=[]
fid_hist=[]
loss_hist_test=[]
fid_hist_test=[]
fraud_fid=[]
for epoch in range(epochs):
batches = iterate_batches(X=training_data, batch_size=batch_size)
for xbatch in batches:
encoder_params = opt.step(cost, encoder_params, X=xbatch)
if epoch%5 == 0:
loss_training = cost(encoder_params, X_training )
fidel = fidelity(encoder_params, X_training )
loss_hist.append(loss_training)
fid_hist.append(fidel)
print("Epoch:{} | Loss:{} | Fidelity:{}".format(epoch, loss_training, fidel))
loss_test = cost(encoder_params, X_tes )
fidel = fidelity(encoder_params, X_tes )
loss_hist_test.append(loss_test)
fid_hist_test.append(fidel)
print("Test-Epoch:{} | Loss:{} | Fidelity:{}".format(epoch, loss_test, fidel))
f_fidel = fidelity(encoder_params, fraud_data )
fraud_fid.append(f_fidel)
print("Fraud Fidelity:{}".format(f_fidel))
# +
fig = plt.figure()
plt.plot([x for x in range(0,len(loss_hist)*5,5)],np.array(fid_hist),label="train fidelity")
plt.plot([x for x in range(0,len(loss_hist)*5,5)],np.array(fid_hist_test),label="test fidelity")
plt.plot([x for x in range(0,len(loss_hist)*5,5)],np.array(fraud_fid),label="fraud fidelity")
plt.legend()
plt.title("Fraud 4-3-4 Compression Fidelity e2",)
plt.xlabel("Epoch")
plt.ylabel("Fidelity")
plt.savefig("Fraud 4-3-4 Compression Fidelity e2")
print("fidelity:",fid_hist[-1])
# +
fig = plt.figure()
plt.plot([x for x in range(0,len(loss_hist)*5,5)],np.array(loss_hist),label="train loss")
plt.plot([x for x in range(0,len(loss_hist)*5,5)],np.array(loss_hist_test),label="test loss")
plt.legend()
plt.title("Fraud 4-3-4 Compression Loss e2",)
plt.xlabel("Epoch")
plt.ylabel("Loss")
plt.savefig("Fraud 4-3-4 Compression Loss e2")
print("loss:",loss_hist[-1])
# +
name = "fraud_434_training_e2"
Circuit_prop={ "shots":shots, "nr_trash":nr_trash, "nr_latent":nr_latent, "nr_ent":nr_ent }
Training_param = { "num_samples" == num_samples,
"batch_size" == batch_size,
#"nr_layers"== nr_layers,
"epochs" == epochs,
"learning_rate" == learning_rate,
"beta1" == beta1,
"beta2 "== beta2,
"optimizer"=="Adam"}
performance={"loss_hist":loss_hist, "fid_hist":fid_hist,
"loss_hist_test":loss_hist_test, "fid_hist_test":fid_hist_test,
"encoder_params":encoder_params}
experiment_data={"Circuit_prop":Circuit_prop,
"Training_param":Training_param,
"performance:":performance,
"Name":name}
# open file for writing
f = open(name+".txt","w")
f.write( str(experiment_data) )
# -
experiment_parameters={"autoencoder":"e2","params":encoder_params}
f=open("Params_Fraud_encoder_e2-CorrelatedFeautures.txt","w")
f.write(str(experiment_parameters))
# +
branch = df
non_fraud_df = branch.loc[branch["Class"]!=1][:492]
non_fraud = non_fraud_df[feature_list]
np_non_fraud = non_fraud.to_numpy()
non_fraud_data = [ torch.tensor([np_non_fraud[i]]) for i in range(len(non_fraud.to_numpy()))]
non_fraud_flist=[]
for b in non_fraud_data:
f=fidelity(encoder_params, [b])
non_fraud_flist.append(f.item())
print(min(non_fraud_flist))
print(max(non_fraud_flist))
# +
#np_ilegal= ilegal.to_numpy()
#ilegal_data = [ torch.tensor([np_ilegal[i]]) for i in range(len(ilegal.to_numpy()))]
fraud_flist=[]
for b in fraud_data:
f=fidelity(encoder_params, [b])
fraud_flist.append(f.item())
print(min(fraud_flist))
print(max(fraud_flist))
# +
plt.hist(non_fraud_flist, bins =100 ,label="non_fraud",color = "red",alpha=0.4)
plt.hist(fraud_flist, bins = 100 ,label="fraud", color = "skyblue",alpha=0.4)
plt.title("Compression fidelity",)
plt.legend()
plt.savefig("Compression_fidelity")
plt.show()
# -
| Use-case_Fraud_detection/BEST_fraud_detection/.ipynb_checkpoints/QuantumCreditFraud-Qiskit-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Number of clicks prediction
# ---
import pandas as pd
import numpy as np
import requests
from io import StringIO
import matplotlib.pyplot as plt
import time
from pandas.plotting import scatter_matrix
from sklearn.preprocessing import MinMaxScaler, PowerTransformer
from sklearn.pipeline import Pipeline
# ## EDA
# +
# function for reading data from the url
# -
def loading_data(file_url):
"""
Reading data from a given url
inputs:
------
url : link to the data
returns
-----:
data : data frame
"""
file_id = file_url.split('/')[-2]
dwn_url='https://drive.google.com/uc?export=download&id=' + file_id
url = requests.get(dwn_url).text
csv_raw = StringIO(url)
df = pd.read_csv(csv_raw)
return df
# +
# Let's load the training data
# -
hotel_url = 'https://drive.google.com/file/d/1c85h1hzgzLvAeYSh-EVpY6Gz3dYLsd6R/view?usp=sharing'
hotel_data = loading_data(hotel_url )
# +
#hotel_data = hotel_data.sample(100)
# -
# first we will have a quick look at the data head and tail
hotel_data.head()
# +
# we see that we have only numerical predictors
# -
hotel_data.tail()
# +
# We can see right a way that our data has Nan in it which indicate we will need some prepocessing and cleaning
# +
# let's get a quick description of the dat set
hotel_data.info()
# -
# > We have 12 predictors, alll of them are float except the n_clicks which is integer
# > The data contains 396478 row. From the comparision of the counts, it appears that `avg_rating` has the maximum number of NaNs.
# Now let's have a look at the statistics of values
hotel_data.describe()
# > There are a couple of interesting observations:
# - `Content score` ranging from 7 to 71 with a mean of 50.99.
# - We have some negative `n_images` which have to be removed.
# - Median `avg_rating` is 83.0 but median `stars` is 0.
# - We have a median of 189 for `n_reviews`.
# - The `avg_price` ranges from 4 to 8000 and the `avg_saving` ranges from 0 to 99%.
# - Hotels have avg `n_clicks` of 13.78
# +
# Now let's look at a histogram for the data
hotel_data.drop(columns = ['hotel_id']).hist(bins = 50, figsize=(10,15))
plt.show()
# -
# - > There is big difference between variables in terms of scale.
# - > we have many features with heavy tails distribution.
# +
# To solve this issue we can use power transformation.
# -
# ### Power Transformation
# perform a box-cox transform of the dataset
scaler = MinMaxScaler(feature_range=(1, 2))
power = PowerTransformer(method='box-cox')
pipeline = Pipeline(steps=[('s', scaler),('p', power)])
data = pipeline.fit_transform(hotel_data.drop(columns = ['hotel_id']))
# convert the array back to a dataframe
dataset = pd.DataFrame(data)
# histograms of the variables
dataset.hist(bins = 50, figsize=(10,15));
# > Some of the features such as avg_price really benefited from this transformation
# Let's calculate Pearson's correlation to see if there is linear relationship between variables
corr_hotel = hotel_data.corr()
corr_hotel ['n_clicks'].sort_values(ascending = False)
# now let's see how the numerical variables are correlated, we will drop city_id since it's categorical variable and hotel_id
# because it's just an identifier
numerical_vars = ['content_score','n_images','distance_to_center','avg_rating','stars','n_reviews','avg_rank','avg_price','avg_saving_percent',
'n_clicks']
scatter_matrix(hotel_data[numerical_vars],figsize = (16,12));
# > Both correlations and scatter matrix indicate that there is no single variable that has a string linear correlation with out target.
# +
## Now le't look at features one by one
# +
#1. hote_id : this column will be dropped
# -
original_size = len(hotel_data)
original_size
# 2 content score
hotel_data[['content_score']].describe()
hotel_data[hotel_data['content_score'].isnull()]
# > We have 508 rows that have not only content_score as Nan but also `n_images`,`distance_to_center`, `avg_rating`, `stars`, and `n_reviews`
hotel_data = hotel_data[hotel_data['content_score'].notnull()]
len(hotel_data)
# 3 n_images
hotel_data[['n_images']].describe()
# > We have some negative `n_images` here we need to remove. Let's first check how many of these we have
#
hotel_data[hotel_data['n_images'] < 0]
# > 3361 rows, we notice that `avg_rating` for those rows is also NAN, so we will go agead and remove them.
#
hotel_data = hotel_data[hotel_data['n_images'] >= 0]
len(hotel_data)
# 4 content score
hotel_data[['distance_to_center']].describe()
# > No negative values here!
hotel_data[hotel_data['distance_to_center'].isnull()]
# > We see that the same rows have NaN's in many other columns, we will drop these
hotel_data = hotel_data[hotel_data['distance_to_center'].notnull()]
# 5 avg_rating
hotel_data[['avg_rating']].describe()
# > No negative values here!
hotel_data[hotel_data['avg_rating'].isnull()]
# - > Well this is big chunck of the data with NaN in `avg rating`. Instead of removing these rows, we have to replace them with a suitable value
# - > We could use mean, median or 0. For me, I think it makes more sense to replace these value with 0. A missing `avg_rating` is more likely to be 0
hotel_data['avg_rating'] = hotel_data['avg_rating'].fillna(0)
# 6 stars
hotel_data[['stars']].describe()
# > No negative values
hotel_data[hotel_data['stars'].isnull()]
# > Only 8 rows that can be safely removed!
hotel_data = hotel_data[hotel_data['stars'].notnull()]
# 7 avg_rating
hotel_data[['n_reviews']].describe()
# > No negative values here
hotel_data[hotel_data['n_reviews'].isnull()]
# > We don't have Nans here!
# 8 avg_rank
hotel_data[['avg_rank']].describe()
# > No ngative values
hotel_data[hotel_data['avg_rank'].isnull()]
# > We don't have Nans here
# 9 avg_price
hotel_data[['avg_price']].describe()
# > No negative values here.
hotel_data[hotel_data['avg_price'].isnull()]
# > We have 153 rows, we can see that in the same raws we have Nan in avg_saving_percent, we can go ahead and remove those
#
hotel_data = hotel_data[hotel_data['avg_price'].notnull()]
len(hotel_data)
# 20 avg_price
hotel_data[['avg_saving_percent']].describe()
# > No negative values here!
hotel_data[hotel_data['avg_saving_percent'].isnull()]
# > No NaNs any more!
# +
# 11 city_id : This is categorical variable. Usually we use one-hot encoding with this type variable. Let's see if it is feasible to apply this strategy here
# -
city_id_count = hotel_data['city_id'].value_counts().reset_index(name='count').rename(columns={'index': 'city_id'})
city_id_count
# > we have 33,157 unique values for city_id
city_id_count .hist();
# > We note here that many of these city_ids have low frequency.
#
# - Encoding this categorical variable will result in high dimentional data that was too big to my pc memory. And as we noted above most of these `city_id's` have low frequency, therefore I am going to replace each `cit_id` with it's count,i.e how much it appears in the dat set and will then treat it as a numerical variable
# ### Creating new numerical feature for city_id
# +
# we will merge the original data with the city_id_count from above
# -
hotel_data = pd.merge(hotel_data,city_id_count ,on=['city_id'], how='left' )
hotel_data
# now we can drop the original city_id columns
hotel_data = hotel_data.drop(columns = ['city_id'])
# +
# Finally let's look at our target variable
# -
hotel_data[['n_clicks']].describe()
# let's check the distribution of n_clicks
hotel_data[['n_clicks']].hist(bins=7);
hotel_data[['n_clicks']].boxplot();
# +
# it seems we have a lot of outliers, let's see how many of those is are out side the range between the 5 and 95 quantiles
y = hotel_data['n_clicks']
removed_outliers = y.between(y.quantile(.05), y.quantile(.95))
print(str(y[removed_outliers].size) + "/" + str(len(hotel_data)) + " data points remain.")
# -
index_names = hotel_data[~removed_outliers].index
hotel_data.drop(index_names, inplace=True)
len(hotel_data)
# > We will loose around 20,000 rows. let's go ahead and remove those outliers
# +
### Now let's look at the data type for each column
# -
hotel_data.dtypes
# +
# let's convert n_clicks to float as we expect out model to output float numbers for n_clicks
# -
hotel_data['count'] = hotel_data['count'].astype(float)
hotel_data['n_clicks'] = hotel_data['n_clicks'].astype(float)
hotel_data.to_csv('cleaned.csv', index = False)
| 1.eda_preprocessing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/enesdemirag/cifar10-classification/blob/main/cifar10_deep_cnn.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="n_R3RiRl0ANl"
# Dependencies
import random
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from datetime import datetime as dt
from sklearn.metrics import confusion_matrix
from tensorflow.keras.layers import Flatten, Dense, Conv2D, Dropout, MaxPooling2D, Activation, BatchNormalization
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import RMSprop, Adam
from tensorflow.keras.losses import MeanSquaredError, CategoricalCrossentropy
from tensorflow.keras.utils import plot_model, to_categorical
from tensorflow.keras.metrics import Precision, Recall, CategoricalAccuracy, AUC
from tensorflow.keras.datasets import cifar10
from tensorflow.keras.callbacks import TensorBoard, LearningRateScheduler
from keras import regularizers
# %matplotlib inline
# %load_ext tensorboard
# + id="KS8Dc2M16pRE"
# Class labels
classes = {
0: "airplane",
1: "automobile",
2: "bird",
3: "cat",
4: "deer",
5: "dog",
6: "frog",
7: "horse",
8: "ship",
9: "truck",
}
# + id="HQFa-VaHZkna"
def lr_schedule(epoch):
lrate = 0.001
if epoch > 75:
lrate = 0.0005
elif epoch > 100:
lrate = 0.0003
return lrate
# + id="P6d5AX5NZp52"
class DeepCNN(object):
def __init__(self):
self.model = Sequential()
self.model.add(Conv2D(32, (3,3), padding='same', kernel_regularizer=regularizers.l2(1e-4), input_shape=(32, 32, 3)))
self.model.add(Activation('elu'))
self.model.add(BatchNormalization())
self.model.add(Conv2D(32, (3,3), padding='same', kernel_regularizer=regularizers.l2(1e-4)))
self.model.add(Activation('elu'))
self.model.add(BatchNormalization())
self.model.add(MaxPooling2D(pool_size=(2,2)))
self.model.add(Dropout(0.2))
self.model.add(Conv2D(64, (3,3), padding='same', kernel_regularizer=regularizers.l2(1e-4)))
self.model.add(Activation('elu'))
self.model.add(BatchNormalization())
self.model.add(Conv2D(64, (3,3), padding='same', kernel_regularizer=regularizers.l2(1e-4)))
self.model.add(Activation('elu'))
self.model.add(BatchNormalization())
self.model.add(MaxPooling2D(pool_size=(2,2)))
self.model.add(Dropout(0.3))
self.model.add(Conv2D(128, (3,3), padding='same', kernel_regularizer=regularizers.l2(1e-4)))
self.model.add(Activation('elu'))
self.model.add(BatchNormalization())
self.model.add(Conv2D(128, (3,3), padding='same', kernel_regularizer=regularizers.l2(1e-4)))
self.model.add(Activation('elu'))
self.model.add(BatchNormalization())
self.model.add(MaxPooling2D(pool_size=(2,2)))
self.model.add(Dropout(0.4))
self.model.add(Flatten())
self.model.add(Dense(10, activation='softmax'))
self.model.compile(
optimizer = RMSprop(lr=0.001,decay=1e-6),
loss = CategoricalCrossentropy(),
metrics = [Precision(name="precision"), Recall(name="recall"), CategoricalAccuracy(name="accuracy"), AUC(name="auc")]
)
def train(self, features, labels, validation_data, batch_size=32, epochs=100, shuffle=True):
log_dir = "logs/fit/" + dt.now().strftime("%Y%m%d-%H%M%S")
tensorboard_callback = TensorBoard(log_dir=log_dir, histogram_freq=1)
history = self.model.fit(features, labels, batch_size, epochs, shuffle=shuffle, verbose=1, validation_data=validation_data, callbacks=[tensorboard_callback, LearningRateScheduler(lr_schedule)])
self.epochs = history.epoch
self.hist = pd.DataFrame(history.history)
return self.epochs, self.hist
def test(self, features, labels):
self.loss, self.presicion, self.recall, self.accuracy, self.auc = self.model.evaluate(features, labels, verbose=0)
return self.loss, self.presicion, self.recall, self.accuracy, self.auc
def predict(self, img):
return self.model.predict(img)
def save(self, path="./saved_models/"):
timestamp = dt.timestamp(dt.now())
filename = path + "CNN_" + str(timestamp)
plot_model(self.model, to_file=filename + ".png", show_shapes=True, show_layer_names=True)
self.model.save(filename + ".h5")
def summary(self):
return self.model.summary()
# + id="46ijnu6K032I"
# Dataset
def prepare_data():
(images_train, labels_train), (images_test, labels_test) = cifar10.load_data()
# normalize the pixel values
images_train = images_train.astype('float32')
images_test = images_test.astype('float32')
images_train /= 255
images_test /= 255
# one hot encoding
labels_train = to_categorical(labels_train, 10)
labels_test = to_categorical(labels_test, 10)
return images_train, labels_train, images_test, labels_test
# + id="VNTkJ1O809_C" colab={"base_uri": "https://localhost:8080/"} outputId="68b9ec1c-54a0-477e-9dc0-8352893b8d9b"
# Preprocessing
images_train, labels_train, images_test, labels_test = prepare_data()
# + colab={"base_uri": "https://localhost:8080/"} id="7vJFI1L25W2D" outputId="3106f76d-df2c-450b-9064-0d95e810774b"
# Creating model
model = DeepCNN()
model.summary()
# + id="-CdKUP-57LWu" colab={"base_uri": "https://localhost:8080/"} outputId="ba820e9c-b408-440b-b739-f682b6d4d2f4"
# Training Deep CNN Model
epochs, hist = model.train(images_train, labels_train, validation_data=(images_test, labels_test), epochs=100)
# + colab={"base_uri": "https://localhost:8080/", "height": 367} id="U7L1YT6d758G" outputId="62bbf82f-eb10-410d-abac-7faae13f8c88"
# Plotting the training metrics
fig, ax = plt.subplots(1, 5, figsize=(30, 5))
ax[0].set_xlabel("Epoch")
ax[0].set_ylabel("Value")
ax[0].set_title("Loss")
ax[1].set_xlabel("Epoch")
ax[1].set_ylabel("Value")
ax[1].set_title("Presicion")
ax[2].set_xlabel("Epoch")
ax[2].set_ylabel("Value")
ax[2].set_title("Recall")
ax[3].set_xlabel("Epoch")
ax[3].set_ylabel("Value")
ax[3].set_title("AUC")
ax[4].set_xlabel("Epoch")
ax[4].set_ylabel("Value")
ax[4].set_title("Accuracy")
ax[0].plot(epochs[1:], hist["loss"][1:], color="r")
ax[1].plot(epochs[1:], hist["precision"][1:], color="g")
ax[2].plot(epochs[1:], hist["recall"][1:], color="b")
ax[3].plot(epochs[1:], hist["auc"][1:], color="y")
ax[4].plot(epochs[1:], hist["accuracy"][1:], color="k")
# + colab={"base_uri": "https://localhost:8080/"} id="7ShZ5ooh1ECF" outputId="01e899ee-f642-42f1-ea05-59495d839e99"
# Testing Deep CNN Model
loss, precision, recall, accuracy, auc = model.test(images_test, labels_test)
print("--- Test Results ---")
print("%-10s: %f" %("Loss", loss))
print("%-10s: %f" %("Precision", precision))
print("%-10s: %f" %("Recall", recall))
print("%-10s: %f" %("AUC", auc))
print("%-10s: %f" %("Accuracy", accuracy))
# + colab={"base_uri": "https://localhost:8080/", "height": 160} id="TKfYr_gs8l_7" outputId="955021ea-b8d0-4f50-9ed5-726858837112"
# Prediction Deep CNN Model
fig, ax = plt.subplots(1, 10, figsize=(20, 5))
for i in range(10):
rand_sample = random.randint(0, 1000)
img = images_test[rand_sample].reshape((1, 32, 32, 3))
y_orig = labels_test[rand_sample]
y_pred = model.predict(img)
y_orig = list(y_orig).index(1)
y_pred = [1 if i == max(y_pred[0]) else 0 for i in y_pred[0]].index(1)
ax[i].set_xticks([])
ax[i].set_yticks([])
ax[i].imshow(img[0])
ax[i].set_title("True: %s \nPredict: %s" % (classes[y_orig], classes[y_pred]))
# + id="4d0SdmGqzykF"
def print_confusion_matrix(confusion_matrix, class_names, figsize = (10,7), fontsize=14):
df_cm = pd.DataFrame(confusion_matrix, index=class_names, columns=class_names)
fig = plt.figure(figsize=figsize)
try:
heatmap = sns.heatmap(df_cm, annot=True, fmt="d")
except ValueError:
raise ValueError("Confusion matrix values must be integers.")
heatmap.yaxis.set_ticklabels(heatmap.yaxis.get_ticklabels(), rotation=0, ha='right', fontsize=fontsize)
heatmap.xaxis.set_ticklabels(heatmap.xaxis.get_ticklabels(), rotation=45, ha='right', fontsize=fontsize)
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 498} id="UTyB64J32M8c" outputId="a704130c-04ff-44b8-e671-60f1f05a8c3a"
y_pred = np.argmax(model.predict(images_test), axis=1)
y_truth = np.argmax(labels_test, axis=1)
print_confusion_matrix(confusion_matrix(y_truth, y_pred), list(classes.values()))
# + id="XqQX6HiucvzK"
model.model.save("model.h5")
# + id="LYl6aVP03uTJ"
# %tensorboard --logdir logs/fit
| cifar10_deep_cnn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Two-dimensional advection equation
#
# We want to solve the following PDE:
#
# \begin{equation}
# \frac{\partial \phi}{\partial t} + u \frac{\partial \phi}{\partial x} + v \frac{\partial \phi}{\partial y} = 0
# \end{equation}
#
# The independen variables (i.e, $x$, $y$ and $t$) are used as input values for the NN, and the solution (i.e. $\phi(x,y,t)$) is the output. In order to find the solution, at each step the NN outputs are derived w.r.t the inputs. Then, a loss function that matches the PDE is built and the weights are updated accordingly. If the loss function goes to zero, we can assume that our NN is indeed the solution to our PDE. We will try to find a general solution for different values of $u$, so it will be set also as an input.
# +
# imports
import numpy as np
import matplotlib.pyplot as plt
import nangs
import torch
device = "cuda" if torch.cuda.is_available() else "cpu"
nangs.__version__, torch.__version__
# +
from nangs import PDE
U, V = -0.5, 1
class Adv2d(PDE):
def computePDELoss(self, inputs, outputs):
# compute gradients
grads = self.computeGrads(outputs, inputs)
# compute loss
dpdx, dpdy, dpdt = grads[:, 0], grads[:, 1], grads[:, 2]
return {'pde': dpdt + U*dpdx + V*dpdy}
pde = Adv2d(inputs=('x', 'y', 't'), outputs='p')
# +
# define the mesh
from nangs import Mesh
x = np.linspace(0,1,20)
y = np.linspace(0,1,20)
t = np.linspace(0,1,20)
mesh = Mesh({'x': x, 'y': y, 't': t}, device=device)
pde.set_mesh(mesh)
# +
# initial condition (t = 0)
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
from matplotlib import cm
t0 = np.array([0])
_x, _y = np.meshgrid(x, y)
p0 = np.sin(2*np.pi*_x)*np.sin(2*np.pi*_y)
fig = plt.figure()
ax = fig.gca(projection='3d')
surf = ax.plot_surface(_x, _y, p0.reshape((len(_y),len(_x))), cmap=cm.coolwarm, linewidth=0, antialiased=False)
fig.colorbar(surf, shrink=0.5, aspect=5)
plt.show()
# +
from nangs import Dirichlet, Periodic
# initial condition
initial_condition = Dirichlet({'x': x, 'y': y, 't': t0}, {'p': p0.reshape(-1)}, device=device, name="initial")
pde.add_boco(initial_condition)
# periodic bocos
x1 = np.array([0.])
x2 = np.array([1.])
periodic1 = Periodic({'x': x1, 'y': y, 't': t}, {'x': x2, 'y': y, 't': t}, name="boco_x", device=device)
periodic2 = Periodic({'x': x, 'y': x1, 't': t}, {'x': x, 'y': x2, 't': t}, name="boco_y", device=device)
pde.add_boco(periodic1)
pde.add_boco(periodic2)
# +
from nangs import MLP
BATCH_SIZE = 256
LR = 1e-2
EPOCHS = 30
NUM_LAYERS = 3
NUM_HIDDEN = 128
mlp = MLP(len(pde.inputs), len(pde.outputs), NUM_LAYERS, NUM_HIDDEN).to(device)
optimizer = torch.optim.Adam(mlp.parameters())
scheduler = torch.optim.lr_scheduler.OneCycleLR(optimizer, max_lr=LR, pct_start=0.1, total_steps=EPOCHS)
pde.compile(mlp, optimizer, scheduler)
# %time hist = pde.solve(EPOCHS, BATCH_SIZE)
# +
t = 0.912312
x = np.linspace(0,1,100)
y = np.linspace(0,1,100)
_x, _y = np.meshgrid(x, y)
p0 = np.sin(2*np.pi*(_x - U*t))*np.sin(2*np.pi*(_y - V*t))
eval_mesh = Mesh({'x': x, 'y': y, 't': t}, device=device)
p = pde.eval(eval_mesh)
p = p.cpu().numpy()
fig = plt.figure(figsize=(15,5))
ax1 = fig.add_subplot(1, 2, 1, projection='3d')
ax2 = fig.add_subplot(1, 2, 2, projection='3d')
ax1.plot_surface(_x, _y, p0.reshape((len(_y),len(_x))), cmap=cm.coolwarm, linewidth=0, antialiased=False)
ax2.plot_surface(_x, _y, p.reshape((len(_y),len(_x))), cmap=cm.coolwarm, linewidth=0, antialiased=False)
plt.show()
print("L2 error: ", np.sqrt(sum((p0.ravel()-p.ravel())**2)))
| examples/02_adv2d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Find the markdown blocks that say interaction required! The notebook should take care of the rest!
# # Import libs
# +
import sys
import os
from eflow.foundation import DataPipeline,DataFrameTypes
# from eflow.model_analysis import ClassificationAnalysis
from eflow.utils.modeling_utils import optimize_model_grid
from eflow.utils.eflow_utils import get_type_holder_from_pipeline, remove_unconnected_pipeline_segments
from eflow.utils.pandas_utils import data_types_table
from eflow.utils.sys_utils import get_all_directories_from_path
from eflow.decision_bounds.visualize_multidimensional import VisualizeMultidimensional
import pandas as pd
import numpy as np
import scikitplot as skplt
import matplotlib.pyplot as plt
from sklearn.tree import DecisionTreeClassifier
from sklearn.model_selection import train_test_split
import copy
import pickle
from IPython.display import clear_output
from plotly.offline import init_notebook_mode
from plotly.offline import plot as plotly_html_plot
# -
from sklearn.metrics import mean_absolute_error
# !pip uninstall scipy --yes
# +
# # Additional add ons
# # !pip install pandasgui
# # !pip install pivottablejs
# clear_output()
# -
# %matplotlib notebook
# %matplotlib inline
# ## Declare Project Variables
# ### Interaction required
# +
dataset_path = "Datasets/train.csv"
# -----
dataset_name = "Home Insurance Cross Sell Prediction"
pipeline_name = "Home Insurance Pipeline"
# -----
# -----
notebook_mode = True
# -
# ## Clean out segment space
remove_unconnected_pipeline_segments()
# # Import dataset
df = pd.read_csv(dataset_path)
shape_df = pd.DataFrame.from_dict({'Rows': [df.shape[0]],
'Columns': [df.shape[1]]})
display(shape_df)
display(df.head(30))
data_types_table(df)
# # Loading and init df_features
# +
# Option: 1
# df_features = get_type_holder_from_pipeline(pipeline_name)
# -
# Option: 2
df_features = DataFrameTypes()
df_features.init_on_json_file(os.getcwd() + f"/eflow Data/{dataset_name}/df_features.json")
df_features.display_features(display_dataframes=True,
notebook_mode=notebook_mode)
# # Any extra processing before eflow DataPipeline
# # Setup pipeline structure
# ### Interaction Required
main_pipe = DataPipeline(pipeline_name,
df,
df_features)
main_pipe.perform_pipeline(df,
df_features)
df = df[0:1000]
df["Response"].value_counts()
X = df.drop(columns=df_features.target_feature()).values
y = df[df_features.target_feature()].values
feature_order = list(df.columns)
feature_order.remove(df_features.target_feature())
vis = VisualizeMultidimensional(X,
feature_names=feature_order,
dataset_name=dataset_name,
dataset_sub_dir="3D Vis",
overwrite_full_path=None,
notebook_mode=True,
pca_perc=.95)
scaled = vis.get_scaled_data()
indexes = vis.get_ordered_dp_indexes()
# +
import plotly.graph_objects as go
import random
import pandas as pd
# Read data from a csv
fig = go.Figure(data=[go.Surface(z=scaled[indexes])])
fig.update_traces(contours_z=dict(show=True, usecolormap=False,
project_z=True,))
fig.update_layout(title='3D', autosize=False,
scene_camera_eye=dict(x=1.87, y=0.88, z=-0.64),
width=1000, height=1000,
margin=dict(l=65, r=50, b=65, t=90),
)
fig.show()
# -
scaled.shape
# +
import plotly.graph_objects as go
import pandas as pd
# Read data from a csv
z_data = pd.read_csv('https://raw.githubusercontent.com/plotly/datasets/master/api_docs/mt_bruno_elevation.csv')
fig = go.Figure(data=[go.Surface(z=z_data.values)])
fig.update_traces(contours_z=dict(show=True, usecolormap=True,
highlightcolor="limegreen", project_z=True))
fig.update_layout(title='Mt Bruno Elevation', autosize=False,
scene_camera_eye=dict(x=1.87, y=0.88, z=-0.64),
width=500, height=500,
margin=dict(l=65, r=50, b=65, t=90)
)
fig.show()
# -
z_data
import eflow
help(eflow)
| testing/Home Ins/.ipynb_checkpoints/Visualize 3D Test-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Fast Image Processing with MXNet
#
# Previous tutorials have shown two ways of preprocessing images:
# - `mx.io.ImageRecordIter` is fast but inflexible. It is great for simple tasks like image recognition but won't work for more complex tasks like detection and segmentation.
# - `mx.recordio.unpack_img` (or `cv2.imread`, `skimage`, etc) + `numpy` is flexible but slow. The root of the problem is python's Global Interpreter Lock (GIL). GIL is a complicated topic but the gist is python doesn't really support multi-threading; even if you spawn multiple threads execution will still be serialized. You can workaround it with multi-processing and message passing, but it's hard to program and introduces overhead.
#
# To solve this issue, MXNet provides `mx.image` package. It stores images in [mx.nd.NDArray](./ndarray.ipynb) format and leverages MXNet's [dependency engine](http://mxnet.io/architecture/note_engine.html) to automatically parallelize image processing and circumvent GIL.
#
# Please read [Intro to NDArray](./ndarray.ipynb) first if you are not familar with it.
#
# Setup environment:
# %matplotlib inline
from __future__ import print_function
import os
import time
# set the number of threads you want to use before importing mxnet
os.environ['MXNET_CPU_WORKER_NTHREADS'] = '4'
import mxnet as mx
import numpy as np
import matplotlib.pyplot as plt
import cv2
# download example images
os.system('wget http://data.mxnet.io/data/test_images.tar.gz')
os.system('tar -xf test_images.tar.gz')
# ## Image Loading
# First we load images with mx.image.imdecode. The interface is very similar to opencv. But everything runs in parallel.
#
# We also compare performance agains opencv. You can restart the kernel and change MXNET_CPU_WORKER_NTHREADS to see the performance improvement as more threads are used.
# opencv
N = 1000
tic = time.time()
for i in range(N):
img = cv2.imread('test_images/ILSVRC2012_val_00000001.JPEG', flags=1)
img = cv2.cvtColor(img, cv2.COLOR_BGR2RGB)
print(N/(time.time()-tic), 'images decoded per second with opencv')
plt.imshow(img); plt.show()
# mx.image
tic = time.time()
for i in range(N):
img = mx.image.imdecode(open('test_images/ILSVRC2012_val_00000001.JPEG','rb').read())
mx.nd.waitall()
print(N/(time.time()-tic), 'images decoded per second with mx.image')
plt.imshow(img.asnumpy()); plt.show()
# ## Image Transformations
#
# Once images are loaded as NDArray, you can then use `mx.nd.*` operators to transform them. mx.image provides utility functions for some typical transformations:
# resize to w x h
tmp = mx.image.imresize(img, 100, 70)
plt.imshow(tmp.asnumpy()); plt.show()
# resize shorter edge to size while preserving aspect ratio
tmp = mx.image.resize_short(img, 100)
plt.imshow(tmp.asnumpy()); plt.show()
# crop a random w x h region from image
tmp, coord = mx.image.random_crop(img, (150, 200))
print(coord)
plt.imshow(tmp.asnumpy()); plt.show()
# Other utility functions include `fixed_crop`, `center_crop`, `color_normalize`, and `random_size_crop`.
# ## ImageIter
#
# Given the above functionalities, it's easy to write a custom data iterator. As an example, we provide `mx.image.ImageIter`, which is similar to Torch's [resnet](https://github.com/facebook/fb.resnet.torch/blob/master/dataloader.lua) image loading pipeline. For more details, please see `mxnet/python/mxnet/image.py`.
| python/basic/advanced_img_io.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # [Learn Quantum Computing with Python and Q#](https://www.manning.com/books/learn-quantum-computing-with-python-and-q-sharp?a_aid=learn-qc-granade&a_bid=ee23f338)<br>Chapter 6 Exercise Solutions
# ----
# > Copyright (c) <NAME> and <NAME>.
# > Code sample from the book "Learn Quantum Computing with Python and Q#" by
# > <NAME> and <NAME>, published by Manning Publications Co.
# > Book ISBN 9781617296130.
# > Code licensed under the MIT License.
# ### Exercise 6.1
# **Change the definition of `HelloWorld` to say your name instead of "classical world," and then run `%simulate` again using your new definition.**
function HelloWorld() : Unit {
Message("Hello, Sarah!");
}
%simulate HelloWorld
# ----
# ### Exercise 6.2
# **Use the `%simulate` magic command to run the `NextRandomBit` operation a few times; do you get the results you'd expect?**
operation NextRandomBit() : Result {
using (qubit = Qubit()) {
H(qubit);
return M(qubit);
}
}
%simulate NextRandomBit
%simulate NextRandomBit
%simulate NextRandomBit
# ----
# ### Exercise 6.3
# **What's the type of your new definition of `NextRandomBit`?**
# +
open Microsoft.Quantum.Measurement;
operation NextRandomBit(
statePreparation : (Qubit => Unit)
) : Result {
using (qubit = Qubit()) {
statePreparation(qubit);
return Microsoft.Quantum.Measurement.MResetZ(qubit);
}
}
# -
# This version of `NextRandomBit` is an operation that takes one input of type `(Qubit => Unit)`, and that returns a single output of type `Result`.
# Thus, `NextRandomBit` has type `((Qubit => Unit) => Result)`.
# ----
# ### Exercise 6.4
# **Partial application works for functions as well as operations!
# Try it out by writing a function `Plus` that adds two integers, `n` and `m`, and another function `PartialPlus` that takes an input `n` and returns a function that adds `n` to its input.**
#
# *HINT*: You can get started using the following code snippet as a template.
#
# ```Q#
# function Plus(n : Int, m : Int) : Int {
# // fill in this part
# }
#
# function PartialPlus(n : Int) : (Int -> Int) {
# // fill in this part
# }
# ```
# +
function Plus(n : Int, m : Int) : Int {
return n + m;
}
function PartialPlus(n : Int) : (Int -> Int) {
return Plus(n, _);
}
# -
function Example() : Unit {
let plusTwo = PartialPlus(2);
let three = plusTwo(1);
let five = plusTwo(3);
Message($"three = {three}, five = {five}");
}
%simulate Example
# ----
# ### Epilogue
#
# _The following cell logs what version of the components this was last tested with._
%version
| ch06/ch06-exercise-solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Testing out the reparameterization trick
#
# Just a simple implementation to test if it will be appropriate for the GLM, if it is, we can use Auto-Encoding Variational Bayes inference.
#
# The basic premise is we can construct a differenctiable Monte-Carlo estimator,
# $$
# \mathbb{E}_{q(z)}[f(z)] = \int q_{\theta}(z|x) f(z) dz
# \approx \frac{1}{L} \sum^L_{l=1} f(g_{\theta}(x, \epsilon^{(l)})),
# $$
# where
# $$
# z^{(l)} = g_{\theta}(x, \epsilon^{(l)}) \qquad \text{and} \qquad \epsilon^{(l)} \sim p(\epsilon),
# $$
# that results in lower variance derivatives than Monte-Carlo sampling the derivatives using, e.g. variational black box methods.
#
# ## Test 1: $f(z)$ is a log-Normal
#
# ### Likelihood approximation
#
# Let's start with a really simple example,
# $$
# \begin{align}
# f(z) &= \log \mathcal{N}(x|z, \sigma^2), \\
# q_\theta(z | x) &= \mathcal{N}(z | \mu, \lambda).
# \end{align}
# $$
# We can solve this integral analytically,
# $$
# \int \mathcal{N}(z | \mu, \lambda) \log \mathcal{N}(x|z, \sigma^2) dz
# = \log \mathcal{N}(x | \mu, \sigma^2) - \frac{\lambda}{2 \sigma^2}
# $$
# So we can test how this compares to the reparameterization trick results.
# lets use the following deterministic function for reparameterization,
# $$
# g_{(\mu, \lambda)}(\epsilon^{(l)}) = \mu + \sqrt{\lambda}\epsilon^{(l)}
# $$
# where
# $$
# p(\epsilon) = \mathcal{N}(0, 1)
# $$
# Now let's test:
# $$
# \log \mathcal{N}(x | \mu, \sigma^2) - \frac{\lambda}{2 \sigma^2} \stackrel{?}{\approx}
# \frac{1}{L} \sum^L_{l=1} \log \mathcal{N}(x|,g_{(\mu, \lambda)}(\epsilon^{(l)}), \sigma^2)
# $$
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as pl
pl.style.use('ggplot')
from scipy.stats import norm
from scipy.special import expit
from scipy.integrate import quadrature
from scipy.misc import derivative
from revrand.mathfun.special import softplus
from revrand.optimize import sgd, Adam
# -
# Initial values
x = 0
mu = 2
sigma = 3
lambd = 0.5
L = 50
# +
# The test
exact = norm.logpdf(x, loc=mu, scale=sigma) - lambd / (2 * sigma**2)
print("Exact expectation = {}".format(exact))
# Normal Monte Calo estimation
z = norm.rvs(loc=mu, scale=np.sqrt(lambd), size=(L,))
approx_mc = norm.logpdf(x, loc=z, scale=sigma)
print("MC Approx expectation = {} ({})".format(approx_mc.mean(), approx_mc.std()))
# Reparameterised Sampling
g = lambda e: mu + np.sqrt(lambd) * e
e = norm.rvs(loc=0, scale=1, size=(L,))
approx_re = norm.logpdf(x, loc=g(e), scale=sigma)
print("Reparameterized Approx expectation = {} ({})".format(approx_re.mean(), approx_re.std()))
# -
# We would expect a trivial relationship here between exact monte-carlo and the reparameterization trick, since they are doing the same thing. Lets see if gradient estimates have lower variances now.
#
# ### Gradient approximation
# Let's evaluate the exact gradient for $\mu$,
# $$
# \frac{\partial}{\partial \mu} \left(\log \mathcal{N}(x | \mu, \sigma^2) - \frac{\lambda}{2 \sigma^2} \right) =
# \frac{1}{\sigma^2} (x - \mu)
# $$
# Now the approximation
# $$
# \begin{align}
# \frac{\partial}{\partial \mu} \left(
# \frac{1}{L} \sum^L_{l=1} \log \mathcal{N}(x|,g_{(\mu, \lambda)}(\epsilon^{(l)}), \sigma^2) \right) &=
# \frac{1}{L} \sum^L_{l=1} \frac{1}{\sigma^2} (x - g_{(\mu, \lambda)}(\epsilon^{(l)}))
# \frac{\partial g_{(\mu, \lambda)}(\epsilon^{(l)})}{\partial \mu}, \\
# &= \frac{1}{L} \sum^L_{l=1} \frac{1}{\sigma^2} (x - g_{(\mu, \lambda)}(\epsilon^{(l)})).
# \end{align}
# $$
# +
# A range of mu's
N = 100
mu = np.linspace(-5, 5, N)
# Exact
dmu = (x - mu) / sigma**2
# Approx
e = norm.rvs(loc=0, scale=1, size=(L, N))
approx_dmu = (x - g(e)) / sigma**2
Edmu = approx_dmu.mean(axis=0)
Sdmu = approx_dmu.std(axis=0)
# plot
pl.figure(figsize=(15, 10))
pl.plot(mu, dmu, 'b', label='Exact')
pl.plot(mu, Edmu, 'r', label= 'Approx')
pl.fill_between(mu, Edmu - 2 * Sdmu, Edmu + 2 * Sdmu, edgecolor='none', color='r', alpha=0.3)
pl.legend()
pl.title("Derivatives of expected log Gaussian")
pl.xlabel('$\mu$')
pl.ylabel('$\partial f(z)/ \partial \mu$')
pl.show()
# -
# ## Test 2: $f(z)$ is log Bernoulli
#
# Now let's try the following function with the same posterior and $g$ as before,
#
# $$
# f(z) = \log \text{Bern}(x | \text{logistic}(z)) = x z - \log(1 + exp(z))
# $$
#
# We can get an "exact" expectation using quadrature. First of all, likelihoods,
#
# ### Likelihood Approximation
# +
# Quadrature
def qlogp(z, mu):
q = norm.pdf(z, loc=mu, scale=np.sqrt(lambd))
logp = x * z - softplus(z)
return q * logp
def quadELL(mu):
return quadrature(qlogp, a=-10, b=10, args=(mu,))[0]
ELL = [quadELL(m) for m in mu]
# Reparam
e = norm.rvs(loc=0, scale=1, size=(L, N))
approx_ELL = x * g(e) - softplus(g(e))
EELL = approx_ELL.mean(axis=0)
SELL = approx_ELL.std(axis=0)
# -
# plot
pl.figure(figsize=(15, 10))
pl.plot(mu, ELL, 'b', label='Quadrature')
pl.plot(mu, EELL, 'r', label= 'Approx')
pl.fill_between(mu, EELL - 2 * SELL, EELL + 2 * SELL, edgecolor='none', color='r', alpha=0.3)
pl.legend()
pl.title("ELL with log Bernoulli")
pl.xlabel('$\mu$')
pl.ylabel('$\mathbb{E}[\log Bern(x | z)]$')
pl.show()
# ### Gradient approximation
#
# $$
# \begin{align}
# \frac{\partial}{\partial \mu} \mathbb{E}_q \left[\frac{\partial f(z)}{\partial \mu} \right]
# &\approx \frac{1}{L} \sum^L_{l=1} (x - \text{logistic}(g(\epsilon^{(l)})))
# \frac{\partial g(\epsilon^{(l)})}{\partial \mu} \\
# &= \frac{1}{L} \sum^L_{l=1} x - \text{logistic}(g(\epsilon^{(l)}))
# \end{align}
# $$
# +
# Quadrature
dmu = [derivative(quadELL, m) for m in mu]
# Reparam
e = norm.rvs(loc=0, scale=1, size=(L, N))
approx_dmu = x - expit(g(e))
Edmu = approx_dmu.mean(axis=0)
Sdmu = approx_dmu.std(axis=0)
# -
# plot
pl.figure(figsize=(15, 10))
pl.plot(mu, dmu, 'b', label='Quadrature')
pl.plot(mu, Edmu, 'r', label= 'Approx')
pl.fill_between(mu, Edmu - 2 * Sdmu, Edmu + 2 * Sdmu, edgecolor='none', color='r', alpha=0.3)
pl.legend()
pl.title("Derivative of $\mu$ with log Bernoulli")
pl.xlabel('$\mu$')
pl.ylabel('$\partial f(z)/ \partial \mu$')
pl.show()
# ## Optimisation test
#
# Now let's see if we can optimise Expected log likelihood using SG!
#
# +
data = np.ones((100, 1), dtype=bool)
mu_rec, dmu_rec = [], []
def ell_obj(mu, x, samples=100):
e = norm.rvs(loc=0, scale=1, size=(samples,))
g = mu + np.sqrt(lambd) * e
ll = (x * g - softplus(g)).mean()
dmu = (x - expit(g)).mean()
mu_rec.append(float(mu))
dmu_rec.append(float(dmu))
return -ll, -dmu
res = sgd(ell_obj, x0=np.array([-4]), data=data, maxiter=1000, updater=Adam(), eval_obj=True)
# +
# plot
niter = len(mu_rec)
fig = pl.figure(figsize=(15, 10))
ax1 = fig.add_subplot(111)
ax1.plot(range(niter), res.norms, 'b', label='gradients')
ax1.plot(range(niter), res.objs, 'g', label='negative ELL')
ax1.set_ylabel('gradients/negative ELL')
ax1.legend()
for t in ax1.get_yticklabels():
t.set_color('b')
ax2 = ax1.twinx()
ax2.set_ylabel('$\mu$')
ax2.plot(range(niter), mu_rec, 'r', label='$\mu$')
for t in ax2.get_yticklabels():
t.set_color('r')
pl.show()
# -
| demos/reparameterization_trick.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from qutip import*
import numpy as np
import matplotlib.pyplot as plt
# # Bloch Sphere
Q=1; gamma=np.arctan(Q);t=np.linspace(0,2*np.pi,100)
p=len(t);
nx=np.cos(t)*np.sin(gamma)
ny=np.sin(t)*np.sin(gamma)
nz=np.cos(gamma)*np.ones(p)
N=[nx,ny,nz];
P=[nx,ny,-nz]
vx=np.cos(np.pi/2)*np.sin(gamma);
vy=np.sin(np.pi/2)*np.sin(gamma);
vz=np.cos(gamma);
vp=[vx,vy,vz];
vn=[vx,-vy,-vz];
# +
b = Bloch(view=[-60,15]);
b.add_points(N,'l');
b.add_points(P,'l');
b.add_vectors(vp)
b.add_vectors(vn)
b.frame_alpha=0.1; b.frame_num=2
b.sphere_alpha=0.08
b.vector_color=['b','r']; b.vector_width=2
b.add_annotation([0,0.1,0.2],'$\gamma $');
b.make_sphere()
plt.savefig('AnilloNA.png',dpi=600,bbox_inches='tight')
# +
gamma=np.pi/2;t=np.linspace(0,2*np.pi,100)
p=len(t);
nx=np.cos(t)*np.sin(gamma)
ny=np.sin(t)*np.sin(gamma)
nz=np.cos(gamma)*np.ones(p)
N=[nx,ny,nz];
P=[nx,ny,-nz]
vx=np.cos(np.pi/2)*np.sin(gamma);
vy=np.sin(np.pi/2)*np.sin(gamma);
vz=np.cos(gamma);
vp=[vx,vy,vz];
vn=[vx,-vy,-vz];
# +
b = Bloch(view=[-60,15]);
b.add_points(N,'l');
#b.add_points(P,'l');
b.add_vectors(vp)
b.add_vectors(vn)
b.frame_alpha=0.1; b.frame_num=2
b.sphere_alpha=0.08
b.vector_color=['b','r']; b.vector_width=2
b.add_annotation([0,0.1,0.2],'$\gamma=\pi/2$');
b.make_sphere()
plt.savefig('AnilloA.png',dpi=600,bbox_inches='tight')
# +
Q=1; gamma=np.arctan(Q);t=np.linspace(0,2*np.pi,100)
p=len(t);
nx=np.cos(t)*np.sin(gamma)
ny=np.sin(t)*np.sin(gamma)
nz=np.cos(gamma)*np.ones(p)
N=[nx,ny,nz];
vx=np.cos(np.pi/2)*np.sin(gamma);
vy=np.sin(np.pi/2)*np.sin(gamma);
vz=np.cos(gamma);
vp=[vx,vy,vz];
b = Bloch(view=[-60,15]);
b.add_points(N,'l');
b.add_vectors(vp)
b.frame_alpha=0.1; b.frame_num=2
b.sphere_alpha=0.08
b.vector_color=['b','r']; b.vector_width=2
b.make_sphere()
plt.savefig('EjemploFAA.png',dpi=600,bbox_inches='tight')
# -
# # Phase Graphs
# +
QR=np.linspace(0,10,20); gamma=np.arctan(QR)
fDA=QR*np.sin(gamma)/2
fGA=(1-np.cos(gamma))/2
fTA=fDA+fGA
# +
fig, (ax1,ax2,ax3) = plt.subplots(nrows=3, ncols=1,sharex=True)
ax1.plot(QR,fTA,'+r')
ax1.set_ylabel('$\delta/2\pi$')
ax1.set_title('')
ax2.plot(QR,fDA,'+y')
ax2.set_ylabel('$\delta_{d}/2\pi$')
ax3.plot(QR,fGA,'+c')
ax3.set_ylabel('$\delta_{g}/2\pi$')
ax3.set_xlabel('$Q_{R}$')
plt.savefig('Fases.eps' ,dpi=600,bbox_inches='tight')
| GraficasTFG.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Natural Language Processing (NLP)
# ### University of California, Berkeley - Spring 2022
# ## Today's subjects:
#
# - Natural Language Processing (NLP)
# - Gensim's introduction
# - Word2Vec
# Natural language processing (NLP) is one of the most important technologies of the information age. Understanding complex language utterances is also a crucial part of artificial intelligence. Applications of NLP are everywhere because people communicate most everything in language: web search, advertisement, emails, customer service, language translation, radiology reports, etc. There are a large variety of underlying tasks and machine learning models behind NLP applications. In recent years, deep learning approaches have obtained very high performance across many different NLP tasks. These can solve tasks with single end-to-end models and do not require traditional, task-specific feature engineering. In this winter quarter course students will learn to implement, train, debug, visualize and invent their own neural network models. The course provides a thorough introduction to cutting-edge research in deep learning applied to NLP. On the model side we will cover word vector representations, window-based neural networks, recurrent neural networks, long-short-term-memory models, recursive neural networks, convolutional neural networks as well as some recent models involving a memory component. Through lectures and programming assignments students will learn the necessary engineering tricks for making neural networks work on practical problems. In this course, students will gain a thorough introduction to cutting-edge research projects in Deep Learning with NLP.
# 
#
# Reference: https://chatbotslife.com/natural-language-principles-65e88e20b94
# __NOTE__: This notebooks requires basic knowledge of Python. If you are not familiar with Python, checkout this course's introduction to python notebooks.
# There are different levels of tasks in NLP, from speech processing to semantic interpretation and discourse processing. The goal of NLP is to be able to design algorithms to allow computers to "understand" the natural language in order to perform some task. Some of the tasks are listed as following:
#
# - Spell Checking
# - Keyword Search
# - Finding Synonyms
# - Parsing information from websites, documents, etc
# - Machine Translation (e.g. English to French Translation)
# - Semantic Analysis (e.g. What is the meaning of this statement?)
# - Question Answering
# ## Word Representations
# The first and arguably most important common denominator across all NLP tasks is how we represent words as input to any of our models. To perform well on most NLP tasks we first need to have some notion of similarity and difference between words. We can encode words into __word vectors__. A word vector is supposed to contain the meaning of the word and represent the word as a point in a multi-dimensional space.
#
# There are an estimated 13 million tokens for the English language. Are they all completely unrelated? Ofcourse they are related. Thus, we want to encode word tokens each into some numeric vector that represents a point in multi-dimensional space which we'll call "word" space. One intuition is that perhaps there actually exists some N-dimensional space (such that N << 13 million) that is sufficient to encode all semantics of our language. Each dimension would encode some meaning that we transfer using speech. For instance, some dimensions will indicate tense (past vs. present vs. future), count (singular vs. plural),
# and gender (masculine vs. feminine).
#
# There are a number of methods to calculate word vectors:
#
# - One-hot vectors
# - SVD based Methods (Out of context of this course)
# - Iteration Based Methods
# * Language Models (N-grams)
# * Continuous Bag of Words (CBOW)
# * SkipGram Model
# ### One-hot Vectors
# In this simple approach, each word vector has the length of the vocabulary (which is really really high!). we will represent each word with a vector of all zeros and one 1 at the index of the word in the sorted english language. As can be inferred from this word vectors, the distance between each word is equal to distance of any other pair of words. Also, each pair of words dot product will result to zero, meaning that we represent each word as a complete independent entity in our "word" space. In this "word" space, Each dimension represents a different word. Indeed, each word is a unit vector streched in the direction its corresponding dimension in the "word" space.
#
#
# As can be seen, the main issue with this approach is that the dimensionality of the "word" space is too high. We can find a much much smaller subspace that encodes the relationship between the words.
# ### Iteration Based Models
# In another well known approach in encoding words into a "word" space, which is "iteration based" family of models, we can try to create a model that will be able to learn one iteration at a time and eventually be able to encode the probability of a word given its context. One of the first and famous approaches is called `Word2Vec`. We will give you a brief introduction about word2vec algorithm in the following sections.
# #### Word2Vec
# The Word2Vec's idea is to design a model whose parameters are the word vectors. Then, train the model on a certain objective. At every iteration
# we run the model, evaluate the errors, and follow an update rule that has some notion of penalizing the model parameters that caused the error. Thus, we learn our word vectors. Word2Vec is developed by Mikolov et. al which includes 2 algorithms:
#
# - Continuous Bag Of Words (CBOW)
# - Skip-gram
#
# Consider this example for the rest of this lecture:
# <center>"The cat jumped over the puddle."</center>
# #### Iteration Based Models: CBOW
# <center>
# <img src="./CBOW.jpg" width="500"/>
# </center>
#
# Reference: https://srishtee-kriti.medium.com/mathematics-behind-continuous-bag-of-words-cbow-model-1e54cc2ecd88
# CBOW aims to predict a center word from the surrounding context in terms of word vectors. In this approach, we treat {"The", "cat", ’over", "the’, "puddle"} as a context and from these words, we aim to predict or generate the center word "jumped". This type of model we call a Continuous Bag of Words (CBOW) Model.
# + [markdown] tags=[]
# #### Iteration Based Models: SkipGram
# -
# <center>
# <img src="./skipgram.png" width=500/>
# </center>
#
# Reference: https://www.researchgate.net/figure/The-architecture-of-Skip-gram-model-20_fig1_322905432
# Skip-gram does the opposite, and predicts the distribution (probability) of context words from a center word. In this approach the goa to create a model such that given the center word "jumped", the model will be able to predict or generate the surrounding words "The", "cat", "over", "the", "puddle". Here we call the word "jumped" the context. We call this type of model a SkipGram model.
# <center>
# <img src="./skipgram-2.png"/>
# <center><p><center><a href="https://trailhead.salesforce.com/fr/content/learn/modules/word-meaning-and-word2vec/get-started-with-word2vec">Word2Vec Skipgram model with window size 2</a></center></p></center>
# </center>
#
# + [markdown] tags=[]
# ## Gensim installation
# -
# We'll use `gensim` package in order to gain some basic insights about NLP. In order to install `gensim`, run the following cell:
# !pip install gensim
# ## Let's start with Word Vectors
# Word Vectors (a.k.a. word embeddings) are often used as a fundamental component for downstream NLP tasks, e.g. question answering, text generation, translation, etc., so it is important to build some intuitions as to their strengths and weaknesses.
#
# Here, you will explore two types of word vectors: those derived from co-occurrence matrices, and those derived via GloVe.
# +
import numpy as np
# Get the interactive Tools for Matplotlib
# %matplotlib inline
import matplotlib.pyplot as plt
plt.style.use('ggplot')
plt.rcParams['figure.dpi'] = 150
from sklearn.decomposition import PCA
from gensim.test.utils import datapath, get_tmpfile
from gensim.models import KeyedVectors
from gensim.scripts.glove2word2vec import glove2word2vec
# -
# For looking at word vectors, I'll use Gensim. We also use it in hw1 for word vectors. Gensim isn't really a deep learning package. It's a package for for word and text similarity modeling, which started with (LDA-style) topic models and grew into SVD and neural word representations. But its efficient and scalable, and quite widely used.
#
# Our homegrown Stanford offering is GloVe word vectors. Gensim doesn't give them first class support, but allows you to convert a file of GloVe vectors into word2vec format. You can download the GloVe vectors from the Glove page. They're inside this zip file
#
# (I use the 100d vectors below as a mix between speed and smallness vs. quality. If you try out the 50d vectors, they basically work for similarity but clearly aren't as good for analogy problems. If you load the 300d vectors, they're even better than the 100d vectors.)
glove_file = datapath('/home/mohsen/data/glove.6B.100d.txt')
word2vec_glove_file = get_tmpfile("glove.6B.100d.word2vec.txt")
glove2word2vec(glove_file, word2vec_glove_file)
model = KeyedVectors.load_word2vec_format(word2vec_glove_file)
model.most_similar('clinton')
model.most_similar('apple')
model.most_similar(negative='apple')
result = model.most_similar(positive=['woman', 'king'], negative=['man'])
print("{}: {:.4f}".format(*result[0]))
def analogy(x2, x1, y1):
result = model.most_similar(positive=[y1, x2], negative=[x1])
return result[0][0]
analogy('iranian', 'iran', 'america')
analogy('tallest', 'tall', 'long')
analogy('fantastic', 'good', 'bad')
analogy('waitress', 'waiter', 'actor')
len(model.key_to_index.keys())
def display_pca_scatterplot(model, words=None, sample=0):
if words == None:
if sample > 0:
words = np.random.choice(list(model.key_to_index.keys()), sample)
else:
words = [ word for word in model.vocab ]
word_vectors = np.array([model[w] for w in words])
twodim = PCA().fit_transform(word_vectors)[:,:2]
plt.figure(figsize=(6,6))
plt.scatter(twodim[:,0], twodim[:,1], edgecolors='k', c='r')
for word, (x,y) in zip(words, twodim):
plt.text(x+0.05, y+0.05, word)
plt.show()
display_pca_scatterplot(model,
['man', 'woman', 'king', 'queen'])
display_pca_scatterplot(model,
['coffee', 'tea', 'beer', 'wine', 'brandy', 'rum', 'champagne', 'water',
'spaghetti', 'borscht', 'hamburger', 'pizza', 'falafel', 'sushi', 'meatballs',
'dog', 'horse', 'cat', 'monkey', 'parrot', 'koala', 'lizard',
'frog', 'toad', 'monkey', 'ape', 'kangaroo', 'wombat', 'wolf',
'france', 'germany', 'hungary', 'luxembourg', 'australia', 'fiji', 'china',
'homework', 'assignment', 'problem', 'exam', 'test', 'class',
'school', 'college', 'university', 'institute'])
display_pca_scatterplot(model, sample=5)
display_pca_scatterplot(model, sample=10)
display_pca_scatterplot(model, sample=50)
display_pca_scatterplot(model, sample=100)
# ## Congrats!
#
# The notebook is available at https://github.com/Naghipourfar/molecular-biomechanics/
| deep learning/Word2Vec.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Space Report
#
#
# <img src="images/polito_logo.png" alt="Polito Logo" style="width: 200px;"/>
#
#
# ## Pittsburgh Bridges Data Set
#
# <img src="images/andy_warhol_bridge.jpg" alt="Andy Warhol Bridge" style="width: 200px;"/>
#
# Andy Warhol Bridge - Pittsburgh.
#
# Report created by Student <NAME> s253666, for A.A 2019/2020.
#
# **Abstract**:The aim of this report is to evaluate the effectiveness of distinct, different statistical learning approaches, in particular focusing on their characteristics as well as on their advantages and backwards when applied onto a relatively small dataset as the one employed within this report, that is Pittsburgh Bridgesdataset.
#
# **Key words**:Statistical Learning, Machine Learning, Bridge Design.
# ### Imports Section <a class="anchor" id="imports-section"></a>
from utils.all_imports import *;
# %matplotlib inline
# Set seed for notebook repeatability
np.random.seed(0)
# +
# READ INPUT DATASET
# =========================================================================== #
dataset_path, dataset_name, column_names, TARGET_COL = get_dataset_location()
estimators_list, estimators_names = get_estimators()
dataset, feature_vs_values = load_brdiges_dataset(dataset_path, dataset_name)
# -
columns_2_avoid = ['ERECTED', 'LENGTH', 'LOCATION']
# Make distinction between Target Variable and Predictors
# --------------------------------------------------------------------------- #
rescaledX, y, columns = prepare_data_for_train(dataset, target_col=TARGET_COL)
# ## Pricipal Component Analysis
show_table_pc_analysis(X=rescaledX)
# #### Major Pros & Cons of PCA
#
#
# ## Learning Models <a class="anchor" id="learning-models"></a>
# +
# Parameters to be tested for Cross-Validation Approach
# -----------------------------------------------------
plots_names = list(map(lambda xi: f"{xi}_learning_curve.png", estimators_names))
pca_kernels_list = ['linear', 'poly', 'rbf', 'cosine', 'sigmoid']
cv_list = list(range(10, 1, -1))
param_grids = []
parmas_logreg = {
'penalty': ('l1', 'l2', 'elastic', None),
'solver': ('newton-cg', 'lbfgs', 'liblinear', 'sag', 'saga'),
'fit_intercept': (True, False),
'tol': (1e-4, 1e-3, 1e-2),
'class_weight': (None, 'balanced'),
'C': (10.0, 1.0, .1, .01, .001, .0001),
# 'random_state': (0,),
}; param_grids.append(parmas_logreg)
parmas_knn_clf = {
'n_neighbors': (2,3,4,5,6,7,8,9,10),
'weights': ('uniform', 'distance'),
'metric': ('euclidean', 'minkowski', 'manhattan'),
'leaf_size': (5, 10, 15, 30),
'algorithm': ('ball_tree', 'kd_tree', 'brute'),
}; param_grids.append(parmas_knn_clf)
params_sgd_clf = {
'loss': ('log', 'modified_huber'), # ('hinge', 'log', 'modified_huber', 'squared_hinge', 'perceptron')
'penalty': ('l2', 'l1', 'elasticnet'),
'alpha': (1e-1, 1e-2, 1e-3, 1e-4),
'max_iter': (50, 100, 150, 200, 500, 1000, 1500, 2000, 2500),
'class_weight': (None, 'balanced'),
'learning_rate': ('optimal',),
'tol': (None, 1e-2, 1e-4, 1e-5, 1e-6),
'random_state': (0,),
}; param_grids.append(params_sgd_clf)
kernel_type = 'svm-rbf-kernel'
params_svm_clf = {
'gamma': (1e-7, 1e-4, 1e-3, 1e-2, 0.1, 1.0, 10, 1e+2, 1e+3, 1e+5, 1e+7),
'max_iter':(1e+2, 1e+3, 2 * 1e+3, 5 * 1e+3, 1e+4, 1.5 * 1e+3),
'degree': (1,2,3,4,5),
'coef0': (.001, .01, .1, 0.0, 1.0, 10.0),
'shrinking': (True, False),
'kernel': ['linear', 'poly', 'rbf', 'sigmoid',],
'class_weight': (None, 'balanced'),
'C': (1e-4, 1e-3, 1e-2, 0.1, 1.0, 10, 1e+2, 1e+3),
'probability': (True,),
}; param_grids.append(params_svm_clf)
parmas_tree = {
'splitter': ('random', 'best'),
'criterion':('gini', 'entropy'),
'max_features': (None, 'sqrt', 'log2'),
'max_depth': (None, 3, 5, 7, 10,),
'splitter': ('best', 'random',),
'class_weight': (None, 'balanced'),
}; param_grids.append(parmas_tree)
parmas_random_forest = {
'n_estimators': (3, 5, 7, 10, 30, 50, 70, 100, 150, 200),
'criterion':('gini', 'entropy'),
'bootstrap': (True, False),
'min_samples_leaf': (1,2,3,4,5),
'max_features': (None, 'sqrt', 'log2'),
'max_depth': (None, 3, 5, 7, 10,),
'class_weight': (None, 'balanced', 'balanced_subsample'),
}; param_grids.append(parmas_random_forest)
# Some variables to perform different tasks
# -----------------------------------------------------
N_CV, N_KERNEL, N_GS = 9, 5, 6;
nrows = N_KERNEL // 2 if N_KERNEL % 2 == 0 else N_KERNEL // 2 + 1;
ncols = 2; grid_size = [nrows, ncols]
# -
# | Learning Technique | Type of Learner | Type of Learning |Classification | Regression | Clustering |
# | --- | --- | --- | --- | --- | --- |
# | *K-Nearest Neighbor* | *Instance-based or Non-generalizing* | *Supervised and Usupervised Learning* | *Supported* | *Supported* | *Supported*|
n_components=9
learning_curves_by_kernels(
# learning_curves_by_components(
estimators_list[:], estimators_names[:],
rescaledX, y,
train_sizes=np.linspace(.1, 1.0, 10),
n_components=9,
pca_kernels_list=pca_kernels_list[0],
verbose=0,
by_pairs=True,
savefigs=True,
scoring='accuracy',
figs_dest=os.path.join('figures', 'learning_curve', f"Pcs_{n_components}"), ignore_func=True,
# figsize=(20,5)
)
# + language="javascript"
# IPython.OutputArea.prototype._should_scroll = function(lines) {
# return false;
# }
# +
plot_dest = os.path.join("figures", "n_comp_9_analysis", "grid_search")
X = rescaledX; pos = 2
df_gs, df_auc_gs, df_pvalue = grid_search_all_by_n_components(
estimators_list=estimators_list[pos], \
param_grids=param_grids[pos-1],
estimators_names=estimators_names[pos], \
X=X, y=y,
n_components=9,
random_state=0, show_plots=False, show_errors=False, verbose=1, plot_dest=plot_dest, debug_var=False)
df_9, df_9_auc = df_gs, df_auc_gs
# -
# Looking at the results obtained running *Knn Classifier* against our dataset splitted into training set and test set and adopting a different kernel trick applied to *kernel-Pca* unsupervised preprocessing method we can state generally speaking that all the such a *Statistical Learning technique* leads to a sequence of results that on average are more than appreciable beacues of the accuracy scores obtained at test time which compared against the same score but related to train phase allows us to understand that during the model creation the resulting classifiers do not overfit to the data, and even when the training score was high it was still lower than the scores in terms of accuracy obtained from the Logisti Regression Models which overfit to the data. Moreover, looking at the weighted values of *Recall, Precision, and F1-Scores* we can notably claim that the classifiers based on Knn obtained good performance and and except for one trial where we got lower and worst results, when *Sigmoid Trick* is selected, in the remaning cases have gotten remarkable results. More precisely we can say what follows:
# - speaking about __Linear kernel Pca based Knn Classifier__, when adoping the default threshold of *.5* for classification purposes we have a model that reaches an accuracy of *71%* at test time against an accuracy of *92%* at train step, while the Auc score reaches a value of *76%* with a Roc Curve that shows a behavior for which the model for a first set of thresholds let *TPR* grows faster than *FPR*, and only when we take into account larger thresholds we can understand that the trend is reversed. Looking at classification report we can see that the model has high precision and recall for class 1, so this means that the classifier has high confidence when predicting class 1 labels, instead it is less certain when predicting class 0 instances because has low precision, even if the model was able to predict correctly all the samples from class 0, leading to high recall.
# - observing __Polynomial kernel Pca based Knn Estimator__, we can notice that such a model exploiting a default threshold of *.5* reaches an accuracy of *74%* at test time against an accuracy of *89%* at train step, while the Auc score reaches a value of *77%*. What we can immediately understand is that the second model we have trained for Knn classifier is able to better generalize because obtained a higher accuracy score for test set which is also less far from train accuracy score, moreover the model has a slightly greater precision and recall when referring to class 1, while the precision and recall about class 0 seem to be more or less the same.
# - review __Rbf kernel Pca based Knn Classifier__, we can notice that such a model exploiting a default threshold of *.5* reaches an accuracy of *79%* at test time against an accuracy of *95%* at train step, while the Auc score reaches a value of *74%*. We can understand that even if this model, having selected *Rbf kernel Trick for kernel-Pca*, corresponds to the estimator that among Knn classifiers is the one with the best performance in terms of accuracy we notice that the corresponding auc score is less than the other first two analyzed trials where we end up saying that such classifiers lead to acceptable results. However, this method set with the hyper-params found by the grid-search algorithm reveals a higher value of precision related to class 0, meaning that *Rbf kernel Pca based Logisti Classifier* has a higher precision than previous models when classifying instances as belonging to class 0, while precisin and recall metrics for class 1 was more or less the same. This classifier is the one that we should select since it has higher precision values for better classifyng new instances.
# - looking at __Cosine kernel Pca based Knn Classifier__, we can notice that such a model exploiting a default threshold of *.5* reaches an accuracy of *59%* at test time against an accuracy of *92%* at train step, while the Auc score reaches a value of *62%*. We can clearly see that that such a model corresponds to the worst solution amongst the models we have trained exploiting *Knn classifier*, because we can state that due to a mostly lower accuracy score obtained at test time than the accuracy score referred to training time the classifier seems to have overfit to the data. In particular speaking about Precision and Recall scores about class 1, from classification report, the model seems to be mostly precise when predicting class 1 as label for the instances to be classified, however was misclassifying nearly half of the samples from class 1. Furthermore, the model also do not obtain fine results in terms of metrics when we focus on class 0 precision and recall. This model is the oen we should avoid, and do not exploit.
# - finally, referring to __Sigmoid kernel Pca based Knn Model__, we can notice that such a model exploiting a default threshold of *.5* reaches an accuracy of *65%* at test time against an accuracy of *92%* at train step, while the Auc score reaches a value of *72%*. It has values for performance metrics such as *precisio, recall, and F1-Score* which are more or less quyite similar to those of the first models trained for Knn classifier, that are those which are exploiting lienar and polynomial tricks, however this model misclassifyes larger number of class 1 instances lowering down the precision related to class 0, as well as lowering down recall from class 1. Also such a classifier with respect the first three trial is not sufficiently fine to be accepted so that we can exclude it from our choice.
# create_widget_list_df([df_gs, df_auc_gs]) #print(df_gs); print(df_auc_gs)
show_table_summary_grid_search(df_gs, df_auc_gs, df_pvalue)
# Looking at the table dispalyed just above that shows the details about the selected values for hyper-parameters specified during grid search, in the different situations accordingly to the fixed kernel-trick for kernel Pca unsupervised method we can state that, referring to the first two columns of *Train and Test Accuracy*, we can recognize which trials lead to more overfit results such as for *Cosine, and Sigmoid Tricks* or less overfit solution such as in the case of *Linear, Polynomial, and Rbf Trick*. Speaking about the hyper-parameters, we can say what follows:
# - looking at the *algorithm paramter*, which can be set alternatively as *brute, kd-tree, and ball-tree* where each algorithm represents an differen strategy for implementing neighbor-based learning with pros and cons in terms of requested training time, memory usage and inference performance in terms of elapsed time, we can clearly understand that the choice of the kind of kernel trick for performing kernel-Pca does not care since all the trials preferred and selectd *ball-tree* strategy to solve the problem. It means that the grid search algorithm, when forced to try all the possible combination recorded such a choice as the best hyper-param which leads to building an expensive data strucuture which aims at integrating somehow distance information to achieve better performance scores. So, this should make us reason about the fact that it is still a good choice or we should re-run the procedure excluding such algorithm. In fact the answer to such a issue depend on the forecast about the number of queryes we aim to solve. If it will be huge in future than ball-tree algorthm was a good choice and a goo parameter included amongst the hyper-params grid of values, otherwise we should get rid of it.
# - referring to *leaf_size parameter*, we can notice that also here the choice of a specific kernel trick for performing kernel-Pca algorithm does not affect the value tha such a parameter has assumed amongst those proposed. However, recalling that leaf size hyper-param is used to monitor and control the tree-like structure of our solutions we can understand that since the value is pretty low the obtained trees were allowed to grow toward maximum depth.
# - speaking about *distance parameter*, the best solution through different trials was *Euclidean distance*, which also corresponds to the default choice, furthermore the choice of a kernel trick in the context of the other grid values was not affecting the choice of *distance parameter*.
# - *n_neighbors parameter* is the one which is most affected and influenced by the choice of kernel trick for performing the kernel-Pca preprocessing method, since three out of five trials found 3 as the best value which are *Linear, Poly and Cosine tricks*, however only the first two examples using such a low number of neighbors still obtained fine results, instead the best trial which is the classifier characterized from Rbf kernel trick for kernel-Pca has selected 7 as the best value for the number of neighbors meanning that such a classifier required a greater number of neighbor before estimating class label and also that the query time t solve the inference would be longer.
# - lastly, the *weights param* is involved when we want to assign a certain weight to examples used during classification, where usually farway points will have less effect and nearby point grow their importance. The most frequent choice was represented by the *distance strategy*, which assign a weigh value to each sample of train set involved during classification a value proportional to the inverse of the distance of that sample from the query point. Only the Sigmoid kernel trick case instead adopted a weights strategy which corresponds to the default choice which is the uniform strategy.
#
# If we imagine to build up an *Ensemble Classifier* from the family of *Average Methods*, which state that the underlying principle leading their creation requires to build separate and single classifiers than averaging their prediction in regression context or adopting a majority vote strategy for the classification context, we can claim that amongst the purposed Knn classifier, for sure, we could employ the classifier foudn from the first three trials because of their performance metrics and also because Ensemble Methods such as Bagging Classifier, usually work fine exploiting an ensemble of independent and fine tuned classifier differently from Boosting Methods which instead are based on weak learners.
# +
# show_histogram_first_sample(Xtrain_transformed, ytrain_, estimators_)
# -
# ### Improvements and Conclusions <a class="anchor" id="Improvements-and-conclusions"></a>
#
# Extension that we can think of to better improve the analyses we can perform on such a relative tiny dataset many include, for preprocessing phases:
# - Selecting different *Feature Extraction ant Dimensionality Reduction Techniques* other than Pca or kernel Pca such as:
# *linear discriminant analysis (LDA)*, or *canonical correlation analysis (CCA) techniques* as a pre-processing step.
#
# Extension that we can think of to better improve the analyses we can perform on such a relative tiny dataset many include, for training phases:
#
# - Selecting different *Ensemble Methods, investigating both Average based and Boosting based Statistical Learning Methods*.
#
# Extension that we can think of to better improve the analyses we can perform on such a relative tiny dataset many include, for diagnostic analyses after having performed train and test phases:
#
# - Using other measures, indicators and ghraphical plots such as the *Total Operating Characteristic (TOC)*, since also such a measure characterizes diagnostic ability while revealing more information than the ROC. In fact for each threshold, ROC reveals two ratios, TP/(TP + FN) and FP/(FP + TN). In other words, ROC reveals hits/(hits + misses) and false alarms/(false alarms + correct rejections). On the other hand, TOC shows the total information in the contingency table for each threshold. Lastly, the TOC method reveals all of the information that the ROC method provides, plus additional important information that ROC does not reveal, i.e. the size of every entry in the contingency table for each threshold.
# ## References section <a class="anchor" id="references"></a>
# ### Main References
# - Data Domain Information part:
# - (Deck) https://en.wikipedia.org/wiki/Deck_(bridge)
# - (Cantilever bridge) https://en.wikipedia.org/wiki/Cantilever_bridge
# - (Arch bridge) https://en.wikipedia.org/wiki/Deck_(bridge)
# - Machine Learning part:
# - (Theory Book) https://jakevdp.github.io/PythonDataScienceHandbook/
# - (Feature Extraction: PCA) https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.PCA.html
# - (Linear Model: Logistic Regression) https://scikit-learn.org/stable/modules/linear_model.html#logistic-regression
# - (Neighbor-based Learning: Knn) https://scikit-learn.org/stable/modules/neighbors.html
# - (Stochastc Learning: SGD Classifier) https://scikit-learn.org/stable/modules/sgd.html#sgd
# - (Discriminative Model: SVM) https://scikit-learn.org/stable/modules/svm.html
# - (Non-Parametric Learning: Decsion Trees) https://scikit-learn.org/stable/modules/tree.html#tree
# - (Ensemble, Non-Parametric Learning: RandomForest) https://scikit-learn.org/stable/modules/ensemble.html#forest
# - Metrics:
# - (F1-Accuracy-Precision-Recall) https://towardsdatascience.com/beyond-accuracy-precision-and-recall-3da06bea9f6c
# - Statistics:
# - (Correlation and dependence) https://en.wikipedia.org/wiki/Correlation_and_dependence
# - (KDE) https://jakevdp.github.io/blog/2013/12/01/kernel-density-estimation/
# - Chart part:
# - (Seaborn Charts) https://acadgild.com/blog/data-visualization-using-matplotlib-and-seaborn
# - Third Party Library:
# - (sklearn) https://scikit-learn.org/stable/index.html
# - (statsmodels) https://www.statsmodels.org/stable/index.html#
#
# ### Others References
# - Plots:
# - (Python Plot) https://www.datacamp.com/community/tutorials/matplotlib-tutorial-python?utm_source=adwords_ppc&utm_campaignid=898687156&utm_adgroupid=48947256715&utm_device=c&utm_keyword=&utm_matchtype=b&utm_network=g&utm_adpostion=&utm_creative=255798340456&utm_targetid=aud-299261629574:dsa-473406587955&utm_loc_interest_ms=&utm_loc_physical_ms=1008025&gclid=Cj0KCQjw-_j1BRDkARIsAJcfmTFu4LAUDhRGK2D027PHiqIPSlxK3ud87Ek_lwOu8rt8A8YLrjFiHqsaAoLDEALw_wcB
# - Markdown Math part:
# - (Math Symbols Latex) https://oeis.org/wiki/List_of_LaTeX_mathematical_symbols
# - (Tutorial 1) https://share.cocalc.com/share/b4a30ed038ee41d868dad094193ac462ccd228e2/Homework%20/HW%201.2%20-%20Markdown%20and%20LaTeX%20Cheatsheet.ipynb?viewer=share
# - (Tutorial 2) https://jupyter-notebook.readthedocs.io/en/stable/examples/Notebook/Typesetting%20Equations.html
| pittsburgh-bridges-data-set-analysis/models-analyses/grid_search_analyses/.ipynb_checkpoints/Data Space Report (Official) - Knn Analysis.v1.0.1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/yibiaojason/A-Fast-implemetation-of-Confluence/blob/main/hw2_classification.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="OSL2CMEzmQvB"
# #**Homework 2 - Classification**
# 若有任何問題,歡迎來信至助教信箱 <EMAIL>
# + [markdown] id="ox7joE3aZkh-"
# Binary classification is one of the most fundamental problem in machine learning. In this tutorial, you are going to build linear binary classifiers to predict whether the income of an indivisual exceeds 50,000 or not. We presented a discriminative and a generative approaches, the logistic regression(LR) and the linear discriminant anaysis(LDA). You are encouraged to compare the differences between the two, or explore more methodologies. Although you can finish this tutorial by simpliy copying and pasting the codes, we strongly recommend you to understand the mathematical formulation first to get more insight into the two algorithms. Please find [here](http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2016/Lecture/Logistic%20Regression%20(v3).pdf) and [here](http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2016/Lecture/Classification%20(v3).pdf) for more detailed information about the two algorithms.
#
# 二元分類是機器學習中最基礎的問題之一,在這份教學中,你將學會如何實作一個線性二元分類器,來根據人們的個人資料,判斷其年收入是否高於 50,000 美元。我們將以兩種方法: logistic regression 與 generative model,來達成以上目的,你可以嘗試了解、分析兩者的設計理念及差別。針對這兩個演算法的理論基礎,可以參考李宏毅老師的教學投影片 [logistic regression](http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2016/Lecture/Logistic%20Regression%20(v3).pdf) 與 [generative model](http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2016/Lecture/Classification%20(v3).pdf)。
#
# 若有任何問題,歡迎來信至助教信箱 <EMAIL>
# + [markdown] id="nkNW5cQmohoo"
# #Dataset
#
# This dataset is obtained by removing unnecessary attributes and balancing the ratio between positively and negatively labeled data in the [**Census-Income (KDD) Data Set**](https://archive.ics.uci.edu/ml/datasets/Census-Income+(KDD)), which can be found in [**UCI Machine Learning Repository**](https://archive.ics.uci.edu/ml/index.php). Only preprocessed and one-hot encoded data (i.e. *X_train*, *Y_train* and *X_test*) will be used in this tutorial. Raw data (i.e. *train.csv* and *test.csv*) are provided to you in case you are interested in it.
#
# 這個資料集是由 [**UCI Machine Learning Repository**](https://archive.ics.uci.edu/ml/index.php) 的 [**Census-Income (KDD) Data Set**](https://archive.ics.uci.edu/ml/datasets/Census-Income+(KDD)) 經過一些處理而得來。為了方便訓練,我們移除了一些不必要的資訊,並且稍微平衡了正負兩種標記的比例。事實上在訓練過程中,只有 X_train、Y_train 和 X_test 這三個經過處理的檔案會被使用到,train.csv 和 test.csv 這兩個原始資料檔則可以提供你一些額外的資訊。
# + id="Ww4-VJoJqE-_" colab={"base_uri": "https://localhost:8080/", "height": 238} outputId="398da30b-8e74-4e94-fe37-6e9f2c6af2b2"
# !gdown --id '1KSFIRh0-_Vr7SdiSCZP1ItV7bXPxMD92' --output data.tar.gz
# !tar -zxvf data.tar.gz
# !ls
# + [markdown] id="WRXI0kf0W4Bd"
# #Logistic Regression
#
# In this section we will introduce logistic regression first. We only present how to implement it here, while mathematical formulation and analysis will be omitted. You can find more theoretical detail in [Prof. Lee's lecture](https://www.youtube.com/watch?v=hSXFuypLukA).
#
# 首先我們會實作 logistic regression,針對理論細節說明請參考[李宏毅老師的教學影片](https://www.youtube.com/watch?v=hSXFuypLukA)
#
# ###Preparing Data
#
# Load and normalize data, and then split training data into training set and development set.
#
# 下載資料,並且對每個屬性做正規化,處理過後再將其切分為訓練集與發展集。
# + id="7NzAmkzU2MAS" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="61610be3-295e-4ff8-befe-8044938141c1"
import numpy as np
np.random.seed(0)
X_train_fpath = './data/X_train'
Y_train_fpath = './data/Y_train'
X_test_fpath = './data/X_test'
output_fpath = './output_{}.csv'
# Parse csv files to numpy array
with open(X_train_fpath) as f:
next(f)
X_train = np.array([line.strip('\n').split(',')[1:] for line in f], dtype = float)
with open(Y_train_fpath) as f:
next(f)
Y_train = np.array([line.strip('\n').split(',')[1] for line in f], dtype = float)
with open(X_test_fpath) as f:
next(f)
X_test = np.array([line.strip('\n').split(',')[1:] for line in f], dtype = float)
def _normalize(X, train = True, specified_column = None, X_mean = None, X_std = None):
# This function normalizes specific columns of X.
# The mean and standard variance of training data will be reused when processing testing data.
#
# Arguments:
# X: data to be processed
# train: 'True' when processing training data, 'False' for testing data
# specific_column: indexes of the columns that will be normalized. If 'None', all columns
# will be normalized.
# X_mean: mean value of training data, used when train = 'False'
# X_std: standard deviation of training data, used when train = 'False'
# Outputs:
# X: normalized data
# X_mean: computed mean value of training data
# X_std: computed standard deviation of training data
if specified_column == None:
specified_column = np.arange(X.shape[1])
if train:
X_mean = np.mean(X[:, specified_column] ,0).reshape(1, -1)
X_std = np.std(X[:, specified_column], 0).reshape(1, -1)
X[:,specified_column] = (X[:, specified_column] - X_mean) / (X_std + 1e-8)
return X, X_mean, X_std
def _train_dev_split(X, Y, dev_ratio = 0.25):
# This function spilts data into training set and development set.
train_size = int(len(X) * (1 - dev_ratio))
return X[:train_size], Y[:train_size], X[train_size:], Y[train_size:]
# Normalize training and testing data
X_train, X_mean, X_std = _normalize(X_train, train = True)
X_test, _, _= _normalize(X_test, train = False, specified_column = None, X_mean = X_mean, X_std = X_std)
# Split data into training set and development set
dev_ratio = 0.1
X_train, Y_train, X_dev, Y_dev = _train_dev_split(X_train, Y_train, dev_ratio = dev_ratio)
train_size = X_train.shape[0]
dev_size = X_dev.shape[0]
test_size = X_test.shape[0]
data_dim = X_train.shape[1]
print('Size of training set: {}'.format(train_size))
print('Size of development set: {}'.format(dev_size))
print('Size of testing set: {}'.format(test_size))
print('Dimension of data: {}'.format(data_dim))
# + [markdown] id="imgCeBDoApdb"
#
# ###Some Useful Functions
#
# Some functions that will be repeatedly used when iteratively updating the parameters.
#
# 這幾個函數可能會在訓練迴圈中被重複使用到。
# + id="hSDAw5LTAs2o"
def _shuffle(X, Y):
# This function shuffles two equal-length list/array, X and Y, together.
randomize = np.arange(len(X))
np.random.shuffle(randomize)
return (X[randomize], Y[randomize])
def _sigmoid(z):
# Sigmoid function can be used to calculate probability.
# To avoid overflow, minimum/maximum output value is set.
return np.clip(1 / (1.0 + np.exp(-z)), 1e-8, 1 - (1e-8))
def _f(X, w, b):
# This is the logistic regression function, parameterized by w and b
#
# Arguements:
# X: input data, shape = [batch_size, data_dimension]
# w: weight vector, shape = [data_dimension, ]
# b: bias, scalar
# Output:
# predicted probability of each row of X being positively labeled, shape = [batch_size, ]
return _sigmoid(np.matmul(X, w) + b)
def _predict(X, w, b):
# This function returns a truth value prediction for each row of X
# by rounding the result of logistic regression function.
return np.round(_f(X, w, b)).astype(np.int)
def _accuracy(Y_pred, Y_label):
# This function calculates prediction accuracy
acc = 1 - np.mean(np.abs(Y_pred - Y_label))
return acc
# + [markdown] id="OxJdfhEEOYwg"
# ### Functions about gradient and loss
#
# Please refers to [Prof. Lee's lecture slides](http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2016/Lecture/Logistic%20Regression%20(v3).pdf)(p.12) for the formula of gradient and loss computation.
#
# 請參考[李宏毅老師上課投影片](http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2016/Lecture/Logistic%20Regression%20(v3).pdf)第 12 頁的梯度及損失函數計算公式。
# + id="DqYkUgLjOWi1"
def _cross_entropy_loss(y_pred, Y_label):
# This function computes the cross entropy.
#
# Arguements:
# y_pred: probabilistic predictions, float vector
# Y_label: ground truth labels, bool vector
# Output:
# cross entropy, scalar
cross_entropy = -np.dot(Y_label, np.log(y_pred)) - np.dot((1 - Y_label), np.log(1 - y_pred))
return cross_entropy
def _gradient(X, Y_label, w, b):
# This function computes the gradient of cross entropy loss with respect to weight w and bias b.
y_pred = _f(X, w, b)
pred_error = Y_label - y_pred
w_grad = -np.sum(pred_error * X.T, 1)
b_grad = -np.sum(pred_error)
return w_grad, b_grad
# + [markdown] id="XXEFuqydaA34"
# ### Training
#
# Everything is prepared, let's start training!
#
# Mini-batch gradient descent is used here, in which training data are split into several mini-batches and each batch is fed into the model sequentially for losses and gradients computation. Weights and bias are updated on a mini-batch basis.
#
# Once we have gone through the whole training set, the data have to be re-shuffled and mini-batch gradient desent has to be run on it again. We repeat such process until max number of iterations is reached.
#
# 我們使用小批次梯度下降法來訓練。訓練資料被分為許多小批次,針對每一個小批次,我們分別計算其梯度以及損失,並根據該批次來更新模型的參數。當一次迴圈完成,也就是整個訓練集的所有小批次都被使用過一次以後,我們將所有訓練資料打散並且重新分成新的小批次,進行下一個迴圈,直到事先設定的迴圈數量達成為止。
# + id="s6yNUeG9aBR1" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="ab2d6731-1d43-4298-b278-5bc4f907b096"
# Zero initialization for weights ans bias
w = np.zeros((data_dim,))
b = np.zeros((1,))
# Some parameters for training
max_iter = 10
batch_size = 8
learning_rate = 0.2
# Keep the loss and accuracy at every iteration for plotting
train_loss = []
dev_loss = []
train_acc = []
dev_acc = []
# Calcuate the number of parameter updates
step = 1
# Iterative training
for epoch in range(max_iter):
# Random shuffle at the begging of each epoch
X_train, Y_train = _shuffle(X_train, Y_train)
# Mini-batch training
for idx in range(int(np.floor(train_size / batch_size))):
X = X_train[idx*batch_size:(idx+1)*batch_size]
Y = Y_train[idx*batch_size:(idx+1)*batch_size]
# Compute the gradient
w_grad, b_grad = _gradient(X, Y, w, b)
# gradient descent update
# learning rate decay with time
w = w - learning_rate/np.sqrt(step) * w_grad
b = b - learning_rate/np.sqrt(step) * b_grad
step = step + 1
# Compute loss and accuracy of training set and development set
y_train_pred = _f(X_train, w, b)
Y_train_pred = np.round(y_train_pred)
train_acc.append(_accuracy(Y_train_pred, Y_train))
train_loss.append(_cross_entropy_loss(y_train_pred, Y_train) / train_size)
y_dev_pred = _f(X_dev, w, b)
Y_dev_pred = np.round(y_dev_pred)
dev_acc.append(_accuracy(Y_dev_pred, Y_dev))
dev_loss.append(_cross_entropy_loss(y_dev_pred, Y_dev) / dev_size)
print('Training loss: {}'.format(train_loss[-1]))
print('Development loss: {}'.format(dev_loss[-1]))
print('Training accuracy: {}'.format(train_acc[-1]))
print('Development accuracy: {}'.format(dev_acc[-1]))
# + [markdown] id="RJuoQ_R2jUmX"
# ### Plotting Loss and accuracy curve
# + id="DH3AJtvHjVJ7" colab={"base_uri": "https://localhost:8080/", "height": 545} outputId="f3fc5d1b-ddcc-4cf6-eea5-ebf23026edc5"
import matplotlib.pyplot as plt
# Loss curve
plt.plot(train_loss)
plt.plot(dev_loss)
plt.title('Loss')
plt.legend(['train', 'dev'])
plt.savefig('loss.png')
plt.show()
# Accuracy curve
plt.plot(train_acc)
plt.plot(dev_acc)
plt.title('Accuracy')
plt.legend(['train', 'dev'])
plt.savefig('acc.png')
plt.show()
# + [markdown] id="HzIcYAfvkUZ_"
# ###Predicting testing labels
#
# Predictions are saved to *output_logistic.csv*.
#
# 預測測試集的資料標籤並且存在 *output_logistic.csv* 中。
# + id="ZEAKhugPkUyH" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="97c3eb12-a9c5-4c43-bc62-54f7f5f4797d"
# Predict testing labels
predictions = _predict(X_test, w, b)
with open(output_fpath.format('logistic'), 'w') as f:
f.write('id,label\n')
for i, label in enumerate(predictions):
f.write('{},{}\n'.format(i, label))
# Print out the most significant weights
ind = np.argsort(np.abs(w))[::-1]
with open(X_test_fpath) as f:
content = f.readline().strip('\n').split(',')
features = np.array(content)
for i in ind[0:10]:
print(features[i], w[i])
# + [markdown] id="1C6sqhUbLMGe"
# # Porbabilistic generative model
#
# In this section we will discuss a generative approach to binary classification. Again, we will not go through the formulation detailedly. Please find [Prof. Lee's lecture](https://www.youtube.com/watch?v=fZAZUYEeIMg) if you are interested in it.
#
# 接者我們將實作基於 generative model 的二元分類器,理論細節請參考[李宏毅老師的教學影片](https://www.youtube.com/watch?v=fZAZUYEeIMg)。
#
# ### Preparing Data
#
# Training and testing data is loaded and normalized as in logistic regression. However, since LDA is a deterministic algorithm, there is no need to build a development set.
