code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="WbiEDNVkEG3T"
# # SETUP
# + colab={"base_uri": "https://localhost:8080/"} id="Hp8vMTsAGani" outputId="265ed585-88f4-4deb-8312-25aeeedc3e36"
# !git clone 'https://github.com/radiantearth/mlhub-tutorials.git'
# + colab={"base_uri": "https://localhost:8080/"} id="l_MzJvZxGakE" outputId="fb919f15-d6be-42ec-d5e5-821361323034"
# !pip install -r '/content/mlhub-tutorials/notebooks/South Africa Crop Types Competition/requirements.txt' -q
# + id="8YFLc43syf6x"
exit(0)
# + [markdown] id="T-RSB5omEInc"
# # LIBRARIES
# + id="V-xKDNhkyhTZ"
# Required libraries
import os
import tarfile
import json
from pathlib import Path
from radiant_mlhub.client import _download as download_file
import datetime
import rasterio
import numpy as np
import pandas as pd
from sklearn.ensemble import RandomForestClassifier
from sklearn.metrics import log_loss
from sklearn.model_selection import StratifiedShuffleSplit
os.environ['MLHUB_API_KEY'] = '96f33e4c9510d0d369d881c6fdefa91502829db09f41e0c92cba8b02fede920b'
# + [markdown] id="2S1h19sfEKEr"
# # DOWNLOAD DATA
# + id="dBqBu3cMyjQA"
DOWNLOAD_S1 = True # If you set this to true then the Sentinel-1 data will be downloaded which is not needed in this notebook.
# Select which imagery bands you'd like to download here:
DOWNLOAD_BANDS = {
'B01': False,
'B02': False,
'B03': False,
'B04': False,
'B05': False,
'B06': False,
'B07': False,
'B08': False,
'B8A': False,
'B09': False,
'B11': False,
'B12': False,
'CLM': False
}
# In this model we will only use Green, Red and NIR bands. You can select to download any number of bands.
# Our choice relies on the fact that vegetation is most sensitive to these bands.
# We also donwload the CLM or Cloud Mask layer to exclude cloudy data from the training phase.
# You can also do a feature selection, and try different combination of bands to see which ones will result in a better accuracy.
# + id="g68iGaSgylS4"
FOLDER_BASE = 'ref_south_africa_crops_competition_v1'
def download_archive(archive_name):
if os.path.exists(archive_name.replace('.tar.gz', '')):
return
print(f'Downloading {archive_name} ...')
download_url = f'https://radiant-mlhub.s3.us-west-2.amazonaws.com/archives/{archive_name}'
download_file(download_url, '.')
print(f'Extracting {archive_name} ...')
with tarfile.open(archive_name) as tfile:
tfile.extractall()
os.remove(archive_name)
for split in ['test']:
# Download the labels
labels_archive = f'{FOLDER_BASE}_{split}_labels.tar.gz'
download_archive(labels_archive)
# Download Sentinel-1 data
if DOWNLOAD_S1:
s1_archive = f'{FOLDER_BASE}_{split}_source_s1.tar.gz'
download_archive(s1_archive)
for band, download in DOWNLOAD_BANDS.items():
if not download:
continue
s2_archive = f'{FOLDER_BASE}_{split}_source_s2_{band}.tar.gz'
download_archive(s2_archive)
def resolve_path(base, path):
return Path(os.path.join(base, path)).resolve()
def load_df(collection_id):
split = collection_id.split('_')[-2]
collection = json.load(open(f'{collection_id}/collection.json', 'r'))
rows = []
item_links = []
for link in collection['links']:
if link['rel'] != 'item':
continue
item_links.append(link['href'])
for item_link in item_links:
item_path = f'{collection_id}/{item_link}'
current_path = os.path.dirname(item_path)
item = json.load(open(item_path, 'r'))
tile_id = item['id'].split('_')[-1]
for asset_key, asset in item['assets'].items():
rows.append([
tile_id,
None,
None,
asset_key,
str(resolve_path(current_path, asset['href']))
])
for link in item['links']:
if link['rel'] != 'source':
continue
source_item_id = link['href'].split('/')[-2]
if source_item_id.find('_s1_') > 0 and not DOWNLOAD_S1:
continue
elif source_item_id.find('_s1_') > 0:
for band in ['VV', 'VH']:
asset_path = Path(f'{FOLDER_BASE}_{split}_source_s1/{source_item_id}/{band}.tif').resolve()
date = '-'.join(source_item_id.split('_')[10:13])
rows.append([
tile_id,
f'{date}T00:00:00Z',
's1',
band,
asset_path
])
if source_item_id.find('_s2_') > 0:
for band, download in DOWNLOAD_BANDS.items():
if not download:
continue
asset_path = Path(f'{FOLDER_BASE}_{split}_source_s2_{band}/{source_item_id}_{band}.tif').resolve()
date = '-'.join(source_item_id.split('_')[10:13])
rows.append([
tile_id,
f'{date}T00:00:00Z',
's2',
band,
asset_path
])
return pd.DataFrame(rows, columns=['tile_id', 'datetime', 'satellite_platform', 'asset', 'file_path'])
competition_test_df = load_df(f'{FOLDER_BASE}_test_labels')
# + id="C-dUI1_AzOHw" colab={"base_uri": "https://localhost:8080/"} outputId="10cbf2f0-0156-48ba-fa67-e1093fec4479"
import gc
gc.collect()
# + [markdown] id="fDHGa05LEPJD"
# # CREATE DATA
# + id="JxPaJy6m6EBg"
# This DataFrame lists all types of assets including documentation of the data.
# In the following, we will use the Sentinel-2 bands as well as labels.
tile_ids_test = competition_test_df['tile_id'].unique()
# + id="mG5fN4cI65qQ"
from tqdm import tqdm_notebook
import gc
import warnings
warnings.simplefilter('ignore')
# + id="harn9qwG65oB"
n_obs = 5
# + id="YB2CV1T96Uuh"
# %%time
competition_test_df['Month'] = pd.to_datetime(competition_test_df['datetime']).dt.month
X = np.empty((0, 2*8),dtype=np.float16)
y = np.empty((0, 1),dtype=np.float16)
field_ids = np.empty((0, 1),np.float16)
for tile_id in tqdm_notebook(tile_ids_test[0:tile_ids_test.shape[0]]):
tile_df = competition_test_df[competition_test_df['tile_id']==tile_id]
field_id_src = rasterio.open(tile_df[tile_df['asset']=='field_ids']['file_path'].values[0])
field_id_array = field_id_src.read(1).flatten()
nonzeroidx = np.nonzero(field_id_array)[0]
field_ids = np.append(field_ids, field_id_array[nonzeroidx])
tile_date_times = tile_df[tile_df['satellite_platform']=='s1']['Month'].unique().tolist()
X_tile = np.empty((nonzeroidx.shape[0], 0),dtype=np.float16)
n_X = 0
for date_time_idx in range(0,len(tile_date_times)):
vv_src = rasterio.open(tile_df[(tile_df['Month']==month) & (tile_df['asset']=='VV')]['file_path'].values[0])
vv_array = np.expand_dims(vv_src.read(1).flatten()[nonzeroidx], axis=1)
vh_src = rasterio.open(tile_df[(tile_df['Month']==month) & (tile_df['asset']=='VH')]['file_path'].values[0])
vh_array = np.expand_dims(vh_src.read(1).flatten()[nonzeroidx], axis=1)
X_tile = np.append(X_tile,vv_array, axis = 1)
X_tile = np.append(X_tile,vh_array, axis = 1)
del vv_array,vh_array
del vv_src,vh_src
X = np.append(X, X_tile, axis=0)
del X_tile , field_id_array , field_id_src
gc.collect()
# + colab={"base_uri": "https://localhost:8080/"} id="nStutoS0Po_N" outputId="8578ba50-6b6a-423c-8e28-54de56c8709d"
gc.collect()
# + [markdown] id="pONjdGDAW4BM"
# # Data
# + id="5otf8f9CDM85"
data = pd.DataFrame(X)
data['field_id'] = field_ids
# + [markdown] id="4VXCeF0LPLvN"
# * **Reduce Memory Usage**
# + id="BF8FnGvhPAuF"
def reduce_mem_usage(df, verbose=True):
numerics = ['int16', 'int32', 'int64', 'float16', 'float32', 'float64']
start_mem = df.memory_usage().sum() / 1024**2
for col in df.columns:
col_type = df[col].dtypes
if col_type in numerics:
c_min = df[col].min()
c_max = df[col].max()
if str(col_type)[:3] == 'int':
if c_min > np.iinfo(np.int8).min and c_max < np.iinfo(np.int8).max:
df[col] = df[col].astype(np.int8)
elif c_min > np.iinfo(np.int16).min and c_max < np.iinfo(np.int16).max:
df[col] = df[col].astype(np.int16)
elif c_min > np.iinfo(np.int32).min and c_max < np.iinfo(np.int32).max:
df[col] = df[col].astype(np.int32)
elif c_min > np.iinfo(np.int64).min and c_max < np.iinfo(np.int64).max:
df[col] = df[col].astype(np.int64)
else:
if c_min > np.finfo(np.float16).min and c_max < np.finfo(np.float16).max:
df[col] = df[col].astype(np.float16)
elif c_min > np.finfo(np.float32).min and c_max < np.finfo(np.float32).max:
df[col] = df[col].astype(np.float32)
else:
df[col] = df[col].astype(np.float64)
end_mem = df.memory_usage().sum() / 1024**2
print('Memory usage after optimization is: {:.2f} MB'.format(end_mem))
print('Decreased by {:.1f}%'.format(100 * (start_mem - end_mem) / start_mem))
return df
# + id="5UWYy1FHPArw"
data = reduce_mem_usage(data)
# + id="tNQPTwawf9ul"
data.head()
# + id="8kHqXT3FRxeE"
gc.collect()
# + id="2-K4HNAaXYph"
# Each field has several pixels in the data. Here our goal is to build a Random Forest (RF) model using the average values
# of the pixels within each field. So, we use `groupby` to take the mean for each field_id
data_grouped = data.groupby('field_id').mean().reset_index()
data_grouped = reduce_mem_usage(data_grouped)
data_grouped
# + id="wn2qm85MVHYr"
feat = ["VV","VH"]
columns = [x + '_Month4' for x in feat] + [x + '_Month5' for x in feat] + \
[x + '_Month6' for x in feat] + [x + '_Month7' for x in feat] + \
[x + '_Month8' for x in feat] + [x + '_Month9' for x in feat] + \
[x + '_Month10' for x in feat] + [x + '_Month11' for x in feat]
columns = ['field_id'] + columns
# + id="DU-QIMg0V2tU"
data_grouped.columns = columns
data_grouped
# + colab={"base_uri": "https://localhost:8080/"} id="UDGsQHL5R7Ip" outputId="341e5aa1-f2ab-4d8a-f27e-7d69edcb9034"
from google.colab import drive
drive.mount('/content/drive')
# + id="XYtA3TmeUvly"
data_grouped.to_csv('S1TestObs1.csv',index=False)
os.makedirs('/content/drive/MyDrive/RadiantEarth',exist_ok=True)
os.makedirs('/content/drive/MyDrive/RadiantEarth/Data',exist_ok=True)
os.makedirs('/content/drive/MyDrive/RadiantEarth/Data/TestS1',exist_ok=True)
# !cp 'S1TestObs1.csv' "/content/drive/MyDrive/RadiantEarth/Data/TestS1/"
|
3rd place - ASSAZZIN/XL/Data Creation/S1-Test/S1Test_Observation1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ridge Regression
#
# **Ridge Regression** extends linear regression by providing L2 regularization of the coefficients. It can reduce the variance of the predictors, and improves the conditioning of the problem.
#
# The model can take array-like objects, either in host as NumPy arrays or in device (as Numba or cuda_array_interface-compliant), as well as cuDF DataFrames as the input.
#
# For information about cuDF, refer to the [cuDF documentation](https://docs.rapids.ai/api/cudf/stable).
#
# For information about cuML's ridge regression API: https://rapidsai.github.io/projects/cuml/en/stable/api.html#cuml.Ridge.
# ## Imports
import cudf
from cuml import make_regression, train_test_split
from cuml.metrics.regression import r2_score
from cuml.linear_model import Ridge as cuRidge
from sklearn.linear_model import Ridge as skRidge
# ## Define Parameters
# +
n_samples = 2**20
n_features = 399
random_state = 23
# -
# ## Generate Data
# +
# %%time
X, y = make_regression(n_samples=n_samples, n_features=n_features, random_state=0)
X = cudf.DataFrame.from_gpu_matrix(X)
y = cudf.DataFrame.from_gpu_matrix(y)[0]
X_cudf, X_cudf_test, y_cudf, y_cudf_test = train_test_split(X, y, test_size = 0.2, random_state=random_state)
# -
# Copy dataset from GPU memory to host memory.
# This is done to later compare CPU and GPU results.
X_train = X_cudf.to_pandas()
X_test = X_cudf_test.to_pandas()
y_train = y_cudf.to_pandas()
y_test = y_cudf_test.to_pandas()
# ## Scikit-learn Model
#
# ### Fit, predit and evaluate
# +
# %%time
ridge_sk = skRidge(fit_intercept=False, normalize=True, alpha=0.1)
ridge_sk.fit(X_train, y_train)
# -
# %%time
predict_sk= ridge_sk.predict(X_test)
# %%time
r2_score_sk = r2_score(y_cudf_test, predict_sk)
# ## cuML Model
#
# ### Fit, predit and evaluate
# +
# %%time
# Run the cuml ridge regression model to fit the training dataset.
# Eig is the faster algorithm, but svd is more accurate.
# In general svd uses significantly more memory and is slower than eig.
# If using CUDA 10.1, the memory difference is even bigger than in the other supported CUDA versions
ridge_cuml = cuRidge(fit_intercept=False, normalize=True, solver='eig', alpha=0.1)
ridge_cuml.fit(X_cudf, y_cudf)
# -
# %%time
predict_cuml = ridge_cuml.predict(X_cudf_test)
# %%time
r2_score_cuml = r2_score(y_cudf_test, predict_cuml)
# ## Compare Results
print("R^2 score (SKL): %s" % r2_score_sk)
print("R^2 score (cuML): %s" % r2_score_cuml)
|
cuml/ridge_regression_demo.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/lopez-isaac/DS-Unit-2-Kaggle-Challenge/blob/master/module4/LS_DS_224_assignment.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="PxGvNaaSgr52" colab_type="text"
# Lambda School Data Science
#
# *Unit 2, Sprint 2, Module 4*
#
# ---
# + [markdown] colab_type="text" id="nCc3XZEyG3XV"
# # Classification Metrics
#
# ## Assignment
# - [ ] If you haven't yet, [review requirements for your portfolio project](https://lambdaschool.github.io/ds/unit2), then submit your dataset.
# - [ ] Plot a confusion matrix for your Tanzania Waterpumps model.
# - [ ] Continue to participate in our Kaggle challenge. Every student should have made at least one submission that scores at least 70% accuracy (well above the majority class baseline).
# - [ ] Submit your final predictions to our Kaggle competition. Optionally, go to **My Submissions**, and _"you may select up to 1 submission to be used to count towards your final leaderboard score."_
# - [ ] Commit your notebook to your fork of the GitHub repo.
# - [ ] Read [Maximizing Scarce Maintenance Resources with Data: Applying predictive modeling, precision at k, and clustering to optimize impact](https://towardsdatascience.com/maximizing-scarce-maintenance-resources-with-data-8f3491133050), by Lambda DS3 student <NAME>. His blog post extends the Tanzania Waterpumps scenario, far beyond what's in the lecture notebook.
#
#
# ## Stretch Goals
#
# ### Reading
# - [Attacking discrimination with smarter machine learning](https://research.google.com/bigpicture/attacking-discrimination-in-ml/), by Google Research, with interactive visualizations. _"A threshold classifier essentially makes a yes/no decision, putting things in one category or another. We look at how these classifiers work, ways they can potentially be unfair, and how you might turn an unfair classifier into a fairer one. As an illustrative example, we focus on loan granting scenarios where a bank may grant or deny a loan based on a single, automatically computed number such as a credit score."_
# - [Notebook about how to calculate expected value from a confusion matrix by treating it as a cost-benefit matrix](https://github.com/podopie/DAT18NYC/blob/master/classes/13-expected_value_cost_benefit_analysis.ipynb)
# - [Simple guide to confusion matrix terminology](https://www.dataschool.io/simple-guide-to-confusion-matrix-terminology/) by <NAME>, with video
# - [Visualizing Machine Learning Thresholds to Make Better Business Decisions](https://blog.insightdatascience.com/visualizing-machine-learning-thresholds-to-make-better-business-decisions-4ab07f823415)
#
#
# ### Doing
# - [ ] Share visualizations in our Slack channel!
# - [ ] RandomizedSearchCV / GridSearchCV, for model selection. (See module 3 assignment notebook)
# - [ ] More Categorical Encoding. (See module 2 assignment notebook)
# - [ ] Stacking Ensemble. (See below)
#
# ### Stacking Ensemble
#
# Here's some code you can use to "stack" multiple submissions, which is another form of ensembling:
#
# ```python
# import pandas as pd
#
# # Filenames of your submissions you want to ensemble
# files = ['submission-01.csv', 'submission-02.csv', 'submission-03.csv']
#
# target = 'status_group'
# submissions = (pd.read_csv(file)[[target]] for file in files)
# ensemble = pd.concat(submissions, axis='columns')
# majority_vote = ensemble.mode(axis='columns')[0]
#
# sample_submission = pd.read_csv('sample_submission.csv')
# submission = sample_submission.copy()
# submission[target] = majority_vote
# submission.to_csv('my-ultimate-ensemble-submission.csv', index=False)
# ```
# + colab_type="code" id="lsbRiKBoB5RE" colab={}
# %%capture
import sys
# If you're on Colab:
if 'google.colab' in sys.modules:
DATA_PATH = 'https://raw.githubusercontent.com/LambdaSchool/DS-Unit-2-Kaggle-Challenge/master/data/'
# !pip install category_encoders==2.*
# !pip install matplotlib==3.1.0
# If you're working locally:
else:
DATA_PATH = '../data/'
# + colab_type="code" id="BVA1lph8CcNX" colab={}
import pandas as pd
# Merge train_features.csv & train_labels.csv
train = pd.merge(pd.read_csv(DATA_PATH+'waterpumps/train_features.csv'),
pd.read_csv(DATA_PATH+'waterpumps/train_labels.csv'))
# Read test_features.csv & sample_submission.csv
test = pd.read_csv(DATA_PATH+'waterpumps/test_features.csv')
sample_submission = pd.read_csv(DATA_PATH+'waterpumps/sample_submission.csv')
# + id="say_7eiSgr6C" colab_type="code" outputId="2266f29d-bc16-4488-d65c-86fef62e6e90" colab={"base_uri": "https://localhost:8080/", "height": 34}
import matplotlib
print(matplotlib.__version__)
# + [markdown] id="ktV060KljTH6" colab_type="text"
# #cleanup
# + id="lh1cc4evixBw" colab_type="code" colab={}
## split train and val data sets
from sklearn.model_selection import train_test_split
train, val = train_test_split(train, train_size=.80, test_size=.20,
stratify=train['status_group'], random_state=42)
# + id="C1_AwGgvjain" colab_type="code" colab={}
import numpy as np
def wrangle(X):
"""Wrangle train, validate, and test sets in the same way"""
# Prevent SettingWithCopyWarning
X = X.copy()
# About 3% of the time, latitude has small values near zero,
# outside Tanzania, so we'll treat these values like zero.
X['latitude'] = X['latitude'].replace(-2e-08, 0)
# When columns have zeros and shouldn't, they are like null values.
# So we will replace the zeros with nulls, and impute missing values later.
# Also create a "missing indicator" column, because the fact that
# values are missing may be a predictive signal.
cols_with_zeros = ['longitude', 'latitude', 'construction_year',
'gps_height', 'population']
for col in cols_with_zeros:
X[col] = X[col].replace(0, np.nan)
X[col+'_MISSING'] = X[col].isnull()
# Drop duplicate columns
duplicates = ['quantity_group', 'payment_type']
X = X.drop(columns=duplicates)
# Drop recorded_by (never varies) and id (always varies, random)
unusable_variance = ['recorded_by', 'id']
X = X.drop(columns=unusable_variance)
# Convert date_recorded to datetime
X['date_recorded'] = pd.to_datetime(X['date_recorded'], infer_datetime_format=True)
# Extract components from date_recorded, then drop the original column
X['year_recorded'] = X['date_recorded'].dt.year
X['month_recorded'] = X['date_recorded'].dt.month
X['day_recorded'] = X['date_recorded'].dt.day
X = X.drop(columns='date_recorded')
# Engineer feature: how many years from construction_year to date_recorded
X['years'] = X['year_recorded'] - X['construction_year']
X['years_MISSING'] = X['years'].isnull()
# return the wrangled dataframe
return X
train = wrangle(train)
val = wrangle(val)
test = wrangle(test)
# + id="cAEUcAFJjkPX" colab_type="code" colab={}
## clean the nan of construction year
train['construction_year'].median()
#apply change to all data sets
data_sets = [train,val,test]
for x in data_sets:
x['construction_year'] = x['construction_year'].replace(np.NaN,1986)
#how many years in service
for x in data_sets:
x['pump_age'] = (x['year_recorded'] - x['construction_year'])
# + id="iPMYCV2YjoUa" colab_type="code" outputId="aa5723c8-5ab5-4a80-fbae-0b873d4a4505" colab={"base_uri": "https://localhost:8080/", "height": 54}
### make features and target
target = 'status_group'
train_features = train.drop(columns=[target])
# Get a list of the numeric features
numeric_features = train_features.select_dtypes(include='number').columns.tolist()
#Get a series with the cardinality of the nonnumeric features
cardinality = train_features.select_dtypes(exclude='number').nunique()
# Get a list of all categorical features with cardinality <= 50
low_categorical_features = cardinality[cardinality <= 50].index.tolist()
#get a list of high categorical features with cardinality >= 50
high_categorical_features = cardinality[cardinality >= 50].index.tolist()
# Combine the lists
features = numeric_features + low_categorical_features + high_categorical_features
print(features)
# + id="nzQ8KdHHmE8D" colab_type="code" colab={}
X_train = train[features]
y_train = train[target]
X_val = val[features]
y_val = val[target]
X_test = test[features]
# + id="0qt98s6skPDf" colab_type="code" colab={}
#generare y_pred
import category_encoders as ce
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import make_pipeline
from sklearn.ensemble import RandomForestClassifier
# + id="Emj3xZKFkYJP" colab_type="code" colab={}
Working_condition = make_pipeline(
ce.OneHotEncoder(cols=low_categorical_features),
ce.OrdinalEncoder(),
SimpleImputer(missing_values=np.nan,strategy='median'),
RandomForestClassifier(n_estimators=100,max_depth=20, random_state=42, n_jobs=-1)
)
# + id="T86WyQ0bkkIn" colab_type="code" outputId="7a8f4888-d9f8-4c2f-fdca-0065430aee1b" colab={"base_uri": "https://localhost:8080/", "height": 34}
from sklearn.metrics import accuracy_score
# Fit on train, score on val
Working_condition.fit(X_train, y_train)
y_pred = Working_condition.predict(X_val)
print('Validation Accuracy', accuracy_score(y_val, y_pred))
# + [markdown] id="o7ifOlNJjysE" colab_type="text"
# #confusion matrix
# + id="Q1IK56Vuj1vS" colab_type="code" colab={}
from sklearn.metrics import confusion_matrix
# + id="VmeBcyGyj-Pg" colab_type="code" outputId="be78993a-8aa4-4ede-8229-458c19336332" colab={"base_uri": "https://localhost:8080/", "height": 68}
#very basic matix
confusion_matrix(y_val, y_pred)
# + id="9AVsJgxMnTDr" colab_type="code" outputId="4e4945c5-276e-4fa2-df25-f24d9fbccbc8" colab={"base_uri": "https://localhost:8080/", "height": 51}
# We need to get labels
from sklearn.utils.multiclass import unique_labels
unique_labels(y_val)
# + id="ov_kpM7HpDRL" colab_type="code" outputId="df8344c8-2349-4f77-981c-b89b8abdf8d5" colab={"base_uri": "https://localhost:8080/", "height": 119}
# 1. Check that our labels are correct
# add predicted and actual before lables
def plot_confusion_matrix(y_true, y_pred):
labels = unique_labels(y_true)
columns = [f'Predicted {label}' for label in labels]
index = [f'Actual {label}' for label in labels]
return columns, index
plot_confusion_matrix(y_val, y_pred)
# + id="tFJULzkVqs4h" colab_type="code" outputId="c03083b3-3172-4f0b-c8e6-395af42dd043" colab={"base_uri": "https://localhost:8080/", "height": 142}
#1st way = Make it a pandas dataframe
def plot_confusion_matrix(y_true, y_pred):
labels = unique_labels(y_true)
columns = [f'Predicted {label}' for label in labels]
index = [f'Actual {label}' for label in labels]
#top cell with this new line added
table = pd.DataFrame(confusion_matrix(y_true, y_pred),
columns=columns, index=index)
return table
plot_confusion_matrix(y_val, y_pred)
# + id="Op0jqQeyrKQ-" colab_type="code" outputId="0537c4bb-a536-4015-f917-584b3f832cc8" colab={"base_uri": "https://localhost:8080/", "height": 422}
import seaborn as sns
#2nd way = heatmap
def plot_confusion_matrix(y_true, y_pred):
labels = unique_labels(y_true)
columns = [f'Predicted {label}' for label in labels]
index = [f'Actual {label}' for label in labels]
table = pd.DataFrame(confusion_matrix(y_true, y_pred),
columns=columns, index=index)
#build of pandas DF from top cell
return sns.heatmap(table, annot=True, fmt='d', cmap='viridis')
plot_confusion_matrix(y_val, y_pred);
|
module4/LS_DS_224_assignment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# Description:
# A simple and well formatted jupyter notebook for extracting necessary
# information from csv of sound-activity project under Think-IOT Lab in
# Dr. B. C. Roy Engineering College.
#
# Author: <NAME>
#
# Dependency:
# pandas
#
#
import pandas as pd
# +
# Initializations and operation Specifications:
file_name = "data_files/raw_data.csv"
header = None # Default: 'infer'
col_to_get = '0, 1-2, 5' # To retain all collumns in DataFrame set: col_to_get = None / ''
col_name = "sound, date-time, reading #1, distance"
# +
# Function definitions:
def dataGlimse(DataFrm, dataAbt = "RAW DATA"):
print("\n\n-------- DATA INFO --------\n")
print(DataFrm.info())
print("\n\nDataFrame Shape: ",DataFrm.shape)
print("\n\nDataFrame Indexing: ",DataFrm.index)
print("\n\n--------",dataAbt,"GLIMSE --------")
print("\n# Head #\n")
print(DataFrm.head())
print("\n# Tail #\n")
print(DataFrm.tail())
print("\n\n--------",dataAbt,"GLIMSE END --------")
# -
reviews = pd.read_csv(file_name, header=header)
dataGlimse(reviews, "RAW DATA")
new_reviews = reviews.dropna()
# +
if col_to_get != '' and col_to_get != None:
col_nos = []
for cols in col_to_get.split(','):
cols = cols.replace(' ', '').split('-')
if len(cols) == 1:
col_nos = col_nos+[int(cols[0])]
else:
col_nos = col_nos+list(range(int(cols[0]),int(cols[1]) + 1))
new_reviews = new_reviews[col_nos]
if col_name != None:
new_reviews.columns = [name.strip() for name in col_name.split(',')]
dataGlimse(new_reviews, "SLICED DATA")
# -
|
Sound data Analyzer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # NetworkX
# NetworkX is a Python language software package for the creation, manipulation, and study of the structure, dynamics, and function of complex networks.
#
# With NetworkX you can load and store networks in standard and nonstandard data formats, generate many types of random and classic networks, analyze network structure, build network models, design new network algorithms, draw networks, and much more.
#
# Library documentation: <a>https://networkx.github.io/</a>
import networkx as nx
G = nx.Graph()
# basic add nodes
G.add_node(1)
G.add_nodes_from([2, 3])
# add a group of nodes at once
H = nx.path_graph(10)
G.add_nodes_from(H)
# add another graph itself as a node
G.add_node(H)
# add edges using similar methods
G.add_edge(1, 2)
e = (2, 3)
G.add_edge(*e)
G.add_edges_from([(1, 2), (1, 3)])
G.add_edges_from(H.edges())
# can also remove or clear
G.remove_node(H)
G.clear()
# repeats are ignored
G.add_edges_from([(1,2),(1,3)])
G.add_node(1)
G.add_edge(1,2)
G.add_node('spam') # adds node "spam"
G.add_nodes_from('spam') # adds 4 nodes: 's', 'p', 'a', 'm'
# get the number of nodes and edges
G.number_of_nodes(), G.number_of_edges()
# access graph edges
G[1]
G[1][2]
# set an attribute of an edge
G.add_edge(1,3)
G[1][3]['color'] = 'blue'
FG = nx.Graph()
FG.add_weighted_edges_from([(1, 2, 0.125), (1, 3, 0.75), (2, 4, 1.2), (3, 4, 0.375)])
for n, nbrs in FG.adjacency():
for nbr, eattr in nbrs.items():
data = eattr['weight']
if data < 0.5: print('(%d, %d, %.3f)' % (n, nbr, data))
# graph attribte
G = nx.Graph(day='Friday')
G.graph
# modifying an attribute
G.graph['day'] = 'Monday'
G.graph
# node attributes
G.add_node(1, time='5pm')
G.add_nodes_from([3], time='2pm')
G.node[1]['room'] = 714
G.nodes(data=True)
# edge attributes (weight is a special numeric attribute)
G.add_edge(1, 2, weight=4.7)
G.add_edges_from([(3, 4), (4, 5)], color='red')
G.add_edges_from([(1, 2 ,{'color': 'blue'}), (2, 3, {'weight' :8})])
G[1][2]['weight'] = 4.7
# directed graph
DG = nx.DiGraph()
DG.add_weighted_edges_from([(1, 2 ,0.5), (3, 1, 0.75)])
DG.out_degree(1, weight='weight')
DG.degree(1, weight='weight')
DG.successors(1)
DG.predecessors(1)
# convert to undirected graph
H = nx.Graph(G)
# basic graph drawing capability
# %matplotlib inline
import matplotlib.pyplot as plt
nx.draw(G)
Tested; Gopal
|
tests/ipython-notebooks/NetworkX.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.3
# language: julia
# name: julia-1.6
# ---
# # Item Collaborative Filtering
# * This notebook implements item-based collaborative filtering
# * Prediction is $\tilde r_{ij} = \dfrac{\sum_{k \in N(j)} w_{kj}^{\lambda_w}r_{ik}^{\lambda_r}}{\sum_{k \in N(j)} w_{kj}^{\lambda_w} + \lambda}$ for item-based collaborative filtering
# * $r_{ij}$ is the rating for user $i$ and item $j$
# * $w_{kj}$ is the similarity between items $j$ and $k$
# * $N(j)$ is the largest $K$ sorted by $w_{kj}$
# * $\lambda_w, \lambda_r, \lambda$ are regularization parameters
residual_alphas = [];
@nbinclude("Alpha.ipynb");
# ## Determine the neighborhoods for each user and item
function read_similarity_matrix(outdir)
read_params(outdir)["S"]
end;
function get_abs_neighborhood(item, S, K)
weights = S[:, item]
# ensure that the neighborhood for an item does not include itself
weights[item] = Inf
K = Int(min(K, length(weights) - 1))
order = partialsortperm(abs.(weights), 2:K+1, rev = true)
order, weights[order]
end;
# +
isnonzero(x) = !isapprox(x, 0.0, atol=eps(Float64))
# exponentially decay x
function decay(x, a)
isnonzero(x) ? sign(x) * abs(x)^a : zero(eltype(a))
end
# each prediction is just the weighted sum of all items in the neighborhood
# we apply regularization terms to decay the weights, ratings, and final prediction
function make_prediction(item, users, R, get_neighborhood, λ)
if item > size(R)[2]
# the item was not in our training set; we have no information
return zeros(length(item))
end
items, weights = get_neighborhood(item)
weights = decay.(weights, λ[1])
predictions = zeros(eltype(weights), length(users))
weight_sum = zeros(eltype(weights), length(users))
for u = 1:length(users)
for (i, weight) in zip(items, weights)
if isnonzero(R[users[u], i])
predictions[u] += weight * decay(R[users[u], i], λ[2])
weight_sum[u] += abs(weight)
end
end
end
for u = 1:length(users)
if isnonzero(weight_sum[u] + λ[3])
predictions[u] /= (weight_sum[u] + λ[3])
end
end
predictions
end;
# -
function collaborative_filtering(training, inference, get_neighborhood, λ)
R = sparse(
training.user,
training.item,
training.rating,
maximum(training.user),
maximum(training.item),
)
preds = zeros(eltype(λ), length(inference.rating))
@tprogress Threads.@threads for item in collect(Set(inference.item))
mask = inference.item .== item
preds[mask] =
make_prediction(item, inference.user[mask], R, get_neighborhood, λ)
end
preds
end;
# +
Base.@kwdef mutable struct cf_params
name::Any
training_residuals::Any
validation_residuals::Any
neighborhood_type::Any
S::Any # the similarity matrix
K::Any # the neighborhood size
λ::Vector{Float64} = [1.0, 1.0, 0.0] # [weight_decay, rating_decay, prediction_decay]
end;
to_dict(x::T) where {T} = Dict(string(fn) => getfield(x, fn) for fn ∈ fieldnames(T));
# -
# ## Item based CF
# +
function get_training(residual_alphas)
get_residuals("training", residual_alphas)
end
function get_validation(residual_alphas)
get_residuals("validation", residual_alphas)
end
function get_inference()
training = get_split("training")
validation = get_split("validation")
test = get_split("test")
RatingsDataset(
user = [training.user; validation.user; test.user],
item = [training.item; validation.item; test.item],
rating = fill(
0.0,
length(training.rating) + length(validation.item) + length(test.item),
),
)
end;
# -
function optimize_model(param)
# unpack parameters
training = get_training(param.training_residuals)
validation = get_validation(param.validation_residuals)
S = read_similarity_matrix(param.S)
K = param.K
neighborhood_types = Dict("abs" => get_abs_neighborhood)
neighborhoods = i -> neighborhood_types[param.neighborhood_type](i, S, K)
# optimize hyperparameters
function validation_mse(λ)
pred = collaborative_filtering(training, validation, neighborhoods, λ)
truth = validation.rating
β = pred \ truth
loss = mse(truth, pred .* β)
@debug "loss: $loss β: $β: λ $λ"
loss
end
res = optimize(
validation_mse,
param.λ,
LBFGS(),
autodiff = :forward,
Optim.Options(show_trace = true, extended_trace = true),
)
param.λ = Optim.minimizer(res)
# save predictions
inference = get_inference()
preds = collaborative_filtering(training, inference, neighborhoods, param.λ)
sparse_preds = sparse(inference.user, inference.item, preds)
function model(users, items, predictions)
result = zeros(length(users))
for i = 1:length(users)
if users[i] <= size(predictions)[1] && items[i] <= size(predictions)[2]
result[i] = predictions[users[i], items[i]]
end
end
result
end
write_predictions(
(users, items) -> model(users, items, sparse_preds),
outdir = param.name,
residual_alphas = param.validation_residuals,
save_training = true,
)
write_params(to_dict(param), outdir = param.name)
end;
|
notebooks/TrainingAlphas/ItemCollaborativeFilteringBase.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Forecasting Energy Demand
#
# ## Data Wrangling
#
# The project consists of two data sets:
# * Hourly electricity demand data from the EIA;
# * Hourly observed weather data from LCD/NOAA.
#
# Additionally to demand and weather data, I'll create features based on time to see how the trends are impacted by day of week, hour, week of year, if is holiday, etc.
#
# To limit the scope of the project, I'll use data from Los Angeles exclusively to validate if is possible to improve electricity demand forecasting using weather data.
# +
import boto3
import io
from sagemaker import get_execution_role
role = get_execution_role()
bucket ='sagemaker-data-energy-demand'
# +
S3_CLIENT = boto3.client('s3')
files_list = S3_CLIENT.list_objects_v2(Bucket=bucket, Prefix='raw_data/weather/')
s3_files = files_list['Contents']
latest_weather_data = max(s3_files, key=lambda x: x['LastModified'])
weather_data_location = 's3://{}/{}'.format(bucket, latest_weather_data['Key'])
# +
import requests
import json
import datetime
import pandas as pd
from scipy import stats
from pandas.io.json import json_normalize
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import warnings
warnings.filterwarnings('ignore')
# -
# ### Electricity data
# Electricity data were retrieved using EIA’s API and then unpacked into a dataframe. The API contain hourly entries from July 2015 to present.
#
# The electricity data required just simple cleaning. There were few null values in the set and a very small number of outliers. Removing outliers cut only ~.01% of the data.
# +
EIA5__API_KEY = '1d48c7c8354cc4408732174250d3e8ff'
REGION_CODE = 'LDWP'
CITY = 'LosAngeles'
def str_to_isodatetime(string):
'''
This function transforms strings to an ISO Datetime.
'''
year = string[:4]
month = string[4:6]
day = string[6:8]
time = string[8:11] + ':00:00+0000'
return year + month + day + time
def eia2dataframe(response):
'''
This function unpacks the JSON file from EIA API into a pandas dataframe.
'''
data = response['series'][0]['data']
dates = []
values = []
for date, demand in data:
if demand is None or demand <= 0:
continue
dates.append(str_to_isodatetime(date))
values.append(float(demand))
df = pd.DataFrame({'datetime': dates, 'demand': values})
df['datetime'] = pd.to_datetime(df['datetime'])
df.set_index('datetime', inplace=True)
df = df.sort_index()
return df
electricity_api_response = requests.get('http://api.eia.gov/series/?api_key=%s&series_id=EBA.%s-ALL.D.H' % (EIA__API_KEY, REGION_CODE)).json()
electricity_df = eia2dataframe(electricity_api_response)
# -
electricity_df.isnull().sum()
print(res)
# ### Observed weather data
# LCD data are not available via NOAA’s API so I manually downloaded from the website as a CSV file which I imported to a pandas DataFrame. As common in data that come from physical sensors, LCD data required extensive cleansing.
#
# The main challenges in cleaning the LCD data was that there were in some cases multiple entries for the same hour. I wanted to have just one entry per hour such that I could eventually align LCD data with the hourly entries in the electricity data.
#
# I wrote a function that group weather data by hour and the mode of the entries for same hour. I performed the cleaning this way because either way, the values for multiple per-hour entries are very similar, so the choice of which entry to keep doesn’t make a real difference.
#
def fix_date(df):
'''
This function goes through the dates in the weather dataframe and if there is more than one record for each
hour, we pick the record closest to the hour and drop the rows with the remaining records for that hour.
This is so we can align this dataframe with the one containing electricity data.
input: Pandas DataFrame
output:
'''
df['date'] = pd.to_datetime(df['date']).dt.tz_localize('UTC')
df['date_rounded'] = df['date'].dt.floor('H')
df.drop('date', axis=1, inplace=True)
df.rename({"date_rounded": "datetime"}, axis=1, inplace=True)
df.set_index('datetime', inplace=True)
last_of_hour = df[~df.index.duplicated(keep='last')]
last_of_hour.sort_index(ascending=True, inplace=True, kind='mergesort')
return last_of_hour
# +
def clean_sky_condition(df):
'''
This function cleans the hourly sky condition column by assigning the hourly sky condition to be the one at the
top cloud layer, which is the best determination of the sky condition, as described by the documentation.
input: Pandas DataFrame
output:
'''
conditions = df['hourlyskyconditions']
new_condition = []
for k, condition in enumerate(conditions):
if type(condition) != str and np.isnan(condition):
new_condition.append(np.nan)
else:
colon_indices = [i for i, char in enumerate(condition) if char == ':']
n_layers = len(colon_indices)
try:
colon_position = colon_indices[n_layers - 1]
if condition[colon_position - 1] == 'V':
condition_code = condition[colon_position - 2 : colon_position]
else:
condition_code = condition[colon_position - 3 : colon_position]
new_condition.append(condition_code)
except:
new_condition.append(np.nan)
df['hourlyskyconditions'] = new_condition
df['hourlyskyconditions'] = df['hourlyskyconditions'].astype('category')
return df
def hourly_degree_days(df):
'''
This function adds hourly heating and cooling degree days to the weather DataFrame.
'''
df['hourlycoolingdegrees'] = df['hourlydrybulbtemperature'].apply(lambda x: x - 65. if x >= 65. else 0.)
df['hourlyheatingdegrees'] = df['hourlydrybulbtemperature'].apply(lambda x: 65. - x if x <= 65. else 0.)
return df
# import csv
weather_df = pd.read_csv(weather_data_location, usecols=['DATE', 'DailyCoolingDegreeDays', 'DailyHeatingDegreeDays', 'HourlyDewPointTemperature', 'HourlyPrecipitation', 'HourlyRelativeHumidity', 'HourlySeaLevelPressure', 'HourlySkyConditions', 'HourlyStationPressure', 'HourlyVisibility', 'HourlyDryBulbTemperature', 'HourlyWindSpeed'],
dtype={
'DATE': object,
'DailyCoolingDegreeDays': object,
'DailyHeatingDegreeDays': object,
'HourlyDewPointTemperature': object,
'HourlyPrecipitation': object,
'HourlyRelativeHumidity': object,
'HourlySeaLevelPressure': object,
'HourlySkyConditions': object,
'HourlyStationPressure': object,
'HourlyVisibility': object,
'HourlyDryBulbTemperature': object,
'HourlyWindSpeed': object
})
# make columns lowercase for easier access
weather_df.columns = [col.lower() for col in weather_df.columns]
# clean dataframe so that there's only one record per hour
weather_df = fix_date(weather_df)
# fill the daily heating and cooling degree days such that each hour in an individual day has the same value
weather_df['dailyheatingdegreedays'] = weather_df['dailyheatingdegreedays'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
weather_df.dailyheatingdegreedays.astype('float64')
weather_df['dailycoolingdegreedays'] = weather_df['dailycoolingdegreedays'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
weather_df.dailycoolingdegreedays.astype('float64')
weather_df['dailyheatingdegreedays'] = weather_df['dailyheatingdegreedays'].bfill()
weather_df['dailycoolingdegreedays'] = weather_df['dailycoolingdegreedays'].bfill()
weather_df = clean_sky_condition(weather_df)
# clean other columns by replacing string based values with floats
# values with an 's' following indicate uncertain measurments. we simply change those to floats and include them like normal
weather_df['hourlyvisibility'] = weather_df['hourlyvisibility'].apply(lambda x: float(x) if str(x)[-1] != 'V' else float(str(x)[:-1]))
weather_df['hourlydrybulbtemperature'] = weather_df['hourlydrybulbtemperature'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
weather_df['hourlydewpointtemperature'] = weather_df['hourlydewpointtemperature'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
# set trace amounts equal to zero and change data type
weather_df['hourlyprecipitation'].where(weather_df['hourlyprecipitation'] != 'T', 0.0, inplace=True)
weather_df['hourlyprecipitation'] = weather_df['hourlyprecipitation'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
weather_df['hourlystationpressure'] = weather_df['hourlystationpressure'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
weather_df['hourlywindspeed'] = weather_df['hourlywindspeed'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
weather_df['hourlyrelativehumidity'] = weather_df['hourlyrelativehumidity'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
weather_df['hourlysealevelpressure'] = weather_df['hourlysealevelpressure'].apply(lambda x: float(x) if str(x)[-1] != 's' else float(str(x)[:-1]))
weather_df.hourlyprecipitation.astype('float64')
weather_df.hourlyvisibility.astype('float64')
weather_df.hourlyrelativehumidity.astype('float64')
weather_df.hourlysealevelpressure.astype('float64')
weather_df.hourlystationpressure.astype('float64')
weather_df.hourlywindspeed.astype('float64')
weather_df = hourly_degree_days(weather_df)
# -
weather_df.hourlyrelativehumidity.astype('float64')
weather_df.hourlysealevelpressure.astype('float64')
weather_df.dtypes
## Cut dataframes based on date to align sources
cut_electricity = electricity_df[:weather_df.index.max()]
cut_weather = weather_df[electricity_df.index.min():]
# ## Dealing with outliers and NaN values
#
# The plot distributions bof the features below is used to determine what columns should be filled by using the median
# and which should be filled according to ffill. The features whose ```medians``` and ```means``` are close together suggest that the ```median``` is a good choice for NaNs.Conversely features whose median and means are further apart suggest the presence of outliers and in this case I use ```ffill``` because we are dealing with time series and values in previous time steps are useful in predicting values for later time steps
# +
diff = max(cut_weather.index) - min(cut_electricity.index)
days, seconds = diff.days, diff.seconds
hours = days * 24 + seconds // 3600
minutes = (seconds % 3600) // 60
seconds = seconds % 60
number_of_steps = hours + 1
# -
print('*** min ***')
print(min(cut_electricity.index))
print(min(cut_weather.index))
print(cut_weather.index.min() == cut_electricity.index.min())
print('*** max ***')
print(max(cut_electricity.index))
print(max(cut_weather.index))
print(cut_weather.index.max() == cut_electricity.index.max())
print('*** instances quantity is equal? ***')
print(cut_weather.shape[0] == cut_electricity.shape[0])
print('*** weather, demand, expected ***')
print(cut_weather.shape[0], cut_electricity.shape[0], number_of_steps)
# +
fill_dict = {'median': ['dailyheatingdegreedays', 'hourlyaltimetersetting', 'hourlydrybulbtemperature', 'hourlyprecipitation', 'hourlysealevelpressure', 'hourlystationpressure', 'hourlywetbulbtempf', 'dailycoolingdegreedays', 'hourlyvisibility', 'hourlywindspeed', 'hourlycoolingdegrees', 'hourlyheatingdegrees'], 'ffill': ['demand', 'hourlydewpointtemperature', 'hourlyrelativehumidity']}
# fill electricity data NaNs
for col in cut_electricity.columns:
if col in fill_dict['median']:
cut_electricity[col].fillna(cut_electricity[col].median(), inplace=True)
else:
cut_electricity[col].fillna(cut_electricity[col].ffill(), inplace=True)
# fill weather data NaNs
for col in cut_weather.columns:
if col == 'hourlyskyconditions':
cut_weather[col].fillna(cut_weather[col].value_counts().index[0], inplace=True)
elif col in fill_dict['median']:
cut_weather[col].fillna(cut_weather[col].median(), inplace=True)
else:
cut_weather[col].fillna(cut_weather[col].ffill(), inplace=True)
# -
print(cut_weather.shape[0] == cut_electricity.shape[0])
electricity_set = set(cut_electricity.index)
weather_set = set(cut_weather.index)
print(len(electricity_set.difference(weather_set)))
# finally merge the data to get a complete dataframe for LA, ready for training
merged_df = cut_weather.merge(cut_electricity, right_index=True, left_index=True, how='inner')
merged_df = pd.get_dummies(merged_df)
merged_df.head()
merged_df.index.name = 'datetime'
if 'hourlyskyconditions_VV' in list(merged_df.columns):
merged_df.drop('hourlyskyconditions_VV', axis=1, inplace=True)
if 'hourlyskyconditions_' in list(merged_df.columns):
merged_df.drop('hourlyskyconditions_', axis=1, inplace=True)
# +
# save as csv file to continue in another notebook
csv_buffer = io.StringIO()
s3_resource = boto3.resource('s3')
key = 'dataframes/%s_dataset.csv' % CITY
merged_df.to_csv(csv_buffer, compression=None)
s3_resource.Object(bucket, key).put(Body=csv_buffer.getvalue())
# -
|
0_WRANGLING.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Model
#
# In this notebook we read in the previously created dataset with features engineered, and then perform the following tasks:
#
# * Split into train and test data. Also use reduced size subsets for quicker evaluation.
# * Evaluate some classification algorithms and paramters.
# * Perform simple cross validation to find the best model.
# * Measure the model performance.
# ## Read in dataset and verify
# Imports
import findspark
findspark.init()
findspark.find()
import pyspark
# Imports for creating spark session
from pyspark import SparkContext, SparkConf
from pyspark.sql import SparkSession
conf = pyspark.SparkConf().setAppName('sparkify-capstone-model').setMaster('local')
sc = pyspark.SparkContext(conf=conf)
spark = SparkSession(sc)
# Imports for modelling, tuning and evaluation
from pyspark.ml.classification import LogisticRegression, RandomForestClassifier, GBTClassifier
from pyspark.ml.evaluation import BinaryClassificationEvaluator, MulticlassClassificationEvaluator
# Imports for visualization and output
import matplotlib.pyplot as plt
from IPython.display import HTML, display
# Read in dataset
conf.set("spark.driver.maxResultSize", "0")
path = "out/features.parquet"
df = spark.read.parquet(path)
# Look at some values about the data to confirm it was read in correctly
print("Dataset set rows/cols: {},{}".format(df.count(), len(df.columns)))
df.printSchema()
df.show(10)
# ## Split
#
# Try three ways to split the data to have an initial idea about how some algorithms perform.
# +
# First, we are going to use just a subset of the dataset becuase doing a lot of tuning and cross validation
# would take too long otherwise.
def createSubset(df, factor):
"""
INPUT:
df: The dataset to split
factor: How much of the dataset to return
OUTPUT:
df_subset: The split subset
"""
df_subset, df_dummy = df.randomSplit([factor, 1 - factor])
return df_subset
df_subset = createSubset(df, .2)
df_subset.count()
# -
# Now we split the subset into train and test.
# Note: Best split factor of 90% was determined by trial and error.
df_train, df_test = df_subset.randomSplit([0.9, 0.1])
print(df_train.count())
print(df_test.count())
# ## Algorithm selection
#
# We will be looking at some of pysparks classificatoin algorhtims:
#
# * Logistic Regression (single)
# * Random Forest (parallel)
# * Gradient-Boosted Tree (sequential)
#
# The most basic, logistic regression is a good start.
# ### LogisticRegression
#
# First we will be creating a function to fit and train the LR model, which we may use many times.
# Then we also create some helpful functions for showing the evaluation of the model with some metrics.
# +
def logisticRegressionPredictions(df_train, df_test, threshold = 0.5, labelCol = "churn", featuresCol = "features"):
""" Fit, evaluate and show results for LogisticRegression
INPUT:
df_train: The training data set
df_test: The testing data set
threshold: The algorithm's threshold for classification.
labelCol: The label column name, "churn" by default.
featuresCol: The label column name, "features" by default.
OUTPUT:
predictions: The model's predictions
"""
# Fit and train model
logreg = LogisticRegression(labelCol = labelCol, featuresCol = featuresCol, threshold = threshold).fit(df_train)
return logreg.evaluate(df_test).predictions
def printConfusionMatrix(tp, fp, tn, fn):
""" Simple function to output a confusion matrix from f/t/n/p values as html table.
INPUT:
data: The array to print as table
OUTPUT:
Prints the array as html table.
"""
html = "<table><tr><td></td><td>Act. True</td><td>False</td></tr>"
html += "<tr><td>Pred. Pos.</td><td>{}</td><td>{}</td></tr>".format(tp, fp)
html += "<tr><td>Negative</td><td>{}</td><td>{}</td></tr>".format(fn, tn)
html += "</table>"
display(HTML(html))
def showEvaluationMetrics(predictions):
""" Calculate and print the some evaluation metrics for the passed predictions.
INPUT:
predictions: The predictions to evaluate and print
OUTPUT:
Just prints the evaluation metrics
"""
# Calculate true, false positives and negatives to calculate further metrics later:
tp = predictions[(predictions.churn == 1) & (predictions.prediction == 1)].count()
tn = predictions[(predictions.churn == 0) & (predictions.prediction == 0)].count()
fp = predictions[(predictions.churn == 0) & (predictions.prediction == 1)].count()
fn = predictions[(predictions.churn == 1) & (predictions.prediction == 0)].count()
printConfusionMatrix(tp, fp, tn, fn)
# Calculate and print metrics
f1 = MulticlassClassificationEvaluator(labelCol = "churn", metricName = "f1") \
.evaluate(predictions)
accuracy = float((tp + tn) / (tp + tn + fp + fn))
recall = float(tp / (tp + fn))
precision = float(tp / (tp + fp))
print("F1: ", f1)
print("Accuracy: ", accuracy)
print("Recall: ", recall)
print("Precision: ", precision)
def plotROC(df_model):
"""
Plot the Receiver Operator Curve for the evaluated model
INPUT:
df_model: The trained model
OUTPUT:
Plots the curve
"""
plt.figure(figsize = (5,5))
plt.plot([0, 1], [0, 1], 'r--')
plt.plot(df_model.summary.roc.select('FPR').collect(),
df_model.summary.roc.select('TPR').collect())
plt.xlabel('FPR')
plt.ylabel('TPR')
plt.show()
def printAUC(predictions, labelCol = "churn"):
""" Print the area under curve for the predictions.
INPUT:
predictions: The predictions to get and print the AUC for
OUTPU:
Prints the AUC
"""
print("Area under curve: ", BinaryClassificationEvaluator(labelCol = labelCol).evaluate(predictions))
# -
predictions = logisticRegressionPredictions(df_train, df_test)
showEvaluationMetrics(predictions)
# +
# Evaluation results:
# The confusion matrix looks pretty good but a bit worrying is the high count of false negatives (Type 2 errors).
# This is possibly due to our label set being unbalanced.
# F1 score is looking good, I can be happy with over 80% at first try.
# But as expected, the high false negative count lead to a bad recall score.
# In the case of churn, a false negative is not a desirable prediction outcome, because this means we miss customers
# who actually churn and do not get a chance to change their minds. Which is the point of the excersize.
# So we will try to tune this and also watch the overall score.
# But before, let's plot the ROC and show AUC:
model = LogisticRegression(labelCol = "churn").fit(df_train)
model.evaluate(df_test).predictions
plotROC(model)
printAUC(predictions)
# +
# This looks like a lot of RO curves I have seen so that and also high AUC seems promising as well.
# Lookig the the curve though, we see there is some room for improvement.
# -
# ### Optimization: Threshold
#
# Since our input data is skewed on the label, we can try to undersample negative cases by lowering the threshold.
# Hopefully, this will help significantly with our bad recall rate.
predictions = logisticRegressionPredictions(df_train, df_test, 0.4)
showEvaluationMetrics(predictions)
# We see that our f1 score has slightly improved. Recall increased by quite a bit but precision also went down.
# Let's try one more time with an only slighty smaller threshold:
predictions = logisticRegressionPredictions(df_train, df_test, 0.45)
showEvaluationMetrics(predictions)
# +
# Recall went down and precision up.
# So there is a tradeoff here. Maybe if we change the input data itself?
# -
# ### Optimization: Undersampling negatives in the input data
#
# As an alternative to penalize false negatives, we can try to undersample them.
# Check distributions of churn
zeros = df_subset.filter(df["churn"] == 0)
ones = df_subset.filter(df["churn"] == 1)
zerosCount = zeros.count()
onesCount = ones.count()
print("Ones: {}, Zeros: {}".format(onesCount, zerosCount))
print(onesCount / zerosCount * 100)
# +
# As a "quick and dirty" check, we will just hack off some 0's and see what happens.
# Note: Normally we would use something like KStratified sampling or similar but that is beyond the scode of this project.
def undersampleNegatives(df, ratio, labelCol = "churn"):
"""
Undersample the negatives (0's) in the given dataframe by ratio.
NOTE: The "selection" method here is of course very crude and in a real version should be randomized and shuffled.
INPUT:
df: dataframe to undersample negatives from
ratio: Undersampling ratio
labelCol: LAbel column name in the input dataframe
OUTPUT:
A new dataframe with negatives undersampled by ratio
"""
zeros = df.filter(df[labelCol] == 0)
ones = df.filter(df[labelCol] == 1)
zeros = createSubset(zeros, ratio)
return zeros.union(ones)
df_undersampled = undersampleNegatives(df_subset, .8)
# -
# Check distribution again
zeros = df_undersampled.filter(df["churn"] == 0)
ones = df_undersampled.filter(df["churn"] == 1)
zerosCount = zeros.count()
onesCount = ones.count()
print("Ones: {}, Zeros: {}".format(onesCount, zerosCount))
print(onesCount / zerosCount * 100)
df_train, df_test = df_undersampled.randomSplit([0.91, 0.09])
predictions = logisticRegressionPredictions(df_train, df_test)
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# As we can see, our recall score did go up a little, but at the cost of precision.
# Same as when we modified te threshold.
# The f1 score stayed about the same, but the AUC went up a little.
#
# So while we do have some small optimization, it will be a business decision to decide which way to
# tune the model - either more precision or more recall.
# Another thing we can try is to modify our input data in the ETL stage.
# -
# ### RandomForest
#
# Logistic regression performed reasonably well but there was a very low recall rate.
# Let's see how a RandomForesClassifier does on this dataset.
# Since we have some high correlation, an ensemble learning model
# like RFC might do a little better.
# +
def randomForestPredictions(df_train, df_test, numTrees = 50, labelCol = "churn", featuresCol = "features"):
""" Fit, evaluate and show results for RandomForestClassifier
INPUT:
df_train: The training data set.
df_test: The testing data set.
numTrees: Number of trees in the forest.
labelCol: The label column name, "churn" by default.
featuresCol: The label column name, "features" by default.
OUTPUT:
predictions: The model's predictions.
"""
# Fit and train model
rfc = RandomForestClassifier(labelCol = labelCol, featuresCol = featuresCol, numTrees = numTrees).fit(df_train)
return rfc.transform(df_test)
predictions = randomForestPredictions(df_train, df_test)
# -
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# While there is a very low recall rate, the Precision is perfect (This could be due to our reduced dataset size),
# and the AUC is pretty high as well.
# Let's try another optimization.
# -
# ### Optimization: numTrees
#
# We will try to increase the number of decision trees in the forest.
predictions = randomForestPredictions(df_train, df_test, 100)
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# Not a whole lot has improved.
# Lastly we try to run the RF on the undersampled input data.
# Since it severly penalized the recall, we will remove a larger portion of negatives this time.
# -
# ### Optimization: Undersampled negatives
#
# Same as above with LogisticRegression.
# ratio was derived by trial and error
df_undersampled = undersampleNegatives(df_subset, .264)
df_train, df_test = df_undersampled.randomSplit([0.9, 0.1])
predictions = randomForestPredictions(df_train, df_test, 100)
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# It seems that with severely reduced negatives in the data, the RandomForest classifier was able to converge better.
# The f1 score has worsened, but now recall is on a higher level, meaning we would not miss
# as many customers churning as before.
# But will these good results from the undersampled data hold for the real test data? Let's see:
rfc = RandomForestClassifier(labelCol = "churn", featuresCol = "features", numTrees = 100).fit(df_train)
df_train, df_test = df_subset.randomSplit([0.9, 0.1])
predictions = rfc.transform(df_test)
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# Again we see that there is a tradeoff to be expected between recall and precision.
# But the f1 score is good, as is precision, and now recall is looking better, so it's an improvement.
# Anyway this concludes the preliminary experiments for RandomForest tuning, in conclusion I like the model better as it seems
# to perform better and also gives more leeway when tuning as compared to LogisticRegression.
# -
# ### Gradient Boost
#
# As last algorithm we try a gradient boosted tree classifier.
# +
def gbtPredictions(df_train, df_test, maxIter = 10, labelCol = "churn", featuresCol = "features"):
""" Fit, evaluate and show results for GBTClassifier
INPUT:
df_train: The training data set.
df_test: The testing data set.
maxIter: Number of maximum iterations in the gradeint boost.
labelCol: The label column name, "churn" by default.
featuresCol: The label column name, "features" by default.
OUTPUT:
predictions: The model's predictions
"""
# Fit and train model
gbt = GBTClassifier(labelCol = labelCol, featuresCol = featuresCol, maxIter = maxIter).fit(df_train)
return gbt.transform(df_test)
predictions = gbtPredictions(df_train, df_test)
# -
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# The GBT results look very promising. There are very high rates right off the bat, and recall is nearing 50%.
# -
# ### Optimization: Unersampling negatives
#
# With our experience from above, let's try undersampling as first optimization.
# What happens here is largeley analogous to similar procedures above.
df_undersampled = undersampleNegatives(df_subset, .6)
df_train, df_test = df_undersampled.randomSplit([0.9, 0.1])
predictions = gbtPredictions(df_train, df_test)
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# This is a significant improvement. There is a strongly improved recall rate at a high precision.
# Let's see how it does on the normal sampled test set:
gbt = GBTClassifier(labelCol = "churn", featuresCol = "features", maxIter = 10).fit(df_train)
df_train, df_test = df_subset.randomSplit([0.9, 0.1])
predictions = gbt.transform(df_test)
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# The model was able to translate it's performance very well to the normal sampled dataset.
# I am convinced that this is the best algorithm to use.
# -
# ### Full undersampled dataset
#
# Let us run and evalute the algorithm on the full (undersampled) dataset.
# +
df_train, df_test = df.randomSplit([0.9, 0.1])
df_undersampled = undersampleNegatives(df_train, .6)
gbt = GBTClassifier(labelCol = "churn", featuresCol = "features", maxIter = 10).fit(df_undersampled)
predictions = gbt.transform(df_test)
showEvaluationMetrics(predictions)
printAUC(predictions)
# +
# The f1 score is very good, in fact all scores but recall are very good.
# But recall is still very acceptable under these conditions.
# Now we can use automated optimization to search for the best GBT model.
# -
# Output the notebook to an html file
from subprocess import call
call(['python', '-m', 'nbconvert', 'model.ipynb'])
|
udacity/data-scientist-nanodegree/sparkify/model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: visualization-curriculum-gF8wUgMm
# language: python
# name: visualization-curriculum-gf8wugmm
# ---
# + [markdown] papermill={"duration": 0.010032, "end_time": "2020-03-28T16:40:23.793109", "exception": false, "start_time": "2020-03-28T16:40:23.783077", "status": "completed"} tags=[]
# # COVID-19 Tracking U.S. Cases
# > Tracking coronavirus total cases, deaths and new cases in US by states.
#
# - comments: true
# - author: <NAME>
# - categories: [overview, interactive, usa]
# - hide: true
# - permalink: /covid-overview-us/
# + papermill={"duration": 0.018147, "end_time": "2020-03-28T16:40:23.817848", "exception": false, "start_time": "2020-03-28T16:40:23.799701", "status": "completed"} tags=[]
#hide
print('''
Example of using jupyter notebook, pandas (data transformations), jinja2 (html, visual)
to create visual dashboards with fastpages
You see also the live version on https://gramener.com/enumter/covid19/united-states.html
''')
# + papermill={"duration": 0.356603, "end_time": "2020-03-28T16:40:24.180685", "exception": false, "start_time": "2020-03-28T16:40:23.824082", "status": "completed"} tags=[]
#hide
import numpy as np
import pandas as pd
from jinja2 import Template
from IPython.display import HTML
# + papermill={"duration": 0.013482, "end_time": "2020-03-28T16:40:24.200306", "exception": false, "start_time": "2020-03-28T16:40:24.186824", "status": "completed"} tags=[]
#hide
from pathlib import Path
if not Path('covid_overview.py').exists():
# ! wget https://raw.githubusercontent.com/pratapvardhan/notebooks/master/covid19/covid_overview.py
# + papermill={"duration": 0.045632, "end_time": "2020-03-28T16:40:24.252035", "exception": false, "start_time": "2020-03-28T16:40:24.206403", "status": "completed"} tags=[]
#hide
import covid_overview as covid
# + papermill={"duration": 0.070888, "end_time": "2020-03-28T16:40:24.330257", "exception": false, "start_time": "2020-03-28T16:40:24.259369", "status": "completed"} tags=[]
#hide
COL_REGION = 'Province/State'
kpis_info = [
{'title': 'New York', 'prefix': 'NY'},
{'title': 'Washington', 'prefix': 'WA'},
{'title': 'California', 'prefix': 'CA'}]
data = covid.gen_data_us(region=COL_REGION, kpis_info=kpis_info)
# + papermill={"duration": 0.037772, "end_time": "2020-03-28T16:40:24.375120", "exception": false, "start_time": "2020-03-28T16:40:24.337348", "status": "completed"} tags=[]
#hide
data['table'].head(5)
# + papermill={"duration": 0.135637, "end_time": "2020-03-28T16:40:24.517739", "exception": false, "start_time": "2020-03-28T16:40:24.382102", "status": "completed"} tags=[]
#hide_input
template = Template(covid.get_template(covid.paths['overview']))
html = template.render(
D=data['summary'], table=data['table'],
newcases=data['newcases'].iloc[:, -15:],
COL_REGION=COL_REGION,
KPI_CASE='US',
KPIS_INFO=kpis_info,
LEGEND_DOMAIN=[5, 50, 500, np.inf],
np=np, pd=pd, enumerate=enumerate)
HTML(f'<div>{html}</div>')
# + [markdown] papermill={"duration": 0.009551, "end_time": "2020-03-28T16:40:24.538412", "exception": false, "start_time": "2020-03-28T16:40:24.528861", "status": "completed"} tags=[]
# Visualizations by [<NAME>](https://twitter.com/PratapVardhan)[^1]
#
# [^1]: Source: ["The New York Times"](https://github.com/nytimes/covid-19-data). Link to [notebook](https://github.com/pratapvardhan/notebooks/blob/master/covid19/covid19-overview-us.ipynb), [orignal interactive](https://gramener.com/enumter/covid19/united-states.html)
|
_notebooks/2020-03-21-covid19-overview-us.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: scMyositis
# language: python
# name: scmyositis
# ---
# # Get amino acids from TraCeR assemble
# This notebook looks for TraCeR assemble output and builds a dataframe with cell data.<br>
# It assumes the following data folder structure:
# ```bash
# ├── AB
# │ ├── <batch_1>
# │ │ ├── <cell_1a>
# │ │ │ ├── filtered_TCR_seqs
# │ │ │ │ ├── filtered_TCRs.txt
# │ │ │ │ └── ...
# │ │ │ └── ...
# │ │ ├── <cell_1b>
# │ │ │ ├── filtered_TCR_seqs
# │ │ │ │ ├── filtered_TCRs.txt
# │ │ │ │ └── ...
# │ │ │ └── ...
# │ │ └── ...
# │ ├── <batch_2>
# │ │ ├── <cell_2a>
# │ │ │ ├── filtered_TCR_seqs
# │ │ │ │ ├── filtered_TCRs.txt
# │ │ │ │ └── ...
# │ │ │ └── ...
# │ │ ├── <cell_2b>
# │ │ │ ├── filtered_TCR_seqs
# │ │ │ │ ├── filtered_TCRs.txt
# │ │ │ │ └── ...
# │ │ │ └── ...
# │ │ └── ...
# │ └── ...
# ├── GD
# │ ├── <batch_1>
# │ │ ├── <cell_1a>
# │ │ │ ├── filtered_TCR_seqs
# │ │ │ │ ├── filtered_TCRs.txt
# │ │ │ │ └── ...
# │ │ │ └── ...
# │ │ ├── <cell_1b>
# │ │ │ ├── filtered_TCR_seqs
# │ │ │ │ ├── filtered_TCRs.txt
# │ │ │ │ └── ...
# │ │ │ └── ...
# │ │ └── ...
# │ ├── <batch_2>
# │ │ ├── <cell_2a>
# │ │ │ ├── filtered_TCR_seqs
# │ │ │ │ ├── filtered_TCRs.txt
# │ │ │ │ └── ...
# │ │ │ └── ...
# │ │ ├── <cell_2b>
# │ │ │ ├── filtered_TCR_seqs
# │ │ │ │ ├── filtered_TCRs.txt
# │ │ │ │ └── ...
# │ │ │ └── ...
# │ │ └── ...
# │ └── ...
# ```
# **Author: <NAME>**<br>
# 22/02/2021<br>
# Kernel: `scMyocitis`<br>
import os
import argparse
import pandas as pd
from objects import Cell, Chain
from objects import AlphaChain, BetaChain, GammaChain, DeltaChain
from objects import create_cell_from_AB, append_GD_data
pd.set_option('display.max_columns',None)
in_path = './data'
out_file = 'results/results_tracer.csv'
AB_path = os.path.join(in_path,os.path.normpath('AB'))
GD_path = os.path.join(in_path,os.path.normpath('GD'))
# ### Looping over AB cell files
cells = {}
for batch in os.listdir(AB_path):
path = os.path.join(AB_path,batch)
cont = 1
L = len(os.listdir(path))
for folder in os.listdir(path):
file = os.path.join(path,folder,os.path.normpath('filtered_TCR_seqs/filtered_TCRs.txt'))
cell = create_cell_from_AB(file)
print("Cell {}, {}/{} in batch {}".format(cell.name,cont,L,batch))
cell.add_batch(batch)
cells[cell.name] = cell
cont = cont + 1
# ### Looping over GD files
for batch in os.listdir(GD_path):
path = os.path.join(GD_path,batch)
cont = 1
L = len(os.listdir(path))
for folder in os.listdir(path):
file = os.path.join(path,folder,os.path.normpath('filtered_TCR_seqs/filtered_TCRs.txt'))
print("Cell {}/{} in batch {}".format(cont,L,batch))
append_GD_data(file,cells)
cont = cont +1
# # Dataframe generation
DF = pd.DataFrame(index=cells.keys())
cont = 1
L = len(cells)
for name,cell in cells.items():
print("Cell {}, {}/{}".format(name,cont,L))
DF.loc[name,'seq_batch'] = cell.batch
chains = cell.A_chains + cell.B_chains + cell.G_chains + cell.D_chains
for chain in chains:
meta_colnames = [chain.allele + '_' + s for s in list(chain.__dict__.keys())][2:]
values = list(chain.__dict__.values())[2:]
DF.loc[name,meta_colnames] = values
cont = cont +1
DF
# # Export data
DF.to_csv(out_file,sep=',')
|
collect_assemble/get_aa.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from json import loads
import numpy as np
import xarray as xr
import zarr
from dask.distributed import Client
from fsspec.implementations.http import HTTPFileSystem
from xarray.testing import assert_chunks_equal, assert_equal, assert_identical
import xpublish
# -
# Let's check to make sure our server started alright
# !head logfile.txt
# Start a dask cluster for use on the client side
client = Client(n_workers=4, dashboard_address=43757)
client
# You can now open three more browser tabs/windows:
#
# _Note that you will have to modify the url prefix slightly, to do this, just copy the first part of your browser's URL_
#
# 1. Xpublish Web App: e.g. https://hub.gke.mybinder.org/user/jhamman-xpublish-gbbqbxfi/proxy/9000
# 2. Xpublish's Dask Cluster Dashboard: e.g. https://hub.gke.mybinder.org/user/jhamman-xpublish-gbbqbxfi/proxy/8787/status
# 3. This Notebook's Dask Cluster Dashboard: e.g. https://hub.gke.mybinder.org/user/jhamman-xpublish-gbbqbxfi/proxy/43757/status
#
# _Also note that this port numbers may change. The server side ports are available in `logfile.txt` (see above) and the client-side port is in the cell above._
# +
# We can access our API using fsspec's HTTPFileSystem
fs = HTTPFileSystem()
# The http mapper gives us a dict-like interface to the API
http_map = fs.get_mapper("http://0.0.0.0:9000")
# -
# We can access API enpoints by key now...
for key in [".zmetadata", "keys"]:
print(key, http_map[key], "\n")
# The .zmetadata key returns the json dictionary of consolidated zarr metadata
# We can load/decode that and access one array's attributes
d = loads(http_map[".zmetadata"])
d["metadata"]["air/.zattrs"]
# We can pass that mapper object directly to zarr's open_consolidated function
# This returns a zarr groups
zg = zarr.open_consolidated(http_map, mode="r")
zg.tree()
# And we can do the same with xarray's open_zarr function
ds = xr.open_zarr(http_map, consolidated=True)
ds
# The rest of this notebook applies some simple tests to show that the served dataset is indentical to the
# "air_temperature" dataset in xarray's tutorial dataset.
ds_tutorial = xr.tutorial.open_dataset(
"air_temperature", chunks=dict(lat=5, lon=5), decode_cf=True
)
ds_tutorial.air.attrs
def test(actual, expected, index):
"""a simple equality test with index as a parameter"""
assert np.array_equal(actual[index].values, expected[index].values)
# test a bunch of indexing patterns
for index in [
(0, 0, 0),
(slice(0, 4), 0, 0),
(slice(0, 4), slice(0, 4), 0),
(slice(0, 4), slice(0, 4), slice(0, 4)),
(slice(-4), slice(0, 4), slice(0, 4)),
(slice(None), slice(0, 4), slice(0, 4)),
(slice(None), slice(None), slice(0, 4)),
(slice(None), slice(None), slice(None)),
]:
print(index)
test(ds_tutorial["air"], ds["air"], index)
assert_equal(ds, ds_tutorial)
assert_chunks_equal(ds, ds_tutorial)
assert_identical(ds, ds_tutorial)
|
examples/open_dataset.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Computer Vision Nanodegree
#
# ## Project: Image Captioning
#
# ---
#
# In this notebook, you will train your CNN-RNN model.
#
# You are welcome and encouraged to try out many different architectures and hyperparameters when searching for a good model.
#
# This does have the potential to make the project quite messy! Before submitting your project, make sure that you clean up:
# - the code you write in this notebook. The notebook should describe how to train a single CNN-RNN architecture, corresponding to your final choice of hyperparameters. You should structure the notebook so that the reviewer can replicate your results by running the code in this notebook.
# - the output of the code cell in **Step 2**. The output should show the output obtained when training the model from scratch.
#
# This notebook **will be graded**.
#
# Feel free to use the links below to navigate the notebook:
# - [Step 1](#step1): Training Setup
# - [Step 2](#step2): Train your Model
# - [Step 3](#step3): (Optional) Validate your Model
# # <a id='step1'></a>
# ## Step 1: Training Setup
#
# In this step of the notebook, you will customize the training of your CNN-RNN model by specifying hyperparameters and setting other options that are important to the training procedure. The values you set now will be used when training your model in **Step 2** below.
#
# You should only amend blocks of code that are preceded by a `TODO` statement. **Any code blocks that are not preceded by a `TODO` statement should not be modified**.
#
# ### Task #1
#
# Begin by setting the following variables:
# - `batch_size` - the batch size of each training batch. It is the number of image-caption pairs used to amend the model weights in each training step.
# - `vocab_threshold` - the minimum word count threshold. Note that a larger threshold will result in a smaller vocabulary, whereas a smaller threshold will include rarer words and result in a larger vocabulary.
# - `vocab_from_file` - a Boolean that decides whether to load the vocabulary from file.
# - `embed_size` - the dimensionality of the image and word embeddings.
# - `hidden_size` - the number of features in the hidden state of the RNN decoder.
# - `num_epochs` - the number of epochs to train the model. We recommend that you set `num_epochs=3`, but feel free to increase or decrease this number as you wish. [This paper](https://arxiv.org/pdf/1502.03044.pdf) trained a captioning model on a single state-of-the-art GPU for 3 days, but you'll soon see that you can get reasonable results in a matter of a few hours! (_But of course, if you want your model to compete with current research, you will have to train for much longer._)
# - `save_every` - determines how often to save the model weights. We recommend that you set `save_every=1`, to save the model weights after each epoch. This way, after the `i`th epoch, the encoder and decoder weights will be saved in the `models/` folder as `encoder-i.pkl` and `decoder-i.pkl`, respectively.
# - `print_every` - determines how often to print the batch loss to the Jupyter notebook while training. Note that you **will not** observe a monotonic decrease in the loss function while training - this is perfectly fine and completely expected! You are encouraged to keep this at its default value of `100` to avoid clogging the notebook, but feel free to change it.
# - `log_file` - the name of the text file containing - for every step - how the loss and perplexity evolved during training.
#
# If you're not sure where to begin to set some of the values above, you can peruse [this paper](https://arxiv.org/pdf/1502.03044.pdf) and [this paper](https://arxiv.org/pdf/1411.4555.pdf) for useful guidance! **To avoid spending too long on this notebook**, you are encouraged to consult these suggested research papers to obtain a strong initial guess for which hyperparameters are likely to work best. Then, train a single model, and proceed to the next notebook (**3_Inference.ipynb**). If you are unhappy with your performance, you can return to this notebook to tweak the hyperparameters (and/or the architecture in **model.py**) and re-train your model.
#
# ### Question 1
#
# **Question:** Describe your CNN-RNN architecture in detail. With this architecture in mind, how did you select the values of the variables in Task 1? If you consulted a research paper detailing a successful implementation of an image captioning model, please provide the reference.
#
# **Answer:** The CNN Encoder is pre-trained ResNet-50 architecture without fully connected layer to extract the features from the images as specified in the 1_Preliminaries. The model and it's parameters have been specified with the reference from the paper(1411.4555). The paper specifies the dimensions for the embeddings and the size of the LSTM memory as 512 and tokenization based on threshold value for vocabulary as 5 as specified in 1_Preliminaries.
#
#
# ### (Optional) Task #2
#
# Note that we have provided a recommended image transform `transform_train` for pre-processing the training images, but you are welcome (and encouraged!) to modify it as you wish. When modifying this transform, keep in mind that:
# - the images in the dataset have varying heights and widths, and
# - if using a pre-trained model, you must perform the corresponding appropriate normalization.
#
# ### Question 2
#
# **Question:** How did you select the transform in `transform_train`? If you left the transform at its provided value, why do you think that it is a good choice for your CNN architecture?
#
# **Answer:** The provided transformations were used as it is since Resize, RandomCrop, ToTensor, Normalize are usually the standard transforms used. Also, RandomHorizontalFlip used provides random flipping of the images with the probability of 0.5.
#
# ### Task #3
#
# Next, you will specify a Python list containing the learnable parameters of the model. For instance, if you decide to make all weights in the decoder trainable, but only want to train the weights in the embedding layer of the encoder, then you should set `params` to something like:
# ```
# params = list(decoder.parameters()) + list(encoder.embed.parameters())
# ```
#
# ### Question 3
#
# **Question:** How did you select the trainable parameters of your architecture? Why do you think this is a good choice?
#
# **Answer:** As the encoder is pretrained we do not need to retrain the encoder model. Thus using the value suggested 'params = list(decoder.parameters()) + list(encoder.embed.parameters())' the embedding layer of CNN and decoder parameters were set.
#
# ### Task #4
#
# Finally, you will select an [optimizer](http://pytorch.org/docs/master/optim.html#torch.optim.Optimizer).
#
# ### Question 4
#
# **Question:** How did you select the optimizer used to train your model?
#
# **Answer:** Adam optimizer depends on adaptive learning and it is well suited for the problems involving large data or parameters.
# +
import torch
import torch.nn as nn
from torchvision import transforms
import sys
sys.path.append('/opt/cocoapi/PythonAPI')
from pycocotools.coco import COCO
from data_loader import get_loader
from model import EncoderCNN, DecoderRNN
import math
from workspace_utils import active_session
## TODO #1: Select appropriate values for the Python variables below.
batch_size = 128 # batch size
vocab_threshold = 5 # minimum word count threshold
vocab_from_file = True # if True, load existing vocab file
embed_size = 512 # dimensionality of image and word embeddings
hidden_size = 512 # number of features in hidden state of the RNN decoder
num_epochs = 3 # number of training epochs
save_every = 1 # determines frequency of saving model weights
print_every = 100 # determines window for printing average loss
log_file = 'training_log.txt' # name of file with saved training loss and perplexity
# (Optional) TODO #2: Amend the image transform below.
transform_train = transforms.Compose([
transforms.Resize(256), # smaller edge of image resized to 256
transforms.RandomCrop(224), # get 224x224 crop from random location
transforms.RandomHorizontalFlip(), # horizontally flip image with probability=0.5
transforms.ToTensor(), # convert the PIL Image to a tensor
transforms.Normalize((0.485, 0.456, 0.406), # normalize image for pre-trained model
(0.229, 0.224, 0.225))])
# Build data loader.
data_loader = get_loader(transform=transform_train,
mode='train',
batch_size=batch_size,
vocab_threshold=vocab_threshold,
vocab_from_file=vocab_from_file)
# The size of the vocabulary.
vocab_size = len(data_loader.dataset.vocab)
# Initialize the encoder and decoder.
encoder = EncoderCNN(embed_size)
decoder = DecoderRNN(embed_size, hidden_size, vocab_size)
# Move models to GPU if CUDA is available.
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
encoder.to(device)
decoder.to(device)
# Define the loss function.
criterion = nn.CrossEntropyLoss().cuda() if torch.cuda.is_available() else nn.CrossEntropyLoss()
# TODO #3: Specify the learnable parameters of the model.
params = list(encoder.embed.parameters())+list(decoder.parameters())
# TODO #4: Define the optimizer.
optimizer = torch.optim.Adam(params)
# Set the total number of training steps per epoch.
total_step = math.ceil(len(data_loader.dataset.caption_lengths) / data_loader.batch_sampler.batch_size)
# -
# <a id='step2'></a>
# ## Step 2: Train your Model
#
# Once you have executed the code cell in **Step 1**, the training procedure below should run without issue.
#
# It is completely fine to leave the code cell below as-is without modifications to train your model. However, if you would like to modify the code used to train the model below, you must ensure that your changes are easily parsed by your reviewer. In other words, make sure to provide appropriate comments to describe how your code works!
#
# You may find it useful to load saved weights to resume training. In that case, note the names of the files containing the encoder and decoder weights that you'd like to load (`encoder_file` and `decoder_file`). Then you can load the weights by using the lines below:
#
# ```python
# # Load pre-trained weights before resuming training.
# encoder.load_state_dict(torch.load(os.path.join('./models', encoder_file)))
# decoder.load_state_dict(torch.load(os.path.join('./models', decoder_file)))
# ```
#
# While trying out parameters, make sure to take extensive notes and record the settings that you used in your various training runs. In particular, you don't want to encounter a situation where you've trained a model for several hours but can't remember what settings you used :).
#
# ### A Note on Tuning Hyperparameters
#
# To figure out how well your model is doing, you can look at how the training loss and perplexity evolve during training - and for the purposes of this project, you are encouraged to amend the hyperparameters based on this information.
#
# However, this will not tell you if your model is overfitting to the training data, and, unfortunately, overfitting is a problem that is commonly encountered when training image captioning models.
#
# For this project, you need not worry about overfitting. **This project does not have strict requirements regarding the performance of your model**, and you just need to demonstrate that your model has learned **_something_** when you generate captions on the test data. For now, we strongly encourage you to train your model for the suggested 3 epochs without worrying about performance; then, you should immediately transition to the next notebook in the sequence (**3_Inference.ipynb**) to see how your model performs on the test data. If your model needs to be changed, you can come back to this notebook, amend hyperparameters (if necessary), and re-train the model.
#
# That said, if you would like to go above and beyond in this project, you can read about some approaches to minimizing overfitting in section 4.3.1 of [this paper](http://ieeexplore.ieee.org/stamp/stamp.jsp?arnumber=7505636). In the next (optional) step of this notebook, we provide some guidance for assessing the performance on the validation dataset.
# +
import torch.utils.data as data
import numpy as np
import os
import requests
import time
with active_session():
# Open the training log file.
f = open(log_file, 'w')
old_time = time.time()
response = requests.request("GET",
"http://metadata.google.internal/computeMetadata/v1/instance/attributes/keep_alive_token",
headers={"Metadata-Flavor":"Google"})
for epoch in range(1, num_epochs+1):
for i_step in range(1, total_step+1):
if time.time() - old_time > 60:
old_time = time.time()
requests.request("POST",
"https://nebula.udacity.com/api/v1/remote/keep-alive",
headers={'Authorization': "STAR " + response.text})
# Randomly sample a caption length, and sample indices with that length.
indices = data_loader.dataset.get_train_indices()
# Create and assign a batch sampler to retrieve a batch with the sampled indices.
new_sampler = data.sampler.SubsetRandomSampler(indices=indices)
data_loader.batch_sampler.sampler = new_sampler
# Obtain the batch.
images, captions = next(iter(data_loader))
# Move batch of images and captions to GPU if CUDA is available.
images = images.to(device)
captions = captions.to(device)
# Zero the gradients.
decoder.zero_grad()
encoder.zero_grad()
# Pass the inputs through the CNN-RNN model.
features = encoder(images)
outputs = decoder(features, captions)
# Calculate the batch loss.
loss = criterion(outputs.view(-1, vocab_size), captions.view(-1))
# Backward pass.
loss.backward()
# Update the parameters in the optimizer.
optimizer.step()
# Get training statistics.
stats = 'Epoch [%d/%d], Step [%d/%d], Loss: %.4f, Perplexity: %5.4f' % (epoch, num_epochs, i_step, total_step, loss.item(), np.exp(loss.item()))
# Print training statistics (on same line).
print('\r' + stats, end="")
sys.stdout.flush()
# Print training statistics to file.
f.write(stats + '\n')
f.flush()
# Print training statistics (on different line).
if i_step % print_every == 0:
print('\r' + stats)
# Save the weights.
if epoch % save_every == 0:
torch.save(decoder.state_dict(), os.path.join('./models', 'decoder-%d.pkl' % epoch))
torch.save(encoder.state_dict(), os.path.join('./models', 'encoder-%d.pkl' % epoch))
# Close the training log file.
f.close()
# -
# <a id='step3'></a>
# ## Step 3: (Optional) Validate your Model
#
# To assess potential overfitting, one approach is to assess performance on a validation set. If you decide to do this **optional** task, you are required to first complete all of the steps in the next notebook in the sequence (**3_Inference.ipynb**); as part of that notebook, you will write and test code (specifically, the `sample` method in the `DecoderRNN` class) that uses your RNN decoder to generate captions. That code will prove incredibly useful here.
#
# If you decide to validate your model, please do not edit the data loader in **data_loader.py**. Instead, create a new file named **data_loader_val.py** containing the code for obtaining the data loader for the validation data. You can access:
# - the validation images at filepath `'/opt/cocoapi/images/train2014/'`, and
# - the validation image caption annotation file at filepath `'/opt/cocoapi/annotations/captions_val2014.json'`.
#
# The suggested approach to validating your model involves creating a json file such as [this one](https://github.com/cocodataset/cocoapi/blob/master/results/captions_val2014_fakecap_results.json) containing your model's predicted captions for the validation images. Then, you can write your own script or use one that you [find online](https://github.com/tylin/coco-caption) to calculate the BLEU score of your model. You can read more about the BLEU score, along with other evaluation metrics (such as TEOR and Cider) in section 4.1 of [this paper](https://arxiv.org/pdf/1411.4555.pdf). For more information about how to use the annotation file, check out the [website](http://cocodataset.org/#download) for the COCO dataset.
# +
# (Optional) TODO: Validate your model.
|
2_Training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#Week 1 Assignment : Data Visualization Project - <NAME>
#Goal: analyzing the relationship between fertility and life expectancy worldwide
#Step 1: Load, inspect and format data
#Step 2: Static Analysis with Boxplots for the time period between 2000 and 2015
#Step 3: Dynamic Analysis with Animation (variable evolution across the years)
# -
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import imageio
# +
## Step 1: Load and format data
# Read Files
fert = pd.read_csv('/home/rita/Documents/spiced/spiced-projects/convex_capers_student_code/week01/data/gapminder_total_fertility.csv', index_col=0)
life = pd.read_excel('/home/rita/Documents/spiced/spiced-projects/convex_capers_student_code/week01/data/gapminder_lifeexpectancy.xlsx', index_col=0)
pop = pd.read_excel('/home/rita/Documents/spiced/spiced-projects/convex_capers_student_code/week01/data/gapminder_population.xlsx', index_col = 0)
cont = pd.read_csv('/home/rita/Documents/spiced/spiced-projects/convex_capers_student_code/week01/data/continents.csv', sep =';')
#Fertility Table
fert.columns = fert.columns.astype(int)
fert.index.name = 'country'
fert = fert.reset_index()
fert = fert.melt(id_vars ='country', var_name='year',value_name='fertility_rate')
#Life Expectancy Table
life.index.name = 'country'
life = life.reset_index()
life = life.melt(id_vars='country', var_name='year', value_name='life expectancy')
#Population Table
pop.columns = pop.columns.astype(int)
pop.index.name = 'country'
pop = pop.reset_index()
pop = pop.melt(id_vars ='country', var_name='year',value_name='Total_Population')
#Merge Tables
df = fert.merge(pop)
df = df.merge(life)
df = df.merge(cont)
df.head()
# +
#Step 2: Static Analysis (2000 vs 2015) with Boxplots
#Subsetting the data frame
df_00_15 = df[(df['year'] == 2000) | (df['year'] == 2015)]
# Descriptive Statistics 2000 vs 2015
round(df_00_15.groupby(['year'])[['fertility_rate', 'life expectancy']].describe(),2)
# +
# World Fertility Rate Chart 2000 vs 2015
plt.figure(figsize=(12,8))
sns.boxplot(x='continent', y="fertility_rate", hue="year", data=df_00_15, palette="Set3")
# +
# World Life Expectancy Chart 2000 vs 2015
plt.figure(figsize=(12,8))
sns.boxplot(x='continent', y="life expectancy", hue="year",data=df_00_15, palette="Set3")
# +
#Step 3: Dynamic Analysis with Animation : World Life expectancy and Fertility rate across time
#Note: this code was written based on the following tutorials:
#https://www.python-graph-gallery.com/340-scatterplot-animation
#https://www.python-graph-gallery.com/341-python-gapminder-animation
# First, create the chart for one year
df_subset = df.loc[df['year'] == 2000]
sns.set_theme(style="whitegrid")
plt.figure(figsize=(16,14))
ax = sns.scatterplot(x='life expectancy', y='fertility_rate',
size ='Total_Population',
sizes=(150,2050),
hue='continent',
#legend = False,
data=df_subset,
alpha=0.6)
plt.title('World Fertility and Life expectancy in year 2000', fontsize=24)
plt.xlabel('Life expectancy', fontsize=16)
plt.ylabel('Fertility', fontsize=16)
# Locate the legend outside of the plot
plt.legend(bbox_to_anchor=(1, 1)
, loc='upper left'
, fontsize=12
, borderpad=0.5
, labelspacing = 2
, ncol=2
, handleheight = 3
)
# +
# Second, write a loop that creates a chart for every year
df_subset = df[(df['year'] >= 2000)]
df_subset.head(5)
dpi=96
for i in df_subset['year']:
plt.ioff()
fig = plt.figure(figsize=(680/dpi, 480/dpi), dpi=dpi)
df_subset = df.loc[df['year'] == i]
sns.set_theme(style="whitegrid")
plt.figure(figsize=(12,8))
ax = sns.scatterplot(x='life expectancy', y='fertility_rate',
size ='Total_Population',
sizes=(150,2050),
hue='continent',
#legend = False,
data=df_subset,
alpha=0.6)
plt.title('Year '+str(i), fontsize=20)
plt.xlabel('Life expectancy', fontsize=16)
plt.ylabel('Fertility', fontsize=16)
# Locate the legend outside of the plot
plt.legend(bbox_to_anchor=(1 , 1), loc='upper left', fontsize=10)
#plt.show()
filename= 'lifeexp_'+str(i)+'.png'
plt.savefig(fname=filename, dpi=96)
plt.gca()
plt.close(fig)
plt.close('all')
# +
# Third, we save the pictures from the previous loop and create a gif animation
images = []
for i in df_1960_2015['year']:
filename = 'lifeexp_{}.png'.format(i)
images.append(imageio.imread(filename))
imageio.mimsave('output.gif', images, fps=20)
|
week01/week01_ Assignment_Ana_Rita_Santos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Load python libraries**
# +
# %matplotlib inline
import datetime
import matplotlib.pyplot as plt
import os.path
import xarray as xr
import numpy as np
import netCDF4
import cartopy.crs as ccrs
import cartopy.feature as cfeature
from cartopy.mpl.gridliner import LONGITUDE_FORMATTER, LATITUDE_FORMATTER
import matplotlib.ticker as mticker
from IPython.display import Image, display
from glob import glob
from modules.GOESR_functions import goes_lat_lon_reproj, get_s3_keys
from modules.plotter import loadCPT
import matplotlib
matplotlib.use("Agg")
from matplotlib import colors as c
import matplotlib.animation as animation
# +
filename_pattern = '../data/GOES17_ABI_L2_CMI13/OR_ABI*.nc'
GOES_files = glob(filename_pattern) # search files
GOES_files.sort()
DS = xr.open_mfdataset(GOES_files, concat_dim='t')
long_name = DS.CMI.long_name
lats, lons = goes_lat_lon_reproj(DS)
# put reprojected GOES HT data into new dataset
DS_goes = xr.Dataset({'CMI': (['time', 'y', 'x'], DS.CMI.values)},
coords={'time': DS.t.values,
'lat': (['y', 'x'], lats),
'lon': (['y', 'x'], lons)})
DS_goes
# -
DS_goes.time.values[-1]
# +
def drawmap(ax, x, y, VO, cmap, title):
datacrs = ccrs.PlateCarree() ## the projection the data is in
mapcrs = ccrs.PlateCarree() ## the projection you want your map displayed in
ax = fig.add_subplot(1,1,1, projection=mapcrs)
ax.set_title(title, fontsize=14)
ax.set_extent(ext, crs=mapcrs)
# Add Border Features
coast = ax.coastlines(linewidths=1.0, resolution='10m')
ax.add_feature(states_provinces, edgecolor='k')
ax.add_feature(cfeature.BORDERS)
# Add grid lines
gl = ax.gridlines(crs=datacrs, draw_labels=True,
linewidth=.5, color='black', alpha=0.5, linestyle='--')
gl.xlocator = mticker.FixedLocator(np.arange(-140., -110., 4))
gl.ylocator = mticker.FixedLocator(np.arange(24, 44, 2))
gl.xformatter = LONGITUDE_FORMATTER
gl.yformatter = LATITUDE_FORMATTER
gl.xlabels_top = False
gl.ylabels_right = False
# Add contour plot
cs = ax.pcolormesh(lons, lats, VO, transform=datacrs, cmap=cmap, vmin=-103, vmax=84, zorder=1)
return cs
def myanimate(i, ax, DS, cmap):
ax.clear()
VO = DS.CMI.values[i]- 273.15 ## Convert to Celsius
x = DS.lat
y = DS.lon
title = 'GOES-17 {0} at {1}'.format(long_name, DS_goes.time.values[i])
new_contour = drawmap(ax,x,y,VO,cmap, title)
return new_contour
# -
lats = DS_goes.lat
lons = DS_goes.lon
VO = DS_goes.CMI.values - 273.15 ## Convert to Celsius
# +
# band_info = '(Band ' + str(DS.band_id.values[0]) + ', ' \
# # + str(DS.band_wavelength.values[0]) + DS.band_wavelength.units + ')'
# title = 'GOES-17 {0} {1} on {2}'.format(DS.CMI.long_name, band_info, DS.time_coverage_end)
title = 'GOES-17 {0} at {1}'.format(long_name, DS_goes.time.values[0])
FFMpegWriter = animation.writers['ffmpeg']
metadata = dict(title=title,
comment='Movie for GOES-17 Cloud Brightness Temp starting at 4 March 2019 18 UTC')
writer = FFMpegWriter(fps=20, metadata=metadata)
datacrs = ccrs.PlateCarree() ## the projection the data is in
mapcrs = ccrs.PlateCarree() ## the projection you want your map displayed in
ext = [-115.0, -135.0, 25.0, 40.0]
# Import cmap from CPT
cmap = loadCPT('../data/Colortables/IR4AVHRR6.cpt')
# Create a feature for States/Admin 1 regions at 1:50m from Natural Earth
states_provinces = cfeature.NaturalEarthFeature(
category='cultural',
name='admin_1_states_provinces_lines',
scale='50m',
facecolor='none')
fig = plt.figure(figsize=[16,12]) # a new figure window
ax = fig.add_subplot(1,1,1, projection=mapcrs)
ax.set_extent(ext, crs=mapcrs)
# Add Border Features
coast = ax.coastlines(linewidths=1.0, resolution='10m')
ax.add_feature(states_provinces, edgecolor='k')
ax.add_feature(cfeature.BORDERS)
# Add grid lines
gl = ax.gridlines(crs=datacrs, draw_labels=True,
linewidth=.5, color='black', alpha=0.5, linestyle='--')
gl.xlocator = mticker.FixedLocator(np.arange(-140., -110., 4))
gl.ylocator = mticker.FixedLocator(np.arange(24, 44, 2))
gl.xformatter = LONGITUDE_FORMATTER
gl.yformatter = LATITUDE_FORMATTER
gl.xlabels_top = False
gl.ylabels_right = False
first_contour = ax.pcolormesh(lons, lats, VO[0], transform=datacrs, cmap=cmap, vmin=-103, vmax=84)
ax.set_title(title)
# Add a color bar
cbar = fig.colorbar(first_contour, orientation='vertical', cmap=cmap, shrink=0.7)
Unit = "Brightness Temperature ($\mathrm{\degree C}$)"
cbar.set_label(Unit, fontsize=12)
ani = animation.FuncAnimation(fig, myanimate, frames=np.arange(len(DS_goes.CMI)), fargs=(ax, DS_goes, cmap), interval=50)
ani.save("GOES17_CMI.mp4")
# -
len(DS_goes.CMI)
|
Supplemental Video.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from scipy.stats import norm
import numpy as np
df = pd.read_csv('user_info.csv')
print(df.shape)
df = df.drop('username',axis=1)
df['won'] = [0]*df.shape[0]
kd = [i/(1+sum([df.qps_elims,df.qps_deaths])) for i in [df.qps_elims,df.qps_deaths]]
df['kill_ratio'] = kd[0]
df['death_ratio'] = kd[1]
df.head(10)
##get rid of 1 row 1644 / 6 = 274
# +
def get_team():
index = [i for i in np.random.randint(0,df.shape[0],size=12)]
team1 = df.iloc[index[0:6]].mean(axis=0)
team2 = df.iloc[index[6:12]].mean(axis=0)
t1 = 0
t2 = 0
for col in df.columns:
if 'deaths' in col:
if team1[col] > team2[col]:
t1 = t1 - 1
t2 = t2 + 1
else:
t1 = t1 + 1
t2 = t2 - 1
else:
if team1[col] > team2[col]:
t1 = t1 + 1
t2 = t2 - 1
else:
t1 = t1 - 1
t2 = t2 + 1
if np.random.randint(0,100) >= 90:
t1 = t1 + 10
elif np.random.randint(0,100) <= 10:
t2 = t2 + 10
if t1 > t2:
team1['won'] = 1
team2['won'] = 0
elif t2 > t1:
team1['won'] = 0
team2['won'] = 1
else:
team1['won'] = 1
team2['won'] = 1
return pd.DataFrame([team1,team2], columns=df.columns)
team_averages = pd.DataFrame(columns=df.columns)
for i in range(1000):
team_averages = team_averages.append(get_team())
# -
team_averages.shape
# +
# df.qps_elims.plot(kind='hist', normed=True)
plt.hist(team_averages.qps_elims);
# +
from sklearn.model_selection import train_test_split
from sklearn.ensemble import RandomForestClassifier
from xgboost import XGBClassifier
from sklearn.pipeline import make_pipeline
from sklearn.metrics import accuracy_score
y_pred = [1]*len(team_averages)
accuracy_score(team_averages.won,y_pred)
# df.head()
# +
train, test = train_test_split(team_averages, train_size=0.80, test_size=0.20,
stratify=team_averages['won'], random_state=42)
target = 'won'
X_train = train.drop(columns=target)
y_train = train[target]
X_test = test.drop(columns=target)
y_test = test[target]
train.shape, test.shape
# +
sample_weight = [1]*len(X_train)
for i in range(len(sample_weight)):
if i in [5,6,7,8]:
sample_weight[i] *= 15
len(sample_weight)
# +
rfc = RandomForestClassifier(n_estimators=1000, random_state=42, n_jobs=-1,max_depth=20)
# Fit on train, score on val
rfc.fit(X_train, y_train)
rfc_y_pred = rfc.predict(X_test)
print('Accuracy Score', accuracy_score(y_test, rfc_y_pred), ' - Model Score', rfc.score(X_test,y_test))
# +
# fit model no training data
xgb = XGBClassifier(n_estimators=1000,random_state=42,max_depth=10, n_jobs=-1)
xgb.fit(X_train, y_train, sample_weight = sample_weight)
xgb_y_pred = xgb.predict(X_test)
print('Accuracy Score', accuracy_score(y_test, xgb_y_pred), ' - Model Score', xgb.score(X_test,y_test))
# +
from sklearn.neighbors import KNeighborsClassifier
knc = KNeighborsClassifier(n_neighbors=25, leaf_size=30,weights='distance')
knc.fit(X_train, y_train)
knc_y_pred = knc.predict(X_test)
print('Accuracy Score', accuracy_score(y_test, knc_y_pred), ' - Model Score', knc.score(X_test,y_test))
# +
from sklearn import svm
clf = svm.SVC(gamma=1)
clf.fit(X_train, y_train, sample_weight=sample_weight)
clf_y_pred = clf.predict(X_test)
print('Accuracy Score', accuracy_score(y_test, clf_y_pred), ' - Model Score', clf.score(X_test,y_test))
# -
# ## Assignment
#
# Complete these tasks for your project, and document your decisions.
#
# - [ ] Choose your target. Which column in your tabular dataset will you predict?
# - [ ] Choose which observations you will use to train, validate, and test your model. And which observations, if any, to exclude.
