code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: df0893f56f349688326838aaeea0de204df53a132722cbd565e54b24a8fec5f6
# name: python3
# ---
import comet_ml
from comet_ml import Experiment
import torch_optimizer as opt
# +
import seaborn as sns
import pandas as pd
import time
import cv2
import os
import albumentations
import timm
from pytorch_lightning.loggers import CometLogger
import pickle
from sklearn.model_selection import StratifiedKFold
from pytorch_lightning.callbacks import ModelCheckpoint,EarlyStopping
from torchvision.models import resnet50,resnet18
from sklearn.preprocessing import LabelEncoder,MultiLabelBinarizer
from albumentations.pytorch import ToTensorV2
import json
import torch.nn.functional as F
import numpy as np
import random
import pytorch_lightning as pl
import torch.optim as optim
import torch.nn as nn
from torch.utils.data import Dataset,DataLoader
import torch
from sklearn.utils.class_weight import compute_class_weight
# -
class cfg:
img_size = 512
model_name = "resnet50"
max_epochs = 100
batch_size = 32
classes = np.array(['complex','frog_eye_leaf_spot','powdery_mildew','rust','scab'])
train_root_dir = "preprocessed"
val_root_dir = "train_images"
patience = [5,2]
factor= .1
folds=5
min_lr=1e-8
def seeding(seed=2021):
np.random.seed(seed)
os.environ['PYTHONHASHSEED'] = str(seed)
torch.random.manual_seed(seed)
random.seed(seed)
seeding()
@torch.jit.script
def mish(input):
return input * F.tanh(F.softplus(input))
class Mish(nn.Module):
def __init__(self):
super().__init__()
def forward(self,input):
return mish(input)
def train_augs():
return albumentations.Compose([
albumentations.RandomResizedCrop(cfg.img_size, cfg.img_size, scale=(0.9, 1), p=1),
albumentations.HorizontalFlip(p=0.5),
albumentations.VerticalFlip(p=0.5),
albumentations.ShiftScaleRotate(p=0.5),
albumentations.HueSaturationValue(hue_shift_limit=10, sat_shift_limit=10, val_shift_limit=10, p=0.7),
albumentations.RandomBrightnessContrast(brightness_limit=(-0.2,0.2), contrast_limit=(-0.2, 0.2), p=0.7),
albumentations.CLAHE(clip_limit=(1,4), p=0.5),
albumentations.OneOf([
albumentations.OpticalDistortion(distort_limit=1.0),
albumentations.GridDistortion(num_steps=5, distort_limit=1.),
albumentations.ElasticTransform(alpha=3),
], p=0.2),
albumentations.OneOf([
albumentations.GaussNoise(var_limit=[10, 50]),
albumentations.GaussianBlur(),
albumentations.MotionBlur(),
albumentations.MedianBlur(),
], p=0.2),
albumentations.Resize(cfg.img_size, cfg.img_size),
albumentations.OneOf([
albumentations.JpegCompression(),
albumentations.Downscale(scale_min=0.1, scale_max=0.15),
], p=0.2),
albumentations.IAAPiecewiseAffine(p=0.2),
albumentations.IAASharpen(p=0.2),
albumentations.Cutout(max_h_size=int(cfg.img_size * 0.1), max_w_size=int(cfg.img_size * 0.1), num_holes=5, p=0.5),
albumentations.Normalize(p=1.0),
ToTensorV2()
])
# +
def val_augs():
return albumentations.Compose([
albumentations.Resize(cfg.img_size,cfg.img_size),
albumentations.Normalize(p=1.0),
ToTensorV2()
])
class FGVC(Dataset):
def __init__(self,df,mode='train',transforms=None):
self.transforms = transforms
self.dataf = df
self.mode = mode
def __len__(self):
return len(self.dataf)
def __getitem__(self,idx):
img_name = self.dataf.loc[idx,'image']
if self.mode == 'train':
img = cv2.imread(os.path.join(cfg.train_root_dir,img_name))
elif self.mode =='val':
img = cv2.imread(os.path.join(cfg.val_root_dir,img_name))
#print(img_name)
label = self.dataf.iloc[idx][cfg.classes]
if self.transforms is not None:
img = self.transforms(image=img)['image']
return img,torch.tensor(label)
# -
mis = Mish()
class FGVCNet(pl.LightningModule):
def __init__(self):
super(FGVCNet,self).__init__()
if "resnet" in cfg.model_name:
self.model = eval(cfg.model_name)(pretrained=False)
for params in self.model.parameters():
params.require_grad = True
self.model = nn.Sequential(*list(self.model.children())[:-2])
self.model.add_module("average_pool",nn.AdaptiveAvgPool2d(output_size=(1,1)))
self.model.add_module("Flatten",nn.Flatten())
self.model.add_module("FC1",nn.Linear(2048,1024))
self.model.add_module("ACT1",mis)
self.model.add_module("FC2",nn.Linear(1024,1024))
self.model.add_module("ACT2",mis)
self.model.add_module("CLASSIFIER",nn.Linear(1024,5))
if "efficient" in cfg.model_name:
self.model = timm.create_model(cfg.model_name,pretrained=True)
num_ftrs = self.model.classifier.in_features
self.model.classifier = nn.Linear(num_ftrs,5)
def forward(self,x):
return self.model(x)
def configure_optimizers(self):
op = opt.Ranger(self.parameters(),lr=0.01)
scheduler = {
'scheduler':optim.lr_scheduler.CosineAnnealingWarmRestarts(optimizer=op,T_0=10),
'monitor':'val_f1',
'interval':'epoch',
'frequency':1,
'strict':True
}
self.op = op
self.scheduler = scheduler
return [op],[scheduler]
def training_step(self,batch,batch_idx):
X,y = batch
y_hat = self.model(X)
loss_tr = F.binary_cross_entropy_with_logits(y_hat,y.float())
f1_tr = pl.metrics.functional.f1(y_hat.sigmoid(),y,5)
self.log("TrainLoss",loss_tr,prog_bar=True,on_step=False,on_epoch=True,logger=True)
self.log("TrainF1",f1_tr,prog_bar=True,on_epoch=True,on_step=False,logger=True)
return loss_tr
def validation_step(self,batch,batch_idx):
X,y = batch
y_hat = self.model(X)
loss_val = F.binary_cross_entropy_with_logits(y_hat,y.float())
f1_val = pl.metrics.functional.f1(y_hat.sigmoid(),y,5)
self.log("val_loss",loss_val,prog_bar=True,on_step=False,on_epoch=True,logger=True)
self.log("val_f1",f1_val,prog_bar=True,on_epoch=True,on_step=False,logger=True)
return loss_val
train_data = pd.read_csv("train2.csv")
# + tags=[]
def main():
i=0
comet = CometLogger(api_key="<KEY>",workspace="thirurjst",save_dir='comet',project_name="fgvc8",experiment_name=f"{cfg.model_name}_{cfg.img_size}_Model Fold - {i+1}/5"+"\t"+"Time:"+str(time.time()))
estp = EarlyStopping(monitor="val_f1",mode="max",patience=cfg.patience[0])
if os.path.isdir(f"{cfg.model_name}_{cfg.img_size}_Model_Fold-{i+1}_2"):
print("Already Exists")
else:
os.makedirs(f"{cfg.model_name}_{cfg.img_size}_Model_Fold-{i+1}_2")
mod_ckpt = ModelCheckpoint(
monitor='val_loss',
mode='min',
dirpath=f"{cfg.model_name}_{cfg.img_size}_Model_Fold-{i+1}_2",
filename='Resnet18_512_Checkpoint-ValLoss:{val_loss:.4f}-F1:{val_f1:.4f}',
save_top_k=1,
)
trainer = pl.Trainer(gpus=1,precision=16,max_epochs=cfg.max_epochs,progress_bar_refresh_rate=30,deterministic=True,benchmark=True,callbacks=[mod_ckpt,estp],logger=comet,accumulate_grad_batches=1)
print(f"Initializing model Fold - {i+1}/5")
model = FGVCNet()
print("*** Model Initialization Completed ***")
train_recs = train_data[train_data.folds != i].reset_index(drop=True)
val_recs = train_data[train_data.folds == i].reset_index(drop=True)
train_dataset = FGVC(df=train_recs,mode='train')
val_dataset = FGVC(df=val_recs,mode='val',transforms=val_augs())
train_loader = DataLoader(train_dataset,batch_size=32,num_workers=4)
val_loader = DataLoader(val_dataset,batch_size=32,num_workers=4)
trainer.fit(model,train_loader,val_loader)
print("Uploading Model to Comet")
comet.experiment.log_model(f"Model{i+1}",f"./{cfg.model_name}_{cfg.img_size}_Model_Fold-{i+1}_2")
print("Upload OK...")
comet.experiment.end()
# + tags=[]
main()
# -
| resnet50-training-fgvc8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from qiskit import *
# %config InlineBackend.figure_format = 'svg' # Makes the images look nice
# +
#qr = QuantumRegister(3)
#cr = ClassicalRegister(3)
# -
circuit = QuantumCircuit(3,3)
# %matplotlib inline
# %matplotlib inline
circuit.draw(output='mpl')
circuit.h(0)
circuit.draw(output='mpl')
circuit.cx(0,1)
circuit.draw(output='mpl')
circuit.cx(1,2)
circuit.draw(output='mpl')
circuit.measure([0,1,2], [0,1,2])
circuit.draw(output='mpl')
# Greenberger–Horne–Zeilinger state (GHZ state)
simulator = Aer.get_backend('qasm_simulator')
result = execute(circuit, backend = simulator).result()
from qiskit.tools.visualization import plot_histogram
plot_histogram(result.get_counts(circuit))
from qiskit import IBMQ
IBMQ.save_account()
IBMQ.load_account()
provider = IBMQ.get_provider(hub = 'ibm-q')
qcomp = provider.get_backend('ibmq_vigo')
# + active=""
# import qiskit.tools.jupyter
# %qiskit_job_watcher
# -
job = execute(circuit, backend=qcomp)
from qiskit.tools.monitor import job_monitor
job_monitor(job)
result = job.result()
plot_histogram(result.get_counts(circuit))
def make_ghz3(shots,device):
circuit = QuantumCircuit(3,3)
circuit.h(0)
circuit.cx(0,1)
circuit.cx(1,2)
circuit.measure([0,1,2], [0,1,2])
IBMQ.load_account()
provider = IBMQ.get_provider(hub = 'ibm-q')
qcomp = provider.get_backend(device)
execute(circuit, backend=qcomp, shots=shots) # shots is number of repitions of each circuit
result = job.result()
return result
make_ghz3(1024,'ibmq_ourense')
| assignment-01/AbsoluteWIMPs/Assignment01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Imports
# +
# # !git clone https://github.com/Holstrup/MetricLearning
# import os
# os.chdir("MetricLearning/MetricLearning")
# # !git pull
# -
import sklearn.metrics as sk_metrics
import sklearn.decomposition as sk_decomp
import sklearn.datasets as datasets
from sklearn.model_selection import train_test_split
from sklearn.metrics import confusion_matrix
import numpy as np
import Model
import Plots
import matplotlib.pyplot as plt
import knnClassifier
from scipy.special import softmax
import plot_confusion_matrix as plots
import random
from knnClassifier import calc_mode
import os
os.chdir("../")
# We ignore divisions by 0. We will do that in the chi square distance, but we don't care.
np.seterr(divide='ignore', invalid='ignore')
# ## Hyperparameters
# Underneath you can define the hyperparameters, l, mu and K.
# * $l$ is the margin parameter
# * $\mu$ is a trade-off parameter between the push and pull in the loss function
# * $K$ is the number of target neighbors
# * $D$ is the dimensionality of our data. We choose that arbitrarily
# * $L$ is the linear transformation
#
# L is set to be a diagonal matrix of ones to begin with. However, it would be interesting to experiement with other initial L matrices, since the problem is non-convex. We could also try to implement the constrained non-convex version of LMNN.
l = 0.5
mu = 0.5
K = 5
D = 800
L = softmax(10 * np.eye(D, dtype=float) + 0.001 * D, axis = 1)
alpha = 0.001
# ## Functions
# This block defines two functions that we will use to compute the distances. The first function calculates the distance given two points xi and xj. The second function calculates the distance between some point xi and all the other points X, through the L space. This is very important to note.
# * $chi-square-distance(xi, xj)$ corresponds to (3) in the [Non-linear metric learning paper](http://www.cs.cornell.edu/~kilian/papers/chilmnn.pdf)
# * $distance(xi, X)$ is an extension to $chi-square-distance(xi, xj)$, namely finding the chi square distance from one point xi, to all the other points
# +
def chi_square_distance(xi, xj):
"""
Chi square distance
:param xi: Embedding (1, D)
:param xj: Target Neighbor (1, D)
:return: Distance
"""
return 1 / 2 * np.nansum(np.square(xi - xj) / (xi + xj))
def distance(xi, X, L):
"""
Chi square distance from one point xi, to all other points
:param xi: Embedding (1, D)
:param X: Data (N, D)
:return: Distances (1, N)
"""
N, K = np.shape(X)
Distances = np.zeros(N)
for i in range(N):
Distances[i] = chi_square_distance(L @ xi, L @ X[i, :])
return Distances
# -
# These next functions are calculating the target neighbors and the imposters. Read through the comments, and it should make sense how it is done. One little note: In the target neighbours function, we find the distance through the L space. This is meant as the initial L space. We only find these target neighbours once, and should therefore not update what target neighbors a specific point xi has.
# +
def find_target_neighbors(X, Y, L):
"""
Find target neighbours for all points
:param X: Data Matrix (N, D)
:param Y: Labels (1, N)
:return: TN_lookup_table (N, K)
:return: TN_distance_table (N, K)
"""
global TN_lookup_table
global TN_distance_table
N, _ = np.shape(X)
TN_lookup_table = np.zeros((N, K))
TN_distance_table = np.zeros((N, K))
for i in range(N):
xi = X[i,:]
yi = Y[i]
# Find distance from xi to all other points
TN_Distances = distance(xi, X, L)
TN_Indicies = np.argsort(TN_Distances)
j = k = 0
#Loop to add indicies of target neighbours to lookup table
while j < K:
# if yi and target neighbour have the same label AND it is not the same point
if Y[TN_Indicies[k]] == yi and TN_Indicies[k] != i:
# Add to lookup table and distance table
TN_lookup_table[i, j] = TN_Indicies[k]
TN_distance_table[i, j] = TN_Distances[TN_Indicies[k]]
j += 1
k += 1
TN_lookup_table = TN_lookup_table.astype(int)
return TN_lookup_table, TN_distance_table
#Check if the impostor is within the margin of the target neighbor + marginal distance l
def check(L, xi, xj, xk):
return (chi_square_distance(L @ xi, L @ xj) + l >= chi_square_distance(L @ xi, L @ xk))
# -
# ## LMNN Functions
#
# These next functions, are made from looking at this [Chi square Metric Learning for Nearest Neighbor Classification and Its Analysis paper](https://projet.liris.cnrs.fr/imagine/pub/proceedings/ICPR-2012/media/files/1795.pdf). Specifically, if you scroll to the last page you can see the equations that are implemented below. Here are some notes, that helps when you read it:
# * $L_{\alpha, \beta}$ refers to the alpha'th row and the beta'th column in the L matrix. Same goes for all the other places there is subscript.
# * $\psi_{i, j, \alpha}$ in this paper is the same as $t_{i,j}$ in the Yang paper
# * In this paper they refer to imposters with the letter l (as opposed to k in the other papers)
# * $\xi_{i, j, l}$ I interpret as a check that the point $x_{l}$ is actually an imposter. That should always be the case since, we calculate the imposters this way in the imposters function.
# * $gradient-function$ is the main function that deals with computing the gradient
# * $gradient-element$ sustains $gradient-function$ and corresponds to (26) and (27) in the above stated [paper](https://projet.liris.cnrs.fr/imagine/pub/proceedings/ICPR-2012/media/files/1795.pdf)
# * $outer-loss$ corresponds to (11) - for target neighbors
# * $inner-loss-function$ corresponds to (12) - for impostors
# * $tau-function$ ($\Psi$ in the text) corresponds to (25) in the above stated [paper](https://projet.liris.cnrs.fr/imagine/pub/proceedings/ICPR-2012/media/files/1795.pdf)
#Tau Function
def tau_function(X_Matrix, L_Matrix, i, j, alpha):
N, D = np.shape(X_Matrix)
numerator = 0
denominator = 0
for k in range(D):
numerator += L_Matrix[alpha, k] * (X_Matrix[i, k] - X_Matrix[j, k])
denominator += L_Matrix[alpha, k] * (X_Matrix[i, k] + X_Matrix[j, k])
return numerator / denominator
# +
def gradient_and_loss_function(X, Y, L_Matrix):
D, D = np.shape(L_Matrix)
gradient_matrix = np.zeros((D,D))
for alpha in range(D):
for beta in range(D):
gradient_matrix[alpha, beta], loss = gradient_and_loss_element(X, Y, L_Matrix, alpha, beta)
return gradient_matrix, loss
def gradient_and_loss_element(X_Matrix, Y, L_Matrix, alpha, beta):
global mu
N, _ = np.shape(X_Matrix)
gradient = 0
outer_sum = 0
Inner_sum = 0
loss = 0
for i in range(N):
Pull = 0
for j in TN_lookup_table[i, :]:
tauij = tau_function(X_Matrix, L_Matrix, i, j, alpha)
Lij = 2 * tauij * (X_Matrix[i, beta] - X_Matrix[j, beta]) - (tauij**2) * (X_Matrix[i, beta] + X_Matrix[j, beta])
outer_sum += Lij
for k in range(N):
# We need to update the distance to our target neighbours and compute the max distance
if (check(L, X_Matrix[i], X_Matrix[j], X_Matrix[k]) and (Y[i] != Y[k])):
tauik = tau_function(X_Matrix, L_Matrix, i, k, alpha)
Lik = 2 * tauik * (X_Matrix[i, beta] - X_Matrix[k, beta]) - (tauik**2) * (X_Matrix[i, beta] + X_Matrix[k, beta])
Inner_sum += Lij - Lik
# Calculate loss
loss += (1 - mu) * pullLoss(X_Matrix, L_Matrix, i, j) + mu * pushLoss(X_Matrix, Y, L_Matrix, i, j)
gradient = (1 - mu) * outer_sum + mu * Inner_sum
return gradient, loss
#Loss for pull
def pullLoss(X_Matrix, L_Matrix, i, j):
return chi_square_distance(L_Matrix @ X_Matrix[i], L_Matrix @ X_Matrix[j])
#Loss for push
def pushLoss(X_Matrix, Y, L_Matrix, i, j):
loss = 0
N, _ = np.shape(X_Matrix)
for k in range(N):
if (check(L_Matrix, X_Matrix[i], X_Matrix[j], X_Matrix[k]) and (Y[i] != Y[k])):
loss += max(0, l + chi_square_distance(L_Matrix @ X_Matrix[i], L_Matrix @ X_Matrix[j]) - chi_square_distance(L_Matrix @ X_Matrix[i], L_Matrix @ X_Matrix[k]))
return loss
# -
# ## Separate implementation of the loss function, gradient for pulling and gradient for pushing
# ### Implementation of the loss function
# +
def loss_function(X, Y, L_Matrix):
loss = 0
D, D = np.shape(L_Matrix)
for alpha in range(D):
for beta in range(D):
loss += loss_element(X, Y, L_Matrix, alpha, beta)
return loss
def loss_element(X, Y, L_Matrix, alpha, beta):
loss = 0
global mu
N, _ = np.shape(X)
for i in range(N):
Pull = 0
for j in TN_lookup_table[i, :]:
# Calculate loss
loss += (1 - mu) * pull_loss(X, L_Matrix, i, j) + mu * push_loss(X, Y, L_Matrix, i, j)
return loss
#Loss for pull
def pull_loss(X_Matrix, L_Matrix, i, j):
return chi_square_distance(L_Matrix @ X_Matrix[i], L_Matrix @ X_Matrix[j])
#Loss for push
def push_loss(X_Matrix, Y, L_Matrix, i, j):
loss = 0
N, _ = np.shape(X_Matrix)
for k in range(N):
if (check(L_Matrix, X_Matrix[i], X_Matrix[j], X_Matrix[k]) and (Y[i] != Y[k])):
loss += max(0, l + chi_square_distance(L_Matrix @ X_Matrix[i], L_Matrix @ X_Matrix[j]) - chi_square_distance(L_Matrix @ X_Matrix[i], L_Matrix @ X_Matrix[k]))
return loss
# -
# ### Implementation of the gradient for pulling and pushing
# * $Pull(XMatrix, LMatrix)$ corresponds to (26)
# * $Push(XMatrix, LMatrix, Y)$ corresponds to (27)
# +
### PULL FUNCTIONS
def Pull(X_Matrix, L_Matrix):
D, D = np.shape(L_Matrix)
gradient_matrix = np.zeros((D,D))
for alpha in range(D):
for beta in range(D):
gradient_matrix[alpha, beta] = Pull_matrix_element(X_Matrix, L_Matrix, alpha, beta)
return gradient_matrix
def Pull_matrix_element(X_Matrix, L_Matrix, alpha, beta):
N, D = np.shape(X_Matrix)
gradient = 0
for i in range(N):
for j in TN_lookup_table[i, :]:
tau = tau_function(X_Matrix, L_Matrix, i, j, alpha)
gradient += 2 * tau * (X_Matrix[i, beta] - X_Matrix[j, beta]) - (tau**2) * (X_Matrix[i, beta] + X_Matrix[j, beta])
return gradient
# +
### PUSH FUNCTIONS
def Push(X_Matrix, L_Matrix, Y):
D, D = np.shape(L_Matrix)
gradient_matrix = np.zeros((D,D))
for alpha in range(D):
for beta in range(D):
gradient_matrix[alpha, beta] = Push_matrix_element(X_Matrix, L_Matrix, Y, alpha, beta)
return gradient_matrix
def Push_matrix_element(X_Matrix, L_Matrix, Y, alpha, beta):
N, D = np.shape(X_Matrix)
gradient = 0
for i in range(N):
for j in TN_lookup_table[i, :]:
for k in range(N):
if (check(L, X_Matrix[i], X_Matrix[j], X_Matrix[k]) and (Y[i] != Y[k])):
tauij = tau_function(X_Matrix, L_Matrix, i, j, alpha)
tauik = tau_function(X_Matrix, L_Matrix, i, k, alpha)
Lij = 2 * tauij * (X_Matrix[i, beta] - X_Matrix[j, beta]) - (tauij**2) * (X_Matrix[i, beta] + X_Matrix[j, beta])
Lik = 2 * tauik * (X_Matrix[i, beta] - X_Matrix[k, beta]) - (tauik**2) * (X_Matrix[i, beta] + X_Matrix[k, beta])
gradient += Lij - Lik
return gradient
# -
# The update for the gradient is done based on the predefined learning rate.
# The gradient should not be negative and its values should sum up to 1. Thus, we have made use of the softmax function for updating the gradient.
def update_step(G):
"""
Update L
:param G: Computed gradient for a given iteration
"""
global L
new_L = L - alpha * G
L = softmax(new_L, axis = 1)
# + [markdown] heading_collapsed=true
# ## Amazon dataset
# + hidden=true
folder_path = "Data_Amazon/"
Data_Matrix = np.zeros(800)
label = []
for _, dirs, _ in os.walk(folder_path, topdown=True):
for directory in dirs:
sub_folder_path = os.path.join(folder_path, directory)
for _, _, files in os.walk(sub_folder_path):
for name in files:
if name != '.DS_Store':
2+2
vec = scipy.io.loadmat(os.path.join(sub_folder_path, name))['histogram']
vec = vec / np.sum(vec)
label.append(name)
Data_Matrix = np.vstack((Data_Matrix, vec))
Data_Matrix = Data_Matrix[1:, :]
# + [markdown] heading_collapsed=true
# ## Dataset - IRIS
# + [markdown] hidden=true
# Shape of IRIS dataset: (150,4)
# + hidden=true
iris = datasets.load_iris()
X_train, X_test, y_train, y_test = train_test_split(iris.data, iris.target, test_size=0.3, random_state=0)
# Normalize data
X_train = X_train / np.sum(X_train, axis = 1)[:,None]
X_test = X_test / np.sum(X_test, axis = 1)[:,None]
# + hidden=true
mu = 0.5
K = 3
_, D = np.shape(X_train)
L = np.eye(D, dtype=float)
alpha = 0.001
# + hidden=true
TN_lookup_table, TN_distance_table = find_target_neighbors(X_train, y_train, L)
l = np.median(TN_distance_table)
# -
# ## Dataset - Fruits
# +
X_train, y_train = Model.get_data('data.db')
X_test, y_test = Model.get_data('data_test.db')
s_score = sk_metrics.silhouette_score(X_train.T, y_train, metric='euclidean')
print("Shape of trained embeddings is: {}".format(np.shape(X_train)))
print("Shape of trained labels is: {} ".format(np.shape(y_train)))
print("Shape of test embeddings is: {}".format(np.shape(X_test)))
print("Shape of test labels is: {} ".format(np.shape(y_test)))
print("Silhouette Score of data is: {}".format(s_score))
# +
#By increasing the number of components we deal with the check-function issue (To be fixed)
pca = sk_decomp.PCA(n_components=10)
pca.fit(X_train.T)
X_train = pca.transform(X_train.T)
X_train = abs(X_train / np.linalg.norm(X_train))
X_test = pca.transform(X_test.T)
X_test = abs(X_test / np.linalg.norm(X_test))
s_score = sk_metrics.silhouette_score(X_train, y_train, metric='euclidean')
print("Explained variance of low dimensional data is: {}".format(sum(pca.explained_variance_ratio_)))
print("Silhouette Score of data is: {}".format(s_score))
# -
#It's very important to set the margin very low as the data points are quite densly packed
mu = 0.7
K = 3
alpha = 0.001
_, D = np.shape(X_train)
L = np.eye(D, dtype=float)
TN_lookup_table, TN_distance_table = find_target_neighbors(X_train, y_train, L)
l = 0.0001
# ### Histograms using the initial linear transformation matrix L
# +
new_X = (L @ X_train.T).T
plot_items = new_X[35:50:5,:]
plot_labels = y_train[35:50:5]
Plots.plot_some_data(plot_items, plot_labels)
# -
# ## Run
# +
ITERATIONS = 5
lossList = []
#X_train = Data_Matrix
#y_train = label
for i in range(ITERATIONS):
print(i)
# Gt = (1 - mu) * Pull(X_train, L) + mu * Push(X_train, L, y_train)
# loss = loss_function(X_train, y_train, L)
Gt, loss = gradient_and_loss_function(X_train, y_train, L)
lossList.append(loss)
update_step(Gt)
print(L)
print(loss)
print("\n")
plt.plot(lossList, '-')
plt.show()
# -
# As expected, the loss follows an exponentially decreasing function. Having as component the loss from pushing the impostors and pulling the target neighbors, this means that the target neighbors are pulled closer and the impostors pushed away.
# ### Plot of the linear trasformation matrix L
Plots.plot_kernel(L)
# ### Histograms using the updated linear transformation matrix L
#new_X = X_train
new_X = (L @ X_train.T).T
plot_items = new_X[10:20:4,:]
plot_labels = y_train[10:20:4]
Plots.plot_some_data(plot_items, plot_labels)
new_X[10:20:4,:]
# ## Testing
# Further, in order to check the increase in accuracy, we have implemented a knnClassifier
# +
prediction = []
for i in range(len(X_test)):
prediction.append(random.choice(calc_mode(knnClassifier.knn(X_test[i], X_train, y_train, L, 3))))
confusion_matrix = plots.plot_confusion_matrix(y_test, prediction, normalize=False)
# -
| MetricLearning/LMNN2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
## Figures adapted from: http://sociograph.blogspot.com/2012/11/visualizing-adjacency-matrices-in-python.html
# +
from __future__ import division
import os
import igraph
import numpy as np
import pandas as pd
from matplotlib.backends.backend_pdf import PdfPages
from scipy.stats.kde import gaussian_kde
from matplotlib.ticker import FixedLocator,LinearLocator,MultipleLocator, FormatStrFormatter
from mpl_toolkits.axes_grid1 import Grid
import networkx as nx
from matplotlib import pyplot, patches
import numpy as np
from collections import defaultdict
# relevant python libraries
execfile('../functions/python_libraries.py')
execfile('../functions/create_oSBM_class_specific_alternative.py')
execfile('../functions/compute_homophily.py')
execfile('../functions/compute_monophily.py')
# -
def draw_adjacency_matrix_modified(adjacency_matrix, plt_name=None,
partitions=[], colors=[],
partition_indicator = False):
plt_name.imshow(adjacency_matrix,
cmap="binary",
interpolation="nearest")
plt_name.spines["right"].set_linewidth(0.5)
plt_name.spines["left"].set_linewidth(0.5)
plt_name.spines["bottom"].set_linewidth(0.5)
plt_name.spines["top"].set_linewidth(0.5)
plt_name.spines["right"].set_color('lightgray')
plt_name.spines["left"].set_color('lightgray')
plt_name.spines["bottom"].set_color('lightgray')
plt_name.spines["top"].set_color('lightgray')
if partition_indicator:
assert len(partitions) == len(colors)
for partition, color in zip(partitions, colors):
current_idx = 0
for module in partition:
plt_name.add_patch(patches.Rectangle((current_idx, current_idx),
len(module), # Width
len(module), # Height
facecolor="none",
edgecolor=color,
linewidth="1"))
current_idx += len(module)
# +
def assignmentArray_to_lists(assignment_array):
by_attribute_value = defaultdict(list)
for node_index, attribute_value in enumerate(assignment_array):
by_attribute_value[attribute_value].append(node_index)
return by_attribute_value.values()
# -
test = pd.read_csv('../../figures/toyA.csv', header=-1)
test.head()
adj = np.matrix(np.nan_to_num(test))
np.shape(adj)
print np.mean(adj!=adj.T)
y_toy = np.array([1,1,1,1,1,1,2,2,2,2,2,2])
from __future__ import division
print len(y_toy)
print np.shape(adj)
# %matplotlib inline
plt.hist(adj[y_toy==1,:] * np.matrix(y_toy==1).T/(np.sum(adj[y_toy==1],1)),
normed=True)
plt.hist(adj[y_toy==2,:] * np.matrix(y_toy==2).T/(np.sum(adj[y_toy==2],1)),
normed=True)
plt.show()
homophily_index_Jackson_alternative(adj, y_toy)
monophily_index_overdispersion_Williams(adj, y_toy)
adj2 = np.matrix(adj)**2
adj2[range(adj2.shape[0]),range(adj2.shape[0])]=0
adj2 = (adj2 >=1)+0
y_toy_assignment = y_toy.copy()
y_toy_lists = assignmentArray_to_lists(y_toy_assignment)
# +
# %matplotlib inline
f, (ax1, ax2) = plt.subplots(1, 2,
sharey=False, sharex=False,
figsize=(8, 3))
draw_adjacency_matrix_modified(adj,#[idx_amherst,:][idx_amherst,:], #np.concatenate([idx1,idx2]),
partitions=[y_toy_lists],
colors=["red"],
plt_name=ax1,
partition_indicator=False)
ax1.set_xticklabels([])
ax1.set_yticklabels([])
ax1.set_title('$A$')
ax1.axvline(5.5, color='red')
ax1.axhline(5.5, color='red')
draw_adjacency_matrix_modified(adj2,#[idx_amherst,:][:,idx_amherst], #np.concatenate([idx1,idx2]),
partitions=[y_toy_lists],
colors=["red"],
plt_name=ax2,
partition_indicator=False)
ax2.set_xticklabels([])
ax2.set_yticklabels([])
ax2.axvline(5.5, color='red')
ax2.axhline(5.5, color='red')
ax2.set_title('$A^2$')
plt.tight_layout()
#plt.show()
pp = PdfPages('../../figures/figure1b_toy_example.pdf')
pp.savefig(dpi = 300)
pp.close()
# +
## khop spot-check
k_hop = np.array([1,2,3,4,5])
class_values = np.unique(y_toy)
prop_same_array = []
for k in k_hop:
print k
adj_amherst_k= np.matrix(adj)**k
adj_amherst_k[range(adj_amherst_k.shape[0]),range(adj_amherst_k.shape[0])]=0 ## remove self-loops
mv_g1 = (adj_amherst_k[y_toy==class_values[0],:] * np.matrix((y_toy==class_values[0])+0).T)/np.sum(adj_amherst_k[y_toy==class_values[0],:],1)
mv_g2 = (adj_amherst_k[y_toy==class_values[1],:] * np.matrix((y_toy==class_values[1])+0).T)/np.sum(adj_amherst_k[y_toy==class_values[1],:],1)
proportion_majority_same = np.mean(np.concatenate((np.array(mv_g1).T[0]>np.mean(y_toy==class_values[0]),
np.array(mv_g2).T[0]>np.mean(y_toy==class_values[1]))))
prop_same_array.append( proportion_majority_same)
print prop_same_array
# -
| code/0_oSBM/Fig 1b.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Kvaibhav1997/Cancer-Prediction/blob/main/Vaccination_Tweets.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="BtBpNF60lpnN"
import numpy as np
import pandas as pd
import string
import seaborn as sns
import matplotlib.pyplot as plt
from datetime import date
import plotly.express as px
from collections import Counter
from textblob import TextBlob
from wordcloud import WordCloud
# + colab={"base_uri": "https://localhost:8080/", "height": 513} id="YxxZfHoPnREB" outputId="0ede5c8a-b653-413c-97df-03122a2fb501"
dataset=pd.read_csv('vaccination_tweets.csv')
dataset.head()
# + colab={"base_uri": "https://localhost:8080/"} id="DT94HtxcnggX" outputId="e4a8ddbf-009e-4271-c763-5263c92e0ea1"
dataset.columns
# + colab={"base_uri": "https://localhost:8080/"} id="BnrIHAfOnoxc" outputId="651d4e41-4ccb-4f33-8de9-7cf31bcd9253"
dataset.dtypes
# + colab={"base_uri": "https://localhost:8080/"} id="IGoGgk70n8MD" outputId="b698345a-0492-4a30-d440-607ce5a87e51"
dataset.info()
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="Bdi5lsKsoRRB" outputId="f88db369-1744-4247-e6bb-f9c18e4373df"
dataset.describe()
# + colab={"base_uri": "https://localhost:8080/"} id="qBBX22CX9qXS" outputId="2e4eabd5-18d4-4c4d-b355-260f64d77082"
dataset.shape
# + colab={"base_uri": "https://localhost:8080/"} id="SSvbvwNpnuyQ" outputId="2743d5e2-da77-498e-86c3-8bc8bbe1a523"
dataset.isnull().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="dB-wrfWBJex8" outputId="3fd84768-636f-412b-8b84-5a3d5111f999"
dataset['user_verified'].value_counts()
# + [markdown] id="9Cb9ioIw_c2O"
# # Data Manipulation
# + id="_AuAZnir_gpF"
#verified account or not
dataset['user_verified']=dataset['user_verified'].apply(lambda x:'verified' if x==True else 'not_verified')
# + id="FEfywgW9_6NS"
#account age
from datetime import date
dataset['today']=date.today()
dataset['user_created']=pd.to_datetime(dataset['user_created']).dt.year
dataset['today']=pd.to_datetime(dataset['today'])
dataset['today']=dataset['today'].dt.year
dataset['acc_age']= dataset['today']-dataset['user_created']
# + id="M107HhwXAR_Q"
#converting date to date format
dataset['date']=pd.to_datetime(dataset['date'])
# + colab={"base_uri": "https://localhost:8080/"} id="c_Lra4xQA46O" outputId="b30313d2-c681-43ee-f448-b25f11cf8c1c"
L = ['year', 'month', 'day', 'dayofweek', 'dayofyear', 'weekofyear', 'quarter']
dataset = dataset.join(pd.concat((getattr(dataset['date'].dt, i).rename(i) for i in L), axis=1))
# + id="7hK_YfZvBG4S"
#total enagement
dataset['total_engagement']=dataset['retweets']+dataset['favorites']
# + colab={"base_uri": "https://localhost:8080/"} id="qyp90J5YCiwZ" outputId="8bb8f6c0-b5cc-411c-f5ed-b0bbaa71da83"
dataset['tweet_lenght']=dataset['text'].apply(lambda x:len(x))
dataset['tweet_lenght'].describe()
# + colab={"base_uri": "https://localhost:8080/"} id="Uc3pc_lsBtsC" outputId="18a08133-0276-47a8-cc67-9f84b5f65f24"
#The tweet length can be no longer than 280 characters so let's check tweets length
dataset['tweet_lenght']=dataset['text'].apply(lambda x:len(x))
dataset['tweet_lenght'].describe()
# + id="myUg5zjgCzTL"
#tweet length cases
dataset['tweet_length']=dataset['text'].apply(lambda x:'short' if len(x)<=130 else 'long')
# + id="L5V9mFEEDgRD"
#fix country location
loc_dataset = dataset['user_location'].str.split(',',expand=True)
loc_dataset=loc_dataset.rename(columns={0:'fst_loc',1:'snd_loc'})
# + colab={"base_uri": "https://localhost:8080/"} id="fmkTFRe3DzO7" outputId="81caa5f0-9b40-4401-a2d2-b3e9620dc141"
#fixing states with country shortcut
# Remove Spaces
loc_dataset['snd_loc'] = loc_dataset['snd_loc'].str.strip()
# Rename States
state_fix = {'Ontario': 'Canada','United Arab Emirates': 'UAE','TX': 'USA','NY': 'USA'
,'FL': 'USA','England': 'UK','Watford': 'UK','GA': 'USA','IL': 'USA'
,'Alberta': 'Canada','WA': 'USA','NC': 'USA','British Columbia': 'Canada','MA': 'USA','ON':'Canada'
,'OH':'USA','MO':'USA','AZ':'USA','NJ':'USA','CA':'USA','DC':'USA','AB':'USA','PA':'USA','SC':'USA'
,'VA':'USA','TN':'USA','New York':'USA','Dubai':'UAE','CO':'USA'}
loc_dataset = loc_dataset.replace({"snd_loc": state_fix})
loc_dataset['snd_loc'].value_counts()[:10]
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="G7D4wOzaHZjy" outputId="66f73540-35a8-4d0c-faea-58f0daa62828"
#engament
line = dataset.groupby('date',as_index=False).agg({'total_engagement':'sum'})
fig = go.Figure()
fig.add_trace(go.Scatter(x=line.date, y=line.total_engagement,
mode='lines+markers'))
# + id="0zqoNuyf8Pvn"
dataset['user_location'] = dataset['user_location'].fillna('NaN')
dataset['source'] = dataset['source'].fillna('NaN')
dataset['hashtags'] = dataset['hashtags'].fillna('NaN')
locations = pd.Series(Counter(dataset['user_location'])).sort_values(ascending=False)
locations = locations.drop('NaN')[locations>5]
source = Counter(dataset['source'])
source = pd.Series(source).sort_values(ascending=False)
verified = Counter(dataset['user_verified'])
verified = pd.Series(verified).sort_values(ascending=False)
name = Counter(dataset['user_name'])
name = pd.Series(name).sort_values(ascending=False)
name = name[name>4]
hashtags = []
for i in dataset['hashtags']:
for j in i.translate(str.maketrans('', '', string.punctuation)).split(' '):
hashtags.append(j)
hashtags = Counter(hashtags)
hashtags = pd.Series(hashtags).sort_values(ascending=False)
hashtags = hashtags[hashtags>8]
followers = dataset.sort_values(by='user_followers', ascending=False)
followers_i = followers['user_name'].drop_duplicates().keys()[:15]
friends = dataset.sort_values(by='user_friends', ascending=False)[:15]
friends_i = friends['user_name'].drop_duplicates().keys()
favourites = dataset.sort_values(by='user_favourites', ascending=False)
favourites_i = favourites['user_name'].drop_duplicates().keys()[:15]
# + [markdown] id="kSCLLNBZ9qEO"
# # Visualisation of Data
# + [markdown] id="mc61vuY-9wy5"
# ## Bar Plot
# + id="JRYVfxbx92vb"
def barplot(x, y, title, xlabel, ylabel, capsize=12, fontsize=12, labelsize=9,
palette='twilight', rotation=0, figsize=(13, 13), bartext_size=15):
fig, ax = plt.subplots(1, 1, figsize=figsize)
bars = sns.barplot(x, y, palette=palette, capsize=capsize)
for bar in bars.patches:
bars.annotate(format(bar.get_height(), '.0f'),
(bar.get_x()+bar.get_width()/2., bar.get_height()), ha='center', va='bottom',
size=bartext_size)
plt.title(title, fontsize=fontsize)
plt.xlabel(xlabel, fontsize=fontsize)
plt.ylabel(ylabel, fontsize=fontsize)
plt.xticks(rotation=rotation)
plt.tick_params(axis='x', labelsize=labelsize)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="4DQ_rs0R_UK6" outputId="9ef6d563-da39-4a66-cd83-f1658f66dfc8"
barplot(locations.keys(), locations, title='Tweets per location', xlabel='Location',
ylabel='Number of tweets', rotation=90, bartext_size=10)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="MiF2WY5AFn4U" outputId="0c0f5f4e-5866-4c17-b33a-f82f3fd053d9"
barplot(source.keys(), source, title='Tweets per source', xlabel='Source',
ylabel='Number of tweets', palette='Accent', rotation=90)
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="ZZt9Q1cXIOhh" outputId="034c90dd-5e3b-48b6-ce10-be3f4950ec6b"
barplot(name.keys(), name, 'People who tweeted', 'Username', 'Number of tweets', rotation=90,
palette='autumn', bartext_size=11)
# + colab={"base_uri": "https://localhost:8080/", "height": 962} id="hCv4ZfYoIQuX" outputId="fb84f064-5097-4473-8d02-5a8892760d5e"
barplot(hashtags.keys(), hashtags, 'Hashtags for tweets', 'Hashtags', 'Number of tweets',
rotation=90, bartext_size=10)
# + colab={"base_uri": "https://localhost:8080/", "height": 875} id="ID9gVI2cIVPO" outputId="b2141bc2-6d69-48f4-f68f-5580ecedc378"
barplot(verified.keys(), verified, 'Verified twitter acounts',
'Whether an account is verified or not', 'Number of accounts')
# + [markdown] id="YqZf9pJQIjxq"
# ## Pie Plots
# + id="T_hsWz-BF4vV"
def piechart(x, y, title, figsize=(10, 13)):
fig, ax = plt.subplots(1, 1, figsize=figsize)
plt.pie(x, labels=y, autopct=lambda p:f'{p:.2f}%')
plt.title(title)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 605} id="By375jMOIjOP" outputId="dd9a278c-43a6-40a9-d5e2-fdc79db70e7d"
piechart(followers['user_followers'][followers_i], followers['user_name'][followers_i],'Followers per source')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="y4vBiFBQIwt5" outputId="08963581-204f-4d88-b893-29a7c64ef3c1"
piechart(friends['user_friends'][friends_i], friends['user_name'][friends_i],'Number of friends per source')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="gXx8RgMrI0Fw" outputId="901de6e8-ec9a-444f-a122-8dc8c0125172"
piechart(favourites['user_favourites'][favourites_i], favourites['user_name'][favourites_i],'Amount of favourites per source')
# + [markdown] id="BYcGj2HkI6GV"
# ## Line Graphs
# + id="KD_hQrb2JKif"
def time_plot(time, title, x, y):
count = Counter(time)
count = pd.Series(dict(zip(count.values(), [int(i) for i in count.keys()]))).sort_values()
fig, ax = plt.subplots(1, 1, figsize=(13, 10))
ax.plot([str(i) for i in count], count.keys())
ax.set_title(title)
ax.set_xlabel(x)
ax.set_ylabel(y)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 621} id="Tt07oaLeI4Be" outputId="2f52b2ae-395a-4d15-8845-56772ff82652"
time_plot([str(i)[8:10] for i in dataset['date']], 'Number of tweets over the days','Day in December', 'Number of tweets')
# + colab={"base_uri": "https://localhost:8080/", "height": 621} id="ebM97Bz0JP52" outputId="32a95ec3-9528-4493-d6f0-2f5ad2743fdc"
time_plot([str(i)[11:13] for i in dataset['date']], 'Number of tweets over the hours','Hour when tweet was released', 'Number of tweets')
# + [markdown] id="AdAWjG-2My67"
# ## Heatmap
# + colab={"base_uri": "https://localhost:8080/", "height": 531} id="09NMcGITMr1B" outputId="79a9258d-3847-43c1-edc3-e543203b8f82"
corr=dataset.corr()
plt.figure(figsize=(10,7))
sns.heatmap(corr,annot=True)
# + [markdown] id="3sSM14XKNExh"
# ## Countplot
# + colab={"base_uri": "https://localhost:8080/", "height": 443} id="63OLJoagNAcA" outputId="60aeb29a-d6c1-4536-dbdb-78128f9189af"
dataset['tweet_length']=dataset['text'].apply(lambda x:'short' if len(x)<=130 else 'long')
plt.figure(figsize=(7,7))
sns.countplot(x='tweet_length',data=dataset);
# + colab={"base_uri": "https://localhost:8080/", "height": 669} id="lchvGjjVEqZc" outputId="fc71274c-3519-44a9-fa75-861864654b37"
df3=pd.DataFrame(loc_dataset['snd_loc'].value_counts()[:20]).reset_index()
df3
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="SwwxhHTGEdrS" outputId="af25966f-9708-4305-a2ea-688808c432f9"
fig = px.choropleth(df3, locations = df3['index'],
color = df3['snd_loc'],locationmode='country names',hover_name = df3['snd_loc'],
color_continuous_scale = px.colors.sequential.Inferno)
fig.update_layout(title='Sales tracking')
fig.show()
# + [markdown] id="UZygSToZ03Ry"
# # Sentiment Analysis
# + id="DtaITx3k1ZCf"
tweets = dataset['text']
# + id="skS45NNe2Xug"
all_sentences = []
for word in tweets:
all_sentences.append(word)
all_sentences
lines = list()
for line in all_sentences:
words = line.split()
for w in words:
lines.append(w)
# + id="vLC9rJXn3LCa"
import re
lines = [re.sub(r'[^A-Za-z0-9]+', '', x) for x in lines]
lines
lines2 = []
for word in lines:
if word != '':
lines2.append(word)
# + colab={"base_uri": "https://localhost:8080/"} id="pjD1hUhp022H" outputId="92cdb296-1aa7-49d4-d295-886d3ec400c9"
features=tweets.values
features
# + id="GzGEzb7a3DFO"
processed_features = []
for sentence in range(0, len(features)):
# Remove all the Http: urls
processed_feature = re.sub('(https?://\S+)', '', str(features[sentence]))
# Remove all the special characters
processed_feature = re.sub(r'\W', ' ', processed_feature)
# Remove all single characters
processed_feature= re.sub(r'\s+[a-zA-Z]\s+', ' ', processed_feature)
# Remove single characters from the start
processed_feature = re.sub(r'\^[a-zA-Z]\s+', ' ', processed_feature)
# Substituting multiple spaces with single space
processed_feature = re.sub(r'\s+', ' ', processed_feature, flags=re.I)
# Removing prefixed 'b'
processed_feature = re.sub(r'^b\s+', '', processed_feature)
# Converting to Lowercase
processed_feature = processed_feature.lower()
processed_features.append(processed_feature)
# + colab={"base_uri": "https://localhost:8080/"} id="VScSlnPV3TeS" outputId="98e44cfd-f913-4b23-9c7a-a90dc197f711"
features[:5]
# + colab={"base_uri": "https://localhost:8080/"} id="Ayu74FRI3XAj" outputId="3dd12f79-c944-49a9-fb02-db52b6774ea6"
processed_features[:5]
# + [markdown] id="z3hpACvO3h6k"
# Adding Subjectivity and Polarity
# + id="F_sfZ3wl5HpM"
ds=pd.DataFrame()
ds['Tweets']=processed_features
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="IMBmD01B5TSb" outputId="81ee8e9b-b468-41c4-bfa0-80edcad08b22"
# Create a function to get the subjectivity
def getSubjectivity(text):
return TextBlob(text).sentiment.subjectivity
# Create a function to get the polarity
def getPolarity(text):
return TextBlob(text).sentiment.polarity
# Create two new columns 'Subjectivity' & 'Polarity'
ds['Subjectivity'] = ds['Tweets'].apply(getSubjectivity)
ds['Polarity'] = ds['Tweets'].apply(getPolarity)
# Show the new dataframe with columns 'Subjectivity' & 'Polarity'
ds
# + [markdown] id="FwiAYrAk5g7k"
# Creating Sentiment Analysis
# + colab={"base_uri": "https://localhost:8080/", "height": 419} id="lMyJa5G1680-" outputId="8bc379e9-184d-46de-e09b-82ac27198f17"
# Create a function to compute negative (-1), neutral (0) and positive (+1) analysis
def getAnalysis(score):
if score < 0:
return 'Negative'
elif score == 0:
return 'Neutral'
else:
return 'Positive'
ds['Analysis'] = ds['Polarity'].apply(getAnalysis)
ds
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="V5P1E6EV7DvV" outputId="a005a7d4-c998-4f43-e54e-574022f059ab"
Neutral = len(ds[ds['Analysis']=='Neutral'])
Negative = len(ds[ds['Analysis']=='Negative'])
Positive = len(ds[ds['Analysis']=='Positive'])
labels = ['Negative','Positive','Neutral']
values = [Negative,Positive,Neutral]
#====
import plotly.graph_objects as go
colors = ['darkred','green', 'darkblue' ]
fig = go.Figure(data=[go.Pie(labels=labels,
values=values)])
fig.update_traces(hoverinfo='label+percent', textinfo='percent', textfont_size=20,textposition='inside',
marker=dict(colors=colors, line=dict(color='black', width=1)))
fig.show()
| Vaccination_Tweets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.8 ('base')
# language: python
# name: python3
# ---
from bs4 import BeautifulSoup as bs
import requests
from splinter import Browser
from webdriver_manager.chrome import ChromeDriverManager
import pandas as pd
import time
# ### NASA Mars News
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
url= "https://redplanetscience.com/"
browser.visit(url)
time.sleep(3)
# +
html = browser.html
soup = bs(html,'html.parser')
news = soup.find_all('div',class_='list_text')
latest_title = []
latest_body = []
for n in news:
title = n.find('div',class_='content_title')
paragraph = n.find('div',class_='article_teaser_body')
latest_title.append(title)
latest_body.append(paragraph)
# -
# Latest Title and Paragraph
latest_title1 = latest_title[0].text
latest_body1 = latest_body[0].text
print(f'Latest Title: {latest_title1}')
print(f'Latest Paragraph: {latest_body1}')
# ### JPL Mars Space Images - Featured Image
url1= "https://spaceimages-mars.com/"
browser.visit(url1)
# +
# Use splinter to navigate the site and find the image url for the
# current Featured Mars Image and assign the url string to a variable called
# `featured_image_url`.
# Make sure to find the image url to the full size `.jpg` image.Make sure to save a complete url string for this image.
html = browser.html
soup = bs(html, 'html.parser')
# -
featured_image_url = url1 + soup.find('img', class_='headerimage fade-in')['src']
print(f'Featured Image URL: {featured_image_url}')
# ### Mars Facts
url = "https://galaxyfacts-mars.com"
tables = pd.read_html(url)
tables
df = tables[1]
df
html_table = df.to_html()
html_table
html_table.replace('\n','')
print(html_table)
# ### Mars Hemispheres
url = "https://marshemispheres.com/"
browser.visit(url)
html = browser.html
soup = bs(html, 'html.parser')
hemisphere_items = soup.find_all('div', class_='item')
hemisphere_image_urls = []
# +
# Save both the image url string for the full resolution hemisphere image,
# and the Hemisphere title containing the hemisphere name.
# Use a Python dictionary to store the data using the keys `img_url` and `title`.
# Append the dictionary with the image url string and the hemisphere title to a list.
# This list will contain one dictionary for each hemisphere.
# -
for item in hemisphere_items:
try:
# Hemisphere title
title = item.find('div',class_='description').h3.text
# hemisphere_title.append(title)
# print(hemisphere_title)
# Hemisphere Image URL
thumbImage_url = item.find('a',class_='itemLink product-item')['href']
# print(thumbImage_url)
# Visit each thumbnail link
browser.visit(url + thumbImage_url)
time.sleep(3)
html = browser.html
soup = bs(html, 'html.parser')
image_url = url + soup.find('img', class_='wide-image')['src']
#Append
hemisphere_image_urls.append({"Title" : title, "img_url" : image_url})
except Exception as e:
print(e)
print(hemisphere_image_urls)
browser.quit()
| mission_to_mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Defining Multiple Subdomains
#
# One of the features in OpenPNM is the ability to model heterogeneous materials by applying different pore-scale models to different regions. This is done by (a) creating a unique **Geometry** object for each region (i.e. small pores vs big pores) and (b) creating unique **Physics** object for each region as well (i.e. Knudsen diffusion vs Fickian diffusion). One consequence of this segregation of properties is that a *single* array containing values for all locations in the domain does not exist. OpenPNM offers a shortcut for this, known as ``interleave_data``, which happens *automatically*, and makes it possible to query **Geometry** properties via the **Network** object, and **Physics** properties from the associated **Phase** object:
#
# Let's demonstrate this by creating a network and assigning two separate geometries to each half of the network:
import openpnm as op
pn = op.network.Cubic([5, 5, 5])
geo1 = op.geometry.GenericGeometry(network=pn, pores=range(0, 75),
throats=range(0, 150))
geo2 = op.geometry.GenericGeometry(network=pn, pores=range(75, 125),
throats=range(150, 300))
geo1['pore.diameter'] = 1.0
geo2['pore.diameter'] = 0.1
# Each of the Geometry objects has a 'pore.diameter' array with different values. To obtain a single array of 'pore.diameter' with values in the correct locations, we can use the Network as follows:
Dp = pn['pore.diameter']
print(Dp[70:80])
# As can be seen, the 'pore.diameter' array contains values from both Geometry objects, and they are in their correction locations in terms of the domain number system. This is referred to as ``interleave_data``. It also works to obtain Physics values via their associated Phase object.
#
# Interleaving of data also works in the reverse direction, so that data only present on the network can be accessed via the Geometry objects:
coords = geo1['pore.coords']
print(coords[0:3])
# Finally, ``interleave_data`` works between objects of the same type, so that if 'pore.volume' is present on one but not another Geometry object, you will get an array of NaNs when asking for it on the object that does not have it:
geo1['pore.volume'] = 3.0
print(geo2['pore.volume'][:5])
# ### Points to Note
#
# * Data **cannot** be written in this way, so that you cannot write 'pore.diameter' values to the Network if 'pore.diameter is already present on a Geometry (e.g. pn['pore.diameter'] = 2.0 will result in an error)
# * Interleaving data is automatically attempted if the requested key is not found. For instance, when you request ``pn['pore.diameter']`` it is not found, so a search is made of the associated Geometry objects and if found an array is built.
# * If an array named 'pore.foo' is already present on the Network or Phase, it cannot be created on a Geometry or Physics, resepctively, since this would break the automated ``interleave_data`` mechanism, which searches for arrays called 'pore.foo' on all associated objects
| examples/tutorials/defining_multiple_subdomains.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Tutorial 4: Scenario database
#
# In this tutorial, we will cover the following items:
#
# 1. Loading and saving the database
# 2. Obtaining objects from the database
# 3. Deleting an object from the database
# 4. Add item to the database
# 5. Looping through all scenarios
# Before starting, let us do the necessary imports
import os
import matplotlib.pyplot as plt
import numpy as np
from domain_model import DocumentManagement, StateVariable
# ## 1. Loading and saving the database
#
# A real-life application of the object-oriented framework is the scenario database of [TNO's StreetWise](https://www.tno.nl/en/focus-areas/traffic-transport/expertise-groups/research-on-integrated-vehicle-safety/scenario-based-safety-validation-for-connected-and-automated-driving/). The code contains a class that is used to interface with the database. This class (`DocumentManagement`) can also be used to interact in a similar fashion with a locally stored database. This is practical if the local database is not too large.
#
# In this tutorial, we will use `DocumentManagement` using a locally stored set of scenarios in which the leading vehicle is decelerating. Loading a "database" from a locally stored file is straightforward. When passing a file path when creating an object from the `DocumentManagement` class, the file is loaded.
DM = DocumentManagement(os.path.join("examples", "lvd_scenarios.json"))
# The JSON code of all objects are now contained by `DM`. More specifically, `DM.collections` contains dictionaries of all objects. For example, `DM.collections["scenario"]` contains all scenarios. This dictionary, which consists for (key, value) pairs, uses the scenario IDs as "keys" and the JSON code of the scenario as "value".
#
# Using `DM.collections`, we can see how much data we have:
print("We have:")
print("{:d} scenarios,".format(len(DM.collections["scenario"])))
print("{:d} activity categories,".format(len(DM.collections["activity_category"])))
print("{:d} activities,".format(len(DM.collections["activity"])))
print("{:d} events,".format(len(DM.collections["event"])))
print("{:d} actor categories,".format(len(DM.collections["actor_category"])))
print("{:d} actors,".format(len(DM.collections["actor"])))
print("{:d} physical element categories,".format(len(DM.collections["physical_element_category"])))
print("{:d} physical elements, and".format(len(DM.collections["physical_element"])))
print("{:d} models.".format(len(DM.collections["model"])))
# Saving all objects is also straightforward. Use the `to_json` function to store the "database" into a JSON file.
DM.to_json(os.path.join("examples", "lvd_scenarios2.json"))
# ## 2. Obtaining objects from the database
#
# The `DM` contains all JSON codes of the objects, but what we actually want is to have all objects. To retrieve an actual object, the `get_item` function is used. It takes two arguments:
#
# - The type of object you want (specified using a string, e.g., `"scenario"` or `"actor"`)
# - The key value, i.e., the ID of the object.
#
# For example, let us retrieve the first scenario in the database:
key = list(DM.collections["scenario"].keys())[0]
scenario1 = DM.get_item("scenario", key)
# Now `scenario1` is a `Scenario` object. All of its attributes are automatically instantiated using `get_item`. For example, here are the actors:
for actor in scenario1.actors:
print("{:s}: {}".format(actor.name, actor))
# When using `get_item` to retrieve an object that has already been instantiated, then the object that has been instantiated is returned instead. This is desired, because otherwise two objects of the same, e.g., actor, is returned. To see what we mean, let's look at the following example:
key = list(DM.collections["scenario"].keys())[1]
scenario2 = DM.get_item("scenario", key)
for actor in scenario2.actors:
print("{:s}: {}".format(actor.name, actor))
# This is a different scenario, but one actor is the same as for the first scenario (the ego vehicle). When having a close look at the pointer's address, we can see that the ego vehicle of the second scenario is actually the same object as for the first scenario. The target vehicle, however, is a different object this time.
# ## 3. Deleting an object from the database
#
# Deleting an item from the database can be done using the `delete_item` function. It uses the same arguments as the `get_item` function.
DM.delete_item("scenario", scenario1.uid)
scenario1.uid in DM.collections["scenario"]
# As shown above, the ID of the deleted scenario is not found anymore in the collections. Note, however, that the attribute objects of the scenario are not deleted from the database. For example, the actor is still in the database:
scenario1.actors[0].uid in DM.collections["actor"]
# For demonstrating another functionality, let us also delete the target vehicle of the first scenario:
DM.delete_item("actor", scenario1.actors[1].uid)
# ## 4. Add item to the database
#
# To add an object to the database, use the `add_item` function with as only argument the object that has to be added to the database. For example, we can add the deleted scenario to the database.
DM.add_item(scenario1)
# Note, however, that when trying to retrieve this scenario that we just added, an error will occur:
try:
scenario3 = DM.get_item("scenario", scenario1.uid)
except KeyError as e:
print("Key not found: {}".format(e))
print("ID of actor that we deleted: {:d}".format(scenario1.actors[1].uid))
# What happened here? While we are trying to retrieve the scenario, the `get_item` function automatically tries to retrieve the attribute objects of the scenario. However, it fails to retrieve the target vehicle, because that object has not been added to the database after we deleted it. In other words, by default, only the object that is passed to `add_item` is stored in the database, while its attribute objects are not stored.
#
# In order to also store the attributes, add `include_atributes=True` to the function call:
DM.add_item(scenario1, include_attributes=True)
scenario3 = DM.get_item("scenario", scenario1.uid)
# Now there is no error when retrieving the scenario from the database.
# ## 5. Looping through all scenarios
#
# In the following example, we demonstrate how to loop through all scenarios. In principle, the same can be done with the other objects.
# +
for key in DM.collections["scenario"]:
scenario = DM.get_item("scenario", key)
# To get the speed, we need:
# 1. The right actor, which is always the second (index=1) actor of the scenario.
# 2. Select the right state variable (StateVariable.LON_TARGET).
# 3. Select the right time window.
# Note: StateVariable.LON_TARGET contains two variables: speed and distance (hence the "[:, 0]").
time = np.linspace(scenario.get_tstart(), scenario.get_tend(), 100)
leading_speed = scenario.get_state(scenario.actors[1], StateVariable.LON_TARGET, time)[:, 0]
plt.plot(time-scenario.get_tstart(), leading_speed, color=(.5, .5, .5))
plt.xlabel("Time [s]")
plt.ylabel("Speed [m/s]")
_ = plt.title("Speed of leading vehicle during scenario")
# -
# You reached the end of the third tutorial. In the [next tutorial](./Tutorial%205%20Scenario%20category%20including%20I2V%20communication.ipynb), we will see how we can use the domain model to create a scenario that involves communication.
| Tutorial 4 Scenario database.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2
import numpy as np
from pynput.mouse import Button, Controller
import wx
lower_blue = np.array([110,100,100])
upper_blue = np.array([130,255,255])
lower_red = np.array([170,120,70])
upper_red = np.array([180,255,255])
mouse = Controller()
app=wx.App(False)
(sx,sy)=wx.GetDisplaySize()
(camx,camy)=(320,240)
kernelOpen=np.ones((5,5))
kernelClose=np.ones((20,20))
# +
cap = cv2.VideoCapture(0)
pinchFlag = 0
while True:
_, img=cap.read()
img=cv2.resize(img,(camx,camy))
imgHSV= cv2.cvtColor(img,cv2.COLOR_BGR2HSV)
# mask_red=cv2.inRange(imgHSV,lower_red,upper_red)
mask_blue=cv2.inRange(imgHSV,lower_blue,upper_blue)
# maskOpen_red=cv2.morphologyEx(mask_red,cv2.MORPH_OPEN,kernelOpen)
# maskClose_red=cv2.morphologyEx(maskOpen_red,cv2.MORPH_CLOSE,kernelClose)
maskOpen_blue=cv2.morphologyEx(mask_blue,cv2.MORPH_OPEN,kernelOpen)
maskClose_blue=cv2.morphologyEx(maskOpen_blue,cv2.MORPH_CLOSE,kernelClose)
# conts_red,h=cv2.findContours(maskClose_red.copy(),cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_NONE)
conts_blue,h=cv2.findContours(maskClose_blue.copy(),cv2.RETR_EXTERNAL,cv2.CHAIN_APPROX_NONE)
if(len(conts_blue)==2):
if(pinchFlag==1):
pinchFlag=0
mouse.release(Button.left)
hull1 = cv2.convexHull(conts_blue[0],False)
img = cv2.drawContours(img,[hull1],-1,(0,0,255),1)
M1 = cv2.moments(hull1)
c1X = int(M1["m10"] / M1["m00"])
c1Y = int(M1["m01"] / M1["m00"])
hull2 = cv2.convexHull(conts_blue[1],False)
img = cv2.drawContours(img,[hull2],-1,(0,0,255),1)
M2 = cv2.moments(hull2)
c2X = int(M2["m10"] / M2["m00"])
c2Y = int(M2["m01"] / M2["m00"])
cX = (c1X +c2X)/2
cY = (c1Y +c2Y)/2
mouseLoc=(sx-(cX*sx/camx), cY*sy/camy)
mouse.position=mouseLoc
while mouse.position!=mouseLoc:
pass
elif(len(conts_blue)==1):
if(pinchFlag==0):
pinchFlag=1
mouse.press(Button.left)
hull = cv2.convexHull(conts_blue[0],False)
img = cv2.drawContours(img,[hull],-1,(0,0,255),1)
M = cv2.moments(hull)
cX = int(M["m10"] / M["m00"])
cY = int(M["m01"] / M["m00"])
mouseLoc=(sx-(cX*sx/camx), cY*sy/camy)
mouse.position=mouseLoc
while mouse.position!=mouseLoc:
pass
cv2.imshow("cam",img)
key = cv2.waitKey(1)
if key & 0xFF == ord('q'):
break
cap.release()
cv2.destroyAllWindows()
# -
cap.release()
| trials/mouse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from my_packages.Operations import Math
Math.get_average([1,3,4,5,6,7,88,9,4,5,66,3,44,46,7,9])
from my_packages.Transformers import StringTransformer
StringTransformer.reverse_string('em si siht olleH')
from my_packages.Transformers import StringTransformer
StringTransformer.capitalizestring('text')
| Notebook for my_packages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ECY85RClqCAm" colab={"base_uri": "https://localhost:8080/"} outputId="73e7f08b-d83b-4986-ab7b-9bbaf30b9654"
import nltk
from nltk.stem import *
nltk.download('punkt')
# + id="cB1q1C5h-h0Z"
text = """For the second time this year, the coronavirus has found its way to the very top of British politics and forced Prime Minister <NAME> into self-quarantine. On Sunday night, Johnson tweeted that he must "self-isolate for two weeks, after being in contact with someone with Covid-19." "It doesn't matter that I've had the disease and I'm bursting with antibodies," he said in a Monday video message, adding that he "felt great" and would keep leading the UK virus response, as well as his government's plans to "#BuildBackBetter." Yet the optimism in that message, including the hashtag, masks the reality of exactly how enormous a week this is for the Johnson premiership, and how much of a blow it is for the PM to be trapped in solitude. Downing Street had spent the weekend dealing with the fallout from three straight days of chaos, in which two of his most senior advisers dramatically resigned following allegations that they had been briefing viciously against both Johnson himself and his fiancée, <NAME>. The advisers in question, <NAME> and <NAME>, were among the most controversial and disliked members of Johnson's inner circle and have been accused by numerous people in government of being power hungry and self-interested. Before Johnson's self-quarantining, the turmoil in Downing Street had dominated five days of coverage in the UK, overshadowing what is arguably an even bigger headache for the PM than the coronavirus. Brexit really is now on the home stretch. The current transition period -- which was designed to prevent a sudden halt of the flow of goods, among other things, between the UK and the European Union -- ends on December 31. If the two sides are unable to strike a free trade agreement before that date, then the chaotic no-deal cliff edge -- which many fear would lead to shortages in things like food, toilet paper and medicine -- would be the new reality. Thursday's video conference of the EU27 is the penultimate time that the heads of government from the bloc's member states are scheduled to meet before the end of the year. The final meeting of 2020, on December 10, is considered too late in the day. As has been the case for months, a deal is in sight and the areas of agreement vastly outweigh the areas of disagreement. However, the key stumbling blocks that have prevented a deal remain. The first and most important is Brussels' insistence on a level playing field in exchange for access to the EU's single market. This, for some time, has been a red line for the UK, which objects to being bound by EU competition rules prohibiting how the government could use state aid to help the growth of British enterprise. The two other key sticking points -- fishing rights and the involvement of EU law in the arbitration of any deal -- are also difficult, though it's easier to see a path to agreement in both. It has for some time been assumed that when the talks reached the stages that they are at now (with legal texts on the table and a deal within grasp), the negotiators, who are civil servants acting on the mandate of their political leadership, would make way for political leaders to bridge the final gaps."""
# + id="f8nJ_ejm-sH7"
from nltk.tokenize import RegexpTokenizer
tokenizer = RegexpTokenizer(r'\w+')
words = tokenizer.tokenize(text)
# + id="qbxA8rZ4-5-z" colab={"base_uri": "https://localhost:8080/"} outputId="4a3d32b8-7a4e-4e9d-aa56-46bcbf93faa4"
nltk.sent_tokenize(text)
# + id="Inu0BjwS-9iF" colab={"base_uri": "https://localhost:8080/"} outputId="13ef97f6-de9d-4aaa-ee63-1054b9e2f73a"
from nltk.probability import FreqDist
freq = FreqDist(words)
freq.most_common(10)
# + id="0ZBxbaDK_PFB" colab={"base_uri": "https://localhost:8080/", "height": 330} outputId="41dc82b6-27bb-4713-8049-d00d5d39b756"
import matplotlib.pyplot as plt
freq.plot(100,cumulative=False)
plt.show()
# + [markdown] id="BxoNoGm0_2jd"
# Check for "Stop Words"
# + id="66Max7Y1_V49" colab={"base_uri": "https://localhost:8080/"} outputId="85781920-852c-405a-cc79-0ee5559eecde"
import nltk
nltk.download('stopwords')
from nltk.corpus import stopwords
stopwords.words('english')
print(stopwords.words() [620:680])
# + id="15ttRyqa__Wd" colab={"base_uri": "https://localhost:8080/"} outputId="c67b9b65-de2f-458f-8a80-ebde1a4a2822"
print(stopwords.fileids())
# + id="E-iY1mztAapy"
en_stops = set(stopwords.words('english'))
filterd = []
for x in words:
if x not in en_stops:
filterd.append(x)
# + id="D1TLubVHBA-b" colab={"base_uri": "https://localhost:8080/"} outputId="88865cf3-7f7f-4ab0-ec92-5d81c61660d9"
filterd_dist = FreqDist(filterd)
filterd_dist.most_common(10)
# + [markdown] id="uuBj_pg3SXMb"
# Checking for Stemmerize words
# + id="9YeZQL1jBf03"
from nltk.stem import PorterStemmer
ps = PorterStemmer()
stemmered = []
for x in filterd:
stemmered.append(ps.stem(x))
# + id="zzIRl7ZHHcee" colab={"base_uri": "https://localhost:8080/"} outputId="16bc6dd1-7d86-43ea-a53e-51036d2aff79"
stemmered_dist = FreqDist(stemmered)
stemmered_dist.most_common(10)
# + [markdown] id="GhwTwGDkShPI"
# Checking for Lemmatizeried words
# + id="mlzbmhjbXAUS" colab={"base_uri": "https://localhost:8080/"} outputId="7e05356e-34c5-4497-8ab8-a405e6fc49ed"
# %pip install -U textblob
# + id="yj-vZ9WqHvmU" colab={"base_uri": "https://localhost:8080/", "height": 205} outputId="f7b57535-b114-40ab-c10c-1bc0126c6c80"
import nltk
nltk.download('wordnet')
nltk.download('averaged_perceptron_tagger')
from nltk.stem import WordNetLemmatizer
from textblob import TextBlob, Word
def lemmatize_with_postag(sentence):
sent = TextBlob(sentence)
tag_dict = {"J": 'a',
"N": 'n',
"V": 'v',
"R": 'r'}
words_and_tags = [(w, tag_dict.get(pos[0], 'n')) for w, pos in sent.tags]
lemmatized_list = [wd.lemmatize(tag) for wd, tag in words_and_tags]
return " ".join(lemmatized_list)
# Lemmatize
sentence = text
lemmatize_with_postag(sentence)
# + id="Uhm5BmH3Z8I9"
new = lemmatize_with_postag(sentence)
new_words = tokenizer.tokenize(new)
# + id="aqjYxOToVAlf" colab={"base_uri": "https://localhost:8080/"} outputId="600ed462-e022-40a8-98ed-d0163009ee60"
new_list = []
for x in new_words:
if x not in en_stops:
new_list.append(x)
freq_new = FreqDist(new_list)
freq_new.most_common(10)
# + [markdown] id="fLOtq4UJZXEo"
# Sentiment Analysis
# + id="0KSeaHC6ZXoW"
from nltk import sentiment
from nltk.sentiment.vader import SentimentIntensityAnalyzer
def sentiment_scores(sentence):
sid_obj = SentimentIntensityAnalyzer()
sentiment_dict = sid_obj.polarity_scores(sentence)
print("Overall sentiment dictionary is : ", sentiment_dict)
print("sentence was rated as ", sentiment_dict['neg']*100, "% Negative")
print("sentence was rated as ", sentiment_dict['neu']*100, "% Neutral")
print("sentence was rated as ", sentiment_dict['pos']*100, "% Positive")
print("Sentence Overall Rated As", end = " ")
if sentiment_dict['compound'] >= 0.05 :
print("Positive")
elif sentiment_dict['compound'] <= - 0.05 :
print("Negative")
else :
print("Neutral")
# + id="QNIEm2v9Z9mk" colab={"base_uri": "https://localhost:8080/"} outputId="59299498-e0f0-4ae0-b959-9e552749c957"
import nltk
nltk.download('vader_lexicon')
if __name__ == "__main__" :
print("\n1st statement :")
sentence = new
sentiment_scores(sentence)
print("\n2nd Statement :")
sentence = text
sentiment_scores(sentence)
print("\n3rd Statement :")
sentence = text2
sentiment_scores(sentence)
# + [markdown] id="xbtvZgaWbnX1"
# Bag of Words
# + id="f3S6AaZNbGQ5"
text2 = """President <NAME> is facing a barrage of calls to permit potentially life-saving transition talks between his health officials and incoming President-elect <NAME>'s aides on a fast-worsening pandemic he is continuing to ignore in his obsessive effort to discredit an election that he clearly lost. The increasingly urgent pleas are coming from inside his administration, the President-elect's team and independent public health experts as Covid-19 cases rage out of control countrywide, claiming more than 1,000 US lives a day. More than 246,000 Americans have now died from the disease, and a bitter winter lies ahead even amid encouraging news such as Monday's announcement that a vaccine developed by Moderna is demonstrating a high success rate in early clinical trials, the second such positive vaccine news in about a week. But instead of listening or mobilizing to tackle what some medical experts warn is becoming a "humanitarian" crisis, Trump spent the weekend during which the US passed 11 million infections amplifying lies and misinformation about his election loss. At one point, he appeared to acknowledge Sunday in a tweet that Biden won, before backtracking with a stream of defiance on Twitter. This came as the nation's top infectious disease expert, Dr. <NAME>, said on CNN's "State of the Union" Sunday that "of course it would be better if we could start working with" the Biden team that will take office on January 20. Biden's incoming White House chief of staff <NAME> said Sunday that the President-elect's team had been unable to talk to current top health officials like Fauci about the pandemic owing to Trump's refusal to trigger ascertainment — the formal process of opening a transition to a new administration. "<NAME>'s going to become president of the United States in the midst of an ongoing crisis. That has to be a seamless transition," Klain said on NBC's "Meet the Press," adding that while the new administration planned to contact top pharmaceutical firms making the vaccine like Pfizer, it was particularly key to get in touch with Department of Health and Human Services officials responsible for rolling it out in the coming months. But the official who is currently most influential with the President, Dr. <NAME>, who critics say favors a herd immunity approach that could lead to thousands of deaths, wrote an inflammatory tweet on Sunday that exemplified the White House's contempt for unifying leadership during the pandemic. Atlas called on the people of Michigan to "rise up" against new Covid-19 restrictions introduced in schools, theaters and restaurants by Democratic Gov. <NAME> -- who was recently the target of an alleged domestic terrorism kidnapping plot."""
# + id="0Wiin2GebvNz"
words2 = tokenizer.tokenize(text2)
filtered2 = [w for w in words2 if w not in en_stops]
stemmered2 = [ps.stem(w) for w in filtered2]
freq_words2 = FreqDist(stemmered2)
# + id="NZvUtNqZb-3d" colab={"base_uri": "https://localhost:8080/", "height": 160} outputId="946dc5ce-5e3d-4541-f6df-6c6e8856284d"
import pandas as pd
df = pd.DataFrame([dict(stemmered_dist),dict(freq_words2)])
df.fillna(0,inplace=True)
df.head()
# + id="_AYlncYXceCt" colab={"base_uri": "https://localhost:8080/"} outputId="11bad6ab-014a-43e6-a6cb-481175b09e60"
from sklearn.metrics.pairwise import cosine_similarity
cosine_similarity(df)
# + id="-VsCwmM8ZOja"
| assets/EMSE6574/Week10_Assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv-datascience
# language: python
# name: venv-datascience
# ---
# # K Nearest Neighbors
#
# You've been given a classified data set from a company! They've hidden the feature column names but have given you the data and the target classes.
#
# We'll try to use KNN to create a model that directly predicts a class for a new data point based off of the features.
# # Import Libraries
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# # Data
df = pd.read_csv('Classified Data', index_col=0)
df.head()
df.info()
df.describe().T
# # Standardize the Variables
#
# **IMPORTANT:**
#
# Because the KNN classifier predicts the class of a given test observation by identifying the observations that are nearest to it, the scale of the variables matters. Any variables that are on a large scale will have a much larger effect on the distance between the observations, and hence on the KNN classifier, than variables that are on a small scale.
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaled_features = scaler.fit_transform(df.drop('TARGET CLASS', axis=1))
scaled_features[:5]
type(scaled_features)
# re-create features dataframe, column name is without TARGET CLASS
X = pd.DataFrame(scaled_features, columns=df.columns[:-1])
X.head()
# # Train Test Split
from sklearn.model_selection import train_test_split
y = df['TARGET CLASS']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3, random_state=101)
X_train.shape, X_test.shape, y_train.shape, y_test.shape
# # Using KNN
#
# Remember that we are trying to come up with a model to predict whether someone will TARGET CLASS or not. We'll start with k=1.
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train, y_train)
# # Predictions and Evaluations
predictions = knn.predict(X_test)
predictions[:5]
from sklearn.metrics import classification_report, plot_confusion_matrix
print(classification_report(y_test, predictions))
plot_confusion_matrix(knn, X_test, y_test);
# # Choosing a K Value (uisng Elbow method)
#
# Let's go ahead and use the elbow method to pick a good K Value:
# +
error_rate = []
for i in range(1, 40):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train, y_train)
i_pred = knn.predict(X_test)
error_rate.append(np.mean(i_pred != y_test))
# -
plt.figure(figsize=(10,7))
plt.plot(range(1, 40), error_rate, color='blue', linestyle='--', marker='o', markerfacecolor='red', markersize=10)
plt.title('Error Rate vs K Value')
plt.xlabel('K')
plt.ylabel('Error Rate');
# Here we can see that that after arouns K>23 the error rate just tends to hover around 0.06-0.05 Let's retrain the model with that and check the classification report!
# # Retrain the model with chosen model
knn = KNeighborsClassifier(n_neighbors=17)
knn.fit(X_train, y_train)
predictions = knn.predict(X_test)
print(classification_report(y_test, predictions))
plot_confusion_matrix(knn, X_test, y_test);
# # Using RandomizedSearchCV
from sklearn.model_selection import RandomizedSearchCV
knn = KNeighborsClassifier()
parameters = {
'n_neighbors': range(1, 100)
}
cv_model = RandomizedSearchCV(knn, parameters, cv=5, random_state=101, refit=True)
cv_model.fit(X_train, y_train)
cv_model.best_params_
cv_predictions = cv_model.predict(X_test)
print(classification_report(y_test, cv_predictions))
plot_confusion_matrix(cv_model, X_test, y_test);
| Data Science and Machine Learning Bootcamp - JP/14-K-Nearest-Neighbors/01-K Nearest Neighbors with Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This tutorial will demonstrate how to make a diagnostic plot for a planet candidate in TESS data.
import matplotlib.pyplot as plt
import sys
sys.path.append('../code')
# Provide some basic information on the star and its planet. In this example, we'll look at TIC-25375553, the host star of confirmed planet TOI-143.01 (WASP-111b).
# +
star = {}
star['id'] = 25375553
planet = {}
planet['period'] = 2.31097
planet['epoch'] = 1325.58249
planet['durationHours'] = 3.129
planet['radius'] = 17.54
# -
# We need the lightcurve (time, raw flux, detrended flux) of TIC-25375553 to plot it out. You can get this from your favourite source. In this example let's get the SPOC 2 minute cadence data from Sectors 1 and 28 for this target via lightkurve.
# +
import lightkurve as lk
import numpy as np
lcfs = lk.search_lightcurvefile('TIC %i' % star['id'], mission='TESS').download_all()
lc_raw = lcfs.PDCSAP_FLUX.stitch()
lc_clean = lc_raw.remove_outliers(sigma=20, sigma_upper=4)
# Mask out transit from detrending (optional)
temp_fold = lc_clean.fold(planet['period'], t0=planet['epoch'])
fractional_duration = (planet['durationHours'] / 24.0) / planet['period']
phase_mask = np.abs(temp_fold.phase) < (fractional_duration * 1.5)
transit_mask = np.in1d(lc_clean.time, temp_fold.time_original[phase_mask])
lc_flat = lc_clean.flatten(mask=transit_mask)
star['time'] = lc_clean.time
star['raw'] = lc_clean.flux
star['flux'] = lc_flat.flux
# -
# We also need the difference image data for signal. We already have that from running the first tutorial (example_diffimages.ipynb), so let's just load that back in:
# +
import pickle
fname = 'tic25375553/imageData_planet0_sector28_camera1.pickle'
with open(fname, 'rb') as f:
imageData = pickle.load(f)
# -
# Now we have all we need to make a basic report for the planet signal.
# +
from plots import plot_report
plot_report(star, planet, imageData)
# -
# Description of plots:
#
# First column, first row: raw lightcurve, with odd and even transits marked with blue and orange ticks, respectively. Large gaps in the data are skipped over for plotting to ease visibility of data.
#
# First column, second row: detrended lightcurve
#
# First column, third row: phase diagram showing all data
#
# First column, fourth row: close-up of phase diagram centred on transit
#
# Second column, second row: close-up of phase diagram centred on only odd and even transits for comparison
#
# Second column, third row: close-up of phase diagram centred on most significant secondary in lightcurve (default at phase = 0.5; set `planet['phs_sec'] = X` to centre the plot elsewhere)
#
# Second column, fourth row: phase diagram centred on phase = 0.5
#
# Third column, first row: full difference image SNR
#
# Third column, second row: full direct image
#
# Third column, third row: close-up of difference image SNR
#
# Third column, fourth row: close-up of direct image
#
# Fourth column: summary of information about planet and star. Stellar properties are queryied from the TIC catalogue automatically.
| sardis/.ipynb_checkpoints/example_diagnosticplot-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nuclio - prediction function
# ## Setup the environment
# nuclio: ignore
import nuclio
# ### Set environment variables
import os
# nuclio: ignore
os.environ['CUSTOMERS_TABLE'] = os.path.join(os.getenv('V3IO_USERNAME', 'iguazio'), 'customers')
os.environ['PREDICTIONS_TABLE'] = os.path.join(os.getenv('V3IO_USERNAME', 'iguazio'), 'predictions')
# +
# Iguazio access
# %nuclio env FRAMESD=${V3IO_FRAMESD}
# %nuclio env V3IO_USERNAME=${V3IO_USERNAME}
# %nuclio env V3IO_ACCESS_KEY=${V3IO_ACCESS_KEY}
# Model handling
# %nuclio env MODEL_FILE=lgb.model
# %nuclio env -c MODEL_FILEPATH=/tmp/mlmodel/${MODEL_FILE}
# %nuclio env -l MODEL_FILEPATH=models/trained/${MODEL_FILE}
# Function variables
# %nuclio env NUM_OF_PRODUCTS_TO_RETURN=4
# %nuclio env CUSTOMERS_TABLE= ${CUSTOMERS_TABLE}
# %nuclio env PREDICTIONS_TABLE= ${PREDICTIONS_TABLE}
# -
# ### Base image
# %nuclio config spec.build.baseImage = "python:3.6-jessie"
# ### Installations
# When installing packages while working, Please reset the kernel to allow Jupyter to load the new packages.
# %%nuclio cmd -c
pip install requests
pip install pandas
pip install lightgbm
pip install v3io_frames
# ### Get the model
# nuclio: ignore
# Verify the model is in the shared data directory
os.environ['MODEL_SHARED_FILEPATH'] = f'/users/{os.environ["V3IO_USERNAME"]}/recommendation_demo/models/'
os.environ['FULL_LOCAL_MODEL_PATH'] = f'{os.path.join(os.getcwd(), os.environ["MODEL_FILEPATH"])}'
# !mkdir -p /v3io${MODEL_SHARED_FILEPATH}
# !cp ${FULL_LOCAL_MODEL_PATH} /v3io${MODEL_SHARED_FILEPATH}
# %nuclio env MODEL_SHARED_FILEPATH = ${MODEL_SHARED_FILEPATH}
# %%nuclio cmd -c
apt-get update && apt-get install -y wget
# mkdir -p /tmp/mlmodel
wget -O /tmp/mlmodel/${MODEL_FILE} --header "x-v3io-session-key: ${V3IO_ACCESS_KEY}" http://${V3IO_WEBAPI_SERVICE_HOST}:8081${MODEL_SHARED_FILEPATH}${MODEL_FILE}
# ### Imports
# +
# Util
import json
import requests
import datetime
# Function
import pandas as pd
import lightgbm as lgb
# DB
import v3io_frames as v3f
# -
# ## Function code
# ### Init context
def init_context(context):
# Define DB
client = v3f.Client('framesd:8081', container='users')
setattr(context, 'client', client)
setattr(context, 'customers_table', os.environ['CUSTOMERS_TABLE'])
setattr(context, 'predictions', os.environ['PREDICTIONS_TABLE'])
try:
context.client.delete('tsdb', context.predictions, if_missing=1)
except:
context.logger.debug(f'couldnt delete {context.predictions}')
try:
context.client.create('tsdb', context.predictions, attrs={'rate': '1/s'})
except:
context.logger.debug(f'couldnt create {context.predictions}')
# define Model
model = lgb.Booster(model_file=os.environ['MODEL_FILEPATH'])
setattr(context, 'model', model)
# vars
setattr(context, 'k', int(os.environ['NUM_OF_PRODUCTS_TO_RETURN']))
# ### Format dataframe for prediction
def prepare_df(df):
# Extract features col
keep_cols = ['products']
df = df.loc[:, keep_cols]
# Create features df
df = pd.read_json(df.values[0][0])
return df
def handler(context, event):
# Get user
customer_id = event.body['id']
store = event.body['store']
context.logger.debug(f'Predicting for: {customer_id} in {store}')
# Get user parameters
df = context.client.read('kv', context.customers_table, filter=f'id=={customer_id}').reset_index()
# Do we have features for the user?
if not df.empty:
# Create features df for the user
df = prepare_df(df)
# Predict
df["prediction"] = context.model.predict(df)
# Eliminate low confidence results
df = df.loc[df.prediction > 0.01, ['prediction', 'product_id']]
# Keep top products
df.sort_values(by='prediction', ascending=False, inplace=True)
best_products = df.iloc[:context.k, :]
best_products = best_products.reset_index()
best_products = best_products.reset_index()
best_products = best_products.rename(columns={'level_0': 'prediction_num'})
best_products['customer_id'] = int(customer_id)
best_products['store'] = store
best_products['time'] = datetime.datetime.now()
best_products = best_products.drop('index', axis=1)
best_products = best_products.set_index(['time', 'store', 'prediction_num', 'product_id'])
context.logger.debug(f'Predicted:\n{best_products}')
# Save results to DB
context.client.write('tsdb', context.predictions, best_products)
# nuclio: ignore
init_context(context)
# nuclio: ignore
event = nuclio.Event(body={'id': '1232', 'store': '03311311313011021022'})
handler(context, event)
# nuclio: ignore
context.client.read('tsdb', context.predictions)
# %nuclio deploy -n prediction_server -p recommendation_engine -c
| demos/location-based-recommendation/03-nuclio-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Report 5: Cellular Automata
# ## <NAME>
# ### Introduction
# A cellular automaton is a self-contained universe where its space is divided into a grid of cells, and has its own "laws of physics". Each cell in the grid has its own state and a set of neighbors. Time in this universe is a discrete set of steps. At each step of time, a transition rule updates the state of every cell based on the current state of the cell and its neighbors.
#
# The concept of cellular automata was discovered in the 1940s by <NAME> and <NAME> (who also discovered the prime spiral). However, it wasn't until the 1970s when the cellular automaton the "Game of Life" was created by <NAME> that interest in cellular automata grew. Today, cellular automata are normally used to model large, complex physical systems in areas such as theoretical biology, microstructure modeling, and computer processors.
# ### Task Statement
# In this report, the goal is to implement the cellular automaton that was defined in class and explore all the possible behaviors that different transition rules can lead to. The cellular automaton has the following characteristics:
# * A one dimensional grid with each cell having just two neighbors, one on each side.
# * There are four possible cell states: 1, 2, 3, 4. Each state is represented by a different color.
# * There are periodic edge conditions; the rightmost and leftmost cells are neighbors.
# * The transition rule is that the state of each cell is updated based on the sum of the current cell state and those of its neighbors.
# ### Methods
# %pylab inline
from numpy.random import randint
def CellAuto():
nGen=25
nCells=25
nrow=3
ncol=3
Colors=array([[1,0,0], [0,1,1], [.5,0,1], [1,1,0]], dtype=float) #red, cyan, purple, yellow
figure(figsize=(15,15))
for j in range(1,nrow*ncol+1):
CellState=empty((nGen, nCells),dtype=int)
CellState[0]=randint(4, size=nCells)
Rule=randint(4,size=10)
for i in range(1,nGen):
SumCells=CellState[i-1]+roll(CellState[i-1],-1)+roll(CellState[i-1],1)
CellState[i]=Rule[SumCells]
CellColor=Colors[CellState]
subplot(nrow,ncol,j)
imshow(CellColor, interpolation='none')
xticks([])
yticks([])
title(j, fontsize='12')
xlabel(Rule, fontsize='12');
# The function **CellAuto()** is used to generate multiple plots of cellular automata all at once. First, some variables are defined:
#
# * **nGen** is the number of generations the function will apply the transition rule to. It is also the number of rows.
# * **nCells** is the number of cells present within this cellular automaton. It is also the number of columns.
# * **nrow** is the number of rows in the subplot.
# * **ncol** is the number of columns in the subplot.
# * **Colors** is an array of shape (4,3) that contains the colors to be used in the images. From left to right, the colors are: red, cyan, purple, and yellow.
# * **figure** is not a variable, but it is used to define the figure size that will be used to hold all the subplotted images.
#
# In this function, **nGen** and **nCells** are both 25. This means that there will be 25 generations, and each generation will have 25 cells. **nrow** and **ncol** are both set to 3, and this means that the subplot will be broken up into a grid of 3 rows and 3 columns, so there will be 9 images total with 3 images per row. As stated, **Colors** is the array that holds the colors to be used. The keyword argument *dtype* is used to set the data type in the array to be floats. Lastly, a figure size of 15 by 15 will be used to subplot the images. (Note: The reason nGen and nCells are 25 is that my computer doesn't really like to run anything larger than this for some reason. When I ran 50 it blue screened. Luckily, it is enough to see the end behaviors.)
#
# Next is a *for loop* that runs from range 1 up to (but not including) **nrow x ncol +1**, or 10. This *for loop* is used to plot different cellular automata in a 3 by 3 grid, thus all code that creates the automata must be put within it.
#
# First, the **CellState** array must be created. This is an initially empty integer array that is the size **nGen** by **nCells**, or 25 by 25. We then set the first row (index 0) of this array as a row of random integers ranging from 0 to 3. By doing this, every time the *for loop* begins again, the initial generation of cell states is randomly set. After that, the transition rule, **Rule**, is generated. The rule is also an array of random integers from 0 to 3, but *size=10* means that it only contains 10 numbers.
#
# Another *for loop* is used to apply the transition rule to the intitial or current cell states, and it runs from 1 to **nGen**. The reason it starts at 1 and not 0 is because we are generating the next, or ith, generation of cell states. We already defined what state 0 is, so we do not want to overwrite it. In this cellular automata, the transition rule is applied to the sum of a cell and its neighbors to the right and left. To obtain the sum of the current cell states (the **SumCells** array), we use the *roll* function to shift the array elements by 1 to the left or right. When using *roll*, we shift the ith-1 generation, since this the generation right before the one we are trying to compute. Also, since the loop range must start at 1, i-1 allows for the initial generation to be used. Then, to get the next generation of cell states, we use array indexing to apply the transition rule to the **SumCells** array. The array indexing does this by replacing values in the **SumCells** array with values in the **Rule** array based on the index in the **Rule** array. For example, let's say one of the values in **SumCells** is 4. This number will be replaced by whatever number is indexed 4 in **Rule** (which would be the 5th number because indexing starts at 0). When the transition rule is done being applied, the new array of number becomes the ith generation in **CellStates**.
#
# The rest of the code is outside the 2nd *for loop*, but still inside the 1st.
#
# Now that we have generated all 25 generations, we must apply a color to each individual cell state. We once again use array indexing to replace the values in **CellStates** with the colors defined in the **Colors** array, and we call this new array **CellColor**. The following values are replaced with these colors:
# * A state of 0 is replaced with the color red.
# * A state of 1 is replaced with the color cyan.
# * A state of 2 is replaced with the color purple.
# * A state of 3 is replaced with the color yellow.
#
# Finally, now that we have an array of only colors, we can plot the image. First, *subplot* is called, and the 3 by 3 grid is created. As the first loop runs, each image will be subplotted in the jth position in the grid. Next, *imshow* is called, and this is what displays the color array **CellColor**. The keyword argument *interpolation='none'* is used to prevent the colors from blurring into one another. The x- and y-axis tick marks are turned off, and each image is given a number, j, to identify it (this is also its position number in the grid). Additionally, the transition rule is put below each image by making it the x-axis label.
# ### Results
CellAuto()
# After running the **CellAuto** function, we obtain the above images, and make the following observations:
#
# 1. Images 2, 6, and 9 all have downward pointing triangles of various sizes. Inspection of all the rules reveals that the 1st postiton might be important since the color of the triangles is the number in index 0. Index 5, and possibly 6, might also be important because they correspond to colors found in the background. (Note: I know the images dont have backgrounds per se, but it is easier for me to describe the most prominent color, if there is one, as the background color.) Additionally, indices 1 through 3 might also have an influence, since all the colors in these positions make up the outline of the triangles.
#
# 2. Images 1 and 4 contain patterns with no discernable order.
#
# 3. Image 8 has an end behavior of all purple except for a cyan stripe. Cyan replaces cell states of 1, and since there is only a single 1 present within the transition rule, the location of the 1 in index 4 must be very important. That is, if the 1 is anywhere else, we would not get this vertical stripe. Importance in position can also be seen in this rule because there are two 3's, but we see that the color yellow is gone by the 2nd generation. Thus, it can be concluded that the 3's are not in an important position.
#
# 4. Image 7 has a ladder-like pattern of purple and yellow horizontal stripes. Even though there are 0's and 1's, red and cyan are gone after several generations. The rule sequence [...2,2,3,2,2,2] is what is most likely causing this pattern, since 2=purple and 3=yellow, and the yellow lines are surrounded by purple.
#
# 5. Image 3 has a main background color (red) with horizontal (cyan) stripes of various lengths. In the rule, half of the numbers in the rule are 0 (0=red), so this could explain the red background. There is only a single 1, and the position of this 1 must be important because this is what would cause the cyan stripes. Even though there are a lot of 3's, they must not be in important positions, because the amount of yellow and purple looks almost the same, and there is only one 2 versus three 3's.
#
# 6. Image 5 has a combination of all the other images. It has yellow downward triangles, ladder-like structures, horizontal stripes, and random patterns.
CellAuto()
# I ran the function a second time and found the following:
# (Note: when I reference the other images and their rules, I will refer to them as number # above)
#
# 1. Images 1 and 5 are like the images in number 2 above; They contain no patterns.
#
# 2. Image 8 is like the image in number 3 above (also image 8). They both have an end behavior of one color, but this image does not have the stripe. Index 4 in this image is 2, which is purple, and this confirms the above observation that this index is important. In this image index 4 is the same color as the end behavior, no stripe appears, while in the other image, index 4 was 1 which was the only source of the cyan stripe.
#
# 3. Image 9 is like the images in number 1 with the triangles. The rule upholds most of the observations made about indicies 0 through 3, but not 5 and 6 since there is not a background color.
#
# 4. I could have put image 7 in with image 8, but I felt that this type of stripe deserved its own category because it depends on multiple indices and not just index 4. In the other 2 images, it seems that the number in indices 2 and 9 do not matter, but in this case it does because one of them must be the source of the purple.
#
# 5. Image 4 has that ladder-like pattern seen in number 4 above, but the rules do not follow the same pattern. In this case, it seems that [...1,2,2,2,1,...] located in the middle of the array might be responsible. In the other image, the sequence was located at the end of the array.
#
# 6. Image 2 is like number 6 above in that is has almost all of the patterns seen so far present.
#
# 7. Image 6 is another one of those with an end behavior of mostly 1 color, but in this case, there are two stripes of alternating colors. These stripes are most likely the result of the [...1,3,1,3] sequence at the end of the rule.
#
# 8. Image 3 has several different patterns going on, but it is not like image 2. It has 2 main colors, cyan and purple, and this could be due to the fact that the rule has mainly 1's and 2's. Yellow is also on a diagonal and is a vertical stripe, and this could be related to the indicies of the 3's (index 0 and 7).
# ### Textile Cone Pattern
# The last part of the report is to try to find a rule that would produce the Textile Cone Shell shown in the picture (if you have time). While I don't have time to actually implement the rule, I did some research and found the following:
#
# The cellular automaton that makes this pattern is called "Rule 30" and it was introduced by <NAME> in 1983. It is called "Rule 30" because 30 is the smallest Wolfram code that can describe the rule (Wolfram code is a code system created by him too). "Rule 30" is also a Class 3 rule, which means that it displays chaotic and aperiodic behavior.
#
# The rule to make the pattern depends on the current state of the cell and its neighbors on each side. It is the following:
#
# * Current State Pattern: 111 -> New State: 0
# * Current State Pattern: 110 -> New State: 0
# * Current State Pattern: 101 -> New State: 0
# * Current State Pattern: 100 -> New State: 1
# * Current State Pattern: 011 -> New State: 1
# * Current State Pattern: 010 -> New State: 1
# * Current State Pattern: 001 -> New State: 1
# * Current State Pattern: 000 -> New State: 0
# ### Conclusion
# In this report, a simple cellular automaton was created and explored. In the automata, both the transition rule and initial state were both randomly created using the *randint* function in NumPy. Then, the code was run twice to create 18 images with different transition rules. The images were then grouped together by the patterns they contained, and the rules were analyzed in order to see if there were any features that led to that specific pattern. Lastly, how to create the Textile Cone shell pattern was discussed.
