code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
my_data = np.random.random((2,8)) #recfromcsv('LIAB.ST.csv', delimiter='\t')
new_col = my_data.sum(1)[...,None] # None keeps (n, 1) shape
new_col.shape
#(210,1)
print(my_data)
all_data = np.append(my_data, new_col, 1)
all_data.shape
print("\n\n\n\n",all_data)
#(210,9)
# +
import numpy as np
no_train_mat_col = 16
attributes = np.arange(0,no_train_mat_col-1,1,int)
#print(attributes)
a =[]
a.append([5,6])
a.append([7,8])
#print(a)
b=[]
b.append([0,12,5,1,2,0,1,0,2])
b.append([0,1,2,3,4,5,6,8,9])
b.append([0,1,2,3,4,5,6,8,9])
b= np.array(b)
#print(b)
g = np.where(b == 0)
#print(len(g[0]))
#b = np.eye(2)
#print(b[0][-1])
print(len(np.unique(b[:,-1])))
# -
a =[2,5,3,6,4]
path = np.where(a==np.amax(a))
print(path[0][0])
import numpy as np
from scipy import stats
arr = np.array([1,2,4,5,0,3,0,6,2])
list_sorted = arr.argsort()[:6]
print(list_sorted)
print(arr[list_sorted])
mode_class = stats.mode(arr[list_sorted])
print(mode_class[0])
import statistics as s
import numpy as np
arr = np.array([0,9,4,5,2,3,0,6,2])
multi = s.multimode(arr)
print(multi)
| CSE4309/src/program_garage.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Extracting a Custom Property
from chemdataextractor import Document
from chemdataextractor.model import Compound
from chemdataextractor.doc import Paragraph, Heading
# ## Example Document
# Let's create a simple example document with a single heading followed by a single paragraph:
d = Document(
Heading(u'Synthesis of 2,4,6-trinitrotoluene (3a)'),
Paragraph(u'The procedure was followed to yield a pale yellow solid (b.p. 240 °C)')
)
# What does this look like:
d
# ## Default Parsers
#
# By default, ChemDataExtractor won't extract the boiling point property:
d.records.serialize()
# ## Defining a New Property Model
#
# The first task is to define the schema of a new property, and add it to the `Compound` model:
# +
from chemdataextractor.model import BaseModel, StringType, ListType, ModelType
class BoilingPoint(BaseModel):
value = StringType()
units = StringType()
Compound.boiling_points = ListType(ModelType(BoilingPoint))
# -
# ## Writing a New Parser
#
# Next, define parsing rules that define how to interpret text and convert it into the model:
# +
import re
from chemdataextractor.parse import R, I, W, Optional, merge
prefix = (R(u'^b\.?p\.?$', re.I) | I(u'boiling') + I(u'point')).hide()
units = (W(u'°') + Optional(R(u'^[CFK]\.?$')))(u'units').add_action(merge)
value = R(u'^\d+(\.\d+)?$')(u'value')
bp = (prefix + value + units)(u'bp')
# +
from chemdataextractor.parse.base import BaseParser
from chemdataextractor.utils import first
class BpParser(BaseParser):
root = bp
def interpret(self, result, start, end):
compound = Compound(
boiling_points=[
BoilingPoint(
value=first(result.xpath('./value/text()')),
units=first(result.xpath('./units/text()'))
)
]
)
yield compound
# -
Paragraph.parsers = [BpParser()]
# ## Running the New Parser
# +
d = Document(
Heading(u'Synthesis of 2,4,6-trinitrotoluene (3a)'),
Paragraph(u'The procedure was followed to yield a pale yellow solid (b.p. 240 °C)')
)
d.records.serialize()
| examples/extracting_a_custom_property.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#implementation of logistic regression with 2 features and 2 classes
import numpy as np
from numpy.random import randn, uniform
from numpy import matmul
import matplotlib.pyplot as plt
def vector_sigmoid(vector):
# apply the sigmoid function to all elements of a vector
return 1 / (1 + 2.718 ** -vector)
def normalized(vector):
return (vector - min(vector)) / (max(vector) - min(vector))
def get_derivative(X, y, theta, theta_index):
# get the partial derivative of the cost function relative to x[theta_index] (for each training example)
return (1 / m) * sum((vector_sigmoid(matmul(X, theta)) - y) * X[:, theta_index])
m = 100
m_div = m // 2 # number of examples in each cluster
X = np.array((np.ones(m),
np.append(randn(m_div) + uniform(0, 10), randn(m_div) + uniform(4, 6)),
np.append(randn(m_div) + uniform(0, 10), randn(m_div) + uniform(4, 6)),))
y = np.append(np.zeros(m_div), np.ones(m_div))
X[1:] = np.array([normalized(x) for x in X[1:]]) # normalize X
y = normalized(y) # normalize y
X = X.T # for easier computation
theta = np.full(len(X[0]), 0.5)
delta_theta = np.ones(theta.shape)
a = 10 # learning rate
# i = 0
while abs(sum(delta_theta)) > 5 * 10 ** -4: # controls the needed decrease in cost to continue
for t in range(len(delta_theta)):
delta_theta[t] = - a * get_derivative(X, y, theta, t)
theta += delta_theta
# i += 1
# print(i, theta, delta_theta)
# plt.scatter(X[:, 1][:m_div], X[:, 2][:m_div], color='b')
# plt.scatter(X[:, 1][m_div:], X[:, 2][m_div:], color='r')
# accuracy = 100
# divisor_x_values = np.arange(0, 1, 1 / accuracy)
# divisor_y_values = (-theta[0] - divisor_x_values * theta[1]) / theta[2]
# plt.plot(divisor_x_values, divisor_y_values, color='black', linewidth='1') # show point where the sigmoid function reaches 0.5
# plt.xlabel('x1')
# plt.ylabel('x2')
# plt.show()
# plt.pause(0.001)
# plt.clf()
plt.scatter(X[:, 1][:m_div], X[:, 2][:m_div], color='b')
plt.scatter(X[:, 1][m_div:], X[:, 2][m_div:], color='r')
accuracy = 100
divisor_x_values = np.arange(0, 1, 1 / accuracy)
divisor_y_values = (-theta[0] - divisor_x_values * theta[1]) / theta[2] # derived from sigmoid(X*theta) = 0.5
plt.plot(divisor_x_values, divisor_y_values, color='black', linewidth='1') # show point where the sigmoid function reaches 0.5
plt.xlabel('x1')
plt.ylabel('x2')
plt.show()
| logistic_regression/log_reg_2_features.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
import time
sys.path.append('/home/kevinteng/Desktop/BrainTumourSegmentation')
import numpy as np
import tensorflow as tf
import matplotlib.pyplot as plt
import os, random
import utils
from utils_vis import plot_comparison, plot_labels_color
from utils import compute_metric_dc
import nibabel as nib
from sklearn.model_selection import KFold
# %matplotlib inline
# - Blue => Label 1 (Necrotic and Non-enhancing Tumor Core)
# - Yellow => Label 2 (Peritumoral Edema)
# - Green => Label 3/4 (GD-Enhancing Tumor)
# ---
# * Core => Label 1 & 3
# * Enhancing => Label 3
# * Complete => Label 1,2, 3
# ---
# # Hyperparameter
SHUFFLE_BUFFER = 4000
max_epochs = 20
BATCH_SIZE = 8
lr = 0.001
opt = tf.keras.optimizers.Adam(lr)
ver = 'model_self_attention_03' #save version
dropout=0.2 #dropout rate
hn = 'he_normal' #kernel initializer
tfrecords_read_dir = '/home/kevinteng/Desktop/ssd02/BraTS20_tfrecords05/'
stack_npy = "/home/kevinteng/Desktop/ssd02/BraTS2020_stack05/"
# ---
# # Helper Functions
# +
xent = tf.keras.losses.CategoricalCrossentropy()
def generalized_dice(y_true, y_pred, smooth = 1e-5):
"""
Generalized Dice Score
https://arxiv.org/pdf/1707.03237
https://github.com/Mehrdad-Noori/Brain-Tumor-Segmentation/blob/master/loss.py
"""
y_true = tf.reshape(y_true,shape=(-1,4))
y_pred = tf.reshape(y_pred,shape=(-1,4))
sum_p = tf.reduce_sum(y_pred, -2)
sum_r = tf.reduce_sum(y_true, -2)
sum_pr = tf.reduce_sum(y_true * y_pred, -2)
weights = tf.math.pow(tf.math.square(sum_r) + smooth, -1)
generalized_dice = (2 * tf.reduce_sum(weights * sum_pr)) / (tf.reduce_sum(weights * (sum_r + sum_p)))
return generalized_dice
def generalized_dice_loss(y_true, y_pred):
return 1-generalized_dice(y_true, y_pred)
def custom_loss(y_true, y_pred):
"""
The final loss function consists of the summation of two losses "GDL" and "CE"
with a regularization term.
"""
return generalized_dice_loss(y_true, y_pred) + 1.25 * xent(y_true, y_pred)
def data_aug(imgs):
choice = np.random.randint(0,4)
#no augmentation
if choice==0:
x = imgs
#flip up and down
if choice==1:
x = tf.image.flip_up_down(imgs)
#flip left and right
if choice==2:
x = tf.image.flip_left_right(imgs)
#rotation based on angle
if choice==3:
n_rot = np.random.randint(1,4)
x = tf.image.rot90(imgs, k=n_rot)
return x
# + [markdown] pycharm={"name": "#%% md\n"}
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Layer Names
# + pycharm={"name": "#%%\n"}
# template for guided attention block
layer_name_p01 = ['pam01_conv01', 'pam01_conv02', 'pam01_softmax', 'pam01_conv03',
'pam01_alpha','pam01_add']
layer_name_c01 = ['cam01_softmax', 'cam01_alpha','cam01_add']
layer_name_p02 = ['pam02_conv01', 'pam02_conv02', 'pam02_softmax', 'pam02_conv03',
'pam02_alpha', 'pam02_add']
layer_name_c02 = ['cam02_softmax', 'cam02_alpha','cam02_add']
layer_name_template = [layer_name_p01, layer_name_c01, layer_name_p02, layer_name_c02]
layer_name_ga = []
for b in range(1,4):
layer_block = []
for layer in layer_name_template:
layer_internal = [i+'block0{}'.format(b) for i in layer]
layer_block.append(layer_internal)
layer_name_ga.append(layer_block)
# -
# ----
# # Model
# +
from tensorflow.keras import Model
from tensorflow.keras.layers import Input, Conv2D, UpSampling2D, Activation, Add, Multiply, GaussianNoise
from tensorflow.keras.layers import SeparableConv2D, BatchNormalization, Dropout, concatenate
from tensorflow.keras.layers import GlobalAveragePooling2D, Dense, Layer, Softmax, ReLU, PReLU
from tensorflow_addons.layers import GroupNormalization
from utils_model import *
from attention import *
def conv_block_sep_v2(x, filters, layer_name, norm_fn='bn', kernel_size=(3, 3),
kernel_initializer='glorot_uniform', acti_fn='relu', dropout_rate=None):
'''
Dual convolution block with [full pre-activation], Norm -> Acti -> Conv
:param x: Input features
:param filters: A list that contains the number of filters for 1st and 2nd convolutional layer
:param layer_name: A list that contains the name for the 1st and 2nd convolutional layer
:param norm_fn: Tensorflow function for normalization, 'bn' for Batch Norm, 'gn' for Group Norm
:param kernel_size: Kernel size for both convolutional layer with 3x3 as default
:param kernel_initializer: Initializer for kernel weights with 'glorot uniform' as default
:param acti_fn: Tensorflow function for activation, 'relu' for ReLU, 'prelu' for PReLU
:param dropout_rate: Specify dropouts for layers
:return: Feature maps of same size as input with number of filters equivalent to the last layer
'''
assert type(filters)==list, "Please input filters of type list."
assert type(layer_name)==list, "Please input filters of type list."
assert acti_fn!= None, 'There should be an activation functino specified'
#1st convolutional block
if norm_fn=='bn':
x = BatchNormalization()(x)
elif norm_fn=='gn':
x = GroupNormalization()(x)
if acti_fn=='relu':
x = ReLU()(x)
elif acti_fn=='prelu':
x = PReLU(shared_axes=[1,2])(x)
if dropout_rate != None:
x = Dropout(dropout_rate)(x)
x = SeparableConv2D(filters[0], kernel_size, padding='same', kernel_initializer=kernel_initializer, name = layer_name[0])(x)
#2nd convolutional block
if norm_fn=='bn':
x = BatchNormalization()(x)
elif norm_fn=='gn':
x = GroupNormalization()(x)
if acti_fn=='relu':
x = ReLU()(x)
elif acti_fn=='prelu':
x = PReLU(shared_axes=[1,2])(x)
x = SeparableConv2D(filters[1], kernel_size, padding='same', kernel_initializer=kernel_initializer, name = layer_name[1])(x)
return x
def down_sampling_sep_v2(x, filters, layer_name, norm_fn='bn', kernel_size=(3, 3), acti_fn='relu',
kernel_initializer='glorot_uniform', dropout_rate=None, mode ='coord', x_dim=None, y_dim=None):
'''
Down sampling function version 2 with Convolutional layer of stride 2 as downsampling operation, with
[full pre-activation], Norm -> Acti -> Conv
:param x: Input features
:param filters: Number of filters for Convolutional layer of stride 2
:param layer_name: Layer name for convolutional layer
:param norm_fn: Tensorflow function for normalization, 'bn' for Batch Norm, 'gn' for Group Norm
:param kernel_size: Kernel size for both convolutional layer with 3x3 as default
:param acti_fn: Tensorflow function for activation, 'relu' for ReLU, 'prelu' for PReLU
:param kernel_initializer: Initializer for kernel weights with 'glorot uniform' as default
:param dropout_rate: Specify dropouts for layers
:param mode: 'coord' for Seperable Coord Conv, 'normal' for Seperable Conv
:param x_dim: x dimension for coord conv
:param y_dim: y dimension for coord conv
:return: Feature maps of size scaled down by 2 with number of filters specified
'''
assert mode=='coord' or mode=='normal', "Use 'coord' or 'normal' for mode!"
assert acti_fn!= None, 'There should be an activation functino specified'
if norm_fn=='bn':
x = BatchNormalization()(x)
elif norm_fn=='gn':
x = GroupNormalization()(x)
if acti_fn=='relu':
x = ReLU()(x)
elif acti_fn=='prelu':
x = PReLU(shared_axes=[1,2])(x)
if dropout_rate != None:
x = Dropout(dropout_rate)(x)
if mode=='coord':
#seperable coordconv
assert (x_dim!=None and y_dim!=None), "Please input dimension for CoordConv!"
x = Conv2D(1, kernel_size, strides=(2, 2), padding='same', kernel_initializer=kernel_initializer)(x)
x = CoordConv(x_dim=x_dim, y_dim=y_dim, with_r=False, filters=filters, strides=(1,1),
kernel_size = 3, padding='same', kernel_initializer=kernel_initializer, name=layer_name)(x)
else:
#normal mode
x = SeparableConv2D(filters, kernel_size, strides=(2, 2), padding='same', kernel_initializer=kernel_initializer, name=layer_name)(x)
return x
def res_block_sep_v2(x_in, filters, layer_name, norm_fn='gn', kernel_size=(3, 3),
kernel_initializer='glorot_uniform', acti_fn='prelu', dropout_rate=None):
assert len(filters)==2, "Please assure that there is 2 values for filters."
assert len(layer_name)==3, "Please assure that there is 3 values for layer name"
layer_name_conv = [layer_name[i] for i in range(len(layer_name)-1)]
output_conv_block = conv_block_sep_v2(x_in, filters, layer_name_conv, norm_fn=norm_fn, kernel_size=kernel_size,
kernel_initializer = kernel_initializer, acti_fn = acti_fn, dropout_rate=dropout_rate)
output_add = Add(name = layer_name[-1])([output_conv_block, x_in])
return output_add
def guided_attention_block(inp_feature, layer_name_p, layer_name_c):
'''
Guided attention block that takes feature as input and concatenates features
from PAM and CAM as output
:param inp_feature: Input features
:param layer_name_p: layer name list for PAM
:param layer_name_c: layer name list for CAM
:return: squeezed concatenated features of PAM and CAM
'''
pam_feature = PAM(inp_feature, layer_name_p, kernel_initializer=hn)
cam_feature = CAM(inp_feature, layer_name_c)
add = Add()([pam_feature,cam_feature]) #[60,60,128]
up = UpSampling2D(size=(4,4))(add) #[240,240,128]
squeeze = Conv2D(filters=64, kernel_size=1, padding='same', kernel_initializer=hn,
activation='relu')(up)
#pam and cam features
feature_pc = [pam_feature, cam_feature]
return squeeze, feature_pc
def guided_attention(res_feature, ms_feature, layer_name):
'''
Guided attention module
:param res_feature: Upsampled Feature maps from Res Block
:param ms_feature: Multi scale feature maps result from Res Block
:param layer_name: Layer Name should consist be a list contating 4 list
Example:
layer_name_p01 = ['pam01_conv01', 'pam01_conv02', 'pam01_softmax', 'pam01_conv03',
'pam01_alpha','pam01_add']
layer_name_c01 = ['cam01_softmax', 'cam01_alpha','cam01_add']
layer_name_p02 = ['pam02_conv01', 'pam02_conv02', 'pam02_softmax', 'pam02_conv03',
'pam02_alpha', 'pam02_add']
layer_name_c02 = ['cam02_softmax', 'cam02_alpha','cam02_add']
layer_name = [layer_name_p01, layer_name_c01, layer_name_p02, layer_name_c02]
:return: guided attention module with shape same as input
'''
assert len(layer_name)==4, "Layer name should be a list consisting 4 lists!"
#self attention block01
concat01 = concatenate([res_feature, ms_feature], axis=-1)
squeeze01, feature_pc01 = guided_attention_block(concat01, layer_name[0], layer_name[1])
multi01 = Multiply()([squeeze01, ms_feature])
#self attention block02
# concat02 = concatenate([multi01, res_feature],axis=-1)
# squeeze02 = guided_attention_block(concat02, layer_name[2], layer_name[3])
return multi01, feature_pc01
def forward(x):
'''
Resnet as backbone for multiscale feature retrieval.
Each resblock output(input signal), next resblock output(gated signal) is
feed into the gated attention for multi scale feature refinement.
Each gated attention output is pass through a bottle neck layer to standardize
the channel size by squashing them to desired filter size of 64.
The features are upsampled at each block to the corresponding [wxh] dimension
of w:240, h:240.
The upsampled features are concat and squash to corresponding channel size of 64
which yield multiscale feature.
:param x: batched images
:return: feature maps of each res block
'''
#inject noise
gauss1 = GaussianNoise(0.01)(x)
#retrieve input dimension
b,w,h,c = x.shape
#---- ResNet and Multiscale Features----
#1st block
conv01 = CoordConv(x_dim=w, y_dim=h, with_r=False, filters=64, strides=(1,1),
kernel_size = 3, padding='same', kernel_initializer=hn, name='conv01')(gauss1)
res_block01 = res_block_sep_v2(conv01, filters=[128, 64], layer_name=["conv02", "conv03", "add01"], dropout_rate=dropout)
#2nd block
down_01 = down_sampling_sep_v2(res_block01, filters=128, layer_name = 'down_01', kernel_initializer=hn,
mode='normal',x_dim=w//2, y_dim=w//2)
res_block02 = res_block_sep_v2(down_01, filters=[256, 128], layer_name=["conv04", "conv05", "add02"], dropout_rate=dropout)
#3rd block
down_02 = down_sampling_sep_v2(res_block02, filters=256, layer_name = 'down_02', kernel_initializer=hn,
mode='normal',x_dim=w//4, y_dim=h//4)
res_block03 = res_block_sep_v2(down_02, filters=[512, 256], layer_name=["conv06", "conv07", "add03"], dropout_rate=dropout)
#4th block
down_03 = down_sampling_sep_v2(res_block03, filters=512, layer_name = 'down_03', kernel_initializer=hn,
mode='normal',x_dim=w//8, y_dim=h//8)
res_block04 = res_block_sep_v2(down_03, filters=[1024, 512], layer_name=["conv08", "conv09", "add04"], dropout_rate=dropout)
# *apply activation function for the last output
res_block04 = PReLU(shared_axes=[1,2])(res_block04)
#grid attention blocks
att_block01, g_att01 = attention_block(res_block01,res_block02,64,'grid_att01')
att_block02, g_att02 = attention_block(res_block02,res_block03,128,'grid_att02')
att_block03, g_att03 = attention_block(res_block03, res_block04,256,'gird_att03')
#bottle neck => layer squash all attention block to same filter size 64
bottle01 = Conv2D(filters=64, kernel_size=1, padding='same', kernel_initializer=hn)(att_block01)
bottle02 = Conv2D(filters=64, kernel_size=1, padding='same', kernel_initializer=hn)(att_block02)
bottle03 = Conv2D(filters=64, kernel_size=1, padding='same', kernel_initializer=hn)(att_block03)
#upsampling for all layers to same (wxh) dimension=>240x240
up01 = bottle01 #[240,240,64]
up02 = UpSampling2D(size=(2, 2), interpolation='bilinear')(bottle02) #[120,120,64]=>[240,240,64]
up03 = UpSampling2D(size=(4,4), interpolation='bilinear')(bottle03) #[60,60,64]=>[240,240,64]
#multiscale features
concat_all = concatenate([up01,up02,up03],axis=-1) #[240,240,3*64]
#squeeze to have the same channel as upsampled features [240,240,3*64] => [240,240,64]
ms_feature = Conv2D(filters=64, kernel_size=1, padding='same', kernel_initializer=hn)(concat_all)
#Segmentations from multiscale features *without softmax activation
seg_01 = Conv2D(4, (1,1), name='seg_01')(up01)
seg_02 = Conv2D(4, (1,1), name='seg_02')(up02)
seg_03 = Conv2D(4, (1,1), name='seg_03')(up02)
#----self guided attention blocks-----
ga_01, f_pc01 = guided_attention(up01, ms_feature, layer_name_ga[0])
ga_02, f_pc02 = guided_attention(up02, ms_feature, layer_name_ga[1])
ga_03, f_pc03 = guided_attention(up03, ms_feature, layer_name_ga[2])
#Segmentations from guided attention features *without softmax activation
seg_ga01 = Conv2D(4, (1,1), name='seg_ga01')(ga_01)
seg_ga02 = Conv2D(4, (1,1), name='seg_ga02')(ga_02)
seg_ga03 = Conv2D(4, (1,1), name='seg_ga03')(ga_03)
#outputs for xent losses
output_xent = [seg_01, seg_02, seg_03, seg_ga01, seg_ga02, seg_ga03]
#output for dice coefficient loss
pred_seg = Add()(output_xent)
output_dice = Softmax()(pred_seg/len(output_xent))
#output for feature visualization
#gated attention
gated_attention = [g_att01, g_att02, g_att03]
#pam and cam features
f_pc = [f_pc01, f_pc02, f_pc03]
return output_xent, output_dice, gated_attention, f_pc
# + pycharm={"name": "#%%\n"}
#Build Model
input_layer = Input(shape=(200,200,4))
model = Model(input_layer, forward(input_layer))
# +
xent_logit = tf.keras.losses.CategoricalCrossentropy(from_logits=True)
@tf.function
def train_fn(image, label):
with tf.GradientTape() as tape:
output_xent, output_dice,_,_ = model(image, training=True)
loss_dice = generalized_dice_loss(label, output_dice)
loss_xents=[]
for seg in output_xent:
loss_xent = xent_logit(label, seg)
loss_xents.append(loss_xent)
loss_total = sum(loss_xents)+loss_dice
gradients = tape.gradient(loss_total, model.trainable_variables)
opt.apply_gradients(zip(gradients, model.trainable_variables))
return output_dice, loss_total, gradients
@tf.function
def val_fn(image, label):
model_output = model(image, training=False)
loss = custom_loss(label, model_output)
return model_output, loss
# -
# ---
epochs = 1
#list
loss_list = []
start_runtime = time.time()
while epochs <= max_epochs:
start = time.time()
print()
print("Epochs {:2d}".format(epochs))
steps = 1
ds = os.listdir(tfrecords_read_dir)
#shuffle directory list of tfrecords
shuffle = random.shuffle(ds)
loss_inner = []
for tf_re in ds:
tf_dir = os.path.join(tfrecords_read_dir+tf_re)
dataset = utils.parse_tfrecord(tf_dir).shuffle(SHUFFLE_BUFFER).batch(BATCH_SIZE)
for imgs in dataset:
#data augmentation
imgs = data_aug(imgs)
#crop images
image = imgs[:,20:220,20:220,:4]
#unprocessed label for plotting (cropped)
label = imgs[:,20:220,20:220,-1]
#for simplicity label 4 will be converted to 3 for sparse encoding
label = tf.where(label==4,3,label)
label = tf.keras.utils.to_categorical(label, num_classes=4)
#--------------<training function>----------------------------
img_seg, loss, gradients = train_fn(image, label)
#map from sparse to label
img_seg = tf.math.argmax(img_seg,-1,output_type=tf.int32)
label = tf.math.argmax(label,-1,output_type=tf.int32)
#accuracy of the output values for that batch
acc = tf.reduce_mean(tf.cast(tf.equal(img_seg,label), tf.float32))
#store lost for every steps
loss_inner.append(loss)
#save weights for every 5 epochs
if epochs%5==0:
model.save_weights('/home/kevinteng/Desktop/model_weights/model_{}.h5'.format(ver))
#output
if steps%5000==0:
input_img = [image[0,:,:,0], plot_labels_color(label[0]), plot_labels_color(img_seg[0])]
caption = ['Input Image', 'Ground Truth', 'Model Output']
plot_comparison(input_img, caption, n_col = 3, figsize=(10,10))
acc_stp = tf.reduce_mean(tf.cast(tf.equal(img_seg[0],label[0]), tf.float32))
dc_list_stp =compute_metric_dc(label[0],img_seg[0])
print("Steps: {}, Loss:{}".format(steps, loss))
print("Accurary: {}".format(acc_stp))
print("Seq: TC, ET, WT")
print("Dice coefficient: {}".format(dc_list_stp))
print("Gradient min:{}, max:{}".format(np.min(gradients[0]), np.max(gradients[0])))
steps+=1
loss_list.append(np.mean(loss_inner))
#end time per epochs
elapsed_time =(time.time()-start)/60 #unit in mins
print("Compute time per epochs: {:.2f} mins".format(elapsed_time))
epochs+=1
#end time for total epochs
elapsed_time_runtime = (time.time()-start_runtime)/60
print()
print('----------------------------------<END>---------------------------------')
print("Total run time for {} epochs: {:.2f} mins".format(epochs, elapsed_time_runtime))
# ---
# # Save Weights
model.save_weights('/home/kevinteng/Desktop/model_weights/model_{}.h5'.format(ver))
# ---
# # Validation
model.load_weights('/home/kevinteng/Desktop/model_weights/model_{}.h5'.format(ver))
def output_fn(image):
b,w,h,c = image.shape
model.trainable = False
_, model_output,_,_ = model(image)
# we need [240,240,155] to input into cloud validation
if w!=240:
#padding constant
p = int(240-w)//2
padding = tf.constant([[0,0],[p,p],[p,p],[0,0]]) #p=20
model_output = tf.pad(model_output, padding, "CONSTANT")
return model_output
ds = '/home/kevinteng/Desktop/ssd02/BraTS2020_preprocessed05/'
save_path = '/home/kevinteng/Desktop/ssd02/submission/'
actual_label = '/home/kevinteng/Desktop/ssd02/MICCAI_BraTS2020_TrainingData/BraTS20_Training_001/BraTS20_Training_001_seg.nii.gz'
#all brain affine are the same just pick one
brain_affine = nib.load(actual_label).affine
for train_or_val in sorted(os.listdir(ds)):
save_dir = save_path + train_or_val+'_'+ver
if not os.path.exists(save_dir):
os.makedirs(save_dir)
merge01 = os.path.join(ds+train_or_val)
for patient in sorted(os.listdir(merge01)):
patient_id = patient.split('.')[0]
merge02 = os.path.join(merge01,patient)
imgs = np.load(merge02)
image = imgs[:,20:220,20:220,:4]
seg_output = 0 #flush RAM
seg_output = np.zeros((240,240,155))
for i in range(image.shape[0]):
inp = tf.expand_dims(image[i],0)
img_seg = output_fn(inp) #validation function
#map from sparse to label
seg_output[:,:,i] = np.argmax(img_seg,-1)
#convert label from 4 to 3 and np array and cast as int
seg_output= np.where(seg_output==3,4,seg_output).astype(np.uint8)
prediction_ni = nib.Nifti1Image(seg_output, brain_affine)
prediction_ni.to_filename(save_dir+'/{}.nii.gz'.format(patient_id))
# ---
# # Model Summary
model.summary()
| Model/BraTS20_SGANet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="QjEYP4VMAJs_"
#
# # **Final Project**
# + [markdown] id="5qtYDGc6yJH4"
#
# Team members:
#
# * <NAME>
# * <NAME>
# * <NAME>
# * <NAME>
#
#
#
# + [markdown] id="BDcMnRNxAdE-"
# ## **Domain knowledge**
# + [markdown] id="6uvKOsk3-cHE"
#
#
# <p align="center">
# <img src="
# https://drive.google.com/uc?export=view&id=1CFPAD1qpdSpGBaEjNc1eQXSqjzuiAD5f">
#
# </p>
#
# **Mandoob Tuwaiq** is an app that provides delivery services. whether you want to send a gift to friends and family, or you want to increase your store customers and provide a delivery service for your product to them. Hence through the application, an integrated team of delivery drivers work to carry out the task of delivery in an easy and simple way.
#
# The aim of **Mandoob Tuwaiq** is to Increase drivers' income and provide a great delivery service for buyers and stores.
#
#
# + [markdown] id="mK1SumA1-sTb"
# **In this project, we focusing to help app founders to achieve their objectives by building a customer segmentation model. That's will help them to use different approaches for marketing.**
# <br> </br>
#
# We **thank** the founders of the application for giving us the opportunity to work on the data.
# + [markdown] id="3lJSMLgMjFbE"
# # **Dash Installation**
# + colab={"base_uri": "https://localhost:8080/"} id="e4tgtNYqiR1K" outputId="9090ec57-e851-4db2-f065-33dbd08108da"
#pip install --upgrade dash dash-core-components dash-html-components dash-renderer
# #!apt-get -qq install -y libfluidsynth1
# #!pip install matplotlib-venn
# !pip install jupyter-dash
# !pip install matplotlib-venn
# + colab={"base_uri": "https://localhost:8080/"} id="XL0qetVsiQFK" outputId="06e5a58e-bdd8-42ab-acdb-109f4eda8594"
pip install dash
# + colab={"base_uri": "https://localhost:8080/"} id="bjQujrQc808U" outputId="9ecb5c61-2837-4a69-fa25-9c4ccf5876f8"
pip install pandas
# + [markdown] id="_8vKj3xOqYMx"
# ## **Importing**
# + id="LuW_zknn8idL"
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# libraries for build dashboard
import dash
import dash_core_components as dcc
import dash_html_components as html
from dash.dependencies import Input, Output
import plotly.express as px
from jupyter_dash import JupyterDash
# Set the plot style to a seaborn theme
plt.style.use("seaborn")
sns.set_style('white')
#sns.set_palette("YlGn_r",4)
# + [markdown] id="5dJCJArS68d6"
# # **Loading Datasets**
#
# We creating dataFrame for each sheets from excel file for dataset with read data from drive csv file
#
# + id="hET0WNZs4fjB"
# read data from the drive csv file
##### Choose Orders Sheet #####
df = pd.read_excel(path, sheet_name='Orders')
# + id="0rbNTAFhHUqx"
# read data from the drive csv file
##### Choose Users Sheet #####
df_users = pd.read_excel(path, sheet_name='Users')
# + id="TOuGEp_tsumT"
# read data from the drive csv file
##### Choose OrderPayment Sheet #####
df_payment = pd.read_excel(path, sheet_name='OrderPayment')
# + [markdown] id="as-AaKwtjquT"
# ### **Drop Values**
# Drop Values from 'UserType' column
# + id="BOJKzgQnY8yG"
df_users.drop(df_users.index[df_users['UserType'] == 0], inplace = True)
# + colab={"base_uri": "https://localhost:8080/"} id="ccP9l4T4iZTh" outputId="ad9eb64f-6da4-47a0-ff0d-8ce5cf4070dc"
complate_orders = df[(~ df['DeliveryDateTime'].isna())& (df['Status'] == 6)]
complate_orders['hour'] = complate_orders['DeliveryDateTime'].dt.hour
complate_orders['date'] = complate_orders['DeliveryDateTime'].dt.date
complate_orders['day'] = complate_orders['DeliveryDateTime'].dt.day
complate_orders['month'] = complate_orders['DeliveryDateTime'].dt.month
# + [markdown] id="kaOIteAueN0F"
# #### Loading cluster **dataset**
# + colab={"base_uri": "https://localhost:8080/", "height": 149} id="AZQ3yh38uCER" outputId="150011fc-552e-43e5-e5c3-012a4ac268d8"
df_customer = pd.read_csv(path_)
df_customer.head(2)
# + [markdown] id="sKiLCDhqd9HW"
#
# # **Merge data**
# + id="AMtvrnxJ8Vft"
# rename the order id in the tow sheets order and payment ( the confirmed order only here)
df = df.rename(columns={"Code":"MandoubakOrder_Id"})
df_all = df.merge(df_payment, on='MandoubakOrder_Id')
# + [markdown] id="_0sbu1P86SZ3"
# ## **Preprocessing data**
# + [markdown] id="G7gsjDYX8eV5"
# ### **Check missing value :**
# + colab={"base_uri": "https://localhost:8080/"} id="8UCFR7UR8nho" outputId="d3e4d540-ff4e-4925-f399-3cd5f84a831c"
# missing values in data
df.isna().sum()
# + colab={"base_uri": "https://localhost:8080/"} id="BZVwugJJHPqS" outputId="6df9d196-f3f0-45d2-d9a2-c845baa83121"
# data type for each column
df.dtypes
# + [markdown] id="OrtQwlHOGyQn"
# ### Fill missing values
# + id="TM4LULPeGQ_s"
# DeliveryLocationId and ReceivingLocationId
# 0 mean the loaction is empty
df.DeliveryLocationId = df.DeliveryLocationId.fillna(0)
df.ReceivingLocationId = df.ReceivingLocationId.fillna(0)
# DeliveryDateTime and ReceivingDateTime
# N/A mean DateTime not availble
df.ReceivingDateTime = df.ReceivingDateTime.fillna('N/A')
df.DeliveryDateTime = df.DeliveryDateTime.fillna('N/A')
# DriverId
# 0 mean there isn't assigned Driver for this order
df.DriverId = df.DriverId.fillna(0)
# ProductDescription
df.ProductDescription = df.ProductDescription.fillna('Other')
# + colab={"base_uri": "https://localhost:8080/"} id="IEfi-wkPLbVx" outputId="b478a12e-de9b-4e55-943a-3f8a672120b0"
df.isnull().sum()
# + [markdown] id="MaJIAbbH6ZGO"
# ### Add new column for **Product Categorie**
# + colab={"base_uri": "https://localhost:8080/"} id="ptCQyRY3HJei" outputId="dfd71cac-08d5-49da-bf19-78b57f1bc563"
# Product Cat
# Gift and flowers, Food, Clothes, Accessories, Beauty and care, Other
df.ProductCategories.value_counts()
# + [markdown] id="kxJ0fj3nj8F3"
# ## **Converting columns from int To string,**
# ## **and changes variable**
# + [markdown] id="0wlnJTHe-emy"
# Convert 'AccountActive' columns To string, and changes Values
# + id="romYK_5P98Kb"
# Convert columns To string
df_users.AccountActive = df_users.AccountActive.astype(str)
# changes Values
df_users.AccountActive = df_users.AccountActive.replace({'-1': "Active Account"})
df_users.AccountActive = df_users.AccountActive.replace({'0': "inactive Account "})
# + [markdown] id="L6ZnQ8fq7fBk"
# Convert UserType columns To string, and changes Values
# + id="-D7YzWAZ87hJ"
# Convert columns To string
df_users.UserType =df_users.UserType.astype(str)
# changes Values
df_users.UserType = df_users.UserType.replace({'1': "Store"})
df_users.UserType = df_users.UserType.replace({'2': "Buyer"})
df_users.UserType = df_users.UserType.replace({'3': "Driver"})
# + [markdown] id="LPhcnxl8woSh"
# Convert Cluster to string
# + id="DEdlzx2SpS05"
df_customer.pred_cluster_DBSCAN =df_customer.pred_cluster_DBSCAN.astype(str)
df_customer.pred_cluster_DBSCAN =df_customer.pred_cluster_DBSCAN.replace({'-1': "cluster 1"})
df_customer.pred_cluster_DBSCAN =df_customer.pred_cluster_DBSCAN.replace({'0': "cluster 2"})
df_customer.pred_cluster_DBSCAN =df_customer.pred_cluster_DBSCAN.replace({'1': "cluster 3"})
df_customer.pred_cluster_DBSCAN =df_customer.pred_cluster_DBSCAN.replace({'2': "cluster 4"})
df_customer.pred_cluster_DBSCAN =df_customer.pred_cluster_DBSCAN.replace({'3': "cluster 5"})
# + [markdown] id="tVA0l6xH8tHj"
# Convert 'Status' columns to string, changes Values
#
# + id="BqMeBZD-8_XE"
# Convert columns To string
df.Status =df.Status.astype(str)
# changes Values
df.Status = df.Status.replace({'1': "New"})
df.Status = df.Status.replace({'2': "confirmed"})
df.Status = df.Status.replace({'3': "Reserved"})
df.Status = df.Status.replace({'4': "Delivering"})
df.Status = df.Status.replace({'6': "Confirmed_Delivery "})
df.Status = df.Status.replace({'7': "Cancelled"})
df.Status = df.Status.replace({'8': " Closed"})
df.Status= df.Status.replace({'10': "Expired"})
# + [markdown] id="5RsxbhuqVA-a"
# Rename Id colum in df User
# + colab={"base_uri": "https://localhost:8080/", "height": 422} id="eQ8C2_G6UO5a" outputId="89c43e57-9871-464c-c1d4-a0777c35bb93"
df_users = df_users.rename(columns={'Id': 'UserId'})
df_users.head(2)
# + [markdown] id="6AjnM1hAx0w0"
# ### Count registred Users per month
# + id="25herSImiVxN"
df_users['Registration_Month'] = [i for i in df_users.CreatedDate.dt.month]
df_users['Registration_Day'] = [i for i in df_users.CreatedDate.dt.day]
df_users['Registration_Year'] = [i for i in df_users.CreatedDate.dt.year]
# + [markdown] id="SYzEAn_59wBq"
# # **Dashboard**
#
# -Visual at least 6 plots show insights from data (either static or interactive)
# Note: If we just create 6 plots, all of them should be related to the target.
#
# -Think about important insights related to users (buyers and stores), and can add insights related to orders if we plot more than 6 plots.
#
# -Determine which tools will be use for build dashboard
#
# -Make sure to use different types of plot (scatter, bar, count, hist, line, pie …)
#
# -Write a short description about each plot. These plots should answer the business questions.
#
# + colab={"base_uri": "https://localhost:8080/"} id="C_EPktgRin4Z" outputId="4ba0ec88-0803-4ceb-ef11-69eaca646c26"
# !pip install matplotlib-venn
# + id="zpkVtwWbFL-_"
import dash
import dash_core_components as dcc
import dash_html_components as html
import pandas as pd
import plotly.graph_objs as go
import random
sns.set_palette("PuBuGn_r",4) # Choose Color
# + [markdown] id="vfH17OU_ygcJ"
# # The **dashboard**
# + colab={"base_uri": "https://localhost:8080/", "height": 671} id="p8NYsaauJ4tR" outputId="c425fd99-1386-4188-f4ca-b9b4eeec6fc7"
# Instatiate the App
app = JupyterDash(__name__ )
# Create layout that will hold all visuals
app.layout = html.Div([
html.H1(
'Mandoob Tuwaiq app',
style={'color':'#FFFFFF','background-color': '#148F77','text-align':'center','padding':10}
), # Change colors and style
###########################
################ Input 1
html.Div([
html.Label([
"Choose Product Category ", # Label title
# create Dropdown list
dcc.Dropdown(
id = 'ProductCategories', # Dropdown id
clearable = False,
value = "Food", # default value
style={'color': '#0B5345'},
# return the unique values from ProductCategories column
options = [{"label":x, "value":x} for x in df.ProductCategories.unique()]
)
],
style={'fontSize':18}), # change the options size
# create a graph
dcc.Graph(id='graph1'),
],
# add style for the div
style={'width': '47%', 'float': 'left', 'display': 'inline-block',
'background-color': '#F3F3F3','margin-right':1,'padding':10,'margin-top':1}),
###########################
############ Input 2
html.Div([
html.Label([ ## Create Sliider
"Month",
dcc.Slider(
id='Registration_Month',
min=1,
max=6,
value=2,
#style='color': '#67001F',
marks={ # Changes Monthe from int to text
1: {"label": "Jan"},
2: {"label": "Feb"},
3: {"label": "March"},
4: {"label": "April"},
5: {"label": "May"},
6: {"label": "Jun"},
}
),
]),
html.Br([]),
#style={'fontSize':18}, # change the options size
# create a graph
dcc.Graph(id='graph2'),
],
# add style for the div
style={'width': '47%', 'float': 'right', 'display': 'inline-block',
'background-color': '#F3F3F3','margin-left':1,'padding':10,'margin-top':1}),
###########################
########### Input 3
html.Div([
html.Label([
"Chose Cluster from DBSCAN", # Label title
# create Dropdown list
dcc.RadioItems(
id = 'pred_cluster_DBSCAN', # Dropdown id
value = "cluster 2", # default value
style={'color': '#000000'},
# return the unique values from brand_name column
options = [{"label":x, "value":x} for x in df_customer.pred_cluster_DBSCAN.unique()]
)
],
style={'fontSize':18}), # change the options size
# create a graph
dcc.Graph(id='graph3'),
],
# add style for the div
style={'width': '47%', 'float': 'left', 'display': 'inline-block',
'background-color': '#F3F3F3','margin-right':1,'padding':10,'margin-top':1}),
###########################
########## Input 4
html.Div([
html.Label([
"Chose Cluster from k-mean", # Label title
# create Dropdown list
dcc.Dropdown(
id = 'pred_cluster', # Dropdown id
clearable = False,
value = 3, # default value
style={'color': '#0B5345'},
# return the unique values from ProductCategories column
options = [{"label":x, "value":x} for x in df_customer.pred_cluster.unique()]
)
],
style={'fontSize':18}), # change the options size
# create a graph
dcc.Graph(id='graph4'),
],
# add style for the div
style={'width': '47%', 'float': 'left', 'display': 'inline-block',
'background-color': '#F3F3F3','margin-right':1,'padding':10,'margin-top':1}),
###########################
########## Input 5
html.Div([
html.Label([
"Who's Uesre?", # Label title
# create Dropdown list
dcc.RadioItems(
id = 'UserType', # Dropdown id
value = "Store", # default value
style={'color': '#000000'},
# return the unique values from brand_name column
options = [{"label":x, "value":x} for x in df_users.UserType.unique()]
)
],
style={'fontSize':18}), # change the options size
#style={'fontSize':18}, # change the options size
# create a graph
dcc.Graph(id='graph5'),
],
# add style for the div
style={'width': '47%', 'float': 'left', 'display': 'inline-block',
'background-color': '#F3F3F3','margin-right':1,'padding':10,'margin-top':1}),
###########################
###### Input 6
html.Div([
# Create slider
html.Label([
"OrdersMonth",
dcc.Slider(
id='orders_month',
min=2,
max=6,
value=3,
marks={ # Changes Monthe from int to text
2: {"label": "Feb"},
3: {"label": "March"},
4: {"label": "April"},
5: {"label": "May"},
6: {"label": "Jun"},
}
),
]),
html.Br([]),
# create a graph
dcc.Graph(id='graph6'),
],
# add style for the div
style={'width': '47%', 'float': 'righ', 'display': 'inline-block',
'background-color': '#F3F3F3','margin-left':1,'padding':10,'margin-top':1}),
])
###########################
###### 1st plot Call Back
@app.callback(
Output('graph1','figure'),
Input('ProductCategories','value')
)
def update_figures(ProductCategories):
# filter data by category
test = df_all[df_all['ProductCategories']== ProductCategories]
# create Pie plot for PaymentMode shows anothers Payment methods
fig = px.pie(test, test['PaymentMode'],color_discrete_sequence=px.colors.sequential.PuBuGn_r)
fig.update_layout(title_text='Most Payment Methods') # add title
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# return plot
return fig
###########################
##### 2nd plot Call Back
@app.callback(
Output('graph2','figure'),
Input('Registration_Month','value')
)
def update_figures(month):
df_regestertion = df_users[(df_users['UserType']=="Buyer") & ( df_users['Registration_Month'] == month)]['CreatedDate'].dt.date.value_counts().sort_index()
fig = px.line(df_regestertion,
x= df_regestertion.index,
y= df_regestertion.values,
color_discrete_sequence=px.colors.sequential.PuBuGn_r,
labels = {"x":"Date","y":"Count of registred user"},)
fig.update_layout(title_text='Number of registred user')
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# return plot
return fig
###########################
############ 3rd plot Call back
@app.callback(
Output('graph3','figure'),
Input('pred_cluster_DBSCAN','value')
)
def update_figures(pred_cluster_DBSCAN_chosen):
dfDBSCAN=df_customer[(df_customer['pred_cluster_DBSCAN']==pred_cluster_DBSCAN_chosen)]
fig = px.box(dfDBSCAN,
x=dfDBSCAN["Number_of_Orders"],
y=dfDBSCAN["Total_Payment"],
color_discrete_sequence=px.colors.sequential.PuBuGn_r,
)
fig.update_layout(title_text='DBSCAN cluter')
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# return plot
return fig
###########################
###### 4 plot Call Back
@app.callback(
Output('graph4','figure'),
Input('pred_cluster','value')
)
def update_figures(pred_cluster_chosen):
dfKmean=df_customer[(df_customer['pred_cluster']==pred_cluster_chosen)]
fig = px.scatter(dfKmean,
x=dfKmean["Number_of_Orders"],
y=dfKmean["Total_Payment"],
color_discrete_sequence=px.colors.sequential.PuBuGn_r,
)
fig.update_layout(title_text='k-mean cluter')
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# return plot
return fig
###########################
###### 5 plot Call Back
@app.callback(
Output('graph5','figure'),
Input('UserType','value')
)
def update_figures(UserTypeChosen):
dfUserType=df_users[(df_users['UserType']==UserTypeChosen)]
fig = px.bar(dfUserType,
x=dfUserType["AccountActive"].value_counts().index,
y=dfUserType["AccountActive"].value_counts().values,
# hue='UserActive',
color_discrete_sequence=px.colors.sequential.PuBuGn_r,
labels = {"x":"Account","y":"Count of Account Active "},)
fig.update_layout(title_text='Account Active')
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#000000', # change the plot title color
})
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#000000', # change the plot title color
})
# return plot
return fig
###########################
# 6 plot Call Back
@app.callback(
Output('graph6','figure'),
Input('orders_month','value')
)
def update_figures(month):
complate_orders_ = complate_orders[(complate_orders['month'] == month)].pivot_table(index='hour',columns='day',aggfunc='count')['Id']
complate_orders_ = complate_orders_.reindex(
index=range(24),
columns=range(1,max(complate_orders[complate_orders['month'] == month].day)),
fill_value=np.nan
)
fig = go.Figure(data=go.Heatmap(
z = complate_orders_.values,
y = complate_orders_.index,
x = complate_orders_.columns,
xgap = 3 ,
ygap = 3,
colorscale =px.colors.sequential.Mint,
hoverongaps = True))
fig.update_traces(
hovertemplate="<br>".join([
"Hour: %{y}",
"Day: %{x}",
"Number of completed orders: %{z}",
])
)
fig.update_layout(title_text='Number of completed orders')
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# transparent background
fig.update_layout({
'plot_bgcolor': 'rgba(0, 0, 0, 0)', # change plot background color
'paper_bgcolor': 'rgba(0, 0, 0, 0)', # change paper background color
'title_font_color': '#0B5345', # change the plot title color
})
# return plot
return fig
# Run the App Server
app.run_server(mode="inline")
# + [markdown] id="DDQRM5bccI1h"
# ## **Insights**
#
# - We created Pie chart to Shows The most Payment methods based on Product category. Is shown when choose product category that you want see most payment method use for it.
# - Line chart shows Timeline for each months and see what that day have more New Registration in 2021. you should Select Month to see what the most day have more new Registration in the month.
# - The scatter graph shows the relationship between Number of Orders and Total Payment based on k-mean cluster, User Chose Cluster from k-mean to see the result based on the entry.
# - The box graph shows the relationship between Number of Orders and Total Payment based on DBSCAN cluster, User Chose Cluster from DBSCAN to see the result based on the entry.
# - The Last graph shows the Number of completed orders for each months, User Chose Month from 2021 to see the result based on the entry.
# - About bar chart is Show us the status of the account, whether it is active or inactive, is shown when selecting the user type If the user Driver or Buyer or Store.
#
# -
| dashboard.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# +
import os
import sys
import cv2
import glob
import numpy as np
import tensorflow as tf
from sklearn.model_selection import train_test_split
EPOCHS = 10
IMG_WIDTH = 30
IMG_HEIGHT = 30
NUM_CATEGORIES = 43
TEST_SIZE = 0.4
# -
data_dir = 'gtsrb'
images = []
labels = []
for i in range(NUM_CATEGORIES):
for path in glob.glob(f'{data_dir}/{i}/*.ppm'):
img = cv2.imread(path)
re_img = cv2.resize(img, (IMG_HEIGHT, IMG_WIDTH))
images.append(re_img)
labels.append(i)
labels = tf.keras.utils.to_categorical(labels)
x_train, x_test, y_train, y_test = train_test_split(np.array(images), np.array(labels), test_size=TEST_SIZE)
model = tf.keras.models.Sequential([
tf.keras.layers.Conv2D(32, (3, 3), activation="relu", input_shape=(IMG_WIDTH, IMG_HEIGHT, 3)),
tf.keras.layers.MaxPooling2D(pool_size=(2, 2)),
tf.keras.layers.Conv2D(64, (3, 3), activation="relu"),
tf.keras.layers.MaxPooling2D(pool_size=(2, 2)),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(128, activation="relu"),
tf.keras.layers.Dense(64, activation="relu"),
tf.keras.layers.Dropout(0.33),
tf.keras.layers.Dense(NUM_CATEGORIES, activation="softmax")
])
model.compile(
optimizer="adam",
loss="categorical_crossentropy",
metrics=["accuracy"]
)
model.fit(x_train, y_train, epochs=EPOCHS)
model.evaluate(x_test, y_test, verbose=2)
model.evaluate(x_test, y_test, verbose=2)
| 5_neural_networks/traffic/traffic_test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <font style="font-size:6pt"> <i>
# All of these python notebooks are available at https://gitlab.erc.monash.edu.au/andrease/Python4Maths.git </i>
# </font>
# # Getting started
#
# Python can be used like a calculator. Simply type in expressions to get them evaluated.
#
# ## Basic syntax for statements
# The basic rules for writing simple statments and expressions in Python are:
# * No spaces or tab characters allowed at the start of a statement: Indentation plays a special role in Python (see the section on control statements). For now simply ensure that all statements start at the beginning of the line.
# * The '#' character indicates that the rest of the line is a comment
# * Statements finish at the end of the line:
# * Except when there is an open bracket or paranthesis:
# ```python
# 1+2
# +3 #illegal continuation of the sum
# (1+2
# + 3) # perfectly OK even with spaces
# ```
# * A single backslash at the end of the line can also be used to indicate that a statement is still incomplete
# ```python
# 1 + \
# 2 + 3 # this is also OK
# ```
# The jupyter notebook system for writting Python intersperses text (like this) with Python statements. Try typing something into the cell (box) below and press the 'Run' button above to execute it. We will discuss operators that can be used in such operations further below, but for numbers these are mostly fairly obvious.
1 + 2 * 3
# Python has extensive help built in. You can execute `help()` for an overview or `help(x)` for any library, object or type `x`. Try using `help("topics")` to get a list of help pages built into the help system.
help("topics")
# ## Variables & Values
# A name that is used to denote something or a value is called a variable. In python, variables can be declared and values can be assigned to it as follows,
x = 2 # anything after a '#' is a comment
y = 5
xy = 'Hey'
print(x+y, xy) # not really necessary as the last value in a bit of code is displayed by default
# Multiple variables can be assigned with the same value.
x = y = 1
print(x,y)
# The basic types build into Python include `float` (floating point numbers), `int` (integers), `str` (unicode character strings) and `bool` (boolean). Some examples of each:
2.0 # a simple floating point number
1e100 # a googol as floating point number
-1234567890 # an integer
True or False # the two possible boolean values
'This is a string'
"It's another string"
print("""Triple quotes (also with '''), allow strings to break over multiple lines.
Alternatively \n is a newline character (\t for tab, \\ is a single backslash)""")
# Python also has complex numbers that can be written as follows. Note that the brackets are required.
complex(1,2)
(1.0+2j) # the same number as above
# ## Operators
# ### Arithmetic Operators
# | Symbol | Task Performed |
# |----|---|
# | + | Addition |
# | - | Subtraction |
# | / | Division |
# | // | Integer division |
# | % | Modulus (remainder) |
# | * | Multiplication |
# | ** | Exponentiation (power) |
#
# As expected these operations generally promote to the most general type of any of the numbers involved i.e. int -> float -> complex.
1+2.0
3-1
2 * (3+0j) * 1.0
3/4
# In many languages (and older versions of python) 1/2 = 0 (truncated division). In Python 3 this behaviour is captured by a separate operator that rounds down: (ie a // b$=\lfloor \frac{a}{b}\rfloor$)
3//4.0
15%10
# Python natively allows (nearly) infinite length integers while floating point numbers are double precision numbers:
11**300
11.0**300
# ### Relational Operators
# | Symbol | Task Performed |
# |----| :--- |
# | == | True, if it is equal |
# | != | True, if not equal to |
# | < | less than |
# | > | greater than |
# | <= | less than or equal to |
# | >= | greater than or equal to |
#
# Note the difference between `==` (equality test) and `=` (assignment)
z = 2
z == 2
z > 2
# Comparisons can also be chained in the mathematically obvious way. The following will work as expected in Python (but not in other languages like C/C++):
0.5 < z <= 1
# ### Boolean and Bitwise Operators
# |Operator|Meaning | \| | Symbol | Task Performed |
# |----|--- | - |----|---|
# |`and`| Logical and | \| | & | Bitwise And |
# |`or` | Logical or | \| | $\mid$ | Bitwise OR |
# |`not` | Not | \| | ~ | Negate |
# | | | \| | ^ | Exclusive or |
# | | | \| | >> | Right shift |
# | | | \| | << | Left shift |
#
a = 2 #binary: 10
b = 3 #binary: 11
print('a & b =',a & b,"=",bin(a&b))
print('a | b =',a | b,"=",bin(a|b))
print('a ^ b =',a ^ b,"=",bin(a^b))
print('b << a =',b<<a,"=",bin(b<<a))
print( not (True and False), "==", not True or not False)
# ### Assignment operators
#
# The binary operators can be combined with assignment to modify a variable value.
# For example:
x = 1
x += 2 # add 2 to x
print("x is",x)
x <<= 2 # left shift by 2 (equivalent to x *= 4)
print('x is',x)
x **= 2 # x := x^2
print('x is',x)
# # Built-in Functions
# Python comes with a wide range of functions. However many of these are part of stanard libraries like the `math` library rather than built-in.
# ## Converting values
#
# Conversion from hexadecimal to decimal is done by adding prefix **0x** to the hexadecimal value or vice versa by using built in `hex( )`, Octal to decimal by adding prefix **0** to the octal value or vice versa by using built in function `oct( )`.
hex(171) # hexadecmial value as string
0xAB
# `int( )` converts a number to an integer. This can be a single floating point number, integer or a string. For strings the base can optionally be specified:
print(int(7.7), int('111',2),int('7'))
# Similarly, the function `str( )` can be used to convert almost anything to a string
print(str(True),str(1.2345678),str(-2))
# ## Mathematical functions
# Mathematical functions include the usual suspects like logarithms, trigonometric fuctions, the constant $\pi$ and so on.
import math
math.sin(math.pi/2)
from math import * # avoid having to put a math. in front of every mathematical function
sin(pi/2) # equivalent to the statement above
# ## Simplifying Arithmetic Operations
# `round( )` function rounds the input value to a specified number of places or to the nearest integer.
print( round(5.6231) )
print( round(4.55892, 2) )
# `abs( )` provides the absolute value of any number (including the magnitude of a complex number).
c =complex('5+2j')
print("|5+2i| =", abs(c) , "\t |-5| =", abs(-5) )
# `divmod(x,y)` outputs the quotient and the remainder in a tuple (you will be learning about tuples in the further chapters) in the format (quotient, remainder).
divmod(9,2)
# ## Accepting User Inputs
# `input(prompt)`, prompts for and returns input as a string. A useful function to use in conjunction with this is `eval()` which takes a string and evaluates it as a python expression.
#
# *Note:* In notebooks it is often easier just to modify the code than to prompt for input.
abc = input("abc = ")
abcValue=eval(abc)
print(abc,'=',abcValue)
| Intro-to-Python/01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1>Previsão de receita por pessoa com base no Censo</h1>
# Neste notebook contém um projeto desenvolvido durante o curso "Machine Learning e Data Science com Python de A a Z" da instituição Udemy.
#
# OBJETIVO DA ANÁLISE:
# - Comparar algoritmos de Machine Learning que possam prever se a receita ultrapassa 50 mil/ano com base nos dados do censo.
# Também conhecido como conjunto de dados "Census Income".
# - > 50K, <= 50K.
#
# Informações do conjunto de dados:
# - Data set pode ser encontrado no link: http://archive.ics.uci.edu/ml/datasets/Adult
# - A extração foi feita por <NAME> do banco de dados do Censo de 1994.
# - A tarefa de previsão é determinar se uma pessoa ganha mais de 50K por ano.
#
# Algoritmos de Classificação utilizados:
# - Naive Bayes
# - Árvore de Decisão
# - Random Forest
# - KNN
# - Regressão Logística
# - SVM
# - Redes Neurais Artificiais
#
import warnings
warnings.filterwarnings('ignore')
# <h1>Importando a base de dados</h1>
import pandas as pd
# importando a base de dados
# para uma melhor compreensão, será alterado o nome das colunas
base = pd.read_csv('census.csv', names = ['idade', 'tipo_emprego', 'caracteristica', 'educacao', 'anos_estudo', \
'estado_civil', 'ocupacao', 'parentesco', 'raça', 'sexo', 'ganho_capital', \
'perda_capital','hrs_trabalhada_semana', 'pais_origem', 'renda_anual'], header = 0)
base.head(10)
# <h1>Explorando os dados</h1>
# verificando o tipo de dados de cada coluna
base.dtypes
# quantidade de linhas e colunas do dataFrame
base.shape
print("O dataset tem {} linhas e {} colunas".format(base.shape[0], base.shape[1]))
# utilizando a função Info para obter um resumo sobre o dataset
base.info()
# <h1>Atributos previsores e classe</h1>
# criando uma variável para armazenar os atributos previsores
previsores = base.iloc[:, 0:14].values
previsores
# criando uma variável para armazenar o atributo classe
classe = base.iloc[:, 14].values
classe
# <h1>Transformação de variáveis categóricas</h1>
# LabelEncoder para transformar as variáveis categóricas em numéricas
from sklearn.preprocessing import LabelEncoder
# +
# transformando as variáveis categóricas dos atributos previsores
labelencoder_previsores = LabelEncoder()
previsores[:, 1] = labelencoder_previsores.fit_transform(previsores[:, 1])
previsores[:, 3] = labelencoder_previsores.fit_transform(previsores[:, 3])
previsores[:, 5] = labelencoder_previsores.fit_transform(previsores[:, 5])
previsores[:, 6] = labelencoder_previsores.fit_transform(previsores[:, 6])
previsores[:, 7] = labelencoder_previsores.fit_transform(previsores[:, 7])
previsores[:, 8] = labelencoder_previsores.fit_transform(previsores[:, 8])
previsores[:, 9] = labelencoder_previsores.fit_transform(previsores[:, 9])
previsores[:, 13] = labelencoder_previsores.fit_transform(previsores[:, 13])
# -
previsores
# transformando a variável categórica do atributo classe
labelencoder_classe = LabelEncoder()
classe = labelencoder_classe.fit_transform(classe)
# <= 50K tem o valor = 0
# > 50K tem o valor = 1
classe
# <h1>Escalonamento dos atributos</h1>
# +
# Aplicar uma fórmula para os atributos na mesma escala
# Utilizando a Padronização x= x - média(x) / desvioPadrão(x)
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
previsores = scaler.fit_transform(previsores)
# -
previsores
# <h1>Spliting</h1>
# Dividindo as variáveis em treino e teste
# 85% dados para treino e 15% dados para teste
from sklearn.model_selection import train_test_split
previsores_treinamento, previsores_teste, classe_treinamento, classe_teste = train_test_split(previsores, classe, \
test_size=0.15, random_state=0)
# Imprimindo os resultados
print("{0:0.2f}% nos dados de treino".format((len(previsores_treinamento)/len(base.index)) * 100))
print("{0:0.2f}% nos dados de teste".format((len(previsores_teste)/len(base.index)) * 100))
len(previsores_treinamento)
len(classe_treinamento)
len(previsores_teste)
len(classe_teste)
# <h1>Naive Bayes</h1>
#
# O algoritmo “Naive Bayes” é um classificador probabilístico.
# Ele recebe o nome de “naive” (ingênuo) porque desconsidera a correlação entre as variáveis (features).
#
# Classificação de textos, filtragem de SPAM e análise de sentimento em redes sociais são algumas das muitas aplicações para esse algoritmo.
# O algoritmo é muito robusto para previsões em tempo real, pois precisa de poucos dados para realizar a classificação. Se existe a necessidade de correlacionar fatores, esse algoritmo tende a falhar nas predições.
# ### Construindo e treinando o modelo
# Importando o algoritmo
from sklearn.naive_bayes import GaussianNB
# Criando o modelo preditivo
classificador = GaussianNB()
# Treinando o modelo
classificador.fit(previsores_treinamento, classe_treinamento)
# testando o modelo
previsoes = classificador.predict(previsores_teste)
# ### Verificando a exatidão no modelo
# precisão do modelo
from sklearn.metrics import confusion_matrix, accuracy_score
precisao = accuracy_score(classe_teste, previsoes)
precisao
# matriz de confusão
# diagonal principal (quantidade de acertos)
matriz = confusion_matrix(classe_teste, previsoes)
matriz
# <h1>Árvore de Decisão</h1>
#
# Uma árvore de decisão utiliza base de dados histórica e geralmente começa com um único nó, que se divide em possíveis resultados. Cada um desses resultados leva a nós adicionais, que se ramificam em outras possibilidades. Assim, cria-se uma forma de árvore. Esses algoritmos são considerados um dos melhores e mais utilizados métodos de aprendizagem supervisionada, pois nos dão modelos preditivos de alta precisão, estabilidade e facilidade de interpretação. Ao contrário dos modelos lineares, eles mapeiam muito bem relações não-lineares.
# ### Construindo e treinando o modelo
# Importando o algoritmo
from sklearn.tree import DecisionTreeClassifier
# Criando o modelo preditivo
classificador = DecisionTreeClassifier(criterion = 'entropy', random_state = 0)
# Treinando o modelo
classificador.fit(previsores_treinamento, classe_treinamento)
# testando o modelo
previsoes = classificador.predict(previsores_teste)
# ### Verificando a exatidão no modelo
# precisão do modelo
precisao = accuracy_score(classe_teste, previsoes)
precisao
# matriz de confusão
# diagonal principal (quantidade de acertos)
matriz = confusion_matrix(classe_teste, previsoes)
matriz
# <h1>Random Forest</h1>
#
# Esse algoritmo irá criar muitas árvores de decisão, de maneira aleatória, formando o que podemos enxergar como uma floresta, onde cada árvore será utilizada na escolha do resultado final. É um método de aprendizagem de máquina versátil e capaz de executar tarefas de regressão e de classificação. Ele também aplica métodos de redução dimensional, trata valores faltantes, valores anómalos (‘outliers’) e outras etapas essenciais da exploração de dados.
# É um tipo de método de aprendizado de ‘ensemble’, onde um grupo de modelos fracos são combinados para formar um modelo mais forte.
# ### Construindo e treinando o modelo
# Importando o algoritmo
from sklearn.ensemble import RandomForestClassifier
# Criando o modelo preditivo
classificador = RandomForestClassifier(n_estimators = 40, criterion = 'entropy', random_state = 0)
# Treinando o modelo
classificador.fit(previsores_treinamento, classe_treinamento)
# testando o modelo
previsoes = classificador.predict(previsores_teste)
# ### Verificando a exatidão no modelo
# precisão do modelo
precisao = accuracy_score(classe_teste, previsoes)
precisao
# matriz de confusão
# diagonal principal (quantidade de acertos)
matriz = confusion_matrix(classe_teste, previsoes)
matriz
# <h1>KNN</h1>
#
# O KNN (K-Nearest Neighbor) tem o objetivo de determinar a qual grupo uma determinada amostra vai pertencer com base nas amostras vizinhas. Os exemplos de treinamento são armazenados e a previsão é feita somente quando um novo registro precisa ser classificado. Diferente dos outros algoritmos, ele não constrói um modelo, apenas faz o cálculo da distância.
# ### Construindo e treinando o modelo
# Importando o algoritmo
from sklearn.neighbors import KNeighborsClassifier
classificador = KNeighborsClassifier(n_neighbors = 5, metric = 'minkowski', p = 2)
classificador.fit(previsores_treinamento, classe_treinamento)
# testando
previsoes = classificador.predict(previsores_teste)
# ### Verificando a exatidão
# precisão
precisao = accuracy_score(classe_teste, previsoes)
precisao
# matriz de confusão
# diagonal principal (quantidade de acertos)
matriz = confusion_matrix(classe_teste, previsoes)
matriz
# <h1>Regressão Logística</h1>
#
# Esse algoritmo mede a relação entre a variável dependente categórica e uma ou mais variáveis independentes, estimando as probabilidades usando uma função logística. Analisa diferentes aspectos ou variáveis de um objeto para depois determinar uma classe na qual ele se encaixa melhor.
# ### Construindo e treinando o modelo
# Importando o algoritmo
from sklearn.linear_model import LogisticRegression
# Criando o modelo preditivo
classificador = LogisticRegression(solver='lbfgs')
# Treinando o modelo
classificador.fit(previsores_treinamento, classe_treinamento)
# testando o modelo
previsoes = classificador.predict(previsores_teste)
# ### Verificando a exatidão no modelo
# precisão do modelo
precisao = accuracy_score(classe_teste, previsoes)
precisao
# matriz de confusão
# diagonal principal (quantidade de acertos)
matriz = confusion_matrix(classe_teste, previsoes)
matriz
# <h1>SVM - Máquinas de Vetores de Suporte</h1>
#
# Uma máquina de vetores de suporte (SVM) desenvolve o modelo tomando as entradas de treinamento, mapeando elas no espaço multidimensional e utilizando regressão para encontrar um hiperplano (um hiperplano é uma superfície em espaço de n dimensões que o separa em duas metades de espaço) que melhor separa duas classes de entradas. Uma vez que esse modelo tenha sido treinada, ele é capaz de avaliar novas entradas em relação ao hiperplano divisor e classificá-las em uma entre duas categorias.
# ### Construindo e treinando o modelo
# Importando o algoritmo
from sklearn.svm import SVC
# Criando o modelo preditivo
classificador = SVC(kernel = 'linear', random_state = 1)
# Treinando o modelo
classificador.fit(previsores_treinamento, classe_treinamento)
# testando o modelo
previsoes = classificador.predict(previsores_teste)
# ### Verificando a exatidão no modelo
# precisão do modelo
precisao = accuracy_score(classe_teste, previsoes)
precisao
# matriz de confusão
# diagonal principal (quantidade de acertos)
matriz = confusion_matrix(classe_teste, previsoes)
matriz
# <h1>Redes Neurais Artificiais</h1>
#
# Redes Neurais Artificiais são técnicas computacionais que apresentam um modelo matemático inspirado na estrutura neural de organismos inteligentes e que adquirem conhecimento através da experiência. As redes neurais são compostas por várias unidades de processamento. Mesmo sendo usadas para resolverem problemas complexos, será utilizado como um exemplo.
# ## Redes Neurais com sklearn
# ### Construindo e treinando o modelo
# Importando o algoritmo
from sklearn.neural_network import MLPClassifier
# Criando o modelo preditivo
classificador = MLPClassifier(verbose=True, max_iter=1000, tol= 0.000010)
# Treinando o modelo
classificador.fit(previsores_treinamento, classe_treinamento)
# testando o modelo
previsoes = classificador.predict(previsores_teste)
# ### Verificando a exatidão no modelo
# precisão do modelo
precisao = accuracy_score(classe_teste, previsoes)
precisao
# matriz de confusão
# diagonal principal (quantidade de acertos)
matriz = confusion_matrix(classe_teste, previsoes)
matriz
# ## Redes Neurais com Keras
# ### Construindo e treinando o modelo
from keras.models import Sequential
from keras.layers import Dense
# Criando o modelo preditivo
classificador = Sequential()
# Criação das camadas ocultas e de saída
classificador.add(Dense(units = 8, activation = 'relu', input_dim = 14))
classificador.add(Dense(units = 8, activation = 'relu'))
classificador.add(Dense(units = 1, activation = 'sigmoid'))
# Compilando a rede neural
classificador.compile(optimizer = 'adam', loss = 'binary_crossentropy', metrics = ['accuracy'])
# Treinando o modelo
classificador.fit(previsores_treinamento, classe_treinamento, batch_size = 10, epochs = 100)
# testando o modelo
previsoes = classificador.predict(previsores_teste)
previsoes = (previsoes > 0.5)
# ### Verificando a exatidão no modelo
# precisão do modelo
precisao = accuracy_score(classe_teste, previsoes)
precisao
# matriz de confusão
# diagonal principal (quantidade de acertos)
matriz = confusion_matrix(classe_teste, previsoes)
matriz
# <h1>Conclusão</h1>
#
# Realizando apenas um teste podemos observar que os Algoritmos de Redes Neurais Artificiais e Random Forest tiveram um maior percentual de acerto nessa base de dados. Isso não significa que esses algoritmos sejam os melhores a serem utilizados para prever a renda anual por pessoa dessa base de dados. Para uma melhor compreensão desse resultado, deve ser feito mais testes a fim de comprovar a eficácia desses algoritmos.
| previsao_receita_censo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 3_Modifying_an_idf
# ## Intro
#
# This notebook uses Eppy to read and modify an idf file.
#
# The '1ZoneUncontrolled.idf' is read and the orientation changed to 0, 90, 180 and 270.
# Each variant is saved as a separate file.
#
# This demo uses the following idf file, which is available in the ExampleFiles in the EnergyPlus installation.
with open('1ZoneUncontrolled.idf','r') as f:
print(f.read())
# ## Set up eppy
from eppy.modeleditor import IDF
iddfile = r'C:\EnergyPlusV8-9-0\Energy+.idd'
IDF.setiddname(iddfile)
# ## Create an instance and read in an .idf file
idf=IDF()
idf.idfname='1ZoneUncontrolled.idf'
idf.read()
idf
# ## Modify the idf instance and save with new filenames
orientations=[0,90,180,270]
building_object=idf.idfobjects['BUILDING'][0]
for orientation in orientations:
idf.idfobjects['BUILDING'][0].North_Axis=orientation
fp='1ZoneUncontrolled_{}.idf'.format(orientation)
idf.save(fp)
# ## Check it has worked
orientations=[0,90,180,270]
for orientation in orientations:
fp='1ZoneUncontrolled_{}.idf'.format(orientation)
idf=IDF(fp)
print('file:',fp,'\n',idf.idfobjects['BUILDING'][0])
| demo/3_Modifying_an_idf/3_Modifying_an_idf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MAT281 - Laboratorio N°01
# <a id='p1'></a>
#
# ## Problema 01
#
# ### a) Calcular el número $\pi$
#
# En los siglos XVII y XVIII, <NAME> y <NAME> descubrieron una serie infinita que sirve para calcular $\pi$:
#
# $$\displaystyle \pi = 4 \sum_{k=1}^{\infty}\dfrac{(-1)^{k+1}}{2k-1} = 4(1-\dfrac{1}{3}+\dfrac{1}{5}-\dfrac{1}{7} + ...) $$
#
# Desarolle un programa para estimar el valor de $\pi$ ocupando el método de Leibniz, donde la entrada del programa debe ser un número entero $n$ que indique cuántos términos de la suma se utilizará.
#
#
# * **Ejemplo**: *calcular_pi(3)* = 3.466666666666667, *calcular_pi(1000)* = 3.140592653839794
#
# ### Definir Función
#
def calcular_pi(n:int)->float:
"""
calcular_pi(n)
Aproximacion del valor de pi mediante el método de Leibniz
Parameters
----------
n : int
Numero de terminos.
Returns
-------
output : float
Valor aproximado de pi.
Examples
--------
>>> calcular_pi(3)
3.466666666666667
>>> calcular_pi(1000)
3.140592653839794
"""
pi = 0 # valor incial
for k in range(1,n+1):
numerador = (-1)**(k+1) # numerador de la iteracion i
denominador = 2*k-1 # denominador de la iteracion i
pi+=numerador/denominador # suma hasta el i-esimo termino
return 4*pi
# Acceso a la documentación
help(calcular_pi)
# ### Verificar ejemplos
# ejemplo 01
assert calcular_pi(3) == 3.466666666666667, "ejemplo 01 incorrecto"
calcular_pi(3.0)
# ejemplo 02
assert calcular_pi(1000) == 3.140592653839794, "ejemplo 02 incorrecto"
# **Observación**:
#
# * Note que si corre la línea de comando `calcular_pi(3.0)` le mandará un error ... ¿ por qué ?
# * En los laboratorio, no se pide ser tan meticuloso con la documentacion.
# * Lo primero es definir el código, correr los ejemplos y luego documentar correctamente.
# calcular_pi (3.0) manda error ya que la funcion esta definida en int (entero) y 3.0 es float (real)
# ### b) Calcular el número $e$
#
# Euler realizó varios aportes en relación a $e$, pero no fue hasta 1748 cuando publicó su **Introductio in analysin infinitorum** que dio un tratamiento definitivo a las ideas sobre $e$. Allí mostró que:
#
#
# En los siglos XVII y XVIII, <NAME> y <NAME> descubrieron una serie infinita que sirve para calcular π:
#
# $$\displaystyle e = \sum_{k=0}^{\infty}\dfrac{1}{k!} = 1+\dfrac{1}{2!}+\dfrac{1}{3!}+\dfrac{1}{4!} + ... $$
#
# Desarolle un programa para estimar el valor de $e$ ocupando el método de Euler, donde la entrada del programa debe ser un número entero $n$ que indique cuántos términos de la suma se utilizará.
#
#
# * **Ejemplo**: *calcular_e(3)* =2.5, *calcular_e(1000)* = 2.7182818284590455
# ### Definir función
def fac (x:int)-> int:
if x==1:
return 1
else:
return (x * fac(x-1)) #definición clásica
def calcular_e (n:int)-> float:
sum=0
for i in range (1,n):
sum=sum+(1/fac(i))
return (sum+1)
calcular_e(3)
calcular_e(1000)
# ### Verificar ejemplos
# ejemplo 01
assert calcular_e(3) == 2.5, "ejemplo 01 incorrecto"
# ejemplo 02
assert calcular_e(1000) == 2.7182818284590455, "ejemplo 02 incorrecto"
# <a id='p2'></a>
#
# ## Problema 02
#
#
# Sea $\sigma(n)$ definido como la suma de los divisores propios de $n$ (números menores que n que se dividen en $n$).
#
# Los [números amigos](https://en.wikipedia.org/wiki/Amicable_numbers) son enteros positivos $n_1$ y $n_2$ tales que la suma de los divisores propios de uno es igual al otro número y viceversa, es decir, $\sigma(n_1)=\sigma(n_2)$ y $\sigma(n_2)=\sigma(n_1)$.
#
#
# Por ejemplo, los números 220 y 284 son números amigos.
# * los divisores propios de 220 son 1, 2, 4, 5, 10, 11, 20, 22, 44, 55 y 110; por lo tanto $\sigma(220) = 284$.
# * los divisores propios de 284 son 1, 2, 4, 71 y 142; entonces $\sigma(284) = 220$.
#
#
# Implemente una función llamada `amigos` cuyo input sean dos números naturales $n_1$ y $n_2$, cuyo output sea verifique si los números son amigos o no.
#
# * **Ejemplo**: *amigos(220,284)* = True, *amigos(6,5)* = False
#
# ### Definir Función
def amigos(a:int,b:int) ->bool:
sum_a=0
sum_b=0
for i in range(1,a):
if a%i==0:
sum_a+=i
for j in range(1,b):
if b%j==0:
sum_b+=j
if sum_a==b and sum_b==a:
return ("True")
else:
return ("False")
amigos(220,284)
amigos(6,5)
# ### Verificar ejemplos
# ejemplo 01
assert amigos(220,284) == True, "ejemplo 01 incorrecto"
# ejemplo 02
assert amigos(6,5) == False, "ejemplo 02 incorrecto"
# <a id='p3'></a>
#
# ## Problema 03
#
# La [conjetura de Collatz](https://en.wikipedia.org/wiki/Collatz_conjecture), conocida también como conjetura $3n+1$ o conjetura de Ulam (entre otros nombres), fue enunciada por el matemático <NAME> en 1937, y a la fecha no se ha resuelto.
#
# Sea la siguiente operación, aplicable a cualquier número entero positivo:
# * Si el número es par, se divide entre 2.
# * Si el número es impar, se multiplica por 3 y se suma 1.
#
# La conjetura dice que siempre alcanzaremos el 1 (y por tanto el ciclo 4, 2, 1) para cualquier número con el que comencemos.
#
# Implemente una función llamada `collatz` cuyo input sea un número natural positivo $N$ y como output devulva la secuencia de números hasta llegar a 1.
#
# * **Ejemplo**: *collatz(9)* = [9, 28, 14, 7, 22, 11, 34, 17, 52, 26, 13, 40, 20, 10, 5, 16, 8, 4, 2, 1]
# ### Definir Función
def collatz(n:int)->list:
if n<1:
return ("el número debe ser positivo")
else:
lista=[n]
while n!=1:
if n%2==0:
n=n/2
else:
n=3*n+1
lista.append(n)
return(lista)
collatz(9)
# ### Verificar ejemplos
# ejemplo 01
assert collatz(9) == [9, 28, 14, 7, 22, 11, 34, 17, 52, 26, 13, 40, 20, 10, 5, 16, 8, 4, 2, 1], "ejemplo 01 incorrecto"
# <a id='p4'></a>
#
# ## Problema 04
#
# La [conjetura de Goldbach](https://en.wikipedia.org/wiki/Goldbach%27s_conjecture) es uno de los problemas abiertos más antiguos en matemáticas. Concretamente, <NAME>, en 1921, en su famoso discurso pronunciado en la Sociedad Matemática de Copenhague, comentó que probablemente la conjetura de Goldbach no es solo uno de los problemas no resueltos más difíciles de la teoría de números, sino de todas las matemáticas. Su enunciado es el siguiente:
#
# $$\textrm{Todo número par mayor que 2 puede escribirse como suma de dos números primos - <NAME> (1742)}$$
#
# Implemente una función llamada `goldbach` cuyo input sea un número natural positivo $N$ y como output devuelva la suma de dos primos ($N1$ y $N2$) tal que: $N1+N2=N$.
#
# * **Ejemplo**: goldbash(4) = (2,2), goldbash(6) = (3,3) , goldbash(8) = (3,5)
# ### Definir función
def primo(n:int)->bool:
if n==1:
return("1 no es primo")
else:
for i in range(2,n):
if n%i==0:
return("False")
return("True")
def primo_men(n:int)->list:
if n==1:
return("1 no considera primo")
else:
lista=[]
for i in range(2,n):
if primo(i)=="True":
lista.append(i)
return(lista)
def goldbach (n:int)->tuple:
if n<=2: return ("error, ingrese número mayor a 2")
elif n%2!=0: return("error, ingrese número par")
else:
lista=primo_men(n)
for i in lista:
for j in lista:
if i+j==n:
return((i,j))
goldbach(4)
goldbach(6)
goldbach(8)
# ### Verificar ejemplos
# ejemplo 01
assert goldbash(4) == (2,2), "ejemplo 01 incorrecto"
# ejemplo 02
assert goldbash(6) == (3,3), "ejemplo 02 incorrecto"
# ejemplo 03
assert goldbash(8) == (3,5), "ejemplo 03 incorrecto"
| labs/lab_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # RMSprop with `Gluon`
#
#
# +
import mxnet as mx
from mxnet import autograd
from mxnet import gluon
from mxnet import ndarray as nd
import numpy as np
import random
mx.random.seed(1)
random.seed(1)
# Generate data.
num_inputs = 2
num_examples = 1000
true_w = [2, -3.4]
true_b = 4.2
X = nd.random_normal(scale=1, shape=(num_examples, num_inputs))
y = true_w[0] * X[:, 0] + true_w[1] * X[:, 1] + true_b
y += .01 * nd.random_normal(scale=1, shape=y.shape)
dataset = gluon.data.ArrayDataset(X, y)
net = gluon.nn.Sequential()
net.add(gluon.nn.Dense(1))
square_loss = gluon.loss.L2Loss()
# +
# %matplotlib inline
import matplotlib as mpl
mpl.rcParams['figure.dpi']= 120
import matplotlib.pyplot as plt
def train(batch_size, lr, gamma, epochs, period):
assert period >= batch_size and period % batch_size == 0
net.collect_params().initialize(mx.init.Normal(sigma=1), force_reinit=True)
# RMSProp.
trainer = gluon.Trainer(net.collect_params(), 'rmsprop',
{'learning_rate': lr, 'gamma1': gamma})
data_iter = gluon.data.DataLoader(dataset, batch_size, shuffle=True)
total_loss = [np.mean(square_loss(net(X), y).asnumpy())]
for epoch in range(1, epochs + 1):
for batch_i, (data, label) in enumerate(data_iter):
with autograd.record():
output = net(data)
loss = square_loss(output, label)
loss.backward()
trainer.step(batch_size)
if batch_i * batch_size % period == 0:
total_loss.append(np.mean(square_loss(net(X), y).asnumpy()))
print("Batch size %d, Learning rate %f, Epoch %d, loss %.4e" %
(batch_size, trainer.learning_rate, epoch, total_loss[-1]))
print('w:', np.reshape(net[0].weight.data().asnumpy(), (1, -1)),
'b:', net[0].bias.data().asnumpy()[0], '\n')
x_axis = np.linspace(0, epochs, len(total_loss), endpoint=True)
plt.semilogy(x_axis, total_loss)
plt.xlabel('epoch')
plt.ylabel('loss')
plt.show()
# -
train(batch_size=10, lr=0.03, gamma=0.9, epochs=3, period=10)
# ## Next
# [AdaDalta from scratch](../chapter06_optimization/adadelta-scratch.ipynb)
# For whinges or inquiries, [open an issue on GitHub.](https://github.com/zackchase/mxnet-the-straight-dope)
| chapter06_optimization/rmsprop-gluon.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# Chain: **define-by-run** scheme
# ---
import numpy as np
import chainer
from chainer import cuda, Function, gradient_check, report, training, utils, Variable
from chainer import datasets, iterators, optimizers, serializers
from chainer import Link, Chain, ChainList
import chainer.functions as F
import chainer.links as L
from chainer.training import extensions
# +
x_data = np.array([5], dtype=np.float32)
x= Variable(x_data)
y= x**2 - 2*x + 1
# -
# y is also Variable
y.data
# error propagation
y.backward()
# gradient is computed and stored in `grad`
x.grad
z = 2*x
y = x**2 - z + 1
# set return_grad = True
y.backward(retain_grad=True)
z.grad
z = 2*x
y = x**2 - z + 1
# did no set return_grad, no
y.backward()
z.grad is None
x = Variable(np.array([[1, 2, 3], [4, 5, 6]], dtype=np.float32))
y = x**2 - 2*x + 1
y.grad = np.ones((2, 3), dtype=np.float32)
y.backward()
x.grad
# links
# ---
#from 3dim to 2dim
# f = x.dot(W.T) + b
f = L.Linear(3,2)
print('f.W.data.shape', f.W.data.shape)
print("f.b.data:", f.b.data)
x = Variable(np.array([[1, 2, 3], [4, 5, 6]], dtype=np.float32))
y = f(x)
y.data
# Example MINIST
# ---
train, test = datasets.get_mnist()
train_iter = iterators.SerialIterator(train, batch_size=100, shuffle=True)
# repeat=False, which means we stop iteration when all examples are visited. This option is usually required for the test/validation datasets
test_iter = iterators.SerialIterator(test, batch_size=100, repeat=False, shuffle=False)
class MLP(Chain):
def __init__(self, n_units, n_out):
super(MLP, self).__init__()
with self.init_scope():
# the size of the inputs to each layer will be inferred
self.l1 = L.Linear(None, n_units) # n_in -> n_units
self.l2 = L.Linear(None, n_units) # n_units -> n_units
self.l3 = L.Linear(None, n_out) # n_units -> n_out
def __call__(self, x):
h1 = F.relu(self.l1(x))
h2 = F.relu(self.l2(h1))
y = self.l3(h2)
return y
model = L.Classifier(MLP(100, 10)) # the input size, 784, is inferred
optimizer = optimizers.SGD()
optimizer.setup(model)
updater = training.StandardUpdater(train_iter, optimizer)
trainer = training.Trainer(updater, (20, 'epoch'), out='result')
trainer.run()
trainer.extend(extensions.Evaluator(test_iter, model))
trainer.extend(extensions.LogReport())
trainer.extend(extensions.PrintReport(['epoch', 'main/accuracy', 'validation/main/accuracy']))
trainer.extend(extensions.ProgressBar())
trainer.run()
# +
#RNN
import numpy as np
import chainer.links as L
from chainer import Variable, Chain, optimizers
l = L.LSTM(100, 50)
l.reset_state()
x = Variable(np.random.rand(10, 100).astype(np.float32))
y = l(x)
class RNN(Chain):
def __init__(self):
super(RNN, self).__init__()
with self.init_scope():
self.embed = L.EmbedID(1000, 100) # word embedding
self.mid = L.LSTM(100, 50) # 1st LSTM layer
self.out = L.Linear(50, 1000) # deef-forward output layer
def reset_state(self):
self.mid.reset_state()
def __call__(self,cur_word):
# Given the current word ID, predict the next word
x = self.embed(cur_word)
h = self.mid(x)
y = self.mid(h)
return y
rnn = RNN()
model = L.Classifier(rnn)
optimizer = optimizers.SGD()
optimizer.setup(model)
| Chainer_notes/Chainer_basic_tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Advanced: Iterating over genomes with PatMatch
#
# The [previous notebook, 'Advanced: Sending PatMatch output directly to Python'](Sending%20PatMatch%20output%20directly%20to%20Python.ipynb),
# covered leveraging the the Jupyter environment to skip over needing to save a file to actually pass results from shell scripts into Python. This notebook will demonstrate using one of those approaches to iterate over several genomes.
# This builds on other aspects seen in the early notebooks, too.
#
# ## Preparing
#
# Similar to the previous notebook, in order to insure everything is all set, act as if this is a new session in this Jupyter environment, and run the next cell so that you can start stepping through the preparation steps by first getting a sequence file. Plus, you'll get the files for scripts to convert it to dataframe and plot sites across a chromosome and import the main functions of those scripts.
#
# Repeating these steps if you had already done so this session will cause no harm, and so go ahead and run this cell.
# !curl -O http://sgd-archive.yeastgenome.org/sequence/S288C_reference/chromosomes/fasta/chrmt.fsa
# !curl -O https://raw.githubusercontent.com/fomightez/sequencework/master/patmatch-utilities/patmatch_results_to_df.py
from patmatch_results_to_df import patmatch_results_to_df
# !curl -O https://raw.githubusercontent.com/fomightez/sequencework/master/plot_sites/plot_sites_position_across_chromosome.py
from plot_sites_position_across_chromosome import plot_sites_position_across_chromosome
# Additionally, the sequences of two other mitochondrial genomes will be retrieved.
#
# Reference for the additional sequence data:
# - [Contrasting evolutionary genome dynamics between domesticated and wild yeasts.
# <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. Nat Genet. 2017 Jun;49(6):913-924. doi: 10.1038/ng.3847. Epub 2017 Apr 17. PMID: 28416820](https://www.ncbi.nlm.nih.gov/pubmed/28416820)
# Prepare for getting PacBio (Yue et al 2017 sequences)
#make a list of the strain designations
yue_et_al_strains = ["N44","UFRJ50816"]
# Get & unpack the genome sequences from strains
import os
genomes = []
expected_resulting_file = "N44_mito.genome.fsa"
if not os.path.isfile(expected_resulting_file):
for s in yue_et_al_strains:
# !curl -OL http://yjx1217.github.io/Yeast_PacBio_2016/data/Mitochondrial_Genome/{s}.mt.genome.fa.gz
# !gunzip -f {s}.mt.genome.fa.gz
# rename the files to follow the convention used for SGD reference
# !mv {s}.mt.genome.fa {s}_mito.genome.fsa
genomes.append(s+"_mito.genome.fsa")
# Add identifiers to each description line so results for each strain clear later. The reference from the Saccharomyces Genome database will be tagged 'SGD_REFmito'.
# +
# %%capture
import sys
import os
# add identifiers to each description line so results for each strain clear later
def add_strain_id_to_description_line(file,strain_id):
'''
Takes a file and edits every description line to add
strain_id after the caret.
Saves the fixed file
'''
import sys
output_file_name = "temp.txt"
# prepare output file for saving so it will be open and ready
with open(output_file_name, 'w') as output_file:
# read in the input file
with open(file, 'r') as input_handler:
# prepare to give feeback later or allow skipping to certain start
lines_processed = 0
for line in input_handler:
lines_processed += 1
if line.startswith(">"):
rest_o_line = line.split(">")
new_line = ">"+strain_id +"\n"
else:
new_line = line
# Send text to output
output_file.write(new_line)
# replace the original file with edited
# !mv temp.txt {file}
# Feedback
sys.stderr.write("\n{} has had identifiers added.".format(file))
files_tagged = 0
for g in genomes:
add_strain_id_to_description_line(g, g.split('.genome.fsa')[0])
files_tagged += 1
# Feedback
sys.stderr.write("\n{} sets of strain identifiers added.".format(files_tagged))
# Edit the description line for the SGD reference to be clear
# !sed -i '1s/.*/>SGD_REF_mito/' chrmt.fsa
# -
# Make a list of the genomes based on the name.
fn_pat_to_check = ".fsa"
genomes = []
import os
import fnmatch
for file in os.listdir('.'):
if fnmatch.fnmatch(file, '*'+fn_pat_to_check):
genomes.append(file)
genomes
# ## Iteraring over the genomes with PatMatch searching for sequence patterns
# %%capture
import pandas as pd
promoter_pattern = "DDWDWTAWAAGTARTADDDD"
dfs = []
for seq_file in genomes:
# !perl ../patmatch_1.2/unjustify_fasta.pl {seq_file}
# output = !perl ../patmatch_1.2/patmatch.pl -c {promoter_pattern} {seq_file+".prepared"}
# !rm {seq_file+".prepared"}
df_pat = patmatch_results_to_df(output.n, pattern=promoter_pattern, name="promoter")
typical_file_suffix = "_mito.genome.fsa"
if typical_file_suffix in seq_file:
strain_info = seq_file.split(typical_file_suffix)[0]
else:
strain_info = "SGD_REF"
df_pat["strain"] = strain_info
cols = df_pat.columns.tolist()
n = int(cols.index('strain'))
cols = [cols[n]] + cols[:n] + cols[n+1:]
df_pat = df_pat[cols]
dfs.append(df_pat)
df = pd.concat(dfs)
# See the previous advanced notebook, [Using brackets or other strange characters to make complex patterns on command line or with Python](Using%20brackets%20or%20other%20strange%20characters%20to%20make%20complex%20patterns%20on%20command%20line%20or%20with%20Python.ipynb), if your pattern is complex and involves brackets as you'll need to edit the following line in the above code:
#
# ```python
# # output = !perl ../patmatch_1.2/patmatch.pl -c {promoter_pattern} {seq_file+".prepared"}`
# ```
#
# Because `%%capture` is used in the above cell to stop the output from accumulating to a large size when many genomes are analyzed, the results are checked in the cells following.
len(df)
for dataf in dfs:
print(len(dataf))
# A more informative way to look at the number of hits per strain.
df.sort_values('hit_number', ascending=False, inplace=True)
largest_hit_num_by_id_df = df.groupby('FASTA_id').head(1)
largest_hit_num_by_id_df = largest_hit_num_by_id_df.groupby('strain').head(1).reset_index(drop=True)
largest_hit_num_by_id_df
# Visualize each genomes positions with x-axis matching particular genome:
# (Have to do each in separate cell or keeps overlaying past points the current way the plot script works.)
dataf = dfs[0]
dataf = dataf.rename(columns={'hit_id':'sys_gene_id'})
plot_sites_position_across_chromosome(dataf);
print (dataf.FASTA_id[0] + " Plot:");
dataf = dfs[1]
dataf = dataf.rename(columns={'hit_id':'sys_gene_id'})
plot_sites_position_across_chromosome(dataf);
print (dataf.FASTA_id[0] + " Plot:");
dataf = dfs[2]
dataf = dataf.rename(columns={'hit_id':'sys_gene_id'})
plot_sites_position_across_chromosome(dataf);
print (dataf.FASTA_id[0] + " Plot:");
# Note the different x-axis spanned in each of the three plots.
#
# -----
#
# Enjoy!
| notebooks/Iterating over genomes with PatMatch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %load_ext autoreload
# %autoreload 2
# %pylab inline
import jax
import jax.numpy as np
import tensorflow_probability as tfp; tfp = tfp.experimental.substrates.jax
tfd = tfp.distributions
from nsec.samplers import ScoreHamiltonianMonteCarlo, ScoreMetropolisAdjustedLangevinAlgorithm
from nsec.datasets.two_moons import get_two_moons
from nsec.tempered_sampling import TemperedMC
# +
from functools import partial
def logp(x, sigma):
return get_two_moons(sigma+0.02).log_prob(x).squeeze()
dlogp = jax.jit(jax.grad(logp))
# -
def score_fn(x, sigma):
@partial(jax.vmap, in_axes=(0,0))
def inner_fn(x, sigma):
return dlogp(x, sigma)
return inner_fn(x, sigma)
logp(np.zeros([10,1,2]).astype('float32'),
np.ones([10,1,1]).astype('float32')).shape
score_fn(np.zeros([10,2]).astype('float32'),
np.ones([10]).astype('float32')).shape
def make_kernel_fn(target_log_prob_fn, target_score_fn):
return ScoreHamiltonianMonteCarlo(
target_log_prob_fn=target_log_prob_fn,
target_score_fn=target_score_fn,
step_size=0.01,
num_leapfrog_steps=3,
num_delta_logp_steps=4)
tmc = TemperedMC(
target_score_fn=score_fn,
inverse_temperatures=np.ones([10]),
make_kernel_fn=make_kernel_fn,
gamma=0.7,
min_steps_per_temp=20,
num_delta_logp_steps=4)
num_results = int(1e3)
num_burnin_steps = int(1)
samples, trace = tfp.mcmc.sample_chain(
num_results=num_results,
current_state=np.zeros([10,2])+randn(10,2),
kernel=tmc,
num_burnin_steps=num_burnin_steps,
#trace_fn=lambda _, pkr: pkr.pre_swap_replica_results.is_accepted,
seed=jax.random.PRNGKey(0))
trace.post_tempering_inverse_temperatures.shape
trace.pre_tempering_results.is_accepted
plot(trace.post_tempering_inverse_temperatures[:,0])
plot(trace.post_tempering_inverse_temperatures[:,1])
plot(trace.post_tempering_inverse_temperatures[:,2])
plot(np.exp(trace.tempering_log_accept_ratio[:,0]))
samples.shape
hist2d(samples[:,0,0],samples[:,0,1],128,
range=[[-1.1,2.1],[-0.6,1.1]],cmap='gist_stern');
scatter(samples[:,0,0],samples[:,0,1], c=log10(trace.post_tempering_inverse_temperatures[:,0])); colorbar()
hist2d(samples[:,1,0],samples[:,1,1],128,
range=[[-1.1,2.1],[-0.6,1.1]],cmap='gist_stern');
scatter(samples[:,1,0],samples[:,1,1], c=log10(trace.post_tempering_inverse_temperatures[:,1])); colorbar()
hist2d(samples[:,2,0],samples[:,2,1],128,
range=[[-1.1,2.1],[-0.6,1.1]],cmap='gist_stern');
hist2d(samples[:,:,0].flatten(),samples[:,:,1].flatten(),128,
range=[[-1.1,2.1],[-0.6,1.1]],cmap='gist_stern');
hist2d(samples[:,0],samples[:,1],128,
range=[[-1.1,2.1],[-0.6,1.1]],cmap='gist_stern');
hist2d(samples[:,0],samples[:,1],128,
range=[[-1.1,2.1],[-0.6,1.1]],cmap='gist_stern');
hist2d(samples[:,0],samples[:,1],64,
range=[[-1.1,2.1],[-0.6,1.1]],);
def make_replica_target_log_prob_fn(log_prob_fn, inverse_temp):
def fn(x):
y = log_prob_fn(x) * inverse_temp
return fn
def make_kernel_fn(target_log_prob_fn, target_score_fn):
return ScoreHamiltonianMonteCarlo(
target_log_prob_fn=target_log_prob_fn,
target_score_fn=target_score_fn,
step_size=step_size,
num_leapfrog_steps=3,
num_delta_logp_steps=4)
remc = ReplicaExchangeMC(
target_score_fn=score_fn,
inverse_temperatures=sigmas,
make_kernel_fn=make_kernel_fn,
num_delta_logp_steps=32)
def trace_swaps(unused_state, results):
return (results.is_swap_proposed_adjacent,
results.is_swap_accepted_adjacent)
num_results = int(1e4)
num_burnin_steps = int(1e2)
samples, trace = tfp.mcmc.sample_chain(
num_results=num_results,
current_state=np.zeros([1,2]),
kernel=remc,
num_burnin_steps=num_burnin_steps,
trace_fn=lambda _, pkr: pkr.pre_swap_replica_results.is_accepted,
seed=jax.random.PRNGKey(0))
[len(where(trace[:,i,0])[0]) for i in range(4)]
hist2d(samples[:,0,0],samples[:,0,1],64,
range=[[-1.1,2.1],[-0.6,1.1]],);
hist2d(samples[:,0,0],samples[:,0,1],64,
range=[[-1.1,2.1],[-0.6,1.1]],);
dist_target=get_two_moons(sigmas[0]+0.02)
samps = dist_target.sample(10000, seed=jax.random.PRNGKey(0))
hist2d(samps[:,0],samps[:,1],64,
range=[[-1.1,2.1],[-0.6,1.1]],);
# And just to compare, what would have happened if we only ran a normal
# HMC in the same conditions
hmc = tfp.mcmc.HamiltonianMonteCarlo(
target_log_prob_fn=dist_target.log_prob,
num_leapfrog_steps=3,
step_size=0.0125)
samples_hmc, is_accepted = tfp.mcmc.sample_chain(
num_results=num_results,
num_burnin_steps=num_burnin_steps,
current_state=np.zeros([1,2]),
kernel=hmc,
trace_fn=lambda _, pkr: pkr.is_accepted,
seed=jax.random.PRNGKey(0))
hist2d(samples_hmc[:,0,0],samples_hmc[:,0,1],64,
range=[[-1.1,2.1],[-0.6,1.1]],);
| notebooks/TemperedScoreMC.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: R
# language: R
# name: ir
# ---
# # H2O Use Case - Predictive Maintenance
#
# - Source: https://archive.ics.uci.edu/ml/datasets/SECOM
# - H2O Basics: train a default Gradient Boosting Machine (GBM) for binary classification.
# Load h2o library
suppressPackageStartupMessages(library(h2o))
# Start and connect to a local H2O cluster
h2o.init(nthreads = -1)
# Importing data from local CSV
h_secom <- h2o.importFile(path = "secom.csv", destination_frame = "h_secom")
# Print out column names
colnames(h_secom)
# Look at "Classification"
summary(h_secom$Classification, exact_quantiles=TRUE)
# "Classification" is a column of numerical values
# Convert "Classification" in secom dataset from numerical to categorical value
h_secom$Classification <- as.factor(h_secom$Classification)
# Look at "Classification" again
summary(h_secom$Classification, exact_quantiles=TRUE)
# Define target (y) and features (x)
target <- "Classification"
features <- setdiff(colnames(h_secom), target)
print(features)
# Splitting dataset into training and test
h_split <- h2o.splitFrame(h_secom, ratios = 0.7, seed = 1234)
h_train <- h_split[[1]] # 70%
h_test <- h_split[[2]] # 30%
# Look at the size
dim(h_train)
dim(h_test)
# Check Classification in each dataset
summary(h_train$Classification, exact_quantiles = TRUE)
summary(h_test$Classification, exact_quantiles = TRUE)
# H2O Gradient Boosting Machine with default settings
model <- h2o.gbm(x = features,
y = target,
training_frame = h_train,
seed = 1234)
# Print out model summary
summary(model)
# Check performance on test set
h2o.performance(model, h_test)
# # Making Predictions
# Use the model for predictions
yhat_test <- h2o.predict(model, h_test)
# Show first 10 rows
head(yhat_test, 10)
| use_cases/predictive_maintenance/step_01_basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # + 
# # **Word Count Lab: Building a word count application**
# #### This lab will build on the techniques covered in the Spark tutorial to develop a simple word count application. The volume of unstructured text in existence is growing dramatically, and Spark is an excellent tool for analyzing this type of data. In this lab, we will write code that calculates the most common words in the [Complete Works of William Shakespeare](http://www.gutenberg.org/ebooks/100) retrieved from [Project Gutenberg](http://www.gutenberg.org/wiki/Main_Page). This could also be scaled to find the most common words on the Internet.