# - [ ] Determine whether your problem is regression or classification.
# - [ ] Choose your evaluation metric.
# - [ ] Begin with baselines: majority class baseline for classification, or mean baseline for regression, with your metric of choice.
# - [ ] Begin to clean and explore your data.
# - [ ] Choose which features, if any, to exclude. Would some features "leak" information from the future?
# ### Target: Win or lose rate
# ### I will train on individual players based on their past games and heroes.
|
notebooks/OW_Project.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: geo_dev
# language: python
# name: geo_dev
# ---
# # Generating spatial weights
#
# `momepy` is using `libpysal` to handle spatial weights, but also builds on top of it. This notebook will show how to use different weights.
import momepy
import geopandas as gpd
import matplotlib.pyplot as plt
# We will again use `osmnx` to get the data for our example and after preprocessing of building layer will generate tessellation layer.
# + tags=["hide_output"]
import osmnx as ox
gdf = ox.geometries.geometries_from_place('Kahla, Germany', tags={'building':True})
buildings = ox.projection.project_gdf(gdf)
buildings['uID'] = momepy.unique_id(buildings)
limit = momepy.buffered_limit(buildings)
tessellation = momepy.Tessellation(buildings, unique_id='uID', limit=limit).tessellation
# -
# ## Queen contiguity
#
# Morphological tessellation allows using contiguity-based weights matrix. While `libpysal.weights.contiguity.Queen` will do the standard Queen contiguity matrix of the first order; it might not be enough to capture proper context. For that reason, we can use `momepy.sw_high` to capture all neighbours within set topological distance `k`. It generates spatial weights of higher orders under the hood and joins them together.
sw3 = momepy.sw_high(k=3, gdf=tessellation, ids='uID')
# Queen contiguity of morphological tessellation can capture the comparable level of information across the study area - the number of the neighbour is relatively similar and depends on the morphology of urban form. We can visualize it by counting the number of neighbours (as captured by `sw3`).
tessellation['neighbours'] = momepy.Neighbors(tessellation, sw3,'uID').series
# + tags=["hide_input"]
f, ax = plt.subplots(figsize=(10, 10))
tessellation.plot(ax=ax, column='neighbours', legend=True, cmap='Spectral_r')
buildings.plot(ax=ax, color="white", alpha=0.4)
ax.set_axis_off()
plt.show()
# -
# ## Distance
#
# Often we want to define the neighbours based on metric distance. We will look at two options - distance band and k-nearest neighbour.
#
# ### Distance band
#
# We can imagine distance band as a buffer of a set radius around centroid of each object, for example, 400 meters. For that, we can use `libpysal.weights.DistanceBand`:
# + tags=["hide_output"]
import libpysal
dist400 = libpysal.weights.DistanceBand.from_dataframe(buildings, 400,
ids='uID')
# -
# Because we have defined spatial weights using uID, we can use `dist400` generated on buildings and use it on tessellation:
tessellation['neighbours400'] = momepy.Neighbors(tessellation, dist400, 'uID').series
# + tags=["hide_input"]
f, ax = plt.subplots(figsize=(10, 10))
tessellation.plot(ax=ax, column='neighbours400', legend=True, cmap='Spectral_r')
buildings.plot(ax=ax, color="white", alpha=0.4)
ax.set_axis_off()
plt.show()
# -
# ### K nearest neighbor
#
# If we want fixed number of neighbours, we can use `libpysal.weights.KNN`:
knn = libpysal.weights.KNN.from_dataframe(buildings, k=200, ids='uID')
tessellation['neighboursKNN'] = momepy.Neighbors(tessellation, knn,'uID').series
# + [markdown] tags=["popout"]
# **Note**: As all tessellation cells have the same number of neighbours (due to KNN), they all have the same colour.
# + tags=["hide_input"]
f, ax = plt.subplots(figsize=(10, 10))
tessellation.plot(ax=ax, column='neighboursKNN', legend=True, cmap='Spectral_r')
buildings.plot(ax=ax, color="white", alpha=0.4)
ax.set_axis_off()
plt.show()
# -
# All of them can be used within morphometric analysis. Theoretical and practical differences are discussed in Fleischmann, Romice and Porta (2019).
#
# For the other options on generating spatial weights see [lipysal API](https://pysal.org/libpysal/api.html).
|
docs/user_guide/weights/weights_nb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (OpenSimEnv_V2)
# language: python
# name: opensimenv_v2
# ---
# +
from __future__ import print_function
from __future__ import unicode_literals
from __future__ import division
from __future__ import absolute_import
from future import standard_library
standard_library.install_aliases() # NOQA
import argparse
import logging
import sys
#import box2d
from logging import getLogger
import chainer
from chainer import optimizers
import gym
gym.undo_logger_setup() # NOQA
from gym import spaces
import gym.wrappers
import numpy as np
from matplotlib import style
import argparse
import logging
import matplotlib.pyplot as plt
from chainer import functions as F
from chainerrl.agents.trpo import TRPO
from chainerrl.agents.acer import ACERSeparateModel
from chainerrl.optimizers.nonbias_weight_decay import NonbiasWeightDecay
from chainerrl.optimizers import rmsprop_async
from chainerrl.action_value import QuadraticActionValue #Q-function output for continuous action space
#from chainerrl.action_value import DiscreteActionValue
import chainerrl
from chainerrl import experiments
from chainerrl import explorers
from chainerrl import misc
from chainerrl import policy
from chainerrl import q_functions
from chainerrl import replay_buffer
from chainerrl import v_functions
#import osim
#from osim.env import L2RunEnv
style.use('ggplot')
# -
# environment settings
env_name='Pendulum-v0'
# Chainer's settings
parser = argparse.ArgumentParser()
#logging.basicConfig(level=args.logger_level)
seed=0
gpu=-1
# +
number_of_episodes=10000
max_episode_length=200
replay_start_size=5000
number_of_update_times=1
target_update_interval=1
target_update_method='soft'
soft_update_tau=1e-2
update_interval=4
number_of_eval_runs=100
eval_interval=10 ** 5
final_exploration_steps=10 ** 6
gamma=0.995
minibatch_size=128
# +
# Helper's functions
def clip_action_filter(a):
return np.clip(a, action_space.low, action_space.high)
def reward_filter(r):
return r
def phi(obs):
return obs.astype(np.float32)
def random_action():
a = action_space.sample()
if isinstance(a, np.ndarray):
a = a.astype(np.float32)
return a
def make_env(test,env_name,render=True):
env = gym.make(env_name)
# Use different random seeds for train and test envs
env_seed = 2 ** 32 - 1 - seed if test else seed
env.seed(env_seed)
#if args.monitor:
#env = gym.wrappers.Monitor(env, args.outdir)
if isinstance(env.action_space, spaces.Box):
misc.env_modifiers.make_action_filtered(env, clip_action_filter)
if not test:
misc.env_modifiers.make_reward_filtered(env, reward_filter)
if render and not test:
misc.env_modifiers.make_rendered(env)
return env
# -
# Set a random seed used in ChainerRL
misc.set_random_seed(seed)
# +
#env = gym.make(env_name)
env = make_env(test=False,env_name=env_name,render=False)
#timestep_limit = env.spec.tags.get('wrapper_config.TimeLimit.max_episode_steps')
#timestep_limit=300
obs_size = np.asarray(env.observation_space.shape).prod()
action_space = env.action_space
action_size = np.asarray(action_space.shape).prod()
# +
""""
q_func = q_functions.FCSAQFunction(
obs_size,
action_size,
n_hidden_channels=critic_hidden_units,
n_hidden_layers=critic_hidden_layers)
"""
obs_space = env.observation_space
v_fuc = chainerrl.v_functions.FCVFunction(
obs_space.low.size,
n_hidden_channels=64,
n_hidden_layers=3,
last_wscale=0.01,
nonlinearity=F.tanh,
)
v_fuc_opt = chainer.optimizers.Adam()
v_fuc_opt.setup(v_fuc)
pi = chainerrl.policies.FCGaussianPolicyWithStateIndependentCovariance(
obs_space.low.size,
action_space.low.size,
n_hidden_channels=64,
n_hidden_layers=3,
mean_wscale=0.01,
nonlinearity=F.tanh,
var_type='diagonal',
var_func=lambda x: F.exp(2 * x), # Parameterize log std
var_param_init=0, # log std = 0 => std = 1
)
# +
#model
#rbuf = replay_buffer.ReplayBuffer(5 * 10 ** 5)
rbuf = replay_buffer.ReplayBuffer(5 * 10)
ou_sigma = (action_space.high - action_space.low) * 0.2
explorer = explorers.AdditiveOU(sigma=ou_sigma)
obs_normalizer = chainerrl.links.EmpiricalNormalization(obs_space.low.size)
#vf_opt = chainer.optimizers.Adam()
#vf_opt.setup(vf)
# -
# The agent
agent = TRPO(
policy=pi,
vf=v_fuc,
vf_optimizer = v_fuc_opt,
obs_normalizer=obs_normalizer,
phi=lambda x: np.array(x).astype(np.float32, copy=False),
update_interval=update_interval,
conjugate_gradient_max_iter=20,
conjugate_gradient_damping=1e-3,
gamma=gamma,
lambd=0.97,
vf_epochs=5,
entropy_coef=0,
)
# +
G=[]
G_mean=[]
for ep in range(1, number_of_episodes+ 1):
if ep%100:
agent.save("TRPO_Pendulum_last_one_10000")
obs = env.reset()
reward = 0
done = False
R = 0 # return (sum of rewards)
t = 0 # time step
episode_rewards=[]
while not done and t < max_episode_length:
# Uncomment to watch the behaviour
#env.render()
action = agent.act_and_train(obs, reward)
obs, reward, done, _ = env.step(action)
R += reward
episode_rewards.append(reward)
t += 1
if done or t >= max_episode_length :
# Calculate sum of the rewards
episode_rewards_sum = sum(episode_rewards)
G.append(episode_rewards_sum)
total_G = np.sum(G)
maximumReturn = np.amax(G)
print("%f" % (episode_rewards_sum), file=open("TRPO_Pendulum_last_one_reward_10000.txt", "a"))
#print("%i" % (episode_rewards_sum))
if ep % 10 == 0:
print("==========================================")
print("Episode: ", ep)
print("Rewards: ", episode_rewards_sum)
print("Max reward so far: ", maximumReturn)
# Mean reward
total_reward_mean = np.divide(total_G, ep+1)
G_mean.append(total_reward_mean)
print("Mean Reward", total_reward_mean)
print("%f" % (total_reward_mean), file=open("TRPO_Pendulum_last_one_MEAN_Reward_10000.txt", "a"))
agent.stop_episode_and_train(obs, reward, done)
print('Finished.')
plt.xlabel('episdes')
plt.ylabel('reword')
plt.plot(G)
plt.savefig('trpo_Pendulum_last_one_10000.png',dpi=1000)
plt.plot(G_mean)
plt.ylabel('Average of Returns')
plt.xlabel('Number of episodes/10')
plt.savefig("ReturnsAverage_VS_Episodes_trpo_Pendulum_last_one_10000",dpi=1000)
# -
plt.plot(G_mean)
plt.ylabel('Average of Returns')
plt.xlabel('Number of episodes/10')
plt.title('ReturnsAverage_VS_Episodes trpo_prosthetic_edited_parameters_1')
plt.savefig("ReturnsAverage_VS_Episodes trpo_prosthetic_edited_parameters_1")
plt.xlabel('episdes')
plt.ylabel('reword')
plt.title('trpo_prosthetic_edited_parameters_1')
plt.plot(G)
plt.savefig('trpo_prosthetic_edited_parameters_1.png',dpi=1000)
# +
plt.xlabel('episdes')
plt.ylabel('reword')
plt.title('trpo_prosthetic_edited_parameters_1')
plt.plot(G)
plt.savefig('trpo_prosthetic_edited_parameters_1.png')
# -
plt.plot(G_mean)
plt.ylabel('Average of Returns trpo_Walker2D')
plt.xlabel('Number of episodes/10')
plt.title('ReturnsAverage_VS_Episodes trpo_Walker2D')
plt.savefig("ReturnsAverage_VS_Episodes trpo_Walker2D_2_4000",dpi=1000)
|
Pendulum/TRPO_Pendulum.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# <a href='http://www.pieriandata.com'><img src='../Pierian_Data_Logo.png'/></a>
# ___
# <center><em>Copyright by Pierian Data Inc.</em></center>
# <center><em>For more information, visit us at <a href='http://www.pieriandata.com'>www.pieriandata.com</a></em></center>
# # Linear Regression Project Exercise - Solutions
# Now that we have learned about feature engineering, cross validation, and grid search, let's test all your new skills with a project exercise in Machine Learning. This exercise will have a more guided approach, later on the ML projects will begin to be more open-ended. We'll start off with using the final version of the Ames Housing dataset we worked on through the feature engineering section of the course. Your goal will be to create a Linear Regression Model, train it on the data with the optimal parameters using a grid search, and then evaluate the model's capabilities on a test set.
# ---
# ---
# ---
# ## Complete the tasks in bold
#
# **TASK: Run the cells under the Imports and Data section to make sure you have imported the correct general libraries as well as the correct datasets. Later on you may need to run further imports from scikit-learn.**
#
# ### Imports
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# ### Data
df = pd.read_csv("../DATA/AMES_Final_DF.csv")
df.head()
df.info()
# **TASK: The label we are trying to predict is the SalePrice column. Separate out the data into X features and y labels**
X = df.drop('SalePrice',axis=1)
y = df['SalePrice']
# **TASK: Use scikit-learn to split up X and y into a training set and test set. Since we will later be using a Grid Search strategy, set your test proportion to 10%. To get the same data split as the solutions notebook, you can specify random_state = 101**
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.10, random_state=101)
# **TASK: The dataset features has a variety of scales and units. For optimal regression performance, scale the X features. Take carefuly note of what to use for .fit() vs what to use for .transform()**
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaled_X_train = scaler.fit_transform(X_train)
scaled_X_test = scaler.transform(X_test)
# **TASK: We will use an Elastic Net model. Create an instance of default ElasticNet model with scikit-learn**
from sklearn.linear_model import ElasticNet
base_elastic_model = ElasticNet()
# **TASK: The Elastic Net model has two main parameters, alpha and the L1 ratio. Create a dictionary parameter grid of values for the ElasticNet. Feel free to play around with these values, keep in mind, you may not match up exactly with the solution choices**
param_grid = {'alpha':[0.1,1,5,10,50,100],
'l1_ratio':[.1, .5, .7, .9, .95, .99, 1]}
# **TASK: Using scikit-learn create a GridSearchCV object and run a grid search for the best parameters for your model based on your scaled training data. [In case you are curious about the warnings you may recieve for certain parameter combinations](https://stackoverflow.com/questions/20681864/lasso-on-sklearn-does-not-converge)**
from sklearn.model_selection import GridSearchCV
# verbose number a personal preference
grid_model = GridSearchCV(estimator=base_elastic_model,
param_grid=param_grid,
scoring='neg_mean_squared_error',
cv=5,
verbose=1)
grid_model.fit(scaled_X_train,y_train)
# **TASK: Display the best combination of parameters for your model**
grid_model.best_params_
# **TASK: Evaluate your model's performance on the unseen 10% scaled test set. In the solutions notebook we achieved an MAE of $\$$14149 and a RMSE of $\$$20532**
y_pred = grid_model.predict(scaled_X_test)
from sklearn.metrics import mean_absolute_error,mean_squared_error
mean_absolute_error(y_test,y_pred)
np.sqrt(mean_squared_error(y_test,y_pred))
np.mean(df['SalePrice'])
# ## Great work!
#
# ----
|
Data Science Resources/Jose portila - ML/10-Cross-Val-and-LinReg-Project/.ipynb_checkpoints/03-Linear-Regression-Project-Exercise-Solutions-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 64-bit (''qc'': conda)'
# name: python3
# ---
# +
import numpy as np
# Importing standard Qiskit libraries
from qiskit import *
from qiskit.visualization import *
from qiskit.providers.aer import QasmSimulator
from qiskit.providers.ibmq import *
from qiskit.tools.monitor import job_monitor
from qiskit.ignis.mitigation.measurement import (complete_meas_cal,CompleteMeasFitter)
IBMQ.load_account()
provider = IBMQ.get_provider(hub='ibm-q')
sim = Aer.get_backend('qasm_simulator')
# -
def oracleS(s):
n = len(s)
qc = QuantumCircuit(2*n)
for i in range(n):
if s[i] == '1':
for j in range(n):
if s[j] == '1':
qc.cx(i,n+j)
return qc
def simons(s):
n = len(s)
h = QuantumCircuit(2*n,2*n)
h.h(range(n))
h.barrier()
qc = h.compose(oracleS(s))
qc.barrier()
r = range(n,2*n)
qc.measure(r,r)
qc.barrier()
qc.compose(h,inplace=True)
r = range(0,n)
qc.measure(r,r)
return qc
s = "11"
n = len(s)
qc = simons(s)
qc.draw('mpl')
backend = least_busy(provider.backends(filters=lambda x: x.configuration().n_qubits >= n
and not x.configuration().simulator
and x.status().operational==True and x.name()!='ibmq_bogota'))
print("least busy backend: ", backend)
m = execute(qc,backend=sim,shots=1024).result()
print(m)
plot_histogram(
m.get_counts()
#execute(qc,backend=sim,shots=1024).result().get_counts()
)
job = execute(qc,backend=backend,shots=1024)
job_monitor(job)
plot_histogram(
job.result().get_counts()
)
meas_fitter = CompleteMeasFitter(cal_results, state_label)
meas_fitter.plot_calibration()
mit_res = meas_fitter.filter.apply(results)
|
simons.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # I/O handling and settings #
#
# The raredecay package contains simple functions to save all the output produced during the run and even provides the simple possibility to add your own output.
#
# To be able to save the output, and also to tweak a few other settings like the logger verbosity, we have to call *initialize()* right at the beginning before any import statements occur.
#
# If we ommit this step and go right to our desired functions, a standard *initialize()* will be called automatically with no output-path defined.
#
# ### Why do we want output saved to file? ###
#
# Not every information should be displayed during a run in the console and not every figure should be plotted. But if the output_path is specified, every information (more then displayed) and nearly every plot (some issues there, but most of the plots work) gets saved to file anyway. So you still have the possibility to lookup things later on which you though may are not of interest.
# And not to forget: if you run it as a job, it is often the easiest way to save the output directly instead of redericting the prints to the console.
from raredecay import settings
settings.initialize(output_path='tmp_output', # any valid directory, best use absolute path.
#If None -> no output saved
run_name="My first run", # Names the folder
overwrite_existing=False, # False by default. Adds a number if folder already exists.
# If True, overwrite existing folder.
run_message="This is my first run, just checking out the settings module",
# appears at the beginning of the saved output file
verbosity=3, # Default value. between 1 and 5. The higher, the more the code will tell you
plot_verbosity=3, # Default value. same as verbosity but for showing plots
prompt_for_input=True, # if True, the script asks for an addition to the run-name
# as well as before *show()*
logger_console_level='warning', # Default value. What kind of logger messages you want to see
#in the console
logger_file_level='debug', # Default value. Values:'debug', 'info', 'warning', 'error', 'critical'
# The "higher" (more serious) the level, the less will be logged (from the level specified and up)
n_cpu=1, # Default value. The estimated number of cores/threads to use. Can be changed later
gpu_in_use=False # Default value, advanced. Can be changed later.
#Only use if gpu-parallelization is in use
)
# #### Prompt_for_input ####
# As prompt_for_input was set to True, the script asked for a name extension. It is useful to enter your "instant thoughts about the run", like in the example: "n_estimators 20 with L2 0.1" to remember why you did this run. It will be added to the file name, as we will see later on.
# ## Writing your script ##
# So far, everything is done for the initialization and all the functions from the package can be imported. To make sure that the output will be saved, we have to call *finalize()* at the very end.
#
#
# ### Changing settings during the script ###
# It is possible, for some settings, to be changed during the script (and therefore between diferent function calls).
settings.set_verbosity(verbosity=3, # Default value
plot_verbosity=3 # Default value
)
settings.parallel_profile(n_cpu=-1, # Default value. Will choose all available cpus.
# -2 takes all but one, -3 all but 2 and so on...
gpu_in_use=False # Default value.
)
settings.figure_save_config(file_formats=['png', 'svg'], #Default value.
# The formats to save the figure to. See the docs for available formats
to_pickle=True, # Default value. Pickles the matplotlib-figure for possible re-plot
dpi=150 # Default value. The image resolution
)
# ### Adding additional output ###
# During a script, we may want to add output by ourselves; comments, figures, dictionaries, data...
# The easiest way to do that is to get the output-handler of the package.
out = settings.get_output_handler()
# To add any kind of output, we just have to call *add_output()* and give all the output inside of an iterable.
out.add_output(["Hi, I am ", 42, "without", {'me': 42, 'he': 41}])
# It concatenates the objects and creates a nice representation (e.g. for dicts).
# We can specify several options:
out.add_output(["Hi, I am ", 42, "without", {'me': 42, 'he': 41}],
to_end=False, # Default value. If True, the output will be written at the very end.
# This should be done with the important results in order to see them all at the end.
importance=3, # Default value. The higher, the more likely it will be **printed** (it is saved anyway).
# Whether something is printed/plotted depends on the importances as well as on the verbosity
# There is a trade-off between verbosity and importance
title="We can set a big title here",
subtitle="Or (also) a subtitle",
section="This is the smallest title", # We do not need to have all three kinds of title at once;)
obj_separator=" ", # Default value. The separation between the printed objects.
data_separator="\n\n", # Default value. The strings to separate from the next *add_output()*
force_newline=False # Default value. If True, this output will be written on a newline, no matter
# how the last output ended and not be concatenated on the same line as the last *add_output()*
)
# Beside objects off all kind, we can also save figures. This does the same as plt.figure() but saves it in addition (at the end of the run, so you can plot into the same figure several times)
# +
my_figure = out.save_fig("My first figure")
# as an example:
import matplotlib.pyplot as plt
plt.hist([1,4, 5, 4, 2, 4, 2])
# -
# Again, we can specify a couple of parameters:
my_figure2 = out.save_fig("My second figure",
importance=3, # same as for *add_output()*
file_format=None, # Default value. If None, will take the default ones.
to_pickle=True, # Default value. If True, pickle the figure for a possible re-plot
)
# as an example, we may also plot something
plt.hist([1,1,1,1,2,2,3,4,5,5,5])
# ### Advanced feature: rederict I/O-stream ###
# You may encounter a package, that prints its output directly to the console, but you want to save it to a file too.
# Therefore you can use the *IO_to_string()* method, which redericts the output, and *IO_to_sys()* to have normal output again.
out.IO_to_string()
print "You cannot see that... yet"
# The output is currently collected by the output-handler.
out.IO_to_sys(importance=3, # Default value. Whether to plot the collected output or not
# We can add several keyword-arguments, which will be given directly to *add_output()*
# Example:
section="Example of IO_to_sys and back")
# ## Finalize the run ##
# At the end of every run, we should (and have to in case we specified an output_path) finalize it.
# As we choose *prompt_for_input* in the *initialize()* function, we will be asked to press enter (an empty input) before the plots will be showed. This way we won't be suprised by popping up windows.
settings.finalize(show_plots=True, # Default value. The *lazy* way for plt.show() ;)
play_sound_at_end=False # Default value. If True, a beep will sound at the end
#to let you know it's at the end.
)
# The function *finalize()* returns all the output nicely formatted (as seen on the screen above the plots) and can be used for whatever.
|
howto/IO_handling_and_settings.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# * Vectorization is converting iterative statements into a vector based operation.
# * Vectorization in numpy is implemented by ufuncs -"Universal Functions".
# * vectorization is way faster than iterating over elements.
# * they ALso provide broadcasting and additional methods like reduce, accumulate , etc. that are very helpful for computation.
#
# ### Add the Elements of Two Lists
# iterate over both of the lists and then sum each elements
# +
a = [1, 2, 3, 4]
b = [4, 5, 6, 7]
z = []
for i, j in zip(a,b):
z.append(i+j)
print(z)
# -
# #### Simple Arithmatic functions
# * add()
# * substract()
# * multiply()
# * divide()
# * power()
# * mod() / remainder()
# * divmod()
# * absolute() / abs()
# *
|
Numpy Vectorization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pre-trained weight usage
#
# A minimum example of how to load and use the CEM500K pre-trained weights for classification or segmentation tasks.
#
# Before getting started download CEM500K data and models from EMPIAR:
# - EMPIAR entry: https://www.ebi.ac.uk/pdbe/emdb/empiar/entry/10592/
# - Download help: https://www.ebi.ac.uk/pdbe/emdb/empiar/faq#question_Download
#
import os
# ## PyTorch
# First, let's consider a simple binary classification model.
import torch
import torch.nn as nn
import segmentation_models_pytorch as smp
from copy import deepcopy
from torchvision.models import resnet50
path_to_empiar_download = '' #fill this in
state_path = os.path.join(path_to_empiar_download, 'pretrained_models/cem500k_mocov2_resnet50_200ep_pth.tar')
state = torch.load(state_path, map_location='cpu')
#take a look at what's inside the state
print(list(state.keys()))
# - Epoch: the training epoch when state was recorded
# - Arch: the model architecture: "resnet50"
# - State_dict: state dict for the complete pretrained model (both query and key encoders)
# - Optimizer: state of the optimizer at save (useful for resuming training)
# - Norms: the mean and std pixel values used during training
state_dict = state['state_dict']
#format the parameter names to match torchvision resnet50
resnet50_state_dict = deepcopy(state_dict)
for k in list(resnet50_state_dict.keys()):
#only keep query encoder parameters; discard the fc projection head
if k.startswith('module.encoder_q') and not k.startswith('module.encoder_q.fc'):
resnet50_state_dict[k[len("module.encoder_q."):]] = resnet50_state_dict[k]
#delete renamed or unused k
del resnet50_state_dict[k]
# +
#create model and load the pretrained weights
model = resnet50()
#overwrite the first conv layer to accept single channel grayscale image
#overwrite the fc layer for binary classification
model.conv1 = nn.Conv2d(1, 64, kernel_size=7, stride=2, padding=3, bias=False)
model.fc = nn.Linear(2048, 1, bias=True)
#loads all parameters but those for the fc head
#those parameters need to be trained
model.load_state_dict(resnet50_state_dict, strict=False)
# -
# Now let's load the parameters into a simple binary segmentation UNet.
#as before we need to update parameter names to match the UNet model
#for segmentation_models_pytorch we simply and the prefix "encoder."
#format the parameter names to match torchvision resnet50
unet_state_dict = deepcopy(resnet50_state_dict)
for k in list(unet_state_dict.keys()):
unet_state_dict['encoder.' + k] = unet_state_dict[k]
del unet_state_dict[k]
model = smp.Unet('resnet50', in_channels=1, encoder_weights=None, classes=1)
#all encoder parameters are loaded
#parameters in the decoder must be trained on task data
model.load_state_dict(unet_state_dict, strict=False)
# The segmentation_models_pytorch module comes with a selection of state-of-the-art semantic segmentation models. The weight loading procedure is the same for all of these architectures. For a full list, see https://github.com/qubvel/segmentation_models.pytorch#models.
|
notebooks/pretrained_weights.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Best Model i.e LSTM on Hist. of pixels (16 bin) is used to make predictions.
#
# Success and Failure Cases are shown.
#
# Visualization of Training and Test data both ground truth and predictions on google maps is also shown.
#
# Please replace plot.api_key with your own key.
# +
import math
from pandas import DataFrame
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
from numpy import array
from keras.layers import Convolution2D, MaxPooling2D, Flatten, Reshape,Conv2D
from keras.models import Sequential
from keras.layers.wrappers import TimeDistributed
from keras.utils import np_utils
import numpy as np
import cv2
from keras.preprocessing.image import img_to_array
import matplotlib.pyplot as plt
# %matplotlib inline
import pandas
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten,LSTM
from keras.optimizers import Adam
from keras.layers.normalization import BatchNormalization
from keras.utils import np_utils
from keras.layers import Conv2D, MaxPooling2D, ZeroPadding2D, GlobalAveragePooling2D
from keras.preprocessing.image import ImageDataGenerator
from keras import models
from numpy import array
from keras import backend as K
from sklearn.metrics import mean_absolute_error
from keras import optimizers
from keras.layers import Bidirectional
# +
results_rmse = []
results_mae = []
results_std = []
import numpy
# Model 1 : LSTM with 16 bins hist of pixel vals
num_bins=16
# -
# # Model 1 : LSTM with 16 bins hist of pixel vals
# +
model = Sequential()
trainX=[]
trainY=[]
testX=[]
testY=[]
numpy.random.seed(3)
time_steps=19
# load the dataset
dataframe = pandas.read_csv('./Trainold.csv')
dataset = dataframe.values
scaler = MinMaxScaler(feature_range=(0, 1))
# we group by day so we can process a video at a time.
grouped = dataframe.groupby(dataframe.VidName)
per_vid = []
for _, group in grouped:
per_vid.append(group)
print(len(per_vid))
# generate sequences a vid at a time
for i,vid in enumerate(per_vid):
histValuesList=[]
scoreList=[]
# if we have less than 20 datapoints for a vid we skip over the
# vid assuming something is missing in the raw data
total = vid.iloc[:,4:20].values
vidImPath=vid.iloc[:,0:2].values
if len(total) < time_steps :
continue
scoreVal=vid["Score"].values[0] + 1
max_total_for_vid = scoreVal.tolist()
#max_total_for_vid = vid["Score"].values[0].tolist()
for i in range(0,time_steps):
videoName=vidImPath[i][0]
imgName=vidImPath[i][1]
path="./IMAGES/Train/"+videoName+"/"+imgName
image = cv2.imread(path,0)
hist = cv2.calcHist([image],[0],None,[num_bins],[0,256])
hist_arr = hist.flatten()
#img_arr = img_to_array(image)
histValuesList.append(hist_arr)
#scoreList.append(max_total_for_vid)
trainX.append(histValuesList)
trainY.append([max_total_for_vid])
#trainY.append(scoreList)
print(len(trainX[0]))
#trainX = np.array([np.array(xi) for xi in trainX])
trainX=numpy.array(trainX)
trainY=numpy.array(trainY)
print(trainX.shape,trainY.shape)
vid_names_maps=[]
time_steps=19
# load the dataset
dataframe = pandas.read_csv('./Test.csv')
dataset = dataframe.values
#print(dataset)
# we group by day so we can process a video at a time.
grouped = dataframe.groupby(dataframe.VidName)
per_vid = []
for _, group in grouped:
per_vid.append(group)
print(len(per_vid))
# generate sequences a vid at a time
for i,vid in enumerate(per_vid):
histValuesList=[]
scoreList=[]
# if we have less than 20 datapoints for a vid we skip over the
# vid assuming something is missing in the raw data
total = vid.iloc[:,4:20].values
vidImPath=vid.iloc[:,0:2].values
if len(total)<time_steps :
vid_names_maps.append()
continue
scoreVal=vid["Score"].values[0] + 1
max_total_for_vid = scoreVal.tolist()
#max_total_for_vid = vid["Score"].values[0].tolist()
for i in range(0,time_steps):
#histValuesList.append(total[i])
#print("Vid and Img name")
#print(req[i][0],req[i][1])
videoName=vidImPath[i][0]
imgName=vidImPath[i][1]
path="./IMAGES/Test/"+videoName+"/"+imgName
image = cv2.imread(path,0)
hist = cv2.calcHist([image],[0],None,[num_bins],[0,256])
hist_arr = hist.flatten()
histValuesList.append(hist_arr)
#scoreList.append(max_total_for_vid)
testX.append(histValuesList)
testY.append([max_total_for_vid])
#testY.append(scoreList)
print(len(testX[0]))
#trainX = np.array([np.array(xi) for xi in trainX])
testX=numpy.array(testX)
testY=numpy.array(testY)
print(testX.shape,testY.shape)
trainX=numpy.array(trainX)
trainX=trainX.reshape(-1,num_bins)
trainX=trainX.reshape(-1,19,num_bins)
print(numpy.max(trainX))
testX=numpy.array(testX)
testX=testX.reshape(-1,num_bins)
testX=testX.reshape(-1,19,num_bins)
print(numpy.max(testX))
trainX=numpy.array(trainX)
trainX=trainX.reshape(-1,num_bins)
trainX = trainX/numpy.max(trainX)
trainX=trainX.reshape(-1,19,num_bins)
print(trainX.shape,trainY.shape)
testX=numpy.array(testX)
testX=testX.reshape(-1,num_bins)
testX = testX/numpy.max(testX)
testX=testX.reshape(-1,19,num_bins)
print(testX.shape,testY.shape)
print(trainX.shape,trainY.shape)
print(testX.shape,testY.shape)
#print(valX.shape,valY.shape)
adam1 = optimizers.Adam(lr=0.001)
sgd1 = optimizers.SGD(lr=0.005) #0.005 or 6,100 neurons (1.24,1.12 with 0.003 and 0.2 )
print('Build model...')
# Build Model
#model = Sequential()
model.add(LSTM(100, input_shape=(19, num_bins))) #100
model.add(Dense(1))
model.add(Dropout(0.1))
model.compile(loss='mse', optimizer=sgd1, metrics=['mse'])
#model.compile(loss='mse', optimizer=sgd1,metrics=['mean_squared_error'])
history =model.fit(trainX, trainY, nb_epoch=500, batch_size=20, verbose=2,shuffle=True) #500 batch =2
# make predictions
trainPredict = model.predict(trainX)
trainScore = mean_squared_error(trainY, trainPredict)
print('Train Score: %.2f MSE' % (trainScore))
from keras import backend as K
def root_mean_squared_error(y_true, y_pred):
return K.sqrt(K.mean(K.square(y_pred - y_true), axis=-1))
pred=model.predict(testX)
print(pred.shape)
print(testY.shape)
# calculate root mean squared error
#trainScore = math.sqrt(mean_squared_error(trainY[0], trainPredict[:,0]))
#print('Train Score: %.2f RMSE' % (trainScore))
testScore = mean_squared_error(testY, pred)
print('Test Score: %.2f MSE' % (testScore))
#maeScore = root_mean_squared_error(testY, pred)
#print('RMSE Score: %.2f MAE' % (maeScore))
rmse = np.sqrt(((pred - testY) ** 2).mean(axis=0))
print('RMSE Score: %.2f rmse' % (rmse))
mae = mean_absolute_error(testY, pred)
print('MAE Score: %.2f mae' % (mae))
list1=[]
list2=[]
diff=[]
for i in range(0,len(pred)):
print(testY[i],pred[i])
list1.append(testY[i])
list2.append(pred[i])
diff.append(abs(testY[i]-pred[i]))
print(numpy.mean(diff))
stdVals=numpy.std(diff)
results_rmse.append(rmse)
results_mae.append(mae)
#stdVals = np.std(testY-pred)
print(stdVals)
results_std.append(stdVals)
# -
# ## Predictions on training data
# +
# make predictions on training data
trainPredict = model.predict(trainX)
trainScore = mean_squared_error(trainY, trainPredict)
print('Train Score: %.2f MSE' % (trainScore))
rmse = np.sqrt(((trainPredict - trainY) ** 2).mean(axis=0))
print('RMSE Score: %.2f rmse' % (rmse))
mae = mean_absolute_error(trainY, trainPredict)
print('MAE Score: %.2f mae' % (mae))
list1=[]
list2=[]
diff=[]
for i in range(0,len(trainPredict)):
print(trainY[i],trainPredict[i])
list1.append(trainY[i])
list2.append(trainPredict[i])
diff.append(abs(trainY[i]-trainPredict[i]))
print(numpy.mean(diff))
stdVals=numpy.std(diff)
results_rmse.append(rmse)
results_mae.append(mae)
#stdVals = np.std(testY-pred)
print(stdVals)
results_std.append(stdVals)
# -
# trainY
list1
# trainPredict
list2
# ## Predictions on test data
# +
pred=model.predict(testX)
print(pred.shape)
print(testY.shape)
testScore = mean_squared_error(testY, pred)
print('Test Score: %.2f MSE' % (testScore))
rmse = np.sqrt(((pred - testY) ** 2).mean(axis=0))
print('RMSE Score: %.2f rmse' % (rmse))
mae = mean_absolute_error(testY, pred)
print('MAE Score: %.2f mae' % (mae))
list1=[]
list2=[]
diff=[]
for i in range(0,len(pred)):
print(testY[i],pred[i])
list1.append(testY[i])
list2.append(pred[i])
diff.append(abs(testY[i]-pred[i]))
print(numpy.mean(diff))
stdVals=numpy.std(diff)
results_rmse.append(rmse)
results_mae.append(mae)
#stdVals = np.std(testY-pred)
print(stdVals)
results_std.append(stdVals)
# -
# testY
list1
# pred
list2
# abs(testY-pred)
diff
# ## Finding the index of success and failure videos
success=[]
failure=[]
for i in range(0,len(diff)):
if diff[i]<0.8:
success.append(i)
else:
failure.append(i)
success
failure
print(len(success))
print(len(failure))
# # Identifying success videos
# +
testX=[]
testY=[]
goodVideos=[]
numpy.random.seed(3)
time_steps=19
# load the dataset
dataframe = pandas.read_csv('./Test.csv')
dataset = dataframe.values
#print(dataset)
# we group by day so we can process a video at a time.
grouped = dataframe.groupby(dataframe.VidName)
per_vid = []
for _, group in grouped:
per_vid.append(group)
print(len(per_vid))
# generate sequences a vid at a time
for i,vid in enumerate(per_vid):
histValuesList=[]
scoreList=[]
# if we have less than 20 datapoints for a vid we skip over the
# vid assuming something is missing in the raw data
total = vid.iloc[:,4:20].values
vidImPath=vid.iloc[:,0:2].values
if len(total)<time_steps :
print(i)
continue
scoreVal=vid["Score"].values[0] + 1
max_total_for_vid = scoreVal.tolist()
goodVideos.append(vid["VidName"].values[0])
#max_total_for_vid = vid["Score"].values[0].tolist()
for i in range(0,time_steps):
#histValuesList.append(total[i])
#print("Vid and Img name")
#print(req[i][0],req[i][1])
videoName=vidImPath[i][0]
imgName=vidImPath[i][1]
path="./IMAGES/Test/"+videoName+"/"+imgName
image = cv2.imread(path,0)
hist = cv2.calcHist([image],[0],None,[num_bins],[0,256])
hist_arr = hist.flatten()
histValuesList.append(hist_arr)
#scoreList.append(max_total_for_vid)
testX.append(histValuesList)
testY.append([max_total_for_vid])
#testY.append(scoreList)
print(len(testX[0]))
#trainX = np.array([np.array(xi) for xi in trainX])
testX=numpy.array(testX)
testY=numpy.array(testY)
print(testX.shape,testY.shape)
testX=numpy.array(testX)
testX=testX.reshape(-1,num_bins)
testX = testX/numpy.max(testX)
testX=testX.reshape(-1,19,num_bins)
print(testX.shape,testY.shape)
print(testX.shape,testY.shape)
# -
# Videos considered after preprocessing
goodVideos
print(len(goodVideos))
# +
sucX=[]
sucName=[]
failX=[]
failName=[]
sucY=[]
failY=[]
for i in range(0,len(testX)):
if i in success:
sucX.append(testX[i])
sucY.append(testY[i])
sucName.append(goodVideos[i])
elif i in failure:
failX.append(testX[i])
failY.append(testY[i])
failName.append(goodVideos[i])
print(len(sucX),len(failX))
print(len(sucY),len(failY))
# +
print(len(sucX[0]))
#trainX = np.array([np.array(xi) for xi in trainX])
sucX=numpy.array(sucX)
sucY=numpy.array(sucY)
print(sucX.shape,sucY.shape)
sucX=numpy.array(sucX)
sucX=sucX.reshape(-1,num_bins)
sucX = sucX/numpy.max(sucX)
sucX=sucX.reshape(-1,19,num_bins)
print(sucX.shape,sucY.shape)
print(sucX.shape,sucY.shape)
# -
# # Successful cases (VideoName,GroundTruth,Predictions)
# +
pred=model.predict(sucX)
print(pred.shape)
print(sucX.shape)
for i in range(0,len(pred)):
print(sucName[i],sucY[i],pred[i])
# -
# # Identifying failure videos
# +
print(len(failX[0]))
#trainX = np.array([np.array(xi) for xi in trainX])
failX=numpy.array(failX)
failY=numpy.array(failY)
print(failX.shape,failY.shape)
failX=numpy.array(failX)
failX=failX.reshape(-1,num_bins)
failX = failX/numpy.max(failX)
failX=failX.reshape(-1,19,num_bins)
print(failX.shape,failY.shape)
print(failX.shape,failY.shape)
# -
# # Failure cases (VideoName,GroundTruth,Predictions)
# +
pred=model.predict(failX)
print(pred.shape)
print(failX.shape)
for i in range(0,len(pred)):
print(failName[i],failY[i],pred[i])
# -
# # Preparing data for maps
# ## Preparing training data
# +
import os
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import misc
import matplotlib.image as mpimg
import csv
import sys
from collections import defaultdict
import sys
import re
import time
import random
import cv2
from pylab import imread, imshow, gray, mean
gray()
# column names same as in csv file
col_names = ['VidName','ImgName','Lati','Longi']
np.random.seed(7)
time_steps=19
# load the dataset
# path of the video file csv with lat and longi score values
video_scores_path = "./Trainold.csv"
dataframe = pd.read_csv(video_scores_path)
dataset = dataframe.values
# we group by day so we can process a video at a time.
grouped = dataframe.groupby(dataframe.VidName)
per_vid = []
for _, group in grouped:
per_vid.append(group)
print(len(per_vid))
trainX=[]
trainY=[]
# generate sequences a vid at a time
for i,vid in enumerate(per_vid):
histValuesList=[]
# if we have less than 20 datapoints for a vid we skip over the
# vid assuming something is missing in the raw data
total = vid[col_names].values
if len(total) < time_steps :
continue
scoreVal=vid["Score"].values[0] + 1
max_total_for_vid = scoreVal.tolist()
#max_total_for_vid = vid["Score"].values[0].tolist()
histValuesList.append(total[1])
trainX.append(histValuesList)
#trainX.append(total[0:time_steps+1])
trainY.append([max_total_for_vid])
print(len(trainX[0]))
#trainX = np.array([np.array(xi) for xi in trainX])
trainX=np.array(trainX)
trainY=np.array(trainY)
print(trainX.shape,trainY.shape)
# +
prediction=[[0,0.8657829],[0,0.87993103],[1,1.3973285],[4,1.5053711],[3,1.2717153],[3,1.379099],[2,1.639708],[2,1.663112],[2,1.5343771],[1,1.4163165],[2,1.1997833],[1,1.0789253],[4,1.2343893],[1,0.9072609],[4,1.2738577],[3,1.1652279],[0,0.67907596],[0,0.790267],[0,0.9799684],[2,1.4247246],[4,1.3236446],[1,1.075547],[1,0.9333824],[1,1.370743],[1,1.5718254],[1,1.6793518],[1,0.88987446],[2,1.4067743],[3,1.6134661],[1,1.448578],[1,0.74892867],[0,0.6371925],[3,1.5682616],[3,1.563446],[4,1.6133012],[3,1.9407429],[2,1.9993695],[2,1.8576354],[2,1.0853736],[2,1.5776322],[1,1.2538174],[1,1.2942755],[2,1.3595243],[2,1.3074241],[2,1.049606],[1,0.74621785],[0,0.4598062],[1,0.9922364],[0,0.6146336],[0,0.36708295],[1,0.66955113],[1,1.0811235],[1,1.4658868],[2,1.6368428],[2,1.5438807],[1,1.7842182],[2,1.5640081],[2,1.1591837],[3,1.6423346],[3,1.9664043],[3,1.7660748],[2,1.1584783],[3,0.9957206],[1,0.88221407],[1,1.4206612],[2,1.1734943],[2,1.3031573],[2,1.2647648],[1,0.6225684],[2,0.9201188],[1,0.8307609],[2,1.2336228],[2,1.3905258],[2,1.1744916],[0,1.4268484],[1,1.4361352],[2,1.1651131],[3,0.8364122],[2,0.62921],[1,1.1430522],[1,0.97430265],[1,0.7059832],[1,0.8587964],[1,0.8164649],[1,0.70572674],[1,0.8964597],[1,1.0888579],[0,1.0548699],[2,0.99313796],[0,0.70063215],[1,1.3648108],[2,1.1785933],[2,1.0186093],[3,1.1419646],[1,1.0615672],[1,1.0875626],[1,0.9385246],[1,1.2282915],[1,1.3046808],[0,1.159186],[1,1.2628542],[0,1.0115659],[1,1.6899377],[1,1.5836076],[3,1.0448731],[2,0.74293053],[2,0.7788062],[1,1.1646601],[0,0.8948417],[2,1.330603],[1,1.1767646],[1,1.1695198],[2,1.0223768],[0,0.73476326]]
truth=[]
pred=[]
for item in zip(trainX,prediction):
#print(item[0][0][2],item[1][0])
truthTemp=[]
predTemp=[]
truthTemp.append(item[0][0][2])
truthTemp.append(item[0][0][3])
truthTemp.append(item[1][0])
predTemp.append(item[0][0][2])
predTemp.append(item[0][0][3])
predTemp.append(item[1][1])
truth.append(truthTemp)
pred.append(predTemp)
truth
print("pred")
truth
# +
# visualizing ground truth first
tskip=[]
tnew=[]
tslworn=[]
tworn=[]
theavy=[]
for elem in truth:
if(elem[0]==0 or elem[1]==0):
continue
if(elem[2]==0):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
tskip.append(tempL)
elif(elem[2]==1):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
tnew.append(tempL)
elif(elem[2]==2):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
tslworn.append(tempL)
elif(elem[2]==3):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
tworn.append(tempL)
elif(elem[2]==4):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
theavy.append(tempL)
# +
print(tskip)
lat0=[]
long0=[]
for item in tskip:
print(item[0],item[1])
lat0.append(item[0])
long0.append(item[0])
lat0=[]
long0=[]
for item in tskip:
lat0.append(item[0])
long0.append(item[1])
print(lat0)
print(long0)
# -
# # Visualizing Training data ground truth on google maps.
#
# #### Please replace plot.api_key with your own key
# +
# ADDING ONE AT A TIME
# ['#440154', '#404387', '#29788E', '#22A784', '#79D151', '#FDE724']
from bokeh.io import output_file, output_notebook, show
from bokeh.models import (
GMapPlot, GMapOptions, ColumnDataSource, Circle, LogColorMapper, BasicTicker, ColorBar,
DataRange1d, PanTool, WheelZoomTool, BoxSelectTool
)
from bokeh.models.mappers import ColorMapper, LinearColorMapper
from bokeh.palettes import Viridis5
from bokeh.io import output_file, show
from bokeh.models import (
GMapPlot, GMapOptions, ColumnDataSource, Circle, Range1d, PanTool, WheelZoomTool, BoxSelectTool
)
map_options = GMapOptions(lat=37.5324, lng=-121.9687, map_type="roadmap", zoom=11)
plot = GMapPlot(x_range=Range1d(), y_range=Range1d(), map_options=map_options)
plot.title.text = "San Francisco Bay Area"
# For GMaps to function, Google requires you obtain and enable an API key:
#
# https://developers.google.com/maps/documentation/javascript/get-api-key
#
# Replace the value below with your personal API key:
plot.api_key = "key"
lat0=[]
long0=[]
for item in tskip:
lat0.append(item[0])
long0.append(item[1])
source = ColumnDataSource(
data=dict(
lat=lat0,
lon=long0,
)
)
circle = Circle(x="lon", y="lat", size=6, fill_color="black", fill_alpha=1.8, line_color=None)
plot.add_glyph(source, circle)
#output_notebook()
#plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
#show(plot)
# *****************************************************************************************************
# # FOR NEW ROAD
lat1=[]
long1=[]
for item in tnew:
lat1.append(item[0])
long1.append(item[1])
source1 = ColumnDataSource(
data=dict(
lat1=lat1,
lon1=long1,
)
)
circle1 = Circle(x="lon1", y="lat1", size=16, fill_color="green", fill_alpha=1.8, line_color=None)
plot.add_glyph(source1, circle1)
#output_notebook()
#plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
#show(plot)
# *****************************************************************************************************
# # FOR SLIGHTLY WORN ROAD
lat2=[]
long2=[]
for item in tslworn:
lat2.append(item[0])
long2.append(item[1])
source2 = ColumnDataSource(
data=dict(
lat2=lat2,
lon2=long2,
)
)
circle2 = Circle(x="lon2", y="lat2", size=13, fill_color="yellow", fill_alpha=1.8, line_color=None)
plot.add_glyph(source2, circle2)
#output_notebook()
#plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
#show(plot)
# *****************************************************************************************************
# # FOR WORN ROAD
lat3=[]
long3=[]
for item in tworn:
lat3.append(item[0])
long3.append(item[1])
source3 = ColumnDataSource(
data=dict(
lat3=lat3,
lon3=long3,
)
)
circle3 = Circle(x="lon3", y="lat3", size=10, fill_color="purple", fill_alpha=1.8, line_color=None)
plot.add_glyph(source3, circle3)
#output_notebook()
#plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
#show(plot)
# *****************************************************************************************************
# # FOR HEAVY ROAD
lat4=[]
long4=[]
for item in theavy:
lat4.append(item[0])
long4.append(item[1])
source4 = ColumnDataSource(
data=dict(
lat4=lat4,
lon4=long4,
)
)
circle4 = Circle(x="lon4", y="lat4", size=8, fill_color="red", fill_alpha=1.8, line_color=None)
plot.add_glyph(source4, circle4)
output_notebook()
plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
show(plot)
# -
pred
# +
from bokeh.io import output_file, output_notebook, show
from bokeh.models import (
GMapPlot, GMapOptions, ColumnDataSource, Circle, LogColorMapper, BasicTicker, ColorBar,
DataRange1d, PanTool, WheelZoomTool, BoxSelectTool
)
from bokeh.models.mappers import ColorMapper, LinearColorMapper
from bokeh.palettes import Viridis5
from bokeh.io import output_file, show
from bokeh.models import (
GMapPlot, GMapOptions, ColumnDataSource, Circle, Range1d, PanTool, WheelZoomTool, BoxSelectTool
)
# visualizing predictions
pskip=[]
pnew=[]
pslworn=[]
pworn=[]
pheavy=[]
for elem in pred:
if(elem[0]==0 or elem[1]==0):
continue
if(elem[2]>=0 and elem[2]<=0.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pskip.append(tempL)
elif(elem[2]>0.5 and elem[2]<=1.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pnew.append(tempL)
elif(elem[2]>1.5 and elem[2]<=2.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pslworn.append(tempL)
elif(elem[2]>2.5 and elem[2]<=3.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pworn.append(tempL)
elif(elem[2]>3.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pheavy.append(tempL)
# -
# # Visualizing Training data predictions on google maps.
#
# #### Please replace plot.api_key with your own key
# +
# predictions plot_pred
# ADDING ONE AT A TIME
# ['#440154', '#404387', '#29788E', '#22A784', '#79D151', '#FDE724']
map_options = GMapOptions(lat=37.5324, lng=-121.9687, map_type="roadmap", zoom=11)
plot_pred = GMapPlot(x_range=Range1d(), y_range=Range1d(), map_options=map_options)
plot_pred.title.text = "San Francisco Bay Area"
# For GMaps to function, Google requires you obtain and enable an API key:
#
# https://developers.google.com/maps/documentation/javascript/get-api-key
#
# Replace the value below with your personal API key:
plot_pred.api_key = "key"
lat0=[]
long0=[]
for item in pskip:
lat0.append(item[0])
long0.append(item[1])
source = ColumnDataSource(
data=dict(
lat=lat0,
lon=long0,
)
)
circle = Circle(x="lon", y="lat", size=6, fill_color="black", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source, circle)
#output_notebook()
#plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
#show(plot_pred)
# *****************************************************************************************************
# # FOR NEW ROAD
lat1=[]
long1=[]
for item in pnew:
lat1.append(item[0])
long1.append(item[1])
source1 = ColumnDataSource(
data=dict(
lat1=lat1,
lon1=long1,
)
)
circle1 = Circle(x="lon1", y="lat1", size=16, fill_color="green", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source1, circle1)
#output_notebook()
#plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
#show(plot_pred)
# *****************************************************************************************************
# # FOR SLIGHTLY WORN ROAD
lat2=[]
long2=[]
for item in pslworn:
lat2.append(item[0])
long2.append(item[1])
source2 = ColumnDataSource(
data=dict(
lat2=lat2,
lon2=long2,
)
)
circle2 = Circle(x="lon2", y="lat2", size=13, fill_color="yellow", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source2, circle2)
#output_notebook()
#plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
#show(plot_pred)
# *****************************************************************************************************
# # FOR WORN ROAD
lat3=[]
long3=[]
for item in pworn:
lat3.append(item[0])
long3.append(item[1])
source3 = ColumnDataSource(
data=dict(
lat3=lat3,
lon3=long3,
)
)
circle3 = Circle(x="lon3", y="lat3", size=10, fill_color="purple", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source3, circle3)
#output_notebook()
#plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
#show(plot_pred)
# *****************************************************************************************************
# # FOR HEAVY ROAD
lat4=[]
long4=[]
for item in pheavy:
lat4.append(item[0])
long4.append(item[1])
source4 = ColumnDataSource(
data=dict(
lat4=lat4,
lon4=long4,
)
)
circle4 = Circle(x="lon4", y="lat4", size=8, fill_color="red", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source4, circle4)
output_notebook()
plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
show(plot_pred)
# -
# ## Preparing test data
# +
import os
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
from scipy import misc
import matplotlib.image as mpimg
import csv
import sys
from collections import defaultdict
import sys
import re
import time
import random
import cv2
from pylab import imread, imshow, gray, mean
gray()
# column names same as in csv file
col_names = ['VidName','ImgName','Lati','Longi']
np.random.seed(7)
time_steps=19
# load the dataset
# path of the video file csv with lat and longi score values
video_scores_path = "./Test.csv"
dataframe = pd.read_csv(video_scores_path)
dataset = dataframe.values
# we group by day so we can process a video at a time.
grouped = dataframe.groupby(dataframe.VidName)
per_vid = []
for _, group in grouped:
per_vid.append(group)
print(len(per_vid))
trainX=[]
trainY=[]
# generate sequences a vid at a time
for i,vid in enumerate(per_vid):
histValuesList=[]
# if we have less than 20 datapoints for a vid we skip over the
# vid assuming something is missing in the raw data
total = vid[col_names].values
if len(total) < time_steps :
continue
scoreVal=vid["Score"].values[0] + 1
max_total_for_vid = scoreVal.tolist()
#max_total_for_vid = vid["Score"].values[0].tolist()
histValuesList.append(total[1])
trainX.append(histValuesList)
#trainX.append(total[0:time_steps+1])
trainY.append([max_total_for_vid])
print(len(trainX[0]))
#trainX = np.array([np.array(xi) for xi in trainX])
trainX=np.array(trainX)
trainY=np.array(trainY)
print(trainX.shape,trainY.shape)
# -
# +
prediction=[[0,2.2048342],
[0,1.1126734],
[0,1.3997675],
[0,0.5157764],
[1,0.6071486],
[3,2.0375755],
[0,0.99859166],
[4,1.7473166],
[0,1.1756531],
[0,1.6052155],
[3,1.8819863],
[4,1.265181],
[1,1.3298031],
[2,1.2512382],
[1,1.0782294],
[2,1.5557319],
[1,1.3530238],
[2,1.0890144],
[1,1.8473151],
[0,0.8526046],
[0,1.0932784],
[0,1.2102916],
[2,1.2795185],
[0,0.67705584],
[0,0.6167114],
[0,0.6345362],
[2,0.8060204],
[1,1.1223748],
[3,1.4328588],
[0,1.0436226],
[0,0.7542808],
[0,0.81572336],
[0,1.2461239],
[2,0.83412206],
[2,1.0093734],
[1,0.8660643],
[1,2.1815915],
[2,1.4881321],
[1,1.6342332],
[2,1.5071036],
[1,1.5950464],
[1,1.2154874],
[4,1.5704111],
[1,1.4999061],
[1,0.95844793],
[2,1.0333613],
[0,0.82135975],
[1,1.3967812],
[1,1.0393807],
[1,1.1608542],
[2,0.669296],
[2,0.7003008]]
# +
truth=[]
pred=[]
for item in zip(trainX,prediction):
#print(item[0][0][2],item[1][0])
truthTemp=[]
predTemp=[]
truthTemp.append(item[0][0][2])
truthTemp.append(item[0][0][3])
truthTemp.append(item[1][0])
predTemp.append(item[0][0][2])
predTemp.append(item[0][0][3])
predTemp.append(item[1][1])
truth.append(truthTemp)
pred.append(predTemp)
truth
print("pred")
truth
# +
# visualizing ground truth first
tskip=[]
tnew=[]
tslworn=[]
tworn=[]
theavy=[]
for elem in truth:
if(elem[0]==0 or elem[1]==0):
continue
if(elem[2]==0):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
tskip.append(tempL)
elif(elem[2]==1):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
tnew.append(tempL)
elif(elem[2]==2):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
tslworn.append(tempL)
elif(elem[2]==3):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
tworn.append(tempL)
elif(elem[2]==4):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
theavy.append(tempL)
# +
print(tskip)
lat0=[]
long0=[]
for item in tskip:
print(item[0],item[1])
lat0.append(item[0])
long0.append(item[0])
lat0=[]
long0=[]
for item in tskip:
lat0.append(item[0])
long0.append(item[1])
print(lat0)
print(long0)
# -
# # Visualizing Test data ground truth on google maps.
#
# #### Please replace plot.api_key with your own key
# +
# ADDING ONE AT A TIME
# ['#440154', '#404387', '#29788E', '#22A784', '#79D151', '#FDE724']
from bokeh.io import output_file, output_notebook, show
from bokeh.models import (
GMapPlot, GMapOptions, ColumnDataSource, Circle, LogColorMapper, BasicTicker, ColorBar,
DataRange1d, PanTool, WheelZoomTool, BoxSelectTool
)
from bokeh.models.mappers import ColorMapper, LinearColorMapper
from bokeh.palettes import Viridis5
from bokeh.io import output_file, show
from bokeh.models import (
GMapPlot, GMapOptions, ColumnDataSource, Circle, Range1d, PanTool, WheelZoomTool, BoxSelectTool
)
map_options = GMapOptions(lat=37.5324, lng=-121.9687, map_type="roadmap", zoom=11)
plot = GMapPlot(x_range=Range1d(), y_range=Range1d(), map_options=map_options)
plot.title.text = "San Francisco Bay Area"
# For GMaps to function, Google requires you obtain and enable an API key:
#
# https://developers.google.com/maps/documentation/javascript/get-api-key
#
# Replace the value below with your personal API key:
plot.api_key = "key"
lat0=[]
long0=[]
for item in tskip:
lat0.append(item[0])
long0.append(item[1])
source = ColumnDataSource(
data=dict(
lat=lat0,
lon=long0,
)
)
circle = Circle(x="lon", y="lat", size=6, fill_color="black", fill_alpha=1.8, line_color=None)
plot.add_glyph(source, circle)
#output_notebook()
#plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
#show(plot)
# *****************************************************************************************************
# # FOR NEW ROAD
lat1=[]
long1=[]
for item in tnew:
lat1.append(item[0])
long1.append(item[1])
source1 = ColumnDataSource(
data=dict(
lat1=lat1,
lon1=long1,
)
)
circle1 = Circle(x="lon1", y="lat1", size=16, fill_color="green", fill_alpha=1.8, line_color=None)
plot.add_glyph(source1, circle1)
#output_notebook()
#plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
#show(plot)
# *****************************************************************************************************
# # FOR SLIGHTLY WORN ROAD
lat2=[]
long2=[]
for item in tslworn:
lat2.append(item[0])
long2.append(item[1])
source2 = ColumnDataSource(
data=dict(
lat2=lat2,
lon2=long2,
)
)
circle2 = Circle(x="lon2", y="lat2", size=13, fill_color="yellow", fill_alpha=1.8, line_color=None)
plot.add_glyph(source2, circle2)
#output_notebook()
#plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
#show(plot)
# *****************************************************************************************************
# # FOR WORN ROAD
lat3=[]
long3=[]
for item in tworn:
lat3.append(item[0])
long3.append(item[1])
source3 = ColumnDataSource(
data=dict(
lat3=lat3,
lon3=long3,
)
)
circle3 = Circle(x="lon3", y="lat3", size=10, fill_color="purple", fill_alpha=1.8, line_color=None)
plot.add_glyph(source3, circle3)
#output_notebook()
#plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
#show(plot)
# *****************************************************************************************************
# # FOR HEAVY ROAD
lat4=[]
long4=[]
for item in theavy:
lat4.append(item[0])
long4.append(item[1])
source4 = ColumnDataSource(
data=dict(
lat4=lat4,
lon4=long4,
)
)
circle4 = Circle(x="lon4", y="lat4", size=8, fill_color="red", fill_alpha=1.8, line_color=None)
plot.add_glyph(source4, circle4)
output_notebook()
plot.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot.html")
show(plot)
# -
pred
# +
from bokeh.io import output_file, output_notebook, show
from bokeh.models import (
GMapPlot, GMapOptions, ColumnDataSource, Circle, LogColorMapper, BasicTicker, ColorBar,
DataRange1d, PanTool, WheelZoomTool, BoxSelectTool
)
from bokeh.models.mappers import ColorMapper, LinearColorMapper
from bokeh.palettes import Viridis5
from bokeh.io import output_file, show
from bokeh.models import (
GMapPlot, GMapOptions, ColumnDataSource, Circle, Range1d, PanTool, WheelZoomTool, BoxSelectTool
)
# visualizing predictions
pskip=[]
pnew=[]
pslworn=[]
pworn=[]
pheavy=[]
for elem in pred:
if(elem[0]==0 or elem[1]==0):
continue
if(elem[2]>=0 and elem[2]<=0.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pskip.append(tempL)
elif(elem[2]>0.5 and elem[2]<=1.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pnew.append(tempL)
elif(elem[2]>1.5 and elem[2]<=2.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pslworn.append(tempL)
elif(elem[2]>2.5 and elem[2]<=3.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pworn.append(tempL)
elif(elem[2]>3.5):
tempL=[]
tempL.append(elem[0])
tempL.append(elem[1])
pheavy.append(tempL)
# -
# # Visualizing Test data predictions on google maps.
#
# #### Please replace plot.api_key with your own key
# +
# predictions plot_pred
# ADDING ONE AT A TIME
# ['#440154', '#404387', '#29788E', '#22A784', '#79D151', '#FDE724']
map_options = GMapOptions(lat=37.5324, lng=-121.9687, map_type="roadmap", zoom=11)
plot_pred = GMapPlot(x_range=Range1d(), y_range=Range1d(), map_options=map_options)
plot_pred.title.text = "San Francisco Bay Area"
# For GMaps to function, Google requires you obtain and enable an API key:
#
# https://developers.google.com/maps/documentation/javascript/get-api-key
#
# Replace the value below with your personal API key:
plot_pred.api_key = "key"
lat0=[]
long0=[]
for item in pskip:
lat0.append(item[0])
long0.append(item[1])
source = ColumnDataSource(
data=dict(
lat=lat0,
lon=long0,
)
)
circle = Circle(x="lon", y="lat", size=6, fill_color="black", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source, circle)
#output_notebook()
#plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
#show(plot_pred)
# *****************************************************************************************************
# # FOR NEW ROAD
lat1=[]
long1=[]
for item in pnew:
lat1.append(item[0])
long1.append(item[1])
source1 = ColumnDataSource(
data=dict(
lat1=lat1,
lon1=long1,
)
)
circle1 = Circle(x="lon1", y="lat1", size=16, fill_color="green", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source1, circle1)
#output_notebook()
#plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
#show(plot_pred)
# *****************************************************************************************************
# # FOR SLIGHTLY WORN ROAD
lat2=[]
long2=[]
for item in pslworn:
lat2.append(item[0])
long2.append(item[1])
source2 = ColumnDataSource(
data=dict(
lat2=lat2,
lon2=long2,
)
)
circle2 = Circle(x="lon2", y="lat2", size=13, fill_color="yellow", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source2, circle2)
#output_notebook()
#plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
#show(plot_pred)
# *****************************************************************************************************
# # FOR WORN ROAD
lat3=[]
long3=[]
for item in pworn:
lat3.append(item[0])
long3.append(item[1])
source3 = ColumnDataSource(
data=dict(
lat3=lat3,
lon3=long3,
)
)
circle3 = Circle(x="lon3", y="lat3", size=10, fill_color="purple", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source3, circle3)
#output_notebook()
#plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
#show(plot_pred)
# *****************************************************************************************************
# # FOR HEAVY ROAD
lat4=[]
long4=[]
for item in pheavy:
lat4.append(item[0])
long4.append(item[1])
source4 = ColumnDataSource(
data=dict(
lat4=lat4,
lon4=long4,
)
)
circle4 = Circle(x="lon4", y="lat4", size=8, fill_color="red", fill_alpha=1.8, line_color=None)
plot_pred.add_glyph(source4, circle4)
output_notebook()
plot_pred.add_tools(PanTool(), WheelZoomTool(), BoxSelectTool())
#output_file("gmap_plot_pred.html")
show(plot_pred)
|
Visualization_Best_Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="JSjG64ra4aFu"
# from google.colab import drive
# drive.mount('/content/drive')
# # !pip install torch
# + id="V8-7SARDZErK"
import torch.nn as nn
import torch.nn.functional as F
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import torch
import torchvision
import torchvision.transforms as transforms
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, utils
import torch.optim as optim
from matplotlib import pyplot as plt
import copy
import pickle
# Ignore warnings
import warnings
warnings.filterwarnings("ignore")
torch.backends.cudnn.deterministic = True
torch.backends.cudnn.benchmark = False
# + colab={"base_uri": "https://localhost:8080/"} id="vwJv7Y8Rewez" outputId="09fc47d8-9952-4425-e954-aa84e192c426"
transform = transforms.Compose(
[transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
trainset = torchvision.datasets.CIFAR10(root='./data', train=True, download=True, transform=transform)
testset = torchvision.datasets.CIFAR10(root='./data', train=False, download=True, transform=transform)
trainloader = torch.utils.data.DataLoader(trainset, batch_size=10, shuffle=False)
testloader = torch.utils.data.DataLoader(testset, batch_size=10, shuffle=False)
classes = ('plane', 'car', 'bird', 'cat', 'deer', 'dog', 'frog', 'horse', 'ship', 'truck')
foreground_classes = {'plane', 'car', 'bird'}
background_classes = {'cat', 'deer', 'dog', 'frog', 'horse','ship', 'truck'}
# print(type(foreground_classes))
dataiter = iter(trainloader)
background_data=[]
background_label=[]
foreground_data=[]
foreground_label=[]
batch_size=10
for i in range(5000): #5000*batch_size = 50000 data points
images, labels = dataiter.next()
for j in range(batch_size):
if(classes[labels[j]] in background_classes):
img = images[j].tolist()
background_data.append(img)
background_label.append(labels[j])
else:
img = images[j].tolist()
foreground_data.append(img)
foreground_label.append(labels[j])
foreground_data = torch.tensor(foreground_data)
foreground_label = torch.tensor(foreground_label)
background_data = torch.tensor(background_data)
background_label = torch.tensor(background_label)
# + id="nId70Q1P602e"
fg1, fg2, fg3 = 0,1,2
# + id="6SnnV31U60zk"
def imshow(img):
img = img / 2 + 0.5 # unnormalize
npimg = img#.numpy()
plt.imshow(np.transpose(npimg, (1, 2, 0)))
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="WK8Gp3u365CD" outputId="e761c6ba-e4e7-4e85-a198-656bdea8bc69"
foreground_data.shape, foreground_label.shape, background_data.shape, background_label.shape
# + id="FWijU1c565AO"
mean_bg = torch.mean(background_data, dim=0, keepdims= True)
std_bg = torch.std(background_data, dim=0, keepdims= True)
# + colab={"base_uri": "https://localhost:8080/"} id="8I2VnepD64-f" outputId="2b105e4d-17f5-448e-f034-cf8142a204b0"
mean_bg.shape, std_bg.shape
# + id="JUCSXEZM646m"
foreground_data = (foreground_data - mean_bg) / std_bg
background_data = (background_data - mean_bg) / std_bg
# + colab={"base_uri": "https://localhost:8080/"} id="S5Z_2PmE6422" outputId="ee8dae71-8ecf-463f-f273-06ee08c044dd"
foreground_data.shape, foreground_label.shape, background_data.shape, background_label.shape
# + colab={"base_uri": "https://localhost:8080/"} id="-D9vhiFj640j" outputId="55a7839d-5360-4ca8-a787-ebf9c1596d8d"
torch.sum(torch.isnan(foreground_data)), torch.sum(torch.isnan(background_data))
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="z0iJyjBu64yA" outputId="f7266dba-8ade-4ce4-edd6-e356ecbc9992"
imshow(foreground_data[0])
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="71tH2MMh64vE" outputId="fd777dbf-5abc-4d84-fcaa-a5e974fb4722"
imshow(background_data[2])
# + id="9nDYhjJse6Qq"
def create_mosaic_img(bg_idx,fg_idx,fg):
"""
bg_idx : list of indexes of background_data[] to be used as background images in mosaic
fg_idx : index of image to be used as foreground image from foreground data
fg : at what position/index foreground image has to be stored out of 0-8
"""
image_list=[]
j=0
for i in range(9):
if i != fg:
image_list.append(background_data[bg_idx[j]])
j+=1
else:
image_list.append(foreground_data[fg_idx])
label = foreground_label[fg_idx] - fg1 # minus fg1 because our fore ground classes are fg1,fg2,fg3 but we have to store it as 0,1,2
#image_list = np.concatenate(image_list ,axis=0)
image_list = torch.stack(image_list)
return image_list,label
# + id="aivGVg14e9iZ"
desired_num = 20000
mosaic_list_of_images =[] # list of mosaic images, each mosaic image is saved as list of 9 images
fore_idx =[] # list of indexes at which foreground image is present in a mosaic image i.e from 0 to 9
mosaic_label=[] # label of mosaic image = foreground class present in that mosaic
list_set_labels = []
for i in range(desired_num):
set_idx = set()
np.random.seed(i)
bg_idx = np.random.randint(0,35000,8)
set_idx = set(background_label[bg_idx].tolist())
fg_idx = np.random.randint(0,15000)
set_idx.add(foreground_label[fg_idx].item())
fg = np.random.randint(0,9)
fore_idx.append(fg)
image_list,label = create_mosaic_img(bg_idx,fg_idx,fg)
mosaic_list_of_images.append(image_list)
mosaic_label.append(label)
list_set_labels.append(set_idx)
# + colab={"base_uri": "https://localhost:8080/"} id="cog5VUzGgE5L" outputId="132dacd2-d144-47e3-a3ae-9079d9889bbe"
print(len(mosaic_list_of_images) , len(mosaic_label), len(mosaic_list_of_images[0:10000]))
print(len(fore_idx))
# + id="xX91RwMy-IP4"
def create_avg_image_from_mosaic_dataset(mosaic_dataset,labels,foreground_index,dataset_number):
"""
mosaic_dataset : mosaic_dataset contains 9 images 32 x 32 each as 1 data point
labels : mosaic_dataset labels
foreground_index : contains list of indexes where foreground image is present so that using this we can take weighted average
dataset_number : will help us to tell what ratio of foreground image to be taken. for eg: if it is "j" then fg_image_ratio = j/9 , bg_image_ratio = (9-j)/8*9
"""
avg_image_dataset = []
for i in range(len(mosaic_dataset)):
img = torch.zeros([3, 32,32], dtype=torch.float64)
for j in range(9):
if j == foreground_index[i]:
img = img + mosaic_dataset[i][j]*dataset_number/9
else :
img = img + mosaic_dataset[i][j]*(9-dataset_number)/(8*9)
avg_image_dataset.append(img)
return avg_image_dataset , labels , foreground_index
# + colab={"base_uri": "https://localhost:8080/"} id="n_TpL_3nRiy1" outputId="b853cd8b-ccc4-42aa-ce2a-c06806b616fd"
mosaic_list_of_images = torch.stack(mosaic_list_of_images)
mosaic_list_of_images.shape
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="WXXH6iF7RJsN" outputId="2dd262f8-bb56-46c2-8c71-f8eb64d6a31e"
imshow(mosaic_list_of_images[0][2])
# + id="pbyUn4M0RNPL"
mean_train = torch.mean(mosaic_list_of_images[0:10000], dim=0, keepdims= True)
std_train = torch.std(mosaic_list_of_images[0:10000], dim=0, keepdims= True)
mosaic_list_of_images = (mosaic_list_of_images - mean_train) / std_train
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="O-KROVzVRgEL" outputId="ac1cd0ab-77cc-4726-96be-81c1cab42567"
imshow(mosaic_list_of_images[0][2])
# + id="LGz8Y88vIZPT"
avg_image_dataset_0 , labels_0, fg_index_0 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 0)
avg_image_dataset_1 , labels_1, fg_index_1 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 1)
avg_image_dataset_2 , labels_2, fg_index_2 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 2)
avg_image_dataset_3 , labels_3, fg_index_3 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 3)
avg_image_dataset_4 , labels_4, fg_index_4 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 4)
avg_image_dataset_5 , labels_5, fg_index_5 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 5)
avg_image_dataset_6 , labels_6, fg_index_6 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 6)
avg_image_dataset_7 , labels_7, fg_index_7 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 7)
avg_image_dataset_8 , labels_8, fg_index_8 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 8)
avg_image_dataset_9 , labels_9, fg_index_9 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[0:10000], mosaic_label[0:10000], fore_idx[0:10000] , 9)
test_dataset_10 , labels_10 , fg_index_10 = create_avg_image_from_mosaic_dataset(mosaic_list_of_images[10000:20000], mosaic_label[10000:20000], fore_idx[10000:20000] , 9)
# avg_image_dataset_1 = torch.stack(avg_image_dataset_1)
# avg_image_dataset_2 = torch.stack(avg_image_dataset_2)
# avg_image_dataset_3 = torch.stack(avg_image_dataset_3)
# avg_image_dataset_4 = torch.stack(avg_image_dataset_4)
# avg_image_dataset_5 = torch.stack(avg_image_dataset_5)
# avg_image_dataset_6 = torch.stack(avg_image_dataset_6)
# avg_image_dataset_7 = torch.stack(avg_image_dataset_7)
# avg_image_dataset_8 = torch.stack(avg_image_dataset_8)
# avg_image_dataset_9 = torch.stack(avg_image_dataset_9)
# test_dataset_10 = torch.stack(test_dataset_10)
# + colab={"base_uri": "https://localhost:8080/", "height": 552} id="UJrZaJfR8s99" outputId="2fa550db-0d1b-4b7f-ca15-2b1b4571a99b"
imshow(avg_image_dataset_1[0]), imshow(avg_image_dataset_9[0])
# + id="9Ay6qqeZ7_I-"
# mean_train = torch.mean(avg_image_dataset_1, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_1, dim=0, keepdims= True)
# avg_image_dataset_1 = (avg_image_dataset_1 - mean_train) / std_train
# avg_image_dataset_1_test = (test_dataset_10 - mean_train) / std_train
# mean_train = torch.mean(avg_image_dataset_2, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_2, dim=0, keepdims= True)
# avg_image_dataset_2 = (avg_image_dataset_2 - mean_train) / std_train
# avg_image_dataset_2_test = (test_dataset_10 - mean_train) / std_train
# mean_train = torch.mean(avg_image_dataset_3, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_3, dim=0, keepdims= True)
# avg_image_dataset_3 = (avg_image_dataset_3 - mean_train) / std_train
# avg_image_dataset_3_test = (test_dataset_10 - mean_train) / std_train
# mean_train = torch.mean(avg_image_dataset_4, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_4, dim=0, keepdims= True)
# avg_image_dataset_4 = (avg_image_dataset_4 - mean_train) / std_train
# avg_image_dataset_4_test = (test_dataset_10 - mean_train) / std_train
# mean_train = torch.mean(avg_image_dataset_5, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_5, dim=0, keepdims= True)
# avg_image_dataset_5 = (avg_image_dataset_5 - mean_train) / std_train
# avg_image_dataset_5_test = (test_dataset_10 - mean_train) / std_train
# mean_train = torch.mean(avg_image_dataset_6, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_6, dim=0, keepdims= True)
# avg_image_dataset_6 = (avg_image_dataset_6 - mean_train) / std_train
# avg_image_dataset_6_test = (test_dataset_10 - mean_train) / std_train
# mean_train = torch.mean(avg_image_dataset_7, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_7, dim=0, keepdims= True)
# avg_image_dataset_7 = (avg_image_dataset_7 - mean_train) / std_train
# avg_image_dataset_7_test = (test_dataset_10 - mean_train) / std_train
# mean_train = torch.mean(avg_image_dataset_8, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_8, dim=0, keepdims= True)
# avg_image_dataset_8 = (avg_image_dataset_8 - mean_train) / std_train
# avg_image_dataset_8_test = (test_dataset_10 - mean_train) / std_train
# mean_train = torch.mean(avg_image_dataset_9, dim=0, keepdims= True)
# std_train = torch.std(avg_image_dataset_9, dim=0, keepdims= True)
# avg_image_dataset_9 = (avg_image_dataset_9 - mean_train) / std_train
# avg_image_dataset_9_test = (test_dataset_10 - mean_train) / std_train
# + id="nSO9SFE25Lrk"
class MosaicDataset(Dataset):
"""MosaicDataset dataset."""
def __init__(self, mosaic_list_of_images, mosaic_label):
"""
Args:
csv_file (string): Path to the csv file with annotations.
root_dir (string): Directory with all the images.
transform (callable, optional): Optional transform to be applied
on a sample.
"""
self.mosaic = mosaic_list_of_images
self.label = mosaic_label
#self.fore_idx = fore_idx
def __len__(self):
return len(self.label)
def __getitem__(self, idx):
return self.mosaic[idx] , self.label[idx] #, self.fore_idx[idx]
# + id="faQWc1-wN3p6"
batch = 250
traindata_0 = MosaicDataset(avg_image_dataset_0, labels_0 )
trainloader_0 = DataLoader( traindata_0 , batch_size= batch ,shuffle=True)
traindata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
trainloader_1 = DataLoader( traindata_1 , batch_size= batch ,shuffle=True)
traindata_2 = MosaicDataset(avg_image_dataset_2, labels_2 )
trainloader_2 = DataLoader( traindata_2 , batch_size= batch ,shuffle=True)
traindata_3 = MosaicDataset(avg_image_dataset_3, labels_3 )
trainloader_3 = DataLoader( traindata_3 , batch_size= batch ,shuffle=True)
traindata_4 = MosaicDataset(avg_image_dataset_4, labels_4 )
trainloader_4 = DataLoader( traindata_4 , batch_size= batch ,shuffle=True)
traindata_5 = MosaicDataset(avg_image_dataset_5, labels_5 )
trainloader_5 = DataLoader( traindata_5 , batch_size= batch ,shuffle=True)
traindata_6 = MosaicDataset(avg_image_dataset_6, labels_6 )
trainloader_6 = DataLoader( traindata_6 , batch_size= batch ,shuffle=True)
traindata_7 = MosaicDataset(avg_image_dataset_7, labels_7 )
trainloader_7 = DataLoader( traindata_7 , batch_size= batch ,shuffle=True)
traindata_8 = MosaicDataset(avg_image_dataset_8, labels_8 )
trainloader_8 = DataLoader( traindata_8 , batch_size= batch ,shuffle=True)
traindata_9 = MosaicDataset(avg_image_dataset_9, labels_9 )
trainloader_9 = DataLoader( traindata_9 , batch_size= batch ,shuffle=True)
# + id="obE1xeyRks1Q"
testdata_0 = MosaicDataset(avg_image_dataset_0, labels_0 )
testloader_0 = DataLoader( testdata_0 , batch_size= batch ,shuffle=False)
testdata_1 = MosaicDataset(avg_image_dataset_1, labels_1 )
testloader_1 = DataLoader( testdata_1 , batch_size= batch ,shuffle=False)
testdata_2 = MosaicDataset(avg_image_dataset_2, labels_2 )
testloader_2 = DataLoader( testdata_2 , batch_size= batch ,shuffle=False)
testdata_3 = MosaicDataset(avg_image_dataset_3, labels_3 )
testloader_3 = DataLoader( testdata_3 , batch_size= batch ,shuffle=False)
testdata_4 = MosaicDataset(avg_image_dataset_4, labels_4 )
testloader_4 = DataLoader( testdata_4 , batch_size= batch ,shuffle=False)
testdata_5 = MosaicDataset(avg_image_dataset_5, labels_5 )
testloader_5 = DataLoader( testdata_5 , batch_size= batch ,shuffle=False)
testdata_6 = MosaicDataset(avg_image_dataset_6, labels_6 )
testloader_6 = DataLoader( testdata_6 , batch_size= batch ,shuffle=False)
testdata_7 = MosaicDataset(avg_image_dataset_7, labels_7 )
testloader_7 = DataLoader( testdata_7 , batch_size= batch ,shuffle=False)
testdata_8 = MosaicDataset(avg_image_dataset_8, labels_8 )
testloader_8 = DataLoader( testdata_8 , batch_size= batch ,shuffle=False)
testdata_9 = MosaicDataset(avg_image_dataset_9, labels_9 )
testloader_9 = DataLoader( testdata_9 , batch_size= batch ,shuffle=False)
testdata_10 = MosaicDataset(test_dataset_10, labels_10 )
testloader_10 = DataLoader( testdata_10 , batch_size= batch ,shuffle=False)
# + id="SadRzWBBZEsP"
class Conv_module(nn.Module):
def __init__(self,inp_ch,f,s,k,pad):
super(Conv_module,self).__init__()
self.inp_ch = inp_ch
self.f = f
self.s = s
self.k = k
self.pad = pad
self.conv = nn.Conv2d(self.inp_ch,self.f,k,stride=s,padding=self.pad)
self.bn = nn.BatchNorm2d(self.f)
self.act = nn.ReLU()
torch.nn.init.xavier_normal_(self.conv.weight)
torch.nn.init.zeros_(self.conv.bias)
def forward(self,x):
x = self.conv(x)
x = self.bn(x)
x = self.act(x)
return x
# + id="IgGYMG_ZZEsT"
class inception_module(nn.Module):
def __init__(self,inp_ch,f0,f1):
super(inception_module, self).__init__()
self.inp_ch = inp_ch
self.f0 = f0
self.f1 = f1
self.conv1 = Conv_module(self.inp_ch,self.f0,1,1,pad=0)
self.conv3 = Conv_module(self.inp_ch,self.f1,1,3,pad=1)
#self.conv1 = nn.Conv2d(3,self.f0,1)
#self.conv3 = nn.Conv2d(3,self.f1,3,padding=1)
def forward(self,x):
x1 = self.conv1.forward(x)
x3 = self.conv3.forward(x)
#print(x1.shape,x3.shape)
x = torch.cat((x1,x3),dim=1)
return x
# + id="thkdqW91Hpju"
class downsample_module(nn.Module):
def __init__(self,inp_ch,f):
super(downsample_module,self).__init__()
self.inp_ch = inp_ch
self.f = f
self.conv = Conv_module(self.inp_ch,self.f,2,3,pad=0)
self.pool = nn.MaxPool2d(3,stride=2,padding=0)
def forward(self,x):
x1 = self.conv(x)
#print(x1.shape)
x2 = self.pool(x)
#print(x2.shape)
x = torch.cat((x1,x2),dim=1)
return x,x1
# + id="u1yVWgR4vFhe"
class inception_net(nn.Module):
def __init__(self):
super(inception_net,self).__init__()
self.conv1 = Conv_module(3,96,1,3,0)
self.incept1 = inception_module(96,32,32)
self.incept2 = inception_module(64,32,48)
self.downsample1 = downsample_module(80,80)
self.incept3 = inception_module(160,112,48)
self.incept4 = inception_module(160,96,64)
self.incept5 = inception_module(160,80,80)
self.incept6 = inception_module(160,48,96)
self.downsample2 = downsample_module(144,96)
self.incept7 = inception_module(240,176,60)
self.incept8 = inception_module(236,176,60)
self.pool = nn.AvgPool2d(5)
self.linear = nn.Linear(236,3)
torch.nn.init.xavier_normal_(self.linear.weight)
torch.nn.init.zeros_(self.linear.bias)
def forward(self,x):
x = self.conv1.forward(x)
#act1 = x
x = self.incept1.forward(x)
#act2 = x
x = self.incept2.forward(x)
#act3 = x
x,act4 = self.downsample1.forward(x)
x = self.incept3.forward(x)
#act5 = x
x = self.incept4.forward(x)
#act6 = x
x = self.incept5.forward(x)
#act7 = x
x = self.incept6.forward(x)
#act8 = x
x,act9 = self.downsample2.forward(x)
x = self.incept7.forward(x)
#act10 = x
x = self.incept8.forward(x)
#act11 = x
#print(x.shape)
x = self.pool(x)
#print(x.shape)
x = x.view(-1,1*1*236)
x = self.linear(x)
return x
# + id="cOWrnzv1fVjD"
def test_all(number, testloader,inc):
correct = 0
total = 0
out = []
pred = []
with torch.no_grad():
for data in testloader:
images, labels = data
images, labels = images.to("cuda"),labels.to("cuda")
out.append(labels.cpu().numpy())
outputs= inc(images)
_, predicted = torch.max(outputs.data, 1)
pred.append(predicted.cpu().numpy())
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 test dataset %d: %d %%' % (number , 100 * correct / total))
# + id="tFfAJZkcZEsY"
def train_all(trainloader, ds_number, testloader_list):
print("--"*40)
print("training on data set ", ds_number)
torch.manual_seed(12)
inc = inception_net().double()
inc = inc.to("cuda")
criterion_inception = nn.CrossEntropyLoss()
optimizer_inception = optim.Adam(inc.parameters(), lr=0.001 ) #, momentum=0.9)
acti = []
loss_curi = []
epochs = 100
for epoch in range(epochs): # loop over the dataset multiple times
ep_lossi = []
running_loss = 0.0
for i, data in enumerate(trainloader, 0):
# get the inputs
inputs, labels = data
inputs, labels = inputs.to("cuda"),labels.to("cuda")
# zero the parameter gradients
optimizer_inception.zero_grad()
# forward + backward + optimize
outputs = inc(inputs)
loss = criterion_inception(outputs, labels)
loss.backward()
optimizer_inception.step()
# print statistics
running_loss += loss.item()
if i % 10 == 9: # print every 10 mini-batches
print('[%d, %5d] loss: %.3f' %(epoch + 1, i + 1, running_loss / 10))
ep_lossi.append(running_loss/10) # loss per minibatch
running_loss = 0.0
loss_curi.append(np.mean(ep_lossi)) #loss per epoch
print("Epoch_Loss: ",np.round(np.mean(ep_lossi),5))
if(np.mean(ep_lossi) <= 0.005):
break
print('Finished Training')
# torch.save(inc.state_dict(),"/content/drive/My Drive/Research/Experiments on CIFAR mosaic/Exp_2_Attention_models_on_9_datasets_made_from_10k_mosaic/weights/train_dataset_"+str(ds_number)+"_"+str(epochs)+".pt")
correct = 0
total = 0
with torch.no_grad():
for data in trainloader:
images, labels = data
images, labels = images.to("cuda"), labels.to("cuda")
outputs = inc(images)
_, predicted = torch.max(outputs.data, 1)
total += labels.size(0)
correct += (predicted == labels).sum().item()
print('Accuracy of the network on the 10000 train images: %d %%' % ( 100 * correct / total))
for i, j in enumerate(testloader_list):
test_all(i+1, j,inc)
print("--"*40)
return loss_curi
# + colab={"base_uri": "https://localhost:8080/"} id="mI-vqhB-fVjJ" outputId="226c9976-3f67-47bb-9e20-6436fa7b9ed2"
train_loss_all=[]
testloader_list= [ testloader_0, testloader_1, testloader_2, testloader_3, testloader_4, testloader_5, testloader_6,
testloader_7, testloader_8, testloader_9, testloader_10]
train_loss_all.append(train_all(trainloader_0, 0, testloader_list))
train_loss_all.append(train_all(trainloader_1, 1, testloader_list))
train_loss_all.append(train_all(trainloader_2, 2, testloader_list))
train_loss_all.append(train_all(trainloader_3, 3, testloader_list))
train_loss_all.append(train_all(trainloader_4, 4, testloader_list))
train_loss_all.append(train_all(trainloader_5, 5, testloader_list))
train_loss_all.append(train_all(trainloader_6, 6, testloader_list))
train_loss_all.append(train_all(trainloader_7, 7, testloader_list))
train_loss_all.append(train_all(trainloader_8, 8, testloader_list))
train_loss_all.append(train_all(trainloader_9, 9, testloader_list))
# + id="AbZaQekCfVjN"
# %matplotlib inline
# + colab={"base_uri": "https://localhost:8080/", "height": 279} id="ouBomi5DfVjR" outputId="c9b15616-388c-4ec4-923a-316eaf1c0320"
fig = plt.figure()
for i,j in enumerate(train_loss_all):
plt.plot(j,label ="dataset "+str(i+1))
plt.xlabel("Epochs")
plt.ylabel("Training_loss")
plt.legend()
fig.savefig("Figure.pdf")
|
CODS_COMAD/CIN/CIFAR_10k_complex_mini_inception_8layers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import matplotlib.pyplot as plt
import tensorflow as tf
import numpy as np
import pandas as pd
import os
from sklearn.preprocessing import MinMaxScaler
from sklearn.metrics import mean_squared_error
from sklearn.metrics import accuracy_score
from sklearn.metrics import r2_score
# from tf.keras.models import Sequential # This does not work!
from tensorflow.python.keras.models import Sequential
from tensorflow.python.keras.layers import Input, Dense, GRU, Embedding
from tensorflow.python.keras.optimizers import RMSprop
from tensorflow.python.keras.callbacks import EarlyStopping, ModelCheckpoint, TensorBoard, ReduceLROnPlateau
from tensorflow.python.keras import metrics
from tensorflow.python.keras.layers import Dropout
from tensorflow.python.keras.layers import LeakyReLU
rawData = pd.read_csv('toyData.csv', sep=',')
rawData.head()
shiftMonths = 36
dfTargets = rawData.shift(-shiftMonths)
dfTargets.head()
xData = rawData.values[0:-shiftMonths]
print(type(xData))
print("Shape:", xData.shape)
yData = dfTargets.values[:-shiftMonths,0]
print(type(yData))
print("Shape:", yData.shape)
xTrain = xData[:(len(yData)-shiftMonths)]
xTest = xData[-shiftMonths:]
yTrain = yData[:(len(yData)-shiftMonths)]
yTest = yData[-shiftMonths:]
print ('Number of samples in training data:',len(xTrain))
print ('Number of samples in test data:',len(xTest))
xScaler = MinMaxScaler()
xTrainScaled = xScaler.fit_transform(xTrain)
print("Min:", np.min(xTrainScaled))
print("Max:", np.max(xTrainScaled))
xTestScaled = xScaler.transform(xTest)
yTrain = yTrain.reshape(-1, 1)
yTest = yTest.reshape(-1, 1)
yScaler = MinMaxScaler(feature_range=(2, 4))
yTrainScaled = yScaler.fit_transform(yTrain)
yTestScaled = yScaler.transform(yTest)
print(xTrainScaled.shape)
print(yTrainScaled.shape)
numXSignals = xData.shape[1]
numYSignals = 1
def batchGenerator(batchSize, sequenceLength):
"""
Generator function for creating random batches of training-data.