# ### References
# 1. https://en.wikipedia.org/wiki/Cellular_automaton
# 2. http://mathworld.wolfram.com/Rule30.html
# 3. https://en.wikipedia.org/wiki/Rule_30
| report_05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import random
from pyBedGraph import BedGraph
from pybedtools import BedTool
import scipy.stats
def read_cf(directory, file_name):
with open(directory + file_name) as f:
chrom = {}
for line in f:
tmp = line.strip().split("\t")
if tmp[0] != 'chrM':
chrom[tmp[0]] = int(tmp[1])
return chrom
def read_peak(directory, file_name):
with open(directory + file_name) as f:
seen = {}
specific = []
common = []
for line in f:
tmp = line.strip().split("\t")
region = ','.join([tmp[0], tmp[1], tmp[2]])
if region not in seen.keys():
if tmp[3] != '.':
common.append([tmp[0], int(tmp[1]), int(tmp[2])])
else:
specific.append([tmp[0], int(tmp[1]), int(tmp[2])])
seen[region] = 1 # add region to seen list
return specific, common
def plot_boxplot(dataset, dlabel, clr, tit, ylab, fig_name):
fig = plt.figure(figsize = (8,6))
medianprops = dict(linewidth = 3, color=clr)
i=0
boxprops = dict(linewidth = 1.5)
toplot = [np.asarray([]) for i in range(len(dataset))]
for d in dataset:
#medianprops = dict(linewidth = 3, color=colcode[i])
datax = toplot
datax[i] = np.asarray(dataset[i])
plt.boxplot(datax, widths = 0.6, medianprops = medianprops, boxprops = boxprops)
i +=1
plt.xticks([i for i in range(1, len(dataset)+1)], dlabel, fontsize = 18)
plt.yticks(fontsize = 18)
plt.ylabel(ylab, fontsize = 18)
#plt.ylim(bottom=2.5)
plt.title(tit, fontsize = 18)
plt.savefig(fig_name+'.pdf', dpi=150, bbox_inches="tight")
plt.show()
plt.close()
def get_cov(interval_list, bgobj):
tmpvals = []
for x in interval_list:
value = list(bgobj.stats(stat = 'max', intervals = [x]))
tmpvals.append(value[0])
return tmpvals
def write_result(directory, out_list, out_name):
with open(directory+out_name, 'a') as file1:
for i in range(len(out_list)):
file1.write('\t'.join(map(str, out_list[i])) + '\n')
file1.close()
directory='/Users/kimm/Desktop/GM12878_files/'
#ctcf_cov='CDH0002NR_hg38_CTCF_FDR_0.1_pseudoGEM_5000_enrichTest_master_PASS.bedgraph'
ctcf_cov='GM12878-CTCF-pooled_comp_sing_FDR_0.2_PASS.bedgraph'
ctcf_peak='GM12878-CTCF-pooled_comp_sing_FDR_0.2_PASS_thresh70_merge3kb_peaks_GM12878-cohesin-pooled_comp_sing_FDR_0.2_PASS_thresh400_merge3kb_peaks_intersect_wao.bed'
#cohesin_cov='SHG0180-181-182NR_hg38_cohesin_FDR_0.1_pseudoGEM_5000_enrichTest_master_PASS.bedgraph'
cohesin_cov='GM12878-cohesin-pooled_comp_sing_FDR_0.2_PASS.bedgraph'
cohesin_peak='GM12878-cohesin-pooled_comp_sing_FDR_0.2_PASS_thresh400_merge3kb_peaks_GM12878-CTCF-pooled_comp_sing_FDR_0.2_PASS_thresh70_merge3kb_peaks_intersect_wao.bed'
chromfile = read_cf(directory, 'hg38.chrom.sizes')
#repet = BedTool(directory+'hg38PAM.sameChr.tx.sorted.legal.6To12Copies.within5kb_col1-4.bed')
bgctcf = BedGraph(directory+'hg38.chrom.sizes', directory+ctcf_cov)
bgcohesin = BedGraph(directory+'hg38.chrom.sizes', directory+cohesin_cov)
for key,val in chromfile.items():
bgctcf.load_chrom_data(key)
bgcohesin.load_chrom_data(key)
cohesin_spec, cohesin_comm = read_peak(directory, cohesin_peak)
ctcf_spec, ctcf_comm = read_peak(directory, ctcf_peak)
len(cohesin_comm)
len(cohesin_spec)
len(ctcf_comm)
len(ctcf_spec)
cohesin_comm_vals = list(bgcohesin.stats(stat = 'max', intervals = cohesin_comm))
cohesin_spec_vals = list(bgcohesin.stats(stat = 'max', intervals = cohesin_spec))
ctcf_comm_vals = list(bgctcf.stats(stat = 'max', intervals = ctcf_comm))
ctcf_spec_vals = list(bgctcf.stats(stat = 'max', intervals = ctcf_spec))
cohesin_comm_vals = get_cov(cohesin_comm, bgcohesin)
cohesin_spec_vals = get_cov(cohesin_spec, bgcohesin)
ctcf_comm_vals = get_cov(ctcf_comm, bgctcf)
ctcf_spec_vals = get_cov(ctcf_spec, bgctcf)
cohesin_stat = scipy.stats.mannwhitneyu([np.log10(x) for x in cohesin_comm_vals], [np.log10(x) for x in cohesin_spec_vals])
cohesin_title = "Cohesin \n Common: median = " + str(int(np.median(cohesin_comm_vals))) + "; n = " + str(len(cohesin_comm_vals)) + "\n" + "Specific: median = " + str(int(np.median(cohesin_spec_vals))) + "; n = " + str(len(cohesin_spec_vals)) + "\n" + "Mann-Whitney stat = " + str(cohesin_stat[0]) + "; pval = " + str(cohesin_stat[1])
plot_boxplot([[np.log10(x) for x in cohesin_comm_vals], [np.log10(x) for x in cohesin_spec_vals]], ['Cohesin common', 'Cohesin specific'], '#006600', cohesin_title, 'log10(Binding intensity)', 'cohesin_common_vs_spec_boxplot')
np.log10(400)
ctcf_stat = scipy.stats.mannwhitneyu([np.log10(x) for x in ctcf_spec_vals], [np.log10(x) for x in ctcf_comm_vals])
ctcf_title = "CTCF \n Common: median = " + str(int(np.median(ctcf_comm_vals))) + "; n = " + str(len(ctcf_comm_vals)) + "\n" + "Specific: median = " + str(int(np.median(ctcf_spec_vals))) + "; n = " + str(len(ctcf_spec_vals)) + "\n" + "Mann-Whitney stat = " + str(ctcf_stat[0]) + "; pval = " + str(ctcf_stat[1])
plot_boxplot([[np.log10(x) for x in ctcf_spec_vals], [np.log10(x) for x in ctcf_comm_vals]], ['CTCF specific', 'CTCF common'], '#0000B2', ctcf_title, 'log10(Binding intensity)', 'ctcf_common_vs_spec_boxplot')
| ipynb/GM12878_CTCF_vs_cohesin_binding_boxplot_20200607.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_pytorch_p36)
# language: python
# name: conda_pytorch_p36
# ---
# +
# %reload_ext autoreload
# %autoreload 2
# %cd /home/ubuntu/dione-sr/
# -
import os
import imageio
import numpy as np
import pandas as pd
from fs_s3fs import S3FS
from matplotlib import pyplot as plt
from tqdm.auto import tqdm
import cv2 as cv
from hrnet.src.train import resize_batch_images
from sr.data_loader import ImagesetDataset
from sr.metrics import METRICS, minshift_loss
from torch.utils.data import DataLoader
aws_access_key_id = ''
aws_secret_access_key = ''
filesystem = S3FS(bucket_name='',
aws_access_key_id=aws_access_key_id,
aws_secret_access_key=aws_secret_access_key)
# +
norm_deimos = {k: v for k, v in np.load(filesystem.openbin('metadata/deimos_min_max_norm.npz')).items()}
norm_s2 = {k: v for k, v in np.load(filesystem.openbin('metadata/s2_min_max_norm.npz')).items()}
data_df = pd.read_parquet(filesystem.openbin('metadata/npz_info_small.pq'))
country_norm_df = pd.read_parquet(filesystem.openbin('metadata/s2_norm_per_country.pq'))
# -
NPZ_FOLDER = ''
data_df.head()
# +
dataset = ImagesetDataset(imset_dir=NPZ_FOLDER,
imset_npz_files=data_df.singleton_npz_filename.values,
filesystem=filesystem,
country_norm_df=country_norm_df,
normalize=True,
norm_deimos_npz=norm_deimos,
norm_s2_npz=norm_s2,
time_first=True
)
dataloader = DataLoader(dataset,
batch_size=256,
shuffle=False,
num_workers=16,
pin_memory=True)
# -
SHIFTS = 6
# ### test run on a single batch
batch = next(iter(dataloader))
batch.keys()
lrs = batch['lr']
hrs = batch['hr']
names = batch['name']
alphas = batch['alphas']
interpolated = resize_batch_images(lrs[:, -1, [-1], ...],
fx=3, fy=3, interpolation=cv.INTER_CUBIC)
mse = METRICS['MSE'](hrs[:, [-1], ...], interpolated.float())
mse_shift, mse_ids = minshift_loss(hrs[:, [-1], ...], interpolated.float(),
shifts=SHIFTS, metric='MSE')
mse_shift_c, mse_ids_c = minshift_loss(hrs[:, [-1], ...], interpolated.float(),
metric='MSE', shifts=SHIFTS, apply_correction=True)
fig, ax = plt.subplots(figsize=(10, 10))
ax.scatter(mse_shift.numpy(), mse_shift_c.numpy(), alpha=.3, label='MSE shifted corrected')
ax.scatter(mse_shift.numpy(), mse.numpy(), alpha=.3, label='MSE')
ax.plot([0, 1], [0, 1], 'k')
ax.grid()
ax.legend()
ax.set_xlabel('MSE shifted')
np.where(mse_shift_c.numpy() > .35)
np.where(mse_shift_c.numpy() < .02)
# +
idx = 224
img_de = hrs[idx, [-1], ...].numpy().squeeze()
img_s2 = interpolated[idx].numpy().squeeze()
# +
ids = mse_ids_c[idx, :].numpy().astype(np.uint8)
print(ids)
img_s2 = img_s2[SHIFTS//2:-SHIFTS//2, SHIFTS//2:-SHIFTS//2]
img_de = img_de[ids[0]:ids[1], ids[2]:ids[3]]
img_s2 = 255*(img_s2-img_s2.min())/(img_s2.max()-img_s2.min())
img_de = 255*(img_de-img_de.min())/(img_de.max()-img_de.min())
giffile = f's2-deimos-{names[idx]}.gif'
imageio.mimsave(giffile,
[img_s2.astype(np.uint8), img_de.astype(np.uint8)],
duration=0.5)
# -
# ## Compute scores on entire dataset of patchlets
# +
pq_filename = 'scores-bicubic-32x32.pq'
if not os.path.exists(pq_filename):
scores = []
for sample in tqdm(dataloader):
hrs = sample['hr'][:, [-1], ...]
interpolated = resize_batch_images(sample['lr'][:, -1, [-1], ...],
fx=3, fy=3, interpolation=cv.INTER_CUBIC)
mse_ = METRICS['MSE'](hrs.float(), interpolated.float())
mse_shift, _ = minshift_loss(hrs.float(), interpolated.float(),
metric='MSE', shifts=SHIFTS)
mse_shift_c, _ = minshift_loss(hrs.float(), interpolated.float(),
metric='MSE', shifts=SHIFTS, apply_correction=True)
psnr_shift_c, _ = minshift_loss(hrs.float(), interpolated.float(),
metric='PSNR', shifts=SHIFTS, apply_correction=True)
ssim_shift_c, _ = minshift_loss(hrs.float(), interpolated.float(),
metric='SSIM', shifts=SHIFTS, apply_correction=True)
for name, mse, mse_s, mse_sc, psnr, ssim in zip(sample['name'],
mse_,
mse_shift,
mse_shift_c,
psnr_shift_c,
ssim_shift_c):
scores.append({'name': name,
'MSE': mse.numpy().astype(np.float32),
'MSE_s': mse_s.numpy().astype(np.float32),
'MSE_s_c': mse_sc.numpy().astype(np.float32),
'PSNR_s_c': psnr.numpy().astype(np.float32),
'SSIM_s_c': ssim.numpy().astype(np.float32)})
df = pd.DataFrame(scores)
print(len(df))
df.MSE = df.MSE.astype(np.float32)
df.MSE_s = df.MSE_s.astype(np.float32)
df.MSE_s_c = df.MSE_s_c.astype(np.float32)
df.PSNR_s_c = df.PSNR_s_c.astype(np.float32)
df.SSIM_s_c = df.SSIM_s_c.astype(np.float32)
df.to_parquet(pq_filename)
else:
df = pd.read_parquet(pq_filename)
# -
len(df)
df.head()
fig, ax = plt.subplots(figsize=(15, 10))
df.MSE.hist(ax=ax, alpha=.3, bins=50, range=(0, 1), label='MSE')
df.MSE_s.hist(ax=ax, alpha=.3, bins=50, range=(0, 1), label='MSE_s')
df.MSE_s_c.hist(ax=ax, alpha=.3, bins=50, range=(0, 1), label='MSE_s_c')
ax.legend()
fig, ax = plt.subplots(figsize=(15, 10))
ax.scatter(df.MSE_s_c, df.SSIM_s_c, alpha=.1)
fig, ax = plt.subplots(figsize=(15, 10))
ax.scatter(df.PSNR_s_c, df.SSIM_s_c, alpha=.1)
data_df.rename(columns={'singleton_npz_filename': 'name'}, inplace=True)
scores_df = pd.merge(df, data_df, on='name')
scores_df.head()
| notebooks/05d-calculate-scores.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
r3=r2.drop_duplicates(subset='hash', keep='last', inplace=False)
# +
r = pd.DataFrame(pd.read_csv("test_with_new_features.csv"))
r1= pd.DataFrame(pd.read_csv("data_test.csv"))
target
# -
r['hash']=r1['hash']
r['target']=r['center']
r=r.drop_duplicates(subset='hash', keep='last', inplace=False)
r=r.drop(['time_entry','time_exit','x_entry','y_entry','x_exit','y_exit','time_interval','distance','center'], axis=1)
r=r.drop(['id'], axis=1)
r = r.rename(columns={'trajectory_id': 'id'})
r.head(10)
r.to_csv("submission_0905.csv",index=False,sep=',')
| tocsv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.model_selection import train_test_split
import numpy as np
# X = m x n
# theta = n x k
# score = softmax_function = m x k
# y = m x 1
class SoftmaxReg():
"""
eta = learning rate.
num_iters =
early_stopping =
"""
def __init__(self, eta=0.1, num_iters=1000, early_stopping=False, penalty=None, alpha=0.1):
self.eta = eta
self.num_iters = num_iters
self.early_stopping = early_stopping
self.penalty = penalty
self.alpha = alpha
self.best_theta = None
def score(self, X, theta):
return X @ theta
def softmax_function(self, X, theta):
results = np.exp(self.score(X, theta)) / np.sum(np.exp(self.score(X, theta)), axis=1, keepdims=True)
return results
def cross_entropy(self, X, y, theta):
m = len(y)
J = - (1 / m) * np.sum(np.log(self.softmax_function(X, theta)).T @ y)
if self.penalty == "l2":
J += (self.alpha / 2) * np.sum(theta[1:] ** 2)
return J
def fit(self, X, y):
m, n = X.shape
X = np.c_[np.ones((m, 1)), X]
labels = len(np.unique(y))
new_y = np.zeros((m, labels))
y = y.reshape(-1, 1)
for label in range(labels):
new_y[:, label: label + 1] = (y == label).astype(np.int)
all_theta = np.zeros((n + 1, labels))
if self.early_stopping:
X_train, X_val, y_train, y_val = train_test_split(X, new_y, test_size=0.2, random_state=2042)
best_theta = None
minimum_error = float("inf")
for i in range(self.num_iters):
h = self.softmax_function(X_train, all_theta)
gradients = (1 / m) * (X_train.T @ (h - y_train))
if self.penalty == "l2":
gradients[1:] += self.alpha * gradients[1:]
all_theta -= self.eta * gradients
val_cost = self.cross_entropy(X_val, y_val, all_theta)
if val_cost < minimum_error:
minimum_error = val_cost
best_theta = all_theta
self.best_theta = best_theta
else:
for i in range(self.num_iters):
h = self.softmax_function(X, all_theta)
gradients = (1 / m) * (X.T @ (h - new_y))
if self.penalty == "l2":
gradients[1:] += (self.alpha) * all_theta[1:]
all_theta -= self.eta * gradients
self.best_theta = all_theta
return all_theta
def predict(self, X):
X = np.c_[np.ones((len(X), 1)), X]
opt_theta = self.best_theta
preds = np.argmax(self.softmax_function(X, opt_theta), axis=1)
return preds
def predict_proba(self, X):
X = np.c_[np.ones((len(X), 1)), X]
opt_theta = self.best_theta
return self.softmax_function(X, opt_theta)
# +
# testing the algorithm on iris datatset.
from sklearn import datasets
iris = datasets.load_iris()
# -
X = iris["data"][:, (2, 3)]
y = iris["target"]
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.figure(figsize=(10, 5))
plt.plot(X[y==2, 0], X[y==2, 1], "g^", label="Iris virginica")
plt.plot(X[y==1, 0], X[y==1, 1], "bs", label="Iris versicolor")
plt.plot(X[y==0, 0], X[y==0, 1], "ro", label="Iris setosa")
# +
# splitting the data into train, test and validation set
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=2042)
X_tr, X_val, y_tr, y_val = train_test_split(X_train, y_train, test_size=0.2, random_state=2042)
# +
#training the model
soft_reg = SoftmaxReg(eta=0.01, num_iters=5001)
soft_reg.fit(X_tr, y_tr)
# +
# predicting on the validation set
val_preds = soft_reg.predict(X_val)
np.mean(y_val == val_preds)
# +
# trying l2 regularization
soft_reg_l2 = SoftmaxReg(eta=0.1, num_iters=5001, alpha=0.1, penalty="l2")
soft_reg_l2.fit(X_tr, y_tr)
# +
# predicting on validation set with l2 regularization.
val_preds = soft_reg_l2.predict(X_val)
np.mean((val_preds == y_val))
# +
# trying early stopping
soft_reg_early_stop = SoftmaxReg(eta=0.1, num_iters=5001, alpha=0.1, penalty="l2", early_stopping=True)
soft_reg_early_stop.fit(X_train, y_train)
# -
soft_reg_early_stop.best_theta
val_preds = soft_reg_early_stop.predict(X_val)
np.mean(val_preds == y_val)
# +
# predicting on test set
test_preds = soft_reg_early_stop.predict(X_test)
np.mean(test_preds == y_test)
# -
soft_reg = SoftmaxReg(eta=0.01, num_iters=5001)
soft_reg.fit(X, y.reshape(-1, 1))
# +
x0, x1 = np.meshgrid(
np.linspace(X[:, 0].min(), X[:, 0].max()),
np.linspace(X[:, 1].min(), X[:, 1].max())
)
x_new = np.c_[x0.ravel(), x1.ravel()]
preds = soft_reg_early_stop.predict(x_new)
zz = preds.reshape(x0.shape)
import matplotlib.pyplot as plt
plt.figure(figsize=(10, 4))
plt.plot(X[y==2, 0], X[y==2, 1], "g^", label="Iris virginica")
plt.plot(X[y==1, 0], X[y==1, 1], "bs", label="Iris versicolor")
plt.plot(X[y==0, 0], X[y==0, 1], "ro", label="Iris setosa")
plt.legend(loc="upper left")
plt.xlabel("petal length")
plt.ylabel("petal width")
plt.contourf(x0, x1, zz)
# -
| .ipynb_checkpoints/softmax_reg-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Science In One* Hour: Edge Detection
# \**for sufficiently large values of one*
#
# **<NAME>**
# **Data Science**
# **Spring 2020**
# **Assignment 1**
#
# For our first assignment, I've decided to write a very simple edge-detection algorithm for analyzing arbitrary images. It's not a particularly complicated implementation, but it's something I can put together in a short time fairly easily.
#
# This entire exercise took a few hours to complete, as it included getting my environment on tensor.hood.edu up and running, familiarizing myself with Jupyter notebooks, some light research on which libraries I'd want to use, and shifting mental gears from C# syntax (where I spend most of my time) to Python syntax (where I know what's going on but have considerably less experience than other languages.) While the guts of the algorithm took right around an hour, additional time was spent debugging, improving, and documenting this notebook.
#
# Stack Exchange helped considerably in the completion of this exercise.
#
# ## Supporting Libraries
#
# I'll be using the Computer Vision library (cv2) to load images, as image loading isn't the interesting part of what I'll be doing today.
#
# To install cv2, run:
#
# conda install -c anaconda opencv
#
# We'll also be using pyplot to show images in this notebook.
# +
import numpy as np
import cv2
import urllib
from matplotlib import pyplot as plt
# -
# ## User Options
#
# In the following block of code, there are options you can adjust to change how this script works.
#
# The `deltaThreshold` value allows you to adjust the sensitivity of the edge detection algorithm. Simply put, if the value of the current pixel is `deltaThreshold` more or less than that of the previous pixel, that pixel is marked as an edge pixel. Trial and error has shown 14 to be a generally good value for a variety of applications.
#
# the `pixelAdjustment` value defines what type of interpolation to apply to plotted images. Since this notebook's output is smaller than most of our source images, I've gone with `bilinear` interpolation as the default; this setting generates output pixels that are representative of the averages of multiple source pixels. The other recommended setting, `nearest` neighbor, uses a single pixel out of a group of source pixels to set the output pixel's value.
#
# Nearest neighbor will result in brighter, choppier edges that may not convey all edge data; bilinear will result in fainter but more accurate edge representations.
# +
deltaThreshold = 12
#pixelAdjustment = 'nearest'
pixelAdjustment = 'bilinear'
# -
# ## Dataset
#
# My dataset will be an image from the Internet. Specifically, I'll be using a photo of my dog, Latte. I've provided a few other examples from a variety of sources, as well; uncomment different `urllib.request.urlopen` lines below to change the image, or enter your own at the bottom.
# +
# a photo of my dog with strong edges
loadedRawData = urllib.request.urlopen('https://pbs.twimg.com/media/ENn1JwVXkAcpNjY?format=jpg&name=4096x4096').read()
############################
##### ALTERNATE IMAGES #####
############################
# a photo of my dog with weak edges
#loadedRawData = urllib.request.urlopen('https://pbs.twimg.com/media/EO5jMShXsAM5sMZ?format=jpg&name=4096x4096').read()
# Saturn V liftoff
#loadedRawData = urllib.request.urlopen('https://www.nasa.gov/sites/default/files/styles/side_image/public/62295main_liftoff_full.jpg?itok=3qa1O7vU').read()
# the Google Logo: an outstanding source for edge detection
#loadedRawData = urllib.request.urlopen('https://cdn.vox-cdn.com/thumbor/Pkmq1nm3skO0-j693JTMd7RL0Zk=/0x0:2012x1341/1200x800/filters:focal(0x0:2012x1341)/cdn.vox-cdn.com/uploads/chorus_image/image/47070706/google2.0.0.jpg').read()
# "Starry Night": a sub-optimal source for edge detection
#loadedRawData = urllib.request.urlopen('https://upload.wikimedia.org/wikipedia/commons/thumb/e/ea/Van_Gogh_-_Starry_Night_-_Google_Art_Project.jpg/757px-Van_Gogh_-_Starry_Night_-_Google_Art_Project.jpg').read()
# Choose your own image:
#loadedRawData = urllib.request.urlopen('{URL GOES HERE}').read()
# -
# Once I've loaded an image, I convert the loaded data into a cv2 image. I'll also convert to greyscale at this point to reduce dimensionality and make the data easier to work with.
#
# *Note that I'm doing a weird back-conversion from color to grayscale and back again. This lets us add edge data to the source image later; it also makes cv2 play nicely with the plotting library. There's some channel weirdness going on that I can fix with an ugly kludge easily enough, so.*
#
# *Note, too, that I'm using bilinear interpolation when displaying photos. I'm not good enough with Python and matplotlib yet to know how to easily resize these outputs, so I've turned on bilinear interpolation to ensure that all the edge data is at least visible in the outputs. With the default "nearest neighbor" interpolation, a lot of data points were being dropped, resulting in choppier-looking results that left out key bits of data.*
# +
sourceImage = np.asarray(bytearray(loadedRawData), dtype="uint8")
sourceImageColor = cv2.imdecode(sourceImage, cv2.COLOR_BGR2RGB)
sourceImage = cv2.cvtColor(sourceImageColor, cv2.COLOR_RGB2GRAY)
sourceImage = cv2.cvtColor(sourceImage, cv2.COLOR_GRAY2RGB)
sourceImageColor = cv2.cvtColor(sourceImageColor, cv2.COLOR_BGR2RGB)
plt.imshow(sourceImageColor, interpolation=pixelAdjustment)
plt.title("Source image")
plt.show()
plt.imshow(sourceImage, interpolation=pixelAdjustment)
plt.title("Source image (grayscale)")
plt.show()
# -
# We can use cv2's *shape* property to find the dimensions of the image, like so:
print("Loaded image dimensions: " + str(sourceImage.shape[0]) + "x" + str(sourceImage.shape[1]))
# ## Basic Edge Detection
#
# Now that we've loaded the image into an array of greyscale values, we can begin performing edge detection. First, we'll create a clone of the image to use as our destination image.
targetImage = np.asarray(bytearray(loadedRawData), dtype="uint8")
targetImage = cv2.imdecode(targetImage, cv2.COLOR_BGR2GRAY)
print("target image created: " + str(targetImage.shape[0]) + "x" + str(targetImage.shape[1]))
# Then, we get to our edge detection algorithm. This is a simple matter of scanning through each row of pixels in the image and determining how much the value of the current pixel has changed from the value of the previous pixel we examined. If the difference in pixel values exceeds a certain threshold (indicated in `deltaThreshold`,) then we count that as an edge pixel.
#
# I've added some supporting data structures that I may use in future to do things like automatically choose thresholds, but that doesn't happen in an hour.
# +
xMax = targetImage.shape[0]-1
yMax = targetImage.shape[1]-1
maxPixelValue = 0
minPixelValue = 255
pixelCounts = np.zeros(255)
xIndex = 0
yIndex = 0
previousPixel = 0
while(xIndex < xMax):
yIndex = 0
previousPixel = sourceImage[xIndex,yIndex,0];
while (yIndex < yMax):
delta = abs(int(previousPixel) - int(sourceImage[xIndex, yIndex,0]))
maxPixelValue = max(delta, maxPixelValue)
minPixelValue = min(delta, minPixelValue)
if (delta < deltaThreshold):
delta = 0
else:
delta = 254
targetImage[xIndex, yIndex,0] = delta
targetImage[xIndex, yIndex,1] = 0
targetImage[xIndex, yIndex,2] = 0
pixelCounts[delta] += 1
previousPixel = sourceImage[xIndex,yIndex,0];
yIndex += 1
xIndex += 1
print("maxPixelValue = " + str(maxPixelValue) + ", minPixelValue = " + str(minPixelValue))
print("max occurrance count: " + str(np.max(pixelCounts)));
plt.imshow(targetImage, interpolation=pixelAdjustment)
plt.title("Edge detection, horizontal pass")
plt.show()
# -
# After I finish my row scan, I perform a column scan. Note that I'm simply adding to the thresholds calculated in my first pass. This will allow me to rapidly identify values that were found in both the horizontal and vertical passes.
# +
maxPixelValue = 0
minPixelValue = 255
pixelCounts = np.zeros(255)
xIndex = 0
yIndex = 0
previousPixel = 0
while(yIndex < yMax):
xIndex = 0
previousPixel = sourceImage[xIndex,yIndex,0];
while (xIndex < xMax):
delta = abs(int(previousPixel) - int(sourceImage[xIndex, yIndex,0]))
maxPixelValue = max(delta, maxPixelValue)
minPixelValue = min(delta, minPixelValue)
if (delta < deltaThreshold):
delta = 0
else:
delta = 254
targetImage[xIndex, yIndex,0] += 0
targetImage[xIndex, yIndex,1] += delta
targetImage[xIndex, yIndex,2] += 0
pixelCounts[delta] += 1
previousPixel = sourceImage[xIndex,yIndex,0];
xIndex += 1
yIndex += 1
plt.imshow(targetImage, interpolation=pixelAdjustment)
plt.title("Edge detection, vertical + horizontal pass")
plt.show()
# -
# Having run both my horizontal and vertical pass, we can see the results of both passes on the same image. Green pixels indicate vertical pass hits, red pixels indicate horizontal pass hits, and yellow pixels indicate hits from both directions. By combining the two results, we get a pretty good edge detection result below:
# +
xIndex = 0
yIndex = 0
previousPixel = 0
while(xIndex < xMax):
yIndex = 0
while (yIndex < yMax):
if ((int(targetImage[xIndex, yIndex, 0]) + int(targetImage[xIndex, yIndex, 1]) + int(targetImage[xIndex, yIndex, 2])) < 250):
targetImage[xIndex, yIndex, 0] = 0
targetImage[xIndex, yIndex, 1] = 0
targetImage[xIndex, yIndex, 2] = 0
else:
targetImage[xIndex, yIndex, 0] = 254
targetImage[xIndex, yIndex, 1] = 254
targetImage[xIndex, yIndex, 2] = 254
yIndex += 1
xIndex += 1
# invert the image to make it look better
targetImage = cv2.bitwise_not(targetImage)
plt.imshow(targetImage, interpolation=pixelAdjustment)
plt.title("Edge detection, all hits")
plt.show()
# re-invert image for further processing below
targetImage = cv2.bitwise_not(targetImage)
# -
# Now, I can take my final basic edge detection and overlay it on top of the original color image. The algorithm clearly highlights edge features, but it also catches things like bold patterns, shadows and other noise. It's far from an optimal solution for edge detection, but it's a start.
# +
xIndex = 0
yIndex = 0
while(xIndex < xMax):
yIndex = 0
while (yIndex < yMax):
if (targetImage[xIndex, yIndex, 0] > 200):
sourceImage[xIndex, yIndex, 0] = 0
sourceImage[xIndex, yIndex, 1] = 254
sourceImage[xIndex, yIndex, 2] = 254
sourceImageColor[xIndex, yIndex, 0] = 0
sourceImageColor[xIndex, yIndex, 1] = 254
sourceImageColor[xIndex, yIndex, 2] = 254
yIndex += 1
xIndex += 1
plt.imshow(sourceImage, interpolation=pixelAdjustment)
plt.title("Original image (grayscale), edge detection hits in teal")
plt.show()
plt.imshow(sourceImageColor, interpolation=pixelAdjustment)
plt.title("Original image, edge detection hits in teal")
plt.show()
# -
# ## And that's it!
# That's my quick edge detection algorithm, all done in one* hour! Future work on this might include additional steps, such as are used in the Canny algorithm (which is where I drew initial inspiration for this exercise.) Things like Gaussian blurring, clustering, and line completion could all help build a much more powerful edge detector.
#
# *\*margin of error ±400%*
| Assignment 1 - Edge Detection in 1 Hour.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#dependecies
import pandas as pd
import numpy as np
import os
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import LabelEncoder, MinMaxScaler
from tensorflow.keras.utils import to_categorical
df = pd.read_csv(os.path.join('..', "arun/Resources/", 'diagnosis-of-covid-19-and-its-clinical-spectrum.csv'))
#Data pre-processing
data = df[['sars_cov_2_exam_result','patient_age_quantile', 'leukocytes', 'platelets', 'monocytes', 'hematocrit', 'eosinophils', 'red_blood_cells', 'hemoglobin', 'lymphocytes', 'mean_platelet_volume']]
feature_names = ['patient_age_quantile', 'leukocytes', 'platelets', 'monocytes', 'hematocrit', 'eosinophils', 'red_blood_cells', 'hemoglobin', 'lymphocytes', 'mean_platelet_volume']
y = data['sars_cov_2_exam_result'].values
X = data.drop('sars_cov_2_exam_result', axis=1).values
print(X.shape, y.shape)
#handle missing data
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(missing_values=np.nan, strategy='mean')
imputer.fit(X[:, 1:len(feature_names)])
X[:, 1:len(feature_names)] = imputer.transform(X[:, 1:len(feature_names)])
X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=42)
X_scaler = MinMaxScaler().fit(X_train)
X_train_scaled = X_scaler.transform(X_train)
X_test_scaled = X_scaler.transform(X_test)
#Label-encode data set
label_encoder = LabelEncoder()
label_encoder.fit(y_train)
encoded_y_train = label_encoder.transform(y_train)
encoded_y_test = label_encoder.transform(y_test)
# Convert encoded labels to one-hot-encoding
y_train_categorical = to_categorical(encoded_y_train)
y_test_categorical = to_categorical(encoded_y_test)
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Dense
# Creating model and adding layers
model = Sequential()
model.add(Dense(units=100, activation='relu', input_dim=10))
model.add(Dense(units=100, activation='relu'))
model.add(Dense(units=2, activation='softmax'))
# Compile and fit the model
model.compile(optimizer='adam',
loss='categorical_crossentropy',
metrics=['accuracy'])
model.summary()
model.fit(
X_train_scaled,
y_train_categorical,
epochs=60,
shuffle=True,
verbose=2
)
# +
#Quantifying the model
model_loss, model_accuracy = model.evaluate(
X_test_scaled, y_test_categorical, verbose=2)
print(
f"Normal Neural Network - Loss: {model_loss}, Accuracy: {model_accuracy}")
# -
#Make some predictions
encoded_predictions = model.predict_classes(X_test_scaled[:5])
prediction_labels = label_encoder.inverse_transform(encoded_predictions)
print(f"Predicted classes: {prediction_labels}")
print(f"Actual Labels: {list(y_test[:5])}")
#Save model
model.save('tensorflow_model.pkl')
#import model and make prediction
import tensorflow as tf
model_tensor = tf.keras.models.load_model("tensorflow_model.pkl")
prediction=model_tensor.predict_classes([[ 19, -1.288428, -0.906829, 0.567652, 0.694287, -0.835508, 0.578024, 0.541564, -0.295726, -0.325903]])
print(prediction)
print(model_tensor.predict_classes([[ 19, -1.288428, -0.906829, 0.567652, 0.694287, -0.835508, 0.578024, 0.541564, -0.295726, -0.325903]]))
| grant/tensorflow_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Final project, Numerical Analysis 2018-2019
#
#
# ## Project description
#
# In this project, we would like to compare the performance of some embarassingly simple algorithms to solve a classification problem based on the MNIST database.
#
# The abstract aim of the program is to write a function:
#
# ```
# result = classify(image)
# ```
#
# that takes as input a small grey scale image of a hand-written digit (from the MNIST database), and returns the digit corresponding to the content of the image.
#
# An example of the images we'll be working on is the following:
#
# 
#
# Some background on the MNIST database (from wikipedia):
#
#
# ## MNIST database
#
# *From Wikipedia, the free encyclopedia*
#
# The MNIST database (Modified National Institute of Standards and Technology database) is a large database of handwritten digits that is commonly used for training various image processing systems. The database is also widely used for training and testing in the field of machine learning. It was created by "re-mixing" the samples from NIST's original datasets. The creators felt that since NIST's training dataset was taken from American Census Bureau employees, while the testing dataset was taken from American high school students, it was not well-suited for machine learning experiments. Furthermore, the black and white images from NIST were normalized to fit into a 28x28 pixel bounding box and anti-aliased, which introduced grayscale levels.
#
# ## MNIST sample images.
#
# The MNIST database contains 60,000 training images and 10,000 testing images. Half of the training set and half of the test set were taken from NIST's training dataset, while the other half of the training set and the other half of the test set were taken from NIST's testing dataset. There have been a number of scientific papers on attempts to achieve the lowest error rate; one paper, using a hierarchical system of convolutional neural networks, manages to get an error rate on the MNIST database of 0.23%. The original creators of the database keep a list of some of the methods tested on it. In their original paper, they use a support vector machine to get an error rate of 0.8%. An extended dataset similar to MNIST called EMNIST has been published in 2017, which contains 240,000 training images, and 40,000 testing images of handwritten digits and characters.
#
# ## Algorithm
#
# We start by defining the distance between two images. Ideally, a distance function between two images is zero when the images are the same, and greater than zero when the images are different.
#
# The bigger the distance, the more different the images should be. Ideally, the distance between an image of the number `9` should be closer to an image of the number `8` than to an image of the number `1` (the digits `9` and `8`, as images, differ by the fact that the first has one closed loop, while the second has two closed loops, while the digit `1` is mostly a straight line). Two different images representing the same number should be even closer (i.e., the distance function should return a "small" number).
#
# Given a distance and a training set of images for which we know everything, the simplest algorithm we can think of to classify an image `z`, is the following: given a set of train images (`x_train`) for which we know the digit they represent (`y_train`), measure the distance between `z` and all images in `x_train`, and classify the image `z` to represent the same digit of the image that is closest to `z` in `x_train`:
#
# Parameters of the algorithm:
#
# - `x_train`
# - `y_train`
# - a distance function `dist`
#
# Input of the function
#
# - `z`
#
# Output of the function
#
# - `digit`
#
# where
#
# ```
# def classify(z):
# all_distances = array([dist(x, z) for x in x_train])
# digit = y_train[argmin(all_distances)]
# return digit
# ```
#
# We will experiment with different distances, and we will try to improve the algorithm above in a step by step fashon.
#
# ## Data description
#
# Each image in the MNIST dataset represents a hand written digit, in the form of a matrix of `28x28` values between zero and one, representing gray scale values (zero = white, one = black).
#
# We use an array of `60.000x28x28` floating point values to collect all training images, and an array of `60.000` digits containing the (correct) value of the training digits (between 0 and 9 inclusive).
#
# The testing images are instead collected into two arrays of size `10.000x28x28` and `10.0000` respectively.
# +
# %pylab inline
# # input image dimensions
# img_rows, img_cols = 28, 28
# # Uncomment the following lines if you have keras installed. Otherwise you can
# # use the file I uploaded: mnist.npz
#import keras
#from keras.datasets import mnist
# # the data, split between train and test sets
#(x_train, y_train), (x_test, y_test) = mnist.load_data()
# if K.image_data_format() == 'channels_first':
# x_train = x_train.reshape(x_train.shape[0], 1, img_rows, img_cols)
# x_test = x_test.reshape(x_test.shape[0], 1, img_rows, img_cols)
# input_shape = (img_rows, img_cols)
# else:
# x_train = x_train.reshape(x_train.shape[0], img_rows, img_cols)
# x_test = x_test.reshape(x_test.shape[0], img_rows, img_cols)
# input_shape = (img_rows, img_cols)
# x_train = x_train.astype('float32')
# x_test = x_test.astype('float32')
# x_train /= 255
# x_test /= 255
# np.savez_compressed('mnist.npz', x_train, y_train, x_test, y_test)
arc = load('mnist.npz')
x_train = arc['arr_0']
y_train = arc['arr_1']
x_test = arc['arr_2']
y_test = arc['arr_3']
print(x_train.shape, y_train.shape)
print(x_test.shape, y_test.shape)
# -
# # Plotting one image
#
# How do we plot the images? `pyplot`, which has been imported by the first line of the previous cell, contains a command called `imshow`, that can be used to plot images.
#
# In this case we know it is a greyscale image, with zero representing white and one representing black, so we use a colormap that goes from white to black, i.e., `gray_r` where `_r` stands for reversed.
# Show image number 15, and write in the title what digit it should correspond to
N=3
imshow(x_train[N], cmap='gray_r')
_ = title('Hand written digit '+str(y_train[N]))
# **IF YOU DON'T HAVE ENOUGH COMPUTATIONAL POWER, RUN THE EXERCISES ONLY UP TO WHAT IS SUSTAINABLE FOR YOUR PC**
#
# General guidelines:
#
# - Time all functions you construct, and try to make them run as fast as possible by precomputing anything that can be precomputed
# - Extra points are gained if you reduce the complexity of the given algorithms in any possible way, for example by exploiting linearity, etc.
# - If something takes too long to execute, make sure you time it on a smaller set of input data, and give estimates of how long it would take to run the full thing (without actually running it). Plot only the results you manage to run on your PC.
#
# # Assignment 1
#
# Implement the following distance functions
#
# - d_infty $$ d_{\infty}(a,b) := \max_{i,j} |b_{ij}-a_{ij}|$$
# - d_one $$ d_1(a,b) := \sum_{i,j} |b_{ij}-a_{ij}|$$
# - d_two $$ d_2(a,b) := \sqrt{\sum_{i,j} |b_{ij}-a_{ij}|^2}$$
#
# that take two `(28,28)` images in input, and return a non-negative number.
#
# # Assignment 2
#
# Write a function that, given a number `N`, and a distance function `dist`, computes the distance matrix D of shape `(N,N)` between the first `N` entries of `x_train`:
#
# ```
# D[i,j] = dist(x_train[i], x_train[j])
# ```
#
# performing the **minimum** number of operations (i.e., avoid computing a distance if it has already been computed before, i.e., keep in mind that dist(a,b) = dist(b,a)).
#
# # Assignment 3
#
# Compute and plot the three distance matrices
#
# - Dinfty
# - D1
# - D2
#
# for the first 100 images of the training set, using the function `imshow` applied to the three matrices
#
# # Assignment 4
#
# Using only a distance matrix, apply the algorithm described above and compute the efficency of the algorithm, i.e., write a function that:
#
# Given a distance matrix with shape `(N,N)`, constructed on the first `N` samples of the `x_train` set, count the number of failures of the **leave one out** strategy, i.e.,
#
# - set `error_counter` to zero
#
# - for every line `i` of the matrix:
#
# - find the index `j` (different from `i`) for which `D[i,k] >= D[i,j]` for all `k` different from `i` and `j`.
#
# - if `y_train[j]` is different from `y_train[i]`, increment by one `error_counter`
#
# - return the error: error_counter/N
#
# - apply the function above to the 3 different distance matrices you computed before
#
#
# # Assignment 5
#
# Run the algorithm implemented above for N=100,200,400,800,1600 on the three different distances, and plot the three error rate as a function of N (i.e., compute the distance matrix, and compute the efficiency associated to the distance matrix).
#
# You should get an error like:
# ```
# [[ 0.58 0.17 0.17 ]
# [ 0.52 0.145 0.135 ]
# [ 0.4425 0.15 0.135 ]
# [ 0.4 0.145 0.12875 ]
# [ 0.369375 0.1025 0.09375 ]]
# ```
# where each column represents a different norm.
#
# ** In the next assignments, optional points are given if you manage to make the algorithm run faster, by pre-computing everything you can precompute in advance **
#
# # Assignment 6
#
# In principle, it should be possible to decrease the error by using a better norm. From the table above, it is clear that the L2 distance works better than the L1 distance, which works better than the Linfty distance.
#
# However, *none of these distances exploit the fact that the image is a two-dimensional object*, and that there is information also in the **neighboring** information of the pixels.
#
# One way to exploit this, is to interpret the image as a continuous function with values between zero and one, defined on a square domain `\Omega=[0,27]x[0,27]`.
#
# $$ f: \Omega \to R $$
#
# - Implement a function that computes an approximation of the $H^1$ norm distance on the renormalized images. Given two images $f_1$ and $f_2$
# - Compute $$a = \frac{f_1}{\int_\Omega f_1}$$, $$b=\frac{f_2}{\int_\Omega f_2}$$
# - Define the $H^1$ distance as
# $$
# d_{H^1}(f_1,f_2) := \sqrt{\int_\Omega |\nabla(a-b)|^2+ (a-b)^2}
# $$
# using the algorithm you prefer (or the library you prefer) to compute the gradients and the integrals. Notice that $\nabla f = (\partial f/\partial x, \partial f/\partial y)$ is a vector valued function, and $|\nabla g|^2 := (\partial g/\partial x)^2 + (\partial g/\partial y)^2$
#
# - Compute the distance matrix and the efficiency for this distance for N=100,200,400,800,1600
#
# ## Assignment 7
#
# An even better improvement on the previous distance function is given by the following algorithm
#
# - Given two images $f1$ and $f2$:
# - Compute $$a = \frac{f_1}{\int_\Omega f_1}$$, $$b=\frac{f_2}{\int_\Omega f_2}$$
# - Solve
# $$
# -\Delta \phi = a - b \qquad \text{ in } \Omega
# $$
# $$
# \phi = 0 \text{ on } \partial\Omega
# $$
# - Define the *Monge Ampere* distance
# $$
# d_{MA}(f_1,f_2) = \int_\Omega (a+b)|\nabla \phi|^2
# $$
#
# - Compute the distance matrix and the efficiency for this distance for N=100,200,400,800,1600
#
# ## Assigment 8 (optional for DSSC, PhD and LM, Mandatory for MHPC)
#
# Use the `BallTree` algorithm (https://en.wikipedia.org/wiki/Ball_tree), from the `sklearn` package, and construct a tree data structure **that uses one of the custom distances defined above**.
#
# For each N in 3200,6400,12800,25600,51200, and for each distance defined above
#
# - Build a tree using the first N entries of the training set `x_train`
# - Construct a function that tests the efficiency on all the entries of the test set `x_test`:
# - for any image in `x_test`, call it `x_test[i]`, query the tree for the nearest neighbor (call it `k`), and assign as predicted digit the digit of the `x_train[k]` image, i.e., `y_train[k]`
# - check if `y_train[k]` is equal to the corresponding entry in `y_test[i]`. If not, increment a counter of the error
# - return the efficiency, i.e., `error_counter/len(x_test)`
# - Plot, in a single graph, the error of each distance as a function of `N` (including labels, titles, etc.)
#
# - Once you have the tree, experiment with different nearest neighbor algorithms, i.e., instead of taking only one nearest neighbor, take a larger number (a small number of your choice), and instead of returning the single closest digit, return the one with the largest number of occurrences. Plot the same graph you gave before, and see if you gain an improvement. Motivate all choices you have to make to get to the final answer.
#
#
# **IF YOU DON'T HAVE ENOUGH COMPUTATIONAL POWER, RUN THE EXERCISES ONLY UP TO WHAT IS SUSTAINABLE FOR YOUR PC**
# +
# Assignement 1
#The whole program needs at least a 6 GB RAM machine
from numpy import linalg
def d_infy(im1,im2):
''' Calculates the l-infinity norm'''
return np.max(np.abs(im2 - im1))
#slower
#return np.linalg.norm(np.ravel(im2) - np.ravel(im1),inf)
def d_one(im1,im2):
''' Calculates l-one norm'''
return np.sum(np.abs(im2 - im1))
#return np.linalg.norm(np.ravel(im2) - np.ravel(im1),1)
def d_two(im1,im2):
''' Calculates the l-2 norm'''
return sqrt(np.sum((im2 - im1)**2))
#return np.linalg.norm(np.ravel(im2) - np.ravel(im1),2)
# +
# Assignement 2
def calc_train_dist(N,t_set,dist_fun):
'''Calculates the distance matrix'''
final = np.zeros((N,N))
for n in range(N):
for k in range(n+1,N):
final[n,k] = dist_fun(t_set[n,:,:],t_set[k,:,:])
return final + final.T
def calc_train_dist_ind(N,dist_fun,norm_set,grad_set):
'''Calculate the distance matrix with indexes'''
final = np.zeros((N,N))
for n in range(N):
for k in range(n+1,N):
final[n,k] = dist_fun(n, k, norm_set, grad_set)
return final + final.T
# +
# Assignement 3
#A preview of the similarity matrix
# %time Dinf = calc_train_dist(100, x_train, d_infy)
# %time Done = calc_train_dist(100, x_train, d_one)
# %time Dtwo = calc_train_dist(100, x_train, d_two)
imshow(Dinf)
_ = title('Dinf distance matrix ')
show()
imshow(Done)
_ = title('Done distance matrix')
show()
imshow(Dtwo)
_ = title('Dtwo distance matrix')
show()
# +
# Assignement 4
def LOOC(d_mat,res_train, N = 100):
'''Calculates the error of the NN classifier'''
error_counter = 0
mat = d_mat[:N,:N].copy()
for i in range(N):
#exclude the distance with himself, and correct the resulting index
ind = np.argmin(mat[i,np.arange(mat.shape[1]) != i])
if ind >= i:
ind += 1
if res_train[ind] != res_train[i]:
error_counter += 1
return error_counter/N
# %time cross_Done = LOOC(Done, y_train)
# %time cross_Dtwo = LOOC(Dtwo, y_train)
# %time cross_Dinf = LOOC(Dinf, y_train)
# -
# Assignement 5.1
DN = 1600
# precomputation of the distance matrices
# %time Dinf2 = calc_train_dist(DN, x_train, d_infy)
# %time Done2 = calc_train_dist(DN, x_train, d_one)
# %time Dtwo2 = calc_train_dist(DN, x_train, d_two)
# +
# %time
# Assignement 5.2
# Errors for different norms
result_dist = np.zeros((5,3))
Num = [100,200,400,800,1600]
for i in range(5):
result_dist[i,0] = LOOC(Done2, y_train, N = Num[i])
result_dist[i,1] = LOOC(Dtwo2, y_train, N = Num[i])
result_dist[i,2] = LOOC(Dinf2, y_train, N = Num[i])
plot(Num, result_dist)
_ = legend(["Done","Dtwo","Dinf"])
# -
# Assignement 5.3
result_dist
# +
# Assignement 6.1
def normalize(im):
'''Returns the image normalized by its integral'''
return im.reshape(28,28)/np.sum(im)
def precalc_grad(data_set):
'''precalculate the gradient'''
fin = [np.zeros((2,28,28))] * len(data_set)
for n in range(len(fin)):
fin[n]= np.gradient(data_set[n])
return fin
def precalc_norm(data_set):
'''precalculate the normalized images'''
fin = [np.zeros((28,28))] * data_set.shape[0]
for n in range(len(fin)):
fin[n] = normalize(data_set[n])
return fin
def d_H1(ind1,ind2, norm_set,grad_table):
'''H1 distance'''
#Just use index and precomputed values
ind1 = int(ind1)
ind2 = int(ind2)
a1,a2 = grad_table[ind1][0], grad_table[ind1][1]
b1,b2 = grad_table[ind2][0],grad_table[ind2][1]
return sqrt(np.sum(((a1-b1)**2 + (a2-b2)**2) + (norm_set[ind1]-norm_set[ind2])**2))
#I also precalculate those of the test to use them in the balltree
norms = precalc_norm(np.vstack((x_train,x_test)))
grads = precalc_grad(norms)
len(norms)
# %time DH1_p = calc_train_dist_ind(100, d_H1,norms,grads)
imshow(DH1_p)
_ = title('DH1 distance matrix')
# +
# Assignement 6.2
#errors for H1 norm
# %time DH1 = calc_train_dist_ind(DN, d_H1,norms, grads)
result_dist2 = np.zeros((5,))
for i in range(5):
result_dist2[i] = LOOC(DH1, y_train, N = Num[i])
plot(Num, result_dist2)
# +
# Assignement 7.1
from scipy.sparse import diags, csc_matrix
from scipy.sparse.linalg import splu
from scipy.linalg import lu_solve
def build_laplacian(N):
'''construct the matrix for finite differences approximation of the laplacian'''
mat = diags([1,1,-4,1,1],[-N,-1,0,1,N], shape = (N*N,N*N)).toarray()
for i in range(0,N*N,N):
mat[i,i-1] = 0
mat[i-1,i] = 0
return mat
lap = build_laplacian(26)
#use sparse matric format
lu = splu(csc_matrix(lap))
def precalc_lapl(norm_set,lu):
'''Precalculates the different functions phi'''
fin = [np.zeros((28,28))] * len(norm_set)
for n in range(len(fin)):
phi = np.zeros((28,28))
phi[1:27,1:27] = lu.solve((norm_set[n][1:27,1:27].ravel())).reshape((26,26))
fin[n] = phi
return fin
def d_ma(ind1,ind2, norm_set,grad_set2):
'''Morge-Ampere distance with precalculated phi'''
#thi is required when we use indexes from the balltree
ind1 = int(ind1)
ind2 = int(ind2)
a = norm_set[ind1]
b = norm_set[ind2]
aphi1d, aphi2d = grad_set2[ind1][0], grad_set2[ind1][1]
bphi1d, bphi2d = grad_set2[ind2][0], grad_set2[ind2][1]
return np.sum(((aphi1d-bphi1d)**2 + (aphi2d-bphi2d)**2)*(a+b))
#precalc phi and gradients
lapl = precalc_lapl(norms,lu)
grads2 = precalc_grad(lapl)
del lapl
# %time DMA = calc_train_dist_ind(100, d_ma,norms, grads2)
imshow(DMA)
_ = title('DMA distance matrix')
# +
# Assignement 7.2
#errors for Morge-Ampere distance
# %time DMA = calc_train_dist_ind(DN, d_ma,norms, grads2)
result_dist3 = np.zeros((5,))
for i in range(5):
result_dist3[i] = LOOC(DMA, y_train, N = Num[i])
plot(Num, result_dist3)
# +
# Assignement 8.1
# Basically I tried during the exercise to precompute as much as possible,
# as a result this scripts are wasting a lot of memory.
# The BallTree computation is pretty intensive and we are in the order of hours,
# the last part is a better implementation using the wrapper functions in scikit learn.
# The bigger advantage of the KNeighborsClassifier() consists in its parallelization;
# however my pc has only a mid-range-2015 two core CPU, so it does not benefit at all from
# the multicore support(even though I think it can make the difference on a bigger machine)
# To gain data in a suitable time I have used only 1000 train samples
from sklearn.neighbors import BallTree
def Ballfunctions(i,j):
'''Construct different balltrees for the different metrics'''
if(i < 3):
return BallTree(x_train[:j,:,:].reshape(j,28*28),metric="pyfunc", func = DistanceList[i])
elif (i == 3):
return BallTree(np.arange(j).reshape(-1,1),metric="pyfunc",
func = DistanceList[i],norm_set = norms,grad_table = grads)
else:
return BallTree(np.arange(j).reshape(-1,1),metric="pyfunc",
func = DistanceList[i],norm_set = norms,grad_set2 = grads2)
Num = [3200,6400,12800,25600,51200]
DistanceList = [d_one, d_two, d_infy, d_H1, d_ma ]
#create a nested dict with key = distance and value a dict with key = number of samples and the balltree
DistanceDict = {k:v for k,v in zip(("D1","D2","Dinf","DH1","DMA"), [{}] * 5)}
for i,k in enumerate(DistanceDict.keys()):
BallTrees = [Ballfunctions(i,j) for j in Num]
DistanceDict[k] = {k:v for k,v in zip(Num,BallTrees)}
# +
# Assignement 8.2
def ball_LOOCV(res_train,data_test,res_test,dlist,Nlist, k=1, indx = False):
'''Classification error of NN for the ball tree, when k > 1, we use the mode of the neighbors'''
error_counter_list = []
for n in Nlist:
error_counter = 0
if k == 1:
for i in range(0,len(res_test),10):
idx = 0
# change from index two images distance to precalculated
if indx:
idx = dlist[n].query(np.array([i + 60000]).reshape(1,-1),return_distance = False)[0][0]
else:
idx = dlist[n].query(data_test[i].reshape(1,-1),return_distance = False)[0][0]
if res_train[idx] != res_test[i]:
error_counter += 1
error_counter_list.append(error_counter/(len(res_test)/10))
else:
for i in range(0,len(res_test),10):
idx = 0
if indx:
idx = dlist[n].query(np.array([i + 60000]).reshape(1,-1),return_distance = False, k = k)[0]
else:
idx = dlist[n].query(data_test[i].reshape(1,-1),return_distance = False, k =k)[0]
#take the mode
counts = np.bincount(res_train[idx])
num = np.argmax(counts)
if num != res_test[i]:
error_counter += 1
error_counter_list.append(error_counter/(len(res_test)/10))
return error_counter_list
# -
#Pretty slow
plot_list = []
for i,dist in enumerate(("D1","D2","Dinf","DH1","DMA")):
flag = False
if dist == "DH1" or dist == "DMA":
flag = True
# %time plot_list.append(ball_LOOCV(y_train,x_test,y_test,DistanceDict[dist],Num,1, flag))
plot(Num,np.array(plot_list).T)
_ = title("KNN with K=1")
_ = legend(["D1","D2","Dinf","DH1","DMA"])
# Assignement 8.3
#senza dubbio non è molto elegante
plot_list2 = []
for i,dist in enumerate(("D1","D2","Dinf","DH1","DMA")):
flag = False
if dist == "DH1" or dist == "DMA":
flag = True
# %time plot_list2.append(ball_LOOCV(y_train,x_test,y_test,DistanceDict[dist],Num,100, flag))
plot(Num,np.array(plot_list2).T)
_ = title("KNN with K=100")
_ = legend(["D1","D2","Dinf","DH1","DMA"])
# Assignement 8 bis
#Attention this snippet uses all of the available cores, set n_jobs accordingly
#This is the same thing as before, just using the wrapper interface from sklearn
from sklearn.neighbors import BallTree
from sklearn.neighbors import KNeighborsClassifier
def Ballfunctions(i,j,k = 1):
'''Construct different balltrees for the different metrics'''
if(i < 3):
return KNeighborsClassifier(n_neighbors=k, leaf_size = 100,
algorithm = "ball_tree",
metric= DistanceList[i], n_jobs = -1).fit(x_train[:j,:,:].reshape(j,28*28),y_train[:j])
elif (i == 3):
return KNeighborsClassifier(n_neighbors=k, leaf_size = 100,
algorithm = "ball_tree",
metric = DistanceList[i], metric_params = {
"norm_set" : norms,
"grad_table" : grads}, n_jobs = -1).fit(np.arange(j).reshape(-1,1), y_train[:j])
else:
return KNeighborsClassifier(n_neighbors=k, leaf_size = 100,
algorithm = "ball_tree",
metric = DistanceList[i],
metric_params = {"norm_set" : norms,
"grad_set2" : grads2}, n_jobs = -1).fit(np.arange(j).reshape(-1,1), y_train[:j])
Num = [3200,6400,12800,25600,51200]
DistanceList = [d_one, d_two, d_infy, d_H1, d_ma ]
#create a nested dict with key = distance and value a dict with key = number of samples and the balltree
DistanceDict = {k:v for k,v in zip(("D1","D2","Dinf","DH1","DMA"), [{}] * 5)}
for i,k in enumerate(DistanceDict.keys()):
BallTrees = [Ballfunctions(i,j) for j in Num]
DistanceDict[k] = {k:v for k,v in zip(Num,BallTrees)}
plot_list = []
for i in ("D1","D2","Dinf","DH1","DMA"):
dist_list = []
if i == "D1" or i == "D2" or i == "Dinf":
for n in Num:
# %time dist_list.append(DistanceDict[i][n].score(x_test.reshape(-1,28*28), y_test))
else:
for n in Num:
# %time dist_list.append(DistanceDict[i][n].score(np.arange(60000,70000).reshape(-1,1), y_test))
plot_list.append(dist_list)
| final_project/final_project_2018-2019_SALVATORE_MILITE.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import sympy
x, y = sympy.symbols('x y')
# -
# - $\alpha$ (그리스 문자 알파), a (영어)
# - $\beta$ (그리스 문자 베타), b (영어)
# - $\gamma$ (그리스 문자 감마), r (영어)
# - $\epsilon$ (그리스 문자 엡실론), e (영어)
# - $\kappa$ (그리스 문자 카파), k (영어)
# - $\nu$ (그리스 문자 뉴), v (영어)
# - $\rho$ (그리스 문자 로), p (영어)
# - $\sigma$ (그리스 문자 시그마), 6 (숫자)
# - $\omega$ (그리스 문자 오메가), w (영어)
# ## Numpy를 활용한 선형대수 입문 (김도형 박사님 블로그) - datascienceshool.net에 가면 엄청 도움됨!
# - 선형대수 (linear Algebra)는 데이터 분석에 필요한 각종 계산을 돕기 위한 학문이다. 데이터 분석을 하려면 수 많은 숫자로 이루어진 데이터를 다루어야 한다. 하나의 데이터가 수십 개에서 수만 개의 숫자로 이루어져 있을 수도 있고, 또 이러한 데이터 수만 개가 하나의 집합을 이루고 있을 수도 있다.
# - 선형대수를 사용하면 대량의 데이터를 포함하는 복잡한 계산 과정을 몇 글자 되지 않는 간단한 수식으로 서술할 수 있다. 따라서, 데이터를 다루는 과정을 정확하고 간단하게 서술할 수 있다. 이를 위해 우선 선형대수에서 사용되는 여러 가지 기호와 개념을 익혀보자.
# ## Data Type
# - 선형대수에서 다루는 데이터는 개수나 형태에 따라 크게 **스칼라(scalar), 벡터(vector), 행렬(matrix)**의 세 가지 유형으로 나누어진다. 간단하게 말하자면 스칼라는 숫자 하나로 이루어진 데이터이고, 벡터는 여러 개의 숫자로 이루어진 데이터 레코드(data record)이며, 행렬은 이러한 벡터, 즉 데이터 레코드가 여러 개 있는 데이터 집합이라고 볼 수 있다.
# ## 스칼라 (Scalar)
# - 스칼라는 하나의 숫자만으로 이루어진 데이터를 말한다. 예를 들어 어떤 붓꽃 한 송이의 꽃잎 길이를 측정하면 숫자가 하나 나올 것이다. 이 데이터는 스칼라이다. 스칼라는 보통 x와 같이 알파벳 소문자로 표기하며 실수(real number)인 숫자 중의 하나이므로 실수 집합 R의 원소라는 의미에서 다음처럼 표기한다.
# - x $\in$ R
# ## 벡터 (Vector)
# - 벡터는 여러 개의 숫자가 특정한 순서대로 모여 있는 것을 말한다. 사실 대부분의 데이터 레코드는 여러 개의 숫자로 이루어진 경우가 많다. 예를 들어 붓꽃의 종을 알아내기 위해 크기를 측정할 때, 꽃잎의 길이 x1뿐 아니라 꽃잎의 폭 x2, 꽃 받침의 길이 x3, 꽃받침의 폭 x4라는 4개의 숫자를 측정할 수도 있다. 이렇게 측정된 4개의 숫자는 한 송이의 붓 꽃에서 나온 데이터이므로 따로따로 다루기보다는 하나의 쌍으로 묶어 놓는 것이 좋다. 이때 숫자의 순서가 바뀌면 어떤 숫자가 꽃잎의 길이이고 어떤 숫자가 꽃받침의 폭인지 알 수 없으므로 숫자의 순서를 유지하는 것이 중요하다. 이런 데이터 묶음을 선형대수에서는 벡터라고 부른다.
# - 붓꽃의 예에서는 4개의 데이터(x1, x2, x3, x4)가 하나로 묶여 있는데, 이를 선형 대수 기호로는 다음 처럼 하나의 문자 x로 표기한다.
# - x = $[[x1],[
# x2],
# [x3],
# [x4]]$
# - 이 때 벡터는 복수의 가로줄, 즉 행(row)을 가지고 하나의 세로줄, 즉 열(column)을 가지는 형태로 위에서 아래로 내려써서 표기해야 한다.
# - 하나의 벡터를 이루는 데이터의 개수를 차원(dimension)이라고 한다. 위에서 예로 든 x는 4개의 실수로 이루어져 있으므로 4차원 실수 벡터라고 한다.
# - 벡터의 원소가 되는 스칼라 값음 1, 2 등의 자연수 아래 첨자(sub-script)를 붙여서 원소의 위치를 표시한다. 하지만, 자연수 아래 첨자가 되어 있다고 무조건 스칼라는 아니다. 마찬가지로 벡터가 여러 개 있으면 서로 다른 벡터를 구별하기 위해 벡터 이름에 아래 첨자를 붙일 수도 있다. 따라서 아래 첨자를 가진 알파벳 소문자 기호는 스칼라일 수도 있고 벡터일 수도 있다. 두 경우는 문맥에 따라 구별해야 한다. 때에 따라서는 스칼라가 아닌 벡터라는 점을 강조하기 위해 볼드체 기호 **x**를 사용하기도 한다.
# - 예를 들어 어떤 붓꽃 표본 한 송이를 꺽어 측정하였더니, 꽃잎의 길이가 5.1cm, 꽃잎의 폭이 3.5cm, 꽃받침의 길이가 1.5cm, 꽃받침의 폭이 0.2cm 였다면 이 데이터 쌍을 x1이라고 이름 붙이고 다음처럼 나타낼 수도 있다.
# - x1 = [5.1\n3.5\n1.4\n0.2]
# - x2 = [4.9\n3.0\n1.4\n0.2]
# - 만약 이 데이터를 이용하여 붓꽃의 종을 결정하는 예측 문제를 풀고 있다면 이러한 데이터 벡터를 특징벡터(feature vector)이라고 한다. Numpy를 사용하여 벡터를 표현할 떄는 벡터를 열의 개수가 하나인 2차원 배열 객체로 표현하는 것이 올바르다. 예를들어 위의 벡터 x1을 numpy로 표기하면 다음과 같다.
x1 = np.array([[5.1], [3.5], [1.4], [0.2]])
x1
# - 하지만 Numpy는 1차원 배열 객체도 대부분 벡터로 인정한다. 이 때는 벡터가 마치 하나의 행처럼 표시되어도 실제로는 열이라는 점에 주의한다.
x1 = np.array([5.1, 3.5, 1.4, 0.2])
x1
# - 그러나 벡터 데이터를 처리하는 프로그램에 따라서 두 가지 표현법 중 열 표기를 정확하게 요구하는 경우도 있으므로 주의해야 한다. 예를 들어 Scikit-learn 패키지에서 벡터를 요구하는 경우에는 반드시 열의 개수가 1개인 2차원 배열 객체를 넣어야 한다.
# #### 연습 문제 1
# - Numpy를 사용해서 다음에 제시한 붓꽃 데이터 x2에 대한 벡터 변수 x2를 만든다.
# - x2 = [4.9\n3.0\n1.4\n0.2]
x2 = np.array([[4.9], [3.0], [1.4], [0.2]])
x2
# - 예측 문제의 입력 데이터는 대부분 벡터로 표시한다. 예를 들어 숫자 이미지를 입력받아 어떤 숫자인지 분류하는 문제에서 이미지는 원래 2차원 데이터이지만 1차원 벡터로 변환할 수 있다. 다음은 8 x 8 차원의 숫자 0과 1 이미지 4개를 64차원의 벡터로 펼친 것이다. 같은 숫자에 대한 벡터가 서로 닮았다는 점에 주의하라.
# +
from sklearn.datasets import load_digits
digits = load_digits()
d0 = digits.images[0]
d1 = digits.images[10]
d2 = digits.images[1]
d3 = digits.images[11]
f, ax = plt.subplots(1, 4)
ax[0].imshow(d0, interpolation = 'nearest', cmap=plt.cm.Blues)
ax[0].grid(False)
ax[1].imshow(d1, interpolation = 'nearest', cmap=plt.cm.Blues_r)
ax[1].grid(False)
ax[2].imshow(d2, interpolation = 'nearest', cmap=plt.cm.Blues)
ax[2].grid(False)
ax[3].imshow(d3, interpolation = 'nearest', cmap=plt.cm.Blues_r)
ax[3].grid(False)
plt.show()
# +
v0 = d0.reshape(64, 1)
v1 = d1.reshape(64, 1)
v2 = d2.reshape(64, 1)
v3 = d3.reshape(64, 1)
f, ax = plt.subplots(4, 1, figsize=(12,2))
ax[0].imshow(v0.T, interpolation='nearest', cmap=plt.cm.Blues)
ax[0].grid(False); ax[0].set_xticks([]); ax[0].set_yticks([])
ax[1].imshow(v1.T, interpolation='nearest', cmap=plt.cm.Blues)
ax[1].grid(False); ax[0].set_xticks([]); ax[0].set_yticks([])
ax[2].imshow(v2.T, interpolation='nearest', cmap=plt.cm.Blues)
ax[2].grid(False); ax[0].set_xticks([]); ax[0].set_yticks([])
ax[3].imshow(v3.T, interpolation='nearest', cmap=plt.cm.Blues)
ax[3].grid(False); ax[0].set_xticks([]); ax[0].set_yticks([])
plt.tight_layout()
plt.show()
# -
# ## 행렬 (Matrix)
#
# - 행렬은 복수의 차원을 가지는 데이터 레코드가 다시 여러 개 있는 경우의 데이터를 합쳐서 표기한 것이다. 예를 들어 앞서 말한 붓꽃 예에서 6개의 붓꽃에 대해 크기를 측정하였다면 4차원 붓꽃 데이터가 6개가 있다. 즉, 4 X 6 = 24개의 실수 숫자가 있는 것이다.
#
# - 행렬의 우너소 하나하나는x2,3처럼 두 개의 숫자 쌍을 아래첨자(sub-script)로 붙여서 표기한다. 첫 번째 숫자가 행을 뜻하고 두 번째 숫자가 열을 뜻한다. 예를 들어 x2,3는 두 번째 행(위에서 아래로 두번째), 세번째 열(왼쪽에서 오른쪽으로 세번째)의 숫자를 뜻한다. 행/열의 개수가 10보다 적을떄는 쉼표 없이 x23라고 표기할 때도 있다.
# - 벡터는 열의 수가 1인 행렬이라고 볼 수 있으므로 벡터를 다른 말로 **열 벡터(column vector)**라고도 한다.
# - 데이터를 행렬로 묶어서 표시할 때는 붓꽃 하나에 대한 데이터 레코드, 즉 하나의 벡터가 열이 아닌 행(row)으로 표시한다. 붓꽃의 예에서는 하의 데이터 레코드가 4차원 데이터 였다는 점을 기억하다.
#
# - **하나의 데이터 레코드를 단독으로 벡터로 나타낼 때는 하나의 열(column)** 나타내고 **복수의 데이터 레코드 집합을 행렬로 나타낼 때는 하나의 데이터 레코드가 하나의 행(row)**으로 표기하는 것은 얼핏 보기에는 일관서잉 없어 보이지만, 추후 다른 연산을 할 때 이런 모양이 필요하기 때문이다. 데이터 분석에서 쓰는 일반적인 관례이므로 외워두어야 한다.
# - 만약 이 데이터를 이용하여 붓꽃의 종을 결정하는 예측 문제를 풀고 있다면 이 행을 특징 행렬(feature matrix)이라고 하기도 한다.
# - 앞에서 예로 들었던 두 송이의 붓꽃 데이터를 하나의 행렬로 합친 Numpy는 아래와같다.
A = np.array([[11,12,13],[21,22,23]])
A
# #### 연습 문제 2
# - Numpy를 사용해서 다음에 제시한 붓 꽃 데이터 X에 대한 행렬 변수 x를 만든다.
x = np.array([[5.1, 3.5, 1.4, 0.2], [4.9, 3.0, 1.4, 0.2]])
x
# ## 전치 연산 (Transpose)
# - 이번에는 이러한 스칼러, 벡터, 행렬 데이터를 변형시키는 방법 즉, 연산(operation)에 대해서 알아보자. **전치(Transpose)** 연산은 행렬에서 가장 기본이 되는 연산으로 행렬의 **행과 열을 바꾸는 연산**을 말한다. 전치 연산은 벡터나 행렬에 T라는 위첨자(super-script)를 붙여서 표기한다. 예를 들어 앞에서 보인 6 x 4 차원의 행렬을 전치 연산하면 4 x 6 차원의 행렬이 된다.
# - 전치 연산으로 만든 행렬을 원래 행렬에 대한 전치행렬이라고 한다. (열)벡터 x에 대해 전치 연산을 적용하여 만든 x transpose는 행의 수가 1인 행렬이므로 행 벡터(row vecotr)이라고 한다.
# - Numpy에서는 ndarray 객체의 T라는 속성을 이용하여 전치 행렬을 구한다. 이 때 T 메서드(method)가 아닌 속성(attribute)이므로 소괄호 ()를 붙여서 호출하면 안된다.
A.T
# - 다만 1차원 ndarray는 전치 연산이 정의되지 않는다.
x1
x1.T
# #### 연습문제 3
# - 1. Numpy를 사용해서 다음에 제시한 붓꽃 데이터 X의 전치행렬 X Transpose를 구한다.
# - 2. 전치행렬을 다시 전치한 행렬 (X transpose) of transpose를 구한다. 이 행렬과 원래 행렬X를 비교한다.
X = np.array([[5.1, 3.5, 1.4, 0.2], [4.9, 3.0, 1.4, 0.2]])
X.T
X.T.T
# ## 특수한 벡터와 행렬
# - 몇가지 특수한 벡터와 행렬은 별도의 기호나 이름이 붙는다.
#
# ### 영 벡터
# - 모든 원소가 0인 N차원 벡터이다.
# ### 일 벡터
# - 모든 원소가 1인 N차원 벡터이다.
# ### 정방 행렬
# - 행의 개수와 열의 개수가 같은 행렬을 정방 행렬(square Matrix)이라고 한다.
# ### 대각 행렬
# - 행렬에서 행과 열이 같은 위치를 주 대각 또는 간단히 대각(diagonal)이라고 한다. 대각 위치에 있지 않는 것들은 비대각이라고 한다. 모든 비대각 요소가 0인 행렬을 대각 행렬이라고 한다.
# - 대각 행렬이 되려면 대각 성분이 0이기만 하면 된다. 반드시 정방행렬일 필요는 없다.
# - Numpy로 대각 정방 행렬을 생성하려면 diag 명령을 사용한다.
np.diag([1,2,3])
# ### 단위행렬 (Identity Matrix)
# - 대각 행렬 중에서도 모든 대각 성분의 값이 1인 대각행렬을 단위 행렬이라고 한다. 단위 행렬은 보통 알파벳 대문자 I로 표기한다.
# - Numpy로 단위행렬을 생성하려면, identity or eye 명령을 사용한다.
np.identity(5)
np.eye(10)
# ### 대칭행렬
# - 만약 전치 연산을 통해서 얻은 전치 행렬과 원래의 행렬이 같으면 대칭 행렬(symmetric matrix)이라고 한다. 정방 행렬만 대칭 행렬이 될 수 있다.
# #### 연습문제 4
# 1. 영 벡터, 일벡터, 정방행렬, 대각행렬, 단위행렬, 대칭 행렬의 예를 하나씩 만들어 본다.
# 2. 위의 벡터와 행렬을 Numpy로 나타내 본다.
# 영벡터
zero = np.array([[0], [0], [0], [0]])
zero
# 일벡터
one = np.array([[1], [1], [1]])
one
# 정방행렬
X = np.array([[1,2,3],[4,5,6], [7,8,9]])
X.T
# 대각행렬
Y = np.array([[1,0,0], [0,3,0], [0,0,10]])
Y
# 단위행렬
np.array([[1,0,0], [0,1,0], [0,0,1]]), np.eye(3)
# 대칭 행렬
np.array([[1,2,3], [2,1,4], [3,4,1]])
| 2018_06_04_(1)Review_Numpy_Linear_Algebra.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#/usr/bin/python
# -*- coding: utf-8 -*-
import os
import sys
import json
import keys
import glob
import re
from PIL import Image
from io import BytesIO
from requests_oauthlib import OAuth1Session
# +
# 画像を保存するディレクトリ
script_dir_path = os.path.dirname(os.path.abspath("__file__"))
image_dir_path = os.path.normpath(os.path.join(script_dir_path, "../images"))
path = os.path.exists(image_dir_path)
if not path:
os.mkdir(image_dir_path)
since_id_path = script_dir_path + "/since_id.txt"
with open(since_id_path) as f:
since_id = f.read()
# +
#API認証情報
CK = keys.CK
CS = keys.CS
AT = keys.AT
ATS = keys.ATS
twitter = OAuth1Session(CK,CS,AT,ATS)
#API取得
search = "https://api.twitter.com/1.1/search/tweets.json?tweet_mode=extended"
# ツイートの取得設定
num = 10
keyword = '#VRChat' + ' -filter:retweets'
params = {'q' : keyword, 'count' : num, 'since_id' : since_id}
req = twitter.get(search, params = params)
# -
def generate_image_num():
image_num_max = 50
image_path_list = sorted(glob.glob(image_dir_path + '/[0-9][0-9][0-9].jpg'))
image_name_list = [os.path.basename(p) for p in sorted(image_path_list) if os.path.isfile(p)]
if len(image_name_list) == 0:
image_num = 0
else:
image_num_last = int(re.sub(r"\D", "", image_name_list[-1]))
if image_num_last < image_num_max:
image_num = image_num_last + 1
else:
# 一番古いファイルパス
image_path_oldest = sorted(image_path_list, key=lambda f: os.stat(f).st_mtime, reverse=True)[-1]
image_num = int(re.sub(r"\D", "", image_path_oldest))
return image_num
# +
if req.status_code == 200:
tweets = json.loads(req.content)
if tweets['statuses']:
since_id = tweets['statuses'][0]['id']
print(tweets['statuses'][0]['full_text'])
with open(since_id_path, mode='w') as f:
f.write(str(since_id))
for tweet in tweets['statuses']:
if 'extended_entities' in tweet:
if 'media' in tweet['extended_entities']:
for media in tweet['extended_entities']['media']:
if media['type'] == 'photo':
# 画像を保存
media_urls = media['media_url']
downloads = twitter.get(media_urls).content
image = Image.open(BytesIO(downloads))
image_resize = image.resize((1024, 1024))
image_num = generate_image_num()
image_name = '{0:03d}'.format(image_num) + '.jpg'
image_path = image_dir_path + '/' + image_name
image_resize.save(image_path)
else:
print("ERROR: %d" % req.status_code)
| scripts/otameshi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pyspark
from pyspark import SparkContext
sc = SparkContext.getOrCreate();
import findspark
findspark.init()
from pyspark.sql import SparkSession
spark = SparkSession.builder.master("local[*]").getOrCreate()
spark.conf.set("spark.sql.repl.eagerEval.enabled", True) # Property used to format output tables better
spark
# +
sc.stop()
# +
import pyspark
from pyspark.sql import SparkSession
spark = SparkSession.builder.master("local[*]").getOrCreate()
spark.conf.set("spark.sql.repl.eagerEval.enabled", True) # Property used to format output tables better
spark
from pyspark.sql import SparkSession
spark = SparkSession.builder.appName('Twitter Data Analysis').getOrCreate()
# -
import os
import re
from datetime import date, datetime
import pandas as pd
file_path_name = 'JoeBidenTweets.csv'
def open_file(file_path_name):
return pd.read_csv(file_path_name, index_col=[0])
# +
print(open_file(file_path_name).head())
# -
new_df = open_file(file_path_name)
count = new_df['tweet'].str.split().str.len()
count.index = count.index.astype(str) + ' words:'
count.sort_index(inplace=True)
words_count = []
for i, content in new_df['tweet'].items():
new_values =[]
new_values = content.split()
words_count.append(len(new_values))
print("Total number of words: ", sum(words_count), "words")
count = new_df['likes']
likes_count = []
for i, content in new_df['likes'].items():
new_values =[]
new_values = content
likes_count.append(new_values)
print("Total number of words: ", sum(likes_count), "words")
count = new_df['quotes']
quotes_count = []
for i, content in new_df['quotes'].items():
new_values =[]
new_values = content
quotes_count.append(new_values)
print("Total number of words: ", sum(quotes_count), "words")
print("Average number of words per tweet: ", round(count.mean(),2), "words")
print("Max number of words per tweet: ", count.max(), "words")
print("Min number of words per tweet: ", count.min(), "words")
new_df['tweet_length'] = new_df['tweet'].str.len()
print("Total length of a dataset: ", new_df.tweet_length.sum(), "characters")
print("Average length of a tweet: ", round(new_df.tweet_length.mean(),0), "characters")
| historical data (1).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Stop Words
# Words like "a" and "the" appear so frequently that they don't require tagging as thoroughly as nouns, verbs and modifiers. We call these *stop words*, and they can be filtered from the text to be processed. spaCy holds a built-in list of some 305 English stop words.
# Perform standard imports:
import spacy
nlp = spacy.load('en_core_web_sm')
# Print the set of spaCy's default stop words (remember that sets are unordered):
print(nlp.Defaults.stop_words)
len(nlp.Defaults.stop_words)
# ## To see if a word is a stop word
nlp.vocab['myself'].is_stop
nlp.vocab['mystery'].is_stop
# ## To add a stop word
# There may be times when you wish to add a stop word to the default set. Perhaps you decide that `'btw'` (common shorthand for "by the way") should be considered a stop word.
# +
# Add the word to the set of stop words. Use lowercase!
nlp.Defaults.stop_words.add('btw')
# Set the stop_word tag on the lexeme
nlp.vocab['btw'].is_stop = True
# -
len(nlp.Defaults.stop_words)
nlp.vocab['btw'].is_stop
# <font color=green>When adding stop words, always use lowercase. Lexemes are converted to lowercase before being added to **vocab**.</font>
# ## To remove a stop word
# Alternatively, you may decide that `'beyond'` should not be considered a stop word.
# +
# Remove the word from the set of stop words
nlp.Defaults.stop_words.remove('beyond')
# Remove the stop_word tag from the lexeme
nlp.vocab['beyond'].is_stop = False
# -
len(nlp.Defaults.stop_words)
nlp.vocab['beyond'].is_stop
# Great! Now you should be able to access spaCy's default set of stop words, and add or remove stop words as needed.
# ## Next up: Vocabulary and Matching
| 08-deep-learning/labs/03_NLP/04-Stop-Words.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Application to data classification
# + tags=["hide-input"]
from IPython.display import IFrame
IFrame(src="https://cdnapisec.kaltura.com/p/2356971/sp/235697100/embedIframeJs/uiconf_id/41416911/partner_id/2356971?iframeembed=true&playerId=kaltura_player&entry_id=1_iyo3sdca&flashvars[streamerType]=auto&flashvars[localizationCode]=en&flashvars[leadWithHTML5]=true&flashvars[sideBarContainer.plugin]=true&flashvars[sideBarContainer.position]=left&flashvars[sideBarContainer.clickToClose]=true&flashvars[chapters.plugin]=true&flashvars[chapters.layout]=vertical&flashvars[chapters.thumbnailRotator]=false&flashvars[streamSelector.plugin]=true&flashvars[EmbedPlayer.SpinnerTarget]=videoHolder&flashvars[dualScreen.plugin]=true&flashvars[hotspots.plugin]=1&flashvars[Kaltura.addCrossoriginToIframe]=true&&wid=1_5yyqw8l9" ,width='800', height='500')
# -
| _build/jupyter_execute/Module3/m3_06.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: SpringRTS
# language: ''
# name: spring_kernel
# ---
# %help
_p(5)
_p(CMD)
_p(Spring.GetAllUnits())
_p(Spring.GetUnitCommands(6512, -1)[1])
_p(CMD[29])
for i = 1, 30 do _p(i, CMD[i]) end
| issues/cmdids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from dataset import Dictionary, VQAFeatureDataset, VisualGenomeFeatureDataset
from dataset_gqa import GQAFeatureDataset
from dataset_gqa import tfidf_from_questions_gqa
from os.path import join, exists
import torch
import numpy as np
dictionary = Dictionary.load_from_file(join('data', 'glove/dictionary.pkl'))
train_dset = GQAFeatureDataset('testdev_all', dictionary, 'semantic', adaptive=False, pos_emb_dim=64,dataroot='data')
dictionary = Dictionary.load_from_file(join('data', 'glove/dictionary.pkl'))
train_dset = VQAFeatureDataset('train', dictionary, 'semantic', adaptive=False, pos_emb_dim=64,dataroot='data')
# +
features, normalized_bb, question, target, question_id, image_id, bb, spatial_adj_matrix,semantic_adj_matrix = train_dset.__getitem__(100)
#{'question_id': 142001, 'image_id': 142, 'image': 6372, 'question': 'Is this a banana toast?', 'answer': {'labels': tensor([9, 3]), 'scores': tensor([1., 1.])}, 'q_token': tensor([ 1, 2, 14, 352, 684, 19901, 19901, 19901, 19901, 19901,
# 19901, 19901, 19901, 19901])}
# -
print (features)
print (features.shape)
#(100,2048)
print (normalized_bb)
print (normalized_bb.shape)
#(2048,7,7)
print (question)
print (question.shape)
print (target)
print (target.shape)
print (question_id)
print (image_id)
print (bb)
print (bb.shape)
#(100,4)
print (spatial_adj_matrix)
print (spatial_adj_matrix.shape)
print (semantic_adj_matrix)
print (semantic_adj_matrix.shape)
dictionary = Dictionary.load_from_file(join('data', 'glove/dictionary.pkl'))
tfidf, weights = tfidf_from_questions_gqa(['test_all', 'val_all'], dictionary, 'data', 'gqa')
# +
a = np.random.randn(1, 1, 2, 3)
t1 = torch.tensor(a)
t2 = torch.tensor(a)
t3 = torch.tensor(a, device=torch.device('cpu'))
t1 = t3.to(torch.device('cpu'))
# -
device = torch.device("cpu")
print (device)
| research_experiments/test_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
import pandas_profiling
birth=pd.read_csv('https://raw.githubusercontent.com/reddyprasade/Data-Sets-For-Machine-Learnig-and-Data-Science/master/DataSets/births.csv')
# + jupyter={"outputs_hidden": true}
birth
# -
birth.head()
# + jupyter={"outputs_hidden": true}
birth.tail()
# + jupyter={"outputs_hidden": true}
birth['decade']=10*(birth['year']//10)
birth
# + jupyter={"outputs_hidden": true}
birth.pivot_table('births',index='decade',columns='gender',aggfunc='sum')
# + jupyter={"outputs_hidden": true}
birth.pivot_table('births',index='decade',columns='gender',aggfunc='sum').plot()
# + jupyter={"outputs_hidden": true}
birth.pivot_table('births',index='year',columns='gender',aggfunc='max').plot()
# -
x=np.arange(1,9)
x
# + jupyter={"outputs_hidden": true}
plt.boxplot(x)
# -
quartiles=np.percentile(x,25,interpolation='midpoint') # Take mean value ord
quartiles
quartiles=np.percentile(x,75,interpolation='linear') # Even
quartiles
birth['births'].median()
quartiles=np.percentile(birth['births'],[25,50,75])
quartiles
mu=quartiles[1]
mu
sig=0.74*(quartiles[2]-quartiles[0])
sig
birth=birth.query('(births>@mu-5*@sig) & (births<@mu+5*@sig)')
# + jupyter={"outputs_hidden": true}
birth
# -
birth['day']=birth['day'].astype(int)
# + jupyter={"outputs_hidden": true}
birth.head()
# + jupyter={"outputs_hidden": true}
birth.isnull().describe()
# + jupyter={"outputs_hidden": true}
birth.isnull().sum()
# + jupyter={"outputs_hidden": true}
birth.index=pd.to_datetime(10000*birth.year+
100*birth.month+
birth.day,format='%Y%m%d')
birth['dayofweek']=birth.index.dayofweek
birth
# -
#import matplotlib as mpl
birth.pivot_table('births',index='dayofweek',columns='decade',aggfunc='mean').plot()
plt.gca().set_xticklabels(['Mon','Tues','Wed','Thurs','Fri','Sat','Sun'])
plt.ylabel('mean birth by day')
# + jupyter={"outputs_hidden": true}
birth_by_date=birth.pivot_table('births',[birth.index.month,birth.index.day])
birth_by_date
# + jupyter={"outputs_hidden": true}
birth_by_date.index=[pd.datetime(2012,month,day) for (month,day) in birth_by_date.index]
birth_by_date.head()
# + jupyter={"outputs_hidden": true}
birth_by_date.tail()
# + jupyter={"outputs_hidden": true}
fig,ax=plt.subplots(figsize=(12,6))
birth_by_date.plot(ax=ax);
# -
| pivot_table.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# #### New to Plotly?
# Plotly's Python library is free and open source! [Get started](https://plot.ly/python/getting-started/) by downloading the client and [reading the primer](https://plot.ly/python/getting-started/).
# <br>You can set up Plotly to work in [online](https://plot.ly/python/getting-started/#initialization-for-online-plotting) or [offline](https://plot.ly/python/getting-started/#initialization-for-offline-plotting) mode, or in [jupyter notebooks](https://plot.ly/python/getting-started/#start-plotting-online).
# <br>We also have a quick-reference [cheatsheet](https://images.plot.ly/plotly-documentation/images/python_cheat_sheet.pdf) (new!) to help you get started!
# +
import plotly.plotly as py
import plotly.graph_objs as go
import pandas as pd
# -
# #### Customize Hover for Spikelines
#
# By default, Plotly's 3D plots display lines called "spikelines" while hovering over a point.
# These lines project from the hover point to each of the three axes' normal planes and
# then extend from those projection data points to the planes' wall boundaries.
# +
import plotly.plotly as py
import plotly.graph_objs as go
import numpy as np
data = [go.Scatter3d(
x=np.random.randn(50),
y=np.random.randn(50),
z=np.random.randn(50),
)]
layout = go.Layout(
scene=go.layout.Scene(
xaxis=go.layout.scene.XAxis(
spikecolor='#1fe5bd',
spikesides=False,
spikethickness=6,
),
yaxis=go.layout.scene.YAxis(
spikecolor='#1fe5bd',
spikesides=False,
spikethickness=6,
),
zaxis=go.layout.scene.ZAxis(
spikecolor='#1fe5bd',
spikethickness=6,
),
)
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='hover-spikelines')
# -
# #### Customize Hover for Surface Contours
#
# In addition to spikelines, Plotly 3D Surface plots also display surface contours on hover by default.
# These are customized by styling the [`contours`](https://plot.ly/python/reference/#surface-contours)
# attribute in the surface trace.
# +
import plotly.plotly as py
import plotly.graph_objs as go
x = [10,20,30,40]
y = [0,1,2,3]
z = [[2,2,2,3],
[1,1,1,1],
[1,1,0,0],
[0,0,0,0]]
data = [go.Surface(
opacity=0.9,
x=x,
y=y,
z=z,
contours=go.surface.Contours(
x=go.surface.contours.X(
highlight=True,
highlightcolor="#41a7b3",
),
y=go.surface.contours.Y(highlight=False),
z=go.surface.contours.Z(highlight=False),
)
)]
layout = go.Layout(
scene=go.layout.Scene(
xaxis = go.layout.scene.XAxis(showspikes=False),
yaxis = go.layout.scene.YAxis(showspikes=False),
zaxis = go.layout.scene.ZAxis(showspikes=False),
)
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='hover-surface-contours')
# -
# #### Reference
# See [https://plot.ly/python/reference/#layout-scene](https://plot.ly/python/reference/#layout-scene) and [https://plot.ly/python/reference/#surface-contours](https://plot.ly/python/reference/#surface-contours) for more information and options!