# #### ** During this lab we will cover: **
# #### *Part 1:* Creating a base RDD and pair RDDs
# #### *Part 2:* Counting with pair RDDs
# #### *Part 3:* Finding unique words and a mean value
# #### *Part 4:* Apply word count to a file
# #### Note that, for reference, you can look up the details of the relevant methods in [Spark's Python API](https://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD)
# ### ** Part 1: Creating a base RDD and pair RDDs **
# #### In this part of the lab, we will explore creating a base RDD with `parallelize` and using pair RDDs to count words.
# #### ** (1a) Create a base RDD **
# #### We'll start by generating a base RDD by using a Python list and the `sc.parallelize` method. Then we'll print out the type of the base RDD.
wordsList = ['cat', 'elephant', 'rat', 'rat', 'cat']
wordsRDD = sc.parallelize(wordsList, 4)
# Print out the type of wordsRDD
print type(wordsRDD)
# #### ** (1b) Pluralize and test **
# #### Let's use a `map()` transformation to add the letter 's' to each string in the base RDD we just created. We'll define a Python function that returns the word with an 's' at the end of the word. Please replace `<FILL IN>` with your solution. If you have trouble, the next cell has the solution. After you have defined `makePlural` you can run the third cell which contains a test. If you implementation is correct it will print `1 test passed`.
# #### This is the general form that exercises will take, except that no example solution will be provided. Exercises will include an explanation of what is expected, followed by code cells where one cell will have one or more `<FILL IN>` sections. The cell that needs to be modified will have `# TODO: Replace <FILL IN> with appropriate code` on its first line. Once the `<FILL IN>` sections are updated and the code is run, the test cell can then be run to verify the correctness of your solution. The last code cell before the next markdown section will contain the tests.
# +
# TODO: Replace <FILL IN> with appropriate code
def makePlural(word):
"""Adds an 's' to `word`.
Note:
This is a simple function that only adds an 's'. No attempt is made to follow proper
pluralization rules.
Args:
word (str): A string.
Returns:
str: A string with 's' added to it.
"""
return word + 's'
print makePlural('cat')
# +
# One way of completing the function
def makePlural(word):
return word + 's'
print makePlural('cat')
# -
# Load in the testing code and check to see if your answer is correct
# If incorrect it will report back '1 test failed' for each failed test
# Make sure to rerun any cell you change before trying the test again
from test_helper import Test
# TEST Pluralize and test (1b)
Test.assertEquals(makePlural('rat'), 'rats', 'incorrect result: makePlural does not add an s')
# #### ** (1c) Apply `makePlural` to the base RDD **
# #### Now pass each item in the base RDD into a [map()](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.map) transformation that applies the `makePlural()` function to each element. And then call the [collect()](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.collect) action to see the transformed RDD.
# TODO: Replace <FILL IN> with appropriate code
pluralRDD = wordsRDD.map(makePlural)
print pluralRDD.collect()
# TEST Apply makePlural to the base RDD(1c)
Test.assertEquals(pluralRDD.collect(), ['cats', 'elephants', 'rats', 'rats', 'cats'],
'incorrect values for pluralRDD')
# #### ** (1d) Pass a `lambda` function to `map` **
# #### Let's create the same RDD using a `lambda` function.
# TODO: Replace <FILL IN> with appropriate code
pluralLambdaRDD = wordsRDD.map(lambda word: word + 's')
print pluralLambdaRDD.collect()
# TEST Pass a lambda function to map (1d)
Test.assertEquals(pluralLambdaRDD.collect(), ['cats', 'elephants', 'rats', 'rats', 'cats'],
'incorrect values for pluralLambdaRDD (1d)')
# #### ** (1e) Length of each word **
# #### Now use `map()` and a `lambda` function to return the number of characters in each word. We'll `collect` this result directly into a variable.
# TODO: Replace <FILL IN> with appropriate code
pluralLengths = (pluralRDD
.map(lambda word: len(word))
.collect())
print pluralLengths
# TEST Length of each word (1e)
Test.assertEquals(pluralLengths, [4, 9, 4, 4, 4],
'incorrect values for pluralLengths')
# #### ** (1f) Pair RDDs **
# #### The next step in writing our word counting program is to create a new type of RDD, called a pair RDD. A pair RDD is an RDD where each element is a pair tuple `(k, v)` where `k` is the key and `v` is the value. In this example, we will create a pair consisting of `('<word>', 1)` for each word element in the RDD.
# #### We can create the pair RDD using the `map()` transformation with a `lambda()` function to create a new RDD.
# TODO: Replace <FILL IN> with appropriate code
wordPairs = wordsRDD.map(lambda word: (word, 1))
print wordPairs.collect()
# TEST Pair RDDs (1f)
Test.assertEquals(wordPairs.collect(),
[('cat', 1), ('elephant', 1), ('rat', 1), ('rat', 1), ('cat', 1)],
'incorrect value for wordPairs')
# ### ** Part 2: Counting with pair RDDs **
# #### Now, let's count the number of times a particular word appears in the RDD. There are multiple ways to perform the counting, but some are much less efficient than others.
# #### A naive approach would be to `collect()` all of the elements and count them in the driver program. While this approach could work for small datasets, we want an approach that will work for any size dataset including terabyte- or petabyte-sized datasets. In addition, performing all of the work in the driver program is slower than performing it in parallel in the workers. For these reasons, we will use data parallel operations.
# #### ** (2a) `groupByKey()` approach **
# #### An approach you might first consider (we'll see shortly that there are better ways) is based on using the [groupByKey()](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.groupByKey) transformation. As the name implies, the `groupByKey()` transformation groups all the elements of the RDD with the same key into a single list in one of the partitions. There are two problems with using `groupByKey()`:
# + #### The operation requires a lot of data movement to move all the values into the appropriate partitions.
# + #### The lists can be very large. Consider a word count of English Wikipedia: the lists for common words (e.g., the, a, etc.) would be huge and could exhaust the available memory in a worker.
#
# #### Use `groupByKey()` to generate a pair RDD of type `('word', iterator)`.
# TODO: Replace <FILL IN> with appropriate code
# Note that groupByKey requires no parameters
wordsGrouped = wordPairs.groupByKey()
for key, value in wordsGrouped.collect():
print '{0}: {1}'.format(key, list(value))
# TEST groupByKey() approach (2a)
Test.assertEquals(sorted(wordsGrouped.mapValues(lambda x: list(x)).collect()),
[('cat', [1, 1]), ('elephant', [1]), ('rat', [1, 1])],
'incorrect value for wordsGrouped')
# #### ** (2b) Use `groupByKey()` to obtain the counts **
# #### Using the `groupByKey()` transformation creates an RDD containing 3 elements, each of which is a pair of a word and a Python iterator.
# #### Now sum the iterator using a `map()` transformation. The result should be a pair RDD consisting of (word, count) pairs.
# TODO: Replace <FILL IN> with appropriate code
wordCountsGrouped = wordsGrouped.map(lambda (k,v): (k, sum(v)))
print wordCountsGrouped.collect()
# TEST Use groupByKey() to obtain the counts (2b)
Test.assertEquals(sorted(wordCountsGrouped.collect()),
[('cat', 2), ('elephant', 1), ('rat', 2)],
'incorrect value for wordCountsGrouped')
# #### ** (2c) Counting using `reduceByKey` **
# #### A better approach is to start from the pair RDD and then use the [reduceByKey()](http://spark.apache.org/docs/latest/api/python/pyspark.html#pyspark.RDD.reduceByKey) transformation to create a new pair RDD. The `reduceByKey()` transformation gathers together pairs that have the same key and applies the function provided to two values at a time, iteratively reducing all of the values to a single value. `reduceByKey()` operates by applying the function first within each partition on a per-key basis and then across the partitions, allowing it to scale efficiently to large datasets.
# +
# TODO: Replace <FILL IN> with appropriate code
# Note that reduceByKey takes in a function that accepts two values and returns a single value
wordCounts = wordPairs.reduceByKey(lambda a,b: a+b)
print wordCounts.collect()
# -
# TEST Counting using reduceByKey (2c)
Test.assertEquals(sorted(wordCounts.collect()), [('cat', 2), ('elephant', 1), ('rat', 2)],
'incorrect value for wordCounts')
# #### ** (2d) All together **
# #### The expert version of the code performs the `map()` to pair RDD, `reduceByKey()` transformation, and `collect` in one statement.
# TODO: Replace <FILL IN> with appropriate code
wordCountsCollected = (wordsRDD
.map(lambda word: (word, 1))
.reduceByKey(lambda a,b: a+b)
.collect())
print wordCountsCollected
# TEST All together (2d)
Test.assertEquals(sorted(wordCountsCollected), [('cat', 2), ('elephant', 1), ('rat', 2)],
'incorrect value for wordCountsCollected')
# ### ** Part 3: Finding unique words and a mean value **
# #### ** (3a) Unique words **
# #### Calculate the number of unique words in `wordsRDD`. You can use other RDDs that you have already created to make this easier.
# TODO: Replace <FILL IN> with appropriate code
uniqueWords = wordsRDD.map(lambda word: (word, 1)).distinct().count()
print uniqueWords
# TEST Unique words (3a)
Test.assertEquals(uniqueWords, 3, 'incorrect count of uniqueWords')
# #### ** (3b) Mean using `reduce` **
# #### Find the mean number of words per unique word in `wordCounts`.
# #### Use a `reduce()` action to sum the counts in `wordCounts` and then divide by the number of unique words. First `map()` the pair RDD `wordCounts`, which consists of (key, value) pairs, to an RDD of values.
# +
# TODO: Replace <FILL IN> with appropriate code
from operator import add
totalCount = (wordCounts
.map(lambda (a,b): b)
.reduce(add))
average = totalCount / float(wordCounts.distinct().count())
print totalCount
print round(average, 2)
# -
# TEST Mean using reduce (3b)
Test.assertEquals(round(average, 2), 1.67, 'incorrect value of average')
# ### ** Part 4: Apply word count to a file **
# #### In this section we will finish developing our word count application. We'll have to build the `wordCount` function, deal with real world problems like capitalization and punctuation, load in our data source, and compute the word count on the new data.
# #### ** (4a) `wordCount` function **
# #### First, define a function for word counting. You should reuse the techniques that have been covered in earlier parts of this lab. This function should take in an RDD that is a list of words like `wordsRDD` and return a pair RDD that has all of the words and their associated counts.
# TODO: Replace <FILL IN> with appropriate code
def wordCount(wordListRDD):
"""Creates a pair RDD with word counts from an RDD of words.
Args:
wordListRDD (RDD of str): An RDD consisting of words.
Returns:
RDD of (str, int): An RDD consisting of (word, count) tuples.
"""
return (wordListRDD
.map(lambda a : (a,1))
.reduceByKey(lambda a,b: a+b))
print wordCount(wordsRDD).collect()
# TEST wordCount function (4a)
Test.assertEquals(sorted(wordCount(wordsRDD).collect()),
[('cat', 2), ('elephant', 1), ('rat', 2)],
'incorrect definition for wordCount function')
# #### ** (4b) Capitalization and punctuation **
# #### Real world files are more complicated than the data we have been using in this lab. Some of the issues we have to address are:
# + #### Words should be counted independent of their capitialization (e.g., Spark and spark should be counted as the same word).
# + #### All punctuation should be removed.
# + #### Any leading or trailing spaces on a line should be removed.
#
# #### Define the function `removePunctuation` that converts all text to lower case, removes any punctuation, and removes leading and trailing spaces. Use the Python [re](https://docs.python.org/2/library/re.html) module to remove any text that is not a letter, number, or space. Reading `help(re.sub)` might be useful.
# TODO: Replace <FILL IN> with appropriate code
import re
def removePunctuation(text):
"""Removes punctuation, changes to lower case, and strips leading and trailing spaces.
Note:
Only spaces, letters, and numbers should be retained. Other characters should should be
eliminated (e.g. it's becomes its). Leading and trailing spaces should be removed after
punctuation is removed.
Args:
text (str): A string.
Returns:
str: The cleaned up string.
"""
return re.sub("[^a-zA-Z0-9 ]", "", text.strip(" ").lower())
print removePunctuation('Hi, you!')
print removePunctuation(' No under_score!')
# TEST Capitalization and punctuation (4b)
Test.assertEquals(removePunctuation(" The Elephant's 4 cats. "),
'the elephants 4 cats',
'incorrect definition for removePunctuation function')
# #### ** (4c) Load a text file **
# #### For the next part of this lab, we will use the [Complete Works of William Shakespeare](http://www.gutenberg.org/ebooks/100) from [Project Gutenberg](http://www.gutenberg.org/wiki/Main_Page). To convert a text file into an RDD, we use the `SparkContext.textFile()` method. We also apply the recently defined `removePunctuation()` function using a `map()` transformation to strip out the punctuation and change all text to lowercase. Since the file is large we use `take(15)`, so that we only print 15 lines.
# +
# Just run this code
import os.path
baseDir = os.path.join('data')
inputPath = os.path.join('cs100', 'lab1', 'shakespeare.txt')
fileName = os.path.join(baseDir, inputPath)
shakespeareRDD = (sc
.textFile(fileName, 8)
.map(removePunctuation))
print '\n'.join(shakespeareRDD
.zipWithIndex() # to (line, lineNum)
.map(lambda (l, num): '{0}: {1}'.format(num, l)) # to 'lineNum: line'
.take(15))
# -
# #### ** (4d) Words from lines **
# #### Before we can use the `wordcount()` function, we have to address two issues with the format of the RDD:
# + #### The first issue is that that we need to split each line by its spaces.
# + #### The second issue is we need to filter out empty lines.
#
# #### Apply a transformation that will split each element of the RDD by its spaces. For each element of the RDD, you should apply Python's string [split()](https://docs.python.org/2/library/string.html#string.split) function. You might think that a `map()` transformation is the way to do this, but think about what the result of the `split()` function will be.
# TODO: Replace <FILL IN> with appropriate code
shakespeareWordsRDD = shakespeareRDD.flatMap(lambda a: a.split(" "))
shakespeareWordCount = shakespeareWordsRDD.count()
print shakespeareWordsRDD.top(5)
print shakespeareWordCount
# TEST Words from lines (4d)
# This test allows for leading spaces to be removed either before or after
# punctuation is removed.
Test.assertTrue(shakespeareWordCount == 927631 or shakespeareWordCount == 928908,
'incorrect value for shakespeareWordCount')
Test.assertEquals(shakespeareWordsRDD.top(5),
[u'zwaggerd', u'zounds', u'zounds', u'zounds', u'zounds'],
'incorrect value for shakespeareWordsRDD')
# #### ** (4e) Remove empty elements **
# #### The next step is to filter out the empty elements. Remove all entries where the word is `''`.
# TODO: Replace <FILL IN> with appropriate code
shakeWordsRDD = shakespeareWordsRDD.filter(lambda word: len(word) > 0)
shakeWordCount = shakeWordsRDD.count()
print shakeWordCount
# TEST Remove empty elements (4e)
Test.assertEquals(shakeWordCount, 882996, 'incorrect value for shakeWordCount')
# #### ** (4f) Count the words **
# #### We now have an RDD that is only words. Next, let's apply the `wordCount()` function to produce a list of word counts. We can view the top 15 words by using the `takeOrdered()` action; however, since the elements of the RDD are pairs, we need a custom sort function that sorts using the value part of the pair.
# #### You'll notice that many of the words are common English words. These are called stopwords. In a later lab, we will see how to eliminate them from the results.
# #### Use the `wordCount()` function and `takeOrdered()` to obtain the fifteen most common words and their counts.
# TODO: Replace <FILL IN> with appropriate code
top15WordsAndCounts = wordCount(shakeWordsRDD).takeOrdered(15, lambda (a,b): -b)
print '\n'.join(map(lambda (w, c): '{0}: {1}'.format(w, c), top15WordsAndCounts))
# TEST Count the words (4f)
Test.assertEquals(top15WordsAndCounts,
[(u'the', 27361), (u'and', 26028), (u'i', 20681), (u'to', 19150), (u'of', 17463),
(u'a', 14593), (u'you', 13615), (u'my', 12481), (u'in', 10956), (u'that', 10890),
(u'is', 9134), (u'not', 8497), (u'with', 7771), (u'me', 7769), (u'it', 7678)],
'incorrect value for top15WordsAndCounts')
| Week 2 - Introduction to Apache Spark/lab1_word_count_student.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
a = pd.read_csv("normal_texts_final.csv") #reading all data
b = pd.read_csv("final_predated - data.csv")
a
b
c= pd.concat([a,b]) #Combining normal texts and predated texts
c
c =c.sample(frac=1).reset_index(drop=True) #Shuffling Data and Reseting index
c
c.to_csv('Combined_data.csv', index=False)
| data/Data_Source_preprocessing/combine_normal_predated/Combine_All_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # R to python
#
#
# with the rpy2 we can run R code into an txt file or just plain text, then setting into a <i> robjects.r() </i> function
#
#
# to run the following it is necesary to have installed R on your computer https://www.r-project.org/ <br>
# and install rpy2 by <code> pip install rpy2 </code>
# # LINEAR MODEL IN R
# los chunks a continuacion son una implementacion de Rpy2 para correr un <i>lm()</i> en R y desplegar los resultados y plots en python
# +
import pandas as pd
import numpy as np
import datetime
from pathlib import Path
from rpy2 import robjects
#from tadpole_algorithms.models import BenchmarkLastVisit
#from tadpole_algorithms.preprocessing.split import split_test_train_tadpole
#import os
#os.environ['R_HOME'] = "C:/Program Files/R/R-3.6.3"
#import sys
# insert at 1, 0 is the script path (or '' in REPL)
#sys.path.insert(1, '/Users/Eider/Documents/GitHub/tadpole-algorithms')
import tadpole_algorithms
from tadpole_algorithms.models import BenchmarkSVM_R
# -
#instanciate the model to get the functions
model = BenchmarkSVM_R()
#load a csv and do some preprosess so R can handle it
df_diab = pd.read_csv("diabetes.csv")
#call the preprocess function to create a R dataframe with a python dataframe
df_diab_r = model.preprocess_df_R(df_diab)
df_diab_r
#model the data with a linear model and get the results
diab_lm = model.modelfitting_R(model="lm",
formula="y~.",
dataframe=df_diab_r) #can also use a 'dumb' formula and pass a dataframe
#diab_lm
#more detail results
print(diab_lm.names) # view all names
# +
#create a new dataframe
predict_df = pd.DataFrame({'age': np.random.normal(9, 4, 100),
'acidity':np.random.normal(-8,7,100) } )
#parse python df to R df
predict_df_R = model.preprocess_df_R(predict_df)
#predict using our model on new data
predictions = model.predict_R(model=diab_lm,
test_df=predict_df_R)
predictions
# -
# # BWSIMS (UNDER CONSTRUCTION)
# +
bswimstext = ""
with open('./BSWIMS.txt', 'r') as file:
#this file contains the BSWIMS function
bswimstext = file.read()
robjects.r(bswimstext)
#robjects.r(''' BSWiMS.model(formula = paste(theOutcome," ~ 1"),data = theData,NumberofRepeats = bswimsReps) ''')
# -
# # SVM IN R(UNDER CONSTRUCTION)
diab_svm= model.SVM_fitting_R(formula="y~.",dataframe=df_diab_r)
#predict using our model on new data
predictions = model.predict_R(model=diab_svm,
test_df=predict_df_R)
predictions
FRESA = importr('FRESA.CAD')
FRESA
df = model.tadpole_tidyng()
df
#predictions using svm over tadpole tidy dataframe
tadpole_svm= model.SVM_fitting_R(formula="status~.",dataframe=df)
#predict using our model on the same data
predictions = model.predict_R(model=tadpole_svm,
test_df=df)
predictions
crear funcion de xgb en r y hacerla funcionar en el pipeline
gmb_r= model.caret_gmb_modelfitting_R()
gmb_r.names
predictions_gmb = model.predict_R(model=gmb_r,
test_df=df)
predictions_gmb
| R to python.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#
# Note, this is copied from https://gist.github.com/bbengfort/044682e76def583a12e6c09209c664a1
# I'm trying to learn from it and apply the same patterns to the movie classification project
#
from nltk.corpus import wordnet
from sklearn.feature_extraction.text import CountVectorizer,TfidfTransformer
from sklearn.metrics import f1_score
from sklearn.model_selection import cross_val_score
from sklearn.svm import LinearSVC
from sklearn.pipeline import Pipeline
from sklearn.base import BaseEstimator, TransformerMixin
import pandas as pd
import numpy as np
import string
from nltk.corpus import stopwords as sw
from nltk.corpus import wordnet as wn
from nltk import wordpunct_tokenize
from nltk import WordNetLemmatizer
from nltk import sent_tokenize
from nltk import pos_tag
# +
# Load training set (train.tsv - tab separated values)
# cols => PhraseId, SentenceId, Phrase, Sentiment
train_df = pd.read_csv('data/train.csv', sep='|')
train_df.head()
# +
import os
import time
import string
import pickle
from operator import itemgetter
from nltk.corpus import stopwords as sw
from nltk.corpus import wordnet as wn
from nltk import wordpunct_tokenize
from nltk import WordNetLemmatizer
from nltk import sent_tokenize
from nltk import pos_tag
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import LabelEncoder
from sklearn.linear_model import SGDClassifier
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.metrics import classification_report as clsr
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.cross_validation import train_test_split as tts
def timeit(func):
"""
Simple timing decorator
"""
def wrapper(*args, **kwargs):
start = time.time()
result = func(*args, **kwargs)
delta = time.time() - start
return result, delta
return wrapper
def identity(arg):
"""
Simple identity function works as a passthrough.
"""
return arg
class NLTKPreprocessor(BaseEstimator, TransformerMixin):
"""
Transforms input data by using NLTK tokenization, lemmatization, and
other normalization and filtering techniques.
"""
def __init__(self, stopwords=None, punct=None, lower=True, strip=True):
"""
Instantiates the preprocessor, which make load corpora, models, or do
other time-intenstive NLTK data loading.
"""
self.lower = lower
self.strip = strip
self.stopwords = set(stopwords) if stopwords else set(sw.words('english'))
self.punct = set(punct) if punct else set(string.punctuation)
self.lemmatizer = WordNetLemmatizer()
def fit(self, X, y=None):
"""
Fit simply returns self, no other information is needed.
"""
return self
def inverse_transform(self, X):
"""
No inverse transformation
"""
return X
def transform(self, X):
"""
Actually runs the preprocessing on each document.
"""
return [
list(self.tokenize(doc)) for doc in X
]
def tokenize(self, document):
"""
Returns a normalized, lemmatized list of tokens from a document by
applying segmentation (breaking into sentences), then word/punctuation
tokenization, and finally part of speech tagging. It uses the part of
speech tags to look up the lemma in WordNet, and returns the lowercase
version of all the words, removing stopwords and punctuation.
"""
# Break the document into sentences
for sent in sent_tokenize(document):
# Break the sentence into part of speech tagged tokens
for token, tag in pos_tag(wordpunct_tokenize(sent)):
# Apply preprocessing to the token
token = token.lower() if self.lower else token
token = token.strip() if self.strip else token
token = token.strip('_') if self.strip else token
token = token.strip('*') if self.strip else token
# If punctuation or stopword, ignore token and continue
if token in self.stopwords or all(char in self.punct for char in token):
continue
# Lemmatize the token and yield
lemma = self.lemmatize(token, tag)
yield lemma
def lemmatize(self, token, tag):
"""
Converts the Penn Treebank tag to a WordNet POS tag, then uses that
tag to perform much more accurate WordNet lemmatization.
"""
tag = {
'N': wn.NOUN,
'V': wn.VERB,
'R': wn.ADV,
'J': wn.ADJ
}.get(tag[0], wn.NOUN)
return self.lemmatizer.lemmatize(token, tag)
@timeit
def build_and_evaluate(X, y, classifier=SGDClassifier, outpath=None, verbose=True):
"""
Builds a classifer for the given list of documents and targets in two
stages: the first does a train/test split and prints a classifier report,
the second rebuilds the model on the entire corpus and returns it for
operationalization.
X: a list or iterable of raw strings, each representing a document.
y: a list or iterable of labels, which will be label encoded.
Can specify the classifier to build with: if a class is specified then
this will build the model with the Scikit-Learn defaults, if an instance
is given, then it will be used directly in the build pipeline.
If outpath is given, this function will write the model as a pickle.
If verbose, this function will print out information to the command line.
"""
@timeit
def build(classifier, X, y=None):
"""
Inner build function that builds a single model.
"""
if isinstance(classifier, type):
classifier = classifier()
model = Pipeline([
('preprocessor', NLTKPreprocessor()),
('vectorizer', TfidfVectorizer(tokenizer=identity, preprocessor=None, lowercase=False)),
('classifier', classifier),
])
model.fit(X, y)
return model
# Label encode the targets
labels = LabelEncoder()
y = labels.fit_transform(y)
# Begin evaluation
if verbose: print("Building for evaluation")
X_train, X_test, y_train, y_test = tts(X, y, test_size=0.2)
model, secs = build(classifier, X_train, y_train)
if verbose: print("Evaluation model fit in {:0.3f} seconds".format(secs))
if verbose: print("Classification Report:\n")
y_pred = model.predict(X_test)
print(clsr(y_test, y_pred, target_names=labels.classes_))
if verbose: print("Building complete model and saving ...")
model, secs = build(classifier, X, y)
model.labels_ = labels
if verbose: print("Complete model fit in {:0.3f} seconds".format(secs))
if outpath:
with open(outpath, 'wb') as f:
pickle.dump(model, f)
print("Model written out to {}".format(outpath))
return model
def show_most_informative_features(model, text=None, n=20):
"""
Accepts a Pipeline with a classifer and a TfidfVectorizer and computes
the n most informative features of the model. If text is given, then will
compute the most informative features for classifying that text.
Note that this function will only work on linear models with coefs_
"""
# Extract the vectorizer and the classifier from the pipeline
vectorizer = model.named_steps['vectorizer']
classifier = model.named_steps['classifier']
# Check to make sure that we can perform this computation
if not hasattr(classifier, 'coef_'):
raise TypeError(
"Cannot compute most informative features on {} model.".format(
classifier.__class__.__name__
)
)
if text is not None:
# Compute the coefficients for the text
tvec = model.transform([text]).toarray()
else:
# Otherwise simply use the coefficients
tvec = classifier.coef_
# Zip the feature names with the coefs and sort
coefs = sorted(
zip(tvec[0], vectorizer.get_feature_names()),
key=itemgetter(0), reverse=True
)
topn = zip(coefs[:n], coefs[:-(n+1):-1])
# Create the output string to return
output = []
# If text, add the predicted value to the output.
if text is not None:
output.append("\"{}\"".format(text))
output.append("Classified as: {}".format(model.predict([text])))
output.append("")
# Create two columns with most negative and most positive features.
for (cp, fnp), (cn, fnn) in topn:
output.append(
"{:0.4f}{: >15} {:0.4f}{: >15}".format(cp, fnp, cn, fnn)
)
return "\n".join(output)
if __name__ == "__main__":
PATH = "model.pickle"
if not os.path.exists(PATH):
# Time to build the model
from nltk.corpus import movie_reviews as reviews
X = [reviews.raw(fileid) for fileid in reviews.fileids()]
y = [reviews.categories(fileid)[0] for fileid in reviews.fileids()]
model = build_and_evaluate(X,y, outpath=PATH)
else:
with open(PATH, 'rb') as f:
model = pickle.load(f)
print(show_most_informative_features(model))
# -
# +
from nltk.corpus import movie_reviews as reviews
# X = [reviews.raw(fileid) for fileid in reviews.fileids()]
# y = [reviews.categories(fileid)[0] for fileid in reviews.fileids()]
X = [train_df.raw(SentenceId) for SentenceId in train_df.SentenceId()]
y = [train_df.categories(SentenceId)[0] for SentenceId in train_df.SentenceId()]
model = build_and_evaluate(X,y, outpath='results')
# +
with open(PATH, 'rb') as f:
model = pickle.load(f)
print(show_most_informative_features(model))
# -
| MSRA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import time
k=0
e=0
i=10
print('Введи своё имя')
klikuha = input()
print('Введи свою фамилию')
familiya = input()
print('Введи свой возраст')
age = int(input())
print('Введи свой пол: жен или муж')
pol=input()
if pol!=str('жен') and pol!=str('муж'):
print('Соблюдай условия')
pol=input()
print('Введи свой вес')
ves=int(input())
print('Введи свой рост')
H=int(input())
print('Добро пожаловать в систему, ' + klikuha + ' ' + familiya)
print('Твои показания на данный момент:')
c = ves/((H/100)*(H/100))
if c<16:
print('У тебя острый дефицит массы тела')
if c>=16 and c<18.5:
print('У тебя дефицит массы тела')
if c>=18.5 and c<25:
print('Нормальный вес')
k=k+1
if c>=25 and c<30:
print('Предожирение')
if c>=30 and c<35:
print('Ожирение 1 степени')
if c>=35 and c<40:
print('Ожирение 2 степени')
if c>=40:
print('Ожирение 3 степени')
print('Давай проверим твои умственные способности. Реши пример: 44*444444')
R=int(input())
if R==19555536:
print('Умственные способности в норме')
e=e+1
else:
print('У нас для тебя плохие новости...')
print('Идет дополнительный анализ...')
for i in range(10):
time.sleep(1)
print('...')
print('Делаем выводы...')
for i in range(10):
time.sleep(1)
print('...')
if k>0:
print('Твое физическое здоровье в норме')
else:
print('Ты в плохой форме, чувак')
if e>0:
print('Твои умственные способности в норме')
else:
print('Ты отстаешь в развитии')
print('Выход из аккаунта: ' + klikuha + ' ' + familiya)
while i>0:
time.sleep(1)
print(i)
i=i-1
print('Выход завершен')
for i in range(10):
time.sleep(1)
print('...')
# ##
| BMI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="evSOEz99Mvop"
# ## Exploring Text Data
#
# Working with text is generally more challenging than working with numerical data. Hence, any kind of technique that helps in generating an intuition of the existing dataset is welcome. One of the simplest approach to understand any text document or to compare multiple documents can be to compute a frequency table of individual words present in the document/documents and use it to conduct further experiements like: finding top words per document, finding top common words among documents etc.
#
# In this notebook, I have taken the challenge of Analyzing Sentiments from Twitter data, so I'll focus on how to generate word frequencies and use it to create **Word Clouds** to help me get a better overall understanding of the dataset.
#
# **Note:-** We are using the same dataset from `tweets.csv` that we used earlier. You can get it [here.](https://studio.trainings.analyticsvidhya.com/assets/courseware/v1/aa0ae6514e0be95f11be85b84d4fd6d2/asset-v1:AnalyticsVidhya+NLP101+2018_T1+type@asset+block/tweets.csv)
#
# ### Table of Contents
# 1. About the Dataset
# 2. Generating Word Frequency
# 3. EDA using Word Clouds
# 4. Why to Preprocess text data?
# 5. Challenge
# + [markdown] id="_Yx4W6n8MvpA"
# ### 1. About the Dataset
#
# The dataset that we are going to use is the same dataset of tweets from Twitter. You can download it from [here.](https://studio.trainings.analyticsvidhya.com/assets/courseware/v1/aa0ae6514e0be95f11be85b84d4fd6d2/asset-v1:AnalyticsVidhya+NLP101+2018_T1+type@asset+block/tweets.csv)
# Let's load the dataset using pandas and have a quick look at some sample tweets.
# + id="u85tEt4fMvpC" colab={"base_uri": "https://localhost:8080/", "height": 538} outputId="a9c4f142-65c6-46f7-a420-cb807aa7e44d"
#Load the dataset
import pandas as pd
dataset = pd.read_csv('tweets.csv', encoding = 'ISO-8859-1')
dataset.head()
# + [markdown] id="xBQo5GPIMvpF"
# As can be seen above, **text** column is of interest to us as it contains the tweet. At this point, you don't have to worry about other columns as that will be handled in future modules. Let's go ahead and inspect some of the tweets.
#
# ### 2. Generating Word Frequency
#
# Let's first generate a frequency table of all the words present in all the tweets combined.
# + id="FcKrM5JuMvpG" outputId="91c22f08-4f15-48bb-af4c-6324c5b571a8"
def gen_freq(text):
#Will store the list of words
word_list = []
#Loop over all the tweets and extract words into word_list
for tw_words in text.split():
word_list.extend(tw_words)
#Create word frequencies using word_list
word_freq = pd.Series(word_list).value_counts()
#Print top 20 words
word_freq[:20]
return word_freq
gen_freq(dataset.text.str)
# + [markdown] id="MuO-mmRmMvpI"
# ### 3. EDA using Word Clouds
#
# Now that you have succesfully created a frequency table, you can use that to create multiple **visualizations** in the form of word clouds. Sometimes, the quickest way to understand the context of the text data is using a word cloud of top 100-200 words. Let's see how to create that in Python.
#
# **Note:-** You'll use the `WordCloud` library of Python. You can install it by -
#
# `pip install wordcloud`
# + id="G1_0GDUgMvpJ" outputId="c0a06c1d-0236-4c3f-bd12-3e395925649f"
#Import libraries
import matplotlib.pyplot as plt
from wordcloud import WordCloud
#Generate word cloud
wc = WordCloud(width=400, height=330, max_words=100, background_color='white').generate_from_frequencies(word_freq)
plt.figure(figsize=(12, 8))
plt.imshow(wc, interpolation='bilinear')
plt.axis('off')
plt.show()
# + [markdown] id="IpeJ9xWXMvpL"
# **Few things to Note:-**
#
# 1. There is noise in the form of "RT" and "&" which can be removed from the word frequency.
# 2. Stop words like "the", "in", "to", "of" etc. are obviously ranking among the top frequency words but these are just constructs of the English language and are not specific to the people's tweets.
# 3. Words like "demonetization" have occured multiple times. The reason for this is that the current text is not **Normalized** so words like "demonetization", "Demonetization" etc. are all considered as different words.
#
# The above are some of the problems that we need to address in order to make better visualization. Let's solve some of the problems!
#
# #### Text Cleaning
#
# I utilize Regex to do text cleaning.
# + id="VgOmoqlGMvpM"
import re
def clean_text(text):
#Remove RT
text = re.sub(r'RT', '', text)
#Fix &
text = re.sub(r'&', '&', text)
#Remove punctuations
text = re.sub(r'[?!.;:,#@-]', '', text)
#Convert to lowercase to maintain consistency
text = text.lower()
return text
# + [markdown] id="FECQ1Gk5MvpO"
# The above will solve problems related to RT, & and also the problem of counting same word twice due to case difference. Yet we can do better, let's remove the common stop words.
#
# #### Stop words Removal
# WordCloud provides its own stopwords list. They are shown as:
#
# + id="XJAgH09iMvpP" outputId="a99469e4-12cc-40e0-b2ba-77b3bfc2b679"
#Import list of stopwards
from wordcloud import STOPWORDS
print(STOPWORDS)
# + [markdown] id="ThSFdqTYMvpQ"
# Now that you know what all has to be changed to improve our word cloud, let's make some wordclouds. We'll call the previous functions of `clean_text()` and `gen_freq()` to perform cleaning and frequency computation operation respectively and drop the words present in `STOPWORDS` from the `word_freq` dictionary.
# + id="PopEtCUuMvpR" outputId="c7b584e2-290e-4929-8693-57573be4bf67"
text = dataset.text.apply(lambda x: clean_text(x))
word_freq = gen_freq(text.str)*100
word_freq = word_freq.drop(labels=STOPWORDS, errors='ignore')
#Generate word cloud
wc = WordCloud(width=450, height=330, max_words=200, background_color='white').generate_from_frequencies(word_freq)
plt.figure(figsize=(12, 14))
plt.imshow(wc, interpolation='bilinear')
plt.axis('off')
plt.show()
# + [markdown] id="0hYGssQ-MvpS"
# Now that you have succesfully created a wordcloud, you can get some insight into the areas of interest of the general twitter users:
#
# - It is evident that people are talking about govt. policies like **demonetization**, **J&K**.
# - There are some personalitites that are mentioned numerous times like **evanspiegel**, **PM <NAME>**, **Dr <NAME>** etc.
# - There are also talks about **oscars**, **youtube** and **terrorists**
# - There are many sub-topics that revolve around demonetization like **atms**, **bank**, **cash**, **paytm** etc. Which tells that many people are concerned about it.
#
# ### 4. Why to Preprocess text data?
#
# We've already seen that without performing preprocessing operations like cleaning, removing stopwords and changing case in the dataset the representation always comes out wrong. In this case, it was that the wordcloud was full of noise but in other cases it might be your Machine Learning model that is going to suffer.
#
# Also something to note is even now some words are misreperesented for example: **modi**, **narendra** and **narendramodi** all refer to the same person. This can eaisly be solved by **Normalizing** our text.
| Exploring_Text_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Implementando la regresión lineal
# +
import torch
import numpy as np
import pandas as pd
import torch.nn as nn
import torch.optim as optim
import matplotlib.pyplot as plt
# +
car_prices = [5,6,7,8,9,10]
units_sold = [8.5, 8, 7.5, 7, 6.5, 6]
plt.scatter(car_prices, units_sold)
# -
prices_array = np.array(car_prices).reshape(-1,1)
units_array = np.array(units_sold).reshape(-1,1)
prices_array, units_array
# +
prices = torch.from_numpy(prices_array).float().requires_grad_(True)
units = torch.from_numpy(units_array).float()
prices, prices.shape
# +
model = nn.Linear(1,1)
loss_function = nn.MSELoss()
optimizer = optim.SGD(model.parameters(), lr=0.015)
losses = []
iterations = 2000
for i in range(iterations):
pred = model(prices)
loss = loss_function(pred, units)
losses.append(loss.data)
optimizer.zero_grad()
loss.backward()
optimizer.step()
print(loss)
plt.plot(range(iterations), losses)
# -
x = torch.Tensor([[14.0]])
p = model(x)
p
| files/pytorch/algoritmos-ml/regresion-lineal/regresion-lineal.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 64-bit (''test'': conda)'
# name: python3710jvsc74a57bd04374d16277cd59720eda5e9a892d33ee7e53ac8b7c0031fbe42f60839aa8916a
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# ## Hysteresis and log-log calibration example
#
# Given a calibration of an instrument for an increasing and decreasing input $x$ [mV] and output of the instrument $y$ [mV]
# + slideshow={"slide_type": "skip"}
import numpy as np
import matplotlib.pyplot as pl
# %matplotlib inline
import matplotlib as mpl
mpl.rcParams['figure.figsize'] = 8, 6
mpl.rcParams['font.size'] = 18
# + slideshow={"slide_type": "slide"}
from IPython.display import Image
Image(filename='../../img/hysteresis_example.png',width=400)
# + slideshow={"slide_type": "skip"}
x = np.array([0.0, 1.0, 2.0, 3.0, 4.0, 5.0, 5.0, 4.0, 3.0, 2.0, 1.0, 0.0])
y = np.array([0.1, 1.1, 2.1, 3.0, 4.1, 5.0, 5.0, 4.2, 3.2, 2.2, 1.2, 0.2])
# + slideshow={"slide_type": "slide"}
pl.plot(x,y,'o')
pl.xlabel('$x$ [mV]')
pl.ylabel('$y$ [mV]')
# -
# 1. We see the error, but we do not know if it is a random or not
# 2. In order to see the hysteresis, we have to set the plot with the lines connecting points:
pl.plot(x,y,'--o')
pl.xlabel('$x$ [mV]')
pl.ylabel('$y$ [mV]')
# ### Estimate the hysteresis error:
#
# $e_h = y_{up} - y_{down}$
#
# $e_{h_{max}} = max(|e_h|)$
#
# $e_{h_{max}}\% = 100\% \cdot \frac{e_{h_{max}}}{y_{max}-y_{min}} $
e_h = y[:6]-np.flipud(y[6:])
print ("e_h =", e_h,"[mV]")
e_hmax = np.max(np.abs(e_h))
print ("e_hmax= %3.2f %s" % (e_hmax,"[mV]"))
e_hmax_p = 100*e_hmax/(np.max(y) - np.min(y))
print ("Relative error = %3.2f%s FSO" % (e_hmax_p,"%"))
# # Sensitivity error example
from IPython.core.display import Image
Image(filename='../../img/sensitivity_error_example.png',width=400)
x = np.array([0.5, 1.0, 2.0, 5.0, 10.0, 20.0, 50.0, 100.0])
y = np.array([0.4, 1.0, 2.3, 6.9, 15.8, 36.4, 110.1, 253.2])
pl.plot(x,y,'--o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$y$ [V]')
pl.title('Calibration curve')
# Sensitivity, $K$ is:
#
# $ K_i = \left( \frac{\partial y}{\partial x} \right)_{x_i} $
K = np.diff(y)/np.diff(x)
print K
pl.plot(x[1:],K,'--o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$K$ [V/cm]')
pl.title('Sensitivity')
# Instead of working with non-linear curve of sensitivity we can use the usual trick: the logarithmic scale
pl.loglog(x,y,'--o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$y$ [V]')
pl.title('Logarithmic scale')
logK = np.diff(np.log(y))/np.diff(np.log(x))
print( logK)
pl.plot(x[1:],logK,'--o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$K$ [V/cm]')
pl.title('Logarithmic sensitivity')
pl.plot([x[1],x[-1]],[1.2,1.2],'r--')
pl.loglog(x,y,'o',x,x**(1.2))
pl.xlabel('$x$ [cm]')
pl.ylabel('$y$ [V]')
pl.title('Logarithmic scale')
pl.legend(('$y$','$x^{1.2}$'),loc='best')
pl.plot(x,y-x**(1.2),'o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$y - y_c$ [V]')
pl.title('Deviation plot')
# pl.legend(('$y$','$x^{1.2}$'),loc='best')
# ## Regression analysis
# Following the recipe of http://www.answermysearches.com/how-to-do-a-simple-linear-regression-in-python/124/
# +
from scipy.stats import t
def linreg(X, Y):
"""
Summary
Linear regression of y = ax + b
Usage
real, real, real = linreg(list, list)
Returns coefficients to the regression line "y=ax+b" from x[] and y[], and R^2 Value
"""
N = len(X)
if N != len(Y): raise(ValueError, 'unequal length')
Sx = Sy = Sxx = Syy = Sxy = 0.0
for x, y in zip(X, Y):
Sx = Sx + x
Sy = Sy + y
Sxx = Sxx + x*x
Syy = Syy + y*y
Sxy = Sxy + x*y
det = Sx * Sx - Sxx * N # see the lecture
a,b = (Sy * Sx - Sxy * N)/det, (Sx * Sxy - Sxx * Sy)/det
meanerror = residual = residualx = 0.0
for x, y in zip(X, Y):
meanerror = meanerror + (y - Sy/N)**2
residual = residual + (y - a * x - b)**2
residualx = residualx + (x - Sx/N)**2
RR = 1 - residual/meanerror
# linear regression, a_0, a_1 => m = 1
m = 1
nu = N - (m+1)
sxy = np.sqrt(residual / nu)
# Var_a, Var_b = ss * N / det, ss * Sxx / det
Sa = sxy * np.sqrt(1/residualx)
Sb = sxy * np.sqrt(Sxx/(N*residualx))
# We work with t-distribution, ()
# t_{nu;\alpha/2} = t_{3,95} = 3.18
tvalue = t.ppf(1-(1-0.95)/2, nu)
print("Estimate: y = ax + b")
print("N = %d" % N)
print("Degrees of freedom $\\nu$ = %d " % nu)
print("a = %.2f $\\pm$ %.3f" % (a, tvalue*Sa/np.sqrt(N)))
print("b = %.2f $\\pm$ %.3f" % (b, tvalue*Sb/np.sqrt(N)))
print("R^2 = %.3f" % RR)
print("Syx = %.3f" % sxy)
print("y = %.2f x + %.2f $\\pm$ %.3f V" % (a, b, tvalue*sxy/np.sqrt(N)))
return a, b, RR, sxy
# -
print (linreg(np.log(x),np.log(y)))
pl.loglog(x,y,'o',x,x**(1.21)-0.0288)
pl.xlabel('$x$ [cm]')
pl.ylabel('$y$ [V]')
pl.title('Logarithmic scale')
pl.legend(('$y$','$x^{1.2}$'),loc='best')
pl.plot(x,y-(x**(1.21)-0.0288),'o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$y - y_c$ [V]')
pl.title('Deviation plot')
# pl.legend(('$y$','$x^{1.2}$'),loc='best')
| notebooks/calibration/hysteresis_error_analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/masvgp/math_3280/blob/main/CS246_Colab_6.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="kPt5q27L5557"
# # CS246 - Colab 6
# ## node2vec
# + [markdown] id="p0-YhEpP_Ds-"
# ### Setup
# + [markdown] id="82kmj9fIVlq0"
# First of all, we install the [graph2vec library](https://github.com/VHRanger/graph2vec) which offers a fast implementation of the node2vec method.
#
# If you are curious to learn how to implement fast random walks on graphs, I recommend you to [read the blog post](https://www.singlelunch.com/2019/08/01/700x-faster-node2vec-models-fastest-random-walks-on-a-graph/) which explains some of the design choices behind this library.
# + id="Ld017NzXCo-g"
# !pip install nodevectors
# + [markdown] id="pUCNE4-5Wc8b"
# We now import the library, and create a small wrapper class which will expose only the few hyperparameters we will need to tune in this Colab
# + id="QiIPtudrGlTf"
from nodevectors import Node2Vec
import networkx as nx
class Node2VecNew(Node2Vec):
"""
Parameters
----------
p : float
p parameter of node2vec
q : float
q parameter of node2vec
d : int
dimensionality of the embedding vectors
"""
def __init__(self, p=1, q=1, d=32):
super().__init__(
n_components=d,
walklen=10,
epochs=50,
return_weight=1.0/p,
neighbor_weight=1.0/q,
threads=0,
w2vparams={'window': 4,
'negative': 5,
'iter': 10,
'ns_exponent': 0.5,
'batch_words': 128})
# + [markdown] id="qwtlO4_m_LbQ"
# Lastly, let's import some of the common libraries needed for our task.
# + id="twk-K-jilWK7"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# + [markdown] id="kAYRX2PMm0L6"
# ### Example
#
# In the example below, we will try to reproduce the plot in slide 8 of [CS246 - Lecture 12](http://web.stanford.edu/class/cs246/slides/12-graphs2.pdf).
# + id="2PsVt5f2CsmP" colab={"base_uri": "https://localhost:8080/"} outputId="39b658db-b266-4aca-9017-dba506207acf"
# Load the Zachary's Karate Club as a NetworkX Graph object
KCG = nx.karate_club_graph()
# Fit embedding model to the Karate Club graph
n2v = Node2VecNew(1, 1, 2)
n2v.fit(KCG)
# + id="ZEjzhwEiC1HD"
embeddings = []
for node in KCG.nodes:
embedding = list(n2v.predict(node))
club = KCG.nodes[node]['club']
embeddings.append(embedding + [club])
# Construct a pandas dataframe with the 2D embeddings from node2vec,
# plus the club name that each node belongs to after the split
df = pd.DataFrame(embeddings, columns=['x', 'y', 'club'])
# + id="fJBTilMGLInb"
# Nodes who stayed with the Mr. Hi will be plotted in red, while nodes
# who moved with the Officer will be plotted in blue
colors = ['red' if x == 'Mr. Hi' else 'blue' for x in df.club]
df.plot.scatter(x='x', y='y', s=50, c=colors)
# + [markdown] id="e6vmpmj6Ylph"
# If our example trained correctly, you should notice a clear separation between the blue and red nodes. Solely from the graph structure, node2vec could predict how the Zachary's Karate Club split!
#
# Tune the hyperparameters ```p``` and ```q```, and notice how they affect the resulting embeddings.
# + [markdown] id="vbmr23B2rJKR"
# ### Your Task
# + [markdown] id="x15OQeyys1xd"
# Now we will study the behavior of node2vec on [barbell graphs](https://en.wikipedia.org/wiki/Barbell_graph).
#
# Below you can see a toy example of a barbell graph generated with NetworkX.
# + id="xUYxs4E8aHgG"
toy_barbell = nx.barbell_graph(7, 0)
nx.draw_kamada_kawai(toy_barbell)
# + [markdown] id="G7K3-57FyD8D"
#
#
#
# Generate a larger barbell graph, where each complete graph has exactly 1000 nodes, and the path length between the complete graphs is equal to 1 (i.e., all the nodes in the barbell graph belong to either one of the two complete graphs, and the connecting path does not have any internal node).
#
# Then, learn node2vec embeddings on this graph, setting ```p = 1, q = 1``` and ```d = 10```.
# + id="k_rmp-SvaIOt"
# YOUR CODE HERE
# + [markdown] id="fyxZBVwhzLQe"
# Write a function that takes as input a node id ```n``` in the graph (e.g., ```5```) and returns a list containing the cosine similarity between the node2vec vector of the input node ```n``` and all the nodes in the given barbell graph (including the similarity with ```n``` itself).
# + id="aq7advtkjXSp"
# YOUR CODE HERE
# + [markdown] id="zw1oNDDD2dZO"
# Generate another barbell graph, this time adding a path of length 51 between the two complete graphs. To find out how, refer to the NetworkX documentation: [https://networkx.github.io/documentation/stable/reference/generated/networkx.generators.classic.barbell_graph.html#networkx.generators.classic.barbell_graph](https://networkx.github.io/documentation/stable/reference/generated/networkx.generators.classic.barbell_graph.html#networkx.generators.classic.barbell_graph)
#
# Learn the node2vec embeddings for the nodes of this new graph, using the same hyperparameters as before.
# + id="OlPg6k6z65Gk"
# YOUR CODE HERE
# + [markdown] id="SIrXJyVNP2AI"
# Once you have working code for each cell above, **head over to Gradescope, read carefully the questions, and submit your solution for this Colab**!
#
| CS246_Colab_6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Date and Time data Types and Tools
#
# The Python standard library includes data types for date and time data, as well as calendar-related functionality.
#
# The `datetime.time`, and `calendar` modules are the main places to start.
from datetime import datetime
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
now = datetime.now()
now
print(now.year, now.month, now.day)
# `datetime` store both the date and time down to the microsecond.
#
# `datetime.timedelta` represents the temproal different two datetime objects.
delta = datetime(2011, 1, 7) - datetime(2008, 6, 24, 8, 15)
delta
print("different day: %s, different second: %s" % (delta.days, delta.seconds))
# ### `timedelta`
#
# You can add(or subtract) a `timedelta` or multiple thereof to a `datetime` object to yield a new shifted object.
#
# ### Types in datetime module
#
# Types | Description |
# --------------|:------------------------------------------------------------------------------------------:|
# **`date`** | Store calendar date(year, month, day) using the Gregorian calendar |
# **`time`** | Store time of days as hours, minutesm and microseconds |
# **`datetime`** | Stroes both date and time |
# **`timedelta`** | Represents the difference between two datetime values(as days, secondsm and microsenconds) |
from datetime import timedelta
start = datetime(2011, 1, 7)
start + timedelta(days = 3, hours = 3, minutes = 3, seconds = 12)
start - 2 * timedelta(days = 12)
# ### Convert between string and datetime
#
# `datetime` objects and pandas `Timestamp` objects, which I'll introduce later, can be ***formatted as string using `str` or the `strftime` method***, passing a format specification.
#
# `strftime`(string format output time)
#
# ***These same format codes can be used to convert strings to dates using `datetime.strptime`(new datetime parsed from a string).***
stamp = datetime(2013, 1, 3)
stamp
stamp.strftime("%Y-%m-%d")
value = '2011-01-03'
datetime.strptime(value, "%Y-%m-%d")
datestr = ["7/6/2011", '8/6/2011']
[datetime.strptime(x, "%m/%d/%Y") for x in datestr]
# ***`datetime.strptime` is the best way to parse a date with a known format.***
#
# However, it can be a bit annoying to have to write a format spec each time, especially date formats.
#
# ***You can use the `parser.parse` method in the third party `dateutil` package.***
#
# ***`dateutil` is capable of parsing almost an human-intelligible date represention.***
from dateutil.parser import parse
parse("2011-01-03")
parse('Jan 31, 1997 10:45 PM')
# In international locales, day appearing before month is very common, so you can pass `dayfirst = True` to indicate this.
parse("6/12/2011", dayfirst = True)
# pandas is generally oriented toward working with arrays of dates, whether used as an axis index or a column in a DataFrame.
#
# ***The `to_datetime` method parses many different kinds of date representayions.***
import pandas as pd
datestr = ["7/6/2011", '8/6/2011']
pd.to_datetime(datestr)
# ***`dateutil.parser.parse` method get `datetime` object, then use `strftime` method output specified format.****
[parse(date).strftime("%Y-%m-%d") for date in datestr]
# ***Using `parse` method can parse string of the diffent datetime format.***
temp = ["12/6/2016", "2016-08-06"]
[parse(date).strftime("%Y-%m-%d") for date in temp]
# ***`NaT`*** (Not a Time) is pandas's NA value for timestamp data.
idx = pd.to_datetime(datestr + [None])
idx
print(idx[2])
print(pd.isnull(idx))
# ### Time Series Basics
#
# The most basic kind of time series object in pandas is a Series indexed by timestamps, which is often represented external to pandas as Python strings or `datetime` object.
dates = [datetime(2011, 1, 2), datetime(2011, 1, 5), datetime(2011, 1, 7),
datetime(2011, 1, 8), datetime(2011, 1, 10), datetime(2011, 1, 12)]
ts = pd.Series(data = np.random.randn(6), index = dates)
ts[-3:]
ts.index
# ***Like other Series, arithmetic operations between differently-indexed time series automatically align on the dates.***
#
# A timestamp can be substituted anywhere you would use a `datetime` object.
#
# ***Additionally, it can store frequency information(if any) and understands how time zone conversions and other kinds of maniplations.***
print(ts)
ts + ts[::2]
stamp = ts.index[0]
stamp
# ### Indexing, Selection, Subseting
#
# ***As a convenience, you can also pass a string that is interpretable as a date.***
#
# For longer time series, a year or only a year and month can be passed to easily select of data.
stamp = ts.index[2]
ts[stamp]
print(ts['1/10/2011'])
print(ts['20110110'])
longer_ts = pd.Series(data = np.random.randn(1000),
index = pd.date_range(start = '1/1/2000', periods = 1000))
print(len(longer_ts))
longer_ts[-5:]
# You can just pass year or month to slice with data.
print('Select 2001 :', longer_ts['2001'][-3:])
print("-"*50)
print('Select 2001-5:', longer_ts['2001-05'][-3:])
# ***Slice with dataset works just like a regular Series.***
longer_ts[(longer_ts.index >= datetime(2000, 2, 1)) & (longer_ts.index <= datetime(2000, 2, 27))][-5:]
longer_ts[longer_ts.index < '2000-01-15'][-3:]
ts['1/6/2011':'1/11/2011']
# As before you can pass either a string data, datetime, ot Timestamp.
#
# Remember that slicing this manner prodeces views on the source time series just like slicing NumPy array.
#
# There is an equivalient instance method `truncate` which slices a TimeSeries between two dates.
ts.truncate(after = '2011-01-09')
dates = pd.date_range(start = "1/1/2000", periods = 1000, freq = "W-WED")
long_df = pd.DataFrame(data = np.random.randn(1000, 4),
columns=['Colorado', 'Texas', 'New York', 'Ohio'],
index = dates)
long_df['5-2001']
# ### Time Series with Duplicat Indices
#
# In some applications, there may be multiple data observations falling on a particular timestamp.
#
# We can tell that the index isn't unique by checking its `is_unique` property.
dates = pd.DatetimeIndex(data = ['1/1/2000', '1/2/2000', '1/2/2000',
'1/2/2000', '1/3/2000'])
dup_ts = pd.Series(data = np.arange(5), index = dates)
dup_ts
dup_ts.index.is_unique
# ***Indexing into this time series will now either produce scalar values or slices depending on whether a timestamp is duplicated.***
#
# ***Suppose you wanted to aggregate the data having non-unique timestamp. One way to do this is to use `groupby` and pass `level = 0`(the only level of indexing)***
print("not duplicated:", dup_ts['1/3/2000'])
print('-'*50)
print("duplicated:", dup_ts['1/2/2000'])
grouped = dup_ts.groupby(level = 0)
grouped.count()
# ### Data Ranges, Frequencies, and Shifting
#
# Generic time series in pandas are assumed to be irregular, they have no fixed frequency.
#
# It often desirable to work relative to fixed frequency, duch as daily, monthly or every 15 minutes, even if that means introducing missing values into time series.
#
# Fortunately pandas has a full suite of standard time series frequencies and tools for resampling, inferring frequencies, and generating fix frequency data range.
#
# ***Converting it to be fixed daily frequency can be a accomplished by calling `resample`.***
# +
dates = [datetime(2011, 1, 2), datetime(2011, 1, 5), datetime(2011, 1, 7),
datetime(2011, 1, 8), datetime(2011, 1, 10), datetime(2011, 1, 12)]
ts = pd.Series(data = np.random.randn(6), index = dates)
ts
# -
ts.resample('1D').asfreq()[:5]
# ### Generating Date Ranges
#
# `pd.date_range` generates daily timestamps.
#
# If you pass only a start or end date, you must pass a number of periods to generate.
#
# The start and end dates defind strict boundaries for the generated date index.
pd.date_range(start = "4/1/2012", periods = 20)
pd.date_range(end = '6/1/2012', periods = 20)
pd.date_range(start = '1/1/2000', end = '12/1/2000', freq = 'BM')
# ***Sometimes you will have start or end dates with time information but want to generate a set of timestamps `normalized` to midnight as a convention.***
pd.date_range('5/2/2012 12:56:31', periods = 5, normalize = True)
# ### Frequencies and Data Offsets
#
# Frequencies in pandas are composed of a base frequency and multiplier.
#
# Base frequencies are typically refferred to by a sting alias, like `M` for monthly or `H` for hourly.
#
# For each base frequency, there is an object defind generally referred to as a `data offset`.
from pandas.tseries.offsets import Hour, Minute
hour = Hour(n = 1)
hour
# In most applications, you would never need to explicitly create one of these objects, instead using string alias like "H" or "4H".
#
# Similary, you can pass frequency like `2h30min` whcih will effectively be parsrd to the same expression.
pd.date_range(start = '1/1/2000', end = '1/3/2000', freq = '4H')
Hour(2) + Minute(30)
pd.date_range("1/1/2000", periods = 10, freq = 'MS') + Hour(15)
# ***Once useful frequency class is `week of month`, starting with `WOM`.***
#
# This enbles you to get dates like the third Friday of each month.
# WOM = week of month
rng = pd.date_range("1/1/2012", "9/1/2012", freq = 'WOM-3FRI')
rng
# ### Shifting (Leading and Lagging) Data
#
# ***`Shifting` refers to moving data backward and forward through time.***
#
# Both `Series` and `DataFrame` have a `shift` method for doing naive shifts forward or backward, leaving the index unmodified.
ts = pd.Series(data = np.random.randn(4),
index = pd.date_range("1/1/2000", periods = 4, freq = 'M'))
ts
# ***A common use of `shift` is computing percent changes in a time series or multiple time series as `DataFrame` columns.***
ts.shift(periods = 1)
ts/ts.shift(1) -1
# ***Because naive shifts leave the index unmodified, some data is discarded.***
#
# Thus if the frequency is known, it can be passed to `shift` to advance the timestamps instead of simply the data.
#
# Other frequencies can be passed, too, giving you a lot of flexibility in how to lead and lag the data.
ts.shift(periods = 2, freq = 'M')
ts.shift(3, freq = '3D')
# ### Shifting dates with offsets
#
# The pandas date offsets can also be used with `datetime` or `timestamp` objects.
from pandas.tseries.offsets import Day, MonthEnd
now = datetime.now()
now + 3*Day(n = 1)
# If you add an anchored offset like `MonthEnd`, the first increment will roll forward a date to the next date accroding to the frequency rule.
#
# ***Anchored offset can explicitly `roll` dates forward or backward using their `rollforward` and `rollback` methods.***
now + MonthEnd(1)
offset = MonthEnd(n = 3)
print("rollforward:", offset.rollforward(now))
print("-"*50)
print('rollback :', offset.rollback(now))
# ***A clever use of date offsets is to use these method with `groupby`.***
#
# ***An easier and faster way to do this is using `resample`.***
ts = pd.Series(data = np.random.randn(20),
index = pd.date_range('1/15/2000', periods = 20, freq = "4d"))
ts[-10:]
print(ts[ts.index < datetime(2000, 2, 1)].mean())
print(ts[ts.index < "2/1/2000"].mean())
print(ts[ts.index < "2000/2/1"].mean())
ts.groupby(offest.rollforward).mean()
ts.resample("M").mean()
# ### Time Zone Handling
import pytz
pytz.common_timezones[-5:]
tz = pytz.timezone(zone = 'Asia/Taipei')
tz
# ### Localization and Conversion
#
# `date_range` can be generated with a time zone set.
#
# `tz_convert` are also instance method on `DateIndex`.
rng = pd.date_range('3/9/2012 9:30', periods = 6, freq = 'D', tz = tz)
ts = pd.Series(data = np.random.randn(len(rng)), index = rng)
print(ts)
print("-"*50)
print(ts.index.tz)
# ***Conversion from naive to localized is handled by the `tz_convert` method.***
ts_utc = ts.tz_convert(tz = 'UTC')
ts_utc
ts_utc.tz_convert(tz = 'US/Eastern')
# ### Operations with Time Zone - aware Timestamp Objects
#
# ***Similar to time series and date ranges, individual Timestamp objects similarly can be localized from naive to time zone-aware and converted from one time zone to another.***
#
# If you don't defind time-zone when creating the `timestamp` or `date_range`, you can use `tz_localized` to setting time-zone.
#
# `tz_convert` use to change time zone from one to another.
stamp = pd.Timestamp('2011-03-12 04:00')
stamp_utc = stamp.tz_localize('utc')
stamp_utc
stamp_utc.tz_convert('US/Eastern')
# You can also pass time zone when creating the timestamp.
stamp_moscow = pd.Timestamp('2011-03-12 04:00', tz = 'Europe/Moscow')
stamp_moscow
# ***Time zone aware `Timestamp` objects internally store a UTC timestamp value as nanoseconds since the UNIX epoch(January, 1, 1970).***
#
# ***This UTC value is invariant between time zone conversions.***
stamp_utc.value
stamp_utc.tz_convert("US/Eastern").value
# When performing time arithmetic using pandas's `DateOffset` objects, daylight savings time transitions are respected where possible.
from pandas.tseries.offsets import Hour
stamp = pd.Timestamp('2012-03-12 01:30', tz = 'US/Eastern')
stamp
stamp + Hour(2)
# ### Operations between Different Time Zones
#
# ***If two time series with different time zones are combined, the result will be UTC.***
#
# Since the timestamps are stored under the hood in UTC, this is a straightforward operation and requires no conversion to happen.
rng = pd.date_range('3/7/2012 9:30', periods = 10, freq = "B")
rng
ts = pd.Series(np.random.randn(len(rng)), index = rng)
ts[-3:]
ts1 = ts[:7].tz_localize('Europe/London')
print("ts1:", ts1)
print("ts1 dtype : ", ts1.index.dtype)
print("-"*50)
ts2 = ts1[2:].tz_convert('Europe/Moscow')
print("ts2:", ts2)
print("ts2 dtype : ", ts2.index.dtype)
print("-"*50)
result = ts1 + ts2
print('result:', result)
print('result dtype:', result.index.dtype)
# ### Periods and Period Arithmetic
#
# `Periods` represent time span, like days, months, quarters, or years.
#
# The `Period` class represents this day type, requiring a string or integer and a frequency from the above table.
#
# In this case, the `Period` object represents the full timespan from January 1, 2007 at December 31, 2007, inclusive.
#
# Conveniently, adding and subtracting integers from periods has the effect of shifting by their frequency.
p = pd.Period(value = 2007, freq = 'A-DEC')
p
p + 5
# If two periods have the **same frequency**, their difference is the number of units between them.
pd.Period(value = '2014', freq = 'A-DEC') - p
# Regular ranges of periods can be constructed using the `period_range` function.
#
# ***The `PeriodIndex` class store a sequence of period and can serve as an axis index in any `pandas` data structure.***
rng = pd.period_range(start = '1/1/2000', end = '6/30/2000', freq = 'M')
rng
pd.Series(data = np.random.randn(6), index = rng)
# ***If you have any array of strings, you can also appeal to the `PeriodIndex` class itself.***
values = ['2001Q3', '2002Q2', '2003Q1']
index = pd.PeriodIndex(values, freq = 'Q-DEC')
index
# ### Period Frequency Conversion
#
# ***`Period` and `PeriodIndex` objects can be converted to another frequency using their `asfreq` method.***
#
# You can think of `Period("2007", "A-DEC")` as being a cursor pointing to a span of time, subdivided by monthly periods.
#
# For a `fiscal year` ending on a month other than December, the monthly subperiods belonging are different.
p = pd.Period('2007', freq = 'A-JUN')
p
p.asfreq(freq = "M", how = 'start')
p.asfreq(freq = 'M', how = 'end')
p = pd.Period(value = '2007-08', freq = 'M')
p.asfreq("A-JAN")
rng = pd.period_range('2006', '2009', freq='A-DEC')
ts = pd.Series(np.random.randn(len(rng)), index = rng)
ts
ts.asfreq(freq = 'M', how = 'start')
ts.asfreq(freq = 'B', how = 'end')
# ### Quarterly Period Frequencies
#
# Quarterly data is standard in accounting, finance, and other fields.
#
# Much quarterly data is reported relative to a `fiscal year end`, typeically the last calender or business day of one of 12 months of the year.
#
# As such, the period `2012Q4` has a different meaning depending on fiscal year end.
p = pd.Period('2012Q4', freq = "Q-JAN")
p
p.asfreq('D', 'start')
p.asfreq('D', 'end')
p4pm = (p.asfreq("B", 'e') - 1).asfreq("T", 's') + 16*60
p4pm
p4pm.to_timestamp()
# Generating quarterly ranges works as you would expect using `period-range`.
rng = pd.period_range(start = '2011Q3', end = '2012Q4', freq = "Q-JAN")
ts = pd.Series(np.arange(len(rng)), index=rng)
ts
new_rng = (rng.asfreq("B", 'e') - 1).asfreq("T", "s") + 16*60
ts.index = new_rng.to_timestamp()
ts
# ### Converting Timestamps to Periods (and Back)
#
# `pd.Series` and `pd.DataFrame` objects indexed by timestamps can be converted to periods using the `to_period` method.
rng = pd.date_range(start = '1/1/2000', periods = 3, freq = 'M')
ts = pd.Series(data = np.random.randn(3), index=rng)
ts
pts = ts.to_period()
pts
# Since periods always refer to non-overlapping timespans, a timestamp can only belong to a single period for a given frequency.
#
# While the frequency of the new `PeriodIndex` is inferred from the timestamp by default, you can specify any frequency you want.
rng = pd.date_range('1/29/2000', periods = 3, freq = 'M')
ts2 = pd.Series(np.random.randn(3), index=rng)
ts2
pts = ts2.to_period('M')
pts
# ***To convert back to timestamp, use `to_timestamp`***
print(pts.to_timestamp(how = 'start'))
print("-"*50)
print(pts.to_timestamp(how = 'end'))
# ### Creating a PeriodIndex from array
#
# Fixed frequency data sets are sometimes stored with timespan information spread across multiple columns.
#
# ***By passing these arrays to `PeriodIndex` with a frequency, they can be combined to form an index for the `DataFrame`.***
data = pd.read_csv(".\\pydata-book\\examples\\macrodata.csv")
data[-3:]
index = pd.PeriodIndex(year = data['year'],
quarter = data['quarter'], freq = 'Q-DEC')
index
data.index = index
data[-5:]
# ### Resampling and Frequency Conversion
#
# `Resampling` refers to the process of converting a time series from one frequency to another.
#
# ***Aggregating higher frequency data to lower frequency is called `downsampling`, while converting lower frequency to higher frequency is called `upsampling`.***
#
# Not all resampling falls into either of these categories.
#
# For example, converting `W-WED`(weekly on Wednesday) to `W-FRI` is neither unsampling nor downsampling.
rng = pd.date_range(start = "1/1/2000", periods = 100, freq = 'D')
rng
# ***`resample` is flexible and high-performance method that can be used to process very large time series.***
#
ts = pd.Series(data = np.random.randn(len(rng)), index = rng)
ts[-5:]
ts.resample(rule = 'M').mean()
ts.resample(rule = "M", kind = 'period').mean()
# ### Downsampling
#
# Aggregating data to regular, lower frequency is a pretty normal time series task.
#
# The data you're aggreagting doesn't need to be fixed frequently.
#
# The desired frequency defines `bin edges` that are used to slice the time series into pieces to aggregate.
#
# To convert to monthly, `M` or `BM`, the data need to be chopped up into one month interval, and the union of intervals must make up the whole time frame.
#
# There are a couple things to think about when using `resample` to downsample data:
# * Which side of each interval is closed
# * How to lable each aggregated bin, wither with the start of the interval or the end
rng = pd.date_range("1/1/2000", periods = 12, freq = 'T')
rng
ts = pd.Series(data = np.arange(12), index = rng)
ts
# Suppose you wanted to aggregate this data into five-minute chucks or bars by taking the sum of each group.
#
# The frequency you pass define bin edges in five-minute increments.
#
# **The `right bin` edge is inclusive, so the 00:05 value is included in the 00:00 to 00:05 interval.***
#
# Passing `closed = 'ledt'` changes the interval to be closed on the left.
# +
print("-"*10 + 'Include left bin' + "-"*10)
print(ts.resample(rule = '5min', closed = 'left', label = 'right').sum())
print("-"*10 + 'Include right bin' + "-"*10)
print(ts.resample(rule = '5min', closed = 'right', label = 'right').sum())
# -
# ***Lastly, you might want to shift the result index by some amount, say subtracting one second from the right rdge to make it more clear which interval the timestamp refers to.***
#
# To do this, pass a string or date offset to `loffset`.
ts.resample(rule = '5min', loffset = '-1s', label = 'right').sum()
# the same as `loffset`
temp = ts.resample(rule = '5min', label = 'right').sum()
temp.shift(periods = -1, freq = 's')
# ### Open-Hight-Low-Close (OHLC) resampling
#
# In finance, an ubiquitous way to aggregate a time series is to compute four values for each bucket.
#
# The first(open), last(close), maximum(hight) and minimal(low) values.
#
# By passing `ohlc` method you will obtain a DataFrame having colunms containing these four aggregates, which are efficiently computed in a single sweep of the data.
#
# ***The default setting of the argument `closed` and `label` are `left`.***
ts.resample(rule = '5min', closed = 'right', label = 'right').ohlc()
ts.resample(rule = '5min', closed = 'left', label = 'left').ohlc()
# ### Resampling with Groupby
#
# An alternate way to downsample is to use pandas's `groupby` functionality.
#
# ***You can groupy by month or weekday by passing a function that accesses those fields on the time series's index.***
rng = pd.date_range('1/1/2000', periods = 100, freq = 'D')
ts = pd.Series(data = np.random.randn(len(rng)), index = rng)
ts[-5:]
print('-'*25 + 'Groupby by month' + '-'*25)
print(ts.groupby(lambda x: x.month).mean())
print('-'*25 + 'Groupby by weekday' + '-'*25)
print(ts.groupby(lambda x: x.weekday).mean())
ts.resample(rule = 'M').mean()
# ### Unsampling and Interpolation
#
# When converting from a low frequency to a highter frequency, no aggregation is needed.
frame = pd.DataFrame(data = np.random.randn(2, 4),
index = pd.date_range('2000', periods = 2, freq = 'W-WED'),
columns = ['Colorado', 'Texas', 'New York', 'Ohio'])
frame[-2:]
# Suppose you wanted to fill forward each weekly value on the non-Wednesday.
#
# ***The same filling or interpolation methods available in `fillna` and `reindex` methods are available for resampling.***
# +
df_daily = frame.resample('D').asfreq()
print(df_daily)
df_daily1 = frame.resample('D').ffill()
print(df_daily1)
# -
df_daily.fillna(method = 'ffill') # forward fill, `bfill` => backward fill
frame.resample('D').bfill()
# You can similarly choose to only fill a certain number of periods forward to limit how far to continue using an observed value.
temp = frame.resample('D')
temp.ffill(limit = 2)
print(frame.resample('W-THU').asfreq())
print("-"*25 + 'forward fill' + "-"*25)
print(frame.resample('W-THU').ffill())
print("-"*25 + 'backward fill' + "-"*25)
print(frame.resample('W-THU').ffill())
# ### Resampling with Periods
frame = pd.DataFrame(np.random.randn(24, 4),
index=pd.period_range('1-2000', '12-2001', freq='M'),
columns=['Colorado', 'Texas', 'New York', 'Ohio'])
frame[-5:]
annual_frame = frame.resample(rule = "A-DEC").mean()
annual_frame
# Upsampling is more nuanced as you must make a decision about which end of the timespan in the new frequency to place the value before resampling, just like the `asfreq` method.
# Q-DEC : Quarterly, year ending in Decmber
print("-"*20 + 'convention = "start"' + "-"*20)
print(annual_frame.resample(rule = 'Q-DEC', convention = 'start').ffill())
print("-"*20 + 'convention = "start"' + "-"*20)
print(annual_frame.resample(rule = 'Q-DEC', convention = 'end').ffill())
# Since periods refer to timespans, the rules about upsampling and downsampling are more rigid:
#
# * ***In downsampling, the target frequency must be a subperiod of the source frequency.***
# * ***In upsampling, the target frequency must be a superpriod of the source frequency.***
# The original index start from 1-2000, so the index after transforming will start from 2000Q4.
annual_frame.resample('Q-MAR').ffill()
# ### Time Series Plotting
close_px_all = pd.read_csv(".\\pydata-book\\examples\\stock_px.csv", parse_dates = True, index_col = 0)
close_px_all.head(5)
close_px = close_px_all[['AAPL', "MSFT", "XOM"]]
close_px = close_px.resample('B').ffill()
close_px.head(5)
# ***When called on a DataFrame all of the time series are drawn on a single subplot with a legend indicating which is which.***
close_px['AAPL'].plot(grid = True)
close_px.loc['2009'].plot(grid = True)
close_px['AAPL'].loc["01-2011":"03-2011"].plot()
appl_q = close_px['AAPL'].resample("Q-DEC").ffill()
appl_q['2009':].plot(grid = True)
# ### Moving Window Functions
#
# A common class of array transformations intended for time series operations are statistic and other functions evaluated over a sliding window or with exponentially decaying weights.
#
# ***Like other statistical functionsm these also automatically exclude missing data.***
#
# `rolling_mean` is one of the simplest such functions.
#
# It takes a TimeSeries or DataFrame along with a window(expressed as a number of periods).
close_px[['AAPL']].plot()
# By default functions like `rolling` require the indicated number of non-NA observations.
#
# ***This behavior can be changed to account for missing data and, in particular, the fact that you will have fewer than `window` periods of data at the begining of the time series.***
close_px['AAPL'].rolling(window = 250, center = False).mean().plot()
close_px['AAPL'].plot(grid = True, title = 'Apple Price with 250-day MA')
appl_std250 = close_px['AAPL'].rolling(window = 250, min_periods = 10).std()
print(close_px[close_px.index <= '1990-02-14']['AAPL'].std())
appl_std250[:11]
appl_std250.plot(grid = True,
title = 'Apple 250-day daily return standard deviation')
# To compute an `expanding window mean`, you can see that an expanding window is just a special case where the window is the length of the time series, but only one or more period is required to compute a value.
# Define expanding mean in terms of rolling mean
expanding_mean = lambda x: rolling_mean(x, len(x), min_periods = 1)
close_px.rolling(window = 60).mean().plot(logy = True)
# ### Exponentially-weighted functions
#
# An alternative to using a static window size with equally-weighted observations is to specify a constant `decay factor` to give more weight to more recent observations.
#
# In mathematical terms, if $ma_t$ is the moving average result at time t and x is the time series in question, each value in the reslut is computed as $ma_t = a^* ma_{t-1} + (a-1)*x_t$, where a is the decay factor.
#
# ***Since an exponentially-weighted statistic place more weight on more recent observations, it `adapts` faster to changes compared with the equal-weighted version.***
# +
fig, axes = plt.subplots(nrows=2, ncols=1, sharex=True,
sharey=True, figsize=(12, 7))
aapl_px = close_px.AAPL['2005':'2009']
ma60 = aapl_px.rolling(window = 60, min_periods = 50).mean()
ewma60 = aapl_px.ewm(span = 60).mean()
aapl_px.plot(style = 'k--', ax = axes[0], grid = True)
ma60.plot(style = 'k--', ax = axes[0], grid = True)
aapl_px.plot(style='k-', ax=axes[1])
ewma60.plot(style='k--', ax=axes[1])
axes[0].set_title('Simple MA')
axes[1].set_title('Exponentially-weight MA')
# -
# ### Binary Moving Window Functions
#
# Some statistical operator, like correlation and covariance, need to operate on two time series.
spx_px = pd.read_csv(".\\pydata-book\\examples\\spx.csv", parse_dates = True, index_col = 0)
spx_px[:3]
spx_rets = spx_px/spx_px.shift(1)-1 # compute percent changes
returns = close_px.pct_change()
spx_px.pct_change()[:3] # the same as the above
corr = returns['AAPL'].rolling(window = 125, min_periods = 100).corr(spx_rets)
corr.plot(grid = True, title = 'Six-month AAPL return correlation to S&P 500')
# need to transform to Series
spx_ser = pd.Series(spx_rets.values.flatten(), index = spx_rets.index)
corr = returns.rolling(window = 125, min_periods = 100).corr(spx_ser)
corr.plot(grid = True, title = 'Six-month return correlation to S&P 500')
# ### User-Defined Moving Window Functions
#
# ***The `rolling_apply` function provides a means to apply an array function of your own devising over a moving window.***
from scipy.stats import percentileofscore
scrore_at_2percent = lambda x: percentileofscore(x, score = 0.02, kind = 'rank')
result = returns['AAPL'].rolling(window = 250).apply(scrore_at_2percent)
result.plot(grid = True,
title = 'Percentile rank of 2% AAPL return over 1 year window')
# ### Reference
#
# - ***[Difference between `asfreq` and `resample`(by stack overflow)](https://stackoverflow.com/questions/18060619/difference-between-asfreq-and-resample)***
| Ch10 Time Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Load the data
# +
import numpy as np
import pandas as pd
table = pd.read_csv("positionData.csv", low_memory=False)
display(table.head())
# -
# ### Prepare the data extracting x and y
# +
import time
import itertools
b = time.time()
train_ratio = 0.8
array = table.values[:len(table)]
i = np.arange(len(table))
#1st position, 2nd game, 3rd parameter, but then transposed
x_cac = np.array([array[np.ix_(i,range(4))],
array[np.ix_(i,range(4, 8))],
array[np.ix_(i,range(8, 12))],
array[np.ix_(i,range(12, 16))],
array[np.ix_(i,range(16, 20))]]).transpose(1, 0, 2)
y_cac = np.array([[1, 0, 0, 0, 0], [0, 1, 0, 0, 0], [0, 0, 1, 0, 0], [0, 0, 0, 1, 0], [0, 0, 0, 0, 1]])
perms = np.array(list(itertools.permutations([0, 1, 2, 3, 4])))
e = np.arange(5)
perm = np.arange(len(perms))
#GO
x_train = np.zeros((int(len(array)*train_ratio)*120, 20))
y_train = np.zeros((int(len(array)*train_ratio)*120, 25))
for i in range(int(len(array)*train_ratio)):
x_train[i*120+perm] = x_cac[i][perms[np.ix_(perm,e)]].reshape(len(perms), 20)
y_train[i*120+perm] = y_cac[perms[np.ix_(perm,e)]].reshape(len(perms), 25)
x_test = np.zeros((int(len(array)*(1-train_ratio))*120, 20))
y_test = np.zeros((int(len(array)*(1-train_ratio))*120, 25))
for i in range(int(len(array)*(1-train_ratio))):
x_test[i*120+perm] = x_cac[i+int(len(array)*train_ratio)][perms[np.ix_(perm,e)]].reshape(len(perms), 20)
y_test[i*120+perm] = y_cac[perms[np.ix_(perm,e)]].reshape(len(perms), 25)
print("Execution time:", round(time.time()-b, 2), "seconds")
# -
# ### Shuffle the data and check it is ok
# +
def unison_shuffled_copies(a, b):
assert len(a) == len(b)
p = np.random.permutation(len(a))
return a[p], b[p]
x_train, y_train = unison_shuffled_copies(x_train, y_train)
x_test, y_test = unison_shuffled_copies(x_test, y_test)
print(x_train[-1])
print(y_train[-1])
# -
# ### Create the loaders and separate training and testing data
# +
import torch
import torch.utils.data
bs = 64
device = torch.device("cuda:0")
train = torch.utils.data.TensorDataset(torch.from_numpy(x_train), torch.from_numpy(y_train))
trainloader = torch.utils.data.DataLoader(train, batch_size=bs, shuffle=True, num_workers=0)
test = torch.utils.data.TensorDataset(torch.from_numpy(x_test), torch.from_numpy(y_test))
testloader = torch.utils.data.DataLoader(test, batch_size=bs, shuffle=True, num_workers=0)
# -
# ### Define the Neural Network
# +
import torch.nn as nn
import torch.nn.functional as F
class Net(nn.Module):
def __init__(self):
super(Net, self).__init__()
self.i = torch.arange(5).view(1, 5).to(device)
self.id = torch.zeros(5, bs, 141).to(device)
self.rune = torch.zeros(5, bs, 17).to(device)
self.spells = torch.zeros(5, bs, 10).to(device)
self.dropEmb = nn.Dropout(p=0.).train()
self.dropLoc = nn.Dropout(p=0.).train()
self.dropHid = nn.Dropout(p=0.1).train()
self.idC = nn.Linear(141, 60) #50 best
self.runeC = nn.Linear(17, 10) #9 best
self.spellsC = nn.Linear(10, 10)#10 best
self.c = nn.Linear(60+10+10, 70) #40 best
self.fc2 = nn.Linear((70)*5, 250) #40, 200 best
self.fc3 = nn.Linear(250, 25) #200 best
#self.fc4 = nn.Linear(400, 25)
def forward(self, x):
self.id.zero_()
self.rune.zero_()
self.spells.zero_()
k1 = x[:, self.i*4].long().transpose(0, 2).view(5, bs, 1)
k2 = x[:, self.i*4+1].long().transpose(0, 2).view(5, bs, 1)
k3 = x[:, self.i*4+2].long().transpose(0, 2).view(5, bs, 1)
k4 = x[:, self.i*4+3].long().transpose(0, 2).view(5, bs, 1)
self.id.scatter_(2, k1, 1)
self.rune.scatter_(2, k2, 1)
self.spells.scatter_(2, k3, 1)
self.spells.scatter_(2, k4, 1)
x = torch.cat((
self.dropLoc(F.relu(self.c(torch.cat((
self.dropEmb(F.relu(self.idC(self.id[0]))),
self.dropEmb(F.relu(self.runeC(self.rune[0]))),
self.dropEmb(F.relu(self.spellsC(self.spells[0])))
), dim=1)))),
self.dropLoc(F.relu(self.c(torch.cat((
self.dropEmb(F.relu(self.idC(self.id[1]))),
self.dropEmb(F.relu(self.runeC(self.rune[1]))),
self.dropEmb(F.relu(self.spellsC(self.spells[1])))
), dim=1)))),
self.dropLoc(F.relu(self.c(torch.cat((
self.dropEmb(F.relu(self.idC(self.id[2]))),
self.dropEmb(F.relu(self.runeC(self.rune[2]))),
self.dropEmb(F.relu(self.spellsC(self.spells[2])))
), dim=1)))),
self.dropLoc(F.relu(self.c(torch.cat((
self.dropEmb(F.relu(self.idC(self.id[3]))),
self.dropEmb(F.relu(self.runeC(self.rune[3]))),
self.dropEmb(F.relu(self.spellsC(self.spells[3])))
), dim=1)))),
self.dropLoc(F.relu(self.c(torch.cat((
self.dropEmb(F.relu(self.idC(self.id[4]))),
self.dropEmb(F.relu(self.runeC(self.rune[4]))),
self.dropEmb(F.relu(self.spellsC(self.spells[4])))
), dim=1))))
), dim=1)
"""
x = torch.cat((
F.relu(self.c(torch.cat((
F.relu(self.idC(self.id[0])),
F.relu(self.runeC(self.rune[0])),
F.relu(self.spellsC(self.spells[0]))
), dim=1))),
F.relu(self.c(torch.cat((
F.relu(self.idC(self.id[1])),
F.relu(self.runeC(self.rune[1])),
F.relu(self.spellsC(self.spells[1]))
), dim=1))),
F.relu(self.c(torch.cat((
F.relu(self.idC(self.id[2])),
F.relu(self.runeC(self.rune[2])),
F.relu(self.spellsC(self.spells[2]))
), dim=1))),
F.relu(self.c(torch.cat((
F.relu(self.idC(self.id[3])),
F.relu(self.runeC(self.rune[3])),
F.relu(self.spellsC(self.spells[3]))
), dim=1))),
F.relu(self.c(torch.cat((
F.relu(self.idC(self.id[4])),
F.relu(self.runeC(self.rune[4])),
F.relu(self.spellsC(self.spells[4]))
), dim=1)))
), dim=1)
"""
#x = self.dropHid(F.relu(self.fc2(x)))
x = F.relu(self.fc2(x))
x = F.sigmoid(self.fc3(x))
return x
net = Net()
net.to(device)
# -
# ### Define a loss function and a optimizer
# +
import torch.optim as optim
#optimizer = optim.SGD(net.parameters(), lr=0.2, momentum=0.9)
optimizer = optim.Adam(net.parameters(), amsgrad=True)
# -
# ### Train the network
# +
for epoch in range(12): # loop over the dataset multiple times
testiter = iter(testloader)
running_loss = 0.0 # mean loss in the 2000 batches between printing and printing
bTime = time.time()
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to(device), labels.to(device)
# zero the parameter gradients
optimizer.zero_grad()
# forward + backward + optimize
outputs = net(inputs)
loss = F.binary_cross_entropy(outputs, labels.float())
loss.backward()
optimizer.step()
# print statistics
running_loss += loss.item()
if i % 200 == 199: # print every 100 mini-batches
#print(time.time()-bTime, "iter")
#bTime = time.time()
print('[%d, %5d] loss: %.3f' %
(epoch + 1, i + 1, running_loss / 200))
running_loss = 0.0
if i % 4000 == 3999:
lossc = 0
count = 0
net.train(False)
for i in range(1000):
images, labels = testiter.next()
images, labels = images.to(device), labels.to(device)
outputs = net(images)
lossc += F.binary_cross_entropy(outputs, labels.float()).item()
count += 1
print('test loss: '+str(round(lossc/count, 4)))
net.train(True)
if i > len(trainloader)-2*bs:
break
print('Finished Training')
# -
# ### Test the network
# +
lossc = 0
correct = torch.zeros(6).to(device)
cache = torch.zeros(bs).to(device).long()
count = 0
bTime = time.time()
net.train(False)
for data in testloader:
images, labels = data
images, labels = images.to(device), labels.to(device)
outputs = net(images)
lossc += F.binary_cross_entropy(outputs, labels.float()).item()
cache.zero_()
for i in np.arange(0, 25, 5):
cache += (torch.argmax(outputs[:, i:i+5], 1) == torch.argmax(labels[:, i:i+5], 1)).long()
correct += torch.bincount(cache).float()
if count > 2000:
break
count += 1
net.train(True)
print(round(time.time()-bTime, 2))
print('loss: '+str(round(lossc/count, 5)))
print('5 correct: '+str(round(correct[5].item()/(count*bs)*100, 2))+'%')
print('4 correct: '+str(round(correct[4].item()/(count*bs)*100, 2))+'%')
print('3 correct: '+str(round(correct[3].item()/(count*bs)*100, 2))+'%')
print('2 correct: '+str(round(correct[2].item()/(count*bs)*100, 2))+'%')
print('1 correct: '+str(round(correct[1].item()/(count*bs)*100, 2))+'%')
print('0 correct: '+str(round(correct[0].item()/(count*bs)*100, 2))+'%')
# -
# ### Save the weights
# +
import pickle
with open('99.71', 'wb') as fp:
pickle.dump(list(net.parameters()), fp)
# -
| position inference/NNwithPERMS.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Run trained model on test data and visualize
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" executionInfo={"elapsed": 883, "status": "ok", "timestamp": 1578784778530, "user": {"displayName": "<NAME> 1320262643", "photoUrl": "", "userId": "12069756592370329757"}, "user_tz": 300} id="PqrxTSb8pEXX" outputId="28b38dc5-4bdd-45f8-af78-6f324c136b15"
# Import libs
import os
import time
import cv2
from tqdm import tqdm
import numpy as np
import skimage.draw
import random
import keras
import matplotlib.pyplot as plt
import matplotlib.colors
from skimage.transform import resize
import efficientnet.tfkeras
from tensorflow.keras.models import load_model
from scipy import ndimage as ndi
from skimage.segmentation import watershed
from skimage.feature import peak_local_max
from PIL import Image, ImagePalette
NUCLEI_PALETTE = ImagePalette.random()
def create_directory(directory):
'''
Creates a new folder in the specified directory if the folder doesn't exist.
INPUT
directory: Folder to be created, called as "folder/".
OUTPUT
New folder in the current directory.