"""
# Infinite loop.
while True:
# Allocate a new array for the batch of input-signals.
numXSignals = xData.shape[1]
xShape = (batchSize, sequenceLength, numXSignals)
xBatch = np.zeros(shape=xShape, dtype=np.float16)
# Allocate a new array for the batch of output-signals.
numYSignals = 1
yShape = (batchSize, sequenceLength, numYSignals)
yBatch = np.zeros(shape=yShape, dtype=np.float16)
# Fill the batch with random sequences of data.
for i in range(batchSize):
# Get a random start-index.
# This points somewhere into the training-data.
numTrain = len(yTrain)
idx = np.random.randint(numTrain - sequenceLength)
# Copy the sequences of data starting at this index.
xBatch[i] = xTrainScaled[idx:idx+sequenceLength]
yBatch[i] = yTrainScaled[idx:idx+sequenceLength]
yield (xBatch, yBatch)
batchSize = 4
sequenceLength = 18
generator = batchGenerator(batchSize=batchSize,
sequenceLength=sequenceLength)
xBatch, yBatch = next(generator)
print(xBatch.shape)
print(yBatch.shape)
validationData = (np.expand_dims(xTestScaled, axis=0),
np.expand_dims(yTestScaled, axis=0))
model = Sequential()
model.add(GRU(units=1024,
activation='tanh',
return_sequences=True,
input_shape=(None, numXSignals,)))
model.add(Dropout(0.3))
model.add(Dense(numYSignals, activation='relu'))
# model.add(Dense(numYSignals))
# +
# model.add(LeakyReLU(alpha=.3))
# -
if False:
from tensorflow.python.keras.initializers import RandomUniform
# Maybe use lower init-ranges.
init = RandomUniform(minval=-0.05, maxval=0.05)
model.add(Dense(numYSignals,
activation='linear',
kernel_initializer=init))
optimizer = RMSprop(lr=1e-3)
model.compile(loss='mae',
optimizer=optimizer,
metrics = ['mse'])
model.summary()
pathCheckpoint = 'checkpoint.keras'
callbackCheckpoint = ModelCheckpoint(filepath=pathCheckpoint,
monitor='val_loss',
verbose=1,
save_weights_only=True,
save_best_only=True)
callbackEarlyStopping = EarlyStopping(monitor='val_loss',
patience=5, verbose=1)
# +
# callback_tensorboard = TensorBoard(log_dir='./23_logs/',
# histogram_freq=0,
# write_graph=False)
# -
callbackReduceLR = ReduceLROnPlateau(monitor='val_loss',
factor=0.1,
min_lr=1e-4,
patience=0,
verbose=1)
callbacks = [callbackEarlyStopping,
callbackCheckpoint,
# callback_tensorboard,
callbackReduceLR]
# %%time
history = model.fit_generator(generator=generator,
epochs=50,
steps_per_epoch=18,
validation_data=validationData,
callbacks=callbacks)
try:
model.load_weights(pathCheckpoint)
except Exception as error:
print("Error trying to load checkpoint.")
print(error)
# plot history
plt.plot(history.history['loss'], label='train')
plt.plot(history.history['val_loss'], label='test')
plt.legend()
plt.show()
result = model.evaluate(x=np.expand_dims(xTestScaled, axis=0),
y=np.expand_dims(yTestScaled, axis=0))
print("loss (test-set):", result)
# If you have several metrics you can use this instead.
if False:
for res, metric in zip(result, model.metrics_names):
print("{0}: {1:.3e}".format(metric, res))
x1 = np.expand_dims(xTrainScaled, axis=0)
yPred1 = model.predict(x1)
yPredRescaled1 = yScaler.inverse_transform(yPred1[0])
x2 = np.expand_dims(xTestScaled, axis=0)
yPred2 = model.predict(x2)
yPredRescaled2 = yScaler.inverse_transform(yPred2[0])
t = np.arange(len(rawData['Price']) - shiftMonths)
# +
# Make the plotting-canvas bigger.
plt.figure(figsize=(25,8))
# Plot and compare the two signals.
plt.plot(t[:len(yTrain)], yTrain, label='true train')
plt.plot(t[:len(yTrain)], yPredRescaled1, label='prediction train')
plt.plot(t[len(yTrain):], yTest, label='true test')
plt.plot(t[len(yTrain):], yPredRescaled2, label='prediction test')
plt.ylim([0,20])
# Plot labels etc.
plt.xlabel('Time Steps') # in number of months
plt.ylabel('Price')
plt.legend()
plt.show()
# -
np.sqrt(mean_squared_error(yTest, yPredRescaled2))
|
GRU.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="-3d28Q2JC6Rb" colab_type="text"
# ## Test the precision at k metric on SGD model trained on 25%
#
#
#
#
#
#
#
# + id="v0_omYmKgPYs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="8042704a-1ca3-4294-c192-8a5e5460b88e"
import joblib
import pandas as pd
import numpy as np
import logging
import nltk
import matplotlib.pyplot as plt
import re
from numpy import random
from sklearn.feature_extraction.text import CountVectorizer, TfidfVectorizer
from sklearn.metrics import accuracy_score, confusion_matrix
from nltk.corpus import stopwords
from sklearn.model_selection import train_test_split
from sklearn.linear_model import SGDClassifier
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import FunctionTransformer
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import accuracy_score
from sklearn.metrics import classification_report
from bs4 import BeautifulSoup
nltk.download('stopwords')
# + id="wrzBTJkWioyk" colab_type="code" colab={}
# + id="nrt_BHGdio13" colab_type="code" colab={}
df= pd.read_csv('/content/drive/My Drive/Colab Notebooks/data/reddit/df25.csv')
df.shape
labels = df['subreddit'].drop_duplicates()
# + id="KnA-WoZulwRo" colab_type="code" colab={}
X = df['title'].str.cat(df['selftext'], sep=' ')
y = df.subreddit
X_train, X_test, y_train,y_test = train_test_split(X,y,test_size=.2,random_state=42)
# label encode y
le = LabelEncoder()
y_train = le.fit_transform(y_train)
y_test = le.transform(y_test)
# + id="EcSYuHZjD0jB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6edd86e3-24b4-4b23-9be1-278c373d15cd"
y_test
# + [markdown] id="4uOgmkitkBBS" colab_type="text"
#
# + id="iTRMT0B5iGHH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ca846030-bf85-477a-dd56-52ddb35992b4"
X = df['title'].str.cat(df['selftext'], sep=' ')
y = df.subreddit
X.shape, y.shape
# + id="YZCsD96mhWJG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="c873359d-10be-48e0-82ee-9affe5ab71e3"
# %%time
# try sasaanas way
model = joblib.load('/content/drive/My Drive/Colab Notebooks/models/sgd25_trained.joblib')
# + id="Gb2V0HBO9RJa" colab_type="code" colab={}
#ref model
model = SGDClassifier(alpha=0.0001, average=False, class_weight=None,
early_stopping=True, epsilon=0.1, eta0=0.0, fit_intercept=True,
l1_ratio=0.15, learning_rate='optimal', loss='modified_huber',
max_iter=1000, n_iter_no_change=3, n_jobs=-1, penalty='l2',
power_t=0.5, random_state=42, shuffle=True, tol=0.001,
validation_fraction=0.1, verbose=0, warm_start=True)
# + id="4XFPpK_89Hqo" colab_type="code" colab={}
from sklearn.feature_selection import chi2, SelectKBest
chi2_selector = SelectKBest(chi2, k=30000)
NUM_FEATURES = 100000
sgdpipe = Pipeline([
('vect', TfidfVectorizer(
min_df=5,
ngram_range=(1,2),
stop_words=None,
token_pattern='(?u)\\b\\w+\\b',
)),
# ('select', chi2_selector),
('clf', model ) ## pay attn to transformer vs vectorizer
])
parameters = {
'vect__max_df': ( 0.7, 0.75, 0.8),
'vect__min_df': (.01, .02, .03),
'vect__max_features': (3000,30000),
'clf__loss': ['modified_huber'],
'clf__penalty': ['l2'],
'clf__alpha': [1e-3],
'clf__random_state': [42],
'clf__early_stopping': [True],
'clf__n_iter_no_change': [3],
'clf__max_iter': [100],
'clf__class_weight': [None],
'clf__warm_start': [False],
'clf__verbose': [0]
}
grid_search = GridSearchCV(sgdpipe, parameters, cv=3, n_jobs=-1, verbose=3)
# + id="AdoXVmyrJBEr" colab_type="code" colab={}
# + id="epuAfjl1I52y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 377} outputId="9b55d616-941f-4e36-9795-19cc8ce8e99a"
grid_search.fit(X_train,y_train)
grid_search.best_score_,grid_search.best_params_
# + id="iX3iVfPOJ3yw" colab_type="code" colab={}
# Needed imports
import numpy as np
from IPython.display import Audio
from scipy.io import wavfile
from google.colab import output
output.eval_js('new Audio("https://github.com/R6500/Python-bits/raw/master/Colaboratory/Sounds/Bicycle%20bell%203.wav").play()')
# + id="j65ikq689aTT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="172c68b1-f130-4b6e-b7d3-d11e92e1b33d"
pipe.fit(X_train,y_train)
# + [markdown] id="_vKX8wMijM8m" colab_type="text"
# The model specified below is just for reference, when run on the complete 1M rows dataset it achieved
#
#
# data_array = np.load('data.npy')
#
# + [markdown] id="JfZnTMbFkChl" colab_type="text"
#
# + id="B61lyjwhiGDV" colab_type="code" colab={}
# baseline linear svm with stochastic gradient descent training
sgd = SGDClassifier(loss='hinge',
penalty='l2',
alpha=1e-3,
random_state=42,
early_stopping=True,
max_iter =100, # num epochs
validation_fraction=0.1,
n_iter_no_change=3,
class_weight=None,
warm_start=True,
n_jobs=-1,
verbose=3)
# + id="SBiiVkOiB_NT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 51} outputId="34edb35b-383e-4842-a5d6-b374afedbd30"
# %%time
model = pipe.named_steps['clf']
y_pred_proba = pipe.predict_proba(X_test)
y_pred = np.argmax(y_pred_proba, axis=1)
def precision_at_k(y_true, y_pred, k=5):
y_true = np.array(y_true)
y_pred = np.array(y_pred)
y_pred = np.argsort(y_pred, axis=1)
y_pred = y_pred[:, ::-1][:, :k]
arr = [y in s for y, s in zip(y_true, y_pred)]
return np.mean(arr)
print('accuracy %s' % accuracy_score(y_pred, y_test))
# + id="XyDpIubpCVOa" colab_type="code" colab={}
# + id="MI2Dnm7VDWgJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 68} outputId="2423625f-bc9c-4bc1-afe0-e3796e855076"
print('precision@1 =', np.mean(y_test == y_pred))
print('precision@3 =', precision_at_k(y_test, y_pred_proba, 3))
print('precision@5 =', precision_at_k(y_test, y_pred_proba, 5))
# + id="nxeHeG_ukFdT" colab_type="code" colab={}
X_train = np.load('/content/drive/My Drive/Colab Notebooks/data/reddit/X25_trans.npy', allow_pickle=True)
X_test = np.load('/content/drive/My Drive/Colab Notebooks/data/reddit/X25_test_trans.npy', allow_pickle=True)
y_train = np.load('/content/drive/My Drive/Colab Notebooks/data/reddit/y25_trans_.npy', allow_pickle=True)
y_test = np.load('/content/drive/My Drive/Colab Notebooks/data/reddit/y25_test_trans_.npy', allow_pickle=True)
print(X_train.shape, y_train.shape, X_test.shape, y_test.shape)
# + [markdown] id="WEvpL4cGoG7U" colab_type="text"
# This saved model was trained on 25% of the dataset. and achieved
# accuracy 0.7334452122408687
#
# on test
# + id="1RURhx-gmqt3" colab_type="code" colab={}
# %%time
sgd25 = joblib.load('/content/drive/My Drive/Colab Notebooks/models/sgd25_trained.joblib')
# + id="cty9YOSjnIv0" colab_type="code" colab={}
y_pred25 = sgd25.predict(X_test)
print('accuracy %s' % accuracy_score(y_pred25, y_test))
class_report25 = classification_report(y_test, y_pred25)
|
notebooks/BW4_sgdc_precision_at_k.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Match Analysis
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
# -
# ## Data Cleaning and Exploration
matches = pd.read_csv("matches.csv" , index_col = "id")
matches = matches.iloc[:,:-3]
matches.shape
matches.winner.unique()
# ## Taking in consideration only KKR VS RR matches
KR = matches[np.logical_or(np.logical_and(matches['team1']=='Kolkata Knight Riders',matches['team2']=='Rajasthan Royals'),
np.logical_and(matches['team2']=='Kolkata Knight Riders',matches['team1']=='Rajasthan Royals'))]
KR.head()
KR.shape
KR.season.unique()
KR.isnull().sum()
KR.describe().iloc[:,2:].T
# ## Head To Head
KR.groupby("winner")["winner"].count()
sns.countplot(KR["winner"])
plt.text(-0.1,9,str(KR['winner'].value_counts()['Kolkata Knight Riders']),size=25,color='white')
plt.text(0.9,9,str(KR['winner'].value_counts()['Rajasthan Royals']),size=25,color='white')
plt.xlabel('Winner',fontsize=20)
plt.ylabel('No. of Matches',fontsize=20)
plt.title('RR VS KKR-head to head',fontsize = 20)
# +
Season_wise_Match_Winner = pd.DataFrame(KR.groupby(["season","winner"])["winner"].count())
Season_wise_Match_Winner.columns = ["winner count"]
print("Season wise winner of matches between RR VS KKR :")
Season_wise_Match_Winner
# -
# ## Winning Percentage
Winning_Percentage = KR['winner'].value_counts()/len(KR['winner'])
print("KKR winning percentage against RR(overall) : {}%".format(int(round(Winning_Percentage[0]*100))))
print("RR winning percentage against KKR(overall) : {}%".format(int(round(Winning_Percentage[1]*100))))
# ## Performance Analysis
def performance( team_name , given_df ):
for value in given_df.groupby('winner'):
if value[0] == team_name:
total_win_by_runs = sum(list(value[1]['win_by_runs']))
total_win_by_wickets = sum(list(value[1]['win_by_wickets']))
if 0 in list(value[1]['win_by_runs'].value_counts().index):
x = value[1]['win_by_runs'].value_counts()[0]
else:
x = 0
if 0 in list(value[1]['win_by_wickets'].value_counts().index):
y = value[1]['win_by_wickets'].value_counts()[0]
else:
y = 0
number_of_times_given_team_win_while_defending = (len(value[1]) - x )
number_of_times_given_team_win_while_chasing = (len(value[1]) - y )
average_runs_by_which_a_given_team_wins_while_defending = total_win_by_runs / number_of_times_given_team_win_while_defending
average_wickets_by_which_a_given_team_wins_while_chasing = total_win_by_wickets / number_of_times_given_team_win_while_chasing
print('Number of times given team win while defending :' , number_of_times_given_team_win_while_defending )
print('Number of times given team win while chasing :' , number_of_times_given_team_win_while_chasing )
print()
print('Average runs by which a given team wins while defending : ' ,round(average_runs_by_which_a_given_team_wins_while_defending))
print('Average wickets by which a given team wins while chasing : ' ,round(average_wickets_by_which_a_given_team_wins_while_chasing))
performance("Kolkata Knight Riders",KR)
performance("Rajasthan Royals",KR)
# ## Toss Analysis
# +
Toss_Decision = pd.DataFrame(KR.groupby(['toss_winner',"toss_decision"])["toss_decision"].count())
print ("Toss winner decision :")
Toss_Decision
# -
sns.set(style='whitegrid')
plt.figure(figsize = (18,8))
sns.countplot(KR['toss_winner'],palette='Set2',hue=KR['toss_decision'])
plt.title('Toss decision statistics for both team',fontsize=15)
plt.yticks(fontsize=15)
plt.xticks(fontsize=15)
plt.xlabel('Toss winner',fontsize=15)
plt.ylabel('Count',fontsize=15)
plt.legend(loc='best',fontsize=15)
plt.show()
# +
Toss_Decision_based_Winner = pd.DataFrame(KR.groupby(['toss_winner',"toss_decision","winner"])["winner"].count())
print(" No of times toss winning decision leading to match winning : ")
Toss_Decision_based_Winner
# -
sns.set(style='whitegrid')
plt.figure(figsize = (18,9))
sns.countplot(KR['toss_winner'],hue=KR['winner'])
plt.title('Match Winner vs Toss Winner statistics for both team',fontsize=15)
plt.yticks(fontsize=15)
plt.xticks(fontsize=15)
plt.xlabel('Toss winner',fontsize=15)
plt.ylabel('Match Winner',fontsize=15)
plt.legend(loc="best",fontsize=15)
# ### Toss Decision based Analysis of both the teams seperately :
KKR = KR[KR["toss_winner"]=="<NAME>"]
RR = KR[KR["toss_winner"]=="<NAME>"]
sns.set(style='whitegrid')
plt.figure(figsize = (18,9))
sns.countplot(KKR['toss_decision'],hue=KKR['winner'])
plt.title('Match Winner vs Toss Winner statistics for KKR',fontsize=15)
plt.yticks(fontsize=15)
plt.xticks(fontsize=15)
plt.xlabel('Toss decision of KKR',fontsize=15)
plt.ylabel('Match Winner',fontsize=15)
plt.legend(loc="best",fontsize=15)
sns.set(style='whitegrid')
plt.figure(figsize = (18,9))
sns.countplot(RR['toss_decision'],hue=RR['winner'])
plt.title('Match Winner vs Toss Winner statistics for RR',fontsize=15)
plt.yticks(fontsize=15)
plt.xticks(fontsize=15)
plt.xlabel('Toss decision of RR',fontsize=15)
plt.ylabel('Match Winner',fontsize=15)
plt.legend(loc="best",fontsize=15)
# +
player_of_the_match = pd.DataFrame(KR.player_of_match.value_counts())
print("Man of the match :")
player_of_the_match
# -
# ## Recent Year Performance Analysis
cond2 = KR["season"] == 2018
cond3 = KR["season"] == 2019
final = KR[cond2 | cond3]
final
final.shape
# +
player = pd.DataFrame(final.player_of_match.value_counts())
print("Man of the match :")
player
# +
plt.figure(figsize = (10,6))
sns.countplot(final['winner'])
plt.title('Match won in recent years',fontsize=15)
plt.yticks(fontsize=15)
plt.xticks(fontsize=15)
plt.xlabel('Team',fontsize=15)
plt.ylabel('Win Count',fontsize=15)
plt.show()
# -
# From all the above analysis related to Head to Head, Toss analysis,Performance Analysis, Recent year analysis etc we saw that KKR has better chances of winning the match specially if they chase the score.
|
KKR VS RR/Match Analysis KKR VS RR.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + pycharm={"name": "#%%\n"}
Pid_Set = 27
Pzr_pre = np.arange(Pid_Set-3,Pid_Set+3, 0.001)
# + pycharm={"name": "#%%\n"}
for A in [0, 1]:
rlist = []
for get_pre in Pzr_pre:
r = 0
if abs(get_pre - Pid_Set) < 0.25 and A == 0:
r += 1
else:
r += (2 - abs(get_pre - Pid_Set))/2
rlist.append(r)
plt.plot(rlist)
plt.plot(Pzr_pre/Pzr_pre.max())
# + pycharm={"name": "#%%\n"}
Pid_Set_Level = 30
# Pzr_lev = np.arange(0,100, 0.1)
Pzr_lev = np.arange(25, 35, 0.01)
for A in [0, 1]:
rlist = []
for get_pre in Pzr_lev:
r = 0
if abs(get_pre - Pid_Set_Level) < 0.25 and A == 0:
r += 1
else:
r_ = np.clip((2 - abs(get_pre - Pid_Set_Level))/2, 0, 1)
r += r_
rlist.append(r)
plt.plot(rlist)
plt.plot(Pzr_lev/100)
# + pycharm={"name": "#%%\n"}
db = pd.read_csv('DB/3_76.txt')
plt.plot(db['ZINST63'])
# + pycharm={"name": "#%%\n"}
|
PZR_bubblegeneration/Reward_Tester.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from DatasetHandler.BiwiBrowser import *
biwi = readBIWIDataset(subjectList = [s for s in range(1, 3)])#
print(now())
for inputMatrix, labels in biwi:
print(inputMatrix.shape, labels.shape)
print(now())
print(now())
for subj, (inputMatrix, labels) in biwi.items():
print(subj, inputMatrix.shape, labels.shape)
snippedBiwi = readBIWIDataset(frameTarFile = BIWI_SnippedData_file, labelsTarFile = BIWI_Lebels_file_Local)
|
DeepRL_For_HPE/TestingBiwiBrowser.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# %load_ext autoreload
# %autoreload 2
# # Two LJ particles
import molsysmt as msm
import numpy as np
from tqdm import tqdm
import openmm as mm
import openmm.app as app
import openmm.unit as unit
# +
# System definition.
## First atom: Argon
mass_1 = 39.948 * unit.amu
sigma_1 = 3.404 * unit.angstroms
epsilon_1 = 0.238 * unit.kilocalories_per_mole
charge_1 = 0.0 * unit.elementary_charge
## Second atomo: Xenon
mass_2 = 131.293 * unit.amu
sigma_2 = 3.961 * unit.angstroms
epsilon_2 = 0.459 * unit.kilocalories_per_mole
charge_2 = 0.0 * unit.elementary_charge
# System creation.
system = mm.System()
non_bonded_force = mm.NonbondedForce()
reduced_sigma = 0.5*(sigma_1+sigma_2)
cutoff_distance = 4.0*reduced_sigma
switching_distance = 3.0*reduced_sigma
non_bonded_force.setNonbondedMethod(mm.NonbondedForce.CutoffPeriodic)
non_bonded_force.setUseSwitchingFunction(True)
non_bonded_force.setCutoffDistance(cutoff_distance)
non_bonded_force.setSwitchingDistance(switching_distance)
# Adding atom 1
system.addParticle(mass_1)
non_bonded_force.addParticle(charge_1, sigma_1, epsilon_1)
# Adding atom 2
system.addParticle(mass_2)
non_bonded_force.addParticle(charge_2, sigma_2, epsilon_2)
# Periodic box
system.setDefaultPeriodicBoxVectors([3.0, 0.0, 0.0]*unit.nanometers, [0.0, 3.0, 0.0]*unit.nanometers, [0.0, 0.0, 3.0]*unit.nanometers)
_ = system.addForce(non_bonded_force)
# +
# Thermodynamic state and integrator.
step_size = 2*unit.femtoseconds
temperature = 300*unit.kelvin
friction = 1.0/unit.picosecond
integrator = mm.LangevinIntegrator(temperature, friction, step_size)
# +
# Platform.
platform_name = 'CUDA'
platform = mm.Platform.getPlatformByName(platform_name)
# +
# Contexto.
context = mm.Context(system, integrator, platform)
# +
# Initial conditions.
initial_positions = np.zeros([2, 3], np.float32) * unit.angstroms
initial_velocities = np.zeros([2, 3], np.float32) * unit.angstroms/unit.picoseconds
initial_positions[1, 0] = 1.0 * unit.nanometers
context.setPositions(initial_positions)
context.setVelocities(initial_velocities)
# +
# Simulation parameters.
simulation_time = 20.0*unit.nanosecond
saving_time = 1.0*unit.picoseconds
n_steps_per_saving_period = int(saving_time/step_size)
n_saving_periods = int(simulation_time/saving_time)
# +
# Numpy arrays as simulation reporters.
time = np.zeros([n_saving_periods], np.float32) * unit.picoseconds
position = np.zeros([n_saving_periods, 2, 3], np.float32) * unit.nanometers
velocity = np.zeros([n_saving_periods, 2, 3], np.float32) * unit.nanometers/unit.picosecond
potential_energy = np.zeros([n_saving_periods], np.float32) * unit.kilocalories_per_mole
kinetic_energy = np.zeros([n_saving_periods], np.float32) * unit.kilocalories_per_mole
box = np.zeros([n_saving_periods, 3, 3], np.float32) * unit.nanometers
# +
# Saving data for time 0 in reporters
state = context.getState(getPositions=True, getVelocities=True, getEnergy=True)
time[0] = state.getTime()
position[0] = state.getPositions()
velocity[0] = state.getVelocities()
kinetic_energy[0]=state.getKineticEnergy()
potential_energy[0]=state.getPotentialEnergy()
box[0] = state.getPeriodicBoxVectors()
# +
# Running the simulation
for ii in tqdm(range(n_saving_periods)):
context.getIntegrator().step(n_steps_per_saving_period)
state = context.getState(getPositions=True, getVelocities=True, getEnergy=True)
time[ii] = state.getTime()
position[ii] = state.getPositions()
velocity[ii] = state.getVelocities()
kinetic_energy[ii]=state.getKineticEnergy()
potential_energy[ii]=state.getPotentialEnergy()
box[ii] = state.getPeriodicBoxVectors()
# -
# ## molsysmt.TrajectoryDict class and trjdpk file
trajdict={
'time' : time,
'coordinates' : position,
'box' : box
}
msm.get_form(trajdict)
msm.convert(trajdict, to_form='Ar_Xe_pbc_vacuum.trjpk')
trajdict2 = msm.convert('Ar_Xe_pbc_vacuum.trjpk', to_form='molsysmt.TrajectoryDict')
trajdict2
|
docs/contents/demo/two_LJ_particles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="ELQ-Q3GCD0Cg" colab_type="code" colab={}
import pandas as pd
import numpy as np
from sklearn.tree import DecisionTreeRegressor
from sklearn.metrics import mean_absolute_error
from sklearn.model_selection import cross_val_score
# + id="SRCA0pZ4EJ5F" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="73458b1c-7b91-4ef2-a3fb-0b3db8af84cc" executionInfo={"status": "ok", "timestamp": 1581606167332, "user_tz": -60, "elapsed": 7699, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
# cd "/content/drive/My Drive/Colab Notebooks/dw_matrix"
# + id="rdq05GxIEsQY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="91487a24-919c-407b-e3e7-8d63ec6c47c8" executionInfo={"status": "ok", "timestamp": 1581606236637, "user_tz": -60, "elapsed": 1275, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
df = pd.read_csv('data/men_shoes.csv', low_memory = False)
df.shape
# + id="Dc1sPq4PE0Uj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="2f996028-2c0b-4d46-eb6b-419d3c0ca173" executionInfo={"status": "ok", "timestamp": 1581606259869, "user_tz": -60, "elapsed": 687, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
df.columns
# + id="RKuz-_DDFEZX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5ada3aa9-e00c-4529-94fc-253617c1d657" executionInfo={"status": "ok", "timestamp": 1581606326206, "user_tz": -60, "elapsed": 607, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
mean_price = np.mean(df['prices_amountmin'])
mean_price
# + id="wApnYCRCFUxL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="14603976-c4b0-4a60-a61c-76dbc901881f" executionInfo={"status": "ok", "timestamp": 1581606504777, "user_tz": -60, "elapsed": 923, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
y_true = df['prices_amountmin']
y_pred = [mean_price] * y_true.shape[0]
mean_absolute_error(y_true,y_pred)
# + id="Kt1Vv9krFqSX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="8ec62a4a-2297-4357-ee51-effcf6719f94" executionInfo={"status": "ok", "timestamp": 1581606557803, "user_tz": -60, "elapsed": 882, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
df['prices_amountmin'].hist(bins=100)
# + id="mAf-Auv_FtZf" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="6f36f955-ebcf-4ee0-c2a6-92203642327c" executionInfo={"status": "ok", "timestamp": 1581606620180, "user_tz": -60, "elapsed": 976, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
np.log1p(df['prices_amountmin']).hist(bins=100)
# + id="VOtyYgaTGUzB" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8120a607-da45-48c1-ee63-ebe2ddf03bf0" executionInfo={"status": "ok", "timestamp": 1581606695627, "user_tz": -60, "elapsed": 560, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
y_true = df['prices_amountmin']
y_pred = [np.median(y_true)] * y_true.shape[0]
mean_absolute_error(y_true,y_pred)
# + id="ZO_FrzTBGu_V" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4575251d-38b8-4437-b243-677ec45251d0" executionInfo={"status": "ok", "timestamp": 1581606933218, "user_tz": -60, "elapsed": 943, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
y_true = df['prices_amountmin']
price_log_mean = np.expm1(np.mean(np.log1p(y_true)))
y_pred = [price_log_mean] * y_true.shape[0]
mean_absolute_error(y_true,y_pred)
# + id="S75DhXivHo5A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="e97340ae-39b2-4b30-9343-9c4a42031163" executionInfo={"status": "ok", "timestamp": 1581606969861, "user_tz": -60, "elapsed": 1253, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
df.columns
# + id="RgbtRQY4Hxve" colab_type="code" colab={}
df['brand_cat'] = df['brand'].factorize()[0]
# + id="b9aHqnA8H41u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="8db97867-7151-4d90-89c8-b268e03f6817" executionInfo={"status": "ok", "timestamp": 1581607354142, "user_tz": -60, "elapsed": 709, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
feats = ['brand_cat']
X = df[feats].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, scoring = 'neg_mean_absolute_error')
np.mean(scores), np.std(scores)
# + id="C51-PmbhJExC" colab_type="code" colab={}
def run_model(feats):
X = df[feats].values
y = df['prices_amountmin'].values
model = DecisionTreeRegressor(max_depth=5)
scores = cross_val_score(model, X, y, scoring = 'neg_mean_absolute_error')
return np.mean(scores), np.std(scores)
# + id="_QZFijSQJjec" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="38e580e3-5f2f-4101-a89e-6bdb5c1c33d1" executionInfo={"status": "ok", "timestamp": 1581607460805, "user_tz": -60, "elapsed": 1001, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
run_model(['brand_cat'])
# + id="DOIVlosmJps0" colab_type="code" colab={}
df['manufacturer_cat'] = df['manufacturer'].factorize()[0]
# + id="doSYkTjrJ66m" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="07259b69-2d92-405d-8300-f34ca6e7e91d" executionInfo={"status": "ok", "timestamp": 1581607571666, "user_tz": -60, "elapsed": 1082, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
run_model(['manufacturer_cat'])
# + id="zkkVjjvIKj90" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b6c9d815-5d50-48cf-9ef6-7d06bcf6f98d" executionInfo={"status": "ok", "timestamp": 1581607718506, "user_tz": -60, "elapsed": 740, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
run_model(['brand_cat', 'manufacturer_cat'])
# + id="F6p12uqMKEuz" colab_type="code" colab={}
df['dimension_cat'] = df['dimension'].factorize()[0]
# + id="rD1O0CokKRPX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="70de9c0d-4b38-4e8d-bbf0-c7fbcdf51281" executionInfo={"status": "ok", "timestamp": 1581607640542, "user_tz": -60, "elapsed": 595, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
run_model(['dimension_cat'])
# + id="blzg0QXjKVp0" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7d7169a4-4d96-4a3d-f694-78cd3e9d7328" executionInfo={"status": "ok", "timestamp": 1581607750755, "user_tz": -60, "elapsed": 1839, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
run_model(['brand_cat', 'manufacturer_cat' , 'dimension_cat'])
# + id="0WO5lcYyKwQN" colab_type="code" colab={}
# !git add matrix_one/day4.ipynb
# + id="QhHk6B8ILbXk" colab_type="code" colab={}
# !git config --global user.email "<EMAIL>"
# !git config --global user.name "Monika"
# + id="v5EOiCfaLvtk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 69} outputId="f14d808e-e80c-47e3-f4af-c72390355b4b" executionInfo={"status": "ok", "timestamp": 1581608050659, "user_tz": -60, "elapsed": 10144, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "16179400812149074115"}}
# !git commit -m "Men's Shoe Prices - model"
# + id="XpXZkvmoL3c6" colab_type="code" colab={}
|
matrix_one/day4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Prepare per observation details
# +
import collections
import json
import bz2
import os
import configparser
import pandas
import numpy
from scipy.special import logit
from neo4j.v1 import GraphDatabase
import hetio.readwrite
import hetio.neo4j
# -
coef_df = pandas.read_table('model/coefficient.tsv')
feature_df = pandas.read_table('features/transformed-features.tsv.bz2', low_memory=False)
drop = {'prior_logit', 'intercept'}
coef_df = coef_df.query("feature not in @drop")
coef_df = coef_df.query("coef != 0")
coef_df = coef_df.sort_values('coef')
# ## Term contribution
coef_series = pandas.Series(data=coef_df.coef.tolist(), index=coef_df.feature)
contrib_df = feature_df[coef_df.feature].mul(coef_series, axis='columns')
contrib_df = feature_df[['compound_id', 'disease_id']].join(contrib_df)
contrib_df.head(2)
with bz2.open('./predictions/term-contribution.tsv.bz2', 'wt') as write_file:
contrib_df.to_csv(write_file, float_format='%.5g', sep='\t', index=False)
# ## Metapath contribution
pos_dwpc_coef_df = coef_df[(coef_df.feature.str.startswith('dwpc_')) & (coef_df.coef > 0)].copy()
pos_dwpc_coef_df['metapath'] = pos_dwpc_coef_df.feature.map(lambda x: x.split('_')[1])
pos_dwpc_coef_df.head(2)
# Estimate of percent contribution of each positive term with a positive coefficient
pos_contrib_df = contrib_df[pos_dwpc_coef_df.feature].copy()
pos_contrib_df[pos_contrib_df < 0] = 0
observation_total = pos_contrib_df.sum(axis='columns')
pos_contrib_df = pos_contrib_df.div(observation_total, axis='index')
pos_contrib_df.columns = [x.split('_')[1] for x in pos_contrib_df.columns]
pos_contrib_df = contrib_df[['compound_id', 'disease_id']].join(pos_contrib_df)
#pos_contrib_df = predict_df.merge(pos_contrib_df)
#pos_contrib_df.sort_values('prediction', inplace=True, ascending=True)
pos_contrib_df.head(2)
with bz2.open('./predictions/metapath-contribution.tsv.bz2', 'wt') as write_file:
pos_contrib_df.to_csv(write_file, float_format='%.4g', sep='\t', index=False)
# ## Contribution by path
predict_df = (
pandas.read_table('predictions/probabilities.tsv', low_memory=False)
# .query("prediction > 0.50") # Reduce the prediction set for easy testing
)
predict_df.head()
len(predict_df)
# +
# Create a dictionary of (compound_id, disease_id, metapath) keys to lookup untransformed DWPCs
untran_df = pandas.read_table('features/features.tsv.bz2', low_memory=False)
dwpc_melt_df = pandas.melt(untran_df, id_vars=['compound_id', 'disease_id'],
value_vars=list(pos_dwpc_coef_df.feature), var_name='metapath', value_name='dwpc')
untran_dwpc_map = dict()
for row in dwpc_melt_df.itertuples():
key = row.compound_id, row.disease_id, row.metapath
untran_dwpc_map[key] = row.dwpc
# -
# ## Prepare a list of observations
# +
obj = list()
pos_contrib_df
for i, row in predict_df.merge(pos_contrib_df).iterrows():
observation = collections.OrderedDict()
compound_id = row['compound_id']
disease_id = row['disease_id']
for key in ['compound_id', 'compound_name', 'disease_id', 'disease_name', 'category', 'status', 'prediction', 'training_prediction', 'compound_percentile', 'disease_percentile', 'n_trials']:
value = row[key]
if pandas.notnull(value):
observation[key] = value
contribs = collections.OrderedDict()
for metapath in pos_dwpc_coef_df.metapath:
percent_contrib = row[metapath]
if percent_contrib == 0 or pandas.isnull(percent_contrib):
continue
contribs[metapath] = percent_contrib
observation['metapath_contribution'] = contribs
obj.append(observation)
len(obj)
# -
predict_df.head()
# ## Load metapaths info
# +
def create_path_template(metarels):
# Create cypher query
q = '(:' + metarels[0][0] + ' {{identifier: {}}})'
for i, (source_label, target_label, rel_type, direction) in enumerate(metarels):
kwargs = {
'i': i + 1,
'rel_type': rel_type,
'target_label': ':{}'.format(target_label),
'dir0': '<-' if direction == 'backward' else '-',
'dir1': '->' if direction == 'forward' else '-',
}
q += '{dir0}[:{rel_type}]{dir1}({target_label} {{{{identifier: {{}}}}}})'.format(**kwargs)
return q
def get_paths(elem):
c_id = elem['compound_id']
d_id = elem['disease_id']
dfs = list()
for metapath, contribution in elem['metapath_contribution'].items():
untran_dwpc = untran_dwpc_map[(c_id, d_id, 'dwpc_' + metapath)]
pdp_query = metapath_to_query[metapath]
parameters = {
'source': c_id,
'target': d_id,
'dwpc': untran_dwpc,
'metapath_contribution': elem['metapath_contribution'][metapath],
}
result = session.run(pdp_query, parameters)
df = pandas.DataFrame((x.values() for x in result), columns=result.keys())
df['source_edge'] = df['nodes'].map(lambda x: '—'.join([x[0], metapath_to_source_metaedge[metapath], x[1]]))
df['target_edge'] = df['nodes'].map(lambda x: '—'.join([x[-1], metapath_to_target_metaedge[metapath], x[-2]]))
df['nodes'] = df['nodes'].map(lambda x: '—'.join(x))
df['metapath'] = metapath
dfs.append(df)
if not dfs:
return None
df = pandas.concat(dfs).sort_values('percent_of_prediction', ascending=False)
return df
def format_property(x):
if isinstance(x, int):
return str(x)
if isinstance(x, str):
return '"{}"'.format(x)
assert False
def get_summary_cypher(path_df, n_max = 5):
path_df = path_df.iloc[:n_max, :]
if path_df.empty:
return None
path_queries = list()
for i, row in enumerate(path_df.itertuples()):
path_template = metapath_to_cypher[row.metapath]
path_query = path_template.format(*map(format_property, row.node_ids))
path_query = 'MATCH p{} = {}'.format(i, path_query)
path_queries.append(path_query)
return_query = 'RETURN [{}]'.format(', '.join('p{}'.format(i) for i in range(len(path_df))))
return '\n'.join(path_queries) + '\n' + return_query
def get_directory(compound_id, disease_id):
base_path = '../../het.io-rep-data/prediction-info'
directory = os.path.join(base_path, compound_id, disease_id.replace(':', '_'))
os.makedirs(directory, exist_ok=True)
return directory
# -
config = configparser.ConfigParser()
config.read('../config.ini')
commit = config['hetnet']['integrate_commit']
url = 'https://github.com/dhimmel/integrate/raw/{}/data/metagraph.json'.format(commit)
metagraph = hetio.readwrite.read_metagraph(url)
with open('features/metapaths.json') as read_file:
metapaths = json.load(read_file)
metapath_to_cypher = dict()
metapath_to_source_metaedge = dict()
metapath_to_target_metaedge = dict()
for metapath in metapaths:
metapath['object'] = metagraph.metapath_from_abbrev(metapath['abbreviation'])
metapath['metarels'] = hetio.neo4j.metapath_to_metarels(metapath['object'])
metapath['path_template'] = create_path_template(metapath['metarels'])
abbrev = metapath['abbreviation']
metapath_to_cypher[abbrev] = metapath['path_template']
metapath_obj = metapath['object']
metapath_to_source_metaedge[abbrev] = metapath_obj[0].kind
metapath_to_target_metaedge[abbrev] = metapath_obj[-1].kind
# +
metapath_to_query = dict()
for metapath in metapaths:
dwpc_query = metapath['dwpc_query']
pdp_query = dwpc_query.split('RETURN')[0] + \
'''\
WITH
extract(n in nodes(path)| n.name) AS nodes,
extract(n in nodes(path)| n.identifier) AS node_ids,
sum(reduce(pdp = 1.0, d in degrees| pdp * d ^ -0.4)) / { dwpc } AS percent_of_DWPC
WITH
nodes, node_ids,
percent_of_DWPC,
percent_of_DWPC * { metapath_contribution } AS percent_of_prediction
RETURN nodes, percent_of_prediction, percent_of_DWPC, node_ids
'''
metapath_to_query[metapath['abbreviation']] = pdp_query
# -
driver = GraphDatabase.driver("bolt://neo4j.het.io")
session = driver.session()
# +
# %%time
for elem in obj:
directory = get_directory(elem['compound_id'], elem['disease_id'])
path = os.path.join(directory, 'info.json')
with open(path, 'wt') as write_file:
json.dump(elem, write_file, indent=2, sort_keys=True)
# Save path_df
path_df = get_paths(elem)
if path_df is None:
continue
path = os.path.join(directory, 'paths.tsv')
path_df.drop('node_ids', axis='columns').to_csv(path, sep='\t', index=False, float_format='%.3g')
# Create a cypher query with the most influential paths
path = os.path.join(directory, 'highlights.cyp')
summary_cypher = get_summary_cypher(path_df, 10)
with open(path, 'wt') as write_file:
write_file.write(summary_cypher)
# -
elem
# Close Neo4j driver session
session.close()
|
prediction/5-contribution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
from sklearn.tree import DecisionTreeClassifier
from sklearn import tree
music_data = pd.read_csv('music.csv')
X = music_data.drop(columns=['genre'])
y = music_data['genre']
model = DecisionTreeClassifier()
model.fit(X, y)
tree.export_graphviz(model, out_file='music-recommender.dot',
feature_names=['age', 'geender'], class_names=sorted(y.unique()),
label='all',
rounded=True, filled=True)
# -
music_data
|
HelloWorld.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import torch
from torch import nn
from sklearn.preprocessing import OneHotEncoder, StandardScaler
from sklearn import preprocessing
import numpy as np
import matplotlib.pyplot as plt
import scipy.io
from sklearn.metrics import euclidean_distances
from cave.cavefacade import CAVE
from ConfigSpace.util import impute_inactive_values
from ConfigSpace import CategoricalHyperparameter, UniformFloatHyperparameter, UniformIntegerHyperparameter, OrdinalHyperparameter
from cave.utils.helpers import combine_runhistories
from cave.utils.helpers import create_random_runhistories, combine_random_local, create_new_rhs # Julia BA
import torchvision
from torch.autograd import Variable
import warnings
import pickle
import collections
# +
cave = CAVE(folders=["../DataForTesting/cplex_regions200/SMAC3/run-1/smac3-output_2019-03-15_09:55:14_185212/run_1/"],
output_dir="./CAVE/Output/June/AutoEncoder/03_06_CAVE_cplex_jupyter_autoencoder/",
ta_exec_dir=["../DataForTesting/cplex_regions200/SMAC3/run-1/"],
file_format='SMAC3',
# verbose_level='DEBUG'
)
print("Finished")
# +
scenario = cave.scenario
from copy import deepcopy
configspace = deepcopy(scenario.cs)
runhistory = cave.global_original_rh
training, transform = create_random_runhistories(runhistory)
dicti = configspace._hyperparameters.items()
# +
training.save_json('training.json')
import pickle
pickle.dump(training, open("training.pkl", 'wb'), protocol=0)
pickle.load(open("training.pkl", 'rb'))
# -
# ### Review Configspace
# +
configspace.get_hyperparameters() # List of all Hyperparameter with name, type, choices/interval
configspace.get_hyperparameters()[0] # Index return hyperparamter at this place
configspace.get_hyperparameters()[0].name # Specification what is needed of this hyperparameter
configspace.get_hyperparameter_names() # List of the names of all hyperparameter
training.get_all_configs_combined() # list of all configurations
training.get_all_configs_combined()[0] # Returns the configuration at the place of index
name = configspace.get_hyperparameters()[0].name
training.get_all_configs_combined()[0].get(name) # Get value of the configuration of the defined hyperparameter
# -
# # Convert Data
#
# ## Case 1
#
# * standardize continual Data
# * replace nan with -1
# +
with warnings.catch_warnings():
warnings.simplefilter("ignore")
for hp in configspace.get_hyperparameters():
if type(hp) is CategoricalHyperparameter:
hp.choices = hp.choices + (-1234, )
values = [OneHotEncoder(categories='auto').fit((np.sort(np.array(hp.choices)).reshape((-1,1))))
if type(hp) is CategoricalHyperparameter
else (StandardScaler().fit(np.array([confi.get(hp.name) for confi in training.get_all_configs_combined()]).reshape(-1, 1))
if type(hp) in {UniformFloatHyperparameter, UniformIntegerHyperparameter, OrdinalHyperparameter}
else None)
for hp in configspace.get_hyperparameters()]
# -
for i in range(len(values)):
if type(values[i]) == StandardScaler and type(values[i]) != OneHotEncoder:
pass
elif type(values[i]) == OneHotEncoder and type(values[i]) != StandardScaler:
pass
else:
print("Fehler")
# +
config = training.get_all_configs_combined()[0]
for hp in configspace.get_hyperparameters():
if type(hp) is CategoricalHyperparameter:
print(hp.name, hp.choices)
# print(config)
# print(hp)
# OneHotEncoder(categories='auto').fit(np.vstack((np.sort(np.array(hp.choices)).reshape((-1,1)), [[-1]])))
#one_hot_encode(training.get_all_configs_combined()[0])
# -
# one hot encoding
def one_hot_encode(config):
# Code from PhMueller
# Create array with one hot encoded values
result_vec = np.array([]).reshape((-1, 1)) # .astype(object)
for i, hp in enumerate(configspace.get_hyperparameters()):
val = np.array(config.get(hp.name)).reshape((-1, 1))
# print(val)
# case if categorical
if type(values[i]) is OneHotEncoder:
if val == [[None]]:
# val = np.array(['-1']).reshape((-1, 1))
val = np.array([['-1234']])
if len(result_vec) == 0:
result_vec = values[i].transform(val).toarray() # .astype(object)
else:
result_vec = np.hstack((result_vec,
values[i].transform(val).toarray()))
# if it is continous
else:
if val == [[None]]:
if len(result_vec) == 0:
result_vec = np.array([-1234]).reshape((-1, 1))
else:
result_vec = np.hstack((result_vec, [[-1234]]))
elif len(result_vec) == 0:
result_vec = values[i].transform(val)
else:
result_vec = np.hstack((result_vec,
values[i].transform(val)))
return result_vec
for i in range(len(values)):
if i == None:
print("Error")
# +
convert_data = np.array([]).reshape((-1, 1))
for confi in range(len(training.config_ids)):
if confi % 500 == 0:
print(confi)
if len(convert_data) == 0:
convert_data = one_hot_encode(training.get_all_configs_combined()[confi])
continue
convert_data = np.vstack((convert_data, one_hot_encode(training.get_all_configs_combined()[confi])))
print(len(convert_data))
# [one_hot_encode(training.get_all_configs_combined()[confi]) for confi in range(len(training.config_ids))]
# +
convert_data_transform = np.array([]).reshape((-1, 1))
for confi in range(len(transform.config_ids)):
if confi % 10 == 0:
print(confi)
if len(convert_data_transform) == 0:
convert_data_transform = one_hot_encode(transform.get_all_configs_combined()[confi])
continue
convert_data_transform = np.vstack((convert_data_transform, one_hot_encode(transform.get_all_configs_combined()[confi])))
print(len(convert_data_transform))
# -
convert_data.shape[1] == convert_data_transform.shape[1]
np.save("convert_data.npy", convert_data)
convert_data.shape
np.load("convert_data.npy")
# # AutoEncoder
class Softmax_BA(nn.Module):
"""My own class with softmax and crossentropy to transform tensor back in original strucutre"""
__constants__ = ['dim']
def __init__(self, num_category, transform_list, confi, dim=None):
super(Softmax_BA, self).__init__()
self.num_cat = num_category
self.transform_list = transform_list
self.configspace = confi
self.dim = dim
def forward(self, x):
indexing = 0
x_ = x.clone()
softmax = nn.Softmax(dim=1)
for num in range(len(self.configspace.get_hyperparameters())):
if type(self.transform_list[num]) == OneHotEncoder:
x_[:, indexing:indexing+self.num_cat[num]] = softmax(x[:, indexing:indexing+self.num_cat[num]])
indexing += self.num_cat[num]
else:
indexing += 1
x = x_
return x# Variable(x.data, requires_grad=True)
class Autoencoder(nn.Module):
""" Our autoencoder class. """
def __init__(self, length, act_f, num_layers):
super(Autoencoder, self).__init__()
if act_f.lower() == 'relu':
self.act_f = torch.nn.ReLU()
else:
self.act_f = torch.nn.Tanh()
assert num_layers > 1
self.encoder_layer = nn.ModuleList(
[nn.Linear(int(length/(i+1)), int(length/(i+2))) for i in range(num_layers-1)]
)
self.encoder_layer.extend([nn.Linear(int(length/(num_layers)), 2)])
self.decoder_layer = nn.ModuleList(
[nn.Linear(2, int(length/(num_layers)))]
)
self.decoder_layer.extend(
[nn.Linear(int(length/(i+1)), int(length/(i))) for i in range(num_layers-1, 0, -1)]
)
def encoder(self, x):
for i, layer in enumerate(self.encoder_layer):
x = layer(x)
x = self.act_f(x) if i < len(self.encoder_layer) - 1 else x
return x
def decoder(self, x):
for i, layer in enumerate(self.decoder_layer):
x = layer(x)
x = self.act_f(x) if i < len(self.decoder_layer) - 1 else x
return x
def forward(self, x):
x = self.encoder(x)
x = self.decoder(x)
x = Softmax_BA(num_cat, values, configspace, dim=1)(x)
return x
def give_latent_image(self, x):
x = self.encoder(x)
return x
num_cat = []
for hp in configspace.get_hyperparameters():
if type(hp) == CategoricalHyperparameter:
num_cat.append(len(hp.choices))
else:
num_cat.append(False)
# +
def train(model, X_train, X_test, num_epochs, learning_rate, weight_decay=1e-5, plot_interval=10, verbose=False):
loss_history = list()
test_loss_history = list()
# criterion = loss_function()
optimizer = torch.optim.Adam(model.parameters(),
lr=learning_rate,
weight_decay=weight_decay)
for epoch in range(num_epochs):
# Get a new batch of data, 64 key-value pairs in it
ids = np.random.choice(X_train.shape[0], 64, replace=False)
X_train = X_train[ids]
# Convert to torch tensor, usually you also need to convert to float as in here.
# X_train = torch.tensor(X_train).float()
# X_test = torch.tensor(X_test).float()
# Forward. Encodes and decodes and gives us the model's prediction.
# model() actually calls 'forward()'
output = model(X_train)
output_test = model(X_test)
# Calculate loss, defined above as mean squared error
loss = loss_function(output, X_train, num_cat)
loss_test = loss_function(output_test, X_test, num_cat)
# === The backpropagation
# Reset the gradients
optimizer.zero_grad()
# Calculate new gradients with backpropagation
loss.backward()
# Tune weights accoring to optimizer (it has the learnrate and weight decay as defined above)
optimizer.step()
# To do output stuff with loss and image, we have to detach() and convert back to numpy.
loss = loss.detach().numpy()
loss_test = loss_test.detach().numpy()
# Append to loss history
loss_history.append(loss)
test_loss_history.append(loss_test)
if verbose:
print('Epoch: ' + str(epoch) + ". Train loss: " + str(loss.item()) + " Test loss: " + str(loss_test.item()))
if epoch % plot_interval == 0 and epoch != 0:
print('First 5x5 Dimension of prediction \n ')
print(X_train[0, 22:31])
print(output[0, 22:31])
print("-"*100)
"""low_dim_train = model.give_latent_image(X_train)
low_dim_test = model.give_latent_image(X_test)
low_dim_train = low_dim_train.detach().numpy()
low_dim_test = low_dim_test.detach().numpy()
plt.scatter(low_dim_train[:, 0], low_dim_train[:, 1], s=10.0,label="train points")
plt.scatter(low_dim_test[:, 0], low_dim_test[:, 1], s=10.0,label="test points")
plt.legend()
plt.show()"""
return loss_history, test_loss_history, model
def test(trained_model, X, num_plot):
""" Test our autoencoder. """
for i in range(num_plot):
"""index = 0
for cats in num_cat:
if cats == False:
index += 1
continue
plt.bar(np.arange(cats), X[i][index:index+cats], label="true", alpha=0.3)
plt.bar(np.arange(cats), output[i][index:index+cats], label="prediction", alpha=0.3)
plt.legend()
plt.show()
index += cats
print("last index true: " + str(X[i][-1]) + ", prediction: " + str(output[i][-1]))"""
ids = np.random.choice(X.shape[0], 100)
X = X[ids]
X = torch.tensor(X).float()
output = trained_model(X)
loss = loss_function(output, X, num_cat)
loss = loss.detach().numpy()
X = X.detach().numpy()
output = output.detach().numpy()
print("Input: \n %s \n Output: \n %s" % (X[:2, 15:25], output[:2, 15:25]))
print("Train loss: " + str(loss.item()))
print("-" * 10)
# +
import random
division = int(len(training.config_ids)* 0.75)
ids = np.arange(convert_data.shape[0])
np.random.shuffle(ids)
train_ids = ids[:division]
test_ids = ids[division:]
# -
def cross_entropy_one_hot(input, target):
_, labels = target.max(dim=1)
return nn.CrossEntropyLoss()(input, labels)
def loss_function(label, predition, num_category):
indexing = 0
categorical_Loss = 0
mse = nn.MSELoss()
mse_Loss = 0
for num, hp in enumerate(configspace.get_hyperparameters()):
if type(hp) == CategoricalHyperparameter:
confi_pred = predition[:, indexing:indexing+num_category[num]]
confi_lable = label[:, indexing:indexing+num_category[num]]
categorical_Loss += cross_entropy_one_hot(confi_lable, confi_pred)
indexing += num_category[num]
else:
mse_Loss += mse(label[:, indexing], predition[:, indexing])
indexing += 1
#print("MSE: %s" % mse_Loss)
#print("CE: %s" % categorical_Loss)
return mse_Loss + categorical_Loss
# +
# New model
model = Autoencoder(convert_data.shape[1], "tanh", 3)
# Train the model and return loss history
loss_history, test_loss_history, new_model = train(model,
X_train=torch.tensor(convert_data[train_ids]).float(),
X_test=torch.tensor(convert_data[test_ids]).float(),
num_epochs=1000,
learning_rate=1e-5,
weight_decay=1e-5,
plot_interval=100,
verbose=True)
# Plot the loss history. Careful: It's the train loss
plt.plot(loss_history, label="train")
plt.plot(test_loss_history, label="test")
plt.legend()
plt.show()
# -
# Print the test loss and plot some example images
test(new_model, convert_data_transform, num_plot=100)
# +
X = torch.tensor(convert_data).float()
Z = torch.tensor(convert_data_transform).float()
low_dim_rand = model.give_latent_image(X)
low_dim_rand = low_dim_rand.detach().numpy()
low_dim_local = model.give_latent_image(Z)
low_dim_local = low_dim_local.detach().numpy()
plt.scatter(low_dim_rand[:, 0], low_dim_rand[:, 1], s=10.0,label="random points")
plt.scatter(low_dim_local[:, 0], low_dim_local[:, 1], s=10.0,label="random points")
plt.show()
# +
from ConfigSpace.read_and_write import json
with open('./config_space.json', 'w') as f:
f.write(json.write(configspace))
# +
X = torch.tensor(convert_data).float()
low_dim = model.give_latent_image(X)
low_dim = low_dim.detach().numpy()
# +
plt.scatter(low_dim[:, 0], low_dim[:, 1],) # label="local points")
# plt.legend()
plt.show()
# -
def calculate_costvalue(dists, red_dists):
"""
Helpfunction to calculate the costvalue to test how big the difference of distance is in the embedding
and original space.
Parameters
----------
dists: np.array, shape(n_samples, n_samples)
Matrix of the distances in the original space.
red_dists: np.array, shape(n_samples, k_dimensions)
Koordinates o
Returns
--------
costvalue: float
Costvalue of the distances of the two spaces.
costvalues = sum_i sum_j=i+1 (distance_low_space_ij - distance_high_space_ij)
"""
n_conf = dists.shape[0]
low_dists = euclidean_distances(red_dists)
costvalue = []
mean_actual = []
for i in range(n_conf - 1):
for j in range(i+1, n_conf):
costvalue.append((dists[i][j] - low_dists[i][j])**2)
mean_actual.append(low_dists[i][j])
mean_actual_value = sum(mean_actual) / len(mean_actual)
actual = [(mean_actual_value - dif)**2 for dif in mean_actual]
pred_actual = sum(costvalue)
rse = pred_actual / sum(actual)
costvalue = sum(costvalue) / len(costvalue)
print("costvalue")
print(costvalue)
print("rse")
print(rse)
return costvalue
# +
# Softmax
m = nn.Softmax(dim=1)
test = torch.randn(2, 3)
output = m(test)
print(test)
print(output)
# -
loss = nn.CrossEntropyLoss()
input = torch.randn(4, 1, requires_grad=True)
target = torch.empty(4, dtype=torch.long).random_(1)
output = loss(input, target)
output.backward()
input
torch.empty(4, dtype=torch.long).random_(1)
image = output.detach().numpy()
# image = image[0].reshape(image.shape[1])
plt.imshow(image)
# # Misc
# One-hot-encoder version with -1 for each one-hot dimension → nan by categorical with 4 choices is [-1, -1, -1, -1]
# one hot encoding
def one_hot_encode(config):
# Code from PhMueller
# Create array with one hot encoded values
result_vec = np.array([]).reshape((-1, 1)) # .astype(object)
for i, name in enumerate(configspace.get_hyperparameter_names()):
val = np.array(config.get(name)).reshape((-1, 1))
# Case if this value is not given in the configuration
if val == [[None]]:
# Test, maybe this is not working
if len(result_vec) == 0 and type(configspace.get_hyperparameter(name)) == CategoricalHyperparameter:
cats = len(configspace.get_hyperparameters()[i].choices)
result_vec = np.array([-1] * cats).reshape((1, len(np.array([-1] * cats))))
elif len(result_vec) == 0 and type(configspace.get_hyperparameter(name)) != CategoricalHyperparameter:
result_vec = np.array([-1]).reshape((-1, 1))
elif len(result_vec) > 0 and type(configspace.get_hyperparameter(name)) == CategoricalHyperparameter:
cats = len(configspace.get_hyperparameters()[i].choices)
result_vec = np.hstack((result_vec, np.array([-1] * cats).reshape((1, len(np.array([-1] * cats))))))
else:
result_vec = np.hstack((result_vec, [[-1]]))
# case if categorical
elif type(values[i]) is OneHotEncoder:
if len(result_vec) == 0:
result_vec = values[i].transform(val).toarray() # .astype(object)
else:
result_vec = np.hstack((result_vec,
values[i].transform(val).toarray()))
# if it is one
else:
if len(result_vec) == 0:
result_vec = values[i].transform(val)
else:
result_vec = np.hstack((result_vec,
values[i].transform(val)))
return result_vec
# +
oe = OneHotEncoder(categories='auto').fit(np.array([1,2,'-1']).reshape((-1,1)))
oe.categories_
oe.transform(np.array(1).reshape((-1, 1))).toarray()
# -
|
AutoEncoder_CAVE_new.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.9 64-bit (''3dunet'': conda)'
# language: python
# name: python3
# ---
import h5py
import matplotlib.pyplot as plt
def describe(im):
print(f"\tdtype: {im.dtype}")
print(f"\tshape: {im.shape}")
print(f"\tmin: {im.min()}")
print(f"\tmean: {im.mean()}")
print(f"\tmax: {im.max()}")
file_path = "/scratch/ottosson/datasets/FM_SMALL/plantseg_training/0h_h5.h5"
with h5py.File(file_path, 'r') as f:
print(f.keys())
raw = f['raw'][:]
label = f['label'][:]
print("raw")
describe(raw)
print("label")
describe(label)
z_slice = 4
fig, axs = plt.subplots(1,2)
axs[0].imshow(raw[z_slice])
axs[1].imshow(label[z_slice])
|
src/datasets/setup_datasets/FM/create_plantseg_training/view_result.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="4f3CKqFUqL2-" slideshow={"slide_type": "slide"}
# # Hyperparameter tuning with Cloud ML Engine
# -
# **Learning Objectives:**
# * Improve the accuracy of a model by hyperparameter tuning
import os
PROJECT = 'cloud-training-demos' # REPLACE WITH YOUR PROJECT ID
BUCKET = 'cloud-training-demos-ml' # REPLACE WITH YOUR BUCKET NAME
REGION = 'us-central1' # REPLACE WITH YOUR BUCKET REGION e.g. us-central1
# for bash
os.environ['PROJECT'] = PROJECT
os.environ['BUCKET'] = BUCKET
os.environ['REGION'] = REGION
os.environ['TFVERSION'] = '1.8' # Tensorflow version
# + language="bash"
# gcloud config set project $PROJECT
# gcloud config set compute/region $REGION
# + [markdown] colab_type="text" id="6TjLjL9IU80G"
# ## Create command-line program
#
# In order to submit to Cloud ML Engine, we need to create a distributed training program. Let's convert our housing example to fit that paradigm, using the Estimators API.
# + language="bash"
# rm -rf house_prediction_module
# mkdir house_prediction_module
# mkdir house_prediction_module/trainer
# touch house_prediction_module/trainer/__init__.py
# +
# %%writefile house_prediction_module/trainer/task.py
import argparse
import os
import json
import shutil
from . import model
if __name__ == '__main__' and "get_ipython" not in dir():
parser = argparse.ArgumentParser()
# TODO: Add learning_rate and batch_size as command line args
parser.add_argument(
'--output_dir',
help = 'GCS location to write checkpoints and export models.',
required = True
)
parser.add_argument(
'--job-dir',
help = 'this model ignores this field, but it is required by gcloud',
default = 'junk'
)
args = parser.parse_args()
arguments = args.__dict__
# Unused args provided by service
arguments.pop('job_dir', None)
arguments.pop('job-dir', None)
# Append trial_id to path if we are doing hptuning
# This code can be removed if you are not using hyperparameter tuning
arguments['output_dir'] = os.path.join(
arguments['output_dir'],
json.loads(
os.environ.get('TF_CONFIG', '{}')
).get('task', {}).get('trial', '')
)
# Run the training
shutil.rmtree(arguments['output_dir'], ignore_errors=True) # start fresh each time
# Pass the command line arguments to our model's train_and_evaluate function
model.train_and_evaluate(arguments)
# +
# %%writefile house_prediction_module/trainer/model.py
import numpy as np
import pandas as pd
import tensorflow as tf
tf.logging.set_verbosity(tf.logging.INFO)
# Read dataset and split into train and eval
df = pd.read_csv("https://storage.googleapis.com/ml_universities/california_housing_train.csv", sep = ",")
df['num_rooms'] = df['total_rooms'] / df['households']
np.random.seed(seed = 1) #makes split reproducible
msk = np.random.rand(len(df)) < 0.8
traindf = df[msk]
evaldf = df[~msk]
# Train and eval input functions
SCALE = 100000
def train_input_fn(df, batch_size):
return tf.estimator.inputs.pandas_input_fn(x = traindf[["num_rooms"]],
y = traindf["median_house_value"] / SCALE, # note the scaling
num_epochs = None,
batch_size = batch_size, # note the batch size
shuffle = True)
def eval_input_fn(df, batch_size):
return tf.estimator.inputs.pandas_input_fn(x = evaldf[["num_rooms"]],
y = evaldf["median_house_value"] / SCALE, # note the scaling
num_epochs = 1,
batch_size = batch_size,
shuffle = False)
# Define feature columns
features = [tf.feature_column.numeric_column('num_rooms')]
def train_and_evaluate(args):
# Compute appropriate number of steps
num_steps = (len(traindf) / args['batch_size']) / args['learning_rate'] # if learning_rate=0.01, hundred epochs
# Create custom optimizer
myopt = tf.train.FtrlOptimizer(learning_rate = args['learning_rate']) # note the learning rate
# Create rest of the estimator as usual
estimator = tf.estimator.LinearRegressor(model_dir = args['output_dir'],
feature_columns = features,
optimizer = myopt)
#Add rmse evaluation metric
def rmse(labels, predictions):
pred_values = tf.cast(predictions['predictions'], tf.float64)
return {'rmse': tf.metrics.root_mean_squared_error(labels * SCALE, pred_values * SCALE)}
estimator = tf.contrib.estimator.add_metrics(estimator, rmse)
train_spec = tf.estimator.TrainSpec(input_fn = train_input_fn(df = traindf, batch_size = args['batch_size']),
max_steps = num_steps)
eval_spec = tf.estimator.EvalSpec(input_fn = eval_input_fn(df = evaldf, batch_size = len(evaldf)),
steps = None)
tf.estimator.train_and_evaluate(estimator, train_spec, eval_spec)
# + language="bash"
# rm -rf house_trained
# export PYTHONPATH=${PYTHONPATH}:${PWD}/house_prediction_module
# gcloud ml-engine local train \
# --module-name=trainer.task \
# --job-dir=house_trained \
# --package-path=$(pwd)/trainer \
# -- \
# --batch_size=30 \
# --learning_rate=0.02 \
# --output_dir=house_trained
# -
# # Create hyperparam.yaml
# %%writefile hyperparam.yaml
trainingInput:
hyperparameters:
goal: MINIMIZE
maxTrials: 5
maxParallelTrials: 1
hyperparameterMetricTag: rmse
params:
- parameterName: batch_size
type: INTEGER
minValue: 8
maxValue: 64
scaleType: UNIT_LINEAR_SCALE
- parameterName: learning_rate
type: DOUBLE
minValue: 0.01
maxValue: 0.1
scaleType: UNIT_LOG_SCALE
# + language="bash"
# OUTDIR=gs://${BUCKET}/house_trained # CHANGE bucket name appropriately
# gsutil rm -rf $OUTDIR
# export PYTHONPATH=${PYTHONPATH}:${PWD}/house_prediction_module
# gcloud ml-engine jobs submit training house_$(date -u +%y%m%d_%H%M%S) \
# --config=hyperparam.yaml \
# --module-name=trainer.task \
# --package-path=$(pwd)/house_prediction_module/trainer \
# --job-dir=$OUTDIR \
# --runtime-version=$TFVERSION \
# -- \
# --output_dir=$OUTDIR
# -
# !gcloud ml-engine jobs describe house_180912_195904 # CHANGE jobId appropriately
# ## Challenge exercise
# Add a few engineered features to the housing model, and use hyperparameter tuning to choose which set of features the model uses.
#
# <p>
# Copyright 2018 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
|
courses/machine_learning/deepdive/05_artandscience/labs/b_hyperparam.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# name: python3
# ---
from Integrais import Integrais as it
from EDO import EDO
from sympy import var, Lambda, cos, sin, tan, exp, log, ln, sqrt, solve
import pandas as pd
import numpy as np
x = var('x')
y = var('y')
# +
# Questão 01
# Letra A
x0 = 1
y0 = -1
dy = Lambda((x, y), 1/x**2 - y/x - y**2)
print('f(x, y) = ', dy(x,y))
I = (1, 2)
N = 10
Y = EDO(dy, y0, x0)
y1, y_steps, x_steps = Y.euler(I, N, dp=4)
print(y1)
# Letra B
print(f'\nSolução Exata: y(2) = {-0.5}')
print(f'Solução Aproximada: y(2) = {y1}')
print(f'Erro Absoluto: {abs(-0.5 - y1):.4f}')
# -
# Questão 02
x0 = 1
y0 = 2
dy = Lambda((x, y), -x * y**2)
print('f(x, y) = ', dy(x,y))
I = (1, 1.5)
N = 5
Y = EDO(dy, y0, x0)
y1, y_steps, x_steps = Y.euler(I, N, dp=4)
print(f'Solução Aproximada: y(1.5) = {y1}')
# +
# Questão 03
# Letra A - Euler
x0 = 0
y0 = 1
dy = Lambda((x, y), x * y ** (1/3))
print('f(x, y) = ', dy(x,y))
I = (0, 2)
N = 5
Y = EDO(dy, y0, x0)
y1_euler, y_steps_euler, x_steps = Y.euler(I, N, dp=4)
print('Metodo de Euler')
print(f'Solução aproximada: y(2) = {y1_euler}')
# Letra B - Euler Aprimorado
y1_aprimorado, y_steps_apri, x_steps = Y.runge_kutta2(I, N, dp=4)
print('\n Método de Euler Aprimorado')
print(f'Solução aproximada: y(2) = {y1_aprimorado}')
# Letra C - Erro e Tabela
y_ex = Lambda(x, ((x**2 + 3) / 3)**(3/2))
print('f(y) = ', y_ex(x))
y_ex_values = [np.round(float(y_ex(value)), 4) for value in x_steps]
comparacao = {'Euler': y_steps_euler,
'Ruge-Kutta 2': y_steps_apri,
'Exato': y_ex_values}
comparacao_df = pd.DataFrame(comparacao, index=x_steps)
comparacao_df
# +
# Questão 04
x0 = 1
y0 = 2
dy = Lambda((x, y), -x*y**2)
print('f(x, y) = ', dy(x,y))
N = 2
I = (0, 1.5)
Y = EDO(dy, y0, x0)
y1, y_steps, x_steps = Y.runge_kutta4(I, N, dp=4)
print(f'Solução aproximada: y(1.5) = {y1}')
# -
# Questão 05
x0 = 0
y0 = 1
dy = Lambda((x, y), y*cos(x))
print('f(x, y) = ', dy(x,y))
N = 3
I = (0, 0.6)
Y = EDO(dy, y0, x0)
y1, y_steps, x_steps = Y.runge_kutta4(I, N, dp=4)
print(f'Solução aproximada: y(0.6) = {y1}')
# Questão 06
x0 = 0
y0 = 1000
dy = Lambda((x, y), -0.1 * y)
print('f(x, y) = ', dy(x,y))
N = 2000
I = (0, 2)
Y = EDO(dy, y0, x0)
y1, y_steps, x_steps = Y.euler(I, N, dp=4)
k = 1
while y1 > 500:
y1, y_steps, x_steps = Y.euler((I[0], I[1]+k), N, dp=4)
k += 1
k -= 1
print(f'Solução aproximada: y({k+I[1]}) = {y1}')
# +
# Questão 7
# dy/dx = r(x) y**2 + a(x)y + b(x)
# como queremos a resposta para x = 0.2
# valor aproximado
x0 = 0
y0 = 3
dy = Lambda((x, y), 1)
print('f(x, y) = ', dy(x,y))
N = 2
I = (0, 0.2)
Y = EDO(dy, y0, x0)
y1, y_steps, x_steps = Y.runge_kutta4(I, N, dp=4)
print(f'Solução aproximada: y({I[1]}) = {y1}')
# -
|
MAT 271/listaV.ipynb
|
# ##### Copyright 2020 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # nqueens
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/nqueens.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/nqueens.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
# Copyright 2010 <NAME> <EMAIL>
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
"""
n-queens problem in Google CP Solver.
N queens problem.
This model was created by <NAME> (<EMAIL>)
Also see my other Google CP Solver models:
http://www.hakank.org/google_or_tools/
"""
from ortools.constraint_solver import pywrapcp
# Create the solver.
solver = pywrapcp.Solver("n-queens")
#
# data
#
# n = 8 # size of board (n x n)
# declare variables
q = [solver.IntVar(0, n - 1, "x%i" % i) for i in range(n)]
#
# constraints
#
solver.Add(solver.AllDifferent(q))
for i in range(n):
for j in range(i):
solver.Add(q[i] != q[j])
solver.Add(q[i] + i != q[j] + j)
solver.Add(q[i] - i != q[j] - j)
# for i in range(n):
# for j in range(i):
# solver.Add(abs(q[i]-q[j]) != abs(i-j))
# symmetry breaking
# solver.Add(q[0] == 0)
#
# solution and search
#
solution = solver.Assignment()
solution.Add([q[i] for i in range(n)])
collector = solver.AllSolutionCollector(solution)
# collector = solver.FirstSolutionCollector(solution)
# search_log = solver.SearchLog(100, x[0])
solver.Solve(
solver.Phase([q[i] for i in range(n)], solver.INT_VAR_SIMPLE,
solver.ASSIGN_MIN_VALUE), [collector])
num_solutions = collector.SolutionCount()
print("num_solutions: ", num_solutions)
if num_solutions > 0:
for s in range(num_solutions):
qval = [collector.Value(s, q[i]) for i in range(n)]
print("q:", qval)
for i in range(n):
for j in range(n):
if qval[i] == j:
print("Q", end=" ")
else:
print("_", end=" ")
print()
print()
print()
print("num_solutions:", num_solutions)
print("failures:", solver.Failures())
print("branches:", solver.Branches())
print("WallTime:", solver.WallTime())
else:
print("No solutions found")
n = 8
|
examples/notebook/contrib/nqueens.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
class InsufficientBalance(Exception):
def __init__(self, *args):
if args:
self.message = args[0]
self.deficit = args[1]
else:
self.message = None
self.deficit = 0.0
def __str__(self):
return f"InsufficientBalance [message: {self.message}, deficit: {self.deficit}]"
class Account:
def __init__(self, iban, balance=50.0):
self.iban = iban
self.balance = balance
def deposit(self, amount):
if amount <= 0: # validation
raise ValueError(f"Amount ({amount}) must be positive.")
self.balance += amount # business logic
def withdraw(self, amount):
print("Account::withdraw")
if amount <= 0: # validation
raise ValueError(f"Amount ({amount}) must be positive.")
if amount > self.balance: # business rule
message = f"Balance {self.balance} is less than amount {amount}"
deficit = amount - self.balance
raise InsufficientBalance(message,deficit)
self.balance -= amount
def __str__(self):
return f"Account [iban: {self.iban}, balance: {self.balance}]"
try:
acc1 = Account(iban="tr1", balance=100000.0)
print(acc1)
acc1.withdraw(75000.0) # withdraw(acc1,75000.0 )
print(acc1)
acc1.deposit(5000.0)
print(acc1)
acc1.withdraw(40000)
print(acc1)
except ValueError as err:
print(err)
except InsufficientBalance as err:
print(err)
# Account -> super-class / Base Class
# CheckingAccount -> sub-class / Derived clas
class CheckingAccount(Account):
def __init__(self,iban,balance,overdraft_amount=500):
super().__init__(iban,balance) # Account's Constructor
self.overdraft_amount = overdraft_amount
def withdraw(self, amount): # overriding
print("CheckingAccount::withdraw")
if amount <= 0: # validation
raise ValueError(f"Amount ({amount}) must be positive.")
if amount > (self.balance+self.overdraft_amount): # business rule
message = f"Balance {self.balance} is less than amount {amount}"
deficit = amount - self.balance - self.overdraft_amount
raise InsufficientBalance(message,deficit)
self.balance -= amount
# super().withdraw(amount)
def __str__(self):
return f"CheckingAccount [iban: {self.iban}, balance: {self.balance}]"
try:
acc1 = CheckingAccount(iban="tr1", balance=1000.0)
print(acc1)
acc1.withdraw(750.0) # withdraw(acc1,750.0 )
print(acc1)
acc1.deposit(50.0)
print(acc1)
acc1.withdraw(801.0)
print(acc1)
except ValueError as err:
print(err)
except InsufficientBalance as err:
print(err)
acc2 = Account("TR1", 2000)
acc3 = CheckingAccount("TR2", 3000, 1000)
isinstance(acc2, Account)
isinstance(acc2, CheckingAccount)
isinstance(acc3, Account)
isinstance(acc3, CheckingAccount)
accounts = [
Account("TR1", 2000),
CheckingAccount("TR2", 3000, 1000),
Account("TR3", 4000),
CheckingAccount("TR4", 5000, 2000),
]
for acc in accounts:
print(isinstance(acc,Account))
for acc in accounts:
acc.withdraw(50)
def get_total_balance(accounts):
total_balance = 0
for acc in accounts:
total_balance += acc.balance
return total_balance
print(f"{get_total_balance(accounts):.2f}")
class Pet: # interface/abstract class
def play(self):
pass
def setName(self):
pass
class Cat(Pet):
def __init__(self, name= "Garfield"):
self.name = name
def play(self): # override
print(f"{self.name} is playing now...")
def setName(self): # override
return self.name
cat1 = Cat("Tekir")
cat1.play()
# +
# operator overloading
# # +, -, /, *, %, ... -> int, str, float
# Class -> Object -> operator
# -
class fraction:
def __init__(self, pay, payda):
self.pay = pay
self.payda = payda
def __add__(self,right):
new_pay = self.pay * right.payda + self.payda * right.pay
return fraction( new_pay , self.payda * right.payda)
def __truediv__(self,right): # /
print("/")
new_pay = self.pay * right.payda
new_payda = self.payda * right.pay
return fraction( new_pay , new_payda)
def __floordiv__(self,right): # //
print("//")
new_pay = self.pay * right.payda
new_payda = self.payda * right.pay
return fraction( new_pay , new_payda)
def __str__(self):
return f"{self.pay}/{self.payda}"
f1 = fraction(3,5) # 3 / 5
f2 = fraction(1,2) # 1 / 2
f3 = f1 + f2 + f1 # f3 = f1.add(f2).add(f1)
f4 = f1 / f2
f5 = f1 // f2
print(f1)
print(f2)
print(f3)
print(f4)
print(f5)
|
module08-class.and.oop.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:ds]
# language: python
# name: conda-env-ds-py
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# <img src="https://upload.wikimedia.org/wikipedia/commons/4/47/Logo_UTFSM.png" width="200" alt="utfsm-logo" align="left"/>
#
# # MAT281
# ### Aplicaciones de la Matemática en la Ingeniería
# + [markdown] slideshow={"slide_type": "slide"}
# ## Módulo 02
# ## Clase 03: Combinando Datos
# + [markdown] slideshow={"slide_type": "slide"}
# ## Objetivos
#
# * Añadir nuevas columnas a un dataframe.
# * Combinar distintos dataframes según reglas.
# + [markdown] slideshow={"slide_type": "subslide"}
# ## Contenidos
# * [Motivación](#motivation)
# * [Nuevas columnas](#assign)
# * [Concatenar](#concat)
# * [Unir](#merge)
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='motivation'></a>
# ## Motivación
#
# ¿Te imaginas como las grandes compañías o gobiernos almacenan sus datos? No, no es en un excel gigante en un pendrive.
# + [markdown] slideshow={"slide_type": "slide"}
# ### Base de Datos
#
# Una __Base de Datos__ es un conjunto de datos almacenados en una computadora (generalmente un servidor). Estos datos poseen una estructura con tal que sean de fácil acceso.
# -
# ### Base de Datos Relacional
#
# Es el tipo de base de datos más ampliamente utilizado, aunque existen otros tipos de bases de datos para fines específicos. Utiliza una estructura tal que es posible identificar y acceder a datos relacionados entre si. Generalmente una base de datos relacional está organizada en __tablas__.
#
# Las tablas están conformadas de filas y columnas. Cada columna posee un nombre y tiene un tipo de dato específico, mientras que las filas son registros almacenados.
#
# Por ejemplo, la siguiente tabla tiene tres columnas y cuatro registros. En particular, la columna ```age``` tiene tipo ```INTEGER``` y las otras dos tipo ```STRING```.
#
# 
#
# __¿Este formato de datos te parece familar?__
# ### ¿Qué es SQL?
#
# Sus siglas significan _Structured Query Language_ (Lenguaje de Consulta Estructurada) es un lenguaje de programación utilizado para comunicarse con datos almacenados en un Sistema de Gestión de Bases de Datos Relacionales (_Relational Database Management System_ o RDBMS). Posee una sintaxis muy similar al idioma inglés, con lo cual se hace relativamente fácil de escribir, leer e interpretar.
#
# Hay distintos RDBMS entre los cuales la sintaxis de SQL difiere ligeramente. Los más populares son:
#
# - SQLite
# - MySQL / MariaDB
# - PostgreSQL
# - Oracle DB
# - SQL Server
# ### ¿Y esto en qué afecta a un matemático?
#
# En una empresa de tecnología hay cargos especialmente destinados a todo lo que tenga que ver con bases de datos, por ejemplo: creación, mantención, actualización, obtención de datos, transformación, seguridad y un largo etc.
#
# Los matemáticos en la industria suelen tener cargos como _Data Scientist_, _Data Analyst_, _Data Statistician_, _Data X_ (reemplace _X_ con tal de formar un cargo que quede bien en Linkedin), en donde lo importante es otorgar valor a estos datos. Por ende, lo mínimo que deben satisfacer es:
#
# - Entendimiento casi total del modelo de datos (tablas, relaciones, tipos, etc.)
# - Seleccionar datos a medida (_queries_).
# ### Modelo de datos
#
# Es la forma en que se organizan los datos. En las bases de datos incluso es posible conocer las relaciones entre tablas. A menudo se presentan gráficamente como en la imagen de abajo (esta será la base de datos que utilizaremos en los ejericios del día de
#
# 
# Esta base de datos se conoce con el nombre de _**chinook database**_. La descripción y las imágenes se pueden encontrar [aquí](http://www.sqlitetutorial.net/sqlite-sample-database/).
#
# En la figura anterior, existen algunas columnas _especiales_ con una llave al lado de su nombre. ¿Qué crees que significan?
#
# Las 11 tablas se definen de la siguiente forma (en inglés):
#
# - ```employees``` table stores employees data such as employee id, last name, first name, etc. It also has a field named ReportsTo to specify who reports to whom.
# - ```customers``` table stores customers data.
# - ```invoices``` & ```invoice_items``` tables: these two tables store invoice data. The ```invoices``` table stores invoice header data and the ```invoice_items``` table stores the invoice line items data.
# - ```artists``` table stores artists data. It is a simple table that contains only artist id and name.
# - ```albums``` table stores data about a list of tracks. Each album belongs to one artist. However, one artist may have multiple albums.
# - ```media_types``` table stores media types such as MPEG audio file, ACC audio file, etc.
# - ```genres``` table stores music types such as rock, jazz, metal, etc.
# - ```tracks``` table store the data of songs. Each track belongs to one album.
# - ```playlists``` & ```playlist_track tables```: ```playlists``` table store data about playlists. Each playlist contains a list of tracks. Each track may belong to multiple playlists. The relationship between the ```playlists``` table and ```tracks``` table is many-to-many. The ```playlist_track``` table is used to reflect this relationship.
# ### Manos a la obra!
import os
import pandas as pd
import sqlite3
# En la carpeta data se encuentra el archivo chinook.db, que es básicamente una base de datos sql. Definiremos una simple función con tal de recibir una _query_ en formato ```str``` de ```python``` y retorne el resultado de la _query_ en un dataframe de pandas.
def chinook_query(query):
# Crear un conector
conn = sqlite3.connect(os.path.join('data', 'chinook.db'))
# Retorna un dataframe
return pd.read_sql_query(query, con=conn)
# Ver todas las tablas de la base de datos
chinook_query(
"""
SELECT name
FROM sqlite_master
WHERE type='table'
"""
)
# En el ejemplo anterior resulta muy importante no mezclar tipos de comillas distintos. Si se define un string comenzando y terminando con `"` todas las comillas a usar en el interior deben de ser `'`, o viceversa.
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='assign'></a>
# ## Nuevas Columnas
#
# Es usual que desde una misma tabla/dataframe se quiera agregar columnas dependiendo de ciertas reglas.
# -
# Veamos la tabla de empleados para motivar los ejemplos.
employees = chinook_query("select * from employees")
employees.head()
#
# __Ejemplo:__ Crear una nueva columna que sea `Title - FirstName Lastname`
# Si fuera un solo empleado la tarea es sencilla
title, fname, lname = employees.loc[0, ["Title", "FirstName", "Lastname"]].values
print(f"{title} - {fname} {lname}")
# __Idea:__ Iterar por cada fila
for idx, row in employees.iterrows():
print(f"idx is: {idx} with type {type(idx)}\n")
print(f"row is: \n{row}\nwith type {type(row)}\n")
break
# %%timeit
for idx, row in employees.iterrows():
full_emp = f"{row['Title']} - {row['FirstName']} {row['LastName']}"
employees.loc[idx, "full_employee"] = full_emp
employees.head()
# Una mejor forma es utilizar `apply`
employees = chinook_query("select * from employees")
employees.apply(lambda row: f"{row['Title']} - {row['FirstName']} {row['LastName']}", axis=1)
# Para asignarlo basta con:
employees["full_employee"] = employees.apply(lambda row: f"{row['Title']} - {row['FirstName']} {row['LastName']}", axis=1)
employees.head()
# Midamos cuanto demora la ejecución:
# %%timeit
employees["full_employee"] = employees.apply(lambda row: f"{row['Title']} - {row['FirstName']} {row['LastName']}", axis=1)
# Algo así como un tercio del tiempo. ¿Bien o no?
# Si estás en medio de una concatenación de métodos el método `assign` es genial. Al principio puede confundir pero básicamente asigna nuevas columnas sin necesidad del paso anterior. Usualmente yo las ocupo así:
employees.assign(
full_employee=lambda x: x.apply(lambda row: f"{row['Title']} - {row['FirstName']} {row['LastName']}", axis=1)
).head()
# Pero incluso puedes operar entre series, y como los dataframes se componen de series todo es más fácil!
employees["Title"] + " - " + employees["FirstName"] + " " + employees["LastName"]
# %%timeit
employees["full_employee"] = employees["Title"] + " - " + employees["FirstName"] + " " + employees["LastName"]
# Pero más elegante y compatible con métodos concatenados:
# %%timeit
employees.assign(
full_employee=lambda x: x["Title"] + " - " + x["FirstName"] + " " + x["LastName"]
)
# La diferencia de tiempo es despreciable con este dataset, pero la elegancia es superior.
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='concat'></a>
# ## Concatenar
#
# Imagina que tienes varias tablas con las mismas columnas y quieres unirlas en una grande. Pandas posee la función `concat` para esta labor.
# -
# <img src="https://pandas.pydata.org/pandas-docs/stable/_images/merging_concat_basic.png" width="360" height="240" align="center"/>
# +
df1 = pd.DataFrame({'A': ['A0', 'A1', 'A2', 'A3'],
'B': ['B0', 'B1', 'B2', 'B3'],
'C': ['C0', 'C1', 'C2', 'C3'],
'D': ['D0', 'D1', 'D2', 'D3']},
index=[0, 1, 2, 3])
df2 = pd.DataFrame({'A': ['A4', 'A5', 'A6', 'A7'],
'B': ['B4', 'B5', 'B6', 'B7'],
'C': ['C4', 'C5', 'C6', 'C7'],
'D': ['D4', 'D5', 'D6', 'D7']},
index=[4, 5, 6, 7])
df3 = pd.DataFrame({'A': ['A8', 'A9', 'A10', 'A11'],
'B': ['B8', 'B9', 'B10', 'B11'],
'C': ['C8', 'C9', 'C10', 'C11'],
'D': ['D8', 'D9', 'D10', 'D11']},
index=[8, 9, 10, 11])
pd.concat([df1, df2, df3])
# -
# Un __método__ similar es `.append()`, el cual concatena a lo largo del axis=0, es decir, a través de los índices.
df1.append([df2, df3])
# Si los dataframes tienen distintas columnas no es impedimento para concatenarlas.
# <img src="https://pandas.pydata.org/pandas-docs/stable/_images/merging_append2.png" width="360" height="480" align="center"/>
# +
df4 = pd.DataFrame({'B': ['B2', 'B3', 'B6', 'B7'],
'D': ['D2', 'D3', 'D6', 'D7'],
'F': ['F2', 'F3', 'F6', 'F7']},
index=[2, 3, 6, 7])
pd.concat([df1, df4], axis=0, sort=False)
# -
# ¿Quieres que se agreguen columnas dependiendo de los index que tienen cada dataframe?
# <img src="https://pandas.pydata.org/pandas-docs/stable/_images/merging_concat_axis1.png" width="540" height="480" align="center"/>
pd.concat([df1, df4], axis=1, sort=False)
# + [markdown] slideshow={"slide_type": "slide"}
# <a id='merge'></a>
# ## Unir
#
# En pandas es posible unir dos tablas sin la necesidad de iterar fila por fila. La funcionalidad la entrega el método `merge`.
#
# Ejemplo: En la base datos chinook existen las tablas `albums` y `artists`. ¿Cómo agregar el nombre del artista a la tabla de álbumes?
# +
artists = chinook_query("select * from artists")
albums = chinook_query("select * from albums")
display(artists.head())
display(albums.head())
# -
# Por el modelo de datoa anterior sabemos que ambas tablas están relacionadas a través de la columna `ArtistId`. Iterando haríamos algo así:
# %%timeit
for idx, row in artists.iterrows():
artist_name = artists.loc[lambda x: x["ArtistId"] == row["ArtistId"], "Name"].iloc[0] # Acceder al string
albums.loc[idx, "ArtistName"] = artist_name
albums = chinook_query("select * from albums")
# %%timeit
albums_merge = albums.merge(artists, how="left", on="ArtistId")
albums.merge(artists, how="left", on="ArtistId").head()
# ### Tipo de merge
#
# El ejemplo anterior utiliza un ```left join```. Los cuatro tipos de cruces más comunes:
#
# - ```inner```: (_default_) Retorna aquellos registros donde los valors de columnas utilizadas para los cruces se encuentran en ambas tablas.
# - ```left```: Retorna todos los registros de la tabla colocada a la izquierda, aunque no tengan correspondencia en la tabla de la derecha.
# - ```right```: Retorna todos los registros de la tabla colocada a la derecha, aunque no tengan correspondencia en la tabla de la izquierda.
# - ```outer```: Retorna todos los valores de ambas tablas, tengan correspondencia o no.
#
# La siguiente imagen explica el resultado que se obtiene con los distintos tipos de cruces.
#
# 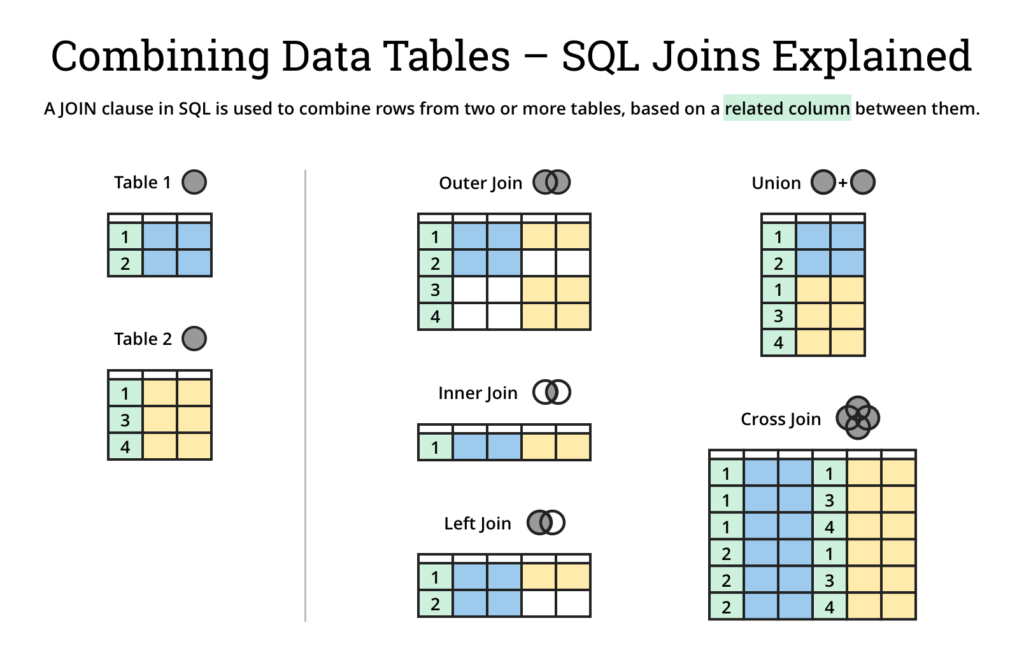
# 
# ### Ejemplitos
# +
left = pd.DataFrame({'key1': ['K0', 'K0', 'K1', 'K2'],
'key2': ['K0', 'K1', 'K0', 'K1'],
'A': ['A0', 'A1', 'A2', 'A3'],
'B': ['B0', 'B1', 'B2', 'B3']})
right = pd.DataFrame({'key1': ['K0', 'K1', 'K1', 'K2'],
'key2': ['K0', 'K0', 'K0', 'K0'],
'C': ['C0', 'C1', 'C2', 'C3'],
'D': ['D0', 'D1', 'D2', 'D3']})
# -
# #### Left Merge
#
# <img src="https://pandas.pydata.org/pandas-docs/stable/_images/merging_merge_on_key_left.png" width="540" height="480" align="center"/>
pd.merge(left, right, on=['key1', 'key2'])
# #### Right Merge
# <img src="https://pandas.pydata.org/pandas-docs/stable/_images/merging_merge_on_key_right.png" width="540" height="480" align="center"/>
pd.merge(left, right, how='right', on=['key1', 'key2'])
# #### Outer Merge
#
# <img src="https://pandas.pydata.org/pandas-docs/stable/_images/merging_merge_on_key_outer.png" width="540" height="480" align="center"/>
pd.merge(left, right, how='outer', on=['key1', 'key2'])
# #### Inner Merge
# <img src="https://pandas.pydata.org/pandas-docs/stable/_images/merging_merge_on_key_inner.png" width="540" height="480" align="center"/>
pd.merge(left, right, how='inner', on=['key1', 'key2'])
# ### Problemas de llaves duplicadas
#
# Cuando se quiere realizar el cruce de dos tablas, pero an ambas tablas existe una columna (key) con el mismo nombre, para diferenciar la información entre la columna de una tabla y otra, pandas devulve el nombre de la columna con un guión bajo x (key_x) y otra con un guión bajo y (key_y)
# +
left2 = pd.DataFrame({'A': [1, 2], 'B': [2, 2]})
right2 = pd.DataFrame({'A': [4, 5, 6], 'B': [2, 2, 2]})
pd.merge(left2, right2, on='B', how='outer')
# -
# `Merge` también se puede usar como método, por lo que es posible concatener varias operaciones.
#
# Ejemplo: Retornar un dataframe con el nombre de todas las canciones, su álbum y artista, ordenados por nombre de artista, album y canción.
# +
tracks = chinook_query("select * from tracks")
albums = chinook_query("select * from albums")
artists = chinook_query("select * from artists")
tracks.head()
# -
artists.head()
(
tracks.rename(columns={"Name": "TrackName"})
.merge(
albums.rename(columns={"Title": "AlbumName"}),
how="left",
on="AlbumId"
)
.merge(
artists.rename(columns={"Name": "ArtistName"}),
how="left",
on="ArtistId"
)
.loc[:, ["TrackName", "AlbumName", "ArtistName"]]
.sort_values(["ArtistName", "AlbumName", "TrackName"])
)
|
m02_data_analysis/m02_c03_combining_data/m02_c03_combining_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Peer-graded Assignment Part B
# ## Instructions
#
# B. Normalize the data (5 marks)
#
# Repeat Part A but use a normalized version of the data. Recall that one way to normalize the data is by subtracting the mean from the individual predictors and dividing by the standard deviation.
#
# How does the mean of the mean squared errors compare to that from Step A?
# +
# import all required libraries
import numpy as np
import pandas as pd
from sklearn.model_selection import train_test_split
from keras.models import Sequential
from keras.layers import Dense
from sklearn.metrics import mean_squared_error
#import data
concrete_data = pd.read_csv('https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/DL0101EN/labs/data/concrete_data.csv')
#Normalize and split the data into training/test sets
y = concrete_data.Strength
X = concrete_data.drop('Strength', axis=1)
X_norm = (X - X.mean()) / X.std()
n_cols = X_norm.shape[1]
X_train, X_test, y_train, y_test = train_test_split(X_norm, y, test_size=0.3)
# +
# function to build our model, with 1 hidden layer that has 10 nodes, one output, using the adam optimizer and mean_squared_error for the loss function.
def regression_model():
model = Sequential()
model.add(Dense(10, activation='relu', input_shape=(n_cols,)))
model.add(Dense(1))
model.compile(optimizer='adam', loss='mean_squared_error')
return model
# function to build a trained model
def trained_model():
model = regression_model()
model.fit(X_train, y_train, validation_split=0.3, epochs=50, verbose=0)
return model
#Build a model and print the mse
model = trained_model()
y_predicted = model.predict(X_test)
mse = mean_squared_error(y_test, y_predicted)
print(mse)
# +
# function to train a model and return its MSE
def mse():
model = trained_model()
y_predicted = model.predict(X_test)
mse = mean_squared_error(y_test, y_predicted)
return mse
# build list of MSEs
# note: this takes a minute.
z = []
for i in range(50):
z.append(mse())
#print(z[i])
# mean of z
mn = np.mean(z)
print("Mean: ", mn)
# standard deviation of z
sd = np.std(z)
print("Standard Deviation: ", sd)
# -
# Results:
# Mean: 696.7189404593732
# Standard Deviation: 144.11727330401794
#
# From part A: Mean: 516.1412238566875
# Standard Deviation: 710.1154005722235
|
Course3/Capstone/PeerGradedPartB.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="xlWqj4sv8Kqm"
# #**Linear Regression**
# ### * Created by <NAME>
# ### * Created on Agosto 31, 2019
# + [markdown] colab_type="text" id="g__6St9K9eDM"
# ## For the Iris data set (https://archive.ics.uci.edu/ml/datasets/Iris)
# + [markdown] colab_type="text" id="n0U-OOOi-ILm"
# ### 1. Will the real iris data please stand up? Bezdek et. al. IEEE Transactions on Fuzzy Systems, Vol 7, Issue 3, pp. 368-369. June 1999.
# Many papers have been written that have used the iris dataset obtained from different sources, and some of these datasets contain errors in their data, which has caused a discrepancy between machine learning models created by different authors, in addition to some of them they do not provide the information from where they obtained the dataset, it is proposed to create a central repository for known datasets, although it has already been performed before, was not successfully, so far it is recommended to take the values directly from the Fisher’s paper.
# + [markdown] colab_type="text" id="M8ib_kkY_ntp"
# ### 2. In this exercise we will apply the Simple Linear Regression method in a simplified manner with the data. For this we will consider the total set of the three types of iris as a single class and we will use only two variables: the length of the petals as an independent (or input) variable and the width of the petal as the dependent (or output) variable. That is, the existence of a linear relationship between the two variables will be sought, regardless of the type of flower.
# + colab={} colab_type="code" id="AfE8Lt3f_yKT"
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import math
# + colab={"base_uri": "https://localhost:8080/", "height": 202} colab_type="code" id="PeBEb8nFFW0w" outputId="7b933b56-a12f-4f34-ee0d-b0778c9da4b3"
# The iris dataset was uploaded to the directory on my github URL
# After the reading we assign the columns to the panda dataframe
df1 = pd.read_csv('https://raw.githubusercontent.com/Wittline/Machine_Learning/master/Linear%20Regression/iris.data',
sep=',',
names=["sepallength", "sepalwidth", "petallength", "petalwidth", 'class']);
df1.head()
# + colab={} colab_type="code" id="DgXZ9TlYNp2z"
#Dropping the sepalwidth and sepallength columns
df1.drop(df1.columns[[0, 1]], axis = 1, inplace = True)
setosa= df1[df1['class']=='Iris-setosa']
versicolor =df1[df1['class']=='Iris-versicolor']
virginica =df1[df1['class']=='Iris-virginica']
# + colab={"base_uri": "https://localhost:8080/", "height": 202} colab_type="code" id="KdQ5jGouOZr0" outputId="15d9b04d-23b3-4b48-c0b9-3d2c8813d474"
df1.head()
# + [markdown] colab_type="text" id="e6mlXTMGBVQ0"
# ### i. Obtaining the summary of the 5 numbers of each of the two variables indicated: length and width of the petal. Also obtaining the variance and standard deviation of each one.
