code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Reads one file
# This jupyter notebook shows how to use the **open_dataset** function of **GOES** package, to read and get the information from **one** GOES-16/17 file.
#
# Index:
# - [Reads file](#reads_file)
# - [Gets attribute from file](#gets_attribute)
# - [Gets variable from file](#gets_variable)
# - [Gets image from file](#gets_image)
# - [Gets dimension from file](#gets_dimension)
# <a id='reads_file'></a>
# ## Reads file
# Sets path and name of file that will be read.
path = '/home/joao/Downloads/GOES-16/ABI/'
file = 'OR_ABI-L2-CMIPF-M6C13_G16_s20200782000176_e20200782009496_c20200782010003.nc'
# Import the GOES package.
import GOES
# Reads file.
ds = GOES.open_dataset(path+file)
# Display the content of file.
print(ds)
# <a id='gets_attribute'></a>
# ## Gets attribute from file
# **Attribute** is a string parameters with information about the file. To get one attribute from file, write the follow:
title = ds.attribute('title')
print(title)
orbital = ds.attribute('orbital_slot')
print(orbital)
resol = ds.attribute('spatial_resolution')
print(resol)
# <a id='gets_variable'></a>
# ## Gets variable from file
# **Variable** is a python class that content a parameter with theirs attributes. Write the follow to get a variable:
hsat = ds.variable('nominal_satellite_height')
print(hsat)
# Print one attribute of parameter.
print(hsat.units)
# Print the parameter value:
print(hsat.data)
# In the above example, the parameter is a simple number, for this reason, when the method **data** was used, the value was printed directly.
# \
# **Get other variable:**
times = ds.variable('time_bounds')
print(times)
# Print the parameter value:
print(times.data)
# In this case, the parameter is an array and has two elements. Use the position index to select one of them.
print(times.data[1])
# Print one attribute of parameter.
print(times.long_name)
# <a id='gets_image'></a>
# ## Gets image from file
# **Image** is a Python class that contains a parameter that has the dimensions ('y', 'x'). Write the follow to get a image:
CMI, Lons, Lats = ds.image('CMI')
print(CMI)
# Print the parameter value:
print(CMI.data)
# Print one attribute of parameter.
print(CMI.standard_name)
# <a id='gets_dimension'></a>
# ## Gets dimension from file
# **Dimensions** is a class with the spatial attributes of the variables. To get one dimension from file, write the follow:
dim_x = ds.dimension('x')
print(dim_x)
# Theirs attributes are **name** and **size**.
print(dim_x.name)
print(dim_x.size)
| examples/v3.2/reads_one_file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Matplotlib.pyplot.boxplot Python Library Package Investigation - AOC G00364756
#
# ## Date created: 18/11/2018
#
# ## Introduction¶
# This is a project tasked to students of the Data Analytics Higher Diploma as delivered by Dr. <NAME> and provided by GMIT. The objective of this project is to investigate the "boxplot" python package of the "matplotlib.pyplot" python library by describing the purpose of boxplots and describing alternatives to the boxplot. This will be done through use of a Jupyter Notebook and Python with the aim of demonstrating how the functions operate in a clear and engaging format. A clear history of the work conducted in this project will be available on the Github repository "52446---Fundamentals-of-Data-Analysis---Project" submitted to the "Fundamentals of Data Analysis" GMIT moodle page. All works drawn upon to aid in the creation of this submission will be referenced within the "References" section of this Jupyter Notebook (please see the table of contents). The due date for this assignment is the 14/12/2018.
#
# [1](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.boxplot.html?highlight=boxplot#matplotlib.pyplot.boxplot) : See References
# ## Contents
# 1. Problem statement
# 2. History of the "boxplot" and situations in which it is used
# 3. Demonstrate the use of the box plot using selected data
# 4. Explain any relevant terminology such as the terms quartile and percentile
# 5. Compare the box plot to alternatives
# * Violinplots (Iris Dataset)
# * Beanplot (Iris Dataset)
# 6. Results and Conclusions
# * Results
# * Conclusions
# 7. References
# ## 1. Problem statement
# The box plot is common in data analysis for investigating individual numerical variables.
# In this project, you will investigate and explain box plots and their uses. The boxplot
# function from the Python package matplotlib.pyplot can be used to create box plots.
# Your submission should be in the form of a repository containing a Jupyter notebook in
# which you detail your findings. In your notebook, you should:
# * Summarise the history of the box plot and situations in which it used.
# * Demonstrate the use of the box plot using data of your choosing.
# * Explain any relevant terminology such as the terms quartile and percentile.
# * Compare the box plot to alternatives.
# ## 2. History of the "boxplot" and situations in which it is used
#
# The boxplot was invented by mathematician <NAME>. Tukey first introduced the boxplot in 1970 as a tool for data analysis but the boxplot didn't gain notoriety until his formal publication "Exploratory Data Analysis" in 1977. The boxplot is a form of summary statistic that has been widely adopted in data analytics since its inception. The boxplot is a graphical representation of the shape of a distribution, its central value (the median), and its variability. The boxplot is used in situations where the mean and standard deviation are not appropriate to use. The mean and standard deviation are most appropriate in representing the centre and spread for symmetrical distributions with no outliers. Where distributions contain extreme values and are highly skewed, the boxplot is often considered a more appropriate statistical tool to use as it is less influenced by these conditions, however it should not be used as a sole measure of spread as scaling of data and missing data points can lead to misleading boxplots. The boxplot has evolved over the years. Experts in statistics have contributed enhancements to the boxplot since its acceptance into the world of statistics. An example of an enhancement is the variable width boxplot, this boxplot gives tha analyst a visual representation of the size of the distribution by varying the width of the "box". Conventional boxplots did not include this element as they were drawn by hand whereas nowadays computers have encouraged creativity, allowing data analysts to show more information about a distribution through an evolved form of the boxplot, all the while keeping the compact nature of the boxplot. This is important as the original idea of the boxplot was to be a simple summary statistic representing the main features of a distribution.
#
# <img src="Boxplot_explained.JPG" width="600" height="500" align="center">
#
# [2](https://www150.statcan.gc.ca/n1/edu/power-pouvoir/ch12/5214889-eng.htm) : See References
#
# [3](http://mathworld.wolfram.com/Box-and-WhiskerPlot.html) : See References
#
# [4](https://en.wikipedia.org/wiki/Box_plot) : See References
#
# [5](https://en.wikipedia.org/wiki/John_Tukey) : See References
#
# [6](http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.48.4362&rep=rep1&type=pdf) : See References
#
# [7](http://vita.had.co.nz/papers/boxplots.pdf) : See References
# ## 3. Demonstrate the use of the box plot using selected data
# Below is a demonstration of the use of boxplots on the Iris Flower dataset.
#
# [8](https://en.wikipedia.org/wiki/Iris_flower_data_set) : See References
# +
# Import all python packages necessary to analyse the dataset
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# Read the iris dataset csv file.
df = pd.read_csv("iris.csv", names = ["sepal_length", "sepal_width", "petal_length", "petal_width", "class"])
# +
# magic code that debugs some issues observed with matplotlib
# %matplotlib inline
# Plotting boxplots of sepal length, sepal width, petal length and petal width for each Iris class.
# Setting the colours for the different classes of iris, adapted from: https://python-graph-gallery.com/33-control-colors-of-boxplot-seaborn/
my_pal = {"Iris-versicolor": "b", "Iris-setosa": "r", "Iris-virginica":"g"}
# Creates subplots in a 1 row by 4 column format, the '1' means that the code preceeding will apply to the 1st subplot.
# Subplot 1: Boxplot of Sepal Length for the different species of Iris in the Iris dataset.
plt.subplot(141)
sns.boxplot(x=df['class'], y=df["sepal_length"], whis=1.5, flierprops=dict(markerfacecolor='r', marker='s'), orient="v", palette=my_pal)
plt.title("Sepal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
# Subplot 2: Boxplot of Sepal width for the different species of Iris in the Iris dataset.
plt.subplot(142)
sns.boxplot(x=df['class'], y=df["sepal_width"], whis=1.5, flierprops= dict(markerfacecolor='r', marker='s'), orient="v", palette=my_pal)
plt.title("Sepal Width", fontsize=25)
plt.xlabel("Iris class",fontsize=20)
plt.ylabel("centimetres")
# Subplot 3: Boxplot of Petal Length for the different species of Iris in the Iris dataset.
plt.subplot(143)
sns.boxplot(x=df['class'], y=df['petal_length'], whis=1.5, orient="v", palette=my_pal)
plt.title("Petal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
# Subplot 4: Boxplot of Petal Width for the different species of Iris in the Iris dataset.
plt.subplot(144)
sns.boxplot(x=df['class'], y=df['petal_width'], whis=1.5, orient="v", palette=my_pal)
plt.title("Petal Width", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
plt.subplots_adjust(left=0, bottom=0.1, right=3, top=2, wspace=0.30, hspace=0.30)
plt.show()
# -
# ## 4. Explain any relevant terminology such as the terms quartile and percentile
# This sections describes some of the definitions and terminology encountered in the investigation of boxplots and alternative to the boxplot. Please see below an explaination of some of these definitions:
#
# A **quartile** is a definition of the four equal groups into which a population can be divided according to the distribution of values of a particular variable.
#
# [9](https://www.investopedia.com/terms/q/quartile.asp) : See References
#
# A **percentile** is the definition given to each of the 100 equal groups into which a population can be divided according to the distribution of values of a particular variable.
#
# [10](https://www.statisticshowto.datasciencecentral.com/probability-and-statistics/percentiles-rank-range/) : See References
#
# **Outliers** are individual points that lie outside the extremes of the dataset.
#
# [7](http://vita.had.co.nz/papers/boxplots.pdf) : 40 years of boxplots - <NAME> and <NAME>
#
# The **median** is the middle value of a dataset, its the value that separates the top half of the dataset from the bottom half of the dataset e.g. (1,1,6,8,14,15,19), in this instance 8 is the median for the given dataset.
#
# [11](https://en.wikipedia.org/wiki/Median) : See References
#
# A **rug** plot is a one dimensional scatter plot, displayed as marks along an axis, used to visualise the distribution of the data.
#
# [12](https://en.wikipedia.org/wiki/Rug_plot) : See References
#
# **Multimodality** in statistics, is when a distribution has more than one peak.
#
# [13](https://www.statisticshowto.datasciencecentral.com/multimodal-distribution/) : See References
# ## 5. Compare the box plot to alternatives
# Violinplots and Beanplots are alternatives to the use of Boxplots. They are evolutions of the boxplot containing more information about a distribution, while keeping the compact nature of the boxplot. Each are described in detail in the following sections...
# ### Violinplots (Iris Dataset)
# Like boxplots, violinplots contain the quartiles and median of a dataset but combine these with a density trace which is mirrored to form a polygon shape. This allows a visual representation of the distribution. Individual outliers are not visible in a violin plot. This should be taken into account when using a violinplot to discribe a distribution, to ensure an accurate representation of the distribution. Violinplots can be more useful than boxplots when comparing datasets, depending on their complexity.
#
# [14](http://pyinsci.blogspot.com/2009/09/violin-plot-with-matplotlib.html) : See References
#
# [16](https://cran.r-project.org/web/packages/beanplot/vignettes/beanplot.pdf) : See References
#
# <h3 align="center">Violinplot</h3>
# <img src="violinplot.png" alt="Violinplot" style="width:50%">
#
# Below is a demonstration of the use of violinplots on the Iris Flower dataset...
# +
# Plotting violinplots of sepal length, sepal width, petal length and petal width for each Iris class.
# Setting the colours for the different classes of iris, adapted from: https://python-graph-gallery.com/33-control-colors-of-boxplot-seaborn/
my_pal = {"Iris-versicolor": "b", "Iris-setosa": "r", "Iris-virginica":"g"}
# Creates subplots in a 1 row by 4 column format, the '1' means that the code preceeding will apply to the 1st subplot.
# Subplot 1: Violinplot of Sepal Length for the different species of Iris in the Iris dataset.
plt.subplot(141)
sns.violinplot( inner="quart", x=df['class'], y=df["sepal_length"], whis=1.5, orient="v", palette=my_pal)
plt.title("Sepal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
# Subplot 2: Boxplot of Sepal width for the different species of Iris in the Iris dataset.
plt.subplot(142)
sns.violinplot( inner="quart", x=df['class'], y=df["sepal_width"], whis=1.5, orient="v", palette=my_pal)
plt.title("Sepal Width", fontsize=25)
plt.xlabel("Iris class",fontsize=20)
plt.ylabel("centimetres")
# Subplot 3: Boxplot of Petal Length for the different species of Iris in the Iris dataset.
plt.subplot(143)
sns.violinplot( inner="quart", x=df['class'], y=df['petal_length'], whis=1.5, orient="v", palette=my_pal)
plt.title("Petal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
# Subplot 4: Boxplot of Petal Width for the different species of Iris in the Iris dataset.
plt.subplot(144)
sns.violinplot( inner="quart", x=df['class'], y=df['petal_width'], whis=1.5, orient="v", palette=my_pal)
plt.title("Petal Width", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
plt.subplots_adjust(left=0, bottom=0.1, right=3, top=2, wspace=0.30, hspace=0.30)
plt.show()
# -
# ### Beanplots (Iris Dataset)
# A beanplot, so the name suggests, is a plot made up of multiple batches or "beans". Each bean consists of a density trace, which is mirrored to form a polygon shape. Violinplots are very similar to beanplots in this regard. The key difference between the violinplot and the beanplot is that the beanplot contains a rug plot, (please see section 4 of this jupyter notebook which contains definitions of certain terminology), within the polygon shape, showing the individual data points within the dataset as lines on an axes. A boxplot will not clearly indicate the difference between distributions through use of the inter-quartile range. However if a beanplot is used the analyst can clearly see the difference between the distributions, due to the polygons formed by the density trace and the idividual data points from the rug plot within.
#
# [15](https://jnlnet.files.wordpress.com/2008/11/beanplots.jpg) : See References
# [7](http://vita.had.co.nz/papers/boxplots.pdf) : See References
# [16](https://cran.r-project.org/web/packages/beanplot/vignettes/beanplot.pdf) : See References
#
# <img src="beanplots.jpg" alt="Beanplot" style="width:40%">
#
# Below is a demonstration of the use of beanplots on the Iris Flower dataset...
# +
#Plotting a beanplots of sepal length, sepal width, petal length and petal width for each Iris class.
sns.violinplot(data=df['sepal_length'], scale="count", inner="stick")
# Setting the colours for the different classes of iris, adapted from: https://python-graph-gallery.com/33-control-colors-of-boxplot-seaborn/
my_pal = {"Iris-versicolor": "b", "Iris-setosa": "r", "Iris-virginica":"g"}
# Creates subplots in a 1 row by 4 column format, the '1' means that the code preceeding will apply to the 1st subplot.
# Subplot 1: Beanplot of Sepal Length for the different species of Iris in the Iris dataset.
plt.subplot(141)
sns.violinplot(x=df['class'], y=df["sepal_length"], whis=1.5, scale="count", inner="stick", orient="v", palette=my_pal)
plt.title("Sepal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
# Subplot 2: Beanplot of Sepal width for the different species of Iris in the Iris dataset.
plt.subplot(142)
sns.violinplot( x=df['class'], y=df["sepal_width"], whis=1.5, scale="count", inner="stick", orient="v", palette=my_pal)
plt.title("Sepal Width", fontsize=25)
plt.xlabel("Iris class",fontsize=20)
plt.ylabel("centimetres")
# Subplot 3: Beanplot of Petal Length for the different species of Iris in the Iris dataset.
plt.subplot(143)
sns.violinplot( x=df['class'], y=df['petal_length'], whis=1.5,scale="count", inner="stick", orient="v", palette=my_pal)
plt.title("Petal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
# Subplot 4: Beanplot of Petal Width for the different species of Iris in the Iris dataset.
plt.subplot(144)
sns.violinplot( x=df['class'], y=df['petal_width'], whis=1.5, orient="v", scale="count", inner="stick", palette=my_pal)
plt.title("Petal Width", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
plt.subplots_adjust(left=0, bottom=0.1, right=3, top=2, wspace=0.30, hspace=0.30)
plt.show()
# -
# ## 6. Results and Conclusions
# ### Results
# Below we can compare a boxplot, a violinplot and a beanplot of the Sepal Length for the Iris dataset.<br/>
# From the boxplot, the position of the whiskers and the box convey that the datasets are not majorly skewed. The Virginica dataset is the slightly skewed in the 0cm direction and contains one possible outlier representated as a red square outside the whisker.<br/>
# The violinplot describes the distributions a little more showing the density trace of the datasets, the density of the datasets can be interpreted from the varying shapes of the polygons. It seems that the majority of the values for the Setosa class lie around the value 5 which appears to be the median value also. It is also interesting to point out that the violinplot does not represent the possible outlier in the Virginica dataset that was previously identified by the boxplot. <br/>
# The beanplot is identical to the violinplot but represents the individual data points as a rug plot inside the polygon, we can observe the spread of the data points in this way. The setosa data points seem to be evenly spread whereas the Virginica values seem to be skewed in the 0cm direction. The rug plot information while interesting does not represent multimodality effectively. It is also interesting to point out that the beanplot does not represent the possible outlier in the Virginica dataset that was previously identified by the boxplot. <br/>
# <br/>
# Boxplots are beneficial in identifying possible outliers but not the overall density of a distribution or multimodality of the distribution.<br/>
# Violinplots and beanplots are beneficial in giving a visual representation of the density of the distribution, the violinplot also shows the inter-quartile range and the median of the dataset.<br/>
# The beanplot shows each individual data point giving granularity to the distribution but lacks the mean and interquartile range of the boxplot and violinplot which can be important information. Violinplots and beanplots are also poor at identifying outliers.
#
# [17](https://greenbookblog.org/2018/03/21/replacing-boxplots-and-histograms-with-rugs-violins-and-bean-plots/) : See References
# +
# Plotting a boxplot, violinplot and beanplot for sepal length to easily compare the three.
# Subplot 1: Boxplot of Sepal width for the different species of Iris in the Iris dataset.
plt.subplot(131)
sns.boxplot(x=df['class'], y=df["sepal_length"], whis=1.5, flierprops=dict(markerfacecolor='r', marker='s'), orient="v", palette=my_pal)
plt.title("Sepal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
# Subplot 2: Violinplot of Sepal Length for the different species of Iris in the Iris dataset.
plt.subplot(132)
sns.violinplot( inner="quart", x=df['class'], y=df["sepal_length"], whis=1.5, orient="v", palette=my_pal)
plt.title("Sepal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
# Subplot 3: Beanplot of Sepal Length for the different species of Iris in the Iris dataset.
plt.subplot(133)
sns.violinplot(x=df['class'], y=df["sepal_length"], whis=1.5, scale="count", inner="stick", orient="v", palette=my_pal)
plt.title("Sepal Length", fontsize=25)
plt.xlabel("Iris class", fontsize=20)
plt.ylabel("centimetres")
plt.subplots_adjust(left=0, bottom=0.1, right=3, top=2, wspace=0.30, hspace=0.30)
plt.show()
# -
# ### Conclusions
# In an era before the pandemic use of computers for data analytics, boxplots were an extremely useful tool in analysing data as they were quick to create by hand and represented some of the main attributes of a dataset in visual form. While the existance of alternatives has seen a reduction in the use of the traditional boxplot, it still remains a valuable tool in analysing data and in certain scenarios may still be the most appropriate tool for summarising datasets. The benefits and limiting factors of each method described in this investigation are clearly demonstrated above and the term "horses for courses" is applicable when choosing any of these methods to represent a dataset (each method needs to be matched to a purpose).
# ## 7.References
#
# [1](https://matplotlib.org/api/_as_gen/matplotlib.pyplot.boxplot.html?highlight=boxplot#matplotlib.pyplot.boxplot) : Matplotlib.pyplot.boxplot documentation
#
# [2](https://www150.statcan.gc.ca/n1/edu/power-pouvoir/ch12/5214889-eng.htm) : Statistics Canada - Constructing boxplots and whisper plots
#
# [3](http://mathworld.wolfram.com/Box-and-WhiskerPlot.html) : Wolfram Mathworld - Box and whisper plots
#
# [4](https://en.wikipedia.org/wiki/Box_plot) : Wikipedia - Boxplots webpage
#
# [5](https://en.wikipedia.org/wiki/John_Tukey) : Wikipedia - John W. Tukey webpage
#
# [6](http://citeseerx.ist.psu.edu/viewdoc/download?doi=10.1.1.48.4362&rep=rep1&type=pdf) : Discussion paper - Misleading or Confusing Boxplots, <NAME>
#
# [7](http://vita.had.co.nz/papers/boxplots.pdf) : 40 years of boxplots - <NAME> and <NAME>
#
# [8](https://en.wikipedia.org/wiki/Iris_flower_data_set) : Wikipedia - Iris Flower Datset
#
# [9](https://www.investopedia.com/terms/q/quartile.asp) : Definition of Quartile
#
# [10](https://www.statisticshowto.datasciencecentral.com/probability-and-statistics/percentiles-rank-range/) : Definition of Percentile
#
# [11](https://en.wikipedia.org/wiki/Median) : Wikipedia - Median
#
# [12](https://en.wikipedia.org/wiki/Rug_plot) : Wikipedia - Rug plot
#
# [13](https://www.statisticshowto.datasciencecentral.com/multimodal-distribution/) : Statisticshowto - Multimodal distributions
#
# [14](http://pyinsci.blogspot.com/2009/09/violin-plot-with-matplotlib.html) : Pyinsci - Violin Plot Image
#
# [15](https://jnlnet.files.wordpress.com/2008/11/beanplots.jpg) : JNLnet - Bean PLot Image
#
# [16](https://cran.r-project.org/web/packages/beanplot/vignettes/beanplot.pdf) : Beanplot: A Boxplot Alternative for Visual
# Comparison of Distributions - <NAME>
#
# [17](https://greenbookblog.org/2018/03/21/replacing-boxplots-and-histograms-with-rugs-violins-and-bean-plots/) : Greenbook blog post, <NAME> - Replacing Boxplots and Histograms, with Rugs, Violins & Bean Plots
# # END
| AOC_G00364756_52466---Fundamentals of Data Analysis---Project 2018.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''work'': conda)'
# language: python
# name: python3
# ---
# + [markdown] id="ID2g9XHMvAXO"
# # Generate PDF/A files from docTR output
#
# These files have also a readable text layer on top of the image, which can be used to search in a document with any PDF-Viewer.
# + id="sPMz5UYUvAXQ" outputId="6b4284e0-0efe-4443-911b-840e90145716"
# !pip3 install git+https://github.com/mindee/doctr.git
# !pip3 install reportlab>=3.6.2
# optional if you want to merge multiple pdfs
# !pip3 install PyPDF2==1.26.0
# + id="ZC8sIbEZvAXS"
# Imports
import base64
import re
from tempfile import TemporaryDirectory
from math import atan, cos, sin
from typing import Dict, Optional, Tuple
from xml.etree import ElementTree as ET
from xml.etree.ElementTree import Element
import numpy as np
import PyPDF2
from PyPDF2 import PdfFileMerger
from doctr.io import DocumentFile
from doctr.models import ocr_predictor
from PIL import Image
from reportlab.lib.colors import black
from reportlab.lib.units import inch
from reportlab.lib.utils import ImageReader
from reportlab.pdfgen.canvas import Canvas
# + [markdown] id="Aa9PoeOdvAXS"
# ## Define the hOCR (xml) parser
# At the beginning, we have to define a HocrParser which is able to parse the exported XML-Tree/File.
#
# [hOCR convention](https://github.com/kba/hocr-spec/blob/master/1.2/spec.md)
# + id="pblMOnCovAXT"
class HocrParser():
def __init__(self):
self.box_pattern = re.compile(r'bbox((\s+\d+){4})')
self.baseline_pattern = re.compile(r'baseline((\s+[\d\.\-]+){2})')
def _element_coordinates(self, element: Element) -> Dict:
"""
Returns a tuple containing the coordinates of the bounding box around
an element
"""
out = out = {'x1': 0, 'y1': 0, 'x2': 0, 'y2': 0}
if 'title' in element.attrib:
matches = self.box_pattern.search(element.attrib['title'])
if matches:
coords = matches.group(1).split()
out = {'x1': int(coords[0]), 'y1': int(
coords[1]), 'x2': int(coords[2]), 'y2': int(coords[3])}
return out
def _get_baseline(self, element: Element) -> Tuple[float, float]:
"""
Returns a tuple containing the baseline slope and intercept.
"""
if 'title' in element.attrib:
matches = self.baseline_pattern.search(
element.attrib['title']).group(1).split()
if matches:
return float(matches[0]), float(matches[1])
return (0.0, 0.0)
def _pt_from_pixel(self, pxl: Dict, dpi: int) -> Dict:
"""
Returns the quantity in PDF units (pt) given quantity in pixels
"""
pt = [(c / dpi * inch) for c in pxl.values()]
return {'x1': pt[0], 'y1': pt[1], 'x2': pt[2], 'y2': pt[3]}
def _get_element_text(self, element: Element) -> str:
"""
Return the textual content of the element and its children
"""
text = ''
if element.text is not None:
text += element.text
for child in element:
text += self._get_element_text(child)
if element.tail is not None:
text += element.tail
return text
def export_pdfa(self,
out_filename: str,
hocr: ET.ElementTree,
image: Optional[np.ndarray] = None,
fontname: str = "Times-Roman",
fontsize: int = 12,
invisible_text: bool = True,
dpi: int = 300):
"""
Generates a PDF/A document from a hOCR document.
"""
width, height = None, None
# Get the image dimensions
for div in hocr.findall(".//div[@class='ocr_page']"):
coords = self._element_coordinates(div)
pt_coords = self._pt_from_pixel(coords, dpi)
width, height = pt_coords['x2'] - \
pt_coords['x1'], pt_coords['y2'] - pt_coords['y1']
# after catch break loop
break
if width is None or height is None:
raise ValueError("Could not determine page size")
pdf = Canvas(out_filename, pagesize=(width, height), pageCompression=1)
span_elements = [element for element in hocr.iterfind(".//span")]
for line in span_elements:
if 'class' in line.attrib and line.attrib['class'] == 'ocr_line' and line is not None:
# get information from xml
pxl_line_coords = self._element_coordinates(line)
line_box = self._pt_from_pixel(pxl_line_coords, dpi)
# compute baseline
slope, pxl_intercept = self._get_baseline(line)
if abs(slope) < 0.005:
slope = 0.0
angle = atan(slope)
cos_a, sin_a = cos(angle), sin(angle)
intercept = pxl_intercept / dpi * inch
baseline_y2 = height - (line_box['y2'] + intercept)
# configure options
text = pdf.beginText()
text.setFont(fontname, fontsize)
pdf.setFillColor(black)
if invisible_text:
text.setTextRenderMode(3) # invisible text
# transform overlayed text
text.setTextTransform(
cos_a, -sin_a, sin_a, cos_a, line_box['x1'], baseline_y2)
elements = line.findall(".//span[@class='ocrx_word']")
for elem in elements:
elemtxt = self._get_element_text(elem).strip()
# replace unsupported characters
elemtxt = elemtxt.translate(str.maketrans(
{'ff': 'ff', 'ffi': 'ffi', 'ffl': 'ffl', 'fi': 'fi', 'fl': 'fl'}))
if not elemtxt:
continue
# compute string width
pxl_coords = self._element_coordinates(elem)
box = self._pt_from_pixel(pxl_coords, dpi)
box_width = box['x2'] - box['x1']
font_width = pdf.stringWidth(elemtxt, fontname, fontsize)
# Adjust relative position of cursor
cursor = text.getStartOfLine()
dx = box['x1'] - cursor[0]
dy = baseline_y2 - cursor[1]
text.moveCursor(dx, dy)
# suppress text if it is 0 units wide
if font_width > 0:
text.setHorizScale(100 * box_width / font_width)
text.textOut(elemtxt)
pdf.drawText(text)
# overlay image if provided
if image is not None:
pdf.drawImage(ImageReader(Image.fromarray(image)),
0, 0, width=width, height=height)
pdf.save()
# + [markdown] id="hzpt5nHFvAXe"
# ## OCR the files and show the results
#
# Now we are ready to start the OCR process and show the results.
# + id="HjhRaZ6tvAXe"
# Download a sample
# !wget https://www.allianzdirect.de/dam/documents/home/Versicherungsbedingungen-08-2021.pdf
# Read the file
docs = DocumentFile.from_pdf("Versicherungsbedingungen-08-2021.pdf").as_images()
model = ocr_predictor(det_arch='db_resnet50', reco_arch='crnn_vgg16_bn', pretrained=True)
# we will grab only the first two pages from the pdf for demonstration
result = model(docs[:2])
result.show(docs)
# + [markdown] id="MiA8N21GvAXf"
# ## Export as PDF/A
# In this section we will export our documents as PDF/A files.
#
# We show 3 possible options for this.
# -
# ### All as single PDF/A file
# All files will be saved as single file.
# + id="2dDy5t1UvAXf"
# returns: list of tuple where the first element is the (bytes) xml string and the second is the ElementTree
xml_outputs = result.export_as_xml()
# init the above parser
parser = HocrParser()
# iterate through the xml outputs and images and export to pdf/a
# the image is optional else you can set invisible_text=False and the text will be printed on a blank page
for i, (xml, img) in enumerate(zip(xml_outputs, docs)):
xml_element_tree = xml[1]
parser.export_pdfa(f'{i}.pdf', hocr=xml_element_tree, image=img)
# -
# ### All merged into one PDF/A file
# All PDF/A files will be merged into one PDF/A file.
# + id="SkEZrL-hvAXg"
# returns: list of tuple where the first element is the (bytes) xml string and the second is the ElementTree
xml_outputs = result.export_as_xml()
# init the above parser
parser = HocrParser()
# you can also merge multiple pdfs into one
merger = PdfFileMerger()
for i, (xml, img) in enumerate(zip(xml_outputs, docs)):
xml_element_tree = xml[1]
with TemporaryDirectory() as tmpdir:
parser.export_pdfa(f'{tmpdir}/{i}.pdf', hocr=xml_element_tree, image=img)
merger.append(f'{tmpdir}/{i}.pdf')
merger.write(f'docTR-PDF.pdf')
# -
# ### All as base64 encoded PDF/A files
# All PDF/A files will be saved as base64 strings in a list.
# + id="owWVIPcKvAXg" outputId="ad9b4eda-146b-4ab1-c613-06af4afce64b"
# returns: list of tuple where the first element is the (bytes) xml string and the second is the ElementTree
xml_outputs = result.export_as_xml()
# init the above parser
parser = HocrParser()
# or encode the pdfs into base64 (Rest API usage)
base64_encoded_pdfs = list()
for i, (xml, img) in enumerate(zip(xml_outputs, docs)):
xml_element_tree = xml[1]
with TemporaryDirectory() as tmpdir:
parser.export_pdfa(f'{tmpdir}/{i}.pdf',
hocr=xml_element_tree, image=img)
with open(f'{tmpdir}/{i}.pdf', 'rb') as f:
base64_encoded_pdfs.append(base64.b64encode(f.read()))
print(f'{len(base64_encoded_pdfs)} pdfs encoded')
# -
# ## How can I use a PDF/A?
# You can open the saved pdf's with any PDF-Viewer and type some words you are searching for in the document.
#
# Matches will be highlighted in the text layer.
#
# Or you use Python to search, for example words in the text layer.
# +
# search specific words in the pdf and print all matches
pattern = "Allianz"
file_name = "docTR-PDF.pdf"
reader = PyPDF2.PdfFileReader(file_name)
num_pages = reader.getNumPages()
for i in range(0, num_pages):
page = reader.getPage(i)
text = page.extractText()
for match in re.finditer(pattern, text):
print(f'Page no: {i} | Match: {match}')
# -
# ## To go further
# [Wikipedia PDF/A](https://en.wikipedia.org/wiki/PDF/A)
#
# [Difference between PDF/A and PDF](https://askanydifference.com/difference-between-pdf-a-and-pdf/)
#
# ### Happy Coding :)
| doctr/export_as_pdfa.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#Child Nutrient Calculator - Project
Name = str(input("Enter Name:"))
Age=int(input("Enter Age:"))
Gender=str(input("Enter Gender:"))
Height=float(input("Enter Height(cms):"))#input will be in cms
Weight=float(input("Enter Weight(kgs):")) #input will be in kgs because user should be feel comfortable
#we have given user inputs normal details
print()
print('\033[1m'+ "Calorie Consumption"+ '\033[0m')
print()
Milk=int(input("Enter quantity of milk intake:"))
Vegetable= int(input("Enter quantity of vegetables intake: "))
Lentils= int(input("Enter quantity of lentils intake: "))
Egg= int(input("Enter quantity of egg intake: "))
Rice= int(input("Enter quantity of rice intake: "))
Meat= int(input("Enter quantity of meat intake: "))
#we have given user input for calorie consumption in grams
cal_Milk=(100/100)*Milk;
cal_Vegetable=(85/100)*Vegetable;
cal_Lentils=(113/100)*Lentils;
cal_Egg=(155/100)*Egg;
cal_Rice=(130/100)*Rice;
cal_Meat=(143/100)*Meat;
#we have converted into calories
Inches=Height/2.54
Pounds= Weight*2.20462262
BMI=Pounds/(Inches**2)*703
#BMI formula used
if BMI < 16:
x="Severely Underweight"
elif BMI>=16 and BMI< 18.5:
x="Underweight"
elif BMI>=18.5 and BMI<25:
x="Healthy"
elif BMI>=25 and BMI<30:
x="Overweight"
elif BMI>=30:
x="Obese"
else:
print("You have entered incorrect inputs")
print("Please check you inputs again")
formatted_BMI="{:.2f}".format(BMI)#To represent float in 2 decimals we have used this.
print(f"BMI of {Name} is {formatted_BMI} and he(she) is {x}.")
# this are condition for BMI
calories = cal_Milk+cal_Vegetable+cal_Lentils+cal_Egg+cal_Rice+cal_Meat
#Calorie calculations
if (Age>=0 and Age<=2) and calories < 800:
print(f"The daily calorie consumption of {Name} is {calories} and child is under-nourished.")
elif (Age>=0 and Age<=2) and calories >= 800:
print(f"The daily calorie consumption of {Name} is {calories} and child is well-nourished.")
if (Age>=3 and Age<=4) and calories < 1400:
print(f"The daily calorie consumption of {Name} is {calories} and child is under-nourished.")
elif (Age>=3 and Age<=4) and calories >= 1400:
print(f"The daily calorie consumption of {Name} is {calories} and child is well-nourished.")
if (Age>=5 and Age<=8) and calories < 1800:
print(f"The daily calorie consumption of {Name} is {calories} and child is under-nourished.")
elif (Age>=5 and Age<=8) and calories>= 1800:
print(f"The daily calorie consumption of {Name} is {calories} and child is well-nourished.")
#this are conditions for calories consumption
# -
| Child Nutrient Calculator - Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# importing Libraries
import math
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.preprocessing import LabelEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression
from sklearn.ensemble import RandomForestRegressor
from sklearn.metrics import mean_squared_error
from sklearn.metrics import accuracy_score , r2_score
from sklearn.model_selection import GridSearchCV
# -
# Read the data
data=pd.read_csv("insurance.csv")
data.head()
# Get the shape of data
data.shape
# data.describe()
#finding the null values
data.isnull().sum()
# Droping the duplicate values.
data.drop_duplicates()
#In this graph we are calculating the sum of all the charges based on the regions
charges = data['charges'].groupby(data.region).sum().sort_values(ascending = True)
f, ax = plt.subplots(1, 1, figsize=(8, 6))
ax = sns.barplot(charges.head(), charges.head().index, palette='Blues')
#In this graph we are counting the age based on the sex
f, ax = plt.subplots(1, 1, figsize=(15, 10))
sns.countplot(x='age',hue='sex',data=data,palette="Set1")
plt.title('countplot of age based on sex')
f, ax = plt.subplots(1, 1, figsize=(12, 8))
ax = sns.barplot(x='region', y='charges', hue='sex', data=data, palette='Blues')
#As sex and smoker are categorical applied labelEncoder to convert to numerical
label = LabelEncoder()
data['sex']= label.fit_transform(data['sex'])
data['smoker'] = label.fit_transform(data['smoker'])
data.head()
#appling one hot encoding to convert the categorical to numerical
var= pd.get_dummies(data['region'])
#Appending the one hot encoded value to the original dataframe
data[['northeast','northwest','southeast']]=var[['northeast','northwest','southeast']]
data=data.drop('region',axis=1)
#as other column standarized between 0 and 1 so converting age,bmi in the range
sc = StandardScaler()
data[['age','bmi','charges']]=sc.fit_transform(data[['age','bmi','charges']])
#Seperating the features and label
x=data.drop('charges',axis=1)
y=data['charges']
#Spliting the data into train and test
x_train, x_test, y_train, y_test = train_test_split(x, y, test_size=0.3, random_state=0)
# +
# Extracting best parameter values of RandomForestRegressor by using GridSearchCV.
rfr_parameters = {'n_estimators' : [10, 20, 50, 100],
'max_depth' : [3, 5, 7, 9, 10]
}
grid_search_rfr = GridSearchCV(estimator = RandomForestRegressor(),
param_grid = rfr_parameters,
cv = 10,
n_jobs = -1)
grid_search_rfr.fit(x_train, y_train)
rfr = grid_search_rfr.best_estimator_
# -
# printing best parameter values.
print(rfr)
regressor = RandomForestRegressor(n_estimators= 50
,random_state=0, max_depth=3)
regressor.fit(x_train,y_train)
predict=regressor.predict(x_test)
# Checking MSE, RMSE and r2_Score
print(mean_squared_error(y_test, predict))
print(math.sqrt(mean_squared_error(y_test, predict)))
print(r2_score(y_test,predict))
plt.scatter(predict,y_test)
| RFR.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # Aprendizado Supervisionado
#
# Vamos começar o estudo da aprendizagem de máquina com o tipo de aprendizado denominado de **Supervisionado**.
#
# > * **Aprendizado Supervisionado (*Supervised Learning*):** A training set of examples with the correct responses (targets) is provided and, based on this training set, the algorithm generalises to respond correctly to all possible inputs. This also colled learning from exemplars.
#
# Um algoritmo supervisionado é uma função que, dado um conjunto de exemplos rotulados, constrói um *preditor*. Os rótulos atribuídos aos exemplos são definidos a partir de um domínio conhecido. Se este domínio for um conjunto de valores nominais, estamos lidando com um problema de *classificação*. Agora se este domínio for um conjunto infinito e ordenado de valores, passamos a lidar com um problema de *regressão*. O preditor construído recebe nomes distintos a depender da tarefa. Chamamos de classificador (para o primeiro tipo de rótulo) ou regressor (para o segundo tipo).
#
# Um classificador (ou regressor) também é uma função que recebe um exemplo não rotulado e é capaz de definir um rótulo dentro dos valores possíveis. Se estivermos trabalhando com um problema de regressão este rótulo está dentro do intervalo real assumido no problema. Se for uma tarefa de classificação, esse rótulo é uma das classes definidas.
#
# Podemos definir formalmente da seguinte maneira, segundo (FACELI, et. al, 2011):
#
# *Uma definição formal seria, dado um conjunto de observações de pares $D=\{(x_i, f(x_i)), i = 1, ..., n\}$, em que $f$ representa uma função desconhecida, um algoritmo de AM preditivo (ou supervisionado) aprende uma aproximação $f'$ da função desconhecida $f$. Essa função aproximada, $f'$, permite estimar o valor de $f$ para novas observações de $x$.*
#
# Temos duas situações para $f$:
#
# * **Classificação:** $y_i = f(x_i) \in \{c_1,...,c_m\}$, ou seja, $f(x_i)$ assume valores em um conjunto discreto, não ordenado;
# * **Regressão:** $y_i = f(x_i) \in R$, ou seja, $f(x_i)$ assume valores em um cojunto infinito e ordenado de valores.
#
#
# ## Regressão Linear
#
# Vamos mostrar como a regressão funciona através de um método denominado regressão linear. Esse tutorial é baseado em 3 materiais:
#
# * Tutorial de Regressão Linear: https://github.com/justmarkham/DAT4/blob/master/notebooks/08_linear_regression.ipynb por
# * Slides sobre Regressão Linear: http://pt.slideshare.net/perone/intro-ml-slides20min
# * Cap. 3 do Livro "An Introduction to Statistical Learning" disponível em: http://www-bcf.usc.edu/~gareth/ISL/
# * Livro "Inteligência Artificial - Uma Abordagem de Aprendizado de Máquina" disponível em: https://www.amazon.com.br/dp/8521618808/ref=cm_sw_r_tw_dp_x_MiGdybV5B9TTT
#
# Para o nosso trabalho, vamos trabalhar com a base de *Advertising* disponibilizada pelo livro *"An Introduction to Statistical Learning"*. Essa base consiste de 3 atributos que representam os gastos de propaganda (em milhares de dólares) de um determinado produto na TV, Rádio e Jornal. Além disso, é conhecido a quantidade de vendas realizadas (em milhares de unidades) para cada instância. Vamos explorar a base de ados a seguir:
# + deletable=true editable=true
# Imports necessários para a parte de Regressão Linear
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.model_selection import cross_val_score
# %matplotlib inline
# + [markdown] deletable=true editable=true
# O primeiro passo é carregar a base de dados. Ela está disponível a partir do link: http://www-bcf.usc.edu/~gareth/ISL/Advertising.csv. Para carregar a base vamos utilizar a biblioteca [**Pandas**](http://pandas.pydata.org/). Detalhes dessa biblioteca estão fora do escopo destes tutoriais. Sendo assim, apenas vamos usá-la sem tecer muitos detalhes sobre as operações realizadas. Basicamente, vamos utiliza-la para carregar a base de arquivos e plotar dados nos gráficos. Mais informações podem ser encontradas na documentação da biblioteca.
# + deletable=true editable=true
# Carrega a base e imprime os dez primeiros registros da base
data = pd.read_csv("http://www-bcf.usc.edu/~gareth/ISL/Advertising.csv", index_col=0)
print(data.head(10))
# + [markdown] deletable=true editable=true
# O *dataset* possui 3 atributos: *TV*, *Radio* e *Newspaper*. Cada um deles corresponde a quantidade de dólares gastos em propaganda em cada uma das mídias para um produto específico. Já a responsta (*Sales*) consiste da quantidade de produtos vendidos para cada produto. Esse *dataset* possui 200 instâncias.
#
# Para melhor visualizar, vamos plotar as informações da base de dados.
# + deletable=true editable=true
fig, axs = plt.subplots(1, 3, sharey=True)
data.plot(kind='scatter', x='TV', y='sales', ax=axs[0], figsize=(16, 8))
data.plot(kind='scatter', x='radio', y='sales', ax=axs[1])
data.plot(kind='scatter', x='newspaper', y='sales', ax=axs[2])
# + [markdown] deletable=true editable=true
# O nosso objetivo é analisar os dados e tirar certas conclusões a partir deles. Basicamente, queremos responder as seguintes perguntas:
#
# *** Com base nestes dados, como poderíamos gastar o dinheiro designado para propaganda no futuro? ***
#
# Em outras palavras:
#
# * *Existe uma relação entre o dinheiro gasto em propaganda e a quantidade de vendas?*
# * *Quão forte é esse relacionamento?*
# * *Quais são os tipos de propaganda que contribuem para as vendas?*
# * *Qual o efeito de cada tipo de propaganda nas vendas?*
# * *Dado um gasto específico em propaganda, é possível prever quanto será vendido?*
#
# Para explorar essas e outras questões, vamos utilizar, inicialmente, uma **Regressão Linear Simples**.
#
# + [markdown] deletable=true editable=true
# ### Regressão Linear Simples
#
# Como o próprio nome diz, a regressão linear simples é um método muito (muito++) simples para prever valores **(Y)** a partir de uma única variável **(X)**. Para este modelo, é assumido que existe uma aproximação linear entre X e Y. Matematicamente, podemos escrever este relacionamento a partir da seguinte função:
#
# $Y \approx \beta_0 + \beta_1X$, onde $\approx$ pode ser lido como *aproximadamente*.
#
# $\beta_0$ e $\beta_1$ são duas constantes desconhecidas que representam a intercepção da reta com o eixo vertical ($\beta_0$) e o declive (coeficiente angular) da reta ($\beta_1$). As duas constantes são conhecidas como coeficientes ou parâmetros do modelo. O propósito da regressão linear é utilizar o conjunto de dados conhecidos para estimar os valores destas duas variáveis e definir o modelo aproximado:
#
# $\hat{y} = \hat{\beta_0} + \hat{\beta_1}x$,
#
# onde $\hat{y}$ indica um valor estimado de $Y$ baseado em $X = x$. Com essa equação podemo prever, neste caso, as vendas de um determinado produto baseado em um gasto específico em propaganda na TV.
#
# Mas como podemos estimar estes valores?
# + [markdown] deletable=true editable=true
# ### Estimando Valores
#
# Na prática, $\beta_0$ e $\beta_1$ são desconhecidos. Para que a gente possa fazer as estimativas, devemos conhecer os valores destes atributos. Para isso, vamos utilizar os dados já conhecidos.
#
# Considere,
#
# $(x_1,y_1), (x_2,y_2), ..., (x_n, y_n)$ $n$ pares de instâncias observadas em um conjunto de dados. O primeiro valor consiste de uma observação de $X$ e o segundo de $Y$. Na base de propagandas, esses dados consistem dos 200 valores vistos anteriormente.
#
# O objetivo na construção do modelo de regressão linear é estimar os valores de $\beta_0$ e $\beta_1$ tal que o modelo linear encontrado represente, da melhor foma, os dados disponibilizados. Em outras palavras, queremos encontrar os valores dos coenficientes de forma que a reta resultante seja a mais próxima possível dos dados utilizados.
#
# Basicamente, vamos encontrar várias retas e analisar qual delas se aproxima mais dos dados apresentados. Existem várias maneiras de medir essa "proximidade". Uma delas é a RSS (*residual sum of squares*), que é representada pela equação:
#
# $\sum_{i=1}^{N}{(\hat{y_i}-y_i)^2}$, onde $\hat{y_i}$ o valor estimado de y e $y_i$, o valor real.
#
# A figura a seguir apresenta um exemplo que mostra os valores estimados e a diferença residual.
#
# 
#
# Os pontos vermelhos representam os dados observados; a linha azul, o modelo construído e as linhas cinzas, a diferença residual entre o que foi estimado e o que era real.
#
# Vamos estimar tais parâmetros utilizando *scikit-learn*.
# + [markdown] deletable=true editable=true
# ### Aplicação do modelo de regressão linear
# + [markdown] deletable=true editable=true
# O primeiro passo é separar dos dados (*features*) das classes (*labels*) dos dados que serão utilizados para treinar nosso modelo.
# + deletable=true editable=true
# Carregando os dados de treinamento e os labels
feature_cols = ['TV']
X = data[feature_cols] # Dados de Treinamento
y = data.sales # Labels dos dados de Treinamento
# + [markdown] deletable=true editable=true
# Em seguida, vamos instanciar o modelo de Regressão Linear do ScikitLearn e treina-lo com os dados.
# + deletable=true editable=true
lm = LinearRegression() # Instanciando o modelo
lm.fit(X, y) # Treinando com os dados de treinamento
# + [markdown] deletable=true editable=true
# Como dito anteriormente, o modelo aprendeu, baseado no conjunto de dados, valores para $\beta_0$ e $\beta_1$. Vamos visualizar os valores encontrados.
# + deletable=true editable=true
#Imprimindo beta_0
print("Valor de Beta_0: " + str(lm.intercept_))
#Imprimindo beta_1
print("Valor de Beta_1: " + str(lm.coef_[0]))
# + [markdown] deletable=true editable=true
# Esse valores representam os valores de $\beta_0$ e $\beta_1$ da nossa equação que representa um modelo simples de regressão linear onde é levado em consideração somente um atributo.
#
# Com esses valores é possível estimar quanto será vendido dado um determinado gasto em propaganda de TV. Além disso, o coeficiente $\beta_1$ nos conta mais sobre o problema.
#
# O valor de $0.047536640433$ indica que cada unidade que aumentarmos em propaganda de TV implica em um aumento de $0.047536640433$ nas vendas. Em outras palavras, cada $1,000$ gastos em TV está associado com um aumento de 47.537 de unidades nas vendas.
#
# Vamos usar esses valores para estimar quanto será vendido se gastarmos $50000$ em TV.
#
# $y = 7.03259354913 + 0.047536640433 \times 50$
#
#
# + deletable=true editable=true
7.03259354913+0.047536640433*50
# + [markdown] deletable=true editable=true
# Desta forma, poderíamos prever a venda de 9409 unidades.
#
# No entanto, nosso objetivo não é fazer isso manualmente. A idéia é construir o modelo e utiliza-lo para fazer a estimativa de valores. Para isso, vamos utilizar o método *predict*.
#
# Podemos estimar para uma entrada apenas:
# + deletable=true editable=true
lm.predict([[50]])
# + [markdown] deletable=true editable=true
# Ou várias:
# + deletable=true editable=true
lm.predict([[50], [200], [10]])
# + [markdown] deletable=true editable=true
# Para entender melhor como a Regressão Linear funciona vamos visualizar no gráfico o modelo construído.
# + deletable=true editable=true
'''
O código a seguir faz a predição para o menor e maior valores de X na base de treinamento. Estes valores
são utilizados para construir uma reta que é plotada sobre os dados de treinamento.
'''
X_new = pd.DataFrame({'TV': [data.TV.min(), data.TV.max()]}) # Menor e Maior valores de X na base de treinamento
preds = lm.predict(X_new) # Predição destes valores
data.plot(kind='scatter', x='TV', y='sales') # Plotagem dos valores da base de treinamento
plt.plot(X_new, preds, c='red', linewidth=2) # Plotagem da reta
# + [markdown] deletable=true editable=true
# A reta em vermelho representa o modelo de regressão linear construído a partir dos dados passados.
# + [markdown] deletable=true editable=true
# ### Avaliando o Modelo Construído
#
# Para avaliar o modelo construído vamos utilizar uma métrica denominada de $R^2$ (*R-squared* ou coeficiente de determinação).
#
# (By [Wikipedia](https://pt.wikipedia.org/wiki/R%C2%B2))
# *O coeficiente de determinação, também chamado de R², é uma medida de ajustamento de um modelo estatístico linear generalizado, como a Regressão linear, em relação aos valores observados. O R² varia entre 0 e 1, indicando, em percentagem, o quanto o modelo consegue explicar os valores observados. Quanto maior o R², mais explicativo é modelo, melhor ele se ajusta à amostra. Por exemplo, se o R² de um modelo é 0,8234, isto significa que 82,34\% da variável dependente consegue ser explicada pelos regressores presentes no modelo.*
#
# Para entender melhor a métrica, vamos analisar o gráfico a seguir:
# 
# *Fonte da Imagem: https://github.com/justmarkham/DAT4/ *
#
#
# Observe que a função representada pela cor vemelha se ajusta melhor aos dados do que as retas de cor azul e verde. Visualmente podemos ver que, de fato, a curva vemelha descreve melhor a distribuição dos dados plotados.
#
# Vamos calcular o valor do *R-squared* para o modelo construído utilizando o método *score* que recebe como parâmetro os dados de treinamento.
#
# + deletable=true editable=true
lm.score(X, y)
# + [markdown] deletable=true editable=true
# Sozinho esse valor não nos conta muito. No entanto, ele será bastante útil quando formos comparar este modelo com outros mais à frente.
# + [markdown] deletable=true editable=true
# ### Multiple Linear Regression
#
# Podemos estender o modelo visto anteriormente para trabalhar com mais de um atributo, a chamada *Multiple Linear Regression*. Matematicamente, teríamos:
#
# $y \approx \beta_0 + \beta_1 x_1 + ... \beta_n x_n$
#
# Cada $x$ representa um atribuito e cada atributo possui seu próprio coeficiente. Para nossa base de dados, teríamos:
#
# $y \approx \beta_0 + \beta_1 \times TV + \beta_2 \times Radio + \beta_3 \times Newspaper$
#
# Vamos construir nosso modelo para este caso:
# + deletable=true editable=true
# Carregando os dados de X e y do dataset
feature_cols = ['TV','radio','newspaper']
X = data[feature_cols]
y = data.sales
#Instanciando e treinando o modelo de regressão linear
lm = LinearRegression()
lm.fit(X, y)
#Imprimindo os coeficientes encontrados
print("Valor de Beta_0: ")
print(str(lm.intercept_))
print()
print("Valores de Beta_1, Beta_2, ..., Beta_n: ")
print(list(zip(feature_cols, lm.coef_)))
# + [markdown] deletable=true editable=true
# O modelo construído foi:
#
# $y \approx 2.93888936946 + 0.045764645455397601_1 \times TV + 0.18853001691820448 \times Radio -0.0010374930424762578 \times Newspaper$
# + [markdown] deletable=true editable=true
# Assim como fizemos no primeiro exemplo, podemos utilzar o método *predict* para prever valores não conhecidos.
# + deletable=true editable=true
lm.predict([[100, 25, 25], [200, 10, 10]])
# + [markdown] deletable=true editable=true
# Avaliando o modelo, temos o valor para o $R^2$:
# + deletable=true editable=true
lm.score(X, y)
# + [markdown] deletable=true editable=true
# ### Entendendo os resultados obtidos
#
# Vamos analisar alguns resultados obtidos nos dois modelos construídos anteriormente. A primeira coisa é verificar o valor de $\beta_1$. Esse valor deu positivo para os atributos *TV* e *Radio* e negativo para o atributo *Newspaper*. Isso significa que o gasto em propaganda está relacionado positivamente às vendas nos dois primeiros atributos. Diferente do que acontece com o *Newspaper*: o gasto está negativamente associado às vendas.
#
# Uma outra coisa que podemos perceber é que o *R-squared* aumentou quando aumentamos o número de atributos. Isso normalmente acontece com essa métrica. Basicamente, podemos concluir que este último modelo tem um valor mais alto para o *R-squared* que o modelo anterior que considerou apenas a TV como atributo. Isto significa que este modelo fornece um melhor "ajuste" aos dados fornecidos.
#
# No entanto, o *R-squared* não é a melhor métrica para avaliar tais modelos. Se fizermos um análise estatística mais aprofundada (essa análise foge do escopo desta disciplina. Detalhes podem ser encontrados [aqui](https://github.com/justmarkham/DAT4/blob/master/notebooks/08_linear_regression.ipynb)) vamos perceber que o atributo *Newspaper* não influencia (estatisticamente) no total de vendas. Teoricamente, poderíamos descartar tal atributo. No entanto, se calcularmos o valor do *R-squared* para um modelo sem *Newspaper* e para o modelo com *Newspaper*, o valor do segundo será maior que o primeiro.
#
# ** Essa tarefa fica como atividade ;) **
# + [markdown] deletable=true editable=true
# ## KNN: k-Nearest Neighbors
# + [markdown] deletable=true editable=true
# No início desse tutorial tratamos o problema de aprendizado supervisionado a partir de dois pontos de vista. O primeiro da regressão. Mostramos como trabalhar com a regressão linear para prever valores em um intervalo. O segundo, como o problema de classificação de instâncias em classes. Para exemplificar esse problema, vamos trabalhar com o KNN, uma das técnicas mais simples de classificação.
#
# A idéia básica do KNN é que podemos classificar uma instância desconhecida com base nas informações dos vizinhos mais próximos. Para isso, exergamos os dados como pontos marcados em um sistema cartesiano e utilizamos a distância entre pontos para identificar quais estão mais próximoas.
#
# Para entender um pouco mais do KNN, assista [este vídeo](https://www.youtube.com/watch?v=UqYde-LULfs)
#
# Para começar, vamos analisar o conjunto de dados a seguir:
# + deletable=true editable=true
data = pd.read_csv("http://www.data2learning.com/datasets/basehomemulher.csv", index_col=0)
data
# + [markdown] deletable=true editable=true
# Os dados representam informações de altura e peso coletadas de homens e mulheres. Se plotarmos tais informações no gráfico, temos:
# + deletable=true editable=true
colors = {0:'red', 1:'blue'}
# Plotagem dos valores da base de treinamento
data.plot(kind='scatter', x='peso', y='altura',c=data['classe'].apply(lambda x: colors[x]))
# + [markdown] deletable=true editable=true
# Considere que com base nestes dados, desejamos classificar uma nova instância. Vamos considerar uma instância onde a altura seja 1.70 e o peso 50. Se plotarmos esse ponto no gráfico, temos (a nova instância está representada pelo x):
# + deletable=true editable=true
colors = {0:'red', 1:'blue'}
# Plotagem dos valores da base de treinamento
data.plot(kind='scatter', x='peso', y='altura',c=data['classe'].apply(lambda x: colors[x]))
plt.plot([50], [1.70], 'x', c='green')
# + [markdown] deletable=true editable=true
# O KNN vai classificar a nova instância com base nos vizinhos mais próximos. Neste caso, a nova instância seria classificada como mulher. Essa comparação é feita com os $k$ vizinhos mais próximos.
#
# Por exemplo, se considerarmos 3 vizinhos mais próximos e destes 3, dois são mulheres e 1 homem, a instância seria classificada como mulher já que corresponde a classe da maioria dos vizinhos.
# + [markdown] deletable=true editable=true
# A distância entre dois pontos pode ser calculada de diversas formas. A biblioteca do ScikitLearn lista [uma série de métricas de distância](http://scikit-learn.org/stable/modules/generated/sklearn.neighbors.DistanceMetric.html) que podem ser usadas. Vamos considerar um novo ponto e simular o que o algoritmo do KNN faz.
# + deletable=true editable=true
colors = {0:'red', 1:'blue'}
# Plotagem dos valores da base de treinamento
data.plot(kind='scatter', x='peso', y='altura',c=data['classe'].apply(lambda x: colors[x]))
plt.plot([77], [1.68], 'x', c='green')
# + [markdown] deletable=true editable=true
# Vamos trabalhar com o ponto **{'altura': 1.68, 'peso':77}** e calcular sua distância para todos os demais pontos. No exemplo vamos usar a distância euclideana: $\sqrt{\sum{(x_1 - x_2)^2 + (y_1 - y_2)^2}}$. Para simplificar, vamos utilizar nossa própria implementação da distância euclideana.
# + deletable=true editable=true
import math
# Calcula a distância euclideana de dois pontos
def euclideanDistance(instance1, instance2, length):
distance = 0
for x in range(length):
distance += pow((instance1[x] - instance2[x]), 2)
return math.sqrt(distance)
# Só para visualização: converte os valores para labels em String
def convert_label(value):
if value == 0.0: return 'Mulher'
else: return 'Homem'
# 0 = mulher , 1 = homem
for index, row in data.iterrows():
print(convert_label(row['classe']), '%0.2f' % euclideanDistance([row['peso'], row['altura']],[77, 1.68], 2))
# + [markdown] deletable=true editable=true
# Uma vez que calculamos a distância do novo ponto a todos os demais pontos da base, devemos verificar os $k$ pontos mais próximos e ver qual classe predomina nestes pontos. Considerando os 3 vizinhos mais próximos ($k=3$), temos:
#
# * Homem: 10.0
# * Homem: 14.0
# * Mulher: 15.0
#
# Sendo assim, a instância selecionada seria classificada como **Homem**.
#
# E se considerássemos $k=5$?
#
# * Homem: 10.0
# * Homem: 14.0
# * Mulher: 15.0
# * Mulher: 17.0
# * Mulher: 24.0
#
# Neste caso, a instância seria classificada como **Mulher**.
# + deletable=true editable=true
import math
# Calcula a distância euclideana de dois pontos
def euclideanDistance(instance1, instance2, length):
distance = 0
for x in range(length):
distance += pow((instance1[x] - instance2[x]), 2)
return math.sqrt(distance)
# Só para visualização: converte os valores para labels em String
def convert_label(value):
if value == 0.0: return 'Mulher'
else: return 'Homem'
# 0 = mulher , 1 = homem
for index, row in data.iterrows():
print(convert_label(row['classe']), '%0.2f' % euclideanDistance([row['peso'], row['altura']],[77, 1.68], 2))
# + [markdown] deletable=true editable=true
# Dá para perceber que o valor de $k$ influencia bastante na classificação dos objetos. Mais para frente usaremos a precisão do modelo para determinar o melhor valor de $k$. Quando o valor de $k$ é muito pequeno, o modelo está mais sensível aos pontos de ruído da base. Quando o $k$ é muito grande, a vizinhança pode incluir elementos de outra classe. Vale ressaltar que, normalmente, escolhemos valores de $k$ ímpar para evitar empates.
# + [markdown] deletable=true editable=true
# Um caso especial do KNN é quando utilizamos K = 1. Vamos considerar o exemplo da imagem a seguir:
#
# ** Dataset de treinamento **
#
# 
#
# + [markdown] deletable=true editable=true
# Ao considerarmos K = 1, podemos construir um mapa de classificação, como é mostrado a seguir:
#
# ** Mapa de classificação para o KNN (K=1) **
#
# 
# > *Image Credits: Data3classes, Map1NN, Map5NN by Agor153. Licensed under CC BY-SA 3.0*
#
# Uma nova instância será classificada de acordo com a região na qual ela se encontra.
# + [markdown] deletable=true editable=true
# Para finalizar, vale ressaltar dois pontos. O primeiro é que em alguns casos faz-se necessário normalizar os valores da base de treinamento por conta da discrepância entre as escalas dos atributos. Por exemplo, podemos ter altura em um intervalo de 1.50 à 1.90, peso no intervalo de 60 à 100 e salário no intervalo de 800 à 1500. Essa diferença de escalas pode fazer com que as medidas das distâncias sejam influenciadas por um único atributo.
#
# Um outro pronto é em relação as vantagens e desvantagens desta técnica. A principal vantagem de se usar um KNN é que ele é um modelo simples de ser implementado. No entanto, ele possui um certo custo computacional no cálculo da distância entre os pontos. Um outro problema é que a qualidade da classificação pode ser severamente prejudicada com a presença de ruídos na base.
#
#
# + [markdown] deletable=true editable=true
# ### Implementando o KNN com ScikitLearn
#
# Vamos implementar o KNN utilizando o ScikitLearn e realizar as tarefas de classificação para a base da Iris.
# + deletable=true editable=true
# Importando o Dataset
from sklearn.datasets import load_iris
data_iris = load_iris()
X = data_iris.data
y = data_iris.target
# + [markdown] deletable=true editable=true
# Ao instanciar o modelo do KNN devemos passar o parâmetro *n_neighbors* que corresponde ao valor $k$, a quantidade de vizinhos próximos que será considerada.
# + deletable=true editable=true
# Importando e instanciando o modelo do KNN com k = 1
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
# + [markdown] deletable=true editable=true
# Instanciado o modelo, vamos treina-lo com os dados de treinamento.
# + deletable=true editable=true
knn.fit(X, y)
# + [markdown] deletable=true editable=true
# Assim como fizemos com a regressão linear, podemos utilizar o modelo construído para fazer a predição de dados que ainda não foram analisados.
# + deletable=true editable=true
# O método predict retorna a classe na qual a instância foi classificada
predict_value = knn.predict([[3, 5, 4, 2],[1,2,3,4]])
print(predict_value)
print(data_iris.target_names[predict_value[0]])
print(data_iris.target_names[predict_value[1]])
# + [markdown] deletable=true editable=true
# ### Avaliando e escolhendo o melhor modelo
#
# Para que possamos escolher o melhor modelo, devemos primeiro avalia-los. A avaliação do modelo de classificação é feita por meio da métrica denominada de **acurácia**. A acurácia corresponde a taxa de acerto do modelo. Um modelo que possui $90\%$ de acurácia acertou a classe em $90\%$ dos casos que foram analisados.
#
# Vale ressaltar que a escolha do melhor modelo depende de vários fatores por isso precisamos testar diferentes modelos para diferentes bases com diferentes parâmetros. Isso será melhor abordado mais a frente no nosso curso. Para simplificar, vamos trabahar com dois modelos do KNN para a base Iris. O primeiro com um K=3 e outro com K = 10.
# + deletable=true editable=true
knn_3 = KNeighborsClassifier(n_neighbors=3)
knn_3.fit(X, y)
knn_10 = KNeighborsClassifier(n_neighbors=10)
knn_10.fit(X, y)
# + deletable=true editable=true
accuracy_3 = knn_3.score(X, y)
accuracy_10 = knn_10.score(X, y)
# + deletable=true editable=true
print('Acurácia com k = 3: ', '%0.4f'% accuracy_3)
print('Acurácia com k = 10: ', '%0.4f'% accuracy_10)
# + [markdown] deletable=true editable=true
# Neste caso, o modelo com k = 10 é um modelo que acertou mais casos que o modelo com k = 3.
# + [markdown] deletable=true editable=true
# A acurácia calculada nos exemplos de Regressão Linear e KNN são chamadas de **acurácia no treinamento**. A acurácia recebe esse nome pois o modelo foi treinado e testado na mesma base. Quando treinamos e testamos nosso modelo com a mesma base, caímos no risco de construir um modelo que não seja capaz de generalizar aquele conhecimento adquirido. Normalmente, quando isso acontece, lidamos com um problema chamado de *overfit*. O correto é treinarmos nosso modelo com uma base e testar com um conjunto de dados novo para o modelo. Desta forma, aumentamos as chances de construir um modelo capaz de generalizar o "conhecimento" extraído da base de dados.
#
# O ideal é que pudéssemos treinar e testar o modelo em bases distintas. No próximo tutorial, vamos mostrar como podemos dividir a base em treino/teste e trabalhar com o que chamamos de **validação cruzada**.
#
# Até o próximo tutorial ;)
| SupervisedLearning/Tutorial01_RegressaoLinear_KNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tensorflow-gpu
# language: python
# name: tensorflow-gpu
# ---
# # Pure CNN (with cross-entropy)
import random
import numpy as np
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import Input, Model, layers
import tensorflow_addons as tfa
import matplotlib.pyplot as plt
gpu_available = tf.test.is_gpu_available()
print(gpu_available)
is_cuda_gpu_available = tf.test.is_gpu_available(cuda_only=True)
print(is_cuda_gpu_available)
# 清除暫存model
tf.keras.backend.clear_session()
# # Hyperparameters
# +
epochs = 20
batch_size = 16
margin = 1.
'''Margin for constrastive loss. (數值通過sigmoid 範圍於0~1)'''
SEED = 2022
#rng = np.random.default_rng(SEED)
#new_seed = rng.random()
'''fix random seed'''
np.random.seed(SEED)
tf.random.set_seed(SEED)
# -
# # Dataset
# +
# Load the MNIST dataset
(x_train_val, y_train_val), (x_test, y_test) = keras.datasets.mnist.load_data()
# Change the data type to a floating point format
x_train_val = x_train_val.astype("float32")
x_test = x_test.astype("float32")
# +
"""
## Define training and validation sets
"""
# Keep 50% of train_val in validation set
x_train, x_val = x_train_val[:30000], x_train_val[30000:]
y_train, y_val = y_train_val[:30000], y_train_val[30000:]
del x_train_val, y_train_val
# +
"""
## Create pairs of images
We will train the model to differentiate between digits of different classes. For
example, digit `0` needs to be differentiated from the rest of the
digits (`1` through `9`), digit `1` - from `0` and `2` through `9`, and so on.
To carry this out, we will select N random images from class A (for example,
for digit `0`) and pair them with N random images from another class B
(for example, for digit `1`). Then, we can repeat this process for all classes
of digits (until digit `9`). Once we have paired digit `0` with other digits,
we can repeat this process for the remaining classes for the rest of the digits
(from `1` until `9`).
"""
def make_pairs(x, y):
"""Creates a tuple containing image pairs with corresponding label.
Arguments:
x: List containing images, each index in this list corresponds to one image.
y: List containing labels, each label with datatype of `int`.
Returns:
Tuple containing two numpy arrays as (pairs_of_samples, labels),
where pairs_of_samples' shape is (2len(x), 2,n_features_dims) and
labels are a binary array of shape (2len(x)).
"""
num_classes = max(y) + 1
digit_indices = [np.where(y == i)[0] for i in range(num_classes)]
pairs = []
labels = []
pairs_answers = []
for idx1 in range(len(x)):
# add a matching example
x1 = x[idx1]
label1 = y[idx1]
idx2 = random.choice(digit_indices[label1])
x2 = x[idx2]
pairs += [[x1, x2]]
labels += [1]
# add a non-matching example
label2 = random.randint(0, num_classes - 1)
while label2 == label1:
label2 = random.randint(0, num_classes - 1)
idx2 = random.choice(digit_indices[label2])
x2 = x[idx2]
pairs += [[x1, x2]]
labels += [0]
pairs_answers += [[label1,label2]]
return np.array(pairs), np.array(labels).astype("float32"), np.array(pairs_answers)
# make train pairs
pairs_train, labels_train, pairs_train_answer = make_pairs(x_train, y_train)
# make validation pairs
pairs_val, labels_val, pairs_val_answer = make_pairs(x_val, y_val)
# make test pairs
pairs_test, labels_test, pairs_test_answer = make_pairs(x_test, y_test)
# +
"""
We get:
**pairs_train.shape = (60000, 2, 28, 28)**
- We have 60,000 pairs
- Each pair contains 2 images
- Each image has shape `(28, 28)`
"""
"""
Split the training pairs
"""
x_train_1 = pairs_train[:, 0] # x_train_1.shape is (60000, 28, 28)
x_train_2 = pairs_train[:, 1]
"""
Split the validation pairs
"""
x_val_1 = pairs_val[:, 0] # x_val_1.shape = (60000, 28, 28)
x_val_2 = pairs_val[:, 1]
"""
Split the test pairs
"""
x_test_1 = pairs_test[:, 0] # x_test_1.shape = (20000, 28, 28)
x_test_2 = pairs_test[:, 1]
# +
"""
## Visualize pairs and their labels
"""
def visualize(pairs, labels, to_show=6, num_col=3, predictions=None, test=False):
"""Creates a plot of pairs and labels, and prediction if it's test dataset.
Arguments:
pairs: Numpy Array, of pairs to visualize, having shape
(Number of pairs, 2, 28, 28).
to_show: Int, number of examples to visualize (default is 6)
`to_show` must be an integral multiple of `num_col`.
Otherwise it will be trimmed if it is greater than num_col,
and incremented if if it is less then num_col.
num_col: Int, number of images in one row - (default is 3)
For test and train respectively, it should not exceed 3 and 7.
predictions: Numpy Array of predictions with shape (to_show, 1) -
(default is None)
Must be passed when test=True.
test: Boolean telling whether the dataset being visualized is
train dataset or test dataset - (default False).
Returns:
None.
"""
# Define num_row
# If to_show % num_col != 0
# trim to_show,
# to trim to_show limit num_row to the point where
# to_show % num_col == 0
#
# If to_show//num_col == 0
# then it means num_col is greater then to_show
# increment to_show
# to increment to_show set num_row to 1
num_row = to_show // num_col if to_show // num_col != 0 else 1
# `to_show` must be an integral multiple of `num_col`
# we found num_row and we have num_col
# to increment or decrement to_show
# to make it integral multiple of `num_col`
# simply set it equal to num_row * num_col
to_show = num_row * num_col
# Plot the images
fig, axes = plt.subplots(num_row, num_col, figsize=(5, 5))
for i in range(to_show):
# If the number of rows is 1, the axes array is one-dimensional
if num_row == 1:
ax = axes[i % num_col]
else:
ax = axes[i // num_col, i % num_col]
ax.imshow(tf.concat([pairs[i][0], pairs[i][1]], axis=1), cmap="gray")
ax.set_axis_off()
if test:
ax.set_title("True: {} | Pred: {:.5f}".format(labels[i], predictions[i][0]))
else:
ax.set_title("Label: {}".format(labels[i]))
if test:
plt.tight_layout(rect=(0, 0, 1.9, 1.9), w_pad=0.0)
else:
plt.tight_layout(rect=(0, 0, 1.5, 1.5))
plt.show()
# +
"""
Inspect training pairs
"""
visualize(pairs_train[:-1], labels_train[:-1], to_show=4, num_col=4)
# +
"""
Inspect validation pairs
"""
visualize(pairs_val[:-1], labels_val[:-1], to_show=4, num_col=4)
# +
"""
Inspect test pairs
"""
visualize(pairs_test[:-1], labels_test[:-1], to_show=4, num_col=4)
# -
# # Model
# +
# Tensorflow tutorial cnn model for mnist
class simple_cnn():
'''Simple Kuihao's Subclass Style'''
def __init__(self, input_shape, embedding_dim=2, num_classes=10):
self.input_shape = input_shape
self.embedding_dim = embedding_dim
self.num_classes = num_classes
self.layers_EmbeddingNet = None
self.layers_ClassifierNet = None
def define_cnn_embeddings(self):
EmbeddingNet_input = layers.Input(self.input_shape)
x = layers.Conv2D(32, (3, 3), activation='relu')(EmbeddingNet_input)
x = layers.MaxPooling2D((2, 2))(x)
x = layers.Conv2D(64, (3, 3), activation='relu')(x)
x = layers.MaxPooling2D((2, 2))(x)
x = layers.Conv2D(64, (3, 3), activation='relu')(x)
x = layers.Flatten()(x)
x = layers.Dense(64, activation='relu')(x)
EmbeddingNet_output = layers.Dense(self.embedding_dim, activation=None)(x)
EmbeddingNet = keras.Model(name='EmbeddingNet_Subclass', inputs=EmbeddingNet_input, outputs=EmbeddingNet_output)
print(EmbeddingNet.summary())
self.layers_EmbeddingNet = EmbeddingNet
def define_classfier(self):
ClassificationNet_input = layers.Input(self.input_shape)
embeddnig = self.layers_EmbeddingNet(ClassificationNet_input)
x = layers.PReLU()(embeddnig)
ClassificationNet_output = layers.Dense(self.num_classes, activation='softmax')(x)
ClassificationNet = keras.Model(name='ClassificationNet_Subclass', inputs=ClassificationNet_input, outputs=ClassificationNet_output)
print(ClassificationNet.summary())
self.layers_ClassifierNet = ClassificationNet
def forward(self):
self.define_cnn_embeddings()
self.define_classfier()
# compile
self.layers_ClassifierNet.compile(loss='SparseCategoricalCrossentropy', optimizer='adam', metrics=['accuracy'])
def get_whole_model(self):
return self.layers_ClassifierNet
def get_EmbeddingNet(self):
return self.layers_EmbeddingNet
CNN_object = simple_cnn((28,28,1),2,10)
CNN_object.forward()
MyCNN = CNN_object.get_whole_model()
# -
MyCNN.fit(x_train, y_train,
validation_data=(x_val, y_val),
batch_size=batch_size,
epochs=epochs)
results = MyCNN.evaluate(x_test,y_test)
print("test loss, test acc:", results)
# +
# Testing Set Embeddnig 分布
MyCNN_EmbeddingNet = CNN_object.get_EmbeddingNet()
test_embeddnig = MyCNN_EmbeddingNet.predict(x_test)
print(test_embeddnig.shape)
CNN_fig = plt.figure(figsize=(20,10))
plt.scatter(test_embeddnig[:, 0], test_embeddnig[:, 1], s=5, c=y_test, cmap='Spectral')
CNN_fig.gca().set_aspect('equal', 'datalim')
plt.colorbar(boundaries=np.arange(11)-0.5).set_ticks(np.arange(10))
plt.xlabel('embeddnig dim 1')
plt.ylabel('embeddnig dim 2')
plt.show()
# -
| Twins_FL/Trailer/PureCNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import torch
# #!pip install pyannote.audio==1.1.1
# #!pip install pyannote.core[notebook]
# #!pip install pyannote.pipeline
# #!pip install pyannote.core
# !pip install ipywidgets
#pipeline = torch.hub.load('pyannote/pyannote-audio', 'dia_ami')
pipeline = torch.hub.load('pyannote/pyannote-audio', 'dia')
# %time diarization = pipeline({'audio': '/home/reves/test-dt.wav'})
for turn, _, speaker in diarization.itertracks(yield_label=True):
print(f'Speaker "{speaker}" speaks between t={turn.start:.1f}s and t={turn.end:.1f}s.')
from ipywidgets import Audio
# + tags=[]
Audio(value=open('/home/reves/test-dt.wav','rb').read(),format="wav")
# test = Audio.from_file('/home/reves/test-dt.wav', autoplay=False)
# test.play()
# -
from plume.utils.transcribe import chunk_transcribe_meta_gen, transcribe_rpyc_gen
base_transcriber, base_prep = transcribe_rpyc_gen()
transcriber, prep = chunk_transcribe_meta_gen(
base_transcriber, base_prep, method="chunked")
import pydub
audio_file = '/home/reves/test-dt.wav'
aseg = pydub.AudioSegment.from_file(audio_file)
aseg
for turn, _, speaker in diarization.itertracks(yield_label=True):
#print(f'Speaker "{speaker}" speaks between t={turn.start:.1f}s and t={turn.end:.1f}s.')
speaker_label = "Agent" if speaker == "B" else "Customer"
tscirpt = transcriber(prep(aseg[turn.start*1000:turn.end*1000]))
print(f'#{speaker_label}[{turn.start:.1f}s-{turn.end:.1f}s]: {tscirpt}')
| notebooks/Diarization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# **Função de ativação e sua derivada**
# +
def sigmoid(x):
return 1 / (1 + np.exp(-x))
def sigmoid_derivative(x):
return x * (1 - x)
# -
# **Função de custo**
# +
#def MSE(Y_target, Y_pred):
# return np.mean( (Y_target - Y_pred) ** 2 )
# -
# **Definindo o dataset**
X = np.array([
[0, 0],
[0, 1],
[1, 0],
[1, 1]
])
X
Y = np.array([
[0],
[1],
[1],
[0]
])
Y
# **Taxa de aprendizagem**
N = 0.5
# **Quantidade de épocas**
#10000
EPOCHS = 6
# **Vetor da função de custo**
cost = np.array([])
# **Arquitetura da rede**
n_neurons_input_layer = 2
n_neurons_hidden_layer_1 = 3
#n_neurons_hidden_layer_2 = 3
n_neurons_output_layer = 1
# **Pesos**
w_hidden_layer_1 = np.random.rand(n_neurons_input_layer, n_neurons_hidden_layer_1)
w_hidden_layer_1
# +
#w_hidden_layer_2 = np.random.rand(n_neurons_hidden_layer_1, n_neurons_hidden_layer_2)
#w_hidden_layer_2
# -
w_output_layer = np.random.rand(n_neurons_hidden_layer_1, n_neurons_output_layer)
w_output_layer
# **Vieses**
b_hidden_layer_1 = np.zeros(n_neurons_hidden_layer_1)
b_hidden_layer_1
# +
#b_hidden_layer_2 = np.zeros(n_neurons_hidden_layer_2)
#b_hidden_layer_2
# -
b_output_layer = np.zeros(n_neurons_output_layer)
b_output_layer
# **Treino da rede**
def MSE(Y_target, Y_pred):
return np.mean( (Y_target - Y_pred) ** 2 )
w_hidden_layer_1_ant = w_hidden_layer_1
w_output_layer_ant = w_output_layer
aux_1
aux_o
for epoch in range(EPOCHS):
activation_hidden_layer_1 = sigmoid( np.dot(X, w_hidden_layer_1) + b_hidden_layer_1 )
activation_output_layer = sigmoid(np.dot(activation_hidden_layer_1,w_hidden_layer_1) + b_output_layer)
cost = np.append(cost, MSE(Y, activation_output_layer))
delta_output_layer = (Y - activation_output_layer) * sigmoid_derivative(activation_output_layer)
delta_hidden_layer_1 = np.dot(delta_output_layer, w_output_layer.T) * sigmoid_derivative(activation_hidden_layer_1)
aux_1 = w_hidden_layer_1
aux_o = w_output_layer
w_output_layer += N * np.dot(activation_hidden_layer_1.T, delta_output_layer)
w_hidden_layer_1 += N * np.dot(X.T, delta_hidden_layer_1)
w_hidden_layer_1_ant = aux_1
w_output_layer_ant = aux_o
print('layer 1\n{}'.format())
print('layer o\n{}'.format())
print('ant layer 1\n{}'.format())
print('ant layer 1\n{}'.format())
# **Gráfico da função de custo**
plt.plot(cost)
plt.grid(color = 'g', linestyle=':', linewidth=.1)
plt.title('Função de custo da rede')
plt.xlabel('Épocas')
plt.ylabel('Custo')
plt.show()
print('oi')
| 1mexer opiaperceptron_multicamadas-Copy1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="A-HpugGZklca"
# ## Baixando séries históricas do Yahoo! Finance
#
# **Topicos:**
#
# * Como obter uma lista de ações
# * Como baixar séries históricas de uma lista de ações
# * Como salvar os DataFrames do Pandas no formato pickle<br><br>
#
# **Considerações**
#
# A ferramenta usada para baixar é o fix_yahoo_finance que é uma modificação feita a partir do pandas_datareader<br><br>
#
# **1. Obtendo a lista de ações** (opcional)
#
# O objetivo desta etapa é apenas obter uma lista de tickers.<br><br>
#
# **2. Lista de ações da Composição Atual do IBOV**
#
# Esta etapa obtém a lista de ações que compõem o índice Bovespa atual (2021).
#
# Será usada uma sequência de comandos bash para extrair a lista da página Composição Atual do IBOV - Índice Bovespa, que serão executados a dentro de uma rotina Python, que posteriormente irá salvar a lista em disco.<br><br>
#
# **3. Webscrapping: baixando a lista da página**
#
# Poderia ser feito usando BeautifulSoup e/ou Scrapy?
#
# Sim, no entanto para este caso utilizaremos comandos bash.<br><br>
# + id="-Tq0ZKOvkXrJ"
import subprocess
# + id="BdFuGKMKmiWp"
# bash command line to be exectuted inside python
commands = """
# Baixando o código html da página
wget https://br.advfn.com/indice/ibovespa -O tmp0.tmp
# Extraindo as colunas de tickers e nomes
cat tmp0.tmp | head -n434 | tail -n80 > tmp1.tmp
cat tmp1.tmp | grep 'br.advfn.com' | cut -c1-200 | cut -d. -f3- | cut -d'"' -f1,3 > tmp2.tmp
cat tmp2.tmp | cut -d'/' -f4-6 | sed -e 's./cotacao"Cotação .,.g' | cut -d',' -f1 | rev | cut -d'-' -f1 | rev > tmp4.tmp
cat tmp2.tmp | cut -d'/' -f4-6 | sed -e 's./cotacao"Cotação .,.g' | cut -d',' -f2 > tmp5.tmp
# Salvando a lista final
paste -d, tmp4.tmp tmp5.tmp > lista_ibovespa.csv
# Removendo arquivos temporários
rm -f tmp*.tmp
"""
# + id="p-aZRlF7mlEk"
p = subprocess.Popen(commands, shell=True, stdout=subprocess.PIPE)
msg, err = p.communicate()
# + [markdown] id="df1EqxePm7i9"
# **Modificações adicionais**
#
# Carregando a lista anterior como numpy.array:
# + colab={"base_uri": "https://localhost:8080/"} id="dBUg9Nycm6uN" outputId="6be0e72e-f166-4f68-dda7-bec344370e7f"
import numpy as np
# ibovespa stock tickers
lst_stocks = np.loadtxt('./lista_ibovespa.csv', delimiter=',', dtype=str)
print('Number of stocks listed on iBovespa:', len(lst_stocks))
# + colab={"base_uri": "https://localhost:8080/"} id="JxGtbbO5nMoM" outputId="27944f1e-bc23-4db0-f525-b7034832a3ed"
for ticker, name in lst_stocks[:41]:
print('Ticker: {} | Stock name: {}'.format(ticker, name))
# + [markdown] id="EmDFiEmOntrc"
# O Yahoo! Finance emprega um sufixo para ações de bolsas fora dos EUA. Para as ações da Bovespa, por exemplo, aplica o sufixo **.SA** no símbolo de cada ação. Ou seja, a ação ABEV3 da Ambev é referenciada como 'ABEV3**.SA**'.