'''
if not os.path.exists(directory):
os.makedirs(directory)
# Name experiment
experiment_name = "exp-1"
# Define paths
dataset_name = "test_images"
base_path = os.path.abspath(".")
test_dataset_path = os.path.join(base_path, "dataset", dataset_name)
log_path = os.path.join(base_path, "logs", experiment_name)
print(test_dataset_path)
# -
model = None
model = load_model('{}/{}.h5'.format(log_path, experiment_name), compile=False)
#model.summary()
# +
def pad(img, pad_size=96):
"""
Load image from a given path and pad it on the sides, so that eash side is divisible by 96 (network requirement)
if pad = True:
returns image as numpy.array, tuple with padding in pixels as(x_min_pad, y_min_pad, x_max_pad, y_max_pad)
else:
returns image as numpy.array
"""
if pad_size == 0:
return img
height, width = img.shape[:2]
if height % pad_size == 0:
y_min_pad = 0
y_max_pad = 0
else:
y_pad = pad_size - height % pad_size
y_min_pad = int(y_pad / 2)
y_max_pad = y_pad - y_min_pad
if width % pad_size == 0:
x_min_pad = 0
x_max_pad = 0
else:
x_pad = pad_size - width % pad_size
x_min_pad = int(x_pad / 2)
x_max_pad = x_pad - x_min_pad
img = cv2.copyMakeBorder(img, y_min_pad, y_max_pad, x_min_pad, x_max_pad, cv2.BORDER_REFLECT_101)
return img, (x_min_pad, y_min_pad, x_max_pad, y_max_pad)
def unpad(img, pads):
"""
img: numpy array of the shape (height, width)
pads: (x_min_pad, y_min_pad, x_max_pad, y_max_pad)
@return padded image
"""
(x_min_pad, y_min_pad, x_max_pad, y_max_pad) = pads
height, width = img.shape[:2]
return img[y_min_pad:height - y_max_pad, x_min_pad:width - x_max_pad]
def read_nuclei(path):
"read raw data"
# Load 4-channel image
img = skimage.io.imread(path)
# input image
if len(img.shape) > 2:
img = img[:,:,:3]
# mask
else:
# do nothing
pass
return img
def save_nuclei(path, img):
"save image"
skimage.io.imsave(path, img)
def sliding_window(image, step, window):
x_loc = []
y_loc = []
cells = []
for y in range(0, image.shape[0], step):
for x in range(0, image.shape[1], step):
cells.append(image[y:y + window[1], x:x + window[0]])
x_loc.append(x)
y_loc.append(y)
return x_loc, y_loc, cells
def extract_patches(image, step, patch_size):
patches = []
# Get locations
x_pos, y_pos, cells = sliding_window(image, step, (patch_size[0], patch_size[1]))
for (x, y, cell) in zip(x_pos, y_pos, cells):
# Get patch
patch = image[y:y + patch_size[0], x:x + patch_size[0]]
# Get size
raw_dim = (patch.shape[1], patch.shape[0]) # W, H
#print(raw_dim)
#print(patch.shape)
if raw_dim != (patch_size[0], patch_size[1]):
# Resize to 64x64
#patch = cv2.resize(patch, (64, 64), interpolation = cv2.INTER_AREA)
patch, pad_locs = pad(patch, pad_size=patch_size[0])
# Do stuffffff
patches.append(patch)
else:
# Do stuffffff
patches.append(patch)
patches = np.array(patches)
return patches
# Compute Panoptic quality metric for each image
def Panoptic_quality(ground_truth_image,predicted_image):
TP = 0
FP = 0
FN = 0
sum_IOU = 0
matched_instances = {}# Create a dictionary to save ground truth indices in keys and predicted matched instances as velues
# It will also save IOU of the matched instance in [indx][1]
# Find matched instances and save it in a dictionary
for i in np.unique(ground_truth_image):
if i == 0:
pass
else:
temp_image = np.array(ground_truth_image)
temp_image = temp_image == i
matched_image = temp_image * predicted_image
for j in np.unique(matched_image):
if j == 0:
pass
else:
pred_temp = predicted_image == j
intersection = sum(sum(temp_image*pred_temp))
union = sum(sum(temp_image + pred_temp))
IOU = intersection/union
if IOU> 0.5:
matched_instances [i] = j, IOU
# Compute TP, FP, FN and sum of IOU of the matched instances to compute Panoptic Quality
pred_indx_list = np.unique(predicted_image)
pred_indx_list = np.array(pred_indx_list[1:])
# Loop on ground truth instances
for indx in np.unique(ground_truth_image):
if indx == 0:
pass
else:
if indx in matched_instances.keys():
pred_indx_list = np.delete(pred_indx_list, np.argwhere(pred_indx_list == [indx][0]))
TP = TP+1
sum_IOU = sum_IOU+matched_instances[indx][1]
else:
FN = FN+1
FP = len(np.unique(pred_indx_list))
PQ = sum_IOU/(TP+0.5*FP+0.5*FN)
return PQ
# +
# SAME CODE BLOCK AS IN 6_inference.ipynb
import numpy as np
from skimage.transform import resize
# Helper function for data visualization
def visualize(**images):
"""Plot images in one row."""
norm=plt.Normalize(0,4) # 5 classes including BG
map_name = matplotlib.colors.LinearSegmentedColormap.from_list("", ["black", "red","yellow","blue", "green"])
n = len(images)
plt.figure(figsize=(18, 16))
for i, (name, image) in enumerate(images.items()):
plt.subplot(1, n, i + 1)
plt.xticks([])
plt.yticks([])
plt.title(' '.join(name.split('_')).title())
plt.imshow(image, cmap=map_name, norm=norm)
plt.show()
def prep(img):
img = img.astype('float32')
img = (img > 0.5).astype(np.uint8) # threshold
img = resize(img, (image_cols, image_rows), preserve_range=True)
return img
def visualize_results(image, mask):
f, axarr = plt.subplots(1,2, figsize=(16, 16))
norm=plt.Normalize(0,4) # 5 classes including BG
map_name = matplotlib.colors.LinearSegmentedColormap.from_list("", ["black", "red","yellow","blue", "green"])
axarr[0].imshow(image)
axarr[1].imshow(mask, cmap=map_name, norm=norm)
def vis_gray(image, mask):
f, axarr = plt.subplots(1,2, figsize=(16, 16))
axarr[0].imshow(image)
axarr[1].imshow(mask, cmap='gray')
def predict(im):
"""Predict on patch"""
im = np.expand_dims(im, axis=0)
im = model.predict(im)
im = np.argmax(im.squeeze(), axis=-1)
#assert im.shape == (96, 96), "Wrong shape, {}!".format(im.shape)
return im
def instance_seg(image):
distance = ndi.distance_transform_edt(image)
local_maxi = peak_local_max(distance, indices=False, footprint=np.ones((3, 3)), labels=image)
markers = ndi.label(local_maxi)[0]
labels = watershed(-distance, markers, mask=image)
return labels
def whole_slide_predict(whole_image):
#import pdb; pdb.set_trace()
# If input image less than patch, infer on whole image
if whole_image.shape[0] < 96 or whole_image.shape[1] < 96:
# Get size
raw_dim = (whole_image.shape[1], whole_image.shape[0]) # W, H
# Resize to 64x64 for prediction
#whole_image_rs = cv2.resize(whole_image, (64, 64), interpolation = cv2.INTER_AREA)
whole_image_rs, pad_locs = pad(whole_image, pad_size=96)
# Infer
pred = predict(whole_image_rs)
# Resize back to original shape
#pred = cv2.resize(pred, raw_dim, interpolation = cv2.INTER_AREA)
pred = unpad(pred, pad_locs)
# Change dtype for resizing back to original shape
pred = pred.astype(np.uint8)
else:
# Get patch locations
x_pos, y_pos, cells = sliding_window(whole_image, 96, (96, 96))
# Array for storing predictions
pred = np.zeros((whole_image.shape[0], whole_image.shape[1])).astype(np.uint8)
# Slide over each patch
for (x, y, cell) in zip(x_pos, y_pos, cells):
# Get patch
patch = whole_image[y:y + 96, x:x + 96]
# Get size
raw_dim = (patch.shape[1], patch.shape[0]) # W, H
# If less than patch size, resize and then run prediction
if raw_dim != (96, 96):
# Resize to 64x64
#patch_rs = cv2.resize(patch, (64, 64), interpolation = cv2.INTER_AREA)
patch_rs, pad_locs = pad(patch, pad_size=96)
#print(patch.dtype, processed.dtype)
assert patch.dtype == patch_rs.dtype, "Wrong data type after resizing!"
# Infer
processed = predict(patch_rs)
# Resize back to original shape
#processed = cv2.resize(processed, raw_dim, interpolation = cv2.INTER_AREA)
processed = unpad(processed, pad_locs)
# Change dtype
processed = processed.astype(np.uint8)
assert patch.shape[:2] == processed.shape, "Wrong shape!"
assert patch.dtype == processed.dtype, "Wrong data type in prediction!"
else:
# Infer
processed = predict(patch)
# Change dtype
processed = processed.astype(np.uint8)
#print(patch.dtype, processed.dtype)
assert patch.shape[:2] == processed.shape, "Wrong shape!"
assert patch.dtype == processed.dtype, "Wrong data type in prediction!"
# Add in image variable
pred[y:y + 96, x:x + 96] = processed
processed = None
return pred
# -
# ### Run inference on a random single image
# +
image_fns = sorted(next(os.walk(test_dataset_path))[2])
idx = random.randrange(len(image_fns))
print("Index: ",idx)
image = skimage.io.imread(os.path.join(test_dataset_path, image_fns[idx]))
print("Image shape:", image.shape)
pred = whole_slide_predict(image)
print(pred.dtype)
# Post processing to refine predictions
pred_filt = cv2.medianBlur(pred.astype(np.uint8), 5)
print(image.shape, pred.shape)
print("Uniques predicted", np.unique(pred))
assert image.shape[:2] == pred.shape, "Image missmatch"
#visualize_results(image, pred)
visualize(
image=image,
Predicted_mask = pred,
Filtered_mask = pred_filt
)
# +
#inst_mask = instance_seg(pred_filt)
#print(inst_mask.shape)
#im = Image.fromarray(inst_mask.astype(np.uint8), mode='P')
#im.putpalette(NUCLEI_PALETTE)
#im
# +
# Dummy mask
zero_mask = np.zeros((pred_filt.shape[0], pred_filt.shape[1])).astype(np.uint8)
# Overlay target class
epi_mask = np.where(pred_filt != 1, zero_mask, 1)
lym_mask = np.where(pred_filt != 2, zero_mask, 2)
neu_mask = np.where(pred_filt != 3, zero_mask, 3)
macro_mask = np.where(pred_filt != 4, zero_mask, 4)
# Get uniques for (debugging)
print(epi_mask.shape, lym_mask.shape, neu_mask.shape, macro_mask.shape)
# Get instances for each class using watershed
epi_mask = instance_seg(epi_mask)
lym_mask = instance_seg(lym_mask)
neu_mask = instance_seg(neu_mask)
macro_mask = instance_seg(macro_mask)
print(epi_mask.shape, lym_mask.shape, neu_mask.shape, macro_mask.shape)
# Add color to instances
epi_mask = Image.fromarray(epi_mask.astype(np.uint8), mode='P')
epi_mask.putpalette(NUCLEI_PALETTE)
lym_mask = Image.fromarray(lym_mask.astype(np.uint8), mode='P')
lym_mask.putpalette(NUCLEI_PALETTE)
neu_mask = Image.fromarray(neu_mask.astype(np.uint8), mode='P')
neu_mask.putpalette(NUCLEI_PALETTE)
macro_mask = Image.fromarray(macro_mask.astype(np.uint8), mode='P')
macro_mask.putpalette(NUCLEI_PALETTE)
# +
def visualize_inst(**images):
"""Plot images in one row."""
norm=plt.Normalize(0,4) # 5 classes including BG
map_name = matplotlib.colors.LinearSegmentedColormap.from_list("", ["black", "red","yellow","blue", "green"])
n = len(images)
plt.figure(figsize=(30, 24))
for i, (name, image) in enumerate(images.items()):
if name == "GT_mask" or name == "Predicted_mask":
plt.subplot(1, n, i + 1)
plt.xticks([])
plt.yticks([])
plt.title(' '.join(name.split('_')).title())
plt.imshow(image, cmap=map_name, norm=norm)
else:
plt.subplot(1, n, i + 1)
plt.xticks([])
plt.yticks([])
plt.title(' '.join(name.split('_')).title())
plt.imshow(image)
plt.savefig("others/result2.png", bbox_inches = 'tight', pad_inches = 0.2, dpi=300)
plt.show()
visualize_inst(
image=image,
#GT_mask = gt,
Predicted_mask = pred_filt,
Epithelial_mask = epi_mask,
Lymphocyte_mask = lym_mask,
Macrophage_mask = neu_mask,
Neutrophil_mask = macro_mask)
# -
# ### Infer all test images
for i in tqdm(range(len(image_fns[:]))):
image = skimage.io.imread(os.path.join(test_dataset_path, image_fns[i]))
pred = whole_slide_predict(image)
#visualize_results(image, pred)
# Post processing to refine predictions
pred_filt = cv2.medianBlur(pred.astype(np.uint8), 5)
print(image.shape, pred.shape)
print("Uniques predicted", np.unique(pred))
visualize(
image=image,
Predicted_mask = pred,
Filtered_mask = pred_filt
)
assert image.shape[:2] == pred.shape, "Image missmatch"
# +
from pynotify import send_email, send_email_with_attachment
subject = "Experiment results"
message = "Done!"
dest = "<EMAIL>"
# sends an email
send_email(dest, subject, message)
| 4_inference.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
##EXTRACTING SCRIPT -> IMAGECOLLECTION -> IMAGE -> GOOGLEDRIVE
# +
import ee
import geemap
import os
Map = geemap.Map()
Map
# -
def maskL8sr(image):
# Bits 3 and 5 are cloud shadow and cloud, respectively.
cloudShadowBitMask = (1 << 3)
cloudsBitMask = (1 << 5)
# Get the pixel QA band.
qa = image.select('pixel_qa')
# Both flags should be set to zero, indicating clear conditions.
mask = qa.bitwiseAnd(cloudShadowBitMask).eq(0) \
.And(qa.bitwiseAnd(cloudsBitMask).eq(0))
return image.updateMask(mask)
#MASKING FUNCTION FOR L5 AND L7
def cloudMaskL457(image):
qa = image.select('pixel_qa')
# If the cloud bit (5) is set and the cloud confidence (7) is high
# or the cloud shadow bit is set (3), then it's a bad pixel.
cloud = qa.bitwiseAnd(1 << 5) \
.And(qa.bitwiseAnd(1 << 7)) \
.Or(qa.bitwiseAnd(1 << 3))
# Remove edge pixels that don't occur in all bands
mask2 = image.mask().reduce(ee.Reducer.min())
return image.updateMask(cloud.Not()).updateMask(mask2)
# +
#setting variables
pointA = [12.37558, 45.51725]
pointB = [12.46914, 45.47393]
region = ee.Geometry.Rectangle([pointA[0], pointA[1], pointB[0], pointB[1]])
folder = "Test_Folder"
scale = 30
################ CHANGE THIS FOR HOW MANY ITERATION WILL BE MADE #####################
starting_year = 2019
# -
def exportLandsatImage(year):
if year<1999:
dataset = ee.ImageCollection('LANDSAT/LT05/C01/T1_SR') \
.filterDate(str(year)+'-01-01', str(year)+'-12-31') \
.map(cloudMaskL457) \
image = dataset.select(['B3', 'B2', 'B1']).median()
vis = {
'bands': ['B3', 'B2', 'B1'],
'gamma': 1.4,
'max': 3000,
'min': 0
}
elif year<2013 :
dataset = ee.ImageCollection('LANDSAT/LE07/C01/T1_SR') \
.filterDate(str(year)+'-01-01', str(year)+'-12-31') \
.map(cloudMaskL457) \
image = dataset.select(['B3', 'B2', 'B1']).median()
vis = {
'bands': ['B3', 'B2', 'B1'],
'gamma': 1.4,
'max': 3000,
'min': 0
}
else :
dataset = ee.ImageCollection('LANDSAT/LC08/C01/T1_SR') \
.filterDate(str(year)+'-01-01', str(year)+'-12-31') \
.map(maskL8sr) \
dataset_bands = dataset.select(['B4', 'B3', 'B2'])
image = ee.ImageCollection.toBands(dataset_bands)
vis = {
'bands': ["B4","B3","B2"],
'gamma': 1.4,
'max': 3000,
'min': 0
}
out_dir = os.path.join(os.path.expanduser('~'), 'Downloads')
name = str(year) + '_landsat.tif'
filename = os.path.join(out_dir, name)
image = image.clip(region).unmask()
print("wooooooooooop")
geemap.ee_export_image(image, filename = filename, scale=scale, region=region, file_per_band=False)
for year in range(starting_year,2020,1):
#gets right landset and exports image
exportLandsatImage(year)
#creates image of water map and exports
dataset_water = ee.ImageCollection("JRC/GSW1_2/YearlyHistory") \
.filterDate(str(year)+'-01-01', str(year)+'-12-31')
image_water = dataset_water.select(['waterClass']).median()
| Jupyter/.ipynb_checkpoints/ExtractLocalEachYear-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.6 64-bit
# language: python
# name: python38664bitdc5bcfc3f08f4be2983655d129be0c8b
# ---
# +
import numpy as np
import pandas as pd
from sklearn import preprocessing
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
# %matplotlib inline
matplotlib.style.use('ggplot')
np.random.seed(34)
# -
#create columns of various distributions
df = pd.DataFrame({
'beta': np.random.beta(5, 1, 1000) * 60, # beta
'exponential': np.random.exponential(10, 1000), # exponential
'normal_p': np.random.normal(10, 2, 1000), # normal platykurtic
'normal_l': np.random.normal(10, 10, 1000), # normal leptokurtic
})
df.head()
# +
# make bimodal distribution
first_half = np.random.normal(20, 3, 500)
second_half = np.random.normal(-20, 3, 500)
bimodal = np.concatenate([first_half, second_half])
df['bimodal'] = bimodal
# create list of column names to use later
col_names = list(df.columns)
# +
# plot original distribution plot
fig, (ax1) = plt.subplots(ncols=1, figsize=(10, 8))
ax1.set_title('Original Distributions')
sns.kdeplot(df['beta'], ax=ax1)
sns.kdeplot(df['exponential'], ax=ax1)
sns.kdeplot(df['normal_p'], ax=ax1)
sns.kdeplot(df['normal_l'], ax=ax1)
sns.kdeplot(df['bimodal'], ax=ax1);
# -
df.mean()
df.info()
df.describe()
df.plot()
# # Add a feature with much larger values
normal_big = np.random.normal(1000000, 10000, (1000,1)) # normal distribution of large values
df['normal_big'] = normal_big
col_names.append('normal_big')
df['normal_big'].plot(kind='kde')
df.normal_big.mean()
# +
# plot original distribution plot with larger value feature
fig, (ax1) = plt.subplots(ncols=1, figsize=(10, 8))
ax1.set_title('Original Distributions')
sns.kdeplot(df['beta'], ax=ax1)
sns.kdeplot(df['exponential'], ax=ax1)
sns.kdeplot(df['normal_p'], ax=ax1)
sns.kdeplot(df['normal_l'], ax=ax1)
sns.kdeplot(df['bimodal'], ax=ax1);
sns.kdeplot(df['normal_big'], ax=ax1);
# -
df.plot()
df.describe()
# # MinMaxScaler
# +
mm_scaler = preprocessing.MinMaxScaler()
df_mm = mm_scaler.fit_transform(df)
df_mm = pd.DataFrame(df_mm, columns=col_names)
fig, (ax1) = plt.subplots(ncols=1, figsize=(10, 8))
ax1.set_title('After MinMaxScaler')
sns.kdeplot(df_mm['beta'], ax=ax1)
sns.kdeplot(df_mm['exponential'], ax=ax1)
sns.kdeplot(df_mm['normal_p'], ax=ax1)
sns.kdeplot(df_mm['normal_l'], ax=ax1)
sns.kdeplot(df_mm['bimodal'], ax=ax1)
sns.kdeplot(df_mm['normal_big'], ax=ax1);
# -
df_mm['beta'].min()
df_mm['beta'].max()
mins = [df[col].min() for col in df.columns]
mins
mixs = [df[col].max() for col in df.columns]
mixs
# # Let's check the minimums and maximums for each column after MinMaxScaler.
mins = [df_mm[col].min() for col in df_mm.columns]
mins
mixs = [df_mm[col].max() for col in df_mm.columns]
mixs
# # RobustScaler
# +
r_scaler = preprocessing.RobustScaler()
df_r = r_scaler.fit_transform(df)
df_r = pd.DataFrame(df_r, columns=col_names)
fig, (ax1) = plt.subplots(ncols=1, figsize=(10, 8))
ax1.set_title('After RobustScaler')
sns.kdeplot(df_r['beta'], ax=ax1)
sns.kdeplot(df_r['exponential'], ax=ax1)
sns.kdeplot(df_r['normal_p'], ax=ax1)
sns.kdeplot(df_r['normal_l'], ax=ax1)
sns.kdeplot(df_r['bimodal'], ax=ax1)
sns.kdeplot(df_r['normal_big'], ax=ax1);
# -
mins = [df_r[col].min() for col in df_r.columns]
mins
mixs = [df_r[col].max() for col in df_r.columns]
mixs
# # StandardScaler
# +
s_scaler = preprocessing.StandardScaler()
df_s = s_scaler.fit_transform(df)
df_s = pd.DataFrame(df_s, columns=col_names)
fig, (ax1) = plt.subplots(ncols=1, figsize=(10, 8))
ax1.set_title('After StandardScaler')
sns.kdeplot(df_s['beta'], ax=ax1)
sns.kdeplot(df_s['exponential'], ax=ax1)
sns.kdeplot(df_s['normal_p'], ax=ax1)
sns.kdeplot(df_s['normal_l'], ax=ax1)
sns.kdeplot(df_s['bimodal'], ax=ax1)
sns.kdeplot(df_s['normal_big'], ax=ax1);
# -
mins = [df_s[col].min() for col in df_s.columns]
mins
maxs = [df_s[col].max() for col in df_s.columns]
maxs
# # Normalizer
#
# +
n_scaler = preprocessing.Normalizer()
df_n = n_scaler.fit_transform(df)
df_n = pd.DataFrame(df_n, columns=col_names)
fig, (ax1) = plt.subplots(ncols=1, figsize=(10, 8))
ax1.set_title('After Normalizer')
sns.kdeplot(df_n['beta'], ax=ax1)
sns.kdeplot(df_n['exponential'], ax=ax1)
sns.kdeplot(df_n['normal_p'], ax=ax1)
sns.kdeplot(df_n['normal_l'], ax=ax1)
sns.kdeplot(df_n['bimodal'], ax=ax1)
sns.kdeplot(df_n['normal_big'], ax=ax1);
# -
mins = [df_n[col].min() for col in df_n.columns]
mins
mixs = [df_n[col].max() for col in df_n.columns]
mixs
# # Combined Plot
# +
# Combined plot.
fig, (ax0, ax1, ax2, ax3) = plt.subplots(ncols=4, figsize=(20, 8))
ax0.set_title('Original Distributions')
sns.kdeplot(df['beta'], ax=ax0)
sns.kdeplot(df['exponential'], ax=ax0)
sns.kdeplot(df['normal_p'], ax=ax0)
sns.kdeplot(df['normal_l'], ax=ax0)
sns.kdeplot(df['bimodal'], ax=ax0)
sns.kdeplot(df['normal_big'], ax=ax0);
ax1.set_title('After MinMaxScaler')
sns.kdeplot(df_mm['beta'], ax=ax1)
sns.kdeplot(df_mm['exponential'], ax=ax1)
sns.kdeplot(df_mm['normal_p'], ax=ax1)
sns.kdeplot(df_mm['normal_l'], ax=ax1)
sns.kdeplot(df_mm['bimodal'], ax=ax1)
sns.kdeplot(df_mm['normal_big'], ax=ax1);
ax2.set_title('After RobustScaler')
sns.kdeplot(df_r['beta'], ax=ax2)
sns.kdeplot(df_r['exponential'], ax=ax2)
sns.kdeplot(df_r['normal_p'], ax=ax2)
sns.kdeplot(df_r['normal_l'], ax=ax2)
sns.kdeplot(df_r['bimodal'], ax=ax2)
sns.kdeplot(df_r['normal_big'], ax=ax2);
ax3.set_title('After StandardScaler')
sns.kdeplot(df_s['beta'], ax=ax3)
sns.kdeplot(df_s['exponential'], ax=ax3)
sns.kdeplot(df_s['normal_p'], ax=ax3)
sns.kdeplot(df_s['normal_l'], ax=ax3)
sns.kdeplot(df_s['bimodal'], ax=ax3)
sns.kdeplot(df_s['normal_big'], ax=ax3);
# -
| LogisticRegression/Scale & Standardize or Normalize with scikit-learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ベイズ最適化入門
# https://github.com/Ma-sa-ue/practice/blob/master/machine%20learning(python)/bayeisan_optimization.ipynb
# The original code is based on python2. A few modifications to fit it to python3 are needed.
# %matplotlib inline
# %run ../common/homemade_GPR.py
# %run ../common/homemade_BO.py
import sys
import matplotlib.pyplot as plt
np.random.seed(seed=123)
#Define data, supervised data
def x2y(x):
f = 40.0*np.sin(x/1.0) - (0.3*(x+6.0))**2 - (0.2*(x-4.0))**2 - 1.0*np.abs(x+2.0) + np.random.normal(0,1,1)
return f
#
xmin = -20
xmax = 20
Nx = 1000
x = np.linspace(xmin, xmax, Nx)
y = list(map(x2y,x)) #for python3
y = np.array(y)
plt.plot(x, y) #### plot true data
plt.show()
#Define GPR and Bayesian opt.
GPR = Gaussian_Process_Regression(alpha = 1.0e-8)
#GPR.a1_RBF = 0.0
typical_scale=0.1
GPR.a1_RBF = 1.0
GPR.a2_RBF = typical_scale**2
GPR.a1_exp = 0.0
GPR.a2_exp = typical_scale
GPR.a1_const = 0.0
print(GPR.a1_RBF, GPR.a2_RBF, GPR.a1_exp, GPR.a2_exp, GPR.a1_const)
#
BO = Bayesian_opt()
BO.acqui_name = 'EI'
#BO.acqui_name = 'PI'
#BO.acqui_name = 'UCB'
print('# The choice of acquisition function: ',BO.acqui_name)
# +
#Definition of array as the initial condition
x_sample_init = np.array([])
y_sample_init = np.array([])
Ninitial = 2
for i in range(Ninitial):
x_point = np.random.uniform(xmin,xmax) #Initial point is randomely chosen
x_sample_init = np.append(x_sample_init,x_point)
y_point = x2y(x_point)
y_sample_init = np.append(y_sample_init,y_point)
#
Nepoch = 16 #Number of optimization
nplotevery = Nepoch//16 #Plot the results in every this number
mean, std, x_point, y_point, maxval_list = DO_BO(GPR, BO, x2y, x, x_sample_init, y_sample_init, Nepoch, nplotevery, answer_is_there=True)
plt.figure()
plt.plot(maxval_list)
plt.grid()
plt.show()
# -
| Project2/Bopt_EI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 2D Ferromagnetic Ising on a PBC Square Lattice
#
# $$H = -J\sum_{\langle ij\rangle} \sigma_i\sigma_j$$
#
# By inspection, we see that the ground states of this system correspond to all spins pointing in the same direction.
# +
import numpy as np
import matplotlib.pyplot as plt
from ising_animator import IsingAnimator
from abstract_ising import AbstractIsing
# %matplotlib inline
# -
class Ising2DPBC(AbstractIsing):
def __init__(self, Nx, Ny, J=1.):
self.J, self.Nx, self.Ny = J, Nx, Ny
self.num_spins = self.Nx * self.Ny
# initialize system at infinite temperature
# i.e. spins are completely random and uncorrelated
self.spins = 2*(np.random.rand(self.Nx, self.Ny) < 0.5) - 1
def energy(self):
"""Returns the energy of the current spin configuration"""
horizontal = np.sum(np.roll(self.spins, 1, axis=0) * self.spins)
vertical = np.sum(np.roll(self.spins, 1, axis=1) * self.spins)
total = -self.J * (horizontal + vertical)
return total
def energy_diff(self, i, j):
"""Returns the energy difference resulting from flipping the site at coordinates (i,j)"""
# sum the nearest neighbour sites
nn_sum = (
self.spins[i-1, j]
+ self.spins[(i+1) % self.Nx, j]
+ self.spins[i, j-1]
+ self.spins[i, (j+1) % self.Ny]
)
return 2 * self.J * self.spins[i, j] * nn_sum
def rand_site(self):
"""Selects a site in the lattice at random"""
i = np.random.randint(self.Nx)
j = np.random.randint(self.Ny)
return i, j
ising = Ising2DPBC(10, 10, J=1)
ising.spins
# perform 1000 MC steps
for t in range(1000):
# take a look at the abstract_ising.py file to see how mc_step works
E = ising.mc_step(T=1.0)
if t % 50 == 0:
print(E)
ising.spins
# +
n_steps = 100
t = np.arange(n_steps+1)
T0 = 100
T_min = 0.01
T = T0 * ((T_min/T0) ** (t/n_steps))
T
# -
plt.plot(t, T)
plt.title("Exponential Decay Annealing")
plt.ylabel("$T$")
plt.xlabel("time");
# +
# reinitialize so we're back at a T=infinity state
ising = Ising2DPBC(10, 10, J=1)
print(ising.spins) # print the starting configuration
# NOTE: sometimes the animation gets stuck displaying a single image
# The most reliable way to fix this is by restarting the notebook.
# Initializing the Ising Model in the same cell as the one where you
# run the animation also seems to work.
animator = IsingAnimator(ising)
animator.run_animation(T)
# -
# If the ground state was found, we should see the plot completely fill up with a single color. Since the 2D Ising Model is fairly simple, this happens quite often. In some cases however, we may see the formation of distinct domains, in which there is a straight line separating two domains of opposite spin. An example of this is shown in the video below:
# +
from IPython.display import Video
Video("domain_wall.mp4")
# -
# States with domain separation correspond to local minima of the energy. Ideally, our annealer should be able to help us escape these local minima and find the global minimum (the ground state).
| Project_4_Ising_Annealer/Task_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# # In The Name Of ALLAH
# - Scrapping The Datatable of https://www.worldometers.info/coronavirus/
#
# ## Imports
# - Selenium to get the website content by web driver
# - BeutifulSoup to scrap the data from the Html content
# - time to simulate in specific time {Showing Us before its implement}
# - Pandas to save the data scrapped as a CSV
# selenium
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from selenium.webdriver.chrome.options import Options
#bs
from bs4 import BeautifulSoup
from bs4 import element
# time
import time
# Pandas
import pandas as pd
# ## Creating The Driver and adding the Browser driver options
# Options
#Options = webdriver.ChromeOptions()
#Options.add_argument('--ignore-certificate-error')
#Options.add_argument('--incognitio')
#Options.add_argument('--headless')
#driver = webdriver.Chrome(options= Options)
chromeOption = Options()
#chromeOption.add_argument('--kiosk')
driver = webdriver.Chrome(options= chromeOption)
# # Open the Site to Scrap it
# +
driver.get('https://www.worldometers.info/coronavirus/')
# -
#if you need to getting the data of yesterday
# ## Getting the Table Content
PGsrc = driver.page_source
PageSrc = BeautifulSoup( PGsrc , "lxml")
HtmlTable =PageSrc.find(id ='main_table_countries_today')
PGsrc = driver.page_source
PageSrc = BeautifulSoup( PGsrc , "lxml")
HtmlTable =PageSrc.find(id ='main_table_countries_yesterday')
# HtmlTable.prettify() show the content of TableHtml
# ## Scrapping The Thead to get the attributes names
# - Getting table headers as list
# - trying to extract the strings from them
#
# +
Covid_headers = HtmlTable.thead.find_all('th')
Covid_headers
# -
# > Observing a `br` element in the middile of each header
for header in Covid_headers:
header.br.decompose()
print(header)
# +
HL =[]
for header in Covid_headers:
HL.append(header.text)
HL
# -
# > Good we made it, But the names must be so clear enouph, removing the mistakes
def remove_mis(word):
return str(word).replace(',','').replace('Other','').replace('\xa0','').replace('/','_').replace('\n','')
HL = list(map(remove_mis, HL))
HL
# ## Scrap the Body of the table
Covid_body = HtmlTable.tbody.find_all('tr')
Covid_body[0]
# > We need a List of Lists to apply all the body
#
BL =[[]]
i=0
for trow in Covid_body:
x = []
for tdata in trow.find_all('td'):
x.append(tdata.text)
i = i+1
BL.append(x)
BL[1]
BL[0]
BL.remove([])
BL[0]
# > Good Let's save it as CSV file
# - But first we need columns to save not rows
#
# +
table = {}
BC =[[]]
for col in range(len(HL)):
BC.append([])
for row in BL:
BC[col].append(row[col])
# -
# ## We just need some of prints
print(BC[0])
print('\n')
print(len(BC[0]))
print(BC[1])
print('\n')
print(len(BC[1]))
# > creating the Dictionary
CovidTuple = []
for col in range(len(HL)):
x =(HL[col] ,BC[col])
CovidTuple.append(x)
CovidT = dict(CovidTuple)
CovidT['Country']
# ## amazing so work no let's craete the dataframe
CovidDF = pd.DataFrame(CovidT , columns= HL)
CovidDF
name = str(time.ctime()).replace(' ', '').replace(':','_') +'.csv'
CovidDF.to_csv(name)
name = '8_4_2020Covid19.csv'
CovidDF.to_csv(name)
| Scrapping/Covid19.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Q3pharIQvnBr" outputId="57a20471-e7d7-4821-ed5b-bd9e9090ba45"
import pandas as pd
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import numpy as np
def cross_validation(datapts,gtsplit):
datapts_split = np.array_split((datapts),10)
gtsplit = np.array_split((gtsplit),10)
return datapts_split,gtsplit
def inputdata(index,datasplit,gtsplit):
train = np.array(np.vstack([x for i,x in enumerate(datasplit) if i != index]))
trainclasslabel = np.array(np.concatenate([x for i,x in enumerate(gtsplit) if i != index]))
test = np.array(datasplit[index])
testclasslabel = np.array(gtsplit[index])
# print("cv train type",train.dtype)
return train,trainclasslabel,test,testclasslabel
#normalization definition
def normalize(df): #normalizaion function
result = df.copy()
for column in df.columns:
max_value = df[column].max()
min_value = df[column].min()
result[column] = (df[column] - min_value) / (max_value - min_value)
return result
def Accuracy(y_true,y_pred):
count = 0
for i in range(len(y_pred)):
if y_pred[i] == y_true[i]:
count +=1
accuracy = count / float(len(y_true))
print("accuray:",accuracy)
return accuracy
def Recall(y_true,y_pred):
cm = ConfusionMatrix(y_true,y_pred)
recall = np.diag(cm) / np.sum(cm, axis = 1)
recall = np.mean(recall)
print("recall:",recall)
return recall
def Precision(y_true,y_pred):
cm = ConfusionMatrix(y_true,y_pred)
precision = np.diag(cm) / np.sum(cm, axis = 0)
precision = np.mean(precision)
print("precision:",precision)
return precision
def fscore(prec,recall):
# cm = ConfusionMatrix(y_true,y_pred)
f1score = 2 * (prec * recall) / (prec + recall)
print("f1score:",f1score)
return f1score
def ConfusionMatrix(y_true,y_pred):
y_pred = np.array(y_pred)
y_true = np.asarray(y_true)
target = len(np.unique(y_true))
a = (y_true * target) + y_pred
g = np.sort(a)
sq = target * target
hist, bin_edges = np.histogram(a, bins=range(g[0].astype('int'), g[0].astype('int')+ sq + 1))
hist = np.reshape(hist, (target, target))
return hist
def KNN(X_train,X_test,Y_train, k):
n = len(X_test)
dists = np.zeros((n, len(X_train)))
y_pred = np.zeros(n)
dists = np.sqrt((X_test**2).sum(axis=1)[:, np.newaxis] + (X_train**2).sum(axis=1) - 2 * X_test.dot(X_train.T))
for i in range(n):
k_neighbour = []
index_arr = np.argsort(dists[i, :], axis = 0)
k_neighbour = Y_train[index_arr[:k]].tolist()
y_pred[i] = (max(set(k_neighbour), key = k_neighbour.count))
return y_pred
#demo execution
# pf1 = pd.read_csv("/content/project3_dataset3_train.txt", delimiter = "\t", header=None)
# pf2 = pd.read_csv("/content/project3_dataset3_test.txt", delimiter = "\t", header=None)
# x_train = pf1.iloc[:,:-1]
# y_train = pf1.iloc[:,-1]
# x_test = pf2.iloc[:,:-1]
# y_test = pf2.iloc[:,-1]
# x_train = x_train.values
# y_train = y_train.values
# x_test = x_test.values
# y_test = y_test.values
# k = [2,7,8,9]
# for i in k:
# print("k =",i)
# y_pred = KNN(x_train,x_test,y_train,int(i))
# Accuracy(y_test,y_pred)
# prec = Precision(y_test,y_pred)
# rec = Recall(y_test,y_pred)
# fscore(prec,rec)
#project dataset exection
pf1 = pd.read_csv("/content/project3_dataset1.txt", delimiter = "\t", header=None) #change the file path for new file
features = pf1.iloc[:,:-1] #train features split
train_labels = pf1.iloc[:,-1]
k = 3 #change the k value as required
#processing to resolve categorical values
for index,element in pf1.iteritems():
if element.dtype == np.object:
features = pd.concat([features,pd.get_dummies(element,prefix = index)],axis = 1)
features.drop([index],axis=1,inplace=True)
features = features.astype(float)
#train labels split
features = normalize(features) #normalizing the train data
# print(features)
datafold,classfold = cross_validation(features,train_labels)
avg_accuracy = avg_precision = avg_recall = avg_f1score = 0
for index in range(10):
x_train, y_train, x_test, y_test = inputdata(index,datafold,classfold)
result = KNN(x_train,x_test,y_train,k)
knn_confusion_matrix = ConfusionMatrix(y_test, result)
acc = Accuracy(y_test, result)
prec = Precision(y_test, result)
recall = Recall(y_test, result)
f1score = fscore(prec,recall)
print("\n")
# print(result)
avg_accuracy += acc
avg_precision += prec
avg_recall += recall
avg_f1score += f1score
print("average accuracy:",avg_accuracy * 10 )
print("average pecision:",avg_precision * 10)
print("average recall:",avg_recall * 10)
print("average f1score:",avg_f1score * 0.1)
| project3/project3/Codes/KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:PythonData] *
# language: python
# name: conda-env-PythonData-py
# ---
import pandas as pd
from IPython.display import HTML
Cities=pd.read_csv("cities.csv")
Cities
C=Cities.to_html(classes='table table-responsive table-sm table-dark')
C("cities2.html", index=False)
| Resources/to html.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .sh
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Bash
# language: bash
# name: bash
# ---
# <img style="float: right;" src="img/git.png">
#
# Git kata cz. 2 - poznajmy się bliżej
# ==============
# <br>
#
# ### *Developer Days, 22.01.2019*
# <br><br>
# <NAME>
# <br><br>
# [@wyhasany](https://twitter.com/wyhasany)
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Git stash~~
# * [x] ~~Git add interactive~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [ ] Git rebase
# * [ ] `git rebase`
# * [ ] co oznacza `fast forward`
# * [ ] Git bisect
# * [ ] Git fetch vs pull
# * [ ] Praca z wieloma repozytoriami
# * [ ] ...
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="img/basic-rebase.png" />
#
# > Źródło Git Pro 2nd Edition
# + slideshow={"slide_type": "subslide"}
cd ~/ && rm -rf rebase/ && mkdir rebase && cd rebase && git init && git config commit.gpgsign false
echo C0 > file
git add file
git commit -am C0
echo C1 >> file
git commit -am C1
echo C2 >> file
git commit -am C2
echo C3 >> file
git commit -am C3
git checkout -b experiment
echo C4 >> file
git commit -am C4
git log --all --graph --oneline
# + slideshow={"slide_type": "subslide"}
#git checkout master
#git rebase experiment
#git log --oneline --graph --all
# + slideshow={"slide_type": "fragment"}
git checkout master
echo C5 >> file
git commit -am C5
git log --oneline --graph --all
# + slideshow={"slide_type": "subslide"}
git checkout experiment
git rebase master
#git rebase experiment
# + slideshow={"slide_type": "fragment"}
cat file
# + slideshow={"slide_type": "subslide"}
sed -i '5d' file #usuń czwartą linię itd
sed -i '6d' file
sed -i '7d' file
git add file
git rebase --continue
git log --oneline --all --graph
# + slideshow={"slide_type": "fragment"}
git checkout master
git merge experiment
echo
git log --oneline --all --graph
# + slideshow={"slide_type": "subslide"}
cd ~/ && rm -rf rebase/
# -
# ## Ćwiczenia
#
# 1. Stwórz nowe repozytorium, załóż branch master z plikiem a.txt i dodaj w kolejnych commitach linie `1`, `2`, `3` z takimi samymi nazwami commitów. Załóż brach `experiment` i dołóż kolejne commity z liniami `4` i `5`. Przenieś się na branch master i wykonaj `git rebase experiment` powiedz co się stało i dlaczego.
# + [markdown] slideshow={"slide_type": "subslide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Merge (łączenie zmian)~~
# * [x] ~~Git stash~~
# * [x] ~~Git add interactive~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~`git rebase`~~
# * [x] ~~co oznacza `fast forward`~~
# * [ ] Git bisect
# * [ ] Git fetch vs pull
# * [ ] Praca z wieloma repozytoriami
# * [ ] Git push
# * [ ] ...
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [ ] Git bisect
# * [ ] `git bisect`
# * [ ] `^` operator
# * [ ] `~` operator
# * [ ] `git show`
# * [ ] Git fetch vs pull
# * [ ] Praca z wieloma repozytoriami
# * [ ] Git push
# * [ ] ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Przykład
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf bisect && mkdir bisect && cd bisect && git init && git config commit.gpgsign false
touch projectfile
git add projectfile
cat > test.sh << EOF
if grep 63 projectfile
then
echo BAD
else
echo GOOD
fi
EOF
chmod +x test.sh
git add test.sh
for ((i=1;i<=100;i++))
do
echo $i >> projectfile
git commit -am "A$i"
done
git log --oneline
# + slideshow={"slide_type": "subslide"}
git bisect start
./test.sh
git bisect bad
git status
# + slideshow={"slide_type": "fragment"}
git checkout HEAD~99
git status
./test.sh
git bisect good
git log --oneline
# + slideshow={"slide_type": "subslide"}
git status
./test.sh
git bisect good
git log --oneline
./test.sh
git bisect bad
# + slideshow={"slide_type": "subslide"}
./test.sh
git bisect good
./test.sh
git bisect bad
./test.sh
git bisect bad
./test.sh
git bisect bad
./test.sh
git bisect bad
# + slideshow={"slide_type": "subslide"}
git log --oneline
# + slideshow={"slide_type": "subslide"}
git diff HEAD^ HEAD
# + [markdown] slideshow={"slide_type": "fragment"}
# <center><img src="img/git-blame.jpg" width="600px" alt="Hunk WTF?!" style="position:center" /></center>
# + slideshow={"slide_type": "subslide"}
tree .git -L 1
tree .git/refs
# + slideshow={"slide_type": "subslide"}
find .git/ -type f | grep BISECT | xargs -I % sh -c "echo '\n>> % <<' ; cat %"
# + slideshow={"slide_type": "fragment"}
git status
git bisect reset
git status
tree .git -L 1
tree .git/refs
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Bisect automatyczny
# + slideshow={"slide_type": "fragment"}
cat > ../test_authomatic.sh << EOF
if ./test.sh | grep BAD
then
exit 1
else
exit 0
fi
EOF
chmod +x ../test_authomatic.sh
git bisect start HEAD HEAD~99 --
git bisect run ./../test_authomatic.sh
git status
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf bisect && rm test_authomatic.sh
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Operator `~` i `^`
# + slideshow={"slide_type": "fragment"}
rm -rf bisect-2 && mkdir bisect-2 && cd bisect-2 && git init && git config commit.gpgsign false
touch aplik
git add aplik
git commit -am 'Init'
git checkout -b agalaz
echo 'agalaz bla'>> aplik
git commit -am 'agalaz bla'
git checkout master
echo 'master bla 1' >> aplik
git commit -am 'master bla 1'
echo 'master bla 2' >> aplik
git commit -am 'master bla 2'
git merge -X ours -m merged agalaz
git log --oneline --all --graph
# + slideshow={"slide_type": "subslide"}
git show --quiet --oneline HEAD^
git show --quiet --oneline HEAD^^
git show --quiet --oneline HEAD^^^
git show --quiet --oneline HEAD^2
git show --quiet --oneline HEAD^2^
git show --quiet --oneline HEAD~1
git show --quiet --oneline HEAD~2
git show --quiet --oneline HEAD~2^1
git show --quiet --oneline HEAD^2~1
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf bisect-2
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Ćwiczenia
#
# Do domu
#
# Przygotuj analogiczny kod (jak w przykładzie) w języku programowania, którego używasz na co dzień, zrób test wskazujący na wystąpienie błędu i znajdź automatycznie commit, który wprowadził błąd.
#
# lub
#
# Poczytaj o git bisect
#
# https://git-scm.com/docs/git-bisect
#
# https://git-scm.com/docs/git-bisect-lk2009.html
#
# ...ale w domu i tak spróbuj zrobić przykład wyżej! 🤘
# + [markdown] slideshow={"slide_type": "subslide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~`git bisect`~~
# * [x] ~~`^` operator~~
# * [x] ~~`~` operator~~
# * [x] ~~`git show`~~
# * [ ] Git fetch vs pull
# * [ ] Praca z wieloma repozytoriami
# * [ ] Git push
# * [ ] Git submodule
# * [ ] ...
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Git add interactive~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [ ] Git fetch vs pull
# * [ ] `git fetch`
# * [ ] `git pull`
# * [ ] Praca z wieloma repozytoriami
# * [ ] Git push
# * [ ] Git submodule
# * [ ] ...
# + slideshow={"slide_type": "subslide"}
cd ~/ && rm -rf pull && mkdir pull && cd pull
mkdir git_origin && cd git_origin && git init && git config commit.gpgsign false
echo 'pierwszy commit' > plik
git add plik
git commit -am plik
cd ..
git clone git_origin git_cloned
tree
diff git_origin/plik git_cloned/plik
# + slideshow={"slide_type": "subslide"}
diff git_origin/.git/config git_cloned/.git/config
# + slideshow={"slide_type": "fragment"}
cd git_cloned
git config commit.gpgsign false
git remote
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Fetch vs Pull
#
# * `fetch` pobiera dane nie ruszając naszego repo w oddzielne miejsce (szczegóły w krótce)
# * `pull` pobiera zmiany ze zdalnego repo dla wybranego brancha i merge'uje do aktualnego brancha
# + slideshow={"slide_type": "fragment"}
pwd
git remote -v
# + slideshow={"slide_type": "fragment"}
cd ../git_origin
echo 'fetchable change' >> plik
git commit -am fetchable
git log --oneline --graph --all
# + slideshow={"slide_type": "subslide"}
cd ../git_cloned
git fetch origin master
# + slideshow={"slide_type": "fragment"}
git log --oneline --graph --all
git branch --all
tree .git/refs
# + slideshow={"slide_type": "subslide"}
git merge origin/master
git log --oneline --graph --all
# + slideshow={"slide_type": "fragment"}
cd ../git_origin
echo 'pullable change' >> plik
git commit -am 'pull'
git log --oneline --graph --all
cd ../git_cloned
git pull origin master
git log --oneline --graph --all
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf pull
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Ćwiczenia
#
# Do domu:
#
# 1. Pobierz dowolne repo z githuba, które ma więcej niż dwa branche
# 2. Utwórz nową gałąź `atom`
# 3. Zrób `git fetch origin jakaś_gałąż`
# 4. Zrób merge do aktualnej gałęzi
# 5. Zrób `git pull origin inna_gałąź`
# 6. Odpowiedz sobie jakie są różnice w obu podejściach.=
# + [markdown] slideshow={"slide_type": "subslide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Git add interactive~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~`git fetch`~~
# * [x] ~~`git pull`~~
# * [ ] Praca z wieloma repozytoriami
# * [ ] Git push
# * [ ] Git submodule
# * [ ] ...
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] Praca z wieloma repozytoriami
# * [ ] tworzenie wielu remote'ów
# * [ ] przerzucanie zmian pomiędzy wieloma remote'ami
# * [ ] Git push
# * [ ] Git submodule
# * [ ] Git log
# * [ ] Git squash
# * [ ] ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Przykład
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf pushing && mkdir pushing && cd pushing
mkdir git-origin && cd git-origin && git init && git config commit.gpgsign false
echo 'first commit' > file
git add file
git commit -am file
cd ..
git clone git-origin git-clone
# + slideshow={"slide_type": "fragment"}
cd git-origin
git branch -a -v
cd ../git-clone
git config commit.gpgsign false
git branch -a -v
cd ..
# + slideshow={"slide_type": "subslide"}
cd git-clone
git checkout -b abranch
echo 'cloned abranch commit' >> file
git commit -am 'cloned abranch commit'
git push origin abranch
# + [markdown] slideshow={"slide_type": "fragment"}
# ### A co jeśli remote ma inny content?
# + slideshow={"slide_type": "subslide"}
cd ..
rm -rf git-origin git-clone
mkdir git-origin && cd git-origin
git init && git config commit.gpgsign false
echo 'first commit' > file
git add file
git commit -am file
cd ..
git clone git-origin git-clone
cd git-clone
git config commit.gpgsign false
git checkout -b abranch
echo 'cloned abranch commit' >> file
git commit -am 'cloned abranch commit'
cd ../git-origin
git checkout -b abranch
echo 'origin abranch commit' >> file
git commit -am 'origin abranch commit'
cd ../git-clone
git push origin abranch:abranch
# + slideshow={"slide_type": "subslide"}
git fetch origin
git branch -v -a
echo
git log --oneline --graph --all
# + slideshow={"slide_type": "fragment"}
git merge remotes/origin/abranch
# + slideshow={"slide_type": "subslide"}
cat file
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Trzy połączone repozytoria - przykład
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf remotes && mkdir remotes && cd remotes
mkdir git-origin && cd git-origin && git init && git config commit.gpgsign false
echo 'first commit' > file
git add file
git commit -am file
cd ..
git clone git-origin alice-cloned
git clone git-origin bob-cloned
#Alice dodaje zmiany na swoim masterze
cd alice-cloned
git config commit.gpgsign false
echo alice-change >> file
git commit -am 'alice change'
git log --oneline --graph --all
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Jak Bob może pobrać zmiany Alice?
# + [markdown] slideshow={"slide_type": "fragment"}
# 1. DODAJ repo Alice jako remote u Boba
# 2. FETCH zmiany od Alice z gałęzi master
# 3. MERGE tych zmian w lokalnym repozytorium Boba (lub zamiast 2. i 3. użyć `git pull`)
# + slideshow={"slide_type": "subslide"}
cd ../bob-cloned
git config commit.gpgsign false
git remote add alice ../alice-cloned
git remote -v
# + slideshow={"slide_type": "fragment"}
git fetch alice
# + slideshow={"slide_type": "fragment"}
git branch -vv -a
# + slideshow={"slide_type": "subslide"}
git merge alice/master
#git merge remotes/alice/master
# + slideshow={"slide_type": "fragment"}
cat file
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf remotes
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Ćwiczenia
#
# 1. Stwórz repozytorium gita i sklonuj je obok. Zrób zmianę w pierwszym repozytorium na gałęzi master, sfetch'uj ją, a następnie zmerge'uj
# 2. Stwórz zmianę w jednym pliku na obu repozytoriach, która wywoła konflikt. Rozwiąż konflikt po obu stronach tak, aby repozytoria były spójne.
# 3. Zrób to samo co w 2. tylko na trzech repozytoriach (np. na dwóch klonach)
# + [markdown] slideshow={"slide_type": "subslide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~tworzenie wielu remote'ów~~
# * [x] ~~przerzucanie zmian pomiędzy wieloma remote'ami~~
# * [ ] Git push
# * [ ] Git submodule
# * [ ] Git log
# * [ ] ...
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [ ] Git push
# * [ ] `git push`
# * [ ] błędy przy `git push`
# * [ ] Git submodule
# * [ ] Git log
# * [ ] ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Przykład
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf pushing && mkdir pushing && cd pushing
mkdir git-origin && cd git-origin && git init && git config commit.gpgsign false
echo 'first commit' > file
git add file
git commit -am file
cd ..
git clone git-origin git-clone
# + slideshow={"slide_type": "fragment"}
cd git-origin
git branch -a -v
cd ../git-clone
git config commit.gpgsign false
git branch -a -v
cd ..
# + slideshow={"slide_type": "subslide"}
cd git-clone
git checkout -b abranch
echo 'cloned abranch commit' >> file
git commit -am 'cloned abranch commit'
git push origin abranch
# + [markdown] slideshow={"slide_type": "fragment"}
# ### A co jeśli remote ma inny content?
# + slideshow={"slide_type": "subslide"}
cd ..
rm -rf git-origin git-clone
mkdir git-origin && cd git-origin
git init && git config commit.gpgsign false
echo 'first commit' > file
git add file
git commit -am file
cd ..
git clone git-origin git-clone
cd git-clone
git config commit.gpgsign false
git checkout -b abranch
echo 'cloned abranch commit' >> file
git commit -am 'cloned abranch commit'
cd ../git-origin
git checkout -b abranch
echo 'origin abranch commit' >> file
git commit -am 'origin abranch commit'
cd ../git-clone
git push origin abranch:abranch
# + slideshow={"slide_type": "subslide"}
git fetch origin
git branch -v -a
echo
git log --oneline --graph --all
# + slideshow={"slide_type": "fragment"}
git merge remotes/origin/abranch
# + slideshow={"slide_type": "subslide"}
cat file
# + slideshow={"slide_type": "fragment"}
sed -i '2d' file
sed -i '3d' file
sed -i '4d' file
cat file
git status
# + slideshow={"slide_type": "fragment"}
git commit -am 'merged conflict on file'
git log --oneline --graph --all
# + [markdown] slideshow={"slide_type": "subslide"}
# ### a co gdy gałąź istnieje tylko na remote?
# + slideshow={"slide_type": "fragment"}
cd ..
rm -rf git-origin git-clone
mkdir git-origin
cd git-origin
git init && git config commit.gpgsign false
echo 'first commit' > file
git add file
git commit -am file
cd ..
git clone git-origin git-clone
cd git-origin
git checkout -b abranch
echo 'origin abranch commit' >> file
git commit -am 'cloned abranch commit'
git branch -a
# + slideshow={"slide_type": "fragment"}
cd ../git-clone
git config commit.gpgsign false
git branch -a
echo
git remote -v
# + slideshow={"slide_type": "subslide"}
git fetch origin
# + slideshow={"slide_type": "fragment"}
git branch -a
# + slideshow={"slide_type": "fragment"}
git checkout abranch
git branch -a -vv
echo
git log --oneline --graph --all
# + [markdown] slideshow={"slide_type": "subslide"}
# ### a jak śledzić zdalne gałęzie z różnymi nazwami?
# + slideshow={"slide_type": "fragment"}
cd ..
rm -rf git-origin git-clone
mkdir git-origin
cd git-origin
git init
git config commit.gpgsign off
echo 'first commit' > file
git add file
git commit -am file
cd ..
git clone git-origin git-clone
cd git-clone
git config commit.gpgsign off
git checkout -b abranch
echo 'origin abranch commit' >> file
git commit -am 'cloned abranch commit'
git push
# + slideshow={"slide_type": "subslide"}
git branch -a -vv
# + slideshow={"slide_type": "subslide"}
#git push --set-upstream origin abranch
#git push -u origin abranch
git push origin abranch
echo
git branch -a -vv
# + slideshow={"slide_type": "fragment"}
git push -u origin abranch
echo
git branch -a -vv
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf pushing
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Ćwiczenia
#
# 1. Stwórz repozytorium na GitHub
# 2. Dodaj do niego jakąś treść (prosty plik płaski zrobiony w GUI)
# 3. Sklonuj repozytorium, stwórz gałąź i wypchmij ją na serwer zdalny
# 4. Zobacz gałąź na GitHubie
# + [markdown] slideshow={"slide_type": "subslide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [x] ~~`git push`~~
# * [x] ~~błędy przy `git push`~~
# * [ ] Git submodule
# * [ ] Git log
# * [ ] ...
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [ ] Git submodule
# * [ ] Git log
# * [ ] Git squash
# * [ ] ...
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Przykład - biblioteka
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf submodules && mkdir submodules && cd submodules && git init && git config commit.gpgsign false
echo 'A' > file
git add file
git commit -am 'a'
git checkout -b experimental
echo 'C - EXPERIMENTAL' >> file
git commit -am EXPERIMENTAL
git checkout master
echo 'B' >> file
git commit -am 'B'
git log --oneline --all --graph
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Użycie biblioteki z konkretnego brancha `experimental`
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf repo && mkdir repo && cd repo && git init && git config commit.gpgsign false
echo 'source lib' > file
git add file
git commit -am 'source lib'
echo 'do something with experimental' >> file
cat file
# + slideshow={"slide_type": "subslide"}
git submodule init
echo
git submodule add ../submodules
echo
ls -a
echo
ls -a submodules/
echo
cat .gitmodules
echo
git submodule status
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Pytania jakie powinniśmy sobie zadać
#
# 1. Jak przełączyć się na gałąż `experimental`?
# 2. Co się stanie jak coś się zmieni na `experimental`? Czy mój kod sam się zaktualizuje?
# 3. Co się stanie jeśli wprowadzę zmiany w submodule? Czy mogę pushować? Czy mogę trzymać te zmiany prywatnie w moim repo?
# 4. Co jeśli zdarzą się konflikty pomiędzy tymi repozytoriami?
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Przełączmy się na branch `experimental`
# + slideshow={"slide_type": "fragment"}
cd submodules
git branch -a -vv
# + slideshow={"slide_type": "fragment"}
git checkout experimental
#git checkout -b my_local_master --track origin/master
echo
git branch -a -vv
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Git śledzi (track) stan submodułu
# + slideshow={"slide_type": "fragment"}
cd ..
git diff
# + slideshow={"slide_type": "fragment"}
git commit -am 'lib moved to experimental'
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Wprowadzamy zmianę w bibliotece
# + slideshow={"slide_type": "fragment"}
cd ../submodules
git checkout experimental
echo 'D' >> file
git commit -am 'D - a fix added'
# + slideshow={"slide_type": "fragment"}
cd ../repo/submodules
git status
# + slideshow={"slide_type": "fragment"}
git pull
# + slideshow={"slide_type": "subslide"}
git log --oneline --graph --all
cd ..
git diff
# + [markdown] slideshow={"slide_type": "fragment"}
# ### Klonowanie projektu z submodułami
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf repo-cloned
git clone repo repo-cloned
cd repo-cloned
tree
# + slideshow={"slide_type": "fragment"}
git submodule status
# + slideshow={"slide_type": "fragment"}
git submodule init
# + slideshow={"slide_type": "subslide"}
git submodule update
echo
git submodule status
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Czy da się to zrobić szybciej? `git clone --recursive`
# + slideshow={"slide_type": "fragment"}
cd ~/
rm -rf repo-recursive
git clone --recursive repo repo-recursive
echo
tree repo-recursive
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf repo && rm -rf repo-cloned && rm -rf repo-recursive && rm -rf submodules
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Ćwiczenia
#
# Stwórz dwa repozytoria, gdzie jedno ma drugie (nawzajem) jako submoduł
# + [markdown] slideshow={"slide_type": "subslide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [x] ~~Git submodule~~
# * [ ] Git log
# * [ ] Git squash
# * [ ] ...
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Reflog~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [x] ~~Git submodule~~
# * [ ] Git log
# * [ ] Git squash
# * [ ] Repozytoria bare
# * [ ] ...
# + slideshow={"slide_type": "subslide"}
cd ~/ && rm -rf keepassxc
git clone https://github.com/keepassxreboot/keepassxc.git
cd keepassxc
git log --graph --oneline --all --topo-order
# + slideshow={"slide_type": "subslide"}
git log
# + slideshow={"slide_type": "subslide"}
git log --oneline
# + slideshow={"slide_type": "subslide"}
git log --oneline --graph
# + slideshow={"slide_type": ""}
git log --graph --oneline --all > all_output
git log --graph --oneline > noall_output
diff all_output noall_output
# + slideshow={"slide_type": "fragment"}
rm all_output noall_output
# + slideshow={"slide_type": "subslide"}
git log --graph --oneline --all --decorate --source
# + slideshow={"slide_type": "subslide"}
git log --graph --oneline --all --simplify-by-decoration
# + slideshow={"slide_type": "subslide"}
git log --graph --oneline --all --simplify-by-decoration --pretty='%ar %s %h'
# + slideshow={"slide_type": "subslide"}
git log --graph --oneline --all --simplify-by-decoration --pretty=
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf keepassxc
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Ćwiczenie
#
# Użyj `git log --pretty`aby wypisać logi w keepassxc od 2018-09-01 w takim formacie:
#
# Commit Hash: #numer hasha#, Author: #autor name#, Date: #author date, RFC2822 style#
#
# hint: zajrzyj go `man git log` tam znajdziesz podpowiedzi jak to zrobić. Jak znajdziesz czas to przyjrzyj się filtrowaniu printowanych commitów
# + [markdown] slideshow={"slide_type": "subslide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [x] ~~Git submodule~~
# * [x] ~~Git log~~
# * [ ] Git squash
# * [ ] Repozytoria bare
# * [ ] ...
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [x] ~~Git submodule~~
# * [x] ~~Git log~~
# * [ ] Git squash
# * [ ] git rebase -i
# * [ ] git rev-list
# * [ ] Repozytoria bare
# * [ ] ...
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="img/99-pull-request.png" width="800px" alt="Hunk WTF?!" style="float:left; margin-right: 30px;" />
#
# ## Motywacja
#
# https://github.com/bendews/ansible-cloudflared/pull/2#issuecomment-436852893
# + slideshow={"slide_type": "subslide"}
cd ~/ && rm -rf squashing && mkdir squashing && cd squashing && git init && git config commit.gpgsign false
touch file
git add file
git commit -m 'Init'
for i in {1..10}
do
echo $i >> file
git commit -am "commit:${i}"
done
git status
git log
# + slideshow={"slide_type": "subslide"}
cd ..
rm -rf squashing-clone
git clone squashing squashing-clone
cd squashing-clone
git config commit.gpgsign false
git remote -v
echo
git log
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Aby zesquashować commity potrzebujemy
#
# * referencję do najnowszego commita ze zbiotu commitów które chcemy squashować
# * referencję do najstarszego commita...
# + slideshow={"slide_type": "fragment"}
git rev-list --max-parents=0 HEAD
# + slideshow={"slide_type": "fragment"}
git rev-list HEAD
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Terminal
#
# ```
# cd ~/squashing-clone
# git rebase -i $(git rev-list --max-parents=0 HEAD) HEAD
#
# 1 pick 8c283fa commit:1
# 2 pick 5720f0f commit:2
# 3 pick d6f975e commit:3
# 4 pick 880f333 commit:4
# 5 pick 3c8b11f commit:5
# 6 pick 7a13bc7 commit:6
# 7 pick d036f47 commit:7
# 8 pick 739a99a commit:8
# 9 pick 7fa1e78 commit:9
# 10 pick ac30399 commit:10
# 11
# 12 # Rebase 6aadfc6..ac30399 onto 6aadfc6 (10 commands)
# 13 #
# 14 # Commands:
# 15 # p, pick = use commit
# 16 # r, reword = use commit, but edit the commit message
# 17 # e, edit = use commit, but stop for amending
# 18 # s, squash = use commit, but meld into previous commit
# 19 # f, fixup = like "squash", but discard this commit's log message
# 20 # x, exec = run command (the rest of the line) using shell
# 21 # d, drop = remove commit
# 22 #
# 23 # These lines can be re-ordered; they are executed from top to bottom.
# 24 #
# 25 # If you remove a line here THAT COMMIT WILL BE LOST.
# 26 #
# 27 # However, if you remove everything, the rebase will be aborted.
# 28 #
# 29 # Note that empty commits are commented out
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ```
# 1 r 8c283fa commit:1
# 2 s 5720f0f commit:2
# 3 s d6f975e commit:3
# 4 s 880f333 commit:4
# 5 s 3c8b11f commit:5
# 6 s 7a13bc7 commit:6
# 7 s d036f47 commit:7
# 8 s 739a99a commit:8
# 9 s 7fa1e78 commit:9
# 10 s ac30399 commit:10
# 11
# 12 # Rebase 6aadfc6..ac30399 onto 6aadfc6 (10 commands)
# 13 #
# 14 # Commands:
# 15 # p, pick = use commit
# 16 # r, reword = use commit, but edit the commit message
# 17 # e, edit = use commit, but stop for amending
# 18 # s, squash = use commit, but meld into previous commit
# 19 # f, fixup = like "squash", but discard this commit's log message
# 20 # x, exec = run command (the rest of the line) using shell
# 21 # d, drop = remove commit
# 22 #
# 23 # These lines can be re-ordered; they are executed from top to bottom.
# 24 #
# 25 # If you remove a line here THAT COMMIT WILL BE LOST.
# 26 #
# 27 # However, if you remove everything, the rebase will be aborted.
# 28 #
# 29 # Note that empty commits are commented out
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ```
# 1 commit:all
# 2
# 3 # Please enter the commit message for your changes. Lines starting
# 4 # with '#' will be ignored, and an empty message aborts the commit.
# 5 #
# 6 # Date: Thu Dec 6 21:25:56 2018 +0100
# 7 #
# 8 # interactive rebase in progress; onto 6aadfc6
# 9 # Last command done (1 command done):
# 10 # reword 8c283fa commit:1
# 11 # Next commands to do (9 remaining commands):
# 12 # squash 5720f0f commit:2
# 13 # squash d6f975e commit:3
# 14 # You are currently editing a commit during a rebase.
# 15 #
# 16 # Changes to be committed:
# 17 # modified: file
# 18 #
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ```
# 1 # This is a combination of 10 commits.
# 2 # This is the 1st commit message:
# 3
# 4 commit:all
# 5
# 6 # This is the commit message #2:
# 7
# 8 commit:2
# 9
# 10 # This is the commit message #3:
# 11
# 12 commit:3
# 13
# 14 # This is the commit message #4:
# 15
# 16 commit:4
# 17
# 18 # This is the commit message #5:
# 19
# 20 commit:5
# 21
# 22 # This is the commit message #6:
# 23
# 24 commit:6
# 25
# 26 # This is the commit message #7:
# 27
# 28 commit:7
# 29
# 30 # This is the commit message #8:
# 31
# 32 commit:8
# 33
# 34 # This is the commit message #9:
# 35
# 36 commit:9
# 37
# 38 # This is the commit message #10:
# 39
# 40 commit:10
# 41
# 42 # Please enter the commit message for your changes. Lines starting
# 43 # with '#' will be ignored, and an empty message aborts the commit.
# 44 #
# 45 # Date: Thu Dec 6 21:25:56 2018 +0100
# 46 #
# 47 # interactive rebase in progress; onto 6aadfc6
# 48 # Last commands done (10 commands done):
# 49 # squash 7fa1e78 commit:9
# 50 # squash ac30399 commit:10
# 51 # No commands remaining.
# 52 # You are currently rebasing.
# 53 #
# 54 # Changes to be committed:
# 55 # modified: file
# 56 #
# ```
# + [markdown] slideshow={"slide_type": "subslide"}
# ```
# git branch -f master
# git checkout master
# ```
# + slideshow={"slide_type": "fragment"}
git log --oneline --graph --all
# + slideshow={"slide_type": "subslide"}
git push origin
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Co się stanie jak spullujemy repo i spróbujemy pchnąć je na origin?
# + slideshow={"slide_type": "fragment"}
git push -f origin
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf squashing*
# + [markdown] slideshow={"slide_type": "subslide"}
# <img src="img/not_rebase_pushed_commits.jpg" width="800px" alt="Hunk WTF?!" style="float:left; margin-right: 30px;" />
#
# ### A jak squashować kod, który jest na repo? (seems dangerous)
#
# ```
# git rebase -i origin/master~4 master
# git push origin +master
# ```
#
# * `+` robi force na jednym branchu
# * `--force` robi force na wszystkich branchach
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Ćwiczenie
#
# 1. Wykmiń jak działają pozostałe funkcje przy `git rebase -i`
# 2. Spróbuj użyć komendy `exec`
# 3. Przećwicz squashowanie
# + [markdown] slideshow={"slide_type": "subslide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [x] ~~Git submodule~~
# * [x] ~~Git log~~
# * [x] ~~Git squash~~
# * [x] ~~git rebase -i~~
# * [x] ~~git rev-list~~
# * [ ] Repozytoria bare
# * [ ] Cherry picking and Three-Way Merges
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Cherry picking~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [x] ~~Git submodule~~
# * [x] ~~Git log~~
# * [x] ~~Git squash~~
# * [ ] Repozytoria bare
# * [ ] Cherry picking and Three-Way Merges
# + slideshow={"slide_type": "subslide"}
cd ~/ && rm -rf bare-repo && mkdir bare-repo && cd bare-repo && git init --bare
tree -a
echo
cat config
# + slideshow={"slide_type": "subslide"}
cd ~/
rm -rf bare-repo-alice
rm -rf bare-repo-bob
git clone bare-repo bare-repo-alice
git clone bare-repo bare-repo-bob
# + slideshow={"slide_type": "fragment"}
cd ~/bare-repo-alice
git config commit.gpgsign false
git remote -v
touch file
git add file
git commit -m 'init'
for i in {1..10}
do
echo $i >> file
git commit -am "commit:${i}"
done
git log --oneline
git push
# + slideshow={"slide_type": "subslide"}
cd ../bare-repo-bob
git log --oneline
# + slideshow={"slide_type": "fragment"}
git fetch origin
git merge origin/master
git log --oneline
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Alice znów squashuje :)
# + slideshow={"slide_type": "fragment"}
cd ~/bare-repo-alice
git log --oneline
# + slideshow={"slide_type": "fragment"}
git push origin master
# + slideshow={"slide_type": "subslide"}
git push -f origin master
git log --oneline --graph --all
# + [markdown] slideshow={"slide_type": "fragment"}
# ## A co na to Bob?
# + slideshow={"slide_type": "subslide"}
cd ~/bare-repo-bob
git status
git pull --no-edit
# + slideshow={"slide_type": "fragment"}
git log --oneline --graph --all
# + slideshow={"slide_type": "subslide"}
git push
# + slideshow={"slide_type": "subslide"}
cd ../bare-repo-alice
git pull --no-edit
echo
git log --oneline --graph --all
echo
git status
# + slideshow={"slide_type": "fragment"}
#WTF per minute?
cd ~/
rm -rf bare-repo*
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Ćwiczenie
#
# 1. Załóż repo na GitHubie i spróbuj je pobrać przez ssh (`git://...`)
# + [markdown] slideshow={"slide_type": "slide"}
# # Status
#
# * [x] ~~...~~
# * [x] ~~Git rebase~~
# * [x] ~~Git bisect~~
# * [x] ~~Git fetch vs pull~~
# * [x] ~~Praca z wieloma repozytoriami~~
# * [x] ~~Git push~~
# * [x] ~~Git submodule~~
# * [x] ~~Git log~~
# * [x] ~~Git squash~~
# * [x] ~~Repozytoria bare~~
# * [ ] Cherry picking and Three-Way Merges
# * [ ] `cherry pick` może zrobić coś czego się nie spodziewasz
# * [ ] `git diff-tree` i `git apply` komendy
# * [ ] co to `three-way merge`
# + slideshow={"slide_type": "subslide"}
cd ~/ && rm -rf patch-and-apply && mkdir patch-and-apply && cd patch-and-apply && git init && git config commit.gpgsign false
touch afile
git add afile
git commit -m 'file added'
echo First change, on master >> afile
git commit -am 'First change, on master added'
git branch abranch
echo Second change, on master >> afile
git commit -am 'Second change, on master added'
git checkout abranch
echo First change, on abranch >> afile
git commit -am 'First change, on abranch added'
echo Second change, on abranch >> afile
echo New file, on abranch >> newfile
git add newfile
git commit -am 'Second change, on abranch added'
git tag abranchtag
echo
cat afile
echo
git checkout master
echo
cat afile
echo
git log --oneline --graph --all
# + slideshow={"slide_type": "subslide"}
git show abranchtag
# + slideshow={"slide_type": "subslide"}
git cherry-pick abranchtag
echo
cat afile
echo
cat newfile
# + slideshow={"slide_type": "subslide"}
git cherry-pick --abort
#create patch
git diff-tree -p abranchtag > abranchtag.patch
cat abranchtag.patch
# + slideshow={"slide_type": "fragment"}
cat abranchtag.patch | git apply
git status
git log --oneline --graph --all
# + slideshow={"slide_type": "subslide"}
cat abranchtag.patch | git apply --reject
# + slideshow={"slide_type": "subslide"}
cat afile.rej
# + slideshow={"slide_type": "fragment"}
cd ~/patch-and-apply
sed -i '8d' abranchtag.patch
cat abranchtag.patch | git apply
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="img/33-sha-1-collision.png" width="800px" alt="Hunk WTF?!" style="float:left; margin-right: 30px;" />
#
# ## Jak liczone są skróty commitów?
#
# Liczymy SHA1 z:
# - całego kodu źródłowego bez ignorowanych i niekomitowanych plików
# - z SHA1 ostatniego commitu
# - z informacji o autorze
# - z informacji o commiterze (tak mogą być różni 😅)
# - z treści commita
# + [markdown] slideshow={"slide_type": "subslide"}
# # Przykład
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf tmp-sha1
mkdir tmp-sha1
cd tmp-sha1
git init
touch A
git config commit.gpgsign false
git add A
git commit -am 'A'
git log
# + slideshow={"slide_type": "fragment"}
git log --format=format:%H
# + slideshow={"slide_type": "subslide"}
git cat-file commit HEAD
# + slideshow={"slide_type": "fragment"}
printf "commit %s\0" $(git cat-file commit HEAD | wc -c)
# + slideshow={"slide_type": "fragment"}
(printf "commit %s\0" $(git cat-file commit HEAD | wc -c); git cat-file commit HEAD) | sha1sum
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf tmp-sha1
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Polskie znaki w nazwach plików
# + slideshow={"slide_type": "fragment"}
cd ~/ && rm -rf git-diacritics && mkdir git-diacritics && cd git-diacritics
git init
git config commit.gpgsign false
touch Źródło
git status
cd ~/ && rm -rf git-diacritics
# + [markdown] slideshow={"slide_type": "slide"}
# ## Aliasy dla komend gita?
# + slideshow={"slide_type": "fragment"}
alias | grep git
# + [markdown] slideshow={"slide_type": "slide"}
# ## Nakładki na gita, np. IntellIJ (demo)
# + [markdown] slideshow={"slide_type": "slide"}
# # Przydatne linki
#
# <br />
#
# 1. [To nie działa WTF?!](http://justinhileman.info/article/git-pretty/git-pretty.png)
# 2. [Cheat sheet powiązany z fazami życia kontentu w gicie](http://ndpsoftware.com/git-cheatsheet.html)
# + [markdown] slideshow={"slide_type": "subslide"}
# <center><img src="img/git-pretty.png" width="900px" alt="Orange Logo" style="position:center" /></center>
| Git-kata2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # Keras Functional API
# ## Recall: All models (layers) are callables
# ```python
# from keras.layers import Input, Dense
# from keras.models import Model
#
# # this returns a tensor
# inputs = Input(shape=(784,))
#
# # a layer instance is callable on a tensor, and returns a tensor
# x = Dense(64, activation='relu')(inputs)
# x = Dense(64, activation='relu')(x)
# predictions = Dense(10, activation='softmax')(x)
#
# # this creates a model that includes
# # the Input layer and three Dense layers
# model = Model(inputs=inputs, outputs=predictions)
# model.compile(optimizer='rmsprop',
# loss='categorical_crossentropy',
# metrics=['accuracy'])
# model.fit(data, labels) # starts training
# ```
# # Multi-Input Networks
# ## Keras Merge Layer
# Here's a good use case for the functional API: models with multiple inputs and outputs.
#
# The functional API makes it easy to manipulate a large number of intertwined datastreams.
#
# Let's consider the following model.
# ```python
# from keras.layers import Dense, Input
# from keras.models import Model
# from keras.layers.merge import concatenate
#
# left_input = Input(shape=(784, ), name='left_input')
# left_branch = Dense(32, input_dim=784, name='left_branch')(left_input)
#
# right_input = Input(shape=(784,), name='right_input')
# right_branch = Dense(32, input_dim=784, name='right_branch')(right_input)
#
# x = concatenate([left_branch, right_branch])
# predictions = Dense(10, activation='softmax', name='main_output')(x)
#
# model = Model(inputs=[left_input, right_input], outputs=predictions)
# ```
# Resulting Model will look like the following network:
# <img src="../imgs/multi_input_model.png" />
# Such a two-branch model can then be trained via e.g.:
# ```python
# model.compile(optimizer='rmsprop', loss='categorical_crossentropy', metrics=['accuracy'])
# model.fit([input_data_1, input_data_2], targets) # we pass one data array per model input
# ```
# ## Try yourself
# #### Step 1: Get Data - MNIST
# +
# let's load MNIST data as we did in the exercise on MNIST with FC Nets
# +
# # %load ../solutions/sol_821.py
# -
# #### Step 2: Create the Multi-Input Network
## try yourself
# +
## `evaluate` the model on test data
# -
# Keras supports different Merge strategies:
#
# * `add`: element-wise sum
# * `concatenate`: tensor concatenation. You can specify the concatenation axis via the argument concat_axis.
# * `multiply`: element-wise multiplication
# * `average`: tensor average
# * `maximum`: element-wise maximum of the inputs.
# * `dot`: dot product. You can specify which axes to reduce along via the argument dot_axes. You can also specify applying any normalisation. In that case, the output of the dot product is the cosine proximity between the two samples.
# You can also pass a function as the mode argument, allowing for arbitrary transformations:
# ```python
# merged = Merge([left_branch, right_branch], mode=lambda x: x[0] - x[1])
# ```
# ---
# # Even more interesting
#
# Here's a good use case for the functional API: models with multiple inputs and outputs.
#
# The functional API makes it easy to manipulate a large number of intertwined datastreams.
#
# Let's consider the following model (from: [https://keras.io/getting-started/functional-api-guide/](https://keras.io/getting-started/functional-api-guide/) )
# ## Problem and Data
#
# We seek to predict how many retweets and likes a news headline will receive on Twitter.
#
# The main input to the model will be the headline itself, as a sequence of words, but to spice things up, our model will also have an auxiliary input, receiving extra data such as the time of day when the headline was posted, etc.
#
# The model will also be supervised via two loss functions.
#
# Using the main loss function earlier in a model is a good regularization mechanism for deep models.
#
# <img src="https://s3.amazonaws.com/keras.io/img/multi-input-multi-output-graph.png" width="40%" />
# +
from keras.layers import Input, Embedding, LSTM, Dense
from keras.models import Model
# Headline input: meant to receive sequences of 100 integers, between 1 and 10000.
# Note that we can name any layer by passing it a "name" argument.
main_input = Input(shape=(100,), dtype='int32', name='main_input')
# This embedding layer will encode the input sequence
# into a sequence of dense 512-dimensional vectors.
x = Embedding(output_dim=512, input_dim=10000, input_length=100)(main_input)
# A LSTM will transform the vector sequence into a single vector,
# containing information about the entire sequence
lstm_out = LSTM(32)(x)
# -
# Here we insert the auxiliary loss, allowing the LSTM and Embedding layer to be trained smoothly even though the main loss will be much higher in the model.
auxiliary_output = Dense(1, activation='sigmoid', name='aux_output')(lstm_out)
# At this point, we feed into the model our auxiliary input data by concatenating it with the LSTM output:
# +
from keras.layers import concatenate
auxiliary_input = Input(shape=(5,), name='aux_input')
x = concatenate([lstm_out, auxiliary_input])
# We stack a deep densely-connected network on top
x = Dense(64, activation='relu')(x)
x = Dense(64, activation='relu')(x)
x = Dense(64, activation='relu')(x)
# And finally we add the main logistic regression layer
main_output = Dense(1, activation='sigmoid', name='main_output')(x)
# -
# ### Model Definition
model = Model(inputs=[main_input, auxiliary_input], outputs=[main_output, auxiliary_output])
# We compile the model and assign a weight of 0.2 to the auxiliary loss.
#
# To specify different **loss_weights or loss** for each different output, you can use a list or a dictionary. Here we pass a single loss as the loss argument, so the same loss will be used on all outputs.
# #### Note:
# Since our inputs and outputs are named (we passed them a "name" argument),
# We can compile&fit the model via:
model.compile(optimizer='rmsprop',
loss={'main_output': 'binary_crossentropy', 'aux_output': 'binary_crossentropy'},
loss_weights={'main_output': 1., 'aux_output': 0.2})
# ```python
#
# # And trained it via:
# model.fit({'main_input': headline_data, 'aux_input': additional_data},
# {'main_output': labels, 'aux_output': labels},
# epochs=50, batch_size=32)
# ```
# ### Hands On - Resnet
# Deep residual networks took the deep learning world by storm when Microsoft Research released Deep Residual Learning for Image Recognition. These networks led to 1st-place winning entries in all five main tracks of the ImageNet and COCO 2015 competitions, which covered image classification, object detection, and semantic segmentation. The robustness of ResNets has since been proven by various visual recognition tasks and by non-visual tasks involving speech and language.
#
# #### Motivation
#
# Network depth is of crucial importance in neural network architectures, but deeper networks are more difficult to train. The residual learning framework eases the training of these networks, and enables them to be substantially deeper — leading to improved performance in both visual and non-visual tasks. These residual networks are much deeper than their ‘plain’ counterparts, yet they require a similar number of parameters (weights).
# The (degradation) problem:
# With network depth increasing, accuracy gets saturated (which might be unsurprising) and then degrades rapidly. Unexpectedly, such degradation is not caused by overfitting, and adding more layers to a suitably deep model leads to higher training error.
# The core insight:
# Let us consider a shallower architecture and its deeper counterpart that adds more layers onto it. There exists a solution to the deeper model by construction: the layers are copied from the learned shallower model, and the added layers are identity mapping. The existence of this constructed solution indicates that a deeper model should produce no higher training error than its shallower counterpart.
# #### The proposed solution:
#
#
# <img src="https://cdn-images-1.medium.com/max/1600/1*pUyst_ciesOz_LUg0HocYg.png">
#
#
# A residual block — the fundamental building block of residual networks. Figure 2: https://arxiv.org/pdf/1512.03385.pdf
# Instead of hoping each stack of layers directly fits a desired underlying mapping, we explicitly let these layers fit a residual mapping. The original mapping is recast into F(x)+x. We hypothesize that it is easier to optimize the residual mapping than to optimize the original, unreferenced mapping. To the extreme, if an identity mapping were optimal, it would be easier to push the residual to zero than to fit an identity mapping by a stack of nonlinear layers.
# We have reformulated the fundamental building block (figure above) of our network under the assumption that the optimal function a block is trying to model is closer to an identity mapping than to a zero mapping, and that it should be easier to find the perturbations with reference to an identity mapping than to a zero mapping. This simplifies the optimization of our network at almost no cost. Subsequent blocks in our network are thus responsible for fine-tuning the output of a previous block, instead of having to generate the desired output from scratch.
# ### Hands On - Build Resnet
#
# By the time you got here, you should be able to build Resnet and train it on MNIST.
#
# Do do :)
| day03/Advanced_Keras_Tutorial/.ipynb_checkpoints/1.0 Multi-Modal Networks-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Chess AI using Deep Learning
# > Following Victor Sim's tutorial
#
# - toc: false
# - badges: true
# - comments: true
# - author: <NAME>
# - categories: [comparison, jupyter]
# - image: images/normal-distribution-tests.png
#hide
import os
import chess
import numpy as np
import pandas as pd
from tensorflow import keras
from tensorflow.keras import layers
#hide
os.chdir('XXXXXXXXXXX')
df = pd.read_csv('chess_normalized.csv')
data = df['moves'].tolist()[:500]
split_data = []
indice = 500
#hide
chess_dict = {
'p' : [1,0,0,0,0,0,0,0,0,0,0,0],
'P' : [0,0,0,0,0,0,1,0,0,0,0,0],
'n' : [0,1,0,0,0,0,0,0,0,0,0,0],
'N' : [0,0,0,0,0,0,0,1,0,0,0,0],
'b' : [0,0,1,0,0,0,0,0,0,0,0,0],
'B' : [0,0,0,0,0,0,0,0,1,0,0,0],
'r' : [0,0,0,1,0,0,0,0,0,0,0,0],
'R' : [0,0,0,0,0,0,0,0,0,1,0,0],
'q' : [0,0,0,0,1,0,0,0,0,0,0,0],
'Q' : [0,0,0,0,0,0,0,0,0,0,1,0],
'k' : [0,0,0,0,0,1,0,0,0,0,0,0],
'K' : [0,0,0,0,0,0,0,0,0,0,0,1],
'.' : [0,0,0,0,0,0,0,0,0,0,0,0],
}
alpha_dict = {
'a' : [0,0,0,0,0,0,0],
'b' : [1,0,0,0,0,0,0],
'c' : [0,1,0,0,0,0,0],
'd' : [0,0,1,0,0,0,0],
'e' : [0,0,0,1,0,0,0],
'f' : [0,0,0,0,1,0,0],
'g' : [0,0,0,0,0,1,0],
'h' : [0,0,0,0,0,0,1],
}
number_dict = {
1 : [0,0,0,0,0,0,0],
2 : [1,0,0,0,0,0,0],
3 : [0,1,0,0,0,0,0],
4 : [0,0,1,0,0,0,0],
5 : [0,0,0,1,0,0,0],
6 : [0,0,0,0,1,0,0],
7 : [0,0,0,0,0,1,0],
8 : [0,0,0,0,0,0,1],
}
#hide
def make_matrix(board):
pgn = board.epd()
foo = []
pieces = pgn.split(" ", 1)[0]
rows = pieces.split("/")
for row in rows:
foo2 = []
for thing in row:
if thing.isdigit():
for i in range(0, int(thing)):
foo2.append('.')
else:
foo2.append(thing)
foo.append(foo2)
return foo
def translate(matrix,chess_dict):
rows = []
for row in matrix:
terms = []
for term in row:
terms.append(chess_dict[term])
rows.append(terms)
return rows
# +
#hide
for point in data[:indice]:
point = point.split()
split_data.append(point)
data = []
for game in split_data:
board = chess.Board()
for move in game:
board_ready = board.copy()
data.append(board.copy())
board.push_san(move)
trans_data = []
for board in data:
matrix = make_matrix(board)
trans = translate(matrix,chess_dict)
trans_data.append(trans)
pieces = []
alphas = []
numbers = []
# -
#hide
true_data = flatten(split_data)
for i in range(len(true_data)):
try:
term = flatten(split_data)[i]
original = term[:]
term = term.replace('x','')
term = term.replace('#','')
term = term.replace('+','')
if len(term) == 2:
piece = 'p'
else:
piece = term[0]
alpha = term[-2]
number = term[-1]
pieces.append(chess_dict[piece])
alphas.append(alpha_dict[alpha])
numbers.append(number_dict[int(number)])
except:
pass
#hide
board_inputs = keras.Input(shape=(8, 8, 12))
conv1= layers.Conv2D(10, 3, activation='relu')
conv2 = layers.Conv2D(10, 3, activation='relu')
pooling1 = layers.MaxPooling2D(pool_size=(2, 2), strides=None, padding="valid", data_format=None,)
pooling2 = layers.MaxPooling2D(pool_size=(2, 2), strides=None, padding="valid", data_format=None,)
flatten = keras.layers.Flatten(data_format=None)
x = conv1(board_inputs)
x = pooling1(x)
x = conv2(x)
x = flatten(x)
piece_output = layers.Dense(12,name = 'piece')(x)
model_pieces = keras.Model(inputs=board_inputs, outputs=piece_output, name="chess_ai_v3")
earlystop = keras.callbacks.EarlyStopping(monitor='loss', min_delta=0, patience=250, verbose=0, mode='auto', baseline=None, restore_best_weights=True)
model_pieces.compile(
loss=keras.losses.mse,
optimizer=keras.optimizers.Adam(),
metrics=None,
)
model_pieces.fit(trans_data[:len(pieces)],pieces[:len(pieces)],batch_size=64, epochs=100,callbacks = [earlystop])
clear_output()
board_inputs = keras.Input(shape=(8, 8, 12))
conv1= layers.Conv2D(10, 3, activation='relu')
conv2 = layers.Conv2D(10, 3, activation='relu')
pooling1 = layers.MaxPooling2D(pool_size=(2, 2), strides=None, padding="valid", data_format=None,)
pooling2 = layers.MaxPooling2D(pool_size=(2, 2), strides=None, padding="valid", data_format=None,)
flatten = keras.layers.Flatten(data_format=None)
x = conv1(board_inputs)
x = pooling1(x)
x = conv2(x)
x = flatten(x)
alpha_output = layers.Dense(7,name = 'alpha')(x)
model_alpha = keras.Model(inputs=board_inputs, outputs=alpha_output, name="chess_ai_v3")
earlystop = keras.callbacks.EarlyStopping(monitor='loss', min_delta=0, patience=250, verbose=0, mode='auto', baseline=None, restore_best_weights=True)
model_alpha.compile(
loss=keras.losses.mse,
optimizer=keras.optimizers.Adam(),
metrics=None,
)
model_alpha.fit(trans_data[:len(alphas)],alphas[:len(alphas)],batch_size=64, epochs=100,callbacks = [earlystop])
clear_output()
board_inputs = keras.Input(shape=(8, 8, 12))
conv1= layers.Conv2D(10, 3, activation='relu')
conv2 = layers.Conv2D(10, 3, activation='relu')
pooling1 = layers.MaxPooling2D(pool_size=(2, 2), strides=None, padding="valid", data_format=None,)
pooling2 = layers.MaxPooling2D(pool_size=(2, 2), strides=None, padding="valid", data_format=None,)
flatten = keras.layers.Flatten(data_format=None)
x = conv1(board_inputs)
x = pooling1(x)
x = conv2(x)
x = flatten(x)
numbers_output = layers.Dense(7,name = 'number')(x)
model_number = keras.Model(inputs=board_inputs, outputs=numbers_output, name="chess_ai_v3")
earlystop = keras.callbacks.EarlyStopping(monitor='loss', min_delta=0, patience=250, verbose=0, mode='auto', baseline=None, restore_best_weights=True)
model_number.compile(
loss=keras.losses.mse,
optimizer=keras.optimizers.Adam(),
metrics=None,
)
model_number.fit(trans_data[:len(numbers)],numbers[:len(numbers)],batch_size=64, epochs=100,callbacks = [earlystop])
clear_output()
# +
#hide
new_chess_dict = {}
new_alpha_dict = {}
new_number_dict = {}
for term in chess_dict:
definition = tuple(chess_dict[term])
new_chess_dict[definition] = term
new_chess_dict[term] = definition
for term in alpha_dict:
definition = tuple(alpha_dict[term])
new_alpha_dict[definition] = term
new_alpha_dict[term] = definition
for term in number_dict:
definition = tuple(number_dict[term])
new_number_dict[definition] = term
new_number_dict[term] = definition
data = np.reshape(trans_data[0],(1,8,8,12))
pred = model_pieces.predict(data)
def translate_pred(pred):
translation = np.zeros(pred.shape)
index = pred[0].tolist().index(max(pred[0]))
translation[0][index] = 1
return translation[0]
piece = translate_pred(model_pieces.predict(data))
alpha = translate_pred(model_alpha.predict(data))
number = translate_pred(model_alpha.predict(data))
piece_pred = new_chess_dict[tuple(piece)]
alpha_pred = new_alpha_dict[tuple(alpha)]
number_pred = new_number_dict[tuple(number)]
move =str(piece_pred)+str(alpha_pred)+str(number_pred)
# -
# Developed by [<NAME>](https://cleberjamaral.github.io/).
#
# Sources:
# * [Creating A Chess AI using Deep Learning - Victor Sim's article](https://towardsdatascience.com/creating-a-chess-ai-using-deep-learning-d5278ea7dcf)
# * [Portable Game Notation](https://en.wikipedia.org/wiki/Portable_Game_Notation)
| _notebooks/2021-01-19-a-chess-ai-using-deep-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.1 64-bit ('3.8')
# name: python3
# ---
import pandas as pd
import numpy as np
# Importing the data as a dataframe.
dataset = pd.read_csv("data/Data_Lv3_UMDOTS_Escooters.csv")
# Lets take a look at a few data point
#dataset.head()
dataset.tail()
dataset.shape
| practice2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine learning with Scala Spark linear regression
#
# > "How to do linear regression with Spark in a Scala application"
#
# - toc:true
# - branch: master
# - badges: false
# - comments: false
# - author: <NAME>
# - categories: [Spark, Scala, API, data-analysis, machine-learning]
# ## <a name="overview"></a> Overview
# In a <a href="https://pockerman.github.io/qubit_opus/machine-learning/scala/linear-regression/2021/06/27/ml-with-scala-linear-regression.html">previous post</a> I developed a trivial Scala application that performs linear regression with only one feature. In this post, I want to go a bit further, I want to use Spark's <a href="https://spark.apache.org/docs/latest/ml-guide.html">MLlib</a> to develop a linear regression model using two features this time.