#
# 訓練集與測試集的處理方法跟 logistic regression 一模一樣,然而因為 generative model 有可解析的最佳解,因此不必使用到 development set。
# + id="czWXO7qML8DU"
# Parse csv files to numpy array
with open(X_train_fpath) as f:
next(f)
X_train = np.array([line.strip('\n').split(',')[1:] for line in f], dtype = float)
with open(Y_train_fpath) as f:
next(f)
Y_train = np.array([line.strip('\n').split(',')[1] for line in f], dtype = float)
with open(X_test_fpath) as f:
next(f)
X_test = np.array([line.strip('\n').split(',')[1:] for line in f], dtype = float)
# Normalize training and testing data
X_train, X_mean, X_std = _normalize(X_train, train = True)
X_test, _, _= _normalize(X_test, train = False, specified_column = None, X_mean = X_mean, X_std = X_std)
# + [markdown] id="L8NGKl-EPvok"
# ### Mean and Covariance
#
# In generative model, in-class mean and covariance are needed.
#
# 在 generative model 中,我們需要分別計算兩個類別內的資料平均與共變異。
# + id="iQrzXXKUPwHT"
# Compute in-class mean
X_train_0 = np.array([x for x, y in zip(X_train, Y_train) if y == 0])
X_train_1 = np.array([x for x, y in zip(X_train, Y_train) if y == 1])
mean_0 = np.mean(X_train_0, axis = 0)
mean_1 = np.mean(X_train_1, axis = 0)
# Compute in-class covariance
cov_0 = np.zeros((data_dim, data_dim))
cov_1 = np.zeros((data_dim, data_dim))
for x in X_train_0:
cov_0 += np.dot(np.transpose([x - mean_0]), [x - mean_0]) / X_train_0.shape[0]
for x in X_train_1:
cov_1 += np.dot(np.transpose([x - mean_1]), [x - mean_1]) / X_train_1.shape[0]
# Shared covariance is taken as a weighted average of individual in-class covariance.
cov = (cov_0 * X_train_0.shape[0] + cov_1 * X_train_1.shape[0]) / (X_train_0.shape[0] + X_train_1.shape[0])
# + [markdown] id="kifW1pFxXXA5"
# ### Computing weights and bias
#
# Directly compute weights and bias from in-class mean and shared variance. [Prof. Lee's lecture slides](http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2016/Lecture/Classification%20(v3).pdf)(p.33) gives a concise explanation.
#
# 權重矩陣與偏差向量可以直接被計算出來,算法可以參考[李宏毅老師教學投影片](http://speech.ee.ntu.edu.tw/~tlkagk/courses/ML_2016/Lecture/Classification%20(v3).pdf)第 33 頁。
# + id="UghOxYrUXXPU" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c71a5d93-4c19-48e5-b0ea-38c3e1e132ca"
# Compute inverse of covariance matrix.
# Since covariance matrix may be nearly singular, np.linalg.inv() may give a large numerical error.
# Via SVD decomposition, one can get matrix inverse efficiently and accurately.
u, s, v = np.linalg.svd(cov, full_matrices=False)
inv = np.matmul(v.T * 1 / s, u.T)
# Directly compute weights and bias
w = np.dot(inv, mean_0 - mean_1)
b = (-0.5) * np.dot(mean_0, np.dot(inv, mean_0)) + 0.5 * np.dot(mean_1, np.dot(inv, mean_1))\
+ np.log(float(X_train_0.shape[0]) / X_train_1.shape[0])
# Compute accuracy on training set
Y_train_pred = 1 - _predict(X_train, w, b)
print('Training accuracy: {}'.format(_accuracy(Y_train_pred, Y_train)))
# + [markdown] id="RDKWzBy0bi3c"
# ###Predicting testing labels
#
# Predictions are saved to *output_generative.csv*.
#
# 預測測試集的資料標籤並且存在 *output_generative.csv* 中。
# + id="T3QjToT_Sq9J" colab={"base_uri": "https://localhost:8080/", "height": 187} outputId="955f3181-82a9-4701-d57f-5e524cacd34e"
# Predict testing labels
predictions = 1 - _predict(X_test, w, b)
with open(output_fpath.format('generative'), 'w') as f:
f.write('id,label\n')
for i, label in enumerate(predictions):
f.write('{},{}\n'.format(i, label))
# Print out the most significant weights
ind = np.argsort(np.abs(w))[::-1]
with open(X_test_fpath) as f:
content = f.readline().strip('\n').split(',')
features = np.array(content)
for i in ind[0:10]:
print(features[i], w[i])
| hw2_classification.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# # Neural ordinary differential equation model
# <NAME> (@sdwfrost) 2022-03-31
#
# A neural ODE is an ODE where a neural network defines its derivative function. In this simple example, we train a neural ODE on a standard SIR model described by an ODE, and generate a forecast of the dynamics.
#
# ## Libraries
using OrdinaryDiffEq
using DiffEqFlux, Flux
using Random
using Plots;
Random.seed!(123);
# ## Transitions of ODE system
#
# To assist in numerical stability, we consider the proportion of individuals in the population (`s,i,r`) rather than the number of individuals (`S,I,R`). As the neural ODEs are defined out-of-place (i.e. they return a new derivative `du`), we use this syntax for our 'true' ODE model.
function sir_ode(u,p,t)
(s,i,r) = u
(β,γ) = p
ds = -β*s*i
di = β*s*i - γ*i
dr = γ*i
[ds,di,dr]
end;
# ## Parameters, initial conditions, etc.
#
# Note that our parameter values and initial conditions reflect that in this example, we assume we are modeling proportions (i.e. `s+i+r=1`).
p = [0.5,0.25]
u0 = [0.99, 0.01, 0.0]
tspan = (0.0, 40.0)
δt = 1;
# ## Solving the true model
#
# To derive trajectories for training, we first solve the true model.
solver = Rodas5();
sir_prob = ODEProblem(sir_ode, u0, tspan, p)
sir_sol = solve(sir_prob, solver, saveat = δt);
# This simple example assumes we have accurate values for all state variables, which we obtain from the solution of the ODE over the training time period.
#
# For training, we use a short timespan of `(0,30)`, and will forecast for an additional 10 time units, with training using all three state variables every `δt` time units.
train_time = 30.0
train_data = Array(sir_sol(0:δt:train_time));
# ## Defining a neural ODE
#
# To define a neural ODE, we need to decide on an architecture. Here is a simple multilayer perceptron that takes three inputs (the three state variables `s,i,r`) and generates three outputs (the derivatives, `ds,di,dr`).
nhidden = 8
sir_node = FastChain(FastDense(3, nhidden, tanh),
FastDense(nhidden, nhidden, tanh),
FastDense(nhidden, nhidden, tanh),
FastDense(nhidden, 3));
# As we are using a Flux.jl `FastChain`, we could write our neural ODE as follows (see [this page](https://diffeqflux.sciml.ai/dev/examples/neural_ode_sciml/) under 'Usage Without the Layer Function').
p_ = Float64.(initial_params(sir_node));
function dudt_sir_node(u,p,t)
s,i,r = u
ds,di,dr = ann_node([s,i,r],p)
[ds,di,dr]
end
prob_node = ODEProblem(dudt_sir_node, u0, tspan, p_);
# However, `DiffEqFlux.jl` offers a simpler interface where we can just pass a neural network, without generating the initial parameters and writing the gradient function.
tspan_train = (0,train_time)
prob_node = NeuralODE(sir_node,
tspan_train,
solver,
saveat=δt,
sensealg = InterpolatingAdjoint(autojacvec=ReverseDiffVJP()))
nump = length(prob_node.p)
# The following uses the sum of squared differences between the neural ODE predictions and the true state variables (`train_data`, above) as a loss function. As described [here](https://diffeqflux.sciml.ai/dev/examples/divergence/), it is important to be able to handle failed integrations.
function loss(p)
sol = prob_node(u0,p)
pred = Array(sol)
sum(abs2, (train_data .- pred)), pred
end;
# The following is a simple callback function that displays the current value of the loss every 50 steps. We'll keep an array of losses to plot later.
const losses = []
callback = function (p, l, pred)
push!(losses, l)
numloss = length(losses)
if numloss % 50 == 0
display("Epoch: " * string(numloss) * " Loss: " * string(l))
end
return false
end;
# ### Training
res_node = DiffEqFlux.sciml_train(loss,
prob_node.p,
cb = callback);
plot(losses, yaxis = :log, xaxis = :log, xlabel = "Iterations", ylabel = "Loss", legend = false)
# ## Plotting
#
# We generate a new problem with the parameters from the above fit.
prob_node = NeuralODE(sir_node,
tspan_train,
solver,
saveat=δt,
sensealg = InterpolatingAdjoint(autojacvec=ReverseDiffVJP()),
p = res_node)
sol_node = prob_node(u0);
# A plot of the dynamics shows a good fit.
scatter(sir_sol, label=["True Susceptible" "True Infected" "True Recovered"])
plot!(sol_node, label=["Estimated Susceptible" "Estimated Infected" "Estimated Recovered"])
# ## Forecasting
#
# We can also run the fitted model forward in time in order to assess its ability to forecast.
tspan_test = (0.0, 40.0)
prob_node_test = NeuralODE(sir_node,
tspan_test,
solver,
saveat=δt,
sensealg = InterpolatingAdjoint(autojacvec=ReverseDiffVJP()),
p = res_node)
sol_node_test = prob_node_test(u0);
p_node = scatter(sol_node_test, legend = :topright, label=["True Susceptible" "True Infected" "True Recovered"], title="Neural ODE Extrapolation: training until t=30")
plot!(p_node,sol_node_test, lw=5, label=["Estimated Susceptible" "Estimated Infected" "Estimated Recovered"])
# ## Performance for different initial conditions
#
# Here, we evaluate the fit for a different initial condition than that used for training.
newu0 = [0.95, 0.05, 0.0]
sir_prob_u0 = remake(sir_prob,u0=newu0)
sir_sol_u0 = solve(sir_prob_u0, solver, saveat = δt)
node_sol_u0 = prob_node(newu0)
p_node = scatter(sir_sol_u0, legend = :topright, label=["True Susceptible" "True Infected" "True Recovered"], title="Neural ODE with different initial conditions")
plot!(p_node,node_sol_u0, lw=5, label=["Estimated Susceptible" "Estimated Infected" "Estimated Recovered"])
# Even though the neural network is only training the derivatives, there is some dependency on the initial conditions used to generate the training data.
#
# ## Performance with a shorter training dataset
#
# Here is an example of training with more limited data - training only on times `t=0:20`. The values of `abstol` and `reltol` are reduced in order to avoid numerical problems.
tspan_train2 = (0.0,20.0)
prob2 = ODEProblem(sir_ode, u0, tspan_train2, p)
sol2 = solve(prob2, solver, saveat = δt)
data2 = Array(sol2)
solver2 = ExplicitRK()
prob_node2 = NeuralODE(sir_node,
tspan_train2,
solver2,
saveat=δt,
sensealg = InterpolatingAdjoint(autojacvec=ReverseDiffVJP()))
function loss2(p)
sol = prob_node2(u0,p)
pred = Array(sol)
sum(abs2, (data2 .- sol)), pred
end
const losses2 = []
callback2 = function (p, l, pred)
push!(losses2, l)
numloss = length(losses2)
if numloss % 50 == 0
display("Epoch: " * string(numloss) * " Loss: " * string(l))
end
return false
end
res_node2 = DiffEqFlux.sciml_train(loss2,
prob_node2.p,
cb = callback2);
# We now solve the new model over the full testing time period.
prob_node2_test = NeuralODE(sir_node,
tspan_test,
solver,
saveat=δt,
sensealg = InterpolatingAdjoint(autojacvec=ReverseDiffVJP()),
p = res_node2)
sol_node2_test = prob_node2_test(u0);
# A plot of the forecast shows that the model still predicts well in the very short term, but fails in the longer term due to more limited data, and shows unrealistic increases in susceptibles. With even more limited data (training up to `t=15.0`), the extrapolation becomes worse, and even shows negative population sizes.
p_node2 = scatter(sir_sol, legend = :topright, label=["True Susceptible" "True Infected" "True Recovered"], title="Neural ODE Extrapolation: training until t=20")
plot!(p_node2, sol_node2_test, lw=5, label=["Estimated Susceptible" "Estimated Infected" "Estimated Recovered"])
# ## Discussion
#
# Neural ODEs provide a way to fit and extrapolate in a data-driven way, and they perform well, at least in terms of fit and short-term forecasts, for this simple example. With more limited data, the goodness-of-fit to the training data may be misleading. Numerical issues can also arise in the fitting of neural ODEs; potential solutions include changing the solver and decreasing the tolerances. In addition, we rarely have access to all the state variables of a system. Many of these deficiencies can be addressed through combining neural network approaches with domain-specific knowledge e.g. using [universal differential equations](https://arxiv.org/abs/2001.04385).
| notebook/node/node.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# ### Import Packages
library(readxl)
library(stats)
library(data.table)
library(MASS)
library(moments)
library(car)
library(dplyr)
library(nortest)
# ### Ingest Data
data <- read.csv('./sf_airbnb_clean_subset.csv')
for_ratings <- read.csv('./sf_airbnb_clean.csv')
colnames(for_ratings)
# ### Subsetting Data
for_ratings <- for_ratings %>% select(-id, -host_id, -number_of_reviews, -number_of_reviews_ltm, -review_scores_accuracy,
-review_scores_cleanliness, -review_scores_checkin, -review_scores_communication,
-review_scores_location, -review_scores_value)
colnames(data)
subset_data <- data %>% select(-id,
-host_id, -X)
for_ratings <- na.omit(for_ratings)
head(subset_data)
colnames(subset_data)
subset_data <- na.omit(subset_data)
# ### Model 1
model_1 <- lm(log(price + 1) ~ ., data = subset_data)
summary(model_1)
stepAIC(model_1, direction = 'both', trace = FALSE)
model_2 <- lm(formula = log(price + 1) ~ host_is_superhost + host_has_profile_pic +
neighbourhood_cleansed + property_type +
room_type + accommodates + bathrooms + bedrooms + beds +
security_deposit + cleaning_fee + guests_included + extra_people +
maximum_nights + minimum_minimum_nights + maximum_minimum_nights +
minimum_maximum_nights + maximum_nights_avg_ntm +
availability_30 + availability_365 + instant_bookable + require_guest_phone_verification +
calculated_host_listings_count
+ reviews_per_month +
with_wifi + with_air_conditioning + with_cable_tv + pets_allowed,
data = subset_data)
summary(model_2)
capture.output(summary(model_2), file = "myfile4.txt")
residuals <- resid(model_2)
qqnorm(residuals)
qqline(residuals)
ad.test(residuals)
hist(log(subset_data$price))
model_3 <- lm(formula = log(price + 1) ~ host_listings_count + neighbourhood_cleansed +
latitude + property_type + room_type + accommodates + bathrooms +
bedrooms + beds + security_deposit + cleaning_fee + guests_included +
extra_people + minimum_minimum_nights + minimum_maximum_nights +
maximum_nights_avg_ntm + availability_30 + number_of_reviews +
number_of_reviews_ltm + review_scores_rating + review_scores_cleanliness +
review_scores_value + require_guest_profile_picture + calculated_host_listings_count +
calculated_host_listings_count_entire_homes + calculated_host_listings_count_private_rooms +
reviews_per_month + with_air_conditioning + pets_allowed,
data = subset_data)
summary(model_3)
residuals <- resid(model_3)
ad.test(residuals)
qqnorm(residuals)
colnames(for_ratings)
ratings_model <- lm(review_scores_rating ~ ., data = for_ratings)
summary(ratings_model)
stepAIC(ratings_model, direction = 'both', trace = FALSE)
ratings_model_2 <- lm(formula = review_scores_rating ~ host_response_time + host_is_superhost +
host_listings_count + host_has_profile_pic + neighbourhood_cleansed +
property_type + room_type + accommodates + beds + price +
guests_included + extra_people + minimum_nights + maximum_nights +
minimum_minimum_nights + maximum_minimum_nights + maximum_maximum_nights +
minimum_nights_avg_ntm + calendar_updated + availability_30 +
availability_365 + cancellation_policy + calculated_host_listings_count +
calculated_host_listings_count_entire_homes + calculated_host_listings_count_private_rooms +
calculated_host_listings_count_shared_rooms + with_wifi +
with_air_conditioning + with_cable_tv + pets_allowed, data = for_ratings)
summary(ratings_model_2)
max(for_ratings$review_scores_rating)
hist(1 / (101 - for_ratings$review_scores_rating))
install.packages('randomForest')
library(randomForest)
random_forest <- randomForest(review_scores_rating ~ host_response_time + host_is_superhost +
host_listings_count + host_has_profile_pic + neighbourhood_cleansed +
property_type + room_type + accommodates + beds + price +
guests_included + extra_people + minimum_nights + maximum_nights +
minimum_minimum_nights + maximum_minimum_nights + maximum_maximum_nights +
minimum_nights_avg_ntm + availability_30 +
availability_365 + cancellation_policy + calculated_host_listings_count +
calculated_host_listings_count_entire_homes + calculated_host_listings_count_private_rooms +
calculated_host_listings_count_shared_rooms + with_wifi +
with_air_conditioning + with_cable_tv + pets_allowed, data = for_ratings)
random_forest
| EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Getting test data from Elasticsearch
#
# Here we get a fixed number of documents from Elasticsearch using a random seed and store them in JSON format.
# +
# %load_ext autoreload
# %autoreload 2
import sys
sys.path.append("../../..")
from heritageconnector.datastore import es, index
from elasticsearch import helpers
from tqdm.auto import tqdm
from itertools import islice
import json
# +
limit = None
# only consider docs with both a label and description field
query = {
"query": {
"function_score": {
"query": {
"bool": {
"must": [
{"exists": {"field": "graph.@rdfs:label"}},
{
"exists": {
"field": "data.http://www.w3.org/2001/XMLSchema#description"
}
},
]
}
},
"random_score": {"seed": 42, "field": "_seq_no"},
}
}
}
doc_generator = helpers.scan(
client=es,
index=index,
query=query,
preserve_order=True,
)
if limit:
doc_generator = islice(doc_generator, limit)
# +
# run export
output_path = f"./test_data_{limit}.jsonl"
with open(output_path, "w") as f:
for doc in tqdm(doc_generator):
json.dump(doc, f)
f.write("\n")
# -
| experiments/NEL/data/get_raw_data_from_es.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Goal:
#
# Outline the process of producing shared QC metric schema that delegates to picard names when they are adequately descriptive of what they measure. The following two workflows were used to extract metrics, and files were downloaded to `picard_metric_dir` and `optimus_metric_dir`:
# ```
# https://job-manager.mint-dev.broadinstitute.org/jobs/a39b92db-bed0-40d4-83de-3ca0505dc5a8 # 10x v2
# https://job-manager.mint-dev.broadinstitute.org/jobs/b9ff68b4-2434-4909-8275-850cb84ebb13 # ss2
# ```
import os
from crimson import picard
# ## Examine SS2 pipeline metrics outputs
#
# Listed below are the file names of metrics files emitted by a smart-seq2 workflow
picard_metric_dir = os.path.expanduser('~/Desktop/picard')
# !ls $picard_metric_dir
# This method parses a few of the files that are in a consistent format
# +
metric_files = [os.path.join(picard_metric_dir, f) for f in os.listdir(picard_metric_dir)]
def parse_picard(metric_file):
with open(metric_file, 'r') as f:
json_data = picard.parse(f)
metric_class_name = json_data['metrics']['class']
metrics = {}
for d in json_data['metrics']['contents']:
for k, v in d.items():
metrics[k] = type(v)
del metrics['SAMPLE_ALIAS'], metrics['LIBRARY']
return metric_class_name, metrics
# -
# This is a map between the metric class and the names of metrics calculated by each class, mapped to the output type.
#
# Caveat: 5 of the files don't decode. Those are printed in full below.
all_metrics_and_names = {}
for m in metric_files[:-2]:
try:
all_metrics_and_names.__setitem__(*parse_picard(m))
except:
print(m)
all_metrics_and_names
# Below, files that didn't convert are just printed to console to get a sense of their metric names
# !cat $picard_metric_dir/SRR1294925_qc.base_distribution_by_cycle_metrics.txt
# !cat $picard_metric_dir/SRR1294925_qc.gc_bias.summary_metrics.txt
# !cat $picard_metric_dir/SRR1294925_qc.gc_bias.detail_metrics.txt
# !cat $picard_metric_dir/SRR1294925_qc.error_summary_metrics.txt
# !cat $picard_metric_dir/SRR1294925_qc.quality_by_cycle_metrics.txt
# !cat $picard_metric_dir/SRR1294925_qc.alignment_summary_metrics.txt
# ## Optimus Metrics
#
# Now, do the same for Optimus metrics. Optimus has all of the metrics in one file, although may not have the depth of analysis that the picard ones do. We could use picard + user research to identify missing metrics and expand our complement as recommended.
# +
import pandas as pd
optimus_metric_dir = os.path.expanduser('~/Desktop/optimus')
print('cell metrics\n')
for c in pd.read_csv(os.path.join(optimus_metric_dir, 'merged-cell-metrics.csv.gz')).columns[1:]:
print(c)
print('\ngene metrics\n')
for c in pd.read_csv(os.path.join(optimus_metric_dir, 'merged-gene-metrics.csv.gz')).columns[1:]:
print(c)
# -
# ## Schema
#
# Picard does not appear to follow a consistent schema, although one can map their results into JSON using Crimson, which is helpful. The matrix service will require the data be shipped in the same storage format recommend for sequencing data. Currently this appears to be Zarr.
#
# ## ACs for ticket humancellatlas/secondary-analysis#105:
# 1. List of metrics to be updated:
# 1. All, both pipelines. This is a small epic that will span multiple tickets to implement in a forward-compatible fashion.
# 2. names to be changed:
# 2. Both SS2 and Optimus metrics will need to be mapped to a schema that we generate. Many of the picard names will work as is, however some will need to change by nature of their specificity to the files that generated them. For example, "REF_BASE" and "ALT_BASE" lose substantial meaning when combined into a columnar file.
# 3. Generally, the conclusion here is that a simple mapping from Optimus -> picard is neither possible nor practical, and that a larger effort should be undertaken to future-proof this aspect of secondary analysis.
# 4. conclusion on this effort: 1.5
#
#
# ## Further Implementation Suggestions:
# 1. Approach Crimson developer about willingness to receive PRs that expand his tool to encompass additional formats (as we receive them).
# 1. Is BSD format OK for HCA? Ask about licensing if no.
# 2. Expand it to read any other metrics.
# 3. Write some kind of json-metric-to-zarr converter. Keep the 'to-zarr' part separated, in case we need to change things around later.
# 4. In the process of doing this, determine our own internal schema and a series of maps from other tools' QC metrics.
# 5. Around our schema, design a glossary that describes each metric.
#
# ## Concrete Proposal: QC metrics Tool
# 1. Maps of tool-specific names to HCA schema: enables conversion of names to internal notation
# 2. Metric Converter library (Crimson?): enables conversion of metric data files to internal notation
# 3. Metric formatter library: dumps the json internal representation to disk using a swappable format backend
#
# It is possible that (3) could be shared by the software used for the matrix service. Most of the code is hopefully embedded in library-specific writers (zarr has a fairly good one).
| analyses/metrics/determine_qc_metric_schema.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// ## Introduction
// https://opendsa-server.cs.vt.edu/ODSA/Books/CS3/html/IntroDSA.html
// - Some problems to think about...
// 1. What is the fastest route from Los Angeles, CA to New York City, NY?
// - how long does it take to get there?
// 2. What is the most reliable source on the Internet to learn about data structures and algorithms?
// 3. Who was the most influential celebrity/politician of last year?
// 4. What is an average salary of a software engineer?
// 5. What is the cheapest way to travel from Grand Junction, CO, to Kathmandu, Nepal?
// 6. How does Apple's Siri know what appointment you have next when you ask it?
// + [markdown] heading_collapsed=true
// ## Computer Science (CS) fundamentals
// + [markdown] hidden=true
// - the core of CS is representing data so that it can be efficiently stored and retrieved
// - many computer programs sole functionality is to do just that, but as fast as possible. e.g., search engines like, Google, Bing, etc.
// - some programs may do heavy mathematical computation as fast as possible, e.g., wolframalpha computational intelligence (https://www.wolframalpha.com)
// - find factorial(10000) or 10000!
//
// **the study of data structures and the alogrithms that manipulate them to solve a given problem in feasible time and resource is the heart of compuer science**
// -
// ## Goals of Data Structure (DS) and Algorithm courses
// 1. present a collection of commonly used data structures and algorithms, programmer's basic "toolkit"
// - for many problems these toolkit out-of-the box will provide efficient solutions
// - "toolkit" forms the basis/foundations to build more advanced data structures
// 2. introduce the idea of trade-offs between the costs and benefits associated with every data structure or algorithm, e.g., trade-offs between memory and time
// 3. learn to measure the effectiveness of a data structure or algorithm so you can pick the right ones for the job
// - also allows you to quickly judge the merits of new data structures or algorithms that you or others might invent
// ## Solving problems
// - the primary job of computer scientists is to solve problems!
// - there are often too many approaches to solve a problem
// - at the heart of computer program designs are two (sometimes conflicting) goals:
// 1. to design an algorithm that is easy to understand, code, and debug
// 2. to design an algorithm that makes efficient use of the computer's resources
// - "elegant" solutions meet both these goals!
// - software engineering focuses on goal 1, though we emphasize it from CS1!
// - CS2 and CS3 usually focuses on goal 2.
// ## Why faster DS and algorithms when we've faster computers?
// - according to Moore's law https://en.wikipedia.org/wiki/Moore%27s_law, no. of transistors in computer's circuit board doubles every two years.
// - so, if processor speed and memory size continue to improve, won't today's hard problem be solved by tomorrow's hardware?
// - additional computing power enables us to tackle more complex problems, such as sophisticated user interfaces such as in mobile devices, bigger problem sizes (big data), or new problems previously deemed computationally infeasible
// - resources are always limited...
// - efficient solution solves the problem within the required *resource constraints*.
// - may require fewer resources than known alternatives, regardless of whether it meets any particular requirements
// - **cost** of a solution is the amount of resources that the solution consumes
// - measured typically in terms of one key resource such as time implying it meets all other resource constraints
// - e.g., fastest solutions on open.kattis.com problems meeting memory requirements: https://open.kattis.com/problems/cd/statistics
// ## Selecting a Data Structure
// 1. analyze your problem to determine the **basic operations** e.g., inserting data item into the data structure, deleting, finding a specified data item
// - quantify the resource constraints for each operation
// - select the data structure that best meets these requirements
//
// ### Some questions to think about to determine the importance of operations
// 1. is the application static or dynamic
// - in static applications, data is loaded at the beginning and new data are not inserted
// - in dynamic applications, new data items are inserted and may be inserted in any locations
// - can data items be deleted? this may make the implementation more complicated
// - how are the data items processed? in some well-defined order, random access?
// ## Exercises
// 1. Which of these is NOT a definition for efficiency in a computer program?
// - it solves the problem within the required resource constraints
// - it requires fewer resources than known alternatives
// - it runs in linear time
| Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Mutation Analysis
#
# In the [chapter on coverage](Coverage.ipynb), we showed how one can identify which parts of the program are executed by a program, and hence get a sense of the effectiveness of a set of test cases in covering the program structure. However, coverage alone may not be the best measure for the effectiveness of a test, as one can have great coverage without ever checking a result for correctness. In this chapter, we introduce another means for assessing the effectiveness of a test suite: After injecting *mutations* – _artificial faults_ – into the code, we check whether a test suite can detect these artificial faults. The idea is that if it fails to detect such mutations, it will also miss real bugs.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# **Prerequisites**
#
# * You need some understanding of how a program is executed.
# * You should have read [the chapter on coverage](Coverage.ipynb).
# + [markdown] slideshow={"slide_type": "skip"}
# ## Synopsis
# <!-- Automatically generated. Do not edit. -->
#
# To [use the code provided in this chapter](Importing.ipynb), write
#
# ```python
# >>> from fuzzingbook.MutationAnalysis import <identifier>
# ```
#
# and then make use of the following features.
#
#
# This chapter introduces two methods of running *mutation analysis* on subject programs. The first class `MuFunctionAnalyzer` targets individual functions. Given a function `gcd` and two test cases evaluate, one can run mutation analysis on the test cases as follows:
#
# ```python
# >>> for mutant in MuFunctionAnalyzer(gcd, log=True):
# >>> with mutant:
# >>> assert gcd(1, 0) == 1, "Minimal"
# >>> assert gcd(0, 1) == 1, "Mirror"
# >>> mutant.pm.score()
# ```
# The second class `MuProgramAnalyzer` targets standalone programs with test suites. Given a program `gcd` whose source code is provided in `gcd_src` and the test suite is provided by `TestGCD`, one can evaluate the mutation score of `TestGCD` as follows:
#
# ```python
# >>> class TestGCD(unittest.TestCase):
# >>> def test_simple(self):
# >>> assert cfg.gcd(1, 0) == 1
# >>>
# >>> def test_mirror(self):
# >>> assert cfg.gcd(0, 1) == 1
# >>> for mutant in MuProgramAnalyzer('gcd', gcd_src):
# >>> mutant[test_module].runTest('TestGCD')
# >>> mutant.pm.score()
# ```
# The mutation score thus obtained is a better indicator of the quality of a given test suite than pure coverage.
#
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Why Structural Coverage is Not Enough
#
# One of the problems with [structural coverage](Coverage.ipynb) measures is that it fails to check whether the program executions generated by the test suite were actually _correct_. That is, an execution that produces a wrong output that is unnoticed by the test suite is counted exactly the same as an execution that produces the right output for coverage. Indeed, if one deletes the assertions in a typical test case, the coverage would not change for the new test suite, but the new test suite is much less useful than the original one. As an example, consider this "test":
# -
def ineffective_test():
execute_the_program_as_a_whole()
assert True
# The final assertion here will always pass, no matter what `execute_the_program_as_a_whole()` will do. Okay, if `execute_the_program_as_a_whole()` raises an exception, the test will fail, but we can also get around that:
def ineffective_test():
try:
execute_the_program_as_a_whole()
except:
pass
assert True
# The problem with these "tests", however, is that `execute_the_program_as_a_whole()` may achieve 100% code coverage (or 100% of any other structural coverage metric). Yet, this number of 100% does not reflect the ability of the test to discover bugs, which actually is 0%.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# This is indeed, not an optimal state of affairs. How can we verify that our tests are actually useful? One alternative (hinted in the chapter on coverage) is to inject bugs into the program, and evaluate the effectiveness of test suites in catching these injected bugs. However, that introduces another problem. How do we produce these bugs in the first place? Any manual effort is likely to be biased by the preconceptions of the developer as to where the bugs are likely to occur, and what effect it would have. Further, writing good bugs is likely to take a significant amount of time, for a very indirect benefit. Hence such a solution is not sufficient.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Seeding Artificial Faults with Mutation Analysis
#
# Mutation Analysis offers an alternative solution to assess the effectiveness of a test suite. The idea of mutation analysis is to seed _artificial faults_, known as *mutations*, into the program code, and to check whether the test suite finds them. Such a mutation could, for instance, replace a `+` by a `-` somewhere within `execute_the_program_as_a_whole()`. Of course, the above ineffective tests would not detect this, as they do not check any of the results. An effective test would, however; and the assumption is that the more effective a test is in finding _artificial_ faults, the more effective it would be in finding _real_ faults.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# The insight from Mutation Analysis is to consider the probability of insertion of a bug from the perspective of a programmer. If one assumes that the attention received by each program element in the program is sufficiently similar, one can further assume that each token in the program has a similar probability of being incorrectly transcribed. Of course, the programmer will correct any mistakes that gets detected by the compilers (or other static analysis tools). So the set of valid tokens different from the original that make it past the compilation stage is considered to be its possible set of _mutations_ that represent the _probable faults_ in the program. A test suite is then judged by its capability to detect (and hence prevent) such mutations. The proportion of such mutants detected over all _valid_ mutants produced is taken as the mutation score. In this chapter, we see how one can implement Mutation Analysis in Python programs. The mutation score obtained represents the ability of any program analysis tools to prevent faults, and can be used to judge static test suites, test generators such as fuzzers, and also static and symbolic execution frameworks.
# -
# It might be intuitive to consider a slightly different perspective. A test suite is a program that can be considered to accept as its input, the program to be tested. What is the best way to evaluate such a program (the test suite)? We can essentially *fuzz* the test suite by applying small mutations to the input program, and verifying that the test suite in question does not produce unexpected behaviors. The test suite is supposed to only allow the original through; and hence any mutant that is not detected as faulty represents a bug in the test suite.
# ## Structural Coverage Adequacy by Example
#
# Let us introduce a more detailed example to illustrate both the problems with coverage as well as how mutation analysis works. The `triangle()` program below classifies a triangle with edge lengths $a$, $b$, and $c$ into the proper triangle category. We want to verify that the program works correctly.
def triangle(a, b, c):
if a == b:
if b == c:
return 'Equilateral'
else:
return 'Isosceles'
else:
if b == c:
return "Isosceles"
else:
if a == c:
return "Isosceles"
else:
return "Scalene"
# Here are a few test cases to ensure that the program works.
def strong_oracle(fn):
assert fn(1, 1, 1) == 'Equilateral'
assert fn(1, 2, 1) == 'Isosceles'
assert fn(2, 2, 1) == 'Isosceles'
assert fn(1, 2, 2) == 'Isosceles'
assert fn(1, 2, 3) == 'Scalene'
# Running them actually causes all tests to pass.
strong_oracle(triangle)
# However, the statement that "all tests pass" has value only if we know that our tests are effective.
# What is the effectiveness of our test suite? As we saw in the [chapter on coverage](Coverage.ipynb), one can use structural coverage techniques such as statement coverage to obtain a measure of effectiveness of the test case.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import bookutils
# -
from Coverage import Coverage
import inspect
# We add a function `show_coverage()` to visualize the coverage obtained.
class Coverage(Coverage):
def show_coverage(self, fn):
src = inspect.getsource(fn)
name = fn.__name__
covered = set([lineno for method,
lineno in self._trace if method == name])
for i, s in enumerate(src.split('\n')):
print('%s %2d: %s' % ('#' if i + 1 in covered else ' ', i + 1, s))
with Coverage() as cov:
strong_oracle(triangle)
cov.show_coverage(triangle)
# Our `strong_oracle()` seems to have adequately covered all possible conditions.
# That is, our set of test cases is reasonably good according to structural coverage. However, does the coverage obtained tell the whole story? Consider this test suite instead:
def weak_oracle(fn):
assert fn(1, 1, 1) == 'Equilateral'
assert fn(1, 2, 1) != 'Equilateral'
assert fn(2, 2, 1) != 'Equilateral'
assert fn(1, 2, 2) != 'Equilateral'
assert fn(1, 2, 3) != 'Equilateral'
# All that we are checking here is that a triangle with unequal sides is not equilateral. What is the coverage obtained?
with Coverage() as cov:
weak_oracle(triangle)
cov.show_coverage(triangle)
# Indeed, there does not seem to be _any_ difference in coverage.
# The `weak_oracle()` obtains exactly the same coverage as that of `strong_oracle()`. However, a moment's reflection should convince one that the `weak_oracle()` is not as effective as `strong_oracle()`. However, _coverage_ is unable to distinguish between the two test suites. What are we missing in coverage?
# The problem here is that coverage is unable to evaluate the _quality_ of our assertions. Indeed, coverage does not care about assertions at all. However, as we saw above, assertions are an extremely important part of test suite effectiveness. Hence, what we need is a way to evaluate the quality of assertions.
# ## Injecting Artificial Faults
#
# Notice that in the [chapter on coverage](Coverage.ipynb), coverage was presented as a _proxy_ for the likelihood of a test suite to uncover bugs. What if we actually try to evaluate the likelihood of a test suite to uncover bugs? All we need is to inject bugs into the program, one at a time, and count the number of such bugs that our test suite detects. The frequency of detection will provide us with the actual likelihood of the test suite to uncover bugs. This technique is called _fault injection_. Here is an example for _fault injection_.
def triangle_m1(a, b, c):
if a == b:
if b == c:
return 'Equilateral'
else:
# return 'Isosceles'
return None # <-- injected fault
else:
if b == c:
return "Isosceles"
else:
if a == c:
return "Isosceles"
else:
return "Scalene"
# Let us see if our test suites are good enough to catch this fault. We first check whether `weak_oracle()` can detect this change.
from ExpectError import ExpectError
with ExpectError():
weak_oracle(triangle_m1)
# The `weak_oracle()` is unable to detect any changes. What about our `strong_oracle()`?
with ExpectError():
strong_oracle(triangle_m1)
# Our `strong_oracle()` is able to detect this fault, which is evidence that `strong_oracle()` is probably a better test suite.
# _Fault injection_ can provide a good measure of effectiveness of a test suite, provided we have a list of possible faults. The problem is that collecting such a set of _unbiased_ faults is rather expensive. It is difficult to create good faults that are reasonably hard to detect, and it is a manual process. Given that it is a manual process, the generated faults will be biased by the preconceptions of the developer who creates it. Even when such curated faults are available, they are unlikely to be exhaustive, and likely to miss important classes of bugs, and parts of the program. Hence, _fault injection_ is an insufficient replacement for coverage. Can we do better?
# Mutation Analysis provides an alternative to a curated set of faults. The key insight is that, if one assumes that the programmer understands the program in question, the majority of errors made are very likely small transcription errors (a small number of tokens). A compiler will likely catch most of these errors. Hence, the majority of residual faults in a program is likely to be due to small (single token) variations at certain points in the structure of the program from the correct program (This particular assumption is called the *Competent Programmer Hypothesis* or the *Finite Neighborhood Hypothesis*).
#
# What about the larger faults composed of multiple smaller faults? The key insight here is that, for a majority of such complex faults, test cases that detect a single smaller fault in isolation is very likely to detect the larger complex fault that contains it. (This assumption is called the *Coupling Effect*.)
#
# How can we use these assumptions in practice? The idea is to simply generate *all* possible *valid* variants of the program that differs from the original by a small change (such as a single token change) (Such variants are called *mutants*). Next, the given test suite is applied to each variant thus generated. Any mutant detected by the test suite is said to have been *killed* by the test suite. The effectiveness of a test suite is given by the proportion of mutants killed to the valid mutants generated.
# We next implement a simple mutation analysis framework and use it to evaluate our test suites.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Mutating Python Code
#
# To manipulate a Python program, we work on the _abstract syntax tree_ (AST) representation – which is the internal representation compilers and interpreters work on after reading in the program text.
#
# Briefly speaking, we convert the program into a tree, and then _change parts of this tree_ – for instance, by changing `+` operators into `-` or vice versa, or actual statements into `pass` statements that do nothing. The resulting mutated tree can then be processed further; it can be passed on to the Python interpreter for execution, or we can _unparse_ it back into a textual form.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# We begin by importing the AST manipulation modules.
# -
import ast
import astor
import inspect
# We can get the source of a Python function using `inspect.getsource()`. (Note that this does not work for functions defined in other notebooks.)
triangle_source = inspect.getsource(triangle)
triangle_source
# To view these in a visually pleasing form, our function `print_content(s, suffix)` formats and highlights the string `s` as if it were a file with ending `suffix`. We can thus view (and highlight) the source as if it were a Python file:
from bookutils import print_content
print_content(triangle_source, '.py')
# Parsing this gives us an abstract syntax tree (AST) – a representation of the program in tree form.
triangle_ast = ast.parse(triangle_source)
# What does this AST look like? The helper functions `astor.dump_tree()` (textual output) and `showast.show_ast()` (graphical output with [showast](https://github.com/hchasestevens/show_ast)) allow us to inspect the structure of the tree. We see that the function starts as a `FunctionDef` with name and arguments, followed by a body, which is a list of statements; in this case, the body contains only an `If`, which itself contains other nodes of type `If`, `Compare`, `Name`, `Str`, and `Return`.
print(astor.dump_tree(triangle_ast))
# Too much text for you? This graphical representation may make things simpler.
from bookutils import rich_output
if rich_output():
import showast
showast.show_ast(triangle_ast)
# The function `astor.to_source()` converts such a tree back into the more familiar textual Python code representation.
print_content(astor.to_source(triangle_ast), '.py')
# ## A Simple Mutator for Functions
#
# Let us now go and mutate the `triangle()` program. A simple way to produce valid mutated version of this program is to replace some of its statements by `pass`.
#
# The `MuFunctionAnalyzer` is the main class responsible for mutation analysis of the test suite. It accepts the function to be tested. It normalizes the source code given by parsing and unparsing it once, using the functions discussed above. This is required to ensure that later `diff`s between the original and mutant are not derailed by differences in whitespace, comments, etc.
class MuFunctionAnalyzer:
def __init__(self, fn, log=False):
self.fn = fn
self.name = fn.__name__
src = inspect.getsource(fn)
self.ast = ast.parse(src)
self.src = astor.to_source(self.ast) # normalize
self.mutator = self.mutator_object()
self.nmutations = self.get_mutation_count()
self.un_detected = set()
self.mutants = []
self.log = log
def mutator_object(self, locations=None):
return StmtDeletionMutator(locations)
def register(self, m):
self.mutants.append(m)
def finish(self):
pass
# The `get_mutation_count()` fetches the number of possible mutations available. We will see later how this can be implemented.
class MuFunctionAnalyzer(MuFunctionAnalyzer):
def get_mutation_count(self):
self.mutator.visit(self.ast)
return self.mutator.count
# The `Mutator` provides the base class for implementing individual mutations. It accepts a list of locations to mutate. It assumes that the method `mutable_visit()` is invoked on all nodes of interest as determined by the subclass. When the `Mutator` is invoked without a list of locations to mutate, it simply loops through all possible mutation points and retains a count in `self.count`. If it is invoked with a specific list of locations to mutate, the `mutable_visit()` method calls the `mutation_visit()` which performs the mutation on the node. Note that a single location can produce multiple mutations. (Hence the hashmap).
class Mutator(ast.NodeTransformer):
def __init__(self, mutate_location=-1):
self.count = 0
self.mutate_location = mutate_location
def mutable_visit(self, node):
self.count += 1 # statements start at line no 1
if self.count == self.mutate_location:
return self.mutation_visit(node)
return self.generic_visit(node)
# The `StmtDeletionMutator` simply hooks into all the statement processing visitors. It performs mutation by replacing the given statement with `pass`. As you can see, it visits all kinds of statements.
class StmtDeletionMutator(Mutator):
def visit_Return(self, node): return self.mutable_visit(node)
def visit_Delete(self, node): return self.mutable_visit(node)
def visit_Assign(self, node): return self.mutable_visit(node)
def visit_AnnAssign(self, node): return self.mutable_visit(node)
def visit_AugAssign(self, node): return self.mutable_visit(node)
def visit_Raise(self, node): return self.mutable_visit(node)
def visit_Assert(self, node): return self.mutable_visit(node)
def visit_Global(self, node): return self.mutable_visit(node)
def visit_Nonlocal(self, node): return self.mutable_visit(node)
def visit_Expr(self, node): return self.mutable_visit(node)
def visit_Pass(self, node): return self.mutable_visit(node)
def visit_Break(self, node): return self.mutable_visit(node)
def visit_Continue(self, node): return self.mutable_visit(node)
# The actual mutation consists of replacing the node with a `pass` statement:
class StmtDeletionMutator(StmtDeletionMutator):
def mutation_visit(self, node): return ast.Pass()
# For `triangle()`, this visitor produces five mutations – namely, replacing the five `return` statements with `pass`:
MuFunctionAnalyzer(triangle).nmutations
# We need a way to obtain the individual mutants. For this, we convert our `MuFunctionAnalyzer` to an *iterable*.
class MuFunctionAnalyzer(MuFunctionAnalyzer):
def __iter__(self):
return PMIterator(self)
# The `PMIterator`, which is the *iterator* class for `MuFunctionAnalyzer` is defined as follows.
class PMIterator:
def __init__(self, pm):
self.pm = pm
self.idx = 0
# The `next()` method returns the corresponding `Mutant`:
class PMIterator(PMIterator):
def __next__(self):
i = self.idx
if i >= self.pm.nmutations:
self.pm.finish()
raise StopIteration()
self.idx += 1
mutant = Mutant(self.pm, self.idx, log=self.pm.log)
self.pm.register(mutant)
return mutant
# The `Mutant` class contains logic for generating mutants when given the locations to mutate.
class Mutant:
def __init__(self, pm, location, log=False):
self.pm = pm
self.i = location
self.name = "%s_%s" % (self.pm.name, self.i)
self._src = None
self.tests = []
self.detected = False
self.log = log
# Here is how it can be used:
for m in MuFunctionAnalyzer(triangle):
print(m.name)
# These names are a bit generic yet. Let's see whether we can get more insights into the mutations produced.
# The `generate_mutant()` simply calls the `mutator()` method, and passes the mutator a copy of the AST.
class Mutant(Mutant):
def generate_mutant(self, location):
mutant_ast = self.pm.mutator_object(
location).visit(ast.parse(self.pm.src)) # copy
return astor.to_source(mutant_ast)
# The `src()` method returns the mutated source.
class Mutant(Mutant):
def src(self):
if self._src is None:
self._src = self.generate_mutant(self.i)
return self._src
# Here is how one can obtain the mutants, and visualize the difference from the original:
import difflib
for mutant in MuFunctionAnalyzer(triangle):
shape_src = mutant.pm.src
for line in difflib.unified_diff(mutant.pm.src.split('\n'),
mutant.src().split('\n'),
fromfile=mutant.pm.name,
tofile=mutant.name, n=3):
print(line)
# In this `diff` output, lines prefixed with `+` are added, whereas lines prefixed with `-` are deleted. We see that each of the five mutants indeed replaces a return statement with a `pass` statement.
# We add the `diff()` method to `Mutant` so that it can be called directly.
class Mutant(Mutant):
def diff(self):
return '\n'.join(difflib.unified_diff(self.pm.src.split('\n'),
self.src().split('\n'),
fromfile='original',
tofile='mutant',
n=3))
# ## Evaluating Mutations
#
# We are now ready to implement the actual evaluation. We define our mutant as a _context manager_ that verifies whether all assertions given succeed. The idea is that we can write code such as
#
# ```python
# for mutant in MuFunctionAnalyzer(function):
# with mutant:
# assert function(x) == y
# ```
#
# and while `mutant` is active (i.e., the code block under `with:`), the original function is replaced by the mutated function.
# The `__enter__()` function is called when the `with` block is entered. It creates the mutant as a Python function and places it in the global namespace, such that the `assert` statement executes the mutated function rather than the original.
class Mutant(Mutant):
def __enter__(self):
if self.log:
print('->\t%s' % self.name)
c = compile(self.src(), '<mutant>', 'exec')
eval(c, globals())
# The `__exit__()` function checks whether an exception has occurred (i.e., the assertion failed, or some other error was raised); if so, it marks the mutation as `detected`. Finally, it restores the original function definition.
class Mutant(Mutant):
def __exit__(self, exc_type, exc_value, traceback):
if self.log:
print('<-\t%s' % self.name)
if exc_type is not None:
self.detected = True
if self.log:
print("Detected %s" % self.name, exc_type, exc_value)
globals()[self.pm.name] = self.pm.fn
if self.log:
print()
return True
# The `finish()` method simply invokes the method on the mutant, checks if the mutant was discovered, and returns the result.
from ExpectError import ExpectTimeout
class MuFunctionAnalyzer(MuFunctionAnalyzer):
def finish(self):
self.un_detected = {
mutant for mutant in self.mutants if not mutant.detected}
# The mutation score – the ratio of mutants detected by the test suite - is computed by `score()`. A score of 1.0 means that all mutants were discovered; a score of 0.1 means that only 10% of mutants were detected.
class MuFunctionAnalyzer(MuFunctionAnalyzer):
def score(self):
return (self.nmutations - len(self.un_detected)) / self.nmutations
# Here is how we use our framework.
import sys
for mutant in MuFunctionAnalyzer(triangle, log=True):
with mutant:
assert triangle(1, 1, 1) == 'Equilateral', "Equal Check1"
assert triangle(1, 0, 1) != 'Equilateral', "Equal Check2"
assert triangle(1, 0, 2) != 'Equilateral', "Equal Check3"
mutant.pm.score()
# Only one out of five mutations resulted in a failing assertion. Hence, the `weak_oracle()` test suite gets a mutation score of 20%.
for mutant in MuFunctionAnalyzer(triangle):
with mutant:
weak_oracle(triangle)
mutant.pm.score()
# Since we are modifying the global namespace, we do not have to refer to the function directly within the for loop of mutant.
def oracle():
strong_oracle(triangle)
for mutant in MuFunctionAnalyzer(triangle, log=True):
with mutant:
oracle()
mutant.pm.score()
# That is, we were able to achieve `100%` mutation score with the `strong_oracle()` test suite.
#
# Here is another example. `gcd()` computes the greatest common divisor of two numbers.
def gcd(a, b):
if a < b:
c = a
a = b
b = c
while b != 0:
c = a
a = b
b = c % b
return a
# Here's a test for it. How effective is it?
for mutant in MuFunctionAnalyzer(gcd, log=True):
with mutant:
assert gcd(1, 0) == 1, "Minimal"
assert gcd(0, 1) == 1, "Mirror"
mutant.pm.score()
# We see that our `TestGCD` test suite is able to obtain a mutation score of 42%.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Mutator for Modules and Test Suites
#
# Consider the `triangle()` program we discussed previously. As we discussed, a simple way to produce valid mutated version of this program is to replace some of its statements by `pass`.
# -
# For demonstration purposes, we would like to proceed as though the program was in a different file. We can do that by producing a `Module` object in Python, and attaching the function to it.
import types
def import_code(code, name):
module = types.ModuleType(name)
exec(code, module.__dict__)
return module
# We attach the `triangle()` function to the `shape` module.
shape = import_code(shape_src, 'shape')
# We can now invoke triangle through the module `shape`.
shape.triangle(1, 1, 1)
# We want to test the `triangle()` function. For that, we define a `StrongShapeTest` class as below.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
import unittest
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
class StrongShapeTest(unittest.TestCase):
def test_equilateral(self):
assert shape.triangle(1, 1, 1) == 'Equilateral'
def test_isosceles(self):
assert shape.triangle(1, 2, 1) == 'Isosceles'
assert shape.triangle(2, 2, 1) == 'Isosceles'
assert shape.triangle(1, 2, 2) == 'Isosceles'
def test_scalene(self):
assert shape.triangle(1, 2, 3) == 'Scalene'
# -
# We define a helper function `suite()` that looks through a given class and identifies the test functions.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
def suite(test_class):
suite = unittest.TestSuite()
for f in test_class.__dict__:
if f.startswith('test_'):
suite.addTest(test_class(f))
return suite
# -
# The tests in `TestTriangle` class can be invoked with different test runners. The simplest is to directly invoke the `run()` method of the `TestCase`.
suite(StrongShapeTest).run(unittest.TestResult())
# The `TextTestRunner` class provides ability to control the verbosity of execution. It also allows one to return on the *first* failure.
# + button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"}
runner = unittest.TextTestRunner(verbosity=0, failfast=True)
runner.run(suite(StrongShapeTest))
# -
# Running the program under coverage is accomplished as follows:
with Coverage() as cov:
suite(StrongShapeTest).run(unittest.TestResult())
# The coverage obtained is given by:
cov.show_coverage(triangle)
class WeakShapeTest(unittest.TestCase):
def test_equilateral(self):
assert shape.triangle(1, 1, 1) == 'Equilateral'
def test_isosceles(self):
assert shape.triangle(1, 2, 1) != 'Equilateral'
assert shape.triangle(2, 2, 1) != 'Equilateral'
assert shape.triangle(1, 2, 2) != 'Equilateral'
def test_scalene(self):
assert shape.triangle(1, 2, 3) != 'Equilateral'
# How much coverage does it obtain?
with Coverage() as cov:
suite(WeakShapeTest).run(unittest.TestResult())
cov.show_coverage(triangle)
# The `MuProgramAnalyzer` is the main class responsible for mutation analysis of the test suite. It accepts the name of the module to be tested, and its source code. It normalizes the source code given by parsing and unparsing it once. This is required to ensure that later `diff`s between the original and mutant are not derailed by differences in whitespace, comments, etc.
class MuProgramAnalyzer(MuFunctionAnalyzer):
def __init__(self, name, src):
self.name = name
self.ast = ast.parse(src)
self.src = astor.to_source(self.ast)
self.changes = []
self.mutator = self.mutator_object()
self.nmutations = self.get_mutation_count()
self.un_detected = set()
def mutator_object(self, locations=None):
return AdvStmtDeletionMutator(self, locations)
# We now extend the `Mutator` class.
class AdvMutator(Mutator):
def __init__(self, analyzer, mutate_locations=None):
self.count = 0
self.mutate_locations = [] if mutate_locations is None else mutate_locations
self.pm = analyzer
def mutable_visit(self, node):
self.count += 1 # statements start at line no 1
return self.mutation_visit(node)
# The `AdvStmtDeletionMutator` simply hooks into all the statement processing visitors. It performs mutation by replacing the given statement with `pass`.
class AdvStmtDeletionMutator(AdvMutator, StmtDeletionMutator):
def __init__(self, analyzer, mutate_locations=None):
AdvMutator.__init__(self, analyzer, mutate_locations)
def mutation_visit(self, node):
index = 0 # there is only one way to delete a statement -- replace it by pass
if not self.mutate_locations: # counting pass
self.pm.changes.append((self.count, index))
return self.generic_visit(node)
else:
# get matching changes for this pass
mutating_lines = set((count, idx)
for (count, idx) in self.mutate_locations)
if (self.count, index) in mutating_lines:
return ast.Pass()
else:
return self.generic_visit(node)
# Aagin, we can obtain the number of mutations produced for `triangle()` as follows.
MuProgramAnalyzer('shape', shape_src).nmutations
# We need a way to obtain the individual mutants. For this, we convert our `MuProgramAnalyzer` to an *iterable*.
class MuProgramAnalyzer(MuProgramAnalyzer):
def __iter__(self):
return AdvPMIterator(self)
# The `AdvPMIterator`, which is the *iterator* class for `MuProgramAnalyzer` is defined as follows.
class AdvPMIterator:
def __init__(self, pm):
self.pm = pm
self.idx = 0
# The `next()` method returns the corresponding `Mutant`
class AdvPMIterator(AdvPMIterator):
def __next__(self):
i = self.idx
if i >= len(self.pm.changes):
raise StopIteration()
self.idx += 1
# there could be multiple changes in one mutant
return AdvMutant(self.pm, [self.pm.changes[i]])
# The `Mutant` class contains logic for generating mutants when given the locations to mutate.
class AdvMutant(Mutant):
def __init__(self, pm, locations):
self.pm = pm
self.i = locations
self.name = "%s_%s" % (self.pm.name,
'_'.join([str(i) for i in self.i]))
self._src = None
# Here is how it can be used:
shape_src = inspect.getsource(triangle)
for m in MuProgramAnalyzer('shape', shape_src):
print(m.name)
# The `generate_mutant()` simply calls the `mutator()` method, and passes the mutator a copy of the AST.
class AdvMutant(AdvMutant):
def generate_mutant(self, locations):
mutant_ast = self.pm.mutator_object(
locations).visit(ast.parse(self.pm.src)) # copy
return astor.to_source(mutant_ast)
# The `src()` method returns the mutated source.
class AdvMutant(AdvMutant):
def src(self):
if self._src is None:
self._src = self.generate_mutant(self.i)
return self._src
# Again, we visualize mutants as difference from the original:
import difflib
# We add the `diff()` method to `Mutant` so that it can be called directly.
class AdvMutant(AdvMutant):
def diff(self):
return '\n'.join(difflib.unified_diff(self.pm.src.split('\n'),
self.src().split('\n'),
fromfile='original',
tofile='mutant',
n=3))
for mutant in MuProgramAnalyzer('shape', shape_src):
print(mutant.name)
print(mutant.diff())
break
# We are now ready to implement the actual evaluation. For doing that, we require the ability to accept the module where the test suite is defined, and invoke the test method on it. The method `getitem` accepts the test module, fixes the import entries on the test module to correctly point to the mutant module, and passes it to the test runner `MutantTestRunner`.
class AdvMutant(AdvMutant):
def __getitem__(self, test_module):
test_module.__dict__[
self.pm.name] = import_code(
self.src(), self.pm.name)
return MutantTestRunner(self, test_module)
# The `MutantTestRunner` simply calls all `test_` methods on the test module, checks if the mutant was discovered, and returns the result.
from ExpectError import ExpectTimeout
class MutantTestRunner:
def __init__(self, mutant, test_module):
self.mutant = mutant
self.tm = test_module
def runTest(self, tc):
suite = unittest.TestSuite()
test_class = self.tm.__dict__[tc]
for f in test_class.__dict__:
if f.startswith('test_'):
suite.addTest(test_class(f))
runner = unittest.TextTestRunner(verbosity=0, failfast=True)
try:
with ExpectTimeout(1):
res = runner.run(suite)
if res.wasSuccessful():
self.mutant.pm.un_detected.add(self)
return res
except SyntaxError:
print('Syntax Error (%s)' % self.mutant.name)
return None
raise Exception('Unhandled exception during test execution')
# The mutation score is computed by `score()`.
class MuProgramAnalyzer(MuProgramAnalyzer):
def score(self):
return (self.nmutations - len(self.un_detected)) / self.nmutations
# Here is how we use our framework.
import sys
test_module = sys.modules[__name__]
for mutant in MuProgramAnalyzer('shape', shape_src):
mutant[test_module].runTest('WeakShapeTest')
mutant.pm.score()
# The `WeakShape` test suite resulted in only `20%` mutation score.
for mutant in MuProgramAnalyzer('shape', shape_src):
mutant[test_module].runTest('StrongShapeTest')
mutant.pm.score()
# On the other hand, we were able to achieve `100%` mutation score with `StrongShapeTest` test suite.
#
# Here is another example, `gcd()`.
gcd_src = inspect.getsource(gcd)
class TestGCD(unittest.TestCase):
def test_simple(self):
assert cfg.gcd(1, 0) == 1
def test_mirror(self):
assert cfg.gcd(0, 1) == 1
for mutant in MuProgramAnalyzer('cfg', gcd_src):
mutant[test_module].runTest('TestGCD')
mutant.pm.score()
# We see that our `TestGCD` test suite is able to obtain `42%` mutation score.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "slide"}
# ## The Problem of Equivalent Mutants
#
# One of the problems with mutation analysis is that not all mutants generated need to be faulty. For example, consider the `new_gcd()` program below.
# -
def new_gcd(a, b):
if a < b:
a, b = b, a
else:
a, b = a, b
while b != 0:
a, b = b, a % b
return a
# This program can be mutated to produce the following mutant.
def gcd(a, b):
if a < b:
a, b = b, a
else:
pass
while b != 0:
a, b = b, a % b
return a
for i, mutant in enumerate(MuFunctionAnalyzer(new_gcd)):
print(i,mutant.src())
# While other mutants are faulty compared to the original, `mutant 1` is indistinguishable from the original in terms of its semantics because it removes an inconsequential assignment. This means that `mutant 1` does not represent a fault. These kinds of mutants that do not represent a fault are called *Equivalent mutants*. The problem with equivalent mutants is that it becomes very difficult to judge the mutation score in the presence of equivalent mutants. For example, with a mutation score of 70%, anywhere from 0 to 30% of the mutants may be equivalent. Hence, without knowing the actual number of equivalent mutants, it is impossible to judge how much the tests can be improved. We discuss two methods to deal with equivalent mutants.
# ### Statistical Estimation of Number of Equivalent Mutants
#
# If the number of mutants that are alive is small enough, one may rely on simply inspecting them manually. However, if the number of mutants are sufficiently large (say > 1000), one may choose a smaller number of mutants from the alive mutants randomly and manually evaluate them to see whether they represent faults. The sample size determination is governed by the following formula for a binomial distribution (approximated by a normal distribution):
#
# $$
# n \ge \hat{p}(1-\hat{p})\bigg(\frac{Z_{\frac{\alpha}{2}}}{\Delta}\bigg)^2
# $$
# where $n$ is the number of samples, $p$ is the parameter for the probability distribution, $\alpha$ is the accuracy desired, $\Delta$ the precision. For an accuracy of $95\%$, $Z_{0.95}=1.96$. we have the following values (the maximum value of $\hat{p}(1-\hat{p}) = 0.25$) and $Z$ is the critical value for normal distribution:
# $$
# n \ge 0.25\bigg(\frac{1.96}{\Delta}\bigg)^2
# $$
# For $\Delta = 0.01$, (that is for a maximum error of 1%), we need to evaluate $9604$ mutants for equivalence. If one relaxes the constraint to $\Delta = 0.1$ (that is an error of $10\%$), then one needs to evaluate only $96$ mutants for equivalence.
# ### Statistical Estimation of the Number of Immortals by Chao's Estimator
#
# While the idea of sampling only a limited number of mutants is appealing, it is still limited in that manual analysis is necessary. If computing power is cheap, another way to estimate the number of true mutants (and hence the number of equivalent mutants) is by means of Chao's estimator. As we will see in the chapter on [when to stop fuzzing](WhenToStopFuzzing.ipynb), the formula is given by:
#
# $$
# \hat S_\text{Chao1} = \begin{cases}
# S(n) + \frac{f_1^2}{2f_2} & \text{if $f_2>0$}\\
# S(n) + \frac{f_1(f_1-1)}{2} & \text{otherwise}
# \end{cases}
# $$
# The basic idea is to compute the result of the complete test matrix $T \times M$ of each test against each mutant. The variable $f_1$ represents the number of mutants that were killed exactly once, and the variable $f_2$ represents the number of variables that were killed exactly twice. $S(n)$ is the total number of mutants killed. Here, $\hat{S}_{Chao1}$ provides the estimate of the true number of mutants. If $M$ is the total mutants generated, then $M - \hat{S}_{Chao1}$ represents the number of **immortal** mutants. Note that these **immortal** mutants are somewhat different from the traditional equivalent mutants in that the **mortality** depends on the oracle used to distinguish variant behavior. That is, if one uses a fuzzer that relies on errors thrown to detect killing, it will not detect mutants that produce different output but does not throw an error. Hence, the *Chao1* estimate will essentially be the asymptote value of mutants the fuzzer can detect if it is given an infinite amount of time. The **immortal** mutant estimate will approach true **equivalent** mutant estimate when the oracle used is sufficiently strong.
# For more details see the chapter on [when to stop fuzzing](WhenToStopFuzzing.ipynb). A comprehensive guide to species discovery in testing is the paper by Boehme \cite{boehme2018species}.
# ## Synopsis
# This chapter introduces two methods of running *mutation analysis* on subject programs. The first class `MuFunctionAnalyzer` targets individual functions. Given a function `gcd` and two test cases evaluate, one can run mutation analysis on the test cases as follows:
for mutant in MuFunctionAnalyzer(gcd, log=True):
with mutant:
assert gcd(1, 0) == 1, "Minimal"
assert gcd(0, 1) == 1, "Mirror"
mutant.pm.score()
# The second class `MuProgramAnalyzer` targets standalone programs with test suites. Given a program `gcd` whose source code is provided in `gcd_src` and the test suite is provided by `TestGCD`, one can evaluate the mutation score of `TestGCD` as follows:
class TestGCD(unittest.TestCase):
def test_simple(self):
assert cfg.gcd(1, 0) == 1
def test_mirror(self):
assert cfg.gcd(0, 1) == 1
for mutant in MuProgramAnalyzer('gcd', gcd_src):
mutant[test_module].runTest('TestGCD')
mutant.pm.score()
# The mutation score thus obtained is a better indicator of the quality of a given test suite than pure coverage.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Lessons Learned
#
# * We have learned why structural coverage is insufficient to evaluate the quality of test suites.
# * We have learned how to use Mutation Analysis for evaluating test suite quality.
# * We have learned the limitations of Mutation Analysis -- Equivalent and Redundant mutants, and how to estimate them.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Next Steps
#
# * While naive fuzzing generates poor quality oracles, techniques such as [symbolic](SymbolicFuzzer.ipynb) and [concolic](ConcolicFuzzer.ipynb) can enhance the quality oracles used in fuzzing.
# * [Dynamic invariants](DynamicInvariants.ipynb) can also be of great help in improving the quality of oracles.
# * The chapter on [when to stop fuzzing](WhenToStopFuzzing.ipynb) provides a detailed overview of the Chao estimator.
# -
# ## Background
#
# The idea of Mutation Analysis was first introduced by Lipton et al. \cite{lipton1971fault}. An excellent survey of mutation analysis research was published by Jia et al. \cite{jia2011analysis}. The chapter on Mutation Analysis by Papadakis et al \cite{papadakis2019mutation} is another excellent overview of the current trends in mutation analysis.
# + [markdown] button=false new_sheet=true run_control={"read_only": false}
# ## Exercises
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Exercise 1: Arithmetic Expression Mutators
#
# Our simple statement deletion mutation is only one of the ways in which a program could be mutated. Another category of mutants is _expression mutation_ where arithmetic operators such as `{+,-,*,/}` etc are replaced for one another. For example, given an expression such as
# ```
# x = x + 1
# ```
# One can mutate it to
# ```
# x = x - 1
# ```
# and
# ```
# x = x * 1
# ```
# and
# ```
# x = x / 1
# ```
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# First, we need to find out which node types we want to mutate. We get these via the ast functions and find that the node type is named BinOp
# + button=false new_sheet=false run_control={"read_only": false}
print(astor.dump_tree(ast.parse("1 + 2 - 3 * 4 / 5")))
# + [markdown] button=false new_sheet=false run_control={"read_only": false} solution2="hidden" solution2_first=true
# To mutate the tree, you thus need to change the `op` attribute (which has one of the values `Add`, `Sub`, `Mult`, and `Div`)
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** First, we need to find out which node types we want to mutate. We get these via the `ast` functions and find that the node type is named `BinOp`
# + slideshow={"slide_type": "skip"} solution2="hidden"
print(astor.dump_tree(ast.parse("1 + 2 - 3 * 4 / 5")))
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden" solution2_first=true
# To mutate the tree, we need to change the `op` attribute (which has one of the values `Add`, `Sub`, `Mult`, and `Div`). Write a class `BinOpMutator` that does the necessary mutations, and then create a class `MuBinOpAnalyzer` as subclass of `MuFunctionAnalyzer` which makes use of `BinOpMutator`.
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# **Solution.** As with `StmtDeletionMutator`, we have to visit the AST – in this case, all `BinOp` nodes – and make appropriate changes:
# + slideshow={"slide_type": "skip"} solution2="hidden"
class BinOpMutator(Mutator):
def visit_BinOp(self, node): return self.mutable_visit(node)
# + slideshow={"slide_type": "skip"} solution2="hidden"
class BinOpMutator(BinOpMutator):
def mutation_visit(self, node):
replacement = {
type(ast.Add()): ast.Sub(),
type(ast.Sub()): ast.Add(),
type(ast.Mult()): ast.Div(),
type(ast.Div()): ast.Mult()
}
try:
node.op = replacement[type(node.op)]
except KeyError:
pass # All other binary operators (and, mod, etc.)
return node
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# We hook this into our own analyzer:
# + slideshow={"slide_type": "skip"} solution2="hidden"
class MuBinOpAnalyzer(MuFunctionAnalyzer):
def mutator_object(self, locations=None):
return BinOpMutator(locations)
# + [markdown] slideshow={"slide_type": "skip"} solution2="hidden"
# Here's how it mutates a function:
# + slideshow={"slide_type": "skip"} solution2="hidden"
def arith_expr():
return 1 + 2 - 3 * 4 / 5
# + slideshow={"slide_type": "skip"} solution2="hidden"
for mutant in MuBinOpAnalyzer(arith_expr, log=True):
print(mutant.diff())
# + [markdown] button=false new_sheet=false run_control={"read_only": false} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# ### Exercise 2: Optimizing Mutation Analysis
#
# Our technique for mutation analysis is somewhat inefficient in that we run the tests even on mutants that have mutations in code not covered by the test case. Test cases have no possibility of detecting errors on portions of code they do not cover. Hence, one of the simplest optimizations is to first recover the coverage information from the given test case, and only run the test case on mutants where the mutations lie in the code being covered by the test case. Can you modify the `MuFunctionAnalyzer` to incorporate recovering coverage as the first step?
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden"
# **Solution.** Left to the astute reader.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# ### Exercise 3: Byte Code Mutator
#
# We have seen how to mutate the AST given the source. One of the deficiencies with this approach is that the Python bytecode is targeted by other languages too. In such cases, the source may not be readily converted to a Python AST, and it is desirable to mutate the bytecode instead. Can you implement a bytecode mutator for Python function that mutates the bytecode instead of fetching the source and then mutating it?
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden"
# **Solution.** Left to the astute reader.
# + [markdown] button=false new_sheet=false run_control={"read_only": false} solution="hidden" solution2="hidden" solution2_first=true solution_first=true
# ### Exercise 4: Estimating Residual Defect Density
#
# The defect density of a program is the number of defects in a program that that were detected before release divided by the program size. The residual defect density is the percentage of defects that escaped detection. While estimation of the real residual defect density is difficult, mutation analysis can provide an upper bound. The number of mutants that remain undetected is a plausible upper bound on the number of defects that remain within the program. However, this upper bound may be too wide. The reason is that some of the remaining faults can interact with each other, and if present together, can be detected by the available test suite. Hence, a tighter bound is the number of mutants that can exist *together* in a given program without being detected by the given test suite. This can be accomplished by starting with the complete set of mutations possible, and applying delta-debugging from [the chapter on reducing](Reducer.ipynb) to determine the minimum number of mutations that need to be removed to make the mutant pass undetected by the test suite. Can you produce a new `RDDEstimator` by extending the `MuFunctionAnalyzer` that estimates the residual defect density upper bound using this technique?
# + [markdown] button=false new_sheet=false run_control={"read_only": false} slideshow={"slide_type": "skip"} solution="hidden" solution2="hidden"
# **Solution.** Left to the astute reader.
| notebooks/MutationAnalysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GW Example
#
# ## v2 -- Refactor with localization
# ## v3 -- Refactor with PATH
# %matplotlib notebook
# +
# imports
from importlib import reload
import os
from pkg_resources import resource_filename
import numpy as np
import healpy as hp
import pandas
from astropy.io import fits
from astropy.table import Table
from astropy.coordinates import SkyCoord
from astropath import bayesian
from astropath import localization
from astropath import path
# -
# ## LIGO example -- GW170817
lfile = os.path.join(resource_filename('astropath', 'data'), 'gw_examples',
'GW170817_skymap.fits.gz')
gw170817 = hp.read_map(lfile)
header = fits.open(lfile)[1].header
header
hp.mollview(
gw170817,
coord=["C"],
title="GW170817",
#unit="mK",
#norm="hist",
#min=-1,
#max=1,
)
hp.graticule()
# ## Galaxies
# https://vizier.u-strasbg.fr/viz-bin/VizieR?-source=VII/275
galfile = os.path.join(resource_filename('astropath', 'data'), 'gw_examples',
'GW170817_galaxies.csv')
cut_galaxies = pandas.read_csv(galfile, index_col=0)
cut_galaxies.head()
# ### Coordinates
cut_gal_coord = SkyCoord(ra=cut_galaxies.RAJ2000, dec=cut_galaxies.DEJ2000, unit='deg')
ngc_4993 = SkyCoord('13h09m47.706s -23d23m01.79s', frame='icrs')
np.min(ngc_4993.separation(cut_gal_coord).to('arcmin'))
np.argmin(ngc_4993.separation(cut_gal_coord))
cut_galaxies.iloc[11]
# ## PATH time
# ### Instantiate
Path = path.PATH()
# ### Candidates
Path.init_candidates(cut_galaxies.RAJ2000.values,
cut_galaxies.DEJ2000.values,
cut_galaxies.maj.values,
mag=cut_galaxies.Bmag.values)
# ### Priors
# Candidates
Path.init_cand_prior('inverse', P_U=0.)
# Offsets
Path.init_theta_prior('exp', 6.)
# ### Localization
Path.init_localization('healpix',
healpix_data=gw170817,
healpix_nside=header['NSIDE'],
healpix_ordering='NESTED',
healpix_coord='C')
# ### Calculate Priors
P_O = Path.calc_priors()
# ## Calculate Posteriors
P_Ox, P_Ux = Path.calc_posteriors('local', box_hwidth=30.)
Path.candidates.sort_values('P_Ox', ascending=False)
# ## F'ing A
# ----
# ## Multi order
# https://emfollow.docs.ligo.org/userguide/tutorial/multiorder_skymaps.html
mofile = os.path.join(resource_filename('astropath', 'data'), 'gw_examples',
'GW190814_PublicationSamples.multiorder.fits')
# ----
| docs/nb/GW_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computer Vision, Lab 6: Two-View Reconstruction
#
# Today we'll take a look at how to perform 3D reconstruction of a scene using point correspondences between two calibrated views of that scene.
#
# We'll explore keypoint detection and matching, estimating the essential matrix, estimating the camera rotation and translation, and resolving the scale ambiguity using extrinsic camera parameters.
#
# ## Data needed for the lab
#
# Grab the two home robot navigation videos we've already worked with. You may also want to just use the sequence of frames
# we segmented from. You'll also need the intrinsic parameters of the camera you obtained last time in order to undistort images from
# the camera.
# - [Video 1 (nighttime, LED lighting)](https://drive.google.com/file/d/1K2EjcMJifDUOkSP_amlg8wcHmv_jh44V/view?usp=sharing)
# - [Video 2 (daytime, ambient lighting)](https://drive.google.com/file/d/1LKH5zPhZRPKSHF287apsaOL5ZMN3c7JB/view?usp=sharing)
# - [Segmented frames from video 1](https://drive.google.com/drive/folders/1V0GyVhnrO9NgXLRzJNFVpLOqFJJScUg2?usp=sharing)
# - [Calibration images for the camera used for all video/image data](https://github.com/dsai-asia/CV/tree/master/Labs/05-Calibration/sample-calib-images-jetson-rpicam)
#
# ## Feature matching: AKAZE vs. ORB?
#
# We'll look at two feature point matchers today. They are both similar to the original idea
# of wide baseline matching with SIFT, first invented by <NAME> at the University of British Colombia around 2000.
# SIFT (and its faster successor SURF) are free for academic or individual use, but they are patent protected, so you have
# to license the algorithms if you want to make money with them! For that reason, the OpenCV community has implemented quite
# a few other feature point detectors and matchers so you have wide range of choices that are patent-free. We'll look at
# AKAZE and ORB a bit.
#
# Reference: [Comparing ORB and AKAZE for visual odometry
# of unmanned aerial vehicles](http://www.epacis.net/ccis2016/papers/paper_121.pdf)
#
# In **ORB**, the detection step is based on the FAST keypoint detector,
# which is an efficient corner detector suitable for real-time applications due
# to its computation properties. Since FAST does not include an orientation
# operator, ORB adds an orientation component to it, which
# is called oFAST (oriented FAST).
#
# **AKAZE** makes use of a "Fast Explicit Diffusion" (FED) scheme embedded in a pyramidal framework in order to build an accelerated feature detection system in nonlinear scale spaces. By means of FED schemes, a nonlinear scale space can be built much faster than with any other kind of discretization scheme.
#
# ## Keypoint detection and matching
#
# Study the [ORB/AKAZE OpenCV tutorial](https://docs.opencv.org/4.3.0/dc/d16/tutorial_akaze_tracking.html).
# It shows us how to do the following:
#
# - Detect and describe keypoints on the first frame, manually set object boundaries
# - For every next frame:
# 1. Detect and describe keypoints
# 2. Match them using bruteforce matcher
# 3. Estimate homography transformation using RANSAC
# 4. Filter out the outliers among the matches
# 5. Apply homography transformation to the bounding box to find the object
# 6. Draw bounding box and inliers and compute the inlier ratio as an evaluation metric
#
# While this is useful for tracking a 2D planar object with a fixed camera, the keypoint
# matching method is appropriate for full 3D point correspondence estimation, estimation of F or E,
# and so on. We'll just have to replace the homography transformation with F or E.
#
# First we'll talk about feature matching a bit, get the tutorial code running (code is replicated below and nicely translated
# into Python by Alisa), then we'll
# get AKAZE and ORB keypoints from the first two frames with motion in the sequence of frames from Video 1.
#
# When you adapt the tutorial code to our situation,
# note that the it has some things such as setting the ROI and tracking from a video that are not relevant for us.
# Focus on the keypoint detector setup and keypoint matcher setup.
#
# ## C++ / main.cpp
#include <opencv2/opencv.hpp>
#include <vector>
#include <iostream>
#include <iomanip>
#include "stats.h" // Stats structure definition
#include "utils.h" // Drawing and printing functions
using namespace std;
using namespace cv;
const double akaze_thresh = 3e-4; // AKAZE detection threshold set to locate about 1000 keypoints
const double ransac_thresh = 2.5f; // RANSAC inlier threshold
const double nn_match_ratio = 0.8f; // Nearest-neighbour matching ratio
const int bb_min_inliers = 100; // Minimal number of inliers to draw bounding box
const int stats_update_period = 10; // On-screen statistics are updated every 10 frames
namespace example {
class Tracker
{
public:
Tracker(Ptr<Feature2D> _detector, Ptr<DescriptorMatcher> _matcher) :
detector(_detector),
matcher(_matcher)
{}
void setFirstFrame(const Mat frame, vector<Point2f> bb, string title, Stats& stats);
Mat process(const Mat frame, Stats& stats);
Ptr<Feature2D> getDetector() {
return detector;
}
protected:
Ptr<Feature2D> detector;
Ptr<DescriptorMatcher> matcher;
Mat first_frame, first_desc;
vector<KeyPoint> first_kp;
vector<Point2f> object_bb;
};
void Tracker::setFirstFrame(const Mat frame, vector<Point2f> bb, string title, Stats& stats)
{
cv::Point* ptMask = new cv::Point[bb.size()];
const Point* ptContain = { &ptMask[0] };
int iSize = static_cast<int>(bb.size());
for (size_t i = 0; i < bb.size(); i++) {
ptMask[i].x = static_cast<int>(bb[i].x);
ptMask[i].y = static_cast<int>(bb[i].y);
}
first_frame = frame.clone();
cv::Mat matMask = cv::Mat::zeros(frame.size(), CV_8UC1);
cv::fillPoly(matMask, &ptContain, &iSize, 1, cv::Scalar::all(255));
detector->detectAndCompute(first_frame, matMask, first_kp, first_desc);
Mat res;
drawKeypoints(first_frame, first_kp, res, Scalar(255, 0, 0), DrawMatchesFlags::DRAW_RICH_KEYPOINTS);
imshow("key points", res);
waitKey(0);
destroyWindow("key points");
stats.keypoints = (int)first_kp.size();
drawBoundingBox(first_frame, bb);
putText(first_frame, title, Point(0, 60), FONT_HERSHEY_PLAIN, 5, Scalar::all(0), 4);
object_bb = bb;
delete[] ptMask;
}
Mat Tracker::process(const Mat frame, Stats& stats)
{
TickMeter tm;
vector<KeyPoint> kp;
Mat desc;
tm.start();
detector->detectAndCompute(frame, noArray(), kp, desc);
stats.keypoints = (int)kp.size();
vector< vector<DMatch> > matches;
vector<KeyPoint> matched1, matched2;
matcher->knnMatch(first_desc, desc, matches, 2);
for (unsigned i = 0; i < matches.size(); i++) {
if (matches[i][0].distance < nn_match_ratio * matches[i][1].distance) {
matched1.push_back(first_kp[matches[i][0].queryIdx]);
matched2.push_back(kp[matches[i][0].trainIdx]);
}
}
stats.matches = (int)matched1.size();
Mat inlier_mask, homography;
vector<KeyPoint> inliers1, inliers2;
vector<DMatch> inlier_matches;
if (matched1.size() >= 4) {
homography = findHomography(Points(matched1), Points(matched2),
RANSAC, ransac_thresh, inlier_mask);
}
tm.stop();
stats.fps = 1. / tm.getTimeSec();
if (matched1.size() < 4 || homography.empty()) {
Mat res;
hconcat(first_frame, frame, res);
stats.inliers = 0;
stats.ratio = 0;
return res;
}
for (unsigned i = 0; i < matched1.size(); i++) {
if (inlier_mask.at<uchar>(i)) {
int new_i = static_cast<int>(inliers1.size());
inliers1.push_back(matched1[i]);
inliers2.push_back(matched2[i]);
inlier_matches.push_back(DMatch(new_i, new_i, 0));
}
}
stats.inliers = (int)inliers1.size();
stats.ratio = stats.inliers * 1.0 / stats.matches;
vector<Point2f> new_bb;
perspectiveTransform(object_bb, new_bb, homography);
Mat frame_with_bb = frame.clone();
if (stats.inliers >= bb_min_inliers) {
drawBoundingBox(frame_with_bb, new_bb);
}
Mat res;
drawMatches(first_frame, inliers1, frame_with_bb, inliers2,
inlier_matches, res,
Scalar(255, 0, 0), Scalar(255, 0, 0));
return res;
}
}
int main(int argc, char** argv)
{
string video_name = "robot.mp4";
VideoCapture video_in;
video_in.open(video_name);
if (!video_in.isOpened()) {
cerr << "Couldn't open " << video_name << endl;
return 1;
}
Stats stats, akaze_stats, orb_stats;
Ptr<AKAZE> akaze = AKAZE::create();
akaze->setThreshold(akaze_thresh);
Ptr<ORB> orb = ORB::create();
Ptr<DescriptorMatcher> matcher = DescriptorMatcher::create("BruteForce-Hamming");
example::Tracker akaze_tracker(akaze, matcher);
example::Tracker orb_tracker(orb, matcher);
Mat frame;
namedWindow(video_name, WINDOW_NORMAL);
cout << "\nPress any key to stop the video and select a bounding box" << endl;
while (waitKey(1) < 1)
{
video_in >> frame;
cv::resizeWindow(video_name, frame.size());
imshow(video_name, frame);
}
vector<Point2f> bb;
cv::Rect uBox = cv::selectROI(video_name, frame);
bb.push_back(cv::Point2f(static_cast<float>(uBox.x), static_cast<float>(uBox.y)));
bb.push_back(cv::Point2f(static_cast<float>(uBox.x + uBox.width), static_cast<float>(uBox.y)));
bb.push_back(cv::Point2f(static_cast<float>(uBox.x + uBox.width), static_cast<float>(uBox.y + uBox.height)));
bb.push_back(cv::Point2f(static_cast<float>(uBox.x), static_cast<float>(uBox.y + uBox.height)));
akaze_tracker.setFirstFrame(frame, bb, "AKAZE", stats);
orb_tracker.setFirstFrame(frame, bb, "ORB", stats);
Stats akaze_draw_stats, orb_draw_stats;
Mat akaze_res, orb_res, res_frame;
int i = 0;
for (;;) {
i++;
bool update_stats = (i % stats_update_period == 0);
video_in >> frame;
// stop the program if no more images
if (frame.empty()) break;
akaze_res = akaze_tracker.process(frame, stats);
akaze_stats += stats;
if (update_stats) {
akaze_draw_stats = stats;
}
orb->setMaxFeatures(stats.keypoints);
orb_res = orb_tracker.process(frame, stats);
orb_stats += stats;
if (update_stats) {
orb_draw_stats = stats;
}
drawStatistics(akaze_res, akaze_draw_stats);
drawStatistics(orb_res, orb_draw_stats);
vconcat(akaze_res, orb_res, res_frame);
cv::imshow(video_name, res_frame);
if (waitKey(1) == 27) break; //quit on ESC button
}
akaze_stats /= i - 1;
orb_stats /= i - 1;
printStatistics("AKAZE", akaze_stats);
printStatistics("ORB", orb_stats);
return 0;
}
# ## C++ / stats.h
# +
#ifndef STATS_H
#define STATS_H
struct Stats
{
int matches;
int inliers;
double ratio;
int keypoints;
double fps;
Stats() : matches(0),
inliers(0),
ratio(0),
keypoints(0),
fps(0.)
{}
Stats& operator+=(const Stats& op) {
matches += op.matches;
inliers += op.inliers;
ratio += op.ratio;
keypoints += op.keypoints;
fps += op.fps;
return *this;
}
Stats& operator/=(int num)
{
matches /= num;
inliers /= num;
ratio /= num;
keypoints /= num;
fps /= num;
return *this;
}
};
#endif // STATS_H#pragma once
# -
# ## C++ / utils.h
# +
#ifndef UTILS_H
#define UTILS_H
#include <opencv2/opencv.hpp>
#include <vector>
#include "stats.h"
using namespace std;
using namespace cv;
void drawBoundingBox(Mat image, vector<Point2f> bb);
void drawStatistics(Mat image, const Stats& stats);
void printStatistics(string name, Stats stats);
vector<Point2f> Points(vector<KeyPoint> keypoints);
Rect2d selectROI(const String& video_name, const Mat& frame);
void drawBoundingBox(Mat image, vector<Point2f> bb)
{
for (unsigned i = 0; i < bb.size() - 1; i++) {
line(image, bb[i], bb[i + 1], Scalar(0, 0, 255), 2);
}
line(image, bb[bb.size() - 1], bb[0], Scalar(0, 0, 255), 2);
}
void drawStatistics(Mat image, const Stats& stats)
{
static const int font = FONT_HERSHEY_PLAIN;
stringstream str1, str2, str3, str4;
str1 << "Matches: " << stats.matches;
str2 << "Inliers: " << stats.inliers;
str3 << "Inlier ratio: " << setprecision(2) << stats.ratio;
str4 << "FPS: " << std::fixed << setprecision(2) << stats.fps;
putText(image, str1.str(), Point(0, image.rows - 120), font, 2, Scalar::all(255), 3);
putText(image, str2.str(), Point(0, image.rows - 90), font, 2, Scalar::all(255), 3);
putText(image, str3.str(), Point(0, image.rows - 60), font, 2, Scalar::all(255), 3);
putText(image, str4.str(), Point(0, image.rows - 30), font, 2, Scalar::all(255), 3);
}
void printStatistics(string name, Stats stats)
{
cout << name << endl;
cout << "----------" << endl;
cout << "Matches " << stats.matches << endl;
cout << "Inliers " << stats.inliers << endl;
cout << "Inlier ratio " << setprecision(2) << stats.ratio << endl;
cout << "Keypoints " << stats.keypoints << endl;
cout << "FPS " << std::fixed << setprecision(2) << stats.fps << endl;
cout << endl;
}
vector<Point2f> Points(vector<KeyPoint> keypoints)
{
vector<Point2f> res;
for (unsigned i = 0; i < keypoints.size(); i++) {
res.push_back(keypoints[i].pt);
}
return res;
}
#endif // UTILS_H#pragma once
# -
# ## Python / stats.py
#
# Some quick tips:
# - Use a multiline comment (""" some data """) after a class namde or function name declaration to make your intellisense checker happy.
# - use <tt>:type</tt> to define the type of a parameter to a function or method.
# - You can overload operators like <tt>+</tt>, <tt>-</tt>, <tt>*</tt>, and <tt>/</tt> yourself. Just declare a method with <tt>__method__(self,...)</tt>.