#
# **Referências:**
#
# [Exchanges and data providers on Yahoo Finance](https://help.yahoo.com/kb/SLN2310.html)
#
# [Yahoo Finance Exchanges And Suffixes](https://sites.google.com/a/stockhistoricaldata.com/stock-historical-data/yahoo-finance-suffixes)
# + [markdown] id="mYKQtJWun_Rn"
# **Adicionando o sufixo nos simbolos:**
# + colab={"base_uri": "https://localhost:8080/"} id="4B_s8bYnoD3U" outputId="6114e0d9-bce6-4e02-f1b1-0ef329f84ac8"
# ticker symbols with Bovespa's suffix
lst_tickers = np.asarray([ '{}.SA'.format(x) for x in lst_stocks[:,0]], dtype=str)
#
for ticker in lst_tickers[1:41]:
print('Ticker: {}'.format(ticker))
# + [markdown] id="8PaNBbYCoDOT"
# **Incorporando BVMF3, Ibovespa e Dólar**<br><br>
#
# * Até 2017 a ação B3 ON tinha o símbolo BVMF3 e em 2018 passou a usar o símbolo B3SA3. Assim a BVMF3.SA será adicionada manualmente à lista de ações a serem baixadas.
#
# * O índice Bovespa (^BVSP) e a cotação do Dólar em reais (USDBRL=X) também serão adicionadas. (Perceba o prefixo '^' e o sufixo '=X' usados.)
# + colab={"base_uri": "https://localhost:8080/"} id="o7ejYObEpGvt" outputId="a6e7e818-8fa5-401a-a2d7-034436e308da"
# adding BVMF3.SA
lst_tickers = np.sort(np.concatenate((lst_tickers, ['BVMF3.SA']))) # this stock changed the name to B3SA3 in 2018
# adding ^BVSP and USDBRL=X
lst_tickers = np.concatenate((lst_tickers, ['^BVSP', 'USDBRL=X'])) # this stock changed the name to B3SA3 in 2018
# checking the last ones
for ticker in lst_tickers[-2:]:
print('Ticker: {}'.format(ticker))
# saving the list
np.savetxt('list_tickers_yahoo.txt', lst_tickers, fmt='%s')
# + [markdown] id="Mt7Qfbf9qzAt"
# ## Baixando as séries históricas
#
# O API do Yahoo! Finance não funciona mais como antes, causando falhas no uso da biblioteca pandas_datareader.<br><br>
#
# O recente mal funcionamento com algumas APIs é descrito na página de desenvolvimento do pandas_datareader:<br><br>
#
#
# **Yahoo!, Google Options, Google Quotes and EDGAR have been immediately deprecated.**
#
# > Immediate deprecation of Yahoo!, Google Options and Quotes and EDGAR. The end points behind these APIs have radically changed and the existing readers require complete rewrites. In the case of most Yahoo! data the endpoints have been removed. PDR would like to restore these features, and pull requests are welcome.<br><br>
#
# **Existe porém uma solução temporária para isto, o [fix-yahoo-finance](https://github.com/ranaroussi/fix-yahoo-finance).**<br><br>
#
# O fix_yahoo_finance não está disponível na distribuição Anaconda, mas é possível o instalar a partir do pip:
#
# `$ pip install fix_yahoo_finance --upgrade --no-cache-dir`<br><br>
#
# **Usando o fix_yahoo_finance**
#
# Abaixo é definida uma função que utiliza o módulo fix_yahoo_finance para baixar séries históricas do API do Yahoo! Finance.<br><br>
#
# A função método download_stocks_from_yahoo recebe a lista de símbolos, baixa cada elemento da lista como DataFrame do Pandas e os salva no formato pickle na pasta indicada pela variável output_path. O nome do arquivo salvo para cada ação da lista é df_XXXXX.pickle onde XXXXX representa o símbolo da ação em questão, onde os prefixos e sufixos são removidos.
# + id="KxLAANf7sapU"
import numpy as np
import os
import subprocess
#from pandas_datareader import data as pdr
import fix_yahoo_finance as yf
# See https://github.com/ranaroussi/fix-yahoo-finance/blob/master/README.rst
yf.pdr_override() # <== that's all it takes :-)
def download_stocks_from_yahoo(tickers, start, end, output_path='', verbose=1):
'''
Downloads stocks from Yahoo! Finance and saves each stock as a Pandas DataFrame object
in the pickle data format: df_XXXXX.pickle, where XXXXX is the ticker of a particular stock.
Prefixes and suffixes are removed from the output name.
Inputs:
tickers: list/array of tickers
start/end: datetime.datetime.date objects
output_path: string
Outputs:
failed: list of the tickers whose download failed
'''
failed = []
# creates the output folder path if it doesnt exist yet
command = 'mkdir -p {}'.format(output_path)
p = subprocess.Popen(command, shell=True, stdout=subprocess.PIPE)
msg, err = p.communicate()
for ticker in tickers:
ticker = ticker.upper()
# deleting Yahoo's prefixes and suffixes from the name
stock_name = ticker.replace('^', '')
stock_name = stock_name.split('=')[0]
stock_name = stock_name.replace('.SA', '')
# setting the full path for the output file
fname_output = os.path.join(output_path,'df_{}.pickle'.format(stock_name))
try:
if verbose:
print('\n Attempting to download {} from {} to {}.'.format(ticker, start, end))
df = yf.download(ticker, start=start, end=end, as_panel=False)
except:
failed.append(ticker)
print('* Unable to download {}. * \n'.format(ticker))
else:
try:
df.to_pickle(fname_output)
except:
print('* Error when trying to save on disk {}. * \n'.format(fname_output))
return failed
# + [markdown] id="a9hQcj4fs7Wi"
# **Download das ações**
#
# Serão baixadas as séries históricas das ações do período de 01/01/2001 até a data presente. Os DataFrames serao salvos no formato pickle no diretório 'raw'.
# + id="G0k0Gvgrs4bY"
import numpy as np
import datetime
# loading the list of tickers as a np.array
tickers = np.loadtxt('list_tickers_yahoo.txt', dtype=str)
# setting the start and end dates
start = datetime.datetime(2001, 1, 1).date()
end = datetime.datetime.today().date()
# setting folder name where dataframes will be saved
output_path = 'raw'
# + colab={"base_uri": "https://localhost:8080/"} id="ZZP0JzfbtXXp" outputId="59ed46b3-bdf0-4a71-9489-88b04d02f729"
# downloading list of tickers
lst_failed = download_stocks_from_yahoo(tickers[:], start, end, output_path)
# + colab={"base_uri": "https://localhost:8080/"} id="Gd9eXHx-tg5m" outputId="aae62633-3167-4e89-9f95-8104eef53088"
# Checking for errors
if len(lst_failed) > 0:
print('Unable to download the following stocks:')
print(lst_failed)
#print('\n Trying one more time:')
#lst_failed = download_stocks_from_yahoo(lst_failed, start, end, output_path)
else:
print('All tickers downloaded successfully')
# + [markdown] id="_H317MqRuFmp"
# **Concatenação da BVMF3 e B3SA3 (opcional)**
#
# Como comentado anteriormente, esta ação mudou de nome em 2018. Neste passo, os DataFrames correspondentes a estas ações serão concatenados em um novo que será salvo em disco.
# + colab={"base_uri": "https://localhost:8080/", "height": 252} id="1Z_cjl4VuDDa" outputId="7fc6c2bd-e68b-43a6-a45f-2e27d9baca70"
import pandas as pd
import os
picklepath = os.path.join(output_path, 'df_{}.pickle')
#df1 = pd.read_pickle( picklepath.format('BVMF3') )
df2 = pd.read_pickle( picklepath.format('B3SA3') )
#
#print(df1.shape, df2.shape)
print(df2.shape)
df2.tail()
# + colab={"base_uri": "https://localhost:8080/"} id="raHZlyufusmH" outputId="94dfd3f5-964e-4f74-c7f4-febb6c500342"
#df3 = pd.concat([df1, df2], axis=0)
#print(df1.shape, df2.shape, df3.shape)
#print(df3.columns)
print(df2.columns)
# + id="xnMpN-FJuSRz"
#df3.tail() # there are few days missing
df2.tail() # there are few days missing
# re-writing on disk
#df3.to_pickle(picklepath.format('B3SA3'))
df2.to_pickle(picklepath.format('B3SA3'))
# deleting from disk
#status = os.system('rm -f {}'.format(picklepath.format('BVMF3')))
# + [markdown] id="674YARVbw_Lr"
# ## Carrengando os dados (Loading the data)
#
# Os dados serão armazenados em dois dataframes:
#
# * **df_stocks**: all the stocks
# * **df_bench**: only the benchmarks
# + [markdown] id="z8WEdjksxbqe"
# 1. Importando Bibliotecas
# + id="tI2Wre-Fw2mO"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import datetime
import dateutil
import glob
import os
# + [markdown] id="8ZweHXqew_jZ"
# 2. Listando dataframes previamente armazenados
# + id="v8AHAE44w_5x"
# listing pandas dataframes previously saved
lst_df_path = glob.glob(os.path.join('/content/raw', 'df_*.pickle'))
# + id="pYQLIq5gxAjh" colab={"base_uri": "https://localhost:8080/"} outputId="6b03a668-c050-4a04-ee5c-58aee79c67e7"
# checking the path and file names
#lst_df_path[:3]
lst_df_path[:]
# + id="9vExDHzeyBEm"
# remove the ticker that will be used for Benchmarks later
lst_df_path.remove('/content/raw/df_BVSP.pickle')
lst_df_path.remove('/content/raw/df_USDBRL.pickle')
# + id="bgxcdpF9ySIf"
# creating a separed list for the Benchmarks
lst_df_path_bench = ['/content/raw/df_BVSP.pickle', '/content/raw/df_USDBRL.pickle']
# + colab={"base_uri": "https://localhost:8080/"} id="F8La_mRevgwu" outputId="9768bd65-a13f-42b9-fd14-93508316f80d"
lst_df_path_bench[:]
# + id="zBzcS1IzyUlR"
# concatenating all stocks into one dataframe
lst_df_stocks = []
for fname in lst_df_path:
df = pd.read_pickle(fname)
# keeping only Adj Close
df.drop(columns=['Open', 'High', 'Low', 'Close', 'Volume'], inplace=True)
ticker = fname.split('/content/raw/')[1].split('df_')[1].split('.')[0]
df.columns = [ticker]
lst_df_stocks.append(df)
df_stocks = pd.concat(lst_df_stocks, axis=1)
# + id="Byq4JsIYyWzt"
df_stocks = pd.concat(lst_df_stocks, axis=1)
# + id="ie2cw3C7yZGn" colab={"base_uri": "https://localhost:8080/"} outputId="399de68b-5a89-4e0e-d652-9e0e9a7ebfd1"
# checking column names
df_stocks.columns
# + id="lD-6M7UiydhW"
# concatenating the benchmarks into one dataframe
lst_df_bench = []
for fname in lst_df_path_bench:
df = pd.read_pickle(fname)
# keeping only Adj Close
df.drop(columns=['Open', 'High', 'Low', 'Close', 'Volume'], inplace=True)
ticker = fname.split('/content/raw/')[1].split('df_')[1].split('.')[0]
df.columns = [ticker]
lst_df_bench.append(df)
df_bench = pd.concat(lst_df_bench, axis=1)
# + id="SHBGJ297ygV9" colab={"base_uri": "https://localhost:8080/"} outputId="c961b3b3-7f9a-44f1-e808-de195429e16b"
df_bench.columns
# + id="oiUquaeKyhV_" colab={"base_uri": "https://localhost:8080/", "height": 235} outputId="c6a7dd80-8c12-4839-86a0-472398e6e132"
df_bench.head()
# + [markdown] id="zw0iLfokxBA_"
# ## Portfólio Otimizado Mensal
#
# O objetivo é compor uma carteira com bom desempenho utilizando apenas uma pequena quantidade de ações da lista.
#
# A cada mês será elaborada uma nova carteira com base no Índice Sharpe dos meses anteriores, e seu desempenho será comparado com três benchmarks:
#
# * iBovespa: Índice oficial da Bovespa (composto por +60 ações)
#
# * Média BVSP: média simples de todas as ações disponíveis da iBovespa
#
# * Dolar: O valor atual dos dólares americanos em reais
#
# **Restrições adicionais ao portfólio:**
#
# O peso máximo de uma ação é de 25%
# O peso mínimo de uma ação é 2%
#
# **Resultados esperados:**
#
# * desempenho aprimorado no longo prazo
# * maior volatilidade que o iBovespa, devido ao pequeno número de ações que compõem a carteira
#
# **Configurando a otimização**
#
# *Baseado no curso Udemy de Jose Portilla em [Python para algoritmo financeiro e comercial.](https://www.udemy.com/python-for-finance-and-trading-algorithms/learn/v4/)*
# + id="yRUArlKVxBUr"
from scipy.optimize import minimize
# + id="oPzVVzYAobk9"
# utility function to obtain the expected Return, expected Volatity, and Sharpe Ration from the log returns, given the weights
def get_ret_vol_sr(weights):
global log_ret
weights = np.array(weights)
ret = np.sum( log_ret.mean() * weights * 252)
vol = np.sqrt( np.dot(weights.T, np.dot(log_ret.cov()*252, weights)))
sr = ret/vol
return np.array([ret, vol, sr])
# + id="P5lY2a6Cocnh"
# the actual function to be minimized
def neg_sharpe(weights):
return -1.*get_ret_vol_sr(weights)[2]
# + id="ByzTjpbEogum"
# contraint function
def check_sum(weights):
return np.sum(weights) - 1.
# + id="wOVOdhSLoipS"
# contraint function
def check_max_weight(weights):
global max_weight
return np.minimum(weights.max(), max_weight) - weights.max()
# + id="0VVWYsXZokVh"
# contraint function
def check_weights(weights):
global max_weight
w1 = np.sum(weights) - 1.
w2 = np.minimum(weights.max(), max_weight) - weights.max()
return np.abs(w1) + np.abs(w2)
# + id="YkBr7YIWomJc"
# constraint tuple
#cons = ({'type' : 'eq', 'fun' : check_sum})
#cons = ({'type' : 'eq', 'fun' : check_sum}, {'type' : 'eq', 'fun' : check_max_weight}) # did not work
cons = ({'type' : 'eq', 'fun' : check_weights}) # using this workaround instead
# + id="9rp2cX83onma"
n_stocks = df_stocks.shape[1]
# + id="qscX04Z8ooB6"
bounds = tuple([(0,1) for i in range(n_stocks)])
# + id="5eJXj1B4opwi"
init_guess = np.ones(n_stocks) / n_stocks
# + [markdown] id="-Ep3QM3kow23"
# ## Definir parâmetros de previsão
# + id="x8WGdb1zorgr"
# the start date of the fist prediction (year, month, day)
day_start = datetime.datetime(2020,1,1).date()
# total number of months to run the prediction
n_months_run = 16
# training months before current prediction
n_months_train = 12
# portfolio weights (before re-balancing)
max_weight = 0.25 # used in the constraint function
min_weight = 0.02 # used in the running prediction
# + [markdown] id="-b6uPwH-o1qg"
# # Previsão mensal em execução
# + id="OJ0ZL3zno53L" colab={"base_uri": "https://localhost:8080/"} outputId="53943a72-37e9-4fa7-ec33-b7798c183415"
delta_month = dateutil.relativedelta.relativedelta(months=+1)
delta_day = dateutil.relativedelta.relativedelta(days=+1)
valid_start = day_start
valid_end = valid_start + delta_month - delta_day
train_start = valid_start - n_months_train*delta_month
train_end = valid_start - delta_day
time = []
p = []
b1 = []
b2 = []
b3 = []
#
for i in range(n_months_run):
# dataframes
df_train = df_stocks.truncate(before=train_start, after=train_end)
df_valid = df_stocks.truncate(before=valid_start, after=valid_end)
df_valid_bench = df_bench.truncate(before=valid_start, after=valid_end)
# calculating log returns of the training data
log_ret = np.log( df_train.divide(df_train.shift(1, axis=0), axis=0) ).iloc[2:]
# notice that log_ret is used by the function `get_ret_vol_sr` and, consequently,
# the `neg_sharpe` function
# calculating optimized weights
opt_results = minimize(neg_sharpe, init_guess, method='SLSQP', bounds=bounds, constraints=cons)
weights = opt_results.x
# Weight Re-balancing
idx = np.where(opt_results.x>=min_weight)[0]
weights = weights[idx]
weights /= weights.sum()
labels = log_ret.columns[idx]
# using the portfolio weights on the validation data
df1 = df_valid[labels]
df1 = df1/df1.iloc[0] # percentage return of the portfolio
df2 = (df1 * weights).sum(axis=1)
df2 = df2/df2.iloc[0] # percentage return of the portfolio
# percentage return of the benchmarks
df2b = df_valid_bench/df_valid_bench.iloc[0]
time.append(valid_start.strftime('%Y/%m'))
p.append(df2.iloc[-1])
b1.append(df2b['BVSP'].iloc[-1])
b2.append(df2b['USDBRL'].iloc[-1])
b3.append(df1.mean(axis=1).iloc[-1]) # Simple average of all stocks
print('\nStart: {}, Portfolio: {:.2f}, iBovespa: {:.2f}, Dolar: {:.2f}, Avg. : {:.2f}'.format(time[-1], p[-1],
b1[-1], b2[-1], b3[-1]))
for l,w in zip(labels, weights):
print(' > {} : {:.2f}'.format(l, w))
# time update for the next loop
valid_start += delta_month
valid_end = valid_start + delta_month - delta_day
train_start += delta_month
train_end = valid_start - delta_day
# + [markdown] id="xPvjA6IVpgPb"
# ## Apresentando os resultados
# + id="wT2hJkJRqQNx"
d = {'Date' : pd.to_datetime(time),
'Portfolio' : p,
'iBovespa' : b1,
'Dolar' : b2,
'Avg. BVSP' : b3}
df_results = pd.DataFrame(data=d)
df_results.set_index('Date', inplace=True)
# + id="DDBvahyGqQ4Y" colab={"base_uri": "https://localhost:8080/"} outputId="52553663-f4cf-4ef6-86f1-3a3310208dce"
print('Average - Monthly returns:')
df_results.mean(axis=0)
# + id="mKnAdxLdqS8Q" colab={"base_uri": "https://localhost:8080/"} outputId="87d7e428-9393-4ad9-c82e-4a5620d71322"
print('std - Monthly returns:')
df_results.std(axis=0)
# + id="L35WvQXyqYCI" colab={"base_uri": "https://localhost:8080/", "height": 308} outputId="aac34117-5977-4dd9-f753-5decab007edd"
ax = df_results.plot(style='-o')
ax.axhline(y=1.0, color='gray', linestyle='--', lw=0.5)
| bovespa_tickers_download_portfolio.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# # German Traffic Sign Classification
#
# ## Step 0: Load The Data
# +
# Load pickled data
import pickle
# TODO: Fill this in based on where you saved the training and testing data
training_file = "traffic-signs-data/train.p"
validation_file="traffic-signs-data/valid.p"
testing_file = "traffic-signs-data/test.p"
with open(training_file, mode='rb') as f:
train = pickle.load(f)
with open(validation_file, mode='rb') as f:
valid = pickle.load(f)
with open(testing_file, mode='rb') as f:
test = pickle.load(f)
X_train, y_train = train['features'], train['labels']
X_valid, y_valid = valid['features'], valid['labels']
X_test, y_test = test['features'], test['labels']
# -
# ---
#
# ## Step 1: Dataset Summary & Exploration
# +
### Replace each question mark with the appropriate value.
### Use python, pandas or numpy methods rather than hard coding the results
import numpy as np
# TODO: Number of training examples
n_train = len(X_train)
# TODO: Number of validation examples
n_validation = len(X_valid)
# TODO: Number of testing examples.
n_test = len(X_test)
# TODO: What's the shape of an traffic sign image?
image_shape = X_train[0].shape
# TODO: How many unique classes/labels there are in the dataset.
n_classes = len(np.unique(y_train))
print("Number of training examples =", n_train)
print("Number of testing examples =", n_test)
print("Image data shape =", image_shape)
print("Number of classes =", n_classes)
# -
# ### Exploratory visualizations of the dataset
# +
### Data exploration visualization code goes here.
### Feel free to use as many code cells as needed.
import random
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
# %matplotlib inline
#Random image
index = random.randint(0, len(X_train))
image= X_train[index].squeeze()
plt.imshow(image, cmap="gray")
print(image.shape)
print(y_train[index])
# +
#plot one image from each of the class
#The index of the image can be mapped to the name of the sign using the
#csv file included in the repository
u, indices = np.unique(y_train, return_index=True)
fig = plt.figure(figsize=(15, 15))
fig.suptitle("All Traffic Signs")
columns = 8
rows = (len(indices)/columns) + 1
for i, index in enumerate(indices, 1):
fig.add_subplot(rows, columns, i)
plt.imshow(X_train[index].squeeze())
plt.xlabel(y_train[index])
plt.tick_params(axis='both',
which='both',
bottom='off',
top='off',
labelbottom='off',
right='off',
left='off',
labelleft='off')
plt.show()
# -
#plot number of unique samples per class
histogram = plt.figure()
hist, bins = np.histogram(y_train, bins=u)
width = 0.7 * (bins[1] - bins[0])
center = (bins[:-1] + bins[1:]) / 2
plt.bar(center, hist, align='center', width=width)
plt.title("Number of samples per class")
plt.show()
# ----
#
# ## Step 2: Design and Test a Model Architecture
#
# Design and implement a deep learning model that learns to recognize traffic signs. Train and test your model on the [German Traffic Sign Dataset](http://benchmark.ini.rub.de/?section=gtsrb&subsection=dataset).
#
# The LeNet-5 implementation shown in the [classroom](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) at the end of the CNN lesson is a solid starting point. You'll have to change the number of classes and possibly the preprocessing, but aside from that it's plug and play!
#
# With the LeNet-5 solution from the lecture, you should expect a validation set accuracy of about 0.89. To meet specifications, the validation set accuracy will need to be at least 0.93. It is possible to get an even higher accuracy, but 0.93 is the minimum for a successful project submission.
#
# There are various aspects to consider when thinking about this problem:
#
# - Neural network architecture (is the network over or underfitting?)
# - Play around preprocessing techniques (normalization, rgb to grayscale, etc)
# - Number of examples per label (some have more than others).
# - Generate fake data.
#
# Here is an example of a [published baseline model on this problem](http://yann.lecun.com/exdb/publis/pdf/sermanet-ijcnn-11.pdf). It's not required to be familiar with the approach used in the paper but, it's good practice to try to read papers like these.
# ### Pre-process the Data Set (normalization, grayscale, etc.)
# Minimally, the image data should be normalized so that the data has mean zero and equal variance. For image data, `(pixel - 128)/ 128` is a quick way to approximately normalize the data and can be used in this project.
#
# Other pre-processing steps are optional. You can try different techniques to see if it improves performance.
#
# Use the code cell (or multiple code cells, if necessary) to implement the first step of your project.
# +
#helper functions
#convert images to singlechannel:
def single_channel(images, mode):
#convert either to gray or y channel images
c1_images = np.empty([len(images), 32, 32, 1])
for i in range(len(images)):
if mode == 'Y':
c1, c2, c3 = cv2.split(cv2.cvtColor(images[i], cv2.COLOR_RGB2YUV))
elif mode == 'G':
c1 = cv2.cvtColor(images[i], cv2.COLOR_RGB2GRAY)
c1_images[i] = np.expand_dims(c1, axis=3)
return c1_images
#add noise to the images
def add_jitter(images):
jitter_images = np.empty(images.shape)
for i in range(len(images)):
img = images[i]
h, w, c = img.shape
noise = np.random.randint(-2, 2, (h, w))
jitter = np.zeros_like(img)
jitter[:, :, 0] = noise
noise_added = np.expand_dims(cv2.add(img, jitter), axis=3)
jitter_images[i] = noise_added
return jitter_images
#rotate the images
def rotate(images):
#rotate the image between a random angle of [-15, 15] deg
rotated_images = np.empty(images.shape)
for i in range(len(images)):
(h, w) = images[i].shape[:2]
center = (w / 2, h / 2)
rand_angle = random.uniform(-15.0, 15.0)
M = cv2.getRotationMatrix2D(center, rand_angle, 1.0)
rotated_image = np.expand_dims(cv2.warpAffine(images[i], M, (w, h)), axis=3)
#print(rotated_image.shape)
rotated_images[i] = rotated_image
return rotated_images
#concatenate the images together
def concatenate_images(c1_images, jitter_images, rotated_images, labels):
X_train_final = np.empty([len(c1_images)*3, 32, 32, 1])
for i in range(len(X_train)):
X_train_final[i] = c1_images[i]
X_train_final[i + len(c1_images)] = jitter_images[i]
X_train_final[i + len(c1_images)*2] = rotated_images[i]
#concatenate the labels together
print(labels.shape)
labels_length = len(labels)
y_train_final = np.empty([labels_length*3],)
for i in range(labels_length):
y_train_final[i] = labels[i]
y_train_final[i + labels_length] = labels[i]
y_train_final[i + labels_length*2] = labels[i]
return X_train_final, y_train_final
# +
import cv2
from skimage import exposure
#experimental pipeline tried to augment data: not used in final result.
def pipeline(images, labels, mode):
"""
Preprocess the image by passing it through the pipeline
:param images: The initial images to be processed
:param labels: The respective labels of the images
:param mode: Y = Y channel from the YUV spectrum | G = Grayscale
:param augment: 1 = augment the data | 0 = don't augment the data
:return: Preprocessed and concatenated images, concatednated labels
"""
c1_images = single_channel(images, mode)
print(c1_images.shape)
#add noise to the image
jitter_images = add_jitter(c1_images)
#rotate the images
rotated_images = rotate(c1_images)
print(rotated_images.shape)
#concatenate the images
X_train_final, y_train_final = concatenate_images(c1_images,
jitter_images,
rotated_images,
labels)
return X_train_final, y_train_final
### Normalise the image data
def normalize(image_data):
"""
Normalize the image data by equalizing histogram
:param image_data: The image data to be normalized
:return: Normalized image data
"""
normalized_data = []
for i in range(len(image_data)):
normalized_data.append(exposure.equalize_hist(image_data[i]))
return normalized_data
# -
X_train_final= single_channel(X_train, 'Y')
X_valid_final = single_channel(X_valid, 'Y')
X_test_final = single_channel(X_test, 'Y')
from keras.preprocessing.image import ImageDataGenerator
#augmenting data using keras
def augment_data(x_train, y_train):
datagen = ImageDataGenerator(
featurewise_center=False,
featurewise_std_normalization=False,
rotation_range=15,
width_shift_range=0.2,
height_shift_range=0.2,
zoom_range=0.2,
shear_range=0.2,
horizontal_flip=False,
vertical_flip=False)
augmented_images = []
augmented_labels = []
datagen.fit(x_train)
batch = 0
for x_batch, y_batch in datagen.flow(x_train, y_train, batch_size=len(x_train)):
augmented_images.append(x_batch)
augmented_labels.append(y_batch)
batch += 1
if batch == 2:
break
return augmented_images, augmented_labels
augmented_images, augmented_labels = augment_data(X_train_final, y_train)
#concatenate all the augmented data and the initial training data
def concatenate(train_images, augmented_images, train_labels, augmented_labels):
org_len = len(train_labels)
final_images = np.empty([org_len * (len(augmented_images) + 1), 32, 32, 1])
final_labels = np.empty([org_len * (len(augmented_images) + 1)])
for i in range(org_len):
final_images[i] = train_images[i]
final_labels[i] = train_labels[i]
for i in range(len(augmented_images)):
for j in range(org_len):
final_images[j + org_len * (i + 1)] = augmented_images[i][j]
final_labels[j + org_len * (i + 1)] = augmented_labels[i][j]
return final_images, final_labels
X_train_final, y_train_final = concatenate(X_train_final, augmented_images, y_train, augmented_labels)
#normalise the training, validation and test data using equalize histogram
X_train_final = normalize(X_train_final)
X_valid_final = normalize(X_valid_final)
X_test_final = normalize(X_test_final)
# +
import scipy
import scipy.misc
from PIL import Image
#tried out global contrast normalization. Accuracy was lower than histogram eq.
def global_contrast_normalization(images, s, lmda, epsilon):
normalized_images = []
for i in range(len(images)):
# replacement for the loop
X_average = np.mean(images[i])
# print('Mean: ', X_average)
X = images[i] - X_average
# `su` is here the mean, instead of the sum
contrast = np.sqrt(lmda + np.mean(X**2))
X = s * X / max(contrast, epsilon)
normalized_images.append(X)
return normalized_images
# scipy can handle it
#X_train_final = global_contrast_normalization(X_train_final, 1, 10, 0.000000001)
#X_valid_final = global_contrast_normalization(X_valid_final, 1, 10, 0.000000001)
#X_test_final = global_contrast_normalization(X_test_final, 1, 10, 0.000000001)
# +
from sklearn.utils import shuffle
X_train_final, y_train_final = shuffle(X_train_final,y_train_final)
# -
# ### Model Architecture
import tensorflow as tf
#reduced the number of epochs from 30 to 20 as the validation accuracy peaks around 20
#definitely overfitting with 30 epochs as test accuracy is less than validation accuracy
EPOCHS = 40
BATCH_SIZE = 256
#increased the number of features
def get_weights_biases(mu, sigma):
weights = {
'wc1' : tf.Variable(tf.truncated_normal([5, 5, 1, 108], mu, sigma)),
'wc2' : tf.Variable(tf.truncated_normal([5, 5, 108, 108], mu, sigma)),
'wd1' : tf.Variable(tf.truncated_normal([7992, 1024], mu, sigma)),
'out' : tf.Variable(tf.truncated_normal([1024, n_classes], mu, sigma))
}
biases = {
'bc1' : tf.Variable(tf.zeros([108])),
'bc2' : tf.Variable(tf.zeros([108])),
'bd1' : tf.Variable(tf.zeros([1024])),
'out' : tf.Variable(tf.zeros([n_classes]))
}
return weights, biases
def conv2d(x, W, b, s=1):
conv = tf.nn.conv2d(x, W, strides=[1, s, s, 1], padding='VALID')
conv = tf.nn.bias_add(conv, b)
return tf.nn.relu(conv)
def maxpooling2d(x, k=2):
conv = tf.nn.max_pool(x,
ksize=[1, k, k, 1],
strides=[1, k, k, 1],
padding='VALID')
return conv
# +
from tensorflow.contrib.layers import flatten
def LeNet(x, keep_prob):
mu = 0
sigma = 0.1
W, b = get_weights_biases(mu, sigma)
#first layer
#Input = 32 x 32 x 1
#Output = 14 x 14 x 108
conv1 = conv2d(x, W['wc1'], b['bc1'])
conv1 = maxpooling2d(conv1)
print("1st layer shape : ", conv1.get_shape().as_list())
#second layer
#Input = 14 x 14 x 108
#Output = 7 x 7 x 108
conv1_subsample = maxpooling2d(conv1, k=2)
print("1st layer shape after subsample : ", conv1_subsample.get_shape().as_list())
#second layer
#Input = 14 x 14 x 108
#Output = 5 x 5 x 108
conv2 = conv2d(conv1, W['wc2'], b['bc2'])
conv2 = maxpooling2d(conv2)
print("2nd layer shape : ", conv2.get_shape().as_list())
#concatenated layer
#Output = 7992
conv2_shape = conv2.get_shape().as_list()
conv2_reshaped = tf.reshape(conv2, [-1, conv2_shape[1] * conv2_shape[2] * conv2_shape[3]])
conv1_subsample_shape = conv1_subsample.get_shape().as_list()
conv1_subsample_reshaped = tf.reshape(conv1_subsample, [-1,
conv1_subsample_shape[1] * conv1_subsample_shape[2] * conv1_subsample_shape[3]])
concatenated_layer = tf.concat(1, [conv2_reshaped, conv1_subsample_reshaped])
print("Concatenated layer shape : ", concatenated_layer.get_shape().as_list())
#third layer
#Input = 7992
#Output = 1024
fd1 = tf.add(tf.matmul(concatenated_layer, W['wd1']), b['bd1'])
fd1 = tf.nn.relu(fd1)
fd1 = tf.nn.dropout(fd1, keep_prob)
print("Third layer shape : ", fd1.get_shape().as_list())
#output layer
#Input = 1024
#Output = n_classes
out = tf.add(tf.matmul(fd1, W['out']), b['out'])
return out
# -
# ### Train, Validate and Test the Model
# A validation set can be used to assess how well the model is performing. A low accuracy on the training and validation
# sets imply underfitting. A high accuracy on the training set but low accuracy on the validation set implies overfitting.
x = tf.placeholder(tf.float32, (None, 32, 32, 1))
y = tf.placeholder(tf.int32, (None))
keep_prob = tf.placeholder(tf.float32)
one_hot_y = tf.one_hot(y, n_classes)
# +
rate = 0.0001
logits = LeNet(x, keep_prob)
cost = tf.reduce_mean(\
tf.nn.softmax_cross_entropy_with_logits(labels=one_hot_y,
logits=logits))
optimizer = tf.train.AdamOptimizer(learning_rate=rate).minimize(cost)
# +
correct_prediction = tf.equal(tf.argmax(logits, 1), tf.argmax(one_hot_y, 1))
accuracy_operation = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
saver = tf.train.Saver()
def evaluate(X_data, y_data):
num_examples = len(X_data)
total_accuracy = 0
sess = tf.get_default_session()
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_data[offset:offset + BATCH_SIZE], y_data[offset:offset + BATCH_SIZE]
accuracy = sess.run(accuracy_operation, feed_dict={x:batch_x,
y:batch_y,
keep_prob: 1.0})
total_accuracy += (accuracy * len(batch_x))
return total_accuracy / num_examples
# -
#train the model
with tf.Session() as sess:
# saver.restore(sess, './lenet-norm-gray')
sess.run(tf.global_variables_initializer())
num_examples = len(X_train_final)
for epoch in range(EPOCHS):
X_train_final, y_train_final = shuffle(X_train_final, y_train_final)
for offset in range(0, num_examples, BATCH_SIZE):
batch_x, batch_y = X_train_final[offset:offset+BATCH_SIZE], y_train_final[offset:offset+BATCH_SIZE]
sess.run(optimizer, feed_dict={x:batch_x,
y:batch_y,
keep_prob: 0.5})
validation_accuracy = evaluate(X_valid_final, y_valid)
print("EPOCH {}....".format(epoch+1))
print("Validation Accuracy = {:.3f}".format(validation_accuracy))
print()
saver.save(sess, './lenet-max-datax20')
print("Model saved")
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
test_accuracy = evaluate(X_test_final, y_test)
print("Test Accuracy = {:.3f}".format(test_accuracy))
# ---
#
# ## Step 3: Test a Model on New Images
#
# To give yourself more insight into how your model is working, download at least five pictures of German traffic signs from the web and use your model to predict the traffic sign type.
#
# You may find `signnames.csv` useful as it contains mappings from the class id (integer) to the actual sign name.
# ### Load and Output the Images
# +
import os
from PIL import Image
import random
import matplotlib.pyplot as plt
# Visualizations will be shown in the notebook.
# %matplotlib inline
image_list = os.listdir('new-signs/')
resized_images = []
fig = plt.figure(figsize=(20, 20))
columns = 2
rows = (len(image_list)/columns) + 1
for i, image_name in enumerate(image_list, 1):
image = Image.open('new-signs/' + image_name)
resized_image = image.resize((32, 32), Image.ANTIALIAS)
resized_images.append(resized_image)
fig.add_subplot(rows, columns, i)
plt.imshow(image)
plt.xlabel(image_name)
plt.show()
# +
#the first step would be to resize the images to 32x32, which we did while opening the image
#plotting the resized images
fig = plt.figure(figsize=(10, 10))
columns = 2
rows = (len(resized_images)/columns) + 1
for i, resized_image in enumerate(resized_images, 1):
fig.add_subplot(rows, columns, i)
plt.imshow(resized_image)
plt.xlabel(image_list[i - 1])
plt.show()
# -
# ### Predict the Sign Type for Each Image
# +
#convert pil images to numpy array
for i in range(len(resized_images)):
resized_images[i] = np.array(resized_images[i])
# +
image_labels = [14, 12, 21, 25, 11]
resized_images_final = single_channel(resized_images, 'Y')
resized_images_final = normalize(resized_images_final)
#plot pre-processed images images
fig = plt.figure(figsize=(10, 10))
columns = 2
rows = (len(resized_images_final)/columns) + 1
for i, resized_image in enumerate(resized_images_final, 1):
fig.add_subplot(rows, columns, i)
plt.imshow(resized_image.squeeze(), cmap="gray")
plt.xlabel(image_list[i - 1])
plt.show()
# +
predictions = tf.argmax(logits, 1)
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
# saver.restore(sess, './lenet-batch-size-128')
model_predictions = sess.run(predictions, feed_dict = { x: resized_images_final,
y: image_labels,
keep_prob: 1.0})
print(model_predictions)
# -
# ### Analyze Performance
### Calculate the accuracy for these 5 new images.
### For example, if the model predicted 1 out of 5 signs correctly, it's 20% accurate on these new images.
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
new_test_accuracy = evaluate(resized_images_final, image_labels)
print("Test Accuracy = {:.3f}".format(new_test_accuracy))
# ### Output Top 5 Softmax Probabilities For Each Image Found on the Web
# For each of the new images, print out the model's softmax probabilities to show the **certainty** of the model's predictions (limit the output to the top 5 probabilities for each image). [`tf.nn.top_k`](https://www.tensorflow.org/versions/r0.12/api_docs/python/nn.html#top_k) could prove helpful here.
#
# The example below demonstrates how tf.nn.top_k can be used to find the top k predictions for each image.
#
# `tf.nn.top_k` will return the values and indices (class ids) of the top k predictions. So if k=3, for each sign, it'll return the 3 largest probabilities (out of a possible 43) and the correspoding class ids.
#
# Take this numpy array as an example. The values in the array represent predictions. The array contains softmax probabilities for five candidate images with six possible classes. `tf.nn.top_k` is used to choose the three classes with the highest probability:
#
# ```
# # (5, 6) array
# a = np.array([[ 0.24879643, 0.07032244, 0.12641572, 0.34763842, 0.07893497,
# 0.12789202],
# [ 0.28086119, 0.27569815, 0.08594638, 0.0178669 , 0.18063401,
# 0.15899337],
# [ 0.26076848, 0.23664738, 0.08020603, 0.07001922, 0.1134371 ,
# 0.23892179],
# [ 0.11943333, 0.29198961, 0.02605103, 0.26234032, 0.1351348 ,
# 0.16505091],
# [ 0.09561176, 0.34396535, 0.0643941 , 0.16240774, 0.24206137,
# 0.09155967]])
# ```
#
# Running it through `sess.run(tf.nn.top_k(tf.constant(a), k=3))` produces:
#
# ```
# TopKV2(values=array([[ 0.34763842, 0.24879643, 0.12789202],
# [ 0.28086119, 0.27569815, 0.18063401],
# [ 0.26076848, 0.23892179, 0.23664738],
# [ 0.29198961, 0.26234032, 0.16505091],
# [ 0.34396535, 0.24206137, 0.16240774]]), indices=array([[3, 0, 5],
# [0, 1, 4],
# [0, 5, 1],
# [1, 3, 5],
# [1, 4, 3]], dtype=int32))
# ```
#
# Looking just at the first row we get `[ 0.34763842, 0.24879643, 0.12789202]`, you can confirm these are the 3 largest probabilities in `a`. You'll also notice `[3, 0, 5]` are the corresponding indices.
# +
### Print out the top five softmax probabilities for the predictions on the German traffic sign images found on the web.
### Feel free to use as many code cells as needed.
softmax_probs = tf.nn.softmax(logits)
with tf.Session() as sess:
saver.restore(sess, tf.train.latest_checkpoint('.'))
softmax_pred = sess.run(softmax_probs, feed_dict = { x: resized_images_final,
y: image_labels,
keep_prob: 1.0} )
top_5_preds = sess.run(tf.nn.top_k(tf.constant(softmax_pred), k=5))
print(top_5_preds)
# -
# ### Project Writeup
#
# Once you have completed the code implementation, document your results in a project writeup using this [template](https://github.com/udacity/CarND-Traffic-Sign-Classifier-Project/blob/master/writeup_template.md) as a guide. The writeup can be in a markdown or pdf file.
# > **Note**: Once you have completed all of the code implementations and successfully answered each question above, you may finalize your work by exporting the iPython Notebook as an HTML document. You can do this by using the menu above and navigating to \n",
# "**File -> Download as -> HTML (.html)**. Include the finished document along with this notebook as your submission.
# ---
#
# ## Step 4 (Optional): Visualize the Neural Network's State with Test Images
#
# This Section is not required to complete but acts as an additional excersise for understaning the output of a neural network's weights. While neural networks can be a great learning device they are often referred to as a black box. We can understand what the weights of a neural network look like better by plotting their feature maps. After successfully training your neural network you can see what it's feature maps look like by plotting the output of the network's weight layers in response to a test stimuli image. From these plotted feature maps, it's possible to see what characteristics of an image the network finds interesting. For a sign, maybe the inner network feature maps react with high activation to the sign's boundary outline or to the contrast in the sign's painted symbol.
#
# Provided for you below is the function code that allows you to get the visualization output of any tensorflow weight layer you want. The inputs to the function should be a stimuli image, one used during training or a new one you provided, and then the tensorflow variable name that represents the layer's state during the training process, for instance if you wanted to see what the [LeNet lab's](https://classroom.udacity.com/nanodegrees/nd013/parts/fbf77062-5703-404e-b60c-95b78b2f3f9e/modules/6df7ae49-c61c-4bb2-a23e-6527e69209ec/lessons/601ae704-1035-4287-8b11-e2c2716217ad/concepts/d4aca031-508f-4e0b-b493-e7b706120f81) feature maps looked like for it's second convolutional layer you could enter conv2 as the tf_activation variable.
#
# For an example of what feature map outputs look like, check out NVIDIA's results in their paper [End-to-End Deep Learning for Self-Driving Cars](https://devblogs.nvidia.com/parallelforall/deep-learning-self-driving-cars/) in the section Visualization of internal CNN State. NVIDIA was able to show that their network's inner weights had high activations to road boundary lines by comparing feature maps from an image with a clear path to one without. Try experimenting with a similar test to show that your trained network's weights are looking for interesting features, whether it's looking at differences in feature maps from images with or without a sign, or even what feature maps look like in a trained network vs a completely untrained one on the same sign image.
#
# <figure>
# <img src="visualize_cnn.png" width="380" alt="Combined Image" />
# <figcaption>
# <p></p>
# <p style="text-align: center;"> Your output should look something like this (above)</p>
# </figcaption>
# </figure>
# <p></p>
#
# +
### Visualize your network's feature maps here.
### Feel free to use as many code cells as needed.
# image_input: the test image being fed into the network to produce the feature maps
# tf_activation: should be a tf variable name used during your training procedure that represents the calculated state of a specific weight layer
# activation_min/max: can be used to view the activation contrast in more detail, by default matplot sets min and max to the actual min and max values of the output
# plt_num: used to plot out multiple different weight feature map sets on the same block, just extend the plt number for each new feature map entry
def outputFeatureMap(image_input, tf_activation, activation_min=-1, activation_max=-1 ,plt_num=1):
# Here make sure to preprocess your image_input in a way your network expects
# with size, normalization, ect if needed
# image_input =
# Note: x should be the same name as your network's tensorflow data placeholder variable
# If you get an error tf_activation is not defined it may be having trouble accessing the variable from inside a function
activation = tf_activation.eval(session=sess,feed_dict={x : image_input})
featuremaps = activation.shape[3]
plt.figure(plt_num, figsize=(15,15))
for featuremap in range(featuremaps):
plt.subplot(6,8, featuremap+1) # sets the number of feature maps to show on each row and column
plt.title('FeatureMap ' + str(featuremap)) # displays the feature map number
if activation_min != -1 & activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin =activation_min, vmax=activation_max, cmap="gray")
elif activation_max != -1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmax=activation_max, cmap="gray")
elif activation_min !=-1:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", vmin=activation_min, cmap="gray")
else:
plt.imshow(activation[0,:,:, featuremap], interpolation="nearest", cmap="gray")
| Traffic_Sign_Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="GAtifAnTu5xK" colab_type="text"
# # LDA Model for Visualization
# + id="TfUtCrFZvAOL" colab_type="code" colab={}
# imports needed for data
import pandas as pd
import numpy as np
import pickle
from sklearn.feature_extraction.text import CountVectorizer
# + id="hpDm-vVHbQB9" colab_type="code" outputId="3481ae57-5<PASSWORD>-45ef-fb2f-<PASSWORD>" colab={"base_uri": "https://localhost:8080/", "height": 207}
# read in the data with pandas
data = pd.read_parquet('clean_review_0.parquet')
data = data[['business_id', 'token']]
print(data.shape)
data.head()
# + id="1hlagXSP06eP" colab_type="code" outputId="bb8e6df6-1e05-4c58-b1db-9fa695bb1827" colab={"base_uri": "https://localhost:8080/", "height": 33}
# create a variable for later inputs
token = data['token']
token.shape
# + id="LHQy6odTq6JW" colab_type="code" colab={}
# Fit and transform the processed titles
cv = CountVectorizer(stop_words='english')
cvdata = cv.fit_transform(data['token'].astype(str))
# + id="VcWSwjxOq6SZ" colab_type="code" outputId="cb402cf9-6301-4f5e-a7c6-d1027308184e" colab={"base_uri": "https://localhost:8080/", "height": 310}
print(cvdata[0])
# + [markdown] id="GJs98sJCbm2h" colab_type="text"
# After fitting we can set up the corpus and dictionary
# + id="q8JCgLSHc-sV" colab_type="code" colab={}
# imports for LDA with Gensim
from gensim import matutils, models
import scipy.sparse
# + id="OSHXm5DQdcL4" colab_type="code" colab={}
# we're going to put the data into a new gensim format
sparse_counts = scipy.sparse.csr_matrix(cvdata)
corpus = matutils.Sparse2Corpus(sparse_counts)
# + id="UELR1ZTKdwI_" colab_type="code" colab={}
# gensim also requires a dictionary of all the terms, and possibly their location.
# cv = pickle.load(open("SOMETHING.pkl", "rb"))
id2word = dict((v, k) for k, v in cv.vocabulary_.items())
# + [markdown] id="j6by_GAsenyp" colab_type="text"
# now that we have the corpus (TDM) and id2word (dictionary of location: term) we will need to specify 2 other parameters - The nunber of Topics and The number of Passes. We'll start the number of topics at 2, see if it makes sense and adjust from there
# + id="NV12LYs5e-zo" colab_type="code" outputId="0863bc3a-c180-428d-dd43-4d83531345e6" colab={"base_uri": "https://localhost:8080/", "height": 82}
# set the lda model and the parameters
# 2 topics
lda = models.LdaModel(corpus=corpus, id2word=id2word, num_topics=2, passes=10)
lda.print_topics()
# + id="x5i4TYFCheoe" colab_type="code" outputId="b696b97f-64a3-4a74-b4bb-e4f1d702e143" colab={"base_uri": "https://localhost:8080/", "height": 115}
# 3 topics
lda = models.LdaModel(corpus=corpus, id2word=id2word, num_topics=3, passes=10)
lda.print_topics()
# + id="6mWvhjA2hewZ" colab_type="code" outputId="8e9c8f3d-8ef0-487d-a527-c4077b417098" colab={"base_uri": "https://localhost:8080/", "height": 147}
# 4 topics
lda = models.LdaModel(corpus=corpus, id2word=id2word, num_topics=4, passes=10)
lda.print_topics()
# + [markdown] id="mAWQ_gYNhNP9" colab_type="text"
# The output: first row shows the top words for the 1st topic, then below will be the rows for the 2nd topic, etc
#
# + [markdown] id="xLpNqgfjiP__" colab_type="text"
# The next level will be to get Nouns and Adjectives only. This will polish the topics being found.
# + id="nfGJ64ro11zU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="e17d233a-e014-4beb-ed55-4f936e750507"
# There was an error message later that said this install and download was required in order to move on
# !pip install nltk
# + id="vO6v-g6t2BP2" colab_type="code" colab={}
import nltk
# + id="vp0aEAsj1-uq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 82} outputId="f500b630-d0e4-4a5e-eaad-5766aacf315e"
nltk.download('averaged_perceptron_tagger')
# + [markdown] id="LtUe8nY72MZi" colab_type="text"
# Now that nltk was installed and imported
# + id="0r0KQB5He_1y" colab_type="code" colab={}
# Let's create a function to pull out the nouns and adj from the text.
# NN is used for nouns and JJ is used for Adjectives
from nltk import pos_tag
def nouns_adj(text):
is_noun_adj = lambda pos: pos[:2] == 'NN' or pos[:2] == 'JJ'
tokenized = token
nouns_adj = [word for (word, pos) in pos_tag(tokenized) if is_noun_adj]
return ' '.join(nouns_adj)
# + id="FoRCCFBc1X7Q" colab_type="code" colab={}
# read in the cleaned data, before the vectorizer step
data_clean = token
# + id="VMWbZMs-e_9k" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 361} outputId="63a46646-00cf-4cae-ff10-1617a2475857"
# apply the nouns adj function to the transcripts to filter
data_nouns_adj = pd.DataFrame(data_clean.apply(nouns_adj))
data_nouns_adj
# + [markdown] id="M1uOFvQ2kira" colab_type="text"
# the output will be each doc with their transcript
# + id="EvC_hf7Yktei" colab_type="code" colab={}
# create a new DTM only using the nouns and adj
data_cv = data_nouns_adj.transcript
data_dtm = pd.DataFrame(data_cv.toarray(), columns = data_cv.get_feature_names)
data_dtm.index = data_nouns_adj.index
data_dtm
# + [markdown] id="hC_SUa0Cm71X" colab_type="text"
# now we can recreate everything to include what we've made
#
# + id="rmpfKiFFnBDR" colab_type="code" colab={}
# create the gensim corpus
corpusna = matutils.Sparse2Corpus(scipy.sparse,scr_matrix(data_dtm.transpose()))
# create the vocabulary dictionary
id2wordna = dict((v, k) for k, v in data_cv.vocabulary_.items())
# + id="jrnIJ2uBn8F_" colab_type="code" colab={}
# start with 2 topics again
ldana = models.LdaModel(corpus=corpusna, num_topics=2, id2word=id2wordna, passes=10)
ldna.print_topics()
# + id="auITsU2LoTk4" colab_type="code" colab={}
# try 3 topics
ldana = models.LdaModel(corpus=corpusna, num_topics=3, id2word=id2wordna, passes=10)
ldna.print_topics()
# + id="eUHzC_wnojFP" colab_type="code" colab={}
# try 4 topics
ldana = models.LdaModel(corpus=corpusna, num_topics=4, id2word=id2wordna, passes=10)
ldna.print_topics()
# + [markdown] id="6jMdno48owh3" colab_type="text"
# When the topics start looking different we can go with that to the next step.
# + id="GV06Miy9ojNc" colab_type="code" colab={}
# run more iterations on our "final model"
# what increasing the passes does is it stabalizes which words falls into a topic
ldana = models.LdaModel(corpus=corpusna, num_topics=4, id2word=id2wordna, passes=80)
ldna.print_topics()
# + id="tYQNpxvrpTN4" colab_type="code" colab={}
# now we can look at which topic each doc or transcript contains
corpus_transformed = ldna[corpusna]
list(zip([a for [(a,b)] in corpus_transformed], data_dtm.index))
| NLP/LDA_Template_La_Teran_Evans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # UHFQA
# Just like the driver for the HDAWG in the previous example, we now use the `tk.UHFQA` instrument driver.
# +
import numpy as np
import matplotlib.pyplot as plt
import qcodes as qc
import zhinst.qcodes as ziqc
uhfqa = ziqc.UHFQA("qa1", "dev2266", interface="1gbe", host="10.42.0.226")
# -
print([k for k in uhfqa.submodules.keys()])
print([k for k in uhfqa.parameters.keys()])
# ## AWG core of the UHFQA
#
# Also the UHFQA features one *AWG Core*.
print([k for k in uhfqa.awg.parameters.keys()])
# ## Readout channels of the UHFQA
#
# The UHFQA comes with signal processing streams for up to ten channels in parallel. The settings for the readout are grouped by channel in a list of all ten `channels`. Each item in the `channels` property of the UHFQA is an *Instrument Channel* that represent the signal processing path for one of the ten channels.
print([k for k in uhfqa.channels[0].parameters.keys()])
# Each of the channels follows the following signal processing steps:
#
# 1. Demodulation of the input signal
# 2. Rotation in the complex plane
# 3. Thresholding for binary result values
#
#
# The values for the rotation and thresholding stages can be set using the `rotation` and `threshold` parameter of the *channel*.
#
# The standard mode for the demodulation of input signals is the *weighted integration* mode. This corresponds to setting the integration weights for the two quadratures of the input signal to oscillate at a given demodulation frequency. When enabling the weighted integration with `ch.enable()`, the integration weights for the two quadratures are set. The demodulation frequency is set to the parameter `readout_frequency`.
#
# Enabling weighted integration for the first four channels of the UHFQA and setting their demodulation frequency could look like this:
# +
freqs = [85.6e6, 101.3e6, 132.8e6]
for ch in uhfqa.channels[:3]:
ch.enable()
ch.readout_frequency(freqs[ch.index])
# -
# The resut vector of each channel can be retrieved from the instrument by calling the read-only parameter *result*.
print(uhfqa.channels[0].result.__doc__)
# ## Readout parameters
#
# There are readout parameters taht are not specific to one isngle channel but affect all ten readout channels. These are
#
# * the `integration_time`: the time in seconds used for integrating the input signals
# * the `result_source` lets the user select at which point in the signal processing path the `result` value should be taken
# * the `averaging_mode` specifies if the hardware averages on the device should be taken in a *sequential* or *cyclic* way
# * the `crosstalk_matrix` specifies a 10 x 10 matrix that can be calibrated to compensate for crosstalk betweeen the channels
#
# These three *parameters* are attributes of the UHFQA instrument driver.
print(uhfqa.integration_time.__doc__)
print(uhfqa.result_source.__doc__)
print(uhfqa.averaging_mode.__doc__)
print(uhfqa.crosstalk_matrix.__doc__)
# Other important readout parameters can be accessed through the *nodetree*, for example the
#
# * *result length*: the number of points to acquire
# * *result averages*: the number of hardware averages
print(uhfqa.qas[0].result.length.__doc__)
print(uhfqa.qas[0].result.averages.__doc__)
# ## *Arm* the UHFQA readout
#
# The `arm(...)` method of the UHFQA prepares the device for data acquisition. It enables the *Results Acquisition* and resets the acquired points to zero. This should be done before every measurement. The method also includes a shortcut to setting the values *result length* and *result averages*. They can be specified as keyword arguments. If the keyword arguemnts are not specified, nothing is changed.
uhfqa.arm(length=1e3, averages=2**5)
| examples/example2-2_UHFQA.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflowGPU]
# language: python
# name: conda-env-tensorflowGPU-py
# ---
# +
# %pylab inline
import numpy as np
import tensorflow as tf
from scipy import integrate
from mpl_toolkits.mplot3d import Axes3D
import keras
from keras import optimizers
from keras.models import Model,Sequential,load_model
from keras.layers import Input,Dense, Activation
from keras import backend as K
from keras.utils.generic_utils import get_custom_objects
from keras.utils import plot_model
from IPython.display import clear_output
# +
class PlotLosses(keras.callbacks.Callback):
def on_train_begin(self, logs={}):
self.i = 0
self.x = []
self.losses = []
self.val_losses = []
self.fig = plt.figure()
self.logs = []
def on_epoch_end(self, epoch, logs={}):
self.logs.append(logs)
self.x.append(self.i)
self.losses.append(logs.get('loss'))
self.val_losses.append(logs.get('val_loss'))
self.i += 1
clear_output(wait=True)
plt.plot(self.x, self.losses, label="loss")
plt.plot(self.x, self.val_losses, label="val_loss")
plt.yscale('log')
plt.legend()
plt.show();
plot_losses = PlotLosses()
# -
def progress_bar(percent):
length = 40
pos = round(length*percent)
clear_output(wait=True)
print('['+'█'*pos+' '*(length-pos)+'] '+str(int(100*percent))+'%')
# ## Set up Trajectories
# define functions to help create trajectories to given Lorenz equation using random initial conditions
sigma=10; beta=8/3; rho=28;
def lrz_rhs(t,x):
return [sigma*(x[1]-x[0]), x[0]*(rho-x[2]), x[0]*x[1]-beta*x[2]];
end_time = 8
sample_rate = 100
t = np.linspace(0,end_time,sample_rate*end_time,endpoint=True)
def lrz_trajectory():
x0 = 20*(np.random.rand(3)-.5)
sol = integrate.solve_ivp(lrz_rhs,[0,end_time],x0,t_eval=t,rtol=1e-10,atol=1e-11)
return sol.y
x = lrz_trajectory()
plt.figure()
plt.gca(projection='3d')
plt.plot(x[0],x[1],x[2])
plt.show()
# ## Generate Data
# `Y` is composed of position vectors one step forward in time from those in `X`. Data comes from `N` trajectories each with `traj_length` entries
from scipy.io import loadmat
xy = loadmat('xy.mat')
X = xy['input']
Y = xy['output']
N = 200
D = np.zeros((N,3,len(t)))
for i in range(N):
progress_bar((i+1)/N)
D[i] = lrz_trajectory()
#np.savez('trajectories',D=D)
# +
#D = np.load('trajectories.npz')['D']
# -
X = np.transpose(D[:,:,:-1],axes=[0,2,1]).reshape(-1,3)
Y = np.transpose(D[:,:,1:],axes=[0,2,1]).reshape(-1,3)
i=231
X[i]==Y[i-1]
np.shape(X)
# +
num_epochs = 500
input_shape = (X.shape[1],)
inputs = Input(shape = input_shape)
x = Dense(output_dim = 100, activation = 'sigmoid')(inputs)
#x = Dense(output_dim = 512, activation = 'selu')(inputs)
#x = Dense(output_dim = 200, activation = 'sigmoid')(x)
#x = Dense(output_dim = 500, activation = 'elu')(x)
x = Dense(output_dim = 3, activation = 'linear')(x)
output = x
model = Model(input=inputs,output=output)
lr = 0.01
#decay = lr/num_epochs-1e-9 #optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
optimizer = optimizers.Adam(lr=lr)#optimizers.rmsprop(lr=lr)
model.compile(optimizer=optimizer, loss='mean_squared_error') #compiling here
epoch = num_epochs
model.fit(X, Y, batch_size=600, epochs=num_epochs, validation_split=0.05, callbacks=[], verbose=1)
# -
# ## Create Neural Net Model
# How do we pick this?
# +
def rad_bas(x):
return K.exp(-x**2)
get_custom_objects().update({'rad_bas': Activation(rad_bas)})
def tan_sig(x):
return 2/(1+K.exp(-2*x))-1
get_custom_objects().update({'tan_sig': Activation(tan_sig)})
# +
x = keras.layers.Input(shape=(3,))
l1 = Dense(32, activation='sigmoid', use_bias=True)(x)
l2 = Dense(64, activation='rad_bas', use_bias=True)(l1)
l3 = Dense(32, activation='sigmoid', use_bias=True)(l2)
l4 = Dense(16, activation='linear', use_bias=True)(l3)
y = Dense(3)(l1)
model = Model(inputs=x,outputs=y)
# -
model = Sequential()
model.add(Dense(100, activation='sigmoid', use_bias=True, input_shape=(3,)))
#model.add(Dense(10, activation='tan_sig', use_bias=True))
#model.add(Dense(10, activation='linear', use_bias=True))
model.add(Dense(3))
# +
x = keras.layers.Input(shape=(3,))
b0 = Dense(3, use_bias=True)(x)
b1 = keras.layers.Multiply()([x,b0])
y = Dense(3)(b1)
#c0 = Dense(3, activation='sigmoid')(x)
#y = keras.layers.Add()([c0,b2])
model = Model(inputs=x, outputs=y)
# +
inputs = Input(shape = input_shape)
x = Dense(output_dim = 100, activation = 'sigmoid')(inputs)
#x = Dense(output_dim = 512, activation = 'selu')(inputs)
#x = Dense(output_dim = 200, activation = 'sigmoid')(x)
#x = Dense(output_dim = 500, activation = 'elu')(x)
x = Dense(output_dim = 3, activation = 'linear')(x)
output = x
model = Model(input=inputs,output=output)
lr = 0.01
#decay = lr/num_epochs-1e-9 #optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
optimizer = optimizers.Adam(lr=lr)#optimizers.rmsprop(lr=lr)
model.compile(optimizer=optimizer, loss='mean_squared_error') #compiling here
# -
# ## Compile Model
sgd1 = optimizers.SGD(lr=0.001, decay=1e-15, momentum=1, nesterov=True)
adam1 = optimizers.Adam(lr=.01)
nadam1 = keras.optimizers.Nadam(lr=0.02, beta_1=0.9, beta_2=0.999, epsilon=None, schedule_decay=0.004)
rmsprop1 = keras.optimizers.RMSprop(lr=0.01, rho=0.9, epsilon=None, decay=0.0)
model.compile(loss='mean_squared_error', optimizer=adam1, metrics=['accuracy'])
#plot_model(model, to_file='model.pdf', show_shapes=True)
lr = 0.01
#decay = lr/num_epochs-1e-9 #optimizers.SGD(lr=0.01, decay=1e-6, momentum=0.9, nesterov=True)
optimizer = optimizers.Adam(lr=lr)#optimizers.rmsprop(lr=lr)
model.compile(optimizer=optimizer, loss='mean_squared_error') #compiling here
# ## Fit Model
model.fit(X, Y, epochs=300, batch_size=800, shuffle=True, callbacks=[], validation_split=0.0, verbose=1)
model.save('200adam_3.h5')
model = load_model('lrz_model_basic.h5')
x = np.zeros((3,end_time*sample_rate))
x[:,0] = 30*(np.random.rand(3)-1/2)
for i in range(end_time*sample_rate-1):
x[:,i+1] = model.predict(np.array([x[:,i]]))
xsol = integrate.solve_ivp(lrz_rhs,[0,end_time],x[:,0],t_eval=t,rtol=1e-10,atol=1e-11).y
plt.figure()
plt.gca(projection='3d')
plt.plot(x[0],x[1],x[2])
plt.plot(xsol[0],xsol[1],xsol[2])
plt.show()
for i in range(3):
plt.figure()
plt.plot(t,x[i])
plt.plot(t,xsol[i])
plt.show()
len(X)
x[1]
len(t)
| amath563/hw2/.ipynb_checkpoints/NN_lrz-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#This are the most common imports
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from datetime import datetime, timedelta
# %matplotlib inline
# # Dataset basic walkthrough
df = pd.read_csv('hmofaltasadministrativas.csv', encoding="latin1")
# Make all letters uppercase
df = df.applymap(lambda s:s.upper() if type(s) == str else s)
# The dataset is from 01/01/2015 through 05/07/2018
# We will delete the data from 2018
#Obtenemos dia, mes y año de la columna de fecha_presentación del dataframe
df['AÑO'] = pd.DatetimeIndex(df['fecha_presentacion']).year
df['MES'] = pd.DatetimeIndex(df['fecha_presentacion']).month
df['DIA'] = pd.DatetimeIndex(df['fecha_presentacion']).day
df['HORA'] = pd.DatetimeIndex(df['fecha_presentacion']).hour
df['MINUTE'] = pd.DatetimeIndex(df['fecha_presentacion']).minute
df.drop('fecha_presentacion', axis= 1, inplace = True)
df.tail()
df = df[df['AÑO'] < 2018]
df.info()
# # Top 10 Colonias con mas delitos
colonias_delitos = df['colonia_delito'].value_counts()
colonias_delitos_top10 = df['colonia_delito'].value_counts()[0:10]
colonias_delitos_plt = colonias_delitos.plot(kind='barh')
plt.close()
df_colonias_delitos_top10 = pd.DataFrame(colonias_delitos_top10)
df_colonias_delitos_top10
colonias_delitos_top10_plt = colonias_delitos_top10.plot(kind='barh',figsize = (14,8))
colonias_delitos_top10_plt.set_alpha(0.8)
colonias_delitos_top10_plt.set_title("Top 10 de colonias con mas delitos 2015-2017", fontsize=18)
# create a list to collect the data
totals = []
# find the values and append to list
for i in colonias_delitos_plt.patches:
totals.append(i.get_width())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in colonias_delitos_top10_plt.patches:
# get_width pulls left or right; get_y pushes up or down
colonias_delitos_top10_plt.text(i.get_width()+.3, i.get_y()+.38, \
str(round((i.get_width()/total)*100, 2))+'%', fontsize=10,
color='black')
#Invert the direction of the plot
colonias_delitos_top10_plt.invert_yaxis()
# Since colonia Centro (N) has de highest percentage of crimes we will focus on that one.
# # Colonia Centro (N)
# ## Dataset
df_colonia_centro = df[df['colonia_delito'] == 'CENTRO (N)']
df_colonia_centro.head(2)
# ## Top 10 Motivos de remisión en colonia Centro (N)
colonia_centro = df_colonia_centro['motivo_remision'].value_counts()
colonia_centro_top10 = colonia_centro[0:10]
colonia_centro_plt = colonia_centro.plot(kind='barh')
plt.close()
df_colonia_centro_top10 = pd.DataFrame(colonia_centro_top10)
df_colonia_centro_top10
colonia_centro_top10_plt = colonia_centro_top10.plot(kind='barh',figsize = (20,12))
colonia_centro_top10_plt.set_alpha(0.8)
colonia_centro_top10_plt.set_title("Top 10 motivos de remision en colonia Centro (N) 2015-2017",
fontsize=24)
colonia_centro_top10_plt.tick_params(labelsize=21)
# create a list to collect the data
totals = []
# find the values and append to list
for i in colonia_centro_plt.patches:
totals.append(i.get_width())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in colonia_centro_top10_plt.patches:
# get_width pulls left or right; get_y pushes up or down
colonia_centro_top10_plt.text(i.get_width()+.3, i.get_y()+.38, \
str(round((i.get_width()/total)*100, 2))+'%', fontsize=19,
color='black')
#Invert the direction of the plot
colonia_centro_top10_plt.invert_yaxis()
# ## Cuando se dieron los delitos de "Causar o provocar escandalo en lugares públicos o privados en la colonia Centro (N)
df_colonia_centro.head(2)
# +
df_escandalo_colcentro = df_colonia_centro[df_colonia_centro['motivo_remision'] ==
'CAUSAR O PROVOCAR ESCANDALO EN LUGARES PUBLICOS O PRIVADOS']
plt.figure(figsize=(18,12))
escandalo_colcentro_plt = sns.countplot(x='MES',data=df_escandalo_colcentro, palette="muted")
plt.title('Frecuencia de CAUSAR O PROVOCAR ESCANDALO EN LUGARES PUBLICOS O PRIVADOS en colonia Centro (N)')
plt.xlabel('Mes')
plt.ylabel('Frequency')
labels = ["Enero", "Febrero", "Marzo", "Abril", "Mayo", 'Junio', 'Julio', 'Agosto', 'Septiembre',
"Octubre", 'Noviembre', 'Diciembre']
escandalo_colcentro_plt.set_xticklabels(labels, rotation = 45 , fontsize=18)
# -
# ### Porcentaje de reincidencias del delito CAUSAR O PROVOCAR ESCANDALO EN LUGARES PUBLICOS O PRIVADOS en colonia CENTRO (N)
df_escandalo_colcentro_reincidencias = df_escandalo_colcentro.drop_duplicates(subset=
['colonia_detenido','nacimiento',
'estatura', 'peso','sexo'],
keep = 'last')
(1-(len(df_escandalo_colcentro_reincidencias)/len(df_escandalo_colcentro)))*100
# +
df_escandalo_colcentro_reincidencias = df_escandalo_colcentro[df_escandalo_colcentro.duplicated(['colonia_detenido',
'nacimiento', 'estatura', 'peso',
'sexo']) == True]
plt.figure(figsize=(16,10))
df_escandalo_colcentro_reincidencias_plt = sns.countplot(x='MES',data=df_escandalo_colcentro_reincidencias, palette="muted")
plt.title('Frecuencia de reincidencias de CAUSAR O PROVOCAR ESCANDALO EN LUGARES PUBLICOS O PRIVADOS en colonia Centro (N)')
plt.xlabel('Mes')
plt.ylabel('Frequency')
labels = ["Enero", "Febrero", "Marzo", "Abril", "Mayo", 'Junio', 'Julio', 'Agosto', 'Septiembre',
"Octubre", 'Noviembre', 'Diciembre']
df_escandalo_colcentro_reincidencias_plt.set_xticklabels(labels, rotation = 45 , fontsize=18)
# -
# ##
colonia_centro_colonia_detenido = df_escandalo_colcentro['colonia_detenido'].value_counts()
colonia_centro_colonia_detenido_top10 = colonia_centro_colonia_detenido[0:10]
colonia_centro_colonia_detenido_plt = colonia_centro.plot(kind='barh')
plt.close()
colonia_centro_colonia_detenido_top10 = pd.DataFrame(colonia_centro_colonia_detenido_top10)
colonia_centro_colonia_detenido_top10
colonia_centro_colonia_detenido_top10_plt = colonia_centro_colonia_detenido_top10.plot(kind='barh',figsize = (20,12))
colonia_centro_colonia_detenido_top10_plt.set_alpha(0.8)
colonia_centro_colonia_detenido_top10_plt.set_title("Donde viven las personas que han sido detenidas por Causar escandalo en la colonia CENTRO(N) ",
fontsize=24)
colonia_centro_colonia_detenido_top10_plt.tick_params(labelsize=18)
# create a list to collect the data
totals = []
# find the values and append to list
for i in colonia_centro_colonia_detenido_top10_plt.patches:
totals.append(i.get_width())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in colonia_centro_colonia_detenido_top10_plt.patches:
# get_width pulls left or right; get_y pushes up or down
colonia_centro_colonia_detenido_top10_plt.text(i.get_width()+.3, i.get_y()+.38, \
str(round((i.get_width()/total)*100, 2))+'%', fontsize=18,
color='black')
#Invert the direction of the plot
colonia_centro_colonia_detenido_top10_plt.invert_yaxis()
# ## Top 10 Motivos de remision
motivo_remision = df['motivo_remision'].value_counts()
motivo_remision_top10 = df['motivo_remision'].value_counts()[0:10]
motivo_remision_plt = motivo_remision.plot(kind='barh')
plt.close()
df_motivo_remision_top10 = pd.DataFrame(motivo_remision_top10)
df_motivo_remision_top10
motivo_remision_top10_plt = motivo_remision_top10.plot(kind='barh',figsize = (20,16))
motivo_remision_top10_plt.set_alpha(0.8)
motivo_remision_top10_plt.set_title("Top 10 de colonias con mas delitos 2015-2017", fontsize=14)
# create a list to collect the data
totals = []
# find the values and append to list
for i in motivo_remision_plt.patches:
totals.append(i.get_width())
# set individual bar lables using above list
total = sum(totals)
# set individual bar lables using above list
for i in motivo_remision_top10_plt.patches:
# get_width pulls left or right; get_y pushes up or down
motivo_remision_top10_plt.text(i.get_width()+.3, i.get_y()+.38, \
str(round((i.get_width()/total)*100, 2))+'%', fontsize=20,
color='black')
#Invert the direction of the plot
motivo_remision_top10_plt.invert_yaxis()
# +
df_motivo_remision_mes = df[df['motivo_remision'] ==
'DEAMBULAR EN LA VIA PUBLICA EN ESTADO DE EMBRIAGUEZ O DROGADO']
plt.figure(figsize=(18,12))
motivo_remision_mes_plt = sns.countplot(x='MES',data=df_motivo_remision_mes, palette="muted")
plt.title('Frecuencia de DEAMBULAR EN LA VIA PUBLICA EN ESTADO DE EMBRIAGUEZ O DROGADO')
plt.xlabel('Mes')
plt.ylabel('Frecuencia')
labels = ["Enero", "Febrero", "Marzo", "Abril", "Mayo", 'Junio', 'Julio', 'Agosto', 'Septiembre',
"Octubre", 'Noviembre', 'Diciembre']
motivo_remision_mes_plt.set_xticklabels(labels, rotation = 45 , fontsize=18)
#for item in motivo_remision_mes_plt.patches:
# height = item.get_height()
# df_motivo_remision_mes.text(item.get_x()+item.get_width()/2.,
# height + 3,
# '{:1.2f}'.format(height/total*100)+'%',
# ha="center")
# -
# # Predictions
df_colonia_centro.head(2)
new_df = df_colonia_centro.copy()
#new_df['motivo_remision'] = pd.factorize(new_df['motivo_remision'], sort=True)[0] + 1
new_df['zona'] = pd.factorize(new_df['zona'], sort=True)[0] + 1
new_df['colonia_delito'] = pd.factorize(new_df['colonia_delito'], sort=True)[0] + 1
new_df['sexo'] = pd.factorize(new_df['sexo'], sort=True)[0] + 1
new_df['zona'] = pd.factorize(new_df['zona'], sort=True)[0] + 1
new_df['motivo_remision'] = pd.factorize(new_df['motivo_remision'], sort=True)[0] + 1
new_df['colonia_detenido'] = pd.factorize(new_df['colonia_detenido'], sort=True)[0] + 1
new_df.head()
new_df['Reincidencia'] = new_df.duplicated(subset= ['colonia_detenido', 'motivo_remision', 'nacimiento', 'estatura', 'peso', 'sexo'])
new_df.head()
new_df['Reincidencia'].sum()
new_df.dropna(inplace= True)
# +
#This is for splitting the train data vs the test data
from sklearn.model_selection import train_test_split
#X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.3, random_state = 101)
# This is for Logistic Regression
from sklearn.linear_model import LogisticRegression
#This is a import for classification_report, it will show the accuracy of the training model
from sklearn.metrics import classification_report, confusion_matrix
#This is a import for Random Forest
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
#Steps to predict
from sklearn.ensemble import RandomForestRegressor
from IPython.display import display
# -
X = new_df.drop('Reincidencia',
axis = 1)
y = new_df['Reincidencia']
X_train, X_test, y_train, y_test = train_test_split(X,y, test_size=0.3, random_state = 101)
# Escalar los datos
from sklearn.preprocessing import StandardScaler
standardScaler = StandardScaler()
X_train = standardScaler.fit_transform(X_train)
X_test = standardScaler.transform(X_test)
# ## Random Forest
rfc = RandomForestClassifier(n_estimators = 10,criterion = 'gini')
rfc.fit(X_train, y_train)
prediction = rfc.predict(X_test)
#prediction = rfc.predict(X_test)
print(confusion_matrix(y_test, prediction))
print('\n')
print(classification_report(y_test, prediction))
# Create the parameter grid to optimize
param_frst = [
{
'criterion': ['gini', 'entropy'],
'bootstrap': [True, False],
'n_estimators': [50, 100],
'max_depth': [2, 5],
'max_leaf_nodes': [10, 20]
}
]
# Escoger hiperparametros para modelo Random Forest
from sklearn.model_selection import GridSearchCV
#param_frst = [{"n_estimators": [100,500,750,1000], "criterion": ["gini", "entropy"]}]
grid_search_frst = GridSearchCV(estimator=rfc,
param_grid=param_frst,
scoring = 'accuracy',
cv=3,
n_jobs=-1)
grid_search_frst = grid_search_frst.fit(X_train, y_train)
# Calcular accuracy para Random Forest
best_acc_frst = grid_search_frst.best_score_
best_acc_frst
# Calcular los mejores parametros para el modelo Random Forest
best_params_frst = grid_search_frst.best_params_
best_params_frst
rfc = RandomForestClassifier(bootstrap= False,
criterion= 'gini',
max_depth= 5,
max_leaf_nodes = 20,
n_estimators= 50)
rfc.fit(X_train, y_train)
prediction = rfc.predict(X_test)
#prediction = rfc.predict(X_test)
print(confusion_matrix(y_test, prediction))
print('\n')
print(classification_report(y_test, prediction))
#Confusion matrix
from sklearn.metrics import confusion_matrix
frst_cm = confusion_matrix(y_test, prediction)
fig = plt.figure(figsize = (5,5))
sns.heatmap(frst_cm,annot=True,fmt='5.0f',cmap="coolwarm")
plt.title('Confusion matrix para Random Forest', y=2.05, size=16)
# +
conf_mat = confusion_matrix(y_test, prediction, labels=np.sort(y_test.unique()))
conf_mat_df = pd.DataFrame(
conf_mat)
conf_mat_props = pd.DataFrame(
conf_mat_df.values / conf_mat_df.sum(axis=1)[:,None])
fig = plt.figure(figsize = (6,6))
sns.heatmap(conf_mat_props, annot=True, cmap= 'coolwarm');
# -
# ## Logistic Regression
# +
# iniciar y ajustar el modelo de regresión logistica en el dataset de entrenamiento.
# primero importamos la libreria de Regresion Logistica de SkLearn
from sklearn.linear_model import LogisticRegression
log_rg = LogisticRegression()
log_rg.fit(X_train, y_train)
# Predicción
y_log_rg = log_rg.predict(X_test)
# -
# Confusion matrix
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_log_rg)
sns.heatmap(cm,annot=True,fmt='3.0f',cmap="Greens")
plt.title('Confusion matrix para Regresión Logistica', y=1.05, size=15)
# Reporte de Clasificación
from sklearn.metrics import classification_report
cr = classification_report(y_test, y_log_rg)
print(cr)
# +
# Grid search cross validation
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import LogisticRegression
grid={"C":np.logspace(-3,3,7), "penalty":["l1","l2"]}# l1 lasso l2 ridge
logreg=LogisticRegression()
logreg_cv=GridSearchCV(logreg,grid,cv=10)
logreg_cv.fit(X_train,y_train)
print("tuned hpyerparameters :(best parameters) ",logreg_cv.best_params_)
print("accuracy :",logreg_cv.best_score_)
# -
# ## Naive bayes
#
# +
# Iniciar y ajustar el modelo de Naive Bayes al dataset de entrenamiento.
from sklearn.naive_bayes import GaussianNB
naive_b = GaussianNB()
naive_b.fit(X_train, y_train)
#Predicción
y_naive = naive_b.predict(X_test)
# -
#Confusion matrix
from sklearn.metrics import confusion_matrix
naive_cm = confusion_matrix(y_test, y_naive)
sns.heatmap(naive_cm,annot=True,fmt='3.0f',cmap="Blues")
plt.title('Confusion matrix para Naive Bayes', y=1.05, size=15)
# Reporte de Clasificación
from sklearn.metrics import classification_report
naive_cr = classification_report(y_test, y_naive)
print(naive_cr)
| Faltas Administrativas Hermosillo/HMO Faltas administrativas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # %matplotlib inline
from matplotlib import style
style.use('fivethirtyeight')
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
import datetime as dt
# # Reflect Tables into SQLAlchemy ORM
# Python SQL toolkit and Object Relational Mapper
import sqlalchemy
from sqlalchemy.ext.automap import automap_base
from sqlalchemy.orm import Session
from sqlalchemy import create_engine, func, inspect
engine = create_engine("sqlite:///Resources/hawaii.sqlite", echo=False)
# +
# Could not read sqlite file so the following is to help me understand the data
# -
engine.execute('SELECT * FROM measurement LIMIT 5').fetchall()
inspector = inspect(engine)
columns = inspector.get_columns('station')
for c in columns:
print(c['name'], c['type'])
inspector = inspect(engine)
columns = inspector.get_columns('measurement')
for c in columns:
print(c['name'], c['type'])
# +
# reflect an existing database into a new model
Base = automap_base()
# reflect the tables
Base.prepare(engine, reflect = True)
# -
# We can view all of the classes that automap found
Base.classes.keys()
# Save references to each table
Measurement = Base.classes.measurement
Station = Base.classes.station
# Create our session (link) from Python to the DB
session = Session(engine)
# # Exploratory Climate Analysis
# +
# Design a query to retrieve the last 12 months of precipitation data and plot the results
# +
# Calculate the date 1 year ago from the last data point in the database
# Last data point in the database
last_date = session.query(Measurement.date).order_by(Measurement.date.desc()).first()
# # last_date = 2017-08-23
# 1 year data from last data point
a_year_ago = dt.date(2017,8,23) - dt.timedelta(days=365)
a_year_ago
# +
# Perform a query to retrieve the data and precipitation scores
scores = session.query(Measurement.date, func.sum(Measurement.prcp)).\
filter(func.strftime("%Y-%m-%d", Measurement.date) >= a_year_ago).\
group_by(Measurement.date).\
order_by(Measurement.date).all()
# scores
# +
# Save the query results as a Pandas DataFrame and set the index to the date column
# Sort the dataframe by date
scores_df = pd.DataFrame(scores, columns = ['Date', 'Precipitation']).set_index(['Date']).sort_index(ascending = True)
scores_df.head()
# +
# Use Pandas Plotting with Matplotlib to plot the data
fig, ax = plt.subplots(figsize=[10,5])
ax.bar(scores_df.index.values, scores_df['Precipitation'], color= "green" , label = 'Precipitation')
# importing additional packages
import matplotlib.dates as mdates
plt.ylim(0, 15)
ax.xaxis.set_major_locator(mdates.WeekdayLocator(interval=5))
plt.xticks(rotation='vertical')
plt.xlabel('Date')
plt.ylabel('Total Precipitation (Inches)')
# plt.title()
ax.legend()
plt.tight_layout()
plt.show()
# -
# Use Pandas to calcualte the summary statistics for the precipitation data
scores_df.describe()
# ### QUERY - above count results should be 365 days????
# # Station Analysis
# Design a query to show how many stations are available in this dataset?
station_count = session.query(Measurement.station).group_by(Measurement.station).count()
station_count
# +
# What are the most active stations? (i.e. what stations have the most rows)?
# List the stations and the counts in descending order.
most_active = session.query(Measurement.station, func.count(Measurement.station)).\
group_by(Measurement.station).\
order_by(func.count(Measurement.station).desc()).\
all()
most_active
# +
# Using the station id from the previous query, calculate the lowest temperature recorded,
# highest temperature recorded, and average temperature of the most active station?
stats_most_active = session.query(Measurement.station, func.min(Measurement.tobs), func.max(Measurement.tobs), func.avg(Measurement.tobs)).\
filter(Measurement.station == 'USC00519281').all()
stats_most_active
# +
# Choose the station with the highest number of temperature observations.
# Query the last 12 months of temperature observation data for this station and plot the results as a histogram
# CHECK - whether station with the highest number of temp observation is still the same as USC00519281. The instruction for this bit is a bit vague.....