# + colab={"base_uri": "https://localhost:8080/", "height": 294} colab_type="code" id="Pdk_C4X8PhuK" outputId="ba2d3c8f-61ae-4aed-efc1-a9b015b399cf"
df1.describe()
# + [markdown] colab_type="text" id="toNtZATyB17x"
# ### ii. For each of the two variables obtain the histograms and the box diagrams.
# * Both variables are in the same units, correctly distributed.
# * The quartiles are correctly appreciated and there is no presence of outliers
#
# + colab={"base_uri": "https://localhost:8080/", "height": 449} colab_type="code" id="GtvpqKdHTkOd" outputId="8eb07ecd-711d-4a94-cfb7-66e8033b6da0"
x = df1.petallength
y = df1.petalwidth
plt.figure(figsize = (10, 7))
plt.hist(x, bins=20, alpha=0.6, label='Petal Length' )
plt.hist(y, bins=20, alpha=0.6, label='Petal Width')
plt.grid(True, alpha=0.6)
plt.ylabel("Frequency")
plt.title("Histogram", fontsize=20)
plt.legend(loc='upper right')
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 458} colab_type="code" id="AMAZQ9rUa2TS" outputId="c36d5192-04ed-4286-9078-96471a7907f7"
plt.figure(1, figsize=(10, 7))
bp = plt.boxplot([df1.petallength, df1.petalwidth], vert=True, patch_artist=True,
flierprops={'alpha':0.6, 'markersize': 6,
'markeredgecolor': '#555555','marker': 'd',
'markerfacecolor': "#555555"},
capprops={'color': '#555555', 'linewidth': 2},
boxprops={'color': '#555555', 'linewidth': 2},
whiskerprops={'color': '#555555', 'linewidth': 2},
medianprops={'color': '#555555', 'linewidth': 2},
meanprops={'color': '#555555', 'linewidth': 2})
plt.grid(True, alpha=0.6)
plt.title("Box Plot", fontsize=20)
plt.ylabel("Frequency", fontsize=20)
plt.xticks(ticks=[1, 2], labels=['petallength', 'petalwidth'], fontsize=20)
bp['boxes'][0].set(facecolor='blue', alpha= 0.6)
bp['boxes'][1].set(facecolor="orange",alpha= 0.6 )
plt.show()
# + [markdown] colab_type="text" id="Mf1lUQSdCwFy"
# ### iii. Obtaining the scatter plot of the variable length against the variable width.
# * For each kind of flower it is observed that while the length of the petal grows, the width grows in another proportion.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 472} colab_type="code" id="LzWF79c-pc2I" outputId="f35d3e00-5c72-438d-a5c5-ea2b9dad3b5e"
plt.figure(figsize=(10, 7))
plt.scatter(x = setosa.petallength, y = setosa.petalwidth, label="setosa", color='orange')
plt.scatter(x = versicolor.petallength, y = versicolor.petalwidth, label="versicolor", color='b')
plt.scatter(x = virginica.petallength, y = virginica.petalwidth, label="virginica", color='g')
plt.grid(True, alpha=0.6)
plt.title("Scatter", fontsize=20)
plt.xlabel("petallength", fontsize=20)
plt.ylabel("petalwidth", fontsize=20)
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="inWV0-NcDHmA"
# ### iv. Obtaining the linear regression model (equation) 𝑦 = 𝑎 + 𝑏𝑥 using the least squares method.
# 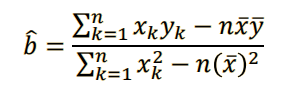
#
# 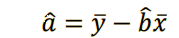
#
#
#
#
#
#
# # **y = -0.36651 + 0.4164*PetalLength**
# + colab={} colab_type="code" id="7L9dWICWANaW"
#Least square method, this will return the Variance and covariance of X and Y (X,Y)= (PetalLength, PetalWidth)
def Least_square(x,y):
mx = x.mean()
my = y.mean()
u=0
d=0
e=0
i=0;
while(i< len(x)):
u += ((x[i] - mx)*(y[i] - my))
d += ((x[i] - mx)**2)
e += ((y[i] - my)**2)
i +=1;
# 𝑦 = 𝑎 + 𝑏𝑥
#returning a and b
b = u/d
a = my - b*mx
return a, b, u, d, e
# + colab={"base_uri": "https://localhost:8080/", "height": 432} colab_type="code" id="HOKnIawb5RTr" outputId="3d95c338-cb76-4bbd-addc-c676479f5111"
x= df1.petallength
y= df1.petalwidth
a, b, covxy, vx, vy = Least_square(x, y)
yp= a + b*x
plt.figure(figsize=(10, 7))
plt.scatter(x,y,color="black", label= "(PetalLength, PetalWidth)")
plt.plot([min(x), max(x)], [min(yp), max(yp)], color="blue", label="y = -0.36651 + 0.4164x")
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="f_sSFkLIF6IJ"
# ### v. Find and interpret the p-value of the coefficients 𝑎 and 𝑏 found.
# 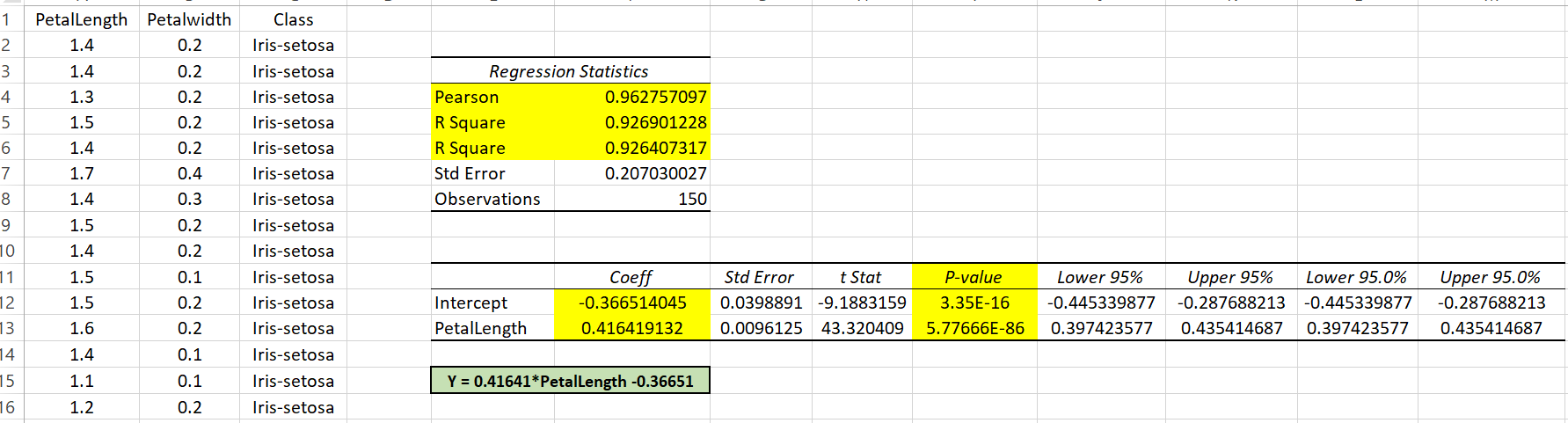
#
# + [markdown] colab_type="text" id="ljJUYcd0GBQ_"
# ### vi. Obtaining the Pearson correlation coefficient and interpret it.
# 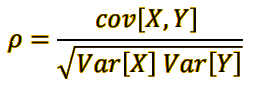
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="nvpg-7IL17BW" outputId="11ffacb1-e6c7-41b1-805a-843256c2fa3f"
pearson = (covxy/math.sqrt((vx*vy)))
pearson
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="teXv7UO1Gw5C" outputId="81a9bac6-9a9a-4ace-a3d0-0f10cc899d1d"
# Coefficient of determination
R2 = (pearson**2)
R2
# + [markdown] colab_type="text" id="yqaqIcXIHMGc"
# ### vii. Based on the results of items v and vi, what can you say about the model found? Do you consider that it is a good model and could be used to make predictions or inferences afterwards?
# * In the case of the p-values an excel hypothesis test was performed, both probabilities tell us how likely it is to reject each coefficient, it is the probability that it is equal to zero, to remove it from the equation, both probabilities are very small therefore both coefficients are significant and should not be removed from the linear equation. "The larger the number, the more likely it is to eliminate the coefficient"
# * If the variable pearson (r) ≈ +1, there is a direct linear relationship (with slope increasing) between petalLength and petalWidth. That is, if the petalLength values are increased, then the PetalWidth values are also increased proportionally.
#
#
#
|
Linear Regression/Code/Linear_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="uSUfDrbZywvc"
# # Offline Replayer Evaluation II - Recogym small
# > Running recogym for offline simulation and evaluation with small number of users and items
#
# - toc: true
# - badges: true
# - comments: true
# - categories: [bandit]
# - image:
# + [markdown] id="B7BGtRmWYACb"
# ## Environment setup
# + id="lE0TYT5zVVjy"
# !pip install -q recogym
# + id="cUH6Vc1OU89n"
import numpy as np
from numpy.random.mtrand import RandomState
from scipy.special import logsumexp
import scipy
import pandas as pd
from scipy.stats.distributions import beta
from copy import deepcopy
from scipy.sparse import csr_matrix
from scipy.sparse.linalg import svds
from itertools import chain
from sklearn.neighbors import NearestNeighbors
from IPython.display import display, HTML
from matplotlib.ticker import FormatStrFormatter
import gym, recogym
from recogym import env_1_args, Configuration
from recogym.agents import OrganicUserEventCounterAgent, organic_user_count_args
from recogym.agents.organic_count import OrganicCount, organic_count_args, to_categorical
from recogym import Configuration
from recogym.agents import Agent
from recogym.envs.observation import Observation
from recogym.agents import RandomAgent, random_args
from recogym import verify_agents, verify_agents_IPS
from recogym.evaluate_agent import plot_verify_agents, verify_agents_recall_at_k
from recogym.envs.session import OrganicSessions
from recogym.envs.context import DefaultContext
from recogym.envs.observation import Observation
import matplotlib.pyplot as plt
# %matplotlib inline
P = 50 # Number of Products
U = 50 # Number of Users
# + id="vk3eavl0VN_n"
# You can overwrite environment arguments here
env_1_args['random_seed'] = 42
env_1_args['num_products']= P
env_1_args['phi_var']=0.0
env_1_args['number_of_flips']=P//2
env_1_args['sigma_mu_organic'] = 0.1
env_1_args['sigma_omega']=0.05
# + id="8ShuzevnVI7b"
# Initialize the gym for the first time by calling .make() and .init_gym()
env = gym.make('reco-gym-v1')
env.init_gym(env_1_args)
# + id="p3ZfkuZcVMZT"
env.reset()
# + colab={"base_uri": "https://localhost:8080/", "height": 703} id="zdnvCn74VnMn" outputId="e6f5d948-bf69-4167-e04a-25bd575b0db6"
# Generate RecSys logs for U users
reco_log = env.generate_logs(U)
reco_log.head(20)
# + colab={"base_uri": "https://localhost:8080/"} id="xbB6tEBPVuDZ" outputId="1eed5eff-6834-45c0-ced3-d895be1f9aac"
n_events = reco_log.shape[0]
n_organic = reco_log.loc[reco_log['z'] == 'organic'].shape[0]
print('Training on {0} organic and {1} bandit events'.format(n_organic, n_events - n_organic))
# + [markdown] id="gX5Gve7eYG_r"
# ## Defining evaluation methods
# + [markdown] id="485rufJ-X8Ju"
# ### Traditional evaluation
# + id="44GbKe76WZAV"
def leave_one_out(reco_log, agent, last = False, N = 1, folds = 10):
# 1. Extract all organic events
reco_log = reco_log.loc[reco_log['z'] == 'organic']
# 2. For every user sequence - randomly sample out an item
hits = []
for _ in range(folds):
user_id = 0
history = []
session = OrganicSessions()
agent.reset()
for row in reco_log.itertuples():
# If we have a new user
if row.u != user_id:
if last:
# Sample out last item
index = len(history) - 1
else:
# Sample out a random item from the history
index = np.random.choice(len(history),
replace = False)
test = history[index]
train = history[:index] + history[index + 1:]
# 3. Recreate the user sequence without these items - Let the agent observe the incomplete sequence
for t, v in list(train):
session.next(DefaultContext(t, user_id), int(v))
# 4. Generate a top-N set of recommendations by letting the agent act
# TODO - For now only works for N = 1
try:
prob_a = agent.act(Observation(DefaultContext(t + 1, user_id), session), 0, False)['ps-a']
except:
prob_a = [1 / P] * P
# 5. Compute metrics checking whether the sampled test item is in the top-N
try:
hits.append(np.argmax(prob_a) == int(test[1]))
except:
hits.append(0)
# Reset variables
user_id = row.u
history = []
session = OrganicSessions()
agent.reset()
# Save the organic interaction to the running average for the session
history.append((row.t,row.v))
# Error analysis
mean_hits = np.mean(hits)
serr_hits = np.std(hits) / np.sqrt(len(hits))
low_bound = mean_hits - 1.96 * serr_hits
upp_bound = mean_hits + 1.96 * serr_hits
return mean_hits, low_bound, upp_bound
def verify_agents_traditional(reco_log, agents, last = False, N = 1, folds = 10):
# Placeholder DataFrame for result
stat = {
'Agent': [],
'0.025': [],
'0.500' : [],
'0.975': [],
}
# For every agent
for agent_id in agents:
# Compute HR@k
mean, low, upp = leave_one_out(reco_log, agents[agent_id], last = last, N = N, folds = folds)
stat['Agent'].append(agent_id)
stat['0.025'].append(low)
stat['0.500'].append(mean)
stat['0.975'].append(upp)
return pd.DataFrame().from_dict(stat)
# + [markdown] id="Ka70n5JcYPYE"
# ### Counterfactual evaluation
# + id="_HoADS4uX54y"
def compute_ips_weights(agent, reco_log):
# Placeholder for return values
rewards = [] # Labels for actions
t_props = [] # Treatment propensities
l_props = [] # Logging propensities
# For every logged interaction
user_id = 0
session = OrganicSessions()
agent.reset()
for row in reco_log.itertuples():
# If we have a new user
if row.u != user_id:
# Reset
session = OrganicSessions()
agent.reset()
user_id = row.u
# If we have an organic event
if row.z == 'organic':
session.next(DefaultContext(row.t, row.u), int(row.v))
else:
prob_a = agent.act(Observation(DefaultContext(row.t, row.u), session), 0, False)['ps-a']
rewards.append(row.c)
try:
t_props.append(prob_a[int(row.a)])
except:
t_props.append(0)
l_props.append(row.ps)
session = OrganicSessions()
return np.asarray(rewards), np.asarray(t_props), np.asarray(l_props)
def verify_agents_counterfactual(reco_log, agents, cap = 3):
# Placeholder DataFrame for results
IPS_stat = {
'Agent': [],
'0.025': [],
'0.500' : [],
'0.975': [],
}
CIPS_stat = {
'Agent': [],
'0.025': [],
'0.500' : [],
'0.975': [],
}
SNIPS_stat = {
'Agent': [],
'0.025': [],
'0.500' : [],
'0.975': [],
}
# For every agent
for agent_id in agents:
# Get the rewards and propensities
rewards, t_props, l_props = compute_ips_weights(agents[agent_id], reco_log)
# Compute the sample weights - propensity ratios
p_ratio = t_props / l_props
# Effective sample size for E_t estimate (from <NAME>)
n_e = len(rewards) * (np.mean(p_ratio) ** 2) / (p_ratio ** 2).mean()
n_e = 0 if np.isnan(n_e) else n_e
print("Effective sample size for agent {} is {}".format(str(agent_id), n_e))
# Critical value from t-distribution as we have unknown variance
alpha = .00125
cv = scipy.stats.t.ppf(1 - alpha, df = int(n_e) - 1)
###############
# VANILLA IPS #
###############
# Expected reward for pi_t
E_t = np.mean(rewards * p_ratio)
# Variance of the estimate
var = ((rewards * p_ratio - E_t) ** 2).mean()
stddev = np.sqrt(var)
# C.I. assuming unknown variance - use t-distribution and effective sample size
min_bound = E_t - cv * stddev / np.sqrt(int(n_e))
max_bound = E_t + cv * stddev / np.sqrt(int(n_e))
# Store result
IPS_stat['Agent'].append(agent_id)
IPS_stat['0.025'].append(min_bound)
IPS_stat['0.500'].append(E_t)
IPS_stat['0.975'].append(max_bound)
##############
# CAPPED IPS #
##############
# Cap ratios
p_ratio_capped = np.clip(p_ratio, a_min = None, a_max = cap)
# Expected reward for pi_t
E_t_capped = np.mean(rewards * p_ratio_capped)
# Variance of the estimate
var_capped = ((rewards * p_ratio_capped - E_t_capped) ** 2).mean()
stddev_capped = np.sqrt(var_capped)
# C.I. assuming unknown variance - use t-distribution and effective sample size
min_bound_capped = E_t_capped - cv * stddev_capped / np.sqrt(int(n_e))
max_bound_capped = E_t_capped + cv * stddev_capped / np.sqrt(int(n_e))
# Store result
CIPS_stat['Agent'].append(agent_id)
CIPS_stat['0.025'].append(min_bound_capped)
CIPS_stat['0.500'].append(E_t_capped)
CIPS_stat['0.975'].append(max_bound_capped)
##############
# NORMED IPS #
##############
# Expected reward for pi_t
E_t_normed = np.sum(rewards * p_ratio) / np.sum(p_ratio)
# Variance of the estimate
var_normed = np.sum(((rewards - E_t_normed) ** 2) * (p_ratio ** 2)) / (p_ratio.sum() ** 2)
stddev_normed = np.sqrt(var_normed)
# C.I. assuming unknown variance - use t-distribution and effective sample size
min_bound_normed = E_t_normed - cv * stddev_normed / np.sqrt(int(n_e))
max_bound_normed = E_t_normed + cv * stddev_normed / np.sqrt(int(n_e))
# Store result
SNIPS_stat['Agent'].append(agent_id)
SNIPS_stat['0.025'].append(min_bound_normed)
SNIPS_stat['0.500'].append(E_t_normed)
SNIPS_stat['0.975'].append(max_bound_normed)
return pd.DataFrame().from_dict(IPS_stat), pd.DataFrame().from_dict(CIPS_stat), pd.DataFrame().from_dict(SNIPS_stat)
# + [markdown] id="bvr-NmAdYUSg"
# ## Creating agents
# + [markdown] id="MTAIbCveYWTy"
# ### SVD agent
# + id="6xcs7TqWYSbI"
class SVDAgent(Agent):
def __init__(self, config, U = U, P = P, K = 5):
super(SVDAgent, self).__init__(config)
self.rng = RandomState(self.config.random_seed)
assert(P >= K)
self.K = K
self.R = csr_matrix((U,P))
self.V = np.zeros((P,K))
self.user_history = np.zeros(P)
def train(self, reco_log, U = U, P = P):
# Extract all organic user logs
reco_log = reco_log.loc[reco_log['z'] == 'organic']
# Generate ratings matrix for training, row-based for efficient row (user) retrieval
self.R = csr_matrix((np.ones(len(reco_log)),
(reco_log['u'],reco_log['v'])),
(U,P))
# Singular Value Decomposition
_, _, self.V = svds(self.R, k = self.K)
def observe(self, observation):
for session in observation.sessions():
self.user_history[session['v']] += 1
def act(self, observation, reward, done):
"""Act method returns an Action based on current observation and past history"""
self.observe(observation)
scores = self.user_history.dot(self.V.T).dot(self.V)
action = np.argmax(scores)
prob = np.zeros_like(scores)
prob[action] = 1.0
return {
**super().act(observation, reward, done),
**{
'a': action,
'ps': prob[action],
'ps-a': prob,
},
}
def reset(self):
self.user_history = np.zeros(P)
# + [markdown] id="CwOGMwwyYjTQ"
# ### Item-KNN agent
# + id="YHIXb-KHYejQ"
class itemkNNAgent(Agent):
def __init__(self, config, U = U, P = P, k = 5, greedy = False, alpha = 1):
super(itemkNNAgent, self).__init__(config)
self.rng = RandomState(self.config.random_seed)
self.k = min(P,k)
self.greedy = greedy
self.alpha = alpha
self.Rt = csr_matrix((P,U))
self.user_history = np.zeros(P)
self.S = np.eye(P)
def train(self, reco_log, U = U, P = P):
# Extract all organic user logs
reco_log = reco_log.loc[reco_log['z'] == 'organic']
# Generate ratings matrix for training, row-based for efficient row (user) retrieval
self.R_t = csr_matrix((np.ones(len(reco_log)),
(reco_log['v'],reco_log['u'])),
(P,U))
# Set up nearest neighbours module
nn = NearestNeighbors(n_neighbors = self.k,
metric = 'cosine')
# Initialise placeholder for distances and indices
distances = []
indices = []
# Dirty fix for multiprocessing backend being unable to pickle large objects
nn.fit(self.R_t)
distances, indices = nn.kneighbors(self.R_t, return_distance = True)
# Precompute similarity matrix S
data = list(chain.from_iterable(1.0 - distances))
rows = list(chain.from_iterable([i] * self.k for i in range(P)))
cols = list(chain.from_iterable(indices))
# (P,P)-matrix with cosine similarities between items
self.S = csr_matrix((data,(rows, cols)), (P,P)).todense()
def observe(self, observation):
for session in observation.sessions():
self.user_history[session['v']] += 1
def act(self, observation, reward, done):
"""Act method returns an Action based on current observation and past history"""
self.observe(observation)
scores = self.user_history.dot(self.S).A1
if self.greedy:
action = np.argmax(scores)
prob = np.zeros_like(scores)
prob[action] = 1.0
else:
scores **= self.alpha
prob = scores / np.sum(scores)
action = self.rng.choice(self.S.shape[0], p = prob)
return {
**super().act(observation, reward, done),
**{
'a': action,
'ps': prob[action],
'ps-a': prob,
},
}
def reset(self):
self.user_history = np.zeros(P)
# + [markdown] id="qK82zI6qYspo"
# ### User-KNN agent
# + id="uoRDZ8mNYpLo"
class userkNNAgent(Agent):
def __init__(self, config, U = U, P = P, k = 5, greedy = False, alpha = 1):
super(userkNNAgent, self).__init__(config)
self.rng = RandomState(self.config.random_seed)
self.k = min(P,k)
self.greedy = greedy
self.alpha = alpha
self.U = U
self.P = P
self.R = csr_matrix((U,P))
self.user_history = np.zeros(P)
self.nn = NearestNeighbors(n_neighbors = self.k, metric = 'cosine')
def train(self, reco_log, U = U, P = P):
# Extract all organic user logs
reco_log = reco_log.loc[reco_log['z'] == 'organic']
# Generate ratings matrix for training, row-based for efficient row (user) retrieval
self.R = csr_matrix((np.ones(len(reco_log)),
(reco_log['u'],reco_log['v'])),
(U,P))
# Fit nearest neighbours
self.nn.fit(self.R)
def observe(self, observation):
for session in observation.sessions():
self.user_history[session['v']] += 1
def act(self, observation, reward, done):
"""Act method returns an Action based on current observation and past history"""
self.observe(observation)
# Get neighbouring users based on user history
distances, indices = self.nn.kneighbors(self.user_history.reshape(1,-1))
scores = np.add.reduce([dist * self.R[idx,:] for dist, idx in zip(distances,indices)])
if self.greedy:
action = np.argmax(scores)
prob = np.zeros_like(scores)
prob[action] = 1.0
else:
scores **= self.alpha
prob = scores / np.sum(scores)
action = self.rng.choice(self.P, p = prob)
return {
**super().act(observation, reward, done),
**{
'a': action,
'ps': prob[action],
'ps-a': prob,
},
}
def reset(self):
self.user_history = np.zeros(P)
# + [markdown] id="CcVL6ih6Y0p8"
# ### Agent initializations
# + id="qewN2myJ0Qvu"
# SVD Agent
SVD_agent = SVDAgent(Configuration(env_1_args), U, P, 30)
# item-kNN Agent
itemkNN_agent = itemkNNAgent(Configuration(env_1_args), U, P, 500, greedy = True)
# user-kNN Agent
userkNN_agent = userkNNAgent(Configuration(env_1_args), U, P, 20, greedy = True)
# Generalised Popularity agent
GPOP_agent = OrganicCount(Configuration({
**env_1_args,
'select_randomly': True,
}))
# Generalised Popularity agent
GPOP_agent_greedy = OrganicCount(Configuration({
**env_1_args,
'select_randomly': False,
}))
# Peronalised Popularity agent
PPOP_agent = OrganicUserEventCounterAgent(Configuration({
**organic_user_count_args,
**env_1_args,
'select_randomly': True,
}))
# Peronalised Popularity agent
PPOP_agent_greedy = OrganicUserEventCounterAgent(Configuration({
**organic_user_count_args,
**env_1_args,
'select_randomly': False,
}))
# Random Agent
random_args['num_products'] = P
RAND_agent = RandomAgent(Configuration({**env_1_args, **random_args,}))
# + id="wYX3_5fYYumd"
SVD_agent.train(reco_log)
itemkNN_agent.train(reco_log)
userkNN_agent.train(reco_log)
# + [markdown] id="qRqbHqMJY9vL"
# ## Offline evaluation
# + [markdown] id="E9r-zhlAZZ9M"
# ### Generating test logs
# + colab={"base_uri": "https://localhost:8080/"} id="pnzTANe1Y3lW" outputId="ac6746cc-cac4-4ffd-d9bf-185be390bfca"
# %%time
# Placeholder for agents
agents = {
' Random': RAND_agent,
' Popular': GPOP_agent_greedy,
' User-pop': PPOP_agent,
' SVD': SVD_agent,
' User-kNN': userkNN_agent,
'Item-kNN': itemkNN_agent,
}
agent_ids = sorted(list(agents.keys()))#['SVD','GPOP','PPOP','RAND']
# Generate new logs, to be used for offline testing
n_test_users = 50 # U
test_log = env.generate_logs(n_test_users)
n_events = test_log.shape[0]
n_organic = test_log.loc[test_log['z'] == 'organic'].shape[0]
print('Testing on {0} organic and {1} bandit events'.format(n_organic, n_events - n_organic))
# + [markdown] id="igubir8vaB0m"
# ### (Util) helper function to plot barchart
# + id="zCpbb9K-ZB-b"
def plot_barchart(result, title, xlabel, col = 'tab:red', figname = 'fig.eps', size = (6,2), fontsize = 12):
fig, axes = plt.subplots(figsize = size)
plt.title(title, size = fontsize)
n_agents = len(result)
yticks = np.arange(n_agents)
mean = result['0.500']
lower = result['0.500'] - result['0.025']
upper = result['0.975'] - result['0.500']
plt.barh(yticks,
mean,
height = .25,
xerr = (lower, upper),
align = 'center',
color = col,)
plt.yticks(yticks, result['Agent'], size = fontsize)
plt.xticks(size = fontsize)
plt.xlabel(xlabel, size = fontsize)
plt.xlim(.0,None)
plt.gca().xaxis.set_major_formatter(FormatStrFormatter('%.2f'))
plt.savefig(figname, bbox_inches = 'tight')
plt.show()
# + [markdown] id="w9r6FSeDZdyM"
# ### Leave-one-out evaluation
# + colab={"base_uri": "https://localhost:8080/", "height": 479} id="zJSfQ5RDZpab" outputId="cc6ff56c-29d7-4dbb-8c0f-8f2cc97eca86"
# %%time
result_LOO = verify_agents_traditional(test_log, deepcopy(agents))
display(result_LOO)
plot_barchart(result_LOO, 'Evaluate on Organic Feedback', 'HR@1', 'tab:red', 'traditional_eval.eps')
# + [markdown] id="KMEG7NLGZ5Kp"
# ### IPS Estimators
# + colab={"base_uri": "https://localhost:8080/"} id="RuQ5m3goeHnR" outputId="2112c1ff-c0d3-4ae9-e6d1-a97c054756a7"
# Generate new logs, to be used for offline testing
test_log_ppop = env.generate_logs(n_test_users, agent = deepcopy(PPOP_agent))
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="Gjj31s8ge6qZ" outputId="c5372867-642f-41df-94b7-ff3a5a02068f"
test_log_ppop.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 975} id="SdKK38bSZ6Bl" outputId="d2356d2e-a0d9-46b6-ee08-6d9dea3d31ef"
# %%time
cap = 15
result_IPS, result_CIPS, result_SNIPS = verify_agents_counterfactual(test_log_ppop, deepcopy(agents), cap = cap)
display(result_IPS)
plot_barchart(result_IPS, 'IPS', 'CTR', 'tab:blue', 'bandit_eval_noclip.eps')
display(result_CIPS)
plot_barchart(result_CIPS, 'Clipped IPS', 'CTR', 'tab:blue', 'bandit_eval_clip{0}.eps'.format(cap))
# + [markdown] id="NqDH9cq_b4vh"
# ### A/B tests
# + id="Pvc_5gPg0HA3"
n_test_users = 50 # U
agents = {
' Random': RAND_agent,
' Popular': GPOP_agent_greedy,
' User-pop': PPOP_agent,
' SVD': SVD_agent,
' User-kNN': userkNN_agent,
'Item-kNN': itemkNN_agent,
}
# + colab={"base_uri": "https://localhost:8080/", "height": 649} id="DsK98lKNb6RB" outputId="52732828-5c5e-4c9b-b64e-796b6449b611"
# %%time
result_AB = verify_agents(env, n_test_users, deepcopy(agents))
display(result_AB)
plot_barchart(result_AB, 'A/B-test', 'CTR', 'tab:green', 'ABtest_eval.eps')
# + colab={"base_uri": "https://localhost:8080/", "height": 553} id="ieFcpaa-b8Tt" outputId="73f390e5-27d0-494b-9d78-7bf439e68bd0"
def combine_barchart(resultAB, resultCIPS, title, xlabel, figname = 'fig.eps', size = (10,8), fontsize = 12):
fig, axes = plt.subplots(figsize = size)
plt.title(title, size = fontsize)
n_agents = len(resultAB)
for i, (name, colour, result) in enumerate([('A/B-test', 'tab:green', result_AB),('CIPS', 'tab:blue', result_CIPS)]):
mean = result['0.500']
lower = result['0.500'] - result['0.025']
upper = result['0.975'] - result['0.500']
height = .25
yticks = [a + i * height for a in range(n_agents)]
plt.barh(yticks,
mean,
height = height,
xerr = (lower, upper),
align = 'edge',
label = name,
color = colour)
plt.yticks(yticks, result['Agent'], size = fontsize)
plt.xticks(size = fontsize)
plt.xlabel(xlabel, size = fontsize)
plt.legend(loc = 'lower right')
plt.xlim(.0,None)
plt.gca().xaxis.set_major_formatter(FormatStrFormatter('%.3f'))
plt.savefig(figname, bbox_inches = 'tight')
plt.show()
combine_barchart(result_AB, result_CIPS, 'Evaluate on Bandit Feedback', 'CTR', 'ABtest_CIPS.eps')
# + colab={"base_uri": "https://localhost:8080/", "height": 193} id="Q4NB72lgb-Ek" outputId="e253dce2-29ab-483c-c461-6a3a2157cdf6"
plot_barchart(result_LOO, 'Evaluate on Organic Feedback', 'HR@1', 'tab:red', 'traditional_eval.eps')
|
_notebooks/2021-06-17-recostep-tutorial-offline-replayer-eval-recogym-small.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
# ---
# # Подготовка предложений
# > Для английского языка до разрыва строки можно размещать нескольких предложений (абзацы), поскольку в скрипте реализовано разбиение английского текста на предложения и модель en-ru может выполнять пакетную обработку предложений.
#
# > Для арабского языка до разрыва строки можно размещать только одно предложение (без точки ".", т.к. знаки препинания в арабском имеют иное начертание, например: نُقْطَةٌ - точка, عَلامةُ استِفْهامٍ - вопросительный знак, عَلامَةُ تَعَجُّبٍ - восклицательный знак)!
# ## Арабский текст (input.ar)
#
# запись
file = pd.ExcelFile('../../test_dataset/test_dataset.xlsx')
df = file.parse()
text = []
s = 0
for sent in df.iloc[:]['Арабский']:
tmp = sent.replace('\n', ' ').strip()
text.extend(tmp.split('.')) # убираем точку
text = [t.strip() for t in text if t]
with open("input.ar", 'w', encoding="utf-8") as f:
f.write('\n'.join(text) + '\n')
# чтение первых 3 предложений
with open("input.ar", encoding='utf-8') as f:
file = f.readlines()
print(*file[:3])
# ## Английский текст (input.en)
# запись
file = pd.ExcelFile('../../test_dataset/test_dataset.xlsx')
df = file.parse()
text = []
for sent in df.iloc[:]['Английский ']:
text.append(sent.strip())
with open("input.en", 'w', encoding="utf-8") as f:
f.write('\n'.join(text) + '\n')
# чтение первых 3 абзацев
with open("input.en", encoding='utf-8', newline='') as f:
file = f.readlines()
print(*file[:3])
# ## Русский текст (input.ru)
# запись
file = pd.ExcelFile('../../test_dataset/test_dataset.xlsx')
df = file.parse()
text = []
for sent in df.iloc[:]['Русский']:
text.append(sent.replace('\n', ' ').strip())
with open("input.ru", 'w', encoding="utf-8") as f:
f.write('\n'.join(text) + '\n')
# чтение первых 3 абзацев
with open("input.ru", encoding='utf-8', newline='') as f:
file = f.readlines()
print(*file[:3])
# ---
# # Предобработка текста, поступающая в предобученные модели
# ## Арабский
# ## Английский
t = '▁This ▁case ▁was ▁resolved ▁with ▁the ▁same ▁evidence ▁that ▁they ▁had ▁at ▁least ▁10 ▁years ▁before . ▁If ▁justice ▁is ▁a ▁form ▁of ▁dis s ua sion ▁to ▁prevent ▁people ▁from ▁committing ▁crimes , ▁impunity ▁in ▁Colombia ▁is ▁almost ▁an ▁incentive , ▁because ▁it ▁is ▁really ▁difficult ▁that ▁someone ▁is ▁convicted ▁even ▁though ▁they ▁are ▁guilty .'
ord(t[0])
ord(t[5])
print(ord(' '))
# ## Русский
# ---
# # Постобработка текста, поступающая из модели
# ## Арабский
# ## Английский
# ## Русский
# ---
# # Результаты переводов
# ## Англо-русский
# | Английский | Модель en-ru | Русский |
# | --- | --- | --- |
# | This case was resolved with the same evidence that they had at least 10 years before. If justice is a form of dissuasion to prevent people from committing crimes, impunity in Colombia is almost an incentive, because it is really difficult that someone is convicted even though they are guilty. | Это дело было урегулировано с использованием тех же доказательств, что и по меньшей мере 10 лет назад. Если правосудие является одной из форм устрашения, чтобы не допустить совершения людьми преступлений, то безнаказанность в Колумбии является практически стимулом, поскольку действительно трудно, чтобы кто-то был осужден, даже несмотря на то, что он виновен. | Это дело закрыли с теми же доказательствами, что у них имелись по крайней мере 10 лет назад. Если правосудие — это форма сдерживания людей от совершения преступлений, то безнаказанность в Колумбии — это почти стимул для злоумышленников, потому что крайне трудно осудить человека, даже если известно, что он виновен. |
# ## Арабско-русский
|
reports/pretrained_model_test/preparation_and_evaluation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from sklearn import preprocessing
from keras.models import Sequential
from keras.layers import Dense
from keras.utils import to_categorical
from keras.callbacks import EarlyStopping
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.decomposition import PCA
dfg = pd.read_csv('wat-r2-good.csv')
dfm = pd.read_csv('wat-r2-mals.csv')
dfg.head()
dfm.head()
df = dfg.append(dfm, ignore_index=True)
df.head()
df = df.sort_values('time')
df.head(50)
df.to_csv('wat-r2-pkt.csv',index=False)
df = pd.read_csv("wat-r2-pkt.csv")
df
df.corr()
train_X = df.drop(columns=['packet_address','time','target','pkt_count_all','pkt_count_all','pkt_max_all','src_router','src_ni'])
train_X
#normalization
x = train_X.values
min_max_scaler = preprocessing.MinMaxScaler()
x_scaled = min_max_scaler.fit_transform(x)
train_X = pd.DataFrame(x_scaled)
train_X
corr_df = pd.concat([train_X, df[['target']]], axis = 1)
corr_df.corr()
train_Y = df['target']
train_Y
pca = PCA(0.95)
pca.fit(train_X)
principal_components = pca.transform(train_X)
principal_components
pca.explained_variance_ratio_
features = range(pca.n_components_)
plt.bar(features, pca.explained_variance_)
plt.xticks(features)
plt.xlabel("PCs")
plt.ylabel("Variance")
principal_df = pd.DataFrame(data = principal_components)
principal_df
final_df = pd.concat([principal_df, df[['target']]], axis = 1)
final_df
final_df.corr()
fig = plt.figure(figsize = (8,8))
ax = fig.add_subplot(1,1,1)
ax.set_xlabel('Principal Component 1', fontsize = 15)
ax.set_ylabel('Principal Component 2', fontsize = 15)
ax.set_title('2D PCA', fontsize = 20)
targets = [1, 0]
colors = ['r', 'g']
for target, color in zip(targets,colors):
indicesToKeep = final_df['target'] == target
ax.scatter(final_df.loc[indicesToKeep, 0]
, final_df.loc[indicesToKeep, 1]
, c = color
, s = 50)
ax.legend(targets)
ax.grid()
dup_df = df.drop(columns=['packet_address','time'])
dup_df
dup_df[dup_df.duplicated()].shape
dup_df[dup_df.duplicated()].count()
# + jupyter={"source_hidden": true}
seed = 7
np.random.seed(seed)
# -
x_train, x_test, y_train, y_test = train_test_split(train_X, train_Y, test_size=0.32, random_state=seed, shuffle=True)
model = Sequential()
n_cols = train_X.shape[1]
n_cols
model.add(Dense(32, activation='relu', input_shape=(n_cols,)))
model.add(Dense(16, activation='relu'))
model.add(Dense(8, activation='relu'))
model.add(Dense(4, activation='relu'))
model.add(Dense(1, activation='sigmoid'))
model.compile(optimizer='sgd', loss='mean_squared_error', metrics=['accuracy'])
early_stopping_monitor = EarlyStopping(patience=20)
model.fit(x_train, y_train, epochs=50, validation_split=0.4)
scores = model.evaluate(x_test, y_test, verbose=0)
print("%s: %.2f%%" % (model.metrics_names[1], scores[1]*100))
dff = pd.read_csv('wat-r2-pkt.csv',nrows=500)
sns.distplot(df['src_router'], kde = False, bins=30, color='blue')
sns.distplot(df['dst_router'], kde = False, bins=30, color='red')
sns.distplot(df['inport'], kde = False, bins=30, color='green')
sns.distplot(df['outport'], kde = False, bins=30, color='green')
sns.distplot(df['packet_type'], kde = False, bins=30, color='red')
|
[02 - Modeling]/dos ver 5.1/wat-r2-model-v1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# This notebook was prepared by [<NAME>](https://github.com/donnemartin). Source and license info is on [GitHub](https://github.com/donnemartin/interactive-coding-challenges).
# # Challenge Notebook
# ## Problem: Determine the number of bits to flip to convert int a to int b'.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
#
# * Can we assume A and B are always ints?
# * Yes
# * Is the output an int?
# * Yes
# * Can we assume A and B are always the same number of bits?
# * Yes
# * Can we assume the inputs are valid (not None)?
# * No
# * Can we assume this fits memory?
# * Yes
# ## Test Cases
#
# * A or B is None -> Exception
# * General case
# <pre>
# A = 11101
# B = 01111
# Result: 2
# <pre>
# ## Algorithm
#
# Refer to the [Solution Notebook](). If you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
a = int('11101')
b = int('01101')
a ^ b
class Bits(object):
def bits_to_flip(self, a, b):
if a is None or b is None:
raise TypeError('a or b cannot be None')
# TODO: Implement me
counter = 0
pass
# ## Unit Test
# **The following unit test is expected to fail until you solve the challenge.**
# +
# # %load test_bits_to_flip.py
import unittest
class TestBits(unittest.TestCase):
def test_bits_to_flip(self):
bits = Bits()
a = int('11101', base=2)
b = int('01111', base=2)
expected = 2
self.assertEqual(bits.bits_to_flip(a, b), expected)
print('Success: test_bits_to_flip')
def main():
test = TestBits()
test.test_bits_to_flip()
if __name__ == '__main__':
main()
# -
# ## Solution Notebook
#
# Review the [Solution Notebook]() for a discussion on algorithms and code solutions.
|
bit_manipulation/bits_to_flip/bits_to_flip_challenge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Problem Set 3
# ## Running and Interpreting Regressions
# ## Due Date: Friday 3/8, 8:00 AM
# Put picture / instructions / grading info here.
# +
import random
import pandas as pd
import numpy as np
from scipy import stats
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
sns.set()
# -
# ## Problem 1: Recognizing Patterns in the Data
# The two graphs below shows the empirical distribution for daily earnings for 10,000 individuals (these are simulated rather than real data points). The first graph shows the distribution for all workers pooled. The second graph shows three separate distributions where workers are stratified into three separate skill groups: low skilled, medium skilled, and high skilled.
# +
## Simulating random data
daily_earnings = [random.gauss(300, 100) for x in range(10000)]
## Setting up plot
sns.distplot(daily_earnings);
# +
skill_low = [random.gauss(200, 40) for x in range(3334)]
skill_med = [random.gauss(300, 40) for x in range(3333)]
skill_high = [random.gauss(400, 40) for x in range(3333)]
sns.distplot(skill_low)
sns.distplot(skill_med)
sns.distplot(skill_high);
# -
# **Part 1**: Comparing these two graphs, does this stratification seem meaningful? That is to say, does stratifying the sample by skill group explain variance in daily earnings? Why or why not?
# *Answer here*
# **Part 2**: Suppose you wished to formally test the hypothesis that mean earning differ across these three groups. How would you do so? Be explicit in terms of formally stating the hypothesis, how you would construct the test statistic and which distribution you would use. Suppose you have 3,334 observations for low skilled workers, and 3,333 for medium and high skilled workers.
# *Answer here*
# **Part 3**: In our discussion of the bivariate model, we laid out a series of assumptions regarding the data generating process that we are assuming to be true when we fit a bivariate regression model. These assumptions include that the population regression function is linear in the x variable and a host of assumptions regarding the error term of the equation that generates the data. The following two graphs (again using data I simulated) display scatter plots and fitted bivariate regression lines for two separate relationships. In each, one of the assumptions that we commonly make is violated. **Identify and describe the violation for each graph**.
# +
# Create a heteroscedastic graph here
# -
# *Answer here*
# +
# Create a graph with uneven residuals here
# -
# *Answer here*
# The graph below presents a scatter plot of the number of recorded earthquakes that are magnitude 8.0 or higher for each year between 1916 and 2015. The data also shows a fitted regression line displaying the linear trend in large earthquakes (note, this is actual and not simulated data). I have connected consecutive data points by a line to visibly connect consecutive annual counts of large earthquakes. Again, one of the assumptions that we commonly make regarding the data generating process is violated in this graph. **Which one? What does this violation mean in terms of predicting patterns in seismic activity?**
# +
# Reading in data
earthquakes = pd.read_csv('earthquake.csv')
# Converting time column to datetime object
earthquakes['time'] = pd.to_datetime(earthquakes['time'])
# Filtering the dataset to only earthquakes larger than 8.0
earthquakes_eight = earthquakes[earthquakes['mag'] >= 8.0]
# Setting up figure
plt.figure(figsize=(12,6))
# Plotting figure
plt.plot(earthquakes_eight['time'], earthquakes_eight['mag'], marker = 'o')
# Adjusting figure presentation
plt.title('Earthquakes Greater than 8.0 Magnitude, 1916 - 2015', fontsize = 20)
plt.xlabel('Year', fontsize = 14)
plt.ylabel('Magnitude', fontsize = 14);
# Would like to agg a line of best fit
# Would like to get space between title and plot
# -
# *Answer here*
# ## Problem 2: Testing for a Difference in Means
# Using the earthquake data, I created a dummy variable equal to one for all years after 1965. This splits the sample into 50 early years and 50 later years. The table below presents the results of running a t-test on that dummy variable:
# +
# Creating dummy variable where 1966 and later is 1, 1965 and earlier is 0
earthquakes['years66plus'] = np.where(earthquakes['time'] >= pd.to_datetime('1966-01-01 00:00:00'), 1, 0)
# Separating data based on dummy
earthquakes_pre = earthquakes[earthquakes['years66plus'] == 0]
earthquakes_post = earthquakes[earthquakes['years66plus'] == 1]
# Running basic t-test
stats.ttest_ind(earthquakes_pre['mag'], earthquakes_post['mag'], equal_var = True)
# Need to adjust this so it puts out more information for 3/4
# or just make clear that that they need to grab that information on their own
# -
# **Part 1**: What hypothesis am I testing exactly by running this command? (State it in terms of a null and alternative.)
# *Answer here*
# **Part 2**: Using a two-tailed test, what is the likelihood of observing the differences in average annual earthquakes under the null hypothesis of no difference?
# *Answer here*
# **Part 3**: Compare the confidence intervals for the average number of quakes in the early years and the later years. What do you notice about these two intervals that strongly suggests that there is a statistically significant difference in average earthquakes across these two time periods?
# *Answer here*
# **Part 4**: Using the standard deviation estimates and the sample size estimates, calculate the standard error for the average for each year grouping (*Note: The answer is in the output, so you should be able to directly verify that have calculated the correct answer. You should use the formulas presented in lecture 2*).
# *Answer here*
# ## Problem 3: Comparing Bivariate and Multivariate Regression Results
# The figure below shows three scatter plots (these are simulated rather than actual data). Suppose the data pertains to full time workers in the U.S. between the ages of 30 and 50.
#
# 1. The first scatter plot shows annual earnings against the percent of people who are poor in one’s zip code of residence at birth.
# 2. The second scatter plot shows the relationship between one’s annual earnings as an adult and the educational attainment of each person’s mother as recorded on their birth certificate.
# 3. The final scatter plot shows the relationship between percent poor in birth zip code and mother’s education.
#
# From these three figures we see that...
# 1. People born into poor zip codes tend to have lower earnings as an adult;
# 2. People with more educated mothers make more money as adults; and
# 3. People with more educated mothers tend to be raised in wealthier zip codes.
# +
# Scatter plots here
# -
# **Part 1**: Suppose we first run a regression of annual earnings on the poverty rate in one’s birth zip code. The output from this regression follows:
# +
# Regression here
# -
# We then run a second regression where the dependent variable is again annual earnings but our control variables include both poverty in birth zip code as well as maternal educational attainment. Using the omitted variables formula, what do you predict will be the effect of adding a control for maternal education on our estimate of the marginal effect of higher poverty in one’s birth zip on adult earnings? Alternatively stated, how will the coefficient on zip poverty rates differ in these two regressions?
# *Answer here*
# **Part 2**: The output below presents the regression output from a model where the dependent variable is the annual earnings and the two explanatory variables are the percent poor in one’s birth zip code and maternal educational attainment. Using this output, how many additional years of maternal education would it take to offset the disadvantage of growing up in a zip code that is 100 percent poor relative to a zip code is 0 percent poor? **(Remembers, this regression is estimated using a simulated (i.e., made up) data set. So please do not read anything into the actual numbers)**.
# *Answer here*
# **Part 3**: Compare the output from these two regressions:
#
# - Why is the root mean square error smaller in the multivariate regression relative to the bivariate regression?
# - Why is the total sum of squares the same in both regressions?
# - The standard error for the coefficient on zip_poverty is larger in the multivariate regression than in the bivariate regression. However, the root mean square error (the numerator in the equation for the standard error for this coefficient) is smaller in the multivariate equation? How can this be?
# - Describe the simple hypothesis test that would basically test the first model against the second (note the first model is nested within the second in that you can impose a restriction on the parameters of the second model that would leave us with the first). Can you reject the model of the simple bivariate model in favor of the multivariate model using the output above?
# *Answer here*
# ## Problem 4: Analyzing Gender Differentials in Weekly Earnings
# **Note: This section is more barebones than in the problem set, where Steve dedicates a little bit of time to explaining how to upload the data and then how to run regressions. Could create interactive example to replace that.**
# The data set from this question is an extract from the 2000 March Current Population Survey and contains data on weekly earnings, gender, age, age squared, and a set of dummy variables indicating level of educational attainment for approximately 13,000 randomly selected working adults in the United States.
# +
cps_march = pd.read_csv('ps3data.csv')
cps_march.shape
# -
cps_march.head()
# Using this dataset, do the following:
# **Part 1**: Calculate the sample averages of all of the variables in the data set. For the dummy variables in the data set, what are these averages telling us?
# *Answer here*
# **Part 2**: Estimate a regression where the dependent variable is weekly earnings and the only independent variable is a female dummy variable. What is the mean difference in earnings between men and women?
# *Answer here*
# **Part 3**: Add age, age squared, and the educational-attainment dummy variables to the previous specification. Do gender differences in age and education explain the gender earnings differential?
# *Answer here*
# **Part 4**: Interact the female dummy variable with all of the other explanatory variables in the regression and add these interaction terms to the specification. Do the returns (that is to say, the effects) to education and age differ by gender? What happens to the coefficient on female?
# *Answer here*
# **Part 5**: Test the null hypothesis that the effects of education and age are the same for men and women. (You’ll need to use an F-test).
# *Answer here*
# **Part 6**: Use the final specification to predict the earnings of...
#
# 1. A 30 year old women with a high school degree
# 2. A 30 year old women with a college degree
# 3. A 40 year old women with a college degree
# 4. A 30 year old man with a high school degree
# 5. A 30 year old man with a college degree, and
# 6. A 40 year old man with a college degree.
#
# What do the prediction indicate happens to the gender earnings differential as educational attainment increases (holding age constant) and as age increases (holding educational attainment constant)?
# *Answer here*
|
GSPP_240B_PS3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Kvotekurver
# ### Norge
# ***
# %matplotlib inline
import pandas as pd
import numpy as np
np.random.seed(666)
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
# ### Dict med alle kvoter, inkl. forslag til kvotekurver, med kilde
# # %load kvotekurver_NO.py
no_curves = [
{'Year': 2012, 'Value': 0.030, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2013, 'Value': 0.049, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2014, 'Value': 0.069, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2015, 'Value': 0.088, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2016, 'Value': 0.108, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2017, 'Value': 0.127, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2018, 'Value': 0.146, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2019, 'Value': 0.165, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2020, 'Value': 0.183, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2021, 'Value': 0.182, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2022, 'Value': 0.181, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2023, 'Value': 0.180, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2024, 'Value': 0.179, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2025, 'Value': 0.176, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2026, 'Value': 0.164, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2027, 'Value': 0.151, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2028, 'Value': 0.132, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2029, 'Value': 0.113, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2030, 'Value': 0.094, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2031, 'Value': 0.075, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2032, 'Value': 0.056, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2033, 'Value': 0.037, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2034, 'Value': 0.018, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2035, 'Value': 0.009, 'Type': 'kvote_norge_0', 'source': {
'url': 'https://www.regjeringen.no/contentassets/033aadf584be44449591210145deb12f/no/pdfs/prp201020110101000dddpdfs.pdf',
'page': 61,
'desc': 'Prop. 101 L, (2010–2011), Proposisjon til Stortinget (forslag til lovvedtak), Lov om elsertifikater'
}
},
{'Year': 2012, 'Value': 0.030, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2013, 'Value': 0.049, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2014, 'Value': 0.069, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2015, 'Value': 0.088, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2016, 'Value': 0.108, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2017, 'Value': 0.127, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2018, 'Value': 0.146, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2019, 'Value': 0.165, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2020, 'Value': 0.183, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2021, 'Value': 0.182, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2022, 'Value': 0.181, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2023, 'Value': 0.180, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2024, 'Value': 0.179, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2025, 'Value': 0.176, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2026, 'Value': 0.164, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2027, 'Value': 0.151, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2028, 'Value': 0.132, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2029, 'Value': 0.113, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2030, 'Value': 0.094, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2031, 'Value': 0.075, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2032, 'Value': 0.056, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2033, 'Value': 0.037, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2034, 'Value': 0.018, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2035, 'Value': 0.009, 'Type': 'kvote_norge_2012', 'source': {
'url': 'https://www.nve.no/Media/4097/elsertifikat-%C3%A5rsrapport-2012_publisering_no.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2012'
}
},
{'Year': 2012, 'Value': 0.030, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2013, 'Value': 0.049, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2014, 'Value': 0.069, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2015, 'Value': 0.088, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2016, 'Value': 0.108, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2017, 'Value': 0.127, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2018, 'Value': 0.146, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2019, 'Value': 0.165, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2020, 'Value': 0.183, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2021, 'Value': 0.182, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2022, 'Value': 0.181, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2023, 'Value': 0.180, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2024, 'Value': 0.179, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2025, 'Value': 0.176, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2026, 'Value': 0.164, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2027, 'Value': 0.151, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2028, 'Value': 0.132, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2029, 'Value': 0.113, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2030, 'Value': 0.094, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2031, 'Value': 0.075, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2032, 'Value': 0.056, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2033, 'Value': 0.037, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2034, 'Value': 0.018, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2035, 'Value': 0.009, 'Type': 'kvote_norge_2013', 'source': {
'url': 'https://www.nve.no/Media/4095/elsertifikat-%C3%A5rsrapport-n-2013web.pdf',
'page': 34,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2013'
}
},
{'Year': 2012, 'Value': 0.030, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2013, 'Value': 0.049, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2014, 'Value': 0.069, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2015, 'Value': 0.088, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2016, 'Value': 0.108, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2017, 'Value': 0.127, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2018, 'Value': 0.146, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2019, 'Value': 0.165, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2020, 'Value': 0.183, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2021, 'Value': 0.182, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2022, 'Value': 0.181, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2023, 'Value': 0.180, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2024, 'Value': 0.179, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2025, 'Value': 0.176, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2026, 'Value': 0.164, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2027, 'Value': 0.151, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2028, 'Value': 0.132, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2029, 'Value': 0.113, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2030, 'Value': 0.094, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2031, 'Value': 0.075, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2032, 'Value': 0.056, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2033, 'Value': 0.037, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2034, 'Value': 0.018, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2035, 'Value': 0.009, 'Type': 'kvote_norge_2014', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2012, 'Value': 0.030, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2013, 'Value': 0.049, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2014, 'Value': 0.069, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2015, 'Value': 0.088, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2016, 'Value': 0.119, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2017, 'Value': 0.137, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2018, 'Value': 0.154, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2019, 'Value': 0.172, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2020, 'Value': 0.197, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2021, 'Value': 0.196, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2022, 'Value': 0.196, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2023, 'Value': 0.195, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2024, 'Value': 0.193, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2025, 'Value': 0.186, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2026, 'Value': 0.174, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2027, 'Value': 0.156, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2028, 'Value': 0.131, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2029, 'Value': 0.109, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2030, 'Value': 0.090, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2031, 'Value': 0.072, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2032, 'Value': 0.054, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2033, 'Value': 0.036, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2034, 'Value': 0.018, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2035, 'Value': 0.009, 'Type': 'kvote_norge_99', 'source': {
'url': 'https://www.nve.no/media/2205/rapport2015_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2014'
}
},
{'Year': 2012, 'Value': 0.030, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2013, 'Value': 0.049, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2014, 'Value': 0.069, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2015, 'Value': 0.088, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2016, 'Value': 0.119, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2017, 'Value': 0.137, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2018, 'Value': 0.154, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2019, 'Value': 0.172, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2020, 'Value': 0.197, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2021, 'Value': 0.196, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2022, 'Value': 0.196, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2023, 'Value': 0.195, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2024, 'Value': 0.193, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2025, 'Value': 0.186, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2026, 'Value': 0.174, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2027, 'Value': 0.156, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2028, 'Value': 0.131, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2029, 'Value': 0.109, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2030, 'Value': 0.090, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2031, 'Value': 0.072, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2032, 'Value': 0.054, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2033, 'Value': 0.036, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2034, 'Value': 0.018, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2035, 'Value': 0.009, 'Type': 'kvote_norge_2015', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2016/rapport2016_51.pdf',
'page': 31,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2015'
}
},
{'Year': 2012, 'Value': 0.030, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2013, 'Value': 0.049, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2014, 'Value': 0.069, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2015, 'Value': 0.088, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2016, 'Value': 0.119, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2017, 'Value': 0.137, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2018, 'Value': 0.154, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2019, 'Value': 0.172, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2020, 'Value': 0.197, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2021, 'Value': 0.196, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2022, 'Value': 0.196, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2023, 'Value': 0.195, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2024, 'Value': 0.193, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2025, 'Value': 0.186, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2026, 'Value': 0.174, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2027, 'Value': 0.156, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2028, 'Value': 0.131, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2029, 'Value': 0.109, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2030, 'Value': 0.090, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2031, 'Value': 0.072, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2032, 'Value': 0.054, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2033, 'Value': 0.036, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2034, 'Value': 0.018, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2035, 'Value': 0.009, 'Type': 'kvote_norge_2016', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2017/rapport2017_52.pdf',
'page': 32,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2016'
}
},
{'Year': 2012, 'Value': 0.030, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2013, 'Value': 0.049, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2014, 'Value': 0.069, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2015, 'Value': 0.088, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2016, 'Value': 0.119, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2017, 'Value': 0.137, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2018, 'Value': 0.153, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2019, 'Value': 0.167, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2020, 'Value': 0.189, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2021, 'Value': 0.189, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2022, 'Value': 0.189, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2023, 'Value': 0.188, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2024, 'Value': 0.186, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2025, 'Value': 0.183, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2026, 'Value': 0.168, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2027, 'Value': 0.153, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2028, 'Value': 0.128, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2029, 'Value': 0.107, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2030, 'Value': 0.083, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2031, 'Value': 0.066, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2032, 'Value': 0.049, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2033, 'Value': 0.033, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2034, 'Value': 0.016, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2035, 'Value': 0.008, 'Type': 'kvote_norge_2017', 'source': {
'url': 'http://publikasjoner.nve.no/rapport/2018/rapport2018_53.pdf',
'page': 30,
'desc': 'Et norsk-svensk elsertifikatmarked, ÅRSRAPPORT FOR 2017'
}
},
{'Year': 2017, 'Value': 0.137, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2018, 'Value': 0.153, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2019, 'Value': 0.167, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2020, 'Value': 0.189, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2021, 'Value': 0.189, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2022, 'Value': 0.189, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2023, 'Value': 0.188, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2024, 'Value': 0.186, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2025, 'Value': 0.183, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2026, 'Value': 0.168, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2027, 'Value': 0.153, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2028, 'Value': 0.128, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2029, 'Value': 0.107, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2030, 'Value': 0.083, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2031, 'Value': 0.066, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2032, 'Value': 0.049, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2033, 'Value': 0.033, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2034, 'Value': 0.016, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2035, 'Value': 0.008, 'Type': 'kvote_norge_2018', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2017-11-27-1835',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2017-11-27-1835'
}
},
{'Year': 2017, 'Value': 0.137, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2018, 'Value': 0.153, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2019, 'Value': 0.171, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2020, 'Value': 0.189, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2021, 'Value': 0.188, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2022, 'Value': 0.188, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2023, 'Value': 0.187, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2024, 'Value': 0.184, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2025, 'Value': 0.177, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2026, 'Value': 0.162, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2027, 'Value': 0.146, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2028, 'Value': 0.125, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2029, 'Value': 0.101, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2030, 'Value': 0.083, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2031, 'Value': 0.066, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2032, 'Value': 0.049, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2033, 'Value': 0.033, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2034, 'Value': 0.016, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
},
{'Year': 2035, 'Value': 0.008, 'Type': 'kvote_norge_2019', 'source': {
'url': 'https://lovdata.no/dokument/LTI/forskrift/2018-10-31-1637',
'page': 'NA',
'desc': 'Forskrift om endring i forskrift om elsertifikater, FOR-2018-10-31-1637'
}
}
]
# +
df_no = pd.DataFrame(no_curves)
df_no['Value'] = df_no['Value'] * 100 # Prosent
df_no
# -
# ### Norge: Plot av alle verdier
# +
plt.rcParams['figure.figsize'] = 30, 10
sns.set_style('ticks')
sns.set_context('poster')
sns.barplot(x='Year', y='Value', data=df_no,
hue='Type')
sns.despine()
plt.xlabel('År')
plt.ylabel('Elsertifikatkvote')
plt.ylim(0, 20)
# -
# ### Norge: Plott av alle verdier (år < 2019)
# +
plt.rcParams['figure.figsize'] = 30, 10
sns.set_style('ticks')
sns.set_context('poster')
sns.barplot(x='Year', y='Value', data=df_no[df_no['Year'] < 2019],
hue='Type')
sns.despine()
plt.xlabel('År')
plt.ylabel('Elsertifikatkvote til og med 2018')
plt.ylim(0, 20)
# -
# ### Norge: Plott av "kvote_norge_2017" (år < 2019)
# <br>
# "kvote_norge_2017": Historisk kvote
# Norwegian flag colors
flag_red = '#EF2B2D' # Deep Carmine Pink
flag_blue = '#002868' # Cool Black
# +
plt.rcParams['figure.figsize'] = 20, 10
sns.set_style('ticks')
sns.set_context('poster')
sns.barplot(x='Year', y='Value',
data=df_no.loc[(df_no['Year'] < 2019) & (df_no['Type'] == "kvote_norge_2017")],
color=flag_blue)
sns.despine()
plt.xlabel('År')
plt.ylabel('%')
plt.title('Elsertifikatkvote Norge: 2012 - 2018')
plt.ylim(0, 20)
# -
# ### Norge: Verdiene fra forrige plott
df_no.loc[(df_no['Year'] < 2019) & (df_no['Type'] == "kvote_norge_2017")]
|
jup_notebooks/kvotekurver/kvotekurver_norge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python2
# name: python2
# ---
# # Discussion 10
#
# ## Web Visualization
#
# Web browsers are ubiquitous and support interactivity through JavaScript. This means the web is an excellent platform for visualizations! The Mozilla Developer Network is a good source for [learning more about web development][web-intro].
#
# When making web visualizations, it helps to know a little bit of JavaScript. Here's a [brief intro][js-intro] and a [more detailed guide][js-guide].
#
# [js-intro]: https://learnxinyminutes.com/docs/javascript/
# [js-guide]: https://developer.mozilla.org/en-US/docs/Web/JavaScript/Guide
# [web-intro]: https://developer.mozilla.org/en-US/docs/Learn
#
# Here are the most popular JavaScript libraries used for web visualizations:
#
# <table><tr>
# <th>Library</th><th>Based On</th><th>Python Support</th><th>Description</th>
# </tr><tr>
# <td>[D3.js](https://d3js.org/)</td><td>-</td><td>[mpld3](http://mpld3.github.io/)</td>
# <td>
# Short for Data-Driven Documents, D3 allows you to bind data to HTML tags.
# In other words, you can use data to control the structure and style of a
# web page.
# </td>
# </tr><tr>
# <td>[Vega](https://vega.github.io/vega/)</td><td>D3.js</td><td>~~vincent~~</td>
# <td>
# A visualization grammar (the same idea as ggplot) built on top of D3. You
# write a description of what you want in JSON, and Vega produces a D3
# visualization.
# </td>
# </tr><tr>
# <td>[Vega Lite](https://vega.github.io/vega-lite/)</td><td>Vega</td><td>[altair](https://altair-viz.github.io/)</td>
# <td>
# A visualization grammar for _common statistical graphics_ built on top of
# Vega. You write a JSON description which is translated to Vega and then D3.
# </td>
# </tr><tr>
# <td>[plotly.js](https://plot.ly/javascript/)</td><td>D3.js</td><td>[plotly](https://plot.ly/python/)</td>
# <td>
# A visualization library that supports the Python, R, Julia, and MATLAB
# plotly packages. Although this is an open-source library, development
# is controlled by Plotly (a private company).
# </td>
# </tr><tr>
# <td>[BokehJS](http://bokeh.pydata.org/en/latest/docs/dev_guide/bokehjs.html)</td><td>-</td><td>[bokeh](http://bokeh.pydata.org/)</td>
# <td>
# A visualization library designed to be used from other (non-JavaScript)
# languages. You write Python, R, or Scala code to produce visualizations.
# </td>
# </tr><tr>
# <td>[Leaflet](http://leafletjs.com/)</td><td>-</td><td>[folium](https://github.com/python-visualization/folium)</td>
# <td>
# An interactive maps library that can display GeoJSON data.
# </td>
# </tr></table>
#
# Also worth mentioning is the [pygal](http://www.pygal.org/en/stable/) package, which produces SVG plots that can be viewed in a web browser and does not rely on any JavaScript library.
# ## Static Visualizations
#
# Let's use Bokeh to make a scatterplot of the diamonds data.
# +
import pandas as pd
diamonds = pd.read_csv("diamonds.csv")
diamonds.head()
# -
# To display Bokeh plots in a Jupyter notebook, you must first call the setup function `output_notebook()`. You don't have to do this if you're going to save your plots to HTML instead.
# +
import bokeh.io
bokeh.io.output_notebook()
# -
# Now we can make a plot. The `bokeh.charts` submodule has functions to create common statistical plots. You can also use functions in the `bokeh.models` submodule to fine-tune plots.
#
# Bokeh's plotting functions work with data frames in [tidy](http://vita.had.co.nz/papers/tidy-data.pdf) form.
# +
import bokeh.charts
plt = bokeh.charts.Scatter(diamonds, x = "carat", y = "price", color = "cut",
webgl = True, tools = "wheel_zoom,pan", active_scroll = "wheel_zoom"
)
bokeh.charts.show(plt)
# Optional: save the plot to a standalone HTML file.
#bokeh.io.output_file("MY_PLOT.html")
# -
# ## Maps
# +
import folium
# Make a map.
m = folium.Map(location = [45.5236, -122.6750])
# Optional: set up a Figure to control the size of the map.
fig = folium.Figure(width = 800, height = 400)
fig.add_child(m)
# Optional: save the map to a standalone HTML file.
# fig.save("MY_MAP.html")
# -
# The Bay Area Rapid Transit (BART) system publishes [data about where its stations are located](http://www.bart.gov/schedules/developers/geo). The data is in KML format, which is an XML format for geospatial data. We can extract the information directly or find a suitable KML reader for Python.
# +
import lxml.etree as lx
# Extract the names and coordinates from the KML file.
xml = lx.parse("bart.kml")
# XML files use namespaces.
ns = {"d": "http://www.opengis.net/kml/2.2"}
places = xml.findall("//d:Placemark", ns)
places = [(p.find("./d:name", ns).text, p.find(".//d:coordinates", ns).text) for p in places]
# Convert to a dataframe, then split the longitude and latitude.
places = pd.DataFrame(places, columns = ["name", "lonlat"])
places.lonlat = places.lonlat.str.split(",", 2)
places["lon"] = places.lonlat.str.get(0).astype(float)
places["lat"] = places.lonlat.str.get(1)
# Latitude is sometimes malformed, with a space and an extra coordinate.
places.lat = places.lat.str.split(" ", 1).str.get(0).astype(float)
places.drop("lonlat", axis = 1, inplace = True)
places.head()
# -
# A GeoDataFrame would also be appropriate. Now we can plot the points on a map.
# +
m = folium.Map(location = [37.8, -122.3], zoom_start = 11)
for name, lon, lat in places.itertuples(index = False):
folium.Marker([lat, lon], popup = name).add_to(m)
fig = folium.Figure(width = 800, height = 400)
fig.add_child(m)
# -
# Folium can also display boundaries stored in GeoJSON files. See the README for more info.
#
# You can use GeoPandas to convert shapefiles to GeoJSON files.
#
# Let's display the distribution of the walrus using [data from the International Union for Conservation of Nature](http://www.iucnredlist.org/technical-documents/spatial-data).
#
# 
# +
m = folium.Map()
m.choropleth(geo_path = "walrus.geojson")
fig = folium.Figure(width = 800, height = 400)
fig.add_child(m)
# -
# ## Interactive Visualizations
#
# In order to make a visualization interactive, you need to run some code when the user clicks on a widget. The code can run _client-side_ on the user's machine, or _server-side_ on your server.
#
# For client-side interactivity:
#
# * Your code must be written in JavaScript.
# * You can host your visualization on any web server. No special setup is needed.
# * Your visualization will use the user's CPU and memory.
#
# For server-side interactivity:
#
# * Your code can be written in any language the server supports. This may require special setup.
# * Your visualization will use the server's CPU and memory.
# * You can update the data in real-time.
# * You can save data submitted by the user.
#
# Shiny is a server-side framework for R. There are lots of server-side frameworks for Python. Two of the most popular are [Django][django] and [Flask][flask].
#
# [django]: https://www.djangoproject.com/
# [flask]: http://flask.pocoo.org/
# ### Client-side
#
# Client-side interactivity is cheaper to get started with because you can use a free web server (like GitHub Pages).
#
# Let's make the diamonds plot interactive so that the user can select which variables get plotted. Unfortunately, Bokeh charts don't work with interactivity, so we have to build the plot with simpler functions. We'll lose the color-coding, although you could still add that with a bit more work.
diamonds.head()
# +
import bokeh.layouts
import bokeh.models
import bokeh.plotting
original = bokeh.models.ColumnDataSource(diamonds)
source = bokeh.models.ColumnDataSource({"x": diamonds.carat, "y": diamonds.price})
plt = bokeh.plotting.figure(tools = [], webgl = True)
plt.circle("x", "y", source = source)
# Set up selector boxes.
numeric_cols = ["carat", "depth", "table", "price", "x", "y", "z"]
sel_x = bokeh.models.widgets.Select(title = "x-axis", options = numeric_cols, value = "carat")
sel_y = bokeh.models.widgets.Select(title = "y-axis", options = numeric_cols, value = "price")
# Callback for x selector box.
callback_x = bokeh.models.CustomJS(args = {"original": original, "source": source}, code = """
// This is the JavaScript code that will run when the x selector box is changed.
// You can use the alert() function to "print" values.
alert(cb_obj.value);
source.data['x'] = original.data[cb_obj.value];
source.trigger('change');
""")
sel_x.js_on_change("value", callback_x)
# Callback for y selector box.
callback_y = bokeh.models.CustomJS(args = {"original": original, "source": source}, code = """
// This is the JavaScript code that will run when the y selector box is changed.
source.data['y'] = original.data[cb_obj.value];
source.trigger('change');
""")
sel_y.js_on_change("value", callback_y)
# Position the selector boxes to the right of the plot.
layout = bokeh.layouts.column(sel_x, sel_y)
layout = bokeh.layouts.row(plt, layout)
bokeh.charts.show(layout)
# -
# ### Server-side
#
# Server-side interactivity is a lot more flexible. Flask is a simple framework with great documentation, so it's easy to get started with.
#
# A demo flask website is available at: <https://github.com/nick-ulle/flask-demo>
#
# The core of a flask website (or "app") is a script with functions that return the text that should be displayed on each page.
#
# ```python
# # gh_barplot.py
# import flask
# from flask import Flask
#
# import gh_events
#
# # Set up a Flask app.
# app = Flask(__name__)
#
# # This function returns the "/" page.
# @app.route("/")
# def index():
# events = gh_events.fetch()
# events = gh_events.parse_events(events)
# script, div = gh_events.bar_plot_types(events)
# # Substitute values into the `index.html` template file.
# return flask.render_template("index.html", script = script, div = div)
#
# # This function returns the "/hello1" page.
# @app.route("/hello<int:n>")
# def hello(n):
# if n == 1:
# return "Hello, world!"
# else:
# return "Hello, all {} worlds!".format(n)
# ```
# This website also uses another script, `gh_events.py` to fetch data from GitHub's API. The `gh_events.py` script is a regular Python script and doesn't contain any flask code.
#
# ```python
# # gh_events.py
# import bokeh, bokeh.charts
# from bokeh.plotting import figure
# from bokeh.embed import components
# import pandas as pd
# import requests
#
# # Fetch events from the GitHub API.
# def fetch():
# response = requests.get("https://api.github.com/events")
# response.raise_for_status()
#
# return response.json()
#
# # Parse the event data into a data frame.
# def parse_events(events):
# data = (
# (evt['type'], evt['actor']['login'], evt['repo']['name'])
# for evt in events
# )
#
# return pd.DataFrame.from_records(data, columns = ["Type", "User", "Repo"])
#
# # Make a Bokeh bar plot of the event types.
# def bar_plot_types(events):
# plot = bokeh.charts.Bar(events, "Type")
#
# return bokeh.embed.components(plot)
# ```
# The website's homepage is based on the template file `index.html`. This file uses [Jinja](http://jinja.pocoo.org/) syntax to indicate where substitutions should be made.
#
# ```html
# <!-- index.html -->
# <html>
# <head>
# <!-- Bokeh CSS & JavaScript Files -->
# <link
# href="http://cdn.pydata.org/bokeh/release/bokeh-0.12.4.min.css"
# rel="stylesheet" type="text/css">
# <link
# href="http://cdn.pydata.org/bokeh/release/bokeh-widgets-0.12.4.min.css"
# rel="stylesheet" type="text/css">
#
# <script src="http://cdn.pydata.org/bokeh/release/bokeh-0.12.4.min.js">
# </script>
# <script src="http://cdn.pydata.org/bokeh/release/bokeh-widgets-0.12.4.min.js">
# </script>
# <!-- End of Bokeh Files -->
# {{script|safe}}
# </head>
#
# <body>
# This is the display.
# {{div|safe}}
# </body>
# </html>
# ```
|
sta141b/2017/discussion10.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Django Shell-Plus
# language: python
# name: django_extensions
# ---
# +
import sys
sys.path.insert(0, '../src/')
sys.path.insert(0, '../')
import django
django.setup()
# +
from importlib import reload
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import ipywidgets as widgets
from IPython.display import display, clear_output
from ipywidgets import interact, interact_manual, Layout
# +
import luminol.anomaly_detector as lad
from apps.utils.time import UTC_P0100
import apps.ad.anomaly_detection as ad
from apps.mc.api.util import get_topics, get_property, get_time_slots, get_features_of_interest, get_aggregating_process, get_observation_getter
from psycopg2.extras import DateTimeTZRange
# -
phenomenon_date_from = pd.to_datetime("2019-01-01")
phenomenon_date_to = pd.to_datetime("2019-03-30")
d_topic = get_topics()[0]
d_prop = get_property(d_topic)[0]
d_feature = get_features_of_interest(d_topic, d_prop)[0]
d_time_slot = get_time_slots(d_topic)[0]
# + pixiedust={"displayParams": {}}
def detect_anomalies(
phenomenon_date_from = phenomenon_date_from,
phenomenon_date_to = phenomenon_date_to,
detector_method='bitmap_mod',
use_baseline=True,
shift=True,
extend_range=True,
detector_params={
"precision": 6,
"lag_window_size": 96,
"future_window_size": 96,
"chunk_size": 2
},
topic = d_topic,
prop = d_prop,
feature = d_feature,
time_slot = d_time_slot
):
pt_range_z = DateTimeTZRange(
pd.to_datetime(phenomenon_date_from).replace(tzinfo=UTC_P0100),
pd.to_datetime(phenomenon_date_to).replace(tzinfo=UTC_P0100)
)
get_func, feature_time_slots = get_observation_getter(
topic,
prop,
time_slot,
feature,
pt_range_z
)
anoms = ad.get_timeseries(
phenomenon_time_range=pt_range_z,
num_time_slots=len(feature_time_slots),
get_observations=get_func,
detector_method=detector_method,
detector_params=detector_params,
shift=shift,
use_baseline=use_baseline,
extend_range=extend_range,
)
anoms["feature_time_slots"] = feature_time_slots
return anoms
# -
def highlight(indices, alpha, color, ax):
i=0
while i<len(indices):
ax.axvspan(indices[i]-0.5, indices[i]+0.5, facecolor=color, edgecolor='none', alpha=alpha)
i+=1
# +
colors = ['r', 'g', 'c', 'm', 'y', 'k']
results = []
plt.ioff()
def plot(detectors, hlt_detector):
results = detectors
plt.close()
fig, ax1 = plt.subplots(figsize=(20,7))
hs = pd.DataFrame({
'anomalies': detectors[hlt_detector]["property_anomaly_rates"]
})
if detectors[hlt_detector]["property_anomaly_percentiles"]:
perc = detectors[hlt_detector]["property_anomaly_percentiles"]
color = colors[list(detectors.keys()).index(hlt_detector)]
for p in perc.keys():
highlight(hs[hs['anomalies'] > perc[p]].index, p*0.0025, color, ax1)
first_result = detectors[list(detectors.keys())[0]]
lns = []
if first_result["property_values"]:
ts = pd.DataFrame({
'values': [float(n) if n is not None else n for n in first_result["property_values"]]
}, index=[n.lower.strftime("%-d.%-m.%Y") for n in first_result["feature_time_slots"]])
values_line = ts['values'].plot.line(ax=ax1, color='b')
ax1.set_ylabel('values', color='b')
ax1.tick_params('y', colors='b')
lns.append(values_line.get_lines()[0])
for i in range(len(detectors.keys())):
detector = list(detectors.keys())[i]
color = colors[i]
anomalies = detectors[detector]["property_anomaly_rates"]
if anomalies:
ts = pd.DataFrame({
'anomalies': anomalies
})
ax2 = ax1.twinx()
anomalies_line = ts['anomalies'].plot.line(ax=ax2, color=color, label=detector)
lns.append(anomalies_line.get_lines()[0])
ax2.tick_params('y', colors=color)
if lns:
labs = [ln.get_label() for ln in lns]
ax1.legend(lns, labs, loc=1)
# baseName = f"{baserange.lower.date()}..{baserange.upper.date()}"
# if first_result['phenomenon_time_range']:
# rangeName = f"{first_result['phenomenon_time_range'].lower.date()}..{first_result['phenomenon_time_range'].upper.date()}"
# plt.savefig(f"graphs/{baseName}_{rangeName}_window-{str(window_size)}_prec-{str(detector_params['precision'])}.png", format="png")
# plt.savefig(f"graphs/{rangeName}_window-{str(window_size)}_prec-{str(detector_params['precision'])}.png", format="png")
return fig
# -
detectors = {
"Bitmap mod": "bitmap_mod",
"Bitmap diminishing": "bitmap_diminishing",
"Bitmap diminishing baseline": "bitmap_diminishing_bl",
"Bitmap mod shift": "bitmap_mod_shift",
"LinkedIn bitmap": "bitmap_detector",
"Default": "default_detector",
"Derivative": "derivative_detector",
"Exponential average": "exp_avg_detector",
# "Absolute threshold": "absolute_threshold",
# "Diff Percent": "diff_percent_threshold",
# "Sign test": "sign_test",
}
# +
def val_bitmap_mod(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(start_date, end_date, "bitmap_mod", shift=False, feature=feature, prop=prop, time_slot=time_slot, detector_params=detector_params)
def val_bitmap_diminishing(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(start_date, end_date, "bitmap_diminishing", shift=False, use_baseline=False, feature=feature, prop=prop, time_slot=time_slot, detector_params=detector_params)
def val_bitmap_diminishing_bl(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(start_date, end_date, "bitmap_diminishing", shift=False, feature=feature, prop=prop, time_slot=time_slot, detector_params=detector_params)
def val_bitmap_mod_shift(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(start_date, end_date, "bitmap_mod_shift", feature=feature, prop=prop, time_slot=time_slot, detector_params=detector_params)
def val_bitmap_detector(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(start_date, end_date, "bitmap_detector", shift=False, use_baseline=False, feature=feature, prop=prop, time_slot=time_slot, detector_params=detector_params)
def val_default_detector(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(start_date, end_date, "default_detector", feature=feature, prop=prop, time_slot=time_slot, detector_params={}, use_baseline=False, extend_range=False, shift=False)
def val_derivative_detector(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(start_date, end_date, "derivative_detector", feature=feature, prop=prop, time_slot=time_slot, detector_params={}, use_baseline=False, extend_range=False, shift=False)
def val_exp_avg_detector(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(start_date, end_date, "exp_avg_detector", feature=feature, prop=prop, time_slot=time_slot, detector_params={}, use_baseline=False, extend_range=False, shift=False)
def val_absolute_threshold(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(t_from, t_to, "absolute_threshold", feature=feature, prop=prop, time_slot=time_slot, detector_params={}, use_baseline=False, extend_range=False, shift=False)
def val_diff_percent_threshold(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(t_from, t_to, "diff_percent_threshold", feature=feature, prop=prop, time_slot=time_slot, detector_params={}, use_baseline=False, extend_range=False, shift=False)
def val_sign_test(feature, prop, time_slot, start_date, end_date, detector_params):
return detect_anomalies(t_from, t_to, "sign_test", feature=feature, prop=prop, time_slot=time_slot, detector_params={}, use_baseline=False, extend_range=False, shift=False)
def plot_anomalies(feature, prop, time_slot, start_date, end_date, precision, window_size, chunk_size, hlt_detector, bitmap_mod, bitmap_diminishing, bitmap_diminishing_bl, bitmap_mod_shift, bitmap_detector, default_detector, derivative_detector, exp_avg_detector):
args = [feature, prop, time_slot, start_date, end_date, {
"precision": precision,
"lag_window_size": window_size,
"future_window_size": window_size,
"chunk_size": chunk_size
}]
anomalies = {}
if bitmap_mod:
anomalies["bitmap_mod"] = val_bitmap_mod(*args)
if bitmap_diminishing:
anomalies["bitmap_diminishing"] = val_bitmap_diminishing(*args)
if bitmap_diminishing_bl:
anomalies["bitmap_diminishing_bl"] = val_bitmap_diminishing_bl(*args)
if bitmap_mod_shift:
anomalies["bitmap_mod_shift"] = val_bitmap_mod_shift(*args)
if bitmap_detector:
anomalies["bitmap_detector"] = val_bitmap_detector(*args)
if default_detector:
anomalies["default_detector"] = val_default_detector(*args)
if derivative_detector:
anomalies["derivative_detector"] = val_derivative_detector(*args)
if exp_avg_detector:
anomalies["exp_avg_detector"] = val_exp_avg_detector(*args)
# if absolute_threshold:
# anomalies["absolute_threshold"] = val_absolute_threshold(*args)
# if diff_percent_threshold:
# anomalies["diff_percent_threshold"] = val_diff_percent_threshold(*args)
# if sign_test:
# anomalies["sign_test"] = val_sign_test(*args)
if len(anomalies.keys()) > 1:
i = 0
while hlt_detector not in anomalies.keys():
hlt_detector = detectors[i]
i += 1
return plot(anomalies, hlt_detector)
# +
t_from = "2019-01-01"
t_to = "2019-03-30"
def hlt_detectors():
d = {}
if bitmap_mod_widget.value: d["Bitmap mod"] = "bitmap_mod"
if bitmap_diminishing_widget.value: d["Bitmap diminishing"] = "bitmap_diminishing"
if bitmap_diminishing_bl_widget.value: d["Bitmap diminishing baseline"] = "bitmap_diminishing_bl"
if bitmap_mod_shift_widget.value: d["Bitmap mod shift"] = "bitmap_mod_shift"
if bitmap_detector_widget.value: d["LinkedIn bitmap"] = "bitmap_detector"
if default_detector_widget.value: d["Default"] = "default_detector"
if derivative_detector_widget.value: d["Derivative"] = "derivative_detector"
if exp_avg_detector_widget.value: d["Exponential average"] = "exp_avg_detector"
# if absolute_threshold_widget.value: d["Absolute threshold"] = "absolute_threshold"
# if diff_percent_threshold_widget.value: d["Diff Percent"] = "diff_percent_threshold"
# if sign_test_widget.value: d["Sign test"] = "sign_test"
return d
def update_hlt_detectors(*args):
hlt_detector_widget.options = hlt_detectors()
def update_property_widget(*args):
property_widget.options = get_property(topic_widget.value)
def update_feature_widget(*args):
feature_widget.options = get_features_of_interest(topic_widget.value, property_widget.value)
def update_time_slots_widget(*args):
time_slots_widget.options = get_time_slots(topic_widget.value)
# +
bitmap_mod_widget = widgets.Checkbox(value=False,description="Bitmap mod")
bitmap_mod_widget.observe(update_hlt_detectors, "value")
bitmap_diminishing_widget = widgets.Checkbox(value=True,description="Bitmap diminishing")
bitmap_diminishing_widget.observe(update_hlt_detectors, "value")
bitmap_diminishing_bl_widget = widgets.Checkbox(value=True,description="Bitmap diminishing baseline")
bitmap_diminishing_bl_widget.observe(update_hlt_detectors, "value")
bitmap_mod_shift_widget = widgets.Checkbox(value=False,description="Bitmap mod shift")
bitmap_mod_shift_widget.observe(update_hlt_detectors, "value")
bitmap_detector_widget = widgets.Checkbox(value=False,description="LinkedIn bitmap")
bitmap_detector_widget.observe(update_hlt_detectors, "value")
default_detector_widget = widgets.Checkbox(value=False,description="Default")
default_detector_widget.observe(update_hlt_detectors, "value")
derivative_detector_widget = widgets.Checkbox(value=False,description="Derivative")
derivative_detector_widget.observe(update_hlt_detectors, "value")
exp_avg_detector_widget = widgets.Checkbox(value=False,description="Exponential average")
exp_avg_detector_widget.observe(update_hlt_detectors, "value")
# absolute_threshold_widget = widgets.Checkbox(value=False,description="Absolute threshold")
# absolute_threshold_widget.observe(update_hlt_detectors, "value")
# diff_percent_threshold_widget = widgets.Checkbox(value=False,description="Diff Percent")
# diff_percent_threshold_widget.observe(update_hlt_detectors, "value")
# sign_test_widget = widgets.Checkbox(value=False,description="Sign test")
# sign_test_widget.observe(update_hlt_detectors, "value")
detector_widgets = widgets.HBox([
widgets.VBox([
bitmap_mod_widget,
bitmap_diminishing_widget,
bitmap_diminishing_bl_widget,
bitmap_detector_widget,
]),
widgets.VBox([
bitmap_mod_shift_widget,
default_detector_widget,
derivative_detector_widget,
exp_avg_detector_widget,
]),
# widgets.VBox([
# absolute_threshold_widget,
# diff_percent_threshold_widget,
# sign_test_widget
# ])
])
hlt_detector_widget = widgets.Dropdown(options=hlt_detectors(), value=detectors["Bitmap diminishing"], description="Highlight")
precision_widget = widgets.IntSlider(value=6, min=2, max=16, step=1, description="Precision")
window_size_widget = widgets.BoundedIntText(value=96, min=4, max=256, step=1, description="Window size")
chunk_size_widget = widgets.IntSlider(value=2, min=2, max=16, step=1, description="Chunk size")
topic_widget = widgets.Dropdown(options=get_topics(), description="Topic")
topic_widget.observe(update_property_widget, "value")
topic_widget.observe(update_feature_widget, "value")
topic_widget.observe(update_time_slots_widget, "value")
property_widget = widgets.Dropdown(options=get_property(topic_widget.value), description="Property")
property_widget.observe(update_feature_widget, "value")
feature_widget = widgets.Dropdown(options=get_features_of_interest(topic_widget.value, property_widget.value), description="Station")
time_slot_widget = widgets.Dropdown(options=get_time_slots(topic_widget.value), description="Aggregate to")
start_date_widget = widgets.DatePicker(value=pd.to_datetime(t_from).date(), description="Start date")
end_date_widget = widgets.DatePicker(value=pd.to_datetime(t_to).date(), description="End date")
date_widgets = widgets.HBox([
start_date_widget,
end_date_widget
])
# ui = widgets.Tab(children=[
widget_accordion = widgets.Accordion(children=[
widgets.VBox([
widgets.HBox([topic_widget, property_widget]),
widgets.HBox([feature_widget, time_slot_widget]),
date_widgets
]),
widgets.VBox([
precision_widget,
window_size_widget,
chunk_size_widget
]),
widgets.VBox([
detector_widgets,
hlt_detector_widget
])
],
# layout=Layout(
# height='300px',
# display='flex',
# align_items='center',
# justify_content='center'
# )
)
widget_accordion.set_title(0, "General")
widget_accordion.set_title(1, "Detector parameters")
widget_accordion.set_title(2, "Detectors used")
widget_accordion.selected_index = 0
out = widgets.Output()
plot_button = widgets.Button(
description="Plot"
)
def click(b):
fig = plot_anomalies(
bitmap_mod=bitmap_mod_widget.value,
bitmap_diminishing=bitmap_diminishing_widget.value,
bitmap_diminishing_bl=bitmap_diminishing_bl_widget.value,
bitmap_mod_shift=bitmap_mod_shift_widget.value,
bitmap_detector=bitmap_detector_widget.value,
default_detector=default_detector_widget.value,
derivative_detector=derivative_detector_widget.value,
exp_avg_detector=exp_avg_detector_widget.value,
# absolute_threshold=absolute_threshold_widget.value,
# diff_percent_threshold=diff_percent_threshold_widget.value,
# sign_test=sign_test_widget.value,
feature=feature_widget.value,
prop=property_widget.value,
time_slot=time_slot_widget.value,
start_date=start_date_widget.value,
end_date=end_date_widget.value,
precision=precision_widget.value,
window_size=window_size_widget.value,
chunk_size=chunk_size_widget.value,
hlt_detector=hlt_detector_widget.value
)
with out:
clear_output(wait=True)
display(fig)
plot_button.on_click(click)
ui = widgets.VBox([
widget_accordion,
plot_button,
out
])
def init():
reload(ad)
reload(lad)
display(ui)
click(None)
print("Aggregating process:", get_aggregating_process(topic_widget.value, property_widget.value, feature_widget.value))
# -
init()
|
jupyter-notebooks/AD Experimentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# # Graph Fraud Detection with DGL on Amazon SageMaker
# +
import sagemaker
from sagemaker import get_execution_role
role = get_execution_role()
sess = sagemaker.Session()
# -
# ## Data Preprocessing and Feature Engineering
# ### Upload raw data to S3
#
# The dataset we use is the [IEEE-CIS Fraud Detection dataset](https://www.kaggle.com/c/ieee-fraud-detection/data?select=train_transaction.csv) which is a typical example of financial transactions dataset that many companies have. The dataset consists of two tables:
#
# * **Transactions**: Records transactions and metadata about transactions between two users.
# * **Identity**: Contains information about the identity users performing transactions
#
# Now let's move the raw data to a convenient location in the S3 bucket for this proejct, where it will be picked up by the preprocessing job and training job.
#
# If you would like to use your own dataset for this demonstration. Replace the `raw_data_location` with the s3 path or local path of your dataset.
# +
# Replace with an S3 location or local path to point to your own dataset
raw_data_location = 's3://sagemaker-solutions-us-west-2/Fraud-detection-in-financial-networks/data'
bucket = 'SAGEMAKER_S3_BUCKET'
prefix = 'dgl'
input_data = 's3://{}/{}/raw-data'.format(bucket, prefix)
# !aws s3 cp --recursive $raw_data_location $input_data
# Set S3 locations to store processed data for training and post-training results and artifacts respectively
train_data = 's3://{}/{}/processed-data'.format(bucket, prefix)
train_output = 's3://{}/{}/output'.format(bucket, prefix)
# -
# ### Build container for Preprocessing and Feature Engineering
#
# Data preprocessing and feature engineering is an important component of the ML lifecycle, and Amazon SageMaker Processing allows you to do these easily on a managed infrastructure. Now, we'll create a lightweight container that will serve as the environment for our data preprocessing. The container can also be easily customized to add in more dependencies if our preprocessing job requires it.
# +
import boto3
region = boto3.session.Session().region_name
account_id = boto3.client('sts').get_caller_identity().get('Account')
ecr_repository = 'sagemaker-preprocessing-container'
ecr_repository_uri = '{}.dkr.ecr.{}.amazonaws.com/{}:latest'.format(account_id, region, ecr_repository)
# !bash data-preprocessing/container/build_and_push.sh $ecr_repository docker
# -
# ### Run Preprocessing job with Amazon SageMaker Processing
#
# The script we have defined at `data-preprocessing/graph_data_preprocessor.py` performs data preprocessing and feature engineering transformations on the raw data. Some of the data transformation and feature engineering techniques include:
#
# * Performing numerical encoding for categorical variables and logarithmic transformation for transaction amount
# * Constructing graph edgelists between transactions and other entities for the various relation types
# +
from sagemaker.processing import ScriptProcessor, ProcessingInput, ProcessingOutput
script_processor = ScriptProcessor(command=['python3'],
image_uri=ecr_repository_uri,
role=role,
instance_count=1,
instance_type='SAGEMAKER_PROCESSING_INSTANCE_TYPE')
script_processor.run(code='data-preprocessing/graph_data_preprocessor.py',
inputs=[ProcessingInput(source=input_data,
destination='/opt/ml/processing/input')],
outputs=[ProcessingOutput(destination=train_data,
source='/opt/ml/processing/output')],
arguments=['--id-cols', 'card1,card2,card3,card4,card5,card6,ProductCD,addr1,addr2,P_emaildomain,R_emaildomain',
'--cat-cols','M1,M2,M3,M4,M5,M6,M7,M8,M9'])
# -
# ### View Results of Data Preprocessing
#
# Once the preprocessing job is complete, we can take a look at the contents of the S3 bucket to see the transformed data. We have a set of bipartite edge lists between transactions and different device id types as well as the features, labels and a set of transactions to validate our graph model performance.
# +
from os import path
from sagemaker.s3 import S3Downloader
processed_files = S3Downloader.list(train_data)
print("===== Processed Files =====")
print('\n'.join(processed_files))
# optionally download processed data
# S3Downloader.download(train_data, train_data.split("/")[-1])
# -
# ## Train Graph Neural Network with DGL
#
# Graph Neural Networks work by learning representation for nodes or edges of a graph that are well suited for some downstream task. We can model the fraud detection problem as a node classification task, and the goal of the graph neural network would be to learn how to use information from the topology of the sub-graph for each transaction node to transform the node's features to a representation space where the node can be easily classified as fraud or not.
#
# Specifically, we will be using a relational graph convolutional neural network model (R-GCN) on a heterogeneous graph since we have nodes and edges of different types.
# ### Hyperparameters
#
# To train the graph neural network, we need to define a few hyperparameters that determine:
#
# * The kind of graph we're constructing
# * The class of graph neural network models we will be using
# * The network architecture
# * The optimizer and optimization parameters
#
# +
edges = ",".join(map(lambda x: x.split("/")[-1], [file for file in processed_files if "relation" in file]))
params = {'nodes' : 'features.csv',
'edges': 'relation*',
'labels': 'tags.csv',
'model': 'rgcn',
'num-gpus': 1,
'batch-size': 10000,
'embedding-size': 64,
'n-neighbors': 1000,
'n-layers': 2,
'n-epochs': 10,
'optimizer': 'adam',
'lr': 1e-2
}
print("Graph will be constructed using the following edgelists:\n{}" .format('\n'.join(edges.split(","))))
# -
# ### Create and Fit SageMaker Estimator
#
# With the hyperparameters defined, we can kick off the training job. We will be using the Deep Graph Library (DGL), with MXNet as the backend deep learning framework, to define and train the graph neural network. Amazon SageMaker makes it do this with the Framework estimators which have the deep learning frameworks already setup. Here, we create a SageMaker MXNet estimator and pass in our model training script, hyperparameters, as well as the number and type of training instances we want.
#
# We can then `fit` the estimator on the the training data location in S3.
# +
from sagemaker.mxnet import MXNet
estimator = MXNet(entry_point='train_dgl_mxnet_entry_point.py',
source_dir='dgl-fraud-detection',
role=role,
train_instance_count=1,
train_instance_type='SAGEMAKER_TRAINING_INSTANCE_TYPE',
framework_version="1.6.0",
py_version='py3',
hyperparameters=params,
output_path=train_output,
code_location=train_output,
sagemaker_session=sess)
estimator.fit({'train': train_data})
# -
# Once the training is completed, the training instances are automatically saved and SageMaker stores the trained model and evaluation results to a location in S3.