# Try it, it's very useful!
# +
import numpy as np
class Stats:
"""
Statistic class
Attributes
----------
matches=0 (int):
total number of matching
inliers=0 (int):
number of inliner matching
ratio=0. (float):
Nearest-neighbour matching ratio
keypoints=0 (int):
Wall
fps=0. (float):
frame per 1 sec
Methods
-------
add(Stats) - overload + function:
plus the information into this class
divide(Stats) - overload + function:
divide the information into this class
"""
matches:int
inliers:int
ratio:float
keypoints:int
fps:float
def __init__(self, matches = 0, inliers = 0, ratio = 0., keypoints = 0, fps = 0.):
self.matches = matches
self.inliers = inliers
self.ratio = ratio
self.keypoints = keypoints
self.fps = fps
def __add__(self, op:"Stats") -> "Stats":
self.matches += op.matches
self.inliers += op.inliers
self.ratio += op.ratio
self.keypoints += op.keypoints
self.fps += op.fps
return self
def __truediv__(self, num:int) -> "Stats":
self.matches //= num
self.inliers //= num
self.ratio /= num
self.keypoints //= num
self.fps /= num
return self
def __str__(self) -> str:
return "matches({0}) inliner({1}) ratio({2:.2f}) keypoints({3}) fps({4:.2f})".format(self.matches, self.inliers, self.ratio, self.keypoints, self.fps)
__repr__ = __str__
def to_strings(self):
"""
Convert to string set of matches, inliners, ratio, and fps
"""
str1 = "Matches: {0}".format(self.matches)
str2 = "Inliers: {0}".format(self.inliers)
str3 = "Inlier ratio: {0:.2f}".format(self.ratio)
str4 = "Keypoints: {0}".format(self.keypoints)
str5 = "FPS: {0:.2f}".format(self.fps)
return str1, str2, str3, str4, str5
def copy(self):
return Stats(self.matches, self.inliers, self.ratio, self.keypoints, self.fps)
# +
# test the class
#from stats import Stats
test1 = Stats(5, 2, 9, 4, 1.5)
test2 = Stats(2, 1, 0, 8, 9)
test1 + test2
print(test1)
test1 / 3
print(test1)
# -
# ## Python / Utils.py
# +
from stats import Stats
import cv2
from typing import List #use it for :List[...]
def drawBoundingBox(image, bb):
"""
Draw the bounding box from the points set
Parameters
----------
image (array):
image which you want to draw
bb (List):
points array set
"""
color = (0, 0, 255)
for i in range(len(bb) - 1):
b1 = (int(bb[i][0]), int(bb[i][1]))
b2 = (int(bb[i + 1][0]), int(bb[i + 1][1]))
cv2.line(image, b1, b2, color, 2)
b1 = (int(bb[len(bb) - 1][0]), int(bb[len(bb) - 1][1]))
b2 = (int(bb[0][0]), int(bb[0][1]))
cv2.line(image, b1, b2, color, 2)
def drawStatistics(image, stat: Stats):
"""
Draw the statistic to images
Parameters
----------
image (array):
image which you want to draw
stat (Stats):
statistic values
"""
font = cv2.FONT_HERSHEY_PLAIN
str1, str2, str3, str4, str5 = stat.to_strings()
shape = image.shape
cv2.putText(image, str1, (0, shape[0] - 120), font, 2, (0, 0, 255), 3)
cv2.putText(image, str2, (0, shape[0] - 90), font, 2, (0, 0, 255), 3)
cv2.putText(image, str3, (0, shape[0] - 60), font, 2, (0, 0, 255), 3)
cv2.putText(image, str5, (0, shape[0] - 30), font, 2, (0, 0, 255), 3)
def printStatistics(name: str, stat: Stats):
"""
Print the statistic
Parameters
----------
name (str):
image which you want to draw
stat (Stats):
statistic values
"""
print(name)
print("----------")
str1, str2, str3, str4, str5 = stat.to_strings()
print(str1)
print(str2)
print(str3)
print(str4)
print(str5)
print()
def Points(keypoints):
res = []
for i in keypoints:
res.append(i)
return res
# -
# ## Python / main.py
# +
import cv2
import numpy as np
import time
from stats import Stats
from utils import drawBoundingBox, drawStatistics, printStatistics, Points
akaze_thresh:float = 3e-4 # AKAZE detection threshold set to locate about 1000 keypoints
ransac_thresh:float = 2.5 # RANSAC inlier threshold
nn_match_ratio:float = 0.8 # Nearest-neighbour matching ratio
bb_min_inliers:int = 100 # Minimal number of inliers to draw bounding box
stats_update_period:int = 10 # On-screen statistics are updated every 10 frames
class Tracker:
def __init__(self, detector, matcher):
self.detector = detector
self.matcher = matcher
def setFirstFrame(self, frame, bb, title:str):
iSize = len(bb)
stat = Stats()
ptContain = np.zeros((iSize, 2))
i = 0
for b in bb:
#ptMask[i] = (b[0], b[1])
ptContain[i, 0] = b[0]
ptContain[i, 1] = b[1]
i += 1
self.first_frame = frame.copy()
matMask = np.zeros(frame.shape, dtype=np.uint8)
cv2.fillPoly(matMask, np.int32([ptContain]), (255,0,0))
# cannot use in ORB
# self.first_kp, self.first_desc = self.detector.detectAndCompute(self.first_frame, matMask)
# find the keypoints with ORB
kp = self.detector.detect(self.first_frame,None)
# compute the descriptors with ORB
self.first_kp, self.first_desc = self.detector.compute(self.first_frame, kp)
# print(self.first_kp[0].pt[0])
# print(self.first_kp[0].pt[1])
# print(self.first_kp[0].angle)
# print(self.first_kp[0].size)
res = cv2.drawKeypoints(self.first_frame, self.first_kp, None, color=(255,0,0), flags=cv2.DRAW_MATCHES_FLAGS_DRAW_RICH_KEYPOINTS)
stat.keypoints = len(self.first_kp)
drawBoundingBox(self.first_frame, bb);
cv2.imshow("key points of {0}".format(title), res)
cv2.waitKey(0)
cv2.destroyWindow("key points of {0}".format(title))
cv2.putText(self.first_frame, title, (0, 60), cv2.FONT_HERSHEY_PLAIN, 5, (0,0,0), 4)
self.object_bb = bb
return stat
def process(self, frame):
stat = Stats()
start_time = time.time()
kp, desc = self.detector.detectAndCompute(frame, None)
stat.keypoints = len(kp)
matches = self.matcher.knnMatch(self.first_desc, desc, k=2)
matched1 = []
matched2 = []
matched1_keypoints = []
matched2_keypoints = []
good = []
for i,(m,n) in enumerate(matches):
if m.distance < nn_match_ratio * n.distance:
good.append(m)
matched1_keypoints.append(self.first_kp[matches[i][0].queryIdx])
matched2_keypoints.append(kp[matches[i][0].trainIdx])
matched1 = np.float32([ self.first_kp[m.queryIdx].pt for m in good ]).reshape(-1,1,2)
matched2 = np.float32([ kp[m.trainIdx].pt for m in good ]).reshape(-1,1,2)
stat.matches = len(matched1)
homography = None
if (len(matched1) >= 4):
homography, inlier_mask = cv2.findHomography(matched1, matched2, cv2.RANSAC, ransac_thresh)
dt = time.time() - start_time
stat.fps = 1. / dt
if (len(matched1) < 4 or homography is None):
res = cv2.hconcat([self.first_frame, frame])
stat.inliers = 0
stat.ratio = 0
return res, stat
inliers1 = []
inliers2 = []
inliers1_keypoints = []
inliers2_keypoints = []
for i in range(len(good)):
if (inlier_mask[i] > 0):
new_i = len(inliers1)
inliers1.append(matched1[i])
inliers2.append(matched2[i])
inliers1_keypoints.append(matched1_keypoints[i])
inliers2_keypoints.append(matched2_keypoints[i])
inlier_matches = [cv2.DMatch(_imgIdx=0, _queryIdx=idx, _trainIdx=idx,_distance=0) for idx in range(len(inliers1))]
inliers1 = np.array(inliers1, dtype=np.float32)
inliers2 = np.array(inliers2, dtype=np.float32)
stat.inliers = len(inliers1)
stat.ratio = stat.inliers * 1.0 / stat.matches
bb = np.array([self.object_bb], dtype=np.float32)
new_bb = cv2.perspectiveTransform(bb, homography)
frame_with_bb = frame.copy()
if (stat.inliers >= bb_min_inliers):
drawBoundingBox(frame_with_bb, new_bb[0])
res = cv2.drawMatches(self.first_frame, inliers1_keypoints, frame_with_bb, inliers2_keypoints, inlier_matches, None, matchColor=(255, 0, 0), singlePointColor=(255, 0, 0))
return res, stat
def getDetector(self):
return self.detector
def main():
video_name = "robot.mp4"
video_in = cv2.VideoCapture()
video_in.open(video_name)
if (not video_in.isOpened()):
print("Couldn't open ", video_name)
return -1
akaze_stats = Stats()
orb_stats = Stats()
akaze = cv2.AKAZE_create()
akaze.setThreshold(akaze_thresh)
orb = cv2.ORB_create()
matcher = cv2.DescriptorMatcher_create("BruteForce-Hamming")
akaze_tracker = Tracker(akaze, matcher)
orb_tracker = Tracker(orb, matcher)
cv2.namedWindow(video_name, cv2.WINDOW_NORMAL);
print("\nPress any key to stop the video and select a bounding box")
key = -1
while(key < 1):
_, frame = video_in.read()
w, h, ch = frame.shape
cv2.resizeWindow(video_name, (h, w))
cv2.imshow(video_name, frame)
key = cv2.waitKey(1)
print("Select a ROI and then press SPACE or ENTER button!")
print("Cancel the selection process by pressing c button!")
uBox = cv2.selectROI(video_name, frame);
bb = []
bb.append((uBox[0], uBox[1]))
bb.append((uBox[0] + uBox[2], uBox[0] ))
bb.append((uBox[0] + uBox[2], uBox[0] + uBox[3]))
bb.append((uBox[0], uBox[0] + uBox[3]))
stat_a = akaze_tracker.setFirstFrame(frame, bb, "AKAZE",);
stat_o = orb_tracker.setFirstFrame(frame, bb, "ORB");
akaze_draw_stats = stat_a.copy()
orb_draw_stats = stat_o.copy()
i = 0
video_in.set(cv2.CAP_PROP_POS_FRAMES, 0)
while True:
i += 1
update_stats = (i % stats_update_period == 0)
_, frame = video_in.read()
if frame is None:
# End of video
break
akaze_res, stat = akaze_tracker.process(frame)
akaze_stats + stat
if (update_stats):
akaze_draw_stats = stat
orb.setMaxFeatures(stat.keypoints)
orb_res, stat = orb_tracker.process(frame)
orb_stats + stat
if (update_stats):
orb_draw_stats = stat
drawStatistics(akaze_res, akaze_draw_stats)
drawStatistics(orb_res, orb_draw_stats)
res_frame = cv2.vconcat([akaze_res, orb_res])
# cv2.imshow(video_name, akaze_res)
cv2.imshow(video_name, res_frame)
if (cv2.waitKey(1) == 27): # quit on ESC button
break
akaze_stats / (i - 1)
orb_stats / (i - 1)
printStatistics("AKAZE", akaze_stats);
printStatistics("ORB", orb_stats);
return 0
main()
# -
# ## Exercises
#
# ### ORB/AKAZE Tutorial
#
# Get the tutorial running and play with it.
#
# ### Feature points
#
# Select a pair of frames with motion from the Video 1 frame sequence.
#
# Detect ORB and AKAZE features and use
# the OpenCV [<code>drawKeypoints()</code>](https://docs.opencv.org/4.3.0/d4/d5d/group__features2d__draw.html#ga5d2bafe8c1c45289bc3403a40fb88920) function to display the keypoints detected in the two images. Your result should look something like this:
#
# <img src="img/lab06-1.png" width="600"/>
#
# ### Undistortion
#
# Using the parameters you got and saved in Lab 05,
# use <tt>undistortPoints()</tt> to obtain "ideal" undistorted points for each of the input point sets.
#
# Be careful about the Mat object resulting from <code>undistortPoints()</code>. It is a Nx1 2 channel, 64-bit image, so to access it, you use code such as
# (C++):
#
# // Example use of undistortPoints function
#
# Mat xy_undistorted; // leave empty, opencv will fill it.
# undistortPoints(match_points, xy_undistorted, camera_matrix, dist_coeffs);
#
# Point2f point;
# for (int i = 0;i<nPoints;i++)
# {
# point.x = xy_undistorted.at<cv::Vec2d>(i, 0)[0];
# point.y = xy_undistorted.at<cv::Vec2d>(i, 0)[1];
# // do something
# }
#
# It's easier in Python:
#
# xy_undistorted = cv2.undistortPoints(match_points, camera_matrix, dist_coeffs)
#
# x = xy_undistorted[i][0]
# y = xy_undistorted[i][1]
#
# Knowing this in advance will save you some time.
#
# ### Feature point matching
#
# Next, get matches using the brute force Hamming matcher, remove indistinct matches (matches for which the ratio of distances for the first and second match is greater than 0.8) and use the OpenCV [<code>drawMatches()</code>](https://docs.opencv.org/4.3.0/d4/d5d/group__features2d__draw.html#gad8f463ccaf0dc6f61083abd8717c261a) function to display the result for AKAZE and ORB.
#
# In your report, discuss which keypoint detector seems to work best in terms of number of matches and number of accurate matches.
#
# ### Essential matrix
#
# Next, let's find an essential matrix relating these two images using the better keypoint matching algorithm from the previous experiment.
# Use <code>findEssentialMat</code> to get an essential matrix with RANSAC.
# Check carefully about normalization of the point correspondences.
# After that, replot your correspondences with inliers only, obtaining something like the following:
#
# <img src="img/lab06-2.png" width="600"/>
#
# Pick two pairs of corresponding points in the two images and verify that $X^T K^{-T} E K^{-1} X' = 0$, approximately.
#
# Hint: you can tell <code>drawMatches</code> to only draw inliers by constructing a vector of vector of char like this:
#
# std::vector<std::vector<char> > vvMatchesMask;
# for (int i = 0, j = 0; i < matched1.size(); i++) {
# if (vMatched[i]) {
# if (inlier_mask.at<uchar>(j)) {
# vvMatchesMask.push_back( { 1, 0 } );
# } else {
# vvMatchesMask.push_back( { 0, 0 });
# }
# j++;
# } else {
# vvMatchesMask.push_back( { 0, 0 });
# }
# }
#
# Here's the Python:
#
# matchesMask = []
# j = 0
# for i in range(len(good)):
# if vMatched[i]:
# if inlier_mask[j] > 0:
# matchesMask.append( ( 1, 0 ) )
# else:
# matchesMask.append( ( 0, 0 ) )
# j += 1
# else:
# matchesMask.append( ( 0, 0 ))
#
# Here <code>vMatched</code> is a vector of <code>bool</code> that I constructed while selecting matches according to the distance ratio.
#
# Using undistorted images and undistorted points (see note above about how to access the undistorted point array) you should get something like this:
#
# <img src="img/lab06-3.png" width="600"/>
#
# ### Epipolar lines
#
# Finally, draw a couple corresponding epipolar lines in each undistorted image. You should get something like this:
#
# For frame 1:
#
# <img src="img/lab06-4.png" width="600"/>
#
# For frame 2:
#
# <img src="img/lab06-5.png" width="600"/>
#
# Next, perform factorization of E to get R and t.
#
# In your report, show your analysis of the number of keypoints, matched keypoints, matched unique keypoints (those that pass the distance ratio test), and inliers according to the estimated essential matrix.
#
# ### Recover relative pose
#
# Use <code>correctMatches()</code> and <code>recoverPose()</code> to "clean up" your image points (adjust each corresponding pair of points to be on corresponding epipolar lines according to E/F) and get the rotation and translation between the two camera frames. Understand the rotation and translation vectors you get and the scale ambiguity inherent in a metric 3D reconstruction.
#
# Construct the two projection matrices and use <code>triangulatePoints()</code> to obtain 3D points from the corrected 2D points. Visualize the 3D point cloud in Octave to see if it is sensible.
#
# You should get something similar to this:
#
# <img src="img/lab06-6.png" width="600"/>
#
# Here the points have been transformed from the first camera's coordinate frame to the robot frame for the first camera, using the rotation matrix and translation matrix from the extrinsic calibration.
#
# ### Find absolute scale
#
# We know that after scaling then transforming the 3D points into the world coordinate system, the points with the smallest 'Z' values should be the ones on the floor. Can you come up with a scale factor that pushes the "bottom" of the point cloud to the floor (Z=0) in the world frame?
# For that you'll need the extrinsic parameters of the camera. We'll provide them.
# Show your solution and a visualization of the points.
#
# After scaling the points in the camera frame (or re-triangulating after scaling the translation vector from <code>recoverPose()</code>), you should have a structure similar to what's shown in [this video](https://drive.google.com/file/d/16lwooQ4rIGJJ1cLM-hUxb_m-tmmyWddY/view).
| Labs/06-Two-View-Reconstruction/06-Two-View-Reconstruction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import GraphQL Client
from os.path import expanduser
from gql import gql, Client
from gql.transport.requests import RequestsHTTPTransport
with open(expanduser('~/Desktop/.api/yelp_apikey.txt')) as f: # Grab API Key
api_key = f.readline().strip()
# -
# # Set Params
# +
# build the request framework
transport = RequestsHTTPTransport(url='https://api.yelp.com/v3/graphql',
headers={'Authorization': 'bearer {}'.format(api_key),
'Content-Type': 'application/json'},
use_json=True)
client = Client(transport=transport, fetch_schema_from_transport=True) # Create the client
# +
query = """
{
business(id: "mission-cliffs-climbing-and-fitness-san-francisco") {
name
id
alias
reviews {
user {
name
id
}
id
rating
text
time_created
}
}
}
"""
response_query = client.execute(gql(query)) # Get the response
# -
# print the query
print(response_query)
| practice/yelp-graphql-api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="NVGjeOv_TZ7j"
# # Salad Documentation
#
# Group 22: <NAME>, <NAME>, <NAME>, <NAME>
#
# December, 2021
#
# ---
# + [markdown] id="vEPjEB1VTWIW"
# ### Table of Contents
#
# * [Introduction](#c1)
# * [Background](#c2)
# * [Computational Graph](#c21)
# * [Chain Rule](#c22)
# * [Forward Mode](#c23)
# * [Gradient & Jacobian](#c24)
# * [Dual Numbers](#c25)
# * [Software Organization](#c4)
# * [Directory Structure](#c41)
# * [Modules](#c42)
# * [Test Suite](#c43)
# * [Packaging and Distribution](#c44)
# * [How to Use Salad](#c3)
# * [Installation](#c31)
# * [Methods](#c32)
# * [1 input to 1 output](#c321)
# * [m inputs to 1 output](#c322)
# * [m inputs to n outputs](#s1)
# * [Implementation](#c5)
# * [Data Structure](#c51)
# * [Classes and Methods](#c52)
# * [Variable](#c521)
# * [Forward](#c522)
# * [Extensions](#c6)
# * [Gradient Descent (GD)](#c61)
# * [Broyden–Fletcher–Goldfarb–Shanno (BFGS)](#c62)
# * [Stochastic Gradient Descent (SGD)](#c63)
# * [Broader Impact](#c7)
# * [Software Inclusivity](#c8)
# * [Licensing](#c9)
# * [Future Directions](#c10)
# + [markdown] id="LnjCm3B52bhI"
#
#
# ## <a name='c1'></a>1 Introduction
#
# Differentiation is a central operation in science and engineering, used in various settings from financial markets to physics labs. For example, to optimize a function we must find the minima or maxima by using differentiation tests. There are multiple computational mechanisms to find derivatives. For example, numerical approximations like the Finite-Differences method involve utilizing the definition of a derivative and plugging in small values of h to evaluate the function. However, this method suffers from poor accuracy and stability issues due to floating point errors. Symbolic computation is another method, more accurate than numerical approximations, but often not applicable due to its unweildly and complex overhead. Here, we focus on automatic differentiation (AD), which overcomes the drawbacks of these two issues: it has less overhead than symbolic differentiation while calculating derivatives at machine precision. This makes it an integral part of machine learning algorithms, which demand computational efficiency as well as high precision.
# + [markdown] id="-a2bzELoW9QM"
# ## <a name='c2'></a>2 Background
#
# Automatic differentiation can be approached in two ways, known as $\textit{forward mode}$ and $\textit{ reverse mode}$. Both methods depend on the decomposition of a complex function into elementary operations. Then, we can take advantage of the chain rule to compute the derivatives of complex functions from trivially computed derivatives of these elementary functions.
# + [markdown] id="ESJMFwb9XM4k"
# ### <a name='c21'></a>2.1 Computational Graph
#
# The first step of AD is to construct a computational graph which relates the elementary inputs to the final function. Each node represents an intermediate computation $v_n$, and edges connect nodes via elementary functions $V_n$. For example, given a multivariate scalar function $f(x_1, x_2) = sin(x_1 + x_2)$, the computational graph looks like the following:
#
# <img src="https://github.com/cs107-DaLand/cs107-FinalProject/raw/final/docs/images/comp_graph.png" width="60%"/>
#
# Elementary functions include but are not limited to the following: algebraic operations, exponentials, trigonometrics, and their inverses. AD takes advantage of the fact that derivatives of elementary functions are trivial to compute, allowing the differentiation of complex functions via the chain rule.
# + [markdown] id="qzwAsGUiYENQ"
# ### <a name='c22'></a>2.2 Chain Rule
#
# Suppose we want to calculate $\frac{\partial f}{\partial x_1}$. From the computational graph, we can rewrite $f$ in terms of nested intermediate functions:
#
# \begin{equation*}
# f(x_1, x_2) = V_2(V_1(V_{-1}(x_1), V_0(x_2)))
# \end{equation*}
#
# where $V_n$ is the function that gives the value of node $v_n$. The chain rule states that
#
# \begin{equation*}
# \frac{\partial f}{\partial x_1} = \frac{\partial V_2}{\partial V_1} \frac{\partial V_1}{\partial V_{-1}} \frac{\partial V_{-1}}{\partial x_1}
# \end{equation*}
#
# For a multivariate function $f(g_1(x), g_2(x), ..., g_n(x))$, the chain rule generalizes to:
#
# \begin{equation*}
# \frac{\partial f}{\partial x} = \sum_{i = 1}^{n} \Big( \frac{\partial f}{\partial g_i} \frac{\partial g_i}{\partial x} \Big)
# \end{equation*}
# + [markdown] id="5Yw6dLhDYHO7"
# ### <a name='c23'></a>2.3 Forward Mode
#
# Forward mode, which is the conceptually simpler method, involves computing the chain rule from inside to outside. For the above example $f(x_1, x_2) = sin(x_1 + x_2)$, the $\textit{evaluation trace}$ at the point $(1, 2)$ is generated using the computational graph as a guide:
#
# <img src="https://github.com/cs107-DaLand/cs107-FinalProject/raw/final/docs/images/table1.png" width="60%"/>
#
# The evaluation trace essentially computes a value (primal trace) and derivative (tangent trace) for each node of the computational graph. The desired final results are $\frac{\partial f}{\partial x_1}$ and $\frac{\partial f}{\partial x_2}$, shown in the last row of the trace using the gradient symbol $\nabla$. Forward mode is advantageous when there are few inputs ($m$) and many outputs ($n$). The opposite is true for reverse mode, which motivates its use in deep learning applications with many inputs.
# + [markdown] id="uK_nq9QxYJXS"
# ### <a name='c24'></a>2.4 Gradient & Jacobian
#
# The gradient of a multivariate scalar-valued function $f(g_1, g_2, ..., g_m)$ gives all of its partial derivatives in a vector of size $m \times 1$:
#
# \begin{equation*}
# \nabla f = \nabla f_g = \begin{bmatrix}
# \frac{\partial f}{\partial g_1} \\
# \frac{\partial f}{\partial g_2} \\
# \vdots \\
# \frac{\partial f}{\partial g_m}
# \end{bmatrix}
# \end{equation*}
#
# If the inputs $g_i$ are themselves multivariate functions with inputs $x \in \mathbb{R}^m$, the chain rule gives
#
# \begin{equation*}
# \nabla_x f = \sum_{i = 1}^{n} \Bigg( \frac{\partial f}{\partial g_i} \nabla g_i (x) \Bigg)
# \end{equation*}
#
# Finally, we reach the most general form of the gradient: for multivariate vector-valued functions with $m$ inputs and $n$ outputs, the derivative of outputs with respect to inputs can be represented in a $\textit{Jacobian matrix}$ of dimensions $n \times m$:
#
# \begin{gather*}
# f(x_1, x_2, ..., x_m) = \begin{bmatrix} y_1 \\ y_2 \\ \vdots \\ y_n \end{bmatrix} \\
# J_f
# =
# \begin{bmatrix} \frac{\partial y_1}{\partial x_1} & \ldots & \frac{\partial y_1}{\partial x_m} \\
# \vdots & \ddots & \vdots \\
# \frac{\partial y_n}{\partial x_1} & \ldots & \frac{\partial y_n}{\partial x_m} \end{bmatrix}
# \end{gather*}
# + [markdown] id="Ph4qaGGtYLMc"
# ### <a name='c25'></a>2.5 Dual Numbers
#
# Dual numbers offer a convenient way to store and compute the primal and tangent traces in forward mode AD. A dual number has the form
#
# \begin{equation*}
# a + b \epsilon,
# \end{equation*}
#
# where $a, b \in \mathbb{R}$ and $\epsilon^2 = 0$. Combined with Taylor series expansion of a function $f$ centered at $a$, dual numbers give rise to the property
#
# \begin{equation*}
# f(a + b \epsilon) = f(a) + f' (a) b \epsilon
# \end{equation*}
#
# because all terms with powers of $\epsilon$ higher than $2$ are equal to zero. Thus, the Taylor series approximation of a function about a dual number gives both its value and derivative, in a single calculation. In forward mode AD it is convenient to represent both the primal and tangent trace of each node as a dual number, where the real part corresponds to the primal and the dual part corresponds to the tangent. As shown below, functions (operations) on these nodes preserve the relationship between the real and dual parts:
#
# \begin{equation*}
# g\big( f(a + \epsilon) \big) = g \big( f(a) + f' (a) \epsilon \big) = g \big( f(a) \big) + g' \big( f(a) \big) f'(a) \epsilon
# \end{equation*}
#
# Note that the coefficient of $\epsilon$ is simply the derivative of the real part $g(f(a))$, as given by the chain rule.
# + [markdown] id="E6REGhAf_Vwf"
# ## <a name='c4'></a>3 Software Organization
# + [markdown] id="SgYsWY4bZeF9"
# ### <a name='c41'></a>3.1 Directory Structure
#
# Our directory structure looks like the following:
#
# cs107-FinalProject/
# ├── LICENSE
# ├── README.md
# ├── pyproject.toml
# ├── requirements.txt
# ├── setup.py
# ├── docs/
# │ ├── images/
# │ │ ├── comp_graph.png
# │ │ └── table1.png
# │ ├── milestone1.pdf
# │ ├── milestone2 progress.pdf
# │ ├── milestone2.pdf
# │ ├── Documentation.ipynb
# │ └── Documentation.html
# ├── tests/
# │ ├── run_test.sh
# │ └── test_forward.py
# │ └── test_optimize.py
# └── src/
# └── cs107_salad/
# ├── __init__.py
# ├── Forward/
# │ ├── __init__.py
# │ ├── salad.py
# │ └── utils.py
# └── Optimization/
# ├── __init__.py
# └── optimize.py
#
# + [markdown] id="Sj8f8jsiZhH6"
# ### <a name='c42'></a>3.2 Modules
#
# Three external modules that we will rely on in our implementation will be NumPy, pytest, and coverage.
#
# # + NumPy offers support for large, multi-dimensional arrays as well a wide range of optimized mathematical functions and operators for these arrays. Numerical calculation of function values in Salad will rely extensively on this module.
# # + pytest is a feature rich testing framework in python that makes it easy to write unit tests that scale for full libraries.
# # + coverage is a commonly-used library used to compute code coverage during test execution.
# + [markdown] id="VEWwad3NZjQc"
# ### <a name='c3'></a>3.3 Test Suite
#
# The test suite will live inside the $\texttt{tests}$ directory.
#
# To run the test suite, find $\texttt{run_tests.sh}$ in the $\texttt{tests}$ folder. Run the following command to run test and generate coverage report.
#
# ```
# sh run_tests.sh
# ```
#
# Local test result:
#
# <table class="index" data-sortable="">
# <thead>
# <tr class="tablehead" title="Click to sort">
# <th class="name left" aria-sort="none" data-shortcut="n">Module</th>
# <th aria-sort="none" data-default-sort-order="descending" data-shortcut="s">statements</th>
# <th aria-sort="none" data-default-sort-order="descending" data-shortcut="m">missing</th>
# <th aria-sort="none" data-default-sort-order="descending" data-shortcut="x">excluded</th>
# <th class="right" aria-sort="none" data-shortcut="c">coverage</th>
# </tr>
# </thead>
# <tbody>
# <tr class="file">
# <td class="name left"><a href="d_86ea9ce4ec7f5104___init___py.html">/home/ddv/projects/CS107/final-project/cs107-FinalProject/src/cs107_salad/Forward/__init__.py</a></td>
# <td>0</td>
# <td>0</td>
# <td>0</td>
# <td class="right" data-ratio="0 0">100%</td>
# </tr>
# <tr class="file">
# <td class="name left"><a href="d_86ea9ce4ec7f5104_salad_py.html">/home/ddv/projects/CS107/final-project/cs107-FinalProject/src/cs107_salad/Forward/salad.py</a></td>
# <td>380</td>
# <td>31</td>
# <td>0</td>
# <td class="right" data-ratio="349 380">92%</td>
# </tr>
# <tr class="file">
# <td class="name left"><a href="d_86ea9ce4ec7f5104_utils_py.html">/home/ddv/projects/CS107/final-project/cs107-FinalProject/src/cs107_salad/Forward/utils.py</a></td>
# <td>28</td>
# <td>2</td>
# <td>0</td>
# <td class="right" data-ratio="26 28">93%</td>
# </tr>
# <tr class="file">
# <td class="name left"><a href="d_3e139fac06414ae6___init___py.html">/home/ddv/projects/CS107/final-project/cs107-FinalProject/src/cs107_salad/Optimization/__init__.py</a></td>
# <td>0</td>
# <td>0</td>
# <td>0</td>
# <td class="right" data-ratio="0 0">100%</td>
# </tr>
# <tr class="file">
# <td class="name left"><a href="d_3e139fac06414ae6_optimize_py.html">/home/ddv/projects/CS107/final-project/cs107-FinalProject/src/cs107_salad/Optimization/optimize.py</a></td>
# <td>128</td>
# <td>5</td>
# <td>0</td>
# <td class="right" data-ratio="123 128">96%</td>
# </tr>
# <tr class="file">
# <td class="name left"><a href="d_e085b66f4602d46f___init___py.html">/home/ddv/projects/CS107/final-project/cs107-FinalProject/src/cs107_salad/__init__.py</a></td>
# <td>0</td>
# <td>0</td>
# <td>0</td>
# <td class="right" data-ratio="0 0">100%</td>
# </tr>
# <tr class="file">
# <td class="name left"><a href="test_forward_py.html">test_forward.py</a></td>
# <td>984</td>
# <td>29</td>
# <td>0</td>
# <td class="right" data-ratio="955 984">97%</td>
# </tr>
# <tr class="file">
# <td class="name left"><a href="test_optimize_py.html">test_optimize.py</a></td>
# <td>95</td>
# <td>3</td>
# <td>0</td>
# <td class="right" data-ratio="92 95">97%</td>
# </tr>
# </tbody>
# <tfoot>
# <tr class="total">
# <td class="name left">Total</td>
# <td>1615</td>
# <td>70</td>
# <td>0</td>
# <td class="right" data-ratio="1545 1615">96%</td>
# </tr>
# </tfoot>
# </table>
#
# Test coverage is currently at 96%.
# + [markdown] id="SJjfqOAKZk9O"
# ### <a name='c44'></a>3.4 Packaging and Distribution
#
# We have distributed the package through the Test Python Package Index (PyPi). Users will be able to install our package just by doing:
#
# ```
# pip install cs107-salad --extra-index-url=https://test.pypi.org/simple/
# ```
#
# Because the package is on test PyPi, the extra index url flag allows numpy, one of package dependecies, to be installed from main PyPi instead of test PyPi. Test PyPi does not have the most up to date version of numpy.
#
# Eventually, when the package has even more features, we would aim to distribute the package on the main PyPi.
#
# + [markdown] id="QBBOyusK2x6U"
# ## 4 <a name='c3'></a>How to Use $\texttt{Salad}$
# + [markdown] id="i2ggHLcpYo2d"
# ### <a name='c31'></a>4.1 Installation
#
# The users will pull $\texttt{Salad}$ from PyPI using command line
# + id="9X-6wg0v2RtM" colab={"base_uri": "https://localhost:8080/"} outputId="8d1c880b-d6b8-4c79-c899-6ffc479a5551"
# !pip install cs107-salad --extra-index-url=https://test.pypi.org/simple/
# + [markdown] id="styvKsxF9V0r"
# Then import $\texttt{salad}$ and other dependencies (e.g. $\texttt{numpy})$ in python script
# + id="gH7-zmZq9TVD"
import cs107_salad.Forward.salad as ad
import numpy as np
# + [markdown] id="HO9sGi409sUo"
# ### <a name='c32'></a>4.2 Methods
#
# Users can use our package to deal with 3 different scenarios: 1 input to 1 output; multiple inputs to 1 output; multiple inputs to multiple outputs.
#
# + [markdown] id="Wmso3GnZYyth"
# #### <a name='c321'></a>4.2.1 1 input to 1 output
#
# Users may specify a single function with one input variable. See the example below with $f(x) = \exp(2x)$ with $x=2$.
# + id="ayr5nBZ69qbr"
x = ad.Variable(2, label="x") # specifies the input variable
f = ad.exp(2 * x) # specifies functions involving the variable
# + [markdown] id="2Gpj1AVf95qZ"
# We make two observations at this stage: First, arithmetic operations ($\exp$ in our example) need to be implemented within $\texttt{salad}$. Second, $\texttt{f}$ is of type $\texttt{Variable}$.
#
# To retrieve the value and derivative, call
# + colab={"base_uri": "https://localhost:8080/"} id="Ik0e3rOL93JG" outputId="0086669a-7112-4ce2-d6f0-98d4248d1f76"
val = f.val # array(exp(4)), numpy.array
val
# + colab={"base_uri": "https://localhost:8080/"} id="FD1-LLpf-Fm5" outputId="2f0a7ba7-a75a-4b52-b769-d0e4f5c55a12"
der = f.der["x"] # 2exp(4), float
der
# + [markdown] id="PlivG653klKI"
# The input value of the variable can also be a list consisting of multiple points. See the example below with $f(x) = \exp(2x)$ with $x=1,2,3,4$.
# + id="srzP7q_9ku0W"
x = ad.Variable([1,2,3,4], label="x") # specifies the input variable consisting of 4 points
f = ad.exp(2 * x) # specifies functions involving the variable
# + [markdown] id="fNGeUajXk145"
# The observations are stored in an array of the same length.
# + colab={"base_uri": "https://localhost:8080/"} id="DHrlb_Gck9kw" outputId="aab5057f-0896-4841-cf80-d92317a003dd"
val = f.val # value: exp(2x)
val
# + colab={"base_uri": "https://localhost:8080/"} id="AfnBRH6AlANu" outputId="3171c336-6b5b-4c07-fa5f-c69a55d0040c"
der = f.der["x"] # derivatives: 2exp(2x)
der
# + [markdown] id="Uu70p43F-RXQ"
# #### <a name='c322'></a>4.2.2 $m$ inputs to 1 output
#
# When there are multiple input variables, users can specify the name of the variable when initializing the instance. The name usually matches the variable name on the left hand side. See the example below with $f(x, y) = \exp(x+2y)$, with $x=1, y=1$.
# + id="H3LCiCrH-IS1"
x = ad.Variable(1, label='x') # specifies input variable x
y = ad.Variable(1, label='y') # specifies input variable y
f = ad.exp(x + 2 * y) # specifies functions involving the variables
# + colab={"base_uri": "https://localhost:8080/"} id="32kLH8l8-Yw_" outputId="5eae3ad8-027d-4677-b236-9f7985f6df96"
val = f.val # array(exp(3)), numpy.array
val
# + [markdown] id="OoYGPsHH-tD7"
#
#
# To get the derivative with respect to $x$, one simply uses "$x$" as the key to get the value from the dict $\texttt{f.der}$. Calling $\texttt{f.der}$ gives the gradient with respect to all input variables inside a dictionary.
# + colab={"base_uri": "https://localhost:8080/"} id="Ps-7E2yQ-Z1X" outputId="7b121fce-eef6-4720-99cc-99e1a18227df"
dx = f.der["x"] # exp(3), float
dx
# + colab={"base_uri": "https://localhost:8080/"} id="W2QEGPDT-bRk" outputId="562f980a-fe1c-4d8e-c1de-2bdc5cb29e34"
dy = f.der["y"] # 2exp(3), float
dy
# + colab={"base_uri": "https://localhost:8080/"} id="K-Qr5hFh-ceI" outputId="55c804a4-eac5-414a-aa5f-00a67b531e87"
grad = f.der # {"x": exp(3), "y": 2exp(3)}, dict
grad
# + [markdown] id="FKIU6EIYlqEd"
# Similarly, the users can calculate the values and derivatives of the function at multiple data points by using a list as input. See the example below with $f(x, y) = \exp(x+2y)$ at $(1, 1)$ and $(3, 0)$
# + id="hnLBi8Z-l9fc"
x = ad.Variable([1,3], label='x') # specifies input variable x
y = ad.Variable([1,0], label='y') # specifies input variable y
f = ad.exp(x + 2 * y) # specifies functions involving the variables
# + colab={"base_uri": "https://localhost:8080/"} id="NBwt7nBVmOZq" outputId="47da2e2f-3257-4722-b709-31ec86e609f1"
val = f.val # array(exp(3), exp(3)), numpy.array
val
# + colab={"base_uri": "https://localhost:8080/"} id="3Y7zXxQDmSVn" outputId="6cd82f30-d6f4-413b-f9ce-a198b9d84c9d"
grad = f.der # {"x": [exp(3), exp(3)], "y": [2exp(3), 2exp(3)]}, dict
grad
# + [markdown] id="mhPDp0CK-4kC"
# #### <a name='s1'></a>4.2.3 $m$ inputs to $n$ outputs
#
# We recognize that users may sometimes want to compute Jacobians. Consider the following case, $x = [x_1~x_2]^T, y = [y_1~y_2]^T, f(x, y) = [2x + \exp(y) ~ 3x+2\sin(y)]^T$.
#
# We provide another class $\texttt{Forward}$ to do the calculation.
# + colab={"base_uri": "https://localhost:8080/"} id="rAqhnBsK-df6" outputId="ae0c8f85-fcaf-4e41-cfe5-5ccab6d1dbb0"
variables = {'x': 3, 'y': 5} # specifies input values
functions = ['2*x + exp(y)', '3*x + 2*sin(y)'] # specifies functions
f = ad.Forward(variables, functions)
print(f)
# + [markdown] id="z_x6w5Yl_H0k"
# In this case, all computed results will be saved in $\texttt{f.results}$, which is a list of values and derivatives of the two functions. The user can obtain the value and the derivatives of the $i^{th}$ function by calling
# + colab={"base_uri": "https://localhost:8080/"} id="JxfmeaoO_FIM" outputId="33547823-61d8-40ef-a295-5c9b198d7e26"
val1 = f.results[0].val # array(11)
val1
# + colab={"base_uri": "https://localhost:8080/"} id="ca7kOGgW_M9b" outputId="e052494b-109c-4737-eb1f-145440410893"
der1 = f.results[0].der # {'x': array(2.), 'y': array(1.)}
der1
# + colab={"base_uri": "https://localhost:8080/"} id="yc9qBskw_QKG" outputId="905c1a79-e767-4ac9-e1c8-264b169d8a43"
val2 = f.results[1].val # array(19)
val2
# + colab={"base_uri": "https://localhost:8080/"} id="dI8TS8nX_RxO" outputId="7e0d41eb-f793-4bbe-e6a8-b4f1636f2f23"
der2 = f.results[1].der # {'x': array(3.), 'y': array(2.)}
der2
# + [markdown] id="z_wcX2E-me2n"
# The input can also be a list of values.
# + colab={"base_uri": "https://localhost:8080/"} id="GPdl-oH3ma3c" outputId="04c01167-79e0-4e65-b79e-90026a407cae"
variables = {'x': [3,4], 'y': [2,3]} # specifies input values
functions = ['2*x + exp(y)', '3*x + 2*sin(y)'] # specifies functions
f = ad.Forward(variables, functions)
print(f)
# + [markdown] id="WVp6TPX4FR5-"
# ## <a name='c5'></a>5 Implementation
# + [markdown] id="mh6oTIHAZsi1"
# ### <a name='c51'></a>5.1 Data Structure
#
# The core data structures used in our implementation are $\texttt{list}$, $\texttt{numpy.array}$, and $\texttt{dict}$. See the [m inputs to n outputs](#s1) example from above as an illustration. The input variables are in a $\texttt{dict}$. The input functions are in a $\texttt{dict}$. The Jacobian with respect to each vector-form input is in $\texttt{numpy.array}$. To easily match the Jacobian to the corresponding input vector, we use a $\texttt{dict}$, where the key is the name of the corresponding input variable, and the value is a $\texttt{numpy.array}$ of the Jacobian.
# + [markdown] id="_sgkZaYEZuXu"
# ### <a name='c52'></a>5.2 Classes and Methods
#
# The baseline implementation currently includes two classes: 1) $\texttt{Variable}$, 2) $\texttt{Forward}$
# + [markdown] id="iIblG_jIZwMM"
# #### <a name='c521'></a>5.2.1 $\texttt{Variable}$
#
# In $\texttt{Variable}$, we re-define dunder methods and other arithmetic operations. See below for the structure of the class and method definition.
# + id="UmrUikCpE8OS"
class Variable(object):
counter = 0
def __init__(self, val, der=None, label=None, ad_mode="forward", increment_counter=True):
pass
def __add__(self, other):
pass
def __str__(self):
pass
def __radd__(self, other):
pass
def __sub__(self, other):
pass
def __rsub__(self, other):
pass
def __mul__(self, other):
pass
def __rmul__(self, other):
pass
def __truediv__(self, other):
pass
def __rtruediv__(self, other):
pass
def __neg__(self, other):
pass
def __pow__(self, other):
pass
def __rpow__(self, other):
pass
def __eq__(self, other):
pass
def __ne__(self, other):
pass
def __lt__(self, other):
pass
def __le__(self, other):
pass
def __gt__(self, other):
pass
def __ge__(self, other):
pass
# + [markdown] id="G1iRG-OwGq8a"
# On a high level, each instance of this class corresponds to a node in the computation graph in the background section. We keep track of the primal trace ($\texttt{self.val}$) and the tangent trace ($\texttt{self.der}$) as attributes for all instances.
#
# We also implement elementary functions like $\texttt{sin}$, $\texttt{sqrt}$, $\texttt{log}$, and $\texttt{exp}$.
# + id="RF0Ml0qpGo5H"
def sin(x):
pass
def cos(x):
pass
def tan(x):
pass
def exp(x):
pass
def log(x):
pass
def ln(x):
pass
def sqrt(x):
pass
def logistic(x):
pass
def arcsin(x):
pass
def arccos(x):
pass
def arctan(x):
pass
def sinh(x):
pass
def cosh(x):
pass
def tanh(x):
pass
# + [markdown] id="IcLBtPVwG38J"
# Users can use these functions for simple numeric calculations:
# + colab={"base_uri": "https://localhost:8080/"} id="icjsHly7Gz3m" outputId="d306d49d-58af-40eb-cd22-d34216657e39"
x = 3
ans = ad.sin(x)
print(f'x = {ans}, \ntype of x: {type(ans)}')
# + [markdown] id="tTS89pOMG90J"
# Or, users can use these functions to compute function derivatives. In this case, the inputs should be of the type $\texttt{Variable}$.
# + colab={"base_uri": "https://localhost:8080/"} id="fRkxw6-MG7Zg" outputId="da6150b3-38cf-4b2b-f95b-e8a0eecfe888"
x = ad.Variable(3, label="x")
ans = ad.sin(x)
print(f'x = {ans.val}, x.der = {ans.der}')
# + colab={"base_uri": "https://localhost:8080/"} id="exviNmWnHCtP" outputId="ceb6c58d-ad37-43f8-bc35-9318890d8ef5"
print(type(ans))
# + [markdown] id="zwhn6mIoHF0N"
# More arithemetic operations and functions to be included if time allows and as we see fit.
#
# #### <a name='c522'></a>5.2.2 $\texttt{Forward}$
# We also include a $\texttt{Forward}$ class for easy handling of multiple function input. The variables are input in a $\texttt{dict}$ and the functions are input in a $\texttt{list}$. The $\texttt{__init__}$ function within $\texttt{Forward}$ simply loops through and evaluate each functions one by one. See the [m inputs to n outputs](#s1) example from above as an illustration.
# + [markdown] id="Gq1vGCePbris"
# ## <a name='c6'></a>6 Extensions
#
# We implement an optimization toolkit as the extension feature. These optimization methods are useful in numerous contexts such as statistics, machine learning, operations research, and economics. The `Optimize` module includes the following optimization schemes:
#
# 1. Gradient Descent (GD)
# 2. Broyden-Fletcher-Goldfarb-Shanno (BFGS)
# 3. Stochastic Gradient Descent (SGD)
#
# The intended use of the first two methods, GD and BFGS is as follows:
# 1. users input an arbitrary function supported by `Salad`, $f(\textbf{x})$, where $\textbf{x} \in \mathbb R^n$
# 2. users supply a starting value, $\textbf{x}_0 \in \mathbb R^n$, at which the method starts to search for a local minimum
# 3. the optimization method returns $\mathbf{x}^* \in \mathbb R^n$ that achieves a local minimum of the function $f(x)$.
#
# The third method, SGD, is implemented in the context of a linear regression, $\mathbf{y}$ regressed on $\mathbf{X}$.
#
# The intended uses of SGD is as follows:
# 1. users input covariate matrix $\mathbf{X} \in \mathbb R^{nxp}$ with $n$ rows ($n$ observations) and $p$ columns ($p$ covariates); and input response vector $\mathbf{y} \in \mathbb R^n$ with $n$ entries ($n$ observations).
# 2. users supply the batch size used in each SGD iteration
# 3. user specify the starting position of the search for coefficient estimates
# 4. SGD returns the regression coefficient $\mathbf{b}^* \in \mathbb R^b$.
# + [markdown] id="WUPflnkrG8NE"
# Users first need to import the optimization toolkit at `cs107_salad.Optimization.optimize`.
# + id="uZLBHhyruEwm"
import cs107_salad.Optimization.optimize as optimize
# + [markdown] id="spu4WajJr9ph"
# ### <a name='c61'></a>6.1 [Gradient Descent (GD)](https://en.wikipedia.org/wiki/Gradient_descent)
#
# Gradient descent is an iterative optimization algorithm to find a local minimum of some function. The method iteratively calculates the gradient at each given point, and goes toward the direction of steepest descent.
#
# Specifically, consider a differentiable multi-variable function $f(\textbf{x})$. Starting with an initial guess $\textbf{x}_0$, at each step $k$, the method updates $x_{k+1}$ in the following way:
#
# $$\textbf{x}_{k+1} = \textbf{x}_k - \lambda \nabla F(\textbf{x}_k)$$
#
# The algorithm terminates when the magnitude of the update $\lambda \nabla F(\textbf{x}_k)$ is smaller than a tolerance level or the maximum number of iterations is reached. $\lambda$ is the learning rate that adjusts for how much the method goes in the negative gradient direction.
# + [markdown] id="0srk8RjruLuC"
# Below is an demonstration of calling GD to find the minimin of a convex function $x^2 + 2y^2$. `min_params` is the $\mathbf{x}$ value that gives a local minimum, `val` is the function value at this local minimum, and `der` is the gradient at this local minimum.
# + id="0J-aFluIuXvu"
f = "x**2 + 2*y**2"
starting_pos = {"x": 5, "y": 2}
GD = optimize.GradientDescent()
min_params, val, der, hist = GD.optimize(f, starting_pos, full_history=True)
# + colab={"base_uri": "https://localhost:8080/"} id="VknNNiB8ucaZ" outputId="96a1758b-15a6-401b-a3dd-026ec0550e92"
print('optimized parameter:', min_params)
print('function value at minimum', val)
print('gradient at minimum:', der)
# + [markdown] id="3RHDcYJowAoX"
# Users may ask `GD.optimize` to output the search history by including a `full_history` flag. The full history allows users to easily visualize the search path.
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="d4hJEMr-jLdQ" outputId="943280a9-324b-4779-b790-b8614d0d6e43"
import matplotlib.pyplot as plt
def f(x):
x, y = x
return x**2 + y**2
def plot_f(x,y,canvas, title):
X, Y = np.meshgrid(x,y)
Z = f([X,Y])
canvas.contour(X,Y,Z,levels=np.linspace(0,100,80), alpha=0.3)
canvas.plot([0],[0],'ro',lw=2, label='global min')
canvas.set_xlabel('x')
canvas.set_ylabel('y')
canvas.set_title(title)
canvas.legend(loc=3)
_, axs = plt.subplots(1,1,figsize=(8,6))
x = np.linspace(-6,6,100)
y = np.linspace(-6,6,100)
plot_f(x,y,axs,title='Gradient Descent Path')
axs.plot(hist['x'][0], hist['y'][0], 'bo', lw=1, label='starting position');
axs.plot(hist['x'], hist['y'], 'b--', lw=1, label='search path');
axs.legend();
# + [markdown] id="DC6rDBjnuoom"
# ### <a name='c62'></a>6.2 [Broyden–Fletcher–Goldfarb–Shanno (BFGS)](https://en.wikipedia.org/wiki/Broyden%E2%80%93Fletcher%E2%80%93Goldfarb%E2%80%93Shanno_algorithm)
#
# Broyden–Fletcher–Goldfarb–Shanno (BFGS) is a popular quasi-Newton Method that iteratively finds a local minimum of some functions using approximated Hessian.
#
# Specifically, consider a differentiable multi-variable function $f(\textbf{x})$. BFGS starts with an initial guess $\textbf{x}_0$, and $\mathbf{B}_0 = \mathbf{I}$ before the first iteration. The algorithm computes the following in iteration $k$:
#
# 1. Solve for $\mathbf{B}_k$ that satisfies $\mathbf{B}_k \mathbf{s}_k = -\nabla f(\mathbf{x}_k)$
# 2. Update $\mathbf{x}_{k+1} = \mathbf{x}_k + \mathbf{s}_k$
# 3. Compute $\mathbf{y}_k = \nabla f(\mathbf{x}_{k+1}) - \nabla f(\mathbf{x}_k)$
# 4. Update $\mathbf{B}_{k+1} = \mathbf{B}_k + \Delta \mathbf{B}_k$
#
# where $\Delta B_k = \frac{\textbf{y}_{k}\textbf{y}_{k}^T}{\textbf{y}_{k}^T\textbf{s}_{k}} - \frac{B_k\textbf{s}_{k}\textbf{s}_{k}^TB_k}{\textbf{s}_{k}^TB_k\textbf{s}_{k}}$
#
# The algorithm terminates when the update $\mathbf{s}_k$ is smaller than a tolerance or if the maximum number of iterations has been reached.
# + [markdown] id="fzpb45RkxNg0"
# The example below finds the local (global) minimum for a convex function $x^2 + 2y^2$. `min_params` stores the $\mathbf{x}$ value that attains the local (global) minimum, `val` stores the function value at the minimum, and `hist` stores the entire search path of $\mathbf{x}$.
# + id="KJmlP7gzxEL1"
f = "x**2 + 2*y**2"
starting_pos = {"x": 5.0, "y": 2.0}
BFGS = optimize.BFGS()
min_params, val, hist = BFGS.optimize(f, starting_pos, full_history=True)
# + colab={"base_uri": "https://localhost:8080/"} id="VPko50P5xbf3" outputId="b6eb706e-fe85-4775-b4fe-7b1436c7da31"
print('optimized parameter:', min_params)
print('function value at minimum', val)
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="lQZrInU1m4kC" outputId="6b553da8-a47f-4757-cefc-f9c0a7754680"
import matplotlib.pyplot as plt
def f(x):
x, y = x
return x**2 + y**2
def plot_f(x,y,canvas, title):
X, Y = np.meshgrid(x,y)
Z = f([X,Y])
canvas.contour(X,Y,Z,levels=np.linspace(0,100,80), alpha=0.3)
canvas.plot([0],[0],'ro',lw=2, label='global min')
canvas.set_xlabel('x')
canvas.set_ylabel('y')
canvas.set_title(title)
canvas.legend(loc=3)
_, axs = plt.subplots(1,1,figsize=(8,6))
x = np.linspace(-6,6,100)
y = np.linspace(-6,6,100)
plot_f(x,y,axs,title='Gradient Descent Path')
axs.plot(hist['x'][0], hist['y'][0], 'bo', lw=1, label='starting position');
axs.plot(hist['x'], hist['y'], 'bx--', lw=1, label='search path');
axs.legend();
# + [markdown] id="kP5hGJCxxn_c"
# ### <a name='c63'></a>6.3 [Stochastic Gradient Descent (SGD)](https://en.wikipedia.org/wiki/Stochastic_gradient_descent)
#
# Stochastic gradient descent (SGD) algorithm iteratively finds a local minimum of some functions with suitable smoothness properties. It is a more generalized version of gradient descent that involves randomness in gradient calculation.
#
# In each iteration of SGD, it randomly samples a subset (batch) of the entire input dataset, and conducts one step of gradient descent using the gradient calculated from the subset to reduce computational burden. SGD terminates when the maximum number of iterations is reached.
#
# Our implementation of SGD has a limited use case: linear regression. In particular, users specify a covariate matrix $\mathbf{X}$ and a response vector $\mathbf{y}$, our SGD function aims to minimize the following function:
#
# $$\mathbf{b}^* = \arg\min_{\mathbf{b}} = \frac{1}{n} \sum_{i=1}^n \left(y_i - \mathbf{X}_i \mathbf{b}\right)^2$$
#
# where $\mathbf{X} \in \mathbb R^{n\times p}, \mathbf{y} \in \mathbb R^n, \mathbf{b}\in \mathbb R^p$. Note that the objective as a function of the estimated coefficients $\mathbf{b}$ is convex. Therefore, SGD in this case actually leads to the global minimum. $\mathbf{b}^*$ is the estimated coefficient by minimizing mean squared error.
#
# + [markdown] id="oAJPPmYfzDXA"
# The folloing example is to calculate the coefficient of regressing $\mathbf{y}$ on $\mathbf{X}$. `min_params` is the estimated cofficients, `val` is the loss attained at the estiamted coefficients, and `der` is the gradient at the minimum loss.
# + id="91ibOTltzO5p"
X = np.random.rand(100, 3)
y = X @ np.array([2, 0, 3]) + 0.001 * np.random.rand(X.shape[0]) # ground truth data generating process
SGD = optimize.StochasticGradientDescent(X, y, batch_size=10)
min_params, val, der = SGD.optimize([0, 0, 0], max_iter=5000, learning_rate=0.01)
# + colab={"base_uri": "https://localhost:8080/"} id="JZpm7WvuzWcI" outputId="07630054-c6f6-40c8-9432-49b1c9a30824"
print('optimized parameter:', min_params)
# + [markdown] id="zC16wjuBS2oo"
# ## <a name='c7'></a>7 Broader Impact
#
# Our extension involves several optimization methods: gradient descent, stochastic gradient descent, BFGS, and Newton’s method. By itself, these are facially neutral algorithms – they are not discriminatory on their face – yet they have clear potential discriminatory applications and effects. Our subpackages could easily be extended and combined with other software modules to create bad, biased, or unethical AI models. For instance, Amazon was forced to scrap its automated recruiting tool when it was revealed that the algorithm was biased against women. The algorithm was biased because it was trained on resumes previously submitted to Amazon. However, reflecting unequal trends pervasive throughout the tech industry, most candidates that made it through the recruitment process were men. Amazon’s model, when optimized to this data, learned that resumes from men were preferable to resumes from women. One could imagine how our package, when used to optimize on biased data from any setting would have negative disparate impacts.
#
#
#
# + [markdown] id="X0pRsLzoS9nE"
#
# ## <a name='c8'></a>8 Software Inclusivity
#
# The salad package warmly welcomes all people regardless of background to contribute to the development of our package. Our package is open-source, easily accessible through Github and PyPi, and licensed under the MIT License. Nonetheless, we recognize there are many structural barriers for the average person to interact with and contribute to our software package. For instance, while we aimed to have high code quality with clear comments, our comments are all in English. As a result, non-native English speakers are at a disadvantage in contributing to our package. Moreover, computer science/programming education and education in general are not shared equally across society. BIPOC, the poor, women, and other minorities are all not afforded the same educational opportunities as the cis-male White and Asian men that currently make up the majority of programmers in the US. Such underrepresented minorities face more constraints in contributing to open source software projects like ours.
#
#
#
# + [markdown] id="KAc_mchmZ5OV"
# ## <a name='c9'></a>9 Licensing
#
# We will use the MIT License. Future developers are welcome to extend our library and distribute closed source versions. Because we are using NumPy, a copyright licensed library, we are unable to use a license like GNU GPLv3 (as it requires all libraries used by the library to be copyleft).
# + [markdown] id="KdbEXA36S_BX"
# ## <a name='c10'></a>10 Future Directions
#
# There is much room for improvement of our package from supporting higher and mixed derivatives to building out support for differential programming.
#
# The four areas that we are most interested in, however, are the following:
#
# 1. Support for reverse mode. Our package currently supports computing derivatives to machine precision of functions $f:\mathbb{R}^m\to\mathbb{R}^n$. However, forward mode is most efficient when $n \gg m$, ie the number of functions we are evaluating is much greater than the number of inputs. When $n \ll m$, i.e. the number of functions we are evaluating is much smaller than the number of inputs, reverse mode is actually more efficient.
#
# - Support for backpropagation. Backpropagation is a special case of reverse mode that is used for scalar objective functions where our objective function represents the error between our output and the true value. Implementing backpropagation would allow us to extend our package to be used for developing fully connected neural networks.
#
# 2. Building out support for more elementary functions such as the inverse hyperbolic functions.
#
# 3. Building out more user friendly tools such as a GUI that allows the user to visualize the computational graph of the functions they have entered.
#
# 4. The optimization toolkit is also preliminary at the moment. A couple ideas for future extensions include:
#
# a. Implementation of the Newton's Method: Newton's Method for finding the local minimum of a function uses the Hessian, which requires a further extension to our `salad` implementation. We could include the computation of the Hessian in our implementation, potentially as a third variable within the `Variable` class.
#
# b. Improvement of the current implementation of SGD: Currently, the only use case of our SGD is for regression (i.e. minimize the mean squared error). The method could benefit from
# - Allowing users to specify any loss function of their choice
# - Automic hyper-parameter tuning
# - Repeated search from multiple independent starting point to achieve a lower local minimum
#
# c. Implementation of other optimization fucntions such as `Adam`, `RMSprop`, and `FTRL` which are more stable and more efficient optimizers commonly used for deep learning and neural networks.
#
# + id="l0jBVBBpzzWG"
| docs/Documentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# First of all, the necessary libraries:
# %matplotlib inline
#pytorch packages
import math
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from torch.autograd import Variable
#To download the dataset for torchvision
import torchvision
from torchvision import datasets, transforms
#For plots
import matplotlib.pyplot as plt
# # Download and look at the data
# For this example, we will try to recognize hand-written digits, specifically the ones of the MNIST dataset, that contains overall 70,000 28-by-28-pixels pictures of hand-written digits. This dataset is easily accessible in pytorch via dataset.MNSIT. You just have to specify you want to download it if it's not already in the directory, and pytorch will process it to create a DataSet.
#Change to the directory of your choice.
PATH = './Mnist/'
# Grab the training and the test set.
trn_set = datasets.MNIST(PATH, train=True, download=True)
tst_set = datasets.MNIST(PATH, train=False, download=True)
# Let's have a look at the data in the training set first.
len(trn_set.data), len(tst_set.test_data)
# Each image is represented by a tensor of size 28 by 28, each value represents the color of the corresponding pixel, from 0 (black) to 255 (white). Torch tensors are the equivalent of numpy ndarrays.
trn_set.data[0]
# It's easy to convert a torch tensor to a numpy array via the .numpy() command.
#
# Conversely, you can create a torch Tensor from a numpy array x via torch.Tensor(x)
trn_set.data[0].numpy()
# It's then easy to see the corresponding picture via plt.
plt.imshow(trn_set.data[0].numpy(), cmap='Greys')
# Let's have a look at the corresponding label...
trn_set.targets[0]
# # Preparing the data
# A pytorch neural network will expect the data to come in the form of minibatches of tensors. To do that, we use a pytorch object called DataLoader. It will randomly separate the pictures (with the associated label) in minibatches. If you have multiple GPUs, it also prepares the work to be parallelized between them (just change num_workers from 0 to your custom value). We only shuffle the data randomly for the training.
#
# First we need to explicitely ask our dataset to transform the images in tensors.
tsfms = transforms.ToTensor()
trn_set = datasets.MNIST(PATH, train=True, download=True, transform=tsfms)
tst_set = datasets.MNIST(PATH, train=False, download=True, transform=tsfms)
trn_loader = torch.utils.data.DataLoader(trn_set, batch_size=64, shuffle=True, num_workers=0)
tst_loader = torch.utils.data.DataLoader(tst_set, batch_size=64, shuffle=False, num_workers=0)
# Let's have a look at an example. A data loader can be converted into an iterator and we can then ask him for a minibatch.
mb_example = next(iter(trn_loader))
# Such a minibacth containts two torch tensors: the first one contains the data (here our pictures) and the second one the expected labels.
mb_example[0].size(), mb_example[1].size()
# Note that pytorch has automatically added one dimension to our images (the 1 in second position). It would be 3 if we had had the three usual channels for the colors (RGB). Pytorch puts this channel in the second dimension and not the last because it simplifies some computation.
#
# Let's see the first tensor.
mb_example[0][0,0]
# Note that pytorch transformed the values that went from 0 to 255 into floats that go from 0. to 1.
#
# We can have a look at the first pictures and draw them.
fig = plt.figure()
for i in range(0,4):
sub_plot = fig.add_subplot(1,4,i+1)
sub_plot.axis('Off')
plt.imshow(mb_example[0][i,0].numpy(), cmap='Greys')
sub_plot.set_title(str(mb_example[1][i].item()))
# Another usual transformation we do before feeing the pictures to our neural network is to normalize the input. This means subtracting the mean and dividing by the standard deviation. We can either search for the usual values on Google or compute them from scratch.
mean = torch.mean(trn_set.data.type(torch.FloatTensor))/255.
std = torch.std(trn_set.data.type(torch.FloatTensor))/255.
mean,std
# We divide by 255 to get the means of our data when it's convereted into floats from 0. to 1.
#
# Then we go back to creating a transfrom and add the normalization. Note that we use the same mean and std for the test set. Afterward, we reload our datasets, adding this transform.
tsfms = transforms.Compose([transforms.ToTensor(), transforms.Normalize((mean,), (std,))])
trn_set = datasets.MNIST(PATH, train=True, download=True, transform=tsfms)
tst_set = datasets.MNIST(PATH, train=False, download=True, transform=tsfms)
trn_loader = torch.utils.data.DataLoader(trn_set, batch_size=64, shuffle=True, num_workers=0)
tst_loader = torch.utils.data.DataLoader(tst_set, batch_size=64, shuffle=False, num_workers=0)
# Now if we want to plot our digits, we will have to denormalize the images.
mb_example = next(iter(trn_loader))
fig = plt.figure()
for i in range(0,4):
sub_plot = fig.add_subplot(1,4,i+1)
sub_plot.axis('Off')
plt.imshow(mb_example[0][i,0].numpy() * std.numpy() + mean.numpy(), cmap='Greys', interpolation=None)
sub_plot.set_title(str(mb_example[1][i].item()))
# # Create a model
# It's always a good idea to create a model as a subclass of nn.Module. That way, we can use all the features this class provides.
#
# We override the init function (but still call the init function of nn.Module) to define our custom layers (here two linear layers) and we have to define the forward function, which explains how to compute the output.
#
# The first line of the forward function is to flatten our input, since we saw it has four dimensions: minibatch by channel by height by width. We only keep the minibatch size as our first dimension (x.size(0)) and the -1 is to tell pytorch to determine the right number for the second dimension.
class SimpleNeuralNet(nn.Module):
def __init__(self, n_in, n_hidden, n_out):
super().__init__()
self.linear1 = nn.Linear(n_in, n_hidden)
self.linear2 = nn.Linear(n_hidden, n_out)
def forward(self,x):
x = x.view(x.size(0),-1)
x = F.relu(self.linear1(x))
return F.log_softmax(self.linear2(x), dim=-1)
# Then we can instanciate the class with our input size (28 * 28), an hidden size of 100 layers and 10 outputs (as many as digits).
#
# The optimizer will automatically do the Stochastic Gradient Descent for us (or any of its variant if we want).
net = SimpleNeuralNet(28*28,100,10)
optimizer = optim.SGD(net.parameters(),lr=1e-2)
# Now we're ready to write our training loop. To compute the gradient automatically, pytorch requires us to put the torch tensors with our inputs and labels into Variable objects, that way it'll remember the transformation these go through until we arrive at our loss function. We then call loss.backward() to compute all the gradients (which will then be in the grad field of any variable).
#
# The optimizer takes care of the step of our gradient descent in the optimizer.step() function. Since the gradients are accumulated, we have to tell pytorch when to reinitialize them (which the purpose of the optimizer.zero_grad() command at the beginning).
def train(nb_epoch):
for epoch in range(nb_epoch):
running_loss = 0.
corrects = 0
print(f'Epoch {epoch+1}:')
for data in trn_loader:
#separate the inputs from the labels
inputs,labels = data
#wrap those into variables to keep track of how they are created and be able to compute their gradient.
inputs, labels = Variable(inputs), Variable(labels)
#Put the gradients back to zero
optimizer.zero_grad()
#Compute the outputs given by our model at this stage.
outputs = net(inputs)
_,preds = torch.max(outputs.data,1)
#Compute the loss
loss = F.nll_loss(outputs, labels)
running_loss += loss.data * inputs.size(0)
corrects += torch.sum(labels.data == preds)
#Backpropagate the computation of the gradients
loss.backward()
#Do the step of the SGD
optimizer.step()
print(f'Loss: {running_loss/len(trn_set)} Accuracy: {100.*corrects/len(trn_set)}')
# Now we're ready to train our model.
train(10)
# 96.3% accuracy is good, but that's on the training set and we may be overfitting. Let's try on the test set now to see if we're doing well or not.
# +
def validate():
running_loss = 0.
corrects = 0
for data in tst_loader:
#separate the inputs from the labels
inputs,labels = data
#wrap those into variables to keep track of how they are created and be able to compute their gradient.
#Even if we don't require the gradient here, a nn.Module expects a variable.
inputs, labels = Variable(inputs), Variable(labels)
#Compute the outputs given by our model at this stage.
outputs = net(inputs)
_,preds = torch.max(outputs.data,1)
#Compute the loss
loss = F.nll_loss(outputs, labels)
running_loss += loss.data * inputs.size(0)
corrects += torch.sum(labels.data == preds)
print(f'Loss: {running_loss/len(tst_set)} Accuracy: {100.*corrects/len(tst_set)}')
def validate_2():
net.eval()
running_loss = 0.
corrects = 0
for data in tst_loader:
#separate the inputs from the labels
inputs,labels = data
#wrap those into variables to keep track of how they are created and be able to compute their gradient.
#Even if we don't require the gradient here, a nn.Module expects a variable.
inputs, labels = Variable(inputs), Variable(labels)
#Compute the outputs given by our model at this stage.
outputs = net(inputs)
_,preds = torch.max(outputs.data,1)
#Compute the loss
loss = F.nll_loss(outputs, labels)
running_loss += loss.data * inputs.size(0)
corrects += torch.sum(labels.data == preds)
print(f'Loss: {running_loss/len(tst_set)} Accuracy: {100.*corrects/len(tst_set)}')
# -
validate()
validate_2()
# So we weren't overfitting!
# # Learning rate finder
# The details of how this code has been built are all explained in this [blog article](https://sgugger.github.io/how-do-you-find-a-good-learning-rate.html).
def find_lr(init_value = 1e-8, final_value=10., beta = 0.98):
num = len(trn_loader)-1
mult = (final_value / init_value) ** (1/num)
lr = init_value
optimizer.param_groups[0]['lr'] = lr
avg_loss = 0.
best_loss = 0.
batch_num = 0
losses = []
log_lrs = []
for data in trn_loader:
batch_num += 1
#As before, get the loss for this mini-batch of inputs/outputs
inputs,labels = data
inputs, labels = Variable(inputs), Variable(labels)
optimizer.zero_grad()
outputs = net(inputs)
loss = criterion(outputs, labels)
#Compute the smoothed loss
avg_loss = beta * avg_loss + (1-beta) *loss.data
smoothed_loss = avg_loss / (1 - beta**batch_num)
#Stop if the loss is exploding
if batch_num > 1 and smoothed_loss > 4 * best_loss:
return log_lrs, losses
#Record the best loss
if smoothed_loss < best_loss or batch_num==1:
best_loss = smoothed_loss
#Store the values
losses.append(smoothed_loss)
log_lrs.append(math.log10(lr))
#Do the SGD step
loss.backward()
optimizer.step()
#Update the lr for the next step
lr *= mult
optimizer.param_groups[0]['lr'] = lr
return log_lrs, losses
# Now we can define our neural net as before.
net = SimpleNeuralNet(28*28,100,10)
optimizer = optim.SGD(net.parameters(),lr=1e-1)
criterion = F.nll_loss
# Then plot the losses versus the logs of the learning rate
logs,losses = find_lr()
plt.plot(logs[10:-5],losses[10:-5])
# This suggests the best learning rate is $10^{-1}$ so we can use test this one after defining a new network.
net = SimpleNeuralNet(28*28,100,10)
optimizer = optim.SGD(net.parameters(),lr=1e-1)
train(1)
# We are already at 92.21% accuracy when the learning rate used before gave us 84.86% in one epoch!
| Pytorch/how_to_find_a_good_learning_rate/Learning rate finder.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_pytorch_p36
# language: python
# name: conda_pytorch_p36
# ---
# ## Benchmark model
# In this notebook a Simple Moving Average model will be created in order to have a benchmark for my DeepAR based model (to be tuned in the next notebook).
# ### Hyperparameters
# DeepAR is the model of choice for this project.
# This model expects input data to be already test-train split.
# A big part of the model design has to be done looking close at data.
# More specifically, defining these two hyperparameters about the data:
# * Context length
# * Prediction length
# ### Prediction length
# This is the length of the time series future predictions in days. It will be conveniently set to 20 days (exactly 4 weeks of trading hours). Smaller intervals could also be analyzed.
# Anyway, any interval shorter than 5 days (one week of trading hours) would be of little significance.
# A longer interval could be interesting from an application point of view, but it can be challenging in terms of model performances.
# ### Context length
# Context length can be either:
# * designed on patterns or seasonality observed in the data, if any is present;
# * chosen as a fixed value. This will be my choice, and it will be the same as the moving average window, in order to have a good reference metrics, applicable to both this model and the benchmark model.
# To explore this second option, we will refer to what we've found during the EDA stage.
# declaring prediction legnth and context length
prediction_length = [10, 20, 50]
context_length = [10, 20, 50]
# ### Train, test and validation split
# Time series will be all trimmed from `data_start` in `1.ExploratoryDataAnalysis.ipy`, according to DeepAR documentation, train time series should be set as the size of the entire time series less the prediction length.
# Validation length will exactly equal to prediction length.
from utils.data_prepare import train_test_valid_split
# #### IBM Stock train, test and validation split
# +
#df_ibm = df_ibm.loc[data_start:].copy()
# -
df_ibm_train, df_ibm_test, df_ibm_valid = train_test_valid_split(df_ibm, prediction_length=prediction_length[1])
print(len(df_ibm), len(df_ibm_train), len(df_ibm_test), len(df_ibm_valid))
df_ibm_train.head()
df_ibm_test.head()
df_ibm_valid.head()
# #### Apple Inc. Stock train test split
# +
#df_aapl = df_aapl.loc[data_start:].copy()
# -
df_aapl_train, df_aapl_test, df_aapl_valid = train_test_valid_split(df_aapl, prediction_length=prediction_length[1])
print(len(df_aapl), len(df_aapl_train), len(df_aapl_test), len(df_aapl_valid))
df_aapl_train.head()
df_aapl_test.head()
df_aapl_valid.head()
# #### Amazon Stock train test split
# +
#df_amzn = df_amzn.loc[data_start:].copy()
# -
df_amzn_train, df_amzn_test, df_amzn_valid = train_test_valid_split(df_amzn, prediction_length=prediction_length[1])
print(len(df_amzn), len(df_amzn_train), len(df_amzn_test), len(df_amzn_valid))
df_amzn_train.head()
df_amzn_test.head()
df_amzn_valid.head()
# #### Alphabet Inc. Stock train test split
# +
#df_googl = df_googl.loc[data_start:].copy()
# -
df_googl_train, df_googl_test, df_googl_valid = train_test_valid_split(df_googl, prediction_length=prediction_length[1])
print(len(df_googl), len(df_googl_train), len(df_googl_test), len(df_googl_valid))
df_googl_train.head()
df_googl_test.head()
df_googl_valid.head()
# ### Test and Validation Data sets Retro-fix
# It is clear that the above initialized values has introduced some kind of data leakage into test and validation data sets, from the moment that SMA computation should be stopped at the end of train time series. In order to solve this issue, I will set all the simple moving average values in the validation set and in the last part (last 20 values) of test set to the value of the last one in the training set.
# #### IBM Data
# set moving average and Bollinger bands columns values to same values in last test set record
for mds in win_szs:
ma_column_name = "%s_ac_ma" %(str(mds))
df_ibm_valid[ma_column_name] = df_ibm_train[ma_column_name][-1]
bb_column_name_u = "%s_ac_bb_u" %(str(mds))
df_ibm_valid[bb_column_name_u] = df_ibm_train[bb_column_name_u][-1]
bb_column_name_l = "%s_ac_bb_l" %(str(mds))
df_ibm_valid[bb_column_name_l] = df_ibm_train[bb_column_name_l][-1]
# #### AAPL Validation Data
# set moving average and Bollinger bands columns values to same values in last test set record
# set moving average and Bollinger bands columns values to same values in last test set record
for mds in win_szs:
ma_column_name = "%s_ac_ma" %(str(mds))
df_aapl_valid[ma_column_name] = df_aapl_train[ma_column_name][-1]
bb_column_name_u = "%s_ac_bb_u" %(str(mds))
df_aapl_valid[bb_column_name_u] = df_aapl_train[bb_column_name_u][-1]
bb_column_name_l = "%s_ac_bb_l" %(str(mds))
df_aapl_valid[bb_column_name_l] = df_aapl_train[bb_column_name_l][-1]
df_aapl_valid.head()
# #### AMZN Validation Data
# set moving average and Bollinger bands columns values to same values in last test set record
for mds in win_szs:
ma_column_name = "%s_ac_ma" %(str(mds))
df_amzn_valid[ma_column_name] = df_amzn_train[ma_column_name][-1]
bb_column_name_u = "%s_ac_bb_u" %(str(mds))
df_amzn_valid[bb_column_name_u] = df_amzn_train[bb_column_name_u][-1]
bb_column_name_l = "%s_ac_bb_l" %(str(mds))
df_amzn_valid[bb_column_name_l] = df_amzn_train[bb_column_name_l][-1]
df_amzn_valid.head()
# #### GOOGL Validation Data
# set moving average and Bollinger bands columns values to same values in last test set record
for mds in win_szs:
ma_column_name = "%s_ac_ma" %(str(mds))
df_googl_valid[ma_column_name] = df_googl_train[ma_column_name][-1]
bb_column_name_u = "%s_ac_bb_u" %(str(mds))
df_googl_valid[bb_column_name_u] = df_googl_train[bb_column_name_u][-1]
bb_column_name_l = "%s_ac_bb_l" %(str(mds))
df_googl_valid[bb_column_name_l] = df_googl_train[bb_column_name_l][-1]
df_googl_valid.head()
n = 20
ma_str = str(n)+'_ac_ma'
# Now I'll initialize an array of moving average values on test and validation time series, to be used in future model comparison:
ser_valid_ma_bmk = [df_ibm_valid[:][ma_str], df_aapl_valid[:][ma_str], df_amzn_valid[:][ma_str], df_googl_valid[:][ma_str]]
# For test set, the last `prediction_length[1] = 20` days will be used for benchmark:
ser_test_ma_bmk = [df_ibm_valid[:][ma_str], df_aapl_valid[:][ma_str], df_amzn_valid[:][ma_str], df_googl_valid[:][ma_str]]
# Fixing time index in test data:
ser_test_ma_bmk[0].index = df_ibm_test[-prediction_length[1]:].index
ser_test_ma_bmk[1].index = df_aapl_test[-prediction_length[1]:].index
ser_test_ma_bmk[2].index = df_amzn_test[-prediction_length[1]:].index
ser_test_ma_bmk[3].index = df_googl_test[-prediction_length[1]:].index
# # Metrics computation
# Computing metrics on the benchmark model will give me a good reference to evaluate the deep learning model after training.
# In the cas
# As can be presumed and seen from data prints, first `n` values of validation data are `NaN` so, it will be better just to exclude the first `n-1` values from the validation set to avoid misleading values to be catch in metrics evaluation.
from sklearn.metrics import mean_absolute_error, mean_squared_error, mean_absolute_percentage_error, r2_score
# ### Metrics performances on test data
# #### IBM stock
# +
#n = 10
#ma_str = str(n)+'_ac_ma'
# +
#ibm_ma_mse_loss = mean_squared_error(df_ibm_valid.iloc[n-1:]['Adj Close'], df_ibm_valid.iloc[n-1:][ma_str])
# +
#print(ibm_ma_mse_loss)
# -
n = 20
ma_str = str(n)+'_ac_ma'
# Mean Absolute Error
ibm_ma_mae_loss = mean_absolute_error(df_ibm_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[0])
print(ibm_ma_mae_loss)
# Root Mean Squared Error
ibm_ma_mse_loss = mean_squared_error(df_ibm_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[0], squared=False)
print(ibm_ma_mse_loss)
# Mean Absolute Percentage Error
ibm_ma_map_loss = mean_absolute_percentage_error(df_ibm_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[0])
print(ibm_ma_map_loss)
# R<sup>2</sup> score
ibm_ma_r2_score = r2_score(df_ibm_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[0])
print(ibm_ma_r2_score)
# +
#n = 50
#ma_str = str(n)+'_ac_ma'
# +
#ibm_ma_mse_loss = mean_squared_error(df_ibm_valid.iloc[n-1:]['Adj Close'], df_ibm_valid[n-1:][ma_str])
# +
#print(ibm_ma_mse_loss)
# -
# #### Apple Inc. stock
# +
#n = 10
#ma_str = str(n)+'_ac_ma'
# +
#aapl_ma_mse_loss = mean_squared_error(df_aapl_valid.iloc[n-1:]['Adj Close'], df_aapl_valid.iloc[n-1:][ma_str])
# +
#print(aapl_ma_mse_loss)
# -
n = 20
ma_str = str(n)+'_ac_ma'
# Mean Absolute Error
aapl_ma_mae_loss = mean_absolute_error(df_aapl_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[1])
print(aapl_ma_mae_loss)
# ##### Root Mean Squared Error
aapl_ma_rmse_loss = mean_squared_error(df_aapl_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[1], squared=False)
print(aapl_ma_rmse_loss)
# ##### Mean Absolute Percentage Error
aapl_ma_map_loss = mean_absolute_percentage_error(df_aapl_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[1])
print(aapl_ma_map_loss)
# R<sup>2</sup> score
aapl_ma_r2_score = r2_score(df_aapl_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[1])
print(aapl_ma_r2_score)
# +
#n = 50
#ma_str = str(n)+'_ac_ma'
# +
#aapl_ma_mse_loss = mean_squared_error(df_aapl_valid.iloc[n-1:]['Adj Close'], df_aapl_valid[n-1:][ma_str])
# +
#print(aapl_ma_mse_loss)
# -
# #### Amazon.com stock
# +
#n = 10
#ma_str = str(n)+'_ac_ma'
# +
#amzn_ma_mse_loss = mean_squared_error(df_amzn_valid.iloc[n-1:]['Adj Close'], df_amzn_valid.iloc[n-1:][ma_str])
# +
#print(amzn_ma_mse_loss)
# -
n = 20
ma_str = str(n)+'_ac_ma'
# Mean Absolute Error
amzn_ma_mae_loss = mean_absolute_error(df_amzn_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[2])
print(amzn_ma_mae_loss)
# Root Mean Squared Error
amzn_ma_rmse_loss = mean_squared_error(df_amzn_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[2], squared=False)
print(amzn_ma_rmse_loss)
# ##### Mean Absolute Percentage Error
amzn_ma_map_loss = mean_absolute_percentage_error(df_amzn_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[2])
print(amzn_ma_map_loss)
# R<sup>2</sup> score
amzn_ma_r2_score = r2_score(df_amzn_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[2])
print(amzn_ma_r2_score)
# +
#n = 50
#ma_str = str(n)+'_ac_ma'
# +
#amzn_ma_mse_loss = mean_squared_error(df_amzn_valid.iloc[n-1:]['Adj Close'], df_amzn_valid[n-1:][ma_str])
# +
#print(amzn_ma_mse_loss)
# -
# #### Alphabet Inc. stock
# +
#n = 10
#ma_str = str(n)+'_ac_ma'
# +
#googl_ma_mse_loss = mean_squared_error(df_googl_valid.iloc[n-1:]['Adj Close'], df_googl_valid.iloc[n-1:][ma_str])
# +
#print(googl_ma_mse_loss)
# -
n = 20
ma_str = str(n)+'_ac_ma'
# Mean Absolute Error
googl_ma_mae_loss = mean_absolute_error(df_googl_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[3])
print(googl_ma_mae_loss)
# Root Mean Squared Error
googl_ma_rmse_loss = mean_squared_error(df_googl_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[3], squared=False)
print(googl_ma_rmse_loss)
# ##### Mean Absolute Percentage Error
googl_ma_map_loss = mean_absolute_percentage_error(df_googl_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[3])
print(googl_ma_map_loss)
# R<sup>2</sup> score
googl_ma_r2_score = r2_score(df_googl_test.iloc[-prediction_length[1]:]['Adj Close'], ser_test_ma_bmk[3])
print(googl_ma_r2_score)
# ### Metrics performances on validation data
# #### IBM stock
# +
#n = 10
#ma_str = str(n)+'_ac_ma'
# +
#ibm_ma_mse_loss = mean_squared_error(df_ibm_valid.iloc[n-1:]['Adj Close'], df_ibm_valid.iloc[n-1:][ma_str])
# +
#print(ibm_ma_mse_loss)
# -
n = 20
ma_str = str(n)+'_ac_ma'
# Mean Absolute Error
ibm_ma_mae_loss = mean_absolute_error(df_ibm_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[0])
print(ibm_ma_mae_loss)
# Root Mean Squared Error
ibm_ma_mse_loss = mean_squared_error(df_ibm_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[0], squared=False)
print(ibm_ma_mse_loss)
# Mean Absolute Percentage Error
ibm_ma_map_loss = mean_absolute_percentage_error(df_ibm_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[0])
print(ibm_ma_map_loss)
# R<sup>2</sup> score
ibm_ma_r2_score = r2_score(df_ibm_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[0])
print(ibm_ma_r2_score)
# +
#n = 50
#ma_str = str(n)+'_ac_ma'
# +
#ibm_ma_mse_loss = mean_squared_error(df_ibm_valid.iloc[n-1:]['Adj Close'], df_ibm_valid[n-1:][ma_str])
# +
#print(ibm_ma_mse_loss)
# -
# #### Apple Inc. stock
# +
#n = 10
#ma_str = str(n)+'_ac_ma'
# +
#aapl_ma_mse_loss = mean_squared_error(df_aapl_valid.iloc[n-1:]['Adj Close'], df_aapl_valid.iloc[n-1:][ma_str])
# +
#print(aapl_ma_mse_loss)
# -
n = 20
ma_str = str(n)+'_ac_ma'
# Mean Absolute Error
aapl_ma_mae_loss = mean_absolute_error(df_aapl_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[1])
print(aapl_ma_mae_loss)
# ##### Root Mean Squared Error
aapl_ma_rmse_loss = mean_squared_error(df_aapl_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[1], squared=False)
print(aapl_ma_rmse_loss)
# ##### Mean Absolute Percentage Error
aapl_ma_map_loss = mean_absolute_percentage_error(df_aapl_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[1])
print(aapl_ma_map_loss)
# R<sup>2</sup> score
aapl_ma_r2_score = r2_score(df_aapl_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[1])
print(aapl_ma_r2_score)
# +
#n = 50
#ma_str = str(n)+'_ac_ma'
# +
#aapl_ma_mse_loss = mean_squared_error(df_aapl_valid.iloc[n-1:]['Adj Close'], df_aapl_valid[n-1:][ma_str])
# +
#print(aapl_ma_mse_loss)
# -
# #### Amazon.com stock
# +
#n = 10
#ma_str = str(n)+'_ac_ma'
# +
#amzn_ma_mse_loss = mean_squared_error(df_amzn_valid.iloc[n-1:]['Adj Close'], df_amzn_valid.iloc[n-1:][ma_str])
# +
#print(amzn_ma_mse_loss)
# -
n = 20
ma_str = str(n)+'_ac_ma'
# Mean Absolute Error
amzn_ma_mae_loss = mean_absolute_error(df_amzn_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[2])
print(amzn_ma_mae_loss)
# Root Mean Squared Error
amzn_ma_rmse_loss = mean_squared_error(df_amzn_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[2], squared=False)
print(amzn_ma_rmse_loss)
# ##### Mean Absolute Percentage Error
amzn_ma_map_loss = mean_absolute_percentage_error(df_amzn_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[2])
print(amzn_ma_map_loss)
# R<sup>2</sup> score
amzn_ma_r2_score = r2_score(df_amzn_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[2])
print(amzn_ma_r2_score)
# +
#n = 50
#ma_str = str(n)+'_ac_ma'
# +
#amzn_ma_mse_loss = mean_squared_error(df_amzn_valid.iloc[n-1:]['Adj Close'], df_amzn_valid[n-1:][ma_str])
# +
#print(amzn_ma_mse_loss)
# -
# #### Alphabet Inc. stock
# +
#n = 10
#ma_str = str(n)+'_ac_ma'
# +
#googl_ma_mse_loss = mean_squared_error(df_googl_valid.iloc[n-1:]['Adj Close'], df_googl_valid.iloc[n-1:][ma_str])
# +
#print(googl_ma_mse_loss)
# -
n = 20
ma_str = str(n)+'_ac_ma'
# Mean Absolute Error
googl_ma_mae_loss = mean_absolute_error(df_googl_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[3])
print(googl_ma_mae_loss)
# Root Mean Squared Error
googl_ma_rmse_loss = mean_squared_error(df_googl_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[3], squared=False)
print(googl_ma_rmse_loss)
# ##### Mean Absolute Percentage Error
googl_ma_map_loss = mean_absolute_percentage_error(df_googl_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[3])
print(googl_ma_map_loss)
# R<sup>2</sup> score
googl_ma_r2_score = r2_score(df_googl_valid.iloc[:]['Adj Close'], ser_valid_ma_bmk[3])
print(googl_ma_r2_score)
# +
#n = 50
#ma_str = str(n)+'_ac_ma'
# +
#googl_ma_mse_loss = mean_squared_error(df_googl_valid.iloc[n-1:]['Adj Close'], df_googl_valid[n-1:][ma_str])
# +
#print(googl_ma_mse_loss)
# -
# Volatility
print(volatility(df_ibm_valid['Adj Close'], n))
print(volatility(df_aapl_valid['Adj Close'], n))
print(volatility(df_amzn_valid['Adj Close'], n))
print(volatility(df_googl_valid['Adj Close'], n))
# As expected, loss augments as we observe moving average on larger windows.
# Also, we can observe that Amazon.com and Alphabet Inc. have greater losses, that also corresponds to higher volatility.
| 2.BenchmarkModel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import anndata
import numpy as np
import pandas as pd
import importlib
import sys
sys.path.append("..")
import autogenes
# -
data = pd.read_csv('../datasets/GSE75748_bulk_data.csv',index_col='index')
data = data.T.iloc[:,:20].values
ag = autogenes.AutoGeneS(data)
ag.run()
ag.run(ngen=5,seed=2,population_size=50,offspring_size=50)
ag.run(ngen=5,seed=2)
ag.run(ngen=15,seed=2,crossover_pb=0,mutation_pb=0)
ag.run(ngen=5,seed=2,population_size=2,offspring_size=2)
ag.run(ngen=10,seed=2,ind_standard_pb=0.01)
ag.run(ngen=10,seed=2,ind_standard_pb=0.7)
ag.run(ngen=10,seed=2,mutate_flip_pb=0.1)
ag.run(ngen=10,seed=2,mutate_flip_pb=0.9)
# ## Compare the effect of crossover_thres
ag.run(ngen=20,seed=2)
ag.run(ngen=20,seed=2,crossover_thres=100)
| tests_jupyter/genetic_algorithm_parameters.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
import pandas as pd
import numpy as np
import pickle
from collections import Counter
# %matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
# -
# Defualt image size
import matplotlib.pylab as pylab
pylab.rcParams['figure.figsize'] = 16, 9
# %%time
stations_dict = pickle.load(open('stations_dict.p', "rb"))
stations_latlng = pickle.load(open('stations_latlng.p', "rb"))
#df = pd.read_feather('2017_all')
df = pd.read_feather('df_')
#df = pd.read_feather('df_train')
df.shape
df.columns
df.tripduration.median()
df.tripduration.quantile([.25, .5, .75, .8])
# 3248, 3480 special stations
sp_stations = [3248, 3480, 3247, 3215, 3478]
idx = df[(df['start station id'].isin(sp_stations))
| (df['end station id'].isin(sp_stations))].index
df.drop(idx,0,inplace=True)
df.shape
df.head(3)
df.tail(3)
# 2% of the trips ends up in the same station, ignore these for our purpose
print(df[df['start station id'] == df['end station id']].shape)
df = df[df['start station id'] != df['end station id']]
df.tripduration.quantile([0, .1, .25, .5, .75, .99, .999, .9999])
# only look those trips are less or equal to one hour
df = df[df.tripduration <= 3600]
print(df.shape)
d = df.tripduration
sns.distplot(d, bins = 100);
sns.distplot(np.log(d), bins = 100);
del d
Counter(df.usertype)
# %%time
df['date'] = df['starttime'].apply(lambda x: x.date())
# %%time
S = set(df['date'])
d = dict()
for s in S:
d[s] = s.weekday() + 1
df['weekday'] = df['date'].map(d)
sns.barplot(x="weekday", y="tripduration", data=df.sample(500000))
sns.barplot(x="weekday", y="tripduration", data=df[df.usertype == 'Customer'].sample(100000))
sns.barplot(x="weekday", y="tripduration", data=df[df.usertype == 'Subscriber'].sample(100000))
df.columns
# +
tmp.head()
# +
# # number of trips vs. dow
tmp = df.groupby(['weekday','weekend'])\
.agg({'tripduration': 'size',
'date': lambda x: x.nunique()}).reset_index()
tmp['avg_trip_num'] = tmp['tripduration']/tmp['date']
sns.barplot(x="weekday", y="avg_trip_num", hue='weekend', data=tmp)
# -
# consumer
# number of trips vs. dow
tmp = df[df.usertype == 'Customer'].groupby(['weekday']).tripduration.size().reset_index()
sns.regplot(x="weekday", y="tripduration", data=tmp,
scatter_kws={"s": 50},
order=2, ci=None, truncate=True, fit_reg=False)
# %%time
df['weekend'] = df['weekday'].map(lambda x: 0 if x < 6 else 1)
S = set(df.date)
d = dict()
for s in S:
d[s] = s.month
df['month'] = df['date'].map(d)
# number of trips vs. hours
tmp = df.groupby(['starthour']).tripduration.size().reset_index()
display(tmp)
sns.regplot(x="starthour", y="tripduration", data=tmp,
scatter_kws={"s": 50}, ci=None, fit_reg=False);
def hour_min(time):
t = time.split(':')
return int(t[0])*100 + int(t[1])/60*100
# %%time
df['time'] = df['starttime'].astype(str).apply(lambda x: x[11:])
df['time'] = df['time'].map(lambda x: hour_min(x))
# +
# # number of trips vs. dow
tmp = df.groupby(['time'])\
.agg({'tripduration': 'size',
'date': lambda x: x.nunique()}).reset_index()
tmp['avg_trip_num'] = tmp['tripduration']/tmp['date']
s = sns.regplot(x="time", y="avg_trip_num", data=tmp,
scatter_kws={"s": 10}, ci=None, fit_reg=False);
axes = s.axes
axes.set_xlim(0,2400)
# -
# Customer
# number of trips vs. HH%MM
tmp = df[df.usertype == 'Customer'].groupby(['time']).tripduration.size().reset_index()
sns.regplot(x="time", y="tripduration", data=tmp,
scatter_kws={"s": 10}, ci=None, fit_reg=False);
# Customer
# number of trips vs. HH%MM
tmp = df[df.usertype == 'Subscriber'].groupby(['time']).tripduration.size().reset_index()
sns.regplot(x="time", y="tripduration", data=tmp,
scatter_kws={"s": 10}, ci=None, fit_reg=False);
# %%time
plt.figure(figsize=(24,16))
sns.barplot(x="starthour", y="tripduration",
data=df[(df.usertype == 'Subscriber') & (df.weekend == 0)].sample(300000))
plt.figure(figsize=(24,16))
sns.barplot(x="starthour", y="tripduration",
data=df[(df.usertype == 'Customer') & (df.weekend == 1)].sample(300000))
tmp = df.groupby(['month', 'usertype']).tripduration.size().reset_index()
plt.figure(figsize=(24,13.5))
sns.barplot(x="month", y="tripduration", hue="usertype",
data=tmp);
# +
# number of trips vs. day
tmp = df[df['month']==8].groupby(['date', 'usertype']).tripduration.size().reset_index()
tmp['date'] = tmp['date'].apply(lambda x: str(x)[-2:])
plt.figure(figsize=(24,13.5))
sns.barplot(x="date", y="tripduration", hue="usertype",
data=tmp);
# -
from datetime import datetime
def display_all(df):
"""
display more than 20 rows/cols
"""
with pd.option_context("display.max_rows", 1000):
with pd.option_context("display.max_columns", 1000):
display(df)
nyc_temp = pd.read_csv('nyc_temp_2017.csv')
nyc_temp['2017'] = nyc_temp['2017'].apply(lambda x: datetime.strptime(x, "%Y-%m-%d").date())
nyc_temp.columns = ['date', 'Temp_high', 'Temp_avg', 'Temp_low', 'Precip', 'Rain', 'Snow', 'Fog']
nyc_temp.sample(5)
# %%time
df = pd.merge(df, nyc_temp, 'left', on='date')
df.Precip.quantile(np.clip(np.arange(.7, 1., .05), 0, 1))
# +
df['rain_vol'] = 0
# v light, medium, heavy
df.loc[df['Precip'] >= 0.001, 'rain_vol'] = 1
df.loc[df['Precip'] >= 0.03, 'rain_vol'] = 2
df.loc[df['Precip'] >= 0.2, 'rain_vol'] = 3
df['temp_level'] = 0
df.loc[df['Temp_high'] >= 56, 'temp_level'] = 1
df.loc[df['Temp_high'] >= 67, 'temp_level'] = 2
df.loc[df['Temp_high'] >= 76, 'temp_level'] = 3
df.loc[df['Temp_high'] >= 83, 'temp_level'] = 4
# +
tmp = df[df.usertype == 'Subscriber']\
.groupby(['temp_level','rain_vol'])\
.agg({'tripduration': 'size',
'date': lambda x: x.nunique()}).reset_index()
tmp['avg_trip_num'] = tmp['tripduration']/tmp['date']
g = sns.barplot(x="temp_level",
y="avg_trip_num",
data=tmp,
hue="rain_vol",
palette=sns.cubehelix_palette(8, start=.9, rot=-.75))
# scatter_kws={"s": 10}, ci=None, fit_reg=False);
g.figure.set_size_inches(16, 9)
# +
tmp = df[df.usertype == 'Subscriber']\
.groupby(['temp_level','Rain'])\
.agg({'tripduration': 'size',
'date': lambda x: x.nunique()}).reset_index()
tmp['avg_trip_num'] = tmp['tripduration']/tmp['date']
g = sns.barplot(x="temp_level", y="avg_trip_num", data=tmp, hue="Rain",)
# scatter_kws={"s": 10}, ci=None, fit_reg=False);
g.figure.set_size_inches(16, 9)
# +
tmp = df[df.usertype == 'Subscriber']\
.groupby(['temp_level','Snow'])\
.agg({'tripduration': 'size',
'date': lambda x: x.nunique()}).reset_index()
tmp['avg_trip_num'] = tmp['tripduration']/tmp['date']
g = sns.barplot(x="temp_level", y="avg_trip_num", data=tmp, hue="Snow",)
# scatter_kws={"s": 10}, ci=None, fit_reg=False);
g.figure.set_size_inches(16, 9)
# +
tmp = df[df.usertype == 'Subscriber']\
.groupby(['temp_level','Fog'])\
.agg({'tripduration': 'size',
'date': lambda x: x.nunique()}).reset_index()
tmp['avg_trip_num'] = tmp['tripduration']/tmp['date']
g = sns.barplot(x="temp_level", y="avg_trip_num", data=tmp, hue="Fog",)
# scatter_kws={"s": 10}, ci=None, fit_reg=False);
g.figure.set_size_inches(16, 9)
# -
# %%time
df['lat1'] = df['start station id'].map(lambda x: stations_latlng[x][0])
df['lon1'] = df['start station id'].map(lambda x: stations_latlng[x][1])
df['lat2'] = df['end station id'].map(lambda x: stations_latlng[x][0])
df['lon2'] = df['end station id'].map(lambda x: stations_latlng[x][1])
from math import sin, cos, sqrt, atan2, radians
def manhattan_distance(latlon1, latlon2):
R = 6371
lat1 = radians(latlon1[0])
lon1 = radians(latlon1[1])
lat2 = radians(latlon2[0])
lon2 = radians(latlon2[1])
dlon = lon2 - lon1
dlat = lat2 - lat1
a1 = sin(dlat / 2)**2
c1 = 2 * atan2(sqrt(a1), sqrt(1 - a1))
d1 = R * c1
a2 = sin(dlon / 2)**2
c2 = 2 * atan2(sqrt(a2), sqrt(1 - a2))
d2 = R * c2
return d1+d2
# +
d1 = stations_latlng[523]
d1
d2 = stations_latlng[428]
d2
a = abs(d1[0]-d2[0])
b = abs(d1[1]-d2[1])
(a+b)*111.195
# -
manhattan_distance(d1, d2)
d1
d2
# %%time
tmp = df.groupby(['start station id', 'end station id']).size().reset_index()
tmp.columns = ['start station id', 'end station id', 'size']
tmp = tmp.sort_values('size', ascending=False).reset_index()
# +
# %%time
tmp['lat1'] = tmp['start station id'].map(lambda x: stations_latlng[x][0])
tmp['lon1'] = tmp['start station id'].map(lambda x: stations_latlng[x][1])
tmp['lat2'] = tmp['end station id'].map(lambda x: stations_latlng[x][0])
tmp['lon2'] = tmp['end station id'].map(lambda x: stations_latlng[x][1])
lat2 = tmp['lat2'].values
lon2 = tmp['lon2'].values
# -
plt.figure(figsize = (10,10))
plt.plot(lon2,lat2,'.', alpha = 0.8, markersize = 0.1)
plt.show()
def latlon2pos(lat, lon, size=240):
return int(round((40.84-lat)*size/240*1000-1)), int(round((lon+74.12)*size/240*1000-1))
# +
# %%time
tmp = df.groupby(['start station id']).size().reset_index()
tmp.columns = ['start station id', 'size']
tmp = tmp.sort_values('size', ascending=False).reset_index()
tmp['lat1'] = tmp['start station id'].map(lambda x: stations_latlng[x][0])
tmp['lon1'] = tmp['start station id'].map(lambda x: stations_latlng[x][1])
print(tmp.shape)
# show the log density of pickup and dropoff locations
s = 200
imageSize = (s,s)
locationDensityImage = np.zeros(imageSize)
for i in range(len(tmp)):
t = tmp.loc[i]
locationDensityImage[latlon2pos(t['lat1'], t['lon1'], s)] += t['size']#np.log1p(t['size'])
fig, ax = plt.subplots(nrows=1,ncols=1,figsize=(12,12))
ax.imshow(np.log1p(locationDensityImage), cmap='hot')
ax.set_axis_off()
# +
# %%time
tmp = df.groupby(['end station id']).size().reset_index()
tmp.columns = ['end station id', 'size']
tmp = tmp.sort_values('size', ascending=False).reset_index()
tmp['lat1'] = tmp['end station id'].map(lambda x: stations_latlng[x][0])
tmp['lon1'] = tmp['end station id'].map(lambda x: stations_latlng[x][1])
print(tmp.shape)
# show the log density of pickup and dropoff locations
s = 200
imageSize = (s,s)
locationDensityImage1 = np.zeros(imageSize)
for i in range(len(tmp)):
t = tmp.loc[i]
locationDensityImage1[latlon2pos(t['lat1'], t['lon1'], s)] += t['size']#np.log1p(t['size'])
fig, ax = plt.subplots(nrows=1,ncols=1,figsize=(12,12))
ax.imshow(np.log1p(locationDensityImage1), cmap='hot')
ax.set_axis_off()
# -
from sklearn.cluster import KMeans
# +
# # %%time
# tmp = df[['start station id']].sample(200000)
# loc_df = pd.DataFrame()
# loc_df['longitude'] = tmp['start station id'].map(lambda x: stations_latlng[x][1])
# loc_df['latitude'] = tmp['start station id'].map(lambda x: stations_latlng[x][0])
# Ks = range(5, 50)
# km = [KMeans(n_clusters=i) for i in Ks]
# score = [km[i].fit(loc_df).score(loc_df) for i in range(len(km))]
# score = [abs(i) for i in score]
# -
plt.plot((score))
# +
# %%time
tmp = df[['start station id']].sample(200000)
loc_df = pd.DataFrame()
loc_df['longitude'] = tmp['start station id'].map(lambda x: stations_latlng[x][1])
loc_df['latitude'] = tmp['start station id'].map(lambda x: stations_latlng[x][0])
kmeans = KMeans(n_clusters=16, random_state=2, n_init = 10).fit(loc_df)
loc_df['label'] = kmeans.labels_
plt.figure(figsize = (10,10))
for label in loc_df.label.unique():
plt.plot(loc_df.longitude[loc_df.label == label],loc_df.latitude[loc_df.label == label],'.', alpha = 0.3, markersize = 1)
plt.title('Clusters of New York (and New Jersey)')
plt.show()
# -
fig,ax = plt.subplots(figsize = (10,10))
for label in loc_df.label.unique():
ax.plot(loc_df.longitude[loc_df.label == label],loc_df.latitude[loc_df.label == label],'.', alpha = 0.4, markersize = 0.1, color = 'gray')
ax.plot(kmeans.cluster_centers_[label,0],kmeans.cluster_centers_[label,1],'o', color = 'r')
ax.annotate(label, (kmeans.cluster_centers_[label,0],kmeans.cluster_centers_[label,1]), color = 'b', fontsize = 20)
ax.set_title('Cluster Centers')
plt.show()
# %%time
df['start_cluster'] = kmeans.predict(df[['lon1','lat1']])
df['end_cluster'] = kmeans.predict(df[['lon2','lat2']])
clusters = pd.DataFrame()
clusters['x'] = kmeans.cluster_centers_[:,0]
clusters['y'] = kmeans.cluster_centers_[:,1]
clusters['label'] = range(len(clusters))
loc_df = loc_df.sample(5000)
import os
from matplotlib.pyplot import *
import matplotlib.pyplot as plt
from matplotlib import animation
from sklearn.cluster import KMeans
from IPython.display import HTML
from subprocess import check_output
import io
import base64
# +
# %%time
fig, ax = plt.subplots(1, 1, figsize = (10,10))
df_ = df.sample(5000000)
def animate(hour):
ax.clear()
ax.set_title('Absolute Traffic - Hour ' + str(hour))
plt.figure(figsize = (10,10));
for label in loc_df.label.unique():
ax.plot(loc_df.longitude[loc_df.label == label],loc_df.latitude[loc_df.label == label],'.', alpha = 1, markersize = 2, color = 'gray');
ax.plot(kmeans.cluster_centers_[label,0],kmeans.cluster_centers_[label,1],'o', color = 'r');
for label in clusters.label:
for dest_label in clusters.label:
num_of_rides = len(df_[(df_.start_cluster == label) & (df_.end_cluster == dest_label) & (df_.starthour == hour)])
dist_x = clusters.x[clusters.label == label].values[0] - clusters.x[clusters.label == dest_label].values[0]
dist_y = clusters.y[clusters.label == label].values[0] - clusters.y[clusters.label == dest_label].values[0]
pct = np.true_divide(num_of_rides,len(df_))
arr = Arrow(clusters.x[clusters.label == label].values, clusters.y[clusters.label == label].values, -dist_x, -dist_y, edgecolor='white', width = 15*pct)
ax.add_patch(arr)
arr.set_facecolor('g')
ani = animation.FuncAnimation(fig,animate,sorted(df.starthour.unique()), interval = 1000);
plt.close();
ani.save('Absolute.gif', writer='imagemagick', fps=2);
filename = 'Absolute.gif'
video = io.open(filename, 'r+b').read();
encoded = base64.b64encode(video);
HTML(data='''<img src="data:image/gif;base64,{0}" type="gif" />'''.format(encoded.decode('ascii')));
# +
# %%time
fig, ax = plt.subplots(1, 1, figsize = (10,10))
def animate(hour):
ax.clear()
ax.set_title('Relative Traffic - Hour ' + str(hour))
plt.figure(figsize = (10,10))
for label in loc_df.label.unique():
ax.plot(loc_df.longitude[loc_df.label == label],loc_df.latitude[loc_df.label == label],'.', alpha = 1, markersize = 2, color = 'gray')
ax.plot(kmeans.cluster_centers_[label,0],kmeans.cluster_centers_[label,1],'o', color = 'r')
for label in clusters.label:
for dest_label in clusters.label:
num_of_rides = len(df_[(df_.start_cluster == label) & (df_.end_cluster == dest_label) & (df_.starthour == hour)])
dist_x = clusters.x[clusters.label == label].values[0] - clusters.x[clusters.label == dest_label].values[0]
dist_y = clusters.y[clusters.label == label].values[0] - clusters.y[clusters.label == dest_label].values[0]
pct = np.true_divide(num_of_rides,len(df_[df_.starthour == hour]))
arr = Arrow(clusters.x[clusters.label == label].values, clusters.y[clusters.label == label].values, -dist_x, -dist_y, edgecolor='white', width = pct)
ax.add_patch(arr)
arr.set_facecolor('g')
ani = animation.FuncAnimation(fig,animate,sorted(df_.starthour.unique()), interval = 1000)
plt.close()
ani.save('Relative.gif', writer='imagemagick', fps=2)
filename = 'Relative.gif'
video = io.open(filename, 'r+b').read()
encoded = base64.b64encode(video)
HTML(data='''<img src="data:image/gif;base64,{0}" type="gif" />'''.format(encoded.decode('ascii')))
# -
df.tripduration.quantile([0, .25, .5, .75, 1.])
df.to_feather('df_')
def col_encode(col):
"""Encodes a pandas column with continous ids.