highest_tobs = session.query(Measurement.station, func.count(Measurement.tobs)).\
group_by(Measurement.station).\
order_by(func.count(Measurement.station).desc()).\
all()
# highest_tobs
# CHECK!
# Last 12 months of tobs
most_active_station = session.query(Measurement.station, Measurement.tobs).\
filter(Measurement.station == 'USC00519281').\
filter(func.strftime("%Y-%m-%d", Measurement.date) >= a_year_ago).\
group_by(Measurement.date).\
order_by(Measurement.date).all()
# most_active_station
waihee_df = pd.DataFrame(most_active_station, columns = ['Station_ID', 'Temp_Obs']).set_index(['Station_ID'])
# waihee_df.head()
hist = plt.hist(waihee_df['Temp_Obs'], bins=12, label = 'tobs')
plt.xlabel('Temperature')
plt.ylabel('Frequency')
plt.legend()
# plt.xlim(55, 90)
plt.ylim(0, 70)
plt.show()
# -
# ## Bonus Challenge Assignment
# +
# This function called `calc_temps` will accept start date and end date in the format '%Y-%m-%d'
# and return the minimum, average, and maximum temperatures for that range of dates
def calc_temps(start_date, end_date):
"""TMIN, TAVG, and TMAX for a list of dates.
Args:
start_date (string): A date string in the format %Y-%m-%d
end_date (string): A date string in the format %Y-%m-%d
Returns:
TMIN, TAVE, and TMAX
"""
return session.query(func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)).\
filter(Measurement.date >= start_date).\
filter(Measurement.date <= end_date).all()
# function usage example
print(calc_temps('2012-02-28', '2012-03-05'))
# +
# Use your previous function `calc_temps` to calculate the tmin, tavg, and tmax
# for your trip using the previous year's data for those same dates.
# Start date = '2016-12-16'
# End date = '2016-12-31'
print(calc_temps('2015-12-16', '2015-12-31'))
# +
# Plot the results from your previous query as a bar chart.
# Use "Trip Avg Temp" as your Title
# Use the average temperature for the y value
# Use the peak-to-peak (tmax-tmin) value as the y error bar (yerr)
trip_temp_df = pd.DataFrame(calc_temps('2015-12-16', '2015-12-31'), columns = ['Min Temp', 'Avg Temp', 'Max Temp'])
# trip_temp_df
y_value = trip_temp_df['Avg Temp']
plt.bar(y_value, figsize=[10,5])
plt.title('Trip Avg Temp')
# plt.ylim(0, 15)
# plt.xticks(rotation='vertical')
# plt.xlabel('')
# plt.ylabel('Temp (F)')
# plt.show()
# +
# Calculate the total amount of rainfall per weather station for your trip dates using the previous year's matching dates.
# Sort this in descending order by precipitation amount and list the station, name, latitude, longitude, and elevation
# +
# Create a query that will calculate the daily normals
# (i.e. the averages for tmin, tmax, and tavg for all historic data matching a specific month and day)
def daily_normals(date):
"""Daily Normals.
Args:
date (str): A date string in the format '%m-%d'
Returns:
A list of tuples containing the daily normals, tmin, tavg, and tmax
"""
sel = [func.min(Measurement.tobs), func.avg(Measurement.tobs), func.max(Measurement.tobs)]
return session.query(*sel).filter(func.strftime("%m-%d", Measurement.date) == date).all()
daily_normals("01-01")
# +
# calculate the daily normals for your trip
# push each tuple of calculations into a list called `normals`
# Set the start and end date of the trip
# Use the start and end date to create a range of dates
# Stip off the year and save a list of %m-%d strings
# Loop through the list of %m-%d strings and calculate the normals for each date
# -
# Load the previous query results into a Pandas DataFrame and add the `trip_dates` range as the `date` index
# Plot the daily normals as an area plot with `stacked=False`
| climate_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tensorflow model - 92% accuracy
#
# #### Adapted from https://www.tensorflow.org/get_started/mnist/beginners
#
# Due to the increibly time consuming and downright depressing outcome of my attempt at bettering the 99% accuracy model provided by tensorflow [here](https://www.tensorflow.org/get_started/mnist/pros) I've decided to scale down the model and use this instead.
# +
#Imports
import tensorflow as tf
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("MNIST_data/", one_hot=True)
# +
#placeholder
x_placeholder = tf.placeholder(tf.float32, [None, 784])
#weights
weights = tf.Variable(tf.zeros([784, 10]))
#biases
biases = tf.Variable(tf.zeros([10]))
# -
y_placeholder = tf.nn.softmax(tf.matmul(x_placeholder, weights) + biases)
y_cep = tf.placeholder(tf.float32, [None, 10])
cross_entropy = tf.reduce_mean(-tf.reduce_sum(y_cep * tf.log(y_placeholder), reduction_indices=[1]))
train_step = tf.train.GradientDescentOptimizer(0.5).minimize(cross_entropy)
# +
sess = tf.InteractiveSession()
saver = tf.train.Saver()
# save model
tf.global_variables_initializer().run()
# -
for _ in range(20000):
batch_xs, batch_ys = mnist.train.next_batch(100)
sess.run(train_step, feed_dict={x_placeholder: batch_xs, y_cep: batch_ys})
correct_prediction = tf.equal(tf.argmax(y_placeholder,1), tf.argmax(y_cep,1))
accuracy = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
print(sess.run(accuracy, feed_dict={x_placeholder: mnist.test.images, y_cep: mnist.test.labels}))
path = saver.save(sess, "./mnist92/mnist92.ckpt")
print("Model saved at: %s" % path)
| Tensorflow-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
from parcels import FieldSet, Field, ParticleSet, JITParticle, AdvectionRK4, ErrorCode, Variable
import cartopy
from glob import glob
import matplotlib.pyplot as plt
from matplotlib import colors, cm
import numpy as np
import xarray as xr
from netCDF4 import Dataset
import math as math
import matplotlib.animation as animation
from collections import Counter
sys.path.insert(1, '../../functions/')
from ParticlePlotFunctions import *
import pandas as pd
# %matplotlib inline
def variability_beaching(namefile,fieldtype='beaching'):
"""This function returns the number of particles beaching at each island per month"""
Traj = ReadTrajectories(namefile)
total_particles = Traj['lon'].shape[0]
ptime= Traj['time'][:]
island = Traj['island'][:]
ptime[isnat(ptime)]=Traj['time'][0,0]
simtimes = np.arange(np.min(ptime), np.max(ptime), dtype='datetime64[6h]').astype('datetime64[ns]')
arrivals = np.zeros((11,simtimes.shape[0]))
if fieldtype == 'beaching':
beached = Traj['beached'][:]
for p in range(total_particles):
if beached[p,-1]==1:
beaching_index = np.where(beached[p,:]==1)[0][0]
time_index = np.where(simtimes == ptime[p,beaching_index].astype('datetime64[6h]').astype('datetime64[ns]'))[0]
island_beached = int(island[p,beaching_index])
arrivals[0,time_index] += 1
arrivals[island_beached,time_index] += 1
else:
distance = Traj['distance'][:]
release_loc = 675 #number of release locations
distance_from_coast = 4 #in km, threshold for beaching
for p in range(total_particles):
index_loc = p%release_loc
beached = np.where((distance[p,:] < distance_from_coast) & (distance[p,:] != 0))[0]
if beached.any():
beaching_index = beached[0]
time_index = np.where(simtimes == ptime[p,beaching_index].astype('datetime64[6h]').astype('datetime64[ns]'))[0]
island_beached = int(island[p,beaching_index])
arrivals[0,time_index] += 1
arrivals[island_beached,time_index] += 1
data = {'Time': simtimes,
'Number of particles beaching in total': arrivals[0,:],
'Number of particles beaching Espanola': arrivals[1,:],
'Number of particles beaching Floreana': arrivals[2,:],
'Number of particles beaching Isabela': arrivals[3,:],
'Number of particles beaching San Cristobal': arrivals[4,:],
'Number of particles beaching Santa Fe': arrivals[5,:],
'Number of particles beaching Santa Cruz': arrivals[6,:],
'Number of particles beaching Fernandina': arrivals[7,:],
'Number of particles beaching Santiago': arrivals[8,:],
'Number of particles beaching Marchena': arrivals[9,:],
'Number of particles beaching Pinta': arrivals[10,:],}
timeseries = pd.DataFrame(data, columns = ['Time',
'Number of particles beaching in total',
'Number of particles beaching Espanola',
'Number of particles beaching Floreana',
'Number of particles beaching Isabela',
'Number of particles beaching San Cristobal',
'Number of particles beaching Santa Fe',
'Number of particles beaching Santa Cruz',
'Number of particles beaching Fernandina',
'Number of particles beaching Santiago',
'Number of particles beaching Marchena',
'Number of particles beaching Pinta'])
timeseries.Time = pd.to_datetime(timeseries.Time)
beaching_monthly = timeseries.resample('M', on='Time').sum()
return beaching_monthly
# +
# FIGURE: Beaching variability
filepath = '../../input/particles/Beaching_200826.nc'
onlyadvection_beaching = variability_beaching(filepath,fieldtype='other')
filepath = '../../input/particles/Beaching_200826_wstokes.nc'
stokes_beaching = variability_beaching(filepath)
filepath = '../../input/particles/Beaching_200826_wind0010.nc'
wind_beaching = variability_beaching(filepath)
filepath = '../../input/particles/Beaching_200826_wstokes_wind0010.nc'
all_beaching = variability_beaching(filepath)
figsize=(16,10)
fig, axs = plt.subplots(2, 1, figsize=figsize, sharey = False, sharex = True)
axs = axs.ravel()
axs[0].plot(onlyadvection_beaching["Number of particles beaching in total"],'-k',label='only advection')
axs[0].plot(stokes_beaching["Number of particles beaching in total"],'--k',label='+ Stokes Drift')
axs[0].plot(wind_beaching["Number of particles beaching in total"],'-.k',label='+ Wind')
axs[0].plot(all_beaching["Number of particles beaching in total"],':k',label='+ Stokes Drift + Wind')
axs[0].legend()
axs[0].set_ylim([0, 2500])
axs[0].set_ylabel('Number of particles')
axs[0].set_title('beaching variability')
axs[1].plot(wind_beaching["Number of particles beaching Espanola"],label='Espanola')
axs[1].plot(wind_beaching["Number of particles beaching Floreana"],label='Floreana')
axs[1].plot(wind_beaching["Number of particles beaching Isabela"],label='Isabela')
axs[1].plot(wind_beaching["Number of particles beaching San Cristobal"],label='San Cristobal')
axs[1].plot(wind_beaching["Number of particles beaching Santa Fe"],label='Santa Fe')
axs[1].plot(wind_beaching["Number of particles beaching Santa Cruz"],label='Santa Cruz')
axs[1].plot(wind_beaching["Number of particles beaching Fernandina"],label='Fernandina')
axs[1].plot(wind_beaching["Number of particles beaching Santiago"],label='Santiago')
axs[1].plot(wind_beaching["Number of particles beaching Marchena"],label='Marchena')
axs[1].plot(wind_beaching["Number of particles beaching Pinta"],label='Pinta')
axs[1].legend()
axs[1].set_ylim([0, 600])
axs[1].set_ylabel('Number of particles')
axs[1].set_xlabel('Time (months)')
axs[1].set_title('beaching variability at each island for + Stokes Drift simulation')
plt.rcParams.update({'font.size': 14})
plt.savefig('variability_arrival.png', dpi = 300)
# -
# ### Figure on beaching variability
#
# There is clearly a seasonal cycle:
# - limited beaching in the months Jul - Aug - Sep (cold/dry season, strong winds and currents)
# - enhanced beaching in the months Mar - Apr - May (warm/wet season, weak winds and currents)
#
# Do we expect to have more beaching with strong winds and currents or less?
#
# (Isabela by far largest island so most particles beach there, second figure could be better interpreted when scaled by coastline-length)
# +
# FIGURE: Pie diagram of how many particles beach in total at each island
labels = 'Espanola', 'Floreana', 'Isabela', 'San Cristobal', 'Santa Fe', 'Santa Cruz', 'Fernandina', 'Santiago', 'Marchena', 'Pinta'
sizes1 = [stokes_beaching["Number of particles beaching Espanola"].sum(),
stokes_beaching["Number of particles beaching Floreana"].sum(),
stokes_beaching["Number of particles beaching Isabela"].sum(),
stokes_beaching["Number of particles beaching San Cristobal"].sum(),
stokes_beaching["Number of particles beaching Santa Fe"].sum(),
stokes_beaching["Number of particles beaching Santa Cruz"].sum(),
stokes_beaching["Number of particles beaching Fernandina"].sum(),
stokes_beaching["Number of particles beaching Santiago"].sum(),
stokes_beaching["Number of particles beaching Marchena"].sum(),
stokes_beaching["Number of particles beaching Pinta"].sum()]
sizes2 = [onlyadvection_beaching["Number of particles beaching Espanola"].sum(),
onlyadvection_beaching["Number of particles beaching Floreana"].sum(),
onlyadvection_beaching["Number of particles beaching Isabela"].sum(),
onlyadvection_beaching["Number of particles beaching San Cristobal"].sum(),
onlyadvection_beaching["Number of particles beaching Santa Fe"].sum(),
onlyadvection_beaching["Number of particles beaching Santa Cruz"].sum(),
onlyadvection_beaching["Number of particles beaching Fernandina"].sum(),
onlyadvection_beaching["Number of particles beaching Santiago"].sum(),
onlyadvection_beaching["Number of particles beaching Marchena"].sum(),
onlyadvection_beaching["Number of particles beaching Pinta"].sum()]
explode = (0, 0, 0.1, 0, 0, 0, 0, 0, 0, 0)
figsize=(10,20)
fig, axs = plt.subplots(2, 1, figsize=figsize)
im = axs[0].pie(sizes1, explode = explode, labels=labels, autopct='%1.1f%%', shadow=False, startangle=0)
text = axs[0].set_title('Percentage of particles beaching on each island - with Stokes Drift')
im2 = axs[1].pie(sizes2, explode = explode, labels=labels, autopct='%1.1f%%', shadow=False, startangle=0)
text = axs[1].set_title('Percentage of particles beaching on each island - only advection')
plt.savefig('arrivalpercentage_per_island.png', dpi = 300)
# -
# 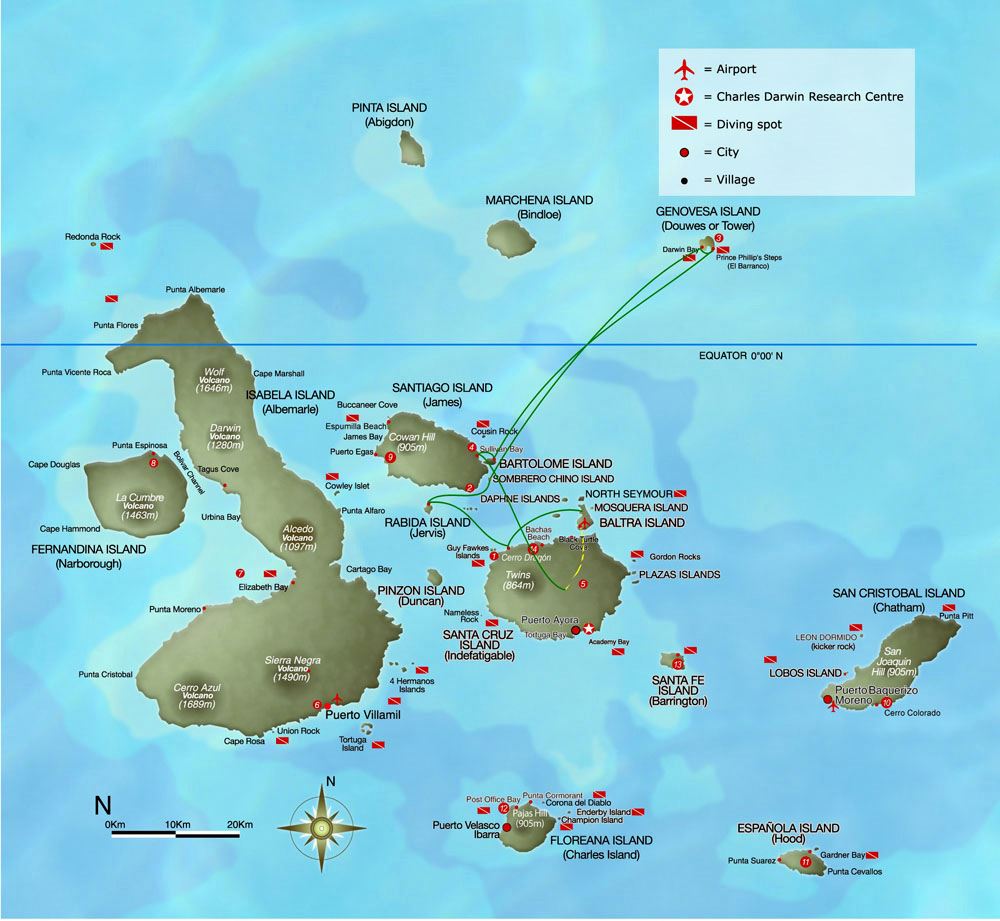
# +
## Make map where particles beach
# need landborder and loop through particles to find closest landborder point and then scatter
| documentation/20.09_BeachingVariability/beaching_variability.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Ch4.2 Histograms, Binnings, and Density
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-white')
data = np.random.randn(1000)
# -
plt.hist(data);
# The ``hist()`` function with options
plt.hist(data, bins=200, density=True, alpha=0.5,
histtype='stepfilled', color='steelblue',
edgecolor='none');
# combination of ``histtype='stepfilled'`` along with some transparency ``alpha``
# +
x1 = np.random.normal(0, 0.8, 1000)
x2 = np.random.normal(-2, 1, 1000)
x3 = np.random.normal(3, 2, 1000)
kwargs = dict(histtype='stepfilled', alpha=0.3, normed=True, bins=40)
plt.hist(x1, **kwargs)
plt.hist(x2, **kwargs)
plt.hist(x3, **kwargs);
# -
# simply compute the histogram
counts, bin_edges = np.histogram(data, bins=5)
print(counts)
counts, bin_edges = np.histogram(data, bins=10)
print(counts)
# ## Two-Dimensional Histograms and Binnings
mean = [0, 0]
cov = [[1, 1], [1, 2]]
x, y = np.random.multivariate_normal(mean, cov, 10000).T
# ### ``plt.hist2d``: Two-dimensional histogram
plt.hist2d(x, y, bins=30, cmap='Blues')
cb = plt.colorbar()
cb.set_label('counts in bin')
# ### ``plt.hexbin``: Hexagonal binnings
plt.hexbin(x, y, gridsize=30, cmap='Blues')
cb = plt.colorbar(label='count in bin')
| III_DataEngineer_BDSE10/1905_Python/TeacherCode/datascience/Ch4.2_Histograms_and_Binnings.ipynb |
# ---
# jupyter:
# jupytext:
# split_at_heading: true
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Command line functions
#
# > Console commands added by the nbdev library
# +
# default_exp cli
# -
# export
from nbdev.imports import *
from nbdev.export import *
from nbdev.sync import *
from nbdev.merge import *
from nbdev.export2html import *
from nbdev.test import *
from fastscript import call_parse, Param
# `nbdev` comes with the following commands. To use any of them, you muse be in one of the subfolder of your project: they will search for the `settings.ini` recursively in the parent directory but need to accessit to be able to work. Their names all begin by nbdev so you can easily get a list with tab completion.
# - `nbdev_build_lib` builds the library from the notebooks
# - `nbdev_update_lib` propagates any change in the library back to the notebooks
# - `nbdev_diff_nbs` gives you the diff between the notebooks and the exported library
# - `nbdev_build_docs` builds the documentation from the notebooks
# - `nbdev_nb2md` to convert a notebook to a markdown file
# - `nbdev_clean_nbs` removes all superfluous metadata form the notebooks, to avoid merge conflicts
# - `nbdev_read_nbs` read all notebooks to make sure none are broken
# - `nbdev_trust_nbs` trust all notebooks (so that the HTML content is shown)
# - `nbdev_fix_merge` will fix merge conflicts in a notebook file
# - `nbdev_install_git_hooks` install the git hooks that use the last two command automatically on each commit/merge.
# ## Navigating from notebooks to script and back
#export
@call_parse
def nbdev_build_lib(fname:Param("A notebook name or glob to convert", str)=None):
"Export notebooks matching `fname` to python modules"
write_tmpls()
notebook2script(fname=fname)
# By default (`fname` left to `None`), the whole library is built from the notebooks in the `lib_folder` set in your `settings.ini`.
#export
@call_parse
def nbdev_update_lib(fname:Param("A notebook name or glob to convert", str)=None):
"Propagates any change in the modules matching `fname` to the notebooks that created them"
script2notebook(fname=fname)
# By default (`fname` left to `None`), the whole library is treated. Note that this tool is only designed for small changes such as typo or small bug fixes. You can't add new cells in notebook from the library.
#export
@call_parse
def nbdev_diff_nbs():
"Prints the diff between an export of the library in notebooks and the actual modules"
diff_nb_script()
# ## Extracting tests
# export
def _test_one(fname, flags=None, verbose=True):
print(f"testing: {fname}")
start = time.time()
try:
test_nb(fname, flags=flags)
return True,time.time()-start
except Exception as e:
if "Kernel died before replying to kernel_info" in str(e):
time.sleep(random.random())
_test_one(fname, flags=flags)
if verbose: print(f'Error in {fname}:\n{e}')
return False,time.time()-start
# export
@call_parse
def nbdev_test_nbs(fname:Param("A notebook name or glob to convert", str)=None,
flags:Param("Space separated list of flags", str)=None,
n_workers:Param("Number of workers to use", int)=None,
verbose:Param("Print errors along the way", bool)=True,
timing:Param("Timing each notebook to see the ones are slow", bool)=False):
"Test in parallel the notebooks matching `fname`, passing along `flags`"
if flags is not None: flags = flags.split(' ')
if fname is None:
files = [f for f in Config().nbs_path.glob('*.ipynb') if not f.name.startswith('_')]
else: files = glob.glob(fname)
files = [Path(f).absolute() for f in sorted(files)]
if len(files)==1 and n_workers is None: n_workers=0
# make sure we are inside the notebook folder of the project
os.chdir(Config().nbs_path)
results = parallel(_test_one, files, flags=flags, verbose=verbose, n_workers=n_workers)
passed,times = [r[0] for r in results],[r[1] for r in results]
if all(passed): print("All tests are passing!")
else:
msg = "The following notebooks failed:\n"
raise Exception(msg + '\n'.join([f.name for p,f in zip(passed,files) if not p]))
if timing:
for i,t in sorted(enumerate(times), key=lambda o:o[1], reverse=True):
print(f"Notebook {files[i].name} took {int(t)} seconds")
# By default (`fname` left to `None`), the whole library is tested from the notebooks in the `lib_folder` set in your `settings.ini`.
# ## Building documentation
# The following functions complete the ones in `export2html` to fully build the documentation of your library.
#export
import time,random,warnings
#export
def _leaf(k,v):
url = 'external_url' if "http" in v else 'url'
#if url=='url': v=v+'.html'
return {'title':k, url:v, 'output':'web,pdf'}
#export
_k_names = ['folders', 'folderitems', 'subfolders', 'subfolderitems']
def _side_dict(title, data, level=0):
k_name = _k_names[level]
level += 1
res = [(_side_dict(k, v, level) if isinstance(v,dict) else _leaf(k,v))
for k,v in data.items()]
return ({k_name:res} if not title
else res if title.startswith('empty')
else {'title': title, 'output':'web', k_name: res})
#export
_re_catch_title = re.compile('^title\s*:\s*(\S+.*)$', re.MULTILINE)
#export
def _get_title(fname):
"Grabs the title of html file `fname`"
with open(fname, 'r') as f: code = f.read()
src = _re_catch_title.search(code)
return fname.stem if src is None else src.groups()[0]
#hide
test_eq(_get_title(Config().doc_path/'export.html'), "Export to modules")
#export
from nbdev.export2html import _nb2htmlfname
#export
def create_default_sidebar():
"Create the default sidebar for the docs website"
dic = {"Overview": "/"}
files = [f for f in Config().nbs_path.glob('*.ipynb') if not f.name.startswith('_')]
fnames = [_nb2htmlfname(f) for f in sorted(files)]
dic.update({_get_title(f):f'/{f.stem}' for f in fnames if f.stem!='index'})
dic = {Config().lib_name: dic}
json.dump(dic, open(Config().doc_path/'sidebar.json', 'w'), indent=2)
# The default sidebar lists all html pages with their respective title, except the index that is named "Overview". To build a custom sidebar, set the flag `custom_sidebar` in your `settings.ini` to `True` then change the `sidebar.json` file in the `doc_folder` to your liking. Otherwise, the sidebar is updated at each doc build.
# +
#hide
#create_default_sidebar()
# -
#export
def make_sidebar():
"Making sidebar for the doc website form the content of `doc_folder/sidebar.json`"
if not (Config().doc_path/'sidebar.json').exists() or Config().custom_sidebar == 'False': create_default_sidebar()
sidebar_d = json.load(open(Config().doc_path/'sidebar.json', 'r'))
res = _side_dict('Sidebar', sidebar_d)
res = {'entries': [res]}
res_s = yaml.dump(res, default_flow_style=False)
res_s = res_s.replace('- subfolders:', ' subfolders:').replace(' - - ', ' - ')
res_s = f"""
#################################################
### THIS FILE WAS AUTOGENERATED! DO NOT EDIT! ###
#################################################
# Instead edit {'../../sidebar.json'}
"""+res_s
open(Config().doc_path/'_data/sidebars/home_sidebar.yml', 'w').write(res_s)
# export
_re_index = re.compile(r'^(?:\d*_|)index\.ipynb$')
# export
def make_readme():
"Convert the index notebook to README.md"
index_fn = None
for f in Config().nbs_path.glob('*.ipynb'):
if _re_index.match(f.name): index_fn = f
assert index_fn is not None, "Could not locate index notebook"
convert_md(index_fn, Config().config_file.parent, jekyll=False)
n = Config().config_file.parent/index_fn.with_suffix('.md').name
shutil.move(n, Config().config_file.parent/'README.md')
# export
@call_parse
def nbdev_build_docs(fname:Param("A notebook name or glob to convert", str)=None,
force_all:Param("Rebuild even notebooks that haven't changed", bool)=False,
mk_readme:Param("Also convert the index notebook to README", bool)=True,
n_workers:Param("Number of workers to use", int)=None):
"Build the documentation by converting notebooks mathing `fname` to html"
notebook2html(fname=fname, force_all=force_all, n_workers=n_workers)
if fname is None: make_sidebar()
if mk_readme: make_readme()
# By default (`fname` left to `None`), the whole documentation is build from the notebooks in the `lib_folder` set in your `settings.ini`, only converting the ones that have been modified since the their corresponding html was last touched unless you pass `force_all=True`. The index is also converted to make the README file, unless you pass along `mk_readme=False`.
# export
@call_parse
def nbdev_nb2md(fname:Param("A notebook file name to convert", str),
dest:Param("The destination folder", str)='.',
jekyll:Param("To use jekyll metadata for your markdown file or not", bool)=True,):
"Convert the notebook in `fname` to a markdown file"
convert_md(fname, dest, jekyll=jekyll)
# ## Other utils
# export
@call_parse
def nbdev_read_nbs(fname:Param("A notebook name or glob to convert", str)=None):
"Check all notebooks matching `fname` can be opened"
files = Config().nbs_path.glob('**/*.ipynb') if fname is None else glob.glob(fname)
for nb in files:
try: _ = read_nb(nb)
except Exception as e:
print(f"{nb} is corrupted and can't be opened.")
raise e
# By default (`fname` left to `None`), the all the notebooks in `lib_folder` are checked.
# export
@call_parse
def nbdev_trust_nbs(fname:Param("A notebook name or glob to convert", str)=None,
force_all:Param("Trust even notebooks that haven't changed", bool)=False):
"Trust noteboks matching `fname`"
check_fname = Config().nbs_path/".last_checked"
last_checked = os.path.getmtime(check_fname) if check_fname.exists() else None
files = Config().nbs_path.glob('**/*.ipynb') if fname is None else glob.glob(fname)
for fn in files:
if last_checked and not force_all:
last_changed = os.path.getmtime(fn)
if last_changed < last_checked: continue
nb = read_nb(fn)
if not NotebookNotary().check_signature(nb): NotebookNotary().sign(nb)
check_fname.touch(exist_ok=True)
# By default (`fname` left to `None`), the all the notebooks in `lib_folder` are trusted. To speed things up, only the ones touched since the last time this command was run are trusted unless you pass along `force_all=True`.
# export
@call_parse
def nbdev_fix_merge(fname:Param("A notebook filename to fix", str),
fast:Param("Fast fix: automatically fix the merge conflicts in outputs or metadata", bool)=True,
trust_us:Param("Use local outputs/metadata when fast mergning", bool)=True):
"Fix merge conflicts in notebook `fname`"
fix_conflicts(fname, fast=fast, trust_us=trust_us)
# When you have merge conflicts after a `git pull`, the notebook file will be broken and won't open in jupyter notebook anymore. This command fixes this by changing the notebook to a proper json file again and add markdown cells to signal the conflict, you just have to open that notebook again and look for `>>>>>>>` to see those conflicts and manually fix them. The old broken file is copied with a `.ipynb.bak` extension, so is still accessible in case the merge wasn't sucessful.
#
# Moreover, if `fast=True`, conflicts in outputs and metadata will automatically be fixed by using the local version if `trust_us=True`, the remote one if `trust_us=False`. With this option, it's very likely you won't have anything to do, unless there is a real conflict.
#export
def bump_version(version, part=2):
version = version.split('.')
version[part] = str(int(version[part]) + 1)
for i in range(part+1, 3): version[i] = '0'
return '.'.join(version)
test_eq(bump_version('0.1.1' ), '0.1.2')
test_eq(bump_version('0.1.1', 1), '0.2.0')
# export
@call_parse
def nbdev_bump_version(part:Param("Part of version to bump", int)=2):
"Increment version in `settings.py` by one"
cfg = Config()
print(f'Old version: {cfg.version}')
cfg.d['version'] = bump_version(Config().version, part)
cfg.save()
update_version()
print(f'New version: {cfg.version}')
# ## Git hooks
# export
import subprocess
# export
@call_parse
def nbdev_install_git_hooks():
"Install git hooks to clean/trust notebooks automatically"
path = Config().config_file.parent
fn = path/'.git'/'hooks'/'post-merge'
#Trust notebooks after merge
with open(fn, 'w') as f:
f.write("""#!/bin/bash
echo "Trusting notebooks"
nbdev_trust_nbs
"""
)
os.chmod(fn, os.stat(fn).st_mode | stat.S_IEXEC)
#Clean notebooks on commit/diff
with open(path/'.gitconfig', 'w') as f:
f.write("""# Generated by nbdev_install_git_hooks
#
# If you need to disable this instrumentation do:
#
# git config --local --unset include.path
#
# To restore the filter
#
# git config --local include.path .gitconfig
#
# If you see notebooks not stripped, checked the filters are applied in .gitattributes
#
[filter "clean-nbs"]
clean = nbdev_clean_nbs --read_input_stream True
smudge = cat
required = true
[diff "ipynb"]
textconv = nbdev_clean_nbs --disp True --fname
""")
cmd = "git config --local include.path ../.gitconfig"
print(f"Executing: {cmd}")
result = subprocess.run(cmd.split(), shell=False, check=False, stderr=subprocess.PIPE)
if result.returncode == 0:
print("Success: hooks are installed and repo's .gitconfig is now trusted")
else:
print("Failed to trust repo's .gitconfig")
if result.stderr: print(f"Error: {result.stderr.decode('utf-8')}")
with open(Config().nbs_path/'.gitattributes', 'w') as f:
f.write("""**/*.ipynb filter=clean-nbs
**/*.ipynb diff=ipynb
"""
)
# This command install git hooks to make sure notebooks are cleaned before you commit them to GitHub and automatically trusted at each merge. To be more specific, this creates:
# - an executable '.git/hooks/post-merge' file that contains the command `nbdev_trust_nbs`
# - a `.gitconfig` file that uses `nbev_clean_nbs` has a filter/diff on all notebook files inside `nbs_folder` and a `.gitattributes` file generated in this folder (copy this file in other folders where you might have notebooks you want cleaned as well)
# ## Export -
#hide
from nbdev.export import *
notebook2script()
| nbs/06_cli.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Time: O(logn)
# Space: O(1)
def find_max_value(arr):
n = len(arr)
low, high = 0, n - 1
while low <= high:
mid = (low + high) // 2
if (not mid or mid == n - 1) or arr[mid - 1] < arr[mid] > arr[mid + 1]:
return arr[mid]
if arr[mid - 1] < arr[mid] < arr[mid + 1]:
low = mid + 1
else:
high = mid - 1
if __name__=='__main__':
test_cases = [[3, 5,15, 50, 11, 10, 8, 6],
[10, 20, 30, 40, 50],
[8, 10, 20, 80, 100, 200, 400, 500, 3, 2, 1],
[120, 100, 80, 20, 0]]
for tc in test_cases:
print(find_max_value(tc))
# -
| assignments/array/Maximum Value in an array of Increasing and Decreasing using Binary Search.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Environment (conda_tensorflow_p36)
# language: python
# name: conda_tensorflow_p36
# ---
# ### Read sheet 2
# +
import pandas as pd
import numpy as np
# Define cols to fetch
cols = ["Fake Applicant ID", "Age (Birthday Masked)", "Income", "Promise Zone?", "Primary Interest In Course", "Education", "Hours Coded", "How Many Hours A Week Can You Commit To Class", "Hacker Rank Score", "Completed?"]
# Get app data
df_sheet2_filt = pd.read_csv("https://s3-eu-west-1.amazonaws.com/ai-hack-q3-orion/datasets/app_demo_data.csv",
skip_blank_lines=True,
index_col=0,
na_values='No Data',
usecols=cols
)
df_sheet2_filt.head()
# -
# Define cols to fetch
cols = ["Fake Applicant ID", "Attendance", "Course Id"]
# Get completion data
df_sheet1 = pd.read_csv("https://s3-eu-west-1.amazonaws.com/ai-hack-q3-orion/datasets/completion_data.csv",
skip_blank_lines=True,
index_col=1,
na_values='No Data',
usecols=cols,
dtype = {"Course Id" : "object", "Fake Applicant ID" : "int32", "Attendance" : "str"}
)
df_sheet1_filt = df_sheet1.dropna(axis=0)
df_sheet1_filt.head()
total_number_of_classes = df_sheet1_filt.groupby('Fake Applicant ID')['Attendance'].count()
total_number_of_classes.head()
series_attended = df_sheet1_filt.groupby(['Fake Applicant ID'])['Attendance'].apply(lambda x: x[x.str.contains('Attended')].count())
series_attended.head()
attendance_percentage = series_attended.div(total_number_of_classes)
# +
#pd.concat([df_sheet1_filt, attendance_percentage], axis=1)
test_dataframe = pd.merge(df_sheet1_filt, attendance_percentage,how='inner', on='Fake Applicant ID')
test_dataframe_temp = test_dataframe.drop_duplicates(subset=['Course Id'])
test_dataframe_temp.head(25)
# -
| kevin/init_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Boolean Expressions
#
# Boolean expressions are those that evaluate to either `True` or `False`. Some simple examples:
5 < 6
7 < 6
6 < 6
6 <= 6
# ### Relational Operators
#
# These include `<`, `<=`, `>`, `>=`, and `==`. (`==` means equal to, while `=` means assignment to a variable.) They ask whether a *relation* between two objects holds or not. So, as we see above, 6 is *not* less than 6 (the expressions yields `False`) but 6 *is* less than or equal to 6 (the expressions yields `True`). These operators can be *chained*:
n = 4
3 < n < 5 < 6
# #### What is equality?
#
# Two objects can be equal by being the *same*, or by having the *same value*. My friend Joan and I might have equal incomes (same value) but we are not the *same person*. On the other hand, **George Washington** and **the first president of the United States** *are* the same person, named in two different ways.
#
# Python has a ways to ask each question. `==` asks whether two things have the same value, while `is` asks if they are the same object. So, let's ask do the "boxes" `a` and `b` hold the same value:
a = 4.5
b = 4.5
a == b
# Now let's ask if `a` is *the same box* as `b`:
a is b
# But if we make `c` another label on the box labelled `a`, then `is` returns `True`:
c = a
a is c
# Why did we use `float` numbers above, and not `int`?
a = 4
b = 4
c = a
a == b
d = 4
a is b
a is d
a = None
a is None
# ### Boolean Operators
# We will be using the *boolean operators*:
#
# - `and`
# - `or`
# - `not`
# For an `and` expression to be `True`, both the expression to its right and the one to its left must be `True`:
#
3 < 4 and 5 < 6
3 < 4 and 6 < 5
# For an `or` expression to be `True`, either the expression to its right or the one to its left must be `True`:
3 < 4 or 6 < 5
1 > 0 or 2 > 1
1 < 0 or 2 < 1
# `not` simply reverses the boolean value to which it is applied:
6 < 5
not 6 < 5
# What is the value of this boolean expression?
not (((6 < 7) and (4 < 3)) or (7 >= 7))
# a) True
#
# b) False
# What is the value of the boolean expression on the 3rd line below?
x = 7
y = 6
(x < y) or (x < 10) and (y > 3)
# a) True
#
# b) False
# Finally, let us update our order of precedence to include the boolean operators:
#
# 1. parentheses: `()`
# 2. exponentiation: `**`
# 3. negation: `-`
# 4. multiplication, both divisions, modulus: `*`, `/`, `//`, `%`
# 5. addition and subtraction: `+`, `-`
# 6. comparison operators (`==`, `!=`, `>`, `>=`, `<`, `<=`)
# 7. `not`
# 8. `and`
# 9. `or`
# 10. position (left to right among equal precedence operators)
#
# So what is the value of this boolean expression?
5 + 3 * 2 < 12 and 4**2 < 16 or 3 + 4 > 6
# a) True
#
# b) False
| notebooks/BooleanExpr.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 作業 : (Kaggle)鐵達尼生存預測
# https://www.kaggle.com/c/titanic
# # 作業1
# * 參考範例,將鐵達尼的艙位代碼( 'Cabin' )欄位使用特徵雜湊 / 標籤編碼 / 目標均值編碼三種轉換後,
# 與其他數值型欄位一起預估生存機率
# +
# 做完特徵工程前的所有準備 (與前範例相同)
import pandas as pd
import numpy as np
import copy, time
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import cross_val_score
from sklearn.linear_model import LogisticRegression
data_path = 'data/'
df_train = pd.read_csv(data_path + 'titanic_train.csv')
df_test = pd.read_csv(data_path + 'titanic_test.csv')
train_Y = df_train['Survived']
ids = df_test['PassengerId']
df_train = df_train.drop(['PassengerId', 'Survived'] , axis=1)
df_test = df_test.drop(['PassengerId'] , axis=1)
df = pd.concat([df_train,df_test])
df.head()
# +
#只取類別值 (object) 型欄位, 存於 object_features 中
object_features = []
for dtype, feature in zip(df.dtypes, df.columns):
if dtype == 'object':
object_features.append(feature)
print(f'{len(object_features)} Numeric Features : {object_features}\n')
# 只留類別型欄位
df = df[object_features]
df = df.fillna('None')
train_num = train_Y.shape[0]
df.head()
# -
# # 作業2
# * 承上題,三者比較效果何者最好?ans:計數編碼
# 對照組 : 標籤編碼 + 邏輯斯迴歸
df_temp = pd.DataFrame()
for c in df.columns:
df_temp[c] = LabelEncoder().fit_transform(df[c])
train_X = df_temp[:train_num]
estimator = LogisticRegression()
print(cross_val_score(estimator, train_X, train_Y, cv=5).mean())
df_temp.head()
# 加上 'Cabin' 欄位的計數編碼
count_df = df.groupby(['Cabin'])['Name'].agg({'Cabin_Count':'size'}).reset_index()
df = pd.merge(df, count_df, on=['Cabin'], how='left')
count_df.sort_values(by=['Cabin_Count'], ascending=False).head(10)
# 'Cabin'計數編碼 + 邏輯斯迴歸
df_temp = pd.DataFrame()
for c in object_features:
df_temp[c] = LabelEncoder().fit_transform(df[c])
df_temp['Cabin_Count'] = df['Cabin_Count']
train_X = df_temp[:train_num]
estimator = LogisticRegression()
print(cross_val_score(estimator, train_X, train_Y, cv=5).mean())
df_temp.head()
# 'Cabin'特徵雜湊 + 邏輯斯迴歸
df_temp = pd.DataFrame()
for c in object_features:
df_temp[c] = LabelEncoder().fit_transform(df[c])
df_temp['Cabin_Hash'] = df['Cabin'].map(lambda x:hash(x) % 10)
train_X = df_temp[:train_num]
estimator = LogisticRegression()
print(cross_val_score(estimator, train_X, train_Y, cv=5).mean())
df_temp.head()
# 'Cabin'計數編碼 + 'Cabin'特徵雜湊 + 邏輯斯迴歸
df_temp = pd.DataFrame()
for c in object_features:
df_temp[c] = LabelEncoder().fit_transform(df[c])
df_temp['Cabin_Hash'] = df['Cabin'].map(lambda x:hash(x) % 10)
df_temp['Cabin_Count'] = df['Cabin_Count']
train_X = df_temp[:train_num]
estimator = LogisticRegression()
print(cross_val_score(estimator, train_X, train_Y, cv=5).mean())
df_temp.head()
| Day_024_CountEncoderandFeatureHash.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import gpflow
from vbpp.model import VBPP
from data_up_events_training import make_estimate_data_for_up
def build_data(user, start_line, end_line, filename):
#events_oral = make_estimate_data(user,start_line,end_line,filename)
#events = np.unique(events_oral.flatten())
events = make_estimate_data_for_up(user, start_line, end_line, filename)
num_observations = len(events)
#print(num_observations)
return events, num_observations
def domain_grid(domain, num_points): #域grid
return np.linspace(domain.min(axis=1), domain.max(axis=1), num_points)
def domain_area(domain): #域面积
return np.prod(domain.max(axis=1) - domain.min(axis=1))
def build_model(events, domain, num_observations, M=20, variance = 1.0, lengthscales = 0.5 ):
#kernel = gpflow.kernels.SquaredExponential()
kernel = gpflow.kernels.SquaredExponential(variance = variance, lengthscales = lengthscales)
Z = domain_grid(domain, M) #均匀切分domain,和events无关
feature = gpflow.inducing_variables.InducingPoints(Z) #inducing point(将均匀切分的点作为inducing point)
q_mu = np.zeros(M) #均值为0?
q_S = np.eye(M) #单位矩阵
#print (events)
num_events = len(events)
beta0 = np.sqrt(num_events / domain_area(domain)) # 事件数/域面积 的开方,是什么? 是第二个模型的offset
model = VBPP(feature, kernel, domain, q_mu, q_S, beta0=beta0, num_events=num_events, num_observations = num_observations)
return model
# +
N = 100 #预测点(lambda)
#目标用户信息,用于模型训练
object_user = 's052'
object_data_str = 0
object_data_end = 50
#kernel param
variance = 1.9
lengthscales = 0.53
inducing_num = 4
filename = "./data/DSL-StrongPasswordData.xls"
events,num_observations = build_data(object_user,object_data_str, object_data_end,filename)
events = np.array(events, float).reshape(-1, 1)
domain_max = max(events) + 0.03
domain = [0,domain_max]
domain = np.array(domain, float).reshape(1, 2)
model = build_model(events, domain, num_observations, M=inducing_num, variance=variance, lengthscales = lengthscales) #M是 inducing point的数量
def objective_closure(): #目标函数
return - model.elbo(events)
gpflow.optimizers.Scipy().minimize(objective_closure, model.trainable_variables)
# +
#画强度的估值图
X = domain_grid(domain, N)
lambda_mean, lower, upper = model.predict_lambda_and_percentiles(X)
lower = lower.numpy().flatten()
upper = upper.numpy().flatten()
title = object_user+'-'+str(object_data_str)+'_'+str(object_data_end)+'-M'+ str(inducing_num) + '-var'+ str(variance) + '-len' + str(lengthscales)
plt.title(title)
plt.xlim(X.min(), X.max())
#plt.ylim(0, 30)
plt.plot(X, lambda_mean)
plt.fill_between(X.flatten(), lower, upper, alpha=0.3)
plt.plot(events, np.zeros_like(events), '|')
plt.show()
# +
#目标用户信息,用于模型训练
object_data_str = 50
object_data_end = 100
'''
#kernel param
variance = 1.9
lengthscales = 0.52
inducing_num = 4
'''
events,num_observations = build_data(object_user,object_data_str, object_data_end,filename)
events = np.array(events, float).reshape(-1, 1)
domain_max = max(events) + 0.03
domain = [0,domain_max]
domain = np.array(domain, float).reshape(1, 2)
model = build_model(events, domain, num_observations, M=inducing_num, variance=variance, lengthscales = lengthscales) #M是 inducing point的数量
def objective_closure(): #目标函数
return - model.elbo(events)
gpflow.optimizers.Scipy().minimize(objective_closure, model.trainable_variables)
# +
#画强度的估值图
X = domain_grid(domain, N)
lambda_mean, lower, upper = model.predict_lambda_and_percentiles(X)
lower = lower.numpy().flatten()
upper = upper.numpy().flatten()
title = object_user+'-'+str(object_data_str)+'_'+str(object_data_end)+'-M'+ str(inducing_num) + '-var'+ str(variance) + '-len' + str(lengthscales)
plt.title(title)
plt.xlim(X.min(), X.max())
#plt.ylim(0, 30)
plt.plot(X, lambda_mean)
plt.fill_between(X.flatten(), lower, upper, alpha=0.3)
plt.plot(events, np.zeros_like(events), '|')
plt.show()
# +
#目标用户信息,用于模型训练
object_data_str = 100
object_data_end = 150
#kernel param
variance = 1.9
lengthscales = 0.53
inducing_num = 4
events,num_observations = build_data(object_user,object_data_str, object_data_end,filename)
events = np.array(events, float).reshape(-1, 1)
domain_max = max(events) + 0.03
domain = [0,domain_max]
domain = np.array(domain, float).reshape(1, 2)
model = build_model(events, domain, num_observations, M=inducing_num, variance=variance, lengthscales = lengthscales) #M是 inducing point的数量
def objective_closure(): #目标函数
return - model.elbo(events)
gpflow.optimizers.Scipy().minimize(objective_closure, model.trainable_variables)
# +
#画强度的估值图
X = domain_grid(domain, N)
lambda_mean, lower, upper = model.predict_lambda_and_percentiles(X)
lower = lower.numpy().flatten()
upper = upper.numpy().flatten()
title = object_user+'-'+str(object_data_str)+'_'+str(object_data_end)+'-M'+ str(inducing_num) + '-var'+ str(variance) + '-len' + str(lengthscales)
plt.title(title)
plt.xlim(X.min(), X.max())
#plt.ylim(0, 30)
plt.plot(X, lambda_mean)
plt.fill_between(X.flatten(), lower, upper, alpha=0.3)
plt.plot(events, np.zeros_like(events), '|')
plt.show()
# +
#目标用户信息,用于模型训练
object_data_str = 150
object_data_end = 200
'''
#kernel param
variance = 1.0
lengthscales = 0.5
inducing_num = 3
'''
events,num_observations = build_data(object_user,object_data_str, object_data_end,filename)
events = np.array(events, float).reshape(-1, 1)
domain_max = max(events) + 0.03
domain = [0,domain_max]
domain = np.array(domain, float).reshape(1, 2)
model = build_model(events, domain, num_observations, M=inducing_num, variance=variance, lengthscales = lengthscales) #M是 inducing point的数量
def objective_closure(): #目标函数
return - model.elbo(events)
gpflow.optimizers.Scipy().minimize(objective_closure, model.trainable_variables)
# +
#画强度的估值图
X = domain_grid(domain, N)
lambda_mean, lower, upper = model.predict_lambda_and_percentiles(X)
lower = lower.numpy().flatten()
upper = upper.numpy().flatten()
title = object_user+'-'+str(object_data_str)+'_'+str(object_data_end)+'-M'+ str(inducing_num) + '-var'+ str(variance) + '-len' + str(lengthscales)
plt.title(title)
plt.xlim(X.min(), X.max())
plt.ylim(0, 30)
plt.plot(X, lambda_mean)
plt.fill_between(X.flatten(), lower, upper, alpha=0.3)
plt.plot(events, np.zeros_like(events), '|')
plt.show()
# +
#目标用户信息,用于模型训练
object_data_str = 200
object_data_end = 250
'''
#kernel param
variance = 1.6
lengthscales = 0.5
inducing_num =3
'''
events,num_observations = build_data(object_user,object_data_str, object_data_end,filename)
events = np.array(events, float).reshape(-1, 1)
domain_max = max(events) + 0.03
domain = [0,domain_max]
domain = np.array(domain, float).reshape(1, 2)
model = build_model(events, domain, num_observations, M=inducing_num, variance=variance, lengthscales = lengthscales) #M是 inducing point的数量
def objective_closure(): #目标函数
return - model.elbo(events)
gpflow.optimizers.Scipy().minimize(objective_closure, model.trainable_variables)
# +
#画强度的估值图
X = domain_grid(domain, N)
lambda_mean, lower, upper = model.predict_lambda_and_percentiles(X)
lower = lower.numpy().flatten()
upper = upper.numpy().flatten()
title = object_user+'-'+str(object_data_str)+'_'+str(object_data_end)+'-M'+ str(inducing_num) + '-var'+ str(variance) + '-len' + str(lengthscales)
plt.title(title)
plt.xlim(X.min(), X.max())
#plt.ylim(0, 30)
plt.plot(X, lambda_mean)
plt.fill_between(X.flatten(), lower, upper, alpha=0.3)
plt.plot(events, np.zeros_like(events), '|')
plt.show()
# +
#目标用户信息,用于模型训练
object_data_str = 250
object_data_end = 300
#kernel param
variance = 1.9
lengthscales = 0.51
inducing_num = 4
events,num_observations = build_data(object_user,object_data_str, object_data_end,filename)
events = np.array(events, float).reshape(-1, 1)
domain_max = max(events) + 0.03
domain = [0,domain_max]
domain = np.array(domain, float).reshape(1, 2)
model = build_model(events, domain, num_observations, M=inducing_num, variance=variance, lengthscales = lengthscales) #M是 inducing point的数量
def objective_closure(): #目标函数
return - model.elbo(events)
gpflow.optimizers.Scipy().minimize(objective_closure, model.trainable_variables)
# +
#画强度的估值图
X = domain_grid(domain, N)
lambda_mean, lower, upper = model.predict_lambda_and_percentiles(X)
lower = lower.numpy().flatten()
upper = upper.numpy().flatten()
title = object_user+'-'+str(object_data_str)+'_'+str(object_data_end)+'-M'+ str(inducing_num) + '-var'+ str(variance) + '-len' + str(lengthscales)
plt.title(title)
plt.xlim(X.min(), X.max())
#plt.ylim(0, 30)
plt.plot(X, lambda_mean)
plt.fill_between(X.flatten(), lower, upper, alpha=0.3)
plt.plot(events, np.zeros_like(events), '|')
plt.show()
# +
#目标用户信息,用于模型训练
object_data_str = 300
object_data_end = 350
#kernel param
variance = 1.9
lengthscales = 0.53
inducing_num = 4
events,num_observations = build_data(object_user,object_data_str, object_data_end,filename)
events = np.array(events, float).reshape(-1, 1)
domain_max = max(events) + 0.03
domain = [0,domain_max]
domain = np.array(domain, float).reshape(1, 2)
model = build_model(events, domain, num_observations, M=inducing_num, variance=variance, lengthscales = lengthscales) #M是 inducing point的数量
def objective_closure(): #目标函数
return - model.elbo(events)
gpflow.optimizers.Scipy().minimize(objective_closure, model.trainable_variables)
# +
#画强度的估值图
X = domain_grid(domain, N)
lambda_mean, lower, upper = model.predict_lambda_and_percentiles(X)
lower = lower.numpy().flatten()
upper = upper.numpy().flatten()
title = object_user+'-'+str(object_data_str)+'_'+str(object_data_end)+'-M'+ str(inducing_num) + '-var'+ str(variance) + '-len' + str(lengthscales)
plt.title(title)
plt.xlim(X.min(), X.max())
plt.ylim(0, 30)
plt.plot(X, lambda_mean)
plt.fill_between(X.flatten(), lower, upper, alpha=0.3)
plt.plot(events, np.zeros_like(events), '|')
plt.show()
# +
#目标用户信息,用于模型训练
object_data_str = 350
object_data_end = 400
#kernel param
variance = 1.9
lengthscales = 0.53
inducing_num = 4
events,num_observations = build_data(object_user,object_data_str, object_data_end,filename)
events = np.array(events, float).reshape(-1, 1)
domain_max = max(events) + 0.03
domain = [0,domain_max]
domain = np.array(domain, float).reshape(1, 2)
model = build_model(events, domain, num_observations, M=inducing_num, variance=variance, lengthscales = lengthscales) #M是 inducing point的数量
def objective_closure(): #目标函数
return - model.elbo(events)
gpflow.optimizers.Scipy().minimize(objective_closure, model.trainable_variables)
# +
#画强度的估值图
X = domain_grid(domain, N)
lambda_mean, lower, upper = model.predict_lambda_and_percentiles(X)
lower = lower.numpy().flatten()
upper = upper.numpy().flatten()
title = object_user+'-'+str(object_data_str)+'_'+str(object_data_end)+'-M'+ str(inducing_num) + '-var'+ str(variance) + '-len' + str(lengthscales)
plt.title(title)
plt.xlim(X.min(), X.max())
plt.ylim(0, 30)
plt.plot(X, lambda_mean)
plt.fill_between(X.flatten(), lower, upper, alpha=0.3)
plt.plot(events, np.zeros_like(events), '|')
plt.show()
# -
| up_event_estimate_lambda.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# PCovR-Inspired Feature Selection
# ==============================
# +
from sklearn.linear_model import RidgeCV
from sklearn.preprocessing import StandardScaler
from matplotlib import pyplot as plt
from matplotlib import cm
from tqdm.notebook import tqdm
import numpy as np
from skcosmo.feature_selection import PCovCUR, PCovFPS, CUR, FPS
from skcosmo.datasets import load_csd_1000r
from skcosmo.preprocessing import StandardFlexibleScaler
cmap = cm.brg
# -
# For this, we will use the provided csd dataset, which has 100 features to select from.
X, y = load_csd_1000r(return_X_y=True)
X = StandardFlexibleScaler(column_wise=False).fit_transform(X)
y = StandardScaler().fit_transform(y.reshape(X.shape[0], -1))
n = X.shape[-1]//2
lr = RidgeCV(cv=2, alphas=np.logspace(-10,1), fit_intercept=False)
# ## Feature Selection with CUR + PCovR
# First, let's demonstrate CUR feature selection, and show the ten features chosen with a mixing parameter of 0.0, 0.5, and 1.0 perform.
# +
for m in np.arange(0, 1.01, 0.5, dtype=np.float32):
if m < 1.0:
idx = PCovCUR(mixing=m, n_to_select=n).fit(X, y).selected_idx_
else:
idx = CUR(n_to_select=n).fit(X, y).selected_idx_
plt.loglog(
range(1, n + 1),
np.array(
[
lr.fit(X[:, idx[: ni + 1]], y).score(X[:, idx[: ni + 1]], y)
for ni in range(n)
]
),
label=m,
c=cmap(m),
marker="o",
)
plt.xlabel("Number of Features Selected")
plt.ylabel(r"$R^2$")
plt.legend(title="Mixing \nParameter")
plt.show()
# -
# ### Non-iterative feature selection with CUR + PCovR
# Computing a non-iterative CUR is more efficient, although can result in poorer performance for larger datasets. you can also use a greater number of eigenvectors to compute the feature importance by varying `k`, but `k` should not exceed the number of targets, for optimal results.
# +
m = 0.0
idx = PCovCUR(mixing=m, n_to_select=n).fit(X, y).selected_idx_
idx_non_it = PCovCUR(mixing=m, iterative=False, n_to_select=n).fit(X, y).selected_idx_
plt.loglog(
range(1, n + 1),
np.array(
[
lr.fit(X[:, idx[: ni + 1]], y).score(X[:, idx[: ni + 1]], y)
for ni in range(n)
]
),
label='Iterative',
marker="o",
)
plt.loglog(
range(1, n + 1),
np.array(
[
lr.fit(X[:, idx_non_it[: ni + 1]], y).score(X[:, idx_non_it[: ni + 1]], y)
for ni in range(n)
]
),
label='Non-Iterative',
marker="s",
)
plt.xlabel("Number of Features Selected")
plt.ylabel(r"$R^2$")
plt.legend()
plt.show()
# -
# ## Feature Selection with FPS + PCovR
# Next, let's look at FPS. We'll choose the first index from CUR at m = 0, which is 46.
# +
for m in np.arange(0, 1.01, 0.5, dtype=np.float32):
if m < 1.0:
idx = PCovFPS(mixing=m, n_to_select=n, initialize=46).fit(X, y).selected_idx_
else:
idx = FPS(n_to_select=n, initialize=46).fit(X, y).selected_idx_
plt.loglog(
range(1, n + 1),
np.array(
[
lr.fit(X[:, idx[: ni + 1]], y).score(X[:, idx[: ni + 1]], y)
for ni in range(n)
]
),
label=m,
c=cmap(m),
marker="o",
)
plt.xlabel("Number of Features Selected")
plt.ylabel(r"$R^2$")
plt.legend(title="Mixing \nParameter")
plt.show()
# -
| examples/FeatureSelection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Exploring the WMS with OWSLib
# +
from owslib.wms import WebMapService
url = "https://pae-paha.pacioos.hawaii.edu/thredds/wms/dhw_5km?service=WMS"
web_map_services = WebMapService(url)
print("\n".join(web_map_services.contents.keys()))
# -
# ### Layer metadata
# +
layer = "CRW_SST"
wms = web_map_services.contents[layer]
name = wms.title
lon = (wms.boundingBox[0] + wms.boundingBox[2]) / 2.0
lat = (wms.boundingBox[1] + wms.boundingBox[3]) / 2.0
center = lat, lon
time_interval = "{0}/{1}".format(
wms.timepositions[0].strip(), wms.timepositions[-1].strip()
)
style = "boxfill/sst_36"
if style not in wms.styles:
style = None
# -
# ### Single layer
# +
import folium
from folium import plugins
lon, lat = -50, -40
m = folium.Map(location=[lat, lon], zoom_start=5, control_scale=True)
w = folium.raster_layers.WmsTileLayer(
url=url,
name=name,
styles=style,
fmt="image/png",
transparent=True,
layers=layer,
overlay=True,
COLORSCALERANGE="1.2,28",
)
w.add_to(m)
time = plugins.TimestampedWmsTileLayers(w, period="PT1H", time_interval=time_interval)
time.add_to(m)
folium.LayerControl().add_to(m)
m
# -
# ### Multiple layers
# +
m = folium.Map(location=[lat, lon], zoom_start=5, control_scale=True)
w0 = folium.raster_layers.WmsTileLayer(
url=url,
name="sea_surface_temperature",
styles=style,
fmt="image/png",
transparent=True,
layers="CRW_SST",
overlay=True,
)
w1 = folium.raster_layers.WmsTileLayer(
url=url,
name="analysed sea surface temperature anomaly",
styles=style,
fmt="image/png",
transparent=True,
layers="CRW_SSTANOMALY",
overlay=True,
)
w0.add_to(m)
w1.add_to(m)
time = folium.plugins.TimestampedWmsTileLayers(
[w0, w1], period="PT1H", time_interval=time_interval
)
time.add_to(m)
folium.LayerControl().add_to(m)
m
| examples/WmsTimeDimension.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Apriori Algorithm:
#
# The apriori algorithm is an algorithm for mining frequent itemset for boolean association rules.'
#
# Apriori uses a "botton up" approach, where frequent subsets are extended one item at a time which is known as candidate generation, and groups of candidates are tested against the data.
#
# Apriori algorithm is a data mining techniques which helps us mine basket data or data about transaction for association rule.
# Basket has 2 meaning:
# 1.Single transaction: All items brought in one single transaction.
# 2.Items brought by user for short period of time like for 1 or 2 month
#
# When a set of basket is fed into Apriori algorithm it will generate set of rules which tells what kind of products are purchased in same basket. Each of these rules has to satisfy minimum support and confidence.