#
|
source/sagemaker/dgl-fraud-detection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/ranma-26/github-slideshow/blob/master/Untitled0.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pnwlAq3cgSeh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fdb4ac48-903a-4771-ec47-8733591ad590"
1+1
# + id="OFWk8ydCjJ28" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a1e212a3-d5f0-4c78-d00d-7203ef032dd5"
100-1
# + id="Yf1MRXXwjW5c" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="e26394fb-4ce1-4bad-a3b7-0c69242466ef"
28*28
# + id="W6pjsShsjaJT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="aeb52f6f-0298-4660-bb7d-1efda11ecee7"
8473/53
# + id="u1ZpLj-_ks6n" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="38e90571-c056-4d8e-b255-6efac6238077"
((4890+2448)*5 + 998)*1.1
# + [markdown] id="rwZxN4g9laay" colab_type="text"
# 4,890円のRaspberry Piを5つと、 2,448円のSDを5つ、998円のHDMIケーブルを1つ買った場合の合計金額(消費税10%込)はいくらでしょうか?
# + id="chtl1deXlgvj" colab_type="code" colab={}
# + [markdown] id="F75HnPEZlros" colab_type="text"
# プログラムは小数点以下を自動では四捨五入してくれないので端数が出てしまいました。小数点以下を四捨五入するにはround関数を使います。
# + [markdown] id="jyUKgDyelx8A" colab_type="text"
#
# + id="d9eMRaAVlyVU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="964f486a-8df2-42d4-8184-00e80f027f20"
round(((4890+2448)*5 + 998)*1.1)
# + [markdown] id="hMhEsvnpnZNM" colab_type="text"
# >>> raspi = 4890
# >>> sdcard = 2448
# >>> hdmicable = 998
#
# + id="BPnte0ZnneFA" colab_type="code" colab={}
raspi = 4890
sdcard = 2448
hdmicable = 998
# + id="0shqtNaWqN_1" colab_type="code" colab={}
# + [markdown] id="TuzIfHdMqOik" colab_type="text"
# プログラムで元々ある言葉を予約語という
# + id="Pmpp85QprpTk" colab_type="code" colab={}
cart = raspi * 5
# + id="aiXCIIylrsI-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="73e8f9d4-4f02-43dc-a8d1-d9fa47981aaa"
cart
# + id="6rOwI2OBr3px" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="b3a89db6-95d9-46d1-d866-856043ff0c13"
cart += sdcard * 5
cart
# + id="Tmv1dtVqsD3B" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fdb1127e-f16c-48a4-f02f-97a7e87905fb"
cart
# + id="03Sb_rYCsMuM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="394352b2-0c50-4eac-d8cb-a069ede94d06"
cart += hdmicable * 1
cart
# + id="qMc62NbjsZPt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c2ed127f-1231-4365-808a-430875e4de40"
total = cart * 1.1
total
# + id="8TG53q3it72j" colab_type="code" colab={}
# + id="vZ7G-gA9sfGr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6e8c7374-7298-4cba-bcfd-7d4e9ce16d05"
# + id="WnGHBlKEtzXr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="69c385e9-18b7-42f2-8c97-b96d62e9364d"
type(cart)
# + id="LgYfaoK5t3Rp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="dc7c9941-50f0-431f-8804-b72c4a14da31"
type(total)
# + id="Rdb-93SHt9rr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="ac907f5d-e790-447f-c1ed-a3c1019f53f4"
total = round(cart * 1.1)
total
# + id="S04Zn4JDuDPJ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bc8bd510-6e03-406a-a163-f8aaf88e1ef5"
type(cart)
|
Untitled0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Likevektskonsentrasjoner for en diprotisk syre
# Her skal vi gjøre eksempel 16.10 fra læreboken (side 567) og finne likevektskonsentrasjoner for 0.10 M av den diprotiske syren oksalsyre ($\text{H}_2 \text{C}_2 \text{O}_4$):
#
# $$\text{H}_2 \text{C}_2 \text{O}_4 \rightleftharpoons \text{HC}_2 \text{O}_4^{-} + \text{H}^{+},\quad K_{a,1} = 6.5 \times 10^{-2}$$
#
# $$\text{HC}_2 \text{O}_4^- \rightleftharpoons \text{C}_2 \text{O}_4^{2-} + \text{H}^{+},\quad K_{a,1} = 6.1 \times 10^{-5}$$
#
# Vi skal løse denne oppgaven ved å bruke Python. For å kunne regne symbolsk skal vi bruke et bibliotek som heter [SymPy](https://www.sympy.org/).
import sympy as sym # Importer SymPy
# Definer størrelsene vi kjenner
START_KONSENTRASJON = 0.10
KA1 = 6.5e-2
KA2 = 6.1e-5
Kw = 1e-14 # For vann
# Over har vi listet opp hva vi kjenner. La oss også liste opp alle de ukjente som vi skal bestemme (ved likevekt):
# - $[\text{H}_2 \text{C}_2 \text{O}_4]$
# - $[\text{HC}_2 \text{O}_4^{-}]$
# - $[\text{H}^{+}]$
# - $[\text{C}_2 \text{O}_4^{2-}]$
# - $[\text{OH}^-]$
#
# Vi har altså 5 ukjente. La oss definere de som størrelser (spesifikt som [SymPy-symboler](https://docs.sympy.org/latest/tutorial/intro.html#a-more-interesting-example)) slik at vi kan regne med de (dette blir litt som når vi introduserer $x$ osv. for ukjente størrelser i ligninger vi skriver for hånd):
# Vi definerer de ukjente størrelsene. For å spare litt skriving bruker vi
# - HHA for syren H2C2O4
# - HA for syren HCO4
# - A for den korresponderende basen C2O4^2-
# - H for H+ :-)
# - Vi tar også med OH for OH-, siden eksempelet i læreboka bruker den
c_HHA, c_HA, c_A, c_H, c_OH = sym.symbols('c_HHA c_HA c_A c_H c_OH')
# Vi har nå definert konsentrasjonene. Disse er foreløpig ukjente. For å bestemme de, så trenger vi noen ligninger som relaterer de til hverandre. Mulige slike ligninger er:
# - syre-basekonstanten
# - elektronøytralitet
# - massebalanser
# La oss begynne med syre-basekonstantene:
ligning1 = sym.Eq((c_HA * c_H)/c_HHA, KA1)
ligning2 = sym.Eq((c_A * c_H)/c_HA, KA2)
ligning3 = sym.Eq(c_H * c_OH, Kw)
ligning1
ligning2
ligning3
# Den neste ligningen vi kan benytte oss av, er at det må være like mye negativ og positiv ladning. Her er det 4 ladede forbindelser:
# - negative: $\text{HC}_2 \text{O}_4^{-}$, $\text{C}_2 \text{O}_4^{2-}$ (merk at denne har ladning $-2$) og $[\text{OH}^-]$
# - positive: $\text{H}^+$
#
# Summen av ladninger er null. Det betyr at konsentrasjonen av positive ladninger er like stor som
# konsentrasjonen av negative ladninger:
#
# $$[\text{H}]^+ = 2 [\text{C}_2 \text{O}_4]^{2-} + [\text{HC}_2 \text{O}_4]^- + [\text{OH}]^-.$$
#
# (Merk igjen faktoren $2$ som tar hensyn til ladningen på $-2$)
#
# La oss skrive det som en ligning med symbolene vi har definert:
# Elektronøytralitet:
ligning4 = sym.Eq(c_HA + 2 * c_A + c_OH, c_H)
ligning4
# Når det gjelder massebalanse, så har vi mange valg. La oss bruke massen av karbon. Vi vet at det ikke dannes noe ekstra masse i denne reaksjonen. Det betyr at massen av karbon vi startet med er lik massen av karbon ved likevekt. Skrevet med konsentrasjoner blir dette:
#
# $$2 [\text{H}_2 \text{C}_2 \text{O}_4]_{\text{start}} = 2 [\text{H}_2 \text{C}_2 \text{O}_4]_{\text{likevekt}} + 2 [\text{HC}_2 \text{O}_4^-]_{\text{likevekt}} + 2 [\text{C}_2 \text{O}_4^{2-}]_{\text{likevekt}}$$
#
# Faktoren $2$ tar hensyn til at det er to stk. karbon in hver forbindelse. Her er dette en felles faktor, så vi kan
# dele den bort:
#
# $$[\text{H}_2 \text{C}_2 \text{O}_4]_{\text{start}} = [\text{H}_2 \text{C}_2 \text{O}_4]_{\text{likevekt}} + [\text{HC}_2 \text{O}_4^-]_{\text{likevekt}} + [\text{C}_2 \text{O}_4^{2-}]_{\text{likevekt}}$$
#
# La oss formulere det som en ligning:
# Massebalanse for karbon:
ligning5 = sym.Eq(START_KONSENTRASJON, c_HHA + c_HA + c_A)
ligning5
# Vi har nå 5 ligninger og vi har 5 ukjente. Dette kan vi (eller i dette tilfellet, SymPy) løse. Her skal vi be om en numerisk løsning siden dette er raskere enn å få SymPy til å regne symbolsk.
#
# For å finne en numerisk løsning, må vi gjette på hva konsentrasjonene kan være. Disse gjetningene bruker SymPy for å finne en bedre løsning. Her prøver vi oss med at:
# - lite $\text{H}_2 \text{C}_2 \text{O}_4$ dissosierer, så denne er kanskje ca. lik startkonsentrasjonen på 0.1 M
# - noe $\text{HC}_2 \text{O}_4^{-}$, $\text{C}_2 \text{O}_4^{2-}$ og $\text{H}^{+}$ dannes. La oss bare si at de er ca. 1/10 av startkonsentrasjonen (0.01 M).
# - det vil være lite $\text{OH}^-$ ved likevekt siden vi ser på en syre. For å være konsistent med konsentrasjonen vi satte over for $\text{H}^+$, la oss sette den til $10^{-12}$.
løsning = sym.nsolve(
[ligning1, ligning2, ligning3, ligning4, ligning5],
[c_HHA, c_HA, c_A, c_H, c_OH],
[0.1, 0.01, 0.01, 0.01, 1e-12],
)
løsning
# OK, her ser vi at de løsningene jeg gjettet på ikke var så veldig gode. Spesielt bommet jeg veldig på $[\text{C}_2 \text{O}_4^{2-}]$ (hvis jeg hadde vært litt mer observant, så hadde jeg sett at $K_{a,2}$ er liten, slik at det sannsynligvis er lite $\text{C}_2 \text{O}_4^{2-}$ ved likevekt). Men SymPy fant likevel en løsning!
#
# Når vi løser ligninger numerisk på denne måten, kan svaret avhenge av hva vi gjetter på at konsentrasjonene ca. er. SymPy bruker disse for å finne bedre løsninger, men hvis vi er uheldige med gjetningen, så kan vi ende opp med f.eks. negative konsentrasjoner. Her gikk det heldigvis bra. La oss oppsummere løsningen:
print(f'[H2C2O4]: {løsning[0]:.3g} M')
print(f'[HC2O4^-]: {løsning[1]:.3g} M')
print(f'[H^+]: {løsning[3]:.3g} M')
print(f'[C2O4^2-]: {løsning[2]:.3e} M')
print(f'[OH^-]: {løsning[4]:.3g} M')
# Til sammenligning sier læreboken:
# - $[\text{H}_2 \text{C}_2 \text{O}_4] = 0.046$ M
# - $[\text{HC}_2 \text{O}_4^{-}] = 0.054$ M
# - $[\text{H}^{+}] = 0.054$ M
# - $[\text{C}_2 \text{O}_4^{2-}] = 6.1 \times 10^{-5}$ M
# - $[\text{OH}^{-}] = 1.9 \times 10^{-13}$ M
#
# Vi fant altså samme løsning!
# Her har vi ikke fått noe informasjon om at SymPy faktisk konvergerte. Vi bør derfor i det minste sjekke at alle ligningene vi definerte over er oppfylt for løsningen vi har funnet.
#
# Vi kan gjøre dette ved å sette inn verdier og sjekke at venstre side av ligningene er ca. lik høyre side av ligningene. Vi kan bruke `ligning.rhs` for å få tilgang til høyre side ("right-hand-side") og `ligning.lhs` for å få tilgang til venstre side ("left-hand-side").
#
# La oss trekke venstre fra høyre side for alle ligningene og sjekke at forskjellen blir ca. 0:
test1 = ligning1.lhs.evalf(subs={c_HHA: løsning[0], c_HA: løsning[1], c_H: løsning[3]}) - ligning1.rhs
test2 = ligning2.lhs.evalf(subs={c_HA: løsning[1], c_A: løsning[2], c_H: løsning[3]}) - ligning2.rhs
test3 = ligning3.lhs.evalf(subs={c_H: løsning[3], c_OH: løsning[4]}) - ligning3.rhs
test4 = (ligning4.lhs.evalf(subs={c_HA: løsning[1], c_A: løsning[2], c_OH: løsning[4]}) -
ligning4.rhs.evalf(subs={c_H: løsning[3]}))
test5 = ligning5.lhs - ligning5.rhs.evalf(subs={c_HHA: løsning[0], c_HA: løsning[1],c_A: løsning[2]})
sum_feil = 0.0
for i, test in enumerate((test1, test2, test3, test4, test5)):
print(f'lhs - rhs, ligning{i+1}: {test}')
sum_feil += abs(test)
print(f'Summert feil: {sum_feil}')
# Alle disse er forskjellene er små (største er $6.9 \times 10^{-18}$ på min datamaskin) og vi er derfor fornøyde med den numeriske løsningen!
|
jupyter/syrebase/diprotisk.ipynb
|
# Solution to: [Day 3: Drawing Marbles](https://www.hackerrank.com/challenges/s10-mcq-6/problem)
# <h1 id="tocheading">Table of Contents</h1>
# <div id="toc"></div>
#
#
# - Table of Contents
# - Math Solution
# - Facts
# - Monte Carlo Solution
# - Imports
# - Constants
# - Auxiliary functions
# - Main
# + language="javascript"
# $.getScript('https://kmahelona.github.io/ipython_notebook_goodies/ipython_notebook_toc.js')
# -
# This script contains 2 sections:
# 1. Math solution to the problem
# 2. Monte Carlo simulation of the problem
# # Math Solution
# A bag contains 3 red marbles and 4 blue marbles. Then, 2 marbles are drawn from the bag, at random, without replacement.
# If the first marble drawn is red, what is the probability that the second marble is blue?
#
#
# ## Facts
# - 7 marbles in the bag
# - 1st one is always red
# - P(B)?
#
# If the first one is always red, we don't calculate the probability of the 1st draw.
# Thus, we use the 6 remaining marbles.
# \begin{equation}
# \large
# P(B) = \frac{4}{6} = \frac{2}{3}
# \end{equation}
# # Monte Carlo Solution
# ## Imports
from typing import List
import random
# ## Constants
MARBLE_DICT = {
'r' : 3,
'b' : 4
}
FIRST_MARBLE = 'r'
SECOND_MARBLE = 'b'
# ## Auxiliary functions
def create_marble_bag(marbles: dict) -> List[str]:
"""Returns list of marbles to draw from."""
bag = []
for k, v in marbles.items():
m = [k for _ in range(v)]
bag += m
return bag
def remove_first_marble(bag: List[str], marble: str) -> List[str]:
"""Returns bag after removing marble."""
bag.remove(marble)
return bag
def check_second_marble(bag: List[str], marble: str) -> bool:
"""Returns boolean if sample from bag is the marble."""
return random.choice(bag) == marble
def get_ratio(bag: List[str], marble: str, iterations: int) -> float:
"""Returns ratio of times sample from bag is marble."""
was_marble = 0
for _ in range(iterations):
if check_second_marble(bag, marble):
was_marble += 1
return was_marble / iterations
# ## Main
def main():
bag = create_marble_bag(MARBLE_DICT)
bag = remove_first_marble(bag, FIRST_MARBLE)
iterations = 1000000
ratio = get_ratio(bag, SECOND_MARBLE, iterations)
print(ratio)
if __name__ == "__main__":
main()
|
statistics/10_days/11_day3drawingmarbles.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# ## Business Understanding
# Purpose: Ask relevant questions and define objectives for the problem that needs to be tackled
#
# ## Background
# In recent years, the range of funding options for projects created by individuals and small companies has expanded considerably. In addition to savings, bank loans, friends & family funding and other traditional options, crowdfunding has become a popular and readily available alternative.
#
# Kickstarter, founded in 2009, is one particularly well-known and popular crowdfunding platform. It has an all-or-nothing funding model, whereby a project is only funded if it meets its goal amount; otherwise no money is given by backers to a project. A huge variety of factors contribute to the success or failure of a project — in general, and also on Kickstarter. Some of these are able to be quantified or categorized, which allows for the construction of a model to attempt to predict whether a project will succeed or not. The aim of this project is to construct such a model and also to analyse Kickstarter project data more generally, in order to help potential project creators assess whether or not Kickstarter is a good funding option for them, and what their chances of success are.
#
# ### Final Deliverables
#
#
# * Well designed presentation for non-technical stakeholders outlining findings and recommendations, as well as future work (10min presentation).
# * Jupyter notebook following Data Science Lifecycle
#
# ### Things to think about
#
# * Try different (at least 3) machine learning algorithms to check which performs best on the problem at hand
# * What would be right performance metric: Precision, recall, accuracy, F1 score, or something else? (Check TPR?)
# * Check for data imbalance
#
#
# ## Key Question
#
# We currently hold a task by Kickstarter to come up with a model to predict in a first step whether is project is likely to be successful, given certain project parameters. In a second step (out of scope), Kickstarter would like to be able to provide a good goal recommendation for creators( for example using staff picks etc.)
#
# * Given certain project parameters, __is a campaign likely to succeed or fail?__ --> classification
# * what would e a __reasonable goal reccomendation for creators__ --> regression
#
#
#
# ## Feature Glossary
#
# Features included in model
#
# * Target : state
# *
# *
# *
#
# ## Dataset Description
#
# - **backers_count**: Amount of people who backed this project
# - **category**:
# - **country**: Country the project owner lives in
# - **created_at**: Date when the prjoect was created
# - **currency**: Currency of the country where the owner lives in
# - **currency_trailing_code**:
# - **current_currency**:
# - **deadline**: Date until the project can be backed
# - **disable_communication**: If the communication with owner was disabled or not
# - **fx_rate**: Foreign exchange rate
# - **goal**: Project is only funded when the goal amount is reached
# - **launched_at**: Date when the project was launced
# - **spotlight**: Highlighted projects (available to all projects that are successfully funded)
# - **staff_pick**: Promissing project picked by Kickstarter employees
# - **state**: Project status
# - **state_changed_at**: Date when state changed the last time
# - **static_usd_rate**: static USD Convergen rate at time
# - **usd_pledged**: pledge amount converted to USD using Static_usd_rate
#
#
# ## Dataset New/Added Feature Description
#
# - **campaign_days**: Days the Project was live
# - **pledged_over**: Amount Pledged surpassing the Goal(with converted pledge amount)
# - **pre_launched_days**: Days before the Project was launched
#
#
# ## Target Metric
#
# * F1 score — Since creators wouldn’t want the model to predict too many success that will turn out to be a failure (minimize False Positives) and backers would want to make sure the model capture as many success as possible (minimize False Negatives), I want a balance between precision and recall
#
# ## Outcome / Reccomendations
# *
# *
# *
# + [markdown] tags=[]
# ## Import Libraries
# + tags=[]
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.ticker as ticker
from matplotlib.gridspec import GridSpec
import scipy as sc
from scipy.stats import kstest
import seaborn as sns
import math
import warnings
warnings.filterwarnings("ignore")
#Data mining
import os, glob
#Preprocessing
from sklearn.model_selection import train_test_split, RandomizedSearchCV
from sklearn.dummy import DummyClassifier
from sklearn.preprocessing import OneHotEncoder, MinMaxScaler, StandardScaler, PolynomialFeatures, LabelEncoder
from sklearn.linear_model import LinearRegression, LogisticRegression
from sklearn.model_selection import cross_val_score
from sklearn.metrics import classification_report, confusion_matrix, f1_score, accuracy_score
import imblearn
from imblearn.over_sampling import RandomOverSampler
## AdaBoost
from sklearn.ensemble import AdaBoostClassifier
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import make_scorer
from sklearn.tree import DecisionTreeClassifier
# + [markdown] tags=[]
# ## Dashboard
# Purpose : Define global variables and visuals
# -
random_state = 100
test_size = 0.3
sns.set(style = "white")
# + [markdown] tags=[]
# ## Data Mining
# +
# Import multiple Kickstarter csv files and merge into one dataframe
path = "data-2"
all_files = glob.glob(os.path.join(path, "*.csv"))
all_df = []
for f in all_files:
df = pd.read_csv(f, sep=',')
df['file'] = f.split('/')[-1]
all_df.append(df)
merged_df = pd.concat(all_df, ignore_index=True, sort=True)
# +
#
# merged_df = pd.read_csv('data-2/Kickstarter_all.csv') ### brauche ich wenn ich den Anderen Kram nicht laufen lassen will
# + [markdown] tags=[]
# ## Inspection and Data Cleaning
# -
merged_df.info()
# +
#save the merged data as .zip
#compression_opts = dict(method='zip', archive_name='out.csv')
#merged_df.to_csv('out.zip', index=False, compression=compression_opts)
# -
# Display shape of "data"
merged_df.shape
merged_df.head()
merged_df.columns
merged_df.groupby('state').count()
pd.isnull(merged_df).sum()
# + [markdown] tags=[]
# ## Data Handling
# -
# create a dataset for Inspection
final = merged_df.copy()
# + [markdown] tags=[]
# ### Dropping Data
# -
drop_list = []
# #### Dropping features with missing values
drop_missing_values = ['blurb', 'friends', 'is_backing', 'is_starred', 'permissions', 'usd_type', 'location']
drop_list.extend(drop_missing_values)
final = final.drop(drop_missing_values, axis = 1)
# #### Dropping useless features
drop_useless_features = ['creator', 'currency_symbol', 'name', 'photo', 'profile', 'slug', 'source_url', 'urls', 'file']
drop_list.extend(drop_useless_features)
final = final.drop(drop_useless_features, axis = 1)
# #### Dropping redundant features
drop_redundant_features = ['pledged', 'usd_pledged']
drop_list.extend(drop_redundant_features)
final = final.drop(drop_redundant_features, axis = 1)
drop_list
# #### Replacing features
def clean_category(DataFrame):
cat_list = []
subcat_list = []
for e in DataFrame.category:
string_list = e.split(',')
if '/' in string_list[2]:
cat_list.append(string_list[2].split('/')[0][8:])
subcat_list.append(string_list[2].split('/')[1][:-1])
else:
cat_list.append(string_list[2][8:-1])
subcat_list.append('None')
DataFrame['category'] = cat_list
DataFrame['sub_category'] = subcat_list
return DataFrame
modified_list = ['category','state']
final = clean_category(final)
final.category.unique()
# +
#replace successful and failed with 1 and 0
final.state.replace(['successful','failed'], [1,0],inplace=True)
final = final.query('state == [1,0]')
final.state = final.state.astype(int)
#
final.is_starrable = final.is_starrable.astype(int)
final.disable_communication = final.disable_communication.astype(int)
final.currency_trailing_code = final.currency_trailing_code.astype(int)
final.staff_pick = final.staff_pick.astype(int)
final.spotlight = final.spotlight.astype(int)
#drop live,susspended,cancelled
#final = final[final['state'] == [1,0]]
# + [markdown] tags=[]
# ### Time conversions
#
#
# -
modified_list.extend(['launched_at', 'deadline', 'created_at', 'state_changed_at'])
#converting unix time
final.launched_at = pd.to_datetime(final.launched_at,unit='s',infer_datetime_format=True)
final.deadline = pd.to_datetime(final.deadline,unit='s',infer_datetime_format=True)
final.created_at = pd.to_datetime(final.created_at,unit='s',infer_datetime_format=True)
final.state_changed_at = pd.to_datetime(final.state_changed_at,unit='s',infer_datetime_format=True)
# ### Writing df changes
# +
feature_list = list(merged_df.columns)
df_features = pd.DataFrame(feature_list,columns =['features'])
df_features['dropped'] = df_features.features.isin(drop_list)
df_features['drop_reason'] = ['missing_values' if x in drop_missing_values \
else 'useless' if x in drop_useless_features \
else 'redundant' if x in drop_redundant_features \
else 'None' for x in df_features['features']]
df_features['modified'] = df_features.features.isin(modified_list)
# -
df_features
# + [markdown] tags=[]
# # Data Exploration
# Purpose: we gotta form a hypotheses / story about our defined problem by visually analyzing the data
# -
#new dataset for exploration
data_exp = final.copy()
# +
#years
#final['launched_at_yr'] = [date.year for date in final['launched_at']]
# -
final.info()
# Seperate continious vs. categorical variables
data_cat_col = ['category','country','sub_category','country','currency','current_currency','is_starrable','disable_communication','state']
data_cont_col = [x for x in final if x not in data_cat_col]
data_cat = final[data_cat_col]
data_cont = final[data_cont_col]
# Check if scaling is needed ( we can do this by looking at the .skew()
final.skew()
# +
#Plot correlation heatmap for continious values
mask = np.triu(np.ones_like(data_cont.corr(), dtype=np.bool))
f, ax = plt.subplots(figsize=(11, 9))
cmap = sns.diverging_palette(220, 10, as_cmap=True)
sns.heatmap(data_cont.corr(), mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5}, annot=True, fmt=".1g");
# -
#Plot a histogram of our Target 'state' and see if it needs scaling for future work
data_exp['state'].value_counts(ascending=True).plot(kind='bar')
# * imbalanced data!!
# +
#plt.figure(figsize=(14,8))
#sns.countplot(x='launched_at_yr', hue='state', data=data_exp);
# -
plt.figure(figsize=(14,10))
sns.countplot(x='category', hue='state', data=data_exp);
# # Feature Engineering
new_features_list = ['pledged_over', 'campaign_days', 'pre_launched_days', 'launched_at_yr', 'goal_converted']
# +
#create new features
final['pledged_over'] = final.converted_pledged_amount - final.goal
final['campaign_days'] = ((final.deadline - final.launched_at)/ np.timedelta64(1, 'h')).astype(int)
final['pre_launched_days'] = ((final.launched_at - final.created_at)/ np.timedelta64(1, 'h')).astype(int)
final['launched_at_yr'] = [date.year for date in final['launched_at']]
final['launched_at_mth'] = [date.month for date in final['launched_at']]
final['goal_converted'] = final["goal"] * final["static_usd_rate"]
#use log on goal_converted
# -
#use log on stuff
final['goal_converted_log'] = [(math.log(el)+1) for el in final['goal_converted']]
final['converted_pledged_amount_log'] = np.log(final['converted_pledged_amount']+1)
final['backers_count_log'] = np.log(final['backers_count']+1)
final['goal_converted_log'].unique()
# # Preprocessing (Train/Test Split and Basemodel)
# In order to apply modelling on different dataset types, we should consider a nice way to do the splits.
#
#
# +
#define predictors and target variable X,y
#X = final.drop(["state"], axis=1)
#y = final["state"]
# -
final.info()
# +
# Get dummies for object variables: category, sub_category, currency, country
category_dummies = pd.get_dummies(final['category'], prefix='cat', drop_first=True)
subcategory_dummies = pd.get_dummies(final['sub_category'], prefix='subcat', drop_first=True)
currency_dummies = pd.get_dummies(final['currency'], prefix='cur', drop_first=True)
country_dummies = pd.get_dummies(final['country'], prefix='country', drop_first=True)
final = final.drop(['category', 'sub_category', 'currency', 'country'], axis=1)
final = pd.concat([final, category_dummies, subcategory_dummies, currency_dummies, country_dummies], axis=1)
# -
X = final.drop(["state", 'goal_converted', 'launched_at_yr', 'pledged_over', 'spotlight', 'currency_trailing_code', 'current_currency', 'created_at', 'deadline', 'fx_rate', 'goal', 'id', 'launched_at', 'state_changed_at', 'backers_count', 'backers_count_log', 'static_usd_rate', 'converted_pledged_amount_log', 'converted_pledged_amount'], axis=1)
y = final["state"]
#Split data into training and testing sets
X_train, X_test, y_train, y_test= train_test_split(X,y,test_size=test_size,
random_state=random_state,
shuffle=True,
stratify=y)
# X_train = np.array(X_train)
# X_test = np.array(X_test)
# y_train = np.array(y_train)
# y_test = np.array(y_test)
# +
# create a dummy classifier model as Basemodel
dum_clf = DummyClassifier(strategy='constant',constant=1).fit(X_train,y_train)
y_pred_dum_clf = dum_clf.predict(X_test)
print(confusion_matrix(y_test,y_pred_dum_clf))
print(classification_report(y_test,y_pred_dum_clf))
# +
#for future work
#scores = cross_val_score(dummy_clf, X_train, y_train, scoring='f1', cv=10, n_jobs=-1)
# +
#use oversampling
# define oversampling strategy
oversample = RandomOverSampler(sampling_strategy='minority', random_state=random_state)
# fit and apply the transform
X_train_over, y_train_over = oversample.fit_resample(np.array(X_train), np.array(y_train))
# -
sum(y_train_over)
len(y_train_over)
features_scalable_list = ['goal_converted']
data_cont_col
# +
#X_train[features_scalable_list]
# -
X_train_scaled = X_train.copy()
X_test_scaled = X_test.copy()
# +
#use standard scaler on X_train and y_train
scaler = StandardScaler()
X_train_scaled[features_scalable_list] = scaler.fit_transform(np.array(X_train[features_scalable_list])) # Scaler is fitted to training data _only_
X_test_scaled[features_scalable_list] = scaler.transform(np.array(X_test[features_scalable_list])) # Already fitted scaler is applied to test data
#data_cat_col = ['category','country','sub_category','country','currency','current_currency','is_starrable','disable_communication']
#data_cont_col = [x for x in final if x not in data_cat_col]
# +
#use standard scaler on X_train_over and y_train_over
# + [markdown] tags=[]
# # Future Work
# +
# To do: save final df as csv
#compression_opts = dict(method='zip', archive_name='Kickstarter_all_clean.csv')
#final.to_csv('Kickstarter_all_clean.zip', index=False, compression=compression_opts)
# -
# # Predictive Modelling
# Purpose: Train machine learning models (supervised), evaluate their performance and use them to make predictions
# * using f1 score as our metric
# +
#logistic regression
# +
#Random Forest Classifier
# +
#Support Vector Machines (use classifier)
# -
#AdaBoost
X_train.head()
adaB = AdaBoostClassifier(base_estimator=DecisionTreeClassifier(), n_estimators=100, learning_rate=0.4, random_state=1)
model = adaB.fit(X_train_scaled, y_train)
X_train.info()
def generic_clf(X_train, Y_train, X_test, Y_test, clf):
mod = clf.fit(X_train,Y_train)
pred_train = clf.predict(X_train)
pred_test = clf.predict(X_test)
print(f1_score(Y_test, pred_test))
print(classification_report(Y_test, pred_test))
print(confusion_matrix(Y_test, pred_test))
sns.heatmap(confusion_matrix(Y_test, pred_test), annot=True, cmap='YlGn', fmt='d');
return pred_train, pred_test
# + jupyter={"source_hidden": true} tags=[]
# the following 2 modells use the features
first_features_list = ['disable_communication',
'is_starrable',
'staff_pick',
'campaign_days',
'pre_launched_days',
'launched_at_yr',
'launched_at_mth',
'goal_converted',
'goal_converted_log',
'cat_comics',
'cat_crafts',
'cat_dance',
'cat_design',
'cat_fashion',
'cat_film & video',
'cat_food',
'cat_games',
'cat_journalism',
'cat_music',
'cat_photography',
'cat_publishing',
'cat_technology',
'cat_theater',
'subcat_None',
'subcat_academic',
'subcat_accessories',
'subcat_action',
'subcat_animals',
'subcat_animation',
'subcat_anthologies',
'subcat_apparel',
'subcat_apps',
'subcat_architecture',
'subcat_art books',
'subcat_audio',
'subcat_bacon',
'subcat_blues',
'subcat_calendars',
'subcat_camera equipment',
'subcat_candles',
'subcat_ceramics',
"subcat_children's books",
'subcat_childrenswear',
'subcat_chiptune',
'subcat_civic design',
'subcat_classical music',
'subcat_comedy',
'subcat_comic books',
'subcat_community gardens',
'subcat_conceptual art',
'subcat_cookbooks',
'subcat_country & folk',
'subcat_couture',
'subcat_crochet',
'subcat_digital art',
'subcat_diy',
'subcat_diy electronics',
'subcat_documentary',
'subcat_drama',
'subcat_drinks',
'subcat_electronic music',
'subcat_embroidery',
'subcat_events',
'subcat_experimental',
'subcat_fabrication tools',
'subcat_faith',
'subcat_family',
'subcat_fantasy',
"subcat_farmer's markets",
'subcat_farms',
'subcat_festivals',
'subcat_fiction',
'subcat_fine art',
'subcat_flight',
'subcat_food trucks',
'subcat_footwear',
'subcat_gadgets',
'subcat_gaming hardware',
'subcat_glass',
'subcat_graphic design',
'subcat_graphic novels',
'subcat_hardware',
'subcat_hip-hop',
'subcat_horror',
'subcat_illustration',
'subcat_immersive',
'subcat_indie rock',
'subcat_installations',
'subcat_interactive design',
'subcat_jazz',
'subcat_jewelry',
'subcat_kids',
'subcat_knitting',
'subcat_latin',
'subcat_letterpress',
'subcat_literary journals',
'subcat_literary spaces',
'subcat_live games',
'subcat_makerspaces',
'subcat_metal',
'subcat_mixed media',
'subcat_mobile games',
'subcat_movie theaters',
'subcat_music videos',
'subcat_musical',
'subcat_narrative film',
'subcat_nature',
'subcat_nonfiction',
'subcat_painting',
'subcat_people',
'subcat_performance art',
'subcat_performances',
'subcat_periodicals',
'subcat_pet fashion',
'subcat_photo',
'subcat_photobooks',
'subcat_places',
'subcat_playing cards',
'subcat_plays',
'subcat_poetry',
'subcat_pop',
'subcat_pottery',
'subcat_print',
'subcat_printing',
'subcat_product design',
'subcat_public art',
'subcat_punk',
'subcat_puzzles',
'subcat_quilts',
'subcat_r&b',
'subcat_radio & podcasts',
'subcat_ready-to-wear',
'subcat_residencies',
'subcat_restaurants',
'subcat_robots',
'subcat_rock',
'subcat_romance',
'subcat_science fiction',
'subcat_sculpture',
'subcat_shorts',
'subcat_small batch',
'subcat_software',
'subcat_sound',
'subcat_space exploration',
'subcat_spaces',
'subcat_stationery',
'subcat_tabletop games',
'subcat_taxidermy',
'subcat_television',
'subcat_textiles',
'subcat_thrillers',
'subcat_translations',
'subcat_typography',
'subcat_vegan',
'subcat_video',
'subcat_video art',
'subcat_video games',
'subcat_wearables',
'subcat_weaving',
'subcat_web',
'subcat_webcomics',
'subcat_webseries',
'subcat_woodworking',
'subcat_workshops',
'subcat_world music',
'subcat_young adult',
'subcat_zines',
'cur_CAD',
'cur_CHF',
'cur_DKK',
'cur_EUR',
'cur_GBP',
'cur_HKD',
'cur_JPY',
'cur_MXN',
'cur_NOK',
'cur_NZD',
'cur_SEK',
'cur_SGD',
'cur_USD',
'country_AU',
'country_BE',
'country_CA',
'country_CH',
'country_DE',
'country_DK',
'country_ES',
'country_FR',
'country_GB',
'country_HK',
'country_IE',
'country_IT',
'country_JP',
'country_LU',
'country_MX',
'country_NL',
'country_NO',
'country_NZ',
'country_SE',
'country_SG',
'country_US']
# + tags=[]
adamodel = generic_clf(X_train, y_train, X_test, y_test, adaB)
# -
adamodel_scaled = generic_clf(X_train_scaled, y_train, X_test_scaled, y_test, adaB)
print(model.feature_importances_)
len(model.feature_importances_)
important_features = X_train.columns.to_list()
print(important_features[3], important_features[5], important_features[8], important_features[9])
# +
adaB_opt = AdaBoostClassifier(base_estimator=DecisionTreeClassifier(max_depth= 3, min_samples_split = 4), \
n_estimators=120, learning_rate=0.5, random_state=1,)
model_opt = adaB_opt.fit(X_train, y_train)
adamodel_opt = generic_clf(X_train, y_train, X_test, y_test, adaB_opt)
# -
features_used = X_train.columns.tolist()
my_project = pd.read_csv('Future_projects_first.csv', sep = ';')
my_project.head()
predict_me = my_project.iloc[0]
predict_me
my_project.values.reshape(1, -1)
type(predict_me)
np.asarray(predict_me)
adaB_opt.predict(predict_me)
future_projects = pd.DataFrame(features_used)
future_projects
future_projects_tr = future_projects.transpose(copy=False)
to_predict = X_train.copy()
to_predict = to_predict.drop(to_predict.index.to_list()[1:] ,axis = 0 )
to_predict.to_csv('to_predict_df.csv')
# +
#to_predict_new = pd.read_csv('to_predict_df_new.csv', sep = ';')
# -
adaB_opt.predict(np.array(to_predict_new))
np.array(to_predict_new)
to_predict_new.head()
#compression_opts = dict(method='zip', archive_name='Kickstarter_all_clean.csv')
future_projects_tr.to_csv('data-2/Future_projects.csv', index=False)
important_features_opt = model_opt.feature_importances_.tolist()
feature_names_opt = X_train.columns.to_list()
feature_df = pd.DataFrame(important_features_opt, feature_names_opt)
feature_ranking = feature_df.sort_values(0, ascending=False)
feature_ranking.head(25)
# +
# GridSearch
parameters = {'n_estimators':[50, 120],
'learning_rate':[0.1, 0.5, 1.],
'base_estimator__min_samples_split' : np.arange(2, 8, 2),
'base_estimator__max_depth' : np.arange(1, 4, 1)
}
clf = AdaBoostClassifier(base_estimator=DecisionTreeClassifier())
scorer = make_scorer(f1_score)
grid = GridSearchCV(clf, parameters, verbose=True, scoring = scorer)
result = grid.fit(X_train, y_train)
print('Best Parameters:', result.best_params_)
print('Best Score:', result.best_score_)
# -
# Best Parameters: {'base_estimator__max_depth': 3, 'base_estimator__min_samples_split': 2, 'learning_rate': 0.5, 'n_estimators': 120}
#
# Best Score: 0.8554056873636828
# + tags=[]
# Further GridSearch
parameters2 = {'n_estimators':[120, 150],
'learning_rate':[0.4, 0.5],
'base_estimator__min_samples_split' : np.arange(2, 8, 2),
'base_estimator__max_depth' : np.arange(1, 4, 1)
}
clf = AdaBoostClassifier(base_estimator=DecisionTreeClassifier())
scorer = make_scorer(f1_score)
grid2 = GridSearchCV(clf, parameters2, verbose=True, scoring = scorer)
result2 = grid.fit(X_train, y_train)
print('Best Parameters:', result2.best_params_)
print('Best Score:', result2.best_score_)
# + jupyter={"outputs_hidden": true, "source_hidden": true} tags=[]
X_train.columns.to_list()
# + [markdown] tags=[]
# #### TO DO:
# - check ob monat drin is,
# - sub cat raus, year raus
# - eine goal var raus
# + [markdown] tags=[]
# # Ensemble Methods
# +
#use KNN,SVC,DTC,Randomforestclassifier,XGB....
# -
# # Future Work
# +
#use maybe RandomizedSearchCV on RandomForest or any given Algorithm
# -
# # Data Visualisation
# Purpose: Communicate the findings with stakeholders using plots and interactive visualisations
# ideas for stakeholder communication:
# - which are the top (5) categories for successfull kickstarter projects?
# - goal amount as important feature: bin the amounts and plot them over successfull projects count.
# - add month variable to show which month for launching the project is the most promising.
# - show successrates of projects over the years: Kickstarter, still worth it?
#
# - outlook for future research: what is the role of marketing campaigns? (further data needed)
# # Findings
# Purpose: Summarize the key results and findings
|
03_Model.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
from __future__ import division, print_function
import numpy as np
from collections import OrderedDict
import logging
from IPython.display import display
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
from astropy.io import fits
import astropy.wcs
from astropy import coordinates
import astropy.units as apu
from astropy import table
import astropyp
from astropyp.wrappers.astromatic import ldac
from astropyp.phot import stack
import bd_search
alogger = logging.getLogger('astropyp')
alogger.setLevel(logging.INFO)
idx_connect = 'sqlite:////media/data-beta/users/fmooleka/decam/decam.db'
ref_path = '/media/data-beta/users/fmooleka/decam/catalogs/ref/'
# SExtractor 'extract' detection parameters
conv_filter = np.load('/media/data-beta/users/fmooleka/2016decam/5x5gauss.npy')
sex_params = {
'extract': {
'thresh': 40,
#'err':,
'minarea': 3, # default
'conv': conv_filter,
#'deblend_nthresh': 32, #default
'deblend_cont': 0.001,
#'clean': True, #default
#'clean_param': 1 #default
},
'kron_k': 2.5,
'kron_min_radius': 3.5,
'filter': conv_filter,
#'thresh': 1.5 # *bkg.globalrms
}
obj='F100'
refname = '2MASS'
#refname = 'UCAC4'
fullref = ldac.get_table_from_ldac(ref_path+'{0}-{1}.fits'.format(obj, refname))
# +
import warnings
from astropy.utils.exceptions import AstropyWarning
warnings.simplefilter('ignore', category=AstropyWarning)
def get_exp_files(expnum, night, filtr, idx_connect):
sql = 'select * from decam_obs where expnum={0} and filter like "{1}%" and dtcaldat="{2}"'.format(
expnum, filtr, night)
exp_info = astropyp.db_utils.index.query(sql, idx_connect)
img_filename = exp_info[exp_info['PRODTYPE']=='image'][0]['filename']
img = fits.open(img_filename)
dqmask_filename = exp_info[exp_info['PRODTYPE']=='dqmask'][0]['filename']
dqmask = fits.open(dqmask_filename)
return img, dqmask
min_flux = 1000
min_amplitude = 1000
good_amplitude = 50
calibrate_amplitude = 200
frame = 1
explist = [442433, 442434, 442435]
aper_radius = 8
ccds = []
for expnum in explist:
#img, dqmask = get_exp_files(expnum, "2015-05-26", "i", idx_connect)
img, dqmask = get_exp_files(expnum, "2015-05-26", "z", idx_connect)
header = img[frame].header
wcs = astropy.wcs.WCS(header)
img_data = img[frame].data
dqmask_data = dqmask[frame].data
ccd = astropyp.phot.phot.SingleImage(header, img_data, dqmask_data,
wcs=wcs, gain=4., exptime=30, aper_radius=aper_radius)
ccds.append(ccd)
# -
ccd_stack = stack.Stack(ccds, 1)
ccd_stack.detect_sources(min_flux=min_flux, good_amplitude=good_amplitude,
calibrate_amplitude=calibrate_amplitude, psf_amplitude=1000, sex_params=sex_params,
subtract_bkg=True, windowed=False)
ccd_stack.get_transforms()
reload(astropyp.phot.stack)
ccd_stack.stack_images(pool_size=1, slices=[slice(0,500),slice(0,500)])
print(ccd_stack.wcs)
print(ccd_stack.ccds[1].wcs)
ccd_stack.detect_sources(ccd_stack.stack, min_flux=100,
good_amplitude=100, calibrate_amplitude=200, psf_amplitude=1000,
sex_params=sex_params, subtract_bkg=True, windowed=False, wcs=ccd_stack.wcs,
aper_radius=8, exptime=30.)
cat1 = ccd_stack.stack.catalog
cat2 = ccd_stack.ccds[1].catalog
cat2 = cat2.sources[(cat2.x<500) & (cat2.y<500)]
print(len(ccd_stack.stack.catalog.sources))
print(len(ccd_stack.ccds[1].catalog.sources))
import astropy.units as apu
((cat1['ra'][:4]-cat2['ra'][:4])*apu.deg).to('arcsec')
ccd_stack.wcs.to_header()
reload(astropyp.catalog)
astropyp.catalog.match_all_catalogs([cat1,cat2], 'ra','dec', combine=False, separation=2*apu.arcsec)
ccd_stack.ccds[0].wcs
imgs = [ccd.img for ccd in ccd_stack.ccds]
dqmasks = [ccd.dqmask for ccd in ccd_stack.ccds]
tx_solutions = [ccd_stack.tx_solutions[(1,0)], None, ccd_stack.tx_solutions[(1,2)]]
# %time stack, dqmask, patches = stack_full_images(imgs, 1, tx_solutions, dqmasks)
imgs = [ccd.img[:1000,:500] for ccd in ccd_stack.ccds]
dqmasks = [ccd.dqmask[:1000,:500] for ccd in ccd_stack.ccds]
tx_solutions = [ccd_stack.tx_solutions[(1,0)], None, ccd_stack.tx_solutions[(1,2)]]
# %time stack, dqmask, patches = stack_full_images(imgs, 1, tx_solutions, dqmasks)
# +
imgs = [ccd.img for ccd in ccd_stack.ccds]
dqmasks = [ccd.dqmask for ccd in ccd_stack.ccds]
tx_solutions = [ccd_stack.tx_solutions[(1,0)], None, ccd_stack.tx_solutions[(1,2)]]
# %time stack, dqmask, patches=stack_full_images(imgs, 1, tx_solutions, dqmasks, order=5)
# +
imgs = [ccd.img for ccd in ccd_stack.ccds]
dqmasks = [ccd.dqmask for ccd in ccd_stack.ccds]
tx_solutions = [ccd_stack.tx_solutions[(1,0)], None, ccd_stack.tx_solutions[(1,2)]]
# %time stack, dqmask, patches=stack_full_images(imgs, 1, tx_solutions, dqmasks, order=3)
# -
stk = stack.filled(0)
max_offset=3
ccd = astropyp.phot.phot.SingleImage(
img=stk, dqmask=dqmask, gain=4., exptime=30, aper_radius=8)
ccd.detect_sources(sex_params, subtract_bkg=True)
ccd.select_psf_sources(min_flux, min_amplitude, edge_dist=aper_radius+max_offset)
psf_array = ccd.create_psf()
ccd.show_psf()
good_idx = ccd.catalog.sources['peak']>calibrate_amplitude
#good_idx = ccd.catalog.sources['peak']>good_amplitude
good_idx = good_idx & (ccd.catalog.sources['pipeline_flags']==0)
result = ccd.perform_psf_photometry(indices=good_idx)
# +
good_idx = ccd.catalog.sources['peak']>calibrate_amplitude
good_idx = good_idx & (ccd.catalog.sources['pipeline_flags']==0)
good_idx = good_idx & np.isfinite(ccd.catalog.sources['psf_mag'])
good_sources = ccd.catalog.sources[good_idx]
print('rms', np.sqrt(np.sum(good_sources['psf_mag_err']**2/len(good_sources))))
print('mean', np.mean(good_sources['psf_mag_err']))
print('median', np.median(good_sources['psf_mag_err']))
print('stddev', np.std(good_sources['psf_mag_err']))
bad_count = np.sum(good_sources['psf_mag_err']>.05)
print('bad psf error: {0}, or {1}%'.format(bad_count, bad_count/len(good_sources)*100))
print('Better than 5%: {0} of {1}'.format(np.sum(good_sources['psf_mag_err']<=.05), len(good_sources)))
print('Better than 2%: {0} of {1}'.format(np.sum(good_sources['psf_mag_err']<=.02), len(good_sources)))
good_sources['aper_flux','psf_flux','peak','psf_mag_err'][good_sources['psf_mag_err']>.05]
# -
max_offset=3
ccd = astropyp.phot.phot.SingleImage(
img=stk, dqmask=dqmask, gain=4., exptime=30, aper_radius=8)
ccd.detect_sources(sex_params, subtract_bkg=True)
ccd.select_psf_sources(min_flux, min_amplitude, edge_dist=aper_radius+max_offset)
psf_array = ccd.create_psf()
ccd.show_psf()
#good_idx = ccd.catalog.sources['peak']>calibrate_amplitude
good_idx = ccd.catalog.sources['peak']>good_amplitude
good_idx = good_idx & (ccd.catalog.sources['pipeline_flags']==0)
result = ccd.perform_psf_photometry(indices=good_idx)
# +
#good_idx = ccd.catalog.sources['peak']>calibrate_amplitude
good_idx = ccd.catalog.sources['peak']>good_amplitude
good_idx = good_idx & (ccd.catalog.sources['pipeline_flags']==0)
good_idx = good_idx & np.isfinite(ccd.catalog.sources['psf_mag'])
good_sources = ccd.catalog.sources[good_idx]
print('rms', np.sqrt(np.sum(good_sources['psf_mag_err']**2/len(good_sources))))
print('mean', np.mean(good_sources['psf_mag_err']))
print('median', np.median(good_sources['psf_mag_err']))
print('stddev', np.std(good_sources['psf_mag_err']))
bad_count = np.sum(good_sources['psf_mag_err']>.05)
print('bad psf error: {0}, or {1}%'.format(bad_count, bad_count/len(good_sources)*100))
print('Better than 5%: {0} of {1}'.format(np.sum(good_sources['psf_mag_err']<=.05), len(good_sources)))
print('Better than 2%: {0} of {1}'.format(np.sum(good_sources['psf_mag_err']<=.02), len(good_sources)))
good_sources['aper_flux','psf_flux','peak','psf_mag_err'][good_sources['psf_mag_err']>.05]
# -
import os
os.path.basename(os.path.join('this','is','a','test.fits'))
# +
idx_connect = 'sqlite:////media/data-beta/users/fmooleka/decam/decam.db'
def get_exp_info(obj, night, filtr, idx_connect):
sql = 'select * from decam_obs where object like "{0}%" '.format(obj)
sql += ' and filter like "{0}%" and dtcaldat="{1}"'.format(filtr, night)
if filtr=='i':
sql += ' and exptime>31'
else:
sql += ' and exptime>8'
exp_info = astropyp.db_utils.index.query(sql, idx_connect)
return exp_info
tbl = get_exp_info('F100', '2015-05-26', 'i', idx_connect)
#tbl[tbl['PRODTYPE']=='image']
# -
tbl['new filename'] = np.zeros((len(tbl), 1), dtype="U{0}".format(100))
tbl
tbl['new filename'][(tbl['EXPNUM']==442431) & (tbl['PRODTYPE']=='image')] = "If I were a rich man"
tbl
't:'+'/test/filename'
import bd_search
idx_connect = 'sqlite:////media/data-beta/users/fmooleka/decam/decam.db'
exp_info = bd_search.pipeline.get_all_exp_info('F100', '2015-05-26', 'i', idx_connect,
'/scratch/temp', '<EMAIL>:')
exp_info
|
examples/test_full_img_stacking.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from gensim.models import KeyedVectors
from gensim.scripts.glove2word2vec import glove2word2vec
# +
glove_file = 'vectors.txt'
word2vec_file = "word2Vec.txt"
glove2word2vec(glove_file, word2vec_file)
# To load glove word embeddings on gensim model we need to convert "vectors.txt" file to Word2Vec format.
# -
model = KeyedVectors.load_word2vec_format(word2vec_file)
# Similar Words
model.most_similar("kedi")
# +
# Synonyms & Antonyms
w1 = "fakir"
w2 = "yoksul"
w3 = "zengin"
w1_w2_dist = model.distance(w1, w2)
w1_w3_dist = model.distance(w1, w3)
print(f"Synonyms {w1}, {w2} have cosine distance: {w1_w2_dist}")
print(f"Antonyms {w1}, {w3} have cosine distance: {w1_w3_dist}")
# -
# Solving Analogies with Word Vectors
model.most_similar(positive=['fransa', 'berlin'], negative=['almanya'])
# Solving Analogies with Word Vectors
model.most_similar(positive=['geliyor', 'gitmek'], negative=['gelmek'])
|
src/examples/Exploring Word Vectors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # [UIUC Black Hole](https://arxiv.org/abs/1001.4077) Initial data
#
# ## Authors: <NAME>, <NAME>, & <NAME>
#
# ### Formatting improvements courtesy <NAME>
#
# ## This module sets up UIUC Black Hole initial data ([Liu, Etienne, & Shapiro, PRD 80 121503, 2009](https://arxiv.org/abs/1001.4077)).
#
# ### We can convert from spherical to any coordinate system defined in [reference_metric.py](../edit/reference_metric.py) (e.g., SinhSpherical, Cylindrical, Cartesian, etc.) using the [Exact ADM Spherical-or-Cartesian-to-BSSNCurvilinear converter module](Tutorial-ADM_Initial_Data-Converting_Exact_ADM_Spherical_or_Cartesian_to_BSSNCurvilinear.ipynb)
#
# **Module Status:** <font color='green'><b> Validated </b></font>
#
# **Validation Notes:** This module has been validated to exhibit convergence to zero of the Hamiltonian and momentum constraint violation at the expected order to the exact solution (see plots at bottom of [the exact initial data validation start-to-finish tutorial notebook](Tutorial-Start_to_Finish-BSSNCurvilinear-Setting_up_Exact_Initial_Data.ipynb); momentum constraint violation in non-$\phi$ directions is zero), and all quantities have been validated against the [original SENR code](https://bitbucket.org/zach_etienne/nrpy).
#
# ### NRPy+ Source Code for this module: [BSSN/UIUCBlackHole.py](../edit/BSSN/UIUCBlackHole.py)
#
# ## Introduction:
# UIUC black holes have the advantage of finite coordinate radius in the maximal spin limit. It is therefore excellent for studying very highly spinning black holes. This module sets the UIUC black hole at the origin.
# <a id='toc'></a>
#
# # Table of Contents:
# $$\label{toc}$$
#
# 1. [Step 1](#initializenrpy): Set up the needed NRPy+ infrastructure and declare core gridfunctions
# 1. [Step 2](#bl_radius): The Boyer-Lindquist Radius
# 1. [Step 2.a](#define_inner_outer_radii): Define the inner and outer radii
# 1. [Step 2.b](#define_bl_radius): Define the Boyer-Lindquist radius
# 1. [Step 3](#line_element): Define the line element, and extract components of $\gamma_{ij}$
# 1. [Step 4](#extrinsic_curvature): Define and construct nonzero components of the extrinsic curvature $K_{ij}$
# 1. [Step 5](#lapse_shift): Construct Lapse function $\alpha$ and components of shift vector $\beta$
# 1. [Step 6](#code_validation): Code Validation against `BSSN.UIUCBlackHole` NRPy+ module
# 1. [Step 7](#latex_pdf_output) Output this notebook to $\LaTeX$-formatted PDF file
#
#
# <a id='initializenrpy'></a>
#
# # Step 1: Set up the needed NRPy+ infrastructure and declare core gridfunctions \[Back to [top](#toc)\]
# $$\label{initializenrpy}$$
#
# First, we will import the core modules of Python/NRPy+ and specify the main gridfunctions that we will need.
# Second, we set some basic NRPy+ parameters. E.g., set the spatial dimension parameter to 3.
#
# **Inputs for initial data**:
#
# * The black hole mass $M$.
# * The dimensionless spin parameter $\chi = a/M$
#
# **Additional variables needed for spacetime evolution**:
#
# * Desired coordinate system Boyer-Lindquist coordinates $(r_{BL}, \theta, \phi)$
# <br>
# * Desired initial lapse $\alpha$ and shift $\beta^i$. We will choose our gauge conditions as $\alpha=1$ and $\beta^i=B^i=0$. $\alpha = \psi^{-2}$ will yield much better behavior, but the conformal factor $\psi$ depends on the desired *destination* coordinate system (which may not be spherical coordinates).
# +
# Step P0: Load needed modules
import sympy as sp
import NRPy_param_funcs as par
import indexedexp as ixp
import grid as gri
from outputC import *
import reference_metric as rfm
# All gridfunctions will be written in terms of spherical coordinates (r, th, ph):
r,th,ph = sp.symbols('r th ph', real=True)
thismodule = "UIUCBlackHole"
# Step 0: Set spatial dimension (must be 3 for BSSN)
DIM = 3
par.set_parval_from_str("grid::DIM",DIM)
# Step 1: Set psi, the conformal factor:
# The UIUC initial data represent a Kerr black hole with mass M
# and dimensionless spin chi in UIUC quasi-isotropic coordinates,
# see https://arxiv.org/abs/1001.4077
# Input parameters:
M,chi = par.Cparameters("REAL", thismodule, ["M","chi"],[1.0,0.99])
# Spin per unit mass
a = M*chi
# -
# <a id='bl_radius'></a>
#
# # Step 2: The Boyer-Lindquist Radius \[Back to [top](#toc)\]
# $$\label{bl_radius}$$
#
# <a id='define_inner_outer_radii'></a>
#
# ## Step 2.a: Defining the Inner and Outer Radii \[Back to [top](#toc)\]
# $$\label{define_inner_outer_radii}$$
#
#
#
# Boyer-Lindquist radii of the outer (+) and inner (−) horizons of the BH, defined under equation 1 in [Liu, Etienne, & Shapiro (2009)](https://arxiv.org/abs/1001.4077) as
# $$ r_{\pm} = M \pm \sqrt{M^2 - a^2}$$
# +
# Defined under equation 1 in Liu, Etienne, & Shapiro (2009)
# https://arxiv.org/pdf/1001.4077.pdf
# Boyer - Lindquist outer horizon
rp = M + sp.sqrt(M**2 - a**2)
# Boyer - Lindquist inner horizon
rm = M - sp.sqrt(M**2 - a**2)
# -
# <a id='define_bl_radius'></a>
#
# ## Step 2.b: Define the Boyer-Lindquist Radius \[Back to [top](#toc)\]
# $$\label{define_bl_radius}$$
#
# Define $r_{BL}$, equation 11 of [Liu, Etienne, & Shapiro (2009)](https://arxiv.org/abs/1001.4077), using the radial coordinate $r$:
#
# $$ r_{BL} = r \left( 1 + \frac{r_+}{4r}\right)^2. $$
# Boyer - Lindquist radius in terms of UIUC radius
# Eq. 11
# r_{BL} = r * ( 1 + r_+ / 4r )^2
rBL = r*(1 + rp / (4*r))**2
# Quantities used to calculate the spatial metric $\gamma_{ij}$, found under equation 2 of [<NAME>, & Shapiro (2009)](https://arxiv.org/abs/1001.4077):
# $$ \Sigma = r_{BL}^2 + a^2 \cos^2 \theta, $$
#
# $$ \Delta = r_{BL}^2 - 2Mr_{BL} + a^2, $$
#
# $$ A = \left(r_{BL}^2 + a^2\right)^2 - \Delta a^2 \sin^2 \theta. $$
# +
# Expressions found below Eq. 2
# Sigma = r_{BL}^2 + a^2 cos^2 theta
SIG = rBL**2 + a**2*sp.cos(th)**2
# Delta = r_{BL}^2 - 2Mr_{BL} + a^2
DEL = rBL**2 - 2*M*rBL + a**2
# A = (r_{BL}^2 + a^2)^2 - Delta a^2 sin^2 theta
AA = (rBL**2 + a**2)**2 - DEL*a**2*sp.sin(th)**2
# -
# <a id='line_element'></a>
#
# # Step 3: Define the Line element and extract components of $\gamma_{ij}$ \[Back to [top](#toc)\]
# $$\label{line_element}$$
#
# The line element, defined in equation 13 of [<NAME>, & Shapiro (2009)](https://arxiv.org/abs/1001.4077):
#
# $$ ds^2 = \frac{\Sigma\left(r + \frac{r_+}{4}\right)^2 } {r^3 \left(r_{BL} - r_- \right)} dr^2 + \Sigma d\theta^2 + \frac{ A \sin^2 \theta }{\Sigma} d\phi^2 $$
# +
# *** The ADM 3-metric in spherical basis ***
gammaSphDD = ixp.zerorank2()
# Declare the nonzero components of the 3-metric (Eq. 13):
# ds^2 = Sigma (r + r_+/4)^2 / ( r^3 (r_{BL} - r_- ) * dr^2 +
# Sigma d theta^2 + (A sin^2 theta) / Sigma * d\phi^2
gammaSphDD[0][0] = ((SIG*(r + rp/4)**2)/(r**3*(rBL - rm)))
gammaSphDD[1][1] = SIG
gammaSphDD[2][2] = AA/SIG*sp.sin(th)**2
# -
# <a id='extrinsic_curvature'></a>
#
# # Step 4: Define and construct nonzero components of extrinsic curvature $K_{ij}$ \[Back to [top](#toc)\]
# $$\label{extrinsic_curvature}$$
#
#
#
# Nonzero components of the extrinsic curvature, equation 14 of [Liu, Etienne, & Shapiro (2009)](https://arxiv.org/abs/1001.4077):
#
# $$ K_{r\phi} = K_{\phi r} = \frac{Ma\sin^2\theta}{\Sigma\sqrt{A\Sigma}} \
# \left[3r^4_{BL} + 2a^2 r^2_{BL} - a^4 - a^2 \left(r^2_{BL} - a^2\right) \sin^2 \theta\right] \
# \left(1 + \frac{r_+}{4r}\right) \frac{1}{\sqrt{r(r_{BL} - r_-)}} $$
# +
# *** The physical trace-free extrinsic curvature in spherical basis ***
# Nonzero components of the extrinsic curvature K, given by
# Eq. 14 of Liu, Etienne, & Shapiro, https://arxiv.org/pdf/1001.4077.pdf:
KSphDD = ixp.zerorank2() # K_{ij} = 0 for these initial data
# K_{r phi} = K_{phi r} = (Ma sin^2 theta) / (Sigma sqrt{A Sigma}) *
# [3r^4_{BL} + 2a^2 r^2_{BL} - a^4 - a^2 (r^2_{BL} - a^2) sin^2 theta] *
# (1 + r_+ / 4r) (1 / sqrt{r(r_{BL} - r_-)})
KSphDD[0][2] = KSphDD[2][0] = (M*a*sp.sin(th)**2)/(SIG*sp.sqrt(AA*SIG))*\
(3*rBL**4 + 2*a**2*rBL**2 - a**4- a**2*(rBL**2 - a**2)*\
sp.sin(th)**2)*(1 + rp/(4*r))*1/sp.sqrt(r*(rBL - rm))
# -
# Nonzero components of the extrinsic curvature, equation 15 of [Liu, Etienne, & Shapiro (2009)](https://arxiv.org/abs/1001.4077):
#
# $$ K_{\theta\phi} = K_{\phi\theta} = -\frac{2a^3 Mr_{BL}\cos\theta \sin^3\theta} {\Sigma \sqrt{A\Sigma} } \left(r - \frac{r_+}{4}\right) \sqrt {\frac{r_{BL} - r_-}{r} } $$
# +
# Components of the extrinsic curvature K, given by
# Eq. 15 of Liu, Etienne, & Shapiro, https://arxiv.org/pdf/1001.4077.pdf:
# K_{theta phi} = K_{phi theta} = -(2a^3 Mr_{BL} cos theta sin^3 theta) /
# (Sigma sqrt{A Sigma}) x (r - r_+ / 4) sqrt{(r_{BL} - r_-) / r }
KSphDD[1][2] = KSphDD[2][1] = -((2*a**3*M*rBL*sp.cos(th)*sp.sin(th)**3)/ \
(SIG*sp.sqrt(AA*SIG)))*(r - rp/4)*sp.sqrt((rBL - rm)/r)
# -
# <a id='lapse_shift'></a>
#
# # Step 5: Construct Lapse function $\alpha$ and components of shift vector $\beta$ \[Back to [top](#toc)\]
# $$\label{lapse_shift}$$
#
# $$\alpha=1$$
# <br>
# $$\beta^i=B^i=0$$
# +
alphaSph = sp.sympify(1)
betaSphU = ixp.zerorank1() # We generally choose \beta^i = 0 for these initial data
BSphU = ixp.zerorank1() # We generally choose B^i = 0 for these initial data
# Validated against original SENR: KSphDD[0][2], KSphDD[1][2], gammaSphDD[2][2], gammaSphDD[0][0], gammaSphDD[1][1]
# print(sp.mathematica_code(gammaSphDD[1][1]))
# -
# <a id='code_validation'></a>
#
# # Step 6: Code Validation against `BSSN.UIUCBlackHole` NRPy+ module \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
#
# Here, as a code validation check, we verify agreement in the SymPy expressions for UIUC black hole initial data between
#
# 1. this tutorial and
# 2. the NRPy+ [BSSN.UIUCBlackHole](../edit/BSSN/UIUCBlackHole.py) module.
# +
# Reset the list of gridfunctions, as registering a gridfunction
# twice will spawn an error.
gri.glb_gridfcs_list = []
# First we import needed core NRPy+ modules
import reference_metric as rfm
import grid as gri
# Unused, but needed because BrillLindquist() also does tensor rescalings.
rfm.reference_metric()
import BSSN.UIUCBlackHole as uibh
uibh.UIUCBlackHole()
print("Consistency check between Brill-Lindquist tutorial and NRPy+ BSSN.BrillLindquist module. ALL SHOULD BE ZERO.")
print("alphaSph - uibh.alphaSph = "+str(sp.simplify(alphaSph - uibh.alphaSph)))
for i in range(DIM):
print("betaSphU["+str(i)+"] - uibh.betaSphU["+str(i)+"] = "+\
str(sp.simplify(betaSphU[i] - uibh.betaSphU[i])))
print("BSphU["+str(i)+"] - uibh.BaSphU["+str(i)+"] = "+str(sp.simplify(BSphU[i] - uibh.BSphU[i])))
for j in range(DIM):
print("gammaSphDD["+str(i)+"]["+str(j)+"] - uibh.gammaSphDD["+str(i)+"]["+str(j)+"] = "+\
str(sp.simplify(gammaSphDD[i][j] - uibh.gammaSphDD[i][j])))
print("KSphDD["+str(i)+"]["+str(j)+"] - uibh.KSphDD["+str(i)+"]["+str(j)+"] = "+\
str(sp.simplify(KSphDD[i][j] - uibh.KSphDD[i][j])))
# -
# <a id='latex_pdf_output'></a>
#
# # Step 7: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [Tutorial-ADM_Initial_Data-UIUC_BlackHole.pdf](Tutorial-ADM_Initial_Data-UIUC_BlackHole.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
# !jupyter nbconvert --to latex --template latex_nrpy_style.tplx Tutorial-ADM_Initial_Data-UIUC_BlackHole.ipynb
# !pdflatex -interaction=batchmode Tutorial-ADM_Initial_Data-UIUC_BlackHole.tex
# !pdflatex -interaction=batchmode Tutorial-ADM_Initial_Data-UIUC_BlackHole.tex
# !pdflatex -interaction=batchmode Tutorial-ADM_Initial_Data-UIUC_BlackHole.tex
# !rm -f Tut*.out Tut*.aux Tut*.log
|
notebook/Tutorial-ADM_Initial_Data-UIUC_BlackHole.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Desafio 1
#
# Para esse desafio, vamos trabalhar com o data set [Black Friday](https://www.kaggle.com/mehdidag/black-friday), que reúne dados sobre transações de compras em uma loja de varejo.
#
# Vamos utilizá-lo para praticar a exploração de data sets utilizando pandas. Você pode fazer toda análise neste mesmo notebook, mas as resposta devem estar nos locais indicados.
#
# > Obs.: Por favor, não modifique o nome das funções de resposta.
# ## _Set up_ da análise
import pandas as pd
import numpy as np
black_friday = pd.read_csv("black_friday.csv")
# ## Inicie sua análise a partir daqui
black_friday.head()
df = black_friday
aux = black_friday[black_friday['Product_Category_2'].isnull()]
aux['Product_Category_2'].equals(aux['Product_Category_3'])
black_friday.query('Gender == \"F\" and Age==\"26-35\"').count()
aux[aux['Product_Category_3']]
df.head()
df.info()
# ## Questão 1
#
# Quantas observações e quantas colunas há no dataset? Responda no formato de uma tuple `(n_observacoes, n_colunas)`.
def q1():
# Retorne aqui o resultado da questão 1.
return df.shape
# ## Questão 2
#
# Há quantas mulheres com idade entre 26 e 35 anos no dataset? Responda como um único escalar.
def q2():
# Retorne aqui o resultado da questão 2.
female_filtered = df.loc[(df['Age'] == '26-35') & (df['Gender'] == 'F')]
return int(female_filtered.shape[0])
# ## Questão 3
#
# Quantos usuários únicos há no dataset? Responda como um único escalar.
def q3():
# Retorne aqui o resultado da questão 3.
return df['User_ID'].nunique()
# ## Questão 4
#
# Quantos tipos de dados diferentes existem no dataset? Responda como um único escalar.
def q4():
# Retorne aqui o resultado da questão 4.
return df.dtypes.nunique()
# ## Questão 5
#
# Qual porcentagem dos registros possui ao menos um valor null (`None`, `ǸaN` etc)? Responda como um único escalar entre 0 e 1.
def q5():
# Retorne aqui o resultado da questão 5.
return (df.shape[0] - df.dropna().shape[0]) / df.shape[0]
# ## Questão 6
#
# Quantos valores null existem na variável (coluna) com o maior número de null? Responda como um único escalar.
def q6():
# Retorne aqui o resultado da questão 6.
return int(df.isna().sum().max())
# ## Questão 7
#
# Qual o valor mais frequente (sem contar nulls) em `Product_Category_3`? Responda como um único escalar.
def q7():
# Retorne aqui o resultado da questão 7.
cleaned_column = df['Product_Category_3'].dropna()
return cleaned_column.mode()[0]
# ## Questão 8
#
# Qual a nova média da variável (coluna) `Purchase` após sua normalização? Responda como um único escalar.
def q8():
# Retorne aqui o resultado da questão 8.
min_max = (df['Purchase'] - df['Purchase'].min()) \
/ (df['Purchase'].max() - df['Purchase'].min())
return float(min_max.mean())
# ## Questão 9
#
# Quantas ocorrências entre -1 e 1 inclusive existem da variáel `Purchase` após sua padronização? Responda como um único escalar.
# +
def q9():
# Retorne aqui o resultado da questão 9.
df_normalized = (df['Purchase'] - df['Purchase'].mean()) / df['Purchase'].std()
return int(df_normalized.between(-1,1).sum())
q9()
# -
# ## Questão 10
#
# Podemos afirmar que se uma observação é null em `Product_Category_2` ela também o é em `Product_Category_3`? Responda com um bool (`True`, `False`).
def q10():
# Retorne aqui o resultado da questão 10.
df_aux = df[['Product_Category_2','Product_Category_3']]
df_aux = df_aux[df_aux['Product_Category_2'].isna()]
return df_aux['Product_Category_2'].equals(df_aux['Product_Category_3'])
|
data-science-0/main.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .r
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# + [markdown] colab_type="text" id="copyright-notice"
# #### Copyright 2017 Google LLC.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="copyright-notice2"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="eV16J6oUY-HN" slideshow={"slide_type": "slide"}
# # 神经网络简介
# + [markdown] colab_type="text" id="_wIcUFLSKNdx"
# **学习目标:**
# * 使用 TensorFlow `DNNRegressor` 类定义神经网络 (NN) 及其隐藏层
# * 训练神经网络学习数据集中的非线性规律,并实现比线性回归模型更好的效果
# + [markdown] colab_type="text" id="_ZZ7f7prKNdy"
# 在之前的练习中,我们使用合成特征来帮助模型学习非线性规律。
#
# 一组重要的非线性关系是纬度和经度的关系,但也可能存在其他非线性关系。
#
# 现在我们从之前练习中的逻辑回归任务回到标准的(线性)回归任务。也就是说,我们将直接预测 `median_house_value`。
# + [markdown] colab_type="text" id="J2kqX6VZTHUy"
# ## 设置
#
# 首先加载和准备数据。
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="AGOM1TUiKNdz"
import math
from IPython import display
from matplotlib import cm
from matplotlib import gridspec
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
from sklearn import metrics
import tensorflow as tf
from tensorflow.python.data import Dataset
tf.logging.set_verbosity(tf.logging.ERROR)
pd.options.display.max_rows = 10
pd.options.display.float_format = '{:.1f}'.format
california_housing_dataframe = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_train.csv", sep=",")
california_housing_dataframe = california_housing_dataframe.reindex(
np.random.permutation(california_housing_dataframe.index))
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="2I8E2qhyKNd4"
def preprocess_features(california_housing_dataframe):
"""Prepares input features from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the features to be used for the model, including
synthetic features.
"""
selected_features = california_housing_dataframe[
["latitude",
"longitude",
"housing_median_age",
"total_rooms",
"total_bedrooms",
"population",
"households",
"median_income"]]
processed_features = selected_features.copy()
# Create a synthetic feature.
processed_features["rooms_per_person"] = (
california_housing_dataframe["total_rooms"] /
california_housing_dataframe["population"])
return processed_features
def preprocess_targets(california_housing_dataframe):
"""Prepares target features (i.e., labels) from California housing data set.
Args:
california_housing_dataframe: A Pandas DataFrame expected to contain data
from the California housing data set.
Returns:
A DataFrame that contains the target feature.
"""
output_targets = pd.DataFrame()
# Scale the target to be in units of thousands of dollars.
output_targets["median_house_value"] = (
california_housing_dataframe["median_house_value"] / 1000.0)
return output_targets
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="pQzcj2B1T5dA"
# Choose the first 12000 (out of 17000) examples for training.
training_examples = preprocess_features(california_housing_dataframe.head(12000))
training_targets = preprocess_targets(california_housing_dataframe.head(12000))
# Choose the last 5000 (out of 17000) examples for validation.
validation_examples = preprocess_features(california_housing_dataframe.tail(5000))
validation_targets = preprocess_targets(california_housing_dataframe.tail(5000))
# Double-check that we've done the right thing.
print "Training examples summary:"
display.display(training_examples.describe())
print "Validation examples summary:"
display.display(validation_examples.describe())
print "Training targets summary:"
display.display(training_targets.describe())
print "Validation targets summary:"
display.display(validation_targets.describe())
# + [markdown] colab_type="text" id="RWq0xecNKNeG"
# ## 构建神经网络
#
# 神经网络由 [DNNRegressor](https://www.tensorflow.org/api_docs/python/tf/contrib/learn/DNNRegressor) 类定义。
#
# 使用 **`hidden_units`** 定义神经网络的结构。`hidden_units` 参数会创建一个整数列表,其中每个整数对应一个隐藏层,表示其中的节点数。以下面的赋值为例:
#
# `hidden_units=[3,10]`
#
# 上述赋值为神经网络指定了两个隐藏层:
#
# * 第一个隐藏层包含 3 个节点。
# * 第二个隐藏层包含 10 个节点。
#
# 如果我们想要添加更多层,可以向该列表添加更多整数。例如,`hidden_units=[10,20,30,40]` 会创建 4 个分别包含 10、20、30 和 40 个单元的隐藏层。
#
# 默认情况下,所有隐藏层都会使用 ReLu 激活函数,且是全连接层。
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="ni0S6zHcTb04"
def construct_feature_columns(input_features):
"""Construct the TensorFlow Feature Columns.
Args:
input_features: The names of the numerical input features to use.
Returns:
A set of feature columns
"""
return set([tf.feature_column.numeric_column(my_feature)
for my_feature in input_features])
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="zvCqgNdzpaFg"
def my_input_fn(features, targets, batch_size=1, shuffle=True, num_epochs=None):
"""Trains a linear regression model of one feature.
Args:
features: pandas DataFrame of features
targets: pandas DataFrame of targets
batch_size: Size of batches to be passed to the model
shuffle: True or False. Whether to shuffle the data.
num_epochs: Number of epochs for which data should be repeated. None = repeat indefinitely
Returns:
Tuple of (features, labels) for next data batch
"""
# Convert pandas data into a dict of np arrays.
features = {key:np.array(value) for key,value in dict(features).items()}
# Construct a dataset, and configure batching/repeating
ds = Dataset.from_tensor_slices((features,targets)) # warning: 2GB limit
ds = ds.batch(batch_size).repeat(num_epochs)
# Shuffle the data, if specified
if shuffle:
ds = ds.shuffle(10000)
# Return the next batch of data
features, labels = ds.make_one_shot_iterator().get_next()
return features, labels
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="U52Ychv9KNeH"
def train_nn_regression_model(
learning_rate,
steps,
batch_size,
hidden_units,
training_examples,
training_targets,
validation_examples,
validation_targets):
"""Trains a neural network regression model.
In addition to training, this function also prints training progress information,
as well as a plot of the training and validation loss over time.
Args:
learning_rate: A `float`, the learning rate.
steps: A non-zero `int`, the total number of training steps. A training step
consists of a forward and backward pass using a single batch.
batch_size: A non-zero `int`, the batch size.
hidden_units: A `list` of int values, specifying the number of neurons in each layer.
training_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for training.
training_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for training.
validation_examples: A `DataFrame` containing one or more columns from
`california_housing_dataframe` to use as input features for validation.
validation_targets: A `DataFrame` containing exactly one column from
`california_housing_dataframe` to use as target for validation.
Returns:
A `LinearRegressor` object trained on the training data.
"""
periods = 10
steps_per_period = steps / periods
# Create a linear regressor object.
my_optimizer = tf.train.GradientDescentOptimizer(learning_rate=learning_rate)
my_optimizer = tf.contrib.estimator.clip_gradients_by_norm(my_optimizer, 5.0)
dnn_regressor = tf.estimator.DNNRegressor(
feature_columns=construct_feature_columns(training_examples),
hidden_units=hidden_units
)
# Create input functions
training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value"],
batch_size=batch_size)
predict_training_input_fn = lambda: my_input_fn(training_examples,
training_targets["median_house_value"],
num_epochs=1,
shuffle=False)
predict_validation_input_fn = lambda: my_input_fn(validation_examples,
validation_targets["median_house_value"],
num_epochs=1,
shuffle=False)
# Train the model, but do so inside a loop so that we can periodically assess
# loss metrics.
print "Training model..."
print "RMSE (on training data):"
training_rmse = []
validation_rmse = []
for period in range (0, periods):
# Train the model, starting from the prior state.
dnn_regressor.train(
input_fn=training_input_fn,
steps=steps_per_period
)
# Take a break and compute predictions.
training_predictions = dnn_regressor.predict(input_fn=predict_training_input_fn)
training_predictions = np.array([item['predictions'][0] for item in training_predictions])
validation_predictions = dnn_regressor.predict(input_fn=predict_validation_input_fn)
validation_predictions = np.array([item['predictions'][0] for item in validation_predictions])
# Compute training and validation loss.
training_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(training_predictions, training_targets))
validation_root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(validation_predictions, validation_targets))
# Occasionally print the current loss.
print " period %02d : %0.2f" % (period, training_root_mean_squared_error)
# Add the loss metrics from this period to our list.
training_rmse.append(training_root_mean_squared_error)
validation_rmse.append(validation_root_mean_squared_error)
print "Model training finished."
# Output a graph of loss metrics over periods.
plt.ylabel("RMSE")
plt.xlabel("Periods")
plt.title("Root Mean Squared Error vs. Periods")
plt.tight_layout()
plt.plot(training_rmse, label="training")
plt.plot(validation_rmse, label="validation")
plt.legend()
print "Final RMSE (on training data): %0.2f" % training_root_mean_squared_error
print "Final RMSE (on validation data): %0.2f" % validation_root_mean_squared_error
return dnn_regressor
# + [markdown] colab_type="text" id="2QhdcCy-Y8QR" slideshow={"slide_type": "slide"}
# ## 任务 1:训练神经网络模型
#
# **调整超参数,目标是将 RMSE 降到 110 以下。**
#
# 运行以下代码块来训练神经网络模型。
#
# 我们已经知道,在使用了很多特征的线性回归练习中,110 左右的 RMSE 已经是相当不错的结果。我们将得到比它更好的结果。
#
# 在此练习中,您的任务是修改各种学习设置,以提高在验证数据上的准确率。
#
# 对于神经网络而言,过拟合是一种真正的潜在危险。您可以查看训练数据损失与验证数据损失之间的差值,以帮助判断模型是否有过拟合的趋势。如果差值开始变大,则通常可以肯定存在过拟合。
#
# 由于存在很多不同的可能设置,强烈建议您记录每次试验,以在开发流程中进行参考。
#
# 此外,获得效果出色的设置后,尝试多次运行该设置,看看结果的重复程度。由于神经网络权重通常会初始化为较小的随机值,因此每次运行结果应该存在差异。
#
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="rXmtSW1yKNeK"
dnn_regressor = train_nn_regression_model(
learning_rate=0.01,
steps=500,
batch_size=10,
hidden_units=[10, 2],
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
# + [markdown] colab_type="text" id="O2q5RRCKqYaU"
# ### 解决方案
#
# 点击下方即可查看可能的解决方案
# + [markdown] colab_type="text" id="j2Yd5VfrqcC3"
# **注意**:在本次练习中,参数的选择有点随意。我们尝试了越来越复杂的组合,并进行了较长时间的训练,直到误差降到目标之下。这决不是最佳组合;其他组合可能会获得更低的 RMSE。如果您的目标是找到可以产生最小误差的模型,那么您需要使用更严格的流程,例如参数搜索。
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="IjkpSqmxqnSM"
dnn_regressor = train_nn_regression_model(
learning_rate=0.001,
steps=2000,
batch_size=100,
hidden_units=[10, 10],
training_examples=training_examples,
training_targets=training_targets,
validation_examples=validation_examples,
validation_targets=validation_targets)
# + [markdown] colab_type="text" id="c6diezCSeH4Y" slideshow={"slide_type": "slide"}
# ## 任务 2:用测试数据进行评估
#
# **确认您的验证效果结果经受得住测试数据的检验。**
#
# 获得满意的模型后,用测试数据评估该模型,以与验证效果进行比较。
#
# 提示:测试数据集位于[此处](https://storage.googleapis.com/mledu-datasets/california_housing_test.csv)。
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "test": {"output": "ignore", "timeout": 600}} colab_type="code" id="icEJIl5Vp51r"
california_housing_test_data = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_test.csv", sep=",")
# YOUR CODE HERE
# + [markdown] colab_type="text" id="vvT2jDWjrKew"
# ### 解决方案
#
# 点击下方即可查看可能的解决方案。
# + [markdown] colab_type="text" id="FyDh7Qy6rQb0"
# 与顶部代码类似,我们只需加载合适的数据文件、对其进行预处理并调用预测和 mean_squared_error 即可。
#
# 请注意,由于我们会使用所有记录,因此无需对测试数据进行随机化处理。
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="vhb0CtdvrWZx"
california_housing_test_data = pd.read_csv("https://storage.googleapis.com/mledu-datasets/california_housing_test.csv", sep=",")
test_examples = preprocess_features(california_housing_test_data)
test_targets = preprocess_targets(california_housing_test_data)
predict_testing_input_fn = lambda: my_input_fn(test_examples,
test_targets["median_house_value"],
num_epochs=1,
shuffle=False)
test_predictions = dnn_regressor.predict(input_fn=predict_testing_input_fn)
test_predictions = np.array([item['predictions'][0] for item in test_predictions])
root_mean_squared_error = math.sqrt(
metrics.mean_squared_error(test_predictions, test_targets))
print "Final RMSE (on test data): %0.2f" % root_mean_squared_error
|
ml/cc/exercises/zh-CN/intro_to_neural_nets.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.1.0
# language: julia
# name: julia-1.1
# ---
# # MLP GAN MNIST
# This notebook is a demonstration for a simple GAN training on MNIST by using KnetLayers. Let's get started!
using Pkg; for p in ("Colors","ImageMagick","Images"); haskey(Pkg.installed(),p) || Pkg.add(p); end
using Knet, KnetLayers, Colors, ImageMagick, Images, Statistics
import Knet: Data
import KnetLayers: arrtype
setoptim!(model,optimizer) = for p in params(model) p.opt=Knet.clone(optimizer) end # Easy optimizer setter
#Data
include(Knet.dir("data","mnist.jl"))
# ## Discriminator and Generator Networks
#
# Discriminator and Generator networks are defined as `D` and `G` respectively. Loss functions `𝑱d` and `𝑱g` are defined according to the equation X in GAN paper. Sample noise function `𝒩` is a normal distribution. Loss functions are defined according to the equations in Algorithm 1 section of the [paper](https://arxiv.org/abs/1406.2661 "arXiv"). We use a slightly modified generator loss according to [GAN tricks](https://github.com/soumith/ganhacks#2-a-modified-loss-function "GAN Tricks").
# $$ J_d = -\frac{1}{m} \sum_{i=1}^{m} log(D(x^{(i)}) + log(1-D(G(z^{(i)})))$$
# $$ J_g = -\frac{1}{m} \sum_{i=1}^{m} log(D(G(z^{(i)}))) $$
#
# *`𝜀` is used to prevent log functions from resulting NaN values.
global const 𝜀=Float32(1e-8)
𝑱d(D,x,Gz) = -mean(log.(D(x) .+ 𝜀) .+ log.((1+𝜀) .- D(Gz)))/2 #discriminator loss
𝑱g(D,G,z) = -mean(log.(D(G(z)) .+ 𝜀)) # generator loss
𝒩(input, batch) = arrtype(randn(Float32, input, batch)) #sample noise
# ## Train & Test Function
#
# This `runmodel` function is implementing training procedure described in GAN paper. It first update discriminator with specified optimizer, then update generator network. Same function can be used in test mode by passing `train` argument as false. In the test mode it calculates losses instead of gradients.
function runmodel(D, G, data, 𝞗; dtst=nothing, train=false, saveinterval=20)
gloss = dloss = total= 0.0; B = 𝞗[:batchsize]
if train
Dprms, Gprms, L = params(D), params(G), 𝞗[:epochs]
else
Dprms, Gprms, L = nothing, nothing, 1
end
for i=1:L
for (x,_) in data
Gz = G(𝒩(𝞗[:ginp], B)) #Generate Fake Images
z = 𝒩(𝞗[:ginp], 2B) #Sample z from Noise
if train
jd = @diff 𝑱d(D, x, Gz)
for w in Dprms update!(w,grad(jd,w)) end
jg = @diff 𝑱g(D, G, z)
for w in Gprms update!(w,grad(jg,w)) end
else
jd = 𝑱d(D, x, Gz)
jg = 𝑱g(D, G, z)
end
dloss += 2B*value(jd); gloss += 2B*value(jg); total += 2B
end
train ? runmodel(D, G, dtst, 𝞗; train=false) : println((gloss/total, dloss/total))
i % saveinterval == 0 && generate_and_show(D, G, 100, 𝞗) # save 10 images
end
end
# ## Generate and Display
#
# This function generates a random `number` of images and displays them.
function generate_and_show(D,G,number,𝞗)
Gz = convert(Array,G(𝒩(𝞗[:ginp], number))) .> 0.5
Gz = reshape(Gz, (28, 28, number))
L = floor(Int, sqrt(number))
grid = []
for i = 1:L:number
push!(grid, reshape(permutedims(Gz[:,:,i:i+L-1], (2,3,1)), (L*28,28)))
end
display(Gray.(hcat(grid...)))
end
# ## Model&Data Run
# We will load MNIST data and initalize our MLP models
#
𝞗 = Dict(:batchsize=>32,:epochs=>80,:ginp=>256,:genh=>512,:disch=>512,:optim=>Adam(;lr=0.0002))
G = Chain(MLP(𝞗[:ginp], 𝞗[:genh], 784; activation=ELU()), Sigm())
D = Chain(MLP(784, 𝞗[:disch], 1; activation=ELU()), Sigm())
setoptim!(D, 𝞗[:optim]); setoptim!(G, 𝞗[:optim])
xtrn,ytrn,xtst,ytst = mnist()
global dtrn,dtst = mnistdata(xsize=(784,:),xtype=arrtype, batchsize=𝞗[:batchsize])
# ## RUN!
generate_and_show(D, G, 100, 𝞗)
runmodel(D, G, dtst, 𝞗; train=false) # initial losses
runmodel(D, G, dtrn, 𝞗; train=true, dtst=dtst) # training
|
examples/gan-mlp.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
from model import Unet
import sys
sys.path.append("../tools")
from pycocotools.coco import COCO
import pycocotools.mask
from utils import img_generator
images_train = img_generator('images_train.json')
images_val = img_generator('images_val.json')
coco = COCO("../annotations/instances_train2014.json")
for image in images_train:
annotation = coco.loadAnns(ids = [image["id"]])
break
annotation
image
import skimage.io as io
import matplotlib.pyplot as plt
print(imgIDs[0])
image = coco.loadImgs([imgIDs[0]])
img = io.imread("../train2014/{}".format(img[0]["file_name"]))
plt.imshow(img)
# +
from skimage.transform import resize
img_resized = resize(img, (512,512,3))
plt.imshow(img_resized)
# -
plt.imshow(img)
annIds = coco.getAnnIds(imgIds=imgIDs[0])
anns = coco.loadAnns(annIds)
coco.showAnns(anns)
anns
annIds
imgIDs = coco.getImgIds(catIds=[1])
imgIDs
len(imgIDs)
from pycocotools.mask import decode
sample = anns[1]
sample['segmentation'][0]
plt.imshow(coco.annToMask(anns[0]))
import numpy as np
np.sum(coco.annToMask(anns[0]))
mask_resized = resize(coco.annToMask(anns[0]), (512,512))
plt.imshow(mask_resized)
plt.imshow(mask_resized)
import cv2
mask_resized = cv2.resize(coco.annToMask(anns[0]), (512,512))
plt.imshow(mask_resized)
import numpy as np
np.shape(coco.annToMask(anns[1]))
anns[1]
# +
catIDs = list(range(1,91))
y = np.zeros((image['height'], image['width'], 90))
for cat in catIDs:
annIds = coco.getAnnIds(imgIds=image['id'],catIds=[cat])
anns = coco.loadAnns(annIds)
if len(anns) > 0:
for ann in anns:
mask = coco.annToMask(ann)
y[:,:,cat] = np.logical_or(y[:,:,cat], mask).astype(int)
# -
len(annIds)
y.shape
annIds
x = np.asarray([[0,1,1],[1,0,1]])
y = np.asarray([[0,1,0],[1,1,1]])
z = np.logical_and(x,y)
x
y
z
z = z.astype(int)
z
|
Unet/.ipynb_checkpoints/coco-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/aamini/introtodeeplearning_labs/blob/master/lab2/Part2_debiasing_solution.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="Ag_e7xtTzT1W"
# <table align="center">
# <td align="center"><a target="_blank" href="http://introtodeeplearning.com">
# <img src="http://introtodeeplearning.com/images/colab/mit.png" style="padding-bottom:5px;" />
# Visit MIT Deep Learning</a></td>
# <td align="center"><a target="_blank" href="https://colab.research.google.com/github/aamini/introtodeeplearning_labs/blob/master/lab2/Part2_debiasing_solution.ipynb">
# <img src="http://introtodeeplearning.com/images/colab/colab.png?v2.0" style="padding-bottom:5px;" />Run in Google Colab</a></td>
# <td align="center"><a target="_blank" href="https://github.com/aamini/introtodeeplearning_labs/blob/master/lab2/Part2_debiasing_solution.ipynb">
# <img src="http://introtodeeplearning.com/images/colab/github.png" height="70px" style="padding-bottom:5px;" />View Source on GitHub</a></td>
# </table>
#
#
# # Laboratory 2: Computer Vision
#
# # Part 2: Debiasing Facial Detection Systems
#
# In the second portion of the lab, we'll explore two prominent aspects of applied deep learning: facial detection and algorithmic bias.
#
# Deploying fair, unbiased AI systems is critical to their long-term acceptance. Consider the task of facial detection: given an image, is it an image of a face? [Recent work from the MIT Media Lab](http://proceedings.mlr.press/v81/buolamwini18a/buolamwini18a.pdf) showed that this seemingly simple, but extremely important, task is subject to extreme amounts of algorithmic bias among select demographics. [Another report](https://ieeexplore.ieee.org/document/6327355) analyzed the face detection system used by the US law enforcement and found that it had significantly lower accuracy among dark skinned women between the age of 18-30 years old.
#
# Run the next code block for a short video from Google that explores how and why it's important to consider bias when thinking about machine learning:
# + colab={"base_uri": "https://localhost:8080/", "height": 322} colab_type="code" id="XQh5HZfbupFF" outputId="4a6df36f-b4b5-442c-b735-0cce3ad31e71"
from IPython.display import YouTubeVideo
YouTubeVideo('59bMh59JQDo')
# + [markdown] colab_type="text" id="3Ezfc6Yv6IhI"
# In this lab, we'll investigate [one recently published approach](http://introtodeeplearning.com/AIES_2019_Algorithmic_Bias.pdf) to addressing algorithmic bias. We'll build a facial detection model that learns the *latent variables* underlying face image datasets and uses this to adaptively re-sample the training data, thus mitigating any biases that may be present in order to train a *debiased* model.
#
# Let's get started by installing the relevant dependencies:
# + colab={"base_uri": "https://localhost:8080/", "height": 69} colab_type="code" id="E46sWVKK6LP9" outputId="7fb31396-0d46-4446-dada-87d4f9f9f26f"
import tensorflow as tf
tf.enable_eager_execution()
import functools
import matplotlib.pyplot as plt
import numpy as np
import pdb
# Download the class repository
# #! git clone https://github.com/aamini/introtodeeplearning_labs.git > /dev/null 2>&1
#% cd introtodeeplearning_labs
# #! git pull
#% cd ..
# Import the necessary class-specific utility files for this lab
import util
# + [markdown] colab_type="text" id="V0e77oOM3udR"
# ## 2.1 Datasets
#
# We'll be using three datasets in this lab. In order to train our facial detection models, we'll need a dataset of positive examples (i.e., of faces) and a dataset of negative examples (i.e., of things that are not faces). We'll use these data to train our models to classify images as either faces or not faces. Finally, we'll need a test dataset of face images. Since we're concerned about the potential *bias* of our learned models against certain demographics, it's important that the test dataset we use has equal representation across the demographics or features of interest. In this lab, we'll consider skin tone and gender.
#
# 1. **Positive training data**: [CelebA Dataset](http://mmlab.ie.cuhk.edu.hk/projects/CelebA.html). A large-scale (over 200K images) of celebrity faces.
# 2. **Negative training data**: [ImageNet](http://www.image-net.org/). Many images across many different categories. We'll take negative examples from a variety of non-human categories.
# 3. **Test data**: [Pilot Parliaments Benchmark](http://proceedings.mlr.press/v81/buolamwini18a/buolamwini18a.pdf) (PPB). The PPB dataset consists of images of 1270 male and female parliamentarians from various African and European countries and exhibits parity in both skin tone and gender. The gender of each face is annotated with the sex-based "Male'' and "Female'' labels. Skin tone annotations are based on the Fitzpatrick skin type classification system, with each image labeled as "Lighter'' or "Darker''.
#
# Let's begin by importing these datasets:
# + colab={} colab_type="code" id="RWXaaIWy6jVw"
# Get the training data: both images from CelebA and ImageNet
#path_to_training_data = tf.keras.utils.get_file('train_face.h5', 'https://www.dropbox.com/s/l5iqduhe0gwxumq/train_face.h5?dl=1')
path_to_training_data = './train_face.h5'
# + [markdown] colab_type="text" id="bAY6pDc_Zljt"
# This directly downloads the raw data. We've written two classes that do a bit of data pre-processing and import the results in a usable format: `TrainingDatasetLoader` for the training data and `PPBFaceEvaluator` for the test data.
#
# Let's create a `TrainingDatasetLoader` and use it to take a look at the training data.
# + colab={"base_uri": "https://localhost:8080/", "height": 52} colab_type="code" id="UX-eUcEoazBm" outputId="f91666a3-e67e-4213-e313-e339b6cf4a1f"
# Instantiate a TrainingDatasetLoader using the downloaded dataset
loader = util.TrainingDatasetLoader(path_to_training_data)
# + [markdown] colab_type="text" id="yIE321rxa_b3"
# We can look at the size of the training dataset and grab a batch of size 100:
# + colab={} colab_type="code" id="DjPSjZZ_bGqe"
number_of_training_examples = loader.get_train_size()
(images, labels) = loader.get_batch(100)
# + [markdown] colab_type="text" id="sxtkJoqF6oH1"
# Play around with displaying images to get a sense of what the training data actually looks like!
# + colab={"base_uri": "https://localhost:8080/", "height": 165} colab_type="code" id="Jg17jzwtbxDA" outputId="1e0c923e-f1e5-4ac9-b506-064a0d11568e"
#@title Change the sliders to look at positive and negative training examples! { run: "auto" }
face_images = images[np.where(labels==1)[0]]
not_face_images = images[np.where(labels==0)[0]]
idx_face = 20 #@param {type:"slider", min:0, max:50, step:1}
idx_not_face = 11 #@param {type:"slider", min:0, max:50, step:1}
plt.figure(figsize=(4,2))
plt.subplot(1, 2, 1)
plt.imshow(face_images[idx_face])
plt.title("Face")
plt.grid(False)
plt.subplot(1, 2, 2)
plt.imshow(not_face_images[idx_not_face])
plt.title("Not Face")
plt.grid(False)
# + [markdown] colab_type="text" id="kjuaEdLzb4pP"
# We can also create a `PPBFaceEvaluator` instance for the PPB dataset and display some example images. We'll use this dataset later on in the evaluation step.
# + colab={"base_uri": "https://localhost:8080/", "height": 351} colab_type="code" id="4B4egQZY6wEt" outputId="5d322438-3add-4333-e05d-5384c8d46700"
#@title { run: "auto" }
import util
ppb = util.PPBFaceEvaluator(skip=4) # create the dataset handler
gender = "male" #@param ["male", "female"]
skin_color = "darker" #@param ["lighter", "darker"]
img = ppb.get_sample_faces_from_demographic(gender, skin_color)
plt.imshow(img)
plt.grid(False)
# + [markdown] colab_type="text" id="NDj7KBaW8Asz"
# ### Thinking about bias
#
# Remember we'll be training our facial detection classifiers on the large, well-curated CelebA dataset (and ImageNet), and then evaluating their accuracy by testing them on the PPB dataset. Our goal is to build a model that trains on CelebA *and* achieves high classification accuracy on PPB across all demographics, and to thus show that this model does not suffer from any hidden bias.
#
# What exactly do we mean when we say a classifier is biased? In order to formalize this, we'll need to think about [*latent variables*](https://en.wikipedia.org/wiki/Latent_variable), variables that define a dataset but are not strictly observed. As defined in the generative modeling lecture, we'll use the term *latent space* to refer to the probability distributions of the aforementioned latent variables. Putting these ideas together, we consider a classifier *biased* if its classification decision changes after it sees some additional latent features. This notion of bias may be helpful to keep in mind throughout the rest of the lab.
# + [markdown] colab_type="text" id="AIFDvU4w8OIH"
# ## 2.2 CNN for facial detection
#
# First, we'll define and train a CNN on the facial classification task, and evaluate its accuracy on the PPB dataset. Later, we'll evaluate the performance of our debiased models against this baseline CNN. The CNN model has a relatively standard architecture consisting of a series of convolutional layers with batch normalization followed by two fully connected layers to flatten the convolution output and generate a class prediction.
#
# ### Define and train the CNN model
#
# Like we did in the first part of the lab, we'll define our CNN model, and then train on the CelebA and ImageNet datasets using the `tf.GradientTape` class and the `tf.GradientTape.gradient` method.