# +
from IPython.display import display, HTML
display(HTML('<link href="//fonts.googleapis.com/css?family=Open+Sans:600,400,300,200|Inconsolata|Ubuntu+Mono:400,700" rel="stylesheet" type="text/css" />'))
display(HTML('<link rel="stylesheet" type="text/css" href="http://help.plot.ly/documentation/all_static/css/ipython-notebook-custom.css">'))
# ! pip install git+https://github.com/plotly/publisher.git --upgrade
import publisher
publisher.publish(
'3d-hover-options.ipynb', 'python/3d-hover/', 'Python 3D Hover Options | Plotly',
'How to customize 3d hover options in python with Plotly.',
title= 'Python 3D Hover Options | Plotly',
name = '3D Hover Options',
has_thumbnail='true', thumbnail='thumbnail/3d-hover.png',
language='python',
display_as='3', order=19,
ipynb= '~notebook_demo/257')
# -
| _posts/python-v3/3d/3d-hover/3d-hover-options.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import os
from pathlib import Path
import urllib.request as request
import numpy as np
import matplotlib.pyplot as plt
# +
IMAGE_SIZE = (224, 224)
# -
model = tf.keras.applications.VGG16(
include_top=True, weights="imagenet", input_tensor=None, input_shape=None, classes=1000
)
model.summary()
model.save("VGG16_full_model.h5")
model.layers[1].padding
# +
data_URL = "https://download.pytorch.org/tutorial/hymenoptera_data.zip"
# create a directory
def create_dirs(dir_path):
os.makedirs(dir_path, exist_ok=True)
print(f"{dir_path} directory created")
ROOT_DATA_DIR = "data"
create_dirs(ROOT_DATA_DIR)
data_zip_file = "data.zip"
data_zip_path = os.path.join(ROOT_DATA_DIR, data_zip_file)
if not os.path.isfile(data_zip_file):
print("downloading data...")
filename, headers = request.urlretrieve(data_URL, data_zip_path)
print(f"filename: {filename} created with info \n{headers}")
else:
print(f"file is already present")
# +
# unzip data
from zipfile import ZipFile
unzip_data_dirname = "unzip_data_dir"
unzip_data_dir = os.path.join(ROOT_DATA_DIR,unzip_data_dirname)
if not os.path.exists(unzip_data_dir):
os.makedirs(unzip_data_dir,exist_ok=True)
with ZipFile(data_zip_path) as f:
f.extractall(unzip_data_dir)
else:
print(f"data already extracted")
# -
main_data_dir = Path("./data/unzip_data_dir/hymenoptera_data/train")
main_data_dir
BATCH_SIZE=32
pixels=224
IMAGE_SIZE=(pixels,pixels)
datagen_kwargs= dict(rescale=1./255,validation_split=0.20)
dataflow_kwargs= dict(target_size=IMAGE_SIZE,batch_size=BATCH_SIZE)
# +
valid_datagen = tf.keras.preprocessing.image.ImageDataGenerator(**datagen_kwargs)
valid_generator = valid_datagen.flow_from_directory(
main_data_dir, subset="validation", shuffle=False, **dataflow_kwargs)
do_data_augmentation = False
if do_data_augmentation:
train_datagen = tf.keras.preprocessing.image.ImageDataGenerator(
rotational_range=40,
horizontal_flip=True,
width_shift_range=0.2,
height_shift_range=0.2,
shear_range=0.2,
zoom_range=0.2,
**datagen_kwargs)
else:
train_datagen = valid_datagen
train_generator = train_datagen.flow_from_directory(
main_data_dir, subset="training", shuffle=True, **dataflow_kwargs
)
# +
RGB_IMAGE_SIZE = (pixels, pixels, 3)
vgg = tf.keras.applications.vgg16.VGG16(
input_shape=RGB_IMAGE_SIZE,
weights="imagenet",
include_top=False
)
# -
vgg.summary()
for layer in vgg.layers:
print(f"{layer.name:20s}:{layer.trainable}")
vgg.output
del model
# +
CLASSES = 2
x = tf.keras.layers.Flatten()(vgg.output)
prediction = tf.keras.layers.Dense(CLASSES, activation="softmax")(x)
model = tf.keras.models.Model(inputs=vgg.input, outputs = prediction)
model.summary()
# -
model.compile(
optimizer=tf.keras.optimizer.SGD(learning_rate=0.01,momentum=0.9),
loss=tf.keras.losses.CategoricalCrossentropy(),
metrics=['accuracy']
)
# +
EPOCHS = 10
histroy = model.fit(
train_generator, epochs=EPOCHS, validation_data=valid_generator
)
# -
model.save("trained_model.h5")
label_map={val:key for key, val in train_generator.class_indices.items()}
label_map
test_img = plt.imread("./data/unzip_data_dir/hymenoptera_data/val/ants/800px-Meat_eater_ant_qeen_excavating_hole.jpg")
plt.imshow(test_img)
test_img.shape
model.predict(test_img)
tf.image.resize(test_img,(224,224))
model.predict(resized_img)
input_data = tf.expand_dims(resized_img,axis=0)
pred = model.predict(input_data)
pred
argmax=tf.argmax(pred[0]).numpy()
label_map[argmax]
loaded_model = tf.keras.models.load_model("./trained_model.h5")
preprocessd_img = tf.keras.applications.vgg16.preprocess_input(resized_img)
preprocessd_img.shape
plt.imshow(preprocessd_img)
plt.imshow(test_img)
full_model = 138_357_544
our_model 14_714_688
type(full_model)
our_model/full_model
| notebook/Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''crime-analysis-report'': pipenv)'
# language: python
# name: python37464bitcrimeanalysisreportpipenv0ac31493bf424ed3a94c4babbaf0924e
# ---
# ## DBSCAN Clustering
import numpy as np
import matplotlib.pyplot as plt
import pickle
import pandas as pd
from sklearn.preprocessing import StandardScaler
# **DBSCAN Algorithm**
# <pre>
# For each object p in D
# if p is a core object and not processed then
# C = retrieve all objects density-reachable from p
# mark all objects in C as processed
# report C as a cluster
# else mark p as outlier
# end if
# End For
# </pre>
class DBSCAN(object):
UN_CLASSIFIED = 0
NOISE = -1
UN_VISITED = 0
VISITED = 1
def __init__(self, eps, min_pts):
self.eps = eps
self.minPts = min_pts
self.n_cluster = 1
def fit(self, X):
self._train(X)
def _train(self, X):
n = X.shape[0]
self.labels_ = np.array([DBSCAN.UN_CLASSIFIED] * n)
visited = DBSCAN.UN_VISITED * np.ones(n)
left_classified, = np.where(visited == DBSCAN.UN_VISITED)
while len(left_classified) != 0:
x_idx = left_classified[0]
visited[x_idx] = DBSCAN.VISITED # mark visited
start_point = X[x_idx, :]
n_neighbors, pts = self.get_neighbors(X, start_point)
if n_neighbors < self.minPts:
self.labels_[x_idx] = DBSCAN.NOISE
else:
self.labels_[x_idx] = self.n_cluster
while len(pts) != 0:
neighor_id = pts[0]
pts = np.delete(pts, np.where(pts == neighor_id))
if visited[neighor_id] == DBSCAN.UN_VISITED:
visited[neighor_id] = DBSCAN.VISITED # mark visited
n_new_neighors, new_pts = self.get_neighbors(X, X[neighor_id])
if n_new_neighors >= self.minPts:
pts = np.concatenate((pts, new_pts), axis=None)
if self.labels_[neighor_id] == DBSCAN.UN_CLASSIFIED:
self.labels_[neighor_id] = self.n_cluster
self.n_cluster += 1
left_classified, = np.where(visited == DBSCAN.UN_VISITED)
def get_neighbors(self, X, pt):
dist = np.linalg.norm(X - pt, 2, axis=1)
neighors = np.argsort(dist)
count, = np.where(dist <= self.eps)
return len(count), neighors[:len(count)]
@staticmethod
def k_nearest_neighbour(X, pt, k):
dist = np.linalg.norm(X - pt, 2, axis=1)
neighors = np.argsort(dist)
k_dist = dist[neighors[k]]
return k_dist
@classmethod
def plot_param_est_curve(cls, data, k_nn):
y = sorted([DBSCAN.k_nearest_neighbour(data, pt, k_nn) for pt in data])
# plot
plt.plot([id_ for id_ in range(len(data))], y, marker='o')
plt.xlabel('Points sorted according to k={} nearest neighour'.format(k_nn))
plt.ylabel('k={} Nearest Distance'.format(k_nn))
data = pd.read_pickle('./state_clubbed_df.pickle')
data
all_headers = data.columns
all_headers
cols_to_norm = all_headers[2:-1]
print (cols_to_norm)
data[cols_to_norm] = StandardScaler().fit_transform(data[cols_to_norm])
data
import nbimporter # library used for importing ipnb notebooks
from pca import PCA ## import pca.ipynb as module
n_features = len(cols_to_norm)
n_features
# Using PCA to determine the principal components of the features
pca = PCA(n_components=n_features)
principal_components = pca.fit_transform(data[cols_to_norm].to_numpy())
features = range(pca.n_components_)
plt.bar(features, pca.explained_variance_ratio, color='black')
plt.xlabel('PCA features')
plt.ylabel('variance %')
plt.xticks(features)
PCA_components = pd.DataFrame(principal_components)
PCA_components = pd.concat([data[all_headers[:2]], PCA_components.loc[:, [0,1]]], axis=1)
PCA_components
plt.scatter(PCA_components[0], PCA_components[1])
plt.xlabel('PCA 1')
plt.ylabel('PCA 2')
plt
DBSCAN.plot_param_est_curve(PCA_components.loc[:, [0,1]].to_numpy(), 50)
clusterer = DBSCAN(eps=2, min_pts=50)
clusterer.fit(PCA_components.loc[:, [0,1]].to_numpy())
plt.scatter(PCA_components[0], PCA_components[1], c=clusterer.labels_)
plt.xlabel('PCA 1')
plt.ylabel('PCA 2')
df_dbscan_labels = pd.concat([data[all_headers[:2]], pd.Series(clusterer.labels_)], axis=1)
df_dbscan_labels.columns = ['State', 'Year', 'Label']
df_dbscan_labels
df = pd.read_pickle('./state_clubbed_df.pickle')
concat_df = pd.concat([df, df_dbscan_labels['Label']], axis=1)
outliers = concat_df[concat_df['Label'] == -1]
outliers
outliers.to_pickle('dbscan-outliers.pickle')
| dbscan.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
matplotlib.rcParams['figure.figsize'] = [16, 8]
import pandas as pd
pd.options.display.float_format = "{:.2f}".format
import numpy as np
import warnings
from tqdm import tqdm
warnings.filterwarnings("ignore")
# +
# All variables we care about
FcolumnNames1999_2007 = ['releaseNum', 'familyID', 'composition', 'headCount', 'ageHead', 'maritalStatus', 'own',
'employmentStatus', 'liquidWealth', 'race', 'industry','incomeHead', "incomeWife",
'foodCost', 'houseCost', 'transCost', 'educationCost', 'childCost', 'healthCost', 'education',
'participation', 'investmentAmount', 'annuityIRA', 'wealthWithoutHomeEquity', "wealthWithHomeEquity"]
FcolumnNames2009_2017 = ['releaseNum', 'familyID', 'composition', 'headCount', 'ageHead', 'maritalStatus', 'own',
'employmentStatus', 'liquidWealth', 'race', 'industry' ,'incomeHead', 'incomeWife',
'participation', 'investmentAmount', 'annuityIRA', 'wealthWithoutHomeEquity', 'wealthWithHomeEquity',
'foodCost', 'houseCost', 'transCost', 'educationCost', 'childCost', 'healthCost', 'education']
FcolumnNames2019 = ['releaseNum', 'familyID', 'composition', 'headCount', 'ageHead', 'maritalStatus', 'own',
'employmentStatus', 'liquidWealth_bank', 'liquidWealth_bond', 'race', 'industry' ,'incomeHead', 'incomeWife',
'participation', 'investmentAmount', 'annuityIRA', 'wealthWithoutHomeEquity', 'wealthWithHomeEquity',
'foodCost', 'houseCost', 'transCost', 'educationCost', 'childCost', 'healthCost', 'education']
# The timeline we care about
years = [1999, 2001, 2003, 2005, 2007, 2009, 2011, 2013, 2015, 2017, 2019]
# +
def Fcompile_data_with_features(features, years = years):
df = pd.DataFrame()
# Loading the data through years
for year in years:
df_sub = pd.read_csv(str(year) + ".csv")
if year >= 1999 and year <= 2007:
df_sub.columns = FcolumnNames1999_2007
elif year >= 2009 and year <= 2017:
df_sub.columns = FcolumnNames2009_2017
else:
# In the year 2019, the liquid wealth equals to liquidWealth in cash and liquid bond
df_sub.columns = FcolumnNames2019
df_sub["liquidWealth"] = df_sub['liquidWealth_bank'] + df_sub['liquidWealth_bond']
df_sub['year'] = year
df = pd.concat([df, df_sub[['familyID','year'] + features]])
df = df.reset_index(drop = True)
return df
# The function is used to drop the values we do not like in the dataFrame,
# the input "features" and "values" are both list
def drop_values(features, values, df):
for feature in features:
for value in values:
df = df[df[feature] != value]
df = df.reset_index(drop = True)
return df
# -
# prepare the combined dataset and set up dummy variables for qualitative data
df = Fcompile_data_with_features(['composition', 'headCount', 'ageHead', 'maritalStatus', 'own',
'employmentStatus', 'liquidWealth', 'race', 'industry','incomeHead', 'incomeWife',
'foodCost', 'houseCost', 'transCost', 'educationCost', 'childCost', 'healthCost', 'education',
'participation', 'investmentAmount', 'annuityIRA', 'wealthWithoutHomeEquity', 'wealthWithHomeEquity'], years)
# data clean, drop NA/DK values
df = drop_values(["ageHead"],[999], df)
df = drop_values(["maritalStatus"],[8,9], df)
df = drop_values(["own"],[8,9], df)
df = drop_values(["employmentStatus"],[0,22,8,98, 99], df)
df = drop_values(["liquidWealth"],[999999998,999999999,-400], df)
df = drop_values(["race"],[0,8,9], df)
df = drop_values(["industry"],[999,9999,0], df)
df = drop_values(["education"],[99,0], df)
# calculate the aggregate variables
df["totalExpense"] = df[['foodCost', 'houseCost', 'transCost',
'educationCost', 'childCost', 'healthCost']].sum(axis = 1)
df["laborIncome"] = df["incomeHead"] + df["incomeWife"]
df["costPerPerson"] = df["totalExpense"]/df["headCount"]
df["HomeEquity"] = df["wealthWithHomeEquity"] - df["wealthWithoutHomeEquity"]
# +
maritalStatus = ["Married", "neverMarried", "Widowed", "Divorced", "Separated"]
employmentStatus = ["Working", "temporalLeave", "unemployed", "retired", "disabled", "keepHouse", "student", "other"]
race = ["White", "Black","AmericanIndian","Asian","Latino","otherBW","otherRace"]
# Education
# < 8th grade: middle school
# >= 8 and < 12: high scho0l
# >=12 and < 15: college
# >= 15 post graduate
education = ["middleSchool", "highSchool", "college", "postGraduate"]
# Industry
# < 400 manufacturing
# >= 400 and < 500 publicUtility
# >= 500 and < 680 retail
# >= 680 and < 720 finance
# >= 720 and < 900 service
# >= 900 otherIndustry
industry = ["finance", "noneFinance"]
ownership = ["owner", "renter"]
data = []
for i in tqdm(range(len(df))):
dataCollect = []
# marital status
dataCollect.append(maritalStatus[int(df.iloc[i]["maritalStatus"]-1)])
# employment
dataCollect.append(employmentStatus[int(df.iloc[i]["employmentStatus"]-1)])
# race
dataCollect.append(race[int(df.iloc[i]["race"] - 1)])
# Education variable
if df.iloc[i]["education"] < 8:
dataCollect.append(education[0])
elif df.iloc[i]["education"] >= 8 and df.iloc[i]["education"] < 12:
dataCollect.append(education[1])
elif df.iloc[i]["education"] >= 12 and df.iloc[i]["education"] < 15:
dataCollect.append(education[2])
else:
dataCollect.append(education[3])
# industry variable
if df.iloc[i]["year"] in [1999, 2001]:
if df.iloc[i]["industry"] >= 707 and df.iloc[i]["industry"] <= 718:
dataCollect.append(industry[0])
else:
dataCollect.append(industry[1])
elif df.iloc[i]["year"] in [2003,2005,2007,2009,2011,2013,2015,2015]:
if df.iloc[i]["industry"] >= 687 and df.iloc[i]["industry"] <= 699:
dataCollect.append(industry[0])
else:
dataCollect.append(industry[1])
else:
if df.iloc[i]["industry"] >= 6870 and df.iloc[i]["industry"] <= 6990:
dataCollect.append(industry[0])
else:
dataCollect.append(industry[1])
# ownership status
if df.iloc[i]["own"] == 1:
dataCollect.append(ownership[0])
else:
dataCollect.append(ownership[1])
data.append(dataCollect)
# Categorical dataFrame
df_cat = pd.DataFrame(data, columns = ["maritalStatus", "employmentStatus", "race", "education", "industry", "ownership"])
Fdf = pd.concat([df[["familyID", "year",'composition', 'headCount', 'ageHead', 'liquidWealth', 'laborIncome',
"costPerPerson","totalExpense", 'participation', 'investmentAmount', 'annuityIRA',
'wealthWithoutHomeEquity', "wealthWithHomeEquity", "HomeEquity"]],
df_cat[["maritalStatus", "employmentStatus", "education","race", "industry", "ownership"]]], axis=1)
# -
# Adjust for inflation, all values are in thousand dollor
years = [1999, 2001, 2003, 2005, 2007, 2009, 2011, 2013, 2015, 2017, 2019]
values_at2020 = np.array([1.55, 1.46, 1.40, 1.32, 1.24, 1.20, 1.15, 1.11, 1.09, 1.05, 1.01])
values_at2005 = values_at2020/1.32
quantVariables = ['annuityIRA', 'investmentAmount', 'liquidWealth', 'laborIncome', 'costPerPerson',
'totalExpense', 'wealthWithoutHomeEquity', 'wealthWithHomeEquity', "HomeEquity"]
for i in tqdm(range(len(Fdf))):
for variable in quantVariables:
Fdf.loc[i, variable] = round(Fdf.loc[i, variable] * values_at2005[years.index(Fdf.loc[i,"year"])] / 1000, 2)
Fdf.head()
# drop the outliers
for var in quantVariables:
Fdf = Fdf[Fdf[var] < Fdf[var].quantile(0.999)]
Fdf = Fdf[(Fdf["ageHead"] >= 20) & (Fdf["ageHead"] <= 80)]
Fdf.head()
Fdf.shape
# ### Group population into 4 type of agents
lowSkill = ["middleSchool", "highSchool"]
highSkill = ["college", "postGraduate"]
highFinance = Fdf[(Fdf["education"].isin(highSkill)) & (Fdf["industry"] == "finance")]
lowFinance = Fdf[(Fdf["education"].isin(lowSkill)) & (Fdf["industry"] == "finance")]
highNoneFinance = Fdf[(Fdf["education"].isin(highSkill)) & (Fdf["industry"] == "noneFinance")]
lowNoneFinance = Fdf[(Fdf["education"].isin(lowSkill)) & (Fdf["industry"] == "noneFinance")]
Fdf["skillLevel"] = "High"
Fdf.loc[Fdf["education"].isin(lowSkill), "skillLevel"] = "Low"
Fdf["financeExperience"] = "No"
Fdf.loc[Fdf["industry"] == "finance", "financeExperience"] = "Yes"
Fdf["ageGroup"] = "20"
Fdf["decadeGroup"] = "90's"
for i in range(2,10, 2):
Fdf.loc[Fdf["ageHead"] > i*10, "ageGroup"] = str(i*10)
for year in range(1990,2020,10):
Fdf.loc[Fdf["year"] > year, "decadeGroup"] = str(year) + "s"
Fdf.loc[(Fdf["employmentStatus"] != "Working")&(Fdf["employmentStatus"] != "retired"), "employmentStatus"] = "unemployed"
Fdf.loc[Fdf["employmentStatus"]=="Working", "employmentStatus"] = "employed"
Fdf.loc[Fdf["ageGroup"]== "20", "ageGroup"] = "20-40"
Fdf.loc[Fdf["ageGroup"]== "40", "ageGroup"] = "40-60"
Fdf.loc[Fdf["ageGroup"]== "60", "ageGroup"] = "60-80"
Fdf.head()
Fdf.tail()
Fdf[(Fdf["ownership"] == "renter") & (Fdf["HomeEquity"] != 0)]
Fdf["stockInvestmentRatio"] = Fdf.investmentAmount/Fdf.wealthWithoutHomeEquity
Fdf.loc[-((Fdf["stockInvestmentRatio"] >= 0)&(Fdf["stockInvestmentRatio"] <= 1)), "stockInvestmentRatio"] = 0
# +
# plt.title("Mean Total Wealth Level Empirical Value vs Model Value")
# plt.plot(range(20,81), np.load("modelPoorHigh.npy"), label = "HighSkilled_model")
# plt.plot(range(20,81), np.load("modelRichLow.npy"), label = "LowSkilled_model")
# plt.plot(range(20,81), Fdf[Fdf["skillLevel"] == "High"].groupby("ageHead")["wealthWithHomeEquity"].mean().values, label = "HighSkilled_empirical")
# plt.plot(range(20,81), Fdf[Fdf["skillLevel"] == "Low"].groupby("ageHead")["wealthWithHomeEquity"].mean().values, label = "LowSkilled_empirical")
# plt.legend()
# savefig('EmpiricalVsModel.jpg')
# -
Fdf.to_csv("familyData.csv")
plt.title("Participation ratio over different edcucation backgrounds")
Fdf.loc[Fdf["education"]=="middleSchool", "education"] = "highSchool"
Fdf.groupby("education").participation.mean().reindex(["highSchool", "college", "postGraduate"]).plot.bar(color = "blue",rot=0)
savefig('educationParticipation.jpg')
Fdf.loc[Fdf["employmentStatus"]=="Working", "employmentStatus"] = "employed"
plt.title("Participation ratio over different employment status")
Fdf.groupby("employmentStatus").participation.mean().reindex(["unemployed", "employed", "retired"]).plot.bar(color = "blue",rot=0)
savefig('employmentParticipation.jpg')
plt.title("Participation ratio over different employment industries")
Fdf.groupby("industry").participation.mean().plot.bar(color = "blue",rot=0)
savefig('industryParticipation.jpg')
plt.title("Participation ratio over different age groups")
Fdf.groupby("ageGroup").participation.mean().plot.bar(color = "blue",rot=0)
savefig('ageParticipation.jpg')
plt.title("Participation ratio over housing status")
Fdf.groupby("ownership").participation.mean().plot.bar(color = "blue",rot=0)
savefig('ownership.jpg')
plt.title("Participation ratio over different years")
Fdf.groupby("year").participation.mean().plot.bar(color = "blue",rot=0)
savefig('year.jpg')
Fdf[(Fdf.education == 'highSchool')|(Fdf.education == 'college')].groupby("ageHead")['laborIncome'].mean().plot(label = "highSchool")
Fdf[Fdf.education == 'postGraduate'].groupby("ageHead")['laborIncome'].mean().plot(label = "postGraduate")
# +
from scipy.signal import savgol_filter
lowIncome = Fdf[(Fdf.education == 'highSchool')|(Fdf.education == 'college')].groupby("ageHead")['laborIncome'].mean().values
highIncome = Fdf[Fdf.education == 'postGraduate'].groupby("ageHead")['laborIncome'].mean().values
lowIncome_median = Fdf[(Fdf.education == 'highSchool')|(Fdf.education == 'college')].groupby("ageHead")['laborIncome'].median().values
highIncome_median = Fdf[Fdf.education == 'postGraduate'].groupby("ageHead")['laborIncome'].median().values
highIncomeSmooth = savgol_filter(highIncome,window_length=9,polyorder=1)
lowIncomeSmooth = savgol_filter(lowIncome,window_length=9,polyorder=1)
ages = list(range(20,81))
# -
plt.plot(ages, highIncome, label = "high income")
plt.plot(ages, lowIncome, label = "low income")
plt.plot(ages, highIncome_median, label = "high income_median")
plt.plot(ages, lowIncome_median, label = "low income_median")
plt.plot(ages, highIncomeSmooth, label = "high income smoothed")
plt.plot(ages, lowIncomeSmooth, label = "low income smoothed")
plt.legend()
highIncomeSmooth[46:] = np.mean(highIncomeSmooth[46:])
lowIncomeSmooth[46:] = np.mean(lowIncomeSmooth[46:])
plt.plot(ages, highIncomeSmooth, label = "high income smoothed")
plt.plot(ages, lowIncomeSmooth, label = "low income smoothed")
plt.legend()
highIncomeSmooth
lowIncomeSmooth
np.save("../shutDownRetirement/constant/highIncomeDetermined", highIncomeSmooth)
np.save("../shutDownRetirement/constant/lowIncomeDetermined", lowIncomeSmooth)
c_bar = Fdf.groupby("ageHead")['totalExpense'].quantile(0.25).values
c_bar_smooth = savgol_filter(c_bar,window_length=11,polyorder=1)
plt.plot(ages, c_bar, label = "lower 25th percentile of the consumption")
plt.plot(ages, c_bar_smooth, label = "lower 25th percentile of the consumption smoothed")
plt.legend()
np.save("../constant/c_bar", c_bar_smooth)
Fdf[Fdf.skillLevel == 'High'].groupby("ageHead")['wealthWithHomeEquity'].median().plot(label = "HighSkill")
Fdf[Fdf.skillLevel == 'Low'].groupby("ageHead")['wealthWithHomeEquity'].median().plot(label = "LowSkill")
Fdf[Fdf.skillLevel == 'High'].groupby("ageHead")['wealthWithHomeEquity'].mean().plot(label = "HighSkill_mean")
Fdf[Fdf.skillLevel == 'Low'].groupby("ageHead")['wealthWithHomeEquity'].mean().plot(label = "LowSkill_mean")
plt.legend()
# + tags=[]
for year in years:
plt.figure()
plt.title("age distribution in the year: " + str(year))
plt.hist(Fdf[Fdf["year"] == year].ageHead, bins = 100)
# -
for year in years:
highSkill = plt.scatter(year, Fdf[(Fdf["year"] == year)&(Fdf["skillLevel"] == "High")].shape[0], color = 'r')
lowSkill = plt.scatter(year, Fdf[(Fdf["year"] == year)&(Fdf["skillLevel"] == "Low")].shape[0], color = 'g')
plt.legend((highSkill,lowSkill), ("highSkill", "lowSkill"))
highSkillRatios = []
lowSkillRatios = []
for year in years:
highRatio = Fdf[(Fdf["year"] == year)&(Fdf["skillLevel"] == "High")].shape[0]/Fdf[(Fdf["year"] == year)].shape[0]
lowRatio = Fdf[(Fdf["year"] == year)&(Fdf["skillLevel"] == "Low")].shape[0]/Fdf[(Fdf["year"] == year)].shape[0]
highSkillRatios.append(highRatio)
lowSkillRatios.append(lowRatio)
highSkill = plt.scatter(year, highRatio, color = 'r')
lowSkill = plt.scatter(year, lowRatio, color = 'g')
plt.legend((highSkill,lowSkill), ("highSkill", "lowSkill"))
highSkillRatios
lowSkillRatios
financeRatios = []
noneFinanceRatios= []
for year in years:
fRatio = Fdf[(Fdf["year"] == year)&(Fdf["financeExperience"] == "Yes")].shape[0]/Fdf[(Fdf["year"] == year)].shape[0]
nfRatio = Fdf[(Fdf["year"] == year)&(Fdf["financeExperience"] == "No")].shape[0]/Fdf[(Fdf["year"] == year)].shape[0]
financeRatios.append(fRatio)
noneFinanceRatios.append(nfRatio)
finance = plt.scatter(year, fRatio, color = 'r')
noneFinance = plt.scatter(year, nfRatio, color = 'g')
plt.legend((finance,noneFinance), ("finance", "noneFinance"))
for year in years:
finance = plt.scatter(year, Fdf[(Fdf["year"] == year)&(Fdf["financeExperience"] == "Yes")].shape[0], color = 'r')
nonFinance = plt.scatter(year, Fdf[(Fdf["year"] == year)&(Fdf["financeExperience"] == "No")].shape[0], color = 'g')
plt.legend((finance,noneFinance), ("finance", "noneFinance"))
financeRatios
noneFinanceRatios
| 20211218/family/Household.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Annotated Data
#
# The AnnData library is the primary protocol that is used to store imaging data in an efficient, multi-functional format. It is created using the `anndata` sub-module and can be accessed using `trialobj.data`. By default, `trialobj.data` is a data array generated from Suite2p processed data.
# For all guidance on AnnData objects, visit: https://anndata.readthedocs.io/en/latest/index.html.
#
# The AnnData object is built around the raw Flu matrix of each `trialobj` . In keeping with AnnData conventions, the data structure is organized in *n* observations (obs) x *m* variables (var), where observations are suite2p ROIs and variables are imaging frame timepoints.
#
# + pycharm={"name": "#%%\n"}
display.Image("/home/pshah/Documents/code/packerlabimaging/files/packerlabimaging-anndata-integration-01.jpg")
# + pycharm={"name": "#%%\n"}
import packerlabimaging as pli
# + pycharm={"name": "#%%\n"}
expobj: pli.Experiment = pli.import_obj(pkl_path='/home/pshah/Documents/code/packerlabimaging/tests/RL109_analysis.pkl')
trialobj = expobj.load_trial(trialID=expobj.trialIDs[0])
# + pycharm={"name": "#%%\n"}
trialobj.data # this is the anndata object for this trial
# -
# ### storage of Flu data
#
# The raw data is stored in `.X`
# + pycharm={"name": "#%%\n"}
print(trialobj.data.X)
print('shape: ', trialobj.data.X.shape)
# -
# Processed data is added to `trialobj.data` as a unique `layers` key.
# + pycharm={"name": "#%%\n"}
trialobj.data.layers
# + pycharm={"name": "#%%\n"}
print(trialobj.data.layers['dFF'])
print('shape: ', trialobj.data.layers['dFF'].shape)
# -
# The rest of the AnnData data object is built according to the dimensions of the original Flu data input.
# ### observations (Suite2p ROIs metadata and associated processing info)
#
# For instance, the metadata for each suite2p ROI stored in Suite2p’s stat.npy output is added to `trialobject.data` under `obs` and `obsm` (1D and >1-D observations annotations, respectively).
# + pycharm={"name": "#%%\n"}
trialobj.data.obs
# + pycharm={"name": "#%%\n"}
trialobj.data.obsm
# -
# The `.obsm` includes the ypix and xpix outputs for each suite2p ROI which represent the pixel locations of the ROI mask.
# + pycharm={"name": "#%%\n"}
print('ypix:', trialobj.data.obsm['ypix'][:5], '\n\nxpix: \t', trialobj.data.obsm['xpix'][:5])
# -
# ### variables (temporal synchronization of paq channels and imaging)
#
# And the temporal synchronization data of the experiment collected in .paq output is added to the variables annotations under `var`. These variables are timed to the imaging frame clock timings. The total # of variables is the number of imaging frames in the original Flu data input.
# + pycharm={"name": "#%%\n"}
trialobj.data.var
# -
# ### Creating or Modifying AnnData arrays of trialobj
#
# There are a number of helper functions to create anndata arrays or modify existing anndata arrays.
# + pycharm={"name": "#%%\n"}
# creating new anndata object. This is identical to the base AnnData library.
# the example below is from the Getting Started Tutorial for AnnData:
# any given anndata object is created from constituent data arrays.
# 1) Primary data matrix
import numpy as np
import pandas as pd
n_rois, n_frames = 10, 10000
X = np.random.random((n_rois, n_frames)) # create random data matrix
df = pd.DataFrame(X, columns=range(n_frames), index=np.arange(n_rois, dtype=int).astype(str))
df # show the dataframe
# + pycharm={"name": "#%%\n"}
#2) Observations matrix
obs_meta = pd.DataFrame({
'cell_type': np.random.choice(['exc', 'int'], n_rois),
},
index=np.arange(n_rois, dtype=int).astype(str), # these are the same IDs of observations as above!
)
obs_meta
# + pycharm={"name": "#%%\n"}
#3) Variables matrix
var_meta = pd.DataFrame({
'exp_group': np.random.choice(['A','B', 'C'], n_frames),
},
index=np.arange(n_frames, dtype=int).astype(str), # these are the same IDs of observations as above!
)
var_meta
# + pycharm={"name": "#%%\n"}
#4) Creating a new anndata attribute for the trialobj
import packerlabimaging.processing.anndata as ad # from the processing module, import anndata submodule
trialobj.new_anndata = ad.AnnotatedData(X=df,obs=obs_meta, var=var_meta)
print(trialobj.new_anndata)
# + pycharm={"name": "#%%\n"}
# adding an 'obs' to existing anndata object
new_obs = pd.DataFrame({
'cell_loc_x': np.random.random_integers(0, 512, n_rois),
'cell_loc_y': np.random.random_integers(0, 512, n_rois),
},
index=np.arange(n_rois, dtype=int).astype(str), # these are the same IDs of observations as above!
)
cell_loc_x = np.random.random_integers(0, 512, n_rois)
cell_loc_y = np.random.random_integers(0, 512, n_rois)
trialobj.new_anndata.add_obs(obs_name='cell_loc_x', values=cell_loc_x)
trialobj.new_anndata.add_obs(obs_name='cell_loc_y', values=cell_loc_y)
print(trialobj.new_anndata)
# + pycharm={"name": "#%%\n"}
# deleting an 'obs' to existing anndata object
# uses the pop method
trialobj.new_anndata.del_obs('cell_type')
print(trialobj.new_anndata)
# -
# *Note: adding and deleting an 'var' to existing anndata object can be done in the exact same manner as demonstrated above for 'obs' using .add_var() and .del_var() methods on an anndata object.*
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Plotting quickly from anndata
# + [markdown] pycharm={"name": "#%% md\n"}
# To quickly plot from data contained within an `anndata` object, we convert the `anndata` object to a long-form `pandas` dataframe.
#
# Using this format of the data, we can quickly access the excellent `seaborn` library for plotting.
# + pycharm={"name": "#%%\n"}
trialobj.data.df = trialobj.data.convert_to_df()
# + pycharm={"name": "#%%\n"}
import seaborn as sns
import matplotlib.pyplot as plt
sns.scatterplot(data=df, x="group", y="test_data")
plt.show()
| docs/source/Tutorials/Tutorial-7-anndata-module.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This notebook creates nuclio serverless function for train data preprocessing.
# The datapreprocessing functions are defined in widsutil.py
# The encoded data is saved as a dataset artifact
# +
import mlrun
from os import path
project_name = "widsdb2"
project_dir = "conf"
widsdb2_proj = mlrun.new_project(project_name,
context=project_dir,
init_git=True,
user_project=False)
#wids_prj, artifact_path = mlrun.set_environment('http://mlrun-api:8080',
# project=project_name, user_project=False)
#widsdb2_proj = mlrun.projects.load_project(project_dir, clone=True)
# +
# nuclio: start-code
# +
import os
import sys
sys.path.append('/v3io/projects/widsdb2/util')
import json
import pandas as pd
import numpy as np
from collections import defaultdict
import widsutil as util
from cloudpickle import dumps, dump, load
from mlrun.execution import MLClientCtx
from mlrun.datastore import DataItem
def trdata_prep(
context:MLClientCtx,
src: DataItem,
file_ext: str = "csv",
train_enc: str = "train_enc"
):
"""process a raw icu data file
* `encoded` is input for a cross validation and training function
widsutil module has various transform functions
steps (not necessarily in correct order, some parallel)
* column name maps
* deal with nans and other types of missings/junk
* label encode binary and ordinal category columns
* create category ranges from numerical columns
And finally,
* test
TODO:
* parallelize where possible
* more abstraction (more parameters, chain sklearn transformers)
* convert to marketplace function
:param context: the function execution context
:param src: an artifact or file path
:param file_ext: file type for artifacts
:param train_enc: key of encoded data table in artifact store
"""
df = src.as_df()
datapreprocess =util.DataPreprocess(df)
df = datapreprocess.preprocess()
#df = df.fillna(0)
df.fillna(df.mean(), inplace=True)
#np.where(df.values >= np.finfo(np.float32).max)
context.log_dataset(train_enc, df=df, format=file_ext, index=False)
# +
# nuclio: end-code
# -
train_data_prep_func = mlrun.code_to_function(name='prep',
handler='trdata_prep',
kind='job',
image='mlrun/ml-models')
train_data_prep_func.save()
widsdb2_proj.set_function(train_data_prep_func)
widsdb2_proj.save()
# + jupyter={"outputs_hidden": true}
#widsdb2_proj.functions
# +
import mlrun
import pandas as pd
from mlrun.run import get_dataitem
train_df = pd.read_csv('TrainingWiDS2021.csv', index_col=[0])
print(train_df.shape)
widsdb2_proj.log_dataset(key='raw_train_data', df=train_df, index=False, format='csv')
# +
from mlrun.platforms import auto_mount
import sys
sys.path.append('/v3io/projects/widsdb2/util')
#Run the nuclio function on cluster
train_data_prep_func = train_data_prep_func.apply(auto_mount())
train_data_prep_run = train_data_prep_func.run(
inputs={"src" :'store://raw_train_data', 'train-enc': 'train_enc' })
# -
#Run the nuclio function locally
train_data_prep_run = train_data_prep_func.run(
inputs={"src" :'store://raw_train_data', 'train-enc': 'train_enc' }, local=True)
# local=True, artifact_path=artifact_path)
#save the project with function added
widsdb2_proj.save()
#Get the preprocessed dataframe which is output of nuclio function run above
dfa = mlrun.get_dataitem(train_data_prep_run.outputs['train_enc']).as_df()
dfa
train_data_prep_run.outputs['train_enc']
data = f'store://widsdb2/prep-trdata_prep_train_enc'
dataset = mlrun.run.get_dataitem(data).as_df()
dataset.head()
#dataset.isnull().any() == True
dataset.columns[dataset.isnull().any()]
# +
from mlrun.mlutils.data import get_sample
raw, labels, header = get_sample(dataset, sample=-1, label='diabetes_mellitus')
len(raw)
| wids-preptrain.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computational Astrophysics
# ## 03. Fundamentals of Python. Classes and Objects
#
# ---
# ## <NAME>
#
# Observatorio Astronómico Nacional\
# Facultad de Ciencias\
# Universidad Nacional de Colombia
#
# ---
# ### About this notebook
#
# In this notebook we present some of the fundamentals of classes and objects using `python`.
#
# ---
# ### Defining a Class
# #### Attributes
# To begin, we define a class called 'Planet' with three *attributes*. We use the class initialization definition to introduce these attributes.
class Planet(object):
def __init__ (self, planet_name, mass, orbit_period):
self.planet_name = planet_name
self.mass = mass # in units of Earth mass
self.orbit_period = orbit_period # in Earth years
# Now we can create an object using this class.
mars = Planet('Mars', 0.107, 1.88)
# The attributes of this object are easily obtained,
mars.mass
mars.orbit_period
# #### Methods
# Now we will define a class with both attributes and methods.
class Planet(object):
def __init__ (self, planet_name, mass, orbit_period):
self.planet_name = planet_name
self.mass = mass # in units of Earth mass
self.orbit_period = orbit_period # in Earth years
def semimajor_axis(self):
'''
------------------------------------------
semimajor_axis()
------------------------------------------
Returns the value of the semimajor axis of
the planet in AU, calculated using
Kepler's third law.
------------------------------------------
'''
return self.orbit_period**(2./3.)
# We define a planet using this class.
mars = Planet('Mars', 0.107, 1.88)
# The method is obtained by
mars.semimajor_axis()
# For the Earth, we have
earth = Planet('Earth', 1., 1.)
earth.semimajor_axis()
# Information of the methods of a class is obtained as usual:
# +
# Planet.semimajor_axis?
# -
print(Planet.semimajor_axis.__doc__)
# ---
# ### Defining a SubClass
# Once the class is defined, it can be used to define a subclass:
# +
class Planet(object):
def __init__ (self, planet_name, mass, orbit_period):
self.planet_name = planet_name
self.mass = mass # in units of Earth mass
self.orbit_period = orbit_period # in Earth years
def semimajor_axis(self):
'''
------------------------------------------
semimajor_axis()
------------------------------------------
Returns the value of the semimajor axis of
the planet in AU, calculated using
Kepler's third law.
------------------------------------------
'''
return self.orbit_period**(2./3.)
class Dwarf(Planet):
def description(self):
'''
------------------------------------------
description()
------------------------------------------
Returns a string with the information of
the mass of the dwarf planet.
------------------------------------------
'''
descrip = self.planet_name + ' is a dwarf planet with a mass of ' \
+ str(self.mass) + ' Earth masses.'
return descrip
# -
# Define Pluto as a dwarf planet:
pluto = Dwarf('Pluto', 0.00218, 248.00)
# We can access all the attributes and methods of the 'Planet' class as well as those of the 'Dwarf' class.
pluto.mass
pluto.semimajor_axis()
pluto.description()
| 01._Fundamentals/presentation/Fundamentals03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.6 64-bit (''venv'': venv)'
# name: python_defaultSpec_1598173300831
# ---
# +
import math
import arviz as az
import matplotlib.pyplot as plt
import pandas as pd
from IPython.display import set_matplotlib_formats
import jax.numpy as jnp
from jax import lax, random
from jax.scipy.special import expit, logit
import numpyro
import numpyro.distributions as dist
import numpyro.optim as optim
from numpyro.diagnostics import print_summary
from numpyro.distributions.transforms import OrderedTransform
from numpyro.infer import ELBO, MCMC, NUTS, SVI, Predictive, init_to_value
from numpyro.infer.autoguide import AutoLaplaceApproximation
# %config InlineBackend.figure_formats = ["svg"]
az.style.use("arviz-darkgrid")
numpyro.set_host_device_count(4)
# +
ratings_counts = jnp.array([12, 36, 7, 41])
pr_r = ratings_counts / ratings_counts.sum()
cpr_r = jnp.cumsum(pr_r, -1)
cpr_r
# -
lco_r = logit(cpr_r)
lco_r
plt.plot(range(1, 5), cpr_r, "--o")
plt.gca().set(xlabel="rank", ylabel="cumulative proportion", ylim=(-0.1, 1.1))
plt.show()
# + tags=[]
hurricanes = pd.read_csv("../data/Hurricanes.csv", sep=";")
d = hurricanes
print(d.shape)
display(d.sample(3))
d.describe()
# -
d.category.value_counts().sort_index()
#
# ## Hurricane fatalities and gender of names
#
# ### Description
#
# Data used in Jung et al 2014 analysis of effect of gender of name on hurricane fatalities. Note that hurricanes Katrina (2005) and Audrey (1957) were removed from the data.
#
# ### Format
#
# name : Given name of hurricane
# year : Year of hurricane
# deaths : number of deaths
# category : Severity code for storm
# min_pressure : Minimum pressure, a measure of storm strength; low is stronger
# damage_norm : Normalized estimate of damage in dollars
# female : Indicator variable for female name
# femininity : 1-11 scale from totally masculine (1) to totally feminine (11) for name. Average of 9 scores from 9 raters.
#
import seaborn as sns
sns.pairplot(d[['femininity', 'category', 'min_pressure', 'damage_norm', 'deaths']])
# + tags=[]
def model1(deaths=None, femininity=None):
a = numpyro.sample("a", dist.Normal(0, .3))
bF = numpyro.sample("bF", dist.Normal(0.2, .25))
lambda_ = numpyro.deterministic("lambda", jnp.exp(a + bF * femininity))
numpyro.sample("deaths", dist.Poisson(lambda_), obs=deaths)
# +
x = jnp.linspace(d.femininity.min(), d.femininity.max(), 100)
lmu = Predictive(model1, num_samples=100, return_sites=["lambda"]).get_samples(
random.PRNGKey(53),
femininity=x,
)["lambda"]
plt.axhline(y=d.deaths.min(), c="k", ls="--")
plt.axhline(y=d.deaths.max(), c="k", ls="--")#, lw=0.5)
plt.title("prior pred")
plt.xlabel('femininity')
for i in range(40):
plt.plot(x, lmu[i], "k", alpha=0.3)
# + tags=[]
mcmc = MCMC(NUTS(model1), 500, 500)
mcmc.run(random.PRNGKey(0), femininity=d.femininity.values, deaths=d.deaths.values)
mcmc.print_summary()
post1 = mcmc.get_samples()
# +
x = jnp.linspace(d.femininity.min(), d.femininity.max(), 100)
pred = Predictive(model1, post1, return_sites=["lambda"]).get_samples(random.PRNGKey(33), femininity=x)
lmu = jnp.mean(pred['lambda'], 0)
lci = jnp.percentile(pred['lambda'], q=(5.5, 94.5), axis=0)
plt.scatter(d.femininity.values, d.deaths.values)
plt.plot(x, lmu, 'k')
plt.fill_between(x, lci[0], lci[1], color='k', alpha=0.2)
# -
def model2(deaths=None):
a = numpyro.sample("a", dist.Normal(0, .3))
lambda_ = numpyro.deterministic("lambda", jnp.exp(a))
numpyro.sample("deaths", dist.Poisson(lambda_), obs=deaths)
# + tags=[]
mcmc = MCMC(NUTS(model2), 500, 500)
mcmc.run(random.PRNGKey(0), deaths=d.deaths.values)
mcmc.print_summary()
post2 = mcmc.get_samples()
# -
def model3(deaths=None, femininity=None):
a = numpyro.sample("a", dist.Normal(0, 5))
bF = numpyro.sample("bF", dist.Normal(0.2, .25))
phi = numpyro.sample("phi", dist.Exponential(1))
lambda_ = numpyro.deterministic("lambda", jnp.exp(a + bF * femininity))
numpyro.sample("deaths", dist.GammaPoisson(lambda_ / phi, 1 / phi), obs=deaths)
# + tags=[]
mcmc = MCMC(NUTS(model3), 500, 500)
mcmc.run(random.PRNGKey(0), femininity=d.femininity.values, deaths=d.deaths.values)
mcmc.print_summary()
post3 = mcmc.get_samples()
# +
x = jnp.linspace(d.femininity.min(), d.femininity.max(), 100)
pred = Predictive(model3, post3, return_sites=["lambda"]).get_samples(random.PRNGKey(33), femininity=x)
lmu = jnp.mean(pred['lambda'], 0)
lci = jnp.percentile(pred['lambda'], q=(5.5, 94.5), axis=0)
plt.scatter(d.femininity.values, d.deaths.values)
plt.plot(x, lmu, 'k')
plt.fill_between(x, lci[0], lci[1], color='k', alpha=0.2)
# -
deaths_pred = Predictive(model3, post3)(random.PRNGKey(123), femininity=d.femininity.values)["deaths"]
plt.scatter(d.femininity.values, d.deaths.values)
plt.errorbar(
d.femininity.values,
jnp.mean(deaths_pred, 0),
jnp.std(deaths_pred, 0) / 2,
fmt="o",
c="k",
mfc="none",
ms=7,
elinewidth=1,
)
plt.plot(d.femininity.values, jnp.percentile(deaths_pred, 5.5, 0), "k+")
plt.plot(d.femininity.values, jnp.percentile(deaths_pred, 94.5, 0), "k+")
| practice/12.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predicting Star Temperature
# Using the Open Exoplanet Catalogue database: https://github.com/OpenExoplanetCatalogue/open_exoplanet_catalogue/
#
# ## Data License
# Copyright (C) 2012 <NAME>
#
# Permission is hereby granted, free of charge, to any person obtaining a copy of this database and associated scripts (the "Database"), to deal in the Database without restriction, including without limitation the rights to use, copy, modify, merge, publish, distribute, sublicense, and/or sell copies of the Database, and to permit persons to whom the Database is furnished to do so, subject to the following conditions:
#
# The above copyright notice and this permission notice shall be included in all copies or substantial portions of the Database. A reference to the Database shall be included in all scientific publications that make use of the Database.
#
# THE DATABASE IS PROVIDED "AS IS", WITHOUT WARRANTY OF ANY KIND, EXPRESS OR IMPLIED, INCLUDING BUT NOT LIMITED TO THE WARRANTIES OF MERCHANTABILITY, FITNESS FOR A PARTICULAR PURPOSE AND NONINFRINGEMENT. IN NO EVENT SHALL THE AUTHORS OR COPYRIGHT HOLDERS BE LIABLE FOR ANY CLAIM, DAMAGES OR OTHER LIABILITY, WHETHER IN AN ACTION OF CONTRACT, TORT OR OTHERWISE, ARISING FROM, OUT OF OR IN CONNECTION WITH THE DATABASE OR THE USE OR OTHER DEALINGS IN THE DATABASE.
#
# ## Setup
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import seaborn as sns
stars = pd.read_csv('../../ch_09/data/stars.csv')
stars.head()
# -
# ## EDA
stars.info()
stars.describe()
sns.heatmap(stars.corr(), vmin=-1, vmax=1, center=0, annot=True, fmt='.1f')
# ## Build the model and evaluate it
# +
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
data = stars[['metallicity', 'temperature', 'magJ', 'radius', 'magB', 'magV', 'magK', 'mass', 'planets']].dropna()
y = data.pop('temperature')
X = data
X_train, X_test, y_train, y_test = train_test_split(
X, y, test_size=0.25, random_state=0
)
lm = LinearRegression().fit(X_train, y_train)
lm.score(X_test, y_test) # R-squared
# -
from sklearn.metrics import mean_squared_error
np.sqrt(mean_squared_error(y_test, lm.predict(X_test)))
# Find the linear regression equation:
[(coef, feature) for coef, feature in zip(lm.coef_, X_train.columns)]
lm.intercept_
# Look at the residuals:
from ml_utils.regression import plot_residuals
plot_residuals(y_test, lm.predict(X_test))
# <hr>
# <div>
# <a href="./exercise_1.ipynb">
# <button>← Previous Solution</button>
# </a>
# <a href="./exercise_3.ipynb">
# <button style="float: right;">Next Solution →</button>
# </a>
# </div>
# <hr>
| solutions/ch_09/exercise_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <table>
# <tr><td><img style="height: 150px;" src="images/geo_hydro1.jpg"></td>
# <td bgcolor="#FFFFFF">
# <p style="font-size: xx-large; font-weight: 900; line-height: 100%">AG Dynamics of the Earth</p>
# <p style="font-size: large; color: rgba(0,0,0,0.5);">Jupyter notebooks</p>
# <p style="font-size: large; color: rgba(0,0,0,0.5);"><NAME></p>
# </td>
# </tr>
# </table>
# # Dynamic systems: 12. Shallow-water equations
# ## 1D advection equation revisited (finite volumes)
# ----
# *<NAME>,
# Geophysics Section,
# Institute of Geological Sciences,
# Freie Universität Berlin,
# Germany*
import numpy as np
import matplotlib.pyplot as plt
# Consider a partial differential equation of the form:
# $$
# \frac{\partial h}{\partial t}
# + u \frac{\partial h}{\partial x} = 0
# $$
# with
# - $h$ [m] height,
# - $t$ [s] time,
# - $x$ [m] distance,
# - $u$ [m/s] advection velocity.
#
# We need an initial condition $h(x,t=0)$ to solve the problem.
#
# Define model domain $x \in [xmin,xmax]$ with $nx$ points, maximum time $T$,
# velocity $u$ as constant and positive,
# and spatial ($dx$) as well time ($dt$) increments. Check **Courant number**!
# +
nx = 101
xmin = 0
xmax = 2
T = 5
x = np.linspace(xmin,xmax,nx)
dt = 0.25
dx = (x.max()-x.min()) / (nx-1)
u = 0.05
g = 9.81
print("dx: ",dx," dt: ",dt," u: ",u)
#print(x)
Courant = u*dt/dx
print ('Courant number: ',Courant)
# -
# ----
# ### 1. Example
#
# Consider the initial condition
# $$
# h(x,t=0) = \sin(2\pi x), \quad x \in [0,1]
# $$
# We want to solve for $h(x,t)$ for $t \in [0,T]$.
# ----
# ### Central finite differences (FTCS)
#
# We rewrite the partial differential equation into **finite differences**,
# with forward differences in time (FT), and central differences in space (CS),
# a scheme called **FTCS**:
# $$
# \frac{h_{i}^{n+1}-h_{i}^{n}}{\Delta t}
# + u \frac{h_{i+1}^{n}-h_{i-1}^{n}}{2\Delta x}
# \simeq 0
# $$
# <img style="height: 200px;" src="images/1Dcoord_FTCS.jpg">
# which can be recasted into the scheme:
# $$
# h_{i}^{n+1} \simeq h_{i}^{n}
# - u \frac{\Delta t}{2\Delta x} \left( h_{i+1}^{n}-h_{i-1}^{n} \right)
# $$
# +
# FTCS scheme
# start time
time = 0
dtplot = 1.00
tplot = dtplot
# initial values
h = np.sin(2*np.pi*x)
plt.plot(x,h,linewidth=6,color='lightgray',label='h$_0$')
# solution
while (time < T):
time = time + dt
### PERIODIC BC ###
h[0] = h[-2]
h[-1] = h[1]
hold = h
for i in range(1,h.shape[0]-1):
h[i] = hold[i] - u*dt/2/dx*(hold[i+1]-hold[i-1])
if (time > tplot):
plt.plot(x,h,label='h(t='+str(round(time,2))+' s)')
plt.legend()
tplot = tplot + dtplot
# -
# **Problem:**
# The use of the forward value $h_{i+1}^{n}$ in space uses
# a value we do not know.
# ----
# ### Upwind finite differences
#
# We rewrite the partial differential equation into **finite differences**,
# with forward differences in time (FT), and backward differences in space (BS),
# a scheme called **upwind**:
# $$
# \frac{h_{i}^{n+1}-h_{i}^{n}}{\Delta t}
# + u \frac{h_{i}^{n}-h_{i-1}^{n}}{\Delta x}
# \simeq 0, \quad u>0
# $$
# <img style="height: 200px;" src="images/1Dcoord_upwind.jpg">
# which can be recasted into the scheme:
# $$
# h_{i}^{n+1} \simeq h_{i}^{n}
# - u \frac{\Delta t}{\Delta x} \left( h_{i}^{n}-h_{i-1}^{n} \right)
# $$
#
# **Note:** This upwind scheme is for $u>0$, for $u<0$ we would need a forward difference ...
# +
# Upwind scheme
# start time
time = 0
dtplot = 1.00
tplot = dtplot
# initial values
h = np.sin(2*np.pi*x)
plt.plot(x,h,linewidth=6,color='lightgray',label='h$_0$')
# solution
while (time < T):
time = time + dt
### PERIODIC BC ###
h[0] = h[-2]
h[-1] = h[1]
hold = h
for i in range(1,h.shape[0]-1):
h[i] = hold[i] - u*dt/dx*(hold[i]-hold[i-1])
if (time > tplot):
plt.plot(x,h,label='h(t='+str(round(time,2))+' s)')
plt.legend()
tplot = tplot + dtplot
# -
# **Problem:**
# Stable, but numerical diffusion.
# ----
# ### Finite-volume Gondurov scheme with Rosanov flux
#
# We reformulate the partial differential equation into a pseudo-non-linear
# one:
# $$
# \frac{\partial h}{\partial t}
# + \frac{\partial (hu)}{\partial x} = 0
# $$
# and we identify the term $hu$ as flux term:
# $$
# f(u) = h u
# $$
# <img style="height: 200px;" src="images/1Dcoord_LLF.jpg">
#
# For the **finite volume method**, we integrate over one cell:
# $$
# \int\limits_{t_{n}}^{t_{n+1}}
# \int\limits_{x_{i-\frac{1}{2}}}^{x_{i+\frac{1}{2}}}
# \frac{\partial h}{\partial t}
# + \frac{\partial (hu)}{\partial x} dx dt = 0
# $$
# Integrate by parts:
# $$
# \int\limits_{x_{i-\frac{1}{2}}}^{x_{i+\frac{1}{2}}}
# h(x,t^{n+1}) - h(x,t^{n}) dx
# +
# \int\limits_{t_{n}}^{t_{n+1}}
# f(x_{i+\frac{1}{2}},t) - f(x_{i-\frac{1}{2}},t)dt
# = 0
# $$
# Final integration:
# $$
# \Delta x \left( h_{i}^{n+1} - h_{i}^{n} \right)
# + \Delta t \left( F_{i+\frac{1}{2}} - F_{i-\frac{1}{2}} \right) = 0
# $$
#
#
#
#
#
# Rewriting the non-linear partial differential equation into
# $$
# h_{i}^{n+1} \simeq h_{i}^{n}
# - \frac{\Delta t}{\Delta x} \left( F_{i+\frac{1}{2}} - F_{i-\frac{1}{2}} \right)
# $$
# with the **Rosanov fluxes** $F_{i+\frac{1}{2}}$ and $F_{i-\frac{1}{2}}$ calculated
# at the cell interfaces:
# $$
# F_{i+\frac{1}{2}} =
# \frac{1}{2} \left[ f(h_i^n) + f(h_{i+1}^n) \right]
# - \frac{\lambda_{max}}{2} \left( h_{i+1}^{n} - h_{i}^{n} \right)
# $$
# The eigen value $\lambda_{max}$ is
# - $\lambda_{max}=u$ for linear advection,
# - $\lambda_{max}=\max(f(u))$ for non-linear advection.
# +
# Finite volume with Rusanov flux scheme
# start time
time = 0
dtplot = 1.00
tplot = dtplot
# initial values
h = np.sin(2*np.pi*x)
plt.plot(x,h,linewidth=6,color='lightgray',label='h$_0$')
# solution
while (time < T):
time = time + dt
### PERIODIC BC ###
h[0] = h[-2]
h[-1] = h[1]
Fhup = h[1:]*u
Fhum = h[:-1]*u
Rh = (Fhup+Fhum)/2 - u/2*(h[1:]-h[:-1])
h[1:-1] = h[1:-1] - dt/dx*(Rh[1:]-Rh[:-1])
if (time > tplot):
plt.plot(x,h,label='h(t='+str(round(time,2))+' s)')
plt.legend()
tplot = tplot + dtplot
# -
# ----
# ### 2. Example
#
# Consider the initial condition
# $$
# h(x,t=0) = e^{-\frac{(x-\mu)^2}{\sigma^2}}, \quad x \in [0,10]
# $$
# with $\mu=2$ and $\sigma=1$.
#
# We want to solve for $h(x,t)$ for $t \in [0,T]$.
# +
nx = 101
xmin = 0
xmax = 10
T = 5
x = np.linspace(xmin,xmax,nx)
dt = 0.05
dx = (x.max()-x.min()) / (nx-1)
u = 0.5 # and test with u=1!!!
g = 9.81
mu = 2
sigma = 1
print("dx: ",dx," dt: ",dt," u: ",u)
#print(x)
Courant = u*dt/dx
print ('Courant number: ',Courant)
# +
# FTCS scheme
# start time
time = 0
dtplot = 1.00
tplot = dtplot
# initial values
h = np.exp(-(x-mu)**2/sigma**2)
plt.plot(x,h,linewidth=6,color='lightgray',label='h$_0$')
# solution
while (time < T):
time = time + dt
### PERIODIC BC ###
h[0] = h[-2]
h[-1] = h[1]
hold = h
for i in range(1,h.shape[0]-1):
h[i] = hold[i] - u*dt/2/dx*(hold[i+1]-hold[i-1])
if (time > tplot):
plt.plot(x,h,label='h(t='+str(round(time,2))+' s)')
plt.legend()
tplot = tplot + dtplot
# +
# Upwind scheme
# start time
time = 0
dtplot = 1.00
tplot = dtplot
# initial values
h = np.exp(-(x-mu)**2/sigma**2)
plt.plot(x,h,linewidth=6,color='lightgray',label='h$_0$')
# solution
while (time < T):
time = time + dt
### PERIODIC BC ###
h[0] = h[-2]
h[-1] = h[1]
hold = h
for i in range(1,h.shape[0]-1):
h[i] = hold[i] - u*dt/dx*(hold[i]-hold[i-1])
if (time > tplot):
plt.plot(x,h,label='h(t='+str(round(time,2))+' s)')
plt.legend()
tplot = tplot + dtplot
# +
# Finite volume with Rusanov flux scheme
# start time
time = 0
dtplot = 1.00
tplot = dtplot
# initial values
h = np.exp(-(x-mu)**2/sigma**2)
plt.plot(x,h,linewidth=6,color='lightgray',label='h$_0$')
# solution
while (time < T):
time = time + dt
### PERIODIC BC ###
h[0] = h[-2]
h[-1] = h[1]
Fhup = h[1:]*u
Fhum = h[:-1]*u
Rh = (Fhup+Fhum)/2 - u/2*(h[1:]-h[:-1])
h[1:-1] = h[1:-1] - dt/dx*(Rh[1:]-Rh[:-1])
if (time > tplot):
plt.plot(x,h,label='h(t='+str(round(time,2))+' s)')
plt.legend()
tplot = tplot + dtplot
# -
# ... done
| Dynamics_lab12_advection1D_FiniteVolume_LLF.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Linear Model Tests
# This notebook generates a lasso model and produces scores for each of the countries. The scores are the train/test MAEs and the "MPEs" (Mean Percentage Error: 100*MAE/Population).
#
# Now using standard scaler for the models
import pickle
import os
import urllib.request
from sklearn.linear_model import Lasso
from sklearn.model_selection import train_test_split
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# ## Importing the Training Data
# +
# Main source for the training data
DATA_URL = 'https://raw.githubusercontent.com/OxCGRT/covid-policy-tracker/master/data/OxCGRT_latest.csv'
# Local files
data_path = 'examples/predictors/ryan_predictor/data'
DATA_FILE = data_path + '/OxCGRT_latest.csv'
if not os.path.exists(data_path):
os.mkdir(data_path)
urllib.request.urlretrieve(DATA_URL, DATA_FILE)
# -
df = pd.read_csv(DATA_FILE,
parse_dates=['Date'],
encoding="ISO-8859-1",
dtype={"RegionName": str,
"RegionCode": str},
error_bad_lines=False)
# df[cases_df['RegionName'] == 'California']
HYPOTHETICAL_SUBMISSION_DATE = np.datetime64("2020-07-31")
df = df[df.Date <= HYPOTHETICAL_SUBMISSION_DATE]
# Add RegionID column that combines CountryName and RegionName for easier manipulation of data
df['GeoID'] = df['CountryName'] + '__' + df['RegionName'].astype(str)
# Add new cases column
df['NewCases'] = df.groupby('GeoID').ConfirmedCases.diff().fillna(0)
# Keep only columns of interest
id_cols = ['CountryName',
'RegionName',
'GeoID',
'Date']
cases_col = ['NewCases']
npi_cols = ['C1_School closing',
'C2_Workplace closing',
'C3_Cancel public events',
'C4_Restrictions on gatherings',
'C5_Close public transport',
'C6_Stay at home requirements',
'C7_Restrictions on internal movement',
'C8_International travel controls',
'H1_Public information campaigns',
'H2_Testing policy',
'H3_Contact tracing',
'H6_Facial Coverings']
df = df[id_cols + cases_col + npi_cols]
# Fill any missing case values by interpolation and setting NaNs to 0
df.update(df.groupby('GeoID').NewCases.apply(
lambda group: group.interpolate()).fillna(0))
# Fill any missing NPIs by assuming they are the same as previous day
for npi_col in npi_cols:
df.update(df.groupby('GeoID')[npi_col].ffill().fillna(0))
# ## Making the Model
# +
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestRegressor
scores_df = pd.DataFrame(columns = ['Country', 'TrainMAE', 'TestMAE'])
# Helpful function to compute mae
def mae(pred, true):
return np.mean(np.abs(pred - true))
do_not_scale_list = ['India', 'Mauritania', 'Philippines', 'Costa Rica']
forest_list = ['Italy', 'Egypt', 'Iraq', 'Singapore', 'Poland', 'Pakistan'
'Germany', 'Peru', 'Central African Republic', 'Guinea', 'Palestine',
'France', 'Ecuador', 'Tanzania', 'Kyrgyz Republic']
# The models for these countries were found to perform significantly better using:
# - unscaled data for a linear regression
# and
# - random forest
scaler = StandardScaler()
model = Lasso(alpha=0.1, precompute=True, max_iter=10000, positive=True, selection='random')
for country in df['CountryName'].unique().tolist():
country_df = df[df['CountryName'] == country]
# Set number of past days to use to make predictions
nb_lookback_days = 30
# Create training data across all countries for predicting one day ahead
X_cols = cases_col + npi_cols
y_col = cases_col
X_samples = []
y_samples = []
geo_ids = country_df.GeoID.unique()
for g in geo_ids:
gdf = country_df[country_df.GeoID == g]
all_case_data = np.array(gdf[cases_col])
all_npi_data = np.array(gdf[npi_cols])
# Create one sample for each day where we have enough data
# Each sample consists of cases and npis for previous nb_lookback_days
nb_total_days = len(gdf)
for d in range(nb_lookback_days, nb_total_days - 1):
X_cases = all_case_data[d-nb_lookback_days:d]
# Take negative of npis to support positive
# weight constraint in Lasso.
X_npis = -all_npi_data[d - nb_lookback_days:d]
# Flatten all input data so it fits Lasso input format.
X_sample = np.concatenate([X_cases.flatten(),
X_npis.flatten()])
y_sample = all_case_data[d + 1]
X_samples.append(X_sample)
y_samples.append(y_sample)
X_samples = np.array(X_samples)
y_samples = np.array(y_samples).flatten()
# Split data into train and test sets
X_train, X_test, y_train, y_test = train_test_split(X_samples, y_samples, test_size=0.2, random_state=42)
if country in do_not_scale_list:
model = Lasso(alpha=0.1, precompute=True, max_iter=10000, positive=True, selection='random')
model.fit(X_train, y_train)
elif country in forest_list:
model = RandomForestRegressor(max_depth=2, random_state=0)
model.fit(X_train, y_train)
else:
X_train = scaler.fit_transform(X_train)
X_test = scaler.transform(X_test)
model = Lasso(alpha=0.1, precompute=True, max_iter=10000, positive=True, selection='random')
model.fit(X_train, y_train)
# Evaluate model
train_preds = model.predict(X_train)
train_preds = np.maximum(train_preds, 0) # Don't predict negative cases
# print('Train MAE:', mae(train_preds, y_train))
test_preds = model.predict(X_test)
test_preds = np.maximum(test_preds, 0) # Don't predict negative cases
# print('Test MAE:', mae(test_preds, y_test))
score_df = pd.DataFrame([[country,
mae(train_preds, y_train),
mae(test_preds, y_test)]],
columns=['Country', 'TrainMAE', 'TestMAE'])
scores_df = scores_df.append(score_df)
scores_df
# -
og_df = pd.read_csv(DATA_FILE,
parse_dates=['Date'],
encoding="ISO-8859-1",
dtype={"RegionName": str,
"RegionCode": str},
error_bad_lines=False)
og_df['GeoID'] = og_df['CountryName'] + '__' + og_df['RegionName'].astype(str)
geoid_cases = og_df.groupby('GeoID').agg({'ConfirmedCases':np.median}).reset_index()
geoid_cases = geoid_cases.merge(og_df[['GeoID','CountryName']], how='left', left_on='GeoID', right_on='GeoID')
geoid_cases = geoid_cases.groupby('CountryName').agg({'ConfirmedCases':np.sum}).reset_index()
geoid_cases
scores_df = scores_df.merge(geoid_cases, how='left', left_on='Country', right_on='CountryName').drop(['CountryName'], axis=1)
scores_df
scores_df['TrainMPE'] = 100*scores_df['TrainMAE']/scores_df['ConfirmedCases']
scores_df['TestMPE'] = 100*scores_df['TestMAE']/scores_df['ConfirmedCases']
scores_df.sort_values(by='TestMPE').reset_index()
scores_df.sort_values(by='TestMPE').reset_index().to_csv('case_pred_errors_as_percent.csv', index=False)
scores_df = scores_df.sort_values(by='TestMPE').reset_index()
lin_no_scale = pd.read_csv('lin_vs_rand_forest.csv')
lin_no_scale[['Country','TrainMAE_x','TestMAE_x','ConfirmedCases']]
lin_no_scale[['Country','TrainMAE_x','TestMAE_x','ConfirmedCases']].merge(scores_df, how='left', left_on='Country', right_on='Country')
lin_no_scale[['Country','TrainMAE_x','TestMAE_x','ConfirmedCases']].merge(
scores_df, how='left', left_on='Country', right_on='Country').to_csv('lin_noscale_v_scale.csv')
top_50_country_list = scores_df.sort_values(by='TestMPE')['Country'].tolist()[:50]
import pickle
pickle.dump(top_50_country_list, open( "top_50_country_list.p", "wb" ) )
# ## Evaluating the Scores
scores_df = scores_df[scores_df['TestMAE'] != 0].sort_values(by='TestMAE')
country_pops = pd.read_csv('countrypops.csv')
country_pops = country_pops[['Country', 'Population']]
scores_df.head()
scores_w_pops = scores_df.merge(country_pops, how = 'left', left_on = 'Country', right_on = 'Country')
population_list = scores_w_pops['Population'].tolist()
for i, val in enumerate(population_list):
if type(val) not in [int, float]:
population_list[i] = float(val.replace(',',''))
# else:
# population_list[i] = 0
scores_w_pops['Population'] = population_list
#MPE = Mean Percentage Error (I made this term up)
scores_w_pops['TrainMPE'] = 100*scores_w_pops['TrainMAE']/scores_w_pops['Population']
scores_w_pops['TestMPE'] = 100*scores_w_pops['TestMAE']/scores_w_pops['Population']
scores_w_pops[['Country','TestMAE','Population', 'TestMPE']].sort_values(by='TestMPE')
scores_w_pops['TrainMPE'] = 100*scores_w_pops['TrainMAE']/scores_w_pops['Population']
scores_w_pops['TestMPE'] = 100*scores_w_pops['TestMAE']/scores_w_pops['Population']
scores_w_pops.sort_values(by='TestMPE').to_csv('case_pred_errors_as_percent.csv')
| covid_xprize/RyanModelCountryScoreswScaling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#dependencies
import pandas as pd
from sqlalchemy import create_engine
from config import db_password
# -
# # Extract CSVs into DataFrames
# +
#loading CSV files
file1 = "Drugs_package.csv"
package= pd.read_csv(file1)
package.head()
# -
package = package.drop_duplicates(subset=['PRODUCTID'])
# +
package = package.copy()
#rename the columns
package = package.rename(columns={"PRODUCTID": "productid",
"PRODUCTNDC": "productndc",
"NDCPACKAGECODE": "ndcpackagecode",
"PACKAGEDESCRIPTION": "packagedescription"})
#set index
package.set_index("productid", inplace=True)
package.head()
# +
#package.columns
# +
# file2 = "Drugs_product.csv"
# product = pd.read_csv(file2)
# product.head()
# +
# print(product.columns)
# +
file3 = "Nutritions_US.csv"
nutrition = pd.read_csv(file3)
nutrition.head()
# -
nutrition.columns
nut_transformed = nutrition[["NDB_No", "Shrt_Desc", "Water_(g)", "Energ_Kcal"]].copy()
nut_transformed
# +
#rename columns
nut_transformed = nut_transformed.rename(columns={"NDB_No": "id",
"Shrt_Desc": "description",
"Water_(g)": "water_g",
"Energ_Kcal": "energy_kcal"})
nut_transformed.set_index("id", inplace=True)
nut_transformed.head()
# -
# ### Connect to local database
rds_connection_string = f"postgres:{db_password}@localhost:5432/ETL"
engine = create_engine(f'postgresql://{rds_connection_string}')
engine.table_names()
# ### Load DataFrames into database
package.to_sql(name='drugs_package', con=engine, if_exists='append', index=True)
# +
#pd.read_sql_query('select * from drugs_package', con=engine)
# -
nut_transformed.to_sql(name='nutrition', con=engine, if_exists='append', index=True)
# ### Confirm data has been added by querying the tables
pd.read_sql_query('select * from nutrition', con=engine)
pd.read_sql_query('select * from drugs_package', con=engine)
| .ipynb_checkpoints/ETL process-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
from matplotlib.patches import Circle
import numpy as np
import os
import glob
import cv2
DATA_DIR = "D:\\Data\\Dispertech\\2020-01-27"
for file in glob.glob(os.path.join(DATA_DIR, '*.npy')):
filename = file.split('\\')[-1]
data = np.load(file)
fig, ax = plt.subplots(figsize=(11,10), dpi=100)
plt.imshow(data.T)
plt.title(filename)
plt.savefig(file+'.png')
plt.close()
file = "fiber_end_1902060_0.npy"
filename = os.path.join(DATA_DIR,file)
data = np.load(os.path.join(DATA_DIR, filename))[450:850, 450:850]
# test_img = mask*np.ones(data.shape, dtype=np.uint8)*2**8
circles = cv2.HoughCircles(np.uint8(data),cv2.HOUGH_GRADIENT,10,50,param1=50,param2=30,minRadius=100,maxRadius=0)
fig, ax = plt.subplots(2, figsize=(11,10), dpi=100)
fig.set()
y = 208
x = 238
ax[0].imshow(data.T)
ax[0].axhline(y=y,color='red')
ax[0].axvline(x=x,color='red')
ax[1].plot(data[:,y])
ax[1].plot(data[x,:])
img = np.load(os.path.join(DATA_DIR, filename))
img = gaussian(img, sigma=2)
mask = img>.005
d = np.zeros(img.shape, dtype=np.uint8)
d[mask] = 255
# d = gaussian(d, sigma=2)
fig, ax = plt.subplots(figsize=(11,10), dpi=100)
plt.imshow(d)
circles = cv2.HoughCircles(d,cv2.HOUGH_GRADIENT,200,500,param1=50,param2=50,minRadius=45,maxRadius=350)
for c in circles[0, :]:
circ = Circle((c[0],c[1]),c[2], alpha=0.4, facecolor=None)
ax.add_patch(circ)
from skimage import data, color
from skimage.transform import hough_circle, hough_circle_peaks
from skimage.feature import canny
from skimage.draw import circle_perimeter
from skimage.util import img_as_ubyte
from skimage.filters import gaussian
img = np.load(os.path.join(DATA_DIR, filename))
image = img_as_ubyte(img)
mask = img>130
d = np.zeros(img.shape, dtype=np.uint8)
d[mask] = 255
d = gaussian(d)
plt.imshow(d)
plt.hist(img.reshape(1228800,1))
| notebooks/View_Saved_File.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pymaceuticals, Analysis in Drug testing Data
#
# Pymaceuticals specializes in drug-based, anti-cancer pharmaceuticals. In their most recent efforts, they've since begun screening for potential treatments to squamous cell carcinoma (SCC), a commonly occurring form of skin cancer.
# #
# As Data Analyst, you've been given access to the complete data from their most recent animal study. In this study, 250 mice were treated through a variety of drug regimes over the course of 45 days. Their physiological responses were then monitored over the course of that time. Your objective is to analyze the data to show how four treatments (Capomulin, Infubinol, Ketapril, and Placebo) compare.
#
#
#
# 1- Creating a scatter plot that shows how the tumor volume changes over time for each treatment.
#
# 2- Creating a scatter plot that shows how the number of metastatic (cancer spreading) sites changes over time for each treatment.
#
# 3- Creating a scatter plot that shows the number of mice still alive through the course of treatment (Survival Rate)
#
# +
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import warnings
warnings.filterwarnings('ignore')
path_clinic_data = "Resources/Pymaceuticals_clinicaltrial_data.csv"
path_mouse_data = "Resources/Pymaceuticals_mouse_drug_data.csv"
clinic_data = pd.read_csv(path_clinic_data)
mouse_data = pd.read_csv(path_mouse_data)
clinic_data.head()
mouse_data.head()
merged_data = pd.merge(clinic_data, mouse_data, how="outer",on="Mouse ID" )
#merged_data.head(15)
# -
# # 1 - Scatter plot that shows how the tumor volume changes over time for each treatment.
#
# +
#IMPORT
from scipy.stats import sem
#GROUP THE DATA FRAME BY DRUG AND TIME
grouped_drug = merged_data.groupby(["Drug", "Timepoint"])
#GETTING THE MEAN() OF TUMOR COLUMN AND PUT IN DATA FRAME
Tumor_mean = grouped_drug["Tumor Volume (mm3)"].mean()
df1 = pd.DataFrame({ "Tumor Volume (mm3)" : Tumor_mean})
#GETTING THE STANDARD ERROR OF TUMOR AND PUT IN A DATA FRAME
Standard_error = grouped_drug["Tumor Volume (mm3)"].sem()
df2 = pd.DataFrame({"Standard Error" : Standard_error})
#df1
#df2
# +
#RESENTING THE INDEX
ResetIndex_by_drugs = df1.reset_index()
ResetIndex_by_error = df2.reset_index()
#SETING THE INDEX BY TIME AND SETTING COLUMNS BY DRUGS
Mean_index = ResetIndex_by_drugs.pivot(columns= "Drug", index= "Timepoint")["Tumor Volume (mm3)"]
df3 = pd.DataFrame(Mean_index)
#SETING THE INDEX BY TIME AND SETTING COLUMNS BY ERROR
Error_index = ResetIndex_by_error.pivot(columns= "Drug", index= "Timepoint")["Standard Error"]
df4 = pd.DataFrame(Error_index)
#df3.head(5)
#df4
# +
#PLOTTING THE ERROR BAR USING MEAN AS REFERENCE REMOVING THE NOISE IN THE GRAPH
x_axis = np.arange(0, 50, 10)
plt.errorbar(df3.index, df3["Capomulin"], yerr=df4["Capomulin"],
color="red", marker="o", markersize=5, linestyle="dashed", linewidth=0.3)
plt.errorbar(df3.index, df3["Infubinol"], yerr= df4["Infubinol"],
color="green", marker="x", markersize=5, linestyle="dashed",linewidth=0.3)
plt.errorbar(df3.index, df3["Ketapril"], yerr=df4["Ketapril"],
color="blue", marker="*", markersize=5, linestyle="dashed", linewidth=0.3)
plt.errorbar(df3.index, df3["Placebo"], yerr= df4["Placebo"],
color="brown", marker="x", markersize=5, linestyle="dashed",linewidth=0.3)
#PUTTING LEGEND, LABELS, CUSTOMIZING..
plt.title('Tumor vol(mm3) Increase over time')
plt.xlabel('Treatment Time (Days)')
plt.ylabel("Tumor vol(mm3)")
plt.style.use('seaborn-whitegrid')
plt.grid(linestyle="dashed")
plt.legend(loc='best', fontsize=12, fancybox=True)
plt.savefig("MetastaticSpreadDuringTreatment.png")
plt.figure(figsize=(3,3))
plt.show()
# -
# # 2 - Scatter plot that shows how the number of metastatic (cancer spreading) sites changes over time for each treatment.
# +
#GETTING THE MEAN() OF METASTATIC COLUMN AND PUT IN DATA FRAME
grouped_drug2 = merged_data.groupby(["Drug","Timepoint"])
Tumor_mean2 = grouped_drug2["Metastatic Sites"].mean()
df3 = pd.DataFrame({ "Metastatic Mean()" : Tumor_mean2})
#GETTING THE STANDARD ERROR OF TUMOR AND PUT IN A DATA FRAME
Standard_error2 = grouped_drug2["Metastatic Sites"].sem()
df4 = pd.DataFrame({"Metastatic Sem()" : Standard_error2})
#df3.head(5)
#df4
# +
#RESENTING THE INDEX
ResetIndex_df3 = df3.reset_index()
ResetIndex_df4 = df4.reset_index()
#SETING THE INDEX BY TIME AND SETTING COLUMNS BY DRUGS
Mean_index2 = ResetIndex_df3.pivot(columns= "Drug", index="Timepoint")["Metastatic Mean()"]
df3_met = pd.DataFrame(Mean_index2)
#SETING THE INDEX BY TIME AND SETTING COLUMNS BY ERROR
Error_index2 = ResetIndex_df4.pivot(columns= "Drug", index= "Timepoint")["Metastatic Sem()"]
df4_met = pd.DataFrame(Error_index2)
#df3_met
#df4_met
# +
#PLOTING THE ERROR BAR
plt.errorbar(df3_met.index, df3_met["Capomulin"], yerr=df4_met["Capomulin"],
color="red", marker="o", markersize=5, linestyle="dashed", linewidth=0.50)
plt.errorbar(df3_met.index, df3_met["Infubinol"], yerr= df4_met["Infubinol"],
color="green", marker="x", markersize=5, linestyle="dashed",linewidth=0.3)
plt.errorbar(df3_met.index, df3_met["Ketapril"], yerr=df4_met["Ketapril"],
color="blue", marker="*", markersize=5, linestyle="dashed", linewidth=0.3)
plt.errorbar(df3_met.index, df3_met["Placebo"], yerr= df4_met["Placebo"],
color="brown", marker="x", markersize=5, linestyle="dashed",linewidth=0.3)
# Set x and y axis labels including the title of the chart
plt.title('Metastatic Spread During Treatment') # Give plot main title
plt.xlabel('Treatment Duration (Days)') # set text for the x axis
plt.ylabel('Met. Sites') # set text for the y axis
plt.style.use('seaborn-whitegrid')
plt.grid(linestyle="dashed")
plt.legend(loc='best', fontsize=12, fancybox=True)
# Save the Figure
plt.savefig("MetastaticSpreadDuringTreatment.png")
# Show the Figure
plt.show()
# -
# # 3 - Scatter plot that shows the number of mice still alive through the course of treatment (Survival Rate)
# +
#COUNTING NUMBER OF MOUSES AND PUTTING IN DATA FRAME
grouped_drug3= merged_data.groupby(["Drug","Timepoint"])
mouse_count = grouped_drug3["Mouse ID"].count()
df5 = pd.DataFrame({ "Mouse Count" : mouse_count })
#RESET INDEX --> DATA MUNGING TO HAVE "TIMEPOINT AS INDEX AN "DRUGS AS COLUMNS
ResetIndex_df5 = df5.reset_index()
df5_mouse = ResetIndex_df5.pivot(columns="Drug", index="Timepoint")['Mouse Count']
#df5_mouse
# +
#MAKING A REGULAR PLOT CALCULATING THE PERCENTAGE
plt.plot((100*df5_mouse["Capomulin"]/ 25),
color="red", marker="o", markersize=5, linestyle="dashed",linewidth=0.3)
plt.plot(100*df5_mouse["Infubinol"]/25,
color="green", marker="x", markersize=5, linestyle="dashed",linewidth=0.3)
plt.plot(100* df5_mouse["Ketapril"]/25,
color="blue", marker="*", markersize=5, linestyle="dashed", linewidth=0.3)
plt.plot(100* df5_mouse["Placebo"]/25,
color="brown", marker="x", markersize=5, linestyle="dashed", linewidth=0.3)
plt.title('Mouse Survival rate, overtime')
plt.xlabel('Treatment Duration (Days)') # set text for the x axis
plt.ylabel('Mouse Percentage') # set text for the y axis
plt.style.use('seaborn-whitegrid')
plt.grid(linestyle="dashed")
plt.legend(loc='best', fontsize=12, fancybox=True)
# Save the Figure
plt.savefig("MetastaticSpreadDuringTreatment.png")
plt.show()
# -
#
| Pharmaceutics Analysis, Testing a drugs product.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
import math
from torch import optim
from torch import Tensor
from torch.autograd import Variable
from torch import nn
from torch.nn import functional as F
import dlc_practical_prologue
size=1000;
train_input, train_target, train_classes, test_input, test_target, test_classes = \
dlc_practical_prologue.generate_pair_sets(size)
train_input, train_target, train_classes = Variable(train_input), Variable(train_target), Variable((train_classes))
test_input, test_target, test_classes = Variable(test_input), Variable(test_target), Variable(test_classes)
mini_batch_size = 10
# +
import matplotlib.pyplot as plt
fig = plt.figure()
for i in range(6):
plt.subplot(2,3,i+1)
plt.imshow(test_input[i][0], cmap='gray', interpolation='none')
plt.title("Value: {}".format(test_classes[i][0]))
plt.tight_layout()
plt.xticks([])
plt.yticks([])
# -
for i in range(6):
plt.subplot(2,3,i+1)
plt.imshow(test_input[i][1], cmap='gray', interpolation='none')
plt.title("Value: {}".format(test_classes[i][1]))
plt.tight_layout()
plt.xticks([])
plt.yticks([])
# # First neural net
# Train the model on all the 2000 images in train_input
my_train_input = train_input.reshape([2000,196])
my_train_classes = train_classes.reshape([2000])
my_test_input = test_input.reshape([2000,196])
my_test_classes = test_classes.reshape([2000])
# +
# device
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
# number of epoch
num_epochs = 25
# batch size to compute mini-batch
batch_size = 50
# number of pixels in the image
input_size = 196
# number of possible digit: 0 to 9
num_class = 10
# small step to find a minima
learning_rate = 0.005
# hidden size
hidden_size = 200
# Fully connected neural network with one hidden layer
class NeuralNet(nn.Module):
def __init__(self, input_size, hidden_size, num_class):
super(NeuralNet, self).__init__()
self.layer1 = nn.Linear(input_size, hidden_size)
self.relu = nn.ReLU()
self.layer2 = nn.Linear(hidden_size, hidden_size)
self.layer3 = nn.Linear(hidden_size, num_class)
def forward(self, x):
outputs = self.layer1(x)
outputs = self.relu(outputs)
outputs = self.layer2(outputs)
outputs = self.relu(outputs)
outputs = self.layer3(outputs)
return outputs
# creating neural net
model = NeuralNet(input_size, hidden_size, num_class).to(device)
# CrossEntropyLoss and optimizer which minimize loss with learning rate step
criterion = nn.CrossEntropyLoss()
optimizer = torch.optim.SGD(model.parameters(), lr=learning_rate)
# -
def test_accuracy(model_, my_test_input_, my_test_classes_):
total = my_test_input_.size(0)
outputs = model_(my_test_input_)
_, predictions = torch.max(outputs.data, 1)
well_predicted_count = (predictions == my_test_classes_).sum().item()
return 1 - well_predicted_count / total
# +
def train_model(model_, my_train_input_, my_train_classes_, criterion_, optimizer_,num_epochs_,batch_size_):
train_error = []
test_error = []
# train function
for epoch in range(1, num_epochs_+1):
for i in range(int(len(my_train_input_)/batch_size_)):
# Move tensors to the configured device
images = my_train_input_.narrow(0,i*batch_size_,batch_size_).to(device)
labels = my_train_classes_.narrow(0,i*batch_size_,batch_size_).to(device)
# Forward pass
outputs = model_(images)
loss = criterion_(outputs, labels)
# Backward and optimize
optimizer_.zero_grad()
loss.backward()
optimizer_.step()
train_error.append(test_accuracy(model_, my_train_input_, my_train_classes_))
test_error.append(test_accuracy(model_, my_test_input, my_test_classes))
if(epoch % 5 == 0 or epoch == 1):
print ('Loss: {:.4f} on epoch: {}, train error: {:.5f}, test error: {:.5f}'.format(loss.item(),epoch,train_error[-1],test_error[-1]))
return train_error, test_error
train_error, test_error = train_model(model, my_train_input, my_train_classes, criterion, optimizer,num_epochs,batch_size)
# -
import matplotlib.pyplot as plt
plt.plot(train_error, 'r--', test_error, 'b--')
plt.ylabel('some numbers')
plt.title("Train error in red, test error in blue over the epoch")
plt.show()
# +
#as we got 100% on the train error it seems that we just leared by heart the train dataset
# let's adding regularization:
class NeuralNet_reg(nn.Module):
def __init__(self, input_size, hidden_size, num_class):
super(NeuralNet_reg, self).__init__()
self.layer1 = nn.Linear(input_size, hidden_size)
self.relu = nn.ReLU()
self.layer2 = nn.Linear(hidden_size, hidden_size)
self.layer3 = nn.Linear(hidden_size, hidden_size)
self.layer4 = nn.Linear(hidden_size, num_class)
self.dropout = nn.Dropout(0.7)
def forward(self, x):
outputs = self.layer1(x)
outputs = self.relu(outputs)
outputs = self.layer2(outputs)
outputs = self.relu(outputs)
outputs = self.layer3(outputs)
outputs = self.relu(outputs)
outputs = self.layer4(outputs)
return outputs
# creating neural net
model_reg = NeuralNet_reg(input_size, hidden_size, num_class).to(device)
# CrossEntropyLoss and optimizer which minimize loss with learning rate step
criterion_reg = nn.CrossEntropyLoss()
optimizer_reg = torch.optim.SGD(model_reg.parameters(), lr=learning_rate)
train_error, test_error = train_model(model_reg, my_train_input, my_train_classes, criterion_reg, optimizer_reg, num_epochs, batch_size)
# -
plt.plot(train_error, 'r--', test_error, 'b--')
plt.ylabel('some numbers')
plt.title("Train error in red, test error in blue over the epoch")
plt.show()
# # Testing on the 2 batches
# A model for the two batch of 1000 images
my_test_input1 = test_input[:,0,:]
my_test_input2 = test_input[:,1,:]
my_test_classes1 = test_classes[0:1000,0]
my_test_classes2 = test_classes[0:1000,1]
# verifying the digits and labels
for i in range(6):
plt.subplot(2,3,i+1)
plt.imshow(my_test_input2[i], cmap='gray', interpolation='none')
plt.title("Value: {}".format(my_test_classes2[i]))
plt.tight_layout()
plt.xticks([])
plt.yticks([])
my_test_input1=my_test_input1.reshape(1000,196)
my_test_input2=my_test_input2.reshape(1000,196)
print("Error on batch 1: {}".format(test_accuracy(model, my_test_input1, my_test_classes1)))
print("Error on batch 2: {}".format(test_accuracy(model, my_test_input2, my_test_classes2)))
# # Building neural net for Digit Comparison
# +
# number of epoch
num_epochs_dc = 21
# batch size to compute mini-batch
batch_size_dc = 20
# number of pixels in the image
input_size_dc = 2
# number of possible digit: 0 to 9
num_class_dc = 1
# small step to find a minima
learning_rate_dc = 0.004
# hidden size
hidden_size_dc = 200
# Fully connected neural network with one hidden layer
class NeuralNet_dc(nn.Module):
def __init__(self, input_size, hidden_size, num_class):
super(NeuralNet_dc, self).__init__()
self.layer1 = nn.Linear(input_size, hidden_size)
self.relu = nn.ReLU()
self.layer2 = nn.Linear(hidden_size, hidden_size)
self.layer3 = nn.Linear(hidden_size, num_class)
def forward(self, x, batch_size_):
outputs = self.layer1(x)
outputs = self.relu(outputs)
outputs = self.layer2(outputs)
outputs = self.relu(outputs)
outputs = self.layer3(outputs)
return outputs.reshape([1, batch_size_])[0]
# creating neural net
model_dc = NeuralNet_dc(input_size_dc, hidden_size_dc, num_class_dc).to(device)
# MSELoss and optimizer which minimize loss with learning rate step
criterion_dc = nn.MSELoss()
optimizer_dc = torch.optim.SGD(model_dc.parameters(), lr=learning_rate_dc)
# train function
for epoch in range(num_epochs_dc):
for i in range(int(len(train_classes)/batch_size_dc)):
# Move tensors to the configured device
inputs = train_classes.narrow(0,i*batch_size_dc,batch_size_dc).to(device).float()
labels = train_target.narrow(0,i*batch_size_dc,batch_size_dc).to(device)
# Forward pass
outputs = model_dc(inputs,batch_size_dc)
loss = criterion_dc(outputs, labels.float())
# Backward and optimize
optimizer_dc.zero_grad()
loss.backward()
optimizer_dc.step()
if(epoch % 5 == 0):
print ('Loss: {:.4f} on epoch: {}'.format(loss.item(),epoch+1))
# +
# test function
def test_accuracy_dc(model_, my_test_input_, my_test_classes_):
total = my_test_input_.size(0)
outputs = model_(my_test_input_.float(),total)
t = Variable(torch.Tensor([0.5]))
predictions = (outputs > t).float() * 1
well_predicted_count = (predictions == my_test_classes_.float()).sum().item()
print('Accuracy: {} %'.format(100 * well_predicted_count / total))
test_accuracy_dc(model_dc, test_classes, test_target)
# -
# # Running both neural networks in sequence
# +
# test function
def getting_y(model_, my_test_input_, my_test_classes_):
total = my_test_input_.size(0)
outputs = model_(my_test_input_)
_, predictions = torch.max(outputs.data, 1)
well_predicted_count = (predictions == my_test_classes_).sum().item()
print('Accuracy: {} %'.format(100 * well_predicted_count / total))
return predictions
y1 = getting_y(model_reg, my_test_input1, my_test_classes1)
y2 = getting_y(model_reg, my_test_input2, my_test_classes2)
y_test_classes = torch.tensor([y1.tolist(), y2.tolist()]).transpose_(0, 1)
y_test_classes
# +
# test function
def digit_comparison(model_, my_test_input_, my_test_classes_):
total = my_test_input_.size(0)
outputs = model_(my_test_input_.float())
t = Variable(torch.Tensor([0.5]))
predictions = (outputs > t).float() * 1
well_predicted_count = (predictions == my_test_classes_.float()).sum().item()
print('Accuracy: {} %'.format(100 * well_predicted_count / total))
test_accuracy_dc(model_dc, y_test_classes, test_target)
# -
| .ipynb_checkpoints/2_fullyconnected_neural_net_in_sequence_95.8%-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Setting up a dataset
# +
import patients
number_of_patients = 100
curation_time_treatment = patients.treatment_group(number_of_patients)
curation_time_placebo = patients.placebo_group(number_of_patients)
print(curation_time_treatment)
print(curation_time_placebo)
# -
# # Descriptive statistics
# +
import numpy as np
def do_descriptive_statistics(x):
mean_x = np.mean(x)
standard_deviation_x = np.std(x)
print("Mean: " + str(mean_x) + " +- " + str(standard_deviation_x))
# +
print("Treatment group")
do_descriptive_statistics(curation_time_treatment)
print("Placebo group")
do_descriptive_statistics(curation_time_placebo)
# +
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
ax.hist(curation_time_treatment, bins=10)
ax.set_title('Curation time of ' + str(len(curation_time_treatment)) + ' patients treated')
ax.set_ylabel("count")
ax.set_xlabel("Curation time / days")
plt.show()
fig, ax = plt.subplots()
ax.hist(curation_time_placebo, bins=10)
ax.set_title('Curation time of ' + str(len(curation_time_placebo)) + ' patients receiving a placebo')
ax.set_ylabel("count")
ax.set_xlabel("Curation time / days")
plt.show()
# +
from my_statistics_functions import draw_curation_time_histogram
draw_curation_time_histogram(curation_time_treatment,
"patients receiving a treatment")
draw_curation_time_histogram(curation_time_placebo,
"patients receiving a placebo")
# -
# # Hypothesis testing
#
# * null-hypothesis: Patients receiving the treatment feel better earlier
#
# * alternate hypothesis: Patients receiving the placebo need longer to feel better
#
# ## Two-sample T-test of independent samples
# +
from scipy import stats
presumptive_ripe_time = 25
statistics, p_value = stats.ttest_ind(curation_time_treatment, curation_time_placebo)
print("p-value: " + str(p_value))
# -
#
| docs/44_hypothesis_testing/Testing curation time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # freud.environment.BondOrder
# ## Computing Bond Order Diagrams
# The `freud.environment` module analyzes the local environments of particles. In this example, the `freud.environment.BondOrder` class is used to plot the bond order diagram (BOD) of a system of particles.
import freud
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
from mpl_toolkits.mplot3d import Axes3D
# ### Setup
# Our sample data will be taken from an face-centered cubic (FCC) structure. The array of points is rather large, so that the plots are smooth. Smaller systems may need to gather data from multiple frames in order to smooth the resulting array's statistics, by computing multiple times with `reset=False`.
uc = freud.data.UnitCell.fcc()
box, points = uc.generate_system(40, sigma_noise=0.05)
# Now we create a `BondOrder` compute object and create some arrays useful for plotting.
# +
n_bins_theta = 100
n_bins_phi = 100
bod = freud.environment.BondOrder((n_bins_theta, n_bins_phi))
phi = np.linspace(0, np.pi, n_bins_phi)
theta = np.linspace(0, 2 * np.pi, n_bins_theta)
phi, theta = np.meshgrid(phi, theta)
x = np.sin(phi) * np.cos(theta)
y = np.sin(phi) * np.sin(theta)
z = np.cos(phi)
# -
# ### Computing the Bond Order Diagram
# Next, we use the `compute` method and the `bond_order` property to return the array.
bod_array = bod.compute(
system=(box, points), neighbors={"num_neighbors": 12}
).bond_order
# Clean up polar bins for plotting
bod_array = np.clip(bod_array, 0, np.percentile(bod_array, 99))
plt.imshow(bod_array.T)
plt.show()
# ### Plotting on a sphere
# This code shows the bond order diagram on a sphere as the sphere is rotated. The code takes a few seconds to run, so be patient.
fig = plt.figure(figsize=(12, 8))
for plot_num in range(6):
ax = fig.add_subplot(231 + plot_num, projection="3d")
ax.plot_surface(
x,
y,
z,
rstride=1,
cstride=1,
shade=False,
facecolors=matplotlib.cm.viridis(bod_array / np.max(bod_array)),
)
ax.set_xlim(-1, 1)
ax.set_ylim(-1, 1)
ax.set_zlim(-1, 1)
ax.set_axis_off()
# View angles in degrees
view_angle = 0, plot_num * 15
ax.view_init(*view_angle)
plt.show()
# ### Using Custom Neighbors
# We can also use a custom neighbor query to determine bonds. For example, we can filter for a range of bond lengths. Below, we only consider neighbors between $r_{min} = 2.5$ and $r_{max} = 3$ and plot the resulting bond order diagram.
bod_array = bod.compute(
system=(box, points), neighbors={"r_max": 3.0, "r_min": 2.5}
).bond_order
# Clean up polar bins for plotting
bod_array = np.clip(bod_array, 0, np.percentile(bod_array, 99))
plt.imshow(bod_array.T)
plt.show()
| module_intros/environment.BondOrder.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.4.0
# language: julia
# name: julia-1.4
# ---
# + [markdown] colab_type="text" id="lRjH7kTOuAie"
# # Julia 安裝及簡介
#
#
# ## 1. 查看 Julia 安裝版本及系統資訊
#
# 呼叫 `versioninfo()` 互動式工具 (Interactive Utilities) 函式 (Function)
#
# 確認 Julia, IJulia 環境安裝成功,並顯示 Julia 版本以及系統資訊
# + colab={} colab_type="code" id="dev1xg2IuAim" outputId="db608f87-4488-4684-adeb-02975b6bd624"
versioninfo()
# + [markdown] colab_type="text" id="F0bcpyUWuAjE"
# ## 2. Hello Julia
#
# 第一個 Julia 程式:透過 `println` 函式,印出 Hello Julia 字串
# + colab={} colab_type="code" id="N9QFB3z9uAjK" outputId="df22f8a3-5a62-48f4-c459-f3d83fc4af15"
println("Hello Julia")
# + [markdown] colab_type="text" id="uFVzX7aXuAjh"
# ## 3. 套件管理
#
# 使用內建的套件管理 (Package Management) 工具,查看目前已安裝的套件及版本。
# + colab={} colab_type="code" id="XbZwHdO3uAjn" outputId="8ff5c4be-46fe-4b40-90d9-64712b521ea7"
using Pkg
Pkg.installed()
# + [markdown] colab_type="text" id="JGkXrvNHuAj5"
# ## 安裝套件
#
# 試著用 Pkg 安裝新的套件
#
# 在這邊我們安裝 DataFrames 套件,會在之後的課程中用到
# + [markdown] colab={} colab_type="code" id="Rsn65UPQuAj_" outputId="d31fcff5-8283-4c91-d05a-87580b87c11c"
# Pkg.add("DataFrames")
# + [markdown] colab_type="text" id="8iWdb2JLuAkN"
# ## 移除套件
# + colab={} colab_type="code" id="yWTGGI6yuAkS" outputId="304da5d4-be6f-4dab-f50c-c0a13d0afa5d"
Pkg.rm("DataFrames")
# + [markdown] colab_type="text" id="YG0HB_WIuAkp"
# ## 4. 內建函式 include
#
# 透過 include 來執行 (或包含) 既有的 Julia 程式
#
# 在 "hello_julia.jl" 程式裡面,程式內容是列印字串
# ```julia
# println("Hello Julia")
# ```
# + colab={} colab_type="code" id="MMm-o_UFuAkt" outputId="33d1d6bf-e8a4-48ea-b439-5a10a3b1e17f"
include("hello_julia.jl")
| example/julia_001_example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/dli-invest/dli-invest.github.io/blob/master/ibook/notebooks/analyze/pandas_ta_plots_ip.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="e7hbDsgYThVb" outputId="37e07475-e662-491b-f010-fa0734318f3b" colab={"base_uri": "https://localhost:8080/", "height": 612}
# !pip install mplfinance yfinance
# !pip install -U git+https://github.com/twopirllc/pandas-ta
# + id="yYK61cBETWId"
# %matplotlib inline
import datetime as dt
import random as rnd
import numpy as np
import pandas as pd
import mplfinance as mpf
import pandas_ta as ta
import yfinance as yf
# + id="YA1ijAPgTO64"
class Chart(object):
def __init__(self, df: pd.DataFrame = None, strategy: ta.Strategy = ta.CommonStrategy, *args, **kwargs):
self.verbose = kwargs.pop("verbose", False)
if isinstance(df, pd.DataFrame) and df.ta.datetime_ordered:
self.df = df
if self.df.name is not None and self.df.name != "":
df_name = str(self.df.name)
else:
df_name = "DataFrame"
if self.verbose: print(f"[i] Loaded {df_name}{self.df.shape}")
else:
print(f"[X] Oops! Missing 'ohlcv' data or index is not datetime ordered.\n")
return None
self._validate_ta_strategy(strategy)
self._validate_mpf_kwargs(**kwargs)
self._validate_chart_kwargs(**kwargs)
# Build TA and Plot
self.df.ta.strategy(self.strategy, verbose=self.verbose)
self._plot(**kwargs)
def _validate_ta_strategy(self, strategy):
if strategy is not None or isinstance(strategy, ta.Strategy):
self.strategy = strategy
elif len(self.strategy_ta) > 0:
print(f"[+] Strategy: {self.strategy_name}")
else:
self.strategy = ta.CommonStrategy
def _validate_chart_kwargs(self, **kwargs):
"""Chart Settings"""
self.config = {}
self.config["last"] = kwargs.pop("last", recent_bars(self.df))
self.config["rpad"] = kwargs.pop("rpad", 10)
self.config["title"] = kwargs.pop("title", "Asset")
self.config["volume"] = kwargs.pop("volume", True)
def _validate_mpf_kwargs(self, **kwargs):
# mpf global chart settings
default_chart = mpf.available_styles()[-1]
default_mpf_width = {
'candle_linewidth': 0.6,
'candle_width': 0.525,
'volume_width': 0.525
}
mpfchart = {}
mpf_style = kwargs.pop("style", "")
if mpf_style == "" or mpf_style.lower() == "random":
mpf_styles = mpf.available_styles()
mpfchart["style"] = mpf_styles[rnd.randrange(len(mpf_styles))]
elif mpf_style.lower() in mpf.available_styles():
mpfchart["style"] = mpf_style
mpfchart["figsize"] = kwargs.pop("figsize", (12, 10))
mpfchart["non_trading"] = kwargs.pop("nontrading", False)
mpfchart["rc"] = kwargs.pop("rc", {'figure.facecolor': '#EDEDED'})
mpfchart["plot_ratios"] = kwargs.pop("plot_ratios", (12, 1.7))
mpfchart["scale_padding"] = kwargs.pop("scale_padding", {'left': 1, 'top': 4, 'right': 1, 'bottom': 1})
mpfchart["tight_layout"] = kwargs.pop("tight_layout", True)
mpfchart["type"] = kwargs.pop("type", "candle")
mpfchart["width_config"] = kwargs.pop("width_config", default_mpf_width)
mpfchart["xrotation"] = kwargs.pop("xrotation", 15)
self.mpfchart = mpfchart
def _attribution(self):
print(f"\nPandas v: {pd.__version__} [pip install pandas] https://github.com/pandas-dev/pandas")
print(f"Data from Yahoo Finance v: 1.0.19 [pip install yfinance]")
print(f"Technical Analysis with Pandas TA v: {ta.version} [pip install pandas_ta] https://github.com/twopirllc/pandas-ta")
print(f"Charts by Matplotlib Finance v: {mpf.__version__} [pip install mplfinance] https://github.com/matplotlib/mplfinance\n")
def _right_pad_df(self, rpad: int, delta_unit: str = "D", range_freq: str = "B"):
if rpad > 0:
dfpad = self.df[-rpad:].copy()
dfpad.iloc[:,:] = np.NaN
df_frequency = self.df.index.value_counts().mode()[0] # Most common frequency
freq_delta = pd.Timedelta(df_frequency, unit=delta_unit)
new_dr = pd.date_range(start=self.df.index[-1] + freq_delta, periods=rpad, freq=range_freq)
dfpad.index = new_dr # Update the padded index with new dates
self.df = self.df.append(dfpad)
def _plot(self, **kwargs):
if not isinstance(self.mpfchart["plot_ratios"], tuple):
print(f"[X] plot_ratios must be a tuple")
return
# Override Chart Title Option
chart_title = self.config["title"]
if "title" in kwargs and isinstance(kwargs["title"], str):
chart_title = kwargs.pop("title")
# Override Right Bar Padding Option
rpad = self.config["rpad"]
if "rpad" in kwargs and kwargs["rpad"] > 0:
rpad = int(kwargs["rpad"])
def cpanel():
return len(self.mpfchart['plot_ratios'])
# Last Second Default TA Indicators
linreg = kwargs.pop("linreg", False)
linreg_name = self.df.ta.linreg(append=True).name if linreg else ""
midpoint = kwargs.pop("midpoint", False)
midpoint_name = self.df.ta.midpoint(append=True).name if midpoint else ""
ohlc4 = kwargs.pop("ohlc4", False)
ohlc4_name = self.df.ta.ohlc4(append=True).name if ohlc4 else ""
clr = kwargs.pop("clr", False)
clr_name = self.df.ta.log_return(cumulative=True, append=True).name if clr else ""
rsi = kwargs.pop("rsi", False)
rsi_length = kwargs.pop("rsi_length", None)
if isinstance(rsi_length, int) and rsi_length > 1:
rsi_name = self.df.ta.rsi(length=rsi_length, append=True).name
elif rsi:
rsi_name = self.df.ta.rsi(append=True).name
else: rsi_name = ""
zscore = kwargs.pop("zscore", False)
zscore_length = kwargs.pop("zscore_length", None)
if isinstance(zscore_length, int) and zscore_length > 1:
zs_name = self.df.ta.zscore(length=zscore_length, append=True).name
elif zscore:
zs_name = self.df.ta.zscore(append=True).name
else: zs_name = ""
macd = kwargs.pop("macd", False)
macd_name = ""
if macd:
macds = self.df.ta.macd(append=True)
macd_name = macds.name
squeeze = kwargs.pop("squeeze", False)
lazybear = kwargs.pop("lazybear", False)
squeeze_name = ""
if squeeze:
squeezes = self.df.ta.squeeze(lazybear=lazybear, detailed=True, append=True)
squeeze_name = squeezes.name
ama = kwargs.pop("archermas", False)
ama_name = ""
if ama:
amas = self.df.ta.amat(append=True)
ama_name = amas.name
aobv = kwargs.pop("archerobv", False)
aobv_name = ""
if aobv:
aobvs = self.df.ta.aobv(append=True)
aobv_name = aobvs.name
treturn = kwargs.pop("trendreturn", False)
if treturn:
# Long Trend requires Series Comparison (<=. <, = >, >=)
# or Trade Logic that yields trends in binary.
default_long = self.df["SMA_10"] > self.df["SMA_20"]
long_trend = kwargs.pop("long_trend", default_long)
short_trend = ~long_trend # Opposite/Inverse
self.df["TRl"] = ta.trend_return(self.df["close"], long_trend, cumulative=True)
self.df["TRs"] = ta.trend_return(self.df["close"], short_trend, cumulative=True)
self.df["TR"] = self.df["TRl"] + self.df["TRs"]
# Pad and trim Chart
self._right_pad_df(rpad)
mpfdf = self.df.tail(self.config["last"])
mpfdf_columns = list(self.df.columns)
# BEGIN: Custom TA Plots and Panels
# Modify the area below
taplots = [] # Holds all the additional plots
# Panel 0: Price Overlay
if linreg_name in mpfdf_columns:
taplots += [mpf.make_addplot(mpfdf[linreg_name], type=kwargs.pop("linreg_type", "line"), color=kwargs.pop("linreg_color", "black"), linestyle="-.", width=1.2, panel=0)]
if midpoint_name in mpfdf_columns:
taplots += [mpf.make_addplot(mpfdf[midpoint_name], type=kwargs.pop("midpoint_type", "scatter"), color=kwargs.pop("midpoint_color", "fuchsia"), width=0.4, panel=0)]
if ohlc4_name in mpfdf_columns:
taplots += [mpf.make_addplot(mpfdf[ohlc4_name], ylabel=ohlc4_name, type=kwargs.pop("ohlc4_type", "scatter"), color=kwargs.pop("ohlc4_color", "blue"), alpha=0.85, width=0.4, panel=0)]
if self.strategy.name == ta.CommonStrategy.name:
total_sma = 0 # Check if all the overlap indicators exists before adding plots
for c in ["SMA_10", "SMA_20", "SMA_50", "SMA_200"]:
if c in mpfdf_columns: total_sma += 1
else: print(f"[X] Indicator: {c} missing!")