#
# Support and confidence is matrixes that are defined in a way that says how valid a rule is and how strong the association is. Support tells us how many out of all the transaction contain the items in the rule. Confidence tells us how many of the transactions out of all the transaction which contains those items exhibit these association
#
# #Data Setup
# https://github.com/marshallshen/nyc_restaurants_inspection
# +
import pandas as pd
df = pd.read_csv("tesco_dataset.csv")
#df1 = pd.read_csv("Restaurant-Dataset.csv")
df
# +
# # %load apriori.py
import sys
from itertools import chain, combinations
from collections import defaultdict
from optparse import OptionParser
import numpy as np
import matplotlib.pyplot as plt
from numpy import *
def subsets(arr):
""" Returns non empty subsets of arr"""
return chain(*[combinations(arr, i + 1) for i, a in enumerate(arr)])
def returnItemsWithMinSupport(itemSet, transactionList, minSupport, freqSet):
"""calculates the support for items in the itemSet and returns a subset
of the itemSet each of whose elements satisfies the minimum support"""
_itemSet = set()
localSet = defaultdict(int)
for item in itemSet:
for transaction in transactionList:
if item.issubset(transaction):
freqSet[item] += 1
localSet[item] += 1
for item, count in localSet.items():
support = float(count)/len(transactionList)
if support >= minSupport:
_itemSet.add(item)
return _itemSet
def joinSet(itemSet, length):
"""Join a set with itself and returns the n-element itemsets"""
return set([i.union(j) for i in itemSet for j in itemSet if len(i.union(j)) == length])
def getItemSetTransactionList(data_iterator):
transactionList = list()
itemSet = set()
for record in data_iterator:
transaction = frozenset(record)
transactionList.append(transaction)
for item in transaction:
itemSet.add(frozenset([item])) # Generate 1-itemSets
return itemSet, transactionList
def runApriori(data_iter, minSupport, minConfidence):
"""
run the apriori algorithm. data_iter is a record iterator
Return both:
- items (tuple, support)
- rules ((pretuple, posttuple), confidence)
"""
itemSet, transactionList = getItemSetTransactionList(data_iter)
freqSet = defaultdict(int)
largeSet = dict()
# Global dictionary which stores (key=n-itemSets,value=support)
# which satisfy minSupport
assocRules = dict()
# Dictionary which stores Association Rules
oneCSet = returnItemsWithMinSupport(itemSet,
transactionList,
minSupport,
freqSet)
currentLSet = oneCSet
k = 2
while(currentLSet != set([])):
largeSet[k-1] = currentLSet
currentLSet = joinSet(currentLSet, k)
currentCSet = returnItemsWithMinSupport(currentLSet,
transactionList,
minSupport,
freqSet)
currentLSet = currentCSet
k = k + 1
def getSupport(item):
"""local function which Returns the support of an item"""
return float(freqSet[item])/len(transactionList)
toRetItems = []
for key, value in largeSet.items():
toRetItems.extend([(tuple(item), getSupport(item))
for item in value])
toRetRules = []
for key, value in largeSet.items()[1:]:
for item in value:
_subsets = map(frozenset, [x for x in subsets(item)])
for element in _subsets:
remain = item.difference(element)
if len(remain) > 0:
confidence = getSupport(item)/getSupport(element)
if confidence >= minConfidence:
toRetRules.append(((tuple(element), tuple(remain)),
confidence))
return toRetItems, toRetRules
def printResults(items, rules):
"""prints the generated itemsets sorted by support and the confidence rules sorted by confidence"""
for item, support in sorted(items, key=lambda (item, support): support):
print "item: %s , %.3f" % (str(item), support)
print "\n------------------------ RULES:"
for rule, confidence in sorted(rules, key=lambda (rule, confidence): confidence):
pre, post = rule
print "Rule: %s ==> %s , %.3f" % (str(pre), str(post), confidence)
def dataFromFile(fname):
"""Function which reads from the file and yields a generator"""
file_iter = open(fname, 'rU')
for line in file_iter:
line = line.strip().rstrip(',') # Remove trailing comma
record = frozenset(line.split(','))
yield record
if __name__ == "__main__":
optparser = OptionParser()
optparser.add_option('-f', '--inputFile',
dest='input',
help='filename containing csv',
default=None)
optparser.add_option('-s', '--minSupport',
dest='minS',
help='minimum support value',
default=0.15,
type='float')
optparser.add_option('-c', '--minConfidence',
dest='minC',
help='minimum confidence value',
default=0.6,
type='float')
(options, args) = optparser.parse_args()
inFile = None
if options.input is None:
inFile = sys.stdin
elif options.input is not None:
inFile = dataFromFile(options.input)
else:
print 'No dataset filename specified, system with exit\n'
sys.exit('System will exit')
minSupport = options.minS
minConfidence = options.minC
items, rules = runApriori(inFile, minSupport, minConfidence)
printResults(items, rules)
#Reference
#https://github.com/asaini/Apriori
# +
# #%run apriori.py -f Restaurant-Dataset.csv -s 0.7 0.8
#or
# %run apriori.py -f tesco_dataset.csv -s 0.5 0.6
# +
# Display in graph and plots
import numpy as np
import matplotlib.pyplot as plt
N = rules
names,values = zip(*N)
ind = np.arange(len(N)) # the x locations for the groups
width = 0.20 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, values, width, color='r')
# add some text for labels, title and axes ticks
ax.set_ylabel('Support')
ax.set_xticks(ind+width/20)
ax.set_xticklabels(names)
plt.show()
#Reference
#https://matplotlib.org/examples/api/barchart_demo.html
# -
# The above bar graph shows the rules generated by running the algorithm with the support.
# Bibiography
#
# https://www.analyticsvidhya.com/blog/2014/08/effective-cross-selling-market-basket-analysis/
#
# https://matplotlib.org/examples/api/barchart_demo.html
#
# https://github.com/asaini/Apriori
#
# https://github.com/marshallshen/nyc_restaurants_inspection
| Homeworks/Homework8/Homework8_Apriori.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] toc=true
# <h1>Table of Contents<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#Importing-libraries" data-toc-modified-id="Importing-libraries-1"><span class="toc-item-num">1 </span>Importing libraries</a></span></li><li><span><a href="#Scraping-data-science-jobs-from-Indeed" data-toc-modified-id="Scraping-data-science-jobs-from-Indeed-2"><span class="toc-item-num">2 </span>Scraping data science jobs from Indeed</a></span></li></ul></div>
# -
# ## Importing libraries
# +
import json
import re
import requests
from selenium import webdriver
from selenium.webdriver.chrome.options import Options
from bs4 import BeautifulSoup
import time
from selenium.webdriver.chrome.service import Service
from selenium.webdriver.common.by import By
s=Service('/usr/local/bin/chromedriver')
option = webdriver.ChromeOptions()
option.add_argument("--incognito")
chrome_prefs = {}
chrome_prefs["profile.default_content_settings"] = {"images": 2}
chrome_prefs["profile.managed_default_content_settings"] = {"images": 2}
option.experimental_options["prefs"] = chrome_prefs
read_driver = webdriver.Chrome(service=s,options=option)
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:100% !important; }</style>"))
display(HTML("<style>.output_result { max-width:100% !important; }</style>"))
#display(HTML('<style>.prompt{width: 0px; min-width: 0px; visibility: collapse}</style>'))
import warnings
warnings.filterwarnings('ignore')
pd.options.display.max_rows = 999
# -
# ## Scraping data science jobs from Indeed
URL_BASE='https://www.indeed.com'
cities = ['New+York', 'Chicago', 'San+Francisco', 'Austin', 'Seattle',
'Los+Angeles', 'Philadelphia', 'Atlanta', 'Dallas', 'Pittsburgh',
'Portland', 'Phoenix', 'Denver', 'Miami', 'El+Paso', 'Boston',
'Palo+Alto%2C+CA', 'Tampa%2C+FL', 'Stamford%2C+CT','San+Jose%2C+CA']
jobs=[]
job_ids_dump=[]
for city in cities:
name_of_city=city
keywords='data+scientist'
scrape_url=URL_BASE+'/jobs?q='+keywords+'&l='+name_of_city+'&radius=25'
counter=0
start=0
job_ids_add=[]
job_ids=[]
#Get all job ids for that city
while counter==0 or len(job_ids_add)>0:
read_driver.get(scrape_url+'&start='+str(counter*10))
content=read_driver.page_source
soup=BeautifulSoup(content,'html.parser')
job_ids_add=[item['data-jk'] for item in soup.findAll('a') if 'data-jk' in item.attrs]
job_ids_add=list(set([j for j in job_ids_add if j not in job_ids]))
job_ids+=job_ids_add
job_ids_dump+=job_ids_add
counter+=1
time.sleep(0.1)
print(city.replace('+',' ').replace('%2C',','),len(job_ids))
#Load job descriptions
counter2=0
for job_id in job_ids:
jobdescurl=URL_BASE+'/viewjob?jk='+job_id
read_driver.get(jobdescurl)
content=read_driver.page_source
time.sleep(0.1)
job_soup=BeautifulSoup(content,'html.parser')
job={}
job['url']=jobdescurl
job['company']=job_soup.find('div',{'class':'icl-u-lg-mr--sm icl-u-xs-mr--xs'}).text
job['city']=city.replace('+',' ').replace('%2C',',')
try:
job['company_rating']=job_soup.find('meta',{'itemprop':'ratingValue'})['content']
except:
job['company_rating']=''
try:
job['company_ratingcount']=job_soup.find('meta',{'itemprop':'ratingCount'})['content']
except:
job['company_ratingcount']=''
job['title']=job_soup.find('div',{'class':'jobsearch-JobInfoHeader-title-container'}).text
job['city']='New York'
job['description']=job_soup.find('div',{'id':'jobDescriptionText'}).text
jobs.append(job)
counter2+=1
if counter2%50==0:
print(city,counter2,'/',len(job_ids))
#Save from where left off
import pickle
with open('indeedjobs.pickle', 'wb') as fp:
pickle.dump(jobs, fp)
with open('indeedjobids.pickle', 'wb') as fp:
pickle.dump(job_ids_dump, fp)
jobs=pd.DataFrame(jobs)
jobs.to_pickle('indeed_jobs.pickle')
| notebooks/2.0-MS-IndeedScraper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Assignment 5
# ### Objective
# + active=""
#
# Learn how to connect to social media network (we will use Twitter as example in this assignment), and collect/preprocess/analyze its data
#
#
# Tweets Data Can be used for different purposes by marketing department and data analytics team. For example:
#
# - Compete with rivals to offer price-match gaurantee policy
# - Offer same coupons like rivals in the market place
#
#
# -
# ### Installation and Setup
# + active=""
# For this assignment you need to do the following setup first:
#
# 1. Create an account on twitter.com.
#
# 2. Generate authentication tokens by following the instructions here :
# https://dev.twitter.com/oauth/overview/application-owner-access-tokens
#
# 3. Add your tokens to the credentials.txt file.
#
#
#
# -
# ### Twitter API
# + active=""
# Twitter API
#
# Two APIs:
#
# REST API: Submit HTTP requests to access specific information (tweets, friends, ...)
# Streaming API: Open a continuous connection to Twitter to receive real-time data.
#
# These APIs are HTTP GET request
#
#
#
#
# Here are the twitter API docs that you must familiarize yourself with
#
# https://dev.twitter.com/rest/reference/get/followers/ids
#
# https://dev.twitter.com/overview/api/twitter-libraries
#
#
# + active=""
# When you search in a text (tweets are text messages), often you need to be aware of something STOP WORDS.
# You could read more about stop-words here:
#
# https://en.wikipedia.org/wiki/Stop_words
#
# -
# # Lets create twitter object and use its API. Code snippets below will show you how to use this API
from TwitterAPI import TwitterAPI, TwitterOAuth, TwitterRestPager
o = TwitterOAuth.read_file('credentials.txt')
o.access_token_key
# Using OAuth1...
twitter = TwitterAPI(o.consumer_key,
o.consumer_secret,
o.access_token_key,
o.access_token_secret)
help(twitter)
# What can we do with this twitter object?
# builtin method `dir` tells us...
dir(twitter)
twitter.auth
# Get help on the `request` method using the builtin method called...`help`
help(twitter.request)
# Let's start by querying the search API
response = twitter.request('search/tweets', {'q': 'big+data'})
# What object is returned?
# builtin type method will tell us.
print type(response)
dir(response)
response.json
response.status_code
# See https://dev.twitter.com/overview/api/response-codes
tweets = [r for r in response]
print('found %d tweets' % len(tweets))
type(tweets)
type(tweets[0])
tweets[0]
help(tweets[0])
tweets[0].keys()
tweets[0]['text']
tweets[0]['created_at']
tweets[14]['text']
tweets[0]['user']
user = tweets[0]['user']
print('screen_name=%s, name=%s, location=%s' % (user['screen_name'], user['name'], user['location']))
# +
# Who follows this person?
# https://dev.twitter.com/docs/api/1.1/get/followers/list
screen_name = user['screen_name']
response = twitter.request('followers/list', {'screen_name': screen_name, 'count':200})
followers = [follower for follower in response]
print 'found %d followers for %s' % (len(followers), screen_name)
# See more about paging here: https://dev.twitter.com/docs/working-with-timelines
# -
print followers[0]['screen_name']
# ## Limitations: Can only search 2 weeks in past But can get up to 3,200 most recent tweets of a user Rate limits! https://dev.twitter.com/docs/rate-limiting/1.1/limits e.g., 180 requests in 15 minute window
# # Get BestBuy timeline for the deals screen-name
#
# # This is the screen name for BestBuy_Deals
# Get BestBuyDeals timeline = ''
screen_name = 'BestBuy_Deals'
timeline = [tweet for tweet in twitter.request('statuses/user_timeline',
{'screen_name': screen_name,
'count': 200})]
print 'got %d tweets for user %s' % (len(timeline), screen_name)
# +
# Print time got created.
timeline[3]['created_at']
# -
# Print the text.
print '\n\n\n'.join(t['text'] for t in timeline)
# Count words
from collections import Counter # This is just a fancy dict mapping from object->int, starting at 0.
counts = Counter()
for tweet in timeline:
counts.update(tweet['text'].lower().split())
print('found %d unique terms in %d tweets' % (len(counts), len(timeline)))
counts.most_common(10)
import re
for tweet in timeline:
deal = tweet['text']
print deal + '\n'
# # Find the deals in the BestBuy_Deal tweets that match products in BestDeal MySQL product table
# +
import re
dealMatchGauranteed=[]
for tweet in timeline:
deal = (tweet['text']).encode('ascii','ignore')
if (len(re.findall('TV|XBOX|PS4',deal)) >= 1):
dealMatchGauranteed = dealMatchGauranteed + [deal]
# -
# Sanity Test that we got some deals
dealMatchGauranteed[:2]
# # Create and write the deals into DealMatches.txt file that will be used by web-app of BestDeal to display two deal matches
# +
dealMatchFile = open('DealMatches.txt', 'w')
for deal in dealMatchGauranteed:
dealMatchFile.write("%s\n" % deal)
dealMatchFile.close()
# -
| Assignment5BestBuyDeals.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
import os,sys
import matplotlib.pyplot as plt
import numpy as np
import helper
import simulation
# Generate some random images
input_images, target_masks = simulation.generate_random_data(192, 192, count=3)
for x in [input_images, target_masks]:
print(x.shape)
print(x.min(), x.max())
# Change channel-order and make 3 channels for matplot
input_images_rgb = [x.astype(np.uint8) for x in input_images]
# Map each channel (i.e. class) to each color
target_masks_rgb = [helper.masks_to_colorimg(x) for x in target_masks]
# Left: Input image, Right: Target mask
helper.plot_side_by_side([input_images_rgb, target_masks_rgb])
# +
from torch.utils.data import Dataset, DataLoader
from torchvision import transforms, datasets, models
class SimDataset(Dataset):
def __init__(self, count, transform=None):
self.input_images, self.target_masks = simulation.generate_random_data(192, 192, count=count)
self.transform = transform
def __len__(self):
return len(self.input_images)
def __getitem__(self, idx):
image = self.input_images[idx]
mask = self.target_masks[idx]
if self.transform:
image = self.transform(image)
return [image, mask]
# use same transform for train/val for this example
trans = transforms.Compose([
transforms.ToTensor(),
transforms.Normalize([0.485, 0.456, 0.406], [0.229, 0.224, 0.225]) # imagenet
])
train_set = SimDataset(2000, transform=trans)
val_set = SimDataset(200, transform=trans)
image_datasets = {
'train': train_set, 'val': val_set
}
batch_size = 25
dataloaders = {
'train': DataLoader(train_set, batch_size=batch_size, shuffle=True, num_workers=0),
'val': DataLoader(val_set, batch_size=batch_size, shuffle=True, num_workers=0)
}
dataset_sizes = {
x: len(image_datasets[x]) for x in image_datasets.keys()
}
dataset_sizes
# +
import torchvision.utils
def reverse_transform(inp):
inp = inp.numpy().transpose((1, 2, 0))
mean = np.array([0.485, 0.456, 0.406])
std = np.array([0.229, 0.224, 0.225])
inp = std * inp + mean
inp = np.clip(inp, 0, 1)
inp = (inp * 255).astype(np.uint8)
return inp
# Get a batch of training data
inputs, masks = next(iter(dataloaders['train']))
print(inputs.shape, masks.shape)
for x in [inputs.numpy(), masks.numpy()]:
print(x.min(), x.max(), x.mean(), x.std())
plt.imshow(reverse_transform(inputs[3]))
# +
from torchvision import models
base_model = models.resnet18(pretrained=False)
list(base_model.children())
# +
# check keras-like model summary using torchsummary
import torch
from torchsummary import summary
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
base_model = base_model.to(device)
summary(base_model, input_size=(3, 224, 224))
# +
import torch
import torch.nn as nn
def convrelu(in_channels, out_channels, kernel, padding):
return nn.Sequential(
nn.Conv2d(in_channels, out_channels, kernel, padding=padding),
nn.ReLU(inplace=True),
)
class ResNetUNet(nn.Module):
def __init__(self, n_class):
super().__init__()
self.base_model = models.resnet18(pretrained=True)
self.base_layers = list(base_model.children())
self.layer0 = nn.Sequential(*self.base_layers[:3]) # size=(N, 64, x.H/2, x.W/2)
self.layer0_1x1 = convrelu(64, 64, 1, 0)
self.layer1 = nn.Sequential(*self.base_layers[3:5]) # size=(N, 64, x.H/4, x.W/4)
self.layer1_1x1 = convrelu(64, 64, 1, 0)
self.layer2 = self.base_layers[5] # size=(N, 128, x.H/8, x.W/8)
self.layer2_1x1 = convrelu(128, 128, 1, 0)
self.layer3 = self.base_layers[6] # size=(N, 256, x.H/16, x.W/16)
self.layer3_1x1 = convrelu(256, 256, 1, 0)
self.layer4 = self.base_layers[7] # size=(N, 512, x.H/32, x.W/32)
self.layer4_1x1 = convrelu(512, 512, 1, 0)
self.upsample = nn.Upsample(scale_factor=2, mode='bilinear', align_corners=True)
self.conv_up3 = convrelu(256 + 512, 512, 3, 1)
self.conv_up2 = convrelu(128 + 512, 256, 3, 1)
self.conv_up1 = convrelu(64 + 256, 256, 3, 1)
self.conv_up0 = convrelu(64 + 256, 128, 3, 1)
self.conv_original_size0 = convrelu(3, 64, 3, 1)
self.conv_original_size1 = convrelu(64, 64, 3, 1)
self.conv_original_size2 = convrelu(64 + 128, 64, 3, 1)
self.conv_last = nn.Conv2d(64, n_class, 1)
def forward(self, input):
x_original = self.conv_original_size0(input)
x_original = self.conv_original_size1(x_original)
layer0 = self.layer0(input)
layer1 = self.layer1(layer0)
layer2 = self.layer2(layer1)
layer3 = self.layer3(layer2)
layer4 = self.layer4(layer3)
layer4 = self.layer4_1x1(layer4)
x = self.upsample(layer4)
layer3 = self.layer3_1x1(layer3)
x = torch.cat([x, layer3], dim=1)
x = self.conv_up3(x)
x = self.upsample(x)
layer2 = self.layer2_1x1(layer2)
x = torch.cat([x, layer2], dim=1)
x = self.conv_up2(x)
x = self.upsample(x)
layer1 = self.layer1_1x1(layer1)
x = torch.cat([x, layer1], dim=1)
x = self.conv_up1(x)
x = self.upsample(x)
layer0 = self.layer0_1x1(layer0)
x = torch.cat([x, layer0], dim=1)
x = self.conv_up0(x)
x = self.upsample(x)
x = torch.cat([x, x_original], dim=1)
x = self.conv_original_size2(x)
out = self.conv_last(x)
return out
# +
# check keras-like model summary using torchsummary
from torchsummary import summary
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
model = ResNetUNet(6)
model = model.to(device)
summary(model, input_size=(3, 224, 224))
# +
from collections import defaultdict
import torch.nn.functional as F
import torch
from loss import dice_loss
def calc_loss(pred, target, metrics, bce_weight=0.5):
bce = F.binary_cross_entropy_with_logits(pred, target)
pred = torch.sigmoid(pred)
dice = dice_loss(pred, target)
loss = bce * bce_weight + dice * (1 - bce_weight)
metrics['bce'] += bce.data.cpu().numpy() * target.size(0)
metrics['dice'] += dice.data.cpu().numpy() * target.size(0)
metrics['loss'] += loss.data.cpu().numpy() * target.size(0)
return loss
def print_metrics(metrics, epoch_samples, phase):
outputs = []
for k in metrics.keys():
outputs.append("{}: {:4f}".format(k, metrics[k] / epoch_samples))
print("{}: {}".format(phase, ", ".join(outputs)))
def train_model(model, optimizer, scheduler, num_epochs=25):
best_model_wts = copy.deepcopy(model.state_dict())
best_loss = 1e10
for epoch in range(num_epochs):
print('Epoch {}/{}'.format(epoch, num_epochs - 1))
print('-' * 10)
since = time.time()
# Each epoch has a training and validation phase
for phase in ['train', 'val']:
if phase == 'train':
scheduler.step()
for param_group in optimizer.param_groups:
print("LR", param_group['lr'])
model.train() # Set model to training mode
else:
model.eval() # Set model to evaluate mode
metrics = defaultdict(float)
epoch_samples = 0
for inputs, labels in dataloaders[phase]:
inputs = inputs.to(device)
labels = labels.to(device)
# zero the parameter gradients
optimizer.zero_grad()
# forward
# track history if only in train
with torch.set_grad_enabled(phase == 'train'):
outputs = model(inputs)
loss = calc_loss(outputs, labels, metrics)
# backward + optimize only if in training phase
if phase == 'train':
loss.backward()
optimizer.step()
# statistics
epoch_samples += inputs.size(0)
print_metrics(metrics, epoch_samples, phase)
epoch_loss = metrics['loss'] / epoch_samples
# deep copy the model
if phase == 'val' and epoch_loss < best_loss:
print("saving best model")
best_loss = epoch_loss
best_model_wts = copy.deepcopy(model.state_dict())
time_elapsed = time.time() - since
print('{:.0f}m {:.0f}s'.format(time_elapsed // 60, time_elapsed % 60))
print('Best val loss: {:4f}'.format(best_loss))
# load best model weights
model.load_state_dict(best_model_wts)
return model
# +
import torch
import torch.optim as optim
from torch.optim import lr_scheduler
import time
import copy
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
print(device)
num_class = 6
model = ResNetUNet(num_class).to(device)
# freeze backbone layers
# Comment out to finetune further
for l in model.base_layers:
for param in l.parameters():
param.requires_grad = False
optimizer_ft = optim.Adam(filter(lambda p: p.requires_grad, model.parameters()), lr=1e-4)
exp_lr_scheduler = lr_scheduler.StepLR(optimizer_ft, step_size=10, gamma=0.1)
model = train_model(model, optimizer_ft, exp_lr_scheduler, num_epochs=15)
# +
#### prediction
import math
model.eval() # Set model to evaluate mode
test_dataset = SimDataset(3, transform = trans)
test_loader = DataLoader(test_dataset, batch_size=3, shuffle=False, num_workers=0)
inputs, labels = next(iter(test_loader))
inputs = inputs.to(device)
labels = labels.to(device)
pred = model(inputs)
pred = torch.sigmoid(pred)
pred = pred.data.cpu().numpy()
print(pred.shape)
# Change channel-order and make 3 channels for matplot
input_images_rgb = [reverse_transform(x) for x in inputs.cpu()]
# Map each channel (i.e. class) to each color
target_masks_rgb = [helper.masks_to_colorimg(x) for x in labels.cpu().numpy()]
pred_rgb = [helper.masks_to_colorimg(x) for x in pred]
helper.plot_side_by_side([input_images_rgb, target_masks_rgb, pred_rgb])
| pytorch_resnet18_unet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: py3
# language: python
# name: py3
# ---
# # Train a multi-layer perceptron (MLP) model
#
# In this notebook, we are going to use PySINGA to train a MLP model for classifying 2-d points into two categories (i.e., positive and negative). We use this example to illustrate the usage of PySINGA's modules. Please refer to the [documentation page](http://singa.apache.org/en/docs/index.html) for the functions of each module.
# +
from __future__ import print_function
from builtins import range
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# To import PySINGA modules
from singa import tensor
from singa import optimizer
from singa import loss
from singa import layer
#from singa.proto import model_pb2
# Task is to train a MLP model to classify 2-d points into the positive and negative categories.
#
# ## Training data generation
#
# The following steps would be conducted to generate the training data.
# 1. draw a boundary line in the 2-d space
# 2. generate data points in the 2-dspace
# 3. label the data points above the boundary line as positive points, and label other points as negative points.
# We draw the boundary line as $y=5x+1$
# generate the boundary
f = lambda x: (5 * x + 1)
bd_x = np.linspace(-1., 1, 200)
bd_y = f(bd_x)
# We generate the datapoints by adding a random noise to the data points on the boundary line
# generate the training data
x = np.random.uniform(-1, 1, 400)
y = f(x) + 2 * np.random.randn(len(x))
# We label the data points above the boundary line as positive points with label 1 and other data points with label 0 (negative).
# +
# convert training data to 2d space
label = np.asarray([5 * a + 1 > b for (a, b) in zip(x, y)])
data = np.array([[a,b] for (a, b) in zip(x, y)], dtype=np.float32)
plt.plot(bd_x, bd_y, 'k', label = 'boundary')
plt.plot(x[label], y[label], 'ro', ms=7)
plt.plot(x[~label], y[~label], 'bo', ms=7)
plt.legend(loc='best')
plt.show()
# -
# ## Create the MLP model
#
# 1. We will create a MLP by with one dense layer (i.e. fully connected layer).
# 2. We use the Softmax function to get compute the probability of each category for every data point.
# 3. We use the cross-entropy as the loss function.
# 4. We initialize the weight matrix following guassian distribution (mean=0, std=0.1), and set the bias to 0.
# 5. We creat a SGD updater to update the model parameters.
#
# 2 and 3 are combined by the SoftmaxCrossEntropy.
# +
# create layers
layer.engine = 'singacpp'
dense = layer.Dense('dense', 2, input_sample_shape=(2,))
p = dense.param_values()
print(p[0].shape, p[1].shape)
# init parameters
p[0].gaussian(0, 0.1) # weight matrix
p[1].set_value(0) # bias
# setup optimizer and loss func
opt = optimizer.SGD(lr=0.05)
lossfunc = loss.SoftmaxCrossEntropy()
# -
# * Each layer is created with a layer name and other meta data, e.g., the dimension size for the dense layer. The last argument is the shape of a single input sample of this layer.
# * **param_values()** returns a list of tensors as the parameter objects of this layer
# * SGD optimzier is typically created with the weight decay, and momentum specified. The learning rate could be specified at creation or passed in when the optimizer is applied.
# ## Train the model
#
# We run 1000 iterations to train the MLP model.
# 1. For each iteration, we compute the gradient of the models parameters and use them to update the model parameters.
# 2. Periodically, we plot the prediction from the model.
# +
tr_data = tensor.from_numpy(data)
tr_label = tensor.from_numpy(label.astype(int))
# plot the classification results using the current model parameters
def plot_status(w, b, title='origin'):
global bd_x, bd_y, data
pr = np.add(np.dot(data, w), b)
lbl = pr[:, 0] < pr[:, 1]
plt.figure(figsize=(6,3));
plt.plot(bd_x, bd_y, 'k', label='truth line')
plt.plot(data[lbl, 0], data[lbl, 1], 'ro', ms=7)
plt.plot(data[~lbl, 0], data[~lbl, 1], 'bo', ms=7)
plt.legend(loc='best')
plt.title(title)
plt.xlim(-1, 1);
plt.ylim(data[:, 1].min()-1, data[:, 1].max()+1)
# sgd
for i in range(1000):
act = dense.forward(True, tr_data)
lvalue = lossfunc.forward(True, act, tr_label)
dact = lossfunc.backward()
dact /= tr_data.shape[0]
_, dp = dense.backward(True, dact)
# update the parameters
opt.apply(i, dp[0], p[0], 'w')
opt.apply(i, dp[1], p[1], 'b')
if (i%100 == 0):
print('training loss = %f' % lvalue.l1())
plot_status(tensor.to_numpy(p[0]), tensor.to_numpy(p[1]),title='epoch %d' % i)
#train(dat, label)
# -
# The layer class has forward and backward functions for back-propagation.
# * forward() accepts two arguments, the first one indicates the phase (training or evaluation); the second one includes the input tensor(s); It outputs the layer values as a single or a list of tensors.
# * backward() accepts two arguments, the first one is not used currently; the second one includes the gradients of the layer values. It outputs a tuple, where the first field includes the gradient tensor(s) of the input(s), and the second field includes a list of gradients for the parameters.
#
# The optimzier class **apply** function updates the parameter values using the gradients. The first argument is the iteration ID, followed by the gradient tensor and the value tensor. Each parameter tensor has a name associated with it, which is used by Optimizer to keep some internal data (e.g., history gradients) for each parameter.
#
# The loss class computes the loss value given the predictions and the ground truth in **forward()** function. It computes the gradients of the predictions w.r.t the loss function and outputs the gradient tensor(s) by **backward()** function.
# ## Observation
#
# We can see that prediction of the data points are getting correct labels with the training going on.
# # Next [CNN example](./cnn.ipynb)
| doc/en/docs/notebook/mlp.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .groovy
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Groovy
// language: groovy
// name: groovy
// ---
def plot = new Plot(title: "Setting 2nd Axis bounds")
def ys = [0, 2, 4, 6, 15, 10]
def ys2 = [-40, 50, 6, 4, 2, 0]
def ys3 = [3, 6, 3, 6, 70, 6]
plot << new YAxis(label:"Spread")
plot << new Line(y: ys)
plot << new Line(y: ys2, yAxis: "Spread")
//plot << new Line(y: ys3, yAxis: "Spread")
//plot.getYAxes()[0].setBound(1,5);
//plot.getYAxes()[1].setBound(3,6) // this should change the bounds of the 2nd, right axis
plot
def plot = new CategoryPlot()
def cs = [new Color(255, 0, 0, 128)] * 5 // transparent bars
def cs1 = [new Color(0, 0, 0, 128)] * 5 // transparent bars
plot << new YAxis(label:"nd")
plot << new CategoryBars(value: [[1, 2, 3], [1, 3, 5]], color: cs)
plot << new CategoryBars(value: [[4, 4, 4], [6,7, 8]], color: cs1, yAxis: "nd")
plot.getYAxes()[0].setBound(0,5)
plot.getYAxes()[1].setBound(-20,20) // this should change the bounds of the 2nd, right axis
plot
def plot = new Plot();
def y1 = [1.5, 1, 6, 5, 2, 8]
def y2 = [10, 10, 60, 50, 20, 50]
plot << new YAxis(label:"nd")
def cs = [Color.black, Color.red, Color.gray, Color.green, Color.blue, Color.pink]
def ss = [StrokeType.SOLID, StrokeType.SOLID, StrokeType.DASH, StrokeType.DOT, StrokeType.DASHDOT, StrokeType.LONGDASH]
plot.getYAxes()[1].setBound(0,20)
plot << new Stems(y: y2, color: cs, width: 2)
plot << new Stems(y: y1, color: cs, style: ss, width: 6, yAxis: "nd")
def plot = new Plot(title: "Bars")
def cs = [new Color(255, 0, 0, 128)] * 5 // transparent bars
def cs1 = [new Color(0, 0, 0, 128)] * 5 // transparent bars
plot << new YAxis(label:"nd")
cs[3] = Color.red // set color of a single bar, solid colored bar
plot << new Bars(x: (1..5), y: [3, 6, 5, 99, 8], color: cs, outlineColor: Color.black, width: 0.3)
plot << new Bars(x: (1..5), y: [1, 2, 3, 4, 11], color: cs1, outlineColor: Color.black, width: 0.3, yAxis: "nd")
//plot.getYAxes()[1].setBound(0,5)
plot
def plot = new Plot(title: "Changing Point Size, Color, Shape")
def y1 = [6, 7, 12, 11, 8, 14]
def y2 = y1.collect { it - 2 }
def y3 = y2.collect { it - 2 }
def y4 = y3.collect { it - 2 }
plot << new YAxis(label:"nd")
plot << new Points(y: y1)
plot << new Points(y: y2, shape: ShapeType.CIRCLE, yAxis: "nd")
plot << new Points(y: y3, size: 8.0, shape: ShapeType.DIAMOND)
plot << new Points(y: y4, size: 12.0, color: Color.orange, outlineColor: Color.red)
plot.getYAxes()[1].setBound(5,10)
plot
def p = new Plot()
p << new YAxis(label:"nd")
p << new Line(y: [3, 6, 12, 24], displayName: "Median")
p << new Area(y: [4, 8, 16, 32], base: [2, 4, 8, 16],
color: new Color(255, 0, 0, 50), displayName: "Q1 to Q3", yAxis: "nd")
p.getYAxes()[1].setBound(0,100)
p
| doc/groovy/2ndYaxis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# [source](../api/alibi_detect.cd.chisquare.rst)
# # Chi-Squared
#
# ## Overview
#
# The drift detector applies feature-wise [Chi-Squared](https://en.wikipedia.org/wiki/Chi-squared_test) tests for the categorical features. For multivariate data, the obtained p-values for each feature are aggregated either via the [Bonferroni](https://mathworld.wolfram.com/BonferroniCorrection.html) or the [False Discovery Rate](http://www.math.tau.ac.il/~ybenja/MyPapers/benjamini_hochberg1995.pdf) (FDR) correction. The Bonferroni correction is more conservative and controls for the probability of at least one false positive. The FDR correction on the other hand allows for an expected fraction of false positives to occur. Similarly to the other drift detectors, a preprocessing steps could be applied, but the output features need to be categorical.
# ## Usage
#
# ### Initialize
#
#
# Parameters:
#
# * `p_val`: p-value used for significance of the Chi-Squared test for each feature. If the FDR correction method is used, this corresponds to the acceptable q-value.
#
# * `X_ref`: Data used as reference distribution.
#
# * `preprocess_X_ref`: Whether to already count and store the number of instances for each possible category of each variable of the reference data `X_ref` when initializing the detector. If a preprocessing step is specified, the step will be applied first. Defaults to *True*. It is possible that it needs to be set to *False* if the preprocessing step requires statistics from both the reference and test data.
#
# * `categories_per_feature`: Optional dictionary with as keys the feature column index and as values the number of possible categorical values for that feature. E.g.: *{0: 5, 1: 9, 2: 7}*. If it is not specified, `categories_per_feature` is inferred from `X_ref`.
#
# * `update_X_ref`: Reference data can optionally be updated to the last N instances seen by the detector or via [reservoir sampling](https://en.wikipedia.org/wiki/Reservoir_sampling) with size N. For the former, the parameter equals *{'last': N}* while for reservoir sampling *{'reservoir_sampling': N}* is passed.
#
# * `preprocess_fn`: Function to preprocess the data before computing the data drift metrics. Typically a dimensionality reduction technique. Needs to return categorical features for the Chi-Squared detector.
#
# * `preprocess_kwargs`: Keyword arguments for `preprocess_fn`.
#
# * `correction`: Correction type for multivariate data. Either *'bonferroni'* or *'fdr'* (False Discovery Rate).
#
# * `n_features`: Number of features used in the Chi-Squared test. No need to pass it if no preprocessing takes place. In case of a preprocessing step, this can also be inferred automatically but could be more expensive to compute.
#
# * `n_infer`: If the number of features needs to be inferred after the preprocessing step, we can specify the number of instances used to infer the number of features from since this can depend on the specific preprocessing step.
#
# * `data_type`: can specify data type added to metadata. E.g. *'tabular'*.
#
# Initialized drift detector example:
#
# ```python
# from alibi_detect.cd import ChiSquareDrift
#
# cd = ChiSquareDrift(p_val=0.05, X_ref=X_ref)
# ```
# ### Detect Drift
#
# We detect data drift by simply calling `predict` on a batch of instances `X`. We can return the feature-wise p-values before the multivariate correction by setting `return_p_val` to *True*. The drift can also be detected at the feature level by setting `drift_type` to *'feature'*. No multivariate correction will take place since we return the output of *n_features* univariate tests. For drift detection on all the features combined with the correction, use *'batch'*. `return_p_val` equal to *True* will also return the threshold used by the detector (either for the univariate case or after the multivariate correction).
#
# The prediction takes the form of a dictionary with `meta` and `data` keys. `meta` contains the detector's metadata while `data` is also a dictionary which contains the actual predictions stored in the following keys:
#
# * `is_drift`: 1 if the sample tested has drifted from the reference data and 0 otherwise.
#
# * `p_val`: contains feature-level p-values if `return_p_val` equals *True*.
#
# * `threshold`: for feature-level drift detection the threshold equals the p-value used for the significance of the Chi-Square test. Otherwise the threshold after the multivariate correction (either *bonferroni* or *fdr*) is returned.
#
# * `distance`: feature-wise Chi-Square test statistics between the reference data and the new batch if `return_distance` equals *True*.
#
#
# ```python
# preds_drift = cd.predict(X, drift_type='batch', return_p_val=True, return_distance=True)
# ```
# ### Saving and loading
#
# The drift detectors can be saved and loaded in the same way as other detectors when using the built-in preprocessing steps (`alibi_detect.cd.preprocess.UAE` and `alibi_detect.cd.preprocess.HiddenOutput`) or no preprocessing at all:
#
# ```python
# from alibi_detect.utils.saving import save_detector, load_detector
#
# filepath = 'my_path'
# save_detector(cd, filepath)
# cd = load_detector(filepath)
# ```
#
# A custom preprocessing step can be passed as follows:
#
# ```python
# cd = load_detector(filepath, **{'preprocess_kwargs': preprocess_kwargs})
# ```
# ## Examples
#
# [Drift detection on income prediction](../examples/cd_chi2ks_adult.nblink)
| doc/source/methods/chisquaredrift.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Q#
# language: qsharp
# name: iqsharp
# ---
# # Superposition Kata Workbook, Part 2
#
# The [Superposition Kata Workbook, Part 1](./Workbook_Superposition.ipynb) includes the solutions of kata tasks 1 - 7. Part 2 continues the explanations for the rest of the tasks.
# To begin, first prepare this notebook for execution (if you skip this step, you'll get "Syntax does not match any known patterns" error when you try to execute Q# code in the next cells):
%package Microsoft.Quantum.Katas::0.11.2003.3107
# > The package versions in the output of the cell above should always match. If you are running the Notebooks locally and the versions do not match, please install the IQ# version that matches the version of the `Microsoft.Quantum.Katas` package.
# > <details>
# > <summary><u>How to install the right IQ# version</u></summary>
# > For example, if the version of `Microsoft.Quantum.Katas` package above is 0.1.2.3, the installation steps are as follows:
# >
# > 1. Stop the kernel.
# > 2. Uninstall the existing version of IQ#:
# > dotnet tool uninstall microsoft.quantum.iqsharp -g
# > 3. Install the matching version:
# > dotnet tool install microsoft.quantum.iqsharp -g --version 0.1.2.3
# > 4. Reinstall the kernel:
# > dotnet iqsharp install
# > 5. Restart the Notebook.
# > </details>
#
# ## <a name="greenberger-horne-zeilinger"></a> Task 8. Greenberger-Horne-Zellinger state.
#
# **Input:** $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state (stored in an array of length $N$).
#
# **Goal:** Change the state of the qubits to the GHZ state $\frac{1}{\sqrt{2}} \big (|0\dots0\rangle + |1\dots1\rangle\big)$.
# ### Solution
#
# The single-qubit GHZ state is the plus state $\frac{1}{\sqrt{2}} \big (|0\rangle + |1\rangle\big)$ that we've discussed in [task 1](./Workbook_Superposition.ipynb#plus-state). As a reminder, that state is prepared by applying a Hadamard gate.
#
# The 2-qubit GHZ state is the Bell state $\frac{1}{\sqrt{2}} \big (|00\rangle + |11\rangle\big)$ that we've discussed in [task 6](./Workbook_Superposition.ipynb#bell-state). That state can been prepared using the following circuit:
#
# <img src="./img/Task6HadamardCNOTCircuit.png"/>
#
# The next one is the 3-qubit GHZ state:
# $$|GHZ\rangle = \frac{1}{\sqrt{2}} \big (|000\rangle + |111\rangle\big)$$
#
# Let's use the 2-qubit circuit as a building block to construct the circuit for 3 qubits. First, let's add a third qubit to the above circuit:
#
# <img src="./img/Task8Hadamardand3rdqubitircuit.png"/>
#
# Comparing the state prepared by this circuit with the desired end state, we see that they differ only in the third (rightmost) qubit:
#
# $$|\Phi^+\rangle |0\rangle = \frac{1}{\sqrt{2}} \big (|000\rangle + |11\color{red}0\rangle\big)$$
# $$|GHZ\rangle = \frac{1}{\sqrt{2}} \big (|000\rangle + |11\color{red}1\rangle\big)$$
#
#
# Applying a controlled NOT operation using the first (leftmost) qubit as the control bit and the third (rightmost) qubit as the target qubit allows us to fix this difference:
#
# <table style="background-color: white; border:0 solid; tr { background-color:white; }">
# <col width=30%>
# <col width=70%>
# <td style="text-align:center; background-color:white; border:0"><img src="./img/Task8HadamardAndCNOTCircuit.png"/></td>
# <td style="text-align:left; background-color:white; border:0"><img src="./img/Task8CNOTFlip.png"/></td>
# </table>
#
# Similarly, the following circuit will prepare the GHZ state on four qubits $\frac{1}{\sqrt2} (|0000\rangle + |1111\rangle$:
# <img src="./img/Task84QubitCircuit.png"/>
#
# Thus we can come to the general solution: apply Hadamard gate to the first qubit and do a series of CNOT gates with the first qubit as control and each of the other qubits as targets.
# +
%kata T08_GHZ_State_Test
open Microsoft.Quantum.Arrays;
operation GHZ_State (qs : Qubit[]) : Unit {
H(qs[0]);
// Library function Rest returns all array elements except for the first one
for (q in Rest(qs)) {
CNOT(qs[0], q);
}
}
# -
# [Return to task 8 of the Superposition kata.](./Superposition.ipynb#greenberger-horne-zeilinger)
# ## <a name="superposition-of-all-basis-vectors"></a> Task 9. Superposition of all basis vectors.
#
# **Input:** $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state.
#
# **Goal:** Change the state of the qubits to an equal superposition of all basis vectors $\frac{1}{\sqrt{2^N}} \big (|0 \dots 0\rangle + \dots + |1 \dots 1\rangle\big)$.
#
# > For example, for $N = 2$ the final state should be $\frac{1}{\sqrt{2}} \big (|00\rangle + |01\rangle + |10\rangle + |11\rangle\big)$.
# ### Solution
#
# As we've seen in [task 4](./Workbook_Superposition.ipynb#superposition-of-all-basis-vectors-on-two-qubits), to prepare a superposition of all basis vectors on 2 qubits we need to apply a Hadamard gate to each of the qubits.
#
# It seems that the solution for the general case might be to apply a Hadamard gate to every qubit as well. Let's check the first few examples:
#
# \begin{align*}
# H|0\rangle &= \frac{1}{\sqrt2}\big(|0\rangle + |1\rangle\big)\\
# H|0\rangle \otimes H|0\rangle &= \frac{1}{\sqrt2} \big(|0\rangle + |1\rangle\big) \otimes \frac{1}{\sqrt2} \big(|0\rangle + |1\rangle\big)\\
# &= \frac{1}{\sqrt{2^2}}\big(|00\rangle + |01\rangle+ |10\rangle+ |11\rangle\big)\\
# H|0\rangle \otimes H|0\rangle \otimes H|0\rangle &= \frac{1}{\sqrt{2^2}}\big(|00\rangle + |01\rangle+ |10\rangle+ |11\rangle\big) \otimes \frac{1}{\sqrt2}\big(|0\rangle + |1\rangle\big)\\
# &= \frac{1}{\sqrt{2^3}}\big(|000\rangle + |001\rangle + |010\rangle+ |100\rangle+ |110\rangle + |101\rangle+ |011\rangle+ |111\rangle\big)\\
# \underset{N}{\underbrace{H|0\rangle \otimes \dots \otimes H|0\rangle}}
# &= \frac{1}{\sqrt{2^{N-1}}} \big( |\underset{N-1}{\underbrace{0 \cdots 0}}\rangle + \cdots + |\underset{N-1}{\underbrace{1 \cdots 1}}\rangle \big) \otimes \frac{1}{\sqrt2}\big(|0\rangle + |1\rangle\big) = \\
# &= \frac{1}{\sqrt{2^N}} \big( |\underset{N}{\underbrace{0 \cdots 0}}\rangle + \cdots + |\underset{N}{\underbrace{1 \cdots 1}}\rangle \big)\\
# \end{align*}
#
# Thus, the solution requires us to iterate over the qubit array and to apply the Hadamard gate to each element as follows:
# +
%kata T09_AllBasisVectorsSuperposition_Test
operation AllBasisVectorsSuperposition (qs : Qubit[]) : Unit {
for (q in qs) {
H(q);
}
}
# -
# [Return to task 9 of the Superposition kata.](./Superposition.ipynb#superposition-of-all-basis-vectors)
# ## <a name="superposition-of-all-even-or-all-odd-numbers"></a>Task 10. Superposition of all even or all odd numbers.
#
# **Inputs:**
#
# 1. $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state (stored in an array of length $N$).
# 2. A boolean `isEven`.
#
# **Goal:** Prepare a superposition of all *even* numbers if `isEven` is `true`, or of all *odd* numbers if `isEven` is `false`.
# A basis state encodes an integer number using [big-endian](https://en.wikipedia.org/wiki/Endianness) binary notation: state $|01\rangle$ corresponds to the integer $1$, and state $|10 \rangle$ - to the integer $2$.
#
# > For example, for $N = 2$ and `isEven = false` you need to prepare superposition $\frac{1}{\sqrt{2}} \big (|01\rangle + |11\rangle\big )$,
# and for $N = 2$ and `isEven = true` - superposition $\frac{1}{\sqrt{2}} \big (|00\rangle + |10\rangle\big )$.
# </details>
# ### Solution
#
# Let’s look at some examples of basis states to illustrate the binary numbering system.
#
# The 4 basis states on $N = 2$ qubits can be split in two columns, where the left column represents the basis states that form the required superposition state for `isEven = false` and the right column - the basis states that form the required superposition state for `isEven = true`:
#
# <img src="./img/Task10_1.png" width="400">
#
# If we do the same basis state split for $N = 3$ qubits, the pattern becomes more obvious:
#
# <img src="./img/Task10_2.png" width="400">
# The two leftmost qubits go through all possible basis states for `isEven = false` and for `isEven = true`, and the rightmost qubit stays in the $|1\rangle$ state for `isEven = false` and in the $|0\rangle$ state for `isEven = true`.
#
# A quick sanity check for $N = 4$ qubits re-confirms the pattern:
#
# <img src="./img/Task10_3.png" width="400">
#
# Again, the three leftmost qubits go through all possible basis states in both columns, and the rightmost qubit stays in the same state in each column.
#
# The solution is to put all qubits except the rightmost one into an equal superposition (similar to what we did in Task 9) and to set the rightmost qubit to $|0\rangle$ or $|1\rangle$ depending on the `isEven` flag, using the X operator to convert $|0\rangle$ to $|1\rangle$ if `isEven = false`.
# +
%kata T10_EvenOddNumbersSuperposition_Test
operation EvenOddNumbersSuperposition (qs : Qubit[], isEven : Bool) : Unit is Adj {
let N = Length(qs);
for (i in 0 .. N-2) {
H(qs[i]);
}
// for odd numbers, flip the last bit to 1
if (not isEven) {
X(qs[N-1]);
}
}
# -
# [Return to task 10 of the Superposition kata.](./Superposition.ipynb#superposition-of-all-even-or-all-odd-numbers)
# ## <a name="threestates-twoqubits"></a>Task 11*. $\frac{1}{\sqrt{3}} \big(|00\rangle + |01\rangle + |10\rangle\big)$ state.
#
# **Input:** Two qubits in the $|00\rangle$ state.
#
# **Goal:** Change the state of the qubits to $\frac{1}{\sqrt{3}} \big(|00\rangle + |01\rangle + |10\rangle\big)$.
# ### Solution
#
# *Coming up soon... Meanwhile, follow Niel's answer at https://quantumcomputing.stackexchange.com/a/2313/*
# +
%kata T11_ThreeStates_TwoQubits_Test
open Microsoft.Quantum.Math;
operation ThreeStates_TwoQubits (qs : Qubit[]) : Unit {
// Rotate first qubit to (sqrt(2) |0⟩ + |1⟩) / sqrt(3) (task 1.4 from BasicGates kata)
let theta = ArcSin(1.0 / Sqrt(3.0));
Ry(2.0 * theta, qs[0]);
// Split the state sqrt(2) |0⟩ ⊗ |0⟩ into |00⟩ + |01⟩
(ControlledOnInt(0, H))([qs[0]], qs[1]);
}
# -
# [Return to task 11 of the Superposition kata.](./Superposition.ipynb#threestates-twoqubits)
# ## <a name="hardy-state"></a>Task 12*. Hardy state.
#
# **Input:** Two qubits in the $|00\rangle$ state.
#
# **Goal:** Change the state of the qubits to $\frac{1}{\sqrt{12}} \big(3|00\rangle + |01\rangle + |10\rangle + |11\rangle \big)$.
# ### Solution
#
# *Coming up soon...*
# +
%kata T12_Hardy_State_Test
open Microsoft.Quantum.Math;
operation Hardy_State (qs : Qubit[]) : Unit {
Ry(2.0 * ArcCos(Sqrt(10.0 / 12.0)), qs[0]);
(ControlledOnInt(0, Ry))([qs[0]], (2.0 * ArcCos(Sqrt(9.0 / 10.0)), qs[1]));
Controlled H([qs[0]], qs[1]);
}
# -
# [Return to task 12 of the Superposition kata.](./Superposition.ipynb#hardy-state)
# ## <a name="superposition-of-zero-and-given-bit-string"></a> Task 13. Superposition of $|0 \dots 0\rangle$ and the given bit string.
#
# **Inputs:**
#
# 1. $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state.
# 2. A bit string of length $N$ represented as `Bool[]`. Bit values `false` and `true` correspond to $|0\rangle$ and $|1\rangle$ states. You are guaranteed that the first bit of the bit string is `true`.
#
# **Goal:** Change the state of the qubits to an equal superposition of $|0 \dots 0\rangle$ and the basis state given by the bit string.
#
# > For example, for the bit string `[true, false]` the state required is $\frac{1}{\sqrt{2}}\big(|00\rangle + |10\rangle\big)$.
# ### Solution
#
# > A common strategy for preparing a superposition state in a qubit register is using an auxiliary qubit (or several, for more complicated states). The auxiliary qubit can be put into a superposition state through the usual means of applying a Hadamard gate (or a rotation about the Y axis for an uneven superposition).
# > Then the basis states of the desired superposition are prepared individually based on the auxiliary qubit state by using it as the control qubit for a CNOT gate. One of the basis states will be prepared controlled on the $|0\rangle$ component of the auxiliary state, and the other - controlled on the $|1\rangle$ component.
# > Finally, you have to return the auxiliary qubit to the $|0\rangle$ state by uncomputing it, i.e., by using the basis state prepared from the $|1\rangle$ component as the control qubits for a CNOT gate with the auxiliary qubit as the target.
# >
# > More details on using this approach can be found in the solution to tasks [15](#superposition-of-four-bit-strings) and [16](#wstate-on-2k-qubits). However, for this task we can come up with a simpler solution.
# > Instead of allocating a new qubit to use as the auxiliary, we can use the first qubit in the register for this purpose, because we are guaranteed that the first bit in the two basis vectors that comprise the required superposition is different.
# > This saves us the need to allocate a new qubit and lets us skip the uncomputing step, as the qubit acting as the control for the next preparation steps is part of the desired result.
#
# Consider the earlier tasks in this kata that asked to prepare Bell states and GHZ state; the structure of the superposition state in this task is a more general case of those scenarios: all of them ask to prepare an equal superposition of two different basis states.
#
# The first step of the solution is the same as in those tasks: put the first qubit in the register into an equal superposition of $|0\rangle$ and $|1\rangle$ using the H gate to get the following state:
#
# $$\frac{1}{\sqrt2} (|0\rangle + |1\rangle) \otimes |0 \dots 0\rangle = \frac{1}{\sqrt2} (|00 \dots 0\rangle + |10 \dots 0\rangle)$$
#
# The first term of the superposition already matches the desired state, so we need to fix the second term.
# To do that, we will walk through the remaining qubits in the register, checking if the bit in the corresponding position of the bit string `bits` is `true`.
# If it is, that qubit's state needs to be adjusted from $0$ to $1$ in the second term of our superposition (and left unchanged in the first term).
# We can do this change using the CNOT gate with the first qubit as the control and the current qubit as the target.
# When we have finished walking through the register like this, the register will be in the desired superposition.
# +
%kata T13_ZeroAndBitstringSuperposition_Test
operation ZeroAndBitstringSuperposition (qs : Qubit[], bits : Bool[]) : Unit {
H(qs[0]);
for (i in 1 .. Length(qs) - 1) {
if (bits[i]) {
CNOT(qs[0], qs[i]);
}
}
}
# -
# [Return to task 13 of the Superposition kata.](./Superposition.ipynb#superposition-of-zero-and-given-bit-string)
# ### <a name="superposition-of-two-bit-strings"></a> Task 14. Superposition of two bit strings.
#
# **Inputs:**
#
# 1. $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state.
# 2. Two bit strings of length $N$ represented as `Bool[]`s. Bit values `false` and `true` correspond to $|0\rangle$ and $|1\rangle$ states. You are guaranteed that the two bit strings differ in at least one bit.
#
# **Goal:** Change the state of the qubits to an equal superposition of the basis states given by the bit strings.
#
# > For example, for bit strings `[false, true, false]` and `[false, false, true]` the state required is $\frac{1}{\sqrt{2}}\big(|010\rangle + |001\rangle\big)$.
# ### Solution
#
# The strategy of using an auxiliary qubit to control the preparation process described in the previous task can be applied to this task as well.
#
# We will start by allocating an auxiliary qubit and preparing it in the $\frac{1}{\sqrt2} (|0\rangle + |1\rangle)$ state using the H gate. The overall state of the system will be
#
# $$\frac{1}{\sqrt2} (|0\rangle + |1\rangle)_a \otimes |0 \dots 0\rangle_r = \frac{1}{\sqrt2} (|0\rangle_a \otimes |0 \dots 0\rangle_r + |1\rangle_a \otimes |0 \dots 0\rangle_r)$$
#
# At this point, we can prepare the two basis states of the target state separately, bit by bit, controlling the preparation of one of them on the $|0\rangle$ state of the auxiliary qubit and the preparation of the other one - on the $|1\rangle$ state.
# If a bit in one of the bit strings is `true`, we will apply a controlled X gate with the auxiliary qubit as control, the qubit in the corresponding position of the register as target, and control it on the $|0\rangle$ or the $|1\rangle$ state depending on which bit string we are considering at the moment.
# Such controlled gate can be implemented using [`ControlledOnInt`](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.canon.controlledonint) library function.
#
# After this the state of the system will be
# $$\frac{1}{\sqrt2} (|0\rangle_a \otimes |bits_1\rangle_r + |1\rangle_a \otimes |bits_2\rangle_r)$$
#
# Finally, we will uncompute the auxiliary qubit by using [`ControlledOnBitString`](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.canon.controlledonbitstring) library function with the second bit string and the `X` operation as arguments, the quantum register as the control, and the auxiliary qubit as the target.
# This will affect only the $|1\rangle_a \otimes |bits_2\rangle_r$ term, flipping the state of the auxiliary qubit in it and bringing the system to its final state:
#
# $$|0\rangle_a \otimes \frac{1}{\sqrt2} (|bits_1\rangle + |bits_2\rangle)_r$$
# +
%kata T14_TwoBitstringSuperposition_Test
operation TwoBitstringSuperposition (qs : Qubit[], bits1 : Bool[], bits2 : Bool[]) : Unit {
using (q = Qubit()) {
H(q);
for (i in 0 .. Length(qs) - 1) {
if (bits1[i]) {
(ControlledOnInt(0, X))([q], qs[i]);
}
if (bits2[i]) {
(ControlledOnInt(1, X))([q], qs[i]);
}
}
// uncompute the auxiliary qubit to release it
(ControlledOnBitString(bits2, X))(qs, q);
}
}
# -
# It is also possible to solve the task without using an extra qubit, if instead we use one of the qubits in the register in this role.
# While walking through the register and bit strings, the first time the bit strings disagreed, the qubit in the corresponding position would take on the role of the auxiliary qubit; we would put it in superposition using the H gate and perform all subsequent bit flips using that qubit as the control.
#
# This saves us an additional qubit and allows to skip the uncomputing step, though the code becomes less elegant.
# We will move the classical logic of comparing two bit strings to find the first position in which they differ to a function `FindFirstDiff`; note that it has to be defined in a separate code cell.
function FindFirstDiff (bits1 : Bool[], bits2 : Bool[]) : Int {
for (i in 0 .. Length(bits1) - 1) {
if (bits1[i] != bits2[i]) {
return i;
}
}
return -1;
}
# +
%kata T14_TwoBitstringSuperposition_Test
operation TwoBitstringSuperposition (qs : Qubit[], bits1 : Bool[], bits2 : Bool[]) : Unit {
// find the index of the first bit at which the bit strings are different
let firstDiff = FindFirstDiff(bits1, bits2);
// Hadamard corresponding qubit to create superposition
H(qs[firstDiff]);
// iterate through the bit strings again setting the final state of qubits
for (i in 0 .. Length(qs) - 1) {
if (bits1[i] == bits2[i]) {
// if two bits are the same, apply X or nothing
if (bits1[i]) {
X(qs[i]);
}
} else {
// if two bits are different, set their difference using CNOT
if (i > firstDiff) {
CNOT(qs[firstDiff], qs[i]);
if (bits1[i] != bits1[firstDiff]) {
X(qs[i]);
}
}
}
}
}
# -
# [Return to task 14 of the Superposition kata.](./Superposition.ipynb#superposition-of-two-bit-strings)
# ### <a name="superposition-of-four-bit-strings"></a>Task 15*. Superposition of four bit strings.
#
# **Inputs:**
#
# 1. $N$ ($N \ge 1$) qubits in the $|0 \dots 0\rangle$ state.
# 2. Four bit strings of length $N$, represented as `Bool[][]` `bits`. `bits` is an $4 \times N$ array which describes the bit strings as follows: `bits[i]` describes the `i`-th bit string and has $N$ elements. You are guaranteed that all four bit strings will be distinct.
#
# **Goal:** Change the state of the qubits to an equal superposition of the four basis states given by the bit strings.
#
# > For example, for $N = 3$ and `bits = [[false, true, false], [true, false, false], [false, false, true], [true, true, false]]` the state required is $\frac{1}{2}\big(|010\rangle + |100\rangle + |001\rangle + |110\rangle\big)$.
# ### Solutions
# #### Solution 1
#
# We are going to use the same trick of auxiliary qubits that we used in [the previous task](#superposition-of-two-bit-strings).
# Since the desired superposition has 4 basis states with equal amplitudes, we are going to need two qubits to define a unique basis to control preparation of each of the basis states in the superposition.
#
# We start by allocating two extra qubits and preparing an equal superposition of all 2-qubit states on them by applying an H gate to each of them:
#
# $$\frac12 (|00\rangle + |01\rangle + |10\rangle + |11\rangle)_a \otimes |0 \dots 0\rangle_r$$
#
# Then, for each of the four given bit strings, we walk through it and prepare the matching basis state on the main register of qubits, using controlled X gates with the corresponding basis state of the auxiliary qubits as control.
# For example, when preparing the bit string `bits[0]`, we apply X gates controlled on the basis state $|00\rangle$; when preparing the bit string `bits[1]`, we apply X gates controlled on $|10\rangle$, and so on.
#
# > We can choose an arbitrary matching of the 2-qubit basis states used as controls and the bit strings prepared on the main register.
# > Since all amplitudes are the same, the result does not depend on which state controlled which bit string preparation.
# > It can be convenient to use indices of the bit strings, converted to little-endian, to control preparation of the bit strings.
# > Q# library function [`ControlledOnInt`](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.canon.controlledonint) does exactly that.
#
# After this the system will be in the state
#
# $$\frac12 (|00\rangle_a |bits_0\rangle_r + |10\rangle_a |bits_1\rangle_r + |01\rangle_a |bits_2\rangle_r + |11\rangle_a |bits_3\rangle_r)$$
#
# As the last step, we must uncompute the auxiliary qubits, i.e., return them to the $|00\rangle$ state to unentangle them from the main register.
# Same as we did in the previous task, we will use [`ControlledOnBitString`](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.canon.controlledonbitstring) with the corresponding bit string and the X operation as arguments, the quantum register as the control, and the auxiliary qubits as the target.
# We will uncompute each of them separately, so one of the auxiliary qubits will be uncomputed with the `bits[1]` and `bits[3]` bit strings as controls, and the other - with the `bits[2]` and `bits[3]`.
# +
%kata T15_FourBitstringSuperposition_Test
operation FourBitstringSuperposition (qs : Qubit[], bits : Bool[][]) : Unit {
using (anc = Qubit[2]) {
// Put two ancillas into equal superposition of 2-qubit basis states
ApplyToEachA(H, anc);
// Set up the right pattern on the main qubits with control on ancillas
for (i in 0 .. 3) {
for (j in 0 .. Length(qs) - 1) {
if (bits[i][j]) {
(ControlledOnInt(i, X))(anc, qs[j]);
}
}
}
// Uncompute the ancillas, using patterns on main qubits as control
for (i in 0 .. 3) {
if (i % 2 == 1) {
(ControlledOnBitString(bits[i], X))(qs, anc[0]);
}
if (i / 2 == 1) {
(ControlledOnBitString(bits[i], X))(qs, anc[1]);
}
}
}
}
# -
# #### Solution 2
#
# We are going to leverage the recursion abilities of Q# to create a superposition of the four bit strings. This solution also extends to an arbitrary number of bit strings with no code changes.
#
# For this process we will look at the first bits of each string and adjust the probability of measuring a $|0\rangle$ or $|1\rangle$ accordingly on the first qubit of our answer. We will then recursively call (as needed) the process again to adjust the probabilities of measurement on the second bit depending on the first bit. This process recurses until no more input bits are provided.
#
# Consider for example the following four bit strings on which to create a superposition:
# $|001\rangle, |101\rangle, |111\rangle, |110\rangle$.
#
# We can rewrite the superposition state we need to prepare as
#
# $$\frac12 \big(|001\rangle + |101\rangle + |111\rangle + |110\rangle \big) = \frac12 |0\rangle \otimes |01\rangle + \frac{\sqrt3}{2} |1\rangle \otimes \frac{1}{\sqrt3} \big(|10\rangle + |11\rangle + |10\rangle \big)$$
#
# As the first step of the solution, we need to prepare a state $\frac12 |0\rangle + \frac{\sqrt3}{2} |1\rangle$ on the first qubit (to measure $|0\rangle$ with $\frac14$ probability and to measure $|1\rangle$ with $\frac34$ probability). To do this, we will apply an [`Ry`](https://docs.microsoft.com/en-us/qsharp/api/qsharp/microsoft.quantum.intrinsic.ry) rotation to the first qubit.
#
# After this, we'll need to prepare the rest of the qubits in appropriate states depending on the state of the first qubit - state $|01\rangle$ if the first qubit is in state $|0\rangle$ and state $\frac{1}{\sqrt3} \big(|10\rangle + |11\rangle + |10\rangle \big)$ if the first qubit is in state $|1\rangle$. We can do this recursively using the same logic. Let's finish walking through this example in detail.
#
# The second qubit of the recursion follows similarly but depends on the first qubit. If the first qubit measures $|0\rangle$, then we want the second qubit to measure $|0\rangle$ with 100% probability, but if it measures $|1\rangle$, we want it to measure $|0\rangle$ with $\frac13$ probability and $|1\rangle$ with $\frac23$ probability. For this, we can do a controlled [`Ry`](https://docs.microsoft.com/en-us/qsharp/api/qsharp/microsoft.quantum.intrinsic.ry) rotation on the second qubit with the first qubit as control.
#
# The third qubit in this example will have three cases because it depends on the first two qubits; this follows naturally from the recursion.
#
# 1. If the first two qubits measure $|00\rangle$, then we need the third qubit to measure $|0\rangle$ with 100% probability.
# 2. If the first two qubits measure $|10\rangle$, then we need the third qubit to measure $|1\rangle$ with 100% probability.
# 3. If the first two qubits measure $|11\rangle$, then we need the third qubit to measure $|0\rangle$ with $\frac12$ probability and $|1\rangle$ with $\frac12$ probability. Just as with the second qubit, a controlled [`Ry`](https://docs.microsoft.com/en-us/qsharp/api/qsharp/microsoft.quantum.intrinsic.ry) rotation on the third qubit will accomplish this goal.
#
# > We will use [ControlledOnBitString](https://docs.microsoft.com/en-us/qsharp/api/qsharp/microsoft.quantum.canon.controlledonbitstring) operation to perform rotations depending on the state of several previous qubits.
# +
%kata T15_FourBitstringSuperposition_Test
open Microsoft.Quantum.Convert;
open Microsoft.Quantum.Math;
operation FourBitstringSuperposition (qs : Qubit[], bits : Bool[][]) : Unit {
FourBitstringSuperposition_Recursive(new Bool[0], qs, bits);
}
operation FourBitstringSuperposition_Recursive (currentBitString : Bool[], qs : Qubit[], bits : Bool[][]) : Unit {
// an array of bit strings whose columns we are considering begin with |0⟩
mutable zeroLeads = new Bool[][0];
// an array of bit strings whose columns we are considering begin with |1⟩
mutable oneLeads = new Bool[][0];
// the number of bit strings we're considering
let rows = Length(bits);
// the current position we're considering
let currentIndex = Length(currentBitString);
if (rows >= 1 and currentIndex < Length(qs)) {
// figure out what percentage of the bits should be |0⟩
for (row in 0..rows-1) {
if (bits[row][currentIndex]) {
set oneLeads = oneLeads + [bits[row]];
} else {
set zeroLeads = zeroLeads + [bits[row]];
}
}
// rotate the qubit to adjust coefficients based on the previous bit string
// for the first path through, when the bit string has zero length,
// the Controlled version of the rotation will perform a regular rotation
let theta = ArcCos(Sqrt(IntAsDouble(Length(zeroLeads)) / IntAsDouble(rows)));
(ControlledOnBitString(currentBitString, Ry))(qs[0 .. currentIndex - 1],
(2.0 * theta, qs[currentIndex]));
// call state preparation recursively based on the bit strings so far
FourBitstringSuperposition_Recursive(currentBitString + [false], qs, zeroLeads);
FourBitstringSuperposition_Recursive(currentBitString + [true], qs, oneLeads);
}
}
# -
# [Return to task 15 of the Superposition kata.](./Superposition.ipynb#superposition-of-two-bit-strings)
# ### <a name="wstate-on-2k-qubits"></a>Task 16**. W state on $2^k$ qubits.
#
# **Input:** $N = 2^k$ qubits in the $|0 \dots 0\rangle$ state.
#
# **Goal:** Change the state of the qubits to the [W state](https://en.wikipedia.org/wiki/W_state) - an equal superposition of $N$ basis states on $N$ qubits which have Hamming weight of 1.
#
# > For example, for $N = 4$ the required state is $\frac{1}{2}\big(|1000\rangle + |0100\rangle + |0010\rangle + |0001\rangle\big)$.
# ### Solution
#
# The problem becomes more manageable if broken down into the simplest cases and built up from there.
#
# 1. The smallest instance of the problem, $N = 1$, requires preparing $|W_1\rangle = |1\rangle$; this can be done trivially using an X gate.
#
# 2. The next instance, $N = 2$, requires preparing $|W_2\rangle = \frac{1}{\sqrt2}\big(|10\rangle + |01\rangle\big)$.
# It matches one of the Bell states we've seen earlier, but preparing it will be more interesting (and more useful for the next steps!) if we think of it in recursive terms.
# Let see how to express $|W_2\rangle$ in terms of $|W_1\rangle$:
#
# $$|W_2\rangle = \frac{1}{\sqrt2}\big(|10\rangle + |01\rangle\big) = \frac{1}{\sqrt2}\big(|W_1\rangle \otimes |0\rangle + |0\rangle \otimes |W_1\rangle\big)$$
#
# This representation suggests us a solution: "split" the starting state $|00\rangle$ in two terms, prepare $|W_1\rangle$ on the first qubit for the first term and on the second qubit - for the second term.
# To do this, we can again use an auxiliary qubit prepared in the $|+\rangle$ state and control the preparation of $|W_1\rangle$ state on the first or the second qubit based on the state of the auxiliary qubit:
#
# $$|0\rangle_{aux} |00\rangle_{reg} \overset{H}{\longrightarrow}
# \frac{1}{\sqrt2}(|0\rangle + |1\rangle)_{aux} \otimes |00\rangle_{reg} =
# \frac{1}{\sqrt2}(|0\rangle_{aux} |00\rangle_{reg} + |1\rangle_{aux} |00\rangle_{reg})
# \overset{CNOT_0}{\longrightarrow} \\ {\longrightarrow}
# \frac{1}{\sqrt2}(|0\rangle_{aux} |W_1\rangle|0\rangle_{reg} + |1\rangle_{aux} |00\rangle_{reg})
# \overset{CNOT_1}{\longrightarrow} \\ {\longrightarrow}
# \frac{1}{\sqrt2}(|0\rangle_{aux} |W_1\rangle|0\rangle_{reg} + |1\rangle_{aux} |0\rangle|W_1\rangle_{reg})
# $$
#
# > The auxiliary qubit is now entangled with the rest of the qubits, so we can't simply reset it without it affecting the superposition we have prepared using it.
#
# The last step to bring the register to the desired state is to uncompute the auxiliary qubit for the term $|1\rangle_{aux} |0\rangle|W_1\rangle_{reg}$ (the other term already has it in state $|0\rangle$).
#
# To do this, we need to consider the explicit expression of the state $|0\rangle|W_1\rangle = |01\rangle$.
# Similarly to the previous tasks, we'll uncompute the auxiliary qubit for this term by using a controlled X gate, with the auxiliary qubit as the target and the main register in the $|01\rangle$ state as a control.
# This will make sure that the gate is applied only for this term and not for any others.
#
# The last step can be simplified to use fewer qubits as controls: we can use just the second qubit of the main register in state $|1\rangle$ as control, since we know that if the second qubit is in state $|1\rangle$, the first one has to be in state $|0\rangle$ (we don't need to use both of them as the control pattern).
#
# 3. If we take this one step further, to $N = 4$, we'll see that the same recursive logic can be applied to the larger and larger sizes of the problem. Indeed,
#
# $$|W_4\rangle = \frac{1}{2}\big(|1000\rangle + |0100\rangle + |0010\rangle + |0001\rangle\big) = \\
# = \frac{1}{\sqrt2} \big(\frac{1}{\sqrt2}(|10\rangle + |01\rangle) \otimes |00\rangle + |00\rangle \otimes \frac{1}{\sqrt2}(|10\rangle + |01\rangle) = \\
# = \frac{1}{\sqrt2} \big(|W_2\rangle \otimes |00\rangle + |00\rangle \otimes |W_2\rangle\big)
# $$
#
# We can use the same approach for this case: prepare an auxiliary qubit in $|+\rangle$ state and use it to control preparation of $W_2$ state on the first and the second half of the register.
# The last step will be uncomputing the $|1\rangle$ state of the auxiliary qubit using two controlled X gates with each of the qubits of the second half of the register in state $|1\rangle$ as controls.
#
# The same recursive approach can be generalized for arbitrary powers of 2 as the register size.
# +
%kata T16_WState_PowerOfTwo_Test
operation WState_PowerOfTwo (qs : Qubit[]) : Unit is Adj+Ctl {
let N = Length(qs);
if (N == 1) {
// base of recursion: |1⟩
X(qs[0]);
} else {
let K = N / 2;
using (anc = Qubit()) {
H(anc);
(ControlledOnInt(0, WState_PowerOfTwo))([anc], qs[0 .. K - 1]);
(ControlledOnInt(1, WState_PowerOfTwo))([anc], qs[K .. N - 1]);
for (i in K .. N - 1) {
CNOT(qs[i], anc);
}
}
}
}
# -
# This implementation of the recursion requires $\log_2 N = k$ extra qubits allocated for controlling the preparation (one per level of recursion).
# We can modify our approach to use just one extra qubit at a time.
#
# To do this, let's notice that to prepare $|W_{N}\rangle$ we need to prepare the $|W_{N-1}\rangle$ state on half of the qubits for both states of the auxiliary qubit, the difference is just in which half of the register we're using.
# This means that we can prepare the $|W_{N-1}\rangle$ state on the first half of the qubits, and use an auxiliary qubit in superposition to control SWAP-ing the first half of the register with the second half.
# The uncomputation of the auxiliary qubit happens in the same way as in the first approach.
# +
%kata T16_WState_PowerOfTwo_Test
operation WState_PowerOfTwo (qs : Qubit[]) : Unit is Adj+Ctl {
let N = Length(qs);
if (N == 1) {
// base of recursion: |1⟩
X(qs[0]);
} else {
let K = N / 2;
WState_PowerOfTwo(qs[0 .. K - 1]);
using (anc = Qubit()) {
H(anc);
for (i in 0 .. K - 1) {
Controlled SWAP([anc], (qs[i], qs[i + K]));
}
for (i in K .. N - 1) {
CNOT(qs[i], anc);
}
}
}
}
# -
# [Return to task 16 of the Superposition kata.](./Superposition.ipynb#wstate-on-2k-qubits)
# ### <a name="wstate-on-arbitray-number-of-qubits"></a>Task 17**. W state on an arbitrary number of qubits.
#
# **Input:** $N$ qubits in the $|0 \dots 0\rangle$ state ($N$ is not necessarily a power of 2).
#
# **Goal:** Change the state of the qubits to the [W state](https://en.wikipedia.org/wiki/W_state) - an equal superposition of $N$ basis states on $N$ qubits which have Hamming weight of 1.
#
# > For example, for $N = 3$ the required state is $\frac{1}{\sqrt{3}}\big(|100\rangle + |010\rangle + |001\rangle\big)$.
# ### Solution
#
# This problem allows a variety of solutions that rely on techniques from arbitrary rotations to recursion to postselection.
#
# The first approach we will describe relies on performing a sequence of controlled rotations.
#
# To prepare a weighted superposition $\cos \theta |0\rangle + \sin \theta |1\rangle$ on a single qubit, we need to start with the $|0\rangle$ state and apply the [Ry gate](https://docs.microsoft.com/qsharp/api/qsharp/microsoft.quantum.intrinsic.ry) to it with the angle parameter equal to $2 \theta$.
# We'll apply the Ry gate with angle $2 \theta_1 = 2\arcsin \frac{1}{\sqrt{N}}$ to the first qubit of the register to prepare the following state:
#
# $$(\cos \theta_1 |0\rangle + \sin \theta_1 |1\rangle) \otimes |0 \dots 0\rangle = \frac{1}{\sqrt{N}}|10 \dots 0\rangle + \frac{\sqrt{N-1}}{\sqrt{N}}|00 \dots 0\rangle $$
#
# The first term $\frac{1}{\sqrt{N}}|10 \dots 0\rangle$ already matches the first term of the $|W_N\rangle$ state; now we need to convert the second term $\frac{\sqrt{N-1}}{\sqrt{N}}|00 \dots 0\rangle$ into the rest of the $|W_N\rangle$ terms.
#
# To prepare a term that matches the second term of the $|W_N\rangle$ state, we can apply another Ry gate to the term $|00 \dots 0\rangle$, this time to the second qubit, with an angle $2 \theta_2 = 2\arcsin \frac{1}{\sqrt{N-1}}$.
# To make sure it doesn't affect the term that we're already happy with, we will apply a controlled version of the Ry gate, with the first qubit of the register in state $|0\rangle$ as control.
# This will change our state to
#
# $$\frac{1}{\sqrt{N}}|10 \dots 0\rangle + \frac{\sqrt{N-1}}{\sqrt{N}} |0\rangle \otimes (\cos \theta_2 |0\rangle + \sin \theta_2 |1\rangle) \otimes |0 \dots 0\rangle = \\
# = \frac{1}{\sqrt{N}}|10 \dots 0\rangle + \frac{\sqrt{N-1}}{\sqrt{N}} \frac{1}{\sqrt{N-1}} |010 \dots 0\rangle + \frac{\sqrt{N-1}}{\sqrt{N}} \frac{\sqrt{N-2}}{\sqrt{N-1}} |000 \dots 0\rangle$$
#
# Now we have two terms that match the terms of the $|W_N\rangle$ state, and need to convert the third term $\frac{\sqrt{N-2}}{\sqrt{N}}|00 \dots 0\rangle$ into the rest of terms.
#
# We will keep going like this, preparing one term of the $|W_N\rangle$ state at a time, until the rotation on the last qubit will be an X gate, controlled on all previous qubits being in the $|0 \dots 0\rangle$ state.
# +
%kata T17_WState_Arbitrary_Test
open Microsoft.Quantum.Convert;
open Microsoft.Quantum.Math;
operation WState_Arbitrary (qs : Qubit[]) : Unit {
let N = Length(qs);
Ry(2.0 * ArcSin(Sqrt(1.0/IntAsDouble(N))), qs[0]);
for (i in 1 .. N-1) {
(ControlledOnInt(0, Ry(2.0 * ArcSin(Sqrt(1.0/IntAsDouble(N - i))), _)))(qs[0 .. i-1], qs[i]);
}
}
# -
# We can express the same sequence of gates using recursion, if we notice that
#
# $$|W_N\rangle = \frac{1}{\sqrt{N}}|10 \dots 0\rangle + \frac{\sqrt{N-1}}{\sqrt{N}}|0\rangle \otimes |W_{N-1}\rangle$$
#
# The first step of the solution would still be applying the Ry gate with angle $2 \theta_1 = 2\arcsin \frac{1}{\sqrt{N}}$ to the first qubit of the register to prepare the following state:
#
# $$\frac{1}{\sqrt{N}}|10 \dots 0\rangle + \frac{\sqrt{N-1}}{\sqrt{N}}|00 \dots 0\rangle $$
#
# But we would express the rest of the controlled rotations as the operation that prepares the $|W_{N-1}\rangle$ state, controlled on the $|0\rangle$ state of the first qubit.
#
# > Note that you don't have to implement the controlled version of the gate yourself; it is sufficient to add `is Adj+Ctl` to the signature of the operation `WState_Arbitrary` to specify that controlled variant has to be generated automatically.
# +
%kata T17_WState_Arbitrary_Test
open Microsoft.Quantum.Convert;
open Microsoft.Quantum.Math;
operation WState_Arbitrary (qs : Qubit[]) : Unit is Adj+Ctl {
let N = Length(qs);
Ry(2.0 * ArcSin(Sqrt(1.0/IntAsDouble(N))), qs[0]);
if (N > 1) {
(ControlledOnInt(0, WState_Arbitrary))(qs[0 .. 0], qs[1 ...]);
}
}
# -
# The last approach we will describe uses a technique that is completely different from the ones you've seen before in this kata: postselection.
#
# > Note that this approach requires familiarity with measurements of quantum systems, in particular with the effect of partial measurement on a multi-qubit system; you might want to return to it after you've covered the relevant tutorials and katas.
#
# *Coming up soon...*
# [Return to task 17 of the Superposition kata.](./Superposition.ipynb#wstate-on-arbitray-number-of-qubits)
| Superposition/Workbook_Superposition_Part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Copyright 2016 The TensorFlow Authors. All Rights Reserved.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# http://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# ==============================================================================
"""Contains a factory for building various models."""
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import tensorflow as tf
from preprocessing import cifarnet_preprocessing
from preprocessing import inception_preprocessing
from preprocessing import lenet_preprocessing
from preprocessing import vgg_preprocessing
from preprocessing import mobilenet_preprocessing
from preprocessing import mobilenetdet_preprocessing
slim = tf.contrib.slim
def get_preprocessing(name, is_training=False):
"""Returns preprocessing_fn(image, height, width, **kwargs).
Args:
name: The name of the preprocessing function.
is_training: `True` if the model is being used for training and `False`
otherwise.
Returns:
preprocessing_fn: A function that preprocessing a single image (pre-batch).
It has the following signature:
image = preprocessing_fn(image, output_height, output_width, ...).
Raises:
ValueError: If Preprocessing `name` is not recognized.
"""
preprocessing_fn_map = {
'cifarnet': cifarnet_preprocessing,
'inception': inception_preprocessing,
'inception_v1': inception_preprocessing,
'inception_v2': inception_preprocessing,
'inception_v3': inception_preprocessing,
'inception_v4': inception_preprocessing,
'inception_resnet_v2': inception_preprocessing,
'lenet': lenet_preprocessing,
'resnet_v1_50': vgg_preprocessing,
'resnet_v1_101': vgg_preprocessing,
'resnet_v1_152': vgg_preprocessing,
'resnet_v2_50': vgg_preprocessing,
'resnet_v2_101': vgg_preprocessing,
'resnet_v2_152': vgg_preprocessing,
'vgg': vgg_preprocessing,
'vgg_a': vgg_preprocessing,
'vgg_16': vgg_preprocessing,
'vgg_19': vgg_preprocessing,
'mobilenet': mobilenet_preprocessing,
'mobilenetdet': mobilenetdet_preprocessing
}
if name not in preprocessing_fn_map:
raise ValueError('Preprocessing name [%s] was not recognized' % name)
def preprocessing_fn(image, output_height, output_width, **kwargs):
return preprocessing_fn_map[name].preprocess_image(
image, output_height, output_width, is_training=is_training, **kwargs)
return preprocessing_fn
| preprocessing/preprocessing_factory.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:kaggle]
# language: python
# name: conda-env-kaggle-py
# ---
# # Table of Contents
# * [Intro](#Intro)
# * [Time-Series](#Time-Series)
# * [Moving Average](#Moving-Average)
# * [Filling missing values](#Filling-missing-values)
# * [Interpolation](#Interpolation)
# * [Stationarity [TOFIX]](#Stationarity-[TOFIX])
# *
# * [Check on more complex example Time Series](#Check-on-more-complex-example-Time-Series)
# * [Correlation](#Correlation)
# * [Arima](#Arima)
#
# # Intro
# Notebook that explores time-series and techniques to analyze them.
#
# Resources:
# * [Data Analysis with Open Source Tools](http://shop.oreilly.com/product/9780596802363.do)
# * [Think Stats 2e - <NAME>](http://greenteapress.com/wp/think-stats-2e/)
# ## Time-Series
# "A time-series is sequence of measurements from a system that varies in time"
#
# A time-series is generally decomposed in three major components:
# * **Trend**: persistent change along time.
# * **Seasonality**: regular periodic variation. There can be multiple seasonalities, and each can span different time-frames (by day, week, month, year, etc.)