# + colab={} colab_type="code" id="82EVTAAW7B_X"
n_outputs = 1 # number of outputs (i.e., face or not face)
n_filters = 12 # base number of convolutional filters
'''Function to define a standard CNN model'''
def make_standard_classifier():
Conv2D = functools.partial(tf.keras.layers.Conv2D, padding='same', activation='relu')
BatchNormalization = tf.keras.layers.BatchNormalization
Flatten = tf.keras.layers.Flatten
Dense = functools.partial(tf.keras.layers.Dense, activation='relu')
model = tf.keras.Sequential([
Conv2D(filters=1*n_filters, kernel_size=[5,5], strides=[2,2], input_shape=(64,64,3)),
BatchNormalization(),
Conv2D(filters=2*n_filters, kernel_size=[5,5], strides=[2,2]),
BatchNormalization(),
Conv2D(filters=4*n_filters, kernel_size=[3,3], strides=[2,2]),
BatchNormalization(),
Conv2D(filters=6*n_filters, kernel_size=[3,3], strides=[1,1]),
BatchNormalization(),
Flatten(),
Dense(1, activation=None),
tf.keras.layers.Dropout(0.5)
])
return model
standard_classifier = make_standard_classifier()
# + [markdown] colab_type="text" id="c-eWf3l_lCri"
# Now let's train the standard CNN!
# + colab={"base_uri": "https://localhost:8080/", "height": 695} colab_type="code" id="jmOBzRgplB-n" outputId="f024e7aa-7e90-480a-8266-49c426875acc"
batch_size = 36
num_epochs = 1 # keep small to run faster
learning_rate = 1e-3
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate) # define our optimizer
#loss_history = util.LossHistory(smoothing_factor=0.99) # to record the evolution of the loss
#plotter = util.PeriodicPlotter(sec=2, scale='semilogy')
# The training loop!
for epoch in range(num_epochs):
#custom_msg = util.custom_progress_text("Epoch: %(epoch).0f Loss: %(loss)2.2f")
#bar = util.create_progress_bar(custom_msg)
#for idx in bar(range(loader.get_train_size()//batch_size)):
for idx in range(loader.get_train_size()//batch_size):
# First grab a batch of training data and convert the input images to tensors
x, y = loader.get_batch(batch_size)
x = tf.convert_to_tensor(x, dtype=tf.float32)
y = tf.convert_to_tensor(y, dtype=tf.float32)
# GradientTape to record differentiation operations
with tf.GradientTape() as tape:
logits = standard_classifier(x) # feed the images into the model
loss_value = tf.nn.sigmoid_cross_entropy_with_logits(labels=y, logits=logits) # compute the loss
#custom_msg.update_mapping(epoch=epoch, loss=loss_value.numpy().mean())
# Backpropagation
grads = tape.gradient(loss_value, standard_classifier.variables)
optimizer.apply_gradients(zip(grads, standard_classifier.variables), global_step=tf.train.get_or_create_global_step())
print(".", end="")
#print(loss_value.numpy().mean())
#loss_history.append(loss_value.numpy().mean())
#plotter.plot(loss_history.get())
# + [markdown] colab_type="text" id="AKMdWVHeCxj8"
# ### Evaluate performance of the standard CNN
#
# Next, let's evaluate the classification performance of our CelebA-trained standard CNN on the training dataset and the PPB dataset. For the PPB data, we'll look at the classification accuracy across four different demographics defined in PPB: dark-skinned male, dark-skinned female, light-skinned male, and light-skinned female.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="35-PDgjdWk6_" outputId="1e93ef65-e63a-43b6-ebbf-391c711d7635"
# Evaluate on a subset of CelebA+Imagenet
(batch_x, batch_y) = loader.get_batch(5000)
y_pred_standard = tf.round(tf.nn.sigmoid(standard_classifier.predict(batch_x)))
acc_standard = tf.reduce_mean(tf.cast(tf.equal(batch_y, y_pred_standard), tf.float32))
print "Standard CNN accuracy on (potentially biased) training set: {:.4f}".format(acc_standard.numpy())
# + colab={"base_uri": "https://localhost:8080/", "height": 138} colab_type="code" id="vfDD8ztGWk6x" outputId="e23992a8-6a9d-4222-9cf3-802d8c4b2ff2"
# Evaluate on PPB dataset (takes ~3 minutes)
standard_cnn_accuracy = []
for skin_color in ['lighter', 'darker']:
for gender in ['male', 'female']:
standard_cnn_accuracy.append( ppb.evaluate([standard_classifier], gender, skin_color, from_logit=True)[0] )
print
print "{} {}: {}".format(gender, skin_color, standard_cnn_accuracy[-1])
# + colab={} colab_type="code" id="SaPPGYdPmcCi"
plt.bar(range(4), standard_cnn_accuracy)
plt.xticks(range(4), ('LM', 'LF', 'DM', 'DF'))
plt.ylim(np.min(standard_cnn_accuracy)-0.1,np.max(standard_cnn_accuracy)+0.1)
plt.ylabel('Accuracy')
# + [markdown] colab_type="text" id="j0Cvvt90DoAm"
# Take a look at the accuracies for this first model across these four groups. What do you observe? Would you consider this model biased or unbiased, and why?
# + [markdown] colab_type="text" id="nLemS7dqECsI"
# ## 2.3 Variational autoencoder (VAE) for learning latent structure
#
# As you saw, the accuracy of the CNN varies across the four demographics we looked at. To think about why this may be, consider the dataset the model was trained on, CelebA. If certain features, such as dark skin or hats, are *rare* in CelebA, the model may end up biased against these as a result of training with a biased dataset. That is to say, its classification accuracy will be worse on faces that have under-represented features, such as dark-skinned faces or faces with hats, relevative to faces have features that are well-represented in the training data! This is a problem.
#
# Our goal is to train a *debiased* version of this classifier -- one that accounts for potential disparities in feature representation within the training data. Specifically, to build a debiased facial classifier, we'll train a model that learns a representation of the underlying latent space to the face training data. The model then uses this information to mitigate unwanted biases by sampling faces with rare features, like dark skin or hats, *more frequently* during training. The key design requirement for our model is that it can learn an *encoding* of the latent features in the face data in an entirely *unsupervised* way. To achieve this, we'll turn to variational autoencoders (VAEs).
#
# 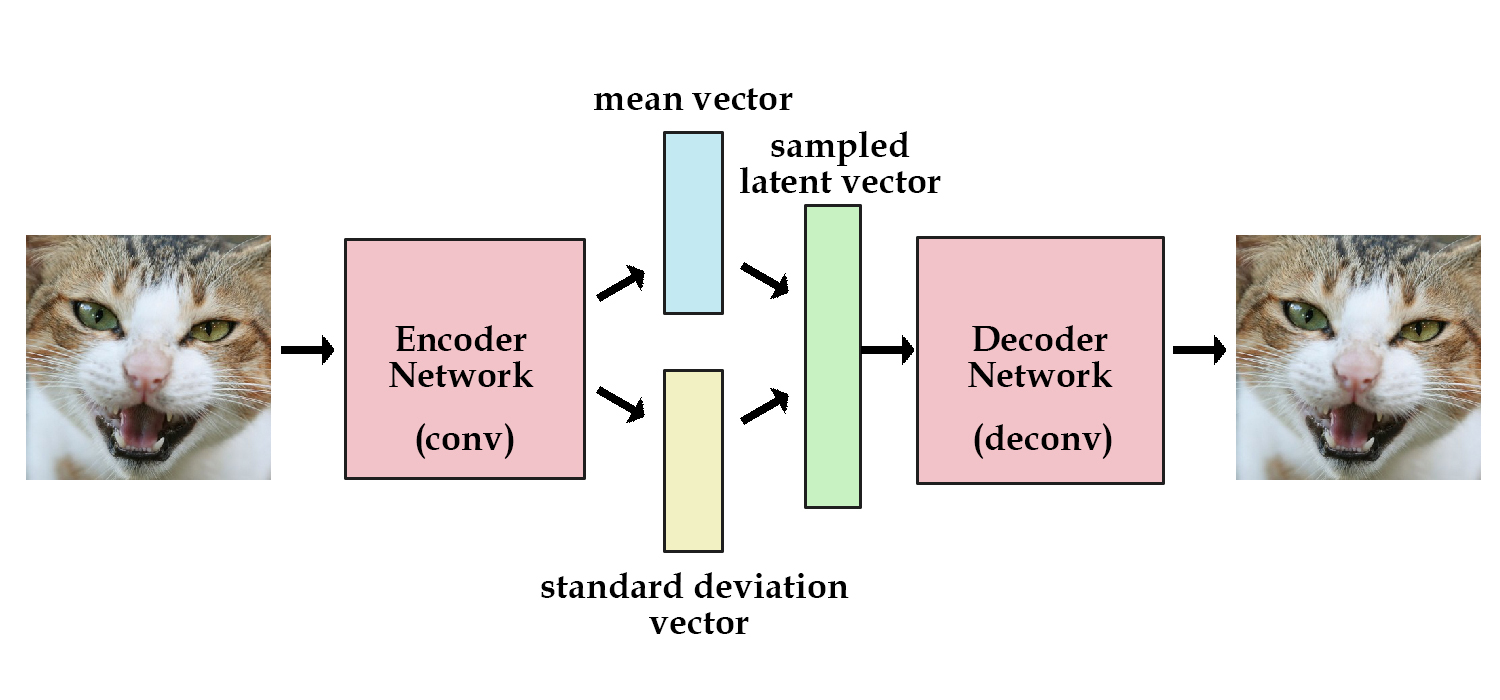
#
# As shown in the schematic above, VAEs rely on an encoder-decoder structure to learn a latent representation of the input data. In the context of computer vision, the encoder network takes in input images, encodes them into a series of variables defined by a mean and standard deviation, and then draws from the distributions defined by these parameters to generate a set of sampled latent variables. The decoder network then "decodes" these variables to generate a reconstruction of the original image, which is used during training to help the model identify which latent variables are important to learn.
#
# Let's formalize two key aspects of the VAE model and define relevant functions for each.
#
# + [markdown] colab_type="text" id="KmbXKtcPkTXA"
# ### Understanding VAEs: loss function
#
# In practice, how can we train a VAE? In doing the reparameterization above, we constrain the means and standard deviations to approximately follow a unit Gaussian. Recall that these are learned parameters, and therefore must factor into the loss computation, and that the decoder portion of the VAE is using these parameters to output a reconstruction that should closely match the input image, which also must factor into the loss. What this means is that we'll have two terms in our VAE loss function:
#
# 1. **Latent loss ($L_{KL}$)**: measures how closely the learned latent variables match a unit Gaussian and is defined by the Kullback-Leibler (KL) divergence. Note that the reparameterization trick is what makes this loss function differentiable!
# 2. **Reconstruction loss ($L_{x}{(x,\hat{x})}$)**: measures how accurately the reconstructed outputs match the input and is given by the $L^2$ norm of the input image and its reconstructed output.
#
# The equations for both of these losses are provided below:
#
# $$ L_{KL}(\mu, \sigma) = \frac{1}{2}\sum\limits_{j=0}^{k-1}\small{(\sigma_j + \mu_j^2 - 1 - \log{\sigma_j})} $$
#
# $$ L_{x}{(x,\hat{x})} = ||x-\hat{x}||_2 $$
#
# Thus for the VAE loss we have:
#
# $$ L_{VAE} = c\cdot L_{KL} + L_{x}{(x,\hat{x})} $$
#
# where $c$ is a weighting coefficient used for regularization.
#
# Now we're ready to define our VAE loss function:
# + colab={} colab_type="code" id="S00ASo1ImSuh"
# Function to calculate VAE loss given an input x, reconstructed output x_pred,
# encoded means mu, encoded log of standard deviation logsigma, and weight parameter for the latent loss
def vae_loss_function(x, x_pred, mu, logsigma, kl_weight=0.0005):
'''TODO: Define the latent loss'''
latent_loss = 0.5 * tf.reduce_sum(tf.exp(logsigma) + tf.square(mu) - 1.0 - logsigma, axis=1) # TODO
'''TODO: Define the reconstruction loss. Hint: you'll need to use tf.reduce_mean'''
reconstruction_loss = tf.reduce_mean((x-x_pred)**2, axis=(1,2,3)) # TODO
'''TODO: Define the VAE loss'''
vae_loss = kl_weight * latent_loss + reconstruction_loss
return vae_loss
# + [markdown] colab_type="text" id="E8mpb3pJorpu"
# Great! Now that we have a more concrete sense of how VAEs work, let's explore how we can leverage this network structure to train a *debiased* facial classifier.
# + [markdown] colab_type="text" id="DqtQH4S5fO8F"
# ### Understanding VAEs: reparameterization
#
# As you may recall from lecture, VAEs use a "reparameterization trick" for sampling learned latent variables. Instead of the VAE encoder generating a single vector of real numbers for each latent variable, it generates a vector of means and a vector of standard deviations that are constrained to roughly follow Gaussian distributions. We then sample from the standard deviations and add back the mean to output this as our sampled latent vector. Formalizing this for a latent variable $z$ where we sample $\epsilon \sim \mathcal{N}(0,(I))$ we have:
#
# $$ z = \mathbb{\mu} + e^{\left(\frac{1}{2} \cdot \log{\Sigma}\right)}\circ \epsilon $$
#
# where $\mu$ is the mean and $\Sigma$ is the covariance matrix. This is useful because it will let us neatly define the loss function for the VAE, generate randomly sampled latent variables, achieve improved network generalization, **and** make our complete VAE network differentiable so that it can be trained via backpropagation. Quite powerful!
#
# Let's define a function to implement the VAE sampling operation:
# + colab={} colab_type="code" id="cT6PGdNajl3K"
"""Reparameterization trick by sampling from an isotropic unit Gaussian.
# Arguments
args (tensor): mean and log of standard deviation of latent distribution (Q(z|X))
# Returns
z (tensor): sampled latent vector
"""
def sampling(args):
z_mean, z_logsigma = args
batch = z_mean.shape[0]
dim = z_mean.shape[1]
# by default, random_normal has mean=0 and std=1.0
epsilon = tf.random_normal(shape=(batch, dim))
'''TODO: Define the reparameterization computation!'''
return z_mean + tf.math.exp(0.5 * z_logsigma) * epsilon # TODO
# + [markdown] colab_type="text" id="qtHEYI9KNn0A"
# ## 2.4 Debiasing variational autoencoder (DB-VAE) for facial detection
#
# Now, we'll use the general idea behind the VAE architecture to build a model, termed a *debiasing variational autoencoder* or DB-VAE, to mitigate (potentially) unknown biases present within the training idea. We'll train our DB-VAE model on the facial detection task, run the debiasing operation during training, evaluate on the PPB dataset, and compare its accuracy to our original, biased CNN model.
#
# ### The DB-VAE model
#
# The key idea behind this debiasing approach is to use the latent variables learned via a VAE to adaptively re-sample the CelebA data during training. Specifically, we will alter the probability that a given image is used during training based on how often its latent features appear in the dataset. So, faces with rarer features (like dark skin, sunglasses, or hats) should become more likely to be sampled during training, while the sampling probability for faces with features that are over-represented in the training dataset should decrease (relative to uniform random sampling across the training data).
#
# A general schematic of the DB-VAE approach is shown here:
#
# 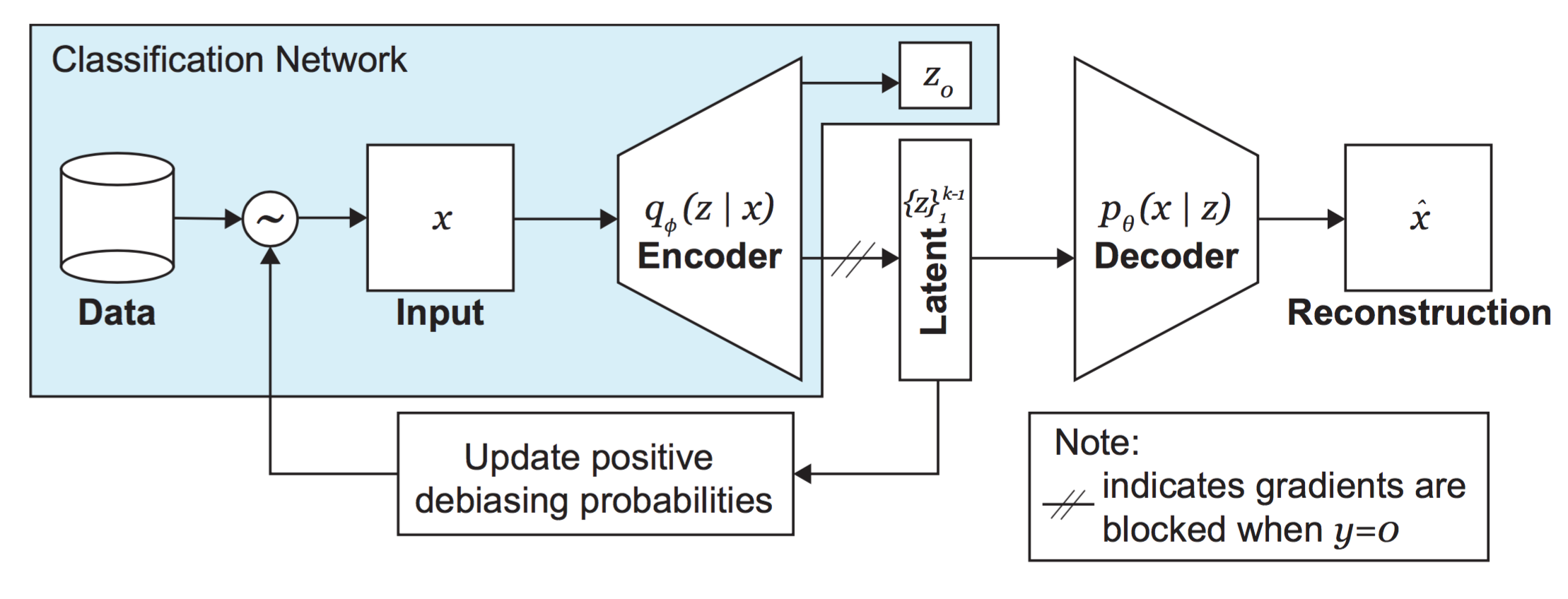
# + [markdown] colab_type="text" id="ziA75SN-UxxO"
# Recall that we want to apply our DB-VAE to a *supervised classification* problem -- the facial detection task. Importantly, note how the encoder portion in the DB-VAE architecture also outputs a single supervised variable, $z_o$, corresponding to the class prediction -- face or not face. Usually, VAEs are not trained to output any supervised variables (such as a class prediction)! This is another key distinction between the DB-VAE and a traditional VAE.
#
# Keep in mind that we only want to learn the latent representation of *faces*, as that's what we're ultimately debiasing against, even though we are training a model on a binary classification problem. We'll need to ensure that, **for faces**, our DB-VAE model both learns a representation of the unsupervised latent variables, captured by the distribution $q_\phi(z|x)$, **and** outputs a supervised class prediction $z_o$, but that, **for negative examples**, it only outputs a class prediction $z_o$.
# + [markdown] colab_type="text" id="XggIKYPRtOZR"
# ### Defining the DB-VAE loss function
#
# This means we'll need to be a bit clever about the loss function for the DB-VAE. The form of the loss will depend on whether it's a face image or a non-face image that's being considered.
#
# For **face images**, our loss function will have two components:
#
#
# 1. **VAE loss ($L_{VAE}$)**: consists of the latent loss and the reconstruction loss.
# 2. **Classification loss ($L_y(y,\hat{y})$)**: standard cross-entropy loss for a binary classification problem.
#
# In contrast, for images of non-faces, our loss function is solely the classification loss.
#
# We can write a single expression for the loss by defining an indicator variable $\mathcal{I}_f$which reflects which training data are images of faces ($\mathcal{I}_f(x) = 1$ ) and which are images of non-faces ($\mathcal{I}_f(x) = 0$). Using this, we obtain:
#
# $$L_{total} = L_y(y,\hat{y}) + \mathcal{I}_f(x)\Big[L_{VAE}\Big]$$
#
# Let's write a function to define the DB-VAE loss function:
#
#
# + colab={} colab_type="code" id="VjieDs8Ovcqs"
# Loss function for DB-VAE
def debiasing_loss_function(x, x_pred, y, y_logit, mu, logsigma):
'''TODO: call the relevant function to obtain VAE loss'''
vae_loss = vae_loss_function(x, x_pred, mu, logsigma) # TODO
'''TODO: define the classification loss'''
classification_loss = tf.nn.sigmoid_cross_entropy_with_logits(labels=y, logits=y_logit) # TODO
# Use the training data labels to create variable face_mask
face_mask = tf.cast(tf.equal(y, 1), tf.float32)
'''TODO: define the DB-VAE total loss! Hint: think about the dimensionality of your output.'''
total_loss = tf.reduce_mean(
classification_loss +
face_mask * vae_loss
)
return total_loss, classification_loss
# + [markdown] colab_type="text" id="YIu_2LzNWwWY"
# ### DB-VAE architecture
#
# Now we're ready to define the DB-VAE architecture. First, let's define some key parameters for our model: the number of latent variables, the number of supervised outputs, and the starting number of filters for the first convolutional layer in the encoder.
# + colab={} colab_type="code" id="ds7o8AuFxUpg"
latent_dim = 100
# + [markdown] colab_type="text" id="amYEnHdJxWYB"
# To build the DB-VAE, we'll define each of the encoder and decoder networks separately, create and initialize the two models, and then construct the end-to-end VAE. We'll go through each of these steps in turn.
# + colab={} colab_type="code" id="4k0tQeW1xpJf"
'''Define the encoder network for the DB-VAE'''
def make_face_encoder_network():
Conv2D = functools.partial(tf.keras.layers.Conv2D, padding='same', activation='relu')
BatchNormalization = tf.keras.layers.BatchNormalization
Flatten = tf.keras.layers.Flatten
Dense = functools.partial(tf.keras.layers.Dense, activation='relu')
inputs = tf.keras.layers.Input(shape=(64,64,3))
hidden = Conv2D(filters=1*n_filters, kernel_size=[5,5], strides=[2,2])(inputs)
hidden = BatchNormalization()(hidden)
hidden = Conv2D(filters=2*n_filters, kernel_size=[5,5], strides=[2,2])(hidden)
hidden = BatchNormalization()(hidden)
hidden = Conv2D(filters=4*n_filters, kernel_size=[3,3], strides=[2,2])(hidden)
hidden = BatchNormalization()(hidden)
hidden = Conv2D(filters=6*n_filters, kernel_size=[3,3], strides=[1,1])(hidden)
hidden = BatchNormalization()(hidden)
hidden = Flatten(name='flatten')(hidden)
# hidden = Dense(128)(hidden)
'''Encoder outputs:
y_logit: supervised class prediction
z_mean: means in the latent space
z_logsigma: standard deviations in the latent space'''
y_logit = Dense(1, activation=None, name='y_logit')(hidden)
z_mean = Dense(latent_dim, name='z_mean')(hidden)
z_logsigma = Dense(latent_dim, name='z_logsigma')(hidden)
# use reparameterization trick to sample from the latent space
z = tf.keras.layers.Lambda(sampling, output_shape=(latent_dim,))([z_mean, z_logsigma])
# define the outputs that the encoder model should return
outputs = [y_logit, z_mean, z_logsigma, z]
# finalize the encoder model
encoder = tf.keras.Model(inputs=inputs, outputs=outputs, name='encoder')
# get the shape of the final convolutional output (right before the flatten)
flatten_layer_idx = encoder.layers.index(encoder.get_layer('flatten'))
pre_flatten_shape = encoder.layers[flatten_layer_idx-1].get_output_at(0).shape[1:]
return encoder, inputs, outputs, pre_flatten_shape
# + [markdown] colab_type="text" id="FlB-Gcot0gYw"
# Similarly, we can define the decoder network, which takes as input the sampled latent variables, runs them through a series of deconvolutional layers, and outputs a reconstruction of the original input image:
# + colab={} colab_type="code" id="JfWPHGrmyE7R"
'''Define the decoder network for the DB-VAE'''
def make_face_decoder_network(pre_flatten_shape):
Conv2DTranspose = functools.partial(tf.keras.layers.Conv2DTranspose, padding='same', activation='relu')
BatchNormalization = tf.keras.layers.BatchNormalization
Flatten = tf.keras.layers.Flatten
Dense = functools.partial(tf.keras.layers.Dense, activation='relu')
latent_inputs = tf.keras.layers.Input(shape=(latent_dim,))
# hidden = Dense(128)(latent_inputs)
hidden = Dense(tf.reduce_prod(pre_flatten_shape))(latent_inputs)
hidden = tf.keras.layers.Reshape(pre_flatten_shape)(hidden)
# series of deconvolutional layers with batch normalization
hidden = Conv2DTranspose(filters=4*n_filters, kernel_size=[3,3], strides=[1,1])(hidden)
hidden = BatchNormalization()(hidden)
hidden = Conv2DTranspose(filters=2*n_filters, kernel_size=[3,3], strides=[2,2])(hidden)
hidden = BatchNormalization()(hidden)
hidden = Conv2DTranspose(filters=1*n_filters, kernel_size=[5,5], strides=[2,2])(hidden)
hidden = BatchNormalization()(hidden)
x_hat = Conv2DTranspose(filters=3, kernel_size=[5,5], strides=[2,2])(hidden)
# instantiate decoder model
decoder = tf.keras.Model(inputs=latent_inputs, outputs=x_hat, name='decoder')
return decoder
# + [markdown] colab_type="text" id="yWCMu12w1BuD"
# Now, call these functions to create the encoder and decoder!
# + colab={} colab_type="code" id="dSFDcFBL13c3"
'''TODO: create the encoder and decoder networks'''
encoder, inputs, ouputs, pre_flatten_shape = make_face_encoder_network() # TODO
decoder = make_face_decoder_network(pre_flatten_shape) # TODO
# initialize the models
encoder_output = encoder(inputs)
y_logit, z_mean, z_logsigma, z = encoder_output
reconstructed_inputs = decoder(z)
# + [markdown] colab_type="text" id="QbRI5_rz2Myy"
# Finally we can construct our network end-to-end.
# + colab={} colab_type="code" id="WITL88Fm2Z0a"
# Construct the end to end vae
vae = tf.keras.Model(inputs, reconstructed_inputs)
# + [markdown] colab_type="text" id="DjdyDDpc01ZZ"
# Let's visualize the architecture of the encoder to get a more concrete understanding of this network,
# + colab={} colab_type="code" id="7yKMwQU606ZR"
util.display_model(encoder)
# + [markdown] colab_type="text" id="M-clbYAj2waY"
# As you can see, the encoder architecture is virtually identical to the CNN from earlier in this lab. Note the outputs of this model: `y_logit, z_mean, z_logsigma, z`. Think carefully about why each of these are outputted and their significance to the problem at hand.
#
#
# + [markdown] colab_type="text" id="nbDNlslgQc5A"
# ### Adaptive resampling for automated debiasing with DB-VAE
#
# So, how can we actually use DB-VAE to train a debiased facial detection classifier? Recall the DB-VAE architecture. As the input images are fed through the network, the encoder learns an estimate $\mathcal{Q}(z|X)$ of the latent space. We want to increase the relative frequency of rare data by increased sampling of under-represented regions of the latent space. We can approximate $\mathcal{Q}(z|X)$ using the frequency distributions of each of the learned latent variables, and then define the probability distribution of selecting a given datapoint $x$ based on this approximation. These probability distributions will be used during training to re-sample the data.
#
# You'll write a function to execute this update of the sampling probabilities, and then call this function within the DB-VAE training loop to actually debias the model.
# + [markdown] colab_type="text" id="Fej5FDu37cf7"
# First, we've defined a short helper function `get_latent_mu` that returns the latent variable means returned by the encoder after a batch of images is inputted to the network:
# + colab={} colab_type="code" id="ewWbf7TE7wVc"
# Function to return the means for an input image batch
def get_latent_mu(images, encoder, batch_size=1024):
N = images.shape[0]
mu = np.zeros((N, latent_dim))
for start_ind in xrange(0, N, batch_size):
end_ind = min(start_ind+batch_size, N+1)
batch = images[start_ind:end_ind]
batch = tf.convert_to_tensor(batch, dtype=tf.float32)/255.
_, batch_mu, _, _ = encoder(batch)
mu[start_ind:end_ind] = batch_mu
return mu
# + [markdown] colab_type="text" id="wn4yK3SC72bo"
# Now, let's define the actual resampling algorithm `get_training_sample_probabilities`. Importantly note the argument `smoothing_fac`. This parameter tunes the degree of debiasing: for `smoothing_fac=0`, the re-sampled training set will tend towards falling uniformly over the latent space.
# + colab={} colab_type="code" id="HiX9pmmC7_wn"
'''Function that recomputes the sampling probabilities for images within a batch
based on how they distribute across the '''
def get_training_sample_probabilities(images, encoder, bins=10, smoothing_fac=0.0):
print "Recomputing the sampling probabilities"
mu = get_latent_mu(images, encoder)
# sampling probabilities for the images
training_sample_p = np.zeros(mu.shape[0])
# consider the distribution for each latent variable
for i in range(latent_dim):
latent_distribution = mu[:,i]
# generate a histogram of the latent distribution
hist_density, bin_edges = np.histogram(latent_distribution, density=True, bins=bins)
# find which latent bin every data sample falls in
# https://docs.scipy.org/doc/numpy-1.13.0/reference/generated/numpy.digitize.html
bin_edges[0] = -float('inf')
bin_edges[-1] = float('inf')
'''TODO: call the digitize function to find which bins in the latent distribution
every data sample falls in to'''
bin_idx = np.digitize(latent_distribution, bin_edges) # TODO
# smooth the density function [Eq. #]
hist_smoothed_density = hist_density + smoothing_fac
hist_smoothed_density = hist_smoothed_density / np.sum(hist_smoothed_density)
'''TODO: invert the density function to compute the sampling probability!
HINT: think carefully about the indexing of the bins! What is the length of bin_edges?'''
p = 1.0/(hist_smoothed_density[bin_idx-1]) # TODO
# normalize all probabilities
p = p / np.sum(p)
# update sampling probabilities
training_sample_p = np.maximum(p, training_sample_p)
# final normalization
training_sample_p /= np.sum(training_sample_p)
return training_sample_p
# + [markdown] colab_type="text" id="pF14fQkVUs-a"
# Now that we've defined the resampling update, we can train our DB-VAE model on the CelebA/ImageNet training data, and run the above operation to re-weight the importance of particular data points as we train the model. Remember again that we only want to debias for features relevant to *faces*, not the set of negative examples.
#
# Complete the code block below to execute the training loop!
# + colab={} colab_type="code" id="9YR8U43FVZ_8"
loss_history = []
optimizer = tf.train.AdamOptimizer(learning_rate=learning_rate)
enable_debiasing = True
all_faces = loader.get_all_train_faces() # parameter from data loader
for epoch in range(num_epochs):
# progress message and bar
custom_msg = util.custom_progress_text("Epoch: %(epoch).0f Iter: %(idx).0f Class Loss: %(class_loss)2.2f Loss: %(loss)2.2f")
bar = util.create_progress_bar(custom_msg)
p_faces = None
if enable_debiasing:
# Recompute data sampling proabilities if debiasing is enabled
'''TODO: write the function call to recompute the sampling probabilities
when debiasing is enabled'''
p_faces = get_training_sample_probabilities(all_faces, encoder) # TODO
for idx in bar(range(loader.get_train_size()//batch_size)):
# load a batch of data
(x, y) = loader.get_batch(batch_size, p_pos=p_faces)
x = tf.convert_to_tensor(x, dtype=tf.float32)
y = tf.convert_to_tensor(y, dtype=tf.float32)
# define GradientTape for automatic differentiation
with tf.GradientTape() as tape:
y_logit, mu, logsigma, z = encoder(x)
x_hat = decoder(z)
'''TODO: call the relevant loss function to compute the loss'''
loss, class_loss = debiasing_loss_function(x, x_hat, y, y_logit, mu, logsigma) # TODO
'''TODO: use the GradientTape.gradient method to compute the gradients'''
grads = tape.gradient(loss, vae.variables) # TODO
# apply gradients to variables
optimizer.apply_gradients(zip(grads, vae.variables),
global_step=tf.train.get_or_create_global_step())
# track the losses
class_loss_value = class_loss.numpy().mean()
loss_value = loss.numpy().mean()
loss_history.append((class_loss_value, loss_value))
custom_msg.update_mapping(epoch=epoch, idx=idx, loss=loss_value, class_loss=class_loss_value)
# plot the progress every 100 steps
if idx%100 == 0:
util.plot_sample(x,y,vae)
# + [markdown] colab_type="text" id="uZBlWDPOVcHg"
# Wonderful! Now we should have a trained and (hopefully!) debiased facial classification model, ready for evaluation!
# + [markdown] colab_type="text" id="Eo34xC7MbaiQ"
# ## 2.4 Evaluation on Pilot Parliaments Benchmark (PPB) Dataset
#
# Finally let's test our DB-VAE model on the[ PPB dataset](http://proceedings.mlr.press/v81/buolamwini18a/buolamwini18a.pdf).
#
# We'll evaluate both the overall accuracy of the DB-VAE as well as its accuracy on each the "Dark Male", "Dark Female", "Light Male", and "Light Female" demographics, and compare the performance of this debiased model against the biased CNN from earlier in the lab.
#
# Here are some example images from the PPB dataset.
#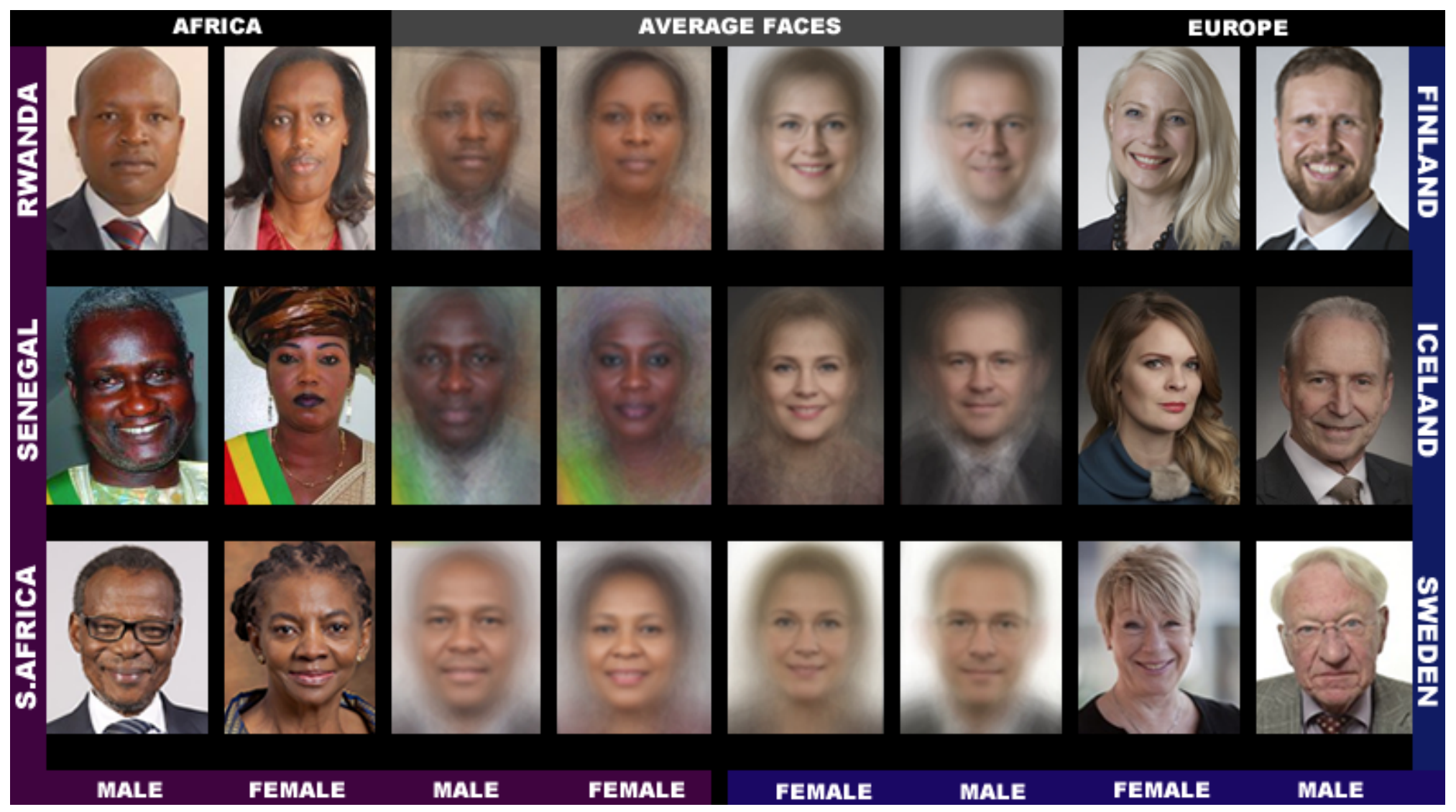 
# + [markdown] colab_type="text" id="ruzxwzo2ko6N"
# To assess performance, we'll measure the classification accuracy of each model, which we define as the fraction of PPB faces detected. By comparing the accuracy of a model without debiasing and our DB-VAE model, we can get a sense of how effectively we were able to debias against features like skin tone and gender.
#
# Let's evaluate our debiased model on the PPB test dataset.
# + colab={} colab_type="code" id="bgK77aB9oDtX"
# Evaluate on PPB dataset (takes ~4 minutes)
accuracy_debiased = []
for skin_color in ['lighter', 'darker']:
for gender in ['male', 'female']:
accuracy_debiased.append( ppb.evaluate([encoder], gender, skin_color, output_idx=0, from_logit=True)[0] )
print
print "{} {}: {}".format(gender, skin_color, accuracy_debiased[-1])
# + [markdown] colab_type="text" id="F-3NzMB0oQtv"
# We can calculate the accuracies of our model on the whole PPB dataset as well as across the four demographics proposed and visualize our results comparing to the standard, biased CNN.
#
# + colab={} colab_type="code" id="zzm-THVJkBjY"
bar_width = 0.3
plt.bar(np.arange(4), standard_cnn_accuracy, width=bar_width)
plt.bar(np.arange(4)+bar_width, accuracy_debiased, width=bar_width)
plt.legend(('Standard Classifier','Debiased Classifier (DB-VAE)'))
plt.xticks(np.arange(4), ('LM', 'LF', 'DM', 'DF'))
plt.ylim(np.min([standard_cnn_accuracy,accuracy_debiased])-0.1,1)
plt.ylabel('Accuracy')
# + [markdown] colab_type="text" id="rESoXRPQo_mq"
# ## 2.5 Conclusion
#
# We encourage you to think about and maybe even address some questions raised by the approach and results outlined here:
#
# * How does the accuracy of the DB-VAE across the four demographics compare to that of the standard CNN? Do you find this result surprising in any way?
# * In which applications (either related to facial detection or not!) would debiasing in this way be desired? Are there applications where you may not want to debias your model?
# * Do you think it should be necessary for companies to demonstrate that their models, particularly in the context of tasks like facial detection, are not biased? If so, do you have thoughts on how this could be standardized and implemented?
# * Do you have ideas for other ways to address issues of bias, particularly in terms of the training data?
#
# Hopefully this lab has shed some light on a few concepts, from vision based tasks, to VAEs, to algorithmic bias. We like to think it has, but we're biased ;).
#
# 
|
old-notes/old-ai/dl/2-COMPUTER-VISION-TRANSFER-LEARNING/mit-lab/2-debiasing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Make graph
file = "out.ego-twitter"
# ! head out.ego-twitter
FIN = open(file, 'r')
left = set()
right = set()
edges = set()
for line in FIN:
if '%' in line or not line:
continue
u, w = map(int, line.strip().split())
left.add(u)
right.add(w)
edges.add(str(u) + ' ' + str(w))
len(left), len(right), len(edges)
FOUT = open('train.in', 'w')
for line in edges:
FOUT.write(line + '\n')
FOUT.close()
# ! tail train.in
|
datasets/directed/ego-twitter/Make graph.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import seaborn as sns
import pandas as pd
import matplotlib.pyplot as plt
import json
import os
# %matplotlib inline
# %reload_ext autoreload
# -
# # Plot
# +
def load_data(filename):
with open(filename, 'r') as f:
lines = f.readlines()
lines = lines[1:]
timesteps = []
mean_rewards = []
best_mean_rewards = []
for i in range(0, len(lines), 6):
timestep = int(lines[i].strip().split(' ')[-1]) // 10000
mean_reward = float(lines[i + 1].strip().split(' ')[-1])
best_mean_reward = lines[i + 2].strip().split(' ')[-1]
best_mean_reward = -21.0 if best_mean_reward == "-inf" else float(best_mean_reward)
timesteps.append(timestep)
mean_rewards.append(mean_reward)
best_mean_rewards.append(best_mean_reward)
return timesteps, mean_rewards, best_mean_rewards
def plot(data, labels=["best mean reward", "mean reward"]):
timesteps, plot_datas = data[0], data[1:]
#plt.plot(timesteps, mean_rewards)
plt.xlabel('Timestep(*10000)')
plt.ylabel('reward')
for d, l in zip(plot_datas, labels):
plt.plot(timesteps, d, label=l)
plt.legend()
plt.grid()
plt.show()
# -
# # Result of different hyperparameters
# Default parameter result data
data = load_data('pong_default_parameter.txt')
plot(data)
# Double batch size result data
data_bz_128 = load_data('pong_bz_128.txt')
plot(data_bz_128)
# target network update every 1000 iteration
data_target_update_1000 = load_data('pong_target_update_1000.txt')
plot(data_target_update_1000)
# +
#
# -
# Plot different result in the same picture
timestep_len = len(data_bz_128[0])
all_data = (data_bz_128[0], data[1][:timestep_len], data_bz_128[1][:timestep_len], data_target_update_1000[1][:timestep_len])
plot(all_data, labels=["default", "bz_128", "target_update_1000"])
|
hw3/homework3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (TensorFlow 2.3 Python 3.7 CPU Optimized)
# language: python
# name: python3__SAGEMAKER_INTERNAL__arn:aws:sagemaker:us-east-2:429704687514:image/tensorflow-2.3-cpu-py37-ubuntu18.04-v1
# ---
# <!-- # pip install -U jedi==0.17.2 -->
# # SageMaker Hosting を利用して TensorFlow のモデルを SageMaker hosting のいろいろな機能を使ってホスティングする
# * ホスティングするモデルは pretrained モデルである MobileNet (とv2)を題材にする
# * SageMaker Studio の以下環境の使用を前提とする
# * `TensorFlow 2.3 Python 3.7 CPU Optimized`
# * `t3.medium` または `m5.large`
# * 以下コンテンツの 1 と 2 は実行必須だが、あとは実行したいところから実行可能
#
# ## contents
# * [1. (必須)使用するモジュールのインストールと読み込み、定数の設定](#1.-(必須)使用するモジュールのインストールと読み込み、定数の設定)
# * [2. (必須)使用するモデルの動作確認と-S3-への転送](#2.-(必須)使用するモデルの動作確認と-S3-への転送)
# * [3. TensorFlow の SageMaker マネージドコンテナ(TensorFlow Serving)を利用した hosting](#3.-TensorFlow-の-SageMaker-マネージドコンテナ(TensorFlow-Serving)を利用した-hosting)
# * [#3-1. SageMaker SDK の場合の手順概要](#3-1.-SageMaker-SDK-の場合の手順概要)
# * [#3-2. boto3 の場合の手順概要](#3-2.-boto3-の場合の手順概要)
# * [4. 前処理/後処理追加](#4.-前処理/後処理追加)
# * [4-1. SageMaker Python SDK で前処理/後処理を追加してホスティングと推論](#4-1.-SageMaker-Python-SDK-で前処理/後処理を追加してホスティングと推論)
# * [4-2. Boto3 で前処理/後処理を追加してホスティングと推論](#4-2.-Boto3-で前処理/後処理を追加してホスティングと推論)
# * [5. マルチモデルエンドポイント](#5.-マルチモデルエンドポイント)
# * [6. 非同期推論](#6.-非同期推論)
# * [7. オートスケール](#7.-オートスケール)
# * [8. サーバーレス推論](#8.-サーバーレス推論)
# * [9. 独自コンテナイメージの持ち込みを利用した推論](#9.-独自コンテナイメージの持ち込みを利用した推論)
# ## 1. (必須)使用するモジュールのインストールと読み込み、定数の設定
pip install -U matplotlib
import tensorflow as tf, os, tarfile, json, numpy as np, base64, sagemaker, boto3
from sagemaker.tensorflow import TensorFlowModel
from io import BytesIO
from matplotlib import pyplot as plt
from PIL import Image
from time import sleep
from glob import glob
sm_client = boto3.client('sagemaker')
smr_client = boto3.client('sagemaker-runtime')
s3_client = boto3.client('s3')
endpoint_inservice_waiter = sm_client.get_waiter('endpoint_in_service')
sm_role = sagemaker.get_execution_role()
sess = sagemaker.session.Session()
bucket = sess.default_bucket()
print(f'使用するロール : {sm_role}')
print(f'使用するバケット : {bucket}')
# ## 2. (必須)使用するモデルの動作確認と S3 への転送
# 1. tensorflow の pre-trained model である mobilenet を読み込む
# 2. 推論用の画像とラベルをダウンロード
# 3. 推論用の画像を前処理
# 4. モデルを tar.gz に固めて S3 にアップロード(SageMaker Hosting するため)
# 5. 使用する推論用コンテナの URI を取得
model = tf.keras.applications.mobilenet.MobileNet()
model.summary()
# +
WORK_DIR = f'{os.getcwd()}/work/'
os.makedirs(WORK_DIR, exist_ok=True)
# サンプル画像をダウンロード
file = tf.keras.utils.get_file(
f'{WORK_DIR}cat.jpg',
'https://gahag.net/img/201608/11s/gahag-0115329292-1.jpg')
# 分類クラスをダウンロード
labels_path = tf.keras.utils.get_file(
f'{WORK_DIR}/ImageNetLabels.txt',
'https://storage.googleapis.com/download.tensorflow.org/data/ImageNetLabels.txt')
labels = list(np.array(open(labels_path).read().splitlines())[1:])
# -
with open('./code/labels.txt','wt') as f:
for txt in labels:
f.write(txt+'\n')
imsize = Image.open(file).size
# 画像のresizeと前処理結果の確認
x,y = imsize[0]-imsize[1],0
img = Image.open(file).crop((x,y,900,537)).resize((model.input_shape[1],model.input_shape[2]))
img_arr = ((np.array(img)-127.5)/127.5).astype(np.float32).reshape(-1,model.input_shape[1],model.input_shape[2],3)
img
# モデルの動作確認
print(labels[np.argmax(model.predict(img_arr))]) # tabby
# +
# 保存ディレクトリを指定
MODEL_DIR = './mobilenet/0001'
# tar.gz の出力先を指定
TAR_DIR = 'MyModel'
os.makedirs(TAR_DIR, exist_ok=True)
TAR_NAME = os.path.join(TAR_DIR, 'model.tar.gz')
# モデルを SavedModel 形式で保存
model.save(MODEL_DIR)
# tar.gz ファイルを出力
with tarfile.open(TAR_NAME, mode='w:gz') as tar:
tar.add(MODEL_DIR)
# S3 にアップロードして、返り値としてS3のURIを受け取る
model_s3_path = f's3://{bucket}/{TAR_DIR}'
model_s3_uri = sagemaker.s3.S3Uploader.upload(
local_path = TAR_NAME,
desired_s3_uri = model_s3_path
)
print(model_s3_uri)
# +
# Sagemaker SDK でマネージドコンテナの URI を取得
container_image_tf24_uri = sagemaker.image_uris.retrieve(
"tensorflow", # TensorFlow のマネージドコンテナを利用
sagemaker.session.Session().boto_region_name, # ECR のリージョンを指定
version='2.4', # TensorFlow のバージョンを指定
instance_type = 'ml.m5.large', # インスタンスタイプを指定
image_scope = 'inference' # 推論コンテナを指定
)
print(container_image_tf24_uri)
# -
# ## 3. TensorFlow の SageMaker マネージドコンテナ(TensorFlow Serving)を利用した hosting
# * コンテナの詳細は[こちら](https://github.com/aws/sagemaker-tensorflow-serving-container)
# * SageMaker SDK と boto3 を利用した場合それぞれ行う
# * 前提として事前に saved model 形式で保存したモデルを tar.gz に固めて S3 に配置しておく(2 で実施済)
#
# ### 3-1. SageMaker SDK の場合の手順概要
# 1. SavedModel 形式でモデルを保存(済)
# 2. モデルを tar.gz で固める(済)
# 3. S3 にモデルをアップロード(済)
# 4. SageMaker SDK の [TensorFlowModel](https://sagemaker.readthedocs.io/en/stable/frameworks/tensorflow/sagemaker.tensorflow.html?highlight=TensorFlowModel#sagemaker.tensorflow.model.TensorFlowModel) API で S3 に配置したモデルを読み込む
# 5. [deploy](https://sagemaker.readthedocs.io/en/stable/frameworks/tensorflow/sagemaker.tensorflow.html?highlight=TensorFlowModel#sagemaker.tensorflow.model.TensorFlowModel.deploy) メソッドで推論エンドポイントを作成
# 6. 推論実行
# 7. 推論エンドポイントを削除(付随するものも合わせて削除)
MODEL_NAME = 'MyTFModelFromSMSDK'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'Endpoint'
ENDPOINT_NAME = ENDPOINT_CONFIG_NAME
# モデルとコンテナの指定
tf_model = TensorFlowModel(
name = MODEL_NAME,
model_data=model_s3_uri, # モデルの S3 URI
role= sm_role, # 割り当てるロール
image_uri = container_image_tf24_uri, # コンテナイメージの S3 URI
)
# デプロイ(endpoint 生成)
predictor = tf_model.deploy(
endpoint_name=ENDPOINT_NAME, # エンドポイントの名前
initial_instance_count=1, # インスタンス数
instance_type='ml.m5.large', # インスタンスタイプ
)
img = Image.open(file).resize((model.input_shape[1],model.input_shape[2]))
img_arr = ((np.array(img)-127.5)/127.5).astype(np.float32).reshape(-1,model.input_shape[1],model.input_shape[2],3)
result = np.argmax(predictor.predict(img_arr)['predictions'][0])
print(labels[result])
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
# ### 3-2. boto3 の場合の手順概要
# 1. SavedModel 形式でモデルを保存(済)
# 2. モデルを tar.gz で固める(済)
# 3. S3 にモデルをアップロード(済)
# 4. boto3 の [create_model](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/sagemaker.html#SageMaker.Client.create_model) メソッドで SageMaker のサービスに S3 にアップロードしたモデルを登録する
# 5. boto3 の [create_endpoint_config](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/sagemaker.html#SageMaker.Client.create_endpoint_config)で推論エンドポイントの設定を作成する
# 6. boto3 の [create_endpoint](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/sagemaker.html#SageMaker.Client.create_endpoint) で推論エンドポイントを作成する
# 7. boto3 の [invoke_endpoint](https://boto3.amazonaws.com/v1/documentation/api/latest/reference/services/sagemaker-runtime.html#SageMakerRuntime.Client.invoke_endpoint) で推論する
# 8. 推論エンドポイントを削除
MODEL_NAME = 'MyTFModelAddProcessFromBoto3'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'EndpointConfig'
ENDPOINT_NAME = MODEL_NAME + 'Endpoint'
response = sm_client.create_model(
ModelName=MODEL_NAME,
PrimaryContainer={
# SageMaker SDK の時と同じ URI を指定
'Image': container_image_tf24_uri,
# SageMaker SDK の時と同じ URI を指定
'ModelDataUrl': model_s3_uri,
},
# SageMaker SDK の時と同じ role を指定
ExecutionRoleArn=sm_role,
)
response = sm_client.create_endpoint_config(
EndpointConfigName=ENDPOINT_CONFIG_NAME,
ProductionVariants=[
{
'VariantName': 'AllTrafic',
'ModelName': MODEL_NAME,
'InitialInstanceCount': 1,
'InstanceType': 'ml.m5.xlarge',
},
],
)
response = sm_client.create_endpoint(
EndpointName=ENDPOINT_NAME,
EndpointConfigName=ENDPOINT_CONFIG_NAME,
)
endpoint_inservice_waiter.wait(
EndpointName=ENDPOINT_NAME,
WaiterConfig={'Delay': 5,}
)
# リストを文字列にして渡すパターン
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'Body' : str(img_arr.tolist())
}
response = smr_client.invoke_endpoint(**request_args)
predictions = json.loads(response['Body'].read().decode('utf-8'))['predictions'][0]
print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
# jsonにして渡すパターン
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'Body' : json.dumps({"instances": img_arr.tolist()})
}
response = smr_client.invoke_endpoint(**request_args)
predictions = json.loads(response['Body'].read().decode('utf-8'))['predictions'][0]
print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
# リソース削除
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
# ## 4. 前処理/後処理追加
# * リスト形式でデータを作成し(た後で json形式に変換し)て predict を行っていたが、 `inference.py` を使うことで前処理/後処理を endpoint 側で行うことも可能。
# * 重い画像の前処理を潤沢なエンドポイントのコンピューティングリソースで実行することで、呼び出し側 (Lambda など)の頻繁に処理するコンピューティングリソースの負荷を低減できる
# * 呼び出し側が前処理を意識せずに実装できるようになる(呼び出し側はデータサイエンティストの領域に入らずに済み、エンドポイントで実行する前処理までをDSの領域にできる)
# * 以下を例に実装する。
# * 前処理)画像のバイナリデータを base64 エンコーディングしたものを直接送りつけて、 endpoint 側でリストに変換
# * 後処理)softmax の結果から一番可能性の高い値を取得し、そのインデックスからラベルに変換
#
# ### 4-1. SageMaker Python SDK で前処理/後処理を追加してホスティングと推論
# 手順は前後の処理無しの場合と同じで、`TensorFlowModel` APIでモデルを読み込む際、前処理/後処理を記載した `inference.py` とそのディレクトリを指定する
# !pygmentize ./code/inference.py
MODEL_NAME = 'MyTFModelAddProcessFromSMSDK'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'Endpoint'
ENDPOINT_NAME = ENDPOINT_CONFIG_NAME
TAR_DIR = 'MyModelAddProcess'
code_dir = './code'
os.makedirs(TAR_DIR, exist_ok=True)
TAR_NAME = os.path.join(TAR_DIR, 'model.tar.gz')
with tarfile.open(TAR_NAME, mode='w:gz') as tar:
tar.add(MODEL_DIR)
# +
model_add_process_s3_path = f's3://{bucket}/{TAR_DIR}'
model_add_process_s3_uri = sagemaker.s3.S3Uploader.upload(
local_path = TAR_NAME,
desired_s3_uri = model_add_process_s3_path
)
print(model_add_process_s3_uri)
# -
# #### inference.py と必要なファイルの設定
# * entry_point 引数で `inference.py` (名前固定)を指定すると `input_handler` と `output_handler` を推論前後に実行してくれる
# * 必要なモジュール等がある場合は `source_dir` 引数に格納してあるディレクトリを指定すると一緒に読み込むが、 inference.py が `source_dir` のルートに存在する必要がある
# * ホスティング先の展開ディレクトリは `/opt/ml/model/code` になるので、他のファイルを読み込む時は絶対パスで指定するとよい(カレントディレクトリは `/sagemaker` で実行される)
# モデルとコンテナの指定
tf_model = TensorFlowModel(
name = MODEL_NAME,
model_data=model_add_process_s3_uri, # モデルの S3 URI
role= sm_role, # 割り当てるロール
image_uri = container_image_tf24_uri, # コンテナイメージの S3 URI
entry_point = './code/inference.py',
source_dir = './code/'
)
# デプロイ(endpoint 生成)
predictor = tf_model.deploy(
endpoint_name=ENDPOINT_NAME,
initial_instance_count=1, # インスタンス数
instance_type='ml.m5.xlarge', # インスタンスタイプ
)
# 推論
with open('./work/cat.jpg', 'rb') as img:
data = img.read()
bio = BytesIO()
bio.write(data)
b64_data = base64.b64encode(bio.getvalue()).decode('utf-8')
json_b64 = json.dumps({'b64_image':b64_data})
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'Body' : json_b64
}
response = smr_client.invoke_endpoint(**request_args)
print(response['Body'].read().decode('utf-8'))
# すでにあった場合の削除
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
# ### 4-2. Boto3 で前処理/後処理を追加してホスティングと推論
MODEL_NAME = 'MyTFModelAddProcessFromBoto3'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'EndpointConfig'
ENDPOINT_NAME = MODEL_NAME + 'Endpoint'
# #### inference.py 他を model.tar.gz に同包
# boto3 から endpoint を作成する場合は、SageMaker SDK のように `entry_point` や `source_dir` の設定ができないため、 必要なファイルは予め `model.tar.gz` に一緒に入れる必要がある
# (SageMaker SDK の場合は裏側で自動で `inference.py` などを model.tar.gz に再度固めて s3 にアップロードしてくれている)
# model.tar.gz にモデルなどを固める
TAR_DIR = 'MyModelAddProcess'
code_dir = './code'
os.makedirs(TAR_DIR, exist_ok=True)
TAR_NAME = os.path.join(TAR_DIR, 'model.tar.gz')
with tarfile.open(TAR_NAME, mode='w:gz') as tar:
tar.add(MODEL_DIR)
tar.add(code_dir) # inference.py などを同包
# +
model_add_process_s3_path = f's3://{bucket}/{TAR_DIR}'
model_add_process_s3_uri = sagemaker.s3.S3Uploader.upload(
local_path = TAR_NAME,
desired_s3_uri = model_add_process_s3_path
)
print(model_add_process_s3_uri)
# +
response = sm_client.create_model(
ModelName=MODEL_NAME,
PrimaryContainer={
# SageMaker SDK の時と同じ URI を指定
'Image': container_image_tf24_uri,
# SageMaker SDK の時と同じ URI を指定
'ModelDataUrl': model_add_process_s3_uri,
},
# SageMaker SDK の時と同じ role を指定
ExecutionRoleArn=sm_role,
)
response = sm_client.create_endpoint_config(
EndpointConfigName=ENDPOINT_CONFIG_NAME,
ProductionVariants=[
{
'VariantName': 'AllTrafic',
'ModelName': MODEL_NAME,
'InitialInstanceCount': 1,
'InstanceType': 'ml.m5.xlarge',
},
],
)
response = sm_client.create_endpoint(
EndpointName=ENDPOINT_NAME,
EndpointConfigName=ENDPOINT_CONFIG_NAME,
)
endpoint_inservice_waiter.wait(
EndpointName=ENDPOINT_NAME,
WaiterConfig={'Delay': 5,}
)
# -
# 推論
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'Body' : json_b64
}
response = smr_client.invoke_endpoint(**request_args)
print(response['Body'].read().decode('utf-8'))
# 削除
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
# ## 5. マルチモデルエンドポイント
# * 1つの推論インスタンスに複数のモデルをデプロイすることが可能
# * モデルごとにtar.gzにかためて、S3 の指定プレフィックス直下に配置する
# * 以下は boto3 の例。SageMaker SDK でもマルチモデルエンドポイントは可能で詳細は[こちら](https://sagemaker.readthedocs.io/en/stable/frameworks/tensorflow/deploying_tensorflow_serving.html?highlight=multi%20model#deploying-more-than-one-model-to-your-endpoint)
# * エンドポイント作成手順はシングルモデルと変わらないが、それぞれのモデルを {モデル名}.tar.gz に固めた上で同じキープレフィックスに配置し、create_model する際の引数に、tar.gzを配置しているプレフィックス(tar.gzのオブジェクトのURIではない)を指定する
# * 呼び出す(invoke_endpoint)する際にモデルのファイル名を指定する
#
# ### モデル準備と動作確認( mobilenetv2 )
# 新しくmobilenetv2を追加し、mobilenetとmobilenetv2の2モデルを1つのエンドポイントでホスティングする準備
model2 = tf.keras.applications.mobilenet_v2.MobileNetV2()
model2.summary()
# モデルの動作確認(v1との比較)
# mobilenet
prediction = model.predict(img_arr)[0]
print(prediction[np.argmax(prediction)],labels[np.argmax(prediction)])
# mobilenetV2
prediction = model2.predict(img_arr)[0]
print(prediction[np.argmax(prediction)],labels[np.argmax(prediction)])
# +
# MobileNetV2 の準備
# 保存ディレクトリを指定
MODEL2_DIR = './mobilenetv2/0001'
# モデルを SavedModel 形式で保存
model2.save(MODEL2_DIR)
# mobilenetv2.tar.gz の出力先を指定
TAR_DIR = 'MyMultiModel'
os.makedirs(TAR_DIR, exist_ok=True)
TAR_NAME = os.path.join(TAR_DIR, 'mobilenetv2.tar.gz')
# tar.gz ファイルを出力
with tarfile.open(TAR_NAME, mode='w:gz') as tar:
tar.add(MODEL2_DIR, arcname="0001")
# MobileNet の準備
# mobilenet.tar.gz の出力先を指定
TAR_NAME = os.path.join(TAR_DIR, 'mobilenet.tar.gz')
# tar.gz ファイルを出力
with tarfile.open(TAR_NAME, mode='w:gz') as tar:
tar.add(MODEL_DIR, arcname="0001")
# +
# mobilenet と mobilenet v2 をそれぞれ S3 にアップロードする
multi_model_s3_path = f's3://{bucket}/{TAR_DIR}/'
# !aws s3 cp ./{TAR_DIR}/ {multi_model_s3_path} --recursive
# -
MODEL_NAME = 'MyMultiModel'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'EndpointConfig'
ENDPOINT_NAME = MODEL_NAME + 'Endpoint'
# ### モデルの作成~エンドポイント作成
# * シングルモデルのときはtar.gzのパスを指定していたが、マルチモデルのときはモデルを保存しているプレフィックスを指定する
# * 他はシングルモデルと同じ
# +
response = sm_client.create_model(
ModelName=MODEL_NAME,
PrimaryContainer={
'Image': container_image_tf24_uri,
'Mode':'MultiModel',
'ModelDataUrl': multi_model_s3_path, # tar.gz を複数配置している s3 のプレフィックスを指定
},
ExecutionRoleArn=sm_role,
)
response = sm_client.create_endpoint_config(
EndpointConfigName=ENDPOINT_CONFIG_NAME,
ProductionVariants=[
{
'VariantName': 'AllTrafic',
'ModelName': MODEL_NAME,
'InitialInstanceCount': 1,
'InstanceType': 'ml.m5.xlarge',
},
],
)
response = sm_client.create_endpoint(
EndpointName=ENDPOINT_NAME,
EndpointConfigName=ENDPOINT_CONFIG_NAME,
)
endpoint_inservice_waiter.wait(
EndpointName=ENDPOINT_NAME,
WaiterConfig={'Delay': 5,}
)
# -
# ### マルチモデルエンドポイントでの推論
# * `TargetModel` 引数にtar.gzに固めたモデルのファイル名を入れればそのモデルが使用される
# mobilenet推論
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'TargetModel' : 'mobilenet.tar.gz',
'Body' : json.dumps({"instances": img_arr.tolist()})
}
response = smr_client.invoke_endpoint(**request_args)
predictions = json.loads(response['Body'].read().decode('utf-8'))['predictions'][0]
print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
# mobilenetv2推論
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'TargetModel' : 'mobilenetv2.tar.gz',
'Body' : json.dumps({"instances": img_arr.tolist()})
}
response = smr_client.invoke_endpoint(**request_args)
predictions = json.loads(response['Body'].read().decode('utf-8'))['predictions'][0]
print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
# ### モデルの追加
# * 同じプレフィックス下に新しくモデルを追加すれば追加したモデルで推論可能
# * ここでは mobilenetv2 を別名に差し替えて(mobilenetv2**_2**)、追加でアップロードしてそちらも機能することを確認する
#
# 注1)モデルの削除は S3 から削除すればできるが、タイムラグがかなりあるので注意。モデルをホスティングをしているインスタンスからモデルが削除されない限り(コントロールできない領域で、ホスティングしているインスタンスのメモリ/ストレージが不足したときのみ自動で読み込んでいるモデルが削除される)S3から削除したモデルで推論できる。
# 注2)同様にモデルの更新についても、S3に配置したモデルを上書き保存しても古いモデルがうごき続けてしまう可能性がある。[公式のメッセージ](https://docs.aws.amazon.com/sagemaker/latest/dg/add-models-to-endpoint.html)としては「上書き保存はするな」
# !aws s3 cp ./{TAR_DIR}/mobilenetv2.tar.gz {multi_model_s3_path}mobilenetv2_2.tar.gz
# mobilenet推論
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'TargetModel' : 'mobilenetv2_2.tar.gz', # 後から追加したモデル
'Body' : json.dumps({"instances": img_arr.tolist()})
}
response = smr_client.invoke_endpoint(**request_args)
predictions = json.loads(response['Body'].read().decode('utf-8'))['predictions'][0]
print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
# ## 6. 非同期推論
# * 非同期推論は、推論リクエストがないときにインスタンス数を 0 にすることコストを削減する
# * バッチ変換(batch transform)と近いが、バッチ変換はデータが溜まっている前提で一気に動かすのに対して、個別のデータに対して数分オーダで推論結果を求められる場合に適する(非同期推論は推論対象データが数 GB でも返せる)
# * 詳細は[こちら](https://aws.amazon.com/jp/about-aws/whats-new/2021/08/amazon-sagemaker-asynchronous-new-inference-option/)
# * 使い方はリアルタイム推論に近いが、`endpoint_config`で非同期独自の設定をする
MODEL_NAME = 'MyTFModelFromBoto3Async'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'EndpointConfig'
ENDPOINT_NAME = MODEL_NAME + 'Endpoint'
# ### モデルの作成
# リアルタイム推論と同じ
response = sm_client.create_model(
ModelName=MODEL_NAME,
PrimaryContainer={
# SageMaker SDK の時と同じ URI を指定
'Image': container_image_tf24_uri,
# SageMaker SDK の時と同じ URI を指定
'ModelDataUrl': model_s3_uri,
},
# SageMaker SDK の時と同じ role を指定
ExecutionRoleArn=sm_role,
)
# ### 推論エンドポイントの設定
# `AsyncInferenceConfig` という引数で、推論結果を配置するS3の出力先を指定する
response = sm_client.create_endpoint_config(
EndpointConfigName=ENDPOINT_CONFIG_NAME,
ProductionVariants=[
{
'VariantName': 'AllTrafic',
'ModelName': MODEL_NAME,
'InitialInstanceCount': 1,
'InstanceType': 'ml.m5.xlarge',
},
],
AsyncInferenceConfig={
"OutputConfig": {
"S3OutputPath": f"s3://{bucket}/async_inference/output"
},
}
)
# ### 推論エンドポイントの作成
# リアルタイム推論と同じ
# +
response = sm_client.create_endpoint(
EndpointName=ENDPOINT_NAME,
EndpointConfigName=ENDPOINT_CONFIG_NAME,
)
endpoint_inservice_waiter.wait(
EndpointName=ENDPOINT_NAME,
WaiterConfig={'Delay': 5,}
)
# -
# ### 非同期推論実行
# 事前にS3に推論データを配置して、`invoke_endpoint_async`で非同期推論を実行する
json_name = './tabby.json'
with open(json_name,'wt') as f:
f.write(json.dumps({"instances": img_arr.tolist()}))
tabby_s3_uri = sagemaker.s3.S3Uploader.upload(
local_path = json_name,
desired_s3_uri = f"s3://{bucket}/async_inference/input"
)
# %%time
response = smr_client.invoke_endpoint_async(
EndpointName=ENDPOINT_NAME,
InputLocation=tabby_s3_uri,
ContentType='application/json'
)
output_s3_uri = response['OutputLocation']
output_key = output_s3_uri.replace(f's3://{bucket}/','')
while True:
result = s3_client.list_objects(Bucket=bucket, Prefix=output_key)
exists = True if "Contents" in result else False
if exists:
print('!')
obj = s3_client.get_object(Bucket=bucket, Key=output_key)
predictions = json.loads(obj['Body'].read().decode())['predictions'][0]
print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
break
else:
print('.',end='')
sleep(0.1)
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
# ## 7. オートスケール
# * Endpoint はオートスケールさせることができる
# * 推論が増えたら自動で増強、減ったら削減、など
# * スケーリング対象のメトリクスは[こちら](https://docs.aws.amazon.com/sagemaker/latest/dg/monitoring-cloudwatch.html#cloudwatch-metrics-endpoint-invocation)
# * 以下は非同期推論を用いてオートスケールをした場合だが、同期推論もやり方は同じ
# * オートスケールはエンドポイントを立てた後、 AWS の アプリケーションオートスケーリングサービスを利用して実現する
MODEL_NAME = 'MyTFModelFromBoto3AsyncWithAutoScaling'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'EndpointConfig'
ENDPOINT_NAME = MODEL_NAME + 'Endpoint'
# ### モデルの作成
response = sm_client.create_model(
ModelName=MODEL_NAME,
PrimaryContainer={
'Image': container_image_tf24_uri,
'ModelDataUrl': model_s3_uri,
},
ExecutionRoleArn=sm_role,
)
# ### 推論エンドポイントの設定
VARIANT_NAME = 'MyVariant'
response = sm_client.create_endpoint_config(
EndpointConfigName=ENDPOINT_CONFIG_NAME,
ProductionVariants=[
{
'VariantName': VARIANT_NAME,
'ModelName': MODEL_NAME,
'InitialInstanceCount': 1,
'InstanceType': 'ml.m5.xlarge',
},
],
AsyncInferenceConfig={
"OutputConfig": {
"S3OutputPath": f"s3://{bucket}/async_inference_with_autoscaling/output"
},
}
)
# +
response = sm_client.create_endpoint(
EndpointName=ENDPOINT_NAME,
EndpointConfigName=ENDPOINT_CONFIG_NAME,
)
endpoint_inservice_waiter.wait(
EndpointName=ENDPOINT_NAME,
WaiterConfig={'Delay': 5,}
)
# +
# オートスケーリングの設定
aa_client = boto3.client('application-autoscaling')
SCALABLE_DIMENSION = 'sagemaker:variant:DesiredInstanceCount'
resource_id = (f'endpoint/{ENDPOINT_NAME}/variant/{VARIANT_NAME}')
response = aa_client.register_scalable_target(
ServiceNamespace="sagemaker",
ResourceId=resource_id,
ScalableDimension=SCALABLE_DIMENSION,
MinCapacity=1,
MaxCapacity=2,
)
response = aa_client.put_scaling_policy(
PolicyName="Invocations-ScalingPolicy",
ServiceNamespace="sagemaker",
ResourceId=resource_id,
ScalableDimension=SCALABLE_DIMENSION,
PolicyType="TargetTrackingScaling",
TargetTrackingScalingPolicyConfiguration={
"TargetValue": 1.0,
"CustomizedMetricSpecification": {
"MetricName": "ApproximateBacklogSizePerInstance",
"Namespace": "AWS/SageMaker",
"Dimensions": [{"Name": "EndpointName", "Value": ENDPOINT_NAME}],
"Statistic": "Average",
},
"ScaleInCooldown": 10,
"ScaleOutCooldown": 10
},
)
# -
# インスタンス数の確認
instance_count=sm_client.describe_endpoint(EndpointName=ENDPOINT_NAME)['ProductionVariants'][0]['CurrentInstanceCount']
print(f'現在稼動しているインスタンス数: {instance_count}')
# ### 推論を50000回行って負荷をかけてオートスケーリングするかを確認する
# +
# %%time
json_name = './tabby.json'
with open(json_name,'wt') as f:
f.write(json.dumps({"instances": img_arr.tolist()}))
tabby_s3_uri = sagemaker.s3.S3Uploader.upload(
local_path = json_name,
desired_s3_uri = f"s3://{bucket}/async_inference/input"
)
output_key_list = []
# 推論
for _ in range(50000):
response = smr_client.invoke_endpoint_async(
EndpointName=ENDPOINT_NAME,
InputLocation=tabby_s3_uri,
ContentType='application/json'
)
output_s3_uri = response['OutputLocation']
output_key = output_s3_uri.replace(f's3://{bucket}/','')
output_key_list.append(output_key)
# 全ての結果を確認する
for output_key in output_key_list:
while True:
result = s3_client.list_objects(Bucket=bucket, Prefix=output_key)
exists = True if "Contents" in result else False
if exists:
# print('!',end='')
# # 結果確認
# obj = s3_client.get_object(Bucket=bucket, Key=output_key)
# predictions = json.loads(obj['Body'].read().decode())['predictions'][0]
# print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
break
else:
print('.',end='')
sleep(1)
# インスタンス数の確認
instance_count=sm_client.describe_endpoint(EndpointName=ENDPOINT_NAME)['ProductionVariants'][0]['CurrentInstanceCount']
print(f'現在稼動しているインスタンス数: {instance_count}')
# -
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
# ## 8. サーバーレス推論(2021/12時点ではパブリックプレビュー)
# * インスタンスを意識せずにエンドポイントだけ建てる
# * 実態は推論イベント発生時に都度コンピューティングリソースが立ち上がる
# * 通常のリアルタイム推論との違いは `create_endpoint_config` する際に、インスタンス数やインスタンスタイプの設定はしなくなり、代わりに`ServerlessConfig`の中でメモリと最大同時期同数を設定する
# +
MODEL_NAME = 'MyTFModelFromBoto3Serverless'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'EndpointConfig'
ENDPOINT_NAME = MODEL_NAME + 'Endpoint'
response = sm_client.create_model(
ModelName=MODEL_NAME,
PrimaryContainer={
'Image': container_image_tf24_uri,
'ModelDataUrl': model_s3_uri,
},
ExecutionRoleArn=sm_role,
)
response = sm_client.create_endpoint_config(
EndpointConfigName=ENDPOINT_CONFIG_NAME,
ProductionVariants=[
{
'VariantName': 'AllTrafic',
'ModelName': MODEL_NAME,
# インスタンスカウントやインスタンスタイプはなくなる
'ServerlessConfig': { # 通常のリアルタイム推論とは違い、ServerlessConfig というキーで設定する
'MemorySizeInMB': 1024, # メモリサイズは 1024 , 2048, 3072, 4096, 5120, 6144 から選ぶ
'MaxConcurrency': 3 # 最大同時起動数
}
},
],
)
response = sm_client.create_endpoint(
EndpointName=ENDPOINT_NAME,
EndpointConfigName=ENDPOINT_CONFIG_NAME,
)
endpoint_inservice_waiter.wait(
EndpointName=ENDPOINT_NAME,
WaiterConfig={'Delay': 5,}
)
# -
# jsonにして渡すパターン
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'Body' : json.dumps({"instances": img_arr.tolist()})
}
response = smr_client.invoke_endpoint(**request_args)
predictions = json.loads(response['Body'].read().decode('utf-8'))['predictions'][0]
print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
# ## 9. 独自コンテナイメージの持ち込みを利用した推論
# * SageMaker のマネージドコンテナイメージ以外に、独自のコンテナイメージを持ち込める
# * コンテナイメージをビルドし、ECR にプッシュしてその URL を指定する以外はマネージドコンテナイメージと使い方が一緒
# * SageMaker Studio の場合はコンテナイメージのビルドに `sm-docker build` コマンドを使う必要がある
# * `sm-docker build` する前にロールの信頼関係とポリシーを追加する必要がある
# `sm-docker build` が裏側で AWS CodeBuild を使うために必要
# * SageMaker Notebook (classic) の場合はそのまま `docker build` コマンドを使えばよい
# 定数の設定
IMAGE_NAME = 'sagemaker_byoc_tf_inference-cpu'
TAG = ':1'
# ### ビルドするコンテナイメージと、依存関係のあるモジュールの確認
# !cat container/Dockerfile
# !cat container/requirements.txt
# ### **SageMaker Studio を使っている場合のみ**以下を実行
# #### sm-docker コマンドのインストール
# !pip install sagemaker-studio-image-build
# #### Studio アタッチされているロールに以下の信頼関係を追加
# 以下のコマンドの出力結果をコピーして追加する
# !cat container/trust_relationships.json
# #### Studio アタッチされているロールにインラインポリシー(もしくはポリシーを別途作成して)アタッチする
# 以下のコマンドの出力結果をコピーしてアタッチする
# !cat container/inline_policy.json
# #### コンテナイメージのビルド
# %cd container
# !chmod +x tfserve/serve
# !sm-docker build . --repository {IMAGE_NAME}{TAG}
# %cd ../
account_id = boto3.client('sts').get_caller_identity()['Account']
region_name = boto3.session.Session().region_name
tf_own_image_uri = f'{account_id}.dkr.ecr.{region_name}.amazonaws.com/{IMAGE_NAME}{TAG}'
print(tf_own_image_uri)
# ### **SageMaker Notebook (classic) をつかってる場合のみ**以下を実行
# %cd container
# !chmod +x tfserve/serve
# !docker build -t {IMAGE_NAME}{TAG} .
# %cd ../
# +
# boto3の機能を使ってリポジトリ名に必要な情報を取得する
account_id = boto3.client('sts').get_caller_identity().get('Account')
region = boto3.session.Session().region_name
ecr_endpoint = f'{account_id}.dkr.ecr.{region}.amazonaws.com/'
repository_uri = f'{ecr_endpoint}{IMAGE_NAME}'
tf_own_image_uri = f'{repository_uri}{TAG}'
# !aws ecr get-login-password --region {region} | docker login --username AWS --password-stdin {ecr_endpoint}
# !docker tag {image_name}{tag} {image_uri}
# 同名のリポジトリがあった場合は削除
# !aws ecr delete-repository --repository-name $image_name --force
# リポジトリを作成
# !aws ecr create-repository --repository-name $image_name
# イメージをプッシュ
# !docker push {image_uri}
# -
# ### 以下共通で推論エンドポイントの作成
# +
MODEL_NAME = 'MyTFModelFromBoto3BYOC'
ENDPOINT_CONFIG_NAME = MODEL_NAME + 'EndpointConfig'
ENDPOINT_NAME = MODEL_NAME + 'Endpoint'
response = sm_client.create_model(
ModelName=MODEL_NAME,
PrimaryContainer={
'Image': tf_own_image_uri,
'ModelDataUrl': model_s3_uri,
},
ExecutionRoleArn=sm_role,
)
response = sm_client.create_endpoint_config(
EndpointConfigName=ENDPOINT_CONFIG_NAME,
ProductionVariants=[
{
'VariantName': 'AllTrafic',
'ModelName': MODEL_NAME,
'InitialInstanceCount': 1,
'InstanceType': 'ml.m5.xlarge',
},
],
)
response = sm_client.create_endpoint(
EndpointName=ENDPOINT_NAME,
EndpointConfigName=ENDPOINT_CONFIG_NAME,
)
endpoint_inservice_waiter.wait(
EndpointName=ENDPOINT_NAME,
WaiterConfig={'Delay': 5,}
)
# -
# jsonにして渡すパターン
request_args = {
'EndpointName': ENDPOINT_NAME,
'ContentType' : 'application/json',
'Accept' : 'application/json',
'Body' : json.dumps({"instances": img_arr.tolist()})
}
response = smr_client.invoke_endpoint(**request_args)
predictions = json.loads(response['Body'].read().decode('utf-8'))['predictions'][0]
print(labels[np.argmax(predictions)],predictions[np.argmax(predictions)])
r = sm_client.delete_endpoint(EndpointName=ENDPOINT_NAME)
r = sm_client.delete_endpoint_config(EndpointConfigName=ENDPOINT_CONFIG_NAME)
r = sm_client.delete_model(ModelName=MODEL_NAME)
|
sagemaker-inference-features-tensorflow.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] graffitiCellId="id_dwjdx8h"
# # Recursion
# ## Introduction
#
# Recursion is a technique for solving problems where the solution to a particular problem depends on the solution to a smaller instance of the same problem.
#
# Consider the problem of calculating $\mathtt{2^5}$. Let's assume to calculate this, you need to do one multiplication after another. That's $2 * 2 * 2 * 2 * 2$. We know that $2^5 = 2 * 2^4$. If we know the value of $2^4$, we can easily calculate $2^5$.
#
# We can use recursion to solve this problem, since the solution to the original problem ($2^n$) depends on the solution to a smaller instance ($2^{n-1}$) of the same problem. The recursive solution is to calculate $2 * 2^{n-1}$ for all n that is greater than 0. If n is 0, return 1. We'll ignore all negative numbers.
#
# Let's look at what the recursive steps would be for calculating $2^5$.
#
# $2^5 = 2 * 2^4$
#
# $2^5 = 2 * 2 * 2^3$
#
# $2^5 = 2 * 2 * 2 * 2^2$
#
# $2^5 = 2 * 2 * 2 * 2 * 2^1$
#
# $2^5 = 2 * 2 * 2 * 2 * 2 * 2^0$
#
# $2^5 = 2 * 2 * 2 * 2 * 2 * 1$
#
# ## Code
# Let's look at the recursive function `power_of_2`, which calculates $2^n$.
# + graffitiCellId="id_szy5ni3"
def power_of_2(n):
if n == 0:
return 1
return 2 * power_of_2(n - 1)
print(power_of_2(5))
# + [markdown] graffitiCellId="id_npqrqt3"
# As you can see, the function calls itself to calculate the smaller instance of the solution. Let's break down the `power_of_2` function, starting with the first two lines.
# ```
# if n == 0:
# return 1
# ```
# These lines contain the base case. This is where you catch edge cases that don't fit the problem ($2 * 2^{n-1}$). Since we aren't considering any $n < 0$ valid, $2 * 2^{n-1}$ can't be used when $n$ is $0$. This section of the code returns the solution to $2^0$ without using $2 * 2^{n-1}$.
# ```
# return 2 * power_of_2(n - 1)
# ```
# This code is where it breaks the problem down into smaller instances. Using the formula $2^{n} = 2 * 2^{n-1}$, the `power_of_2` function calls itself to calculate $2^{n-1}$. To better understand what is happening, let's look at the call stack with an example.
# + [markdown] graffitiCellId="id_g25qq7v"
# ## Call Stack
# Let's follow the [call stack](https://en.wikipedia.org/wiki/Call_stack) when calling `power_of_2(5)`:
#
# First `power_of_2(5)` is called.
#
# Then `power_of_2(5)` calls `power_of_2(4)`
#
# Then `power_of_2(4)` calls `power_of_2(3)`
#
# ...
#
# Then `power_of_2(1)` calls `power_of_2(0)`
#
# At this point, the call stack will look something like this:
# ```
# ...
# File "<ipython-input-27-9e8459c7465f>", line 5, in power_of_2
# return 2 * power_of_2(n - 1)
# File "<ipython-input-27-9e8459c7465f>", line 5, in power_of_2
# return 2 * power_of_2(n - 1)
# File "<ipython-input-27-9e8459c7465f>", line 5, in power_of_2
# return 2 * power_of_2(n - 1)
# File "<ipython-input-27-9e8459c7465f>", line 5, in power_of_2
# return 2 * power_of_2(n - 1)
# File "<ipython-input-27-9e8459c7465f>", line 3, in power_of_2
# return 1
# ```
# Let's look at a cleaner view of the stack:
# ```
# ...
# -> power_of_2(5)
# -> power_of_2(4)
# -> power_of_2(3)
# -> power_of_2(2)
# -> power_of_2(1)
# -> power_of_2(0)
# ```
# Each function is waiting on the function it called to complete. So, `power_of_2(5)` is waiting for `power_of_2(4)`, `power_of_2(4)` is waiting for `power_of_2(3)`, etc..
#
# The function `power_of_2(0)` will return $1$
#
# Using the 1 returned from `power_of_2(0)`, `power_of_2(1)` will return $2 * 1$
#
# Using the 2 returned from `power_of_2(1)`, `power_of_2(2)` will return $2 * 2$
#
# ...
#
#
# Using the 16 returned from `power_of_2(4)`, `power_of_2(5)` will return $2 * 16$
#
# Finally, the result of $2^5$ is returned! $2^5 = 2 * 2^4 = 2 * 16 = 32$
#
# ### Practice Problem
# Implement `sum_integers(n)` to calculate the sum of all integers from $1$ to $n$ using recursion. For example, `sum_integers(3)` should return $6$ ($1 + 2 + 3$).
# + graffitiCellId="id_3937ww3"
def sum_integers(n):
if n == 1:
return 1
return n + sum_integers(n -1)
print(sum_integers(4))
# + [markdown] graffitiCellId="id_i8ruo8b"
# ## Gotchas
# When using recursion, there are a few things to look out for that you don't have to worry about when running a loop (iteratively). Let's go over a few of those items.
#
# ### Call Stack
# We went over an example of the call stack when calling `power_of_2(5)` above. In this section, we'll cover the limitations of recursion on a call stack. Run the cell below to create a really large stack. It should raise the error `RecursionError: maximum recursion depth exceeded in comparison`.
# + graffitiCellId="id_snmtgsf"
print(power_of_2(10000))
# + [markdown] graffitiCellId="id_wlikaoq"
# Python has a limit on the depth of recursion to prevent a [stack overflow](https://en.wikipedia.org/wiki/Stack_overflow). However, some compilers will turn [tail-recursive functions](https://en.wikipedia.org/wiki/Recursion_(computer_science)#Tail-recursive_functions) into an iterative loop to prevent recursion from using up the stack. Since Python's compiler doesn't do this, you'll have to watch out for this limit.
# ### Slicing
# Let's look at recursion on arrays and how you can run into the problem of slicing the array. If you haven't heard the term slicing, it's the operation of taking a subset of some data. For example, the list `a` can be sliced using the following operation: `a[start:stop]`. This will return a new list from index `start` (inclusive) to index `stop` (exclusive).
#
# Let's look at an example of a recursive function that takes the sum of all numbers in an array. For example, the array of `[5, 2, 9, 11]` would sum to 27 (5 + 2 + 9 + 11).
# + graffitiCellId="id_v9ovfz1"
#solution with print
def sum_array(array):
print('\narray',array)
print('len(array)',len(array))
print('array[1:]',array[1:])
print('array[0]',array[0])
# Base Case
if len(array) == 1:
return array[0]
return array[0] + sum_array(array[1:])
arr = [5, 2, 9, 11]
print(sum_array(arr))
print('\nans11+[ ]')
print('9+11')
print('2+20')
print('5+22')
# +
#solution with print
def sum_array(array):
# Base Case
if len(array) == 1:
return array[0]
return array[0] + sum_array(array[1:])
arr = [1, 2, 3, 4]
print(sum_array(arr))
# + [markdown] graffitiCellId="id_54i7x5u"
# Looking at this, you might think it has a running time of O($n$), but that isn't correct due to the slice operation `array[1:]`. This operation will take O($k$) time to run where $k$ is the number of elements to copy. So, this function is actually O($k*n$) running time complexity and O($k*n$) space complexity.
#
# To visualize this, let's plot the time it takes to slice.
# + graffitiCellId="id_ocuve8r"
import matplotlib.pyplot as plt
import statistics
import time
# %matplotlib inline
n_steps = 10
step_size = 1000000
array_sizes = list(range(step_size, n_steps*step_size, step_size))
big_array = list(range(n_steps*step_size))
times = []
# Calculate the time it takes for the slice function to run with different sizes of k
for array_size in array_sizes:
start_time = time.time()
big_array[:array_size]
times.append(time.time() - start_time)
# Graph the results
plt.scatter(x=array_sizes, y=times)
plt.ylim(top=max(times), bottom=min(times))
plt.xlabel('Array Size')
plt.ylabel('Time (seconds)')
plt.plot()
# + [markdown] graffitiCellId="id_pvlssjf"
# As you can see, it's linear time to slice.
#
# Instead of slicing, we can pass the index for the element that we want to use for addition. That will give us the following function:
# + graffitiCellId="id_14u4wyj"
def sum_array_index(array, index):
# Base Cases
if len(array) - 1 == index:
return array[index]
return array[index] + sum_array_index(array, index + 1)
arr = [1, 2, 3, 4]
print(sum_array_index(arr, 0))
# + [markdown] graffitiCellId="id_cxz3ww6"
# That eliminates the need to do slicing. With the two different functions implemented, let's compare the running times.
# + graffitiCellId="id_h6g7uc7"
import matplotlib.pyplot as plt
import statistics
import time
n_steps = 10
step_size = 200
array_sizes = list(range(step_size, n_steps*step_size, step_size))
big_array = list(range(n_steps*step_size))
sum_array_times = []
sum_array_index_times = []
for array_size in array_sizes:
subset_array = big_array[:array_size]
start_time = time.time()
sum_array(subset_array)
sum_array_times.append(time.time() - start_time)
start_time = time.time()
sum_array_index(subset_array, 0)
sum_array_index_times.append(time.time() - start_time)
plt.scatter(x=array_sizes, y=sum_array_times, label='sum_array')
plt.scatter(x=array_sizes, y=sum_array_index_times, label='sum_array_index')
plt.ylim(
top=max(sum_array_times + sum_array_index_times),
bottom=min(sum_array_times + sum_array_index_times))
plt.legend()
plt.xlabel('Array Size')
plt.ylabel('Time (seconds)')
plt.plot()
# + [markdown] graffitiCellId="id_wgssm9u"
# As you can see, the function `sum_array` is a polynomial and `sum_array_index` is linear as we predicted.
#
# However, in our pursuit to use recursion we actually made things worse. Let's look at an iterative solution to this problem:
# + graffitiCellId="id_z5yodhr"
def sum_array_iter(array):
result = 0
for x in array:
result += x
return result
arr = [1, 2, 3, 4]
print(sum_array_iter(arr))
# + [markdown] graffitiCellId="id_ogosv6y"
# The `sum_array_iter` function is a lot more straightforward than the two recursive functions, which is important. Second, to help ensure an answer that is correct and bug free, you generally want to pick the solution that is more readable. In some cases recursion is more readable and in some cases iteration is more readable. As you gain experience reading other people’s code, you’ll get an intuition for code readability.
|
recursion/.ipynb_checkpoints/simple_recursion-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:good_robot]
# language: python
# name: conda-env-good_robot-py
# ---
import numpy as np
import matplotlib.pyplot as plt
import networkx as nx
# +
def make_graphs(n=2, i=None, j=None, js=None):
"""Make a graph, recursively, by either including, or skipping an edge.
Edges are given in lexicographical order, by construction."""
out = []
if i is None: # First call
out = [[(0,1)]+r for r in make_graphs(n=n, i=0, j=1, js=set([1]))]
elif j<n-1:
out += [[(i,j+1)]+r for r in make_graphs(n=n, i=i, j=j+1, js=js | set([j+1]))]
out += [ r for r in make_graphs(n=n, i=i, j=j+1, js=js)]
elif i<n-1:
out = make_graphs(n=n, i=i+1, j=i+1, js=js)
else:
out = [[]]
return out
gs = make_graphs(3)
print('Drawing...')
# plot_graphs(gs)
# +
def perm(n, s=None):
"""All permutations of n elements."""
if s is None: return perm(n, tuple(range(n)))
if not s: return [[]]
return [[i]+p for i in s for p in perm(n, tuple([k for k in s if k!=i]))]
perm(3)
# +
def permute(g, n):
"""Create a set of all possible isomorphic codes for a graph,
as nice hashable tuples. All edges are i<j, and sorted lexicographically."""
ps = perm(n)
out = []
for p in ps:
out.append(tuple(sorted([(p[i],p[j]) if p[i]<p[j] else (p[j],p[i])for i,j in g])))
return out
permute([(0, 1), (0, 2)], 3)
# +
def connected(g):
"""Check if the graph is full connected, with Union-Find."""
nodes = set([i for e in g for i in e])
roots = {node: node for node in nodes}
def _root(node, depth=0):
if node==roots[node]: return (node, depth)
else: return _root(roots[node], depth+1)
for i,j in g:
ri,di = _root(i)
rj,dj = _root(j)
if ri==rj: continue
if di<=dj: roots[ri] = rj
else: roots[rj] = ri
return len(set([_root(node)[0] for node in nodes]))==1
assert connected([(0, 1), (0, 2), (1, 3)])
assert not connected([(0, 1), (2, 3)])
# +
def filter(gs, target_nv):
"""Filter all improper graphs: those with not enough nodes,
those not fully connected, and those isomorphic to previously considered."""
mem = set({})
gs2 = []
for g in gs:
nv = len(set([i for e in g for i in e]))
if nv != target_nv:
continue
if not connected(g):
continue
if tuple(sorted(g)) not in mem:
gs2.append(g)
mem |= set(permute(g, target_nv))
#print('\n'.join([str(a) for a in mem]))
return gs2
filter(gs, 3)
# +
def plot_graphs(graphs, figsize=14, dotsize=20):
"""Utility to plot a lot of graphs from an array of graphs.