# ## Machine learning with Scala Spark linear regression
# The first thing I need to do in order to use MLlib in my Scala application is to update the dependencies in the
# ```build.sbt``` script. These should now look as
# ```
# libraryDependencies += "org.apache.spark" % "spark-core_2.12" % "3.0.1"
# libraryDependencies += "org.apache.spark" % "spark-sql_2.12" % "3.0.1"
# libraryDependencies += "org.apache.spark" % "spark-mllib_2.12" % "3.0.1"
# ```
# ```
# package train.spark
#
# import org.apache.spark.ml.regression.LinearRegression
# import org.apache.spark.SparkContext
# import org.apache.spark.SparkContext._
# import org.apache.spark.SparkConf
# import org.apache.spark.sql.SparkSession
#
# import org.apache.spark.ml.feature.VectorAssembler
# import org.apache.spark.ml.linalg.Vectors
# import org.apache.spark.sql.types.DoubleType
#
#
# object LinearRegressionApp {
#
# def main(args: Array[String]) {
#
# val conf = new SparkConf().setAppName("Linear regression Spark")
# val sc = new SparkContext(conf)
#
# val session = SparkSession.builder().appName("Linear regression Spark").master("local[4]").getOrCreate()
#
# // Should be some file on your system
# val csvFile = "/home/alex/qi3/spark_scala/data/spark_regression.csv"
# val inputTrainigSet = session.read.format("csv").load(csvFile)
#
# println("Number of Partitions: "+inputTrainigSet.rdd.getNumPartitions)
# println("Action: First element: "+inputTrainigSet.rdd.first())
#
# val analysisData = inputTrainigSet.withColumn("x1", inputTrainigSet("_c0").cast(DoubleType))
# .withColumn("x2", inputTrainigSet("_c1").cast(DoubleType))
# .withColumn("y", inputTrainigSet("_c2").cast(DoubleType))
# .drop("_c0")
# .drop("_c1")
# .drop("_c2")
#
#
# //creating features column
# val assembler = new VectorAssembler()
# .setInputCols(Array("x1","x2"))
# .setOutputCol("features")
#
# // create the model
# val lr = new LinearRegression()
# .setMaxIter(10)
# .setRegParam(0.3)
# .setElasticNetParam(0.8)
# .setFeaturesCol("features")
# .setLabelCol("y")
#
# val trainigSet = assembler.transform(analysisData)
#
# // Fit the model
# val lrModel = lr.fit(trainigSet)
#
# // Print the coefficients and intercept for linear regression
# println(s"Coefficients: ${lrModel.coefficients} Intercept: ${lrModel.intercept}")
#
# // Summarize the model over the training set and print out some metrics
# val trainingSummary = lrModel.summary
#
# println(s"numIterations: ${trainingSummary.totalIterations}")
#
# // there is sth wrong with my scala/spark version and this
# // throws an excpetion
# //println(s"objectiveHistory: [${trainingSummary.objectiveHistory.mkString(",")}]")
#
# trainingSummary.residuals.show()
# println(s"RMSE: ${trainingSummary.rootMeanSquaredError}")
# println(s"r2: ${trainingSummary.r2}")
# }
# }
# ```
# ```
# 21/08/25 12:36:15 WARN Utils: Your hostname, LT-2R0620-101 resolves to a loopback address: 127.0.1.1; using 192.168.0.71 instead (on interface wlp58s0)
# 21/08/25 12:36:15 WARN Utils: Set SPARK_LOCAL_IP if you need to bind to another address
# WARNING: An illegal reflective access operation has occurred
# WARNING: Illegal reflective access by org.apache.spark.unsafe.Platform (file:/home/alex/MySoftware/spark-3.0.1-bin-hadoop2.7/jars/spark-unsafe_2.12-3.0.1.jar) to constructor java.nio.DirectByteBuffer(long,int)
# WARNING: Please consider reporting this to the maintainers of org.apache.spark.unsafe.Platform
# WARNING: Use --illegal-access=warn to enable warnings of further illegal reflective access operations
# WARNING: All illegal access operations will be denied in a future release
# 21/08/25 12:36:16 WARN NativeCodeLoader: Unable to load native-hadoop library for your platform... using builtin-java classes where applicable
# 21/08/25 12:36:17 WARN SparkContext: Using an existing SparkContext; some configuration may not take effect.
# Number of Partitions: 1
# Action: First element: [0.0,4.0,4.357400305044133]
# 21/08/25 12:36:22 WARN BLAS: Failed to load implementation from: com.github.fommil.netlib.NativeSystemBLAS
# 21/08/25 12:36:22 WARN BLAS: Failed to load implementation from: com.github.fommil.netlib.NativeRefBLAS
# Coefficients: [1.2545846367230242,0.7527507820338242] Intercept: 1.305736977601481
# numIterations: 3
# +--------------------+
# | residuals|
# +--------------------+
# | 0.04066019930735543|
# | -0.6631570819021908|
# | 0.8844468485401586|
# |-0.27725408848247746|
# | 1.523792089069631|
# | 0.9081058052618962|
# | 0.6154843963633212|
# | -1.5426210882366824|
# | -1.116750516169644|
# | -0.5438006575317718|
# |-0.41191237820348237|
# |-0.10423573938951769|
# | -0.7720329729420263|
# | -0.5175509972153742|
# | 0.5066514385552212|
# | 0.28386941829179424|
# | -1.7266735995448794|
# | -0.7963013580643907|
# | -0.8306208671329927|
# | -0.7913153349720496|
# +--------------------+
# only showing top 20 rows
#
# RMSE: 1.0241722775198268
# r2: 0.8486882566011
#
# ```
| _notebooks/2021-08-25-ml-scala-spark-linear-regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import math
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.pipeline import Pipeline
import sys
sys.path.append("..")
from source.df_pipeline import df_imputer, df_scaler, dummify
pd.set_option('max_columns', 200)
pd.set_option('max_rows', 80)
# +
df_train = pd.read_csv('../data_processed/train_processed.csv', dtype={'WindSpeed': 'object'})
df_train.head()
# -
class transformation(TransformerMixin, BaseEstimator):
def __init__(self, mean_weight=10):
self.columns = None
self.mean_weight = mean_weight
self.smooth_team = {}
def fit(self, X, y=None):
return self
def stats_by_play(self, data):
avg_by_play = data.groupby(['PlayId',
'Team',
'offense_team'], as_index=False)[['PlayerHeight',
'PlayerWeight',
'age',
'S', 'A']].mean()
spread = data.groupby(['PlayId',
'Team',
'offense_team'])[['X', 'Y']].std().reset_index()
tot_momentum = data.groupby(['PlayId',
'Team',
'offense_team'], as_index=False)[['X_speed', 'Y_speed',
'PlayerWeight',
'X_acceleration','Y_acceleration']].sum()
tot_momentum['x_momentum'] = tot_momentum['X_speed'] * tot_momentum['PlayerWeight']
tot_momentum['y_momentum'] = tot_momentum['Y_speed'] * tot_momentum['PlayerWeight']
tot_momentum['x_force'] = tot_momentum['X_acceleration'] * tot_momentum['PlayerWeight']
tot_momentum['y_force'] = tot_momentum['Y_acceleration'] * tot_momentum['PlayerWeight']
tot_momentum.drop(['X_speed', 'Y_speed',
'PlayerWeight',
'X_acceleration','Y_acceleration'], axis=1, inplace=True)
avg_by_play = pd.merge(avg_by_play, tot_momentum, on=['PlayId', 'Team', 'offense_team'])
avg_by_play = pd.merge(avg_by_play, spread, on=['PlayId', 'Team', 'offense_team'])
poss_team = avg_by_play[avg_by_play.Team == avg_by_play.offense_team].copy()
def_team = avg_by_play[avg_by_play.Team != avg_by_play.offense_team].copy()
poss_team.rename(columns={'PlayerHeight': 'poss_avg_height',
'PlayerWeight': 'poss_avg_weight',
'age': 'poss_avg_age',
'X': 'poss_std_X',
'Y': 'poss_std_Y',
'S': 'poss_avg_S',
'A': 'poss_avg_A',
'x_momentum': 'poss_x_momentum',
'y_momentum': 'poss_y_momentum',
'x_force': 'poss_x_force',
'y_force': 'poss_y_force'}, inplace=True)
def_team.rename(columns={'PlayerHeight': 'def_avg_height',
'PlayerWeight': 'def_avg_weight',
'age': 'def_avg_age',
'X': 'def_std_X',
'Y': 'def_std_Y',
'S': 'def_avg_S',
'A': 'def_avg_A',
'x_momentum': 'def_x_momentum',
'y_momentum': 'def_y_momentum',
'x_force': 'def_x_force',
'y_force': 'def_y_force'}, inplace=True)
avg_by_play = pd.merge(poss_team.drop('Team', axis=1),
def_team.drop('Team', axis=1), on=['PlayId', 'offense_team'])
avg_by_play['tot_x_momenumt'] = avg_by_play['poss_x_momentum'] - avg_by_play['def_x_momentum']
avg_by_play['tot_x_force'] = avg_by_play['poss_x_force'] - avg_by_play['def_x_force']
avg_by_play['height_diff'] = avg_by_play['poss_avg_height'] - avg_by_play['def_avg_height']
avg_by_play['weight_diff'] = avg_by_play['poss_avg_weight'] - avg_by_play['def_avg_weight']
avg_by_play['age_diff'] = avg_by_play['poss_avg_age'] - avg_by_play['def_avg_age']
avg_by_play['X_diff'] = avg_by_play['poss_std_X'] - avg_by_play['def_std_X']
avg_by_play['Y_diff'] = avg_by_play['poss_std_Y'] - avg_by_play['def_std_Y']
return avg_by_play
def process_play(self, X):
cols_by_play = ['GameId', 'PlayId', 'YardLine',
'Quarter', 'GameClock', 'Down', 'Distance',
'OffenseFormation', 'DefendersInTheBox',
'Location', 'StadiumType', 'Turf',
'GameWeather','Temperature', 'Humidity', 'WindSpeed', 'WindDirection',
'PlayDirection', 'HomeScoreBeforePlay', 'VisitorScoreBeforePlay']
train_play = X[cols_by_play].drop_duplicates()
avg_by_play = self.stats_by_play(X)
train_play = pd.merge(train_play, avg_by_play.drop('offense_team', axis=1), on=['PlayId'])
return train_play
def transform(self, X, y=None):
train_play = self.process_play(X)
carriers = X[X.has_ball].copy()
to_drop = ['GameId', 'NflId', 'Team', 'Orientation','YardLine', 'Quarter', 'GameClock', 'PossessionTeam',
'Down', 'FieldPosition', 'HomeScoreBeforePlay',
'VisitorScoreBeforePlay', 'NflIdRusher', 'OffensePersonnel','DefensePersonnel',
'PlayDirection', 'Position', 'HomeTeamAbbr',
'VisitorTeamAbbr', 'Location', 'StadiumType', 'GameWeather',
'Temperature', 'Humidity', 'WindSpeed', 'WindDirection', 'to_left',
'has_ball', 'offense_team', 'Distance',
'OffenseFormation', 'DefendersInTheBox', 'Turf']
carriers.drop(to_drop, axis=1, inplace=True)
full_train = pd.merge(carriers, train_play, on='PlayId')
full_train.drop(['GameId', 'WindDirection', 'WindSpeed', 'GameWeather',
'PlayDirection', 'StadiumType', 'Turf', 'Location', 'GameClock'], axis=1, inplace=True)
self.columns = full_train.columns
return full_train
def get_features_name(self):
return self.columns
# +
tmp = transformation().transform(df_train)
tmp.head()
# -
df_train.head()
# +
transf_pipe = Pipeline([('trsf', transformation()),
('dummifier', dummify(drop_first=True)),
('Imputer', df_imputer()),
('scl', df_scaler(method='standard'))])
tmp = transf_pipe.fit_transform(df_train)
tmp.head()
# -
# # Targets preparation
#
# These are targets that ultimatively were not used but might have led to richer models.
def create_targets(data):
unique_plays = data[['PlayId', 'Yards']].drop_duplicates()
simple = unique_plays['Yards'].reset_index(drop=True)
# As total distance of the rusher
rushers = data[data.has_ball].copy().reset_index(drop=True)
tot_dist = rushers['Yards'] + rushers['from_yardline']
# As percentage of Yards remaining to be gained
unique_plays = data[['PlayId', 'YardLine', 'Yards']].drop_duplicates().reset_index(drop=True)
perc_gained = unique_plays['Yards'] / (110 - unique_plays['YardLine'])
# As both
perc_dist = tot_dist / (110 - rushers['X'])
return simple, tot_dist, perc_gained, perc_dist
y1, y2, y3, y4 = create_targets(df_train)
# +
targets = pd.DataFrame({'simple': y1, 'total_distance': y2, 'perc_gained': y3, 'perc_dist': y4})
targets.head()
# -
targets.describe()
| nfl_2019/notebooks/02 - Transformation Pipeline.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/harnalashok/deeplearning/blob/main/pretrained_layers_autoencoder_II.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="VKSFbYrgfA-y"
# <table align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/harnalashok/deeplearning/blob/main/pretrained_layers_autoencoder_II.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# </table>
# + id="8hUNNtAJhWE7"
# Last amended: 21st Jan, 2021
# Myfolder:
#
# Objectives
# i) Building autoencoder using Model class subclassing
# ii) Training autoencoder with gaussian noise added
# iii) Using pre-trained autoencoder layers in a classifier
# iv) Comparing Classifer performance with and without pre-trained
# v) Using keras model as a layer
# vi) A pre-trained model using autoencoder-with-noise added gives
# better classification
#
#
# Ref: https://www.tensorflow.org/tutorials/generative/autoencoder#first_example_basic_autoencoder
# https://www.tensorflow.org/tutorials/generative/autoencoder#third_example_anomaly_detection
# Practical Recommendations for Gradient-Based Training of DeepArchitectures by <NAME>
#
# + id="WoU-r_hoe4qD"
# 1.0 Import libraries
import tensorflow as tf
from tensorflow.keras import layers
from tensorflow.keras.datasets import fashion_mnist
from tensorflow.keras.models import Model
# + id="lZygQFzuHUo4"
# 1.1 Display outputs from multiple commands in a colab cell
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = "all"
# + colab={"base_uri": "https://localhost:8080/"} id="LR9cqRU3e--E" outputId="2e42bb71-67cc-4c14-fff1-7ee8eb5e655a"
# 2.0 Get fashion mnist data
(x_train, y_train), (x_test, y_test) = fashion_mnist.load_data()
# Normalize data
x_train = x_train.astype('float32') / 255.
x_test = x_test.astype('float32') / 255.
# Data shape
print (x_train.shape)
print (x_test.shape)
# + id="M1WD0i_-Gwfl"
# 2.1 Reshape data for feeding it to NN model
x_train = x_train.reshape((-1, 784))
x_test = x_test.reshape((-1, 784))
# + id="Hgm7Ld0UeAVh"
# 2.2 Design an Autoencoder with Subclassing
# Encoder has noise added
# Ref: https://www.tensorflow.org/guide/keras/custom_layers_and_models
# Page 313, Book: Hands-on Machine Learning witgh Scitkit-Learn, Keras, and Tensorflow
latent_dim = 64
class Autoencoder(Model):
# 2.2.1 Design all layers
def __init__(self, latent_dim, noise_level=0.1):
super(Autoencoder, self).__init__()
self.latent_dim = latent_dim
self.noise_level = noise_level
# 2.2.2 This is our encoder
self.encoder = tf.keras.Sequential(
[
layers.Input(shape=(784,)),
layers.Dense(self.latent_dim, activation='relu'),
layers.Dense(self.latent_dim, activation='relu'),
layers.GaussianNoise(0.1), # Add some noise
layers.Dense(self.latent_dim, activation='relu')
]
)
# 2.2.3 This is our decoder
self.decoder = tf.keras.Sequential(
[
layers.Dense(self.latent_dim, activation='relu'),
layers.Dense(self.latent_dim, activation='relu'),
layers.Dense(784, activation='sigmoid'),
#layers.Reshape((28, 28))
]
)
# 2.2.4 Call function with just one parameter
def call(self, inputs):
encoded = self.encoder(inputs)
decoded = self.decoder(encoded)
return decoded
# + colab={"base_uri": "https://localhost:8080/"} id="AVI1uxltfQzP" outputId="361dacf6-e176-4d98-d1d0-6e4561568795"
# 3.0 Instantiate, compile and train autoencoder
autoencoder = Autoencoder(100, 0.1)
autoencoder.compile(optimizer='adam', loss="mse")
autoencoder.fit(x_train, x_train,
epochs=100,
shuffle=True,
validation_data=(x_test, x_test))
# + id="EfoYTxPNdX1d"
# 3.1 Layer-wise summary
autoencoder.summary()
# + id="VgTclMTsdd1x"
# 3.2 Just look at layers
autoencoder.layers
autoencoder.layers[-2]
# + id="Xu4uMR9IU4vf"
# 4.0 Design an Autoencoder with Subclassing
# BUT Encoder has NO noise added
latent_dim = 64
class Autoencoder_n(Model):
# 4.0.1 Design all layers
def __init__(self, latent_dim, noise_level=0.1):
super(Autoencoder_n, self).__init__()
self.latent_dim = latent_dim
self.noise_level = noise_level
# 4.0.2 This is our encoder
self.encoder = tf.keras.Sequential(
[
layers.Input(shape=(784,)),
layers.Dense(self.latent_dim, activation='relu'),
layers.Dense(self.latent_dim, activation='relu'),
#layers.GaussianNoise(0.1),
layers.Dense(self.latent_dim, activation='relu')
]
)
# 4.0.3 This is our decoder
self.decoder = tf.keras.Sequential(
[
layers.Dense(self.latent_dim, activation='relu'),
layers.Dense(self.latent_dim, activation='relu'),
layers.Dense(784, activation='sigmoid'),
#layers.Reshape((28, 28))
]
)
# 4.0.4 Call function with just one parameter
def call(self, inputs):
encoded = self.encoder(inputs)
decoded = self.decoder(encoded)
return decoded
# + colab={"base_uri": "https://localhost:8080/"} id="ezFcbBWjVQGT" outputId="42719a02-4c73-471b-d6a8-89f8a0ec2d11"
# 5.0 As our model has been built using subclassing API,
# to intantiate the model, we have to fit it.
# Of course, this training is of no use as we will
# replace the encoder weights by the learned weights
# of earlier autoencoder
autoencoder_n = Autoencoder_n(100)
autoencoder_n.compile(optimizer='adam', loss="mse")
autoencoder_n.fit(x_train, x_train,
epochs=10,
shuffle=True,
validation_data=(x_test, x_test))
# + id="tMkLkswaWxY2"
# 5.1 Replace 'encoder' weights of autoencoder_n
autoencoder_n.layers[1].set_weights(autoencoder.layers[1].get_weights())
# + id="C8eSAC0NfQ6i"
# 6.0 So now we have two autoencoders. One which was trained with noise added
# to input. And the other whose 'encoder' has the same weights as of earlier
# autoencoder. BUT this autoencoder does NOT have, so-to-say GaussianNoise layer.
# + [markdown] id="IEGs9nThiPr9"
# # Classification
# Using autoencoder pre-trained weights while performing classification
# + id="o36gyv5nIODO"
# 7.0 Define Classification model function
#
def class_model(trainable = False):
model1 = tf.keras.models.Sequential()
# 7.1 Add autoencoder_n as a layer
# But only the 'encoder' part
# WE ADD THAT autoencoder that
# DOES NOT have gauusian noise layer
model1.add(autoencoder_n.layers[-2])
# 7.2 This is the output layer of our model
model1.add(layers.Dense(10,activation = "softmax"))
# 7.3 No training for autoencoder
autoencoder.layers[-2].trainable = trainable
model1.layers[0].trainable = trainable
return model1
# + colab={"base_uri": "https://localhost:8080/"} id="pJyB92zeJVL2" outputId="d278eb52-760a-4623-bfa3-c681568ad001"
# 8.0 Instantiate classification model and train it
model1 = class_model(False)
# 8.1
model1.compile(loss = "sparse_categorical_crossentropy", metrics = "accuracy")
# 8.2
model1.fit(x_train, y_train,
epochs=100,
shuffle=True,
validation_data=(x_test, y_test)
)
# + colab={"base_uri": "https://localhost:8080/"} id="Q0_gpKiokzp9" outputId="22af28de-fad6-4cd0-a557-0d369faa97ae"
# 8.3 Evaluate model
model1.evaluate(x_test, y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="oi0kq3elIpq9" outputId="6c55cd6f-6ac7-4b00-e2c9-6d3fbc987849"
# 8.4 Also get its summary
model1.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="5mYAfU_zObRU" outputId="eefd07bb-8552-4c1f-de7d-425115979d37"
# 9.0 Run the classification model again but
# this time train the autoencoder layer
model2 = class_model(True)
# 9.1
model2.compile(loss = "sparse_categorical_crossentropy", metrics = "accuracy")
# 9.2
model2.fit(x_train, y_train,
epochs=100,
shuffle=True,
validation_data=(x_test, y_test)
)
# + colab={"base_uri": "https://localhost:8080/"} id="a_NIL4F3OgCq" outputId="f050cadf-e79f-451d-da95-e7b788c9978c"
# 9.3 Evaluate the model
# Observe that a pre-trained model using
# autoencoder gives better classification
model2.evaluate(x_test,y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="vSt7xaHkjduo" outputId="b625bb88-fc84-46<PASSWORD>-ad<PASSWORD>"
# 10.0 If you evaluate model1 again, we get very low accuracy
# as autoencoder weights have changed
model1.evaluate(x_test,y_test)
# + id="eQ91niuYxFM1"
| pretrained_layers_autoencoder_II.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="TWvhl5uAx6lr" colab_type="code" outputId="f97442b0-cc86-4971-babe-b154b846db4e" colab={"base_uri": "https://localhost:8080/", "height": 122}
from google.colab import drive
drive.mount('/content/gdrive')
# + id="Yloj8NxbyJcb" colab_type="code" outputId="b984d905-903b-487b-e884-250914925632" colab={"base_uri": "https://localhost:8080/", "height": 34}
import sys,random,math
from collections import Counter
import numpy as np
f = open('gdrive/My Drive/Grokking/tasksv11/en/qa1_single-supporting-fact_train.txt','r')
raw = f.readlines() #len(raw): 3000; raw[:3] ['1 Mary moved to the bathroom.\n', '2 John went to the hallway.\n', '3 Where is Mary? \tbathroom\t1\n']
f.close()
tokens = list()
for line in raw[0:1000]:
tokens.append(line.lower().replace("\n","").split(" ")[1:])
print(len(tokens),tokens[0:3])
# + id="JSYrGh0eXEa0" colab_type="code" outputId="99f4eca9-e2f4-46c2-ea05-b279e04c5eed" colab={"base_uri": "https://localhost:8080/", "height": 34}
vocab = set()
for sent in tokens:
for word in sent:
vocab.add(word)
vocab = list(vocab) #vocab len: 82
print("sent len:",len(sent),"; sent:",sent)
word2index = {}
for i,word in enumerate(vocab):
word2index[word]=i
def words2indices(sentence):
idx = list()
for word in sentence:
idx.append(word2index[word])
return idx
def softmax(x):
e_x = np.exp(x - np.max(x))
return e_x / e_x.sum(axis=0)
# + id="JOcvo3nyagk9" colab_type="code" colab={}
np.random.seed(1)
embed_size = 10
# word embeddings
embed = (np.random.rand(len(vocab),embed_size) - 0.5) * 0.1 #embed.shape: (82, 10)
# embedding -> embedding (initially the identity matrix)
recurrent = np.eye(embed_size) #recurrent.shape: (10, 10)
# sentence embedding for empty sentence
start = np.zeros(embed_size)
# embedding -> output weights
decoder = (np.random.rand(embed_size, len(vocab)) - 0.5) * 0.1 #decoder.shape: (10, 82)
# one hot lookups (for loss function)
one_hot = np.eye(len(vocab))
# + id="30YXC8M6bsSD" colab_type="code" colab={}
def predict(sent):
layers = list()
layer = {}
layer['hidden'] = start
layers.append(layer)
loss = 0
# forward propagate
preds = list()
for target_i in range(len(sent)):
layer = {}
# try to predict the next term
layer['pred'] = softmax(layers[-1]['hidden'].dot(decoder)) #layer['pred']).shape: (82,)
loss += -np.log(layer['pred'][sent[target_i]])
# generate the next hidden state
layer['hidden'] = layers[-1]['hidden'].dot(recurrent) + embed[sent[target_i]] #layer['hidden'].shape: (10,)
layers.append(layer)
return layers, loss
#a = [10, 5,20, 81]
#predict(a)
# + id="ebmwwJ_ot8hy" colab_type="code" outputId="bd9bfebb-4387-4012-eafb-6fc764bc616c" colab={"base_uri": "https://localhost:8080/", "height": 527}
# forward
for iter in range(30000):
alpha = 0.001
sent = words2indices(tokens[iter%len(tokens)][1:]) # eg tokens ['moved', 'to', 'the', 'bathroom.'] | eg sent [7, 21, 29, 34]
layers,loss = predict(sent)
# back propagate
for layer_idx in reversed(range(len(layers))): # eg len(layers): 5
layer = layers[layer_idx]
target = sent[layer_idx-1]
if(layer_idx > 0): # if not the first layer
layer['output_delta'] = layer['pred'] - one_hot[target]
new_hidden_delta = layer['output_delta'].dot(decoder.transpose())
# if the last layer - don't pull from a later one becasue it doesn't exist
if(layer_idx == len(layers)-1):
layer['hidden_delta'] = new_hidden_delta
else:
layer['hidden_delta'] = new_hidden_delta + layers[layer_idx+1]['hidden_delta'].dot(recurrent.transpose())
else: # if the first layer
layer['hidden_delta'] = layers[layer_idx+1]['hidden_delta'].dot(recurrent.transpose())
# update weights
start -= layers[0]['hidden_delta'] * alpha / float(len(sent))
for layer_idx,layer in enumerate(layers[1:]):
decoder -= np.outer(layers[layer_idx]['hidden'], layer['output_delta']) * alpha / float(len(sent))
embed_idx = sent[layer_idx]
embed[embed_idx] -= layers[layer_idx]['hidden_delta'] * alpha / float(len(sent))
recurrent -= np.outer(layers[layer_idx]['hidden'], layer['hidden_delta']) * alpha / float(len(sent))
if(iter % 1000 == 0):
print("Perplexity:" + str(np.exp(loss/len(sent))))
# + id="d3cYSI61Td_b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="729beaf6-c9d2-4225-ce22-bc94eb444f29"
sent_index = 4
l,_ = predict(words2indices(tokens[sent_index]))
print(tokens[sent_index])
for i,each_layer in enumerate(l[1:-1]):
input = tokens[sent_index][i]
true = tokens[sent_index][i+1]
pred = vocab[each_layer['pred'].argmax()]
print("Prev Input:" + input + (' ' * (12 - len(input))) +\
"True:" + true + (" " * (15 - len(true))) + "Pred:" + pred)
| Recurrence/Predicting_Next_Word_RNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + jupyter={"outputs_hidden": false}
# %matplotlib inline
# -
#
# # Using the SMAC interface tuned for hyperparameter optimization for black-box optimization
#
#
# + jupyter={"outputs_hidden": false}
import logging
import numpy as np
print(np.version.version)
from ConfigSpace.hyperparameters import UniformFloatHyperparameter
# Import ConfigSpace and different types of parameters
from smac.configspace import ConfigurationSpace
from smac.facade.smac_hpo_facade import SMAC4HPO
# Import SMAC-utilities
from smac.scenario.scenario import Scenario
def rosenbrock_2d(x):
""" The 2 dimensional Rosenbrock function as a toy model
The Rosenbrock function is well know in the optimization community and
often serves as a toy problem. It can be defined for arbitrary
dimensions. The minimium is always at x_i = 1 with a function value of
zero. All input parameters are continuous. The search domain for
all x's is the interval [-5, 10].
"""
x1 = x["x0"]
x2 = x["x1"]
val = 100. * (x2 - x1 ** 2.) ** 2. + (1 - x1) ** 2.
return val
logging.basicConfig(level=logging.INFO) # logging.DEBUG for debug output
# Build Configuration Space which defines all parameters and their ranges
cs = ConfigurationSpace()
x0 = UniformFloatHyperparameter("x0", -5, 10, default_value=-3)
x1 = UniformFloatHyperparameter("x1", -5, 10, default_value=-4)
cs.add_hyperparameters([x0, x1])
# Scenario object
scenario = Scenario({"run_obj": "quality", # we optimize quality (alternatively runtime)
"runcount-limit": 10, # max. number of function evaluations; for this example set to a low number
"cs": cs, # configuration space
"deterministic": "true"
})
# Example call of the function
# It returns: Status, Cost, Runtime, Additional Infos
def_value = rosenbrock_2d(cs.get_default_configuration())
print("Default Value: %.2f" % def_value)
# Optimize, using a SMAC-object
print("Optimizing! Depending on your machine, this might take a few minutes.")
smac = SMAC4HPO(scenario=scenario,
rng=np.random.RandomState(42),
tae_runner=rosenbrock_2d)
smac.optimize()
# -
1+1
| examples/SMAC4HPO_rosenbrock.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Rock Paper Scissors Code Along
#
# In this code along, we will build a game or rock, paper, scissors. This is intended for novice programmers who should have finished basic tutorials on python and coding. However, I provided some walk through and solutions for anyone to follow this code along. It is recommended that the users have basic understanding of data types, loops, boolean statements, and function building. If you are not familiar with these topics, it is recommeded to complete the free lesson available on [kaggle.com](https://www.kaggle.com/learn/python).
#
# 
#
# ## Objective
#
# This notebook is to provide a code along for creating rock, paper, scissors game in python. In this project following concepts will be covered. <br>
#
# - Using built-in `input` function to communicate with a user
# - Create logics to validate input
# - Use python's `random` libraries
# - Practice while loops and if statements
# - Organize code by making functions
# - Practice object oriented programming
#
# ## Author
# - [<NAME>](https://github.com/yunghanjeong/rockpaperscissors)
# ## User Input
#
# Prompting a user for an input in python can be done with the built-in function, `input`. Try running the cell below.
# run the cell, provide some input, press enter. The cell will display the input
# the input can also be saved into a variable
user_input = input("type something here: ")
# run this cell to confirm the correct storage of variable
print("your input was: ", user_input)
# As seen above the input function asks the user and ask for an input. You can pass a string in the `input()` function to add a prompt. The input function will also store the value directly and we can chain a method relevant to the data type we are expecting. We can test this out with a string method `upper()` with the input function. Run the cell below for example.
# type something in lower case
user_input = input("type something in lower case here: ").upper()
print("your input was: ", user_input)
# If you run the cell above, but provide a numeric value as an input python will automatically skip the `upper()` method since that is not a method found within `int` for `float` data types. This will be useful later, because processing the input by chaining a method allows us to keep the user input consistent. We don't need to account for varying capitalization.
# ## Checking Against a List
#
# Now that we can ask the user for their input let's start building the logic to validate the user input. Since this is a game of rock, paper, scissors we only have to deal with 3 valid input types. You can build a series of `if/elif` statements to check for the value, but python provides with a fantastic operator called `in`. This will allow us to check a value against a list of values. Run the cell below for a demonstration
print(1 in [1, 2, 3])
print(12 in [11, 13, 14])
print("hello" in ["hello", "world"])
# The `in` operator can also be combined with `not` operator.
print("python" not in ["hello", "world"])
# ## Building Input Validation
#
# Now we are ready to build the logic for user input validation. We want to build a logic that will prompt the user for rock, paper, or scissors. Then, we would want to check the input against a list of valid inputs. If the input is not valid, prompt the user until a valid response is given. Since this should run continuously a `while` loop should be utilized rather than an `if` statement.
# +
# build a list of valid input
# initialize a user input that is a blank strings
# create a while loop which asks for a valid response
# +
# uncomment below to run a provide solution
# choices = ["rock", "paper", "scissors"] #list of valid inputs
# player_input = "" #blank initialization
# while player_input not in choices: #while the proper choice is not provide, this will run at least once by default
# player_input = input("Rock, Paper, or Scissors? ").lower() #ask for input
# -
# ## Saving the Logic
#
# Once the logic building is complete, it's alwaysa a good idea to save the logic in a function. This will allow us to utilize the logic again by calling the fuction. You can define a function with `def` and providing a name followed with necessary arguments (0 or more). You can `return` a value within a function, but it is not mandatory. Run the next two cells for a demonstration on building functions.
# +
# this function will always return the string when called
def no_argument_fx():
return "This function returns a string"
# this function will not return any values but has a print out
def no_return_fx(user_input):
print(f"You provided {user_input} as the function argument")
# +
no_argument = no_argument_fx()
print(no_argument)
print(no_return_fx(1987))
# -
# Also run the cell below to see how a function with no return statement works. Note that you can assign a variable, which can initialize the function, but it will have a data type of NoneType. This is because the function does not return anything.
noreturn = no_return_fx(1234)
type(noreturn) #NoneType, because the function does not return anything!
# This function in particular does not require any arguments, but we will `return` the user input.
#
# **NOTE** Initialize the user input and valid input list as the local variable of the function!
# build a function using the logic built previously
# write the function with no arguments and return the user input
# +
# uncomment below to run a provided solution
# function
# def user_selection():
# player_input = "" #initialize user input
# valid_selection = ["rock", "paper", "scissors"] #initialize valid input list
# while player_input not in valid_selection:
# player_input = input("Rock, Paper, or Scissors? ").lower()
# return player_input
# -
# ***
# ## Random Library and Computer's Selection
#
# Unfortunately, you can't program a computer to select a true random value. However, python has a `random` library that simulates a random selection. To pick a value from a list we can use the `.choice()` method. Run the cell below for example.
# try running this cell multiple times!
import random
print(random.choice([1,2,3]))
# Now we can utilize this method to simulate a computer choosing their rock, paper, or scissors. Try this with a valid input list you've built above.
# +
# using the random.choice() function, make the computer select a value from the valid input list
# -
# ## Comparing the Result
#
# 
#
# There are total 7 outcomes in a game of rock paper scissors. There are 3 ways to win, 3 ways to lose, and 1 way to tie. Technically there are 3 ways to tie, but we can combine all ties into a single comparison logic. There are only limited case of logic check, so this is a perfect candidate for `if` statements. Remember that you can chain if statements with `elif` statements. If you need a refresher, run the cell below.
# +
# run this multiple times to see different results
the_number = 3 # the answer
list_num = [0, 1, 2, 3, 4, 5] # the choices
# to chain multiple if statements, use elif
if the_number == random.choice(list_num):
print("The number is: ", the_number)
elif the_number > random.choice(list_num):
print("The number is greater than our guess")
else:
print("the number is lower than our guess")
# -
# Now let's try building the logic, which compares the computer's choice against the user choice. Remember that we built a function asking for user's choice above and can utilize that. Now this should start to resemble a game!
# +
# call the player input function above and store the input into a variable
# save a computer's random selection into a variable
# build if/elif statements to check for who won the game. Announce the result using print statements.
# +
# uncomment below to run a provided solution.
# Make sure to uncomment and run other solutions above to ensure that this runs properly!
# import random #this is redudant, but repeated for safety
# valid_selection = ["rock", "paper", "scissors"]
# user_input = user_selection()
# computer_selection = random.choice(valid_selection)
# print("Your Choice: ", user_input.title())
# print("Computer's Choice: ", computer_selection.title())
# if computer_selection == user_input: #draw condition
# print("It's a draw!")
# elif computer_selection == "rock" and user_input == "paper":
# print("You win!")
# elif computer_selection == "rock" and user_input == "scissors":
# print("Computer wins!")
# elif computer_selection == "paper" and user_input == "scissors":
# print("You win!")
# elif computer_selection == "paper" and user_input == "rock":
# print("Computer wins!")
# elif computer_selection == "scissors" and user_input == "paper":
# print("Computer wins!")
# elif computer_selection == "scissors" and user_input == "rock":
# print("You win!")
# -
# ## Saving the Logic (Again!)
#
# As it was done before, let's save this logic as a function. This function will have 1 argument, which is the player choice. The function will make the computer choose from a valid input, should be another local variable, and print out the result. This function does not need to return a value.
# +
# define the function name with 1 argument (player choice)
# the function should make the computer pick a random choice
# then compare the computer's choice and player's choice and announce the result
# +
# uncomment below to run a provided solution.
# import random #could be redudant
# def who_won(player):
# valid_selection = ["rock", "paper", "scissors"]
# computer_selection = random.choice(valid_selection)
# print("Your Choice: ", user_input.title())
# print("Computer's Choice: ", computer_selection.title())
# if computer_selection == user_input: #draw condition
# print("It's a draw!")
# elif computer_selection == "rock" and user_input == "paper":
# print("You win!")
# elif computer_selection == "rock" and user_input == "scissors":
# print("Computer wins!")
# elif computer_selection == "paper" and user_input == "scissors":
# print("You win!")
# elif computer_selection == "paper" and user_input == "rock":
# print("Computer wins!")
# elif computer_selection == "scissors" and user_input == "paper":
# print("Computer wins!")
# elif computer_selection == "scissors" and user_input == "rock":
# print("You win!")
# -
# ## Build the Game
#
# Now we are ready to build the game! Before we build the game, however, ensure that the cells containing your functions ran, so we have access to them. Building the game is as simple as running the first function and saving the user input and passing the user input to the result comparison function.
# +
# create a variable and store the user input function return value
# call the result comparison function and pass the variable above to run
# +
# Example implementation, ensure to run all cells with provided solution to run this cell.
# user_input = user_selection()
# who_won(user_input)
# -
# ## Refactoring
#
# Congratulations on finishing the code along! However, the project is far from over, it is always recommended for programmers to refactor their code to reduce runtime and redudancy and add additional features. In this section I would like to recommend following steps for refacotring. When you are done please check out the bonus section below.
#
# - Call the user function within result comparison to reduce function call
# - Add a function that tracks the score
# - Format the print statements for better readability
# - Create a function that asks player if they want to play again and loop the game accordingly
# ## BONUS: Object Oriented Programming
#
# As you were following along this notebook and programming this, I hope you were able to spot some redudancies in the coding. You can utilize object oriented programming to overcome some of the redudancies and organize the project by separating your functions and the actual game file. Try combining all the functions under a single class and initialize all local variables under self. Then, import the game class into a new folder and build your game using the functions under the class. You'll be amazing how clean the final implementation would be.
#
# You can check the implementation below for the example and reference the `src` folder for the class that I have built.
#
# Good luck!
#
# ### Reference Implementation
#
# Below is a reference for object oriented programming practice of this code. The `rockpaperscissors.py` located in `src` folder contains `game` class, which contains all the functions that have been called. Please check out the `src` folder in the repo for the code reference. You can also check out `rock_paper_scissors_game.py` for a complete python implementation.
# +
import os #this might be necessary for some systems
from src.rockpaperscissors import game
play = True #set true to run first
gm = game() #initialize the game
while play:
user_score, computer_score = gm.who_won() # play and get score
print(f"Current Score is Player: {user_score} Computer: {computer_score}") #print score
play = gm.play_again() #check for replay
# -
| rock_paper_scissors_code_along.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Python Introduction with some detours
# 
# 
# # Table of Contents <a class="anchor" id="toc">
#
# * [Hello World](#hello-world)
# * [Python History](#python-history)
# * [What is Programming?](#what-is-programming)
# * [Jupyter Basics](#jupyter-basics)
# * [Variables](#variables)
# * [Arithmetic](#arithmetic)
# * [Program Flow Control](#flow-control)
# * [Functions](#functions)
# * [Libraries](#libraries)
# * [Crucial Python Ideas](#python-ideas)
# * [Learning Resources](#learning-resources)
# ## Hello World <a class="anchor" id="hello-world">
#
# [Back to Table of Contents](#toc)
print("Hello world!")
# +
### Try printing a greeting of your own!
# +
### What Happens when you get an error?
# -
# ## Python History <a class="anchor" id="python-history">
#
# [Back to Table of Contents](#toc)
#
# ## Python created by <NAME> in early 1990s (later at Google,Dropbox)
# ## Language reference: https://docs.python.org/3/index.html
#
# 
# # Why Python?
# ## Why Python but not Julia or R or some other language or even VBA in Excel?
# ### Readability, Glue Language(APIs), From Startups to Google and Facebook, Pipeline
#
# Prime numbers in J language used at some finance institutions **1: I. 2= +/ 0= (] |/ ]) i.y**
# 
# ### Python is now programming language # 3 in TIOBE language index (as of September 2018)
# https://www.tiobe.com/tiobe-index/
#
# https://developers.slashdot.org/story/18/09/08/1722213/python-displaces-c-in-tiobe-index-top-3
# 
# https://www.anaconda.com/download/
# ## Batteries included principle
# 
# ## What is Programming? <a class="anchor" id="what-is-programming">
#
# [Back to Table of Contents](#toc)
#
# * Egg algorithm
# * Computers are stupid, they only do what they are told to do
# * If it is stupid but it works, then it is not stupid
# * Make it work, make it right, make it fast(last two steps often not required in real life)
# * GIGO principle
#
# * Error messages are nothing to be afraid of, usually the message will explain what needs fixing!
# +
# Our first comment
# Real Program Comments should generally describe why
# Here comments will describe extra information not covered or needed for starting out
# REPL(Read,Eval,Print, Loop)
# Python - Interpreted Language(commands executed as they come)
# -
# ## Jupyter Basics <a class="anchor" id="jupyter-basics">
#
# [Back to Table of Contents](#toc)
#
#
# * Esc-M turns cell into Markdown cell for formatting (https://guides.github.com/pdfs/markdown-cheatsheet-online.pdf)
# * Esc-Y turns cell into code cell(default)
#
#
# * Ctrl-Enter runs code of cell in place
# * Alt-Enter runs code for current cell and creates a new cell below
# * Esc-A creates a new cell above current cell
# * Esc-B creates a new cell below current cell
# * Esc-dd deletes current cell
# +
# Try Esc-B to create a new cell,
# Enter print("Hello RCS!")
# Press Ctrl-Enter
# Did you get any error messages?
# +
# Try Esc-B then Esc-M to creata Markdown cell and write some text
# -
# # Text (Markdown) cell
#
# [Markdown Basics](https://guides.github.com/features/mastering-markdown/)
#
# I want to write a *novel*
# ## 2nd Level Headline
#
# * Unordered **List** item 1
# * List Item 2
#
#
# 1. Item 1
# 2. Item 2
#
#
# # Esc - Y makes cell a code cell
# # Esc - M makes cell a markdown cell
# ## Variables <a class="anchor" id="variables">
#
# [Back to Table of Contents](#toc)
myname="Valdis"
# Creating our first variable will persist through this workbook once it is run
print(myname)
y = 2019
y
theAnswer = 42
myPi = 3.14159
isHot = True
isHot
# type(variableName) will return variable data type
type(theAnswer)
# +
# What is the data type of myName ?
# How about data type of isHot ?
# -
theAnswer = "My answer"
type(theAnswer)
# Variables cannot be reserved keywords
help("keywords")
# +
# Reserved Keywords
False class from or
None continue global pass
True def if raise
and del import return
as elif in try
assert else is while
async except lambda with
await finally nonlocal yield
break for not
# -
#
# ### Data types in Python 3.x
#
# * Integers type(42) int
# * Floating Point type(3.14) float
# * Boolean type(True),type(False) bool
# * String(ordered, immutable char sequence) type("OyCaramba") str
# * List type([1,2,63,"aha","youcanmixtypeinsidelist", ["even","nest"]]) list
# * Dictionary(key:value pairs) type({"foo":"bar", "favoriteday":"Friday"}) dict
# * Tuple - ordered immutable sequence type("sup",7,"dwarves") tup
# * Set (unordered collection of uniques) ("k","a","r","t","u","p","e","l","i","s")
# ## More on variables
# https://realpython.com/python-variables
# ## Strings
# * immutable
# * Unicode support
#
#
# * implement all common sequence operators
# https://docs.python.org/3/library/stdtypes.html#typesseq-common
#
# * string specific methods
# https://docs.python.org/3/library/stdtypes.html#string-methods
name = "Valdis"
print(name)
# String length
len(name)
# How is this different from name ?
len("name")
# Getting Individual characters
name[0]
# Getting last char
name[-1]
name[-2]
name
name[3]
# ### String Slicing
# Slicing syntax
# Start at 0 an go until but not including 3
name[0:3]
name[:3]
name[1:3]
name
name[0:6:2]
name[::2]
shortName = name[::2]
shortName
# lets play with food!
food = "potatoes"
food[::2]
food[1::2]
food[1:6:2]
# Pythonic way of reversing a string
food[::-1]
# modifying strings
# unmutability
# food[2]="x" is not allowed
newfood = food[:2] + "x" + food[3:]
newfood
# '', "" work the same
# ''' For multiline '''
longstring = ''' This will be a very long string
and a very long day
and a very long summer
'''
# print out the longstring
print(longstring)
# ## "f-strings", “formatted string literals”
#
# In some other languages also known as string interpolation
# Create myname and favfood variables with appropriate text
# Then run the cell below
# What would happen if you did not assign variables ?
favfood = "potatoes"
print(f"My name is {myname} and my favorite food is {favfood} ")
# f strings in Python 3.6+ older formatting methods not covered in this course
# https://realpython.com/python-f-strings/
# Old string concatation method
print("My name is " + myname + " and my favorite food is " + favfood)
# ## Python Lists
#
# * Ordered
# * Mutable(can change individual members!)
# * Comma separated between brackets [1,3,2,5,6,2]
# * Can have duplicates
# * Can be nested
#
newlist = [1,2,3,"Liftoff!"]
newlist
mylist = list(range(11,21+1,1))
# mylist = [11, 12, 13, 14, 15, 16, 17, 18, 19, 20] would work too but not practical for longer ranges...
print(mylist)
# ### Slice notation
#
# somestring[start:end:step]
#
# somelist[start:end:step]
#
# start is at index 0(first element), end is -1 the actual index
# #### Examples below
mylist[0]
mylist[3:]
mylist[-2:]
mylist[:-2]
mylist[::2]
"Valdis"[2:5]
myname[-1]
myname[::-1]
mylist
#
# ### Common list methods.
# * list.append(elem) -- adds a single element to the end of the list. Common error: does not return the new list, just modifies the original.
# * list.insert(index, elem) -- inserts the element at the given index, shifting elements to the right.
# * list.extend(list2) adds the elements in list2 to the end of the list. Using + or += on a list is similar to using extend().
# * list.index(elem) -- searches for the given element from the start of the list and returns its index. Throws a ValueError if the element does not appear (use "in" to check without a ValueError).
# * list.remove(elem) -- searches for the first instance of the given element and removes it (throws ValueError if not present)
# * list.sort() -- sorts the list in place (does not return it). (The sorted() function shown later is preferred.)
# * list.reverse() -- reverses the list in place (does not return it)
# * list.pop(index)-- removes and returns the element at the given index. Returns the rightmost element if index is omitted (roughly the opposite of append()).
mylist.append(42)
mylist
mylist.pop()
mylist
mylist.count(14)
mystr = "this is a string"
mystr.upper()
help(mystr.upper)
words = mystr.split()
words
words[-1]
# ## Dictionaries
#
# * Collection of Key - Value pairs
# * also known as associative array
# * unordered
# * keys unique in one dictionary
# * storing, extracting
mydict = {"country":"Latvia"} #Key-Value store, also knows as Hashmaps, Keys must be unique
mydict["food"]="potatoes"
mydict["food"]
mydict["country"]
len(mydict)
mydict
mydict.keys()
mydict.values()
mydict.values()
"potatoes" in mydict.values()
for key,value in mydict.items():
print(key,value)
mydict['food'] = ['potatoes', 'cheese']
mydict
mydict['food'] = mydict['food'] + ['milk']
mydict
# ## Sets
#
# * unordered
# * uniques only
# * curly braces {3, 6, 7}
s={3,3,6,1,3,6,7}
print(s)
# ## Tuples
#
# * ordered
# * immutable (cannot be changed!)
# * Can be used as a collection of fields
mytuple = 6, 4, 9
print(mytuple)
# ## Arithmetic Operators <a class="anchor" id="arithmetic">
#
# [Back to Table of Contents](#toc)
# * `+ - * / `
# * `**(power)`
# * `% modulus`
# * `//(integer division)`
# * `() parenthesis for order`
#
5+4*3-(6/2)
5+4*3-(6//2)
7//2
int(9.0//2)
int(9.0)//2
4%3
15%2
2554545%10
type(1)
type(14.0)
5**33
11**120 # no maximum anymore
import math
# Tab on math. to see what functions are available
# Shift-Tab inside parenthesis to see what the particular function does
math.ceil(7.8)
math.sqrt(2)
2**0.5
int(3.33)
# ## Flow Control <a class="anchor" id="flow-control">
#
# [Back to Table of Contents](#toc)
# +
# With Flow Control we can tell our program/cells to choose different paths
# -
# ## Conditional operators
#
# `< > <= >= == != and or not`
# +
# What is truth in computer language?
# -
2*2 == 4
myTruth = 2*2 == 4
myTruth
5 > 7
int(' 055555 ')
# +
print(5 == int('5'))
print(5 <= 6)
# +
print(5 <= 5)
# check if 5 is NOT equal to 6
print(5 != 6)
print(5 != 5)
# +
# We check each letter from left side
# on mismatch we check ASCII (UTF-8) tables for values
'VALDIS' < 'VOLDEMARS'
# -
5 < 6
2*2
555+5
myname = "Valdis"
print(myname)
myname
a = True
print(a)
a
len('VALDIS') < len('VOLDEMARS')
True and True
True and False
True or False
False or False or False or True
not True
not False
# ## If Statement
## Conditional execution
# if 4 is larger than 5 then do something
if 4 > 5:
print("4 is larger than 5 wow!")
print("MOre text")
print("Always")
if 5 >= 5:
print("hello")
if 5 <= 5:
print("hello")
if 5 == 6:
print("hello thats magic")
if 5 != 6:
print("hello thats not magic")
if 2*2 == 4:
print("Do one thing if if is True")
print("DO more things if if is True")
print("Do this always")
c = float(input("Enter temperature in Celsius "))
f = c * 9/5 + 32
print("Farenheit Temperature is", f)
if f > 100:
print("You are too hot, find a doctor?")
# +
# Try reversing the above program to create a Farenheit to Celsius converter
# -
a = -55
if a > 5:
print('a is larger than 5')
# do more stuff
else:
print('a is NOT larger than 5')
# do more stuff if a is not larger than 5
print("Continue as normal")
#elif comes from else if
x = int(input("Enter an integer please! "))
if x > 42:
print("Too ambitious an answer!")
elif x < 42:
print("You dream too little!")
else:
print("That is the answer to everything!")
#These lines below will execute always
print('Your number is', x)
# ## Loops
# +
# How would be perform the same/similar action multiple times?
# -
i = 0
print("<NAME> ")
while i < 5: # notice the colon
print("talk")
i+= 1 # same as i = i + 1
i
# +
# What would happen if we did not have i+=1 in our above program ?
# -
for x in range(10):
print(x)
for c in "Valdis":
print(c)
mylist
for item in mylist[:5]:
print(item)
# I could more stuff here
print('This will happen at the end always')
for k,v in mydict.items():
print(k,v)
## Splitting a line of text into words
mytext = "A quick brown fox jumped over a sleeping dog"
words = mytext.split()
print(words)
## Print first letter of each word
for w in words:
print(w[0], w[0].isupper(), w.istitle()) ## istitle() checks every word in a string so not good for unsplit strings
myline="Mr. <NAME>, who was usually very late in the mornings"
words=myline.split()
words
words[1][0].isupper()
# Enumerate for showing index of item when going through many items
for i, x in enumerate(range(10,15)):
print(i, x)
for i, c in enumerate(myname):
print(i, c)
# ## What is a function? <a class="anchor" id="functions">
#
# [Back to Table of Contents](#toc)
# * A function is a block of organized, reusable code that is used to perform a single, related action.
# * Single, organized, related always ? :)
# ### DRY - Do not Repeat Yourself principle
# * Every piece of knowledge must have a single, unambiguous, authoritative representation within a system. http://wiki.c2.com/?DontRepeatYourself
#
# * Contrast WET - We Enjoy Typing, Write Everything Twice, Waste Everyone's Time
# Here we define our first function
def myFirstFunc():
print("Running My first func")
# function has to be defined before it is called
myFirstFunc()
# Passing parameters(arguments)
def printName(name):
print(f"Maybe my name is: {name}")
printName("Valdis")
def add(a, b):
print(a+b)
add(4,6)
add(9,233)
add("Hello ","Riga")
add([1,2,7],list(range(6,12)))
# Try calling add function with other parameters
# We make Docstrings with '''Helpful function description inside'''
def mult(a, b):
'''Returns
multiple from first two arguments'''
print("Look ma I am multiplying!", a, b, a*b)
return a*b
res = mult(4,5)
res
def printnum(num):
if num > 10:
print(f"This number {num} is too unwieldy for me to print")
else:
print(f"This {num} is a nice number")
# +
def isEven(num):
if num%2 == 0:
print(f"{num} is even")
else:
print(f"{num} is odd")
isEven(3)
isEven(4)
# -
def processLine(line):
words = line.split()
linegood=False
for word in words:
if word[0].isupper():
print(word, end='\t')
linegood=True
if linegood == True:
print('')
# ## Libraries <a class="anchor" id="libraries">
#
# [Back to Table of Contents](#toc)
# +
# Python and Batteries Included Philosophy
## Why reinvent the wheel?
# -
# Try importing this
import this
import math
# notice the . syntax helper
math.cos(3.14)
math.
from collections import Counter
magic = "abracadabra"
cnt = Counter(magic)
cnt
# There are hundreds of useful Python libraries
## Crucial libraries are collected in Standard Library
# https://docs.python.org/3/library/
# Batteries included
dir(cnt)
cnt.most_common()
# ### Installing libraries
#
# [Back to Table of Contents](#toc)
#
# Some "batteries" you have to add as you go:
# * there is a huge repository of Python libraries at https://pypi.org/ (and more on Github and other sources)
# * Anaconda already has many of them pre-installed
#
# To install Python libraries from command line, use the `pip` tool:
# * example: `pip install requests`
#
#
# #### "Requests III: HTTP for Humans and Machines, alike"
#
# https://3.python-requests.org/
import requests
# +
# let's get "raw" wiki code from Latvian wikipedia
url = "https://lv.wikipedia.org/w/index.php?title=Rīga&action=raw"
response = requests.get(url)
response.status_code
# +
data = response.text
data[:300]
# -
data[5892:6200]
# ## Most important Python ideas <a class="anchor" id="python-ideas">
#
# [Back to Table of Contents](#toc)
#
# * dir(myobject) to find what can be done (most decent text editors/IDEs will offer autocompletion and hints though)
# * help(myobject) general help
# * type(myobject) what type it is
#
# ## Slicing Syntax for sequences(strings,lists and more)
# `
# myname[start:end:step]
# myname[:5]`
#
# ## : indicates a new indentation level
#
# `if x > 5:
# print("Do Work when x > 5")
# print("Always Do this")`
# # Python Resources <a class="anchor" id="learning-resources">
#
# [Back to Table of Contents](#toc)
# ## Wiki for Tutorials
#
# https://wiki.python.org/moin/BeginnersGuide/NonProgrammers
# ## Tutorials Begginner to Intermediate
#
#
#
#
# * https://automatetheboringstuff.com/ - Anything by <NAME> is great
# * http://newcoder.io/tutorials/ - 5 sets of practical tutorials
# * [Think Like a Computer Scientist](http://interactivepython.org/runestone/static/thinkcspy/index.html) full tutorial
# * [Non-Programmers Tutorial for Python 3](https://en.wikibooks.org/wiki/Non-Programmer%27s_Tutorial_for_Python_3) quite good for wikibooks
# * [Real Python](https://realpython.com/) Python Tutorials for all levels
#
#
# * [Learn Python 3 the Hard Way](https://learnpythonthehardway.org/python3/intro.html) controversial author but very exhaustive, some like this approach
# ## More Advanced Python Specific Books
#
# * [Python Cookbook](https://www.amazon.com/Python-Cookbook-Third-David-Beazley/dp/1449340377) Recipes for specific situations
#
# * [Effective Python](https://effectivepython.com/) best practices
# * [Fluent Python](http://shop.oreilly.com/product/0636920032519.do) **highly recommended**, shows Python's advantages
# ## General Best Practices Books
# #### (not Python specific)
#
# * [Code Complete 2](https://www.goodreads.com/book/show/4845.Code_Complete) - Fantastic best practices
# * [The Mythical Man-Month](https://en.wikipedia.org/wiki/The_Mythical_Man-Month) - No silver bullet even after 40 years.
# * [The Pragmatic Programmer](https://www.amazon.com/Pragmatic-Programmer-Journeyman-Master/dp/020161622X) - More practical advice
# * [Clean Code](https://www.amazon.com/Clean-Code-Handbook-Software-Craftsmanship/dp/0132350882) - more towards agile
# ## Blogs / Personalities / forums
#
# * [<NAME>](https://dbader.org/)
# * [Reddit Python](https://www.reddit.com/r/python)
# ## Exercises/Challenges
# * http://www.pythonchallenge.com/ - first one is easy but after that...
# * [Advent of Code](https://adventofcode.com/) - yearly programming challenges
# * https://projecteuler.net/ - gets very mathematical but first problems are great for testing
# ## Explore Public Notebooks on Github
# Download them and try them out for yourself
#
# https://github.com/jupyter/jupyter/wiki/A-gallery-of-interesting-Jupyter-Notebooks
# ## Questions / Suggestions ?
#
# Pull requests welcome
#
# e-mail **<EMAIL> at gmail.com**
| Python Introduction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ur8xi4C7S06n"
# Copyright 2021 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="JAPoU8Sm5E6e"
# <table align="left">
#
# <td>
# <a href="https://colab.research.google.com/github/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/unofficial/pipelines/pipelines_intro_kfp.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/colab-logo-32px.png" alt="Colab logo"> Run in Colab
# </a>
# </td>
# <td>
# <a href="https://github.com/GoogleCloudPlatform/ai-platform-samples/blob/master/ai-platform-unified/notebooks/unofficial/pipelines/pipelines_intro_kfp.ipynb">
# <img src="https://cloud.google.com/ml-engine/images/github-logo-32px.png" alt="GitHub logo">
# View on GitHub
# </a>
# </td>
# <td>
# <a href="https://console.cloud.google.com/ai/platform/notebooks/deploy-notebook?download_url=https://github.com/GoogleCloudPlatform/ai-platform-samples/raw/master/ai-platform-unified/notebooks/unofficial/pipelines/pipelines_intro_kfp.ipynb">
# Open in Google Cloud Notebooks
# </a>
# </td>
# </table>
# + [markdown] id="o2rM_9Ml7-W2"
# # Introduction to Vertex Pipelines using the KFP SDK
# + [markdown] id="tvgnzT1CKxrO"
# ## Overview
#
# This notebook provides an introduction to using [Vertex Pipelines](https://cloud.google.com/vertex-ai/docs/pipelines) with [the Kubeflow Pipelines (KFP) SDK](https://www.kubeflow.org/docs/components/pipelines/).
#
# ### Objective
#
# In this example, you'll learn:
#
# - The basics of defining and compiling a pipeline.
# - How to schedule recurring pipeline runs.
# - How to specify which service account to use for a pipeline run.
#
#
# ### Costs
#
# This tutorial uses billable components of Google Cloud:
#
# * Vertex AI Training
# * Cloud Storage
# * Cloud Functions
# * Cloud Scheduler
#
#
# Learn about pricing for [Vertex AI](https://cloud.google.com/ai-platform-unified/pricing), [Cloud Storage](https://cloud.google.com/storage/pricing), [Cloud Functions](https://cloud.google.com/functions/pricing), and [Cloud Scheduler](https://cloud.google.com/scheduler/pricing), and use the [Pricing
# Calculator](https://cloud.google.com/products/calculator/)
# to generate a cost estimate based on your projected usage.
# + [markdown] id="ze4-nDLfK4pw"
# ### Set up your local development environment
#
# **If you are using Colab or Google Cloud Notebooks**, your environment already meets
# all the requirements to run this notebook. You can skip this step.
# + [markdown] id="gCuSR8GkAgzl"
# **Otherwise**, make sure your environment meets this notebook's requirements.
# You need the following:
#
# * The Google Cloud SDK
# * Git
# * Python 3
# * virtualenv
# * Jupyter notebook running in a virtual environment with Python 3
#
# The Google Cloud guide to [Setting up a Python development
# environment](https://cloud.google.com/python/setup) and the [Jupyter
# installation guide](https://jupyter.org/install) provide detailed instructions
# for meeting these requirements. The following steps provide a condensed set of
# instructions:
#
# 1. [Install and initialize the Cloud SDK.](https://cloud.google.com/sdk/docs/)
#
# 1. [Install Python 3.](https://cloud.google.com/python/setup#installing_python)
#
# 1. [Install
# virtualenv](https://cloud.google.com/python/setup#installing_and_using_virtualenv)
# and create a virtual environment that uses Python 3. Activate the virtual environment.
#
# 1. To install Jupyter, run `pip install jupyter` on the
# command-line in a terminal shell.
#
# 1. To launch Jupyter, run `jupyter notebook` on the command-line in a terminal shell.
#
# 1. Open this notebook in the Jupyter Notebook Dashboard.
# + [markdown] id="i7EUnXsZhAGF"
# ### Install additional packages
#
# Install the KFP SDK.
# + id="IaYsrh0Tc17L"
import sys
if "google.colab" in sys.modules:
USER_FLAG = ""
else:
USER_FLAG = "--user"
# + id="aR7LNYMUCVKc"
# !python3 -m pip install {USER_FLAG} kfp --upgrade
# + [markdown] id="hhq5zEbGg0XX"
# ### Restart the kernel
#
# After you install the additional packages, you need to restart the notebook kernel so it can find the packages.
# + id="EzrelQZ22IZj"
# Automatically restart kernel after installs
import os
if not os.getenv("IS_TESTING"):
# Automatically restart kernel after installs
import IPython
app = IPython.Application.instance()
app.kernel.do_shutdown(True)
# + [markdown] id="6GPgNN7eeX1l"
# Check the version of the package you installed. The KFP SDK version should be >=1.6.
# + id="NN0mULkEeb84"
# !python3 -c "import kfp; print('KFP SDK version: {}'.format(kfp.__version__))"
# + [markdown] id="lWEdiXsJg0XY"
# ## Before you begin
#
# This notebook does not require a GPU runtime.
# + [markdown] id="BF1j6f9HApxa"
# ### Set up your Google Cloud project
#
# **The following steps are required, regardless of your notebook environment.**
#
# 1. [Select or create a Google Cloud project](https://console.cloud.google.com/cloud-resource-manager). When you first create an account, you get a $300 free credit towards your compute/storage costs.
#
# 1. [Make sure that billing is enabled for your project](https://cloud.google.com/billing/docs/how-to/modify-project).
#
# 1. [Enable the AI Platform (Unified), Cloud Storage, and Compute Engine APIs](https://console.cloud.google.com/flows/enableapi?apiid=aiplatform.googleapis.com,compute_component,storage-component.googleapis.com).
#
# 1. Follow the "**Configuring your project**" instructions from the AI Platform Pipelines documentation.
#
# 1. If you are running this notebook locally, you will need to install the [Cloud SDK](https://cloud.google.com/sdk).
#
# 1. Enter your project ID in the cell below. Then run the cell to make sure the
# Cloud SDK uses the right project for all the commands in this notebook.
#
# **Note**: Jupyter runs lines prefixed with `!` as shell commands, and it interpolates Python variables prefixed with `$` into these commands.
# + [markdown] id="WReHDGG5g0XY"
# #### Set your project ID
#
# **If you don't know your project ID**, you may be able to get your project ID using `gcloud`.
# + id="oM1iC_MfAts1"
import os
PROJECT_ID = ""
# Get your Google Cloud project ID from gcloud
if not os.getenv("IS_TESTING"):
# shell_output=!gcloud config list --format 'value(core.project)' 2>/dev/null
PROJECT_ID = shell_output[0]
print("Project ID: ", PROJECT_ID)
# + [markdown] id="qJYoRfYng0XZ"
# Otherwise, set your project ID here.
# + id="riG_qUokg0XZ"
if PROJECT_ID == "" or PROJECT_ID is None:
PROJECT_ID = "python-docs-samples-tests" # @param {type:"string"}
# + [markdown] id="06571eb4063b"
# #### Timestamp
#
# If you are in a live tutorial session, you might be using a shared test account or project. To avoid name collisions between users on resources created, you create a timestamp for each instance session, and append it onto the name of resources you create in this tutorial.
# + id="697568e92bd6"
from datetime import datetime
TIMESTAMP = datetime.now().strftime("%Y%m%d%H%M%S")
# + [markdown] id="dr--iN2kAylZ"
# ### Authenticate your Google Cloud account
#
# **If you are using AI Platform Notebooks**, your environment is already
# authenticated. Skip this step.
# + [markdown] id="sBCra4QMA2wR"
# **If you are using Colab**, run the cell below and follow the instructions
# when prompted to authenticate your account via oAuth.
#
# **Otherwise**, follow these steps:
#
# 1. In the Cloud Console, go to the [**Create service account key**
# page](https://console.cloud.google.com/apis/credentials/serviceaccountkey).
#
# 2. Click **Create service account**.
#
# 3. In the **Service account name** field, enter a name, and
# click **Create**.
#
# 4. In the **Grant this service account access to project** section, click the **Role** drop-down list. Type "AI Platform"
# into the filter box, and select
# **AI Platform Administrator**. Type "Storage Object Admin" into the filter box, and select **Storage Object Admin**.
#
# 5. Click *Create*. A JSON file that contains your key downloads to your
# local environment.
#
# 6. Enter the path to your service account key as the
# `GOOGLE_APPLICATION_CREDENTIALS` variable in the cell below and run the cell.
# + id="PyQmSRbKA8r-"
import os
import sys
# If you are running this notebook in Colab, run this cell and follow the
# instructions to authenticate your GCP account. This provides access to your
# Cloud Storage bucket and lets you submit training jobs and prediction
# requests.
# If on AI Platform, then don't execute this code
if not os.path.exists("/opt/deeplearning/metadata/env_version"):
if "google.colab" in sys.modules:
from google.colab import auth as google_auth
google_auth.authenticate_user()
# If you are running this notebook locally, replace the string below with the
# path to your service account key and run this cell to authenticate your GCP
# account.
elif not os.getenv("IS_TESTING"):
# %env GOOGLE_APPLICATION_CREDENTIALS ''
# + [markdown] id="NxhCPW6e46EF"
# ### Create a Cloud Storage bucket as necessary
#
# You will need a Cloud Storage bucket for this example. If you don't have one that you want to use, you can make one now.
#
#
# Set the name of your Cloud Storage bucket below. It must be unique across all
# Cloud Storage buckets.
#
# You may also change the `REGION` variable, which is used for operations
# throughout the rest of this notebook. Make sure to [choose a region where AI Platform (Unified) services are
# available](https://cloud.google.com/ai-platform-unified/docs/general/locations#available_regions). You may
# not use a Multi-Regional Storage bucket for training with AI Platform.
# + id="MzGDU7TWdts_"
BUCKET_NAME = "gs://[your-bucket-name]" # @param {type:"string"}
REGION = "us-central1" # @param {type:"string"}
# + id="cf221059d072"
if BUCKET_NAME == "" or BUCKET_NAME is None or BUCKET_NAME == "gs://[your-bucket-name]":
BUCKET_NAME = "gs://" + PROJECT_ID + "aip-" + TIMESTAMP
# + [markdown] id="-EcIXiGsCePi"
# **Only if your bucket doesn't already exist**: Run the following cell to create your Cloud Storage bucket.
# + id="NIq7R4HZCfIc"
# ! gsutil mb -l $REGION $BUCKET_NAME
# + [markdown] id="ucvCsknMCims"
# Finally, validate access to your Cloud Storage bucket by examining its contents:
# + id="vhOb7YnwClBb"
# ! gsutil ls -al $BUCKET_NAME
# + [markdown] id="XoEqT2Y4DJmf"
# ### Import libraries and define constants
# + [markdown] id="YYtGjGG45ELJ"
# Define some constants.
# + id="5zmD19ryCre7"
# PATH=%env PATH
# %env PATH={PATH}:/home/jupyter/.local/bin
USER = "your-user-name" # <---CHANGE THIS
PIPELINE_ROOT = "{}/pipeline_root/{}".format(BUCKET_NAME, USER)
PIPELINE_ROOT
# + [markdown] id="IprQaSI25oSk"
# Do some imports:
#
# + id="UFDUBveR5UfJ"
from typing import NamedTuple
from kfp import dsl
from kfp.v2 import compiler
from kfp.v2.dsl import component
from kfp.v2.google.client import AIPlatformClient
# + [markdown] id="IbN_49SUW7b7"
# ## Define a simple pipeline
#
# This example defines a simple pipeline with three steps.
# The following sections define some pipeline *components*, and then defines a pipeline that uses them.
#
# + [markdown] id="nbV9dECIePV8"
# ### Create Python-function-based pipeline components
#
# First, create a component based on a very simple Python function. It takes a string input parameter and returns that value as output.
#
# Note the use of the `@component` decorator, which compiles the function to a component when evaluated. For example purposes, this example specifies a base image to use for the component (`python:3.9`), and a component YAML file, `hw.yaml` to which to write the compiled component. (The default base image is `python:3.7`, which would of course work just fine too).
#
# After you run the cell below, you can view the generated component yaml file in your local directory.
# + id="Ye7mr8WPW7b8"
@component(output_component_file="hw.yaml", base_image="python:3.9")
def hello_world(text: str) -> str:
print(text)
return text
# + [markdown] id="2dTdxgSm17Rk"
# As you'll see below, evaluation of this component creates a task factory function (called `hello_world`) that you can use in defining a pipeline step.
#
# While we don't show it here, if you want to share this component definition, or use it in another context, you could also load it from its yaml file like this:
# `hello_world_op = components.load_component_from_file('./hw.yaml')`.
# You can also use the `load_component_from_url` method, if your component yaml file is stored online. (For GitHub URLs, load the 'raw' file.)
# + [markdown] id="fkKFuQYDeFLB"
# Next, define two additional pipeline components.
#
# For example purposes, the first component below, `two_outputs`, installs the given `google-cloud-storage` package.
# (To keep the example code simple, the component function code won't actually use this import).
#
# This is one way that you can install the necessary package components.
# Alternatively, you can specify a base image that includes the necessary installations.
#
# The `two_outputs` component returns two named outputs. In the next section, you'll see how those outputs can be consumed by other pipeline steps.
#
# The second component below, `consumer`, takes three string inputs and prints them out.
# + id="HQNJpcK_m7YN"
@component(packages_to_install=["google-cloud-storage"])
def two_outputs(
text: str,
) -> NamedTuple(
"Outputs",
[
("output_one", str), # Return parameters
("output_two", str),
],
):
# the import is not actually used for this simple example, but the import
# is successful, as it was included in the `packages_to_install` list.
from google.cloud import storage # noqa: F401
o1 = f"output one from text: {text}"
o2 = f"output two from text: {text}"
print("output one: {}; output_two: {}".format(o1, o2))
return (o1, o2)
@component
def consumer(text1: str, text2: str, text3: str):
print(f"text1: {text1}; text2: {text2}; text3: {text3}")
# + [markdown] id="YTh_chI1PEpb"
# ### Define a pipeline that uses the components
#
# Next, define a pipeline that uses these three components.
#
# By evaluating the component definitions above, you've created task factory functions that you can use in the pipeline definition to create pipeline steps.
#
# The pipeline takes an input parameter, and passes that parameter as an argument to the first two pipeline steps (`hw_task` and `two_outputs_task`).
#
# Then, the third pipeline step (`consumer_task`) consumes the outputs of the first and second steps. Because the `hello_world` component definition just returns one unnamed output, it can be referred to as `hw_task.output`. The `two_outputs` task returns two named outputs, which you can access as `two_outputs_task.outputs["<output_name>"]`.
#
# Note that in the `@dsl.pipeline` decorator, you're defining the `PIPELINE_ROOT` Cloud Stora path to use. If you had not included that info here, it would be required to specify it when creating the pipeline run, as you'll see below.
# + id="UZ32XAM_PEpf"
@dsl.pipeline(
name="hello-world-v2",
description="A simple intro pipeline",
pipeline_root=PIPELINE_ROOT,
)
def intro_pipeline(text: str = "hi there"):
hw_task = hello_world(text)
two_outputs_task = two_outputs(text)
consumer_task = consumer( # noqa: F841
hw_task.output,
two_outputs_task.outputs["output_one"],
two_outputs_task.outputs["output_two"],
)
# + [markdown] id="2Hl1iYEKSzjP"
# ## Compile and run the pipeline
#
# Now, you're ready to compile the pipeline:
# + id="7PwUFV-MleGs"
from kfp.v2 import compiler # noqa: F811
compiler.Compiler().compile(
pipeline_func=intro_pipeline, package_path="hw_pipeline_job.json"
)
# + [markdown] id="qfNuzFswBB4g"
# The pipeline compilation generates the `hw_pipeline_job.json` job spec file.
#
# Next, instantiate an API client object:
# + id="Hl5Q74_gkW2c"
from kfp.v2.google.client import AIPlatformClient # noqa: F811
api_client = AIPlatformClient(
project_id=PROJECT_ID,
region=REGION,
)
# + [markdown] id="_jrn6saiQsPh"
# Then, you run the defined pipeline like this:
# + id="R4Ha4FoDQpkd"
response = api_client.create_run_from_job_spec(
job_spec_path="hw_pipeline_job.json",
# pipeline_root=PIPELINE_ROOT # this argument is necessary if you did not specify PIPELINE_ROOT as part of the pipeline definition.
)
# + [markdown] id="GvBTCP318RKs"
# Click on the generated link to see your run in the Cloud Console. It should look like this:
#
# <a href="https://storage.googleapis.com/amy-jo/images/mp/intro_pipeline.png" target="_blank"><img src="https://storage.googleapis.com/amy-jo/images/mp/intro_pipeline.png" width="60%"/></a>
# + [markdown] id="3QdnVXcwpZAi"
# ## Recurring pipeline runs: create a scheduled pipeline job
#
# This section shows how to create a **scheduled pipeline job**. You'll do this using the pipeline defined in the previous section.
#
# Under the hood, the scheduled jobs are supported by the Cloud Scheduler and a Cloud Functions function. Check first that the APIs for both of these services are enabled.
# You will need to first enable the [enable the Cloud Scheduler API](http://console.cloud.google.com/apis/library/cloudscheduler.googleapis.com) and the [Cloud Functions and Cloud Build APIs](https://console.cloud.google.com/flows/enableapi?apiid=cloudfunctions,cloudbuild.googleapis.com) if you have not already done so. Note also that you may need to [create an App Engine app for your project](https://cloud.google.com/scheduler/docs/quickstart) if one does not already exist.
#
#
# See the [Cloud Scheduler](https://cloud.google.com/scheduler/docs/configuring/cron-job-schedules) documentation for more on the cron syntax.
#
# + [markdown] id="kjksVdC_0xLt"
# Create a scheduled pipeline job, passing as an arg the job spec file that you compiled above.
#
# Note that you can pass a `parameter_values` dict that specifies the pipeline input parameters you want to use.
# + id="RvUu12Rj6G8Z"
# adjust time zone and cron schedule as necessary
response = api_client.create_schedule_from_job_spec(
job_spec_path="hw_pipeline_job.json",
schedule="2 * * * *",
time_zone="America/Los_Angeles", # change this as necessary
parameter_values={"text": "Hello world!"},
# pipeline_root=PIPELINE_ROOT # this argument is necessary if you did not specify PIPELINE_ROOT as part of the pipeline definition.
)
# + [markdown] id="6Dvbvc_ksuxE"
# Once the scheduled job is created, you can see it listed in the [Cloud Scheduler](https://console.cloud.google.com/cloudscheduler/) panel in the Console.
#
# <a href="https://storage.googleapis.com/amy-jo/images/kf-pls/pipelines_scheduler.png" target="_blank"><img src="https://storage.googleapis.com/amy-jo/images/kf-pls/pipelines_scheduler.png" width="95%"/></a>
#
# You can test the setup from the Cloud Scheduler panel by clicking 'RUN NOW'.
#
# > **Note**: The implementation is using a Cloud Functions function, which you can see listed in the [Cloud Functions](https://console.cloud.google.com/functions/list) panel in the console as `templated_http_request-v1`.
# Don't delete this function, as it will prevent the Cloud Scheduler jobs from actually kicking off the pipeline run. If you do delete it, create a new scheduled job in order to recreate the function.
#
# When you're done experimenting, you probably want to **PAUSE** your scheduled job from the Cloud Scheduler panel, so that the recurrent jobs do not keep running.
# + [markdown] id="O4gCLMgYNlJb"
# ## Specifying a service account to use for a pipeline run
#
# By default, the [service account](https://cloud.google.com/iam/docs/service-accounts) used for your pipeline run is your [default compute engine service account](https://cloud.google.com/compute/docs/access/service-accounts#default_service_account).
# However, you might want to run pipelines with permissions to access different roles than those configured for your default SA (e.g. perhaps using a more restricted set of permissions).
# If you want to execute your pipeline using a different service account, this is straightforward to do. You just need to give the new service account the correct permissions.
# + [markdown] id="ZIStAYOVRfOl"
# ### Create a service account (quick start version)
#
# See the documentation for more details on the process of creating and configuring a service account to work with AI Platform Pipelines. In brief:
#
# - Go to the Cloud Console -> IAM & Admin -> Service Accounts, click “**CREATE SERVICE ACCOUNT**”.
# - Enter a name and click “**CREATE**”.
# - In the second step of the process, give the service account “AI Platform User” and “Storage Admin” roles
# - In the third step, add yourself as the Service account User and Admin.
# - Open the service account permission tab, and click “**Grant Access**”.
# - Give the “AI Platform Custom Code Service Agent” (`<EMAIL>-<PROJECT_<EMAIL>>@<EMAIL>` & `service-<PROJECT_<EMAIL>>@<EMAIL>`) the “Service Account Token Creator” role.
#
# + [markdown] id="XrH_JBGETX4g"
# Once your service account is configured, you can pass it as an arg to the `create_run_from_job_spec` method, as follows:
# + id="V_0rTpLNAEt2"
SERVICE_ACCOUNT = (
"<service-account-<EMAIL>" # <--- CHANGE THIS
)
api_client = AIPlatformClient(
project_id=PROJECT_ID,
region=REGION,
)
response = api_client.create_run_from_job_spec(
job_spec_path="hw_pipeline_job.json", # <--- CHANGE THIS IF YOU WANT TO RUN OTHER PIPELINES
pipeline_root=PIPELINE_ROOT,
service_account=SERVICE_ACCOUNT,
)
# + [markdown] id="ZjweeCVdgB5w"
# The pipeline job runs with the permissions of the given service account.
# + [markdown] id="Xm46quDiBp7M"
# ## Using the Pipelines REST API
#
# At times you may want to use the REST API instead of the Python KFP SDK. Below are examples of how to do that.
#
# Where a command requires a pipeline ID, you can get that info from the "Run" column in the pipelines list— as shown below— as well as from the 'details' page for a given pipeline. You can also see that info when you list pipeline job(s) via the API.
#
# <a href="https://storage.googleapis.com/amy-jo/images/mp/pipeline_id.png" target="_blank"><img src="https://storage.googleapis.com/amy-jo/images/mp/pipeline_id.png" width="80%"/></a>
#
# > Note: Currently, the KFP SDK does not support all the API calls below.
#
# + id="sby6SIGXB_3O"
ENDPOINT = REGION + "-aiplatform.googleapis.com"
# + [markdown] id="oPPjZYXnCa_W"
# ### List pipeline jobs
#
# (This request may generate a large response if you have many pipeline runs).
# + id="hAC_ZCMZJKTJ"
# ! curl -X GET -H "Authorization: Bearer $(gcloud auth print-access-token)" -H "Content-Type: application/json" https://{ENDPOINT}/v1beta1/projects/{PROJECT_ID}/locations/{REGION}/pipelineJobs
# + [markdown] id="Je-HJUUp0MAi"
# ### Create a pipeline job
#
# For this API request, you need to submit a compiled pipeline job spec. We're using the one we generated above, "hw_pipeline_job.json".
# + id="fXDF-br4r220"
# ! curl -X POST -H "Authorization: Bearer $(gcloud auth print-access-token)" -H "Content-Type: application/json; charset=utf-8" https://{ENDPOINT}/v1beta1/projects/{PROJECT_ID}/locations/{REGION}/pipelineJobs --data "@sample_pipeline.json"
# + [markdown] id="cjuuG8GRCiNe"
# ### Get a pipeline job from its ID
# + id="BFtoTo64gnqV"
PIPELINE_RUN_ID = "xxxx" # <---CHANGE THIS
# ! curl -X GET -H "Authorization: Bearer $(gcloud auth print-access-token)" -H "Content-Type: application/json" https://{ENDPOINT}/v1beta1/projects/{PROJECT_ID}/locations/{REGION}/pipelineJobs/{PIPELINE_RUN_ID}
# + [markdown] id="RDKhnFrNCol3"
# ### Cancel a pipeline job given its ID
# + id="6efo3RjWhFDp"
PIPELINE_RUN_ID_TO_CANCEL = "xxxx" # <---CHANGE THIS
# ! curl -X POST -H "Authorization: Bearer $(gcloud auth print-access-token)" -H "Content-Type: application/json" https://{ENDPOINT}/v1beta1/projects/{PROJECT_ID}/locations/{REGION}/pipelineJobs/{PIPELINE_RUN_ID_TO_CANCEL}:cancel
# + [markdown] id="s9Tc0cO8C1OF"
# ### Delete a pipeline job given its ID
# + id="lu2IAHhWijDY"
PIPELINE_RUN_ID_TO_DELETE = "xxxx" # <---CHANGE THIS
# ! curl -X DELETE -H "Authorization: Bearer $(gcloud auth print-access-token)" -H "Content-Type: application/json" https://{ENDPOINT}/v1beta1/projects/{PROJECT_ID}/locations/{REGION}/pipelineJobs/{PIPELINE_RUN_ID_TO_DELETE}
# + [markdown] id="6yRt6JlmrGuG"
# ## Cleaning up
#
# To clean up all Google Cloud resources used in this project, you can [delete the Google Cloud
# project](https://cloud.google.com/resource-manager/docs/creating-managing-projects#shutting_down_projects) you used for the tutorial.
#
# Otherwise, you can delete the individual resources you created in this tutorial:
# - delete Cloud Storage objects that were created. Uncomment and run the command in the cell below **only if you are not using the `PIPELINE_ROOT` path for any other purpose**.
#
# + id="mkdPFlpwrGuP"
# Warning: this command will delete ALL Cloud Storage objects under the PIPELINE_ROOT path.
# # ! gsutil -m rm -r $PIPELINE_ROOT
| ai-platform-unified/notebooks/unofficial/pipelines/pipelines_intro_kfp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Sistema: dS/dt=-bS, dI/dt=bI, b es positivo (b=beta)
from sympy import *
from sympy.abc import S,I,t,b
# + tags=[]
#puntos criticos
P=-b*S
Q=b*I
#establecer P(S,I)=0 y Q(S,I)=0
Peqn=Eq(P,0)
Qeqn=Eq(Q,0)
print(solve((Peqn,Qeqn),S,I))
#Eigenvalores y eigenvectores
M=Matrix([[-b,0],[0,b]])
print(M.eigenvals())
pprint(M.eigenvects())
# -
# El sistema tiene un punto critico en el origen el cual tiene eigenvalores -b y b, ya que b es postivo los eigenvalores son reales distintos uno positivo y otro negativo por lo que el punto critico es un punto silla, el cual es inestable.
# Ya que el sistema tiene un punto critico que es un punto silla, por la definicion 9 el indice del punto critico es -1 y por el teorema 1 entonces no hay ciclos limites en el sistema, es decir no existe una solucion periodica aislada del sistema.
import matplotlib.pyplot as plt
import numpy as np
from scipy.integrate import odeint
import pylab as pl
import matplotlib
def dx_dt(x,t):
return [ -0.5*x[0] , 0.5*x[1] ]
#trayectorias en tiempo hacia adelante
ts=np.linspace(0,10,500)
ic=np.linspace(20000,100000,3)
for r in ic:
for s in ic:
x0=[r,s]
xs=odeint(dx_dt,x0,ts)
plt.plot(xs[:,0],xs[:,1],"-", color="orangered", lw=1.5)
#trayectorias en tiempo hacia atras
ts=np.linspace(0,-10,500)
ic=np.linspace(20000,100000,3)
for r in ic:
for s in ic:
x0=[r,s]
xs=odeint(dx_dt,x0,ts)
plt.plot(xs[:,0],xs[:,1],"-", color="orangered", lw=1.5)
#etiquetas de ejes y estilo de letra
plt.xlabel('S',fontsize=20)
plt.ylabel('I',fontsize=20)
plt.tick_params(labelsize=12)
plt.ticklabel_format(style="sci", scilimits=(0,0))
plt.xlim(0,100000)
plt.ylim(0,100000)
#campo vectorial
X,Y=np.mgrid[0:100000:15j,0:100000:15j]
u=-0.5*X
v=0.5*Y
pl.quiver(X,Y,u,v,color='dimgray')
plt.savefig("SI.pdf",bbox_inches='tight')
plt.show()