# * **Noise**: random variation
# +
# %matplotlib notebook
import numpy as np
import seaborn as sns
import pandas as pd
sns.set_context("paper")
# -
# # Moving Average
# Moving Averages (also rolling/running average or moving/running mean) is a technique that helps to extract the trend from a series. It reduces noise and decreases impact of outliers. Is consists in dividing the series in overlapping windows of fixed size $N$, and for each considering the average value. It follows that the first $N-1$ values will be undefined, given that they don't have enough predecessors to compute the average.
#
# **Exponentially-Weighted Moving Average (EWMA)** is an alternative that gives more importance to recent values.
# rolling mean basic example
series = np.arange(10)
pd.Series(series).rolling(3).mean()
# ewma basic example
series = np.arange(10)
pd.Series(series).ewm(3).mean()
# ewm on partial long series of 0s
series = [1,1,1,1,1,1,1,0,0,0,0,0,0,0,0,0]
pd.Series(series).ewm(span=2).mean()
pd.Series(series).ewm(span=2).mean().plot()
# # Filling missing values
# Basic Pandas methods:
# * pad/ffill: fill values forward
# * bfill/backfill: fill values backward
# Random arrays to play with
a = np.arange(20)
b = a*a
b_empty = np.array(a*a).astype('float')
# Add missing values and get a Pandas Series
b_empty[[0, 5, 6, 15]] = np.nan
c = pd.Series(b_empty)
# Visualize how filling method works
fig, axes = sns.plt.subplots(2)
sns.pointplot(np.arange(20), c, ax=axes[0])
sns.pointplot(np.arange(20), c.fillna(method='bfill'), ax=axes[1])
sns.plt.show()
# ## Interpolation
# http://pandas.pydata.org/pandas-docs/stable/generated/pandas.Series.interpolate.html
#
# 'linear' ignores the index, ‘time’: interpolation works on daily and higher resolution data to interpolate given length of interval
# # Stationarity [TOFIX]
# [Link](https://www.analyticsvidhya.com/blog/2016/02/time-series-forecasting-codes-python/)
df = pd.read_csv("time_series.csv")
df['datetime'] = pd.to_datetime(df['datetime'])
df.set_index("datetime", inplace=True)
df.fillna(method='pad', axis=0, inplace=True)
df.head()
# +
#Determing rolling statistics
rolmean = pd.rolling_mean(new_df, window=12)
rolstd = pd.rolling_std(new_df, window=12)
#Plot rolling statistics:
sns.plt.plot(new_df, color='blue',label='Original')
sns.plt.plot(rolmean, color='red', label='Rolling Mean')
sns.plt.plot(rolstd, color='black', label = 'Rolling Std')
sns.plt.show()
# -
new_df = df.copy()
new_df['val'] = new_df['val'] - pd.ewma(new_df, halflife=12)['val']
new_df['val'] = new_df['val'] - pd.ewma(new_df, halflife=12)['val']
new_df.plot()
sns.plt.show()
# ### Check on more complex example Time Series
df = pd.read_csv("time_series.csv")
df['datetime'] = pd.to_datetime(df['datetime'])
df.set_index("datetime", inplace=True)
df.head()
df.plot()
sns.plt.show()
df.fillna(method='pad', axis=0).plot()
sns.plt.show()
null_indexes = [i for i, isnull in enumerate(pd.isnull(df['val'].values)) if isnull]
# missing_values_correct
y = [32.69,32.15,32.61,29.3,28.96,28.78,31.05,29.58,29.5,30.9,31.26,31.48,29.74,29.31,29.72,28.88,30.2,27.3,26.7,27.52]
filled = df['val'].interpolate(method='time').values
predict = filled[null_indexes]
len(predict)==len(y)
d = sum([abs((y[i]-predict[i])/y[i]) for i in range(len(y))])
d
# # Correlation
# [Link](https://anomaly.io/understand-auto-cross-correlation-normalized-shift)
#
# There exists different methods to analyze correlation for time-series. If we compare two different time series we are talking about **cross-correlation**, while in **auto-correlation** a time-series is compared with itself (can detect seasonality). Both of the previous mentioned categories can use normalization (useful for example when series characterized by different scales, also good for values = zero).
#
# Correlation between two time-series $y$ and $x$ is defined as
#
# $$ corr(x, y) = \sum_{n=0}^{n-1} x[n]*y[n] $$
#
# while normalized correlation is defined as
#
# $$norm\_corr(x,y)=\dfrac{\sum_{n=0}^{n-1} x[n]*y[n]}{\sqrt{\sum_{n=0}^{n-1} x[n]^2 * \sum_{n=0}^{n-1} y[n]^2}}$$
#
# For auto-correlation we shift the time-series by an interval called **lag**, and then compare the shifted version with the original one to understand the strength of the correlation (process sometime also called serial-correlation, especially when lag=1).
# The idea is that a series values are not random independent event, but should have some level of dependency with preceding values. This dependency is the pattern we are trying to discover.
#
# Suggestions: check correlation after removing the trend. Understand the seasonality more appropriate for your case.
a = np.array([1,2,-2,4,2,3,1,0])
b = np.array([2,3,-2,3,2,4,1,-1])
c = np.array([-2,0,4,0,1,1,0,-2])
print("a and b correlate value = {}".format(np.correlate(a, b)[0]))
print("a and c correlate value = {}".format(np.correlate(a, c)[0]))
def normalized_cross_correlation(a, v):
# cross-correlation is simply the dot product of our arrays
cross_cor = np.dot(a, v)
norm_term = np.sqrt(np.sum(a**2) * np.sum(v**2))
return cross_cor/norm_term
normalized_cross_correlation(a, c)
print("a and a/2 correlate value = {}".format(np.correlate(a, a/2)[0]))
print("a and a/2 normalized correlate value = {}".format(normalized_cross_correlation(a, a/2)))
# # Arima
| miscellaneous/Time Series.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## TODO
# * Add O2C and C2O seasonality
# * Look at diff symbols
# * Look at fund flows
# ## Key Takeaways
# * ...
#
#
# In the [first post](sell_in_may.html) of this short series, we covered several seasonality patterns for large cap equities (i.e, SPY), most of which continue to be in effect.
#
# The findings of that exercise sparked interest in what similar seasonal patterns may exist in other asset classes. This post will pick up where that post left off, looking at "risk-off" assets which exhibit low (or negative) correlation to equities.
#
#
# +
## Replace this section of imports with your preferred
## data download/access interface. This calls a
## proprietary set of methods (ie they won't work for you)
import sys
sys.path.append('/anaconda/')
import config
sys.path.append(config.REPO_ROOT+'data/')
from prices.eod import read
####### Below here are standard python packages ######
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
from IPython import display
import seaborn as sns
from IPython.core.display import HTML,Image
## Load Data
symbols = ['SPY','IWM','AGG','LQD','IEF','MUB','GLD']
#symbols = ['SPY','IWM','AGG','LQD','JNK','IEF']
prices = read.get_symbols_close(symbols,adjusted=True)
returns = prices.pct_change()
log_ret = np.log(prices).diff()
# -
# ### Month-of-year seasonality
#
# Again, we'll start with month-of-year returns for several asset classes. Note that I'm making use of the seaborn library's excellent `clustermap()` method to both visually represent patterns in asset classes _and_ to group the assets by similarity (using Euclidean distance between the average monthly returns vectors of each column).
#
# _Note that the values plotted are z-score values (important for accurate clustering)_.
# +
by_month = log_ret.resample('BM').sum()
by_month[by_month==0.0] = None
# because months prior to fund launch are summed to 0.0000
avg_monthly = by_month.groupby(by_month.index.month).mean()
sns.clustermap(avg_monthly[symbols],row_cluster=False,z_score=True, metric='euclidean',\
cmap=sns.diverging_palette(10, 220, sep=20, n=7))
## Notes:
# should use either z_score =True or standard_scale = True for accurate clustering
# Uses Euclidean distance as metric for determining cluster
# -
# Clearly, the seasonal patterns we saw in the [last post](sell_in_may.html) do not generalize across all instruments - which is a very good thing! IWM (small cap equities) do more or less mimic the SPY patterns, but the "risk-off" assets generally perform well in the summer months of July and August, when equities had faltered.
#
# We might consider a strategy of shifting from risk-on (e.g., SPY) to risk-off (e.g., IEF) for June to September.
rotation_results = pd.Series(index=avg_monthly.index)
rotation_results.loc[[1,2,3,4,5,10,11,12]] = avg_monthly['SPY']
rotation_results.loc[[6,7,8,9]] = avg_monthly['IEF']
#
print("Returns:")
print(avg_monthly.SPY.sum())
print(rotation_results.sum())
print()
print("Sharpe:")
print(avg_monthly.SPY.sum()/(by_month.std()['SPY']*12**0.5))
print(rotation_results.sum()/(rotation_results.std()*12**0.5))
avg_monthly.SPY.std()*12**0.5
#
# Next, I'll plot the same for day-of-month.
avg_day_of_month = log_ret.groupby(log_ret.index.day).mean()
sns.clustermap(avg_day_of_month[symbols],row_cluster=False,z_score= True,metric='euclidean',\
cmap=sns.diverging_palette(10, 220, sep=20, n=7))
# This is a bit messy, but I think the dominant pattern is weakness within all "risk-off" assets (treasurys, etc...) for the first 1/3 to 1/2 of the month, followed by a very strong end of month rally.
#
# Finally, plot a clustermap for day-of-week:
avg_day_of_week = log_ret.groupby(log_ret.index.weekday+1).mean()
sns.clustermap(avg_day_of_week[symbols],row_cluster=False,z_score= True,metric='euclidean',\
cmap=sns.diverging_palette(10, 220, sep=20, n=7))
# Again, a bit messy. However, the most consistent pattern is "avoid Thursday" for risk-off assets like AGG, LQD, and IEF. Anyone with a hypothesis as to why this might be, please do share!
#
#
# ### Observations
# * Clusters form about as you'd expect. The "risk-off" assets like Treasurys (IEF), munis (MUB), gold (GLD), and long volatility (VXX) tend to cluster together. The "risk-on" assets like SPY, EEM, IXUS, and JNK tend to cluster together.
# * Risk-off assets (Treasurys etc...) appear to follow the opposite of "sell in May", with weakness in November and December, when SPY and related were strongest.
# * Within day-of-month, there are some _very_ strong patterns for fixed income, with negative days at the beginning of month and positive days at end of month.
# * Day of week shows very strong clustering of risk-off assets (outperform on Fridays). There's an interesting clustering of underperformance on Mondays. This may be a false correlation since some of these funds have much shorter time histories than others and may be reflecting that
# +
risk_off_symbols = ['IEF','MUB','AGG','LQD']
df = log_ret[symbols_1].mean(axis=1).dropna().to_frame(name='pct_chg')
by_month = df.resample('BM').sum()
by_month['month'] = by_month.index.month
title='Avg Log Return (%): by Calendar Month \nfor Risk-off Symbols {}'.format(risk_off_symbols)
s = (by_month.groupby('month').pct_chg.mean()*100)
my_colors = ['r','r','r','r','g','g','g','g','g','g','r','r',]
ax = s.plot(kind='bar',color=my_colors,title=title)
ax.axhline(y=0.00, color='grey', linestyle='--', lw=2)
# -
# Wow, maybe there's some truth to this myth! It appears that there is a strong difference between the summer months (June to September) and the rest.
#
# From the above chart, it appears than we'd be well advised to sell on June 1st and buy back on September 30th. However, to follow the commonly used interpretation of selling on May 1st and repurchasing on Oct 31st. I'll group the data into those two periods and calculate the monthly average:
# +
by_month['season'] = None
by_month.loc[by_month.month.between(5,10),'season'] = 'may_oct'
by_month.loc[~by_month.month.between(5,10),'season'] = 'nov_apr'
(by_month.groupby('season').pct_chg.mean()*100).plot.bar\
(title='Avg Monthly Log Return (%): \nMay-Oct vs Nov_Apr (1993-present)'\
,color='grey')
# -
# A significant difference. The "winter" months are more than double the average return of the summer months. But has this anomaly been taken out of the market by genius quants and vampire squid? Let's look at this breakout by year:
# Of these, the most interesting patterns, to me, are the day-of-week and day-of-month cycles.
#
# ### Day of Week
# I'll repeat the same analysis pattern as developed in the prior post (["Sell in May"](sell_in_may.html)), using a composite of four generally "risk-off" assets. You may choose create composites differently.
# +
risk_off_symbols = ['IEF','MUB','AGG','LQD']
df = log_ret[risk_off_symbols].mean(axis=1).dropna().to_frame(name='pct_chg')
by_day = df
by_day['day_of_week'] = by_day.index.weekday+ 1
ax = (by_day.groupby('day_of_week').pct_chg.mean()*100).plot.bar\
(title='Avg Daily Log Return (%): by Day of Week \n for {}'.format(risk_off_symbols),color='grey')
plt.show()
by_day['part_of_week'] = None
by_day.loc[by_day.day_of_week ==4,'part_of_week'] = 'thurs'
by_day.loc[by_day.day_of_week !=4,'part_of_week'] = 'fri_weds'
(by_day.groupby('part_of_week').pct_chg.mean()*100).plot.bar\
(title='Avg Daily Log Return (%): Mon vs Tue-Fri \n for {}'.format(risk_off_symbols)\
,color='grey')
title='Avg Daily Log Return (%) by Part of Week\nFour Year Moving Average\n for {}'.format(risk_off_symbols)
by_day['year'] = by_day.index.year
ax = (by_day.groupby(['year','part_of_week']).pct_chg.mean().unstack().rolling(4).mean()*100).plot()
ax.axhline(y=0.00, color='grey', linestyle='--', lw=2)
ax.set_title(title)
# -
# The "avoid Thursday" for risk-off assets seemed to be remarkably useful until about 4 years ago, when it ceased to work. I'll call this one busted. Moving on to day-of-month, and following the same grouping and averaging approach:
risk_off_symbols = ['IEF','MUB','AGG','LQD']
by_day = log_ret[risk_off_symbols].mean(axis=1).dropna().to_frame(name='pct_chg')
by_day['day_of_month'] = by_day.index.day
title='Avg Daily Log Return (%): by Day of Month \nFor: {}'.format(symbols_1)
ax = (by_day.groupby('day_of_month').pct_chg.mean()*100).plot.bar(xlim=(1,31),title=title,color='grey')
ax.axhline(y=0.00, color='grey', linestyle='--', lw=2)
# Here we see the same pattern as appeared in the clustermap. I wonder if the end of month rally is being driven by the ex-div date, which I believe is usually the 1st of the month for these funds.
#
# _Note: this data is dividend-adjusted so there is no valid reason for this - just dividend harvesting and behavioral biases, IMO._
# +
by_day['part_of_month'] = None
by_day.loc[by_day.index.day <=10,'part_of_month'] = 'first_10d'
by_day.loc[by_day.index.day >10,'part_of_month'] = 'last_20d'
(by_day.groupby('part_of_month').pct_chg.mean()*100).plot.bar\
(title='Avg Daily Log Return (%): \nDays 1-10 vs 11-31\nfor risk-off assets {}'.format(risk_off_symbols)\
,color='grey')
title='Avg Daily Log Return (%) \nDays 1-10 vs 11-31\nfor risk-off assets {}'.format(risk_off_symbols)
by_day['year'] = by_day.index.year
ax = (by_day.groupby(['year','part_of_month']).pct_chg.mean().unstack().rolling(4).mean()*100).plot(title=title)
ax.axhline(y=0.00, color='grey', linestyle='--', lw=2)
# -
# In contrast to the day-of-week anomaly, this day-of-month pattern seems to hold extremely well. It's also an extremely tradeable anomaly, considering that it requires only one round-trip per month.
baseline = by_day.resample('A').pct_chg.sum()
only_last_20 = by_day[by_day.part_of_month=='last_20d'].resample('A').pct_chg.sum()
pd.DataFrame({'baseline':baseline,'only_last_20':only_last_20}).plot.bar()
print(pd.DataFrame({'baseline':baseline,'only_last_20':only_last_20}).mean())
# Going to cash in the first 10 days of each month actually _increased_ annualized returns (log) by about 0.60%, while simultaneously lowering capital employed and volatility of returns. Of the seasonality anomalies we've reviewed in this post and the previous, this appears to be the most robust and low risk.
#
#
#
# ## Conclusion
# ...
#
# If the future looks anything like the past (insert standard disclaimer about past performance...) then rules of thumb might be:
# * Sell on Labor Day and buy on Halloween - especially do this on election years! This assumes that you've got a productive use for the cash!
# * Do your buying at Friday's close, do your selling at Wednesday's close
# * Maximize your exposure at the end/beginning of months and during the early-middle part of the month, lighten up.
# * Remember that, in most of these anomalies, _total_ return would decrease by only participating in part of the market since any positive return is better than sitting in cash. Risk-adjusted returns would be significantly improved by only participating in the most favorable periods. It's for each investor to decide what's important to them.
#
# I had intended to extend this analysis to other asset classes, but will save that for a future post. I'd like to expand this to small caps, rest-of-world developed/emerging, fixed income, growth, value, etc...
#
#
# ### One last thing...
#
# If you've found this post useful, please follow [@data2alpha](https://twitter.com/data2alpha) on twitter and forward to a friend or colleague who may also find this topic interesting.
#
# Finally, take a minute to leave a comment below. Share your thoughts on this post or to offer an idea for future posts. Thanks for reading!
| content-draft/sell_in_may_part2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: spatial-networks
# language: python
# name: python3
# ---
# +
#Load libs
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import os
from pathlib import Path
from tysserand import tysserand as ty
from PIL import Image
Image.MAX_IMAGE_PIXELS = 1000000000
from PIL import Image, ImageOps
import fcsparser
from os import listdir
from os.path import isfile, join
#set up working dir
import sys
sys.path.extend([
'../tysserand/tysserand',
'../mosna',
])
import seaborn as sns
from time import time
import copy
from skimage import color
import matplotlib as mpl
import napari
from sklearn.preprocessing import StandardScaler
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC, LinearSVC
from sklearn.metrics import accuracy_score
from sklearn.model_selection import cross_validate, GridSearchCV, RandomizedSearchCV
from scipy.stats import loguniform
import umap
# if not installed run: conda install -c conda-forge umap-learn
import hdbscan
from sklearn.cluster import OPTICS, cluster_optics_dbscan
from skimage import io
from scipy.stats import ttest_ind # Welch's t-test
from scipy.stats import mannwhitneyu # Mann-Whitney rank test
from scipy.stats import ks_2samp # Kolmogorov-Smirnov statistic
sys.path.append("/home/mouneem/mosna/")
from tysserand import tysserand as ty
from mosna import mosna
import glob
import re
# +
# GENERATE CSV FILE OF EACH LAYERS
pathC1 = '/mnt/SERVER-CRCT-STORAGE/CRCT_Imagin/CORDELIER Pierre/HaloData/21-003.IMMCORE.C2v1/Halo archive 2021-12-13 16-38 - v3.3.2541/ObjectData/'
csvs = [f for f in listdir(pathC1) if isfile(join(pathC1, f))]
for csvFile in csvs:
csvData = pd.read_csv(pathC1 + csvFile)
print(csvData.columns)
csvData[['x']] = ( csvData[['XMax']] + csvData[['XMax']] ) / 2
csvData[['y']] = ( csvData[['YMax']] + csvData[['YMax']] ) / 2
# +
img_path = '/home/mouneem/tysserand/CORDELIER_PIERRE/C2v1/'
edg_path = '/home/mouneem/tysserand/CORDELIER_PIERRE/edg/'
coords_path = '/home/mouneem/tysserand/CORDELIER_PIERRE/CRDS/'
nets_path = '/home/mouneem/tysserand/CORDELIER_PIERRE/nets/'
imgs = [f for f in listdir(img_path) if isfile(join(img_path, f))]
coords = [f for f in listdir(coords_path) if isfile(join(coords_path, f))]
edges = [f for f in listdir(edg_path) if isfile(join(edg_path, f))]
# -
print(len(edges))
# +
for fileedg in edges[5:7]:
print(fileedg)
filecoords = "coords."+".".join(fileedg.split(".")[1:])
filenet = ""+".".join(fileedg.split(".")[1:])
pattern = filenet.split("czi")[0]
edg = pd.read_csv(edg_path + fileedg , header = None)
crd = pd.read_csv(coords_path + filecoords , header = None)
plt.figure(figsize=(30, 30), dpi=80)
crd=crd.drop([0,1])
edg=edg.drop([0,1])
img_found = pattern+"jpg" in imgs
if img_found:
img = plt.imread ( img_path + pattern+"jpg")
fig, ax = ty.showim(img, figsize=(30, 30))
else:
fig, ax = plt.subplots(1,1,figsize=(30, 30))
ax.scatter(crd.iloc[:,1], crd.iloc[:,2], c ="blue")
Xs = list(edg.iloc[:,1 ])
Ys = list(edg.iloc[:,1 ])
print(Xs)
print(type(edg))
[x0, y0], [x1, y1] = coords[edg]
ax.plot([x0, x1], [y0, y1], c=cmap(dist), zorder=0, alpha=alpha_edges, linewidth=linewidth)
#fig.show()
fig.savefig(nets_path+filenet+'.png')
# +
mosna_path = '/home/mouneem/tysserand/CORDELIER_PIERRE/Mixmat/'
mosna_output = str("/home/mouneem/tysserand/CORDELIER_PIERRE/mosna_output/")
mosnas = [f for f in listdir(mosna_path) if isfile(join(mosna_path, f))]
for mosnafile in mosnas[:2]:
mixmat = pd.read_csv(mosna_path + mosnafile ,index_col=0 )
print(mixmat)
title = "Assortativity by cell types:"
print(title)
fig, ax = plt.subplots(figsize=(9, 6))
sns.heatmap(mixmat, center=0, cmap="vlag", annot=True, linewidths=.5, ax=ax)
plt.xticks(rotation=30, ha='right');
# plt.xticks(rotation=30, ha='right', fontsize=20);
# plt.yticks(fontsize=20);
plt.savefig(mosna_output + mosnafile+"assortativity.png", bbox_inches='tight', facecolor='white')
# -
mosna_path = '/home/mouneem/tysserand/CORDELIER_PIERRE/Mixmat/'
# +
Layer1 = '/home/mouneem/tysserand/CORDELIER_PIERRE/tummors/'
tummors = [ ".".join(f.split(".")[:-3]) for f in listdir(Layer1) if isfile(join(Layer1, f))]
mosnas = [f for f in listdir(mosna_path) if isfile(join(mosna_path, f))]
mosna_output = str("/home/mouneem/tysserand/CORDELIER_PIERRE/mosna_output/")
print(file)
FullMatrix = MAT
for mosnafile in mosnas:
file = ".".join(mosnafile.split(".")[1:-4])
if file in tummors:
print(file, mosnafile)
mixmat = pd.read_csv(mosna_path + mosnafile ,index_col=0 )
print(mixmat)
keep = np.triu(np.ones(mixmat.shape)).astype('bool').reshape(mixmat.size)
MAT = pd.DataFrame(mixmat.stack())
MAT.to_csv('out.csv')
MAT = pd.read_csv('out.csv')
MAT.columns = ['X','Y','Value']
di = {'C1': "Cancer", "C2": 'CD8 T-Cell', 'C3' : 'CD4 T-Cell', 'C4':'B Cell', 'Other':"Other",'C5':'CD3+CD20+' }
MAT = MAT.replace( {"Y": di })
MAT = MAT.replace( {"X": di })
MAT["comb"] = MAT["X"].astype(str) + " / " + MAT["Y"].astype(str)
MAT["Value"]=(MAT["Value"]-MAT["Value"].min())/(MAT["Value"].max()-MAT["Value"].min())
MAT['sample'] = file
FullMatrix = FullMatrix.append(pd.DataFrame(data = MAT))
print(FullMatrix)
# -
# +
FullMatrix.to_csv('FullMatrix.csv')
FullMatrix = FullMatrix[['Value', 'comb', 'sample']]
FullMatrix2 = FullMatrix[ FullMatrix['comb'].isin(['Other / Other' , 'Other / CD8 T-Cell' , 'Other / CD4 T-Cell', 'Other / Cancer',
'CD8 T-Cell / CD8 T-Cell' , 'CD8 T-Cell / CD4 T-Cell', 'CD8 T-Cell / Cancer',
'CD4 T-Cell / CD4 T-Cell', 'CD4 T-Cell / Cancer',
'Cancer / Cancer', ]) ]
FullMatrix2.index = FullMatrix2[['sample']]
print(FullMatrix2)
Matrix = FullMatrix2.pivot_table(index=["sample"],
columns='comb',
values='Value')
plt.figure(figsize=(20, 5))
sns.heatmap(Matrix)
# -
plt.figure(figsize=(20, 5))
sns.clustermap(Matrix, yticklabels=False, center = 0, z_score =1)
| workflows/genrate_nets.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('..')
# +
from mnist import MNIST
import numpy as np
mndata = MNIST('./mnist')
raw_x, raw_y = mndata.load_training()
raw_x = np.array(raw_x).astype(np.float32)/255.0
raw_y = np.array(raw_y)
CLASS_NAMES = np.arange(0,10)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
ind = 55
print(raw_y[ind])
img = np.reshape(raw_x[ind,:],(28,28))
plt.imshow(img)
# +
from utils import print_np_ram
print('raw_x',raw_x.shape,raw_x.dtype)
print_np_ram('raw_x',raw_x)
print('raw_y',raw_y.shape,raw_y.dtype)
print_np_ram('raw_y',raw_y)
# +
from sklearn.model_selection import train_test_split
random_state = 0
x_train, x_test, y_train, y_test = train_test_split(raw_x,raw_y,test_size=0.3, random_state=random_state)
print('x_train',x_train.shape)
print('x_test',x_test.shape)
print('y_train',y_train.shape)
print('y_test',y_test.shape)
# +
# %load_ext autoreload
# %autoreload 2
import tinyFlame
from tinyFlame.handler import ModelHandler, SAVE_OPTIONS, TrainHandler
from tinyFlame.losses import cross_entropy
from tinyFlame.metrics import accuracy, dummy_accuracy
from tinyFlame.models import SimpleMLP
def generate_model_name():
num_features = len(feature_names)-1
name = ''
for i in range(num_features-1):
name += feature_names[i]#+'.'
name += feature_names[num_features-1]+'_'
name += preproc+'_DROP01_'
name += 'L'+str(latent_layers)
name +='_U'+str(latent_dim)
name += '_bi'*bi_lstm
return name
batch_size = 50 # BATCH
epochs = 20 # 100
#model_name = generate_model_name()
model_name = 'nan'
input_dim = [x_train.shape[-1]]
output_dim = [len(CLASS_NAMES)]
model_parameters = {
'output_dim' : output_dim,
'epochs' : epochs,
'input_dim' : input_dim,
'hidden_units' : 200,
'hidden_layers' : 3, # 3
'drop_out' : 0.2,
'name' : model_name,
}
myModel = SimpleMLP(model_parameters)
train_parameters = {
'input_dim' : input_dim,
'epochs' : epochs,
'batch_size' : batch_size,
'data_path' : 'save',
'id' : 0,
}
print("output_dim:",model_parameters['output_dim'])
print("input_dim:",model_parameters['input_dim'])
myModel = ModelHandler(myModel, myModel.name, train_parameters)
myModel.build()
myModel.set_loss(cross_entropy)
myModel.set_metrics({'accuracy': accuracy,'dummy_accuracy': dummy_accuracy,})
#myModel.set_optimizer(optim.Adam(self.model.parameters(), lr=0.001))
load = 0
train = 1
debug = 0
save_mode = SAVE_OPTIONS.BEST_METRI_CFINAL
save_mode = SAVE_OPTIONS.BEST_LOSS_FINAL
patience = 2
myModel.set_train_handler(TrainHandler(save_mode, target_saved_metric=0, patience_epochs=patience))
if load:
myModel.load_model()
if train:
#myModel.fit((x_train,y_train),(x_train,y_train),debug=debug,save_mode=save_mode)
myModel.fit((x_train,y_train),(x_test,y_test), debug=debug)
#myModel.fit((x_train,y_train),(x_train,y_train), debug=debug)
# -
myModel.show_train()
pass
ind = 55
print(raw_y[ind])
img = np.reshape(raw_x[ind,:],(28,28))
plt.imshow(img)
'''
BENCHMARK MINS 20 epocas:
> PC i7 NO-GPU oscar: 3.0066
'''
# +
import importlib
import metricUtils
importlib.reload(metricUtils)
from metricUtils import get_cm_accuracy
from tinyFlame.predictions import *
pred_arr, true_arr = predict(myModel, x_test, y_test)
cm, accuracy = get_cm_accuracy(pred_arr, true_arr)
# +
from metricUtils import plot_custom_confusion_matrix, ModelResults, ResultsID
results = ModelResults(myModel)
results.append(ResultsID(accuracy, cm))
#results.print()
plot_custom_confusion_matrix(cm, CLASS_NAMES)
pass
| tests_to_move/.ipynb_checkpoints/tinyFlame_example-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import drama as drm
import numpy as np
from time import time
import matplotlib.pylab as plt
from matplotlib import gridspec
# %matplotlib inline
# +
i_sig = 1 # signal number
n_ftrs = 100
noise = 0.2
scl = 0.01
sft = 0.01
X, y = drm.synt_event(i_sig,n_ftrs,sigma = noise,n1 = scl,n2 = sft,n3 = scl,n4 = sft)
gs = gridspec.GridSpec(1, 2)
plt.figure(figsize=(8,3))
ax1 = plt.subplot(gs[0, 0])
ax2 = plt.subplot(gs[0, 1])
ax1.set_title('Inliers')
ax2.set_title('Outliers')
inliers = X[y==0]
outliers = X[y==1]
for i in range(10):
ax1.plot(inliers[i],'b')
ax2.plot(outliers[i],'r')
# -
t0 = time()
for _ in range(10):
auc,mcc,rws,conf = drm.grid_run_drama(X,y)
print((time()-t0)/(10*5*10))
t0 = time()
for _ in range(10):
auc,mcc,rws,conf = drm.grid_run_lof(X,y)
print((time()-t0)/(10*3*3*20))
t0 = time()
for _ in range(10):
auc,mcc,rws,conf = drm.grid_run_iforest(X,y)
print((time()-t0)/(10*3*3*2))
| notebooks/old_set/runtime_benchmark.ipynb |
# +
"""
GPA Calculator
"""
import io
import pytest
from unittest import TestCase
from unittest.mock import patch
from p1 import *
@pytest.mark.describe('asserts True if conversion is correct')
def test_simple_gpa():
gpa = simple_gpa('A+')
assert gpa == 4.0
gpa = simple_gpa('A')
assert gpa == 4.0
gpa = simple_gpa('A-')
assert gpa == 3.7
gpa = simple_gpa('B+')
assert gpa == 3.3
gpa = simple_gpa('B')
assert gpa == 3.0
gpa = simple_gpa('B-')
assert gpa == 2.7
gpa = simple_gpa('C+')
assert gpa == 2.3
gpa = simple_gpa('C')
assert gpa == 2.0
gpa = simple_gpa('C-')
assert gpa == 1.7
gpa = simple_gpa('D+')
assert gpa == 1.3
gpa = simple_gpa('D')
assert gpa == 1.0
gpa = simple_gpa('D-')
assert gpa == 0.7
gpa = simple_gpa('F')
assert gpa == 0.0
| pset_functions/db_search/tests/nb/test_p1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python3 (TopoDevel)
# language: python
# name: topodevel
# ---
# # rstoolbox - a Python library for large-scale analysis of computational protein design data and structural bioinformatics
#
# [](https://doi.org/10.1101/428045)
#
# []()
#
# This notebook contains full details about the code examples presented in the paper.
#
# ## Imports
# +
# Default Libraries
import os
# External Libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
# Own Libraries
import rstoolbox as rs
#import readme
# Global matplotlib Parameters to match BMC figure requirements
plt.rcParams['svg.fonttype'] = 'none'
plt.rcParams.update({'font.size': 6.5,
'font.sans-serif': 'Arial',
'font.family': 'sans-serif'})
# rstoolbox configuration options.
rs.core.set_option('system', 'overwrite', True)
# rstoolbox configuration options. Put this one at the end of a cell to alter styling features when printing
# tabulated data (amongst other things).
rs.utils.format_Ipython()
# +
# This examples have all the possible Rosetta runs pre-calculated so it can run in a server.
# When trying to force-run the Rosetta functions, paths to the Rosetta instalation must be provided:
# rs.core.set_option('rosetta', 'path', '$YOUR_PATH_TO_ROSETTA_BINARIES')
# rs.core.set_option('rosetta', 'compilation', '$YOUR_ROSETTA_BINARIES_EXTENSION')
# -
# ## Analysis of protein backbone features
# ### Load Data
# +
# This function would normally execute Rosetta. As the file 'mota_1kx8_d2.dssp.minisilent' already exists, it will
# skip the execution. To force the function to re-calculate, delete the mentioned file.
ref = rs.io.get_sequence_and_structure('data/example01/mota_1kx8_d2.pdb')
# add_quality_measure would normally execute Rosetta. As the files 't001_.200.9mers.qual' and
# 'wauto.200.9mers.qual' already exist, it will skip the execution. To force the function to re-calculate,
# delete the mentioned files.
seqfrags = rs.io.parse_rosetta_fragments('data/example01/t001_.200.9mers')
seqfrags = seqfrags.add_quality_measure(None, 'data/example01/mota_1kx8_d2.pdb')
strfrags = rs.io.parse_rosetta_fragments('data/example01/wauto.200.9mers')
strfrags = strfrags.add_quality_measure(None, 'data/example01/mota_1kx8_d2.pdb')
# Default read of silent files loads all score terms but no sequence or structure information.
abseq = rs.io.parse_rosetta_file('data/example01/abinitio_seqfrags.minsilent.gz')
abstr = rs.io.parse_rosetta_file('data/example01/abinitio_strfrags.minsilent.gz')
# -
# ### Evaluate and Plot
# +
fig = plt.figure(figsize=(170 / 25.4, 170 / 25.4))
grid = (3, 6)
ax1 = plt.subplot2grid(grid, (0, 0), fig=fig, colspan=2)
rs.plot.plot_ramachandran_single(ref.iloc[0], 'A', ax1, scatter_s=2, line_linewidth=.4)
ax1.tick_params('y', labelrotation=90)
ax1 = plt.subplot2grid(grid, (0, 2), fig=fig, colspan=2)
rs.plot.plot_ramachandran_single(ref.iloc[0], 'A', ax1, 'PRE-PRO', scatter_s=2, line_linewidth=.4)
ax1.tick_params('y', labelrotation=90)
ax1 = plt.subplot2grid(grid, (0, 4), fig=fig, colspan=2)
rs.plot.plot_ramachandran_single(ref.iloc[0], 'A', ax1, 'PRO', scatter_s=2, line_linewidth=.4)
ax1.tick_params('y', labelrotation=90)
ax1 = plt.subplot2grid(grid, (1, 0), fig=fig, colspan=3)
ax2 = plt.subplot2grid(grid, (1, 3), fig=fig, colspan=3, sharey=ax1)
rs.plot.plot_fragments(seqfrags.slice_region(11, 46), strfrags.slice_region(11, 46), ax1, ax2,
titles=None, showfliers=False, linewidth=1)
ax2.axes.get_yaxis().set_visible(False)
rs.utils.add_top_title(ax1, 'sequence-based 9mers')
rs.utils.add_top_title(ax2, 'structure-based 9mers')
ax1.set_xlabel('residues')
ax2.set_xlabel('residues')
ax1 = plt.subplot2grid(grid, (2, 0), fig=fig, colspan=3)
sns.scatterplot(x="rms", y="score", data=abseq[abseq['score']<0], ax=ax1, linewidth=.2, s=4)
ax2 = plt.subplot2grid(grid, (2, 3), fig=fig, colspan=3, sharey=ax1, sharex=ax1)
sns.scatterplot(x="rms", y="score", data=abstr[abstr['score']<0], ax=ax2, linewidth=.2, s=4)
ax2.axes.get_yaxis().set_visible(False)
rs.utils.add_top_title(ax1, 'sequence-based fragments')
rs.utils.add_top_title(ax2, 'structure-based fragments')
ax1.set_xlabel('RMSD')
ax2.set_xlabel('RMSD')
plt.tight_layout(w_pad=0)
plt.savefig('images/example01_folding.png', dpi=300)
plt.show()
# -
# ## Guiding iterative CPD workflows
# Refreshing matplotlib styling parameters, as sometimes matplotlib gets confused.
plt.rcParams.update({'font.size': 6.5, 'font.sans-serif': 'Arial', 'font.family': 'sans-serif'})
# ### Load Data
# Make the file, which was split to be able to upload it to github
# !cat data/example02/1kx8_silent2.split_* > data/example02/1kx8_silent2.silent.gz
# By request, sequences for one or multiple chain identifiers can be also loaded while reading
# the silent file.
df = rs.io.parse_rosetta_file('data/example02/1kx8_silent2.silent.gz', {'sequence': '*'})
# ### Analysis and New Mutants
# +
# Select the top 5% decoys by score.
dftop = df[df['score'] < df['score'].quantile(.05)]
# Create a SequenceFrame with the frequencies of each residue type in each position for both the top set
# and the full population.
fstop = rs.analysis.sequential_frequencies(dftop, 'A', 'sequence', 'protein')
fs = df.sequence_frequencies('A') # Shorcut to utils.sequential_frequencies
# Calculate the difference between both.
fsdiff = (fstop - fs)
# Select the best residue type in each position in which a residue type is 20% more represented in the top
# popultaion than in the full population
muts = fsdiff[(fsdiff.T > 0.20).any()].idxmax(axis=1)
muts = list(zip(muts.index, muts.values))
# Select the best scored sequence that does NOT contain ANY of those residues.
pick = df.get_sequence_with('A', muts, confidence=0.25, invert=True).sort_values('score').iloc[:1]
seq = pick.iloc[0].get_sequence('A')
# And add itself as a reference sequence of the population (reference sequence is used when generating and
# identifying mutants).
pick.add_reference_sequence('A', seq)
# Generate the mutant variants that add the overepresented variants to the pick.
muts = [(muts[i][0], muts[i][1] + seq[muts[i][0] - 1]) for i in range(len(muts))]
variants = pick.generate_mutant_variants('A', muts)
variants.add_reference_sequence('A', seq)
# The resfiles are necessary to guide the mutation process.
variants = variants.make_resfile('A', 'NATAA', 'data/example02/mutants.resfile')
# This function would normally execute Rosetta. As the file 'variants.silent' already exists, it will
# skip the execution. To force the function to re-calculate, delete the mentioned file.
variants = variants.apply_resfile('A', 'data/example02/variants.silent')
variants = variants.identify_mutants('A')
# -
# ### Plot
# +
fig = plt.figure(figsize=(170 / 25.4, 170 / 25.4))
grid = (12, 4)
# Visualize over-represented residues in the top 5%
ax = plt.subplot2grid(grid, (0, 0), fig=fig, colspan=4, rowspan=6)
cbar_ax = plt.subplot2grid(grid, (6, 0), fig=fig, colspan=4, rowspan=1)
sns.heatmap(fsdiff.T, cmap="Blues", ax=ax, vmin=0, yticklabels=True, cbar_ax=cbar_ax, cbar_kws={"orientation": "horizontal"})
rs.utils.add_top_title(ax, 'Top scoring enrichment')
# Compare query positions in initial sequence and after mutant generation
ax = plt.subplot2grid(grid, (7, 0), fig=fig, colspan=2, rowspan=2)
rs.plot.logo_plot_in_axis( pick, 'A', ax=ax, refseq=False, key_residues=[_[0] for _ in muts] )
ax.set_ylabel('freq')
ax = plt.subplot2grid(grid, (7, 2), fig=fig, colspan=2, rowspan=2)
rs.plot.logo_plot_in_axis( variants, 'A', ax=ax, refseq=True, key_residues=[_[0] for _ in muts] )
ax.set_ylabel('freq')
# Check which mutations perform better
ax = plt.subplot2grid(grid, (9, 0), fig=fig, colspan=2, rowspan=3)
sns.scatterplot('mutant_count_A', 'score', data=variants, ax=ax)
ax.tick_params('y', labelrotation=90)
# Show distribution in best performing decoys
ax = plt.subplot2grid(grid, (9, 2), fig=fig, colspan=2, rowspan=3)
rs.plot.logo_plot_in_axis( variants.sort_values('score').head(3), 'A', ax=ax, refseq=True, key_residues=[_[0] for _ in muts] )
ax.set_ylabel('freq')
plt.tight_layout()
plt.savefig('images/example02_mutants.png', dpi=300)
plt.show()
# -
# Clean big created files
# !rm data/example02/1kx8_silent2.silent.gz
# ## Evaluation of designed proteins
# Refreshing matplotlib styling parameters, as sometimes matplotlib gets confused.
plt.rcParams.update({'font.size': 6.5, 'font.sans-serif': 'Arial', 'font.family': 'sans-serif'})
# ### Load Data
# +
# This function would normally execute Rosetta. As the file '1kx8.dssp.minisilent' already exists, it will
# skip the execution. To force the function to re-calculate, delete the mentioned file.
baseline = rs.io.get_sequence_and_structure('data/example03/1kx8.pdb', minimize=True)
slen = len(baseline.iloc[0].get_sequence('A'))
# CATH 70% ID. We filter structures with scores over 0 to improve image visibility.
cath = rs.utils.load_refdata('cath', 70)
cath = cath[cath['score']<=0]
cath = cath[(cath['length']>=slen-5) & (cath['length']<=slen+5)]
# Load designs for gen1 and gen2
gen1 = rs.io.parse_rosetta_file('data/example03/gen1.minisilent.gz')
gen2 = rs.io.parse_rosetta_file('data/example03/gen2.silent')
# Identifiers of selected decoys
decoys = ['138_188_1kx8_0033_0018_0001', '158_188_1kx8_0033_0028_0001', '158_188_1kx8_0033_0044_0001',
'215_188_1kx8_0033_0024_0001', '72_188_1kx8_0033_0018_0001', '85_188_1kx8_0033_0016_0001']
# Load sample experimental data
dfcd = rs.io.read_CD('data/example03/CD/', model='J-815')
dfspr = rs.io.read_SPR('data/example03/spr_data.txt')
# -
# ### Plot and Analyse
# +
fig = plt.figure(figsize=(170 / 25.4, 170 / 25.4))
grid = (3, 4)
axs = rs.plot.multiple_distributions(gen2, fig, (3, 4), values=['score', 'hbond_bb_sc', 'hbond_sc', 'rmsd'],
refdata=gen1, violins=False, legends=True, showfliers=False, linewidth=1)
for l in range(0, len(axs) - 1):
axs[l].get_legend().remove()
rs.utils.edit_legend_text(axs[-1], ['gen2', 'gen1'])
axs[-1].set_xlabel('RMSD')
axs = rs.plot.plot_in_context(gen2[gen2['description'].isin(decoys)], fig, (3, 2), cath, (1, 0),
['score', 'cav_vol'], ref_equivalences={'cavity': 'cav_vol'})
axs[0].axvline(baseline.iloc[0]['score'], color='k', linestyle='--', linewidth=1)
axs[1].axvline(baseline.iloc[0]['cavity'], color='k', linestyle='--', linewidth=1)
data_axs1 = axs[1].get_lines()
data_axs1[1].set_linestyle('None')
axs[1].legend([data_axs1[0], data_axs1[1], data_axs1[-1]], ['cath', 'designs', '1kx8'], loc='upper right')
ax = plt.subplot2grid(grid, (2, 0), fig=fig, colspan=2)
rs.plot.plot_CD(dfcd, ax, sample=5)
ax.tick_params('y', labelrotation=90)
ax = plt.subplot2grid(grid, (2, 2), fig=fig, colspan=2)
rs.plot.plot_SPR(dfspr, ax, fitcolor='black')
ax.tick_params('y', labelrotation=90)
data_ax = ax.get_lines()
ax.legend([data_ax[0], data_ax[1]], ['SPR data', 'model fits'], loc='upper right')
plt.tight_layout(w_pad=0)
plt.savefig('images/example03_evaluation.png', dpi=300)
plt.savefig('images/example03_evaluation.svg', dpi=300)
plt.show()
# -
# ## Comparison and benchmarking of design protocols
# Refreshing matplotlib styling parameters, as sometimes matplotlib gets confused.
plt.rcParams.update({'font.size': 6.5, 'font.sans-serif': 'Arial', 'font.family': 'sans-serif'})
# ### Load Data
# +
# This function would normally execute Rosetta. As the file '4oyd.dssp.minisilent' already exists, it will
# skip the execution. To force the function to re-calculate, delete the mentioned file.
baseline = rs.io.get_sequence_and_structure('data/example04/4oyd.pdb.gz', minimize=True)
# Combine data from multiple, consecutive analysis. This part is greatly simplified in the main text, in which
# only two files are combined; in reality, there are 4 files for each experiment. Also, some definitions are
# used to limit the amount of loaded data. To ease comprehension, the first file ('designs') is the actual silent
# file, while the second ('evals') will be a CSV dump of the other 3.
experiments = ['no_target', 'static', 'pack', 'packmin']
df = []
for experiment in experiments:
# Load Rosetta silent file from decoy generation
ds = rs.io.parse_rosetta_file('data/example04/{}.design.gz'.format(experiment),
'data/example04/description.json')
# Load the processed scores. Casting DataFrame into DesignFrame is as easy as shown here.
ev = rs.components.DesignFrame(pd.read_csv('data/example04/{}.evals.gz'.format(experiment)))
# Different outputs for the same decoys can be combined through their ‘description’ field (decoy identifier)
df.append(ds.merge(ev, on='description'))
df = pd.concat(df)
df.add_reference_sequence('B', baseline.iloc[0].get_sequence('B')[:-1])
# -
# ### Plot and Analyse
# +
fig = plt.figure(figsize=(170 / 25.4, 170 / 25.4))
grid = (12, 4)
axs = rs.plot.multiple_distributions(df, fig, grid, values=['score', 'LocalRMSDH', 'post_ddg', 'bb_clash'],
labels=['score', 'RMSD', 'ddG', 'bb_clash'], x='binder_state',
order=experiments, showfliers=False, linewidth=1, rowspan=3)
for ax in axs:
ax.tick_params('x', labelrotation=35)
ax.set_xlabel('')
ax = plt.subplot2grid(grid, (3, 0), fig=fig, colspan=4, rowspan=4)
rs.plot.per_residue_matrix_score_plot(df[df['binder_state']=='no_target'].sort_values('score').iloc[0],
'B', ax, 'BLOSUM62', add_alignment=False, linewidth=1, color=0)
rs.plot.per_residue_matrix_score_plot(df[df['binder_state']=='pack'].sort_values('score').iloc[0],
'B', ax, 'BLOSUM62', add_alignment=False, linewidth=1, color=2,
selections=[('43-64', 'red')])
rs.utils.add_top_title(ax, 'no_target (blue) - pack (green)')
ax = plt.subplot2grid(grid, (7, 0), fig=fig, colspan=2, rowspan=4)
rs.plot.sequence_frequency_plot(df[df['binder_state']=='no_target'], 'B', ax, key_residues='43-64',
cbar=False, border_width=1, clean_unused=0.05, xrotation=90)
rs.utils.add_top_title(ax, 'no_target')
ax = plt.subplot2grid(grid, (7, 2), fig=fig, colspan=2, rowspan=4)
ax_cbar = plt.subplot2grid(grid, (11, 0), fig=fig, colspan=4)
rs.plot.sequence_frequency_plot(df[df['binder_state']=='pack'], 'B', ax, key_residues='43-64',
border_width=1, clean_unused=0.05, xrotation=90, cbar_ax=ax_cbar)
rs.utils.add_top_title(ax, 'pack')
plt.tight_layout()
plt.savefig('images/example04_benchmark.png', dpi=300)
plt.show()
# -
| notebook/rstoolbox_paper.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
from past.builtins import xrange
# +
# NAND gate features
# note: x0 is a dummy variable for the bias term
# x0 x1 x2
x = [[1., 0., 0.],
[1., 0., 1.],
[1., 1., 0.],
[1., 1., 1.]]
# Desired outputs
y = [1.,
1.,
1.,
0.]
# // ---- STATE RECAP ----
# // NAND Gate + X0 Bias and Y-true
# // X0 // X1 // X2 // Y
# // 1 // 0 // 0 // 1
# // 1 // 0 // 1 // 1
# // 1 // 1 // 0 // 1
# // 1 // 1 // 1 // 0
# -
def f_activation(F,z):
if F > z:
yhat = 1.
else:
yhat = 0.
return yhat
# eta - learning rate
# t - iterations
# z - threshold for activation function
eta = 0.1
t = 50
z = 0
# initalize weight vector with all zeros
w = np.zeros(len(x[0])) # weights
w
# +
A0 = 1.0
A1 = 0.0
A2 = 0.0
B0 = 1.0
B1 = 0.0
B2 = 1.0
C0 = 1.0
C1 = 1.0
C2 = 0.0
D0 = 1.0
D1 = 1.0
D2 = 1.0
print("X0,X1,X2")
print(A0,A1,A2)
print(B0,B1,B2)
print(C0,C1,C2)
print(D0,D1,D2)
print("Y")
print(A3)
print(B3)
print(C3)
print(D3)
# -
W0 = w[0]
W1 = w[1]
W2 = w[2]
A3 = y[0]
B3 = y[1]
C3 = y[2]
D3 = y[3]
A3,B3,C3,D3
#ts1e1
# Dot product of the weight vector and the first row of features
F0 = (W0*A0) + (W1*A1) + (W2+A2)
yhatA = f_activation(F0,z)
match = yhatA == A3
print(F0,yhatA,match)
# +
# y-pred false; update weights
W0 = W0 + eta * (A3 - yhatA) * A0
W1 = W1 + eta * (A3 - yhatA) * A1
W2 = W2 + eta * (A3 - yhatA) * A2
print(W0)
print(W1)
print(W2)
# -
F1_test = np.dot(x[1], [W0,W1,W2])
F1_test
F1 = (W0*B0) + (W1*B1) + (W2*B2)
F1
F1_wrong = (W0*B0) + (W1*B1) + (W2+B2)
F1_wrong
# ..............
F1 = (W0*B0) + (W1*B1) + (W2+B2)
yhatB = f_activation(F1,z)
matchB = yhatB == B3
print(F1, yhatB,match)
F2 = (W0*C0) + (W1*C1) + (W2+C2)
yhatC = f_activation(F2,z)
matchC = yhatC == C3
print(F2, yhatC)
F2
F3 = (W0*D0) + (W1*D1) + (W2+D2)
yhatD = f_activation(F3,z)
matchD = yhatD == D3
print(F3, yhatD)
for i in xrange(0,len(y)):
errors[i] = (y[i]-yhat_vec[i])**2
for i in xrange(0, len(x)):
# summation step
f = np.dot(x[i], w)
print(f)
# weight vector
w
| nand_lbl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] tags=["meta", "draft"]
# # Seaborn
# -
# Source: https://github.com/jdhp-docs/notebooks/blob/master/python_seaborn_en.ipynb
# <a href="https://colab.research.google.com/github/jdhp-docs/notebooks/blob/master/python_seaborn_en.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
# <a href="https://mybinder.org/v2/gh/jdhp-docs/notebooks/master?filepath=python_seaborn_en.ipynb"><img align="left" src="https://mybinder.org/badge.svg" alt="Open in Binder" title="Open and Execute in Binder"></a>
# Official documentation: https://seaborn.pydata.org/index.html
# + tags=["hide"]
import seaborn as sns
import pandas as pd
import math
# -
sns.__version__
# ## Aspect
sns.set_context('talk')
#sns.set_context('poster')
# ### Figsize
df = sns.load_dataset("fmri")
df.head()
sns.relplot(x="timepoint", y="signal", kind="line", data=df);
sns.relplot(x="timepoint", y="signal", kind="line", data=df,
height=6, aspect=2);
# ### Legend
# +
l = []
for run in range(100):
for a in (1., 3.):
for x in range(10):
y = a * x + 10. * np.random.normal()
row = [x, y, a, run]
l.append(row)
df = pd.DataFrame(l, columns=["x", "y", "a", "run"])
df.head()
# -
sns.catplot(x="x", y="y", hue="a", data=df,
kind="point",
height=6, aspect=2);
# +
g = sns.catplot(x="x", y="y", hue="a", data=df,
kind="point",
height=6, aspect=2);
g._legend.set_title("Slope")
# + [markdown] toc-hr-collapsed=false
# ## Relplot
# -
# ### Scatter plot
tips = sns.load_dataset("tips")
tips.head()
sns.relplot(x="total_bill", y="tip", data=tips);
sns.scatterplot(x="total_bill", y="tip", data=tips);
sns.relplot(x="total_bill", y="tip", hue="size", size="day", style="time", row="sex", col="smoker", data=tips);
# + [markdown] toc-hr-collapsed=true
# ### Line plot
# -
# Official documentation: https://seaborn.pydata.org/tutorial/relational.html#aggregation-and-representing-uncertainty
#
# "The default behavior in seaborn is to aggregate the multiple measurements at each x value by plotting the mean and the 95% confidence interval around the mean."
# #### First example
# +
l = []
sigma = 1.
for run in range(1000):
for x in np.linspace(-10, 10, 100):
row = [x, np.random.normal(loc=0., scale=sigma), run]
l.append(row)
df = pd.DataFrame(l, columns=["x", "y", "run"])
df.head()
# +
sns.relplot(x="x", y="y", kind="line", data=df,
height=6, aspect=2)
plt.axhline(0, color="r", linestyle=":", label="Actual mean")
plt.legend();
# +
sns.relplot(x="x", y="y", kind="line", data=df,
height=6, aspect=2,
units="run", estimator=None, alpha=0.1)
plt.axhline(0, color="r", linestyle=":", label="Actual mean")
plt.legend();
# +
sns.relplot(x="x", y="y", data=df,
height=6, aspect=2, marker=".",
estimator=None, alpha=0.15)
plt.axhline(2. * sigma, color="k", linestyle=":", label=r"$2 \sigma$")
plt.axhline(0, color="r", linestyle=":", label="Actual mean")
plt.axhline(-2. * sigma, color="k", linestyle=":", label=r"$2 \sigma$")
plt.legend();
# +
sns.relplot(x="x", y="y", kind="line", data=df,
height=6, aspect=2,
estimator=np.median)
plt.axhline(0, color="r", linestyle=":", label="Actual median")
plt.legend();
# -
# #### Second example
# +
l = []
for run in range(100):
for func in ("sin", "cos"):
for x in np.linspace(-10, 10, 100):
y = math.sin(x) if func == "sin" else math.cos(x)
row = [x, y + np.random.normal(), func, run]
l.append(row)
df = pd.DataFrame(l, columns=["x", "y", "func", "run"])
df.head()
# -
sns.relplot(x="x", y="y", kind="line", hue="func", data=df,
height=6, aspect=2);
# #### Third example
fmri = sns.load_dataset("fmri")
fmri.head()
sns.relplot(x="timepoint", y="signal", data=fmri,
height=6, aspect=2);
sns.catplot(x="timepoint", y="signal", data=fmri, aspect=3);
sns.relplot(x="timepoint", y="signal", kind="line", data=fmri,
height=6, aspect=2);
# #### Fourth example
# +
l = []
for run in range(100):
for a in (1., 3.):
for x in range(10):
y = a * x + 10. * np.random.normal()
row = [x, y, a, run]
l.append(row)
df = pd.DataFrame(l, columns=["x", "y", "a", "run"])
df.head()
# -
sns.relplot(x="x", y="y", hue="a", data=df,
kind="line",
height=6, aspect=2);
# The legend is bad because relplot() is made for real values, even for the "hue" variable... Here, catplot would be more adapted.
# ## Catplot
# +
l = []
for run in range(100):
for a in (1., 3.):
for x in range(10):
y = a * x + 10. * np.random.normal()
row = [x, y, a, run]
l.append(row)
df = pd.DataFrame(l, columns=["x", "y", "a", "run"])
df.head()
# -
sns.catplot(x="x", y="y", hue="a", data=df,
kind="point",
height=6, aspect=2);
sns.catplot(x="x", y="y", hue="a", data=df,
kind="point",
markers=".",
scale=0.7,
linestyles=":",
capsize=0.1,
height=6, aspect=2);
# + [markdown] toc-hr-collapsed=true
# ## Pairplot
# +
# https://seaborn.pydata.org/tutorial/distributions.html#visualizing-pairwise-relationships-in-a-dataset
iris = sns.load_dataset("iris")
iris.head()
# -
sns.pairplot(iris, hue="species");
# +
# https://seaborn.pydata.org/tutorial/distributions.html#visualizing-pairwise-relationships-in-a-dataset
titanic = sns.load_dataset("titanic")
titanic.head()
# -
sns.pairplot(titanic, vars=["survived", "pclass", "fare"], hue="survived");
| nb_dev_python/python_seaborn_en.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pyspark.sql import SparkSession
spark=SparkSession.builder.appName("LogisticRegression").getOrCreate()
from pyspark.ml.classification import LogisticRegression
from pyspark.ml.feature import VectorAssembler
from pyspark.sql.types import *
from pyspark.sql.functions import *
my_data=spark.read.csv("C:/Users/User/Desktop/Data/flights.csv", header=True, inferSchema=True)
my_data.show(5)
my_data.printSchema()
flightschema=StructType([
StructField ("DayofMonth", IntegerType(),False),
StructField("DayofWeek", IntegerType(),False),
StructField("Carrier", StringType(),False),
StructField ("OriginAirportID", IntegerType(),False),
StructField("DestAirportID", IntegerType(),False),
StructField("DepDelay", IntegerType(),False),
StructField("ArrDelay", IntegerType(),False)
]
)
df=spark.read.csv("C:/Users/User/Desktop/Data/flights.csv", header=True, schema=flightschema)
df.show(5)
df.printSchema()
# # select Some import data for Classification features and change arrival delay into binary class
# * late
# * not late
df1=df.select("DayofMonth","DayofWeek","originAirportID","DestAirportID","DepDelay",\
((col("ArrDelay") > 15).cast("Int").alias("Late")))
df1.show()
# # Dividing Data into Train and Test
train_data,test_data=df1.randomSplit([0.7,0.3])
train_data.count()
test_data.count()
# # Preparing Data
# Vector Assembler
assembler=VectorAssembler(inputCols=["DayofMonth","DayofWeek","originAirportID","DestAirportID","DepDelay"]\
,outputCol="features")
tran_data=assembler.transform(df1)
tran_data.show(5)
# # Final DataSet
tran_data=tran_data.select("features",tran_data["Late"].alias("label"))
tran_data.show(5)
train_data,test_data=tran_data.randomSplit([0.7,0.3])
train_data.count()
test_data.count()
train_data.show(2)
# # Training Data
lr=LogisticRegression(featuresCol="features",labelCol="label",predictionCol="prediction",maxIter=10, regParam=0.3)
lrmodel=lr.fit(train_data)
print("Model is trained")
lrmodel.transform(train_data).show(10)
lrmodel.coefficients
# Grab the Correct prediction
train_pred=lrmodel.transform(train_data)
train_pred.show(5)
correct_prediction=train_pred.filter(train_pred["label"]==train_pred["prediction"]).count()
print("Accuracy for training-data :,",correct_prediction/(train_data.count()))
# # testing Data
test=lrmodel.transform(test_data)
correct_prediction_test=test.filter(test["label"]==test["prediction"]).count()
print("Accuracy for test-data :,",correct_prediction_test/(test_data.count()))
| .ipynb_checkpoints/10b.Random Forest Classifier-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Nearest-neighbour indexing using xoak
#
# This notebook experiments subsetting datasets using `TLONG, TLAT, ULONG, ULAT` by making use of the [xoak](https://xoak.readthedocs.io/en/latest/) package
# +
import matplotlib as mpl
import numpy as np
import xarray as xr
import xoak
import pop_tools
# +
# open sample data
filepath = pop_tools.DATASETS.fetch('Pac_POP0.1_JRA_IAF_1993-12-6-test.nc')
ds = xr.open_dataset(filepath)
# get DZU and DZT, needed for operations later on
filepath_g = pop_tools.DATASETS.fetch('Pac_grid_pbc_1301x305x62.tx01_62l.2013-07-13.nc')
ds_g = xr.open_dataset(filepath_g)
ds.update(ds_g)
# -
grid, xds = pop_tools.to_xgcm_grid_dataset(ds)
# ## Set the "index"
#
# This is what allows the indexing magic.
xds.xoak.set_index(['TLAT', 'TLONG'], 'scipy_kdtree')
# ## Extracting sections
# +
# have to know that 0.1 is a reasonable choice
lons = xr.Variable("points", np.arange(150, 270, 0.1))
lats = xr.zeros_like(lons)
eqsection = xds.xoak.sel(TLONG=lons, TLAT=lats)
# plot
eqsection.TEMP.sel(z_t=slice(50000)).plot(y="z_t", cmap=mpl.cm.Spectral_r, yincrease=False)
# -
# ## Extracting multiple points
# +
moorings = xds.xoak.sel(
TLONG=xr.Variable("moor", [360 - 140, 360 - 110]),
TLAT=xr.Variable("moor", [0, 0]),
)
moorings.TEMP.plot(hue="TLONG", y="z_t", yincrease=False)
# -
# ## Extracting single points
# + tags=[]
xds.xoak.sel(TLONG=360 - 140, TLAT=0).TEMP
# -
# xoak expectes "trajectories" to sample along. For a single point we create a 1D variable representing the coordinate location we want: in this case `TLONG=220, TLAT=0`. Seems like this could be fixed: https://github.com/xarray-contrib/xoak/issues/37
xds.xoak.sel(
TLONG=xr.Variable("points", [360 - 140]), TLAT=xr.Variable("points", [0])
).TEMP.squeeze("points")
# ## Limitations
# - [ ] cannot simply index by point
# - [ ] Can only set one index at a time (so only, TLONG, TLAT or ULONG, ULAT)
# - [ ] indexes are not propagated so `xds.TEMP.xoak` will only work after `xds.TEMP.set_index` is called
xds.TEMP.xoak.sel(TLONG=220, TLAT=0)
xds.xoak.sel(TLONG=220, TLAT=0)
# %load_ext watermark
# %watermark -d -iv -m -g
| docs/source/examples/xoak-example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.3 64-bit
# name: python38364bitb680b9ea85ba4b86b35b18465fd75df3
# ---
# # Solving AT1 Q1 - finding shortest path and diameter in weighted edges graph
#
# 
# +
# construct the graph
import networkx as nx
import matplotlib.pyplot as plt
G = nx.Graph()
G.add_nodes_from("ABCDEFGHIJK")
# +
weighted_edges = [
('A','B',2),('A','C',1),('A','E',3),
('B','E',1),
('C','I',2),('C','F',3),('C','D',4),
('D','F',1),('D','J',2),('D','K',3),
('E','I',4),
('F','J',3),('F','G',3),
('G','I',1),('G','J',1),('G','H',5),
('H','J',1),('H','K',4),
#('I'),
('J','K',2)#,
#('K')
]
pos = {
"A": (2,8),
"B": (1,5),
"C": (5,8.5),
"D": (6,4),
"E": (4,4.5),
"F": (8,6),
"G": (11,7),
"H": (13, 6),
"I": (8.5, 8.5),
"J": (10, 3.5),
"K": (12,1)
}
# -
G.add_weighted_edges_from(weighted_edges)
edge_labels = nx.get_edge_attributes(G,'weight')
# pos = nx.circular_layout(G)
nx.draw_networkx_nodes(G, pos, node_size=500, node_color='grey')
nx.draw_networkx_labels(G,pos,font_size=15,font_family='sans-serif')
nx.draw_networkx_edges(G,pos)
nx.draw_networkx_edge_labels(G,pos,edge_labels=edge_labels)
plt.show()
print(f"Number of nodes: {G.order()}")
print(f"Number of edges: {G.size()}")
# find shortest path from A to K
A_K_shortestpath = nx.dijkstra_path(G, source='A',target='K')
A_K_shortestpath_length = nx.dijkstra_path_length(G, source='A', target=A_K_shortestpath[-1])
print(f"shortest path from A to K is {A_K_shortestpath_length} long: {'-'.join(A_K_shortestpath)}")
# +
nodes_list = list(G.nodes)
diameter = 0
diameter_path = None
for i, start_n in enumerate(G.nodes):
print(i, start_n)
nodes_list.pop(0) # remove current node from nodes list
str = ""
diameter_updated = False
for j, target_n in enumerate(nodes_list):
shortest_path = nx.dijkstra_path(G, source=start_n, target=target_n)
shortest_path_length = nx.dijkstra_path_length(G, source=start_n, target=target_n)
str = str + f"({j}, {target_n}, {'-'.join(shortest_path)}, {shortest_path_length})"
if shortest_path_length > diameter:
diameter = shortest_path_length
diameter_path = shortest_path
diameter_updated = True
print(str)
print(f"{'update diameter' if diameter_updated else ''}")
# -
print(f"network diamter is {diameter} along path {'-'.join(diameter_path)}")
| AT1-Q1-shortestpath-diameter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import cv2 as cv
import numpy as np
# +
#캠 없어서 에러
capture = cv.VideoCapture(0)
count = 0
fps = 0
t0 = cv.getTickCount()
while True:
ret, frame = capture.read()
cv.putText(frame, str(fps), (10, 60), cv.FONT_HERSHEY_SIMPLEX, 2, (0,255,0), 3)
cv.imshow('frame', frame)
if cv.waitKey(1) == 27:
break
count += 1
if count == 10:
t = cv.getTickCount()
time = (t - t0) / cv.getTickFrequency()
fps = int(np.round(count / time))
count = 0
capture.release()
# -
| AI 이노베이션 스퀘어 시각지능 과정/202004/20200424/20200424_opencv_example01_video_capture.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## _*Quantum SVM (quantum kernel method)*_
#
# ### Introduction
#
# Please refer to [this file](./svm_qkernel.ipynb) for introduction.
#
# This file shows an example how to use Aqua API to build SVM classifier and keep the instance for future prediction.
from datasets import *
from qiskit_aqua.utils import split_dataset_to_data_and_labels, map_label_to_class_name
from qiskit_aqua.input import get_input_instance
from qiskit_aqua import run_algorithm, get_feature_map_instance, get_algorithm_instance, get_multiclass_extension_instance
# First we prepare the dataset, which is used for training, testing and the finally prediction.
#
# *Note: You can easily switch to a different dataset, such as the Breast Cancer dataset, by replacing 'ad_hoc_data' to 'Breast_cancer' below.*
# +
n = 2 # dimension of each data point
training_dataset_size = 20
testing_dataset_size = 10
sample_Total, training_input, test_input, class_labels = ad_hoc_data(training_size=training_dataset_size,
test_size=testing_dataset_size,
n=n, gap=0.3, PLOT_DATA=False)
datapoints, class_to_label = split_dataset_to_data_and_labels(test_input)
print(class_to_label)
# -
# With the dataset ready we initialize the necessary inputs for the algorithm:
# - build all components required by SVM
# - feature_map
# - multiclass_extension (optional)
# +
svm = get_algorithm_instance("QSVM.Kernel")
svm.random_seed = 10598
svm.setup_quantum_backend(backend='local_statevector_simulator')
feature_map = get_feature_map_instance('SecondOrderExpansion')
feature_map.init_args(num_qubits=2, depth=2, entanglement='linear')
svm.init_args(training_input, test_input, datapoints[0], feature_map)