Each graphs is a list of edges; each edge is a tuple."""
n = len(graphs)
fig = plt.figure(figsize=(figsize,figsize))
fig.patch.set_facecolor('white') # To make copying possible (no transparent background)
k = int(np.sqrt(n))
for i in range(n):
plt.subplot(k+1,k+1,i+1)
g = nx.Graph()
for e in graphs[i]:
g.add_edge(e[0],e[1])
nx.draw_kamada_kawai(g, node_size=dotsize)
print('.', end='')
#plot_graphs([[(0,1),(1,2),(1,3)]])
# -
len(gs)
NV = 6
print('Building...')
gs = make_graphs(NV)
print('Filtering...')
gs = filter(gs, NV)
print('Drawing...')
graphs = gs
n = len(graphs)
f, axs = plt.subplots(8, 14, figsize=(14, 8))
axs = axs.flatten()
for i, ax in enumerate(axs):
plt.sca(ax)
g = nx.Graph()
for e in graphs[i]:
g.add_edge(e[0],e[1])
nx.draw_kamada_kawai(g, node_size=0)
f.savefig('images/0019_ngraphs.svg', dpi=300)
n = len(graphs)
point_sets = []
for i, ax in enumerate(axs):
g = nx.Graph()
for e in graphs[i]:
g.add_edge(e[0],e[1])
x = nx.kamada_kawai_layout(g)
point_sets.append(x)
from matplotlib.collections import LineCollection
import bezier
def gen_bezier(nodes, bez_eval_start=0, bez_eval_end=1, n_eval_points=1000):
nodes = np.asfortranarray(nodes)
curve = bezier.Curve(nodes, degree=(nodes.shape[1]-1))
eval_points = np.linspace(bez_eval_start, bez_eval_end, n_eval_points)
x, y = curve.evaluate_multi(eval_points)
return np.stack([x, y])
# +
xs, ys = np.meshgrid(np.arange(8), np.arange(14))
xs = xs.flatten() * 2.5
ys = ys.flatten() * 2.5
edge_sets = []
bez_noise_scale = 0.4
for i in range(n):
x = xs[i]
y = ys[i]
center = np.array([x, y])
graph = graphs[i]
ps = point_sets[i]
edges = []
for e0,e1 in graph:
p0 = ps[e0] + center
p1 = ps[e1] + center
pmid = np.mean([p0, p1], axis=0) + np.random.randn(2) * bez_noise_scale
edges.append(np.stack([p0, pmid, p1]).T)
edge_sets += edges
bez_lines = [gen_bezier(edge).T for edge in edge_sets]
f,ax = plt.subplots(figsize=(8,14))
lc = LineCollection(bez_lines, color='k', alpha=0.2)
ax.add_collection(lc)
ax.axis('tight')
ax.axis('off')
# +
xs, ys = np.meshgrid(np.arange(14), np.arange(8))
xs = xs.flatten() * 2.5
ys = ys.flatten() * 2.5
edge_sets = []
bez_noise_scale = 0.3
for i in range(n):
x = xs[i]
y = ys[i]
center = np.array([x, y])
graph = graphs[i]
ps = point_sets[i]
edges = []
for e0,e1 in graph:
p0 = ps[e0] + center
p1 = ps[e1] + center
pmid = np.mean([p0, p1], axis=0) + np.random.randn(2) * bez_noise_scale
edges.append(np.stack([p0, pmid, p1]).T)
edge_sets += edges
bez_lines = [gen_bezier(edge).T for edge in edge_sets]
f,ax = plt.subplots(figsize=(14,8))
lc = LineCollection(bez_lines, color='k', alpha=0.2)
ax.add_collection(lc)
ax.axis('tight')
ax.axis('off')
# -
from pathlib import Path
savedir = Path('/mnt/c/code/side/good_robot/images')
filename = '0055_kamada_kawaii.png'
save_filepath = savedir.joinpath(filename)
f.savefig(save_filepath)
|
04_graph_grams.ipynb
|
# ---
# jupyter:
# jupytext:
# formats: Rmd,ipynb
# notebook_metadata_filter: jupytext_format_version,jupytext_formats,language_info
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# language_info:
# codemirror_mode:
# name: ipython
# version: 3
# file_extension: .py
# mimetype: text/x-python
# name: python
# nbconvert_exporter: python
# pygments_lexer: ipython3
# version: 3.6.5
# ---
# # Literary characters
# + tags=["hide-cell"]
# HIDDEN
# The standard set of libraries we need
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# Make plots look a little bit more fancy
plt.style.use('fivethirtyeight')
# The standard library for data in tables
import pandas as pd
# A tiny function to read a file directly from a URL
from urllib.request import urlopen
def read_url(url):
return urlopen(url).read().decode()
# + tags=["hide-cell"]
# HIDDEN
# Read the text of Pride and Prejudice, split into chapters.
book_url = 'http://www.gutenberg.org/ebooks/42671.txt.utf-8'
book_text = read_url(book_url)
# Break the text into Chapters
book_chapters = book_text.split('CHAPTER ')
# Drop the first "Chapter" - it's the Project Gutenberg header
book_chapters = book_chapters[1:]
# -
# [Pride and Prejudice](https://en.wikipedia.org/wiki/Pride_and_Prejudice) is
# the story of five sisters: Jane, Elizabeth, Mary, Kitty and Lydia, and their
# journey through the social life of the mid-17th century. You may remember
# that Elizabeth ends up marrying the dashing and aloof Mr Darcy, but along the
# way, the feckless Lydia runs off with the equally feckless Mr Wickham, and the
# slightly useless Mr Bingley wants to marry Jane, the most beautiful of the
# sisters.
#
# We can see when these characters appear in the book, by counting how many
# times their names are mentioned in each chapter.
# +
# Count how many times the characters appear in each chapter.
counts = pd.DataFrame.from_dict({
'Elizabeth': np.char.count(book_chapters, 'Elizabeth'),
'Darcy': np.char.count(book_chapters, 'Darcy'),
'Lydia': np.char.count(book_chapters, 'Lydia'),
'Wickham': np.char.count(book_chapters, 'Wickham'),
'Bingley': np.char.count(book_chapters, 'Bingley'),
'Jane': np.char.count(book_chapters, 'Jane')},
)
# The cumulative counts:
# how many times in Chapter 1, how many times in Chapters 1 and 2, and so on.
cum_counts = counts.cumsum()
# Add the chapter numbers
number_of_chapters = len(book_chapters)
cum_counts['Chapter'] = np.arange(number_of_chapters)
# Do the plot
cum_counts.plot(x='Chapter')
plt.title('Cumulative Number of Times Each Name Appears');
# -
# In the plot above, the horizontal axis shows chapter numbers and the vertical
# axis shows how many times each character has been mentioned up to and
# including that chapter.
#
# Notice first that Elizabeth and Darcy are the main characters. Around chapter
# 13 we see Wickham and Lydia spike up, as they run away together, and mentions
# of Darcy flatten off, when he goes to look for them. Around chapter 50 we see
# Jane and Bingley being mentioned at a very similar rate, as Bingley proposes,
# and Jane accepts.
#
# {ucb-page}`Literary_Characters`
|
intro/Literary_Characters.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 64-bit (''python39-2vM0bQN-'': pipenv)'
# name: python3
# ---
# install syft if we are on google colab
# %load_ext autoreload
# %autoreload 2
import syft as sy
print(f"You're running syft version: {sy.__version__}")
import numpy as np
import pandas as pd
from tqdm import tqdm
import time
domain_client = sy.login(email='<EMAIL>', password='<PASSWORD>',port = 8081)
# +
data = domain_client.datasets[-1]
X_train = data["train_images"]
Y_train = data["train_labels"]
X_dev = data["train_images"]
Y_dev = data["train_labels"]
m,n = X_train.public_shape
X_train = X_train.T
X_train = X_train *(1/ 255.0)
X_dev = X_dev.T
X_dev = X_dev *(1/ 255.0)
# +
from syft.core.node.common.node_service.user_manager.user_messages import (
UpdateUserMessage,
)
# Upgrade admins budget
content = {"user_id": 1, "budget": 9_999_999}
domain_client._perform_grid_request(grid_msg=UpdateUserMessage, content=content)
domain_client.privacy_budget
# +
def init_params(input_size: int):
print(f"Using input size: {input_size}")
W1 = np.random.rand(8, input_size) - 0.5
b1 = np.random.rand(8, 1) - 0.5
W2 = np.random.rand(8, 8) - 0.5
b2 = np.random.rand(8, 1) - 0.5
return W1, b1, W2, b2
def ReLU(Z):
return Z*(Z>0)
def softmax(Z):
exp_cache = Z.exp()
inv = (exp_cache.sum().reciprocal())
A = exp_cache * inv
return A
def forward_prop(W1, b1, W2, b2, X):
Z1 = X.__rmatmul__(W1) + b1
A1 = ReLU(Z1)
Z2 = A1.__rmatmul__(W2) + b2
A2 = softmax(Z2)
return Z1, A1, Z2, A2
def ReLU_deriv(Z):
return Z > 0
def one_hot(Y):
one_hot_Y = np.zeros((Y.size, Y.max() + 1))
one_hot_Y[np.arange(Y.size),Y] = 1
one_hot_Y = one_hot_Y.T
return one_hot_Y
def backward_prop(Z1, A1, Z2, A2, W1, W2, X, Y):
one_hot_Y = Y.one_hot()
dZ2 = A2 - one_hot_Y
dW2 = dZ2@(A1.T) * (1/m)
db2 = dZ2.sum() * (1/m)
dZ1 = dZ2.__rmatmul__(W2.T) * ReLU_deriv(Z1)
dW1 = dZ1@(X.T) * (1/m)
db1 = dZ1.sum()*(1/m)
return dW1, db1, dW2, db2
def update_params(W1, b1, W2, b2, dW1, db1, dW2, db2, alpha):
W1 = (dW1 * alpha - W1) * -1
b1 = (db1 * alpha - b1) * -1
W2 = (dW2 * alpha - W2) * -1
b2 = (db2 * alpha - b2) * -1
return W1, b1, W2, b2
def get_predictions(A2):
return np.argmax(A2, 0)
def get_accuracy(predictions, Y):
return np.sum(predictions == Y) / Y.size
def gradient_descent(X, Y, alpha, iterations):
W1, b1, W2, b2 = init_params(X.public_shape[0])
print("[INFO]: Starting training!\n")
for i in tqdm(range(iterations)):
Z1, A1, Z2, A2 = forward_prop(W1, b1, W2, b2, X)
dW1, db1, dW2, db2 = backward_prop(Z1, A1, Z2, A2, W1, W2, X, Y)
W1, b1, W2, b2 = update_params(W1, b1, W2, b2, dW1, db1, dW2, db2, alpha)
# if i % 40 == 0:
# print("Predicition will be availabe in the next iteration.....Thank you for your support.")
# # predictions = get_predictions(A2)
# # print("Iteration: ", i, " Accuracy : " ,get_accuracy(predictions, Y))
return W1, b1, W2, b2
# -
W1, b1, W2, b2 = gradient_descent(X_train, Y_train, 0.10, 1)
for ptr in [W1, b1, W2, b2]:
ptr.block_with_timeout(60)
print(W1.exists)
print(b1.exists)
print(W2.exists)
print(b2.exists)
# +
# DEBUG CELLS
# W1_O = W1.get_copy()
# print(type(W1_O))
# b1_O = b1.get_copy()
# print(type((b1_O.child)))
# W2_O = W2.get_copy()
# print(type((W2_O.child)))
# b2_O = b2.get_copy()
# print(type((b2_O.child)))
# -
W1 = W1.publish(sigma=1000)
b1 = b1.publish(sigma=1000)
W2 = W2.publish(sigma=1000)
b2 = b2.publish(sigma=1000)
a,b,c,d = init_params(X_train.public_shape[0])
print(a.shape)
print(b.shape)
print(c.shape)
print(d.shape)
t1,t2,t3,t4 = W1.get_copy(),b1.get_copy(),W2.get_copy(),b2.get_copy()
print(t1.shape)
print(t2.shape)
print(t3.shape)
print(t4.shape)
t4
|
notebooks/Experimental/Rasswanth/TissueMNIST_demo/TM-data-scientist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# # !sudo pip install git+https://github.com/anttttti/Wordbatch.git
# -
import wordbatch
from wordbatch.extractors import WordHash
from wordbatch.models import FM_FTRL
# from wordbatch.data_utils import *
import threading
import pandas as pd
from sklearn.metrics import roc_auc_score
import time
import numpy as np
import gc
from contextlib import contextmanager
# +
@contextmanager
def timer(name):
t0 = time.time()
yield
print(f'[{name}] done in {time.time() - t0:.0f} s')
def df_add_counts(df, cols):
arr_slice = df[cols].values
unq, unqtags, counts = np.unique(np.ravel_multi_index(arr_slice.T, arr_slice.max(0) + 1),
return_inverse=True, return_counts=True)
df["_".join(cols)+'_count'] = counts[unqtags]
def df2csr(wb, df, pick_hours=None):
df.reset_index(drop=True, inplace=True)
with timer("Adding counts"):
df['click_time']= pd.to_datetime(df['click_time'])
dt= df['click_time'].dt
df['day'] = dt.day.astype('uint8')
df['hour'] = dt.hour.astype('uint8')
del(dt)
df_add_counts(df, ['ip', 'day', 'hour'])
df_add_counts(df, ['ip', 'app'])
df_add_counts(df, ['ip', 'app', 'os'])
df_add_counts(df, ['ip', 'device'])
df_add_counts(df, ['app', 'channel'])
#cpuStats()
with timer("Adding next click times"):
D= 2**26
df['category'] = (df['ip'].astype(str) + "_" + df['app'].astype(str) + "_" + df['device'].astype(str) \
+ "_" + df['os'].astype(str)).apply(hash) % D
click_buffer= np.full(D, 3000000000, dtype=np.uint32)
df['epochtime']= df['click_time'].astype(np.int64) // 10 ** 9
next_clicks= []
for category, time in zip(reversed(df['category'].values), reversed(df['epochtime'].values)):
next_clicks.append(click_buffer[category]-time)
click_buffer[category]= time
del(click_buffer)
df['next_click']= list(reversed(next_clicks))
for fea in ['ip_day_hour_count','ip_app_count','ip_app_os_count','ip_device_count',
'app_channel_count','next_click']: df[fea]= np.log2(1 + df[fea].values).astype(int)
with timer("Generating str_array"):
str_array= ("I" + df['ip'].astype(str) \
+ " A" + df['app'].astype(str) \
+ " D" + df['device'].astype(str) \
+ " O" + df['os'].astype(str) \
+ " C" + df['channel'].astype(str) \
+ " WD" + df['day'].astype(str) \
+ " H" + df['hour'].astype(str) \
+ " AXC" + df['app'].astype(str)+"_"+df['channel'].astype(str) \
+ " OXC" + df['os'].astype(str)+"_"+df['channel'].astype(str) \
+ " AXD" + df['app'].astype(str)+"_"+df['device'].astype(str) \
+ " IXA" + df['ip'].astype(str)+"_"+df['app'].astype(str) \
+ " AXO" + df['app'].astype(str)+"_"+df['os'].astype(str) \
+ " IDHC" + df['ip_day_hour_count'].astype(str) \
+ " IAC" + df['ip_app_count'].astype(str) \
+ " AOC" + df['ip_app_os_count'].astype(str) \
+ " IDC" + df['ip_device_count'].astype(str) \
+ " AC" + df['app_channel_count'].astype(str) \
+ " NC" + df['next_click'].astype(str)
).values
#cpuStats()
if 'is_attributed' in df.columns:
labels = df['is_attributed'].values
weights = np.multiply([1.0 if x == 1 else 0.2 for x in df['is_attributed'].values],
df['hour'].apply(lambda x: 1.0 if x in pick_hours else 0.5))
else:
labels = []
weights = []
return str_array, labels, weights
# +
batchsize = 10000000
D = 2 ** 25
wb = wordbatch.WordBatch(
None,
extractor=(
WordHash,
{
"ngram_range": (1, 1),
"analyzer": "word",
"lowercase": False,
"n_features": D,
"norm": None,
"binary": True
}),
minibatch_size=batchsize // 80,
procs=8,
freeze=True,
timeout=1800,
verbose=0
)
clf = FM_FTRL(
alpha=0.05,
beta=0.1,
L1=0.0,
L2=0.0,
D=D,
alpha_fm=0.02,
L2_fm=0.0,
init_fm=0.01,
weight_fm=1.0,
D_fm=8,
e_noise=0.0,
iters=3,
inv_link="sigmoid",
e_clip=1.0,
threads=24,
use_avx=1,
verbose=0
)
# +
validate_filename = '../data/interim/train_2017-11-09_0400.csv'
df_val = pd.read_csv(validate_filename, engine='c', sep=",", dtype=dtypes)
str_array_val, labels_val, weights_val = df2csr(wb, df_val, pick_hours={4, 5, 10, 13, 14})
X_val = wb.transform(str_array_val)
del df_val, str_array_val
gc.collect()
# -
df = pd.read_hdf('../data/raw/train.hdf.compress')
str_array, y, weights= df2csr(wb, df, pick_hours={4, 5, 10, 13, 14})
with timer("transform"):
X = wb.transform(str_array)
with timer("fit_batch"):
clf.partial_fit(X, y, sample_weight=weights)
print("Train:")
with timer("evaluate_batch train"):
print(i, "ROC AUC:", roc_auc_score(y, clf.predict(X)))
|
notebooks/wordbatch-Copy1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1A.2 - Classes, méthodes, attributs, opérateurs et carré magique (correction)
#
# Correction.
from jyquickhelper import add_notebook_menu
add_notebook_menu()
# ### Exercice 1 : carré magique
# +
class CarreMagique :
def __init__(self, coef) :
self.mat = [ [ coef[i+j*3] for i in range(3) ] for j in range(3) ]
def __str__(self) :
return "\n".join ( [ ",".join( [ str(n) for n in row ] ) for row in self.mat ] )
def __add__ (self, carre) :
coef = []
for i in range(3) :
for j in range(3) :
coef.append ( self.mat[i][j] + carre.mat[i][j])
return CarreMagique(coef)
c = CarreMagique ( [ 1,3,4, 2,6,9, 8,7,5 ] )
print (c)
print("--")
print (c + c)
# -
# ### Exercice 2 : à faire à trois, carré magique (suite)
# +
class CarreMagique :
def __init__(self, coef) :
self.mat = [ [ coef[i+j*3] for i in range(3) ] for j in range(3) ]
def __str__(self) :
return "\n".join ( [ ",".join( str(n) for n in row ) for row in self.mat ] )
def __add__ (self, carre) :
coef = []
for i in range(3) :
for j in range(3) :
coef.append ( self.mat[i][j] + carre.mat[i][j])
return CarreMagique(coef)
def somme_ligne_colonne_diagonale(self):
tout = [ sum ( ligne ) for ligne in self.mat ] + \
[ sum ( self.mat[i][j] for i in range(3) ) for j in range(3) ] + \
[ sum ( self.mat[i][i] for i in range(3) ) ] + \
[ sum ( self.mat[2-i][i] for i in range(3) ) ]
return tout
def coefficient_unique(self):
d = { }
for ligne in self.mat :
for c in ligne :
d [c] = d.get(c,0) + 1
return len(d) == 9
def est_magique(self):
unique = self.coefficient_unique()
if not unique : return False
somme = self.somme_ligne_colonne_diagonale()
return min(somme) == max(somme)
c = CarreMagique ( [ 1,1,1, 1,1,1, 1,1,1 ] )
print (c.est_magique())
c = CarreMagique ( [ 1,4,8, 5,2,6, 7,9,3 ] )
print (c.est_magique())
c = CarreMagique ( [ 1,6,8, 7,5,3, 2,4,9 ] )
print (c.est_magique())
c = CarreMagique ( [ 2,7,6, 9,5,1, 4,3,8 ] )
print (c.est_magique())
# -
# ### Exercice 3 : trouver tous les carrés magiques
#
# La première version est fastidieuse à écrire mais simple à comprendre.
# +
def tous_les_carre_naif() :
res = []
for a1 in range(9) :
for a2 in range(9) :
for a3 in range(9) :
for b1 in range(9) :
for b2 in range(9) :
for b3 in range(9) :
for c1 in range(9) :
for c2 in range(9) :
for c3 in range(9) :
carre = CarreMagique( [a1,a2,a3, b1,b2,b3, c1,c2,c3 ])
if carre.est_magique() :
res.append (carre)
print (carre)
return res
# tous_les_carre_naif() (c'est très long)
# -
# La seconde version n'est pas plus rapide mais elle contient moins de boucles.
# +
def tous_les_carre_naif2() :
# on choisit l'ensemble de tous les tableaux de 9 chiffres compris entre 1 et 9
coef = [ 1 ] * 9
res = [ ]
while coef [0] < 10 :
carre = CarreMagique(coef)
if carre.est_magique() :
res.append (carre)
print (carre)
coef[-1] += 1
if coef[-1] >= 10 :
i = len(coef)-1
while coef[i] >= 10 and i > 0 :
coef[i] = 1
coef[i-1] += 1
i -= 1
# tous_les_carre_naif2() (c'est très long)
# -
# La troisième version utilise le fait que les chiffres d'un carré magique sont tous différents. Il suffit de regarder seulement tous les permutations. La variable ``stop_after`` permet de se limiter seulement aux premiers.
# +
def tous_les_carres_permutation( permut = None, pos = 0, stop_after = 3):
if pos == 9 :
carre = CarreMagique (permut)
if carre.est_magique() :
print (carre)
print ()
return [ carre ]
else :
return []
else :
res = [ ]
if permut == None :
permut = [ i+1 for i in range(9) ]
for i in range (pos,9) :
# on permute les éléments i et pos
a = permut[i]
permut[i] = permut[pos]
permut[pos] = a
res += tous_les_carres_permutation(permut, pos+1)
if stop_after > 0 and len(res) >= stop_after :
return res
# on effectue la permutation inverse
a = permut[i]
permut[i] = permut[pos]
permut[pos] = a
return res
res = tous_les_carres_permutation()
print ("nombre de carrés", len(res))
# -
# Le langage Python propose une fonction qui parcourt toutes les permutations d'un ensemble : [itertools.permutation](https://docs.python.org/3.4/library/itertools.html#itertools.permutations). Cela réduit de beaucoup la longueur du programme.
# +
import itertools
def tous_les_carres_permutation( stop_after = 3):
res = [ ]
firstn = list(range(1,10))
for permut in itertools.permutations(firstn) :
carre = CarreMagique (permut)
if carre.est_magique() :
res.append( carre )
if stop_after >= 0 :
print (carre)
print ()
if len(res) >= stop_after :
return res
return res
res = tous_les_carres_permutation()
print ("nombre de carrés", len(res))
# -
# ### Exercice 4 : faire plus rapide
# Est-il possible d'aller plus vite que de parcourir l'ensemble des permutations ? La réponse est oui. En parcourant les permutations, la fonction qui teste si les chiffres sont uniques est devenue inutile. Pour vérifier qu'on va plus vite, on peut mesurer le temps que met la fonction pour trouver tous les carrés :
import time
d = time.clock()
res = tous_les_carres_permutation(-1)
d = time.clock() - d
print ("nombre de carrés", len(res), " en ", d, "seconds")
# Pour aller plus vite, il faut utiliser la contrainte des sommes. Comment ? Lorsqu'on permute les nombres, on peut simplement vérifier que les deux premières lignes ont la même somme. L'utilisation de cette contrainte nous permet de d'aller 10 fois plus vite et d'obtenir le résultat en moins d'une seconde. L'inconvénient est que l'optimisation fonctionne parce qu'on ne parcourt pas toutes les permutations. On ne peut plus utiliser la fonction [itertools.permutation](https://docs.python.org/3.4/library/itertools.html#itertools.permutations).
# +
def tous_les_carres_permutation_ligne12_meme_somme( permut = None, pos = 0):
if pos == 9 :
carre = CarreMagique (permut)
if carre.est_magique() :
#print (carre)
#print ()
return [ carre ]
else :
return []
else :
if pos >= 6 : # ajout
if sum ( permut[:3]) != sum(permut[3:6]) : # ajout
return [ ] # ajout
res = [ ]
if permut == None :
permut = [ i+1 for i in range(9) ]
for i in range (pos,9) :
# on permute les éléments i et pos
a = permut[i]
permut[i] = permut[pos]
permut[pos] = a
res += tous_les_carres_permutation_ligne12_meme_somme(permut, pos+1) # changé
# on effectue la permutation inverse
a = permut[i]
permut[i] = permut[pos]
permut[pos] = a
return res
import time
d = time.clock()
res = tous_les_carres_permutation_ligne12_meme_somme()
d = time.clock() - d
print ("nombre de carrés", len(res), " en ", d)
# -
# ### Programme complet
# +
class CarreMagique :
def __init__(self, coef) :
self.mat = [ [ coef[i+j*3] for i in range(3) ] for j in range(3) ]
def __str__(self) :
return "\n".join ( [ ",".join( [ str(n) for n in row ] ) for row in self.mat ] )
def __add__ (self, carre) :
coef = []
for i in range(3) :
for j in range(3) :
coef.append ( self.mat[i][j] + carre.mat[i][j])
return CarreMagique(coef)
def somme_ligne_colonne_diagonale(self):
tout = [ sum ( ligne ) for ligne in self.mat ] + \
[ sum ( self.mat[i][j] for i in range(3) ) for j in range(3) ] + \
[ sum ( self.mat[i][i] for i in range(3) ) ] + \
[ sum ( self.mat[2-i][i] for i in range(3) ) ]
return tout
def coefficient_unique(self):
d = { }
for ligne in self.mat :
for c in ligne :
d [c] = d.get(c,0) + 1
return len(d) == 9
def est_magique(self):
unique = self.coefficient_unique()
if not unique : return False
somme = self.somme_ligne_colonne_diagonale()
return min(somme) == max(somme)
def tous_les_carres_permutation_ligne12_meme_somme( permut = None, pos = 0):
if pos == 9 :
carre = CarreMagique (permut)
if carre.est_magique() :
#print (carre)
#print ()
return [ carre ]
else :
return []
else :
if pos >= 6 : # ajout
if sum ( permut[:3]) != sum(permut[3:6]) : # ajout
return [ ] # ajout
res = [ ]
if permut == None :
permut = [ i+1 for i in range(9) ]
for i in range (pos,9) :
# on permute les éléments i et pos
a = permut[i]
permut[i] = permut[pos]
permut[pos] = a
res += tous_les_carres_permutation_ligne12_meme_somme(permut, pos+1) # changé
# on effectue la permutation inverse
a = permut[i]
permut[i] = permut[pos]
permut[pos] = a
return res
import time
d = time.clock()
res = tous_les_carres_permutation_ligne12_meme_somme()
d = time.clock() - d
print ("nombre de carrés", len(res), " en ", d)
# -
# On peut faire encore plus rapide en utilisant les contraintes pour inférer les autres coefficients (solution venant d'un élève) :
def tous_les_carres():
for a1 in range(1,10):
for a2 in range(1,10):
for a3 in range(1,10):
for b1 in range(1,10):
somme = a1 + a2 + a3
c1 = somme - a1 - b1
b2 = somme - a3 - c1
b3 = somme - b1 - b2
c2 = somme - a2 - b2
c3 = somme - c1 - c2
M = CarreMagique([a1,a2,a3,b1,b2,b3,c1,c2,c3])
if M.est_magique() and 0 < b2 < 10 and 0 < b3 < 10 and 0 < c1 < 10 and 0 < c2 < 10 and 0 < c3 < 10 :
#print(M)
#print("---------------")
pass
# %timeit tous_les_carres()
|
_doc/notebooks/td1a/td1a_correction_session5.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Collaborative filtering on the MovieLense Dataset
# ## Learning Objectives
# 1. Know how to explore the data using BigQuery
# 2. Know how to use the model to make recommendations for a user
# 3. Know how to use the model to recommend an item to a group of users
#
# ###### This notebook is based on part of Chapter 9 of [BigQuery: The Definitive Guide](https://www.oreilly.com/library/view/google-bigquery-the/9781492044451/ "http://shop.oreilly.com/product/0636920207399.do") by <NAME> Tigani.
# ### MovieLens dataset
# To illustrate recommender systems in action, let’s use the MovieLens dataset. This is a dataset of movie reviews released by GroupLens, a research lab in the Department of Computer Science and Engineering at the University of Minnesota, through funding by the US National Science Foundation.
#
# Download the data and load it as a BigQuery table using:
# +
import os
import tensorflow as tf
PROJECT = "qwiklabs-gcp-04-8722038efd75" # REPLACE WITH YOUR PROJECT ID
# Do not change these
os.environ["PROJECT"] = PROJECT
os.environ["TFVERSION"] = '2.1'
# + language="bash"
# rm -r bqml_data
# mkdir bqml_data
# cd bqml_data
# curl -O 'http://files.grouplens.org/datasets/movielens/ml-20m.zip'
# unzip ml-20m.zip
# yes | bq rm -r $PROJECT:movielens
# bq --location=US mk --dataset \
# --description 'Movie Recommendations' \
# $PROJECT:movielens
# bq --location=US load --source_format=CSV \
# --autodetect movielens.ratings ml-20m/ratings.csv
# bq --location=US load --source_format=CSV \
# --autodetect movielens.movies_raw ml-20m/movies.csv
# -
# ## Exploring the data
# Two tables should now be available in <a href="https://console.cloud.google.com/bigquery">BigQuery</a>.
#
# Collaborative filtering provides a way to generate product recommendations for users, or user targeting for products. The starting point is a table, <b>movielens.ratings</b>, with three columns: a user id, an item id, and the rating that the user gave the product. This table can be sparse -- users don’t have to rate all products. Then, based on just the ratings, the technique finds similar users and similar products and determines the rating that a user would give an unseen product. Then, we can recommend the products with the highest predicted ratings to users, or target products at users with the highest predicted ratings.
# +
# JOSE: we run this cell to avoid an error in the previous one
# %%bigquery
CREATE OR REPLACE TABLE movielens.ratings AS
SELECT * FROM `cloud-training-demos`.movielens.movielens_ratings;
CREATE OR REPLACE TABLE movielens.movies AS
SELECT * FROM `cloud-training-demos`.movielens.movielens_movies;
# -
# %%bigquery --project $PROJECT
SELECT *
FROM movielens.ratings
LIMIT 10
# A quick exploratory query yields that the dataset consists of over 138 thousand users, nearly 27 thousand movies, and a little more than 20 million ratings, confirming that the data has been loaded successfully.
# %%bigquery --project $PROJECT
SELECT
COUNT(DISTINCT userId) numUsers,
COUNT(DISTINCT movieId) numMovies,
COUNT(*) totalRatings
FROM movielens.ratings
# On examining the first few movies using the query following query, we can see that the genres column is a formatted string:
# %%bigquery --project $PROJECT
SELECT *
FROM movielens.movies_raw
WHERE movieId < 5
# We can parse the genres into an array and rewrite the table as follows:
# %%bigquery --project $PROJECT
CREATE OR REPLACE TABLE movielens.movies AS
SELECT * REPLACE(SPLIT(genres, "|") AS genres)
FROM movielens.movies_raw
# %%bigquery --project $PROJECT
SELECT *
FROM movielens.movies
WHERE movieId < 5
# ## Matrix factorization
# Matrix factorization is a collaborative filtering technique that relies on factorizing the ratings matrix into two vectors called the user factors and the item factors. The user factors is a low-dimensional representation of a user_id and the item factors similarly represents an item_id.
#
#
# %%bigquery --project $PROJECT
SELECT *
-- Note: remove cloud-training-demos if you are using your own model:
FROM ML.TRAINING_INFO(MODEL `cloud-training-demos.movielens.recommender`)
# What did you get? Our model took an hour to train, and the training loss starts out extremely bad and gets driven down to near-zero over next the four iterations:
#
# <table>
# <tr>
# <th>Iteration</th>
# <th>Training Data Loss</th>
# <th>Evaluation Data Loss</th>
# <th>Duration (seconds)</th>
# </tr>
# <tr>
# <td>4</td>
# <td>0.5734</td>
# <td>172.4057</td>
# <td>180.99</td>
# </tr>
# <tr>
# <td>3</td>
# <td>0.5826</td>
# <td>187.2103</td>
# <td>1,040.06</td>
# </tr>
# <tr>
# <td>2</td>
# <td>0.6531</td>
# <td>4,758.2944</td>
# <td>219.46</td>
# </tr>
# <tr>
# <td>1</td>
# <td>1.9776</td>
# <td>6,297.2573</td>
# <td>1,093.76</td>
# </tr>
# <tr>
# <td>0</td>
# <td>63,287,833,220.5795</td>
# <td>168,995,333.0464</td>
# <td>1,091.21</td>
# </tr>
# </table>
#
# However, the evaluation data loss is quite high, and much higher than the training data loss. This indicates that overfitting is happening, and so we need to add some regularization. Let’s do that next. Note the added l2_reg=0.2:
#THIS CELL WILL YIELD AN ERROR BECAUSE THE MODEL DOES NO LONGER EXIST
# %%bigquery --project $PROJECT
SELECT *
-- Note: remove cloud-training-demos if you are using your own model:
FROM ML.TRAINING_INFO(MODEL `cloud-training-demos.movielens.recommender_l2`)
# Now, we get faster convergence (three iterations instead of five), and a lot less overfitting. Here are our results:
#
# <table>
# <tr>
# <th>Iteration</th>
# <th>Training Data Loss</th>
# <th>Evaluation Data Loss</th>
# <th>Duration (seconds)</th>
# </tr>
# <tr>
# <td>2</td>
# <td>0.6509</td>
# <td>1.4596</td>
# <td>198.17</td>
# </tr>
# <tr>
# <td>1</td>
# <td>1.9829</td>
# <td>33,814.3017</td>
# <td>1,066.06</td>
# </tr>
# <tr>
# <td>0</td>
# <td>481,434,346,060.7928</td>
# <td>2,156,993,687.7928</td>
# <td>1,024.59</td>
# </tr>
# </table>
#
# By default, BigQuery sets the number of factors to be the log2 of the number of rows. In our case, since we have 20 million rows in the table, the number of factors would have been chosen to be 24. As with the number of clusters in K-Means clustering, this is a reasonable default but it is often worth experimenting with a number about 50% higher (36) and a number that is about a third lower (16):
#
#
# %%bigquery --project $PROJECT
SELECT *
-- Note: remove cloud-training-demos if you are using your own model:
FROM ML.TRAINING_INFO(MODEL `cloud-training-demos.movielens.recommender_16`)
# When we did that, we discovered that the evaluation loss was lower (0.97) with num_factors=16 than with num_factors=36 (1.67) or num_factors=24 (1.45). We could continue experimenting, but we are likely to see diminishing returns with further experimentation.
#
# ## Making recommendations
#
# With the trained model, we can now provide recommendations. For example, let’s find the best comedy movies to recommend to the user whose userId is 903. In the query below, we are calling ML.PREDICT passing in the trained recommendation model and providing a set of movieId and userId to carry out the predictions on. In this case, it’s just one userId (903), but all movies whose genre includes Comedy.
# %%bigquery --project $PROJECT
SELECT * FROM
ML.PREDICT(MODEL `cloud-training-demos.movielens.recommender_16`, (
SELECT
movieId, title, 903 AS userId
FROM movielens.movies, UNNEST(genres) g
WHERE g = 'Comedy'
))
ORDER BY predicted_rating DESC
LIMIT 5
# ## Filtering out already rated movies
# Of course, this includes movies the user has already seen and rated in the past. Let’s remove them.
#
# **TODO 1**: Make a prediction for user 903 that does not include already seen movies.
# %%bigquery --project $PROJECT
SELECT * FROM
ML.PREDICT(MODEL `cloud-training-demos.movielens.recommender_16`, (
WITH seen AS (
SELECT ARRAY_AGG(movieId) AS movies
FROM movielens.ratings
WHERE userId = 903
)
SELECT
movieId, title, 903 AS userId
FROM movielens.movies, UNNEST(genres) g, seen
WHERE
g = 'Comedy' AND movieId NOT IN UNNEST(seen.movies)
# TODO: Complete this WHERE to remove seen movies.
))
ORDER BY predicted_rating DESC
LIMIT 5
# For this user, this happens to yield the same set of movies -- the top predicted ratings didn’t include any of the movies the user has already seen.
#
# ## Customer targeting
#
# In the previous section, we looked at how to identify the top-rated movies for a specific user. Sometimes, we have a product and have to find the customers who are likely to appreciate it. Suppose, for example, we wish to get more reviews for movieId = 96481 (American Mullet) which has only one rating and we wish to send coupons to the 5 users who are likely to rate it the highest.
#
# **TODO 2**: Find the top five users who will likely enjoy *American Mullet (2001)*
# %%bigquery --project $PROJECT
SELECT * FROM
ML.PREDICT(MODEL `cloud-training-demos.movielens.recommender_16`, (
WITH allUsers AS (
SELECT DISTINCT userId
FROM movielens.ratings
)
SELECT
96481 AS movieId,
(SELECT title FROM movielens.movies WHERE movieId=96481) title,
userId
FROM
allUsers
# TODO: Select all users
))
ORDER BY predicted_rating DESC
LIMIT 5
# ### Batch predictions for all users and movies
# What if we wish to carry out predictions for every user and movie combination? Instead of having to pull distinct users and movies as in the previous query, a convenience function is provided to carry out batch predictions for all movieId and userId encountered during training. A limit is applied here, otherwise, all user-movie predictions will be returned and will crash the notebook.
# %%bigquery --project $PROJECT
SELECT *
FROM ML.RECOMMEND(MODEL `cloud-training-demos.movielens.recommender_16`)
LIMIT 10
# As seen in a section above, it is possible to filter out movies the user has already seen and rated in the past. The reason already seen movies aren’t filtered out by default is that there are situations (think of restaurant recommendations, for example) where it is perfectly expected that we would need to recommend restaurants the user has liked in the past.
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
|
notebooks/recommendation_systems/labs/2_als_bqml.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Merge & Concat
#
# En muchas ocasiones nos podemos encontrar con que los conjuntos de datos no se encuentran agregados en una única tabla. Cuando esto sucede, existen dos formas para unir la información de distintas tablas: **merge** y **concat**.
# ## Concat
#
# La función `concat()` realiza todo el trabajo pesado de realizar operaciones de concatenación a lo largo de un eje mientras realiza la lógica de conjunto opcional (unión o intersección) de los índices (si los hay) en los otros ejes. Tenga en cuenta que digo "si hay alguno" porque solo hay un único eje posible de concatenación para Series.
#
# ### Concatenar varias tablas con las mismas columnas
import os
import numpy as pd
import pandas as pd
# +
df1 = pd.DataFrame({'A': ['A0', 'A1', 'A2', 'A3'],
'B': ['B0', 'B1', 'B2', 'B3'],
'C': ['C0', 'C1', 'C2', 'C3'],
'D': ['D0', 'D1', 'D2', 'D3']},
index=[0, 1, 2, 3])
df2 = pd.DataFrame({'A': ['A4', 'A5', 'A6', 'A7'],
'B': ['B4', 'B5', 'B6', 'B7'],
'C': ['C4', 'C5', 'C6', 'C7'],
'D': ['D4', 'D5', 'D6', 'D7']},
index=[4, 5, 6, 7])
df3 = pd.DataFrame({'A': ['A8', 'A9', 'A10', 'A11'],
'B': ['B8', 'B9', 'B10', 'B11'],
'C': ['C8', 'C9', 'C10', 'C11'],
'D': ['D8', 'D9', 'D10', 'D11']},
index=[8, 9, 10, 11])
frames = [df1, df2, df3]
result = pd.concat(frames)
# -
#
# <img src="./images/merge_01.png" align="center"/>
#
# ### Concatenar varias tablas con distintas columnas (por filas)
# +
df4 = pd.DataFrame({'B1': ['hola', 'B3', 'B6', 'B7'],
'D': ['D2', 'D3', 'D6', 'D7'],
'F': ['F2', 'F3', 'F6', 'F7']},
index=[2, 3, 6, 7])
result = pd.concat([df1, df4], axis=0, sort=False)
# -
# <img src="./images/merge_02.png" align="center"/>
#
# ### Concatenar varias tablas con distintas columnas (por columnas)
result = pd.concat([df1, df4], axis=1, sort=False)
# <img src="./images/merge_03.png" align="center"/>
#
# ## Merge
#
# La función `merge()` se usa para combinar dos (o más) tablas sobre valores de columnas comunes (keys).
# +
left = pd.DataFrame({'key': ['K0', 'K1', 'K2', 'K3'],
'A': ['A0', 'A1', 'A2', 'A3'],
'B': ['B0', 'B1', 'B2', 'B3']})
right = pd.DataFrame({'key': ['K0', 'K1', 'K2', 'K3'],
'C': ['C0', 'C1', 'C2', 'C3'],
'D': ['D0', 'D1', 'D2', 'D3']})
result = pd.merge(left, right, on='key')
# -
# <img src="./images/merge_04.png" align="center"/>
#
# En este ejemplo, se especifica en la opción `on` las columnas (keys) donde se realizará el cruce de información de ambas tablas.
# ### Tipos de merge
#
# La opción *how* especificica el tipo de cruce que se realizará.
#
# * **left**: usa las llaves solo de la tabla izquierda
# * **right**: usa las llaves solo de la tabla derecha
# * **outer**: usa las llaves de la unión de ambas tablas.
# * **inner**: usa las llaves de la intersección de ambas tablas.
#
# <img src="images/joins2.png" width="480" height="480" align="center"/>
#
#
#
# +
left = pd.DataFrame({'key1': ['K0', 'K0', 'K1', 'K2'],
'key2': ['K0', 'K1', 'K0', 'K1'],
'A': ['A0', 'A1', 'A2', 'A3'],
'B': ['B0', 'B1', 'B2', 'B3']})
right = pd.DataFrame({'key1': ['K0', 'K1', 'K1', 'K2'],
'key2': ['K0', 'K0', 'K0', 'K0'],
'C': ['C0', 'C1', 'C2', 'C3'],
'D': ['D0', 'D1', 'D2', 'D3']})
# -
# #### Merge left
merge_left = pd.merge(left, right,how= 'left', on=['key1', 'key2'])
# <img src="images/merge_05.png" align="center"/>
#
# #### Merge right
merge_rigth = pd.merge(left, right, how='right', on=['key1', 'key2'])
# <img src="images/merge_06.png" align="center"/>
#
# #### Merge outer
merge_outer = pd.merge(left, right, how='outer', on=['key1', 'key2'])
# <img src="images/merge_07.png" align="center"/>
#
# #### Merge inner
merge_inner = pd.merge(left, right, how='inner', on=['key1', 'key2'])
# <img src="images/merge_08.png" align="center"/>
#
# ### Problemas de llaves duplicadas
#
# Cuando se quiere realizar el cruce de dos tablas, pero an ambas tablas existe una columna (key) con el mismo nombre, para diferenciar la información entre la columna de una tabla y otra, pandas devulve el nombre de la columna con un guión bajo x (key_x) y otra con un guión bajo y (key_y)
# +
left = pd.DataFrame({'A': [1, 2], 'B': [2, 2]})
right = pd.DataFrame({'A': [4, 5, 6], 'B': [2, 2, 2]})
result = pd.merge(left, right, on='B', how='outer')
# -
# <img src="images/merge_09.png" align="center"/>
#
# ## Referencia
#
# 1. [Merge, join, and concatenate](https://pandas.pydata.org/pandas-docs/stable/user_guide/merging.html)
#
|
lectures/data_manipulation/data_manipulation/modulos_pandas/merge_concat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Figure(s) in the manuscript created by this notebook: Fig.4C, 3D, 3E.
#
# This notebook fits and plots FRAP data both from clustered proteins and diffuse (unclustered) proteins. The data that this notebook parses comes from the outputs of the "Extract_two_radii_TrackMate.ijm" and "Manual_FRAP_ROI.ijm" ImageJ macros.
# +
# User-defined parameters for analysis:
# Plotting and figure saving params
save_figs = True
save_dir = '../reports/figures/Fig4C-F_FRAP'
plot_settings = '../src/plotting_settings.py'
# Source data metadata
frame_interval = 1 # in seconds
bleach_n_frames = 8 # how many frames bleaching takes (max)
# Source data location
# Path to diffuse FRAP data (proteins diffusing freely, outside of large clusters)
# Output of "Manual_FRAP_ROI.ijm" ImageJ macro
data_dir_noclust = '../data/processed/Fig4C-F_IRE1_FRAP/manual_ROI_FRAP_non-clustered/intensities'
# Path to cluster FRAP data (text files saved by "Extract_two_radii_TrackMate.ijm")
data_dir = '../data/processed/Fig4C-F_IRE1_FRAP/spot_radii'
# The excluded trace file allows you to manually remove bad traces from the analysis.
# It should be a simple csv with each line containing the file name followed by trace ID
excluded_trace_file = '../data/processed/Fig4C-F_IRE1_FRAP/2018-10-11_excluded_traces.csv'
# +
# load the built-in and custom modules
# uncomment for debugging
"""
%load_ext autoreload
%autoreload 2
"""
import os, sys, inspect
import matplotlib
import matplotlib.pylab as plt
from scipy import stats
import numpy as np
from pprint import pprint
import glob
import pandas as pd
import seaborn as sns
# Add source code directory (src) to path to enable module import
module_dir = '../src'
os.sys.path.insert(0, module_dir)
# import custom modules
import fraptools as frap
import diffusion as dif
# +
# Set up figure save dirs and load plotting style
if save_figs:
# %matplotlib
# %run $plot_settings save
# Make directories for saving figures
save_dir_clust = os.path.join(save_dir, 'clusters')
if not os.path.exists(save_dir_clust):
os.makedirs(save_dir_clust)
save_dir_diffuse = os.path.join(save_dir, 'diffuse')
if not os.path.exists(save_dir_diffuse):
os.makedirs(save_dir_diffuse)
save_dir_summary = os.path.join(save_dir, 'summary')
if not os.path.exists(save_dir_summary):
os.makedirs(save_dir_summary)
else:
# %matplotlib inline
# %run $plot_settings plot_only
# +
# Load FRAP data from clusters
# Populate excluded trace file
if excluded_trace_file:
excluded_files_df = pd.read_csv(excluded_trace_file)
excluded_traces = [tuple(x) for x in excluded_files_df.values]
else:
excluded_traces = (None, None)
# Generate list of all valid cluster FRAP files
frap_files = sorted(glob.glob(os.path.join(data_dir,'*.txt')))
# list of all FRAP data:
frap_data_by_file = []
filenames_no_ext = []
# Go file by file and read data
for file in frap_files:
# Read data from the provided source file
data = pd.read_csv(file, delimiter='\t')
filename_no_ext = os.path.split(os.path.splitext(file)[0])[1]
frap_data_by_file.append(data)
filenames_no_ext.append(filename_no_ext)
# Extract individual traces from the raw data
df_by_trace, corr_ints, trace_IDs = frap.get_traces_from_df_list(frap_data_by_file,
filenames_no_ext, exclude=excluded_traces)
# +
# Load FRAP data from non-clustered proteins
frap_files_noclust = sorted(glob.glob(os.path.join(data_dir_noclust,'*.csv')))
# list of all FRAP data:
frap_data_by_file_noclust = []
filenames_no_ext_noclust = []
# Go file by file and read data
for file in frap_files_noclust:
# Read data from the provided source file
data = pd.read_csv(file, delimiter=',')
filename_no_ext = os.path.split(os.path.splitext(file)[0])[1]
frap_data_by_file_noclust.append(data)
filenames_no_ext_noclust.append(filename_no_ext)
# break up data into smaller data frames, one per trace
df_by_trace_noclust, corr_ints_noclust, trace_IDs_noclust = \
frap.read_nonclust_frap_data(frap_data_by_file_noclust,
filenames_no_ext_noclust, exclude=excluded_traces)
# +
# Analyze and plot the FRAP data from clusters
# Fit the individual FRAP traces
fit, data = frap.fit_frap_smart(corr_ints, frame_interval, bleach_n_frames)
# Plot results
for f,d,trace_ID in zip(fit, data, trace_IDs):
file_name = trace_ID[0]
trace_num = trace_ID[1]
full_name = file_name + '_trace-ID_' + str(trace_num)
fig, axarr = frap.plot_fit_results(f,d)
fig.canvas.set_window_title(full_name)
plt.suptitle(full_name)
if save_figs:
fig_filename_pdf = os.path.join(save_dir_clust, (full_name+'.pdf'))
plt.savefig(fig_filename_pdf)
plt.close(fig)
print("done")
# +
# Analyze and plot FRAP data from diffuse (non-clustered) proteins
# Fit the individual FRAP traces
fit_noclust, data_noclust = frap.fit_frap_smart(corr_ints_noclust, frame_interval, bleach_n_frames)
# Plot results
for f,d,trace_ID in zip(fit_noclust, data_noclust, trace_IDs_noclust):
file_name = trace_ID[0]
trace_num = trace_ID[1]
full_name = file_name + '_trace-ID_' + str(trace_num)
fig, axarr = frap.plot_fit_results(f,d)
fig.canvas.set_window_title(full_name)
plt.suptitle(full_name)
if save_figs:
fig_filename_pdf = os.path.join(save_dir_diffuse, (full_name+'.pdf'))
plt.savefig(fig_filename_pdf)
plt.close()
print("Processed trace ID", trace_ID)
print("done")
# +
# Summarize fit results
# Prepare data for plotting
frap_fits_foci = fit
thalf_foci = [f['thalf'] for f in frap_fits_foci]
mobile_f_foci = [f['mobile_fraction'] for f in frap_fits_foci]
thalf_noclust = [f['thalf'] for f in fit_noclust]
mobile_f_noclust = [f['mobile_fraction'] for f in fit_noclust]
print(np.mean(thalf_foci), np.mean(mobile_f_foci), stats.sem(mobile_f_foci))
print(np.mean(thalf_noclust), np.mean(mobile_f_noclust))
print('t-test for thalf: ', stats.ttest_ind(thalf_foci, thalf_noclust))
# Create summary figure
fig, axes = plt.subplots(nrows=1, ncols=2, figsize=(2.6, 1.7))
fig.tight_layout(pad=2)
df1 = pd.DataFrame({'Free' : thalf_noclust})
df2 = pd.DataFrame({'Clustered' : thalf_foci})
df_thalf = pd.concat([df1, df2], axis = 1)
df3 = pd.DataFrame({'Free' : mobile_f_noclust})
df4 = pd.DataFrame({'Clustered' : mobile_f_foci})
df_mobile = pd.concat([df3, df4], axis = 1)
sns.boxplot(data=df_thalf, linewidth=0.5, showfliers = False,
boxprops={'facecolor':'None'}, ax=axes[0])
sns.swarmplot(data=df_thalf, zorder=0.5, ax=axes[0], size=2)
axes[0].set_ylabel('Half-time of recovery (s)')
sns.boxplot(data=df_mobile, linewidth=0.5, showfliers = False,
boxprops={'facecolor':'None'}, ax=axes[1])
sns.swarmplot(data=df_mobile, zorder=0.5, ax=axes[1], size=2)
axes[1].set_ylabel('Mobile fraction')
if save_figs:
fig_filename_pdf = os.path.join(save_dir_summary, 'Clusters_vs_noClusters_recovery.pdf')
plt.savefig(fig_filename_pdf)
plt.show()
# +
# Estimate diffusion constant (from doi:10.1111/tra.12008)
rn = 5 # In microns, effective radius of the bleach spot (st.dev. of the Gaussian beam)
d_foci = [0.25*rn**2/t for t in thalf_foci]
d_er = [0.25*rn**2/t for t in thalf_noclust]
print(np.mean(d_foci), stats.sem(d_foci))
print(np.mean(d_er), stats.sem(d_er))
# Estimate diffusion using Guigas-Weiss model
T = 310 # 37C in Kelvin
c = 6 # in nm; 4 for slip, 6 for stick boundary conditions
ire1_radius = 0.7 # estimated in-plane radius of IRE1 in nm
print('Estimated IRE1 D, um^2/s: ', dif.diffconst_gw(ire1_radius,T,c))
print('Saffman-Delbruck IRE1 D, um^2/s: ', dif.diffconst_sd(ire1_radius,T,c))
|
notebooks/04_FRAP-with-and-without-clustering.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: jsl250
# language: python
# name: jsl250
# ---
# + [markdown] colab_type="text" id="sQFJ6aT4nfhw"
# [](https://colab.research.google.com/github/JohnSnowLabs/spark-nlp-workshop/blob/master/jupyter/enterprise/healthcare/ChunkMergeClinicalMultiple.ipynb)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="MdE588BiY3z1" outputId="cec54ac1-ef24-46cb-e856-a192a94ad900"
import json
with open('keys.json') as f:
license_keys = json.load(f)
license_keys.keys()
# + colab={"base_uri": "https://localhost:8080/", "height": 408} colab_type="code" id="FVFdvGChZDDP" outputId="0c692a99-059b-4aa1-ffee-464add2e5503"
import os
# Install java
# ! apt-get update
# ! apt-get install -y openjdk-8-jdk-headless -qq > /dev/null
os.environ["JAVA_HOME"] = "/usr/lib/jvm/java-8-openjdk-amd64"
os.environ["PATH"] = os.environ["JAVA_HOME"] + "/bin:" + os.environ["PATH"]
# ! java -version
secret = license_keys.get("secret",license_keys.get('SPARK_NLP_SECRET', ""))
os.environ['SPARK_NLP_LICENSE'] = license_keys['SPARK_NLP_LICENSE']
os.environ['JSL_OCR_LICENSE'] = license_keys['JSL_OCR_LICENSE']
os.environ['AWS_ACCESS_KEY_ID']= license_keys['AWS_ACCESS_KEY_ID']
os.environ['AWS_SECRET_ACCESS_KEY'] = license_keys['AWS_SECRET_ACCESS_KEY']
version = license_keys.get("version",license_keys.get('SPARK_NLP_PUBLIC_VERSION', ""))
jsl_version = license_keys.get("jsl_version",license_keys.get('SPARK_NLP_VERSION', ""))
# ! python -m pip install pyspark
# ! python -m pip install --upgrade spark-nlp-jsl==$jsl_version --extra-index-url https://pypi.johnsnowlabs.com/$secret -qq
import sparknlp
import sparknlp_jsl
from sparknlp.base import *
from sparknlp.annotator import *
from sparknlp_jsl.annotator import *
import pyspark
from pyspark.ml import Pipeline
from pyspark.sql import SparkSession
print (sparknlp.version())
print (sparknlp_jsl.version())
spark = sparknlp_jsl.start(secret, gpu=False, spark23=False)
# + colab={} colab_type="code" id="1zgsiTxjaiMd"
# Sample data. If really training a new NER from the output of multiple prertained NERs,
# this data should have thousands of documents
data_chunk_merge = spark.createDataFrame([
(1,"""A 63-year-old man presents to the hospital with a history of recurrent infections that include cellulitis, pneumonias, and upper respiratory tract infections. He reports subjective fevers at home along with unintentional weight loss and occasional night sweats. The patient has a remote history of arthritis, which was diagnosed approximately 20 years ago and treated intermittently with methotrexate (MTX) and prednisone. On physical exam, he is found to be febrile at 102°F, rather cachectic, pale, and have hepatosplenomegaly. Several swollen joints that are tender to palpation and have decreased range of motion are also present. His laboratory values show pancytopenia with the most severe deficiency in neutrophils.
""")]).toDF("id","text")
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="weY5V9h7ZDf0" outputId="60599668-3ee6-4011-caf5-d6f991224668"
# Preprocessing pipeline
da = DocumentAssembler().setInputCol("text").setOutputCol("document")
sd = SentenceDetector().setInputCols("document").setOutputCol("sentence")
tk = Tokenizer().setInputCols("sentence").setOutputCol("token")
emb = WordEmbeddingsModel.pretrained("embeddings_clinical","en","clinical/models").setOutputCol("embs")
# + colab={} colab_type="code" id="ku85AU_Rnfir"
# Ners names and column names for each ner
from collections import OrderedDict
ners_to_merge = OrderedDict({"ner_deid_large":"deid", "ner_bionlp":"bio", "ner_jsl":"jsl"})
# + colab={"base_uri": "https://localhost:8080/", "height": 170} colab_type="code" id="7LYvNOPZnfi2" outputId="979cf279-9363-48a0-a702-9654c7283dda"
# Build the graph
ner_pl = []
for ner, out in ners_to_merge.items():
first = len(ner_pl)==0
ner_pl.append(NerDLModel.pretrained(ner,"en","clinical/models").setInputCols("sentence","token","embs").setOutputCol(out))
ner_pl.append(NerConverter().setInputCols("sentence","token",out).setOutputCol(out+"_chunk"))
if not first:
ner_pl.append(ChunkMergeApproach().setInputCols(prev+"_chunk", out+"_chunk").setOutputCol(out+"_chunk"))
prev = out
out_col = list(ners_to_merge.values())[-1]+"_chunk"
# + colab={} colab_type="code" id="2-3bBy3yP88k"
iob_tagger = IOBTagger().setInputCols("token", out_col).setOutputCol("ner_label")
# + colab={} colab_type="code" id="JMI6QDcunfjI"
pl = Pipeline().setStages([da,sd,tk,emb] + ner_pl + [iob_tagger])
# + colab={} colab_type="code" id="K9lX8WOwnfjP"
merged_data = pl.fit(data_chunk_merge).transform(data_chunk_merge).cache()
# + colab={"base_uri": "https://localhost:8080/", "height": 527} colab_type="code" id="dpLba4tAbPiW" outputId="9bdcf461-ede4-4f1c-be4f-150dbb946a91"
merged_data.selectExpr("id",f"explode({out_col}) as a")\
.selectExpr("id","a.begin","a.end","a.result as chunk","a.metadata.entity as entity").show(100, False)
# + colab={"base_uri": "https://localhost:8080/", "height": 629} colab_type="code" id="NYsZ3KsCnw0y" outputId="3e51ab7f-e51a-4950-b88f-172f6b743995"
#As we can see, ner_label column is ready to train a NerDLApproach
merged_data.selectExpr("id",f"explode(ner_label) as a")\
.selectExpr("id","a.begin","a.end","a.result as chunk","a.metadata.word as word").where("chunk!='O'").show(1000, False)
# + colab={} colab_type="code" id="9KIz9vFoRgp0"
|
jupyter/enterprise/healthcare/NER_from_NERs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (tensorflow)
# language: python
# name: rga
# ---
# # T81-558: Applications of Deep Neural Networks
# **Module 2: Python for Machine Learning**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# # Module 2 Material
#
# Main video lecture:
#
# * Part 2.1: Introduction to Pandas [[Video]](https://www.youtube.com/watch?v=bN4UuCBdpZc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_1_python_pandas.ipynb)
# * **Part 2.2: Categorical Values** [[Video]](https://www.youtube.com/watch?v=4a1odDpG0Ho&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_2_pandas_cat.ipynb)
# * Part 2.3: Grouping, Sorting, and Shuffling in Python Pandas [[Video]](https://www.youtube.com/watch?v=YS4wm5gD8DM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_3_pandas_grouping.ipynb)
# * Part 2.4: Using Apply and Map in Pandas for Keras [[Video]](https://www.youtube.com/watch?v=XNCEZ4WaPBY&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_4_pandas_functional.ipynb)
# * Part 2.5: Feature Engineering in Pandas for Deep Learning in Keras [[Video]](https://www.youtube.com/watch?v=BWPTj4_Mi9E&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_02_5_pandas_features.ipynb)
# # Part 2.2: Categorical and Continuous Values
#
# Neural networks require their input to be a fixed number of columns. This is very similar to spreadsheet data. This input must be completely numeric.
#
# It is important to represent the data in a way that the neural network can train from it. In class 6, we will see even more ways to preprocess data. For now, we will look at several of the most basic ways to transform data for a neural network.
#
# Before we look at specific ways to preprocess data, it is important to consider four basic types of data, as defined by [Stanley Smith Stevens](https://en.wikipedia.org/wiki/Stanley_Smith_Stevens). These are commonly referred to as the [levels of measure](https://en.wikipedia.org/wiki/Level_of_measurement):
#
# * Character Data (strings)
# * **Nominal** - Individual discrete items, no order. For example: color, zip code, shape.
# * **Ordinal** - Individual discrete items that can be ordered. For example: grade level, job title, Starbucks(tm) coffee size (tall, vente, grande)
# * Numeric Data
# * **Interval** - Numeric values, no defined start. For example, temperature. You would never say "yesterday was twice as hot as today".
# * **Ratio** - Numeric values, clearly defined start. For example, speed. You would say that "The first car is going twice as fast as the second."
# ### Encoding Continuous Values
#
# One common transformation is to normalize the inputs. It is sometimes valuable to normalization numeric inputs to be put in a standard form so that two values can easily be compared. Consider if a friend told you that he received a $10 discount. Is this a good deal? Maybe. But the value is not normalized. If your friend purchased a car, then the discount is not that good. If your friend purchased dinner, this is a very good discount!
#
# Percentages are a very common form of normalization. If your friend tells you they got 10% off, we know that this is a better discount than 5%. It does not matter how much the purchase price was. One very common machine learning normalization is the Z-Score:
#
# $z = \frac{x - \mu}{\sigma} $
#
# To calculate the Z-Score you need to also calculate the mean($\mu$) and the standard deviation ($\sigma$). The mean is calculated as follows:
#
# $\mu = \bar{x} = \frac{x_1+x_2+\cdots +x_n}{n}$
#
# The standard deviation is calculated as follows:
#
# $\sigma = \sqrt{\frac{1}{N} \sum_{i=1}^N (x_i - \mu)^2}, {\rm \ \ where\ \ } \mu = \frac{1}{N} \sum_{i=1}^N x_i$
#
# The following Python code replaces the mpg with a z-score. Cars with average MPG will be near zero, above zero is above average, and below zero is below average. Z-Scores above/below -3/3 are very rare, these are outliers.
# +
import os
import pandas as pd
from scipy.stats import zscore
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/auto-mpg.csv",
na_values=['NA','?'])
df['mpg'] = zscore(df['mpg'])
display(df[0:5])
# -
# ### Encoding Categorical Values as Dummies
# The classic means of encoding categorical values is to make them dummy variables. This is also called one-hot-encoding. Consider the following data set.
# +
import pandas as pd
df = pd.read_csv(
"https://data.heatonresearch.com/data/t81-558/jh-simple-dataset.csv",
na_values=['NA','?'])
display(df[0:5])
# -
areas = list(df['area'].unique())
print(f'Number of areas: {len(areas)}')
print(f'Areas: {areas}')
# There are four unique values in the areas column. To encode these to dummy variables we would use four columns, each of which would represent one of the areas. For each row, one column would have a value of one, the rest zeros. This is why this type of encoding is sometimes called one-hot encoding. The following code shows how you might encode the values "a" through "d". The value A becomes [1,0,0,0] and the value B becomes [0,1,0,0].
dummies = pd.get_dummies(['a','b','c','d'],prefix='area')
print(dummies)
# To encode the "area" column, we use the following.
# It is necessary to merge these dummies back into the data frame.
dummies = pd.get_dummies(df['area'],prefix='area')
print(dummies[0:10]) # Just show the first 10
df = pd.concat([df,dummies],axis=1)
# Displaying select columns from the dataset we can see the dummy variables added.
display(df[0:10][['id','job','area','income','area_a',
'area_b','area_c','area_d']])
# Usually, you will remove the original column ('area'), because it is the goal to get the dataframe to be entirely numeric for the neural network.
df.drop('area', axis=1, inplace=True)
display(df[0:10][['id','job','income','area_a',
'area_b','area_c','area_d']])
# ### Target Encoding for Categoricals
#
# Target encoding can sometimes increase the predictive power of a machine learning model. However, it also greatly increases the risk of overfitting. Because of this risk, care must be take if you are going to use this method. It is a popular technique for Kaggle competitions.
#
# Generally, target encoding can only be used on a categorical feature when the output of the machine learning model is numeric (regression).
#
# The concept of target encoding is actually very simple. For each value
# +
# Create a small sample dataset
import pandas as pd
import numpy as np
np.random.seed(43)
df = pd.DataFrame({
'cont_9': np.random.rand(10)*100,
'cat_0': ['dog'] * 5 + ['cat'] * 5,
'cat_1': ['wolf'] * 9 + ['tiger'] * 1,
'y': [1, 0, 1, 1, 1, 1, 0, 0, 0, 0]
})
display(df)
# -
# Rather than creating dummy variables for dog and cat, we would like to change it to a number. We could just use 0 for cat, 1 for dog. However, we can encode more information than just that. The simple 0 or 1 would also only work for one animal. Consider what the mean target value is for cat and dog.
means0 = df.groupby('cat_0')['y'].mean().to_dict()
means0
# The danger is that we are now using the target value for training. This will potentially overfit. The possibility of overfitting is even greater if there are a small number of a particular category. To prevent this from happening, we use a weighting factor. The stronger the weight the more than categories with a small number of values will tend towards the overall average of y, which is calculated as follows.
df['y'].mean()
# The complete function for target encoding is given here.
# Source: https://maxhalford.github.io/blog/target-encoding-done-the-right-way/
def calc_smooth_mean(df1, df2, cat_name, target, weight):
# Compute the global mean
mean = df[target].mean()
# Compute the number of values and the mean of each group
agg = df.groupby(cat_name)[target].agg(['count', 'mean'])
counts = agg['count']
means = agg['mean']
# Compute the "smoothed" means
smooth = (counts * means + weight * mean) / (counts + weight)
# Replace each value by the according smoothed mean
if df2 is None:
return df1[cat_name].map(smooth)
else:
return df1[cat_name].map(smooth),df2[cat_name].map(smooth.to_dict())
# The following code encodes these two categories.
WEIGHT = 5
df['cat_0_enc'] = calc_smooth_mean(df1=df, df2=None, cat_name='cat_0', target='y', weight=WEIGHT)
df['cat_1_enc'] = calc_smooth_mean(df1=df, df2=None, cat_name='cat_1', target='y', weight=WEIGHT)
display(df)
# ### Encoding Categorical Values as Ordinal
#
# Typically categoricals will be encoded as dummy variables. However, there might be other techniques to convert categoricals to numeric. Any time there is an order to the categoricals, a number should be used. Consider if you had a categorical that described the current education level of an individual.
#
# * Kindergarten (0)
# * First Grade (1)
# * Second Grade (2)
# * Third Grade (3)
# * Fourth Grade (4)
# * Fifth Grade (5)
# * Sixth Grade (6)
# * Seventh Grade (7)
# * Eighth Grade (8)
# * High School Freshman (9)
# * High School Sophomore (10)
# * High School Junior (11)
# * High School Senior (12)
# * College Freshman (13)
# * College Sophomore (14)
# * College Junior (15)
# * College Senior (16)
# * Graduate Student (17)
# * PhD Candidate (18)
# * Doctorate (19)
# * Post Doctorate (20)
#
# The above list has 21 levels. This would take 21 dummy variables. However, simply encoding this to dummies would lose the order information. Perhaps the easiest approach would be to assign simply number them and assign the category a single number that is equal to the value in parenthesis above. However, we might be able to do even better. Graduate student is likely more than a year, so you might increase more than just one value.
|
t81_558_class_02_2_pandas_cat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Assignment 02 - The simplest model order selection
#
# The assignment extends Assignment 01 by adding two ingredients:
# 1. considering not only Least Squares but also Maximum Likelihood
# 2. adding the possibility of choosing different model orders
# # task 2.1
#
# Consider the same situation of tasks 1.1 and 1.2 in assignment 1, but with the additional possibility of ‘faulty measurements’ in the following sense: with a probability $\gamma$ the sensor returns something meaningless, i.e., with probability $\gamma$ $e_t$ is so that $y_t = 100$. With probability $1 - \gamma$, instead, the sensor works fine and thus the same probabilistic model for generating $e_t$ applies. Note: the outlier events are i.i.d., i.e., the fact that one has an outlier at time $t$ does not modify the probability of having an outlier also at time $t+1$.
# importing the right packages
import numpy as np
import matplotlib.pyplot as plt
import scipy.optimize as optimize
# define the function that creates an arbitrary polynomial
# that we will use also to create the scalar quadratic
# 'true model'
#
# note that this function uses both 'lambda functions' and
# the 'enumerate' function -- two features of python
def arbitrary_poly(theta):
noiseless_y = lambda x: sum([p*(x**i) for i, p in enumerate(theta)])
return noiseless_y
# +
# Non-random generation between code executions.
# Comment out if you want to repeat the same experiment over and over again
# np.random.seed(123)
# -
# define the function for the mixture distribution
def noise(gamma, alpha, beta, sigma, N):
# generate the noises as before
if np.random.uniform() < alpha:
noise = np.random.normal(0, sigma, N)
else:
noise = np.random.laplace(0, beta, N)
# use \infty as a placeholder for the iid events 'faulty measurement'
for i in range(N):
if np.random.uniform() < gamma:
noise[i] = np.inf
return noise
# # task 2.2
#
# With respect to assignment 1, though, modify the $u_t$'s so to be 201 uniformly spaced values in $[-3, 3]$, and set $\alpha = 0$ so that we have Laplacian noises (more precisely, with scale $\beta = 1$). Keep the "true model" quadratic.
# +
# for readability
min_u = -3
max_u = 3
N = 201 # number of samples
# create the u_t's
u = np.linspace(min_u, max_u, N)
# set the required parameters
theta_true = [1, -8, 4]
gamma = 0.2
beta = 1
sigma = 1
alpha = 0
# create the noisy dataset
noiseless_y = arbitrary_poly(theta_true) # use the lambda function
y = noiseless_y( u ) + noise( gamma, alpha, beta, sigma, len(u) )
# remember to account for the 'faulty measurements'!
y[np.isinf(y)] = 100
# plot the dataset, for debugging purposes
x = np.linspace(start = min_u, stop = max_u, num = 100)
plt.figure()
plt.scatter(u, y, label = 'dataset')
plt.plot(x, noiseless_y(x), 'k:', label = 'true model')
plt.xlabel('u')
plt.ylabel('y')
plt.legend();
# -
# # task 2.3
#
# Code both a Least Squares and a Maximum Likelihood estimator whose underlying assumption on the measurement noise is Laplacian.
#
# Add also the possibility of testing generic model structures (i.e., affine, quadratic, cubic, and so-on models).
# define the function solving the LS problem asked in the assignment
def LS_solver( u, y, model_order ):
# create the features matrix
U = u[:, np.newaxis] ** np.arange( model_order )
# compute the LS estimate by directly solving the normal equations
theta_hat = np.linalg.solve(U.T @ U, U.T @ y)
return theta_hat
def laplace_log_likelihood(theta, u, y):
# probability distribution p, "know" beta = 1 but leave for expm.
beta = 1
U = u[:, np.newaxis] ** np.arange(len(theta))
mu = theta@U.T
negative_log_likelihood = -sum(np.log((1.0/(2*beta)*np.exp(-np.abs(y - mu)/beta))))
return negative_log_likelihood
# define the function solving the ML problem asked in the assignment
def ML_solver( u, y, model_order ):
# compute the ML estimate by directly minimizing the log likelihood
opt_res = optimize.minimize(
fun = laplace_log_likelihood,
x0 = np.zeros( model_order ),
args = (u, y),
options = {'disp': False} )
# save the result in a more readable format
theta_hat = opt_res.x
# debug
#print('estimated parameters ML = ', theta_hat)
return theta_hat
# # task 2.4
#
# Split the dataset so that the first half is the training set, the second 25% is the test set, and the last 25% is the validation set.
#
# Code a function that returns the "performance index" of a generic estimate $\widehat{\theta}$ on a generic dataset $y$. The index shall be equal to the sum of the absolute deviations between the actually measured $y$'s and the predicted ones $\widehat{y}_t$, i.e.,
#
# $$
# \widehat{y}_t
# =
# \left[ \widehat{\theta}_0, \widehat{\theta}_1, \widehat{\theta}_2, \ldots \right]
# \left[ 1, u_t, u_t^2, \ldots \right]^T .
# $$
# split the dataset
training_u = u[:100]
training_y = y[:100]
#
testing_u = u[101:150]
testing_y = y[101:150]
#
validating_u = u[151:]
validating_y = y[151:]
# define the performance index
def performance_index( measured_y, estimated_y ):
return sum([abs(y - y_hat) for y, y_hat in zip(measured_y, estimated_y)])
# +
# DEBUG
# check that everything works
# set a model order, just to check that things work
guessed_model_order = 3
# solve the LS and ML problems
theta_hat_LS = LS_solver( training_u, training_y, guessed_model_order )
theta_hat_ML = ML_solver( training_u, training_y, guessed_model_order )
# compute the estimated models
modeled_y_LS = arbitrary_poly(theta_hat_LS) # use the lambda function
modeled_y_ML = arbitrary_poly(theta_hat_ML) # use the lambda function
# plot the true model against the measurements and the estimated model
x = np.linspace(start = min_u, stop = max_u, num = 100)
plt.figure()
plt.scatter(u, y, label = 'dataset')
plt.plot(x, noiseless_y(x), 'k:', label = 'true model')
plt.plot(x, modeled_y_LS(x), 'b', label = 'estimated model LS')
plt.plot(x, modeled_y_ML(x), 'r', label = 'estimated model ML')
plt.xlabel('u')
plt.ylabel('y')
plt.legend();
# print the performance indexes
print("PI = Performance Index - lower is better")
print( 'PI LS on training set: ', performance_index( training_y, modeled_y_LS(training_u) ) )
print( 'PI ML on training set: ', performance_index( training_y, modeled_y_ML(training_u) ) )
print( 'PI LS on test set: ', performance_index( testing_y, modeled_y_LS(testing_u) ) )
print( 'PI ML on test set: ', performance_index( testing_y, modeled_y_ML(testing_u) ) )
# -
# # task 2.5
#
# Comment the plot above. Why are these behaviours happening?
# The least-squares estimator is purely geometric, and does not rely on any assumptions about the data set and its distribution. The estimator thus struggles with the outliers, since they are weighed equally when minimizing the distance between the dataset and the thetas in the manifold of the assumed model.
#
# The maximum likelihood estimator however, does utilize assumptions about the distribution of the data. Here we assume laplacian noise (which is true), although we do not directly account for the outliers per say. However, the effect that the outliers have on the resulting estimate is greatly reduced since we "believe" less in the outliers as compared to the LS approach, and thus they affect the resulting thetas less.
# # task 2.6
#
# Create a table summarizing how the performance indexes vary as a function of the model order of the estimator on the training and test sets.
# +
# choose the max model order
max_model_order = 5
ls_pis = []
ml_pis = []
# cycle on the various model orders
for current_order in range(1, max_model_order+1):
# train on the training set
theta_hat_LS = LS_solver( training_u, training_y, current_order )
theta_hat_ML = ML_solver( training_u, training_y, current_order )
# create the estimated models
modeled_y_LS = arbitrary_poly(theta_hat_LS) # use the lambda function
modeled_y_ML = arbitrary_poly(theta_hat_ML) # use the lambda function
# predict the test set
predicted_testing_y_LS = modeled_y_LS(testing_u)
predicted_testing_y_ML = modeled_y_ML(testing_u)
# compute the performance indexes on the predictive capabilities on the test set
PI_LS = performance_index(predicted_testing_y_LS, testing_y)
PI_ML = performance_index(predicted_testing_y_ML, testing_y)
ls_pis.append(PI_LS)
ml_pis.append(PI_ML)
for o, pi in enumerate(ls_pis):
# print debug info
print(f"PI for LS w. order {o+1}: {pi}")
print("\n\n")
for o, pi in enumerate(ml_pis):
print(f"PI for ML w. order {o+1}: {pi}")
# -
# # task 2.7
#
# Rerun the code for different values of $\gamma$, and check how the performance capabilities of the LS and ML estimators on the training set vary. Explicitly say:
#
# 1. Which estimator seems more robust, in average? And why?
# 2. Which estimator is so that its performance degrade least gracefully with increasing $\gamma$? And why?
# 3. Which model order would you choose, given the simulations that you have run? And why?
# 1: Thinking of the term "robustness" as "less prone to error given changes in the model and/or dataset" then I would consider the ML estimator as more robust than the LS estimator. This is reasonable given that we, as the programmer in the driver seat, have added laplacian noise, and are correctly assuming that the model is corrupted by the laplacian noise. Because of the same arguments as above, our "belief" of the laplacian distribution leads to the outliers affecting the estimator less.
#
# 2: I would argue that the ML estimator degrades least gracefully, since it suddenly "jumps" to following the outliers at a certain point. The LS estimator performs worse over most gammas, although the performance doesn't jump around. This happens since, at a certain point, the outliers "win" over our assumption that the data is laplacian distributed - "winning" as in, the outliers are more prevalent in the dataset than the modelled data, such that the argmax of the distribution of the data describes the outliers themselves, and not the model generating the actual data.
#
# 3: I would choose order 3 or 4 since the performance for the ML estimator across most gammas is the best for these orders. I prioritize the ML estimator performing well over the LS estimator due to the ML estimator being more robut/a better fit for this problem (as it is defined at least).
|
assignment_02.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
1+1
20/3
20.0/3
type (20.0 / 3)
# # Let's load SageMath's hooks
# %load_ext sage
1+1
type(20.0/3)
# # Algebraic Structures
ZZ, QQ, RR, CC
Zmod(30)
GF(31)
# # Most algebraic structures are organised as parent/elements
GF(17).list()
a = 2
a.parent()
ZZ is a.parent()
a.is_unit()
# # Conversions
b = QQ(a)
b == a
b.parent()
b.is_unit()
# # Working with notebooks
a, b = 2, 3
c = a + b
c
# # Exploring objects
#
# Completion
ZZ.<tab>
# Documentation
# +
# ZZ.CartesianProduct?
# +
# ZZ.cardinality??
# -
# # Polynomial rings
a = x^2 - 1
b = (x-1)*(x+1)
b
a == b
bool(a == b)
# ## x is just a variable name
x = 2
x^2 - 1
# ## Forget about x!
# We don't need symbolic calculus, we need polynomials
# We can reset symbolic variables with `x = SR.var('x')`
QQ['x']
R.<x> = QQ[]
R
a = x^2 -1
a
a.parent()
b = (x-1)*(x+1)
b
a == b
# # Finite fields
p = next_prime(2^20)
R.<x> = GF(p)[]
R
P = R.irreducible_element(20)
P
K.<z> = GF(p^20, modulus=P)
K
z^20
# ## If we don't care about the modulus
L.<z> = GF(p^2)
L
L.modulus()
z.multiplicative_order()
z^3
e = L.primitive_element()
e.multiplicative_order()
e
e.minpoly()
# ## Elliptic Curves
E = EllipticCurve([1,2])
E
E.j_invariant()
E.torsion_order()
E.torsion_points()
G = E.torsion_subgroup()
G
G.gens()
# ## Elliptic curves over finite fields
E = EllipticCurve(GF(p), [0, 1, 0, 2, 3])
E
E.cardinality().factor()
P = E.random_point()
Q = E.lift_x(123)
P, Q
P+Q
Q.order().factor()
R = 9*23*241*Q
R
# ## Isogenies
phi = E.isogeny([R])
phi
S = phi(P)
S
S.parent()
phi(R)
phi.rational_maps()
phi.kernel_polynomial()
phi.kernel_polynomial()(R[0])
# ## Pairings
L.<z> = GF(p^2)
EE = E.change_ring(L)
EE
EE.cardinality().factor()
T = EE.lift_x(912851*z + 87136)
T.order()
w = T.weil_pairing(EE(R), 3)
w
w.multiplicative_order()
T.weil_pairing(2*T, 3)
|
SummerSchool/SageTutorial/WelcomeNotebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="V8-yl-s-WKMG"
# # EfficientDet Tutorial: inference, eval, and training
#
#
#
# <table align="left"><td>
# <a target="_blank" href="https://github.com/google/automl/blob/master/efficientdet/tutorial.ipynb">
# <img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on github
# </a>
# </td><td>
# <a target="_blank" href="https://colab.sandbox.google.com/github/google/automl/blob/master/efficientdet/tutorial.ipynb">
# <img width=32px src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td></table>
# + [markdown] id="muwOCNHaq85j" colab_type="text"
# # 0. Install and view graph.
# + [markdown] colab_type="text" id="dggLVarNxxvC"
# ## 0.1 Install package and download source code/image.
#
#
# + colab_type="code" id="hGL97-GXjSUw" colab={}
# %%capture
#@title
# Install tensorflow and pycocotools
# !pip install tensorflow
# !pip install pytype
# The default pycocotools doesn't work for python3: https://github.com/cocodataset/cocoapi/issues/49
# !pip install -U 'git+https://github.com/cocodataset/cocoapi.git#subdirectory=PythonAPI'
import os
import sys
import tensorflow.compat.v1 as tf
# Download source code.
if "efficientdet" not in os.getcwd():
# !git clone --depth 1 https://github.com/google/automl
os.chdir('automl/efficientdet')
sys.path.append('.')
else:
# !git pull
# + id="Tow-ic7H3d7i" colab_type="code" outputId="be86cdc5-682d-4aff-b9f4-04d836ba8c1f" colab={"base_uri": "https://localhost:8080/", "height": 221}
MODEL = 'efficientdet-d0' #@param
def download(m):
if m not in os.listdir():
# !wget https://storage.googleapis.com/cloud-tpu-checkpoints/efficientdet/coco/{m}.tar.gz
# !tar zxf {m}.tar.gz
ckpt_path = os.path.join(os.getcwd(), m)
return ckpt_path
# Download checkpoint.
ckpt_path = download(MODEL)
print('Use model in {}'.format(ckpt_path))
# Prepare image and visualization settings.
image_url = 'https://user-images.githubusercontent.com/11736571/77320690-099af300-6d37-11ea-9d86-24f14dc2d540.png'#@param
image_name = 'img.png' #@param
# !wget {image_url} -O img.png
import os
img_path = os.path.join(os.getcwd(), 'img.png')
min_score_thresh = 0.4 #@param
max_boxes_to_draw = 200 #@param
line_thickness = 2#@param
import PIL
# Get the largest of height/width and round to 128.
image_size = max(PIL.Image.open(img_path).size)
# + [markdown] id="GvdjcYpUVuQ5" colab_type="text"
# ## 0.2 View graph in TensorBoard
# + id="U2oz3r1LUDzr" colab_type="code" colab={}
# !python model_inspect.py --model_name={MODEL} --logdir=logs &> /dev/null
# %load_ext tensorboard
# %tensorboard --logdir logs
# + [markdown] id="vZk2dwOxrGhY" colab_type="text"
# # 1. inference
# + [markdown] id="_VaF_j7jdVCK" colab_type="text"
# ## 1.1 Benchmark networrk latency
# There are two types of latency:
# network latency and end-to-end latency.
#
# * network latency: from the first conv op to the network class and box prediction.
# * end-to-end latency: from image preprocessing, network, to the final postprocessing to generate a annotated new image.
#
# + id="R_3gL01UbDLH" colab_type="code" outputId="7fcf42cf-e751-4ec3-b33b-41ce4f36898c" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# benchmaak network latency
# !python model_inspect.py --runmode=bm --model_name=efficientdet-d3
# With colab + Tesla T4 GPU, here are the batch size 1 latency summary:
# D0: 14.9ms, FPS = 67.2 (batch size 8 FPS=92.8)
# D1: 29.0ms, FPS = 34.4 (batch size 8 FPS=41.6)
# D2: 43.2ms, FPS = 23.1 (batch size 8 FPS=27.2)
# D3: 76.7ms, FPS = 13.0 (batch size 8 FPS=14.4)
# + [markdown] id="VW95IodKovEu" colab_type="text"
# ## 1.2 Benchmark end-to-end latency
# + id="NSf6SrZcdavN" colab_type="code" outputId="e4ce89f2-8a0c-43ba-8e06-364dc3ca44f7" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Benchmark end-to-end latency (: preprocess + network + posprocess).
#
# With colab + Tesla T4 GPU, here are the batch size 1 latency summary:
# D0 (AP=33.5): 23.8ms, FPS = 42.1 (batch size 4, FPS=79.3)
# D1 (AP=39.6): 36.1ms, FPS = 27.7 (batch size 4, FPS=39.1)
# D2 (AP=43.0): 50.7ms, FPS = 19.7 (batch size 4, FPS=26.0)
# D3 (AP=45.8): 84.6ms, FPS = 11.8 (batch size 4, FPS=13.3)
# D4 (AP=49.4): 140ms, FPS = 7.1 (batch size 4, FPS=7.5)
# D5 (AP=50.7): 298ms, FPS = 3.6
# D6 (AP=51.7): 386ms, FPS = 2.6
m = 'efficientdet-d0' # @param
batch_size = 1# @param
m_path = download(m)
saved_model_dir = 'savedmodel'
# !rm -rf {saved_model_dir}
# !python model_inspect.py --runmode=saved_model --model_name={m} \
# --ckpt_path={m_path} --saved_model_dir={saved_model_dir} \
# --batch_size={batch_size}
# !python model_inspect.py --runmode=saved_model_benchmark --model_name={m} \
# --ckpt_path={m_path} --saved_model_dir={saved_model_dir} \
# --batch_size={batch_size} --input_image=testdata/img1.jpg
# + [markdown] id="jGKs3w2_ZXnu" colab_type="text"
# ## 1.3 Inference images.
#
# ---
#
#
# + id="tlh_S6M9ahe5" colab_type="code" outputId="420baca8-7863-41cb-ed78-7aca8b9e5919" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# first export a saved model.
saved_model_dir = 'savedmodel'
# !rm -rf {saved_model_dir}
# !python model_inspect.py --runmode=saved_model --model_name={MODEL} \
# --ckpt_path={ckpt_path} --saved_model_dir={saved_model_dir}
# Then run saved_model_infer to do inference.
# Notably: batch_size, image_size must be the same as when it is exported.
serve_image_out = 'serve_image_out'
# !mkdir {serve_image_out}
# !python model_inspect.py --runmode=saved_model_infer \
# --saved_model_dir={saved_model_dir} \
# --model_name={MODEL} --input_image=testdata/img1.jpg \
# --output_image_dir={serve_image_out}
# + id="1q2x8s8GpUJz" colab_type="code" colab={}
from IPython import display
display.display(display.Image(os.path.join(serve_image_out, '0.jpg')))
# + id="fHU46tfckaZo" colab_type="code" outputId="415aebc8-5a9b-40f4-a074-b847f14982ef" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# In case you need to specify different image size or batch size or #boxes, then
# you need to export a new saved model and run the inferernce.
serve_image_out = 'serve_image_out'
# !mkdir {serve_image_out}
saved_model_dir = 'savedmodel'
# !rm -rf {saved_model_dir}
# Step 1: export model
# !python model_inspect.py --runmode=saved_model \
# --model_name=efficientdet-d0 --ckpt_path=efficientdet-d0 \
# --hparams="image_size=1920x1280" --saved_model_dir={saved_model_dir}
# Step 2: do inference with saved model.
# !python model_inspect.py --runmode=saved_model_infer \
# --model_name=efficientdet-d0 --ckpt_path=efficientdet-d0 \
# --hparams="image_size=1920x1280" --saved_model_dir={saved_model_dir} \
# --input_image=img.png --output_image_dir={serve_image_out}
from IPython import display
display.display(display.Image(os.path.join(serve_image_out, '0.jpg')))
# + [markdown] id="Vxm-kvfuAZne" colab_type="text"
# ## 1.4 Inference video
# + id="3Pdnd1kQAgKY" colab_type="code" outputId="acb01fa7-c2da-4f6f-efa3-273ef1371fd2" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# step 0: download video
video_url = 'https://storage.googleapis.com/cloud-tpu-checkpoints/efficientdet/data/video480p.mov' # @param
# !wget {video_url} -O input.mov
# Step 1: export model
saved_model_dir = 'savedmodel'
# !rm -rf {saved_model_dir}
# !python model_inspect.py --runmode=saved_model \
# --model_name=efficientdet-d0 --ckpt_path=efficientdet-d0 \
# --saved_model_dir={saved_model_dir}
# Step 2: do inference with saved model using saved_model_video
# !python model_inspect.py --runmode=saved_model_video \
# --model_name=efficientdet-d0 --ckpt_path=efficientdet-d0 \
# --saved_model_dir={saved_model_dir} \
# --input_video=input.mov --output_video=output.mov
# Then you can view the output.mov
# + [markdown] id="RW26DwfirQQN" colab_type="text"
# # 3. COCO evaluation
# + [markdown] colab_type="text" id="cfn_tRFOWKMO"
# ## 3.1 COCO evaluation on validation set.
# + id="2s6E8IsVN0pB" colab_type="code" outputId="6799da8b-6637-4a11-97c8-18b1d22c6eb1" colab={"base_uri": "https://localhost:8080/", "height": 1000}
if 'val2017' not in os.listdir():
# !wget http://images.cocodataset.org/zips/val2017.zip
# !wget http://images.cocodataset.org/annotations/annotations_trainval2017.zip
# !unzip -q val2017.zip
# !unzip annotations_trainval2017.zip
# !mkdir tfrecrod
# !PYTHONPATH=".:$PYTHONPATH" python dataset/create_coco_tfrecord.py \
# --image_dir=val2017 \
# --caption_annotations_file=annotations/captions_val2017.json \
# --output_file_prefix=tfrecord/val \
# --num_shards=32
# + id="eLHZUY3jQpZr" colab_type="code" outputId="6e8c30eb-3d16-4985-c5bb-b44d9e385c21" colab={"base_uri": "https://localhost:8080/", "height": 1000}
# Evalute on validation set (takes about 10 mins for efficientdet-d0)
# !python main.py --mode=eval \
# --model_name={MODEL} --model_dir={ckpt_path} \
# --validation_file_pattern=tfrecord/val* \
# --val_json_file=annotations/instances_val2017.json \
# --use_tpu=False
# + [markdown] id="mDp_acD1pUcx" colab_type="text"
# ## 3.2 COCO evaluation on test-dev.
# + id="9RI_dvx5pbBK" colab_type="code" colab={}
# Eval on test-dev is slow (~40 mins), please be cautious.
RUN_EXPENSIVE_TEST_DEV_EVAL = True #@param
if RUN_EXPENSIVE_TEST_DEV_EVAL == True:
# !rm *.zip *.tar tfrecord/ val2017/ # Cleanup disk space
# Download and convert test-dev data.
if "test2017" not in os.listdir():
# !wget http://images.cocodataset.org/zips/test2017.zip
# !unzip -q test2017.zip
# !wget http://images.cocodataset.org/annotations/image_info_test2017.zip
# !unzip image_info_test2017.zip
# !mkdir tfrecrod
# !PYTHONPATH=".:$PYTHONPATH" python dataset/create_coco_tfrecord.py \
# --image_dir=test2017 \
# --image_info_file=annotations/image_info_test-dev2017.json \
# --output_file_prefix=tfrecord/testdev \
# --num_shards=32
# Evalute on validation set: non-empty testdev_dir is the key pararmeter.
# Also, test-dev has 20288 images rather than val 5000 images.
# !mkdir testdev_output
# !python main.py --mode=eval \
# --model_name={MODEL} --model_dir={ckpt_path} \
# --validation_file_pattern=tfrecord/testdev* \
# --use_tpu=False --eval_batch_size=8 \
# --testdev_dir='testdev_output' --eval_samples=20288
# !rm -rf test2017 # delete images to release disk space.
# Now you can submit testdev_output/detections_test-dev2017_test_results.json to
# coco server: https://competitions.codalab.org/competitions/20794#participate
# + [markdown] id="RW90fiMiyg4n" colab_type="text"
# # 4. Training EfficientDets on PASCAL.
# + [markdown] id="C98Ye0MEyuKD" colab_type="text"
# ## 4.1 Prepare data
# + id="6PC6QrMlylOF" colab_type="code" colab={}
# Get pascal voc 2012 trainval data
import os
if 'VOCdevkit' not in os.listdir():
# !wget http://host.robots.ox.ac.uk/pascal/VOC/voc2012/VOCtrainval_11-May-2012.tar
# !tar xf VOCtrainval_11-May-2012.tar
# !mkdir tfrecord
# !PYTHONPATH=".:$PYTHONPATH" python dataset/create_pascal_tfrecord.py \
# --data_dir=VOCdevkit --year=VOC2012 --output_path=tfrecord/pascal
# Pascal has 5717 train images with 100 shards epoch, here we use a single shard
# for demo, but users should use all shards pascal-*-of-00100.tfrecord.
file_pattern = 'pascal-00000-of-00100.tfrecord' # @param
images_per_epoch = 57 * len(tf.io.gfile.glob('tfrecord/' + file_pattern))
images_per_epoch = images_per_epoch // 8 * 8 # round to 64.
print('images_per_epoch = {}'.format(images_per_epoch))
# + [markdown] id="ZcxDDCCW0ndv" colab_type="text"
# ## 4.2 Train Pascal VOC 2012 from ImageNet checkpoint for Backbone.
# + id="SHPgm9Q13X-l" colab_type="code" colab={}
# Train efficientdet from scratch with backbone checkpoint.
backbone_name = {
'efficientdet-d0': 'efficientnet-b0',
'efficientdet-d1': 'efficientnet-b1',
'efficientdet-d2': 'efficientnet-b2',
'efficientdet-d3': 'efficientnet-b3',
'efficientdet-d4': 'efficientnet-b4',
'efficientdet-d5': 'efficientnet-b5',
'efficientdet-d6': 'efficientnet-b6',
'efficientdet-d7': 'efficientnet-b6',
}[MODEL]
# generating train tfrecord is large, so we skip the execution here.
import os
if backbone_name not in os.listdir():
# !wget https://storage.googleapis.com/cloud-tpu-checkpoints/efficientnet/ckptsaug/{backbone_name}.tar.gz
# !tar xf {backbone_name}.tar.gz
# !mkdir /tmp/model_dir
# key option: use --backbone_ckpt rather than --ckpt.
# Don't use ema since we only train a few steps.
# !python main.py --mode=train_and_eval \
# --training_file_pattern=tfrecord/{file_pattern} \
# --validation_file_pattern=tfrecord/{file_pattern} \
# --val_json_file=tfrecord/json_pascal.json \
# --model_name={MODEL} \
# --model_dir=/tmp/model_dir/{MODEL}-scratch \
# --backbone_ckpt={backbone_name} \
# --train_batch_size=8 \
# --eval_batch_size=8 --eval_samples={images_per_epoch} \
# --num_examples_per_epoch={images_per_epoch} --num_epochs=1 \
# --hparams="num_classes=20,moving_average_decay=0" \
# --use_tpu=False
# + [markdown] id="SKHu-3lBwTiM" colab_type="text"
# ## 4.3 Train Pascal VOC 2012 from COCO checkpoint for the whole net.
# + id="SD59rsZJc1WW" colab_type="code" colab={}
# generating train tfrecord is large, so we skip the execution here.
import os
if MODEL not in os.listdir():
# !wget https://storage.googleapis.com/cloud-tpu-checkpoints/efficientdet/coco/{MODEL}.tar.gz
# !tar xf {MODEL}.tar.gz
# !mkdir /tmp/model_dir/
# key option: use --ckpt rather than --backbone_ckpt.
# !python main.py --mode=train_and_eval \
# --training_file_pattern=tfrecord/{file_pattern} \
# --validation_file_pattern=tfrecord/{file_pattern} \
# --val_json_file=tfrecord/json_pascal.json \
# --model_name={MODEL} \
# --model_dir=/tmp/model_dir/{MODEL}-finetune \
# --ckpt={MODEL} \
# --train_batch_size=8 \
# --eval_batch_size=8 --eval_samples={images_per_epoch} \
# --num_examples_per_epoch={images_per_epoch} --num_epochs=1 \
# --hparams="num_classes=20,moving_average_decay=0" \
# --use_tpu=False
# + [markdown] id="QcBGPMCXRC8q" colab_type="text"
# ## 4.4 View tensorboard for loss and accuracy.
#
# + id="Vrkty06SRD0k" colab_type="code" colab={}
# %load_ext tensorboard
# %tensorboard --logdir /tmp/model_dir/
# Notably, this is just a demo with almost zero accuracy due to very limited
# training steps, but we can see finetuning has smaller loss than training
# from scratch at the begining.
|
efficientdet/tutorial.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: ipykernel_py2
# ---
# ## Lists
# Create a list, called "Numbers". Let it contain the numbers 10, 25, 40, and 50.
# Print the second element from the list.
# Print the 0th element.
# Print the third-to-last element using a minus sign in the brackets.
# Substitute the number 10 with the number 15.
# Delete the number 25 from the Numbers list.
|
Python for Finance - Code Files/33 Lists/Python 2/Lists - Exercise_Py2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="Za8-Nr5k11fh"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" id="Eq10uEbw0E4l"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="_3bi1D2IiCyW"
# # Naive forecasting
# + [markdown] id="m_6H00ELiA57"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c02_naive_forecasting.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/examples/blob/master/courses/udacity_intro_to_tensorflow_for_deep_learning/l08c02_naive_forecasting.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] id="vidayERjaO5q"
# ## Setup
# + id="gqWabzlJ63nL" executionInfo={"status": "ok", "timestamp": 1608980368610, "user_tz": -60, "elapsed": 937, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08618602917479372870"}}
import numpy as np
import matplotlib.pyplot as plt
# + id="sJwA96JU00pW" executionInfo={"status": "ok", "timestamp": 1608980369072, "user_tz": -60, "elapsed": 1391, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08618602917479372870"}}
def plot_series(time, series, format="-", start=0, end=None, label=None):
plt.plot(time[start:end], series[start:end], format, label=label)
plt.xlabel("Time")
plt.ylabel("Value")
if label:
plt.legend(fontsize=14)
plt.grid(True)
def trend(time, slope=0):
return slope * time
def seasonal_pattern(season_time):
"""Just an arbitrary pattern, you can change it if you wish"""
return np.where(season_time < 0.4,
np.cos(season_time * 2 * np.pi),
1 / np.exp(3 * season_time))
def seasonality(time, period, amplitude=1, phase=0):
"""Repeats the same pattern at each period"""
season_time = ((time + phase) % period) / period
return amplitude * seasonal_pattern(season_time)
def white_noise(time, noise_level=1, seed=None):
rnd = np.random.RandomState(seed)
return rnd.randn(len(time)) * noise_level
# + [markdown] id="yVo6CcpRaW7u"
# ## Trend and Seasonality
# + id="BLt-pLiZ0nfB" colab={"base_uri": "https://localhost:8080/", "height": 388} executionInfo={"status": "ok", "timestamp": 1608980373944, "user_tz": -60, "elapsed": 1157, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08618602917479372870"}} outputId="59bd1dcb-d6eb-42d1-abdb-8ed88697d3ba"
time = np.arange(4 * 365 + 1)
slope = 0.05
baseline = 10
amplitude = 40
series = baseline + trend(time, slope) + seasonality(time, period=365, amplitude=amplitude)
noise_level = 5
noise = white_noise(time, noise_level, seed=42)
series += noise
plt.figure(figsize=(10, 6))
plot_series(time, series)
plt.show()
# + [markdown] id="a1sQpPjhtj0G"
# All right, this looks realistic enough for now. Let's try to forecast it. We will split it into two periods: the training period and the validation period (in many cases, you would also want to have a test period). The split will be at time step 1000.
# + id="_w0eKap5uFNP" executionInfo={"status": "ok", "timestamp": 1608980378271, "user_tz": -60, "elapsed": 897, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08618602917479372870"}}
split_time = 1000
time_train = time[:split_time]
x_train = series[:split_time]
time_valid = time[split_time:]
x_valid = series[split_time:]
# + [markdown] id="bjD8ncEZbjEW"
# ## Naive Forecast
# + id="Pj_-uCeYxcAb" executionInfo={"status": "ok", "timestamp": 1608980386429, "user_tz": -60, "elapsed": 859, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08618602917479372870"}}
naive_forecast = series[split_time - 1:-1]
# + id="JtxwHj9Ig0jT" colab={"base_uri": "https://localhost:8080/", "height": 388} executionInfo={"status": "ok", "timestamp": 1608980389130, "user_tz": -60, "elapsed": 1257, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08618602917479372870"}} outputId="e86434e1-bbd3-4941-f56f-47e8b2db7920"
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid, label="Series")
plot_series(time_valid, naive_forecast, label="Forecast")
# + [markdown] id="fw1SP5WeuixH"
# Let's zoom in on the start of the validation period:
# + id="D0MKg7FNug9V" colab={"base_uri": "https://localhost:8080/", "height": 388} executionInfo={"status": "ok", "timestamp": 1608980583272, "user_tz": -60, "elapsed": 1325, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08618602917479372870"}} outputId="733e40cc-c975-428a-b4fe-6497978c44e3"
plt.figure(figsize=(10, 6))
plot_series(time_valid, x_valid, start=0, end=150, label="Series")
plot_series(time_valid, naive_forecast, start=1, end=151, label="Forecast")
# + [markdown] id="35gIlQLfu0TT"
# You can see that the naive forecast lags 1 step behind the time series.
# + [markdown] id="Uh_7244Gsxfx"
# Now let's compute the mean absolute error between the forecasts and the predictions in the validation period:
# + id="LjpLQeWY11H8" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1608980588746, "user_tz": -60, "elapsed": 892, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "08618602917479372870"}} outputId="64e3cf99-593c-4777-cfbc-af5a58d8585b"
errors = naive_forecast - x_valid
abs_errors = np.abs(errors)
mae = abs_errors.mean()
mae
# + [markdown] id="WGPBC9QttI1u"
# That's our baseline, now let's try a moving average.
|
Copia di l08c02_naive_forecasting.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from urllib.request import urlopen
from bs4 import BeautifulSoup
import requests
import re
def no_space(text):
text1 = re.sub(" | |\n|\t|\r","",text)
text2 = re.sub("\n\n","",text1)
return text2
def get_text(url):
item_name_list=[]
item_price_list=[]
try:
html= urlopen(url)
bs= BeautifulSoup(html, "html.parser")
title = bs.findAll('p', {"class":"list_info"}, recurisive=False)
for i in title:
text = no_space(i.get_text())
item_name_list.append(text)
print(text)
price = bs.findAll('p', {"class":"price"})
for i in price:
text2 = no_space(i.get_text())
item_price_list.append(text2)
print(text2)
except:
print("실패")
def get_title(url):
html= urlopen(url)
bs= BeautifulSoup(html, "html.parser")
i=0
item_name_list=[]
for item_name in bs.find_all('p', {"class":"list_info"}):
item_name_list.append(no_space(item_name.get_text()))
return item_name_list
# +
#실제 카테고리별 번호는 001010, 002003 ...