"""
uniq = np.unique(col)
name2idx = {o:i for i,o in enumerate(uniq)}
return name2idx#, np.array([name2idx[x] for x in col]), len(uniq)
col_encode(df['usertype'])
# %%time
df['user_enc'] = df['usertype'].map(col_encode(df['usertype']))
display_all(df.head(5))
# %%time
# naive distance
df['est_dist'] = abs(df['lat1'] - df['lat2']) + abs(df['lon1'] - df['lon2'])
df['est_dist'] = df['est_dist'] * 111195
# %%time
d = df.est_dist
sns.distplot(d, bins = 50);
del d
np.array(df.est_dist).reshape(1, -1)
# %%time
for i in ['starttime', 'stoptime', 'bikeid', 'usertype']:
try:
df.drop([i], 1, inplace=True)
except:
pass
df.est_dist.quantile([.5, .95, .97, .98, .99, 1.])
display_all(df.sample(5))
df['speed'] = df.est_dist/df.tripduration
df.speed.quantile([0, .1, .2, .3, .4, .5, .6, .7 ,.8, .9, 1.])
df.speed.quantile([.9, .92, .94 ,.96, .98, .99, .995, 1.])
idx = df[df.speed > 10].index
df.drop(idx, 0 ,inplace=True)
df = df.reset_index()
# %%time
d = df.speed
sns.distplot(d, bins = 100);
del d
# %%time
for i in ['index', 'lat1', 'lon1', 'lat2', 'lon2', 'time', 'speed']:
try:
df.drop([i], 1, inplace=True)
except:
pass
display_all(df.head())
date_temp = df.groupby(['month']).Temp_high.mean().reset_index()
date_temp['month'] = date_temp['month']-1
date_temp.columns = ['month', 'temp']
# +
tmp = df.groupby(['month']).tripduration.size().reset_index()
#tmp = pd.merge(tmp, date_temp, 'left', 'month')
fig, ax = plt.subplots(figsize=(24,13.5))
ax2 = ax.twinx()
sns.barplot(x="month", y="tripduration", data=tmp, color="#fff89e", ax=ax);
sns.regplot(x="month", y="temp", data=date_temp, ax=ax2, fit_reg=False);
ax.set_ylim(0, None)
ax2.set_ylim(32, 90)
plt.title('Trip numbers in each month along average temperature', fontsize=20)
plt.show()
# -
tmp = df[['date', 'Temp_high']].groupby(['date']).first().reset_index()
tmp['diff'] = 0
tmp.loc[1:, 'diff'] = np.diff(tmp.Temp_high)
tmp.head()
temp_d = dict(zip(tmp['date'], tmp['diff']))
# %%time
df['temp_diff'] = df['date'].map(temp_d)
df['est_dist'] = df['est_dist'].astype(int)
# +
tmp = df[df.month==2].groupby(['date']).tripduration.size().reset_index()
tmp['date'] = tmp['date'].map(col_encode(tmp['date']))+1
fig, ax = plt.subplots(figsize=(24,13.5))
ax2 = ax.twinx()
sns.barplot(x="date", y="tripduration", data=tmp, color="#fff89e", ax=ax);
ax.set_ylim(0, None)
ax2.set_ylim(32, 90)
#plt.title('Trip numbers in each month along average temperature', fontsize=20)
plt.show()
# -
display_all(df.sample(5))
tmp = pd.read_csv('nyc_temp_2017.csv')
tmp['2017'] = pd.to_datetime(tmp['2017'])
tmp.columns = ['date', 'Temp_high', 'Temp_avg',
'Temp_low', 'Precip', 'Rain',
'Snow', 'Fog', 'off_work',
'snow_plus_1']
tmp.sample(10)
tmp = tmp[['date', 'off_work', 'snow_plus_1']]
tmp.sample(3)
# %%time
df = pd.merge(df, tmp, 'left', on='date')
df.sample(5)
df.to_feather('df_train')
| notebook/data_EDA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Linear Regression - Overview
# ==================================
# ***
# ### How can I make predictions about real-world quantities, like sales or life expectancy?
#
# Most often in real world applications we need to understand how one variable is determined by a number of others.
#
# For example:
#
# * How does sales volume change with changes in price. How is this affected by changes in the weather?
#
# * How does the amount of a drug absorbed vary with dosage and with body weight of patient? Does it depend on blood pressure?
#
# * How are the conversions on an ecommerce website affected by two different page titles in an A/B comparison?
#
# * How does the energy released by an earthquake vary with the depth of it's epicenter?
#
# * How is the interest rate charged on a loan affected by credit history and by loan amount?
#
# Answering questions like these, requires us to create a **model**.
#
# A model is a formula where one variable (response) varies depending on one or more independent variables (covariates). For the loan example, interest rate might depend on FICO score, state, loan amount, and loan duration amongst others.
#
# One of the simplest models we can create is a **Linear Model** where we start with the assumption that the dependent variable varies linearly with the independent variable(s).
#
# While this may appear simplistic, many real world problems can be modeled usefully in this way. Often data that don't appear to have a linear relationship can be transformed using simple mappings so that they do now show a linear relationship. This is very powerful and Linear Models, therefore, have wide applicability.
#
# They are one of the foundational tools of Data Science.
#
# Creating a Linear Model involves a technique known as **Linear Regression**. It's a tool you've most probably already used without knowing that's what it was called.
#
# ---
#
#
# #### Linear Regression in the high school physics lab
# Remember a typical physics lab experiment from high school? We had some input X (say force) which gave some output Y (say acceleration).
#
# You made a number of pairs of observations x, y and plotted them on graph paper.
# <img src="files/images/a1fig1_labexperiment.png" />
# Then you had to fit a straight line through the set of observations using a visual "best fit".
# <img src="images/a1fig2_labexperiment_withline.png" />
# And then you read off 'm' the slope, and 'b', the y-intercept from the graph, hoping it was close to the expected answer. By drawing the "best fit" line you were, in effect, visually estimating m and b without knowing it.
#
# You were doing informal Linear Regression. We're going to do this a little more formally. And then make it more sophisticated.
# ### Now for a bit of math
#
# Let's start with the basics.
#
# Remember the equation for a straight line from high school?
#
# $$Y = mX + b$$
#
# where $m$ is the slope and $b$ is the y-intercept.
#
# Very briefly and simplistically, Linear Regression is a class of techniques for
#
# **_Fitting a straight line to a set of data points_**.
#
# This could also be considered reverse engineering a formula from the data.
#
# We'll develop this idea starting from first principles and adding mathematical sophistication as we go along. But before that, you're probably curious what were the 'm' and 'b' values for this graph. We use modeling software to generate this for us and we get:
#
# ---
#
# <img src="images/a1fig3_labexperiment_slopeintercept.png" />
# ---
# We see two numbers, "Intercept" and "Slope".
# Independent of what software we use to do our linear regression for us, it will report these two numbers in one form or another.
# The "Intercept" here is the "b" in our equation.
# And the "Slope" is the slope of Y with respect to the independent variable.
#
# To summarize, we have a dataset (the observations) and a model (our guess for a formula that fits the data) and we have to figure out the parameters of the model (the coefficients m and b in our best fit line) so that the model fits the data the "best".
# We want to use our data to find coefficients for a formula so that the formula will fit the data the "best".
#
# As we continue, we'll actually run the modeling software and generate these numbers from real data. Here we just saw pictures of the results.
# ---
# ### Using the model for prediction
#
# Once you had your visual best fit line and had read off the m and b you probably said something to the effect:
#
# "The data follows a linear equation of the form Y = mX + b where m (slope)=(somenumber) and b (y intercept)=(someothernumber)"
#
# You may recall that the equation is not an exact representation because most probably your data points are not all in a perfectly straight line. So there is some error varying from one data point to the next data point. Your visual approach subjectively tried to minimize some intuitive "total error" over all the data.
#
# What you did was intuitive "Linear Regression". You estimated m and b by the "looks right to me" algorithm.
# We will start with this intuitive notion and rapidly bring some heavy machinery to bear that will allow us to solve pretty sophisticated problems.
#
# At this point your lab exercise may well ask you to approximate what Y will be when X is some number outside the range of your measurements.
# Then you use the equation above where m and b are now actual numbers say 2.1 and 0.3 respectively i.e the equation is Y = 2.1X + 0.3
#
# This equation is your "model"
#
# And you plug in an X to get a Y.
#
# This is where you are using your model to predict a value or, in other words, you are saying that I didn't use this value of X in my experiment and I don't have it in my data but I'd like to know what this value of X will map to on the Y axis.
#
# Based on my model Y = 2.1X + 0.3 if I had used this value in my experiment then I believe I would have got an output Y of approximately what the straight line suggests.
#
# You also want to be able to say "my error is expected to be (some number), so I believe the actual value will lie between Y-error and Y+error".
#
# When used like this we call the X variable the "predictor" as values of Y are **predicted** based one values of X, and the Y variable the "response".
# But before we do that let's take another trip back to the physics lab and peek over at the neighboring lab group's plots.
# We might see a different plot.
# So which one is "correct"?
#
#
# <img src="files/images/a1fig4_twolabexperiments.png" />
# ### A notion of total error
#
#
# Visually we can see that our plot (the first one) is the "better" one. But why?
# Because intuitively we feel that the line is closer to the points in the first one.
# So let's try to understand formally why that might be correct. Or not.
# Actually the graphs above were plotted by software that generated some points with random variation and then plotted a line through them.
#
# What the software did was compute a function called a "loss function", a measure of error. Then, it "tried out" multiple straight lines until it found one that minimized the "loss function" value for that choice -- then it read off the Intercept and X-slope for that line.
#
# Because this error estimation is an important part of our modeling we're going to take a more detailed look at it.
#
#
# We want to create a simple formula for the error or difference between the value of Y given by our straight line, and the actual value of Y from our data set. Unless our line happens to pass through a particular point, this error will be non-zero. It may be positive or negative. We take the square of this error (we can do other things like take the abs value, but here we take the square.....patience, all will be revealed) and then we add up such error terms for each data point to get the total error for this straight line and this data set.
#
# **Important**: for a different set of samples of the **very same** experiment we will get a different data set and possibly a different staright line and so almost certainly a different total error.
#
# The squared error we used is a very commonly used form of the total error previously know as "quadratic error". It also has the property that errors in the negative direction and positive direction are treated the same and this "quadratic error" or "square error" is always have a positive value.
#
# So for now we will use the "squared error" as our representation of error. [1]
#
# So Regression in general is any approach we might use to estimate the coefficients of a model using the data to estimate the coefficients by minimizing the "squared error". Statistical software uses sophisticated numerical techniques using multivariable calculus to minimize this error and give us estimated values for the coefficients.
#
# **Let's try this on some real data.**
#
# We're going to look at a data set of Loan data from [Lending Club](http://www.lendingclub.com), a peer lending web site.
# They have anonymized data on borrowers and loans that have been made. Loan data has many attributes and we'll explore the whole data set in a bit but for now we'll just look at how borrower FICO score affects interest rate charged.
#
# +
# %pylab inline
import pandas as pd
# we have to clean up the raw data set which we will do
# in the next lesson. But for now let's look at the cleaned up data.
# import the cleaned up dataset into a pandas data frame
df = pd.read_csv('../datasets/loanf.csv')
# extract FICO Score and Interest Rate and plot them
# FICO Score on x-axis, Interest Rate on y-axis
intrate = df['Interest.Rate']
fico = df['FICO.Score']
p = plot(fico,intrate,'o')
ax = gca()
xt = ax.set_xlabel('FICO Score')
yt = ax.set_ylabel('Interest Rate %')
# -
# Here we see a distinct downward linear trend where Interest Rate goes down with increasing FICO score. But we also see that for the same FICO score there is a range of Interest rates. This suggests that FICO by itself might not be enough to predict Interest Rate.
# ### Multivariate Linear Regression
#
# So the natural question that arises is what happens if Y depends on more than one variable.
# And this is where the power of mathematical generalization comes in. The same principle applies but in multiple dimensions. Not just two or three but much larger numbers. Twenty, thirty or even hundred independent variables are not out of question if we want to model real world data.
#
# But for now let's look at $Y$ as a function of two independent variables, $X_1$ and $X_2$, so
#
# $$ Y = a_0 + a_1X_1 + a_2X_2 $$
#
# Here $a_0$ is the Intercept term and $a_1, a_2$ are the coefficients of $X_1, X_2$, the independent variables respectively.
#
# So to look at a real data set with potentially multiple independent variables we're going to use the Lending Club data set in the next step.
# ---
#
# ## References
#
# [1] Squared Error <http://en.wikipedia.org/wiki/Residual_sum_of_squares>
from IPython.core.display import HTML
def css_styling():
styles = open("../styles/custom.css", "r").read()
return HTML(styles)
css_styling()
| notebooks/A1. Linear Regression - Overview.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Accessing Forest Inventory and Analysis data with the Planetary Computer STAC API
#
# This notebook demonstrates accessing [Forest Inventory and Analysis](https://planetarycomputer.microsoft.com/dataset/fia) (FIA) data from the Planetary Computer STAC API.
#
# The Forest Inventory and Analysis collection contains many tables, and each STAC table corresponds to one STAC item in the [FIA collection](http://planetarycomputer.microsoft.com/api/stac/v1/collections/fia). In this example, we'll use a few of the tables to estimate the total amount of aboveground carbon, in pounds, per US county.
#
# This example builds on the [plot estimation](https://rfia.netlify.app/courses/plt_est/) example from the [rfia](https://rfia.netlify.app/) package.
from cartopy import crs as ccrs
from dask_gateway import GatewayCluster
import dask_geopandas
import dask.dataframe as dd
import geopandas
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import planetary_computer
# The `tree` table below is relatively large, so we'll process it in parallel on a Dask cluster. The example will still run without a cluster, it will just take longer.
cluster = GatewayCluster()
cluster.scale(16)
client = cluster.get_client()
cluster
# ### Data Access
#
# We'll use three datsaets
#
# * `tree`: Information on each tree 1″ in diameter or larger, linked to `plot` and `cond`.
# * `plot`: Information relevant to each 1-acre field plot where the samples were collected.
# * `cond`: Information on the discrete combination of landscape attributes that define the plot condition.
#
# All of these are available in Azure Blob Storage as parquet datasets that can be read, for example, by `dask.dataframe`.
storage_options = {"account_name": "cpdataeuwest"}
plot = dd.read_parquet(
"az:///cpdata/raw/fia/plot.parquet",
columns=["CN", "STATECD", "COUNTYCD"],
storage_options=storage_options,
)
cond = dd.read_parquet(
"az:///cpdata/raw/fia/cond.parquet",
columns=["PLT_CN", "CONDID"],
storage_options=storage_options,
)
tree = dd.read_parquet(
"az:///cpdata/raw/fia/tree.parquet",
columns=["PLT_CN", "CONDID", "TREE", "DRYBIO_AG", "CARBON_AG", "TPA_UNADJ"],
storage_options=storage_options,
)
# ### Join the datasets
#
# The three datasets can be joined on their various keys. Since `tree` is relatively large, we'll join the other (smaller, in-memory) dataframes to it.
df = (
tree.merge(cond.compute(), on=["PLT_CN", "CONDID"])
.merge(plot.assign(PLT_CN=plot.CN).compute(), on="PLT_CN")
.assign(
bio=lambda df: df.DRYBIO_AG * df.TPA_UNADJ / 2000,
carbon=lambda df: df.CARBON_AG * df.TPA_UNADJ / 2000,
)
)
df
# ### Compute per-county summaries
#
# The `df` dataframe now includes the state and county FIPS codes, and the (adjusted) aboveground carbon and biomass. We'll group by the geographic boundaries and sum the aboveground carbon and biomass.
result = (
df.groupby(["STATECD", "COUNTYCD"])[["bio", "carbon"]]
.sum()
.compute()
.reset_index()
.assign(
STATE=lambda df: df["STATECD"].astype("string").str.pad(2, fillchar="0"),
COUNTY=lambda df: df["COUNTYCD"].astype("string").str.pad(3, fillchar="0"),
)
.drop(columns=["STATECD", "COUNTYCD"])
)
result.head()
# ### Plot the results
#
# Now we'll make a chloropleth for the results. We just need to join in the actual geographic boundaries of the datasets, which we can get with geopandas.
#
# Finally, we'll slice the data down to the continental United States (the dataset covers Hawaii, Alaska, and several other territories).
# +
# TODO: replace with STAC, sign
sas_token = planetary_computer.sas.get_token("<PASSWORD>", "us-census").token
counties = (
dask_geopandas.read_parquet(
"abfs://us-census/2020/cb_2020_us_county_500k.parquet",
storage_options={"account_name": "ai4edataeuwest", "credential": sas_token},
columns=["STATEFP", "COUNTYFP", "geometry"],
).rename(columns={"STATEFP": "STATE", "COUNTYFP": "COUNTY"})
).compute()
# -
gdf = geopandas.GeoDataFrame(pd.merge(result, counties, on=["STATE", "COUNTY"]))
df_conus = gdf.cx[-124.784:-66.951, 24.744:49.346]
df_conus.head()
# Finally, we'll plot the (log) of the estimated carbon stored above ground by the trees.
# +
crs = ccrs.LambertConformal()
fig, ax = plt.subplots(subplot_kw={"projection": crs}, figsize=(16, 9))
df_conus.assign(carbon=np.log(df_conus.carbon)).to_crs(crs.proj4_init).plot(
column="carbon",
cmap="Greens",
edgecolor="k",
scheme="natural_breaks",
k=8,
ax=ax,
linewidths=0.1,
legend=True,
)
# Shift the legend
bbox = ax.legend_.get_bbox_to_anchor().transformed(ax.transAxes.inverted())
bbox.x0 += 0.075
bbox.x1 += 0.075
bbox.y0 -= 0.4
bbox.y1 -= 0.4
ax.legend_.set_bbox_to_anchor(bbox)
ax.axis("off");
# -
# ### Next Steps
#
# Now that you've an introduction to the Forest Inventory and Analysis dataset, learn more with
#
# * The [Reading tabular data quickstart](https://planetarycomputer.microsoft.com/docs/quickstarts/reading-tabular-data/) for an introduction to tabular data on the Planeatry Computer
| datasets/fia/fia-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!--BOOK_INFORMATION-->
# <img align="left" style="padding-right:10px;" src="figures/PDSH-cover-small.png">
#
# *This notebook contains an excerpt from the [Python Data Science Handbook](http://shop.oreilly.com/product/0636920034919.do) by <NAME>; the content is available [on GitHub](https://github.com/jakevdp/PythonDataScienceHandbook).*
#
# *The text is released under the [CC-BY-NC-ND license](https://creativecommons.org/licenses/by-nc-nd/3.0/us/legalcode), and code is released under the [MIT license](https://opensource.org/licenses/MIT). If you find this content useful, please consider supporting the work by [buying the book](http://shop.oreilly.com/product/0636920034919.do)!*
# <!--NAVIGATION-->
# < [Density and Contour Plots](04.04-Density-and-Contour-Plots.ipynb) | [Contents](Index.ipynb) | [Customizing Plot Legends](04.06-Customizing-Legends.ipynb) >
#
# <a href="https://colab.research.google.com/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/04.05-Histograms-and-Binnings.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
# # Histograms, Binnings, and Density
# A simple histogram can be a great first step in understanding a dataset.
# Earlier, we saw a preview of Matplotlib's histogram function (see [Comparisons, Masks, and Boolean Logic](02.06-Boolean-Arrays-and-Masks.ipynb)), which creates a basic histogram in one line, once the normal boiler-plate imports are done:
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-white')
data = np.random.randn(1000)
# -
plt.hist(data);
# The ``hist()`` function has many options to tune both the calculation and the display;
# here's an example of a more customized histogram:
plt.hist(data, bins=30, normed=True, alpha=0.5,
histtype='stepfilled', color='steelblue',
edgecolor='none');
# The ``plt.hist`` docstring has more information on other customization options available.
# I find this combination of ``histtype='stepfilled'`` along with some transparency ``alpha`` to be very useful when comparing histograms of several distributions:
# +
x1 = np.random.normal(0, 0.8, 1000)
x2 = np.random.normal(-2, 1, 1000)
x3 = np.random.normal(3, 2, 1000)
kwargs = dict(histtype='stepfilled', alpha=0.3, normed=True, bins=40)
plt.hist(x1, **kwargs)
plt.hist(x2, **kwargs)
plt.hist(x3, **kwargs);
# -
# If you would like to simply compute the histogram (that is, count the number of points in a given bin) and not display it, the ``np.histogram()`` function is available:
counts, bin_edges = np.histogram(data, bins=5)
print(counts)
# ## Two-Dimensional Histograms and Binnings
#
# Just as we create histograms in one dimension by dividing the number-line into bins, we can also create histograms in two-dimensions by dividing points among two-dimensional bins.
# We'll take a brief look at several ways to do this here.
# We'll start by defining some data—an ``x`` and ``y`` array drawn from a multivariate Gaussian distribution:
mean = [0, 0]
cov = [[1, 1], [1, 2]]
x, y = np.random.multivariate_normal(mean, cov, 10000).T
# ### ``plt.hist2d``: Two-dimensional histogram
#
# One straightforward way to plot a two-dimensional histogram is to use Matplotlib's ``plt.hist2d`` function:
plt.hist2d(x, y, bins=30, cmap='Blues')
cb = plt.colorbar()
cb.set_label('counts in bin')
# Just as with ``plt.hist``, ``plt.hist2d`` has a number of extra options to fine-tune the plot and the binning, which are nicely outlined in the function docstring.
# Further, just as ``plt.hist`` has a counterpart in ``np.histogram``, ``plt.hist2d`` has a counterpart in ``np.histogram2d``, which can be used as follows:
counts, xedges, yedges = np.histogram2d(x, y, bins=30)
# For the generalization of this histogram binning in dimensions higher than two, see the ``np.histogramdd`` function.
# ### ``plt.hexbin``: Hexagonal binnings
#
# The two-dimensional histogram creates a tesselation of squares across the axes.
# Another natural shape for such a tesselation is the regular hexagon.
# For this purpose, Matplotlib provides the ``plt.hexbin`` routine, which will represents a two-dimensional dataset binned within a grid of hexagons:
plt.hexbin(x, y, gridsize=30, cmap='Blues')
cb = plt.colorbar(label='count in bin')
# ``plt.hexbin`` has a number of interesting options, including the ability to specify weights for each point, and to change the output in each bin to any NumPy aggregate (mean of weights, standard deviation of weights, etc.).
# ### Kernel density estimation
#
# Another common method of evaluating densities in multiple dimensions is *kernel density estimation* (KDE).
# This will be discussed more fully in [In-Depth: Kernel Density Estimation](05.13-Kernel-Density-Estimation.ipynb), but for now we'll simply mention that KDE can be thought of as a way to "smear out" the points in space and add up the result to obtain a smooth function.
# One extremely quick and simple KDE implementation exists in the ``scipy.stats`` package.
# Here is a quick example of using the KDE on this data:
# +
from scipy.stats import gaussian_kde
# fit an array of size [Ndim, Nsamples]
data = np.vstack([x, y])
kde = gaussian_kde(data)
# evaluate on a regular grid
xgrid = np.linspace(-3.5, 3.5, 40)
ygrid = np.linspace(-6, 6, 40)
Xgrid, Ygrid = np.meshgrid(xgrid, ygrid)
Z = kde.evaluate(np.vstack([Xgrid.ravel(), Ygrid.ravel()]))
# Plot the result as an image
plt.imshow(Z.reshape(Xgrid.shape),
origin='lower', aspect='auto',
extent=[-3.5, 3.5, -6, 6],
cmap='Blues')
cb = plt.colorbar()
cb.set_label("density")
# -
# KDE has a smoothing length that effectively slides the knob between detail and smoothness (one example of the ubiquitous bias–variance trade-off).
# The literature on choosing an appropriate smoothing length is vast: ``gaussian_kde`` uses a rule-of-thumb to attempt to find a nearly optimal smoothing length for the input data.
#
# Other KDE implementations are available within the SciPy ecosystem, each with its own strengths and weaknesses; see, for example, ``sklearn.neighbors.KernelDensity`` and ``statsmodels.nonparametric.kernel_density.KDEMultivariate``.
# For visualizations based on KDE, using Matplotlib tends to be overly verbose.
# The Seaborn library, discussed in [Visualization With Seaborn](04.14-Visualization-With-Seaborn.ipynb), provides a much more terse API for creating KDE-based visualizations.
# <!--NAVIGATION-->
# < [Density and Contour Plots](04.04-Density-and-Contour-Plots.ipynb) | [Contents](Index.ipynb) | [Customizing Plot Legends](04.06-Customizing-Legends.ipynb) >
#
# <a href="https://colab.research.google.com/github/jakevdp/PythonDataScienceHandbook/blob/master/notebooks/04.05-Histograms-and-Binnings.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
#
| PythonDataScienceHandbook/notebooks/04.05-Histograms-and-Binnings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # [Fourier Transforms With scipy.fft: Python Signal Processing (Real Python)](https://realpython.com/python-scipy-fft/)
#
# - Author: <NAME> [\[e-mail\]](mailto:'Israel%20Oliveira%20'<<EMAIL>>)
# %load_ext watermark
# +
import matplotlib.pyplot as plt
# Algumas configurações para o matplotlib.
# %matplotlib inline
from IPython.core.pylabtools import figsize
figsize(12, 8)
#pd.set_option("max_columns", None)
#pd.set_option("max_rows", None)
# -
# Run this cell before close.
# %watermark -d --iversion -b -r -g -m -v
# !cat /proc/cpuinfo |grep 'model name'|head -n 1 |sed -e 's/model\ name/CPU/'
# !free -h |cut -d'i' -f1 |grep -v total
# +
import numpy as np
from matplotlib import pyplot as plt
SAMPLE_RATE = 44100 # Hertz
DURATION = 5 # Seconds
def generate_sine_wave(freq, sample_rate, duration):
x = np.linspace(0, duration, sample_rate * duration, endpoint=False)
frequencies = x * freq
# 2pi because np.sin takes radians
y = np.sin((2 * np.pi) * frequencies)
return x, y
# Generate a 2 hertz sine wave that lasts for 5 seconds
x, y = generate_sine_wave(2, SAMPLE_RATE, DURATION)
plt.plot(x, y)
plt.show()
# +
_, nice_tone = generate_sine_wave(400, SAMPLE_RATE, DURATION)
_, noise_tone = generate_sine_wave(4000, SAMPLE_RATE, DURATION)
noise_tone = noise_tone * 0.3
mixed_tone = nice_tone + noise_tone
normalized_tone = np.int16((mixed_tone / mixed_tone.max()) * 32767)
plt.plot(normalized_tone[:1000])
plt.show()
# +
from scipy.io.wavfile import write
# Remember SAMPLE_RATE = 44100 Hz is our playback rate
write("mysinewave.wav", SAMPLE_RATE, normalized_tone)
# +
from scipy.fft import fft, fftfreq
# Number of samples in normalized_tone
N = SAMPLE_RATE * DURATION
yf = fft(normalized_tone)
xf = fftfreq(N, 1 / SAMPLE_RATE)
plt.plot(xf, np.abs(yf))
plt.show()
# -
yf = fft(normalized_tone)
xf = fftfreq(N, 1 / SAMPLE_RATE)
plt.plot(xf, np.abs(yf))
plt.show()
# +
yf = fft(normalized_tone)
xf = fftfreq(N, 1 / SAMPLE_RATE)
from scipy.fft import rfft, rfftfreq
# Note the extra 'r' at the front
yf = rfft(normalized_tone)
xf = rfftfreq(N, 1 / SAMPLE_RATE)
plt.plot(xf, np.abs(yf))
plt.show()
# +
# The maximum frequency is half the sample rate
points_per_freq = len(xf) / (SAMPLE_RATE / 2)
# Our target frequency is 4000 Hz
target_idx = int(points_per_freq * 4000)
yf[target_idx - 1 : target_idx + 2] = 0
plt.plot(xf, np.abs(yf))
plt.show()
# +
from scipy.fft import irfft
new_sig = irfft(yf)
plt.plot(new_sig[:1000])
plt.show()
# +
norm_new_sig = np.int16(new_sig * (32767 / new_sig.max()))
write("clean.wav", SAMPLE_RATE, norm_new_sig)
# -
| FFT/Real_Python_Tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # 04 - Full waveform inversion with Dask and Devito pickling
# ## Introduction
#
# Here, we revisit [04_dask.ipynb: Full Waveform Inversion with Devito and Dask](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/04_dask.ipynb), but with a twist: we now want to show that it is possible to use pickle to serialize (deserialize) a Devito object structure into (from) a byte stream. This is specially useful in our example as the geometry of all source experiments remains essentially the same; only the source location changes. In other words, we can convert a `solver` object (built on top of generic Devito objects) into a byte stream to store it. Later on, this byte stream can then be retrieved and de-serialized back to an instance of the original `solver` object by the dask workers, and then be populated with the correct geometry for the i-th source location. We can still benefit from the simplicity of the example and create **only one `solver`** object which can be used to both generate the observed data set and to compute the predicted data and gradient in the FWI process. Further examples of pickling can be found [here](https://github.com/devitocodes/devito/blob/master/tests/test_pickle.py).
# The tutorial roughly follows the structure of [04_dask.ipynb](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/04_dask.ipynb). Technical details about [Dask](https://dask.pydata.org/en/latest/#dask) and [scipy.optimize.minimize](https://docs.scipy.org/doc/scipy/reference/generated/scipy.optimize.minimize.html) will therefore treated only superficially.
# ## What is different from 04_dask.ipynb
#
# * **The big difference between [04_dask.ipynb](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/04_dask.ipynb) and this tutorial is that in the former is created a `solver` object for each source in both forward modeling and FWI gradient kernels. While here only one `solver` object is created and reused along all the optimization process. This is done through pickling and unpickling respectively.**
#
#
# * Another difference between the tutorials is that the in [04_dask.ipynb](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/04_dask.ipynb) is created a list with the observed shots, and then each observed shot record of the list is passed as parameter to a single-shot FWI objective function executed in parallel using the `submit()` method. Here, a single observed shot record along information of its source location is stored in a dictionary, which is saved into a pickle file. Later, dask workers retrieve the corresponding pickled data when computing the gradient for a single shot. The same applies for the `model` object in the optimization process. It is serialized each time the model's velocity is updated. Then, dask workers unpickle data from file back to `model` object.
#
#
# * Moreover, there is a difference in the way that the global functional-gradient is obtained. In [04_dask.ipynb](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/04_dask.ipynb) we had to wait for all computations to finish via `wait(futures)` and then we sum the function values and gradients from all workers. Here, it is defined a type `fg_pair` so that a reduce function `sum` can be used, such function takes all the futures given to it and after they are completed, combine them to get the estimate of the global functional-gradient.
# ## scipy.optimize.minimize
#
# As in [04_dask.ipynb](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/04_dask.ipynb), here we are going to focus on using L-BFGS via [scipy.optimize.minimize(method=’L-BFGS-B’)](https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html#optimize-minimize-lbfgsb)
#
# ```python
# scipy.optimize.minimize(fun, x0, args=(), method='L-BFGS-B', jac=None, bounds=None, tol=None, callback=None, options={'disp': None, 'maxls': 20, 'iprint': -1, 'gtol': 1e-05, 'eps': 1e-08, 'maxiter': 15000, 'ftol': 2.220446049250313e-09, 'maxcor': 10, 'maxfun': 15000})```
#
# The argument `fun` is a callable function that returns the misfit between the simulated and the observed data. If `jac` is a Boolean and is `True`, `fun` is assumed to return the gradient along with the objective function - as is our case when applying the adjoint-state method.
# ## Dask
#
# [Dask](https://dask.pydata.org/en/latest/#dask) is task-based parallelization framework for Python. It allows us to distribute our work among a collection of workers controlled by a central scheduler. Dask is [well-documented](https://docs.dask.org/en/latest/), flexible, an currently under active development.
#
# In the same way as in [04_dask.ipynb](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/04_dask.ipynb), we are going to use it here to parallelise the computation of the functional and gradient as this is the vast bulk of the computational expense of FWI and it is trivially parallel over data shots.
# ## Forward modeling
#
# We define the functions used for the forward modeling, as well as the other functions used in constructing and deconstructing Python/Devito objects to/from binary data as follows:
# +
#NBVAL_IGNORE_OUTPUT
# Set up inversion parameters.
param = {'t0': 0.,
'tn': 1000., # Simulation last 1 second (1000 ms)
'f0': 0.010, # Source peak frequency is 10Hz (0.010 kHz)
'nshots': 5, # Number of shots to create gradient from
'shape': (101, 101), # Number of grid points (nx, nz).
'spacing': (10., 10.), # Grid spacing in m. The domain size is now 1km by 1km.
'origin': (0, 0), # Need origin to define relative source and receiver locations.
'nbl': 40} # nbl thickness.
import numpy as np
import scipy
from scipy import signal, optimize
from devito import Grid
from distributed import Client, LocalCluster, wait
import cloudpickle as pickle
# Import acoustic solver, source and receiver modules.
from examples.seismic import Model, demo_model, AcquisitionGeometry, Receiver
from examples.seismic.acoustic import AcousticWaveSolver
from examples.seismic import AcquisitionGeometry
# Import convenience function for plotting results
from examples.seismic import plot_image
from examples.seismic import plot_shotrecord
def get_true_model():
''' Define the test phantom; in this case we are using
a simple circle so we can easily see what is going on.
'''
return demo_model('circle-isotropic', vp_circle=3.0, vp_background=2.5,
origin=param['origin'], shape=param['shape'],
spacing=param['spacing'], nbl=param['nbl'])
def get_initial_model():
'''The initial guess for the subsurface model.
'''
# Make sure both model are on the same grid
grid = get_true_model().grid
return demo_model('circle-isotropic', vp_circle=2.5, vp_background=2.5,
origin=param['origin'], shape=param['shape'],
spacing=param['spacing'], nbl=param['nbl'],
grid=grid)
def wrap_model(x, astype=None):
'''Wrap a flat array as a subsurface model.
'''
model = get_initial_model()
v_curr = 1.0/np.sqrt(x.reshape(model.shape))
if astype:
model.update('vp', v_curr.astype(astype).reshape(model.shape))
else:
model.update('vp', v_curr.reshape(model.shape))
return model
def load_model(filename):
""" Returns the current model. This is used by the
worker to get the current model.
"""
pkl = pickle.load(open(filename, "rb"))
return pkl['model']
def dump_model(filename, model):
''' Dump model to disk.
'''
pickle.dump({'model':model}, open(filename, "wb"))
def load_shot_data(shot_id, dt):
''' Load shot data from disk, resampling to the model time step.
'''
pkl = pickle.load(open("shot_%d.p"%shot_id, "rb"))
return pkl['geometry'], pkl['rec'].resample(dt)
def dump_shot_data(shot_id, rec, geometry):
''' Dump shot data to disk.
'''
pickle.dump({'rec':rec, 'geometry': geometry}, open('shot_%d.p'%shot_id, "wb"))
def generate_shotdata_i(param):
""" Inversion crime alert! Here the worker is creating the
'observed' data using the real model. For a real case
the worker would be reading seismic data from disk.
"""
# Reconstruct objects
with open("arguments.pkl", "rb") as cp_file:
cp = pickle.load(cp_file)
solver = cp['solver']
# source position changes according to the index
shot_id=param['shot_id']
solver.geometry.src_positions[0,:]=[20, shot_id*1000./(param['nshots']-1)]
true_d = solver.forward()[0]
dump_shot_data(shot_id, true_d.resample(4.0), solver.geometry.src_positions)
def generate_shotdata(solver):
# Pick devito objects (save on disk)
cp = {'solver': solver}
with open("arguments.pkl", "wb") as cp_file:
pickle.dump(cp, cp_file)
work = [dict(param) for i in range(param['nshots'])]
# synthetic data is generated here twice: serial(loop below) and parallel (via dask map functionality)
for i in range(param['nshots']):
work[i]['shot_id'] = i
generate_shotdata_i(work[i])
# Map worklist to cluster, We pass our function and the dictionary to the map() function of the client
# This returns a list of futures that represents each task
futures = c.map(generate_shotdata_i, work)
# Wait for all futures
wait(futures)
# +
#NBVAL_IGNORE_OUTPUT
from examples.seismic import plot_shotrecord
# Client setup
cluster = LocalCluster(n_workers=2, death_timeout=600)
c = Client(cluster)
# Generate shot data.
true_model = get_true_model()
# Source coords definition
src_coordinates = np.empty((1, len(param['shape'])))
# Number of receiver locations per shot.
nreceivers = 101
# Set up receiver data and geometry.
rec_coordinates = np.empty((nreceivers, len(param['shape'])))
rec_coordinates[:, 1] = np.linspace(param['spacing'][0], true_model.domain_size[0] - param['spacing'][0], num=nreceivers)
rec_coordinates[:, 0] = 980. # 20m from the right end
# Geometry
geometry = AcquisitionGeometry(true_model, rec_coordinates, src_coordinates,
param['t0'], param['tn'], src_type='Ricker',
f0=param['f0'])
# Set up solver
solver = AcousticWaveSolver(true_model, geometry, space_order=4)
generate_shotdata(solver)
# -
# ## Dask specifics
#
# Previously in [03_fwi.ipynb](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/03_fwi.ipynb), we defined a function to calculate the individual contribution to the functional and gradient for each shot, which was then used in a loop over all shots. However, when using distributed frameworks such as Dask we instead think in terms of creating a worklist which gets *mapped* onto the worker pool. The sum reduction is also performed in parallel. For now however we assume that the scipy.optimize.minimize itself is running on the *master* process; this is a reasonable simplification because the computational cost of calculating (f, g) far exceeds the other compute costs.
# Because we want to be able to use standard reduction operators such as sum on (f, g) we first define it as a type so that we can define the `__add__` (and `__radd__` method).
# Define a type to store the functional and gradient.
class fg_pair:
def __init__(self, f, g):
self.f = f
self.g = g
def __add__(self, other):
f = self.f + other.f
g = self.g + other.g
return fg_pair(f, g)
def __radd__(self, other):
if other == 0:
return self
else:
return self.__add__(other)
# ## Create operators for gradient based inversion
# To perform the inversion we are going to use [scipy.optimize.minimize(method=’L-BFGS-B’)](https://docs.scipy.org/doc/scipy/reference/optimize.minimize-lbfgsb.html#optimize-minimize-lbfgsb).
#
# First we define the functional, ```f```, and gradient, ```g```, operator (i.e. the function ```fun```) for a single shot of data. This is the work that is going to be performed by the worker on a unit of data.
# +
#NBVAL_IGNORE_OUTPUT
from devito import Function
# Create FWI gradient kernel for a single shot
def fwi_gradient_i(param):
# Load the current model and the shot data for this worker.
# Note, unlike the serial example the model is not passed in
# as an argument. Broadcasting large datasets is considered
# a programming anti-pattern and at the time of writing it
# it only worked reliably with Dask master. Therefore, the
# the model is communicated via a file.
model0 = load_model(param['model'])
dt = model0.critical_dt
nbl = model0.nbl
# Get src_position and data
src_positions, rec = load_shot_data(param['shot_id'], dt)
# Set up solver -- load the solver used above in the generation of the syntethic data.
with open("arguments.pkl", "rb") as cp_file:
cp = pickle.load(cp_file)
solver = cp['solver']
# Set attributes to solver
solver.geometry.src_positions=src_positions
solver.geometry.resample(dt)
# Compute simulated data and full forward wavefield u0
d, u0 = solver.forward(vp=model0.vp, dt=dt, save=True)[0:2]
# Compute the data misfit (residual) and objective function
residual = Receiver(name='rec', grid=model0.grid,
time_range=solver.geometry.time_axis,
coordinates=solver.geometry.rec_positions)
#residual.data[:] = d.data[:residual.shape[0], :] - rec.data[:residual.shape[0], :]
residual.data[:] = d.data[:] - rec.data[0:d.data.shape[0], :]
f = .5*np.linalg.norm(residual.data.flatten())**2
# Compute gradient using the adjoint-state method. Note, this
# backpropagates the data misfit through the model.
grad = Function(name="grad", grid=model0.grid)
solver.gradient(rec=residual, u=u0, vp=model0.vp, dt=dt, grad=grad)
# Copying here to avoid a (probably overzealous) destructor deleting
# the gradient before Dask has had a chance to communicate it.
g = np.array(grad.data[:])[nbl:-nbl, nbl:-nbl]
# return the objective functional and gradient.
return fg_pair(f, g)
# -
# Define the global functional-gradient operator. This does the following:
# * Maps the worklist (shots) to the workers so that the invidual contributions to (f, g) are computed.
# * Sum individual contributions to (f, g) and returns the result.
def fwi_gradient(model, param):
# Dump a copy of the current model for the workers
# to pick up when they are ready.
param['model'] = "model_0.p"
dump_model(param['model'], wrap_model(model))
# Define work list
work = [dict(param) for i in range(param['nshots'])]
for i in range(param['nshots']):
work[i]['shot_id'] = i
# Distribute worklist to workers.
fgi = c.map(fwi_gradient_i, work, retries=1)
# Perform reduction.
fg = c.submit(sum, fgi).result()
# L-BFGS in scipy expects a flat array in 64-bit floats.
return fg.f, fg.g.flatten().astype(np.float64)
# ## FWI with L-BFGS-B
# Equipped with a function to calculate the functional and gradient, we are finally ready to define the optimization function.
# +
from scipy import optimize
# Many optimization methods in scipy.optimize.minimize accept a callback
# function that can operate on the solution after every iteration. Here
# we use this to monitor the true relative solution error.
relative_error = []
def fwi_callbacks(x):
# Calculate true relative error
true_vp = get_true_model().vp.data[param['nbl']:-param['nbl'], param['nbl']:-param['nbl']]
true_m = 1.0 / (true_vp.reshape(-1).astype(np.float64))**2
relative_error.append(np.linalg.norm((x-true_m)/true_m))
# FWI with L-BFGS
ftol = 0.1
maxiter = 5
def fwi(model, param, ftol=ftol, maxiter=maxiter):
# Initial guess
v0 = model.vp.data[param['nbl']:-param['nbl'], param['nbl']:-param['nbl']]
m0 = 1.0 / (v0.reshape(-1).astype(np.float64))**2
# Define bounding box constraints on the solution.
vmin = 1.4 # do not allow velocities slower than water
vmax = 4.0
bounds = [(1.0/vmax**2, 1.0/vmin**2) for _ in range(np.prod(model.shape))] # in [s^2/km^2]
result = optimize.minimize(fwi_gradient,
m0, args=(param, ), method='L-BFGS-B', jac=True,
bounds=bounds, callback=fwi_callbacks,
options={'ftol':ftol,
'maxiter':maxiter,
'disp':True})
return result
# -
# We now apply our FWI function and have a look at the result.
# +
#NBVAL_IGNORE_OUTPUT
model0 = get_initial_model()
# Baby steps
result = fwi(model0, param)
# Print out results of optimizer.
print(result)
# +
#NBVAL_SKIP
# Plot FWI result
from examples.seismic import plot_image
slices = tuple(slice(param['nbl'],-param['nbl']) for _ in range(2))
vp = 1.0/np.sqrt(result['x'].reshape(true_model.shape))
plot_image(true_model.vp.data[slices], vmin=2.4, vmax=2.8, cmap="cividis")
plot_image(vp, vmin=2.4, vmax=2.8, cmap="cividis")
# +
#NBVAL_SKIP
import matplotlib.pyplot as plt
# Plot model error
plt.plot(range(1, maxiter+1), relative_error); plt.xlabel('Iteration number'); plt.ylabel('L2-model error')
plt.show()
# -
# As can be observed in last figures, the results we obtain are exactly the same to the ones obtained in [04_dask.ipynb](https://github.com/devitocodes/devito/blob/master/examples/seismic/tutorials/04_dask.ipynb)
| original examples/seismic/tutorials/04_dask_pickling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Initialization
# +
import spacy
from chemdataextractor import Document
from chemdataextractor.model import Compound
from chemdataextractor.doc import Paragraph, Heading
# -
# ## Playing with two different synthetic paragraphs
#
# `text` is from the Angewandte paper out of Stanford. The synthetic paragraph is not very straightforward.
#
# `text2` is taken from the Methods section of the Nature Nanotech paper out of the University of Toronto. The synthetic portion is much more straightforward in this case.
#
# `text3` is taken from the Methods section of the Nature Materials paper from Korea. The synthesis portion is somewhat readable, but lies in between the previous two texts.
nlp = spacy.load('en_core_web_sm') # Load English dictionary from spaCy
# +
# Angewandte
#text = 'Layered perovskites can be structurally derived from the 3D analogue by slicing along specific crystallographic planes.4 The interlayer separation and thickness of the inorganic layers can be controlled through the choice of organic cations.5 The inorganic layers of most layered perovskites comprise a single sheet (n=1) of corner‐sharing metal–halide octahedra sandwiching layers of organic cations.4 These 2D materials do not have electronic properties typically associated with good solar‐cell absorbers. Along with larger bandgaps compared to the 3D analogue (n=∞), the spatial confinement of the 2D structure and dielectric mismatch between organic and inorganic layers lead to strongly bound excitons with low mobility.6 Such tightly bound excitons are difficult to dissociate into free carriers at room temperature and the localized charge carriers are unlikely to reach the electron/hole selective contacts in a typical solar‐cell geometry. To access the more favorable electronic properties of the 3D structure, we sought an intermediate structure between the n=1 and n=∞ materials. We synthesized the n=3 member of the series (PEA)2(MA)n−1[PbnI3n+1] (n=number of Pb–I sheets in each inorganic layer), by combining (PEA)I, (MA)I, and PbI2 in a 2:2:3 stoichiometric ratio in a solvent mixture of nitromethane/acetone. Slow solvent evaporation afforded dark red crystals of (PEA)2(MA)2[Pb3I10] (1), the first crystallographically characterized n=3 lead perovskite (Figure 1).'
# -
# Nature Nanotechnology
text2 = "Perovskite film fabrication Different dimensionality perovskite [(PEA)2(CH3NH3)n−1PbnI3n+1] solutions was prepared by dissolving stoichiometric quantities of lead iodide (PbI2), methylammonium idodide (MAI) and PEAI in a dimethyl sulfoxide (DMSO)/ɣ-butyrolactone (1:1 volume ratio) mixture at 70 °C for 1 h with continuous stirring. The resulting solution was then filtered through a polytetrafluoroethylene (PTFE) filter (0.2 µm). The resulting solution was spin-coated onto the substrate via a two-step process at 1,000 r.p.m. and 5,000 r.p.m. for 10 s and 60 s, respectively. During the second spin step, 100 µl of chlorobenzene were poured onto the substrate. The resulting films were then annealed at 70 °C for 10 min to improve crystallization."
# Nature Materials
text3 = u"A dense blocking layer of TiO2 (bl-TiO2, ∼70 nm in thickness) was deposited onto a F-doped SnO2 (FTO, Pilkington, TEC8) substrate by spray pyrolysis, using a 20 mM titanium diisopropoxide bis(acetylacetonate) solution (Aldrich) at 450 °C to prevent direct contact between the FTO and the hole-conducting layer. A 200–300-nm-thick mesoporous TiO2 (particle size: about 50 nm, crystalline phase: anatase) film was spin-coated onto the bl-TiO2/FTO substrate using home-made pastes14 and calcining at 500 °C for 1 h in air to remove organic components. CH3NH3I (MAI) and CH3NH3Br (MABr) were first synthesized by reacting 27.86 ml CH3NH2 (40% in methanol, Junsei Chemical) and 30 ml HI (57 wt% in water, Aldrich) or 44 ml HBr (48 wt% in water, Aldrich) in a 250 ml round-bottom flask at 0 °C for 4 h with stirring, respectively. The precipitate was recovered by evaporation at 55 °C for 1 h. MAI and MABr were dissolved in ethanol, recrystallized from diethyl ether, and dried at 60 °C in a vacuum oven for 24 h. The prepared MAI and MABr powders, PbI2 (Aldrich) and PbBr2 (Aldrich) for 0.8 M MAPb(I1 − xBrx)3 (x = 0.1–0.15) solution were stirred in a mixture of GBL and DMSO (7:3 v/v) at 60 °C for 12 h. The resulting solution was coated onto the mp-TiO2/bl-TiO2/FTO substrate by a consecutive two-step spin-coating process at 1,000 and 5,000 r.p.m for 10 and 20 s, respectively. During the second spin-coating step, the substrate (around 1 cm × 1 cm) was treated with toluene drop-casting. A detailed time-rotation profile for the spin-coating is represented in Supplementary Fig. 1c. The substrate was dried on a hot plate at 100 °C for 10 min. A solution of poly(triarylamine) (15 mg, PTAA, EM Index, Mw = 17,500 g mol−1) in toluene (1.5 ml) was mixed with 15 μl of a solution of lithium bistrifluoromethanesulphonimidate (170 mg) in acetonitrile (1 ml) and 7.5 μl 4-tert-butylpyridine and spin-coated on the MAPb(I1 − xBrx)3 (x = 0.1–0.15)/mp-TiO2/bl-TiO2/FTO substrate at 3,000 r.p.m for 30 s. Finally, a Au counterelectrode was deposited by thermal evaporation. The active area of this electrode was fixed at 0.16 cm2."
# ### Nature Materials paragraph: Playing with ChemDataExtractor
# Formatting the Nature Nanotech paragraph according to ChemDataExtractor
nat = Document(Heading(u'Solar cell fabrication'), Paragraph(text3))
nat.records.serialize()
nat.paragraphs[0].pos_tagged_tokens
# +
sp_nm = nlp(text3)
for token in sp_nm:
print(token.text, token.lemma_, token.pos_, token.tag_, token.dep_,
token.shape_, token.is_alpha, token.is_stop)
# +
#sp_ang = nlp(text)
#for token in sp_ang:
# print(token.text, token.lemma_, token.pos_, token.tag_, token.dep_,
# token.shape_, token.is_alpha, token.is_stop)
# +
#angew = Document(text)
# +
#angew
# -
# ## Defining Custom Properties in CDE
#
# I'm using mostly code from the CDE notebook to define a new property for spin-coating step(s).
nat
# +
from chemdataextractor.model import BaseModel, StringType, ListType, ModelType
class SpinCoat(BaseModel):
value = StringType()
units = StringType()
Compound.spin_coat_spd = ListType(ModelType(SpinCoat))
# +
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
from __future__ import unicode_literals
import logging
#from chemdataextractor.model.cem import chemical_name
# +
# logging.getLogger?
# -
# hao 2 edit 2 make more useful
def extract_units(tokens, start, result):
"""Extract units from bracketed after nu"""
for e in result:
for child in e.iter():
if 'rpm' or 'r.p.m.' or 'r.p.m' or 'rcf' or 'r.c.f.' in child.text:
return [E('units', 'whatever the unit is')]
return []
# +
import re
from chemdataextractor.parse import R, I, W, Optional, merge, ZeroOrMore, OneOrMore
from chemdataextractor.parse.cem import chemical_name
solvent = (I('GBL') | R('γ-[Bb]utyrolactone') | chemical_name('solvent'))
units = Optional(R(u'^\b?r(\.)?p(\.)?m(\.)?\b?$') | R(u'^r(\.)?c(\.)?f(\.)?$') | R(u'^([x×]?)(\s?)?g$'))(u'units')
#Optional(W('/')).hide() + W(u'^r\.?p\.?m\.?')
#R('^(re)?crystalli[sz](ation|ed)$', re.I)
value = R(u'^\d+(,\d+)?$')(u'value')
spinspd = (value + units)(u'spinspd')
from chemdataextractor.parse.base import BaseParser
from chemdataextractor.utils import first
def extract_units(tokens, start, result):
"""Extract units from bracketed after nu"""
for e in result:
for child in e.iter():
if R(u'^\b?r(\.)?p(\.)?m(\.)?\b?$') in child.text:
return [E('units', 'rpm')]
elif R(u'^r(\.)?c(\.)?f(\.)?$') in child.text:
return [E('units', 'rcf')]
elif R(u'^([x×]?)(\s?)?g$') in child.text:
return [E('units', 'g')]
return []
class SpinCoatParser(BaseParser):
root = spinspd
def interpret(self, result, start, end):
compound = Compound(
spin_coat_spd=[
SpinCoat(
solvent=first(result.xpath('./solvent/text()')),
value=first(result.xpath('./value/text()')),
units=first(result.xpath('./units/text()'))
)
]
)
yield compound
# -
Paragraph.parsers = [SpinCoatParser()]
# +
d = Document(u'The resulting solution was coated onto the mp-TiO2/bl-TiO2/FTO substrate by a consecutive two-step spin-coating process at 1,000 and 5,000 r.p.m for 10 and 20 s, respectively.')
d.records.serialize()
# -
| development_notebooks/dev_synthesis_parsers/Synthesis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 08 - Introduction to APIs
#
# by [<NAME>](albahnsen.com/) and [<NAME>](https://github.com/jesugome)
#
# version 1.5, June 2020
#
# ## Part of the class [Advanced Methods in Data Analysis](https://github.com/albahnsen/AdvancedMethodsDataAnalysisClass)
#
#
#
# This notebook is licensed under a [Creative Commons Attribution-ShareAlike 3.0 Unported License](http://creativecommons.org/licenses/by-sa/3.0/deed.en_US).
#
# Adapted with permision from https://restful.io/an-introduction-to-api-s-cee90581ca1b
# + [markdown] slideshow={"slide_type": "slide"}
# API stands for application programming interfaces.
#
# An API is the tool that makes a website’s data digestible for a computer. Through it, a computer can view and edit data, just like a person can by loading pages and submitting forms.
#
# 
#
# When systems link up through an API, we say they are integrated. One side the server that serves the API, and the other side the client that consumer the API and can manipulate it.
# + [markdown] slideshow={"slide_type": "slide"}
# # HTTP Request
# Communication in HTTP (Hyper Text Transfer Protocol) center around a concept called the Request-Response Cycle.
#
# 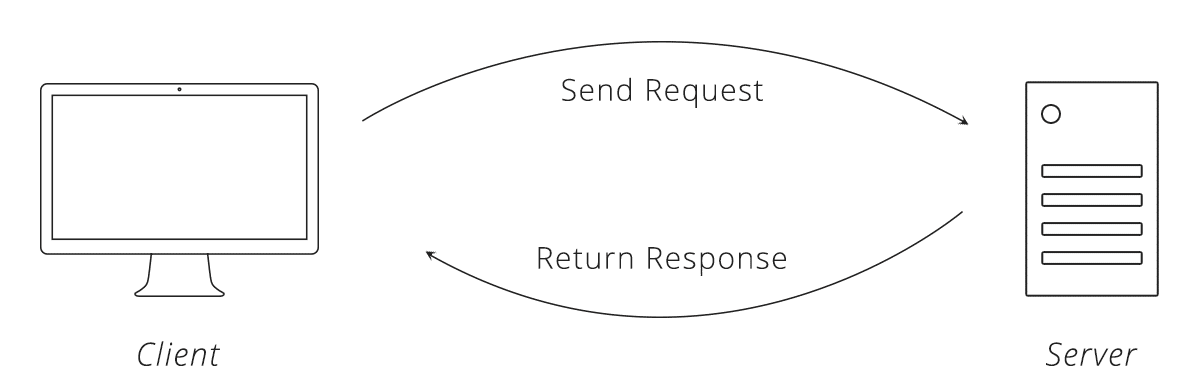
#
# To make a valid request, the client needs to include four things:
#
# 1. URL ( Uniform Resource Locator)
# 2. Method
# 3. List of Headers
# 4. Body
# + [markdown] slideshow={"slide_type": "subslide"}
# ## URL
# URLs become an easy way for the client to tell the server which things it wants to interact with, called resources.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Method
# The method request tells the server what kind of action the client wants the server to take in. The four most commonly seen in API’s are:
#
# * GET — Asks the server to retrieve a resource.
# * POST — Asks the server to create a new resource.
# * PUT — Asks the server to edit/update an existing resource.
# * DELETE — Asks the server to delete a resource.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Headers
# Headers provide meta-information about a request.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Body
# The request body contains the data the client wants to send the server.
#
# 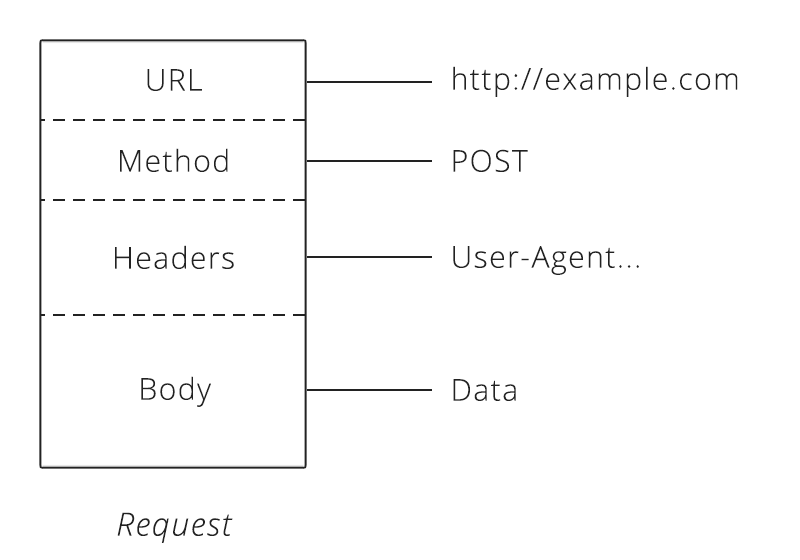
# + [markdown] slideshow={"slide_type": "slide"}
# # HTTP Response
# The server responds with a status code. Status code are three-digit numbers.
#
# 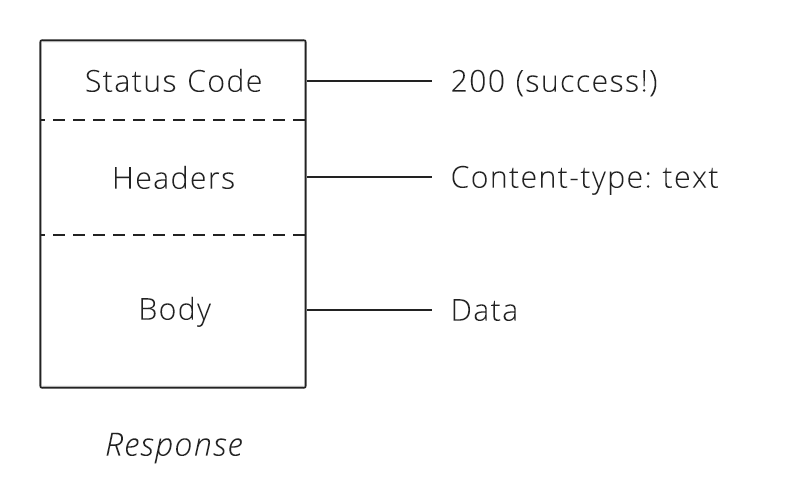
# + [markdown] slideshow={"slide_type": "slide"}
# # Data Formats
# A well-designed format is dictated by what makes the information the easiest for the intended audience to understand. The most common formats found in APIs are JSON and XML.
# + [markdown] slideshow={"slide_type": "subslide"}
# **JSON** is very simple format that has two pieces: key and value.
#
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# **XML** provides a few building blocks. The main block is called a node. XML always starts with a root node, inside there are more “child” nodes. The name of the node tells us the attribute of the order and the data inside is the actual details.
#
# 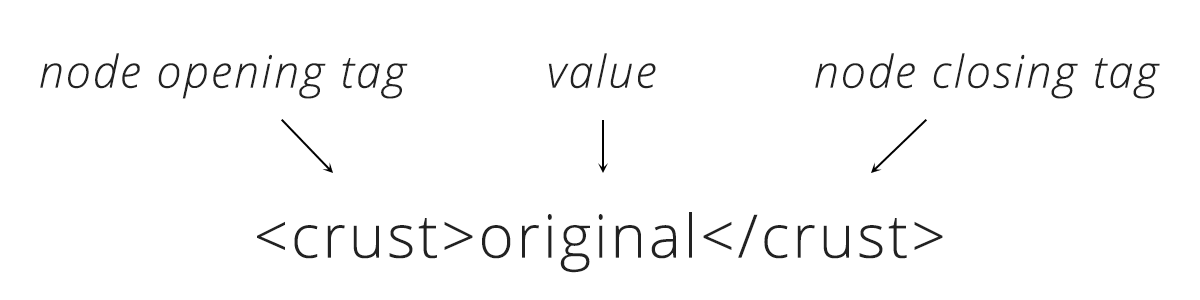
# + [markdown] slideshow={"slide_type": "subslide"}
# ## How Data Formats Are Used In HTTP
# Using Headers we can tell the server what information we are sending to it and what we expect in return.
#
# Content-type: When the clients send the Content-type its saying what format the data is.
#
# Accept: The Accept header tells the server what data-format it is able to accept.
#
# 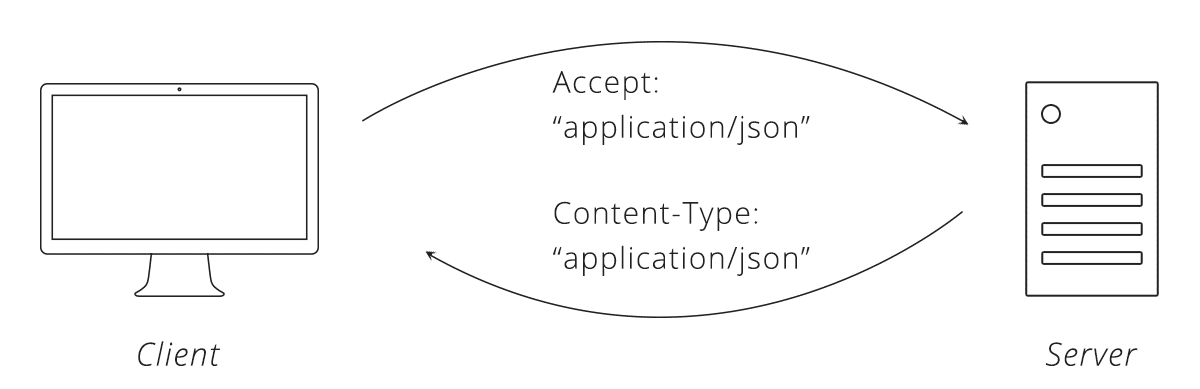
# + [markdown] slideshow={"slide_type": "slide"}
# ## Authentication, Part 1
# There are several techniques APIs use to athenticate a client. These are called authentication schemes.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Basic Authentication
# Also referred as Basic Auth. Basic Auth only requires a user name and password. The client takes these two credentials, converts them to a single value and passes that along in the HTTP header called Authentication.
#
# 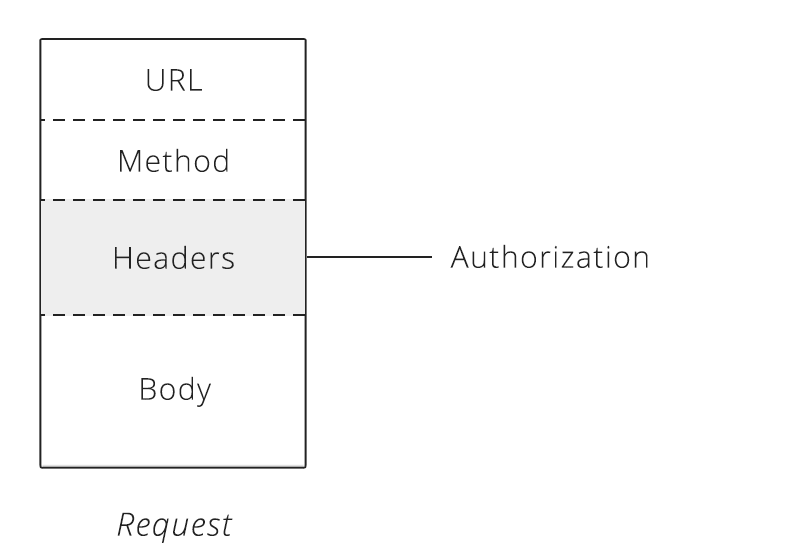
#
# The server compares the Authorization header and compares it to the credential it has stored. If it matches, the server fulfills the request. If there is no match, the server returns status code 401.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### API Key Authentication
#
# API Key Authentication is a technique that overcomes the weakness of using shared credentials. by requiring the API to be accessed with a unique key. Unlike Basic Auth, API keys were conceived at multiple companies in the early days of the web. As a result, API Key Authentication has no standard and everybody has its own way of doing it.
#
# The most common approach has been to include it onto the URL(http://example.com?apikey=mysecret_key).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Authentication, Part 2
# Open Authorization (OAuth) automates the key exchange. OAuth only requires user credentials, then behind the scenes, the client and server are chatting back and forth to get the client a valid key.
#
# Currently there are two versions of OAuth, OAuth1 and OAuth2.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### OAuth2
#
# The players involved are:
# - The User - A person that wants to connect to the website
# - The Client - The website that will be grated the access to the user's data
# - The Server - The website that has the user's data
# + [markdown] slideshow={"slide_type": "slide"}
# # Authentication Example
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Step 1 — User Tells Client to Connect to Server
#
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Step 2 — Client Directs User to Server
#
# The client sends the user over to the server’s website, along with a URL that the server will send the user back to once the user authenticates, called the callback URL.
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Step 3 — User Logs-in to Server and Grants Client Access
#
# With their normal user name and password, the user authenticates with the server.
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Step 4 — Server Sends User Back to Client, Along with Code
#
# 
#
# The server sends the user back to the client using the callback URL. Hidden in the response is a unique authorization code for the client.
#
# 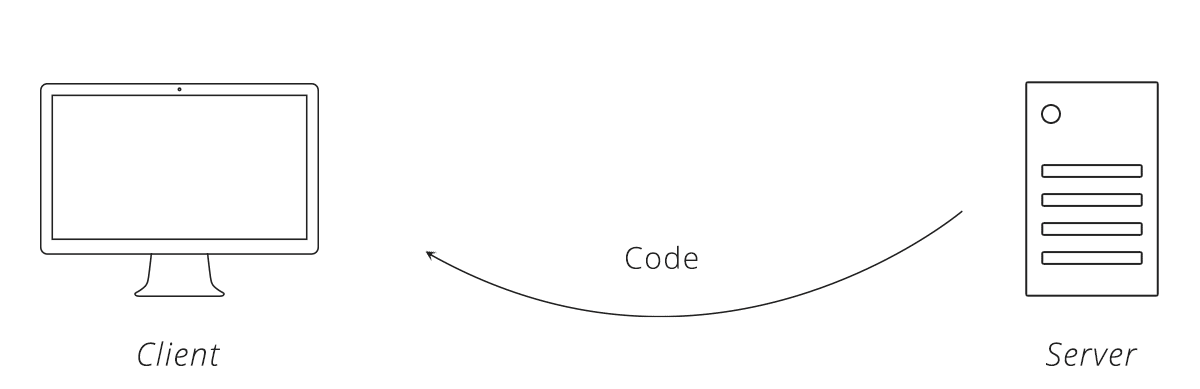
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Step 5 — Client Exchange Code + Secret Key for Access Token
#
# The client takes the authorization code it receives and makes another request to the server. This request includes the client’s secret key. When the server sees a valid authorization code and a trusted client secret key, it is certain that the client is who it claims. The server responds back with an access token.
#
# 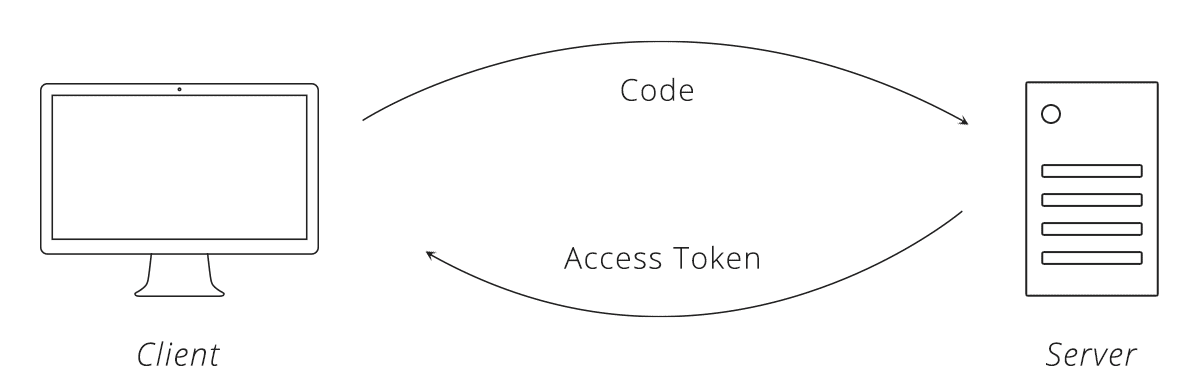
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Step 6 — Client Fetches Data from Server.
#
# 
#
# The access token from Step 5 is essentially another password into the user’s account on the server. The client includes the access token with every request so it can authenticate directly with the server.
#
# Client Refresh Token (Optional)
#
# A feature in OAuth 2 is the option to have access tokens expire. The lifespan of a token is set by the server.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Authorization
# Authorization is the concept of limiting access. In Step 2, when the user allows the client access, buried in the fine print are the exact permissions the client is asking for. Those permission are called scope.
#
# What makes scope powerful is that is client-based restrictions. OAuth scope allows one client to have permission X and another to have permission X and Y.
# + [markdown] slideshow={"slide_type": "slide"}
# # API Design
# ## Start with an Architectural Style
# The two most common architectures for web-based APIs are SOAP, which is an XML-based design that has a standardized structures for requests and responses and REST (Representation State Transfer), is a more open approach, providing lots of conventions, but leaving many decisions to the person designing the API.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Our First Resource
# Resources are the nouns of APIs (customers and pizza). These are things we want the world to interact with.
#
# Let’s try to design an API for a pizza parlour. For the client to be able to talk pizza with us, we need to do several things:
#
# 1. Decide what resource(s) need to be available
# 2. Assign URLs to the resources
# 3. Decide what actions the client should be allowed to perform on those resources
# 4. Figure out what pieces of data are required for each action and what format they should be in.
# + [markdown] slideshow={"slide_type": "subslide"}
# Lets get started with orders. The next step is assigning URLs to the resources. In a typical REST API, a resource will have two URL patterns assigned to it. The first is the plural of the resource name, like orders/. The second is the plural of the resources name plus a unique identifier to specify a single resource, like orders/, where is the unique identifier for an order. These two parameters make up the first endpoints that our API will support. These are called endpoints simply because they go at the end of the URL, as in http://example.com/
# + [markdown] slideshow={"slide_type": "subslide"}
# Now that we picked our resources and assigned it URLs, we need to decide what actions the client can perform. Following REST conventions, we say that the plural endpoint (orders/) is for listing existing orders and creating new ones. The plural with a unique identifier endpoint (orders/), is for retrieving, updating, or canceling a specific order. The client tells the server which action to perform by passing the appropriate HTTP verb (GET, POST, PUT or DELETE) in the request.
# + [markdown] slideshow={"slide_type": "subslide"}
# Our API now looks like this:
#
# 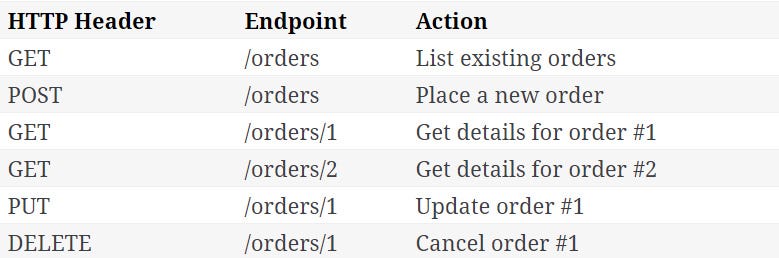
# + [markdown] slideshow={"slide_type": "subslide"}
# Now we decide what data needs to be exchanged. Coming from our pizza parlour, we can say an order of pizza needs a crust and toppings. We also need to select a data format, lets go with JSON. Here is what an interaction between the client and server might look like using this API:
# + [markdown] slideshow={"slide_type": "subslide"}
# 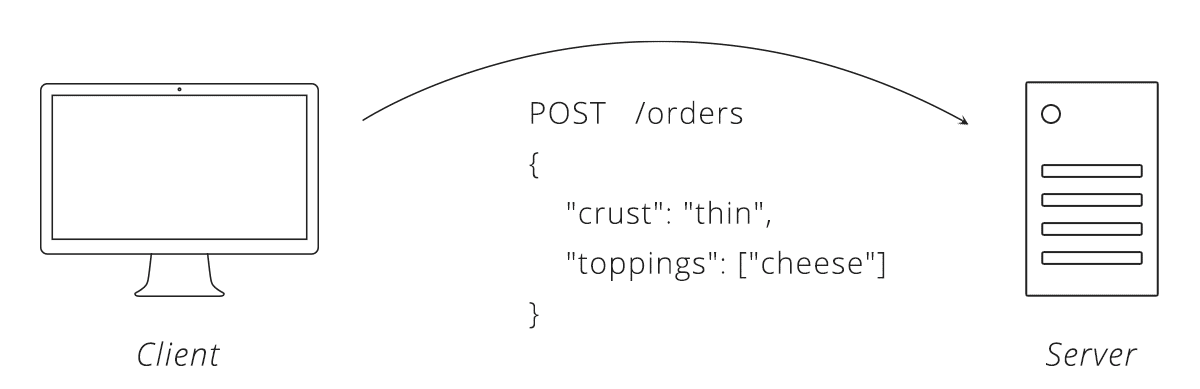
# + [markdown] slideshow={"slide_type": "subslide"}
# 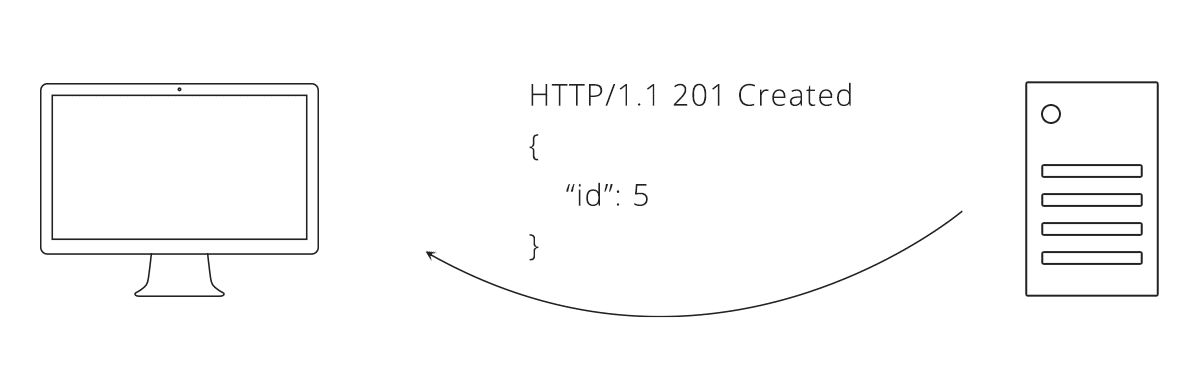
# + [markdown] slideshow={"slide_type": "subslide"}
# 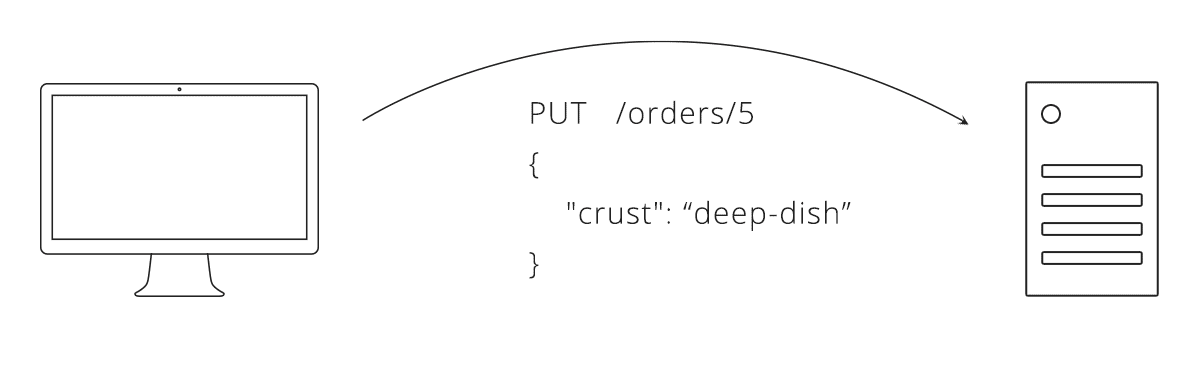
# + [markdown] slideshow={"slide_type": "subslide"}
# 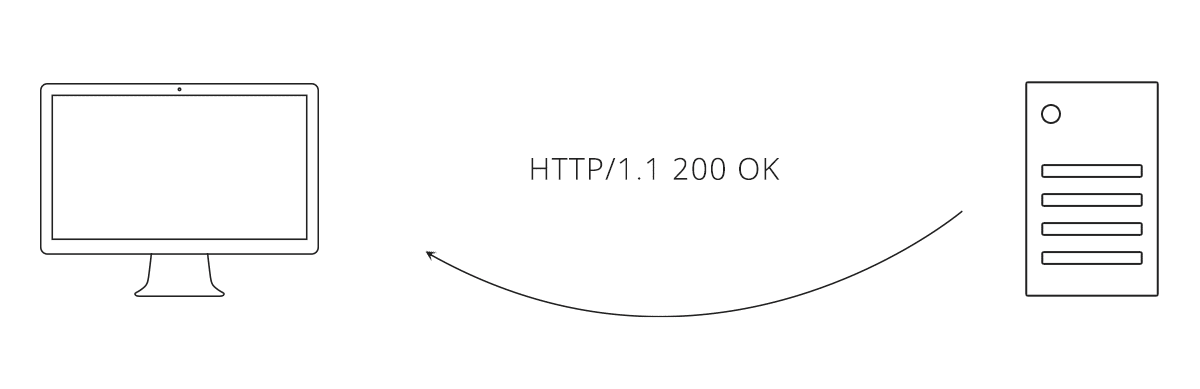
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Linking Resources Together
# We want add a new customer resource to track orders by customer. Just like with orders, our customer resource needs some endpoint. Following convention, /customer and /customer/.
#
# How do we associate orders with customers?
# + [markdown] slideshow={"slide_type": "subslide"}
# REST practitioners are split on how to solve the problem of associating resources. Some say that the hierarchy should continue to grow, giving endpoints like /customers/5/orders for all of customer #5's orders and/customers/5/orders/3 for customer #5's third order. Others argue to keep things flat by including associated details in the data for a resource. Under this paradigm, creating an order requires a customer_id field to be sent with the order details. Both solutions are used by REST APIs in the wild, so it is worth knowing about each.
# + [markdown] slideshow={"slide_type": "subslide"}
# 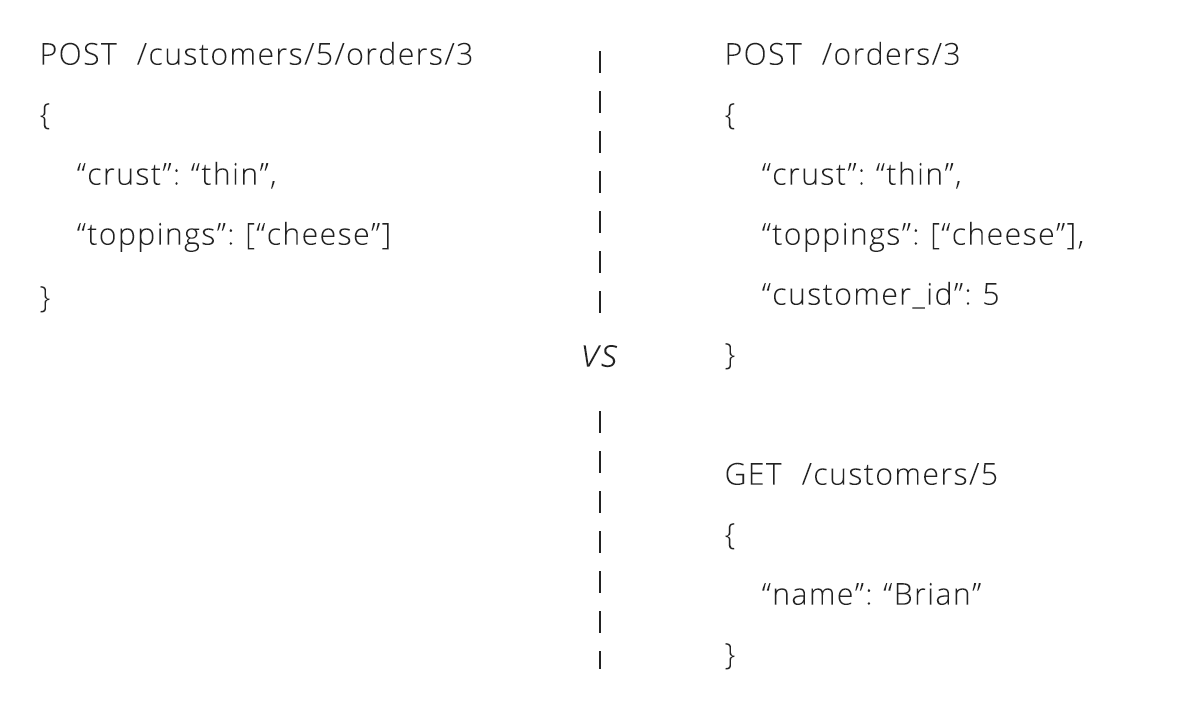
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Searching Data
# URLs have another component called the query string. For example:http://example.com/orders?key=value. REST APIs use the query string to define details of a search. These details are called query parameters. The API dictates what parameters it will accept, and the exact names of those parameters need to be used for them to effect the search. In our case key can allow search by topping and/or curst and you can concatenate the search with an ampersand(&).
# + [markdown] slideshow={"slide_type": "subslide"}
# For example: http://example.com/orders?topping=pepperoni&crust=thin
#
# Another use of the query string is to limit the amount of data returned in each request. The process of splitting data is called pagination. If the client makes a request like GET /orders?page=2&size=200, we know they want the second page of results, with 200 results per page, so order 201–400.
# + [markdown] slideshow={"slide_type": "slide"}
# # Glossary
# - Server: A powerful computer that runs an API
# - API: The “hidden” portion of a website that is meant for computer consumption
# - Client: A program that exchanges data with a server through an API
# - Request — consists of a URL (http://...), a method (GET, POST, PUT, DELETE), a list of headers (User-Agent…), and a body (data).
# - Response — consists of a status code (200, 404…), a list of headers, and a body.
# - JSON: JavaScript Object Notation
# + [markdown] slideshow={"slide_type": "subslide"}
# # Glossary
# - Object: a thing or noun (person, pizza order…)
# - Key: an attribute about an object (color, toppings…)
# - Value: the value of an attribute (blue, pepperoni…)
# - Associative array: a nested object
# - XML: Extensible Markup Language
# - Authentication: process of the client proving its identity to the server
# + [markdown] slideshow={"slide_type": "subslide"}
# # Glossary
# - Credentials: secret pieces of info used to prove the client’s identity (username, password…)
# - Basic Auth: scheme that uses an encoded username and password for credentials
# - API Key Auth: scheme that uses a unique key for credentials
# - Authorization Header: the HTTP header used to hold credentials
# - OAuth: an authentication scheme that automates the key exchange between client and server.
# - Access Token: a secret that the client obtains upon successfully completing the OAuth process.
# + [markdown] slideshow={"slide_type": "subslide"}
# # Glossary
# - Scope: permissions that determine what access the client has to user’s data.
# - SOAP: API architecture known for standardized message formats
# - REST: API architecture that centers around manipulating resources
# - Resource: API term for a business noun like customer or order
# - Endpoint: A URL that makes up part of an API. In REST, each resource gets its own endpoints
# + [markdown] slideshow={"slide_type": "subslide"}
# # Glossary
# - Query String: A portion of the URL that is used to pass data to the server
# - Query Parameters: A key-value pair found in the query string (topping=cheese)
# - Pagination: Process of splitting up results into manageable chunks
| notebooks/08-IntroductionToAPIs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Capstone: Analisando a temperatura do Brasil
#
# Nessa prática iremos juntar os conhecimentos que vimos durante toda a matéria. Vamos analisar os dados de temperatura do Brasil que vimos nas aulas de Python. Vamos utilizar uma [regressão linear](https://pt.wikipedia.org/wiki/Regress%C3%A3o_linear) para estimar a taxa de variação da temperatura ao longo dos anos. Finalmente, vamos visualizar esses valores em um mapa do Brasil.