# -
# With everything setup, we can now run the algorithm.
#
# The run method includes training, testing and predict on unlabeled data.
#
# For the testing, the result includes the success ratio.
#
# For the prediction, the result includes the predicted class names for each data.
#
# After that the trained model is also stored in the svm instance, you can use it for future prediction.
result = svm.run()
# +
print("kernel matrix during the training:")
kernel_matrix = result['kernel_matrix_training']
img = plt.imshow(np.asmatrix(kernel_matrix),interpolation='nearest',origin='upper',cmap='bone_r')
plt.show()
print("testing success ratio: ", result['testing_accuracy'])
print("predicted classes:", result['predicted_classes'])
# -
#
# Use the trained model to evaluate data directly, and we store a `label_to_class` and `class_to_label` for helping converting between label and class name
# +
predicted_labels = svm.predict(datapoints[0])
predicted_classes = map_label_to_class_name(predicted_labels, svm.label_to_class)
print("ground truth: {}".format(datapoints[1]))
print("preduction: {}".format(predicted_labels))
# -
| artificial_intelligence/qsvm_kernel_directly.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import csv
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
df_pos = pd.read_csv('positive_emotions.csv')
df_pos = df_pos.replace(r'^s*$', float('NaN'), regex = True)
df_pos = df_pos.replace(r' ', float('NaN'), regex = True)
df_pos = df_pos[df_pos['positive_premeasure'].notna()]
df_pos = df_pos[df_pos['positive_pep1'].notna()]
uids_list = df_pos['uid'].unique()
print(len(uids_list))
# ### Group Assignments
# 1. High anger, High anxiety
# 2. Low anger, High anxiety
# 3. High anger, Low anxiety
# 4. Low anger, Low anxiety
mean_anger = np.mean([float(elem) for elem in df_anx['anger_premeasure'].to_numpy()])
mean_anx = np.mean([float(elem) for elem in df_anx['anxiety_premeasure'].to_numpy()])
uid_to_anger_anx = {}
uid_to_group = {}
for index, row in df_anx.iterrows():
uid = row['uid']
anger = float(row['anger_premeasure'])
anx = float(row['anxiety_premeasure'])
if anger >= mean_anger:
if anx >= mean_anx:
group = 1
else:
group = 3
else:
if anx >= mean_anx:
group = 2
else:
group = 4
uid_to_anger_anx[uid] = {'anger': anger, 'anxiety': anx, 'group': group}
uid_to_group[uid] = group
valid_uids = list(uid_to_group.keys())
# ## Compute Metrics
robot_X =[84, 84, 84, 84, 84, 84, 83, 83, 82, 82, 81, 81, 80, 80, 79, 79, 78, 78, 77, 77, 76, 76, 75, 75, 74, 74, 73, 73, 72, 72, 72, 72, 72, 72, 71, 71, 70, 70, 69, 69, 68, 68, 68, 68, 68, 68, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 70, 70, 71, 71, 72, 72, 72, 72, 72, 72, 71, 71, 70, 70, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 68, 68, 68, 68, 68, 68, 67, 67, 66, 66, 65, 65, 64, 64, 63, 63, 62, 62, 61, 61, 60, 60, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 61, 61, 62, 62, 62, 62, 62, 62, 62, 62, 63, 63, 63, 63, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 65, 65, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 66, 65, 65, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 63, 63, 62, 62, 62, 62, 61, 61, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 59, 59, 59, 59, 59, 59, 60, 60, 61, 61, 62, 62, 63, 63, 64, 64, 65, 65, 66, 66, 67, 67, 68, 68, 69, 69, 70, 70, 71, 71, 72, 72, 73, 73, 74, 74, 75, 75, 76, 76, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 79, 79, 80, 80, 80, 80, 80, 80, 80, 80, 81, 81, 81, 81, 82, 82, 82, 82, 82, 82, 82, 82, 82, 82, 83, 83, 82, 82, 82, 82, 81, 81, 81, 81, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 80, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 81, 80, 80, 80, 80, 80, 80, 79, 79, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 77, 78, 78, 79, 79, 80, 80, 81, 81, 82, 82, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 83, 82, 82, 81, 81, 80, 80, 79, 79, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 78, 77, 77, 77, 77, 77, 77, 77, 77, 76, 76, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 75, 74, 74, 73, 73, 72, 72, 71, 71, 70, 70, 69, 69, 68, 68, 67, 67, 66, 66, 65, 65, 64, 64, 63, 63, 62, 62, 61, 61, 60, 60, 59, 59, 58, 58, 57, 57, 56, 56, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 56, 56, 57, 57, 58, 58, 58, 58, 59, 59, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 60, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 58, 58, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 59, 58, 58, 57, 57, 56, 56, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 55, 56, 56, 57, 57, 58, 58, 59, 59, 60, 60, 61, 61, 62, 62, 63, 63, 64, 64, 65, 65, 66, 66, 67, 67, 68, 68, 69, 69, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 70, 71, 71, 72, 72, 73, 73, 74, 74, 75, 75, 76, 76, 77, 77, 78, 78, 78, 78, 78, 78, 78, 78, 79, 79, 80, 80, 81, 81, 82, 82, 83, 83, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 84, 85, 85, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 86, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 87, 88, 88, 88, 88, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 89, 88, 88, 87, 87, 86, 86, 86, 86, 86, 86, 86, 86, 85, 85, 84, 84, 83, 83, 82, 82, 81, 81, 80, 80, 79, 79, 78, 78, 77, 77, 77, 77, 76, 76, 76, 76, 75, 75, 75, 75, 74, 74, 74, 74, 73, 73, 72, 72, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 71, 72, 72, 72, 72, 72, 72, 72, 72, 71, 71, 70, 70, 70, 70, 69, 69, 68, 68, 67, 67, 66, 66, 65, 65, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 64, 65, 65, 66, 66, 67, 67, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 69, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 68, 69, 69]
robot_Y =[38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 38, 39, 39, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 41, 41, 42, 42, 42, 42, 43, 43, 44, 44, 45, 45, 46, 46, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 43, 43, 42, 42, 42, 42, 41, 41, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 41, 41, 42, 42, 42, 42, 42, 42, 42, 42, 43, 43, 44, 44, 45, 45, 46, 46, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 44, 44, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 42, 42, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 44, 44, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 44, 44, 45, 45, 46, 46, 46, 46, 46, 46, 46, 46, 46, 46, 47, 47, 46, 46, 46, 46, 46, 46, 46, 46, 46, 46, 46, 46, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 43, 43, 42, 42, 42, 42, 41, 41, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 40, 41, 41, 42, 42, 42, 42, 42, 42, 42, 42, 43, 43, 44, 44, 45, 45, 46, 46, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 44, 44, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 43, 44, 44, 44, 44, 44, 44, 44, 44, 44, 44, 44, 44, 45, 45, 44, 44, 44, 44, 44, 44, 45, 45, 45, 45, 45, 45, 45, 45, 45, 45, 46, 46, 45, 45, 45, 45, 45, 45, 46, 46, 47, 47, 47, 47, 47, 47, 46, 46, 45, 45, 44, 44, 43, 43, 42, 42, 42, 42, 41, 41, 40, 40, 39, 39, 38, 38, 37, 37, 36, 36, 35, 35, 34, 34, 33, 33, 32, 32, 31, 31, 30, 30, 29, 29, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 30, 30, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 29, 30, 30, 31, 31, 32, 32, 33, 33, 34, 34, 34, 34, 34, 34, 34, 34, 34, 34, 35, 35, 34, 34, 34, 34, 34, 34, 35, 35, 36, 36, 37, 37, 37, 37, 37, 37, 36, 36, 35, 35, 34, 34, 33, 33, 32, 32, 31, 31, 30, 30, 29, 29, 28, 28, 27, 27, 26, 26, 25, 25, 24, 24, 23, 23, 22, 22, 21, 21, 20, 20, 19, 19, 18, 18, 17, 17, 16, 16, 15, 15, 14, 14, 13, 13, 12, 12, 11, 11, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 9, 9, 8, 8, 7, 7, 6, 6, 5, 5, 4, 4, 3, 3, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 4, 4, 3, 3, 3, 3, 3, 3, 4, 4, 5, 5, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 6, 5, 5, 5, 5, 5, 5, 5, 5, 4, 4, 4, 4, 4, 4, 4, 4, 4, 4, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 10, 9, 9, 8, 8, 7, 7, 6, 6, 5, 5, 4, 4, 3, 3, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 2, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 4, 4, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 9, 9, 9, 9, 8, 8, 7, 7, 6, 6, 5, 5, 4, 4, 3, 3, 2, 2, 1, 1, 1, 1, 1, 1, 2, 2, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 3, 4, 4, 3, 3, 3, 3, 3, 3, 4, 4, 4, 4, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 10, 10, 11, 11, 12, 12, 12, 12, 12, 12, 12, 12, 12, 12, 13, 13, 12, 12, 12, 12, 12, 12, 12, 12, 12, 12, 11, 11, 10, 10, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 9, 8, 8, 8, 8, 7, 7, 7, 7, 6, 6, 6, 6, 5, 5, 5, 5, 5, 5, 5, 5, 6, 6, 7, 7, 8, 8, 9, 9, 10, 10, 11, 11, 12, 12, 13, 13, 13, 13, 14, 14, 15, 15, 16, 16, 16, 16, 16, 16, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 15, 16, 16, 17, 17, 17, 17, 17, 17, 17, 17, 17, 17, 18, 18, 17, 17, 17, 17, 17, 17, 17, 17, 17, 17, 17, 17, 18, 18, 19, 19, 20, 20, 21, 21, 22, 22, 23, 23, 24, 24, 25, 25, 26, 26, 26, 26, 26, 26, 26, 26, 26, 26, 27, 27, 26, 26, 26, 26, 26, 26, 27, 27, 27, 27, 27, 27, 27, 27, 27, 27, 28, 28, 27, 27, 27, 27, 27, 27, 28, 28, 28, 28, 28, 28, 28, 28, 28, 28, 29, 29, 28, 28, 28, 28, 27, 27, 26, 26, 25, 25, 24, 24, 23, 23, 22, 22, 21, 21, 20, 20, 20, 20, 20, 20, 20, 20, 20, 20, 20, 20]
assert len(robot_X) == len(robot_Y)
# +
# if (minutes ==4 && seconds < 45 && seconds >= 41) {
# otherNumSteps += 0;
# }
# else if (minutes ==4 && seconds < 26 && seconds >= 24) {
# otherNumSteps += 0;
# }
# else if (minutes ==3 && seconds < 46 && seconds >= 43) {
# otherNumSteps += 0;
# }
# else if (minutes ==2 && seconds < 50 && seconds >= 48) {
# otherNumSteps += 0;
# }
# else if (minutes ==1 && seconds < 32 && seconds >= 30) {
# otherNumSteps += 0;
# }
# -
plt.scatter(robot_X, robot_Y)
# # Process Robot Trajectory
N = len(robot_X)
print(N)
difference = int(N/10)
print(difference)
# +
min_to_robot_traj = {(5,0):[]}
counter = 0
for minute in [4,3,2, 1, 0]:
for second in [30, 0]:
counter += 1
keyname = (minute, second)
t_robot_x = robot_X[difference*(counter-1): difference*(counter)]
t_robot_y = robot_Y[difference*(counter-1): difference*(counter)]
traj_robot = [(t_robot_x[i], t_robot_y[i]) for i in range(len(t_robot_x))]
min_to_robot_traj[keyname] = traj_robot
# -
min_to_robot_traj
# # Get Human Data
df = pd.read_csv('minimap_study_3_data.csv')
# +
i = 40
p_uid = valid_uids[i]
df_traj = df[(df['userid']==p_uid) & (df['episode']==1)]
df_traj = df_traj.sort_values(by=['time_spent'], ascending=False)
# -
# +
def compute_minute_to_traj(df_traj):
traj = df_traj['trajectory'].to_numpy()
times = df_traj['time_spent'].to_numpy()
sent_msgs = df_traj['sent_messages'].to_numpy()
victims = df_traj['target'].to_numpy()
to_include = True
minute_to_traj = {}
minute_to_msgs = {}
minute_to_victims = {}
curr_min = 5
curr_sec = 0
where_start = np.where(times == 'start')[0]
if len(where_start) == 0:
minute_to_traj[(curr_min, curr_sec)] = []
else:
# print("where_start = ", where_start)
start_idx = where_start[0]
t = str(times[start_idx])
traj_t = str(traj[start_idx])
msgs_t = str(sent_msgs[start_idx])
vic_t = str(victims[start_idx])
if traj_t == 'nan':
curr_traj = []
else:
curr_traj = [eval(x) for x in traj_t.split(';')]
if msgs_t == 'nan':
curr_msgs = []
else:
curr_msgs = msgs_t
minute_to_traj[(curr_min, curr_sec)] = curr_traj
# minute_to_msgs[(curr_min, curr_sec)] = minute_to_msgs
# minute_to_victims[(curr_min, curr_sec)] = []
if curr_sec == 0:
curr_min -= 1
curr_sec = 30
curr_traj = []
prev_min = 5
prev_seconds = 60
for i in range(len(traj)):
t = str(times[i])
traj_t = str(traj[i])
msgs_t = sent_msgs[i]
vic_t = victims[i]
if ':' not in t:
continue
t_min = int(t.split(":")[0])
t_sec = int(t.split(":")[1])
if traj_t != 'nan':
curr_t_traj = [eval(x) for x in traj_t.split(';')]
# print("INPUTS TO ROUND")
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
if t_min == curr_min:
if t_sec == curr_sec:
curr_traj.extend(curr_t_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
if curr_sec == 0:
curr_min -= 1
curr_sec = 30
else:
curr_sec = 0
curr_traj = []
elif t_sec > curr_sec:
curr_traj.extend(curr_t_traj)
elif t_sec < curr_sec and curr_sec == 30:
diff_in_past_section = abs(prev_seconds - curr_sec) # 2:45-2:30
diff_in_next_section = abs(curr_sec - t_sec) #2:30-2
# 2- 1:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_next_section))
next_section_idx = past_section_idx + 1
curr_traj.extend(curr_t_traj[:past_section_idx+1])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = []
curr_traj.extend(curr_t_traj[next_section_idx:])
elif t_sec == 0 and curr_sec == 30:
diff_in_past_section = abs(prev_seconds - curr_sec) # 2:45 - 2:30
diff_in_next_section = abs(curr_sec - t_sec) # 2:30- 2
# 2 - 1:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_next_section))
next_section_idx = past_section_idx + 1
curr_traj.extend(curr_t_traj[:past_section_idx+1])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = []
curr_traj.extend(curr_t_traj[next_section_idx:])
curr_min -= 1
curr_sec = 30
curr_traj = []
elif t_sec == 30 and curr_sec == 0:
diff_in_past_section = abs(prev_seconds - curr_sec) # 2:15 - 2:00
diff_in_next_section = abs(curr_sec - t_sec) # 2:00- 1:30
# 1:30-1
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_next_section))
next_section_idx = past_section_idx + 1
curr_traj.extend(curr_t_traj[:past_section_idx+1])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
curr_traj.extend(curr_t_traj[next_section_idx:])
curr_sec = 0
curr_traj = []
elif t_min == curr_min-1:
if t_sec > curr_sec and curr_sec == 0:
diff_in_past_section = abs(prev_seconds - curr_sec)
diff_in_next_section = abs(curr_sec - t_sec)
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_next_section))
next_section_idx = past_section_idx + 1
curr_traj.extend(curr_t_traj[:past_section_idx+1])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
elif t_sec == 30 and curr_sec ==30:
diff_in_past_section = abs(prev_seconds - 30) #2:40-2:30
diff_in_mid_section = 30 # 2:30-2
diff_in_next_section = 30 # 2-1:30
#1:30-1
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:next_section_idx+1]
next_section_traj = curr_t_traj[next_section_idx+1:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = []
elif t_sec == 0 and curr_sec == 0:
diff_in_past_section = abs(prev_seconds - 0) #2:04 -2
diff_in_mid_section = 30 # 2 - 1:30
diff_in_next_section = 30 # 1:30 - 1
#1-0:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:next_section_idx+1]
next_section_traj = curr_t_traj[next_section_idx+1:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = next_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
elif t_sec == 0 and curr_sec ==30:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:45-3:30
diff_in_mid_section = 30 #3:30 - 3
# diff_in_next_section = abs(30 - t_sec) # 3 - 2:30
# 2:30 - 2
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section))
# next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:]
# next_section_traj = curr_t_traj[next_section_idx+1:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
# curr_traj = next_section_traj
# minute_to_traj[(curr_min, curr_sec)] = curr_traj
# curr_sec = 0
curr_traj = []
elif t_sec == 30 and curr_sec == 0:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:15-3
diff_in_mid_section = 30 #3 - 2:30
# diff_in_next_section = abs(30 - t_sec) # 2:30 - 2
# 2 - 1:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section))
# next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:]
# next_section_traj = curr_t_traj[next_section_idx+1:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
# curr_traj = next_section_traj
# minute_to_traj[(curr_min, curr_sec)] = curr_traj
# curr_min -= 1
# curr_sec = 30
curr_traj = []
elif t_sec > curr_sec and curr_sec == 30:
# 2 sections off
diff_in_past_section = abs(prev_seconds - curr_sec)
diff_in_mid_section = 30
diff_in_next_section = abs(60 - t_sec)
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:mid_section_idx+1]
next_section_traj = curr_t_traj[next_section_idx:]
# print("past_section_traj = ", past_section_traj)
# print("mid_section_traj = ", mid_section_traj)
# print("next_section_traj = ", next_section_traj)
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
elif t_sec == 0 and curr_sec == 30:
diff_in_past_section = abs(prev_seconds - 0)
diff_in_mid_section = 30
diff_in_next_section = abs(60 - t_sec)
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
mid_section_idx = past_section_idx + 1+ int(diff_in_mid_section/(diff_in_past_section + diff_in_mid_section+ diff_in_next_section))
next_section_idx = mid_section_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_traj = curr_t_traj[past_section_idx+1:]
next_section_traj = []
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
elif t_sec < curr_sec and curr_sec == 30:
# 3 sections off, 2:30 --> 1:14, 2:40-2:30, 2:30-2, 2-1:30, 1:30-1:14
# print("PROBLEMMM")
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
diff_in_past_section = abs(prev_seconds - 30)
diff_in_mid_1_section = 30
diff_in_mid_2_section = 30
diff_in_next_section = abs(30 - t_sec)
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
next_section_idx = mid_section_2_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
next_section_traj = curr_t_traj[mid_section_2_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = next_section_traj
# elif t_sec <= curr_sec and curr_sec == 0:
# # 3 sections off, 2:30 --> 1:14, 2:40-2:30, 2:30-2, 2-1:30, 1:30-1:14
# print("PROBLEMMM 2")
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
elif t_min == curr_min-2:
### 4:30 --> 2:30, 4:00 --> 2:00, 4:00 --> 2:30
# if t_sec > curr_sec and curr_sec == 0:
# print("MAJOR PROBLEM 2: ", t)
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
if t_sec == 30 and curr_sec == 0:
diff_in_past_section = abs(prev_seconds - 30) # 3:15-3
diff_in_mid_1_section = 30 # 3-2:30
diff_in_mid_2_section = 30 # 2:30- 2
diff_in_next_section = abs(30 - t_sec) #2 - 1:30
# 1:30-1
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
next_section_idx = mid_section_2_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
next_section_traj = curr_t_traj[mid_section_2_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
curr_sec = 0
curr_traj = []
elif t_sec == 0 and curr_sec == 30:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:45-3:30
diff_in_mid_1_section = 30 # 3:30-3
diff_in_mid_2_section = 30 # 3- 2:30
diff_in_next_section = abs(30 - t_sec) #2:30 - 2
# 2-1:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_next_section))
next_section_idx = mid_section_2_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
next_section_traj = curr_t_traj[mid_section_2_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = next_section_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
elif t_sec == 30 and curr_sec == 30:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:45-3:30
diff_in_mid_1_section = 30 # 3:30-3
diff_in_mid_2_section = 30 # 3- 2:30
diff_in_mid_3_section = 30 # 2:30 - 2
diff_in_next_section = abs(30 - t_sec) #2 - 1:30
# 1:30-1
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_3_idx = mid_section_2_idx + 1+ int(diff_in_mid_3_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
next_section_idx = mid_section_3_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
mid_section_3_traj = curr_t_traj[mid_section_2_idx+1:mid_section_3_idx+1]
next_section_traj = curr_t_traj[next_section_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_3_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = next_section_traj
curr_sec = 0
curr_traj = []
elif t_sec == 0 and curr_sec == 0:
# To Do
diff_in_past_section = abs(prev_seconds - 30) # 3:15-3:00
diff_in_mid_1_section = 30 # 3:0-2:30
diff_in_mid_2_section = 30 # 2:30- 2
diff_in_mid_3_section = 30 # 2 - 1:30
diff_in_next_section = abs(30 - t_sec) #1:30 - 1
# 1 - 0:30
past_section_idx = int(diff_in_past_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_1_idx = past_section_idx + 1+ int(diff_in_mid_1_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_2_idx = mid_section_1_idx + 1+ int(diff_in_mid_2_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
mid_section_3_idx = mid_section_2_idx + 1+ int(diff_in_mid_3_section/(diff_in_past_section + diff_in_mid_1_section + diff_in_mid_2_section + diff_in_mid_3_section + diff_in_next_section))
next_section_idx = mid_section_3_idx + 1
past_section_traj = curr_t_traj[:past_section_idx+1]
mid_section_1_traj = curr_t_traj[past_section_idx+1:mid_section_1_idx+1]
mid_section_2_traj = curr_t_traj[mid_section_1_idx+1:mid_section_2_idx+1]
mid_section_3_traj = curr_t_traj[mid_section_2_idx+1:mid_section_3_idx+1]
next_section_traj = curr_t_traj[next_section_idx:]
curr_traj.extend(past_section_traj)
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_1_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = mid_section_2_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_min -= 1
curr_sec = 30
curr_traj = mid_section_3_traj
minute_to_traj[(curr_min, curr_sec)] = curr_traj
curr_sec = 0
curr_traj = next_section_traj
curr_min -= 1
curr_sec = 30
curr_traj = []
else:
# print("MAJOR PROBLEM: ", t)
# print("t = ", t)
# print("current = ", (curr_min, curr_sec))
# print("previous = ", (prev_min, prev_seconds))
# print()
to_include = False
prev_min = t_min
prev_seconds = t_sec
stop_indices = np.where(times == 'stop')[0]
if len(stop_indices) > 0:
stop_idx = stop_indices[0]
t = str(times[start_idx])
traj_t = str(traj[start_idx])
msgs_t = str(sent_msgs[start_idx])
vic_t = str(victims[start_idx])
if traj_t != 'nan':
curr_traj.extend([eval(x) for x in traj_t.split(';')])
minute_to_traj[(curr_min, curr_sec)] = curr_traj
return minute_to_traj, to_include
# +
def compute_minute_to_msgs(df_traj):
traj = df_traj['trajectory'].to_numpy()
times = df_traj['time_spent'].to_numpy()
sent_msgs = df_traj['sent_messages'].to_numpy()
victims = df_traj['target'].to_numpy()
to_include = True
minute_to_msgs = {}
curr_min = 5
curr_sec = 0
# minute_to_msgs[(curr_min, curr_sec)] = []
# minute_to_victims[(curr_min, curr_sec)] = []
prev_time = (5,0)
for i in range(len(traj)):
t = str(times[i])
msgs_t = str(sent_msgs[i])
if ':' in t:
t_min = int(t.split(":")[0])
t_sec = int(t.split(":")[1])
prev_time = (t_min, t_sec)
if msgs_t == 'nan':
continue
prev_t_min = prev_time[0]
prev_t_sec = prev_time[1]
window_sec = 0
window_min = prev_t_min
if prev_t_sec > 30:
window_sec = 30
# print((prev_t_min, prev_t_sec), 't = ', (window_min, window_sec))
# print("msgs_t: ", msgs_t)
# print()
keyname = (window_min, window_sec)
if keyname not in minute_to_msgs:
minute_to_msgs[keyname] = []
minute_to_msgs[keyname].append(msgs_t)
# minute_to_traj[(curr_min, curr_sec)] = curr_traj
return minute_to_msgs
# +
def compute_minute_to_victims(df_traj):
traj = df_traj['trajectory'].to_numpy()
times = df_traj['time_spent'].to_numpy()
sent_msgs = df_traj['sent_messages'].to_numpy()
victims = df_traj['target'].to_numpy()
to_include = True
minute_to_victims = {}
curr_min = 5
curr_sec = 0
# minute_to_msgs[(curr_min, curr_sec)] = []
minute_to_victims[(curr_min, curr_sec)] = []
prev_time = (5,0)
for i in range(len(traj)):
t = str(times[i])
victims_t = str(victims[i])
if ':' in t:
t_min = int(t.split(":")[0])
t_sec = int(t.split(":")[1])
prev_time = (t_min, t_sec)
if victims_t in ['nan', 'door']:
continue
prev_t_min = prev_time[0]
prev_t_sec = prev_time[1]
window_sec = 0
window_min = prev_t_min
if prev_t_sec > 30:
window_sec = 30
# print((prev_t_min, prev_t_sec), 't = ', (window_min, window_sec))
# print("victims_t: ", victims_t)
# print()
keyname = (window_min, window_sec)
if keyname not in minute_to_victims:
minute_to_victims[keyname] = []
minute_to_victims[keyname].append(victims_t)
return minute_to_victims
# -
# ## Get Data Dictionaries
uid_to_minute_victims = {}
for i in range(len(valid_uids)):
p_uid = valid_uids[i]
df_traj = df[(df['userid']==p_uid) & (df['episode']==1)]
df_traj = df_traj.sort_values(by=['created_at'], ascending=False)
minute_to_victims = compute_minute_to_victims(df_traj)
# if to_include:
uid_to_minute_victims[p_uid] = minute_to_victims
uid_to_minute_msgs = {}
for i in range(len(valid_uids)):
p_uid = valid_uids[i]
df_traj = df[(df['userid']==p_uid) & (df['episode']==1)]
df_traj = df_traj.sort_values(by=['created_at'], ascending=False)
minute_to_msgs = compute_minute_to_msgs(df_traj)
# if to_include:
uid_to_minute_msgs[p_uid] = minute_to_msgs
uid_to_minute_traj = {}
for i in range(len(valid_uids)):
p_uid = valid_uids[i]
df_traj = df[(df['userid']==p_uid) & (df['episode']==1)]
df_traj = df_traj.sort_values(by=['time_spent'], ascending=False)
minute_to_traj, to_include = compute_minute_to_traj(df_traj)
if to_include:
uid_to_minute_traj[p_uid] = minute_to_traj
# ## Process Data Into Metrics
def l2(x1, x2):
return np.sqrt((x1[0] - x2[0])**2 + (x1[1] - x2[1])**2)
# +
def compute_effort(traj_list, robot_list):
eff = 0
for i in range(len(traj_list)-1):
eff += l2(traj_list[i], traj_list[i+1])
eff_robot = 0
for i in range(len(robot_list)-1):
eff_robot += l2(robot_list[i], robot_list[i+1])
eff = eff + eff_robot
return eff
# +
uid_min_to_data = {}
for puid in uid_to_minute_traj:
uid_min_to_data[puid] = {}
human_traj = uid_to_minute_traj[puid]
robot_traj = min_to_robot_traj
human_msgs = uid_to_minute_msgs[puid]
human_victims = uid_to_minute_victims[puid]
counter = 0
for minute in [4,3,2, 1, 0]:
for second in [30, 0]:
keyname = (minute, second)
uid_min_to_data[puid][counter] = {}
if keyname not in human_traj:
curr_human_traj = []
else:
curr_human_traj = human_traj[keyname]
curr_robot_traj = robot_traj[keyname]
effort = compute_effort(curr_human_traj, curr_robot_traj)
uid_min_to_data[puid][counter]['effort'] = effort
if keyname not in human_msgs:
num_msgs = 0
else:
num_msgs = len(human_msgs[keyname])
uid_min_to_data[puid][counter]['msgs'] = num_msgs
if keyname not in human_victims:
num_victims = 0
else:
num_victims = len(human_victims[keyname])
uid_min_to_data[puid][counter]['victims'] = num_victims
counter += 1
# -
uid_min_to_data
# # Split Into Groups
data_by_group = {1:{}, 2:{}, 3:{}, 4:{}}
for p_uid in uid_min_to_data:
# print(p_uid)
# print(uid_min_to_data[puid])
group_no = uid_to_group[p_uid]
data_by_group[group_no][p_uid] = uid_min_to_data[p_uid]
# break
# # Binarize Data
# +
group_to_means = {1:{}, 2:{}, 3:{}, 4:{}}
for group in data_by_group:
group_data = data_by_group[group]
all_effort = {i:[] for i in range(10)}
all_victims = {i:[] for i in range(10)}
all_msgs = {i:[] for i in range(10)}
for p_uid in group_data:
for i in range(10):
all_effort[i].append(group_data[p_uid][i]['effort'])
all_msgs[i].append(group_data[p_uid][i]['msgs'])
all_victims[i].append(group_data[p_uid][i]['victims'])
# print(all_victims)
final_effort = {i:np.mean(all_effort[i]) for i in range(10)}
final_victims = {i:np.mean(all_victims[i]) for i in range(10)}
final_msgs = {i:np.mean(all_msgs[i]) for i in range(10)}
group_to_means[group]['effort'] = final_effort
group_to_means[group]['victims'] = final_victims
group_to_means[group]['msgs'] = final_msgs
# +
group_to_binary_data = {}
for group in data_by_group:
group_data = data_by_group[group]
mean_effort = group_to_means[group]['effort']
mean_victims = group_to_means[group]['victims']
mean_msgs = group_to_means[group]['msgs']
new_group_data = {}
for p_uid in group_data:
p_uid_data = []
for i in range(10):
effort_binary = 0 if group_data[p_uid][i]['effort'] < mean_effort[i] else 1
msgs_binary = 0 if group_data[p_uid][i]['msgs'] < mean_msgs[i] else 1
victims_binary = 0 if group_data[p_uid][i]['victims'] < mean_victims[i] else 1
p_uid_data.append((effort_binary, msgs_binary, victims_binary))
new_group_data[p_uid] = p_uid_data
group_to_binary_data[group] = new_group_data
# +
group_to_binary_state_data = {}
group_to_state_list = {}
for group in data_by_group:
group_data = data_by_group[group]
group_to_state_list[group] = []
mean_effort = group_to_means[group]['effort']
mean_victims = group_to_means[group]['victims']
mean_msgs = group_to_means[group]['msgs']
new_group_data = {}
for p_uid in group_data:
p_uid_data = []
for i in range(9):
effort_binary = 0 if group_data[p_uid][i]['effort'] < mean_effort[i] else 1
msgs_binary = 0 if group_data[p_uid][i]['msgs'] < mean_msgs[i] else 1
victims_binary = 0 if group_data[p_uid][i]['victims'] < mean_victims[i] else 1
effort_binary_next = 0 if group_data[p_uid][i+1]['effort'] < mean_effort[i+1] else 1
msgs_binary_next = 0 if group_data[p_uid][i+1]['msgs'] < mean_msgs[i+1] else 1
victims_binary_next = 0 if group_data[p_uid][i+1]['victims'] < mean_victims[i+1] else 1
state_vector = (effort_binary_next, msgs_binary_next, victims_binary_next, effort_binary, msgs_binary, victims_binary)
p_uid_data.append(state_vector)
if state_vector not in group_to_state_list[group]:
group_to_state_list[group].append(state_vector)
new_group_data[p_uid] = p_uid_data
group_to_binary_state_data[group] = new_group_data
# +
group_to_state_mapping = {}
for group in group_to_state_list:
group_to_state_mapping[group]= {}
state_id_to_state = dict(enumerate(group_to_state_list[group]))
state_to_state_id = {v: k for k, v in state_id_to_state.items()}
group_to_state_mapping[group]['id_to_vec'] = state_id_to_state
group_to_state_mapping[group]['vec_to_id'] = state_to_state_id
# -
# # Generate States
# +
group_to_state_id_data_w_pid = {}
group_to_state_id_data = {}
for group in group_to_binary_state_data:
group_data = group_to_binary_state_data[group]
state_to_state_id = group_to_state_mapping[group]['vec_to_id']
all_data = []
all_data_w_pid = {}
for p_uid in group_data:
state_data = [state_to_state_id[elem] for elem in group_data[p_uid]]
all_data.append(state_data)
all_data_w_pid[p_uid] = state_data
group_to_state_id_data[group] = all_data
group_to_state_id_data_w_pid[group] = all_data_w_pid
# -
group_to_state_id_data
# +
import pickle
with open('minimap_group_to_state_data.pickle', 'rb') as handle:
group_to_state_id_data = pickle.load(handle)
with open('minimap_group_to_state_data_w_pid.pickle', 'rb') as handle:
group_to_state_id_data_w_pid = pickle.load(handle)
with open('minimap_group_to_binary_state_data.pickle', 'rb') as handle:
group_to_binary_state_data = pickle.load(handle)
with open('minimap_group_to_state_mapping.pickle', 'rb') as handle:
group_to_state_mapping = pickle.load(handle)
# +
import pickle
with open('minimap_group_to_state_data.pickle', 'wb') as handle:
pickle.dump(group_to_state_id_data, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('minimap_group_to_state_data_w_pid.pickle', 'wb') as handle:
pickle.dump(group_to_state_id_data_w_pid, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('minimap_group_to_binary_state_data.pickle', 'wb') as handle:
pickle.dump(group_to_binary_state_data, handle, protocol=pickle.HIGHEST_PROTOCOL)
with open('minimap_group_to_state_mapping.pickle', 'wb') as handle:
pickle.dump(group_to_state_mapping, handle, protocol=pickle.HIGHEST_PROTOCOL)
# -
| minimap_data/ProcessMinimapData-PositiveEmotion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: vocabulary_learning
# language: python
# name: vocabulary_learning
# ---
# +
# %load_ext nb_black
# %load_ext autoreload
# %autoreload 2
import os
print(os.getcwd())
def update_working_directory():
from pathlib import Path
p = Path(os.getcwd()).parents[0]
os.chdir(p)
print(p)
update_working_directory()
# -
path_dataset_train = "data/raw/20201009/dataset_train.pkl"
path_dataset_valid = "data/raw/20201009/dataset_valid.pkl"
# # Import
# +
import dill
import numpy as np
import pandas as pd
pd.set_option("display.max_columns", None)
from src.models.logistic_regression import ModelLogisticRegression
import src.models.performance_metrics as performance_metrics
# -
# # Dataset
# +
with open(path_dataset_train, "rb") as input_file:
dataset_train = dill.load(input_file)
with open(path_dataset_valid, "rb") as input_file:
dataset_valid = dill.load(input_file)
# -
# # Overall
# +
model = ModelLogisticRegression()
model.version
# -
dataset_train = model.preprocessing_training(dataset_train)
model.train(dataset_train)
model.version
with open(f"models/{model.version}__model.pkl", "wb") as file:
dill.dump(model, file)
# + [markdown] heading_collapsed=true
# # Data Transformation
# + hidden=true
vardict.keys()
# + [markdown] hidden=true
# ## Target
# + hidden=true
dataset_train[vardict["target"]].describe()
# + [markdown] hidden=true
# ## Numerical
# + hidden=true
dataset_train[vardict["numerical"]].isnull().sum()
# + hidden=true
def data_transform_numerical(dataset, vardict):
dataset["previous_levenshtein_distance_guess_answer"].fillna(-1, inplace=True)
dataset["previous_question_time"].fillna(-1, inplace=True)
dataset["previous_write_it_again_german"].fillna(-1, inplace=True)
dataset["previous_write_it_again_english"].fillna(-1, inplace=True)
return dataset, vardict
# + [markdown] hidden=true
# ## Diff time
# + hidden=true
dataset_train[vardict["diff_time"]].isnull().sum()
# + hidden=true
def data_transform_diff_time(dataset, vardict):
dataset["days_since_last_occurrence_same_language"].fillna(-1, inplace=True)
dataset["days_since_last_occurrence_any_language"].fillna(-1, inplace=True)
dataset["days_since_last_success_same_language"].fillna(-1, inplace=True)
dataset["days_since_last_success_any_language"].fillna(-1, inplace=True)
dataset["days_since_first_occur_same_language"].fillna(-1, inplace=True)
dataset["days_since_first_occur_any_language"].fillna(-1, inplace=True)
return dataset, vardict
# + [markdown] hidden=true
# ## Boolean
# + hidden=true
dataset_train[vardict["boolean"]]
# + hidden=true
def data_transform_boolean(dataset, vardict):
# Transform to dummies
vardict["dummy_boolean"] = []
for i_var_boolean in vardict["boolean"]:
# possible improvement: pandas.get_dummies(drop_first=False)
i_dummy_boolean = pd.get_dummies(
dataset[i_var_boolean],
prefix=i_var_boolean,
prefix_sep="__",
dummy_na=True,
)
del dataset_train[i_var_boolean]
vardict["dummy_boolean"] = (
vardict["dummy_boolean"] + i_dummy_boolean.columns.tolist()
)
dataset = pd.concat([dataset, i_dummy_boolean], axis=1)
dataset[vardict["dummy_boolean"]].describe()
return dataset, vardict
# + [markdown] hidden=true
# ## Categorical
# + hidden=true
dataset_train[vardict["categorical"]]
# + hidden=true
def data_transform_categorical(dataset, vardict):
# Transform to dummies
vardict["dummy_categorical"] = []
for i_var_categorical in vardict["categorical"]:
# possible improvement: pandas.get_dummies(drop_first=False)
i_dummy_categorical = pd.get_dummies(
dataset[i_var_categorical],
prefix=i_var_categorical,
prefix_sep="__",
dummy_na=True,
)
del dataset[i_var_categorical]
vardict["dummy_categorical"] = (
vardict["dummy_categorical"] + i_dummy_categorical.columns.tolist()
)
dataset = pd.concat([dataset, i_dummy_categorical], axis=1)
return dataset, vardict
# + [markdown] hidden=true
# ## Overall
# + hidden=true
dataset_train, vardict = data_transform_numerical(dataset_train, vardict)
dataset_train, vardict = data_transform_diff_time(dataset_train, vardict)
dataset_train, vardict = data_transform_boolean(dataset_train, vardict)
dataset_train, vardict = data_transform_categorical(dataset_train, vardict)
# + [markdown] hidden=true
# ### vardict
# + hidden=true
vardict["all"] = (
vardict["numerical"]
+ vardict["diff_time"]
+ vardict["dummy_boolean"]
+ vardict["dummy_categorical"]
)
# + [markdown] heading_collapsed=true
# # 1st model
# + hidden=true
X_train = dataset_train[vardict["all"]]
y_train = dataset_train[vardict["target"]]
# + hidden=true
X_train
# + hidden=true
y_train
# + hidden=true
model = LogisticRegression(random_state=0)
# + hidden=true
model.fit(X_train, y_train)
# + hidden=true
with open(f"data/processed/{model_name}_model.pkl", "wb") as file:
dill.dump(model, file)
with open(f"data/processed/{model_name}_vardict.pkl", "wb") as file:
dill.dump(vardict, file)
# + hidden=true
# -
# # Validation results
dataset_valid = model.preprocessing_inference(dataset_valid)
predictions = model.predict(dataset=dataset_valid)
# +
binary_classification_results = performance_metrics.get_binary_classification_results(
predictions, model_name=f"{model.version}_valid"
)
binary_classification_results
# +
regression_results = performance_metrics.get_regression_results(
predictions, model_name=f"{model.version}_valid"
)
regression_results
# -
performance_metrics.plot_roc_auc_curve(predictions, model_name=f"{model.version}_valid")
performance_metrics.plot_precision_recall_curve(
predictions, binary_classification_results, model_name=f"{model.version}_valid"
)
| notebooks/past/Modeling - 1.1 - Logistic Regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# +
def Conv(arr,filt,stride,pad_width,pad_value=0):
width = arr.shape[0]
height = arr.shape[1]
filt_size = filt.shape[0]
row = col = 0
if(len(arr.shape) == 2):
arr = arr.reshape(arr.shape[0],arr.shape[1],1)
if(len(filt.shape) == 2):
filt = filt.reshape(filt.shape[0],filt.shape[1],1)
assert(filt.shape[2]==arr.shape[2]) , "different no of channels : " + str(filt.shape[2]) + " , " + str(arr.shape[2])
#padding
arr = np.pad(arr,((pad_width,pad_width),(pad_width,pad_width),(0,0)),'constant',constant_values=(pad_value,pad_value))
# determine the size of the array obtained after convolution
new_width = np.int(((width + (2*pad_width) - filt_size)/stride) + 1)
new_height = np.int(((height + (2*pad_width) - filt_size)/stride) + 1)
new_arr = np.zeros(( new_width, new_height))
# perform the convolution operation
for i in range(0,width,stride):
col = 0
for j in range(0,height,stride):
conv = arr[i:i+filt_size,j:j+filt_size,:]
if(conv.shape == filt.shape):
conv = conv * filt
conv = conv.reshape(conv.shape[0]*conv.shape[1]*conv.shape[2])
new_arr[row][col] = np.sum(conv)
col = (col+1)%new_height
row+=1
return new_arr
def Pool(arr,filt_size,method,stride):
width = arr.shape[0]
height = arr.shape[1]
row = col = 0
if(len(arr.shape) == 2):
arr = arr.reshape(arr.shape[0],arr.shape[1],1)
# determine the size of the array obtained after convolution
new_width = np.int(((width - filt_size)/stride) + 1)
new_height = np.int(((height - filt_size)/stride) + 1)
new_arr = np.zeros(( new_width, new_height , arr.shape[2]))
# perform max pooling
for ch in range(arr.shape[2]):
row = 0
for i in range(0,width,stride):
col = 0
for j in range(0,height,stride):
conv = arr[i:i+filt_size,j:j+filt_size,ch]
if(conv.shape[:2] == (filt_size,filt_size)):
if(method =="max"):
pool = np.max(conv)
elif(method =="mean"):
pool = np.mean(conv)
new_arr[row , col , ch] = pool
col = (col+1)%new_height
row+=1
if(new_arr.shape[2]==1):
new_arr = new_arr.reshape(new_arr.shape[0],new_arr.shape[1])
return new_arr
def forward(layer_info,X):
if(len(X.shape) == 3):
X = X.reshape(1,X.shape[0],X.shape[1],X.shape[2])
if(len(X.shape) == 2):
X = X.reshape(1,X.shape[0],X.shape[1])
array = X
cache = []
for layer , info in layer_info.items():
cache.append(array)
if(info['layer'] == "Conv"):
if(len(info['filters'].shape) == 3):
info['filters'] = info['filters'].reshape(1,info['filters'].shape[0],info['filters'].shape[1],info['filters'].shape[2])
elif(len(info['filters'].shape) == 2):
info['filters'] = info['filters'].reshape(1,info['filters'].shape[0],info['filters'].shape[1])
array = np.array([np.array([Conv(arr,k,stride=info['stride'],pad_width=info['pad'],pad_value=info['pad value']).T for k in info['filters']]).T for arr in array]) + info['bias']
elif(info['layer'] == "Pool"):
array = np.array([Pool(arr,filt_size=info['filt size'],method = info["method"],stride=info['stride']) for arr in array])
return array , cache
def backward(dZ , layer_info , cache , l_rate):
keys = list(reversed(list(layer_info.keys())))
cache = list(reversed(cache))
for key , A in zip(keys , cache):
dA = np.zeros(A.shape)
if(layer_info[key]['layer'] == 'Conv'):
# get info of hyperparameters
pad_width ,pad_value ,stride = (layer_info[key]['pad'] , layer_info[key]['pad value'] , layer_info[key]['stride'])
W = layer_info[key]['filters']
dW = np.zeros(layer_info[key]['filters'].shape)
filt_size = W.shape[1]
# pad both X and dX and store them in new set of variables
A_pad = np.pad(A,((0,0),(pad_width,pad_width),(pad_width,pad_width),(0,0)),'constant',constant_values=0)
dA_pad = np.pad(dA,((0,0),(pad_width,pad_width),(pad_width,pad_width),(0,0)),'constant',constant_values=0)
(m , height , width , ch) = dZ.shape
#loop over samples , height and width of each image
for i in range(m):
for h in range(0,height,stride):
for w in range(0,width,stride):
# slice the array needed for convolution
sliced = A_pad[i ,h:h+filt_size , w:w+filt_size, :]
#calculate the derivatives dW and dA
dW+= sliced * np.sum(dZ[i,h,w,:])
dA_pad[i ,h:h+filt_size , w:w+filt_size, :]+= np.sum(W[:,:,:,:] * np.sum(dZ[i,h,w,:]),axis=0)
# once dx_pad is calculated , assign only the unpadded part to the derivative dX
dA[i,:,:,:] = dA_pad[i , pad_width:-pad_width, pad_width:-pad_width, :]
db = dZ.sum(axis=(0,1,2))
layer_info[key]['filters'] -= (l_rate / m) * dW
layer_info[key]['bias'] -= (l_rate / m) * db
dZ = dA
elif(layer_info[key]['layer'] == 'Pool'):
# get info of hyperparameters
filt_size , stride , method = (layer_info[key]['filt size'] , layer_info[key]['stride'] , layer_info[key]['method'])
# get dimensions of previous layer's derivatives
(m , height , width , ch) = dZ.shape
for i in range(m):
for h in range(0,height,stride):
for w in range(0,width,stride):
# do backward pass for max pooling
if(method == 'max'):
# slice the array needed for convolution
sliced = A[i ,h:h+filt_size , w:w+filt_size, :]
#create a mask for the sliced array
mask = (sliced == np.max(sliced))
dA[i ,h:h+filt_size , w:w+filt_size, :] += mask * dA[i, h, w,:].max()
#do backward pass for mean pooling
elif(method == 'mean'):
# slice the array needed for convolution
sliced = dA[i,h,w,:]
# obtain the distributed value for the sliced array
shape = (filt_size , filt_size)
average = sliced / (h * w)
dis_value = np.ones(shape) * average
dA[i ,h:h+filt_size , w:w+filt_size, :] += dis_value
dZ = dA
return layer_info
# +
np.random.seed(1)
layer_info = {
"l1" : {
"layer" : "Conv" ,
"pad" : 1 ,
"pad value" : 0 ,
"filters" : np.random.rand(10,3,3,3) , #(no of samples , height , width , channels) , min input to be given -> (height , width)
"bias" : np.random.rand(1,1,1,10) ,
"stride" : 2
} ,
"l2" : {
"layer" : "Pool" ,
"filt size" : 3 ,
"stride" : 1 ,
"method" : "max" #mention the method to pool i.e "max" or "mean" pooling
}
}
X = np.random.randn(40,10,10,3)
output,cache = forward(layer_info , X) #(no of samples , height , width , channels) , min input to be given -> (height , width)
# -
output.shape
# +
np.random.seed(1)
y_true = np.random.randn(output.shape[0],output.shape[1],output.shape[2],output.shape[3])
dZ = y_true - output
layer_info_mod = backward(dZ ,layer_info , cache , 1)
| ConvNet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Sales commissions forecast
# ## Problem Definition
# A company with 5 sales managers wants to provision budget to pay sales commissions to sales managers. The company applies a commission rate based on the percentage of target sales obtained, given by the following table:
#
# | Sales / Target ratio | Commission Rate |
# |---------------------- |----------------- |
# | 0-90 % | 2% |
# | 90-100% | 3% |
# | >= 100% | 4% |
#
# + [markdown] slideshow={"slide_type": "fragment"}
# Each salesman will be compensanted with the commission rate times the total sales obtained. The following table shows the target sales for the five sales managers:
#
# | Sales Manager | Sales Target (€) |
# |--------------- |------------------ |
# | 1 | 100,000 |
# | 2 | 200,000 |
# | 3 | 75,000 |
# | 4 | 400,000 |
# | 5 | 500,000 |
#
#
# + [markdown] slideshow={"slide_type": "subslide"}
# **a)** Estimate the budget for sales commissions the company has to pay in the scenario where all sales managers get exactly the 100% of the sales target (naive approach).
#
# **b)** The company has a historic record of sales for the five sales managers and from this record, it can estimate that the Percent to Plan (The ratio between the actual sales and the sales target) can be modelled by a normal distribution with a mean of 100% and standard deviation of 10%. Use this insight to estimate the budget for sales commissions using a MonteCarlo distribution.
# + [markdown] slideshow={"slide_type": "slide"}
# ## Solution
# **a)** In the requested scenario, the sales obtained by each sales manager are represented in the table below:
#
#
# | Sales Manager | Sales Target (€) | Actual Sales (€) | Percent to Plan (%) | Commission Rate (€) | Commission Amount (€) |
# |--------------- |------------------ |------------------ |--------------------- |--------------------- |----------------------- |
# | 1 | 100,000 | 100,000 | 100 | 4 | 4,000 |
# | 2 | 200,000 | 200,000 | 100 | 4 | 8,000 |
# | 3 | 75,000 | 75,000 | 100 | 4 | 3,000 |
# | 4 | 400,000 | 400,000 | 100 | 4 | 16,000 |
# | 5 | 500,000 | 500,000 | 100 | 4 | 20,000 |
#
# The total budget for sales commission can be obtained with the summation of the last column (51,000€)
# + [markdown] slideshow={"slide_type": "subslide"}
# **b)** In order to estimate the budget using Montecarlo, we are going to use the Python numpy package to calculate the probability distribution.
# + [markdown] slideshow={"slide_type": "subslide"}
# First we import the libraries we are going to use:
# + slideshow={"slide_type": "fragment"} pycharm={"is_executing": false}
import pandas as pd
import numpy as np
# + [markdown] slideshow={"slide_type": "subslide"}
# Then we initialise the data needed to model the problem
# + slideshow={"slide_type": "fragment"} pycharm={"is_executing": false}
avg = 1
std_dev = .1
num_simulations = 1000
sales_target_values = np.array([100000, 200000, 75000, 400000, 500000])
# Define a function to calculate the commission rate depending on the rate to target
def calc_com_rate(x):
if x <= 0.9:
return 0.02
elif x <= 1:
return 0.03
else:
return 0.04
# You can also use a lambda:
# calc_com_rate = lambda x: 0.02 if x <= 0.9 else 0.03 if x <= 0.99 else 0.04
# Vectorize the function so that we can apply it to vectors and matrices
v_calc_com_rate = np.vectorize(calc_com_rate)
# Define a list to keep all the results from each simulation that we want to analyze
all_stats = np.zeros((num_simulations, 3))
# + [markdown] slideshow={"slide_type": "subslide"}
# Now we run the simulations in a for loop:
# + slideshow={"slide_type": "fragment"} pycharm={"is_executing": false}
# Loop through simulations
for i in range(num_simulations):
# Choose random inputs for the sales targets and percent to target
pct_to_target = np.random.normal(avg, std_dev, len(sales_target_values))
#Calculate actual sales
sales = pct_to_target*sales_target_values
# Determine the commissions rate and calculate it
commission_rate = v_calc_com_rate(np.array(pct_to_target))
# Calculate the commission
commission = sales*commission_rate
# We want to track sales,commission amounts and sales targets over all the simulations
# Sum values among sales managers and calculate the mean commission rate
all_stats[i,:] = [np.sum(sales),
np.sum(commission),
np.mean(commission_rate)]
results_df = pd.DataFrame.from_records(all_stats, columns=['Sales',
'Commission_Amount',
'Commission_Rate'])
# + [markdown] slideshow={"slide_type": "subslide"}
# Finally, we represent the results and calculate the confidence interval:
# + slideshow={"slide_type": "fragment"} pycharm={"is_executing": false}
results_df.describe()
# + pycharm={"is_executing": false}
hist = results_df.hist(bins=100)
# + slideshow={"slide_type": "subslide"} pycharm={"is_executing": false}
import scipy.stats as st
#Calculate the 95% confidence interval
# We collect the results from the data frame
a = np.array(results_df['Commission_Amount'])
# loc is used to center distribution at mean of array
# scale is used to scale the distribution according to the standard error
# of the mean (st.sem)
arr_standard_dev = np.std(a)/(len(a)**0.5)
arr_mean = np.mean(a)
interval = (st.norm.ppf(0.025, loc= arr_mean, scale=arr_standard_dev), st.norm.ppf(0.975, loc= arr_mean, scale=arr_standard_dev))
# + slideshow={"slide_type": "skip"} pycharm={"is_executing": false}
import pandas as pd
import numpy as np
avg = 1
std_dev = .1
num_reps = 500
num_simulations = 1000
sales_target_values = np.array([100000, 200000, 75000, 400000, 500000])
# Define a lambda function to calculate the ratio
calc_com_rate = lambda x: 0.02 if x <= 0.9 else 0.03 if x <= 0.99 else 0.04
v_calc_com_rate = np.vectorize(calc_com_rate)
# Choose random inputs for the sales targets and percent to target,
# this time create a matrix with as many rows as simulations
pct_to_target = np.random.normal(avg, std_dev, (num_simulations, len(sales_target_values)))
# Reshape the sales target values into a matrix of adequate size
stv = np.broadcast_to(sales_target_values, (num_simulations, len(sales_target_values)))
# Calculate the sales applying the ratio
sales = pct_to_target*stv
# Calculate commission rate
commission_rate = v_calc_com_rate(pct_to_target)
# And commission
commission = sales*commission_rate
# Sum values among sales managers and calculate the mean commission rate
all_stats = [np.sum(sales, axis=1), np.sum(commission, axis=1), np.mean(commission_rate, axis=1)]
results_df = pd.DataFrame.from_records(np.transpose(all_stats), columns=['Sales',
'Commission_Amount',
'Commission_Rate'])
results_df.describe()
| docs/source/Simulation/Solved/Sales commissions forecast.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 1
# From `math` package import function `ceil()`. `ceil(x)` is equivalent of ceiling function in mathematics where it gives the closest integer bigger than x.
# Your answer goes here
ceil(5.1)
ceil(-1.5)
# Note: If you are getting an error in evaluating the above expressions it could be because you haven't explicitly imported the module content.
# # Exercise 2
# Import package `peewee`.
# Your answer goes here
# You will most likely get an error indicating that this packages does not exist. Try to install it by
#
# `conda install peewee` or `pip install peewee`. Note that these commands are for command line.
#
# Hint: You can use a `!` prior to any command in a Jupyter cell to make that line a command-line command.
# Your answer goes here
# After a successful installation you should be able to import it.
| 00-Python-Basics/Exercise-08-Modules-and-Packages.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Sentiment Analysis of Product Reviews
# ### Import necessary libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import re, string
import nltk
from nltk.corpus import stopwords
from nltk.tokenize import word_tokenize
from nltk.corpus import wordnet
from nltk import pos_tag
from nltk.stem import WordNetLemmatizer
import spacy
from wordcloud import WordCloud, STOPWORDS, ImageColorGenerator
from collections import Counter
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.model_selection import cross_validate
from sklearn.model_selection import train_test_split
from sklearn.model_selection import StratifiedKFold
from sklearn.metrics import classification_report
from sklearn.metrics import confusion_matrix
from sklearn.metrics import roc_auc_score
from sklearn.utils import shuffle
from matplotlib.colors import LinearSegmentedColormap
import seaborn as sns
from sklearn.naive_bayes import MultinomialNB
from xgboost import XGBClassifier
from nltk.stem.porter import PorterStemmer
from tqdm import tqdm
from sklearn.feature_selection import chi2
from sklearn.feature_selection import SelectKBest
from sklearn.model_selection import GridSearchCV
import time, datetime
from sklearn.metrics import make_scorer, accuracy_score, precision_score, recall_score, f1_score
from sklearn.svm import LinearSVC
from sklearn.neighbors import KNeighborsClassifier
# ### Read the Dataset
# +
#reading already processede data
df = pd.read_csv("preprocessed-dataset.csv")
df = df.dropna(how='any',axis=0)
# -
# ## Feature Engineering and Selection
# ### Create TF-IDF
vectorizer = TfidfVectorizer(max_features=7000)
features = vectorizer.fit_transform(df['text'])
tf_idf = pd.DataFrame(features.toarray(), columns=vectorizer.get_feature_names())
# ### Splitting Dataset into Train and Test Set
# We did 80:20 split for training and test
X_train, X_test, y_train, y_test = train_test_split(tf_idf, df['sentiment'], test_size=0.2, random_state=42)
yy=pd.DataFrame(y_train)
train_data = pd.concat([X_train,yy],axis=1)
# ## Oversampling the Train Data
target_count = train_data['sentiment'].value_counts()
negative_class = train_data[train_data['sentiment'] == 0]
positive_class = train_data[train_data['sentiment'] == 1]
negative_over = negative_class.sample(target_count[1], replace=True)
df_train_over = pd.concat([positive_class, negative_over], axis=0)
df_train_over = shuffle(df_train_over)
counts=df_train_over['sentiment'].value_counts()
plt.title("Train Classes count after Oversampling")
plt.bar(counts.index, counts.values)
plt.show()
# # Final Data for Train-Testing
X_train=df_train_over.iloc[:,:-1]
y_train=df_train_over['sentiment']
# ## Modeling
Here we are defining our models with list of values for parameters to find the best value using GridSearchCV
models_with_default_params=[{'mod' : MultinomialNB(), 'param': {'alpha': [10**-5,10**-4,10**-3,10**-2,10**-1,1,1.5,2]}},
{'mod': LinearSVC(), 'param': {'C': [0.1, 1, 10]}},
{'mod': KNeighborsClassifier(), 'param': {'n_neighbors': [1,2,3]}},
{'mod': XGBClassifier(), 'param': {'n_estimators': [100]}}]
X_train.replace(np.NaN, 0, inplace=True)
# # Without Chi-Square Feature Reduction
# +
X_train_vect = X_train
X_test_vect = X_test
for mwdp in models_with_default_params:
SVM_grid_search = GridSearchCV(mwdp['mod'], mwdp['param'], refit=True, verbose=3)
SVM_grid_search.fit(X_train_vect, y_train)
#considering the best model using GridSearchCV
model = SVM_grid_search.best_estimator_
print('---------'+'Model: '+model.__class__.__name__+'---------')
print('Feature Vector Size:',X_train_vect.shape)
print('Best Model: ',model)
train_start_time = datetime.datetime.now()
model.fit(X_train_vect, y_train)
#Calculating the training time
print('TRAIN TIME: ', datetime.datetime.now() - train_start_time)
y_pred = model.predict(X_test_vect)
print('Accuracy: ',accuracy_score(y_test, y_pred))
print('Precision: ',precision_score(y_test, y_pred, average="macro"))
print('Recall: ',recall_score(y_test, y_pred, average="macro"))
print('F1 Score: ',f1_score(y_test, y_pred, average="macro"))
# -
# # With Chi-Square Feature Reduction
# +
X_train_vect = X_train
X_test_vect = X_test
#Reducing no of features with chi-square
chi_selector = SelectKBest(score_func=chi2, k=500)
X_train_vect_chi=chi_selector.fit_transform(X_train_vect, y_train)
X_test_vect_chi=chi_selector.transform(X_test_vect)
for mwdp in models_with_default_params:
SVM_grid_search = GridSearchCV(mwdp['mod'], mwdp['param'], refit=True, verbose=3)
SVM_grid_search.fit(X_train_vect_chi, y_train)
#considering the best model using GridSearchCV
model = SVM_grid_search.best_estimator_
print('---------'+'Model: '+model.__class__.__name__+'---------')
print('Feature Vector Size:',X_train_vect_chi.shape)
print('Best Model: ',model)
train_start_time = datetime.datetime.now()
model.fit(X_train_vect_chi, y_train)
#Calculating the training time
print('TRAIN TIME: ', datetime.datetime.now() - train_start_time)
y_pred = model.predict(X_test_vect_chi)
print('Accuracy: ',accuracy_score(y_test, y_pred))
print('Precision: ',precision_score(y_test, y_pred, average="macro"))
print('Recall: ',recall_score(y_test, y_pred, average="macro"))
print('F1 Score: ',f1_score(y_test, y_pred, average="macro"))
| src/classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %config Completer.use_jedi = False
import networkx as nx
import community as c
import pandas as pd
import matplotlib.pyplot as plt
import os as os
import numpy as np
import random
import seaborn as sns
import time
import operator
from plotly.offline import download_plotlyjs, init_notebook_mode, iplot
import plotly.graph_objs as go
import plotly
from IPython import display
random.seed(246)
# -
df2019q3 = pd.read_csv("2019Q3_Filtered.csv")
investors=pd.read_csv("investors.csv")
df2019q3[df2019q3.name.isin(investors["investor"])]
c=df2019q3.groupby(['issuers','name'])['shares'].sum().rename('count')
p=c/c.groupby(level=0).sum()
p.df = p.reset_index()
onepercent = p.df[p.df["count"]>.01]
c=df2019q3.groupby(['issuers','name'])['shares'].sum().rename('count')
onep_list = onepercent[["name","issuers","count"]]
G = nx.Graph(seed=15)
for i in range(len(onep_list)):
e1=onep_list.iloc[i, 0]
e2=onep_list.iloc[i, 1]
e3=onep_list.iloc[i, 2]
G.add_nodes_from([e1], bipartite=0)
G.add_nodes_from([e2], bipartite=1)
G.add_edges_from([(e1, e2)])
# +
# Create pos: dictionary keyed by node with node positions as values.
pos = nx.spring_layout(G, k=0.1, seed=2)
for n, p in pos.items():
G.nodes[n]['pos'] = p
# +
val_map = {'Pfizer':2.0,
'Astrazeneca': 2.0,
'Moderna': 2.0,
'Sinovac': 3.0,
'Curevac': 1.0,
'Merck': 1.0,
'Themis': 1.0,
'Glaxosmithkline': 1.0,
'Novartis': 1.0,
'Roche': 1.0,
'Sanofi': 1.0,
'Bayer':1.0
}
values = [val_map.get(node, 0.25) for node in G.nodes()]
# +
# Add edges as disconnected lines in a single trace and nodes as a scatter trace.
edge_x = []
edge_y = []
for edge in G.edges():
x0, y0 = G.nodes[edge[0]]['pos']
x1, y1 = G.nodes[edge[1]]['pos']
edge_x.append(x0)
edge_x.append(x1)
edge_x.append(None)
edge_y.append(y0)
edge_y.append(y1)
edge_y.append(None)
edge_trace = go.Scatter(
x=edge_x, y=edge_y,
line=dict(width=0.4, color='#888'),
hoverinfo='text',
mode='lines')
node_x = []
node_y = []
for node in G.nodes():
x, y = G.nodes[node]['pos']
node_x.append(x)
node_y.append(y)
node_trace = go.Scatter(
x=node_x, y=node_y,
mode='markers',
hoverinfo='text',
marker=dict(
showscale=False,
reversescale=False,
color=values,
size=10,
line_width=2))
# +
# Color Node Points by the number of connections.
node_text = []
weights = nx.get_edge_attributes(G,'count').values()
weights2 =list(weights)
weights2 = [i * 15 for i in weights2]
for node, adjacencies in enumerate(G.adjacency()):
node_text.append(adjacencies[0] + ', # of connections: '+str(len(adjacencies[1])))
node_trace.marker.color = values
node_trace.text = node_text
# Node size by betweenness centrality
betCent = nx.betweenness_centrality(G, normalized=True, endpoints=True)
node_size = [v * 80 for v in betCent.values()]
node_trace.marker.size=node_size
# +
# Color Node Points by the number of connections.
node_text = []
for node, adjacencies in enumerate(G.adjacency()):
node_text.append(adjacencies[0] + ', # of connections: '+str(len(adjacencies[1])))
node_trace.marker.color = values
node_trace.text = node_text
# Node size by betweenness centrality
betCent = nx.betweenness_centrality(G, normalized=True, endpoints=True)
node_size = [v * 80 for v in betCent.values()]
node_trace.marker.size=node_size
# +
# Create Network Graph
fig = go.Figure(data=[edge_trace, node_trace],
layout=go.Layout(
title='<br>2019 Q3',
titlefont_size=12,
showlegend=False,
hovermode='closest',
margin=dict(b=20,l=5,r=5,t=40),
annotations=[ dict(
text=' ',
showarrow=False,
xref="paper", yref="paper",
x=0.005, y=-0.002 ) ],
xaxis=dict(showgrid=False, zeroline=False, showticklabels=False),
yaxis=dict(showgrid=False, zeroline=False, showticklabels=False)),
)
fig.update_layout(width=1000, height=600, plot_bgcolor='rgb(255, 255, 255)')
fig.show()
# +
import dash
import dash_core_components as dcc
import dash_html_components as html
app = dash.Dash()
app.layout = html.Div([
dcc.Graph(figure=fig)
])
app.run_server(debug=True, use_reloader=False)
| network_app.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Ji-lASE7BLUo"
#
# # <font color=#770000>ICPE 639 Introduction to Machine Learning </font>
#
# ## ------ With Energy Applications
#
# Some of the examples and exercises of this course are based on several books as well as open-access materials on machine learning, including [Hands-on Machine Learning with Scikit-Learn, Keras and TensorFlow](https://www.oreilly.com/library/view/hands-on-machine-learning/9781492032632/)
#
#
# <p> © 2021: <NAME> </p>
#
# [Homepage](http://xqian37.github.io/)
#
# **<font color=blue>[Note]</font>** This is currently a work in progress, will be updated as the material is tested in the class room.
#
# All material open source under a Creative Commons license and free for use in non-commercial applications.
#
# Source material used under the Creative Commons Attribution-NonCommercial 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc/3.0/ or send a letter to Creative Commons, PO Box 1866, Mountain View, CA 94042, USA.
# + [markdown] id="h8CLzQygBLUt"
# # Perceptrons, Boosting, \& Artificial Neural Networks (ANNs)
#
#
# Here we cover some basics that may serve as the introduction to deep neural networks. This section will cover the content listed below:
#
# - [1 Perceptron](#1-Perceptron)
# - [2 AdaBoost](#2-AdaBoost)
# - [3 ANNs](#3-ANNs)
# - [4 Hands-on Exercise](#4-Hands-on-Exercise)
# - [Reference](#Reference)
#
#
# + id="P78hJOpbBLUu" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632780853929, "user_tz": 300, "elapsed": 1067, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="1266ad98-b45f-453e-d14c-e22f5750db5c"
# required modules
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from sklearn.ensemble import AdaBoostClassifier
from sklearn.tree import DecisionTreeClassifier, export_graphviz, export_text
from sklearn.externals.six import StringIO
import pydot
from IPython.display import Image
from IPython.core.display import HTML
from sklearn.metrics import accuracy_score
import seaborn as sns
from sklearn import neighbors, datasets
from matplotlib.colors import ListedColormap
from sklearn.model_selection import train_test_split
from sklearn.neighbors import KNeighborsClassifier
# + [markdown] id="CzW3wV_zKTbk"
# ## 1 Perceptron
#
#
#
# + [markdown] executionInfo={"elapsed": 1294, "status": "ok", "timestamp": 1616527812431, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "11420231255337685715"}, "user_tz": 300} id="agkk34mwfSW1" outputId="dfc8c64f-d800-428a-a4aa-6b6de7147a9a"
# ### 1.1 Basics
#
# Perceptrons are simply linear halfspaces for binary classification:
# $$sign(w^Tx),$$
# where $w$ are model parameters (often including the intercept $b$ if needed to make explicit) to consider all features $x$ to predict class label $y\in\{-1, 1\}$.
#
# The intuition is that if all the data points are predicted correctly, then we have $y_n w^Tx_n \geq 0$. Hence, we can just construct the constrained optimization formulation to solve for $w$ under these constraints. Note that if we set the objective function to be $|w|^2$, it is similar as SVM.
# + [markdown] id="hW7sE500KdST"
# #### A simple example with a few lines of Python code
#
# [reference link](https://github.com/MaviccPRP/perceptron/blob/master/perceptron.ipynb)
# + id="zRIFOwDsidt7" colab={"base_uri": "https://localhost:8080/"} executionInfo={"status": "ok", "timestamp": 1632781915625, "user_tz": 300, "elapsed": 194, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="b63c3ea7-697e-4848-c74b-73ada28a0c1e"
X = np.array([
[-2,4,-1],
[4,1,-1],
[1, 6, -1],
[2, 4, -1],
[6, 2, -1],
])
y = np.array([-1,-1,1,1,1])
def perceptron_sgd(X, Y):
w = np.zeros(len(X[0])) # initialize to 0's
eta = 1 # learning rate
epochs = 20 # number of iterations
for t in range(epochs):
for i, x in enumerate(X):
if (np.dot(X[i], w)*Y[i]) <= 0:
w = w + eta*X[i]*Y[i] # update if not correctly classified
return w
w = perceptron_sgd(X,y)
print(w)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="JHerhFCOijnp" executionInfo={"status": "ok", "timestamp": 1632781931279, "user_tz": 300, "elapsed": 376, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="6dec3d5a-8cbf-4c93-829e-49c7e8387d3e"
# visualize
for d, sample in enumerate(X):
# Plot the negative samples
if d < 2:
plt.scatter(sample[0], sample[1], s=120, c="r",marker='_', linewidths=2)
# Plot the positive samples
else:
plt.scatter(sample[0], sample[1], s=120, c="b", marker='+', linewidths=2)
# Print a possible hyperplane, that is seperating the two classes.
plt.plot([-2,6],[6,0.5])
# + colab={"base_uri": "https://localhost:8080/", "height": 297} id="WlKMz3h_KjiN" executionInfo={"status": "ok", "timestamp": 1632672163467, "user_tz": 300, "elapsed": 364, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="a542c510-fe0c-4c92-e2b2-2e97fcd45614"
# visualize the training procedure
def perceptron_sgd_plot(X, Y):
'''
train perceptron and plot the total loss in each epoch.
:param X: data samples
:param Y: data labels
:return: weight vector as a numpy array
'''
w = np.zeros(len(X[0]))
eta = 1
n = 30
errors = []
for t in range(n):
total_error = 0
for i, x in enumerate(X):
if (np.dot(X[i], w)*Y[i]) <= 0:
total_error += (np.dot(X[i], w)*Y[i])
w = w + eta*X[i]*Y[i]
errors.append(total_error*-1)
plt.plot(errors)
plt.xlabel('Epoch')
plt.ylabel('Total Loss')
return w
print(perceptron_sgd_plot(X,y))
# + [markdown] id="mFy14PRRKkhC"
# ### 1.2 Perceptron Algorithm
#
# As shown in the previous example, here is the summary of the algorithm:
#
# 1. Initialize $w$ (often set it to all zero).
# 2. Update: $w \leftarrow w + \eta y_n x_n$ if $x_n$ is predicted wrong, where $\eta$ is the learning rate.
#
# Note that this is an **online** learning algorithm. One important theoretical results is that if the training set is linearly separable, the perceptron algorithm converges in a finite number of iterations.
#
# One possible derivation is to consider the *hinge loss* of misclassification:
# $$ l(y_n, x_n; w) = (- y_n w^T x_n)_+ $$
# As we are minimizing the loss, we would update by **gradient descent**:
# $$ w \leftarrow w - \eta \nabla_w l(y_n, x_n; w), $$
# where if we have straight-through simplification for computing the hinge-loss gradient, we can have the updating rule:
# $$ w \leftarrow w + \eta y_n x_n, \mbox{ if wrong prediction}. $$
#
# **<font color=blue>[Note]</font>** Compare with the updating equation for deriving **logistic regression (LR)** (Note the different outspace: Here, $y\in\{-1, 1\}$; while in LR, $y\in\{0, 1\}$.
# + colab={"base_uri": "https://localhost:8080/"} id="I1iRjLuW8wL3" executionInfo={"status": "ok", "timestamp": 1632782436438, "user_tz": 300, "elapsed": 322, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="a5d0b711-c4bd-4868-dcfd-afb3ab0e28ae"
from sklearn.linear_model import Perceptron
clf = Perceptron(fit_intercept=False, random_state=2)#(tol=1e-5, fit_intercept=False)#, random_state=0)
# Not sure why the results change for different random_state as supposed to converge easily in this case...
clf.fit(X, y)
Perceptron()
clf.score(X, y)
print(clf.coef_)
print(X)
print(clf.predict(X))
print(y)
# + [markdown] id="3e5tNaqGKq3L"
#
# ### 1.3 An Application
# + [markdown] id="zDZygSpqHqsh"
# #### Background
# * `Heart` dataset contains a binary outcome `HD` for 303 patients who presented with chest pain. An outcome value of `Yes` indicates the presence of hear disease based on an angiographic test, while `No` means no heart disease.
#
# * There are 13 predictors including `Age`, `Sex`, `Chol` (a cholesterol measurement), `Thal`( Thallium stress test) and other heart and lung function measurements.
#
# + colab={"base_uri": "https://localhost:8080/"} id="yZ_XqXdcKvz7" executionInfo={"status": "ok", "timestamp": 1632783408065, "user_tz": 300, "elapsed": 694, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="872db53c-2b77-4599-af2a-1247545eb892"
Heart = pd.read_csv('https://raw.githubusercontent.com/XiaomengYan/MachineLearning_dataset/main/Heart.csv').drop('Unnamed: 0', axis=1).dropna()
Heart.info()
# + id="O8Yq_fFqji8F" executionInfo={"status": "ok", "timestamp": 1632783410909, "user_tz": 300, "elapsed": 341, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}}
Heart.ChestPain = pd.factorize(Heart.ChestPain)[0]
Heart.Thal = pd.factorize(Heart.Thal)[0]
X2 = Heart.drop('AHD', axis=1) # explanatory variables
y2 = pd.factorize(Heart.AHD)[0] # response variables AHD
# + colab={"base_uri": "https://localhost:8080/"} id="IbFFfNSjjlNk" executionInfo={"status": "ok", "timestamp": 1632783413889, "user_tz": 300, "elapsed": 551, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="44999a62-c938-48f6-926b-c240893764fe"
clf = Perceptron(fit_intercept=False, random_state=2)
clf.fit(X2,y2)
clf.score(X2,y2)
# + [markdown] id="J4FBmcX0I-Wo"
# ## 2 AdaBoost
#
# **Reminder** Ensemble learning: Ensemble learning methods are meta-algorithms that combine several *weak* learners (homogenous or heterogeneous) into a single predictive model to improve prediction performance. Ensemble methods can decrease variance using bagging, reduce bias by boosting, or improve predictions using stacking.
#
#
#
# **<font color=blue>[Note]</font>** **Boosting**, **bagging**, and **stacking**
#
# * *Boosting*: an ensemble method for improving the model predictions. The idea is to train weak learners sequentially, each trying to correct its predecessor. When an input is misclassified, its contribution to the training loss is increased so that it's more likely to be classified correctly. Models in boosting are related to the previous derived models.
# * *Bagging*: (Bootstrap Aggregating) a machine learning ensemble strategy to improve performance, which uses bootstrap to get samples from the original training data, builds the models on each sampled dataset, and aggregates the results of all the models. Bagging can be parallelized as models are trained by different bootstrapped samples.
# * *Stacking*: a strategy that combines multiple base models' predictions into a new data set, which serves as the input data for another model to predict. (similar as deep network architectures)
#
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 432} id="RmDwYGCtNhDJ" executionInfo={"status": "ok", "timestamp": 1631919776787, "user_tz": 300, "elapsed": 11, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09056639542636885950"}} outputId="41078b12-bb7c-4dc4-867f-7db8d66fa9ad"
Image(url= "https://res.cloudinary.com/dyd911kmh/image/upload/f_auto,q_auto:best/v1542651255/image_2_pu8tu6.png")
# + [markdown] id="NnDbAfJOkjsk"
#
#
# ### 2.1 Algorithm
#
#
# Take classification to illustrate the algorithmic procedure for AdaBoost (Adaptive Boosting).
#
# 1. Initially, AdaBoost selects a training subset randomly.
# 2. It iteratively trains the weak learner $h_t$ by selecting the training set based on the accurate prediction of the last training.
# 3. It assigns the weight to the trained classifier in each iteration according to the accuracy of the classifier. The more accurate classifier will get high weight:
# $$ \alpha_t = \frac{1}{2}\ln{(\frac{1}{\epsilon_t}-1)},$$
# where $\epsilon_t = \sum_n \rho_n^t \mathbb{1}(y_n \neq h_t(x_n))$ denotes the weighted training classification error.
# 4. It assigns the higher weight to wrong classified observations so that in the next iteration these observations will get the high probability for classification:
# $$ \rho^{t+1}_n = \frac{\rho^{t}_n e^{-y_n \alpha_t h_t(x_n)}}{\sum_i \rho^{t}_i e^{-y_i \alpha_t h_t(x_i)}}. $$
# 5. This process iterate until the complete training data fits without any error or until reached to the specified maximum number of estimators.
# 6. Output the final prediction $f(x_n) = \sum_t \alpha_t h_t(x_n)$.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 321} id="lpXd-ix3Od3j" executionInfo={"status": "ok", "timestamp": 1632672190070, "user_tz": 300, "elapsed": 92, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="2794cb04-bbdb-4090-9788-64f54d2f96cd"
Image(url= "https://res.cloudinary.com/dyd911kmh/image/upload/f_auto,q_auto:best/v1542651255/image_3_nwa5zf.png")
# + [markdown] id="6UMo8YxQg3t5"
# ### 2.2 AdaBoost with the `Heart` dataset
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="KyMbSRqZMR7n" executionInfo={"status": "ok", "timestamp": 1632783423829, "user_tz": 300, "elapsed": 231, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="b89e7c8d-1a08-48e0-c9c5-ad6326d626a9"
clf = AdaBoostClassifier(n_estimators=50, learning_rate=1)
clf.fit(X2,y2)
# + colab={"base_uri": "https://localhost:8080/"} id="hA4Wg9qWMYfJ" executionInfo={"status": "ok", "timestamp": 1632672195889, "user_tz": 300, "elapsed": 96, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="2415fe55-8806-494f-98aa-9aa5e5edb1b8"
clf.score(X2,y2) #Return the mean accuracy on the given data and labels.
# + colab={"base_uri": "https://localhost:8080/"} id="vFU4hTnwM0GV" executionInfo={"status": "ok", "timestamp": 1632783481799, "user_tz": 300, "elapsed": 292, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="0a2bd5b4-0db5-46ef-c88e-89637c1ecdd8"
# Create adaboost classifer object with Perceptron
abc = AdaBoostClassifier(n_estimators=50, algorithm='SAMME', base_estimator=Perceptron(), learning_rate=1) #algorithm='SAMME.R',
abc.fit(X2,y2)
abc.score(X2,y2)
# + colab={"base_uri": "https://localhost:8080/"} id="5DR1xoYoPTIh" executionInfo={"status": "ok", "timestamp": 1632783920453, "user_tz": 300, "elapsed": 384, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="5a2faae8-e352-40c1-eea9-6d3f6092b519"
from sklearn import datasets
# Load data
iris = datasets.load_iris()
X = iris.data
y = iris.target
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3) # 70% training and 30% test
# Create adaboost classifer object
abc = AdaBoostClassifier(n_estimators=50, learning_rate=1)
# Train Adaboost Classifer
model = abc.fit(X_train, y_train)
#Predict the response for test dataset
y_pred = model.predict(X_test)
# Model Accuracy, how often is the classifier correct?
print("Accuracy:", accuracy_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="JZt9oR7PlBHl" executionInfo={"status": "ok", "timestamp": 1632783925219, "user_tz": 300, "elapsed": 673, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="a08fb9d8-b529-4a64-edc6-a3b9f26fafea"
# Create adaboost classifer object
abc = AdaBoostClassifier(n_estimators=50, algorithm='SAMME', base_estimator=Perceptron(), learning_rate=1) #algorithm='SAMME.R',
# Train Adaboost Classifer
model = abc.fit(X_train, y_train)
#Predict the response for test dataset
y_pred = model.predict(X_test)
print(y_pred)
# Model Accuracy, how often is the classifier correct?
print("Accuracy:", accuracy_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="hIhxRmqVn-Gp" executionInfo={"status": "ok", "timestamp": 1632783937259, "user_tz": 300, "elapsed": 154, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="906f0bb1-a5ec-4385-a367-0ae7fc423f3b"
# Create Perceptron classifer object
clf = Perceptron(tol=1e-3, random_state=0)
# Train Perceptron Classifer
model = clf.fit(X_train, y_train)
#Predict the response for test dataset
y_pred = model.predict(X_test)
print(y_pred)
# Model Accuracy, how often is the classifier correct?
print("Accuracy:", accuracy_score(y_test, y_pred))
# + id="AVCKuLtcgewd" executionInfo={"status": "ok", "timestamp": 1632783963589, "user_tz": 300, "elapsed": 538, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}}
# Perceptron??
# + [markdown] id="luWvkSDo-HxG"
# **SAMME** (Stagewise Additive Modeling using a Multi-class Exponential loss function): https://web.stanford.edu/~hastie/Papers/samme.pdf
#
# http://article.sapub.org/10.5923.j.ajis.20130302.02.html
# + [markdown] id="zXYjeWDeyZcv"
#
# ### 2.3 Math derivations
#
# We will just make hand-waving explanation here: If we have seens that AdaBoost assumes the prediction model:
# $$f(x) = \sum_t \alpha_t h_t(x),$$
# and for each iteration, the model updates:
# $$f_{t}(x) = f_{t-1}(x) + \alpha_t h_t(x), $$
# the model training is to minimize the training loss with the exponential loss function (upperbounding the zero-one loss):
# $$ \min_{\alpha, h}\sum_n e^{-y_n f_t(x_n)}. $$
#
# The choice of the expential loss also provides the theoretical proof that AdaBoost will achieve infinitely small empricial risk asymptotically. Another interesting theoretical result is that AdaBoost somehow is immune to overfitting.
# + [markdown] id="hquPBahNBLU3"
# ## 3 ANNs
#
#
# + [markdown] id="w-SDYByDBLU3"
# ### 3.1 Basics
#
# We have introduced the perceptron algorithm, which is the basic unit for most of the artificial neural networks, including many popular deep network models. Here we focus on the **multilayer perceptron (MLP)**. MLP is a feed-forward ANN model that maps sets of input data onto a set of appropriate outputs. MLP often consists of multiple layers and each layer is fully connected to the following one. The nodes of the layers are neurons using nonlinear activation functions, except for the nodes of the input layer. There can be one or more non-linear hidden layers between the input and the output layer. The training is through backpropagation (chain rule to compute gradients for involved network parameters) based on the corresponding loss function(s).
#
# MLP can be mathematically represented as:
# $$ h^l = g (W^l h^{l-1}), $$
# where $W^l$ denotes the weighting parameters at the $l$th layer and $g(\cdot)$ is the activation function. The hidden variables $h^l$ can be considered as the output for each hidden layer, taking the previous layer's output $h^{l-1}$ as the input. $h^0 = X$ as the input feature vector and the output of the output layer is often used to predict the outcome of interest:
# $$\sum_i W^{L}_i h^{L-1}_i \rightarrow \hat{y}$$ for regression or
# $$\sigma(\sum_i W^{L}_i h^{L-1}_i) \rightarrow \hat{y}$$ for classification.
#
# **<font color=blue>[Note]</font>** All the deep network architectures, convolutional NNs (CNNs), recurrent NNs (RNNs), and graph NNs (GNNs) can all be considered as more "structured" neural networks, customized to the input feature relationships.
# + [markdown] id="Al_jGUMAs9mL"
# ### 3.2 Backpropagation
#
# Essentially, training neural networks is mostly based on (stochastic) gradient descent algorithms. Recently, there have also been high-order methods.
#
# Similar as training perceptrons, the solution of the network parameters $W^l$'s can be solved by (error) **backpropagation** with gradients of the corresponding loss functions:
# $$\mathcal{L}(\hat{y}, y; X, W^l),$$
# where $\nabla_{W^l} \mathcal{L}$ is always computed from the last layer to the input layer through the chain rule with the hidden variables $h^l$. Note that the updating $W^L$ is easy (We have covered it in linear regression or logistic regression). $\nabla_{h^{L-1}} \mathcal{L}$ can also be computed easily.