CategoryId_list = [
# 상의 0~7
['반팔 티셔츠', 1001],
['셔츠/블라우스', 1002],
['피케/카라 티셔츠', 1003],
['후드 티셔츠', 1004],
['맨투맨/스웨트셔츠',1005],
['니트/스웨터', 1006],
['긴팔 티셔츠', 1010],
['민소매 티셔츠', 1011],
# 아우터 8~15
['슈트/블레이저', 2003],
['후드 집업', 2022],
['블루종/MA-1', 2001],
['환절기 코트', 2008],
['래더/라이더스 재킷', 2002],
['숏패딩/숏헤비 아우터', 2012],
['롱패딩/롱헤비 아우터', 2013],
['기타 아우터', 2015],
# 하의 16~21
['데님 팬츠',3002],
['슈트 팬츠/슬랙스', 3008],
['코튼 팬츠', 3007],
['트레이닝/조거 팬츠',3004],
['숏 팬츠', 3009],
['레깅스', 3005],
# 원피스 22~24
['미니 원피스', 20006],
['미디 원피스', 20007],
['맥시 원피스', 20008],
# 스커트 25~27
['미니스커트', 22001],
['미디스커트', 22002],
['롱 스커트', 22003]
]
# -
a= list(str(CategoryId_list[i][1]) for i in range(len(CategoryId_list)))
print(a)
if a[27].split('2200')[0] == '':
print('Y')
else:
print('N')
len(CategoryId_list)
import time
time.sleep(3)
# +
#import pymysql
#db = pymysql.connect(host='localhost', user='root', password='<PASSWORD>', db='musinsa')
#curs = db.cursor()
item_name_list=[]
item_price_list=[]
item_brand_list=[]
item_type_list=[]
for i in range(len(CategoryId_list)):
CategoryId = str(CategoryId_list[i][1])
for k in range(1,6):
Limit=k
if CategoryId.split('2000')[0] =='':
url= f"https://search.musinsa.com/category/"\
+'0'+ CategoryId+'?d_cat_cd='+CategoryId+f"&brand=&rate=&page_kind=search&list_kind=small&sort=pop&sub_sort=&page={Limit}"+'&display_cnt=90'
elif CategoryId.split('2200')[0]=='':
url= f"https://search.musinsa.com/category/"\
+'0'+ CategoryId+'?d_cat_cd='+CategoryId+f"&brand=&rate=&page_kind=search&list_kind=small&sort=pop&sub_sort=&page={Limit}"+'&display_cnt=90'
else:
url= f"https://search.musinsa.com/category/"\
+'00'+ CategoryId+'?d_cat_cd='+CategoryId+f"&brand=&rate=&page_kind=search&list_kind=small&sort=pop&sub_sort=&page={Limit}"+'&display_cnt=90'
#실제 제품의 항목번호를 위하여 00을 추가, 판매순으로 sorting
html= urlopen(url)
bs= BeautifulSoup(html, "html.parser")
for item_name in bs.find_all('p', {"class":"list_info"}):
item_name_list.append(no_space(item_name.get_text()).replace(" ", ""))
item_type_list.append(CategoryId_list[i][0])
# print(i, item_name_list)
# time.sleep(3)
for item_price in bs.find_all('p',{"class":"price"}):
item_price=no_space(item_price.get_text())
item_price=item_price.replace(" ", "")
item_price=item_price.replace("원", '.')
item_price=item_price.split('.')
if len(item_price)>2:
item_price_list.append(item_price[1])
else:
item_price_list.append(item_price[0])
# print(i, item_price_list)
# time.sleep(3)
for item_brand in bs.find_all('p',{"class":"item_title"}):
item_brand_list.append(no_space(item_brand.get_text()))
# print(i, item_brand_list)
# for j in range(len(item_name_list)):
# query = "INSERT INTO products(name,brand, price, type, created_date) VALUES('"\
# + item_name_list[j]+"','"+item_brand_list[j]+"',"+item_price_list[j].replace(',','')+", '"\
# + CategoryId_list[i][0]+"', curdate());"
# time.sleep(3)
# curs.execute(query)
# db.commit()
# db.close
# -
item_price_list
# +
# import pandas as pd
# columns=['item_name_list','item_price_list','item_brand_list','item_type_list']
# item_name_list_df= pd.DataFrame(item_name_list)
# item_price_list_df=pd.DataFrame(item_price_list)
# item_brand_list_df=pd.DataFrame(item_brand_list)
# item_type_list_df=pd.DataFrame(item_type_list)
# df= pd.concat(item_name_list_df, item_price_list_df, item_brand_list_df, item_type_list_df)
# print(df.head())
# -
print(len(item_name_list))
print(len(item_price_list))
print(len(item_brand_list))
print(len(item_type_list))
print(item_name_list[1],'\n',
item_price_list[1],'\n',
item_brand_list[1],'\n',
item_type_list[1])
for i in range(len(item_name_list)):
item_name_list[i]=item_name_list[i].replace(" ", "")
item_name_list
# # 올바른 가격 정보 가져오기(세일 가격이면 세일가격으로)
url="https://search.musinsa.com/category/002012"
html= urlopen(url)
bs= BeautifulSoup(html, "html.parser")
ex_price=[]
for item_price in bs.find_all('p',{"class":"price"}):
ex_price.append(no_space(item_price.get_text()))
for i in range(len(ex_price)):
ex_price[i]=ex_price[i].replace(" ", "")
ex_price
for i in ex_price:
a=i.replace(" ",'')
b=a.replace("원", '.')
c=b.split('.')
if len(c)>2:
print(c[1])
else:
print(c[0])
for i in range(len(ex_price)):
if len(ex_price[i].split('원'))> 2:
ex_price[i]=ex_price[i].split('원')[1]
# +
for i in range(len(ex_price)):
numbers = re.sub(r'[^0-9]', '', ex_price[i])
print(numbers)
# -
# # AWS RDS에 접속하여 쿼리 보내기
query=[]
for j in range(len(item_name_list)):
query.append("INSERT INTO products(name,brand, price, type) VALUES('"+ item_name_list[j]+"','"+item_brand_list[j]+"',"+item_price_list[j].replace(',','')+", '"+ item_type_list[j]+"');")
import pymysql
db = pymysql.connect(host='musinsa-database.crtzc6j6k6ch.ap-northeast-2.rds.amazonaws.com', user='admin', password='<PASSWORD>', port=3306, db='musinsa')
cursor= db.cursor()
for j in range(len(query)):
sql=query[j]
cursor.execute(sql)
result=cursor.fetchall()
print(result)
db.commit()
db.close
print(query[:10])
|
Code/musinsa_products.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import warnings
warnings.filterwarnings('ignore')
import pandas as pd
import numpy as np
abbr = 'NC'
file = 'north_carolina2018.xlsx'
# Read in federal level data
fiscal = pd.read_sas('../../data/fiscal2018', format = 'sas7bdat', encoding='iso-8859-1')
# Generate list of districts in the state in the federal data
state_fiscal = fiscal[(fiscal['STABBR'] == abbr) & (fiscal['GSHI'] == '12')]
len(state_fiscal)
# Read in state level data
state_grads = pd.read_excel('../../data/state_data_raw/' + file, sheet_name='4-year Graduation Rate')
state_grads
# Filter results.
state_grads = state_grads[(state_grads['reporting_year'] == 2018) &
(state_grads['subgroup'] == 'ALL')]
state_grads
# Select and rename columns.
state_grads = state_grads[['school_name', 'denominator', 'pct']]
state_grads.columns = ['District Name', 'Total', 'Graduation Rate']
state_grads.info()
# Convert data types.
state_grads = state_grads.replace('*', '')
state_grads['Total'] = state_grads['Total'].astype(str).str.replace('<', '')
state_grads['Graduation Rate'] = state_grads['Graduation Rate'].astype(str).str.replace('>', '')
state_grads['Graduation Rate'] = state_grads['Graduation Rate'].astype(str).str.replace('<', '')
state_grads['Total'] = pd.to_numeric(state_grads['Total'])
state_grads['Graduation Rate'] = pd.to_numeric(state_grads['Graduation Rate']) / 100
# Check for matches and non-matches in the two lists.
# +
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.upper().str.strip()
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.upper().str.strip()
# +
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(' SCHOOL DISTRICT', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace('SCH DIST', 'SCHOOL DISTRICT')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(' CS', ' CHARTER SCHOOL')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'\sHIGH\sSCHOOL$', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'\sSCHOOL$', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(' PUBLIC SCHOOLS', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(' CO ', ' COUNTY ')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(' BORO', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(' CITY', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace('TWP', 'TOWNSHIP')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'\sREG(\s|$)', ' REGIONAL ')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'\sTOWN$', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'\sTOWNSHIP$', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'\sBORO$', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'\sPUBLIC$', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace('H.S DIST.', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'^N\s', 'NORTH ')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(r'^W\s', 'WEST ')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace("’", "'")
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.replace(' REGIONAL', '')
# state_grads['District Name'] = state_grads['District Name'].astype(str).str.strip()
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(' BOARD OF EDUCATION', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(' VOCATIONAL TECHNICAL', ' VOCATIONAL')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(' CITY', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(' PUBLIC SCHOOLS', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(' PUBLIC', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(' SCHOOL DISTRICT', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(r'\sHIGH\sSCHOOL$', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(r'\sSCHOOL$', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(r'\sTOWNSHIP$', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(r'\sTOWN$', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(r'\sBOROUGH$', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(r'\sBORO$', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(r'\sDISTRICT$', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace('VOCATIONAL AND TECHNICAL', 'VOCATIONAL')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace('VOCATIONAL-TECHNICAL', 'VOCATIONAL')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace('TECHNICAL', 'VOCATIONAL')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace('INSTITUTE OF TECHNOLOGY', 'VOCATIONAL')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace('SCHOOLS OF TECHNOLOGY', 'VOCATIONAL')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(r'\sHIGH$', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.replace(' REGIONAL', '')
# state_fiscal['NAME'] = state_fiscal['NAME'].astype(str).str.strip()
# -
matches = [name for name in list(state_grads['District Name']) if name in list(state_fiscal['NAME'])]
matches.sort()
len(matches)
A = [name for name in list(state_grads['District Name']) if name not in list(state_fiscal['NAME'])]
A.sort()
A
B = [name for name in list(state_fiscal['NAME']) if name not in list(state_grads['District Name'])]
B.sort()
B
# Match any remaining samples I can find.
state_fiscal_rename = {
#'<NAME>',
'Central Wake Charter High School' : 'Central Wake Charter High',
'Charlotte Secondary School' : 'Charlotte Secondary',
'Cherokee Central Schools' : 'Cherokee County Schools',
'Commonwealth High School' : 'Commonwealth High',
#'DPS Education Services (fka Div Prisons)',
#'Deaf and Blind Schools',
'Elizabeth City-Pasquotank Public Schools' : 'Pasquotank County Schools',
'Gaston College Preparatory' : 'KIPP Gaston College Preparatory',
'Gray Stone Day School' : 'Gray Stone Day',
'Mooresville Graded School District' : 'Mooresville City Schools',
#'Mountain Island Charter',
#'NCDPS Juvenile Education Services',
'North East Carolina Preparatory School' : 'North East Carolina Prep',
'Oxford Preparatory School' : 'Oxford Preparatory',
'Piedmont Community Charter' : 'Piedmont Community Charter School',
'Stewart Creek High School' : 'Stewart Creek High',
'The College Preparatory and Leadership A' : 'College Prep and Leadership Academy',
'Thomas Jefferson Classical Academy' : 'Thomas Jefferson Class Academy',
#'Union Academy Charter School' : 'Union Academy',
'<NAME> / Forsyth County Schools' : 'Forsyth County Schools',
'Woods Charter School' : 'Woods Charter'
}
state_fiscal = state_fiscal.replace(state_fiscal_rename)
# Merge federal and state data, keeping only matches between the two.
state_grads_merged = pd.merge(state_fiscal, state_grads, how='inner', left_on='NAME', right_on='District Name')
# Save cleaned data.
state_grads_merged.to_csv('../../data/state_data_merged/' + abbr + '.csv', index=False)
|
cleaning/01_cleaning_code_by_state/NC_cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Predict the magnitude of the next eathquake
import Utils
import pydoc
from Utils import DataSet
from keras.models import Sequential
from keras.layers import Dense
from keras.layers import Dropout
from keras.layers import LSTM
from keras import optimizers
from keras.utils import to_categorical
from keras.callbacks import EarlyStopping
from keras.layers.embeddings import Embedding
from keras.preprocessing import sequence
import matplotlib.pyplot as plt
import matplotlib.patches as mpatches
from matplotlib.pyplot import cm
import pandas as pd
import numpy as np
# ## Name of current file
current_file = 'mag-disc'
setting = 5
# # Settings, formatData:
#
# | **config:** (input data) |
# |---------------------------------------------------------------------------------------------------------|
# | **0: Includes (9 features)** | **1: Includes (9 + 4 = 13 features)** | **2: Includes (9 + 4 + 9 = 22 features)** |
# | - Time | - Latitude | - Eigenvalue, 3 values |
# | - Magnitude | - Longitude | - Plunge, 3 values |
# | - Moment Tensor Exponent | - Depth | - Azimuth, 3 values |
# | - Moment Tensor, 6 values | - Scalar moment | |
#
#
# ## Variable input
call = []
call.append([])
call.append([EarlyStopping(patience=2)])
# +
act, day, epo, cal, ear = Utils.experiment_settings(setting)
activation = act
maxdays = day
epochs = epo
callbacks = call[cal]
early = ear
# -
# ## Constant in experiments
scale = 1
shape = 1
categories_in = 10
categories_out = 10
# ## Get data from database, and format predictors and target
ds = DataSet('1972/01/01', '2019/01/01', maxdays=maxdays)
# ## Predictors
n_predictors = 0
predict_txt = []
for k in range(3):
exec(f'predictors{n_predictors} = ds.formatData(config={k}, norm=scale, shape=shape)')
exec(f'predict_txt.append(\"Data config {k}: predictors{n_predictors} - input values \
between -1 and 1\")')
print(predict_txt[n_predictors])
n_predictors += 1
exec(f'predictors{n_predictors} = ds.formatDataDiscrete(categories=categories_in, \
config={k}, norm=scale, shape=shape)')
exec(f'predict_txt.append(\"Data config {k}: predictors{n_predictors} - discrete input \
values ({categories_in} categories)\")')
print(predict_txt[n_predictors])
n_predictors += 1
# +
pred_shape = []
for k in range(n_predictors):
exec(f'pred_shape.append(predictors{k}.shape[2])')
print("Feature size of predictors:", pred_shape)
# -
# ## Targets
n_targets = 1
target_txt = []
target_txt.append('targets0 - Predict the magitude of the next earthquake. Divide output into ' + \
str(categories_out) + ' categories.')
print(target_txt[0])
targets0 = ds.gt_mag_disc(categories=categories_out)
# # Split data for training and testing
splitat = int(len(predictors0) * .7)
for k in range(n_predictors):
exec(f'input_train{k}, input_test{k} = predictors{k}[ : splitat] , predictors{k}[splitat : ]')
for k in range(n_targets):
exec(f'output_train{k}, output_test{k} = targets{k}[ : splitat] , targets{k}[splitat : ]')
# # Train and test
for i in range(n_predictors):
for j in range(n_targets):
exec(f'model{i}{j} = Sequential()')
exec(f'model{i}{j}.add(LSTM((100), return_sequences=False, use_bias=True, \
input_shape=(1, pred_shape[{i}])))')
exec(f'model{i}{j}.add(Dense(categories_out, activation=activation))')
exec(f'model{i}{j}.compile(loss=\'binary_crossentropy\', optimizer=\'adam\', metrics=[\'mae\'])')
exec(f'model_training{i}{j} = model{i}{j}.fit(input_train{i}, output_train{j}, \
validation_data = (input_test{i}, output_test{j}), epochs = epochs, \
verbose = 0, callbacks = callbacks)')
# # Analyze learning
# +
p = []
for i in range(n_predictors):
patches = []
title = 'predictions' + str(i)
color = iter(cm.rainbow(np.linspace(0,1,n_targets)))
for j in range(n_targets):
c = next(color)
exec(f'plt.plot(model_training{i}{j}.history[\'val_loss\'], c=c)')
exec(f'p.append(mpatches.Patch(color=c, label=\'targets{j}\'))')
patches.append(p[j])
plt.title(title)
plt.xlabel('Epochs')
plt.ylabel('Validation score')
plt.legend(handles=patches)
plt.show()
# -
# ## Predict based on model
for i in range(n_predictors):
for j in range(n_targets):
exec(f'modpred{i}{j} = model{i}{j}.predict(input_test{i})')
# # Plot raw output
# +
plt.style.use('fivethirtyeight')
types = []
n_types = 0
for i in range(n_predictors):
for j in range(n_targets):
title = 'type ' + str(i) + ': predictions ' + str(i) + ' and target ' + str(j)
exec(f'plt.plot(modpred{i}{j}[:],"bx")')
types.append(title)
plt.title(title)
plt.show()
n_types += 1
# -
# # Measure accuracy
# ## Create set based on normal distributed average
#
# (Baseline)
n_dummy = 10
dummyset = ds.getGroundTruthNextMag()
minim = min(dummyset)
maxim = max(dummyset)
# +
dummy = []
dummy_disc = []
for i in range(n_dummy):
dummy.append(Utils.S_model_Predict(dummyset, enable_random_normal=True))
dummy[i] = dummy[i][splitat : ]
dummy_disc.append(Utils.getDiscr3t3(categories_out, dummy[i], minim, maxim))
# -
# ## Plot baseline prediction
perc_avg_list = []
# +
for i in range(n_dummy):
plt.title('Baseline ' + str(i))
perc_avg_list.append(Utils.accu_plot_disc(dummy_disc[i], output_test0))
dummy_avg = np.average(perc_avg_list)
print('-----------------------------------------')
print('Average of baseline: ' + str(dummy_avg) + '%')
print('-----------------------------------------')
# -
# ## Plot accuracy of models
# +
print('-----------------------------------------')
print('Plotting models:')
print('-----------------------------------------')
percentage = []
n_types = 0
for i in range(n_predictors):
for j in range(n_targets):
title = types[n_types]
plt.title(title)
exec(f'percent = Utils.accu_plot_disc(modpred{i}{j}, output_test{j})')
percentage.append(percent)
n_types += 1
# -
# ## Print summary
best_percentage = max(percentage)
best_per_pos = [i for i, j in enumerate(percentage) if j == best_percentage]
best_pp = ','.join(str(n) for n in best_per_pos)
diff = best_percentage - dummy_avg
print('Model: Percentage list for diffent types of input:', percentage)
print('Model: Best percentage: ' + str(best_percentage) + '%')
print('Model: Best performing network type: ' + str(best_pp))
print('-----------------------------------------')
print('Baseline: Average percentage: ' + str(int(dummy_avg)) + '%')
print('-----------------------------------------')
print('Difference, baseline vs. model: ' + str(int(diff)) + '%')
print('-----------------------------------------')
print('Types:')
print('-----------------------------------------')
for line in types:
print(line)
print('-----------------------------------------')
print('Predictors:')
print('-----------------------------------------')
for line in predict_txt:
print(line)
print('-----------------------------------------')
print('Targets:')
print('-----------------------------------------')
for line in target_txt:
print(line)
# # Backup the notebook
input("Press enter to backup the notebook (save first)")
path = './output/' + current_file + '/'
epochs = str(epochs) + early
output_file = Utils.join_list([current_file, activation, maxdays, epochs, best_percentage], '-')
# +
i = 0
type_perc = Utils.join_list(percentage, '|')
infolist = [i, best_percentage, dummy_avg, diff, best_pp, activation, maxdays, epochs, categories_out, type_perc]
out_org = Utils.org_table([infolist])
# + magic_args="-s \"$current_file\" \"$path\" \"$output_file\" \"$out_org\"" language="bash"
# jupyter nbconvert --to html $1.ipynb
# mv $1.html $2$(date +"%Y%m%d-%H%M%S")-$3.html
# jupyter nbconvert --to latex $1.ipynb
# mv $1.tex $2backup/$(date +"%Y%m%d-%H%M%S")-$3.tex
# mv $1_files $2backup/$(date +"%Y%m%d-%H%M%S")-$3
# cp $1.ipynb $2backup/notebook/$(date +"%Y%m%d-%H%M%S")-$3.ipynb
# echo $4 >> $2info.org
|
output/mag-disc/backup/notebook/20180520-162116-mag-disc-relu-180-200es-46.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="ENx3XAmY6l5l"
# ## Bi-directional LSTM on IMDb dataset with Keras
#
# Source: https://keras.io/examples/nlp/bidirectional_lstm_imdb/
# + colab={"base_uri": "https://localhost:8080/"} id="gk9dKIcIoOdk" outputId="ff334243-7b10-4725-f8bb-004c3d1a6eb0"
# Setup
import numpy as np
import warnings
# Ignore FutureWarning from NumPy
warnings.simplefilter(action = 'ignore', category = FutureWarning)
from tensorflow import keras
from tensorflow.keras import layers
from keras.datasets import imdb
from keras.preprocessing import sequence
from keras.layers import Dense, LSTM
from keras.models import Sequential
from keras.layers.embeddings import Embedding
from sklearn.metrics import f1_score, classification_report
max_features = 20000 # Only consider the top 20k words
max_len = 200 # Only consider the first 200 words of each movie review
# Build the model
# Input for variable-length sequences of integers
inputs = keras.Input(shape = (None,), dtype = "int32")
# Embed each integer in a 128-dimensional vector
x = layers.Embedding(max_features, 128)(inputs)
# Add 2 bidirectional LSTMs
x = layers.Bidirectional(layers.LSTM(64, return_sequences = True))(x)
x = layers.Bidirectional(layers.LSTM(64))(x)
# Add a classifier
outputs = layers.Dense(1, activation = "sigmoid")(x)
model = keras.Model(inputs, outputs)
model.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="gFI85VWMqQYI" outputId="2c26413d-f34f-4b26-9a14-d9f7d615aede"
# Load the IMDB movie review sentiment data
(x_train, y_train), (x_test, y_test) = keras.datasets.imdb.load_data(
num_words = max_features, seed = 123)
print(len(x_train), "Training sequences")
print(len(x_test), "Test sequences")
# Use pad_sequence to standardize sequence length:
# This will truncate sequences longer than 200 words and zero-pad sequences
# shorter than 200 words.
x_train = keras.preprocessing.sequence.pad_sequences(x_train, maxlen = max_len)
x_test = keras.preprocessing.sequence.pad_sequences(x_test, maxlen = max_len)
# + id="ey6TP-in7IJs"
# Split into validation since the load_data function won't allow us to do it manually
np.random.seed(123)
val_ind = np.random.randint(0, 24999, 5000)
x_val = x_train[val_ind]
y_val = y_train[val_ind]
# + colab={"base_uri": "https://localhost:8080/"} id="pGNTyKsnqR9a" outputId="49ed1d37-6a00-41c2-a86e-3c7082393911"
# Train and evaluate the model
model.compile(optimizer = "adam", loss = "binary_crossentropy", metrics = ["accuracy"])
history = model.fit(x_train,
y_train,
batch_size = 32,
epochs = 2,
validation_data = (x_val, y_val))
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="mDebbuPRsKRs" outputId="f47a377c-9d02-4896-fbaf-2dcf7284a45c"
history_dict = history.history
acc = history_dict['accuracy']
val_acc = history_dict['val_accuracy']
loss = history_dict['loss']
val_loss = history_dict['val_loss']
epochs = range(1, len(acc) + 1)
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams["figure.figsize"] = (10.0, 6.0)
plt.plot(epochs, acc, '-r', label = 'Training accuracy')
plt.plot(epochs, val_acc, '-b', label = 'Validation accuracy')
plt.title('Training & validation accuracy')
plt.xlabel('Epochs')
plt.ylabel('Accuracy')
plt.legend(loc = 'best')
plt.show()
# + colab={"base_uri": "https://localhost:8080/"} id="XD3Q_WOCxnti" outputId="82a94317-da6c-4e71-c9d2-b9e974a58051"
# Get validation accuracy + loss
scores = model.evaluate(x_val, y_val, batch_size = 32, verbose = 0)
print('Validation loss: %.4f' % scores[0])
print('Validation accuracy: %.4f' % scores[1])
# + colab={"base_uri": "https://localhost:8080/"} id="kZcnlHL4y04L" outputId="ae12171c-5d06-4e39-cc43-c29995faad46"
# Predict classes for test set
yhat_classes = model.predict(x_test, verbose = 0)
yhat_classes = np.concatenate([1 - yhat_classes, yhat_classes], axis = 1)
yhat_classes = np.argmax(yhat_classes, axis = 1)
# Get classification report
print("Classification Report")
print(classification_report(y_test, yhat_classes))
|
source/BiLSTM_with_Keras.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Problem 1 (Morbidelli 8.1)
#
# Prove the following results:
#
# 1. $\mathscr{L}\{cos{(at)}\} = \frac{s}{s^2 + a^2}$
# 2. $\mathscr{L}\{sinh{(at)}\} = \frac{a}{s^2 - a^2}$
# 3. $\mathscr{L}\{cosh{(at)}\} = \frac{s}{s^2 - a^2}$
# 4. $\mathscr{L}\{t^n\} = \frac{n!}{s^{(n+1)}}$
# ### 1.1
# $$\mathscr{L}\{cos(at)\}=\int_0^\infty e^{-st}cos{(at)}dt\\
# =\frac{1}{a}sin{(at)}e^{-st}\bigg|_0^\infty-\int_0^\infty[\frac{1}{a}sin(at)*(-s)e^{-st}]dt
# =0+\frac{a}{s}\int_0^\infty sin(at)=
# \frac{s}{a}[\frac{1}{a}cos(at)e^{-st}\bigg|_0^\infty-\int_0^\infty\frac{-1}{a}cos(at)(-s)e^{-st}dt]\\
# =\frac{s}{a}[\frac{1}{a}-\frac{s}{a}\int_0^\infty cos(at)e^{-st}dt]
# $$
# set $\mathscr{L}\{cos(at)\}=\int_0^\infty e^{-st}cos{(at)}dt=A$ $$\\$$
# We have $$A=\frac{s}{a}[\frac{1}{a}-\frac{s}{a}A]$$
# It follows that $$A=\frac{s}{s^2+a^2}$$
# $$\mathscr{L}\{cos(at)\}=\frac{s}{s^2+a^2}$$
# ### 1.2
# $$
# \mathscr{L}\{sinh(at)\}=\int_0^\infty e^{-st}\frac{(e^{at}-e^{-at})}{2}dt
# =\frac{1}{2}[\int_0^\infty e^{(a-s)t}dt-\int_0^\infty e^{-(a+s)t}dt]
# =\frac{1}{2}[\frac{1}{a-s} e^{(a-s)t} \bigg|_0^\infty+\frac{1}{a+s}e^{-(a+s)t} \bigg|_0^\infty]
# =\frac{1}{2}[\frac{1}{s-a}-\frac{1}{s+a}]
# =\frac{a}{s^2-a^2}
# $$
# ### 1.3
# $$
# \mathscr{L}\{cosh(at)\}=\int_0^\infty e^{-st}\frac{(e^{at}+e^{-at})}{2}dt
# =\frac{1}{2}[\int_0^\infty e^{(a-s)t}dt+\int_0^\infty e^{-(a+s)t}dt]
# =\frac{1}{2}[\frac{1}{a-s} e^{(a-s)t} \bigg|_0^\infty-\frac{1}{a+s}e^{-(a+s)t} \bigg|_0^\infty]
# =\frac{1}{2}[\frac{1}{s-a}+\frac{1}{s+a}]\\
# =\frac{s}{s^2-a^2}
# $$
# ### 1.4
# $$
# \mathscr{L}\{t^n\}=\int_0^\infty e^{-st}t^ndt
# =\frac{-1}{s}e^{-st}t^n \bigg|_0^\infty -\int_0^\infty\frac{-1}{s}e^{-st}nt^{n-1}dt
# =0+\frac{n}{s}\int_0^\infty e^{-st}t^{n-1}
# =\frac{n}{s}\mathscr{L}\{t^{n-1}\}$$
# Iterate for N times till 1, we have:
# $$
# \mathscr{L}\{t^n\}=\frac{n(n-1)(n-2)\cdots(n-i)\cdots1}{s^n}\mathscr{L}\{1\}=\frac{n!}{s^n}\mathscr{L}\{1\}\\
# It gives out:
# \mathscr{L}\{1\}=\int_0^\infty e^{-st}dt=\frac{1}{s}
# $$
# Substitute:
# $$
# \mathscr{L}\{t^n\}=\frac{n!}{s^(n+1)}
# $$
# #### Problem 2 (Morbidelli 8.5)
#
# Solve by LaPlace transform:
#
# $$y''+ y' -2y = 3\cos{3t} - 11\sin{3t}, y(0) = 0, y'(0) = 6 $$
#
#
# ### 2
# Begin with the equation:
# $$\mathscr{L}\{f'(x)\}=s\mathscr{L}\{f(x)\}-f(0)$$
# apply laplace transform to each term together with the initial conditions:
# $$
# \mathscr{L}\{y''\}=s\mathscr{L}\{y'\}-y'(0)=s[s\mathscr{L}\{y\}-y(0)]-y'(0)=s^2\mathscr{L}\{y\}-6\\
# \mathscr{L}\{y'\}=s\mathscr{L}\{y\}-y(0)=s\mathscr{L}\{y\}\\
# \mathscr{L}\{2y\}=2\mathscr{L}\{y\}\\
# \mathscr{L}\{3cos3t\}=\frac{3s}{s^2+9}\\
# \mathscr{L}\{11sin3t\}=\frac{33}{s^2+9}\\
# $$
# We have the Laplace form of the ODE:
# $$
# s^2\mathscr{L}\{y\}-6+s\mathscr{L}\{y\}-2\mathscr{L}\{y\}=\frac{3s}{s^2+9}-\frac{33}{s^2+9}
# $$
# therefore
# $$
# \mathscr{L}\{y\}=\frac{6s^2+3s+21}{(s^2+9)(s^2+s-2)}\\
# =\frac{3}{s^2+9}+\frac{3}{s^2+s-2}\\
# =\frac{3}{s^2+9}+\frac{1}{s-1}-\frac{1}{s+2}
# $$
# Then apply inverse Laplace transform to each term:
# $$
# \mathscr{L^{-1}}\{\frac{3}{s^2+9}\}=sin3t\\
# \mathscr{L^{-1}}\{\frac{1}{s-1}\}=e^t\\
# \mathscr{L^{-1}}\{\frac{1}{s+2}\}=e^{-2t}
# $$
# therefore, we get the solution for y:
# $$
# y=sin3t+e^t-e^{-2t}
# $$
# #### Problem 3 (Morbidelli 8.9 modified)
#
# Solve by LaPlace transform the system of ODEs:
#
# $$y_1'' + y_1' + 2y_2' - y_1 = 1 $$
# $$y_2''' + y_1'' - 2y_1 + y_2 = e^{-t} $$
# $$y_1(0) = 2, y_1'(0) = 0, y_2(0) = 2, y_2'(0) = 0, y_2''(0) = 1 $$
#
# Rather than solving completely just find the matrix representation in the LaPlace domain.
# ### 3
# $$
# \mathscr{L}\{y'''\}=s\mathscr{L}\{y''\}-y''(0)=s[s\mathscr{L}\{y'\}-y'(0)]-y''(0)=s[s[s\mathscr{L}\{y\}-y(0)]-y'(0)]-y''(0)\\
# \mathscr{L}\{y''\}=s\mathscr{L}\{y'\}-y'(0)=s[s\mathscr{L}\{y\}-y(0)]-y'(0)\\
# \mathscr{L}\{y'\}=s\mathscr{L}\{y\}-y(0)\\
# $$
#
# in this case:
# $$
# y_1''=s[s\mathscr{L}\{y_1\}-2]-0=s^2\mathscr{L}\{y_1\}-2s\\
# y_1'=s\mathscr{L}\{y_1\}-2=s\mathscr{L}\{y_1\}-2\\
# y_2'''=s[s[s\mathscr{L}\{y_2\}-2]-0]-1=s^3\mathscr{L}\{y_2\}-2s^2-1\\
# y_2'=s\mathscr{L}\{y_2\}-2=s\mathscr{L}\{y_2\}-2\\
# \mathscr{L}\{1\}=\frac{1}{s}\\
# \mathscr{L}\{e^{-t}\}=\frac{1}{s+1}\\
# $$
# substitute them into original ODEs to get the first order form:
# $$
# s^2\mathscr{L}\{y_1\}-2s+s\mathscr{L}\{y_1\}-2+2s\mathscr{L}\{y_2\}-4-\mathscr{L}\{y_1\}=\frac{1}{s}\\
# s^3\mathscr{L}\{y_2\}-2s^2-1+s^2\mathscr{L}\{y_1\}-2s-2\mathscr{L}\{y_1\}+\mathscr{L}\{y_2\}=\frac{1}{s+1}
# $$
# It follows that:
# $$
# \mathscr{L}\{y_1\}[s^2+s-1]+\mathscr{L}\{y_2\}2s=\frac{1}{s}+2s+6\\
# \mathscr{L}\{y_1\}[s^2-2]+\mathscr{L}\{y_2\}[s^3+1]=\frac{1}{s+1}+2s^2+2s+1
# $$
# The solution can be expressed by the following matrix form:
# $$
# \left[ {\begin{array}{cc}
# {s^2+s-1} & {2s}\\
# {s^2-2} & {s^3+1}\\
# \end{array} } \right]
# \left[ {\begin{array}{cc}
# \mathscr{L}\{y_1\}\\
# \mathscr{L}\{y_2\}\\
# \end{array} } \right]
# =
# \left[ {\begin{array}{cc}
# {\frac{1}{s}+2s+6} \\
# \frac{1}{s+1}+2s^2+2s+1\\
# \end{array} } \right]
# $$
# **Problem 4 (Morbidelli 8.10 modified)**
#
# Solve the diffusion-reaction problem given by (8.6.1) and (8.6.2) in the case where the non-homogeneous term is given by $f(t) = e^{-t}$. Also identify the impulse response.
#
# $$D \frac{\partial^2 c}{\partial x^2} + f(t) = \frac{\partial c}{\partial t}, x \in (0, L), t>0, c(x, 0)=0, c(0,t)=0, c(L,t)=0 $$
#
# **Plot the final solution**.
# ##### 4
# Apply Laplace transform to each term:
# $$
# \mathscr{L}\{D\frac{\partial^2c}{\partial x^2}\}=D\frac{\partial^2}{\partial x^2}\int_0^\infty e^{-st}c(x,t)dt
# =D\frac{d^2C(x,s)}{dx^2}\\
# \mathscr{L}\{f(t)\}=\int_0^\infty e^{-st}e^{-t}dt=\frac{1}{s+1}\\
# \mathscr{L}\{\frac{\partial c(x,t)}{\partial x}\}=\int_0^\infty e^{-st}\frac{\partial c(x,t)}{\partial t}dt
# =s\mathscr{L}\{c(x,t)\}-c(x,0)=sC(x,s)
# $$
# and to the BCs and IC:
# $$
# \mathscr{L}\{c(0,t)\}=0\\
# \mathscr{L}\{c(L,t)\}=0\\
# \mathscr{L}\{c(x,0)\}=0\\
# $$
# now we apply the Laplace transform to the PDE and rearrange:
# $$
# D\frac{d^2C(x,s)}{dx^2}-sC(x,s)=-\frac{1}{s+1}
# $$
# with BCs and IC:
# $$
# \mathscr{L}\{c(0,t)\}=0\\
# \mathscr{L}\{c(L,t)\}=0\\
# \mathscr{L}\{c(x,0)\}=0\\
# $$
# For this non-homogeneous ODE, the solution can be expressed as $C=C_c+C_p$ \\
# First, we solve for $C_c$:
# $$
# \frac{d^2C(x,s)}{dx^2}-\frac{s}{D}C(x,s)=0\\
# $$
# using characteristic functions:
# $$
# \lambda^2-\frac{s}{D}=0
# $$
# we can obtain:
# $$
# \lambda=\pm \sqrt{\frac{s}{D}}
# $$
# since there are 2 distinct real roots:
# $$
# C_c(x,s)=d_1sinh{(\sqrt{\frac{s}{D}}x)}+d_2cosh{(\sqrt{\frac{s}{D}}x)}
# $$
# Then we focus our attention on$c_p$, because $\frac{1}{s+1}$is independent on $t$, we can treat it as a constant,so we guess $C_p$=A and plug it into the ODE:
# $$
# sA=\frac{1}{s+1}
# $$
# so
# $$
# C_p=A=\frac{1}{s(s+1)}
# $$
# Next,
# $$
# C(x,s)=C_c+C_p=d_1sinh{(\sqrt{\frac{s}{D}}x)}+d_2cosh{(\sqrt{\frac{s}{D}}x)}+\frac{1}{s(s+1)}
# $$
# and we plug into BCs, we can obtain:
# $$
# d_1=\frac{1}{s(s+1)}
# \frac{cosh(\sqrt{\frac{s}{D}}L)-1}{sinh(\sqrt{\frac{s}{D}}L)}
# $$
# $$
# d_2=-\frac{1}{s(s+1)}
# $$
# Then we substitute $d_1,d_2$ into $C(x,s):$
# $$
# C(x,s)=\frac{1}{s+1}\frac{sinh(\sqrt{\frac{s}{D}}L)-sinh(\sqrt{\frac{s}{D}}x)-sinh(\sqrt{\frac{s}{D}}L-x)}{sinh(\sqrt{\frac{s}{D}}L)*s}
# $$
# we express $C(X,s)=F(s)G(s), G(s)=\frac{p(s)}{q(s)}$,in this case,
# $$
# F(s)=\frac{1}{1+s}\\
# G(s)=\frac{sinh(\sqrt{\frac{s}{D}}L)-sinh(\sqrt{\frac{s}{D}}x)-sinh(\sqrt{\frac{s}{D}}L-x)}{sinh(\sqrt{\frac{s}{D}}L)*s}\\
# p(s)=sinh(\sqrt{\frac{s}{D}}L)-sinh(\sqrt{\frac{s}{D}}x)-sinh(\sqrt{\frac{s}{D}}L-x)\\
# q(s)=sinh(\sqrt{\frac{s}{D}}L)*s\\
# q'(s)=sinh(\sqrt{\frac{s}{D}}L)+\frac{sL}{2\sqrt{sD}}cosh(\sqrt{\frac{s}{D}}L)
# $$
# now we try to apply the partial fraction decompostion to $q(s)$, when $q(s)=0$
# $$
# s=0 \ or \ sinh(\sqrt{\frac{s}{D}}L)=0
# $$
# and $s=0$ is also a root for $p(s)$, so 0 is a common root, thus the root for $q(s)$ is:
# $$
# \sqrt{\frac{s_n}{D}}L=z=a+ib
# sinh(a+ib)=sinh(a)cosh(ib)+sinh(ib)cosh(a)\\
# =sinh(a)cos(b)+isin(b)cosh(a)\\
# sinh(a+ib)=0
# $$
# only when the following 2 terms satisfy simultaneously:
# $$
# sinh(a)cos(b)=0\\
# isin(b)cosh(a)=0
# $$
# therefore
# $$
# a=0\\
# b=\pm n\pi, n=1,2,3\ldots
# $$
# then
# $$
# \sqrt{\frac{s_n}{D}}L=\pm in\pi \\
# s_n=-\frac{n^2\pi^2D}{L^2}, \ n=1,2,3\ldots
# $$
# substitute $s_n$ into $p(s)$ and $q'(s)$:
# $$
# p(s)=sinh(in\pi)-sinh(\frac{in\pi x}{L})-sinh[in\pi \frac{(L-x)}{L}]\\
# =isin(n\pi)-isin(\frac{n\pi x}{L})-isin[n\pi \frac{(L-x)}{L}]\\
# =-isin(\frac{n\pi x}{L})-i[sin(n\pi)cos(\frac{n\pi x}{L})-sin(\frac{n\pi x}{L})cos(n\pi)]\\
# =-isin(\frac{n\pi x}{L})[1-(-1)^n], \ n=1,2,3\ldots
# $$
# we can simply the expression for $p(s)$:
# $$
# p(s)=0,\ when\ n=2,4,6\ldots\\
# p(s)=-2isin(\frac{n\pi x}{L}),\ when\ n=1,3,5\ldots\\
# $$
# and
# $$
# q'(s)=sinh(in\pi)+in\pi cos(n\pi)\\
# =\frac{in\pi}{2}(-1)^n,\ n=1,2,3\ldots
# $$
# so when $n=1,3,5\ldots$,
# $$
# q'(s)=-\frac{in\pi}{2}
# $$
# Finally, we get the solution for $g(x,t)$:
# $$
# g(x,t)=\frac{4}{\pi}\sum_{n=1,3,5\ldots}^\infty \frac{1}{n} sin(\frac{n\pi x}{L})e^{-\frac{n^2\pi^2D}{L^2}t}
# $$
# in the end,
# $$
# c(x,t)=\int_0^\infty f(t-z)g(z)dz\\
# =\int_0^t e^{-(t-z)}\frac{4}{\pi}\sum_{n=1,3,5\ldots}^\infty \frac{1}{n} sin(\frac{n\pi x}{L})e^{-\frac{n^2\pi^2D}{L^2}z}dz\\
# =\sum_{n=1,3,5\ldots}^\infty \frac{4}{(-\frac{n^2\pi^2D}{L^2}+1)n\pi}sin(\frac{n\pi x}{L})(e^{-\frac{n^2\pi^2D}{L^2}t}-e^{-t})
# $$
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import axes3d
from matplotlib import cm
# +
L = 10
T=100
D = 0.1
d=100
result = 0
x = np.linspace(0, L, d)
t = np.linspace(0, T, d)
u = np.zeros((x.shape[0], t.shape[0]))
xv, tv = np.meshgrid(x, t)
for i in range(1, 100):
n = 2*i - 1
s = ((-n**2*np.pi**2*D)/(L**2))
A = 4/((s+1)*n*np.pi)
B = np.sin(n*np.pi*xv/L)
C = np.exp(s*tv)-np.exp(-tv)
result += A*B*C
result
# +
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111, projection='3d')
ax.plot_surface(xv, tv, result,cmap=cm.coolwarm)
ax.view_init(azim=50,elev=20)
ax.set_xlabel('x')
ax.set_ylabel('t')
ax.set_zlabel('c')
#ax.set_zlim(20, 210)
# -
# #### Problem 5 (Morbidelli 8.12 modified)
#
# Consider the transient heat conduction problem in a one-dimensional rod with no internal heat generation:
#
# $$\frac{a}{\kappa} \frac{\partial T}{\partial t} = \frac{\partial^2 T}{\partial x^2} - u^2 T $$
#
# with the boundary/initial conditions:
#
# $$T(0, t) = T_0 $$
# $$\frac{\partial T}{\partial x}(L, t) = 0 $$
# $$T(x, 0) = 0 $$
#
# Instead of completely solving the problem using LaPlace transforms (which is actually quite a difficult task), just find the solution in the LaPlace domain without performing the reverse transform. Remember to solve for the coefficients using the BCs and ICs too. This is to demonstrate how LaPlace transforms theoretically can simplify a PDE to an ODE, but with the caveat that you may be stuck with a **really** ugly time transforming the solution back to the time domain.
# apply Laplace transform to the PDE:
# $$
# \mathscr{L}\{\frac{a}{\kappa} \frac{\partial T}{\partial t}\}=\frac{a}{\kappa}\mathscr{L}\{\frac{\partial T}{\partial t}\} =\frac{as}{\kappa}\mathscr{L}\{T\}\\
# \mathscr{L}\{ \frac{\partial^2 T}{\partial x^2}\}=\frac{d^2}{dx^2}\mathscr{L}\{T\}\\
# \mathscr{L}\{u^2T\}=u^2\mathscr{L}\{T\}
# $$
# and to the BCs and IC:
# $$
# \mathscr{L}\{T(0,t)\}=\frac{T_0}{s}\\
# \mathscr{L}\{\frac{\partial T}{\partial x}(L,t)\}=s\mathscr{L}\{T(L,t)\}-T(L,0)=s\mathscr{L}\{T(L,t)\}=0 \rightarrow \mathscr{L}\{T(L,t)\}=0\\
# \mathscr{L}\{T(x,0)\}=0\\
# $$
# now, using Laplace transform, we successfully simply the PDE to an ODE:
# $$
# \frac{as}{\kappa}\mathscr{L}\{T\}=\frac{d^2}{dx^2}\mathscr{L}\{T\}-u^2\mathscr{L}\{T\}\\
# $$
# viz
# $$
# \frac{d^2}{dx^2}\mathscr{L}\{T\}-(u^2+\frac{as}{\kappa})\mathscr{L}\{T\}=0
# $$
# with BCs:
# $$
# \mathscr{L}\{T(0,t)\}=\frac{T_0}{s}\\
# \mathscr{L}\{T(L,t)\}=0\\
# $$
# characteristic functions:
# $$
# r^2-(u^2+\frac{as}{\kappa})=0\\
# $$
# the root for r are:
# $$
# r_1= \sqrt{u^2+\frac{as}{\kappa}}\\
# r_2=-\sqrt{u^2+\frac{as}{\kappa}}
# $$
# as $r_1, \ r_2$ are distinct real roots:
# $$
# \mathscr{L}\{T\}=d_1sinh(\sqrt{u^2+\frac{as}{\kappa}}x)+d_2cosh(\sqrt{u^2+\frac{as}{\kappa}}x)\\
# $$
# plug into the BCs:
# $$
# d_1=-\frac{T_0}{s}\frac{cosh(\sqrt{u^2+\frac{as}{\kappa}}L)}{sinh(\sqrt{u^2+\frac{as}{\kappa}}L)}\\
# d_2=\frac{T_0}{s}\\
# $$
# Therefore, in the Laplace domain, the solution for $\mathscr{L}\{T\}$ is:
# $$
# \mathscr{L}\{T\}=-\frac{T_0}{s}\bigg[\frac{cosh(\sqrt{u^2+\frac{as}{\kappa}}L)}{sinh(\sqrt{u^2+\frac{as}{\kappa}}L)}sinh(\sqrt{u^2+\frac{as}{\kappa}}x)-cosh(\sqrt{u^2+\frac{as}{\kappa}}x)\bigg]
# $$
# #### Problem 6 (Morbidelli 8.15)
#
# Consider the axial dispersion of a tracer in an empty tube:
#
# $$-\frac{1}{Pe} \frac{\partial ^2 u}{\partial x^2} + \frac{\partial u}{\partial x} + \frac{\partial u}{\partial t} = 0, 0 < x< 1, t>0, u(x,0)=0, u(0,t)=\delta (t), u(\infty,t)=finite$$
#
# Determine the average residence time and variance of the tracer concentration distribution at the tube outlet.
# Apply the Laplace transform to the PDE:
# $$
# \mathscr{L}\{-\frac{1}{Pe}\frac{\partial^2 u}{\partial x^2}\}=-\frac{1}{Pe}\frac{d^2\mathscr{L}\{u\}}{dx^2}=-\frac{1}{Pe}\frac{d^2U(x,s)}{dx^2}\\
# \mathscr{L}\{\frac{\partial u}{\partial x}\}=\frac{d\mathscr{L}\{u\}}{dx}=\frac{dU(x,s)}{dx}\\
# \mathscr{L}\{\frac{\partial u}{\partial t}\}=s\mathscr{L}\{u\}-u(x,0)=s\mathscr{L}\{u\}=sU(x,s)\\
# \mathscr{L}\{u(0,t)\}=\mathscr{L}\{\delta(t)\}=1\\
# \mathscr{L}\{u(\infty,t)\}=\mathscr{L}\{finite\}=\frac{c}{s}\\
# $$
# thus, we simply the PDE to an ODE:
# $$
# \frac{d^2U}{dx^2}-Pe\frac{dU}{dx}-sPeU=0
# $$
# with BCs:
# $$
# U(0,t)=1\\
# U(\infty,t)=finite\\
# $$
# characteristic functions:
# $$
# r^2-rPe-sPe=0\\
# r=\frac{Pe}{2}(1\pm \sqrt{1+\frac{4s}{Pe}})
# $$
# since there are two distinct real roots:
# $$
# U(x,s)=d_1e^{[\frac{Pe}{2}(1+ \sqrt{1+\frac{4s}{Pe}})x]}+d_2e^{[\frac{Pe}{2}(1-\sqrt{1+\frac{4s}{Pe}})x]}
# $$
# plug into BCs:
# $$
# d_1=0\\
# d_2=1\\
# $$
# thus, the solution for $U(x,s)$ is:
# $$
# U(x,s)=e^{[\frac{Pe}{2}(1- \sqrt{1+\frac{4s}{Pe}})x]}\\
# \frac{\partial U}{\partial s}=-x{(1+\frac{4s}{Pe})}^{-\frac{1}{2}}U\\
# \frac{\partial^2 U}{\partial s^2}=x(1+\frac{4s}{Pe})^{-1}U[\frac{2}{Pe}(1+\frac{4s}{Pe})^{-\frac{1}{2}}+x]\\
# $$
# therefore,
# $$
# \mu_0(x)=1\\
# \mu_1(x)=-\frac{\partial U}{\partial s}(s=0)=x\\
# \mu_2(x)=\frac{2}{Pe}x+x^2\\
# $$
# then,
# $$
# t_{av}=\frac{\mu_1}{\mu_0}=x\\
# \sigma^2=\frac{\mu_2}{\mu_0}-(\frac{\mu_1}{\mu_0})^2=\frac{2}{Pe}x
# $$
# at the oultet of tube,$x=1$, so $$ \\ $$
# the average ersidence time: $$t_{av}=1$$
#
# the variance of the tracer concentration distribution :$$\sigma^2=\frac{2}{Pe}$$
#
|
hw9-Renyu Zheng.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Regression Plots
# Seaborn hat viele eingebaute Fähigkeiten um Diagramme für Regressionen zu erstellen. Jedoch werden wir Regressionen erst richtig im Machine Learning Teil des Kurses besprechen. Deshalb werden wir an dieser Stelle nur die `lmplot()` Funktion kennenlernen.
#
# *lmplot* ermöglicht es uns lineare Modelle zu visualisieren. Zusätzlich werden aber auch Funktionen zum Aufteilen und Einfärben von Daten anhand einzelner Variablen gegeben. Schauen wir uns das im Detail an!
import seaborn as sns
# %matplotlib inline
tips = sns.load_dataset("tips")
tips.head()
# ## lmplot()
sns.lmplot(x="total_bill",y="tip", data=tips)
sns.lmplot(x='total_bill',y='tip',data=tips,hue='sex')
sns.lmplot(x='total_bill',y='tip',data=tips,hue='sex',palette='coolwarm')
# ### Mit Markierungen arbeiten
# lmplot wird durch *regplot* verarbeitet. Das ist einfach eine etwas allgemeinere Form des *lmplot*. *regplot* wiederum hat einen `scatter_kws` Parameter. Innerhalb dessen definieren wir *s* in einem Dictionary, um die Markierungsgröße festzulegen (etwas verwirrend).
# http://matplotlib.org/api/markers_api.html
sns.lmplot(x='total_bill',y='tip',data=tips,hue='sex',palette='coolwarm',
markers=['o','v'],scatter_kws={'s':100})
# ## Ein Grid nutzen
#
# Wir können weitere Differenzierung durch Spalten und Zeilen hinzufügen, indem wir ein Grid nutzen. Dazu einfach die `col` und `row` Parameter spezifizieren:
sns.lmplot(x='total_bill',y='tip',data=tips,col='sex')
sns.lmplot(x="total_bill", y="tip", row="sex", col="time",data=tips)
sns.lmplot(x='total_bill',y='tip',data=tips,col='day',hue='sex',palette='coolwarm')
# ## Perspektive und Größe
# Seaborn Diagramme können in ihrer Größe und Perspektive angepasst werden:
sns.lmplot(x='total_bill',y='tip',data=tips,col='day',hue='sex',palette='coolwarm',
aspect=0.6,size=8)
# Ihr wundert euch evtl. wie sich Schriftgröße und andere optische Aspekte anpassen lassen. Dazu gibt es die Lektion "Style und Farbe"!
# # Gut gemacht!
|
3-Visualization/2-Seaborn/4-Regressionsdiagramme.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#default_exp flask_hello_world
# -
# # Hello World flask app
#
# > Getting started with Flask in nbdev.
# # Getting started
#
# The goal of this notebook is to show how you can create and run `smart_on_fhir_client_py_demo/flask_hello_world.py` on your own machine.
#
# Please see index.ipynb and set things up to run on your own machine.
# ## Code and test a simple flask app
#export
from flask import Flask
#export
def create_app(test_config=None):
"Create and configure an instance of the Flask application."
app = Flask(__name__, instance_relative_config=True)
app.config.from_mapping(
# a default secret that should be overridden by instance config
SECRET_KEY="dev"
)
if test_config is None:
# load the instance config, if it exists, when not testing
app.config.from_pyfile("config.py", silent=True)
else:
# load the test config if passed in
app.config.update(test_config)
@app.route("/hello")
def hello_world():
return f"Hello, World!"
@app.route("/hello/<human_name>")
def hello(human_name):
return f"Hello, {human_name}!"
app.add_url_rule("/", endpoint="hello_world")
return app
# ↓ make sure we can create the app with/without specifying config
assert not create_app().testing
assert create_app({"TESTING": True}).testing
# ↓ make requests against a test client to show that we can `GET` the `/hello` route with/without specifying a name
app = create_app({"TESTING": True})
client = app.test_client()
response = client.get("/hello/<NAME>, MSc")
assert response.data == b"Hello, <NAME>, MSc!"
response = client.get("/")
assert response.data == b"Hello, World!"
# The next cell just says "run the app if we're running from the command line". We use `IN_NOTEBOOK` to avoid running the app when we're in the notebook.
#export
try: from nbdev.imports import IN_NOTEBOOK
except: IN_NOTEBOOK = False
if __name__ == "__main__" and not IN_NOTEBOOK:
create_app().run(debug=True, port=8000)
# ## Convert this notebook to a python module
#
# After making changes to this notebook, we need to get nbdev to re-create the `flask_hello_world.py` file.
#
# We can do this in code ↓ or from the command line with `nbdev_build_lib`
from nbdev.export import notebook2script
notebook2script('50_flask_app.ipynb')
# ## Run the flask app
#
# Now we can run `python smart_on_fhir_client_py_demo/flask_hello_world.py` from the command line and hit URLs like
# - http://localhost:8000/ and
# - http://localhost:8000/hello/Pieter%20van%20de%20Heuvel,%20MSc
|
50_flask_hello_world.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BenNicart/CPEN-21A-CPE-1-1/blob/main/Control_Structure.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="DpAnXlkMm-9p"
# ##If statement
# + colab={"base_uri": "https://localhost:8080/"} id="TX4jm5TLnLGu" outputId="909a003e-9395-48b9-f1a7-18dec6f9cbf4"
a = 12
b = 100
if b>a:
print("b is greater than a")
# + [markdown] id="GnCk75nto0NU"
# ##Elif condition
# + colab={"base_uri": "https://localhost:8080/"} id="oGWJ2Q2Io3jw" outputId="bd411636-18f6-45be-893a-9ffe1130903d"
a = 12
b = 12
if b>a:
print("b is greater than a")
elif a==b:
print("a is equal to b")
# + [markdown] id="5eo4_mDPpjJF"
# ##Else condition
# + colab={"base_uri": "https://localhost:8080/"} id="fVo6Qy_YplNv" outputId="76ff4dde-6b66-44c4-fc03-caf1a9574ff3"
a = 12
b = 120
if b>a:
print("b is greater than a")
elif a==b:
print("a is equal to b")
else:
print("b is less than a")
# + [markdown] id="mLvyDNWpqd_n"
# ##Short Hand if Statement
# + id="gLCz7hFFqkGA"
if a>b: print("a is greater than b")
# + [markdown] id="25F2Zu-frTdO"
# ##Short Hand if else Statement
# + colab={"base_uri": "https://localhost:8080/"} id="04Y1g3QgrWTq" outputId="08490c31-273b-4587-f4ec-d9205226eff6"
print("a is greater than b") if a>b else print("b is greater than a")
# + [markdown] id="dlIQRHjIs4Rv"
# And Conditions
# + colab={"base_uri": "https://localhost:8080/"} id="7ECQ-KYKuTf7" outputId="e74fa8d3-4eec-472a-807b-e4f8baec09c0"
a = 15
b = 35
if b>a and a<b:
print("Both conditions are true")
elif b<a and a>b:
print("a is larger than b")
else:
print("none of the above")
# + [markdown] id="vtiuXYT5vP81"
# Or Conditions
# + colab={"base_uri": "https://localhost:8080/"} id="6Aai8_levSdH" outputId="7c25e7c5-1e21-46e6-acab-03c43627a8be"
a = 35
b = 20
if b>a or a==b:
print("True")
else:
print("False")
# + [markdown] id="CMeSMs7Bw59U"
# ##Nested if
# + colab={"base_uri": "https://localhost:8080/"} id="FYnJHBTUw8Hp" outputId="3069bdf1-2d1c-4f1b-dcd3-3f269529a524"
x = 9
if x>10:
print("Above 10")
if x>20:
print("Above 20")
if x>30:
print("Above 30")
else:
("Not above 30")
else:
print("not above 20")
else:
print("not above 10")
# + [markdown] id="5yQ08svhyKdV"
# Application of if... Else Statement
# + colab={"base_uri": "https://localhost:8080/"} id="gHGo_bRryPOA" outputId="4317f488-017d-4564-b082-5a4f0f254b0b"
#Exanple 1:
age = int(input("Enter your age"))
if age>=18:
print("You are qualified to vote!")
else:
print("You are not qualified to vote!")
# + colab={"base_uri": "https://localhost:8080/"} id="gcWSELwzzfKj" outputId="1848006f-5361-4390-f7da-09f2324aa021"
#Example 2:
number = int(input("Enter your number"))
if number==0:
print("Zero")
elif number>0:
print("Positive")
else:
print("Negative")
# + colab={"base_uri": "https://localhost:8080/"} id="kuK0M2GJ0CIm" outputId="515391e2-223d-4635-fbfe-33364a7fb146"
#Example 3
grade = int(input("Enter your grade"))
if grade>=75:
print("Passed")
elif grade==74:
print("Remedial")
else:
print("Failed")
|
Control_Structure.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <small> <i> This notebook was prepared by <NAME>. For license visit [github](https://github.com/donnemartin/interactive-coding-challenges) </i> </small>
# .
# # Challenge Notebook
#
# ## Problem: Given a string of words, return a string with the words in reverse
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# ## Constraints
# * Can we assume the string is ASCII?
# * Yes
# * Is whitespace important?
# * no the whitespace does not change
# * Is this case sensitive?
# * yes
# * What if the string is empty?
# * return None
# * Is the order of words important?
# * yes
#
# ## Test Cases
# * Empty string -> None
# * "the sun is very hot" -> "eht nus si yrev toh"
#
# ## Algorithm
# * Refer to the [Solution](https://github.com/donnemartin/interactive-coding-challenges/blob/master/arrays_strings/reverse_words/reverse_words_solution.ipynb) if you are stuck and need a hint, the solution notebook's algorithm discussion might be a good place to start.
# ## Code
def reverse_words (S):
#TODO: implement me
pass
# ## Unit Test
# <b> The following unit test is expected to fail until you solve challenge </b>
# +
from nose.tools import assert_equal
class UnitTest (object):
def testReverseWords(self, func):
assert_equal(func('the sun is hot'), 'eht nus si toh')
assert_equal(func(''), None)
assert_equal(func('123 456 789'), '321 654 987')
assert_equal(func('magic'), 'cigam')
print('Success: reverse_words')
def main():
test = UnitTest()
test.testReverseWords(reverse_words)
if __name__=="__main__":
main()
# -
# ## Solution Notebook
# * Review the [Solution Notebook](https://github.com/donnemartin/interactive-coding-challenges/blob/master/arrays_strings/reverse_words/reverse_words_solution.ipynb) for discussion on algorithms and code solutions.
|
staging/arrays_strings/reverse_words/reverse_words_challenge.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
matrix = [[1, 2, 3,], [4, 5, 6], [7, 8, 9]]
print(matrix)
matrix1 = []
for i in range(3):
matrix1.append([i] * 4)
print(matrix1)
matrix2 = [[0] * 3 for i in range(3)]
print(matrix2)
matrix3 = [[1 for col in range(3)] for row in range(3)]
print(matrix3)
# +
array = [[0] * 3 for i in range(3)]
for i in range(3):
for j in range(3):
array[i][j] = i + j
# -
array
array = [0] * 10
array
array[1]
# +
for i in range(10):
array[i] = 2+ 2*i
array
# -
array
[2 * i for i in range(1,11)]
array = [[0] * 4 for i in range(4)]
array
for i in range(4):
for j in range(4):
if i == 0:
array[i][j] = i + j
elif i == 1:
array[i][j] = 3 + i + j
elif i == 2:
array[i][j] = 6 + i + j
else:
array[i][j] = 9 + i + j
for i in range(3):
print(array[i])
for i in range(3):
for j in range(4):
print("%2d" %array[i][j], end = " ")
print()
array = [[0] * 3 for i in range(3)]
array
# +
for i in range(3):
for j in range(3):
array[i][j] = i + j
for i in range (3):
print(array[i])
# -
[2*i for i in range(1,11)]
# +
array = [[0] * 4 for i in range(4)]
for i in range(16):
array[i//4][i%4] = i
for i in range(4):
print(array[i])
# -
for i in range(3):
for j in range(4):
print("%2d" %array[i][j], end = " ")
print()
array = [[0] * 5 for i in range(5)]
array
for i in range(5):
for j in range(5):
array[i][j] = 1+i+j
array
import random
random.randrange(1,25)
# +
import random
array = [[random.randrange(1,25) for i in range(5)] for i in range(5)]
average = []
for i in range(5):
average.append(sum(array[i])/5)
if sum(average)/5 >= 12.5:
print(average,"Big")
else:
print(average,"Small")
# -
sentence = input("Enter the sentence: ")
# +
a = 0
b = 0
for i in range(len(sentence)):
if "A" <= sentence[i] <= "Z":
a += 1
elif "a" <= sentence[i] <= "z":
b += 1
print("sentence:",sentence,
"lower:", a, "upper:",b)
# -
a
b
sentence = input("insert sentence: ")
for i in range(65,91):
if (chr(i+32) not in sentence) and (chr(i) not in sentence):
print(chr(i), end="")
chr(99)
# +
BCDFGJKLNOPQUVWXZ
BDFGJKLNOPQUVWXYZ
C가 없는대신 Y가있네?
# -
sentence
chr(90+32)
# +
n = int(input("insert number: "))
num = [0] * (2 * n)
for i in range(0, n):
num[i] = i*2 + 1
for j in range(0, n):
num[2*n-1-j] = 2 + 2 * j
print(num)
# -
import math
array = [[16, 27], [39, 100], [19, 88]]
for i in range(3):
print(array[i])
# +
#가로 평균
row_avg = []
for i in range(3):
row_avg.append(math.floor(sum(array[i])/2))
row_avg
# +
# 세로 평균
sum1 = 0
for i in range(2):
sum2 = 0
for j in range(3):
sum2 += array[j][i]
sum1 += sum2
print(sum2 // 3, end = " ")
print()
# -
print(sum1/6)
word = input("insert word: 3")
# +
def isHandsome(str):
for i in range(len(word)):
if word[i] == word[len(word)-1-i]:
return True
else:
return False
if isHandsome(word) == True:
print("Handsome")
# -
def isHandsome(str):
for i in range(len(str)):
if str[i] != str
|
problems-review/note3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
dataset = pd.read_csv('C:\\Users\\srushti\\py proj\\student_scores.csv')
dataset.shape
dataset.head()
dataset.describe()
dataset.plot(x='Hours', y='Scores', style='o')
plt.title('Hours vs Percentage')
plt.xlabel('Hours Studied')
plt.ylabel('Percentage Score')
plt.show()
X = dataset.iloc[:, :-1].values
y = dataset.iloc[:, 1].values
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=0)
from sklearn.linear_model import LinearRegression
regressor = LinearRegression()
regressor.fit(X_train, y_train)
print(regressor.intercept_)
print(regressor.coef_)
y_pred = regressor.predict(X_test)
df = pd.DataFrame({'Actual': y_test, 'Predicted': y_pred})
print(df)
from sklearn import metrics
print('Mean Absolute Error:', metrics.mean_absolute_error(y_test, y_pred))
print('Mean Squared Error:', metrics.mean_squared_error(y_test, y_pred))
print('Root Mean Squared Error:', np.sqrt(metrics.mean_squared_error(y_test, y_pred)))
|
StudentsScore.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from keras.models import Sequential
# To initialize neural network
from keras.layers import Convolution2D
# Images are two dimensional, concolution step
from keras.layers import MaxPooling2D
# Pooling step
from keras.layers import Flatten
# Convert pools feature map into this large feature vector
from keras.layers import Dense
#To add fully connected layers
# +
#Initializing the CNN
#there is also a graph option but we'll use sequential ANN Model
classifier = Sequential()
#step 1 - Convolution
#creating the feature map by using feature detector from ınput image
classifier.add( Convolution2D(32,3,3, input_shape=(64,64,3), activation='relu'))
#32 Feature maps&detetctors uses 3 by 3 matrices, we can put 128 in the powerful machines
# +
#step -2 Pooling
classifier.add(MaxPooling2D(pool_size = (2,2)))
classifier.add( Convolution2D(32,3,3, input_shape=(64,64,3), activation='relu'))
classifier.add(MaxPooling2D(pool_size = (2,2)))
# -
#step -3 Flattening
classifier.add(Flatten())
#step-4 Full connection step
classifier.add(Dense(output_dim = 256, activation = 'relu'))
classifier.add(Dense(output_dim = 1, activation = 'sigmoid'))
#binary outcome
# +
#compiling the cnn
classifier.compile(optimizer='adam', loss='binary_crossentropy', metrics = ['accuracy'])
# +
#Fitting to CNN to the images
from keras.preprocessing.image import ImageDataGenerator
train_datagen = ImageDataGenerator(
rescale=1./255,
shear_range=0.2,
zoom_range=0.2,
horizontal_flip=True)
test_datagen = ImageDataGenerator(rescale=1./255)
training_set = train_datagen.flow_from_directory(
'dataset2/two/train',
target_size=(64, 64),
batch_size=32,
class_mode='binary')
test_set = test_datagen.flow_from_directory(
'dataset2/two/test',
target_size=(64, 64),
batch_size=32,
class_mode='binary')
results=classifier.fit_generator(
training_set,
samples_per_epoch=130,
nb_epoch=100,
validation_data=test_set,
nb_val_samples=18)
# +
import matplotlib.pyplot as plt
def plot_acc_loss(results, epochs):
acc = results.history['accuracy']
loss = results.history['loss']
val_acc = results.history['val_accuracy']
val_loss = results.history['val_loss']
plt.figure(figsize=(15, 5))
plt.subplot(121)
plt.plot(range(1,epochs), acc[1:], label='Train_acc')
plt.plot(range(1,epochs), val_acc[1:], label='Test_acc')
plt.title('Accuracy over' + str(epochs) + 'Epochs', size=15)
plt.legend()
plt.grid(True)
plt.subplot(122)
plt.plot(range(1,epochs), loss[1:], label='Train_loss')
plt.plot(range(1,epochs), val_loss[1:], label='Test_loss')
plt.title('Loss over' + str(epochs) + 'Epochs', size=15)
plt.legend()
plt.grid(True)
plt.show()
plot_acc_loss(results, 100)
# +
# Part 3 - Making new predictions
import numpy as np
from keras.preprocessing import image
test_image = image.load_img('dataset2/two/single_prediction/covid.jpeg', target_size = (64, 64))
test_image = image.img_to_array(test_image)
test_image = np.expand_dims(test_image, axis = 0)
result = classifier.predict(test_image)
print(training_set.class_indices)
# +
# %pylab inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
img=mpimg.imread('dataset2/two/single_prediction/covid.jpeg')
imgplot = plt.imshow(img)
plt=plt.title('Covid-19 Positive Chest X-ray ')
# +
if result[0][0] == 1:
prediction = 'normal'
else:
prediction = 'covid'
print("AI's prediction is: "+ prediction)
# -
test_image = image.load_img('dataset2/two/single_prediction/covid2.jpeg', target_size = (64, 64))
test_image = image.img_to_array(test_image)
test_image = np.expand_dims(test_image, axis = 0)
result = classifier.predict(test_image)
print(training_set.class_indices)
# +
# %pylab inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
img=mpimg.imread('dataset2/two/single_prediction/covid2.jpeg')
imgplot = plt.imshow(img)
plt=plt.title('Covid-19 Positive Chest X-ray ')
# +
if result[0][0] == 1:
prediction = 'normal'
else:
prediction = 'covid'
print("AI's prediction is: "+ prediction)
# +
# %pylab inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
test_image = image.load_img('dataset2/two/single_prediction/normal.jpeg', target_size = (64, 64))
test_image = image.img_to_array(test_image)
test_image = np.expand_dims(test_image, axis = 0)
result = classifier.predict(test_image)
img=mpimg.imread('dataset2/two/single_prediction/normal.jpeg')
imgplot = plt.imshow(img)
plt=plt.title('Covid-19 Negative Chest X-ray ')
# +
if result[0][0] == 1:
prediction = 'normal'
else:
prediction = 'covid'
print("AI's prediction is: "+ prediction)
# -
# !pip install opencv-python
# +
test_set.reset()
pred = classifier.predict_generator(test_set,18,verbose=1)
print('Predictions finished')
# -
|
Deep Learning/Image Dehazing using Tensorflow/Covid-CT-Lung.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
from IPython.display import Image
# -
# %matplotlib inline
Image('1584102211.png')
# ## 读取数据
train = pd.read_csv('data/used_car_train_20200313.csv', sep=' ')
train.head(2)
test = pd.read_csv('data/used_car_testA_20200313.csv', sep=' ')
test.head(2)
# 看下数据集样本数
train.shape, test.shape
# ## 探索性数据分析
# ### 缺失值检查
# 求空值数量
train.isnull().sum().sort_values(ascending=False).head()
# ### 数据分布
# 目标值分布,右偏分布
fig, ax = plt.subplots(figsize=(8, 6))
sns.set_style("white")
sns.distplot(train['price'])
train.describe().T
# 查看分类特征的类别数量
for i in ['name', 'model', 'brand', 'regionCode']:
print(i, train[i].nunique())
# 查看特征相关系数
corr = train.corr()
plt.subplots(figsize=(10, 8))
sns.heatmap(corr, cmap="Blues")
# ## 特征工程
# ### 处理倾斜特征
from scipy.stats import norm
# 目标值做log处理
train['price'] = np.log1p(train['price'])
# 查看转化后的分布,有点正态的感觉了
fig, ax = plt.subplots(figsize=(8, 7))
sns.distplot(train['price'], fit=norm)
# ### 移除异常值
# 可以根据计算结果或者其他特征进行移除
train.drop(train[train['price'] < 4].index, inplace=True)
# 整合训练集测试集以便后续特征工程
train_labels = train['price'].reset_index(drop=True)
train_features = train.drop(['price'], axis=1)
test_features = test
all_features = pd.concat([train_features, test_features]).reset_index(drop=True)
# ### 填充缺失值
# 根据最常出现填充,都是零,也可以根据其他来填充,不是瞎填的。。
def fill_missing(df):
df['fuelType'] = df['fuelType'].fillna(0)
df['gearbox'] = df['gearbox'].fillna(0)
df['bodyType'] = df['bodyType'].fillna(0)
df['model'] = df['model'].fillna(0)
return df
all_features = fill_missing(all_features)
all_features.isnull().sum().head()
# ### 数据类型转换
# 处理完
def data_astype(df):
# string
df['SaleID'] = df['SaleID'].astype(int).astype(str)
df['name'] = df['name'].astype(int).astype(str)
df['model'] = df['model'].astype(str)
df['brand'] = df['brand'].astype(str)
df['bodyType'] = df['bodyType'].astype(str)
df['fuelType'] = df['fuelType'].astype(str)
df['gearbox'] = df['gearbox'].astype(str)
df['notRepairedDamage'] = df['notRepairedDamage'].astype(str)
df['regionCode'] = df['regionCode'].astype(int).astype(str)
df['seller'] = df['seller'].astype(int).astype(str)
df['offerType'] = df['offerType'].astype(int).astype(str)
df['regDate'] = df['regDate'].astype(str)
df['creatDate'] = df['creatDate'].astype(str)
# date
df['creatDate'] = pd.to_datetime(df['creatDate'])
return df
all_features = data_astype(all_features)
# ### 编码分类变量
# 先删除掉一些不要的特征
all_features = all_features.drop(['SaleID', 'name', 'regDate', 'model', 'seller',
'offerType', 'creatDate', 'regionCode'], axis=1)
all_features = pd.get_dummies(all_features).reset_index(drop=True)
all_features.shape
# ### 重新创建训练集和测试集
X = all_features.iloc[:len(train_labels), :]
X_test = all_features.iloc[len(train_labels):, :]
X.shape, train_labels.shape, X_test.shape
# ## 训练模型
from sklearn.linear_model import Ridge, RidgeCV
from sklearn.model_selection import KFold, cross_val_score
# ### 设置交叉验证
# K折交叉验证
kf = KFold(n_splits=5, random_state=15, shuffle=True)
# 定义评测指标
def cv_mae(model, X):
mae = -cross_val_score(model, X, train_labels, scoring='neg_mean_absolute_error')
return mae
# ### 定义模型
ridge_alphas = [0.1, 1, 3, 5, 10]
ridge = RidgeCV(alphas=ridge_alphas, cv=kf)
# ### 训练模型
# 查看交叉验证分数
score = cv_mae(ridge, X)
score
score.mean()
ridge.fit(X, train_labels)
# 查看R的平方
ridge.score(X, train_labels)
# ### 模型预测
# 查看预测结果
fig, ax = plt.subplots(figsize=(8, 6))
sns.distplot(ridge.predict(X_test))
# 预测结果
submission = test[['SaleID']].copy()
submission['price'] = np.expm1(ridge.predict(X_test))
submission.shape
submission.to_csv('submission.csv', index=False)
|
baseline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ## Preprocessing of Loan Default Dataset
#
# ### Introduction
#
# We use the dataset of [Tianchi Competetion](https://tianchi.aliyun.com/competition/entrance/531830/information) to train our loan default rate estimation. In this notebook, we preprocess the dataset and generate features, which refers to some execellent work listed as below:
#
# * **Overview**: https://tianchi.aliyun.com/notebook-ai/detail?spm=5176.12586969.1002.6.3b30250fXUZ5fy&postId=129318
# * **EDA**: https://tianchi.aliyun.com/notebook-ai/detail?spm=5176.12586969.1002.12.3b30250fXUZ5fy&postId=129320
# * **Feature Eningeering**: https://tianchi.aliyun.com/notebook-ai/detail?spm=5176.12586969.1002.6.3b30b135z4zdwX&postId=129321
# +
import pandas as pd
import numpy as np
import datetime
import warnings
from tqdm import tqdm
from sklearn.preprocessing import LabelEncoder
pd.set_option('display.max_rows', None)
pd.set_option('display.max_columns', None)
warnings.filterwarnings('ignore')
# -
# ### Read data
#
# To download the dataset to your own s3 bucket:
#
# * Fill {YOUR_S3_BUCKET} and {YOUR_S3_PATH} with your preferred values in the following cell.
# * Uncomment the cell by removing the leading # character.
# * Execute the cell.
# +
# # !aws s3 cp ${MY_S3_BUCKET}/risk/tianchi/train.csv .
# # !aws s3 cp ${MY_S3_BUCKET}/risk/tianchi/testA.csv .
# -
data_train = pd.read_csv('./train.csv')
data_test_a = pd.read_csv('./testA.csv')
# numerical features and categorical features
numerical_fea = list(data_train.select_dtypes(exclude=['object']).columns)
category_fea = list(filter(lambda x: x not in numerical_fea, list(data_train.columns)))
label = 'isDefault'
numerical_fea.remove(label)
numerical_fea
category_fea
# ### Fill null values for numerical and categorical features seperately
data_train[numerical_fea].isnull().sum()
data_train[category_fea].isnull().sum()
# +
# for numerical features we use median values
data_train[numerical_fea] = data_train[numerical_fea].fillna(data_train[numerical_fea].median())
data_test_a[numerical_fea] = data_test_a[numerical_fea].fillna(data_train[numerical_fea].median())
# for categorical features we use mode values
data_train[category_fea] = data_train[category_fea].fillna(data_train[category_fea].mode())
data_test_a[category_fea] = data_test_a[category_fea].fillna(data_train[category_fea].mode())
# -
data_train.isnull().sum()
# ffill for null values `employmentLength`
data_train = data_train.fillna(axis=0, method='ffill')
data_test_a = data_test_a.fillna(axis=0, method='ffill')
data_train.isnull().sum()
# ### Transform `issueDate` into numerical values
for data in [data_train, data_test_a]:
data['issueDate'] = pd.to_datetime(data['issueDate'],format='%Y-%m-%d')
startdate = datetime.datetime.strptime('2007-06-01', '%Y-%m-%d')
data['issueDateDT'] = data['issueDate'].apply(lambda x: x-startdate).dt.days
data[:3]
# ### Transform `employmentLength` into numerical values
data_train['employmentLength'].value_counts(dropna=False).sort_index()
# +
def employmentLength_to_int(s):
if pd.isnull(s):
return s
else:
return np.int8(s.split()[0])
for data in [data_train, data_test_a]:
data['employmentLength'].replace(to_replace='10+ years', value='10 years', inplace=True)
data['employmentLength'].replace('< 1 year', '0 years', inplace=True)
data['employmentLength'] = data['employmentLength'].apply(employmentLength_to_int)
data['employmentLength'].value_counts(dropna=False).sort_index()
# -
# ### Transform `earliesCreditLine` into numerical values
data_train['earliesCreditLine'].sample(5)
# tranform earliesCreditLine into numerical values
for data in [data_train, data_test_a]:
data['earliesCreditLine'] = data['earliesCreditLine'].apply(lambda s: int(s[-4:]))
# ### Encode the categorical features
cate_features = ['grade', 'subGrade', 'employmentTitle', 'homeOwnership', 'verificationStatus', 'purpose', 'postCode', 'regionCode', \
'applicationType', 'initialListStatus', 'title', 'policyCode']
for f in cate_features:
print(f, 'different values:', data[f].nunique())
for data in [data_train, data_test_a]:
data['grade'] = data['grade'].map({'A':1,'B':2,'C':3,'D':4,'E':5,'F':6,'G':7})
for data in [data_train, data_test_a]:
data = pd.get_dummies(data, columns=['subGrade', 'homeOwnership', 'verificationStatus', 'purpose', 'regionCode'], drop_first=True)
# ### Outliers processing
def find_outliers_by_3segama(data,fea):
data_std = np.std(data[fea])
data_mean = np.mean(data[fea])
outliers_cut_off = data_std * 3
lower_rule = data_mean - outliers_cut_off
upper_rule = data_mean + outliers_cut_off
data[fea + '_outliers'] = data[fea].apply(lambda x:str('ExceptionValue') if x > upper_rule or x < lower_rule else 'NormalValue')
return data
for fea in numerical_fea:
data_train = find_outliers_by_3segama(data_train,fea)
print(data_train[fea + '_outliers'].value_counts())
print(data_train.groupby(fea + '_outliers')['isDefault'].sum())
print('-' * 60)
# Filter the exception values
for fea in numerical_fea:
data_train = data_train[data_train[fea+'_outliers']=='NormalValue']
data_train = data_train.reset_index(drop=True)
# ### Feature binning
# Feature binning
data['loanAmnt_bin1'] = np.floor_divide(data['loanAmnt'], 1000)
data['loanAmnt_bin2'] = np.floor(np.log10(data['loanAmnt']))
data['loanAmnt_bin3'] = pd.qcut(data['loanAmnt'], 10, labels=False)
for col in ['grade', 'subGrade']:
temp_dict = data_train.groupby([col])['isDefault'].agg(['mean']).reset_index().rename(columns={'mean': col + '_target_mean'})
temp_dict.index = temp_dict[col].values
temp_dict = temp_dict[col + '_target_mean'].to_dict()
data_train[col + '_target_mean'] = data_train[col].map(temp_dict)
data_test_a[col + '_target_mean'] = data_test_a[col].map(temp_dict)
# ### Feature interaction
# Mean and std
for df in [data_train, data_test_a]:
for item in ['n0','n1','n2','n4','n5','n6','n7','n8','n9','n10','n11','n12','n13','n14']:
df['grade_to_mean_' + item] = df['grade'] / df.groupby([item])['grade'].transform('mean')
df['grade_to_std_' + item] = df['grade'] / df.groupby([item])['grade'].transform('std')
# ### High dimensional feature encoding
# Label-encoding: subGrade, postCode, title
for col in tqdm(['employmentTitle', 'postCode', 'title','subGrade']):
le = LabelEncoder()
le.fit(list(data_train[col].astype(str).values) + list(data_test_a[col].astype(str).values))
data_train[col] = le.transform(list(data_train[col].astype(str).values))
data_test_a[col] = le.transform(list(data_test_a[col].astype(str).values))
print('Label Encoding Completed')
# ### Remove useless features
# Remove issueDate, id
for data in [data_train, data_test_a]:
data.drop(['issueDate','id'], axis=1,inplace=True)
features = [f for f in data_train.columns if f not in ['id','issueDate'] and '_outliers' not in f]
fg_data_train = data_train[features]
fg_data_train[:10]
fg_data_train.to_csv('fg_train_data.csv', sep=',', index=False, encoding='utf-8')
# ### Upload data
#
# To upload the dataset to your own s3 bucket:
#
# * Fill {YOUR_S3_BUCKET} and {YOUR_S3_PATH} with your preferred values in the following cell.
# * Uncomment the cell by removing the leading # character.
# * Execute the cell.
# +
# # !aws s3 cp ./fg_train_data.csv ${MY_S3_BUCKET}/risk/tianchi/
|
demo/dataset/tianchi_loan/fg.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Implementing a neural network in low-level tensorflow without eager execution
# This notebook implements a simple multilayer perceptron for the Boston Housing data.
#
# First we create placeholder tensors for the inputs and output variables.
import tensorflow as tf
x = tf.placeholder(dtype=tf.float32, shape=(None, 13))
y = tf.placeholder(dtype=tf.float32, shape=(None, 1))
# Next we create a function which will perform a linear transformation of its input. This takes care of creating the variable tensors for the biases and weights. Tensorflow keeps track of all the variable tensors we create, so there is no need for us to store them in a data structure. We could store them if we particularly wanted to refer to them later e.g. to inspect their value or to use them in a penalisation variable. (An alternative method is to use [tensor names](https://www.tensorflow.org/programmers_guide/variables).)
def linear_transform(inputs, ninputs, noutputs):
bias = tf.Variable(tf.random_normal([noutputs], stddev=0.05))
weights = tf.Variable(tf.random_normal([ninputs, noutputs], stddev=0.05))
return tf.matmul(inputs, weights) + bias
# Now we create our neural network. I'm using the $tanh$ activation function for the intermediate layers as this small neural network is especially vulnerable to dead ReLUs. For the final layer I use a softplus activation function to ensure the output is positive.
sizes = [5, 3, 1]
ninputs = 13
layer1 = tf.nn.tanh(linear_transform(x, ninputs, sizes[0]))
layer2 = tf.nn.tanh(linear_transform(layer1, sizes[0], sizes[1]))
pred = tf.nn.softplus(linear_transform(layer2, sizes[1], sizes[2]))
# Note that `pred` ends up being a 2-dimensional tensor, although it has only 1 column.
print(pred)
# We'll use mean squared error loss which can be implemented in Tensorflow as follows.
loss = tf.reduce_mean(tf.square(pred - y))
# Now we create optimiser and training operators.
optimiser = tf.train.AdamOptimizer()
train = optimiser.minimize(loss)
init = tf.global_variables_initializer()
# Next we start a Tensorflow session and initialise the variables.
sess = tf.Session()
sess.run(init)
# Below we get the Boston Housing data, normalise it and reshape the outputs into 2-dimensional tensors (which are required because they must match our `pred` tensor which is 2-dimensional as noted above).
# Download data
import numpy as np
from keras.datasets import boston_housing
(x_train, y_train), (x_val, y_val) = boston_housing.load_data()
# Normalise data
x_mean = np.std(x_train, 0)
x_sd = np.std(x_train, 0)
for i in range(13):
x_train[:,i] -= x_mean[i]
x_train[:,i] /= x_sd[i]
x_val[:,i] -= x_mean[i]
x_val[:,i] /= x_sd[i]
# Reshape data
y_train = y_train.reshape((404,1))
y_val = y_val.reshape((102,1))
# Let's split the data into batches.
nbatches = 10
xbatches = np.array_split(x_train, nbatches)
ybatches = np.array_split(y_train, nbatches)
# Finally we can train the neural network.
for e in range(1, 5000):
train_loss = 0.
for (x_,y_) in zip(xbatches, ybatches):
next_loss, _ = sess.run([loss, train], {x:x_, y:y_})
train_loss += next_loss
if e % 250 == 0:
train_loss /= nbatches
val_loss = sess.run(loss, {x:x_val, y:y_val})
print("Epochs {:3d} Train loss {:.1f} Validation loss {:.1f}".format(e, train_loss, val_loss))
# This run has achieved much better training loss than for linear regression and a similar validation loss - compare with Figure 2.1 of the notes. The difference between training and validation loss suggests significant overfitting. (On other training runs the evidence is even stronger due to a large validation loss!)
#
# Also note that compared to the eager execution version, this runs much faster and gives similar results.
#
# Let's look at our predictions.
predictions_train = sess.run(pred, {x:x_train})
predictions_val = sess.run(pred, {x:x_val})
import matplotlib.pyplot as plt
plt.plot(y_train, predictions_train, "bo", label="train")
plt.plot(y_val, predictions_val, "ro", label="validation")
plt.legend()
# The training predictions look significantly improved compared to the linear regression values in Figure 2.2. One improvement is that we've prevented any negative predictions by using a softplus activation. Another is that house values close to 50 are no longer badly underestimated.
#
# On the other hand the validation predictions contain several outlying predictions corroborating the earlier suspicion of overfitting. Regularisation of some sort would probably improve validation loss.
#
# This example illustrates that neural networks can improve on linear regression even for small datasets. However the major benefits are for big datasets!
|
practical exercises/solutions/E Boston Housing with low-level Tensorflow (not eager).ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/chitaha/Awesome-Geospatial/blob/master/satellite_geojson.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="OmDltBEAb1D_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 768} outputId="94b0f55f-e2bc-4066-998d-46b0bcc26dd6"
# !pip install geopandas
# !pip install geojson
# !pip install geojsonio
# + id="dbJSnh2Wb_mt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 615} outputId="a93575e0-9c4c-4ec1-a011-9f0847d0bc6b"
import geopandas as gpd
import geojsonio
states = gpd.read_file('states.geojson')
m_states = states[states['Name'].str.startswith('M')]
geojsonio.display(m_states.to_json())
|
satellite_geojson.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from alignment.optimizations import optimize_dim_subspaces
# ## Optimizing for finding $k_{X}^{*}$ and $k_{\hat{A}}^{*}$ ##
# This code runs on the fly, i.e., we don't save the results of randomizations of matrices.<br>
# Parameters in the function `optimize_dim_subspaces`:<br>
# `num_rdm` : how many realizations of randomization for each percent we consider.<br>
# `num_k` : how many possible $k_{X}$ and $k_{\hat{A}}$ we consider.
#
# Results give us:<br>
# i) a heatmap which indicates the difference of norm of distance matrix between no randomization and full randomization.<br>
# ii) $k_{X}^{*}$, $k_{\hat{A}}^{*}$ and $k_{Y}^{*}$
optimize_dim_subspaces(
dataset="constructive_example",
num_rdm=2,
num_k=5,
num_scanning=1,
norm_type="Frobenius-Norm",
log=False,
heatmap=True
)
|
alignment/demo.ipynb
|
# ##### Copyright 2020 Google LLC.
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
#
# # slitherlink
# <table align="left">
# <td>
# <a href="https://colab.research.google.com/github/google/or-tools/blob/master/examples/notebook/contrib/slitherlink.ipynb"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/colab_32px.png"/>Run in Google Colab</a>
# </td>
# <td>
# <a href="https://github.com/google/or-tools/blob/master/examples/contrib/slitherlink.py"><img src="https://raw.githubusercontent.com/google/or-tools/master/tools/github_32px.png"/>View source on GitHub</a>
# </td>
# </table>
# First, you must install [ortools](https://pypi.org/project/ortools/) package in this colab.
# !pip install ortools
# +
from ortools.constraint_solver import pywrapcp
from collections import deque
small = [[3, 2, -1, 3], [-1, -1, -1, 2], [3, -1, -1, -1], [3, -1, 3, 1]]
medium = [[-1, 0, -1, 1, -1, -1, 1, -1], [-1, 3, -1, -1, 2, 3, -1, 2],
[-1, -1, 0, -1, -1, -1, -1, 0], [-1, 3, -1, -1, 0, -1, -1, -1],
[-1, -1, -1, 3, -1, -1, 0, -1], [1, -1, -1, -1, -1, 3, -1, -1],
[3, -1, 1, 3, -1, -1, 3, -1], [-1, 0, -1, -1, 3, -1, 3, -1]]
big = [[3, -1, -1, -1, 2, -1, 1, -1, 1, 2], [1, -1, 0, -1, 3, -1, 2, 0, -1, -1],
[-1, 3, -1, -1, -1, -1, -1, -1, 3, -1],
[2, 0, -1, 3, -1, 2, 3, -1, -1, -1], [-1, -1, -1, 1, 1, 1, -1, -1, 3, 3],
[2, 3, -1, -1, 2, 2, 3, -1, -1, -1], [-1, -1, -1, 1, 2, -1, 2, -1, 3, 3],
[-1, 2, -1, -1, -1, -1, -1, -1, 2, -1],
[-1, -1, 1, 1, -1, 2, -1, 1, -1, 3], [3, 3, -1, 1, -1, 2, -1, -1, -1, 2]]
def NeighboringArcs(i, j, h_arcs, v_arcs):
tmp = []
if j > 0:
tmp.append(h_arcs[i][j - 1])
if j < len(v_arcs) - 1:
tmp.append(h_arcs[i][j])
if i > 0:
tmp.append(v_arcs[j][i - 1])
if i < len(h_arcs) - 1:
tmp.append(v_arcs[j][i])
return tmp
def PrintSolution(data, h_arcs, v_arcs):
num_rows = len(data)
num_columns = len(data[0])
for i in range(num_rows):
first_line = ''
second_line = ''
third_line = ''
for j in range(num_columns):
h_arc = h_arcs[i][j].Value()
v_arc = v_arcs[j][i].Value()
cnt = data[i][j]
first_line += ' ---' if h_arc else ' '
second_line += '|' if v_arc else ' '
second_line += ' ' if cnt == -1 else ' %i ' % cnt
third_line += '| ' if v_arc == 1 else ' '
termination = v_arcs[num_columns][i].Value()
second_line += '|' if termination else ' '
third_line += '|' if termination else ' '
print(first_line)
print(third_line)
print(second_line)
print(third_line)
last_line = ''
for j in range(num_columns):
h_arc = h_arcs[num_rows][j].Value()
last_line += ' ---' if h_arc else ' '
print(last_line)
class BooleanSumEven(pywrapcp.PyConstraint):
def __init__(self, solver, vars):
pywrapcp.PyConstraint.__init__(self, solver)
self.__vars = vars
self.__num_possible_true_vars = pywrapcp.NumericalRevInteger(0)
self.__num_always_true_vars = pywrapcp.NumericalRevInteger(0)
def Post(self):
for i in range(len(self.__vars)):
v = self.__vars[i]
if not v.Bound():
demon = self.Demon(BooleanSumEven.Update, i)
v.WhenBound(demon)
def InitialPropagate(self):
num_always_true = 0
num_possible_true = 0
possible_true_index = -1
for i in range(len(self.__vars)):
var = self.__vars[i]
if var.Min() == 1:
num_always_true += 1
num_possible_true += 1
elif var.Max() == 1:
num_possible_true += 1
possible_true_index = i
if num_always_true == num_possible_true and num_possible_true % 2 == 1:
self.solver().Fail()
if num_possible_true == num_always_true + 1:
self.__vars[possible_true_index].SetValue(num_always_true % 2)
self.__num_possible_true_vars.SetValue(self.solver(), num_possible_true)
self.__num_always_true_vars.SetValue(self.solver(), num_always_true)
def Update(self, index):
solver = self.solver()
value = self.__vars[index].Value()
if value == 0:
self.__num_possible_true_vars.Decr(solver)
else:
self.__num_always_true_vars.Incr(solver)
num_possible = self.__num_possible_true_vars.Value()
num_always = self.__num_always_true_vars.Value()
if num_always == num_possible and num_possible % 2 == 1:
solver.Fail()
if num_possible == num_always + 1:
possible_true_index = -1
for i in range(len(self.__vars)):
if not self.__vars[i].Bound():
possible_true_index = i
break
if possible_true_index != -1:
self.__vars[possible_true_index].SetValue(num_always % 2)
def DebugString(self):
return 'BooleanSumEven'
# Dedicated constraint: There is a single path on the grid.
# This constraint does not enforce the non-crossing, this is done
# by the constraint on the degree of each node.
class GridSinglePath(pywrapcp.PyConstraint):
def __init__(self, solver, h_arcs, v_arcs):
pywrapcp.PyConstraint.__init__(self, solver)
self.__h_arcs = h_arcs
self.__v_arcs = v_arcs
def Post(self):
demon = self.DelayedInitialPropagateDemon()
for row in self.__h_arcs:
for var in row:
var.WhenBound(demon)
for column in self.__v_arcs:
for var in column:
var.WhenBound(demon)
# This constraint implements a single propagation.
# If one point is on the path, it checks the reachability of all possible
# nodes, and zero out the unreachable parts.
def InitialPropagate(self):
num_rows = len(self.__h_arcs)
num_columns = len(self.__v_arcs)
num_points = num_rows * num_columns
root_node = -1
possible_points = set()
neighbors = [[] for _ in range(num_points)]
for i in range(num_rows):
for j in range(num_columns - 1):
h_arc = self.__h_arcs[i][j]
if h_arc.Max() == 1:
head = i * num_columns + j
tail = i * num_columns + j + 1
neighbors[head].append(tail)
neighbors[tail].append(head)
possible_points.add(head)
possible_points.add(tail)
if root_node == -1 and h_arc.Min() == 1:
root_node = head
for i in range(num_rows - 1):
for j in range(num_columns):
v_arc = self.__v_arcs[j][i]
if v_arc.Max() == 1:
head = i * num_columns + j
tail = (i + 1) * num_columns + j
neighbors[head].append(tail)
neighbors[tail].append(head)
possible_points.add(head)
possible_points.add(tail)
if root_node == -1 and v_arc.Min() == 1:
root_node = head
if root_node == -1:
return
visited_points = set()
to_process = deque()
# Compute reachable points
to_process.append(root_node)
while to_process:
candidate = to_process.popleft()
visited_points.add(candidate)
for neighbor in neighbors[candidate]:
if not neighbor in visited_points:
to_process.append(neighbor)
visited_points.add(neighbor)
if len(visited_points) < len(possible_points):
for point in visited_points:
possible_points.remove(point)
# Loop on unreachable points and zero all neighboring arcs.
for point in possible_points:
i = point // num_columns
j = point % num_columns
neighbors = NeighboringArcs(i, j, self.__h_arcs, self.__v_arcs)
for var in neighbors:
var.SetMax(0)
def SlitherLink(data):
num_rows = len(data)
num_columns = len(data[0])
solver = pywrapcp.Solver('slitherlink')
h_arcs = [[
solver.BoolVar('h_arcs[%i][%i]' % (i, j)) for j in range(num_columns)
] for i in range(num_rows + 1)]
v_arcs = [[
solver.BoolVar('v_arcs[%i][%i]' % (i, j)) for j in range(num_rows)
] for i in range(num_columns + 1)]
# Constraint on the sum or arcs
for i in range(num_rows):
for j in range(num_columns):
if data[i][j] != -1:
sq = [h_arcs[i][j], h_arcs[i + 1][j], v_arcs[j][i], v_arcs[j + 1][i]]
solver.Add(solver.SumEquality(sq, data[i][j]))
# Single loop: each node has a degree 0 or 2
zero_or_two = [0, 2]
for i in range(num_rows + 1):
for j in range(num_columns + 1):
neighbors = NeighboringArcs(i, j, h_arcs, v_arcs)
solver.Add(solver.Sum(neighbors).Member(zero_or_two))
# Single loop: sum or arcs on row or column is even
for i in range(num_columns):
column = [h_arcs[j][i] for j in range(num_rows + 1)]
solver.Add(BooleanSumEven(solver, column))
for i in range(num_rows):
row = [v_arcs[j][i] for j in range(num_columns + 1)]
solver.Add(BooleanSumEven(solver, row))
# Single loop: main constraint
solver.Add(GridSinglePath(solver, h_arcs, v_arcs))
# Special rule on corners: value == 3 implies 2 border arcs used.
if data[0][0] == 3:
h_arcs[0][0].SetMin(1)
v_arcs[0][0].SetMin(1)
if data[0][num_columns - 1] == 3:
h_arcs[0][num_columns - 1].SetMin(1)
v_arcs[num_columns][0].SetMin(1)
if data[num_rows - 1][0] == 3:
h_arcs[num_rows][0].SetMin(1)
v_arcs[0][num_rows - 1].SetMin(1)
if data[num_rows - 1][num_columns - 1] == 3:
h_arcs[num_rows][num_columns - 1].SetMin(1)
v_arcs[num_columns][num_rows - 1].SetMin(1)
# Search
all_vars = []
for row in h_arcs:
all_vars.extend(row)
for column in v_arcs:
all_vars.extend(column)
db = solver.Phase(all_vars, solver.CHOOSE_FIRST_UNBOUND,
solver.ASSIGN_MAX_VALUE)
log = solver.SearchLog(1000000)
solver.NewSearch(db, log)
while solver.NextSolution():
PrintSolution(data, h_arcs, v_arcs)
solver.EndSearch()
|
examples/notebook/contrib/slitherlink.ipynb
|