if total_sma == 4:
ta_smas = [
mpf.make_addplot(mpfdf["SMA_10"], color="green", width=1.5, panel=0),
mpf.make_addplot(mpfdf["SMA_20"], color="orange", width=2, panel=0),
mpf.make_addplot(mpfdf["SMA_50"], color="red", width=2, panel=0),
mpf.make_addplot(mpfdf["SMA_200"], color="maroon", width=3, panel=0),
]
taplots += ta_smas
if len(ama_name):
amat_sr_ = mpfdf[amas.columns[-1]][mpfdf[amas.columns[-1]] > 0]
amat_sr = amat_sr_.index.to_list()
else:
amat_sr = None
# Panel 1: If volume=True, the add the VOL MA. Since we know there is only one, we immediately pop it.
if self.config["volume"]:
volma = [x for x in list(self.df.columns) if x.startswith("VOL_")].pop()
max_vol = mpfdf["volume"].max()
ta_volume = [mpf.make_addplot(mpfdf[volma], color="red", width=2, panel=1, ylim=(-.2 * max_vol, 1.5 * max_vol))]
taplots += ta_volume
# Panels 2 - 9
common_plot_ratio = (3,)
if len(aobv_name):
_p = kwargs.pop("aobv_percenty", 0.2)
aobv_ylim = ta_ylim(mpfdf[aobvs.columns[0]], _p)
taplots += [
mpf.make_addplot(mpfdf[aobvs.columns[0]], ylabel=aobv_name, color="black", width=1.5, panel=cpanel(), ylim=aobv_ylim),
mpf.make_addplot(mpfdf[aobvs.columns[2]], color="silver", width=1, panel=cpanel(), ylim=aobv_ylim),
mpf.make_addplot(mpfdf[aobvs.columns[3]], color="green", width=1, panel=cpanel(), ylim=aobv_ylim),
mpf.make_addplot(mpfdf[aobvs.columns[4]], color="red", width=1.2, panel=cpanel(), ylim=aobv_ylim),
]
self.mpfchart["plot_ratios"] += common_plot_ratio # Required to add a new Panel
if clr_name in mpfdf_columns:
_p = kwargs.pop("clr_percenty", 0.1)
clr_ylim = ta_ylim(mpfdf[clr_name], _p)
taplots += [mpf.make_addplot(mpfdf[clr_name], ylabel=clr_name, color="black", width=1.5, panel=cpanel(), ylim=clr_ylim)]
if (1 - _p) * mpfdf[clr_name].min() < 0 and (1 + _p) * mpfdf[clr_name].max() > 0:
taplots += [mpf.make_addplot(mpfdf["0"], color="gray", width=1.2, panel=cpanel(), ylim=clr_ylim)]
self.mpfchart["plot_ratios"] += common_plot_ratio # Required to add a new Panel
if rsi_name in mpfdf_columns:
rsi_ylim = (0, 100)
taplots += [
mpf.make_addplot(mpfdf[rsi_name], ylabel=rsi_name, color=kwargs.pop("rsi_color", "black"), width=1.5, panel=cpanel(), ylim=rsi_ylim),
mpf.make_addplot(mpfdf["20"], color="green", width=1, panel=cpanel(), ylim=rsi_ylim),
mpf.make_addplot(mpfdf["50"], color="gray", width=0.8, panel=cpanel(), ylim=rsi_ylim),
mpf.make_addplot(mpfdf["80"], color="red", width=1, panel=cpanel(), ylim=rsi_ylim),
]
self.mpfchart["plot_ratios"] += common_plot_ratio # Required to add a new Panel
if macd_name in mpfdf_columns:
_p = kwargs.pop("macd_percenty", 0.15)
macd_ylim = ta_ylim(mpfdf[macd_name], _p)
taplots += [
mpf.make_addplot(mpfdf[macd_name], ylabel=macd_name, color="black", width=1.5, panel=cpanel()),#, ylim=macd_ylim),
mpf.make_addplot(mpfdf[macds.columns[-1]], color="blue", width=1.1, panel=cpanel()),#, ylim=macd_ylim),
mpf.make_addplot(mpfdf[macds.columns[1]], type="bar", alpha=0.8, color="dimgray", width=0.8, panel=cpanel()),#, ylim=macd_ylim),
mpf.make_addplot(mpfdf["0"], color="black", width=1.2, panel=cpanel()),#, ylim=macd_ylim),
]
self.mpfchart["plot_ratios"] += common_plot_ratio # Required to add a new Panel
if zs_name in mpfdf_columns:
_p = kwargs.pop("zascore_percenty", 0.2)
zs_ylim = ta_ylim(mpfdf[zs_name], _p)
taplots += [
mpf.make_addplot(mpfdf[zs_name], ylabel=zs_name, color="black", width=1.5, panel=cpanel(), ylim=zs_ylim),
mpf.make_addplot(mpfdf["-3"], color="red", width=1.2, panel=cpanel(), ylim=zs_ylim),
mpf.make_addplot(mpfdf["-2"], color="orange", width=1, panel=cpanel(), ylim=zs_ylim),
mpf.make_addplot(mpfdf["-1"], color="silver", width=1, panel=cpanel(), ylim=zs_ylim),
mpf.make_addplot(mpfdf["0"], color="black", width=1.2, panel=cpanel(), ylim=zs_ylim),
mpf.make_addplot(mpfdf["1"], color="silver", width=1, panel=cpanel(), ylim=zs_ylim),
mpf.make_addplot(mpfdf["2"], color="orange", width=1, panel=cpanel(), ylim=zs_ylim),
mpf.make_addplot(mpfdf["3"], color="red", width=1.2, panel=cpanel(), ylim=zs_ylim)
]
self.mpfchart["plot_ratios"] += common_plot_ratio # Required to add a new Panel
if squeeze_name in mpfdf_columns:
_p = kwargs.pop("squeeze_percenty", 0.6)
sqz_ylim = ta_ylim(mpfdf[squeeze_name], _p)
taplots += [
mpf.make_addplot(mpfdf[squeezes.columns[-4]], type="bar", color="lime", alpha=0.65, width=0.8, panel=cpanel(), ylim=sqz_ylim),
mpf.make_addplot(mpfdf[squeezes.columns[-3]], type="bar", color="green", alpha=0.65, width=0.8, panel=cpanel(), ylim=sqz_ylim),
mpf.make_addplot(mpfdf[squeezes.columns[-2]], type="bar", color="maroon", alpha=0.65, width=0.8, panel=cpanel(), ylim=sqz_ylim),
mpf.make_addplot(mpfdf[squeezes.columns[-1]], type="bar", color="red", alpha=0.65, width=0.8, panel=cpanel(), ylim=sqz_ylim),
mpf.make_addplot(mpfdf["0"], color="black", width=1.2, panel=cpanel(), ylim=sqz_ylim),
mpf.make_addplot(mpfdf[squeezes.columns[4]], ylabel=squeeze_name, color="green", width=2, panel=cpanel(), ylim=sqz_ylim),
mpf.make_addplot(mpfdf[squeezes.columns[5]], color="red", width=1.8, panel=cpanel(), ylim=sqz_ylim),
]
self.mpfchart["plot_ratios"] += common_plot_ratio # Required to add a new Panel
if treturn:
_p = kwargs.pop("treturn_percenty", 0.33)
treturn_ylim = ta_ylim(mpfdf["TR"], _p)
taplots += [
mpf.make_addplot(mpfdf["TRl"], ylabel="Trend Return", type="bar", color="green", alpha=0.45, width=0.8, panel=cpanel(), ylim=treturn_ylim),
mpf.make_addplot(mpfdf["TRs"], type="bar", color="red", alpha=0.45, width=0.8, panel=cpanel(), ylim=treturn_ylim),
mpf.make_addplot(mpfdf["TR"], color="black", width=1.5, panel=cpanel(), ylim=treturn_ylim),
mpf.make_addplot(mpfdf["0"], color="black", width=1.2, panel=cpanel(), ylim=treturn_ylim),
]
self.mpfchart["plot_ratios"] += common_plot_ratio # Required to add a new Panel
_p = kwargs.pop("cstreturn_percenty", 0.33)
trcs = mpfdf["TR"].cumsum()
treturncs_ylim = ta_ylim(trcs, _p)
taplots += [
mpf.make_addplot(trcs, ylabel="Trend B&H", color="black", width=1.5, panel=cpanel(), ylim=treturncs_ylim),
mpf.make_addplot(mpfdf["0"], color="black", width=1.2, panel=cpanel(), ylim=treturncs_ylim),
]
self.mpfchart["plot_ratios"] += common_plot_ratio # Required to add a new Panel
# END: Custom TA Plots and Panels
if self.verbose:
additional_ta = []
chart_title = f"{chart_title} [{self.strategy.name}] (last {self.config['last']} bars)"
chart_title += f"\nSince {mpfdf.index[0]} till {mpfdf.index[-1]}"
if len(linreg_name) > 0: additional_ta.append(linreg_name)
if len(midpoint_name) > 0: additional_ta.append(midpoint_name)
if len(ohlc4_name) > 0: additional_ta.append(ohlc4_name)
if len(additional_ta) > 0:
chart_title += f"\nIncluding: {', '.join(additional_ta)}"
if amat_sr:
vlines_ = dict(vlines=amat_sr, alpha=0.1, colors="red")
else:
# Hidden because vlines needs valid arguments even if None
vlines_ = dict(vlines=mpfdf.index[0], alpha=0, colors="white")
# Create Final Plot
mpf.plot(mpfdf,
title=chart_title,
type=self.mpfchart["type"],
style=self.mpfchart["style"],
datetime_format="%-m/%-d/%Y",
volume=self.config["volume"],
figsize=self.mpfchart["figsize"],
tight_layout=self.mpfchart["tight_layout"],
scale_padding=self.mpfchart["scale_padding"],
panel_ratios=self.mpfchart["plot_ratios"], # This key needs to be update above if adding more panels
xrotation=self.mpfchart["xrotation"],
update_width_config=self.mpfchart["width_config"],
show_nontrading=self.mpfchart["non_trading"],
vlines=vlines_,
addplot=taplots
)
self._attribution()
# + id="ppCoDeXoTWGj" outputId="f299393d-08d1-4825-bd7b-4b369a8962e4" colab={"base_uri": "https://localhost:8080/", "height": 252}
def get_prices_for_stock(stock, start_date="2019-03-01", end_date="2020-05-30"):
data = yf.download(stock, start=start_date, end=end_date)
data = data.fillna(method='ffill')
# Drop columns with no entries
data = data.dropna(axis='columns', how='all')
return data
def recent_bars(df, tf: str = "1y"):
# All Data: 0, Last Four Years: 0.25, Last Two Years: 0.5, This Year: 1, Last Half Year: 2, Last Quarter: 4
yearly_divisor = {"all": 0, "10y": 0.1, "5y": 0.2, "4y": 0.25, "3y": 1./3, "2y": 0.5, "1y": 1, "6mo": 2, "3mo": 4}
yd = yearly_divisor[tf] if tf in yearly_divisor.keys() else 0
return int(ta.RATE["TRADING_DAYS_PER_YEAR"] / yd) if yd > 0 else df.shape[0]
def ta_ylim(series: pd.Series, percent: float = 0.1):
smin, smax = series.min(), series.max()
if isinstance(percent, float) and 0 <= float(percent) <= 1:
y_min = (1 + percent) * smin if smin < 0 else (1 - percent) * smin
y_max = (1 - percent) * smax if smax < 0 else (1 + percent) * smax
return (y_min, y_max)
return (smin, smax)
ntar_df = get_prices_for_stock('NTAR.CN')
# Used for example Trend Return Long Trend Below
ntar_df.columns = ['open', 'high', 'low', 'close', 'adj_close', 'volume']
ntar_df.head()
# + id="8AwDv8i_WGN7" outputId="234202e1-f747-4c61-aeb3-fdb496dd002a" colab={"base_uri": "https://localhost:8080/", "height": 102}
# Calculate Returns and append to the df DataFrame
ntar_df.ta.log_return(cumulative=True, append=True)
ntar_df.ta.percent_return(cumulative=True, append=True)
ntar_df.ta.rsi(cumulative=False, append=True)
ntar_df.head()
ntar_df.columns
# + id="f2ptWp8ig3UT" outputId="af1d3d01-1928-472c-e731-8b2808909b0c" colab={"base_uri": "https://localhost:8080/", "height": 590}
# We create an additional plot placing it on the third panel
roc_plot = mpf.make_addplot(ntar_df['RSI_14'], panel=2, ylabel='ROC')
#We pass the additional plot using the addplot parameter
mpf.plot(ntar_df, type='candle', style='yahoo', addplot=roc_plot, title="Ntar Price Point", volume=True)
# + id="MT_B32j4g3Rt" outputId="9a780b38-0b3f-4d9c-856a-f308cd853608" colab={"base_uri": "https://localhost:8080/", "height": 630}
# IP analysis just before major increases in stock prices
ip_spike1_df = get_prices_for_stock('IP.CN', '2020-05-04', '2020-07-25')
ip_spike1_df.tail()
ip_spike1_df.ta.log_return(cumulative=True, append=True)
ip_spike1_df.ta.percent_return(cumulative=True, append=True)
ip_spike1_df.ta.rsi(cumulative=False, append=True)
ip_spike1_df.ta.pvol(append=True)
print(ip_spike1_df.columns)
roc_plot = mpf.make_addplot(ip_spike1_df['PVOL'], panel=2, ylabel='ROC')
#We pass the additional plot using the addplot parameter
mpf.plot(ip_spike1_df, type='candle', addplot=roc_plot, title="IP Price Point", volume=True)
# + id="QrPIoYjihND3"
# Cielo Analysis just before major stock increases
| ibook/notebooks/analyze/pandas_ta_plots_ip.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib inline
df1 = pd.read_csv('df1',index_col=0)
df1.head()
df2 = pd.read_csv('df2')
df2.head()
df1['A'].hist(bins=50)
df1.plot(figsize=(10,10))
df1['A'].plot(kind='hist',bins=30)
df1['A'].plot.hist()
df2.head()
df2.plot.area()
df2.plot.bar()
df2.plot.bar(stacked=True)
df1.head()
df1.plot.scatter(x='A',y='B',c='C',cmap='coolwarm')
df1.plot.scatter(x='A',y='B',s=df1['C']*100)
df2.plot.box()
df=pd.DataFrame(np.random.randn(1000,2),columns=['a','b'])
df
df.plot.hexbin(x='a',y='b',gridsize=25,cmap='coolwarm')
df2['a'].plot.density()
df2.plot.density()
| Pandas data viz.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pedroblossbraga/MachineLearning-Pre-Processing-with-Python/blob/master/Effects_of_transformations_in_XGBoostRegressor_example.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="dZQgRSayjhWt"
# ## Tests with different data transformations applied to XGBoost
#
# Samples:
# - original data
# - linearly transformated data
# - minmax scaled data
# - pseudo-random data
# + id="Q-VOWFB-j1BD"
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.impute import SimpleImputer
from xgboost import XGBRegressor
from sklearn.metrics import mean_absolute_error
# + id="VlNDJZbUvd9u"
from sklearn.metrics import mean_squared_error
# + id="pkJHjuRbmfox"
import matplotlib.pyplot as plt
import seaborn as sns
import statistics
from IPython.display import display
# + id="7JmxzeYcnWGj"
import warnings
warnings.filterwarnings("ignore")
# + id="YxgnmJ8kj24c"
from sklearn.datasets import load_boston
X, y = load_boston(return_X_y=True)
# + id="4g5PEgYTkRYJ"
train_X, test_X, train_y, test_y = train_test_split(X, y, test_size=0.2)
# + id="f66pvSSLkaOL"
# lidar com os valores ausentes
df_imputer = SimpleImputer()
train_X = df_imputer.fit_transform(train_X)
test_X = df_imputer.transform(test_X)
# + id="if3BPWCvjfU-"
def test_XGBoost(train_X, train_y, test_X, test_y, plot_residuals=True):
# instanciar o modelo XGBoost
model = XGBRegressor()
# chamar o fit para o modelo
model.fit(train_X, train_y, verbose=False)
# fazer previsões em cima do dataset de teste
predictions = model.predict(test_X)
print("Mean Absolute Error: {:.2f}".format(mean_absolute_error(predictions, test_y)))
if plot_residuals:
plt.figure(figsize=(15,3))
plt.subplot(1,2,1)
plt.title("predictions")
sns.distplot(predictions)
plt.axvline(statistics.mean(predictions), color='red')
plt.subplot(1,2,2)
plt.title(r'residuals $\epsilon = |\hat{y} - y|$')
sns.distplot(abs(predictions-test_y))
plt.axvline(statistics.mean(abs(predictions-test_y)), color='red')
plt.show()
return mean_absolute_error(y_pred=predictions, y_true=test_y), mean_squared_error(y_true=test_y, y_pred=predictions)
# + [markdown] id="31CWDEDJuskV"
# ### original data
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="1Kl0zOFhub4Z" outputId="08e1d661-5f12-435c-8f95-aa2941c64cd9"
mae0, mse0 = test_XGBoost(train_X, train_y, test_X, test_y)
# + [markdown] id="dEWLYelhuufc"
# ### linearly transformed
#
# \begin{equation}
# X_1 = \{x_j + k\}_{j=1}^N, k \in \mathbb{N}
# \end{equation}
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="8T5-k3n7lDTb" outputId="ebdb99f6-a80a-44ec-ba98-e3d77789b595"
k=20
mae1, mse1 = test_XGBoost(train_X+k, train_y+k, test_X+k, test_y+k)
# + [markdown] id="X09WSAedvDN7"
# ### Min-Max scaled
#
# \begin{equation}
# \hat{X} = \left\{ \frac{x_j - min \{ X \} }{max \{ X \} - min \{ X \} } \right\}_{j=1}^N
# \end{equation}
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="D9whU9n2lSwN" outputId="c6db5ce1-5c40-4009-c843-a19cf7610012"
def minmaxscale(v):
return (v - v.min(axis=0))/(v.max(axis=0)-v.min(axis=0))
mae2, mse2 = test_XGBoost(
minmaxscale(train_X),
minmaxscale(train_y),
minmaxscale(test_X),
minmaxscale(test_y)
)
# + id="VW4ccwqZmNKU"
import numpy as np
def randomize_matrix(X):
X_ = X.copy()
if len(X_.shape)==1: # vector
for i in range(X_.shape[0]):
X_[i] = np.random.randint(-20, 20)
else: # matrix
for lin in range(X_.shape[0]):
for col in range(X_.shape[1]):
X_[lin][col] = np.random.randint(-20, 20)
return X_
# + [markdown] id="EN3wretfvprk"
# ### Pseudo-random data
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="10u3ktuQsLYg" outputId="d22ed60d-5134-46d2-e8be-6a2b8b88f4e1"
mae3, mse3 = test_XGBoost(
randomize_matrix(train_X),
randomize_matrix(train_y),
randomize_matrix(test_X),
randomize_matrix(test_y)
)
# + colab={"base_uri": "https://localhost:8080/", "height": 173} id="XOXQv236tI-W" outputId="e4a6935c-6e5e-4098-d341-7a8b7df0175f"
erros = {
'MAE': [mae0, mae1, mae2, mae3],
'MSE': [mse0, mse1, mse2, mse3],
'transf': ['original', 'linear', 'minmax', 'pseudo-random']
}
display(pd.DataFrame(erros))
| Effects_of_transformations_in_XGBoostRegressor_example.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp boostmonodepth_utils
# -
# # boostmonodepth_utils
# > Default description (change me)
# +
#export
import glob
import os
import cv2
import imageio
import numpy as np
from pomerantz.MiDaS.MiDaS_utils import write_depth
# -
#export
BOOST_BASE = 'BoostingMonocularDepth'
BOOST_INPUTS = 'inputs'
BOOST_OUTPUTS = 'outputs'
#export
def run_boostmonodepth(img_names, src_folder, depth_folder):
if not isinstance(img_names, list):
img_names = [img_names]
# remove irrelevant files first
clean_folder(os.path.join(BOOST_BASE, BOOST_INPUTS))
clean_folder(os.path.join(BOOST_BASE, BOOST_OUTPUTS))
tgt_names = []
for img_name in img_names:
base_name = os.path.basename(img_name)
tgt_name = os.path.join(BOOST_BASE, BOOST_INPUTS, base_name)
os.system(f'cp {img_name} {tgt_name}')
# keep only the file name here.
# they save all depth as .png file
tgt_names.append(os.path.basename(tgt_name).replace('.jpg', '.png'))
os.system(f'cd {BOOST_BASE} && python run.py --Final --data_dir {BOOST_INPUTS}/ --output_dir {BOOST_OUTPUTS} --depthNet 0')
for i, (img_name, tgt_name) in enumerate(zip(img_names, tgt_names)):
img = imageio.imread(img_name)
H, W = img.shape[:2]
scale = 640. / max(H, W)
# resize and save depth
target_height, target_width = int(round(H * scale)), int(round(W * scale))
depth = imageio.imread(os.path.join(BOOST_BASE, BOOST_OUTPUTS, tgt_name))
depth = np.array(depth).astype(np.float32)
depth = resize_depth(depth, target_width, target_height)
np.save(os.path.join(depth_folder, tgt_name.replace('.png', '.npy')), depth / 32768. - 1.)
write_depth(os.path.join(depth_folder, tgt_name.replace('.png', '')), depth)
#export
def clean_folder(folder, img_exts=['.png', '.jpg', '.npy']):
for img_ext in img_exts:
paths_to_check = os.path.join(folder, f'*{img_ext}')
if len(glob.glob(paths_to_check)) == 0:
continue
print(paths_to_check)
os.system(f'rm {paths_to_check}')
#export
def resize_depth(depth, width, height):
"""Resize numpy (or image read by imageio) depth map
Args:
depth (numpy): depth
width (int): image width
height (int): image height
Returns:
array: processed depth
"""
depth = cv2.blur(depth, (3, 3))
return cv2.resize(depth, (width, height), interpolation=cv2.INTER_AREA)
| nbs/01_boostmonodepth_utils.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import numpy as np
import pandas as pd
res_path = './res'
feat_path = './features'
train_path = './data/train'
labels = os.listdir(train_path)
linux_test_order = np.argsort(np.load('./linux_order_test.npy'))
linux_label_order = np.argsort(np.load('./linux_order_label.npy'))
model_names = ['densenet', 'resnest50', 'resnet50', 'cnn_base', 'lstm_base']
res = []
for model_name in model_names:
if model_name in ['resnest50', 'resnet50']:
for i in range(5):
cur = np.load(os.path.join(res_path, "{}_fold_{}.npy".format(model_name, i)))
res.append(cur[57886:][linux_test_order][:, linux_label_order])
else:
for i in range(5):
res.append(np.load(os.path.join(res_path, "{}_fold_{}.npy".format(model_name, i)))[57886:])
test_names = np.load(os.path.join(feat_path, 'image_128_256.npz'))['test_names']
sub = pd.DataFrame()
sub['file_name'] = test_names
sub['label'] = np.argmax(np.mean(res, axis=0), axis=1)
sub['label'] = sub['label'].map({i:label for i, label in enumerate(labels)})
sub
import time
now = time.strftime("%Y%m%d_%H%M%S",time.localtime(time.time()))
fname="submit_ensemble_" + now + ".csv"
sub.to_csv(fname, index=False)
| ccf_audio/Ensemble Result.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="A3K1aHn-3BdD"
# Imports all in one place
import pandas as pd
import numpy as np
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error, r2_score
from sklearn.model_selection import train_test_split
# + [markdown] colab_type="text" id="n1o9Qe8ilN19" endofcell="--"
# # "Linear" Regression
#
# Which of the following is a linear regression model?
#
# 
#
# -
#
# -
#
# -
#
# -
#
# -
#
#
# **All** of these functional forms can be fit using Linear Regression. The "Linear" in linear regression refers to the linear form of the equation.
# --
# + [markdown] colab_type="text" id="4a5raiR9doDx"
# ## Linear Combinations
#
# Remember when we rewrote vectors as a **linear combination** of scalars and unit vectors?
#
# \begin{align}
# v = \begin{bmatrix}2 \\ 3\end{bmatrix} = 2 \begin{bmatrix} 1 \\ 0 \end{bmatrix} + 3 \begin{bmatrix} 0 \\ 1\end{bmatrix} = 2\hat{i} + 3\hat{j}
# \end{align}
#
# The syntax where we have a scalar (think coefficient) multiplying some vector (unit vector in this case) and all of them being added together is what makes this a linear combination.
# + [markdown] colab_type="text" id="gu-5ZGkFdpCS"
# ## Linear Equations
#
# A "Linear Equation" is any equation that takes the following form:
#
# \begin{align}
# a_1x_1 + \ldots + a_nx_n + b = 0
# \end{align}
#
# Does this look familiar? A linear equation is one where we have $x_1, \ldots, x_n$ unknowns and $b, a_1, \ldots, a_n$ coefficients which are considered parameters of the equation. "The solutions of such an equation are the values that, when substituted to the unknowns, make the equality true."
#
# [Linear Equation Wikipedia](https://en.wikipedia.org/wiki/Linear_equation)
#
# Linear Regression is **linear** not because it can only plot straight lines and fit straight-line patterns in data, but because the form of the equation used to represent our regression is in the form of a **Linear Equation**.
# + [markdown] colab_type="text" id="yUMyYQSM_w-9"
# ## Well how do we fit curved data then?
# + [markdown] colab_type="text" id="_cs8PskIdy9t"
# # Polynomial Regression
#
# Just as multiple regression was an extension of the bivariate case, Polynomial Regression is an extention of multiple regression and can be used to fit data to any (curved) shape. This is one of the reasons why data exploration is so important. You won't know that you need to fit a polynomial function to a feature unless you have examined its distribution.
#
# [Why is polynomial regression considered a special case of multiple linear regression?](https://stats.stackexchange.com/questions/92065/why-is-polynomial-regression-considered-a-special-case-of-multiple-linear-regres)
# + [markdown] colab_type="text" id="ptmg8FRty5Pu"
# ## Lets look at an example!
# + colab={"base_uri": "https://localhost:8080/", "height": 249} colab_type="code" id="9dEQTcwHy8mY" outputId="7928f778-38e2-4a74-f2df-53e32cc4764e"
df = pd.read_csv('https://raw.githubusercontent.com/ryanleeallred/datasets/master/kc_house_data.csv')
pd.set_option('display.max_columns', 100)
print(df.shape)
print("Does this dataset look familiar?")
df.head()
# + [markdown] colab_type="text" id="MCwbKGIOzqbO"
# ## Find a "curved" feature in the dataset
# + colab={"base_uri": "https://localhost:8080/", "height": 226} colab_type="code" id="wTYcFh3ZzQsS" outputId="4af96e7b-ba87-452b-e8c7-4b768bcfcf7d"
sns.set(style="ticks", color_codes=True)
# Generate a list of column headers not including price
x_columns = df.columns.drop('price')
# Only plot the scatterplot of x variables with our y variable
sns.pairplot(data=df, y_vars=['price'], x_vars=x_columns)
# + [markdown] colab_type="text" id="H6X-TDuL2PaG"
# ## Generate or "engineer" a new grade_squared feature
# + colab={"base_uri": "https://localhost:8080/", "height": 215} colab_type="code" id="lKiJSmUn2Ohr" outputId="c8a41385-5613-403c-e193-cf91d31febf3"
df['grade_squared'] = df['grade']**2
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 226} colab_type="code" id="DQ_yEHOtids9" outputId="3ef80f1d-a253-4739-b07f-953138e22c14"
sns.set(style="ticks", color_codes=True)
# Generate a list of column headers not including price
x_columns = df.columns.drop('price')
# Only plot the scatterplot of x variables with our y variable
sns.pairplot(data=df, y_vars=['price'], x_vars=x_columns)
# + [markdown] colab_type="text" id="PTdqhfQY1hLb"
# ## Test the fit of a polynomial regression to that feature
#
# First we'll fit a regular bivariate regression line and calculate its $R^2$ to get a baseline. Since we want to know if this generated feature is improving our model or not we'll first run our code without it so that we have something to compare to.
#
# $price_i = \beta_0 + \beta_1grade_i + \epsilon_i$
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="l5eFe55l2JHT" outputId="578adb74-904b-49c7-ddd2-d7c1600f637f"
# Separate dependent and independent variables
y = df.loc[:, ['price']].values
X = df.loc[:, ['grade']].values
# Slit into test and train datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=.2, random_state=42)
# fit model using train datasets
model = LinearRegression()
model.fit(X_train, Y_train)
# Create new predictions using x_test
y_pred = model.predict(X_test)
# Measure Accuracy using y_test and y_pred
RMSE = (np.sqrt(mean_squared_error(Y_test, y_pred)))
R2 = r2_score(Y_test, y_pred)
print('RMSE is {}'.format(RMSE))
print('R^2 is {}'.format(R2))
# + [markdown] colab_type="text" id="n4dAZb0U4-vE"
# ## Lets try fitting grade_squared as a bivariate model
#
# $price_i = \beta_0 + \beta_1grade^{2}_i + \epsilon_i$
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="Nfe7mTsm4-Bv" outputId="2682e202-bfda-4c81-bded-3dff017e592a"
# Separate dependent and independent variables
y = df.loc[:, ['price']].values
X = df.loc[:, ['grade_squared']].values
# Slit into test and train datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=.5, random_state=42)
# fit model using train datasets
model = LinearRegression()
model.fit(X_train, Y_train)
# Create new predictions using x_test
y_pred = model.predict(X_test)
# Measure Accuracy using y_test and y_pred
RMSE = (np.sqrt(mean_squared_error(Y_test, y_pred)))
R2 = r2_score(Y_test, y_pred)
print('RMSE is {}'.format(RMSE))
print('R^2 is {}'.format(R2))
# + [markdown] colab_type="text" id="9lSw_KYF5wSB"
# ## Multiple Regression using both grade and grade_squred
#
# $price_i = \beta_0 + \beta_1 grade_i + \beta_2grade^{2}_i + \epsilon_i$
# + colab={"base_uri": "https://localhost:8080/", "height": 67} colab_type="code" id="avqSfsJr2h2I" outputId="0d932f23-6c1b-417f-e7ee-8372b4d2561e"
# Separate dependent and independent variables
y = df.loc[:, ['price']].values
X = df.loc[:, ['grade', 'grade_squared']].values
# Slit into test and train datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=.5, random_state=42)
# fit model using train datasets
model = LinearRegression()
model.fit(X_train, Y_train)
print(model.coef_)
# Create new predictions using x_test
y_pred = model.predict(X_test)
# Measure Accuracy using y_test and y_pred
RMSE = (np.sqrt(mean_squared_error(Y_test, y_pred)))
R2 = r2_score(Y_test, y_pred)
print('RMSE is {}'.format(RMSE))
print('R^2 is {}'.format(R2))
# + [markdown] colab_type="text" id="loZ378cZ7mIC"
# # How to find non-linear features
# + [markdown] colab_type="text" id="fM6fKuubBy8b"
# ## 1) Domain Knowledge (think about your variables and hypothesize)
#
# This is why having domain knowledge about the problem that you're trying to solve is something that's so important. In the context of home prices, variables that have a curved structure often are that way due to some form of diminishing returns increases in certain amenities. Lets think about the following variables:
#
# - Lot Size:
#
# The more land you're willing to buy all at once the cheaper it will be on a per-acre basis (Saving money when you buy in bulk). This trend carries through to small to medium sized lots but but with a more shallow curve.
#
# 
#
# - Square Footage:
#
# Square footage of a home sees a similar pattern. The value an additional 100 square feet in small homes (imagine the difference between say a 800 sq foot home and a 900 sq foot home) makes a big difference to buyers, where as an additional 100 square feet in a mansion probably isn't valued quite as highly.
#
# - Age:
#
# Just like how the prices of new cars drop steeply in the first few years, the value of homes due to age drop quickly in the first few years after its built and then less quickly as time goes on. This is not a linear pattern and needs to be fitted by a polynomial model.
# + [markdown] colab_type="text" id="f5cBlL1xBzFw"
# ## 2) Visual Inspection
#
# We already talked about how generating scatterplots or other graphs is vital in our data exploration stage and can lead us to identify possible candidates for polynomials. Here I just wanted to share one more tip that can help you analyze scatterplots when you have a lot of data.
#
# If you have so much data that it's hard to tell what's going on in your scatterplot, then sample your data and regenerate them to get a better idea.
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" id="udIeJAY-B94U" outputId="9cec3697-7aa8-459f-8e39-7e8c2e2cc513"
sns.set(style="ticks", color_codes=True)
# Generate a list of column headers not including price
x_columns = df.columns.drop('price')
# Sample our dataframe to take 1/20th the values
sampled = df.sample(frac=0.05, replace=True, random_state=42)
# print(x_columns)
# Only plot the scatterplot of x variables with our y variable
sns.pairplot(data=sampled, y_vars=['price'], x_vars=x_columns)
# + [markdown] colab_type="text" id="yDyTStZgBzIU"
# ## 3) Inspect the distribution of residuals
#
# 
# + [markdown] colab_type="text" id="SRF9aISzGRyr"
# ## An Aside: The "Hedonic Housing Model"
#
# Using Linear Regression to model home prices is a very common use of predictive linear regression modeling. It's so common fact that it has its own name: The Hedonic Housing Model. In the Hedonic Housing model it is well understood and reiterated that certain features tend to be curved in nature and these polynomial features (like the ones mentioned above) have all been well explored in real estate prediction circles. This is another of how domain knowledge can give you an edge. The best way to gain domain knowledge is to dive in and try and solve one particular kind of problem, and pick up little tips and tidbits as time goes on.
# + [markdown] colab_type="text" id="U1xiwKaUP0Y_"
# # Log-Linear Regression
#
# In a log-linear regression model, we take the natural log of all of our y variable and use that as our y vector instead of the raw y values. Why would we do that?
#
# <https://www.kaggle.com/c/house-prices-advanced-regression-techniques/discussion/60861>
# + [markdown] colab_type="text" id="4kVJde3zZrms"
# ## 1) To reduce skew in y
#
# Where we have variables with lots of relatively low values and few high values (like with home prices) we would expect to see our data more clustered on the left-hand side with a long tail extending to the right up into the expensive homes. The fact of the matter is that we will be able to make better predictions if we can normalize our data to some degree and one way to do this is by taking the natural log of it.
# + colab={} colab_type="code" id="rdS22S_OQ-vr"
## generate our ln(price) variable
df['ln_price'] = np.log(df['price'])
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" id="YqmZNtUOTZoN" outputId="6125f6ed-2df1-404e-d977-32135f41bf00"
df.price.hist()
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" id="p7CDc9_ETeUf" outputId="cd9f3f3f-9936-4e2d-c3fb-2482ca04ddc9"
df.ln_price.hist()
# + [markdown] colab_type="text" id="qXrEJWb8akIL"
# ## New distribution of variables
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" id="whrf7MbDYisJ" outputId="beffe32c-c216-46fd-97f9-cd9619ad3955"
sns.set(style="ticks", color_codes=True)
# Generate a list of column headers not including price
x_columns = df.columns.drop('ln_price')
# Only plot the scatterplot of x variables with our y variable
sns.pairplot(data=df, y_vars=['ln_price'], x_vars=x_columns,
plot_kws={'alpha': 0.1, 'linewidth':0})
# + [markdown] colab_type="text" id="sd4mICl9ZrvR"
# ## 2) Make coefficients easier to interpret
#
# Transforming our price values in this way won't change our model's ability to generate predictions, but what it **will** do is change the interpretation of all of our x-coefficients. This will change our x-coefficients from having an elasticity type interpretation (a raw dollar amount change if there is a 1 unit increase in x) to having a percentage-terms interpretation. Lets demonstrate and talk about this further.
# + [markdown] colab_type="text" id="T-Yt7EcZZr33"
# ## 3) Make our errors easier to interpret
#
# Errors that have been calculated on variables that are in log form can also be interpreted roughly as percentage error. We've been using percentages all our lives and they have immediate meaning to us. This is why I prefer log-linear regression models when possible.
#
# <https://people.duke.edu/~rnau/411log.htm>
# + [markdown] colab_type="text" id="f1ecoaG9K-FB"
# Feature Engineering is a big topic in machine learning. We won't be able to cover every aspect of it today, but hopefully we can give you a strong idea of what it is and how to go about it.
#
# [Understanding Feature Engineering Part 1](https://towardsdatascience.com/understanding-feature-engineering-part-1-continuous-numeric-data-da4e47099a7b)
#
# Feature engineering is key to success in predictive modeling. It is the process by which we take existing features and combine them or alter them in ways that will expose additional signal to our model. Feature engineering is all about making the most of the data that we already had.
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" id="gLC6Zz1pQaef" outputId="11b76865-0d19-4617-8047-6c114f809f0a"
## Log-Linear Regression
# Separate dependent and independent variables
y = df.loc[:, ['ln_price']].values
X = df.loc[:, ['grade']].values
# Slit into test and train datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=.5, random_state=42)
# fit model using train datasets
model = LinearRegression()
model.fit(X_train, Y_train)
# Create new predictions using x_test
y_pred = model.predict(X_test)
# Measure Accuracy using y_test and y_pred
RMSE = (np.sqrt(mean_squared_error(Y_test, y_pred)))
R2 = r2_score(Y_test, y_pred)
print('RMSE is {}'.format(RMSE))
print('R^2 is {}'.format(R2))
print("coefficients: ", model.coef_[0])
print("intercepts: ", model.intercept_)
# + [markdown] colab_type="text" id="YwlCapGPUwFe"
# This means that a one unit increase in the grade of a home increases its sale price by 31.3%. Often it is much easier to interpret coefficients in this manner than in the regular way.
# + colab={"base_uri": "https://localhost:8080/", "height": 84} colab_type="code" id="ZVNHMgpuXo5z" outputId="460e0b13-54ae-4ae9-82d3-09078d3007ee"
## Log-Linear Regression
# Separate dependent and independent variables
y = df.loc[:, ['ln_price']].values
X = df.loc[:, ['grade', 'grade_squared']].values
# Slit into test and train datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=.5, random_state=42)
# fit model using train datasets
model = LinearRegression()
model.fit(X_train, Y_train)
# Create new predictions using x_test
y_pred = model.predict(X_test)
# Measure Accuracy using y_test and y_pred
RMSE = (np.sqrt(mean_squared_error(Y_test, y_pred)))
R2 = r2_score(Y_test, y_pred)
print('RMSE is {}'.format(RMSE))
print('R^2 is {}'.format(R2))
print("coefficients: ", model.coef_[0])
print("intercepts: ", model.intercept_)
# + [markdown] colab_type="text" id="QCdxF6o1Rowb"
# Our RMSE is really small now because it now represents error in percentage terms. WE're on agerage about 37.5% off in our predictions of house prices.
#
# Our coefficients can also be understood in percentage terms which makes the coefficients on our regression much more digestable at a glance.
# + colab={"base_uri": "https://localhost:8080/", "height": 376} colab_type="code" id="dbuqJRvhVWAX" outputId="b82dc3eb-472f-40ce-9c96-e7da2076bbca"
df['grade'].hist()
# + [markdown] colab_type="text" id="36wffNrwZJ6V"
# # Feature Engineering
# + [markdown] colab_type="text" id="4B98U_YoMBTY"
# ## Polynomial Regression requires feature engineering
#
# You've already seen an example of feature engineering today when we created the $grade^2$ variable. We took an existing feature and used it to generate a new feature that exposed the data to the model in a slightly different way.
#
# "Feature engineering is the process of transforming raw data into features that better represent the underlying problem to the predictive models, resulting in improved model accuracy on unseen data." - [<NAME>](https://machinelearningmastery.com/discover-feature-engineering-how-to-engineer-features-and-how-to-get-good-at-it/)
#
# What is some potential feature engineering that we could do on the King County dataset?
# + colab={"base_uri": "https://localhost:8080/", "height": 222} colab_type="code" id="WKs8ww9yj3e3" outputId="9021a38b-cf80-4870-d9d0-a2d2e29ca2cc"
df.head()
# + [markdown] colab_type="text" id="Pk15gYRxMw9_"
# - **[date]** The date is in a format that is not super useful to us. If were to extract the year we could then take the difference between year and yr_built to find the age of the home. We could also include the squared term of the home age since we know (from our hedonic housing model domain knowledge) that home age typically is not linear.
# - **[bedrooms]** & **[bathrooms]** We maybe use a combined measure of bedrooms and bathrooms, or find an average room square footage by taking total number of rooms and dividing by the square footage.
# - **[sqft_living]** **[sqft_lot]** The difference between lot square footage and home square footage ought to also give us a rough measure of the size of the yard. Rough measures are fine as long as the engineered features expose some new shred of meaning to our model.
# - **[floors]** We could calculate an average number of square feet per floor
# - **[lat]** **[long]** There are all kinds of things that we could do with the latitude and longitude especially if we use some kind of outside API or external dataset to bring in new features associated with the location of the homes. This would take a lot of work but these could potentially be very powerful features.
#
# ## Kaggle is one of the best places to get feature engineering ideas!!!
#
# <https://www.kaggle.com/c/house-prices-advanced-regression-techniques>
#
# + [markdown] colab_type="text" id="1il4YYPMOZ4G"
# ## Lets generate some features!
#
# [King County Dataset Variable Meanings](https://www.kaggle.com/harlfoxem/housesalesprediction)
# + colab={"base_uri": "https://localhost:8080/", "height": 215} colab_type="code" id="4bbfD7HEOdo3" outputId="5207554a-002f-4652-cc13-74f9b2679c3d"
# Show columns so that we can reference them.
df.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 316} colab_type="code" id="ESFfcliEgt9y" outputId="31ef7d05-9969-442b-b294-b6a003fa7a25"
df['ln_price'] = np.log(df['price'])
print(df.columns)
df.head()
# + colab={} colab_type="code" id="dGHxukhjHsWu"
# Non-Feature Engineered Model
# Separate dependent and independent variables
y = df.loc[:, ['ln_price']].values
X = df.loc[:, ['bedrooms', 'bathrooms', 'sqft_living',
'sqft_lot', 'floors', 'waterfront', 'view', 'condition', 'grade',
'sqft_above', 'sqft_basement', 'yr_renovated',
'sqft_living15', 'sqft_lot15', 'grade_squared',]].values
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="h2sqokACHZZ1" outputId="f1b81be4-8780-4e6a-99b3-af573af73b98"
# Slit into test and train datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=.2, random_state=42)
# fit model using train datasets
model = LinearRegression()
model.fit(X_train, Y_train)
# Create new predictions using x_test
y_pred = model.predict(X_test)
# Measure Accuracy using y_test and y_pred
RMSE = (np.sqrt(mean_squared_error(Y_test, y_pred)))
R2 = r2_score(Y_test, y_pred)
print('RMSE is {}'.format(RMSE))
print('R^2 is {}'.format(R2))
# print("coefficients: ", model.coef_[0])
# print("intercepts: ", model.intercept_)
# + colab={} colab_type="code" id="ZD0a0JyUInGu"
df['age'] = 2015 - df['yr_built']
# + colab={} colab_type="code" id="w6Eq29eMIv0E"
y = df.loc[:, ['ln_price']].values
X = df.loc[:, ['bedrooms', 'bathrooms', 'sqft_living',
'sqft_lot', 'floors', 'waterfront', 'view', 'condition', 'grade',
'sqft_above', 'sqft_basement', 'yr_renovated',
'sqft_living15', 'sqft_lot15', 'grade_squared', 'age']].values
# + colab={"base_uri": "https://localhost:8080/", "height": 50} colab_type="code" id="7r-5WeZhIzai" outputId="4d3900ea-d21a-4b1c-ea96-bc14aafd5e78"
# Slit into test and train datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=.2, random_state=42)
# fit model using train datasets
model = LinearRegression()
model.fit(X_train, Y_train)
# Create new predictions using x_test
y_pred = model.predict(X_test)
# Measure Accuracy using y_test and y_pred
RMSE = (np.sqrt(mean_squared_error(Y_test, y_pred)))
R2 = r2_score(Y_test, y_pred)
print('RMSE is {}'.format(RMSE))
print('R^2 is {}'.format(R2))
# + [markdown] colab_type="text" id="O5TODtjOJZzr"
# ## A note about $R^2$
#
# $R^2$ If we add any feature to our model (even nearly meaningless ones) our $R^2$ will improve. For this reason raw $R^2$ is not the ultimate measure of goodness of fit. It is informative but completely secondary to our Root-Mean-Squared-Error (in predictive modeling). While a higher $R^2$ is generally better, this isn't the thing that we're trying to optimize. We care more about minimizing RMSE than maximizing $R^2$
#
# ### "Kitchen Sink" models
#
# You may hear the term "kitchen sink" regression model used. This refers to a regression model that throws every available explanatory variable into the model in an attempt to improve it without much thought to whether those variables should really be considered as affecting y. Kitchen Sink models will have a higher $R^2$ than others but will have higher standard errors (estimates about particular coefficients may be less precise).
#
# Therefore, you tend to see "Kitchen Sink" models when the only priority is predictive accuracy and not interpretability.
#
# ### Alternative measures of Goodness-of-fit
#
# Efforts have been made to improve upon $R^2$ due to these limitations. A metric called "Adjusted $R^2$" seeks to account for the number of explanatory variables included in a model and adjust $R^2$ accordingly. This is something that you can look up if you're curious. I just wanted to make you aware of it more than anything.
# + [markdown] colab_type="text" id="eRh1C4qJXWY3"
# ## A note about dirty linear regression data
#
# ### Linear Regression models can only process numeric values
#
# ### Your data must be free of NaN values before being passed to the algorithm
#
# (some data cleaning will be required in today's assignment)
# + [markdown] colab_type="text" id="_Nww7246e4pn"
# ## Interaction Terms
#
# An interaction occurs when an independent variable has a different effect on the outcome depending on the values of another independent variable.
#
# Lets look at an example where we were trying to estimate the level of satisfaction that a person would have when eating some kind of food with a condiment (sauce) on it.
#
# $satisfaction_i = \beta_0 + \beta_1 food_i + \beta_2condiment_i + \epsilon$
#
# Imagine that we have two foods: Ice Cream and Hot Dogs, and we also have to condiments: hot fudge and mustard.
#
# $\beta_1$ in this example is trying to capture the effect of on satisfaction between eating hot dogs vs eating ice cream, and $\beta_2$ is trying to capture the effect of putting hot fudge (chocolate sauce) vs mustard on your food.
#
# $\beta_2$ is a little more problematic in this scenario. If someone were to come up to you and ask if you preferred hot fudge or mustard on your food, how would you answer?
#
# You would probably say something like **"IT DEPENDS ON WHAT THE FOOD IS."** This means that the effect of our x variables on y (satisfaction) depends on the combination of food and condiment. I don't know about you guys, but I wouldn't be as satisfied if I had hot fudge on my hot dog or mustard on my ice cream.
#
# An interaction terms is something that we add to our regression to account for these "It Depends" moments between two x variables. We do this by multiplying the two of them together or *interacting* them with each other to capture the implications of the different combinations taking place.
#
# $satisfaction_i = \beta_0 + \beta_1 food_i + \beta_2condiment_i + \beta_3(food\times condiment_i) + \epsilon$
#
# <http://statisticsbyjim.com/regression/interaction-effects/>
# + [markdown] colab_type="text" id="pm-PTeevMFWV"
# # "Perfect Multicollinearity"
#
# Perfect Multicollinearity is very related to the concept of linear dependence of matrix vectors. Columns that are perfectly multi-collinear can be created via a linear combination of other columns in the dataset. Columns that differ from one another another by a constant amount (scalar) will be perfectly multicollinear with each other.
#
# It is easy to accidentally create perfectly multicollinear columns by doing unit conversions and then not dropping one or the other columns. You should not include columns that are perfectly multicollinear in your regression but should drop one or the other.
#
# Below is an example of us (accidentally) creating perfectly multicollinear columns. This is bad. Don't do this.
# + colab={"base_uri": "https://localhost:8080/", "height": 185} colab_type="code" id="3QXDRelv6nAB" outputId="4081e3f9-c14b-4804-d52a-130618a996f5"
## Perfect Multicollinearity example
df['bedrooms_x5'] = df['bedrooms']*5
## Non-Feature Engineered Model
# Separate dependent and independent variables
y = df.loc[:, ['ln_price']].values
X = df.loc[:, ['bedrooms', 'bathrooms', 'sqft_living',
'sqft_lot', 'floors', 'waterfront', 'view', 'condition', 'grade',
'sqft_above', 'sqft_basement', 'yr_built', 'yr_renovated', 'zipcode',
'sqft_living15', 'sqft_lot15', 'age', 'age_squared',
'total_bed_bath', 'renovation_size', 'hasBasement', 'sqft_lot_squared',
'sqft_living_squared', 'sqft_lot_squared15', 'sqft_living_squared15', 'bedrooms_x5']].values
# 'sqft_per_bedbath',
# Slit into test and train datasets
X_train, X_test, Y_train, Y_test = train_test_split(X, y, test_size=.5, random_state=42)
# fit model using train datasets
model = LinearRegression()
model.fit(X_train, Y_train)
# Create new predictions using x_test
y_pred = model.predict(X_test)
# Measure Accuracy using y_test and y_pred
RMSE = (np.sqrt(mean_squared_error(Y_test, y_pred)))
R2 = r2_score(Y_test, y_pred)
print('RMSE is {}'.format(RMSE))
print('R^2 is {}'.format(R2))
print("coefficients: ", model.coef_[0])
print("intercepts: ", model.intercept_)
# + [markdown] colab_type="text" id="yjaKkCJMM0LF"
# ## Showing Perfect Multi-Collinearity Graphically
#
# What perfect Multicollinearity really means is that two columns are linearly dependant with each other (from a linear algebra perspective). This means that the determinant of the X matrix will be zero causing it to therefore not be invertible. This will cause grave problems for models that are calculated using the linear algebra approach to Ordinary-Least-Squares regression.
#
# We can show that two columns are perfectly multicollinear by plotting the two variables against each other in a scatterplot and seeing that all of the cata points lie on the same line. (They're co-linear)
# + colab={"base_uri": "https://localhost:8080/", "height": 359} colab_type="code" id="0rZWjMjZ63lg" outputId="1d64f63d-b096-4751-d612-5d1bb344dfb7"
import matplotlib.pyplot as plt
plt.scatter(df['bedrooms'], df['bedrooms_x5'])
plt.show()
# + [markdown] colab_type="text" id="iBekeAlTNpIo"
# ## The "Dummy Variable Trap"
#
# Another way that perfectly multicollinear features can be created is by one-hot-ecoding a binary variable. (don't do this) If we one-hot-encode a binary variable we will end up with two columns that are perfect opposites of each other.
#
# With any categorical variable that is represented as dummy variables, perfect multi-collinearity will exist between the dummy-variable (binary) features of that categorical variable. So if you have used one-hot encoding to turn categorical variables into dummy variables, you will need to drop one of the resulting features to avoid this problem.
#
# Research the phrase "The Dummy Variable Trap" to learn more about this topic.
# + [markdown] colab_type="text" id="6ZH-5O0lLbJ7"
# ## Removing Outliers
#
# To remove outliers via the 1.5*Interquartile-Range method. The first step is to calculate the IQR for each variable.
#
# The IQR is the difference between the 25th and 75th percentiles of the feature.
#
# Find the IQR and multiply it by 1.5
#
# Then add the 1.5xIQR to the 3rd quartile (75th percentile). Anything above that range is an outlier.
# Subtract 1.5xIQR from the 1st quartile (25th percentile). Anything below that value is also an outlier.
#
# You want to minimize outliers in your dataset, so remove them by dropping observations that contain outliers in key features.
#
# Typically you will wan to remove outliers before doing anything else with your dataset. We haven't focused on this strongly yet in the class, but coefficients get strongly biased by outliers so if you want to really have accurate predictions, remove outliers before you begin your feature engineering and modeling.
# + [markdown] colab_type="text" id="nQ08zZkxW0hY"
# ## Major Takeaways
#
# - Polynomial Regression
# - Linear Regression can fit curved lines.
# - Including squared and cubed terms can improve fit and accuracy.
#
# - Log-linear Regression
# - ln(y) helps normalize data with a skewed y variable.
# - ln(y) changes interpretation of coefficients and errors to be percentages.
#
# - Feature Engineering
# - Generating Features improves model accuracy if done well.
# - This is where creativity and domain knowledge really pay off.
#
# - Interaction Terms
# - When you think that certain combinations of x variables might affect y differently than how they do separately. Include an interaction term.
| 06-Linear-Regression/02_Polynomial_Log_linear_Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment #01
#
# This assignment is composed of multiple Python and NumPy programming exercises. The deliverable of this assignment is this Jupyter notebook completed with your solution.
#
# The delivery instructions are the following:
# - Create a GitHub repository for your team. You are free to choose its name. This repository will be used to deliver the programming assignments and the code of your team's final project.
# - Create a folder assignment-01 in the repository. Put the Jupyter notebook with your solution in this repository and post the repository link in your team's D2L dropbox for the assignment.
#
# You are being assessed based on:
#
# 1. Code execution - 25%
# 2. Clarity of the code (e.g., easy to follow, has pertinent comments, etc.) - 25%
# 3. Proper usage of the techniques seen in class - 25%
# - Remember to avoid explicit Python loops!
# 4. Quality and correctness of the results - 25%
#
# More details about the assessment are available in the rubric document in the class GitHub repository.
#
# Points distribution:
# - Question 2-21: 3 points per question
# - Questions 22-26: 8 points per question (total 40)
# - Total: 100 points
import matplotlib.pylab as plt
# **1. Import the NumPy package under the name np**
import numpy as np
# **2. Print the NumPy version and the configuration**
# +
# Print the NumPy version
print("NumPy version:", np.__version__)
# Print the NumPy configuration
print("NumPy configuration:")
print(np.__config__.show())
# -
# **3. Create a 1D array of zeros of size 10 of type float32**
# +
# Generate array according to the specifications
q3 = np.zeros((10), dtype='float32')
# Print the array and type
print("Array:", q3)
print("Array type:", q3.dtype)
# -
# **4. Print the memory size of the array below**
# +
Z = np.zeros((32, 12))
# Obtain the memory size of the array in bytes
print("Array memory size: {} bytes".format(Z.nbytes))
# -
# **5. Create a 1D array with values ranging from 10 to 58**
# +
# Generate a 1D array containing values 10 to 58
q5 = np.arange(10, 59)
print("1D array containing values [10, 58]:\n", q5)
# -
# **6. Reverse the order of the values in the vector (first element becomes last)**
# +
a = np.arange(10)
# Reverse the order of the array (note that this is a shallow copy of a)
a_reversed = a[::-1]
print("Original array:", a)
print("Reversed array:", a_reversed)
# -
# **7. Create a 3x3 matrix with values ranging from 0 to 8**
# +
# Generate a 1D array with values of 0 to 8 and reshape it to be 3x3
q7 = np.arange(9).reshape(3, 3)
print("3x3 matrix containing values [0, 8]:\n", q7)
# -
# **8. Create a 3x3 identity matrix**
# +
# Generate a 3x3 identity matrix
q8 = np.identity(3)
print("3x3 identity matrix:\n", q8)
# -
# **9. Find and print the maximum values of each column in the array below**
# +
Z = np.random.random((10, 11))
# Obtain the maximum value of each column
Z_max_col = Z.max(axis=0)
print("Original array:\n", Z)
print("\nMaximum value of each column:\n", Z_max_col)
# -
# **10. Create a $7 \times 7$ array with 1s on the borders (first row and column, last row and column) and 0s everywhere else**
# +
# Generate a 7x7 matrix of ones
q10 = np.ones((7, 7))
# Fill the inner elements with zeros by specifying the proper indices
q10[1:-1, 1:-1] = 0
print("7x7 array with ones on the border and zeros everywhere else:\n", q10)
# -
# **11. Pad the existing array below with zeros (i.e., add additional rows and columns with zeros). Use the NumPy pad function.**
# +
Z = np.ones((5, 5))
# Pad the array with zeros
Z_padded = np.pad(Z, 1)
print("Original array:\n", Z)
print("\nPadded array:\n", Z_padded)
# -
# **12. Extract the integer part of the random array of positive numbers below**
# +
Z = np.random.uniform(0, 10, 10)
# Take the integer part of each element in the array
Z_int = Z.astype(int)
print("Original array:\n", Z)
print("\nInteger array:\n", Z_int)
# -
# **13. Create a 5x5 matrix with rows with equal values ranging from 0 to 4 (i.e., [0, 1, 2, 3, 4])**
#
# *Hint:* consider using NumPy broadcasting or the tile function*
# +
# Generate an array with rows containing values ranging from 0 to 4
q13 = np.tile([0, 1, 2, 3, 4], (5, 1))
print("5x5 matrix with rows containing values [0, 4]:\n", q13)
# -
# **14. Create a vector of size 10 with values ranging from 0 to 1 (0 and 1 not included)**
#
# *Hint: Consider using the linspace function*
# +
# Generate an array ranging from 0 to 1, slicing the array to exclude 0 and setting the linspace() 'endpoint' argument to 'False' to exclude 1
q14 = np.linspace(0, 1, 11, endpoint=False)[1:]
print("Array of size 10 containing values (0, 1):\n", q14)
# -
# **15. Create a random vector of size 10 and sort it**
# +
# Generate randomized array of size 10
q15_random = np.random.rand(10)
# Sort the randomized array
q15_sorted = np.sort(q15_random)
print("Random array:\n", q15_random)
print("\nSorted array:\n", q15_sorted)
# -
# **16. Consider the cartesian coordinates below (X and Y), convert them to polar coordinates (R and T)**
# +
Z = np.random.random((10, 2))
X, Y = Z[:, 0], Z[:, 1]
# Convert Cartesian coordinates to polar coordinates using the corresponding formula for the radius (R) and angle (T)
R = np.sqrt(X ** 2 + Y ** 2)
T = np.arctan2(Y, X)
print("Cartesian coordinates:")
print("X =", X)
print("Y =", Y)
print("\nPolar coordinates:")
print("R =", R)
print("T (radians) =", T)
# -
# **17. Create random vector of size 10 and replace the maximum value by 0**
# +
# Generate randomized array of size 10
q17_random = np.random.rand(10)
# Replace maximum value with 0 using a boolean mask
q17_max_replaced = q17_random.copy()
q17_max_replaced[q17_random.argmax()] = 0
print("Randomized array:\n", q17_random)
print("\nMax value replaced array:\n", q17_max_replaced)
# -
# **18. Find the nearest value from a given value (z) in an array (Z)**
# +
Z = np.random.uniform(0, 1, 10)
z = 0.5
# Retrive the nearest value by obtaining the index of the element in Z with the smallest difference to z
q18 = Z[np.abs(Z - z).argmin()]
print("Original array:\n", Z)
print("Value of z:", z)
print("Nearest value to z:", q18)
# -
# **19. Consider the vector [5, 4, 3, 2, 1], how to build a new vector with 3 consecutive zeros interleaved between each value?**
# +
Z = np.arange(6)[:0:-1]
# Generate a zeros array of appropriate length
Z_interleaved = np.zeros(4 * ((len(Z) - 1)) + 1, dtype=int)
# Replace every fourth element with an element in Z
Z_interleaved[::4] = Z
print("Original array:", Z)
print("Interleaved array:", Z_interleaved)
# -
# **20. Consider an array of dimension (5, 5, 3), how to mulitply it element-wise by an array with dimensions (5, 5) - i.e., (5, 5, 0)$\times$(5, 5), (5, 5, 1)$\times$(5, 5), (5, 5, 2)$\times$(5, 5)**
# +
A = np.ones((5, 5, 3))
B = 2 * np.ones((5, 5))
# Add third dimension to B and multiply with A
q20 = np.multiply(A, B.reshape(5, 5, 1))
print("Element-wise product:\n", q20)
print("\nShape of result:", q20.shape)
# -
# **21. Swap the first and second rows of the 2D array below**
# +
W = np.arange(30).reshape(6, 5)
print("Original array:\n", W)
print()
# Use advanced slicing to swap rows
W[[0, 1], :] = W[[1, 0], :]
print("Array with swapped rows:\n", W)
# -
# **22. Write a function that receives as input a 2D Boolean NumPy array and outputs the coordinates of the minimal bounding box
# that encloses all non-zero elements in the input array**
#
# **Tip:** Search for the functions non-zero and where in the NumPy documentation.
#
# <img src="./bbox.png" width="150" />
# +
def bbox(img):
# Obtain the row and column indices of the non-zero elements as a tuple
bb_idx = np.nonzero(img)
# Retrieve the minimum and maximum row/column indices to get the minimum bounding-box
r1 = bb_idx[0].min()
c1 = bb_idx[1].min()
r2 = bb_idx[0].max()
c2 = bb_idx[1].max()
return (r1, c1), (r2, c2)
from PIL import Image
img = np.array(Image.open('./bin_image.png').convert('L')) > 0
print("Minimum bounding box:", bbox(img))
# -
# **23. Write a function to compute the average over a sliding window of length N over a 1D array**
#
# *Example:* [1, 2, 3, 4, 5, 6, 7, 8, 9], N = 3 -> [2, 5, 8]
# +
def moving_average(x, N=3):
# Pad array with zeros if the size of array x is not a multiple of N
if x.size % N != 0:
x = np.pad(x, (0, x.size % N))
# Reshape the array to be 2D where each row has N elements and calculate the average of each
x = x.reshape(int(x.size / N), N)
x = x.sum(axis=1) / N
# Flatten back to a 1D array
x = x.flatten()
return x
a = np.arange(30)
N = 3
print("Original array:\n", a)
print("\nSliding window moving average:\n", moving_average(a, N))
# -
# **24. Given a sorted 1D array C that corresponds to a bincount, how to produce an array A such that np.bincount(A) == C?**
#
# *Example:* C = [3, 2, 4] -> D = [0, 0, 0, 1, 1, 2, 2, 2, 2]
#
# *Tip:* Consider using the NumPy repeat function
# +
C = np.bincount([1, 1, 2, 3, 4, 4, 6])
print("bincount array C =", C)
# Repeat the elements in A using the values of C
A = np.repeat(np.arange(C.size), C)
print("\nInverse bincount array A =", A)
print("bincount verification:", np.bincount(A))
# -
# **25. Find the most frequent value in the array below?**
# +
Z = np.random.randint(0, 10, 50)
# Record the instances of each number in the array
Z_counts = np.bincount(Z)
# Retrive the number with the maximum number of instances
Z_max = np.argmax(Z_counts)
print("Original array:\n", Z)
print("Array value counts:", Z_counts)
print("Most frequent array value:", Z_max)
# -
# **26. Write a function to return the n largest values of an array**
# +
def nlargest(a, n=5):
# Sort the array in ascending order
a.sort()
# Reverse array to descending order and return the first n elements
return a[::-1][0:n]
Z = np.arange(10000)
np.random.shuffle(Z)
n = 5
print("n largest values of the array:", nlargest(Z, n))
# -
# ## Team Contributions
#
# All team members worked on the Assignment 1 Jupyter notebook individually and met together to discuss and choose the best answer for each question.
#
# | Team Member | Consensus Score |
# | ---------------- | --------------- |
# | <NAME> | 3 |
# | <NAME> | 3 |
# | <NAME> | 3 |
# | <NAME> | 3 |
# | <NAME> | 3 |
# | <NAME> | 3 |
| assignment-work/assignment-1/assignment01-python-numpy-programming.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Machine Learning Engineer Nanodegree
# ## Deep Learning
# ## Project: Build a Digit Recognition Program
#
# In this notebook, a template is provided for you to implement your functionality in stages which is required to successfully complete this project. If additional code is required that cannot be included in the notebook, be sure that the Python code is successfully imported and included in your submission, if necessary. Sections that begin with **'Implementation'** in the header indicate where you should begin your implementation for your project. Note that some sections of implementation are optional, and will be marked with **'Optional'** in the header.
#
# In addition to implementing code, there will be questions that you must answer which relate to the project and your implementation. Each section where you will answer a question is preceded by a **'Question'** header. Carefully read each question and provide thorough answers in the following text boxes that begin with **'Answer:'**. Your project submission will be evaluated based on your answers to each of the questions and the implementation you provide.
#
# >**Note:** Code and Markdown cells can be executed using the **Shift + Enter** keyboard shortcut. In addition, Markdown cells can be edited by typically double-clicking the cell to enter edit mode.
# ----
# ## Step 1: Design and Test a Model Architecture
# Design and implement a deep learning model that learns to recognize sequences of digits. Train the model using synthetic data generated by concatenating character images from [notMNIST](http://yaroslavvb.blogspot.com/2011/09/notmnist-dataset.html) or [MNIST](http://yann.lecun.com/exdb/mnist/). To produce a synthetic sequence of digits for testing, you can for example limit yourself to sequences up to five digits, and use five classifiers on top of your deep network. You would have to incorporate an additional ‘blank’ character to account for shorter number sequences.
#
# There are various aspects to consider when thinking about this problem:
# - Your model can be derived from a deep neural net or a convolutional network.
# - You could experiment sharing or not the weights between the softmax classifiers.
# - You can also use a recurrent network in your deep neural net to replace the classification layers and directly emit the sequence of digits one-at-a-time.
#
# Here is an example of a [published baseline model on this problem](http://static.googleusercontent.com/media/research.google.com/en//pubs/archive/42241.pdf). ([video](https://www.youtube.com/watch?v=vGPI_JvLoN0))
# ### Implementation
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project. Once you have completed your implementation and are satisfied with the results, be sure to thoroughly answer the questions that follow.
# +
### Your code implementation goes here.
### Feel free to use as many code cells as needed.
# -
# ### Question 1
# _What approach did you take in coming up with a solution to this problem?_
# **Answer:**
# ### Question 2
# _What does your final architecture look like? (Type of model, layers, sizes, connectivity, etc.)_
# **Answer:**
# ### Question 3
# _How did you train your model? How did you generate your synthetic dataset?_ Include examples of images from the synthetic data you constructed.
# **Answer:**
# ----
# ## Step 2: Train a Model on a Realistic Dataset
# Once you have settled on a good architecture, you can train your model on real data. In particular, the [Street View House Numbers (SVHN)](http://ufldl.stanford.edu/housenumbers/) dataset is a good large-scale dataset collected from house numbers in Google Street View. Training on this more challenging dataset, where the digits are not neatly lined-up and have various skews, fonts and colors, likely means you have to do some hyperparameter exploration to perform well.
# ### Implementation
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project. Once you have completed your implementation and are satisfied with the results, be sure to thoroughly answer the questions that follow.
# +
### Your code implementation goes here.
### Feel free to use as many code cells as needed.
# -
# ### Question 4
# _Describe how you set up the training and testing data for your model. How does the model perform on a realistic dataset?_
# **Answer:**
# ### Question 5
# _What changes did you have to make, if any, to achieve "good" results? Were there any options you explored that made the results worse?_
# **Answer:**
# ### Question 6
# _What were your initial and final results with testing on a realistic dataset? Do you believe your model is doing a good enough job at classifying numbers correctly?_
# **Answer:**
# ----
# ## Step 3: Test a Model on Newly-Captured Images
#
# Take several pictures of numbers that you find around you (at least five), and run them through your classifier on your computer to produce example results. Alternatively (optionally), you can try using OpenCV / SimpleCV / Pygame to capture live images from a webcam and run those through your classifier.
# ### Implementation
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project. Once you have completed your implementation and are satisfied with the results, be sure to thoroughly answer the questions that follow.
# +
### Your code implementation goes here.
### Feel free to use as many code cells as needed.
# -
# ### Question 7
# _Choose five candidate images of numbers you took from around you and provide them in the report. Are there any particular qualities of the image(s) that might make classification difficult?_
# **Answer:**
# ### Question 8
# _Is your model able to perform equally well on captured pictures or a live camera stream when compared to testing on the realistic dataset?_
# **Answer:**
# ### Optional: Question 9
# _If necessary, provide documentation for how an interface was built for your model to load and classify newly-acquired images._
# **Answer:** Leave blank if you did not complete this part.
# ----
# ### Step 4: Explore an Improvement for a Model
#
# There are many things you can do once you have the basic classifier in place. One example would be to also localize where the numbers are on the image. The SVHN dataset provides bounding boxes that you can tune to train a localizer. Train a regression loss to the coordinates of the bounding box, and then test it.
# ### Implementation
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project. Once you have completed your implementation and are satisfied with the results, be sure to thoroughly answer the questions that follow.
# +
### Your code implementation goes here.
### Feel free to use as many code cells as needed.
# -
# ### Question 10
# _How well does your model localize numbers on the testing set from the realistic dataset? Do your classification results change at all with localization included?_
# **Answer:**
# ### Question 11
# _Test the localization function on the images you captured in **Step 3**. Does the model accurately calculate a bounding box for the numbers in the images you found? If you did not use a graphical interface, you may need to investigate the bounding boxes by hand._ Provide an example of the localization created on a captured image.
# **Answer:**
# ----
# ## Optional Step 5: Build an Application or Program for a Model
# Take your project one step further. If you're interested, look to build an Android application or even a more robust Python program that can interface with input images and display the classified numbers and even the bounding boxes. You can for example try to build an augmented reality app by overlaying your answer on the image like the [Word Lens](https://en.wikipedia.org/wiki/Word_Lens) app does.
#
# Loading a TensorFlow model into a camera app on Android is demonstrated in the [TensorFlow Android demo app](https://github.com/tensorflow/tensorflow/tree/master/tensorflow/examples/android), which you can simply modify.
#
# If you decide to explore this optional route, be sure to document your interface and implementation, along with significant results you find. You can see the additional rubric items that you could be evaluated on by [following this link](https://review.udacity.com/#!/rubrics/413/view).
# ### Optional Implementation
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project. Once you have completed your implementation and are satisfied with the results, be sure to thoroughly answer the questions that follow.
# +
### Your optional code implementation goes here.
### Feel free to use as many code cells as needed.
# -
# ### Documentation
# Provide additional documentation sufficient for detailing the implementation of the Android application or Python program for visualizing the classification of numbers in images. It should be clear how the program or application works. Demonstrations should be provided.
# _Write your documentation here._
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to
# **File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
| digit_recognition/digit_recognition.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Importamos las librerias necesarias
# Libreria para cinematica inversa
import ikpy
from ikpy import plot_utils,geometry_utils
from ikpy.chain import Chain
from ikpy.link import OriginLink, URDFLink
# Graficos
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import animation
from matplotlib.backends.backend_agg import FigureCanvasAgg as FigureCanvas
from matplotlib.figure import Figure
# %matplotlib inline
from IPython.display import HTML
from PIL import Image
# Operaciones
import numpy as np
import math
# Creamos el brazo, basandonos en el brazo izquierdo de Poppy-torso
left_arm_chain = Chain(name='left_arm', links=[
URDFLink(
name="shoulder_y",
translation_vector=[0, 0, 0],
orientation=[0, 0, 0],
rotation=[1, 0, 0],
),
URDFLink(
name="shoulder_x",
translation_vector=[-10, 0, 5],
orientation=[0, 1.57, 0],
rotation=[0, 1, 0],
),
URDFLink(
name="elbow",
translation_vector=[25, 0, 0],
orientation=[0, 0, 1.57],
rotation=[0, 0, 1],
),
URDFLink(
name="wrist",
translation_vector=[22, 0, 0],
orientation=[0, 0, 0],
rotation=[0, 1, 0],
)
])
# Creamos dos variables, una que obtiene el punto original del brazo y otra para establecer un punto objetivo
initial_pos = left_arm_chain.forward_kinematics([0] * 4) # Posicion inicial del brazo (4 enlaces, 4 posiciones)
initial_pos = list(np.around(initial_pos[:3, 3],decimals=2))
#target_pos = [-9.98,13.82,-26.33]
target_pos = [-16.23,-7.61,-15.68]
print((list(initial_pos),target_pos))
# Visualizamos el brazo y el punto objetivo que hemos puesto
plt.style.use('ggplot')
ax = plt.figure().add_subplot(111, projection='3d') # Este objeto almacena la figura
initial_pos = left_arm_chain.inverse_kinematics(geometry_utils.to_transformation_matrix([-9.98,13.82,-26.33]))
left_arm_chain.plot(initial_pos, ax, target=target_pos)
# Propiedades de los ejes
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
ax.set_xlim3d([-50.0, 50.0])
ax.set_ylim3d([-50.0, 50.0])
ax.set_zlim3d([-50.0, 50.0])
# Probamos el movimiento del brazo en cada uno de los grados de libertad
# +
trayectory = []
# Creamos la trayectoria
# Solo tomamos 3 DOF, ya que el ultimo enlace no tiene movilidad
for i in range(3):
for j in range(1,10):
t = np.zeros(4)
t[i]=math.radians(10*j)
trayectory.append(t)
fig = Figure(figsize=(10,10))
width, height = fig.get_size_inches() * fig.get_dpi()
canvas = FigureCanvas(fig)
ax = fig.add_subplot(111, projection='3d') # Este objeto almacena la figura
left_arm_chain.plot(initial_pos, ax, target=target_pos)
# Propiedades de los ejes
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
ax.set_xlim3d([-50.0, 50.0])
ax.set_ylim3d([-50.0, 50.0])
ax.set_zlim3d([-50.0, 50.0])
canvas.draw()
img = Image.fromarray(np.frombuffer(canvas.tostring_rgb(), dtype='uint8').reshape(int(height), int(width), 3), 'RGB')
im = plt.imshow(img, interpolation='none', aspect='auto', vmin=0, vmax=1)
# Funcion de actualizacion de datos
def animate_func(i):
fig.clear()
ax = fig.add_subplot(111, projection='3d')
#step = left_arm_chain.inverse_kinematics(geometry_utils.to_transformation_matrix(trayectory[i]))
step = trayectory[i]
left_arm_chain.plot(step, ax, target=target_pos)
# Propiedades de los ejes
ax.set_xlabel('X')
ax.set_ylabel('Y')
ax.set_zlabel('Z')
ax.set_xlim3d([-50.0, 50.0])
ax.set_ylim3d([-50.0, 50.0])
ax.set_zlim3d([-50.0, 50.0])
canvas.draw()
img = Image.fromarray(np.frombuffer(canvas.tostring_rgb(), dtype='uint8').reshape(int(height), int(width), 3), 'RGB')
im.set_array(img)
return [im]
# Renderizado
anim = animation.FuncAnimation(fig,animate_func,frames = len(trayectory),interval = 10000 / len(trayectory))
plt.close()
HTML(anim.to_html5_video())
# -
| Arm_3D/arm3d.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../../images/qiskit-heading.gif" alt="Note: In order for images to show up in this jupyter notebook you need to select File => Trusted Notebook" width="500 px" align="left">
# # Quantum Process Tomography
#
# * **Last Updated:** April 29, 2019
# * **Requires:** qiskit-terra 0.8, qiskit-ignis 0.1.1, qiskit-aer 0.2
# This notebook contains examples for using the ``ignis.verification.tomography`` process tomography module.
# +
# Needed for functions
import numpy as np
import time
# Import QISKit classes
import qiskit
from qiskit import QuantumRegister, QuantumCircuit, Aer
from qiskit.quantum_info import state_fidelity
from qiskit.tools.qi.qi import outer
# Tomography functions
from qiskit.ignis.verification.tomography import process_tomography_circuits, ProcessTomographyFitter
# -
# ## 1-qubit process tomography example
# +
# Process tomography of a Hadamard gate
q = QuantumRegister(1)
circ = QuantumCircuit(q)
circ.h(q[0])
# Run circuit on unitary simulator to find ideal unitary
job = qiskit.execute(circ, Aer.get_backend('unitary_simulator'))
ideal_unitary = job.result().get_unitary(circ)
# convert to Choi-matrix in column-major convention
choi_ideal = outer(ideal_unitary.ravel(order='F'))
# Generate process tomography circuits and run on qasm simulator
qpt_circs = process_tomography_circuits(circ, q)
job = qiskit.execute(qpt_circs, Aer.get_backend('qasm_simulator'), shots=4000)
# Extract tomography data so that counts are indexed by measurement configuration
qpt_tomo = ProcessTomographyFitter(job.result(), qpt_circs)
qpt_tomo.data
# +
# MLE Least-Squares tomographic reconstruction
t = time.time()
choi_lstsq = qpt_tomo.fit(method='lstsq')
print('Least-Sq Fitter')
print('fit time:', time.time() - t)
print('fit fidelity:', state_fidelity(choi_ideal / 2, choi_lstsq.data / 2))
# CVXOPT Semidefinite-Program tomographic reconstruction
t = time.time()
choi_cvx = qpt_tomo.fit(method='cvx')
print('\nCVXOPT Fitter')
print('fit time:', time.time() - t)
print('fit fidelity:', state_fidelity(choi_ideal / 2, choi_cvx.data / 2))
# -
# ## 1-qubit process tomography of two-qubit swap gate
#
# We will prepare qubit-0 and measure qubit-1 so the reconstructed channel should be an identity
# +
# Process tomography of a Hadamard gate
q = QuantumRegister(2)
circ = QuantumCircuit(q)
circ.swap(q[0], q[1])
# Ideal channel is a unitary
ideal_unitary = np.eye(2)
choi_ideal = outer(ideal_unitary.ravel(order='F'))
# Generate process tomography circuits and run on qasm simulator
# We use the optional prepared_qubits kwarg to specify that the prepared qubit was different to measured qubit
qpt_circs = process_tomography_circuits(circ, q[1], prepared_qubits=q[0])
job = qiskit.execute(qpt_circs, Aer.get_backend('qasm_simulator'), shots=2000)
# Extract tomography data so that counts are indexed by measurement configuration
qpt_tomo = ProcessTomographyFitter(job.result(), qpt_circs)
qpt_tomo.data
# +
# Least-Squares tomographic reconstruction
t = time.time()
choi_lstsq = qpt_tomo.fit(method='lstsq')
print('Least-Sq Fitter')
print('fit time:', time.time() - t)
print('fit fidelity:', state_fidelity(choi_ideal / 2, choi_lstsq.data / 2))
# CVXOPT Semidefinite-Program tomographic reconstruction
t = time.time()
choi_cvx = qpt_tomo.fit(method='cvx')
print('\nCVXOPT Fitter')
print('fit time:', time.time() - t)
print('fit fidelity:', state_fidelity(choi_ideal / 2, choi_cvx.data / 2))
# -
# ## 2-Qubit entangling circuit
# +
# Bell-state entangling circuit
q = QuantumRegister(2)
circ = QuantumCircuit(q)
circ.h(q[0])
circ.cx(q[0], q[1])
# Run circuit on unitary simulator to find ideal unitary
job = qiskit.execute(circ, Aer.get_backend('unitary_simulator'))
ideal_unitary = job.result().get_unitary(circ)
# convert to Choi-matrix in column-major convention
choi_ideal = outer(ideal_unitary.ravel(order='F'))
# Generate process tomography circuits and run on qasm simulator
qpt_circs = process_tomography_circuits(circ, q)
job = qiskit.execute(qpt_circs, Aer.get_backend('qasm_simulator'), shots=2000)
# Extract tomography data so that counts are indexed by measurement configuration
qpt_tomo = ProcessTomographyFitter(job.result(), qpt_circs)
t = time.time()
choi_lstsq = qpt_tomo.fit(method='lstsq')
print('Least-Sq Fitter')
print('fit time:', time.time() - t)
print('fit fidelity:', state_fidelity(choi_ideal / 4, choi_lstsq.data / 4))
t = time.time()
choi_cvx = qpt_tomo.fit(method='cvx')
print('\nCVXOPT Fitter')
print('fit time:', time.time() - t)
print('fit fidelity:', state_fidelity(choi_ideal / 4, choi_cvx.data / 4))
# -
# ## Using SIC-POVM preparation basis
# +
# Process tomography of a Hadamard gate
q = QuantumRegister(1)
circ = QuantumCircuit(q)
circ.h(q[0])
# Run circuit on unitary simulator to find ideal unitary
job = qiskit.execute(circ, Aer.get_backend('unitary_simulator'))
ideal_unitary = job.result().get_unitary(circ)
# convert to Choi-matrix in column-major convention
choi_ideal = outer(ideal_unitary.ravel(order='F'))
# Generate process tomography circuits and run on qasm simulator
qpt_circs = process_tomography_circuits(circ, q, prep_labels='SIC', prep_basis='SIC')
job = qiskit.execute(qpt_circs, Aer.get_backend('qasm_simulator'), shots=2000)
# Extract tomography data so that counts are indexed by measurement configuration
qpt_tomo = ProcessTomographyFitter(job.result(), qpt_circs, prep_basis='SIC')
qpt_tomo.data
# +
# MLE Least-Squares tomographic reconstruction
t = time.time()
choi_lstsq = qpt_tomo.fit(method='lstsq')
print('Least-Sq Fitter')
print('fit time:', time.time() - t)
print('fit fidelity:', state_fidelity(choi_ideal / 2, choi_lstsq.data / 2))
# CVXOPT Semidefinite-Program tomographic reconstruction
t = time.time()
choi_cvx = qpt_tomo.fit(method='cvx')
print('\nCVXOPT Fitter')
print('fit time:', time.time() - t)
print('fit fidelity:', state_fidelity(choi_ideal / 2, choi_cvx.data / 2))
| qiskit/ignis/process-tomography.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] papermill={"duration": 0.053248, "end_time": "2022-03-14T18:25:29.603702", "exception": false, "start_time": "2022-03-14T18:25:29.550454", "status": "completed"} tags=[]
# ## <span style="color:crimson;">WHAT IS A/B TESTING?
#
# A/B testing, also known as split testing, refers to a randomized experimentation process where two or more versions of a variable (web page, page element, etc.) are shown to different segments of website visitors at the same time to determine which version leaves the maximum impact and drive business metrics
#
# 
# + [markdown] papermill={"duration": 0.053449, "end_time": "2022-03-14T18:25:29.715457", "exception": false, "start_time": "2022-03-14T18:25:29.662008", "status": "completed"} tags=[]
# **<span style="color:crimson;">Context**
#
# Facebook recently introduced a new type of bidding, average bidding, as an alternative to the current type of bidding called maximum bidding.
# The system works in such a way that whoever pays more, its advertisement is shown to users more.
#
# We can say that; show this ad but I will pay maximum 10 USD or 6 USD on average per impression. If someone else pays maximum of 9 USD or an average of 5 USD to show their ads to the target customer, our ad will be shown to the user, because of the our higher offer.
#
# In this dataset you can find the analysis result of comparison of these two product regarding observations in last 40 days
#
# **<span style="color:crimson;">Business Problem**
#
# As a company that will advertise on Facebook, we want to understand which option is more advantageous for us.
#
# * Which of these two options will increase our number of clicks and purchases?
# * Is there a significant difference between the two options?
#
# To find the answer to these questions, we are applying the EU Test today.
#
# **<span style="color:crimson;">Task Details**
#
# * Importing Libraries and Data
# * Data Preprocessing
# * Analyse
# * A/B Test
# * Evaluation of the Results
#
# **<span style="color:crimson;">Attribute Information:**
#
# * **Impression :** Ad impressions
# * **Click :** Number of clicks on the displayed ad
# * **Purchase :** The number of products purchased after the ads clicked
# * **Earning:** Earnings after purchased products
# + [markdown] papermill={"duration": 0.050791, "end_time": "2022-03-14T18:25:29.819222", "exception": false, "start_time": "2022-03-14T18:25:29.768431", "status": "completed"} tags=[]
# ## <span style="color:crimson;">1. IMPORTING LIBRARIES AND DATA
# + papermill={"duration": 13.932745, "end_time": "2022-03-14T18:25:43.804399", "exception": false, "start_time": "2022-03-14T18:25:29.871654", "status": "completed"} tags=[]
# !pip install openpyxl
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import statsmodels.stats.api as sms
from scipy.stats import shapiro,levene,ttest_ind, mannwhitneyu
from scipy import stats
from statsmodels.stats.proportion import proportions_ztest
import warnings
warnings.filterwarnings('ignore')
import os
for dirname, _, filenames in os.walk('/kaggle/input'):
for filename in filenames:
print(os.path.join(dirname, filename))
pd.set_option('display.float_format', lambda x: '%.4f' % x)
# + papermill={"duration": 0.359771, "end_time": "2022-03-14T18:25:44.221869", "exception": false, "start_time": "2022-03-14T18:25:43.862098", "status": "completed"} tags=[]
control_df = pd.read_excel("../input/ab-testing-dataset/ab_testing.xlsx", sheet_name='Control Group')# maximum bidding
test_df = pd.read_excel("../input/ab-testing-dataset/ab_testing.xlsx", sheet_name='Test Group') # average bidding
# + papermill={"duration": 0.105475, "end_time": "2022-03-14T18:25:44.385207", "exception": false, "start_time": "2022-03-14T18:25:44.279732", "status": "completed"} tags=[]
# Control Group (Maximum bidding)
control_df.describe().T
# + papermill={"duration": 0.081311, "end_time": "2022-03-14T18:25:44.526180", "exception": false, "start_time": "2022-03-14T18:25:44.444869", "status": "completed"} tags=[]
# Test Goup (Average_bidding)
test_df.describe().T
# + [markdown] papermill={"duration": 0.057195, "end_time": "2022-03-14T18:25:44.641353", "exception": false, "start_time": "2022-03-14T18:25:44.584158", "status": "completed"} tags=[]
# ##### <span style="color:crimson;">Comparison of Purchase and Earning Descriptive Statistics of Two Groups
#
# I'm putting together the metrics I want to compare by creating a dataframe.
# + papermill={"duration": 0.081982, "end_time": "2022-03-14T18:25:44.782201", "exception": false, "start_time": "2022-03-14T18:25:44.700219", "status": "completed"} tags=[]
desc_compare_df = pd.DataFrame ({"Control_Impression": control_df["Impression"].describe (),"Test_Impression": test_df["Impression"].describe (),
"Control_Click": control_df["Click"].describe (),"Test_Click": test_df["Click"].describe (),
"Control_Purchase": control_df["Purchase"].describe (),"Test_Purchase": test_df["Purchase"].describe (),
"Control_Earning": control_df["Earning"].describe (),"Test_Earning": test_df["Earning"].describe ()})
# + papermill={"duration": 0.074578, "end_time": "2022-03-14T18:25:44.917703", "exception": false, "start_time": "2022-03-14T18:25:44.843125", "status": "completed"} tags=[]
desc_compare_df.head()
# + [markdown] papermill={"duration": 0.058254, "end_time": "2022-03-14T18:25:45.034366", "exception": false, "start_time": "2022-03-14T18:25:44.976112", "status": "completed"} tags=[]
# Although the mean and median of the test group are high, let's check if there is a significant difference since the standard deviation is also high. We can check this by calculating confidence interval of Purchase for both group
# + papermill={"duration": 0.070388, "end_time": "2022-03-14T18:25:45.164558", "exception": false, "start_time": "2022-03-14T18:25:45.094170", "status": "completed"} tags=[]
sms.DescrStatsW (control_df["Purchase"]).tconfint_mean ()
# + papermill={"duration": 0.074091, "end_time": "2022-03-14T18:25:45.301287", "exception": false, "start_time": "2022-03-14T18:25:45.227196", "status": "completed"} tags=[]
sms.DescrStatsW (test_df["Purchase"]).tconfint_mean ()
# + [markdown] papermill={"duration": 0.060305, "end_time": "2022-03-14T18:25:45.420849", "exception": false, "start_time": "2022-03-14T18:25:45.360544", "status": "completed"} tags=[]
# It can be seen that the confidence intervals of the Purchases for these two groups overlap. It means they are not significantly different from each other
# + papermill={"duration": 0.079514, "end_time": "2022-03-14T18:25:45.558708", "exception": false, "start_time": "2022-03-14T18:25:45.479194", "status": "completed"} tags=[]
earning_df = pd.DataFrame ({"Control_Earning": control_df["Earning"].describe (),
"Test_Earning": test_df["Earning"].describe ()})
earning_df.head()
# + papermill={"duration": 0.070851, "end_time": "2022-03-14T18:25:45.690029", "exception": false, "start_time": "2022-03-14T18:25:45.619178", "status": "completed"} tags=[]
sms.DescrStatsW (control_df["Earning"]).tconfint_mean ()
# + papermill={"duration": 0.071969, "end_time": "2022-03-14T18:25:45.822464", "exception": false, "start_time": "2022-03-14T18:25:45.750495", "status": "completed"} tags=[]
sms.DescrStatsW (test_df["Earning"]).tconfint_mean ()
# + [markdown] papermill={"duration": 0.060994, "end_time": "2022-03-14T18:25:45.942984", "exception": false, "start_time": "2022-03-14T18:25:45.881990", "status": "completed"} tags=[]
# It can be seen that the confidence intervals of the earnings for these two groups do not overlap. It means they are significantly different from each other. The average earning of the Test Group is higher than that of the Control Group.
# + [markdown] papermill={"duration": 0.062204, "end_time": "2022-03-14T18:25:46.064708", "exception": false, "start_time": "2022-03-14T18:25:46.002504", "status": "completed"} tags=[]
# # <span style="color:crimson;">AB TESTING
# ## <span style="color:crimson;">1. Assumption Check
# ### <span style="color:crimson;">1.1 Normality Assumption
#
# In order to apply an AB Test, the Normality and Variance homogeneity assumptions should be satisfied.
#
# If normality and variance homogeneity is provided, an independent two-sample t-test (parametric test) is applied.
# If normality and homogeneity of variance are not provided, the Mann-Whitney U test (non-parametric test) is performed.
#
# **<span style="color:crimson;">Normality Assumption Check**
#
# The Shapiro-Wilks Test will be applied for the Normality
#
# * **H0 :** There is no statistically significant difference between sample distribution and theoretical normal distribution
# * **H1 :** There is statistically significant difference between sample distribution and theoretical normal distribution
#
# H0 is rejected if the p_value is less than 0.05.
# + [markdown] papermill={"duration": 0.063533, "end_time": "2022-03-14T18:25:46.189484", "exception": false, "start_time": "2022-03-14T18:25:46.125951", "status": "completed"} tags=[]
# ##### <span style="color:crimson;">Graphical observation of the normal distribution
# + papermill={"duration": 1.538127, "end_time": "2022-03-14T18:25:47.788661", "exception": false, "start_time": "2022-03-14T18:25:46.250534", "status": "completed"} tags=[]
cols = ['Impression','Click','Purchase','Earning']
for col in cols:
fig, ax = plt.subplots(1,2)
fig.set_figheight(3.5)
fig.set_figwidth(10)
sns.distplot (control_df[col], hist=False,ax=ax[0])
sns.distplot (test_df[col], hist=False,ax=ax[1])
ax[0].set_title('Control')
ax[1].set_title('Test')
plt.show()
# + [markdown] papermill={"duration": 0.0683, "end_time": "2022-03-14T18:25:47.928962", "exception": false, "start_time": "2022-03-14T18:25:47.860662", "status": "completed"} tags=[]
# A normal distribution tendency is observed. But it may be missleading so it needs to be checked
# + papermill={"duration": 0.083428, "end_time": "2022-03-14T18:25:48.080066", "exception": false, "start_time": "2022-03-14T18:25:47.996638", "status": "completed"} tags=[]
print('Control Group \n')
#cols = ['Earning_Per_Click','Impression_Per_Click','Puchasing_Per_Impression']
for col in cols:
test_stat, pvalue = shapiro(control_df[col])
print(col)
print('Test Stat = %.4f, p-value = %.4f \n' % (test_stat, pvalue))
# + [markdown] papermill={"duration": 0.070707, "end_time": "2022-03-14T18:25:48.222753", "exception": false, "start_time": "2022-03-14T18:25:48.152046", "status": "completed"} tags=[]
# All p-values are higher than 0.05 it means H0 cannot be rejected. The assumption of normality is provided.
# + papermill={"duration": 0.085764, "end_time": "2022-03-14T18:25:48.379116", "exception": false, "start_time": "2022-03-14T18:25:48.293352", "status": "completed"} tags=[]
print('Test Group \n')
#cols = ['Earning_Per_Click','Impression_Per_Click','Puchasing_Per_Impression']
for col in cols:
test_stat, pvalue = shapiro(test_df[col])
print(col)
print('Test Stat = %.4f, p-value = %.4f \n' % (test_stat, pvalue) )
# + [markdown] papermill={"duration": 0.068652, "end_time": "2022-03-14T18:25:48.519500", "exception": false, "start_time": "2022-03-14T18:25:48.450848", "status": "completed"} tags=[]
# All p-values are higher than 0.05 it means H0 cannot be rejected. The assumption of normality is provided.
# + [markdown] papermill={"duration": 0.071588, "end_time": "2022-03-14T18:25:48.662991", "exception": false, "start_time": "2022-03-14T18:25:48.591403", "status": "completed"} tags=[]
# ### <span style="color:crimson;">1.2. Variance Homogeneity
# Levene’s Test will be applied for the Homogeneity of variances
#
# * **H0:** There is no statistically significant difference between the variance of variance of the related variables of the 2 groups.
#
# * **H1:** There is a statistically significant difference between the variance of variance of the related variables of the 2 groups.
#
# H0 is rejected if the p_value is less than 0.05.
#
# + papermill={"duration": 0.082863, "end_time": "2022-03-14T18:25:48.818797", "exception": false, "start_time": "2022-03-14T18:25:48.735934", "status": "completed"} tags=[]
for col in cols:
ttest_lev, p_value_lev = levene (control_df[col], test_df[col])
print(col)
print ("ttest statistics: {}\np_value: {}\n".format (ttest_lev, p_value_lev))
# + [markdown] papermill={"duration": 0.068344, "end_time": "2022-03-14T18:25:48.956772", "exception": false, "start_time": "2022-03-14T18:25:48.888428", "status": "completed"} tags=[]
# All of the p values except those for the Click are higher than 0.05, it means we cannot reject the H0 hypothesis. Therefore, we can say that there is NO statistically significant difference between the variance distributions of the Impression,Purchase and Earning values of the 2 groups.
#
# p value for click is lower than 0.05 so we can reject H0 hypothesis. There is statistically significant difference between the variance distributions of the Click values of the 2 groups.
# + [markdown] papermill={"duration": 0.068236, "end_time": "2022-03-14T18:25:49.092301", "exception": false, "start_time": "2022-03-14T18:25:49.024065", "status": "completed"} tags=[]
# ## <span style="color:crimson;">2. Implementation of the Hypothesis
#
# ### <span style="color:crimson;">2.1 Comparing Two Group Means
#
# For the Impression,Purchase and Earning normality and variance homogeneity is provided, an independent two-sample t-test (parametric test) will be applied.
# For Click normality assuption is provided but homogeneity of variance is not provided,so the Mann-Whitney U test (non-parametric test) will be performed.
#
# #### <span style="color:crimson;">2.1.1 Parametric Comparison (Independent Two-Sample T-Test)
# The Independent Two-Sample T Test compares the means of two independent groups in order to determine whether there is statistical evidence that the associated population means are significantly different.
#
# **Hypotheses**
#
# * **H0:** µ1 = µ2 (the two population means are equal)
#
# * **H1:** µ1 ≠ µ2 (the two population means are not equal)
#
#
#
# + papermill={"duration": 0.096996, "end_time": "2022-03-14T18:25:49.255601", "exception": false, "start_time": "2022-03-14T18:25:49.158605", "status": "completed"} tags=[]
ttest_ind_Imp, p_value_ind_Imp = ttest_ind (control_df["Impression"], test_df["Impression"], equal_var=True)
print ("ttest statistics: {}\np_value: {}".format (ttest_ind_Imp, p_value_ind_Imp))
# + papermill={"duration": 0.084878, "end_time": "2022-03-14T18:25:49.424348", "exception": false, "start_time": "2022-03-14T18:25:49.339470", "status": "completed"} tags=[]
ttest_ind_pur, p_value_ind_pur = ttest_ind (control_df["Purchase"], test_df["Purchase"], equal_var=True)
print ("ttest statistics: {}\np_value: {}".format (ttest_ind_pur, p_value_ind_pur))
# + papermill={"duration": 0.084138, "end_time": "2022-03-14T18:25:49.580214", "exception": false, "start_time": "2022-03-14T18:25:49.496076", "status": "completed"} tags=[]
ttest_ind_earn, p_value_ind_earn = ttest_ind (control_df["Earning"], test_df["Earning"], equal_var=True)
print ("ttest statistics: {}\np_value: {}".format (ttest_ind_earn, p_value_ind_earn))
# + [markdown] papermill={"duration": 0.073776, "end_time": "2022-03-14T18:25:49.726243", "exception": false, "start_time": "2022-03-14T18:25:49.652467", "status": "completed"} tags=[]
# #### <span style="color:crimson;">2.1.2 Nonparametric Comparison (Mann-Whitney U Test)
# + papermill={"duration": 0.082861, "end_time": "2022-03-14T18:25:49.881581", "exception": false, "start_time": "2022-03-14T18:25:49.798720", "status": "completed"} tags=[]
ttest_value_Click, p_value_Click = mannwhitneyu (control_df["Click"], test_df["Click"])
print ("ttest statistics: {}\np_value: {}".format (ttest_value_Click, p_value_Click))
# + [markdown] papermill={"duration": 0.071449, "end_time": "2022-03-14T18:25:50.028724", "exception": false, "start_time": "2022-03-14T18:25:49.957275", "status": "completed"} tags=[]
#
# ### <span style="color:crimson;">2.2 Two Group Ratio Comparison (Two Sample Ratio Test)
# + [markdown] papermill={"duration": 0.072102, "end_time": "2022-03-14T18:25:50.172784", "exception": false, "start_time": "2022-03-14T18:25:50.100682", "status": "completed"} tags=[]
# So far, we have compared the two groups over the features given in the data set. Now, we continue to compare over the proportional features we have derived. We will use Two Sample Ratio Test (z test) for ratio comparisons.