#
# Let's first denote $z^l = W^{l}h^{l-1}$ and $h^{l} = g(z^l)$ as defined above in MLP, we can write $\nabla_{W^l}\mathcal{L} = \nabla_{z^l}\mathcal{L} \nabla_{W^l} z^l = \nabla_{z^l}\mathcal{L} \bigodot h^{l-1}$ by the chain rule. Note that $\nabla_{z^l}\mathcal{L}$ is easy to compute by backpropagation:
# $$\nabla_{z^l}\mathcal{L} = \nabla_{h^{l}}\mathcal{L}\nabla_{z^l} h^{l} = \nabla_{h^{l}}\mathcal{L} \bigodot g'(z^{l}),$$
# where $g'(\cdot)$ is the derivative of the activation function. Note that
# $$\nabla_{h^{l}}\mathcal{L} = \nabla_{z^{l+1}}\mathcal{L} \nabla_{h^{l}} z^{l+1} = \nabla_{z^{l+1}}\mathcal{L} W^{l+1}$$
# and $W^{l+1}$ has been updated at the layer $l+1$ and $\nabla_{z^{l+1}}\mathcal{L}$ can be again backpropagated from the layer $l+2$. Therefore,
# $$\nabla_{W^l }\mathcal{L} = \nabla_{z^{l+1}}\mathcal{L} W^{l+1} \bigodot g'(z^{l})\bigodot h^{l-1}.$$
#
# + colab={"base_uri": "https://localhost:8080/", "height": 329} id="tJ9fYUIWWYBy" executionInfo={"status": "ok", "timestamp": 1631919777228, "user_tz": 300, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09056639542636885950"}} outputId="567d8752-d12a-4db7-fef2-5669e8875aef"
Image(url= "https://www.python-course.eu/images/mlp_example_layer_800w.webp", width=500)
# + [markdown] id="qp6Y7x7QWUbn"
# ### 3.3 A naive example
# + colab={"base_uri": "https://localhost:8080/", "height": 265} id="lPZm0GaBBLU4" executionInfo={"status": "ok", "timestamp": 1632785441549, "user_tz": 300, "elapsed": 482, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="3ed34aff-8dc3-4f6f-eeff-cddedfc3e58c"
# simulated data in two-dimensional space
from sklearn.datasets import make_blobs
n_samples = 200
blob_centers = ([1, 1], [3, 4], [1, 3.3], [3.5, 1.8])
data, labels = make_blobs(n_samples=n_samples,
centers=blob_centers,
cluster_std=0.5,
random_state=0)
colours = ('green', 'orange', "blue", "magenta")
fig, ax = plt.subplots()
for n_class in range(len(blob_centers)):
ax.scatter(data[labels==n_class][:, 0],
data[labels==n_class][:, 1],
c=colours[n_class],
s=30,
label=str(n_class))
datasets = train_test_split(data,
labels,
test_size=0.2)
train_data, test_data, train_labels, test_labels = datasets
# + colab={"base_uri": "https://localhost:8080/"} id="f6Rxvq_wVAq6" executionInfo={"status": "ok", "timestamp": 1632785495666, "user_tz": 300, "elapsed": 494, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="ac804459-f931-4a11-b7d3-06aa4adb51d9"
from sklearn.neural_network import MLPClassifier
clf = MLPClassifier(solver='lbfgs',
alpha=1e-5,
hidden_layer_sizes=(6,),
random_state=1)
clf.fit(train_data, train_labels)
clf.score(train_data, train_labels)
# + colab={"base_uri": "https://localhost:8080/"} id="Q9PaIbUSVULx" executionInfo={"status": "ok", "timestamp": 1632785537131, "user_tz": 300, "elapsed": 168, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="cf4e60ec-354f-4548-fc62-0a3b07b0a30b"
predictions_train = clf.predict(train_data)
predictions_test = clf.predict(test_data)
train_score = accuracy_score(predictions_train, train_labels)
print("score on train data: ", train_score)
test_score = accuracy_score(predictions_test, test_labels)
print("score on train data: ", test_score)
predictions_train[:20]
# + colab={"base_uri": "https://localhost:8080/"} id="lmxlsU8UVf7E" executionInfo={"status": "ok", "timestamp": 1632672234724, "user_tz": 300, "elapsed": 84, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="0eb09853-465a-4cfb-970f-c2fb7006614b"
## A really naive example
X = [[0., 0.], [0., 1.], [1., 0.], [1., 1.]]
y = [0, 0, 0, 1]
clf = MLPClassifier(solver='lbfgs', alpha=1e-5,
hidden_layer_sizes=(5, 2), random_state=1)
print(clf.fit(X, y))
# + colab={"base_uri": "https://localhost:8080/"} id="IBm3Ejv_Vitv" executionInfo={"status": "ok", "timestamp": 1632672238415, "user_tz": 300, "elapsed": 103, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="fd4c55f2-9d57-4129-8352-81d5afdf067c"
print("weights between input and first hidden layer:")
print(clf.coefs_[0])
print("\nweights between first hidden and second hidden layer:")
print(clf.coefs_[1])
# + [markdown] id="9eiGT5R4BLU4"
# ## 4 Hands-on exercise: Iris data
# + colab={"base_uri": "https://localhost:8080/", "height": 205} id="jeJdQzy6lm87" executionInfo={"status": "ok", "timestamp": 1632672243768, "user_tz": 300, "elapsed": 174, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="969cea54-1263-4671-9e1e-2f9fd46ebd01"
from sklearn import datasets
iris = datasets.load_iris()
iris_data = pd.DataFrame({
'sepal length':iris.data[:,0],
'sepal width':iris.data[:,1],
'petal length':iris.data[:,2],
'petal width':iris.data[:,3],
'species':iris.target
})
iris_data.head()
# + colab={"base_uri": "https://localhost:8080/"} id="TXHLWciBBLU5" executionInfo={"status": "ok", "timestamp": 1632672246309, "user_tz": 300, "elapsed": 86, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="18237e61-1387-49e2-baec-e5d2edfd7c61"
iris = datasets.load_iris()
X = iris.data[:, 2:4]
y = iris.target
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3) # 70% training and 30% test
cmap_light = ListedColormap(['orange', 'azure', 'green'])
cmap_bold = ['darkorange', 'darkblue', 'darkgreen']
weights='uniform'
# we create an instance of Perceptron to fit the data:
# Create Perceptron classifer object
clf = Perceptron(tol=1e-8, random_state=0)
# Train Perceptron Classifer
model = clf.fit(X_train, y_train)
clf.score(X_test,y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="RQmxxV9TOqQS" executionInfo={"status": "ok", "timestamp": 1632672250811, "user_tz": 300, "elapsed": 201, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="9f5ddce0-3b9d-468a-fc8e-564162248fb6"
# Create adaboost classifer object
abp = AdaBoostClassifier(n_estimators=50, algorithm='SAMME', base_estimator=Perceptron(), learning_rate=1) #algorithm='SAMME.R',
# Train Adaboost Classifer
model = abp.fit(X_train, y_train)
abp.score(X_test,y_test)
# + colab={"base_uri": "https://localhost:8080/"} id="cK6fDlEwPcBh" executionInfo={"status": "ok", "timestamp": 1632672256704, "user_tz": 300, "elapsed": 215, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="7d182651-9cd5-4aaa-dd8f-f73a962a3906"
# Create adaboost classifer object
abc = AdaBoostClassifier(n_estimators=50, learning_rate=1)
# Train Adaboost Classifer
model = abc.fit(X_train, y_train)
#Predict the response for test dataset
y_pred = model.predict(X_test)
# Model Accuracy, how often is the classifier correct?
print("Accuracy:", accuracy_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/"} id="LN7dk_3LQJc1" executionInfo={"status": "ok", "timestamp": 1632672262297, "user_tz": 300, "elapsed": 99, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="7bfec726-0284-415e-db95-5f98583a357e"
# Create MLP
mlp = MLPClassifier(solver='lbfgs', alpha=1e-5,
hidden_layer_sizes=(15, 12), random_state=1)
model = mlp.fit(X_train, y_train)
y_pred = model.predict(X_test)
print("Accuracy:", accuracy_score(y_test, y_pred))
# + colab={"base_uri": "https://localhost:8080/", "height": 404} id="bayzbi7RN7Jq" executionInfo={"status": "ok", "timestamp": 1632672268119, "user_tz": 300, "elapsed": 555, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "06431978792501680815"}} outputId="4c82c660-5ccd-4a01-d9d0-9b6766892854"
# Plot the decision boundary. For that, we will assign a color to each
# point in the mesh [x_min - 0.5, x_max + 0.5]x[y_min - 0.5, y_max + 0.5].
h = 0.01
xx, yy = np.meshgrid(np.arange(X[:, 0].min() - 0.5, X[:, 0].max() + 0.5, h), np.arange(X[:, 1].min() - 0.5, X[:, 1].max() + 0.5, h))
Z = mlp.predict(np.c_[xx.ravel(), yy.ravel()])
# Put the result into a color plot
Z = Z.reshape(xx.shape)
plt.figure(figsize=(8, 6))
plt.contourf(xx, yy, Z, cmap = cmap_light)
# Plot also the training points
#n_neighbors = 8
sns.scatterplot(x = X[:, 0], y = X[:, 1], hue = iris.target_names[y], palette = cmap_bold, alpha = 0.8, edgecolor = "black")
plt.xlim(xx.min(), xx.max())
plt.ylim(yy.min(), yy.max())
plt.title("3-Class classification")
plt.xlabel(iris.feature_names[2])
plt.ylabel(iris.feature_names[3])
plt.show()
# + [markdown] id="qwQ8Yb-0h__O"
# ## Reference
# * [Perceptron](https://en.wikipedia.org/wiki/Perceptron)
# * [A quick guide to boosting in Machine Learning](https://medium.com/greyatom/a-quick-guide-to-boosting-in-ml-acf7c1585cb5)
# * [AdaBoost from DataCamp](https://www.datacamp.com/community/tutorials/adaboost-classifier-python)
# * [A Primer to Ensemble Learning – Bagging and Boosting](https://analyticsindiamag.com/primer-ensemble-learning-bagging-boosting/)
# * [ANNs with scikitlearn](https://www.python-course.eu/neural_networks_with_scikit.php)
# + [markdown] id="Jz7RJ9HgARGF"
# # # Questions?
# + colab={"base_uri": "https://localhost:8080/", "height": 56} id="O1Y6HcZPBLU6" executionInfo={"status": "ok", "timestamp": 1631919778483, "user_tz": 300, "elapsed": 10, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s64", "userId": "09056639542636885950"}} outputId="540ad1a3-2458-4d10-c268-db9a76f7750a"
Image(url= "https://mirrors.creativecommons.org/presskit/buttons/88x31/png/by-nc-sa.png", width=100)
# + id="euTnNjqSBLU6"
| Mod4-1-ML-SL-PerceptronAdaBoostANN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Решающие деревья
# При подготовке использовались <a href="https://github.com/esokolov/ml-course-hse/blob/master/2017-fall/seminars/sem07-trees.ipynb">материалы</a> <NAME>
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from sklearn.tree import DecisionTreeRegressor
# %matplotlib inline
# -
# # Постановка задачи и примеры
#
# ### Дерево для задачи классификации:
# <img src='img/0_tree.png' Width=900>
# ### Дерево для задачи регрессии:
# Сгенерируем датасет
rng = np.random.RandomState(1)
X = np.sort(5 * rng.rand(80, 1), axis=0)
y = np.sin(X).ravel()
y[::5] += 3 * (0.5 - rng.rand(16))
# Обучим модели
regr_1 = DecisionTreeRegressor(max_depth=2)
regr_2 = DecisionTreeRegressor(max_depth=5)
regr_1.fit(X, y)
regr_2.fit(X, y)
# Предскажем обученными регрессорами
X_test = np.arange(0.0, 5.0, 0.01)[:, np.newaxis]
y_1 = regr_1.predict(X_test)
y_2 = regr_2.predict(X_test)
# Построим график
# +
plt.figure(figsize=(16, 10))
plt.scatter(X, y, s=50, color="black", label="data")
plt.plot(X_test, y_1, color="green", label="max_depth=2", linewidth=3)
plt.plot(X_test, y_2, color="red", label="max_depth=5", linewidth=3)
plt.xlabel("data")
plt.ylabel("target")
plt.title("Decision Tree Regression")
plt.legend()
plt.show()
# -
# ## Красивая визуализация
# http://www.r2d3.us/visual-intro-to-machine-learning-part-1/
# # Построение деревьев
#
# * Обучающая выборка $(x_i,y_i)_{i=1}^l\in X \times Y$
# * Как разбить на две части: $R_1(j,s)=\{x|x_j \leq s\}$ и $R_2(j,s) = \{x | x_j > s \}$ с использованием критерия Q(X, j, s)?
#
# Найдём наилучшие значения $j$ и $s$, создадим корневую вершину дерева, поставив ей в соответствие функцию (предикат) $[x_j \leq s ]$. Объекты выборки будут разбиты на две части и попадут либо в левое, либо в правое поддерево. Продолжим эту процедуру для каждой подвыборки. Если после очередного разбиения в одной из половин окажутся объекты одного из классов, то создадим листовую вершину, которой будет соответствовать класс попавших в неё объектов.
# Жадный алгоритм переусложняет структуру дерева:
#
#
# <img src='img/0_greedy_tree.png' Width=900>
# # Критерии информативности
#
# * $R_m$ - множество объектов обучающей выборки, попавших в вершину $m$,
# * $N_m=|R_m|$.
# * $p_{mk}$ - доля объектов класса $k\in\{1, ..., K\}$, попавших в вершину $m$: $p_{mk}=\frac{1}{N_m} \sum\limits_{x_i\in R_m} [y_i = k]$.
# * $k_m = arg \max\limits_{k} p_{mk}$ - класс, чьих представителей больше всего среди объектов, попавших в вершину $m$.
#
# ## 1. Ошибка классификации
#
# Если бы вершина $m$ была листовой и относила все объекты к классу $k$:
#
# $$
# H(R_m) = \frac{1}{N_m} \sum\limits_{x_i \in R_m} [y_i \neq k_m].
# $$
#
# Критерий информативности при ветвлении вершины $m$: ($l$ и $r$ - правые и левые вершины)
#
# $$
# Q(R_m, j, s) = H (R_m) - \frac{N_l}{N_m} H(R_l) - \frac{N_r}{N_m} H(R_r) \to \max\limits_{j, s}
# $$
# Грубый критерий - учитывает частоту $p_{m, k_m}$ лишь одного класса
# #### Задача 1
# Покажите, что ошибку классификации также можно записать в виде
# $$H(R_m) = 1 - p_{m, k_m}$$
# #### Решение
# $$
# 1 = \frac{1}{N_m}\sum_{(x_i,\,y_i) \in R_m}[y_i \neq k_m] + \frac{1}{N_m}\sum_{(x_i,\,y_i) \in R_m}[y_i = k_m]
# $$
# $$
# H(R_m) = \frac{1}{N_m}\sum_{(x_i,\,y_i) \in R_m}[y_i \neq k_m] = 1 - p_{m, k_m}
# $$
# ## 2. Индекс Джини
# * Функционал имеет вид $$ H(R_m) = \sum\limits_{k \neq k'}p_{mk}p_{mk'}$$
# * Аналогично определяется критерий информативности:
# $$
# Q(R_m, j, s) = H(R_m) - \frac{N_l}{N_m} H(R_l) - \frac{N_r}{N_m} H(R_r).
# $$
# #### Задача 2
#
# Покажите, что индекс Джини $H(R_m)$ также можно записать в виде:
#
# $$H(R_m) = \sum_{k = 1}^{K} p_{mk} (1 - p_{mk}) = 1 - \sum_{k = 1}^K p_{mk}^2$$
# #### Решение
#
# $$
# \sum_{k \neq k'} p_{mk} p_{mk'}
# =
# \sum_{k = 1}^{K} p_{mk} \sum_{k' \neq k} p_{mk'}
# =
# \sum_{k = 1}^{K} p_{mk} (1 - p_{mk}).
# $$
# #### Задача 3
#
# Рассмотрим вершину $m$ и объекты $R_m$, попавшие в нее. Сопоставим в соответствие вершине $m$ алгоритм $a(x)$, который выбирает класс случайно, причем класс $k$ выбирается с вероятностью $p_{mk}$. Покажите, что матожидание частоты ошибок этого алгоритма на объектах из $R_m$ равно индексу Джини.
# #### Решение
#
# \begin{multline*}
# \mathbb E\frac{1}{N_m} \sum_{x_i \in R_m} [y_i \neq a(x_i)]
# =
# \frac{1}{N_m} \sum_{(x_i,\,y_i) \in R_m} \mathbb E[y_i \neq a(x_i)]
# =
# \frac{1}{N_m} \sum_{(x_i,\,y_i) \in R_m} (1 - p_{m,y_i})
# =\\
# =
# \sum_{k = 1}^{K} \frac{\sum_{(x_i,\,y_i) \in R_m} [y_i = k]}{N_m} (1 - p_{mk})
# =
# \sum_{k = 1}^{K} p_{mk} (1 - p_{mk}).
# \end{multline*}
# Выясним теперь, какой смысл имеет максимизация функционала, соответствующего критерию информативности Джини.
# Сразу выбросим из функционала $H(R_m)$, поскольку данная величина не зависит от $j$ и $s$.
# Преобразуем критерий:
#
# \begin{align*}
# &- \frac{N_\ell}{N_m} H(R_\ell) - \frac{N_r}{N_m} H(R_r)=- \frac{1}{N_m} \left(
# N_\ell - \sum_{k = 1}^{K} p_{\ell k}^2 N_\ell + N_r - \sum_{k = 1}^{K} p_{r k}^2 N_r \right)=\\
# &=
# \frac{1}{N_m} \left(\sum_{k = 1}^{K} p_{\ell k}^2 N_\ell +\sum_{k = 1}^{K} p_{r k}^2 N_r - N_m
# \right)= \{\text{$N_m$ не зависит от $j$ и $s$}\} = \\
# &=\sum_{k = 1}^{K} p_{\ell k}^2 N_\ell + \sum_{k = 1}^{K} p_{r k}^2 N_r.
# \end{align*}
#
# Запишем теперь в наших обозначениях число таких пар объектов $(x_i, x_j)$,
# что оба объекта попадают в одно и то же поддерево, и при этом $y_i = y_j$.
# Число объектов класса $k$, попавших в поддерево $\ell$,
# равно $p_{\ell k} N_\ell$;
# соответственно, число пар объектов с одинаковыми метками, попавших в левое
# поддерево, равно $\sum_{k = 1}^{K} p_{\ell k}^2 N_\ell^2$.
# Интересующая нас величина равна
# $$
# \sum_{k = 1}^{K} p_{\ell k}^2 N_\ell^2 + \sum_{k = 1}^{K} p_{r k}^2 N_r^2.
# $$
# Заметим, что данная величина очень похожа на полученное
# выше представление для критерия Джини.
# Таким образом, максимизацию функционала Джини можно <i>условно</i>
# интерпретировать как максимизацию числа пар объектов одного класса,
# оказавшихся в одном поддереве.
# ## 3. Энтропийный критерий (критерий Шеннона)
# Рассмотрим дискретную случайную величину,
# принимающую $K$ значений с вероятностями $p_1, \dots, p_K$
# соответственно.
# *** Энтропия *** этой случайной величины определяется как:
# $$H(p) = -\sum_{k = 1}^{K} p_k \log_2 p_k$$
# #### Задача 4
# Покажите, что энтропия ограничена сверху и достигает своего максимума на
# равномерном распределении $p_1 = \dots = p_K = 1/K$.
# #### Решение
#
# Нам понадобится неравенство Йенсена: для любой вогнутой функции $f$
# выполнено
# $$
# f\left(\sum_{i = 1}^{n} a_i x_i\right) \geq \sum_{i = 1}^{n} a_i f(x_i),
# $$
# если $\sum_{i = 1}^{n} a_i = 1$.
#
# Применим его к логарифму в определении энтропии~(он является вогнутой функцией):
# $$
# H(p) = \sum_{k = 1}^{K} p_k \log_2 \frac{1}{p_k}
# \leq \log_2 \left( \sum_{k = 1}^{K} p_i \frac{1}{p_i} \right)=\log_2 K.
# $$
#
# Наконец, найдем энтропию равномерного распределения:
# $$
# -\sum_{k = 1}^{K} \frac{1}{K} \log_2 \frac{1}{K} = - K \frac{1}{K} \log_2 \frac{1}{K} = \log_2 K.
# $$
# Энтропия ограничена снизу нулем, причем минимум достигается на вырожденных
# распределениях ($p_i = 1$, $p_j = 0$ для $i \neq j$).
#
# Энтропийный критерий информативности определяется как
# $$
# Q(R_m, j, s) = H(p_m) - \frac{N_\ell}{N_m} H(p_\ell) - \frac{N_r}{N_m} H(p_r),
# $$
# где $p_i = (p_{i1}, \dots, p_{iK})$ - распределение классов в $i$-й вершине.
# Видно, что данный критерий отдает предпочтение более "вырожденным" распределениям
# классов.
# +
plt.figure(figsize=(12, 8))
p = np.linspace(0, 1, 100)
plt.plot(p, [2 * x * (1-x) for x in p], label='gini')
plt.plot(p, [4 * x * (1-x) for x in p], label='2*gini')
plt.plot(p, [1 - max(x, 1 - x) for x in p], label='missclass')
plt.plot(p, [2 * (1 - max(x, 1 - x)) for x in p], label='2*missclass')
plt.plot(p, [-x * np.log2(x + 1e-10) - (1 - x) * np.log2(1 - x + 1e-10) for x in p], label='entropy')
plt.xlabel('p+')
plt.ylabel('criterion')
plt.title('Критерии качества как функции от p+ (бинарная классификация)')
plt.legend()
plt.show()
# -
# # Пример: предсказание цвета шарика по его координате
# <img src='img/0_entropy_statement.png' Width=1200>
# * Вероятности вытаскивания синего и жёлтого шариков соответственно: $$ p_1 = \frac{9}{20}, p_2 = \frac{11}{20}$$
# * Энтропия такого состояния: $$ S_0 = -\frac{9}{20} log_2 \frac{9}{20} - \frac{11}{20} log_2 \frac{11}{20} \approx 1$$
# Как изменится энтропия, если разбить шарики на две группы?
# <img src='img/0_entropy_first_split.png' Width=1200>
# * Для первой группы: $$ S_1 = -\frac{8}{13} log_2 \frac{8}{13} - \frac{5}{13} log_2 \frac{5}{13} \approx 0,96 $$
#
# * И для второй: $$ S_2 = -\frac{6}{7} log_2 \frac{6}{7} - \frac{1}{7} log_2 \frac{1}{7} \approx 0,6 $$
# * Энтропия уменьшилась в обеих группах.
# * Мера прироста информации:
# $$IG(Q) = S_0 - \sum\limits_{i=1}^q \frac{N_i}{N} S_i $$
# где q - число групп после разбиения, $N_i$ - число элементов выборки, у которых признак $Q$ имеет $i$-е значение.
# * $$IG(x\leq 12) = S_0 - \frac{13}{20}S_1 - \frac{7}{20}S_2 \approx 0.16$$
# <img src='img/0_entropy_split.png' Width=800>
# * Для правой группы потребовалось всего одно дополнительное разбиение по признаку "координата меньше либо равна 18", для левой – еще три. Очевидно, энтропия группы с шариками одного цвета равна 0 ($log_2 1=0$), что соответствует представлению, что группа шариков одного цвета – упорядоченная.
# ## 4. Критерии в задачах регрессии
# В задачах регрессии, как правило, в качестве критерия выбирают дисперсию ответов в листе:
# $$
# H_R(R_m) = \frac{1}{N_m} \sum_{(x_i,\,y_i) \in R_m} \left(y_i-\frac{1}{N_m}\sum_{(x_i,\,y_i) \in R_m} y_j \right)^2.
# $$
# Можно использовать и другие критерии - например, среднее абсолютное отклонение от медианы.
# # Критерий останова построения дерева
#
# Для любой непротиворечивой обучающей выборки можно построить решающее дерево, которое имеет нулевую ошибку на данной выборке. Если мы рассмотрим объекты, как точки в пространстве признаков, то каждую эту точку можно ограничить n-мерным кубиком, который не будет содержать других точек. n-мерный кубик прекрасно можно задать деревом.
# Однако в этом случае имеет место **переобучение**.
#
# В связи с этим встаёт вопрос: в каком случае вершину следует объявить листовой?
# Рассмотрим модельную задачу регрессии. Объектами будут являться точки на плоскости (т.е. каждый объект описывается 2 признаками), целевой переменной — расстояние от объекта до точки (0, 0).
def get_grid(data):
x_min, x_max = data[:, 0].min() - 1, data[:, 0].max() + 1
y_min, y_max = data[:, 1].min() - 1, data[:, 1].max() + 1
return np.meshgrid(
np.arange(x_min, x_max, 0.01),
np.arange(y_min, y_max, 0.01),
)
# Сгенерируем датасет
data_x = np.random.normal(size=(100, 2))
data_y = (data_x[:, 0] ** 2 + data_x[:, 1] ** 2) ** 0.5
# Визуализируем
plt.figure(figsize=(8, 8))
plt.scatter(data_x[:, 0], data_x[:, 1], c=data_y, s=100, cmap='spring')
# Обучим регрессор
clf = DecisionTreeRegressor()
clf.fit(data_x, data_y)
# Посмотрим как выглядят предсказания
# +
xx, yy = get_grid(data_x)
predicted = clf.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.figure(figsize=(8, 8))
plt.pcolormesh(xx, yy, predicted, cmap='spring')
plt.scatter(data_x[:, 0], data_x[:, 1], c=data_y, s=100, cmap='spring', edgecolor='k')
# -
# Посмотрим как будет выглядить разделение плоскости в зависимости от
# - минимального количества объектов в листе
# - максимальной глубины дерева
# +
plt.figure(figsize=(14, 14))
for i, max_depth in enumerate([2, 4, None]):
for j, min_samples_leaf in enumerate([15, 5, 1]):
clf = DecisionTreeRegressor(max_depth=max_depth, min_samples_leaf=min_samples_leaf)
clf.fit(data_x, data_y)
xx, yy = get_grid(data_x)
predicted = clf.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.subplot2grid((3, 3), (i, j))
plt.pcolormesh(xx, yy, predicted, cmap='spring')
plt.scatter(data_x[:, 0], data_x[:, 1], c=data_y, s=30, cmap='spring', edgecolor='k')
plt.title('max_depth=' + str(max_depth) + ', min_samples_leaf: ' + str(min_samples_leaf))
# -
# - Увеличение максимальной глубины и/или уменьшение минимального количества объектов выборки в листе приводит к увеличению качества на обучающей выборке и переобучению.
# ## Неустойчивость решающих деревьев
#
# Решающие деревья — это алгоритмы, неустойчивые к изменениям обучающей выборки, т.е. при малейших её изменениях итоговый классификатор может радикально измениться.
# Посмотрим, как будет меняться структура дерева при обучении на разных 90%-х подвыборках.
#
# +
plt.figure(figsize=(20, 6))
for i in range(3):
clf = DecisionTreeRegressor(random_state=42)
indecies = np.random.randint(data_x.shape[0], size=int(data_x.shape[0] * 0.9))
clf.fit(data_x[indecies], data_y[indecies])
xx, yy = get_grid(data_x)
predicted = clf.predict(np.c_[xx.ravel(), yy.ravel()]).reshape(xx.shape)
plt.subplot2grid((1, 3), (0, i))
plt.pcolormesh(xx, yy, predicted, cmap='winter')
plt.scatter(data_x[:, 0], data_x[:, 1], c=data_y, s=30, cmap='winter', edgecolor='k')
# -
# ### Категориальные признаки в деревьях
#
# Есть несколько подходов работы с категориальными признаками в деревьях:
#
# * One-hot encoding, Mean target encoding, binary encoding
# * В некоторых фрэймворках (CatBoost) количество исходящих ребер в вершине может быть равно не 2, а количеству категориальных признаком. Таким образом естественно будут учитываться категориальные признаки.
# # Преимущества и Недостатки решающих деревьев:
#
# **Преимущества**
# * хорошо интерпретируются
# * легко обобщаются для регрессии и классификации
# * допускаются разнотипные данные
#
# **Недостатки**
# * Сравнение с линейными алгоритмами на линейно разделимой выборке - фиаско
# * Переобучение
# * Неустойчивость к шуму, составу выборки, критерию
#
# **Способы устранения недостатков**
# * прунинг (усечение)
# * композиции (леса) деревьев
# #### Pruning
#
# Есть разные подходы к прунингу. Самый простой: срезаем листья и делаем родительскую вершину листом с ответом равным самому частому классу, смотрим на изменение качества на выборке, останавливаем, когда оно начинает ухудшаться.
# Или же мы можем находить поддерево, удаление которого не ухудшит нашу ошибку на валидационной выборке, как показано на рисунке:
# <img src='img/pruning.png' Width=800>
| week0_06_trees_and_ensembles/week0_06_decision_trees.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Module 2 Required Coding Activity
# Introduction to Python (Unit 2) Fundamentals
#
# **This Activity is intended to be completed in the jupyter notebook, Required_Code_MOD2_IntroPy.ipynb, and then pasted into the assessment page that follows.**
#
# All course .ipynb Jupyter Notebooks are available from the project files download topic in Module 1, Section 1.
#
# This is an activity based on code similar to the Jupyter Notebook **`Practice_MOD02_IntroPy.ipynb`** which you may have completed.
#
# | Important Assignment Requirements |
# |:-------------------------------|
# | **NOTE:** This program requires creating a function using **`def`** and **`return`**, using **`print`** output, **`input`**, **`if`**, **`in`** keywords, **`.append()`**, **`.pop()`**, **`.remove()`** list methods. As well as other standard Python |
#
# ## Program: list-o-matic
# This program takes string input and checks if that string is in a list of strings
# - if string is in the list it removes the first instance from list
# - if string is not in the list the input gets appended to the list
# - if the string is empty then the last item is popped from the list
# - if the **list becomes empty** the program ends
# - if the user enters "quit" then the program ends
#
# program has 2 parts
# - **program flow** which can be modified to ask for a specific type of item. This is the programmers choice. Add a list of fish, trees, books, movies, songs.... your choice.
# - **list-o-matic** Function which takes arguments of a string and a list. The function modifies the list and returns a message as seen below.
#
# 
#
# **[ ]** initialize a list with several strings at the beginning of the program flow and follow the flow chart and output examples
#
# *example input/output*
# ```
# look at all the animals ['cat', 'goat', 'cat']
# enter the name of an animal: horse
# 1 instance of horse appended to list
#
# look at all the animals ['cat', 'goat', 'cat', 'horse']
# enter the name of an animal: cat
# 1 instance of cat removed from list
#
# look at all the animals ['goat', 'cat', 'horse']
# enter the name of an animal: cat
# 1 instance of cat removed from list
#
# look at all the animals ['goat', 'horse']
# enter the name of an animal: (<-- entered empty string)
# horse popped from list
#
# look at all the animals ['goat']
# enter the name of an animal: (<-- entered empty string)
# goat popped from list
#
# Goodbye!
# ```
#
# *example 2*
# ```
# look at all the animals ['cat', 'goat', 'cat']
# enter the name of an animal: Quit
# Goodbye!
# ```
#
#
# +
# [] create list-o-matic
# [] copy and paste in edX assignment page
# -
# ### Need assignment tips and clarification?
# See the video on the "End of Module coding assignment > Module 2 Required Code Description" course page on [edX](https://courses.edx.org/courses/course-v1:Microsoft+DEV274x+4T2017/course)
#
# # Important: [How to submit code by pasting](https://courses.edx.org/courses/course-v1:Microsoft+DEV274x+2T2017/wiki/Microsoft.DEV274x.2T2017/paste-code-end-module-coding-assignments/)
#
# [Terms of use](http://go.microsoft.com/fwlink/?LinkID=206977) [Privacy & cookies](https://go.microsoft.com/fwlink/?LinkId=521839) © 2017 Microsoft
| Module 2/utf-8''Required_Code_MOD2_IntroPy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Data Processing
# ## Imports
import json
import pandas as pd
# ## Raw Data
# +
def load(tsv_file):
return pd.read_csv(tsv_file, header=0, sep="\t", index_col=False)
amazon = load("../Raw/amazon.tsv")
imdb = load("../Raw/imdb.tsv")
yelp = load("../Raw/yelp.tsv")
# -
# ## Bar Chart Data
# +
def write(json_file):
# It's expected to get 500, 500 for the three data (mentioned in original data source)
sent = {'amazon': {'negative':0, 'positive':0},
'imbd': {'negative':0, 'positive':0},
'yelp': {'negative':0, 'positive':0}}
for idx, row in amazon.iterrows():
if row['sentiment'] == 1:
sent['amazon']['positive'] = sent['amazon']['positive'] + 1
else:
sent['amazon']['negative'] = sent['amazon']['negative'] + 1
for idx, row in imdb.iterrows():
if row['sentiment'] == 1:
sent['imbd']['positive'] = sent['imbd']['positive'] + 1
else:
sent['imbd']['negative'] = sent['imbd']['negative'] + 1
for idx, row in yelp.iterrows():
if row['sentiment'] == 1:
sent['yelp']['positive'] = sent['yelp']['positive'] + 1
else:
sent['yelp']['negative'] = sent['yelp']['negative'] + 1
with open(json_file, 'w') as file:
json.dump(sent, file)
write("bar.json")
# -
| Data/Bar/Bar.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # GKE
#
# ### start the cluster with two nodes
# ```
# gcloud container clusters create cbroker --scopes 'cloud-platform' --num-nodes 2 --enable-basic-auth --issue-client-certificate --enable-ip-alias --zone us-central1-c
# ````
#
# ### install kubectl
# ```
# sudo snap install kubectl --classic
#
# kubectl get nodes
# ```
#
# ### get credentials for console
# ```
# gcloud container clusters get-credentials cbroker --zone us-central1-c --project adtrac-experimental
# ```
#
#
#
# ---
# # Install docker
#
# ### prerequisites
# ```
# sudo apt-get install apt-transport-https ca-certificates curl gnupg-agent software-properties-common
# ````
#
# ### add docker GPG key
# ```
# curl -fsSL https://download.docker.com/linux/ubuntu/gpg | sudo apt-key add -
# ```
#
# ### check fingerprint
# ```
# sudo apt-key fingerprint 0EBFCD88
# ```
#
# ### add repo
# ```
# sudo add-apt-repository "deb [arch=amd64] https://download.docker.com/linux/ubuntu $(lsb_release -cs) stable"
#
# sudo apt-get update
# ```
#
# ### install docker executables
# ```
# sudo apt-get install docker-ce docker-ce-cli containerd.io
# ```
#
#
# ---
#
# # Build
# ### build and run
# ```
# sudo docker build -f Dockerfile.bionic -t gcr.io/adtrac-experimental/cbroker . && sudo docker run -p 8000:8000 cbroker:latest
# ```
#
# ### push to registry
# ```
# sudo gcloud docker -- push gcr.io/adtrac-experimental/cbroker
# ```
# ---
# # Set up prod access
# key file created with IAM/Service Accounts/Create Key
# ```
# kubectl create secret generic cloudsql-oauth-credentials --from-file=cloudsql.json
# ```
#
# ```
# kubectl create secret generic cloudsql --from-literal=username=[PROXY_USERNAME] --from-literal=password=[PASSWORD]
# ```
| mysite/shell_commands.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# **Imports**
import sys
# !{sys.executable} -m pip install --upgrade pip
# !{sys.executable} -m pip install boto3
# **Setup API Keys**
#
# You need 3 environment variables AWS_ACCESS_KEY_ID, AWS_SECRET_ACCESS_KEY and AWS_DEFAULT_REGION,
# if not setup already you can do that using the following inline env setup
# ```
# # %env AWS_DEFAULT_REGION=us-east-2
# # %env AWS_ACCESS_KEY_ID=************
# # %env AWS_SECRET_ACCESS_KEY=**************
# ```
# %env AWS_DEFAULT_REGION=us-east-1
import boto3
import pprint
client = boto3.client('backup')
l = client.list_backup_vaults()
pp = pprint.PrettyPrinter(indent=4)
pp.pprint(l)
| JupyterSamples/hello-world.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import pandas_datareader.data as web
import datetime as dt
start = dt.datetime(2010,1,1)
end = dt.datetime.now()
df = web.DataReader('AAPL', 'yahoo', start, end)
df.head()
df.reset_index(inplace = True)
df.head()
df.columns = [str(col).upper().replace(' ', '_') for col in df.columns]
df.head()
df.shape
df.to_csv('AAPL.csv')
| Getting Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Convolution Neural Network (Inclass - 10/Mar/2018)
import numpy as np
import keras
import tensorflow as tf
import imageio
import matplotlib.pyplot as plt
% matplotlib inline
from keras.models import Sequential
from keras.layers import Dense, Conv2D, Flatten, Dropout, MaxPooling2D, Activation
# ! wget http://bit.do/deepcheetah
# ! mv deepcheetah cheetah.jpg
# ### Convolution Layer
cheetah = imageio.imread("img/cheetah.jpg")
cheetah.shape
plt.imshow(cheetah)
model1 = Sequential()
model1.add(Conv2D(1, (2,2), padding="same",
input_shape = cheetah.shape))
model1.summary()
from helpers import visualise_conv
visualise_conv(cheetah, model1)
# ### With Convultion + Activation
model2 = Sequential()
model2.add(Conv2D(1, (2,2), padding="same",
input_shape = cheetah.shape))
model2.add(Activation("relu"))
model2.summary()
visualise_conv(cheetah, model2)
# ## With Convulution + Activation + Pooling
model3 = Sequential()
model3.add(Conv2D(1, (2,2), padding="same",
input_shape = cheetah.shape))
model3.add(Activation("relu"))
model3.add(MaxPooling2D(pool_size=(16,16)))
visualise_conv(cheetah, model3)
# ## Build a CNN
from keras.datasets import fashion_mnist
from helpers import fashion_mnist_label
(x_train, y_train), (x_test, y_test) = fashion_mnist.load_data()
# ### Steps 1: Preparing the images and labels
from keras import backend as K
K.image_data_format()
x_train_conv = x_train.reshape(x_train.shape[0], 28, 28, 1)
x_test_conv = x_test.reshape(x_test.shape[0], 28, 28, 1)
x_train_conv.shape, x_test_conv.shape
# Normalise
x_train_conv = x_train_conv / 255
x_test_conv = x_test_conv / 255
# convert class vector to binary class matrices
y_train_class = keras.utils.to_categorical(y_train, 10)
y_test_class = keras.utils.to_categorical(y_test, 10)
# ### Step 2: Model - Convolution + Max Pooling + Dropouts
model = Sequential()
model.add(Conv2D(32, (3,3), activation="relu", input_shape= (28,28,1)))
model.add(Conv2D(64, (3,3), activation="relu"))
model.add(MaxPooling2D(pool_size=(2,2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128, activation="relu"))
model.add(Dropout(0.25))
model.add(Dense(10, activation="softmax"))
model.summary()
model.compile(loss="categorical_crossentropy",
optimizer="sgd", metrics=["accuracy"])
# %time
model.fit(x_train_conv, y_train_class, batch_size=128,
epochs=2, verbose=1,
validation_data=(x_test_conv, y_test_class))
| experiments/Version1/InClass-CNN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# MIT License
#
# Copyright (c) 2019 <NAME>, https://orcid.org/0000-0001-9626-8615 (ORCID)
# +
import xarray as xr
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# -
# # Define functions
# +
from scipy.ndimage.filters import gaussian_filter
# band filter
def raster_gamma_range(raster0, g1, g2):
raster = raster0.copy()
raster.values = raster.values.astype(np.float32)
raster.values = gaussian_filter(raster.values,g1) \
- gaussian_filter(raster.values,g2)
return raster
# -
# dem, srtm
def correlogram(raster1, raster2, gammas):
# spatial filtering
rasters1 = []
rasters2 = []
for g in gammas:
print (g,". ", end = '')
_raster1 = raster_gamma_range(raster1, g-.5, g+.5)
rasters1.append(_raster1)
_raster2 = raster_gamma_range(raster2, g-.5, g+.5)
rasters2.append(_raster2)
print ()
corrs = []
for ridx in range(len(gammas)):
print (ridx+1,". ", end = '')
_raster2 = rasters2[ridx]
for didx in range(len(gammas)):
_raster1 = rasters1[didx]
df = pd.DataFrame({'raster1': _raster1.values.flatten(), 'raster2': _raster2.values.flatten()})
corr = round((df.corr()).iloc[0,1],2)
corrs.append(corr)
da_corr = xr.DataArray(np.array(corrs).reshape([len(gammas),len(gammas)]),
coords=[resolution*gammas,resolution*gammas],
dims=['raster2','raster1'])
return (rasters1, rasters2, da_corr)
# # Define parameters
# +
# to load source data
SRTM="srtm90m.Africa20x20.tif"
GRAVITY="WGM2012_Freeair_ponc_2min.Africa20x20.tif"
# rasters below defined in decimal degrees
# this coefficient [km/pixel] for pixel-based filtering and plotting
resolution = 3.7
GAMMA = 28
DGAMMA= 1
# -
# # Load datasets
dem = xr.open_rasterio(SRTM).rename({'x':'lon','y':'lat'})
dem.values = dem.values.astype(float)
dem.values[dem.values == dem.nodatavals[0]] = np.nan
dem
grav = xr.open_rasterio(GRAVITY).rename({'x':'lon','y':'lat'})
grav
# # Compare source datasets
# +
fig, ((ax1,ax2)) = plt.subplots(1, 2, figsize=(10, 4))
dem.plot(ax=ax1, cmap='terrain')
ax1.set_title('SRTM 90m v4.1',fontsize=16)
grav.plot(ax=ax2, cmap='terrain')
ax2.set_title('WGM2012 Free-air\nGravity Anomalies',fontsize=16)
fig.subplots_adjust(hspace=0.2)
plt.show()
# -
#
# # Make correlogram
gammas = np.arange(1,GAMMA+DGAMMA/2,DGAMMA)
(dems,gravs,da_corr) = correlogram(dem, grav, gammas)
float(da_corr.min()),float(da_corr.max())
# +
fig, (ax1,ax2) = plt.subplots(1,2,figsize=(10.5,5))
da_corr.plot(cmap='RdBu_r',ax=ax1, vmin=-1,vmax=1)
ax1.set_xlabel('SRTM Wavelength, km',fontsize=12)
ax1.set_ylabel('WGM2012 Gravity Wavelength, km',fontsize=12)
da_corr.plot.contour(levels=np.linspace(-1,1,41),cmap='RdBu_r',add_colorbar=True, ax=ax2)
ax2.set_xlabel('SRTM Wavelength, km',fontsize=12)
ax2.set_ylabel('WGM2012 Gravity Wavelength, km',fontsize=12)
plt.suptitle('Pearson Correlation Coefficient:\nSRTM 90m v4.1 and WGM2012 Free-air Gravity Anomalies',fontsize=16)
fig.tight_layout(rect=[0.03, 0.0, 1, 0.9])
plt.show()
# +
fig, (ax1,ax2) = plt.subplots(1,2,figsize=(10.5,5))
# define wavelength [km] and index for it
wavelength = 20
gidx = np.argmin((gammas*resolution-wavelength)**2)
gravs[gidx].plot(cmap='RdBu',ax=ax1)
ax1.set_title('WGM2012 Free-air',fontsize=16)
dems[gidx].plot(cmap='RdBu',ax=ax2)
ax2.set_title('SRTM',fontsize=16)
fig.tight_layout(rect=[0.03, 0.0, 1, 0.9])
plt.suptitle('Wavelength %dkm:\nSRTM 90m v4.1 and WGM2012 Free-air Gravity Anomalies\n' % wavelength,fontsize=16)
plt.show()
| Africa/SRTM90m_Freeair_correlogram.Africa20x20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:tensorflow]
# language: python
# name: conda-env-tensorflow-py
# ---
# +
from IPython.core.interactiveshell import InteractiveShell
InteractiveShell.ast_node_interactivity = 'all' # default is ‘last_expr'
# %load_ext autoreload
# %autoreload 2
# +
import json
import os
from collections import Counter
import io
from random import sample
from tqdm import tqdm
import azure.cosmos.cosmos_client as cosmos_client
from azure.storage.blob import BlockBlobService
from PIL import Image
from visualization import visualization_utils
from data_management.annotations import annotation_constants
# -
# # Query for data
#
# This notebook demonstrates the workflow to compile desired images by querying metadata using the database instance and downloading the images stored in blob storage.
# +
# Cosmos DB config
config = {
'ENDPOINT': os.environ.get('COSMOS_ENDPOINT'),
'PRIMARYKEY': os.environ.get('COSMOS_KEY')
}
# Initialize the Cosmos client
client = cosmos_client.CosmosClient(url_connection=config['ENDPOINT'], auth={
'masterKey': config['PRIMARYKEY']})
container_link = 'dbs/camera-trap/colls/images' # database link + container link
# +
with open('datasets.json') as f:
datasets_table = json.load(f)
# this is a json object with the account name as key, and the key to the account as value
with open('blob_account_keys.json') as f:
blob_account_keys = json.load(f)
# -
# ## Select image entries
#
# Example: top 1000 images from a given dataset with bounding boxes, selecting the file name and the dataset so we can plot the labels.
# +
# %%time
dataset_name = 'rspb_gola'
# did not have species in many of these items
query = {'query': '''
SELECT TOP 1000 im.file_name, im.dataset, im.annotations.bbox, im.annotations.species
FROM images im
WHERE im.dataset = "{}" AND ARRAY_LENGTH(im.annotations.bbox) > 0
'''.format(dataset_name)}
options = {
'enableCrossPartitionQuery': True
}
result_iterable = client.QueryItems(container_link, query, options, partition_key='idfg')
# if you want to restrict to one dataset, pass in partition_key=dataset
results = []
for item in iter(result_iterable):
results.append(item)
print('Length of results:', len(results))
# -
len(results)
results[77]
# ## Download images and visualize labels
#
# For large batches, download using `multiprocessing.ThreadPool`.
# +
sample_size = 2
sample_res = sample(results, sample_size)
for im in sample_res:
dataset = im['dataset']
storage_account = datasets_table[dataset]['storage_account']
storage_container = datasets_table[dataset]['container']
path_prefix = datasets_table[dataset]['path_prefix']
print('Creating blob service')
blob_service = BlockBlobService(account_name=storage_account, account_key=blob_account_keys[storage_account])
print('Created')
stream = io.BytesIO()
_ = blob_service.get_blob_to_stream(storage_container, os.path.join(path_prefix, im['file_name']), stream)
print('Downloaded')
image = Image.open(stream)
print('Opened')
boxes = []
classes = []
for i in im['bbox']:
boxes.append(i['bbox_rel'])
classes.append(annotation_constants.bbox_category_name_to_id['animal'])
visualization_utils.render_iMerit_boxes(boxes, classes, image)
print('Visualized')
image
# -
| data_management/cosmos_db/query_for_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Character Theory
#
# To provide the complete classification for $S_n$ we will study the character theory.
#
# In the analysis of the representations of $S_3$, the key thing was to study the eigenvalues of the actions of individual elements of $S_3$. Finding individual eigenvalues is howwever quite difficult. Luckily, it is sufficient to consider their sum, the trace of a matrix, which is much easier to calculate.
#
# ```{admonition} Definition (Trace)
# :class: definition
#
# Let $\phi:V\to V$ be a linear map of a finite dimensional vector space. The trace of $\phi$ is the sume of diagonal entries $a_{11}+a_{22}+\ldots +a_{nn}$ of a matrix representing $\phi$ in any fixed basis for $V$. This is independent of the choice of basis.
#
# ```
#
# ```{admonition} Definition (Character)
# :class: definition
#
# Let $V$ be a finite dimensional representation of a group $G$. The character of the representation is the function:
#
# $$
# \begin{align*}
# &\chi_V:G\to \mathbb{F}\\
# &g\mapsto \mbox{ trace of } g\mbox{ acting } V
# \end{align*}
# $$
#
# ```
#
#
# ```{admonition} Example (Character)
# :class: example
#
# Character of $S_3$: let us compute the characters of the three irreducible representations of $S_3$:
# - the trivial representation $T$ of $S_3$ is one-dimensional and take the value $(1)$ for all elements in $S_3$. Its character is
#
# $$
# \chi_T=(\chi_T(e),\chi_T((12)),\chi_T((13)),\chi_T((23)),\chi_T((123)),\chi_T((132)))=(1,1,1,1,1,1)
# $$
#
# - the alternating representation $A$ of $S_3$ is one-dimensional and takes the value $(1)$ on all even permutations, and $(-1)$ on all odd permutations
#
# $$
# \chi_A=(\chi_A(e),\chi_A((12)),\chi_A((13)),\chi_A((23)),\chi_A((123)),\chi_A((132)))=(1,-1,-1,-1,1,1)
# $$
#
# - the standard representation $S$ characters could be found by writing the matrices for the action of each of the six elements in $S_3$. However, we can find it using the following fact:
#
# > Let $V$ and $W$ be finite dimensional representations of a group $G$. Then
# >
# >$$
# \chi_{V\oplus W}=\chi_V+\chi_W
# $$
# >
# >as functions on $G$.
#
# Now, because the permutation representation $P$ of $S_3$ decomposes as a sum of the trivial representation and the standard representation, we can compute the character of the standard representation using the fact above to get:
#
# $$
# \chi_P=(3,1,1,1,0,0)=\chi_{T\oplus S}=\chi_T+\chi_S
# $$
#
# that implies that
#
# $$
# \chi_S=(3,1,1,1,0,0)-(1,1,1,1,1,1)=(2,0,0,0,-1,-1)
# $$
#
# All characters of $S_3$ are summarized in the table below
#
# $$
# \begin{array}{c|c|c|c|c|c|c}
# &e&(12)&(13)&(23)&(123)&(132)\\
# \hline
# \mbox{trivial}&1&1&1&1&1&1\\
# \hline
# \mbox{alternating}&1&-1&-1&-1&1&1\\
# \hline
# \mbox{standard}&2&0&0&0&-1&-1
# \end{array}
# $$
#
# The characters of any representation $V$ of $S_3$ can be obtained from these three by decomposing into irreducibles:
#
# $$
# V=T^{a}\oplus A^b\oplus S^c
# $$
#
# and using
#
# $$
# \chi_V=a \chi_T+b\chi_A+c\chi_S
# $$
#
# ```
#
# Isomorphic representations have the same character, but also the converse is true: the character completely determines the representation up to an isomorphism. Much stronger statement is true: the rows of the character table are orthonormal to each other. This means that if $\chi_V$ and $\chi_W$ are characters of irreducible representations of $G$ then their scalar product is either $1$ or $0$, depending on whether $V\cong W$ or not.
#
# This implies that the number of different irreducible representations of $G$ must be smaller than $|G|$. In fact, there is a better bound on the number of irreducible representations of a finite group. Notice that each character takes the same valus on elements with the same cycle structure. This comes from a property of traces that implies that
#
# $$
# \chi_V(hgh^{-1})=\chi_V(g)
# $$
#
# In other words, if $g'=h gh^{-1}$ then $\chi_V(g')=\chi_V(g)$. We call such elements conjugate to each other and for symmetric groups there is a general statement that two permutations are conjugated if they have the same cycle structure. To avoid redundancy, we can therefore write a simplified character table for $S_3$:
#
# $$
# \begin{array}{c|c|c|c}
# g&e&(12)&(123)\\
# \hline
# \#&1&3&2\\
# \hline
# \mbox{trivial}&1&1&1\\
# \hline
# \mbox{alternating}&1&-1&1\\
# \hline
# \mbox{standard}&2&0&-1
# \end{array}
# $$
#
# where in the second row we indicated the number of elements in each conjugacy class.
#
# To summarize, we have the following useful statement: for finite-dimensional representations of a finite group $G$ we have:
# - there are at most $t$ irreducible representations of $G$, where $t$ is the number of conjugacy classes of $G$.
# -each representation is determined (up to an isomorphism) by its character
# - a complex representation $V$ is irreducible if and only if $\langle\chi_V,\chi_V\rangle=1$
# - the multiplicity of a complex irreducible representation $W$ in a representation $V$ is given by $\langle \chi_V,\chi_W\rangle$,
#
# where we defined
#
# $$
# \langle\chi_W,\chi_V\rangle =\frac{1}{|G|}\sum_{g\in G}\chi_W(g)\overline{\chi_V(g)}
# $$
# where the overline indicates the complex conjugate.
#
# This provides us with a powerful tool for analysing complex representaions of finite groups. In particular, we can take any representation and decompose it into irreducible representations using the character theory. Let us take the regular representation $R$ of $S_3$. It is a six-dimensional representation obtained by left multiplication by group elements. We already showed that there are only three representation of $S_3$: trivial $T$, alternating $A$ and standard $S$. Therefore, we have a decomposition
#
# $$
# R\cong T^{a}\oplus A^b\oplus S^c
# $$
#
# for some non-negative integers $a,b,c$. This produces the following relation on characters:
#
# $$
# \chi_V=a \chi_T+b\chi_A+c\chi_S
# $$
#
# The character of the regular representation is $(6,0,0)$. This leads to a system of linear equations:
#
# $$
# (6,0,0)=a(1,1,1)+b(1,-1,1)+c(2,0,-1)
# $$
#
# which is solved by: $(a,b,c)=(1,1,2)$. therefore, the regular representation of $S_3$ decomposes as:
#
# $$
# R\cong T\oplus A\oplus S^2
# $$
#
# This statement can be generalised to any finite group:
#
# ```{admonition} Proposition
# :class: proposition
#
# The regular representation $R$ of any finite group $G$ decomposes (over $\mathbb{C}$) as
#
# $$
# R\cong W_1^{\dim W_1}\oplus W_2^{\dim W_2}\oplus \ldots \oplus W_t^{\dim W_t}
# $$
#
# with every irreducible representation $W_i$ appearing exactly $\dim W_i$ times
# ```
#
# This proposition leads to a useful relation:
#
# $$
# |G|=\sum_i(\dim W_i)^2
# $$
#
# where the sum is taken over all irreducible complex representations of $G$.
#
# ```{admonition} Example (Complete classification of irreducible representations of $D_4$)
# :class: example
#
# For $D_4$ we have $5$ conjugacy classes of elements:
#
# $$
# C_1=\{e\}\quad C_2=\{A,A^3\}\quad C_3=\{A^2\}\quad C_4=\{B,A^2 B\}\quad C_5=\{AB,A^3 B\}
# $$
#
# This means that there are $5$ irreducible complex representations of $D_4$. Moreover, we found $2$ of them already:
#
# $$
# \begin{array}{c|c|c|c|c|c}
# g&C_1&C_2&C_3&C_4&C_5\\
# \hline
# \# &1&2&1&2&2\\
# \hline
# \mbox{trivial}&1&1&1&1&1\\
# \hline
# \mbox{tautological}&2&0&-2&0&0
# \end{array}
# $$
#
# Let $W_3$, $W_4$ and $W_5$ be the remaining irreducible representation, with dimensions $w_3,w_4$ and $w_5$. Then
#
# $$
# |D_4|=8=1^2+2^2+w_1^2+w_4^2+w_5^2
# $$
#
# that implies that $w_3=w_4=w_5=1$. All remaining representaions are one-dimensional. They have the following characters:
#
#
# $$
# \begin{array}{c|c|c|c|c|c}
# g&C_1&C_2&c_3&C_4&C_5\\
# \hline
# \#&1&2&1&2&2\\
# \hline
# \mbox{trivial}&1&1&1&1&1\\
# \hline
# \mbox{tautological}&2&0&-2&0&0\\
# \hline
# W_3&1&1&1&-1&-1\\
# \hline
# W_4&1&-1&1&1&-1\\
# \hline
# W_5&1&-1&1&-1&1
# \end{array}
# $$
#
# It is easy to checj that they are orthonormal. For example
#
# $$
# \langle \chi_{W_1},\chi_{W_3}\rangle=\frac{1}{8}(1\cdot 1\cdot 1+2\cdot 1\cdot 1+1\cdot 1\cdot 1+2\cdot 1\cdot (-1)+1\cdot 1\cdot (-1))=0
# $$
#
# Also
#
# $$
# \langle \chi_{W_3},\chi_{W_3}\rangle=\frac{1}{8}(1\cdot 1\cdot 1+2\cdot 1\cdot 1+1\cdot 1\cdot 1+2\cdot (-1)\cdot (-1)+1\cdot (-1)\cdot (-1))=1
# $$
#
# ```
| _build/jupyter_execute/Lectures/Lecture7.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gain Statistical Insights into Your DataTable
#
# Woodwork provides methods on DataTable to allow you to use the typing information inherent in a DataTable to better understand your data.
#
# Follow along to learn how to use `describe` and `mutual_information` on a retail DataTable so that you can see the full capabilities of the functions.
# +
import pandas as pd
from woodwork import DataTable
from woodwork.demo import load_retail
dt = load_retail()
dt
# -
# ## DataTable.describe
#
# Use `dt.describe()` to calculate statistics for the DataColumns in a DataTable in the format of a pandas DataFrame with the relevant calculations done for each DataColumn.
dt.describe()
# There are a couple things to note in the above dataframe:
#
# - The DataTable's index, `order_product_id`, is not included
# - We provide each DataColumn's typing information according to Woodwork's typing system
# - Any statistics that can't be calculated for a DataColumn, say `num_false` on a `Datetime` are filled with `NaN`.
# - Null values do not get counted in any of the calculations other than `nunique`
# ## DataTable.value_counts
#
# Use `dt.value_counts()` to calculate the most frequent values for each Data Columns that has `category` as a standard tag. This returns a dictionary where each DataColumn is associated with a sorted list of dictionaries. Each dictionary contains `value` and `count`.
dt.value_counts()
# ## DataTable.mutual_information
#
# `dt.mutual_information` calculates the mutual information between all pairs of relevant DataColumns. Certain types, like strings, can't have mutual information calculated.
#
# The mutual information between columns `A` and `B` can be understood as the amount of knowledge you can have about column `A` if you have the values of column `B`. The more mutual information there is between `A` and `B`, the less uncertainty there is in `A` knowing `B`, and vice versa.
dt.mutual_information()
# #### Available Parameters
# `dt.mutual_information` provides two parameters for tuning the mutual information calculation.
#
# - `num_bins` - In order to calculate mutual information on continuous data, Woodwork bins numeric data into categories. This parameter allows you to choose the number of bins with which to categorize data.
# - Defaults to using 10 bins
# - The more bins there are, the more variety a column will have. The number of bins used should accurately portray the spread of the data.
# - `nrows` - If `nrows` is set at a value below the number of rows in the DataTable, that number of rows is randomly sampled from the underlying data
# - Defaults to using all the available rows.
# - Decreasing the number of rows can speed up the mutual information calculation on a DataTable with many rows, but you should be careful that the number being sampled is large enough to accurately portray the data.
# Now that you understand the parameters, you can explore changing the number of bins. Note—this only affects numeric Data Columns `quantity` and `unit_price`. Increase the number of bins from 10 to 50, only showing the impacted columns.
mi = dt.mutual_information()
mi[mi['column_1'].isin(['unit_price', 'quantity']) | mi['column_2'].isin(['unit_price', 'quantity'])]
mi = dt.mutual_information(num_bins = 50)
mi[mi['column_1'].isin(['unit_price', 'quantity']) | mi['column_2'].isin(['unit_price', 'quantity'])]
| docs/source/guides/statistical_insights.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Game of Life
import numpy as np
import numba
import skimage
from skimage import data, color, feature
import matplotlib.pyplot as plt
# ## Create image
image = skimage.img_as_float(color.rgb2gray(data.chelsea())).astype(np.float32)
image = skimage.transform.resize(image, (128,128))
image = feature.canny(image, sigma=1)
fig = plt.figure()
plt.imshow(image, cmap=plt.cm.gray)
# ## Cellular Automata Update
@numba.jit
def apply_rules(image, out_image):
# Prepare neighbouring indices
idx = []
for i in range(-1, 2):
for j in range(-1, 2):
if i == 0 and j == 0:
continue
idx.append([i, j])
# Apply the cellular automata from "conways_game_of_life"
for x in range(1, image.shape[0]-1):
for y in range(1, image.shape[1]-1):
num_neighbours = 0
for i in range(len(idx)):
if image[x+idx[i][0], y+idx[i][1]] == 1.0:
num_neighbours += 1
out_image[x, y] = image[x, y]
if out_image[x, y] == 1.0: # live cell
if num_neighbours < 2: # under population.
out_image[x, y] = 0.0
elif num_neighbours > 3: # overpopulation
out_image[x, y] = 0.0
else: # dead cell
if num_neighbours == 3: # reproduction
out_image[x, y] = 1.0
NUM_ITERATIONS = 50
buffer = [np.copy(image), np.copy(image)]
for i in range(NUM_ITERATIONS):
apply_rules(buffer[0], buffer[1])
buffer[0], buffer[1] = buffer[1], buffer[0] # swap buffers
if i%5 == 0:
plt.imshow(buffer[0], cmap=plt.cm.gray)
fig = plt.figure()
plt.show()
# ## References
# - <NAME>, <NAME>, and <NAME>. "Optimizing memory access patterns for cellular automata on GPUs." In GPU Computing Gems Jade Edition, pp. 67-75. 2011.
# - <NAME>. "Mathematical games: The fantastic combinations of <NAME>’s new solitaire game “life”." Scientific American 223, no. 4 (1970): 120-123.
| stencil_codes/game_of_life.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import cv2
import os
from tqdm.auto import tqdm
from IPython.display import display
import IJB_evals as IJB
templates, medias, p1, p2, label, img_names, landmarks, face_scores = IJB.extract_IJB_data_11(
'./', 'IJBC', force_reload=False
)
for img, landmark in tqdm(zip(img_names, landmarks), total=len(img_names)):
cropped_img = IJB.face_align_landmark(cv2.imread(img), landmark)
cv2.imwrite(os.path.join('IJBC/final_crop', img), cropped_img)
# # baseline (identity)
# + jupyter={"outputs_hidden": true}
args = Args(subset='IJBC',
is_bunch=True,
restore_embs_left='IJB_result/MS1MV2-ResNet100-Arcface_IJBC.npz',
save_result='../../../../results/baseline_nomap.npz')
df, fig = IJB.main(args)
# -
class Args:
def __init__(self, subset='IJBC', is_bunch=False, restore_embs_left=None, restore_embs_right=None, fit_mapping=False, fit_flips=False, decay_coef=0.0, pre_template_map=False, save_result="IJB_result/{model_name}_{subset}.npz"):
self.subset = subset
self.is_bunch=is_bunch
self.restore_embs_left = restore_embs_left
self.restore_embs_right = restore_embs_right
self.fit_mapping = fit_mapping
self.fit_flips = fit_flips
self.decay_coef = decay_coef
self.pre_template_map = pre_template_map
self.save_result = save_result
self.save_embeddings = False
self.model_file = None
self.data_path = './'
self.batch_size=64
self.save_label=False
self.force_reload=False
self.is_one_2_N=False
self.plot_only=None
def __str__(self):
return str(self.__class__) + ": " + str(self.__dict__)
| evaluation/IJB/save_crops.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In this notebook, I will work through MNIST data and look at some classification algorithms
# # Importing common libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# # Importing the Data
# We can use `sklearn.datasets.fetch_openml` to get the MNIST data.
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1)
mnist.keys()
# The MNIST data is already broken out into features and labels
X, y = mnist['data'], mnist['target']
X.shape
# From the shape, we can see it has 7000 rows and 784 columns. Each row has 784 features which represents a 28x28 image.
# Let's look at the first digit
some_digit = X[0]
some_digit_image = some_digit.reshape(28, 28)
plt.imshow(some_digit_image, cmap='binary')
plt.show()
# As we can see, it's a '5'. Let's look at the label for this row.
y[0]
# Yay! it's indeed a '5'. Thing to note here is that the target values are stored in string format, so let's convert the target values into integers.
y = y.astype(np.uint8)
# # Splitting the Data into Training and Test sets
#
# The data is already split into training and test data based on the index. First 60000 rows are training set and the remaining ones are test set. So we can easily split the data without the need of any librarires
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
# # Machine Learning Algorithms
#
# ## Binary Algorithms:
# - __Stochastic Gradient Descent (SGD) Classifier__
# - Capable of handling very large datasets efficiently, as SGD deals with training instances independently
# - Suited for online training
#
# ## Multiclass Classification
#
# Multiclass Classifiers _(also called as Multinomial Classifiers)_ can distinguish between more than two classes.
#
# Some algorithms (such as Random Forest Classifiers or Naive Bayes Classifiers) are capable of handling multiple classes directly. Others (such as Support Vector Machine Classifiers or Linear Classifiers) are strictly Binary Classifiers. However, there are various strategies that we can use to perform Multiclass Classification using Binary Classifierrs:
#
# - __One-versus-the-rest__ (OvA):
# - One way to create a system that can classify the digit images into 10 classes (from 0 to 9) is to train 10 Binary Classifierrs, one for each digit (0-detector, 1-detector, and so on).
# - Then when we want to classsify an image, we get the decision score from each classifier for that image and we select the class whose classifier output (decision score) is the highest.
# - For most Binay Classification algorithms, OvA is preferred.
#
# - __One-versus-One__ (OvO):
# - Another strategy is to train a Binary Classifier for every pair of digits: one to distinguish 0s and 1s, another to distinguish 0s and 2s, another for 1s and 2s, and so on.
# - If there are N classes, then we need to train N$*$(N-1)$/$2 classifiers. For our problem, this means training 45 Binary Classifiers.
# - When we want to classify an image, we have to run the image through all 45 classifiers and see which class wins the most duels.
# - The main advantage of OvO is that each classifier only needs to be trained on the part of the training set for the two classes it must distinguish.
# - Some algorithms (such as Support Vector Machine Classifer) scale poorly with the size of training set, so for these algorithms OvO is preferred since it is faster to train many classifiers on small training sets than training few classifiers on large training sets.
#
# Scikit-Learn detects when you try to use a binary classification algorithm for a multiclass classification task, and it automatically runs OvA (except for SVM classifier for which it uses OvO). Let's try this with our `SGDClassifier`:
# Since we are working with Binary Algorithms, let's focus on only one digit '5'
y_train_5 = (y_train == 5)
y_test_y = (y_test == 5)
# #### Stochastic Gradient Descent (SGD) Classifier:
from sklearn.linear_model import SGDClassifier
sgd_clf = SGDClassifier(random_state=42)
sgd_clf.fit(X_train, y_train_5)
# Now let's predict the value of the first row using this model
sgd_clf.predict([some_digit])
# As we can see the classifier predicts the image correctly
# # Evaluating the Model
#
# - __Cross Validation Score__
# - For a classification model, this approach isn't used as often
# - __Confusion Matrix__
# - Much better way to evaluate a classification model
# - We need set of predictions in order to compute confusion matrix
# - Overall understanding about the predictions and errors
# - Using the confusion matrix, we can calculate various evaluating metrics:
# - __Precision:__
# - How many predictions were actually correct
# - True Positive / (True Positive + False Positive)
# - __Recall:__
# - How many true class we are able to catch
# - True Positive / (True Positive + False Negative)
# - __F1 Score:__
# - It is the harmonic mean of precision and reacall, that means it gives much more weight to the lower value
# - 2 * (Precision * Recall) / (Precision + Recall)
# - __Precision vs Recall Graphs__
# - Precision Recall Curve
# - Precision vs Recall
# - Receiver Operator Curve (ROC)
# - Another common tool used with binary classifiers
# - Similar to precision recall cureve, but instead of plotting precision vs recall, the ROC curve plots true positive rate (another name for recall) against false positive rate
# - The FPR is the ratio of negative instances that are incorrectly classified as positive. It is equal to one minus TNR. The TNR is also called specificity.
# - Hence, the ROC curve plots `Sensitivity _(recall)_` versus `1 - Specificity`
# #### Cross Validation Score
# +
from sklearn.model_selection import cross_val_score
sgd_cv_scores = cross_val_score(sgd_clf, X_train, y_train, cv=3, scoring='accuracy')
for accuracy_ in sgd_cv_scores:
print(f'Accuracy: {round(accuracy_, 2)}')
# -
# #### Confusion Matrix
# In order to get the confusion matrix, we need predictions which then can be compared with the original values to get the confusion matrix. So let's get all the predictions, we can use `cross_val_predict()` to get our predictions. This performs K-fold cross-validation, but instead of returning the scores, it returns the predictions made on each test fold.
from sklearn.model_selection import cross_val_predict
y_train_5_pred = cross_val_predict(sgd_clf, X_train, y_train_5, cv=5)
# Now let's get our confusion matrix
from sklearn.metrics import confusion_matrix
confusion_matrix(y_train_5, y_train_5_pred)
# Each row in confusion matrix represents an actual class, while each column represents a predicted class.
# - The first row of this matrix is the Negative Class (non-5 images)
# - 53,115 were correctly classified as non-5s (True Negative)
# - 1,464 were incorrectly classified as 5s (False Positives)
# - The second row of this matrix is the Positive Class (5 images)
# - 916 were incorrectly classified as non-5s (False Negative)
# - 4,505 were correctly classified as 5s (True Positive)
#
# | Actual Values | Predicted Negative | Predicted Positive |
# | ------------- | ------------------ | ------------------ |
# | Negative | True Negative | False Positive |
# | Positive | False Negative | True Positive |
# Now let's calculate various evaluating metrics:
# ##### Precision, Recall and F1 Score:
# +
from sklearn.metrics import precision_score, recall_score, f1_score
sgd_precision = precision_score(y_train_5, y_train_5_pred)
sgd_recall = recall_score(y_train_5, y_train_5_pred)
sgd_f1_score = f1_score(y_train_5, y_train_5_pred)
print(f'Precision: {sgd_precision: .2%}')
print(f'Recall: {sgd_recall: .2%}')
print(f'F1 Score: {sgd_f1_score: .2%}')
# -
# We can get a particular Precision or Recall by changing the threshods. By default Scikit-Learn doesn't let us set the threshold directly, but it does give us access to decision scores that it uses to make the predictions.
# Please note: there is tradeoff between Precision and Recall, meaning if you try to increase one, the other one decreases.
sgd_y_scores = sgd_clf.decision_function([some_digit])
print(f'y_score: {sgd_y_scores}')
threshold_ = 0
y_some_digit_pred = (sgd_y_scores > threshold_)
print(f'Some digit prediction: {y_some_digit_pred}')
# By default, `SGDClassifier` uses the threshold of __0__. That's why the previous code returns same result as `predict()` method. Let's raise the threshold and see the result
threshold_ = 5000
y_some_digit_pred = (sgd_y_scores > threshold_)
print(f'Some digit prediction: {y_some_digit_pred}')
# Let's get scores of all the instances and look at ways to decide the threshold manually
sgd_y_scores = cross_val_predict(sgd_clf, X_train, y_train_5, cv=3, method='decision_function')
# ##### Precision Recall Curve
# Now with these scores, we can compute precision and recall for all the possible thresholds using `precision_recall_curve()` method:
# +
from sklearn.metrics import precision_recall_curve
sgd_precisions, sgd_recalls, sgd_thresholds = precision_recall_curve(y_train_5, sgd_y_scores)
# -
# Now, we can plot the precision and recalls as functions of threshold values:
# +
plt.figure(figsize=(12, 4))
plt.plot(sgd_thresholds, sgd_precisions[:-1], '--b', linewidth=2, label='Prediction')
plt.plot(sgd_thresholds, sgd_recalls[:-1], 'g-', linewidth=2, label='Recall')
plt.legend(loc='center right', fontsize=16)
plt.title('Precision and Recall vs Threshold', fontsize=20)
plt.xlabel('Threshold', fontsize=16)
plt.grid()
plt.axis([-50000, 50000, 0, 1])
sgd_precision_value = 0.90
sgd_threshold_value = sgd_thresholds[np.argmax(sgd_precisions >= sgd_precision_value)]
sgd_recall_value = sgd_recalls[np.argmax(sgd_thresholds >= sgd_threshold_value)]
plt.plot([-50000, sgd_threshold_value], [sgd_precision_value, sgd_precision_value], 'r:')
plt.plot([-50000, sgd_threshold_value], [sgd_recall_value, sgd_recall_value], 'r:')
plt.plot([sgd_threshold_value, sgd_threshold_value], [sgd_precision_value, 0], 'r:')
plt.plot([sgd_threshold_value, sgd_threshold_value], [sgd_precision_value, sgd_precision_value], 'ro')
plt.plot([sgd_threshold_value, sgd_threshold_value], [sgd_recall_value, sgd_recall_value], 'ro')
plt.show()
# -
# Using the chart above:
# - We can see that as Precision increases, Recall decreases
# - Precision curve is a little bumpier, that is because precision may sometimes decrease as we increase threshold
# - We can find the prefect threshold for our analysis, by comparing the precision and recall at that given threshold
# ##### Precision/Recall
# Another way to select a good precision/recall tradeoff is to plot precision directly with recall:
plt.plot(sgd_recalls, sgd_precisions, 'b-', linewidth=2)
plt.xlabel('Recall', fontsize=16)
plt.ylabel('Precisions', fontsize=16)
plt.title('Precision vs Recall', fontsize=20)
plt.axis([0, 1, 0, 1])
plt.grid()
plt.plot()
# Looking at the above chart:
# - Precision falls sharply around 0.8 (80%) recall
# - We would choose a precision/recall tradeoff just before that drop off
# Let's suppose we decided to aim for a 90% precision value. We can look at the first plot to find out the threshold we need. To get a precise value, we can search for the lowest threshold that gives us at least 90% precision value using `np.argmax()` method. This gives us the first index of the maximum value.
sgd_threshold_90_precision = sgd_thresholds[np.argmax(sgd_precisions >= 0.90)]
print(f'Threshold value for 90% Precision: {round(sgd_threshold_90_precision, 2)}')
# Now, in order to make predicions using this threshold, instead of calling the classifier's `predict()` method, we can just compare the scores with the threshold
sgd_y_train_pred_90 = (sgd_y_scores >= sgd_threshold_90_precision)
# Now let's check the scores:
print('Precision: ', precision_score(y_train_5, sgd_y_train_pred_90))
print('Recall: ', recall_score(y_train_5, sgd_y_train_pred_90))
# ##### ROC Curve
# +
from sklearn.metrics import roc_curve
sgd_fpr, sgd_tpr, sgd_thresholds = roc_curve(y_train_5, sgd_y_scores)
# -
# Now let's plot the graph:
# +
plt.plot(sgd_fpr, sgd_tpr, 'b-', linewidth=2)
plt.plot([0, 1], [0, 1], 'k--', alpha=0.4)
plt.xlabel('False Positive Rate (Fall out)', fontsize=16)
plt.ylabel('True Positive Rate (Recall)', fontsize=16)
plt.title('ROC Curve', fontsize=20)
plt.grid()
plt.axis([0, 1, 0, 1])
plt.show()
# -
# One way to compare classifiers is to measure the area under the curve (AUC). A perfect classifier will have ROC AUC = 1, whereas a purely random classifier with have ROC AUC = 0.5
#
# Scikit-Learn provides a method `roc_auc_score()` to compute this value:
from sklearn.metrics import roc_auc_score
sgd_auc_score = roc_auc_score(y_train_5, sgd_y_scores)
print('ROC AUC Score: ', round(sgd_auc_score, 2))
# _**NOTE:**
# Since the ROC curve is so similar to the precision/recall (or PR) curve, you may wonder how to decide which one to use. As a rule of thumb, you should prefer the PR curve whenever the positive class is rare or when you care more about the false positives than the false negatives, and the ROC curve otherwise. For example, looking at the previous ROC curve (and the ROC AUC score), you may think that the classifier is really good. But this is mostly because there are few positives (5s) compared to the negatives (non-5s). In contrast, the PR curve makes it clear that the classifier has room for improvement (the curve could be closer to the topright corner)._
# #### Random Forest Classifer:
#
# _RandomForestClassifier does not have a `decision_function()` method. Instead, it has `predict_proba()` method. Scikit-Learn classifiers generally have one or the other. The `predict_proba()` method returns an array containing a row per instance and a column per class, each containing the probability that the given instance belongs to the given class (eg: 70% chance that the image represents a 5)_
# +
from sklearn.ensemble import RandomForestClassifier
forest_clf = RandomForestClassifier(random_state=42)
forest_y_probas = cross_val_predict(forest_clf, X_train, y_train_5, cv=3, method='predict_proba')
# -
# In order to plot the ROC curve we need scores, not probabilites. A simple solution is to use the positive class's probability as the score:
forest_y_scores = forest_y_probas[:, 1]
forest_fpr, forest_tpr, forest_threshold = roc_curve(y_train_5, forest_y_scores)
# Now let's plot the ROC Curve:
plt.figure(figsize=(8, 6))
plt.plot(forest_fpr, forest_tpr, 'b-', label='Random Forest')
plt.plot(sgd_fpr, sgd_tpr, 'b:', label='SGD Classifier')
plt.xlabel('False Positive Rate', fontsize=16)
plt.ylabel('True Positive Rate', fontsize=16)
plt.title('ROC Curve', fontsize=20)
plt.legend(loc='lower right')
plt.axis([0, 1, 0, 1])
plt.grid()
plt.show()
# From the above chart:
# - We can see that the Random Forest Classifier's ROC is much closer to the corner, suggesting that Random Forest Classifier is better than SGD Classifier
# Now let's calculate the AUC Score and compare with SGD Classifier
forest_auc_score = roc_auc_score(y_train_5, forest_y_scores)
print(f'Forest AUC Score: {forest_auc_score:.3f}\nSGD AUC Score: {sgd_auc_score:.3f}')
# Here we can see:
# - AUC for Random Forest is higher compared to SGD Classifer
# Let's calculate Precision, Recall and F1 Scores for Random Forest:
#
# _In order to these, we need predictions instead of scores_
forest_y_train_pred = cross_val_predict(forest_clf, X_train, y_train_5, cv=3)
# +
forest_precision = precision_score(y_train_5, forest_y_train_pred)
forest_recall = recall_score(y_train_5, forest_y_train_pred)
forest_f1score = f1_score(y_train_5, forest_y_train_pred)
print(f'Precision: {forest_precision:.2%}')
print(f'Recall: {forest_recall:.2%}')
print(f'F1 Score: {forest_f1score:.2%}')
# -
# #### Multiclass Classification
# ##### SGD Classifier
sgd_clf = SGDClassifier(random_state=42)
sgd_clf.fit(X_train, y_train)
sgd_clf.predict([some_digit])
# That was easy! This code trains the `SGDClassifier` on the training set using the original target classes from 0 to 9, instead of the 5-versus-all target classes. Then it makes prediction (a wrong one in this case). Under the hood, Scikit-Learn acutally traiined 10 binary classifiers, got their decision scores for the image, and selected the class with the highest score.
#
# To see that, we can call the `decision_function()` method. Instead of returning just one score per instance, it now returns 10 scores, one per each class:
some_digit_scores = sgd_clf.decision_function([some_digit])
print('Scores:', some_digit_scores)
print('Classes:', sgd_clf.classes_)
print('Output Class:', sgd_clf.classes_[np.argmax(some_digit_scores)])
# Here, we see that we have 10 scores and the highest score belongs to the class 3.
#
# _**NOTE:**
# When a classifier is trained, it stores the list of classes in its `classes` attribute, ordered by value. In this case, the index of each class in the `classes` array conveniently matches the class itself, but in general we won't be so lucky._
#
# If you want to force Scikit-Learn to use OvO or OvA, you can use the `OneVsOneClassifier` or `OneVsRestClassifier` classes. Simply create an instance and pass a binary classifier to its constructor. For example, this code creates a multicalss classifier using OvO strategy, based on `SGDClassifier`:
# +
from sklearn.multiclass import OneVsOneClassifier
ovo_clf = OneVsOneClassifier(estimator=SGDClassifier(random_state=42))
ovo_clf.fit(X_train, y_train)
ovo_clf.predict([some_digit])
# -
# __NOTE:__ When using `SGDClassifier` as OvO it predicted the some_digit correctly
print('Length of the estimator', len(ovo_clf.estimators_))
# ##### Random Forest Classifier
forest_clf = RandomForestClassifier(random_state=42)
forest_clf.fit(X_train, y_train)
forest_clf.predict([some_digit])
# This time Scikit-Learn did not have to run OvA or OvO because Random Forest Classifier can directly classify instances into multiple classes. We can call the `predict_proba()` to get the list of probabilities that the classifier assigned to each instance for each class:
forest_clf.predict_proba([some_digit])
# We can see that the classifier is fairly confident about it's prediction: the 0.9 at the 5th index in the array means that the model estimates a 90% probability that the image represents a 5. It also thinks that the image could instead be a 2, a 3, or a 9, respectively with 1%, 8% and 1% probability.
# Evaluating the performance of this classifier:
# +
from sklearn.model_selection import cross_val_score
cross_val_score(forest_clf, X_train, y_train, cv=3, scoring='accuracy')
# -
# As we can see, Random Forest Classifier is over 96% accurate on all the test folds, which is not a bad score. We can still fine tune it using various hyperparameters.