# Analisis de existencia de Bifurcaciones
# El punto critico del sistema no varia con el cambio en el valor de b (beta) ya que es idependiente de este parametro.
| ModeloSI.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Restricted Boltzman Machine on MNIST
#
# Created by: <NAME> (<EMAIL>)
#
# +
import tensorflow as tf
import numpy as np
from twodlearn.tf_lib.DBM import *
import math
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
from IPython.display import clear_output
# %matplotlib inline
# -
# ## 1. Load MNIST
# +
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
batch_X, batch_y = mnist.train.next_batch(100)
print(batch_X.shape)
print(np.reshape(batch_X, [-1,28,28,1]).shape)
print(batch_y.shape)
print(np.max(batch_X), np.min(batch_X))
# -
# ## 2. Model definition
sess = tf.InteractiveSession()
# +
n_inputs= 28*28
n_hidden= 1000
rbm_layer = RBM(n_inputs, n_hidden)
# +
batch_size= 500
x = tf.placeholder( tf.float32, shape=(batch_size, n_inputs))
cd_step, gen_model= rbm_layer.evaluate_cd_step(x, k= 10, alpha= 0.001)
# -
# ## 3. Train the model
tf.initialize_all_variables().run()
print('Initialized')
# +
num_steps= 5000
n_logging = 100
for step in range(num_steps):
# -------- train RBM -----#
batch_x, _= mnist.train.next_batch(batch_size)
[_, x_g] = sess.run([cd_step, gen_model], feed_dict= {x : batch_x})
# ------- logging -------
if step%n_logging == 0:
clear_output()
x_gaux = np.reshape(x_g, [-1,28,28,1])
imgplot = plt.imshow(x_gaux[1,:,:,0], cmap='Greys_r')
plt.show()
# -
print(x_g.shape)
x_gaux = np.reshape(x_g, [-1,28,28,1])
idx = 0
imgplot = plt.imshow(x_gaux[idx,:,:,0])
idx += 1
# ## Create random samples from the model
# +
h_test = tf.placeholder( tf.float32, shape=(50, n_hidden))
gen_model2 = rbm_layer.gibbs_sampling_given_h(h_test, k=500)
# -
batch_h = np.random.rand(50, n_hidden)
[x_g] = sess.run([gen_model2], feed_dict= {h_test : batch_h})
x_gaux = np.reshape(x_g, [-1,28,28,1])
idx = 0
# +
imgplot = plt.imshow(x_gaux[idx,:,:,0], cmap='Greys_r')
idx += 1
# -
| twodlearn/Examples/rbm_mnist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 5407, "status": "ok", "timestamp": 1622149679245, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10622192417315035630"}, "user_tz": -120} id="9GeMELQL5F25" outputId="1ddf29c4-e09e-4a42-958d-0ebafce91023"
# %%shell
# Download TorchVision repo to use some files from
# references/detection
git clone https://github.com/pytorch/vision.git
# cd vision
git checkout v0.3.0
# cp references/detection/utils.py ../
# cp references/detection/transforms.py ../
# cp references/detection/coco_eval.py ../
# cp references/detection/engine.py ../
# cp references/detection/coco_utils.py ../
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 45931, "status": "ok", "timestamp": 1622149734253, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10622192417315035630"}, "user_tz": -120} id="h_dD8X1qCMWp" outputId="1cfd8d4f-25dd-453b-cfa7-fdac30d51776"
# mount google drive where the zipped file of images located
from google.colab import drive
drive.mount('/content/gdrive')
# unzip the file
# !unzip "/content/gdrive/My Drive/OleanderDataset.zip"
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3407, "status": "ok", "timestamp": 1622149741595, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10622192417315035630"}, "user_tz": -120} id="wHgDa7O03a3l" outputId="8737d9e7-810e-421e-e395-35f3c4f5852f"
#Initialisation of dataset and CNN structure
# define the dataset
import os
import numpy as np
import torch
import torch.utils.data
from PIL import Image
class NNDataset(torch.utils.data.Dataset):
def __init__(self, root, transforms=None):
self.root = root
self.transforms = transforms
# load all image files, sorting them to
# ensure that they are aligned
self.imgs = list(sorted(os.listdir(os.path.join(root, "Images"))))
self.masks = list(sorted(os.listdir(os.path.join(root, "Masks"))))
def __getitem__(self, idx):
# load images ad masks
img_path = os.path.join(self.root, "Images", self.imgs[idx])
mask_path = os.path.join(self.root, "Masks", self.masks[idx])
img = Image.open(img_path).convert("RGB")
# convert masks to grayscale mode to distinguish background and objects
mask = Image.open(mask_path).convert("L")
mask = np.array(mask)
# instances are encoded as different colors
obj_ids = np.unique(mask)
# first id is the background, so remove it
obj_ids = obj_ids[1:]
# split the color-encoded mask into a set
# of binary masks
masks = mask == obj_ids[:, None, None]
# get bounding box coordinates for each mask
num_objs = len(obj_ids)
boxes = []
for i in range(num_objs):
pos = np.where(masks[i])
xmin = np.min(pos[1])
xmax = np.max(pos[1])
ymin = np.min(pos[0])
ymax = np.max(pos[0])
boxes.append([xmin, ymin, xmax, ymax])
boxes = torch.as_tensor(boxes, dtype=torch.float32)
# there is only one class
labels = torch.ones((num_objs,), dtype=torch.int64)
masks = torch.as_tensor(masks, dtype=torch.uint8)
image_id = torch.tensor([idx])
area = (boxes[:, 3] - boxes[:, 1]) * (boxes[:, 2] - boxes[:, 0])
# suppose all instances are not crowd
iscrowd = torch.zeros((num_objs,), dtype=torch.int64)
target = {}
target["boxes"] = boxes
target["labels"] = labels
target["masks"] = masks
target["image_id"] = image_id
target["area"] = area
target["iscrowd"] = iscrowd
if self.transforms is not None:
img, target = self.transforms(img, target)
return img, target
def __len__(self):
return len(self.imgs)
# import Mask R-CNN architecture for fine-tuning
import torchvision
from torchvision.models.detection.faster_rcnn import FastRCNNPredictor
from torchvision.models.detection.mask_rcnn import MaskRCNNPredictor
def get_instance_segmentation_model(num_classes):
# load an instance segmentation model pre-trained on COCO
model = torchvision.models.detection.maskrcnn_resnet50_fpn(pretrained=True)
# get the number of input features for the classifier
in_features = model.roi_heads.box_predictor.cls_score.in_features
# replace the pre-trained head with a new one
model.roi_heads.box_predictor = FastRCNNPredictor(in_features, num_classes)
# now get the number of input features for the mask classifier
in_features_mask = model.roi_heads.mask_predictor.conv5_mask.in_channels
hidden_layer = 256
# and replace the mask predictor with a new one
model.roi_heads.mask_predictor = MaskRCNNPredictor(in_features_mask,
hidden_layer,
num_classes)
return model
# determine the version of cuda
# !cat /usr/local/cuda/version.txt
# import helper functions
from engine import train_one_epoch, evaluate
import utils
import transforms as T
def get_transform(train):
transforms = []
# converts the image, a PIL image, into a PyTorch Tensor
transforms.append(T.ToTensor())
if train:
# during training, randomly flip the training images
# and ground-truth for data augmentation
transforms.append(T.RandomHorizontalFlip(0.5))
# NB: may add rotation and resize operations
return T.Compose(transforms)
# use our dataset and defined transformations
dataset = NNDataset('Oleander Plant', get_transform(train=True))
dataset_test = NNDataset('Oleander Plant', get_transform(train=False))
# split the dataset in train and test set
torch.manual_seed(1)
indices = torch.randperm(len(dataset)).tolist()
print(len(dataset))
# NB: change the portion of train and test set according to the size of the whole dataset
dataset = torch.utils.data.Subset(dataset, indices[:-6])
dataset_test = torch.utils.data.Subset(dataset_test, indices[-6:])
print(len(dataset))
# define training and validation data loaders
data_loader = torch.utils.data.DataLoader(
dataset, batch_size=2, shuffle=True, num_workers=2,
collate_fn=utils.collate_fn)
data_loader_test = torch.utils.data.DataLoader(
dataset_test, batch_size=1, shuffle=False, num_workers=2,
collate_fn=utils.collate_fn)
# + colab={"base_uri": "https://localhost:8080/", "height": 117, "referenced_widgets": ["6eae9fa5cd6a4f06b2c0ab9421c17e13", "a266f288e7fd4975936a6095dd871cd6", "92f1758d02b246f1a362c0cfd02b6532", "f18f285e3d7c44f3959d1a980379c537", "f6e45587ce844b4baddaff6be1cdbcb1", "d28b5897a53a489facee7baaffb324f1", "79daa0dba146491f8259935ae9ed6420", "483080375f9d41249ec6e11159f10c8e"]} executionInfo={"elapsed": 2694, "status": "ok", "timestamp": 1622149753639, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10622192417315035630"}, "user_tz": -120} id="PwGf7KUTB8yX" outputId="02c585b5-d090-4a8d-8510-5229caa29eaf"
#
if torch.cuda.is_available():
device = torch.device('cuda')
print(torch.cuda.is_available())
print('cuda')
else:
device = torch.device('cpu')
print(torch.cuda.is_available())
print('cpu')
# our dataset has two classes only - background and label
num_classes = 2
# get the model using our helper function
model = get_instance_segmentation_model(num_classes)
# move model to the right device
model.to(device)
# construct an optimizer
params = [p for p in model.parameters() if p.requires_grad]
optimizer = torch.optim.SGD(params, lr=0.005,
momentum=0.9, weight_decay=0.0005)
# and a learning rate scheduler which decreases the learning rate by
# 10x every 3 epochs
lr_scheduler = torch.optim.lr_scheduler.StepLR(optimizer,
step_size=3,
gamma=0.1)
# + colab={"base_uri": "https://localhost:8080/", "height": 564} executionInfo={"elapsed": 1789243, "status": "error", "timestamp": 1622151549110, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10622192417315035630"}, "user_tz": -120} id="WY6FLoPe4J4t" outputId="925c4d35-a9ef-4a6f-a86e-6e67e6ef2016"
# CNN training
# train for epochs
# NB: change number of epochs according to the size of dataset
# few epochs for small dataset to avoid overfitting
num_epochs = 2
for epoch in range(num_epochs):
# train for one epoch, printing every 5 iteration
train_one_epoch(model, optimizer, data_loader, device, epoch, print_freq=5)
# update the learning rate
lr_scheduler.step()
# evaluate on the test dataset
evaluate(model, data_loader_test, device=device)
# + id="_NcRvbVq4hlb"
#save model
import torch
import torch.onnx as onnx
import torchvision.models as models
# + id="eg-DiQ7R4kiT"
# test
# pick one image from the test set
img, _ = dataset_test[1]
# put the model in evaluation mode
model.eval()
with torch.no_grad():
prediction = model([img.to(device)])
prediction
# + id="HLDO0bsU4sjT"
# view verification image
Image.fromarray(img.mul(255).permute(1, 2, 0).byte().numpy()).convert('L')
# + id="2Nq8kzNM4wx7"
# view prediction
Image.fromarray(prediction[0]['masks'][0, 0].mul(255).byte().cpu().numpy()).convert('L')
# + executionInfo={"elapsed": 199, "status": "aborted", "timestamp": 1622151549097, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10622192417315035630"}, "user_tz": -120} id="tCCSSNNr6FyP"
while True:pass
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3081, "status": "ok", "timestamp": 1621864321801, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "10622192417315035630"}, "user_tz": -120} id="lxHEqlWQOJp_" outputId="eaead1be-8064-4349-b23d-b9dd948dacef"
# !pip freeze
| OleanderCNN(5_5).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 06 网格搜索和更多kNN中的超参数
# + jupyter={"outputs_hidden": true}
import numpy as np
from sklearn import datasets
# + jupyter={"outputs_hidden": true}
digits = datasets.load_digits()
X = digits.data
y = digits.target
# + jupyter={"outputs_hidden": true}
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.2)
# -
from sklearn.neighbors import KNeighborsClassifier
knn_clf = KNeighborsClassifier(n_neighbors=3)
knn_clf.fit(X_train, y_train)
knn_clf.score(X_test, y_test)
# # 网格搜索
# 先创建网格参数,p是明科夫斯基参数
param_grid = [
{
'weights':['uniform'],
'n_neighbors':[i for i in range(1, 11)]
},
{
'weights':['distance'],
'n_neighbors':[i for i in range(1, 11)],
'p':[i for i in range(1, 6)]
}
]
# + jupyter={"outputs_hidden": true}
knn_clf = KNeighborsClassifier()
# + jupyter={"outputs_hidden": true}
from sklearn.model_selection import GridSearchCV
grid_search = GridSearchCV(knn_clf, param_grid)
# -
# %%time
grid_search.fit(X_train, y_train)
grid_search.best_score_
grid_search.best_params_
# + jupyter={"outputs_hidden": true}
knn_clf = grid_search.best_estimator_ # 选取最佳模型
# -
knn_clf.score(X_test, y_test)
# %%time
# 并发进行模型获取
grid_search = GridSearchCV(knn_clf, param_grid, n_jobs=-1, verbose=2) # njobs代表用几个核。-1代表用所有的核.verbose代表信息输出的详细程度
grid_search.fit(X_train, y_train)
# + jupyter={"outputs_hidden": true}
| Part5Improve/04-kNN/06-More-Hyper-Parameters-in-kNN-and-Grid-Search/06-More-Hyper-Parameters-in-kNN-and-Grid-Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
# +
a = tf.constant(100)
b = tf.constant(200)
add_op = a+b
sess = tf.Session()
result = sess.run(add_op)
print(result)
# +
a = " asd fff ";
print(a)
# -
print(a.strip())
# +
#newFile = open("./myText.txt", 'w') # just write from new
newFile = open("./myText.txt", 'a') # add
newFile.write('%s %s' % ('it\'s', 'just'))
newFile.write("\n ok next line.\n")
newFile.close()
# +
# %matplotlib inline
import matplotlib.pyplot as plt
x = [ a for a in range(-100, 101) ]
y = [ ( a**2 ) - 2000 for a in x ]
gab,=plt.plot(x, y, 'r-')
plt.show()
# -
# +
import os
import config as conf
contentsSize = 7
magazinePath = os.path.join(conf.data_root, 'magazine.json')
metadataPath = os.path.join(conf.data_root, 'metadata.json')
usersPath = os.path.join(conf.data_root, 'users.json')
contentsPath = [ os.path.join(conf.data_root, 'contents/data.' + str(i) ) for i in range(0, contentsSize) ]
print(magazinePath)
print(metadataPath)
print(usersPath)
print(contentsPath)
import json
outputFolder = './tmp'
# -
'''
import json
## json 형식 변환
## json_data = json.JSONEncoder().encode(data)
## 딕셔너리 형태로 복구
## data = json.JSONDecoder().decode(json_data)
contents = [ open(contentsPath[i], 'r') for i in range(0, contentsSize) ]
# each file line counts
100000
100000
100000
100000
100000
100000
42190
# 단어 카운트
# words = {}
# 메모리 부족으로 일단 나눠서 저장
outputFolder = './tmp'
MAX_LINE = 1
l = 0;
for i in range(0, contentsSize) :
wordsPath = os.path.join(outputFolder, 'words' + str(i) + '.txt')
wordsFile = open(wordsPath, 'w')
words = {}
for jsonLine in contents[i] :
# ============= for test
# if l >= MAX_LINE : break
# l = l + 1
# ============= for test
# convert json to dictionary
line = json.JSONDecoder().decode(jsonLine)
# { "chars": [[]], "morphs": , "id" }
# print(line['id'])
# print(line['morphs'])
for morphs in line['morphs'] :
for morph in morphs :
words[morph] = words.get(morph, 0) + 1
wordsFile.write(json.JSONEncoder().encode(words))
wordsFile.close()
contents[i].close()
'''
# +
'''
저장한 words0.txt ~ words1.txt 를 순회하며
{ 'word1' : count, 'word2' : count, ... } 를 생성 및 저장 -> wordsAll.txt
'''
allDic = {}
for i in range(0, contentsSize) :
# if i != targetIdx : continue
wordsFile = open(os.path.join(outputFolder, 'words' + str(i) + '.txt'), 'r')
temp_a = json.JSONDecoder().decode(wordsFile.read())
if i == 0 : allDic = temp_a
else :
for k, v in temp_a.items() :
allDic[k] = allDic.get(k, 0) + int(v)
wordsFile.close()
print(len(allDic))
wordsFile = open(os.path.join(outputFolder, 'wordsAll.txt'), 'w')
wordsFile.write(json.JSONEncoder().encode(allDic))
wordsFile.close()
# +
wordsAll = {}
with open(os.path.join(outputFolder, 'wordsAll.txt'), 'r') as fall :
wordsAll = json.JSONDecoder().decode(fall.read())
wordsAll = [ (k, v) for k, v in wordsAll.items() ]
wordsAll.sort(key=lambda x:x[1], reverse=True)
print(wordsAll[:10])
# +
# %matplotlib inline
import matplotlib.pyplot as plt
top50w = wordsAll[:50]
x = [ k for k, _ in top50w ]
y = [ v for _, v in top50w ]
gab,=plt.plot(x, y, 'r-')
plt.show()
# +
# doc2vec
from gensim.models import doc2vec
from collections import namedtuple
import json
## json 형식 변환
## json_data = json.JSONEncoder().encode(data)
## 딕셔너리 형태로 복구
## data = json.JSONDecoder().decode(json_data)
contents = [ open(contentsPath[i], 'r') for i in range(0, contentsSize) ]
# each file line counts
# 100000
# 100000
# 100000
# 100000
# 100000
# 100000
# 42190
outputFolder = './tmp'
targetIndex = 0;
for i in range(0, contentsSize) :
if targetIndex != i : continue
wordsPath = os.path.join(outputFolder, 'vec' + str(i) + '.txt')
wordsFile = open(wordsPath, 'w')
tagged_train_docs = []
for jsonLine in contents[i] :
# convert json to dictionary
line = json.JSONDecoder().decode(jsonLine)
# { "chars": [[]], "morphs": , "id" }
# print(line['id'])
# print(line['morphs'])
docId = line['id']
TaggedDocument = namedtuple('TaggedDocument', 'words tags')
mList = []
for morphs in line['morphs'] :
mList.extend(morphs)
tagged_train_docs.append( TaggedDocument(docId, mList) )
contents[i].close()
print(tagged_train_docs.head())
# -
| mydev1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Import packages
import numpy as np
import time
# ### Define the Sigmoid Function and it's derivative
# +
#Sigmoid Function
def sigmoid(x):
s = 1.0/(1 + np.exp(-x))
return s
#Function to calculate derivative of the Sigmoid
def sigmoid_derivative(s):
ds = s * (1 - s)
return ds
# -
# ### Define the TanH Function and it's derivative
# +
#TanH Function
def tanh(x):
t = np.tanh(x)
return t
#Function to calculate derivative of the TanH
def tanh_derivative(t):
dt = 1 - t**2
return dt
# -
# ### Define the ReLU Function and it's derivative
# +
#ReLU Function
def ReLU(x):
r = np.maximum(0, x)
return r
#Function to calculate derivative of the ReLU
def ReLU_derivative(r):
if r.all() > 0:
dr = 1
if r.all() < 0:
dr = 0
return dr
# -
# ### Input and output values
#Define the XOR inputs and outputs
XOR_input = np.array([[0, 0], [0, 1], [1, 0], [1, 1]])
Y = np.array([[0], [1], [1], [0]])
# ### Initialize Weights and Bias
# +
#Define Weight and Bias of 1st Hidden Layer
w1 = np.random.uniform(size = [2, 2])
b1 = np.random.uniform(size = [1, 2])
#Define Weight and Bias of Output Neuron
w2 = np.random.uniform(size = [2, 1])
b2 = np.random.uniform(size = [1, 1])
# -
# ### Loop for 100001 times for training, uncomment the lines for choosing your preferred activation function (Sigmoid/ReLU/Tanh)
# +
#Define the learning rate
learning_rate = 0.01
s = time.clock()
for i in range(100001):
#Forward Propagation
#Compute the forward propagation of first hidden layer
A1 = np.dot(XOR_input, w1) + b1
#Activation Function
#Choose between Sigmoid/ReLU/tanh
Z1 = sigmoid(A1)
#Z1 = ReLU(A1)
#Z1 = tanh(A1)
#Compute the forward propagation
A2 = np.dot(Z1, w2) + b2
#Activation Function for output neuron
#Choose between Sigmoid/ReLU/tanh
#Yhat = sigmoid(A2)
#Yhat = ReLU(A2)
Yhat = tanh(A2)
#Backpropagation
#Calculate error
E = Y - Yhat
#Derivative of sigmoid at output layer
#Choose between Sigmoid/ReLU/tanh
#DE = E * sigmoid_derivative(Yhat)
#DE = E * ReLU_derivative(Yhat)
DE = E * tanh_derivative(Yhat)
#Calculate error
E2 = DE.dot(w2.T)
#Derivative at hidden layer
#Choose between Sigmoid/ReLU/tanh
DE2 = E2 * sigmoid_derivative(Z1)
#DE = E * ReLU_derivative(Yhat)
#DE2 = E * tanh_derivative(Yhat)
#Update weights and bias of output layer
w2 += Z1.T.dot(DE) * learning_rate
b2 += np.sum(DE, axis = 0, keepdims = True) * learning_rate
#Update weights and bias at hidden layer
w1 += XOR_input.T.dot(DE2) * learning_rate
b1 += np.sum(DE2, axis = 0, keepdims = True) * learning_rate
#Printing the cost and epoch at each thousandth epoch
if i % 1000 == 0:
print('Epoch = {} \n Error = {}'.format(i, E))
e = time.clock()
print('Final prediction = {}'.format(Yhat))
print('Time Elapsed = {}'.format(e-s))
| 03. Activation Functions/XOR - Activation Functions (Numpy).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
'''
File name: convert-trace-csv.ipynb
Author: <NAME>, PhD
Github: github.com/hamadaio
Date created: 2021-04-21
Date last modified: 2021-05-30
Python Version: 3.7.9
'''
from allensdk.core.cell_types_cache import CellTypesCache
# Instantiate the CellTypesCache instance. The manifest_file argument
# tells it where to store the manifest, which is a JSON file that tracks
# file paths. If you supply a relative path it will go into your
# current working directory
ctc = CellTypesCache()
# this saves the NWB file to 'cell_types/specimen_464212183/ephys.nwb'
cell_specimen_id = 571642127 # replace with relevant specimen_id
data_set = ctc.get_ephys_data(cell_specimen_id)
# +
# # %matplotlib inline # allows output graph to be printed inline if running in Jupyter Lab
# import init_CellTypesCache when splitting the blocks into individual files
import numpy as np
import csv
# Input manually the neural voltage trace numbers from the neuron 'Browse Electrophysiological Data' section. The example traces
# below are from a human neuron ID = 571471295 (https://celltypes.brain-map.org/experiment/electrophysiology/571471295)
neuralTraces = [18, 28, 27, 31, 32, 36, 39, 40]
for trace in neuralTraces:
with open(f'cell_id:{cell_specimen_id}--trace--{trace}.csv', "w") as neuralCSV:
writer = csv.writer(neuralCSV)
writer.writerow(["time (s)", "inputCurrent (pA)", "voltageResponse (mV)"])
trace_data = data_set.get_sweep(trace)
index_range = trace_data["index_range"]
inputCurrent = trace_data["stimulus"][0:index_range[1] + 1] # in Ampere
voltageResponse = trace_data["response"][0:index_range[1] + 1] # in Voltage
inputCurrent *= 1e12 # to pico-ampere (pA)
voltageResponse *= 1e3 # to milli-volt (mV)
sampling_rate = trace_data["sampling_rate"] # in Hz
time = np.arange(0, len(voltageResponse)) * (1.0 / sampling_rate) # in seconds
np.prod(inputCurrent.shape)
for i in range(np.prod(inputCurrent.shape)):
writer.writerow([time[i], inputCurrent[i], voltageResponse[i]])
print(sampling_rate / 1000, "kHz") # print sampling rate frequency
| src/convert-trace-csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python2
# name: python2
# ---
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
# # Python for Finance
# **Analyze Big Financial Data**
#
# O'Reilly (2014)
#
# <NAME>
# <img style="border:0px solid grey;" src="http://hilpisch.com/python_for_finance.png" alt="Python for Finance" width="30%" align="left" border="0">
# **Buy the book ** |
# <a href='http://shop.oreilly.com/product/0636920032441.do' target='_blank'>O'Reilly</a> |
# <a href='http://www.amazon.com/Yves-Hilpisch/e/B00JCYHHJM' target='_blank'>Amazon</a>
#
# **All book codes & IPYNBs** |
# <a href="http://oreilly.quant-platform.com">http://oreilly.quant-platform.com</a>
#
# **The Python Quants GmbH** | <a href='http://pythonquants.com' target='_blank'>www.pythonquants.com</a>
#
# **Contact us** | <a href='mailto:<EMAIL>'><EMAIL></a>
# # Financial Time Series
import warnings
warnings.simplefilter('ignore')
# ## pandas Basics
# + uuid="eda2a742-134d-4d47-8b30-557b846b9bb3"
import numpy as np
import pandas as pd
# -
# ### First Steps with DataFrame Class
# + uuid="f3be2d89-829a-49b2-96fc-07c475db1e3f"
df = pd.DataFrame([10, 20, 30, 40], columns=['numbers'],
index=['a', 'b', 'c', 'd'])
df
# + uuid="47b70a7b-710f-4c40-9a70-b09db7af1a12"
df.index # the index values
# + uuid="a36c6695-520d-4df1-a6fa-5f8362af37a3"
df.columns # the column names
# + uuid="c93aed37-21de-429d-86ed-9849e4c3e23c"
df.ix['c'] # selection via index
# + uuid="8c7c2f69-3673-40d9-a568-0471c629810d"
df.ix[['a', 'd']] # selection of multiple indices
# + uuid="c3ce0cc3-26e8-4256-ab8c-9a2d4b181633"
df.ix[df.index[1:3]] # selection via Index object
# + uuid="94b1d846-63df-49f4-8a7f-8fed03e5f4fa"
df.sum() # sum per column
# + uuid="4e73eb4f-352d-4527-b0c5-4f3a6e7eb354"
df.apply(lambda x: x ** 2) # square of every element
# + uuid="75206a83-0154-4be2-88d0-7a82a190fda1"
df ** 2 # again square, this time NumPy-like
# + uuid="49a2633a-b3c0-4d00-a227-e0ff4a8cf81d"
df['floats'] = (1.5, 2.5, 3.5, 4.5)
# new column is generated
df
# + uuid="c49b9aea-417a-4c2b-8e27-0e8771a77c87"
df['floats'] # selection of column
# + uuid="aa892c41-6637-45ed-876b-6a70285e4c0b"
df['names'] = pd.DataFrame(['Yves', 'Guido', 'Felix', 'Francesc'],
index=['d', 'a', 'b', 'c'])
df
# + uuid="584ac18c-161f-4c7b-8ff1-1cd406fb8437"
df.append({'numbers': 100, 'floats': 5.75, 'names': 'Henry'},
ignore_index=True)
# temporary object; df not changed
# + uuid="9068cd04-c6ff-4d0c-bd52-cf04cd89a0e9"
df = df.append(pd.DataFrame({'numbers': 100, 'floats': 5.75,
'names': 'Henry'}, index=['z',]))
df
# + uuid="0d526ac2-8691-4a59-83c5-712c800f0464"
df.join(pd.DataFrame([1, 4, 9, 16, 25],
index=['a', 'b', 'c', 'd', 'y'],
columns=['squares',]))
# temporary object
# + uuid="6558c7ff-f24e-4dd0-b57e-a30196bad1f4"
df = df.join(pd.DataFrame([1, 4, 9, 16, 25],
index=['a', 'b', 'c', 'd', 'y'],
columns=['squares',]),
how='outer')
df
# + uuid="3e863c7f-7875-4911-997b-6e48123dc1e5"
df[['numbers', 'squares']].mean()
# column-wise mean
# + uuid="c52173a0-485d-4eb2-b6b4-407d1ff2c30e"
df[['numbers', 'squares']].std()
# column-wise standard deviation
# -
# ### Second Steps with DataFrame Class
# + uuid="d6f56a00-91e6-4221-a1ec-6093f416d1be"
a = np.random.standard_normal((9, 4))
a.round(6)
# + uuid="450bd14d-7668-4f3f-a863-966f13562818"
df = pd.DataFrame(a)
df
# + uuid="968395a4-12bc-46d2-b486-6c767abce366"
df.columns = [['No1', 'No2', 'No3', 'No4']]
df
# + uuid="68e8d73f-93d3-47ac-a656-1edbdebcd1ff"
df['No2'][3] # value in column No2 at index position 3
# + uuid="a80e1e88-d211-4ee4-a6d3-90403a7739a8"
dates = pd.date_range('2015-1-1', periods=9, freq='M')
dates
# + uuid="d8fef9ed-25ca-4ae0-bd0c-026d340a903b"
df.index = dates
df
# + uuid="bcc38d60-3e1c-49bb-b883-ea7564c136b4"
np.array(df).round(6)
# -
# ### Basic Analytics
# + uuid="f760ea25-c64c-4e70-9f91-b72701d919ce"
df.sum()
# + uuid="3dd9bd77-eb80-46cb-87f3-62c053a8e223"
df.mean()
# + uuid="8e167ea8-09b7-4585-8cac-28fe20eefe66"
df.cumsum()
# + uuid="125980cc-91ec-4ab4-9a4a-cfd772dd1254"
df.describe()
# + uuid="9dfc1e40-c030-4a9c-9e3a-ff28c64a93df"
np.sqrt(df)
# + uuid="a540362b-50d7-4ef0-89ba-0b6ee38033f6"
np.sqrt(df).sum()
# + uuid="4b1834ec-9f9b-41d6-8d06-f2efc8433dc4"
%matplotlib inline
df.cumsum().plot(lw=2.0, grid=True)
# tag: dataframe_plot
# title: Line plot of a DataFrame object
# -
# ### Series Class
# + uuid="e86f82d1-5934-42d3-a986-f01bc829adaa"
type(df)
# + uuid="bcebc814-623d-4e8a-81e9-314ab36a7429"
df['No1']
# + uuid="ca241ef9-5359-4c89-bc92-be6346cb3959"
type(df['No1'])
# + uuid="b3d4cc90-e499-459c-88a5-011fde80d864"
import matplotlib.pyplot as plt
df['No1'].cumsum().plot(style='r', lw=2., grid=True)
plt.xlabel('date')
plt.ylabel('value')
# tag: time_series
# title: Line plot of a Series object
# -
# ### GroupBy Operations
# + uuid="4bc106dd-9590-4566-bc70-d410517c8223"
df['Quarter'] = ['Q1', 'Q1', 'Q1', 'Q2', 'Q2', 'Q2', 'Q3', 'Q3', 'Q3']
df
# + uuid="41c1962a-05ba-4c0f-b017-e6873e2d245e"
groups = df.groupby('Quarter')
# + uuid="804e567f-6b74-4405-a10e-d19d914655e7"
groups.mean()
# + uuid="7eb45e5c-b86f-4464-afd9-d5a3665e0f8e"
groups.max()
# + uuid="a871b95e-5946-4b09-b8dc-bc9503d2ff14"
groups.size()
# + uuid="542cf99a-bbf8-447e-9643-d6887ac74be7"
df['Odd_Even'] = ['Odd', 'Even', 'Odd', 'Even', 'Odd', 'Even',
'Odd', 'Even', 'Odd']
# + uuid="f5144c9f-ff37-4e35-9417-e39debdcd45b"
groups = df.groupby(['Quarter', 'Odd_Even'])
# + uuid="06904508-dbf1-431f-a3a2-681f29f03c51"
groups.size()
# + uuid="b8471956-40fc-4203-a54a-aaa45f5a3c00"
groups.mean()
# -
# ## Financial Data
# + uuid="9805e014-8a17-4e54-b6fd-1c77db7b6b78"
import pandas.io.data as web
# + uuid="53a33e39-a3ff-4c95-b0f2-a94d727ae0da"
DAX = web.DataReader(name='^GDAXI', data_source='yahoo',
start='2000-1-1')
DAX.info()
# + uuid="11984b1c-5248-4640-8f3b-a85040eb5683"
DAX.tail()
# + uuid="6185abc6-54c5-4711-b273-3252938f3e5e"
DAX['Close'].plot(figsize=(8, 5), grid=True)
# tag: dax
# title: Historical DAX index levels
# + active=""
# %%time
# DAX['Ret_Loop'] = 0.0
# for i in range(1, len(DAX)):
# DAX['Ret_Loop'][i] = np.log(DAX['Close'][i] /
# DAX['Close'][i - 1])
# + active=""
# DAX[['Close', 'Ret_Loop']].tail()
# + uuid="9b45b8c2-3b95-4c80-94a0-14f891cdd161"
%time DAX['Return'] = np.log(DAX['Close'] / DAX['Close'].shift(1))
# + uuid="5fbf45e9-dd56-40ba-8a75-086a80a04d5b"
DAX[['Close', 'Return', 'Return']].tail()
# + active=""
# del DAX['Ret_Loop']
# + uuid="8995980e-7fa1-482e-8996-3a0cc1050359"
DAX[['Close', 'Return']].plot(subplots=True, style='b',
figsize=(8, 5), grid=True)
# tag: dax_returns
# title: The DAX index and daily log returns
# + uuid="956890ca-7927-4fac-a99a-af7a15cac58f"
DAX['42d'] = pd.rolling_mean(DAX['Close'], window=42)
DAX['252d'] = pd.rolling_mean(DAX['Close'], window=252)
# + uuid="f5440e3f-b808-4685-9bec-5f6ca39609c5"
DAX[['Close', '42d', '252d']].tail()
# + uuid="281a5820-2a77-46b4-b399-8c0913423bc3"
DAX[['Close', '42d', '252d']].plot(figsize=(8, 5), grid=True)
# tag: dax_trends
# title: The DAX index and moving averages
# + uuid="1b12ae02-5e35-47ff-a9c9-8563e468b489"
import math
DAX['Mov_Vol'] = pd.rolling_std(DAX['Return'],
window=252) * math.sqrt(252)
# moving annual volatility
# + uuid="2e75f8fd-bcf8-4c36-93b9-974ef94366c7"
DAX[['Close', 'Mov_Vol', 'Return']].plot(subplots=True, style='b',
figsize=(8, 7), grid=True)
# tag: dax_mov_std
# title: The DAX index and moving, annualized volatility
# -
# ## Regression Analysis
# + uuid="85bf9df2-d445-4600-a02e-37cf2b7dc9ff"
import pandas as pd
from urllib import urlretrieve
# + uuid="17a2e317-7047-4c9f-9faf-7e0e4132bd01"
es_url = 'https://www.stoxx.com/document/Indices/Current/HistoricalData/hbrbcpe.txt'
vs_url = 'https://www.stoxx.com/document/Indices/Current/HistoricalData/h_vstoxx.txt'
urlretrieve(es_url, './data/es.txt')
urlretrieve(vs_url, './data/vs.txt')
!ls -o ./data/*.txt
# Windows: use dir
# + uuid="3bdd1237-41d5-4e92-8d2c-3c4ffd25e7e9"
lines = open('./data/es.txt', 'r').readlines()
lines = [line.replace(' ', '') for line in lines]
# + uuid="6c7769ea-4fb8-49ef-bdc8-4e06b986fb3e"
lines[:6]
# + uuid="b5edc764-13a4-4e0c-b6d3-ac615b4a530b"
for line in lines[3883:3890]:
print line[41:],
# + uuid="ea43adac-94fb-4b11-8af5-153c6fb4cebe"
new_file = open('./data/es50.txt', 'w')
# opens a new file
new_file.writelines('date' + lines[3][:-1]
+ ';DEL' + lines[3][-1])
# writes the corrected third line of the orginal file
# as first line of new file
new_file.writelines(lines[4:])
# writes the remaining lines of the orginial file
new_file.close()
# + uuid="aca0ad29-cce1-4da5-b39e-ace9bafe3077"
new_lines = open('./data/es50.txt', 'r').readlines()
new_lines[:5]
# + uuid="0bc55f0d-cd99-45b9-955e-3a126360e94f"
es = pd.read_csv('./data/es50.txt', index_col=0,
parse_dates=True, sep=';', dayfirst=True)
# + uuid="73526ac3-4bf0-4455-89b2-f6aa614ffdca"
np.round(es.tail())
# + uuid="e6e3100a-8296-494f-9758-bbb0006c5df4"
del es['DEL']
es.info()
# + uuid="fff2d2a1-dce8-4f4c-bab9-0b990a1f7b5f"
cols = ['SX5P', 'SX5E', 'SXXP', 'SXXE', 'SXXF',
'SXXA', 'DK5F', 'DKXF']
es = pd.read_csv(es_url, index_col=0, parse_dates=True,
sep=';', dayfirst=True, header=None,
skiprows=4, names=cols)
# + uuid="76793f6a-1625-4fc2-8063-38536b46b15e"
es.tail()
# + uuid="3a1920c2-8c61-4720-941e-afdb983350aa"
vs = pd.read_csv('./data/vs.txt', index_col=0, header=2,
parse_dates=True, dayfirst=True)
vs.info()
# + uuid="3a437278-4466-41bf-b7f2-f9c17d2c44a7"
import datetime as dt
data = pd.DataFrame({'EUROSTOXX' :
es['SX5E'][es.index > dt.datetime(1999, 1, 1)]})
data = data.join(pd.DataFrame({'VSTOXX' :
vs['V2TX'][vs.index > dt.datetime(1999, 1, 1)]}))
# + uuid="9223e142-d574-40c9-92e7-149d86628458"
data = data.fillna(method='ffill')
data.info()
# + uuid="fc5fc92a-3475-4e4b-a5fe-145809d35919"
data.tail()
# + uuid="07158c72-907f-4636-ad40-95182b7728e3"
data.plot(subplots=True, grid=True, style='b', figsize=(8, 6))
# tag: es50_vs
# title: The EURO STOXX 50 Index and the VSTOXX volatility index
# + uuid="17ae59ff-9863-4c30-9493-22d44c0e5edf"
rets = np.log(data / data.shift(1))
rets.head()
# + uuid="771c53bf-78fb-4260-865b-39307973cb77"
rets.plot(subplots=True, grid=True, style='b', figsize=(8, 6))
# tag: es50_vs_rets
# title: Log returns of EURO STOXX 50 and VSTOXX
# + uuid="709bc1e8-03a8-47c6-9b21-4efa08052dab"
xdat = rets['EUROSTOXX']
ydat = rets['VSTOXX']
model = pd.ols(y=ydat, x=xdat)
model
# + uuid="ff00a7f1-173c-40f8-b077-9892ad310f4a"
model.beta
# + uuid="24c708df-1e81-48c6-b1c2-890dd52e541f"
plt.plot(xdat, ydat, 'r.')
ax = plt.axis() # grab axis values
x = np.linspace(ax[0], ax[1] + 0.01)
plt.plot(x, model.beta[1] + model.beta[0] * x, 'b', lw=2)
plt.grid(True)
plt.axis('tight')
plt.xlabel('EURO STOXX 50 returns')
plt.ylabel('VSTOXX returns')
# tag: scatter_rets
# title: Scatter plot of log returns and regression line
# + uuid="e1f9009e-5b73-4e04-9b10-deea36f4e508"
rets.corr()
# + uuid="a534d3db-df59-4a31-b0b7-1ab3c19d77d0"
pd.rolling_corr(rets['EUROSTOXX'], rets['VSTOXX'],
window=252).plot(grid=True, style='b')
# tag: roll_corr
# title: Rolling correlation between EURO STOXX 50 and VSTOXX
# -
# ## High Frequency Data
# + uuid="cd3bd5f2-565a-4158-a796-e8ec41f76d88"
import numpy as np
import pandas as pd
import datetime as dt
from urllib import urlretrieve
%matplotlib inline
# + uuid="63521c3e-8197-464a-a5bc-dc9a15550893"
url1 = 'http://www.netfonds.no/quotes/posdump.php?'
url2 = 'date=%s%s%s&paper=NKE.N&csv_format=csv'
url = url1 + url2
# + uuid="8fa05c35-a914-49b2-8da1-841146b43931"
year = '2015'
month = '08'
days = ['03', '04', '05', '06', '07']
# dates might need to be updated
# + uuid="946e1882-04bb-4809-869f-1e7dd8ad396a"
NKE = pd.DataFrame()
for day in days:
NKE = NKE.append(pd.read_csv(url % (year, month, day),
index_col=0, header=0, parse_dates=True))
NKE.columns = ['bid', 'bdepth', 'bdeptht', 'offer', 'odepth', 'odeptht']
# shorter colummn names
# + uuid="1fce3fb2-c664-4f77-80dc-0e4907f86dac"
NKE.info()
# + uuid="b0fa9c58-2087-4f99-a7f1-efeb490bb456"
NKE['bid'].plot(grid=True)
# tag: aapl
# title: Nike stock tick data for a week
# + uuid="73447893-dc77-488a-8252-64c29a92e4c0"
to_plot = NKE[['bid', 'bdeptht']][
(NKE.index > dt.datetime(2015, 8, 4, 0, 0))
& (NKE.index < dt.datetime(2015, 8, 5, 2, 59))]
# adjust dates to given data set
to_plot.plot(subplots=True, style='b', figsize=(8, 5), grid=True)
# tag: aapl_day
# title: Apple stock tick data and volume for a trading day
# + uuid="47a27b89-f6a1-4b3c-9944-8a6109642c72"
NKE_resam = NKE.resample(rule='5min', how='mean')
np.round(NKE_resam.head(), 2)
# + uuid="7730056f-4ce8-4a23-853e-f5206eb86ea7"
NKE_resam['bid'].fillna(method='ffill').plot(grid=True)
# tag: aapl_resam
# title: Resampled Apple stock tick data
# + uuid="d8d2f963-3909-4434-95fa-69f932b49451"
def reversal(x):
return 2 * 95 - x
# + uuid="e6d72fc4-0617-480c-84a3-56892f0e4b01"
NKE_resam['bid'].fillna(method='ffill').apply(reversal).plot(grid=True)
# tag: aapl_resam_apply
# title: Resampled Apple stock tick data with function applied to it
# + uuid="ac44afa5-337a-4aed-b811-1b76db21682d"
!rm ./data/*
# Windows: del /data/*
# -
# ## Conclusions
# ## Further Reading
# <img src="http://hilpisch.com/tpq_logo.png" alt="The Python Quants" width="35%" align="right" border="0"><br>
#
# <a href="http://www.pythonquants.com" target="_blank">www.pythonquants.com</a> | <a href="http://twitter.com/dyjh" target="_blank">@dyjh</a>
#
# <a href="mailto:<EMAIL>"><EMAIL></a>
#
# **Python Quant Platform** |
# <a href="http://oreilly.quant-platform.com">http://oreilly.quant-platform.com</a>
#
# **Derivatives Analytics with Python** |
# <a href="http://www.derivatives-analytics-with-python.com" target="_blank">Derivatives Analytics @ Wiley Finance</a>
#
# **Python for Finance** |
# <a href="http://shop.oreilly.com/product/0636920032441.do" target="_blank">Python for Finance @ O'Reilly</a>
| ipython/06_Financial_Time_Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:gmaps] *
# language: python
# name: conda-env-gmaps-py
# ---
# Dependencies
import pandas as pd
from bs4 import BeautifulSoup
import requests
import pymongo
from splinter import Browser
from splinter.exceptions import ElementDoesNotExist
# Initialize PyMongo to work with MongoDBs
conn = 'mongodb://localhost:27017'
client = pymongo.MongoClient(conn)
# Define database and collection
db = client.mars_db
collection = db.articles
# +
# URL of page to be scraped
url = 'https://mars.nasa.gov/news/'
# Retrieve page with the requests module
response = requests.get(url)
# Create BeautifulSoup object; parse with 'html5lib'
soup = BeautifulSoup(response.text, 'html5lib')
# -
| mission_to_mars_old.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solver Bare Bones IQ
# **author**: <NAME> et <NAME>
#
# **date**: January 30 2019
#
# We will solve the Hubbard Model at half-filling, using the known Hartee approximation for the Green Function.
#
# $$ H = \sum_{ij} t_{ ij} c^\dagger_i c_j + U \sum_i n_{i \uparrow} n_{i \downarrow} $$
# ## Mathematical Description
#
# The project here is to build a dynamical mean field theory solver. Extensive references exists, a good starting point being: [DMFT, A. Georges](https://journals.aps.org/rmp/abstract/10.1103/RevModPhys.68.13).
#
# The basic equations of DMFT are the following (we will stick to two dimensions):
#
# ### Math description of DMFT
#
# \begin{align}
# G_{imp}(z) & = \int \frac{dk}{(2 \pi)^2} G_{L}(z, k) \\
# G_{L}(z, k) & = \bigg[z - \epsilon(k) - \Sigma(z) \bigg]^{-1} \\
# G_{imp}(z) & = \bigg[z - \Delta(z) - \Sigma(z) \bigg]^{-1}
# \end{align}
#
#
# ### In words description of DMFT
# The last three equations are iterated until selfconsistency, let us explain in a bit more detail.
#
# #### The impurity solver
#
# | Input | output |
# |---------------------------------------|------------------------------------------|
# | Hybridization function := $\Delta(z)$ | Impurity Green function := $G_{imp}(z)$ |
#
#
# The Impurity solver's tasks is to compute the Impurity green function $G_{imp}(z)$, with z being a frequency, here a complex frequency, namely a matsubara frequency : $z := i \omega_n = (2n + 1)\frac{\pi}{\beta} $ .
# This can be done in mutiple ways, see the above reference.
#
#
# #### The selfconsistency scheme
#
# Once the calculation of the impurity green function is done, one must update the hybridization function by using the above equations. Once the new hybridization function has been calculated, we go back to solving the impurity once more. If this is a bit mysterious, don't worry too much, it will get clearer as we progress along.
#
#
# ### Picture description of DMFT
#
# 
#
# First let us calculate the first moment of the hybridization function;
# A high frequency expansion of the above equations yields:
#
# \begin{align}
# \Delta(z) & \lim_{z -> \infty} = \frac{\Delta_1}{z} \\
# \Delta_1 & = \int \frac{dk}{(2 \pi)^2} [\epsilon(k)]^2 - \Big[ \int \frac{dk}{(2 \pi)^2} \epsilon(k) \Big]^2 \\
# \end{align}
#
# Calculating this high-frequency is important will introduce many concepts
# ## Introduction to python and functions
import numpy as np
from scipy.integrate import dblquad
def eps_k(kx: float, ky:float )-> float:
"""The dispersion relation.
Parameters
----------
kx : float
The x component of the wave-vector k.
ky : float
The y component of the wave-vector k.
Returns
-------
float
The value of the dispersion relation.
"""
return -2.0*(np.cos(kx) + np.cos(ky))
def get_hyb_fm()-> float:
# define the limits of the brillouin zone in kx
kx_limit_low = -np.pi
kx_limit_high = np.pi
# the limits of the brillouin zone in ky need to be function, here inline functions without name,
# also called lambdas
ky_limit_low = lambda kx: -np.pi
ky_limit_high = lambda kx: np.pi
# dblquad returns a tuple with two elements (result: float, error: float),
# Here, we grab the first element, by indexing with "[0]" to get the result.
int_eps_k = dblquad(eps_k, kx_limit_low, kx_limit_high, ky_limit_low, ky_limit_high)[0]
eps_k_squared = lambda kx, ky: eps_k(kx, ky) * eps_k(kx, ky)
int_eps_k_squared = dblquad(eps_k_squared, kx_limit_low, kx_limit_high, ky_limit_low, ky_limit_high)[0]
return 1.0/(2.0*np.pi)**2.0*(int_eps_k_squared - int_eps_k)
print("Value of the Hybridization first moment = ", get_hyb_fm())
# ## Introduction to classes
# +
# Let us keep the previous elements and define a model class that will be
class Model:
def __init__(self, t: float, beta: float, U: float, hyb):
self.t = t # hopping, usaully set to -1.0
self.beta = beta # 1/Temperature
self.U = U # Hubbard interaction
self.hyb = hyb # hybridization
def eps_k(self, kx: float, ky:float) -> float:
return -2.0*self.t*(np.cos(kx) + np.cos(ky))
# +
class ImpuritySolver:
def __init__(self, model: Model):
self.model = model
def solve(self):
green_impurity = np.zeros(self.model.hyb.shape, dtype=complex)
# We know the result for the Hartee part of the self-energy:
# Sigma(z) = U/2.0
self_energy = self.model.U / 2.0 *np.ones(green_impurity.shape, dtype=complex)
for n in range(green_impurity.shape[0]):
iwn = 1.0j * (2.0 * n + 1.0 ) * np.pi / self.model.beta
# we index the numpy array with the "[]" operator. If the array was two dimensional,
# we could index with two variables.
green_impurity[n] = 1.0/(iwn - self.model.hyb[n] - self_energy[n])
return green_impurity
# -
class SelfConsistency:
def __init__(self, model, green_impurity):
self.model = model
self.green_impurity = green_impurity
def green_lattice_scalar_real(self, kx: float, ky:float, self_energy, n: int)->float:
""" Return the real part of the lattice green function for one frequency (scalar, not array),
given by the index n.
"""
iwn = 1.0j * (2.0 * n + 1.0) * np.pi / self.model.beta
return np.real(1.0/(iwn - self.model.eps_k(kx, ky) - self_energy[n]))
def green_lattice_scalar_imag(self, kx:float, ky:float, self_energy, n:int)->float:
""" Return the imaginary part of the lattice green function for one frequency (scalar, not array),
given by the index n.
"""
iwn = 1.0j * (2.0 * n + 1.0) * np.pi / self.model.beta
return np.imag(1.0/(iwn - self.model.eps_k(kx, ky) - self_energy[n]))
def run_selfconsistency(self):
# 0.) extract the self-energy from the green_impurity
self_energy = np.zeros(self.green_impurity.shape, dtype=complex)
for n in range(self.green_impurity.shape[0]):
iwn = 1.0j * (2.0 * n + 1.0 ) * np.pi / self.model.beta
self_energy[n] = iwn - self.model.hyb[n] - 1.0/self.green_impurity[n]
# 1.) Calculate a new green_impurity using the lattice green function
green_impurity_new = np.zeros(self.green_impurity.shape, dtype=complex)
kx_limit_low = -np.pi
kx_limit_high = np.pi
ky_limit_low = lambda kx: -np.pi
ky_limit_high = lambda kx: np.pi
for n in range(green_impurity_new.shape[0]):
green_impurity_new[n] = dblquad(self.green_lattice_scalar_real, kx_limit_low, kx_limit_high,
ky_limit_low, ky_limit_high, args=(self_energy, n))[0]
green_impurity_new[n] += 1.0j*dblquad(self.green_lattice_scalar_imag, kx_limit_low, kx_limit_high,
ky_limit_low, ky_limit_high, args=(self_energy, n))[0]
green_impurity_new[n] /= (2.0*np.pi)**2.0
# 2.) extract new hyb
hyb_new = np.zeros(self.green_impurity.shape, dtype=complex)
for n in range(hyb_new.shape[0]):
iwn = 1.0j * (2.0 * n + 1.0 ) * np.pi / self.model.beta
hyb_new[n] = iwn - 1.0/green_impurity_new[n] - self_energy[n]
return hyb_new
def main():
# 0.) define the simulation parameters
n_freq: int = 200 # The number of frequencies
t = -1.0
beta = 10.0
U = 0.0
iter_max = 5 # maximum number of iterations
# initial guess for the hybridization, hmmm. No clue so put it to zero !
# (Would be better to set the hyb to the first moment calculated above).
hyb = np.zeros(n_freq, dtype=complex)
for ii in range(iter_max):
model = Model(t, beta, U, hyb)
impurity_solver = ImpuritySolver(model)
green_impurity = impurity_solver.solve()
selfconsistency = SelfConsistency(model, green_impurity)
hyb = selfconsistency.run_selfconsistency()
print(".", end="")
frequencies = np.array([(2.0*n + 1)*np.pi/beta for n in range(n_freq)])
return (frequencies, hyb)
frequencies, hyb = main()
# +
import matplotlib.pyplot as plt
# %matplotlib inline
plt.plot(frequencies, hyb.imag, 'b*', label="hyb", markersize=5)
plt.plot(frequencies, -4.0/frequencies, 'r-', label="hyb_fm" , linewidth=3)
plt.legend()
plt.title("Imaginary part of Hybridization and the first moment.")
# plt.xlim(80, 120)
plt.ylim(-2.0, 0.1);
# -
| solver_iq.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Harmonising Landsat and sentinel using DE Africa Coeefficients
#
# * **Products used:**
#
# + raw_mimetype="text/restructuredtext" active=""
# **Keywords**: :index:`data used; landsat 8`, :index:`analysis; change detection`, :index:`band index; NDVI`, :index:`band index; EVI`, :index:`forestry`
# -
# ### Background
#
# ### Description
#
#
# ***
# ## Getting started
#
# ### Load packages
# Load key Python packages and any supporting functions for the analysis.
# +
import datacube
import datacube.utils.rio
import matplotlib.pyplot as plt
import numpy as np
import seaborn as sns
from scipy import stats
import xarray as xr
import geopandas as gpd
from deafrica_tools.datahandling import load_ard
from deafrica_tools.plotting import display_map, rgb
from deafrica_tools.bandindices import calculate_indices
from deafrica_tools.dask import create_local_dask_cluster
from deafrica_tools.classification import collect_training_data
#This will speed up loading data
datacube.utils.rio.set_default_rio_config(aws='auto', cloud_defaults=True)
# -
create_local_dask_cluster()
# ### Connect to the datacube
# Activate the datacube database, which provides functionality for loading and displaying stored Earth observation data.
dc = datacube.Datacube(app="HLS")
# + [markdown] tags=[]
# ### Analysis parameters
#
# +
# Define the area of interest
# Northern Nigeria
img_name = 'nigeria'
latitude = 12.24142
longitude = 9.33816
crop_mask = 'crop_mask_western'
#Southern Nigeia
# latitude = 8.05477
# longitude = 11.01148
# #Northern Ghana
# img_name = 'northern_ghana'
# latitude = 10.15034
# longitude = 0.34555
# crop_mask = 'crop_mask_western'
#Southen Ghana
# latitude = 6.05653
# longitude = 0.50653
#western ivory coast
# latitude = 9.52254
# longitude = -5.57921
# Guinea Part
# latitude = 10.75481
# longitude = -11.05431
#Guinea Southern
# latitude = 9.45168
# longitude = -13.46149
# Ethiopia
# latitude = 9.70906
# longitude = 38.62793
#Egypt
# img_name = 'egypt'
# latitude= 29.2401
# longitude=30.7371
# crop_mask = 'crop_mask_northern'
# #kenya
# img_name = 'kenya'
latitude = -4.39023
longitude = 33.09082
crop_mask = 'crop_mask_eastern'
buffer = 0.01
# crop_mask = 'crop_mask_western'
# Combine central lat,lon with buffer to get area of interest
lat_range = (latitude-buffer, latitude+buffer)
lon_range = (longitude-buffer, longitude+buffer)
# Set the range of dates for the complete sample
time = ('2019')
dask_chunks = dict(x=1000, y=1000)
# + [markdown] tags=[]
# ## View the selected location
# The next cell will display the selected area on an interactive map.
# The red border represents the area of interest of the study.
# Zoom in and out to get a better understanding of the area of interest.
# Clicking anywhere on the map will reveal the latitude and longitude coordinates of the clicked point.
# +
# display_map(x=lon_range, y=lat_range)
# -
# ## Load and view Landsat data
#Create a query object
query = {
'x': lon_range,
'y': lat_range,
'time': time,
'measurements': ['red','nir'],
'resolution': (-30, 30),
'group_by': 'solar_day',
'output_crs': 'EPSG:6933'
}
# load cloud-masked ls8 using load_ard
ds = load_ard(dc=dc,
products=['ls8_sr'],
min_gooddata=0.95,
mask_filters=(['opening',5], ['dilation',5]),
dask_chunks=dask_chunks,
**query,
)
# +
# load cloud-masked fractional cover using load_ard
ds_sentinel = load_ard(dc=dc,
products=['s2_l2a'],
like=ds.geobox,
time=time,
measurements=['red','nir_2'], #use nir narrow to match with LS8
min_gooddata=0.95,
mask_filters=(['opening',5], ['dilation',5]),
dask_chunks=dask_chunks,
)
#rename nir2 to trick calculate_indices
ds_sentinel = ds_sentinel.rename({'nir_2':'nir'})
# +
cm = dc.load(product=crop_mask,
time=('2019'),
measurements='mask',
resampling='nearest',
like=ds.geobox).mask.squeeze()
cm.plot.imshow(add_colorbar=False, figsize=(5,5))
plt.title('Cropland Extent');
# -
# ### Mask non-croplands
ds = ds.where(cm, np.nan)
ds_sentinel = ds_sentinel.where(cm, np.nan)
ds = calculate_indices(ds, 'NDVI', collection='c2', drop=False)
ds_sentinel = calculate_indices(ds_sentinel, 'NDVI', collection='s2', drop=False)
# ## Match Sentinel-2 and Landsat acquistion times
#
# Tolerance of 3 days
#
ds_sentinel = ds_sentinel.reindex(time=ds.time, method='nearest', tolerance='3D')
# + [markdown] tags=[]
# ### Harmonize Landsat 8 with Sentinel-2 using DE Africa Coefficents
# + tags=[]
# multiplying the red band with slope and adding to intercept
slope = 0.979
intercept = 0.012
y_dea = (ds_sentinel['NDVI'] - intercept) / slope
# -
# ### Reduce to 1D for time-series plotting
ndvi_ls = ds.NDVI.mean(dim=['x','y']).compute()
ndvi_s2 = ds_sentinel.NDVI.mean(dim=['x','y']).compute()
ndvi_s2_adj = y_dea.mean(dim=['x','y']).compute()
# ### time-series plot
# +
fig, ax = plt.subplots(2,1, sharex=True, figsize=(14, 6))
ndvi_ls.plot(marker='.', lw=0, color='red', ms=10, label='Landsat 8', ax=ax[0])
ndvi_s2.plot(marker='*', lw=0, ms=10, color='green', label='Sentinel 2', ax=ax[0])
ax[0].title.set_text('Before Band Adjustment')
ax[0].set_xlabel('x-label', fontsize=0)
ax[0].grid()
ax[0].legend()
ndvi_ls.plot(marker='.', lw=0, color='red', ms=10, label='Landsat 8', ax=ax[1])
ndvi_s2_adj.plot(marker='*', lw=0, ms=10, color='green', label='Adj Sentinel 2', ax=ax[1])
ax[1].legend()
ax[1].title.set_text('After Band Adjustment')
ax[1].set_xlabel('Time')
ax[1].grid()
plt.show()
# -
| testing/6_HLS_timeseries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # MLOps with Seldon and Jenkins Classic
#
# This repository shows how you can build a Jenkins Classic pipeline to enable Continuous Integration and Continuous Delivery (CI/CD) on your Machine Learning models leveraging Seldon for deployment.
# This CI/CD pipeline will allow you to:
#
# - Run unit tests using Jenkins Classic.
# - Run end-to-end tests for your model with KIND (Kubernetes in Docker).
# - Promote your model as a across multiple (staging / prod) environments.
#
# To showcase these features we will implement add continuous integration and delivery to three different models.
# You can find these under the `/models` folder.
# As we shall see, each of them will require a [different approach to deployment](#Use-Cases).
# ## CI/CD Pipeline
#
# The diagram below provides a high level overview of the CI/CD pipeline.
# It includes an overview of all the different types of repositories, together with the stakeholders that are the primary contributors of each, as well as the Kubernetes environments in which the applications are deployed.
#
# The key pieces to note on the diagram are:
#
# - There are different types of environments with different restrictions and behaviours, e.g. staging and production.
# - It’s possible to have more than one environment for each type (as the type is just what would give it a specific type of config/behaviour).
# - The environments are by default in the same cluster (as namespaces), however it’s also possible to configure them across different clusters.
# - Each of the green boxes is a single repository, but it can also have a mono-repo approach, whereby each of the white boxes is a folder within a repo.
# 
# ### Model implementation repository
#
# From a high-level point of view, when a model implementation repository is updated by a Data Scientist or ML Engineer, the Jenkins CI will push changes to the [GitOps repository](#gitops-repository). This enables the following workflow:
#
# 1. A Data Scientist or ML Engineer trains a new model.
# 2. The Data Scientist or ML Engineer pushes the updated configuration to the model implementation repository.
# 3. The CI tool automatically builds and tests the model implementation.
# 4. The CI tool automatically pushes the change into the GitOps staging repository.
# 5. The CI tool automatically opens a PR into the GitOps production repository.
#
# One key point to highlight which may not be obvious by just looking at the diagram is that in this phase of model implementation, the example above showcases how we can leverage a re-usable model server - that is, reusing a pre-built docker image instead of building one every time.
# If there are more custom requirements, the user is in full control of the steps performed by the CI Platform Jenkins.
# This means that it is also possible to build s2i wrapped components which may require training the image every time.
#
# To gain a better understanding of how the CI/CD pipeline is implemented on each model implementation repository you can check the documented [deep dive](#diving-into-our-cicd-pipeline).
#
# #### Why a new repo for every model?
#
# A new model implementation repo is currently created because it provides us with a way to separate the “Model Deployment” phase and the “Model Training/Experimentation” phase, and allows us to use the repo as the integration between any frameworks that can serve as sources of models (MLFlow, Kubeflow, Spark, etc).
# The repo is able to store any metadata, IDs, and configuration files required, and is processed through the CI pipeline every time it is modified.
#
# #### Building a docker image in model implementation repository
#
# Whilst most of the times users of this approach will be leveraging re-usable model servers such as the SKLearn model server, it is also possible to build a docker image every single time (i.e. build a non-reusable model every time a model changes).
# This can be be done by adding the relevant steps which would most often include the s2i utility.
# This may be desired if there are non-standard linux libraries or non-standard depdencies that need to be re-installed every time.
# ### GitOps repository
#
# The state of each of our environments (e.g. production or staging) is stored on a GitOps repository.
# This repository contains all the different Kubernetes resources that have been deployed to each cluster.
# It is linked through [ArgoCD](#ArgoCD) to each of our Kubernetes clusters (or namespaces) so that a change in the repository triggers an update of our environment.
#
# When the deployment configuration of a machine learning model implementation is updated, this will automatically make the changes available through a PR to the respective manager/tech-lead/approver.
# This step will enable the end to end machine learning model promotion to be reviewed and approved by the respective individual.
#
# The manager/tech-lead will have to approve the PR before it can be merged.
# Once it’s approved, it will be merged into the GitOps repo, which will immediately trigger the update in the production namespace/cluster.
#
# You can see an example of a GitOps repository in the [SeldonIO/seldon-gitops](https://github.com/SeldonIO/seldon-gitops) repository.
# #### Re-usable model server repository
#
# If there is a need for a new reusable model server, then it’s possible to do so by creating a repository which would follow a different path.
# This would be different to the model implementation repository as it would only be built once in a while, whilst the model server would be built multiple times.
# ### Set up
#
# As a pre-requisite you need to ensure that have access to a Kubernetes cluster.
# In particular, this guide requires the following pre-requisites:
#
# - A Kubernetes cluster running v1.13+.
# - Jenkins Classic installed in your cluster. You can find instructions on how to install and configure it on the [Installing Jenkins on your K8s cluster](#Installing-Jenkins-on-your-K8s-cluster) section.
# - Seldon Core v0.5.1 installed in your cluster.
# + [markdown] toc-hr-collapsed=true toc-nb-collapsed=true
# ### Use cases
#
# This guide goes through three different methods to build and deploy your model.
# Each of these can be found under the `./models/` of this repository.
#
# - Using Seldon pre-built re-usable model servers (`./models/news_classifier`).
# - Using custom re-usable servers (`./models/images_classifier`).
# - Using custom servers with an embedded model.
# -
# ## Diving into our CI/CD Pipeline
#
# On this section we will dive into the internals of the CI/CD pipeline for our [model implementation repositories](#Model-implementation-repository).
# This includes a detailed description of the `Jenkinsfile`, as well as a look into our suggested testing methodology.
#
# Note that this will cover a generic example.
# However, as we shall see, specialising this approach into any of our [three main use cases](#Use-cases) will be straightforward.
#
# We leverage [Jenkins Pipelines](https://jenkins.io/doc/book/pipeline/) in order to run our continuous integration and delivery automation.
# From a high-level point of view, the pipeline configuration will be responsible for:
#
# - Define a **replicable** test and build environment.
# - Run the unit and integration tests (if applicable).
# - Promote the application into our staging and production environments.
#
# We can see a `Jenkinsfile` below taken from the `./models/news_classifier` example.
# This `Jenkinsfile` defines a pipeline which takes into account all of the points mentioned above.
# The following sections will dive into each of the sections in a much higher detail.
# %%writefile ./models/news_classifier/Jenkinsfile
pipeline {
agent {
kubernetes {
defaultContainer 'core-builder'
yamlFile 'models/news_classifier/podTemplate.yaml'
}
}
stages {
stage('Test') {
steps {
sh '''
cd models/news_classifier
make install_dev test
'''
}
}
stage('Test integration') {
steps {
sh '''
cd models/news_classifier
./integration/kind_test_all.sh
'''
}
}
stage('Promote application') {
steps {
withCredentials([[$class: 'UsernamePasswordMultiBinding',
credentialsId: 'github-access',
usernameVariable: 'GIT_USERNAME',
passwordVariable: '<PASSWORD>']]) {
sh '''
cd models/news_classifier
./promote_application.sh
'''
}
}
}
}
}
# %%writefile ./models/news_classifier/podTemplate.yaml
apiVersion: v1
kind: Pod
metadata:
name: test-pod
spec:
containers:
- name: core-builder
image: seldonio/core-builder:0.8
resources:
limits:
cpu: 500m
memory: 1500Mi
ephemeral-storage: "15Gi"
requests:
cpu: 200m
memory: 1500Mi
ephemeral-storage: "15Gi"
securityContext:
privileged: true
tty: true
volumeMounts:
- mountPath: /lib/modules
name: modules
readOnly: true
- mountPath: /sys/fs/cgroup
name: cgroup
- mountPath: /var/lib/docker
name: dind-storage
volumes:
- name: modules
hostPath:
path: /lib/modules
- name: cgroup
hostPath:
path: /sys/fs/cgroup
- name: dind-storage
emptyDir: {}
# ### Replicable test and build environment
#
# In order to ensure that our test environments are versioned and replicable, we make use of the [Jenkins Kubernetes plugin](https://github.com/jenkinsci/kubernetes-plugin).
# This will allow us to create a Docker image with all the necessary tools for testing and building our models.
# Using this image, we will then spin up a separate pod, where all our build instructions will be ran.
# We will use the `podTemplate()` object in the Jenkins Pipeline configuration to define the requirements of this pod
#
# Since it leverages Kubernetes underneath, this also ensure that our CI/CD pipelines are easily scalable.
# ### Integration tests
#
# Now that we have a model that we want to be able to deploy, we want to make sure that we run end-to-end tests on that model to make sure everything works as expected.
# For this we will leverage the same framework that the Kubernetes team uses to test Kubernetes itself: [KIND](https://kind.sigs.k8s.io/).
#
# KIND stands for Kubernetes-in-Docker, and is used to isolate a Kubernetes environent for end-to-end tests.
# In our case, we will use this isolated environment to test our model.
#
# The steps we'll have to carry out include:
#
# 1. Enable Docker within your CI/CD pod.
# 2. Add an integration test stage.
# 3. Leverage the `kind_test_all.sh` script that creates a KIND cluster and runs the tests.
#
# #### Add integration stage to Jenkins
#
# We can leverage Jenkins Pipelines to manage the different stages of our CI/CD pipeline.
# In particular, to add an integration stage, we can use the `stage()` object:
#
# ```groovy
# stage('Test integration') {
# steps {
# sh '''
# cd models/news_classifier
# ./integration/kind_test_all.sh
# '''
# }
# }
# ```
# #### Enable Docker
#
# To test our models, we will need to build their respective containers, for which we will need Docker.
# In order to do so, we will first need to mount a few volumes into the CI/CD container.
# These basically consist of the core components that docker will need to be able to run.
# To mount them we will add these entries into the `podTemplate.yaml` file.
#
# Please also note that we set container to run in `privileged` mode.
#
#
# ```yaml
# ApiVersion: v1
# ...
# spec:
# containers:
# - name: core-builder
# ...
# securityContext:
# privileged: true
# ...
# volumeMounts:
# - mountPath: /lib/modules
# name: modules
# readOnly: true
# - mountPath: /sys/fs/cgroup
# name: cgroup
# - mountPath: /var/lib/docker
# name: dind-storage
# volumes:
# - name: modules
# hostPath:
# path: /lib/modules
# - name: cgroup
# hostPath:
# path: /sys/fs/cgroup
# - name: dind-storage
# emptyDir: {}
# ```
# #### Run tests in Kind
#
# The `kind_run_all.sh` may seem complicated at first, but it's actually quite simple.
# All the script does is set-up a kind cluster with all dependencies, deploy the model and clean everything up.
# Let's break down each of the components within the script.
# We first start the docker daemon and wait until Docker is running (using `docker ps q` for guidance.
#
# ```bash
# ## FIRST WE START THE DOCKER DAEMON
# service docker start
# ## the service can be started but the docker socket not ready, wait for ready
# WAIT_N=0
# while true; do
# # docker ps -q should only work if the daemon is ready
# docker ps -q > /dev/null 2>&1 && break
# if [[ ${WAIT_N} -lt 5 ]]; then
# WAIT_N=$((WAIT_N+1))
# echo "[SETUP] Waiting for Docker to be ready, sleeping for ${WAIT_N} seconds ..."
# sleep ${WAIT_N}
# else
# echo "[SETUP] Reached maximum attempts, not waiting any longer ..."
# break
# fi
# done
# ```
# Once we're running a docker daemon, we can run the command to create our KIND cluster, and install all the components.
# This will set up a Kubernetes cluster using the docker daemon (using containers as Nodes), and then install Ambassador + Seldon Core.
#
#
# ```bash
# ########################################
# ## AVOID EXIT ON ERROR FOR FOLLOWING CMDS
# set +o errexit
#
# ## START CLUSTER
# make kind_create_cluster
# KIND_EXIT_VALUE=$?
#
# ## Ensure we reach the kubeconfig path
# export KUBECONFIG=$(kind get kubeconfig-path)
#
# ## ONLY RUN THE FOLLOWING IF SUCCESS
# if [[ ${KIND_EXIT_VALUE} -eq 0 ]]; then
# # KIND CLUSTER SETUP
# make kind_setup
# SETUP_EXIT_VALUE=$?
# ```
# We can now run the tests; for this we run all the dev installations and kick off our tests (which we'll add inside of the integration folder).
#
# ```bash
# # BUILD S2I BASE IMAGES
# make build
# S2I_EXIT_VALUE=$?
#
# ## INSTALL ALL REQUIRED DEPENDENCIES
# make install_integration_dev
# INSTALL_EXIT_VALUE=$?
#
# ## RUNNING TESTS AND CAPTURING ERROR
# make test
# TEST_EXIT_VALUE=$?
# fi
# ```
#
# Finally we just clean everything, including the cluster, the containers and the docker daemon.
#
# ```bash
# ## DELETE KIND CLUSTER
# make kind_delete_cluster
# DELETE_EXIT_VALUE=$?
#
# ########################################
# ## EXIT STOPS COMMANDS FROM HERE ONWARDS
# set -o errexit
#
# ## CLEANING DOCKER
# docker ps -aq | xargs -r docker rm -f || true
# service docker stop || true
# ```
# ### Promote your application
#
# After running our integration tests, the last step is to promote our model to our staging and production environments.
# For that, we will leverage our [GitOps repository](#GitOps-repository) where the state of each environment is stored.
#
# In particular, we will:
#
# - Push a change to the staging GitOps repository, which will update the staging environment instantly.
# - Submit a PR to the production GitOps repository, which will wait for a Tech Lead / Manager approval.
#
# This will be handled by the `promote_application.sh` script, which can be seen below.
# +
# %%writefile ./models/news_classifier/promote_application.sh
# ##!/bin/bash
## ENSURE WE ARE IN THE DIR OF SCRIPT
# cd -P -- "$(dirname -- "$0")"
## SO WE CAN MOVE RELATIVE TO THE ACTUAL BASE DIR
export GITOPS_REPO="seldon-gitops"
export GITOPS_ORG="adriangonz"
export STAGING_FOLDER="staging"
export PROD_FOLDER="production"
## This is the user that is going to be assigned to PRs
export GIT_MANAGER="adriangonz"
export UUID=$(cat /proc/sys/kernel/random/uuid)
git clone https://${GIT_USERNAME}:${GIT_PASSWORD}@github.com/${GITOPS_ORG}/${GITOPS_REPO}
# cd ${GITOPS_REPO}
# cp -r ../charts/* ${STAGING_FOLDER}/.
# ls ${STAGING_FOLDER}
## Check if any modifications identified
git add -N ${STAGING_FOLDER}/
git --no-pager diff --exit-code --name-only origin/master ${STAGING_FOLDER}
# STAGING_MODIFIED=$?
if [[ $STAGING_MODIFIED -eq 0 ]]; then
echo "Staging env not modified"
exit 0
fi
## Adding changes to staging repo automatically
git add ${STAGING_FOLDER}/
git commit -m '{"Action":"Deployment created","Message":"","Author":"","Email":""}'
git push https://${GIT_USERNAME}:${GIT_PASSWORD}@github.com/${GITOPS_ORG}/${GITOPS_REPO}
## Add PR to prod
# cp -r ../charts/* production/.
## Create branch and push
git checkout -b ${UUID}
git add ${PROD_FOLDER}/
git commit -m '{"Action":"Moving deployment to production repo","Message":"","Author":"","Email":""}'
git push https://${GIT_USERNAME}:${GIT_PASSWORD}@github.com/${GITOPS_ORG}/${GITOPS_REPO} ${UUID}
## Create pull request
export PR_RESULT=$(curl \
-u ${GIT_USERNAME}:${GIT_PASSWORD} \
-v -H "Content-Type: application/json" \
-X POST -d "{\"title\": \"SeldonDeployment Model Promotion Request - UUID: ${UUID}\", \"body\": \"This PR contains the deployment for the Seldon Deploy model and has been allocated for review and approval for relevant manager.\", \"head\": \"${UUID}\", \"base\": \"master\" }" \
https://api.github.com/repos/$GITOPS_ORG/$GITOPS_REPO/pulls)
export ISSUE_NUMBER=$(echo \
$PR_RESULT |
python -c 'import json,sys;obj=json.load(sys.stdin);print(obj["number"])')
## Assign PR to relevant user
curl \
-u ${GIT_USERNAME}:${GIT_PASSWORD} \
-v -H "Content-Type: application/json" \
-X POST -d "{\"assignees\": [\"${GIT_MANAGER}\"] }" \
https://api.github.com/repos/$GITOPS_ORG/$GITOPS_REPO/issues/$ISSUE_NUMBER
# -
# ## Creating a CI/CD pipeline
#
# In order to add a pipeline to Jenkins, you just have to go to the "Manage Jenkins" configuration dashboard, and click on "New Item" to create a new pipeline.
# 
#
# In the first menu, we'll add a name.
# For example, we can create a new pipeline with name `news_classifier`.
# We will then be able to add the specific details.
# Most of these will remain on "default", but we will need to change a couple of them to add a GitHub trigger, Docker access and to point to the right folder within the repository.
#
# Firstly, we will change the following:
#
# * GitHub hook trigger for GITScm polling.
# * Tick "This project is parameterised", and then when you see the next dialog:
# * Click on the "Add parameter" dropdown, and select "Credential Parameter".
# * This will open yet another box, where you want to provide the following details:
# * name: `docker-access`
# * Credential type "Username and Password"
# * Tick: required
# * Default value: Click on the "Add" dropdown, and then on "Jenkins provider":
# * This has opened another dialog box, where you want to add your docker credentials.
# * For this you need to make sure that the current selected option is "Username and Password".
# * There you have to enter your Docker username, and for password it's advised to use a Docker API Key.
# 
# Lastly, we will need to point to the right `Jenkinsfile`.
# Note that since we are working with a monorepository, where multiple model implementations are tracked, we will need to point our pipeline to the `./models/news_classifier` folder.
# If we were working with a single model implementation repository, we would only need to point to the global repo.
#
# * Select "Pipeline script from SCM" from dropdown.
# * Add the repository as SCM (in this case https://github.com/SeldonIO/sig-mlops-jenkins-classic/)
# * Point to the right `Jenkinsfile` under "Script Path". In this case, `models/news_classifier/Jenkinsfile`.
# * If needed, add credentials that will allow to access private repos.
# 
# ### Running pipeline
#
# In order to trigger a new build, we can do it manually by clicking on "Build with Parameters" and then on "Build" or we can just push a new change to our GitHub repo.
# This will take us to a view where we can see some details about each of the stages of the latest builds.
# 
# ## Installing Jenkins on your K8s cluster
#
# If you already have access to a cluster but which doesn't have Jenkins installed, you can do so easily using Helm.
# In particular, you will need to run the following:
# + language="bash"
# helm install \
# --name "jenkins" stable/jenkins \
# --namespace "jenkins" \
# --set "rbac.create=true" \
# --set "master.adminUser=admin" \
# --set "master.adminPassword=<PASSWORD>" \
# --set "master.serviceType=LoadBalancer"
# -
# This will install Jenkins and all the required services in the cluster.
# To get the Load Balancer where it can be accessed you can run:
# + language="bash"
# kubectl get svc -n jenkins | grep jenkins
# -
# ### Further configuration
#
# If you wish to set up automated pipeline triggers, you will have to install the "GitHub" plugin (there are quite a few github related ones but the one you want is the one called plainly "GitHub", which then will allow for triggering pipelines automatically on commit.
#
# - Install the GitHub Plugin [(for automated webhook triggers)](https://support.cloudbees.com/hc/en-us/articles/115003015691-GitHub-Webhook-Non-Multibranch-Jobs).
# - Provide a GitHub token with read access so it can clone relevant repositories.
# - Set-up webhooks so that GitHub can send push requests.
#
# Additionally, you will need to configure your Git's `name` and `email` as part of Jenkins settings.
#
# 
# #### Make sure plugins are updated
#
# If you try to run a pipeline and you get an error such as "No Such DSL Method", or any strange Java exception when running a pipeline, the most probably reason is due to current plugins not being up to date.
#
# Updating your plugins can be done by going to "Manage Jenkins" -> "Plugins", and then selecct all the plugins and click "Update and load after restart". This will take you to another screen - there you should tick the checkbox that reads "restart after plugins are downloaded and installed".
#
# Once you update our plugins you should be ready to go.
# ## ArgoCD
#
# A key point of this approach to MLOps relies on having a GitOps repository which gets synced with our Kubernetes cluster.
# To achieve this we leverage [ArgoCD](https://argoproj.github.io/argo-cd/), which will take care of setting up webhooks with your GitOps repository so that on every change it triggers a synchronisation between the resources you've pushed and what's deployed on the cluster.
# ### Installation
#
# If you don't have it already, you can install ArgoCD following the [official documentation](https://argoproj.github.io/argo-cd/getting_started/#1-install-argo-cd):
# + language="bash"
# kubectl create namespace argocd
# kubectl apply -n argocd -f https://raw.githubusercontent.com/argoproj/argo-cd/stable/manifests/install.yaml
# -
# Additionally, you will need to install the accompanying CLI tool.
# This tool will allow you to easily link your GitOps repository taking care of the entire process.
# The instructions to install it will vary between different platforms.
# The official documentation shows the [recommended method](https://argoproj.github.io/argo-cd/cli_installation/) on each of the major ones.
# ### Setting up GitOps repository
#
# To set up the GitOps repository so that it's tracked by ArgoCD we will use the `argocd` CLI tool.
# We will assume that the `GITHUB_ORG` and `REPONAME` environment variables have been created and that the repository has already been created and can be found in the `https://github.com/$GITHUB_ORG/$REPONAME` url.
# + language="bash"
# export GITHUB_ORG=SeldonIO
# export REPONAME=seldon-gitops
# -
# #### Private repositories (optional)
#
# If your repository is private, we will first need to provide the right credentials for ArgoCD to use.
# We can do so either using a [user / password login](https://argoproj.github.io/argo-cd/user-guide/private-repositories/#https-username-and-password-credential) or using [SSH keys](https://argoproj.github.io/argo-cd/user-guide/private-repositories/#tls-client-certificates-for-https-repositories).
# Note that, for the former, we can also use a [personal access token](https://help.github.com/en/github/authenticating-to-github/creating-a-personal-access-token-for-the-command-line) instead of the password.
#
# As an example, we will add our GitOps repository using a personal access token.
# We will assume that the environment variables `GITHUB_USER` and `GITHUB_TOKEN` are set.
# + language="bash"
# export GITHUB_USER=john.doe
# export GITHUB_TOKEN=<PASSWORD>
#
# argocd repo add https://github.com/$GITHUB_ORG/$REPONAME --username $GITHUB_USER --password $<PASSWORD>
# -
# #### Create ArgoCD projects
#
# The next step is to create two projects within ArgoCD to manage the staging and production environments respectively.
# Each of them will be linked to a folder within our GitOps repository.
# + language="bash"
# argocd app create seldon-staging \
# --repo https://github.com/$GITHUB_ORG/$REPONAME \
# --path staging \
# --dest-namespace staging
# argocd app create seldon-production \
# --repo https://github.com/$GITHUB_ORG/$REPONAME \
# --path production \
# --dest-namespace production
# -
# Note that we could also sync our `staging` and `production` environment differently.
# For example, we could have them on separate repositories or separate branches.
# In this case we would also need to update the `promote_application.sh` script so that it knows how it should promote the respective model between environments.
| examples/cicd/sig-mlops-jenkins-classic/models/jenkins_classic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Debugging retrofitting code
#
# We're trying to figure out why our results from retrofitting on Sampled Quora and fine-tuning on SST-2 and MR aren't showing the same gains that are in the [Retrofitting Paper](https://aclanthology.org/D19-1113.pdf). We've noticed one key difference, in the embedding distances listed in Table 4.
# - Their table shows average L2 distances between shared words as 3.2 for paraphrases and 4.2 for non-paraphrases before retrofitting, and 1.3 for paraphrases and 5.5 for non-paraphrases (after retrofitting on all three datasets).
# - Our experiments ([example run](https://wandb.ai/jack-morris/rf-bert/runs/eqa5zall?workspace=user-jxmorris12)) show embedding distances of 6.5 for paraphrases and 10.5 for non-paraphrases, both of which are greatly increasing throughout training. Why is this the case?
#
# ## This notebook
# In this notebook I want to:
# 1. Figure out how to get the 3.2 and 4.2 numbers on Quora. We should be able to reproduce this exactly using the pre-trained ELMO model.
# 2. Diagnose why these numbers are miscomputed in our setup. Are we getting the wrong representations? Or are we feeding in the wrong input? Or are we getting the words at the wrong index somehow?
# # Computing L2 distances of base ELMO between words from Quora
#
# ## Using our code
# +
# Pretend we're in the root folder of this project ('retrofitting/')
import sys
sys.path.append('..')
# 1. Get a batch of examples from Quora with shared words, etc.
from dataloaders import ParaphraseDatasetElmo
dataset = ParaphraseDatasetElmo(
'quora',
model_name='elmo', num_examples=1024,
max_length=40, stop_words_file=f'../stop_words_en.txt',
r1=0.5, seed=42, split='train'
)
from dataloaders.helpers import train_test_split
train_dataloader, test_dataloader = train_test_split(
dataset, batch_size=256,
shuffle=True, drop_last=True,
train_split=0.8
)
# +
import torch
from models import ElmoRetrofit
# 2. Load batch
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
batch = tuple(t.to(device) for t in next(iter(train_dataloader)))
# 3. Load ELMO
model = ElmoRetrofit(
num_output_representations = 1,
requires_grad=False,
elmo_dropout=0,
).to(device)
# +
# 4. Do inference
word_rep_pos_1, word_rep_pos_2, word_rep_neg_1, word_rep_neg_2 = (
model(*batch)
)
word_rep_pos_1.shape, word_rep_pos_2.shape, word_rep_neg_1.shape, word_rep_neg_2.shape
# -
def l2norm(t1: torch.Tensor, t2: torch.Tensor):
(t1 - t2).mean(1)
torch.norm(word_rep_pos_1 - word_rep_pos_2, p=2, dim=1)
torch.norm(word_rep_pos_1 - word_rep_pos_2, p=2, dim=1).mean()
torch.norm(word_rep_neg_1 - word_rep_neg_2, p=2, dim=1).mean()
# Ok, the distances with our code look like the ones in W&B (7 and 11) instead of the ones from the paper (3 and 4ish). I'm going to strip this down to just strings and the pytorch ELMO model and build back up, since I'm not sure where things are going wrong.
# ## Trying with original code
# +
from typing import List, Tuple, Set
import numpy as np
def get_stop_words(filename: str) -> Set[int]:
"""
Generates a set of token_ids given a text file of stopwords.
"""
stop_words = np.genfromtxt(filename, dtype='str')
return set(stop_words.tolist())
sw = get_stop_words('../stop_words_en.txt')
# +
from mosestokenizer import MosesTokenizer
tokenizer = MosesTokenizer('en', no_escape=True)
def get_shared_word(q1: List[str], q2: List[str]) -> str:
"""gets a shared word that's not a stopword.
takes list of tokens as input
raises StopIteration when no shared words found
"""
w1 = set([w for w in q1 if w.lower() not in sw])
w2 = set([w for w in q2 if w.lower() not in sw])
# print(w1, '//', w2)
shared_words = w1.intersection(w2)
return next(iter(shared_words))
# +
import torch
from allennlp.modules.elmo import Elmo
# ELMO 1B, 96m
# options_file = "https://s3-us-west-2.amazonaws.com/allennlp/models/elmo/2x4096_512_2048cnn_2xhighway/elmo_2x4096_512_2048cnn_2xhighway_options.json"
# weights_file = "https://s3-us-west-2.amazonaws.com/allennlp/models/elmo/2x4096_512_2048cnn_2xhighway/elmo_2x4096_512_2048cnn_2xhighway_weights.hdf5"
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
# ELMO 5.5B, 96m
options_file = "https://s3-us-west-2.amazonaws.com/allennlp/models/elmo/2x4096_512_2048cnn_2xhighway_5.5B/elmo_2x4096_512_2048cnn_2xhighway_5.5B_options.json"
weights_file = "https://s3-us-west-2.amazonaws.com/allennlp/models/elmo/2x4096_512_2048cnn_2xhighway_5.5B/elmo_2x4096_512_2048cnn_2xhighway_5.5B_weights.hdf5"
elmo_model = Elmo(options_file=options_file, weight_file=weights_file,
num_output_representations=1,
requires_grad=True, dropout=0.5,
scalar_mix_parameters=[1, 1, 1]).to(device)
elmo_model.eval()
# for param_name, param in elmo_model.named_parameters():
# print(param_name, param.requires_grad)
# +
from allennlp.modules.elmo import batch_to_ids
import pandas as pd
import torch
def batched_inference(model: torch.nn.Module, data: torch.Tensor, batch_size: int=256) -> torch.Tensor:
i = 0
output = []
while i < len(data):
data_batch = data[i:i+batch_size].to(device)
i += batch_size
output.append(model(data_batch)['elmo_representations'][0].cpu())
return torch.cat(output, dim=0)
def get_shared_word_reps(paraphrases, lower=False) -> Tuple[torch.Tensor, torch.Tensor]:
paraphrases_and_words = []
for ex in paraphrases:
q1 = ex['questions']['text'][0]
q2 = ex['questions']['text'][1]
if lower:
q1 = q1.lower()
q2 = q2.lower()
q1 = tokenizer(q1)
q2 = tokenizer(q2)
try:
shared_word = get_shared_word(q1, q2)
i1 = q1.index(shared_word)
i2 = q2.index(shared_word)
paraphrases_and_words.append( (q1, q2, shared_word, i1, i2) )
except StopIteration:
continue
p_df = pd.DataFrame(paraphrases_and_words, columns=['q1_tokens', 'q2_tokens', 'shared_word', 'i1', 'i2'])
p_df.head()
# paraphrase IDs
p_q1_ids = batch_to_ids(p_df['q1_tokens'])
p_q2_ids = batch_to_ids(p_df['q2_tokens'])
# using jack scudder's trick here. should be a way to do it with torch.select() though?
B_p = torch.arange(len(p_q1_ids))
p_q1_shared_word_ids = p_q1_ids[B_p, p_df['i1']]
p_q2_shared_word_ids = p_q2_ids[B_p, p_df['i2']]
# make sure shared word indices are the same - this should print True!
print(torch.all(p_q1_shared_word_ids == p_q2_shared_word_ids, 0).all())
with torch.no_grad():
# print(q1_reps.keys()) # dict_keys(['elmo_representations', 'mask'])
# print(len(q1_reps['elmo_representations'])) # list of length 1
# print(q1_reps['elmo_representations'][0].shape) # [727, 33, 1024]
B = torch.arange(len(p_q1_ids))
p_q1_reps = batched_inference(elmo_model, p_q1_ids)
p_q1_shared_word_reps = p_q1_reps[B, p_df['i1']] # [727, 1024]
p_q2_reps = batched_inference(elmo_model, p_q2_ids)
p_q2_shared_word_reps = p_q2_reps[B, p_df['i2']]
return p_q1_shared_word_reps, p_q2_shared_word_reps
# +
import datasets
d = datasets.load_dataset('quora')['train']
d = datasets.Dataset.from_dict(d[:8_000])
quora_paraphrases = d.filter(lambda ex: ex['is_duplicate'])
quora_non_paraphrases = d.filter(lambda ex: not ex['is_duplicate'])
len(quora_paraphrases), len(quora_non_paraphrases)
# -
q_p1, q_p2 = get_shared_word_reps(quora_paraphrases, lower=False)
# I want this to be 3.17
# ELMO 5.5B:
# - with uppercase letters included (lower=False): 4.122
# - without uppercase letters: 3.808
# ELMO 1B:
# - with uppercase letters included (lower=False): 6.737
# - without uppercase letters: 6.463
(q_p1 - q_p2).norm(p=2, dim=1).mean()
q_n1, q_n2 = get_shared_word_reps(quora_non_paraphrases, lower=False)
# I want this to be 4.42
# ELMO 5.5B:
# - with uppercase letters included:
# - without uppercase letters:
# ELMO 1B:
# - with uppercase letters included:
# - without uppercase letters:
(q_n1 - q_n2).norm(p=2, dim=1).mean()
pd.Series((q_n1 - q_n2).norm(p=2, dim=1).cpu()).hist(bins=35)
| notebooks/2022-03-09 BERT Embeddings test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from bokeh.sampledata.autompg import autompg as df
from bokeh.sampledata.olympics2014 import data
from bokeh.sampledata.iris import flowers
from bokeh.sampledata.project_funding import project_funding as pf
from bokeh.charts import Scatter, output_notebook, show, vplot, hplot
from bokeh.charts.operations import blend
from bokeh.charts.utils import df_from_json
import pandas as pd
# -
output_notebook()
# +
scatter0 = Scatter(
df, x='mpg', title="x='mpg'", xlabel="Miles Per Gallon")
scatter1 = Scatter(
df, x='mpg', y='hp', title="x='mpg', y='hp'",
xlabel="Miles Per Gallon", ylabel="Horsepower", legend='top_right')
scatter2 = Scatter(
df, x='mpg', y='hp', color='cyl', title="x='mpg', y='hp', color='cyl'",
xlabel="Miles Per Gallon", ylabel="Horsepower", legend='top_right')
scatter3 = Scatter(
df, x='mpg', y='hp', color='origin', title="x='mpg', y='hp', color='origin'",
xlabel="Miles Per Gallon", ylabel="Horsepower", legend='top_right')
scatter4 = Scatter(
df, x='mpg', y='hp', color='cyl', marker='origin', title="x='mpg', y='hp', color='cyl', marker='origin'",
xlabel="Miles Per Gallon", ylabel="Horsepower", legend='top_right')
# +
# Example with nested json/dict like data, which has been pre-aggregated and pivoted
df2 = df_from_json(data)
df2 = df2.sort('medals.total', ascending=False)
df2 = df2.head(10)
df2 = pd.melt(df2, id_vars=['abbr', 'name'])
scatter5 = Scatter(
df2, x='value', y='name', color='variable', title="x='value', y='name', color='variable'",
xlabel="Medals", ylabel="Top 10 Countries", legend='bottom_right')
scatter6 = Scatter(flowers, x=blend('petal_length', 'sepal_length', name='length'),
y=blend('petal_width', 'sepal_width', name='width'), color='species',
title='x=petal_length+sepal_length, y=petal_width+sepal_width, color=species',
legend='top_right')
# -
show(vplot(
hplot(scatter0, scatter1),
hplot(scatter2, scatter3),
hplot(scatter4, scatter5),
hplot(scatter6)
))
| examples/charts/notebook/scatter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Parameters
# w = 26 <br>
# k = 20 <br>
# e = 3 <br>
# p see secion
# +
import matplotlib.pyplot as plt
import pandas as pd
import os
# path to result folder containing subdirs 100, 150 and 250
path = os.path.abspath("<OUT_FOLDER>/test_out/")
tau_as_string = ["0.000000", "0.050000", "0.100000", "0.150000", "0.200000", "0.250000", "0.300000", "0.350000",
"0.400000", "0.450000", "0.500000", "0.550000", "0.600000", "0.650000", "0.700000", "0.750000",
"0.800000", "0.850000", "0.900000", "0.950000", "1.000000"]
tau_as_float = list(map(float, tau_as_string))
# -
def recall(tp):
return (float(tp) / 100000)
def precision(tp, fp):
return (float(tp) / (fp + tp))
def x_y(pattern_size, method):
assert(pattern_size in [100, 150, 250])
assert(method in ["simple", "indirect", "overlapping", "indirect_overlapping"])
tps = []
fps = []
for tau in tau_as_string:
filename = os.path.join(path, "positive", str(pattern_size), "result_{}_w26_k20_e3_tau{}.csv".format(method, tau))
df = pd.read_csv(filename)
tps.append(int(df["hits"]))
for tau in tau_as_string:
filename = os.path.join(path, "negative", str(pattern_size), "result_{}_w26_k20_e3_tau{}.csv".format(method, tau))
df = pd.read_csv(filename)
fps.append(int(df["hits"]))
recalls = [recall(x) for x in tps]
precisions = [precision(x, y) for (x, y) in zip(tps, fps)]
f1 = [2*x*y/(x+y) for (x,y) in zip(precisions, recalls)]
print("{} <- c({})".format(method, ", ".join(map(str, f1))))
return (tau_as_float, f1)
p = 100
# Simple
(x_simple, y_simple) = x_y(p, "simple")
# Indirect
(x_indirect, y_indirect) = x_y(p, "indirect")
# Overlap
(x_overlap, y_overlap) = x_y(p, "overlapping")
# Both
(x_both, y_both) = x_y(p, "indirect_overlapping")
plt.plot(x_simple, y_simple, 'b', x_indirect, y_indirect, 'g',
x_overlap, y_overlap, 'orange', x_both, y_both, 'r')
p = 150
# Simple
(x_simple, y_simple) = x_y(p, "simple")
# Indirect
(x_indirect, y_indirect) = x_y(p, "indirect")
# Overlap
(x_overlap, y_overlap) = x_y(p, "overlapping")
# Both
(x_both, y_both) = x_y(p, "indirect_overlapping")
plt.plot(x_simple, y_simple, 'b', x_indirect, y_indirect, 'g',
x_overlap, y_overlap, 'orange', x_both, y_both, 'r')
p = 250
# Simple
(x_simple, y_simple) = x_y(p, "simple")
# Indirect
(x_indirect, y_indirect) = x_y(p, "indirect")
# Overlap
(x_overlap, y_overlap) = x_y(p, "overlapping")
# Both
(x_both, y_both) = x_y(p, "indirect_overlapping")
plt.plot(x_simple, y_simple, 'b', x_indirect, y_indirect, 'g',
x_overlap, y_overlap, 'orange', x_both, y_both, 'r')
| script/jupyter_notebooks/FScore_vs_tau.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 0. Load Library
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import train_test_split
from sklearn.metrics import mean_absolute_error
from sklearn.metrics import mean_squared_error
import warnings
warnings.filterwarnings('ignore')
# -
# ## 1. Load the Datasets
df = pd.read_csv('victoria.csv')
# ## 2. Exploratory Data Analysis
df.head()
df.shape
df.describe()
df.dtypes
#Data Cleansing
df['totalprice'] = df['totalprice'].replace(r'[,]','', regex=True)
df['area'] = df['area'].replace(r'[,]','', regex=True)
#Mengubah ke data numerik
df['totalprice'] = pd.to_numeric(df['totalprice'])
df['area'] = pd.to_numeric(df['area'])
# ## 3. Data Visualization
#Univariate Analysis Price
sns.distplot(df['totalprice'])
# - Dapat dilihat bahwa harga dari apartment terpusat di kisaran harga 250000
#Mencari korelasi antar features
df.corr().style.background_gradient().set_precision(2)
# - Korelasi totalprice dan toilets cukup kuat yaitu 0.31.
# - Sedangkan korelasi totalprice dan garage yaitu 0.25.
#Bivariate Analysis antara totalprice dan toilets
sns.barplot(data=df, x=df['toilets'], y=df['totalprice'], color='blue')
# - Dapat dilihat bahwa semakin banyak toilets maka akan memberi dampak semakin mahal sebuah apartment.
#Bivariate Analysis antara totalprice dan toilets
sns.barplot(data=df, x=df['rooms'], y=df['totalprice'], color='green')
# - Semakin banyak rooms maka semakin mahal sebuah apartment
# ## 4. Feature Selection
# - Saya akan memilih variabel dengan type data numerik karena model machine learning hanya menerima type data yang berbentuk numerik.
# - Terkecuali conservation yang akan saya rubah ke dalam bentuk numerikal.
#Mengubah nilai conservation
dicti = {
'2B':3,
'3A':3,
'2A':2,
'1A':1
}
df['conservation'] = df['conservation'].replace(dicti)
new_df = df.copy()
#Menghapus kolom yang tidak perlu
drop = ['zone', 'category', 'out', 'streetcategory', 'heating']
new_df.drop(columns=drop, inplace=True)
#Membuat one hot encoding
new_df['conservation'] = pd.Categorical(new_df['conservation'])
new_df = pd.get_dummies(new_df)
# ## 5. Modelling
#Memakai Linear regression
lin_reg = LinearRegression()
#membuat variabel x dan y
x = new_df.drop(columns = ['totalprice'])
y = new_df['totalprice']
#split data
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.2)
#training model
lin_reg.fit(x_train, y_train)
#melakukan prediksi
y_pred = lin_reg.predict(x_test)
def mean_absolute_percentage_error(y_true, y_pred):
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
#check error
print(f'Nilai MAE {mean_absolute_error(y_pred, y_test)}')
print(f'Nilai MAPE {mean_absolute_percentage_error(y_test, y_pred)})')
# - Harga memiliki error sebesar 33 ribu dengan nilai MAPE sebesar 12%
| .ipynb_checkpoints/Adipta Martulandi - Victoria-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/evaneschneider/parallel-programming/blob/master/numba_intro.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={} colab_type="code" id="cBAVRqT-D8JR"
# %matplotlib inline
import matplotlib.pylab as plt
import numpy as np
import math
from numba import jit, njit, vectorize
# + [markdown] colab_type="text" id="72QKUcHITZXs"
# ## What is Numba?
# Numba is a **just-in-time**, **type-specializing**, **function compiler** for accelerating **numerically-focused** Python. That's a long list, so let's break down those terms:
#
# # + **function compiler**: Numba compiles Python functions, not entire applications, and not parts of functions. Numba does not replace your Python interpreter, but is just another Python module that can turn a function into a (usually) faster function.
# # + **type-specializing**: Numba speeds up your function by generating a specialized implementation for the specific data types you are using. Python functions are designed to operate on generic data types, which makes them very flexible, but also very slow. In practice, you only will call a function with a small number of argument types, so Numba will generate a fast implementation for each set of types.
# # + **just-in-time**: Numba translates functions when they are first called. This ensures the compiler knows what argument types you will be using. This also allows Numba to be used interactively in a Jupyter notebook just as easily as a traditional application.
# # + **numerically-focused**: Currently, Numba is focused on numerical data types, like int, float, and complex. There is very limited string processing support, and many string use cases are not going to work well on the GPU. To get best results with Numba, you will likely be using NumPy arrays.
# + [markdown] colab_type="text" id="SgFcESP_EaNv"
# ### Problem 1 - A First Numba Function
#
# **1a)** To start our exploration of Numba's features, let's write a python function to add two numbers. We'll creatively name it `add`:
# + colab={} colab_type="code" id="yzKiUuSjEeDy"
def add(x, y):
return x + y
# + [markdown] colab_type="text" id="2Fnz9vr0FG7k"
# Now, test the function, first with two scalar integers:
# + colab={} colab_type="code" id="o13KJoxJFDyD"
# %timeit add(5,3)
# + [markdown] colab_type="text" id="7HdCKleiFd6r"
# **1b)** With Numpy, we can use our function to add not just scalars, but vectors as well. Using your favorite array creation routine, create two integer arrays with ten elements each, called `a` and `b`, and use your `add` function to add them.
# + colab={} colab_type="code" id="EWwdk1dYE5pN"
a = np.arange(100)
b = np.arange(100) + 5
# + [markdown] colab_type="text" id="Jwqy1reHM0nA"
# Okay, so our function can add things. Now, let's use Numba's `jit` function to create a Numba version of our addition function:
# + colab={} colab_type="code" id="SfnuyYZ1MjxR"
numba_add = jit(add)
# + [markdown] colab_type="text" id="FW3GvGwKkawy"
# More commonly, you will use `jit` as a decorator, by adding `@jit` to the line above your function definition, but the above version shows you that at heart, `@jit` is just a python function that takes other functions as its argument!
# + [markdown] colab_type="text" id="X726qBEztHg5"
# **1c)** By default, a Numba function saves the original python version of the function in the variable `py_func`. Check that the original python version gives you the same answer as the Numba version.
# + colab={} colab_type="code" id="XS0-gUOWN12L"
numba_add.py_func(5,3)
# + [markdown] colab_type="text" id="QQx121wWGD_Y"
# #### Profiling
# A central feature of parallel programming, Numba, and writing efficient code more generally is **profiling**, or understanding how long various pieces of your program take to run. Profiling tools are becoming ever more sophisticated, but for today we're going to stick with the tried-and-true method of timing things. An easy way to do this in python is using the `%timeit` magic function. Let's try it out on our addition function:
# + colab={} colab_type="code" id="XC1o2Ki8Fx2U"
# %timeit add(1,2)
# + [markdown] colab_type="text" id="krQ0FwkjGw-_"
# What's going on here? `%timeit` is running our function many times, and then reporting the average time it takes to run. This is generally a better approach than timing a single function execution, because it accounts for random events that may cause any given run to perform poorly.
# + [markdown] colab_type="text" id="0xXjAFh-MXiv"
# **1d)** Compare the time it takes to run your function with scalar vs array arguments, then your function vs python's add function (the standard ''+'' operator).
# + colab={} colab_type="code" id="3AgB0xKOHjcS"
# %timeit add(a,b)
# + [markdown] colab_type="text" id="YsZ_clWelcVJ"
# So, scalars are faster than arrays (makes sense), and python's addition function is better than ours (seems reasonable). Now, let's see how fast our pre-compiled Numba addition function is.
# + colab={} colab_type="code" id="pqNzG9mhN-lp"
# %timeit numba_add(a,b)
# + [markdown] colab_type="text" id="pgDvbPLml3Dm"
# Hold on - our new pre-compiled function is running even slower than the original python version! What's going on here?
# + [markdown] colab_type="text" id="szNrfvJxl_Vj"
# ### Problem 2 - A Better Numba Function
#
# (This problem borrowed from [seibert's 2018 gtc numba tutorial](https://github.com/ContinuumIO/gtc2018-numba).)
#
# As we saw in the first example, Numba isn't going to speed up everything. Generally, Numba will help you most in circumstances where python's line-by-line interperability and lack of type casting is slowing it down. We can use a slightly more complicated function to demonstrate this. The following is a function to calculate the hypotenuse of two numbers, that has been carefully designed to compensate for the computer's finite precision representation of numbers (check out https://en.wikipedia.org/wiki/Hypot for more info).
# + [markdown] colab_type="text" id="gFmqy_gjvWJX"
# **2a)** Use the `@jit` decorator to generate a Numba version of this function.
# + colab={} colab_type="code" id="VNDuzJvGlkfg"
@jit(nopython=True)
def hypotenuse(x, y):
x = abs(x);
y = abs(y);
t = min(x, y);
x = max(x, y);
t = t / x;
return x * math.sqrt(1+t*t)
# + [markdown] colab_type="text" id="52cFaUWboRZO"
# **2b)** Use the `%timeit` function to determine whether the Numba version of the hyptonenuse function is better than the original Python implementation.
# + colab={} colab_type="code" id="HS7QNmoUmcYG"
# %timeit hypotenuse(3,4)
# %timeit hypotenuse.py_func(3,4)
# + [markdown] colab_type="text" id="0ZQmD-3Ho5oJ"
# **2c)** Numba functions can call other functions, provided they are also Numba functions. Below is a function that loops through two numpy arrays and puts their sum into an output array. Modify the following function to calculate the hypotenuse instead.
# + colab={} colab_type="code" id="0gO3Pxqbo4Nz"
@njit # this is an alias for @jit(nopython=True)
def ex_func(x, y, out):
for i in range(x.shape[0]):
#out[i] = x[i] + y[i] # change this line
out[i] = hypotenuse(x[i],y[i])
# + colab={} colab_type="code" id="m62ZUF84opUb"
in1 = np.arange(10, dtype=np.float64)
in2 = 2 * in1 + 1
out = np.empty_like(in1)
print('in1:', in1)
print('in2:', in2)
ex_func(in1, in2, out)
print('out:', out)
# + colab={} colab_type="code" id="q1GhkW0LBuV_"
# This test will fail until you fix the ex1 function
np.testing.assert_almost_equal(out, np.hypot(in1, in2))
# + [markdown] colab_type="text" id="zhO1-4Das9cm"
# ### Problem 3 - Fun with Fractals
# Now that we've got the basics of the Numba `jit` decorator down, let's have a little fun. A classic example problem in parallel programming is the calculation of a fractal, because a large fraction of the work can be done in parallel. Below is some code that calculates whether a number is a member of the Julia set, and then computes the set on a discrete domain to calculate a fractal.
# + [markdown] colab_type="text" id="tH6JqqMLw5Hq"
# **3a)** Modify the code below to use Numba and test how much faster it is than the original python implementation.
# + colab={} colab_type="code" id="XormIu6WqCOa"
@njit
def julia(x, y, max_iters):
"""
Given the real and imaginary parts of a complex number,
determine if it is a candidate for membership in the Julia
set given a fixed number of iterations.
"""
i = 0
c = complex(-0.8, 0.156)
a = complex(x,y)
for i in range(max_iters):
a = a*a + c
if (a.real*a.real + a.imag*a.imag) > 1000:
return 0
return 255
# + colab={} colab_type="code" id="LCr_6rlEt6KU"
@njit
def create_fractal(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
for x in range(width):
real = min_x + x * pixel_size_x
for y in range(height):
imag = min_y + y * pixel_size_y
color = julia(real, imag, iters)
image[y, x] = color
return image
# + colab={} colab_type="code" id="fZ-U8-tLuDjl"
image = np.zeros((500, 750), dtype=np.uint8)
# %timeit create_fractal(-2.0, 2.0, -1.0, 1.0, image, 200)
# + [markdown] colab_type="text" id="FI8RpWQ4uYTL"
# Want to see what you made? Run the following code to plot the image. Feel free to pick your favorite matplotlib color map :)
# + colab={} colab_type="code" id="5QY46v6Oua79"
plt.imshow(image)
plt.viridis()
plt.show()
# + [markdown] colab_type="text" id="KvUWJRTJG7r7"
# **3b)** There is more than one type of fractal in the world, however! Below is a function that determines membership in the Mandelbrot set. Modify the function below using Numba, then modify the code above to produce a new pretty picture.
# + colab={} colab_type="code" id="6Ow0_mRBHIxC"
@njit
def mandel(x, y, max_iters):
"""
Given the real and imaginary parts of a complex number,
determine if it is a candidate for membership in the Mandelbrot
set given a fixed number of iterations.
"""
i = 0
c = complex(x,y)
z = 0.0j
for i in range(max_iters):
z = z*z + c
if (z.real*z.real + z.imag*z.imag) >= 4:
return i
return 255
# + colab={} colab_type="code" id="hHu6VBXYHXxv"
@njit
def create_fractal_mandel(min_x, max_x, min_y, max_y, image, iters):
height = image.shape[0]
width = image.shape[1]
pixel_size_x = (max_x - min_x) / width
pixel_size_y = (max_y - min_y) / height
for x in range(width):
real = min_x + x * pixel_size_x
for y in range(height):
imag = min_y + y * pixel_size_y
color = mandel(real, imag, iters)
image[y, x] = color
return image
# + colab={} colab_type="code" id="dlW4gww-HftJ"
image = np.zeros((500, 750), dtype=np.uint8)
# %timeit create_fractal_mandel(-2.0, 2.0, -1.0, 1.0, image, 200)
plt.imshow(image)
plt.viridis()
plt.show()
# + [markdown] colab_type="text" id="GvUi6dVvVeY5"
# ### Problem 4 - Typing
#
# Much of the power of Numba comes from its ability to compile a specific version of a python function based on the **data types** of its arguments. The data type describes what kind of variables the function uses, and in Numpy and Numba, pre-defined type names are based on the kind and size of the number in memory. You can find the type of a variable (or array) using numpy's `dtype` object. For example, let's see what type our `a` array is.
# + colab={} colab_type="code" id="L6LBCgxiuu4Q"
a.dtype
# + [markdown] colab_type="text" id="aF8y1QTW8ZK4"
# This tells us the array contains integers, and each integer has been assigned 64bits in memory (or equivalently, 8 bytes). Most python functions are defined to work on arbitrary types, so that if you use the + operator, for example, you can add integers, floats, complex numbers, or even strings! However, this flexibility comes at a cost, performance-wise. Numba, on the other hand, compiles each function based on the types of its arguments, and infers the type of the result. You can see this if you run the `inspect_types` function on a numba function:
# + colab={} colab_type="code" id="NJoPUPFPZb3w"
numba_add.inspect_types()
# + [markdown] colab_type="text" id="D0mmp24rrKUp"
# **4a)** Numba has inferred the types for this function based on how we've used it. Try out your `numba_add` function with two floating point numbers, then re-inspect the types of the Numba function. Are they the same?
# + colab={} colab_type="code" id="Ce0oK34-rcBW"
numba_add(5., 3.)
# -
numba_add.inspect_types()
# + [markdown] colab_type="text" id="Fnuhv8VGCWFx"
# So far we have been using what Numba refers to as "lazy" (or "call-time") decoration. Basically, we've been letting Numba do the work of figuring out how we're using the function and inferring the types for us. Alternatively, if we know how we are going to use a given function, we can use "eager" (or "compile-time") decoration. To do this, we make use of the `vectorize` decorator. For example, if we want to make an integer-only version of our addition function, we could write:
# + colab={} colab_type="code" id="j5BI_s1NCVN_"
@vectorize(['int64(int64, int64)'], target='cpu')
def add_ufunc(x, y):
return x + y
# + [markdown] colab_type="text" id="Dhm4Kf5HDpJY"
# #### Numpy Universal Functions
# You'll notice a couple of new things here. In the first set of brackets, we have specified both the argument types of the function (those are inside the parentheses), as well as the return type of the function. This is just making explicit what Numba was previously inferring on our behalf. In second set of brackets you'll see that we have specified a 'target' architechture for the function. The default is `cpu`, which means that Numba is optimizing the function to your specific machine. Other options include `parallel`, which allows you to take advantage of multicore processors, and `cuda`, which we'll be discussing more tomorrow. You'll also notice that we called this a 'ufunc', which is short for [Universal Function](https://docs.scipy.org/doc/numpy/reference/ufuncs.html). In brief, universal functions are numpy functions that operate on ndarrays in element by element fashion. So if we pass two vectors to our `add_func`, they will be added together and a return vector of the same shape will be returned.