# + [markdown] papermill={"duration": 0.072508, "end_time": "2022-03-14T18:25:50.319001", "exception": false, "start_time": "2022-03-14T18:25:50.246493", "status": "completed"} tags=[]
# #### <span style="color:crimson;">Derivation of New Features for Test and Control Group
# + papermill={"duration": 0.098356, "end_time": "2022-03-14T18:25:50.491681", "exception": false, "start_time": "2022-03-14T18:25:50.393325", "status": "completed"} tags=[]
groups = [control_df, test_df]
for group in groups:
group["Click_Per_Impression"] = group["Click"] / group["Impression"]
group["Earning_Per_Click"] = group["Earning"] / group["Click"]
group["Puchasing_Per_Impression"] = group["Purchase"] / group["Impression"]
control_df.head ()
# + papermill={"duration": 0.090589, "end_time": "2022-03-14T18:25:50.656813", "exception": false, "start_time": "2022-03-14T18:25:50.566224", "status": "completed"} tags=[]
test_df.head ()
# + papermill={"duration": 0.081338, "end_time": "2022-03-14T18:25:50.811925", "exception": false, "start_time": "2022-03-14T18:25:50.730587", "status": "completed"} tags=[]
test_df["Click_Per_Impression"].mean(),control_df["Click_Per_Impression"].mean()
# + papermill={"duration": 0.082662, "end_time": "2022-03-14T18:25:50.967649", "exception": false, "start_time": "2022-03-14T18:25:50.884987", "status": "completed"} tags=[]
test_df["Earning_Per_Click"].mean(),control_df["Earning_Per_Click"].mean()
# + papermill={"duration": 0.081433, "end_time": "2022-03-14T18:25:51.123592", "exception": false, "start_time": "2022-03-14T18:25:51.042159", "status": "completed"} tags=[]
test_df["Puchasing_Per_Impression"].mean(),control_df["Puchasing_Per_Impression"].mean()
# + [markdown] papermill={"duration": 0.075193, "end_time": "2022-03-14T18:25:51.273966", "exception": false, "start_time": "2022-03-14T18:25:51.198773", "status": "completed"} tags=[]
# * Click_Per_Impression : The averages of the two groups seem to differ.
# * Earning_Per_Click : The averages of the two groups seem to close.
# * Puchasing_Per_Impression : The averages of the two groups seem to be same.
#
# With the two sample ratio test, it is checked whether this difference is statistically significant.
#
# + papermill={"duration": 0.228654, "end_time": "2022-03-14T18:25:51.580453", "exception": false, "start_time": "2022-03-14T18:25:51.351799", "status": "completed"} tags=[]
purchase_sum = np.array([control_df["Purchase"].sum(), test_df["Purchase"].sum()])
click_sum = np.array([control_df["Click"].sum(), test_df["Click"].sum()])
impression_sum = np.array([control_df["Impression"].sum(), test_df["Impression"].sum()])
earning_sum = np.array([control_df["Earning"].sum(), test_df["Earning"].sum()])
# + [markdown] papermill={"duration": 0.076972, "end_time": "2022-03-14T18:25:51.735224", "exception": false, "start_time": "2022-03-14T18:25:51.658252", "status": "completed"} tags=[]
# #### <span style="color:crimson;">Click_Per_Impression</span>
# + papermill={"duration": 0.086562, "end_time": "2022-03-14T18:25:51.899793", "exception": false, "start_time": "2022-03-14T18:25:51.813231", "status": "completed"} tags=[]
ttest_z_click_imp, p_click_imp = proportions_ztest (click_sum, impression_sum)
print('Test Stat = %.4f, p-value = %.4f' % (ttest_z_click_imp, p_click_imp))
# + [markdown] papermill={"duration": 0.075237, "end_time": "2022-03-14T18:25:52.052542", "exception": false, "start_time": "2022-03-14T18:25:51.977305", "status": "completed"} tags=[]
# #### <span style="color:crimson;">Earning_Per_Click</span>
# + papermill={"duration": 0.085942, "end_time": "2022-03-14T18:25:52.214646", "exception": false, "start_time": "2022-03-14T18:25:52.128704", "status": "completed"} tags=[]
ttest_z_earn_click, p_earn_click = proportions_ztest (earning_sum, click_sum)
print('Test Stat = %.4f, p-value = %.4f' % (ttest_z_earn_click, p_earn_click))
# + [markdown] papermill={"duration": 0.070755, "end_time": "2022-03-14T18:25:52.364866", "exception": false, "start_time": "2022-03-14T18:25:52.294111", "status": "completed"} tags=[]
# #### <span style="color:crimson;">Puchasing_Per_Impression</span>
# + papermill={"duration": 0.084569, "end_time": "2022-03-14T18:25:52.523588", "exception": false, "start_time": "2022-03-14T18:25:52.439019", "status": "completed"} tags=[]
ttest_z_click_imp, p_click_imp = proportions_ztest (purchase_sum, impression_sum)
print('Test Stat = %.4f, p-value = %.4f' % (ttest_z_click_imp, p_click_imp))
# + [markdown] papermill={"duration": 0.072396, "end_time": "2022-03-14T18:25:52.667805", "exception": false, "start_time": "2022-03-14T18:25:52.595409", "status": "completed"} tags=[]
# * **Click_Per_Impression :** p < 0.05 so the averages of the two groups seem to differ. Control group which means Maximum bidding has higher average.
# * **Earning_Per_Click :** p < 0.05 so the averages of the two groups seem to differ. Test group which means Average bidding has higher average
# * **Puchasing_Per_Impression :** p < 0.05 so the averages of the two groups seem to differ. Control group which means Maximum bidding has higher average.
# + [markdown] papermill={"duration": 0.090497, "end_time": "2022-03-14T18:25:52.838918", "exception": false, "start_time": "2022-03-14T18:25:52.748421", "status": "completed"} tags=[]
# ### <span style="color:crimson;">3 Conclusion</span>
# + papermill={"duration": 0.090331, "end_time": "2022-03-14T18:25:53.006364", "exception": false, "start_time": "2022-03-14T18:25:52.916033", "status": "completed"} tags=[]
test_df["Purchase"].mean(),control_df["Purchase"].mean()
# + papermill={"duration": 0.088389, "end_time": "2022-03-14T18:25:53.177411", "exception": false, "start_time": "2022-03-14T18:25:53.089022", "status": "completed"} tags=[]
test_df["Click"].mean(),control_df["Click"].mean()
# + papermill={"duration": 0.086633, "end_time": "2022-03-14T18:25:53.343128", "exception": false, "start_time": "2022-03-14T18:25:53.256495", "status": "completed"} tags=[]
test_df["Impression"].mean(),control_df["Impression"].mean()
# + papermill={"duration": 0.085224, "end_time": "2022-03-14T18:25:53.503163", "exception": false, "start_time": "2022-03-14T18:25:53.417939", "status": "completed"} tags=[]
test_df["Earning"].mean(),control_df["Earning"].mean()
# + papermill={"duration": 0.084307, "end_time": "2022-03-14T18:25:53.666172", "exception": false, "start_time": "2022-03-14T18:25:53.581865", "status": "completed"} tags=[]
test_df["Click_Per_Impression"].mean(),control_df["Click_Per_Impression"].mean()
# + papermill={"duration": 0.090116, "end_time": "2022-03-14T18:25:53.830360", "exception": false, "start_time": "2022-03-14T18:25:53.740244", "status": "completed"} tags=[]
test_df["Earning_Per_Click"].mean(),control_df["Earning_Per_Click"].mean()
# + papermill={"duration": 0.09212, "end_time": "2022-03-14T18:25:53.997933", "exception": false, "start_time": "2022-03-14T18:25:53.905813", "status": "completed"} tags=[]
test_df["Puchasing_Per_Impression"].mean(),control_df["Puchasing_Per_Impression"].mean()
# + [markdown] papermill={"duration": 0.07411, "end_time": "2022-03-14T18:25:54.149998", "exception": false, "start_time": "2022-03-14T18:25:54.075888", "status": "completed"} tags=[]
#
# * **Purchase :** Maximum bidding (Control Group) and Average bidding (Test Group) has the same average
# * **Click :** Maximum bidding (Control Group) has higher average.
# * **Impression :** Average bidding (Test Group) has higher average
# * **Earning :** Average bidding (Test Group) has higher average
# * **Click_Per_Impression :** Maximum bidding (Control Group) has higher average.
# * **Earning_Per_Click :** Average bidding (Test Group) has higher average
# * **Puchasing_Per_Impression :**Maximum bidding (Control Group) has higher average.
| AB TEST/ab-test-of-ad-bidding.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Brain Tumor Classification with PyTorch⚡Lightning & EfficientNet 3D
#
# The goal of this challenge is to Predict the status of a genetic biomarker important for brain cancer treatment.
#
# All the code is refered from public repository: https://github.com/Borda/kaggle_brain-tumor-3D
# Any nice contribution is welcome!
# + tags=[]
# ! pip uninstall -y kaggle_brain3d
# ! pip install -q https://github.com/Borda/kaggle_brain-tumor-3D/archive/refs/tags/v0.1.0.zip
# ! pip uninstall -q -y wandb
# ! pip list | grep torch
# ! ls -l /home/jovyan/work/rsna-miccai-brain-tumor
# ! nvidia-smi
# ! mkdir /home/jovyan/temp
# %matplotlib inline
# %reload_ext autoreload
# %autoreload 2
import kaggle_brain3d
print(kaggle_brain3d.__version__)
# -
# ## Data exploration
#
# These 3 cohorts are structured as follows: Each independent case has a dedicated folder identified by a five-digit number.
# Within each of these “case” folders, there are four sub-folders, each of them corresponding to each of the structural multi-parametric MRI (mpMRI) scans, in DICOM format.
# The exact mpMRI scans included are:
#
# - **FLAIR**: Fluid Attenuated Inversion Recovery
# - **T1w**: T1-weighted pre-contrast
# - **T1Gd**: T1-weighted post-contrast
# - **T2w**: T2-weighted
#
# #### according to https://www.aapm.org/meetings/amos2/pdf/34-8205-79886-720.pdf
#
# - T1: weighting weighting better deliniates deliniates anatomy anatomy
# - T2: weighting weighting naturally naturally shows pathology
#
# #### according to https://radiopaedia.org/articles/fluid-attenuated-inversion-recovery
#
# Fluid attenuated inversion recovery (FLAIR) is a special inversion recovery sequence with a long inversion time. This removes signal from the cerebrospinal fluid in the resulting images 1. Brain tissue on FLAIR images appears similar to T2 weighted images with grey matter brighter than white matter but CSF is dark instead of bright.
#
# To null the signal from fluid, the inversion time (TI) of the FLAIR pulse sequence is adjusted such that at equilibrium there is no net transverse magnetization of fluid.
#
# The FLAIR sequence is part of almost all protocols for imaging the brain, particularly useful in the detection of subtle changes at the periphery of the hemispheres and in the periventricular region close to CSF.
# + tags=[]
import os
import glob
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
PATH_DATASET = "/home/jovyan/work/rsna-miccai-brain-tumor"
PATH_TEMP = "/home/jovyan/temp"
SCAN_TYPES = ("FLAIR", "T1w", "T1CE", "T2w")
df_train = pd.read_csv(os.path.join(PATH_DATASET, "train_labels.csv"))
df_train["BraTS21ID"] = df_train["BraTS21ID"].apply(lambda i: "%05d" % i)
display(df_train.head())
# -
# See the dataset label distribution
# + tags=[]
_= df_train["MGMT_value"].value_counts().plot(kind="pie", title="label distribution", autopct="%.1f%%")
# -
# For almost all scans we have all four types
scans = [os.path.basename(p) for p in glob.glob(os.path.join(PATH_DATASET, "train", "*", "*"))]
_= pd.Series(scans).value_counts().plot(kind="bar", grid=True)
# ### Interactive view
#
# showing particular scan in XYZ dimension/slices
# + tags=[]
from ipywidgets import interact, IntSlider
from kaggle_brain3d.utils import load_volume, interpolate_volume, show_volume
from kaggle_brain3d.transforms import crop_volume
def interactive_show(volume_path: str, crop_thr: float):
print(f"loading: {volume_path}")
volume = load_volume(volume_path, percentile=0)
print(f"sample shape: {volume.shape} >> {volume.dtype}")
volume = interpolate_volume(volume)
print(f"interp shape: {volume.shape} >> {volume.dtype}")
volume = crop_volume(volume, crop_thr)
print(f"crop shape: {volume.shape} >> {volume.dtype}")
vol_shape = volume.shape
interact(
lambda x, y, z: plt.show(show_volume(volume, x, y, z)),
x=IntSlider(min=0, max=vol_shape[0], step=5, value=int(vol_shape[0] / 2)),
y=IntSlider(min=0, max=vol_shape[1], step=5, value=int(vol_shape[1] / 2)),
z=IntSlider(min=0, max=vol_shape[2], step=5, value=int(vol_shape[2] / 2)),
)
PATH_SAMPLE_VOLUME = os.path.join(PATH_DATASET, "train", "00005", "FLAIR")
interactive_show(PATH_SAMPLE_VOLUME, crop_thr=1e-6)
# -
# ## Prepare dataset
#
# ### Pytorch Dataset
#
# The basic building block is traforming raw data to Torch Dataset.
# We have here loading particular DICOM images into a volume and saving as temp/cacher, so we do not need to take the very time demanding loading do next time - this boost the IO from about 2h to 8min
#
# At the end we show a few sample images from prepared dataset.
# + tags=[]
import os
import pandas as pd
import torch
from tqdm.auto import tqdm
from kaggle_brain3d.data import BrainScansDataset
from kaggle_brain3d.transforms import resize_volume
# ==============================
ds = BrainScansDataset(
image_dir=os.path.join(PATH_DATASET, "train"),
df_table=os.path.join(PATH_DATASET, "train_labels.csv"),
crop_thr=None, cache_dir=PATH_TEMP,
)
for i in tqdm(range(2)):
img = ds[i * 10]["data"]
img = resize_volume(img[0])
show_volume(img, fig_size=(9, 6))
# -
# ### Lightning DataModule
#
# It is constric to wrap all data-related peaces and define Pytoch dataloder for Training / Validation / Testing phase.
#
# At the end we show a few sample images from the fost training batch.
# + tags=[]
from functools import partial
import rising.transforms as rtr
from rising.loading import DataLoader, default_transform_call
from rising.random import DiscreteParameter, UniformParameter
from kaggle_brain3d.data import BrainScansDM, TRAIN_TRANSFORMS, VAL_TRANSFORMS
from kaggle_brain3d.transforms import RandomAffine, rising_zero_mean
# ==============================
# Dataset >> mean: 0.13732214272022247 STD: 0.24326834082603455
rising_norm = partial(rising_zero_mean, mean=0.137, std=0.243)
# define transformations
TRAIN_TRANSFORMS = [
rtr.Rot90((0, 1, 2), keys=["data"], p=0.5),
rtr.Mirror(dims=DiscreteParameter([0, 1, 2]), keys=["data"]),
RandomAffine(scale_range=(0.9, 1.1), rotation_range=(-10, 10), translation_range=(-0.1, 0.1)),
rising_norm,
]
VAL_TRANSFORMS = [
rising_norm,
]
# ==============================
dm = BrainScansDM(
data_dir=PATH_DATASET,
scan_types=["T2w"],
input_size=224,
crop_thr=1e-6,
batch_size=3,
cache_dir=PATH_TEMP,
# in_memory=True,
num_workers=6,
train_transforms=rtr.Compose(TRAIN_TRANSFORMS, transform_call=default_transform_call),
valid_transforms=rtr.Compose(VAL_TRANSFORMS, transform_call=default_transform_call),
)
# dm.prepare_data(3)
dm.setup()
print(f"Training batches: {len(dm.train_dataloader())} and Validation {len(dm.val_dataloader())}")
# Quick view
for batch in dm.train_dataloader():
for i in range(2):
show_volume(batch["data"][i][0], fig_size=(9, 6), v_min_max=(-1., 3.))
break
# -
# ## Prepare 3D model
#
# LightningModule is the core of PL, it wrappes all model related peaces, mainly:
#
# - the model/architecture/weights
# - evaluation metrics
# - configs for optimizer and LR cheduler
# + tags=[]
from torchsummary import summary
from kaggle_brain3d.models import LitBrainMRI
# ==============================
model = LitBrainMRI(lr=1e-3)
# summary(model, input_size=(1, 128, 128, 128))
# -
# ## Train a model
#
# Lightning forces the following structure to your code which makes it reusable and shareable:
#
# - Research code (the LightningModule).
# - Engineering code (you delete, and is handled by the Trainer).
# - Non-essential research code (logging, etc... this goes in Callbacks).
# - Data (use PyTorch DataLoaders or organize them into a LightningDataModule).
#
# Once you do this, you can train on multiple-GPUs, TPUs, CPUs and even in 16-bit precision without changing your code!
# + tags=[]
import pytorch_lightning as pl
logger = pl.loggers.CSVLogger(save_dir='logs/', name=model.name)
swa = pl.callbacks.StochasticWeightAveraging(swa_epoch_start=0.6)
ckpt = pl.callbacks.ModelCheckpoint(
monitor='valid/f1',
save_top_k=1,
save_last=True,
filename='checkpoint/{epoch:02d}-{valid_acc:.4f}-{valid_f1:.4f}',
mode='max',
)
# ==============================
trainer = pl.Trainer(
# overfit_batches=5,
# fast_dev_run=True,
gpus=1,
callbacks=[ckpt], # , swa
logger=logger,
max_epochs=30,
precision=16,
accumulate_grad_batches=12,
# val_check_interval=0.5,
progress_bar_refresh_rate=1,
log_every_n_steps=5,
weights_summary='top',
auto_lr_find=True,
# auto_scale_batch_size='binsearch',
)
# ==============================
trainer.tune(
model,
datamodule=dm,
lr_find_kwargs=dict(min_lr=2e-5, max_lr=1e-2, num_training=35),
# scale_batch_size_kwargs=dict(max_trials=5),
)
print(f"Batch size: {dm.batch_size}")
print(f"Learning Rate: {model.learning_rate}")
# ==============================
trainer.fit(model=model, datamodule=dm)
# -
# ### Training progress
# +
metrics = pd.read_csv(f'{trainer.logger.log_dir}/metrics.csv')
display(metrics.head())
aggreg_metrics = []
agg_col = "epoch"
for i, dfg in metrics.groupby(agg_col):
agg = dict(dfg.mean())
agg[agg_col] = i
aggreg_metrics.append(agg)
df_metrics = pd.DataFrame(aggreg_metrics)
df_metrics[['train/loss', 'valid/loss']].plot(grid=True, legend=True, xlabel=agg_col)
df_metrics[['train/f1', 'train/auroc', 'valid/f1', 'valid/auroc']].plot(grid=True, legend=True, xlabel=agg_col)
# -
| notebooks/Brain-tumor-classif_PT-Lightning_EfficientNet3D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Conversions décimal-binaire-hexadécimal en langage Python
#
# Dans ce notebook nous allons étudier les fonctions natives de Python qui permettent de réaliser des conversions entre les différents systèmes de réprésentation des nombres.
#
# Nous écrirons ensuite l'intégralité du code permettant, sur la carte BBC MicroBit, de convertir en décimal un nombre entré en binaire à l'aide de deux boutons poussoirs.
#
# Remarque : il faut aussi être capable de faire des conversions "à la main" : revoir le polycopié "Représentation des données : Types et valeurs de base" étudié en cours.
#
# ## 1- Ecriture d'un nombre en Python
# Les activités liées au numérique peuvent nécessiter parfois de travailler en binaire ou en hexadécimal (qui est en fait une forme d’écriture plus compacte) plutôt que dans notre système décimal usuel. On peut même travailler en octal...
# Cela pourrait être source de confusion : par exemple, que signifie l’écriture du nombre « 1000 » ?
# * en décimal : mille
# * en binaire l’écriture 1000 a comme équivalent décimal le nombre 8
# * en hexadécimal, l’écriture 1000 a comme équivalent décimal le nombre 4096
# Pour éviter les risques de confusions, on précisera la base dans laquelle le nombre est écrit. Malheureusement il n’y a pas de convention unique… la précision de la base se faisant par un préfixe ou par un suffixe, et pas toujours le même !!!
#
# En Python, la convention est la suivante :
# * un nombre entier, écrit comme on le fait dans la vie courante est représenté en décimal
# * précédé du préfixe 0b ce nombre est écrit en binaire (il est bien sûr constitué uniquement de 0 et de 1)
# * précédé du préfixe 0x ce nombre est écrit en hexadécimal
# * précédé du préfixe 0o il est écrit en octal
#
#
# écrire un nombre dans la base de son choix puis lancer l'interpréteur Python avec la combinaison
# de touches {Ctrl + Entrée} pour en obtenir l'écriture décimale
# nombre à convertir en décimal :
print(0x1000)
# ## 2- Fonctions natives de conversion :
#
# Le langage Python possède 3 fonctions natives pour réaliser la conversion d'un nombre
# (**fonctions natives** ou "**builtin functions**" en anglais : ce sont des fonctions qui font partie du langage et que l'on n'a donc pas besoin d'aller chercher dans un module externe) :
#
# * bin(ce_nombre) pour convertir ce_nombre en binaire
# * hex(ce_nombre) pour convertir ce_nombre en hexadécimal
# * int(ce_nombre) pour convertir ce_nombre en décimal
#
# ce_nombre pourra être donné en décimal, en binaire ou en hexadécimal en respectant le format vu au paragraphe précédent.
#
# Exemple : pour convertir le nombre décimal 141 en binaire, on entrerait dans l’interpréteur : bin(141), ce qui retournerait '0b10001101'.
#
#
# ### 2.1- Conversion décimal <-> binaire :
#
# Dans la cellule ci-dessous, entrer le code permettant :
#
# 1- de convertir le nombre décimal '125' en binaire
#
# 2- de convertir le nombre binaire '10011101' en décimal
# Remarque : on utilisera la fonction print() pour afficher et garder à l'écran les différents résultats
# +
# Conversion de '125' en binaire :
# Conversion de '10011101' en décimal :
# -
# ### 2.2- Conversion décimal <-> hexadécimal :
#
# Dans la cellule ci-dessous, entrer le code permettant :
#
# 1- de convertir et d'afficher le nombre décimal '125' en hexadécimal
#
# 2- de convertir le nombre hexadécimal '0FA4'en décimal
# +
# Conversion de '125' en hexadécimal :
# Conversion de '0FA4' en décimal :
# -
# ### 2.3- Synthèse :
# **EXERCICE 1:** Réaliser les conversions demandées ci-dessous ; observer le nombre de bits nécessaires à l'affichage en binaire
# +
# Convertir 0xF en décimal puis en binaire
# Convertir 0xFF en décimal puis en binaire
# Convertir 0xFFFF en décimal puis en binaire
# -
# **EXERCICE 2:**
# On veut avoir un programme qui demande à l'utilisateur un nombre **n** en décimal et qui en réalise (sur une ligne d'affichage) la conversion en binaire et en hexadécimal. Le rédiger dans la cellule de code ci-dessous :
# Programme de conversion décimal vers binaire et hexadécimal :
# ## 3- Convertisseur Binaire -> Décimal avec la carte BBC Micro:Bit
#
#
#
# Au paragraphe précédent,lors d'une telle conversion, le nombre binaire était passé en une seule fois à la fonction int(). Ici le problème est différent car l'octet à convertir sera entré bit par bit à l'aide des deux boutons poussoirs A et B présents sur la carte Micro:Bit :
#
# 
#
# **Cahier des charges du code à écrire :**
#
# * au démarrage du programme, on rentre dans une boucle infinie en affichant le symbole ‘?’ (le programme tourne alors en boucle en attendant l'appui sur l'un des deux boutons poussoirs
#
# * le code binaire constituant l’octet à convertir sera entré à l’aide des deux boutons poussoirs :
# - si le bouton A est appuyé, on entre alors un 0 ;
# - si le bouton B est appuyé, ce sera alors un 1 ;
# - le code binaire est entré de la gauche vers la droite. Exemple : pour entrer le code 01100001, on appuiera 1 fois sur le bouton A puis 2 fois sur le bouton B, puis 4 fois sur le bouton A et une fois sur le bouton B.
#
# * A chaque appui sur l’un des boutons l’afficheur devra afficher soit un 0 soit un 1 pendant une demi-seconde puis s’effacer en attendant un nouvel appui sur un des boutons.
#
# * Après le huitième appui, l’octet étant alors constitué, on fait défiler sur l’afficheur la valeur décimale de
# cet octet, puis on retourne au début de la boucle infinie pour pouvoir faire une nouvelle conversion.
#
# Au moins trois techniques différentes pourraient être envisagées :
# * l'une arithmétique basée sur la technique de pondération des chiffres en fonction de la position
# * en utilisant les opérateurs de décalage de bits ( >> ou << )
# * à l'aide d'une séquence ordonnée (liste, chaîne de caractères ...)
#
# Les deux dernières techniques nécessitant des connaissances plus approfondies, nous utiliserons la première méthode, qui correspond aussi à la présentation qui en a été faite dans le cours.
#
#
# ### 3.1- Principe mis en oeuvre :
# Le cahier des charges impose de travailler sur un octet, donc sur 8 bits que l'on va nommer de b0 à b7.
# Chacun de ces bits a un poids tel qu'indiqué ci-dessous :
#
# b7 | b6 | b5 | b4 | b3 | b2 | b1 | b0
# :----: | :----: | :----: |:----: | :----: | :----: |:----: | :----:
# 128 | 64 | 32 | 16 | 8 | 4 | 2 | 1
#
#
# Par exemple la conversion de l'octet 1 1 0 1 0 0 1 0 donnera : 210
#
# Ci-dessous deux méthodes arithmétiques similaires utilisables dans un script Python pour convertir le nombre binaire '0b11010010' en nombre décimal.
#
# Remarques :
#
# - en Python le symbole de multiplication est l'étoile : '*',
# - l'élévation à une puissance se réalise avec la double étoile '**'
# - la deuxième écriture (celle avec les puissances de 2) permet de mieux faire apparaître l'algorithme que l'on pourrait mettre en oeuvre.
#
octet = 128*1 + 64*1 + 32*0 + 16*1 + 8*0 + 4*0 + 2*1 + 1*0
print(octet)
# ou encore :
octet = 2**7*1 + 2**6*1 + 2**5*0 + 2**4*1 + 2**3*0 + 2**2*0 + 2**1*1 + 2**0*0
print(octet)
# ... mais de nouveau, on a entré l'intégralité du nombre binaire en une seule fois avant de le convertir.
#
# Comme l'octet sera entré en 8 étapes, on peut envisager de construire son équivalent décimal en autant d'étapes :
# +
# Compléter le code ci-dessous pour obtenir en sortie un nombre décimal (représenté par la variable 'octet') égal
# à 210 pour une entrée en pas à pas de chacun des bits : 1 1 0 1 0 0 1 0
# - ne pas oublier les commentaires...
# - valider votre code à chaque étape à l'aide de la combinaison de touche {Ctrl + Entrée} pour voir la construction
# progressive de cette conversion arithmétique :
octet = 0
# le bit7 vient d'être entré avec le bouton poussoir B (= 1): l'octet a donc changé :
octet = octet + 2**7
# le bit6 vient d'être entré avec le bouton poussoir ... .... ....
# ???
...
# ???
# ... et à la fin on affiche la valeur de l'octet :
print(octet)
# -
# Le programme précédent nous donne l'algorithme à suivre :
#
# - on définit une variable *octet* et on lui affecte la valeur 0. A la fin du programme elle contient la valeur de l'octet.
# - si on entre un bit à 0, on ne change pas la valeur de la variable *octet*,
# - si on entre un bit à 1, on augmente *octet* de la valeur 2 à la puissance n, où n est le rang du bit (n compris entre 0 et 7)
#
# Commençons par simuler l'entrée d'un seul bit (le bit'n') mis à 0 ou à 1 avec les touches A ou B du clavier du PC (attention : à taper en **majuscules**). Pour cela on va utiliser dans ce notebook la fonction **input( )** déjà rencontrée dans le notebook **"Interaction avec l'utilisateur"**
#
# *Remarque : * On ne cherchera pas à gérer le cas où un utilisateur se trompe de touche. En effet, sur la carte Micro:Bit, il n'y a que deux boutons poussoirs...le A et le B !
# +
octet = 0
# choisir ci-dessous une valeur de n entre 0 et 7 pour définir le bit sur lequel on veut travailler:
n = ???
bt = input('?')
if bt =='A': # en cas d'appui sur la touche A du clavier
pass # pour l'instant on ne fait rien
elif bt =='B': # sinon, si la touche B a été appuyée alors :
octet = ???
print(octet)
# -
# Si cette portion de programme fonctionne sur un bit quelconque, il reste à finaliser l'algorithme qui permettra de **répéter huit fois** cette portion de code. On démarera avec le bit 7 pour finir avec le bit 0 :
# +
octet = 0
n = 7
while n >= ???:
bt = input('?')
if bt =='A': # en cas d'appui sur la touche A du clavier
n = ???
elif bt =='B': # sinon, si la touche B a été appuyée alors :
octet = octet + ???
n = ???
print(octet)
# -
# ### 3.2- Mise en application
#
# Lorsque ce code fonctionne :
#
# - en classe : ouvrir le logiciel mu-editor,
#
# ou
#
# - à la maison : aller sur le site https://create.withcode.uk/
#
# et en **faire une adaptation** pour qu'il tourne sur la carte BBC Micro:Bit comme énoncé dans le cahier des charges (il faudra prendre en charge l'appui sur le bouton A ou sur le bouton B)
| 1nsi_2_bin_dec_hex.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Normal distribution
#
# Given normal density and distribution functions for N(0,1)
#
# \begin{equation}
# f_X(x)=\frac{1}{\sqrt{2\pi}}e^{-x^2/2}
# \end{equation}
#
# \begin{equation}
# F_X(x)=\int_\infty^x \frac{1}{\sqrt{2\pi}}e^{-u^2/2}du
# \end{equation}
#
# What is the probability that $4.0 < x < 4.5$?
print("Probability from table: %.6f" % (0.999997 - 0.999968))
# +
from numpy import pi, sqrt, exp
from scipy.integrate import quad
fx = lambda x: 1/sqrt(2*pi)*exp(-x**2/2)
prob = quad(fx, 4, 4.5)
print("Probability from integration: %.6f (%.2e)" % prob)
| Example 1.7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## README.md
# %%writefile README.md
Implementation of [Conditional Generative Adversarial Nets](https://arxiv.org/abs/1411.1784) combined with [Unsupervised Representation Learning with Deep Convolutional Generative Adversarial Networks](https://arxiv.org/abs/1511.06434).
# ## print_object.py
# %%writefile cdcgan_module/trainer/print_object.py
def print_obj(function_name, object_name, object_value):
"""Prints enclosing function, object name, and object value.
Args:
function_name: str, name of function.
object_name: str, name of object.
object_value: object, value of passed object.
"""
# pass
print("{}: {} = {}".format(function_name, object_name, object_value))
# ## image_utils.py
# +
# %%writefile cdcgan_module/trainer/image_utils.py
import tensorflow as tf
from .print_object import print_obj
def preprocess_image(image, params):
"""Preprocess image tensor.
Args:
image: tensor, input image with shape
[cur_batch_size, height, width, depth].
params: dict, user passed parameters.
Returns:
Preprocessed image tensor with shape
[cur_batch_size, height, width, depth].
"""
func_name = "preprocess_image"
# Convert from [0, 255] -> [-1.0, 1.0] floats.
image = tf.cast(x=image, dtype=tf.float32) * (2. / 255) - 1.0
print_obj("\n" + func_name, "image", image)
return image
def resize_fake_images(fake_images, params):
"""Resizes fake images to match real image sizes.
Args:
fake_images: tensor, fake images from generator.
params: dict, user passed parameters.
Returns:
Resized image tensor.
"""
func_name = "resize_real_image"
print_obj("\n" + func_name, "fake_images", fake_images)
# Resize fake images to match real image sizes.
resized_fake_images = tf.image.resize(
images=fake_images,
size=[params["height"], params["width"]],
method="nearest",
name="resized_fake_images"
)
print_obj(func_name, "resized_fake_images", resized_fake_images)
return resized_fake_images
# -
# ## input.py
# +
# %%writefile cdcgan_module/trainer/input.py
import tensorflow as tf
from . import image_utils
from .print_object import print_obj
def decode_example(protos, params):
"""Decodes TFRecord file into tensors.
Given protobufs, decode into image and label tensors.
Args:
protos: protobufs from TFRecord file.
params: dict, user passed parameters.
Returns:
Image and label tensors.
"""
func_name = "decode_example"
# Create feature schema map for protos.
features = {
"image_raw": tf.FixedLenFeature(shape=[], dtype=tf.string),
"label": tf.FixedLenFeature(shape=[], dtype=tf.int64)
}
# Parse features from tf.Example.
parsed_features = tf.parse_single_example(
serialized=protos, features=features
)
print_obj("\n" + func_name, "features", features)
# Convert from a scalar string tensor (whose single string has
# length height * width * depth) to a uint8 tensor with shape
# [height * width * depth].
image = tf.decode_raw(
input_bytes=parsed_features["image_raw"], out_type=tf.uint8
)
print_obj(func_name, "image", image)
# Reshape flattened image back into normal dimensions.
image = tf.reshape(
tensor=image,
shape=[params["height"], params["width"], params["depth"]]
)
print_obj(func_name, "image", image)
# Preprocess image.
image = image_utils.preprocess_image(image=image, params=params)
print_obj(func_name, "image", image)
# Convert label from a scalar uint8 tensor to an int32 scalar.
label = tf.cast(x=parsed_features["label"], dtype=tf.int32)
print_obj(func_name, "label", label)
return {"image": image}, label
def read_dataset(filename, mode, batch_size, params):
"""Reads TF Record data using tf.data, doing necessary preprocessing.
Given filename, mode, batch size, and other parameters, read TF Record
dataset using Dataset API, apply necessary preprocessing, and return an
input function to the Estimator API.
Args:
filename: str, file pattern that to read into our tf.data dataset.
mode: The estimator ModeKeys. Can be TRAIN or EVAL.
batch_size: int, number of examples per batch.
params: dict, dictionary of user passed parameters.
Returns:
An input function.
"""
def _input_fn():
"""Wrapper input function used by Estimator API to get data tensors.
Returns:
Batched dataset object of dictionary of feature tensors and label
tensor.
"""
# Create list of files that match pattern.
file_list = tf.gfile.Glob(filename=filename)
# Create dataset from file list.
if params["input_fn_autotune"]:
dataset = tf.data.TFRecordDataset(
filenames=file_list,
num_parallel_reads=tf.contrib.data.AUTOTUNE
)
else:
dataset = tf.data.TFRecordDataset(filenames=file_list)
# Shuffle and repeat if training with fused op.
if mode == tf.estimator.ModeKeys.TRAIN:
dataset = dataset.apply(
tf.contrib.data.shuffle_and_repeat(
buffer_size=50 * batch_size,
count=None # indefinitely
)
)
# Decode CSV file into a features dictionary of tensors, then batch.
if params["input_fn_autotune"]:
dataset = dataset.apply(
tf.contrib.data.map_and_batch(
map_func=lambda x: decode_example(
protos=x,
params=params
),
batch_size=batch_size,
num_parallel_calls=tf.contrib.data.AUTOTUNE
)
)
else:
dataset = dataset.apply(
tf.contrib.data.map_and_batch(
map_func=lambda x: decode_example(
protos=x,
params=params
),
batch_size=batch_size
)
)
# Prefetch data to improve latency.
if params["input_fn_autotune"]:
dataset = dataset.prefetch(buffer_size=tf.contrib.data.AUTOTUNE)
else:
dataset = dataset.prefetch(buffer_size=1)
# Create a iterator, then get batch of features from example queue.
batched_dataset = dataset.make_one_shot_iterator().get_next()
return batched_dataset
return _input_fn
# -
# ## networks.py
# +
# %%writefile cdcgan_module/trainer/networks.py
import tensorflow as tf
from .print_object import print_obj
class Network(object):
"""Network that could be for generator or discriminator.
Fields:
name: str, name of `Generator` or `Discriminator`.
kernel_regularizer: `l1_l2_regularizer` object, regularizar for kernel
variables.
bias_regularizer: `l1_l2_regularizer` object, regularizar for bias
variables.
"""
def __init__(self, kernel_regularizer, bias_regularizer, name):
"""Instantiates network.
Args:
kernel_regularizer: `l1_l2_regularizer` object, regularizar for
kernel variables.
bias_regularizer: `l1_l2_regularizer` object, regularizar for bias
variables.
name: str, name of generator or discriminator.
"""
# Set name of generator.
self.name = name
# Regularizer for kernel weights.
self.kernel_regularizer = kernel_regularizer
# Regularizer for bias weights.
self.bias_regularizer = bias_regularizer
def embed_labels(self, labels, params, scope):
"""Embeds labels from integer indices to float vectors.
Args:
labels: tensor, labels to condition on of shape
[cur_batch_size, 1].
params: dict, user passed parameters.
scope: str, variable scope.
Returns:
Embedded labels tensor of shape
[cur_batch_size, label_embedding_dimension].
"""
func_name = "{}_embed_labels".format(scope)
with tf.variable_scope(name_or_scope=scope, reuse=tf.AUTO_REUSE):
# Create trainable label embedding matrix.
label_embedding_matrix = tf.get_variable(
name="label_embedding_matrix",
shape=[
params["num_classes"],
params["label_embedding_dimension"]
],
dtype=tf.float32,
initializer=None,
regularizer=None,
trainable=True
)
# Get embedding vectors for integer label index.
label_embeddings = tf.nn.embedding_lookup(
params=label_embedding_matrix,
ids=labels,
name="embedding_lookup"
)
# Flatten back into a rank 2 tensor.
label_vectors = tf.reshape(
tensor=label_embeddings,
shape=[-1, params["label_embedding_dimension"]],
name="label_vectors"
)
print_obj(func_name, "label_vectors", label_vectors)
return label_vectors
def use_labels(self, features, labels, params, scope):
"""Conditions features using label data.
Args:
features: tensor, features tensor, either Z for generator or X for
discriminator.
labels: tensor, labels to condition on of shape
[cur_batch_size, 1].
params: dict, user passed parameters.
scope: str, variable scope.
Returns:
Feature tensor conditioned on labels.
"""
func_name = "{}_use_labels".format(scope)
with tf.variable_scope(name_or_scope=scope, reuse=tf.AUTO_REUSE):
if params["{}_embed_labels".format(scope)]:
label_vectors = self.embed_labels(
labels=labels, params=params, scope=scope
)
else:
label_vectors = tf.one_hot(
indices=tf.squeeze(input=labels, axis=-1),
depth=params["num_classes"],
axis=-1,
name="label_vectors_one_hot"
)
print_obj(func_name, "label_vectors", label_vectors)
if params["{}_concatenate_labels".format(scope)]:
if scope == "generator":
height = params["generator_projection_dims"][0]
width = params["generator_projection_dims"][1]
else:
height = params["height"]
width = params["width"]
# Project labels into image size dimensions.
label_vectors = tf.layers.dense(
inputs=label_vectors,
units=height * width,
activation=None,
kernel_regularizer=self.kernel_regularizer,
bias_regularizer=self.bias_regularizer,
name="labels_dense_concat"
)
# Reshape into an image.
label_image = tf.reshape(
tensor=label_vectors,
shape=[-1, height, width, 1],
name="labels_image_concat"
)
# Concatenate labels & features along feature map dimension.
network = tf.concat(
values=[features, label_image],
axis=-1,
name="features_concat_labels"
)
print_obj(func_name, "network", network)
else:
label_vectors = tf.layers.dense(
inputs=label_vectors,
units=params["latent_size"],
activation=None,
kernel_regularizer=self.kernel_regularizer,
bias_regularizer=self.bias_regularizer,
name="labels_dense_multiply"
)
print_obj(func_name, "label_vectors", label_vectors)
if scope == "generator":
height = params["generator_projection_dims"][0]
width = params["generator_projection_dims"][1]
depth = params["generator_projection_dims"][2]
else:
height = params["height"]
width = params["width"]
depth = params["depth"]
# Project labels into image size dimensions.
label_vectors = tf.layers.dense(
inputs=label_vectors,
units=height * width * depth,
activation=None,
kernel_regularizer=self.kernel_regularizer,
bias_regularizer=self.bias_regularizer,
name="labels_dense_multiply"
)
# Reshape into an image.
label_image = tf.reshape(
tensor=label_vectors,
shape=[-1, height, width, depth],
name="labels_image_multiply"
)
# Element-wise multiply label vectors with latent vectors.
network = tf.multiply(
x=features, y=label_image, name="features_multiply_labels"
)
print_obj(func_name, "network", network)
return network
# -
# ## generator.py
# +
# %%writefile cdcgan_module/trainer/generator.py
import tensorflow as tf
from . import networks
from .print_object import print_obj
class Generator(networks.Network):
"""Generator that takes latent vector input and outputs image.
"""
def __init__(self, kernel_regularizer, bias_regularizer, name):
"""Instantiates and builds generator network.
Args:
kernel_regularizer: `l1_l2_regularizer` object, regularizar for
kernel variables.
bias_regularizer: `l1_l2_regularizer` object, regularizar for bias
variables.
name: str, name of generator.
"""
# Initialize base class.
super().__init__(kernel_regularizer, bias_regularizer, name)
def get_fake_images(self, Z, labels, mode, params):
"""Creates generator network and returns generated images.
Args:
Z: tensor, latent vectors of shape [cur_batch_size, latent_size].
labels: tensor, labels to condition on of shape
[cur_batch_size, 1].
mode: tf.estimator.ModeKeys with values of either TRAIN or EVAL.
params: dict, user passed parameters.
Returns:
Generated image tensor of shape
[cur_batch_size, height * width * depth].
"""
func_name = "get_fake_images"
print_obj("\n" + func_name, "Z", Z)
print_obj(func_name, "labels", labels)
# Dictionary containing possible final activations.
final_activation_dict = {
"sigmoid": tf.nn.sigmoid, "relu": tf.nn.relu, "tanh": tf.nn.tanh
}
with tf.variable_scope(
name_or_scope="generator", reuse=tf.AUTO_REUSE):
# Project latent vectors.
projection_height = params["generator_projection_dims"][0]
projection_width = params["generator_projection_dims"][1]
projection_depth = params["generator_projection_dims"][2]
# shape = (
# cur_batch_size,
# projection_height * projection_width * projection_depth
# )
projection = tf.layers.dense(
inputs=Z,
units=projection_height * projection_width * projection_depth,
activation=None,
kernel_regularizer=self.kernel_regularizer,
bias_regularizer=self.bias_regularizer,
name="projection_dense_layer"
)
print_obj(func_name, "projection", projection)
projection_leaky_relu = tf.nn.leaky_relu(
features=projection,
alpha=params["generator_leaky_relu_alpha"],
name="projection_leaky_relu"
)
print_obj(
func_name, "projection_leaky_relu", projection_leaky_relu
)
# Add batch normalization to keep the inputs from blowing up.
# shape = (
# cur_batch_size,
# projection_height * projection_width * projection_depth
# )
projection_batch_norm = tf.layers.batch_normalization(
inputs=projection_leaky_relu,
training=(mode == tf.estimator.ModeKeys.TRAIN),
name="projection_batch_norm"
)
print_obj(
func_name, "projection_batch_norm", projection_batch_norm
)
# Reshape projection into "image".
# shape = (
# cur_batch_size,
# projection_height,
# projection_width,
# projection_depth
# )
projection_batch_norm_image = tf.reshape(
tensor=projection_batch_norm,
shape=[
-1, projection_height, projection_width, projection_depth
],
name="projection_reshaped"
)
print_obj(
func_name,
"projection_batch_norm_image",
projection_batch_norm_image
)
# Condition on labels.
if params["generator_use_labels"]:
network = self.use_labels(
features=projection_batch_norm_image,
labels=labels,
params=params,
scope="generator"
)
else:
network = projection_batch_norm_image
# Iteratively build upsampling layers.
for i in range(len(params["generator_num_filters"])):
# Add conv transpose layers with given params per layer.
# shape = (
# cur_batch_size,
# generator_kernel_sizes[i - 1] * generator_strides[i],
# generator_kernel_sizes[i - 1] * generator_strides[i],
# generator_num_filters[i]
# )
network = tf.layers.conv2d_transpose(
inputs=network,
filters=params["generator_num_filters"][i],
kernel_size=params["generator_kernel_sizes"][i],
strides=params["generator_strides"][i],
padding="same",
activation=None,
kernel_regularizer=self.kernel_regularizer,
bias_regularizer=self.bias_regularizer,
name="layers_conv2d_tranpose_{}".format(i)
)
print_obj(func_name, "network", network)
network = tf.nn.leaky_relu(
features=network,
alpha=params["generator_leaky_relu_alpha"],
name="leaky_relu_{}".format(i)
)
print_obj(func_name, "network", network)
# Add batch normalization to keep the inputs from blowing up.
network = tf.layers.batch_normalization(
inputs=network,
training=(mode == tf.estimator.ModeKeys.TRAIN),
name="layers_batch_norm_{}".format(i)
)
print_obj(func_name, "network", network)
# Final conv2d transpose layer for image output.
# shape = (cur_batch_size, height, width, depth)
fake_images = tf.layers.conv2d_transpose(
inputs=network,
filters=params["generator_final_num_filters"],
kernel_size=params["generator_final_kernel_size"],
strides=params["generator_final_stride"],
padding="same",
activation=final_activation_dict.get(
params["generator_final_activation"].lower(), None
),
kernel_regularizer=self.kernel_regularizer,
bias_regularizer=self.bias_regularizer,
name="layers_conv2d_tranpose_fake_images"
)
print_obj(func_name, "fake_images", fake_images)
return fake_images
def get_generator_loss(self, fake_logits):
"""Gets generator loss.
Args:
fake_logits: tensor, shape of
[cur_batch_size, 1].
Returns:
Tensor of generator's total loss of shape [].
"""
func_name = "get_generator_loss"
# Calculate base generator loss.
generator_loss = tf.reduce_mean(
input_tensor=tf.nn.sigmoid_cross_entropy_with_logits(
logits=fake_logits,
labels=tf.ones_like(tensor=fake_logits)
),
name="generator_loss"
)
print_obj("\n" + func_name, "generator_loss", generator_loss)
# Get regularization losses.
generator_reg_loss = tf.losses.get_regularization_loss(
scope="generator",
name="generator_regularization_loss"
)
print_obj(func_name, "generator_reg_loss", generator_reg_loss)
# Combine losses for total losses.
generator_total_loss = tf.math.add(
x=generator_loss,
y=generator_reg_loss,
name="generator_total_loss"
)
print_obj(func_name, "generator_total_loss", generator_total_loss)
# Add summaries for TensorBoard.
tf.summary.scalar(
name="generator_loss", tensor=generator_loss, family="losses"
)
tf.summary.scalar(
name="generator_reg_loss",
tensor=generator_reg_loss,
family="losses"
)
tf.summary.scalar(
name="generator_total_loss",
tensor=generator_total_loss,
family="total_losses"
)
return generator_total_loss
# -
# ## discriminator.py
# +
# %%writefile cdcgan_module/trainer/discriminator.py
import tensorflow as tf
from . import networks
from .print_object import print_obj
class Discriminator(networks.Network):
"""Discriminator that takes image input and outputs logits.
"""
def __init__(self, kernel_regularizer, bias_regularizer, name):
"""Instantiates discriminator network.
Args:
kernel_regularizer: `l1_l2_regularizer` object, regularizar for
kernel variables.
bias_regularizer: `l1_l2_regularizer` object, regularizar for bias
variables.
name: str, name of discriminator.
"""
# Initialize base class.
super().__init__(kernel_regularizer, bias_regularizer, name)
def get_discriminator_logits(self, X, labels, params):
"""Creates discriminator network and returns logits.
Args:
X: tensor, image tensors of shape
[cur_batch_size, height * width * depth].
labels: tensor, labels to condition on of shape
[cur_batch_size, 1].
params: dict, user passed parameters.
Returns:
Logits tensor of shape [cur_batch_size, 1].
"""
func_name = "get_discriminator_logits"
print_obj("\n" + func_name, "X", X)
print_obj(func_name, "labels", labels)
with tf.variable_scope(
name_or_scope="discriminator", reuse=tf.AUTO_REUSE):
# Condition on labels.
if params["discriminator_use_labels"]:
network = self.use_labels(
features=X,
labels=labels,
params=params,
scope="discriminator"
)
else:
network = X
# Iteratively build downsampling layers.
for i in range(len(params["discriminator_num_filters"])):
# Add convolutional layers with given params per layer.
# shape = (
# cur_batch_size,
# discriminator_kernel_sizes[i - 1] / discriminator_strides[i],
# discriminator_kernel_sizes[i - 1] / discriminator_strides[i],
# discriminator_num_filters[i]
# )
network = tf.layers.conv2d(
inputs=network,
filters=params["discriminator_num_filters"][i],
kernel_size=params["discriminator_kernel_sizes"][i],
strides=params["discriminator_strides"][i],
padding="same",
activation=None,
kernel_regularizer=self.kernel_regularizer,
bias_regularizer=self.bias_regularizer,
name="layers_conv2d_{}".format(i)
)
print_obj(func_name, "network", network)
network = tf.nn.leaky_relu(
features=network,
alpha=params["discriminator_leaky_relu_alpha"],
name="leaky_relu_{}".format(i)
)
print_obj(func_name, "network", network)
# Add some dropout for better regularization and stability.
network = tf.layers.dropout(
inputs=network,
rate=params["discriminator_dropout_rates"][i],
name="layers_dropout_{}".format(i)
)
print_obj(func_name, "network", network)
# Flatten network output.
# shape = (
# cur_batch_size,
# (discriminator_kernel_sizes[-2] / discriminator_strides[-1]) ** 2 * discriminator_num_filters[-1]
# )
network_flat = tf.layers.Flatten()(inputs=network)
print_obj(func_name, "network_flat", network_flat)
# Final linear layer for logits.
# shape = (cur_batch_size, 1)
logits = tf.layers.dense(
inputs=network_flat,
units=1,
activation=None,
kernel_regularizer=self.kernel_regularizer,
bias_regularizer=self.bias_regularizer,
name="layers_dense_logits"
)
print_obj(func_name, "logits", logits)
return logits
def get_discriminator_loss(self, fake_logits, real_logits, params):
"""Gets discriminator loss.
Args:
fake_logits: tensor, shape of [cur_batch_size, 1].
real_logits: tensor, shape of [cur_batch_size, 1].
params: dict, user passed parameters.
Returns:
Tensor of discriminator's total loss of shape [].
"""
func_name = "get_discriminator_loss"
# Calculate base discriminator loss.
discriminator_real_loss = tf.reduce_mean(
input_tensor=tf.nn.sigmoid_cross_entropy_with_logits(
logits=real_logits,
labels=tf.multiply(
x=tf.ones_like(tensor=real_logits),
y=params["label_smoothing"]
)
),
name="discriminator_real_loss"
)
print_obj(
"\n" + func_name,
"discriminator_real_loss",
discriminator_real_loss
)
discriminator_fake_loss = tf.reduce_mean(
input_tensor=tf.nn.sigmoid_cross_entropy_with_logits(
logits=fake_logits,
labels=tf.zeros_like(tensor=fake_logits)
),
name="discriminator_fake_loss"
)
print_obj(
func_name, "discriminator_fake_loss", discriminator_fake_loss
)
discriminator_loss = tf.add(
x=discriminator_real_loss,
y=discriminator_fake_loss,
name="discriminator_loss"
)
print_obj(func_name, "discriminator_loss", discriminator_loss)
# Get regularization losses.
discriminator_reg_loss = tf.losses.get_regularization_loss(
scope="discriminator",
name="discriminator_reg_loss"
)
print_obj(func_name, "discriminator_reg_loss", discriminator_reg_loss)
# Combine losses for total losses.
discriminator_total_loss = tf.math.add(
x=discriminator_loss,
y=discriminator_reg_loss,
name="discriminator_total_loss"
)
print_obj(
func_name, "discriminator_total_loss", discriminator_total_loss
)
# Add summaries for TensorBoard.
tf.summary.scalar(
name="discriminator_real_loss",
tensor=discriminator_real_loss,
family="losses"
)
tf.summary.scalar(
name="discriminator_fake_loss",
tensor=discriminator_fake_loss,
family="losses"
)
tf.summary.scalar(
name="discriminator_loss",
tensor=discriminator_loss,
family="losses"
)
tf.summary.scalar(
name="discriminator_reg_loss",
tensor=discriminator_reg_loss,
family="losses"
)
tf.summary.scalar(
name="discriminator_total_loss",
tensor=discriminator_total_loss,
family="total_losses"
)
return discriminator_total_loss
# -
# ## train_and_eval.py
# +
# %%writefile cdcgan_module/trainer/train_and_eval.py
import tensorflow as tf
from . import image_utils
from .print_object import print_obj
def get_logits_and_losses(
features, labels, generator, discriminator, mode, params):
"""Gets logits and losses for both train and eval modes.
Args:
features: dict, feature tensors from input function.
labels: tensor, labels to condition on of shape
[cur_batch_size, 1].
generator: instance of generator.`Generator`.
discriminator: instance of discriminator.`Discriminator`.
mode: tf.estimator.ModeKeys with values of either TRAIN or EVAL.
params: dict, user passed parameters.
Returns:
Real and fake logits and generator and discriminator losses.
"""
func_name = "get_logits_and_losses"
# For training discriminator.
print("\nTraining discriminator.")
# Extract real images from features dictionary.
real_images = features["image"]
print_obj("\n" + func_name, "real_images", real_images)
# Get dynamic batch size in case of partial batch.
cur_batch_size = tf.shape(
input=real_images,
out_type=tf.int32,
name="{}_cur_batch_size".format(func_name)
)[0]
# Create random noise latent vector for each batch example.
Z = tf.random.normal(
shape=[cur_batch_size, params["latent_size"]],
mean=0.0,
stddev=1.0,
dtype=tf.float32,
name="discriminator_Z"
)
print_obj(func_name, "Z", Z)
# Get generated image from generator network from gaussian noise.
print("\nCall generator with Z = {}.".format(Z))
fake_images = generator.get_fake_images(
Z=Z, labels=labels, mode=mode, params=params
)
print_obj(func_name, "fake_images", fake_images)
# Resize fake images to match real image sizes.
fake_images = image_utils.resize_fake_images(fake_images, params)
print_obj(func_name, "fake_images", fake_images)
# Get fake logits from discriminator using generator's output image.
print("\nCall discriminator with fake_images = {}.".format(fake_images))
fake_logits = discriminator.get_discriminator_logits(
X=fake_images, labels=labels, params=params
)
print_obj(func_name, "fake_logits", fake_logits)
# Get real logits from discriminator using real image.
print(
"\nCall discriminator with real_images = {}.".format(real_images)
)
real_logits = discriminator.get_discriminator_logits(
X=real_images, labels=labels, params=params
)
print_obj(func_name, "real_logits", real_logits)
# Get discriminator total loss.
discriminator_total_loss = discriminator.get_discriminator_loss(
fake_logits=fake_logits, real_logits=real_logits, params=params
)
print_obj(func_name, "discriminator_total_loss", discriminator_total_loss)
##########################################################################
##########################################################################
##########################################################################
# For training generator.
print("\nTraining generator.")
# Create random noise latent vector for each batch example.
fake_Z = tf.random.normal(
shape=[cur_batch_size, params["latent_size"]],
mean=0.0,
stddev=1.0,
dtype=tf.float32,
name="generator_Z"
)
print_obj(func_name, "fake_Z", fake_Z)
# Create random (fake) labels.
fake_labels = tf.random.uniform(
shape=[cur_batch_size, 1],
minval=0,
maxval=params["num_classes"],
dtype=tf.int32,
name="fake_labels"
)
print_obj(func_name, "fake_labels", fake_labels)
# Get generated image from generator network from gaussian noise.
print("\nCall generator with fake_Z = {}.".format(fake_Z))
fake_fake_images = generator.get_fake_images(
Z=fake_Z, labels=fake_labels, mode=mode, params=params
)
print_obj(func_name, "fake_fake_images", fake_fake_images)
# Get fake logits from discriminator using generator's output image.
print(
"\nCall discriminator with fake_fake_images = {}.".format(
fake_fake_images
)
)
fake_fake_logits = discriminator.get_discriminator_logits(
X=fake_fake_images, labels=fake_labels, params=params
)
print_obj(func_name, "fake_fake_logits", fake_fake_logits)
# Get generator total loss.
generator_total_loss = generator.get_generator_loss(
fake_logits=fake_fake_logits
)
print_obj(func_name, "generator_total_loss", generator_total_loss)
# Add summaries for TensorBoard.
tf.summary.image(
name="fake_images",
tensor=image_utils.resize_fake_images(
fake_images=generator.get_fake_images(
Z=tf.random.normal(
shape=[params["num_classes"], params["latent_size"]],
mean=0.0,
stddev=1.0,
dtype=tf.float32,
name="image_summary_Z"
),
labels=tf.expand_dims(
input=tf.range(
start=0, limit=params["num_classes"], dtype=tf.int32
),
axis=-1,
name="image_summary_fake_labels"
),
mode=tf.estimator.ModeKeys.PREDICT,
params=params
),
params=params
),
max_outputs=params["num_classes"],
)
return (real_logits,
fake_logits,
generator_total_loss,
discriminator_total_loss)
# -
# ## train.py
# +
# %%writefile cdcgan_module/trainer/train.py
import tensorflow as tf
from .print_object import print_obj
def get_variables_and_gradients(loss, scope):
"""Gets variables and their gradients wrt. loss.
Args:
loss: tensor, shape of [].
scope: str, the network's name to find its variables to train.
Returns:
Lists of variables and their gradients.
"""
func_name = "get_variables_and_gradients"
# Get trainable variables.
variables = tf.trainable_variables(scope=scope)
print_obj("\n{}_{}".format(func_name, scope), "variables", variables)
# Get gradients.
gradients = tf.gradients(
ys=loss,
xs=variables,
name="{}_gradients".format(scope)
)
print_obj("\n{}_{}".format(func_name, scope), "gradients", gradients)
# Add variable names back in for identification.
gradients = [
tf.identity(
input=g,
name="{}_{}_gradients".format(func_name, v.name[:-2])
)
if tf.is_tensor(x=g) else g
for g, v in zip(gradients, variables)
]
print_obj("\n{}_{}".format(func_name, scope), "gradients", gradients)
return variables, gradients
def create_variable_and_gradient_histogram_summaries(loss_dict, params):
"""Creates variable and gradient histogram summaries.
Args:
loss_dict: dict, keys are scopes and values are scalar loss tensors
for each network kind.
params: dict, user passed parameters.
"""
for scope, loss in loss_dict.items():
# Get variables and their gradients wrt. loss.
variables, gradients = get_variables_and_gradients(loss, scope)
# Add summaries for TensorBoard.
for g, v in zip(gradients, variables):
tf.summary.histogram(
name="{}".format(v.name[:-2]),
values=v,
family="{}_variables".format(scope)
)
if tf.is_tensor(x=g):
tf.summary.histogram(
name="{}".format(v.name[:-2]),
values=g,
family="{}_gradients".format(scope)
)
def train_network(loss, global_step, params, scope):
"""Trains network and returns loss and train op.
Args:
loss: tensor, shape of [].
global_step: tensor, the current training step or batch in the
training loop.
params: dict, user passed parameters.
scope: str, the variables that to train.
Returns:
Loss tensor and training op.
"""
func_name = "train_network"
print_obj("\n" + func_name, "scope", scope)
# Create optimizer map.
optimizers = {
"Adam": tf.train.AdamOptimizer,
"Adadelta": tf.train.AdadeltaOptimizer,
"AdagradDA": tf.train.AdagradDAOptimizer,
"Adagrad": tf.train.AdagradOptimizer,
"Ftrl": tf.train.FtrlOptimizer,
"GradientDescent": tf.train.GradientDescentOptimizer,
"Momentum": tf.train.MomentumOptimizer,
"ProximalAdagrad": tf.train.ProximalAdagradOptimizer,
"ProximalGradientDescent": tf.train.ProximalGradientDescentOptimizer,
"RMSProp": tf.train.RMSPropOptimizer
}
# Get optimizer and instantiate it.
if params["{}_optimizer".format(scope)] == "Adam":
optimizer = optimizers[params["{}_optimizer".format(scope)]](
learning_rate=params["{}_learning_rate".format(scope)],
beta1=params["{}_adam_beta1".format(scope)],
beta2=params["{}_adam_beta2".format(scope)],
epsilon=params["{}_adam_epsilon".format(scope)],
name="{}_{}_optimizer".format(
scope, params["{}_optimizer".format(scope)].lower()
)
)
else:
optimizer = optimizers[params["{}_optimizer".format(scope)]](
learning_rate=params["{}_learning_rate".format(scope)],
name="{}_{}_optimizer".format(
scope, params["{}_optimizer".format(scope)].lower()
)
)
print_obj("{}_{}".format(func_name, scope), "optimizer", optimizer)
# Get gradients.
gradients = tf.gradients(
ys=loss,
xs=tf.trainable_variables(scope=scope),
name="{}_gradients".format(scope)
)
print_obj("\n{}_{}".format(func_name, scope), "gradients", gradients)
# Clip gradients.
if params["{}_clip_gradients".format(scope)]:
gradients, _ = tf.clip_by_global_norm(
t_list=gradients,
clip_norm=params["{}_clip_gradients".format(scope)],
name="{}_clip_by_global_norm_gradients".format(scope)
)
print_obj("\n{}_{}".format(func_name, scope), "gradients", gradients)
# Zip back together gradients and variables.
grads_and_vars = zip(gradients, tf.trainable_variables(scope=scope))
print_obj(
"{}_{}".format(func_name, scope), "grads_and_vars", grads_and_vars
)
# Create train op by applying gradients to variables and incrementing
# global step.
train_op = optimizer.apply_gradients(
grads_and_vars=grads_and_vars,
global_step=global_step,
name="{}_apply_gradients".format(scope)
)
return loss, train_op
def get_loss_and_train_op(
generator_total_loss, discriminator_total_loss, params):
"""Gets loss and train op for train mode.
Args:
generator_total_loss: tensor, scalar total loss of generator.
discriminator_total_loss: tensor, scalar total loss of discriminator.
params: dict, user passed parameters.
Returns:
Loss scalar tensor and train_op to be used by the EstimatorSpec.
"""
func_name = "get_loss_and_train_op"
# Get global step.
global_step = tf.train.get_or_create_global_step()
# Determine if it is time to train generator or discriminator.
cycle_step = tf.mod(
x=global_step,
y=tf.cast(
x=tf.add(
x=params["discriminator_train_steps"],
y=params["generator_train_steps"]
),
dtype=tf.int64
),
name="{}_cycle_step".format(func_name)
)
# Create choose discriminator condition.
condition = tf.less(
x=cycle_step, y=params["discriminator_train_steps"]
)
# Needed for batch normalization, but has no effect otherwise.
update_ops = tf.get_collection(key=tf.GraphKeys.UPDATE_OPS)
# Ensure update ops get updated.
with tf.control_dependencies(control_inputs=update_ops):
# Conditionally choose to train generator or discriminator subgraph.
loss, train_op = tf.cond(
pred=condition,
true_fn=lambda: train_network(
loss=discriminator_total_loss,
global_step=global_step,
params=params,
scope="discriminator"
),
false_fn=lambda: train_network(
loss=generator_total_loss,
global_step=global_step,
params=params,
scope="generator"
)
)
return loss, train_op
# -
# ## eval_metrics.py
# +
# %%writefile cdcgan_module/trainer/eval_metrics.py
import tensorflow as tf
from .print_object import print_obj
def get_eval_metric_ops(fake_logits, real_logits, params):
"""Gets eval metric ops.
Args:
fake_logits: tensor, shape of [cur_batch_size, 1] that came from
discriminator having processed generator's output image.
real_logits: tensor, shape of [cur_batch_size, 1] that came from
discriminator having processed real image.
params: dict, user passed parameters.
Returns:
Dictionary of eval metric ops.
"""
func_name = "get_eval_metric_ops"
# Concatenate discriminator logits and labels.
discriminator_logits = tf.concat(
values=[real_logits, fake_logits],
axis=0,
name="discriminator_concat_logits"
)
print_obj("\n" + func_name, "discriminator_logits", discriminator_logits)
discriminator_labels = tf.concat(
values=[
tf.ones_like(tensor=real_logits) * params["label_smoothing"],
tf.zeros_like(tensor=fake_logits)
],
axis=0,
name="discriminator_concat_labels"
)
print_obj(func_name, "discriminator_labels", discriminator_labels)
# Calculate discriminator probabilities.
discriminator_probabilities = tf.nn.sigmoid(
x=discriminator_logits, name="discriminator_probabilities"
)
print_obj(
func_name, "discriminator_probabilities", discriminator_probabilities
)
# Create eval metric ops dictionary.
eval_metric_ops = {
"accuracy": tf.metrics.accuracy(
labels=discriminator_labels,
predictions=discriminator_probabilities,
name="discriminator_accuracy"
),
"precision": tf.metrics.precision(
labels=discriminator_labels,
predictions=discriminator_probabilities,
name="discriminator_precision"
),
"recall": tf.metrics.recall(
labels=discriminator_labels,
predictions=discriminator_probabilities,
name="discriminator_recall"
),
"auc_roc": tf.metrics.auc(
labels=discriminator_labels,
predictions=discriminator_probabilities,
num_thresholds=200,
curve="ROC",
name="discriminator_auc_roc"
),
"auc_pr": tf.metrics.auc(
labels=discriminator_labels,
predictions=discriminator_probabilities,
num_thresholds=200,
curve="PR",
name="discriminator_auc_pr"
)
}
print_obj(func_name, "eval_metric_ops", eval_metric_ops)
return eval_metric_ops
# -
# ## predict.py
# +
# %%writefile cdcgan_module/trainer/predict.py
import tensorflow as tf
from . import image_utils
from .print_object import print_obj
def get_predictions_and_export_outputs(features, generator, params):
"""Gets predictions and serving export outputs.
Args:
features: dict, feature tensors from serving input function.
generator: instance of `Generator`.
params: dict, user passed parameters.
Returns:
Predictions dictionary and export outputs dictionary.
"""
func_name = "get_predictions_and_export_outputs"
# Extract given latent vectors from features dictionary.
Z = features["Z"]
print_obj("\n" + func_name, "Z", Z)
# Extract labels from features dictionary & expand from vector to matrix.
labels = tf.expand_dims(input=features["label"], axis=-1)
print_obj(func_name, "labels", labels)
# Get generated images from generator using latent vector.
generated_images = generator.get_fake_images(
Z=Z, labels=labels, mode=tf.estimator.ModeKeys.PREDICT, params=params
)
print_obj(func_name, "generated_images", generated_images)
# Resize generated images to match real image sizes.
generated_images = image_utils.resize_fake_images(
fake_images=generated_images, params=params
)
print_obj(func_name, "generated_images", generated_images)
# Create predictions dictionary.
predictions_dict = {
"generated_images": generated_images
}
print_obj(func_name, "predictions_dict", predictions_dict)
# Create export outputs.
export_outputs = {
"predict_export_outputs": tf.estimator.export.PredictOutput(
outputs=predictions_dict)
}
print_obj(func_name, "export_outputs", export_outputs)
return predictions_dict, export_outputs
# -
# ## cdcgan.py
# +
# %%writefile cdcgan_module/trainer/cdcgan.py
import tensorflow as tf
from . import discriminator
from . import eval_metrics
from . import generator
from . import predict
from . import train
from . import train_and_eval
from .print_object import print_obj
def cdcgan_model(features, labels, mode, params):
"""Conditional Deep Convolutional GAN custom Estimator model function.
Args:
features: dict, keys are feature names and values are feature tensors.
labels: tensor, label data.
mode: tf.estimator.ModeKeys with values of either TRAIN, EVAL, or
PREDICT.
params: dict, user passed parameters.
Returns:
Instance of `tf.estimator.EstimatorSpec` class.
"""
func_name = "cdcgan_model"
print_obj("\n" + func_name, "features", features)
print_obj(func_name, "labels", labels)
print_obj(func_name, "mode", mode)
print_obj(func_name, "params", params)
# Loss function, training/eval ops, etc.
predictions_dict = None
loss = None
train_op = None
eval_metric_ops = None
export_outputs = None
# Instantiate generator.
cdcgan_generator = generator.Generator(
kernel_regularizer=tf.contrib.layers.l1_l2_regularizer(
scale_l1=params["generator_l1_regularization_scale"],
scale_l2=params["generator_l2_regularization_scale"]
),
bias_regularizer=None,
name="generator"
)
# Instantiate discriminator.
cdcgan_discriminator = discriminator.Discriminator(
kernel_regularizer=tf.contrib.layers.l1_l2_regularizer(
scale_l1=params["discriminator_l1_regularization_scale"],
scale_l2=params["discriminator_l2_regularization_scale"]
),
bias_regularizer=None,
name="discriminator"
)
if mode == tf.estimator.ModeKeys.PREDICT:
# Get predictions and export outputs.
(predictions_dict,
export_outputs) = predict.get_predictions_and_export_outputs(
features=features, generator=cdcgan_generator, params=params
)
else:
# Expand labels from vector to matrix.
labels = tf.expand_dims(input=labels, axis=-1)
print_obj(func_name, "labels", labels)
# Get logits and losses from networks for train and eval modes.
(real_logits,
fake_logits,
generator_total_loss,
discriminator_total_loss) = train_and_eval.get_logits_and_losses(
features=features,
labels=labels,
generator=cdcgan_generator,
discriminator=cdcgan_discriminator,
mode=mode,
params=params
)
if mode == tf.estimator.ModeKeys.TRAIN:
# Create variable and gradient histogram summaries.
train.create_variable_and_gradient_histogram_summaries(
loss_dict={
"generator": generator_total_loss,
"discriminator": discriminator_total_loss
},
params=params
)
# Get loss and train op for EstimatorSpec.
loss, train_op = train.get_loss_and_train_op(
generator_total_loss=generator_total_loss,
discriminator_total_loss=discriminator_total_loss,
params=params
)
else:
# Set eval loss.
loss = discriminator_total_loss
# Get eval metrics.
eval_metric_ops = eval_metrics.get_eval_metric_ops(
real_logits=real_logits,
fake_logits=fake_logits,
params=params
)
# Return EstimatorSpec
return tf.estimator.EstimatorSpec(
mode=mode,
predictions=predictions_dict,
loss=loss,
train_op=train_op,
eval_metric_ops=eval_metric_ops,
export_outputs=export_outputs
)
# -
# ## serving.py
# +
# %%writefile cdcgan_module/trainer/serving.py
import tensorflow as tf
from .print_object import print_obj
def serving_input_fn(params):
"""Serving input function.
Args:
params: dict, user passed parameters.
Returns:
ServingInputReceiver object containing features and receiver tensors.
"""
func_name = "serving_input_fn"
# Create placeholders to accept data sent to the model at serving time.
# shape = (batch_size,)
feature_placeholders = {
"Z": tf.placeholder(
dtype=tf.float32,
shape=[None, params["latent_size"]],
name="serving_input_placeholder_Z"
),
"label": tf.placeholder(
dtype=tf.int32,
shape=[None],
name="serving_input_placeholder_label"
)
}
print_obj("\n" + func_name, "feature_placeholders", feature_placeholders)
# Create clones of the feature placeholder tensors so that the SavedModel
# SignatureDef will point to the placeholder.
features = {
key: tf.identity(
input=value,
name="{}_identity_placeholder_{}".format(func_name, key)
)
for key, value in feature_placeholders.items()
}
print_obj(func_name, "features", features)
return tf.estimator.export.ServingInputReceiver(
features=features, receiver_tensors=feature_placeholders
)
# -
# ## model.py
# +
# %%writefile cdcgan_module/trainer/model.py
import tensorflow as tf
from . import input
from . import serving
from . import cdcgan
from .print_object import print_obj
def train_and_evaluate(args):
"""Trains and evaluates custom Estimator model.
Args:
args: dict, user passed parameters.
Returns:
`Estimator` object.
"""
func_name = "train_and_evaluate"
print_obj("\n" + func_name, "args", args)
# Ensure filewriter cache is clear for TensorBoard events file.
tf.summary.FileWriterCache.clear()
# Set logging to be level of INFO.
tf.logging.set_verbosity(tf.logging.INFO)
# Create a RunConfig for Estimator.
config = tf.estimator.RunConfig(
model_dir=args["output_dir"],
save_summary_steps=args["save_summary_steps"],
save_checkpoints_steps=args["save_checkpoints_steps"],
keep_checkpoint_max=args["keep_checkpoint_max"]
)
# Create our custom estimator using our model function.
estimator = tf.estimator.Estimator(
model_fn=cdcgan.cdcgan_model,
model_dir=args["output_dir"],
config=config,
params=args
)
# Create train spec to read in our training data.
train_spec = tf.estimator.TrainSpec(
input_fn=input.read_dataset(
filename=args["train_file_pattern"],
mode=tf.estimator.ModeKeys.TRAIN,
batch_size=args["train_batch_size"],
params=args
),
max_steps=args["train_steps"]
)
# Create exporter to save out the complete model to disk.
exporter = tf.estimator.LatestExporter(
name="exporter",
serving_input_receiver_fn=lambda: serving.serving_input_fn(args)
)
# Create eval spec to read in our validation data and export our model.
eval_spec = tf.estimator.EvalSpec(
input_fn=input.read_dataset(
filename=args["eval_file_pattern"],
mode=tf.estimator.ModeKeys.EVAL,
batch_size=args["eval_batch_size"],
params=args
),
steps=args["eval_steps"],
start_delay_secs=args["start_delay_secs"],
throttle_secs=args["throttle_secs"],
exporters=exporter
)
# Create train and evaluate loop to train and evaluate our estimator.
tf.estimator.train_and_evaluate(
estimator=estimator, train_spec=train_spec, eval_spec=eval_spec)
return estimator
# -
# ## task.py
# +
# %%writefile cdcgan_module/trainer/task.py
import argparse
import json
import os
from . import model
def convert_string_to_bool(string):
"""Converts string to bool.
Args:
string: str, string to convert.
Returns:
Boolean conversion of string.
"""
return False if string.lower() == "false" else True
def convert_string_to_none_or_float(string):
"""Converts string to None or float.
Args:
string: str, string to convert.
Returns:
None or float conversion of string.
"""
return None if string.lower() == "none" else float(string)
def convert_string_to_none_or_int(string):
"""Converts string to None or int.
Args:
string: str, string to convert.
Returns:
None or int conversion of string.
"""
return None if string.lower() == "none" else int(string)
def convert_string_to_list_of_ints(string, sep):
"""Converts string to list of ints.
Args:
string: str, string to convert.
sep: str, separator string.
Returns:
List of ints conversion of string.
"""
return [int(x) for x in string.split(sep)]
def convert_string_to_list_of_floats(string, sep):
"""Converts string to list of floats.
Args:
string: str, string to convert.
sep: str, separator string.
Returns:
List of floats conversion of string.
"""
return [float(x) for x in string.split(sep)]
if __name__ == "__main__":
parser = argparse.ArgumentParser()
# File arguments.
parser.add_argument(
"--train_file_pattern",
help="GCS location to read training data.",
required=True
)
parser.add_argument(
"--eval_file_pattern",
help="GCS location to read evaluation data.",
required=True
)
parser.add_argument(
"--output_dir",
help="GCS location to write checkpoints and export models.",
required=True
)
parser.add_argument(
"--job-dir",
help="This model ignores this field, but it is required by gcloud.",
default="junk"
)
# Training parameters.
parser.add_argument(
"--train_batch_size",
help="Number of examples in training batch.",
type=int,
default=32
)
parser.add_argument(
"--train_steps",
help="Number of steps to train for.",
type=int,
default=100
)
parser.add_argument(
"--save_summary_steps",
help="How many steps to train before saving a summary.",
type=int,
default=100
)
parser.add_argument(
"--save_checkpoints_steps",
help="How many steps to train before saving a checkpoint.",
type=int,
default=100
)
parser.add_argument(
"--keep_checkpoint_max",
help="Max number of checkpoints to keep.",
type=int,
default=100
)
parser.add_argument(
"--input_fn_autotune",
help="Whether to autotune input function performance.",
type=str,
default="True"
)
# Eval parameters.
parser.add_argument(
"--eval_batch_size",
help="Number of examples in evaluation batch.",
type=int,
default=32
)
parser.add_argument(
"--eval_steps",
help="Number of steps to evaluate for.",
type=str,
default="None"
)
parser.add_argument(
"--start_delay_secs",
help="Number of seconds to wait before first evaluation.",
type=int,
default=60
)
parser.add_argument(
"--throttle_secs",
help="Number of seconds to wait between evaluations.",
type=int,
default=120
)
# Image parameters.
parser.add_argument(
"--height",
help="Height of image.",
type=int,
default=32
)
parser.add_argument(
"--width",
help="Width of image.",
type=int,
default=32
)
parser.add_argument(
"--depth",
help="Depth of image.",
type=int,
default=3
)
# Label parameters.
parser.add_argument(
"--num_classes",
help="Number of image classes.",
type=int,
default=10
)
parser.add_argument(
"--label_embedding_dimension",
help="Number of dimensions to embed label classes into.",
type=int,
default=3
)
# Generator parameters.
parser.add_argument(
"--latent_size",
help="The latent size of the noise vector.",
type=int,
default=3
)
parser.add_argument(
"--generator_projection_dims",
help="The 3D dimensions to project latent noise vector into.",
type=str,
default="8,8,256"
)
parser.add_argument(
"--generator_use_labels",
help="Whether to condition on labels in generator.",
type=str,
default="True"
)
parser.add_argument(
"--generator_embed_labels",
help="Whether to embed labels in generator.",
type=str,
default="True"
)
parser.add_argument(
"--generator_concatenate_labels",
help="Whether to concatenate labels in generator.",
type=str,
default="True"
)
parser.add_argument(
"--generator_num_filters",
help="Number of filters for generator conv layers.",
type=str,
default="128, 64"
)
parser.add_argument(
"--generator_kernel_sizes",
help="Kernel sizes for generator conv layers.",
type=str,
default="5,5"
)
parser.add_argument(
"--generator_strides",
help="Strides for generator conv layers.",
type=str,
default="1,2"
)
parser.add_argument(
"--generator_final_num_filters",
help="Number of filters for final generator conv layer.",
type=int,
default=3
)
parser.add_argument(
"--generator_final_kernel_size",
help="Kernel sizes for final generator conv layer.",
type=int,
default=5
)
parser.add_argument(
"--generator_final_stride",
help="Strides for final generator conv layer.",
type=int,
default=2
)
parser.add_argument(
"--generator_leaky_relu_alpha",
help="The amount of leakyness of generator's leaky relus.",
type=float,
default=0.2
)
parser.add_argument(
"--generator_final_activation",
help="The final activation function of generator.",
type=str,
default="None"
)
parser.add_argument(
"--generator_l1_regularization_scale",
help="Scale factor for L1 regularization for generator.",
type=float,
default=0.0
)
parser.add_argument(
"--generator_l2_regularization_scale",
help="Scale factor for L2 regularization for generator.",
type=float,
default=0.0
)
parser.add_argument(
"--generator_optimizer",
help="Name of optimizer to use for generator.",
type=str,
default="Adam"
)
parser.add_argument(
"--generator_learning_rate",
help="How quickly we train our model by scaling the gradient for generator.",
type=float,
default=0.001
)
parser.add_argument(
"--generator_adam_beta1",
help="Adam optimizer's beta1 hyperparameter for first moment.",
type=float,
default=0.9
)
parser.add_argument(
"--generator_adam_beta2",
help="Adam optimizer's beta2 hyperparameter for second moment.",
type=float,
default=0.999
)
parser.add_argument(
"--generator_adam_epsilon",
help="Adam optimizer's epsilon hyperparameter for numerical stability.",
type=float,
default=1e-8
)
parser.add_argument(
"--generator_clip_gradients",
help="Global clipping to prevent gradient norm to exceed this value for generator.",
type=str,
default="None"
)
parser.add_argument(
"--generator_train_steps",
help="Number of steps to train generator for per cycle.",
type=int,
default=100
)
# Discriminator parameters.
parser.add_argument(
"--discriminator_use_labels",
help="Whether to condition on labels in discriminator.",
type=str,
default="True"
)
parser.add_argument(
"--discriminator_embed_labels",
help="Whether to embed labels in discriminator.",
type=str,
default="True"
)
parser.add_argument(
"--discriminator_concatenate_labels",
help="Whether to concatenate labels in discriminator.",
type=str,
default="True"
)
parser.add_argument(
"--discriminator_num_filters",
help="Number of filters for discriminator conv layers.",
type=str,
default="64, 128"
)
parser.add_argument(
"--discriminator_kernel_sizes",
help="Kernel sizes for discriminator conv layers.",
type=str,
default="5,5"
)
parser.add_argument(
"--discriminator_strides",
help="Strides for discriminator conv layers.",
type=str,
default="1,2"
)
parser.add_argument(
"--discriminator_dropout_rates",
help="Dropout rates for discriminator dropout layers.",
type=str,
default="0.3,0.3"
)
parser.add_argument(
"--discriminator_leaky_relu_alpha",
help="The amount of leakyness of discriminator's leaky relus.",
type=float,
default=0.2
)
parser.add_argument(
"--discriminator_l1_regularization_scale",
help="Scale factor for L1 regularization for discriminator.",
type=float,
default=0.0
)
parser.add_argument(
"--discriminator_l2_regularization_scale",
help="Scale factor for L2 regularization for discriminator.",
type=float,
default=0.0
)
parser.add_argument(
"--discriminator_optimizer",
help="Name of optimizer to use for discriminator.",
type=str,
default="Adam"
)
parser.add_argument(
"--discriminator_learning_rate",
help="How quickly we train our model by scaling the gradient for discriminator.",
type=float,
default=0.001
)
parser.add_argument(
"--discriminator_adam_beta1",
help="Adam optimizer's beta1 hyperparameter for first moment.",
type=float,
default=0.9
)
parser.add_argument(
"--discriminator_adam_beta2",
help="Adam optimizer's beta2 hyperparameter for second moment.",
type=float,
default=0.999
)
parser.add_argument(
"--discriminator_adam_epsilon",
help="Adam optimizer's epsilon hyperparameter for numerical stability.",
type=float,
default=1e-8
)
parser.add_argument(
"--discriminator_clip_gradients",
help="Global clipping to prevent gradient norm to exceed this value for discriminator.",
type=str,
default="None"
)
parser.add_argument(
"--discriminator_train_steps",
help="Number of steps to train discriminator for per cycle.",
type=int,
default=100
)
parser.add_argument(
"--label_smoothing",
help="Multiplier when making real labels instead of all ones.",
type=float,
default=0.9
)
# Parse all arguments.
args = parser.parse_args()
arguments = args.__dict__
# Unused args provided by service.
arguments.pop("job_dir", None)
arguments.pop("job-dir", None)
# Fix input_fn_autotune.
arguments["input_fn_autotune"] = convert_string_to_bool(
string=arguments["input_fn_autotune"]
)
# Fix eval steps.
arguments["eval_steps"] = convert_string_to_none_or_int(
string=arguments["eval_steps"])
# Fix generator_projection_dims.
arguments["generator_projection_dims"] = convert_string_to_list_of_ints(
string=arguments["generator_projection_dims"], sep=","
)
# Fix use_labels.
arguments["generator_use_labels"] = convert_string_to_bool(
arguments["generator_use_labels"]
)
arguments["discriminator_use_labels"] = convert_string_to_bool(
arguments["discriminator_use_labels"]
)
# Fix embed_labels.
arguments["generator_embed_labels"] = convert_string_to_bool(
arguments["generator_embed_labels"]
)
arguments["discriminator_embed_labels"] = convert_string_to_bool(
arguments["discriminator_embed_labels"]
)
# Fix concatenate_labels.
arguments["generator_concatenate_labels"] = convert_string_to_bool(
arguments["generator_concatenate_labels"]
)
arguments["discriminator_concatenate_labels"] = convert_string_to_bool(
arguments["discriminator_concatenate_labels"]
)
# Fix num_filters.
arguments["generator_num_filters"] = convert_string_to_list_of_ints(
string=arguments["generator_num_filters"], sep=","
)
arguments["discriminator_num_filters"] = convert_string_to_list_of_ints(
string=arguments["discriminator_num_filters"], sep=","
)
# Fix kernel_sizes.
arguments["generator_kernel_sizes"] = convert_string_to_list_of_ints(
string=arguments["generator_kernel_sizes"], sep=","
)
arguments["discriminator_kernel_sizes"] = convert_string_to_list_of_ints(
string=arguments["discriminator_kernel_sizes"], sep=","
)
# Fix strides.
arguments["generator_strides"] = convert_string_to_list_of_ints(
string=arguments["generator_strides"], sep=","
)
arguments["discriminator_strides"] = convert_string_to_list_of_ints(
string=arguments["discriminator_strides"], sep=","
)
# Fix discriminator_dropout_rates.
arguments["discriminator_dropout_rates"] = convert_string_to_list_of_floats(
string=arguments["discriminator_dropout_rates"], sep=","
)
# Fix clip_gradients.
arguments["generator_clip_gradients"] = convert_string_to_none_or_float(
string=arguments["generator_clip_gradients"]
)
arguments["discriminator_clip_gradients"] = convert_string_to_none_or_float(
string=arguments["discriminator_clip_gradients"]
)
# Append trial_id to path if we are doing hptuning.
# This code can be removed if you are not using hyperparameter tuning.
arguments["output_dir"] = os.path.join(
arguments["output_dir"],
json.loads(
os.environ.get(
"TF_CONFIG", "{}"
)
).get("task", {}).get("trial", ""))
# Run the training job.
model.train_and_evaluate(arguments)
# -
| machine_learning/gan/cdcgan/tf_cdcgan/tf_cdcgan_module.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <img style="float: left; padding-right: 10px; width: 45px" src="https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/iacs.png"> CS-109A Introduction to Data Science
#
# ## Lab 4: Multiple and Polynomial Regression (September 26, 2019 version)
#
# **Harvard University**<br/>
# **Fall 2019**<br/>
# **Instructors:** <NAME>, <NAME>, and <NAME><br/>
# **Lab Instructor:** <NAME> and <NAME><br/>
# **Authors:** <NAME>, <NAME>, <NAME>, <NAME>, <NAME>
#
# ---
## RUN THIS CELL TO GET THE RIGHT FORMATTING
import requests
from IPython.core.display import HTML
styles = requests.get("https://raw.githubusercontent.com/Harvard-IACS/2018-CS109A/master/content/styles/cs109.css").text
HTML(styles)
# ## Table of Contents
#
# <ol start="0">
# <li> Learning Goals / Tip of the Week / Terminology </li>
# <li> Training/Validation/Testing Splits (slides + interactive warm-up)</li>
# <li> Polynomial Regression, and Revisiting the Cab Data</li>
# <li> Multiple regression and exploring the Football data </li>
# <li> A nice trick for forward-backwards </li>
# <li> Cross-validation</li>
# </ol>
# ## Learning Goals
# After this lab, you should be able to
# - Explain the difference between train/validation/test data and WHY we have each.
# - Implement cross-validation on a dataset
# - Implement arbitrary multiple regression models in both SK-learn and Statsmodels.
# - Interpret the coefficent estimates produced by each model, including transformed and dummy variables
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import statsmodels.api as sm
from statsmodels.api import OLS
from sklearn import preprocessing
from sklearn.preprocessing import PolynomialFeatures
from sklearn.metrics import r2_score
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from pandas.plotting import scatter_matrix
import seaborn as sns
# %matplotlib inline
# -
# ## Extra Tip of the Week
#
# Within your terminal (aka console aka command prompt), most shell environments support useful shortcuts:
#
# <ul>
# <li>press the [up arrow] to navigate through your most recent commands</li>
# <li>press [CTRL + A] to go to the beginning of the line</li>
# <li>press [CTRL + E] to go to the end of the line</li>
# <li>press [CTRL + K] to clear the line</li>
# <li>type `history` to see the last commands you've run</li>
# </ul>
#
#
# ## Terminology
#
# Say we have input features $X$, which via some function $f()$, approximates outputs $Y$. That is, $Y = f(X) + \epsilon$ (where $\epsilon$ represents our unmeasurable variation (i.e., irreducible error).
#
# - **Inference**: estimates the function $f$, but the goal isn't to make predictions for $Y$; rather, it is more concerned with understanding the relationship between $X$ and $Y$.
# - **Prediction**: estimates the function $f$ with the goal of making accurate $Y$ predictions for some unseen $X$.
#
#
# We have recently used two highly popular, useful libraries, `statsmodels` and `sklearn`.
#
# `statsmodels` is mostly focused on the _inference_ task. It aims to make good estimates for $f()$ (via solving for our $\beta$'s), and it provides expansive details about its certainty. It provides lots of tools to discuss confidence, but isn't great at dealing with test sets.
#
# `sklearn` is mostly focused on the _prediction_ task. It aims to make a well-fit line to our input data $X$, so as to make good $Y$ predictions for some unseen inputs $X$. It provides a shallower analysis of our variables. In other words, `sklearn` is great at test sets and validations, but it can't really discuss uncertainty in the parameters or predictions.
#
#
# - **R-squared**: An interpretable summary of how well the model did. 1 is perfect, 0 is a trivial baseline model based on the mean $y$ value, and negative is worse than the trivial model.
# - **F-statistic**: A value testing whether we're likely to see these results (or even stronger ones) if none of the predictors actually mattered.
# - **Prob (F-statistic)**: The probability that we'd see these results (or even stronger ones) if none of the predictors actually mattered. If this probability is small then either A) some combination of predictors actually matters or B) something rather unlikely has happened
# - **coef**: The estimate of each beta. This has several sub-components:
# - **std err**: The amount we'd expect this value to wiggle if we re-did the data collection and re-ran our model. More data tends to make this wiggle smaller, but sometimes the collected data just isn't enough to pin down a particular value.
# - **t and P>|t|**: similar to the F-statistic, these measure the probability of seeing coefficients this big (or even bigger) if the given variable didn't actually matter. Small probability doesn't necessarily mean the value matters
# - **\[0.025 0.975\]**: Endpoints of the 95% confidence interval. This is a interval drawn in a clever way and which gives an idea of where the true beta value might plausibly live. (If you want to understand why "there's a 95% chance the true beta is in the interval" is _wrong_, start a chat with Will : )
#
# ## Part 2: Polynomial Regression, and Revisiting the Cab Data
#
# Polynomial regression uses a **linear model** to estimate a **non-linear function** (i.e., a function with polynomial terms). For example:
#
# $y = \beta_0 + \beta_1x_i + \beta_1x_i^{2}$
#
# It is a linear model because we are still solving a linear equation (the _linear_ aspect refers to the beta coefficients).
# read in the data, break into train and test
cab_df = pd.read_csv("../data/dataset_1.txt")
train_data, test_data = train_test_split(cab_df, test_size=0.2, random_state=42)
cab_df.head()
cab_df.shape
# +
# do some data cleaning
#print(train_data['TimeMin'])
#print(train_data['TimeMin'].values) <- Puts data into list form
#print(train_data['TimeMin'].values.reshape(-1, 1)) <- Puts data into single column matrix form
X_train = train_data['TimeMin'].values.reshape(-1,1)/60 # transforms it to being hour-based
y_train = train_data['PickupCount'].values
X_test = test_data['TimeMin'].values.reshape(-1,1)/60 # hour-based
y_test = test_data['PickupCount'].values
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
def plot_cabs(cur_model, poly_transformer=None):
# build the x values for the prediction line
x_vals = np.arange(0,24,.1).reshape(-1,1)
# optionally use the passed-in transformer
if poly_transformer != None:
dm = poly_transformer.fit_transform(x_vals)
else:
dm = x_vals
# make the prediction at each x value
prediction = cur_model.predict(dm)
# plot the prediction line, and the test data
plt.plot(x_vals,prediction, color='k', label="Prediction")
plt.scatter(X_test, y_test, label="Test Data")
# label your plots
plt.ylabel("Number of Taxi Pickups")
plt.xlabel("Time of Day (Hours Past Midnight)")
plt.legend()
plt.show()
# -
from sklearn.linear_model import LinearRegression
fitted_cab_model0 = LinearRegression().fit(X_train, y_train)
plot_cabs(fitted_cab_model0)
fitted_cab_model0.score(X_test, y_test)
# <div class="exercise"><b>Exercise</b></div>
#
# **Questions**:
# 1. The above code uses `sklearn`. As more practice, and to help you stay versed in both libraries, perform the same task (fit a linear regression line) using `statsmodels` and report the $r^2$ score. Is it the same value as what sklearn reports, and is this the expected behavior?
# +
#### EXERCISE: write code here (feel free to work with a partner)
# 1. Shape X train AND test data
X_train_shaped = sm.add_constant(X_train)
X_test_shaped = sm.add_constant(X_test)
print(X_train_shaped.shape, X_test_shaped.shape)
# 2. Fit model to new model instance
OLSModel = OLS(y_train, X_train_shaped).fit()
# 3. Predict results using model instance
y_predicted = OLSModel.predict(X_test_shaped)
# 4. Retrieve instance score comparing y_test and y_predicted
r2 = r2_score(y_test, y_predicted)
print(r2)
# -
# We can see that there's still a lot of variation in cab pickups that's not being captured by a linear fit. Further, the linear fit is predicting massively more pickups at 11:59pm than at 12:00am. This is a bad property, and it's the conseqeuence of having a straight line with a non-zero slope. However, we can add columns to our data for $TimeMin^2$ and $TimeMin^3$ and so on, allowing a curvy polynomial line to hopefully fit the data better.
#
# We'll be using ``sklearn``'s `PolynomialFeatures()` function to take some of the tedium out of building the expanded input data. In fact, if all we want is a formula like $y \approx \beta_0 + \beta_1 x + \beta_2 x^2 + ...$, it will directly return a new copy of the data in this format!
# +
# 1. Create instance of PolynomialFeatures up to 3 polynomial formula values
transformer_3 = PolynomialFeatures(3, include_bias=False)
# 2. .fit_transform the data using X_train
expanded_train = transformer_3.fit_transform(X_train) # TRANSFORMS it to polynomial features
# 3. DataFrame only contains 3 new polynomial columns (x^{1, 2, 3})
pd.DataFrame(expanded_train).describe() # notice that the columns now contain x, x^2, x^3 values
# -
# A few notes on `PolynomialFeatures`:
#
# - The interface is a bit strange. `PolynomialFeatures` is a _'transformer'_ in sklearn. We'll be using several transformers that learn a transformation on the training data, and then we will apply those transformations on future data. With PolynomialFeatures, the `.fit()` is pretty trivial, and we often fit and transform in one command, as seen above with ``.fit_transform()`.
# - You rarely want to `include_bias` (a column of all 1's), since _**sklearn**_ will add it automatically. Remember, when using _**statsmodels,**_ you can just `.add_constant()` right before you fit the data.
# - If you want polynomial features for a several different variables (i.e., multinomial regression), you should call `.fit_transform()` separately on each column and append all the results to a copy of the data (unless you also want interaction terms between the newly-created features). See `np.concatenate()` for joining arrays.
# +
# 1. Create prediction model instance from LinearRegression. Fit using transformed x_train and y_train.
fitted_cab_model3 = LinearRegression().fit(expanded_train, y_train)
# 2. Print expanded_train created in last code cell for review
print("fitting expanded_train:", expanded_train)
# 3. Chart using earlier defined function, including a new polynomial transformer that uses .fit_transform
plot_cabs(fitted_cab_model3, transformer_3)
# -
# <div class="exercise"><b>Exercise</b></div>
#
# **Questions**:
# 1. Calculate the polynomial model's $R^2$ performance on the test set.
# 2. Does the polynomial model improve on the purely linear model?
# 3. Make a residual plot for the polynomial model. What does this plot tell us about the model?
# +
# ANSWER 1
expanded_test = transformer_3.fit_transform(X_test) # Must transform x_test for accurate predictions
print("Test R-squared:", fitted_cab_model3.score(expanded_test, y_test))
# NOTE 1: unlike statsmodels' r2_score() function, sklearn has a .score() function
# NOTE 2: fit_transform() is a nifty function that transforms the data, then fits it
# -
# ANSWER 2: does it?
print("Yes, it does because it takes into effect the cyclical nature of a day and has a higher R2 score.")
# +
# ANSWER 3 (class discussion about the residuals)
# Transform x_train data to x_matrix with polynomial columns.
x_matrix = transformer_3.fit_transform(X_train)
# Convert to pd.DataFrame before describing. See 3 columns!
print(pd.DataFrame(x_matrix).describe())
prediction = fitted_cab_model3.predict(x_matrix)
# Residuals are y_train - prediction. Why y_train, not y_test?
residual = y_train - prediction
plt.scatter(X_train, residual, label="Residual")
plt.axhline(0, color='k')
plt.title("Residuals for the Cubic Model")
plt.ylabel("Residual Number of Taxi Pickups")
plt.xlabel("Time of Day (Hours Past Midnight)")
plt.legend();
# -
# #### Other features
# Polynomial features are not the only constucted features that help fit the data. Because these data have a 24 hour cycle, we may want to build features that follow such a cycle. For example, $sin(24\frac{x}{2\pi})$, $sin(12\frac{x}{2\pi})$, $sin(8\frac{x}{2\pi})$. Other feature transformations are appropriate to other types of data. For instance certain feature transformations have been developed for geographical data.
#
# ### Scaling Features
# When using polynomials, we are explicitly trying to use the higher-order values for a given feature. However, sometimes these polynomial features can take on values that are drastically large, making it difficult for the system to learn an appropriate bias weight due to its large values and potentially large variance. To counter this, sometimes one may be interested in scaling the values for a given feature.