#
# Faremos funções para cada etapa para podermos testar as partes do nosso código separadamente.
# ## Setup
#
# Abaixo, vamos carregar todas as bibliotecas que precisamos:
#
# * `numpy` para fazer contas com matrizes e vetores
# * `maptlotlib.pyplot` para fazer gráficos
# * `mpl_toolkits.basemap` para fazer mapas
# * `scipy.stats` para a regressão linear
# * `glob` para descobrir os nomes de todos os arquivos em uma pasta
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.basemap import Basemap
from scipy.stats import linregress
from glob import glob
# <div class="alert text-center" style="font-size: 16pt">
# Diferente das outras práticas, eu vou preecher o código inicial ao vivo durante a aula.
# <br><br>
# Façam junto comigo para não se perderem.
# </div>
# <h1>Tarefas:</h1>
# 1 - Ler lat, lon;<p>
# 2 - Ler temp_abs;<p>
# 3 - Ler as anomalias, anos demais;<p>
# 4 - Calcular o temperatura real;<p>
# 5 - Regressão(anos, temperatura real);<p>
# 6 - Mapa(lat, lon e a).
def le_lat_lon(nome_arquivo):
"""
Coleta a latituda(lat) e a longitude(lon) a partir de um arquivo de dados.
"""
arquivo = open(nome_arquivo)
for linhas in range(0,5):
linha = arquivo.readline()
partes = linha.split()
lat = float(partes[1])
if partes[2] == 'S,':
lat = -lat
lon = float(partes[3])
if partes[4] == 'W':
lon = -lon
return lat,lon
lat, lon = le_lat_lon("dados/0.80S-49.02W-TAVG-Trend.txt")
def le_temp_abs(nome_arquivo):
arquivo = open(nome_arquivo)
for linhas in range(0,48):
linha = arquivo.readline()
partes = linha.split()
temp_abs = float(partes[8])
return temp_abs
temp_abs = le_temp_abs("dados/0.80S-49.02W-TAVG-Trend.txt")
def le_anos_anomalias(nome_arquivo):
dados = np.loadtxt(nome_arquivo,comments="%")
anomalia = dados[:,4]
anos = dados[:,0]
meses = dados[:,1]
anos_decimais = ( anos + ( meses / 12))
return anos_decimais, anomalia
anos, anom = le_anos_anomalias("dados/0.80S-49.02W-TAVG-Trend.txt")
plt.figure()
plt.plot(anos,anom,'-k')
temperaturas = temp_abs + anom
anos_sem_nan = []
temperaturas_sem_nan = []
for i in range(len(temperaturas)):
if not np.isnan(temperaturas[i]):
temperaturas_sem_nan.append(temperaturas[i])
anos_sem_nan.append(anos[i])
not_nan = ~np.isnan(temperaturas)
temperaturas_sem_nan = temperaturas[not_nan]
anos_sem_nan = anos[not_nan]
a, b, r_value, p_value, std_err = linregress(anos_sem_nan, temperaturas_sem_nan)
print(a)
arquivos = glob("dados/*.txt")
latitude = []
longitude = []
angulo = []
for arquivo in arquivos:
"Captura a latitude e a longitude e armazena no vetor correspondente"
lat, lon = le_lat_lon(arquivo)
latitude.append(lat)
longitude.append(lon)
"Capturar a temperatura absoluta, os anos decimais e anomalias térmicas"
temp_abs = le_temp_abs(arquivo)
anos, anom = le_anos_anomalias(arquivo)
"Calcula a temperatura absoluta e retira os nans delas"
temperaturas = temp_abs + anom
not_nan = ~np.isnan(temperaturas)
temperaturas_sem_nan = temperaturas[not_nan]
anos_sem_nan = anos[not_nan]
"Calcula a taxa de variação das temperaturas"
a, b, r_value, p_value, std_err = linregress(anos_sem_nan, temperaturas_sem_nan)
angulo.append(a)
print(latitude)
print(longitude)
print(angulo)
plt.figure()
plt.scatter(longitude, latitude,s=200,c=angulo,cmap="Reds")
cb = plt.colorbar()
cb.set_label("°C/ano")
bm = Basemap(projection='merc',llcrnrlat=-35,urcrnrlat=5,llcrnrlon=-65,urcrnrlon=-35)
# +
plt.figure(figsize=[8,9])
bm.scatter(longitude, latitude,s=40,c=angulo,cmap="Reds",latlon=True)
cb = plt.colorbar()
cb.set_label('°C/ano')
bm.drawcoastlines(linewidth=0.5)
bm.drawcountries()
bm.drawstates(linewidth=0.1)
# -
# **Course website**: https://github.com/mat-esp/about
#
# **Note**: This notebook is part of the course "Matemática Especial I" of the [Universidade do Estado do Rio de Janeiro](http://www.uerj.br/). All content can be freely used and adapted under the terms of the
# [Creative Commons Attribution 4.0 International License](http://creativecommons.org/licenses/by/4.0/).
#
# 
| capstone.ipynb |
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ ---
/ + [markdown] cell_id="00000-b0144350-4f8d-4a62-9dd3-65e55c5031f8" deepnote_cell_type="markdown" tags=[]
/ # Ideation
/
/
/ + cell_id="00000-c13a5cd8-24d3-44f4-b64e-6f69327e5979" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=9333 execution_start=1632708205579 source_hash="f1484b6c" tags=[]
# Imports etc
#!pip install scikit-optimize
!pip install eli5
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import mpl_toolkits
from collections import deque
from functools import reduce
from sklearn import preprocessing
from sklearn.model_selection import cross_val_score
from sklearn.model_selection import train_test_split
from sklearn import ensemble
from sklearn.metrics import roc_curve, auc
import eli5
from eli5.sklearn import PermutationImportance
from sklearn.model_selection import GridSearchCV
/ %matplotlib inline
/ + cell_id="00002-0d33c2c4-edff-41b4-80bc-b5a35a01cef7" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=165 execution_start=1632708214916 source_hash="adbb9718" tags=[]
# Get data
global_df = pd.read_csv("nfl_games_and_bets.csv")
global_df = global_df.drop(global_df[global_df.schedule_season == 2021].index)
global_df = global_df.drop(columns=['stadium','weather_temperature', 'weather_wind_mph','weather_humidity','weather_detail'])
global_df = global_df.drop(global_df[global_df.schedule_season < 2010].index)
global_df
/ + cell_id="00003-0078aed8-5e65-4247-be6e-3d39d4f89449" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=151 execution_start=1632708215079 source_hash="28bf552b" tags=[]
# Account for team moves
old_to_new_team_name = {"San Diego Chargers": "Los Angeles Chargers", "St. Louis Rams": "Los Angeles Rams", \
"W<NAME>" : "Washington Football Team", "Oakland Raiders": "Las Vegas Raiders"}
global_df = global_df.replace({"team_away": old_to_new_team_name}).replace({"team_home": old_to_new_team_name})
# Maintain consistency between favourite and team name columns
short_form_to_team_name = {"GB": "Green Bay Packers", "HOU": "Houston Texans", "KC": "Kansas City Chiefs", "BUF": "Buffalo Bills", \
"TEN": "Tennessee Titans", "NO": "New Orleans Saints", "SEA": "Seattle Seahawks", "MIN": "Minnesota Vikings", \
"TB": "Tampa Bay Buccaneers", "LVR": "Las Vegas Raiders", "BAL": "Baltimore Ravens", "LAC": "Los Angeles Chargers", \
"IND": "Indianapol<NAME>", "DET": "Detroit Lions", "CLE": "Cleveland Browns", "JAX": "Jacksonville Jaguars", "MIA": "<NAME>", \
"ARI": "Arizona Cardinals", "PIT": "Pittsburgh Steelers", "CHI": "Chicago Bears","ATL": "Atlanta Falcons", "CAR": "Carolina Panthers", \
"LAR": "Los Angeles Rams", "CIN": "Cincinnati Bengals", "DAL": "Dallas Cowboys", "SF": "San Francisco 49ers", "NYG": "New York Giants", \
"WAS": "Washington Football Team", "DEN": "Denver Broncos", "PHI": "Philadelphia Eagles", "NYJ": "New York Jets", "NE": "New England Patriots"}
team_name_to_short_form = {value: key for key, value in short_form_to_team_name.items()}
global_df = global_df.replace({'team_away': team_name_to_short_form}).replace({"team_home": team_name_to_short_form})
# Note: 'PICK' when spread == 0
global_df
/ + [markdown] cell_id="00004-5643e281-a05c-44dc-a2ac-6414581396f9" deepnote_cell_type="markdown" tags=[]
/ # Columns I'd like to add
/ * Last game result (plus/minus) -- have enough data, but need to account for bye
/ * Last few games results -- have enough data, also need to account for bye (maybe W in last 3, or total point difference in last 3)
/ * Covered spread last game (bool) - have enough data
/ * Who covered spread this game -- already a column but may need to feature engineer a bit
/ * wind direction / speed - not enough data
/ * divisional - have enough data but need to build the dataset manually
/ * prime time - don't have enough data
/ * twitter sentiment - don't have enough data
/ * garbage game ? - can use schedule_date maybe
/ + cell_id="00004-f7aa0641-2cab-44f5-a30f-37e3d8853c53" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=235 execution_start=1632708215228 source_hash="a9c800f3" tags=[]
# Determine if teams are within the same division
# AFC = A, NFC = N
# West = W, etc etc
team_to_division = {"ARI": "NW", "LAR": "NW", "SF": "NW", "SEA": "NW", "CAR": "NS", "TB": "NS", "NO": "NS", "ATL": "NS", \
"GB": "NN", "CHI": "NN", "MIN": "NN", "DET": "NN", "WAS": "NE", "DAL": "NE", "PHI": "NE", "NYG": "NE", \
"TEN": "AS", "HOU": "AS", "IND": "AS", "JAX": "AS", "BUF": "AE", "MIA": "AE", "NE": "AE", "NYJ": "AE", \
"BAL": "AN", "PIT": "AN", "CLE": "AN", "CIN": "AN", "LVR": "AW", "DEN": "AW", "KC": "AW", "LAC": "AW"}
global_df2 = global_df
global_df2['home_division'] = global_df2.apply(lambda row: team_to_division[row.team_home], axis=1)
global_df2['away_division'] = global_df2.apply(lambda row: team_to_division[row.team_away], axis=1)
global_df2['intra_division'] = global_df2.apply(lambda row: row.home_division == row.away_division, axis=1)
global_df2 = global_df2.drop(columns=['home_division', 'away_division'])
global_df2
/ + cell_id="00006-cdfba250-8ce3-4619-8ef5-63e17d8ed6e4" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=729 execution_start=1632708215513 source_hash="ca4b640e" tags=[]
# Create auxillary columns to make calculations easier
global_df3 = global_df2
global_df3['home_point_diff'] = global_df2.apply(lambda row: row.score_home - row.score_away, axis=1)
global_df3['away_point_diff'] = global_df3.apply(lambda row: row.score_away - row.score_home, axis=1)
global_df3['home_spread'] = global_df3.apply(lambda row: row.spread_favorite * -1 if row.team_favorite_id == row.team_away else row.spread_favorite, axis=1)
# Loop, sorry pandas
team_to_games = {}
# Get last one result
for index, row in global_df3.iterrows():
# Update the mapping
if row.team_home not in team_to_games:
team_to_games.update({row.team_home : deque([0,0,0])})
if row.team_away not in team_to_games:
team_to_games.update({row.team_away : deque([0,0,0])})
last_games = team_to_games.get(row.team_home)
home_last_3 = last_games[0] + last_games[1] + last_games[2]
home_last_1 = last_games[0]
last_games.pop()
last_games.appendleft(row.home_point_diff)
last_games = team_to_games.get(row.team_away)
away_last_3 = last_games[0] + last_games[1] + last_games[2]
away_last_1 = last_games[0]
last_games.pop()
last_games.appendleft(row.away_point_diff)
global_df3.at[index, 'home_last_3'] = home_last_3
global_df3.at[index, 'away_last_3'] = away_last_3
global_df3.at[index, 'home_last_1'] = home_last_1
global_df3.at[index, 'away_last_1'] = away_last_1
# Update the DF
team_to_games
#global_df3['home_last_game_result'] = global_df3.groupby('')
#global_df3
#team_to_games.get("TB")[0]
/ + cell_id="00007-41bf27cd-af26-49b0-9d6d-481d65f62390" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=137 execution_start=1632708216239 source_hash="a268d41a" tags=[]
global_df3
/ + cell_id="00008-38e4363f-dcf2-4973-9677-d888bb3e91c0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=820 execution_start=1632708216409 source_hash="3bbadca5" tags=[]
global_df_final = global_df3
global_df_final['home_team_covered'] = global_df_final.apply(lambda row: row.home_point_diff + row.home_spread > 0, axis=1)
global_df_final_no_drop = global_df_final
global_df_final = global_df_final.drop(columns = ['schedule_date', 'schedule_week', 'team_home', 'score_home', 'score_away', 'team_away', \
'team_favorite_id', 'spread_favorite', 'away_point_diff'])
# Correlation
corr = global_df_final.corr()
plt.figure(figsize=(20,20))
ax = sns.heatmap(
corr,
vmin=-1, vmax=1, center=0,
cmap=sns.diverging_palette(20, 220, n=200),
square=True
)
ax.set_xticklabels(
ax.get_xticklabels(),
rotation=45,
horizontalalignment='right'
)
#global_df_final
/ + cell_id="00009-e1bcfab9-bef3-46a8-8af6-1d9cd72facd9" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=41047 execution_start=1632708217230 source_hash="79dd3084" tags=[]
# Regressor
train_data = global_df_final.drop(['home_team_covered', 'home_point_diff'], axis=1)
target_label = global_df_final['home_point_diff']
n_features = train_data.shape[1]
x_train, x_test, y_train, y_test = train_test_split(train_data, target_label, test_size = 0.30)
parameters = {
"n_estimators":[5,50, 100],
"max_depth":[1,3,5,7,9],
"learning_rate":[0.01,0.1,1]
}
def display(results):
print(f'Best parameters are: {results.best_params_}')
print("\n")
mean_score = results.cv_results_['mean_test_score']
std_score = results.cv_results_['std_test_score']
params = results.cv_results_['params']
for mean,std,params in zip(mean_score,std_score,params):
print(f'{round(mean,3)} + or -{round(std,3)} for the {params}')
gbc = ensemble.GradientBoostingRegressor()
cv = GridSearchCV(gbc,parameters,cv=5)
cv.fit(x_train, y_train)
display(cv)
/ + cell_id="00010-0dd1ff58-93ea-4062-8c80-a86011298c56" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=173 execution_start=1632708258321 source_hash="20a6e555" tags=[]
train_data2 = global_df_final.drop(['home_team_covered', 'home_point_diff'], axis=1)
target_label2 = global_df_final['home_point_diff']
y_pred_full = cv.predict(train_data2)
global_df_final_no_drop["predicted_diff"] = y_pred_full
global_df_final_no_drop
#y_pred_full = cv.predict(x_test)
#len(y_pred_full)
#df_with_predictions = global_df3
/ + cell_id="00011-fc00a92b-6802-4a32-854b-d541199da132" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=223 execution_start=1632708258491 source_hash="e272072b" tags=[]
perm = PermutationImportance(cv).fit(x_test, y_test)
eli5.show_weights(perm, feature_names = x_test.columns.tolist())
/ + cell_id="00011-2a5afb88-5d83-4b45-aaf9-0799f2fc66b0" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=47 execution_start=1632708258761 source_hash="31538ba6" tags=[]
# Ultimate backtester!!!!
# integer, last two are the decimals. start with $100.00. Each bet is $3
# assume even -110 odds on everything. so bet $3 = win $2.73
money = 10000
won = 0
loss = 0
push = 0
for row in global_df_final_no_drop.itertuples():
if row.predicted_diff + row.home_spread > 2:
print(row)
if row.home_point_diff + row.home_spread > 0:
money = money + 273
won += 1
elif row.home_point_diff + row.home_spread == 0:
push +=1
else:
money = money - 300
loss += 1
if row.predicted_diff + row.home_spread < -2:
print(row)
if row.away_point_diff - row.home_spread > 0:
money = money - 300
loss += 1
elif row.away_point_diff - row.home_spread == 0:
push +=1
else:
money = money + 273
won += 1
print (money)
print (won)
print (loss)
print (push)
/ + cell_id="00013-86d1eb85-a13b-4c6c-8dfd-98f9253f13ce" deepnote_cell_type="code" deepnote_to_be_reexecuted=false execution_millis=7 execution_start=1632708489210 source_hash="2f091579" tags=[]
# the fields most correlated with 'home_team_covered' is away_last_3 (positively)
# so, what if i always bet the home team when away_last_3 is > some value
global_df_final_no_drop.describe()
#away last 3: 75% = 20
money = 10000
won = 0
loss = 0
push = 0
for row in global_df_final_no_drop.itertuples():
# if (row.away_last_3 < -30 and row.away_last_1 < 0 and row.intra_division):
if (row.away_last_1 <= -14 and row.intra_division and row.home_spread < 0): #bet on road underdog
if row.away_point_diff - row.home_spread > 0:
money = money + 273
won += 1
elif row.away_point_diff - row.home_spread > 0:
push +=1
else:
money = money - 300
loss += 1
print (money)
print (won)
print (loss)
print (push)
/ + [markdown] created_in_deepnote_cell=true deepnote_cell_type="markdown" tags=[]
/ <a style='text-decoration:none;line-height:16px;display:flex;color:#5B5B62;padding:10px;justify-content:end;' href='https://deepnote.com?utm_source=created-in-deepnote-cell&projectId=f0efbe77-01fa-4860-b5ee-e7eac30d44e8' target="_blank">
/ <img alt='Created in deepnote.com' style='display:inline;max-height:16px;margin:0px;margin-right:7.5px;' src='data:image/svg+xml;base64,PD94bWwgdmVyc2lvbj0iMS4wIiBlbmNvZGluZz0iVVRGLTgiPz4KPHN2ZyB3aWR0aD0iODB<KEY> > </img>
/ Created in <span style='font-weight:600;margin-left:4px;'>Deepnote</span></a>
| nfl_spread_exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python3-datasci
# language: python
# name: python3-datasci
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import matplotlib
# %matplotlib inline
matplotlib.rcParams['figure.figsize'] = (12, 8) # set default figure size, 8in by 6in
# # Ensemble Learning
#
# Sometimes aggregrates or ensembles of many different opinions on a question can perform as well or better than asking
# a single expert on the same question. This is known as the *wisdom of the crowd* when the aggregrate opinion of
# people on a question performs as well or better as a single isolated expert in predicting some outcome.
#
# Likewise, for machine learning predictors, a similar effect can also often occur. The aggregrate performance of
# multiple predictors and often make a small but significant improvement on building a classifier or regression
# predictor for a complex set of data. A group of machine learning predictors is called an *ensemble*, and thus
# this technique of combining the predictions of an ensemble is known as *Ensemble Learning* .
#
# For exampl,e we could train a group of Decision Tree classifiers, each on a different random subset of the training
# data. To make an ensemble prediciton, you just obtain the predictions of all individual trees, then predict the
# class that gets the most votes. Such an ensemble of Decision Trees is called a *Random Forest*, and despite the
# relative simplicity of decision tree predictors, it can be surprisingly powerful as a ML predictor.
# # Voting Classifiers
#
# Say you have several classifiers for the same classification problem (say a Logistic Classifier, and SVM,
# a Decision Tree and a KNN classifier and perhaps a few more). The simplest way to create an ensemble classifier
# is to aggregrate the predictions of each classifier and predict the class that gets the most votes. This
# majority-vote classifier is called a *hard voting* classifier.
#
# Somewhat surprisingly, this voting classifier often achieves a higher accuracy than the best classifier in the
# ensemble. In fact, even if each classifier is a *weak learner* (meaning it only does slightly better than
# random guessing), the ensemble can still be a *strong learner* (achieving high accuracy). The key to making
# good ensemble predictors is that you need both a sufficient number of learners (even of weak learners), but
# also maybe more importantly, the learners need to be "sufficiently diverse", where diverse is a bit fuzzy to
# define, but in general the classifiers must be as independent as possible, so that even if they are weak predictors,
# they are weak in different and diverse ways.
#
#
# +
def flip_unfair_coin(num_flips, head_ratio):
"""Simulate flipping an unbalanced coin. We return a numpy array of size num_flips, with 0 to represent
1 to represent a head and 0 a tail flip. We generate a head or tail result using the head_ratio probability
threshold drawn from a standard uniform distribution.
"""
# array of correct size to hold resulting simulated flips
flips = np.empty(num_flips)
# flip the coin the number of indicated times
for flip in range(num_flips):
flips[flip] = np.random.random() < head_ratio
# return the resulting coin flip trials trials
return flips
def running_heads_ratio(flips):
"""Given a sequence of flips, where 1 represents a "Head" and 0 a "Tail" flip, return an array
of the running ratio of heads / tails
"""
# array of correct size to hold resulting heads ratio seen at each point in the flips sequence
num_flips = flips.shape[0]
head_ratios = np.empty(num_flips)
# keep track of number of heads seen so far, the ratio is num_heads / num_flips
num_heads = 0.0
# calculate ratio for each flips instance in the sequence
for flip in range(num_flips):
num_heads += flips[flip]
head_ratios[flip] = num_heads / (flip + 1)
# return the resulting sequence of head ratios seen in the flips
return head_ratios
# +
NUM_FLIPPERS = 10
NUM_FLIPS = 10000
HEAD_PERCENT = 0.51
# create 3 separate sequences of flippers
flippers = np.empty( (NUM_FLIPPERS, NUM_FLIPS) )
for flipper in range(NUM_FLIPPERS):
flips = flip_unfair_coin(NUM_FLIPS, HEAD_PERCENT)
head_ratios = running_heads_ratio(flips)
flippers[flipper] = head_ratios
# create an ensemble, in this case we will average the individual flippers
ensemble = flippers.mean(axis=0)
# plot the resulting head ratio for our flippers
flips = np.arange(1, NUM_FLIPS+1)
for flipper in range(NUM_FLIPPERS):
plt.plot(flips, flippers[flipper], alpha=0.25)
plt.plot(flips, ensemble, 'b-', alpha=1.0, label='ensemble decision')
plt.ylim([0.42, 0.58])
plt.plot([1, NUM_FLIPS], [HEAD_PERCENT, HEAD_PERCENT], 'k--', label='51 %')
plt.plot([1, NUM_FLIPS], [0.5, 0.5], 'k-', label='50 %')
plt.xlabel('Number of coin tosses')
plt.ylabel('Heads ratio')
plt.legend();
# -
# ## Scikit-Learn Voting Classifier
#
# The following code is an example of creating a voting classifier in Scikit-Learn. We are using the moons dataset
# shown.
#
# Here we create 3 separate classifiers by hand, a logistic regressor, a decision tree,
# and a support vector classifier (SVC). Notice we specify 'hard' voting for the voting classifier, which
# as we discussed is the simple method of choosing the class with the most votes.
# (This is a binary classification so 2 out of 3 or 3 out of 3 are the only possibilities. For a multiclass
# classification, in case of a tie vote, the voting classifier may fall back to the probability scores the
# classifiers give, assuming the provide probability/confidence measures of their prediction).
# +
# helper functions to visualize decision boundaries for 2-feature classification tasks
# create a scatter plot of the artificial multiclass dataset
from matplotlib import cm
# visualize the blobs using matplotlib. An example of a funciton we can reuse, since later
# we want to plot the decision boundaries along with the scatter plot data
def plot_multiclass_data(X, y):
"""Create a scatter plot of a set of multiclass data. We assume that X has 2 features so that
we can plot on a 2D grid, and that y are integer labels [0,1,2,...] with a unique integer
label for each class of the dataset.
Parameters
----------
X - A (m,2) shaped number array of m samples each with 2 features
y - A (m,) shaped vector of integers with the labeled classes of each of the X input features
"""
# hardcoded to handle only up to 8 classes
markers = ['o', '^', 's', 'd', '*', 'p', 'P', 'v']
#colors = ['r', 'g', 'b', 'c', 'm', 'y', 'k']
# determine number of features in the data
m = X.shape[0]
# determine the class labels
labels = np.unique(y)
#colors = cm.rainbow(np.linspace(0.0, 1.0, labels.size))
colors = cm.Set1.colors
# loop to plot each
for label, marker, color in zip(labels, markers, colors):
X_label = X[y == label]
y_label = y[y == label]
label_text = 'Class %s' % label
plt.plot(X_label[:,0], X_label[:,1],
marker=marker, markersize=8.0, markeredgecolor='k',
color=color, alpha=0.5,
linestyle='',
label=label_text)
plt.xlabel('Feature 1')
plt.ylabel('Feature 2')
plt.legend();
def plot_multiclass_decision_boundaries(model, X, y):
from matplotlib.colors import ListedColormap
"""Use a mesh/grid to create a contour plot that will show the decision boundaries reached by
a trained scikit-learn classifier. We expect that the model passed in is a trained scikit-learn
classifier that supports/implements a predict() method, that will return predictions for the
given set of X data.
Parameters
----------
model - A trained scikit-learn classifier that supports prediction using a predict() method
X - A (m,2) shaped number array of m samples each with 2 features
"""
# determine the class labels
labels = np.unique(y)
#colors = cm.rainbow(np.linspace(0.0, 1.0, labels.size))
#colors = cm.Set1.colors
newcmp = ListedColormap(plt.cm.Set1.colors[:len(labels)])
# create the mesh of points to use for the contour plot
h = .02 # step size in the mesh
x_min, x_max = X[:, 0].min() - h, X[:, 0].max() + h
y_min, y_max = X[:, 1].min() - h, X[:, 1].max() + h
xx, yy = np.meshgrid(np.arange(x_min, x_max, h),
np.arange(y_min, y_max, h))
# create the predictions over the mesh using the trained models predict() function
Z = model.predict(np.c_[xx.ravel(), yy.ravel()])
# Create the actual contour plot, which will show the decision boundaries
Z = Z.reshape(xx.shape)
plt.contourf(xx, yy, Z, cmap=newcmp, alpha=0.33)
#plt.colorbar()
# +
from sklearn.datasets import make_moons
X, y = make_moons(n_samples=2500, noise=0.3)
# we will split data using a 75%/25% train/test split this time
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=42)
# +
from sklearn.ensemble import VotingClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeClassifier
from sklearn.svm import SVC
log_clf = LogisticRegression(solver='lbfgs', C=5.0)
tree_clf = DecisionTreeClassifier(max_depth=10)
svm_clf = SVC(gamma=100.0, C=1.0)
voting_clf = VotingClassifier(
estimators=[('lr', log_clf), ('tree', tree_clf), ('svc', svm_clf)],
voting='hard'
)
voting_clf.fit(X_train, y_train)
# -
plot_multiclass_decision_boundaries(voting_clf, X, y)
plot_multiclass_data(X, y)
# Lets look at each classifier's accuracy on the test set, including for the ensemble voting classifier:
# +
from sklearn.metrics import accuracy_score
for clf in (log_clf, tree_clf, svm_clf, voting_clf):
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(clf.__class__.__name__, accuracy_score(y_test, y_pred))
# -
# The voting classifier will usually outperform all the individual classifier, if the data is sufficiently
# nonseparable to make it relatively hard (e.g. with less random noise in the moons data set, you can get
# real good performance sometimes with random forest and/or svc, which will exceed the voting classifier).
#
# If all classifiers are able to estimate class probabilities (i.e. in `scikit-learn` they support
# `predict_proba()` method), then you can tell `scikit-learn` to predict the class with the highest class
# probability, averaged over all individual classifiers. You can think of this as each classifier having
# its vote weighted by its confidence of the prediction. This is called *soft voting*. It often achieves
# higher performance than hard voting because it gives more weight to highly confident votes. All you
# need to do is replace `voting='hard'` with `voting='soft'` and ensure that all classifiers can estimate
# clas sprobabilities. If you recall, support vector machine classifiers (`SVC`) do not estimate class probabilities by
# default, but if you set `SVC` `probability` hyperparameter to `True`, the `SVC` class will use cross-validation
# to estimate class probabilities. This slows training, but it makes the `predict_proba()` method valid
# for `SVC`, and since both logistic regression and random forests support this confidence estimate, we
# can then use soft voting for the voting classifier.
# +
log_clf = LogisticRegression(solver='lbfgs', C=5.0)
tree_clf = DecisionTreeClassifier(max_depth=8)
svm_clf = SVC(gamma=1000.0, C=1.0, probability=True) # enable probability estimates for svm classifier
voting_clf = VotingClassifier(
estimators=[('lr', log_clf), ('tree', tree_clf), ('svc', svm_clf)],
voting='soft' # use soft voting this time
)
voting_clf.fit(X_train, y_train)
# -
plot_multiclass_decision_boundaries(voting_clf, X, y)
plot_multiclass_data(X, y)
# +
from sklearn.metrics import accuracy_score
for clf in (log_clf, tree_clf, svm_clf, voting_clf):
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print(clf.__class__.__name__, accuracy_score(y_test, y_pred))
# -
# # Bagging and Pasting
#
# One way to get a diverse set of classifiers is to use very different training algorithms. The previous voting
# classifier was an example of this, where we used 3 very different kinds of classifiers for the voting ensemble.
#
# Another approach is to use the same training for every predictor, but to train them on different random
# subsets of the training set. When sampling is performed with replacement, this method is called
# *bagging* (short for *bootstrap aggregrating*). When sampling is performed without replacement, it is
# called *pasting*.
#
# In other words, both approaches are similar. In both cases you are sampling the training data to build
# multiple instances of a classifier. In both cases a training item could be sampled and used to train
# multiple instances in the collection of classifiers that is produced. In bagging, it is possible for a training
# sample to be sampled multiple times in the training for the same predictor. This type of bootstrap aggregration
# is a type of data enhancement, and it is used in other contexts as well in ML to artificially increase the size
# of the training set.
#
# Once all predictors are trained, the ensemble can make predictions for a new instance by simply aggregating the
# predictions of all the predictors. The aggregration function is typically the *statistical mode* (i.e. the
# most frequent prediction, just like hard voting) for classification, or the average for regression.
#
# Each individual predictor has a higher bias than if it were trained on the original training set (because you don't
# use all of the training data on an individual bagged/pasted classifier). But the aggregration overall should usually
# reduce both bias and variance on the final performance. Generall the net result is that the ensemble has a similar
# bias but a lower variance than a single predictor trained on the whole original training set.
#
# Computationally bagging and pasting are very attractive because in theory and in practice all of the classifiers
# can be trained in parallel. Thus if you have a large number of CPU cores, or even a distributed memory
# computing cluster, you can independently train the individual classifiers all in parallel.
#
# ## Scikit-Learn Bagging and Pasting Examples
#
# The ensemble API in `scikit-learn` for performing bagging and/or pasting is relatively simple. As with the voting
# classifier, we specify which type of classifer we want to use. But since bagging/pasting train multiple
# classifiers all of this type, we only have to specify 1. The `n_jobs` parameter tells `scikit-learn` the number of
# cpu cores to use for training and predictions (-1 tells `scikit-learn` to use all available cores).
#
# The following trains an ensemble of 500 decision tree classifiers (`n_estimators`), each trained on 100 training
# instances randomly sampled from the training set with replacement (`bootstrap=True`). If you want to use pasting
# we simply set `bootstrap=False` instead.
#
# **NOTE**: The `BaggingClassifier` automatically performs soft voting instead of hard voting if the base classifier
# can estimate class probabilities (i.e. it has a `predict_proba()` method).
# +
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
bag_clf = BaggingClassifier(
DecisionTreeClassifier(max_leaf_nodes=20), n_estimators=500,
max_samples=100, bootstrap=True, n_jobs=-1
)
bag_clf.fit(X_train, y_train)
y_pred = bag_clf.predict(X_test)
print(bag_clf.__class__.__name__, accuracy_score(y_test, y_pred))
# -
plot_multiclass_decision_boundaries(bag_clf, X, y)
plot_multiclass_data(X, y)
# ## Out-of-Bag Evaluation
#
# With bagging, some instances may be sampled several times for any given predictor, while others may not be
# sampled at all. By default a `BaggingClassifier` samples `m` training instances with replacement, where `m`
# is the size of the training set. This means that only about 63% of the training instances are sampled on average for
# each predictor. The remaining 37% of the training instances that are not sampled are called *out-of-bag* (oob)
# instances. **NOTE**: they are not the same 37% for each resulting predictor, each predictor has a different oob.
#
# Since a predictor never sees the oob instances during training, it can be evaluated on these instances, without the need
# for a separate validation set or cross-validation. You can evaluate the ensemble itself by averaging out the oob
# evaluations for each predictor.
#
# In `scikit-learn` you can set `oob_score=True` when creating a `BaggingClassifier` to request an automatic oob
# evaluation after training:
# +
bag_clf = BaggingClassifier(
DecisionTreeClassifier(), n_estimators=500,
bootstrap=True, n_jobs=-1, oob_score=True
)
bag_clf.fit(X_train, y_train)
print(bag_clf.oob_score_)
y_pred = bag_clf.predict(X_test)
print(accuracy_score(y_test, y_pred))
# -
# The oob decision function for each training instance is also available through the
# `oob_decision_function_` variable.
bag_clf.oob_decision_function_
# ## Random Patches and Random Subspaces
#
# The default behavior of the bagging/patching classifier is to only sample the training target outputs. However,
# it can also be useful to build classifiers that only use some of the feautres of the input data. We have looked
# at methods for adding features, for example by adding polynomial combinations of the feature inputs. But often for
# big data, we might have thousands or even millions of input features. In that case, it can very well be that some
# or many of the features are not really all that useful, or even somewhat harmful, to building a truly good and
# general classifier.
#
# So one approach when we have large number of features is to build multiple classifiers (using bagging/patching)
# on sampled subsets of the features. In `scikit-learn` `BaggingClassifier` this is controllerd by two
# hyperparameters: `max_features` and `bootstrap_features`. They work the same as `max_samples` and `bootstrap`
# but for feature sampling instead of output instance sampling. Thus each predictor will be trained on a random subset
# of the input features. This is particularly useful when dealing with high-dimensional inputs.
#
# Sampling from both training instances and features simultaneously is called the *Random Patches method*.
# Keeping all training instances, but sampling features is called *Random Subspaces method*.
# # Random Forests
#
# As we have already mentioned, a `RandomForest` is simply an ensemble of decision trees, generally trained via the
# bagging method, typically with `max_samples` set to the size of the training set. We could create a
# random forest by hand using `scikit-learn` `BaggingClassifier` on a DecisionTree, which is in fact what we just
# did in the previous section. Our previous ensemble was an example of a random forest classifier.
#
# But in `scikit-learn` instead of building the ensemble somewhat by hant, you can instead use the
# `RandomForestClassifier` class, which is more convenient and which has default hyperparameter settings
# optimized for random forests.
#
# The following code trains a random forest classifier with 500 treas (each limited to a maximum of 16 nodes),
# using all available CPU cores:
# +
from sklearn.ensemble import RandomForestClassifier
rnd_clf = RandomForestClassifier(n_estimators=500, max_leaf_nodes=16, n_jobs=-1)
rnd_clf.fit(X_train, y_train)
y_pred = rnd_clf.predict(X_test)
print(accuracy_score(y_test, y_pred))
# -
# A random forest classifier has all of the hyperparameters of a `DecisionTreeClassifier` (to control
# how trees are grown), plus all of the hyperparameters of a `BaggingClassifier` to control the ensemble itself.
#
# The random forest algorithm introduces extra randomness when growing trees. Instead of searching
# for the very best feature when splitting a node, it searches for the best feature among a random subset of
# features. This results in a greater tree diversity, which trades a higher bias for a lower variance, generally yielding
# a better overall ensemble model.
#
# The following `BaggingClassifier` is roughly equivalent to the previous `RandomForestClassifier`:
# +
bag_clf = BaggingClassifier(
DecisionTreeClassifier(splitter='random', max_leaf_nodes=16),
n_estimators=500, max_samples=1.0, bootstrap=True, n_jobs=-1
)
bag_clf.fit(X_train, y_train)
y_pred = bag_clf.predict(X_test)
print(accuracy_score(y_test, y_pred))
# -
# ## Extra-Trees
#
# When growing a tree in a random forest, at each node only a random subset of features is considered for splitting as
# we just discussed. It is possible to make trees even more random by also using random thresholds for each
# feature rather than searching for the best possible thresholds.
#
# A forest of such extremely random trees is called an *Extremely Randomized Trees* ensemble (or *Extra-Trees*
# for short.
#
# You can create an extra-trees classifier using `scikit-learn`s `ExtraTreesClassifier` class, its API is identical
# to the `RandomForestClassifier` class.
#
# **TIP:** It is hard to tell in advance whether a random forest or an extra-tree will perform better or worse on a
# given set of data. Generally the only way to know is to try both and compare them using cross-validation.
# ## Feature Importance
#
# Lastly, if you look at a single decision tree, important features are likely to appear closer to the root of the
# tree, while unimportnat features will often appear closer to th eleaves (or not a all). Therefore another
# use of random forests is to get an estimate on the importance of the features when making classification
# predictions.
#
# We can get an estimate of a feature's importance by computing the average depth at which it appears across all
# trees in a random forest. `scikit-learn` computes this automatically for every feature after training. You can
# access the result using the `feature_importances_` variable.
#
# For example, if we build a `RandomForestClassifier` on the iris data set (with 4 features), we can output each
# features estimated importance.
# +
from sklearn.datasets import load_iris
iris = load_iris()
rnd_clf = RandomForestClassifier(n_estimators=500, n_jobs=-1)
rnd_clf.fit(iris['data'], iris['target'])
for name, score in zip(iris['feature_names'], rnd_clf.feature_importances_):
print(name, score)
# -
# It seems the most importan feature is petal length, followed closely by petal width. Sepal length and especially
# sepal width are relatively less important.
# # Boosting
#
# *Boosting* (originally called *hypothesis boosting* refers to any ensemble method that can combine several weak learners
# into a strong learner. But unlike the ensembles we looked at before, the general idea is to train predictors
# sequentially, each trying to correct it predecessor. There are many boosting methods, the most popular being
# *AdaBoost* (short for *Adaptive Boosting*) and *Gradient Boosting*.
#
# ## AdaBoost
#
# ## Gradient Boost
#
#
# # Stacking
#
# Stacking works similar to the voting ensembles we have looked at. Multiple independent classifiers are trained
# in parallel and aggregrated. But instead of using a trivial aggregration method (like hard voting), we train
# yet another model to perform the aggregration. This final model (called a *blender* or *meta learner*) takes
# the other trained predictors's output as input and makes a final prediciton from them.
#
# 'Scikit-learn' does not support stacking directly (unlike voting ensembles and boosting). But it is not too difficult
# to hand roll basic implementations of stacking from `scikit-learn` apis.
# +
import sys
sys.path.append("../../src") # add our class modules to the system PYTHON_PATH
from ml_python_class.custom_funcs import version_information
version_information()
| lectures/ng/Lecture-09-Ensembles-Random-Forests.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Various tests for performance using Numpy
import numpy as np
#
# How to populate Numpy array:
#
# %%timeit
a = np.zeros([100, 1000])
for i in range(100):
a[i] = np.random.randint(0, 8, size=(1000))
# %%timeit
b = []
for i in range(100):
b.append(np.random.randint(0, 8, size=(1000)))
a = np.array(b)
#
# Importance of not breaking a pipeline:
# %%timeit
a = 0
c = 32
for i in range(1024):
if i % c == 0:
a += 1
# %%timeit
a = 0
c = 32
for i in range(int(1024 / c)):
for j in range(c):
pass
a += 1
#
#
# Cost of one `if` statement:
# %%timeit
for i in range(1024):
if False:
pass
# %%timeit
for i in range(1024):
pass
#
#
# Expensive conversion of NaN to 0:
# %%timeit a = np.arange(32, dtype='float'); s = 0;
for i in np.arange(32):
s += a[i]
# %%timeit a = np.arange(32, dtype='float'); s = 0; a[np.random.choice(np.arange(32), size=8, replace=False)] = np.nan
for i in np.arange(32):
s += np.nan_to_num(a[i])
# %%timeit a = np.arange(32, dtype='float'); s = 0; a[np.random.choice(np.arange(32), size=8, replace=False)] = np.nan
a[a == np.nan] = 0
for i in np.arange(32):
s += a[i]
| scribble.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (asa)
# language: python
# name: asa
# ---
# # Advanced Spatial Analysis
# # Module 10: Inferential Spatial Modeling
#
# Statistical inference is the process of using a sample to *infer* the characteristics of an underlying population (from which this sample was drawn) through estimation and hypothesis testing. Contrast this with descriptive statistics, which focus simply on describing the characteristics of the sample itself.
#
# Common goals of inferential statistics include:
#
# - parameter estimation and confidence intervals
# - hypothesis rejection
# - prediction
# - model selection
#
# To conduct statistical inference, we rely on *statistical models*: sets of assumptions plus mathematical relationships between variables, producing a formal representation of some theory. We are essentially trying to explain the process underlying the generation of our data. What is the probability distribution (the probabilities of occurrence of different possible outcome values of our response variable)?
#
# **Spatial inference** introduces explicit spatial relationships into the statistical modeling framework, as both theory-driven (e.g., spatial spillovers) and data-driven (e.g., MAUP) issues could otherwise violate modeling assumptions.
#
# Schools of statistical inference:
#
# - frequentist
# - frequentists think of probability as proportion of times some outcome occurs (relative frequency)
# - given lots of repeated trials, how likely is the observed outcome?
# - concepts: statistical hypothesis testing, *p*-values, confidence intervals
# - bayesian
# - bayesians think of probability as amount of certainty observer has about an outcome occurring (subjective probability)
# - probability as a measure of how much info the observer has about the real world, updated as info changes
# - concepts: prior probability, likelihood, bayes' rule, posterior probability
#
# 
# +
import geopandas as gpd
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import pysal as ps
import seaborn as sns
import statsmodels.api as sm
from scipy import stats
from statsmodels.stats.outliers_influence import variance_inflation_factor as vif
from statsmodels.tools.tools import add_constant
np.random.seed(0)
# %matplotlib inline
# -
# load the data
tracts = gpd.read_file('data/census_tracts_data.geojson')
tracts.shape
# map the data
tracts.plot()
tracts.columns
# ## 1. Statistical inference: introduction
#
# ### 1a. Estimating population parameters
# descriptive stats
tracts['med_household_income'].describe()
# descriptive stat: average tract-level median income
tracts['med_household_income'].mean()
# descriptive stat of a simple random sample
n = 500
sample = tracts['med_household_income'].sample(n)
sample.mean()
# How similar is our sample mean to our population mean? Is it a good estimate?
# +
# calculate confidence interval using t-distribution (bc population std dev is unknown)
sample = sample.dropna() #drop nulls
conf = 0.95 #confidence level
df = len(sample) - 1 #degrees of freedom
loc = sample.mean() #the mean
scale = stats.sem(sample) #the standard error
conf_lower, conf_upper = stats.t.interval(conf, df, loc=loc, scale=scale)
# calculate the margin of error
moe = conf_upper - sample.mean()
# display confidence interval
print(f'{conf_lower:0.0f} – {conf_upper:0.0f} ({conf*100:0.0f}% confidence interval)')
print(f'{loc:0.0f} ± {moe:0.0f} (at {conf*100:0.0f}% confidence level)')
# -
# We are 95% confident that this interval contains the true population parameter value. That is, if we were to repeat this process many times (sampling then computing CI), on average 95% of the CIs would contain the true population parameter value (and 5% wouldn't).
# now it's your turn
# try different sample sizes and alpha levels: how do these change the confidence interval's size?
# now it's your turn
# randomly sample 100 tract-level median home values then calculate the mean and 99% confidence interval
# ### 1b. *t*-tests: difference in means
#
# Is the difference between two groups statistically significant?
# choose a variable
var = 'med_home_value'
# create two data subsets
black_tracts = tracts[tracts['pct_black'] > 50]
group1 = black_tracts[var]
hispanic_tracts = tracts[tracts['pct_hispanic'] > 50]
group2 = hispanic_tracts[var]
# what are the probability distributions of these two data sets?
fig, ax = plt.subplots()
ax = group1.plot.kde(ls='--', c='k', alpha=0.5, lw=2, bw_method=0.7)
ax = group2.plot.kde(ls='-', c='k', alpha=0.5, lw=2, bw_method=0.7, ax=ax)
ax.set_xlim(left=0)
ax.set_ylim(bottom=0)
plt.show()
print(int(group1.mean()))
print(int(group2.mean()))
# calculate difference in means
diff = group1.mean() - group2.mean()
diff
# compute the t-stat and its p-value
t_statistic, p_value = stats.ttest_ind(group1, group2, equal_var=False, nan_policy='omit')
p_value
# is the difference in means statistically significant?
alpha = 0.05 #significance level
p_value < alpha
# now it's your turn
# what is the difference in mean tract-level median home values in majority white vs majority black tracts?
# is it statistically significant?
# what if you randomly sample just 25 tracts from each group: is their difference significant?
# ## 2. Statistical models
#
# Introduction to OLS linear regression.
#
# Lots to cover in a course on regression that we must skip for today's quick overview. But in general you'd want to:
#
# - specify a model (or alternative models) based on theory
# - inspect candidate predictors' relationships with the response
# - inspect the predictors' relationships with each other (and reduce multicollinearity)
# - transform predictors for better linearity
# - identify and handle outlier observations
# - regression diagnostics
# ### 2a. Simple (bivariate) linear regression
#
# OLS regression with a single predictor
# choose a response variable and drop any rows in which it is null
response = 'med_home_value'
tracts = tracts.dropna(subset=[response])
# create design matrix containing predictors (drop nulls), and a response variable vector
predictors = 'med_household_income'
X = tracts[predictors].dropna()
y = tracts.loc[X.index][response]
# estimate a simple linear regression model with scipy
m, b, r, p, se = stats.linregress(x=X, y=y)
print('m={:.4f}, b={:.4f}, r^2={:.4f}, p={:.4f}'.format(m, b, r ** 2, p))
# estimate a simple linear regression model with statsmodels
Xc = add_constant(X)
model = sm.OLS(y, Xc)
result = model.fit()
print(result.summary())
# This single predictor explains about half the variation of the response. To explain more, we need more predictors.
#
# ### 2b. Multiple regression
#
# OLS regression with multiple predictors
# create design matrix containing predictors (drop nulls), and a response variable vector
predictors = ['med_household_income', 'pct_white']
X = tracts[predictors].dropna()
y = tracts.loc[X.index][response]
# estimate a linear regression model
Xc = add_constant(X)
model = sm.OLS(y, Xc)
result = model.fit()
print(result.summary())
# #### statsmodels diagnostic output
#
# We discuss diagnostics and standardized regression in more detail below, but here's a quick summary of the output above:
#
# If we get warnings about multicollinearity, but have good VIF scores and significant variables, then check a standardized regression (below) to see if it's just scaling or the intercept/constant causing it (intercept shouldn't cause high condition number if we center/standardize our predictors). A high condition number indicates multicollinearity.
#
# Durbin-Watson tests for autocorrelation: a value around 1.5 to 2.5 is considered fine.
#
# Omnibus tests for normality of residuals: if prob < 0.05, we reject the null hypothesis that they are normally distributed (skew and kurtosis describe their distribution)
#
# Jarque-Bera tests for normality of residuals: if prob < 0.05, we reject the null hypothesis that they are normally distributed
# #### Now add in more variables...
tracts.columns
# create design matrix containing predictors (drop nulls), and a response variable vector
predictors = ['med_household_income', 'pct_white', 'pct_single_family_home', 'pct_built_before_1940',
'med_rooms_per_home', 'pct_bachelors_degree']
X = tracts[predictors].dropna()
y = tracts.loc[X.index][response]
# estimate a linear regression model
Xc = add_constant(X)
model = sm.OLS(y, Xc)
result = model.fit()
print(result.summary())
# now it's your turn
# try different sets of predictors to increase R-squared while keeping the total number of predictors relatively low and theoretically sound
# ### 2c. Standardized regression
#
# *Beta coefficients* are the estimated regression coefficients when the response and predictors are standardized so that their variances equal 1. Thus, we can interpret these coefficients as how many standard deviations the response changes for each standard deviation increase in the predictor. This tells us about "effect size": which predictors have greater effects on the response by ignoring the variables' different units/scales of measurement. However, it relies on the variables' distributions having similar shapes (otherwise the meaning of a std dev in one will differ from a std dev in another).
# estimate a standardized regression model
y_stdrd = pd.Series(stats.mstats.zscore(y), index=y.index, name=y.name)
X_stdrd = pd.DataFrame(stats.mstats.zscore(X), index=X.index, columns=X.columns)
Xc_stdrd = add_constant(X_stdrd)
model_stdrd = sm.OLS(y_stdrd, Xc_stdrd)
result_stdrd = model_stdrd.fit()
print(result_stdrd.summary())
# ### 2d. Diagnostics
#
# Let's take a step back and think about some of the steps we might take prior to specifying the model, and then to diagnose its fit.
# correlation matrix
# how well are predictors correlated with response... and with each other?
correlations = tracts[[response] + sorted(predictors)].corr()
correlations.round(2)
# visual correlation matrix via seaborn heatmap
# use vmin, vmax, center to set colorbar scale properly
sns.set(style='white')
ax = sns.heatmap(correlations, vmin=-1, vmax=1, center=0,
cmap=plt.cm.coolwarm, square=True, linewidths=1)
# plot pairwise relationships with seaborn
grid = sns.pairplot(tracts[[response] + sorted(predictors)], markers='.')
# **Actual vs Predicted**: how well do our model's predicted y values match up to the actual y values? Is the variance the same throughout (homoskedastic)? Point's distance from line is the residual (difference between actual value and predicted value).
# +
# plot observed (y-axis) vs fitted (x-axis)
observed = model.endog #actual response
fitted = result.fittedvalues #predicted response
fig, ax = plt.subplots(figsize=(6, 6))
ax.scatter(x=fitted, y=observed, s=0.2)
# draw a 45° y=x line
ax.set_xlim((min(np.append(observed, fitted)), max(np.append(observed, fitted))))
ax.set_ylim((min(np.append(observed, fitted)), max(np.append(observed, fitted))))
ax.plot(ax.get_xlim(), ax.get_ylim(), ls='--', c='k', alpha=0.5)
ax.set_xlabel('predicted values')
ax.set_ylabel('actual values')
plt.show()
# -
# **Residual Plot**: plot our residuals to look for heteroskedasticity. We want this plot to resemble a random point pattern with no discernable trend. If the spread grows as you move from left to right, you are seeing heteroskedasticity.
# +
# standardized (internally studentized) residuals
resids_stud = result.get_influence().resid_studentized_internal
fig, ax = plt.subplots(figsize=(6, 6))
ax.scatter(x=result.fittedvalues, y=resids_stud, s=0.2)
ax.axhline(y=0, ls='--', c='k', alpha=0.5)
ax.set_title('residuals vs fitted plot')
ax.set_xlabel('fitted values')
ax.set_ylabel('standardized residuals')
plt.show()
# -
# **QQ-Plot**: are the residuals approximately normally distributed? That is, how well do they match a theoretical normal distribution. We want the points to follow the line.
fig, ax = plt.subplots(figsize=(6, 6))
fig = sm.qqplot(resids_stud, line='45', ax=ax)
ax.set_title('normal probability plot of the standardized residuals')
plt.show()
# ^^ looks like we've got a problem with our model! Can we improve it any with a transformation?
# estimate a linear regression model
Xc = add_constant(X)
model = sm.OLS(np.log(y), Xc)
result = model.fit()
#print(result.summary())
resids_stud = result.get_influence().resid_studentized_internal
fig, ax = plt.subplots(figsize=(6, 6))
fig = sm.qqplot(resids_stud, line='45', ax=ax)
ax.set_title('normal probability plot of the standardized residuals')
plt.show()
# **Multicollinearity**: inspecting correlation among the predictors with condition number and VIF
# calculate condition numbers
print(np.linalg.cond(Xc))
print(np.linalg.cond(X))
print(np.linalg.cond(stats.mstats.zscore(X)))
# A high condition number indicates multicollinearity. Rule of thumb, you want this to be below ~20 (in real-world applied analyses it will often be a bit higher though). Condition number is the ratio of the largest eigenvalue in the design matrix to the smallest. In other words, the large condition number in this case results from scaling rather than from multicollinearity. If we have just one variable with units in the thousands (ie, a large eigenvalue) and add a constant with units of 1 (ie, a small eigenvalue), we'll get a large condition number as the ratio, and statsmodels warns of multicollinearity. If you standardize the design matrix, you see condition number without the scaling effects.
#
# VIF is a measure for the collinearity of one variable with all the others. As a rule of thumb, a VIF > 10 indicates strong multicollinearity. If multicollinearity is present in our regression model, the correlated predictors can have large standard errors and thus become insignificant, even though they are theoretically important. By removing redundant predictors, we'll have more sensible regression results for the ones we left in. In statsmodels, the function expects the presence of a constant in the matrix of explanatory variables.
# calculate VIFs for all predictors then view head
vif_values = [vif(X.values, i) for i in range(len(X.columns))]
vifs = pd.Series(data=vif_values, index=X.columns).sort_values(ascending=False).head()
vifs
# remove the worst offender from the design matrix
# ...but is this theoretically sound?
highest_vif = vifs.index[0]
X = X.drop(highest_vif, axis='columns')
# re-calculate VIFs
vif_values = [vif(X.values, i) for i in range(len(X.columns))]
vifs = pd.Series(data=vif_values, index=X.columns).sort_values(ascending=False).head()
vifs
# estimate a linear regression model
Xc = add_constant(X)
model = sm.OLS(y, Xc)
result = model.fit()
print(result.summary())
# now it's your turn
# try removing variables from the set of predictors, or transforming them, then re-calculate VIFs
# can you find a set of predictors that makes good theoretical sense and has less multicollinearity?
# ## 3. Spatial models
#
# Basic types:
#
# - **Spatial heterogeneity**: account for systematic differences across space without explicitly modeling interdependency (non-spatial estimation)
# - spatial fixed effects (intercept varies for each spatial group)
# - spatial regimes (intercept and coefficients vary for each spatial group)
# - **Spatial dependence**: model interdependencies between observations through space
# - spatial lag model (spatially-lagged endogenous variable added as predictor; because of endogeneity, cannot use OLS to estimate)
# - spatial error model (spatial effects in error term)
# - spatial lag+error combo model
# ### 3a. Spatial fixed effects
#
# Using dummy variables representing the counties into which our observations (tracts) are nested
# create a new dummy variable for each county, with 1 if tract is in this county and 0 if not
for county in tracts['COUNTYFP'].unique():
new_col = f'dummy_county_{county}'
tracts[new_col] = (tracts['COUNTYFP'] == county).astype(int)
# remove one dummy from dummies to prevent perfect collinearity
# ie, a subset of predictors sums to 1 (which full set of dummies will do)
county_dummies = [f'dummy_county_{county}' for county in tracts['COUNTYFP'].unique()]
county_dummies = county_dummies[1:]
# create design matrix containing predictors (drop nulls), and a response variable vector
predictors = ['med_household_income', 'pct_white', 'pct_single_family_home', 'pct_built_before_1940',
'med_rooms_per_home', 'pct_bachelors_degree']
X = tracts[predictors + county_dummies].dropna()
y = tracts.loc[X.index][response]
# estimate a linear regression model
Xc = add_constant(X)
model = sm.OLS(y, Xc)
result = model.fit()
print(result.summary())
# ### 3b. Spatial regimes
#
# Each spatial regime can have different model coefficients. Here, the regimes are counties. We'll take a subset of our data (all the tracts appearing in 3 counties). This subsection just uses OLS for estimation, but you can also combine spatial regimes with spatial autogression models (the latter is introduced later).
# pick 3 counties as the regimes, and only estimate a regimes model for this subset
counties = tracts['COUNTYFP'].value_counts().index[:3]
mask = tracts['COUNTYFP'].isin(counties)
# create design matrix containing predictors (drop nulls), a response variable matrix, and a regimes vector
X = tracts.loc[mask, predictors].dropna() #only take rows in the 3 counties
Y = tracts.loc[X.index][[response]] #notice this is a matrix this time for pysal
regimes = tracts.loc[X.index]['COUNTYFP'] #define the regimes
# estimate spatial regimes model with OLS
olsr = ps.model.spreg.OLS_Regimes(y=Y.values, x=X.values, regimes=regimes.values, name_regimes='county',
name_x=X.columns.tolist(), name_y=response, name_ds='tracts')
print(olsr.summary)
# ### 3c. Spatial diagnostics
#
# So far we've seen two spatial heterogeneity models. Now we'll explore spatial dependence, starting by using queen-contiguity spatial weights to model spatial relationships between observations and OLS to check diagnostics.
# create design matrix containing predictors (drop nulls), and a response variable matrix
predictors = ['med_household_income', 'pct_white', 'pct_single_family_home', 'pct_built_before_1940',
'med_rooms_per_home', 'pct_bachelors_degree']
X = tracts[predictors].dropna()
Y = tracts.loc[X.index][[response]] #notice this is a matrix this time for pysal
# compute spatial weights from tract geometries (but only those tracts that appear in design matrix!)
W = ps.lib.weights.Queen.from_dataframe(tracts.loc[X.index])
W.transform = 'r'
# compute OLS spatial diagnostics to check the nature of spatial dependence
ols = ps.model.spreg.OLS(y=Y.values, x=X.values, w=W, spat_diag=True, moran=True)
# calculate moran's I (for the response) and its significance
mi = ps.explore.esda.Moran(y=Y, w=W, two_tailed=True)
print(mi.I)
print(mi.p_sim)
# moran's I (for the residuals): moran's i, standardized i, p-value
ols.moran_res
# #### Interpreting the results
#
# A significant Moran's *I* suggests spatial autocorrelation, but doesn't tell us which alternative specification should be used. Lagrange Multiplier (LM) diagnostics can help with that. If one LM test is significant and the other isn't, then that tells us which model specification (spatial lag vs spatial error) to use:
# lagrange multiplier test for spatial lag model: stat, p
ols.lm_lag
# lagrange multiplier test for spatial error model: stat, p
ols.lm_error
# #### Interpreting the results
#
# If (and only if) both the LM tests produce significant statistics, try the robust versions (the nonrobust LM tests are sensitive to each other):
# robust lagrange multiplier test for spatial lag model: stat, p
ols.rlm_lag
# robust lagrange multiplier test for spatial error model: stat, p
ols.rlm_error
# #### So... which model specification to choose?
#
# If neither LM test is significant: use regular OLS.
#
# If only one LM test is significant: use that model spec.
#
# If both LM tests are significant: run robust versions.
#
# If only one robust LM test is significant: use that model spec.
#
# If both robust LM tests are significant (this can often happen with large sample sizes):
#
# - first consider if the initial model specification is actually a good fit
# - if so, use the spatial specification corresponding to the larger robust-LM statistic
# - or consider a combo model
#
# ### 3d. Spatial lag model
#
# When the diagnostics indicate the presence of a spatial diffusion process.
#
# Model specification:
#
# $y = \rho W y + X \beta + u$
#
# where $y$ is a $n \times 1$ vector of observations (response), $W$ is a $n \times n$ spatial weights matrix (thus $Wy$ is the spatially-lagged response), $\rho$ is the spatial autoregressive parameter to be estimated, $X$ is a $n \times k$ matrix of observations (exogenous predictors), $\beta$ is a $k \times 1$ vector of parameters (coefficients) to be estimated, and $u$ is a $n \times 1$ vector of errors.
# maximum-likelihood estimation with full matrix expression
mll = ps.model.spreg.ML_Lag(y=Y.values, x=X.values, w=W, method='full', name_w='queen',
name_x=X.columns.tolist(), name_y=response, name_ds='tracts')
print(mll.summary)
# the spatial autoregressive parameter estimate, rho
mll.rho
# ### 3e. Spatial error model
#
# When the diagnostics indicate the presence of spatial error dependence.
#
# Model specification:
#
# $y = X \beta + u$
#
# where $X$ is a $n \times k$ matrix of observations (exogenous predictors), $\beta$ is a $k \times 1$ vector of parameters (coefficients) to be estimated, and $u$ is a $n \times 1$ vector of errors. The errors $u$ follow a spatial autoregressive specification:
#
# $u = \lambda Wu + \epsilon$
#
# where $\lambda$ is a spatial autoregressive parameter to be estimated and $\epsilon$ is the vector of errors.
# maximum-likelihood estimation with full matrix expression
mle = ps.model.spreg.ML_Error(y=Y.values, x=X.values, w=W, method='full', name_w='queen',
name_x=X.columns.tolist(), name_y=response, name_ds='tracts')
print(mle.summary)
# the spatial autoregressive parameter estimate, lambda
mle.lam
# ### 3f. Spatial lag+error combo model
#
# Estimated with GMM (generalized method of moments). Essentially a spatial error model with endogenous explanatory variables.
#
# Model specification:
#
# $y = \rho W y + X \beta + u$
#
# where $y$ is a $n \times 1$ vector of observations (response), $W$ is a $n \times n$ spatial weights matrix (thus $Wy$ is the spatially-lagged response), $\rho$ is the spatial autoregressive parameter to be estimated, $X$ is a $n \times k$ matrix of observations (exogenous predictors), $\beta$ is a $k \times 1$ vector of parameters (coefficients) to be estimated, and $u$ is a $n \times 1$ vector of errors.
#
# The errors $u$ follow a spatial autoregressive specification:
#
# $u = \lambda Wu + \epsilon$
#
# where $\lambda$ is a spatial autoregressive parameter to be estimated and $\epsilon$ is the vector of errors.
gmc = ps.model.spreg.GM_Combo_Het(y=Y.values, x=X.values, w=W, name_w='queen', name_ds='tracts',
name_x=X.columns.tolist(), name_y=response)
print(gmc.summary)
# now it's your turn
# with a new set of predictors, compute spatial diagnostics and estimate a new spatial model accordingly
| modules/module10 - inferential spatial models/module10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow]
# language: python
# name: conda-env-tensorflow-py
# ---
# # Preparing the Data
# +
# Set your working directory
from os import getcwd
import pandas as pd
import numpy as np
import pickle as pk
from nltk import stem
import json
import matplotlib.pyplot as plt
import os
import json
fpath = getcwd()
print (fpath)
# -
datapath = fpath + '/data/'
yjson = json.load(open(datapath + "yelp_2.json", "rb")) # there are 35 reviews in this json
reviews = {}
i=0
for key0, val0 in yjson.items(): # keys are 0, 20 etc..
for val in val0["review"]:
reviews[i]= {"rating" : val["reviewRating"]["ratingValue"],
"review" : val["description"]}
i += 1
print ("Total no of reviews " + str(len(reviews)))
# ## Collate Scraped Yelp Reviews - Train
# +
# Read train data (6 restaurants out of 10 restaurants for each category) and collate
total_reviews_train = {}
directory = './data/train/'
for filename in os.listdir(directory):
if filename.endswith(".json"):
print(filename)
yjson = json.load(open(directory + filename, "rb"))
reviews = {}
i=0
for key0, val0 in yjson.items(): # keys are 0, 20 etc..
for val in val0["review"]:
reviews[i]= {"rating" : val["reviewRating"]["ratingValue"],
"review" : val["description"]}
i += 1
if bool(total_reviews_train)==False:
total_reviews_train = dict(reviews)
else:
# need to re-enumerate the numbers here, else it will overwrite the total_reviews by key!
reviews_to_update = {}
cnt = len(total_reviews_train)
for k,v in reviews.items():
reviews_to_update[str(cnt)] = v
cnt = cnt + 1
total_reviews_train.update(reviews_to_update)
#print(total_reviews)
print("Total train data = " + str(len(total_reviews_train)))
# with open('train_scraped.json', 'w') as outfile:
# json.dump(total_reviews_train, outfile)
# -
# ## Collate Scraped Yelp Reviews - Test (do by category)
# +
# Read train data (6 restaurants out of 10 restaurants for each category) and collate
def create_dict_from_folder(directory):
total_reviews_test = {}
#directory = './data/test/'
for filename in os.listdir(directory):
if filename.endswith(".json"):
print(filename)
yjson = json.load(open(directory + filename, "rb"))
reviews = {}
i=0
for key0, val0 in yjson.items(): # keys are 0, 20 etc..
for val in val0["review"]:
reviews[i]= {"rating" : val["reviewRating"]["ratingValue"],
"review" : val["description"],
"id" : val0["id"],
"name" : val0["name"],
"category" : val0["category"]
}
i += 1
if bool(total_reviews_test)==False:
total_reviews_test = dict(reviews)
else:
# need to re-enumerate the numbers here, else it will overwrite the total_reviews by key!
reviews_to_update = {}
cnt = len(total_reviews_test)
for k,v in reviews.items():
reviews_to_update[str(cnt)] = v
cnt = cnt + 1
total_reviews_test.update(reviews_to_update)
#print(total_reviews)
print("Total test data = " + str(len(total_reviews_test)))
# with open('test_scraped.json', 'w') as outfile:
# json.dump(total_reviews_test, outfile)
return total_reviews_test
chinese = create_dict_from_folder('./data/test/chinese/')
indian = create_dict_from_folder('./data/test/indian/')
japanese = create_dict_from_folder('./data/test/japanese/')
french = create_dict_from_folder('./data/test/french/')
# df_chinese = create_df_from_folder('./data/test/chinese/')
# df_chinese
# -
# ## Create the class labels -1 (neg), 1 (pos)
# Note that we should not use lecturer-supplied train_neg.csv and train_pos.csv as the criteria for -ve and +ve may be different.
#
# We need to build our own training set based on what we scraped as well as the yelp_2.json
#
# Use the ratings 1-3 as negative and 4-5 as positive in the Yelp reviews.
# +
def mapPosNegR(score):
rate = 1
if score <= 3: rate = -1 # the result is dependent on the map fn
return rate
actuals = []
for key, val in total_reviews_train.items(): # going through each review
actuals.append(val['rating'])
actuals = list(map(mapPosNegR, actuals)) # actuals in terms of +1 or -1
# -
df_reviews_train = pd.DataFrame(total_reviews_train).transpose()
df_reviews_train = df_reviews_train.reset_index().drop('index', axis=1)
df_actuals = pd.DataFrame(actuals)
df_reviews_train = pd.concat([df_reviews_train, pd.DataFrame(actuals)], axis=1)
#df_reviews.columns = {'rating', 'review', 'sentiment'}
#df_reviews_train
df_reviews_train.rename(columns={0:'sentiment'},
inplace=True)
df_reviews_train.head()
# Most reviews here are positive according to our rating criteria: it is an imbalanced set.
df_reviews_train.groupby('sentiment').sum()
df_reviews_train.tail()
# +
# now test
def process_to_df(dict_in):
actuals = []
for key, val in dict_in.items(): # going through each review
actuals.append(val['rating'])
actuals = list(map(mapPosNegR, actuals)) # actuals in terms of +1 or -1
df_reviews_test = pd.DataFrame(dict_in).transpose()
df_reviews_test = df_reviews_test.reset_index().drop('index', axis=1)
df_reviews_test = pd.concat([df_reviews_test, pd.DataFrame(actuals)], axis=1)
df_reviews_test.rename(columns={0:'sentiment'},
inplace=True)
return df_reviews_test
df_chinese = process_to_df(chinese)
df_japanese = process_to_df(japanese)
df_indian = process_to_df(indian)
df_french = process_to_df(french)
# -
df_chinese
df_reviews_test.groupby('sentiment').sum()
# +
df_reviews_train.to_csv('./data/df_reviews_train.csv', index=False)
#df_reviews_test.to_csv('./data/df_reviews_test.csv', index=False)
df_chinese.to_csv('./data/df_test_chinese.csv', index=False)
df_japanese.to_csv('./data/df_test_japanese.csv', index=False)
df_indian.to_csv('./data/df_test_indian.csv', index=False)
df_french.to_csv('./data/df_test_french.csv', index=False)
# -
| data_preparation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import findspark
findspark.init()
#spark connection and import all required packages
from pyspark.sql import SparkSession,types
spark = SparkSession.builder.master("local").appName("Create DataFrame").getOrCreate()
sc = spark.sparkContext
from datetime import datetime, date
rdd = spark.sparkContext.parallelize([
(1, 2., 'su1', date(2001, 4, 1), datetime(2001, 4, 1, 12, 0)),
(2, 3., 'su3', date(2002, 6, 2), datetime(2002, 6, 2, 12, 0)),
(3, 4., 'su5', date(2003, 3, 3), datetime(2003, 3, 3, 12, 0))
])
df = spark.createDataFrame(rdd, schema=['a', 'b', 'c', 'd', 'e'])
df
df.show()
| Create PySpark DataFrame from RDD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Chapter 6 - Inferring a Binomial Probability via Exact Mathematical Analysis
# +
import numpy as np
from scipy.stats import beta
from scipy.special import beta as beta_func
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('seaborn-white')
color = '#87ceeb'
# -
# ### Set up
# Here, we create a vector of values that theta can take on. This vector is **for plotting only**. We are calculating exact solutions here, so we will have priors, likelihoods, and posteriors for essentially all of the infinite number of number of values that theta can take on.
# +
n_theta_vals = 1001
theta = np.linspace(0, 1, n_theta_vals)
# -
# ### Prior
# We will use a beta distribution to describe our prior beliefs about the values of $\theta$. The use of a beta distribution to represent our prior belief about theta is critical here, because the beta distribution is the conjugate prior probability distribution for the Bernoulli likelihood that we used in chapter 5 and will reuse below.
#
# In addition, the beta distribution is flexible enough to permit several different patterns including:
#
# - $\alpha=1, \beta=1$ yields a uniform prior
# - $\alpha=3, \beta=3$ yields a prior peaked at $\theta=0.5$ (a bit like the truncated normal we used in chapter 5)
# +
a = 3
b = 3
p_theta = beta.pdf(theta, a, b)
# -
# ### Data
# This constructs a set of flip outcomes. Specify the number of heads (i.e., `n_heads`) and the number of tails (i.e., `n_tails`). There are three scenarios prepared:
#
# 1. 1 flip that comes up heads
# 2. 4 flips, 1 of which comes up heads (25% heads)
# 3. 40 flips, 10 of which come up heads (25% heads)
# +
# example 1
n_heads = 1
n_tails = 0
# example 2
#n_heads = 1
#n_tails = 3
# example 3
#n_heads = 10
#n_tails = 30
data = np.repeat([1, 0], [n_heads, n_tails])
# -
# ### Likelihood
# Note that we are using the vector of theta values here this is because we want to plot the likelihood function below we **do not** need these lines of code in order to arrive at the posterior (as we will see).
# +
# Compute the likelihood of the data:
p_data_given_theta = theta**n_heads * (1-theta)**(n_tails)
# calculate the evidence (P(D), the prior probability of the data)
p_data = beta_func(n_heads + a, n_tails + b)/beta_func(a, b)
# -
# ### Inference
# Here is the magic of using priors that are conjugate with our likelihood. Because we are using a beta prior, we can straightforwardly determine the posterior by adding the number of heads/tails to the $\alpha$ and $\beta$ parameters we used to construct our prior.
# +
post_a = n_heads + a
post_b = n_tails + b
# Compute the posterior for our values of theta for later visualization
p_theta_given_data = beta.pdf(theta, post_a, post_b)
# -
# ### Visualize
# Plot the prior, the likelihood, and the posterior.
# +
fig = plt.figure(figsize=(9, 12))
fig.tight_layout()
plt.subplots_adjust(hspace = .4)
# Plot the prior, the likelihood, and the posterior:
for i,dist in enumerate([p_theta, p_data_given_theta, p_theta_given_data]):
plt.subplot(3, 1, i+1)
plt.plot(theta, dist)
plt.xlim(0, 1)
plt.xlabel('$\\theta$', size=16)
# horizontal location of text labels
locx = 0.1
# prior
plt.axes(fig.axes[0])
plt.title('Prior', weight='bold', size=16)
plt.xlim(0, 1)
plt.ylim(0, np.max(p_theta)*1.2)
plt.ylabel(r'$P(\theta)$', size=16)
plt.text(locx, np.max(p_theta)/2, r'beta(%s,%s)' % (a, b), size=16)
# likelihood
plt.axes(fig.axes[1])
plt.title('Likelihood', weight='bold', size=16)
plt.ylabel('$P(D|\\theta)$', size=16)
plt.text(locx, np.max(p_data_given_theta)/2, 'D = %sH,%sT' % (n_heads, n_tails), size=16)
# posterior
plt.axes(fig.axes[2])
plt.title('Posterior', weight='bold', size=16)
plt.ylabel('$P(\\theta|D)$', size=16)
locy = np.linspace(0, np.max(p_theta_given_data), 5)
plt.text(locx, locy[1], r'beta(%s,%s)' % (post_a, post_b), size=16)
plt.text(locx, locy[2], 'P(D) = %.2f' % p_data, size=16)
| Notebooks/Chapter 6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Research2018
# language: python
# name: research
# ---
# +
import os
import numpy as np
import pandas as pd
from tqdm import tqdm
import torch
from datasets import *
from utils import *
from scipy import sparse
from model.EdgeReg import *
from model.EdgeReg_v2 import *
# -
gpunum = "2"
nbits = 128
os.environ["CUDA_VISIBLE_DEVICES"]=gpunum
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
# +
num_samples = 1
dataset_name = 'pubmed'
data_dir = os.path.join('dataset/clean', dataset_name)
train_batch_size=100
test_batch_size=100
train_set = TextDataset(dataset_name, data_dir, subset='train')
test_set = TextDataset(dataset_name, data_dir, subset='test')
train_loader = torch.utils.data.DataLoader(dataset=train_set, batch_size=train_batch_size, shuffle=True)
test_loader = torch.utils.data.DataLoader(dataset=test_set, batch_size=test_batch_size, shuffle=True)
# -
y_dim = train_set.num_classes()
num_bits = nbits
num_features = train_set[0][1].size(0)
num_nodes = len(train_set)
edge_weight = 1.0
dropout_prob = 0.1
if num_samples == 1:
model = EdgeReg(dataset_name, num_features, num_nodes, num_bits, dropoutProb=dropout_prob, device=device)
else:
print("number of samples (T) = {}".format(num_samples))
model = EdgeReg_v2(dataset_name, num_features, num_nodes, num_bits, dropoutProb=dropout_prob, device=device, T=num_samples)
# +
if num_samples == 1:
saved_model_file = 'saved_models/node2hash.{}.T{}.bit{}.pth'.format(dataset_name, num_samples, nbits)
else:
saved_model_file = 'saved_models/node2hash_v2.{}.T{}.bit{}.pth'.format(dataset_name, num_samples, nbits)
model.load_state_dict(torch.load(saved_model_file))
model.to(device)
# +
import torch.nn.functional as F
# get non-binary code
with torch.no_grad():
train_zy = [(model.encode(xb.to(model.device))[0], yb) for _, xb, yb, _ in train_loader]
train_z, train_y = zip(*train_zy)
train_z = torch.cat(train_z, dim=0)
train_y = torch.cat(train_y, dim=0)
test_zy = [(model.encode(xb.to(model.device))[0], yb) for _, xb, yb, _ in test_loader]
test_z, test_y = zip(*test_zy)
test_z = torch.cat(test_z, dim=0)
test_y = torch.cat(test_y, dim=0)
train_z_batch = train_z.unsqueeze(-1).transpose(2,0)
test_z_batch = test_z.unsqueeze(-1)
# compute cosine similarity
dist = F.cosine_similarity(test_z_batch, train_z_batch, dim=1)
ranklist = torch.argsort(dist, dim=1, descending=True)
top100 = ranklist[:, :100]
prec_at_100 = []
for eval_index in range(0, test_y.size(0)):
top100_labels = torch.index_select(train_y.to(device), 0, top100[eval_index]).type(torch.cuda.ByteTensor)
groundtruth_label = test_y[eval_index].type(torch.cuda.ByteTensor)
matches = (groundtruth_label.unsqueeze(0) & top100_labels).sum(dim=1) > 0
num_corrects = matches.sum().type(torch.cuda.FloatTensor)
prec_at_100.append((num_corrects/100.).item())
print('average prec at 100 = {:.4f}'.format(np.mean(prec_at_100)))
| Evaluation_Draft.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
s1=pd.Series([1,2,3,4,5,6],index=pd.date_range('20171101',periods=6))
s1
df=pd.DataFrame(np.random.randn(6,4),index=pd.date_range('20171101',periods=6),
columns=list('ABCD'))
df
df['F']=s1#Setting a new column automatically aligns the data by the indexes
df
df.at['20171101','A']=0#Setting values by label
df
df.iat[0,1]=0#Setting values by position
df
df.loc[:,'D']=np.array([5]*len(df))#Setting by assigning with a numpy array
df
df2=df.copy()
df2[df2>0]=-df2
df2
dates=pd.date_range('20171101',periods=6)
dates
df1=df.reindex(index=dates[0:4],columns=list(df.columns)+['E'])
df1.loc[dates[0]:dates[1],'E']=1
df1
df1.dropna(how='any')#To drop any rows that have missing data
df1.fillna(value=5)#Filling missing data
pd.isnull(df1)
df.mean()
df.mean(1)
s=pd.Series([1,3,5,np.nan,6,8],index=dates).shift(2)#后2个转换成nan,并变成前2个
s
df.sub(s,axis='index')#按照对应索引减去s
df
df.apply(np.cumsum)#按列累加
df.apply(lambda x :x.max()-x.min())
s=pd.Series(np.random.randint(0,7,size=10))
s
s.value_counts()
df['D'].value_counts()
s=pd.Series(['A','B','C','aVA','Bdc',np.nan,'CVSa'])
s.str.lower()
df5=pd.DataFrame(np.random.randn(5,4))
df5
pieces=[df5[:2],df5[2:4],df5[4:]]
pieces
pd.concat(pieces)
left=pd.DataFrame({'key':['foo','foo'],'lval':[1,2]})
right=pd.DataFrame({'key':['foo','foo'],'lval':[3,4]})
print(left)
print(right)
pd.merge(left,right,on='key')
left=pd.DataFrame({'key':['foo','bar'],'lval':[1,2]})
right=pd.DataFrame({'key':['foo','bar'],'rval':[3,4]})
pd.merge(left,right,on='key')
df.to_excel('test.xls')#Writing to an excel file
r=pd.date_range('20171101',periods=10,freq='H')
r
| pandas-10minutes-2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The art of using pipelines
# Pipelines are a natural way to think about a machine learning system. Indeed with some practice a data scientist can visualise data "flowing" through a series of steps. The input is typically some raw data which has to be processed in some manner. The goal is to represent the data in such a way that is can be ingested by a machine learning algorithm. Along the way some steps will extract features, while others will normalize the data and remove undesirable elements. Pipelines are simple, and yet they are a powerful way of designing sophisticated machine learning systems.
#
# Both [scikit-learn](https://stackoverflow.com/questions/33091376/python-what-is-exactly-sklearn-pipeline-pipeline) and [pandas](https://tomaugspurger.github.io/method-chaining) make it possible to use pipelines. However it's quite rare to see pipelines being used in practice (at least on Kaggle). Sometimes you get to see people using scikit-learn's `pipeline` module, however the `pipe` method from `pandas` is sadly underappreciated. A big reason why pipelines are not given much love is that it's easier to think of batch learning in terms of a script or a notebook. Indeed many people doing data science seem to prefer a procedural style to a declarative style. Moreover in practice pipelines can be a bit rigid if one wishes to do non-orthodox operations.
#
# Although pipelines may be a bit of an odd fit for batch learning, they make complete sense when they are used for online learning. Indeed the UNIX philosophy has advocated the use of pipelines for data processing for many decades. If you can visualise data as a stream of observations then using pipelines should make a lot of sense to you. We'll attempt to convince you by writing a machine learning algorithm in a procedural way and then converting it to a declarative pipeline in small steps. Hopefully by the end you'll be convinced, or not!
#
# In this notebook we'll manipulate data from the [Kaggle Recruit Restaurants Visitor Forecasting competition](https://www.kaggle.com/c/recruit-restaurant-visitor-forecasting). The data is directly available through `river`'s `datasets` module.
# +
from pprint import pprint
from river import datasets
for x, y in datasets.Restaurants():
pprint(x)
pprint(y)
break
# -
# We'll start by building and running a model using a procedural coding style. The performance of the model doesn't matter, we're simply interested in the design of the model.
# +
from river import feature_extraction
from river import linear_model
from river import metrics
from river import preprocessing
from river import stats
means = (
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(7)),
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(14)),
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(21))
)
scaler = preprocessing.StandardScaler()
lin_reg = linear_model.LinearRegression()
metric = metrics.MAE()
for x, y in datasets.Restaurants():
# Derive date features
x['weekday'] = x['date'].weekday()
x['is_weekend'] = x['date'].weekday() in (5, 6)
# Process the rolling means of the target
for mean in means:
x = {**x, **mean.transform_one(x)}
mean.learn_one(x, y)
# Remove the key/value pairs that aren't features
for key in ['store_id', 'date', 'genre_name', 'area_name', 'latitude', 'longitude']:
x.pop(key)
# Rescale the data
x = scaler.learn_one(x).transform_one(x)
# Fit the linear regression
y_pred = lin_reg.predict_one(x)
lin_reg.learn_one(x, y)
# Update the metric using the out-of-fold prediction
metric.update(y, y_pred)
print(metric)
# -
# We're not using many features. We can print the last `x` to get an idea of the features (don't forget they've been scaled!)
pprint(x)
# The above chunk of code is quite explicit but it's a bit verbose. The whole point of libraries such as `river` is to make life easier for users. Moreover there's too much space for users to mess up the order in which things are done, which increases the chance of there being target leakage. We'll now rewrite our model in a declarative fashion using a pipeline *à la sklearn*.
# +
from river import compose
def get_date_features(x):
weekday = x['date'].weekday()
return {'weekday': weekday, 'is_weekend': weekday in (5, 6)}
model = compose.Pipeline(
('features', compose.TransformerUnion(
('date_features', compose.FuncTransformer(get_date_features)),
('last_7_mean', feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(7))),
('last_14_mean', feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(14))),
('last_21_mean', feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(21)))
)),
('drop_non_features', compose.Discard('store_id', 'date', 'genre_name', 'area_name', 'latitude', 'longitude')),
('scale', preprocessing.StandardScaler()),
('lin_reg', linear_model.LinearRegression())
)
metric = metrics.MAE()
for x, y in datasets.Restaurants():
# Make a prediction without using the target
y_pred = model.predict_one(x)
# Update the model using the target
model.learn_one(x, y)
# Update the metric using the out-of-fold prediction
metric.update(y, y_pred)
print(metric)
# -
# We use a `Pipeline` to arrange each step in a sequential order. A `TransformerUnion` is used to merge multiple feature extractors into a single transformer. The `for` loop is now much shorter and is thus easier to grok: we get the out-of-fold prediction, we fit the model, and finally we update the metric. This way of evaluating a model is typical of online learning, and so we put it wrapped it inside a function called `progressive_val_score` part of the `evaluate` module. We can use it to replace the `for` loop.
# +
from river import evaluate
model = compose.Pipeline(
('features', compose.TransformerUnion(
('date_features', compose.FuncTransformer(get_date_features)),
('last_7_mean', feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(7))),
('last_14_mean', feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(14))),
('last_21_mean', feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(21)))
)),
('drop_non_features', compose.Discard('store_id', 'date', 'genre_name', 'area_name', 'latitude', 'longitude')),
('scale', preprocessing.StandardScaler()),
('lin_reg', linear_model.LinearRegression())
)
evaluate.progressive_val_score(dataset=datasets.Restaurants(), model=model, metric=metrics.MAE())
# -
# Notice that you couldn't have used the `progressive_val_score` method if you wrote the model in a procedural manner.
#
# Our code is getting shorter, but it's still a bit difficult on the eyes. Indeed there is a lot of boilerplate code associated with pipelines that can get tedious to write. However `river` has some special tricks up it's sleeve to save you from a lot of pain.
#
# The first trick is that the name of each step in the pipeline can be omitted. If no name is given for a step then `river` automatically infers one.
# +
model = compose.Pipeline(
compose.TransformerUnion(
compose.FuncTransformer(get_date_features),
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(7)),
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(14)),
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(21))
),
compose.Discard('store_id', 'date', 'genre_name', 'area_name', 'latitude', 'longitude'),
preprocessing.StandardScaler(),
linear_model.LinearRegression()
)
evaluate.progressive_val_score(datasets.Restaurants(), model, metrics.MAE())
# -
# Under the hood a `Pipeline` inherits from `collections.OrderedDict`. Indeed this makes sense because if you think about it a `Pipeline` is simply a sequence of steps where each step has a name. The reason we mention this is because it means you can manipulate a `Pipeline` the same way you would manipulate an ordinary `dict`. For instance we can print the name of each step by using the `keys` method.
for name in model.steps:
print(name)
# The first step is a `FeatureUnion` and it's string representation contains the string representation of each of it's elements. Not having to write names saves up some time and space and is certainly less tedious.
#
# The next trick is that we can use mathematical operators to compose our pipeline. For example we can use the `+` operator to merge `Transformer`s into a `TransformerUnion`.
# +
model = compose.Pipeline(
compose.FuncTransformer(get_date_features) + \
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(7)) + \
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(14)) + \
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(21)),
compose.Discard('store_id', 'date', 'genre_name', 'area_name', 'latitude', 'longitude'),
preprocessing.StandardScaler(),
linear_model.LinearRegression()
)
evaluate.progressive_val_score(datasets.Restaurants(), model, metrics.MAE())
# -
# Likewhise we can use the `|` operator to assemble steps into a `Pipeline`.
# +
model = (
compose.FuncTransformer(get_date_features) +
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(7)) +
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(14)) +
feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(21))
)
to_discard = ['store_id', 'date', 'genre_name', 'area_name', 'latitude', 'longitude']
model = model | compose.Discard(*to_discard) | preprocessing.StandardScaler()
model |= linear_model.LinearRegression()
evaluate.progressive_val_score(datasets.Restaurants(), model, metrics.MAE())
# -
# Hopefully you'll agree that this is a powerful way to express machine learning pipelines. For some people this should be quite remeniscent of the UNIX pipe operator. One final trick we want to mention is that functions are automatically wrapped with a `FuncTransformer`, which can be quite handy.
# +
model = get_date_features
for n in [7, 14, 21]:
model += feature_extraction.TargetAgg(by='store_id', how=stats.RollingMean(n))
model |= compose.Discard(*to_discard)
model |= preprocessing.StandardScaler()
model |= linear_model.LinearRegression()
evaluate.progressive_val_score(datasets.Restaurants(), model, metrics.MAE())
# -
# Naturally some may prefer the procedural style we first used because they find it easier to work with. It all depends on your style and you should use what you feel comfortable with. However we encourage you to use operators because we believe that this will increase the readability of your code, which is very important. To each their own!
#
# Before finishing we can take a look at what our pipeline looks graphically.
model.draw()
| docs/examples/the-art-of-using-pipelines.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## < DRAFT > NFHP 2015 Total Stream Kms that were Scored in a Selected Spatial Unit
# #### <NAME> -USGS
#
# This notebook returns the number of stream kms within a spatial unit of interest (from the SFR placeNameLookup table). Calculations include:
# Total Stream Kms in spatial unit:
# Total Stream Kms scored in NFHP:
#
# Eventually I would like to also include
# Stream kms classed as river (> 100km2): Avg NFHP Score, total kms scored vs. not
# Stream kms classed as creek (< 100km2): Avg NFHP Score, total kms scored vs. not
#
#
#
# This code is in progress and may change through time.
#
# #### Generalized Steps
# #1.Access Data from GC2 (user or app passes in placeNameLookup and url specific to instance of gc2)
# Note: In NBM visualizations this uses Elastic Search. A version of this code should be updated to work against the ES endpoint.