# -
c = np.arange(1000000)
d = np.arange(1000000) + 5
# + colab={} colab_type="code" id="rxiVxBslV-0z"
# %timeit add_ufunc(c,d)
# + [markdown] colab_type="text" id="oUbAJTzlFTqA"
# **4b)** Try your ufunc out with a new target, 'parallel'. How does the speed compare? What if the array size is much larger?
# + colab={} colab_type="code" id="X4A_kvi1GOkm"
@vectorize(['int64(int64, int64)'], target='parallel')
def add_ufunc_para(x, y):
return x + y
# -
# %timeit add_ufunc_para(c,d)
# + [markdown] colab_type="text" id="KREEf5ZNMKBX"
# ### Problem 5 - Direct Summation
# (This problem borrowed in its entirety from [gforsyth's scipy17 numba tutorial](https://github.com/gforsyth/numba_tutorial_scipy2017).)
# + [markdown] colab_type="text" id="xMPg_TRvRE8Q"
# Many physical problems require the evaluation of all pairwise interactions of a large number of particles, so-called N-body problems. These problems arise in molecular dynamics, astrodynamics and electromagnetics among others.
#
# Their pairwise interactions can be expressed as:
#
# \begin{equation}
# f_i = \sum_{j=1}^n{P \left(\boldsymbol{x}_i, \boldsymbol{x}_j \right)w_j} \ \ \ \text{for } i=1,2,...,n
# \end{equation}
#
# * where subscripts $i$, $j$ respectively denote *target* and *source*
# * $f_i$ can be a *potential* (or *force*) at target point $i$
# * $w_j$ is the *source weight*
# * $\boldsymbol{x}_i, \boldsymbol{x}_j$ are the *spatial positions* of particles
# * $P \left(\boldsymbol{x}_i, \boldsymbol{x}_j \right)$ is the *interaction kernel*.
#
# In order to evalute the potential $f_i$ at a target point $i$, we have to loop over each source particle $j$. Since there are $n$ target points $i$, this 'brute-force' approach costs $\mathcal{O} \left(n^2 \right)$ operations.
# + [markdown] colab_type="text" id="T8Fz6v8cRZe2"
# One possible approach in this kind of problem is to define a few classes, say `Point` and `Particle` and then loop over the objects and perform the necessary point-to-point calculations.
# + colab={} colab_type="code" id="Kgk_4HeaMNI4"
class Point():
"""
Arguments:
domain: the domain of random generated coordinates x,y,z,
default=1.0
Attributes:
x, y, z: coordinates of the point
"""
def __init__(self, domain=1.0):
self.x = domain * np.random.random()
self.y = domain * np.random.random()
self.z = domain * np.random.random()
def distance(self, other):
return ((self.x - other.x)**2 +
(self.y - other.y)**2 +
(self.z - other.z)**2)**.5
# + colab={} colab_type="code" id="XgKlyjf1Rcyl"
class Particle(Point):
"""
Attributes:
m: mass of the particle
phi: the potential of the particle
"""
def __init__(self, domain=1.0, m=1.0):
Point.__init__(self, domain)
self.m = m
self.phi = 0.
# + [markdown] colab_type="text" id="54LlwJUARoQx"
# Next, we define a function to calculate the particle interaction via direct summation:
# + colab={} colab_type="code" id="eaotXcZzRsI6"
def direct_sum(particles):
"""
Calculate the potential at each particle
using direct summation method.
Arguments:
particles: the list of particles
"""
for i, target in enumerate(particles):
for source in (particles[:i] + particles[i+1:]):
r = target.distance(source)
target.phi += source.m / r
# + [markdown] colab_type="text" id="jaUtRaKKS1ZL"
# All that's left is to create a list of random particles with assigned masses:
# + colab={} colab_type="code" id="R8JmgOBtRlnV"
n = 1000
particles = [Particle(m = 1 / n) for i in range(n)]
# + [markdown] colab_type="text" id="tZPmeLQ2SRF-"
# **5a)** Run the direct summation code and determine how long it takes with 10, 100, 1000 particles. Is there a relationship?
# + colab={} colab_type="code" id="5BxIb_3YRuja"
# add code here
# + [markdown] colab_type="text" id="OeTfGagnTcEd"
# Add answer here.
# + [markdown] colab_type="text" id="BkC-j2c2UXbg"
# **How do we use Numba on this problem?** There is a subtle issue here - Numba doesn't support jitting native Python classes. There is a `jit_class` structure in Numba but it's still in early development. But we'd like to have attributes for readable programming. The solution is to build NumPy custom dtypes.
# + colab={} colab_type="code" id="pwAbPrw3SG67"
particle_dtype = np.dtype({'names':['x','y','z','m','phi'],
'formats':[np.double,
np.double,
np.double,
np.double,
np.double]})
# + colab={} colab_type="code" id="0YQzXLouUsXC"
myarray = np.ones(3, dtype=particle_dtype)
# + colab={} colab_type="code" id="gWtollTtU05Q"
myarray
# + [markdown] colab_type="text" id="irR1wcMOVPzH"
# You can access an individual "attribute" like this:
# + colab={} colab_type="code" id="w4oyOqCRU4Zi"
myarray[0]['x'] = 2.0
# + [markdown] colab_type="text" id="sL-2QZVKVYYd"
# **5b)** Write a `jit` function `create_n_random_particles` that takes the arguments `n` (number of particles), `m` (mass of every particle) and a domain within which to generate a random number (as in the class above).
# It should create an array with `n` elements and `dtype=particle_dtype` and then return that array.
#
# For each particle, the mass should be initialized to the value of `m` and the potential `phi` initialized to zero.
# + [markdown] colab_type="text" id="zuPJFbAFXGZV"
# *Hint: You will probably want to loop over the number of particles within the function to assign attributes.*
# + colab={} colab_type="code" id="6_NGgEa_VbLf"
# add code here
# + [markdown] colab_type="text" id="Vp_EbHgJXl9y"
# Now we'll create our array of particles using the new function.
# + colab={} colab_type="code" id="0KKyJWWoXkgj"
particles = create_n_random_particles(1000, .001, 1)
# + colab={} colab_type="code" id="vcaGjOQIXrjU"
particles[:3]
# + [markdown] colab_type="text" id="Vx32Tp4kWwQ-"
# We don't have a `distance` method anymore, so we need to write a function to take care of that.
#
# **5c)** Write a `jit` function `distance` to calculate the distance between two particles of dtype `particle_dtype`.
# + colab={} colab_type="code" id="1EkEnyJSVp9G"
# add code here
# + colab={} colab_type="code" id="nX8TaDkLXcJK"
distance(particles[0], particles[1])
# + colab={} colab_type="code" id="0y18OqcDX-bR"
# %%timeit
distance(particles[0], particles[1])
# + [markdown] colab_type="text" id="JdZRXDJfYN5t"
# **5d)** Modify the `direct_sum` function above to instead work on a NumPy array of particles. Loop over each element in the array and calculate its total potential. Time the result and compare it to your previous version of this function.
# + colab={} colab_type="code" id="FhglZ1oVYEvL"
# add code here
| Session8/Day4/numba_intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
#Only run this once!
# !rm -f pred.npz.bz2 pred.npz
# !wget https://jpata.web.cern.ch/jpata/2101.08578/v1/pred.npz.bz2
# !bzip2 -d pred.npz.bz2
fi = np.load("pred.npz")
ygen = fi["ygen"]
ycand = fi["ycand"]
ypred = fi["ypred"]
ypred_raw = fi["ypred_raw"]
ygen.shape
# We have 100 events, up to 5120 particles in each event, 7 features per particle. We have 3 types of data matrices for each event:
# - ygen - ground truth from the generator
# - ypred - prediction from the MLPF model
# - ycand - prediction from the standard DelphesPF algorithm
#features are (particle ID, charge, pT, eta, sin phi, cos phi, energy)
ygen[0, 0]
#Same for the prediction
ypred[0, 0]
#particle ID (type is)
#0 - no particle
#1 - charged hadron
#2 - neutral hadron
#3 - photon
#4 - electron
#5 - muon
np.unique(ygen[:, :, 0], return_counts=True)
#We also have the raw logits for the multiclass ID prediction
ypred_raw.shape
# +
#Ground truth vs model prediction particles
plt.figure(figsize=(10,10))
ev = ygen[0, :]
msk = ev[:, 0]!=0
plt.scatter(ev[msk, 3], np.arctan2(ev[msk, 4], ev[msk, 5]), s=2*ev[msk, 2], marker="o", alpha=0.5)
ev = ypred[0, :]
msk = ev[:, 0]!=0
plt.scatter(ev[msk, 3], np.arctan2(ev[msk, 4], ev[msk, 5]), s=2*ev[msk, 2], marker="s", alpha=0.5)
plt.xlabel("eta")
plt.ylabel("phi")
| delphes/uncertainty_calibration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# !pip install pymongo
# !pip install pandas
import os, sys
assert sys.version_info.major == 3
import utils
import pandas as pd
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Download Data Form MathWorks
# +
# docs = utils.download_readings_from_mathworks(raw=True)
# -
# ## Upload Data To DB
# +
# utils.upload_readings_to_db(docs)
# -
# ## Download Data From DB
# downloads as pandas dataframe
# site - name of the site (e.g., Skoltech.213) (if None - any)
# tp - type of the measurement (e.g., temperature) (if None - any)
# time_from - request data from date/time (can be datetime.datetime object or string, e.g. '2017-01-01 20:00:00')
# time_to - request data until date/time (can be datetime.datetime object or string, e.g. '2017-01-01 21:00:00')
# raw - if True, then list of jsons, else pandas dataframe with a smaller number of info
data = utils.download_readings_from_db(site=None, tp=None, time_from=None, time_to=None, raw=False)
df = data[data['sensor_id'] == '1']
time = df['time']
vals = df['value']
fig = plt.figure(figsize=(10,6))
plt.plot_date(time, vals, '-')
plt.ylim([20,30])
plt.xlabel('time')
plt.ylabel('Temperature, oC')
plt.show()
| .ipynb_checkpoints/sensors-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19"
# -*- coding: utf-8 -*-
"""keras_bert_classification_tpu.ipynb
Automatically generated by Colaboratory.
Original file is located at
https://colab.research.google.com/drive/1kzGClSAy-SPo_dvgKPIgP0oJCFdZ_8Pa
"""
# @title Preparation
# !pip install -q keras-bert
# !wget -q https://storage.googleapis.com/bert_models/2018_10_18/uncased_L-12_H-768_A-12.zip
# !unzip -o uncased_L-12_H-768_A-12.zip
# @title Constants
SEQ_LEN = 128
BATCH_SIZE = 32 # 64 seems to be the maximum possible on Kaggle
EPOCHS = 5
LR = 1e-4
# @title Environment
import os
pretrained_path = 'uncased_L-12_H-768_A-12'
config_path = os.path.join(pretrained_path, 'bert_config.json')
checkpoint_path = os.path.join(pretrained_path, 'bert_model.ckpt')
vocab_path = os.path.join(pretrained_path, 'vocab.txt')
# TF_KERAS must be added to environment variables in order to use TPU
os.environ['TF_KERAS'] = '1'
# @title Load Basic Model
import codecs
from keras_bert import load_trained_model_from_checkpoint
token_dict = {}
with codecs.open(vocab_path, 'r', 'utf8') as reader:
for line in reader:
token = line.strip()
token_dict[token] = len(token_dict)
model = load_trained_model_from_checkpoint(
config_path,
checkpoint_path,
training=True,
trainable=True,
seq_len=SEQ_LEN,
)
# @title Download IMDB Data
import tensorflow as tf
dataset = tf.keras.utils.get_file(
fname="aclImdb.tar.gz",
origin="http://ai.stanford.edu/~amaas/data/sentiment/aclImdb_v1.tar.gz",
extract=True,
)
# +
# @title Convert Data to Array
import os
import numpy as np
from tqdm import tqdm
from keras_bert import Tokenizer
tokenizer = Tokenizer(token_dict)
def load_data(path):
global tokenizer
indices, sentiments = [], []
for folder, sentiment in (('neg', 0), ('pos', 1)):
folder = os.path.join(path, folder)
for name in tqdm(os.listdir(folder)):
with open(os.path.join(folder, name), 'r') as reader:
text = reader.read()
ids, segments = tokenizer.encode(text, max_len=SEQ_LEN)
indices.append(ids)
sentiments.append(sentiment)
items = list(zip(indices, sentiments))
np.random.shuffle(items)
indices, sentiments = zip(*items)
indices = np.array(indices)
return [indices, np.zeros_like(indices)], np.array(sentiments)
train_path = os.path.join(os.path.dirname(dataset), 'aclImdb', 'train')
test_path = os.path.join(os.path.dirname(dataset), 'aclImdb', 'test')
train_x, train_y = load_data(train_path)
test_x, test_y = load_data(test_path)
# +
print("train_x list details:")
print(len(train_x))
print(train_x[:5])
print(train_y[:5])
print(train_y.shape)
# @title Build Custom Model
from tensorflow.python import keras
from keras_bert import AdamWarmup, calc_train_steps
inputs = model.inputs[:2]
dense = model.get_layer('NSP-Dense').output
outputs = keras.layers.Dense(units=2, activation='softmax')(dense)
decay_steps, warmup_steps = calc_train_steps(
train_y.shape[0],
batch_size=BATCH_SIZE,
epochs=EPOCHS,
)
model = keras.models.Model(inputs, outputs)
#freeze some layers
for layer in model.layers:
layer.trainable = False
model.layers[-1].trainable = True
model.layers[-2].trainable = True
model.layers[-3].trainable = True
model.compile(
AdamWarmup(decay_steps=decay_steps, warmup_steps=warmup_steps, lr=LR),
loss='sparse_categorical_crossentropy',
metrics=['sparse_categorical_accuracy'],
)
# @title Initialize Variables
import tensorflow as tf
import tensorflow.keras.backend as K
sess = K.get_session()
uninitialized_variables = set([i.decode('ascii') for i in sess.run(tf.report_uninitialized_variables())])
init_op = tf.variables_initializer(
[v for v in tf.global_variables() if v.name.split(':')[0] in uninitialized_variables]
)
sess.run(init_op)
# +
from keras_bert import get_custom_objects
# @title Fit
with tf.keras.utils.custom_object_scope(get_custom_objects()):
model.fit(
train_x,
train_y,
epochs=EPOCHS,
batch_size=BATCH_SIZE,
)
# @title Predict
with tf.keras.utils.custom_object_scope(get_custom_objects()):
predicts = model.predict(test_x, verbose=True).argmax(axis=-1)
# @title Accuracy
print(np.sum(test_y == predicts) / test_y.shape[0])
# -
# !ls
| Nokore/scripts/Keras-Bert Classification of Movie Sentiments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/gauravsmind/PythonLessons/blob/master/Basics.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="nHYpD9ph1CAs" colab_type="text"
# #Example of print commands
# + id="boovzBGNjS5V" colab_type="code" outputId="81bfb0bb-1cb0-48a8-b16a-bca9118c0a66" colab={"base_uri": "https://localhost:8080/", "height": 119}
print("Hello World!")
print("Hello Again")
print ("I like typing this.")
print("This is fun.")
print("Yay! Printing.")
print("I'd rather you 'not'.")
#print('I "said" so not touch this.')
# + id="lHjIgx8pxazW" colab_type="code" outputId="6011d2cb-2405-44af-c223-e60be5fbdb7b" colab={"base_uri": "https://localhost:8080/", "height": 51}
# A comment, like this so that you can read later.
# Anything after the # is ignored by python.
print ("I could have coded like this.") # and the comment after this is ignored.
# you can also use a comment to "disable" or comment out code:
print("This will run.")
# + [markdown] id="g70kI8zAxN8a" colab_type="text"
# Example 2
# + id="kAHuD1KCjXlA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="a98bfe21-eec2-4134-ce7d-eb9abd49c195"
types_of_people = 10
x = f"There are {types_of_people} types of people."
binary = "binary"
do_not = "don't"
y = f"Those who know {binary} and those who {do_not}."
print(x)
print(y)
print(f"I said: {x}")
| Basics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import xml.etree.ElementTree as ET
tree2=ET.parse('../datasets/xml1.xml')
type(tree2)
root=tree2.getroot()
root[0][2]
root[0][2].text
root[0][2].tag
root[0]
root[0].tag
root[0].attrib
| Chapter07/Exercise 7.10/.ipynb_checkpoints/Exercise 7.10-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ticket_history]
# language: python
# name: conda-env-ticket_history-py
# ---
# +
# Set desired ticket ID
ticket_id = 'PY-1706'
# Request history XML
import requests
import auth
url = 'https://youtrack.jetbrains.com/rest/issue/{}/history'.format(ticket_id)
r = requests.get(url, auth=(auth.username,auth.password))
# Load the XML
import xml.etree.ElementTree as ET
tree = ET.fromstring(r.content)
# each version of the issue is in an 'issue' tag
from datetime import datetime
updates = [issue for issue in tree]
votes_over_time = {}
def ms_to_s(ms):
return int(ms) // 1000
def field_by_name(parent, name):
# YouTrack describes a version with a set of 'field' elements which are identified
# by the 'name' attribute
#
# The updated time is stored in field[name=updated]
xpath = "*[@name='{}']".format(name)
fields = parent.findall(xpath)
# Fields names are supposed to be unique, therefore take the 0th element
# of the list of found fields
if len(fields) < 1:
raise KeyError('Could not find key: ' + name)
return fields[0]
# extract updates over time from the XML
for update in updates:
try:
# "Updated" time is in field[name=updated]
updated_time_field = field_by_name(update, 'updated')
# The actual value is stored in a 'value' child element
# with a 'SingleField' like 'updated', there's only one
# 'value' child element
updated_time_ms = updated_time_field.find('value').text
# get the datetime
updated = datetime.utcfromtimestamp(ms_to_s(updated_time_ms))
voter_name_fields = field_by_name(update, 'voterName')
voters = [field.text for field in voter_name_fields]
votes_over_time[updated] = voters
except KeyError as e:
print(e)
# +
import pandas as pd
counts_over_time = {}
for date, votes in votes_over_time.items():
counts_over_time[pd.to_datetime(date)] = len(votes)
# -
counts_series = pd.Series(list(counts_over_time.values()), index=counts_over_time.keys())
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib
matplotlib.style.use('ggplot')
counts_series.plot()
# -
| votes_over_time.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#import dependencies
import pandas as pd
from splinter import Browser
from bs4 import BeautifulSoup
from webdriver_manager.chrome import ChromeDriverManager
#setup path & initialize browser
executable_path = {'executable_path': ChromeDriverManager().install()}
browser = Browser('chrome', **executable_path, headless=False)
#visit Mars news site
url = "https://redplanetscience.com/"
browser.visit(url)
#parse results with BeautifulSoup - collect the latest News Title and Paragraph Text
html = browser.html
news_soup = BeautifulSoup(html,"html.parser")
news_soup
# +
#slide_element = news_soup.select_one("div.content_title").text
#slide_element
# -
news_title = news_soup.find("div", class_="content_title").text
print(news_title)
news_paragraph = news_soup.find("div", class_="article_teaser_body").text
print(news_paragraph)
# ## JPL Mars Space Images - Featured Image
#visit Mars space site
url = "https://spaceimages-mars.com/"
browser.visit(url)
html = browser.html
mars_image_soup = BeautifulSoup(html,"html.parser")
mars_image_soup
mars_image = mars_image_soup.find("img", class_="headerimage fade-in").get("src")
print(mars_image)
featured_image_url = f'https://spaceimages-mars.com/{mars_image}'
featured_image_url
# ## Mars facts
# +
#visit Mars facts site and use Pandas to read
# -
mars_df = pd.read_html ("https://galaxyfacts-mars.com/")[1]
print(mars_df)
mars_df.columns=['description', 'value']
#mars_df.set_index('description', inplace=True)
mars_df
# + active=""
# mars_df.to_html()
# -
# ## <NAME>
#visit the astrogeology site
url = "https://marshemispheres.com/"
browser.visit(url)
#obtain images of all hemispheres
html = browser.html
hemispheres_soup = BeautifulSoup(html,"html.parser")
hemispheres_soup
links = browser.find_by_css("a.product-item h3")
links
# +
hemisphere_image_urls = []
# First, get a list of all of the hemispheres
links = browser.find_by_css("a.product-item h3")
# Next, loop through those links, click the link, find the sample anchor, return the href
for i in range(4):
hemisphere = {}
# We have to find the elements on each loop to avoid a stale element exception
browser.find_by_css("a.product-item h3")[i].click()
# Next, we find the Sample image anchor tag and extract the href
# sample_elem = browser.find_link_by_text('Sample').first
sample_elem = browser.links.find_by_text('Sample').first
hemisphere['img_url'] = sample_elem['href']
# Get Hemisphere title
hemisphere['title'] = browser.find_by_css("h2.title").text
# Append hemisphere object to list
hemisphere_image_urls.append(hemisphere)
# Finally, we navigate backwards
browser.back()
# -
hemisphere_image_urls
browser.quit()
| mission_to_mars.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import tess_cpm
import numpy as np
import matplotlib.pyplot as plt
import lightkurve as lk
from astropy.io import fits
from astropy.wcs import WCS
def cpm_periodogram(fits_file, t_row=32, t_col=32):
cpm = tess_cpm.CPM(fits_file, remove_bad=True)
cpm.set_target(t_row, t_col)
cpm.set_exclusion(10)
cpm.set_predictor_pixels(256, method='cosine_similarity')
cpm.lsq(0.1, rescale=True, polynomials=False)
tess_cpm.summary_plot(cpm, 10)
aperture_lc, lc_matrix = cpm.get_aperture_lc(box=1, show_pixel_lc=True, show_aperture_lc=True)
lc = lk.LightCurve(time=cpm.time, flux=aperture_lc)
pg = lc.to_periodogram(oversample_factor=10)
fig, axs = plt.subplots(2, 1, figsize=(15, 8))
pg.plot(ax=axs[0], c='k')
pg.plot(ax=axs[1], c='k', view='period')
fig.suptitle("Periodogram", fontsize=20, y=0.95)
period = pg.period_at_max_power
print(f"Max Power Period: {period}")
lc.fold(period.value*4).scatter()
plt.title(f"Folded Lightcurve with Period: {period:.4f}", fontsize=20)
return cpm
# +
# fits_file = "provided_sources/tess-s0005-1-3_70.445653_7.275704_64x64_astrocut.fits"
# fits_file = "provided_sources/tess-s0005-1-3_72.267451_8.981257_64x64_astrocut.fits"
# fits_file = "provided_sources/tess-s0005-1-3_71.975577_7.370718_64x64_astrocut.fits"
# fits_file = "provided_sources/tess-s0006-1-1_84.180039_3.774854_64x64_astrocut.fits"
# fits_file = "provided_sources/tess-s0002-1-3_357.154800_-15.211056_64x64_astrocut.fits"
# -
cpm_periodogram("provided_sources/tess-s0005-1-3_70.445653_7.275704_64x64_astrocut.fits")
cpm_periodogram("provided_sources/tess-s0005-1-3_72.267451_8.981257_64x64_astrocut.fits")
cpm_periodogram("provided_sources/tess-s0005-1-3_71.975577_7.370718_64x64_astrocut.fits")
cpm_periodogram("provided_sources/tess-s0006-1-1_84.180039_3.774854_64x64_astrocut.fits")
cpm_periodogram("provided_sources/tess-s0002-1-3_357.154800_-15.211056_64x64_astrocut.fits")
cpm_periodogram("provided_sources/tess-s0015-1-1_321.475220_46.623558_64x64_astrocut.fits")
cpm = cpm_periodogram("provided_sources/tess-s0015-1-1_322.782190_48.644292_64x64_astrocut.fits")
# aperture_lc, lc_matrix = cpm.get_aperture_lc(show_pixel_lc=True, show_aperture_lc=True)
# +
# lc = lk.LightCurve(time=cpm.time, flux=aperture_lc)
# pg = lc.to_periodogram(oversample_factor=10)
# fig, axs = plt.subplots(2, 1, figsize=(15, 8))
# pg.plot(ax=axs[0], c='k')
# pg.plot(ax=axs[1], c='k', view='period')
# fig.suptitle("Periodogram", fontsize=20, y=0.95)
# period = pg.period_at_max_power
# print(f"Max Power Period: {period}")
# lc.fold(period.value*4).scatter()
# plt.title(f"Folded Lightcurve with Period: {period:.4f}", fontsize=20)
# -
cpm = cpm_periodogram("provided_sources/tess-s0015-1-1_322.748930_47.808133_64x64_astrocut.fits")
# aperture_lc, lc_matrix = cpm.get_aperture_lc(show_pixel_lc=True, show_aperture_lc=True)
# +
# lc = lk.LightCurve(time=cpm.time, flux=aperture_lc)
# pg = lc.to_periodogram(oversample_factor=10)
# fig, axs = plt.subplots(2, 1, figsize=(15, 8))
# pg.plot(ax=axs[0], c='k')
# pg.plot(ax=axs[1], c='k', view='period')
# fig.suptitle("Periodogram", fontsize=20, y=0.95)
# period = pg.period_at_max_power
# print(f"Max Power Period: {period}")
# lc.fold(period.value*4).scatter()
# plt.title(f"Folded Lightcurve with Period: {period:.4f}", fontsize=20)
# +
# cpm = cpm_periodogram("provided_sources/tess-s0015-1-1_322.878520_47.920650_64x64_astrocut.fits") # Center
cpm = cpm_periodogram("provided_sources/tess-s0015-1-1_322.878520_47.920650_64x64_astrocut.fits", t_row=30, t_col=31)
# +
# lc = lk.LightCurve(time=cpm.time, flux=aperture_lc)
# pg = lc.to_periodogram(oversample_factor=10)
# fig, axs = plt.subplots(2, 1, figsize=(15, 8))
# pg.plot(ax=axs[0], c='k')
# pg.plot(ax=axs[1], c='k', view='period')
# fig.suptitle("Periodogram", fontsize=20, y=0.95)
# period = pg.period_at_max_power
# print(f"Max Power Period: {period}")
# lc.fold(period.value*4).scatter()
# plt.title(f"Folded Lightcurve with Period: {period:.4f}", fontsize=20)
# -
cpm_periodogram("provided_sources/tess-s0015-1-1_325.106020_50.169629_64x64_astrocut.fits")
| periodogram.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import subprocess
import requests
import re
from bs4 import BeautifulSoup
from selenium import webdriver
from urllib.request import urlopen
# -
# 1. !pip install nsetools
#
# 2. !pip install bsetools
#
# 3. !pip install bsedata
# # 1.
# Write a program to pull - 'Buy(Total Quantity) & Sell(Total Quantity)' from the below url:
# https://beta.nseindia.com/get-quotes/derivatives?symbol=BANKNIFTY&identifier=OPTIDXBANKNIFTY29-08-2019CE28000.00
#
baseURL='https://beta.nseindia.com/get-quotes/derivatives?symbol=BANKNIFTY&identifier=OPTIDXBANKNIFTY29-08-2019CE28000.00 '
html = urlopen(baseURL)
soup = BeautifulSoup(html, 'lxml')
type(soup)
title = soup.title
print(title)
text = soup.get_text()
text
soup.find_all("")
| Data-Science-HYD-2k19/Projects/codes/PROJECT 2 ( mercury) ( Web-Scrapping)/.ipynb_checkpoints/Project_Mercury_Omega-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import warnings
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import statsmodels.api as sm
from pmdarima import auto_arima
from sklearn import metrics
from statsmodels.tsa.seasonal import seasonal_decompose
from statsmodels.tsa.stattools import adfuller
warnings.filterwarnings("ignore")
# -
df = pd.read_csv(r'\Data\FB.csv')
df.head(10)
def timeseries_evaluation_metrics_func(y_true, y_pred):
def mean_absolute_percentage_error(y_true, y_pred):
y_true, y_pred = np.array(y_true), np.array(y_pred)
return np.mean(np.abs((y_true - y_pred) / y_true)) * 100
print('Evaluation metric results:-')
print(f'MSE is : {metrics.mean_squared_error(y_true, y_pred)}')
print(f'MSE is : {metrics.mean_absolute_error(y_true, y_pred)}')
print(f'RMSE is : {np.sqrt(metrics.mean_squared_error(y_true, y_pred))}')
print(f'MAPE is : {mean_absolute_percentage_error(y_true, y_pred)}')
print(f'R2 is : {metrics.r2_score(y_true, y_pred)}',end='\n\n')
def Augmented_Dickey_Fuller_Test_func(series , column_name):
print (f'Results of Dickey-Fuller Test for column: {column_name}')
dftest = adfuller(series, autolag='AIC')
dfoutput = pd.Series(dftest[0:4], index=['Test Statistic','p-value','No Lags Used','Number of Observations Used'])
for key,value in dftest[4].items():
dfoutput['Critical Value (%s)'%key] = value
print (dfoutput)
if dftest[1] <= 0.05:
print("Conclusion:====>")
print("Reject the null hypothesis")
print("Data is stationary")
else:
print("Conclusion:====>")
print("Fail to reject the null hypothesis")
print("Data is non-stationary")
for name, column in df[['Close' ,'Open' ,'High','Low']].iteritems():
Augmented_Dickey_Fuller_Test_func(df[name],name)
print('\n')
X = df[['Close' ]]
actualtrain, actualtest = X[0:-30], X[-30:]
exoX = df[['Open' ]]
exotrain, exotest = exoX[0:-30], exoX[-30:]
# +
for m in [1, 4,7,12,52]:
print("="*100)
print(f' Fitting SARIMAX for Seasonal value m = {str(m)}')
stepwise_model = auto_arima(actualtrain,exogenous =exotrain ,start_p=1, start_q=1,
max_p=7, max_q=7, seasonal=True,start_P=1,start_Q=1,max_P=7,max_D=7,max_Q=7,m=m,
d=None,D=None, trace=True,error_action='ignore',suppress_warnings=True, stepwise=True)
print(f'Model summary for m = {str(m)}')
print("-"*100)
stepwise_model.summary()
forecast,conf_int = stepwise_model.predict(n_periods=30,exogenous =exotest,return_conf_int=True)
df_conf = pd.DataFrame(conf_int,columns= ['Upper_bound','Lower_bound'])
df_conf["new_index"] = range(1229, 1259)
df_conf = df_conf.set_index("new_index")
forecast = pd.DataFrame(forecast, columns=['close_pred'])
forecast["new_index"] = range(1229, 1259)
forecast = forecast.set_index("new_index")
timeseries_evaluation_metrics_func(actualtest, forecast)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams["figure.figsize"] = [15, 7]
plt.plot(actualtrain, label='Train ')
plt.plot(actualtest, label='Test ')
plt.plot(forecast, label=f'Predicted with m={str(m)} ')
plt.plot(df_conf['Upper_bound'], label='Confidence Interval Upper bound ')
plt.plot(df_conf['Lower_bound'], label='Confidence Interval Lower bound ')
plt.legend(loc='best')
plt.show()
print("-"*100)
print(f' Diagnostic plot for Seasonal value m = {str(m)}')
display(stepwise_model.plot_diagnostics());
print("-"*100)
# -
| hands-on-time-series-analylsis-python/Chapter 4/5. SARIMAX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # Installation
# +
# #!pip install matplotlib
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import numpy as np
# for model building
from sklearn.linear_model import LinearRegression
from sklearn.linear_model import Ridge
from sklearn.model_selection import train_test_split
from sklearn import metrics
from sklearn.feature_selection import RFE
# for model evaluation
from sklearn.metrics import r2_score
from sklearn.metrics import mean_squared_error
# -
df = pd.read_csv("/Users/jessicazhang/Desktop/data.csv")
df = df.sample(frac=1, random_state=2021)
df.shape
# # Adding Columns
# ## 1. for x5^0.5
# +
# x5 is the number of bathroom
# Bathroom_sqrt is the result of square of root
df['Bathroom_sqrt'] = np.sqrt(df['Bathroom'])
df.head()
# -
# ## 2. for x5^2
# +
# Bathroom_sqr means Bathroom*Bathroom
# pow() means the square
# i.e. pow(4,2) = 4^2 = 16
df['Bathroom_sqr'] = pow(df['Bathroom'],2)
df.head()
# -
# ## 3. for x8^0.5
# +
# x8 is the building area
# BuildingArea_sqr is the square root.
df['BuildingArea_sqrt'] = np.sqrt(df['BuildingArea'])
df.head()
# -
# ## 4. for x8^2
# +
# BuildingArea_sqr is the square result.
df['BuildingArea_sqr'] = pow(df['BuildingArea'],2)
df.head()
# -
# ## 5. for x9^0.5
# +
# x9 is the distance
# Distance_sqrt is the square root.
df['Distance_sqrt'] = np.sqrt(df['Distance'])
df.head()
# -
# ## 6. for x9^2
# +
# Distance_sqr is the square.
df['Disctance_sqr'] = pow(df['Distance'],2)
df.head()
# -
# ## 7. combination
# +
# X8 is the building area
# pow() means the square of the number
# i.e. pow(4,2) = 4*4 = 16
df['BuildingArea_sqr'] = pow(df['BuildingArea'],2)
df['Distance_sqr'] = pow(df['Distance'], 2)
df['Bathroom_sqrt']= np.sqrt(df['Bathroom'])
df.head()
# -
# # Drop Column
df = df.drop(['Rooms'], axis=1)
df = df.drop(['Landsize'], axis=1)
df.head()
# # Spliting into Train and Test Data
df["YearBuilt"] = 2021 - df["YearBuilt"]
df.head()
# Drop the last column
X = df.drop(['Price'], axis = 1)
X
y = df["Price"]
print(y)
# ## Normalization
from sklearn import preprocessing
min_max_scaler = preprocessing.MinMaxScaler()
X = min_max_scaler.fit_transform(X)
# ## Model
# +
# Divide the data into two groups: training set and testing set
# X_train, X_test, y_train, y_test = train_test_split(X, y, random_state=320, test_size=0.27)
# -
# ### K Fold
from sklearn.model_selection import KFold
kf = KFold(n_splits=5)
for train_index, test_index in kf.split(X):
print("TRAIN:", train_index, "TEST:", test_index)
X_train, X_test = X[train_index], X[test_index]
y_train, y_test = y[train_index], y[test_index]
print(X_train.shape)
print(X_test.shape)
print(y_train.shape)
print(y_test.shape)
# ### Linear Regression
model = LinearRegression()
model.fit(X_train, y_train)
# ## Model Evaluation
# ### Score
from sklearn.model_selection import cross_val_score
# (1) Orginal
orginal_score = cross_val_score(model, X, y, cv=5)
print("Original score:",orginal_score)
# (2) Drop a column ‘Rooms’ and Add a new column: x8^0.5
drop_score = cross_val_score(model, X, y, cv=5)
print("After dropping one column:",drop_score)
# (3) Drop a column & Feature Engineer (use the combination)
combined_score = cross_val_score(model, X, y, cv=5)
print("Combination score:",combined_score)
# ## Based on the Third Model
# ### Coefficients & RMSE
# y_pred_class = model.predict(X_test)
model.predict(X_test)
y_test
# +
# Root Mean Squared Error
def customer_scorer(model, X, y):
predict_y = model.predict(X)
rmse = np.sqrt(np.power(predict_y - y, 2).sum() / len(y))
# rmse = mean_squared_error(predict_y, y, squared=False)
return rmse
# -
original_rmse = cross_val_score(model, X, y, cv=5, scoring=customer_scorer)
print("Original RMSE:" + (', {:.4f}' * 5).format(*original_rmse))
drop_rmse = cross_val_score(model, X, y, cv=5, scoring=customer_scorer)
print("Drop RMSE:" + (', {:.4f}' * 5).format(*drop_rmse))
combined_rmse = cross_val_score(model, X, y, cv=5, scoring=customer_scorer)
print("Combination RMSE:" + (', {:.4f}' * 5).format(*combined_rmse))
# ## Model Comparison
# #### Score
# +
original_score = [0.52330568, 0.5656773, 0.52000582, 0.54640202, 0.47200789]
drop_score = [0.5514069, 0.58787212, 0.53455358, 0.56154552, 0.4869308 ]
combined_score = [0.54541132, 0.59951113, 0.53779354, 0.57049032, 0.49503245]
plt.boxplot([original_score, drop_score, combined_score],vert=False,showmeans=True)
plt.xlabel('Scores')
plt.ylabel('Model')
plt.title('Model Comparison')
plt.show()
# -
# ### Comment: Model 3 is the best one.
# #### RMSE
# +
original_rmse = [439363.8769, 418108.6552, 442942.7409, 418938.8163, 496776.7295]
drop_rmse = [426216.8884, 407285.4468, 436178.6955, 411886.2597, 489706.0840]
combined_rmse = [429055.6920, 401493.1292, 434657.9301, 407663.2127, 485824.3327]
plt.boxplot([original_rmse, drop_rmse, combined_rmse],vert=False,showmeans=True)
plt.xlabel('RMSE')
plt.ylabel('Model')
plt.title('RMSE Comparison')
plt.show()
# -
np.mean(combined_rmse)
| Scripts/Linear_Regression_FinalVersion/LinearRegression_Updated_Performance.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.1
# language: julia
# name: julia-1.6
# ---
DATE = "2021-06-26"
TASK = "1000bp-100x-coverage-0.01-error-rate"
DIR = "$(DATE)-$(TASK)"
DIR = mkpath("$(homedir())/$(DIR)")
# +
pkgs = [
"Graphs",
"MetaGraphs",
"BioSequences",
"Random",
"ProgressMeter",
"Revise",
"FASTX"
]
import Pkg
Pkg.add(pkgs)
for pkg in pkgs
eval(Meta.parse("import $(basename(pkg))"))
end
import Mycelia
# -
# set a random seed
seed = Random.seed!(0)
# randomly generate a dna sequence of 100bp
genome = BioSequences.randdnaseq(seed, 1000)
# define error rate
error_rate = 0.01
# +
# generate 100x coverage fastq file
# put accuracy rate into fastq file
coverage = 100
fastq_file = "$(DIR)/$(DATE)-$(TASK).fastq"
error_free_fastq_file = "$(DIR)/$(DATE)-$(TASK).error-free.fastq"
fastq_io = FASTX.FASTQ.Writer(open(fastq_file, "w"))
error_free_fastq_io = FASTX.FASTQ.Writer(open(error_free_fastq_file, "w"))
for i in 1:coverage
true_sequence = Mycelia.observe(genome, error_rate=0.0)
quality_scores = fill(60, length(true_sequence))
error_free_fastq_record = FASTX.FASTQ.Record("$i", true_sequence, quality_scores)
write(error_free_fastq_io, error_free_fastq_record)
observed_sequence = Mycelia.observe(genome, error_rate=error_rate)
q = -10 * log10(error_rate)
quality_scores = fill(q, length(observed_sequence))
fastq_record = FASTX.FASTQ.Record("$i", observed_sequence, quality_scores)
write(fastq_io, fastq_record)
end
close(fastq_io)
close(error_free_fastq_io)
# -
k = 13
kmer_type = BioSequences.BigDNAMer{k}
function visualize_gfa_file(gfa_file)
run(`/Applications/Bandage.app/Contents/MacOS/Bandage image $(gfa_file) $(gfa_file).svg --depwidth 1 --deppower 1`)
# --nodewidth <float> Average node width (0.5 to 1000, default: 5)
# --depwidth <float> Depth effect on width (0 to 1, default: 0.5)
# --deppower <float> Power of depth effect on width (0 to 1, default: 0.5)
html_path_to_svg = "./" * repeat("../", length(split(pwd(), '/')) - 3)
html_path_to_svg *= replace("$(gfa_file).svg", "$(homedir())/" => "")
x = display("text/html", "<img src=$(html_path_to_svg)>")
end
error_free_simple_kmer_graph = Mycelia.fastx_to_simple_kmer_graph(kmer_type, error_free_fastq_file)
# visualize
gfa_file = error_free_fastq_file * ".k-$k.gfa"
Mycelia.graph_to_gfa(error_free_simple_kmer_graph, gfa_file)
visualize_gfa_file(gfa_file)
simple_kmer_graph = Mycelia.fastx_to_simple_kmer_graph(kmer_type, fastq_file)
gfa_file = fastq_file * ".k-$k.gfa"
Mycelia.graph_to_gfa(simple_kmer_graph, gfa_file)
visualize_gfa_file(gfa_file)
@time polished_fastq_file = Mycelia.simple_polish_fastq(simple_kmer_graph, fastq_file, min_depth=7)
polished_simple_kmer_graph = Mycelia.fastx_to_simple_kmer_graph(kmer_type, polished_fastq_file)
gfa_file = polished_fastq_file * ".k-$k.gfa"
Mycelia.graph_to_gfa(polished_simple_kmer_graph, gfa_file)
visualize_gfa_file(gfa_file)
Mycelia.kmer_graph_distances(error_free_simple_kmer_graph, simple_kmer_graph)
Mycelia.kmer_graph_distances(error_free_simple_kmer_graph, polished_simple_kmer_graph)
| docs/_src/5.Development/2021-06-26-1000bp-100x-coverage-0.01-error-rate.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.0.3
# language: julia
# name: julia-1.0
# ---
push!(LOAD_PATH, "../../src")
using FCA
using Colors, Images, FileIO, Plots, Statistics, LinearAlgebra, Random
function extractpatches(image,patchsize,stride=1)
return Matrix{eltype(image)}[image[i:i+patchsize-1,j:j+patchsize-1]
for i in 1:stride:size(image,1)-(patchsize-1),
j in 1:stride:size(image,2)-(patchsize-1)]
end
image = "../images/hedgehog.jpg"
img = Float64.(Gray.(load(image)))
Gray.(img)
include("patch_displaying_utils.jl")
patches = extractpatches(img,6)
patchmat = hcat(vec.(patches)...);
@show size(patchmat)
viewpatches(rand(patches,(3,18)))
batch_size = 50000
npatches = size(patchmat,2)
rand_idx = randperm(npatches)[1:batch_size]
rand_patchmat = patchmat[:,rand_idx];
# ## ICA patches
# We regard these small patches are linear combinations of $64$ $\sf ICA$ components. Assume that the coefficient of a certain ICA component between different patches are i.i.d random variables. Further assume that the coefficents for different ICA components are independent, this patches can be regarded as generated following
# a ICA model:
#
# $$
# patchmat = W_{ica} X_{ica}
# $$
#
# where columns of $W_{ica}$ is the vectorization of ICA components, $X_{ica}$, which represents the coefficients, is modeled as i.i.d samples of independent random variables.
Zica = [vec(rand_patchmat[i,:]) for i = 1: size(rand_patchmat, 1)];
@time Wica, Xica = icf(Zica; opt = "orth");
Wica = Wica[:, sortperm(-vec(std(Wica, dims = 1)))];
viewpatches(reshape([reshape(normpatch(Wica[:,i]), (6,6)) for i = 1: size(Wica, 2)], (6,6)))
# ## FCA patches
# We can regard those small patches are linear combinations of $63$ $\sf FCA$ components (the uniform one is removed as we assume the centered matrix).
# In contrast to $\sf ICA$ case, we do not make assumption on the coefficients.
# Instead, we regard $\sf FCA$ components are freely independent matrices.
#
# Reshaping the columns of $patchmat$ back to patches, we get an array $patcharr$ of $8 \times 8$ matrices. By above discussion, we have the decomposition
#
# $$
# patcharr = W_{fca} X_{fca},
# $$
#
# where the free components in $X_{fca}$ stand for the $\sf FCA$ components and $W_{fca}$ denotes corresponding coefficients.
patcharr = [reshape(rand_patchmat[:, i], (6,6)) for i = 1: size(rand_patchmat, 2)];
# Note that if we set `batch_size` to be $50000$, $W_{fca}$ will be a $50000 \times 64$ matrix. The whitening process in FCF, whien being applied to `patcharr`, involves the eigenvalue decomposition of a $50000 \times 50000$ matrix. (We will take first $64$ eigenvalues and eigenvectors, as the rest eigenvalues are $0$ and irrelavant to the data.) This is time comsuming and unnecessary, a SVD for matrices is desired here.
#
# Note that the covariance of a pair of centered matrices $X_1, X_2 \in \mathbb R^{N \times M}$ is
#
# $$
# \frac{1}{N} \mathrm{Tr}(X_1 \cdot X_2^T),
# $$
#
# which is equivalent to
#
# $$
# \frac{1}{N} vec(X_1)^T \cdot vec(X_2).
# $$
#
# Therefore,
# The non-zero eigenvalues of
#
# $$
# \left[\frac 1 N Tr.(patcharr[i]*patcharr[j]') \right]_{i,j = 1}^{50000}
# $$
#
# are $\frac1N \sigma_i^2$ where $\sigma_i$ denotes the non-zero singualr value of `patchmat`.
# The corresponding eigenvectors are exactly the right singular vectors of `patchmat`.
#
#
# Let us verify this in the following block.
# +
# sample an array of 64 element
testarr = patcharr[1:64];
# results from free_whitening
Y, U, Σ = free_whiten(testarr; mat = "rec");
# results from applying SVD to the patchmat
testmat = rand_patchmat[:, 1:64]
# center the columns of testmat
testmat = testmat - ones(size(testmat, 1))* mean(testmat; dims = 1)
# SVD
Us, Σs, = svd(testmat');
# -
heatmap(U'*Us)
# +
## we now apply SVD to the 50000 patches
rand_patchmat = rand_patchmat - ones(size(rand_patchmat, 1))* mean(rand_patchmat; dims = 1);
U, Σ, = svd(rand_patchmat');
## take first 63 singular value and singular vectors
r = 35
U = U[:, 1: r]
Σ = Σ[1: r]
Y = pinv(Diagonal(Σ))*U'*[(reshape(rand_patchmat[:, i], (6,6))
- mean(rand_patchmat[:, i]) * ones(6, 6)) for i = 1: size(rand_patchmat, 2)];
# -
Wfca, Xfca = freecf(Y; mat = "rec", opt = "orth", obj = "kurt");
# add the uniform patch if necessary
# Xfca = Xfca[sortperm([-std(vec(Xfca[i])) for i = 1: size(Xfca,1)])];
patchfca = [normpatch(Xfca[i]) for i in 1:size(Xfca, 1)]
prepend!(patchfca, [0.5*ones(6,6)])
viewpatches(reshape(patchfca,(6,6)))
| examples/patch/FCApatch_vs_ICApatch.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Async client examples
#
# More async client examples [here](https://apache-ignite-binary-protocol-client.readthedocs.io/en/latest/async_examples.html)
# +
import asyncio
import pandas as pd
from pyignite import GenericObjectMeta, AioClient
from pyignite.datatypes import String, IntObject, DoubleObject
from pyignite.datatypes.prop_codes import PROP_NAME, PROP_SQL_SCHEMA, PROP_QUERY_ENTITIES
# -
CACHE_CONFIG = {
PROP_NAME: 'sql_public_student',
PROP_SQL_SCHEMA: 'public',
PROP_QUERY_ENTITIES: [
{
'table_name': 'student',
'key_field_name': 'sid',
'key_type_name': 'java.lang.Long',
'field_name_aliases': [],
'query_fields': [
{
'name': 'sid',
'type_name': 'java.lang.Long',
'is_key_field': True,
'is_notnull_constraint_field': True,
},
{
'name': 'name',
'type_name': 'java.lang.String',
},
{
'name': 'login',
'type_name': 'java.lang.String',
},
{
'name': 'age',
'type_name': 'java.lang.Integer',
},
{
'name': 'gpa',
'type_name': 'java.math.Double',
},
],
'query_indexes': [],
'value_type_name': 'sql_public_student_type',
'value_field_name': None,
},
],
}
class Student(
metaclass=GenericObjectMeta,
type_name='sql_public_student_type',
schema={
'name': String,
'login': String,
'age': IntObject,
'gpa': DoubleObject,
}
):
pass
students = [
{'name': '<NAME>', 'login': 'jdoe', 'age': 18, 'gpa': 2.5},
{'name': '<NAME>', 'login': 'janed', 'age': 25, 'gpa': 3.5},
{'name': '<NAME>', 'login': 'evaluator', 'age': 45, 'gpa': 4.9}
]
client = AioClient(partition_aware=True)
loop = asyncio.get_event_loop()
# ### Cache operations example
# +
async def cache_api():
async with client.connect([('127.0.0.1', 10800 + i) for i in range(3)]):
cache = await client.get_or_create_cache(CACHE_CONFIG)
await cache.put_all({i: Student(**st) for i, st in enumerate(students)})
all_data = await cache.get_all([i for i in range(len(students))])
row_names = ['name', 'login', 'age', 'gpa', 'version']
rows = []
for k,v in all_data.items():
row = [k]
row += [getattr(v, name, None) for name in row_names]
rows.append(row)
return pd.DataFrame(rows, columns=['sid'] + row_names)
# We must use task because jupyter uses tornado and event loop is in use.
await loop.create_task(cache_api())
# -
# ### Scan query example
# +
async def scan_query():
async with client.connect([('127.0.0.1', 10800 + i) for i in range(3)]):
cache = await client.get_or_create_cache(CACHE_CONFIG)
await cache.put_all({i: Student(**st) for i, st in enumerate(students)})
row_names = ['name', 'login', 'age', 'gpa', 'version']
rows = []
async with cache.scan() as cursor:
async for k,v in cursor:
row = [k]
row += [getattr(v, name, None) for name in row_names]
rows.append(row)
return pd.DataFrame(rows, columns=['sid'] + row_names)
# We must use task because jupyter uses tornado and event loop is in use.
await loop.create_task(scan_query())
# -
# ### Sql query example
# +
async def sql_query():
async with client.connect([('127.0.0.1', 10800 + i) for i in range(3)]):
async with client.sql('select * from student', include_field_names=True) as cursor:
row_names = await cursor.__anext__()
rows = [row async for row in cursor]
return pd.DataFrame(rows, columns=row_names)
# We must use task because jupyter uses tornado and event loop is in use.
await loop.create_task(sql_query())
| examples/basic_async.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Xgboost Classification
# + _cell_guid="adfbe30e-7ebb-0f88-d917-d9a8f97c638e"
import numpy as np
import pandas as pd
import plotly.offline as py
py.init_notebook_mode(connected=True)
import plotly.graph_objs as go
import plotly.tools as tls
import seaborn as sns
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import accuracy_score, log_loss
from imblearn.over_sampling import SMOTE
import xgboost as xgb # Load this xgboost
from sklearn.model_selection import train_test_split
# Import and suppress warnings
import warnings
warnings.filterwarnings('ignore')
from sklearn.model_selection import GridSearchCV, RandomizedSearchCV, cross_val_score, learning_curve, train_test_split
# + [markdown] _cell_guid="5af03c82-cb84-d943-f82c-fc0a15d46b48"
# # 1. Exploratory Data Analysis
#
# Let us load in the dataset via the trusty Pandas package into a dataframe object which we call **attrition** and have a quick look at the first few rows
# + _cell_guid="e035b071-50f8-43ca-9611-fc47272bb05e"
attrition = pd.read_csv('WA_Fn-UseC_-HR-Employee-Attrition.csv')
attrition.head()
# + _cell_guid="57e2bf45-5920-af03-50c1-b5bba334eb11"
# Looking for NaN
attrition.isnull().any()
# +
# attrition.Age.fillna('')
# + [markdown] _cell_guid="5c5dc2ed-7608-4d84-c4f6-c591a3be7570"
# ### Correlation of Features
#
# -
attrition.corr()
# + [markdown] _cell_guid="112cef65-78b8-7790-e705-b173beea6986"
# # Feature Engineering & Categorical Encoding
#
# Task of Feature engineering and numerically encoding the categorical values in our dataset.
# +
# attrition.shape
# -
attrition.dtypes
# + _cell_guid="937385c7-7b7f-f6d0-d974-0527a7118e98"
# Empty list to store columns with categorical data
categorical = []
for col, value in attrition.iteritems():
if value.dtype == 'object':
categorical.append(col)
# Store the numerical columns in a list numerical
numerical = attrition.columns.difference(categorical)
# -
numerical
categorical
# + _cell_guid="5ec5cd49-f8b3-e36b-75dd-ac95fe0373ac"
# Store the categorical data in a dataframe called attrition_cat
attrition_cat = attrition[categorical]
attrition_cat = attrition_cat.drop(['Attrition'], axis=1) # Dropping the target column
# -
attrition_cat
# + [markdown] _cell_guid="7c3c0c95-3725-80dd-0a73-5c840451a438"
# Applying the **get_dummies** method
# + _cell_guid="7ea5b0d8-1f13-e56b-72cf-bcbe7dd6fad2"
attrition_cat = pd.get_dummies(attrition_cat)
attrition_cat.head(3)
# + _cell_guid="de8b3a57-6aba-eae7-2be3-dbe0ae761d6a"
# Store the numerical features to a dataframe attrition_num
attrition_num = attrition[numerical]
# + [markdown] _cell_guid="9de23a93-10b6-33b8-eea8-0cf44c6e5e08"
# let's concat numerical and caterogial dfs
# + _cell_guid="b90b69ba-f19d-0707-7c2c-183b8d01130f"
# Concat the two dataframes together columnwise
attrition_final = pd.concat([attrition_num, attrition_cat], axis=1)
# -
attrition_final.shape
attrition_final.head()
# + [markdown] _cell_guid="1a295568-fab4-b79a-bc0d-be32ad032b3e"
# **Target variable**
#
# The target in this case is given by the column **Attrition** which contains categorical variables therefore requires numerical encoding. We numerically encode it by creating a dictionary with the mapping given as 1 : Yes and 0 : No
# + _cell_guid="bfa5e82f-2dd3-1bee-5b2b-367468be7040"
# Define a dictionary for the target mapping
target_map = {'Yes':1, 'No':0}
# Use the pandas apply method to numerically encode our attrition target variable
target = attrition["Attrition"].apply(lambda x: target_map[x])
target.head(3)
# + [markdown] _cell_guid="5564e6e1-83ed-75de-2540-0d037e31291b"
#
# **Splitting Data into Train and Test sets**
#
# + _cell_guid="c197f8ee-76b0-7137-f001-83f969637521"
# Split data into train and test sets as well as for validation and testing
train, test, target_train, target_test = train_test_split(attrition_final, target, train_size= 0.75,random_state=0);
# -
# # Implementing Machine Learning Models
#
# + [markdown] _cell_guid="610cfa87-0b9d-4671-cd51-c99ef9c9151d"
# ## Xgboost Classifier
#
#
# -
# ### 1.n_estimators - No of Trees in the Model
#
# ### 2.max_leaf_nodes = The maximum number of terminal nodes or leaves in a tree. If this is defined, max_depth will be ignored
#
# ### 3.min_child_weight - Defines the minimum sum of weights of all observations required in a child.
#
# ### 4.max_depth - Maximum Depth of Tree and can be used to control overfiting
#
# ### 5.subsample- The fraction of samples to be used for fitting the individual base learners
#
# ### 6.learning_rate - Learning rate shrinks the contribution of each tree by learning_rate. There is a trade-off between learning_rate and n_estimators
#
# ## Regularization parameters
#
# ### 7. gamma - A node is split only when the resulting split gives a postive Gain. Gamma specifies the minimum loss reduction required to make a split.
#
# ### 8. lambda - This used to handle the regularization part of XGBoost. It should be explored to reduce overfitting
#
# ## Imbalanced Data Handling
#
# ### 9. scale_pos_weight - A value greater than 0 should be used in case of high class imbalance as it helps in faster convergence
xgb_cfl = xgb.XGBClassifier(n_jobs = -1,objective = 'binary:logistic')
xgb_cfl.get_params()
# + _cell_guid="ed6a837e-2864-291c-be8d-3c8e9ed900b7"
# Fit the model to our train and target
xgb_cfl.fit(train, target_train) # default
# Get our predictions
xgb_predictions = xgb_cfl.predict(test)
# -
xgb_predictions_prob = xgb_cfl.predict_proba(test)
xgb_predictions_prob
accuracy_score(target_test, xgb_predictions)
# # HPT - Random Search
# +
# A parameter grid for XGBoost
params = {
'n_estimators' : [100, 200, 500, 750], # no of trees
'learning_rate' : [0.01, 0.02, 0.05, 0.1, 0.25], # eta
'min_child_weight': [1, 5, 7, 10],
'gamma': [0.1, 0.5, 1, 1.5, 5],
'subsample': [0.6, 0.8, 1.0],
'colsample_bytree': [0.6, 0.8, 1.0],
'max_depth': [3, 4, 5, 10, 12]
}
folds = 5
param_comb = 800
random_search = RandomizedSearchCV(xgb_cfl, param_distributions=params, n_iter=param_comb, scoring='accuracy', n_jobs=-1, cv=5, verbose=3, random_state=42)
# -
random_search.fit(train, target_train)
print('\n Best estimator:')
print(random_search.best_estimator_)
print('\n Best accuracy for %d-fold search with %d parameter combinations:' % (folds, param_comb))
print(random_search.best_score_ )
print('\n Best hyperparameters:')
print(random_search.best_params_)
# + _cell_guid="40c37011-76df-fcc7-9cd0-e689374a8d1a"
xgb_predictions_hpt = random_search.predict(test)
accuracy_score(target_test, xgb_predictions_hpt)
# -
# #### Score here shows that HPT helped improving the Model accuracy
# + [markdown] _cell_guid="21cc0476-b03e-731f-97b4-89d81977c3a7"
# ### Feature Importance Xgboost Model
#
# -
xgb_cfl.feature_importances_
# + _cell_guid="082ca641-ffd2-fc3b-a7b6-9418b08767d9"
# Scatter plot
trace = go.Scatter(
y = xgb_cfl.feature_importances_,
x = attrition_final.columns.values,
mode='markers',
marker=dict(
sizemode = 'diameter',
sizeref = 1.3,
size = 12,
color = xgb_cfl.feature_importances_,
colorscale='Portland',
showscale=True
),
text = attrition_final.columns.values
)
data = [trace]
layout= go.Layout(
autosize= True,
title= 'XGBOOST Model Feature Importance',
hovermode= 'closest',
xaxis= dict(
ticklen= 5,
showgrid=False,
zeroline=False,
showline=False
),
yaxis=dict(
title= 'Feature Importance',
showgrid=False,
zeroline=False,
ticklen= 5,
gridwidth= 2
),
showlegend= False
)
fig = go.Figure(data=data, layout=layout)
py.iplot(fig,filename='scatter')
| 8. Machine Learning-2/6. Boosting/2. GBM/3. XG Boost/Xgboost-Classification+Example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # **Lab I : Signaux analogiques**
#
# + **Cours "Physique du Numérique"** - Portail René Descartes - AMU
#
# Préparé par :
#
# - <NAME> (v. 2021-08-31), Aix-Marseille Université © Contenus à diffusion restreinte, dans le cadre de ce cours.
#
# -------------------------
#
# ## I.A : Méthodologie
# #### Ce notebook Jupyter tient lieu à la fois d'énoncé et de compte-rendu de votre travail. A chaque TP correspond un notebook, que vous récupérez sur votre ordinateur local à partir des pages AMeTICE du cours. Au fur-et-à-mesure de votre travail, vous effectuez les exercices et remplissez les cellules avec du code suffisamment commenté, et les cellules Markdown (texte formatté) avec vos réponses aux questions, vos commentaires, sans oublier vos conclusions personnelles. A la fin de la séance de TP, pour pouvoir terminer le TP à la maison, vous récupérez votre notebook (d'extension *.ipynb*) soit par courriel, soit sur le nuage, ou sur clé USB.
#
# #### Pour travailler les notebooks Jupyter sur votre ordinateur, nous vous conseillons d'installer l'environnement de développement [Anaconda](https://www.anaconda.com/), utilisé aussi lors des TP en séance.
#
# #### Lorsque vous avez terminé le compte-rendu, avant la date limite de remise du travail, vous déposez une copie de votre notebook dans la boîte de dépôt (*dropbox*) dédiée des pages AMeTICE du cours, dans la section qui convient.
#
# -------------------------
#
# ## I.B : Python 3 et les notebooks Jupyter
# #### On présuppose de votre part une familiarité raisonnable avec l'usage de base du langage Python (version 3), que nous utiliserons généralement dans les cellules de code d'un notebook Jupyter. Si vous n'êtes pas à l'aise avec Python ou souhaitez revoir les principes de la programmation dans ce langage de plus en plus répandu, nous vous recommandons de travailler par vous-mêmes le [cours d'introduction à la programmation en Python](https://www.jdbonjour.ch/cours/python/introduction/) de <NAME> de l'EPFL, ou un [cours de Python de l'Université Paris Diderot](https://python.sdv.univ-paris-diderot.fr), ou le cours d'introduction à Python 3 pour le calcul scientifique de [l'université de Montpellier](https://courspython.com/bases-python.html)), ou encore de suivre un des très nombreux tutoriels disponibles sur l'Internet ([celui-ci](https://www.w3schools.com/python/default.asp) par exemple) pour vous remettre en mémoire les éléments de ce langage de programmation, que nous allons utiliser tout au long de ce cours, en particulier des TP.
#
# ##### Sur les pages AMeTICE du cours, vous trouverez une introduction aux notebook Jupyter, ainsi qu'un notebook d'apprentissage aux principales fonctionnalités d'un notebook Jupyter. Nous vous conseillons fortement de les parcourir avant d'avoir effectué ce TP.
#
# ------
#
# ## I.C : Enoncé du Lab I : Signaux analogiques
#
# ##### Outre une prise de contact avec l’environnement de programmation en Python 3 dans un notebook *Jupyter*, l’objectif de ce TP est de vérifier ou d'anticiper les résultats obtenus lors du TD 1.
#
# #### Remarque importante : Dans un notebook Jupyter, on construit séquentiellement l'environnement (bibliothèques de fonctions importées - exemples : *numpy*, *matplotlib.pyplot*, fonctions et variables définies par l'utilisateur) de programmation, il est important d'exécuter séquentiellement (de haut en bas) les différentes cellules de code. En effet, la cellule N peut utiliser des fonctions ou variables définies dans les cellules précédentes (N-1, N-2,...). Ainsi, la première cellule de code ci-dessous charge notamment les bibliothèques [numpy](https://courspython.com/apprendre-numpy.html) (permet l'usage des tableaux de nombres multi-dimensionnels appelés *arrays ou ndarrays*), et [matplotlib.pyplot](https://zestedesavoir.com/tutoriels/469/introduction-aux-graphiques-en-python-avec-matplotlib-pyplot/) (permet le tracé de graphes). Elle contient également la fonction *delta(t,t0)*, qui retourne une impulsion en $t=t_0$.
#
# +
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as widgets
# Disable warning messages to avoid distracting the beginners
import warnings
warnings.filterwarnings('ignore')
def delta(t,t0):
# Use np.arange to generate t in order to insure that t0 is present in t
dt=t[1]-t[0]
NN=t.shape
N=NN[0]
x=np.zeros(N,dtype=float)
ind=np.argwhere(abs(t-t0)<dt/2)
# print(ind)
indd=ind[0]
# print(indd)
x[indd]=1
return x
# -
# ## I.D : Signaux & vecteurs : Réalisez votre premier graphe avec Python 3
#
# Pour bien comprendre ce qui suit, nous vous conseillons de recopier (à la main, plutôt qu'un copier/coller afin de mieux mémoriser la syntaxe du langage) et d'exécuter les instructions dans la cellule de code ci-dessous au fur et à mesure de votre progression.
#
# Les signaux unidimensionnels numériques du type x[n] sont généralement représentés à l'aide d'un vecteur ligne ou d'un vecteur colonne. Un vecteur ligne n'est rien d'autre qu'un tableau de nombres ([array](https://www.w3schools.com/python/numpy/numpy_creating_arrays.asp) de l'extension [numpy](https://courspython.com/apprendre-numpy.html)), par exemple x, comprenant une ligne ou une colonne de N nombres séparés par une virgule, que l'on peut définir dans Python en les énumérant entre crochets, comme dans la cellule-code ci-dessous, où l'instruction `plt.stem(x)` permet de visualiser dans un graphe toutes les sept valeurs de ce vecteur-ligne :
x = np.array([-8,-5,-2.5,0.001,3.75,7,8.8])
plt.stem(x)
plt.show()
# où l'échelle des abscisses (horizontale) représente les indices des éléments du vecteur, en commençant par 1 jusqu'à N=7.
#
# Chaque élément du vecteur-ligne x est adressable séparément ; vous pouvez obtenir la valeur des premier et dernier éléments en exécutant dans la ligne de commande les instructions `print(x[0])` et `print(x[-1])`. La longueur du vecteur est obtenue via l'instruction `len(t)`.
#
# ### Exercice
#
# Analysez le script Python de la cellule ci-dessous et essayez de prévoir (avant son exécution) ou d'expliquer à posteriori les valeurs affichées :
t = np.arange(-1,1,0.01)
print(len(t),t[0],t[-1])
x = delta(t,0)
print(max(x))
print(np.argwhere(x==max(x)))
# #### Le vecteur support temporel
#
# La plupart du temps, les indices en abscisse ne sont pas très parlants. Supposons que les sept valeurs du vecteur représentent une quantité évaluée en t=0 et en des temps bien définis ; si l'on veut rendre compte du rythme d'acquisition de ces valeurs, il est pratique de créer un second vecteur, appelé **vecteur-support**, également composé de N=7 éléments, qui contiendra les instants d'échantillonnage, soit :
#
# `t = np.array([0,10,15,30,45,52,60])`
#
# dans la cellule ci-dessous, vous êtes maintenant inviter à visualiser les valeurs de x (en ordonnée) en fonction de celles de t (en abscisse), avec l'instruction stem ou plot :
#
# `plt.stem(t,x)` ou `plt.plot(t,x)`
#
# où la variable en abscisse apparaît en premier. Notez que les deux vecteurs concernés par le stem ou le plot doivent absolument comporter le même nombre d'éléments.
# Vous pouvez ensuite ajouter à votre graphe le titre et la description de chaque axe, avec les instructions :
#
# ```python
# plt.title('Mon premier graphe')
# plt.xlabel('temps (minute)')
# plt.ylabel('Variable x')
# ```
#
# L'instruction `plt.grid('on')` superpose un quadrillage à la figure, ce qui permet de mieux se rendre compte des variations relatives de la grandeur x.
#
# L'instruction `plt.show()` est normalement requise pour permettre l'affichage de la figure par Python (généralement superfétatoire dans un notebook Jupyter).
#
# **Remarque** : Les variables (vecteurs) x et t existent à présent dans l'environnement actuel, qui comprend l'ensemble des variables créées depuis l'ouverture de ce notebook, jusqu'à ce que vous le quittiez, que vous effaciez l'ensemble des variables à l'aide de l'instruction `%reset`, ou que vous effaciez sélectivement une variable particulière à l'aide de l'instruction `del x`, où x est la variable à supprimer de l'environnement. Vous pouvez visualiser toutes les variables de l'environnement avec l'instruction `%whos`, qui liste tous les objets (variables, fonctions, modules...) utilisés dans le notebook).
#
# Félicitations ! Vous venez de réaliser votre premier graphe avec Python, qui doit ressembler à peu de choses près à celui de la figure ci-dessous.
#
# <img src="https://amubox.univ-amu.fr/s/3zrqBq6B7meRkYN/preview" style="zoom:75%;" />
# A toutes fins utiles, les instructions nécessaires pour produire ce graphe sont reprises dans le listing ci-dessous. Des paramètres optionnels ont été rajoutés dans les instructions graphiques afin d'obtenir un graphe plus facilement lisible.
#
# ```python
# x=np.array([-8,-5,-2.5,0.01,3.75,7,8.8])
# t=np.array([0,10,15,30,45,52,60])
#
# plt.figure(1)
# plt.stem(x)
# plt.xlabel('indices des points')
# plt.ylabel('valeurs de x')
# plt.grid('on')
# plt.show()
#
# plt.figure(2)
# plt.stem(t,x)
# plt.xlabel("instants d'acquisition (min)")
# plt.ylabel('valeurs de x')
# plt.grid('on')
# plt.show()
# ```
# ## I.E Représentation des signaux numériques échantillonnés à période $T_e$ constante
#
# #### **Génération rapide du vecteur support**
#
# Les signaux unidimensionnels numériques résultent souvent de l'échantillonnage d'un signal analogique $x(t)$ à l'aide d'un convertisseur Analogique-Digital, qui réalise $x[n]=x(nTe)$ avec une fréquence d'échantillonnage $f_e=\frac{1}{T_e}$ généralement constante pour un signal donné. Heureusement, la génération du vecteur support ne nécessite pas d'écrire explicitement toutes les valeurs de $t_n = n T_e$ ! Deux instructions (`arange`et `linspace`) permettent de générer très simplement la suite des éléments du vecteur-support, régulièrement espacés, qu'on choisira selon la situation initiale :
#
# ##### De deux choses l'une :
# - Soit **on connaît le nombre total N d'échantillons du vecteur x**, ou que l'on peut le déterminer à partir de x (accessible via l'instruction `N=np.size(x)`) ainsi que les temps de début et de fin (ton et toff) lors de l'acquisition du vecteur x. Dans ce cas, il est généralement avantageux de générer le vecteur support avec l'instruction `t=np.linspace(ton,toff,N)`.
#
# - Soit **on connaît la période d'échantillonnage Te du vecteur x**, ainsi que les temps de début et de fin (ton et toff) du vecteur x. Dans ce cas, on pourra utiliser l'instruction `t = np.arange(ton,toff,Te)`. Cette instruction génère autant de points que nécessaire, séparés de Te, pour atteindre toff-Te.
#
# #### **Exemple : Représentation d'une portion d'un signal sinusoïdal**
#
# Avant de considérer la transmission des signaux, nous allons les considérer à un endroit donné, donc à une position de coordonnée x fixée, que nous prendrons égale à zéro par commodité. dans le reste de ce TP, nous allons donc considérer comme modèles de signaux les sinusoïdes décrites par l'une des deux expressions analytiques suivantes :
#
# > $ s(t) = A \ \cos (2\pi f_0 t + \phi )$ : Sinusoïde analogique
# >
# > $ s[n] = s(n \ t_e) = A \ \cos (2\pi f_0 n \ t_e + \phi )$ : Sinusoïde à temps discret (numérique)
#
# où cos(-) désigne la fonction cosinus bien connue de la trigonométrie. Pour définir un signal à temps continu, on utilise généralement une fonction dont la **variable indépendante** est **t, une variable réelle continue qui représente le temps**.
#
# Une sinusoïde analogique x(t) est une fonction mathématique dans laquelle l'angle (ou l'argument) de la fonction cosinusoïdale est, à son tour, une fonction de la variable t. Puisque nous considérons normalement que le temps augmente uniformément, l'angle de la fonction cosinus augmente également en proportion du temps. Les paramètres A, $f_0$, et ϕ sont des nombres fixes pour un signal cosinus particulier. Plus précisément, A est appelé l'amplitude, $f_0$ la fréquence, et ϕ la phase du signal (co)sinusoïdal. Il en va de même pour une sinusoïde à temps discret, qui se compose d'échantillons d'une sinusoïde analogique pris tous les multiples entiers de la période d'échantillonnage $t_e$.
#
# La figure ci-dessous montre le tracé (la trajectoire) (sur l'intervalle temporel [0,20 ms]) de la sinusoïde à temps continu
# $x(t) = 10 \cos(2\pi 440\ t - 0.4\pi)$,
# où $A = 10$, $f_0 = 440 Hz$, et $\phi = -0.4π$.
#
# 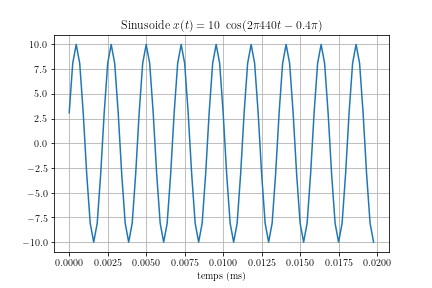
#
# Notez que x(t) oscille entre A et -A, et répète le même modèle d'oscillations tous les 1/440 = 0,00227 s (approximativement). Cet intervalle de temps est appelé la période de la sinusoïde.
#
#
# #### **Exercice**
#
# Dans la cellule de code ci-dessous, représentez sur l'intervalle $[-3 s , 5 s]$ le signal à temps continu sinusoïdal défini par $x(t)=3 cos(2\pi 0.5 t)$, où $A=3$, $f_0=0.5Hz$. On utilise pour cela une sinusoïde à temps discret, qui se compose d'échantillons de la sinusoïde à temps continu $x(t)$ pris tous les multiples entiers de la période d'échantillonnage $t_e$. On commence donc par définir un vecteur support temporel comprenant N éléments.
#
# On choisira N suffisamment grand pour représenter le signal continu avec suffisamment de détails, c'est-à-dire avec suffisamment de points par période de la sinusoïde, qui ici vaut $T_0=\frac{1}{f_0}=2 s$. La section du signal représentée dans le graphe, égale à la durée de la section représentée divisée par la période, sera donc ici de $\frac{5-(-3)}{2}=4$ périodes ; en choisissant 32 points par période, on pourra fixer $N=4 \ 32 = 128$ points. Le vecteur support temporel désigné par la variable t est donc créé avec l'instruction `t=np.linspace(-3,5,128)`.
#
# On crée ensuite la variable x à partir du vecteur support t, dont les dimensions seront automatiquement reproduites (on dit que *x hérite* des caractéristiques de *t*), par l'instruction `x = 3*np.cos(2*np.pi*0.5*t)`.
#
# Le tracé de la section du signal continu x(t) sur l'intervalle $[-3 ;5]$ s'obtient simplement par l'instruction `np.plot(t,x)`.
# ## I.F Usage des sinusoïdes discrètes pour représenter les sinusoïdes à temps continu
#
# Les sinusoïdes (pseudo-)analogiques qui apparaissent dans le graphe ci-dessus sont en réalité des sinusoïdes à temps discret, avec une fréquence d'échantillonnage suffisamment élevée pour qu'elles apparaissent à l'écran comme continues lorsqu'on relie leurs échantillons adjacents par des segments de droite (c'est ce que fait l'instruction `plt.plot(t,x)`).
#
# 1. Pour faire apparaître les échantillons de la sinusoïde, remplacez l'instruction `plt.plot(t,x)` par `plt.plot(t,x,'.')`.
#
# 2. Pour superposer la sinusoïde à temps discret avec la sinusoïde analogique qu'elle réprésente, utilisez l'instruction `plt.plot(t,x,'.',t,x)`.
#
# 3. Pour afficher la sinusoïde à temps discret en fonction de l'indice des échantillons, utilisez l'instruction `plt.plot(x,'.')`.
#
# 2. Transformez le programme ci-dessus pour afficher la sinusoïde avec un axe des abscisses (l'axe horizontal) gradué en ms. Il vous faudra également remplacer l'instruction `plt.xlabel('temps (ms)')` par `plt.xlabel('Indice échantillon')`.
#
# La simulation de la cellule ci-dessous utilise une case à cocher pour afficher la sinusoïde soit uniquement sous son aspect analogique, en fonction du temps, soit son véritable aspect, numérique, en fonction du numéro d'indice, superposée à la sinusoïde analogique qu'elle représente. Cet exemple vous montre comment vous pouvez ajouter dans vos scripts Python des éléments interactifs (des objets html) comme une case à cocher, à l'aide de la bibliothèque [ìpwidgets](https://www.tutorialspoint.com/jupyter/jupyter_notebook_ipywidgets.htm) de Jupyter.
# +
# Définition des paramètres de la sinusoïde
A=10
f0=440
phi=-0.4*np.pi
# définition des autres paramètres
T0=0.02
fe=10*f0 # fréquence d'échantillonnage conforme au critère de Nyquist
te=1/fe
# Définition du vecteur support temporel
t=np.arange(0,T0,1/fe)
# Calcul des valeurs prises par la sinusoïde
x=A*np.cos(2*np.pi*f0*t+phi)
# Construction de la figure
def f(num):
if num:
plt.plot(x,'.r')
plt.plot(x,'b')
plt.xlabel('Index number')
else:
plt.plot(t,x,'b')
plt.xlabel('temps (ms)')
plt.title('Sinusoide')
plt.show()
numW=widgets.Checkbox(value=False,description="Show samples")
widgets.interact(f,num=numW)
# -
# ## I.G : Exercices
#
# ##### **Remarque importante** : Vous n'aurez probablement pas le temps d'effectuer l'ensemble de ces exercices lors de la séance de TP en présentiel. Nous vous conseillons pourtant vivement d'essayer de les résoudre par vous-même, car ils sont très instructifs, et qu'ils inspireront certainement les questions de l'examen de TP terminal.
#
# ### Exercice I.0
#
# 1. Dans la cellule de code ci-dessous, écrivez une fonction Python `mycos(A,f,phi,t)`qui calcule et retourne les valeurs d’une cosinusoïde discrète d’expression analytique $x(t)=A \ \cos (2\pi f_0t+\phi)$ à partir des trois paramètres $(A,f_0,\phi)$ et d’un vecteur support temporel $t$ défini comme un *[array](https://www.w3schools.com/python/numpy/numpy_creating_arrays.asp)*. Ces quatre variables seront fournies en paramètre de la fonction.
#
# 2. Dans la cellule ci-dessous, écrivez une fonction Python qui calcule et retourne les valeurs de la fonction rectangulaire normalisée $rect(t)$ à partir d’un vecteur support temporel $t$ défini comme un *[array](https://www.w3schools.com/python/numpy/numpy_creating_arrays.asp)*.
#
# ### Exercice I.1
#
# #### Aux TD
#
# 1. Déterminez les paramètres $(A,f_0,\phi)$ du signal cosinusoïdal $x(t)$ représenté ci-dessous.
#
# <img src="https://amubox.univ-amu.fr/s/nmRzJEDL3wFWaqS/preview" style="zoom:75%;" />
#
# 2. Tracez schématiquement le spectre unilatéral (module et phase) de ce signal cosinusoïdal.
#
# #### Aux TP
#
# dans la cellule de code ci-dessous, écrivez un script Python pour reproduire le graphe du signal d'après l'expression analytique déterminée plus haut sur l'intervalle $[-15ms,20ms]$, et vérifiez que la sinusoïde tracée ainsi correspond bien à celle de la figure ci-dessus.
# ### Exercice I.2
#
# #### Aux TD
#
# Représentez dans un graphe le signal x(t) décrit par l'expression analytique $x(t)=(2-2t)rect(\frac{t}{2})$.
# A partir du graphe, déterminez la valeur de $x(0)$ ainsi que l'aire sous la courbe.
# Retrouvez ces résultats à partir de l'expression analytique de $x(t)$ et d'une intégrale définie (pour l'aire).
#
# #### Aux TP
#
# Ecrivez un script Python pour reproduire le graphe de $x(t)$ d'après son expression analytique sur l'intervalle qui convient, et calculez numériquement $x(0)$ ainsi que l’aire sous la courbe.
# ### Exercice I.3
#
# #### Aux TD
#
# Représentez le signal $x(t)$ décrit par l'expression analytique suivante : $x(t)=2 rect(2(t-2))$.
# A partir du graphe, déterminez la valeur de $x(0)$ ainsi que l'aire sous la courbe.
# Retrouvez ces résultats à partir de l'expression analytique et d'une intégrale définie (pour l'aire).
#
# **Indication** : Pour tracer le signal en réduisant le risque d'erreurs, on aura intérêt à transformer l'expression de $x(t)$ sous la forme normalisée $x(t)=A \ rect \left( \frac{t-t_0}{T} \right)$.
#
# #### Aux TP
#
# Ecrivez un script Python pour reproduire le graphe de $x(t)$ d'après son expression analytique sur l'intervalle qui convient, et calculez numériquement $x(0)$ ainsi que l’aire sous la courbe.
# ### Exercice I.4
#
# #### Aux TD
#
# On retarde de 2 secondes le signal $x(t)= rect\left( \frac{t}{2}\right)$ pour obtenir un nouveau signal $y(t)$.
#
# - Le signal retardé est-il décalé vers la gauche ou vers la droite ?
# - Sur quelle valeur de $t$ le signal $y(t)$ est-il centré ?
# - Représentez ces deux signaux sur un graphe.
# - Donnez l'expression analytique du signal retardé en fonction de $x(t)$.
#
# #### Aux TP
#
# Ecrivez un script Python pour reproduire les graphes de $x(t)$ et $y(t)$ d'après leur expression analytique sur l'intervalle qui convient.
# ### Exercice I.5
#
# #### Aux TD
#
# Donnez l'expression analytique du signal $y(t)$ dont le graphe est donné dans la figure ci-dessous :
#
# 1. en fonction du signal $x(t)$,
# 2. en fonction du signal $rect(t)$.
#
# <img src="https://amubox.univ-amu.fr/s/fgo4FnsnqoXQJ6d/preview" style="zoom:100%;" />
#
# #### Aux TP
#
# Dans la cellule ci-dessous, écrivez un script Python pour reproduire les graphes de $x(t)$ et $y(t)$ d'après leur expression analytique sur l'intervalle qui convient.
# ### Exercice I.6
#
# #### Aux TD
#
# Ecrivez l'expression analytique du signal x(t) représenté ci-dessous en termes du signal $rect(t)$, de deux manières différentes.
#
# <img src="https://amubox.univ-amu.fr/s/9W7NeHgG6BjF5fH/preview" style="zoom:100%;" />
#
# #### Aux TP
#
# Dans la cellule ci-dessous, en prenant $T_0=5$ et $b=3$, écrivez un script Python pour vérifier l’expression analytique obtenue et reproduire la figure ci-dessus.
# ### Exercice I.7
#
# #### Aux TD
#
# Ecrivez l'expression analytique des six signaux $s_n(t)$ représenté ci-dessous en termes du signal $rect(t)$.
#
# <img src="https://amubox.univ-amu.fr/s/2eyzYfiqXjq7Wma/preview" style="zoom:50%;" />
#
# #### Aux TP
#
# Dans la cellule ci-dessous, écrivez un script Python pour reproduire sur l'intervalle $[-4,4]$ les graphes de ces six signaux $s_n(t)$ d’après l’expression analytique obtenue.
# ### Exercice I.8
#
# #### Aux TD
#
# Ecrivez l'expression du signal $x(t)=rect(t/2)$ en termes du signal "échelon-unité" $u(t)$.
#
# #### Aux TP
#
# Vérifiez cette expression en écrivant un script Python pour reproduire le graphe de $x(t)$.
# ### Exercice I.9
#
# #### Aux TD
#
# Ecrivez les 5 signaux ci-dessous sous la forme générique d'une cosinusoïde $x(t)=A\ \cos(2\pi f_0 t + \phi)$, en précisant dans chaque cas la valeur des paramètres $(A,f_0,\phi)$.
#
# 1. $a(t)=2 \sin (\pi t)$
# 2. $b(t)= 3 \cos (2t)$
# 3. $c(t)=10 \sin(2\pi t+\pi/4)$
# 4. $d(t)= \cos (\pi t + \pi /2)$
# 5. $e(t)= 5 \sin (\pi (t-1))$
# 6. $f(t)= \sin(-t+1)$
#
# **Indication** : On rappelle que le signal $\sin(t)$ est une cosinusoïde retardée de $\frac{\pi}{2}$, *i.e.* $\sin (t) = \cos (t-\frac{\pi}{2})$.
#
# #### Aux TP
#
# Utilisez Python dans la cellule de code ci-dessous pour générer le graphe de chacun de ces six signaux sinusoïdaux sur l'intervalle $[-5,5]$ en vérifiant l'égalité entre les deux expressions de chaque signal.
# ### Exercice I.10
#
# #### Aux TD
#
# 1. Un signal analogique sous forme d'une tension électrique $e(t)$ est amenée à l'entrée d'un amplificateur. Elle en resort sans distorsion après un délais de $1 ms$, amplifiée d'un facteur 10. Ecrivez l'expression analytique du signal de sortie $s_1(t)$ en fonction de $e(t)$.
#
# 2. Un signal analogique sous forme d'une tension électrique $v(t)$ est une cosinusoïde d'amplitude $5V$ et de fréquence $2 kHz$ (on qualifie cette tension *d'alternative*). Le signal voyage à travers un réseau de communication et en émerge après $100\mu s$ sans distorsion notable, bien qu'avec une amplitude atténuée d'un facteur 3. Ecrivez l'expression analytique du signal de sortie $s_2(t)$ en fonction de $v(t)$.
#
# 3. Calculez le déphasage $\phi$ (en radians) du signal de sortie $s_2(t)$ par rapport au signal d'entrée $v(t)$.
#
#
# #### Aux TP
#
# Dans la cellule ci-dessous, tracez à l'aide de Python le graphe des signaux d'entrée et de sortie $v(t)$ et $s_2(t)$, et vérifiez graphiquement la valeur du déphasage.
# ### Exercice I.11
#
# #### Aux TD
#
# On considère les signaux $a(t)$, $b(t)$, $c(t)$ et $d(t)$ représentés ci-dessous.
#
# <img src="https://amubox.univ-amu.fr/s/jnt43r2i5s2FEr9/preview" style="zoom:50%;" />
#
# 1. Exprimez $c(t)$ en termes de $a(t)$ et $b(t)$.
# 2. Exprimez $d(t)$ en termes de $a(t)$ et $b(t)$.
# 3. Pour chacun des quatre signaux, donnez une expression analytique composée d'un signal $rect(t)$ et/ou une cosinusoïde $A \ \cos (2 \pi f_0 t + \phi)$.
#
# #### Aux TP
#
# A l’aide de Python, dans la cellule-code ci-dessous, vérifiez la validité des expressions trouvées en (3) en traçant le graphe des quatre signaux à partir de leur expression analytique. Vérifiez que la figure obtenue est semblable à la figure ci-dessus.
# ### Exercice I.12
#
# #### Aux TD
#
# Un système de traitement des signaux produit un signal de sortie $s(t)$ égal au signal d'entrée $e(t)$ amplifié d'un facteur 4 et retardé de 1 seconde, affecté d'un double écho constitué d'une première réplique du signal d'entrée décalée de 2s par rapport à celui-ci, et d'une seconde réplique décalée de 4s. En supposant que le signal d'entrée est $e(t)=rect \left( \frac{t}{4} \right)$, donnez l'expression analytique du signal de sortie $s(t)$ en fonction du signal d'entrée $e(t)$.
#
# #### Aux TP
#
# A l’aide de Python, dans la cellule ci-dessous, représentez ces deux signaux.
# ------
#
# ## I.H. Conclusions personnelles
#
# Indiquez ci-dessous le temps approximatif que vous passé à travailler ce TP en-dehors de la séance.
#
# > Votre réponse ici :
#
# Ecrivez ci-dessous, en guise de conclusions, quelques phrases décrivant ce que ce TP vous a appris.
#
# > **Conclusions personnelles** (votre réponse ici) :
# ------
# ------
| Enonce_Lab_1_Signaux.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.4 64-bit (''base'': conda)'
# name: python3
# ---
import cv2
vidcap = cv2.VideoCapture('./Data/inazuma_amakumo_peak.mp4')
success,image = vidcap.read()
count = 0
while success:
cv2.imwrite("./Data/amakumo_peak/frame%d.jpg" % count, image) # save frame as JPEG file
success,image = vidcap.read()
print('Read a new frame: ', success)
count += 1
| dataconversion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This is how we automatically aligned the entire EGOCOM dataset - no human aid necessary.
# +
from __future__ import print_function, absolute_import, division, unicode_literals, with_statement # Python 2 compatibility
import os
import numpy as np
import subprocess
# +
# Read in the data
data_loc = "/home/cgn/Downloads/egocom-aligned/"
write_loc = "/home/cgn/Downloads/egocom-aligned-final/"
inter_loc = "/home/cgn/Downloads/egocom-intermediate/"
alignments_fn = "/home/cgn/Downloads/egocom-aligned-final/ALIGNMENT_ALL.txt"
fn_dict = {}
for fn in sorted([v for v in os.listdir(data_loc) if v[-4:] == ".MP4"]):
key = fn[9:23] + fn[32:37] if 'part' in fn else fn[9:21]
fn_dict[key] = fn_dict[key] + [fn] if key in fn_dict else [fn]
with open(alignments_fn, mode='r') as f:
alignments = [z.strip() for z in f.readlines()]
# -
samplerate = 44100
for alignment in alignments:
key, lst = alignment.split(" | ")
# Get alignment array
alignment = (np.array(eval(lst)) / samplerate).round(4)
files = [data_loc + z for z in fn_dict[key]]
inter_files = [inter_loc + z for z in fn_dict[key]]
output_files = [write_loc + z for z in fn_dict[key]]
# Get the duration of each file
d = []
for file in files:
cmd = "ffprobe -i {f} -show_format | grep duration".format(f=file)
d.append(subprocess.getoutput(cmd).split("=")[-1])
duration = np.array(d, dtype=float)
print("Duration of original videos:", duration)
# Find the shortest duration (minus the length we cut from the start)
min_duration = min(duration - alignment)
# Use the shortest duration to find the final duration for all
# videos so that they are the same length.
final_duration = min_duration + alignment
for i in range(len(alignment)):
cmd = "ffmpeg -ss {s} -t {t} -i {i} -c copy {o}".format(
s=alignment[i],
t=min_duration,
i=files[i],
o=output_files[i],
)
print(cmd)
subprocess.getoutput(cmd)
| paper_experiments_work_log/egocom_dataset_creation/auto_align_egocom_dataset.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
from enum import Enum
from collections import defaultdict
import pprint as pp
# 1. Implement the lion and cow example on a 10x10 grid (the lion's
# starting position is at the bottom left square, and the cow is in the
# top right square).
class Action(Enum):
STAY = 0
NORTH = 1
SOUTH = 2
EAST = 3
WEST = 4
# +
reward = {
(1, Action.WEST) : 0,
(2, Action.WEST) : 0,
(2, Action.NORTH) : 0,
(2, Action.EAST) : 0,
(3, Action.NORTH) : 50,
(4, Action.NORTH) : 50,
(5, Action.NORTH) : 0,
(5, Action.EAST) : 0,
(6, Action.STAY) : 0,
(7, Action.EAST) : 100
}
transition = {
(1, Action.WEST) : 2,
(2, Action.WEST) : 3,
(2, Action.NORTH) : 4,
(2, Action.EAST) : 1,
(3, Action.NORTH) : 5,
(4, Action.NORTH) : 6,
(5, Action.NORTH) : 7,
(5, Action.EAST) : 4,
(6, Action.STAY) : 6,
(7, Action.EAST) : 6
}
state_reward = defaultdict(int)
state_reward[5] = 50
state_reward[6] = 100
N_STATES = 7
# -
states = list(range(1, 8))
states
actions = {
1: [Action.WEST],
2: [Action.WEST, Action.NORTH, Action.EAST],
3: [Action.NORTH],
4: [Action.NORTH],
5: [Action.NORTH, Action.EAST],
6: [Action.STAY],
7: [Action.EAST]
}
# ### a) Compute the V* values for each state with discount factor gamma = 0.8.
def value_iteration(gamma, theta):
values = {}
for s in states:
values[s] = 0
niter = 0
while True:
niter += 1
delta = 0
for state in states:
v = values[state]
candidates = [r + gamma * values[ss]
for a in actions[state]
for ss in [transition[(state, a)]] for r in [reward[(state, a)]]]
values[state] = max(candidates)
delta = max(delta, abs(v - values[state]))
if delta < theta:
return values
values = value_iteration(gamma=0.8, theta=1)
print('{0: <10} {1: <6}'.format('state', 'V*'))
for s in states:
print("{0: <10} {1: <5}".format(s, values[s]))
# ### b) What is the optimal policy when gamma = 0.8?
def get_policy(values):
policy = {}
for s in states:
pairs = [(values[transition[s,a]], a) for a in actions[s]]
policy[s] = sorted(pairs)[0][1]
return policy
values = value_iteration(gamma=0.8, theta=1)
get_policy(values)
# ### c) Does the optimal policy change if gamma is set to 0.5 instead? If yes, give the new policy. If not, explain.
values = value_iteration(gamma=0.5, theta=1)
get_policy(values)
# ### d) Compute the Q(s,a) values for the following state action pairs: (S2,West), (S6,Stay), (S3, North). Let gamma = 0.8 and alpha = 1.
def Q_model_based(V_star, state, action, gamma, reward=reward, transition=transition):
return reward[state, action] + gamma * V_star[transition[state, action]]
values = value_iteration(gamma=0.8, theta=1)
print('{: <10} {: <15}'.format('state', 'action'))
for s in states:
print("{0: <10} {1: <5}".format(s, values[s]))
# +
print('{: <10} {: <15} {: <3}'.format('state', 'action', 'Q'))
for state, act in [(2, Action.WEST), (6, Action.STAY), (3, Action.NORTH)]:
print('{: <10} {: <15} {}'.format(state, act, Q_model_based(values, state, act, gamma=0.8)))
# -
# ### e) Consider applying the Q-learning algorithm to the "treasure-hunting" game. Let Q' be the estimate of Q. Initially all Q' values are set to 0, and gamma = 0.8 and alpha = 1. Assume that the agent moves from state S1, via states S2, S3, S5, and S7, to state S6. Show how the Q' values are updated during this episode. Repeat the same episode twice more and show how the Q' values are revised during each episode.
def init_QQ():
QQ = {}
for state in states:
for act in actions[state]:
QQ[(state, act)] = 0
return QQ
def one_run(QQ, s, moves, gamma=0.8):
for move in moves:
print("s", s)
r = reward[(s, move)]
print('action {}'.format(move))
print('reward: {}'.format(r))
ss = transition[s, move]
print('new_state: {}'.format(ss))
QQ[(s, move)] = r + gamma * max([QQ[(ss, a)] for a in actions[ss]])
pp.PrettyPrinter(indent=4).pprint(QQ)
print()
s = ss
return QQ, s
QQ = init_QQ()
start_state = 1
moves = [Action.WEST, Action.WEST, Action.NORTH, Action.NORTH, Action.EAST]
QQ2, s2 = one_run(QQ, start_state, moves)
QQ3, s3 = one_run(QQ2, start_state, moves)
| hw1/hw1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # CLI - Command-line interface
#
# Let's take a look at the `kissim` command-line interface (CLI) to encode a set of structures (from the [KLIFS](https://klifs.net/) database) and perform an all-against-all comparison. The CLI follows the same logic as the quick start Python interface as described in "API - Quick start Python interface".
#
# 
# +
from pathlib import Path
import pandas as pd
# -
# Path to this notebook
HERE = Path(_dh[-1]) # noqa: F821
# ## Encode structures into fingerprints
# + language="bash"
# kissim encode -h
# # flake8-noqa-cell
# -
# ### `kissim encode` command
# + language="bash"
# # Set path to local KLIFS data
# PATH_KLIFS_DOWNLOAD="../../kissim/tests/data/KLIFS_download/"
# kissim encode -i 109 118 12347 1641 3833 9122 -o fingerprints.json -l $PATH_KLIFS_DOWNLOAD -c 2
# # flake8-noqa-cell
# -
# This command generate two files:
#
# - Data: `fingerprints.json`
# - Logs (not under Windows): `fingerprint.log`
# ### Inspect output: `FingerprintGenerator`
#
# You can load the content of the `fingerprints.json` file as `FingerprintGenerator` object.
fingerprints_path = HERE / "fingerprints.json"
fingerprints_path
# +
from kissim.encoding import FingerprintGenerator
fingerprint_generator = FingerprintGenerator.from_json(fingerprints_path)
print(f"Number of fingerprints: {len(fingerprint_generator.data.keys())}")
fingerprint_generator
# -
# ## Compare fingerprints
# + language="bash"
# kissim compare -h
# # flake8-noqa-cell
# -
# ### `kissim compare` command
# + language="bash"
# kissim compare -i fingerprints.json -o . -c 2
# # flake8-noqa-cell
# -
# This command generate the following files:
#
# - Data - fingerprint distances: `fingerprint_distances.csv`
# - Data - feature distances: `feature_distances.csv`
# - Data - default kinase matrix based on minimum fingerprint pair distance per kinase pair: `fingerprint_distances_to_kinase_matrix.csv`
# - Data - default kinase tree based on kinase matrix clustering (hierarchical clustering using Ward's method): `fingerprint_distances_to_kinase_clusters.csv`
# - Logs (not under Windows): `distances.log`
# ### Inspect output: All-against-all fingerprint distances
#
# You can load the content of the `fingerprint_distances.csv` file as `FingerprintDistancesGenerator` object.
# +
from kissim.comparison import FingerprintDistanceGenerator
fingerprint_distance_path = HERE / "fingerprint_distances.csv"
fingerprint_distance_generator = FingerprintDistanceGenerator.from_csv(fingerprint_distance_path)
print(f"Number of pairwise comparisons: {len(fingerprint_distance_generator.data)}")
fingerprint_distance_generator
# -
fingerprint_distance_generator.data
# ### Inspect output: All-against-all feature distances
#
# You can load the content of the `feature_distances.csv` file as `FeatureDistancesGenerator` object.
# +
from kissim.comparison import FeatureDistancesGenerator
feature_distances_path = HERE / "feature_distances.csv"
feature_distances_generator = FeatureDistancesGenerator.from_csv(feature_distances_path)
print(f"Number of pairwise comparisons: {len(feature_distances_generator.data)}")
feature_distances_generator
# -
feature_distances_generator.data
# Clean up output files.
# ### Inspect output: Kinase matrix
kinase_matrix = pd.read_csv(HERE / "fingerprint_distances_to_kinase_matrix.csv", index_col=0)
kinase_matrix
# ### Inspect output: Kinome tree and kinase annotation
# Kinome tree
with open(HERE / "fingerprint_distances_to_kinase_clusters.tree", "r") as f:
newick_string = f.read()
print(newick_string)
# Kinase annotation
kinase_annotations = pd.read_csv(HERE / "kinase_annotation.csv", index_col=0, sep="\t")
kinase_annotations
# The Newick tree and kinase annotation files can be loaded into the external software [FigTree](https://kissim.readthedocs.io/en/latest/tutorials/tree.html#Visualize-tree) to visualize the `kissim`-based kinome tree.