#
# For our ongoing taxi-pickup example, using polynomial features improved our model. If we wished to scale the features, we could use `sklearn`'s StandardScaler() function:
# +
# SCALES THE EXPANDED/POLY TRANSFORMED DATA
# we don't need to convert to a pandas dataframe, but it can be useful for scaling select columns
train_copy = pd.DataFrame(expanded_train.copy())
test_copy = pd.DataFrame(expanded_test.copy())
# Fit the scaler on the training data
scaler = StandardScaler().fit(train_copy)
# Scale both the test and training data.
train_scaled = scaler.transform(expanded_train)
test_scaled = scaler.transform(expanded_test)
# we could optionally run a new regression model on this scaled data
fitted_scaled_cab = LinearRegression().fit(train_scaled, y_train)
fitted_scaled_cab.score(test_scaled, y_test)
# -
# <hr style="height:3px">
#
# ## Part 3: Multiple regression and exploring the Football (aka soccer) data
# Let's move on to a different dataset! The data imported below were scraped by [<NAME>](https://www.kaggle.com/mauryashubham/linear-regression-to-predict-market-value/data) and record various facts about players in the English Premier League. Our goal will be to fit models that predict the players' market value (what the player could earn when hired by a new team), as estimated by https://www.transfermarkt.us.
#
# `name`: Name of the player
# `club`: Club of the player
# `age` : Age of the player
# `position` : The usual position on the pitch
# `position_cat` : 1 for attackers, 2 for midfielders, 3 for defenders, 4 for goalkeepers
# `market_value` : As on transfermrkt.com on July 20th, 2017
# `page_views` : Average daily Wikipedia page views from September 1, 2016 to May 1, 2017
# `fpl_value` : Value in Fantasy Premier League as on July 20th, 2017
# `fpl_sel` : % of FPL players who have selected that player in their team
# `fpl_points` : FPL points accumulated over the previous season
# `region`: 1 for England, 2 for EU, 3 for Americas, 4 for Rest of World
# `nationality`: Player's nationality
# `new_foreign`: Whether a new signing from a different league, for 2017/18 (till 20th July)
# `age_cat`: a categorical version of the Age feature
# `club_id`: a numerical version of the Club feature
# `big_club`: Whether one of the Top 6 clubs
# `new_signing`: Whether a new signing for 2017/18 (till 20th July)
#
# As always, we first import, verify, split, and explore the data.
#
# ## Part 3.1: Import and verification and grouping
# +
league_df = pd.read_csv("../data/league_data.txt")
print(league_df.dtypes)
# QUESTION: what would you guess is the mean age? mean salary?
print("------------------------")
print("Mean age: ", league_df.age.mean())
print("Mean salary: ", np.mean(league_df['market_value']))
#league_df.head()
# -
league_df.shape
league_df.describe()
# ### (Stratified) train/test split
# We want to make sure that the training and test data have appropriate representation of each region; it would be bad for the training data to entirely miss a region. This is especially important because some regions are rather rare.
#
# <div class="exercise"><b>Exercise</b></div>
#
# **Questions**:
# 1. Use the `train_test_split()` function, while (a) ensuring the test size is 20% of the data, and; (2) using 'stratify' argument to split the data (look up documentation online), keeping equal representation of each region. This doesn't work by default, correct? What is the issue?
# 2. Deal with the issue you encountered above. Hint: you may find numpy's `.isnan()` and panda's `.dropna()` functions useful!
# 3. How did you deal with the error generated by `train_test_split`? How did you justify your action?
# *your answer here*:
#
# +
# EXERCISE: feel free to work with a partner
# 1. Use train_test_split with stratify argument on region
#train_df, test_df = train_test_split(league_df, test_size=0.2, stratify=league_df['region'])
# 2. The above doesn't work because there is a single nan value...
#print(list(league_df.region))
# 2. Try with .dropna(). Delete players with no region!
train_df, test_df = train_test_split(league_df.dropna(subset=['region']), test_size=0.2, stratify=league_df['region'].dropna())
# -
train_df.shape, test_df.shape
# Now that we won't be peeking at the test set, let's explore and look for patterns! We'll introduce a number of useful pandas and numpy functions along the way.
# ### Groupby
# Pandas' `.groupby()` function is a wonderful tool for data analysis. It allows us to analyze each of several subgroups.
#
# Many times, `.groupby()` is combined with `.agg()` to get a summary statistic for each subgroup. For instance: What is the average market value, median page views, and maximum fpl for each player position?
train_df.groupby('position').agg({
'market_value': np.mean,
'page_views': np.median,
'fpl_points': np.max
})
train_df.position.unique()
train_df.groupby(['big_club', 'position']).agg({
'market_value': np.mean,
'page_views': np.mean,
'fpl_points': np.mean
})
# <div class="exercise"><b>Exercise</b></div>
#
# **Question**:
# 1. Notice that the `.groupby()` function above takes a list of two column names. Does the order matter? What happens if we switch the two so that 'position' is listed before 'big_club'?
# +
# EXERCISE: feel free to work with a partner
# Yes, order matters in presentation, though not calculation
train_df.groupby(['position', 'big_club']).agg({
'market_value': np.mean,
'page_views': np.mean,
'fpl_points': np.mean
})
# -
# <hr style="height:3px">
#
# ## Part 3.2: Linear regression on the football data
# This section of the lab focuses on fitting a model to the football (soccer) data and interpreting the model results. The model we'll use is
#
# $$\text{market_value} \approx \beta_0 + \beta_1\text{fpl_points} + \beta_2\text{age} + \beta_3\text{age}^2 + \beta_4log_2\left(\text{page_views}\right) + \beta_5\text{new_signing} +\beta_6\text{big_club} + \beta_7\text{position_cat}$$
#
# We're including a 2nd degree polynomial in age because we expect pay to increase as a player gains experience, but then decrease as they continue aging. We're taking the log of page views because they have such a large, skewed range and the transformed variable will have fewer outliers that could bias the line. We choose the base of the log to be 2 just to make interpretation cleaner.
#
# <div class="exercise"><b>Exercise</b></div>
#
# **Questions**:
# 1. Build the data and fit this model to it. How good is the overall model?
# 2. Interpret the regression model. What is the meaning of the coefficient for:
# - age and age$^2$
# - $log_2($page_views$)$
# - big_club
# 3. What should a player do in order to improve their market value? How many page views should a player go get to increase their market value by 10?
# +
# Q1: we'll do most of it for you ...
y_train = train_df['market_value']
y_test = test_df['market_value']
def build_football_data(df):
x_matrix = df[['fpl_points','age','new_signing','big_club','position_cat']].copy()
x_matrix['log_views'] = np.log2(df['page_views'])
# WRITE CODE FOR CREATING THE AGE SQUARED COLUMN
####
x_matrix['age2'] = df['age']**2
# OPTIONALLY WRITE CODE to adjust the ordering of the columns, just so that it corresponds with the equation above
#### Start with x_matrix since that has new columns that df doesn't.
x_matrix = x_matrix[['fpl_points', 'age', 'age2', 'log_views', 'new_signing', 'big_club', 'position_cat']]
# add a constant
x_matrix = sm.add_constant(x_matrix)
return x_matrix
# use build_football_data() to transform both the train_data and test_data
train_transformed = build_football_data(train_df)
test_transformed = build_football_data(test_df)
fitted_model_1 = OLS(endog= y_train, exog=train_transformed, hasconst=True).fit()
print(fitted_model_1.summary())
# WRITE CODE TO RUN r2_score(), then answer the above question about the overall goodness of the model
# Plug in y_test actual values and y_predicted values derived from x_test!!
r2 = r2_score(y_test, fitted_model_1.predict(test_transformed))
print("-----------------------------------------------")
print("R-squared score: ", r2)
# +
# Q2: let's use the age coefficients to show the effect of age has on one's market value;
# we can get the age and age^2 coefficients via:
agecoef = fitted_model_1.params.age
age2coef = fitted_model_1.params.age2
# let's set our x-axis (corresponding to age) to be a wide range from -100 to 100,
# just to see a grand picture of the function
x_vals = np.linspace(-100,100,1000) # Usually when charting prediction line, you should make your own x_vals.
y_vals = agecoef*x_vals + age2coef*x_vals**2
# WRITE CODE TO PLOT x_vals vs y_vals
fig, ax = plt.subplots(2,2, figsize=(20, 20))
ax[0,0].plot(x_vals, y_vals)
ax[0,0].set_title("Effect of Age")
ax[0,0].set_xlabel("Age")
ax[0,0].set_ylabel("Contribution to Predicted Market Value")
ax[0,0].set_xlim(0, 100)
# Q2A: WHAT HAPPENS IF WE USED ONLY AGE (not AGE^2) in our model (what's the r2?); make the same plot of age vs market value
y_vals_2a = agecoef*x_vals
ax[0,1].plot(x_vals, y_vals_2a)
ax[0,1].set_title("Effect of Age (non-Polynomial)")
ax[0,1].set_xlabel("Age")
ax[0,1].set_ylabel("Contribution to Predicted Market Value")
ax[0,1].set_xlim(0, 100)
# Q2B: WHAT HAPPENS IF WE USED ONLY AGE^2 (not age) in our model (what's the r2?); make the same plot of age^2 vs market value
y_vals_2b = agecoef*x_vals**2
ax[1,0].plot(x_vals, y_vals_2b)
ax[1,0].set_title("Effect of Age (just-Polynomial)")
ax[1,0].set_xlabel("Age")
ax[1,0].set_ylabel("Contribution to Predicted Market Value")
ax[1,0].set_xlim(0, 100)
# Q2C: PLOT page views vs market value
pageviewscoef = fitted_model_1.params.log_views
y_vals_2c = pageviewscoef*x_vals
ax[1,1].plot(x_vals, y_vals_2c)
ax[1,1].set_title("Page Views vs Market Value")
ax[1,1].set_xlabel("Page Views")
ax[1,1].set_ylabel("Market Value")
ax[1,1].set_xlim(0, 100)
plt.show()
# -
# <hr style='height:3px'>
#
# ### Part 3.3: Turning Categorical Variables into multiple binary variables
# Of course, we have an error in how we've included player position. Even though the variable is numeric (1,2,3,4) and the model runs without issue, the value we're getting back is garbage. The interpretation, such as it is, is that there is an equal effect of moving from position category 1 to 2, from 2 to 3, and from 3 to 4, and that this effect is probably between -0.5 to -1 (depending on your run).
#
# In reality, we don't expect moving from one position category to another to be equivalent, nor for a move from category 1 to category 3 to be twice as important as a move from category 1 to category 2. We need to introduce better features to model this variable.
#
# We'll use `pd.get_dummies` to do the work for us.
# +
train_design_recoded = pd.get_dummies(train_transformed, columns=['position_cat'], drop_first=True)
test_design_recoded = pd.get_dummies(test_transformed, columns=['position_cat'], drop_first=True)
train_design_recoded.head()
# -
# We've removed the original `position_cat` column and created three new ones.
#
# #### Why only three new columns?
# Why does pandas give us the option to drop the first category?
#
# <div class="exercise"><b>Exercise</b></div>
#
# **Questions**:
# 1. If we're fitting a model without a constant, should we have three dummy columns or four dummy columns?
# 2. Fit a model on the new, recoded data, then interpret the coefficient of `position_cat_2`.
#
# FEEL FREE TO WORK WITH A PARTNER
resu = OLS(y_train, train_design_recoded).fit()
resu.summary()
print("r2:", r2_score(y_test, resu.predict(test_design_recoded)))
print("position_cat_2 coef:", resu.params.position_cat_2)
train_design_recoded.shape, y_train.shape
# **Answers**:
#
# 1. If our model does not have a constant, we must include all four dummy variable columns. If we drop one, we're not modeling any effect of being in that category, and effectively assuming the dropped category's effect is 0.
# 2. Being in position 2 (instead of position 1) has an impact between -1.54 and +2.38 on a player's market value. Since we're using an intercept, the dropped category becomes the baseline and the effect of any dummy variable is the effect of being in that category instead of the baseline category.
#
# ## Part 4: A nice trick for forward-backwards
#
# XOR (operator ^) is a logical operation that only returns true when input differ. We can use it to implement forward-or-backwards selection when we want to keep track of what predictors are "left" from a given list of predictors.
#
# The set analog is "symmetric difference". From the python docs:
#
# `s.symmetric_difference(t) s ^ t new set with elements in either s or t but not both`
#
set() ^ set([1,2,3])
set([1]) ^ set([1,2,3])
set([1, 2]) ^ set([1,2,3])
# <div class="exercise"><b>Exercise</b></div>
#
# Outline a step-forwards algorithm which uses this idea
#
# ## BONUS EXERCISE:
# We have provided a spreadsheet of Boston housing prices (data/boston_housing.csv). The 14 columns are as follows:
# 1. CRIM: per capita crime rate by town
# 2. ZN: proportion of residential land zoned for lots over 25,000 sq.ft.
# 3. INDUS: proportion of non-retail business acres per town
# 4. CHAS: Charles River dummy variable (= 1 if tract bounds river; 0 otherwise)
# 5. NOX: nitric oxides concentration (parts per 10 million) 1https://archive.ics.uci.edu/ml/datasets/Housing 123 20.2. Load the Dataset 124
# 6. RM: average number of rooms per dwelling
# 7. AGE: proportion of owner-occupied units built prior to 1940
# 8. DIS: weighted distances to five Boston employment centers
# 9. RAD: index of accessibility to radial highways
# 10. TAX: full-value property-tax rate per \$10,000
# 11. PTRATIO: pupil-teacher ratio by town
# 12. B: 1000(Bk−0.63)2 where Bk is the proportion of blacks by town
# 13. LSTAT: % lower status of the population
# 14. MEDV: Median value of owner-occupied homes in $1000s We can see that the input attributes have a mixture of units
#
# There are 450 observations.
# <div class="exercise"><b>Exercise</b></div>
#
# Using the above file, try your best to predict **housing prices. (the 14th column)** We have provided a test set `data/boston_housing_test.csv` but refrain from looking at the file or evaluating on it until you have finalized and trained a model.
# 1. Load in the data. It is tab-delimited. Quickly look at a summary of the data to familiarize yourself with it and ensure nothing is too egregious.
# 2. Use a previously-discussed function to automatically partition the data into a training and validation (aka development) set. It is up to you to choose how large these two portions should be.
# 3. Train a basic model on just a subset of the features. What is the performance on the validation set?
# 4. Train a basic model on all of the features. What is the performance on the validation set?
# 5. Toy with the model until you feel your results are reasonably good.
# 6. Perform cross-validation with said model, and measure the average performance. Are the results what you expected? Were the average results better or worse than that from your original 1 validation set?
# 7. Experiment with other models, and for each, perform 10-fold cross-validation. Which model yields the best average performance? Select this as your final model.
# 8. Use this model to evaulate your performance on the testing set. What is your performance (MSE)? Is this what you expected?
| content/labs/lab04/notebook/cs109a_lab4_regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
# + active=""
# Известно, что генеральная совокупность распределена нормально со средним квадратическим отклонением, равным 16. Найти доверительный интервал для оценки математического ожидания a с надежностью 0.95, если выборочная средняя M = 80, а объем выборки n = 256.
# + active=""
# а=0.95 -> z=1.96
# сигма 16
# n = 256
# Xсред = 80
# + active=""
# 80+- (1.96 * (16 / 16))
#
# Доверительный интервал: [78.04;81.96]
# -
int1 = 80-(1.96*16/256**0.5)
int2 = 80+(1.96*16/256**0.5)
print('[' , int1 , ';' , int2,']')
# + active=""
# В результате 10 независимых измерений некоторой величины X, выполненных с одинаковой точностью, получены опытные данные: 6.9, 6.1, 6.2, 6.8, 7.5, 6.3, 6.4, 6.9, 6.7, 6.1 Предполагая, что результаты измерений подчинены нормальному закону распределения вероятностей, оценить истинное значение величины X при помощи доверительного интервала, покрывающего это значение с доверительной вероятностью 0,95. 3,4 задачи решать через тестирование гипотезы
# -
x = [6.9, 6.1, 6.2, 6.8, 7.5, 6.3, 6.4, 6.9, 6.7, 6.1]
a = pd.DataFrame(x)
#стандартное отклонение
std = a.std(ddof=1)
m = a.mean()
z = 2.262
int1 = m - (z * std/10**0.5)
int2 = m + (z * std/10**0.5)
print('[' , int1[0] , ';' , int2[0],']')
# + active=""
# Утверждается, что шарики для подшипников, изготовленные автоматическим станком, имеют средний диаметр 17 мм.
# Используя односторонний критерий с α=0,05, проверить эту гипотезу, если в выборке из n=100 шариков средний диаметр
# оказался равным 17.5 мм, а дисперсия известна и равна 4 кв.мм.
# +
# H0: диаметр = 17мм
# H1: диаметр = 17.5мм
a = 0.05
n = 100
sigma = 4
# дисперсия(sigma) известна - использую z критерий
Z_tablichnoe = 1.654
Z_raschetnoe = (17.5 - 17) / 2 / n**0.5
if Z_tablichnoe > Z_raschetnoe:
right = 'H0'
else:
right = 'H1'
print('верна гипотеза',right, 'на уровне значимости', a)
# + active=""
# Продавец утверждает, что средний вес пачки печенья составляет 200 г.
# Из партии извлечена выборка из 10 пачек. Вес каждой пачки составляет:
# 202, 203, 199, 197, 195, 201, 200, 204, 194, 190.
# Известно, что их веса распределены нормально.
# Верно ли утверждение продавца, если учитывать, что доверительная вероятность равна 99%?
# +
# H0: вес пачки = 200
# H1: вес пачки != 200
a = 0.01
# дисперсия неизвестна -> использую t критерий
x = [202, 203, 199, 197, 195, 201, 200, 204, 194, 190]
x_df = pd.DataFrame(x)
#стандартное отклонение
std = x_df.std(ddof=1)
m = x_df.mean()
T_tablich = 3.250
T_raschet = (m - 200) / std / 10
if T_tablich > T_raschet[0]:
right = 'H0'
else:
right = 'H1'
print('верна гипотеза',right, 'на уровне значимости', a)
# -
| probability/.ipynb_checkpoints/05-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Read file
def read_file(filename):
with open(filename) as infile:
lines = [int(line.strip()) for line in infile.readlines()]
return lines
# +
# Part 1
import collections
def joltage_jumps(numbers):
s = sorted(numbers)
s.insert(0, 0)
s.append(s[-1] + 3)
return [y-x for (x,y) in zip(s[:-1], s[1:])]
# -
# Test part 1
test1 = collections.Counter(joltage_jumps(read_file("test01.txt")))
test2 = collections.Counter(joltage_jumps(read_file("test02.txt")))
(test1[1], test1[3]) == (7, 5) and \
(test2[1], test2[3]) == (22, 10)
# Solve part 1
part1 = collections.Counter(joltage_jumps(read_file("input.txt")))
part1[1] * part1[3]
# Part 2
def combinations(jumps, i=0, memo=None):
if memo == None:
memo = {}
if i in memo:
return memo[i]
if len(jumps) - i <= 1:
return 1
memo[i] = combinations(jumps, i+1, memo)
if jumps[i:i+2] in [[2, 1], [1, 2], [1, 1]]:
memo[i] += combinations(jumps, i+2, memo)
if jumps[i:i+3] == [1, 1, 1]:
memo[i] += combinations(jumps, i+3, memo)
return memo[i]
# Test part 2
combinations(joltage_jumps(read_file("test01.txt"))) == 8 and \
combinations(joltage_jumps(read_file("test02.txt"))) == 19208
# Solve part 2
combinations(joltage_jumps(read_file("input.txt")))
| 10/10.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#hide
# %load_ext autoreload
# %autoreload 2
# !pip install -r ../requirements.txt > /dev/null
# +
# default_exp settings
# +
# export
from deeppavlov.core.common.paths import get_settings_path
from deeppavlov import configs, build_model, train_model
import json
from os import path, popen, mkdir
from shutil import copyfile
import pandas as pd
import numpy as np
from pathlib import Path
import logging
from collections import defaultdict
# +
#hide
logging.basicConfig(
#filename='example.log',
format='%(asctime)s %(levelname)s:%(message)s',
level=logging.DEBUG,
datefmt='%I:%M:%S'
)
logging.info("Hello! Welcome to our automated dialog system!")
logging.debug(" Debug Log Active")
logging.warning(' Warning Log Active')
logging.error(' Error Log Active ')
# -
# # Settings and Configuration
# > Question Answering Automated Dialog System
# +
#export
def change_log_config():
'''Change Deeppavlov configuration files to ERROR mode
'''
settings_file = path.join(get_settings_path(), 'log_config.json')
#logs_key = 'disable_existing_loggers'
settings_json = json.load(open(settings_file))
settings_json['handlers']['file']['level'] = 'ERROR'
settings_json['handlers']['stderr']['level'] = 'ERROR'
settings_json['handlers']['stdout']['level'] = 'ERROR'
settings_json['handlers']['uvicorn_handler']['level'] = 'ERROR'
settings_json['loggers']['deeppavlov']['level'] = 'ERROR'
settings_json['loggers']['deeppavlov']['propagate'] = True
settings_json['loggers']['uvicorn.access']['level'] = 'ERROR'
settings_json['loggers']['uvicorn.access']['propagate'] = True
settings_json['loggers']['uvicorn.error']['level'] = 'ERROR'
settings_json['loggers']['uvicorn.error']['propagate'] = True
json.dump(settings_json, open(settings_file, 'w'))
def run_shell_installs():
''' Run install commands
'''
logging.info(f'..Installing NLP libraries')
change_log_config()
command_strings = (
' pip install deeppavlov', ' python -m deeppavlov install squad',
' python -m deeppavlov install squad_bert',
' python -m deeppavlov install fasttext_avg_autofaq',
' python -m deeppavlov install fasttext_tfidf_autofaq',
' python -m deeppavlov install tfidf_autofaq',
' python -m deeppavlov install tfidf_logreg_autofaq ',
' python -m deeppavlov install tfidf_logreg_en_faq'
)
for command in command_strings:
logging.debug(command)
logging.debug(popen(command).read())
# -
#hide
run_shell_installs()
# +
#export
def action_over_list_f(arr, v):
''' v[0] and v[1] are dictionaries
arr is array of dictionaries
'''
k_id, v_id = next(iter(v[0].items()))
for p, a in enumerate(arr):
if k_id in a.keys() and a[k_id] == v_id:
for k_rep, v_rep in v[1].items():
arr[p][k_rep] = v_rep
def replacement_f(model_config, **args):
'''Replaces the model config dictionary with new values
provided in **args
'''
for k, v in args.items():
if isinstance(v, dict):
replacement_f(model_config[k], **v)
if isinstance(v, str):
model_config[k] = v
if isinstance(model_config[k], list):
action_over_list_f(model_config[k], v)
# +
# test action_over_list_f
from random import randint
def gen_list_keys_for_tests():
'''This function is used for tests
'''
str_n = lambda x: f'{x}_{randint(1,10):1}'
gen_dict_list = lambda: {
'id': str_n('id'),
'key1': str_n('v1'),
'key2': str_n('v2'),
'key3': str_n('v3')
}
pipe_list = [gen_dict_list() for _ in range(randint(3, 10))]
rand_id = pipe_list[randint(0, len(pipe_list) - 1)]['id']
rand_key = f'key{randint(1, 3)}'
new_rand_val = str_n('new')
args = {
'chains': {
'pipe': [{
'id': rand_id
}, {
rand_key : new_rand_val
}]
}
}
return pipe_list, rand_id, rand_key, args, new_rand_val
def test_action_over_list_f():
pipe_list, rand_id, rand_key, args, new_rand_val = gen_list_keys_for_tests()
assert all(
new_rand_val not in pipe_elem.values() for pipe_elem in pipe_list
)
action_over_list_f(pipe_list, args['chains']['pipe'])
assert any(
rand_key in pipe_elem.keys() and
new_rand_val in pipe_elem.values() for pipe_elem in pipe_list
)
def test_replacement_f_list():
pipe_list, rand_id, rand_key, args, new_rand_val = gen_list_keys_for_tests()
mod_conf = {'chains': {'pipe': pipe_list}}
assert all(
new_rand_val not in pipe_elem.values()
for pipe_elem in mod_conf['chains']['pipe']
)
replacement_f(model_config=mod_conf, **args)
assert any(
rand_key in pipe_elem.keys() and
new_rand_val in pipe_elem.values()
for pipe_elem in mod_conf['chains']['pipe']
)
def test_replacement_f_val():
args = {'key3': 'newvalue'}
mod_conf = {'key1': 'val1', 'key2': 'val2', 'key3': 'val3'}
replacement_f(model_config=mod_conf, **args)
assert all(
arg_k in mod_conf.keys() and arg_v in mod_conf.values()
for arg_k, arg_v in args.items()
)
def test_replacement_f_dict():
args = {'1_key_3': {'2_key_2': 'newvalue'}}
mod_conf = {'1_key_3': {'2_key_2': 'oldvalue'}, '0_key_': '0_val'}
replacement_f(model_config=mod_conf, **args)
assert mod_conf['1_key_3']['2_key_2'] == 'newvalue'
test_action_over_list_f()
test_replacement_f_list()
test_replacement_f_val()
test_replacement_f_dict()
# -
#export
def updates_faq_config_file(
configs_path,
**args
):
'''Updates deepplavov json config file
'''
#set FAQ data in config file
model_config = json.load(open(configs_path))
if 'data_url' in model_config['dataset_reader']:
del model_config['dataset_reader']['data_url']
replacement_f(model_config=model_config,**args)
json.dump(model_config, open(configs_path, 'w'))
# +
#test updates_faq_config_file
import tempfile
from shutil import copyfile
def gen_list_keys_for_tests():
str_n = lambda x: f'{x}_{randint(1,10):1}'
gen_dict_list = lambda: {
'id': str_n('id'),
'key1': str_n('v1'),
'key2': str_n('v2'),
'key3': str_n('v3')
}
pipe_list = [gen_dict_list() for _ in range(randint(3, 10))]
rand_id = pipe_list[randint(0, len(pipe_list) - 1)]['id']
rand_key = f'key{randint(1, 3)}'
new_rand_val = str_n('new')
pipe_dict = {'pipe': [{'id': rand_id}, {rand_key: new_rand_val}]}
args = {'chainer': pipe_dict}
return pipe_list, rand_id, rand_key, args, new_rand_val
def test_updates_faq_config_file_update_string():
with tempfile.TemporaryDirectory() as tmpdirname:
tmp_config_file = path.join(tmpdirname, 'tmp_file.json')
copyfile(configs.faq.tfidf_logreg_en_faq, tmp_config_file)
assert path.isfile(tmp_config_file)
updates_faq_config_file(
configs_path=tmp_config_file,
dataset_reader={'data_path': 'fictional_csv_file.csv'}
)
config_json = json.load(open(tmp_config_file))
assert 'data_path' in config_json['dataset_reader']
def test_updates_faq_config_file_update_list():
with tempfile.TemporaryDirectory() as tmpdirname:
tmp_config_file = path.join(tmpdirname, 'tmp_file.json')
pipe_list, rand_id, rand_key, args, new_rand_val = gen_list_keys_for_tests(
)
mod_conf = {
'chainer': {
'pipe': pipe_list
},
'dataset_reader': 'dataset_reader_dictionary'
}
json.dump(mod_conf, open(tmp_config_file, 'w'))
assert path.isfile(tmp_config_file)
updates_faq_config_file(configs_path=tmp_config_file, **args)
config_json = json.load(open(tmp_config_file))
assert any(
rand_key in pipe_elem.keys() and new_rand_val in pipe_elem.values()
for pipe_elem in config_json['chainer']['pipe']
)
test_updates_faq_config_file_update_string()
test_updates_faq_config_file_update_list()
# -
#export
def select_faq_responses(faq_model, question):
'''Calls Deeppavlov FAQ model
'''
return faq_model([question])[0]
# +
#test faq responses
import tempfile
from shutil import copyfile
def gen_mock_csv_file(tmpdirname, faqs):
temp_faq_csv = path.join(tmpdirname, 'tmp_faq.csv')
pd.DataFrame(faqs).to_csv(temp_faq_csv, index=False)
return temp_faq_csv
def gen_mock_vocab_answers(tmpdirname, vocabs):
temp_dict_file = path.join(tmpdirname, 'temp_vocab_answers.dict')
vocabs_text = '\n'.join(
t + '\t' + str(f) for t, f in zip(vocabs['text'], vocabs['freq'])
)
f = open(temp_dict_file, 'w')
f.write(vocabs_text)
f.close()
return temp_dict_file
def gen_faq_config(tmpdirname, vocab_file, faq_file):
temp_configs_faq = path.join(tmpdirname, 'temp_config_faq.json')
copyfile(configs.faq.tfidf_logreg_en_faq, temp_configs_faq)
changes_dict = {'save_path': vocab_file, 'load_path': vocab_file}
id_dict = {'id': 'answers_vocab'}
updates_faq_config_file(
configs_path=temp_configs_faq,
chainer={'pipe': [id_dict, changes_dict]},
dataset_reader={'data_path': faq_file}
)
return temp_configs_faq
def test_faq_response_with_minimum_faqs_in_dataframe_fail_case():
with tempfile.TemporaryDirectory() as tmpdirname:
faqs = {
'Question': ['Is Covid erradicated?'],
'Answer': ['Definitely not!']
}
vocabs = {'text': ['This is a vocab example'], 'freq': [1]}
faq_file = gen_mock_csv_file(tmpdirname, faqs)
vocab_file = gen_mock_vocab_answers(tmpdirname, vocabs)
configs_file = gen_faq_config(tmpdirname, vocab_file, faq_file)
try:
select_faq_responses(
question='Is Enrique the prettiest person in town?',
faq_model=train_model(configs_file, download=True)
)
assert False
except ValueError as e:
assert True
def test_faq_response_with_minimum_faqs_in_dataframe_success_case():
with tempfile.TemporaryDirectory() as tmpdirname:
faqs = {
'Question': ['Is Covid erradicated?', 'Who is the current POTUS?'],
'Answer': ['Definitely not!', '<NAME>']
}
vocabs = {'text': ['This is a vocab example'], 'freq': [1]}
faq_file = gen_mock_csv_file(tmpdirname, faqs)
vocab_file = gen_mock_vocab_answers(tmpdirname, vocabs)
configs_file = gen_faq_config(tmpdirname, vocab_file, faq_file)
assert select_faq_responses(
question='Is Enrique the prettiest person in town?',
faq_model=train_model(configs_file, download=True)
) == ['<NAME>']
def test_faq_response_with_minimum_answers_vocab_success_case():
with tempfile.TemporaryDirectory() as tmpdirname:
faqs = {
'Question': ['Is Covid erradicated?', 'Who is the current POTUS?'],
'Answer': ['Definitely not!', '<NAME>']
}
vocabs = {'text': [], 'freq': []}
faq_file = gen_mock_csv_file(tmpdirname, faqs)
vocab_file = gen_mock_vocab_answers(tmpdirname, vocabs)
configs_file = gen_faq_config(tmpdirname, vocab_file, faq_file)
select_faq_responses(
question='Is Enrique the prettiest person in town?',
faq_model=train_model(configs_file, download=True)
) == ['<NAME>']
test_faq_response_with_minimum_faqs_in_dataframe_fail_case()
test_faq_response_with_minimum_faqs_in_dataframe_success_case()
test_faq_response_with_minimum_answers_vocab_success_case()
# -
#export
def select_squad_responses(
contexts, squad_model, question, best_results=1
):
'''Calls Deeppavlov BERT and RNET Context Question Answering
'''
responses = contexts.context.apply(
lambda context: squad_model([context], [question])
).values
logging.debug(f'Responses: {responses}')
top_responses = [
r[0][0] for r in sorted(responses, key=lambda x: -1 * x[2][0])
[:best_results]
]
logging.debug(f'Top Responses: {top_responses}')
return responses, top_responses
# +
#test select_squad_responses
import tempfile
from shutil import copyfile
empty = {'topic': [], 'context': []}
spacex = {
'topic': ['SpaceX'],
'context':
[
'''Space Exploration Technologies Corp., trading as SpaceX, is an American aerospace manufacturer and space transportation
services company headquartered in Hawthorne, California. It was founded in 2002 by <NAME> with the goal of reducing space
transportation costs to enable the colonization of Mars. SpaceX has developed several launch vehicles, the Starlink satellite
constellation, and the Dragon spacecraft. It is widely considered among the most successful private spaceflight companies.'''
]
}
intekglobal = {
'topic': ['Intekglobal', 'InG'],
'context':
[
'Intekglobal has its headquarters located in TJ',
'Intekglobal is in the north of mexico'
]
}
def assert_squad_model(
contexts, squad_model, question, expected_responses, **args
):
responses, top_responses = select_squad_responses(
contexts=pd.DataFrame(contexts),
squad_model=squad_model,
question=question,
**args
)
assert top_responses == expected_responses
def test_squad_bert():
bert = build_model(configs.squad.squad_bert, download=True)
assert_squad_model(
empty,
bert,
'Is an empty response expected?',
expected_responses=[],
best_results=2
)
assert_squad_model(
spacex, bert, 'Who founded SpaceX?', expected_responses=['<NAME>']
)
assert_squad_model(
intekglobal,
bert,
'Where is Intekglobal located?',
expected_responses=['north of mexico','TJ'],
best_results=2
)
def test_squad_rnet():
bert = build_model(configs.squad.squad, download=True)
assert_squad_model(
empty,
bert,
'Is an empty response expected?',
expected_responses=[],
best_results=5
)
assert_squad_model(
spacex, bert, 'Who founded SpaceX?', expected_responses=['<NAME>']
)
assert_squad_model(
intekglobal,
bert,
'Where is Intekglobal located?',
expected_responses=['north of mexico','TJ'],
best_results=2
)
test_squad_bert()
test_squad_rnet()
del spacex, empty, intekglobal
# +
#export
def load_qa_models(
config_rnet=configs.squad.squad,
config_bert=configs.squad.squad_bert,
config_tfidf=configs.faq.tfidf_logreg_en_faq,
download=True
):
''' Load the squad and faq models
INPUT:\n
- config_rnet -> path to json config file
- config_bert -> path to json config file
- config_tfidf -> path to json config file
- download -> download files (True/False)
The default for the config files are the deeppavlov config files. The default download is True.
'''
qa_models = {
'squad':
{
'rnet': build_model(config_rnet, download=download),
'bert': build_model(config_bert, download=download)
},
'faq': {
'tfidf': train_model(config_tfidf, download=download)
}
}
return qa_models
def get_responses(contexts, question, qa_models, nb_squad_results=1):
''' Generates responses from a question\n
INPUT: \n
- question -> question string \n
- contexts -> list of contexts
- qa_models -> dictionary of available models (see load_qa_models)
'''
responses = {'squad': defaultdict(list), 'faq': defaultdict(list)}
for squad_name, squad_model in qa_models['squad'].items():
responses['squad'][squad_name] = select_squad_responses(
contexts, squad_model, question, best_results=nb_squad_results
)[1]
for faq_name, faq_model in qa_models['faq'].items():
responses['faq'][faq_name] = select_faq_responses(faq_model, question)
return question, responses
# +
# test get_responses
import tempfile
from shutil import copyfile
intekglobal_context = {
'topic': ['Intekglobal', 'InG'],
'context':
[
'Intekglobal has its headquarters located in TJ',
'Intekglobal is in the north of mexico'
]
}
intekglobal_faqs = {
'Question': ['Is Intekglobal an IT company?', 'Where can I apply?'],
'Answer':
['Yes it is!', 'Please refer the our website for further information']
}
def mock_faq_files(tmpdirname, faqs):
faq_files = {
'data': path.join(tmpdirname, 'temp_faq.csv'),
'config': path.join(tmpdirname, 'temp_config_faq.json')
}
pd.DataFrame(faqs).to_csv(faq_files['data'], index=False)
copyfile(configs.faq.tfidf_logreg_en_faq, faq_files['config'])
updates_faq_config_file(
configs_path=faq_files['config'],
dataset_reader={'data_path': faq_files['data']}
)
return faq_files
def test_get_intekglobal_responses():
with tempfile.TemporaryDirectory() as tmpdirname:
faq_files = mock_faq_files(tmpdirname, intekglobal_faqs)
qa_models = load_qa_models(
config_tfidf=faq_files['config'], download=False
)
question, responses = get_responses(
pd.DataFrame(intekglobal_context),
'Where is Intekglobal?',
qa_models,
nb_squad_results=2
)
logging.debug(f' Question: {question}')
logging.debug(f" Responses: {responses}")
assert all(
response in ('north of mexico', 'TJ', 'Yes it is!')
for model_responses in responses['squad'].values()
for response in model_responses
)
def test_get_responses_with_empty_context():
with tempfile.TemporaryDirectory() as tmpdirname:
min_faqs = {
'Question':
['Minimum number of questions?', 'This is the other question?'],
'Answer': ['Two', 'yes']
}
faq_files = mock_faq_files(tmpdirname, min_faqs)
qa_models = load_qa_models(
config_tfidf=faq_files['config'], download=False
)
empty_context = {'topic': [], 'context': []}
question, responses = get_responses(
pd.DataFrame(empty_context),
'What is the minimun number of FAQ questions',
qa_models,
nb_squad_results=2
)
logging.debug(f' Question: {question}')
logging.debug(f' Responses: {responses}')
assert responses['faq']['tfidf'] == ['Two']
test_get_intekglobal_responses()
test_get_responses_with_empty_context()
del intekglobal_context
# +
#export
def format_responses(dict_responses):
'''Format question-response pair\n
INPUT:\n
- dictionary of responses \n
OUTPUT:\n
- list of flattened responses\n
- response as string
'''
flatten_responses = [
res for model_responses in dict_responses.values()
for res in model_responses.values()
]
flatten_responses = [r for res in flatten_responses for r in res]
logging.debug(flatten_responses)
formatted_response = f'\n Answers:\n\n'
for k, res in enumerate(set(flatten_responses)):
formatted_response += f'{k+1}: {res}\n'
logging.debug(formatted_response)
return flatten_responses, formatted_response
# -
#test
def test_format_responses():
dict_responses = {
'sq': {
'1': ['sq_11', 'sq_12'],
'2': ['sq_21']
},
'fq': {
'3': ['fq_11'],
'4': ['fq_21', 'fq_22']
}
}
flatten_responses, formatted_response = format_responses(
dict_responses=dict_responses
)
expected_arr =[
'sq_11', 'sq_12', 'sq_21', 'fq_11', 'fq_21', 'fq_22'
]
assert flatten_responses == expected_arr
assert all(res in formatted_response for res in expected_arr)
test_format_responses()
# +
#export
def get_input(text):
'''This redundancy is needed for testing'''
return input(text)
def question_response(data, qa_models, num_returned_values_per_squad_model=1):
''' Receive response and call get_response()
'''
question = get_input('Introduce question:\n')
_, responses = get_responses(
data['context']['df'], question, qa_models, nb_squad_results=1
)
_,formatted_responses = format_responses(responses)
return question, formatted_responses
# +
##Test FAQ dialog system's part
# +
import tempfile
from unittest.mock import patch
from shutil import copyfile
from collections import defaultdict
def mock_faq_files(tmpdirname, faqs, faq_dic):
faq_dic['path'] = path.join(tmpdirname, 'temp_faq.csv')
faq_dic['config'] = path.join(tmpdirname, 'temp_config_faq.json')
faq_dic['df'] = pd.DataFrame(faqs)
faq_dic['df'].to_csv(faq_dic['path'], index=False)
copyfile(configs.faq.tfidf_logreg_en_faq, faq_dic['config'])
updates_faq_config_file(
configs_path=faq_dic['config'],
dataset_reader={'data_path': faq_dic['path']}
)
def mock_context_file(tmpdirname, contexts, context_dic):
context_dic['path'] = path.join(tmpdirname, 'temp_context.csv')
context_dic['df'] = pd.DataFrame(contexts)
context_dic['df'].to_csv(context_dic['path'], index=False)
@patch('__main__.get_input')
def test_context_response_with_no_updates(mock_input):
mock_input.side_effect = ['Who is <NAME>?']
data = {'context': defaultdict(str), 'faq': defaultdict(str)}
contexts = {
'context':
[
'Intekglobal has its headquarters located in TJ',
'In Intekglobal we care about you',
'''<NAME> is one of the smartest minds on the planet,
he currently works as Intekglobal employee'''
],
'topic': ['headquarters', 'mission', 'Enrique\'s biography']
}
faqs = {
'Question':
['Minimum number of questions?', 'This is the other question?'],
'Answer': ['Two', 'yes']
}
with tempfile.TemporaryDirectory() as tmpdirname:
mock_faq_files(tmpdirname, faqs, data['faq'])
mock_context_file(tmpdirname, contexts, data['context'])
qa_models = load_qa_models(
config_tfidf=data['faq']['config'], download=False
)
question,responses = question_response(data, qa_models)
logging.debug(f' {question}')
logging.debug(f' {responses}')
assert 'Who is <NAME>?' == question
assert 'one of the smartest minds on the planet' in responses
test_context_response_with_no_updates()
# -
#export
def new_question_answer(data, qa_models):
''' Asks for a new question-answer pair; store the result in the
faq dataframe and retrain the faq-model\n
INPUT:\n
- dictionary of data
- dictionary of question-answer models
OUTPUT:\n
- None: Updates the dictionaries of data and models
'''
question = get_input('Introduce question:\n')
new_faq = pd.DataFrame(
{
'Question': [question],
'Answer': [get_input('Introduce the answer:\n')]
}
)
data['faq']['df'] = data['faq']['df'].append(new_faq)
data['faq']['df'].to_csv(data['faq']['path'], index=False)
qa_models['faq']['tfidf'] = train_model(
data['faq']['config'], download=False
)
logging.info('FAQ dataset and model updated..')
# +
#tests
@patch('__main__.get_input')
def test_new_question_answer(mock_input):
question = 'What is Intekglobal?'
new_answer = 'Intekglobal is one of the best companies in the world'
mock_input.side_effect = [question, new_answer]
data = {'context': defaultdict(str), 'faq': defaultdict(str)}
faqs = {
'Question': ['Who owns Tesla Company?', 'Is this is heaven?'],
'Answer': [
'<NAME> is the owner of Tesla', 'No, it is life on earth'
]
}
with tempfile.TemporaryDirectory() as tmpdirname:
mock_faq_files(tmpdirname, faqs, data['faq'])
qa_models = load_qa_models(
config_tfidf=data['faq']['config'], download=False
)
new_question_answer(data, qa_models)
updated_faq = pd.read_csv(data['faq']['path'])
assert updated_faq[updated_faq['Answer'] == new_answer].shape[0] == 1
test_new_question_answer()
# +
#export
def new_context(data):
''' Stores the new context in the context dataframe\n
INPUT:\n
- dictionary of data
OUTPUT:\n
- None: Updates the dictionary of data
'''
new_context = pd.DataFrame(
{
'topic': [get_input('Give context a title:\n')],
'context': [get_input('Introduce the context:\n')]
}
)
data['context']['df'] = data['context']['df'].append(new_context)
data['context']['df'].to_csv(data['context']['path'], index=False)
logging.info('contexts dataset updated..')
# +
@patch('__main__.get_input')
def test_new_context(mock_input):
data = {'context': defaultdict(str), 'faq': defaultdict(str)}
new_topic = 'AI Tool & Chatbot Development'
new_context_str = '''
A chatbot is an important tool for simulating intelligent conversations with humans.
Intekglobal chatbots efficiently live message on platforms such as Facebook Messenger,
Slack, and Telegram. But chatbots are more than just a cool technology advancement.
'''
contexts = {
'context':
[
'''One of the greatest punk rock bands from all the time
is the Ramones.
'''
],
'topic': ['Ramones']
}
mock_input.side_effect = [new_topic, new_context_str]
with tempfile.TemporaryDirectory() as tmpdirname:
logging.debug(str(new_context))
mock_context_file(tmpdirname, contexts, data['context'])
new_context(data)
updated_faq = pd.read_csv(data['context']['path'])
assert updated_faq[updated_faq.topic == new_topic].shape[0] == 1
test_new_context()
# +
#export
def set_minimal_faq_questions(data):
''' Sets the faq configurations that assure a proper operation.
If inexistent, a non-empty dataframe for faq is created with 'Question' and 'Answer as columns'
'''
if data['df'].shape[0] > 1:
return
minimal_questions = [
'Is this the Intekglobal Dialog System?',
'What is the purpose of these two automated questions?'
]
minimal_answers = [
'This is a default reponse of the Dialog System, ' +
'please populate your dataset with better responses',
'The purpose of this automated answer is to initalize the FAQ system, if you are '
+
'seeing this, you probably need to feed your datasets with more samples'
]
minimal_faqs_df = pd.DataFrame(
{
'Question': minimal_questions,
'Answer': minimal_answers
}
)
data['df'] = pd.concat([data['df'], minimal_faqs_df])
data['df'].to_csv(data['path'], index=False)
logging.info(f' File created at {data["path"]}')
def set_minimal_contexts(data):
''' Sets the context configurations that assure a proper operation.
If inexistent, a empy dataframe is created with 'topic' and 'context' as columns
'''
if data['df'].shape[0] > 0:
return
minimal_context_df = pd.DataFrame({'topic': [], 'context': []})
data['df'] = minimal_context_df
data['df'].to_csv(data['path'], index=False)
logging.info(f' File created at {data["path"]}')
def set_data_dict(file, data, question_type, data_dir):
'''Creates unexistent files
'''
data['path'] = file if file is not None else path.join(
data_dir, question_type + '_data.csv'
)
data['df'] = pd.read_csv(data['path']) if path.isfile(data['path']
) else pd.DataFrame()
if question_type == 'faq':
set_minimal_faq_questions(data)
if question_type == 'context':
set_minimal_contexts(data)
def load_and_prepare_data(context_data_file, faq_data_file, data, configs_faq):
'''Calls the context and faq configuration routines.
If dataframe files missing, it will create them in data directory.
If the data frames are provided they must have the following columns for proper functioning:
- context: 'topic', 'context'
- faq: 'Question, 'Answer'
'''
PARENT_DIR = popen('$PWD').read().strip()
if faq_data_file or context_data_file is None:
DATA_DIR = path.join(PARENT_DIR, 'data')
if not path.isdir(DATA_DIR):
mkdir(DATA_DIR)
logging.info(f'Data directory created at {DATA_DIR}')
if configs_faq is None:
configs_faq = configs.faq.tfidf_logreg_en_faq
data['faq']['config'] = configs_faq
set_data_dict(
file=faq_data_file,
data=data['faq'],
question_type='faq',
data_dir=DATA_DIR
)
set_data_dict(
file=context_data_file,
data=data['context'],
question_type='context',
data_dir=DATA_DIR
)
updates_faq_config_file(
configs_path=configs_faq,
dataset_reader={'data_path': data['faq']['path']}
)
# +
#tests
import tempfile,logging
import pandas as pd
from collections import defaultdict
from shutil import rmtree
from os import path,popen
from unittest.mock import patch
def test_set_minimal_faqs_with_more_than_one_question():
with tempfile.TemporaryDirectory() as tmpdirname:
data_file = path.join(tmpdirname, 'tmp_data.csv')
questions = ['a?', 'b?']
answers = ['a', 'b']
df = pd.DataFrame({'Question': questions, 'Answer': answers})
df.to_csv(data_file, index=False)
data = {'df': df, 'path': data_file}
set_minimal_faq_questions(data)
assert data['df'].shape[0] == 2
def test_set_minimal_faqs_with_less_than_two_questions():
with tempfile.TemporaryDirectory() as tmpdirname:
data_file = path.join(tmpdirname, 'tmp_data.csv')
questions = ['a?']
answers = ['a']
df = pd.DataFrame({'Question': questions, 'Answer': answers})
df.to_csv(data_file, index=False)
data = {'df': df, 'path': data_file}
assert data['df'].shape[0] == 1
set_minimal_faq_questions(data)
assert data['df'].shape[0] == 3
assert any(
data['df'].Question == 'Is this the Intekglobal Dialog System?'
)
def test_set_minimal_contexts():
with tempfile.TemporaryDirectory() as tmpdirname:
data_file = path.join(tmpdirname, 'tmp_data.csv')
data = {'df': pd.DataFrame(), 'path': data_file}
set_minimal_contexts(data)
assert path.isfile(data['path'])
assert all(data['df'].columns == ['topic', 'context'])
def test_set_data_dict_no_file():
with tempfile.TemporaryDirectory() as tmpdirname:
data = {'context': defaultdict(str)}
set_data_dict(
file=None,
data=data['context'],
data_dir=tmpdirname,
question_type='context'
)
logging.debug(data)
assert path.isfile(data['context']['path'])
@patch('__main__.popen')
def test_load_and_prepare_data(mock_popen):
with tempfile.TemporaryDirectory() as tmpdirname:
mock_popen("$PWD").read().strip.side_effect = [tmpdirname]
data = {'context': defaultdict(str), 'faq': defaultdict(str)}
load_and_prepare_data(
context_data_file=None,
faq_data_file=None,
data=data,
configs_faq=None
)
data_dir = path.join(tmpdirname, 'data')
assert path.isdir(data_dir)
test_set_minimal_faqs_with_more_than_one_question()
test_set_minimal_faqs_with_less_than_two_questions()
test_set_minimal_contexts()
test_set_data_dict_no_file()
test_load_and_prepare_data()
# -
| nbs/01_settings.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Quiz 1
#
# This is an extension of another notebook: Color Selection. Modify the values of Red,Green and Blue Thresholds so the image you read, looks like below <br/>
#
# <img src="../img/Ftest-after.jpg"/>
# +
# Import required items
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
#Read the Image
image = mpimg.imread('../img/test.jpg')
#print('The following image is:',type(image), 'The dimensions are: ', image.shape)
# Lets grab x and y size and make a copy of image
ysize = image.shape[0]
xsize = image.shape[1]
# ALWAYS make a copy rather than using '=' this will help avoid changing unintended variables
color_image = np.copy(image)
#plt.imshow(color_image)
######
#UPDATE these to get image as defined earlier as RESULT
red_threshold = 0
green_threshold = 0
blue_threshold = 0
#make a list of all three together
rgb_threshold = [red_threshold,green_threshold,blue_threshold]
thresholds = (image[:,:,0] < rgb_threshold[0]) \
|(image[:,:,1] < rgb_threshold[1]) \
|(image[:,:,2] < rgb_threshold[2])
color_image[thresholds] = [0,0,0]
plt.imshow(color_image)
mpimg.imsave('../img/test-after.jpg',color_image)
# -
# BONUS: Print both side by side for better representation
sidebyside,ax = plt.subplots(1,2)
ax[0].imshow(image)
ax[1].imshow(color_image)
| 01-Computer-Vision-Fundamentals/.ipynb_checkpoints/Quiz 1.1 Color Selection-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from selenium import webdriver
from selenium.webdriver.firefox.options import Options
from bs4 import BeautifulSoup
import numpy as np
import pandas as pd
import time
import sys
import requests as re
# +
# options = Options()
# options.set_headless()
# #Firefoxを操作
# driver = webdriver.Firefox(options=options, executable_path="/Users/takayuki/Library/Application Support/WebDriverManager/gecko/v0.26.0/geckodriver-v0.26.0-macos/geckodriver")
# username = "<EMAIL>"
# password = "<PASSWORD>"
# +
# Login Page
# driver.get('https://r.nikkei.com/login')
# time.sleep(2)
# driver.save_screenshot("ss1.png")
# +
# login_username = driver.find_element_by_id("LA7010Form01:LA7010Email")
# login_username.clear()
# login_username.send_keys(username)
# login_password = driver.find_element_by_id("LA7010Form01:LA<PASSWORD>Password")
# login_password.clear()
# login_password.send_keys(password)
# driver.find_element_by_class_name('btnM1').click() # Login button click
# time.sleep(2)
# driver.save_screenshot("ss2.png")
# +
# Search word (ex. 院内感染, 200 articles)
target_url=re.get('https://sitesearch.asahi.com/.cgi/sitesearch/sitesearch.pl?Keywords=院内感染&Searchsubmit2=検索&Searchsubmit=検索&iref=pc_gnavi')
# driver.get(target_url)
time.sleep(2)
# html = driver.page_source
# +
# Articles count
# sel0 = soup.find_all("p",attrs={"class":"search__result-count"})
# for i in sel0:
# count= int(i.text)
# print (count)
# +
# span = soup.find("span", id="SearchResult_Headline" )
# sel0 = soup.find("span", attrs={"class":"SearchResult_Headline"})
# # Article's title
# # class with "digital" or "shimbun"
# sel1 = soup.find_all(True, attrs={"class":["digital", "shimbun"]})
# # Article datetime
# sel2 = soup.find_all("span",attrs={"class":"Date"})
# print(sel0.text)
# -
# +
# CSV file output
out_number = []
out_url = []
out_title = []
out_datetime = []
# if len(sel1) != len(sel2):
# sys.exit()
# -
soup = BeautifulSoup(target_url.content, "html.parser")
elem = soup.find_all(True, attrs={"class":["digital", "shimbun"]})
for i in elem:
link = i.find('a')
out_title_temp = link.text.strip()
out_url_temp = link['href']
out_datetime = i.find_all("span", attrs={"class":"Date"})
out_title.append(out_title_temp)
out_url.append(out_url_temp)
# +
# for i in range(0, len(sel1)):
# cnt = str(i+1)
# out_url_temp = sel1[i].a.get("href")
# out_title_temp = sel1[i].text.strip()
# out_number.append(cnt)
# out_url.append(out_url_temp)
# out_title.append(out_title_temp)
# out_datetime.append(sel2[i].find('Date'))
# -
# print(out_title)
# Result output
resdata = np.vstack([out_number, out_title, out_datetime, out_url])
df = pd.DataFrame(data=resdata).T
df.to_csv('result_asahi.csv', header=False, index=False, mode='a')
driver.quit()
| articles/asahi_scrape.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
# two approaches
# 1. Latent Dirichlet Allocation (LDA)
# 2. Non-Negative Matrix Factorization (NMF)
# -
import os
os.system('export LC_ALL=en_US.UTF-8')
os.system('export LANG=en_US.UTF-8')
# imports
import pandas as pd
import numpy as np
import nltk
import spacy
import matplotlib.pyplot as plt
from sklearn.decomposition import LatentDirichletAllocation, NMF
# %matplotlib inline
nltk.download('stopwords')
from nltk.corpus import stopwords
eng_stopwords = set(stopwords.words('english'))
# read data
local_data_root = os.getcwd()
files = os.listdir(local_data_root)
df_files = [local_data_root+'/'+x for x in files if (x.startswith('t')) & (x.endswith('.zip'))]
print df_files
data = {}
for fn in df_files:
print 'unzip file: %s' % (fn)
os.system('unzip {}'.format(fn))
name = fn.split('.')[0]
name = name.split('/')[-1]
print 'reading file: %s' % (name+'.csv')
df = pd.read_csv(local_data_root+'/'+name+'.csv')
data[name] = df
data.keys()
data['train'].shape
print data['train'].head()
print data['train']['author'].value_counts()
print data['train']['text'][0]
train = data['train'].copy()
import plotly.plotly as py
import plotly
plotly.tools.set_credentials_file(username='nathanv',api_key='<KEY>')
import plotly.graph_objs as go
import plotly.tools as tls
tls.embed('https://plot.ly/~cufflinks/8')
z = {'EAP': '<NAME>', 'MWS': '<NAME>', 'HPL': 'HP Lovecraft'}
# +
data = [go.Bar(
x = train.author.map(z).unique(),
y = train.author.value_counts().values,
marker= dict(colorscale='Jet',
color = train.author.value_counts().values
),
text='Text entries attributed to Author'
)]
layout = go.Layout(
title='Target variable distribution'
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic-bar')
# -
all_words = train['text'].str.split(expand=True)
print type(all_words)
print all_words.shape
print all_words.head()
all_words = train['text'].str.split(expand=True).unstack()
print type(all_words)
print all_words.shape
print all_words.head()
all_words = train['text'].str.split(expand=True).unstack().value_counts()
print type(all_words)
print all_words.shape
print all_words.head()
print all_words[2:50]
# +
data = [go.Bar(
x=all_words.index.values[2:50],
y=all_words.values[2:50],
marker=dict(colorscale='Jet', color=all_words.values[2:50]),
text='Word Counts')]
layout = go.Layout(
title='Top 50 (Uncleaned) Word frequencies in the training dataset')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic-bar')
# -
# WordCloud to visualize each author's work
print train.columns
print train['author'].value_counts()
eap = train[train['author'] == 'EAP']['text'].values
mws = train[train['author'] == 'MWS']['text'].values
hpl = train[train['author'] == 'HPL']['text'].values
print len(eap), len(mws), len(hpl), type(hpl)
from wordcloud import WordCloud, STOPWORDS
# The wordcloud for HP Lovecraft
plt.figure(figsize=(16,13))
wc = WordCloud(background_color="black", max_words=10000,
stopwords=STOPWORDS, max_font_size= 40)
wc.generate(" ".join(hpl))
plt.title("HP Lovecraft", fontsize=20)
# plt.imshow(wc.recolor( colormap= 'Pastel1_r' , random_state=17), alpha=0.98)
plt.imshow(wc.recolor( colormap= 'Pastel2' , random_state=17), alpha=0.98)
plt.axis('off')
# wordcloud for <NAME>
plt.figure(figsize=(20,18))
plt.subplot(211)
wc = WordCloud(background_color="black", max_words=10000,
stopwords=STOPWORDS, max_font_size= 40)
wc.generate(" ".join(eap))
plt.title("<NAME>")
plt.imshow(wc.recolor( colormap= 'PuBu' , random_state=17), alpha=0.9)
plt.axis('off')
# wordcloud for <NAME>
plt.figure(figsize=(20,18))
plt.subplot(211)
wc = WordCloud(background_color="black", max_words=10000,
stopwords=STOPWORDS, max_font_size= 40)
wc.generate(" ".join(mws))
plt.title("<NAME>")
plt.imshow(wc.recolor( colormap= 'viridis' , random_state=17), alpha=0.9)
plt.axis('off')
# Preprocessing
# tokenization
import nltk
nltk.download('punkt')
import string
first_text = train['text'].values[0]
print first_text
print '='*110
print first_text.split(' ')
print '='*110
first_text_tokenized = nltk.word_tokenize(first_text)
print first_text_tokenized
print '='*110
first_text_tokenized_Wpunkt = [x for x in first_text_tokenized if x not in string.punctuation]
print first_text_tokenized_Wpunkt
print '='*110
first_text_tokenized_Wpunkt_Wstop = [x for x in first_text_tokenized_Wpunkt if x not in eng_stopwords]
print first_text_tokenized_Wpunkt_Wstop
print '='*110
print 'length of original text: %d' % (len(first_text_tokenized))
print 'length of text post removal of punctuation: %d' % (len(first_text_tokenized_Wpunkt))
print 'length of text post removal of stopwords and punctuation: %d' % (len(first_text_tokenized_Wpunkt_Wstop))
string.punctuation
# +
# stemming and lematization
# -
from nltk.stem import PorterStemmer, WordNetLemmatizer
nltk.download('wordnet')
base_char = 'The stemmed form of {} is {}'
print base_char.format('running', PorterStemmer().stem('running'))
print base_char.format('runs', PorterStemmer().stem('runs'))
print base_char.format('run', PorterStemmer().stem('run'))
print base_char.format('ran', PorterStemmer().stem('ran'))
print base_char.format('leaves', PorterStemmer().stem('leaves'))
print base_char.format('running', WordNetLemmatizer().lemmatize('running'))
print base_char.format('runs', WordNetLemmatizer().lemmatize('runs'))
print base_char.format('run', WordNetLemmatizer().lemmatize('run'))
print base_char.format('ran', WordNetLemmatizer().lemmatize('ran'))
print base_char.format('leaves', WordNetLemmatizer().lemmatize('leaves'))
# vectorizing text
from sklearn.feature_extraction.text import CountVectorizer
sentence = ["I love to eat Burgers",
"I love to eat Fries"]
vectorizer = CountVectorizer(min_df=0)
sentence_t = vectorizer.fit_transform(sentence)
print sentence_t.toarray()
print vectorizer.get_feature_names()
# Topic Modelling via LDA and NMF
def print_top_words(model, feature_names, n_top_words):
for index, topic in enumerate(model.components_):
message = '\nTopic #{}:'.format(index)
message += " ".join([feature_names[i] for i in topic.argsort()[:-n_top_words - 1: -1]])
print message
print '='*70
# +
# sklearn's countvectroizer class does tokenizing and stopwords removal by removing terms with single characters.
# It also lower cases all terms by default
# +
# Extending the CountVectorizer class with a lemmatizer
# -
lemm = WordNetLemmatizer()
class LemmaCountVectorizer(CountVectorizer):
def build_analyzer(self):
analyzer = super(LemmaCountVectorizer, self).build_analyzer()
return lambda doc: (lemm.lemmatize(w) for w in analyzer(doc))
text = train.text.values.tolist()
print len(text)
print text[0]
tf_vectorizer = LemmaCountVectorizer(max_df=0.95, min_df=2, stop_words='english', decode_error='ignore')
tf = tf_vectorizer.fit_transform(text)
tf_values = tf.toarray()
print type(tf_values)
print len(tf_values)
print tf_values[0]
print len(tf_values[0])
print tf_vectorizer.get_feature_names()[:5]
# +
# top 50 words in terms of frequencies
# -
feat_names = tf_vectorizer.get_feature_names()
count_vec = np.asarray(tf.sum(axis=0)).ravel()
zipped = list(zip(feat_names, count_vec))
zipped = sorted(zipped, key=lambda x: x[1], reverse=True)
for x in zip(*zipped):
print list(x)
x = []
y = []
for item in zipped:
x.append(item[0])
y.append(item[1])
print x[:5]
print y[:5]
x, y = (list(x) for x in zip(*zipped))
print x[:5]
print y[:5]
print len(x)
X = np.concatenate([x[0:15], x[-16:-1]])
Y = np.concatenate([y[0:15], y[-16:-1]])
print X
print Y
# plot top 50 word frequencies
data = [go.Bar(
x = x[:50],
y = y[:50],
marker = dict(colorscale='Jet', color=y[:50]),
text = 'Word Counts'
)]
layout = go.Layout(title='Top 50 Word frequencies after Preprocessing')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic-bar')
# bottom 100 word frequencies
data = [go.Bar(
x = x[-100:],
y = y[-100:],
marker = dict(colorscale='Jet', color=y[-100:]),
text = 'Word Counts')]
layout = go.Layout(title='Bottom 100 word frequencies after preprocessing')
fig = go.Figure(data=data, layout=layout)
py.iplot(fig, filename='basic-bar')
# +
# Latent Dirichlet Allocation
# +
# parameters to tune = n_components (number of topics), alpha (dirichlet prior for document-topic), beta (dirichlet prior for topic-word prior)
# +
# n_components is identified based on KMeans + Latent Semantic Analysis Scheme
# whereby the number of Kmeans clusters and number of LSA dimensions were iterated through and the best silhouette mean score
# -
from sklearn.decomposition import LatentDirichletAllocation
lda = LatentDirichletAllocation(n_components=11, learning_method = 'online',
learning_offset = 50.,
random_state = 0)
lda.fit(tf)
n_top_words = 40
print("\nTopics in LDA model: ")
tf_feature_names = tf_vectorizer.get_feature_names()
print_top_words(lda, tf_feature_names, n_top_words)
first_topic = lda.components_[0]
second_topic = lda.components_[1]
third_topic = lda.components_[2]
fourth_topic = lda.components_[3]
print len(first_topic)
print len(second_topic)
first_topic_words = [tf_feature_names[i] for i in first_topic.argsort()[:-50 - 1 :-1]]
second_topic_words = [tf_feature_names[i] for i in second_topic.argsort()[:-50 - 1 :-1]]
third_topic_words = [tf_feature_names[i] for i in third_topic.argsort()[:-50 - 1 :-1]]
fourth_topic_words = [tf_feature_names[i] for i in fourth_topic.argsort()[:-50 - 1 :-1]]
print first_topic_words
firstcloud = WordCloud(stopwords=STOPWORDS,
background_color='black',
width=2500, height=1000).generate(" ".join(first_topic_words))
plt.imshow(firstcloud)
plt.axis('off')
plt.show()
secondcloud = WordCloud(stopwords=STOPWORDS, background_color='black', width=2500, height=1000).generate(" ".join(second_topic_words)
)
plt.imshow(secondcloud)
plt.axis('off')
plt.show()
| src/topic_modelling_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:py3-TF2.0]
# language: python
# name: conda-env-py3-TF2.0-py
# ---
# # Exercises
#
# ### 9. Adjust the learning rate. Try a value of 0.02. Does it make a difference?
#
# ** Solution **
#
# First, we have to define a custom optimizer (as we did in the TensorFlow intro).
#
# We create the custom optimizer with:
#
# custom_optimizer = tf.keras.optimizers.Adam(learning_rate=0.0001)
#
# Then we change the respective argument in model.compile to reflect this:
#
# model.compile(optimizer=custom_optimizer, loss='sparse_categorical_crossentropy', metrics=['accuracy'])
#
#
# While Adam adapts to the problem, if the orders of magnitude are too different, it may not have time to adjust accordingly. We start overfitting before we can reach a neat solution.
#
# Therefore, for this problem, even 0.02 is a **HIGH** starting learning rate. What if you try a learning rate of = 1?
#
# It's a good practice to try 0.001, 0.0001, and 0.00001. If it makes no difference, pick whatever, otherwise it makes sense to fiddle with the learning rate.
# # Deep Neural Network for MNIST Classification
#
# We'll apply all the knowledge from the lectures in this section to write a deep neural network. The problem we've chosen is referred to as the "Hello World" of deep learning because for most students it is the first deep learning algorithm they see.
#
# The dataset is called MNIST and refers to handwritten digit recognition. You can find more about it on <NAME>'s website (Director of AI Research, Facebook). He is one of the pioneers of what we've been talking about and of more complex approaches that are widely used today, such as covolutional neural networks (CNNs).
#
# The dataset provides 70,000 images (28x28 pixels) of handwritten digits (1 digit per image).
#
# The goal is to write an algorithm that detects which digit is written. Since there are only 10 digits (0, 1, 2, 3, 4, 5, 6, 7, 8, 9), this is a classification problem with 10 classes.
#
# Our goal would be to build a neural network with 2 hidden layers.
# ## Import the relevant packages
# +
import numpy as np
import tensorflow as tf
# TensorFLow includes a data provider for MNIST that we'll use.
# It comes with the tensorflow-datasets module, therefore, if you haven't please install the package using
# pip install tensorflow-datasets
# or
# conda install tensorflow-datasets
import tensorflow_datasets as tfds
# these datasets will be stored in C:\Users\*USERNAME*\tensorflow_datasets\...
# the first time you download a dataset, it is stored in the respective folder
# every other time, it is automatically loading the copy on your computer
# -
# ## Data
#
# That's where we load and preprocess our data.
# +
# remember the comment from above
# these datasets will be stored in C:\Users\*USERNAME*\tensorflow_datasets\...
# the first time you download a dataset, it is stored in the respective folder
# every other time, it is automatically loading the copy on your computer
# tfds.load actually loads a dataset (or downloads and then loads if that's the first time you use it)
# in our case, we are interesteed in the MNIST; the name of the dataset is the only mandatory argument
# there are other arguments we can specify, which we can find useful
# mnist_dataset = tfds.load(name='mnist', as_supervised=True)
mnist_dataset, mnist_info = tfds.load(name='mnist', with_info=True, as_supervised=True)
# with_info=True will also provide us with a tuple containing information about the version, features, number of samples
# we will use this information a bit below and we will store it in mnist_info
# as_supervised=True will load the dataset in a 2-tuple structure (input, target)
# alternatively, as_supervised=False, would return a dictionary
# obviously we prefer to have our inputs and targets separated
# once we have loaded the dataset, we can easily extract the training and testing dataset with the built references
mnist_train, mnist_test = mnist_dataset['train'], mnist_dataset['test']
# by default, TF has training and testing datasets, but no validation sets
# thus we must split it on our own
# we start by defining the number of validation samples as a % of the train samples
# this is also where we make use of mnist_info (we don't have to count the observations)
num_validation_samples = 0.1 * mnist_info.splits['train'].num_examples
# let's cast this number to an integer, as a float may cause an error along the way
num_validation_samples = tf.cast(num_validation_samples, tf.int64)
# let's also store the number of test samples in a dedicated variable (instead of using the mnist_info one)
num_test_samples = mnist_info.splits['test'].num_examples
# once more, we'd prefer an integer (rather than the default float)
num_test_samples = tf.cast(num_test_samples, tf.int64)
# normally, we would like to scale our data in some way to make the result more numerically stable
# in this case we will simply prefer to have inputs between 0 and 1
# let's define a function called: scale, that will take an MNIST image and its label
def scale(image, label):
# we make sure the value is a float
image = tf.cast(image, tf.float32)
# since the possible values for the inputs are 0 to 255 (256 different shades of grey)
# if we divide each element by 255, we would get the desired result -> all elements will be between 0 and 1
image /= 255.
return image, label
# the method .map() allows us to apply a custom transformation to a given dataset
# we have already decided that we will get the validation data from mnist_train, so
scaled_train_and_validation_data = mnist_train.map(scale)
# finally, we scale and batch the test data
# we scale it so it has the same magnitude as the train and validation
# there is no need to shuffle it, because we won't be training on the test data
# there would be a single batch, equal to the size of the test data
test_data = mnist_test.map(scale)
# let's also shuffle the data
BUFFER_SIZE = 10000
# this BUFFER_SIZE parameter is here for cases when we're dealing with enormous datasets
# then we can't shuffle the whole dataset in one go because we can't fit it all in memory
# so instead TF only stores BUFFER_SIZE samples in memory at a time and shuffles them
# if BUFFER_SIZE=1 => no shuffling will actually happen
# if BUFFER_SIZE >= num samples => shuffling is uniform
# BUFFER_SIZE in between - a computational optimization to approximate uniform shuffling
# luckily for us, there is a shuffle method readily available and we just need to specify the buffer size
shuffled_train_and_validation_data = scaled_train_and_validation_data.shuffle(BUFFER_SIZE)
# once we have scaled and shuffled the data, we can proceed to actually extracting the train and validation
# our validation data would be equal to 10% of the training set, which we've already calculated
# we use the .take() method to take that many samples
# finally, we create a batch with a batch size equal to the total number of validation samples
validation_data = shuffled_train_and_validation_data.take(num_validation_samples)
# similarly, the train_data is everything else, so we skip as many samples as there are in the validation dataset
train_data = shuffled_train_and_validation_data.skip(num_validation_samples)
# determine the batch size
BATCH_SIZE = 100
# we can also take advantage of the occasion to batch the train data
# this would be very helpful when we train, as we would be able to iterate over the different batches
train_data = train_data.batch(BATCH_SIZE)
validation_data = validation_data.batch(num_validation_samples)
# batch the test data
test_data = test_data.batch(num_test_samples)
# takes next batch (it is the only batch)
# because as_supervized=True, we've got a 2-tuple structure
validation_inputs, validation_targets = next(iter(validation_data))
# -
# ## Model
# ### Outline the model
# When thinking about a deep learning algorithm, we mostly imagine building the model. So, let's do it :)
# +
input_size = 784
output_size = 10
# Use same hidden layer size for both hidden layers. Not a necessity.
hidden_layer_size = 50
# define how the model will look like
model = tf.keras.Sequential([
# the first layer (the input layer)
# each observation is 28x28x1 pixels, therefore it is a tensor of rank 3
# since we don't know CNNs yet, we don't know how to feed such input into our net, so we must flatten the images
# there is a convenient method 'Flatten' that simply takes our 28x28x1 tensor and orders it into a (None,)
# or (28x28x1,) = (784,) vector
# this allows us to actually create a feed forward neural network
tf.keras.layers.Flatten(input_shape=(28, 28, 1)), # input layer
# tf.keras.layers.Dense is basically implementing: output = activation(dot(input, weight) + bias)
# it takes several arguments, but the most important ones for us are the hidden_layer_size and the activation function
tf.keras.layers.Dense(hidden_layer_size, activation='relu'), # 1st hidden layer
tf.keras.layers.Dense(hidden_layer_size, activation='relu'), # 2nd hidden layer
# the final layer is no different, we just make sure to activate it with softmax
tf.keras.layers.Dense(output_size, activation='softmax') # output layer
])
# -
# ### Choose the optimizer and the loss function
# +
# we define the optimizer we'd like to use,
# the loss function,
# and the metrics we are interested in obtaining at each iteration
custom_optimizer = tf.keras.optimizers.Adam(learning_rate=0.02)
model.compile(optimizer=custom_optimizer, loss='sparse_categorical_crossentropy', metrics=['accuracy'])
# -
# ### Training
# That's where we train the model we have built.
# +
# determine the maximum number of epochs
NUM_EPOCHS = 5
# we fit the model, specifying the
# training data
# the total number of epochs
# and the validation data we just created ourselves in the format: (inputs,targets)
model.fit(train_data, epochs=NUM_EPOCHS, validation_data=(validation_inputs, validation_targets), verbose =2)
# -
# ## Test the model
#
# As we discussed in the lectures, after training on the training data and validating on the validation data, we test the final prediction power of our model by running it on the test dataset that the algorithm has NEVER seen before.
#
# It is very important to realize that fiddling with the hyperparameters overfits the validation dataset.
#
# The test is the absolute final instance. You should not test before you are completely done with adjusting your model.
#
# If you adjust your model after testing, you will start overfitting the test dataset, which will defeat its purpose.
test_loss, test_accuracy = model.evaluate(test_data)
# We can apply some nice formatting if we want to
print('Test loss: {0:.2f}. Test accuracy: {1:.2f}%'.format(test_loss, test_accuracy*100.))
# Using the initial model and hyperparameters given in this notebook, the final test accuracy should be roughly around 97%.
#
# Each time the code is rerun, we get a different accuracy as the batches are shuffled, the weights are initialized in a different way, etc.
#
# Finally, we have intentionally reached a suboptimal solution, so you can have space to build on it.
| 17 - Deep Learning with TensorFlow 2.0/12_Deeper example/12_MNIST - Solutions/9. TensorFlow_MNIST_Learning_rate_Part_2_Solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Ray Serve - Integration with FastAPI
#
# © 2019-2022, Anyscale. All Rights Reserved
#
# 
#
# Ray Serve integrates well with other common [web serving frameworks](https://docs.ray.io/en/latest/serve/tutorials/web-server-integration.html).
#
# In this tutorial, we’ll cover how to deploy [XGBoost](https://xgboost.readthedocs.io/en/stable/) with [FastAPI](https://fastapi.tiangolo.com/) and Ray Serve. We'll use a simple XGBboost classifcation model to train, deploy it on Ray Serve , and access it via HTTP request on a FastAPI endpoint.
#
# FastAPI is a modern, fast (high-performance), web framework for building APIs with Python 3.6+ based on standard Python type hints.
#
# <img src="https://fastapi.tiangolo.com/img/logo-margin/logo-teal.png" width="40%" height="20%">
#
# This XGBoost model will be trained to predict the onset of diabetes using the pima-indians-diabetes dataset from the [UCI Machine Learning Repository website](https://raw.githubusercontent.com/jbrownlee/Datasets/master/pima-indians-diabetes.data.csv). This small dataset contains several numerical medical variables of eight different features related to diabetes, in addition to one target variable — Outcome. So, we’ll use XGBoost to model and solve a simple prediction problem. This tutorial is derived from our [blog](https://www.anyscale.com/blog/deploying-xgboost-models-with-ray-serve).
#
# Let's see how easy it is!
#
#
# +
import numpy as np
from xgboost import XGBClassifier
from sklearn.model_selection import train_test_split
from sklearn.metrics import accuracy_score
import pickle
import ray
from fastapi import FastAPI, Request
from ray import serve
# -
# ### Load the data
# Load the data
dataset = np.loadtxt('pima-indians-diabetes.data.csv', delimiter=",")
# split data into X and y
X = dataset[:, 0:8]
y = dataset[:, 8]
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=7)
# ### Define a utility function:
# * Creates XGBoost classifier
# * trains, fits, and saves the model
# * returns a model
def create_and_save_model():
# Instantiate a model, fit and train
xgb_model = XGBClassifier(use_label_encoder=False)
xgb_model.fit(X_train, y_train)
# saving the model
with open('xgb_model.pkl', 'wb') as f:
pickle.dump(xgb_model, f)
return xgb_model
# ### Create, fit and predict XGBoost model
model = create_and_save_model()
y_pred = model.predict(X_test)
predictions = [round(value) for value in y_pred]
accuracy = accuracy_score(y_test, predictions)
print("Accuracy: %.2f%%" % (accuracy * 100.0))
# ### Create a Ray Serve Deployment with FastAPI
app = FastAPI()
ray.init(namespace="xgbregressor")
serve.start()
@serve.deployment(num_replicas=2, route_prefix="/regressor")
@serve.ingress(app)
class XGBModel:
def __init__(self):
# loading the model
with open("xgb_model.pkl", "rb") as f:
self.model = pickle.load(f)
print("Pickled XGBoost model loaded")
@app.post("/")
async def predict(self, starlette_request:Request):
payload = await starlette_request.json()
print("Worker: received starlette request with data", payload)
# input_vector = [
# payload["Pregnancies"],
# payload["Glucose"],
# payload["BloodPressure"],
# payload["SkinThickness"],
# payload["Insulin"],
# payload["BMI"],
# payload["DiabetesPedigree"],
# payload["Age"],
# ]
# prediction = self.model.predict([np.array(input_vector)])[0]
prediction = round(self.model.predict([np.array(list(payload.values()))])[0])
return {"result": prediction}
XGBModel.deploy()
# ### List current deployments
print(serve.list_deployments())
# ### Send request to the FastAPI endpoint
# +
import requests
sample_request_inputs = [
{"Pregnancies": 6,
"Glucose": 148,
"BloodPressure": 72,
"SkinThickness": 35,
"Insulin": 0,
"BMI": 33.6,
"DiabetesPedigree": 0.625,
"Age": 50,
},
{"Pregnancies": 10,
"Glucose": 168,
"BloodPressure": 74,
"SkinThickness": 0,
"Insulin": 0,
"BMI": 38.0,
"DiabetesPedigree": 0.537,
"Age": 34,
},
{"Pregnancies": 10,
"Glucose": 39,
"BloodPressure": 80,
"SkinThickness": 0,
"Insulin": 0,
"BMI": 27.1,
"DiabetesPedigree": 1.441,
"Age": 57,
},
{"Pregnancies": 1,
"Glucose": 103,
"BloodPressure": 30,
"SkinThickness": 38,
"Insulin": 83,
"BMI": 43.3,
"DiabetesPedigree": 0.183,
"Age": 33,
}
]
# -
# Iterate our requests
for sri in sample_request_inputs:
response = requests.post("http://localhost:8000/regressor/", json=sri)
print(response.text)
ray.shutdown()
| ray-serve/07-Ray-Serve-and-FastAPI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %reload_ext autoreload
# %autoreload 2
# This requires to install fastprogress (pip install fastprogress).
#export
from nb_004a import *
from fastprogress import master_bar,progress_bar
# ## Test with training
#export
def fit(epochs, model, loss_fn, opt, data, callbacks=None, metrics=None, pbar=None):
cb_handler = CallbackHandler(callbacks)
cb_handler.on_train_begin()
if pbar is None: pbar = master_bar(range(epochs))
for epoch in pbar:
model.train()
cb_handler.on_epoch_begin()
for xb,yb in progress_bar(data.train_dl, parent=pbar):
xb, yb = cb_handler.on_batch_begin(xb, yb)
loss,_ = loss_batch(model, xb, yb, loss_fn, opt, cb_handler)
if cb_handler.on_batch_end(loss): break
if hasattr(data,'valid_dl') and data.valid_dl is not None:
model.eval()
with torch.no_grad():
*val_metrics,nums = zip(*[loss_batch(model, xb, yb, loss_fn, cb_handler=cb_handler, metrics=metrics)
for xb,yb in progress_bar(data.valid_dl, parent=pbar)])
val_metrics = [np.sum(np.multiply(val,nums)) / np.sum(nums) for val in val_metrics]
else: val_metrics=None
if cb_handler.on_epoch_end(val_metrics): break
cb_handler.on_train_end()
# +
#export
@dataclass
class Learner():
"Object that wraps together some data, a model, a loss function and an optimizer"
data:DataBunch
model:nn.Module
opt_fn:Callable=AdamW
loss_fn:Callable=F.cross_entropy
metrics:Collection[Callable]=None
true_wd:bool=True
wd:Floats=1e-6
train_bn:bool=True
path:str = 'models'
callback_fns:Collection[Callable]=None
layer_groups:Collection[nn.Module]=None
def __post_init__(self):
self.path = Path(self.path)
self.path.mkdir(parents=True, exist_ok=True)
self.model = self.model.to(self.data.device)
if not self.layer_groups: self.layer_groups = [self.model]
self.callback_fns = listify(self.callback_fns)
self.callbacks = []
def fit(self, epochs:int, lr:Floats, wd:Floats=None, callbacks:Collection[Callback]=None):
if wd is None: wd = self.wd
self.create_opt(lr, wd)
if callbacks is None: callbacks = []
callbacks += [cb(self) for cb in self.callback_fns]
pbar = master_bar(range(epochs))
self.recorder = Recorder(self.opt, epochs, self.data.train_dl, pbar)
callbacks = [self.recorder] + self.callbacks + callbacks
fit(epochs, self.model, self.loss_fn, self.opt, self.data, callbacks=callbacks, metrics=self.metrics, pbar=pbar)
def create_opt(self, lr:Floats, wd:Floats=0.):
lrs = listify(lr, self.layer_groups)
opt = self.opt_fn([{'params': trainable_params(l), 'lr':lr} for l,lr in zip(self.layer_groups, lrs)])
self.opt = OptimWrapper(opt, wd=wd, true_wd=self.true_wd)
def split(self, split_on):
if isinstance(split_on,Callable): split_on = split_on(self.model)
self.layer_groups = split_model(self.model, split_on)
def freeze_to(self, n):
for g in self.layer_groups[:n]:
for l in g:
if not self.train_bn or not isinstance(l, bn_types):
for p in l.parameters(): p.requires_grad = False
for g in self.layer_groups[n:]:
for p in g.parameters(): p.requires_grad = True
def freeze(self):
assert(len(self.layer_groups)>1)
self.freeze_to(-1)
def unfreeze(self): self.freeze_to(0)
def save(self, name): torch.save(self.model.state_dict(), self.path/f'{name}.pth')
def load(self, name): self.model.load_state_dict(torch.load(self.path/f'{name}.pth'))
import nb_004a
nb_004a.Learner = Learner
# +
#export
@dataclass
class Recorder(Callback):
opt: torch.optim
nb_epoch:int
train_dl: DeviceDataLoader = None
pbar: master_bar = None
def on_train_begin(self, **kwargs):
self.losses,self.val_losses,self.lrs,self.moms,self.metrics,self.nb_batches = [],[],[],[],[],[]
def on_batch_begin(self, **kwargs):
self.lrs.append(self.opt.lr)
self.moms.append(self.opt.mom)
def on_backward_begin(self, smooth_loss, **kwargs):
#We record the loss here before any other callback has a chance to modify it.
self.losses.append(smooth_loss)
if self.pbar is not None and hasattr(self.pbar,'child'):
self.pbar.child.comment = f'{smooth_loss:.4f}'
def on_epoch_end(self, epoch, num_batch, smooth_loss, last_metrics, **kwargs):
self.nb_batches.append(num_batch)
if last_metrics is not None:
self.val_losses.append(last_metrics[0])
if len(last_metrics) > 1: self.metrics.append(last_metrics[1:])
self.pbar.write(f'{epoch}, {smooth_loss}, {last_metrics}')
else: self.pbar.write(f'{epoch}, {smooth_loss}')
def plot_lr(self, show_moms=False):
iterations = list(range(len(self.lrs)))
if show_moms:
_, axs = plt.subplots(1,2, figsize=(12,4))
axs[0].plot(iterations, self.lrs)
axs[1].plot(iterations, self.moms)
else: plt.plot(iterations, self.lrs)
def plot(self, skip_start=10, skip_end=5):
lrs = self.lrs[skip_start:-skip_end] if skip_end > 0 else self.lrs[skip_start:]
losses = self.losses[skip_start:-skip_end] if skip_end > 0 else self.losses[skip_start:]
_, ax = plt.subplots(1,1)
ax.plot(lrs, losses)
ax.set_xscale('log')
def plot_losses(self):
_, ax = plt.subplots(1,1)
iterations = list(range(len(self.losses)))
ax.plot(iterations, self.losses)
val_iter = self.nb_batches
val_iter = np.cumsum(val_iter)
ax.plot(val_iter, self.val_losses)
def plot_metrics(self):
assert len(self.metrics) != 0, "There is no metrics to plot."
_, axes = plt.subplots(len(self.metrics[0]),1,figsize=(6, 4*len(self.metrics[0])))
val_iter = self.nb_batches
val_iter = np.cumsum(val_iter)
axes = axes.flatten() if len(self.metrics[0]) != 1 else [axes]
for i, ax in enumerate(axes):
values = [met[i] for met in self.metrics]
ax.plot(val_iter, values)
import nb_004
nb_004.Recorder = Recorder
# -
#export
@dataclass
class ShowGraph(Callback):
learn:Learner
def on_epoch_end(self, last_metrics, **kwargs):
if last_metrics is not None:
rec = learn.recorder
iters = list(range(len(rec.losses)))
val_iter = np.array(rec.nb_batches).cumsum()
x_bounds = (0, (rec.nb_epoch - len(rec.nb_batches)) * rec.nb_batches[-1] + len(rec.losses))
y_bounds = (0, max((max(rec.losses), max(rec.val_losses))))
rec.pbar.update_graph([(iters, rec.losses), (val_iter, rec.val_losses)], x_bounds, y_bounds)
# +
DATA_PATH = Path('data')
PATH = DATA_PATH/'cifar10'
data_mean,data_std = map(tensor, ([0.491, 0.482, 0.447], [0.247, 0.243, 0.261]))
cifar_norm,cifar_denorm = normalize_funcs(data_mean,data_std)
train_tfms = [flip_lr(p=0.5),
pad(padding=4),
crop(size=32, row_pct=(0,1.), col_pct=(0,1.))]
valid_tfms = []
bs = 64
# -
train_ds = ImageDataset.from_folder(PATH/'train', classes=['airplane','dog'])
valid_ds = ImageDataset.from_folder(PATH/'test', classes=['airplane','dog'])
data = DataBunch.create(train_ds, valid_ds, bs=bs, train_tfm=train_tfms, valid_tfm=valid_tfms, num_workers=0, dl_tfms=cifar_norm)
len(data.train_dl), len(data.valid_dl)
model = Darknet([1, 2, 2, 2, 2], num_classes=2, nf=16)
learn = Learner(data, model)
import pdb
learn.fit(5,0.01)
learn.callback_fns = [ShowGraph]
learn.fit(5,0.01)
| deep-learning/fastai-docs/fastai_docs-master/dev_nb/x_004b_implement_fast_progress.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # <center> Data Structure Basic
# ## Jupyter notebook
# 
# +
# kernels
# +
# hotkey
# +
# comment
# +
#markdown
# -
# **Hi, This is markdown cell**
# $$ \alpha = \beta = \frac{\alpha}{\sqrt{\beta}} $$
# # <center> Basic
import this
# ## Arithmetic Operators
# | Symbol | Task Performed |
# |:----:|:---:|
# | + | Addition |
# | - | Subtraction |
# | / | division |
# | % | mod |
# | * | multiplication |
# | // | floor division |
# | ** | to the power of |
2 + 4
6 - 4
5 * 3
5 / 3
5 % 3
3 ** 3
5 + (4 - 3 * 2)** 3 + 1
a = 5 + (4 - 3 * 2)** 3 + 1
a
print(a)
a = 110
a
a = 95
a
x = 2
y = 10
x
print(x+y)
print(x-y)
print(x//y)
# ## Relational Operators
#
# | Symbol | Task Performed |
# |:----: |:---:|
# |= |Assignment|
# |== |True, if it is equal|
# |!= |True, if not equal to|
# |< |less than|
# |> |greater than|
# |<= |less than or equal to|
# |>= |greater than or equal to|
x = 2
y = 6
print(x==y)
print(x==2)
print(x!=y)
print(x>y)
print(y>x)
print(x>=2)
print(x>2)
x > 1 and y < 10
x > 1 or y < 1
1 < x < 3
# ## Type
# | Types | Example |
# | :--: | :--: |
# | string | "Hello" , 'World' |
# | integer | 1 , 2 , 3 |
# | float | 1.2 , 4.6 , 112.6 |
# | boolian | True , Flase |
type
type("Economics")
type(1)
type(1.5)
type(False)
x = 1.3
type(x)
type(1.9)
int(1.9)
y = 5
float(y)
s1 = "Hi, this is python course"
s2 = "It's good to see you here!"
print(s1)
print(s2)
s1.count('p')
s1.find('t')
'-'.join(['1', '2', '3', '4'])
s1
s1.upper()
s1.lower()
s1.replace('i', "0")
s1.split(',')
v = 'aaaa'
print(v)
v * 2
z = 'bbbb'
v + z
# ## List
x = [1,2,3,4,5,6,7,8,9,10]
print(x)
type(x)
x[2]
x[-2]
x[3:6]
# +
# [<begin>:<end>:<step>]
x[0: 5: 3]
# -
x
x*2
x + [4]
y = [1 , 2 , 3]
y.append(5)
y
y.clear()
y
x = [1,2,3,4,5,6]
x
x.remove(1) # number
x
x.pop(2) # index
x
x.reverse()
x
# ## Dictionary
scores = { 'math':19 , 'lit': 18.5 , 'gio' : 20}
type(scores)
scores
scores['math']
scores['lit']
scores.keys()
scores.values()
# #### Other type
tuple
set
a = 3
a = a + 2
a
a += 2
a
import matplotlib.pyplot as plt
x = [1,2,3,4,5,6,7,8,9,10]
y = [10,15,25,85,45,63,87,112,36, 46]
plt.plot(x,y, 'r', lw= 2);
plt.title('Exaple')
plt.grid()
| MSc Planning Economics/1. Data Structure Basic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="djUvWu41mtXa"
# ##### Copyright 2019 The TensorFlow Authors.
# + cellView="form" colab={} colab_type="code" id="su2RaORHpReL"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="NztQK2uFpXT-"
# # Displaying image data in TensorBoard
#
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tensorboard/image_summaries"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/tensorboard/blob/master/docs/image_summaries.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/tensorboard/blob/master/docs/image_summaries.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="eDXRFe_qp5C3"
# ## Overview
#
# Using the **TensorFlow Image Summary API,** you can easily log tensors and arbitrary images and view them in TensorBoard. This can be extremely helpful to sample and examine your input data, or to [visualize layer weights](http://cs231n.github.io/understanding-cnn/) and [generated tensors](https://hub.packtpub.com/generative-adversarial-networks-using-keras/). You can also log diagnostic data as images that can be helpful in the course of your model development.
#
# In this tutorial, you will use learn how to use the Image Summary API to visualize tensors as images. You will also learn how to take an arbitrary image, convert it to a tensor, and visualize it in TensorBoard. You will work through a simple but real example that uses Image Summaries to help you understand how your model is performing.
#
# + [markdown] colab_type="text" id="dG-nnZK9qW9z"
# ## Setup
# + colab={"base_uri": "https://localhost:8080/", "height": 102} colab_type="code" id="3U5gdCw_nSG3" outputId="4a37496e-b653-431d-fb70-f916ae227d99"
try:
# # %tensorflow_version only exists in Colab.
# %tensorflow_version 2.x
except Exception:
pass
# Load the TensorBoard notebook extension.
# %load_ext tensorboard
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="1qIKtOBrqc9Y" outputId="f57988e5-8333-45ba-ef74-1e48995ee1c8"
from datetime import datetime
import io
import itertools
from packaging import version
from six.moves import range
import tensorflow as tf
from tensorflow import keras
import matplotlib.pyplot as plt
import numpy as np
import sklearn.metrics
print("TensorFlow version: ", tf.__version__)
assert version.parse(tf.__version__).release[0] >= 2, \
"This notebook requires TensorFlow 2.0 or above."
# + [markdown] colab_type="text" id="Tq0gyXOGZ3-h"
# # Download the Fashion-MNIST dataset
#
# You're going to construct a simple neural network to classify images in the the [Fashion-MNIST](https://research.zalando.com/welcome/mission/research-projects/fashion-mnist/) dataset. This dataset consist of 70,000 28x28 grayscale images of fashion products from 10 categories, with 7,000 images per category.
#
# First, download the data:
# + colab={"base_uri": "https://localhost:8080/", "height": 153} colab_type="code" id="VmEQwCon3i7m" outputId="41db92f3-905f-4121-d906-a7baa1629dbb"
# Download the data. The data is already divided into train and test.
# The labels are integers representing classes.
fashion_mnist = keras.datasets.fashion_mnist
(train_images, train_labels), (test_images, test_labels) = \
fashion_mnist.load_data()
# Names of the integer classes, i.e., 0 -> T-short/top, 1 -> Trouser, etc.
class_names = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
# + [markdown] colab_type="text" id="qNsjMY0364j4"
# ## Visualizing a single image
#
# To understand how the Image Summary API works, you're now going to simply log the first training image in your training set in TensorBoard.
#
# Before you do that, examine the shape of your training data:
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="FxMPcdmvBn9t" outputId="0de21cb0-3649-4a3b-a293-1b0e4f38c57e"
print("Shape: ", train_images[0].shape)
print("Label: ", train_labels[0], "->", class_names[train_labels[0]])
# + [markdown] colab_type="text" id="4F8zbUKfBuUt"
# Notice that the shape of each image in the data set is a rank-2 tensor of shape (28, 28), representing the height and the width.
#
# However, ```tf.summary.image()``` expects a rank-4 tensor containing ```(batch_size, height, width, channels)```. Therefore, the tensors need to be reshaped.
#
# You're logging only one image, so ```batch_size``` is 1. The images are grayscale, so set ```channels``` to 1.
# + colab={} colab_type="code" id="5yPh-7EWB8IK"
# Reshape the image for the Summary API.
img = np.reshape(train_images[0], (-1, 28, 28, 1))
# + [markdown] colab_type="text" id="JAdJDY3FCCwt"
# You're now ready to log this image and view it in TensorBoard.
# + colab={} colab_type="code" id="IJNpyVyxbVtT"
# Clear out any prior log data.
# !rm -rf logs
# Sets up a timestamped log directory.
logdir = "logs/train_data/" + datetime.now().strftime("%Y%m%d-%H%M%S")
# Creates a file writer for the log directory.
file_writer = tf.summary.create_file_writer(logdir)
# Using the file writer, log the reshaped image.
with file_writer.as_default():
tf.summary.image("Training data", img, step=0)
# + [markdown] colab_type="text" id="rngALbRogXe6"
# Now, use TensorBoard to examine the image. Wait a few seconds for the UI to spin up.
# + colab={} colab_type="code" id="T_X-wIy-lD9f"
# %tensorboard --logdir logs/train_data
# + [markdown] colab_type="text" id="c8n8YqGlT3-c"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/images_single.png?raw=1"/>
# + [markdown] colab_type="text" id="34enxJjjgWi7"
# The "Images" tab displays the image you just logged. It's an "ankle boot".
#
# The image is scaled to a default size for easier viewing. If you want to view the unscaled original image, check "Show actual image size" at the upper left.
#
# Play with the brightness and contrast sliders to see how they affect the image pixels.
# + [markdown] colab_type="text" id="bjACE1lAsqUd"
# ## Visualizing multiple images
#
# Logging one tensor is great, but what if you wanted to log multiple training examples?
#
# Simply specify the number of images you want to log when passing data to ```tf.summary.image()```.
# + colab={} colab_type="code" id="iHUjCXbetIpb"
with file_writer.as_default():
# Don't forget to reshape.
images = np.reshape(train_images[0:25], (-1, 28, 28, 1))
tf.summary.image("25 training data examples", images, max_outputs=25, step=0)
# %tensorboard --logdir logs/train_data
# + [markdown] colab_type="text" id="Fr6LFQG9UD6z"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/images_multiple.png?raw=1"/>
# + [markdown] colab_type="text" id="c-7sZs3XuBBy"
# ## Logging arbitrary image data
#
# What if you want to visualize an image that's not a tensor, such as an image generated by [matplotlib](https://matplotlib.org/)?
#
# You need some boilerplate code to convert the plot to a tensor, but after that, you're good to go.
#
# In the code below, you'll log the first 25 images as a nice grid using matplotlib's ```subplot()``` function. You'll then view the grid in TensorBoard:
# + colab={} colab_type="code" id="F5U_5WKt8bdQ"
# Clear out prior logging data.
# !rm -rf logs/plots
logdir = "logs/plots/" + datetime.now().strftime("%Y%m%d-%H%M%S")
file_writer = tf.summary.create_file_writer(logdir)
def plot_to_image(figure):
"""Converts the matplotlib plot specified by 'figure' to a PNG image and
returns it. The supplied figure is closed and inaccessible after this call."""
# Save the plot to a PNG in memory.
buf = io.BytesIO()
plt.savefig(buf, format='png')
# Closing the figure prevents it from being displayed directly inside
# the notebook.
plt.close(figure)
buf.seek(0)
# Convert PNG buffer to TF image
image = tf.image.decode_png(buf.getvalue(), channels=4)
# Add the batch dimension
image = tf.expand_dims(image, 0)
return image
def image_grid():
"""Return a 5x5 grid of the MNIST images as a matplotlib figure."""
# Create a figure to contain the plot.
figure = plt.figure(figsize=(10,10))
for i in range(25):
# Start next subplot.
plt.subplot(5, 5, i + 1, title=class_names[train_labels[i]])
plt.xticks([])
plt.yticks([])
plt.grid(False)
plt.imshow(train_images[i], cmap=plt.cm.binary)
return figure
# Prepare the plot
figure = image_grid()
# Convert to image and log
with file_writer.as_default():
tf.summary.image("Training data", plot_to_image(figure), step=0)
# %tensorboard --logdir logs/plots
# + [markdown] colab_type="text" id="o_tIghRsXY7S"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/images_arbitrary.png?raw=1"/>
# + [markdown] colab_type="text" id="vZx70BC1zhgW"
# ## Building an image classifier
#
# Now put this all together with a real example. After all, you're here to do machine learning and not plot pretty pictures!
#
# You're going to use image summaries to understand how well your model is doing while training a simple classifier for the Fashion-MNIST dataset.
#
# First, create a very simple model and compile it, setting up the optimizer and loss function. The compile step also specifies that you want to log the accuracy of the classifier along the way.