# #2.Dispaly Data
#
def requestData(url, place):
import requests
query_disturbance = url+"?q=select place_name, scored_km, not_scored_km \
from nfhp.hci2015_summaries_mp where source_id='" + place + "'"
hci_disturbance = requests.get(url=query_disturbance).json()
data = hci_disturbance['features'][0]['properties']
return (data)
# +
#### Step 1: Request Data from GC2
url = 'https://beta-gc2.datadistillery.org/api/v1/sql/bcb'
place = 'doi lands:5324'
data = requestData (url, place)
print (data)
# -
# #### Step 2: Data to display
#
# +
#Dispaly the following information:
#Add scored and unscored kms to get total kms within spatial unit.
total_km = str(float(data['scored_km']) + float(data['not_scored_km']))
print ('Fish habitat condition was scored on ' + data['scored_km'] + ' of ' +total_km + ' NHDPlusV1 stream kms within ' + data['place_name']+'.' )
# -
# #Note within the larger NFHP BAP this could be displayed directly under the title of the BAP to give a user an understanding of the BAP.
| nfhp-2015-total-stream-km-scored-per-spatial-unit.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import matplotlib as mpl
from mpl_toolkits.mplot3d import Axes3D
from sklearn import tree
# ### Loading and sample data
#
# Synthetic Transactions: https://www.kaggle.com/ntnu-testimon/paysim1
# +
# Load file
dataset = pd.read_csv('data/PS_20174392719_1491204439457_log.csv')
trainset = dataset.sample(frac=0.5)
testset = dataset.sample(frac=0.5)
# -
# ### Treat data
# +
# Remove types that never are fraud
trainset = trainset.loc[trainset['type'] != 'PAYMENT']
trainset = trainset.loc[trainset['type'] != 'DEBIT']
trainset = trainset.loc[trainset['type'] != 'CASH_IN']
testset = testset.loc[testset['type'] != 'PAYMENT']
testset = testset.loc[testset['type'] != 'DEBIT']
testset = testset.loc[testset['type'] != 'CASH_IN']
# Map domain to numbers
trainset = trainset.replace("TRANSFER", 0)
trainset = trainset.replace("CASH_OUT", 1)
testset = testset.replace("TRANSFER", 0)
testset = testset.replace("CASH_OUT", 1)
# Treat step in hour, day and week
trainset['hour'] = trainset['step'].map(lambda x: x % 24)
trainset['day'] = trainset['step'].map(lambda x: int(x / 24))
trainset['week'] = trainset['step'].map(lambda x: int(x / 168))
testset['hour'] = testset['step'].map(lambda x: x % 24)
testset['day'] = testset['step'].map(lambda x: int(x / 24))
testset['week'] = testset['step'].map(lambda x: int(x / 168))
# Define labels
trainlabel = trainset['isFraud']
testlabel = testset['isFraud']
# Remove unused data
trainset = trainset.drop(columns=['nameOrig', 'nameDest', 'step', 'isFlaggedFraud', 'isFraud'])
testset = testset.drop(columns=['nameOrig', 'nameDest', 'step', 'isFlaggedFraud', 'isFraud'])
# -
# ### Train dataset information
# +
train_qtd = trainset.shape[0]
train_fraud_qtd = len(list(filter(lambda x: x == 1, trainset['isFraud'])))
train_nonfraud_qtd = len(list(filter(lambda x: x == 0, trainset['isFraud'])))
train_proportion = train_fraud_qtd / train_qtd
print('train_total:', train_qtd)
print('train_fraud:', train_fraud_qtd)
print('train_nonfraud', train_nonfraud_qtd)
print('train_fraud proportion', train_proportion)
trainset.head()
# -
# ### Test dataset information
# +
test_qtd = testset.shape[0]
test_fraud_qtd = len(list(filter(lambda x: x == 1, testset['isFraud'])))
test_nonfraud_qtd = len(list(filter(lambda x: x == 0, testset['isFraud'])))
test_proportion = test_fraud_qtd / test_qtd
print('test_total:', test_qtd)
print('test_fraud:', test_fraud_qtd)
print('test_nonfraud', test_nonfraud_qtd)
print('test_fraud proportion', test_proportion)
testset.head()
# -
# ### Divide Fraud x NonFraud
train_frauds = trainset.loc[trainset['isFraud'] == 1]
train_non_frauds = trainset.loc[trainset['isFraud'] == 0]
# ### Labs
# +
def plot(x_value, y_value, z_value):
fig = plt.figure()
fig.set_size_inches(16, 9)
ax = fig.add_subplot(111, projection='3d')
x_fraud = train_frauds[[x_value]]
y_fraud = train_frauds[[y_value]]
z_fraud = train_frauds[[z_value]]
x_nonfraud = train_non_frauds[[x_value]]
y_nonfraud = train_non_frauds[[y_value]]
z_nonfraud = train_non_frauds[[z_value]]
ax.scatter(x_fraud, y_fraud, z_fraud, c='C3')
ax.scatter(x_nonfraud, y_nonfraud, z_nonfraud, c='C0')
ax.set_xlabel(x_value)
ax.set_ylabel(y_value)
ax.set_zlabel(z_value)
plot('day', 'hour', 'amount')
# -
# ### Learning
clf = tree.DecisionTreeClassifier()
clf = clf.fit(trainset, trainlabel)
# ### Evaluate
accuracy = clf.score(testset, testlabel)
print(accuracy)
| fraud-detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''cref'': conda)'
# language: python
# name: python37664bitcrefconda60993f1b3fc94ad08cb7094813fa06fe
# ---
# %load_ext autoreload
# %autoreload 2
# +
import scipy
import time
import uuid
import matplotlib.pyplot as plt
import networkx as nx
from SPARQLWrapper import SPARQLWrapper, JSON, POST
from req_analysis import *
# -
# # Initialize connection to Neptune
# %run 'req_analysis/libs/neptune-util.py'
# %env NEPTUNE_CLUSTER_ENDPOINT=172.16.17.32
# %env NEPTUNE_CLUSTER_PORT=8182
g = neptune.graphTraversal()
# # Evaluation
sparql = SPARQLWrapper("https://cae-mms-rdf-test-r5-2x.cluster-cw2hjngge6pe.us-gov-west-1.neptune.amazonaws.com:8182/sparql")
ref_targets = ReferenceTargets(sparql)
ref_targets.init_table()
req_evaluator = RequirementEvaluator(sparql)
req_evaluator.init_requirement_elements()
req_evaluator.evaluate_all_requirements(g,
ref_targets,
max_evals=10,
pprint=False)
| 03_full_worker.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6.9 64-bit
# metadata:
# interpreter:
# hash: 4cd7ab41f5fca4b9b44701077e38c5ffd31fe66a6cab21e0214b68d958d0e462
# name: python3
# ---
# +
from pathlib import Path
import pandas as pd
import matplotlib as plt
data_path = Path("tfevents")
# +
networks = ["resnet101"]
# networks = ["resnet101", "resnet101_no_pretrained"]
scenaries = [100, 300, 500, 1000, 3000, 4874]
metric = "DetectionBoxes_Precision/mAP@.75IOU"
training_history = pd.DataFrame()
results = pd.DataFrame(columns = networks, index = scenaries)
for network in networks:
for scenary in scenaries:
data = pd.read_csv(data_path/network/f"{scenary}"/"converted"/f"eval{scenary}.csv")
training_history[f"{scenary}"] = data[metric]
best_result = data[metric].max()
results.loc[scenary, network] = best_result
training_history.index = 1000*(training_history.index + 1)
# -
ax = training_history.plot(figsize = (10, 6))
ax.grid(alpha = 0.3)
ax.set_xlabel("epoch")
ax.set_ylabel(metric)
ax.set_title("Test set evaluations fro ResNet-101")
ax = results.plot(style = "-o", figsize = (10, 6))
ax.grid(alpha = 0.3)
ax.set_title("Best results for each network at each training scenary", fontsize = 16)
ax.set_xlabel("# of training examples per class", fontsize = 14)
ax.set_ylabel(metric, fontsize = 14)
| results_graph.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import csv
import numpy as np
import pandas as pd
# import geopandas as gpd
from IPython.display import Image
# from shapely.geometry import Point, Polygon
from math import factorial
import datetime
import time
import scipy
import os, os.path
from statsmodels.sandbox.regression.predstd import wls_prediction_std
from sklearn.linear_model import LinearRegression
from patsy import cr
# from pprint import pprint
import matplotlib.pyplot as plt
import seaborn as sb
import sys
start_time = time.time()
# +
sys.path.append('/Users/hn/Documents/00_GitHub/Ag/remote_sensing/python/')
sys.path.append('/home/hnoorazar/remote_sensing_codes/')
import remote_sensing_core as rc
import remote_sensing_core as rcp
# +
data_base = "/Users/hn/Documents/01_research_data/remote_sensing/01_NDVI_TS/" + \
"04_Irrigated_eastern_Cloud70/Grant_2018_irrigated/" + \
"Grant_Irrigated_EVI_2018_NassIn_NotCorrectYears/"
data_base = "/Users/hn/Documents/01_research_data/remote_sensing/" + \
"01_NDVI_TS/04_Irrigated_eastern_Cloud70/Grant_2018_irrigated/" + \
"savitzky_EVI/Grant_Irrigated_2018_no_plot/"
param_dir = "/Users/hn/Documents/00_GitHub/Ag/remote_sensing/parameters/"
double_crop_potens = pd.read_csv(param_dir + "double_crop_potential_plants.csv")
# -
# Sav_win_size, sav_order, delt
parameters = [(3, 1, 0.1), (3, 1, 0.2), (3, 1, 0.3), (3, 1, 0.4),
(5, 1, 0.1), (5, 1, 0.2), (5, 1, 0.3), (5, 1, 0.4),
(7, 1, 0.1), (7, 1, 0.2), (7, 1, 0.3), (7, 1, 0.4),
(9, 1, 0.1), (9, 1, 0.2), (9, 1, 0.3), (9, 1, 0.4),
(3, 2, 0.1), (3, 2, 0.2), (3, 2, 0.3), (3, 2, 0.4),
(5, 2, 0.1), (5, 2, 0.2), (5, 2, 0.3), (5, 2, 0.4),
(7, 2, 0.1), (7, 2, 0.2), (7, 2, 0.3), (7, 2, 0.4),
(9, 2, 0.1), (9, 2, 0.2), (9, 2, 0.3), (9, 2, 0.4),
# (3, 3, 0.1), (3, 3, 0.2), (3, 3, 0.3), (3, 3, 0.4),
(5, 3, 0.1), (5, 3, 0.2), (5, 3, 0.3), (5, 3, 0.4),
(7, 3, 0.1), (7, 3, 0.2), (7, 3, 0.3), (7, 3, 0.4),
(9, 3, 0.1), (9, 3, 0.2), (9, 3, 0.3), (9, 3, 0.4)]
# +
# form a data table for output.
output_columns = ['params',
'NassIn_AllFields_AllYears',
'NassOut_AllFields_AllYears',
'NassIn_AllFields_CorrectYear',
'NassOut_AllFields_CorrectYear',
'NassIn_DoublePoten_AllYears',
'NassOut_DoublePoten_AllYears',
'NassIn_DoublePoten_CorrectYear',
'NassOut_DoublePoten_CorrectYear'
]
output_df = pd.DataFrame(data=None,
index=np.arange(len(parameters)),
# index = parameters,
columns = output_columns)
output_df['params'] = parameters
# -
output_df.shape
# +
# for num, param in enumerate(parameters):
# print("Parameter {}: {}".format(num, param))
# +
for num, param in enumerate(parameters):
Sav_win_size = param[0]
sav_order = param[1]
delt = param[2]
data_dir = data_base + "delta" + str(delt) + \
"_Sav_win" + str(Sav_win_size) + "_Order" + str(sav_order) + "/"
curr_table = pd.read_csv(data_dir + "all_poly_and_maxs_savitzky.csv", low_memory=False)
#
# drop last empty row
#
curr_table.drop(curr_table.tail(1).index, inplace=True)
"""
The data table includes all maximum information.
So, each field is repeated several times.
We need to get unique fields.
"""
curr_table.drop(['max_Doy', 'max_value'], axis=1, inplace=True)
curr_table.drop_duplicates(inplace=True)
###
### Pick those with more than two peaks in them
###
curr_table = curr_table[curr_table["max_count"] >= 2]
curr_table["DataSrc"] = curr_table["DataSrc"].str.lower()
##########################################################################################
##########################################################################################
##########################################################################################
##########################################################################################
##########################################################################################
###
### NassIn, AllFields, AllYears
### i.e. Everything other than those with
### no maximum detected on their time series.
###
NassIn_AllFields_AllYears_Acr = np.sum(curr_table['ExctAcr'])
# print ("1) is NASS in? should be ..." )
# print (curr_table.DataSrc.unique())
# print("_____________________________________________")
###
### NassOut_AllFields_AllYears
###
NassOut_AllFields_AllYears = curr_table[curr_table.DataSrc != 'nass'].copy()
# print ("2) is NASS in? should NOT be ...")
# print (NassOut_AllFields_AllYears.DataSrc.unique())
# print("_____________________________________________")
NassOut_AllFields_AllYears_Acr = np.sum(NassOut_AllFields_AllYears['ExctAcr'])
del(NassOut_AllFields_AllYears)
###
### NassIn AllFields CorrectYear
###
NassIn_AllFields_CorrectYear = curr_table[curr_table["LstSrvD"].str.contains("2018", na=False)].copy()
# print ("3) is NASS in? should be ...")
# print ( NassIn_AllFields_CorrectYear.DataSrc.unique())
# print("_____________________________________________")
NassIn_AllFields_CorrectYear_Acr = np.sum(NassIn_AllFields_CorrectYear['ExctAcr'])
del(NassIn_AllFields_CorrectYear)
###
### NassOut AllFields CorrectYear
###
NassOut_AllFields = curr_table[curr_table.DataSrc != 'nass'].copy()
NassOut_AllFields_CorrectYear = \
NassOut_AllFields[NassOut_AllFields["LstSrvD"].str.contains("2018", na=False)].copy()
# print ("4) is NASS in? should NOT be ...")
# print (NassOut_AllFields_CorrectYear.DataSrc.unique())
# print("_____________________________________________")
NassOut_AllFields_CorrectYear_Acr = np.sum(NassOut_AllFields_CorrectYear['ExctAcr'])
del(NassOut_AllFields, NassOut_AllFields_CorrectYear)
###############################################################
#####
##### double potentials
#####
###############################################################
curr_double_poten = curr_table[curr_table.CropTyp.isin(double_crop_potens['Crop_Type'])]
del(curr_table)
###
### NassIn, double potential, AllYears (i.e. Everything other than non-max)
###
NassIn_DoublePoten_AllYears_Acr = np.sum(curr_double_poten['ExctAcr'])
# print ("1) is NASS in? should be ...")
# print (curr_double_poten.DataSrc.unique())
# print("_____________________________________________")
###
### NassOut, double potential, AllYears
###
NassOut_DoublePoten_AllYears = curr_double_poten[curr_double_poten.DataSrc != 'nass'].copy()
NassOut_DoublePoten_AllYears_Acr = np.sum(NassOut_DoublePoten_AllYears['ExctAcr'])
# print ("2) is NASS in? should NOT be ...")
# print (NassOut_DoublePoten_AllYears.DataSrc.unique())
# print("_____________________________________________")
###
### NassIn, double potential, CorrectYear
###
NassIn_DoublePoten_CorrectYear = \
curr_double_poten[curr_double_poten["LstSrvD"].str.contains("2018", na=False)].copy()
NassIn_DoublePoten_CorrectYear_Acr = np.sum(NassIn_DoublePoten_CorrectYear['ExctAcr'])
# print ("3) is NASS in? should be ...")
# print (NassIn_DoublePoten_CorrectYear.DataSrc.unique())
# print("_____________________________________________")
del(NassIn_DoublePoten_CorrectYear)
###
### NassOut, double potential, CorrectYear
###
NassOut_DoublePoten = curr_double_poten[curr_double_poten.DataSrc != 'nass'].copy()
NassOut_DoublePoten_CorrectYear = \
NassOut_DoublePoten[NassOut_DoublePoten["LstSrvD"].str.contains("2018", na=False)].copy()
# print ("4) is NASS in? should NOT be ...")
# print (NassOut_DoublePoten_CorrectYear.DataSrc.unique())
# print("_____________________________________________")
NassOut_DoublePoten_CorrectYear_Acr = np.sum(NassOut_DoublePoten_CorrectYear['ExctAcr'])
del(NassOut_DoublePoten, NassOut_DoublePoten_CorrectYear)
###############################################################
#####
##### assemble the row and put it in output dataframe
#####
###############################################################
row = [NassIn_AllFields_AllYears_Acr, NassOut_AllFields_AllYears_Acr,
NassIn_AllFields_CorrectYear_Acr, NassOut_AllFields_CorrectYear_Acr,
NassIn_DoublePoten_AllYears_Acr, NassOut_DoublePoten_AllYears_Acr,
NassIn_DoublePoten_CorrectYear_Acr, NassOut_DoublePoten_CorrectYear_Acr]
output_df.iloc[num, 1: ] = row
# -
write_path = "/Users/hn/Desktop/"
filename = write_path + "Grant_2018_irrigated_acreages_DoublePeaks.csv"
output_df.to_csv(filename, index = False)
| remote_sensing/python/Local_Jupyter_NoteBooks/confusion_style_tables/zz_Pre_Eastern_WA_Era/Grant_2018_irrigated_acreages_DoublePeaks.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Demo - Fairness Analysis of FICO
#
# Adapted version of:
#
# 1. Fairness and machine learning book - [Chapter 2](https://fairmlbook.org/demographic.html) and [code repository](https://github.com/fairmlbook/fairmlbook.github.io).
#
# 2. <NAME>., <NAME>., & <NAME>. (2016). [Equality of opportunity in supervised learning](https://arxiv.org/abs/1610.02413). In Advances in neural information processing systems (pp. 3315-3323).
#
# 3. [Attacking discrimination with smarter machine learning](https://research.google.com/bigpicture/attacking-discrimination-in-ml/) by Google
# From [Wikipeida](https://en.wikipedia.org/wiki/Credit_score_in_the_United_States):
# > Credit score in the United States is a number representing the creditworthiness of a person, the likelihood that person will pay his or her debts.
# Lenders, such as banks and credit card companies, use credit scores to evaluate the potential risk posed by lending money to consumers. Lenders allege that widespread use of credit scores has made credit more widely available and less expensive for many consumers
#
#
# The analysis is based on data from [Report to the Congress on Credit Scoring and Its Effects on the Availability and Affordability of Credit](https://federalreserve.gov/boarddocs/rptcongress/creditscore/) by the Federal Reserve. The data set provides aggregate statistics from 2003 about a credit score, demographic information (race or ethnicity, gender, marital status), and outcomes (to be defined shortly).
#
#
# In the USA there are three majour creding agencies, which are for-profit organizations. They offer risk score based on the data they collected. Wre are going to look into **FICO** score of TransUnion (called TransRisk). The TransRisk score is in turn based on a proprietary model created by FICO, hence often referred to as FICO scores.
#
#
# 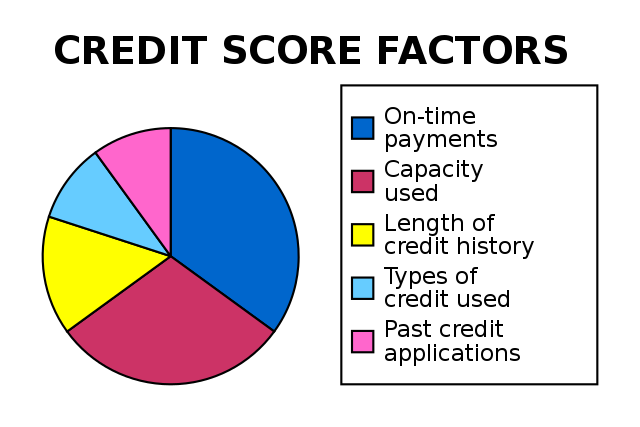
# <small>Source: Wikipedia</small>
#
#
# From Fairness and Machine Learning - Limitations and Opportunities:
# > Regulation of credit agencies in the United States started with the Fair Credit Reporting Act, first passed in 1970, that aims to promote the accuracy, fairness, and privacy of consumer of information collected by the reporting agencies. The Equal Credit Opportunity Act, a United States law enacted in 1974, makes it unlawful for any creditor to discriminate against any applicant the basis of race, color, religion, national origin, sex, marital status, or age.
#
# ### In our analysis we'll use on the joint statistics of score, race, and outcome.
# +
import pandas as pd
import matplotlib.pylab as plt
from responsibly.fairness.metrics import plot_roc_curves
from responsibly.fairness.interventions.threshold import (find_thresholds,
plot_fpt_tpr,
plot_roc_curves_thresholds,
plot_costs,
plot_thresholds)
# -
# ### FICO Dataset
#
# FICO dataset can be loaded directly from `responsibly`. The dataset, in this case, is *aggregate*, i.e., there is no outcome and prediction information per individual, but summarized statistics for each FICO score and race/race/ethnicity group.
# +
from responsibly.dataset import build_FICO_dataset
FICO = build_FICO_dataset()
# -
# `FICO` is a dictionary that holds variaty of data:
FICO.keys()
help(build_FICO_dataset)
# ### Counts by Race or Ethnicity
sum(FICO['totals'].values())
pd.Series(FICO['totals']).plot(kind='barh');
# ### Score Distribution
#
# The score used in the study is based on the TransUnion TransRisk score. TransUnion is a US credit-reporting agency. The TransRisk score is in turn based on a proprietary model created by FICO, hence often referred to as FICO scores. The Federal Reserve renormalized the scores for the study to vary from 0 to 100, with 0 being least creditworthy.
#
# The information on race was provided by the Social Security Administration, thus relying on self-reported values.
#
# The cumulative distribution of these credit scores strongly depends on the group as the next figure reveals.
FICO['cdf'].head()
FICO['cdf'].tail()
# +
f, ax = plt.subplots(1, figsize=(7, 5))
FICO['cdf'].plot(ax=ax)
plt.title('CDF by Group')
plt.ylabel('Cumulative Probability');
# -
# ### Outcome Variable
#
# **Performance variable** that measures a serious delinquency in at least one credit line of a certain time period:
#
# > "(the) measure is based on the performance of new or existing accounts and measures whether individuals have been late 90 days or more on one or more of their accounts or had a public record item or a new collection agency account during the performance period." - *from the Federal Reserve report*
#
# The `FICO['performance']` holds the percentage of non-defaulters for every score value (rows) and race/ethnicity group (columns):
FICO['performance'].head()
FICO['performance'].tail()
# ### Separation Fairness Criterion
# By the separation criterion of a binary classifier, the *FPR* and *TPR* should be equal across the groups.
plot_roc_curves(FICO['rocs'], FICO['aucs'],
figsize=(7, 5));
# The meaning of true positive rate is the rate of predicted positive performance given positive performance. Similarly, false positive rate is the rate of predicted negative performance given a positive performance.
# +
plot_roc_curves(FICO['rocs'], FICO['aucs'],
figsize=(7, 5));
plt.xlim(0, 0.3)
plt.ylim(0.4, 1);
# -
# ### Thresholds vs. FPR and TPR
#
# The ROC is paramaritazied over the thershold, so the same threshold might be related to different (FPR, TPR) pairs for each group. We can observe it by plotting the FPR and the TPR as a function of the threshold by the groups.
plot_fpt_tpr(FICO['rocs'], figsize=(15, 7),
title_fontsize=15, text_fontsize=15);
# Therefore, a naive choice of a single threshold will cause to a violation of the separation fairness criterion, as there will be different in FPR and TPR between the groups.
# ### Comparison of Different Criteria
#
# * Single threshold (Group Unaware)
# * Minimum Cost
# * Independence (Demographic Parity)
# * FNR (Equality of opportunity)
# * Separation (Equalized odds)
#
# #### Cost: $FP = - 5 \cdot TP$
COST_MATRIX = [[0, -5/6],
[0, 1/6]]
thresholds_data = find_thresholds(FICO['rocs'],
FICO['proportions'],
FICO['base_rate'],
FICO['base_rates'],
COST_MATRIX)
plot_roc_curves_thresholds(FICO['rocs'], thresholds_data,
figsize=(7, 7),
title_fontsize=20, text_fontsize=15);
# +
plot_roc_curves_thresholds(FICO['rocs'], thresholds_data,
figsize=(7, 7),
title_fontsize=20, text_fontsize=15)
plt.xlim(0, 0.3)
plt.ylim(0.4, 1);
# -
# ### Thresholds by Strategy and Group
plot_thresholds(thresholds_data,
xlim=(0, 100), figsize=(7, 7),
title_fontsize=20, text_fontsize=15);
# ### Cost by Threshold Strategy
plot_costs(thresholds_data);
# ### Sufficiency Fairness Criterion - Calibration
# +
f, axes = plt.subplots(nrows=1, ncols=2, figsize=(20, 10))
FICO['performance'].plot(ax=axes[0])
axes[0].set_ylabel('Non-default rate')
for group in FICO['cdf'].columns:
axes[1].plot(FICO['cdf'][group], FICO['performance'][group],
label=group)
axes[1].set_ylabel('Non-default rate')
axes[1].set_xlabel('Score')
axes[1].legend();
# -
# Due to the differences in score distribution by group, it could nonetheless be the case that thresholding the score leads to a classifier with different positive predictive values in each group.
| docs/notebooks/demo-fico-analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a href="https://colab.research.google.com/github/Tessellate-Imaging/Monk_Object_Detection/blob/master/example_notebooks/2_pytorch_finetune/Convert%20Pascal%20VOC%20Annotations%20to%20Desired%20Format.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# # Installation
#
# - Run these commands
#
# - git clone https://github.com/Tessellate-Imaging/Monk_Object_Detection.git
#
# - cd Monk_Object_Detection/2_pytorch_finetune/installation
#
# - Select the right requirements file and run
#
# - cat requirements.txt | xargs -n 1 -L 1 pip install
# ! git clone https://github.com/Tessellate-Imaging/Monk_Object_Detection.git
# +
# For colab use the command below
# ! cd Monk_Object_Detection/2_pytorch_finetune/installation && cat requirements_colab.txt | xargs -n 1 -L 1 pip install
# For Local systems and cloud select the right CUDA version
# ! cd Monk_Object_Detection/2_pytorch_finetune/installation && cat requirements.txt | xargs -n 1 -L 1 pip install
# -
# ## Dataset Directory Structure - Required
#
# Parent_Directory (root)
# |
# |-----------Images (img_dir)
# | |
# | |------------------img1.jpg
# | |------------------img2.jpg
# | |------------------.........(and so on)
# |
# |
# |-----------train_labels.csv (anno_file)
#
#
# ## Annotation file format
#
# | Id | Labels |
# | img1.jpg | x1 y1 x2 y2 label1 x1 y1 x2 y2 label2 |
#
# - Labels: xmin ymin xmax ymax label
# - xmin, ymin - top left corner of bounding box
# - xmax, ymax - bottom right corner of bounding box
# # Sample Dataset Credits
#
# - credits: https://github.com/wujixiu/helmet-detection
# ## To Convert pascal voc format to desired format we need to create an annotation csv file from the set of xml annotations
# +
import os
import sys
import numpy as np
import pandas as pd
import xmltodict
import json
from tqdm.notebook import tqdm
from pycocotools.coco import COCO
# -
root_dir = "Monk_Object_Detection/example_notebooks/sample_dataset/GDUT-HWD/";
img_dir = "JPEGImages/";
anno_dir = "Annotations/";
files = os.listdir(root_dir + anno_dir);
combined = [];
for i in tqdm(range(len(files))):
annoFile = root_dir + "/" + anno_dir + "/" + files[i];
f = open(annoFile, 'r');
my_xml = f.read();
anno = dict(dict(xmltodict.parse(my_xml))["annotation"])
fname = anno["filename"];
label_str = "";
if(type(anno["object"]) == list):
for j in range(len(anno["object"])):
obj = dict(anno["object"][j]);
label = anno["object"][j]["name"];
bbox = dict(anno["object"][j]["bndbox"])
x1 = bbox["xmin"];
y1 = bbox["ymin"];
x2 = bbox["xmax"];
y2 = bbox["ymax"];
if(j == len(anno["object"])-1):
label_str += x1 + " " + y1 + " " + x2 + " " + y2 + " " + label;
else:
label_str += x1 + " " + y1 + " " + x2 + " " + y2 + " " + label + " ";
else:
obj = dict(anno["object"]);
label = anno["object"]["name"];
bbox = dict(anno["object"]["bndbox"])
x1 = bbox["xmin"];
y1 = bbox["ymin"];
x2 = bbox["xmax"];
y2 = bbox["ymax"];
label_str += x1 + " " + y1 + " " + x2 + " " + y2 + " " + label;
combined.append([fname, label_str])
combined[:10]
df = pd.DataFrame(combined, columns = ['ID', 'Label']);
df.to_csv(root_dir + "/train_labels.csv", index=False);
# # Author - Tessellate Imaging - https://www.tessellateimaging.com/
#
# # Monk Library - https://github.com/Tessellate-Imaging/monk_v1
#
# Monk is an opensource low-code tool for computer vision and deep learning
#
#
# ## Monk features
# - low-code
# - unified wrapper over major deep learning framework - keras, pytorch, gluoncv
# - syntax invariant wrapper
#
# ## Enables
#
# - to create, manage and version control deep learning experiments
# - to compare experiments across training metrics
# - to quickly find best hyper-parameters
#
# ## At present it only supports transfer learning, but we are working each day to incorporate
#
# - GUI based custom model creation
# - various object detection and segmentation algorithms
# - deployment pipelines to cloud and local platforms
# - acceleration libraries such as TensorRT
# - preprocessing and post processing libraries
#
#
# ## To contribute to Monk AI or Monk Object Detection repository raise an issue in the git-repo or dm us on linkedin
#
# - Abhishek - https://www.linkedin.com/in/abhishek-kumar-annamraju/
# - Akash - https://www.linkedin.com/in/akashdeepsingh01/
#
#
| example_notebooks/2_pytorch_finetune/Convert Pascal VOC Annotations to Desired Format.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # ASTRO 533 - Mid Project 1
#
# **Created:** Sep. 2020
# **Last Edit:** Sep. 2020
#
# **Author:** <NAME>
# **Email:** <EMAIL>
# ## Load packages and read data
# +
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import matplotlib.colors as mcolors
from astropy.table import Table as tb
from astropy.coordinates import SkyCoord # High-level coordinates
from astropy.coordinates import ICRS, Galactic, FK4, FK5 # Low-level frames
from astropy.coordinates import Angle, Latitude, Longitude # Anglesfrom astropy.coordinates
import astropy.units as u
import copy
plt.style.use('bill')
my_YlGnBu = copy.copy(mpl.cm.get_cmap('YlGnBu')) # copy the default cmap
my_YlGnBu.set_bad('w')
my_magma = copy.copy(mpl.cm.get_cmap('magma'))
my_magma.set_bad('k')
gaiarv_cat = tb.read('../glt13.fits', format='fits')
gaiarv_cat_m45 = tb.read('./glt21_m45.fits', format='fits')
# gaiarv_cat_m22 = tb.read('./glt19_m22.fits', format='fits')
# -
# ## Pre-parameters
size_min = 0 * u.pc # in pc
size_max = 100 * u.pc # in pc
# ## Data processing
#
# *There will be several useless warnings.* ***Ignore them!***
# +
m45ra = 15 * (3 + (47/60) + (24/3600)) # RA = 3h 47m 24s
m45dec = 24 + (7/60) + (0/3600) # Dec = 24deg 7min 0sec
gaiarv_cat['d'] = 1000*u.pc*u.mas / (gaiarv_cat['parallax']) # distance in pc
gaiarv_cat['absmag'] = gaiarv_cat['phot_g_mean_mag'] - 5*np.log10(gaiarv_cat['d']/10) # absolute magnitude
gaiarv_cat_m45['d'] = 1000*u.pc*u.mas / (gaiarv_cat_m45['parallax']) # distance in pc
gaiarv_cat_m45['absmag'] = gaiarv_cat_m45['phot_g_mean_mag'] - 5*np.log10(gaiarv_cat_m45['d']/10) # absolute magnitude
# indices of neighborhood stars
ind_nb_pre, = np.where((gaiarv_cat['d'] < size_max) & (gaiarv_cat['d'] > size_min)) # only for plotting
ind_nb, = np.where((gaiarv_cat['d'] < size_max) & (gaiarv_cat['d'] > size_min) &
(gaiarv_cat['absmag'] < 4*gaiarv_cat['bp_rp']+2) &
(((gaiarv_cat['absmag'] > 15*gaiarv_cat['bp_rp']-10.5) & (gaiarv_cat['bp_rp'] < 1)) |
((gaiarv_cat['absmag'] > 2.25*gaiarv_cat['bp_rp']+2.25) & (gaiarv_cat['bp_rp'] > 1))))
print('# of pre-filter neighborhood stars:', len(ind_nb_pre))
print('# of neighborhood stars:', len(ind_nb))
# indices of m45 stars
ind_m45, = np.where((abs(gaiarv_cat_m45['ra']-m45ra) < 3) & (abs(gaiarv_cat_m45['dec']-m45dec) < 3) &
(abs(gaiarv_cat_m45['pmra']-20) < 5) & (abs(gaiarv_cat_m45['pmdec']+45) < 5) &
(abs(gaiarv_cat_m45['parallax']-7.3) < 0.7))
print('# of m45 stars:', len(ind_m45))
# -
# ## CMD
# +
# plot parameters
x_min, x_max = -0.2, 3
y_min, y_max = 0, 12
bins = 100
bins_m45 = 50
# plot
# fig, ax = plt.subplots(figsize=(6,6))
fig, [ax1,ax2] = plt.subplots(1, 2, figsize=(12,6), sharey=True, sharex=True)
fig.subplots_adjust(wspace=0)
ax1.hist2d(gaiarv_cat['bp_rp'][ind_nb_pre], gaiarv_cat['absmag'][ind_nb_pre], range=[[x_min, x_max], [y_min, y_max]],
bins = bins, norm=mcolors.LogNorm(), cmap=my_YlGnBu)
ax1.plot([-0.2,2.5], [1.2,12], c='gray', ls='--') # y < 4x + 2
ax1.plot([0.7,1,3], [0,4.5,9], c='gray', ls='--') # y > 15x - 10.5 (x<1) 2.25x + 2.25 (x>1)
ax1.fill_between([-0.2,2.5], [1.2,12], [12,12], facecolor='gray', alpha=0.1)
ax1.fill_between([0.7,1,3], [0,4.5,9], [0,0,0], facecolor='gray', alpha=0.1)
ax1.set_xlabel(r'$\mathrm{BP-RP}$')
ax1.set_ylabel(r'$\mathrm{G}$')
ax1.set_xlim(x_min, x_max)
ax1.set_ylim(y_max, y_min)
ax1.set_xticks([0, 1, 2, 3])
ax1.set_xticklabels([r'$0$', r'$1$', r'$2$', r'$3$'])
ax1.set_yticks([0, 2, 4, 6, 8, 10, 12])
ax1.set_yticklabels([r'$0$', r'$2$', r'$4$', r'$6$', r'$8$', r'$10$', r'$12$'])
ax1.text(0.96, 0.96, r'$r<%d\ \mathrm{pc}$' % size_max.value, ha='right', va='top', transform=ax1.transAxes, fontsize=18)
ax2.hist2d(gaiarv_cat_m45['bp_rp'][ind_m45], gaiarv_cat_m45['absmag'][ind_m45], range=[[x_min, x_max], [y_min, y_max]],
bins = bins_m45, norm=mcolors.LogNorm(), cmap=my_YlGnBu)
ax2.set_xlabel(r'$\mathrm{BP-RP}$')
ax2.set_ylim(y_max, y_min)
ax2.text(0.96, 0.96, r'$\mathrm{M45}$' % size_max.value, ha='right', va='top', transform=ax2.transAxes, fontsize=18)
plt.savefig('./figures/cmd.pdf')
plt.show()
# -
# ## PDMF
# +
# plot parameters
x_min, x_max = -2, 12
y_min, y_max = 0, 0.4
bins = 40
bin_edges = np.linspace(x_min, x_max, bins+1)
# detection limit
xs = (bin_edges[1:] + bin_edges[:-1])/2
d_lim = np.clip(10**(0.2*(13 - xs) + 1), 0, 100)
correct = (100 / d_lim)**3 # correction factor
# main plot
fig, ax = plt.subplots(figsize=(6,6))
hist_nb, bin_edges = np.histogram(gaiarv_cat['absmag'][ind_nb], bins=bin_edges)
hist_m45, bin_edges = np.histogram(gaiarv_cat_m45['absmag'][ind_m45], bins=bin_edges)
err_nb = np.sqrt(hist_nb) * correct
err_nb = err_nb * bins / (x_max-x_min) / np.sum(hist_nb)
hist_nb = hist_nb * correct
hist_nb = hist_nb * bins / (x_max-x_min) / np.sum(hist_nb)
err_m45 = np.sqrt(hist_m45)
err_m45 = err_m45 * bins / (x_max-x_min) / np.sum(hist_m45)
hist_m45 = hist_m45 * bins / (x_max-x_min) / np.sum(hist_m45)
ax.errorbar(xs, hist_nb, err_nb, fmt='none', alpha=0.5, c='k', elinewidth=1, label=None)
ax.errorbar(xs+0.05, hist_m45, err_m45, fmt='none', alpha=0.5, c='r', elinewidth=1, label=None)
ax.scatter(xs, hist_nb, marker='^', edgecolors='k', facecolor='k', alpha=0.5, s=20, label=r'$r<100\ \mathrm{pc}$')
ax.scatter(xs+0.05, hist_m45, marker='d', edgecolors='r', facecolor='r', alpha=0.5, s=20, label=r'$\mathrm{M45}$')
ax.plot([-1,4,4,-1,-1], [0,0,0.04,0.04,0], c='gray', ls='--')
ax.fill_between([-1,4], [0,0], [0.04,0.04], facecolor='gray', alpha=0.1)
ax.set_xlabel(r'$\mathrm{G}$')
ax.set_ylabel(r'$f\,(\mathrm{G})$')
ax.set_xlim(x_min, x_max)
ax.set_ylim(y_min, y_max)
ax.set_xticks([-2, 0, 2, 4, 6, 8, 10, 12])
ax.set_xticklabels([r'$-2$', r'$0$', r'$2$', r'$4$', r'$6$', r'$8$', r'$10$', r'$12$'])
ax.set_yticks([0, 0.1, 0.2, 0.3, 0.4])
ax.set_yticklabels([r'$0$', r'$0.1$', r'$0.2$', r'$0.3$', r'$0.4$'])
ax.legend(loc=1)
# top ticks
secax = ax.twiny()
secax.set_xlabel(r'$M\,/\,M_\odot$')
secax.set_xlim(x_min, x_max)
secax.set_xticks(-np.array([np.log10(6), np.log10(5), np.log10(4), np.log10(3), np.log10(2), np.log10(1),
np.log10(0.9), np.log10(0.8), np.log10(0.7), np.log10(0.6), np.log10(0.5), np.log10(0.4),
np.log10(0.3), np.log10(0.2)])*8.75+5.2) # G_sun = 5.2
secax.set_xticklabels(['', r'$5$', '', '', r'$2$', r'$1$', '', '', '', '', r'$0.5$', '', '', r'$0.2$'])
# small plot
ax2 = fig.add_axes([0.22,0.40,0.4,0.4])
ax2.errorbar(xs, hist_nb, err_nb, fmt='none', alpha=0.8, c='k', label=None)
ax2.errorbar(xs+0.05, hist_m45, err_m45, fmt='none', alpha=0.8, c='r', label=None)
ax2.scatter(xs, hist_nb, marker='^', edgecolors='k', facecolor='k', alpha=0.8, s=40, label=r'$r<100\ \mathrm{pc}$')
ax2.scatter(xs+0.05, hist_m45, marker='d', edgecolors='r', facecolor='r', alpha=0.8, s=40, label=r'$\mathrm{M45}$')
ax2.set_xlim(-1, 4)
ax2.set_ylim(0, 0.04)
ax2.set_xticks([-1, 0, 1, 2, 3, 4])
ax2.set_xticklabels([r'$-1$', r'$0$', r'$1$', r'$2$', r'$3$', r'$4$'])
ax2.set_yticks([0, 0.01, 0.02, 0.03, 0.04])
ax2.set_yticklabels([r'$0$', r'$0.01$', r'$0.02$', r'$0.03$', r'$0.04$'])
# top ticks
secax2 = ax2.twiny()
secax2.set_xlim(-2, 4)
secax2.set_xticks(-np.array([np.log10(6), np.log10(5), np.log10(4), np.log10(3), np.log10(2)])*8.75+5.2) # G_sun = 5.2
secax2.set_xticklabels([r'$6$', r'$5$', r'$4$', r'$3$', r'$2$'])
plt.savefig('./figures/pdmf.pdf')
plt.show()
# -
# ### Get MF from luminosity functions
# +
# plot parameters
x_min, x_max = np.log10(0.15), np.log10(5)
y_min, y_max = 0, 2
# main plot
fig, ax = plt.subplots(figsize=(6,6))
# get MF from luminosity functions
m_nb = 10**(-(xs[4:]-5.2)/8.75) # corresponding mass
m_edges_nb = 10**(-(10**(-(bin_edges[4:]-5.2)/8.75)-5.2)/8.75) # corresponding mass lags
fm_nb = hist_nb[4:] * 8.75 * 10**((xs[4:]-5.2)/8.75)/np.log(10) # pdmf
fm_err_nb = err_nb[4:] * 8.75 * 10**((xs[4:]-5.2)/8.75)/np.log(10)
fm_m45 = hist_m45[4:] * 8.75 * 10**((xs[4:]-5.2)/8.75)/np.log(10) # imf
fm_err_m45 = err_m45[4:] * 8.75 * 10**((xs[4:]-5.2)/8.75)/np.log(10)
eta = fm_nb / fm_m45
eta_err = eta * np.sqrt((fm_err_nb/fm_nb)**2 + (fm_err_m45/fm_m45)**2)
ax.errorbar(np.log10(m_nb), eta, eta_err, fmt='none', alpha=0.8, c='m', elinewidth=1, label=None)
ax.scatter(np.log10(m_nb), eta, marker='o', edgecolors='m', facecolor='m', alpha=0.8, s=20, label=r'$r<100\ \mathrm{pc}$')
ax.axhline(1, ls='-.', c='gray')
ax.set_xlabel(r'$M\,/\,M_\odot$')
ax.set_ylabel(r'$\eta\,(M)$')
ax.set_xlim(x_min, x_max)
ax.set_ylim(y_min, y_max)
ax.set_xticks(np.array([np.log10(5), np.log10(4), np.log10(3), np.log10(2), np.log10(1),
np.log10(0.9), np.log10(0.8), np.log10(0.7), np.log10(0.6), np.log10(0.5), np.log10(0.4),
np.log10(0.3), np.log10(0.2)])) # G_sun = 5.2
ax.set_xticklabels([r'$5$', '', '', r'$2$', r'$1$', '', '', '', '', r'$0.5$', '', '', r'$0.2$'])
ax.set_yticks([0, 0.5, 1, 1.5, 2])
ax.set_yticklabels([r'$0$', r'$0.5$', r'$1$', r'$1.5$', r'$2$'])
# top ticks
secax = ax.twiny()
secax.set_xlabel(r'$\left.T\,(M)\,\right/\,T_\odot$')
secax.set_xlim(x_min, x_max)
secax.set_xticks(-(np.array([-1, 0,1,2]))/2.5) # G_sun = 5.2
secax.set_xticklabels([r'$0.1$', r'$1$', r'$10$', r'$100$'])
plt.show()
# -
# ### Get SFH from LFs
# +
newx = (m_nb[1:] + m_nb[:-1])/2
psi_list = np.zeros([10000,len(newx)])
for i in range(10000):
test_eta = np.random.normal(eta, eta_err)
d_eta = (test_eta[1:] - test_eta[:-1]) / (m_nb[1:] - m_nb[:-1])
psi_list[i] = -d_eta * newx**3.5
psi = np.mean(psi_list, axis=0)
psi_err = np.std(psi_list, axis=0)
# +
# plot parameters
x_min, x_max = np.log10(0.15), np.log10(5)
y_min, y_max = -70, 70
# main plot
fig, ax = plt.subplots(figsize=(6,6))
ax.errorbar(np.log10(newx), psi, psi_err, fmt='none', alpha=0.8, c='m', elinewidth=1, label=None)
ax.scatter(np.log10(newx), psi, marker='o', edgecolors='m', facecolor='m', alpha=0.8, s=20, label=r'$r<100\ \mathrm{pc}$')
ax.axhline(0, ls='-.', c='gray')
ax.scatter(np.log10(newx[8]), psi[8], marker='*', c='r', s=160)
print('red star mass: %f M_sun' % newx[8], '; time: %f Gyr' % (10*(newx[8])**(-2.5)))
ax.set_xlabel(r'$M\,/\,M_\odot$')
ax.set_ylabel(r'$\psi\,(M)$')
ax.set_xlim(x_min, x_max)
ax.set_ylim(y_min, y_max)
ax.set_xticks(np.array([np.log10(5), np.log10(4), np.log10(3), np.log10(2), np.log10(1),
np.log10(0.9), np.log10(0.8), np.log10(0.7), np.log10(0.6), np.log10(0.5), np.log10(0.4),
np.log10(0.3), np.log10(0.2)])) # G_sun = 5.2
ax.set_xticklabels([r'$5$', '', '', r'$2$', r'$1$', '', '', '', '', r'$0.5$', '', '', r'$0.2$'])
ax.set_yticks([0])
ax.set_yticklabels([r'$0$'])
# top ticks
secax = ax.twiny()
secax.set_xlabel(r'$\left.T\,(M)\,\right/\,T_\odot$')
secax.set_xlim(x_min, x_max)
secax.set_xticks(-np.array([-1, 0,1,2])/2.5) # G_sun = 5.2
secax.set_xticklabels([r'$0.1$', r'$1$', r'$10$', r'$100$'])
plt.savefig('./figures/sfh.pdf')
plt.show()
| project_mid1/project_mid1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/alirezash97/Machine-Learning-Course/blob/main/Final_Project/RANZCR_update.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="0CiPXKgnKjBy"
# # # # trainset
# # !wget 'https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/23870/1781260/compressed/train.zip?GoogleAccessId=<EMAIL>&Expires=1611834692&Signature=sba7KapwtOseu8Fj6Yoyj1tjmhrhBCXWKvdhdOhUWlIapQjsjY6Vw8uT6L7WpLutCsA%2B5y8NN59YMso50eX6tYCai72huJ1CQtpVtr9uZPR9tWuuOYFin8%2BslEGSXVgwooUHZnjUQJppXMxfS7hi70UJqdaiAiIpPv18ILPBG39TBBuOiYgCs%2Bu2yM28dJ5sh3eHjJOElA83FsX4bL8h3PimMScR4sTLaqETj%2Bm0haY%2FjS8fdAeKuSDifxD40RmJz99UsHHkmt%2FWaKWGX5YXlVfwr8rN6pH10CZQ%2FCpQvyRvlkB1CpMiqdl8UrLzukLzdrgrAAp0PiXKwnzrwU%2B7Jw%3D%3D&response-content-disposition=attachment%3B+filename%3Dtrain.zip'
# # # train_tfrecords
# # # # !wget 'https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/23870/1781260/compressed/train_tfrecords.zip?GoogleAccessId=<EMAIL>&Expires=1611835011&Signature=OBxBwfMr9c0gB7Qmiiv9lGIHSqsT2ocsodWh1H56xb%2FYMBjLkiPKxiDXMBPnvOnaUGhMmKTmlpK06O8721DFO1hCNrq9757gZrxaVpm4400ABhzZ86NgLyLfC7Zse6GUlByeDrdd2Dk6KwI%2BjHFPg4TFFov3DW13I2%2FKw9h22tNbssdkfTA7OZgll1EW9Ynh6g%2F2ULQrmTtjkfdLbObPyniLEA5vHLXnK0ySw%2FaNS%2BCICHgGf4ECYqmrdWvzm8uDBrhrDs%2BwEwyMVTa4ZqnI0AS8FoMHexQV5yxbfUmihUDArft4QXrSnyCakAjaPHbknW2gyBfkmE%2FP0AHXAMx7Cg%3D%3D&response-content-disposition=attachment%3B+filename%3Dtrain_tfrecords.zip'
# # # # testset
# # # # !wget 'https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/23870/1781260/compressed/test.zip?GoogleAccessId=<EMAIL>.<EMAIL>.<EMAIL>account.<EMAIL>&Expires=1611411985&Signature=SPlBo05ncQEsd8RLzLUOCGhZ49kM9hcob5WJ1vdJHtGtL6a663HEdtgbwO3mIuv7jGtZYQltdUDZv867XtyOGPuLThK1rKdebC3jRq5DPYnIQPK%2BJ0JX%2FLcTnGiuRgPsxevW0vfjlBsEmJzYHr%2BXsKU6TdOHMaAwyCSX1JVMtO32C3BrgPNujkQ7HiTJ2C7H5bK7mB1Gm0Li%2Bg2wV2IhFl6%2FqW0CvDf3v3eBp9yS8Xt4w18VV5hkebAlCtXts8VU%2BxIgGy8nwIoJSq2vsSUAqx%2BfsuZkOOLfL0YnQ9ziqinMQSuAv8TXcGUhmlz5NS%2Bmmu%2BeRsPbOW9YJw4nkixsow%3D%3D&response-content-disposition=attachment%3B+filename%3Dtest.zip'
# # # # test_tfrecords
# # # # !wget 'https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/23870/1781260/compressed/test_tfrecords.zip?GoogleAccessId=<EMAIL>&Expires=1611412043&Signature=qChPWagoJ3ee9%2FErHTtluS1ojjzeWYALCFY%2BZ9TlU22ED2wIe3p7N98k6RDN3re7EKft5m%2FkeHszfURu59kwAA52o7F8pkvSMXoiuRctQGue4zNza4rKmVLzyvcqbRe7KmEDqiKtT2%2BY2UPaXxc0Yk0fHqqFe%2FZSbP7mqtXFUZHoGv8vW%2BS7Y4DRMXtPNbh5tBX9vHeDNJQ2UhIDOJwzfJCNWsFX5LZBohl7%2BjtzkGee%2FWPCel%2FCbdcddj%2BAvq1CMsRx8vpxRxNnubFCxTY8daCkZm4%2Bi%2FTZl92alLXWeaWda2RcidU2X7oGLbgdYhv3fzSGbBgM18VbJTDO7L71qg%3D%3D&response-content-disposition=attachment%3B+filename%3Dtest_tfrecords.zip'
# # # # train.csv
# # !wget 'https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/23870/1781260/compressed/train.csv.zip?GoogleAccessId=<EMAIL>&Expires=1611835399&Signature=C%2BMv6EdZSpBiCXddrVIhCx0r%2Bkrol9LSH92UL0YXN11VmU8hiYzL%2BTJJMQE2KHJsk7t8s7yFKbZcNvafDmNxkqeD%2BizlVzV1MuP5mTX9jiN30Qam75Ypsmgy5S3hr6%2FxKHalbL5I0VQgeh1mo4Umygb5K91MYwd3sM7wekYTvYEGgDbFF%2BFWk7jCZDIbSuW51pB0q%2FvKhWoh75jZnIlGW48xu2Em1nI5V%2Bfasxh72ezVX5N44X6Ts%2F71sOwfngPwWLkY3cIV3ksBHcuGgWPIbn0usX1CnBOXWtcuu0qbuPqlF3Sh6j31NXbHdjfXj3HZVMT7RNGjCODY2%2Bsk%2FyW48w%3D%3D&response-content-disposition=attachment%3B+filename%3Dtrain.csv.zip'
# # # # submission sample
# # # # !wget "https://www.kaggle.com/c/ranzcr-clip-catheter-line-classification/data?select=sample_submission.csv"
# # # # train annotations
# # # # !wget 'https://storage.googleapis.com/kaggle-competitions-data/kaggle-v2/23870/1781260/compressed/train_annotations.csv.zip?GoogleAccessId=<EMAIL>-<EMAIL>&Expires=1611835498&Signature=DftuTDD7S8H3eTfh%2FLQA50BWtL%2BSATrjxebLTtDXUMN9g4XyzYqCLy6C1pzb9ilL9tS23LI5tUf69e%2B5DxhY2SZBwyQXGss8H69%2F8yewJee93tSn1Asl3z19ExCzGji0rOmritXmVNuLhzApyK6KOw86NfL5pGrXhuBo%2FYSwPDsLpdnYcLsh7m09xfgjtci%2FomGFy3j6hhZxIJyYOwol9wYRWlOFFEtxq0lQ1l8BSv6O6Z1Yl9WV%2FzIV%2FJwA1YUYtHpeAENoGutDAfOAS7ogALme3KOgDMsK4Zv3uJ9Ofkamhz1sMM8g61gdug%2BBpKZY0xmJrrsLQ4NfEL09zRUmaA%3D%3D&response-content-disposition=attachment%3B+filename%3Dtrain_annotations.csv.zip'
# + id="c3f4srgdOJ53"
# # !mkdir /content/trainset
# # !mkdir /content/trainset/data/
# # !mkdir /content/trainset/data/1/
# # !unzip '/content/train.zip' -d /content/trainset/data/1/
# # # # # !unzip '/content/train_tfrecords.zip' -d /content/trainset/tf_records/
# # # # # !unzip '/content/train_annotations.csv.zip' -d /content/trainset/
# # !unzip '/content/train.csv.zip' -d /content/trainset/
# + id="PvDL9xh2Qc6N"
# # !mkdir /content/testset
# # !unzip '/content/test.zip' -d /content/testset/data
# # !unzip '/content/test_tfrecords.zip' -d /content/testset/test_tfrecords/
# + id="c-LgJKyRBziW"
# # !rm /content/test.zip
# # !rm /content/test_tfrecords.zip
# # !rm /content/train.csv.zip
# # !rm /content/train.zip
# # !rm /content/train_annotations.csv.zip
# # !rm /content/train_tfrecords.zip
# # !rm -rf /content/trainset/data/1
# + id="tPQAQU7ncDsr"
import pandas as pd
import numpy as np
# sample_submission = pd.read_csv('/content/sample_submission.csv')
# sample_submission.head()
# + id="ikRE9MDNnQGK"
train_csv = pd.read_csv('/content/trainset/train.csv')
# + id="EgM8GN2e1izJ"
def sampling(trainset, sample_per_class):
index_list = []
validation_index_list = []
threshold = np.full((12, ), sample_per_class)
for index, row in train_csv.iterrows():
sample = row[1:-1].values
flag = True
for i in range(11):
if threshold[i] > 0 :
if sample[i] == 1:
flag = False
threshold[i] -= 1
index_list.append(index)
break
else:
pass
else:
pass
if flag:
if threshold[11] > 0:
threshold[11] -= 1
index_list.append(index)
else:
validation_index_list.append(index)
return index_list, validation_index_list
# + id="RLVFb6FLPtEH" colab={"base_uri": "https://localhost:8080/"} outputId="3ffff68d-6b31-4c09-f4e5-73aab6535dae"
len(train_csv)
# + colab={"base_uri": "https://localhost:8080/"} id="5UdhB_tN9XpN" outputId="e424a10f-163a-4a4e-c482-7ee0222a7786"
sampled_index, validation_index = sampling(train_csv, 1)
validation_names =train_csv.iloc[validation_index, :]
print(len(validation_names))
images_name =train_csv.iloc[sampled_index, :]
len(images_name)
# + id="tbEhf8YFZuho" outputId="fe8ca63c-5bbd-4d96-e0b0-900ffdca05ca" colab={"base_uri": "https://localhost:8080/"}
# !cd test
# + id="sLHK1Y7rwgby" outputId="28dde137-88c5-44ec-c5ec-7736c6837800" colab={"base_uri": "https://localhost:8080/", "height": 346}
from __future__ import print_function, division
import os
import torch
import pandas as pd
import torchvision
from skimage import io, transform
import numpy as np
import matplotlib.pyplot as plt
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
# + id="jUnroASw1rgR"
# we calculate dataset mean and standard derivation only once
# from tqdm import tqdm
# dataset = datasets.ImageFolder('/content/trainset/data', transform=transforms.Compose([transforms.Resize((512, 512)),
# transforms.ToTensor()]))
# loader = torch.utils.data.DataLoader(dataset,
# batch_size=10,
# num_workers=0,
# shuffle=False)
# var = 0.0
# mean = 0.0
# for i, data in tqdm(enumerate(loader)):
# images = data[0]
# batch_samples = images.size(0)
# images = images.view(batch_samples, images.size(1), -1)
# mean += images.mean(2).sum(0)
# var += ((images - mean.unsqueeze(1))**2).sum([0,2])
# std = torch.sqrt(var / (len(loader.dataset)*224*224))
# mean = mean / len(loader.dataset)
# print('dataset mean: ', mean)
# print('dataset std: ', std)
# + id="XaxSOFwmBeWS"
# mean = np.array([0.4823, 0.4823, 0.4823])
# std = np.array([19147.3164, 19147.3164, 19147.3164])
# these results are calculated using above cell
# mean = np.array([0.4823])
# std = np.array([19147.3164])
# mean = np.array([0.4823])
# std = np.array([0.5])
mean = np.array([0.5057, 0.5057, 0.5057])
std = np.array([0.1902, 0.1902, 0.1902])
# + id="oD6dPyTlt_lR" outputId="eddfea14-4220-4d19-d733-ecb7b16847bc" colab={"base_uri": "https://localhost:8080/", "height": 229}
import cv2
class RANZCRDataset(Dataset):
def __init__(self, csv_file='/content/trainset/train.csv', root_dir='/content/trainset/data/1', transform=None, images_name=None):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.Images_name = images_name
self.root_dir = root_dir
self.transform = transform
def __len__(self):
return len(self.Images_name)
def __getitem__(self, idx):
if torch.is_tensor(idx):
idx = idx.tolist()
img_name = os.path.join(self.root_dir,
self.Images_name.iloc[idx, 0])
image = cv2.imread(img_name + '.jpg')
image = np.array(image)
# print(image.shape)
# image = np.expand_dims(image, axis=0)
# print(image.shape)
labels = self.Images_name.iloc[idx, 1:-1].values
labels = labels.astype(np.int)
labels = torch.from_numpy(labels)
sample = {'image': np.array(image), 'label': labels }
if self.transform:
sample['image'] = self.transform(torch.from_numpy(np.array(sample['image'], dtype=float)))
return sample
# + id="RgSZrpo8RAxk"
import torch
from torchvision import datasets, transforms
import torch.nn.functional as F
transform = transforms.Compose([transforms.Resize((1024, 1024)),
transforms.CenterCrop(904),
transforms.Normalize(mean , std),
transforms.ToTensor()])
trainset = RANZCRDataset(csv_file='/content/trainset/train.csv',
root_dir='/content/trainset/data/1', transform=transform, images_name=images_name)
train_data_loader = torch.utils.data.DataLoader(trainset,
batch_size=4,
shuffle=True,
num_workers=0)
validation = RANZCRDataset(csv_file='/content/trainset/train.csv',
root_dir='/content/trainset/data/1', transform=transform, images_name=validation_names)
validation_data_loader = torch.utils.data.DataLoader(validation,
batch_size=4,
shuffle=True,
num_workers=0)
# + colab={"base_uri": "https://localhost:8080/"} id="8muYGHBGwZen" outputId="e057c6b4-0c53-4548-9b02-4fae2f3c4b26"
def imshow(img):
npimg = img.numpy()
npimg = ((npimg * std) + mean) # unnormalize
plt.imshow((np.transpose(npimg, (1, 2, 0)) * 255).astype(np.uint8))
plt.show()
# get some random training images
dataiter = iter(train_data_loader)
sample = dataiter.next()
print(sample['image'].shape)
imshow(torchvision.utils.make_grid(sample['image']))
print(sample['label'])
# + id="cbxvV1q6aAXb"
import torch.nn as nn
import torch.nn.functional as F
class VGG(nn.Module):
def __init__(
self,
features: nn.Module,
num_classes: int = 1000,
init_weights: bool = True
) -> None:
super(VGG, self).__init__()
self.features = features
self.avgpool = nn.AdaptiveAvgPool2d((7, 7))
self.classifier = nn.Sequential(
nn.Linear(512 * 7 * 7, 4096),
nn.ReLU(True),
nn.Dropout(),
nn.Linear(4096, 4096),
nn.ReLU(True),
nn.Dropout(),
nn.Linear(4096, num_classes),
)
if init_weights:
self._initialize_weights()
def forward(self, x: torch.Tensor) -> torch.Tensor:
x = self.features(x)
x = self.avgpool(x)
x = torch.flatten(x, 1)
x = self.classifier(x)
return x
def _initialize_weights(self) -> None:
for m in self.modules():
if isinstance(m, nn.Conv2d):
nn.init.kaiming_normal_(m.weight, mode='fan_out', nonlinearity='relu')
if m.bias is not None:
nn.init.constant_(m.bias, 0)
elif isinstance(m, nn.BatchNorm2d):
nn.init.constant_(m.weight, 1)
nn.init.constant_(m.bias, 0)
elif isinstance(m, nn.Linear):
nn.init.normal_(m.weight, 0, 0.01)
nn.init.constant_(m.bias, 0)
# + id="qw5xyfmQqsZe"
from typing import Union, List, Dict, Any, cast
def make_layers(cfg: List[Union[str, int]], batch_norm: bool = False) -> nn.Sequential:
layers: List[nn.Module] = []
in_channels = 3
for v in cfg:
if v == 'M':
layers += [nn.MaxPool2d(kernel_size=2, stride=2)]
else:
v = cast(int, v)
conv2d = nn.Conv2d(in_channels, v, kernel_size=3, padding=1)
if batch_norm:
layers += [conv2d, nn.BatchNorm2d(v), nn.ReLU(inplace=True)]
else:
layers += [conv2d, nn.ReLU(inplace=True)]
in_channels = v
return nn.Sequential(*layers)
# + id="2uS_Vuo0ra2u"
def _vgg(arch: str, cfg: str, batch_norm: bool, pretrained: bool, progress: bool, **kwargs: Any) -> VGG:
if pretrained:
kwargs['init_weights'] = False
model = VGG(make_layers(cfgs[cfg], batch_norm=batch_norm), **kwargs)
if pretrained:
state_dict = load_state_dict_from_url(model_urls[arch],
progress=progress)
model.load_state_dict(state_dict)
return model
# + id="Y-ERtSFPrkfc"
cfgs: Dict[str, List[Union[str, int]]] = {
'A': [64, 'M', 128, 'M', 256, 256, 'M', 512, 512, 'M', 512, 512, 'M'],
'B': [64, 64, 'M', 128, 128, 'M', 256, 256, 'M', 512, 512, 'M', 512, 512, 'M'],
'D': [64, 64, 'M', 128, 128, 'M', 256, 256, 256, 'M', 512, 512, 512, 'M', 512, 512, 512, 'M'],
'E': [64, 64, 'M', 128, 128, 'M', 256, 256, 256, 256, 'M', 512, 512, 512, 512, 'M', 512, 512, 512, 512, 'M', 1024, 1024,
1024, 1024, 1024, 1024, 'M', 512, 512, 512, 512, 'M'],
}
# + id="23qaqsycr53x"
def vgg19_bn(pretrained: bool = False, progress: bool = True, **kwargs: Any) -> VGG:
r"""VGG 19-layer model (configuration 'E') with batch normalization
`"Very Deep Convolutional Networks For Large-Scale Image Recognition" <https://arxiv.org/pdf/1409.1556.pdf>`._
Args:
pretrained (bool): If True, returns a model pre-trained on ImageNet
progress (bool): If True, displays a progress bar of the download to stderr
"""
return _vgg('vgg19_bn', 'E', True, pretrained, progress, **kwargs)
# + id="XxB6NCvBsRA8"
model_urls = {
'vgg11': 'https://download.pytorch.org/models/vgg11-bbd30ac9.pth',
'vgg13': 'https://download.pytorch.org/models/vgg13-c768596a.pth',
'vgg16': 'https://download.pytorch.org/models/vgg16-397923af.pth',
'vgg19': 'https://download.pytorch.org/models/vgg19-dcbb9e9d.pth',
'vgg11_bn': 'https://download.pytorch.org/models/vgg11_bn-6002323d.pth',
'vgg13_bn': 'https://download.pytorch.org/models/vgg13_bn-abd245e5.pth',
'vgg16_bn': 'https://download.pytorch.org/models/vgg16_bn-6c64b313.pth',
'vgg19_bn': 'https://download.pytorch.org/models/vgg19_bn-c79401a0.pth',
}
# + id="nUoulwIcsclv"
try:
from torch.hub import load_state_dict_from_url
except ImportError:
from torch.utils.model_zoo import load_url as load_state_dict_from_url
# + id="4Dpyq8uYr8-7"
# model = vgg19_bn(pretrained=False)
model = torchvision.models.resnet152(pretrained=False, progress=True)
# + colab={"base_uri": "https://localhost:8080/"} id="_oFre0HvsMRj" outputId="e9e70b18-58ac-440f-c169-5d5fa7195a83"
print(model)
# + id="KLymbGLgMpu_"
import torch.nn as nn
import torch.nn.functional as F
class Network(nn.Module):
def __init__(self):
super(Network, self).__init__()
self.model = model
self.conv1 = nn.Conv2d(3, 3, 5)
self.conv2 = nn.Conv2d(3, 3, 1)
self.pool2 = nn.MaxPool2d(2, 2)
# self.conv2 = nn.Conv2d(8, 16, 5)
# self.pool3 = nn.MaxPool2d(3, 3)
# self.conv3 = nn.Conv2d(16, 64, 6)
# self.conv4 = nn.Conv2d(64, 128, 6)
# self.fc1 = nn.Linear(128 * 54 * 54, 1024)
# self.fc2 = nn.Linear(1024, 512)
# self.fc3 = nn.Linear(512, 11)
self.sigmoid = nn.Sigmoid()
self.fc_final = nn.Linear(1000, 11)
def forward(self, x):
x = self.pool2(F.relu(self.conv1(x)))
x = self.pool2(F.relu(self.conv2(x)))
x = self.model(x)
x = self.sigmoid(self.fc_final(x))
# x = self.pool2(F.relu(self.conv2(x)))
# x = self.pool2(F.relu(self.conv3(x)))
# x = self.pool3(F.relu(self.conv4(x)))
# x = x.view(-1, 128 * 54 * 54)
# x = F.relu(self.fc1(x))
# x = F.relu(self.fc2(x))
# x = self.fc3(x)
# x = self.sigmoid(x)
return x
Network = Network()
# + id="6F8h5at-Ynoc"
import torch.optim as optim
loss_function = nn.BCELoss()
optimizer = optim.SGD(model.parameters(), lr = 0.002, momentum = 0.9, weight_decay = 1e-5)
# optimizer = optim.Adam(Network.parameters(), lr = 0.01)
# + id="DSY9uhoeWodl"
def roc_auc_compute_fn(y_preds, y_targets):
try:
from sklearn.metrics import roc_auc_score
except ImportError:
raise RuntimeError("This contrib module requires sklearn to be installed.")
y_true = y_targets.detach().numpy()
y_pred = y_preds.detach().numpy()
try:
return roc_auc_score(y_true, y_pred, average='micro')
except ValueError:
return None
# + colab={"base_uri": "https://localhost:8080/", "height": 370} id="WjwmUvBRYqR6" outputId="1a7c092f-8732-40eb-92e7-3c1f5554fdf7"
for epoch in range(40):
running_loss = 0.0
for i, data in enumerate(train_data_loader, 0):
print(data['image'].shape)
inputs = data['image'].float()
label = data['label'].float()
optimizer.zero_grad()
outputs = Network(inputs)
loss = loss_function(outputs, label)
loss.backward()
optimizer.step()
auc_score = roc_auc_compute_fn(outputs, label)
running_loss += loss.item()
if i % 1 == 0:
print('[%d, %5d] training loss: %.3f training auc score: %.3f' %( epoch + 1, i + 1, running_loss, auc_score))
dataiter = iter(validation_data_loader)
sample = dataiter.next()
validation_inputs = sample['image'].float()
validation_label = sample['label'].float()
validation_outputs = Network(validation_inputs)
validation_loss = loss_function(validation_outputs, validation_label)
validation_auc_score = roc_auc_compute_fn(validation_outputs, validation_label)
print('[%d, %5d] <validation 10 random sample> loss: %.3f , auc score: %.3f' %( epoch + 1, i + 1, validation_loss, validation_auc_score))
running_loss = 0.0
try:
# torch.save(Network.state_dict(), '/content/trainset/model.pth')
pass
except ValueError:
pass
print('Finished Training Network')
# + colab={"base_uri": "https://localhost:8080/", "height": 354} id="LQkE8mE4ssFm" outputId="3644343c-5644-40d5-bfd1-03887f3096d9"
# Quick evaluation
# get some random training images
dataiter = iter(train_data_loader)
sample = dataiter.next()
inputs = sample['image'].float()
label = sample['label'].float()
print(inputs.shape)
batch_predicted_values = Network(inputs)
imshow(torchvision.utils.make_grid(sample['image']))
print(sample['label'])
print(batch_predicted_values)
# + id="rnhhfYNZJbhc"
| Final_Project/RANZCR_update.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # <font color='navy'>Introducción a Algotrading</font>
# <font color='blue'>Ing. <NAME>, MSF</font><br>
# MF-013 Análisis de Inversión<br>
# Clase del 26 de octubre 2021<br>
#
# Maestría de Finanzas, Facultad de Economía<br>
# UANL<br>
# ## Trabajando con datos financieros
# > _"Claramente los datos le ganan a los algortimos. Sin datos exahustivos tiendes a obtener predicciones no-exahustivas."_ <NAME> (Gerente General la división Analytics Business de IBM).
# Tipos de información financiera (ejemplos).
# <table>
# <th></th>
# <th>Datos Estructurados</th>
# <th>Datos No Estructurados</th>
# <tr>
# <td>Datos históricos</td>
# <td>Precios de cierre</td>
# <td>Noticias financieras</td>
# </tr>
# <tr>
# <td>Datos en tiempo real</td>
# <td>Precios bid/ask de las criptos</td>
# <td>Un tweet de Ellon Musk</td>
#
# ## Tipos de archivos
# Hay muchos formatos de datos que provienen de fuentes externas. Durante el resto del curso trabajaremos con archivos CSV y JSON's.
# ### Archivos CSV
# Son archivos de texto simple, separados por comas. CSV es la abreviación de "Comma Separated Values".<br><br>
# En la mayoría de los archivos CSV, la primer fila representa los encabezados de las columnas. Todas las filas posteriores, representan entradas de datos. En otros casos, las primeras filas representan espcificaciones del archivo en cuestión.
#
# Por lo general es una descarga manual del usuario.
# ### Archivos JSON
# Son archivos que guarda estructura de datos en formato JSON (JavaScript Object Notation). Es un formato utilizado para transmitir datos entre una aplicación y un servidor.
# + [markdown] tags=[]
# ## Archivos CSV y Python
# -
# Python tiene su propia librería para leer archivos CSV. La librería se llama `csv`. Una limitante es que no puedes cargar directamente un archivo de internet como lo hicmos con la función de Numpy `loadtxt( )`. Para poder obtener datos de internet se ocuparía otra librería como `requests` o `urlib`, haciendo la obtención de datos de internet más complicada de lo que es.
#
# Por lo anterior solo utilizaremos pandas para leer archivos csv.
# + [markdown] tags=[]
# ## Importar precios de WALMEX
# -
# Pasos:
# 1. Importar datos de internet, guardarlos en un DataFrame de pandas y gurdarlo como "__*walmex*__"
# 1. Formato del DataFrame:
# * Index: Columna de fechas
# * Fecha más antigua: Index 0
# * Fecha más reciente: Index -1
# * Nombre y órden de las columnas: "Apertura", "Maximo", "Minimo", "Cierre"$^+$
# 1. Crear una columna del DataFrame con los Retornos logarítmicos de los precios de cierre diarios
# 1. Realizar las siguientes gráficas:
# * Precios de cierre
# * Retornos diarios
# * Histograma de los retornos
# <br><br>
#
# __**NOTAS:__<br>
# $+$ No te recomiendo utilizar acentos al momento de definir el nombre de variables, columnas, df, etc.
# ### Lista de emisoras
# <table>
# <tr>
# <th><center></center></th>
# <th><center></center></th>
# <th><center></center></th>
# <th><center></center></th>
# </tr>
#
# <tr>
# <td style="text-align:center;">ac</td>
# <td style="text-align:center;">alfaa</td>
# <td style="text-align:center;">alpeka</td>
# <td style="text-align:center;">alsea</td>
# </tr>
#
# <tr>
# <td style="text-align:center;">amxl</td>
# <td style="text-align:center;">asurb</td>
# <td style="text-align:center;">bimboa</td>
# <td style="text-align:center;">bolsaa</td>
# </tr>
#
# <tr>
# <td style="text-align:center;">cemexcpo</td>
# <td style="text-align:center;">elektra</td>
# <td style="text-align:center;">femsaubd</td>
# <td style="text-align:center;">gapb</td>
# </tr>
#
# <tr>
# <td style="text-align:center;">gcarsoa1</td>
# <td style="text-align:center;">gcc</td>
# <td style="text-align:center;">gmexicob</td>
# <td style="text-align:center;">grumab</td>
# </tr>
#
# <tr>
# <td style="text-align:center;">ienova</td>
# <td style="text-align:center;">kimbera</td>
# <td style="text-align:center;">kofubl</td>
# <td style="text-align:center;">labb</td>
# </tr>
#
# <tr>
# <td style="text-align:center;">livepolc1</td>
# <td style="text-align:center;">megacpo</td>
# <td style="text-align:center;">omab</td>
# <td style="text-align:center;">orbia</td>
# </tr>
#
# <tr>
# <td style="text-align:center;">penoles</td>
# <td style="text-align:center;">pinfra</td>
# <td style="text-align:center;">tlevisacpo</td>
# <td style="text-align:center;">walmex</td>
# </tr>
#
# </table>
# #### Obtener datos de más acciones
# De la liga cambiar "walmex" por la acción de interés, por ejemplo "ac":<br>
# http://bit.ly/oncedos-walmex ---> http://bit.ly/oncedos-ac
# ## Función `read_csv( )` de pandas
# `read_csv( )` nos permite controlar varios parámetros y termina siendo un DataFrame. La clase `DataFrame` tiene varios métodos que tienen muchos usos en el campo de las finanzas.
# https://pandas.pydata.org/docs/reference/api/pandas.read_csv.html
import pandas as pd
import numpy as np
# + [markdown] tags=[]
# ### 1. Importar datos de Walmex
# -
url = 'http://bit.ly/oncedos-walmex'
walmex = pd.read_csv(url)
walmex.head(10)
# + [markdown] tags=[]
# #### Importar datos sin las primeras 6 líneas
# -
walmex = pd.read_csv(url, skiprows=6)
walmex.head()
walmex.info()
# ### 2. Formato del DataFrame
# + [markdown] tags=[]
# #### 2.1 Index: Columna de fechas
# -
# Dos opciones para cambiar la columna "*__Date__*" al index:
# 1. Cambiar la columan "*__Date__*" a index de manera manual.
# 2. Importar los datos espcificando que la columna "_**Date**_" es el index.
# ##### Una vez ya importado el `DataFrame`
walmex.set_index('Date', inplace=True)
walmex.head()
# + [markdown] tags=[]
# ##### Importar datos especificando que la columan "Date" será el index
# -
walmex = pd.read_csv(url, skiprows=6, index_col=0)
walmex.head()
walmex.loc['04/10/2019']
walmex.iloc[0]
# + [markdown] tags=[]
# ### 2.2 / 2.3 Orden de fechas
# -
walmex.sort_index(axis=0, inplace=True)
walmex.head()
walmex.head(20)
walmex = pd.read_csv(url, skiprows=6, index_col=0, parse_dates=True, dayfirst=True)
walmex.info()
# #### Método `sort_index( )`
# #### Atributo `parse_dates`
walmex.sort_index(axis=0, inplace=True)
walmex.head()
walmex.tail()
# ### 2.4 Nombre y órden de las columnas: "Apertura", "Maximo", "Minimo", "Cierre"
# Cambios finales al df:
# 1. Borrar las columnas que no utilizaremos
# 1. Renombrar las columnas a español
# 1. Ordenar las columnas en el formato O-H-L-C
# + [markdown] tags=[]
# #### 2.4.1 Borrar columnas
# -
borrar_columnas = ['Change', '% Change','Change.1', '% Change.1','Change.2', '% Change.2','Change.3', '% Change.3']
walmex.drop(borrar_columnas, axis=1, inplace=True)
walmex.head()
# #### 2.4.2 Renombrar columnas
dicc_renombrar = {'PX_LAST':'Cierre', 'PX_OPEN':'Apertura', 'PX_HIGH':'Maximo', 'PX_LOW':'Minimo'}
walmex.rename(dicc_renombrar, axis=1, inplace=True)
walmex.head()
# + [markdown] tags=[]
# #### 2.4.3 Reordenar columnas
# -
orden_columnas = ['Apertura', 'Maximo', 'Minimo', 'Cierre']
walmex.reindex(columns=orden_columnas, inplace=True)
walmex.reindex(columns=orden_columnas)
walmex.head()
walmex = walmex.reindex(columns=orden_columnas)
walmex.head()
# ### Volver a hacer todo en una sola celda
del walmex
# +
columnas_a_importar = ['Date', 'PX_OPEN', 'PX_HIGH', 'PX_LOW', 'PX_LAST']
walmex = pd.read_csv(url, skiprows = 6, index_col = 0, parse_dates = True, dayfirst = True,
usecols = columnas_a_importar)
walmex.rename({'PX_LAST':'Cierre', 'PX_OPEN':'Apertura', 'PX_HIGH':'Maximo', 'PX_LOW':'Minimo'}, axis = 1,
inplace = True)
orden_columnas = ['Apertura', 'Maximo', 'Minimo', 'Cierre']
walmex = walmex.reindex(columns=orden_columnas)
walmex.sort_index(inplace = True)
# -
walmex.head()
walmex.tail()
# ## Crear funciones
# Las funciones es un bloque de código diseñado para hacer un trabajo específico.
# Las funciones pueden:
# * Recibir valores
# * Tener valores predeterminados *por defalut* para el caso en que no se definan valores
# * NO RECIBIR VALORES!
# * Regresar un resultado
# * No regresar nada
a = 2
b = 3
suma1 = a+b
print(suma1)
i = 8
j = 10
suma2 = i+j
print(suma2)
y = 10
z = 20
suma_n = y + z
print(suma_n)
def funcion_sumar(valor_1, valor_2):
suma = valor_1 + valor_2
print(suma)
return suma
suma_n = funcion_sumar(10, 20)
suma_n
# +
suma1 = funcion_sumar(100,200)
suma2 = funcion_sumar(10,40)
suma3 = funcion_sumar(3,3)
# -
suma2
suma3
# + [markdown] tags=[]
# ### Función para importar datos archivos de Bloomberg
# -
def importar_bloomberg(accion):
url = f'http://bit.ly/oncedos-{accion}'
columnas_a_importar = ['Date', 'PX_OPEN', 'PX_HIGH', 'PX_LOW', 'PX_LAST']
df = pd.read_csv(url, skiprows = 6, index_col = 0, parse_dates = True, dayfirst = True,
usecols = columnas_a_importar)
df.rename({'PX_LAST':'Cierre', 'PX_OPEN':'Apertura', 'PX_HIGH':'Maximo', 'PX_LOW':'Minimo'}, axis = 1,
inplace = True)
orden_columnas = ['Apertura', 'Maximo', 'Minimo', 'Cierre']
df = df.reindex(columns=orden_columnas)
df.sort_index(inplace = True)
return df
ac = importar_bloomberg('ac')
ac.tail()
alfaa = importar_bloomberg('alfaa')
alfaa.head()
# #### Hacer librería HERRAMIENTAS FINANCIERAS
# Esta sección lo hicimos en el archivo de python `herramientas_financieras.py`
# #### Probar librería HERRAMIENTAS FINANCIERAS en otra libreta
# Esta sección la hicimos en otra libreta de jupyter `20211102Clase8PruebasLibreria.ipynb`
# + [markdown] tags=[]
# ### 2.3 Crear columna de retornos diarios
# $$RetLogaritmico = ln(Precio_n) - ln(Precio_{n-1})$$<br>
# $$ =ln\frac{Precio_n}{Precio_{n-1}}$$
# -
walmex.head()
# +
import numpy as np
walmex['Ret'] = np.log(walmex['Cierre'] / walmex['Cierre'].shift(1))
# -
walmex.head()
walmex.tail()
walmex.dropna(inplace=True)
walmex.head()
# #### Retornos en la escala logarítmica
walmex['Ret'].sum()
# #### Retornos en la escala números reales
( walmex['Cierre'].iloc[-1] - walmex['Cierre'].iloc[0] ) / walmex['Cierre'].iloc[0]
np.exp(walmex['Ret'].sum()) - 1
# ### 2.4 Graficar
# #### 2.4.1 Precio de cierre
walmex['Cierre'].plot(figsize=(12,8), title='Precios de cierre WALMEX');
# #### 2.4.2 Retornos
walmex['Ret'].plot(figsize=(12,8), title='Retornos diarios WALMEX');
# #### 2.4.3 Histograma
walmex['Ret'].hist(figsize=(12,8), bins = 10);
walmex['Ret'].plot(kind='hist', figsize=(12,8), bins=18, title='Histograma retornos Walmex');
| Codigo/20211102Clase8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import keras
import matplotlib.pyplot as plt
import cv2
import skimage
from skimage.measure import compare_ssim as ssim
import numpy as np
from keras.models import Sequential
from keras.layers import Conv2D,Dense,MaxPooling2D
from keras.optimizers import Adam,SGD
import math
import os
# %matplotlib inline
# +
# define a PSNR function (Peak Signal to Noise Ratio)
def psnr(target_img,ref_img):
target_data = target_img.astype(float)
ref_data = ref_img.astype(float)
difference = ref_data - target_data
rsme = math.sqrt(np.mean(diff**2))
return 20 * math.log10(255./rmse)
def mse(target_img,ref_img):
err = np.sum((target_img.astype(float) - ref_img.astype(float))**2)
err /= (target_img.shape[0] * target_img[1])
return err
def compare_image(target_img,ref_img):
scores = []
socres.append(psnr(target_img,ref_img))
socres.append(mse(target_img,ref_img))
socres.append((target_img,ref_img,multichannel=True))
return scores
| .ipynb_checkpoints/SRCNN-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="beObUOFyuRjT"
# ##### Copyright 2021 The TF-Agents Authors.
# + cellView="form" id="nQnmcm0oI1Q-"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="s6D70EeAZe-Q"
# # Drivers
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/agents/tutorials/4_drivers_tutorial">
# <img src="https://www.tensorflow.org/images/tf_logo_32px.png" />
# View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/agents/blob/master/docs/tutorials/4_drivers_tutorial.ipynb">
# <img src="https://www.tensorflow.org/images/colab_logo_32px.png" />
# Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/agents/blob/master/docs/tutorials/4_drivers_tutorial.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />
# View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/agents/docs/tutorials/4_drivers_tutorial.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] id="8aPHF9kXFggA"
# ## Introduction
#
# A common pattern in reinforcement learning is to execute a policy in an environment for a specified number of steps or episodes. This happens, for example, during data collection, evaluation and generating a video of the agent.
#
# While this is relatively straightforward to write in python, it is much more complex to write and debug in TensorFlow because it involves `tf.while` loops, `tf.cond` and `tf.control_dependencies`. Therefore we abstract this notion of a run loop into a class called `driver`, and provide well tested implementations both in Python and TensorFlow.
#
# Additionally, the data encountered by the driver at each step is saved in a named tuple called Trajectory and broadcast to a set of observers such as replay buffers and metrics. This data includes the observation from the environment, the action recommended by the policy, the reward obtained, the type of the current and the next step, etc.
# + [markdown] id="t7PM1QfMZqkS"
# ## Setup
# + [markdown] id="0w-Ykwl1bn4v"
# If you haven't installed tf-agents or gym yet, run:
# + id="TnE2CgilrngG"
# !pip install tf-agents
# + id="whYNP894FSkA"
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import tensorflow as tf
from tf_agents.environments import suite_gym
from tf_agents.environments import tf_py_environment
from tf_agents.policies import random_py_policy
from tf_agents.policies import random_tf_policy
from tf_agents.metrics import py_metrics
from tf_agents.metrics import tf_metrics
from tf_agents.drivers import py_driver
from tf_agents.drivers import dynamic_episode_driver
# + [markdown] id="9V7DEcB8IeiQ"
# ## Python Drivers
#
# The `PyDriver` class takes a python environment, a python policy and a list of observers to update at each step. The main method is `run()`, which steps the environment using actions from the policy until at least one of the following termination criteria is met: The number of steps reaches `max_steps` or the number of episodes reaches `max_episodes`.
#
# The implementation is roughly as follows:
#
#
# ```python
# class PyDriver(object):
#
# def __init__(self, env, policy, observers, max_steps=1, max_episodes=1):
# self._env = env
# self._policy = policy
# self._observers = observers or []
# self._max_steps = max_steps or np.inf
# self._max_episodes = max_episodes or np.inf
#
# def run(self, time_step, policy_state=()):
# num_steps = 0
# num_episodes = 0
# while num_steps < self._max_steps and num_episodes < self._max_episodes:
#
# # Compute an action using the policy for the given time_step
# action_step = self._policy.action(time_step, policy_state)
#
# # Apply the action to the environment and get the next step
# next_time_step = self._env.step(action_step.action)
#
# # Package information into a trajectory
# traj = trajectory.Trajectory(
# time_step.step_type,
# time_step.observation,
# action_step.action,
# action_step.info,
# next_time_step.step_type,
# next_time_step.reward,
# next_time_step.discount)
#
# for observer in self._observers:
# observer(traj)
#
# # Update statistics to check termination
# num_episodes += np.sum(traj.is_last())
# num_steps += np.sum(~traj.is_boundary())
#
# time_step = next_time_step
# policy_state = action_step.state
#
# return time_step, policy_state
#
# ```
#
# Now, let us run through the example of running a random policy on the CartPole environment, saving the results to a replay buffer and computing some metrics.
# + id="Dj4_-77_5ExP"
env = suite_gym.load('CartPole-v0')
policy = random_py_policy.RandomPyPolicy(time_step_spec=env.time_step_spec(),
action_spec=env.action_spec())
replay_buffer = []
metric = py_metrics.AverageReturnMetric()
observers = [replay_buffer.append, metric]
driver = py_driver.PyDriver(
env, policy, observers, max_steps=20, max_episodes=1)
initial_time_step = env.reset()
final_time_step, _ = driver.run(initial_time_step)
print('Replay Buffer:')
for traj in replay_buffer:
print(traj)
print('Average Return: ', metric.result())
# + [markdown] id="X3Yrxg36Ik1x"
# ## TensorFlow Drivers
#
# We also have drivers in TensorFlow which are functionally similar to Python drivers, but use TF environments, TF policies, TF observers etc. We currently have 2 TensorFlow drivers: `DynamicStepDriver`, which terminates after a given number of (valid) environment steps and `DynamicEpisodeDriver`, which terminates after a given number of episodes. Let us look at an example of the DynamicEpisode in action.
#
# + id="WC4ba3ObSceA"
env = suite_gym.load('CartPole-v0')
tf_env = tf_py_environment.TFPyEnvironment(env)
tf_policy = random_tf_policy.RandomTFPolicy(action_spec=tf_env.action_spec(),
time_step_spec=tf_env.time_step_spec())
num_episodes = tf_metrics.NumberOfEpisodes()
env_steps = tf_metrics.EnvironmentSteps()
observers = [num_episodes, env_steps]
driver = dynamic_episode_driver.DynamicEpisodeDriver(
tf_env, tf_policy, observers, num_episodes=2)
# Initial driver.run will reset the environment and initialize the policy.
final_time_step, policy_state = driver.run()
print('final_time_step', final_time_step)
print('Number of Steps: ', env_steps.result().numpy())
print('Number of Episodes: ', num_episodes.result().numpy())
# + id="Sz5jhHnU0fX1"
# Continue running from previous state
final_time_step, _ = driver.run(final_time_step, policy_state)
print('final_time_step', final_time_step)
print('Number of Steps: ', env_steps.result().numpy())
print('Number of Episodes: ', num_episodes.result().numpy())
| site/en-snapshot/agents/tutorials/4_drivers_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Optimization with Transaction costs
#
# In this lesson, we’ll show you how to incorporate transaction costs into portfolio optimization. This will give your backtest a more realistic measure of your alpha’s performance. In addition, we’ll show you some additional ways to design your optimization with efficiency in mind. This is really helpful when backtesting, because having reasonably shorter runtimes allows you to test and iterate on your alphas more quickly.
import sys
# !{sys.executable} -m pip install -r requirements.txt
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import pickle
import gzip
import bz2
from statsmodels.formula.api import ols
from scipy.stats import gaussian_kde
import scipy
import scipy.sparse
import patsy
from statistics import median
import datetime
# ## Barra data
#
# We’ll be using factor data that is generated by Barra. This will be good practice because Barra data is used throughout the industry.
#
# Note that we've pre-processed the raw barra data files and stored the data into pickle files. The alternative would be to load the original data, and perform the parsing each time. Since parsing and pre-processing takes time, we recommend doing the pre-processing once and saving the pre-processed data for later use in your backtest.
#
# Choose the number of years to use for the backtest. The data is available for years 2003 to 2008 inclusive.
barra_dir = '../../data/project_8_barra/'
# !ls {barra_dir}
# +
data = {}
for year in [2003]:
fil = barra_dir + "pandas-frames." + str(year) + ".pickle"
data.update(pickle.load( open( fil, "rb" ) ))
covariance = {}
for year in [2003]:
fil = barra_dir + "covariance." + str(year) + ".pickle"
covariance.update(pickle.load( open(fil, "rb" ) ))
daily_return = {}
for year in [2003, 2004]:
fil = barra_dir + "price." + str(year) + ".pickle"
daily_return.update(pickle.load( open(fil, "rb" ) ))
# -
# Notice that the frames variale is a dictionary, where the keys are strings representing each business day.
# ## View the Barra data
#
# We'll take a look at the value stored for a single day (it's a data frame).
#
# As a general reminder of best practices, remember to check what unit of measure your data is in. In some cases, the unit of measure isn’t available in the documentation, so you’ll want to inspect the data to see what makes sense.
#
# For instance, there are volatility fields that are large enough that we can assume they are in percentage units, as opposed to decimal values. In other cases, when we look at daily volume, we may not have documentation about whether the units are in number of shares or in dollars. One way to find this out is to spot check a single stock on a single day, and cross-reference with another source, such as Bloomberg or Yahoo Finance.
# Remember to inspect the data before you use it, as it will help you derive more meaningful results in your portfolio optimization, and in your backtest.
#
# Remember to inspect the data before you use it, as it will help you derive more meaningful results in your portfolio optimization, and in your backtest.
#
# In the exercise, we'll re-scale the data before using it, and there will be comments to point out when we re-scale the data. So don't worry about adjusting anything here, just take a look to get familiar with the data.
data.keys()
data['20030102'].head()
data['20030102'].shape
# #### Factors
#
# Note that the data fields that start with the prefix U-S-F-A-S-T are factor exposures, one column for each factor. We will use some of these as alpha factors, and the rest as risk factors. The reason this makes sense is that, for the time periods in which we’re back-testing, some of these factors were able to produce better than average returns. Barra works with its clients (funds) and gathers information about alphas that worked in the past. These were calculated on historical data to produce the factor exposure data found in the Barra data.
# 
# ## Factors
#
# Here's a partial list of the barra factors in our dataset and their definitions. These are collected from documentation by Barra. There are style factors and industry factors. The industry factors will be used as risk factors. You can consider using the style factors as alpha factors. Any factors not used as alpha factors can be included in the risk factors category.
#
# #### Style factors
# * beta: Describes market risk that cannot be explained by the Country factor. The Beta factor is typically the most important style factor. We calculate Beta by time-series regression of stock excess returns against the market return.
# * 1 day reversal
# * dividend yield: Describes differences in stock returns attributable to stock's historical and predicted dividend-to-price ratios.
# * downside risk (maximum drawdown)
# * earnings quality: Describes stock return differences due to the accrual components of earnings.
# * earnings yield: Describes return differences based on a company’s earnings relative to its price. Earnings Yield is considered by many investors to be a strong value signal. The most important descriptor in this factor is the analyst-predicted 12-month earnings-to-price ratio.
# * growth: Differentiates stocks based on their prospects for sales or earnings growth. The most important descriptor in this factor is the analyst predicted long-term earnings growth. Other descriptors include sales and earnings growth over the previous five years.
# * leverage: Describes return differences between high and low-leverage stocks. The descriptors within this style factor include market leverage, book leverage, and debt-to-assets ratio.
# * liquidity: Describes return differences due to relative trading activity. The descriptors for this factor are based on the fraction of total shares outstanding that trade over a recent window.
# * long-term reversal: Describes common variation in returns related to a long-term (five years ex. recent thirteen months) stock price behavior.
# * management quality
# * Mid capitalization: Describes non-linearity in the payoff to the Size factor across the market-cap spectrum. This factor is based on a single raw descriptor: the cube of the Size exposure. However, because this raw descriptor is highly collinear with the Size factor, it is orthogonalized with respect to Size. This procedure does not affect the fit of the model, but does mitigate the confounding effects of collinearity, while preserving an intuitive meaning for the Size factor. As described by Menchero (2010), the Mid Capitalization factor roughly captures the risk of a “barbell portfolio” that is long mid-cap stocks and short small-cap and large-cap stocks.
# * Momentum – Differentiates stocks based on their performance over the trailing 12 months. When computing Momentum exposures, we exclude the most recent returns in order to avoid the effects of short-term reversal. The Momentum factor is often the second strongest factor in the model, although sometimes it may surpass Beta in importance.
# * Profitability – Combines profitability measures that characterize efficiency of a firm's operations and total activities.
# * Residual Volatility – Measures the idiosyncratic volatility anomaly. It has three descriptors: (a) the volatility of daily excess returns, (b) the volatility of daily residual returns, and (c) the cumulative range of the stock over the last 12 months. Since these descriptors tend to be highly collinear with the Beta factor, the Residual Volatility factor is orthogonalized with respect to the Beta and Size factors.
# * seasonality
# * sentiment
# * Size – Represents a strong source of equity return covariance, and captures return differences between large-cap and small-cap stocks. We measure Size by the log of market capitalization.
# * Short term reversal
# * Value
# * Prospect -- is a function of skewness and maximum drawdown.
# * Management Quality -- is a function of the following:
# * Asset Growth: Annual reported company assets are regressed against time over the past five fiscal years. The slope coefficient is then divided by the average annual assets to obtain the asset growth.
# * Issuance Growth Annual reported company number of shares outstanding regressed against time over the past five fiscal years. The slope coefficient is then divided by the average annual number of shares outstanding.
# * Capital Expenditure Growth: Annual reported company capital expenditures are regressed against time over the past five fiscal years. The slope coefficient is then divided by the average annual capital expenditures to obtain the capital expenditures growth.
# * Capital Expenditure: The most recent capital expenditures are scaled by the average of capital expenditures over the last five fiscal years.
#
#
#
# #### Industry Factors
# * aerospace and defense
# * airlines
# * aluminum and steel
# * apparel
# * Automotive
# * banks
# * beta (market)
# * beverage and tobacco
# * biotech & life science
# * building products
# * chemicals
# * construction & engineering
# * construction & machinery
# * construction materials
# * commercial equipment
# * computer & electronics
# * commercial services
# * industrial conglomerates
# * containers (forest, paper, & packaging)
# * distributors
# * diversified financials
# * electrical equipment
# * electrical utility
# * food & household products & personal
# * food & staples retailing
# * gas & multi-utilities
# * healthcare equipment and services
# * health services
# * home building
# * household durables
# * industry machinery
# * non-life insurance
# * leisure products
# * leisure services
# * life insurance
# * managed healthcare
# * multi-utilities
# * oil & gas conversion
# * oil & gas drilling
# * oil & gas equipment
# * oil and gas export
# * paper
# * pharmaceuticals
# * precious metals
# * personal products
# * real estate
# * restaurants
# * road & rail
# * semiconductors
# * semiconductors equipment
# * software
# * telecommunications
# * transportation
# * wireless
# * SPTY\* and SPLTY\* are various industries
data['20030102'].columns
# ## covariance of factors
#
# Let's look at the covariance of the factors.
covariance.keys()
# View the data for a single day. Notice that the factors are listed in two columns, followed by the covariance between them. We'll use this data later to create a factor covariance matrix.
covariance['20030102'].head()
# ## Daily returns
daily_return.keys()
daily_return['20030102'].head()
# ## Add date for returns
#
# We'll be dealing with two different dates; to help us keep track, let's add an additional column in the daily_return dataframes that stores the date of the returns.
tmp_date = '20030102'
tmp = daily_return[tmp_date]
tmp.head()
tmp_n_rows = tmp.shape[0]
pd.Series([tmp_date]*tmp_n_rows)
tmp['DlyReturnDate'] = pd.Series([tmp_date]*tmp_n_rows)
tmp.head()
# ## Quiz: add daily return date to each dataframe in daily_return dictionary
#
# Name the column `DlyReturnDate`.
# **Hint**: create a list containing copies of the date, then create a pandas series.
for DlyReturnDate, df in daily_return.items():
# TODO
n_rows = df.shape[0]
df['DlyReturnDate'] = pd.Series([DlyReturnDate]*n_rows)
# +
# check results
daily_return['20030102'].head()
# -
# ## Adjust dates to account for trade execution
#
# The data stored in `data` and `covariance` are used to choose the optimal portfolio, whereas the data in `daily_return` represents the the returns that the optimized portfolio would realize, but only after we've received the data, then chosen the optimal holdings, and allowed a day to trade into the optimal holdings. In other words, if we use the data from `data` and `covariance` that is collected at the end of Monday, we'll use portfolio optimization to choose the optimal holdings based on this data, perhaps after hours on Monday. Then on Tuesday, we'll have a day to execute trades to adjust the portfolio into the optimized positions. Then on Wednesday, we'll realize the returns using those optimal holdings.