# ## Delete output files
[i.unlink() for i in HERE.glob("*.json")]
[i.unlink() for i in HERE.glob("*.csv")]
[i.unlink() for i in HERE.glob("*.tree")]
[i.unlink() for i in HERE.glob("*.log")];
| docs/tutorials/cli.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import re
import numpy as np
import json
import datetime
df_movies = pd.read_csv("../data/movies.csv")
df_movies
# +
def extract_movie_title(x):
split_list = re.split('[()]', x)
movie_title = split_list[0]
movie_year = split_list[len(split_list)-2]
return movie_title
def extract_movie_year(x):
split_list = re.split('[()]', x)
movie_title = split_list[0]
movie_year = split_list[len(split_list)-2]
return movie_year
def extract_movie_genres(x):
genre_list = re.split('[|]', x)
return genre_list
# -
df_movies['movie_title']=df_movies['title'].apply(lambda x: extract_movie_title(x))
df_movies['movie_year']=df_movies['title'].apply(lambda x: extract_movie_year(x))
df_movies['movie_genre']=df_movies['genres'].apply(lambda x: extract_movie_genres(x))
df_movies
genre_1 = "Drama"
df_movie_in_genre = df_movies[df_movies['movie_genre'].apply(lambda x: genre_1 in x)]
df_movie_in_genre.shape[0]
df_movie_in_genre
df_links = pd.read_csv("../data/links.csv")
df_links
df_movie_abt_1 = pd.merge(df_movies, df_links,
on=['movieId'],
suffixes=('', '_DROP'),
how='outer').filter(regex='^(?!.*_DROP)')
df_movie_abt_1
df_tmdb = pd.read_csv("../data/tmdb_movies_data.csv")
df_tmdb
# +
df_movie_abt_2 = pd.merge(df_movie_abt_1, df_tmdb,
left_on='tmdbId',
right_on='id',
suffixes=('', '_TMDB'),
how='outer')
df_movie_abt_2.loc[df_movie_abt_2["movie_title"].isnull(),'movie_title'] = df_movie_abt_2["original_title"]
df_movie_abt_2
# -
df_imdb = pd.read_csv("../data/imdb_attributes.csv")
df_imdb
df_movie_abt_3 = pd.merge(df_movie_abt_2, df_imdb,
left_on='imdbId',
right_on='tid',
suffixes=('', '_IMDB'),
how='outer')
df_movie_abt_3
movie_json_list = json.loads(df_movie_abt_3.to_json(orient='records'))
movie_list = []
movie_json_list = json.loads(df_movie_abt_3.to_json(orient='records'))
for i in range(len(movie_json_list)):
if movie_json_list[i]['movie_title'] is None:
if movie_json_list[i]['original_title'] is not None:
movie_json_list[i]['movie_title'] = extract_movie_title(movie_json_list[i]['original_title'])
if movie_json_list[i]['movie_title'] is None:
if movie_json_list[i]['title_IMDB'] is not None:
movie_json_list[i]['movie_title'] = extract_movie_title(movie_json_list[i]['title_IMDB'])
if movie_json_list[i]['movie_year'] is None:
if movie_json_list[i]['release_date'] is not None:
movie_json_list[i]['movie_year'] = movie_json_list[i]['release_year']
if movie_json_list[i]['movie_year'] is None:
if movie_json_list[i]['year'] is not None:
movie_json_list[i]['movie_year'] = extract_movie_year(movie_json_list[i]['year'])
if movie_json_list[i]['movie_title'] not in movie_list:
movie_list.append(movie_json_list[i]['movie_title'])
len(movie_list)
df_movie_abt = pd.DataFrame.from_dict(movie_json_list, orient = 'columns')
df_movie_abt.to_csv("../data/movie_abt.csv")
df_movie_list = pd.DataFrame.from_dict(movie_list, orient='columns')
df_movie_list
df_movie_list.to_csv('../data/movie_list.csv')
| notebooks/merge_dbs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.2
# language: julia
# name: julia-1.6
# ---
# ## Strands for Phase 2, release 0.03 (2.03)
#
# 600 strands in total
#
# This release is identical to 2.02 but with a one week delay.
strengthArray = [0.08, 0.16, 0.48]
radiusArray = [30, 35, 40, 45, 50, 60, 70, 80, 90, 100]
numRepeats_SIR_SEIR = 4;
numRepeats_SI_SEI = 2;
function createRepeats(parentStrand::Dict{Symbol,String}, num)
strands = Array{Dict{Symbol,String}}(undef,0)
for s in strengthArray, r in radiusArray, n in 1:num
strand = deepcopy(parentStrand)
strand[:name] = strand[:name]*"-($s,$r)-repeat$n"
strand[:strength] = "$s"
strand[:radius] = "$r"
push!(strands,strand)
end
strands
end;
baseStrand = Dict{Symbol,String}()
baseStrand[:seedTime] = "2021-08-25T21:00:00Z" #9am on Thursday morning Aug 26 (in Auckland)
baseStrand[:endTime] = "2021-11-04T20:00:00Z"#9pm in the evening of Nov 4 (in Auckland)
baseStrand[:minimumAppVersion] = "60";
### SEI epidemics
SEIstrand = deepcopy(baseStrand)
SEIstrand[:name] = "2.03-SEI"
SEIstrand[:seedingProbability] = "0.1"
SEIstrand[:incubationMean] = "172800" #default for two days incubation
SEIstrand[:incubationShape] = "3"
SEIstrand[:infectionMean] = "31536000" #365 days
SEIstrand[:infectiousPeriodShape] = "10000";
SEIstrand
##SI
SIstrand = deepcopy(SEIstrand)
SIstrand[:name] = "2.03-SI"
SIstrand[:incubationMean] = "1" #1 second incubation
SIstrand[:incubationShape] = "10000"
SIstrand
#The base strand for the SEIR epidemics
SEIRstrand = deepcopy(baseStrand)
SEIRstrand[:name] = "2.03-SEIR"
SEIRstrand[:seedingProbability] = "0.1"
SEIRstrand[:incubationMean] = "172800"
SEIRstrand[:incubationShape] = "3"
SEIRstrand[:infectionMean] = "432000" #5 days should vary
SEIRstrand[:infectiousPeriodShape] = "3";
SEIRstrand
##SIR base strand
SIRstrand = deepcopy(SEIRstrand)
SIRstrand[:name] = "2.03-SIR"
SIRstrand[:incubationMean] = "1" #1 second incubation
SIRstrand[:incubationShape] = "10000"
SIRstrand
SEIRstrandShortDuration = deepcopy(SEIRstrand)
SEIRstrandShortDuration[:name] = SEIRstrandShortDuration[:name]*"-days5";
SEIRstrandShortDuration
SEIRstrandLongDuration = deepcopy(SEIRstrand)
SEIRstrandLongDuration[:infectionMean] = "864000" #10 days
SEIRstrandLongDuration[:name] = SEIRstrandLongDuration[:name]*"-days10";
SEIRstrandLongDuration
SIRstrandShortDuration = deepcopy(SIRstrand)
SIRstrandShortDuration[:name] = SIRstrandShortDuration[:name]*"-days5";
SIRstrandShortDuration
SIRstrandLongDuration = deepcopy(SIRstrand)
SIRstrandLongDuration[:infectionMean] = "864000" #10 days
SIRstrandLongDuration[:name] = SIRstrandLongDuration[:name]*"-days10";
SIRstrandLongDuration
#An array that will hold all strands
strands = Array{Dict{Symbol,String}}(undef,0)
append!(strands, createRepeats(SIstrand, numRepeats_SI_SEI))
append!(strands, createRepeats(SEIstrand, numRepeats_SI_SEI))
append!(strands, createRepeats(SEIRstrandShortDuration, numRepeats_SIR_SEIR))
append!(strands, createRepeats(SEIRstrandLongDuration, numRepeats_SIR_SEIR))
append!(strands, createRepeats(SIRstrandShortDuration, numRepeats_SIR_SEIR))
append!(strands, createRepeats(SIRstrandLongDuration, numRepeats_SIR_SEIR));
length(strands)
# +
function strandJSON(sd::Dict{Symbol,String})
str = """
{\"name\":\"$(sd[:name])\",
\"seedingProbability\":\"$(sd[:seedingProbability])\",
\"infectionProbabilityMapP\":\"$(sd[:strength])\",
\"infectionProbabilityMapK\":\"$(sd[:radius])\",
\"infectionProbabilityMapL\":\"1000000\",
\"incubationPeriodMeanSec\":\"$(sd[:incubationMean])\",
\"incubationPeriodShape\":\"$(sd[:incubationShape])\",
\"infectiousPeriodMeanSec\":\"$(sd[:infectionMean])\",
\"infectiousPeriodShape\":\"$(sd[:infectiousPeriodShape])\",
\"startTime\":\"$(sd[:seedTime])\",
\"endTime\":\"$(sd[:endTime])\",
\"minimumAppVersion\":\"$(sd[:minimumAppVersion])\"}
"""
join(map(x -> isspace(str[x]) ? "" : str[x], 1:length(str))) #how the $%@$% to remove white space in Julia strings?
end
strandStrings = strandJSON.(strands)
if length(filter(x->x=="strands2_03.txt",map(abspath, readdir())))!=0
rm("strands2_03.txt") #remove the text file in the working directory if exists
end
touch("strands2_03.txt") #create the text file in the working directory
open("strands2_03.txt","w") do file
newID = 1764 #last strand in system
for s in strandStrings
newID +=1
write(file,"$newID: $s\n")
println("$newID: $s\n")
end
end
| phase2/strands2_03.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# <a id="top"></a>
# <center>
# <h1>PyCharm + Anaconda</h1>
# <h4><NAME> | jenfly (at) gmail (dot) com </h4>
# </center>
#
# This guide is a companion to https://jenfly.github.io/llc-python-resources/next-steps.html. Jupyter notebook source code and data are available in the [Github repo](https://github.com/jenfly/llc-python-resources).
#
# <i>Please email me with any questions, comments, or suggestions. I'd love to hear from you!</i>
# Now that you've installed Anaconda, let's see how to configure PyCharm to use Python from Anaconda, instead of the default Python that it was using before. To do this, you need to change the <b>project interpreter</b>. The project interpreter is the Python program that PyCharm uses when you execute commands in the console and run scripts. You can set up PyCharm to choose from multiple different interpreters, corresponding to different versions of Python installed on your computer.
#
# Here are the steps to follow to switch to Anaconda Python as the interpreter for our `llc-intro-to-python-master` project.
# #### 1. Open Pycharm
#
# Try typing `import pandas` in the console. You'll probably get an error like the one shown below. Notice also the line that starts with `Python 3.6.1 ...` displayed near the top of the console. This is the project interpreter and it's what we need to change. Your console will display a slightly different variation, depending on what version of Python is currently installed on your computer.
#
# <img src="screenshots/pycharm1.png">
# #### 2. In the top menu, select File -> Settings, and in the window that pops up, select "Project Interpreter"
#
# Or if you're on a Mac, select PyCharm -> Preferences from the top menu, then select "Project Interpreter"
#
# <img src="screenshots/pycharm2.png">
# #### 3. From the dropdown menu on the top right, select "Show All"
#
# <img src="screenshots/pycharm3.png">
# #### 4. In the new window that pops up, select the "+" button on the top right to add a new interpreter
#
# <img src="screenshots/pycharm4.png">
# #### 5. In the new window that pops up, select the "System Interpreter" from the side menu on the left
#
# <img src="screenshots/pycharm5.png">
# #### 6. Click the "..." button on the top right.
#
# In the new window that pops up, navigate through the file system to select the folder where Anaconda is installed. This will likely be a folder called `Anaconda3` that is contained in your main user folder (for me the folder is `C:\Users\jenfl\Anaconda3`). Select the file called `python.exe` within this folder, and click "OK".
#
# <i>Note: These instructions are for Windows. If you're on a Mac, you'll follow the same steps, but the folder and file names might be a bit different.</i>
#
# <img src="screenshots/pycharm6.png">
# #### 7. Now you're back to the previous window, which listed the available interpreters.
#
# The new interpreter that you've just added should show up in the list --- you can find it by looking at the description and seeing that the file path contains `Anaconda`. Let's give this interpreter a new name to help you find it easily.
#
# Make sure your new Anaconda interpreter is selected, and then click on the pencil icon from the side menu to edit it.
#
# <img src="screenshots/pycharm7.png">
# #### 8. In the window that pops up, edit the "Name" field to rename the interpreter (e.g. "Anaconda Python 3.6") and click "OK"
#
# <img src="screenshots/pycharm8.png">
# #### 9. Now you're back to the Settings window
#
# From the dropdown menu on the top right, make sure that your new Anaconda interpreter is selected, and then click "OK".
#
# <img src="screenshots/pycharm9.png">
# #### 10. Close PyCharm and re-open it to implement the changes we made in the previous steps.
#
# It may take a minute or two for the Python console to initialize with the new interpreter, so it might look empty at first. Then, when the console has started up and is displaying some information at the top, check out the line which starts with `Python 3.6.4 | Anaconda custom (64-bit)` --- yours will display something similar. Notice how this line has changed from step 1? This shows that we've switched over to our Anaconda Python interpreter.
#
# Now try typing `import pandas` in the console. If everything worked properly in the previous steps, this command should execute without any errors. You now have access to all the 3rd party libraries that come pre-installed with Anaconda!
#
# <img src="screenshots/pycharm10.png">
# #### 11. An example to test your new interpreter
#
# Try copying and pasting the following code into your console:
# ```
# data = pandas.DataFrame([['meow', 'woof', 'quack'], ['fur', 'fur', 'feathers']], index=['says', 'wears'], columns=['cat', 'dog', 'duck'])
# ```
# And then type `print(data)` in the console to see the new variable you just created. It should look like a little table... intriguing! This is a type of variable called a DataFrame, and it's like a table from a spreadsheet or a .csv file. With `pandas` you can do all the things you would do with your data in Excel or Google Sheets, plus so much more!
#
# For now, don't worry about the details of how the above code works --- you'll get to that in due time as you keep working through the basics and starting learning about the various libraries. Now that you've gone through this process to set up your computer with Anaconda and PyCharm, the great thing is that you have <b>all these libraries immediately available whenever you're ready to dive into them</b>. You won't need to go searching around for the libraries you need and installing them yourself --- they're already installed and ready for you!
#
# <img src="screenshots/pycharm11.png">
| pycharm-anaconda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Recommendation System
#
# Recommendation system is one of the key field for applying machine learning algorithm in most of the technological industries, Netflix (movies, shows, etc), Amazon (books, appliances, etc), Facebook (video, groups, etc), Google (route, recipe, website, etc), and many more. Understanding how to setup a recommendation system is just a starting point getting into the field of data science. Learning the application of the the tools to solve a recommendation problem will provide you the advantage in the field.
#
# Introduction to Recommender System by <NAME>:
# https://www.youtube.com/watch?v=giIXNoiqO_U
#
# ### Types of Recommendation System: Content-Based vs. Collaborative-Filtering
#
# A recommendation system usually has an objective to predict the rating or other ranking or rating metrics for the current users, so the highest predicted rating or ranking content can be recommended to user.
#
# **Typical Data Set:** Rating Book from 0 - 5 stars with missing data "?"
#
# | Book | User_1 | User_2 | User_3 | User_4 | Feature_1 (Sci-Fi) | Feature_2 (Fiction) |
# | :---: | :---: | :---: | :---: | :---: | :---: | :---: |
# | Book_1 | 5 | 4 | 1 | ? | 0.8 | 0.1 |
# | Book_2 | 5 | ? | 0 | 0 | 0.9 | 0.05 |
# | Book_3 | ? | 2 | 4 | 5 | 0.4 | 0.6 |
# | Book_4 | 0 | 0 | ? | 5 | 0.2 | 0.8 |
# | Book_5 | 1 | ? | 5 | 4 | 0.13 | 0.75 |
#
# The table represents the rating vectors from users and the feature vectors for each book.
#
# Note: Features of the content are not always available or well-defined, so this is just one example that the feature of the content is available or defined.
#
# #### Content-Based Recommendation
#
# **Content-Based** recommendation system utilizes the content features which users react positively to for predicting the behavior of a user. During the recommendation process, the **similarity metric** are calculated from the item's feature vectors and the user's preferred feature vectors from their previous record. Then, the top few are recommended by the system. Content-based filtering does not require other users' data during recommendations to one user.
#
# **Pros:**
# - No data is needed from other users
# - Individualize prediction based on user profile and can predict unique taste
# - Well explained recommendation by user profile
#
# **Cons:**
# - Hard to find all appropriate features
# - Overspecialization
# - Cold-Start problem, building user profile for first time user
#
# Content-Based Recommendations by <NAME>:
# https://www.youtube.com/watch?v=9siFuMMHNIA
#
# Let's investigate what to recommend to Bob if we are provided with a feature matrix for some TV shows.
# Import dependencies
import numpy as np
import pandas as pd
from sklearn.metrics.pairwise import cosine_similarity
# +
tv_shows = np.array([[0, 1, 1, 0, 1, 1, 1],
[0, 0, 0, 1, 1, 1, 0],
[1, 1, 1, 0, 0, 1, 1],
[0, 1, 1, 1, 0, 0, 1]])
tv_shows
# -
# Bob likes the TV Show represented by Row \#1. Which show (row) should we recommend to Bob?
#
# **Cosine Similarity**:
#
# One natural way of measuring the similarity between two vectors is by the **cosine of the angle between them**. Two points near one another in feature space will correspond to vectors that nearly overlap, i.e. vectors that describe a small angle $\theta$. And as $\theta$ decreases, $\cos(\theta)$ *increases*. So we'll be looking for large values of the cosine (which ranges between -1 and 1). We can also think of the cosine between two vectors as the *projection of one vector onto the other*:
#
# 
#
# We can use this metric easily if we treat our rows (the items we're comparing for similarity) as vectors: We can calculate the cosine of the angle $\theta$ between two vectors $\vec{a}$ and $\vec{b}$ as follows:
#
# $$\cos(\theta) = \frac{\vec{a}\cdot\vec{b}}{|\vec{a}||\vec{b}|}$$
#
# Example:
#
# $\vec{a} = \begin{bmatrix} 3 & 4 \end{bmatrix}$
#
# $\vec{b} = \begin{bmatrix} 4 \\ 3 \end{bmatrix}$
#
# $\vec{a}\cdot\vec{b} = 3 \cdot 4 + 4 \cdot 3 = 24$
#
# $|\vec{a}| = \sqrt{3^2 + 4^2} = 5$
#
# $|\vec{b}| = \sqrt{4^2 + 3^2} = 5$
#
# $COS(\theta) = \frac{\vec{a}\cdot\vec{b}}{|\vec{a}||\vec{b}|} = \frac{24}{5 \cdot 5} = 0.96$
# +
# Calculate the denominator and numerators of the Cosine Similarity
numerators = np.array([tv_shows[0].dot(tv_show) for tv_show in tv_shows[1:]])
denominators = np.array([np.sqrt(sum(tv_shows[0]**2)) *\
np.sqrt(sum(tv_show**2)) for tv_show in tv_shows[1:]])
# Calculate the Cosine Similarity for each item to item 1
similarity = numerators / denominators
# Results
print(f'Cosine Similarity between Item 1 and 2: {round(similarity[0], 3)}')
print(f'Cosine Similarity between Item 1 and 3: {round(similarity[1], 3)}')
print(f'Cosine Similarity between Item 1 and 4: {round(similarity[2], 3)}')
# -
# Using SKLearm consine_similarity()
cosine_similarity(tv_shows, tv_shows)
# Based on this result, since the cosine similarity to Item 1 is highest for Item 3, we would recommend Item 3 to Bob.
# #### Collaborative-Filtering Recommendation
#
# **Collaborative-Filtering** system does not need the features of the items to be given. Every user and item is described by a feature vector or embedding. It consider other users' reactions while recommending a particular user. It notes which item a particular user likes and also the items that the users with behavior or likings similar to him/her likes, to recommend product to that user.
#
# #### Types of Collaborative Recommendation System:
#
# ##### I. Memory-Based Collaborative Filtering:
#
# Done mainly remembering the user-item interaction matrix, and how a user reacts to it, i.e, the rating that a user gives to an item. There is no dimensionality reduction or model fitting as such. Mainly two sections:
#
# **Item-Based Filtering**:
# Recommend item X to a user based on the similarity of other items Y and Z that you liked. "Because you liked Y and Z, you may also like X."
#
# **User-Based Filtering**:
# Recommend item X to a user based on the similarity of characteristics of other users who also like item X. "The users who like products similar to you also liked those products."
#
# ##### II. Model-Based Collaborative Filtering:
#
# From the matrix, we try to learn how a specific user or an item behaves. We compress the large interaction matrix using dimensional Reduction or using clustering algorithms. In this type, We fit machine learning models and try to predict how many ratings will a user give a product. There are several methods:
#
# **Clustering Algorithms**: Normally use simple clustering Algorithms like K-Nearest Neighbours to find the K closest neighbors or embeddings given a user or an item embedding based on the similarity metrics used.
#
# **Matrix Factorization Based Algorithms**: Like any big number can be factorized into smaller numbers, the user-item interaction table or matrix can also be factorized into two smaller matrices, and these two matrices can also be used to generate back the interaction matrix.
#
# **Deep Learning Methods**: Like convolutional network model.
#
# Collaborative-Filtering Recommendation by <NAME>:
# https://www.youtube.com/watch?v=9AP-DgFBNP4
#
# Let suppose there are three users who like different items,
#
# | Items | User_1 | User_2| User_3 |
# | :---: | :---: | :---: | :---: |
# | Item_1 | 5 | 0 | 0 |
# | Item_2 | 0 | 5 | 0 |
# | Item_3 | 0 | 0 | 5 |
#
# **User-Based Filtering Example**:
#
# What should we recommend to a new user (Bob) based on the similarity of the characteristics between all users. Or, to which is Bob most similar?
# +
# 3 Users Characteristic Vectors
users = np.array([[5, 4, 3, 4, 5],
[3, 1, 1, 2, 5],
[4, 2, 3, 1, 4]])
# Bob's Characteristic Vector
Bob = np.array([5, 0, 0, 0, 0])
# +
# Bob's Characteristic vector
Bob_mag = 5
# Calculate the numerator and demoninators of cosine similarity
numerators = np.array([Bob.dot(user) for user in users])
denominators = np.array([Bob_mag * np.sqrt(sum(user**2))\
for user in users])
# Calculate the cosine similarity
similarity = numerators / denominators
# Results
print(f'Cosine Similarity between Bob and User_1: {round(similarity[0], 3)}')
print(f'Cosine Similarity between Bob and User_2: {round(similarity[1], 3)}')
print(f'Cosine Similarity between Bob and User_3: {round(similarity[2], 3)}')
# -
# Using SKLearn cosine_similarity()
all_users = np.vstack([users, Bob])
cosine_similarity(all_users, all_users)
# Based on this result, we recommend item 3 to Bob because User 3 has the highest cosine similarity score with Bob.
# ### Matrix Factorization & Latent Features
#
# Suppose we start with a matrix $R$ of users and products, where each cell records the ranking the relevant user gave to the relevant product. Very often we'll be able to record this data as a sparse matrix, because many users will not have ranked many items.
#
# **User-Item Rating Matrix**: Matrix R
#
# | Users | Item_1 | Item_2 | Item_3 | Item_4 | ... | Item_m |
# | :---: | :---: | :---: | :---: | :---: | :---: | :---: |
# | User_1 | 5 | ? | 4 | ? | ... | ? |
# | User_2 | ? | 1 | ? | 5 | ... | ? |
# | User_3 | 1 | ? | ? | 4 | ... | 4 |
# | User_4 | ? | ? | 1 | 2 | ... | ? |
# | User_5 | ? | ? | 5 | ? | ... | 1 |
# | User_6 | 3 | 5 | ? | ? | ... | ? |
# | User_7 | 2 | ? | ? | 1 | ... | ? |
# | ... | ... | ... | ... | ... | ... | ... |
# | User_n | ? | 4 | 3 | 3 | ... | 5 |
#
# Note: User-Item Rating Matrix is usually a very **sparse** matrix, which means that many cells in this matrix are empty. In real-world, a single user does not give ratings to even 1% of the total items. Therefore, around 99% of the cells of this matrix are empty.
#
# Imagine factoring this matrix into a user matrix $P$ and an item matrix $Q^T$: $R = PQ^T$. What would the shapes of $P$ and $Q^T$ be? Clearly $P$ must have as many rows as $R$, which is just the number of users who have given ratings. Similarly, $Q^T$ must have as many columns as $R$, which is just the number of items that have received ratings. We also know that the number of columns of $P$ must match the number of rows of $Q^T$ for the factorization to be possible, but this number could really be anything. In practice this will be a small number, and for reasons that will emerge shortly let's refer to these dimensions as **latent features** of the items in $R$. If $p$ is a row of $P$, i.e. a user-vector, and $q$ is a column of $Q^T$, i.e. an item-vector, then $p$ will record the user's particular weights or *preferences* with respect to the latent features, while $q$ will record how the item ranks with respect to those same latent features. This in turn means that we could predict a user's ranking of a particular item simply by calculating the dot-product of $p$ and $q$!
#
# **User Matrix**: Matrix P
#
# | Users | age | gender | ... | Feature_k |
# | :---: | :---: | :---: | :---: | :---: |
# | User_1 | 25 | 0 | ... | 0.3 |
# | User_2 | 33 | 1 | ... | 0.55 |
# | User_3 | 35 | 1 | ... | 0.35 |
# | User_4 | 28 | 0 | ... | 0.44 |
# | User_5 | 46 | 0 | ... | 0.65 |
# | User_6 | 23 | 1 | ... | 0.23 |
# | User_7 | 41 | 0 | ... | 0.32 |
# | ... | ... | ... | ... | ... |
# | User_n | 37 | 1 | ... | 0.68 |
#
# **Item Matrix**: Matrix Q
#
# | Users | Action | Over 60 minutes | Foreign Language | ... | Feature_k |
# | :---: | :---: | :---: | :---: | :---: | :---: |
# | Item_1 | 0 | 1 | 2 | ... | 0 |
# | Item_2 | 1 | 0 | 4 | ... | 1 |
# | Item_3 | 1 | 1 | 1 | ... | 1 |
# | Item_4 | 0 | 1 | 1 | ... | 0 |
# | ... | ... | ... | ... | ... | ... |
# | Item_m | 0 | 0 | 5 | ... | 0 |
#
# Note: The columns in the user matrix ($P$) and rows in the transposed item matrix ($Q^T$)are called "**Latent Factors**" and are an indication of hidden characteristics about the users or the items. The number of latent factors affects the recommendations in a manner where the greater the number of factors, the more personalized the recommendations become. But too many factors can lead to overfitting in the model.
#
# If we could effect such a factorization, $R = PQ^T$, then we could calculate *all* predictions, i.e. fill in the gaps in $R$, by solving for $P$ and $Q$.
#
# 
#
#
# One of the most popular algorithm solving such problem is called "**Alternating Least-Squares**" (**ALS**).
#
#
# ### Alternating Least Square
#
# ALS recommendation systems are often implemented in Spark architectures because of the appropriateness for distributed computing. ALS systems often involve very large datasets (consider how much data the recommendation engine for NETFLIX must have, for example!), and it is often useful to store them as sparse matrices, which Spark's ML library can handle. In fact, Spark's mllib even includes a "Rating" datatype! ALS is **collaborative** and **model-based**, and is especially useful for working with *implicit* ratings.
#
# We're looking for two matrices (a user matrix and an item matrix) into which we can factor our ratings matrix. We can't of course solve for two matrices at once. But here's what we can do:
#
# Make guesses of the values for $P$ and $Q$. Then hold the values of one *constant* so that we can optimize for the values of the other!
#
# Basically this converts our problem into a familiar *least-squares* problem. See [this page](https://textbooks.math.gatech.edu/ila/least-squares.html) and [this page](https://datasciencemadesimpler.wordpress.com/tag/alternating-least-squares/) for more details, but here's the basic idea:
#
# #### ALS in `pyspark`
#
# We'll talk about Big Data and Spark soon, but I'll just note here that Spark has a recommendation submodule inside its ml (machine learning) module. Source code for `pyspark`'s version [here](https://spark.apache.org/docs/latest/api/python/_modules/pyspark/ml/recommendation.html).
#
# #### Non-Negative Matrix Factorization
#
# In SKLearn, there is a matrix factorization method called "Non-Negative Matrix Factorization". We are using this as an example for making prediction for the user-item rating matrix.
#
# SKLearn Documentation: https://scikit-learn.org/stable/modules/generated/sklearn.decomposition.NMF.html
#
# **Python Code**:
#
# ``` Python
# # Import dependencies
# import numpy as np
# from sklearn.decomposition import NMF
#
# # Create the numpy array to store the rating information
# R = np.array(R)
#
# # Create the NMF object
# nmf = NMF()
#
# # Fit transform the rating in numpy array
# user = nmf.fit_transform(R)
#
# # R
# item = nmf.components_
#
# # Dot product of the user and item matrix
# pred_rating = np.dot(user,item)
#
# # Final predicted rating
# pred_rating
# ```
#
# Here is an example for writing a matrix factorization function, which we are going to compare to the result from NMF model from SKLearn.
#
# Source: http://www.quuxlabs.com/blog/2010/09/matrix-factorization-a-simple-tutorial-and-implementation-in-python/
# Create the Matrix Factorization Function
import numpy as np
def matrix_factorization(R, P, Q, K, steps=5000, alpha=0.0002, beta=0.02):
Q = Q.T
for step in range(steps):
for i in range(len(R)):
for j in range(len(R[i])):
if R[i][j] > 0:
eij = R[i][j] - np.dot(P[i,:],Q[:,j])
for k in range(K):
P[i][k] = P[i][k] + alpha * (2 * eij * Q[k][j] - beta * P[i][k])
Q[k][j] = Q[k][j] + alpha * (2 * eij * P[i][k] - beta * Q[k][j])
eR = np.dot(P,Q)
e = 0
for i in range(len(R)):
for j in range(len(R[i])):
if R[i][j] > 0:
e = e + pow(R[i][j] - np.dot(P[i,:],Q[:,j]), 2)
for k in range(K):
e = e + (beta/2) * (pow(P[i][k],2) + pow(Q[k][j],2))
if e < 0.001:
break
return P, Q.T
# +
# Create the rating matrix
R = [
[5,3,0,1],
[4,0,0,1],
[1,1,0,5],
[1,0,0,4],
[0,1,5,4],
]
# Transform the rating matrix into numpy array
R = np.array(R)
# Setting the parameters
N = len(R)
M = len(R[0])
K = 2
P = np.random.rand(N,K)
Q = np.random.rand(M,K)
# Apply the matrix_factorization() function
user, item = matrix_factorization(R, P, Q, K)
# Calculate the predict rating by the dot product of the user and item matrix
pred_rating = np.dot(user, item.T)
print(pred_rating)
# +
import numpy as np
from sklearn.decomposition import NMF
# User-Item Rating Matrix
R = [
[5,3,0,1],
[4,0,0,1],
[1,1,0,5],
[1,0,0,4],
[0,1,5,4],
]
# Create the numpy array to store the rating information
R = np.array(R)
# Create the NMF object
nmf = NMF()
# Fit transform the rating in numpy array
user = nmf.fit_transform(R)
# R
item = nmf.components_
# Dot product of the user and item matrix
pred_rating = np.dot(user,item)
# Final predicted rating
pred_rating
# -
| Recommendation_System_Lecture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from pathlib import Path
import numpy as np
path = Path("data/Metro_ZORI_AllHomesPlusMultifamily_Smoothed.csv")
# +
def start_pipe(dataf:pd.DataFrame) -> pd.DataFrame:
return dataf.copy()
def remove_columns(dataf):
dataf = (
dataf.drop(0)
.drop(["RegionID", "SizeRank"], axis = 1)
)
return dataf
def parse_dates(dataf):
return dataf.assign(Date = pd.to_datetime(dataf['Date'], infer_datetime_format=True))
def melt_df(dataf):
# Melt data and parse dates.
return dataf.melt(["RegionName"], var_name="Date", value_name = "RentIndex")
def interpolate_data_and_lag(dataf, lag=12, targets=6):
cities = dataf['RegionName'].unique()
interpol = []
for c in cities:
mask = dataf['RegionName'] == c
chunk = (dataf[mask]
.drop('RegionName', 1)
.set_index('Date')
.resample(rule="M")
.mean()
.interpolate()
.bfill()
)
for i in range(1,lag+1):
chunk[f't-{i}'] = chunk['RentIndex'].shift(i)
chunk[f't-{i}Diff'] = chunk['RentIndex'] - chunk[f't-{i}']
for i in range(1,targets+1):
chunk[f't+{i}'] = chunk['RentIndex'].shift(-i)
chunk['RegionName'] = c
chunk=chunk.reset_index().reset_index().rename({"index":"TimeIndex"}, axis=1)
interpol.append(chunk)
return pd.concat(interpol, ignore_index=True)
def extract_month_and_year(dataf):
day = 24*60*60
year = (365.2425)*day
return dataf.assign(
MonthSin = np.sin(dataf['Date'].dt.month * (2 * np.pi / 12)),
MonthCos = np.cos(dataf['Date'].dt.month * (2 * np.pi / 12)),
Year = dataf['Date'].dt.year,
Covid = (dataf['Date'].dt.year >= 2020) & (dataf['Date'].dt.month >= 3)
)
from category_encoders import OrdinalEncoder, OneHotEncoder
from sklearn.preprocessing import StandardScaler
encoder = OneHotEncoder()
scaler = StandardScaler()
def scale_and_encode(dataf, df_scaler, df_encoder, n_lag = 12, n_target=6):
# col_to_scale = ['RentIndex', 't-1', 't-1Diff', 't-2', 't-2Diff', 't-3',
# 't-3Diff', 't-4', 't-4Diff', 't-5', 't-5Diff', 't-6', 't-6Diff', 't-7',
# 't-7Diff', 't-8', 't-8Diff', 't-9', 't-9Diff', 't-10', 't-10Diff',
# 't-11', 't-11Diff', 't-12', 't-12Diff', 't+1', 't+2', 't+3', 't+4',
# 't+5', 't+6']
col_to_scale = [f't-{t}' for t in range(1, n_lag+1)] + [f't-{t}Diff' for t in range(1, n_lag+1)] + [f't+{t}' for t in range(1, n_target+1)]
df_scaler.fit_transform(dataf[['RentIndex']])
for col in col_to_scale:
dataf[col] = df_scaler.transform(dataf[[col]])
regions = dataf['RegionName']
dataf = encoder.fit_transform(dataf)
dataf['RegionName'] = regions
return dataf
def drop_id_columns(dataf):
return dataf.drop(['RegionName', 'Date'], axis=1).dropna()
# +
df = pd.read_csv(path)
encoder = OneHotEncoder()
scaler = StandardScaler()
n_lag, n_target = 12,6
cleaned = (df.pipe(start_pipe)
.pipe(remove_columns)
.pipe(melt_df)
.pipe(parse_dates)
.pipe(interpolate_data_and_lag, lag=n_lag, targets=n_target)
.pipe(extract_month_and_year)
.pipe(scale_and_encode, scaler, encoder, n_lag=n_lag, n_target=n_target)
)
df = (cleaned.pipe(drop_id_columns))
df
# +
max_time = max(df['TimeIndex'])
train = df[df['TimeIndex'] < int(max_time - 6)].drop('TimeIndex',1)
val = df[df['TimeIndex'] > int(max_time - 6)].drop('TimeIndex',1)
full = df[df['TimeIndex'] < int(max_time - 1)].drop('TimeIndex',1)
last = df[df['TimeIndex'] > int(max_time - 1)].drop('TimeIndex',1)
# -
def split_target(df, n_target=6):
targets = [f't+{t}' for t in range(1, n_target+1)]
return (df.drop(targets,1), df[targets])
from sklearn.metrics import mean_absolute_percentage_error as mape
from sklearn.ensemble import RandomForestRegressor
val_x, val_y = split_target(val)
train_x, train_y = split_target(train)
train_y
model = RandomForestRegressor(2500, n_jobs=-1)
model.fit(train_x, train_y.dropna())
from sklearn.metrics import mean_squared_error
preds = model.predict(val_x)
mape(scaler.inverse_transform(preds), scaler.inverse_transform(val_y))
val_idx = val_x.index.to_numpy()
ids = cleaned.loc[cleaned.index.isin(val_idx),['RegionName', 'Date']]
projections = pd.DataFrame(scaler.inverse_transform(preds), columns=[f't+{t}' for t in range(1, n_target+1)], index=val_idx)
projections
projections.join(ids)
path = Path("data/Metro_ZORI_AllHomesPlusMultifamily_Smoothed.csv")
df = pd.read_csv(path)
df
base = pd.DataFrame([val_x['t-1'],val_x['t-1'],val_x['t-1'],val_x['t-1'],val_x['t-1'],val_x['t-1']]).T
a = mean_squared_error(scaler.inverse_transform(preds), scaler.inverse_transform(val_y), squared=True)
b = mean_squared_error(scaler.inverse_transform(base), scaler.inverse_transform(val_y), squared=True)
(b-a) / b *100
all_x, all_y = split_target(full)
last_x, last_y = split_target(last)
model.fit(all_x, all_y)
pred = model.predict(last_x)
ny_proj = pd.DataFrame(scaler.inverse_transform(pred)).iloc[0,:]
ny_proj
ny_val = pd.DataFrame(scaler.inverse_transform(last_y)).iloc[0,:]
data= pd.DataFrame([ny_val, ny_proj ], index=['Actual','Projected'])
last_x
prior_data = pd.DataFrame(scaler.inverse_transform(last_x.loc[77][['t-12', 't-11','t-10','t-9','t-8','t-7','t-6','t-5','t-6','t-5','t-4','t-3','t-2','t-1']]))
prior_data
prior_data['Projected'] = np.nan
prior_data=prior_data.rename({0:'Actual'},axis=1)
data.T.set_index(pd.Index([i+15 for i in range(6)]))
combined_data = pd.concat([prior_data,data.T.set_index(pd.Index([i+15 for i in range(6)]))])
combined_data
melted = combined_data.reset_index().melt(id_vars='index')
melted
from plotly import express as px
px.line(melted, x = 'index', y = 'value', color='variable')
cities= pd.DataFrame(all_x['RegionName'].unique())
cities.to_csv("cities.csv")
cities[0].str.split(', ',expand=True).sort_values(1)
pd.DataFrame(cities[0].str.split(', ',expand=True)[1].unique()).to_csv('states.csv')
cleaned
cleaned.shape
cleaned.iloc[8483,:]
latest = cleaned.pipe(drop_id_columns).loc[8477].to_numpy().reshape(1,-1)
model.predict(latest)
latest_pred = model.predict(cleaned.pipe(drop_id_columns).loc[8477].drop([f't+{t}' for t in range(1, n_target+1)]+['TimeIndex']).to_numpy().reshape(1,-1))
scaler.inverse_transform(latest_pred)
set(col1)
val_x.columns
| notebooks/rental.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # 09 Differentiation
# See *Computational Physics* (Landau, Páez, Bordeianu), Chapter 5.1 – 5.6.
#
# These slides include material from *Computational Physics. eTextBook Python 3rd Edition.* Copyright © 2012 Landau, <NAME>. Used under the Creative-Commons Attribution-NonCommerical-ShareAlike 3.0 Unported License.
# + [markdown] slideshow={"slide_type": "subslide"}
# Taking numerical *derivatives* is based on the elementary definition
#
#
# $$
# \frac{dy(t)}{dt} := \lim_{h\rightarrow 0} \frac{y(t+h) - y(t)}{h}
# $$
#
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Problem $h\rightarrow 0$:
# * subtractive cancelations in numerator
# * overflow `number / epsilon_m`
# + [markdown] slideshow={"slide_type": "slide"}
# ## Forward difference
# Expand $y(t)$ in Taylor series
#
# $$
# y(t+h) = y(t) + h y^{(1)}(t) + \frac{h^2}{2!} y^{(2)}(t) + \frac{h^3}{3!} y^{(3)}(t) + \cdots
# $$
#
# (with the derivatives $y^{(n)} \equiv \frac{d^{n}y}{dt^n}$)
# + [markdown] slideshow={"slide_type": "fragment"}
#
# $$
# \frac{y(t+h) - y(t)}{h} = y^{(1)}(t) + \frac{h}{2!} y^{(2)}(t) + \frac{h^2}{3!} y^{(3)}(t) + \cdots
# $$
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Forward difference algorithm
#
# Neglect $\mathcal{O}(h^2)$ terms:
#
# $$
# \frac{dy(t)}{dt} \simeq D_\text{fd} y(t) \equiv \frac{y(t+h) - y(t)}{h}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# $$
# D_\text{fd} y(t) \equiv \frac{y(t+h) - y(t)}{h}
# $$
# 
# <span style="font-size: small; text-align: right">Image from Computational Physics. eTextBook Python 3rd Edition. Copyright © 2012 Landau, <NAME>. Used under the Creative-Commons Attribution-NonCommerical-ShareAlike 3.0 Unported License.</span>
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Approximation error in the forward difference algorithm
# Substitute the Taylor series into $D_\text{fd} y(t)$
#
# $$
# D_\text{fd} y(t) = y^{(1)}(t) - \frac{h}{2} y^{(2)}(t) + \cdots
# $$
# so the error is **first order in $h$**:
# $$
# y^{(1)}(t) - D_\text{fd} y(t) = \frac{1}{2} h y^{(2)}(t) + \mathcal{O}(h^2)
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Example for $D_\text{fd}$
#
# Quadratic function
# $$
# y(t) = a + bt^2 \quad \text{and}\quad y^{(1)}(t) = 2bt
# $$
# Forward difference approximation
# $$
# D_\text{fd} y(t) = \frac{y(t+h) - y(t)}{h} = 2bt + bh
# $$
# is only good for small $h \ll \frac{1}{b}$.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Central difference
# Improve over the (rather bad) error in the *forward difference algorithm* by using the *central difference*.
#
# 
#
# <span style="font-size: small; text-align: right">Image from Computational Physics. eTextBook Python 3rd Edition. Copyright © 2012 Landau, <NAME>. Used under the Creative-Commons Attribution-NonCommerical-ShareAlike 3.0 Unported License.</span>
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Central difference algorithm
# $$
# D_\text{cd} y(t) \equiv \frac{y\Big(t + \frac{h}{2}\Big) - y\Big(t - \frac{h}{2}\Big)}{h}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# Applied to the parabola $y(t) = a + bt^2$ yields the exact result:
# $$
# D_\text{cd} y(t)= 2bt
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Approximation error in $D_\text{cd}$
# Insert Taylor expansion...
#
# $$
# y^{(1)}(t) - D_\text{cd} y(t) = \frac{1}{24} h^2 y^{(3)}(t) + \mathcal{O}(h^4)
# $$
#
# The error is **second order in $h$**.
# + [markdown] slideshow={"slide_type": "fragment"}
# This is generally *much better* than the forward difference.
#
# "Generally" means "when $y(t)$ is sufficiently smooth", i.e.
# \begin{align}
# y^{(3)} \frac{h^2}{24} &\ll y^{(2)} \frac{h}{2} \quad\text{or}\\
# 12\frac{y^{(2)}}{y^{(3)}} &\gg h.
# \end{align}
# + [markdown] slideshow={"slide_type": "fragment"}
# Other difference algorithms exist with even better error behavior (see homework) but all have problems with data that are not smooth. In this case you might have to smooth the data in advance by *interpolation* or *kernel density estimates* (KDE).
# + [markdown] slideshow={"slide_type": "slide"}
# ## Error assessment
# $$
# \epsilon_\text{tot} = \epsilon_\text{app} + \epsilon_\text{ro}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# **Round-off error of differentiation**: difference of two similar numbers cannot be better than machine precision, always divided by small number $h$:
#
# $$
# \epsilon_\text{ro} \approx \frac{\epsilon_m}{h}
# $$
# + [markdown] slideshow={"slide_type": "subslide"}
# **Forward difference algorithmic error**
# $$
# \epsilon_\text{app}^\text{fd} = y^{(2)} \frac{h}{2}
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# What is the *most accurate value of $h$* that we can choose for $D_\text{fd}$?
# + [markdown] slideshow={"slide_type": "fragment"}
# Cross-over from approximation error to round-off error around
#
# $$
# \epsilon_\text{app} = \epsilon_\text{ro}
# $$
#
# For the forward difference
# $$
# y^{(2)} \frac{h_\text{fd}}{2} = \frac{\epsilon_m}{h_\text{fd}}\\
# h_\text{fd}^2 = \frac{2\epsilon_m}{y^{(2)}}
# $$
#
# + [markdown] slideshow={"slide_type": "slide"}
# For rough estimate, assume $y^{(1)} \approx y^{(2)} \approx y^{(3)} \approx 1$:
# + slideshow={"slide_type": "fragment"}
import numpy as np
def h_fd(eps_m, y2=1):
return np.sqrt(2*eps_m/y2)
def eps_fd(eps_m, y2=1.):
h = h_fd(eps_m, y2=y2)
return 2*eps_m/h
# + slideshow={"slide_type": "fragment"}
eps_m = 1e-15
print("h_fd = {0:.2e}".format(h_fd(eps_m, y2=1)))
print("eps_fd = {0:.2e}".format(eps_fd(eps_m, y2=1)))
# + [markdown] slideshow={"slide_type": "fragment"}
# For double precision ($\epsilon_m = 10^{-15}$)
# $$
# h_\text{fd} \approx 4\times 10^{-8}\\
# \epsilon_\text{tot}^\text{fd} \approx 2\epsilon_\text{ro} = \frac{2\epsilon_m}{h_\text{fd}} = h_\text{fd} = 4 \times 10^{-8}
# $$
#
# (Note: simplified because $y^{(2)} \approx 1$.)
# + [markdown] slideshow={"slide_type": "subslide"}
# Do you expect the "best" $h_\text{cd}$ for the *central difference algorithm* to be larger or smaller than $h_\text{fd}$?
# + [markdown] slideshow={"slide_type": "fragment"}
# $$
# h_\text{cd}^3 = \frac{24\epsilon_m}{y^{(3)}}
# $$
#
# Generally $h_\text{cd} > h_\text{fd}$ because the better algorithm allows you to use a bigger step size for a smaller error.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Implementation in Python
#
# \begin{align}
# D_\text{fd} y(t) &\equiv \frac{y(t+h) - y(t)}{h} \\
# D_\text{cd} y(t) &\equiv \frac{y\Big(t + \frac{h}{2}\Big) - y\Big(t - \frac{h}{2}\Big)}{h}
# \end{align}
#
# and also the *extended difference algorithm*
#
# \begin{align}
# D_\text{ep} y(t) &\equiv \frac{4 D_\text{cd}y(t, h/2) - D_\text{cd}y(t, h)}{3} \\
# &= \frac{8\big(y(t+h/4) - y(t-h/4)\big) - \big(y(t+h/2) - y(t-h/2)\big)}{3h}
# \end{align}
#
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Student Problem: Finite Difference Operators
# - implement the three finite difference operators as functions
# - test implementations
# - `git pull` the course resources and work with the notebook **[10_differentiation/10-differentiation-students.ipynb](http://nbviewer.jupyter.org/github/ASU-CompMethodsPhysics-PHY494/PHY494-resources/blob/master/10_differentiation/10-differentiation-students.ipynb)**.
# + slideshow={"slide_type": "-"}
def D_fd(y, t, h):
"""Forward difference"""
return (y(t + h) - y(t))/h
def D_cd(y, t, h):
"""Central difference"""
# implement
def D_ed(y, t, h):
"""Extended difference"""
# implement
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Test your implementations
# Test function: $y(t) = \cos t$
# 1. What is the analytical derivative $\frac{d\cos(t)}{dt}$?
# 1. Calculate the derivative of $y(t) = \cos t$ at $t=0.1, 1, 100$.
# 1. Print derivative and relative error $E = \frac{D y(t) - y^{(1)}(t)}{y^{(1)}(t)}$ (finite difference value $D y(t)$ compared to the analystical value $y^{(1)}(t)$– use numpy functions for "exact" values) as function of $h$.
# 1. Reduce $h$ until you reach machine precision, $h \approx \epsilon_m$
# 1. Plot $\log_{10} |E(h)|$ against $\log_{10} h$.
#
# Try to do the above for all three algorithms
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Function definitions
# -
def D_fd(y, t, h):
"""Forward difference"""
return (y(t + h) - y(t))/h
def D_cd(y, t, h):
"""Central difference"""
return (y(t + h/2) - y(t - h/2))/h
def D_ed(y, t, h):
"""Extended difference"""
return (8*(y(t + h/4) - y(t - h/4)) - (y(t + h/2) - y(t - h/2)))/(3*h)
# + slideshow={"slide_type": "-"}
import numpy as np
# test function: np.cos
# Analytical derivative
def y1(t):
return -np.sin(t)
t_values = np.array([0.1, 1, 100], dtype=np.float64)
# -
# Use numpy functions for everything because then you can operate on all `t_values` at once.
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Evaluate the finite difference derivatives
# Note that we pass *a function* `y` to the forward difference function `D_fd` and we can also pass a whole array of `t_values`!
# + slideshow={"slide_type": "-"}
D_fd(np.cos, t_values, 0.1)
# -
D_fd(np.cos, t_values, 1e-12)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Evaluate the exact derivatives
# Compute the exact derivatives (again, operate on all $t$ together... start thinking in numpy arrays!)
# + slideshow={"slide_type": "-"}
y1(t_values)
# + [markdown] slideshow={"slide_type": "fragment"}
# Calculation of the **absolute error**: subtract the two arrays that you got previously:
# -
D_fd(np.cos, t_values, 0.1) - y1(t_values)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Calculate the relative error $E$
# + slideshow={"slide_type": "-"}
def error(Dxx, y, y1, t, h):
"""Relative error.
Note: will fail for exact 0.
"""
y1_val = y1(t)
# return (Dxx(y, t, h) - y1_val/y1_val
return Dxx(y, t, h)/y1_val - 1
# -
# Note that we pass again a general function for the difference operator so that we can use `error()` with `D_fd()`, `D_cd()` and `D_ep()`.
# + slideshow={"slide_type": "fragment"}
error(D_fd, np.cos, y1, t_values, 0.1)
# -
error(D_fd, np.cos, y1, t_values, 1e-12)
# + [markdown] slideshow={"slide_type": "subslide"}
# #### Plot $|E|$
# Plot $\log_{10} |E(h)|$ against $\log_{10} h$.
# -
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
matplotlib.style.use('ggplot')
h_values = 10**(np.arange(-15, -1, 0.1))
abs_errors = np.abs(error(D_fd, np.cos, y1, 0.1, h_values))
plt.loglog(h_values, abs_errors, label="t=0.1")
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Forward Difference $D_\text{fd}$
# Plot the three different $t$ values together in one plot:
# + slideshow={"slide_type": "-"}
for t in t_values:
abs_errors = np.abs(error(D_fd, np.cos, y1, t, h_values))
plt.loglog(h_values, abs_errors, label=r"$t={}$".format(t))
ax = plt.gca()
ax.legend(loc="best")
ax.set_xlabel(r"$h$")
ax.set_ylabel(r"$|E_\mathrm{fd}|$")
# -
# * error behavior depends on $t$ and on cancellation of errors (e.g. for $t=1$
# * algorithmic error decreases for decreasing $h$ until the round of error starts dominating
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Central Difference $D_\text{cd}$
# Plot the three different $t$ values together in one plot:
# -
for t in t_values:
abs_errors = np.abs(error(D_cd, np.cos, y1, t, h_values))
plt.loglog(h_values, abs_errors, label=r"$t={}$".format(t))
ax = plt.gca()
ax.legend(loc="best")
ax.set_xlabel(r"$h$")
ax.set_ylabel(r"$|E_\mathrm{cd}|$")
# + [markdown] slideshow={"slide_type": "subslide"}
# ##### Extended Difference $D_\text{ed}$
# Plot the three different $t$ values together in one plot:
# -
for t in t_values:
abs_errors = np.abs(error(D_ed, np.cos, y1, t, h_values))
plt.loglog(h_values, abs_errors, label=r"$t={}$".format(t))
ax = plt.gca()
ax.legend(loc="best")
ax.set_xlabel(r"$h$")
ax.set_ylabel(r"$|E_\mathrm{ed}|$")
# + slideshow={"slide_type": "skip"}
| 09_differentiation/09-differentiation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="glA3NV9MRl63" colab_type="text"
# # Particle Filter - Lab 7.3
#
# ## Recap
# This is the Lab on using a Particle Filter in CE6003's Object Tracking. You should complete the tasks in this lab as part of the Particle Filter section of the lesson.
#
# Please remember this lab must be completed before taking the quiz at the end of this lesson.
#
# First, if we haven't already done so, we need to clone the various images and resources needed to run these labs into our workspace.
# + id="md1qGUkYR6H-" colab_type="code" colab={}
# !git clone https://github.com/EmdaloTechnologies/CE6003.git
# #!git clone https://github.com/mcnamarad1971/CE6003.git
# + [markdown] id="Nvp_ecbWSB6V" colab_type="text"
#
# **Program Description**
#
# This program demonstrates a very simple 'tracking' mechanism - derived from a Particle filter. We're going to use our Particle filter to track a single object, namely a person.
#
# + id="Nlei6UBfjbbN" colab_type="code" colab={}
import sys
sys.path.insert(1, "/content/CE6003/code")
# + id="qWvUXFH1iaoJ" colab_type="code" colab={}
import os
import re
import io
import cv2
import time
import numpy as np
import base64
from IPython.display import clear_output, Image, display
from motion_tracking import ParticleFilter # Import pre-cooked Particle filter
# + [markdown] id="_GSU76keSNr7" colab_type="text"
# #The Story So Far
#
# To illustrate how to track something in a video stream, we have used the following technique to generate a set of images for you to work on.
#
# What we did was we generated a short video - just recording one person walking around, on an iPhone.
#
# Then we used ```ffmpeg``` to decompose about 7 seconds of that video down into still images.
#
# ```ffmpeg -i $vid_in -vf fps=30 imgs/daire%03d.png```
#
# We saved those frames as ```imgs/daire%03d.png``` in the git repository in the single-detections directory
#
# We've run yolo3 over those frames to generate bounding boxes and saved those bounding boxes into the same directory.
#
# The file format is comma-separated values and the values are as shown here:
#
# frame index | object-type | centre-x | centre-y | width | height | confidence
# --- | --- | --- | --- | --- | --- | ---
# int | -1 | float | float | float | float | float
#
# * The object type is always a person - that's all we inferred for.
# * The centre-x and width are fractions of the image's width
# * The centre-y and height are fractions of the image's height
# * The confidence is supplied by Yolo3
# + [markdown] id="GYWphuuLX6hr" colab_type="text"
# *What Happens Now*
#
# For each image in the directory, in order,
# * we'll find the centre of the detection in that image (if any)
# * we'll build a bounding box for the detection in that image
# * we'' derive a variance term (crudely) from the Yolo confidence for that image
# * and we'll supply the centre of that bounding box along with the variance term to a Particle Filter implementation
#
# Then, we'll explore how a Particle filter tracks the object in the image stream.
# + [markdown] id="hY4bHjZsuWQ2" colab_type="text"
# **Key Parameters**
#
# We have four key parameters to shape our particle filter. We have the number of particles, and three terms associated with an action estimate;
# xVel, yVel, and velStd
# + id="JujHzAd7iWo_" colab_type="code" colab={}
numParticles = 50
xVel = 5
yVel = 5
velStd = 25
# + [markdown] id="XsQyMnFpZBUP" colab_type="text"
# **Get File Handles**
#
# This function gets the filenames of all the files in the directory, in a reproducible order, and loads in the bounding boxes from file.
# + id="I1lGqvyGZBjj" colab_type="code" colab={}
def get_pngs_and_boxes():
pngdir = "/content/CE6003/images/lab7/single-objects/"
bbdir = "/content/CE6003/images/lab7/single-objects/"
pngfolder = os.fsencode(pngdir)
bbfolder = os.fsencode(bbdir)
pngfiles = []
for filename in os.listdir(pngfolder):
if filename.decode().endswith(".png"):
pngfiles.append(pngdir + filename.decode())
pngfiles.sort()
for filename in os.listdir(bbfolder):
if filename.decode().endswith(".boxes"):
bbfilename = bbdir + filename.decode()
bb = open(bbfilename, "r")
bb_lines = bb.readlines()
bb.close()
return bb_lines, pngfiles
# + [markdown] id="qDJASmq8ZrwJ" colab_type="text"
# **Parse Detections**
#
# We'll use this function in the main loop to wrangle the detections into the format we want to supply to our Particle Filter.
#
# Essentially it takes the name of png file, an img object and the list of bounding boxes as inputs.
#
# It then finds the correct record (if any) for that image in the bounding boxes list and converts the bounding box parameters into a format which we'll use for the rest of the program (it converts back to absolute pixel values).
#
# It returns a centre and a confidence value for the image supplied to it.
# + id="HQw8QFkqZxMD" colab_type="code" colab={}
def parse_detections(bboxes, pngfile, img):
# Sample Line: 400,-1,0.285417,0.241667,0.094792,0.483333,0.999797,-1,-1,-1
# Index, object type,
# x - centre of bounding box (as fraction of image width
# y - centre of bounding box (as fraction of image height
# w - width of bounding box (as fraction of image width)
# h - height of bounding box (as fraction of image height
# prob, _,_,_
# extract the frame index of the png file -
# use it to find the detections for that frame
index = int(re.findall(r'\d+', pngfile)[-1])
imgh, imgw = img.shape[:2]
centre = np.zeros(shape=(2, 1))
P = 0.000001 # hack to avoid div by zero
for line in bboxes:
np_array = np.genfromtxt(io.StringIO(line), delimiter=",")
lineindex = int(np_array[0])
if lineindex == index:
centre = np_array[2:4]
P += np_array[6]
centre[0] *= imgw
centre[1] *= imgh
return centre, P
return centre, P
# + [markdown] id="x4RRg22Hitb5" colab_type="text"
# #Particle Filter
#
# Look back over Particle Filter for an insight into how it is operating.
#
# The concept is:
# * For a low computational cost
# * Generate a set of sampled state update/prediction terms (particles)
# * Generate a measurement prediction from that state
# * Calculate the difference between the predicted measurement of the selected particles and the actual measurement
# * Adjust the state update/prediction particles (through a resampling stage to prevent degeneration of filter), and repeat....
#
# Done on Monte Carlo sampled particles.
#
# One key term to watch is how many particles to use for your specific tracking application.
# + [markdown] id="d7dlyv0Nk7EU" colab_type="text"
# ##Demo
#
# ### Program Execution
# For each file:
# * get centre of detection (if any) and confidence from Yolo
# * feed Particle Filter with these values
# * Print internal Particles used by Particle Filter
#
#
# + id="56tX2yoKlFiS" colab_type="code" colab={}
writer = None
def demo_particle():
global writer
global numParticles
global velStd, xVel, yVel
# Initialise the filter with height of the frame, width of the frame
# and the number of particles
particleFilter = ParticleFilter(1920, 1080, numParticles)
bb_lines, pngfiles = get_pngs_and_boxes()
for pngfile in pngfiles:
#print("handling .." + os.path.basename(pngfile))
img = cv2.imread(pngfile)
# Derive meas-var from yolo confidence level in detection
raw_centre, conf = parse_detections(bb_lines, pngfile, img)
# Crudely derive meas-var. If yolo is confident we want a small
# uncertainty. If yolo isn't confident, translate to
# a large uncertainty.
if(conf > 0.50):
lStd = velStd
else:
lStd = 1
# update weights of particles based on measure
particleFilter.update(raw_centre.item(0), raw_centre.item(1))
# Pretty print particles
for i in range(0, numParticles):
x_part, y_part = particleFilter.returnParticlesCoordinates(i)
cv2.circle(img, (x_part, y_part), 10, (0,255,0),-1)
# Resize and show the image
img2 = cv2.resize(img, (int(img.shape[1]/4), int(img.shape[0]/4)))
# update model - using 5 pixels in x and y and adjusting model variance
# depending on yolo confidence
particleFilter.predict(x_velocity=xVel,y_velocity=yVel, std=lStd)
# estimate the position of the point, based on particle weights
x_est, y_est, _, _ = particleFilter.estimate()
#The resampling draws particles from the current set with a
# probability given by the current weights.
# The new set is an approximation of the distribution which represents the state
# of the particles at time t.
# The resampling solves this problem: after some iterations of the algorithm
# some particles are useless because they do not represent the point
#position anymore, eventually they will be too far away from the real position.
#The resample function removes useless particles and keep the
#useful ones.
particleFilter.resample()
# Build a frame of our output video
if writer is None:
# Initialize our video writer
fourcc = cv2.VideoWriter_fourcc(*'VP80')
writer = cv2.VideoWriter('video.webm', fourcc, 30, (img2.shape[1], img2.shape[0]), True)
# Write the output frame to disk
writer.write(img2)
# Release the file pointers
writer.release()
demo_particle()
# + [markdown] id="8fZJvZXeY15y" colab_type="text"
# **Video**
#
# Thia code plays the video we just made.
#
# The Particles being used by the Filter dot in red.
#
# As you can see, Particle Filtering has a role to play in predicting a reasonable guess for where the object might be while it is off-camera - it should be better than Kalman for non-linear motion.
# + id="mxvG8It3RlWy" colab_type="code" colab={}
# Set this to 1 if video display
# is not working - works with chrome and firefox, not with safari
videoBodge = 0
def arrayShow (imageArray):
ret, png = cv2.imencode('.png', imageArray)
encoded = base64.b64encode(png)
return Image(data=encoded.decode('ascii'))
if(videoBodge == 0):
from IPython.display import HTML
from base64 import b64encode
webm = open('video.webm','rb').read()
data_url = "data:video/webm;base64," + b64encode(webm).decode()
else:
video = cv2.VideoCapture("video.webm")
while(video.isOpened()):
clear_output(wait=True)
ret, frame = video.read()
if(ret == False):
break
lines, columns, _ = frame.shape
img = arrayShow(frame)
display(img)
time.sleep(1)
# + id="nQ3zQNbG86Dq" colab_type="code" colab={}
# Display Video
HTML("""
<video width=200 controls>
<source src="%s" type="video/webm">
</video>
""" % data_url)
# + [markdown] id="OH4IGJkZlO_u" colab_type="text"
# # Conclusion
#
# ## Exercises
# **Exercise 1**
# Simulate occluding the object being detected - for example, only supply every second measurement update to the Particle Filter and observe the Filter behaviour.
#
# **Exercise 2**
# Simply multiply and divide the number of particles and observe how that affects the Particle Filter's predictions.
#
#
# ## Takeaways
# 1. You've seen a Particle Filter used for single object tracking
# 2. You've seen that a Particle Filter can help deal with occlusions - i.e. in this example the object being tracked disappeared for a few frames and the Particle Filter continued to predict motion for it based on its model.
# 3. You've seen that you probably need to tune the Filter to get it working for a particular application.
#
# ## Next Steps
# 1. We've seen a set of tracking filters, from the same 'Kalman' family. Now, we'll look at how to track multiple objects in a scene and then we'll look at other tracking techiques not in the 'Kalman' family such as Optical Flow and CNN Feature based trackers.
| Lab_7_3_ParticleFilter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PaddlePaddle 2.1.2 (Python 3.5)
# language: python
# name: py35-paddle1.2.0
# ---
# # 项目介绍: 基于PaddleLite的树莓派垃圾检测系统
#
# 生活中,垃圾随处可见。道路需要清洁工人们辛苦打扫,并根据一定的规则进行垃圾种类划分。本项目旨在简化该项同类任务中的前置任务,即垃圾智能检测定位与识别,然后搭载小车实现垃圾分类收集。后期,只需要将收集好的垃圾,交于清洁工人们进行简单再分类即可完成路面等地方的垃圾收集分类工作。
#
# ------
#
# 主要框架:
#
# - `垃圾检测功能`: **采用深度学习的目标检测算法实现**,**PaddleDetection开发**
#
# - `硬件部署`: 采用**树莓派4B**,**32位操作系统**,**PaddleLite开发**
#
# - `硬件协同`: (小车结构在该项目的展示中,暂未说明)
# # 一、基于PPDet开发垃圾检测模型
#
# > PPDet: PaddleDetection为飞桨的官方目标检测套件,可以实现众多目标检测模型的训练。
#
# 本项目,基于PPDet开发`PPyolo_r18vd`模型作为垃圾检测项目的深度学习模型,以期望获得视觉中出现的指定垃圾的类别以及相对位置信息,从而辅助目标检测模型的应用。
#
# > 本项目实现的模型,最终落地于`垃圾分拣车`上——`实现垃圾的定位与识别检测,从而进行定位抓取与垃圾识别`。
#
# ---------
#
# 项目所需`模型要求`如下:
#
#
# 1. **模型运行速度**
#
# 2. **模型漏检率优先**
#
# 3. **模型检测识别精度**
#
# -------
#
# 预期`模型选择`: -- `ppyolo_r18_vd` -- , -- `ppyolo_tiny` --
#
#
# | 模型 | 精度(all:%) | 帧率(s) | 漏检率 | 训练成本 |
# | :--------: | :--------: | :--------: | :--------: | :--------: |
# | ppyolo_tiny | 0.5_mAP:95+ | 3-5 | 一般 | 低 |
# | ppyolo_r18_vd | 0.5_mAP:97+ | 1.4-1.6 | 较低 | 低 |
#
# **数据集格式**: COCO/VOC都有尝试, 本项目选用COCO介绍。
#
# > 感兴趣的小伙伴可以观看一个PPDet使用(说明)视频: [PPDet简单使用教程](https://www.bilibili.com/video/BV1vK4y1M728)
#
# > 声音有些小,可能需要带耳机食用~,还望谅解
# ## 1.1 解压PPDet套件
#
# > 本项目基于套件本身进行开发,因此需要导入套件包——已挂载到本项目,可直接使用
# -oq 静默解压
# !unzip -oq data/data99077/PaddleDetection-release-2.1.zip
# !mv PaddleDetection-release-2.1 PaddleDetection
# ## 1.2 解压数据集
#
# > 为方便模型开发训练,因此直接解压到套件中的`dataset`目录下,并新建`diy_coco`来保存
#
# **数据集目录:**
#
# - `PaddleDetection`
#
# - `dataset`
#
# - `diy_coco`
#
# - `Train`
#
# - `Annotations`: **包含coco格式的标注json文件**
#
# - `Images`: **训练图片**
#
# - `Eval`
#
# - `Annotations`: **包含coco格式的标注json文件**
#
# - `Images`: **验证/评估图片**
#
# -------
#
# **部分数据标注展示:**
#
# 
#
# !unzip -oq data/data101886/rubish_det.zip -d PaddleDetection/dataset/diy_coco
# ## 1.3 下载环境依赖(包)
#
# > 主要是补充下载pycocotool,这对解析coco数据格式的标注提供很大的帮助
# %cd /home/aistudio/PaddleDetection
# !pip install -r requirements.txt
# ## 1.4 训练前,明确anchor大小
#
# > 在训练开始前,现在训练数据上,生成一个符合待拟合数据的`anchor`,这将对模型学习合适的特征提供帮助,同时也能更好的框选预测物体的位置!
#
# > 仅限于需要`预置anchor`的模型
#
# 不过,再开始生成`anchor`前,需要先配置好`数据集的加载`。
#
# -------
#
# ### 1.4.1 配置数据加载yml
#
# 因为,本项目的数据格式为coco,因此选择路径: `PaddleDetection/configs/datasets`下的`coco_detection.yml`文件进行修改,使其加载本项目垃圾检测数据!
#
# 修改如下:
#
# metric: COCO
# # 修改num_classes为垃圾分类的数量
# num_classes: 5
#
# TrainDataset:
# # !COCODataSet
# # 2.再配置图片路径 -- 指向Images文件夹
# image_dir: Images
# # 3.最后配置标注文件的路径 -- 指向Annotations下的json文件
# anno_path: Annotations/train.json
# # 1.先配置数据集目录 -- 先指向Train文件夹
# dataset_dir: dataset/diy_coco/Train
# # end: 这里不同改
# data_fields: ['image', 'gt_bbox', 'gt_class', 'is_crowd']
#
# EvalDataset:
# # !COCODataSet
# image_dir: Images
# anno_path: Annotations/val.json
# # 1. 指向另一个文件夹,用于验证评估,其它同上
# dataset_dir: dataset/diy_coco/Eval
#
# TestDataset:
# # !ImageFolder
# # 这里的标注配置,设置为验证的json即可
# anno_path: Annotations/val.json
# ### 1.4.2 选择参与训练的模型
#
# 已经有了配置好的数据加载yml文件,接下来就可以选模型了。
#
# 这里选用`PaddleDetection/configs/ppyolo`下的`ppyolo_r18vd_coco.yml`作为项目要训练的模型。
#
# -------
#
# ### 1.4.3 生成预置anchor
#
# 以上完成了数据加载的配置以及模型的选择之后,我们就可进行预置anchor的自动生成了!
#
# **生成的大致流程:**
#
# 1. 启动时,调用`模型yml`进入参数配置,获取`数据集加载的yml`信息
#
# 2. 生成时,利用数据集中的`所有已有标注信息`进行anchor的`kmeans聚类`生成一个`anchor集合`
#
# 3. 使用时,将生成的anchor收集起来,然后替换模型yml中所有出现anchor列表的地方`即可
# -n: 模型中需要的anchor数量, r18只需要6个
# -s: 生成anchor集合,适用于多大的输入尺寸 —— 会自动生成指定大小下的anchor集合
# -c: 指定使用这些anchor的模型yml
# %cd /home/aistudio/PaddleDetection
# !python tools/anchor_cluster.py -n 6 -s 320 -c configs/ppyolo/ppyolo_r18vd_coco.yml
# ### 1.4.4 整合生成的anchor,并替换模型的anchor
#
# > 替换anchor的地方,对于r18而言有以下两个地方: `configs/ppyolo/ppyolo_r18vd_coco.yml`, `configs/ppyolo/_base_/ppyolo_r18vd.yml`
#
# ------
#
# `ppyolo_r18vd_coco.yml`中的修改如下(**模型yml**):
#
# - Gt2YoloTarget:
# anchor_masks: [[3, 4, 5], [0, 1, 2]]
# # 替换anchor列表为生成的anchor即可
# anchors: [[48, 36], [43, 66], [89, 60], [60, 102], [105, 124], [165, 163]]
# downsample_ratios: [32, 16]
#
# -------
#
# `ppyolo_r18vd.yml`中的修改如下(**模型结构yml**):
#
# YOLOv3Head:
# anchor_masks: [[3, 4, 5], [0, 1, 2]]
# # 替换anchor列表为生成的anchor即可
# anchors: [[48, 36], [43, 66], [89, 60], [60, 102], [105, 124], [165, 163]]
# loss: YOLOv3Loss
#
create_anchors_list = [[59, 45], [54, 82], [112, 74], [75, 127], [131, 154], [206, 204]]
# ## 1.5 配置训练参数
#
# > 对于r18而言,训练参数的修改只需要在 `configs/ppyolo/ppyolo_r18vd_coco.yml`中修改即可
#
# **主要参数修改如下**:
#
# TrainReader:
# sample_transforms:
# ...
# batch_transforms:
# - BatchRandomResize:
# # 原始大小的list对应输入大小为520的预测,现改为320之后,简要修改的这个区间
# # 修改注意事项,每个大小都是32的倍数
# target_size: [224, 256, 288, 320, 352, 384, 416, 448]
# ...
# - Gt2YoloTarget:
# anchor_masks: [[3, 4, 5], [0, 1, 2]]
# # 替换为生成的anchor
# anchors: [[48, 36], [43, 66], [89, 60], [60, 102], [105, 124], [165, 163]]
# downsample_ratios: [32, 16]
# # 根据数据集情况,适当修改即可: 8/16/24/32/48
# batch_size: 32
# mixup_epoch: 500
# shuffle: true
#
#
# EvalReader:
# sample_transforms:
# - Decode: {}
# # target_size改为320
# - Resize: {target_size: [320, 320], keep_ratio: False, interp: 2}
# ...
#
# TestReader:
# inputs_def:
# # 改为320
# image_shape: [3, 320, 320]
# sample_transforms:
# - Decode: {}
# # 改为320
# - Resize: {target_size: [320, 320], keep_ratio: False, interp: 2}
# ...
#
# LearningRate:
# # 原4卡下训练参数,除以4,用于单卡训练
# # 0.004 / 4 == 0.001
# base_lr: 0.001
# ...
# %cd /home/aistudio/PaddleDetection
# !python tools/train.py\
# -c configs/ppyolo/ppyolo_r18vd_coco.yml\
# --eval\
# --use_vdl True
# [08/14 21:03:03] ppdet.engine INFO: Epoch: [196] [20/46] learning_rate: 0.000100 loss_xy: 0.773786 loss_wh: 0.697323 loss_iou: 2.933347 loss_obj: 3.114668 loss_cls: 0.885066 loss: 8.543031 eta: 0:27:02 batch_cost: 0.4652 data_cost: 0.2992 ips: 68.7832 images/s
# [08/14 21:03:12] ppdet.engine INFO: Epoch: [196] [40/46] learning_rate: 0.000100 loss_xy: 0.757029 loss_wh: 0.656280 loss_iou: 2.774072 loss_obj: 3.072931 loss_cls: 0.949183 loss: 8.486620 eta: 0:26:52 batch_cost: 0.4206 data_cost: 0.2787 ips: 76.0866 images/s
# [08/14 21:03:17] ppdet.engine INFO: Epoch: [197] [ 0/46] learning_rate: 0.000100 loss_xy: 0.758142 loss_wh: 0.664071 loss_iou: 2.743285 loss_obj: 3.071552 loss_cls: 1.033830 loss: 8.424139 eta: 0:26:50 batch_cost: 0.4621 data_cost: 0.3208 ips: 69.2533 images/s
# [08/14 21:03:26] ppdet.engine INFO: Epoch: [197] [20/46] learning_rate: 0.000100 loss_xy: 0.736949 loss_wh: 0.639424 loss_iou: 2.764338 loss_obj: 3.022928 loss_cls: 1.026918 loss: 8.329489 eta: 0:26:40 batch_cost: 0.4258 data_cost: 0.2777 ips: 75.1583 images/s
# [08/14 21:03:36] ppdet.engine INFO: Epoch: [197] [40/46] learning_rate: 0.000100 loss_xy: 0.728324 loss_wh: 0.671651 loss_iou: 2.920363 loss_obj: 3.044627 loss_cls: 0.976078 loss: 8.474413 eta: 0:26:30 batch_cost: 0.4600 data_cost: 0.3220 ips: 69.5716 images/s
# [08/14 21:03:40] ppdet.engine INFO: Epoch: [198] [ 0/46] learning_rate: 0.000100 loss_xy: 0.748800 loss_wh: 0.663416 loss_iou: 2.903050 loss_obj: 3.142794 loss_cls: 0.995665 loss: 8.490379 eta: 0:26:27 batch_cost: 0.5249 data_cost: 0.3624 ips: 60.9593 images/s
# [08/14 21:03:50] ppdet.engine INFO: Epoch: [198] [20/46] learning_rate: 0.000100 loss_xy: 0.804090 loss_wh: 0.638163 loss_iou: 2.821011 loss_obj: 3.293034 loss_cls: 0.950222 loss: 8.611068 eta: 0:26:17 batch_cost: 0.4455 data_cost: 0.2798 ips: 71.8259 images/s
# [08/14 21:03:59] ppdet.engine INFO: Epoch: [198] [40/46] learning_rate: 0.000100 loss_xy: 0.729478 loss_wh: 0.671696 loss_iou: 2.855099 loss_obj: 2.954676 loss_cls: 1.013126 loss: 8.109439 eta: 0:26:08 batch_cost: 0.4445 data_cost: 0.3092 ips: 71.9917 images/s
# [08/14 21:04:04] ppdet.engine INFO: Epoch: [199] [ 0/46] learning_rate: 0.000100 loss_xy: 0.729086 loss_wh: 0.640540 loss_iou: 2.748984 loss_obj: 3.005687 loss_cls: 0.877229 loss: 7.902369 eta: 0:26:05 batch_cost: 0.5034 data_cost: 0.3502 ips: 63.5677 images/s
# [08/14 21:04:14] ppdet.engine INFO: Epoch: [199] [20/46] learning_rate: 0.000100 loss_xy: 0.763439 loss_wh: 0.640906 loss_iou: 2.689836 loss_obj: 3.238860 loss_cls: 0.929343 loss: 8.205533 eta: 0:25:56 batch_cost: 0.4675 data_cost: 0.2824 ips: 68.4485 images/s
# [08/14 21:04:24] ppdet.engine INFO: Epoch: [199] [40/46] learning_rate: 0.000100 loss_xy: 0.757755 loss_wh: 0.720121 loss_iou: 2.960909 loss_obj: 3.277584 loss_cls: 0.926977 loss: 8.504792 eta: 0:25:46 batch_cost: 0.4711 data_cost: 0.3046 ips: 67.9259 images/s
# [08/14 21:04:27] ppdet.utils.checkpoint INFO: Save checkpoint: output/ppyolo_r18vd_coco
# [08/14 21:04:27] ppdet.engine INFO: Eval iter: 0
# [08/14 21:04:32] ppdet.metrics.metrics INFO: The bbox result is saved to bbox.json.
# loading annotations into memory...
# Done (t=0.01s)
# creating index...
# index created!
# [08/14 21:04:32] ppdet.metrics.coco_utils INFO: Start evaluate...
# Loading and preparing results...
# DONE (t=0.12s)
# creating index...
# index created!
# Running per image evaluation...
# Evaluate annotation type *bbox*
# DONE (t=1.27s).
# Accumulating evaluation results...
# DONE (t=0.19s).
# Average Precision (AP) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.667
# Average Precision (AP) @[ IoU=0.50 | area= all | maxDets=100 ] = 0.960
# Average Precision (AP) @[ IoU=0.75 | area= all | maxDets=100 ] = 0.836
# Average Precision (AP) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.771
# Average Precision (AP) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = -1.000
# Average Precision (AP) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = -1.000
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 1 ] = 0.394
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets= 10 ] = 0.764
# Average Recall (AR) @[ IoU=0.50:0.95 | area= all | maxDets=100 ] = 0.776
# Average Recall (AR) @[ IoU=0.50:0.95 | area= small | maxDets=100 ] = 0.776
# Average Recall (AR) @[ IoU=0.50:0.95 | area=medium | maxDets=100 ] = -1.000
# Average Recall (AR) @[ IoU=0.50:0.95 | area= large | maxDets=100 ] = -1.000
# [08/14 21:04:34] ppdet.engine INFO: Total sample number: 636, averge FPS: 141.8495250691227
# [08/14 21:04:34] ppdet.engine INFO: Best test bbox ap is 0.668.
# ## 1.6 模型导出
#
# 将模型导出,并且打开`--export_serving_model`,适当能够生成`__model__`, `__params__`格式的模型与参数文件
#
# > 导出前, 需要前往: `configs/ppyolo/_base_/ppyolo_r18vd.yml`这个模型结构文件中,注释掉: `pretrain_weights`后再进行模型导出
#
# 如下:
#
# architecture: YOLOv3
# # pretrain_weights: https://paddledet.bj.bcebos.com/models/pretrained/ResNet18_vd_pretrained.pdparams
# norm_type: sync_bn
# use_ema: true
# ema_decay: 0.9998
# --export_serving_model指令下需要下载该依赖
# !pip install paddle-serving-client
# %cd /home/aistudio/PaddleDetection
# !python tools/export_model.py\
# -c configs/ppyolo/ppyolo_r18vd_coco.yml\
# -o weights='output/ppyolo_r18vd_coco/best_model'\
# --output_dir '/home/aistudio/export_model'\
# --export_serving_model True
# 查看输出结构
# !tree /home/aistudio/export_model -L 3
# 部署需要的内容主要有以下两种
#
# - `*.pdmodel` + `*.pdiparams`
#
# - `__model__` + `__params__`
#
# > 其它可能需要的资料(PaddleLite不直接用,可以作为加载的一些预处理参数的参考): `infer_cfg.yml`, `serving_server_conf.prototxt`
# ## 1.7 模型导出再windows端的部署效果检测
#
# 利用可视化推理验收模型效果:
#
# - **图片推理效果**
#
#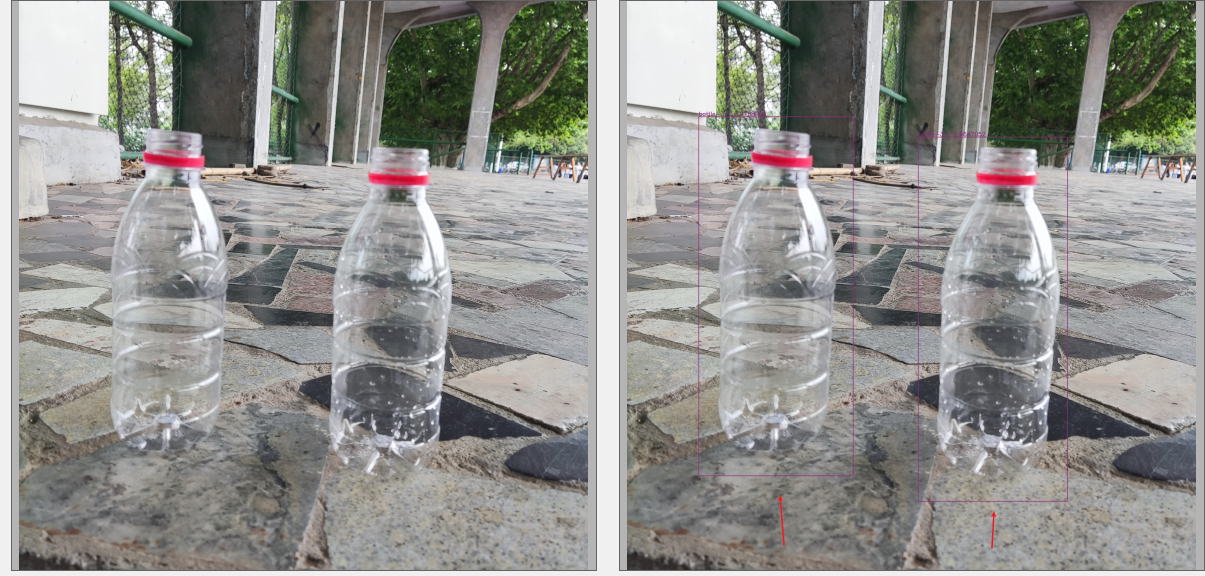 
#
# 
#
# - **视频推理效果**
#
#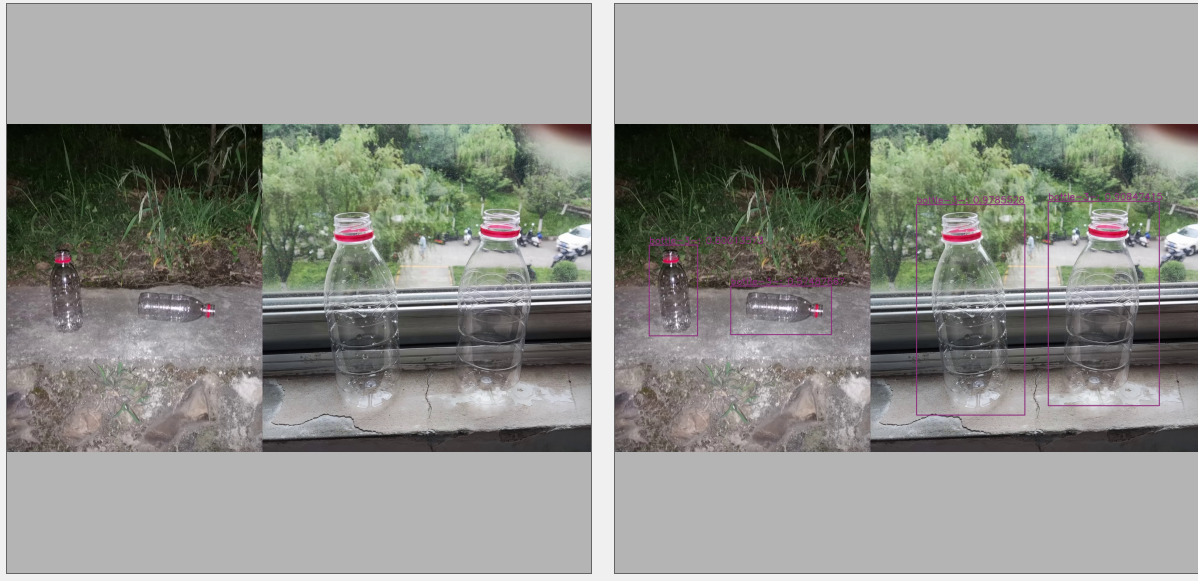 
#
#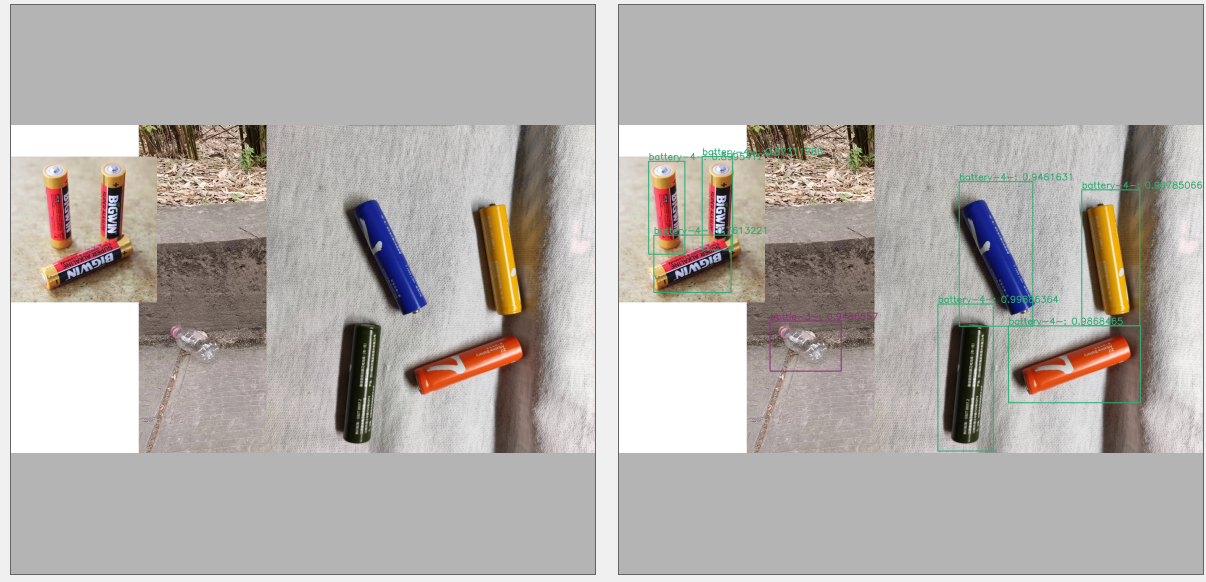 
#
# # 二、基于PPLite实现树莓派端部署
#
# 本项目训练的模型部署到树莓派4B上进行应用,能够实现较为快速准确的垃圾检测!
#
# > 部署说明与部分检测(展示为tiny的效果, 部署代码通用)的效果可以观看视频: [树莓派部署教程与效果展示](https://www.bilibili.com/video/BV1ph411r718?p=4)
#
# 部分效果:
#
# 
#
# ## 2.1 PaddleLite的python包安装
#
# 如果使用32位的操作系统,可以直接使用我编译好的whl(使用与Python3)
#
# 链接:[https://pan.baidu.com/s/1pmULmyNokBcG7EQz2gKWCg](https://pan.baidu.com/s/1pmULmyNokBcG7EQz2gKWCg)
#
# 提取码:plit
#
# ------
#
# 下载好后,上传到树莓派即可 —— 推荐使用`vnc远程服务`的文件传递功能。
#
# ------
#
# 安装指令:
#
# `python3 -m pip install whl_path`
# ## 2.2 部署流程
#
# - 1. 先使用`paddlelite包`中的`opt`这个API实现模型的转换,获取`nb格式`的文件
#
# - 2. 然后使用以下代码进行`模型加载`即可进行模型推理
# ## 2.3 部署代码
#
# 主要处理
#
# - 加载模型,并输出加载时间 `__init__`
#
# - 获取输入数据,配置模型输入 -- `get_input_img`
#
# - **注意不同模型的输入数据**
#
# - 获取绘制好框的图像结果 -- `get_output_img`
#
#
# > 部署代码,来自个人项目: [PPYolo-Tiny树莓派部署实践(一)](https://aistudio.baidu.com/aistudio/projectdetail/2047562)
#
# > 一些注意事项,可以看下面的代码,可以观看树莓派部署视频!
# +
from paddlelite.lite import *
import cv2 as cv
import numpy as np
from matplotlib import pyplot as plt
from time import time
from PIL import Image
from PIL import ImageFont
from PIL import ImageDraw
from PIL import ImageEnhance
class PPYOLO_Detector(object):
def __init__(self, nb_path = None, # nb路径
label_list = None, # 类别list
input_size = [320, 320], # 输入图像大小
img_means = [0., 0., 0.], # 图片归一化均值
img_stds = [0., 0., 0.], # 图片归一化方差
threshold = 0.1, # 预测阈值
num_thread = 1, # ARM CPU工作线程数
work_power_mode = PowerMode.LITE_POWER_NO_BIND # ARM CPU工作模式
):
# 验证必要的参数格式
assert nb_path is not None, \
"Please make sure the model_nb_path has inputed!(now, nb_path is None.)"
assert len(input_size) == 2, \
"Please make sure the input_shape length is 2, but now its length is {0}".format(len(input_size))
assert len(img_means) == 3, \
"Please make sure the image_means shape is [3], but now get image_means' shape is [{0}]".format(len(img_means))
assert len(img_stds) == 3, \
"Please make sure the image_stds shape is [3], but now get image_stds' shape is [{0}]".format(len(img_stds))
assert len([i for i in img_stds if i <= 0]) < 1, \
"Please make sure the image_stds data is more than 0., but now get image_stds' data exists less than or equal 0."
assert threshold > 0. and threshold < 1., \
"Please make sure the threshold value > 0. and < 1., but now get its value is {0}".format(threshold)
assert num_thread > 0 and num_thread <= 4, \
"Please make sure the num_thread value > 1 and <= 4., but now get its value is {0}".format(num_thread)
assert work_power_mode in [PowerMode.LITE_POWER_HIGH, PowerMode.LITE_POWER_LOW,
PowerMode.LITE_POWER_FULL, PowerMode.LITE_POWER_NO_BIND,
PowerMode.LITE_POWER_RAND_HIGH,
PowerMode.LITE_POWER_RAND_LOW], \
"Please make sure the work_power_mode is allowed , which is in \
[PowerMode.LITE_POWER_HIGH, PowerMode.LITE_POWER_LOW, \
PowerMode.LITE_POWER_FULL, PowerMode.LITE_POWER_NO_BIND, \
PowerMode.LITE_POWER_RAND_HIGH, \
PowerMode.LITE_POWER_RAND_LOW], \
but now get its value is {0}"
# 模型nb文件路径
self.model_path = nb_path
# ARM CPU工作线程数
self.num_thread = num_thread
# ARM CPU工作模式
self.power_mode = work_power_mode
# 预测显示阈值
self.threshold = threshold
# 预测输入图像大小
self.input_size = input_size
# 图片归一化参数
# 均值
self.img_means = img_means
# 方差
self.img_stds = img_stds
# 预测类别list
self.label_list = label_list
# 预测类别数
self.num_class = len(label_list) if (label_list is not None) and isinstance(label_list, list) else 1
# 类别框颜色map
self.box_color_map = self.random_colormap()
# 记录模型加载参数的开始时间
self.prepare_time = self.runtime()
# 配置预测
self.config = MobileConfig()
# 设置模型路径
self.config.set_model_from_file(nb_path)
# 设置线程数
self.config.set_threads(num_thread)
# 设置工作模式
self.config.set_power_mode(work_power_mode)
# 构建预测器
self.predictor = create_paddle_predictor(self.config)
# 模型加载参数的总时间花销
self.prepare_time = self.runtime() - self.prepare_time
print("The Prepare Model Has Cost: {0:.4f} s".format(self.prepare_time))
def get_input_img(self, input_img):
'''输入预测图片
input_img: 图片路径或者np.ndarray图像数据 - [h, w, c]
'''
assert isinstance(input_img, str) or isinstance(input_img, np.ndarray), \
"Please enter input is Image Path or numpy.ndarray, but get ({0}) ".format(input_img)
# 装载图像到预测器上的开始时间
self.load_img_time = self.runtime()
if isinstance(input_img, str):
# 读取图片路径下的图像数据
self.input_img = Image.open(input_img)
elif isinstance(input_img, np.ndarray):
# 读取ndarray数据下的图像数据
self.input_img = Image.fromarray(input_img)
# 获取图片原始高宽 : h,w
self.input_shape = np.asarray(self.input_img).shape[:-1]
# 重置图片大小为指定的输入大小
input_data = self.input_img.resize(self.input_size, Image.BILINEAR)
# 转制图像shape为预测指定shape
input_data = np.array(input_data).transpose(2, 0, 1).reshape([1, 3] + self.input_size).astype('float32')
# 将图像数据进行归一化
input_data = self.normlize(input_data)
self.scale_factor = [1., 1.] # [1., 1.]