# + colab={} colab_type="code" id="R74hPWJHzgvZ"
model = keras.models.Sequential([
keras.layers.Flatten(input_shape=(28, 28)),
keras.layers.Dense(32, activation='relu'),
keras.layers.Dense(10, activation='softmax')
])
model.compile(
optimizer='adam',
loss='sparse_categorical_crossentropy',
metrics=['accuracy']
)
# + [markdown] colab_type="text" id="SdT_PpZB1UMn"
# When training a classifier, it's useful to see the [confusion matrix](https://en.wikipedia.org/wiki/Confusion_matrix). The confusion matrix gives you detailed knowledge of how your classifier is performing on test data.
#
# Define a function that calculates the confusion matrix. You'll use a convenient [Scikit-learn](https://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html) function to do this, and then plot it using matplotlib.
# + colab={} colab_type="code" id="rBiXP8-UO8t6"
def plot_confusion_matrix(cm, class_names):
"""
Returns a matplotlib figure containing the plotted confusion matrix.
Args:
cm (array, shape = [n, n]): a confusion matrix of integer classes
class_names (array, shape = [n]): String names of the integer classes
"""
figure = plt.figure(figsize=(8, 8))
plt.imshow(cm, interpolation='nearest', cmap=plt.cm.Blues)
plt.title("Confusion matrix")
plt.colorbar()
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names, rotation=45)
plt.yticks(tick_marks, class_names)
# Normalize the confusion matrix.
cm = np.around(cm.astype('float') / cm.sum(axis=1)[:, np.newaxis], decimals=2)
# Use white text if squares are dark; otherwise black.
threshold = cm.max() / 2.
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
color = "white" if cm[i, j] > threshold else "black"
plt.text(j, i, cm[i, j], horizontalalignment="center", color=color)
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
return figure
# + [markdown] colab_type="text" id="6lOAl_v26QGq"
# You're now ready to train the classifier and regularly log the confusion matrix along the way.
#
# Here's what you'll do:
#
# 1. Create the [Keras TensorBoard callback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/TensorBoard) to log basic metrics
# 2. Create a [Keras LambdaCallback](https://www.tensorflow.org/api_docs/python/tf/keras/callbacks/LambdaCallback) to log the confusion matrix at the end of every epoch
# 3. Train the model using Model.fit(), making sure to pass both callbacks
#
# As training progresses, scroll down to see TensorBoard start up.
# + colab={} colab_type="code" id="utd-vH6hn5RY"
# Clear out prior logging data.
# !rm -rf logs/image
logdir = "logs/image/" + datetime.now().strftime("%Y%m%d-%H%M%S")
# Define the basic TensorBoard callback.
tensorboard_callback = keras.callbacks.TensorBoard(log_dir=logdir)
file_writer_cm = tf.summary.create_file_writer(logdir + '/cm')
# + colab={} colab_type="code" id="bXQ7-9CF0TPA"
def log_confusion_matrix(epoch, logs):
# Use the model to predict the values from the validation dataset.
test_pred_raw = model.predict(test_images)
test_pred = np.argmax(test_pred_raw, axis=1)
# Calculate the confusion matrix.
cm = sklearn.metrics.confusion_matrix(test_labels, test_pred)
# Log the confusion matrix as an image summary.
figure = plot_confusion_matrix(cm, class_names=class_names)
cm_image = plot_to_image(figure)
# Log the confusion matrix as an image summary.
with file_writer_cm.as_default():
tf.summary.image("Confusion Matrix", cm_image, step=epoch)
# Define the per-epoch callback.
cm_callback = keras.callbacks.LambdaCallback(on_epoch_end=log_confusion_matrix)
# + colab={} colab_type="code" id="k6CV7dy-oJZu"
# Start TensorBoard.
# %tensorboard --logdir logs/image
# Train the classifier.
model.fit(
train_images,
train_labels,
epochs=5,
verbose=0, # Suppress chatty output
callbacks=[tensorboard_callback, cm_callback],
validation_data=(test_images, test_labels),
)
# + [markdown] colab_type="text" id="o7PnxGf8Ur6F"
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/images_accuracy.png?raw=1"/>
#
# <img class="tfo-display-only-on-site" src="https://github.com/tensorflow/tensorboard/blob/master/docs/images/images_cm.png?raw=1"/>
# + [markdown] colab_type="text" id="6URWgszz9Jut"
# Notice that accuracy is climbing on both train and validation sets. That's a good sign. But how is the model performing on specific subsets of the data?
#
# Select the "Images" tab to visualize your logged confusion matrices.
# Check "Show actual image size" at the top left to see the confusion matrix at full size.
#
# By default the dashboard shows the image summary for the last logged step or epoch. Use the slider to view earlier confusion matrices. Notice how the matrix changes significantly as training progresses, with darker squares coalescing along the diagonal, and the rest of the matrix tending toward 0 and white. This means that your classifier is improving as training progresses! Great work!
#
# The confusion matrix shows that this simple model has some problems. Despite the great progress, Shirts, T-Shirts, and Pullovers are getting confused with each other. The model needs more work.
#
# If you're interested, try to improve this model with a [convolutional network](https://medium.com/tensorflow/hello-deep-learning-fashion-mnist-with-keras-50fcff8cd74a) (CNN).
| site/en-snapshot/tensorboard/image_summaries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework #2
# # ARE 106 Summer Session II
# ## Name:
# ## SSID:
# Please put your name and SSID in the corresponding cells above.
#
# The homework is worth 13.5 points.
#
# For each of the following questions, show as much of your steps as you can (without going overboard). If you end up getting the wrong answer, but we spot where you made a mistake in the algebra, partial credit will be more readily given. If you only put the final answer, you will be marked either right or wrong.
#
# Answer questions in the correct cell. For problems where you have to input math, make sure that you know that it's a markdown cell (It won't have a `In: []` on the left) and make sure you run the cell by either pressing `Ctrl + Enter` or going to `Cell -> Run Cell`. Alternatively, write all your answers and then go to `Cell -> Run All Cells` after you're done.
#
# **When you are finished export your homework to a PDF by going to `File -> Download as -> PDF`.**
# ### Exercise 1: Statistics Review
#
# For the following problems, write down the answer in the markdown cell below the question. If you have trouble writing math, either double-click on the markdown cell to see how it was written or consult [this cheatsheet](https://math.meta.stackexchange.com/questions/5020/mathjax-basic-tutorial-and-quick-reference
# ):
#
#
# Assume that there are two random variables $X$ and $Y$ with expected values $\mu_x$ and $\mu_y$, and variances $\sigma_x$ and $\sigma_y$. Let the covariance between $X$ and $Y$ be: $\sigma_{xy}$. Note that the covariance between these two variables need not be 0 (i.e. they aren't necessarily independent).
#
# Any others ($a,b,c$ etc...) can be considered non-random parameters.
#
# For each of the following problems, rewrite the expressions in terms of $\mu_x,\mu_y,\sigma_x,\sigma_y$ and $\sigma_{x,y}$ when possible. If not possible, write what you think the answer is and explain why.
#
# - i. $E(aX + bY)$
# - ii. $Var(bY + c)$
# - iii. $Cov(aX + b, c^{2}Y+d)$
# - iv. $E(b)$, remember $b$ is non-random
# - v. $Var(a)$, remember $a$ is non-random
# **Please write your answers here. If you need to use more than one line, you may do so.**
#
# - i. $E(aX) + E(bY) = a\mu_x + b\mu_y$
# - ii. $b^2 \sigma_y$
# - iii. $ac^2\sigma_{xy}$
# - iv. b
# - v. 0
# ### Exercise 2: Statistics Review Continued
#
# This exercise uses the same instructions as question 1.
# - i. What is the definition of $Cov(X,Y)$ in terms of expectations? **Hint: check the statistics review slides from the first class**.
# - ii. Rewrite this expression so that it has $E(XY)$ on one side. Does $E(XY) = \mu_x \mu_y$? Why or why not? **Hint: $\mu_x$ and $\mu_y$ are not random!**
# **Please write your answers here. If you need to use more than one line, you may do so.**
#
# - i. $Cov(X,Y) = E\left[(X-\mu_x)(Y-\mu_y)\right]$
# - ii. $Cov(X,Y) = E(XY) - \mu_x \mu_y - \mu_x \mu_y + \mu_x \mu_y = E(XY) - \mu_x \mu_y$
#
# Therefore: $E(XY) = Cov(X,Y) + \mu_x \mu_y$
#
# In order for this to be true, $Cov(X,Y)=0$.
#
# ### Exercise 3: Deriving the $R^2$.
#
# Suppose that we have an estimated regression model $y_i = \hat{b}_0 + \hat{b}_1 x_i +e_i$, where $\hat{b}_0,\hat{b}_1$ are estimated OLS coefficients. Let $\hat{y}_i = \hat{b}_0 + \hat{b}_1 x_i$, so that:
#
# $$
# y_i = \hat{y}_i + e_i
# $$
#
# Use the definition for $Var(y_i)$ to derive the expression for the $R^2$. What expression has to be equal to 0 in order for this to be true?
# **Please write your answer here. If you need to use more than one line, you may do so.**
#
# $Var(y_i) = Var(\hat{y_i} + e_i) = Var(\hat{y_i})+Var(e_i) + 2Cov(\hat{y_i}, e_i)$
#
# But we know that $Cov(\hat{y_i}, e_i)$ must be equal to 0, so:
#
# $\frac{1}{N}\sum_i^N (y_i - \mu_y)^2 = \frac{1}{N}\sum_{i}^N (\hat{y_i} - \bar{\hat{y_i}})^2 + \frac{1}{N}\sum_i^N (e_i - \bar{e})^2$
#
# We can see that this is the same as $TSS = ESS + SSR$. Since $R^2 = \frac{ESS}{TSS}$, if we divide the above expression by $TSS$, we get:
#
# $1 = R^2 + \frac{SSR}{TSS}$
#
# So $R^2 = 1- \frac{SSR}{TSS}$
# ### Exercise 4: Ordinary Least Squares
#
# Suppose you have some data $x_i$ and you would like to see how it affects $y_i$, your dependent variable. You think that this relationship can be estimated linearly, so the model you write down is:
#
# $$
# y_i = b_0 + b_1 x_i + e_i
# $$
#
# - i. If you wanted to estimate this relationship using OLS, what would be the expressions for $\hat{b}_0$ and $\hat{b}_1$? *You do not need to show your work for this question, you can just write it.*
# - ii. Now suppose that the independent variable, $x_i$, is made up of really small number (ex. many of the observations are .000001). You decide to multiply this variable by $1,000$ so that the variable doesn't have quite so many 0's after the decimal point (making it easier to understand). Let's see how that would change the OLS coefficients.
# - a. What would your estimating equation look like now? (Hint:It will be similar to equation (1), but with one change)
# - b. Write the minimization problem that you need to solve.
# - c. Solve the minimization problem and give the answers for the expressions for the resulting, new OLS coefficients.
# - d. Which coefficient is bigger or smaller?
# **Please write your answer here. If you need to use more than one line, you may do so.**
#
# - i. $b_0 = \bar{y} - b_0 - b_1 \bar{x}$
#
# $b_1$ = $\frac{\sum_i^N (X- \bar{X})(Y-\bar{Y})}{\sum_i^N (X-\bar{X})^2}$
# - ii.
# - a. $y_i = b_0 + b_1 1000 x_i + e_i$
# - b. $\min_{b_0,b_1} \sum_i^N (y_i - b_0 - b_1 1000 x_i)^2$
# - c.
# The first order conditions are:
# $$
# (b_0): 2\sum_i^N (y_i - b_0 - b_1 1000 x_i)(-1)=0
# $$
# $$
# (b_1): 2\sum_i^N (y_i - b_0 - b_1 1000 x_i)(-1000 x_i)=0
# $$
#
# Solving for $b_0$, we get:
#
# $\sum_i^N y_i - \sum_i^N b_0 - \sum_i^N b_1 1000 x_i =0$
#
# $N\bar{y} - Nb_0 - b_1 1000 N\bar{x} =0$
#
# So $\hat{b_0} = \bar{y} - b_1 1000 \bar{x}$
#
# Now substituting that into the $b_1$ first order condition:
# $\sum_i^N (y_i - (\bar{y} - b_1 1000 \bar{x}) - b_1 1000 x_i)(x_i)=0$
#
# $\sum_i^N (y_i - \bar{y})x_i - 1000 b_1 \sum_i^N (x_i -\bar{x})x_i=0$
#
# So we get:
#
# $\hat{b_1} = \frac{1}{1000} \frac{\sum_i^N (y_i - \bar{y})x_i}{\sum_i^N (x_i -\bar{x})x_i}$
#
# - d. From this we can see that the new coefficient is now smaller than before.
# ### Exercise 5: Python Coding
#
# Suppose we have a list of numbers `my_list = [1,3,5,6,7]`. Write a program that takes this list, and multiplies it by 2, but if the number is 3, print out, "this is the number 3."
#
# So what we're looking for, is a function that takes in a list and outputs a list.
#
# Input: `[1,3,5,6,7]`
#
# Output:
#
# ```
# [2,6,10,12,14]
# "this is the number 3"
# ```
#
# **Remember to comment your code and to initialize your lists first (you might need to initialize an empty list first). A comment can be made by using the `#` symbol. Also remember that to tell the function what you want to output, you need to include a `return` statement in the function.**
# +
### Please put your answer here:
## Initialize list
my_list = [1,3,5,6,7]
## define function that multiplies each part of the list by 2 and prints something out for 3.
def f(x):
## initialize empty list for collection of returned values
list_collector = []
for thing in x:
list_collector.append(2*thing)
if thing==3:
print("this is the number 3")
return list_collector
f(my_list)
| Homework/Homework 2/HW2_answers.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test EfficientDet model and BiFPNs
from efficientnet_pytorch import EfficientNet
import torch.nn as nn
import torch
import torch.nn.functional as F
from utils.ssd_model import DBox, Detect
def make_loc_conf(num_classes=21, bbox_aspect_num=[4, 6, 6, 6, 4, 4]):
loc_layers = []
conf_layers = []
for i in range(6):
loc_layers += [nn.Conv2d(256, 256, kernel_size=3, padding=1), nn.BatchNorm2d(256, eps=1e-4, momentum=0.997), nn.ReLU(inplace=True),
nn.Conv2d(256, bbox_aspect_num[i]* 4, kernel_size=3, padding=1), nn.BatchNorm2d(bbox_aspect_num[i]* 4, eps=1e-4, momentum=0.997)]
conf_layers += [nn.Conv2d(256, 256, kernel_size=3, padding=1), nn.BatchNorm2d(256, eps=1e-4, momentum=0.997), nn.ReLU(inplace=True),
nn.Conv2d(256, bbox_aspect_num[i]* num_classes, kernel_size=3, padding=1), nn.BatchNorm2d(bbox_aspect_num[i]* num_classes, eps=1e-4, momentum=0.997)]
return nn.ModuleList(loc_layers), nn.ModuleList(conf_layers)
class BiFPN(nn.Module):
def __init__(self,
num_channels):
super(BiFPN, self).__init__()
self.num_channels = num_channels
def forward(self, inputs):
num_channels = self.num_channels
P3_in, P4_in, P5_in, P6_in, P7_in = inputs
for input in inputs:
print(input.size())
P7_up = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0, groups=num_channels)(P7_in)
scale = (P6_in.size(3)/P7_up.size(3))
P6_up = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0, groups=num_channels)(P6_in+self.Resize(scale_factor=scale)(P7_up))
scale = (P5_in.size(3)/P6_up.size(3))
P5_up = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0, groups=num_channels)(P5_in+self.Resize(scale_factor=scale)(P6_up))
scale = (P4_in.size(3)/P5_up.size(3))
P4_up = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0)(P4_in+self.Resize(scale_factor=scale)(P5_up))
scale = (P3_in.size(3)/P4_up.size(3))
P3_out = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0)(P3_in+self.Resize(scale_factor=scale)(P4_up))
# downsample by interpolation
#print("P6_up scale",scale)
P4_out = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0, groups=num_channels)(P4_in + P4_up+F.interpolate(P3_out, P4_up.size()[2:]))
P5_out = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0, groups=num_channels)(P5_in + P5_up+F.interpolate(P4_out, P5_up.size()[2:]))
P6_out = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0, groups=num_channels)(P6_in + P6_up+F.interpolate(P5_out, P6_up.size()[2:]))
P7_out = self.Conv(in_channels=num_channels, out_channels=num_channels, kernel_size=1, stride=1, padding=0, groups=num_channels)(P7_in + P7_up+F.interpolate(P6_out, P7_up.size()[2:]))
return P3_out, P4_out, P5_out, P6_out, P7_out
@staticmethod
def Conv(in_channels, out_channels, kernel_size, stride, padding, groups = 1):
features = nn.Sequential(
nn.Conv2d(in_channels=in_channels, out_channels=out_channels, kernel_size=kernel_size, stride=stride, padding=padding, groups=groups),
nn.BatchNorm2d(num_features=out_channels, eps=1e-4, momentum=0.997),
nn.ReLU(inplace=True)
)
return features
@staticmethod
def Resize(scale_factor=2, mode='bilinear'):
upsample = nn.Upsample(scale_factor=scale_factor, mode=mode)
return upsample
class EfficientDet(nn.Module):
def __init__(self, phase, cfg, verbose=False, backbone="efficientnet-b0", useBiFPN=True):
super(EfficientDet, self).__init__()
# meta-stuff
self.phase = phase
self.num_classes = cfg["num_classes"]
self.verbose=verbose
# make Dbox
dbox = DBox(cfg)
self.device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
self.dbox_list = dbox.make_dbox_list()
# use Detect if inference
if phase == "inference":
self.detect = Detect()
ratio = 1
# define backbone
model = EfficientNet.from_pretrained(backbone)
self.layer0 = nn.Sequential(model._conv_stem, model._bn0)
if backbone == "efficientnet-b0":
self.layer2 = nn.Sequential(model._blocks[0],model._blocks[1],model._blocks[2],model._blocks[3])
self.layer3 = nn.Sequential(model._blocks[4],model._blocks[5])
self.layer4 = nn.Sequential(model._blocks[6],model._blocks[7],model._blocks[8],model._blocks[9],model._blocks[10],model._blocks[11])
self.layer5 = nn.Sequential(model._blocks[12],model._blocks[13],model._blocks[14],model._blocks[15])
elif backbone == "efficientnet-b2":
self.layer2 = nn.Sequential(model._blocks[0],model._blocks[1],model._blocks[2],model._blocks[3],model._blocks[4],model._blocks[5])
self.layer3 = nn.Sequential(model._blocks[6],model._blocks[7],model._blocks[8])
self.layer4 = nn.Sequential(model._blocks[9],model._blocks[10],model._blocks[11])
self.layer5 = nn.Sequential(model._blocks[12],model._blocks[13],model._blocks[14],model._blocks[15],model._blocks[16],model._blocks[17],model._blocks[18])
# Bottom-up layers
#self.conv5 = nn.Conv2d( 320, 256, kernel_size=1, stride=1, padding=0)
self.conv6 = nn.Conv2d( self.layer5[-1]._block_args.input_filters, 256, kernel_size=3, stride=2, padding=1)
self.conv7 = nn.Conv2d( 256, 256, kernel_size=3, stride=2, padding=1)
self.conv8 = nn.Conv2d( 256, 256, kernel_size=3, stride=1, padding=0)
# Top layer
self.toplayer = nn.Conv2d(self.layer5[-1]._block_args.input_filters, 256, kernel_size=1, stride=1, padding=0) # Reduce channels
# Smooth layers
self.smooth1 = nn.Conv2d(256, 256, kernel_size=3, stride=1, padding=1)
self.smooth2 = nn.Conv2d(256, 256, kernel_size=3, stride=1, padding=1)
# Lateral layers
self.latlayer1 = nn.Conv2d( self.layer4[-1]._block_args.input_filters, 256, kernel_size=1, stride=1, padding=0)
self.latlayer2 = nn.Conv2d( self.layer3[-1]._block_args.input_filters, 256, kernel_size=1, stride=1, padding=0)
# loc, conf layers
self.loc, self.conf = make_loc_conf(self.num_classes, cfg["bbox_aspect_num"])
# FPNs
self.usebifpn=useBiFPN
if BiFPN:
self.BiFPN=BiFPN(256)
def forward(self, x):
# efficientnet layers
x = self.layer0(x)
p3 = self.layer2(x) # 37x37
p4 = self.layer3(p3) # 18x18
p5 = self.layer4(p4)
p5 = self.layer5(p5)
if self.verbose:
print("layerc3:", p3.size())
print("layerc4:", p4.size())
print("layerc5:", p5.size())
# non-efficientnet layers
p6 = self.conv6(p5) # 5x5
p7 = self.conv7(F.relu(p6)) # 3x3
p8 = self.conv8(F.relu(p7)) # 1x1
# TODO: implement BiFPN
if not self.usebifpn:
# Top-down
p5 = self.toplayer(p5) # 10x10
p4 = self._upsample_add(p5, self.latlayer1(p4)) # 19x19
p3 = self._upsample_add(p4, self.latlayer2(p3)) # 38x38
# Smooth
p4 = self.smooth1(p4)
p3 = self.smooth2(p3)
# make loc and confs.
sources = [p3, p4, p5, p6, p7, p8]
else:
# BiFPNs
# Top-down
p5 = self.toplayer(p5) # 10x10
p4 = self._upsample_add(p5, self.latlayer1(p4)) # 19x19
p3 = self._upsample_add(p4, self.latlayer2(p3)) # 38x38
sources = [p3, p4, p5, p6, p7]
sources = self.BiFPN(sources)
# look at source size
if self.verbose:
for source in sources:
print("layer size:", source.size())
# make lists
loc = list()
conf = list()
for (x, l, c) in zip(sources, self.loc, self.conf):
# Permuteは要素の順番を入れ替え
loc.append(l(x).permute(0, 2, 3, 1).contiguous())
conf.append(c(x).permute(0, 2, 3, 1).contiguous())
# locとconfの形を変形
# locのサイズは、torch.Size([batch_num, 34928])
# confのサイズはtorch.Size([batch_num, 183372])になる
loc = torch.cat([o.view(o.size(0), -1) for o in loc], 1)
conf = torch.cat([o.view(o.size(0), -1) for o in conf], 1)
# さらにlocとconfの形を整える
# locのサイズは、torch.Size([batch_num, 8732, 4])
# confのサイズは、torch.Size([batch_num, 8732, 21])
loc = loc.view(loc.size(0), -1, 4)
conf = conf.view(conf.size(0), -1, self.num_classes)
# これで後段の処理につっこめるかたちになる。
output = (loc, conf, self.dbox_list)
if self.phase == "inference":
# Detectのforward
return self.detect(output[0], output[1], output[2].to(self.device))
else:
return output
def _upsample_add(self, x, y):
'''Upsample and add two feature maps.
Args:
x: (Variable) top feature map to be upsampled.
y: (Variable) lateral feature map.
Returns:
(Variable) added feature map.
Note in PyTorch, when input size is odd, the upsampled feature map
with `F.upsample(..., scale_factor=2, mode='nearest')`
maybe not equal to the lateral feature map size.
e.g.
original input size: [N,_,15,15] ->
conv2d feature map size: [N,_,8,8] ->
upsampled feature map size: [N,_,16,16]
So we choose bilinear upsample which supports arbitrary output sizes.
'''
_,_,H,W = y.size()
return F.upsample(x, size=(H,W), mode='bilinear') + y
# +
from utils.efficientdet import EfficientDet
scale = 1
num_class = 21
if scale==1:
ssd_cfg = {
'num_classes': num_class, # 背景クラスを含めた合計クラス数
'input_size': 300*scale, # 画像の入力サイズ
'bbox_aspect_num': [4, 6, 6, 6, 4, 4], # 出力するDBoxのアスペクト比の種類
'feature_maps': [37, 18, 9, 5, 3, 1], # 各sourceの画像サイズ
'steps': [8, 16, 32, 64, 100, 300], # DBOXの大きさを決める
'min_sizes': [30, 60, 111, 162, 213, 264], # DBOXの大きさを決める
'max_sizes': [60, 111, 162, 213, 264, 315], # DBOXの大きさを決める
'aspect_ratios': [[2], [2, 3], [2, 3], [2, 3], [2], [2]],
}
elif scale==2:
ssd_cfg = {
'num_classes': num_class, # 背景クラスを含めた合計クラス数
'input_size': 512, # 画像の入力サイズ
'bbox_aspect_num': [4, 6, 6, 6, 4, 4], # 出力するDBoxのアスペクト比の種類
'feature_maps': [64, 32, 16, 8, 4, 2], # 各sourceの画像サイズ
'steps': [8, 16, 32, 64, 100, 300], # DBOXの大きさを決める
'min_sizes': [30, 60, 111, 162, 213, 264]*scale, # DBOXの大きさを決める
'max_sizes': [60, 111, 162, 213, 264, 315]*scale, # DBOXの大きさを決める
'aspect_ratios': [[2], [2, 3], [2, 3], [2, 3], [2], [2]],
}
back = EfficientDet("train", ssd_cfg, verbose=True, backbone="efficientnet-b0").eval()
x = torch.rand([1,3,300,300])
out = back(x)
print(out[0].size())
print(out[1].size())
# -
print(back)
for param in back.layer0.parameters():
param.requires_grad = False
from utils.efficientdet import EfficientDet
back = EfficientDet("train", ssd_cfg, verbose=True, backbone="efficientnet-b0")
x = torch.rand([1,3,512,512])
out = back(x)
print(out[0].size())
print(out[1].size())
# # Set up MS-coco dataset
# +
from dataset.coco import COCODetection
import torch.utils.data as data
from utils.dataset import VOCDataset, COCODatasetTransform, make_datapath_list, Anno_xml2list, od_collate_fn
batch_size = 2
# +
color_mean = (104, 117, 123) # (BGR)の色の平均値
input_size = 300 # 画像のinputサイズを300×300にする
## DatasetTransformを適応
transform = COCODatasetTransform(input_size, color_mean)
# -
dataset = COCODetection("../data/coco/", image_set="val2014", phase="train", transform=transform)
train_dataloader = data.DataLoader(
dataset, batch_size=batch_size, shuffle=True, num_workers=8, pin_memory=True, collate_fn=od_collate_fn)
# 動作の確認
batch_iterator = iter(train_dataloader) # イタレータに変換
images, targets = next(batch_iterator) # 1番目の要素を取り出す
print(images.size()) # torch.Size([4, 3, 300, 300])
print(len(targets))
print(targets[1].shape) # ミニバッチのサイズのリスト、各要素は[n, 5]、nは物体数
import numpy as np
a = np.random.randn(100,100,3)
np.mean(a,axis=(0,1))
images = images.numpy()[0,:,:,:].transpose([1,2,0])
import matplotlib.pyplot as plt
import cv2
plt.imshow(images) # pltはRGBで表示する.
plt.show()
# + active=""
#
| effnet_testing.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python38
# language: python
# name: python38
# ---
# +
#default_exp secrets
# -
# # Secrets
#export
import boto3
import ujson as json
#export
def getSecret(name="removeBg", region='ap-southeast-1', **kwargs):
'''
retrieve secret information from Amazon's secret manager without revealing the secret
'''
session = boto3.session.Session()
client = session.client(
service_name='secretsmanager',
region_name=region,
**kwargs
)
get_secret_value_response = client.get_secret_value(
SecretId=name
)
return json.loads(get_secret_value_response['SecretString'])
# +
# getSecret()
# -
| nbs/secrets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import tensorflow as tf
# +
coefficients = np.array([[1.], [-20], [100.]])
w = tf.Variable(0,dtype=tf.float32)
x = tf.placeholder(tf.float32,[3, 1])
# cost = tf.add(tf.add(w**2,tf.multiply(-10.,w)),25)
# cost = w**2 -10*w + 25
cost = x[0][0]*w**2 + x[1][0]*w + x[2][0]
train = tf.train.GradientDescentOptimizer(0.01).minimize(cost)
init = tf.global_variables_initializer()
session = tf.Session()
session.run(init)
session.run(w)
print(session.run(w))
# -
session.run(train,feed_dict={x:coefficients})
print(session.run(w))
for i in range(1000):
session.run(train, feed_dict={x:coefficients})
print(session.run(w))
| python/coursera_python/deeplearning_ai_Andrew_Ng/2_IDNN_HTRO/tensorflow_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from plotly.subplots import make_subplots
import plotly.graph_objects as go
import plotly
from datetime import datetime
PATH = "../../"
df_mortalite = pd.read_csv(PATH+'data/france/deces_quotidiens_departement_csv.csv', sep=";", encoding="'windows-1252'")
df_mortalite_2018 = pd.read_csv(PATH+'data/france/deces_quotidiens_departement_csv_avec_2018.csv', sep=";", encoding="'windows-1252'")
# +
#df_mortalite = df_mortalite.merge(df_mortalite_2018[["Date_evenement", "Total_deces_2018"]], left_on="Date_evenement", right_on="Date_evenement", how="left")
# +
df_mortalite_france = df_mortalite[df_mortalite["Zone"] == "France"]
df_mortalite_france_2018 = df_mortalite_2018[df_mortalite_2018["Zone"] == "France"]
window = 7
#df_mortalite_france.loc[:,"Total_deces_2018_diff"] = df_mortalite_france["Total_deces_2018"].diff().rolling(window=window, center=True).mean()
df_mortalite_france_2018.loc[:,"Total_deces_2018_diff"] = df_mortalite_france_2018["Total_deces_2018"].diff().rolling(window=window, center=True).mean()
df_mortalite_france.loc[:,"Total_deces_2019_diff"] = df_mortalite_france["Total_deces_2019"].diff().rolling(window=window, center=True).mean()
df_mortalite_france.loc[:,"Total_deces_2020_diff"] = df_mortalite_france["Total_deces_2020"].diff().rolling(window=window, center=True).mean()
df_mortalite_france.loc[:,"Total_deces_2021_diff"] = df_mortalite_france["Total_deces_2021"].diff().rolling(window=window, center=True).mean()
# -
df_mortalite_france_2018
# +
#### Construction du graphique
fig = make_subplots(specs=[[{"secondary_y": False}]])
# Ajout R_effectif estimé via les urgences au graph
"""fig.add_trace(go.Scatter(x = df_mortalite_france["Date_evenement"], y = df_mortalite_france["Total_deces_2018_diff"],
mode='lines',
line=dict(width=4, color="rgb(96, 178, 219)"),
name="Décès 2018",
marker_size=4,
showlegend=True
))"""
fig.add_trace(go.Scatter(x = df_mortalite_france["Date_evenement"], y = df_mortalite_france["Total_deces_2019_diff"],
mode='lines',
line=dict(width=4, color="rgb(11, 131, 191)"),
name="Décès 2019",
marker_size=4,
showlegend=True
))
fig.add_trace(go.Scatter(x = df_mortalite_france_2018["Date_evenement"], y = df_mortalite_france_2018["Total_deces_2018_diff"],
mode='lines',
line=dict(width=4, color="rgb(96, 178, 219)"),
name="Décès 2018",
marker_size=4,
showlegend=True
))
fig.add_trace(go.Scatter(x = df_mortalite_france["Date_evenement"], y = df_mortalite_france["Total_deces_2020_diff"],
mode='lines',
line=dict(width=4, color="#ffa58f"),
name="Décès 2020",
marker_size=4,
showlegend=True
))
fig.add_trace(go.Scatter(x = df_mortalite_france["Date_evenement"], y = df_mortalite_france["Total_deces_2021_diff"],
mode='lines',
line=dict(width=4, color="red"),
name="Décès 2021",
marker_size=4,
showlegend=True
))
mortalite_now = df_mortalite_france.dropna()["Total_deces_2021_diff"].values[-1]
fig.add_trace(go.Scatter(x = [df_mortalite_france.dropna()["Date_evenement"].values[-1]], y = [mortalite_now],
mode='markers',
name="",
line=dict(width=4, color="red"),
marker_color='red',
marker_size=10,
showlegend=False
))
# Modification du layout
fig.update_layout(
margin=dict(
l=0,
r=0,
b=50,
t=70,
pad=0
),
legend_orientation="h",
title={
'text': "<b>Mortalité en France</b><br><sub>Moyenne mobile de {} jours pour lisser les irrégularités. Derniers jours non consolidés.".format(window),
'y':0.95,
'x':0.5,
'xanchor': 'center',
'yanchor': 'top'},
titlefont = dict(
size=20),
annotations = [
dict(
x=0.5,
y=-0.1,
xref='paper',
yref='paper',
opacity=0.8,
text='Date : {}. Source : INSEE. Auteur : <NAME> - covidtracker.fr.'.format(datetime.now().strftime('%d %B %Y')), showarrow = False
)]
)
fig.update_xaxes(title="", nticks=10)
fig.update_yaxes(title="", rangemode="tozero")
name_fig = "mortalite"
fig.write_image(PATH+"images/charts/france/{}.jpeg".format(name_fig), scale=3, width=900, height=550)
fig.update_layout(
annotations = [
dict(
x=0.5,
y=1.05,
xref='paper',
yref='paper',
xanchor='center',
text='Cliquez sur des éléments de légende pour les ajouter/supprimer',
showarrow = False
)]
)
plotly.offline.plot(fig, filename = PATH+'images/html_exports/france/{}.html'.format(name_fig), auto_open=False)
print("> " + name_fig)
| src/france/covid19_france_insee.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import operator
profit = [60, 40, 100, 120, 150, 200]
obj = [10, 40, 20, 30, 60, 70]
maxweight = 100
n = 6
arr = []
for i in range(n):
arr.append([profit[i], obj[i], round(profit[i]/obj[i],2)])
arr = sorted(arr, reverse = True, key = operator.itemgetter(2))
print(arr)
profitmax = 0
fracobj = 0
for i in range(n):
if maxweight > 0 and arr[i][1] < maxweight:
maxweight -= arr[i][1]
profitmax += arr[i][0]
else:
fracobj = i
break
if maxweight > 0:
profitmax += maxweight * (arr[fracobj][0]/arr[fracobj][1])
print(round(profitmax, 2))
| Greedy/Fractional Knapsack(Greedy Method).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + deletable=true editable=true
# LSTM for Human Activity Recognition
Human activity recognition using smartphones dataset and an LSTM RNN. Classifying the type of movement amongst six categories:
- WALKING,
- WALKING_UPSTAIRS,
- WALKING_DOWNSTAIRS,
- SITTING,
- STANDING,
- LAYING.
## Video dataset overview
Follow this link to see a video of the 6 activities recorded in the experiment with one of the participants:
<a href="http://www.youtube.com/watch?feature=player_embedded&v=XOEN9W05_4A
" target="_blank"><img src="http://img.youtube.com/vi/XOEN9W05_4A/0.jpg"
alt="Video of the experiment" width="400" height="300" border="10" /></a>
<a href="https://youtu.be/XOEN9W05_4A"><center>[Watch video]</center></a>
## Details about input data
I will be using an LSTM on the data to learn (as a cellphone attached on the waist) to recognise the type of activity that the user is doing. The dataset's description goes like this:
> The sensor signals (accelerometer and gyroscope) were pre-processed by applying noise filters and then sampled in fixed-width sliding windows of 2.56 sec and 50% overlap (128 readings/window). The sensor acceleration signal, which has gravitational and body motion components, was separated using a Butterworth low-pass filter into body acceleration and gravity. The gravitational force is assumed to have only low frequency components, therefore a filter with 0.3 Hz cutoff frequency was used.
That said, I will use the almost raw data: only the gravity effect has been filtered out of the accelerometer as a preprocessing step for another 3D feature as an input to help learning.
## What is an RNN?
As explained in [this article](http://karpathy.github.io/2015/05/21/rnn-effectiveness/), an RNN takes many input vectors to process them and output other vectors. It can be roughly pictured like in the image below, imagining each rectangle has a vectorial depth and other special hidden quirks in the image below. **In our case, the "many to one" architecture is used**: we accept time series of feature vectors (one vector per time step) to convert them to a probability vector at the output for classification. Note that a "one to one" architecture would be a standard feedforward neural network.
<img src="http://karpathy.github.io/assets/rnn/diags.jpeg" />
An LSTM is an improved RNN. It is more complex, but easier to train, avoiding what is called the vanishing gradient problem and the exploding gradient problem.
## Results
Scroll on! Nice visuals awaits.
# + deletable=true editable=true
# All Includes
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import tensorflow as tf # Version r0.10
from sklearn import metrics
import os
# + deletable=true editable=true
# Useful Constants
# Those are separate normalised input features for the neural network
INPUT_SIGNAL_TYPES = [
"body_acc_x_",
"body_acc_y_",
"body_acc_z_",
"body_gyro_x_",
"body_gyro_y_",
"body_gyro_z_",
"total_acc_x_",
"total_acc_y_",
"total_acc_z_"
]
# Output classes to learn how to classify
LABELS = [
"WALKING",
"WALKING_UPSTAIRS",
"WALKING_DOWNSTAIRS",
"SITTING",
"STANDING",
"LAYING"
]
# + [markdown] deletable=true editable=true
# ## Let's start by downloading the data:
# + deletable=true editable=true
# Note: Linux bash commands start with a "!" inside those "ipython notebook" cells
DATA_PATH = "data/"
# !pwd && ls
os.chdir(DATA_PATH)
# !pwd && ls
# !python download_dataset.py
# !pwd && ls
os.chdir("..")
# !pwd && ls
DATASET_PATH = DATA_PATH + "UCI HAR Dataset/"
print("\n" + "Dataset is now located at: " + DATASET_PATH)
# + [markdown] deletable=true editable=true
# ## Preparing dataset:
# + deletable=true editable=true
TRAIN = "train/"
TEST = "test/"
# Load "X" (the neural network's training and testing inputs)
def load_X(X_signals_paths):
X_signals = []
for signal_type_path in X_signals_paths:
file = open(signal_type_path, 'rb')
# Read dataset from disk, dealing with text files' syntax
X_signals.append(
[np.array(serie, dtype=np.float32) for serie in [
row.replace(' ', ' ').strip().split(' ') for row in file
]]
)
file.close()
return np.transpose(np.array(X_signals), (1, 2, 0))
X_train_signals_paths = [
DATASET_PATH + TRAIN + "Inertial Signals/" + signal + "train.txt" for signal in INPUT_SIGNAL_TYPES
]
X_test_signals_paths = [
DATASET_PATH + TEST + "Inertial Signals/" + signal + "test.txt" for signal in INPUT_SIGNAL_TYPES
]
X_train = load_X(X_train_signals_paths)
X_test = load_X(X_test_signals_paths)
# Load "y" (the neural network's training and testing outputs)
def load_y(y_path):
file = open(y_path, 'rb')
# Read dataset from disk, dealing with text file's syntax
y_ = np.array(
[elem for elem in [
row.replace(' ', ' ').strip().split(' ') for row in file
]],
dtype=np.int32
)
file.close()
# Substract 1 to each output class for friendly 0-based indexing
return y_ - 1
y_train_path = DATASET_PATH + TRAIN + "y_train.txt"
y_test_path = DATASET_PATH + TEST + "y_test.txt"
y_train = load_y(y_train_path)
y_test = load_y(y_test_path)
# + [markdown] deletable=true editable=true
# ## Additionnal Parameters:
#
# Here are some core parameter definitions for the training.
#
# The whole neural network's structure could be summarised by enumerating those parameters and the fact an LSTM is used.
# + deletable=true editable=true
# Input Data
training_data_count = len(X_train) # 7352 training series (with 50% overlap between each serie)
test_data_count = len(X_test) # 2947 testing series
n_steps = len(X_train[0]) # 128 timesteps per series
n_input = len(X_train[0][0]) # 9 input parameters per timestep
# LSTM Neural Network's internal structure
n_hidden = 32 # Hidden layer num of features
n_classes = 6 # Total classes (should go up, or should go down)
# Training
learning_rate = 0.0025
lambda_loss_amount = 0.0015
training_iters = training_data_count * 300 # Loop 300 times on the dataset
batch_size = 1500
display_iter = 30000 # To show test set accuracy during training
# Some debugging info
print "Some useful info to get an insight on dataset's shape and normalisation:"
print "(X shape, y shape, every X's mean, every X's standard deviation)"
print (X_test.shape, y_test.shape, np.mean(X_test), np.std(X_test))
print "The dataset is therefore properly normalised, as expected, but not yet one-hot encoded."
# + [markdown] deletable=true editable=true
# ## Utility functions for training:
# + deletable=true editable=true
def LSTM_RNN(_X, _weights, _biases):
# Function returns a tensorflow LSTM (RNN) artificial neural network from given parameters.
# Moreover, two LSTM cells are stacked which adds deepness to the neural network.
# Note, some code of this notebook is inspired from an slightly different
# RNN architecture used on another dataset:
# https://tensorhub.com/aymericdamien/tensorflow-rnn
# (NOTE: This step could be greatly optimised by shaping the dataset once
# input shape: (batch_size, n_steps, n_input)
_X = tf.transpose(_X, [1, 0, 2]) # permute n_steps and batch_size
# Reshape to prepare input to hidden activation
_X = tf.reshape(_X, [-1, n_input])
# new shape: (n_steps*batch_size, n_input)
# Linear activation
_X = tf.nn.relu(tf.matmul(_X, _weights['hidden']) + _biases['hidden'])
# Split data because rnn cell needs a list of inputs for the RNN inner loop
_X = tf.split(0, n_steps, _X)
# new shape: n_steps * (batch_size, n_hidden)
# Define two stacked LSTM cells (two recurrent layers deep) with tensorflow
lstm_cell_1 = tf.nn.rnn_cell.BasicLSTMCell(n_hidden, forget_bias=1.0, state_is_tuple=True)
lstm_cell_2 = tf.nn.rnn_cell.BasicLSTMCell(n_hidden, forget_bias=1.0, state_is_tuple=True)
lstm_cells = tf.nn.rnn_cell.MultiRNNCell([lstm_cell_1, lstm_cell_2], state_is_tuple=True)
# Get LSTM cell output
outputs, states = tf.nn.rnn(lstm_cells, _X, dtype=tf.float32)
# Get last time step's output feature for a "many to one" style classifier,
# as in the image describing RNNs at the top of this page
lstm_last_output = outputs[-1]
# Linear activation
return tf.matmul(lstm_last_output, _weights['out']) + _biases['out']
def extract_batch_size(_train, step, batch_size):
# Function to fetch a "batch_size" amount of data from "(X|y)_train" data.
shape = list(_train.shape)
shape[0] = batch_size
batch_s = np.empty(shape)
for i in range(batch_size):
# Loop index
index = ((step-1)*batch_size + i) % len(_train)
batch_s[i] = _train[index]
return batch_s
def one_hot(y_):
# Function to encode output labels from number indexes
# e.g.: [[5], [0], [3]] --> [[0, 0, 0, 0, 0, 1], [1, 0, 0, 0, 0, 0], [0, 0, 0, 1, 0, 0]]
y_ = y_.reshape(len(y_))
n_values = np.max(y_) + 1
return np.eye(n_values)[np.array(y_, dtype=np.int32)] # Returns FLOATS
# + [markdown] deletable=true editable=true
# ## Let's get serious and build the neural network:
# + deletable=true editable=true
# Graph input/output
x = tf.placeholder(tf.float32, [None, n_steps, n_input])
y = tf.placeholder(tf.float32, [None, n_classes])
# Graph weights
weights = {
'hidden': tf.Variable(tf.random_normal([n_input, n_hidden])), # Hidden layer weights
'out': tf.Variable(tf.random_normal([n_hidden, n_classes], mean=1.0))
}
biases = {
'hidden': tf.Variable(tf.random_normal([n_hidden])),
'out': tf.Variable(tf.random_normal([n_classes]))
}
pred = LSTM_RNN(x, weights, biases)
# Loss, optimizer and evaluation
l2 = lambda_loss_amount * sum(
tf.nn.l2_loss(tf_var) for tf_var in tf.trainable_variables()
) # L2 loss prevents this overkill neural network to overfit the data
cost = tf.reduce_mean(tf.nn.softmax_cross_entropy_with_logits(pred, y)) + l2 # Softmax loss
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate).minimize(cost) # Adam Optimizer
correct_pred = tf.equal(tf.argmax(pred,1), tf.argmax(y,1))
accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# + [markdown] deletable=true editable=true
# ## Hooray, now train the neural network:
# + deletable=true editable=true
# To keep track of training's performance
test_losses = []
test_accuracies = []
train_losses = []
train_accuracies = []
# Launch the graph
sess = tf.InteractiveSession(config=tf.ConfigProto(log_device_placement=True))
init = tf.initialize_all_variables()
sess.run(init)
# Perform Training steps with "batch_size" amount of example data at each loop
step = 1
while step * batch_size <= training_iters:
batch_xs = extract_batch_size(X_train, step, batch_size)
batch_ys = one_hot(extract_batch_size(y_train, step, batch_size))
# Fit training using batch data
_, loss, acc = sess.run(
[optimizer, cost, accuracy],
feed_dict={
x: batch_xs,
y: batch_ys
}
)
train_losses.append(loss)
train_accuracies.append(acc)
# Evaluate network only at some steps for faster training:
if (step*batch_size % display_iter == 0) or (step == 1) or (step * batch_size > training_iters):
# To not spam console, show training accuracy/loss in this "if"
print "Training iter #" + str(step*batch_size) + \
": Batch Loss = " + "{:.6f}".format(loss) + \
", Accuracy = {}".format(acc)
# Evaluation on the test set (no learning made here - just evaluation for diagnosis)
loss, acc = sess.run(
[cost, accuracy],
feed_dict={
x: X_test,
y: one_hot(y_test)
}
)
test_losses.append(loss)
test_accuracies.append(acc)
print "PERFORMANCE ON TEST SET: " + \
"Batch Loss = {}".format(loss) + \
", Accuracy = {}".format(acc)
step += 1
print "Optimization Finished!"
# Accuracy for test data
one_hot_predictions, accuracy, final_loss = sess.run(
[pred, accuracy, cost],
feed_dict={
x: X_test,
y: one_hot(y_test)
}
)
test_losses.append(final_loss)
test_accuracies.append(accuracy)
print "FINAL RESULT: " + \
"Batch Loss = {}".format(final_loss) + \
", Accuracy = {}".format(accuracy)
# + [markdown] deletable=true editable=true
# ## Training is good, but having visual insight is even better:
#
# Okay, let's plot this simply in the notebook for now.
# + deletable=true editable=true
# (Inline plots: )
# %matplotlib inline
font = {
'family' : 'Bitstream Vera Sans',
'weight' : 'bold',
'size' : 18
}
matplotlib.rc('font', **font)
width = 12
height = 12
plt.figure(figsize=(width, height))
indep_train_axis = np.array(range(batch_size, (len(train_losses)+1)*batch_size, batch_size))
plt.plot(indep_train_axis, np.array(train_losses), "b--", label="Train losses")
plt.plot(indep_train_axis, np.array(train_accuracies), "g--", label="Train accuracies")
indep_test_axis = np.array(range(batch_size, len(test_losses)*display_iter, display_iter)[:-1] + [training_iters])
plt.plot(indep_test_axis, np.array(test_losses), "b-", label="Test losses")
plt.plot(indep_test_axis, np.array(test_accuracies), "g-", label="Test accuracies")
plt.title("Training session's progress over iterations")
plt.legend(loc='upper right', shadow=True)
plt.ylabel('Training Progress (Loss or Accuracy values)')
plt.xlabel('Training iteration')
plt.show()
# + [markdown] deletable=true editable=true
# ## And finally, the multi-class confusion matrix and metrics!
# + deletable=true editable=true
# Results
predictions = one_hot_predictions.argmax(1)
print "Testing Accuracy: {}%".format(100*accuracy)
print ""
print "Precision: {}%".format(100*metrics.precision_score(y_test, predictions, average="weighted"))
print "Recall: {}%".format(100*metrics.recall_score(y_test, predictions, average="weighted"))
print "f1_score: {}%".format(100*metrics.f1_score(y_test, predictions, average="weighted"))
print ""
print "Confusion Matrix:"
confusion_matrix = metrics.confusion_matrix(y_test, predictions)
print confusion_matrix
normalised_confusion_matrix = np.array(confusion_matrix, dtype=np.float32)/np.sum(confusion_matrix)*100
print ""
print "Confusion matrix (normalised to % of total test data):"
print normalised_confusion_matrix
print ("Note: training and testing data is not equally distributed amongst classes, "
"so it is normal that more than a 6th of the data is correctly classifier in the last category.")
# Plot Results:
width = 12
height = 12
plt.figure(figsize=(width, height))
plt.imshow(
normalised_confusion_matrix,
interpolation='nearest',
cmap=plt.cm.rainbow
)
plt.title("Confusion matrix \n(normalised to % of total test data)")
plt.colorbar()
tick_marks = np.arange(n_classes)
plt.xticks(tick_marks, LABELS, rotation=90)
plt.yticks(tick_marks, LABELS)
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label')
plt.show()
# + deletable=true editable=true
sess.close()
# + [markdown] deletable=true editable=true
# ## Conclusion
#
# Outstandingly, **the accuracy is of 91%**!
#
# This means that the neural networks is almost always able to correctly identify the movement type! Remember, the phone is attached on the waist and each series to classify has just a 128 sample window of two internal sensors (a.k.a. 2.56 seconds at 50 FPS), so those predictions are extremely accurate.
#
# I specially did not expect such good results for guessing between "WALKING" "WALKING_UPSTAIRS" and "WALKING_DOWNSTAIRS" as a cellphone. Thought, it is still possible to see a little cluster on the matrix between those 3 classes. This is great.
#
# It is also possible to see that it was hard to do the difference between "SITTING" and "STANDING". Those are seemingly almost the same thing from the point of view of a device placed on the belly, according to how the dataset was gathered.
#
# I also tried my code without the gyroscope, using only the two 3D accelerometer's features (and not changing the training hyperparameters), and got an accuracy of 87%.
#
#
# ## Improvements
#
# In [another repo of mine](https://github.com/guillaume-chevalier/HAR-stacked-residual-bidir-LSTMs), the accuracy is pushed up to 94% using a special deep bidirectional architecture, and this architecture is tested on another dataset. If you want to learn more about deep learning, I have built a list of ressources that I found to be useful [here](https://github.com/guillaume-chevalier/awesome-deep-learning-resources).
#
#
# ## References
#
# The [dataset](https://archive.ics.uci.edu/ml/datasets/Human+Activity+Recognition+Using+Smartphones) can be found on the UCI Machine Learning Repository.
#
# > <NAME>, <NAME>, <NAME>, <NAME> and <NAME>. A Public Domain Dataset for Human Activity Recognition Using Smartphones. 21th European Symposium on Artificial Neural Networks, Computational Intelligence and Machine Learning, ESANN 2013. Bruges, Belgium 24-26 April 2013.
#
# If you want to cite my work, you can point to the URL of the GitHub repository:
# > https://github.com/guillaume-chevalier/LSTM-Human-Activity-Recognition
#
# ## Connect with me
#
# - https://ca.linkedin.com/in/chevalierg
# - https://twitter.com/guillaume_che
# - https://github.com/guillaume-chevalier/
# + deletable=true editable=true
# Let's convert this notebook to a README as the GitHub project's title page:
# !jupyter nbconvert --to markdown LSTM.ipynb
# !mv LSTM.md README.md
| LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 作業目標:
#
# 通過建立多層的神經網路, 了解權值矩陣更新
# # 作業重點:
#
# 3 層神經網路
#
# 通過增加更多的中間層,以對更多關係的組合進行建模
#
# syn1 權值矩陣將隱層的組合輸出映射到最終結果,
#
# 而在更新 syn1 的同時,還需要更新 syn0 權值矩陣,
#
# 以從輸入資料中更好地產生這些組合
# +
import numpy as np
# Sigmoid 函數可以將任何值都映射到一個位於 0 到 1 範圍內的值。通過它,我們可以將實數轉化為概率值
def nonlin(x,deriv=False):
if(deriv==True):
return x*(1-x)
return 1/(1+np.exp(-x))
X = np.array([ [0,0,1],
[0,1,1],
[1,0,1],
[1,1,1] ])
# define y for output dataset
y = np.array([[0,0,1,1]]).T
# +
# seed random numbers to make calculation
# deterministic (just a good practice)
np.random.seed(1)
#亂數設定產生種子得到的權重初始化集仍是隨機分佈的,
#但每次開始訓練時,得到的權重初始集分佈都是完全一致的。
# initialize weights randomly with mean 0
syn0 = 2*np.random.random((3,2)) - 1
# define syn1
syn1 = 2*np.random.random((2,1)) - 1
iter = 0
syn0_history = [syn0]
syn1_history = [syn1]
#該神經網路權重矩陣的初始化操作。
#用 “syn0” 來代指 (即“輸入層-第一層隱層”間權重矩陣)
#用 “syn1” 來代指 (即“輸入層-第二層隱層”間權重矩陣)
# -
# 神經網路訓練
# for 迴圈反覆運算式地多次執行訓練代碼,使得我們的網路能更好地擬合訓練集
# +
for iter in range(10000):
# forward propagation
l0 = X
l1 = nonlin(np.dot(l0,syn0))
l2 = nonlin(np.dot(l1,syn1))
'''
新增
l2_error 該值說明了神經網路預測時“丟失”的數目。
l2_delta 該值為經確信度加權後的神經網路的誤差,除了確信誤差很小時,它近似等於預測誤差。
'''
# how much did we miss?
l1_error = y - l1
l2_error = y - l2
# multiply how much we missed by the
# slope of the sigmoid at the values in l1
l1_delta = l1_error * nonlin(l1,True)
l2_delta = l2_error * nonlin(l2,True)
# update weights
syn0 += np.dot(l0.T,l1_delta)
syn1 += np.dot(l1.T,l2_delta)
# syn1 update weights
print("Output After Training:")
print(l1)
print("\n\n")
print(l2)
# +
import matplotlib.pyplot as plt
plt.plot(syn0_history[0], ms=3, lw=1.5, color='black')
plt.xlabel(r'$L1$', fontsize=16)
plt.show()
# -
plt.plot(syn1_history[0], ms=3, lw=1.5, color='black')
plt.xlabel(r'$L1$', fontsize=16)
plt.show()
| D75_BackPropagation/Day75-Back_Propagation_HW.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="d5lxxAgsa6go"
# # Metadata
#
# ```
# Course: DS 5001
# Module: 03 Lab
# Topic: Baby Babel
# Author: <NAME>
#
# Purpose: Implements the Library of Babel with a small symbol set and message length.
# ```
# -
# # Set Up
# + colab={} colab_type="code" id="NhxIat0Oa6gz"
import re
import pandas as pd
import numpy as np
import seaborn as sns
from IPython.core.display import HTML
# -
sns.set()
data_home = '../data'
# + [markdown] colab_type="text" id="iWg2R7dFa6gu"
# # Mini Babel
#
# We create a miniature Library of Babel, one based on only four characters, and a message length of six.
# + [markdown] colab_type="text" id="PE1iQpcPa6hG" toc-hr-collapsed=false
# ## The Symbol Set
# + colab={} colab_type="code" id="BpgbcvT8a6hK"
mini_alpha = list('abt ')
# + [markdown] colab_type="text" id="NsTcj8Osa6hV"
# ## All possible Messages of Length 6
#
# This is a clumsy but visually effective way to demonstrate how the Library of Babel might have been constructed. It is essentially the cartesian product of the alphabet, multiplying by the length of the message.
# -
mini_library_list = []
for L1 in mini_alpha:
for L2 in mini_alpha:
for L3 in mini_alpha:
for L4 in mini_alpha:
for L5 in mini_alpha:
for L6 in mini_alpha:
mini_library_list.append(''.join((L1,L2,L3,L4,L5,L6)))
# How many books are in the library?
len(mini_library_list), len(mini_alpha) ** 6
# Can we find a specific book?
mini_library_list.index('at bat')
# ## The Pandas Way
#
# Pandas provides a method -- `pd.MultiIndex.from_product()` -- to create a cartesian product of an arbitrary list of lists.
#
# Let's create a library based on a book length $L = 6$.
L = 6
# + colab={} colab_type="code" id="wqGL3b0Fa6kI"
alpha_lists = [mini_alpha] * L
book_idx = pd.MultiIndex.from_product(alpha_lists)
mini_library = pd.DataFrame(index=book_idx)\
.reset_index()\
.sum(1)\
.to_frame('book')
#.apply(lambda x: ''.join(x), 1)\
# -
mini_library
# Should be the same as $|a|^L$ where $|a|$ is the symbol set size and $L$ is the average message length.
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" executionInfo={"elapsed": 448, "status": "ok", "timestamp": 1549030949938, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-gvKWs7zR4JY/AAAAAAAAAAI/AAAAAAABqfk/Q8O12g6M_T4/s64/photo.jpg", "userId": "11010075019714369526"}, "user_tz": 300} id="iVHotJmNa6iM" outputId="5114039a-ed94-4b8c-ee0f-8df8764b195e"
len(mini_library) == len(mini_alpha)**L
# + colab={"base_uri": "https://localhost:8080/", "height": 201} colab_type="code" executionInfo={"elapsed": 478, "status": "ok", "timestamp": 1549030983594, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-gvKWs7zR4JY/AAAAAAAAAAI/AAAAAAABqfk/Q8O12g6M_T4/s64/photo.jpg", "userId": "11010075019714369526"}, "user_tz": 300} id="7Gd0VNi5a6kR" outputId="2046cb63-79de-4b6b-f077-1b0ac763afae"
mini_library[mini_library.book == 'at bat']
# -
# ## Probability of a book
N = len(mini_library)
assert N == len(mini_alpha)**L
p_book = 1 / N
p_book
# ## Entropy of `mini_library`
#
# Max Entropy: $H_{max} = \sum_N\frac{1}{N}\log_2(\frac{N}{1}) = N\frac{1}{N}\log_2(\frac{N}{1}) = \log_2(N)$
H_max = np.log2(N)
H_max
# + [markdown] colab_type="text" id="S7b2R-qia6lc"
# ## Sample text
# + colab={} colab_type="code" id="re8N-CgYa6le"
mini_text = mini_library.sample(100, replace=True).book.str.cat(sep=' ')
# -
mini_text
# + colab={} colab_type="code" id="re8N-CgYa6le"
display(HTML(mini_text))
# + [markdown] colab_type="text" id="vjbAq589a6mN" toc-hr-collapsed=false
# # Bigger Babel
# -
class UnigramModel():
"""A simple character level language model. A language model is just
the sample space of the symbol system with associated probabilities."""
alpha:[] = list(' abcdefghijklmnopqrstuvwxyz')
def __init__(self):
self.model = pd.DataFrame(index=self.alpha)
self.model.index.name = 'char'
self.model['n'] = 1
self.model['p_x'] = 1 / len(self.alpha)
def update_weights(self, char_str=''):
self.char_str = char_str.lower()
self.chars = pd.Series(list(self.char_str))
self.chars = self.chars[self.chars.isin(self.alpha)]
self.model['n'] = self.chars.value_counts()
self.model['p_x'] = self.model.n / self.model.n.sum()
UGM = UnigramModel()
UGM.model
class Babel():
"""Generate messages based on a character level language model."""
msg_len:int = 40 * 80
use_html = True
def __init__(self, UGM:UnigramModel=UGM):
self.UGM = UGM
def get_message(self):
self.msg = self.UGM.model.sample(self.msg_len, weights='p_x', replace=True).index.str.cat()
if self.use_html:
self.msg = f"<div style='width:6in;font-size:14pt;font-family:monospace;'>{self.msg}</div>"
def print_message(self):
if self.use_html:
display(HTML(self.msg))
else:
display(self.msg)
B1 = Babel(UGM)
B1.get_message()
B1.print_message()
# + [markdown] colab_type="text" id="MOoxIda_a6mg" toc-hr-collapsed=false
# # Add Data to Model
# -
# ## Import corpus
# + colab={} colab_type="code" id="CF72yPC1a6mm" toc-hr-collapsed=true
text_csv = f'{data_home}/output/austen-combo.csv'
# -
text_df = pd.read_csv(text_csv)
text_df.head()
# ## Convert to one big string
text_str = text_df.token_str.str.cat(sep=' ')
text_str[:80]
CHARS = pd.DataFrame(dict(char_token=list(text_str)))
CHARS['char_type'] = CHARS.char_token.str.lower()
CHARSET = CHARS.char_type.value_counts().to_frame('n')
CHARSET.plot.bar(rot=0);
# ## Update weights in model
UGM.update_weights(text_str)
UGM.model.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 880} colab_type="code" executionInfo={"elapsed": 778, "status": "ok", "timestamp": 1549031115347, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-gvKWs7zR4JY/AAAAAAAAAAI/AAAAAAABqfk/Q8O12g6M_T4/s64/photo.jpg", "userId": "11010075019714369526"}, "user_tz": 300} id="9AhzPdhFa6nK" outputId="b3c42708-68a8-4e37-bed6-67963a9a4c4b"
UGM.model.p_x.sort_values().plot(kind='barh', figsize=(5,10));
# -
B1.get_message()
B1.print_message()
# + [markdown] colab_type="text" id="KagqA-bLa6nj" toc-hr-collapsed=true
# # Create Bigram Model
#
# The index is the sample space.
# -
class BigramModel():
alpha:[] = list(' abcdefghijklmnopqrstuvwxyz')
def __init__(self):
self.idx = pd.MultiIndex.from_product([self.alpha, self.alpha], names=['char_x', 'char_y'])
self.model = pd.DataFrame(dict(n=1, p_xy=(1/len(self.idx))), index=self.idx).sort_index()
def update_weights(self, text_str):
char_list = [char for char in text_str if char in self.alpha]
df_cols = dict(
char_x = [' '] + char_list,
char_y = char_list + [' ']
)
self.model['n'] = pd.DataFrame(df_cols).value_counts()\
.to_frame()
self.model.n = self.model.n.fillna(0)
self.model.n += 1 # LaPlace smoothing
self.model['p_xy'] = self.model.n / self.model.n.sum()
def add_conditional_probs(self):
self.model['p_yGx'] = self.model.groupby('char_x')\
.apply(lambda row: row.n / row.n.sum())\
.to_frame('p_yGx').droplevel(0)
def get_conditional_entropy(self):
"""Computes the entropy for each character of the distribution of following characters."""
self.H = self.model.groupby('char_x').apply(lambda row: row.p_yGx * np.log2(1/row.p_yGx))\
.droplevel(0).to_frame('h_yGx')\
.groupby('char_x').h_yGx.sum().to_frame()
BGM = BigramModel()
# ## Get Data to Estimate Model
BGM.update_weights(text_str)
# ## Add Conditional Probabilities
BGM.add_conditional_probs()
# + tags=[]
BGM.model.p_yGx.sort_values(ascending=False).head(10).plot.barh();
# + [markdown] colab_type="text" id="aYKNbehTa6qG"
# ## Get conditional entropy of characters as antecendents
#
# Note that all the vowels have high entropy rates.
# -
BGM.get_conditional_entropy()
BGM.H.h_yGx.sort_values().plot.barh(figsize=(10,10));
# ## Look at Examples
X = BGM.model.p_yGx.unstack()
X = round(X * 100, 2)
X.style.format("{:.2f}").background_gradient(cmap='YlGnBu', axis=None)
sns.set(rc = {'figure.figsize':(15,8)})
sns.heatmap(data=BGM.model.p_yGx.unstack(), cmap='YlGnBu', square=True, vmin=0, vmax=1, cbar=False);
def plot_char(char):
global BGM
h = BGM.H.loc[char].h_yGx.round(2)
title = f"Char {char}, H={h}"
BGM.model.loc[char].p_yGx.sort_values(ascending=False).plot.bar(figsize=(10,2), title=title);
plot_char('q')
plot_char('v')
plot_char('h')
plot_char('p')
plot_char('a')
plot_char(' ')
# + [markdown] colab_type="text" id="GFJO-qWfa6qo" toc-hr-collapsed=true
# # Generate text
# -
english_words = set([word.strip().lower()
for word in open(f"{data_home}/misc/english-words.txt", 'r').readlines()])
class Babel2():
"""Generate messages based on a character level language model."""
msg_len:int = 80 * 40
use_html = True
def __init__(self, BGM:BigramModel, english_words):
self.BGM = BGM
self.english_words = english_words
def get_message(self):
self.msg = ' '
for i in range(self.msg_len):
self.msg += self.BGM.model.loc[self.msg[-1]]\
.sample(weights='p_yGx').index.values[0]
self.tokens = pd.DataFrame(self.msg.split(), columns=['token_str'])
self.vocab = self.tokens.token_str.value_counts().to_frame('n')
self.vocab['en'] = False
self.vocab.loc[self.vocab.index.isin(english_words), 'en'] = True
self.vocab['len'] = self.vocab.index.str.len()
self.tokens['en'] = self.tokens.token_str.map(self.vocab.en)
def print_message(self):
if self.use_html:
html_msg = ''
for token in self.msg.split():
if token in english_words:
token = f"<b style='color:red;'>{token}</b>"
html_msg += ' ' + token
self.msg = f"<p style='color:gray;width:6in;font-size:14pt;font-family:monospace;'>{html_msg}</p>"
display(HTML(self.msg))
else:
display(self.msg)
# + colab={} colab_type="code" id="gQyo_aVla6qr"
B2 = Babel2(BGM, english_words)
B2.get_message()
B2.print_message()
# -
# # Look at Babel Vocab Stats
# ## English words
B2.vocab.loc[B2.vocab.en == True, ['n','len']]\
.sort_values('n', ascending=False)
# ## Type and token ratios
type_rate = round(B2.vocab[B2.vocab.en == True].n.count() / B2.vocab.n.count(), 2)
token_rate = round(B2.vocab[B2.vocab.en == True].n.sum() / B2.vocab.n.sum(), 2)
type_rate, token_rate, round(type_rate/token_rate, 2)
# ## Long words
B2.vocab.query("en == False").sort_values('len', ascending=False).head(20)
# ## Word lengths
# + tags=[]
B2.vocab.len.value_counts().sort_index().plot.bar();
# + tags=[]
B2.vocab.query("en == True").len.value_counts().sort_index().plot.bar();
# -
B2.vocab.len.mean()
B2.vocab[B2.vocab.en == True].len.mean()
# # Memorable passwords?
#
# Words that are not English but which are based on the language model and are of average length.
B2.vocab[(B2.vocab.en == False) & B2.vocab.len.isin([6,7,8])].sample(10)
# # Simplified Polynesian
#
# We implement the Simplified Polynesian example in Manning and Schutz.
poly_chars = list("ptkaiu")
poly_probs = [1/8,1/4,1/8,1/4,1/8,1/8]
POLY = pd.DataFrame(dict(char=poly_chars, p=poly_probs)).set_index('char')
POLY['s'] = 1 / POLY.p
POLY['i'] = np.log2(POLY.s)
POLY['h'] = POLY.p * POLY.i
PH = POLY.h.sum()
POLY
# We can generate words from this -- demonstrating that speech is a **selection process**.
POLY.sample(weights='p', n=5).index.str.cat(sep='')
PH
# So, we can construct a code that translates each character into two or three binary digits
POLY['code'] = ['100','00', '101', '01', '110', '111']
POLY['code_len'] = POLY.code.str.len()
POLY
# Note that the code length equals the information value!
# However, H does not equal mean information. It's close, tho!
round(POLY.i.mean(), 2)
PR = 1 - (PH / np.log2(6))
PR
# ## Conditional
POLYBG = pd.DataFrame(dict(p=0), index = pd.MultiIndex.from_product([list("aiu"), list("ptk")]))
POLYBG
bg_vals = """
a p 1/16
a t 3/8
a k 1/16
i p 1/16
i t 3/16
i k 0/1
u p 0/1
u t 3/16
u k 1/16
""".split('\n')[1:-1]
for bg_val in bg_vals:
x, y, z = bg_val.split()
n, d = z.split('/')
POLYBG.loc[(x,y), 'p'] = int(n) / int(d)
POLYBG.unstack()
POLYBG.reset_index()
PBG = POLYBG.reset_index()
PBG
| M03_LanguageModels/M03_01_Babel.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
BIN = '../../'
sys.path.append(BIN)
import utils
import matplotlib.pyplot as plt
from scipy import stats
import my_matplotlib_style as ms
# %matplotlib inline
# %matplotlib inline
import sys
BIN = '../../'
sys.path.append(BIN)
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
#from sklearn.model_selection import train_test_split
import torch
import torch.nn as nn
#import torch.nn.parallel
import torch.optim as optim
import torch.utils.data
from torch.autograd import Variable
from torch.utils.data import TensorDataset
from torch.utils.data import DataLoader
import my_matplotlib_style as ms
from fastai import data_block, basic_train, basic_data
from fastai.callbacks import ActivationStats
import fastai
import matplotlib as mpl
mpl.rc_file(BIN + 'my_matplotlib_rcparams')
from nn_utils import AE_big, AE_3D_200
from utils import plot_activations
# Load data
train = pd.read_pickle(BIN + 'processed_data/aod/uncompressed_all_jets_train.pkl')
test = pd.read_pickle(BIN + 'processed_data/aod/uncompressed_all_jets_test.pkl')
trainc = pd.read_pickle(BIN + 'processed_data/aod/compressed_all_jets_train.pkl')
testc = pd.read_pickle(BIN + 'processed_data/aod/compressed_all_jets_test.pkl')
branches = ['pt','eta','phi','m'] + ["ActiveArea4vec_eta", "ActiveArea4vec_m", "ActiveArea4vec_phi", "ActiveArea4vec_pt", "Jvt", "JVFCorr" ,"JvtRpt", "FracSamplingMax", "FracSamplingMaxIndex", "Width", "EMFrac","Timing"]
# %matplotlib inline
n_bins = 500
alph = 0.8
#for kk in np.arange(4):
for kk in [0,3]:
plt.figure(kk + 4, figsize=(5,5))
plt.figure(figsize=(6,6))
#plt.ticklabel_format(style='sci', axis='x', scilimits=(0,0))
n_hist_data, bin_edges, _ = plt.hist(train[branches[kk]], color='black', label='Input', alpha=1, bins=n_bins,histtype="step",linewidth=2)
n_hist_datac, _, _ = plt.hist(trainc[branches[kk]], color='orange', label='Compressed Input', alpha=1, bins=bin_edges,histtype="step")
#n_hist_pred, _, _ = plt.hist(pred[:, kk], color=colors[0], label='Output', alpha=alph, bins=bin_edges)
#plt.suptitle(branches[kk])
plt.xlabel(branches[kk])
plt.ylabel('Number of events')
plt.xscale('log')
ms.sciy()
#plt.xlim(-5, 10)
# plt.yscale('log')
#plt.legend(loc=4)
fig_name = 'trainforever_hist_%s' % train.columns[kk]
plt.tight_layout()
plt.savefig("floatcompression_overlaidinput_"+branches[kk])
# -
| examples/float_compression/16D/input_comparison.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# <h1 style='font-size:4rem;color:red;'>Math 267 Project #3
# + [markdown] tags=[]
# ---
#
# ## Solution
#
# ---
# -
# # Spring mass system.
#
# The second order equation of an unforced spring/mass system with mass m, damping constant b and spring constant k is given by:<p>
# $$ mx''(t) + bx'(t) +kx(t) = 0. $$
#
#
# Recall that $x(t)$ represents the displacement of the mass from its resting position ( with x>0 the spring is stretched and x<0 the spring is compressed). Therefore the velocity of the moving mass is $ x'(t)$ and the acceleration is $x’’(t)$. To approximate the solution for this system using Euler’s method we convert the equation into a system by introducing the variable
#
# $$ y = x’.$$
#
# So our two unknowns are: $x$ for the position of the mass and $y$ for the velocity of the mass. As a system the second order equation becomes ( using m = 1)
#
# \begin{align}
# x'(t) &= \; y(t) \\
# y'(t) &= -k \cdot x(t) - b \cdot y(t)
# \end{align}
#
#
# and unlike the Predator-Prey model of exercise 2. this system is linear and can be represented in matrix notation:
#
#
# \begin{align}
# \begin{pmatrix}
# x'(t) \\
# y'(t)
# \end{pmatrix}=
# \begin{pmatrix}
# 0 & 1 \\
# -k & -b
# \end{pmatrix}\cdot
# \begin{pmatrix}
# x(t) \\
# y(t)
# \end{pmatrix}
# \end{align}
#
#
# ---
# ### Collaboration. Students are allowed to work together on this project. Colaboration is encouraged.
# However you final submission must be your own work.
#
# <img src = "https://github.com/rmartin977/math---267-Spring-2022/blob/main/Euclid.png?raw=true" style = "width:500px;height:500px" class="center">
# ### Run the cell below to import the necessary libraries.
# + id="f52331cb-8777-4d71-b19e-b0e890ccf61a"
# import libraries
import numpy as np
import matplotlib.pyplot as plt
from IPython.display import Image
# uncomment the line below if you are running a macbook
# %config InlineBackend.figure_format ='retina'
# -
# ## Exericse 1.
#
# For this exercise you will approximate the solution of an unforced mass-spring system with
# mass m =1, damping constant b = 1 and spring constant k = 2. The second order equation for this system is given by:
#
# $$x''(t) + 1x'(t) + 2x(t) = 0.$$
#
# Use the initial conditions $x(0) = 0$ and $ x'(0)=y(0) = 1$. Use the improved Euler’s method to solve the system. You will need to convert the second order equation to a system as demonstrated above. Choose a Δt=h of 0.1 and a time interval of 12 seconds. Create two plots. One showing x(t) versus time and the second showing the solution curve "trajectory" for the system in the xy (phase) plane.
#
#
# + [markdown] tags=[]
# ## Complete the code cell below to solve the sytstem.
# +
# define the slope functions
def f(x,y):
return y
def g(x,y):
return -1*y -2*x
h = 0.1 # set delta t
t = np.arange(0,12,h)
# Initialeze arrays to store the results.
x = np.zeros_like(t)
y = np.zeros_like(t)
# Set initial conditions:
x[0] = 0
y[0] = 1
# implement Euler's method
for i in range (len(t)-1):
F1 = f(x[i],y[i])
G1 = g(x[i],y[i])
F2 = f(x[i]+F1*h,y[i]+G1*h)
G2 = g(x[i]+F1*h,y[i]+G1*h)
slope1 = (F1+F2)/2
slope2 = (G1+G2)/2
x[i+1] = x[i] + slope1 * h
y[i+1] = y[i] + slope2 * h
# -
# ### Execute the cell below to graph your results.
# You should see a damped sinusoid.
plt.figure(figsize=(8,5))
plt.plot(t,x)
plt.grid()
plt.xlabel('t')
plt.ylabel('x');
plt.title("Damping b = 1");
# ### Execute the cell below to see the trajectory in the phase plane for this problem.
# You should see a spiral.
plt.figure(figsize=(8,5))
plt.plot(x,y,linewidth=2)
plt.xlim(-1.5,1.5)
plt.ylim(-1.5,1.5)
plt.grid()
plt.xlabel('x-displacement')
plt.ylabel('y-velocity');
plt.title("Trajectory for spring mass system b = 1");
# ### Repeat the steps above for b=3 and b=0. Copy and paste cells above and make the necessary modifications. You should show time plots and phase plane plots for b=0 and b=3.
#
# +
# b = 0
# define the slope functions
def f(x,y):
return y
def g(x,y):
return -2*x
h = 0.1 # set delta t
t = np.arange(0,12,h)
# Initialeze arrays to store the results.
x = np.zeros_like(t)
y = np.zeros_like(t)
# Set initial conditions:
x[0] = 0
y[0] = 1
# implement Euler's method
for i in range (len(t)-1):
F1 = f(x[i],y[i])
G1 = g(x[i],y[i])
F2 = f(x[i]+F1*h,y[i]+G1*h)
G2 = g(x[i]+F1*h,y[i]+G1*h)
slope1 = (F1+F2)/2
slope2 = (G1+G2)/2
x[i+1] = x[i] + slope1 * h
y[i+1] = y[i] + slope2 * h
# -
plt.figure(figsize=(8,5))
plt.plot(t,x)
plt.grid()
plt.xlabel('t')
plt.ylabel('x');
plt.xticks(np.arange(0,13,.5))
plt.title("Damping b = 0");
plt.figure(figsize=(8,5))
plt.plot(x,y,linewidth=2)
plt.xlim(-1.5,1.5)
plt.ylim(-1.5,1.5)
plt.grid()
plt.xlabel('x-displacement')
plt.ylabel('y-velocity');
plt.title("Trajectory for spring mass system b = 0");
# +
# b=3
# define the slope functions
def f(x,y):
return y
def g(x,y):
return -3*y -2*x
h = 0.1 # set delta t
t = np.arange(0,12,h)
# Initialeze arrays to store the results.
x = np.zeros_like(t)
y = np.zeros_like(t)
# Set initial conditions:
x[0] = 0
y[0] = 1
# implement Euler's method
for i in range (len(t)-1):
F1 = f(x[i],y[i])
G1 = g(x[i],y[i])
F2 = f(x[i]+F1*h,y[i]+G1*h)
G2 = g(x[i]+F1*h,y[i]+G1*h)
slope1 = (F1+F2)/2
slope2 = (G1+G2)/2
x[i+1] = x[i] + slope1 * h
y[i+1] = y[i] + slope2 * h
# -
plt.figure(figsize=(8,5))
plt.plot(t,x)
plt.grid()
plt.xlabel('t')
plt.ylabel('x');
plt.title("Damping b = 3");
plt.figure(figsize=(8,5))
plt.plot(x,y,linewidth=2)
plt.xlim(-1.5,1.5)
plt.ylim(-1.5,1.5)
plt.grid()
plt.xlabel('x-displacement')
plt.ylabel('y-velocity');
plt.title("Trajectory for spring mass system b = 3");
# ---
# # Exercise 2.
#
# For this exercise you will approximate the solution of an undamped periodically forced spring-mass system with mass m =1 and spring constant k = 1, no damping. The second order equation for this system is given by:
#
# $$ x''(t) + x(t) = Cos(\omega t).$$
#
# Use the initial conditions $x(0) = 0$ and $ x'(0) = y(0) = 0$. Use the improved Euler’s method to solve the system. Note this system is not autonomous so the slope functions could depend on x,y and t. Generate plots of $x(t)$ versus time for ω = 1.1 and ω = 1. Use a time interval of 50 seconds for $ \omega = 1 $ and 150 seconds for $ \omega = 1.1$. For both cases set Δt=0.1. Duplicate and modify the code above to generate the plots. Label the plots appropriately. There are no phase plane plots for Exercise 2.
# ## Solution for $ \omega = 1. $
# +
# define the slope functions
def f(x,y):
return y
def g(x,y,t):
return -1*x + np.cos(t)
h = 0.1 # set delta t
t = np.arange(0,50,h)
# Initialeze arrays to store the results.
x = np.zeros_like(t)
y = np.zeros_like(t)
# Set initial conditions:
x[0] = 0
y[0] = 0
# implement Euler's method
for i in range (len(t)-1):
F1 = f(x[i],y[i])
G1 = g(x[i],y[i],t[i])
F2 = f(x[i]+F1*h,y[i]+G1*h)
G2 = g(x[i]+F1*h,y[i]+G1*h,t[i+1])
slope1 = (F1+F2)/2
slope2 = (G1+G2)/2
x[i+1] = x[i] + slope1 * h
y[i+1] = y[i] + slope2 * h
# -
plt.figure(figsize=(8,5))
plt.plot(t,x)
plt.grid()
plt.xlabel('t')
plt.ylabel('x');
plt.title("Forced Spring-Mass system no damping $\omega = 1.$");
# ## Solution for $ \omega = 1.1 $
# +
# define the slope functions
def f(x,y):
return y
def g(x,y,t):
return -1*x + np.cos(1.1*t)
h = 0.1 # set delta t
t = np.arange(0,150,h)
# Initialeze arrays to store the results.
x = np.zeros_like(t)
y = np.zeros_like(t)
# Set initial conditions:
x[0] = 0
y[0] = 0
# implement Euler's method
for i in range (len(t)-1):
F1 = f(x[i],y[i])
G1 = g(x[i],y[i],t[i])
F2 = f(x[i]+F1*h,y[i]+G1*h)
G2 = g(x[i]+F1*h,y[i]+G1*h,t[i+1])
slope1 = (F1+F2)/2
slope2 = (G1+G2)/2
x[i+1] = x[i] + slope1 * h
y[i+1] = y[i] + slope2 * h
# -
plt.figure(figsize=(12,5))
plt.plot(t,x)
plt.grid()
plt.xlabel('t')
plt.ylabel('x');
plt.title("Forced Srping-Mass system no damping $\omega = 1.1$");
# ## Answer the questions below. Create a text cell to enter your answers.
# 1. Expain and discuss the results of exercise 1.
#
# For exercise \#1 we see the spring behaviour under three damping conditions. <p>
# b = 0 ~ we see no damping and the spring oscillates.<p>
# b = 1 ~ we see an underdamped behaviour and the oscillation decays.<p>
# b = 3 ~ we see overdamped and the spring returns to equilibrium with no oscillations.<p>
#
# 2. Compute the period of oscillation for exerice 1. b=0. Compare to value obtained from the graph.
#
# The period is 2*pi/omega = sqrt(2)*pi = 4.44. This is close to the value from the graph.
#
#
# 3. Discuss the results of exercise 2. Hint: look at the envelopes.
#
# For omega = 1, we are exciting the system at the resonant frequency and thus see RESONANCE, that is the osicllation are increasing.
#
# For omega = 1.1, the exciting frequencey is "close" to the resonant frequency and we observe the BEATING phenomenon.
| Project#3_solution.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Problemas dependientes del tiempo
#
# Nota: a este documento lo podremos ejecutar de manera dinámica si tienen instalado:
#
# - Python 3.5 o más nuevo instalado.
# - [Jupyter Notebook](https://jupyter.readthedocs.io/en/latest/install.html).
# - [FEniCS](https://fenicsproject.org/).
#
# La visualización del mismo es óptima utilizando Jupyter Notebook.
#
# ### Referencias
# - Capítulo 5 de <NAME>, <NAME>, The Finite Element Method: Theory, Implementation, and Applications.
# - Capítulo 6 de <NAME> and <NAME>, [Introduction to Numerical Methods for Variational Problems](https://www.springer.com/gp/book/9783030237875) (2019), Springer.
# ## Introducción
#
# La mayoría de los problemas en la vida real son dependientes del tiempo. Por ejemplo, en el problema que veremos en este tutorial aparece un factor dependiente de la derivada de la temperatura respecto del tiempo $\frac{dT}{dt}$. Si la geometría es una barra unidimensional de largo $L$ y la temperatura es la variable que queremos calcular entonces ahora tendremos dependencia de la posición $x$ y del tiempo $t$. Entonces, podemos tomar tres estrategias:
#
# 1. Usar primero diferencias finitas para estimar la derivada respecto del tiempo y luego se tendrá el problema de discretización espacial, que lo resolveremos por elementos finitos.
# 2. Primero discretizar el espacio y luego el tiempo.
# 3. Discretizar espacio y tiempo a la vez.
#
# Tomaremos la primer opción. Tal como se estudia habitualmente en la resolución de [ecuaciones diferenciales ordinarias](https://es.wikipedia.org/wiki/Ecuaci%C3%B3n_diferencial_ordinaria), existen diferentes opciones para aproximar en diferencias finitas una derivada de primer órden:
#
# - Método de Euler hacia atrás (regresivo, anterior, se puede encontrar de varias maneras, en inglés *backward Euler method*).
#
# $$ \frac{du \left(t_{n}\right)}{dt}=\dot{u}\left(t_{n}\right) \approx \frac{u^{n}-u^{n-1}}{\Delta t} \tag{1}$$
#
# Se llama regresivo porque estamos calculando la derivada del paso $n$ utilizando el mismo paso y el anterior ($n-1$). A este método también se lo denomina *implícito* (ya veremos porque). Lo bueno del método es que es *incondicionalmente estable* (nunca va a crecer de manera indefinida) y $\Delta t$ puede ser elegido arbitrariamente grande (aunque se pierde presición). Es una aproximación de *primer órden*.
#
# - Método de Euler hacia adelante (progresivo, adelantado, en inglés *forward Euler method*).
#
# $$\frac{du \left(t_{n}\right)}{dt}= \dot{u}\left(t_{n}\right) \approx \frac{u^{n+1}-u^{n}}{\Delta t} \tag{2}$$
#
# Note que aquí suponemos que el paso de tiempo es fijo ($\Delta t$) y hemos usado la notación $u^{n}=u\left(t_{n}\right)$. Se llama progresivo porque estamos calculando la derivada del paso $n$ utilizando el mismo paso y el paso futuro ($n+1$). Este método se lo denomina *explícito*. Este método es *condicionalmente estable*, dado que si se elije muy grande el paso puede crecer la solución (además de perder presición). Es también una aproximación de *primer órden*.
#
#
# Existen otros métodos (ver Apéndice 1 de Langtangen and Mardal, 2019), uno de los más conocidos y más aproximado es el Runge-Kuta, que es un *método explícito de cuarto órden*. Nosotros en el ejemplo siguiente utilizaremos el método de Euler hacia atrás, pero antes, haremos la deducción de la Ecuación del Problema Térmico completa.
#
# ### Problema térmico
#
# Tomemos nuevamente el ejemplo de la **temperatura** $T$ en [K] de una barra de longitud $L$ en [m] (1D). Pensemos también que tiene una **fuente** de calor $f$ por unidad de longitud (en [J/sm] es decir [W/m]). También tendremos la **tasa de flujo de calor** $q$ medido en [J/sm$^{2}$] (la energía que fluye por unidad de tiempo por unidad de área, es decir, en [W/m$^{2}$]). A continuación un esquema.
#
# [<img src="barra_termica.png" width="400"/>](barra_termica.png)
#
# Pensemos ahora la energía por unidad de longitud $e$, y su derivada $\dot{e}=\frac{de}{dt}$. La ecuación de balance de energía en toda la barra la lograremos integrando tanto $f$, $\dot{e}$, y sumando la energía que entra y la que sale.
#
# $$\int_{0}^{L} \ \dot{e} \ dx =\underbrace{A\left(0\right)q\left(0\right)}_{\text{Energía que entra}}-\underbrace{A\left(L\right)q\left(L\right)}_{\text{Energía que sale}} +\underbrace{\int_{0}^{L} \ f \ dx.}_{\text{Energía generada}}\tag{3}$$
#
# El Teorema fundamental del cálculo nos da la siguiente expresión:
#
# $$\int_{0}^{L} \ \dot{e} \ dx =-\int_{0}^{L}\left(Aq\right)^{'} \ dx +\int_{0}^{L} \ f \ dx\tag{4}$$
#
# Suponemos también que la energía es proporcional a la temperatura $e=C\cdot T$ (donde $C=\rho \cdot c$ es el producto de la densidad por el calor específico). Si sólamente existe flujo de calor por conducción vale: $ q = - k T^{'}$ donde $k$ es la conductividad térmica. Entonces la Ec. (4) queda:
#
# $$\int_{0}^{L} \ c\rho\frac{dT}{dt} \ dx -\int_{0}^{L}\left(AkT^{'}\right)^{'} \ dx = \int_{0}^{L} \ f \ dx\tag{5}$$
#
# Si hacemos tender a cero $L$, ahora si, llegamos a la Ecuación de calor:
#
# $$ \left \{ \begin{array}{l} c\rho\frac{dT}{dt} -\left(AkT^{'}\right)^{'} = f \ \ \text{ para } \ \ x\in I=\left(0,L \right) \text{ es decir en} \Omega \\ \text{condiciones de borde en } \ \partial \Omega \\ \text{y además condiciones iniciales } \ T\left(x,0\right)=T_{0}\left(x\right) \end{array} \tag{6}\right .$$
#
# ### Ejemplo
#
# Ahora resolveremos la Ec. (6) aunque llamaremos $u$ a la variable $T$ y pasamos dividiendo la constante $c\rho$. Nos queda la ecuación:
#
# $$ \left \{ \begin{array}{l} \dot{u} = a\nabla^{2} u + f \ \ \text{ para } \ \ x\in I=\left(0,1 \right) \text{ es decir en} \Omega \\ u\left(0,t\right)= u\left(1,t\right)\text{ en } \ \partial \Omega \text{ (Dirichlet) } \\ u\left(x,0\right)=u_{0}\left(x\right) \text{ (condiciones iniciales) } \ \end{array} \tag{7}\right .$$
#
# Note que aquí también hemos supuesto la sección de la barra constante y la conductividad térmica constante con la posición, por eso aparece la derivada segunda respecto de la posición (que la hemos denotado como $\nabla^{2}$). Note que ahora las condiciones de borde dependen del tiempo y las condiciones iniciales dependen de la posición.
#
# Comenzaremos con la resolución utilizando el Método de Euler hacia atrás (Ec. (1)).
#
# #### Discretización del tiempo
#
# Utilizando las Ecs. (1) y (7) trabajaremos en la discretización primero del tiempo, y luego obtendremos la formulación variacional para discretizar el espacio. Comencemos con algo de notación y la Ec. (7), la ecuación que sigue significa que todas las funciones dentro de los corchetes están evaluadas en el tiempo $t_{n+1}$:
#
# $$ \left[ \dot{u} = a\nabla^{2} u + f \right]^{n+1}\tag{8}$$
#
# Note que en el lado izquierdo de la ecuación tenemos la derivada de $u$ evaluada en el tiempo $t_{n+1}$, $\dot{u}\left(t_{n+1}\right)$. Utilizando la Ec. (1), podemos reemplazar esto por su aproximación dada por el Método de Euler hacia atrás:
#
# $$\frac{u^{n+1}-u^{n}}{\Delta t} \approx a\nabla^{2} u^{n+1} + f^{n+1}$$
#
# entonces podemos despejar:
#
# $$u^{n+1} \approx u^{n}+ \Delta t a\nabla^{2} u^{n+1} + \Delta t f^{n+1} \tag{9}$$
#
# Aclarando un poco, $u_{e}$ es la solución exacta de la Ec. (7) y $u_{e}^{n+1}$ es la solución discretizando el tiempo de la Ec. (9). Es decir, ahora aproximaremos $u_{e}^{n+1}$ por su solución dada por elementos finitos $u^{n+1}$ ($u_{e}^{n+1}\approx u^{n+1}$).
#
# #### Discretización del espacio
#
# Una vez que tenemos discretizado de alguna manera el tiempo, discretizamos el espacio.
#
# $$u_{e}^{n}\approx u^{n} = \sum_{j=0}^{N} \mathbf{c}_{j}^{n}\xi_{j}\left(x\right)$$
# $$u_{e}^{n+1}\approx u^{n+1} = \sum_{j=0}^{N} \mathbf{c}_{j}^{n+1}\xi_{j}\left(x\right)$$
#
# donde $N$ son los grados de libertad de discretización espacial y $N_{t}$ es el número de discretización en el tiempo, $\mathbf{c}$ son constantes y $\xi_{j}$ son, por ejemplo, las funciones sombrero que vimos al comienzo de este curso.
#
# #### Formulación variacional
#
# Tomamos la Ec. (9) y multiplicamos por la función de prueba ($v$) e integramos.
#
# $$\int u^{n+1} v \ dx \approx \int \left( u^{n}+ \Delta t a\nabla^{2} u^{n+1} + \Delta t f^{n+1}\right) v \ dx \tag{10}$$
#
# Suponiendo que comenzamos con $u^{0} = u_{0}\left(x\right)$ (condición inicial), entonces al paso siguiente $u^{1}$ lo obtendremos a partir de la Ec. (10). Observamos en esta ecuación que aparece $\nabla^{2} u^{n+1}$ que, como ya lo hemos hablado, tiene derivada de orden dos, por lo tanto debemos reducir el órden. Utilizamos como siempre la fórmula de Green y (suponiendo condiciones de Dirichlet) se tiene:
#
# $$\int u^{n+1} v \ dx \approx \int u^{n}\cdot v \ dx- \Delta t a \int \nabla u^{n+1} \cdot \nabla v \ dx + \Delta t \int f^{n+1} v \ dx \tag{11}.$$
#
# Todas las integrales son en $\Omega$.
#
# Resolveremos este problema con la solución de un problema teórico para mostrar la presición del método.
#
# ### Código
#
# El código está subido a la carpeta ejemplos y es el archivo *ejemplo14.py*. Es similar al resuelto en el [tutorial de FEniCS](https://fenicsproject.org/pub/tutorial/html/._ftut1006.html#ch:fundamentals:diffusion).
#
# Continuando con el ejemplo anterior, supondremos que $a=1$, y construiremos una función a partir de la Ec. (7), de tal manera que conozcamos la solución exacta. Por ejemplo, supongamos que la solución exacta es:
#
# $$u_{e} = 1+x^{2}+\beta t$$
#
# Calculamos la derivada con respecto al tiempo y da $\frac{du_{e}}{dt}=\beta$. Asimismo, la derivada respecto del espacio da $\frac{\partial^{2}u_{e}}{\partial x^{2}} = 2$, reemplazando este resultado en la Ec. (7) podemos deducir que $f=\beta -2$ y que la condición de Dirichlet en los bordes está dada por $u_{e}\left(0,t\right)=1+\beta t$ y $u_{e}\left(1,t\right)=2+\beta t$. La condición inicial será $u_{0}\left(x\right)=1+x^{2}$.
#
# Entonces podemos comenzar con el código, como siempre, definiendo algunos parámetros (tiempo total de simulación, pasos de simulación, número de intervalos de la malla, etc.), creando la malla y el espacio de la función de prueba ($v$).
# +
from __future__ import print_function
from fenics import *
import numpy as np
T = 10.0 # tiempo final
num_steps = 40 # número de pasos
dt = T / num_steps # paso de tiempo
nx = 20 #numero de intervalos
minx, maxx = 0.0, 1.0
mesh = IntervalMesh(nx, minx, maxx)#malla en 1D
V = FunctionSpace(mesh, 'P',1)#Lagrange Finite Element
# -
# Luego definiremos la expresión de la solución exacta (que conocemos de antemano, porque la creamos así). Esto me permitirá calcular varias cosas: las condiciones de borde y la condición inicial. Además creándola de esta forma, podremos actualizar su valor fácilmente a medida que evolucione el tiempo.
beta = 1.0
u_D = Expression('1+x[0]*x[0]+beta*t', degree=1, beta=beta, t=0)
# Hemos dado el valor de $\beta=1$ (sin perder generalidad). Definimos las condiciones de borde y las condiciones iniciales.
# +
def boundary(x, on_boundary):
return on_boundary
bc = DirichletBC(V, u_D, boundary)#Condición de borde
u_n = interpolate(u_D, V)#Condición inicial
# -
# Luego, la función ensayo, la de prueba, y la función $f$ (de acuerdo al razonamiento anterior tiene que valer: $f = \beta -2$)
u = TrialFunction(V)
v = TestFunction(V)
f = Constant(beta-2)
# Utilizando la Ec. (11) escribimos la formulación variacional de la forma $F=0$:
#
# $$F=\int u^{n+1} v \ dx - \int u^{n}\cdot v \ dx + \Delta t a \int \nabla u^{n+1} \cdot \nabla v \ dx - \Delta t \int f^{n+1} v \ dx = 0 \tag{12}.$$
#
# Esto en FEniCS lo escribimos casi textualmente:
F = u*v*dx - u_n*v*dx + dt*dot(grad(u), grad(v))*dx - dt*f*v*dx
a, L = lhs(F), rhs(F)
# Hemos reemplazado $u^{n+1}$ por u y $u^{n}$ por u_n.
#
# Ahora tenemos que resolverlo para cada paso de tiempo. Lo hacemos con el siguiente for:
# +
u = Function(V)
u_inter = []
t = 0.0
for nn in range(num_steps):
# actualiza el tiempo
t += dt
u_D.t = t
# calcula la solución
solve(a == L, u, bc)
# Temperatura a la mitad de la barra
u_inter.append(u(0.5))
#Calcula errores
u_exacta = interpolate(u_D,V)
error = np.abs(u_exacta.vector()-u.vector()).max()
print('t= %.2f: error = %.3g' %(t,error))
u_n.assign(u)
# -
# En cada paso, hemos impreso el error en el cálculo. Luego, podemos dibujar la solución.
# +
print('Tipo de variable:',type(u))
import matplotlib.pyplot as plt
import numpy as np
#Extraigo los datos de la solucion u.
uh = u.compute_vertex_values(mesh)
print('Cantidad de celdas:',nx)
print('Cantidad de vertices:',len(uh))
xu = np.linspace(0.0, 1.0, len(uh),endpoint = True)
plt.subplot(2,1,1)
plt.plot(xu,uh,'ro',markersize=10)
##Comparo con solucion exacta
xe = np.arange(0.0,1.0,0.001)
ue = 1.0+xe**2.0+beta*T
plt.plot(xe,ue,'b')
plt.ylabel('Solución en t = 10 s')
plt.xlabel('x')
plt.subplot(2,1,2)
tiempo = np.arange(0.0,T,dt)
plt.plot(tiempo,u_inter,'b')
plt.ylabel('Solución en x = 0.5')
plt.xlabel('t (s)')
plt.show()
# -
# ### Ahora con Euler hacia adelante
#
# Veamos el mismo ejemplo pero ahora con el enfoque Euler hacia adelante, tomamos la Ec. (8) pero ahora en lugar de evaluarla en el tiempo $n+1$ lo haremos en el tiempo $n$:
#
# $$ \left[ \dot{u} = a\nabla^{2} u + f \right]^{n}.$$
#
# Entonces, ahora la derivada de $u$ estará evaluada en el tiempo $t_{n}$, $\dot{u}\left(t_{n}\right)$. Utilizando la Ec. (2), podemos reemplazar esto por su aproximación dada por el Método de Euler hacia adelante:
#
# $$\frac{u^{n+1}-u^{n}}{\Delta t} \approx a\nabla^{2} u^{n} + f^{n}.$$
#
# Realizando el mismo procedimiento, podemos encontrar la formulación variacional y llegaremos a la siguiente expresión:
#
# $$\int u^{n+1} v \ dx - \int u^{n}\cdot v \ dx + \Delta t a \int \nabla u^{n} \cdot \nabla v \ dx - \Delta t \int f^{n} v \ dx = 0.$$
#
# El código será similar, está implementado en el *ejemplo15.py*. Recordemos que el método Euler hacia adelante es condicionalmente esable, por consiguiente, el paso de tiempo que se debe usar será mucho más pequeño (probar con $N_{t}=$ 40, $N_{t}=$ 400 y $N_{t}=$ 4000).
#
# ## Conclusiones
#
# - Hemos introducido la resolución de problemas con funciones variables en el tiempo utilizando elementos finitos.
# - Hemos resuelto la ecuación de calor, la cual es un tipo de ecuación que se denomina [ecuación parabólica](https://en.wikipedia.org/wiki/Parabolic_partial_differential_equation) utilizando dos enfoques diferentes para aproximar la derivada temporal aproximación: Euler hacia adelante (explícito) y hacia atrás (implícito).
# - El método de Euler explícito es condicionalmente estable, hemos visto que el paso de tiempo debe ser mucho más pequeño para obtener la solución. Algunos detalles más sobre este problema se pueden estudiar en el siguiente [enlace](https://hal.archives-ouvertes.fr/hal-01401125/document).