# Example of what we want
data_date_l = sorted(data.keys())
return_date_l = sorted(daily_return.keys())
len(data_date_l)
len(return_date_l)
return_date_l_shifted = return_date_l[2:len(data) + 2]
len(return_date_l_shifted)
# data date
data_date_l[0]
# returns date
return_date_l_shifted[0]
tmp = data['20030102'].merge(daily_return['20030102'], on="Barrid")
tmp.head()
# ## Merge data and daily returns into single dataframe
#
# Use a loop to merge the `data` and `daily_return` tables on the `barrid` column.
# +
frames ={}
# TODO
dlyreturn_n_days_delay = 2
# TODO
date_shifts = zip(
sorted(data.keys()),
sorted(daily_return.keys())[dlyreturn_n_days_delay:len(data) + dlyreturn_n_days_delay])
# TODO
for data_date, price_date in date_shifts:
frames[price_date] = data[data_date].merge(daily_return[price_date], on='Barrid')
# -
# ## Let's work with a single day's data. Later, we'll put this into a loop
#
#
# Notice how the keys are now dates of the returns. So the earliest date in "frames" dictionary is two business days after the earliest date in "data" dictionary.
frames.keys()
df = frames['20030106']
df.head()
# ## Quiz
#
# Filter the stocks so that the estimation universe has stocks with at least 1 billion in market cap. As an aside, it doesn't make much of a difference whether we choose a ">" or ">=", since the threshold we choose is just meant to get a set of relatively liquid assets.
#
# **Hint**: use `.copy(deep=True)` to make an independent copy of the data.
# TODO
estu = df.loc[df.IssuerMarketCap >= 1e9].copy(deep=True)
estu.head()
# For all the columns in the dataframe, the ones with the prefix "USFAST" are factors. We'll use a helper function to get the list of factors.
def factors_from_names(n):
return(list(filter(lambda x: "USFASTD_" in x, n)))
all_factors = factors_from_names(list(df))
all_factors
# ## factors exposures and factor returns
#
# Recall that a factor's factor return times its factor exposure gives the part of a stock's return that is explained by that factor.
#
# The Barra data contains the factor exposure of each factor. We'll use regression to estimate the factor returns of each factor, on each day. The observations will be the cross section of stock factor exposures, as well as the stock returns that are realized two trading days later. Recall from an earlier lesson that this is a cross-sectional regression, because it's a cross section of stocks, for a single time period.
#
# $r_{i,t} = \sum_{j=1}^{k} (\beta_{i,j,t-2} \times f_{j,t})$
# where $i=1...N$ (N assets),
# and $j=1...k$ (k factors).
#
# In the regression, the factor exposure, $\beta_{i,j,t-2}$ is the independent variable, $r_{i,t}$ is the dependent variable, and the factor return $f_{j,t}$ is the coefficient that we'll estimate.
# ## Calculating factor returns
#
# We'll estimate the factor returns $f_{j,t}$ of our chosen alpha factors, using the daily returns of the stocks $r_{i,t}$, where $i=1...N$ and the factor exposure $\beta_{i,j,t-2}$ of each stock to each factor.
#
# Note that we'll use a universe of stocks where the companies have a market capitalization of at least 1 billion. The factor returns estimated would be slightly different depending on which stock universe is chosen, but choosing a market cap of 1 billion or more provides a reasonable estimate of what you'd expect to be tradable. The estimated factor returns would be fairly close to what you'd find if you used the Russell 3000 index as the stock universe.
# ## formula
#
# We'll use a helper function that creates a string that defines which are the independent and dependent variables for a model to use. This string is called a "formula." We'll use this in the regression, and later again when we work with matrices.
def get_formula(factors, Y):
L = ["0"]
L.extend(factors)
return Y + " ~ " + " + ".join(L)
form = get_formula(all_factors, "DlyReturn")
# So, the formula is saying `DlyReturn` is the dependent variable, whereas the `USFAST...` columns are the independent variables.
form
# ## Quiz
#
# Run an ordinary least squares regression
#
# [ols documentation](https://www.statsmodels.org/dev/example_formulas.html)
#
# Here's an example of the syntax.
# ```
# ols(formula='y ~ x1 + x2 + x3', data=dataframe)
# ```
#
# Note that you're free to choose other regression models, such as ridge, lasso, or elastic net. These may give you slightly different estimations of factor returns, but shouldn't be too different from each other.
# +
# TODO
model = ols(formula=form, data=estu)
# TODO
results = model.fit()
# -
# Since the factor data that we're using as the independent variables are the factor exposures, the coefficients estimated by the regression are the estimated factor returns.
results.params
# ## Quiz: winsorize daily returns before calculating factor returns
#
# We're going to use regression to estimate the factor returns of all the factors. To avoid using extreme values in the regression, we'll winsorize, or "clip" the returns. We can check the data distribution using a density plot.
#
# Note that [numpy.where](https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.where.html) has the form
# ```
# numpy.where(<condition>, <value if true>, <value if false>)
# ```
def wins(x,wins_lower,wins_upper):
#TODO
clipped_upper = np.where(x >= wins_upper, wins_upper, x)
clipped_both = np.where(clipped_upper <= wins_lower,wins_lower, clipped_upper)
return clipped_both
# A density plot will help us visually check the effect of winsorizing returns.
def density_plot(data):
density = gaussian_kde(data)
xs = np.linspace(np.min(data),np.max(data),200)
density.covariance_factor = lambda : .25
density._compute_covariance()
plt.plot(xs,density(xs))
plt.show()
# distribution without winsorizing
test = frames['20040102']
density_plot(test['DlyReturn'])
# distribution after winsorizing
test['DlyReturn_wins'] = wins(test['DlyReturn'], wins_lower=-0.1, wins_upper=0.1)
density_plot(test['DlyReturn_wins'])
# ## Quiz
#
# Put the factor returns estimation into a function, so that this can be re-used for each day's data.
def estimate_factor_returns(df, wins_lower=-.25, wins_upper=0.25):
## TODO: build estimation universe based on filters
estu = df.loc[df.IssuerMarketCap > 1e9].copy(deep=True)
## TODO: winsorize returns for fitting
estu['DlyReturn'] = wins(estu['DlyReturn'], wins_lower, wins_upper)
## get a list of all the factors
all_factors = factors_from_names(list(df))
## define a 'formula' for the regression
form = get_formula(all_factors, "DlyReturn")
## create the OLS model, passing in the formula and the estimation universe dataframe
model = ols(formula=form, data=estu)
## return the estimated coefficients
results = model.fit()
return(results.params)
# ## Choose alpha factors
#
# We'll choose the 1 day reversal, earnings yield, value, and sentiment factors as alpha factors. We'll calculate the factor returns of these alpha factors to see how they performed.
alpha_factors = ["USFASTD_1DREVRSL", "USFASTD_EARNYILD", "USFASTD_VALUE", "USFASTD_SENTMT"]
print(alpha_factors)
# ## Quiz: estimate factor returns of alpha factors
#
# Loop through each day, and estimate the factors returns of each factor, that date, in the `frames` dictionary. This may take a minute or more to run per year of data used.
facret = {}
for date in frames:
# TODO: store factor returns as key-value pairs in a dictionary
facret[date] = estimate_factor_returns(frames[date])
type(facret['20040102'])
facret['20040102'].head()
# ## put the factor returns into a dataframe
#
# The pandas series are stored inside a dictionary. We'll put the factor returns into a dataframe where the rows are the dates and the columns are the factor returns (one column for each factor).
#
# First, let's get a list of dates, as Timestamp objects. We'll use [pandas.to_datetime](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.to_datetime.html)
# example of how to convert the keys of the dataframe into Timestamp objects
pd.to_datetime('20040102', format='%Y%m%d')
# ## Quiz
#
# Store the timestamp objects in a list (can use a list comprehension, or for loop).
# TODO
dates_unsorted = [pd.to_datetime(date, format='%Y%m%d') for date in frames.keys()]
# sort the dates in ascending order
my_dates = sorted(dates_unsorted)
# We'll make an empty dataframe with the dates set as the row index.
facret_df = pd.DataFrame(index = my_dates)
facret_df.head()
# The rows are the dates. The columns will be the factor returns.
#
# To convert from Timestamp objects back into a string, we can use [Timestamp.strftime('%Y%m%d')](https://www.programiz.com/python-programming/datetime/strftime).
## example usage of Timestamp.strftime('%Y%m%d')
my_dates[0].strftime('%Y%m%d')
# ## Quiz
# For each date, and for each factor return, get the value from the dictionary and put it into the dataframe.
#
# We can use [pandas.DataFrame.at¶](https://pandas.pydata.org/pandas-docs/stable/reference/api/pandas.DataFrame.at.html),
#
# ```
# DataFrame.at[<index_value>,<column_name>] = <some_value>
# ```
# TODO: for each date (rows), and for each factor (columns),
# store factor return in the dataframe
for dt in my_dates:
for alp in alpha_factors:
facret_df.at[dt, alp] = facret[dt.strftime('%Y%m%d')][alp]
# ## Portfolio optimization for a single period
#
# When we get to the project, we'll want to define the portfolio optimization within a function. For now, let's walk through the steps we'll take in separate cells, so that we can see what's going on.
# The optimization will want to know about the prior trading day's portfolio holdings, also called holdings. The previous day's holdings will be used to estimate the size of the trades due to position changes, which in turn helps us estimate transaction costs. We'll start with an initial holding of zero for a single stock. The reason we'll use a single stock is that the estimation universe chosen on each day will include all stocks that have holdings on the previous day. So we want to keep this list small when we first start out, else we'll keep many stocks that may fall below the 1 billion market cap threshold, just because they were chosen in the initialization of the backtest.
#
# We'll want to choose a stock that is likely to satisfy the 1 billion market cap threshold on any day. So let's choose the stock with the largest market cap.
# we're going to set a single barra id to have a zero portfolio holding,
# so let's pick any barra id of the stock with the largest issuer market cap
estu.sort_values('IssuerMarketCap',ascending=False)[['Barrid','IssuerMarketCap']].head()
# ## Quiz: Intialize previous holdings dataframe
#
# Create a new dataframe and initialize it with a dictionary, where the key is "Barrid" followed by a value that is a pandas.Series containing the barra id of the largest market cap in the stock universe.
#
# Also set another key value pair to "x.opt.previous" and the value is set to a pandas.Series with the value 0.
# TODO
# create a dataframe of previous holdings,
# initializing a single stock (barra id) to zero portfolio holding
previous_holdings = pd.DataFrame(data = {"Barrid" : pd.Series( ["USA0001"]),
"x.opt.previous" : pd.Series(0)})
previous_holdings
# Get a single day's data to be used for the portfolio optimization.
dt = my_dates[0]
date = dt.strftime('%Y%m%d')
print(date)
df = frames[date]
df.head()
# Let's add the previous holdings column to the dataframe
## merge previous portfolio holdings
df = df.merge(previous_holdings, how = 'left', on = 'Barrid')
df.head()
# ## Clean missing and zero values.
#
# First replace missing values with zero.
# +
def na2z(x):
return(np.nan_to_num(x))
def names_numeric_columns(df):
return(df.select_dtypes(include=[np.number]).columns.tolist())
def clean_nas(df):
for x in names_numeric_columns(df):
df[x] = na2z(df[x])
return(df)
# -
df = clean_nas(df)
# ## Quiz: Clean specific risk
# Barra calculates specific risk for each asset. If the value in the data is zero, this may be due to missing data rather than the specific risk actually being zero. So we'll set zero values to the median, to make sure our model is more realistic.
# TODO: if SpecRisk is zero, set it to median
df.loc[df['SpecRisk'] == 0]['SpecRisk'] = median(df['SpecRisk'])
# ## universe
#
# We'll look at stocks that are 1 billion in market cap or greater. An important point here is that we'll need to account for stocks that are already in our portfolio, even if the market cap of the stock is no longer 1 billion on the current day.
# #### Quiz: think about what would happen if we had an existing position in a stock, then the market cap fell below the threshold and the stock was excluded from the stock universe. What would happen to the position on that stock?
# #### Answer
# The stock would not be included in the optimization, which means it would be given a zero position. So this effectively says to sell all holdings in the asset once it falls below the market cap threshold. That's not what we want to do.
# Modify the code to account for the prior day's positions.
## TODO: modify the given code to include the prior day's assets
universe = df.loc[(df['IssuerMarketCap'] >= 1e9)].copy()
universe.head()
# ## Quiz: Nothing here should be allowed to look at returns when forming the portfolio.
# Make this impossible by removing the Daily returns data from the dataframe. Drop the DlyReturn field from the dataframe.
# TODO: drop DlyReturn column
universe = df.loc[(df['IssuerMarketCap'] >= 1e9) | (abs(df['x.opt.previous']) > 0)].copy()
## this will extract all of the factors, including the alphas
# list(universe) gets a list of the column names of the dataframe
all_factors = factors_from_names(list(universe))
all_factors
# ## Alpha factors
#
# Just a reminder that we chose four of these factors that represent previously effective alpha factors. Since these factors became well known over time, they were added to the Barra data set. For the time frame that we're running the back-test, these were effective alpha factors.
alpha_factors #4 alpha factors
# ## Quiz: risk factors
#
# The risk factors we'll use are all the factors that are not alpha factors. Complete the setdiff function so that it takes a superset, a subset, and returns the difference as a set.
#
# diff= SuperSet \ Subset
def setdiff(superset, subset):
# TODO
s = set(subset)
diffset = [x for x in superset if x not in s]
return(diffset)
risk_factors = setdiff(all_factors, alpha_factors)
# 77 risk factors
len(risk_factors)
# Save initial holdings in a variable for easier access. We'll later use it in matrix multiplications, so let's convert this to a numpy array. We'll also use another variable to represent the current holdings, which are to be run through the optimizer. We'll set this to be a copy of the previous holdings. Later the optimizer will continually update this to optimize the objective function.
## initial holdings (before optimization)
# optimal holding from prior day
h0 = np.asarray( universe['x.opt.previous'] )
h = h0.copy()
# ## Matrix of Risk Factor Exposures $\textbf{B}$
#
# The dataframe contains several columns that we'll use as risk factors exposures. Extract these and put them into a matrix.
#
# The data, such as industry category, are already one-hot encoded, but if this were not the case, then using `patsy.dmatrices` would help, as this function extracts categories and performs the one-hot encoding. We'll practice using this package, as you may find it useful with future data sets. You could also store the factors in a dataframe if you prefer to avoid using patsy.dmatrices.
#
# #### How to use patsy.dmatrices
#
# patsy.dmatrices takes in a formula and the dataframe. The formula tells the function which columns to take. The formula will look something like this:
# `SpecRisk ~ 0 + USFASTD_AERODEF + USFASTD_AIRLINES + ...`
# where the variable to the left of the ~ is the "dependent variable" and the others to the right are the independent variables (as if we were preparing data to be fit to a model).
#
# This just means that the pasty.dmatrices function will return two matrix variables, one that contains the single column for the dependent variable `outcome`, and the independent variable columns are stored in a matrix `predictors`.
#
# The `predictors` matrix will contain the matrix of risk factors, which is what we want. We don't actually need the `outcome` matrix; it's just created because that's the way patsy.dmatrices works.
# Note that we chose "SpecRisk" simply because it's not one of the USFAST factors.
# it will be discarded in the next step.
formula = get_formula(risk_factors, "SpecRisk")
formula
# the factors will be in the second returned variable (predictors)
# the outcome variable contains the SpecRisk data, which we don't actually need here
outcome, predictors = patsy.dmatrices(formula,universe)
# `predictors` contains the factor exposures of each asset to each factor.
predictors.shape
# ## Factor exposure matrix $\textbf{B}$
#
#
# Remember, the factor exposure matrix has the exposure of each asset to each factor. Thee number of rows is number of assets, and number of columns is the number of factors.
# +
def NROW(x):
return(np.shape(x)[0])
def NCOL(x):
return(np.shape(x)[1])
# -
# ## Quiz
#
# Set the factor exposure matrix and its transpose, using one of the outputs from calling patsy.dmatrices
# +
## TODO: risk exposure matrix:
B = predictors
BT = B.transpose()
k = NCOL(B) #number of factors (77)
n = NROW(B) #number of assets (2000+)
# -
# ## Factor covariance matrix $\textbf{F}$
#
# We can improve on the factor covariance matrix by reducing noise and also increasing computational efficiency.
#
# If we have, 70 risk factors in our risk model, then the covariance matrix of factors is a 70 by 70 square matrix. The diagonal contains the variances of each factor, while the off-diagonals contain the pairwise covariances of two different risk factors.
# In general, it’s good to have a healthy suspicion of correlations and covariances, and to ask if correlation data adds information or just more noise. One way to be conservative about the information in a covariance matrix is to shrink the covariances, or even reduce them to zero. In other words, we could keep just the variances along the diagonal, and set the covariances in the off-diagonals to zero.
# In the case where we’re using the covariance matrix in a risk factor model, there’s also some additional intuition for why we can try using just the variances, and discard the covariances. The goal of the optimizer is to reduce the portfolio’s exposure to these risk factors. So if the optimizer reduces the portfolio’s exposure to risk factor “one”, and also reduces its exposure to risk factor “two”, then it’s less important to know exactly how factor one varies with factor two.
#
# You may wonder what are the benefits of throwing away the information about the covariances. In addition to making your model more conservative, and limiting possible noise in your data, a diagonal matrix also makes matrix operations more efficient. This theme of computational efficiency is one that you’ll come across in many use cases, including backtesting. Backtesting is a computationally and time-intensive process, so the more efficient you can make it, the more quickly you can test your alphas, and iterate to make improvements.
#
# ## Create Factor covariance matrix $\textbf{F}$
#
# You can try getting all covariances into the matrix. Notice that we'll run into some issues where the covariance data doesn't exist.
#
# One important point to remember is that we need to order the factors in the covariance matrix F so that they match up with the order of the factors in the factor exposures matrix B.
#
# Note that covariance data is in percentage units squared, so to use decimals, so we'll rescale it to convert it to decimal.
# +
## With all covariances
def colnames(X):
if(type(X) == patsy.design_info.DesignMatrix):
return(X.design_info.column_names)
if(type(X) == pandas.core.frame.DataFrame):
return(X.columns.tolist())
return(None)
## extract a diagonal element from the factor covariance matrix
def get_cov_version1(cv, factor1, factor2):
try:
return(cv.loc[(cv.Factor1==factor1) & (cv.Factor2==factor2),"VarCovar"].iloc[0])
except:
print(f"didn't find covariance for: factor 1: {factor1} factor2: {factor2}")
return 0
def diagonal_factor_cov_version1(date, B):
"""
Notice that we'll use the order of column names of the factor exposure matrix
to set the order of factors in the factor covariance matrix
"""
cv = covariance[date]
k = NCOL(B)
Fm = np.zeros([k,k])
for i in range(0,k):
for j in range(0,k):
fac1 = colnames(B)[i]
fac2 = colnames(B)[j]
# Convert from percentage units squared to decimal
Fm[i,j] = (0.01**2) * get_cov_version1(cv, fac1, fac2)
return(Fm)
# -
# Here's an example where the two factors don't have covariance data for the date selected
cv = covariance['20031211']
cv.loc[(cv.Factor1=='USFASTD_AERODEF') & (cv.Factor2=='USFASTD_ALUMSTEL')]
# We can see where all the factor covariances aren't found in the data.
#
# ## Which date?
#
# Recall that there's a DataDate column and DlyReturnDate column in the dataframe. We're going to use a date to access the covariance data. Which date should we use?
df.head()
# ## Answer here
#
#
# ## Quiz
# Choose the correct date, then use the `diagonal_factor_cov_version1` to get the factor covariance matrix of that date.
# TODO
date = str(int(universe['DataDate'][1]))
print(date, end =" ")
F_version1 = diagonal_factor_cov_version1(date, B)
# ## Quiz: Create matrix of factor variances
#
# Just use the factor variances and set the off diagonal covariances to zero.
# +
def colnames(X):
if(type(X) == patsy.design_info.DesignMatrix):
return(X.design_info.column_names)
if(type(X) == pandas.core.frame.DataFrame):
return(X.columns.tolist())
return(None)
## extract a diagonal element from the factor covariance matrix
def get_var(cv, factor):
# TODO
return(cv.loc[(cv.Factor1==factor) & (cv.Factor2==factor),"VarCovar"].iloc[0])
def diagonal_factor_cov(date, B):
"""
Notice that we'll use the order of column names of the factor exposure matrix
to set the order of factors in the factor covariance matrix
"""
# TODO: set the variances only
cv = covariance[date]
k = NCOL(B)
Fm = np.zeros([k,k])
for j in range(0,k):
fac = colnames(B)[j]
Fm[j,j] = (0.01**2) * get_var(cv, fac)
return(Fm)
# -
## factor variances
# gets factor vars into diagonal matrix
# takes B to know column names of B; F will be multipled by B later
# F is square; so row and col names must match column names of B.
F = diagonal_factor_cov(date, B)
F.shape
# Note how the off diagonals are all set to zero.
# ## alpha combination
#
# As a simple alpha combination, combine the alphas with equal weight. In the project, you're welcome to try other ways to combine the alphas. For example, you could calculate some metric for each factor, which indicates which factor should be given more or less weight.
#
# ## Scale factor exposures
#
# Note that the terms that we're calculating for the objective function will be in dollar units. So the expected return $-\alpha^T h$ will be in dollar units. The $h$ vector of portfolio holdings will be in dollar units. The vector of alpha factor exposures $\alpha$ will represent the percent change expected for each stock. Based on the ranges of values in the factor exposure data, which are mostly between -5 and +5 and centered at zero, **we'll make an assumption that a factor exposure of 1 maps to 1 basis point of daily return on that stock.**
#
# So we'll convert the factor values into decimals: 1 factor exposure value $\rightarrow \frac{1}{10,000}$ in daily returns. In other words, we'll rescale the alpha factors by dividing by 10,000.
#
# This is to make the term representing the portfolio's expected return $\alpha^T h$ be scaled so that it represents dollar units.
alpha_factors
def model_matrix(formula, data):
outcome, predictors = patsy.dmatrices(formula, data)
return(predictors)
## matrix of alpha factors
B_alpha = model_matrix(get_formula(alpha_factors, "SpecRisk"), data = universe)
B_alpha
# ## Quiz
#
# Sum across the rows, then re-scale so that the expression $\mathbf{\alpha}^T \mathbf{h}$ is in dollar units.
def rowSums(m):
# TODO
return(np.sum(m, axis=1))
# TODO
scale = 1e-4
alpha_vec = scale * rowSums(B_alpha) #sum across rows (collapse 4 columns into one)
alpha_vec.shape
# ## Original method of calculating common risk term
#
# Recall that the common risk term looks like this:
# $\textbf{h}^T\textbf{BFB}^T\textbf{h}$
#
# Where h is the vector of portfolio holdings, B is the factor exposure matrix, and F is the factor covariance matrix.
#
# We'll walk through this calculation to show how it forms an N by N matrix, which is computationally expensive, and may lead to memory overflow for large values of N.
np.dot( np.dot( h.T, np.matmul( np.matmul(B,F),BT) ), h)
tmp = np.matmul(B,F)
tmp.shape
# this makes an N by matrix (large)
tmp = np.matmul(tmp,BT)
tmp.shape
tmp = np.matmul(h.T,tmp)
tmp.shape
tmp = np.dot(tmp,h)
tmp.shape
tmp
# ## Efficiently calculate common risk term (avoid N by N matrix)
#
# Calculate the portfolio risk that is attributable to the risk factors:
# $\mathbf{h}^T\mathbf{BFB}^T\mathbf{h}$
#
# Note that this can become computationally infeasible and/or slow. Use matrix factorization and carefully choose the order of matrix multiplications to avoid creating an N by N matrix.
#
# #### square root of a matrix.
#
# We can find a matrix $\mathbf{B}$ that's the matrix square root of another matrix $\mathbf{A}$, which means that if we matrix multiply $\mathbf{BB}$, we'd get back to the original matrix $\mathbf{A}$.
#
# Find $\mathbf{Q}$ such that $\mathbf{Q}^T\mathbf{Q}$ is the same as $\mathbf{BFB}^T$. Let's let $\mathbf{G}$ denote the square root of matrix $\mathbf{F}$, so that $\mathbf{GG} = \mathbf{F}$.
#
# Then the expression for the covariance matrix of assets, $\mathbf{BFB}^T$, can be written as $\mathbf{BGGB}^T$.
#
# Let's let $\mathbf{Q}=\mathbf{GB}^T$ and let $\mathbf{Q}^T=\mathbf{BG}$, which means we can rewrite $\mathbf{BGGB}^T = \mathbf{Q}^T\mathbf{Q}$, and the common risk term is $\mathbf{h}^T\mathbf{Q}^T\mathbf{Qh}$
#
# Also, note that we don't have to calculate $\mathbf{BFB}^T$ explicitly, because the actual value we wish to calculate in the objective function will apply the holdings $\mathbf{h}$ to the covariance matrix of assets.
# ## Quiz: matrix square root of F
#
# We'll call this square root matrix $\mathbf{G}$
#
# Use [scipy.linalg.sqrtm](https://docs.scipy.org/doc/scipy-0.15.1/reference/generated/scipy.linalg.sqrtm.html)
#
# TODO
G = scipy.linalg.sqrtm(F)
G.shape
# Double check that multiplying the square root matrix to itself returns us back to the original matrix of factor variances.
np.matmul(G,G) - F
# ## Quiz: calculate $\textbf{Q}$ and $\textbf{Q}^T$
# TODO
# Q = GB'
# Q should be a short and wide matrix
Q = np.matmul(G, BT)
Q.shape
# TODO
# Q' = BG
# Q should be a tall and narrow matrix
QT = np.matmul(B,G)
QT.shape
# notice we could also use the transpose of Q to get Q'
QT - Q.transpose()
# ## Quiz: Include portfolio holdings
#
# So the original formula of
# $h^TBFB^Th$ became
# $h^TBGGB^Th$, where $GG = F$.
#
# And then, if we let $Q^T=BG$ and $Q = GB^T$:
# $h^TQ^TQh$
#
#
# Let $R = Q h$ and $R^T = h^T Q^T$:
#
# The risk term becomes:
# $R^TR$, where $R^T=h^TQ$ and $R=Q^Th$
#
# So an important point here is that we don't want to multiply $Q^TQ$ itself, because this creates the large N by N matrix. We want to multiply $h^TQ^T$ and $Qh$ separately, creating vectors of length k (k is number of risk factors).
# TODO
# R = Qh
R = np.matmul(Q, h)
R.shape
# TODO
# R' = Q'h'
RT = np.matmul(h.T,QT)
RT.shape
# ## Notice how we avoided creating a full N by N matrix
#
# Also, notice that if we have Q, we can take its transpose to get $Q^T$ instead of doing the matrix multiplication.
#
# Similarly, if we have R, which is a vector, we notice that $R^TR$ is the same as taking the dot product. In other words, it's squaring each element in the vector R, and adding up all the squared values.
#
# $R^TR = \sum_{i}^{k}(r_i^2)$
#
# ## Quiz: Put it all together: calculate common risk term efficiently
# +
## TODO: common risk term in term
# TODO: calculate square root of F
G = scipy.linalg.sqrtm(F)
# TODO: calculate Q
Q = np.matmul(G, BT)
# TODO: calculate R
R = np.matmul(Q, h)
# TODO: calculate common risk term
common_risk = np.sum( R ** 2)
# -
# ## Specific Risk term
#
# The portfolio's variance that is specific to each asset is found by combining the holdings with the specific variance matrix:
#
# $h^TSh$, where $h^T$ is a 1 by N vector, S is an N by N matrix, and h is an N by 1 vector.
#
# Recall that S is a diagonal matrix, so all the off-diagonals are zero. So instead of doing the matrix multiplication, we could save computation by working with the vector containing the diagonal values.
#
# $h^TSh = \sum_i^{N}(h_i^2 \times S_i)$ because $S$ is a diagonal matrix.
## check the unit of measure of SpecRisk
# Notice that these are in percent; multiply by .01 to get them back to decimals.aa
universe['SpecRisk'][0:2]
# ## Quiz: Specific Risk term
#
# Given specific risk (volatility), calculate specific variance. First re-scale the specific risk data so that it's in decimal instead of percent.
# +
## TODO: specific variance : rescale it and then square to get specific variance
specVar = (0.01 * universe['SpecRisk']) ** 2
# TODO: specific risk term (include holdings)
spec_risk_term = np.dot(specVar**2, specVar)
# -
# ## Maximize portfolio returns
#
# Since the alpha vector $\mathbf{\alpha}$ is supposed to be indicative of future asset returns, when we look at a portfolio of assets, the weighted sum of these alphas $\mathbf{\alpha}^T \mathbf{h}$ is predictive of the portfolio's future returns. We want to maximize the portfolio's expected future returns, so we want to minimize the negative of portfolio's expected returns $-\mathbf{\alpha}^T \mathbf{h}$
## TODO
expected_return = np.dot(specVar, alpha_vec)
# ## Linear price impact of trading
#
# Assume transaction cost is linearly related to the trade size as a fraction of the average daily volume. Since we won't know the actual daily volume until the day that we're executing, we want to use past data as an estimate for future daily volume. This would be kind of noisy if we simply use the prior day's daily volume, so we'd prefer a more stable estimate like a 30 day rolling average.
#
# A commonly used **estimate for linear market impact is that if a trade size is 1% of the ADV, this moves the price by 10 basis points (1/10,000).**
#
# $Trade size_{i,t}$ is the fraction of your trade relative to the average dollar volume estimated for that stock, for that day.
#
# $Trade_{i,t}$ = dollar amount to trade = $h_{t} - h_{t-1}$, which is the new holding of the asset minus the previous holding.
#
# $ADV_{i,t}$: (average dollar volume) is total dollar amount expected to be traded, based on a moving average of historical daily volume.
#
# $TradeSize_{i,t} = \frac{Trade_{i,t}}{ADV_{i,t}}$: The size of the trade relative to the estimated daily volume.
#
# $\% \Delta Price_{i,t}$ = price change due to trading, as a fraction of the original price (it's a percent change).
# We'll write out the ratio: change in price divided by the trade size.
#
# $ \frac{\% \Delta price_{i,t}}{TradeSize_{i,t}} = \frac{10 bps}{1\%}$
#
# $ \frac{\% \Delta price_{i,t}}{TradeSize_{i,t}} = \frac{10/10^4}{1/100}$
#
# $ \frac{\% \Delta price_{i,t}}{TradeSize_{i,t}} = \frac{10^{-3}}{10^{-2}}$
#
# $ \frac{\% \Delta price_{i,t}}{TradeSize_{i,t}} = 10^{-1}$
#
# Now we'll move things around to solve for the change in price.
#
# $\% \Delta price_{i,t} = 10^{-1} \times TradeSize_{i,t}$
#
# We defined TradeSize to be the Trade divided by ADV.
#
# $\% \Delta price_{i,t} = 10^{-1} \times \frac{Trade_{i,t}}{ADV_{i,t}}$
#
# Note that Trade is the current position minus the prior day's position
#
# $\% \Delta price_{i,t} = 10^{-1} \times \frac{h_{i,t} - h_{i,t-1}}{ADV_{i,t}}$
#
# For convenience, we'll combine the constant $10^{-1}$ and $\frac{1}{ADV_{i}}$ and call it lambda $\lambda_{i}$
#
# $\% \Delta price_{i,t} = \lambda_{i,t} \times (h_{i,t} - h_{i,t-1})$ where $\lambda_{i,t} = 10^{-1}\times \frac{1}{ADV_{i,t}} = \frac{1}{10 \times ADV_{i,t}}$
#
# Note that since we're dividing by $ADV_{i,t}$, we'll want to handle cases when $ADV_{i,t}$ is missing or zero. In those instances, we can set $ADV_{i,t}$ to a small positive number, such as 10,000, which, in practice assumes that the stock is illiquid.
#
# Represent the market impact as $\Delta price_{i} = \lambda_{i} (h_{i,t} - h_{i,t-1})$. $\lambda_{i}$ incorporates the $ADV_{i,t}$. Review the lessons to see how to do this.
#
# Note that since we're dividing by $ADV_{i,t}$, we'll want to handle cases when $ADV_{i,t}$ is missing or zero. In those instances, we can set $ADV_{i,t}$ to a small positive number, such as 10,000, which, in practice assumes that the stock is illiquid.
# ## Quiz
#
# If the ADV field is missing or zero, set it to 10,000.
# +
# TODO: if missing, set to 10000
universe.loc[np.isnan(universe['ADTCA_30']), 'ADTCA_30'] = 1.0e4 ## assume illiquid if no volume information
# TODO: if zero, set to 10000
universe.loc[universe['ADTCA_30'] == 0, 'ADTCA_30'] = 1.0e4 ## assume illiquid if no volume information
# -
# ## Quiz: calculate Lambda
# TODO
adv = universe['ADTCA_30']
Lambda = 0.1 / adv
# ## Quiz: transaction cost term
#
# Transaction cost is change in price times dollar amount traded. For a single asset "i":
#
# $tcost_{i,t} = (\% \Delta price_{i,t}) \times (DollarsTraded_{i,t})$
#
# $tcost_{i,t} = (\lambda_{i,t} \times (h_{i,t} - h_{i,t-1}) ) \times (h_{i,t} - h_{i,t-1})$
#
# Notice that we can simplify the notation so it looks like this:
#
# $tcost_{i,t} = \lambda_{i,t} \times (h_{i,t} - h_{i,t-1})^2$
#
# The transaction cost term to be minimized (for all assets) is:
#
# $tcost_{t} = \sum_i^{N} \lambda_{i,t} (h_{i,t} - h_{i,t-1})^2$
# where $\lambda_{i,t} = \frac{1}{10\times ADV_{i,t}}$
#
# For matrix notation, we'll use a capital Lambda, $\Lambda_{t}$, instead of the lowercase lambda $\lambda_{i,t}$.
#
# $tcost_{t} = (\mathbf{h}_{t} - \mathbf{h}_{t-1})^T \mathbf{\Lambda}_t (\mathbf{h}_{t} - \mathbf{h}_{t-1})$
#
# Note that we'll pass in a vector of holdings as a numpy array. For practice, we'll use the h variable that is initialized to zero.
# TODO
tcost = np.dot( (h - h0) ** 2, Lambda)
# ## objective function
#
# Combine the common risk, idiosyncratic risk, transaction costs and expected portfolio return into the objective function. Put this inside a function.
#
# Objective function is:
# factor risk + idiosyncratic risk - expected portfolio return + transaction costs
# $f(\mathbf{h}) = \frac{1}{2}\kappa \mathbf{h}_t^T\mathbf{Q}^T\mathbf{Q}\mathbf{h}_t + \frac{1}{2} \kappa \mathbf{h}_t^T \mathbf{S} \mathbf{h}_t - \mathbf{\alpha}^T \mathbf{h}_t + (\mathbf{h}_{t} - \mathbf{h}_{t-1})^T \mathbf{\Lambda} (\mathbf{h}_{t} - \mathbf{h}_{t-1})$
#
# ## Risk Aversion $\kappa$
#
# The risk aversion term is set to target a particular gross market value (GMV), or to target a desired volatility. In our case, we tried a few values of the risk aversion term, ran the backtest, and calculated the GMV. Ideally, a quant who is just starting out may have a targeted GMV of 50 million. A risk aversion term of $10^{-6}$ gets the GMV to be in the tens of millions. A higher risk aversion term would decrease the GMV, and a lower risk aversion term would increase the GMV, and also the risk. Note that this isn't necessarily a linear mapping, so in practice, you'll try different values and check the results.
#
# Also, in practice, you'd normally keep the risk aversion term constant, unless your fund is accepting more investor cash, or handling redemptions. In those instances, the fund size itself changes, so the targeted GMV also changes. Therefore, we'd adjust the risk aversion term to adjust for the desired GMV.
#
# Also, note that we would keep this risk aversion term constant, and not adjust it on a daily basis. Adjusting the risk aversion term too often would result in unecessary trading that isn't informed by the alphas.
# ## Quiz
# An important point is to think about what matrices can be multiplied independently of the vector of asset holdings, because those can be done once outside of the objective function. The rest of the objective function that depends on the holdings vector will be evaluated inside the objective function multiple times by the optimizer, as it searches for the optimal holdings.
#
#
# For instance,
#
# $\mathbf{h}^T\mathbf{BFB}^T\mathbf{h}$ became
# $\mathbf{h}^T\mathbf{BGGB}^T\mathbf{h}$, where $\mathbf{GG} = \mathbf{F}$.
#
# And then, if we let $\mathbf{Q}^T=\mathbf{BG}$ and $\mathbf{Q} = \mathbf{GB}^T$:
# $\mathbf{h}^T\mathbf{Q}^T\mathbf{Qh}$
#
# Let $\mathbf{R} = \mathbf{Q h}$ and $\mathbf{R}^T = \mathbf{h}^T \mathbf{Q}^T$:
#
# The risk term becomes:
# $\mathbf{R}^T\mathbf{R}$, where $\mathbf{R}^T=\mathbf{h}^T\mathbf{Q}$ and $\mathbf{R}=\mathbf{Q}^T\mathbf{h}$
#
# * Can we pre-compute Q outside of the objective function?
# * Can we pre-compute R outside of the objective function?
# #### Answer
# Q doesn't depend on h, the holdings vector, so it can be pre-computed once outside of the objective function.
#
# R is created using h, the holdings vector. This should be computed each time the objective function is called, not pre-computed beforehand.
# ## Risk Aversion parameter
#
# The risk aversion term is set to target a particular gross market value (GMV), or to target a desired volatility.
#
# The gross market value is the dollar value of the absolute value of the long and short positions.
#
# $ GMV = \sum_i^N(|h_{i,t}|)$
#
# When we think about what it means to take more risk when investing, taking bigger bets with more money is a way to take on more risk. So the risk aversion term controls how much risk we take by controlling the dollar amount of our positions, which is the gross market value.
#
# In our case, we tried a few values of the risk aversion term, ran the backtest, and calculated the GMV. Ideally, a quant who is just starting out may have a targeted book size of 50 million. In other words, they try to keep their GMV around 50 million.
#
# A risk aversion term of $10^{-6}$ gets the GMV to be in the tens of millions. A higher risk aversion term would decrease the GMV, and a lower risk aversion term would increase the GMV, and also the risk. Note that this isn't necessarily a linear mapping, so in practice, you'll try different values and check the results.
#
# Also, in practice, you'd normally keep the risk aversion term constant, unless your fund is accepting more investor cash, or handling redemptions. In those instances, the fund size itself changes, so the targeted GMV also changes. Therefore, we'd adjust the risk aversion term to adjust for the desired GMV.
#
# Also, note that we would keep this risk aversion term constant, and not adjust it on a daily basis. Adjusting the risk aversion term too often would result in unnecessary trading that isn't informed by the alphas.
#
## Risk aversion
risk_aversion=1.0e-6
# ## Quiz: define objective function
#
# Combine the common risk, idiosyncratic risk, transaction costs and expected portfolio return into the objective function. Put this inside a function.
#
# Objective function is:
# factor risk + idiosyncratic risk - expected portfolio return + transaction costs
# $f(\mathbf{h}) = \frac{1}{2}\kappa \mathbf{h}_t^T\mathbf{Q}^T\mathbf{Q}\mathbf{h}_t + \frac{1}{2} \kappa \mathbf{h}_t^T \mathbf{S} \mathbf{h}_t - \mathbf{\alpha}^T \mathbf{h}_t + (\mathbf{h}_{t} - \mathbf{h}_{t-1})^T \mathbf{\Lambda} (\mathbf{h}_{t} - \mathbf{h}_{t-1})$
#
def func(h):
# TODO: define the objective function, where h is the vector of asset holdings
f = 0.0
f += 0.5 * risk_aversion * np.sum( np.matmul(Q, h) ** 2 )
f += 0.5 * risk_aversion * np.dot(h ** 2, specVar) #since Specific Variance is diagonal, don't have to do matmul
f -= np.dot(h, alpha_vec)
f += np.dot( (h - h0) ** 2, Lambda)
return(f)
# ## Gradient
#
# Before, when we used cvxpy, we didn't have to calculate the gradient, because the library did that for us.
#
# Objective function is:
# factor risk + idiosyncratic risk - expected portfolio return + transaction costs
# $f(\mathbf{h}) = \frac{1}{2}\kappa \mathbf{h}^T\mathbf{Q}^T\mathbf{Qh} + \frac{1}{2} \kappa \mathbf{h}^T \mathbf{S h} - \mathbf{\alpha^T h} + (\mathbf{h}_{t} - \mathbf{h}_{t-1})^T \Lambda (\mathbf{h}_{t} - \mathbf{h}_{t-1})$
#
#
# Let's think about the shape of the resulting gradient. The reason we're interested in calculating the derivative is so that we can tell the optimizer in which direction, and how much, it should shift the portfolio holdings in order to improve the objective function (minimize variance, minimize transaction cost, and maximize expected portfolio return). So we want to calculate a derivative for each of the N assets (about 2000+ in our defined universe). So the resulting gradient will be a row vector of length N.
#
# The gradient, or derivative of the objective function, with respect to the portfolio holdings h, is:
#
# $f'(\mathbf{h}) = \frac{1}{2}\kappa (2\mathbf{Q}^T\mathbf{Qh}) + \frac{1}{2}\kappa (2\mathbf{Sh}) - \mathbf{\alpha} + 2(\mathbf{h}_{t} - \mathbf{h}_{t-1}) \mathbf{\Lambda}$
#
# We can check that each of these terms is a row vector with one value for each asset (1 by N row vector)
# ## Quiz
#
# Calculate the gradient of the common risk term:
#
# $\kappa (\mathbf{Q}^T\mathbf{Qh})$
# TODO: gradient of common risk term
tmp = risk_aversion * np.matmul(QT, np.matmul(Q,h))
# Verify that the calculation returns one value for each asset in the stock universe (about 2000+ )
tmp.shape
# ## Quiz
#
# Calculate gradient of idiosyncratic risk term
#
# $\kappa (\mathbf{Sh})$
# TODO: idiosyncratic risk gradient
tmp = risk_aversion * specVar * h
tmp.shape
# ## Quiz
#
# Calculate the gradient of the expected return
#
# $- \mathbf{\alpha} $
# TODO: expected return gradient
tmp = -alpha_vec
tmp.shape
# ## Quiz
#
# Calculate the gradient of the transaction cost.
#
# $ 2(\mathbf{h}_{t} - \mathbf{h}_{t-1}) \mathbf{\Lambda}$
# transaction cost
tmp = 2 * (h - h0 ) * Lambda
tmp.shape
# ## Quiz: Define gradient function
#
# Put this all together to define the gradient function. The optimizer will use this to make small adjustments to the portfolio holdings.
#
# #### gradient (slightly cleaned up)
#
# We'll simplify the expression a bit by pulling the common $\kappa$ out of the common risk and specific risk. Also, the 1/2 and 2 cancel for both risk terms.
#
# $f'(\mathbf{h}) = \frac{1}{2}\kappa (2\mathbf{Q}^T\mathbf{Qh}) + \frac{1}{2}\kappa (2\mathbf{h}^T\mathbf{S}) - \mathbf{\alpha} + 2(\mathbf{h}_{t} - \mathbf{h}_{t-1})\cdot \Lambda$
#
# becomes
#
# $f'(\mathbf{h}) = \kappa (\mathbf{Q}^T\mathbf{Qh} + \mathbf{Sh}) - \mathbf{\alpha} + 2(\mathbf{h}_{t} - \mathbf{h}_{t-1}) \mathbf{\Lambda}$
# Solution
def grad(x):
# TODO
g = risk_aversion * (np.matmul(QT, np.matmul(Q,h)) + \
(specVar * h) ) - alpha_vec + \
2 * (h-h0) * Lambda
return(np.asarray(g))
# ## Optimizer
#
# Choose an optimizer. You can read about these optimizers:
#
# * L-BFGS
# * Powell
# * Nelder-Mead
# * Conjugate Gradient
#
# In this [page about math optimization](http://scipy-lectures.org/advanced/mathematical_optimization/)
#
# Also read the [scipy.optimize documentation](https://docs.scipy.org/doc/scipy/reference/optimize.html)
#
# Pass in the objective function, prior day's portfolio holdings, and the gradient.
# +
# TODO
optimizer_result = scipy.optimize.fmin_l_bfgs_b("""<your code here>""", """<your code here>""", fprime="""<your code here>""")
h1 = optimizer_result[0]
# -
opt_portfolio = pd.DataFrame(data = {"Barrid" : universe['Barrid'], "h.opt" : h1})
opt_portfolio.head()
# ## risk exposures
# factor exposures times the portfolio holdings for each asset, gives the portfolio's exposure to the factors (portfolio's risk exposure).
#
# $\mathbf{B}^T\mathbf{h}$
# +
# TODO: risk exposures
risk_exposures = np.matmul("""<your code here>""", """<your code here>""")
# put this into a pandas series
pd.Series(risk_exposures, index = colnames(B))
# -
# ## Quiz: alpha exposures
#
# The portfolio's exposures to the alpha factors is equal to the matrix of alpha exposures times the portfolio holdings. We'll use the holdings returned by the optimizer.
#
# $\textbf{B}_{\alpha}^T\mathbf{h}$
# +
# Solution: portfolio's alpha exposure
alpha_exposures = np.matmul("""<your code here>""", """<your code here>""")
# put into a pandas series
pd.Series(alpha_exposures, index = colnames(B_alpha))
# -
# ## Hints for the project
#
# You'll be putting this optimization code into functions so that you can call the optimizer in a loop, as the backtester walks through each day in the data.
# ## Solution notebook
#
# The solution notebook is [here](optimization_with_tcosts_solution.ipynb)
| TradingAI/AI Algorithms in Trading/Lesson 26 - Optimization with Transaction Costs /optimization_with_tcosts.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Explorando la historia
#
# Si queremos ver los cambios que hemos hecho en los distintos pasos, podemos usar `git diff` de nuevo, pero con la notación `HEAD~1`, `HEAD~2`, y así, para referirse a los commits anteriores:
#
# ```bash
# $ git diff HEAD~1 mars.txt
# ```
#
# ```
# diff --git a/mars.txt b/mars.txt
# index 315bf3a..b36abfd 100644
# --- a/mars.txt
# +++ b/mars.txt
# @@ -1,2 +1,3 @@
# Cold and dry, but everything is my favorite color
# The two moons may be a problem for Wolfman
# +But the Mummy will appreciate the lack of humidity
# ```
#
# ```bash
# $ git diff HEAD~2 mars.txt
# ```
#
# ```
# diff --git a/mars.txt b/mars.txt
# index df0654a..b36abfd 100644
# --- a/mars.txt
# +++ b/mars.txt
# @@ -1 +1,3 @@
# Cold and dry, but everything is my favorite color
# +The two moons may be a problem for Wolfman
# +But the Mummy will appreciate the lack of humidity
# ```
#
# De esta forma, podemos construir una cadena de commits.
# El mas reciente final de la cadena es referenciado como `HEAD`;
# podemos hacer referencia a los commits anteriores utilizando la notación `~`
# así `HEAD~1` (pronunciado "head menos uno")
# significa "el commit anterior"
# mientras que `HEAD~123` va 123 commits atrás desde donde estamos ahora.
#
# También nos podemos referir a los commits utilizando esas largas cadenas de digitos y letras que muestra `git log`.
# Estos son IDs únicos par los cambios,
# y "únicos" realmente significan únicos:
# cualquier cambios a cualquier conjunto de archivos en cualquier computadora
# tiene un identificador único de 40 caracteres.
# A nuestro primer commit le fue dado el ID
# f22b25e3<PASSWORD>6<PASSWORD>1e6<PASSWORD>73b,
# así que intentemos lo siguiente:
#
# ```bash
# $ git diff f22b25e3233b4645dabd0d81e651fe074bd8e73b mars.txt
# ```
#
# ```
# diff --git a/mars.txt b/mars.txt
# index df0654a..b36abfd 100644
# --- a/mars.txt
# +++ b/mars.txt
# @@ -1 +1,3 @@
# Cold and dry, but everything is my favorite color
# +The two moons may be a problem for Wolfman
# +But the Mummy will appreciate the lack of humidity
# ```
#
# Esta es la respuesta correcta,
# pero escribir cadenas aleatoria de 40 caracteres es un fastidio,
# de manera que Git nos deja usar tan solo los primeros caracteres:
#
# ```bash
# $ git diff f22b25e mars.txt
# ```
#
# ```
# diff --git a/mars.txt b/mars.txt
# index df0654a..b36abfd 100644
# --- a/mars.txt
# +++ b/mars.txt
# @@ -1 +1,3 @@
# Cold and dry, but everything is my favorite color
# +The two moons may be a problem for Wolfman
# +But the Mummy will appreciate the lack of humidity
# ```
#
# Muy bien! Así podemos guardar cambios los archivos y ver que hemos cambiado—ahora, ¿cómo podemos restaurar versiones antiguas de las cosas?
# Supongamos que accidentalmente sobreescribimos nuestro archivo:
#
# ```bash
# $ nano mars.txt
# $ cat mars.txt
# ```
# ```
# We will need to manufacture our own oxygen
# ```
#
# `git status` ahora nos dice que el archivo ha sido cambiado,
# pero este cambio no ha sido llevado al stage:
#
# ```bash
# $ git status
# ```
#
# ```
# # On branch master
# # Changes not staged for commit:
# # (use "git add <file>..." to update what will be committed)
# # (use "git checkout -- <file>..." to discard changes in working directory)
# #
# # modified: mars.txt
# #
# no changes added to commit (use "git add" and/or "git commit -a")
# ```
#
# Podemos devolver las cosas como estaban usando `git checkout`:
#
# ```bash
# $ git checkout HEAD mars.txt
# $ cat mars.txt
# ```
#
# ```
# Cold and dry, but everything is my favorite color
# The two moons may be a problem for Wolfman
# But the Mummy will appreciate the lack of humidity
# ```
#
# Como pueden adivinar de su nombre, `git checkout` quita el "check" (es decir, restaura) una versión vieja de un archivo.
# En este caso, le estamos diciendo a Gt que queremos recuperar la versión del archivo registrada en `HEAD`,
# la cual es el último commit guardado.
# Si queremos ir aún mas atrás, podemos utilizar en su lugar un identificador de commit:
#
# ```bash
# $ git checkout f22b25e mars.txt
# ```
#
# > ## No pierdas tu HEAD!
# > Arriba utilizamos
# >
# > ```bash
# > $ git checkout f22b25e mars.txt
# > ```
# >
# > para devolver mars.txt a su estado después del commit f22b25e.
# > Si olvidas `mars.txt` en ese comando, git te dirá que "You are in
# > 'detached HEAD' state." En ese estado, no deberías hacer ningún cambio.
# > Puedes arreglarlo volviendo a volviendo a fijar tu "head" usando ``git checkout master``
#
# Es importante recordar que
# debemos usar el número de commit que identifica el estado del respositorio *antes* del cambio que estamos intentando deshacer. Un error frecuente es utilizar el número del commit en el que hicimos el cambio del que nos queremos deshacer. En el ejemplo a continuación, queremos recuperar el estado desde antes del commit mas reciente (`HEAD~1`), el cual es el commit `f22b25e`:
#
# 
#
# Así, para ponerlo todo junto:
#
# > ## Como funciona Git, en forma de comic
# > 
#
# > ## Simplificando el Caso mas Común
# >
# > Si lees la salida de `git status` con cuidado,
# > verás que incluye esta pista:
# >
# > ```bash
# > (use "git checkout -- <file>..." to discard changes in working directory)
# > ```
# >
# > Tal como dice,
# > `git checkout` sin un identificador de versión restaura archivos al estado guardado en `HEAD`.
# > El guión doble `--` es necesario para separar los nombres de los archivos que están siendo recuperados
# > desde el propio comando:
# > sin este,
# > Git podría intentar usar el nombre del archivo como el identificador del commit.
#
# El hecho de que los archivos se pueden revertir uno por uno
# tiende a cambiar la forma en que la gente organiza su trabajo.
# Si todo está en un documento de gran tamaño,
# Es difícil (pero no imposible) deshacer los cambios a la introducción
# sin deshacer también los cambios realizados después a la conclusión.
# Por otra parte, si la introducción y la conclusión se almacenan en archivos separados,
# moverse hacia atrás y hacia adelante en el tiempo se hace mucho más fácil.
| 05-historial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TensorFlow Tutorial - 3. Gradient Descent
#
# 본 문서는 TensorFlow 를 사용하여 Deep Learning을 구현하기 위한 기초적인 실습 자료이다.
#
# The code and comments are written by <NAME> <<EMAIL>><br>
# Upgraed to Tensorflow v1.9 by NamJungGu <<EMAIL>>
#
# <a rel="license" href="http://creativecommons.org/licenses/by-nc-sa/4.0/"><img alt="Creative Commons License" style="border-width:0" src="https://i.creativecommons.org/l/by-nc-sa/4.0/88x31.png" /></a><br />This work is licensed under a <a rel="license" href="http://creativecommons.org/licenses/by-nc-sa/4.0/">Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License</a>.
#
#
#
# # Gradient Descent
#
# 경사 하강법(Gradient Descent)으로 인자 찾아내기
#
# 아래의 예제는 가중치 행렬W와 바이어스b를 경사하강법을 통해서 찾아내는 것을 보여줍니다. 목표값은 간단한 식으로 산출되도록 합니다.
# +
import tensorflow as tf
import numpy as np
# Numpy 랜덤으로 2개 짜리의 가짜 데이터 100개 만들기. (float64 -> float32로 변환)
x_data = np.float32(np.random.rand(100,2))
# 학습 레이블(목표값)은 아래의 식으로 산출. (W = [[1], [2]], b = 3)
y_data = np.dot(x_data,[[1.], [2.]]) + 3.
# -
# 입력 데이터와 W, b를 사용해 선형 모델을 정의합니다.
# b는 0,
b = tf.Variable(tf.zeros([1]))
# W는 1x2 형태의 웨이트 변수 (균등 랜덤값으로 초기화)
W = tf.Variable(tf.random_uniform([2,1], -1.0, 1.0))
y = tf.matmul( x_data,W) + b
# 이제 손실과 학습 함수를 정의 합니다. 평균 제곱 오차가 최소화 되는 지점을 경사하강법으로 구하게 됩니다.
# 손실 함수 정의
loss = tf.reduce_mean(tf.square(y - y_data))
# 경사하강법으로 손실 함수를 최소화 (0.5는 학습 비율)
optimizer = tf.train.GradientDescentOptimizer(0.5)
# 학습 오퍼레이션 정의
train = optimizer.minimize(loss)
# 학습 세션을 시작합니다.
# +
# 모든 변수를 초기화.
init = tf.initialize_all_variables()
# 세션 시작
sess = tf.Session()
sess.run(init)
# 200번 학습.
for step in range(0, 201):
sess.run(train)
if step % 20 == 0:
print("%4d %1.6f %1.6f %1.6f" %(step, sess.run(W)[0],sess.run(W)[1], sess.run(b)) )
# -
# 처음에 설정한 W와 b를 훌륭히 찾아냈습니다. 어떠신가요? 개인적인 느낌은 코드가 깔끔한 것 같습니다. 텐서플로우는 기존에 나와있는 딥러닝 프레임워크에 비해 단순하면서도 표현력이 풍부합니다.
#
# 텐서플로에 대해 좀 더 자세히 살펴보기 위해 MNIST 예제를 보겠습니다. (자세한 설명은 MNIST 예제 페이지를 참고하세요.)
| 1.tensorflow_jupyter/GradientDescent/GradientDescent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
pd.options.display.float_format = '{:.5f}'.format
import numpy as np
np.set_printoptions(precision=4)
np.set_printoptions(suppress=True)
import warnings
warnings.filterwarnings("ignore")
import os.path
def path_base(base_name):
current_dir = os.path.abspath(os.path.join(os.getcwd(), os.pardir))
print(current_dir)
data_dir = current_dir.replace('notebook','data')
print(data_dir)
data_base = data_dir + '\\' + base_name
print(data_base)
return data_base
base = pd.read_csv(path_base('db_plano_saude2.csv'))
base.head(3)
base.shape
X = base.iloc[:,0:1].values
print(X)
y = base.iloc[:,1].values
print(y)
from sklearn.tree import DecisionTreeRegressor
regressor = DecisionTreeRegressor()
regressor.fit(X,y)
# # Score
score = regressor.score(X,y)
score
# # plotar
import matplotlib.pyplot as plt
plt.scatter(X,y)
plt.title('Tree')
plt.xlabel('Idade')
plt.ylabel('Custo plano')
plt.plot(X,regressor.predict(X),color='red')
import numpy as np
X_teste = np.arange(min(X),max(X), 0.1)
X_teste = X_teste.reshape(-1,1)
plt.scatter(X,y)
plt.title('Tree')
plt.xlabel('Idade')
plt.ylabel('Custo plano')
plt.plot(X_teste,regressor.predict(X_teste),color='red')
print(regressor.predict(X))
| notebook/_RegressaoLinear/Tree_PlanoSaude.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (venv)
# language: python
# name: venv
# ---
# # Intro
# This notebook is used to do in-depth tests and exploration of trained models. It supports one of each type of model at a time: memory (classification), time (classification), and time (regression).
# # Notebook options
# There are 3 options for each model:
#
# 1. 'explore_model': Whether the notebook will explore this model at all. If False, all other options can be ignored.
#
# 2. 'path': Either 'auto', or a path to the folder creating the model's data (as saved by the model_saver object from persistence.py). If set to 'auto', the notebook will load the path from model_config.json (the same path used for apply_model.py).
#
# 3. 'training_mode': Either 'normal', 'test', or 'final'. Whether the model was trained on training data only ('normal'), training data and validation data ('test'), or training validation and testing data ('final'). This changes what data the evaluation reports and graphs are based on. In normal mode, the model will be evaluated using validation data; in test mode, the model will be evaluated using testing data; in final mode, we still use testing data, but these evaluations are practically meaningless since the model has trained on that data!
# + tags=[]
options = {'mem': {'explore_model': True,
'path': '/glade/work/jdubeau/model-saves/mem_class_tree_test2021-07-21-17:09/',
'training_mode': 'test'},
'time': {'explore_model': False,
'path': 'auto',
'training_mode': 'final'},
'time_regr': {'explore_model': True,
'path': 'auto',
'training_mode': 'final'}
}
model_types = ['mem', 'time', 'time_regr']
mem_options = options['mem']
time_options = options['time']
time_regr_options = options['time_regr']
# -
# # Imports and display options
from math import exp
import pandas as pd
import pickle
import numpy as np
import matplotlib.pyplot as plt
from itertools import product
from preprocessing import scale
from preprocessing import scale_other
from persistence import model_saver
from evaluation import print_feature_importances, \
plot_cm, print_cr, auc, \
plot_regr_performance, \
score_regressor
from apply_model import get_settings, predict_for_testing, \
custom_predict, predict_regr, \
translate_predictions, \
scale_predictions
from matplotlib import rcParams
rcParams.update({'figure.autolayout': True})
# # Loading models and associated data
# First we load the settings used in apply_model.py. We primarily need the paths to each model and the scaling options.
settings = get_settings()
# +
model_data = {}
for mtype in model_types:
if options[mtype]['path'] == 'auto':
model_path = settings['model_paths'][mtype]
else:
model_path = options[mtype]['path']
ms = model_saver()
ms.load(model_path)
data_names = ['notes', 'model', 'model_df', 'categories_dict',
'X_features', 'X_train', 'y_train_full', 'X_val',
'y_val_full', 'X_test', 'y_test_full']
data = ms.get_all()
model_data[mtype] = {data_names[i]: data[i] for i in range(len(data))}
mem_data = model_data['mem']
time_data = model_data['time']
time_regr_data = model_data['time_regr']
# -
full_names = {'mem': "Memory Classifier",
'time': "Time Classifier",
'time_regr': "Time Regressor"}
# # Exploration
# ## Notes
for mtype in model_types:
if options[mtype]['explore_model']:
print(f"{full_names[mtype]} notes:")
print(model_data[mtype]['notes'])
print("-------")
# ## Feature Importances
for mtype in model_types:
if options[mtype]['explore_model']:
X_features = model_data[mtype]['X_features']
model = model_data[mtype]['model']
print(f"{full_names[mtype]} feature importances:")
print_feature_importances(X_features, model)
print("-------")
# # Performance
# The following cells setup the relevant arrays (X_eval, y_eval, etc). to evaluate the models based on the indicated training modes.
for mtype in model_types:
if options[mtype]['explore_model']:
X_train = model_data[mtype]['X_train']
X_val = model_data[mtype]['X_val']
X_test = model_data[mtype]['X_test']
model_data[mtype]['X_train_norm'], \
model_data[mtype]['X_val_norm'], \
model_data[mtype]['X_test_norm'] = scale(X_train, X_val, X_test)
for mtype in model_types:
if options[mtype]['explore_model']:
y_train_full = model_data[mtype]['y_train_full']
y_val_full = model_data[mtype]['y_val_full']
y_test_full = model_data[mtype]['y_test_full']
if mtype in ['mem', 'time']:
y_train = np.ravel(y_train_full[mtype+'_category'])
y_val = np.ravel(y_val_full[mtype+'_category'])
y_test = np.ravel(y_test_full[mtype+'_category'])
else:
y_train = np.log10(np.ravel(y_train_full))
y_val = np.log10(np.ravel(y_val_full))
y_test = np.log10(np.ravel(y_test_full))
model_data[mtype]['y_train'] = y_train
model_data[mtype]['y_val'] = y_val
model_data[mtype]['y_test'] = y_test
for mtype in model_types:
if options[mtype]['explore_model']:
if options[mtype]['training_mode'] == 'normal':
X = model_data[mtype]['X_val_norm']
y = model_data[mtype]['y_val']
else:
X = model_data[mtype]['X_test_norm']
y = model_data[mtype]['y_test']
model_data[mtype]['X_eval'] = X
model_data[mtype]['y_eval'] = y
# ## Without custom predictions / scaling
# ### Confusion matrix
# The following cell plots the confusion matrix for a classification model. Note that the displayed percentages are taken over each row.
#
# Quick example: If the possible classes are 0, 1, and 2, and row 1 reads [0.30, 0.60, 0.10], that means that when the correct class was 1, the model guessed class 0 30% of the time, class 1 60% of the time, and class 2 10% of the time.
for mtype in ['mem', 'time']:
if options[mtype]['explore_model']:
X = model_data[mtype]['X_eval']
y = model_data[mtype]['y_eval']
plot_cm(X, y, model = model_data[mtype]['model'],
model_name = full_names[mtype],
save=False, path='')
# ### Classification report
# The next cell prints a classification report for each classifying model. To understand the statistics shown, consider the row corresponding to category 2:
#
# Precision = what percentage of the entries for which the model guessed category 2 were actually in category 2
#
# Recall = what percentage of the entries which were in category 2 were guessed as category 2 by the model
#
# f1-Score = a weighted average of precision (P) and recall (R) for category 2, specifically (2PR)/(P+R). Ranges from 0.0 (worst) to 1.0 (best).
#
# Support = how many entries belonged to category 2.
for mtype in ['mem', 'time']:
if options[mtype]['explore_model']:
X = model_data[mtype]['X_eval']
y = model_data[mtype]['y_eval']
print(f"{full_names[mtype]} classification report:")
print_cr(X, y, model = model_data[mtype]['model'])
# ### ROC AUC score
# The following cell prints ROC AUC scores for the two classifying models, that is, the area under the curve (AUC) of the Receiver Operating Characteristic curver (ROC).
#
# This is an average of ROC AUC scores for each individual category. The ROC AUC score for one category, say category 5 for example, represents the probability that when the model is given two random entries, one belonging to category 5 and one not, the model will assign a higher probability of being in category 5 to the entry that is actually in category 5.
#
# Hence the ROC AUC is always a score between 0.0 (worst) and 1.0 (best).
for mtype in ['mem', 'time']:
if options[mtype]['explore_model']:
X = model_data[mtype]['X_eval']
y = model_data[mtype]['y_eval']
print(f"{full_names[mtype]} ROC AUC score:")
auc(X, y, model = model_data[mtype]['model'])
# ### Regression accuracy score
# The next cell prints the training and evaluation score of the regression model.
#
# The score in this case R^2 coefficient, defined as 1 - u/v, where u is the sum over all samples of (y_true - y_pred)^2, and v is the sum over all samples of (y_true - y_mean)^2.
#
# The best possible score is 1.0, and scores can be negative, which would indicate that the model is worse than the strategy of always predicting the average value.
#
# If the training score is much higher than the evaluation score, that's evidence that the model was overfitting.
if options['time_regr']['explore_model']:
X_train_norm = model_data['time_regr']['X_train_norm']
y_train = model_data['time_regr']['y_train']
X_eval = model_data['time_regr']['X_eval']
y_eval = model_data['time_regr']['y_eval']
score_regressor(X_train_norm, y_train, X_eval, y_eval,
model=model_data['time_regr']['model'])
# ### Performance plot
if options['time_regr']['explore_model']:
X = model_data['time_regr']['X_eval']
y = model_data['time_regr']['y_eval']
plot_regr_performance(X, y, model=model_data['time_regr']['model'],
model_name=full_names['time_regr'],
save=False, path='performance-plot.png')
# ## With custom predictions / scaling
# The predictions in this section are designed to get a realistic idea of how the classification models perform, after the predictions are made in a custom way and the values are scaled how we like.
# This next cell sets up several useful arrays for simulating these predictions ('custom_preds', 'custom_values', and 'eval_values'). The first array represents the predicted categories using custom prediction; the second array represents the predicted values (translating and scaling the predicted categories); and the third array represents the actual correct values (used_mem and wall_time from the original data).
for mtype in ['mem', 'time']:
if options[mtype]['explore_model']:
X = model_data[mtype]['X_eval']
custom_preds = custom_predict(X, model_data[mtype]['model'])
model_data[mtype]['custom_preds'] = custom_preds
categories_dict = model_data[mtype]['categories_dict']
custom_values = scale_predictions(translate_predictions(custom_preds,
categories_dict),
**settings[mtype+'_scaling'])
model_data[mtype]['custom_values'] = custom_values
if mtype == 'mem':
target_name = 'used_mem'
else:
target_name = 'wall_time'
if options[mtype]['training_mode'] == 'normal':
eval_values = np.ravel(model_data[mtype]['y_val_full'][target_name])
else:
eval_values = np.ravel(model_data[mtype]['y_test_full'][target_name])
model_data[mtype]['eval_values'] = eval_values
# Here we show some example entries from the arrays we just made.
print(model_data['mem']['y_eval'][:5])
print(model_data['mem']['custom_preds'][:5])
print(model_data['mem']['eval_values'][:5])
print(model_data['mem']['custom_values'][:5])
# The next cell creates a dataframe containing all the relevant information for simulating the classifiers' behavior.
#
# In the evaluation dataframe for the time classifier, we ignore entries where the wall time was reportedly more than 12 hours (we are not interested in predicting those).
def evaluation_df(mtype):
if options[mtype]['training_mode'] == 'normal':
df = model_data[mtype]['y_val_full'].copy()
else:
df = model_data[mtype]['y_test_full'].copy()
if mtype == 'mem':
df['old_delta'] = df.apply(lambda row:
row['req_mem'] - row['used_mem'],
axis=1)
df['pred_cat'] = model_data[mtype]['custom_preds']
df['pred_value'] = model_data[mtype]['custom_values']
df['pred_delta'] = df.apply(lambda row: row['pred_value'] - row['used_mem'],
axis=1)
else:
df['time_delta'] = df.apply(lambda row:
43200 - row['wall_time'],
axis=1)
df['pred_cat'] = model_data[mtype]['custom_preds']
df['pred_value'] = model_data[mtype]['custom_values']
df['pred_delta'] = df.apply(lambda row: row['pred_value'] - row['wall_time'],
axis=1)
df = df[df.wall_time <= 43200]
return df
# Here we show an example of what this dataframe looks like.
evaluation_df('mem').head()
# The next two cells do the simulation we have been preparing for throughout this section.
def total_loss_info(mtype):
df = evaluation_df(mtype)
print(f"Number of entries predicted: {df.shape[0]}")
print("--------")
num_bad_jobs = df[df.pred_delta < 0].shape[0]
print("Number of jobs using more resources than predicted: "
+ f"{num_bad_jobs}")
print("Average delta for jobs using more resources than predicted: "
+ f"{round(df[df.pred_delta<0]['pred_delta'].mean(), 2)}")
print("Sample of jobs using more resources than predicted: ")
if mtype == 'mem':
print(df[df.pred_delta < 0].sample(min(5, num_bad_jobs)))
print("--------")
print("Total unused memory in GB (original): "
+ f"{round(df['old_delta'].sum()/1024)}")
print("Total unused memory in GB (predicted): "
+ f"{round(df[df.pred_delta >= 0]['pred_delta'].sum()/1024)}")
else:
print(df[df.pred_delta < 0].sample(min(5, num_bad_jobs)))
print("--------")
print("Total unused time in years (original): "
+ f"{round(df['time_delta'].sum()/31536000, 4)}")
print("Total unused time in years (predicted): "
+ f"{round(df[df.pred_delta >= 0]['pred_delta'].sum()/31536000, 4)}")
for mtype in ['mem', 'time']:
if options[mtype]['explore_model']:
print(f"Total loss info for {full_names[mtype]}:")
total_loss_info(mtype)
print('------------------')
| notebooks/test-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # SVM with RBF Kernel and Feature Preprocessing
#
# *<NAME>, April 28th 2021*
# +
# Importing our libraries
import pandas as pd
import altair as alt
import numpy as np
from sklearn.tree import DecisionTreeClassifier
from sklearn.dummy import DummyClassifier, DummyRegressor
from sklearn.neighbors import KNeighborsClassifier, KNeighborsRegressor
from sklearn.model_selection import cross_validate, train_test_split
import sys
sys.path.append('code/')
from display_tree import display_tree
from plot_classifier import plot_classifier
import matplotlib.pyplot as plt
# Preprocessing and pipeline
from sklearn.impute import SimpleImputer
from sklearn.metrics.pairwise import euclidean_distances
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import OneHotEncoder, OrdinalEncoder, StandardScaler, MinMaxScaler
# -
# ## House Keeping
# - Assignment due today at 11:59pm!
# - Course feedback!
# - Assignment - things I should know?
# - Assignment2 - before or after the weekend?
# - Polls coming Monday!
# - I hear you don't like breakout rooms, let's try this lecture without them!
# - Per the announcement Monday, download the data for this lecture [here](https://www.kaggle.com/harrywang/housing) and include it in your `data` folder that resides in `lectures`.
# ## Lecture Learning Objectives
#
# - Identify when to implement feature transformations such as imputation and scaling.
# - Describe the difference between normalizing and standardizing and be able to use scikit-learn's `MinMaxScaler()` and `StandardScaler()` to pre-process numeric features.
# - Apply `sklearn.pipeline.Pipeline` to build a machine learning pipeline.
# - Use `sklearn` for applying numerical feature transformations to the data.
# - Discuss the golden rule in the context of feature transformations.
# ## Five Minute Recap/ Lightning Questions
#
# - When using a Dummy Regressor what value does the model predict for unseen data?
# - When using a Dummy Classifier (the one we examined in lecture) what class does the model predict for unseen data?
# - What is the name of the distance metric used in the $k$-nn model we looked at?
# - If a dataset has 14 features and 1 target column, how many dimensions will the feature vector be?
# - What is the hyperparameter name of the $k$-nn classifier we looked at last lecture?
# ### Some lingering questions
#
# - How does a $k$-nn Regressor work?
# - Are we ready to do machine learning on real-world datasets?
# - We've looked at data with numeric features but what do we do if we have features with categories or string values?
# - What happens if we are missing data in our features?
# - Is there a cleaner way to do all the steps we need to do?
# ## Regression with $k$-NN
#
# In $k$-nearest neighbour regression, we take the average of $k$-nearest neighbours instead of the majority vote.
#
# Let's look at an example.
#
# Here we are creating some synthetic data with fifty examples and only one feature.
#
# We only have one feature of `length` and our goal is to predict `weight`.
#
# Regression plots more naturally in 1D, classification in 2D, but of course we can do either for any $d$
#
# Right now, do not worry about the code and only focus on data and our model.
np.random.seed(0)
n = 50
X_1 = np.linspace(0,2,n)+np.random.randn(n)*0.01
X = pd.DataFrame(X_1[:,None], columns=['length'])
X.head()
y = abs(np.random.randn(n,1))*2 + X_1[:,None]*5
y = pd.DataFrame(y, columns=['weight'])
y.head()
snake_X_train, snake_X_test, snake_y_train, snake_y_test = train_test_split(X, y, test_size=0.2, random_state=123)
# Now let's visualize our training data.
# +
source = pd.concat([snake_X_train, snake_y_train], axis=1)
scatter = alt.Chart(source, width=500, height=300).mark_point(filled=True, color='green').encode(
alt.X('length:Q'),
alt.Y('weight:Q'))
scatter
# -
# Now let's try the $k$-nearest neighbours regressor on this data.
#
# Then we create our `KNeighborsRegressor` object with `n_neighbors=1` so we are only considering 1 neighbour and with `uniform` weights.
# +
from sklearn.neighbors import KNeighborsRegressor
knnr_1 = KNeighborsRegressor(n_neighbors=1, weights="uniform")
knnr_1.fit(snake_X_train,snake_y_train);
predicted = knnr_1.predict(snake_X_train)
predicted
# -
# If we scored over regressors we get this perfect score of one since we have `n_neighbors=1` we are likely to overfit.
knnr_1.score(snake_X_train, snake_y_train)
# Plotting this we can see our model is trying to get every example correct since n_neighbors=1. (the mean of 1 point is just going to be the point value)
plt.figure(figsize=(8, 5))
grid = np.linspace(np.min(snake_X_train), np.max(snake_X_train), 1000)
plt.plot(grid, knnr_1.predict(grid), color='orange', linewidth=1)
plt.plot(snake_X_train, snake_y_train, ".r", markersize=10, color='green')
plt.xticks(fontsize= 14);
plt.yticks(fontsize= 14);
plt.xlabel("length",fontsize= 14)
plt.ylabel("weight",fontsize= 14);
# What happens when we use `n_neighbors=10`?
knnr_10 = KNeighborsRegressor(n_neighbors=10, weights="uniform")
knnr_10.fit(snake_X_train, snake_y_train)
knnr_10.score(snake_X_train, snake_y_train)
# Now we can see we are getting a lower score over the training set. Our score decreased from 1.0 when to had `n_neighbors=1` to now having a score of 0.925.
#
# When we plot our model, we can see that it no longer is trying to get every example correct.
plt.figure(figsize=(8, 5))
plt.plot(grid, knnr_10.predict(grid), color='orange', linewidth=1)
plt.plot(snake_X_train, snake_y_train, ".r", markersize=10, color='green')
plt.xticks(fontsize= 16);
plt.yticks(fontsize= 16);
plt.xlabel("length",fontsize= 16)
plt.ylabel("weight",fontsize= 16);
# ## Pros and Cons of 𝑘 -Nearest Neighbours
#
#
# ### Pros:
#
# - Easy to understand, interpret.
# - Simply hyperparameter $k$ (`n_neighbors`) controlling the fundamental tradeoff.
# - Can learn very complex functions given enough data.
# - Lazy learning: Takes no time to `fit`
#
# <br>
#
# ### Cons:
#
# - Can potentially be VERY slow during prediction time.
# - Often not that great test accuracy compared to the modern approaches.
# - Need to scale your features. We'll be looking into this in an upcoming lecture (lecture 4 I think?).
#
# ## Let's Practice
#
# $$ X = \begin{bmatrix}5 & 2\\4 & 3\\ 2 & 2\\ 10 & 10\\ 9 & -1\\ 9& 9\end{bmatrix}, \quad y = \begin{bmatrix}0\\0\\1\\1\\1\\2\end{bmatrix}.$$
#
# If $k=3$, what would you predict for $x=\begin{bmatrix} 0\\0\end{bmatrix}$ if we were doing regression rather than classification?
#
# ```{admonition} Solutions!
# :class: dropdown
#
# 1. 1/3 ($\frac{0 + 0 + 0}{3}$)
# ```
# ## Support Vector Machines (SVMs) with RBF Kernel
#
# Another popular similarity-based algorithm is Support Vector Machines (SVM).
#
# SVMs use a different similarity metric which is called a “kernel” in "SVM land".
#
# We are going to concentrate on the specific kernel called Radial Basis Functions (RBFs).
#
# Back to the good ol' Canadian and USA cities data.
cities_df = pd.read_csv("data/canada_usa_cities.csv")
cities_train_df, cities_test_df = train_test_split(cities_df, test_size=0.2, random_state=123)
cities_train_df.head()
# +
cities_X_train = cities_train_df.drop(columns=['country'])
cities_y_train = cities_train_df['country']
cities_X_test = cities_test_df.drop(columns=['country'])
cities_y_test = cities_test_df['country']
cities_X_train.head()
# -
cities_y_train.head()
# Unlike with $k$-nn, we are not going into detail about how support vector machine classifiers or regressor works but more so on how to use it with `sklearn`.
#
# We can use our training feature table ($X$) and target ($y$) values by using this new SVM model with (RBF) but with the old set up with `.fit()` and `.score()` that we have seen time and time again.
# We import the `SVC` tool from the `sklearn.svm` library (The "C" in SVC represents *Classifier*.
#
# To import the regressor we import `SVR` - R for *Regressor*)
from sklearn.svm import SVC
from sklearn.svm import SVR
# We can cross-validate and score exactly how we saw before.
#
# (For now, ignore `gamma=0.01` we are addressing it coming up)
svm = SVC(gamma=0.01)
scores = cross_validate(svm, cities_X_train, cities_y_train, return_train_score=True)
pd.DataFrame(scores)
svm_cv_score = scores['test_score'].mean()
svm_cv_score
# The biggest thing to know about support vector machines is that superficially, support vector machines are very similar to 𝑘-Nearest Neighbours.
#
# You can think of SVM with RBF kernel as a "smoothed" version of the $k$-Nearest Neighbours.
# +
svm.fit(cities_X_train, cities_y_train);
kn5_model = KNeighborsClassifier(n_neighbors=5)
kn5_model.fit(cities_X_train, cities_y_train);
plt.figure(figsize=(16, 8))
plt.subplot(1, 2, 1)
plt.title("SVC")
plot_classifier(cities_X_train, cities_y_train, svm, ax=plt.gca())
plt.subplot(1, 2, 2)
plt.title("KNN with k = 5")
plot_classifier(cities_X_train, cities_y_train, kn5_model, ax=plt.gca());
# -
# An observation is classified as a positive class if on average it looks more like positive examples. An observation is classified as a negative class if on average it looks more like negative examples.
#
# The primary difference between 𝑘-NNs and SVMs is that:
#
# - Unlike $k$-NNs, SVMs only remember the key examples (Those examples are called **support vectors**).
# - When it comes to predicting a query point, we only consider the key examples from the data and only calculate the distance to these key examples. This makes it more efficient than 𝑘-NN.
# ### Hyperparameters of SVM
#
# There are 2 main hyperparameters for support vector machines with an RBF kernel;
#
# - `gamma`
# - `C`
#
# (told you we were coming back to it!)
#
# We are not equipped to understand the meaning of these parameters at this point but you are expected to describe their relationship to the fundamental tradeoff.
#
# (In short, `C` is the penalty the model accepts for wrongly classified examples, and `gamma` is the curvature (see [here](https://towardsdatascience.com/hyperparameter-tuning-for-support-vector-machines-c-and-gamma-parameters-6a5097416167) for more)
#
# See [`scikit-learn`'s explanation of RBF SVM parameters](https://scikit-learn.org/stable/auto_examples/svm/plot_rbf_parameters.html)
# #### `gamma` and the fundamental trade-off
#
# `gamma` controls the complexity of a model, just like other hyperparameters we've seen.
#
# - higher gamma, higher the complexity.
# - lower gamma, lower the complexity.
plt.figure(figsize=(16, 4))
for i in range(4):
plt.subplot(1, 4, i + 1)
gamma = 10.0 ** (i - 3)
rbf_svm = SVC(gamma=gamma)
rbf_svm.fit(cities_X_train, cities_y_train)
plt.title("gamma = %s" % gamma);
plot_classifier(cities_X_train, cities_y_train, rbf_svm, ax=plt.gca(), show_data=False)
# #### `C` and the fundamental trade-off
#
# `C` also controls the complexity of a model and in turn the fundamental tradeoff.
#
# - higher `C` values, higher the complexity.
# - lower `C` values, lower the complexity.
plt.figure(figsize=(16, 4))
for i in range(4):
plt.subplot(1, 4, i + 1)
C = 10.0 ** (i - 1)
rbf_svm = SVC(C=C, gamma=0.01)
rbf_svm.fit(cities_X_train, cities_y_train)
plt.title("C = %s" % C);
plot_classifier(cities_X_train, cities_y_train, rbf_svm, ax=plt.gca(), show_data=False)
# Obtaining optimal validation scores requires a hyperparameter search between both `gamma` and `C` to balance the fundamental trade-off.
# We will learn how to search over multiple hyperparameters at a time in lecture 5.
# ## Let's Practice
#
# **True or False**
#
# 1\.In Scikit Learn’s SVC classifier, large values of gamma tend to result in higher training scores but probably lower validation scores.
# 2\.If we increase both `gamma` and `C`, we can't be certain if the model becomes more complex or less complex.
#
# ```{admonition} Solutions!
# :class: dropdown
#
# 1. True
# 2. False
# ```
# ## Let's Practice - Coding
#
# Below is some starter code that creates your feature table and target column from the data from the `bball.csv` dataset (in the data folder).
# +
bball_df = pd.read_csv('data/bball.csv')
bball_df = bball_df[(bball_df['position'] =='G') | (bball_df['position'] =='F')]
# Define X and y
X = bball_df.loc[:, ['height', 'weight', 'salary']]
y = bball_df['position']
# -
# 1. Split the dataset into 4 objects: `X_train`, `X_test`, `y_train`, `y_test`. Make the test set 0.2 (or the train set 0.8) and make sure to use `random_state=7`.
# 2. Create an `SVM` model with `gamma` equal to 0.1 and `C` equal to 10.
# 3. Cross-validate using cross_validate() on the objects X_train and y_train specifying the model and making sure to use 5 fold cross-validation and `return_train_score=True`.
# 4. Calculate the mean training and cross-validation scores.
# 1. Split the dataset
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=7)
model = SVC(gamma=0.1, C=10)
# 3. Cross-validate
scores_df = pd.DataFrame(cross_validate(model,X_train,y_train, cv=5, return_train_score=True))
scores_df
# 4. Calculate the mean training and cross-validation scores.
scores_df.mean()
scores_df.mean()['test_score']
scores_df.mean()['train_score']
# ## Preprocessing
# ### The importance of Preprocessing - An Example of Why
#
# So far we have seen:
#
# - Models: Decision trees, 𝑘-NNs, SVMs with RBF kernel.
# - Fundamentals: Train-validation-test split, cross-validation, the fundamental tradeoff, the golden rule.
#
#
#
# Now ...
#
# **Preprocessing**: Transforming input data into a format a machine learning model can use and understand.
#
# #### Basketball dataset
#
# Let's take a look at the `bball.csv` dataset we just used in practice.
#
# - Let's look at the 3 feature columns `height`, `weight` and `salary`.
# - Let's see if these features can help predict the `position` basketball players is.
bball_df = pd.read_csv('data/bball.csv')
bball_df = bball_df[(bball_df['position'] =='G') | (bball_df['position'] =='F')]
X = bball_df[['weight', 'height', 'salary']]
y =bball_df["position"]
X_train, X_test, y_train, y_test =train_test_split(X, y, test_size=0.20, random_state=123)
X_train.head()
y_train.head()
# First, let's see what validations scores we get if we simply predict the most occurring target value in the dataset using the dummy classifier model we saw in the last lecture.
dummy = DummyClassifier(strategy="most_frequent")
scores = cross_validate(dummy, X_train, y_train, return_train_score=True)
print('Mean training score', scores['train_score'].mean().round(2))
print('Mean validation score', scores['test_score'].mean().round(2))
# Here we get a mean validation score for our 5 fold cross_validation (5 is the default) of 57%. Let's now see how much better a $k$-nn model does on the data. We saw that it doesn't do to well on SVM, let's see if there is a difference with $k$-nn.
knn = KNeighborsClassifier()
scores = cross_validate(knn, X_train, y_train, return_train_score=True)
print('Mean training score', scores['train_score'].mean().round(2))
print('Mean validation score', scores['test_score'].mean().round(2))
# Ok, not the score we were hoping for.
#
# We are getting a worse score than the dummy classifier. This can't be right..... and it isn't and we are going to explain why!
#
# Let's have a look at just 2 players.
#
# We can see the values in each column.
two_players = X_train.sample(2, random_state=42)
two_players
# - The values in the `weight` column are around 100.
# - The values in the `height` column are around 2.
# - The values in the `salary` column are much higher at around 2 million.
#
# Let’s now calculate the distance between the two players.
euclidean_distances(two_players)
# So the distance between the players is 117133.0018.
#
# What happens if we only consider the salary column?
euclidean_distances(two_players[["salary"]])
# It looks like it's almost the same distance!
#
# The distance is completely dominated by the `salary` column, the feature with the largest values and the `weight` and `height` columns are being ignored in the distance calculation.
#
# **Does it matter?**
#
# Yes! The scale is based on how data was collected.
#
# Features on a smaller scale can be highly informative and there is no good reason to ignore them.
# We want our model to be robust and not sensitive to the scale.
# **What about for decision trees? Did scale matter then?**
#
# No. In decision trees we ask questions on one feature at a time and so the nodes are created independently without considering others.
#
#
# We have to scale our columns before we use our $k$-nn algorithm (and many others) so they are all using a similar range of values!
#
# And you guessed it - Sklearn has tools called transformers for this.
#
# We'll be using `sklearn`'s [`StandardScaler`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html) for this example.
# We will talk about this type of preprocessing in more detail in a hot minute but for now, concentrate on the syntax.
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler() # Create feature transformer object, can accept hyperparameters like models can!
scaler.fit(X_train) # Fitting the transformer on the train split
X_train_scaled = scaler.transform(X_train) # Transforming the train split
X_test_scaled = scaler.transform(X_test) # Transforming the test split
# `sklearn` uses `fit` and `transform` paradigms for feature transformations. (In model building it was `fit` and `predict` or `score`)
#
# We `fit` the transformer on the train split and then `transform` the train split as well as the test split.
pd.DataFrame(X_train_scaled, columns=X_train.columns).head()
# Now if we look at our features they are all within the same scales as opposed to what it was before:
X_train.head()
# ### Sklearn's *predict* vs *transform*
#
# When we make models, we `fit` and `predict`(`score`) with the syntax:
#
# ```
# model.fit(X_train, y_train)
# X_train_predictions = model.predict(X_train)
# ```
#
# With preprocessing, we replace the `.predict()` step with a `.transform()` step. We can pass `y_train` in `fit` but it's usually ignored. It allows us to pass it just to be consistent with the usual usage of `sklearn`'s `fit` method.
#
# ```
# transformer.fit(X_train, [y_train])
# X_train_transformed = transformer.transform(X_train)
# ```
#
#
# We can also carry out fitting and transforming in one call using `.fit_transform()`, but we must be mindful to use it only on the train split and **NOT** on the test split.
#
# ```
# X_train_transformed = transformer.fit_transform(X_train)
# ```
# Let's scale our features for this basketball dataset and then compare the results with our original score without scaling.
knn_unscaled = KNeighborsClassifier()
knn_unscaled.fit(X_train, y_train);
print('Train score: ', (knn_unscaled.score(X_train, y_train).round(2)))
print('Test score: ', (knn_unscaled.score(X_test, y_test).round(2)))
knn_scaled = KNeighborsClassifier()
knn_scaled.fit(X_train_scaled, y_train);
print('Train score: ', (knn_scaled.score(X_train_scaled, y_train).round(2)))
print('Test score: ', (knn_scaled.score(X_test_scaled, y_test).round(2)))
# The scores with scaled data are now much better compared to the unscaled data in the case of 𝑘-NNs.
#
# We can see now that 𝑘-NN is doing better than the Dummy Classifier when we scaled our features.
#
# We are not carrying out cross-validation here for a reason that we'll look into soon.
#
# We are being a bit sloppy here by using the test set several times for teaching purposes.
#
# But when we build any ML models, we should only assess the test set once.
# ### Common preprocessing techniques
#
# Here are some commonly performed feature transformation techniques we will focus on in this lesson.
# - Imputation
# - Tackling missing values
# - Scaling
# - Scaling of numeric features
# ## Let's Practice
#
# 1\. Name a model that will still produce meaningful predictions with different scaled column values.
# 2\. Complete the following statement: Preprocessing is done ______.
#
# a) To the model but before training
# b) To the data before training the model
# c) To the model after training
# d) To the data after training the model
#
# 3\. `StandardScaler` is a type of what?
# 4\. What data splits does `StandardScaler` alter (Training, Testing, Validation, None, All)?
#
# **True or False**
#
# 5\. Columns with lower magnitudes compared to columns with higher magnitudes are less important when making predictions.
# 6\. A model less sensitive to the scale of the data makes it more robust.
#
# ```{admonition} Solutions!
# :class: dropdown
#
# 1. Decision Tree Algorithm
# 2. b) To the data before training the model
# 3. Transformer
# 4. All
# 5. False
# 6. True
#
# ```
# ## California housing data (A case study)
#
# For the next few examples of preprocessing, we are going to be using a dataset exploring the prices of homes in California to demonstrate feature transformation techniques. The data can be downloaded from this site [here](https://www.kaggle.com/harrywang/housing). Please make sure that you include it in your `data` folder that resides in `lectures`.
#
# This dataset is a modified version of the California Housing dataset available from [<NAME>'s University of Porto website](https://www.dcc.fc.up.pt/~ltorgo/Regression/cal_housing.html)
#
#
# The task is to predict median house values in California districts, given several features from these districts.
#
# +
housing_df = pd.read_csv("data/housing.csv")
train_df, test_df = train_test_split(housing_df, test_size=0.1, random_state=123)
train_df.head()
# -
# Some column values are mean/median but some are not.
#
# Before we use this data we need to do some **feature engineering**.
#
# That means we are going to transform our data into features that may be more meaningful for our prediction.
#
# Let's add some new features to the dataset which could help predict the target: `median_house_value`.
# +
train_df = train_df.assign(rooms_per_household = train_df["total_rooms"]/train_df["households"],
bedrooms_per_household = train_df["total_bedrooms"]/train_df["households"],
population_per_household = train_df["population"]/train_df["households"])
test_df = test_df.assign(rooms_per_household = test_df["total_rooms"]/test_df["households"],
bedrooms_per_household = test_df["total_bedrooms"]/test_df["households"],
population_per_household = test_df["population"]/test_df["households"])
train_df = train_df.drop(columns=['total_rooms', 'total_bedrooms', 'population'])
test_df = test_df.drop(columns=['total_rooms', 'total_bedrooms', 'population'])
train_df.head()
# -
# ### When is it OK to do things before splitting?
#
# - Here it would have been OK to add new features before splitting because we are not using any global information in the data but only looking at one row at a time.
# - But just to be safe and to avoid accidentally breaking the golden rule, it's better to do it after splitting.
# ## Preprocessing: Imputation
#
# Imputation is handling missing values in our data so let's explore this a little.
#
# We can `.info()` we can we all the different column dtypes and also all the number of null values.
train_df.info()
# We see that we have all columns with dtype `float64` except for `ocean_proximity` which appears categorical.
#
# We also notice that the `bedrooms_per_household` column appears to have some `Non-Null` rows.
train_df["bedrooms_per_household"].isnull().sum()
# Knowing this information let's build a model.
#
# When we create our feature table and target objects, we are going to drop the categorical variable `ocean_proximity`. Currently, we don't know how to build models with categorical data, but we will shortly. We will return to this column soon.
# +
X_train = train_df.drop(columns=["median_house_value", "ocean_proximity"])
y_train = train_df["median_house_value"]
X_test = test_df.drop(columns=["median_house_value", "ocean_proximity"])
y_test = test_df["median_house_value"]
knn = KNeighborsRegressor()
# -
# What happens when we try to fit our model with this data?
knn.fit(X_train, y_train)
# > `Input contains NaN, infinity or a value too large for dtype('float64').`
#
# The classifier can't deal with missing values (NaNs).
#
# How can we deal with this problem?
# ### Why we don't drop the rows
#
# We could drop any rows that are missing information but that's problematic too.
#
# Then we would need to do the same in our test set.
#
# And what happens if we get missing values in our deployment data? what then?
#
# Furthermore, what if the missing values don't occur at random and we're systematically dropping certain data?
# Perhaps a certain type of house contributes to more missing values.
#
# Dropping the rows is not a great solution, especially if there's a lot of missing values.
X_train.shape
# +
X_train_no_nan = X_train.dropna()
y_train_no_nan = y_train.dropna()
X_train_no_nan.shape
# -
# ### Why we don't drop the column
#
# If we drop the column instead of the rows, we are throwing away, in this case, 18391 values just because we don't have 185 missing values out of a total of 18567.