# 配置输入tensor
# 输入[[shape, shape]]的图片大小
self.input_tensor0 = self.predictor.get_input(0)
self.input_tensor0.from_numpy(np.asarray([self.input_size], dtype=np.int32))
# 输入[1, 3, shape, shape]的归一化后的图片数据
self.input_tensor1 = self.predictor.get_input(1)
self.input_tensor1.from_numpy(input_data)
# 输入模型处理图像大小与实际图像大小的比例
self.input_tensor2 = self.predictor.get_input(2)
self.input_tensor2.from_numpy(np.asarray(self.scale_factor, dtype=np.int32))
# 装载图像到预测器上的总时间花销
self.load_img_time = self.runtime() - self.load_img_time
print("The Load Image Has Cost: {0:.4f} s".format(self.load_img_time))
def get_output_img(self, num_bbox=1):
'''获取输出标注图片
num_bbox: 最大标注个数
'''
# 预测器开始预测的时间
self.predict_time = self.runtime()
# 根据get_input_img的图像进行预测
self.predictor.run()
# 获取输出预测bbox结果
self.output_tensor = self.predictor.get_output(0)
# 转化为numpy格式
output_bboxes = self.output_tensor.numpy()
# 根据阈值进行筛选,大于等于阈值的保留
output_bboxes = output_bboxes[output_bboxes[:, 1] >= self.threshold]
# 根据预测结果进行框绘制,返回绘制完成的图片
self.output_img = self.load_bbox(output_bboxes, num_bbox)
# 预测器预测的总时间花销
self.predict_time = self.runtime() - self.predict_time
print("The Predict Image Has Cost: {0:.4f} s".format(self.predict_time))
return self.output_img
def normlize(self, input_img):
'''数据归一化
input_img: 图像数据--numpy.ndarray
'''
# 对RGB通道进行均值-方差的归一化
input_img[0, 0] = (input_img[0, 0] / 255. - self.img_means[0]) / self.img_stds[0]
input_img[0, 1] = (input_img[0, 1] / 255. - self.img_means[1]) / self.img_stds[1]
input_img[0, 2] = (input_img[0, 2] / 255. - self.img_means[2]) / self.img_stds[2]
return input_img
def load_bbox(self, input_bboxs, num_bbox):
'''根据预测框在原始图片上绘制框体,并标注
input_bboxs: 预测框
num_bbox: 允许的标注个数
'''
# 创建间绘图参数:[cls_id, score, x1, y1, x2, y2]
self.draw_bboxs = [0] * 6
# 绘图器 -- 根据get_input_img的输入图像
draw = ImageDraw.Draw(self.input_img)
# 根据最大标注个数进行实际标注个数的确定
# input_bboxs.shape[0]: 表示预测到的有效框个数
if len(input_bboxs) != 0: # 存在有效框时
num_bbox = input_bboxs.shape[0] if num_bbox > input_bboxs.shape[0] else num_bbox
else:
num_bbox = 0 # 没有有效框,直接不标注
# 遍历框体,并进行标注
for i in range(num_bbox):
# 类别信息
self.draw_bboxs[0] = input_bboxs[i][0]
# 类别得分
self.draw_bboxs[1] = input_bboxs[i][1]
print(self.label_list[int(self.draw_bboxs[0])], '- score{', self.draw_bboxs[1], "} : ", input_bboxs[i][2], input_bboxs[i][3], input_bboxs[i][4], input_bboxs[i][5])
# 框体左上角坐标
# max(min(input_bboxs[i][2] / self.input_size[0], 1.), 0.):保证当前预测坐标始终在图像内(比例,0.-1.)
# max(min(input_bboxs[i][2] / self.input_size[0], 1.), 0.) * self.input_shape[1]: 直接预测得到的坐标
# min(max(min(input_bboxs[i][2] / self.input_size[0], 1.), 0.) * self.input_shape[1], self.input_shape[1]):保证坐标在图像内(h, w)
self.draw_bboxs[2] = min(max(min(input_bboxs[i][2] / self.input_size[0], 1.), 0.) * self.input_shape[1], self.input_shape[1])
self.draw_bboxs[3] = min(max(min(input_bboxs[i][3] / self.input_size[1], 1.), 0.) * self.input_shape[0], self.input_shape[0])
# 框体右下角坐标
self.draw_bboxs[4] = min(max(min(input_bboxs[i][4] / self.input_size[0], 1.), 0.) * self.input_shape[1], self.input_shape[1])
self.draw_bboxs[5] = min(max(min(input_bboxs[i][5] / self.input_size[1], 1.), 0.) * self.input_shape[0], self.input_shape[0])
# print(self.draw_bboxs[2], self.draw_bboxs[3], self.draw_bboxs[4], self.draw_bboxs[5])
# 绘制框体
# self.box_color_map[int(self.draw_bboxs[i][0])]: 对应类别的框颜色
draw.rectangle(((self.draw_bboxs[2], self.draw_bboxs[3]),
(self.draw_bboxs[4], self.draw_bboxs[5])),
outline = tuple(self.box_color_map[int(self.draw_bboxs[0])]),
width =2)
# 框体位置写上类别和得分信息
draw.text((self.draw_bboxs[2], self.draw_bboxs[3]+1),
"{0}:{1:.4f}".format(self.label_list[int(self.draw_bboxs[0])], self.draw_bboxs[1]),
tuple(self.box_color_map[int(self.draw_bboxs[0])]))
# 返回标注好的图像数据
return np.asarray(self.input_img)
def random_colormap(self):
'''获取与类别数量等量的color_map
'''
np.random.seed(2021)
color_map = [[np.random.randint(20, 255),
np.random.randint(64, 200),
np.random.randint(128, 255)]
for i in range(self.num_class)]
return color_map
def runtime(self):
'''返回当前计时
'''
return time()
# -
# ## 2.4 部署测试代码片段
#
# > 有需要的小伙伴,可以搭载串口通信,实现树莓派与单片机直接的通信哦!
def test():
model_path = "/home/pi/test/ppyolo_tiny/ppyolo_tiny.nb" # 模型参数nb文件 -- 自行修改
img_path = "/home/pi/Desktop/citrus_0005.jpg" # 自己的预测图像
label_list = ['bottle', 'battery', 'cup', 'paper', 'citrus'] # 类别list
input_size = [224, 224] # 输入图像大小
img_means = [0.485, 0.456, 0.406] # 图片归一化均值
img_stds = [0.229, 0.224, 0.225] # 图片归一化方差
threshold = 0.1 # 预测阈值
num_thread = 2 # ARM CPU工作线程数
work_mode = PowerMode.LITE_POWER_NO_BIND # ARM CPU工作模式
max_bbox_num = 1 # 每帧最多标注数
# 创建预测器
detector = PPYOLO_Detector(
nb_path = model_path,
label_list = label_list,
input_size = input_size,
img_means = img_means,
img_stds = img_stds,
threshold = threshold,
num_thread = num_thread,
work_power_mode = PowerMode.LITE_POWER_NO_BIND
)
img = plt.imread(img_path)
img = cv.resize(img,(320, 320)) # 与训练时配置的大小一致
detector.get_input_img(img) # 输入图片数据
img = detector.get_output_img(num_bbox = max_bbox_num) # 得到预测输出后绘制了框的图像
plt.imshow(img)
plt.show()
# ## 2.5 部署效果展示
#
# 请看传送门: [树莓派部署教程与效果展示](https://www.bilibili.com/video/BV1ph411r718?p=4)
# # 三、项目总结
#
# 一直以来,人工智能落地都是一个工业界的热门话题。近年内,有许多优秀的检测算法出现,YOLOV4-PPYOLO-PPYOLOV2等。但在落地时,不光要考虑精度,还需要实时性——也就是考虑部署设备的算力情况。
#
# 因此,本项目就基于PPDet展开了较轻量化模型PPyolo_r18的模型训练来实现垃圾的检测分类,同时利用PaddleLite完成在树莓派端的部署,实现分拣车的视觉关键部分。
#
# 通过本项目,可以对目标检测落地提供一个可行的方案,也是基于python3的PaddleLite部署Paddle模型的一个部署实践方案。
#
# ------
#
# **主要收获的点如下:**
#
# - 1. 数据集不大时,可以先用`大的batch_size`跑`1/3的轮次`稳定损失下降,然后利用`1/2的batch_size`进行接下来的轮次训练,实现继续优化
#
# - 2. 数据输入大小影响着模型处理的快慢,输入图像越小,模型推理越快
#
# - 3. 部署时,根据需要可进行`量化处理``(支持`INT8`, `INT16`),实现更快的模型加载和保存(模型体积减小).
# # 个人介绍
#
# > 姓名:蔡敬辉
#
# > 学历:大三(在读)
#
# > 爱好:喜欢参加一些大大小小的比赛,不限于计算机视觉——有共同爱好的小伙伴可以关注一下哦~后期会持续更新一些自制的竞赛baseline和一些竞赛经验分享
#
# > 主要方向:目标检测、图像分割与图像识别
#
# > 联系方式:qq:3020889729 微信:cjh3020889729
#
# > 学校:西南科技大学
| 2283361.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import trimesh
# load a file by name or from a buffer
mesh = trimesh.load_mesh('../models/featuretype.STL')
# is the current mesh watertight?
mesh.is_watertight
# what's the euler number for the mesh?
mesh.euler_number
# the convex hull is another Trimesh object that is available as a property
# lets compare the volume of our mesh with the volume of its convex hull
np.divide(mesh.volume, mesh.convex_hull.volume)
# since the mesh is watertight, it means there is a
# volumetric center of mass which we can set as the origin for our mesh
mesh.vertices -= mesh.center_mass
# what's the moment of inertia for the mesh?
mesh.moment_inertia
# if there are multiple bodies in the mesh we can split the mesh by
# connected components of face adjacency
# since this example mesh is a single watertight body we get a list of one mesh
mesh.split()
# preview mesh in a pyglet window from a terminal, or inline in a notebook
mesh.show()
# facets are groups of coplanar adjacent faces
# set each facet to a random color
# colors are 8 bit RGBA by default (n,4) np.uint8
for facet in mesh.facets:
mesh.visual.face_colors[facet] = trimesh.visual.random_color()
# transform method can be passed a (4,4) matrix and will cleanly apply the transform
mesh.apply_transform(trimesh.transformations.random_rotation_matrix())
# an axis aligned bounding box is available
mesh.bounding_box.primitive.extents
# a minimum volume oriented bounding box is available
mesh.bounding_box_oriented.primitive.extents
mesh.bounding_box_oriented.primitive.transform
# the bounding box is a trimesh.primitives.Box object, which subclasses
# Trimesh and lazily evaluates to fill in vertices and faces when requested
mesh.bounding_box_oriented.show()
# bounding spheres and bounding cylinders of meshes are also
# available, and will be the minimum volume version of each
# except in certain degenerate cases, where they will be no worse
# than a least squares fit version of the primitive.
print(mesh.bounding_box_oriented.volume,
mesh.bounding_cylinder.volume,
mesh.bounding_sphere.volume)
| examples/quick_start.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import glob
dados[0].columns.values[0]
path = 'C:\\mbafiabigdatat8\\zika-data\\data\\parsed\\brazil\\*.csv'
files = glob.glob(path)
dados_estados = list()
dados_resumo = list()
for name in files:
pd1 = pd.read_csv(name)
if pd1.columns.values[0] == 'no':
dados_estados.append(pd1)
else:
dados_resumo.append(pd1)
dados_resumo
data = pd.read_csv("C:\\mbafiabigdatat8\\zika-data\\data\\parsed\\brazil\\brazil-microcephaly-2016-01-23-table-1.csv")
data
| data/parsed/brazil/parsed_brazil.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TO DO
#
# - [ ] re-estimate without buildings constructed in 1979
# - [ ] re-estimate with diff census attrs
# - [x] re-estimate w/ + w/o buildings built < 2007
# - [ ] check variability of census attrs over time
# - [ ] map of eviction data (hexes)
# - [x] redo table 1 by rates across all addresses
# - [x] redo rd plot for pre 2013 evictions and post 2013 evictions
# - [x] subtract evictions in year 2017??
# - [ ] dedupe evictions at same addr on same date for same tenants? shouldn't be greater than max unit count
# - [ ] use householder exemption in assessor records to infer ownership!
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
import seaborn as sns
import geopandas as gpd
from shapely.geometry import Point
import statsmodels.formula.api as smf
# %matplotlib inline
# # Load data
asr = pd.read_csv('../../../2021_02_summer/evic_paper/data/asr_grouped_by_yr_w_geom.csv', dtype={'fipscd': str})
ev = pd.read_csv('../../../2021_02_summer/evic_paper/data/ev_matched_w_fips.csv')
asr_all = pd.read_csv('../data/assessor_2007-2016_fips_jun_2021.csv')
# # Validate pre-processing
# \# matched evictions should match the number of evictions in the assessor file
assert len(ev[~pd.isnull(ev['asr_index'])]) == asr['ev_count'].sum()
# Only counting evictions after 2007
assert (asr['ev_count'] == asr['ev_count_post_07']).all()
# Make sure we're comparing like to like
assert ev['year'].min() == 2007
assert ev['year'].max() == 2016
assert asr['asr_yr'].min() == 2007
assert asr['asr_yr'].max() == 2016
# # Describe data processing/filtering steps
print('{0} eviction records after cleaning'.format(ev.shape[0]))
print("{0} total assessor records considered".format(asr_all.shape[0]))
print("{0} assessor records after grouping by address and assessed year".format(asr.shape[0]))
print("{0} evictions ({1}%) were matched to assessor records".format(
len(ev[~pd.isnull(ev['asr_index'])]),
np.round(len(ev[~pd.isnull(ev['asr_index'])]) / len(ev) * 100,1)))
asr_rc_eligible = asr[asr['any_rc_eligibility'] == 1] # all rc eligible
print("{0} evictions across {1} rent control eligible assessor records".format(asr_rc_eligible['ev_count'].sum(), len(asr_rc_eligible)))
asr_rc_eligible_valid_units = asr_rc_eligible[asr_rc_eligible['total_units'] > 0]
print("{0} evictions assigned to {1} rent control eligible assessor records with valid unit counts".format(asr_rc_eligible_valid_units['ev_count'].sum(), len(asr_rc_eligible_valid_units)))
print("{0} assessor records with {1} evictions dropped due to unreliable unit counts".format(
asr_rc_eligible.shape[0] - asr_rc_eligible_valid_units.shape[0],
asr_rc_eligible.loc[asr_rc_eligible['total_units']<=0, 'ev_count'].sum()))
asr['any_ev'] = (asr['ev_count'] > 0).astype(int)
asr['pre_1980'] = (asr['year_built_max'] < 1980)
asr['built_1980'] = None
asr.loc[asr['pre_1980'], 'built_1980'] = 'before'
asr.loc[~asr['pre_1980'], 'built_1980'] = 'after'
asr['ev_per_unit'] = asr['ev_count'] / asr['total_units']
ev = ev.merge(asr, left_on='asr_index', right_on='index', suffixes=('_ev', '_asr'))
print(ev.shape)
ev = ev[(ev['total_units'] > 0)]
print(ev.shape)
ev = ev[(ev['any_rc_eligibility'] == 1)]
print(ev.shape)
# #### Table 1
# +
ev.loc[pd.isnull(ev['type']), 'type'] = 'unknown'
type_counts = ev.groupby(['built_1980', 'type']).agg(count=('index_ev', 'nunique')).reset_index()
pre_sums = type_counts.groupby('built_1980')['count'].sum()
ev['ev_type_cat'] = 'breach of lease'
ev.loc[ev['type'].isin([
'OMI', 'Capital Improvement', 'ELLIS', 'Condo Conversion', 'Substantial Rehabilitation',
'Lead Remediation', 'Good Samaritan Tenancy Ends',
'Development Agreement', 'Demolition']), 'ev_type_cat'] = 'no fault'
ev.loc[ev['type'].isin(['unknown', 'Other']), 'ev_type_cat'] = 'unknown/Other'
# -
cat_counts = ev.groupby(['built_1980', 'ev_type_cat']).agg(count=('index_ev', 'nunique')).reset_index()
cat_counts = cat_counts.pivot(index='ev_type_cat', columns='built_1980', values='count')
cat_counts['pct_ev_after'] = cat_counts['after'] / pre_sums['after']
cat_counts['pct_ev_before'] = cat_counts['before'] / pre_sums['before']
cat_counts[['before','pct_ev_before','after','pct_ev_after']]
cat_counts['before'].sum()
cat_counts['after'].sum()
13774 + 189
# #### Table 2
# +
mean_diffs = asr[
(asr['total_units'] > 0)].groupby(['any_rc_eligibility', 'pre_1980']).agg(
total_addresses=('index', 'count'),
total_units=('total_units', 'sum'),
mean_units_per_address=('total_units','mean'),
total_evictions=('ev_count', 'sum'),
mean_any_ev=('any_ev', 'mean'),
mean_ev_per_address=('ev_count','mean'),
mean_ev_per_unit=('ev_per_unit','mean'),
)
# mean_diffs['units_per_address'] = mean_diffs['total_units'] / mean_diffs['total_addresses']
# mean_diffs['evictions_per_address'] = mean_diffs['total_evictions'] / mean_diffs['total_addresses']
# mean_diffs['evictions_per_unit'] = mean_diffs['total_evictions'] / mean_diffs['total_units']
mean_diffs
# -
cat_counts['ev_per_unit_before'] = cat_counts['before'] / mean_diffs.loc[1.0, True]['total_units']
cat_counts['ev_per_unit_after'] = cat_counts['after'] / mean_diffs.loc[1.0, False]['total_units']
cat_counts
cat_counts['mean_diff'] = cat_counts['ev_per_unit_before'] - cat_counts['ev_per_unit_after']
cat_counts
cat_counts['mean_diff'].sum()
# #### Fig 1
sns.set_style({'font.family': 'Times New Roman'})
# +
units_by_yr = asr[
(asr['any_rc_eligibility'] == 1) &
(asr['year_built_max'] > 1900) &
(asr['year_built_max'] < 2100)].groupby('year_built_max').agg({'total_units': 'sum'}).reset_index()
fig, ax = plt.subplots(figsize=(10,5))
plt.rcParams["font.family"] = 'times new roman'
ax.scatter(units_by_yr['year_built_max'], units_by_yr['total_units'], s=25, facecolors='none', edgecolors='r')
ax.plot(units_by_yr['year_built_max'], units_by_yr['total_units'], lw=1, c='k', )
_ = ax.set_xlabel("year built", fontsize=12)
_ = ax.set_ylabel("# new units", fontsize=12)
# _ = ax.set_title("SF New Construction: Rent-control eligible use-codes", fontsize=20)
plt.savefig('../../../2021_02_summer/evic_paper/fig_1.jpg', dpi=300, bbox_inches='tight')
# -
# #### Fig 2
# +
rc_pop = asr[
(asr['any_rc_eligibility'] == 1) & (asr['year_built_max'] > 1953) &
(asr['year_built_max'] < 2007) & (asr['total_units'] > 0)]
yr_vs_ev = rc_pop.groupby('year_built_max').agg({
'ev_per_unit': 'mean',
}).reset_index()
yr_vs_ev1 = yr_vs_ev[(yr_vs_ev['year_built_max'] < 1980) &
(yr_vs_ev['year_built_max'] >= 1953)]
yr_vs_ev2 = yr_vs_ev[(yr_vs_ev['year_built_max'] >= 1980) &
(yr_vs_ev['year_built_max'] <= 2007)]
# -
rc_pop.shape
# +
fig, ax = plt.subplots(figsize=(10,5))
sns.regplot(
x='year_built_max', y='ev_per_unit', data=yr_vs_ev1, ax=ax, truncate=True,
label='rent controlled', line_kws={'color':'k', 'lw':1, 'label':'linear fit (c.i. = 95%)', 'zorder': 0})
sns.regplot(
x='year_built_max', y='ev_per_unit', data=yr_vs_ev2, ax=ax, truncate=True,
label='non-rent controlled', line_kws={'color':'k', 'lw':'1', 'zorder': 0})
ax.axvline(1979.5, ls=':', c='r', label='rent control built-year\n eligibility threshold')
ax.legend()
_ = ax.set_xlabel("property built-year", fontsize=12)
_ = ax.set_ylabel("avg. evictions per unit per year", fontsize=12,
# rotation=0,
# labelpad=70
)
# _ = ax.set_title("SF Eviction Rates (2007-2016)\nfor Multi-family Residential Addresses", fontsize=20)
ax.set_ylim((-0.005, 0.05))
ax.set_xlim((1952, 2008))
# ax.annotate('rent control \nbuilt-year threshold', xy=(1979, 0.04), xycoords='data',
# xytext=(0.3, 0.8), textcoords='axes fraction',
# arrowprops=dict(facecolor='black',headlength=10, width=0.5, headwidth=10),
# horizontalalignment='center', verticalalignment='center', fontsize=12
# )
plt.savefig('../../../2021_02_summer/evic_paper/fig_2.jpg', dpi=300, bbox_inches='tight')
# -
# #### Fig 3
sf = gpd.read_file('../../../2021_02_summer/evic_paper/sf_boundary.shp')
ev['geometry'] = ev.apply(lambda x: Point(x['longitude_asr'], x['latitude_asr']), axis=1)
ev_gpd = gpd.GeoDataFrame(ev, geometry='geometry')
ev_gpd.crs = 'EPSG:4326'
ev_gpd = ev_gpd.to_crs(sf.crs)
ev_gpd['proj_x'] = ev_gpd.geometry.x
ev_gpd['proj_y'] = ev_gpd.geometry.y
fig, ax = plt.subplots(figsize=(10,10))
sf.plot(facecolor='None', edgecolor='k', ax=ax, lw=0.5, alpha=0.8)
hb = ax.hexbin(x=ev_gpd['proj_x'], y=ev_gpd['proj_y'], mincnt=1,
bins=[0,20,40,60,80,100,120,140,160,180,200],
gridsize=18)
cb = fig.colorbar(hb, ax=ax, fraction=0.038,pad=0.01)
cb.set_label('count', fontsize=12)
cb.set_ticks([1,2.25,3.5,4.75,6,7.25,8.5,9.75,11])
cb.set_ticklabels([1,25, 50,75,100,125,150,175,'> 200+'])
ax.axis('off')
plt.savefig('../../../2021_02_summer/evic_paper/fig_3.jpg', dpi=300, bbox_inches='tight')
# #### Appendix A
# +
ev.loc[pd.isnull(ev['type']), 'type'] = 'unknown'
type_counts = ev.groupby(['built_1980', 'type']).agg(count=('index_ev', 'nunique')).reset_index()
pre_sums = type_counts.groupby('built_1980')['count'].sum()
type_counts = type_counts.pivot(index='type', columns='built_1980', values='count')
type_counts['pct_after'] = np.round(type_counts['after'] / pre_sums['after'] * 100, 1)
type_counts['pct_before'] = np.round(type_counts['before'] / pre_sums['before'] * 100, 1)
type_counts.reset_index(inplace=True)
type_counts.set_index('type', inplace=True, drop=True)
# -
type_counts = type_counts.sort_values('pct_before', ascending=False)[['before','pct_before','after','pct_after']]
type_counts = type_counts.astype(str)
type_counts['after'] = type_counts['after'].str.split('.').str[0]
type_counts['before'] = type_counts['before'].str.split('.').str[0]
type_counts = type_counts.replace('nan', '--', regex=True)
type_counts
# # RD
# +
bandwidth = 27
df = asr[
(asr['any_rc_eligibility'] == 1) & (asr['year_built_max'] > 1980 - bandwidth) &
(asr['year_built_max'] < 1980 + bandwidth) & (asr['total_units'] > 0)].copy()
# -
df.shape
54388 - 53493
df['rent_control'] = False
df.loc[df['pre_1980'] == True, 'rent_control'] = True
df['year_built_centered'] = df['year_built_max'] - 1980
df['ev_count'].sum()
# ### Model 1
rd = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control",
data=df)
fitted = rd.fit()
print(fitted.summary())
# ### Model 2: Standard RDD + Property Characteristics
rd2_df = df[(df['total_sqft'] > 50) & (df['total_value'] > 100)].copy()
rd2 = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control + "
"np.log(total_value):np.log(total_sqft) + np.log(total_units)",
data=rd2_df)
fitted2 = rd2.fit()
print(fitted2.summary())
# ### Model 3: Standard RDD + Property Characteristics + Census Tract Attributes
census_df = pd.read_csv('../data/census_tract_acs_data.csv', dtype={'tract_id': str})
asr['tract_id'] = asr['fipscd'].astype(str).str.zfill(15).str[:11]
asr = pd.merge(asr, census_df, on='tract_id', how='left')
dropped_parcels = len(asr[pd.isnull(asr['median_hh_income'])])
dropped_evs = asr[pd.isnull(asr['median_hh_income'])]['ev_count'].sum()
print('Dropped {0} parcels ({1}%) and {2} evictions merging on tract-level census data'.format(
dropped_parcels, np.round(dropped_parcels/len(asr)*100, 2), dropped_evs))
rd3_df = asr[
(asr['any_rc_eligibility'] == 1) & (asr['year_built_max'] > 1980 - bandwidth) &
(asr['year_built_max'] < 1980 + bandwidth) & (asr['total_units'] > 0) &
(asr['total_sqft'] > 50) & (asr['total_value'] > 100)].copy()
rd3_df['rent_control'] = False
rd3_df.loc[rd3_df['pre_1980'] == True, 'rent_control'] = True
rd3_df['year_built_centered'] = rd3_df['year_built_max'] - 1980
rd3_df['pct_non_white'] = 1 - (rd3_df['white_only_pop'] / rd3_df['total_pop_race'])
rd3_df['pct_black'] = (rd3_df['black_only_pop'] / rd3_df['total_pop_race'])
rd3_df['pct_black_multi'] = (rd3_df['black_only_pop'] + rd3_df['multiracial_pop']) / rd3_df['total_pop_race']
rd3_df['pct_latino'] = rd3_df['hispanic_pop'] / rd3_df['total_pop_race']
rd3_df['pct_occ_units_rental'] = rd3_df['total_rental_tenure'] / rd3_df['total_occupied_units']
rd3_df['pct_renter_pop'] = rd3_df['total_renter_pop'] / rd3_df['total_pop_tenure']
rd3_df['pct_rentals_pre2000_movein'] = (
rd3_df['total_rental_tenure'] - rd3_df['renter_movein_2005_later'] - rd3_df['renter_movein_2000_2004']) / (
rd3_df['total_rental_tenure'])
rd3_df['pct_renter_same_house_last_yr'] = rd3_df['renter_non_mover'] / rd3_df['total_renter_pop']
rd3_df = rd3_df[(rd3_df['median_movein_yr_owners'] > 0) & (rd3_df['median_movein_yr_renters'] > 0)]
rd3_df['yrs_before_2011_median_movein_yr_rent'] = rd3_df['median_movein_yr_renters'] - rd3_df['median_movein_yr_renters']
rd3_df['yrs_before_2011_median_movein_yr_own'] = 2011 - rd3_df['median_movein_yr_owners']
rd3_df['median_movein_yr_rent_std'] = (rd3_df['median_movein_yr_renters'] - rd3_df['median_movein_yr_renters'].mean()) / rd3_df['median_movein_yr_renters'].std()
rd3_df['median_movein_yr_own_std'] = (rd3_df['median_movein_yr_owners'] - rd3_df['median_movein_yr_owners'].mean()) / rd3_df['median_movein_yr_owners'].std()
rd3_df['pct_recent_mover_rent'] = 1 - (rd3_df['renter_non_mover'] / rd3_df['total_renter_pop'])
rd3 = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control + "
"np.log(total_value):np.log(total_sqft) + np.log(total_units) + "
"np.log(median_hh_income) + "
"pct_non_white + "
"pct_occ_units_rental + "
"pct_rentals_pre2000_movein",
data=rd3_df)
fitted3 = rd3.fit()
print(fitted3.summary())
rd3b = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control + "
"np.log(total_value):np.log(total_sqft) + np.log(total_units) + "
# "np.log(median_hh_income) + "
"pct_latino +"
# "median_movein_yr_rent_std +"
# "pct_recent_mover_rent +"
"pct_occ_units_rental",
# "pct_rentals_pre2000_movein",
data=rd3_df)
fitted3b = rd3b.fit()
print(fitted3b.summary())
# ### Model 4: Standard RDD + Property Characteristics + Census Tract Attributes + Neighborhood Fixed Effects
rd4_df = rd3_df.copy()
rd4 = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control + "
"np.log(total_value):np.log(total_sqft) + np.log(total_units) + "
"pct_latino + "
"pct_occ_units_rental + "
"nbd_code",
data=rd4_df)
fitted4 = rd4.fit()
print(fitted4.summary())
rd4b = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control + "
"np.log(total_value):np.log(total_sqft) + np.log(total_units) + "
"pct_latino + "
"pct_occ_units_rental + "
"nbd_code",
data=rd4_df)
fitted4b = rd4b.fit()
print(fitted4b.summary())
rd4_df['geometry'] = rd4_df.apply(lambda x: Point(x['longitude'], x['latitude']), axis=1)
rd4_gdf = gpd.GeoDataFrame(rd4_df, geometry='geometry')
rd4_gdf.crs = 'EPSG:4326'
nbd_gdf = gpd.read_file('/Users/max/Documents/cal/2021_02_summer/evic_paper/SF Find Neighborhoods/geo_export_b9533012-1016-4bb8-a58a-ce8a0b05319f.shp')
rd4_nbd_df = gpd.sjoin(rd4_gdf, nbd_gdf,op='within', how='inner')
rd4_nbd_df.shape
nbd_gdf.columns
rd4_nbd_df[rd4_nbd_df['name'] == 'Chinatown']['ev_count'].sum()
# ### rerun models on exact same data
rd_rd4_df = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control",
data=rd4_df)
rd_rd4_df_fitted = rd_rd4_df.fit()
print(rd_rd4_df_fitted.summary())
rd2_rd4_df = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control + "
"np.log(total_value):np.log(total_sqft) + np.log(total_units)",
data=rd4_df)
rd2_rd4_df_fitted = rd2_rd4_df.fit()
print(rd2_rd4_df_fitted.summary())
rd4_df['median_hh_income_std'] = (rd4_df['median_hh_income'] - rd4_df['median_hh_income'])/
rd3_rd4_df = smf.ols(
"ev_per_unit ~ rent_control + year_built_centered*rent_control + "
"np.log(total_value):np.log(total_sqft) + np.log(total_units) + "
"pct_latino +"
"pct_occ_units_rental",
data=rd4_df)
rd3_rd4_df_fitted = rd3_rd4_df.fit()
print(rd3_rd4_df_fitted.summary())
df[~df['index'].isin(rd4_df.index)]['ev_count'].sum()
df[~df['index'].isin(rd4_df.index)].shape
rd4_df.groupby('rent_control')['ev_per_unit'].mean()
| notebooks/analysis_final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
X_train, Y_train = mnist.train.next_batch(5000)
X_test, Y_test = mnist.test.next_batch(200)
xtr = tf.placeholder("float",[None, 784])
xte = tf.placeholder("float",[784])
# +
# Calculate L1 distance
distance = tf.reduce_sum(tf.abs(tf.add(xtr, tf.negative(xte))), reduction_indices = 1)
pred = tf.arg_min(distance, 0)
accuracy = 0.
init = tf.global_variables_initializer()
# -
with tf.Session() as sess:
sess.run(init)
for i in range(len(X_test)):
nn_index = sess.run(pred, feed_dict={xtr:X_train, xte:X_test[i, :]})
print "Test", i, "Prediction:", np.argmax(Y_train[nn_index]), \
"True Class:", np.argmax(Y_test[i])
if np.argmax(Y_train[nn_index])== np.argmax(Y_test[i]):
accuracy += 1./len(X_test)
print accuracy
| nearestNeighbour.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import the Libraries
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
# Load the Dataset
dataset=pd.read_csv('ex1data1.txt')
X=dataset.iloc[:,:-1].values
y=dataset.iloc[:,-1].values
# Split the dataset
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test=train_test_split(X,y,test_size=0.3,random_state=0)
# Train Linear Regression using Skicit Library
from sklearn.linear_model import LinearRegression
model=LinearRegression()
model.fit(X_train,y_train)
# Predict
result=model.predict(X_test)
result
# Plot the Training set
plt.scatter(X_train,y_train,color='red')
plt.plot(X_train,model.predict(X_train),color='blue')
plt.xlabel('Population in 10,000')
plt.ylabel('Profit in $10,000')
plt.title('Training Data')
plt.show()
plt.scatter(X_test,y_test,color='yellow')
plt.plot(X_train,model.predict(X_train),color='blue')
plt.xlabel('Population in 10,000')
plt.ylabel('Profit in $10,000')
plt.title('Test Data')
plt.show()
| Linear Regression/LinearRegressionSingle Variables.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Missing values
#
# Missing data, or Missing values, occur when __no data__ / __no value__ is stored for a certain observation within a variable.
#
# Missing data are a common occurrence both in data science competitions and in data in business settings, and can have a significant effect on the conclusions that can be drawn from the data. Incomplete data is an unavoidable problem in dealing with most data sources.
#
# ### Why is data missing?
#
# The source of missing data can be very different and here are just a few examples:
#
# - A value is missing because it was forgotten or lost or not stored properly
# - For a certain observation, the value of the variable does not exist
# - The value can't be known or identified
#
# Imagine for example that the data comes from a survey, and the data are entered manually into an online form. The data entry could easily forget to complete a field in the form, and therefore, that value for that form would be missing.
#
# The person being asked may not want to disclose the answer to one of the questions, for example, their income. That would be then a missing value for that person.
#
# Sometimes, a certain feature can't be calculated for a specific individual. For example, in the variable 'total debt as percentage of total income' if the person has no income, then the total percentage of 0 does not exist. Therefore it will be a missing value.
#
# Together with understanding the source of missing data, it is important to understand the mechanisms by which missing fields are introduced in a dataset. Depending on the mechanism, we may choose to process the missing values differently. In addition, by knowing the source of missing data, we may choose to take action to control that source, and decrease the number of missing data looking forward during data collection.
#
#
# ### Missing Data Mechanisms
#
# There are 3 mechanisms that lead to missing data, 2 of them involve missing data randomly or almost-randomly, and the third one involves a systematic loss of data.
#
# #### Missing Completely at Random, MCAR:
#
# A variable is missing completely at random (MCAR) if the probability of being missing is the same for all the observations.
# When data is MCAR, there is absolutely no relationship between the data missing and any other values, observed or missing, within the dataset. In other words, those missing data points are a random subset of the data. There is nothing systematic going on that makes some data more likely to be missing than other.
#
# If values for observations are missing completely at random, then disregarding those cases would not bias the inferences made.
#
#
# #### Missing at Random, MAR:
#
# MAR occurs when there is a systematic relationship between the propensity of missing values and the observed data. In other words, the probability an observation being missing depends only on available information (other variables in the dataset). For example, if men are more likely to disclose their weight than women, weight is MAR. The weight information will be missing at random for those men and women that decided not to disclose their weight, but as men are more prone to disclose it, there will be more missing values for women than for men.
#
# In a situation like the above, if we decide to proceed with the variable with missing values (in this case weight), we might benefit from including gender to control the bias in weight for the missing observations.
#
# #### Missing Not at Random, MNAR:
#
# Missing of values is not at random (MNAR) if their being missing depends on information not recorded in the dataset. In other words, there is a mechanism or a reason why missing values are introduced in the dataset.
#
# Examples:
#
# MNAR would occur if people failed to fill in a depression survey because of their level of depression. Here, the missing of data is related to the outcome, depression.
#
# When a financial company asks for bank and identity documents from customers in order to prevent identity fraud, typically, fraudsters impersonating someone else will not upload documents, because they don't have them, precisely because they are fraudsters. Therefore, there is a systematic relationship between the missing documents and the target we want to predict: fraud.
#
# Understanding the mechanism by which data can be missing is important to decide which methods to use to handle the missing values. I will cover how to handle missing values in detail sections 5 and 6.
#
#
#
# ## Real Life example:
#
# ### Predicting Survival on the Titanic: understanding society behaviour and beliefs
#
# Perhaps one of the most infamous shipwrecks in history, the Titanic sank after colliding with an iceberg, killing 1502 out of 2224 people on board. Interestingly, by analysing the probability of survival based on few attributes like gender, age, and social status, we can make very accurate predictions on which passengers would survive. Some groups of people were more likely to survive than others, such as women, children, and the upper-class. Therefore, we can learn about the society priorities and privileges at the time.
#
# ### Peer to peer lending: Finance
#
# Lending Club is a peer-to-peer Lending company based in the US. They match people looking to invest money with people looking to borrow money. When investors invest their money through Lending Club, this money is passed onto borrowers, and when borrowers pay their loans back, the capital plus the interest passes on back to the investors. It is a win for everybody as they can get typically lower loan rates and higher investor returns.
#
# If you want to learn more about Lending Club follow this [link](https://www.lendingclub.com/)
#
# The Lending Club dataset contains complete loan data for all loans issued through the 2007-2015, including the current loan status (Current, Late, Fully Paid, etc.) and latest payment information. Features (aka variables) include credit scores, number of finance inquiries, address including zip codes and state, and collections among others. Collections indicates whether the customer has missed one or more payments and the team is trying to recover their money. The file is a matrix of about 890 thousand observations and 75 variables. More detail on this dataset can be found in [Kaggle's website](https://www.kaggle.com/wendykan/lending-club-loan-data)
#
# ====================================================================================================
#
# To download the Titanic data, go ahead to the [Kaggle website](https://www.kaggle.com/c/titanic/data)
#
# Click on the link 'train.csv', and then click the 'download' blue button towards the right of the screen, to download the dataset. Save it in a folder of your choice.
#
# For the Lending Club dataset. go to this website:
# Go ahead to this [website](https://www.kaggle.com/wendykan/lending-club-loan-data)
#
# Scroll down to the bottom of the page, and click on the link 'loan.csv', and then click the 'download' blue button towards the right of the screen, to download the dataset. Unzip it, and save it to a directory of your choice.
#
# **Note that you need to be logged in to Kaggle in order to download the datasets**.
#
# If you save it in the same directory from which you are running this notebook, and you rename the file to 'titanic.csv' then you can load it the same way I will load it below.
#
# ====================================================================================================
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# to display the total number columns present in the dataset
pd.set_option('display.max_columns', None)
# +
# let's load the titanic dataset
data = pd.read_csv('titanic.csv')
data.head()
# -
# In python, the missing values are stored as NaN, see for example the first row for the variable Cabin.
# +
# you can determine the total number of missing values using
# the isnull method plus the sum method on the dataframe
data.isnull().sum()
# +
# alternatively, you can call the mean method after isnull
# to visualise the percentage of the dataset that
# contains missing values for each variable
data.isnull().mean()
# -
# We can see that there are missing data in the variables Age, Cabin (in which the passenger was travelling) and Embarked, which is the port from which the passenger got into the Titanic.
# ### Missing data Not At Random (MNAR): Systematic missing values
#
# In this dataset, both the missing values of the variables Cabin and Age, were introduced systematically. For many of the people who did not survive, the **age** they had or the **cabin** they were staying in, could not be established. The people who survived could be asked for that information.
#
# Can we infer this by looking at the data?
#
# In a situation like this, we could expect a greater number of missing values for people who did not survive.
#
# Let's have a look.
# +
# we create a dummy variable that indicates whether the value
# of the variable cabin is missing
data['cabin_null'] = np.where(data.Cabin.isnull(), 1, 0)
# find percentage of null values
data.cabin_null.mean()
# -
# As expected, this value coincides with the one observed above when we called the .isnull().mean() method on the dataset.
# +
# and then we evaluate the mean of the missing values in
# cabin for the people who survived vs the non-survivors.
# group data by Survived vs Non-Survived
# and find nulls for cabin
data.groupby(['Survived'])['cabin_null'].mean()
# -
# We observe that the percentage of missing values is higher for people who did not survive (0.87), respect to people that survived (0.60).
# This finding is aligned with our hypothesis that the data is missing because after the people died, the information could not be retrieved.
#
# Having said this, to truly underpin whether the data is missing not at random, we would need to get extremely familiar with the way data was collected. Analysing datasets, can only point us in the right direction or help us build assumptions.
# +
# we repeat the exercise for the variable age:
# First we create a dummy variable that indicates
# whether the value of the variable Age is missing
data['age_null'] = np.where(data.Age.isnull(), 1, 0)
# and then look at the mean in the different survival groups:
# there are more NaN for the people who did not survive
data.groupby(['Survived'])['age_null'].mean()
# -
# Again, we observe an increase in missing data for the people who did not survive the tragedy. The analysis therefore suggests:
#
# **There is a systematic loss of data: people who did not survive tend to have more information missing. Presumably, the method chosen to gather the information, contributes to the generation of these missing data.**
# ### Missing data Completely At Random (MCAR)
#
# In the titanic dataset, there were also missing values for the variable Embarked, let's have a look.
# +
# slice the dataframe to show only those observations
# with missing value for Embarked
data[data.Embarked.isnull()]
# -
# These 2 women were travelling together, <NAME> was the maid of Mrs Stone.
#
# A priori, there does not seem to be an indication that the missing information in the variable Embarked is depending on any other variable, and the fact that these women survived, means that they could have been asked for this information.
#
# Very likely this missingness was generated at the time of building the dataset and therefore we could assume that it is completely random. We can assume that the probability of data being missing for these 2 women is the same as the probability for this variable to be missing for any other person. Of course this will be hard, if possible at all, to prove.
# ### Missing data At Random (MAR)
#
# For this example, I will use the Lending Club loan book. I will look specifically at the variables employer name (emp_title) and years in employment (emp_length), declared by the borrowers at the time of applying for a loan. The former refers to the name of the company for which the borrower works, the second one to how many years the borrower has worked for named company.
#
# Here I will show an example, in which a data point missing in one variable (emp_title) depends on the value entered on the other variable (emp_lenght).
# +
# let's load the columns of interest from the Lending Club loan book dataset
data=pd.read_csv('loan.csv', usecols=['emp_title','emp_length'])
data.head()
# -
# let's check the amount of missing data
data.isnull().mean()
# Around 6% of the observations contain missing data for employment title.
# +
# let's peek at the different employer names
print('Number of different employer names: {}'.format(len(data.emp_title.unique())))
data.emp_title.unique()[0:20]
# -
# let's inspect the variable emp_length
data.emp_length.unique()
# +
# let's look at the percentage of borrowers within
# each label / category of the emp_length variable
data.emp_length.value_counts() / len(data)
# -
# The label 'n/a', which in virtue could be similar to NaN, represents also around 5% of the dataset. So there could be a relationship between missing values in emp_title and 'n/a' in emp_length. Let's have a look.
# +
# the variable emp_length has many categories. I will summarise it
# into 3 for simplicity:'0-10 years' or '10+ years' or 'n/a'
# let's build a dictionary to re-map emp_length to just 3 categories:
length_dict = {k:'0-10 years' for k in data.emp_length.unique()}
length_dict['10+ years']='10+ years'
length_dict['n/a']='n/a'
# let's look at the dictionary
length_dict
# +
# let's re-map the emp_length
data['emp_length_redefined'] = data.emp_length.map(length_dict)
data.emp_length_redefined.unique()
# +
# let's calculate the proportion of working years
# with same employer for those who miss data on employer name
# number of borrowers for whom employer name is missing
value = len(data[data.emp_title.isnull()])
# % of borrowers for whom employer name is missing
# within each category of employment length
data[data.emp_title.isnull()].groupby(['emp_length_redefined'])['emp_length'].count().sort_values() / value
# -
# The majority of the missing values in the job title (emp_title) supplied by the borrower coincides with the label n/a of employment length. This supports the idea that the 2 variables are related.
# +
# let's do the same for those bororwers who reported
# the employer name
# number of borrowers for whom employer name is present
value = len(data.dropna(subset=['emp_title']))
# % of borrowers within each category
data.dropna(subset=['emp_title']).groupby(['emp_length_redefined'])['emp_length'].count().sort_values() / value
# -
# The number of borrowers who have reported an employer name and indicate n/a as employment length are minimal. Further supporting the idea that the missing values in employment length and employment length are related.
#
# 'n/a' in 'employment length' could be supplied by people who are retired, or students, or self-employed. In all of those cases there would not be a number of years at employment to provide, therefore the customer would enter 'n/a' and leave empty the form at the side of 'employer_name'.
#
# In a scenario like this, a missing value in the variable emp_title depends on or is related to the 'n/a' label in the variable emp_length. And, this missing value nature is, in principle, independent of the variable we want to predict (in this case whether the borrower will repay their loan). How this will affect the predictions is unknown.
# **That is all for this demonstration. I hope you enjoyed the notebook, and see you in the next one.**
| Feature Engineering/Feature-Engineering-master/.ipynb_checkpoints/03.1_Missing_values-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Decision trees and random forests
# Decision trees build classification or regression models in the form of a tree structure. Creating a binary decision tree is a process of dividing up the input space. A greedy and recursive approach is used to recursively create binary splits in the input space. At each step, all the values are lined up and different split points are tried and tested using a cost function. The split with the best cost (lowest cost because we minimize cost) is selected. All input variables and all possible split points are evaluated and chosen in a greedy manner as the optimal split point is chosen in each step without checking for whether or not the split will lead to the lowest possible cost several levels down.
#
# In order to make a prediction for a given observation, we typically use the mean (regression) or the mode (classification) of the training observations in the region to which the observation belongs.
# ### A. Classification using decision trees
# 1. Import the make_moons dataset from sklearn using $100$ samples, noise $= 0.35$ and a fixed random state = $42$.
# 2. Fit and visualize a decision tree classifier. Plot the decision boundaries by calling the helper function plot_decision_boundary provided to you.
import numpy as np
import matplotlib.pyplot as plt
from matplotlib.colors import ListedColormap
from sklearn.datasets import make_moons
from sklearn.tree import DecisionTreeClassifier, DecisionTreeRegressor
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import train_test_split, GridSearchCV
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import classification_report
import numpy as np
import pandas as pd
import seaborn as sns
# +
def plot_decision_boundary(tree, X, y, axes=[0, 2.5, 0, 1.2], legend=False, plot_training=True):
x1s = np.linspace(axes[0], axes[1], 100)
x2s = np.linspace(axes[2], axes[3], 100)
x1, x2 = np.meshgrid(x1s, x2s)
X_new = np.c_[x1.ravel(), x2.ravel()]
y_pred = tree.predict(X_new).reshape(x1.shape)
custom_cmap = ListedColormap(['#fafab0','#9898ff','#a0faa0'])
plt.contourf(x1, x2, y_pred, alpha=0.3, cmap=custom_cmap)
custom_cmap2 = ListedColormap(['#7d7d58','#4c4c7f','#507d50'])
plt.contour(x1, x2, y_pred, cmap=custom_cmap2, alpha=0.8)
if plot_training:
plt.plot(X[:, 0][y==0], X[:, 1][y==0], "yo")
plt.plot(X[:, 0][y==1], X[:, 1][y==1], "bs")
plt.plot(X[:, 0][y==2], X[:, 1][y==2], "g^")
plt.axis(axes)
plt.xlabel(r"$x_1$", fontsize=18)
plt.ylabel(r"$x_2$", fontsize=18, rotation=0)
if legend:
plt.legend(["0","1"],loc="upper right", fontsize=14)
# +
##PART A
#Loading data
Xm,ym = make_moons(n_samples=100,random_state=42,noise = 0.35)
#Initializing Decision Tree Model
DcT = DecisionTreeClassifier()
#Training
classifier = DcT.fit(Xm,ym)
#Plotting/Visualization
plt.figure(figsize=(8,5))
plt.title("Decision Tree fit on Moons Dataset with default")
plot_decision_boundary(classifier,Xm,ym,[min(Xm[:,0])-0.015,max(Xm[:,0])+0.015,min(Xm[:,1])-0.015,max(Xm[:,1])+0.015])
#Calculating classification accuracy
acc = classifier.score(Xm,ym)
print("Accuracy = %.f"%(acc*100)+"%")
# -
# #### Q. Comment on the overfitting nature of the Decision trees
# Overfitting is a commonly ocurring phenomenon when it comes to decision trees. Overfitting is a result of high variance in the model and can lead to poor results on unseen data. A naive implementation of decision trees can result in very a complex tree structure with large number of nodes causing overfitting on training data. Since we aim to minimise the impurity of the partioned data at each step of the algorithm, a higher number of partitions are made to achieve a greater depth and hence low or zero impurity. Even if the data is noisy or having outliers, decision trees tend to fit complex boundaries to accomodate noise and outlier points. For the same reason, Decision Trees are very less robust to variation in data. For example, in the decision tree model (with maximum depth set to a default value of 6) trained on the Make Moons dataset, although the accuracy is 100%, a visual inspection of the scatter plot with the generated decision boundaries suggests that the algorithm has partitioned tiny regions in the data space to accommodate just a few yellow and blue points, which is undesirable in general.
# ### B. Overfitting in decision tree classifiers
# 1. List and explain in a few lines, three of the regularization hyperparameters used while building a decision tree and why they might help reduce overfitting of a decision tree model.
# 2. Now fit a decision tree classifier on the same dataset using the min_samples_leaf hyperparameter set to $4$. Plot the decision boundaries and compare with the model fit without any hyperparameters. Which model do you think will generalize better to the test dataset?
# 3. Fit and visualize a decision tree classifier after setting the maximum depth of nodes as $5$. Plot the decision boundaries by calling the helper function plot_decision_boundary provided to you. and compare with the model fit without any hyperparameters. Which model do you think will generalize better to the test dataset?
# **Question B.1**
# The hyperparameters of decision tree models that can be used for regularization are:
# - **Maximum depth of the tree (```max_depth```):** Upon decreasing the maximum depth of the tree to an appropriate value, we can enforce the decision tree algorithm to prevent it from building large and complex trees with a higher depth, thereby preventing overfitting.
# - **Minimum number of samples in the leaf nodes (```min_sample_leaf```):** When we allow smaller number of samples to be allowed inside a leaf, the decision tree algorithm tends to reach there by increasing the depth of the tree, again resulting in overfitting. Setting a higher threshold for the number of samples present to be considered as a leaf node can serve as a good early stopping condition and prevents overfitting.
# - **Pruning - Complexity Parameter (```ccp_alpha```):** Another way to prevent overfitting is pruning of the decision tree. It removes sections of the tree that are uncritical and redundant to classify instances. Pruning is parametrized by the cost complexity parameter $\alpha$, which when assigned a higher value allows higher number of sections of the tree to be pruned.
DcT = DecisionTreeClassifier(random_state=0,min_samples_leaf=4)
#Training
classifier = DcT.fit(Xm,ym)
#Plotting/Visualization
plt.figure(figsize=(8,5))
plt.title("Decision Tree fit on Moons Dataset with min_samples_leaf 4")
plot_decision_boundary(classifier,Xm,ym,[min(Xm[:,0])-0.015,max(Xm[:,0])+0.015,min(Xm[:,1])-0.015,max(Xm[:,1])+0.015])
#Calculating classification accuracy
acc = classifier.score(Xm,ym)
print("Accuracy = %.f"%(acc*100)+"%")
# It can be observed that this model which allows a minimum number of 4 samples in the leaf nodes has attempted to prevent overfitting by making the decision boundaries less complex. However, again through visual inspection, we can say that that this model will generalize marginally better than that trained with default hyperparameters. An accuracy of 95% is achieved on the training dataset.
DcT = DecisionTreeClassifier(random_state=0,max_depth=5)
#Training
classifier = DcT.fit(Xm,ym)
#Plotting/Visualization
plt.figure(figsize=(8,5))
plt.title("Decision Tree fit on Moons Dataset with max depth 5")
plot_decision_boundary(classifier,Xm,ym,[min(Xm[:,0])-0.015,max(Xm[:,0])+0.015,min(Xm[:,1])-0.015,max(Xm[:,1])+0.015])
#Calculating classification accuracy
acc = classifier.score(Xm,ym)
print("Accuracy = %.f"%(acc*100)+"%")
# It can be observed that this model which allows a maximum depth as 5 has attempted to prevent overfitting by making the decision boundaries less complex. Through visual inspection, this model is quite similar to the model trained with maximum depth as 4, and might achieve a slight improvement in terms of generalization on unseen data, compared to the unregularized model. A training accuracy of 96% is achieved.
# #### Q. Comment on the boundary surface of the decision tree.
# Hint: Observe the orthogonality of the boundary
# It can be observed that the decision boundaries generated by the decision tree model are non-linear. However, a single partition (at any intermediate iteration) is a linear boundary that divides the vector space into two. A hierarchical combination of such linear splits generates a non-linear boundary structure. Furthermore, since we split along only one axial direction (corresponding to a single feature) at a time while partitioning the space, orthogonality can be observed in the decision boundary lines.
# ### C. Linear classifiers vs decision tree classifiers
#Generate 100 points [x1, x2] in the 2D plane using np.random.rand() for -0.5<x1<0.5 and -0.5<x2<0.5
np.random.seed(2030)
x = np.random.rand(100,2) - 0.5
y = np.zeros(100)
#Assign a label of 1 to all points having x1>0.0 and 0 otherwise
y[x[:,0]>0] = 1
#Rotate the dataset by 45 degrees in the [x1,x2] plane
rot = [[0.414,-0.414],[0.414,0.414]]
x_rot = np.matmul(rot,x.T)
x_rot = x_rot.T
# 1. Fit a linear Logistic Regression model and Decision Tree on the x_rot
# 2. Fit a linear Logistic Regression model and Decision tree on the make_moons dataset generated in A(1)
# #### Part C.1
log_reg = LogisticRegression().fit(x_rot,y)
plt.figure(figsize=(8,5))
plt.title("Logistic Regression fit on Rotated Dataset")
plot_decision_boundary(log_reg,x_rot,y,[min(x_rot[:,0])-0.015,max(x_rot[:,0])+0.015,min(x_rot[:,1])-0.015,max(x_rot[:,1])+0.015])
acc = log_reg.score(x_rot,y)
print("Accuracy = %.f"%(acc*100)+"%")
DcT_rot = DecisionTreeClassifier(random_state=0).fit(x_rot,y)
plt.figure(figsize=(8,5))
plt.title("Decision Tree fit on Rotated Dataset")
plot_decision_boundary(DcT_rot,x_rot,y,[min(x_rot[:,0])-0.015,max(x_rot[:,0])+0.015,min(x_rot[:,1])-0.015,max(x_rot[:,1])+0.015])
acc = DcT_rot.score(x_rot,y)
print("Accuracy = %.f"%(acc*100)+"%")
# #### Part C.2
# +
#Decision Tree on Moon Dataset
DcT = DecisionTreeClassifier()
#Training
classifier = DcT.fit(Xm,ym)
#Plotting/Visualization
plt.figure(figsize=(8,5))
plt.title("Decision Tree fit on Moons Dataset with default")
plot_decision_boundary(classifier,Xm,ym,[min(Xm[:,0])-0.015,max(Xm[:,0])+0.015,min(Xm[:,1])-0.015,max(Xm[:,1])+0.015])
#Calculating classification accuracy
acc = classifier.score(Xm,ym)
print("Accuracy = %.f"%(acc*100)+"%")
# -
#Logistic Regeression on Moon Dataset
log_reg = LogisticRegression().fit(Xm,ym)
plt.figure(figsize=(8,5))
plt.title("Logistic Regression on Moons Dataset")
plot_decision_boundary(log_reg,Xm,ym,[min(Xm[:,0])-0.015,max(Xm[:,0])+0.015,min(Xm[:,1])-0.015,max(Xm[:,1])+0.015])
acc = log_reg.score(Xm,ym)
print("Accuracy = %.f"%(acc*100)+"%")
# #### Q. Compare the performance of Decision Trees and logistic regression on the two datasets
# It is noted that the decision tree model achieves 100% accuracy on classifying the Rotated data, while the accuracy of Logistic Regression is just 93%. Likeweise, the decision tree model achieves 100% accuracy on classifying the moons data, while the accuracy of Logistic Regression is just 86%. These observations suggest that the decision tree performs very well on the data being trained, using a complex tree structure resulting in nonlinear decision boundary. On the other hand, Logistic Regression generates a linear boundary that is unable to classify all data points accurately. Logistic Regression also uses the probabilily generated by the sigmoid function to classify the points, resulting in a probablisitic assignment of classes to the data points based on a threshold, as opposed to decision trees.
# ### D. Regression using decision trees
# 1. Train a regression tree on the noisy sine dataset given below with three different max_depth values : $2, 3$ and $4$.
# 2. Plot the divisions of the input space and the predicted value in each division of the input space using the helper function plot_regression_predictions provided to you below for each regression tree model in (1).
# +
def plot_regression_predictions(tree_reg, X, y, axes=[0, 1, -0.2, 1], ylabel="$y$"):
x1 = np.linspace(axes[0], axes[1], 500).reshape(-1, 1)
y_pred = tree_reg.predict(x1)
plt.axis(axes)
plt.xlabel("$x_1$", fontsize=18)
if ylabel:
plt.ylabel(ylabel, fontsize=18, rotation=0)
plt.plot(X, y, "b.")
plt.plot(x1, y_pred, "r.-", linewidth=2, label=r"$\hat{y}$")
#Sine + noise training set
np.random.seed(10)
X = np.random.rand(150, 1)
y = np.sin(2*np.pi*X) + np.random.randn(150, 1) / 10
# -
# #### Part D.1
#Decision Tree Regression on Sinusoidal data - maximum depth 2
dec_reg = DecisionTreeRegressor(max_depth=2)
dec_reg.fit(X,y)
#plt.title("Decision Tree Regression on Sinusoidal Data with max depth 2")
plot_regression_predictions(dec_reg,X,y,[min(X)-0.015,max(X)+0.015,min(y)-0.015,max(y)+0.015])
acc = dec_reg.score(X,y)
print("Accuracy = %.2f"%(acc*100)+"%")
plt.title("Decision Tree - Max Depth 2")
#Decision Tree Regression on Sinusoidal data - maximum depth 3
dec_reg = DecisionTreeRegressor(max_depth=3)
dec_reg.fit(X,y)
plot_regression_predictions(dec_reg,X,y,[min(X)-0.015,max(X)+0.015,min(y)-0.015,max(y)+0.015])
acc = dec_reg.score(X,y)
print("Accuracy = %.2f"%(acc*100)+"%")
plt.title("Decision Tree - Max Depth 3")
#Decision Tree Regression on Sinusoidal data - maximum depth 4
dec_reg = DecisionTreeRegressor(max_depth=4)
dec_reg.fit(X,y)
plot_regression_predictions(dec_reg,X,y,[min(X)-0.015,max(X)+0.015,min(y)-0.015,max(y)+0.015])
acc = dec_reg.score(X,y)
print("Accuracy = %.2f"%(acc*100)+"%")
plt.title("Decision Tree - Max Depth 4")
# As observed, increasing the regularizing parameter maximum depth leads to higher training accuracy, and this might lead to overfitting.
# ### E. Overfitting in regression trees
# 1. Fit another regression tree on the dataset used above, with no restrictions on the regularization hyperparameters of the model. Plot the predicted value in all input space divisions.
# 2. What do you observe? Do regression trees too have a tendency to overfit if no regularization is applied?
#Decision Tree Regression on Sinusoidal data - No regularization
dec_reg = DecisionTreeRegressor()
dec_reg.fit(X,y)
plot_regression_predictions(dec_reg,X,y,[min(X)-0.015,max(X)+0.015,min(y)-0.015,max(y)+0.015])
acc = dec_reg.score(X,y)
print("Accuracy = %.2f"%(acc*100)+"%")
plt.title("Decision Tree - No Regularization")
# It is clearly evident that training the decision tree model on the data with no regularization overfits the data using highly complex decision trees, achieving 100% accuracy. Since the maximum depth was not set, this model kept increasing the depth of the tree until the minimum number of samples in the leaves is reached. On the other hand, we impose early stopping by specicifying the maximum depth parameter, and this results less complex models. As the maximum depth increases, the training accuracy increases.
# ### F. Fine-tuning a decision tree
# 1. Generate a moons dataset using make_moons (n_samples = $10000$, noise=$0.5$). Random seed = 42
# 2. Split it into a training set and a test set.
# 3. Use grid search with cross-validation (with the help of GridSearchCV) to find good hyperparameter values for max_leaf_nodes and min_samples_split. Try values ranging from $2$ to $35$ for max_leaf_nodes and $2$ to $10$ for min_samples_split. Report the optimal hyperparameter values.
# 4. Train the decision tree classifier on the full training set using these optimal hyperparameters, and report your model's performance on the test set.
#PART F.1 - Loading 10000 samples of Moons data with noise = 0.5
X_moon,y_moon = make_moons(n_samples=10000,random_state=42,noise=0.5)
#PART F.2 - Train Test Splitting with Test size = 20%
[X_train,X_test,y_train,y_test] = train_test_split(X_moon,y_moon,test_size = 0.2,random_state = 42)
# +
#PART F.3 - Using GridSearchCV for finding the best set of hyperparameters for max_leaf_nodes and min_samples_split of
#the Decision Tree Classifier
parameters = {"max_leaf_nodes":list(range(2,36)),"min_samples_split":list(range(2,11))}
DcT_search = DecisionTreeClassifier()
classifier = GridSearchCV(DcT_search,parameters)
#PART F.4 Training the classifier on the training set and test set and reporting accuracies
classifier.fit(X_train,y_train)
best_classifier = classifier.best_estimator_
print("Reporting Best Model")
print("Maximum Leaf Nodes = %d"%best_classifier.max_leaf_nodes)
print("Minimum Sample Nodes = %d"%best_classifier.min_samples_split)
print("Training Accuracy of best model = %.2f"%(best_classifier.score(X_train,y_train)*100)+"%")
plt.figure(figsize=(8,5))
plt.title("Decision Tree fit on Moons Training Data (8k samples) with best model")
plot_decision_boundary(best_classifier,X_train,y_train,[min(X_train[:,0])-0.015,max(X_train[:,0])+0.015,min(X_train[:,1])-0.015,max(X_train[:,1])+0.015])
# -
#Performance on the test data
print("Test Accuracy of best model = %.2f"%(best_classifier.score(X_test,y_test)*100)+"%")
plt.figure(figsize=(8,5))
plt.title("Decision Tree fit on Moons Test Data (2k samples) with best model")
plot_decision_boundary(best_classifier,X_test,y_test,[min(X_test[:,0])-0.015,max(X_test[:,0])+0.015,min(X_test[:,1])-0.015,max(X_test[:,1])+0.015])
# **As observed, the best model performs equally well on both training and test set data.**
# ### G. Training the Random Forest model
# 1. Load the loans dataset and use one-hot encoding for the column 'purpose'
# 2. Split the data into test and train sets with ratio 70:30 with random seed = 101
# 3. Create an instance of the RandomForestClassifier class and fit it to our training data from the previous step.
# 4. Predict the class of not.fully.paid for the X_test data
# 5. Now create a classification report from the results. Do you get anything strange or some sort of warning?
#PART G.1 - Loading the Loans Dataset and performing OHE
loan_data = pd.read_csv("loan_data.csv")
not_paid = loan_data["not.fully.paid"]
del loan_data["not.fully.paid"]
loan_data = pd.get_dummies(loan_data)
#PART G.2 - Train-Test Split 70:30
[X_train,X_test,y_train,y_test] = train_test_split(loan_data,not_paid,test_size = 0.3,random_state = 101)
#PART G.3 - Training with Random Forest Classifier
RF = RandomForestClassifier()
RF.fit(X_train,y_train)
training_acc = RF.score(X_train,y_train)
print("Training Accuracy of Moon Data = %.2f"%(training_acc*100)+"%")
#PART G.4 - Prediction on test dataset and reporting accuracies
test_acc = RF.score(X_test,y_test)
print("Test Accuracy of Moon Data = %.2f"%(test_acc*100)+"%")
# As noted, even though the Random Forest Classifier achieves a highest accuracy score of 100% on the training data, the accuracy on the test data is only 84.59%.
#PART G.5 - Classification Report
#Classification Report of Training Data
print("Classification Report of Training Data")
print(classification_report(RF.predict(X_train),y_train))
#Classification Report of Test Data
print("Classification Report of Test Data")
print(classification_report(RF.predict(X_test),y_test))
# The prevalence of class imbalance is clearly evident upon observing the classification report of the results obtained from the test data. This is due to very low f1-score and support for the class 1, suggesting that the number of data points in the test data having class label 1 is very less compared to 0-class. A tiny value of precision of class 1 indicates higher number of false positives with respect to that class. On the other hand, class 0 has high values of precision, recall, and f1-score. As the fraction of class-0 samples are really high, there are high chances that the data is classified in that particular class, resulting in higher number of true positives.
print("Percentage of class 1 in the test dataset = %.2f"%(np.sum(y_test==1)*100/len(y_test)))
# This can be confirmed by checking the actual percentage of data points having class label 1, and this turns out to be only 15.41%.
| Assignment 8/G Prashant (BS17B011) - Assignment 8.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: "Python 3.7 (Intel\xAE oneAPI)"
# language: python
# name: c009-intel_distribution_of_python_3_oneapi-beta05-python
# ---
# # Introduction to Numba-dppy
# ## Sections
#
# - [oneAPI Programming Model Overview](#oneAPI-Programming-Model-Overview)
# - [Programming Challenges for Multiple architectures](#Programming-Challenges-for-Multiple-architectures)
# - [Introducing oneAPI](#Introducing-oneAPI)
# - [Introduction to Intel® Distribution for Python*](#Introduction-to-Intel®-Distribution-for-Python*)
# - [Introduction to Numba - Data Parallel Python (numba-dppy)](#Introduction-to-Numba---Data-Parallel-Python-(numba-dppy))
# - [Introduction to Numba](#Numba*)
# - [Parallelism in Python Using Numba](#Parallelism-in-Python-Using-Numba)
# - _Code:_ [Python code running in serial](#Python-code-running-in-serial)
# - _Code:_ [Numba JIT Compilation for CPU](#Numba-JIT-Compilation-for-CPU)
# - [Introduction to numba-dppy](#Introduction-to-numba-dppy)
# - _Code:_ [numba-dppy Example uisng automatic offload](#Automatic-Offload)
# - _Code:_ [Writing Explicit Kernels in numba-dppy](#Writing-Explicit-Kernels-in-numba-dppy)
# ## Learning Objectives
#
# * Explain how the __oneAPI__ programming model can solve the challenges of programming in a heterogeneous world
# * Introduce Intel® Distribution of Python and numba-dppy
# * Utilize __Numba__ and __Numba-dppy__ to write paralle code on CPU and GPU
# * Write explicit kernels using numba-dppy @kernel decorator
# ## oneAPI Programming Model Overview
# The __oneAPI__ programming model provides a comprehensive and unified portfolio of developer tools that can
# be used across hardware targets, including a range of performance libraries spanning several workload
# domains. The libraries include functions custom-coded for each target architecture so the same
# function call delivers optimized performance across supported architectures. __DPC++__ is based on
# industry standards and open specifications to encourage ecosystem collaboration and innovation.
#
# ### oneAPI Distribution
# Intel® oneAPI toolkits are available via multiple distribution channels:
# * Local product installation: install the oneAPI toolkits from the __Intel® Developer Zone__.
# * Install from containers or repositories: install the oneAPI toolkits from one of several supported
# containers or repositories.
# * Pre-installed in the __Intel® DevCloud__: a free development sandbox for access to the latest Intel® SVMS hardware and select oneAPI toolkits.
# ## Programming Challenges for Multiple architectures
# Currently in the data centric space there is growth in specialized workloads. Each kind of data centric hardware typically needs to be programmed using different languages and libraries as there is no common programming language or APIs, this requires maintaining separate code bases. Developers have to learn a whole set of different tools as there is inconsistent tool support across platforms. Developing software for each hardware platform requires a separate investment, with little ability to reuse that work to target a different architecture. You will also have to consider the requirement of the diverse set of data-centric hardware.
#
# <img src="Assets/oneapi1.png">
#
# ## Introducing oneAPI
# __oneAPI__ is a solution to deliver unified programming model to __simplify development__ across diverse architectures. It includes a unified and simplified language and libraries for expressing __parallelism__ and delivers uncompromised native high-level language performance across a range of hardware including __CPUs, GPUs, FPGAs__. oneAPI initiative is based on __industry standards and open specifications__ and is interoperable with existing HPC programming models.
#
# <img src="Assets/oneapi2.png">
#
# ## Introduction to Intel® Distribution for Python*
#
# IDP is a complete Python distribution that includes the necessary Python packages to develop high-performing code targeting Intel® XPUs using Python. The distribution includes the following:
#
# * Optimized Python numerical packages NumPy, Scikit-learn, MKL-FFT that use Intel® oneAPI Math Kernel Library (oneMKL) and Intel® oneAPI Data Analytics Library (oneDAL) to offer near-native performance.
#
# * Customized version of the Numba* JIT compiler that allows generating fast code for Intel® XPUs.
#
# * Data Parallel Python (DPPY) that is a set of packages enabling SYCL-based XPU programming.
# 1. Data Parallel Control (dpctl): A package for controlling execution on SYCL devices and for SYCL USM data management.
# 1. Data Parallel Numeric Python (dpnp): An implementation of the NumPy API using SYCL and oneMKL.
# 1. Numba-dppy: A standalone extension to Numba adding SYCL kernel programming to Numba*.
#
# * Faster machine learning with XGBoost*, scikit-learn, and advanced ML usages, including multiple devices, with daal4py.
#
# * Scikit-ipp for image warping, image filtering, and morphological operations. Includes support for transform function multithreading and partial multithreading for filters using OpenMP.
# ## Numba*
#
# Numba is an open-source, NumPy-aware optimizing compiler for Python developed by Anaconda, Inc in collaboration with an open-source community. It uses the LLVM compiler to generate machine code from Python bytecode.
# Numba can compile a large subset of numerically focused Python, including many NumPy functions. Additionally, Numba has support for automatic parallelization of loops, generation of GPU-accelerated code, and creation of ufuncs and C callbacks. For more information about Numba, see the Numba homepage: http://numba.pydata.org
#
# ### Parallelising Python Programs Using Numba*
#
# Numba has a built in auto-parallelizer that was contributed by Intel. The auto-parallelizer can be enabled by setting the `parallel=True` option in the `@numba.jit` decorator. The auto-parallelizer analyzes data-parallel code regions in the compiled function and schedules them for parallel execution using either OpenMP or TBB.
#
# There are two types of operations that Numba can automatically parallelize: Implicitly data-parallel regions such as NumPy array expressions, NumPy ufuncs, NumPy reduction fucntions. Explicitly data-parallel loops that are specified using the `numba.prange` expression. Please refer https://numba.pydata.org/numba-doc/latest/user/parallel.html for further details on Numba's automatic paralleizer.
# ### Automatic Offload of NumPy Expressions
# A key distinction between Numba-dppy and other GPU backends in Numba is the ability to automatically offload specific data-parallel sections of a Numba JIT function.
#
# ## Parallelism in Python Using Numba
#
# Python has become a pervasive and useful tool in advancing scientific research and computation. Python has a very rich ecosystem of open-source packages for mathematics, science, and engineering, anchored on the performant numerical computation on arrays and matrices, data analysis and visualization capabilities, and an interactive development environment that enables rapid and collaborative iteration of ideas. Python is used to discover new objects in space, calculate thermodynamics, conduct genomic analysis of cancer, estimate the likelihood of earthquakes, simulate musculoskeletal systems, visualize asteroid chemistries, and much more.
#
# Intel’s accelerated Python packages enable scientists to take advantage of the productivity of Python, while taking advantage of the ever-increasing performance of modern hardware. Intel optimized implementations of NumPy and SciPy leverage the Intel Math Kernel Library to achieve highly efficient multithreading, vectorization, and memory management.
# Python as a programming language has enjoyed nearly a decade of use in both industry and academia. This high-productivity language has been one of the most popular abstractions to scientific computing and machine learning, yet the base Python language remains single-threaded. Just how is productivity in these fields being maintained with a single-threaded language?
#
# The Python language’s design, by <NAME>, was meant to trade off type flexibility and predictable, thread-safe behavior against the complexity of having to manage static types and threading primitives. This, in turn, meant having to enforce a global interpreter lock (GIL) to limit execution to a single thread at a time to preserve this design mentality.
#
# Over the last decade, many concurrency implementations have been made for Python―but few in the region of parallelism. Does this mean the language isn’t performant? Let’s explore further.
# The base language’s fundamental constructs for loops and other asynchronous or concurrent calls all abide by the single-threaded GIL, so even list comprehensions such as [x*x for x in range(0,10)] will always be single-threaded. The threading library’s existence in the base language is also a bit misleading, since it provides the behavior of a threading implementation but still operates under the GIL. Many of the features of Python’s concurrent futures to almost parallel tasks also operate under the GIL. Why does such an expressive productivity language restrict the language to these rules?
#
# The reason is the level of abstraction the language design adopted. It ships with many tools to wrap C code, from ctypes to cffi. It prefers multiprocessing over multithreading in the base language, as evidenced by the multiprocessing package in the native Python library. These two design ideas are evident in some of the popular packages, like NumPy and SciPy, which use C code under the Python API to dispatch to a mathematical runtime library such as Intel Math Kernel Library (Intel MKL) or OpenBLAS. The community has adopted the paradigm to dispatch to higher-speed C-based libraries, and has become the preferred method to implement parallelism in Python.
# In the combination of these accepted methods and language limitations are options to escape them and apply parallelism in Python through unique parallelism frameworks:
# * Numba allows for JIT-based compilation of Python code, which can also run LLVM-based Python-compatible code.
# * Cython gives Python-like syntax with compiled modules that can target hardware vectorization as it compiles to a C module.
# * numexpr allows for symbolic evaluation to utilize compilers and advanced vectorization.
#
# These methods escape Python’s GIL in different ways while preserving the original intent of the language, and all three implement different models of parallelism.
# Let’s take the general example of one of the most common language constructs on which we’d want to apply parallelism—the for loop. Looking at the following loop, we can see that it provides a basic service, a vector addition:
# ### Python code running in serial
# The following code demonstrates usage of serial python code. Inspect code; there are no modifications necessary:
# 1. Inspect the following code cell and click Run (▶)to save the code to file.
# 2. Next, run (▶) the cell in the __Build and Run__ section following the code to compile and execute the code.
# +
# %%writefile lab/serial_python.py
# Copyright 2021 Intel Corporation
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from numba import njit
import numpy as np
import dpctl
import timeit
def f1(a, b,c,N):
for i in range(N):
c[i] = a[i] + b[i]
N = 500000
a = np.ones(N, dtype=np.float32)
b = np.ones(N, dtype=np.float32)
c = np.zeros(N,dtype=np.float32)
t = timeit.Timer(lambda: f1(a,b,c,N))
print("Time to calculate the sum in Serial",t.timeit(200),"seconds")
print(c)
# -
# #### Build and Run
# Select the cell below and click run ▶ to compile and execute the code:
# ! chmod 755 q; chmod 755 run_serial_python.sh; if [ -x "$(command -v qsub)" ]; then ./q run_serial_python.sh; else ./run_serial_python.sh; fi
# _If the Jupyter cells are not responsive, or if they error out when you compile the code samples, please restart the Jupyter Kernel:
# "Kernel->Restart Kernel and Clear All Outputs" and compile the code samples again_.
# ### Numba JIT Compilation for CPU
# The previous example handles everything sequentially and doesn’t apply any parallelism to the code. Because of the way this code is written, it’s a good candidate for the Numba framework. Numba uses a decorator (with the @ symbol) to flag functions for just-in-time (JIT) compilation, which we’ll try to apply on this function.
# The following code demonstrates usage of simple DPPY code. Inspect code; there are no modifications necessary:
# 1. Inspect the following code cell and click Run (▶) to save the code to file.
# 2. Next, run (▶) the cell in the __Build and Run__ section following the code to compile and execute the code.
# +
# %%writefile lab/simple_njit_cpu.py
from numba import njit,prange
import numpy as np
import dpctl
import timeit
@njit(parallel=True)
def f1(a, b,c,N):
for i in prange(N):
c[i] = a[i] + b[i]
N = 500000
a = np.ones(N, dtype=np.float32)
b = np.ones(N, dtype=np.float32)
c = np.zeros(N,dtype=np.float32)
t = timeit.Timer(lambda: f1(a,b,c,N))
print("Time to calculate the sum in parallel",t.timeit(200),"seconds")
print(c)
# -
# ! chmod 755 q; chmod 755 run_njit_cpu.sh; if [ -x "$(command -v qsub)" ]; then ./q run_njit_cpu.sh; else ./run_njit_cpu.sh; fi
# _If the Jupyter cells are not responsive, or if they error out when you compile the code samples, please restart the Jupyter Kernel:
# "Kernel->Restart Kernel and Clear All Outputs" and compile the code samples again_.
#
# Including the simple decorator increased performance significantly. This works because the original Python code is written in primitives and datatypes that can be easily compiled and vectorized to a CPU.
#
# The lesson here is that achieving parallelism in Python depends on how the original code is written. Cleanliness of datatypes and the use of vectorizable data structures allow Numba to parallelize code with the insertion of a simple decorator. Being careful about the use of Python dictionaries pays dividends, because historically they don’t vectorize well. Generators and comprehensions suffer from the same problem. Refactoring such code to lists, sets, or arrays can facilitate vectorization.
# ## Introduction to Data Parallel Control (dpctl)
# Dpctl provides a lightweight Python wrapper over a subset of DPC++/SYCL’s API. The goal of dpctl is not (yet) to provide an abstraction for every SYCL function. Dpctl is intended to provide a common runtime to manage specific SYCL resources, such as devices and USM memory, for SYCL-based Python packages and extension modules.
#
# The main features presently provided by dpctl are:
#
# 1. Python wrapper classes for the main SYCL runtime classes mentioned in Section 4.6 of SYCL provisional 2020 spec (https://bit.ly/3asQx07): `platform`, `device`, `context`, `device_selector`, and `queue`.
# 1. A USM memory manager to create Python objects that use SYCL USM for data allocation.
#
#
# Dpctl is available as part of the oneAPI Intel Distribution of Python (IDP). Once oneAPI is installed, dpctl is ready to be used by setting up the IDP that is available inside oneAPI.
# ## Introduction to numba-dppy
#
# Numba-dppy is a standalone extension to the Numba JIT compiler that adds SYCL programming capabilities to Numba. Numba-dppy is packaged as part of the IDP that comes with oneAPI base toolkit, and you don’t need to install any specific Conda packages. The support for SYCL is via DPC++'s SYCL runtime and other SYCL compilers are not supported by Numba-dppy.
#
# Numba-dppy provides two ways to express SYCL parallelism:
#
# * __Automatic offload approach via `@numba.jit`__. The automatic approach extends Numba's existing auto-parallelizer to support generating SYCL kernels from data-parallel code regions. Using the automatic offload approach a programmer needs only minimal changes to their existing code and can try to offload an existing `@numba.jit` decorated function to a SYCL device by invoking the function from a `dpctl.device_context`. We will go into the details of the approach later in the tutorial.
#
# * __Explicit kernel written using the `@numba_dppy.kernel` decorator__. The explicit kernel approach is similar to NUmba's other GPU backends, `numba.cuda` and `numba.roc`. The `@numba_dppy.kernel` decorator is provided in the `numba-dppy` package. Several advanced SYCL features such as _indexing_, _synchronization_, _fences_, _atomcis_ are provided by the `@numba_dppy.kernel` decorator. Thus, using the decorator a relatively low-level SYCL kernel can be written directly in Python. The feature is intended for programmers who already have SYCL and GPU programming experience.
#
# ### Automatic Offload
#
# The automatic offload feature in numba-dppy is triggered when a `@numba.jit` function is invoked inside a `dpctl.device_context` scope.
# + [markdown] tags=[]
# ## numba-dppy - Explicit parallel for-loop
#
# Using Numba's explicit parallel for-loop and the usage of `numba.prange` you can offload the loop to the GPU using 'dpctl.device_context'
# The following code demonstrates usage of numba-dppy's explicit offload using dpctl.device_context. Inspect code; there are no modifications necessary:
# 1. Inspect the following code cell and click Run (▶)to save the code to file.
# 2. Next, run (▶) the cell in the __Build and Run__ section following the code to compile and execute the code.
# -
# _If the Jupyter cells are not responsive, or if they error out when you compile the code samples, please restart the Jupyter Kernel:
# "Kernel->Restart Kernel and Clear All Outputs" and compile the code samples again_.
# +
# %%writefile lab/simple_context.py
# Copyright 2020, 2021 Intel Corporation
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
The numba_dppy extension adds an automatic offload optimizer to
numba. The optimizer automatically detects data-parallel code
regions in a numba.jit function and then offloads the data-parallel
regions to a SYCL device. The optimizer is triggered when a numba.jit
function is invoked inside a dpctl ``device_context`` scope.
This example demonstrates the usage of numba_dppy's automatic offload
functionality. Note that numba_dppy should be installed in your
environment for the example to work.
"""
import numpy as np
from numba import njit, prange
import dpctl
@njit
def add_two_arrays(b, c):
a = np.empty_like(b)
for i in prange(len(b)):
a[i] = b[i] + c[i]
return a
def main():
N = 10
b = np.ones(N)
c = np.ones(N)
# Use the environment variable SYCL_DEVICE_FILTER to change the default device.
# See https://github.com/intel/llvm/blob/sycl/sycl/doc/EnvironmentVariables.md#sycl_device_filter.
device = dpctl.select_default_device()
print("Using device ...")
device.print_device_info()
with dpctl.device_context(device):
result = add_two_arrays(b, c)
print("Result :", result)
print("Done...")
if __name__ == "__main__":
main()
# + [markdown] tags=[]
# #### Build and Run
# Select the cell below and click run ▶ to compile and execute the code:
# -
# ! chmod 755 q; chmod 755 run_simple.sh; if [ -x "$(command -v qsub)" ]; then ./q run_simple.sh; else ./run_simple.sh; fi
# _If the Jupyter cells are not responsive, or if they error out when you compile the code samples, please restart the Jupyter Kernel:
# "Kernel->Restart Kernel and Clear All Outputs" and compile the code samples again_.
# ## numba-dppy Automatic offload using @njit
#
# The following example demonstrates the auto-offload feature of Numba. Numba automatically identifies all the NumPy function calls within a jit-decorated function that have parallel semantics and compiles them for parallel execution on the CPU or the GPU based on the device context specified using `dpctl.device_context`.
#
# ```
# import dpctl
# import numpy as np
# import numba
# from numba import njit, prange
#
# @numba.njit(parallel=True)
# def l2_distance_kernel(a, b):
# sub = a - b
# sq = np.square(sub)
# sum = np.sum(sq)
# d = np.sqrt(sum)
# return d
#
# def main():
# R = 64
# C = 1
#
# X = np.random.random((R,C))
# Y = np.random.random((R,C))
#
# device = dpctl.select_default_device()
# print("Using device ...")
# device.print_device_info()
#
# with dpctl.device_context(device):
# result = l2_distance_kernel(X, Y)
#
# print("Result :", result)
# print("Done...")
#
# if __name__ == "__main__":
# main()
# ```
#
# #### Controllable the fallback behavior during automatic offload
#
# By default, if a section of code cannot be offloaded to the GPU, it is automatically executed on the CPU and a warning is printed. This behavior is only applicable to JIT functions, auto-offloading of NumPy calls, array expressions and prange loops. To disable this functionality and force code running on GPU, set the environment variable NUMBA_DPPY_FALLBACK_OPTION to false (for example, export NUMBA_DPPY_FALLBACK_OPTION=0). In this case the code is not automatically offloaded to the CPU and errors occur if any.
#
# #### Diagnostic reporting for automatic offload
#
# Export NUMBA_DPPY_OFFLOAD_DIAGNOSTICS=1:
#
# Setting the debug environment variable NUMBA_DPPY_OFFLOAD_DIAGNOSTICS provides emission of the parallel and offload diagnostics information based on produced parallel transforms. The level of detail depends on the integer value between 1 and 4 that is set to the environment variable (higher is more detailed). In the "Auto-offloading" section there is the information on which device (device name) this kernel was offloaded.
#
# ## Writing Explicit Kernels in numba-dppy
#
# Writing a SYCL kernel using the `@numba_dppy.kernel` decorator has similar syntax to writing OpenCL kernels. As such, the numba-dppy module provides similar indexing and other functions as OpenCL. The indexing functions supported inside a `numba_dppy.kernel` are:
#
# * numba_dppy.get_local_id : Gets the local ID of the item
# * numba_dppy.get_local_size: Gets the local work group size of the device
# * numba_dppy.get_group_id : Gets the group ID of the item
# * numba_dppy.get_num_groups: Gets the number of gropus in a worksgroup
#
# Refer https://intelpython.github.io/numba-dppy/latest/user_guides/kernel_programming_guide/index.html for more details.
#
# The following code demonstrates usage of simple `@numba_dppy.kernel` code. Inspect code; there are no modifications necessary:
# 1. Inspect the following code cell and click Run (▶) to save the code to file.
# 2. Next, run (▶) the cell in the __Build and Run__ section following the code to compile and execute the code.
# +
# %%writefile lab/simple_2d.py
# #! /usr/bin/env python
# Copyright 2020, 2021 Intel Corporation
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
import dpctl
import numba_dppy as dppy
import numpy as np
@dppy.kernel
def data_parallel_sum(a, b, c):
"""
A two-dimensional vector addition example using the ``kernel`` decorator.
"""
i = dppy.get_global_id(0)
j = dppy.get_global_id(1)
c[i, j] = a[i, j] + b[i, j]
def driver(a, b, c, global_size):
print("before A: ", a)
print("before B: ", b)
data_parallel_sum[global_size, dppy.DEFAULT_LOCAL_SIZE](a, b, c)
print("after C : ", c)
def main():
# Array dimensions
X = 8
Y = 8
global_size = X, Y
a = np.arange(X * Y, dtype=np.float32).reshape(X, Y)
b = np.array(np.random.random(X * Y), dtype=np.float32).reshape(X, Y)
c = np.ones_like(a).reshape(X, Y)
# Use the environment variable SYCL_DEVICE_FILTER to change the default device.
# See https://github.com/intel/llvm/blob/sycl/sycl/doc/EnvironmentVariables.md#sycl_device_filter.
device = dpctl.select_default_device()
print("Using device ...")
device.print_device_info()
with dpctl.device_context(device):
driver(a, b, c, global_size)
print(c)
print("Done...")
if __name__ == "__main__":
main()
# -
# _If the Jupyter cells are not responsive or if they error out when you compile the code samples, please restart the Jupyter Kernel:
# "Kernel->Restart Kernel and Clear All Outputs" and compile the code samples again_
# #### Build and Run
# Select the cell below and click run ▶ to compile and execute the code:
# ! chmod 755 q; chmod 755 run_simple_2d.sh; if [ -x "$(command -v qsub)" ]; then ./q run_simple_2d.sh; else ./run_simple_2d.sh; fi
# ## Lab Exercise: Matrix multiplication using @kernel.decorator
# * In this lab we provide with the source code that computes matrix multiplication using the numba-dppy kernel decorator.
# * Follow the __Step1 and Step 2 and Step 3 in the below code__.
#
# 1. Select the code cell below, __follow the STEPS 1 to 3__ in the code comments, click run ▶ to save the code to file.
# 2. Next run ▶ the cell in the __Build and Run__ section below the code to compile and execute the code.
# +
# %%writefile lab/matrix_mul.py
# #! /usr/bin/env python
# Copyright 2021 Intel Corporation
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
from timeit import default_timer as time
import numpy as np
import numba_dppy as dppy
import dpctl
#***Step1: Uncomment the following lines to enable the dppy.kernel decorator***
@dppy.kernel
def dppy_gemm(a, b, c):
"""
A basic DGEMM implemented as a ``kernel`` function.
"""
i = dppy.get_global_id(0)
j = dppy.get_global_id(1)
if i >= c.shape[0] or j >= c.shape[1]:
return
c[i, j] = 0
for k in range(c.shape[0]):
c[i, j] += a[i, k] * b[k, j]
# Array dimensions
X = 1024
Y = 16
global_size = X, X
griddim = X, X
blockdim = Y, Y
def driver(a, b, c):
# Invoke the kernel
dppy_gemm[griddim, blockdim](a, b, c)
def main():
a = np.arange(X * X, dtype=np.float32).reshape(X, X)
b = np.array(np.random.random(X * X), dtype=np.float32).reshape(X, X)
c = np.ones_like(a).reshape(X, X)
# Use the environment variable SYCL_DEVICE_FILTER to change the default device.
# See https://github.com/intel/llvm/blob/sycl/sycl/doc/EnvironmentVariables.md#sycl_device_filter.
device = dpctl.select_default_device()
print("Using device ...")
device.print_device_info()
#***Step2: Uncomment the following lines to set the device context and target a GPU***
with dpctl.device_context(device):
driver(a, b, c)
# Host compute using standard NumPy
Amat = np.matrix(a)
Bmat = np.matrix(b)
Cans = Amat * Bmat
# Check result
assert np.allclose(c, Cans)
print("Done...",c)
if __name__ == "__main__":
main()
# -
# _If the Jupyter cells are not responsive or if they error out when you compile the code samples, please restart the Jupyter Kernel:
# "Kernel->Restart Kernel and Clear All Outputs" and compile the code samples again_
# #### Build and Run
# Select the cell below and click run ▶ to compile and execute the code:
# ! chmod 755 q; chmod 755 run_matrix_mul.sh; if [ -x "$(command -v qsub)" ]; then ./q run_matrix_mul.sh; else ./run_matrix_mul.sh; fi
# # Summary
# In this module you will have learned the following:
# * How the __oneAPI__ programming model can solve the challenges of programming in a heterogeneous world
# * Intel Distribution of Python and its advantages
# * How to use __Numba__ and __numba-dppy__ to write paralle code on CPU and GPU
# * Write explicit kernels using numba-dppy @kernel decorator
| AI-and-Analytics/Jupyter/Numba_DPPY_Essentials_training/01_DPPY_Intro/DPPY_Intro.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow
# language: python
# name: tflow
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/VMBoehm/DeNoPa/blob/master/TrainNVP_and_measure_Reconstruction_Noise.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="D-Fe5G8m1FTC" outputId="ff7c28d3-7cf0-40e4-a8b8-88eb520ae09c"
import numpy as np
import os
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.cm as cm
from matplotlib import rcParams
import sys
import pickle
plt.rcParams.update({'font.family' : 'lmodern', 'font.size': 16,
'axes.labelsize': 16, 'legend.fontsize': 12,
'xtick.labelsize': 16, 'ytick.labelsize': 16, 'axes.titlesize': 16,
'axes.linewidth': 1.5})
# + colab={"base_uri": "https://localhost:8080/", "height": 168} colab_type="code" id="-AEYmOsH1FTI" outputId="5e5d0d77-378f-4cc2-812f-a69eaf08d888"
import tensorflow as tf
import tensorflow_probability as tfp
import tensorflow_hub as hub
tfd = tfp.distributions
tfb = tfp.bijectors
# -
sys.path.append('../')
from vae.model import get_prior, get_posterior, get_likelihood
import vae.create_datasets as crd
likelihood = 'Gauss'
class_label = -1
network_type= 'fully_connected'
data_set = 'mnist'
latent_size = 8
AE = True
if AE:
params = pickle.load(open('../params/params_%s_%s_%d_%d_%s-AE.pkl'%(data_set,likelihood,class_label,latent_size, network_type),'rb'))
else:
params = pickle.load(open('../params/params_%s_%s_%d_%d_%s.pkl'%(data_set,likelihood,class_label,latent_size,network_type),'rb'))
module_dir = params['module_dir']+'/vae'
model_dir = params['model_dir']
params['add_noise']=False
all_subdirs = [os.path.join(module_dir,d) for d in os.listdir(module_dir+'/') if os.path.isdir(os.path.join(module_dir,d))]
latest_subdir = max(all_subdirs, key=os.path.getmtime)
# + colab={} colab_type="code" id="8puPFE90P0aD"
generator_path = os.path.join(latest_subdir,'decoder')
encoder_path = os.path.join(latest_subdir,'encoder')
# -
nvp_depth = 6
indices = np.arange(params['latent_size'])
permutations = [np.random.permutation(indices) for ii in range(nvp_depth)]
train_input_fn, eval_input_fn = crd.build_input_fns(params,label=params['class_label'],flatten=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 118} colab_type="code" id="yvTEYw44O_5q" outputId="fca6575b-6a27-49b2-bca4-8f5b58feaa0b"
tf.reset_default_graph()
lr = tf.placeholder_with_default(0.001,shape=[])
sigma = tf.placeholder_with_default(0.1,shape=[])
sample_size = tf.placeholder_with_default(16,shape=[])
data = train_input_fn()
testdata = eval_input_fn()
encoder = hub.Module(encoder_path, trainable=False)
generator = hub.Module(generator_path, trainable=False)
optimizer = tf.train.AdamOptimizer(learning_rate=lr)
prior = get_prior(params['latent_size'])
posterior = get_posterior(encoder)
likelihood = get_likelihood(generator,params['likelihood'],sigma)
# inspect the model
z_ = posterior(testdata).mean()
prior_sample = prior.sample(sample_size)
x = likelihood(prior_sample).mean()
recon = likelihood(z_).mean()
### ----------- NVP ----------- ###
def init_once(x, name):
return tf.get_variable(name, initializer=x, trainable=False)
nvp_size = [[512, 512] for ii in range(nvp_depth)]
def get_nvp():
base_dis = get_prior(params['latent_size'])
chain = []
perms = [tfb.Permute(permutation=init_once(permutations[ii],name="permutation%d"%ii)) for ii in range(nvp_depth)]
for i,s in enumerate(nvp_size):
if i==0:
chain.append(perms[i])
if i>2:
chain.append(tfb.RealNVP(num_masked=params['latent_size']//2,shift_and_log_scale_fn=tfb.real_nvp_default_template(hidden_layers=s,name='nvp%d'%i, shift_only=True)))
else:
chain.append(tfb.RealNVP(num_masked=params['latent_size']//2,shift_and_log_scale_fn=tfb.real_nvp_default_template(hidden_layers=s,name='nvp%d'%i)))
chain.append(perms[i])
nvp = tfd.TransformedDistribution(distribution=base_dis,bijector=tfb.Chain(chain),name='mynvp')
return nvp
def nvp_module_spec():
z_sample = tf.placeholder(tf.float32, shape=[None,params['latent_size']])
sample_size = tf.placeholder(tf.int32, shape=[])
u_sample = tf.placeholder(tf.float32, shape=[None,params['latent_size']])
nvp_ = get_nvp()
log_prob = nvp_.log_prob(z_sample)
nvp_sample = nvp_.sample(sample_size)
nvp_fwd_pass = nvp_.bijector.forward(u_sample)
hub.add_signature(inputs={'z_sample':z_sample,'sample_size':sample_size, 'u_sample':u_sample},outputs={'log_prob':log_prob, 'sample':nvp_sample, 'fwd_pass': nvp_fwd_pass})
nvp_spec = hub.create_module_spec(nvp_module_spec)
nvp_funcs = hub.Module(nvp_spec, name='nvp_funcs',trainable=True)
z = posterior(data).mean()
loss = -tf.reduce_mean(nvp_funcs({'z_sample':z,'sample_size':1, 'u_sample':np.zeros((1,params['latent_size']))},as_dict=True)['log_prob'])
opt_op_nvp = optimizer.minimize(loss)
nvp_sample = nvp_funcs({'z_sample':np.zeros((1,params['latent_size'])),'sample_size':1, 'u_sample':prior_sample}, as_dict=True)['fwd_pass']
x_new = likelihood(nvp_sample).mean()
# # ---------------------------end train nvp ----------------- #
# #comment train nvp above section above and uncomment this for loading the trained module
# #nvp_funcs = hub.Module(nvp_func_path, trainable=False)
# + colab={} colab_type="code" id="Soh1tnGH1FTW"
sess = tf.Session()
sess.run(tf.global_variables_initializer())
# -
rec, d = sess.run([recon,data])
x_sample = sess.run(x)
jj=1
plt.figure(figsize=(2*2,4*2))
for ii in range(4):
plt.subplot(4,2,jj)
plt.imshow(d[ii].reshape(28,28),cmap='gray')
plt.axis('off')
jj+=1
plt.subplot(4,2,jj)
plt.imshow(rec[ii].reshape(28,28),cmap='gray',vmin=0, vmax=1)
plt.axis('off')
jj+=1
jj=1
plt.figure(figsize=(4*2,2*2))
for ii in range(8):
plt.subplot(2,4,jj)
plt.imshow(x_sample[ii].reshape(28,28),cmap='gray',vmin=0, vmax=1)
plt.axis('off')
jj+=1
z_sample=[]
for ii in range(1000):
z_sample+=[sess.run(z)]
z_sample=np.asarray(z_sample)
plt.figure(figsize=((latent_size-1)*2+2,2))
for nn in range(latent_size-1):
plt.subplot(1,latent_size-1,nn+1)
plt.scatter(z_sample[:,nn],z_sample[:,nn+1],s=1)
plt.ylabel('latent dim %d'%nn)
plt.xlabel('latent dim %d'%(nn+1))
plt.tight_layout()
# + [markdown] colab_type="text" id="U0xZ_4ActOw9"
# **Training of the NVP**
# + colab={} colab_type="code" id="ggPmR0fs8qEM"
nvp_loss = []
# + colab={} colab_type="code" id="ERmZCW6yBqUR"
ii = 0
nepoch = 0
learning_rate = 1e-4
#train the nvp
while nepoch<1000:
_, ll = sess.run([opt_op_nvp,loss], feed_dict={lr: learning_rate})
nvp_loss+=[ll]
if ii%1000==0:
print(np.mean(nvp_loss[-50::]))
ii+=1
# + colab={} colab_type="code" id="bvfbqZJ4RX9x"
plt.figure()
plt.plot(np.convolve(nvp_loss[::],np.ones((100))/100., mode='valid'),label='nvp loss')
# + colab={} colab_type="code" id="YWmT0AjAeOKs"
s_nvp = sess.run(x_new)
s = sess.run(x)
z_sample_nvp = sess.run(nvp_sample,feed_dict={sample_size:10000})
# -
jj=1
plt.figure(figsize=(4*2,2*2))
for ii in range(8):
plt.subplot(2,4,jj)
plt.imshow(s_nvp[ii].reshape(28,28),cmap='gray',vmin=0, vmax=1)
plt.axis('off')
jj+=1
plt.figure(figsize=((latent_size-1)*2+2,2))
for nn in range(latent_size-1):
plt.subplot(1,latent_size-1,nn+1)
plt.scatter(z_sample_nvp[:,nn],z_sample_nvp[:,nn+1],s=1, label='nvp samples')
plt.scatter(z_sample[:,nn],z_sample[:,nn+1],s=1,alpha=0.8, label='encoded data')
plt.ylabel('latent dim %d'%nn)
plt.xlabel('latent dim %d'%(nn+1))
plt.tight_layout()
plt.legend(loc=(1.01,0.5))
# + colab={} colab_type="code" id="TYtuOw8uB-aO"
nvp_dir = os.path.join(params['module_dir'],'nvp')
if not os.path.isdir(nvp_dir):
os.makedirs(nvp_dir)
nvp_funcs.export(nvp_dir,sess)
# -
| notebooks/InspectResults.ipynb |