#
| Problemas_dependientes_del_tiempo/Introduccion_problema_termico.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %config IPython.matplotlib.backend = "retina"
import matplotlib.pyplot as plt
import numpy as np
# +
from toolkit import trappist1, transit_model, trappist_out_of_transit, trappist1_all_transits
from copy import deepcopy
g = trappist1('g')
original_g = deepcopy(g)
# -
from glob import glob
lc_paths = sorted(glob('all_transits/phot*'))
midtransit_times = len(lc_paths) * [g.t0]
# +
original_t0 = deepcopy(g.t0)
all_times = []
midtransit_time_offsets = []
for i, path, midtransit_time in zip(np.arange(len(lc_paths)), lc_paths, midtransit_times):
for j in range(2):
bjd, flux, err, x, y, fwhm, fwhmx, fwhmy, bg, airmass, exptime = np.loadtxt(path, unpack=True)
all_times.append(bjd)
bjd += 2450000
if j == 1:
g.t0 = midtransit_times[i]
transit_model_g = transit_model(bjd, g)
all_transits = trappist1_all_transits(bjd)
oot = all_transits == 1
X_all = np.vstack([x, y, fwhmx, fwhmy]).T
X = X_all[oot, :]
omega = np.diag(err[oot]**2)
omega_inv = np.linalg.inv(omega)
V = np.linalg.inv(X.T @ omega_inv @ X)
beta = V @ X.T @ omega_inv @ flux[oot]
regressed_lc = flux - (X_all @ beta) + 1
regressed_lc /= np.median(regressed_lc[oot])
from scipy.optimize import fmin_powell, fmin_l_bfgs_b
def minimize_this(p):
return abs(np.sum((regressed_lc[oot] - transit_model_g[oot])**2 /
(p[0] * err[oot])**2)/len(regressed_lc[oot]) - 1)
err_scale = fmin_powell(minimize_this, [1], disp=False)
err *= err_scale
def transit_model_t(p):
params = deepcopy(g)
params.t0 = p[0]
return transit_model(bjd, params)
def chi2(p):
return np.sum((regressed_lc - transit_model_t(p))**2 / err**2)
#result = fmin_powell(chi2, [bjd[np.argmin(regressed_lc)]], disp=False)
#result = fmin_powell(chi2, [midtransit_time], disp=False)
result = fmin_l_bfgs_b(chi2, [bjd[np.argmin(regressed_lc)]], approx_grad=True,
bounds=[(bjd.min(), bjd.max())])[0]
if j == 0:
midtransit_times[i] = result[0]
n_transits = np.round((bjd.mean() - original_g.t0) / g.per)
midtransit_time_offset = midtransit_times[i] - (original_g.t0 + n_transits*g.per)
print(midtransit_time_offset)
midtransit_time_offsets.append(midtransit_time_offset)
plt.errorbar(bjd - midtransit_time_offset, regressed_lc, err, fmt='.')
np.savetxt('reduced_lcs/lightcurve_{}.txt'.format(i), np.vstack([bjd - midtransit_time_offset, regressed_lc, err]).T)
plt.plot(bjd - midtransit_time_offset, transit_model_t(result), 'r')
plt.plot(bjd, transit_model(bjd, original_g), 'm')
# plt.plot(bjd, all_transits)
plt.title(note)
plt.show()
# -
all_lcs = np.vstack(sorted([np.loadtxt(i) for i in glob('reduced_lcs/lightcurve_?.txt')], key=lambda x: x[0][0]))
np.savetxt('reduced_lcs/all_lightcurves.txt', all_lcs)
all_lcs[:, 0].min(), all_lcs[:, 0].ptp(),
# +
import os
import shutil
from subprocess import Popen
lc_paths = glob('reduced_lcs/lightcurve_?.txt')
for i in range(len(lc_paths)):
new_dir = "stsp_{0:d}".format(i)
if not os.path.exists(new_dir):
os.mkdir(new_dir)
shutil.copy('stsp/g.in', os.path.join(new_dir, '.'))
shutil.copy('stsp/stsp_20180302', os.path.join(new_dir, '.'))
shutil.copy(lc_paths[i], os.path.join(new_dir, '.'))
bjd, flux, err = np.loadtxt(lc_paths[i], unpack=True)
original_infile = open(os.path.join(new_dir, 'g.in')).read()
new_infile = original_infile.replace('lightcurve.txt', os.path.basename(lc_paths[i])) # Replace start time
new_infile = new_infile.replace('2457665.2007372407', str(bjd.min())) # Replace start time
new_infile = new_infile.replace('0.21868119994178414', str(bjd.ptp())) # Replace duration
with open(os.path.join(new_dir, 'g.in'), 'w') as w:
w.write(new_infile)
new_infile = new_infile.replace('1\t\t\t\t; number of spots', '0\t\t\t\t; number of spots')
new_infile = '\n'.join(new_infile.splitlines()[:-6] + ['l', '1'])
with open(os.path.join(new_dir, 'g_nospot.in'), 'w') as w:
w.write(new_infile)
#Popen(['./stsp_20180302', 'g.in'], cwd=os.path.abspath(new_dir))
Popen(['./stsp_20180302', 'g_nospot.in'], cwd=os.path.abspath(new_dir))
# +
for i, toffset in enumerate(midtransit_time_offsets):
new_dir = "stsp_{0:d}".format(i)
bjd, flux, err, model, spotinds = np.loadtxt(os.path.join(new_dir, 'g_lcbest.txt'), unpack=True)
bjd_model, _, _, transit_model_g, _ = np.loadtxt(os.path.join(new_dir, 'g_nospot_lcout.txt'), unpack=True)
bjd_int = int(bjd.min())
fig, ax = plt.subplots(1, 2, figsize=(8.5, 4))
ax[0].errorbar(bjd - bjd_int, flux, err, fmt='.', color='k', ecolor='silver', label='Spitzer')
ax[0].plot(bjd_model - bjd_int, transit_model_g, color='k', label='Unspotted')
ax[0].plot(bjd - bjd_int, model, color='r', lw=2, label='STSP')
ax[0].set(xlabel='BJD - {}'.format(bjd_int), ylabel='Flux')
ax[0].legend(loc='lower left')
mcmc = np.loadtxt(os.path.join(new_dir, 'g_mcmc.txt'))
n = 10000
radius = mcmc[-n:, 4]
theta = mcmc[-n:, 5]
phi = mcmc[-n:, 6]
ax[1].hist(radius)
ax[1].set_xlabel('Spot radius')
# ax[1].errorbar(bjd - bjd_int, flux - transit_model_g, err, fmt='.', color='k', ecolor='silver', label='Spitzer')
# ax[1].plot(bjd - bjd_int, model - transit_model_g, lw=2, color='r', label='STSP')
# ax[1].set(xlabel='BJD - {}'.format(bjd_int), ylabel='Residuals')
# ax[1].legend(loc='upper left')
fig.tight_layout()
for axis in ax:
for i in ['right', 'top']:
axis.spines[i].set_visible(False)
fig.savefig('stsp_model_{0}.pdf'.format(i), bbox_inches='tight', dpi=200)
# +
fig, ax = plt.subplots(2, 2, figsize=(10, 5), sharey='row', sharex='col')
dirs = ['stsp_1', 'stsp_4']
for i, new_dir in enumerate(dirs):
bjd, flux, err, model, spotinds = np.loadtxt(os.path.join(new_dir, 'g_lcbest.txt'), unpack=True)
bjd_model, _, _, transit_model_g, _ = np.loadtxt(os.path.join(new_dir, 'g_nospot_lcout.txt'), unpack=True)
bjd_int = int(bjd.min())
ax[0, i].errorbar(bjd - bjd_int, flux, err, fmt='.', color='k', ecolor='silver', label='Spitzer')
ax[0, i].plot(bjd_model - bjd_int, transit_model_g, color='k', label='Expected')
ax[0, i].plot(bjd - bjd_int, model, color='r', lw=2, label='STSP')
ax[1, i].errorbar(bjd - bjd_int, flux - transit_model_g, err, fmt='.', color='k', ecolor='silver', label='Spitzer')
ax[1, i].plot(bjd - bjd_int, model - transit_model_g, color='r', lw=2, label='STSP')
ax[1, i].set(xlabel='BJD - {}'.format(bjd_int))
for axis in fig.axes:
for j in ['top', 'right']:
axis.spines[j].set_visible(False)
axis.grid(ls=':')
ax[0, 0].set_ylabel('Flux')
ax[1, 0].set_ylabel('Residual')
ax[0, 0].legend(loc='lower left')
fig.tight_layout()
fig.savefig('spot_occultations.pdf', bbox_inches='tight')
# -
| lightcurves.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# !pip install torchsummary
# +
import cv2
import pandas.io.clipboard as clipboard
from PIL import ImageGrab
from PIL import Image
import os
import sys
import argparse
import logging
import yaml
import re
import numpy as np
import torch
from torchvision import transforms
from munch import Munch
from transformers import PreTrainedTokenizerFast
from timm.models.resnetv2 import ResNetV2
from timm.models.layers import StdConv2dSame
from dataset.dataset import test_transform
from dataset.latex2png import tex2pil
from models import get_model
from utils import *
last_pic = None
# -
import os
os.chdir("/home/lap14784/Downloads/LaTeX_OCR")
os.getcwd()
# ## Define model
# +
import torch
import torch.nn as nn
import torch.nn.functional as F
# from x_transformers import *
from x_transformers import TransformerWrapper, Decoder
from x_transformers.autoregressive_wrapper import AutoregressiveWrapper, top_k, top_p, entmax, ENTMAX_ALPHA
from timm.models.vision_transformer import VisionTransformer
from timm.models.vision_transformer_hybrid import HybridEmbed
from timm.models.resnetv2 import ResNetV2
from timm.models.layers import StdConv2dSame
from einops import rearrange, repeat
class CustomARWrapper(AutoregressiveWrapper):
def __init__(self, *args, **kwargs):
super(CustomARWrapper, self).__init__(*args, **kwargs)
@torch.no_grad()
def generate(self, start_tokens, seq_len, eos_token=None, temperature=1., filter_logits_fn=top_k, filter_thres=0.9, **kwargs):
device = start_tokens.device
was_training = self.net.training
num_dims = len(start_tokens.shape)
if num_dims == 1:
start_tokens = start_tokens[None, :]
b, t = start_tokens.shape
self.net.eval()
out = start_tokens
mask = kwargs.pop('mask', None)
if mask is None:
mask = torch.full_like(out, True, dtype=torch.bool, device=out.device)
for _ in range(seq_len):
x = out[:, -self.max_seq_len:]
mask = mask[:, -self.max_seq_len:]
# print('arw:',out.shape)
logits = self.net(x, mask=mask, **kwargs)[:, -1, :]
if filter_logits_fn in {top_k, top_p}:
filtered_logits = filter_logits_fn(logits, thres=filter_thres)
probs = F.softmax(filtered_logits / temperature, dim=-1)
elif filter_logits_fn is entmax:
probs = entmax(logits / temperature, alpha=ENTMAX_ALPHA, dim=-1)
sample = torch.multinomial(probs, 1)
out = torch.cat((out, sample), dim=-1)
mask = F.pad(mask, (0, 1), value=True)
if eos_token is not None and (torch.cumsum(out == eos_token, 1)[:, -1] >= 1).all():
break
out = out[:, t:]
if num_dims == 1:
out = out.squeeze(0)
self.net.train(was_training)
return out
class CustomVisionTransformer(VisionTransformer):
def __init__(self, img_size=224, patch_size=16, *args, **kwargs):
super(CustomVisionTransformer, self).__init__(img_size=img_size, patch_size=patch_size, *args, **kwargs)
self.height, self.width = img_size
self.patch_size = patch_size
def forward_features(self, x):
print(np.shape(x))
B, c, h, w = x.shape
x = self.patch_embed(x)
cls_tokens = self.cls_token.expand(B, -1, -1) # stole cls_tokens impl from <NAME>, thanks
x = torch.cat((cls_tokens, x), dim=1)
h, w = h//self.patch_size, w//self.patch_size
pos_emb_ind = repeat(torch.arange(h)*(self.width//self.patch_size-w), 'h -> (h w)', w=w)+torch.arange(h*w)
pos_emb_ind = torch.cat((torch.zeros(1), pos_emb_ind+1), dim=0).long()
x += self.pos_embed[:, pos_emb_ind]
#x = x + self.pos_embed
x = self.pos_drop(x)
for blk in self.blocks:
x = blk(x)
x = self.norm(x)
return x
class Model(nn.Module):
def __init__(self, encoder: CustomVisionTransformer, decoder: CustomARWrapper, args, temp: float = .333):
super().__init__()
self.encoder = encoder
self.decoder = decoder
self.bos_token = args.bos_token
self.eos_token = args.eos_token
self.max_seq_len = args.max_seq_len
self.temperature = temp
@torch.no_grad()
def forward(self, x: torch.Tensor):
print("forward", x)
device = x.device
encoded = self.encoder(x.to(device))
dec = self.decoder.generate(torch.LongTensor([self.bos_token]*len(x))[:, None].to(device), self.max_seq_len,
eos_token=self.eos_token, context=encoded, temperature=self.temperature)
return dec
def get_model(args, training=False):
backbone = ResNetV2(
layers=args.backbone_layers, num_classes=0, global_pool='', in_chans=args.channels,
preact=False, stem_type='same', conv_layer=StdConv2dSame)
min_patch_size = 2**(len(args.backbone_layers)+1)
def embed_layer(**x):
ps = x.pop('patch_size', min_patch_size)
assert ps % min_patch_size == 0 and ps >= min_patch_size, 'patch_size needs to be multiple of %i with current backbone configuration' % min_patch_size
return HybridEmbed(**x, patch_size=ps//min_patch_size, backbone=backbone)
encoder = CustomVisionTransformer(img_size=(args.max_height, args.max_width),
patch_size=args.patch_size,
in_chans=args.channels,
num_classes=0,
embed_dim=args.dim,
depth=args.encoder_depth,
num_heads=args.heads,
embed_layer=embed_layer
).to(args.device)
decoder = CustomARWrapper(
TransformerWrapper(
num_tokens=args.num_tokens,
max_seq_len=args.max_seq_len,
attn_layers=Decoder(
dim=args.dim,
depth=args.num_layers,
heads=args.heads,
**args.decoder_args
)),
pad_value=args.pad_token
).to(args.device)
model = Model(encoder, decoder, args)
# if training:
# # check if largest batch can be handled by system
# im = torch.empty(args.batchsize, args.channels, args.max_height, args.min_height, device=args.device).float()
# seq = torch.randint(0, args.num_tokens, (args.batchsize, args.max_seq_len), device=args.device).long()
# decoder(seq, context=encoder(im)).sum().backward()
# model.zero_grad()
# torch.cuda.empty_cache()
# del im, seq
return model
# -
# +
from torchsummary import summary
from dataset.dataset import test_transform
import cv2
import pandas.io.clipboard as clipboard
from PIL import ImageGrab
from PIL import Image
import os
import sys
import argparse
import logging
import yaml
import re
import numpy as np
import torch
from torchvision import transforms
from munch import Munch
from transformers import PreTrainedTokenizerFast
from timm.models.resnetv2 import ResNetV2
from timm.models.layers import StdConv2dSame
from dataset.latex2png import tex2pil
from models import get_model
from utils import *
last_pic = None
# +
# if arguments is None:
# arguments = Munch({'config': 'settings/config.yaml', 'checkpoint': 'checkpoints/weights.pth', 'no_cuda': True, 'no_resize': False})
arguments = Munch({'epoch': 0, 'backbone_layers': [2, 3, 7], 'betas': [0.9, 0.999], 'batchsize': 10, 'bos_token': 1, 'channels': 1, 'data': 'dataset/data/train.pkl', 'debug': False, 'decoder_args': {'attn_on_attn': True, 'cross_attend': True, 'ff_glu': True, 'rel_pos_bias': False, 'use_scalenorm': False}, 'dim': 256, 'encoder_depth': 4, 'eos_token': 2, 'epochs': 10, 'gamma': 0.9995, 'heads': 8, 'id': None, 'load_chkpt': None, 'lr': 0.001, 'lr_step': 30, 'max_height': 192, 'max_seq_len': 512, 'max_width': 672, 'min_height': 32, 'min_width': 32, 'model_path': 'checkpoints', 'name': 'pix2tex', 'num_layers': 4, 'num_tokens': 8000, 'optimizer': 'Adam', 'output_path': 'outputs', 'pad': False, 'pad_token': 0, 'patch_size': 16, 'sample_freq': 3000, 'save_freq': 5, 'scheduler': 'StepLR', 'seed': 42, 'temperature': 0.2, 'test_samples': 5, 'testbatchsize': 20, 'tokenizer': 'dataset/tokenizer.json', 'valbatches': 100, 'valdata': 'dataset/data/val.pkl', 'wandb': False, 'device': 'cpu', 'max_dimensions': [672, 192], 'min_dimensions': [32, 32], 'out_path': 'checkpoints/pix2tex', 'config': 'settings/config.yaml', 'checkpoint': 'checkpoints/weights.pth', 'no_cuda': False, 'no_resize': False})
# logging.getLogger().setLevel(logging.FATAL)
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
with open(arguments.config, 'r') as f:
params = yaml.load(f, Loader=yaml.FullLoader)
args = parse_args(Munch(params))
args.update(**vars(arguments))
# args.device = "cpu"
args.device = 'cuda' if torch.cuda.is_available() and not args.no_cuda else 'cpu'
model = get_model(args)
# summary(model, (1, 1, 64, 352))
# model.load_state_dict(torch.load(args.checkpoint, map_location=args.device))
# summary(model, (1, 1, 64, 352))
# if 'image_resizer.pth' in os.listdir(os.path.dirname(args.checkpoint)) and not arguments.no_resize:
# image_resizer = ResNetV2(layers=[2, 3, 3], num_classes=max(args.max_dimensions)//32, global_pool='avg', in_chans=1, drop_rate=.05,
# preact=True, stem_type='same', conv_layer=StdConv2dSame).to(args.device)
# image_resizer.load_state_dict(torch.load(os.path.join(os.path.dirname(args.checkpoint), 'image_resizer.pth'), map_location=args.device))
# image_resizer.eval()
# else:
# image_resizer = None
# tokenizer = PreTrainedTokenizerFast(tokenizer_file=args.tokenizer)
# return args, model, image_resizer, tokenizer
# -
from PIL import Image
img = Image.open("./dataset/sample/1000a29807.png")
torch.Size([1, 1, 64, 352])
# +
encoder, decoder = model.encoder, model.decoder
if type(img) is bool:
img = None
if img is None:
if last_pic is None:
print('Provide an image.')
else:
img = last_pic.copy()
else:
last_pic = img.copy()
img = minmax_size(pad(img), args.max_dimensions, args.min_dimensions)
if image_resizer is not None and not args.no_resize:
with torch.no_grad():
input_image = img.convert('RGB').copy()
r, w, h = 1, input_image.size[0], input_image.size[1]
for _ in range(10):
h = int(h * r) # height to resize
img = pad(minmax_size(input_image.resize((w, h), Image.BILINEAR if r > 1 else Image.LANCZOS), args.max_dimensions, args.min_dimensions))
t = test_transform(image=np.array(img.convert('RGB')))['image'][:1].unsqueeze(0)
w = (image_resizer(t.to(args.device)).argmax(-1).item()+1)*32
logging.info(r, img.size, (w, int(input_image.size[1]*r)))
if (w == img.size[0]):
break
r = w/img.size[0]
else:
img = np.array(pad(img).convert('RGB'))
t = test_transform(image=img)['image'][:1].unsqueeze(0)
im = t.to(args.device)
with torch.no_grad():
model.eval()
device = args.device
encoded = encoder(im.to(device))
dec = decoder.generate(torch.LongTensor([args.bos_token])[:, None].to(device), args.max_seq_len,
eos_token=args.eos_token, context=encoded.detach(), temperature=args.get('temperature', .25))
pred = post_process(token2str(dec, tokenizer)[0])
try:
clipboard.copy(pred)
except:
pass
# -
pred
prediction = pred.replace('<', '\\lt ').replace('>', '\\gt ')
prediction
html = str('\\left\\{\\begin{array}{r c l}{{\\delta_{\\epsilon}B}}&{{\\sim}}&{{\\epsilon F\\,,}}\\\\ {{\\delta_{\\epsilon}F}}&{{\\sim}}&{{\\partial\\epsilon+\\epsilon B\\,,}}\\end{array}\\right.')
html
from bs4 import BeautifulSoup
soup = BeautifulSoup(html)
print(soup.get_text())
pageSource = """
<html>
<head><script id="MathJax-script" src="qrc:MathJax.js"></script>
<script>
MathJax.Hub.Config({messageStyle: 'none',tex2jax: {preview: 'none'}});
MathJax.Hub.Queue(
function () {
document.getElementById("equation").style.visibility = "";
}
);
</script>
</head> """ + """
<body>
<div id="equation" style="font-size:1em; visibility:hidden">$${equation}$$</div>
</body>
</html>
""".format(equation=prediction)
from IPython.core.display import display, HTML
display(HTML(pageSource))
| Colab.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + active=""
# 本ページの ipython notebook は :download:`ここ <newmap.ipynb>` からダウンロードできます
# -
# ## 新しい写像を定義し量子論の計算
#
# 古典写像系で示した様SimpleQmapをHarperMapに拡張し,量子論の計算を行う.
# ipython notebookを使わない場合は次の命令は無視して下さい
# %matplotlib inline
# +
import numpy as np
import matplotlib.pyplot as plt
import SimpleQmap as sq
twopi = 2.0*np.pi
class HarperMap(sq.StandardMap):
def __init__(self, k,a):
sq.StandardMap.__init__(self,k)
self.a = a
def func1(self, x):
return a*np.sin(twopi*x)/twopi
def ifunc1(self,x):
return -a*np.cos(twopi*x)/twopi/twopi
def Traj(map,sample=50,tmax=500):
q = np.random.random(sample)
p = (np.random.random(sample) -0.5) * (pmax - pmin) * 2
res = [np.array([])]*2
for i in range(tmax):
pp = p - map.func0(q)
qq = q + map.func1(pp)
q = qq - np.floor(qq)
p = pp - np.floor(pp)
res[0] = np.append(res[0],q)
res[1] = np.append(res[1],p)
return res
dim = 50
k,a = 1,1
qmin, qmax = 0, 1
pmin, pmax = 0, 1
map = HarperMap(k,a)
traj = Traj(map)
domain = [[qmin,qmax],[pmin,pmax]]
qmap = sq.Qmap(map, dim, domain) # defines the quantum system
evals, evecs = qmap.eigen() # return eigenvalues and list of eigenvector of the system.
for i, evec in enumerate(evecs):
fig,axs = plt.subplots(1,2,figsize=(10,5))
#evec.savetxt("data.dat",rep="p") # rep = "q", "p" or "hsm"
theta = np.linspace(-np.pi, np.pi, 100)
z = np.exp(1.j*theta)
axs[0].plot(z.real, z.imag,'-g')
axs[0].plot(evals.real, evals.imag, 'ob')
axs[0].plot(evals[i].real, evals[i].imag, 'or', markersize=10)
x,y,z = evec.hsmrep(row=100,col=100)
axs[1].contour(x,y,z,100)
axs[1].plot(traj[0],traj[1],',k')
fig.suptitle("%d-th eigenstate" % i)
plt.show()
break
# -
| docs/tutorial/newmap.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="LL3kRdXs5zzD"
# ##### Copyright 2018 Google LLC.
#
# Licensed under the Apache License, Version 2.0 (the "License");
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="sAgUgR5Mzzz2"
# # XLA in Python
#
# <img style="height:100px;" src="https://raw.githubusercontent.com/tensorflow/tensorflow/master/tensorflow/compiler/xla/g3doc/images/xlalogo.png"> <img style="height:100px;" src="https://upload.wikimedia.org/wikipedia/commons/c/c3/Python-logo-notext.svg">
#
# _<NAME>_
#
# _The Python XLA client was designed by Roy Frostig._
#
# _JAX was written by <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>._
#
# XLA is the compiler that JAX uses, and the compiler that TF uses for TPUs and will soon use for all devices, so it's worth some study. However, it's not exactly easy to play with XLA computations directly using the raw C++ interface. JAX exposes the underlying XLA computation builder API through a python wrapper, and makes interacting with the XLA compute model accessible for messing around and prototyping.
#
# XLA computations are built as computation graphs in HLO IR, which is then lowered to LLO that is device specific (CPU, GPU, TPU, etc.).
#
# As end users we interact with the computational primitives offered to us by the HLO spec.
# + [markdown] colab_type="text" id="EZK5RseuvZkr"
# ## References
#
# __xla__: the doc that defines what's in HLO - but note that the doc is incomplete and omits some ops.
#
# https://www.tensorflow.org/xla/operation_semantics
#
# more details on ops in the source code.
#
# https://github.com/tensorflow/tensorflow/blob/master/tensorflow/compiler/xla/client/xla_builder.h
#
# __python xla client__: this is the XLA python client for JAX, and what we're using here.
#
# https://github.com/tensorflow/tensorflow/blob/master/tensorflow/compiler/xla/python/xla_client.py
#
# https://github.com/tensorflow/tensorflow/blob/master/tensorflow/compiler/xla/python/xla_client_test.py
#
# __jax__: you can see how jax interacts with the XLA compute layer for execution and JITing in these files.
#
# https://github.com/google/jax/blob/master/jax/lax.py
#
# https://github.com/google/jax/blob/master/jax/lib/xla_bridge.py
#
# https://github.com/google/jax/blob/master/jax/interpreters/xla.py
# + [markdown] colab_type="text" id="3XR2NGmrzBGe"
# ## Colab Setup and Imports
# + [markdown] colab_type="text" id="HMRkxnna8NCN"
# First install jax and jaxlib to get its xla client:
# + colab={} colab_type="code" id="JWCCBdpL8T5t"
# !pip install --upgrade -q https://storage.googleapis.com/jax-wheels/cuda$(echo $CUDA_VERSION | sed -e 's/\.//' -e 's/\..*//')/jaxlib-0.1.21-cp36-none-linux_x86_64.whl
# !pip install --upgrade -q jax
# + colab={} colab_type="code" id="Ogo2SBd3u18P"
# We import as onp to emphasize that we're using vanilla numpy, not jax numpy.
import numpy as onp
# We only need to import JAX's xla_client, not all of JAX.
from jaxlib import xla_client
# Plotting
import matplotlib as mpl
from matplotlib import pyplot as plt
from matplotlib import gridspec
from matplotlib import rcParams
rcParams['image.interpolation'] = 'nearest'
rcParams['image.cmap'] = 'viridis'
rcParams['axes.grid'] = False
# + [markdown] colab_type="text" id="0cf7swaobc5l"
# ## Convenience Functions
# + colab={} colab_type="code" id="5I50k0rhbg6W"
# Here we borrow convenience functions from JAX to convert numpy shape/dtypes
# to XLA appropriate shape/dtypes
def canonicalize_dtype(dtype):
"""We restrict ourselves to 32bit types for this demo."""
_dtype_to_32bit_dtype = {
str(onp.dtype('int64')): onp.dtype('int32'),
str(onp.dtype('uint64')): onp.dtype('uint32'),
str(onp.dtype('float64')): onp.dtype('float32'),
str(onp.dtype('complex128')): onp.dtype('complex64'),
}
dtype = onp.dtype(dtype)
return _dtype_to_32bit_dtype.get(str(dtype), dtype)
def shape_of(value):
"""Given a Python or XLA value, return its canonicalized XLA Shape."""
if hasattr(value, 'shape') and hasattr(value, 'dtype'):
return xla_client.Shape.array_shape(canonicalize_dtype(value.dtype),
value.shape)
elif onp.isscalar(value):
return shape_of(onp.asarray(value))
elif isinstance(value, (tuple, list)):
return xla_client.Shape.tuple_shape(tuple(shape_of(elt) for elt in value))
else:
raise TypeError('Unexpected type: {}'.format(type(value)))
def to_xla_type(dtype):
"Convert to integert xla type, for use with ConvertElementType, etc."
if isinstance(dtype, str):
return xla_client.DTYPE_TO_XLA_ELEMENT_TYPE[dtype]
elif isinstance(dtype, type):
return xla_client.DTYPE_TO_XLA_ELEMENT_TYPE[onp.dtype(dtype).name]
elif isinstance(dtype, onp.dtype):
return xla_client.DTYPE_TO_XLA_ELEMENT_TYPE[dtype.name]
else:
raise TypeError('Unexpected type: {}'.format(type(dtype)))
# + [markdown] colab_type="text" id="odmjXyhMuNJ5"
# ## Simple Computations
# + colab={"height": 33} colab_type="code" executionInfo={"elapsed": 364, "status": "ok", "timestamp": 1549929562036, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-rqjtQZ4KjgQ/AAAAAAAAAAI/AAAAAAAAAAw/5BQt0zmTW5o/s64/photo.jpg", "userId": "09409386770882740563"}, "user_tz": 480} id="UYUtxVzMYIiv" outputId="bd8aa18e-26d9-4df4-ebc3-20026119de17"
# make a computation builder
c = xla_client.ComputationBuilder("simple_scalar")
# define a parameter shape and parameter
param_shape = xla_client.Shape.array_shape(onp.dtype(onp.float32), ())
x = c.ParameterWithShape(param_shape)
# define computation graph
y = c.Sin(x)
# build computation graph
# Keep in mind that incorrectly constructed graphs can cause
# your notebook kernel to crash!
computation = c.Build()
# compile graph based on shape
compiled_computation = computation.Compile([param_shape,])
# define a host variable with above parameter shape
host_input = onp.array(3.0, dtype=onp.float32)
# place host variable on device and execute
device_input = xla_client.LocalBuffer.from_pyval(host_input)
device_out = compiled_computation.Execute([device_input ,])
# retrive the result
device_out.to_py()
# + colab={"height": 33} colab_type="code" executionInfo={"elapsed": 350, "status": "ok", "timestamp": 1549929568548, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-rqjtQZ4KjgQ/AAAAAAAAAAI/AAAAAAAAAAw/5BQt0zmTW5o/s64/photo.jpg", "userId": "09409386770882740563"}, "user_tz": 480} id="rIA-IVMVvQs2" outputId="ce88ec6f-d2ea-4ec2-80b4-ddd1afd36957"
# same as above with vector type:
c = xla_client.ComputationBuilder("simple_vector")
param_shape = xla_client.Shape.array_shape(onp.dtype(onp.float32), (3,))
x = c.ParameterWithShape(param_shape)
# can also use this function to define a shape from an example:
#x = c.ParameterFromNumpy(onp.array([0.0, 0.0, 0.0], dtype=onp.float32))
# which is the same as using our convenience function above:
#x = c.ParameterWithShape(shape_of(onp.array([0.0, 0.0, 0.0],
# dtype=onp.float32)))
# chain steps by reference:
y = c.Sin(x)
z = c.Abs(y)
computation = c.Build()
compiled_computation = computation.Compile([param_shape,])
host_input = onp.array([3.0, 4.0, 5.0], dtype=onp.float32)
device_input = xla_client.LocalBuffer.from_pyval(host_input)
device_out = compiled_computation.Execute([device_input ,])
# retrive the result
device_out.to_py()
# + [markdown] colab_type="text" id="F8kWlLaVuQ1b"
# ## Simple While Loop
# + colab={"height": 33} colab_type="code" executionInfo={"elapsed": 358, "status": "ok", "timestamp": 1549929569852, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-rqjtQZ4KjgQ/AAAAAAAAAAI/AAAAAAAAAAw/5BQt0zmTW5o/s64/photo.jpg", "userId": "09409386770882740563"}, "user_tz": 480} id="MDQP1qW515Ao" outputId="4da894b5-2b0e-455e-a720-3bdadc57d164"
# trivial while loop, decrement until 0
# x = 5
# while x > 0:
# x = x - 1
#
in_shape = shape_of(5)
# body computation:
bcb = xla_client.ComputationBuilder("bodycomp")
x = bcb.ParameterWithShape(in_shape)
const1 = bcb.Constant(onp.int32(1))
y = bcb.Sub(x, const1)
body_computation = bcb.Build()
# test computation:
tcb = xla_client.ComputationBuilder("testcomp")
x = tcb.ParameterWithShape(in_shape)
const0 = tcb.Constant(onp.int32(0))
y = tcb.Gt(x, const0)
test_computation = tcb.Build()
# while computation:
wcb = xla_client.ComputationBuilder("whilecomp")
x = wcb.ParameterWithShape(in_shape)
wcb.While(test_computation, body_computation, x)
while_computation = wcb.Build()
# Now compile and execute:
compiled_computation = while_computation.Compile([in_shape,])
host_input = onp.array(5, dtype=onp.int32)
device_input = xla_client.LocalBuffer.from_pyval(host_input)
device_out = compiled_computation.Execute([device_input ,])
# retrive the result
device_out.to_py()
# + [markdown] colab_type="text" id="7UOnXlY8slI6"
# ## While loops w. tuples - Newton's Method for sqrt
# + colab={"height": 33} colab_type="code" executionInfo={"elapsed": 402, "status": "ok", "timestamp": 1549929572085, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-rqjtQZ4KjgQ/AAAAAAAAAAI/AAAAAAAAAAw/5BQt0zmTW5o/s64/photo.jpg", "userId": "09409386770882740563"}, "user_tz": 480} id="HEWz-vzd6QPR" outputId="6ef10855-232d-4701-a442-0e2667b2fd97"
Xsqr = 2
guess = 1.0
converged_delta = 0.001
maxit = 1000
in_shape = shape_of((1.0, 1.0, 1))
# body computation:
# x_{i+1} = x_{i} - (x_i**2 - y) / (2 * x_i)
bcb = xla_client.ComputationBuilder("bodycomp")
intuple = bcb.ParameterWithShape(in_shape)
y = bcb.GetTupleElement(intuple, 0)
x = bcb.GetTupleElement(intuple, 1)
guard_cntr = bcb.GetTupleElement(intuple, 2)
new_x = bcb.Sub(x, bcb.Div(bcb.Sub(bcb.Mul(x, x), y), bcb.Add(x, x)))
result = bcb.Tuple(y, new_x, bcb.Sub(guard_cntr, bcb.Constant(onp.int32(1))))
body_computation = bcb.Build()
# test computation -- convergence and max iteration test
tcb = xla_client.ComputationBuilder("testcomp")
intuple = tcb.ParameterWithShape(in_shape)
y = tcb.GetTupleElement(intuple, 0)
x = tcb.GetTupleElement(intuple, 1)
guard_cntr = tcb.GetTupleElement(intuple, 2)
criterion = tcb.Abs(tcb.Sub(tcb.Mul(x, x), y))
# stop at convergence criteria or too many iterations
test = tcb.And(tcb.Gt(criterion, tcb.Constant(onp.float32(converged_delta))),
tcb.Gt(guard_cntr, tcb.Constant(onp.int32(0))))
test_computation = tcb.Build()
# while computation:
wcb = xla_client.ComputationBuilder("whilecomp")
intuple = wcb.ParameterWithShape(in_shape)
wcb.While(test_computation, body_computation, intuple)
while_computation = wcb.Build()
# Now compile and execute:
compiled_computation = while_computation.Compile([in_shape,])
y = onp.array(Xsqr, dtype=onp.float32)
x = onp.array(guess, dtype=onp.float32)
maxit = onp.array(maxit, dtype=onp.int32)
device_input = xla_client.LocalBuffer.from_pyval((y, x, maxit))
device_out = compiled_computation.Execute([device_input ,])
host_out = device_out.to_py()
print("square root of {y} is {x}".format(y=y, x=host_out[1]))
# + [markdown] colab_type="text" id="yETVIzTInFYr"
# ## Calculate Symm Eigenvalues
# + [markdown] colab_type="text" id="AiyR1e2NubKa"
# Let's exploit the XLA QR implementation to solve some eigenvalues for symmetric matrices.
#
# This is the naive QR algorithm, without acceleration for closely-spaced eigenvalue convergence, nor any permutation to sort eigenvalues by magnitude.
# + colab={"height": 455} colab_type="code" executionInfo={"elapsed": 1262, "status": "ok", "timestamp": 1549929575801, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-rqjtQZ4KjgQ/AAAAAAAAAAI/AAAAAAAAAAw/5BQt0zmTW5o/s64/photo.jpg", "userId": "09409386770882740563"}, "user_tz": 480} id="wjxDPbqCcuXT" outputId="9683e40b-3c5f-4f3e-c971-0613b182c68c"
Niter = 200
matrix_shape = (10, 10)
in_shape = shape_of(
(onp.zeros(matrix_shape, dtype=onp.float32), 1)
)
# NB: in_shape is the same as the manually constructed:
# xla_client.Shape.tuple_shape(
# (xla_client.Shape.array_shape(onp.dtype(onp.float32), matrix_shape),
# xla_client.Shape.array_shape(onp.dtype(onp.int32), ()))
# )
# body computation -- QR loop: X_i = Q R , X_{i+1} = R Q
bcb = xla_client.ComputationBuilder("bodycomp")
intuple = bcb.ParameterWithShape(in_shape)
x = bcb.GetTupleElement(intuple, 0)
cntr = bcb.GetTupleElement(intuple, 1)
QR = bcb.QR(x)
Q = bcb.GetTupleElement(QR, 0)
R = bcb.GetTupleElement(QR, 1)
RQ = bcb.Dot(R, Q)
bcb.Tuple(RQ, bcb.Sub(cntr, bcb.Constant(onp.int32(1))))
body_computation = bcb.Build()
# test computation -- just a for loop condition
tcb = xla_client.ComputationBuilder("testcomp")
intuple = tcb.ParameterWithShape(in_shape)
cntr = tcb.GetTupleElement(intuple, 1)
test = tcb.Gt(cntr, tcb.Constant(onp.int32(0)))
test_computation = tcb.Build()
# while computation:
wcb = xla_client.ComputationBuilder("whilecomp")
intuple = wcb.ParameterWithShape(in_shape)
wcb.While(test_computation, body_computation, intuple)
while_computation = wcb.Build()
# Now compile and execute:
compiled_computation = while_computation.Compile([in_shape,])
X = onp.random.random(matrix_shape).astype(onp.float32)
X = (X + X.T) / 2.0
it = onp.array(Niter, dtype=onp.int32)
device_in = xla_client.LocalBuffer.from_pyval((X, it))
device_out = compiled_computation.Execute([device_in,])
host_out = device_out.to_py()
eigh_vals = host_out[0].diagonal()
plt.title('D')
plt.imshow(host_out[0])
print('sorted eigenvalues')
print(onp.sort(eigh_vals))
print('sorted eigenvalues from numpy')
print(onp.sort(onp.linalg.eigh(X)[0]))
print('sorted error')
print(onp.sort(eigh_vals) - onp.sort(onp.linalg.eigh(X)[0]))
# + [markdown] colab_type="text" id="FpggTihknAOw"
# ## Calculate Full Symm Eigensystem
# + [markdown] colab_type="text" id="Qos4ankYuj1T"
# We can also calculate the eigenbasis by accumulating the Qs.
# + colab={"height": 1000} colab_type="code" executionInfo={"elapsed": 1569, "status": "ok", "timestamp": 1549929587147, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-rqjtQZ4KjgQ/AAAAAAAAAAI/AAAAAAAAAAw/5BQt0zmTW5o/s64/photo.jpg", "userId": "09409386770882740563"}, "user_tz": 480} id="Kp3A-aAiZk0g" outputId="ebdc1ecc-c9e1-4e95-b989-9645f8648ee0"
Niter = 100
matrix_shape = (10, 10)
in_shape = shape_of(
(onp.zeros(matrix_shape, dtype=onp.float32),
onp.eye(matrix_shape[0]),
1)
)
# body computation -- QR loop: X_i = Q R , X_{i+1} = R Q
bcb = xla_client.ComputationBuilder("bodycomp")
intuple = bcb.ParameterWithShape(in_shape)
X = bcb.GetTupleElement(intuple, 0)
O = bcb.GetTupleElement(intuple, 1)
cntr = bcb.GetTupleElement(intuple, 2)
QR = bcb.QR(X)
Q = bcb.GetTupleElement(QR, 0)
R = bcb.GetTupleElement(QR, 1)
RQ = bcb.Dot(R, Q)
Onew = bcb.Dot(O, Q)
bcb.Tuple(RQ, Onew, bcb.Sub(cntr, bcb.Constant(onp.int32(1))))
body_computation = bcb.Build()
# test computation -- just a for loop condition
tcb = xla_client.ComputationBuilder("testcomp")
intuple = tcb.ParameterWithShape(in_shape)
cntr = tcb.GetTupleElement(intuple, 2)
test = tcb.Gt(cntr, tcb.Constant(onp.int32(0)))
test_computation = tcb.Build()
# while computation:
wcb = xla_client.ComputationBuilder("whilecomp")
intuple = wcb.ParameterWithShape(in_shape)
wcb.While(test_computation, body_computation, intuple)
while_computation = wcb.Build()
# Now compile and execute:
compiled_computation = while_computation.Compile([in_shape,])
X = onp.random.random(matrix_shape).astype(onp.float32)
X = (X + X.T) / 2.0
Omat = onp.eye(matrix_shape[0], dtype=onp.float32)
it = onp.array(Niter, dtype=onp.int32)
device_in = xla_client.LocalBuffer.from_pyval((X, Omat, it))
device_out = compiled_computation.Execute([device_in,])
host_out = device_out.to_py()
eigh_vals = host_out[0].diagonal()
eigh_mat = host_out[1]
plt.title('D')
plt.imshow(host_out[0])
plt.figure()
plt.title('U')
plt.imshow(eigh_mat)
plt.figure()
plt.title('U^T A U')
plt.imshow(onp.dot(onp.dot(eigh_mat.T, X), eigh_mat))
print('sorted eigenvalues')
print(onp.sort(eigh_vals))
print('sorted eigenvalues from numpy')
print(onp.sort(onp.linalg.eigh(X)[0]))
print('sorted error')
print(onp.sort(eigh_vals) - onp.sort(onp.linalg.eigh(X)[0]))
# + [markdown] colab_type="text" id="Ee3LMzOvlCuK"
# ## Convolutions
#
# I keep hearing from the AGI folks that we can use convolutions to build artificial life. Let's try it out.
# + colab={"height": 132} colab_type="code" executionInfo={"elapsed": 1347, "status": "ok", "timestamp": 1549929594704, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/-rqjtQZ4KjgQ/AAAAAAAAAAI/AAAAAAAAAAw/5BQt0zmTW5o/s64/photo.jpg", "userId": "09409386770882740563"}, "user_tz": 480} id="J8QkirDalBse" outputId="73c53980-8dbd-497b-fe56-7e606a29c19f"
Niter=13
matrix_shape = (1,1, 20, 20)
in_shape = shape_of(
(onp.zeros(matrix_shape, dtype=onp.int32), 1)
)
# Body computation -- Conway Update
bcb = xla_client.ComputationBuilder("bodycomp")
intuple = bcb.ParameterWithShape(in_shape)
x = bcb.GetTupleElement(intuple, 0)
cntr = bcb.GetTupleElement(intuple, 1)
# convs require floating-point type
xf = bcb.ConvertElementType(x, to_xla_type('float32'))
stamp = bcb.Constant(onp.ones((1,1,3,3), dtype=onp.float32))
convd = bcb.Conv(xf, stamp, onp.array([1, 1]), xla_client.PaddingType.SAME)
# logic ops require integer types
convd = bcb.ConvertElementType(convd, to_xla_type('int32'))
bool_x = bcb.Eq(x, bcb.ConstantS32Scalar(1))
# core update rule
res = bcb.Or(
# birth rule
bcb.And(bcb.Not(bool_x), bcb.Eq(convd, bcb.ConstantS32Scalar(3))),
# survival rule
bcb.And(bool_x, bcb.Or(
# these are +1 the normal numbers since conv-sum counts self
bcb.Eq(convd, bcb.ConstantS32Scalar(4)),
bcb.Eq(convd, bcb.ConstantS32Scalar(3)))
)
)
# Convert output back to int type for type constancy
int_res = bcb.ConvertElementType(res, to_xla_type('int32'))
bcb.Tuple(int_res, bcb.Sub(cntr, bcb.ConstantS32Scalar(1)))
body_computation = bcb.Build()
# Test computation -- just a for loop condition
tcb = xla_client.ComputationBuilder("testcomp")
intuple = tcb.ParameterWithShape(in_shape)
cntr = tcb.GetTupleElement(intuple, 1)
test = tcb.Gt(cntr, tcb.ConstantS32Scalar(0))
test_computation = tcb.Build()
# While computation:
wcb = xla_client.ComputationBuilder("whilecomp")
intuple = wcb.ParameterWithShape(in_shape)
wcb.While(test_computation, body_computation, intuple)
while_computation = wcb.Build()
# Now compile and execute:
compiled_computation = while_computation.Compile([in_shape,])
# Set up initial state
X = onp.zeros(matrix_shape, dtype=onp.int32)
X[0,0, 5:8, 5:8] = onp.array([[0,1,0],[0,0,1],[1,1,1]])
# Evolve
movie = onp.zeros((Niter,)+matrix_shape[-2:], dtype=onp.int32)
for it in range(Niter):
itr = onp.array(it, dtype=onp.int32)
device_in = xla_client.LocalBuffer.from_pyval((X, itr))
device_out = compiled_computation.Execute([device_in,])
movie[it] = device_out.to_py()[0][0,0]
# Plot
fig = plt.figure(figsize=(15,2))
gs = gridspec.GridSpec(1,Niter)
for i in range(Niter):
ax1 = plt.subplot(gs[:, i])
ax1.axis('off')
ax1.imshow(movie[i])
plt.subplots_adjust(left=0.0, right=1.0, top=1.0, bottom=0.0, hspace=0.0, wspace=0.05)
# + [markdown] colab_type="text" id="9-0PJlqv237S"
# # Fin
#
# There's much more to XLA, but this hopefully highlights how easy it is to play with via the python client!
| notebooks/XLA_in_Python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solutions for NumPy Exercises
# **Question 1**
#
# Write an array by using `np.arange()` and set it to L1.
# Write the array by using python list and set it to L2.
#
# The list should contains number from 1 to 9.
import numpy as np
L1 = np.arange(1,10)
print(L1)
L2 = [i for i in range(1,10)]
print(L2)
# **Question 2**
#
# Find the square of each numbers in the list in *Question 1*.
L1_square = L1 ** 2
print(L1_square)
L2_square = [i ** 2 for i in range (1,10)]
print(L2_square)
# **Question 3**
#
# Create a 2-dimensional matrix L1 by using `.reshape()`.
L1_2d = L1.reshape((3,3))
print(L1_2d)
# **Question 4**
#
# Transpose the matrix in *Question 3* by using `.T`
L1_2d_T = L1_2d.T
print(L1_2d_T)
# **Question 5**
#
# Multiply the matrix in *Question 3* to matrix in *Question 4* by using `np.dot()`.
dot_L = np.dot(L1_2d, L1_2d_T)
print(dot_L)
# **Question 6**
#
# Add both matrix in *Question 3* and *Question 4* by using `np.add()`.
add_L = np.add(L1_2d, L1_2d_T)
print(add_L)
# **Question 7**
#
# Add both matrix in *Question 3* and *Question 4* by using `np.divide()`.
divide_L = np.divide(L1_2d, L1_2d_T)
print(divide_L)
# **Question 8**
#
# Find the eigenvalues for matrix in *Question 3* and matrix in *Question 5* by using `np.linalg.eigvals()`.
eigen_L1 = np.linalg.eigvals(L1_2d)
eigen_L2 = np.linalg.eigvals(dot_L)
print(eigen_L1)
print(eigen_L2)
| Lesson12/Numpy_solutions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/MJVelilla/CS488S21/blob/main/python_materials/learn-python3/notebooks/beginner/exercises/strings_exercise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Z007n2gibqjU"
# # 1. Fill missing pieces
# Fill `____` pieces below to have correct values for `lower_cased`, `stripped` and `stripped_lower_case` variables.
# + id="SohhNnm5bqja"
original = ' Python strings are COOL! '
lower_cased = original.lower()
stripped = original.strip()
stripped_lower_cased = original.strip().lower()
# + [markdown] id="gsHaUyPbbqjb"
# Let's verify that the implementation is correct by running the cell below. `assert` will raise `AssertionError` if the statement is not true.
# + editable=false id="Nr28HMJgbqjc"
assert lower_cased == ' python strings are cool! '
assert stripped == 'Python strings are COOL!'
assert stripped_lower_cased == 'python strings are cool!'
# + [markdown] id="upYcnvBzbqjc"
# # 2. Prettify ugly string
# Use `str` methods to convert `ugly` to wanted `pretty`.
# + editable=false id="E4ooF4r-bqjc"
ugly = ' tiTle of MY new Book\n\n'
# + id="fWeQSFpmbqjd"
# Your implementation:
pretty = ugly.strip().title()
# + [markdown] id="rqXLa7YRbqjd"
# Let's make sure that it does what we want. `assert` raises [`AssertionError`](https://docs.python.org/3/library/exceptions.html#AssertionError) if the statement is not `True`.
# + editable=false id="b3pSU0tNbqjd" outputId="e7bcb3c5-2c26-4498-bf4e-a8e0b892227a" colab={"base_uri": "https://localhost:8080/"}
print('pretty: {}'.format(pretty))
assert pretty == 'Title Of My New Book'
# + [markdown] id="H_PbWVycbqjd"
# # 3. Format string based on existing variables
# Create `sentence` by using `verb`, `language`, and `punctuation` and any other strings you may need.
# + editable=false id="W4DZv25kbqje"
verb = 'is'
language = 'Python'
punctuation = '!'
# + id="Jb37pgPYbqje"
# Your implementation:
sentence = ' '.join(["Learning", language, verb, "fun"]) + punctuation
# + editable=false id="NgTJ_vmObqje" outputId="6b084d89-5adc-4fbd-8a21-fd02d67c84b8" colab={"base_uri": "https://localhost:8080/"}
print('sentence: {}'.format(sentence))
assert sentence == 'Learning Python is fun!'
| python_materials/learn-python3/notebooks/beginner/exercises/strings_exercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [<NAME>](http://sebastianraschka.com)
#
# - [Open in IPython nbviewer](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/python_howtos/scikit_linear_classificationreate=1)
#
# - [Link to this IPython notebook on Github](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/machine_learning/scikit-learn/scikit_linear_classification.ipynb)
#
# - [Link to the GitHub Repository pattern_classification](https://github.com/rasbt/pattern_classification)
# %load_ext watermark
# %watermark -a 'Gopala KR' -u -d -v -p numpy,scikit-learn,matplotlib
# <hr>
# I would be happy to hear your comments and suggestions.
# Please feel free to drop me a note via
# [twitter](https://twitter.com/rasbt), [email](mailto:<EMAIL>), or [google+](https://plus.google.com/+SebastianRaschka).
# <hr>
# # An Introduction to simple linear supervised classification using `scikit-learn`
# In this introduction I want to give a brief overview of how Python's `scikit-learn` machine learning library can be used for simple linear classification.
# <br>
# <br>
# # Sections
# - [About the dataset](#About-the-dataset)
# - [Reading in a dataset from a CSV file](#Reading-in-a-dataset-from-a-CSV-file)
# - [Visualizing the Wine dataset](#Visualizing-the-Wine-dataset)
# - [Splitting into training and test dataset](#Splitting-into-training-and-test-dataset)
# - [Feature Scaling](#Feature-Scaling)
# - [Introduction to Multiple Discriminant Analysis (MDA)](#MDA)
# - [Classification via LDA](#LDA)
# - [Stochastic Gradient Descent (SGD) as linear classifier](#SGD)
# <br>
# <br>
# <br>
# <br>
# ## About the dataset
# [[back to top]](#Sections)
# For the following tutorial, we will be working with the free "Wine" Dataset that is deposited on the UCI machine learning repository
# (http://archive.ics.uci.edu/ml/datasets/Wine).
#
# <br>
#
# <font size="1">
# **Reference:**
# <NAME> al, PARVUS - An Extendible Package for Data
# Exploration, Classification and Correlation. Institute of Pharmaceutical
# and Food Analysis and Technologies, Via Brigata Salerno,
# 16147 Genoa, Italy.</font>
# <br>
# <br>
# The Wine dataset consists of 3 different classes where each row correspond to a particular wine sample.
#
# The class labels (1, 2, 3) are listed in the first column, and the columns 2-14 correspond to the following 13 attributes (features):
#
# 1) Alcohol
# 2) Malic acid
# 3) Ash
# 4) Alcalinity of ash
# 5) Magnesium
# 6) Total phenols
# 7) Flavanoids
# 8) Nonflavanoid phenols
# 9) Proanthocyanins
# 10) Color intensity
# 11) Hue
# 12) OD280/OD315 of diluted wines
# 13) Proline
#
# An excerpt from the wine_data.csv dataset:
#
# <pre>1,14.23,1.71,2.43,15.6,127,2.8,3.06,.28,2.29,5.64,1.04,3.92,1065
# 1,13.2,1.78,2.14,11.2,100,2.65,2.76,.26,1.28,4.38,1.05,3.4,1050
# [...]
# 2,12.37,.94,1.36,10.6,88,1.98,.57,.28,.42,1.95,1.05,1.82,520
# 2,12.33,1.1,2.28,16,101,2.05,1.09,.63,.41,3.27,1.25,1.67,680
# [...]
# 3,12.86,1.35,2.32,18,122,1.51,1.25,.21,.94,4.1,.76,1.29,630
# 3,12.88,2.99,2.4,20,104,1.3,1.22,.24,.83,5.4,.74,1.42,530</pre>
# <br>
# <br>
# ## Reading in a dataset from a CSV file
# [[back to top]](#Sections)
# Since it is quite typical to have the input data stored locally, as mentioned above, we will use the [`numpy.loadtxt`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.loadtxt.html) function now to read in the data from the CSV file.
# (alternatively [`np.genfromtxt()`](http://docs.scipy.org/doc/numpy/reference/generated/numpy.genfromtxt.html) could be used in similar way, it provides some additional options)
# +
import numpy as np
# reading in all data into a NumPy array
all_data = np.loadtxt(open("../../data/wine_data.csv","r"),
delimiter=",",
skiprows=0,
dtype=np.float64
)
# load class labels from column 1
y_wine = all_data[:,0]
# conversion of the class labels to integer-type array
y_wine = y_wine.astype(np.int64, copy=False)
# load the 14 features
X_wine = all_data[:,1:]
# printing some general information about the data
print('\ntotal number of samples (rows):', X_wine.shape[0])
print('total number of features (columns):', X_wine.shape[1])
# printing the 1st wine sample
float_formatter = lambda x: '{:.2f}'.format(x)
np.set_printoptions(formatter={'float_kind':float_formatter})
print('\n1st sample (i.e., 1st row):\nClass label: {:d}\n{:}\n'
.format(int(y_wine[0]), X_wine[0]))
# printing the rel.frequency of the class labels
print('Class label frequencies')
print('Class 1 samples: {:.2%}'.format(list(y_wine).count(1)/y_wine.shape[0]))
print('Class 2 samples: {:.2%}'.format(list(y_wine).count(2)/y_wine.shape[0]))
print('Class 3 samples: {:.2%}'.format(list(y_wine).count(3)/y_wine.shape[0]))
# -
# <br>
# <br>
# ## Visualizing the Wine dataset
# [[back to top]](#Sections)
# There are endless way to visualize datasets for get an initial idea of how the data looks like. The most common ones are probably histograms and scatter plots.
# Scatter plots are useful for visualizing features in more than just one dimension, for example to get a feeling for the correlation between particular features.
# Unfortunately, we can't plot all 13 features here at once, since the visual cortex of us humans is limited to a maximum of three dimensions.
# Below, we will create an example 2D-Scatter plot from the features "Alcohol content" and "Malic acid content".
# Additionally, we will use the [`scipy.stats.pearsonr`](http://docs.scipy.org/doc/scipy/reference/generated/scipy.stats.pearsonr.html) function to calculate a Pearson correlation coefficient between these two features.
#
# %matplotlib inline
# +
from matplotlib import pyplot as plt
from scipy.stats import pearsonr
plt.figure(figsize=(10,8))
for label,marker,color in zip(
range(1,4),('x', 'o', '^'),('blue', 'red', 'green')):
# Calculate Pearson correlation coefficient
R = pearsonr(X_wine[:,0][y_wine == label], X_wine[:,1][y_wine == label])
plt.scatter(x=X_wine[:,0][y_wine == label], # x-axis: feat. from col. 1
y=X_wine[:,1][y_wine == label], # y-axis: feat. from col. 2
marker=marker, # data point symbol for the scatter plot
color=color,
alpha=0.7,
label='class {:}, R={:.2f}'.format(label, R[0]) # label for the legend
)
plt.title('Wine Dataset')
plt.xlabel('alcohol by volume in percent')
plt.ylabel('malic acid in g/l')
plt.legend(loc='upper right')
plt.show()
# -
# <br>
# <br>
# If we want to pack 3 different features into one scatter plot at once, we can also do the same thing in 3D:
# +
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure(figsize=(8,8))
ax = fig.add_subplot(111, projection='3d')
for label,marker,color in zip(
range(1,4),('x', 'o', '^'),('blue','red','green')):
ax.scatter(X_wine[:,0][y_wine == label],
X_wine[:,1][y_wine == label],
X_wine[:,2][y_wine == label],
marker=marker,
color=color,
s=40,
alpha=0.7,
label='class {}'.format(label))
ax.set_xlabel('alcohol by volume in percent')
ax.set_ylabel('malic acid in g/l')
ax.set_zlabel('ash content in g/l')
plt.legend(loc='upper right')
plt.title('Wine dataset')
plt.show()
# -
# <br>
# <br>
# ## Splitting into training and test dataset
# [[back to top]](#Sections)
# It is a typical procedure for machine learning and pattern classification tasks to split one dataset into two: a training dataset and a test dataset.
# The training dataset is henceforth used to train our algorithms or classifier, and the test dataset is a way to validate the outcome quite objectively before we apply it to "new, real world data".
#
# Here, we will split the dataset randomly so that 70% of the total dataset will become our training dataset, and 30% will become our test dataset, respectively.
# +
from sklearn.cross_validation import train_test_split
from sklearn import preprocessing
X_train, X_test, y_train, y_test = train_test_split(X_wine, y_wine,
test_size=0.30, random_state=123)
# -
# Note that since this a random assignment, the original relative frequencies for each class label are not maintained.
# +
print('Class label frequencies')
print('\nTraining Dataset:')
for l in range(1,4):
print('Class {:} samples: {:.2%}'.format(l, list(y_train).count(l)/y_train.shape[0]))
print('\nTest Dataset:')
for l in range(1,4):
print('Class {:} samples: {:.2%}'.format(l, list(y_test).count(l)/y_test.shape[0]))
# -
# <br>
# <br>
# ## Feature Scaling
# [[back to top]](#Sections)
# Another popular procedure is to standardize the data prior to fitting the model and other analyses so that the features will have the properties of a standard normal distribution with
#
# $\mu = 0$ and $\sigma = 1$
#
# where $\mu$ is the mean (average) and $\sigma$ is the standard deviation from the mean, so that the standard scores of the samples are calculated as follows:
#
# \begin{equation} z = \frac{x - \mu}{\sigma}\end{equation}
std_scale = preprocessing.StandardScaler().fit(X_train)
X_train = std_scale.transform(X_train)
X_test = std_scale.transform(X_test)
# +
f, ax = plt.subplots(1, 2, sharex=True, sharey=True, figsize=(10,5))
for a,x_dat, y_lab in zip(ax, (X_train, X_test), (y_train, y_test)):
for label,marker,color in zip(
range(1,4),('x', 'o', '^'),('blue','red','green')):
a.scatter(x=x_dat[:,0][y_lab == label],
y=x_dat[:,1][y_lab == label],
marker=marker,
color=color,
alpha=0.7,
label='class {}'.format(label)
)
a.legend(loc='upper right')
ax[0].set_title('Training Dataset')
ax[1].set_title('Test Dataset')
f.text(0.5, 0.04, 'malic acid (standardized)', ha='center', va='center')
f.text(0.08, 0.5, 'alcohol (standardized)', ha='center', va='center', rotation='vertical')
plt.show()
# -
# <br>
# <br>
# <a id="PCA"></a>
# <br>
# <br>
# <a id='MDA'></a>
# ## Linear Transformation & Classification: Multiple Discriminant Analysis (MDA)
# [[back to top]](#Sections)
# The main purposes of a Multiple Discriminant Analysis is to analyze the data to identify patterns to project it onto a subspace that yields a better separation of the classes. Also, the dimensionality of the dataset shall be reduced with minimal loss of information.
#
# **The approach is very similar to a Principal Component Analysis (PCA), but in addition to finding the component axes that maximize the variance of our data, we are additionally interested in the axes that maximize the separation of our classes (e.g., in a supervised pattern classification problem)**
#
# Here, our desired outcome of the multiple discriminant analysis is to project a feature space (our dataset consisting of n d-dimensional samples) onto a smaller subspace that represents our data "well" and has a good class separation. A possible application would be a pattern classification task, where we want to reduce the computational costs and the error of parameter estimation by reducing the number of dimensions of our feature space by extracting a subspace that describes our data "best".
# #### Principal Component Analysis (PCA) Vs. Multiple Discriminant Analysis (MDA)
# Both Multiple Discriminant Analysis (MDA) and Principal Component Analysis (PCA) are linear transformation methods and closely related to each other. In PCA, we are interested to find the directions (components) that maximize the variance in our dataset, where in MDA, we are additionally interested to find the directions that maximize the separation (or discrimination) between different classes (for example, in pattern classification problems where our dataset consists of multiple classes. In contrast two PCA, which ignores the class labels).
#
# **In other words, via PCA, we are projecting the entire set of data (without class labels) onto a different subspace, and in MDA, we are trying to determine a suitable subspace to distinguish between patterns that belong to different classes. Or, roughly speaking in PCA we are trying to find the axes with maximum variances where the data is most spread (within a class, since PCA treats the whole data set as one class), and in MDA we are additionally maximizing the spread between classes.**
#
# In typical pattern recognition problems, a PCA is often followed by an MDA.
# 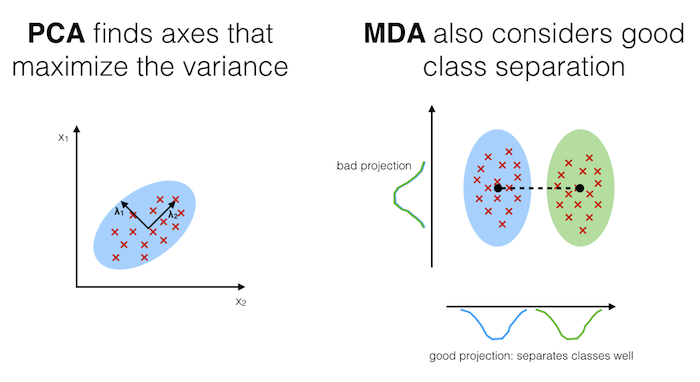
# If you are interested, you can find more information about the MDA in my IPython notebook
# [Stepping through a Multiple Discriminant Analysis - using Python's NumPy and matplotlib](http://nbviewer.ipython.org/github/rasbt/pattern_classification/blob/master/dimensionality_reduction/projection/linear_discriminant_analysis.ipynb?create=1).
# Like we did in the PCA section above, we will use a `scikit-learn` funcion, [`sklearn.lda.LDA`](http://scikit-learn.org/stable/modules/generated/sklearn.lda.LDA.html) in order to transform our training data onto 2 dimensional subspace, where MDA is basically the more generalized form of an LDA (Linear Discriminant Analysis):
# +
from sklearn.discriminant_analysis import LinearDiscriminantAnalysis as LDA
sklearn_lda = LDA(n_components=2)
sklearn_transf = sklearn_lda.fit_transform(X_train, y_train)
plt.figure(figsize=(10,8))
for label,marker,color in zip(
range(1,4),('x', 'o', '^'),('blue', 'red', 'green')):
plt.scatter(x=sklearn_transf[:,0][y_train == label],
y=sklearn_transf[:,1][y_train == label],
marker=marker,
color=color,
alpha=0.7,
label='class {}'.format(label)
)
plt.xlabel('vector 1')
plt.ylabel('vector 2')
plt.legend()
plt.title('Most significant singular vectors after linear transformation via LDA')
plt.show()
# -
# <br>
# <br>
# <br>
# <br>
# ## Classification via LDA
# [[back to top]](#Sections)
# The LDA that we've just used in the section above can also be used as a simple linear classifier.
# +
# fit model
lda_clf = LDA()
lda_clf.fit(X_train, y_train)
LDA(n_components=None, priors=None)
# prediction
print('1st sample from test dataset classified as:', lda_clf.predict(X_test[0,:].reshape(1, -1)))
print('actual class label:', y_test[0])
# -
# Another handy subpackage of sklearn is `metrics`. The [`metrics.accuracy_score`](http://scikit-learn.org/stable/modules/generated/sklearn.metrics.accuracy_score.html), for example, is quite useful to evaluate how many samples can be classified correctly:
# +
from sklearn import metrics
pred_train = lda_clf.predict(X_train)
print('Prediction accuracy for the training dataset')
print('{:.2%}'.format(metrics.accuracy_score(y_train, pred_train)))
# -
# To verify that over model was not overfitted to the training dataset, let us evaluate the classifier's accuracy on the test dataset:
# +
pred_test = lda_clf.predict(X_test)
print('Prediction accuracy for the test dataset')
print('{:.2%}'.format(metrics.accuracy_score(y_test, pred_test)))
# -
# <br>
# <br>
# **Confusion Matrix**
# As we can see above, there was a very low misclassification rate when we'd apply the classifier on the test data set. A confusion matrix can tell us in more detail which particular classes could not classified correctly.
#
# <table cellspacing="0" border="0">
# <colgroup width="60"></colgroup>
# <colgroup span="4" width="82"></colgroup>
# <tr>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" colspan=2 rowspan=2 height="44" align="center" bgcolor="#FFFFFF"><b><font face="Helvetica" size=4><br></font></b></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" colspan=3 align="center" bgcolor="#FFFFFF"><b><font face="Helvetica" size=4>predicted class</font></b></td>
# </tr>
# <tr>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#EEEEEE"><font face="Helvetica" size=4>class 1</font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#EEEEEE"><font face="Helvetica" size=4>class 2</font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#EEEEEE"><font face="Helvetica" size=4>class 3</font></td>
# </tr>
# <tr>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" rowspan=3 height="116" align="center" bgcolor="#F6F6F6"><b><font face="Helvetica" size=4>actual class</font></b></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#EEEEEE"><font face="Helvetica" size=4>class 1</font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#99FFCC"><font face="Helvetica" size=4>True positives</font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#F6F6F6"><font face="Helvetica" size=4><br></font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#F6F6F6"><font face="Helvetica" size=4><br></font></td>
# </tr>
# <tr>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#EEEEEE"><font face="Helvetica" size=4>class 2</font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#FFFFFF"><font face="Helvetica" size=4><br></font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#99FFCC"><font face="Helvetica" size=4>True positives</font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#FFFFFF"><font face="Helvetica" size=4><br></font></td>
# </tr>
# <tr>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#EEEEEE"><font face="Helvetica" size=4>class 3</font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#F6F6F6"><font face="Helvetica" size=4><br></font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#F6F6F6"><font face="Helvetica" size=4><br></font></td>
# <td style="border-top: 1px solid #c1c1c1; border-bottom: 1px solid #c1c1c1; border-left: 1px solid #c1c1c1; border-right: 1px solid #c1c1c1" align="left" bgcolor="#99FFCC"><font face="Helvetica" size=4>True positives</font></td>
# </tr>
# </table>
print('Confusion Matrix of the LDA-classifier')
print(metrics.confusion_matrix(y_test, lda_clf.predict(X_test)))
# As we can see, one sample from class 2 was incorrectly labeled as class 1, from the perspective of class 1, this would be 1 "False Negative" or a "False Postive" from the perspective of class 2, respectively
# <br>
# <a id='SGD'></a>
# ## Stochastic Gradient Descent (SGD) as linear classifier
# [[back to top]](#Sections)
# Let us now compare the classification accuracy of the LDA classifier with a simple classification (we also use the probably not ideal default settings here) via stochastic gradient descent, an algorithm that minimizes a linear objective function.
# More information about the `sklearn.linear_model.SGDClassifier` can be found [here](http://scikit-learn.org/stable/modules/generated/sklearn.linear_model.SGDClassifier.html).
# +
from sklearn.linear_model import SGDClassifier
sgd_clf = SGDClassifier()
sgd_clf.fit(X_train, y_train)
pred_train = sgd_clf.predict(X_train)
pred_test = sgd_clf.predict(X_test)
print('\nPrediction accuracy for the training dataset')
print('{:.2%}\n'.format(metrics.accuracy_score(y_train, pred_train)))
print('Prediction accuracy for the test dataset')
print('{:.2%}\n'.format(metrics.accuracy_score(y_test, pred_test)))
print('Confusion Matrix of the SGD-classifier')
print(metrics.confusion_matrix(y_test, sgd_clf.predict(X_test)))
# -
test complete; Gopal
| tests/others/scikit_linear_classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
lst = ["abc",4125,97.6,True]
lst
lst.count(4125)
lst.copy()
lst.insert(1,2)
lst
lst.pop()
lst
lst.reverse()
lst
tp = (5,"xyz",6.7,"Good Morning")
tp
tp.count("Good Morning")
tp.index(6.7)
s = {57.9,"Hello",0,True,False}
s
s1 = {"Hey",44.9857,709,True,0}
s1
s.difference(s1)
s1.intersection(s)
s1.issubset(s)
s.isdisjoint(s1)
s.symmetric_difference(s1)
s1.intersection_update(s)
s1
s.union(s1)
dt = {"Name":"PQR","Contact":4652748,"Gender":"Female","Blood Group":"O+ve"}
dt
dt.fromkeys("Name")
dt.get("Contact")
dt.update({"Email-id":None})
dt
dt.setdefault("Email-id","<EMAIL>")
dt
dt.setdefault("Color","White")
dt.values()
st = "Hello"
st
st.casefold()
st.encode()
st.ljust(10,"A")
a=st.maketrans("l","p")
st.translate(a)
st.splitlines()
| Day2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gevenbly/TensorAlgs/blob/main/mod_binary_MERA.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="qCml5n60w2Dq"
def define_ham(blocksize):
"""
Define Hamiltonian (quantum critical Ising), perform preliminary blocking
of several sites into an effective site.
"""
# define Pauli matrices
sX = np.array([[0, 1], [1, 0]], dtype=float)
sZ = np.array([[1, 0], [0, -1]], dtype=float)
# define Ising local Hamiltonian
ham_orig = (tprod(sX, sX) - 0.5*tprod(sZ, np.eye(2)) -
0.5*tprod(np.eye(2), sZ))
# shift Hamiltonian to ensure negative defined
en_shift = max(LA.eigh(ham_orig)[0])
ham_loc = ham_orig - en_shift*np.eye(4)
# define block Hamiltonians
d0 = 2 # initial local dim
d1 = d0**blocksize # local dim after blocking
if blocksize==2:
ham_block = (0.5*tprod(ham_loc, np.eye(d0**2)) +
1.0*tprod(np.eye(d0**1), ham_loc, np.eye(d0**1)) +
0.5*tprod(np.eye(d0**2), ham_loc)
).reshape(d0*np.ones(8, dtype=int))
hamAB_init = ham_block.transpose(0,1,4,3,5,6,8,7
).reshape(d1, d1, d1, d1)
hamBA_init = ham_block.transpose(1,0,3,4,6,5,7,8
).reshape(d1, d1, d1, d1)
elif blocksize==3:
ham_block = (1.0*tprod(np.eye(d0**1), ham_loc, np.eye(d0**3)) +
1.0*tprod(np.eye(d0**2), ham_loc, np.eye(d0**2)) +
1.0*tprod(np.eye(d0**3), ham_loc, np.eye(d0**1))
).reshape(d0*np.ones(12, dtype=int))
hamAB_init = ham_block.transpose(0,1,2,5,4,3,6,7,8,11,10,9
).reshape(d1, d1, d1, d1)
hamBA_init = ham_block.transpose(2,1,0,3,4,5,8,7,6,9,10,11
).reshape(d1, d1, d1, d1)
elif blocksize==4:
ham_block = (0.5*tprod(np.eye(d0**1), ham_loc, np.eye(d0**5)) +
1.0*tprod(np.eye(d0**2), ham_loc, np.eye(d0**4)) +
1.0*tprod(np.eye(d0**3), ham_loc, np.eye(d0**3)) +
1.0*tprod(np.eye(d0**4), ham_loc, np.eye(d0**2)) +
0.5*tprod(np.eye(d0**5), ham_loc, np.eye(d0**1))
).reshape(d0*np.ones(16, dtype=int))
hamAB_init = ham_block.transpose(0,1,2,3,7,6,5,4,8,9,10,11,15,14,13,12
).reshape(d1, d1, d1, d1)
hamBA_init = ham_block.transpose(3,2,1,0,4,5,6,7,11,10,9,8,12,13,14,15
).reshape(d1, d1, d1, d1)
return hamAB_init, hamBA_init, en_shift
# + id="3N8zMmAKQYIJ"
def initialize(chi, chimid, hamAB_init, hamBA_init, layers):
""" Initialize the MERA tensors """
# Initialize the MERA tensors
d1 = hamAB_init.shape[0]
iso_temp = orthogonalize(np.random.rand(d1, min(chimid, d1)))
uC = [tprod(iso_temp, iso_temp, do_matricize=False)]
wC = [orthogonalize(np.random.rand(d1, uC[0].shape[2], chi), partition=2)]
vC = [orthogonalize(np.random.rand(d1, uC[0].shape[2], chi), partition=2)]
for k in range(layers-1):
iso_temp = orthogonalize(np.random.rand(chi, chimid))
uC.append(tprod(iso_temp, iso_temp, do_matricize=False))
wC.append(orthogonalize(np.random.rand(chi, chimid, chi), partition=2))
vC.append(orthogonalize(np.random.rand(chi, chimid, chi), partition=2))
# initialize density matrices and effective Hamiltonians
rhoAB = [0]
rhoBA = [0]
hamAB = [hamAB_init]
hamBA = [hamBA_init]
for k in range(layers):
rhoAB.append(np.eye(chi**2).reshape(chi, chi, chi, chi))
rhoBA.append(np.eye(chi**2).reshape(chi, chi, chi, chi))
hamAB.append(np.zeros((chi, chi, chi, chi)))
hamBA.append(np.zeros((chi, chi, chi, chi)))
return hamAB, hamBA, wC, vC, uC, rhoAB, rhoBA
# + id="RSLzMfwC6Jxy"
def define_networks(hamAB, hamBA, wC, vC, uC, rhoAB, rhoBA):
""" Define and plot all principle networks """
# Define the `M` principle network
connects_M = [[3,5,9], [1,5,7], [1,2,3,4], [4,6,10], [2,6,8], [7,8,9,10]]
tensors_M = [vC, vC, hamBA, wC, wC, rhoAB]
order_M = ncon_solver(tensors_M, connects_M)[0]
dims_M = [tensor.shape for tensor in tensors_M]
names_M = ['v', 'v', 'hBA', 'w', 'w', 'rhoAB']
coords_M = [(-0.5,1),(-0.5,-1), (-0.3,-0.2,0.3,0.2),(0.5,1),(0.5,-1),(0.2)]
colors_M = [0,0,1,2,2,3]
# Define the `L` principle network
connects_L = [[3,6,13], [1,8,11], [4,5,6,7], [2,5,8,9], [1,2,3,4],
[10,7,14], [10,9,12], [11,12,13,14]]
tensors_L = [wC, wC, uC, uC, hamAB, vC, vC, rhoBA]
order_L = ncon_solver(tensors_L, connects_L)[0]
dims_L = [tensor.shape for tensor in tensors_L]
names_L = ['w', 'w', 'u', 'u', 'hAB', 'v', 'v', 'rhoBA']
coords_L = [(-0.5, 1.5), (-0.5, -1.5), (-0.3,0.5,0.3,0.9), (-0.3,-0.5,0.3,-0.9),
(-0.6,-0.2,-0.1,0.2), (0.5, 1.5), (0.5, -1.5), (0.2)]
colors_L = [2,2,4,4,1,0,0,3]
# Define the `C` principle network
connects_C = [[5,6,13], [5,9,11], [3,4,6,8], [1,2,9,10], [1,2,3,4], [7,8,14],
[7,10,12], [11,12,13,14]]
tensors_C = [wC, wC, uC, uC, hamBA, vC, vC, rhoBA]
order_C = ncon_solver(tensors_C, connects_C)[0]
dims_C = [tensor.shape for tensor in tensors_C]
names_C = ['w', 'w', 'u', 'u', 'hBA', 'v', 'v', 'rhoBA']
coords_C = [(-0.5, 1.5), (-0.5, -1.5), (-0.3,0.5,0.3,0.9), (-0.3,-0.5,0.3,-0.9),
(-0.3,-0.2,0.3,0.2), (0.5, 1.5), (0.5, -1.5), (0.2)]
colors_C = [2,2,4,4,1,0,0,3]
# Define the `R` principle network
connects_R = [[10,6,13], [10,8,11], [5,3,6,7], [5,1,8,9], [1,2,3,4], [4,7,14],
[2,9,12], [11,12,13,14]]
tensors_R = [wC, wC, uC, uC, hamAB, vC, vC, rhoBA]
order_R = ncon_solver(tensors_R, connects_R)[0]
dims_R = [tensor.shape for tensor in tensors_R]
names_R = ['w', 'w', 'u', 'u', 'hAB', 'v', 'v', 'rhoBA']
coords_R = [(-0.5, 1.5), (-0.5, -1.5), (-0.3,0.5,0.3,0.9), (-0.3,-0.5,0.3,-0.9),
(0.6,-0.2,0.1,0.2), (0.5, 1.5), (0.5, -1.5), (0.2)]
colors_R = [2,2,4,4,1,0,0,3]
# Plot all principle networks
fig = plt.figure(figsize=(24,24))
figM = draw_network(connects_M, order=order_M, dims=dims_M, coords=coords_M,
names=names_M, colors=colors_M, title='M-diagrams',
draw_labels=False, show_costs=True, legend_extend=2.5,
fig=fig, subplot=141, env_pad=(-0.4,-0.4))
figL = draw_network(connects_L, order=order_L, dims=dims_L, coords=coords_L,
names=names_L, colors=colors_L, title='L-diagrams',
draw_labels=False, show_costs=True, legend_extend=2.5,
fig=fig, subplot=142, env_pad=(-0.4,-0.4))
figC = draw_network(connects_C, order=order_C, dims=dims_C, coords=coords_C,
names=names_C, colors=colors_C, title='C-diagrams',
draw_labels=False, show_costs=True, legend_extend=2.5,
fig=fig, subplot=143, env_pad=(-0.4,-0.4))
figR = draw_network(connects_R, order=order_R, dims=dims_R, coords=coords_R,
names=names_R, colors=colors_R, title='R-diagrams',
draw_labels=False, show_costs=True, legend_extend=2.5,
fig=fig, subplot=144, env_pad=(-0.4,-0.4))
# Store `connects` and `order` in a dict for later use
network_dict = {'connects_M': connects_M, 'order_M': order_M,
'connects_L': connects_L, 'order_L': order_L,
'connects_C': connects_C, 'order_C': order_C,
'connects_R': connects_R, 'order_R': order_R,}
return network_dict
# + id="Ec3x6ITl7qrv"
def lift_hamiltonian(hamAB, hamBA, w, v, u, rhoAB, rhoBA, network_dict,
ref_sym=False):
""" Lift the Hamiltonian through one MERA layer """
hamAB_lift = xcon([v, v, hamBA, w, w, rhoAB],
network_dict['connects_M'],
order=network_dict['order_M'], which_envs=5)
hamBA_temp0 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_L'],
order=network_dict['order_L'], which_envs=7)
hamBA_temp1 = xcon([w, w, u, u, hamBA, v, v, rhoBA],
network_dict['connects_C'],
order=network_dict['order_C'], which_envs=7)
if ref_sym is True:
hamBA_temp2 = hamBA_temp0.transpose(1,0,3,2)
else:
hamBA_temp2 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_R'],
order=network_dict['order_R'], which_envs=7)
hamBA_lift = hamBA_temp0 + hamBA_temp1 + hamBA_temp2
return hamAB_lift, hamBA_lift
# + id="Rt89CEM2NS2y"
def lower_density(hamAB, hamBA, w, v, u, rhoAB, rhoBA, network_dict,
ref_sym=False):
""" Lower the density matrix through one MERA layer """
rhoBA_temp0 = xcon([v, v, hamBA, w, w, rhoAB],
network_dict['connects_M'],
order=network_dict['order_M'], which_envs=2)
rhoAB_temp0 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_L'],
order=network_dict['order_L'], which_envs=4)
rhoBA_temp1 = xcon([w, w, u, u, hamBA, v, v, rhoBA],
network_dict['connects_C'],
order=network_dict['order_C'], which_envs=4)
if ref_sym is True:
rhoAB_temp1 = rhoAB_temp0.transpose(1,0,3,2)
else:
rhoAB_temp1 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_R'],
order=network_dict['order_R'], which_envs=4)
rhoAB_lower = 0.5*(rhoAB_temp0 + rhoAB_temp1)
rhoBA_lower = 0.5*(rhoBA_temp0 + rhoBA_temp1)
return rhoAB_lower, rhoBA_lower
# + id="48kn9l27ONxp"
def optimize_w(hamAB, hamBA, w, v, u, rhoAB, rhoBA, network_dict,
ref_sym=False):
""" Optimise the `w` isometry """
w_env0 = xcon([v, v, hamBA, w, w, rhoAB], network_dict['connects_M'],
order=network_dict['order_M'], which_envs=3)
if ref_sym is True:
w_env1, w_env3 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_L'],
order=network_dict['order_L'],
which_envs=[0,5])
else:
w_env1 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_L'], order=network_dict['order_L'],
which_envs=0)
w_env3 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_R'], order=network_dict['order_R'],
which_envs=0)
w_env2 = xcon([w, w, u, u, hamBA, v, v, rhoBA],
network_dict['connects_C'], order=network_dict['order_C'],
which_envs=0)
w_out = orthogonalize(w_env0 + w_env1 + w_env2 + w_env3, partition=2)
return w_out
# + id="Di3CNfrxZWoQ"
def optimize_v(hamAB, hamBA, w, v, u, rhoAB, rhoBA, network_dict,
ref_sym=False):
""" Optimise the `v` isometry """
v_env0 = xcon([v, v, hamBA, w, w, rhoAB], network_dict['connects_M'],
order=network_dict['order_M'], which_envs=0)
if ref_sym is True:
v_env1, v_env3 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_L'],
order=network_dict['order_L'],
which_envs=[0,5])
else:
v_env1 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_L'], order=network_dict['order_L'],
which_envs=5)
v_env3 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_R'], order=network_dict['order_R'],
which_envs=5)
v_env2 = xcon([w, w, u, u, hamBA, v, v, rhoBA],
network_dict['connects_C'], order=network_dict['order_C'],
which_envs=5)
v_out = orthogonalize(v_env0 + v_env1 + v_env2 + v_env3, partition=2)
return v_out
# + id="IGciWMuaZXUB"
def optimize_u(hamAB, hamBA, w, v, u, rhoAB, rhoBA, network_dict,
ref_sym=False):
""" Optimise the `u` disentangler """
u_env0 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_L'], order=network_dict['order_L'],
which_envs=2)
u_env1 = xcon([w, w, u, u, hamBA, v, v, rhoBA],
network_dict['connects_C'], order=network_dict['order_C'],
which_envs=2)
if ref_sym is True:
u_env2 = u_env0.transpose(1,0,3,2)
else:
u_env2 = xcon([w, w, u, u, hamAB, v, v, rhoBA],
network_dict['connects_R'], order=network_dict['order_R'],
which_envs=2)
utot = u_env0 + u_env1 + u_env2
if ref_sym is True:
utot = utot + utot.transpose(1,0,3,2)
u_out = orthogonalize(utot, partition=2)
return u_out
| mod_binary_MERA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Ciclo while
i=1
while i <= 3:
print(i)
i+=1
print("Programa terminado!!!")
condicion = True
i = 10
while condicion:
if i >= 50:
condicion=False
print (i)
i+=10
a = 5
while a < 406:
print(a)
a*=3
# Generar las siguientes
#
# 3,5,7,9
#
# 5,2,-1,-4
a=3
n = int(input("¿Hasta que número?"))
if(n<a):
print("La secuencia inicia en 3, tu numero es MENOR a este")
while a <= n:
print(a)
a+=2
a=5
n = int(input("¿Hasta que número?"))
while a >= n:
print(a)
a-=3
len("Hola mundo")
a="<NAME>"
a[1]
for i in "<NAME>":
print(i,i)
print("Programa terminado")
for i in [5,15,45,135,405]:
print(i==15)
for i in range(2,10):
print(i)
print("Programa terminado")
for i in range(0,10,2):
print(i)
print("Programa terminado")
type( range(5,10,2))
# Convertir a tipo lista
lis= list(range(5,10,2))
print(lis)
for i in "12345":
print(type(i),i+i)
# +
i=1
while i <=8:
print(i)
i+=1
if i == 5:
break
print("Segundo i:",i)
print("Segundo i:",i)
print("Segundo i:",i)
print("Segundo i:",i)
print("Programa terminado")
# +
i=1
while i <=8:
print(i)
i+=1
if i == 5:
continue
print("Segundo i:",i)
print("Segundo i:",i)
print("Segundo i:",i)
print("Segundo i:",i)
print("Programa terminado")
# -
# Aleatorios
import random as ran
# +
# ran.randint?
# -
num_adivinador = ran.randint(1,6)
print(num_adivinador)
# Adivinar el numero que la máquina esta pensado
# SOLO tienes 3 oportunidades
n1 = ran.randint(0,5)
vidas=int(input("¿Cuantass vidas quieres?"))
if vidas>5+1:
print("Seas mamon")
else:
while vidas > 0:
n = int(input("¿En que número estoy pensando?"))
if(n == n1):
print("Adivinaste!!!")
break
else:
vidas-=1
print("Vidas restantes:",vidas)
if vidas==0:
print("Perdiste!!!")
| While.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Importing packages
import fedjax
import jax
import jax.numpy as jnp
import PLM_computation
import FedMix_computation
from grid_search import FedMixGrid, grid_search
from EMNIST_custom import emnist_load_gd_data
import itertools
from matplotlib import pyplot as plt
import pickle
# # Model setup
model = fedjax.models.emnist.create_conv_model(only_digits=False)
def loss(params, batch, rng):
# `rng` used with `apply_for_train` to apply dropout during training.
preds = model.apply_for_train(params, batch, rng)
# Per example loss of shape [batch_size].
example_loss = model.train_loss(batch, preds)
return jnp.mean(example_loss)
def loss_for_eval(params, batch):
preds = model.apply_for_eval(params, batch)
example_loss = model.train_loss(batch, preds)
return jnp.mean(example_loss)
grad_fn = jax.jit(jax.grad(loss))
grad_fn_eval = jax.jit(jax.grad(loss_for_eval))
# + [markdown] tags=[]
# # Grid search setup
# -
# ## Constants
CACHE_DIR = '../data/'
NUM_CLIENTS_GRID_SEARCH = 200
TRAIN_VALIDATION_SPLIT = 0.8
NUM_CLIENTS_PER_PLM_ROUND = 5
NUM_CLIENTS_PER_FEDMIX_ROUND = 10
FEDMIX_ALGORITHM = 'adam'
FEDMIX_NUM_ROUNDS = 500
PLM_NUM_EPOCHS = 100
# ## Datasets and parameters
train_fd, validation_fd = emnist_load_gd_data(
train_val_split=TRAIN_VALIDATION_SPLIT,
only_digits=False,
cache_dir=CACHE_DIR
)
client_ids = set([cid for cid in itertools.islice(
train_fd.client_ids(), NUM_CLIENTS_GRID_SEARCH)])
train_fd = fedjax.SubsetFederatedData(train_fd, client_ids)
validation_fd = fedjax.SubsetFederatedData(validation_fd, client_ids)
plm_init_params = model.init(jax.random.PRNGKey(0))
plm_comp_params = PLM_computation.PLMComputationProcessParams(
plm_init_params, NUM_CLIENTS_PER_PLM_ROUND)
fedmix_init_params = model.init(jax.random.PRNGKey(20))
fedmix_comp_params = FedMix_computation.FedMixComputationParams(
FEDMIX_ALGORITHM, fedmix_init_params, FEDMIX_NUM_ROUNDS)
alpha = 0.7
# ## Grid
fedmix_lrs = 10**jnp.arange(-5., 0.5, 1)
fedmix_batch_sizes = [20, 50, 100, 200]
plm_lrs = 10**jnp.arange(-5., 0.5, 1)
plm_batch_sizes = [10, 20, 50, 100]
grid = FedMixGrid(fedmix_lrs, plm_lrs, fedmix_batch_sizes, plm_batch_sizes)
# # Grid search
SAVE_FILE = '../results/EMNIST_{}_gd.npy'.format(int(10 * alpha))
SAVE_FILE
table = grid_search(
train_fd, validation_fd, grad_fn, grad_fn_eval, model, alpha,
plm_comp_params, fedmix_comp_params, grid, PLM_NUM_EPOCHS,
NUM_CLIENTS_PER_FEDMIX_ROUND, SAVE_FILE
)
best_ind = jnp.unravel_index(jnp.argmax(table), table.shape)
best_ind
plm_batch_size = plm_batch_sizes[best_ind[0]]
plm_lr = plm_lrs[best_ind[1]]
fedmix_batch_size = fedmix_batch_sizes[best_ind[2]]
fedmix_lr = fedmix_lrs[best_ind[3]]
# # FedMix
num_rounds = 3000
# Now we download full train and test datasets.
train_fd, test_fd = fedjax.datasets.emnist.load_data(only_digits=False,
cache_dir='../data/')
plm_comp_hparams = PLM_computation.PLMComputationHParams(PLM_NUM_EPOCHS,
plm_lr,
plm_batch_size)
PLM_dict = PLM_computation.plm_computation(train_fd,
grad_fn,
plm_comp_hparams,
plm_comp_params)
alpha
alpha_dict = {}
for cid in train_fd.client_ids():
alpha_dict[cid] = alpha
len(alpha_dict)
fedmix_hparams = FedMix_computation.FedMixHParams(
fedmix_lr, NUM_CLIENTS_PER_FEDMIX_ROUND, fedmix_batch_size)
fedmix_batch_size
fedmix_comp_params = FedMix_computation.FedMixComputationParams(
FEDMIX_ALGORITHM, fedmix_init_params, num_rounds)
_, stats = FedMix_computation.fedmix_computation_with_statistics(
train_fd, test_fd, grad_fn, grad_fn_eval, model, PLM_dict, alpha_dict,
fedmix_hparams, fedmix_comp_params, 100)
# # FedAvg
client_optimizer = fedjax.optimizers.sgd(learning_rate=10**(-1.5))
server_optimizer = fedjax.optimizers.adam(
learning_rate=10**(-2.5), b1=0.9, b2=0.999, eps=10**(-4))
# Hyperparameters for client local traing dataset preparation.
client_batch_hparams = fedjax.ShuffleRepeatBatchHParams(batch_size=20)
algorithm = fedjax.algorithms.fed_avg.federated_averaging(grad_fn, client_optimizer,
server_optimizer,
client_batch_hparams)
# Initialize model parameters and algorithm server state.
init_params = model.init(jax.random.PRNGKey(17))
server_state = algorithm.init(init_params)
train_client_sampler = fedjax.client_samplers.UniformGetClientSampler(fd=train_fd, num_clients=10, seed=0)
fedavg_test_acc_progress = []
for round_num in range(1, max_rounds + 1):
# Sample 10 clients per round without replacement for training.
clients = train_client_sampler.sample()
# Run one round of training on sampled clients.
server_state, client_diagnostics = algorithm.apply(server_state, clients)
print(f'[round {round_num}]', end='\r')
# Optionally print client diagnostics if curious about each client's model
# update's l2 norm.
# print(f'[round {round_num}] client_diagnostics={client_diagnostics}')
if round_num % 100 == 0:
test_eval_datasets = [cds for _, cds in test_fd.clients()]
test_eval_batches = fedjax.padded_batch_client_datasets(test_eval_datasets, batch_size=256)
test_metrics = fedjax.evaluate_model(model, server_state.params, test_eval_batches)
fedavg_test_acc_progress.append(test_metrics['accuracy'])
print('Test accuracy = {}'.format(test_metrics['accuracy']))
save_file = '../results/test_acc_fedavg.pickle'
with open(save_file, 'wb') as handle:
pickle.dump(fedavg_test_acc_progress, handle)
with open(save_file, 'rb') as handle:
fedavg_test_acc_progress = pickle.load(handle)
fedavg_test_acc_progress = fedavg_test_acc_progress[:30]
# # Plots
accs = [stat['accuracy'] for stat in stats]
round_nums = jnp.linspace(100, 3000, num=30, endpoint=True)
plt.plot(round_nums, accs, label='FLIX')
plt.plot(round_nums, fedavg_test_acc_progress, label='FedAvg')
plt.xlim(left=0)
plt.ylabel('accuracy')
plt.xlabel('rounds')
plt.grid()
plt.title('EMNIST')
plt.legend()
plt.tight_layout()
plt.savefig('../results/plots/EMNIST_preliminary_7.pdf')
| notebooks/EMNIST.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Hinton diagrams
#
#
# Hinton diagrams are useful for visualizing the values of a 2D array (e.g.
# a weight matrix): Positive and negative values are represented by white and
# black squares, respectively, and the size of each square represents the
# magnitude of each value.
#
# Initial idea from <NAME> on the SciPy Cookbook
#
#
# +
import numpy as np
import matplotlib.pyplot as plt
def hinton(matrix, max_weight=None, ax=None):
"""Draw Hinton diagram for visualizing a weight matrix."""
ax = ax if ax is not None else plt.gca()
if not max_weight:
max_weight = 2 ** np.ceil(np.log(np.abs(matrix).max()) / np.log(2))
ax.patch.set_facecolor('gray')
ax.set_aspect('equal', 'box')
ax.xaxis.set_major_locator(plt.NullLocator())
ax.yaxis.set_major_locator(plt.NullLocator())
for (x, y), w in np.ndenumerate(matrix):
color = 'white' if w > 0 else 'black'
size = np.sqrt(np.abs(w) / max_weight)
rect = plt.Rectangle([x - size / 2, y - size / 2], size, size,
facecolor=color, edgecolor=color)
ax.add_patch(rect)
ax.autoscale_view()
ax.invert_yaxis()
if __name__ == '__main__':
# Fixing random state for reproducibility
np.random.seed(19680801)
hinton(np.random.rand(20, 20) - 0.5)
plt.show()
| utils/hinton_demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task Two: Qubits, Bloch Sphere and Basis States
# # Installing Qiskit
# Installing Qiskit:
#
# Step 1: Go to Anaconda Powershell
# Step 2: Type "pip install qiskit"
# Alternatively, pip install qiskit[visualization]
#
# -----OR-----
#
# Uncomment the next line of code to dirtectly install Qiskit from the Jupyter Notebook Code Shell
# +
#Uncomment the below line
# #!pip install qiskit
# #!pip install qiskit[visualization]
# -
import numpy as np
# Only after running above cell or installing Qiskit from the Anaconda Prompt,Run the below cells-->
import qiskit
qiskit.__qiskit_version__
# \begin{align}
# |\psi \rangle = \begin{pmatrix}
# \alpha \\ \beta
# \end{pmatrix}, \quad \text{where } \sqrt{\langle \psi | \psi \rangle} = 1.
# \end{align}
# Think of Qubit as an Electron:
#
# \begin{align}
# \text{spin-up}: \ |0\rangle &= \begin{pmatrix} 1\\0 \end{pmatrix} \\
# \text{spin-down}: \ |1\rangle & = \begin{pmatrix} 0\\1 \end{pmatrix}
# \end{align}
#Import everything from qiskit
from qiskit import *
# Another representation is via Bloch Sphere
# +
from qiskit.visualization import plot_bloch_vector
plot_bloch_vector([0,0,1],title = 'spinup')
# -
# ## Spin + / - :
#
# \begin{align}
# \text{spin +}: \ |+\rangle &= \begin{pmatrix} 1/\sqrt{2} \\ 1/\sqrt{2} \end{pmatrix} = \frac{1}{\sqrt{2}} \left(|0\rangle + |1\rangle\right) \\
# \text{spin -}: \ |-\rangle & = \begin{pmatrix} 1/\sqrt{2} \\ -1/\sqrt{2} \end{pmatrix} = \frac{1}{\sqrt{2}} \left(|0\rangle - |1\rangle\right)
# \end{align}
plot_bloch_vector([1,0,0])
# ## Basis States
#
# \begin{align}
# |0\rangle &= \begin{pmatrix} 1\\0 \end{pmatrix} \\
# |1\rangle & = \begin{pmatrix} 0\\1 \end{pmatrix}
# \end{align}
# Preapring other states from Basis States:
# \begin{align}
# |00 \rangle &= |0\rangle \otimes |0\rangle = \begin{pmatrix} 1\\0 \end{pmatrix} \otimes \begin{pmatrix} 1\\0 \end{pmatrix} = \begin{pmatrix} 1\\0\\0\\0 \end{pmatrix} \\
# |01 \rangle &= |0\rangle \otimes |1\rangle = \begin{pmatrix} 1\\0 \end{pmatrix} \otimes \begin{pmatrix} 0\\1 \end{pmatrix} = \begin{pmatrix} 0\\1\\0\\0 \end{pmatrix} \\
# |10 \rangle &= |1\rangle \otimes |0\rangle = \begin{pmatrix} 0\\1 \end{pmatrix} \otimes \begin{pmatrix} 1\\0 \end{pmatrix} = \begin{pmatrix} 0\\0\\1\\0 \end{pmatrix} \\
# |11 \rangle &= |1\rangle \otimes |1\rangle = \begin{pmatrix} 0\\1 \end{pmatrix} \otimes \begin{pmatrix} 0\\1 \end{pmatrix} = \begin{pmatrix} 0\\0\\0\\1 \end{pmatrix}
# \end{align}
ket_zero = np.array([[1],[0]])
ket_one = np.array([[0],[1]])
np.kron(ket_one,ket_zero)
# **So we have prepared the basis state of a Two Qubit System**
# # That's all for Task 2
# ## Thank You!
# Where can you find me?
#
# LinkedIn : https://www.linkedin.com/in/arya--shah/
#
# Twitter : https://twitter.com/aryashah2k
#
# Github : https://github.com/aryashah2k
#
# If you Like My Work, Follow me/ Connect with me on these platforms.
# Show some Love ❤️ Sponsor me on Github!
| Guided Project - Programming a Quantum Computer with Qiskit - IBM SDK/Task 2/Task 2 - Qubits, Bloch Sphere and Basis States.ipynb |