#
# We are throwing away 99% of the column’s data because we are missing 1%.
#
# But perhaps if we were missing 99.9% of the column values, for example, it would make more sense to drop the column.
#
X_train.shape
# +
X_train_no_col = X_train.dropna(axis=1)
X_train_no_col.shape
# -
# ### Why we use imputation
#
# With **Imputation**, we invent values for the missing data.
#
# Using `sklearn`'s **transformer** `SimpleImputer`, we can impute the `NaN` values in the data with some value.
from sklearn.impute import SimpleImputer
# We can impute missing values in:
#
# - **Categorical columns**:
# - with the most frequent value
# - with a constant of our choosing.
# - **Numeric columns**:
# - with the mean of the column
# - with the median of the column
# - or a constant of our choosing.
# If I sort the values by `bedrooms_per_household` and look at the end of the dataframe, we can see our missing values in the `bedrooms_per_household` column.
#
# Pay close attention to index 7763 since we are going to look at this row after imputation.
X_train.sort_values('bedrooms_per_household').tail(10)
# Using the same `fit` and `transform` syntax we saw earlier for transformers, we can impute the `NaN` values.
#
# Here we specify `strategy="median"` which replaces all the missing values with the column median.
#
# We fit on the training data and transform it on the train and test splits.
#
imputer = SimpleImputer(strategy="median")
imputer.fit(X_train);
X_train_imp = imputer.transform(X_train)
X_test_imp = imputer.transform(X_test)
X_train_imp
# Ok, the output of this isn't a dataframe but a NumPy array!
#
# I can do a bit of wrangling here to take a look at this new array with our previous column labels and as a dataframe.
#
# If I search for our index 7763 which previously contained a `NaN` value, we can see that now I have the median value for the `bedrooms_per_household` column from the `X_train` dataframe.
X_train_imp_df = pd.DataFrame(X_train_imp, columns = X_train.columns, index = X_train.index)
X_train_imp_df.loc[[7763]]
X_train['bedrooms_per_household'].median()
X_train.loc[[7763]]
# Now when we try and fit our model using `X_train_imp`, it works!
knn = KNeighborsRegressor();
knn.fit(X_train_imp, y_train)
knn.score(X_train_imp, y_train)
# ## Preprocessing: Scaling
#
# So we've seen why scaling is important earlier but let's take a little bit of a closer look here.
# There are many ways to scale your data but we are going to look at 2 of them.
#
#
# 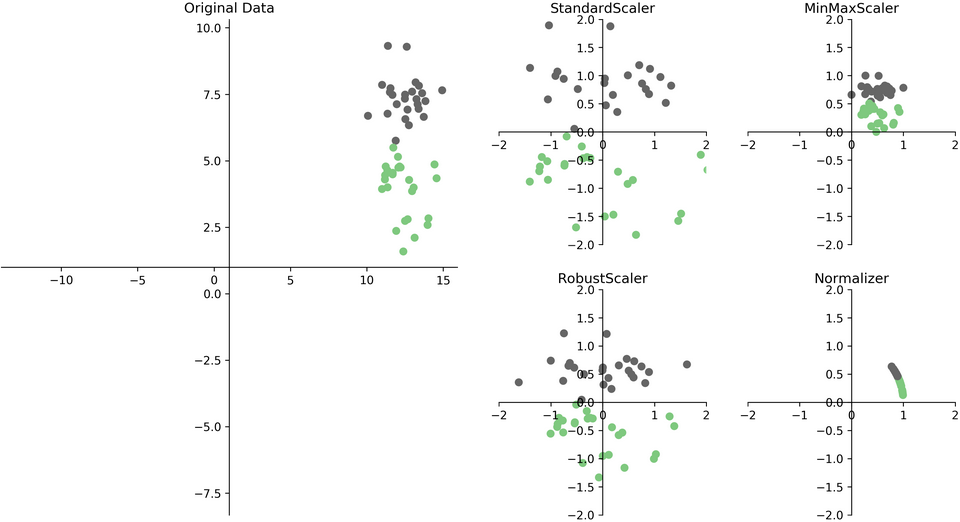
#
# | Approach | What it does | How to update $X$ (but see below!) | sklearn implementation |
# |---------|------------|-----------------------|----------------|
# | normalization | sets range to $[0,1]$ | `X -= np.min(X,axis=0)`<br>`X /= np.max(X,axis=0)` | [`MinMaxScaler()`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.MinMaxScaler.html)
# | standardization | sets sample mean to $0$, s.d. to $1$ | `X -= np.mean(X,axis=0)`<br>`X /= np.std(X,axis=0)` | [`StandardScaler()`](https://scikit-learn.org/stable/modules/generated/sklearn.preprocessing.StandardScaler.html#sklearn.preprocessing.StandardScaler) |
#
# For more resources and articles on this, see [here](http://www.dataminingblog.com/standardization-vs-normalization/) and [here](https://medium.com/@rrfd/standardize-or-normalize-examples-in-python-e3f174b65dfc).
# Let's see what happens when we use each of them.
from sklearn.preprocessing import MinMaxScaler, StandardScaler
# First, let's see how standardization is done first.
scaler = StandardScaler()
X_train_scaled_std = scaler.fit_transform(X_train_imp)
X_test_scaled_std = scaler.transform(X_test_imp)
pd.DataFrame(X_train_scaled_std, columns=X_train.columns, index=X_train.index).head()
# Here, any negative values represent values that are lower than the calculated feature mean and anything positive and greater than 0 are values greater than the original column mean.
knn = KNeighborsRegressor()
knn.fit(X_train_imp, y_train);
print('Unscaled training score :', knn.score(X_train_imp, y_train).round(3))
knn = KNeighborsRegressor()
knn.fit(X_train_scaled_std, y_train)
print('Scaled training score :',knn.score(X_train_scaled_std, y_train))
scaler = MinMaxScaler()
X_train_scaled_norm = scaler.fit_transform(X_train_imp)
X_test_scaled_norm = scaler.transform(X_test_imp)
pd.DataFrame(X_train_scaled_norm, columns=X_train.columns, index=X_train.index).head()
# Looking at the data after normalizing it, we see this time there are no negative values and they all are between 0 and 1.
#
# And the score now?
knn = KNeighborsRegressor()
knn.fit(X_train_scaled_norm, y_train)
print('Scaled training score :',knn.score(X_train_scaled_norm, y_train))
# - Big difference in the KNN training performance after scaling the data.
# - But we saw last week that the training score doesn't tell us much. We should look at the cross-validation score.
#
# So let's see how we can do this but first.... let's practice!
# ## Let's Practice
#
# 1\. When/Why do we need to impute our data?
# 2\. If we have `NaN` values in our data, can we simply drop the column missing the data?
# 3\. Which scaling method will never produce negative values?
# 4\. Which scaling method will never produce values greater than 1?
# 5\. Which scaling method will produce values where the range depends on the values in the data?
#
# **True or False**
#
# 6\. `SimpleImputer` is a type of transformer.
# 7\. Scaling is a form of transformation.
# 8\. We can use `SimpleImputer` to impute values that are missing from numerical and categorical columns.
# ```{admonition} Solutions!
# :class: dropdown
#
# 1. When we have missing data so that sklearn doesn't give an error.
# 2. No but we can if the majority of the values are missing from the column.
# 3. Normalization (`MinMaxScaler`)
# 4. Normalization (`MinMaxScaler`)
# 5. Standardization (`StandardScaler`)
# 6. True
# 7. True
# 8. True
# ```
# ## Feature transformations and the golden rule
#
# How to carry out cross-validation?
#
# - Last week we saw that cross-validation is a better way to get a realistic assessment of the model.
# - Let's try cross-validation with transformed data.
knn = KNeighborsRegressor()
scores = cross_validate(knn, X_train_scaled_std, y_train, return_train_score=True)
pd.DataFrame(scores)
# - Do you see any problem here?
#
# We are using our `X_train_scaled` in our `cross_validate()` function which already has all our preprocessing done.
#
# <img src='imgs/cross-validation.png' width="80%">
# That means that our validation set information is being used to calculate the mean and standard deviation (or min and max values for `MinMaxScaler`) for our training split!
#
# We are allowing information from the validation set to **leak** into the training step.
#
# What was our golden rule of machine learning again? Oh yeah -> ***Our test data should not influence our training data***.
#
# This applies also to our validation data and that it also should not influence our training data.
#
# With imputation and scaling, we are scaling and imputing values based on all the information in the data meaning the training data AND the validation data and so we are not adhering to the golden rule anymore.
#
# Every row in our `x_train_scaled` has now been influenced in a minor way by every other row in `x_train_scaled`.
#
# With scaling every row has been transformed based on all the data before splitting between training and validation.
#
# We need to take care that we are keeping our validation data truly as unseen data.
#
# Before we look at the right approach to this, let's look at the **WRONG** approaches.
# ### Bad methodology 1: Scaling the data separately
#
# We make our transformer, we fit it on the training data and then transform the training data.
#
# Then, we make a second transformer, fit it on the test data and then transform our test data.
# +
scaler = StandardScaler();
scaler.fit(X_train_imp);
X_train_scaled = scaler.transform(X_train_imp)
# Creating a separate object for scaling test data - Not a good idea.
scaler = StandardScaler();
scaler.fit(X_test_imp); # Calling fit on the test data - Yikes!
X_test_scaled = scaler.transform(X_test_imp) # Transforming the test data using the scaler fit on test data ... Bad!
knn = KNeighborsRegressor()
knn.fit(X_train_scaled, y_train);
print("Training score: ", knn.score(X_train_scaled, y_train).round(2))
print("Test score: ", knn.score(X_test_scaled, y_test).round(2))
# -
# This is bad because we are using two different StandardScaler objects but we want to apply the same transformation on the training and test splits.
#
# The test data will have different values than the training data producing a different transformation than the training data.
#
# We should never fit on test data, whether it’s to build a model or with a transforming, test data should never be exposed to the fit function.
# ### Bad methodology 2: Scaling the data together
#
# The next mistake is when we scale the data together. So instead of splitting our data, we are combining our training and testing and scaling it together.
X_train_imp.shape, X_test_imp.shape
# join the train and test sets back together
XX = np.vstack((X_train_imp, X_test_imp))## Don't do it!
XX.shape
scaler = StandardScaler()
scaler.fit(XX)
XX_scaled = scaler.transform(XX)
XX_train = XX_scaled[:18576]
XX_test = XX_scaled[18576:]
knn = KNeighborsRegressor()
knn.fit(XX_train, y_train);
print('Train score: ', (knn.score(XX_train, y_train).round(2))) # Misleading score
print('Test score: ', (knn.score(XX_test, y_test).round(2))) # Misleading score
# Here we are scaling the train and test splits together.
#
# The golden rule says that the test data shouldn’t influence the training in any way.
#
# Information from the test split is now affecting the mean for standardization!
#
# This is a clear violation of the golden rule.
#
# So what do we do? Enter ....
# ## Pipelines
#
# [Scikit-learn Pipeline](https://scikit-learn.org/stable/modules/generated/sklearn.pipeline.Pipeline.html) is here to save the day!
#
# A **pipeline** is a sklearn function that contains a sequence of steps.
#
# Essentially we give it all the actions we want to do with our data such as transformers and models and the pipeline will execute them in steps.
from sklearn.pipeline import Pipeline
# Let's combine the preprocessing and model with pipeline.
#
# we will instruct the pipeline to:
#
# 1. Do imputation using `SimpleImputer()` using a strategy of “median”
# 2. Scale our data using `StandardScaler`
# 3. Build a `KNeighborsRegressor`.
#
# (The last step should be a model and earlier steps should be transformers)
#
# Note: The input for `Pipeline` is a list containing tuples (one for each step).
pipe = Pipeline([
("imputer", SimpleImputer(strategy="median")),
("scaler", StandardScaler()),
("reg", KNeighborsRegressor())
])
pipe.fit(X_train, y_train)
# - Note that we are passing `X_train` and **NOT** the imputed or scaled data here.
#
# When we call `fit` the pipeline is carrying out the following steps:
#
# - Fit `SimpleImputer` on `X_train`.
# - Transform `X_train` using the fit `SimpleImputer` to create `X_train_imp`.
# - Fit `StandardScaler` on `X_train_imp`.
# - Transform `X_train_imp` using the fit `StandardScaler` to create `X_train_imp_scaled`.
# - Fit the model (`KNeighborsRegressor` in our case) on `X_train_imp_scaled`.
#
pipe.predict(X_train)
# When we call `predict` on our data, the following steps are carrying out:
#
# - Transform `X_train` using the fit `SimpleImputer` to create `X_train_imp`.
# - Transform `X_train_imp` using the fit `StandardScaler` to create `X_train_imp_scaled`.
# - Predict using the fit model (`KNeighborsRegressor` in our case) on `X_train_imp_scaled`.
#
# It is not fitting any of the data this time.
#
# <img src='https://amueller.github.io/COMS4995-s20/slides/aml-04-preprocessing/images/pipeline.png' width="50%">
#
# [Source](https://amueller.github.io/COMS4995-s20/slides/aml-04-preprocessing/#18)
# We can’t accidentally re-fit the preprocessor on the test data as we did before.
#
# It automatically makes sure the same transformations are applied to train and test.
#
# Now when we do cross-validation on the pipeline the transformers and the model are refit on each fold.
#
# The pipeline applies the `fit_transform` on the train portion of the data and only `transform` on the validation portion in **each fold**.
#
# This is how to avoid the Golden Rule violation!
scores_processed = cross_validate(pipe, X_train, y_train, return_train_score=True)
pd.DataFrame(scores_processed)
pd.DataFrame(scores_processed).mean()
dummy = DummyRegressor(strategy="median")
scores = cross_validate(dummy, X_train, y_train, return_train_score=True)
pd.DataFrame(scores).mean()
# We can trust here now that the scores are not influenced but the training data and all our steps were done efficiently and easily too.
# ## Let's Practice
#
# 1\. Which of the following steps cannot be used in a pipeline?
#
# a) Scaling
# b) Model building
# c) Imputation
# d) Data Splitting
#
# 2\. Why can't we fit and transform the training and test data together?
#
# **True or False**
#
# 3\. We have to be careful of the order we put each transformation and model in a pipeline.
# 4\. Pipelines will fit and transform on both the training and validation folds during cross-validation.
# ```{admonition} Solutions!
# :class: dropdown
#
# 1. Data Splitting
# 2. It's violating the golden rule.
# 3. True
# 4. False
# ```
# ## Let's Practice - Coding
#
# Let's bring in the basketball dataset again.
# +
# Loading in the data
bball_df = pd.read_csv('data/bball.csv')
bball_df = bball_df[(bball_df['position'] =='G') | (bball_df['position'] =='F')]
# Define X and y
X = bball_df.loc[:, ['height', 'weight', 'salary']]
y = bball_df['position']
# Split the dataset
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.2, random_state=7)
# -
# Build a pipeline named `bb_pipe` that:
# 1. Imputes using "median" as a strategy,
# 2. scale using `StandardScaler`
# 3. builds a `KNeighborsClassifier`.
#
#
# Next, do 5 fold cross-validation on the pipeline using `X_train` and `y_train` and save the results in a dataframe.
# Take the mean of each column and assess your model.
# ## What We've Learned Today<a id="9"></a>
#
# - How the $k$NN algorithm works for regression.
# - How to build an SVM with RBF kernel model.
# - How changing `gamma` and `C` hyperparameters affects the fundamental tradeoff.
# - How to imputer values when we are missing data.
# - Why it's important to scale our features.
# - How to scales our features.
# - How to build a pipeline that executes a number of steps without breaking the golden rule of ML.
#
| _build/html/_sources/lectures/lecture4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
import numpy as np
L = 1.6 # m (overall length)
B = 0.8 # m (overall width)
b = 1.6 / 7.0 # m (demihull width)
T = b / 2.0 # m (draft)
Cb = 0.397 # ul (block coefficient)
V = 3.0 # m/s (velocity)
# +
displacement_length_ratio = 6.27
displacement_per_hull = (L / displacement_length_ratio) ** 3
Fr = V / np.sqrt(9.81 * L)
displacement_total = 2 * displacement_per_hull
mass_displaced = displacement_total * 1000.0
length_demi_width_ratio = L / b
width_length_ratio = (B - b) / L
print Fr
print mass_displaced
# +
V_m = V * np.sqrt(1.6 / L)
Re_m = V_m * 1.6 / 1.14E-6
Re_s = V * L / 1.19E-6
C_FM = 0.075 / (np.log10(Re_m) - 2)**2
C_FS = 0.075 / (np.log10(Re_s) - 2)**2
print C_FM
# +
n = (0.41 + 0.51)/2.0
a = (0.518 + 0.426)/2.0
tau = a * displacement_length_ratio**n
Cr = 6.078 / 1000.0
# -
C_total = C_FS + tau*Cr
print C_total
R_total = 0.5 * 1000.0 * C_total * V**2 * 0.434*2
print R_total
R_total
P = R_total * V
print P
| notebooks/hull_drag/southampton_catamaran_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# default_exp utils
# %load_ext autoreload
# %autoreload 2
import os
os.environ["CUDA_VISIBLE_DEVICES"] = "-1"
# # Utils
#
# Utils functions
# +
# export
import os
import pickle
import re
from typing import Union
from inspect import getmembers
import numpy as np
import tensorflow as tf
import transformers
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.preprocessing import MultiLabelBinarizer
from transformers import AutoTokenizer, PreTrainedTokenizer
from bert_multitask_learning.special_tokens import TRAIN, EVAL, PREDICT
# -
# hide
from bert_multitask_learning.test_base import TestBase
import bert_multitask_learning
import shutil
import numpy as np
test_base = TestBase()
params = test_base.params
# export
def load_transformer_tokenizer(tokenizer_name: str, load_module_name=None):
"""some tokenizers cannot be loaded using AutoTokenizer.
this function served as a util function to catch that situation.
Args:
tokenizer_name (str): tokenizer name
"""
if load_module_name:
tok = getattr(transformers, load_module_name).from_pretrained(
tokenizer_name)
else:
tok = AutoTokenizer.from_pretrained(tokenizer_name)
return tok
load_transformer_tokenizer(
'voidful/albert_chinese_tiny', 'BertTokenizer')
# export
def load_transformer_config(config_name_or_dict, load_module_name=None):
"""Some models need specify loading module
Args:
config_name (str): module name
load_module_name (str, optional): loading module name. Defaults to None.
Returns:
config: config
"""
if load_module_name:
load_module = getattr(transformers, load_module_name)
else:
load_module = transformers.AutoConfig
if isinstance(config_name_or_dict, str):
config = load_module.from_pretrained(
config_name_or_dict, output_attentions=True, output_hidden_states=True)
elif isinstance(config_name_or_dict, dict):
config = load_module.from_dict(
config_name_or_dict, output_attentions=True, output_hidden_states=True)
else:
raise ValueError('config_name_or_dict should be str or dict')
return config
# load config with name
config = load_transformer_config(
'bert-base-chinese')
config_dict = config.to_dict()
# load config with dict
config = load_transformer_config(
config_dict, load_module_name='BertConfig')
# export
def load_transformer_model(model_name_or_config, load_module_name=None):
if load_module_name:
load_module = getattr(transformers, load_module_name)
else:
load_module = transformers.TFAutoModel
if isinstance(model_name_or_config, str):
try:
model = load_module.from_pretrained(
model_name_or_config, output_attentions=True, output_hidden_states=True)
except OSError:
model = load_module.from_pretrained(
model_name_or_config, from_pt=True, output_attentions=True, output_hidden_states=True)
else:
model = load_module.from_config(
model_name_or_config)
return model
# +
# load by name(load weights)
# this is a pt only model
model = load_transformer_model(
'voidful/albert_chinese_tiny')
# load by config (not load weights)
model = load_transformer_model(load_transformer_config(
'bert-base-chinese'))
# +
# export
class LabelEncoder(BaseEstimator, TransformerMixin):
def fit(self, y):
"""Fit label encoder
Parameters
----------
y : array-like of shape (n_samples,)
Target values.
Returns
-------
self : returns an instance of self.
"""
self.encode_dict = {}
self.decode_dict = {}
label_set = set(y)
label_set = sorted(list(label_set))
for l_ind, l in enumerate(label_set):
new_ind = l_ind
self.encode_dict[l] = new_ind
self.decode_dict[new_ind] = l
return self
def fit_transform(self, y):
"""Fit label encoder and return encoded labels
Parameters
----------
y : array-like of shape [n_samples]
Target values.
Returns
-------
y : array-like of shape [n_samples]
"""
self.fit(y)
y = self.transform(y)
return y
def transform(self, y):
"""Transform labels to normalized encoding.
Parameters
----------
y : array-like of shape [n_samples]
Target values.
Returns
-------
y : array-like of shape [n_samples]
"""
encode_y = []
for l in y:
encode_y.append(self.encode_dict[l])
return np.array(encode_y)
def inverse_transform(self, y):
"""Transform labels back to original encoding.
Parameters
----------
y : numpy array of shape [n_samples]
Target values.
Returns
-------
y : numpy array of shape [n_samples]
"""
decode_y = []
for l in y:
decode_y.append(self.decode_dict[l])
return np.array(decode_y)
def dump(self, path):
with open(path, 'wb') as f:
pickle.dump(self.decode_dict, f)
def load(self, path):
with open(path, 'rb') as f:
self.decode_dict = pickle.load(f)
self.encode_dict = {v: k for k, v in self.decode_dict.items()}
def create_path(path):
if not os.path.exists(path):
os.makedirs(path, exist_ok=True)
def get_or_make_label_encoder(params, problem: str, mode: str, label_list=None) -> Union[LabelEncoder, MultiLabelBinarizer, PreTrainedTokenizer]:
"""Function to unify ways to get or create label encoder for various
problem type.
cls: LabelEncoder
seq_tag: LabelEncoder
multi_cls: MultiLabelBinarizer
seq2seq_text: Tokenizer
Arguments:
problem {str} -- problem name
mode {mode} -- mode
Keyword Arguments:
label_list {list} -- label list to fit the encoder (default: {None})
Returns:
LabelEncoder -- label encoder
"""
problem_path = params.ckpt_dir
create_path(problem_path)
problem_type = params.problem_type[problem]
le_path = os.path.join(problem_path, '%s_label_encoder.pkl' % problem)
is_seq2seq_text = problem_type == 'seq2seq_text'
is_multi_cls = problem_type == 'multi_cls'
is_seq = problem_type == 'seq_tag'
is_pretrain = problem_type == 'pretrain'
is_masklm = problem_type == 'masklm'
if problem_type not in params.predefined_problem_type:
is_custom = True
else:
is_custom = False
if is_pretrain:
return None
if mode == TRAIN and not os.path.exists(le_path):
if is_custom:
get_or_make_custom_le_fn = params.get_or_make_label_encoder_fn_dict[
problem_type]
get_or_make_custom_le_fn(params, problem, mode, label_list)
if is_seq2seq_text:
label_encoder = load_transformer_tokenizer(
params.transformer_decoder_tokenizer_name, params.transformer_decoder_tokenizer_loading)
pickle.dump(label_encoder, open(le_path, 'wb'))
elif is_masklm:
label_encoder = load_transformer_tokenizer(
params.transformer_tokenizer_name, params.transformer_tokenizer_loading
)
pickle.dump(label_encoder, open(le_path, 'wb'))
elif is_multi_cls:
label_encoder = MultiLabelBinarizer()
label_encoder.fit(label_list)
pickle.dump(label_encoder, open(le_path, 'wb'))
else:
if isinstance(label_list[0], list):
label_list = [
item for sublist in label_list for item in sublist]
if is_seq:
label_list.append('[PAD]')
label_encoder = LabelEncoder()
label_encoder.fit(label_list)
label_encoder.dump(le_path)
else:
if is_custom:
get_or_make_custom_le_fn = params.get_or_make_label_encoder_fn_dict[
problem_type]
label_encoder = get_or_make_custom_le_fn(
params, problem, mode, label_list)
if is_seq2seq_text or is_multi_cls or is_masklm:
label_encoder = pickle.load(open(le_path, 'rb'))
else:
label_encoder = LabelEncoder()
label_encoder.load(le_path)
if is_custom:
if problem not in params.num_classes:
raise ValueError(
'Seems custom get or make label encoder fn dose not set num_classes to'
'params. Please specify num_classes. Example: params.num_classes[problem] = 100')
return label_encoder
if not is_seq2seq_text and not is_masklm:
if is_multi_cls:
params.num_classes[problem] = label_encoder.classes_.shape[0]
else:
params.num_classes[problem] = len(label_encoder.encode_dict)
else:
try:
params.num_classes[problem] = len(label_encoder.vocab)
except AttributeError:
# models like xlnet's vocab size can only be retrieved from config instead of tokenizer
params.num_classes[problem] = params.bert_decoder_config.vocab_size
return label_encoder
# -
# +
le_train = get_or_make_label_encoder(
params=params, problem='weibo_fake_ner', mode=bert_multitask_learning.TRAIN, label_list=[['a', 'b'], ['c']]
)
# seq_tag will add [PAD]
assert len(le_train.encode_dict) == 4
le_predict = get_or_make_label_encoder(
params=params, problem='weibo_fake_ner', mode=bert_multitask_learning.PREDICT)
assert le_predict.encode_dict==le_train.encode_dict
# list train
le_train = get_or_make_label_encoder(
params=params, problem='weibo_fake_cls', mode=bert_multitask_learning.TRAIN, label_list=['a', 'b', 'c']
)
# seq_tag will add [PAD]
assert len(le_train.encode_dict) == 3
le_predict = get_or_make_label_encoder(
params=params, problem='weibo_fake_cls', mode=bert_multitask_learning.PREDICT)
assert le_predict.encode_dict==le_train.encode_dict
# text
le_train = get_or_make_label_encoder(
params=params, problem='weibo_masklm', mode=bert_multitask_learning.TRAIN)
assert isinstance(le_train, transformers.PreTrainedTokenizer)
le_predict = get_or_make_label_encoder(
params=params, problem='weibo_masklm', mode=bert_multitask_learning.PREDICT)
assert isinstance(le_predict, transformers.PreTrainedTokenizer)
# +
# export
def cluster_alphnum(text: str) -> list:
"""Simple funtions to aggregate eng and number
Arguments:
text {str} -- input text
Returns:
list -- list of string with chinese char or eng word as element
"""
return_list = []
last_is_alphnum = False
for char in text:
is_alphnum = bool(re.match('^[a-zA-Z0-9\[]+$', char))
is_right_brack = char == ']'
if is_alphnum:
if last_is_alphnum:
return_list[-1] += char
else:
return_list.append(char)
last_is_alphnum = True
elif is_right_brack:
if return_list:
return_list[-1] += char
else:
return_list.append(char)
last_is_alphnum = False
else:
return_list.append(char)
last_is_alphnum = False
return return_list
def filter_empty(input_list, target_list):
"""Filter empty inputs or targets
Arguments:
input_list {list} -- input list
target_list {list} -- target list
Returns:
input_list, target_list -- data after filter
"""
return_input, return_target = [], []
for inp, tar in zip(input_list, target_list):
if inp and tar:
return_input.append(inp)
return_target.append(tar)
return return_input, return_target
# +
# export
def infer_shape_and_type_from_dict(inp_dict: dict, fix_dim_for_high_rank_tensor=True):
shape_dict = {}
type_dict = {}
for feature_name, feature in inp_dict.items():
if type(feature) is list:
feature = np.array(feature)
if type(feature) is np.ndarray:
if np.issubdtype(feature.dtype, np.integer):
type_dict[feature_name] = tf.int32
elif np.issubdtype(feature.dtype, np.floating):
type_dict[feature_name] = tf.float32
# this seems not a good idea
if len(feature.shape) > 1 and fix_dim_for_high_rank_tensor:
shape_dict[feature_name] = [
None] + list(feature.shape[1:])
else:
shape_dict[feature_name] = [
None for _ in feature.shape]
elif np.issubdtype(type(feature), np.floating):
type_dict[feature_name] = tf.float32
shape_dict[feature_name] = []
elif np.issubdtype(type(feature), np.integer):
type_dict[feature_name] = tf.int32
shape_dict[feature_name] = []
else:
if isinstance(feature, str):
feature = feature.encode('utf8')
type_dict[feature_name] = tf.string
shape_dict[feature_name] = []
return shape_dict, type_dict
# -
# dose not support nested dict
test_dict = {
'test1': np.random.uniform(size=(64, 32)),
'test2': np.array([1, 2, 3], dtype='int32'),
'test5': 5
}
desc_dict = infer_shape_and_type_from_dict(
test_dict)
assert desc_dict == ({'test1': [None, 32], 'test2': [None], 'test5': []}, {
'test1': tf.float32, 'test2': tf.int32, 'test5': tf.int32})
# +
# export
def get_transformer_main_model(model, key='embeddings'):
"""Function to extrac model name from huggingface transformer models.
Args:
model (Model): Huggingface transformers model
key (str, optional): Key to identify model. Defaults to 'embeddings'.
Returns:
model
"""
for attr_name, attr in getmembers(model):
if attr_name == key:
return model
if hasattr(attr, key):
return attr
# -
model = load_transformer_model(
'voidful/albert_chinese_tiny')
main_model = get_transformer_main_model(model)
isinstance(main_model, transformers.TFAlbertMainLayer)
# +
# export
def get_embedding_table_from_model(model):
base_model = get_transformer_main_model(model)
return base_model.embeddings.word_embeddings
# -
embedding = get_embedding_table_from_model(
model)
assert embedding.shape == (21128, 128)
# +
# export
def get_shape_list(tensor, expected_rank=None, name=None):
"""Returns a list of the shape of tensor, preferring static dimensions.
Args:
tensor: A tf.Tensor object to find the shape of.
expected_rank: (optional) int. The expected rank of `tensor`. If this is
specified and the `tensor` has a different rank, and exception will be
thrown.
name: Optional name of the tensor for the error message.
Returns:
A list of dimensions of the shape of tensor. All static dimensions will
be returned as python integers, and dynamic dimensions will be returned
as tf.Tensor scalars.
"""
shape = tensor.shape.as_list()
non_static_indexes = []
for (index, dim) in enumerate(shape):
if dim is None:
non_static_indexes.append(index)
if not non_static_indexes:
return shape
dyn_shape = tf.shape(input=tensor)
for index in non_static_indexes:
shape[index] = dyn_shape[index]
return shape
def gather_indexes(sequence_tensor, positions):
"""Gathers the vectors at the specific positions over a minibatch."""
sequence_shape = get_shape_list(sequence_tensor, expected_rank=3)
batch_size = sequence_shape[0]
seq_length = sequence_shape[1]
width = sequence_shape[2]
flat_offsets = tf.reshape(
tf.range(0, batch_size, dtype=tf.int32) * seq_length, [-1, 1])
flat_offsets = tf.cast(flat_offsets, tf.int64)
flat_positions = tf.reshape(positions + flat_offsets, [-1])
flat_sequence_tensor = tf.reshape(sequence_tensor,
[batch_size * seq_length, width])
# https://github.com/tensorflow/tensorflow/issues/36236
output_tensor = tf.gather(flat_sequence_tensor*1, flat_positions)
return output_tensor
# -
| source_nbs/01_utils.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# import libraries for data analysis
import pandas as pd
import researchpy as rp
df = pd.read_csv('../data/cleaned_school_distance_learning.csv')
df.head()
df.shape
# ## Demographic summary
# Summary of numeric data
rp.summary_cont(df['age'])
rp.codebook(df['age'])
rp.summary_cat(df[['gender', 'class']])
# ## Basic information regrding online class
class_per_week = df['how_many_days_did_you_have_classes_in_a_week'].astype('category')
rp.summary_cat(class_per_week)
class_per_day = df['how_many_classes_did_you_have_per_day'].astype('category')
rp.summary_cat(class_per_day)
rp.summary_cat(df[['did_you_attend_any_online_class_before_covid-19_pandemic',
'have_you_attended_online_classes_regularly',
'what_was_the_duration_of_each_class',
]])
# Device used to conduct online class
df['what_type_of_device_did_you_use_in_your_online_class']
# Convert response into binary
device = df['what_type_of_device_did_you_use_in_your_online_class'].str.get_dummies(sep=', ')
device.head()
# replace 1 = Yes, 0 = No
device.replace({0: 'No', 1: 'Yes'}, inplace=True)
device.head()
# statistical summary of device
rp.summary_cat(device)
# statistical summary of sources of internet
internet = df['source_of_internet'].str.get_dummies(sep=', ')
internet.head()
# replace 1 = Yes, 0 = No
internet.replace({0: 'No', 1: 'Yes'}, inplace=True)
internet.head()
# summary of sources of internet
rp.summary_cat(internet)
# ## Effects on Health
# summary of health impacts
rp.summary_cat(df['did_you_feel_any_mental_disturbance_and_physical_problem_during_online_class'])
# multiple responses
df['what_kind_of_mental_disturbance_did_you_feel_during_online_class']
# binarize the responses
mental_health = df['what_kind_of_mental_disturbance_did_you_feel_during_online_class'].str.get_dummies(sep=', ')
mental_health.head()
# replace 1 = Yes, 0 = No
mental_health.replace({0: 'No', 1: 'Yes'}, inplace=True)
mental_health.head()
# summary of mental disturbance
rp.summary_cat(mental_health)
# physical problems
physical_health = df['what_kind_of_physical_problem__did_you_face_during_online_class'].str.get_dummies(sep=', ')
physical_health.head()
# replace 1 = Yes, 0 = No
physical_health.replace({0: 'No', 1: 'Yes'}, inplace=True)
physical_health.head()
# summary of physical problems
rp.summary_cat(physical_health)
# ## Perception on Online Class
rp.summary_cat(df['difficulty_during_online_classes'])
rp.summary_cat(df['the_worst_part_of_online_classes'])
rp.summary_cat(df['the_best_part_of_online_classes'])
rp.summary_cat(df['exam_preference'])
| notebooks/Impact of distance learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
import psycopg2
from psycopg2.extras import execute_values
try:
conn = psycopg2.connect(database='parceldatabase', user='techequity', password='<PASSWORD>')
print("successfully connected to database")
except:
print("I am unable to connect to the database")
# +
# cursor
cur = conn.cursor()
get_cols_query = "select center_lat, center_lon, nettaxablevalue from rawparceltable"
sample_data = [];
try:
cur.execute(get_cols_query)
sample_data = cur.fetchall()
print("query successful")
except:
print("query failed")
print(sample_data[0:10])
# -
nettax = [float(item[2]) for item in sample_data]
lat = [float(item[0]) for item in sample_data]
long = [float(item[1]) for item in sample_data]
data = list(zip(lat,long,nettax))
df = pd.DataFrame(data, columns=['latitutde','longitude',
'nettaxablevalue'])
df
from sklearn.model_selection import train_test_split
train, test = train_test_split(df,test_size=0.3)
x_train, y_train, x_test, y_test = train, train, test, test
from sklearn.preprocessing import MinMaxScaler
scaler = MinMaxScaler(feature_range=(0,1))
# +
x_train_scaled = scaler.fit_transform(x_train)
x_train = pd.DataFrame(x_train_scaled)
x_test_scaled = scaler.fit_transform(x_test)
x_test = pd.DataFrame(x_test_scaled)
# -
from sklearn import neighbors
from sklearn.metrics import mean_squared_error
from math import sqrt
rmse_val = []
for K in range(20):
K += 1
model = neighbors.KNeighborsRegressor(n_neighbors=K)
model.fit(x_train,y_train)
pred = model.predict(x_test)
error = sqrt(mean_squared_error(y_test,pred))
rmse_val.append(error)
print('RMSE value for K= ', K, 'is:', error)
# ### We get the lowest RMSE at K=9
curve = pd.DataFrame(rmse_val)
curve.plot()
plt.figure(figsize=(10,6))
plt.plot(range(0,20),rmse_val,color='blue', linestyle='dashed', marker='o',
markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
| data/k-nearest-neighbors.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="QAGIwGzeYXrq"
import pandas as pd
import matplotlib.pyplot as plt
from statsmodels.graphics.tsaplots import plot_acf, plot_pacf
from statsmodels.tsa.arima_model import ARIMA
import statsmodels.api as sm
import itertools
import numpy as np
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 276} id="NJRl_rP9YiAb" outputId="2cb0ea9f-f02d-42f6-9979-d951bcfcb811"
df_orig = pd.read_csv("https://raw.githubusercontent.com/JenBanks8585/Randomdata/main/data/Realty/Zip_ZORI_AllHomesPlusMultifamily_SSA%20(1).csv", parse_dates= True)
print(df_orig.shape)
df_orig.head(3)
# + id="XrPBO3hpYnzm"
# Load data and create a dictionary of csv files per location
def load_data():
url = "https://raw.githubusercontent.com/JenBanks8585/Randomdata/main/data/Realty/Zip_ZORI_AllHomesPlusMultifamily_SSA%20(1).csv"
df_orig = pd.read_csv(url, parse_dates = True)
df= df_orig.copy()
df = df.drop(['RegionID', 'MsaName', 'SizeRank' ], axis = 1)
df = df.fillna(method='ffill')
df_summary = df_orig[['RegionName', 'MsaName']].drop([0])
# get the list of region ID
regionId = list(df['RegionName'])
# Create a dictionary of dataframes for each region ID, with dates as rows.
reg_dict = {}
for id, data in df.groupby('RegionName'):
if id in regionId:
data.to_csv('/content/csvs/{}.csv'.format(id), header = True, index_label = False)
reg_dict[id] = pd.read_csv('/content/csvs/{}.csv'.format(id))
else:
continue
return df, reg_dict, df_summary
df = load_data()[0]
df_dict = load_data()[1]
df_summary = load_data()[2]
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="G8APn4RSY_qj" outputId="2198685d-29c1-4988-ec5d-9dc40225a193"
# Convert a dataframe into long format
def long_format(df_):
tidy = pd.melt(df_, id_vars = ['RegionName'], var_name= 'time')
tidy['time'] = pd.to_datetime(tidy['time'], infer_datetime_format=True)
tidy = tidy.dropna(subset = ['value'])
tidy = tidy.groupby('time').aggregate({'value':'mean'})
return tidy
long_format(df_dict[2474]).head(3)
# + id="iRG8-eMFZMhE"
# Iterate through all dataframes to convert them to long format
def make_tidy_all():
for id in df_dict.keys():
df_dict[id] = long_format(df_dict[id])
return df_dict
tidy_dict = make_tidy_all()
regionId_keys = list(tidy_dict.keys())
# + colab={"base_uri": "https://localhost:8080/", "height": 850} id="ayWXCdFBZQYo" outputId="69621177-923f-4366-da74-838a28312d09"
# Plot some of the location's values places with rolling average
for i in regionId_keys[5:7]:
tidy_dict[i].plot(figsize= (12,3))
tidy_dict[i].rolling(window=12).mean().plot(figsize = (12,3), color = 'red')
# + colab={"base_uri": "https://localhost:8080/", "height": 545} id="iNmHoJhaZcMs" outputId="52fd345a-55c0-402f-928c-8ac98854cb3d"
# plot_acf for index 1801 , this is to get an value for q
plot_acf(tidy_dict[1801][['value']]) #for value of q ( 1-7)
# + colab={"base_uri": "https://localhost:8080/", "height": 545} id="XCfk7UlHZ2Xj" outputId="e4fc5163-e377-4449-b284-1e53626d8209"
plot_pacf(tidy_dict[1801][['value']]) # For value of P
# + id="FpFJRherZ8x1"
# Get AIC scores using p=q=d = 1
def get_aic_across():
AIC = []
for zipcode in tidy_dict.keys():
mod = sm.tsa.statespace.SARIMAX(tidy_dict[zipcode]['value'],
order=(1, 1, 1),
seasonal_order=(1, 1, 1, 12),
enforce_stationarity=False,
enforce_invertibility=False)
res = mod.fit()
AIC.append([zipcode, np.abs(res.aic)])
#Putting information into dataframe
AIC_df = pd.DataFrame(AIC, columns = ["zip", 'aic'])
return AIC_df, res
# + colab={"base_uri": "https://localhost:8080/", "height": 465} id="k6kIg41daFEZ" outputId="1580a323-d743-4d88-e998-637df81eaab9"
results_sarimax = get_aic_across()[1]
results_sarimax.summary()
# + id="EC6AzYisaJPI"
# Creates a dataframe of different models based on location
# Because of the number of locations, the model generation was divided into segments.
# Note there are a total of 2263 locations, in this cell, we are just running the first 20
def get_model(begin_index=0, end_index=20):
model = []
for zipcode in list(tidy_dict.keys())[begin_index:end_index]:
mod = sm.tsa.statespace.SARIMAX(tidy_dict[zipcode]['value'],
order=(1, 1, 1),
seasonal_order=(1, 1, 1, 12),
enforce_stationarity=False,
enforce_invertibility=False)
res = mod.fit()
model.append([zipcode, res])
# Putting information into dataframe
model_df = pd.DataFrame(model, columns = ["zip", 'model'])
return model_df
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="OiOiRvjOcRhg" outputId="839e7469-cd13-464b-a087-f8be8e12c11b"
model_0_20 = get_model(0, 20)
model_0_20.head()
# + id="5DxAGybMa7TF"
# Run the rest of the locations and instantiate them
model_20_500 = get_model(20,500)
model_500_1000 = get_model(500,1000)
model_1000_1500 = get_model(1000,1500)
model_1500_2000 = get_model(1500,2000)
model_last = get_model(2000,2264)
# + id="cKHwFms7fxsZ"
import pickle
with open('/content/pickles/model_500_1000.pkl', 'wb') as f:
pickle.dump(model_500_1000, f)
# + id="lBQ4G78ZgedF"
# Pickle the models
model_list = [model_0_20,
model_20_500,
model_500_1000,
model_1000_1500,
model_1500_2000,
model_last]
for i in range(len(model_list)):
with open(f'/content/pickles/str{model_list[i]}', 'wb') as mod:
pickle.dump(model_list[i], mod)
# + id="MRAsLW93hyyO"
# Access the models
file = open("/content/pickles/model_0_20.pkl", 'rb')
df_20 = pickle.load(file)
file = open("/content/pickles/model_20_500.pkl", 'rb')
df_500 = pickle.load(file)
file = open("/content/pickles/model_500_1000.pkl", 'rb')
df_1000 = pickle.load(file)
file = open("/content/pickles/model_1000_1500.pkl", 'rb')
df_1500 = pickle.load(file)
file = open("/content/pickles/model_1500_2000.pkl", 'rb')
df_2000 = pickle.load(file)
file = open("/content/pickles/model_last.pkl", 'rb')
df_last = pickle.load(file)
# Concatenate the dataframes of zip codes/model
model_df_all= pd.concat([df_20, df_500, df_1000, df_1500, df_2000, df_last])
# + id="6XQ4Vi2Rfyx-"
# Gives forecast given zip code, number of months to forecast and the model dataframe
def get_forecast(zip, steps):
if zip in list(model_df_all['zip']):
forecast = model_df_all.loc[model_df_all['zip'] == zip, 'model'].item().get_forecast(steps = steps)
forecast_conf = forecast.conf_int()
return forecast_conf
return "This location is not available"
# + id="oKLqfzccjjrl" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="225116c3-b4f3-41d9-f412-06a4bc5c1348"
get_forecast( 1752, 6)
# + colab={"base_uri": "https://localhost:8080/", "height": 326} id="a9liPjjmNdzA" outputId="2be82185-44c3-4329-caf0-406b99a334fa"
# Create a dataframe of the next 5-months low/high forecasts.
dfs= []
for zip in model_df_all['zip']:
d = get_forecast(zip, 5)
dfs.append(d)
forecast_5months = pd.concat(dfs, axis =1, names = zip_codes)
# Create a csv file
forecast_5months.to_csv('forecast_5months.csv')
forecast_5months.head()
# + [markdown] id="HfXvqrSjx0b-"
# # Visualization
#
# + id="mN6twP2Nx-pd"
# Get forecast price based on zip and number of forecast values
def plot_forecast(zip, steps):
forecast = model_df_all.loc[model_df_all['zip'] == zip, 'model'].item().get_forecast(steps = steps)
forecast_conf = forecast.conf_int()
ax = tidy_dict[zip]['value'].plot(label='Observed', figsize= (10, 8))
forecast.predicted_mean.plot(ax=ax, label='Forecast')
ax.fill_between(forecast_conf.index,
forecast_conf.iloc[:, 0],
forecast_conf.iloc[:, 1], color='k', alpha=.25)
df_name = df_orig.copy()
df_name = df_name[['RegionName', 'MsaName']]
reg_name = df_name.loc[df_name['RegionName'] == zip, 'MsaName'].item()
ax.set_xlabel('Year')
ax.set_ylabel('Property Rental Price')
plt.title(reg_name)
plt.legend()
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="6-5q7dj_XZmc" outputId="29442c74-e40b-4224-9ae9-a675f851c766"
plot_forecast(2906, 18)
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="LLmpbqlTyAhV" outputId="7a9c6305-30b7-45ae-cf8f-ff93d315a7d9"
plot_forecast(33756, 18)
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="34kM0gSPWuwv" outputId="2c970e63-77a5-4b37-b918-eaa7da0e980a"
plot_forecast(75067,15)
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="rzYefTPzyAce" outputId="fce6f81e-11a4-419c-bdf1-a66360110491"
plot_forecast(92831, 20)
# + [markdown] id="ArQ799toxhvG"
# # Forecasting based on Zipcode
# + id="--OT-Xnmj4nG"
# !pip install uszipcode
# + id="FyZD6XqlkXyd"
# Grabbing the corresponding cities, states and counties of the zipcodes in the dataset
from uszipcode import SearchEngine
search = SearchEngine(simple_zipcode=True)
city = []
state = []
county = []
for zip in list(df_dict.keys()):
search = SearchEngine(simple_zipcode=True)
cityi = search.by_zipcode(zip).city
statei = search.by_zipcode(zip).state
countyi = search.by_zipcode(zip).county
city.append(cityi)
state.append(statei)
county.append(countyi)
# + id="5v2ZlY2_k9ky"
# Doubling each item because we have a low and high values per zipcode
zip_codes =list(df_dict.keys())
zip_codes2 = list(itertools.chain.from_iterable(itertools.repeat(x, 2) for x in zip_codes))
level = ['low', 'high'] * len(zip_codes)
cities = list(itertools.chain.from_iterable(itertools.repeat(x, 2) for x in city))
states = list(itertools.chain.from_iterable(itertools.repeat(x, 2) for x in state))
counties = list(itertools.chain.from_iterable(itertools.repeat(x, 2) for x in county))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="7UKdgy_VlNZR" outputId="fd35addf-9809-41b8-d2fc-3f73f0e8be9a"
# Reading from csv
forecast_5months = pd.read_csv('forecast_5months.csv', index_col = [0])
#Converting index to datetime object
forecast_5months.index = pd.to_datetime(forecast_5months.index, format='%Y%m%d',errors='ignore')
# Casting all values to integer
for col in forecast_5months.columns:
forecast_5months[col] = forecast_5months[col].astype(int)
# Transposing the dataframe
forecast_5months= forecast_5months.T
forecast_5months = forecast_5months.reset_index(drop = True)
forecast_5months.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="JFHu1wLMofso" outputId="8ea3e94d-96c7-4dfe-a54e-09c16250107b"
# Instantiate a new dataframe with other info
label_df = pd.DataFrame()
label_df['zip']= zip_codes2
label_df['city']= cities
label_df['zip']= label_df['zip'].apply(lambda x: str(x).zfill(5))
label_df['county']= counties
label_df['state']= states
label_df['level']= level
# Concatenate other info and forecast
forecast_label = pd.concat([label_df, forecast_5months], axis = 1)
forecast_label.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 111} id="iEKSc8TTpAJf" outputId="0df3992c-a603-486b-8905-00276891ddc7"
# Try it
forecast_label[forecast_label["city"]=="Evanston"]
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="Bx5gIwnKpQhc" outputId="de00046f-c797-4345-ee45-bc5bbea4ea3a"
city_df = forecast_label[['zip', 'level', '2021-01-01', '2021-02-01', '2021-03-01', '2021-04-01', '2021-05-01']]
city_df.columns = ['zip', 'level', 'in_1_month', 'in_2_months', 'in_3_months', 'in_4_months', 'in_5_months']
city_df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="-kBzIOcyp27X" outputId="cec12629-525c-42ac-ddee-011d6ac818e7"
# Grab csv version of database table 'cities'
df = pd.read_csv("/content/city.csv", index_col = [0])
# Take a subset
df_city = df[['city_id', 'city_name', 'state_abbreviation']]
df_city.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="BSD6o680qT51" outputId="f39dc741-7a9b-41a8-def5-8b110ae34d81"
# Get the one word version of the city names
places = []
for place in list(df_city['city_name']):
place = place.split(" ")[0]
places.append(place)
df_city['city'] = places
df_city = df_city.drop(columns = ['city_name'])
df_city.head(3)
# + id="-nxOsnR-qf1l"
# merge the two dataframes to get city_id, this will be used as foreign_key to connect to other tables in the database
new_df = forecast_label.merge(df_city, how = "left", left_on = ['city', 'state'], right_on = ['city', 'state_abbreviation'])
new_df= new_df.dropna()
city_i = []
for i in new_df['city_id']:
id = int(i)
city_i.append(id)
new_df['city_id'] = city_i
# + [markdown] id="0Ev84eDrrsju"
# # Using Zipcode as index
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="u5BzqPjWrUnK" outputId="b6a7f656-8cf3-479b-a98c-fa1f35ea0ae4"
# Renaming columns
zip_df = new_df[['zip', 'city_id', 'level', '2021-01-01', '2021-02-01', '2021-03-01', '2021-04-01', '2021-05-01']]
zip_df.columns = ['zip', 'city_id', 'level', 'in_1_month', 'in_2_months', 'in_3_months', 'in_4_months', 'in_5_months']
zip_df.head(3)
# + colab={"base_uri": "https://localhost:8080/", "height": 266} id="E7v9LwV1r6S4" outputId="36997f5e-fc7d-44d7-e2b1-2a15612b19d3"
# # pivot table
city_d = pd.pivot_table(zip_df, index = ['zip'], columns = 'level', values = ['in_1_month',
'in_2_months',
'in_3_months',
'in_4_months',
'in_5_months'])
city_d.head(5)
# + id="Mg2sZbHvsMUr"
# Grabbing items in the dataframe
index = [int(i) for i in list(city_d.index)]
in_1_month_high = list(city_d[( 'in_1_month', 'high')])
in_1_month_low = list(city_d[( 'in_1_month', 'low')])
in_2_month_high = list(city_d[( 'in_2_months', 'high')])
in_2_month_low = list(city_d[( 'in_2_months', 'low')])
in_3_month_high = list(city_d[( 'in_3_months', 'high')])
in_3_month_low = list(city_d[( 'in_3_months', 'low')])
in_4_month_high = list(city_d[( 'in_4_months', 'high')])
in_4_month_low = list(city_d[( 'in_4_months', 'low')])
in_5_month_high = list(city_d[( 'in_5_months', 'high')])
in_5_month_low = list(city_d[( 'in_5_months', 'low')])
# + colab={"base_uri": "https://localhost:8080/"} id="FaS0zs2A0oBi" outputId="2b378ff5-a7c2-4348-9615-45921009ce80"
in_1_month_high[:3], in_2_month_high[:3]
# + id="mzWJVLuSsUpZ"
# Creating a list of dictionaries zip:forecast
zip_dict =[]
for i in range(len(city_d.index)):
a = {str(index[i]): [{'in_1_month':{'low':in_1_month_low[i], 'high':in_1_month_high[i]}},
{'in_2_months':{'low':in_2_month_low[i], 'high':in_2_month_high[i]}},
{'in_3_months':{'low':in_3_month_low[i], 'high':in_3_month_high[i]}},
{'in_4_months':{'low':in_4_month_low[i], 'high':in_4_month_high[i]}},
{'in_5_months':{'low':in_5_month_low[i], 'high':in_5_month_high[i]}}
]
}
zip_dict.append(a)
# + id="1rJxeWQlzWE8"
zip_dict
# + id="LOKkA7UcvgoI"
# Get forecast given zip code
zips = [list(zip_dict[i].keys())[0] for i in range(len(zip_dict))]
def get_forecast(zip):
if zip in zips:
for i in range(len(zip_dict)):
zip = list(zip_dict[i].keys())[0]
f = zip_dict[i][zip]
return f
else:
return "No forecast for this location"
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="yDbtTegMwgzG" outputId="8865fef5-2354-474f-a85c-b04284c50724"
get_forecast('175332')
# + colab={"base_uri": "https://localhost:8080/"} id="bV6gO6e6xmzy" outputId="39914748-ec39-4c46-9732-fbe421fb5c69"
get_forecast('1752')
# + id="ysfOSg-A5H2_"
# save as json object
import json
with open('zip_dict.json', 'w') as json_file:
json.dump(zip_dict, json_file)
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="Lx3RJ5Jh3r6r" outputId="ff8c882e-ddd2-4f1b-9285-1777b0a2fef8"
# Convert list of dictionary to dataframe format
zips = []
values = []
for i in range(len(zip_dict)):
zip = zip_dict[i].keys()
zips.append(zip)
value = zip_dict[i].values()
values.append(value)
zip_i = [item for sublist in zips for item in sublist]
values_lst = [item for sublist in values for item in sublist]
zip_forecast_df = pd.DataFrame()
zip_forecast_df['zip'] = zip_i
zip_forecast_df['forecast'] = values_lst
zip_forecast_df.head(3)
# + id="FJu8tRlN5NyL"
# Save to csv to be uploaded to github
zip_forecast_df.to_csv('zip_forecast.csv')
# + [markdown] id="afy-3wK-7HCA"
# # Forecasting Based on city_id
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="2kPRyVbI7YDs" outputId="e3c98fa9-006a-498f-9452-6a2bb282fc42"
# # pivot table
city_d= pd.pivot_table(zip_df, index = ['city_id'], columns = 'level', values = ['in_1_month',
'in_2_months',
'in_3_months',
'in_4_months',
'in_5_months'])
city_d.head(3)
# + id="LOajydQo5UuL"
# Grabbing items in the dataframe
index = [int(i) for i in list(city_d.index)]
in_1_month_high = list(city_d[( 'in_1_month', 'high')])
in_1_month_low = list(city_d[( 'in_1_month', 'low')])
in_2_month_high = list(city_d[( 'in_2_months', 'high')])
in_2_month_low = list(city_d[( 'in_2_months', 'low')])
in_3_month_high = list(city_d[( 'in_3_months', 'high')])
in_3_month_low = list(city_d[( 'in_3_months', 'low')])
in_4_month_high = list(city_d[( 'in_4_months', 'high')])
in_4_month_low = list(city_d[( 'in_4_months', 'low')])
in_5_month_high = list(city_d[( 'in_5_months', 'high')])
in_5_month_low = list(city_d[( 'in_5_months', 'low')])
# + id="Vh7FqRr37uL2"
city_id_dict =[]
for i in range(len(city_d.index)):
a = {str(index[i]): [{'in_1_month':{'low':in_1_month_low[i], 'high':in_1_month_high[i]}},
{'in_2_months':{'low':in_2_month_low[i], 'high':in_2_month_high[i]}},
{'in_3_months':{'low':in_3_month_low[i], 'high':in_3_month_high[i]}},
{'in_4_months':{'low':in_4_month_low[i], 'high':in_4_month_high[i]}},
{'in_5_months':{'low':in_5_month_low[i], 'high':in_5_month_high[i]}}
]
}
city_id_dict.append(a)
# + colab={"base_uri": "https://localhost:8080/"} id="-AS-DB_p7x4T" outputId="5b6287bd-3535-48be-b8b7-083548e127c1"
city_id_dict[1]
# + id="BmjMy0t58DYX"
# save json object
with open('city_id_dict.json', 'w') as json_file:
json.dump(city_id_dict, json_file)
# + id="Xq574R178ZUn"
# Get forecast given city_id
ids = [list(city_id_dict[i].keys())[0] for i in range(len(city_id_dict))]
def get_forecast_id(city_id):
if city_id in ids:
for i in range(len(city_id_dict)):
c_id = list(city_id_dict[i].keys())[0]
f = city_id_dict[i][city_id]
return f
else:
return "No forecast for this location"
# + colab={"base_uri": "https://localhost:8080/"} id="KfmJFMhx92pY" outputId="c0cc05ea-ab6c-4952-a3c9-5bccb9aeb346"
get_forecast_id('100820')
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="Yp22GKfz6oDg" outputId="0a45d450-98b2-4c49-8c32-e72b513d52ba"
# Convert list of dictionary to dataframe format
city_ids = []
values2 = []
for i in range(len(city_id_dict)):
id = list(city_id_dict[i].keys())
city_ids.append(id)
value = city_id_dict[i].values()
values2.append(value)
city_i = [item for sublist in city_ids for item in sublist]
values2_lst = [item for sublist in values2 for item in sublist]
city_id_forecast_df = pd.DataFrame()
city_id_forecast_df['city_id'] = city_i
city_id_forecast_df['rental_forecast'] = values2_lst
city_id_forecast_df.head(3)
# + colab={"base_uri": "https://localhost:8080/"} id="C_DXUp8DCAlz" outputId="9116ce8e-2775-448e-e650-0aa6a5e6f386"
type(city_id_forecast_df['rental_forecast'][0])
# + id="_5R-rM_Q7svj"
# Save to csv to be uploaded to github
city_id_forecast_df.to_csv('rental_forecast.csv')
# + [markdown] id="goj2-sH_G-bF"
# # Database
# + id="UOS6t-EoG18W"
import ast
# Create table `rental_forecast` from dataframe
database_url = os.getenv('PRODUCTION_DATABASE_URL')
url = 'https://raw.githubusercontent.com/JenBanks8585/Randomdata/main/data/Realty/rental_forecast.csv'
rental_df = pd.read_csv(url, index_col=[0])
rental_df.columns = ['city_id', 'rental_forecast']
rental_df['rental_forecast'] = rental_df['rental_forecast'].apply(lambda x: x.strip(']['))
rental_df['rental_forecast'] = rental_df['rental_forecast'].apply(lambda x: ast.literal_eval(x))
rental_df['rental_forecast'] = rental_df['rental_forecast'].apply(lambda x: json.dumps(x))
rental_df['city_id'] = rental_df['city_id'].apply(lambda x: str(x).zfill(7))
query = '''
CREATE TABLE "rental_forecasts" (
"city_id" char(7) NOT NULL,
"rental_forecast" json NOT NULL,
CONSTRAINT "rental_forecast_pk" PRIMARY KEY ("city_id"))
WITH (
OIDS=FALSE
);
ALTER TABLE "rental_forecast" ADD CONSTRAINT "rental_forecast_fk0" FOREIGN KEY ("city_id") REFERENCES "cities"("city_id");
'''
rental_df.to_sql('rental_forecast', database_url, if_exists='append', index=False)
| notebooks/sarimax_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Checkpoint 1: Data Cleaning 1
# Load the companies and rounds data (provided on the previous page) into two data frames and name them companies and rounds2 respectively.
import pandas as pd
import numpy as np
companies = pd.read_csv('companies.txt', encoding='ISO-8859-1', sep = '\t')
rounds2 = pd.read_csv('rounds2.csv', encoding='ISO-8859-1')
companies.head()
rounds2.head()
# ### company_permalink and permalink columns in rounds2 and companies dataframes are encoded differently and also not case sensitive
rounds2['company_permalink'] = rounds2['company_permalink'].str.encode('utf-8').str.decode('ascii', 'ignore').str.lower().str.strip()
companies['permalink'] = companies['permalink'].str.encode('utf-8').str.decode('ascii', 'ignore').str.lower().str.strip()
companies.shape
companies.columns
companies.index
companies.describe()
# ### As we can see here permalink column has all rows unique, so it can be a unique and primary column
companies.info()
companies.isnull().sum()
rounds2.info()
rounds2.isnull().sum()
# ### Since approx 20000 rows have nulls in raised_amount_usd column, which turns out to be the fact data
# ### we need to handle these rows
rounds2 = rounds2[~(rounds2.isnull().raised_amount_usd)]
rounds2.isnull().sum()
# # Check to see how many rows have all rows null
rounds2.isnull().all(axis=1).sum()
rounds2[rounds2.isnull().sum(axis=1) > 2]
# ### No row is having more than 2 null values in them
# # Table 1.1 Answers
# ## 1. How many unique companies are present in rounds2?
# - we use drop_duplicates to remove duplicates from rounds2 using company_permalink column which is primary column for companies dataset
# - (Since we have already made this column lowercase so it will exactly match the columns)
len(rounds2.drop_duplicates('company_permalink'))
# ## 2. How many unique companies are present in the companies file?
# - We use drop_duplicates to remove duplicates based on permalink column
# - since it is lowercase it will match all unique
len(companies.drop_duplicates('permalink'))
# ## 3. In the companies data frame, which column can be used as the unique key for each company? Write the name of the column.
companies.describe()
# ### As we can see here permalink column has all rows unique, so it can be a unique and primary column
# ## 4. Are there any companies in the rounds2 file which are not present in companies ? Answer Y/N.
rounds2[~(rounds2.company_permalink.isin(companies.permalink))]
# ### Answer is NO. i.e. rounds2 dataset has all the companies present in companies dataset.
# ## 5. Merge the two data frames so that all variables (columns) in the companies frame are added to the rounds2 data frame. Name the merged frame master_frame. How many observations are present in master_frame ?
master_frame = pd.merge(companies, rounds2, how='inner',
left_on='permalink', right_on='company_permalink')
master_frame.head()
master_frame.info()
# ### 94959 Observations are present in master_frame
# # Table 2.1
# ## Average funding amount of angel, seed, venture and private equity type
group_by_funding = master_frame.groupby(by='funding_round_type')
avg_funding = group_by_funding.mean()
avg_funding.loc[['angel', 'seed', 'venture', 'private_equity']]
avg_funding.loc[['angel', 'seed', 'venture', 'private_equity']].raised_amount_usd.values
avg_funding
# ## Considering that Spark Funds wants to invest between 5 to 15 million USD per investment round, which investment type is the most suitable for them?
avg_funding[avg_funding.raised_amount_usd.between(50_00_000, 1_50_00_000)]
# # Table 3.1
# ## 1. Top english speaking country
master_frame.head()
# ## top nine countries which have received the highest total funding (across ALL sectors for the chosen investment type)
total_funding_agg = master_frame[master_frame.funding_round_type == 'venture'].groupby(by='country_code').sum()
top9 = total_funding_agg.sort_values(by='raised_amount_usd', ascending=False).head(9)
top9
# # Table 4.1
# ## Checkpoint 4: Sector Analysis
main_sector = pd.read_csv('mapping.csv', encoding='ISO-8859-1')
main_sector.head()
main_sector.info()
main_sector[main_sector.Blanks==1]
main_sector = main_sector.loc[~(main_sector.isnull().category_list)]
main_sector.drop('Blanks', inplace=True, axis=1)
main_sector.head()
master_frame.head()
# ## Extract primary sector from category_list
master_frame['primary_sector'] = master_frame.category_list.str.split(pat='|').str.get(0)
# ## We need to unpivot the main_sector (mapping.csv) data so that main_sector can be extracted from 0-1 flags
temp = main_sector.melt(id_vars='category_list', value_name='flag', var_name='main_sector')
main_sector_unpivoted = temp[temp.flag == 1].drop('flag', axis=1)
main_sector_unpivoted.index = range(0, len(main_sector_unpivoted))
main_sector_unpivoted.head()
# ## We then need to merge the master_frame with the main_sector_unpivoted dataset
merged_master = master_frame.merge(main_sector_unpivoted, how='inner', left_on='primary_sector', right_on='category_list')
merged_master.head()
# # Checkpoint 5: Sector Analysis 2
# ## Heavily invested main sectors for each country for venture funcding type and total funding between 5-15 million
venture_funding = merged_master[
(merged_master.funding_round_type == 'venture')
& (merged_master.country_code.isin(['USA', 'GBR', 'IND']))
& (merged_master.raised_amount_usd.between(5_000_000, 15_000_000))].copy()
# ### Changing the amount unit in column `raised_amount_usd` to million-dollor
venture_funding.loc[:, 'raised_amount_usd'] = venture_funding.raised_amount_usd.apply(lambda x: x / 1_000_000)
grp = venture_funding.groupby(by=['country_code', 'main_sector'], as_index=False)
grp_agg = grp.raised_amount_usd.agg({'total_investments': 'count', 'total_investment_amount': 'sum'})
master_final = pd.merge(venture_funding, grp_agg, on=['country_code', 'main_sector'])
master_final.head()
D1 = master_final[master_final.country_code == 'USA'].copy()
D2 = master_final[master_final.country_code == 'GBR'].copy()
D3 = master_final[master_final.country_code == 'IND'].copy()
# ## Table 5.1
# ### 1. Total number of investments (count)
# ### 2. Total amount of investment (USD)
grp1 = venture_funding.groupby(by=['country_code'], as_index=False)
grp1.raised_amount_usd.agg({'total_investments': 'count', 'total_investment_amount': 'sum'}).sort_values(by='total_investment_amount', ascending=False)
# ### 3. Top sector (based on count of investments)
# ### 4. Second-best sector (based on count of investments)
# ### 5. Third-best sector (based on count of investments)
# ### 6. Number of investments in the top sector (refer to point 3)
# ### 7. Number of investments in the second-best sector (refer to point 4)
# ### 8. Number of investments in the third-best sector (refer to point 5)
grp_agg.sort_values(by=['country_code', 'total_investments'], ascending=False)
# ### 9. For the top sector count-wise (point 3), which company received the highest investment?
# #### we need to find, which company recieved highest investment in `Others` sector for all 3 countries
venture_others = venture_funding[venture_funding.main_sector == 'Others'].copy()
venture_others.sort_values(by='raised_amount_usd', ascending=False, inplace=True)
usa_venture_others = venture_others[venture_others.country_code == 'USA']
gbr_venture_others = venture_others[venture_others.country_code == 'GBR']
ind_venture_others = venture_others[venture_others.country_code == 'IND']
print(usa_venture_others.name.head(1))
print(gbr_venture_others.name.head(1))
print(ind_venture_others.name.head(1))
# ### 10. For the second-best sector count-wise (point 4), which company received the highest investment?
venture_cleantech = venture_funding[venture_funding.main_sector == 'Cleantech / Semiconductors'].copy()
venture_news = venture_funding[venture_funding.main_sector == 'News, Search and Messaging'].copy()
venture_cleantech[venture_cleantech.country_code == 'USA'].name.head(1)
venture_cleantech[venture_cleantech.country_code == 'GBR'].name.head(1)
venture_news[venture_news.country_code == 'IND'].name.head(1)
| Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # Week 07, Part 1
#
# ### Topic
# 1. Foundations for Inference
# 1. BACK TO SLIDES FOR WHEN TO USE SAMPLE MEANS
#
# resize
require(repr)
options(repr.plot.width=4, repr.plot.height=4)
# ## 1. Foundations for inference
# Load the fish dataset from week02:
fishdata = read.csv('../week02/undata_fish_2020.csv')
# Take the log of the trade in USD, like before:
log_trade_usd = log10(fishdata$Trade..USD.)
# A quick histogram to remind ourselves what this data looks like:
hist(log_trade_usd)
# Now, let's assume this distribution is a measurement of the population. I.e., this dataset represents all of the trade of fish all over the globe. In reality, it is itself a sample, of which we don't really know the data collection methods, but lets suspend our disbelief for the moment.
#
# Let's pretend we are going out and collecting a random sample of the trade (in units of log10) of fish in the world. We are asking over random years, countries, and import or export - what is the dollar value of the transaction? In short - we are *sampling* from this background population.
#
# Let's create a "mock" sample from our "mock" population data:
nsamples = 10 # Let's start with 10 random samples.
# Next, we'll pull randomly using our "sample" function we used last week:
mysample = log_trade_usd[sample(1:length(log_trade_usd), nsamples, replace=FALSE)]
# Here, we are using sample to pull random *indices* that run the length of the log_trade_usd vector - a total of nsamples.
print(sample(1:length(log_trade_usd), nsamples, replace=FALSE))
print(mysample)
# In opposition to last week we will be setting `replace=FALSE`. This last part just means "sampling without replacement" so we don't double count indicies.
# Let's print out the mean of our sample and compare this to the mean of our "mock" population:
print('Sample Mean, Population Mean')
print(c(mean(mysample), mean(log_trade_usd)))
# Note: if I run this a few times with different random samples, I get different means for my sample. **RUN A FEW TIMES ON YOUR OWN**
# Some questions we might have at this point:
# * So, the means are close, but are they close enough?
# * How to quantify how "close" or "far away" they are?
#
# Let's try to see how good our means are by taking a bunch of samples and calculating their means. We'll store these in a vector of "sample means":
mymeans = c() # where the sample means are stored
nmeans = 5 #start with 5 samples -> 5 sample means
# Use sample function to generate samples of length nsamples (10), take the mean of these, and store it in our sample mean vector. Do this nmeans times:
for (i in 0:nmeans){
# grab the sample like before
mysample = log_trade_usd[sample(1:length(log_trade_usd), nsamples, replace=FALSE)]
# take the mean and store it:
mymeans = append(mymeans, mean(mysample))
}
# Let's make a histogram of these means - this is refered to as our "sampling distribution" in the book:
hist(mymeans, xlab="Sample Mean", prob=TRUE, xlim=c(2,7))
# We'll overplot the actual population mean as well:
hist(mymeans, xlab="Sample Mean", prob=TRUE, xlim=c(2,7))
abline(v=mean(log_trade_usd), col="red", lwd=4) # population mean
# If we start with nmeans = 5, i.e. a sample of means with only 5 measurements, we see a lot of variablity if we run this a few times. **RUN A FEW TIMES ON YOUR OWN**
# What if we change nmeans to 50? 500? 5000?
#
# Q: What do we expect to see? How will our histogram change?
#
# *Take a moment to answer this on your own*
# We see our collection of means is clustered around our population mean and that this clustering gets "tighter" if we increase the number of samplings that we do.
#
# But how can we quantify how "good" our distribution of means is based on the number of sample means we take?
#
# Enter the *standard error*!
#
#
# **Standard Error** - first a calculation:
SE = sd(log_trade_usd)/nsamples**0.5
# Don't worry,we'll talk about this equation in a few minutes.
#
# Here we are calculating the Standard Error based on the population. Let's overplot the theoretical spread on our sample mean distribution based on the mean of our population and the standard error of the population:
# +
hist(mymeans, xlab="Sample Mean", prob=TRUE, xlim=c(2,7))
abline(v=mean(log_trade_usd), col="red", lwd=4) # population mean
x = seq(2,7, length=200)
lines(x, dnorm(x, mean=mean(log_trade_usd), sd=SE), col="green")
# -
# Here, we've overplotted how much we *think* the means should vary from the true, population, mean. We notice that our simulated distribution of sample means lines up nicely with our theoretical. We can see that this is true when we change our the size of our samples (nsamples) **GO BACK AND DO ON YOUR OWN**
#
# Often, don't know anything about the population, only have the sample. Also, we often just have the *one* sample, not a whole bunch where we can calculate the distribution of means. What do we do then? In this case, we can get a handle on how much we *estimate* the mean should vary by using the standard deviation of the values measured in our *sample* instead of the population:
# +
hist(mymeans, xlab="Sample Mean", prob=TRUE, xlim=c(2,7))
abline(v=mean(log_trade_usd), col="red", lwd=4) # population mean
SE_estimate = sd(mysample)/nsamples**0.5
lines(x, dnorm(x,mean=mean(mysample), sd=SE_estimate), col="blue")
# -
# Again, the Standard Error using the sample SD is close to the population measurement, but not exact.
#
# Let's summarize these things a bit more and get a handle on how the Standard Error is used.
# ## 2. BACK TO SLIDES FOR WHEN TO USE SAMPLE MEANS
#
| week07/prep_part1_foundationsForInference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Iterative Verfahren
#
# ### Fixpunktiteration
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Ein Beispiel aus der Rohrhydraulik:
#
# Zur Bestimmung der [Rohrreibungszahl](https://de.wikipedia.org/wiki/Rohrreibungszahl) $\lambda$ kann bei glatten, turbulent durchströmten Rohren die implizite Formel von Prandtl verwendet werden:
#
# $$\frac{1}{\sqrt\lambda} = 2\cdot\log\left(\text{Re}\cdot\sqrt\lambda\right) - 0,8$$
#
# Mathe-Nerds lösen die Gleichung mithilfe der [Lambertschen W-Funktion](https://de.wikipedia.org/wiki/Lambertsche_W-Funktion), Ingenieure jedoch meist mit einem numerischen Verfahren.
#
# Die Lösung für die Prandtl-Formel lässt sich grafisch als Schnittpunkt der linken und rechten Seite finden.
#
# Wir definieren für die linke und rechte Seite separat:
#
# $$LHS\left(\lambda\right) = \frac{1}{\sqrt\lambda}$$
#
# $$RHS\left(\lambda\right) = 2\cdot\log\left(\text{Re}\cdot\sqrt\lambda\right) - 0,8$$
#
# Für eine Reynoldszahl $\text{Re} = 6,4\cdot 10^6$ ergeben sich dann folgende Kurvenverläufe:
# + button=false new_sheet=false run_control={"read_only": false}
import matplotlib.pyplot as plt
import numpy as np
import math
# %config InlineBackend.figure_format = 'svg'
# %matplotlib inline
# linke Seite der Gleichung (left hand side)
def LHS(lamb):
return 1/np.sqrt(lamb)
# rechte Seite der Gleichung (right hand side)
def RHS(lamb, Re):
return 2.0 * np.log10(Re * np.sqrt(lamb)) - 0.8
# Array mit äquidistanten Werten für lambda und Festlegen der Re-Zahl:
lamb = np.arange(0.0001, 0.05, 0.0001);
Re = 6.4e6
plt.plot(lamb, LHS(lamb), label="LHS")
plt.plot(lamb, RHS(lamb, Re), label="RHS")
plt.axis([0, 0.02, 5, 15])
plt.ylabel('LHS, RHS')
plt.xlabel('Rohrreibungszahl $\lambda$')
plt.grid()
plt.legend()
plt.show();
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Als Lösung ("Fixpunkt"), bei dem linke und rechte Seite denselben Wert annehmen lässt sich für $\lambda$ ein Wert von etwa 0,0085 ablesen.
#
# Mithilfe einer Fixpunktiteration lässt sich dieser auch iterativ bestimmen. Hierzu formen wir die Prandtl-Formel so um, dass auf der linken Seite nur noch $\lambda$ steht:
#
# $$\lambda = \frac{1}{\left[RHS\left(\lambda\right)\right]^2}$$
#
# Damit lässt sich nun eine Iterationsvorschrift formulieren:
#
# $$\lambda_{i+1} = \frac{1}{\left[RHS\left(\lambda_i\right)\right]^2}$$
# + button=false new_sheet=false run_control={"read_only": false}
# Startwert für lambda:
lamb_alt = 100
# Liste, um Zwischenergebnisse zu speichern
lambda_i = []
lambda_i.append(lamb_alt)
# Fixpunkt-Algorithmus
for iteration in range(0, 5):
lamb_neu = 1 / (RHS(lamb_alt, Re)**2)
lambda_i.append(lamb_neu)
lamb_alt = lamb_neu
fehler = (RHS(lamb_neu, Re)-LHS(lamb_neu)) / RHS(lamb_neu, Re)
plt.plot(lamb, LHS(lamb))
plt.plot(lamb, RHS(lamb, Re))
plt.plot(lambda_i, LHS(lambda_i), 'o')
for i, txt in enumerate(lambda_i):
plt.annotate(i, (lambda_i[i], LHS(lambda_i[i])))
plt.axis([0, 0.02, 5, 15])
plt.ylabel('LHS, RHS')
plt.xlabel('Rohrreibungszahl $\lambda$')
plt.show();
print ("lambda = ", lamb_neu, ", Fehler in %: ", fehler*100)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Newton-Verfahren
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Das [Newton-Verfahren](https://de.wikipedia.org/wiki/Newton-Verfahren) ist eine weitere Möglichkeit, um Gleichungen iterativ zu lösen. Hierzu wird die Gleichung so umgestellt, dass sich das Problem in eine Nullstellensuche konvertiert. Die oben behandelte Rohrreibungsgleichung ergibt dann:
#
# $$f(\lambda) = \frac{1}{\sqrt\lambda} - 2\cdot\log\left(\text{Re}\cdot\sqrt\lambda\right) + 0,8 = 0$$
#
# Ausgehend von einem geschätzten Startwert für die Nullstelle $\lambda_i$ wird die Steigung $f'(\lambda_i)$ der Funktion berechnet. Die Tangente im Punkt $(\lambda_i,f(\lambda_i))$ ist dann:
#
# $$t(\lambda) = f(\lambda_i) + f'(\lambda_i)\cdot (\lambda - \lambda_i)$$
#
# Der Schnittpunkt dieser Tangente mit der $\lambda$-Achse ergibt den neuen Näherungswert für die Nullstelle und damit die Iterationsvorschrift:
#
# $$\lambda_{i+1} = \lambda_i - \frac{f(\lambda_i)}{f'(\lambda_i)}$$
#
# Im Beispiel mit der Rohrreibungsgleichung ist die Ableitung:
#
# $$f'(\lambda) = -\frac{1}{2\cdot\lambda^{3/2}} - \frac{1}{\lambda}$$
# + button=false new_sheet=false run_control={"read_only": false}
# Startwert für lambda:
lamb_alt = 0.01
# Liste, um Zwischenergebnisse zu speichern
lambda_newton_i = []
lambda_newton_i.append(lamb_alt)
# die Funktion f
def f(lamb, Re):
return 1/np.sqrt(lamb) - 2.0 * np.log10(Re * np.sqrt(lamb)) + 0.8
# die Ableitung von f
def f_strich(lamb):
return -1/(2*lamb**1.5) - 1/(lamb*math.log(10))
# die Tangente an f (nur zur Visualisierung, wird eigentlich nicht benötigt)
def tangente_f(x, lamb, Re):
return f(lamb,Re)+f_strich(lamb)*(x-lamb)
# Newton-Verfahren:
for iteration in range(0, 15):
lamb_neu = lamb_alt - f(lamb_alt, Re)/f_strich(lamb_alt)
lambda_newton_i.append(lamb_neu)
lamb_alt = lamb_neu
fehler = (RHS(lamb_neu, Re)-LHS(lamb_neu)) / RHS(lamb_neu, Re)
# Ergebnisse im Diagramm darstellen:
plt.plot(lamb, f(lamb, Re))
plt.plot(lambda_newton_i, f(lambda_newton_i, Re), 'o')
for i, txt in enumerate(lambda_newton_i):
plt.annotate(i, (lambda_newton_i[i], f(lambda_newton_i[i], Re)))
plt.plot(lamb, tangente_f(lamb, lambda_newton_i[i], Re))
plt.plot([0,0.02],[0,0],'k', linewidth=1)
plt.axis([0, 0.02, -4, 6])
plt.ylabel('f = LHS - RHS')
plt.xlabel('Rohrreibungszahl $\lambda$')
plt.show();
print ("lambda = ", lamb_neu, ", Fehler in %: ", fehler*100)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Verfahren zur Lösung von Gleichungssystemen
#
# Weitere Verfahren, zur Lösung von ganzen Gleichungssystemen werden in Kapitel 4 vorgestellt. Diese Verfahren werden z.B. verwendet, um die riesigen Gleichungssysteme zu lösen, die bei der Diskretisierung von Transportgleichungen mithilfe der Finite-Differenzen- (FDM), Finite-Elemente- (FEM) oder Finite-Volumen-Methode (FVM) entstehen.
#
# Prominente Vertreter sind das Gauß-Verfahren (Gauß-Seidel-Verfahren) und der Thomas-Algorithmus.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# [Hier](1_3-Numerik_Anfangswertaufgaben.ipynb) geht's weiter oder [hier](index.ipynb) zurück zur Übersicht.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ---
# ###### Copyright (c) 2018, <NAME> und <NAME>
#
# Der folgende Python-Code darf ignoriert werden. Er dient nur dazu, die richtige Formatvorlage für die Jupyter-Notebooks zu laden.
# + button=false new_sheet=false run_control={"read_only": false}
from IPython.core.display import HTML
def css_styling():
styles = open('TFDStyle.css', 'r').read()
return HTML(styles)
css_styling()
# -
| 3_1-Numerik_Iterative_Verfahren.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # EXERCISE 1
#Program to print the pattern for a number input by the user.
num = int(input("Enter a number to generate its pattern = "))
for i in range(1,num+1):
for j in range(1,i+1):
print(j,end=" ")
print()
# # EXERCISE 2
#Program to find prime numbers between 2 to 50 using nested for loops.
num = 2
for i in range(2,50):
j = 2
while (j<=(i/2)):
if (i%j==0):
break
j+=1
if (j>i/j):
print(i,"is a prime number")
print("Bye Bye!!")
# # EXERCISE 3
#Program to calculate the factorial of a given number using loop nested iside an if...else.
num = int(input("Enter any number: "))
fact = 1
if num<0:
print("Sorry, factorial does not exist for negative numbers.")
elif num==0:
print("The factorial of 0 is 1")
else:
for i in range(1,num+1):
fact=fact*i
print("factorial of ",num," is ",fact)
# # EXERCISE 4
#Program to check if the input number is prime or not.
num=int(input("Enter the number to be checked: "))
flag = 0 #presumed number is a prime number
if num > 1:
for i in range(2,int(num/2)):
if (num % i == 0):
flag = 1 #num is not a prime number
break
if flag == 1:
print(num, "is not a prime number")
else:
print(num, "is a prime number")
else:
print("Entered number is <= 1, execute again!")
| Week4/Flow of control.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## The goal of this notebook is to present both array and string related challenges and their solutions
# ### Challenge 1: Determine if a string has all unique characters
#
# There are several obvious solutions to this problem, here are just some of them, and their explanatiosn
# Solution Using Hashset
# This solution is easy and short enough, and can be implemented in O(N), assuming the Hashset function will
# have little to no colision
def has_all_unique_characters_hashset(word):
characters = set()
for character in word:
if character in characters:
return False
else:
characters.add(character);
return True
# This solution works by creating an array of 26 elements (one for each possible character of the english alphabet)
# and flagging it's index with a "1" instead of "0" everytime we see a different character.
# If we ever find an index with a number other than "0" it means we found a duplicate
# The total cost is still O(N) + O(N) which is O(N).
# NOTE: the Ord() method converts a character to it's integer value on the ASCII table (a = 97, and it's used as a base for the math of this code)
def has_all_unique_characters_array(word):
characters = [0] * 26
# Lower Case String
word = word.lower()
for character in word:
if characters[ord(character) - 97] is not 0:
return False
else :
characters[ord(character) - 97] = 1
return True
# This solution will sort the string (which can cost O(N LOG N) depending on the sort algorithm) and will simply compare if
# each element it finds is the same as the following one. If any case matches this comparisson, the code will return that
# it found a duplicate.
# The total cost of this solution is O(N LOG N) + O(N) (sorting + iterating over the array), which is O(N LOG N)
def has_all_unique_characters_array_sorting(word):
# Sorting the string
sorted_word = sorted(word)
for i in range(0, len(sorted_word) -1):
if sorted_word[i] is sorted_word[i + 1]:
return False
return True
# +
# Testing it
word_with_duplicates = 'banana'
word_without_duplicates = 'python'
print 'Answers using Hashset'
print has_all_unique_characters_hashset(word_with_duplicates)
print has_all_unique_characters_hashset(word_without_duplicates)
print '\nAnswers using an array of ints'
print has_all_unique_characters_array(word_with_duplicates)
print has_all_unique_characters_array(word_without_duplicates)
print '\nAnswers using sort'
print has_all_unique_characters_array_sorting(word_with_duplicates)
print has_all_unique_characters_array_sorting(word_without_duplicates)
# -
# ### Challenge 2: Detect whether two strings are anagrams or not
#
#
# An anagram is a word that contains exactly the same characters as another word (including the number of times each character appears).
# Sorts both strings, compare them. If they are anagrams, once sorted, they must be the same string
# Time Compleixty = 2 * O(N LOG N) to sort both strings + O(N) to compare both of them, leading to O(N LOG N) time complexity
def is_anagram_sorting(w1, w2):
# Basic test: Length must match
if len(w1) is not len(w2):
return False
# Sorting Strings
w1 = sorted(w1)
w2 = sorted(w2)
return w1 == w2
# This solution is faster than the first one, being able to solve it in O(N) time complexity.
# The trick here is to use an array of size 26 (characters of the english alphabet), initialized with all zeroes.
# For each character of the first string, increment the index of the character in the maps array, by 1.
# Do the same for the second string, but decrement it by 1.
# At the end, the array should still have only zeroes in it
def is_anagram_using_map(w1, w2):
# Basic test: Length must match
if len(w1) is not len(w2):
return False
chars_map = [0] * 26
for idx, _ in enumerate(w1):
chars_map[ord(w1[idx]) - 97] += 1
chars_map[ord(w2[idx]) - 97] -= 1
for character in chars_map:
if character is not 0:
return False
return True
# +
w1 = 'elvis'
w2 = 'lives'
w1_n = 'Apple'
w2_n = 'Banana'
print 'Checking anagrams by sorting:'
print is_anagram_sorting(w1,w2)
print is_anagram_sorting(w1_n,w2_n)
print '\nChecking anagrams using characters map:'
print is_anagram_using_map(w1,w2)
print is_anagram_using_map(w1_n,w2_n)
# -
| Notebooks/Data-Structures/Strings And Arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from numpy.linalg import eigvals
import matplotlib
import matplotlib.pyplot as plt
matplotlib.use('Qt5Agg')
# %matplotlib qt5
#
# extend path by location of the dvr package
#
import sys
sys.path.append('/home/thomas/Current_Work/Jolanta-by-dvr/Python_libs')
import dvr
import jolanta
import read_write
amu_to_au=1822.888486192
au2cm=219474.63068
au2eV=27.211386027
Angs2Bohr=1.8897259886
# +
#
# compute DVR of T and V
# then show the density of states
# in a potential + energy-levels plot
#
rmin=0
rmax=30 # grid from 0 to rmax
thresh = 5 # maximum energy for state in the plot in eV
ppB = 10 # grid points per Bohr
nGrid=int((rmax-rmin)*ppB)
print("nGrid = %d" % nGrid)
rs = dvr.DVRGrid(rmin, rmax, nGrid)
Vs = jolanta.Jolanta_3D(rs, a=0.1, b=1.2, c=0.1, l=1)
Ts = dvr.KineticEnergy(1, rmin, rmax, nGrid)
[energy, wf] = dvr.DVRDiag2(nGrid, Ts, Vs)
n_ene=0
for i in range(nGrid):
print("%3d %12.8f au = %12.5f eV" % (i+1, energy[i], energy[i]*au2eV))
n_ene += 1
if energy[i]*au2eV > thresh:
break
# "DVR normalization", sum(wf[:,0]**2)
# this is correct for plotting
c=["orange", "blue"]
#h=float(xmax) / (nGrid+1.0)
scale=150
plt.plot(rs,Vs*au2eV, '-', color="black")
for i in range(n_ene):
plt.plot(rs, scale*wf[:,i]**2+energy[i]*au2eV, '-', color=c[i%len(c)])
plt.ylim(energy[0]*au2eV-1, energy[n_ene-1]*au2eV+1)
plt.xlabel('$r$ [Bohr]')
plt.ylabel('$E$ [eV]')
plt.show()
# +
""" complex diagonalization example """
theta=40.0/180.0*np.pi
print(theta)
Vs = jolanta.Jolanta_3D(rs*np.exp(1j*complex(theta)), a=0.1, b=1.2, c=0.1, l=1)
H_theta = np.exp(-2j*complex(theta)) * Ts + np.diag(Vs)
energies = eigvals(H_theta)
energies.sort()
energies[:10]*au2eV
# -
r2s = np.linspace(0.5,9.5,num=10)
rts = r2s*np.exp(1j*complex(theta))
(V0as, V0bs, V0cs) = jolanta.Jolanta_3D(r2s, a=0.1, b=1.2, c=0.1, l=1, as_parts=True)
(Vas, Vbs, Vcs) = jolanta.Jolanta_3D(rts, a=0.1, b=1.2, c=0.1, l=1, as_parts=True)
Vas/V0as
# +
#
# above theta = 40 deg the lowest Re(E) are basis set
# representation artifacts, and we should either not go there
# or use a better filter for the states to keep
#
# +
#
# complex scaling loop:
#
# start on the real axis (theta=0) and rotate to theta = theta_max
#
# we keep n_keep energies with the lowest real part
#
n_theta=81
n_keep=16
theta_min=0
theta_max=40.0/180.0*np.pi
thetas=np.linspace(theta_min, theta_max, n_theta, endpoint=True)
run_data = np.zeros((n_theta,n_keep), complex) # array used to collect all theta-run data
for i_theta in range(n_theta):
theta=thetas[i_theta]
Vs = jolanta.Jolanta_3D(rs*np.exp(1j*complex(theta)), a=0.1, b=1.2, c=0.1, l=1)
H_theta = np.exp(-2j*complex(theta)) * Ts + np.diag(Vs)
energies = eigvals(H_theta)
energies.sort()
run_data[i_theta,:] = energies[0:n_keep]
print(i_theta+1, end=" ")
if (i_theta+1)%10==0:
print()
run_data *= au2eV
# +
#run_data[0,1:8]
# -
#
# useful piece of the complex plane
# (if unknown, plot all and zoom with matplotlib)
#
plt.cla()
for i in range(0, n_keep):
plt.plot(run_data[:,i].real, run_data[:,i].imag, 'o', color='blue')
plt.xlim(0,5)
plt.ylim(-1,0)
plt.show()
#
#
# two follow ideas:
#
# - at the last theta compute the angles and compare with 2*theta
# if significantly smaller, then resonance
#
# - once a trajectory has five values, use them
# to establish a Pade[2,2]
# then predict for the next theta
#
def follow_nearest(follow, es):
"""
follow the energy closet to e0 from the real axis into the complex plane
es is a table of theta-run data es[i_theta,j_energies]
the algorithm used is simply nearest to the old energy
"""
(n_thetas, n_energies) = es.shape
trajectory = np.zeros(n_thetas,complex)
for j in range(0,n_thetas):
i = np.argmin(abs(es[j,:]-follow))
follow = trajectory[j] = es[j,i]
return trajectory
n_save = n_keep//2
trajectories = np.zeros((n_theta, n_save), complex)
for j in range(n_save):
trajectories[:,j] = follow_nearest(run_data[0,j], run_data)
for i in range(0, n_save):
plt.plot(trajectories[:,i].real, trajectories[:,i].imag, '-')
plt.show()
#
# save n_save trajectories to file
# csv as real and imag
# (at the moment easier than csv with complex)
# also, include no header, because the energies need to be sorted
# into trajectories first
#
fname="complex_scaling_rmax."+str(int(rmax))+"_ppB."+str(ppB)+".csv"
read_write.write_theta_run(fname,thetas,trajectories)
#
# regarding the question of automatization:
# the resonance clearly stands out in CS/DVR
# use to make a loop over the Jolanata parameters
# and map Eres(a,b,c) in a useful range: 0.1 to 8 eV
#
for i in range(n_save):
print(i, np.angle(trajectories[-1,i],deg=True))
res_traj = trajectories[:,2]
# +
def naive_derivative(xs, ys):
""" naive forward or backward derivative """
return (ys[1]-ys[0])/(xs[1]-xs[0])
def central_derivative(xs, ys):
""" central derivative at x[1] """
return (ys[2]-ys[0])/(xs[2]-xs[0])
def five_point_derivative(xs, ys):
""" five-point derivative at x[2] """
""" (-ys[0] + 8*ys[1] - 8*ys[3] + ys[4])/(12*h) """
return (-ys[0] + 8*ys[1] - 8*ys[3] + ys[4])/(xs[4]-xs[0])/3
# -
abs_der = np.zeros(n_theta)
abs_der[0] = abs(naive_derivative(thetas[0:2], res_traj[0:2]))
abs_der[1] = abs(central_derivative(thetas[0:3], res_traj[0:3]))
for k in range(2,n_theta-2):
abs_der[k] = abs(five_point_derivative(thetas[k-2:k+3], res_traj[k-2:k+3]))
abs_der[-2] = abs(naive_derivative(thetas[-3:], res_traj[-3:]))
abs_der[-1] = abs(naive_derivative(thetas[-2:], res_traj[-2:]))
plt.cla()
plt.plot(thetas*180/np.pi, np.log(abs_der))
plt.show()
#
# get a feeling for the stabilitity of the value
#
j_opt = np.argmin(abs_der)
print(j_opt, thetas[j_opt]*180/np.pi)
print(res_traj[j_opt])
print(res_traj[j_opt-1]-res_traj[j_opt])
print(res_traj[j_opt+1]-res_traj[j_opt])
| notebooks/.ipynb_checkpoints/CS-J3D-checkpoint.ipynb |