# ##### Error Analysis
#
# Let's look at the confusion matrics. In order to do so, we need predictions
y_train_pred = cross_val_predict(forest_clf, X_train, y_train, cv=5)
conf_mx = confusion_matrix(y_train, y_train_pred)
print(conf_mx)
# It's often easier to visualize the confusion matrix. Let's try that:
plt.matshow(conf_mx, cmap='gray')
plt.show()
# The confusion matrix looks good since majority of the values are on the diagonal line.
# Let's focus the plot on the errors. First, we need to divide each value in the confusion matrix by the number of images in the corresponding class, so we can compare error rates instead of absolute number of errors
row_sums = conf_mx.sum(axis=1, keepdims=True)
norm_conf_mx = conf_mx/row_sums
# Now let's fill the diagonal with zeros to keep only the errors, and plot the result:
# +
np.fill_diagonal(norm_conf_mx, 0)
plt.matshow(norm_conf_mx, cmap='gray')
plt.xlabel('Predicted Class')
plt.ylabel('Actual Class')
plt.show()
# -
# ## Multilabel Classification
#
# Until now each instance has always been assigned to just one class. In some cases we may want our classifier to output multiple classes for each instance. For example, consider a face-recognition classifier: what sould it do if it recognizes several people on the same picture? Of course it should attach one tag per person it recognizes. Say the classifier has been trained to recognize three faces, Jack, Jill and John; then when it is shown a picture of Jack and Jill, it should output (1, 1, 0) (meaning "Jack Yes, Jill Yes, John No"). Such a classification system that outputs multiple binary tags is called Multilabel Classification Sysytem.
#
# Let's look at a simple example which detects if the number is greater than 7 and is Odd or Even. For this example, we will be using `KNeighborsClassifier`:
# +
from sklearn.neighbors import KNeighborsClassifier
y_train_large = (y_train >= 7)
y_train_odd = (y_train % 2 == 1)
y_multilabel = np.c_[y_train_large, y_train_odd]
knn_clf = KNeighborsClassifier()
knn_clf.fit(X_train, y_multilabel)
# -
# _**NOTE:** KNeighborsClassifier supports multilabel classification, but not all classifiers do._
# Let's predict the some_digit using the above classifier
knn_clf.predict([some_digit])
# We can see there are two predictions:
# - 1st Prediction (False) is correct as "5" is not larger than "7"
# - 2nd Prediction (True) is correct as "5" is an odd number
# ##### Evaluating Multilabel Classifiers
# There are many ways to evaluate a multilabel classifier, and selecting the right metric really depends on our project. For example, one approach is to measure F1 score for each individual label (or any other binary classifier metric discussed earlier), then simply compute the average score.
#
# We need predictions in order to calculate the F1 Score, so let's get that first:
# The following code took more than 50 mins to run
y_train_knn_pred = cross_val_predict(knn_clf, X_train, y_multilabel, cv=3)
# Now, let's calculate the F1 Score
knn_f1_score = f1_score(y_multilabel, y_train_knn_pred, average='macro')
print(f'KNN F1 Score: {knn_f1_score: .2%}')
# This assumes that all labels are equally important, which may not be the case. In particular, if you have many more pictures of Jill than of Jack and John, you may want to give more weight to the classifier's score on pictures of Jill. One simple option is to give each label a weight equal to its support (i.e., the number of instances with that target label). To do this simply set `average='weighted'` in the preceding code.
# #### Multioutput Classification
#
# It is also known as **Multioutput Multiclass Classification**. It is simply a generalization of multilabel classification where each label can be a multiclass (i.e., it can have more than two possible values).
#
# To illustrate this, let's build a system that removes noise from images. It will take as input a noisy digit image, and it will (hopefully) output a clean digit image, represented as an array of pixel intensities, just like the MNIST images. Notice that the classifier's output is multilabel (one label per pixel) and each label can have multiple values (pixel intensity ranges from 0 to 255). It is thus an example of Multioutput Classification system.
#
# Let's start by creating the training and test sets by taking MNIST images and adding noise to their pixel intensities unsing NumPy's `randint()` function. The target images will be the original images.
noise = np.random.randint(0, 100, (len(X_train), 784))
X_train_mod = X_train + noise
noise = np.random.randint(0, 100, (len(X_test), 784))
X_test_mod = X_test + noise
y_train_mod = X_train
y_test_mod = X_test
# Let's look at the images before the noise reduction algorithm:
# +
some_index = 0
noise_image = X_test_mod[some_index].reshape(28, 28)
image = y_test_mod[some_index].reshape(28, 28)
plt.subplot(121); plt.imshow(noise_image, cmap='binary')
plt.axis('off')
plt.subplot(122); plt.imshow(image, cmap='binary')
plt.axis('off')
# -
# On the left is the noisy input image, and on the right is the clean target image. Now let's train the classifer and make it clean this image
knn_clf.fit(X_train_mod, y_train_mod)
clean_digit = knn_clf.predict([X_test_mod[some_index]]).reshape(28, 28)
plt.imshow(clean_digit, cmap='binary')
| MNIST Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Solar Energy Adoption Model
#
# Our aim is to predict where adoption of residential solar power may be highest in the United States by looking at historical adoption data combined with a variety of demographic, economic, and regulatory features. The labels in the labeled dataset come from the National Renewable Energy Lab's OpenPV project, and the features are derived from a wide variety of sources (census, economic survey, state regulation and incentives data, EIA, etc).
#
# We measure adoption as the number of installations per 100 households in a given zipcode. Our hypothesis is that zipcodes with particularly high rates of adoption have certain attributes in common.
#
# Note that this project focuses on residential adoption, but a very similar process can be executed for commercial customers by expressing adoption as a function of installs per X number of businesses rather than households. We have largely prepared this data in the repo if anyone wants to give it a go.
#
# ##### this notebook casts the problem as binary, rather than multi-class; our previous models had a hard time distinguishing between the high and medium classes, so this combines the two and applies several other estimators
# +
import pandas as pd
import numpy as np
from sklearn.preprocessing import MinMaxScaler
from sqlalchemy import create_engine
from collections import Counter
import matplotlib.pyplot as plt
import seaborn as sns
import yellowbrick
import altair as alt
# -
# #### Get the data.
# Create a connection to the psql instance on Amazon RDS, pull down the residential or commercial adoption data and store it in a dataframe. A handful of the features we plan to use still have non-numeric values in them (income, household size), so we'll want to impute those. Given that the data is already ordered by zipcode, we can assume that contiguous zipcodes share some demographic characteristics and backfill or forward-fill those values (some consecutive zipcodes are non-contiguous, but we'll choose to live with this limitation for now).
#
# We also want to one-hot encode the region column
#
# Note: now that we're not writing to the source files anymore, we can just grab them off the repo, which seems sportier than RDS
# +
# pwd = ''
# engine = create_engine('postgresql+psycopg2://energycosts:'+pwd+'@georgetownenergycosts.cr1legfnv0nf.us-east-1.rds.amazonaws.com:5432/energycosts')
# df = pd.read_sql('residential_adoption_Aug23',engine)
df = pd.read_csv('https://github.com/georgetown-analytics/Energy-Costs/blob/master/residential_adoption_Aug24.csv?raw=true')
df_deploy = pd.read_csv('https://github.com/georgetown-analytics/Energy-Costs/blob/master/zipcode_master_27Aug.csv?raw=true')
# -
df.dtypes
def reg_one_hot(data):
"""One-hot encode the region column - the only gategorical variable in the dataset"""
one_hot_region = pd.get_dummies(data['region'])
data = data.join(one_hot_region)
data.drop('region', axis=1,inplace=True)
return data
def backfiller(data,collist):
"""Define a function that will try to convert objects to numbers
and return a NaN when it can't (the errors='coerce' arg) and then backfill those NaNs
"""
for col in collist:
data[col] = pd.to_numeric(data[col],errors='coerce')
data[col].fillna(method='backfill',inplace=True)
# Clean the labeled data
# +
cols_to_clean = ['avg_hh_size','mean_income','mean_income_earning_hhs','earn_int_div_rent',
'percent_int_div_rent','earning_hhs','percet_1unit']
df = reg_one_hot(df)
backfiller(df,cols_to_clean)
df.head()
# -
# ... and the unlabeled data
# +
# run the same function on the unlabeled data
df_deploy = reg_one_hot(df_deploy)
backfiller(df_deploy,cols_to_clean)
df_deploy.head()
# -
# Convert the Category label to an integer and split the features and the targets from the df.
# +
def enumerate_classes(row): # this seems silly; there must be an easier way
if row == 'High':
return 1
else:
return 0
df['target_class'] = df['Category'].apply(enumerate_classes)
# -
print(list(enumerate(df.columns)))
df = df.sample(frac=1) # re-sample the data, with a sample fraction of 1, returning all the instances in random order. Not sure if train_test_split does this already
X = df.iloc[:,12:-1] # features
y = df['target_class'] # target
X.drop('ZCTA_5',axis=1,inplace=True)
# #### Feature Analysis
# I think population wants to go as a feature. There's a lot of colinerity between it and total households, which are both ingredients, though not directly, in our class calculation. They'd be dead give-aways. The capacity stuff is way less worrisome, though I bet the difference in winter/summer capacity will have a strong relationship with the weather data when we get that integrated
# +
from yellowbrick.features import Rank2D
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111)
visualizer = Rank2D(features=X.columns,algorithm='pearson',ax=ax)
visualizer.fit(X, y)
visualizer.transform(X)
visualizer.poof()
# -
X.drop('total_household',axis=1,inplace=True)
X.drop('population',axis=1,inplace=True)
# +
# seems like I'm experiencing the same issue as described here, but I'm not sure - moving on for now: https://github.com/DistrictDataLabs/yellowbrick/issues/402
# from yellowbrick.features import ParallelCoordinates
# visualizer = ParallelCoordinates(features=X.columns)
# visualizer.fit(X, y)
# visualizer.transform(X)
# visualizer.poof()
# -
# #### Scale the Data
# Our feautres use some wildly different scales (for example, number of installs vs. average annual income). We don't really care about the values themselves, but we do care about their relationship. The distribution of something like income might have a particularly long tail, so we'll use min_max instead of the standard scaler.
# the distribution of mean_income skews right, so we'll want to avoid using anything that relies on standard deviation for this
X['mean_income'].plot(kind='hist')
# +
# note that this will be sensitive to outliers, and there are quite a few in the dataset, particularly among features that describe earnings; I'm not sure this matters
# just yet. Mostly, I don't want to toss these out in case it turns out to be true that people at the very top end of the income spectrum tend to install more solar
scaler = MinMaxScaler()
X_scaled = scaler.fit_transform(X)
X_scaled = pd.DataFrame(X_scaled, columns=X.columns)
X_scaled.head()
# +
from yellowbrick.features.importances import FeatureImportances
from sklearn.ensemble import GradientBoostingClassifier
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot()
viz = FeatureImportances(GradientBoostingClassifier(), ax=ax)
viz.fit(X, y)
viz.poof('feature_importance.png')
# -
# #### Adjust for class imbalance
# Most of the zipcodes in the data are 'low' adoption. We'll create synthetic features using the Synthetic Minority Oversampling method (Adaptive Synthentic focuses, from what I can tell, on generating hard cases. I don't want any of those)
df['target_class'].value_counts().plot(kind='bar')
# +
# do this after train-test split
# from imblearn.over_sampling import SMOTE
# X_sample, y_sample = SMOTE().fit_sample(X_scaled,y)
# -
# #### Train the estimator
# Ensembles of weak learners tend to work best, but KNearestNeighbors gets an honorable mention. We'll leave it in and tune it in case giving a vote later on has a meaningful impact on our score.
# +
from sklearn import model_selection
from sklearn.model_selection import train_test_split
from sklearn.model_selection import cross_val_score
from imblearn.over_sampling import SMOTE
X_train, X_test, y_train, y_test = model_selection.train_test_split(X_scaled,y,test_size = 0.2)
X_train, y_train = SMOTE().fit_sample(X_train,y_train)
# -
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=7,weights='uniform',algorithm='auto')
knn_fitted = knn.fit(X_train, y_train)
knn_fitted.score(X_test,y_test)
from sklearn.ensemble import GradientBoostingClassifier
gbc = GradientBoostingClassifier(loss='exponential')
gbc_fitted = gbc.fit(X_train,y_train)
gbc_fitted.score(X_test, y_test)
# +
from yellowbrick.classifier import DiscriminationThreshold
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111)
visualizer = DiscriminationThreshold(model=gbc,ax=ax)
visualizer.fit(X_train,y_train)
visualizer.poof('discrimination_threshold.png')
# -
# #### Predict the unlabled data
# We'll have to get our unlabled data into the same shape as the training data, dropping a few columns and scaling the numerical features. It would probably have been best to do this through functions up front.
X_deploy = df_deploy.iloc[:,10:].dropna()
X_deploy.drop('ZCTA_5',axis=1,inplace=True)
X_deploy.drop('total_household',axis=1,inplace=True)
X_deploy.drop('population',axis=1,inplace=True)
X_deploy.drop('Territories',axis=1,inplace=True)
X_deploy_scaled = scaler.fit_transform(X_deploy)
X_deploy_scaled = pd.DataFrame(X_deploy_scaled, columns=X_deploy.columns)
X_deploy_scaled.head()
# Our top performing estimators have similar scores on the training data, which are only slightly improved by having them vote. RandomForests, however, produces a higher count in the medium and high classes. The economics of our problem don't really penalize false positives harshly (perhaps in terms of misspent Business Development efforts, but even this is an improvement from the current state), so we'll go with that classifier to predict our unlabled data.
deploy_predict = pd.DataFrame(knn_fitted.predict(X_deploy))
deploy_predict[0].value_counts()
# #### Join the predicted data with the labeled data for the full picture
# What follows is a Rube Goldberg process. First, we identify the list of zipcodes that are in our labeled data and in our predict data. Through this process, we learn that we somehow have 138 invalid zipcodes in our labeled data. We'll drop those, since their primary purpose of training our estimator has already been served.
#
# We'll create a new dataframe that has our existing zip codes and their labels, and we will join to this their LAT/LGN coordinates.
#
# We'll drop the list of labeled zipcodes from the master zipcode list. Then we'll append the labeled zipcodes to the predicted ones.
#
# This process aims to produce a mutually-exclusive, collectively-exhaustive (MECE) list of zipcodes, the coordinates of their centroids, and their actual or predicted classes.
all_zip = df_deploy.iloc[:,1:4]
all_zip = all_zip.join(deploy_predict)
all_zip.columns.values[3] = 'class'
all_zip.head()
# +
# coercing to sting - integer comparison produces strange results, whereas the output of this is correct.
oldzips = [str(z) for z in list(all_zip['ZIP']) if z in list(df['zipcode'])]
# +
# coverting zip to str to match above, and setting as the index so that df.drop can work on the row axis
all_zip['ZIP'] = all_zip['ZIP'].astype(str)
all_zip.set_index('ZIP', inplace=True)
new_zip = all_zip.drop(oldzips,axis=0)
# +
# grab the zipcodes and classes from the labeled data; coerce zipcode to string so all are in same dtype
zips_and_target = ['zipcode','target_class']
old_zips = pd.DataFrame(df[zips_and_target])
old_zips['zipcode'] = old_zips['zipcode'].astype(str)
# +
# left join master zipcode file to associate labeled zipcodes with their LAT/LNG centroids; drop predicted class
# since we have actual
old_zips = old_zips.join(all_zip,how='left',on='zipcode')
old_zips.drop('class',axis=1,inplace=True)
# +
# rename columns so that new_zip and old_zips match
old_zips.columns.values[1] = 'class'
old_zips.columns.values[0] = 'ZIP'
# +
# drop the 138 invalid zipcodes, and set the index of old_zips to zipcode so that column structure matches
old_zips.dropna(inplace=True)
old_zips = old_zips.set_index('ZIP')
# +
# produce MECE zipcode file for mapping
map_zips = old_zips.append(new_zip,sort=True,ignore_index=True)
map_zips.head()
# -
# #### Visualize the output
# We'll start with a one-circle-per zipcode map, which will give the user a sense of the location, number, and density of our predicted classes without giving zipcodes that are larger in area more prominence then they are due.
# +
alt.data_transformers.enable('default', max_rows=None)
#zipcodes = data.zipcodes.url
chart = alt.Chart(map_zips).mark_circle(size=6).encode(
alt.Color('class:N',
scale=alt.Scale(domain=['1', '2'],
range=['red','gray'])),
longitude='LNG:Q',
latitude='LAT:Q'
#color='class:N'
).project(
type='albersUsa'
).properties(
width=1250,
height=800
)
# -
chart.save('mastermap_binary.html')
| Machine Learning/energy_adoption_final-binary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Counts Analysis with Disorder Associations
#
# Co-occurence of terms analysis: check how often pre-selected cognitive terms appear in abstracts with ERP terms.
#
# This analysis searches through pubmed for papers that contain specified ERP and selected association terms.
#
# Data extracted is the count of the number of papers with both terms. This is used to infer the associated terms for each ERP.
#
# This notebook covers the disorder-related association terms.
# +
# %matplotlib inline
import numpy as np
from scipy.stats import normaltest, spearmanr
from lisc import Counts
from lisc.utils import SCDB, load_object
from lisc.utils.io import load_txt_file
from lisc.plts.counts import plot_matrix, plot_clustermap, plot_dendrogram
# -
import seaborn as sns
sns.set_context('talk')
# Import custom project code
import sys
sys.path.append('../code')
from plts import plot_count_hist, plot_time_associations, plot_latency_values
from analysis import get_time_associations
# ## Setup
# +
# Notebook settings
SAVE_FIG = False
SAVE_EXT = '.pdf'
# Set some plot settings for when saving out
# This is because changing these looks a bit odd in the notebook
matrix_linewidths = 0.35 if SAVE_FIG else 0
# -
# Analysis settings
N_ERPS = 150
# Set the file locations
term_dir = '../terms/'
figs_dir = '../data/figures/counts'
db = SCDB('../data/')
# Set the name of the file to load
name = 'disorders'
# Load the counts object
counts = load_object('counts_' + name, directory=db)
# ### Check Database Information
#
# Check the metadata about the data collection, including checking the database data were collected from.
# Check database information
counts.meta_data.db_info
# Check requester details
counts.meta_data.requester
# ## Collection Summaries
# ### ERP Articles
# Check the total number of association papers
print('The total # of ERP papers is \t\t {:.0f}'.format(sum(counts.terms['A'].counts)))
# Check the distribution of ERP papers
print('Test for normality (log-spaced) \t t-val: {:1.2f} \t p-val {:1.2f}'.format(\
*normaltest(np.log10(counts.terms['A'].counts))))
plot_count_hist(counts.terms['A'].counts, bins=12,
save_fig=SAVE_FIG, file_name='erp_hist' + SAVE_EXT, directory=figs_dir)
# ### Association Articles
# Check the total number of association papers
print('The total # of association papers is \t\t {:.0f}'.format(sum(counts.terms['B'].counts)))
# Check the distribution of association papers
print('Test for normality (log-spaced) \t t-val: {:1.2f} \t p-val {:1.2f}'.format(\
*normaltest(np.log10(counts.terms['B'].counts))))
plot_count_hist(counts.terms['B'].counts, bins=12,
save_fig=SAVE_FIG, file_name=name + '_assoc_hist' + SAVE_EXT, directory=figs_dir)
# ### Co-occurence Numbers
# Check how many co-occurence values are zero
n_coocs = np.multiply(*counts.counts.shape)
n_zero = sum(np.ravel(counts.counts) == 0)
percent_zero = (n_zero / n_coocs) * 100
# Print out completeness of the co-occurence matrix
print('Percent zero: \t\t% {:4.2f}'.format(percent_zero))
print('Percent non-zero: \t% {:4.2f}'.format(100 - percent_zero))
# Print out summaries of the co-occurence data
print('The total number of cooc values is: \t{:d}'.format(sum(np.ravel(counts.counts))))
print('The median number of cooc values is: \t{:2.2f}'.format(np.median(np.ravel(counts.counts))))
# Plot the distribution of (non-zero) co-occurence values
plot_count_hist(np.ravel(counts.counts), bins=12, log=True)
# # Check Counts
# Check the terms with the most papers
counts.check_top(dim='A')
counts.check_top(dim='B')
# Check how many papers were found for each ERP term
counts.check_counts(dim='A')
# Check how many papers were found for each association term
counts.check_counts(dim='B')
# Check the most commonly associated association term for each ERP
counts.check_data()
# Check the most commonly associated ERP for each term
counts.check_data(dim='B')
# ## Select ERPs with enough articles
# Check how many ERPs currently
counts.terms['A'].n_terms
# Drop ERPs without a target number of articles
counts.drop_data(N_ERPS, dim='A')
print(counts.terms['A'].n_terms)
# ## Group Level Plots
# Compute the normalized score (percent association)
counts.compute_score('normalize', dim='A')
# Plot the matrix of percent associations - ERPs & terms
plot_matrix(counts, linewidths=matrix_linewidths, figsize=(10, 8),
save_fig=SAVE_FIG, file_name=name + '_associations' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
# Plot a clustermap, clustering ERPs and terms based on similarity
plot_clustermap(counts, attribute='score', cmap='blue',
linewidths=matrix_linewidths, figsize=(12, 10),
save_fig=SAVE_FIG, file_name=name + '_clustermap' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
# ### Similarity Measure
# Calculate similarity between all ERPs (based on term association percents)
counts.compute_score('similarity')
# Plot similarity matrix between ERPs
plot_matrix(counts, linewidths=matrix_linewidths, figsize=(10, 6),
save_fig=SAVE_FIG, file_name=name + '_similarity' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
# Plot a clustermap, clustering ERPs and terms based on similarity
plot_clustermap(counts, attribute='score', cmap='blue',
linewidths=matrix_linewidths, figsize=(12, 10),
save_fig=SAVE_FIG, file_name=name + '_similarity_cluster' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
# ### Association Score
# Calculate association between all ERPs
counts.compute_score('association')
# Plot similarity matrix between terms
plot_matrix(counts, linewidths=matrix_linewidths, figsize=(10, 7),
save_fig=SAVE_FIG, file_name=name + '_associations' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
# ### Dendrograms
# Plot dendrogram of ERPs, based on percent associations with terms
plot_dendrogram(counts, attribute='score', figsize=(6, 8),
save_fig=SAVE_FIG, file_name=name + '_erp_dendro' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
# Plot dendrogram of termss, based on percent associations with ERPs
plot_dendrogram(counts, attribute='score', transpose=True, figsize=(6, 8),
save_fig=SAVE_FIG, file_name=name + '_term_dendro' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
# ## Component correlates across time
# Re-compute normalized score
counts = load_object('counts_' + name, directory=db)
counts.compute_score('normalize')
counts.drop_data(250)
print('Number of kept ERPs for this analysis: {}'.format(len(counts.terms['A'].labels)))
# Load canonical latency information
labels = load_txt_file('erp_labels.txt', term_dir, split_elements=False)
latencies = load_txt_file('latencies.txt', term_dir, split_elements=False)
latency_dict = {label : latency.split(', ') for label, latency in zip(labels, latencies)}
# ### Check the highest association across time
# Get the time and polarity information for the ERPs
time_associations = get_time_associations(counts, latency_dict)
# Set ERPs to drop from this analysis
exclude = ['P3b', 'MMN', 'FRN', 'MRCP', 'BP', 'LRP']
# Exclusion notes:
# - P3b dropped because P3a has same association (schizophrenia) at the same time
# - MMN dropped because N200 has the same association (schizophrenia) at the same time
# - FRN dropped because N2pc has the same association (anxiety) at the same time
# - MRCP, BP, LRP all dropped as preparatory activity (negative latency), all relating to motor
# Plot time associations
plot_time_associations(time_associations, exclude=exclude,
save_fig=SAVE_FIG, file_name=name + '_time' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
# ### Check average association value across time
# Reload the counts object, renormalize, and drop sparse components
counts = load_object('counts_' + name, directory=db)
counts.compute_score('normalize')
counts.drop_data(50)
print('Number of kept ERPs for this analysis: {}'.format(len(counts.terms['A'].labels)))
# Grab the association matrix values, sort and extract latencies
all_time_associations = get_time_associations(counts, latency_dict, 'all')
sorted_associations = sorted(all_time_associations, key=lambda x: x['latency'])
latencies = [el['latency'] for el in sorted_associations]
# Compute the average association value per component across time
avg_func = np.median
avgs = [avg_func(val['association']) for val in sorted_associations]
# Check the correlation between latency and average association score
print('Corr: {:2.4f}, \t p-val: {:2.4f}'.format(*spearmanr(latencies, avgs)))
# Plot the comparison between latency and average association score
plot_latency_values(latencies, avgs,
save_fig=SAVE_FIG, file_name=name + '_latency_corr' + SAVE_EXT,
directory=figs_dir, save_kwargs={'transparent' : True})
| notebooks/05-CountsDisorders.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Garimpagem de Dados
# ## Aula 1 - Python Básico
# ### Nulo
# Em Python, o tipo **None** representa a falta de valor em uma variável
a = None
if a is None:
print('nulo')
# ### Boolean
# Booleanos são escritos com a primeira letra maiúscula
a = True
b = False
c = True
print(a == b)
print(a == c)
# ### Números
# Em python, existem dois tipos básicos de números: Inteiros (int) e Ponto Flutuante (float)
print(type(1))
print(type(1.2))
# Ao dividir dois números (inteiros ou flutuantes), podemos fazer a divisão comum (/) ou divisão inteira (//), onde a última mantém apenas a parte inteira do resultado
print("Divisao: ",1 / 2)
print("Divisao inteira: ",1 // 2)
# Existe também o operador de potência, permitindo elevar um número ao outro.
# **PS**: como as raízes também são potências (porém, invertidas), podemos computá-las com o mesmo operador
print(2 ** 3)
print(25 ** (1/2))
# Python, assim como muitas linguagens, conta com um módulo que implementa diversas funções matemáticas (raíz, logaritmo, topo, piso, etc.
import math
print(math.sqrt(2))
print(math.log(2))
print(math.ceil(2 ** 0.5))
print(math.floor(2 ** 0.5))
# Python também conta um módulo de funções para operações randômicas
import random
print(random.random()) # normalizado entre 0 e 1
print(random.randint(1,10)) # intervalo fechado
# ### Strings
# Em Python, trabalhar com strings é bem simples, uma vez que já existem diversas funções implementadas.
#
# Para criar uma string em Python, basta colocar o conteúdo entre aspas, simples ou duplas (o tipo de aspas que começa a string é o mesmo tipo que irá finalizá-la)
single_quoted_string = 'Data Science'
print(single_quoted_string, type(single_quoted_string))
print(len(single_quoted_string)) # tamanho da string
print(single_quoted_string.upper()) # colocar caracteres em maiusculo
print(single_quoted_string.lower()) # colocar caracteres em minusculo
# Podemos facilmente substituir a ocorrência de um caractere/substring utilizando o método *replace*
str1 = 'Hoje_vai_chover_novamente.'.replace('_', ' ')
print(str1)
# Outra operação interessante de limpeza de strings é o *strip*, o qual remove os espaços em branco no começo e no final da string
print(' Hello '.strip()) # remove no começo e no final
print(' Hello '.lstrip()) # remove no começo
print(' Hello '.rstrip()) # remove no final
# Também podemos criar string mais longas, com múltiplas linhas, utilizando um total de 6 aspas (3 para abrir e 3 para fechar)
multi_line_string = """linha 1
linha 2
linha 3"""
print(multi_line_string)
print(repr(multi_line_string))
# Quando temos strings que contém múltiplas informações e desejamos separá-la, podemos usar a função *split*, informando o caractere/substring a ser utilizado para dividir a string
multi_line_string.split("\n")
# Fato interessante: strings em python podem ser manipuladas com operadores de soma (concatenação) ou multiplicação (repetição)
print("Big" + " " + "Data")
print('Repete ' * 5)
# Existem algumas maneiras de interpolar strings, ou seja, injetar valores dentro dela
# # %d e %f indicam, respectivamente, que um número inteiro ou ponto flutuante será inserido
print("%d/%f/%d" % (2, 4.5, 6))
print("%s/%s/%s" % ('a', 2, False)) # %s indica que algo será inserido como string
# Outra maneira de interpolar strings é usando o *format*
print("{}/{}/{}".format(2, 4, 6))
print("{:2d}/{:2d}/{:2d}".format(2, 4, 16))
print("{:02d}/{:02d}/{:02d}".format(2, 4, 160))
print('{:.2f}'.format(99.8765))
print('{:.0f}'.format(99.8765))
# ### Condições
#
# Estruturas condicionais em Python seguem a mesma lógica de diversas outras linguagens. Uma diferença é que, para os casos em que encadeamos várias condições, para não escrevermos *else if ...* temos o operador *elif*
valor = 99
if valor == 99:
print('veloz')
elif value > 200:
print('muito veloz')
else:
print('lento')
# #### Atribuição ternário
#
# Python também conta com atribuição ternária
x = 5
par_ou_impar = "par" if x % 2 == 0 else "impar"
print(par_ou_impar)
# ### Laços / Loops
# Em Python, a estrutura de repetição *while* funciona de maneira análoga a diversas outras linguagens:
x = 0
while x < 5:
print(x)
x += 1
# Entretanto, o *for* funciona iterando sobre uma coleção/sequência de valores. Existem diversas maneiras de fazer isso:
# +
# definindo um intervalo
for i in range(10): # de 0 a 9, pulando de 1 em 1
print(i, end= " ")
print()
for i in range(5,10): # de 5 a 9, pulando de 1 em 1
print(i, end= " ")
print()
for i in range(0,10,2): # de 0 a 9, pulando de 2 em 2
print(i, end= " ")
print()
# -
# utilizando uma sequência pre-definida
a = [1, 3, 4, 5, 7]
for i in a:
print(i)
# a função enumerate retorna uma lista de pares (i,x), onde:
# i é o índice (0, 1, 2, ...)
# x é o elemento da sequência original
a = [1, 3, 4, 5, 7]
for indice, valor in enumerate(a):
print(indice, valor)
# podemos também percorrer uma coleção mas não utilizar o valor daquele passo
for _ in range(5):
print('oi')
# existem também dois operadores dentro de um for:
# continue - encerra o passo atual e passa para o próximo
# break - encerra o laço
for x in range(10):
if x == 3:
continue
if x == 5:
break
print(x)
# ### Funções
#
# Para definir uma função em python, basta utilizarmos a seguinte estrutua:
#
# ```
# def nome_da_função (argumento1, argumento2, ...):
# corpo da função
# ```
# +
def soma(a, b):
return a + b
print(soma(3, 5))
print(soma('casa ', 'organizada'))
# -
# #### Manipulando Funções
#
# Python pode ser considerada uma linguagem **funcional**. Uma das características do paradigma funcional é permitir que uma função seja manipulada como um valor ou variável, ou seja, podemos passar funções como argumentos para outras funções
# +
def f1():
print("Function 1")
def f2():
print("Function 2")
def chamar_funcao(f):
f()
chamar_funcao(f1)
chamar_funcao(f2)
# -
# Existem também as funções anônimas (chamadas, em Python, de **lambda**). Uma função *anônima* é uma função que contém apenas a assinatura e o corpo, sem um nome definido. Em Python, elas recebem o nome de *lambda* pois elas são funções de um comando só (diferente de linguagens como Javascript e Scala, que permitem funções anônimas de múltiplas linhas).
chamar_funcao(lambda : print("Lambda function"))
# ### Data
# Em Python, podemos manipular informações de data, hora e tempo utilizando o módulo *datetime*
# +
from datetime import datetime
import time
inicio = datetime.now() # obtém o timestamp do momento atual
print(inicio)
for i in range(2):
time.sleep(1) # aplica um delay, em segundos
fim = datetime.now()
print(fim)
print("Tempo de execução: {}".format(fim - inicio))
# -
# ### Listas
#
# Quando desejamos trabalhar com uma sequência de dados, podemos utilizar uma lista
lista = [1, 2, 3, 4, 5]
print(lista)
# Existem diversas maneiras de acessar os elementos de uma lista:
lista[2] #elemento da lista
lista[-1] # último elemento
lista[1:3] #sublista
lista[:3] # 3 primeiros elementos
lista[-2:] # 2 últimos elementos
lista[1:-1] # do segundo ao penúltimo elemento
# O operador *len* pode ser utilizado para medir tamanhos de objetos com múltiplas informações (strings, lista, etc.)
len(lista)
# Podemos somar elementos de uma lista simplesmente usando a função *sum*
sum(lista)
# Podemos também consultar a pertinência de um elemento em uma lista utilizando o operador *in*
print(10 in lista)
print(5 in lista)
# Em Python, as listas são **mutáveis**, ou seja, ela pode ser modificada.
# +
lista.append(7) # adiciona elemento na lista
lista.append(6)
lista.append(7)
print(lista)
lista.remove(7) # remove o primeiro elemento cujo valor é 7
print(lista)
lista.extend([8, 9, -10]) #extende a lista com outra lista
lista
# -
# Por padrão, as variáveis em Python (quando apontando para tipos de dados mais complexos) são **ponteiros**, ou seja, elas são uma referência a um dado alocado na memória. Logo, quando salvamos o valor de uma variável A em uma variável B, estamos apenas dizendo que B **aponta** para a mesma informação que A
lista2 = lista # lista2 referencia lista
lista2[-1] = 10
print(lista)
print(lista2)
# Embora a lista seja mutável, as operações de acesso aos elementos (com o operador \[ \]) retornam *novos* elementos. Logo, podemos fazer uma cópia de uma lista da seguinte maneira
lista3 = lista[:] # lista3 é uma nova lista com o mesmo conteúdo de lista
lista3[-1] = -10
print(lista)
print(lista3)
# Existem duas maneiras de se ordenar uma lista em Python: uma abordagem mutável (que altera a lista original) e uma imutável (que retorna uma nova lista ordenada)
# +
lista4 = [-4, 1, -2, 3]
print(sorted(lista4)) # ordena sem alterar a lista
print(sorted(lista4, reverse=True))
print(sorted(lista4, key=abs))
print(lista4)
lista4.sort() # altera a lista original
print(lista4)
# -
# O módulo **random** contém também um conjunto de funções para trabalharmos com listas. Podemos embaralhá-las, escolher um elemento aleatório ou obter uma amostra de tamanho *n*
# +
import random
a = ['a', 'casa', 'está', 'muito', 'bem', 'organizada']
random.shuffle(a) # mutável
print(a)
random.shuffle(a)
print(a)
random.choice(a) # com repetição
numeros = range(1, 100)
random.sample(numeros, 5) # sem repetição
# -
# A representação padrão de lista em string é uma sequência das strings de seu conteúdo, separado por ', ' e cercados por colchetes. O método **join**, de strings, existe para usar uma string como *separador* de uma lista, conforme o exemplo abaixo
print(a)
print(' '.join(a))
# **OBS**: se um elemento da lista **não for uma string**, o método *join* não funciona
b = [1, 'a', 25, 'abc']
print(b)
print(' '.join(b))
# ### Operações funcionais sobre listas
# Existem algumas funções primitivas para operar sobre os elementos de uma lista. Duas delas são **transformação** (*map*) e filtragem (*filter*).
# #### Transformação
#
# O objetivo desta função é, como o próprio nome sugere, *transformar* os elementos de uma lista, gerando uma nova lista. Em termos mais matemáticos: dada uma função *f(x)* definida para todos os elementos de uma lista *L* [a1, a2, ..., an], será gerada uma nova lista *L2* [b1, b2, ..., bn] tal que *bi = f(ai)*.
#
# Para realizar uma trnasformação, utilizamos a função *map*
# +
a = [1, 2, 3, 4, 5]
doubled_a = map(lambda x: 2*x, a)
doubled_a
# -
# O retorno do *map* é *iterador*, ou seja, os seus valores só existirão qdo forem chamados (seja em um *for*, outro *map* ou fazendo um *cast* para lista) e, uma vez que forem consumidos, o *iterador* fica vazio
# +
#descomente uma alternativa para checar seu funcionamento
doubled_a = map(lambda x: 2*x, a)
# for x in doubled_a:
# print(x, end=" ")
# str_doubled_a = map(lambda x: str(x), doubled_a)
# print(" ".join(str_doubled_a))
# list(doubled_a)
# -
# #### Filtragem
#
# Obviedades a parte, o objetivo dessa operação é gerar uma nova coleção apenas com os elementos que satisfaçam uma condição. De maneira mais matemática: dado um predicado *p(x)* definido para todos os elementos de uma lista *L* [a1, a2, ..., an], será gerada uma nova lista *L2* [b1, b2, ..., bm] tal que *bi $\in$ L* e *p(bi)* é verdade.
even_a = filter(lambda x: x%2 == 0, a)
even_a
# Novamente temos um *iterator* como retorno. Podemos utilizar as mesmas abordagens do *iteratoor* apresentado no *map*
# +
#descomente uma alternativa para checar seu funcionamento
even_a = filter(lambda x: x%2 == 0, a)
# for x in even_a:
# print(x, end=" ")
# str_even_a = map(lambda x: str(x), even_a)
# print(" ".join(str_even_a))
# list(even_a)
# -
# #### Combinando transformação e filtragem: Compressão de Listas
#
# Uma maneira prática para combinar as operações de transformação e filtragem é utilizar uma *compressão de lista*. A anatomia da compressão de lista é a seguinte:
#
# [f(x) for x in lista if p(x)]
# +
quadrados = [x * x for x in range(5)]
print(quadrados)
quadrados_dos_pares = [x * x for x in range(5) if x%2 == 0]
print(quadrados_dos_pares)
# -
# ### Tuplas
#
# Tuplas são grupos ordenados de valores. Esse par é *imutável* e normalmente é utilizado para representar informações estruturadas (linha de um *csv*, tupla de um banco de dados, etc)
tupla = (1, 2, 3, 4)
print(tupla)
# ### Dicionários
#
# Dicionários são pares de chave : valor, utilizados para representar um dado cujos valores tem uma semântica atribuída a ela.
dicionario = { 'nome': 'Fulano', 'sobrenome': '<NAME>' }
print(dicionario['nome'])
print(dicionario['sobrenome'])
# Em Python, podemos acessar, separadamente, suas chaves e seus valores
print(dicionario.keys())
print(dicionario.values())
| 2020/01-python-jupyter-notebook/Python-101.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + papermill={"duration": 0.038668, "end_time": "2020-12-07T00:46:21.357405", "exception": false, "start_time": "2020-12-07T00:46:21.318737", "status": "completed"} tags=[]
RUN_TEST = True
# + papermill={"duration": 0.038004, "end_time": "2020-12-07T00:46:21.424863", "exception": false, "start_time": "2020-12-07T00:46:21.386859", "status": "completed"} tags=[]
from time import time
start_time = time()
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 10.630543, "end_time": "2020-12-07T00:46:32.086911", "exception": false, "start_time": "2020-12-07T00:46:21.456368", "status": "completed"} tags=[]
import tensorflow as tf
import transformers
print(tf.__version__)
print(transformers.__version__)
# + _cell_guid="79c7e3d0-c299-4dcb-8224-4455121ee9b0" _uuid="d629ff2d2480ee46fbb7e2d37f6b5fab8052498a" papermill={"duration": 4.35631, "end_time": "2020-12-07T00:46:36.475209", "exception": false, "start_time": "2020-12-07T00:46:32.118899", "status": "completed"} tags=[]
# Detect hardware, return appropriate distribution strategy
try:
tpu = tf.distribute.cluster_resolver.TPUClusterResolver() # TPU detection
print('Running on TPU ', tpu.cluster_spec().as_dict()['worker'])
except ValueError:
tpu = None
gpus = tf.config.experimental.list_logical_devices("GPU")
if tpu:
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu)
elif len(gpus) > 1: # multiple GPUs in one VM
strategy = tf.distribute.MirroredStrategy(gpus)
else: # default strategy that works on CPU and single GPU
strategy = tf.distribute.get_strategy()
print("REPLICAS: ", strategy.num_replicas_in_sync)
# + papermill={"duration": 0.047044, "end_time": "2020-12-07T00:46:36.555856", "exception": false, "start_time": "2020-12-07T00:46:36.508812", "status": "completed"} tags=[]
PUNCT_SET = set("#《》【】[]") # 保留这些预定义的标点
def is_chinese(uchar: str) -> bool:
# 暂时保留以下字符,看看CV是否提高
if uchar in PUNCT_SET:
return True
if uchar >= '\u4e00' and uchar <= '\u9fa5':
return True
else:
return False
def reserve_chinese(content: str, threshold: int = 512) -> str:
content_str = ''
c = 0
for i in content:
if c == threshold:
break
if is_chinese(i):
content_str += i
c += 1
return content_str
# + papermill={"duration": 1.326057, "end_time": "2020-12-07T00:46:37.926628", "exception": false, "start_time": "2020-12-07T00:46:36.600571", "status": "completed"} tags=[]
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import re
import os
import pickle
from tqdm.notebook import tqdm
import seaborn as sns
import plotly.express as px
import matplotlib.pyplot as plt
# %matplotlib inline
from sklearn.metrics import *
from sklearn.model_selection import StratifiedKFold
from sklearn.model_selection import train_test_split
from sklearn.utils import shuffle
# TENSORFLOW
from tensorflow.keras.layers import Dense, Input, Dropout
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.models import Model
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.utils import to_categorical
from tensorflow.keras.layers import Dense, Input, GlobalAveragePooling1D, GlobalMaxPooling1D
# HUGGINGFACE
from tokenizers import BertWordPieceTokenizer
from transformers import TFAutoModel, AutoTokenizer, TFBertModel
from transformers import TFBertForSequenceClassification, TFTrainer, TFTrainingArguments
from transformers import AdamWeightDecay
# + papermill={"duration": 0.049373, "end_time": "2020-12-07T00:46:38.009038", "exception": false, "start_time": "2020-12-07T00:46:37.959665", "status": "completed"} tags=[]
AUTO = tf.data.experimental.AUTOTUNE
# Configuration
EPOCHS = 2
N_FOLDS = 10
BATCH_SIZE = 32 * strategy.num_replicas_in_sync
MAX_LEN = 192
NUM_AUG = 8
MODEL_NAME = 'bert-base-chinese'
# + papermill={"duration": 2.28214, "end_time": "2020-12-07T00:46:40.472328", "exception": false, "start_time": "2020-12-07T00:46:38.190188", "status": "completed"} tags=[]
tokenizer = AutoTokenizer.from_pretrained(MODEL_NAME)
# + papermill={"duration": 0.04794, "end_time": "2020-12-07T00:46:40.555561", "exception": false, "start_time": "2020-12-07T00:46:40.507621", "status": "completed"} tags=[]
labels = ['文化休闲', '医疗卫生', '教育科技', '城乡建设', '工业', '交通运输', '生态环境', '经济管理',
'政法监察', '农业畜牧业', '文秘行政', '劳动人事', '信息产业', '民政社区', '旅游服务', '商业贸易',
'气象水文测绘地震地理', '资源能源', '财税金融', '外交外事']
label_map, inv_label_map = {}, {}
for idx, label in enumerate(labels):
label_map[label] = idx
inv_label_map[idx] = label
# + papermill={"duration": 65.255362, "end_time": "2020-12-07T00:47:45.847979", "exception": false, "start_time": "2020-12-07T00:46:40.592617", "status": "completed"} tags=[]
train_df_aug = pd.read_csv("/kaggle/input/onecity/train_df_processed_1206_aug_5_chinese.csv")
text_df = pd.DataFrame(train_df_aug['text'].apply(eval).to_list(), columns=[f'text{i}' for i in range(1, NUM_AUG+1)])
text_df = text_df.fillna("").astype(str)
for col in [f'text{i}' for i in range(1, NUM_AUG+1)]:
text_df[col] = text_df[col].apply(lambda x: "" if x == 'ERROR' else x.lower())
text_df[col] = text_df[col].apply(reserve_chinese)
text_df[col] = text_df[col].apply(lambda x: x[:MAX_LEN-2])
text_df['filename'] = train_df_aug['filename']
text_df['label'] = train_df_aug['label']
text_df = text_df[text_df.text1 != '无访问权限']
# + papermill={"duration": 0.870752, "end_time": "2020-12-07T00:47:46.915228", "exception": false, "start_time": "2020-12-07T00:47:46.044476", "status": "completed"} tags=[]
df_text_counts = text_df['text1'].value_counts()
top_freq_texts = set(df_text_counts[df_text_counts > 200].index)
df_sub = text_df[text_df['text1'].apply(lambda x: x not in top_freq_texts)]
print(df_sub.shape)
for text in list(top_freq_texts):
df_sub2 = text_df[text_df['text1'] == text].head(20)
df_sub = pd.concat([df_sub, df_sub2])
print(df_sub.shape)
train_df = df_sub.reset_index(drop=True)
train_df = train_df.sample(frac=1., random_state=2020)
train_df = train_df.reset_index(drop=True)
# + papermill={"duration": 1.055043, "end_time": "2020-12-07T00:47:48.006641", "exception": false, "start_time": "2020-12-07T00:47:46.951598", "status": "completed"} tags=[]
if RUN_TEST:
test_df = pd.read_csv("/kaggle/input/onecity/rest_df_content_only_1206_chinese.csv")
# + papermill={"duration": 464.739998, "end_time": "2020-12-07T00:55:32.785116", "exception": false, "start_time": "2020-12-07T00:47:48.045118", "status": "completed"} tags=[]
# %%time
x_train = []
for idx in range(1, NUM_AUG+1):
col = rf"text{idx}"
print(f"Encode Train: Part {idx}...")
train_aug_encoded = tokenizer.batch_encode_plus(
train_df[col].values,
pad_to_max_length=True,
max_length=MAX_LEN
)
x_train.append(np.array(train_aug_encoded['input_ids']))
y = train_df['label'].map(label_map).values
# + papermill={"duration": 0.426332, "end_time": "2020-12-07T00:55:33.346847", "exception": false, "start_time": "2020-12-07T00:55:32.920515", "status": "completed"} tags=[]
test_df = test_df.fillna("").astype(str)
for col in [f'text{i}' for i in range(1, NUM_AUG+1)]:
test_df[col] = test_df[col].apply(lambda x: "" if x == 'ERROR' else x.lower())
test_df[col] = test_df[col].apply(lambda x: x[:MAX_LEN-2])
# + papermill={"duration": 145.102106, "end_time": "2020-12-07T00:57:58.490993", "exception": false, "start_time": "2020-12-07T00:55:33.388887", "status": "completed"} tags=[]
# %%time
if RUN_TEST:
test_datasets = []
for idx in range(1, NUM_AUG+1):
col = rf"text{idx}"
print(f"Encode Test: Part {idx}...")
test_aug_encoded = tokenizer.batch_encode_plus(
test_df[col].values,
pad_to_max_length=True,
max_length=MAX_LEN
)
x_test = np.array(test_aug_encoded['input_ids'])
test_dataset = (
tf.data.Dataset
.from_tensor_slices(x_test)
.batch(BATCH_SIZE)
)
test_datasets.append(test_dataset)
# + papermill={"duration": 0.055667, "end_time": "2020-12-07T00:57:58.590964", "exception": false, "start_time": "2020-12-07T00:57:58.535297", "status": "completed"} tags=[]
len(x_train[0]), len(y)
# + papermill={"duration": 0.059288, "end_time": "2020-12-07T00:57:58.696195", "exception": false, "start_time": "2020-12-07T00:57:58.636907", "status": "completed"} tags=[]
def build_model(model_name, max_len):
# First load the transformer layer
if MODEL_NAME == 'bert-base-chinese':
transformer_encoder = TFAutoModel.from_pretrained(model_name)
else:
transformer_encoder = TFBertModel.from_pretrained(model_name, from_pt=True)
# This will be the input tokens
input_ids = Input(shape=(max_len,), dtype=tf.int32, name="input_ids")
# Now, we encode the text using the transformers we just loaded
sequence_output = transformer_encoder(input_ids)[0]
# Only extract the token used for classification, which is <s>
cls_token = sequence_output[:, 0, :]
# Finally, pass it through a 3-way softmax, since there's 3 possible laels
out = Dense(20, activation='softmax')(cls_token)
# It's time to build and compile the model
model = Model(inputs=input_ids, outputs=out)
model.compile(
Adam(lr=3e-5),
loss='sparse_categorical_crossentropy',
metrics=['accuracy']
)
return model
# + papermill={"duration": 0.053924, "end_time": "2020-12-07T00:57:58.794992", "exception": false, "start_time": "2020-12-07T00:57:58.741068", "status": "completed"} tags=[]
kfold = StratifiedKFold(n_splits=N_FOLDS)
# + papermill={"duration": 0.056946, "end_time": "2020-12-07T00:57:58.898765", "exception": false, "start_time": "2020-12-07T00:57:58.841819", "status": "completed"} tags=[]
train_df['y_pred'] = ""
train_df['proba'] = 0.0
# + papermill={"duration": 5494.800551, "end_time": "2020-12-07T02:29:33.745002", "exception": false, "start_time": "2020-12-07T00:57:58.944451", "status": "completed"} tags=[]
all_accs = []
test_pred_results = []
for ii, (tr, tt) in enumerate(kfold.split(X=y, y=y)):
if tpu:
tf.config.experimental_connect_to_cluster(tpu)
tf.tpu.experimental.initialize_tpu_system(tpu)
strategy = tf.distribute.experimental.TPUStrategy(tpu)
elif len(gpus) > 1: # multiple GPUs in one VM
strategy = tf.distribute.MirroredStrategy(gpus)
else: # default strategy that works on CPU and single GPU
strategy = tf.distribute.get_strategy()
# Prepare KFold data
y_train, y_valid = y[tr], y[tt]
x_train_combined = np.concatenate([x[tr] for x in x_train])
y_train_combined = np.concatenate([y_train] * len(x_train))
# Shuffle augmented data
idxs = np.arange(len(y_train_combined))
idxs = shuffle(idxs, random_state=2020)
x_train_combined = x_train_combined[idxs]
y_train_combined = y_train_combined[idxs]
train_dataset = (
tf.data.Dataset
.from_tensor_slices((x_train_combined, y_train_combined))
.repeat()
.shuffle(2048)
.batch(BATCH_SIZE)
.prefetch(AUTO)
)
valid_datasets = []
for x in x_train:
valid_dataset = (
tf.data.Dataset
.from_tensor_slices((x[tt], y_valid))
.batch(BATCH_SIZE)
.cache()
.prefetch(AUTO)
)
valid_datasets.append(valid_dataset)
with strategy.scope():
model = build_model(MODEL_NAME, MAX_LEN)
n_steps = len(x_train_combined) // BATCH_SIZE
train_history = model.fit(
train_dataset,
steps_per_epoch=n_steps,
validation_data=valid_datasets[0],
epochs=EPOCHS
)
# Predict on validation set
valid_aug_probs = [model.predict(valid_dataset, verbose=1) for valid_dataset in valid_datasets]
valid_probs = np.mean(valid_aug_probs, axis=0)
y_valid_preds = np.argmax(valid_probs, axis=1)
acc = accuracy_score(y_valid, y_valid_preds)
print(f"Accuracy for KFold {ii}: {acc}")
all_accs.append(acc)
train_df.loc[tt, 'y_pred'] = np.vectorize(inv_label_map.get)(y_valid_preds)
train_df.loc[tt, 'proba'] = valid_probs.max(axis=1)
if RUN_TEST:
# Prediction on test set
test_aug_probs = [model.predict(test_dataset, verbose=1) for test_dataset in test_datasets]
test_probs = np.mean(test_aug_probs, axis=0)
test_pred_results.append(test_probs)
# + papermill={"duration": 6.994657, "end_time": "2020-12-07T02:29:47.758782", "exception": false, "start_time": "2020-12-07T02:29:40.764125", "status": "completed"} tags=[]
# test_aug_encoded = tokenizer.batch_encode_plus(
# # ['工作单位新办序号广饶县环卫处延华文男华泰集团有限公司杜滨男华泰集团有限公司倪鹤女广饶县丰源纺织有限公司燕荣凤女原广饶县供销贸易公司宋福志男广饶县山水水泥有限公司王光诚男花官镇洛程幼儿园王芬女科达集团高孟海男广饶科力达石化科技有限公司田美岗男山东华星石油化工集团有限公司谢文杰男华泰集团有限公司傅建武男原服装厂崔向亮男'],
# # ['出生年月性别青岛市市北区台东三路号单元户刘宽男青岛市市北区顺兴路号户臧丽娜女青岛市市北区东光路号单元宋降龙男青岛市市北区华阳路号大成公馆号楼户王新男青岛市埕口一路三单元户王英光男青岛市市北区瑞海北路号号楼户瑞海馨园肖中权男青岛市市北区台东八路号户周嵩智男山东省青岛市市北区东仲小区号单元户李龙男青岛市市北区长春路东兴市场号号楼单元乔安钢男青岛市市北区台东六路号户鲍习平男山东省莱阳市穴坊镇西富山村孙辉女黄岛路号户蒲英玲女青岛市市北区康宁路号北舍号楼单元户张瑛女青岛市徐州路号号楼单元室苏兆楷男青岛市市北区台东三路号单元户关永斌男无棣县金羚华府号楼一单元室程兵男青岛市市北区标山路号户王玉台男青岛市市北区威海路号户段京兵男山东省青岛市市北区台东七路号楼户梁尧庆男青岛市市北区芙蓉路号号楼单元户连红男通化路号单元户姜腾飞男'],
# ['出生年月民族参加工作时间性别汉大本男张宪印党组成员副局长山东政法干部管理学院汉大专男高峰科员河北燕山大学汉大本女张双双主任山东省委党校汉大本男陈尚平党组副书记局长莱阳农学院汉硕士男胡金庆科员中国海洋大学汉大本男刘勇党组书记副局长山东工业大学汉大本女段琪琪科员长江大学汉初中男顾兆强科员北长山联中汉大专男高峰主任河北燕山大学汉初中男董仁科员蓬莱市大季家中学汉大本男李强副主任山东省委党校汉中专男娄兆军科员烟台水校汉大专男刘玉伟党组成员副局长烟台师范学院汉初中男于庆科员龙口市大王中学汉大本男刘勇党组书记副局长山东工业大学汉大本男李强科员山东省委党校汉大本女隋婷婷科员山东农业大学汉大本女刘宗云科员山东科技大学汉中专男娄兆军科员烟台水校汉大本女刘宗云科员山东科技大学汉大本男吴忠进科员山东函授大学汉初中男董仁科员蓬莱市大季家中学汉初中男于庆副主任龙口市大王中学汉硕士男胡金庆科员中国海洋大学汉大本女张双双主任山东省委党校汉大本女于咏文科员山东省委党校汉大本女隋婷婷科员山东农业大学汉大本男霍延虎科员山东理工大学汉大本女魏童童科员济宁学院汉大本女段琪琪科员长江大学汉初中男顾兆强科员北长山联中汉大本男王海亮党组成员山东省委党校汉大专男刘玉伟党组成员副局长烟台师范学院汉大本女于咏文科员山东省委党校汉大本女王美丁副主任聊城大学汉大本女乔婕科员鲁东大学汉大本男王黎明副主任长岛县委党校汉大本女乔婕科员鲁东大学汉大本男霍延虎科员山东理工大学汉大本女王美丁副主任聊城大学汉大本男王海亮党组成员山东省委党校汉大本男张宪印党组成员副局长山东政法干部管理学院汉大本男王黎明科员长岛县委党校汉大本女魏童童科员济宁学院汉大本男吴忠进科员山东函授大学汉大本男陈尚平党组副书记局长莱阳农学院'],
# pad_to_max_length=True,
# max_length=MAX_LEN
# )
# x_test = np.array(test_aug_encoded['input_ids'])
# test_dataset = (
# tf.data.Dataset
# .from_tensor_slices(x_test)
# .batch(BATCH_SIZE)
# )
# probs = model.predict(test_dataset, verbose=1)
# + papermill={"duration": 6.93249, "end_time": "2020-12-07T02:30:01.677521", "exception": false, "start_time": "2020-12-07T02:29:54.745031", "status": "completed"} tags=[]
# plt.plot(probs[0])
# + papermill={"duration": 6.985826, "end_time": "2020-12-07T02:30:15.473202", "exception": false, "start_time": "2020-12-07T02:30:08.487376", "status": "completed"} tags=[]
# inv_label_map[np.argmax(probs[0])]
# + papermill={"duration": 6.771855, "end_time": "2020-12-07T02:30:29.160716", "exception": false, "start_time": "2020-12-07T02:30:22.388861", "status": "completed"} tags=[]
# for probs in valid_aug_probs:
# _pred = np.argmax(probs, axis=1)
# print(accuracy_score(y_valid, _pred))
# + papermill={"duration": 7.031422, "end_time": "2020-12-07T02:30:43.264826", "exception": false, "start_time": "2020-12-07T02:30:36.233404", "status": "completed"} tags=[]
# for idx in range(2, len(valid_aug_probs)):
# _probs = np.mean(valid_aug_probs[:idx], axis=0)
# _pred = np.argmax(_probs, axis=1)
# print(accuracy_score(y_valid, _pred))
# + papermill={"duration": 6.965447, "end_time": "2020-12-07T02:30:57.079242", "exception": false, "start_time": "2020-12-07T02:30:50.113795", "status": "completed"} tags=[]
# for idx in range(2, len(valid_aug_probs)):
# _probs = np.mean(valid_aug_probs[-idx:], axis=0)
# _pred = np.argmax(_probs, axis=1)
# print(accuracy_score(y_valid, _pred))
# + papermill={"duration": 6.873689, "end_time": "2020-12-07T02:31:11.031149", "exception": false, "start_time": "2020-12-07T02:31:04.157460", "status": "completed"} tags=[]
# np.mean(valid_aug_probs[:3], axis=0)
# + papermill={"duration": 6.936902, "end_time": "2020-12-07T02:31:24.888096", "exception": false, "start_time": "2020-12-07T02:31:17.951194", "status": "completed"} tags=[]
print(all_accs)
print(np.mean(all_accs))
# + papermill={"duration": 7.246427, "end_time": "2020-12-07T02:31:38.962445", "exception": false, "start_time": "2020-12-07T02:31:31.716018", "status": "completed"} tags=[]
test_preds = np.argmax(np.sum(test_pred_results, axis=0), axis=-1)
test_df['label'] = np.vectorize(inv_label_map.get)(test_preds)
test_df[['filename', 'label']].to_csv("content_only_prediction.csv", index=False, encoding='utf-8')
# + papermill={"duration": 6.869928, "end_time": "2020-12-07T02:31:52.731081", "exception": false, "start_time": "2020-12-07T02:31:45.861153", "status": "completed"} tags=[]
with open("test_probs.pkl", 'wb') as f:
pickle.dump(test_pred_results, f)
# + papermill={"duration": 8.895686, "end_time": "2020-12-07T02:32:08.470093", "exception": false, "start_time": "2020-12-07T02:31:59.574407", "status": "completed"} tags=[]
train_df.to_csv("train_error_analysis.csv", index=False)
# + papermill={"duration": 6.925227, "end_time": "2020-12-07T02:32:22.203346", "exception": false, "start_time": "2020-12-07T02:32:15.278119", "status": "completed"} tags=[]
print(f"Total Running Time: {time() - start_time:.3f} seconds")
| Bert_content_only.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="cPdDT4EfNu8P"
# <font color="steelblue">To use this notebook on Colaboratory, you will need to make a copy of it. Go to File > Save a Copy in Drive. You can then use the new copy that will appear in a seperate tab.</font>
#
# + [markdown] colab_type="text" id="dqhVm_8kN2tl"
# # Practice Notebook: Python for Data Science - Data Types
# + [markdown] colab_type="text" id="qLLe0H-WN7az"
# ## 1. Data Types
# + [markdown] colab_type="text" id="VtTVm3cvN_lP"
# #### <font color="blue">Examples</font>
# + [markdown] colab_type="text" id="qIxhQwTHOIjm"
# ##### <font color="blue">Example 1
# + colab={} colab_type="code" id="n7APi3z1NtN0"
# Example 1
# ---
# Dynamically-inferred types
# ---
#
x = 20
print(type(x))
x = '20'
print(type(x))
x = 20.0
print(type(x))
# + [markdown] colab_type="text" id="JdKpybEFOJOM"
# ##### <font color="blue">Example 2
# + colab={} colab_type="code" id="2Al_wNeZOE7t"
# Example 2
# ---
# Manual type-conversion (string to int)
# ---
#
x = 20
y = '5'
print(x + int(y))
# + [markdown] colab_type="text" id="XJoETlR_OKHt"
# ##### <font color="blue">Example 3
# + colab={} colab_type="code" id="CmPAy-efOEx-"
# Example 3
# ---
# Automatic type-conversion (int to float)
# ---
#
x = 20
print(type(x))
x += 5.0
print(x), type(x)
# + [markdown] colab_type="text" id="8w3U0HVEOccy"
# ##### <font color="blue">Example 4
# + colab={} colab_type="code" id="o_qQW55lOeY5"
# Example 4
# ---
# Dividing Integers
# ---
#
a = 20
b = 5
print(a/b)
print(b/a)
# + [markdown] colab_type="text" id="hDZ9eDtmOgLi"
# ##### <font color="blue">Example 5
# + colab={} colab_type="code" id="RqtHFRB1OkU-"
# Example 5
# ---
# Forcing float division
# ---
#
print(b/float('5'))
# How can you correct this?
# + [markdown] colab_type="text" id="SxIaU06sOuYQ"
# ##### <font color="blue">Example 6
# + colab={} colab_type="code" id="Ho7QuNWqOvNs"
# Examaple 6
# ---
# String "arithmetic" (actually concatenation)
# ---
a = 'John '
b = 'Doe'
print(a + b)
# + [markdown] colab_type="text" id="GuR57OHgOBnM"
# #### <font color="green">Challenges</font>
# + [markdown] colab_type="text" id="eaq4UrFLONmb"
# ##### <font color="green">Challenge 1</font>
# + colab={} colab_type="code" id="<KEY>"
# Challenge 1
# ---
# Question: Concatenate and print your full names.
# ---
#
#OUR CODE GOES HERE
a = 'Jessica'
b = 'Wolfe'
a + b
# + [markdown] colab_type="text" id="bSurXHq_OOjc"
# ##### <font color="green">Challenge 2</font>
# + colab={} colab_type="code" id="fQjDSfP1OEPH"
# Challenge 2
# ---
# Question: Run and correct the following code.
# ---
#
x = 10
y = '5'
print(x + int(y))
# + [markdown] colab_type="text" id="hnyRRASVOPq4"
# ##### <font color="green">Challenge 3</font>
# + colab={} colab_type="code" id="7qUmDv5NOFl0"
# Challenge 3
# ---
# Question: Perform integer division of 199 and 3 with the result being a float.
# ---
#
#OUR CODE GOES HERE
a = 199
b = 3
c = (a/(float(b)))
c
# + [markdown] colab_type="text" id="okwTjEZ_PCiN"
# ##### <font color="green">Challenge 4</font>
# + colab={} colab_type="code" id="YThUHgEzPEfP"
# Challenge 4
# ---
# Question: Concatenate and print your postal address with the City and Country.
# ---
#
#OUR CODE GOES HERE
a = 'mansfield'
b = 'ohio'
c = 'United States'
d = a + b + c
d
# -
| Practice_Notebook_Python_for_Data_Science_Data_Types.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pickle
import matplotlib.pyplot as plt
import numpy as np
import data_manager as dm
import config as cg
# # Aclaracion
# Este jupyter notebook no es parte del repositorio y solo esta para propositos evaluativos, que puedan cargar y observar como funciona esto, por falta de tiempo no se pudo experimentar propiamente las funciones requeridas.
# +
data_loader = dm.image_loader(cg.images_folder, cg.bounding_boxes, cg.train_txt, cg.val_txt, cg.test_txt)
data_loader.set_test_mode()
index = 5
img = data_loader.get_image(index)
# print("shape: ",img.shape)
fig = plt.imshow(img)
plt.show()
mask = data_loader.get_mask(index)
# print("shape: ",img.shape)
fig = plt.imshow(mask,cmap=plt.cm.BuPu_r)
plt.show()
# +
def load_element(path_save, name):
file = open(path_save + '/' + name, 'rb')
element = pickle.load(file)
file.close()
return element
path = 'models'
name = 'population_{:d}_{:d}.pickle'.format(1,1)
population = load_element(path,name)
best_filter = population.get_best_individual()
# -
filter_procesor = dm.Filter_processor(loader = data_loader)
filters = best_filter.get_filters()
mean = best_filter.mean
std = best_filter.var
output_mask = filter_procesor.predict_img(filters,mean,std,index)
fig = plt.imshow(output_mask.reshape(480, 640),cmap=plt.cm.BuPu_r)
plt.show()
# ## Map of heat parameters
fitness = np.empty([3,3])
img_manager = dm.Filter_processor()
# +
path_save = 'models'
for mut in range(3):
for pop in range(3):
print('ind_{:d}_{:d}.pickle'.format(mut,pop))
name_ind = 'ind_{:d}_{:d}.pickle'.format(mut,pop)
file = open(path_save+'/'+name_ind, 'rb')
individual = pickle.load(file)
file.close()
individual.fitness(img_manager)
fitness[mut,pop] = individual.get_fitness()
# +
## population : [5,15,50]
## mutation: [0.01,0.1,0.5]
import numpy as np
import matplotlib.pyplot as plt
# sphinx_gallery_thumbnail_number = 2
mutation = ["0.01", "0.1", "0.4"]
population = ["5", "15",'50']
fig, ax = plt.subplots()
im = ax.imshow(fitness)
# We want to show all ticks...
ax.set_xticks(np.arange(len(population)))
ax.set_yticks(np.arange(len(mutation)))
# ... and label them with the respective list entries
ax.set_xticklabels(population)
ax.set_yticklabels(mutation)
# Rotate the tick labels and set their alignment.
plt.setp(ax.get_xticklabels(), rotation=45, ha="right",
rotation_mode="anchor")
# Loop over data dimensions and create text annotations.
for i in range(len(mutation)):
for j in range(len(population)):
text = ax.text(j, i, '{:.3f}'.format(fitness[i, j]),
ha="center", va="center", color="w")
ax.set_title("Heat map: -Entropy")
fig.tight_layout()
plt.show()
| plotting example.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # 2-Dimensional Frame Analysis - Version 04
# This program performs an elastic analysis of 2-dimensional structural frames. It has the following features:
# 1. Input is provided by a set of CSV files (and cell-magics exist so you can specifiy the CSV data
# in a notebook cell). See the example below for an, er, example.
# 1. Handles concentrated forces on nodes, and concentrated forces, concentrated moments, and linearly varying distributed loads applied transversely anywhere along the member (i.e., there is as yet no way to handle longitudinal
# load components).
# 1. It handles fixed, pinned, roller supports and member end moment releases (internal pins). The former are
# handled by assigning free or fixed global degrees of freedom, and the latter are handled by adjusting the
# member stiffness matrix.
# 1. It has the ability to handle named sets of loads with factored combinations of these.
# 1. The DOF #'s are assigned by the program, with the fixed DOF #'s assigned after the non-fixed. The equilibrium
# equation is then partitioned for solution. Among other advantages, this means that support settlement could be
# easily added (there is no UI for that, yet).
# 1. A non-linear analysis can be performed using the P-Delta method (fake shears are computed at column ends due to the vertical load acting through horizontal displacement differences, and these shears are applied as extra loads
# to the nodes).
# 1. A full non-linear (2nd order) elastic analysis will soon be available by forming the equilibrium equations
# on the deformed structure. This is very easy to add, but it hasn't been done yet. Shouldn't be too long.
# 1. There is very little no documentation below, but that will improve, slowly.
# +
from salib import extend, import_notebooks
from Frame2D_Base import Frame2D, ResultSet
import Frame2D_Input
import Frame2D_Output
import Frame2D_Display
import Frame2D_SolveFirstOrder
# -
| Devel/V05/Frame2D.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Molecular Dynamics
# ### University of California, Berkeley - Spring 2022
# The goal of today’s lecture is to present Molecular Dynamics (MD) simulations of macromolecules and how to run them using Python programmming language. In this lecture, `openmm` package is used for molecular dynamics visualizations.
#
# The following concepts are covered in this notebooks:
#
# * __Newton's Laws of Motion__
# * __Simulation of dynamics of particles__
# * __Proteins and levels of their structure__
# * __Molecular Mechanics__
# * __MD simulations of proteins__
# !pip install MDAnalysis
# !pip install numpy==1.20.1
import os
os.chdir('/home/mohsen/projects/molecular-biomechanics/proteomics/')
from md1 import simulate_apple_fall, simulate_three_particles
from IPython.display import Video
# + [markdown] tags=[]
# ## Newton's Laws of Motion
# -
# Newton's 2nd law connects the kinematics (movements) of a body with its mechanics (total force acting on it) and defines the dynamic evolution of its position:
#
# $$m\frac{d^2r(t)}{dt^2} = F = - \nabla{U(r)},$$
#
# where $m$ is the mass, $r$ is the position, $F$ is the force and $U(r)$ is the potential energy, which depends only on the position of the body.
# If one knows the forces acting upon the body, one can find the position of the body at any moment $r(t)$, i.e. predict its dynamics. This can be done by solving Newton's equation of motion. It is a second order ODE that can be solved analytically for a few simple cases: constant force, harmonic oscillator, periodic force, drag force, etc.
# However, a more general approach is to use computers in order to solve the ODE numerically.
# ---
# ## Simulation of Dynamics of Particles
# There are [many methods](https://en.wikipedia.org/wiki/Numerical_methods_for_ordinary_differential_equations#Methods) for solving ODEs. The second order ODE is transformed to the system of two first order ODEs as follows:
#
# $$\frac{dr(t)}{dt} = v(t)$$
#
# $$m\frac{dv(t)}{dt} = F(t)$$
#
# We use a finite difference approximation that comes to a simple forward Euler Algorithm:
#
# $$ v_{n+1} = v_n + \frac{F_n}{m} dt$$
#
# $$ r_{n+1} = r_n + v_{n+1} dt$$
#
# Here we discretize time t with time step $dt$, so $t_{n+1} = t_n + dt$, and $r_{n} = r(t_n)$, $v_{n} = v(t_n)$, where $n$ is the timestep number. Using this method, computing dynamics is straightforward.
# ---
# ### Example 3.1. Simulation of a projectile on Earth.
# We want to know the dynamics of a green apple ($m = 0.3$ kg) tossed horizontally at 10 cm/s speed from the top of the Toronto CN Tower (553 m) for the first 10 seconds.
# 
simulate_apple_fall(
total_time=10,
mass=0.3,
initial_velocity=-0.1,
height=553,
timestep=0.05,
)
Video('./media/apple_fall.mp4')
# When a closed system of particles are interacting through pairwise potentials, the force on each particle $i$ depends on its position with respect to every other particle $j$:
#
# $$m_i\frac{d^2r_i(t)}{dt^2} = \sum_jF_{ij}(t) = -\sum_j\nabla_i{U(|r_{ij}(t)|)}$$
#
# where $r_{ij} = \sqrt{(x_i - x_j)^2 + (y_i - y_j)^2 + (z_i - z_j)^2}$ is the distance between particle $i$ and $j$, and $i,j \in (1,N)$.
# ---
# ### Example 3.2. Simulation of 3-body problem with Hooke's law:
#
# We want to know the dynamics of 3 particles $m = 1$ kg connected to each other with invisible springs with $K_s = 5$ N/m, and $r_0 = 1$ m initially located at (0, 2), (2, 0) and (-1, 0) on the 2D plane for the first 10 seconds of their motion.
#
# **Hint:**
# The pairwise potential is (Hooke's Law): $$U(r_{ij}) = \frac{K_s}{2}(r_{ij} - r_0)^2$$
#
# The negative gradient of the potential is a force from $j$-th upon $i$-th:
#
# $$\mathbf{F_{ij}} = - \nabla_i{U(r_{ij})} = - K_s (r_{ij} - r_0) \nabla_i r_{ij} = - K_s (r_{ij} - r_0) \frac{\mathbf{r_{ij}}}{|r_{ij}|}$$
#
simulate_three_particles(
total_time=26, mass=1.0, ks=5, r0=1.0, timestep=0.05
)
Video('./media/3particles.mp4')
# ---
# ## Proteins, structure and functions
# <img src="./media/protein_structure.png" width="400" align="right">
#
# While we now have a basic knowledge of the purpose and methodology of simulations, we still need to understand what proteins are and why they are important.
#
# [Protein structure](https://en.wikipedia.org/wiki/Protein_structure) is the three-dimensional arrangement of atoms in a protein, which is a chain of amino acids. Proteins are polymers – specifically polypeptides – formed from sequences of 20 types of amino acids, the monomers of the polymer. A single amino acid monomer may also be called a residue, indicating a repeating unit of a polymer. To be able to perform their biological function, proteins fold into one or more specific spatial conformations driven by a number of non-covalent interactions such as:
#
# - hydrogen bonding
# - ionic interactions
# - Van der Waals forces
# - hydrophobic packing
#
# To understand the functions of proteins at a molecular level, it is often necessary to determine their three-dimensional structure using techniques such as X-ray crystallography, NMR spectroscopy, and others.
#
# ### 4.1 Levels of structure:
#
# **Primary structure** of a protein refers to the sequence of amino acids in the polypeptide chain.
#
# **Secondary structure** refers to highly regular local sub-structures of the actual polypeptide backbone chain. There are two main types of secondary structure: the α-helix and the β-strand or β-sheets.
#
# **Tertiary structure** refers to the three-dimensional structure of monomeric and multimeric protein molecules. The α-helixes and β-sheets are folded into a compact globular structure.
#
# **Quaternary structure** is the three-dimensional structure consisting of two or more individual polypeptide chains (subunits) that operate as a single functional unit (multimer).
#
#
# ### 4.2 Functions:
#
# - *Antibodies* - bind to specific foreign particles, ex: IgG
# - *Enzymes* - speed up chemical reactions, ex: Lysozyme
# - *Messengers* - transmit signals, ex: Growth hormone
# - *Structural components* - support for cells, ex: Tubulin
# - *Transport/storage* - bind and carry small molecules, ex: Hemoglobin
#
#
# **Lysozyme** is a protein-enzyme (found in tears, saliva, mucus and egg white) that is a part of the innate immune system with antimicrobial activity characterized by the ability to damage the cell wall of bacteria. Bacteria have polysaccharides (sugars) in their cell wall, that bind to the groove, and lysozyme cuts the bond and destroys bacteria.
#
# <!-- |  |  |  |
# |:-:|:-:|:-:|
# | Sequence | Structure | Function |
#
# Figure credit: [C.Ing](https://github.com/cing/HackingStructBiolTalk) and [wikipedia](https://en.wikipedia.org/wiki/Protein_structure) -->
# ---
# ## Molecular Mechanics
# Since we now know what proteins are and why these molecular machines are important, we consider the method to model them. The basic idea is to create the same kind of approach as we used in the 3-body simulation. Now, our system consists of thousands particles (atoms of the protein plus atoms of surrounding water) and they all are connected via a complex potential energy function.
#
# An all-atom potential energy function $V$ is usually given by the sum of the bonded terms ($V_b$) and non-bonded terms ($V_{nb}$), i.e.
#
# $$V = V_{b} + V_{nb},$$
#
# where the bonded potential includes the harmonic (covalent) bond part, the harmonic angle and
# the two types of torsion (dihedral) angles: proper and improper. As it can be seen, these functions are mostly harmonic potentials
#
# $$V_{b} = \sum_{bonds}\frac{1}{2}K_b(b-b_0)^2 + \sum_{angles}K_{\theta}(\theta-\theta_0)^2 + \sum_{dihedrals}K_{\phi}(1-cos(n\phi - \phi_0)) + \sum_{impropers}K_{\psi}(\psi-\psi_0)^2$$
#
# For example, $b$ and $\theta$ represent the distance between two atoms and the angle between two
# adjacent bonds; $\phi$ and $\psi$ are dihedral (torsion) angles. These can be evaluated for all the
# atoms from their current positions. Also, $K_b$, $K_\theta$, $K_\phi$, and $K_\psi$ are the spring constants, associated
# with bond vibrations, bending of bond angles, and conformational fluctuations in dihedral and
# improper angles around some equilibrium values $b_0$, $\theta_0$, $\phi_0$, and $\psi_0$, respectively.
#
# The non-bonded part of the potential energy function is represented by the electrostatic and van der Waals potentials, i.e.
#
# $$V_{nb} = \sum_{i,j}\left(\frac{q_{i}q_{j}}{4\pi\varepsilon_{0}\varepsilon r_{ij}} + \varepsilon_{ij}\left[\left(\frac{\sigma^{min}_{ij}}{r_{ij}}\right)^{12}-2\left(\frac{\sigma^{min}_{ij}}{r_{ij}}\right)^{6}\right]\right)$$
#
# where $r_{ij}$ is the distance between two interacting atoms, $q_i$ and $q_j$ are their electric charges; $\varepsilon$ and
# $\varepsilon_0$ are electric and dielectric constant; $\varepsilon_{ij} = \sqrt{\varepsilon_i\varepsilon_j}$ and
# $\sigma_{ij} = \frac{\sigma_i + \sigma_j}{2}$ are van der Waals parameters for atoms $i$ and $j$.
#
# **Importantly, each force field has its own set of parameters, which are different for different types of atoms.**
#
# 
#
# ---
# ## Molecular dynamics of proteins <a id='l_md'></a>
# [**Molecular dynamics (MD)**](https://en.wikipedia.org/wiki/Molecular_dynamics) is a computer simulation method for studying the physical movements of atoms and molecules, i.e. their dynamical evolution.
#
# In the most common version, the trajectories of atoms and molecules are determined by numerically solving Newton's equations of motion for a system of interacting particles, where forces between the particles and their potential energies are often calculated using molecular mechanics force fields.
#
#
#
# Now with all that intellectual equipment, we can start running legit Molecular Dynamics simulations. All we need is an initial structure of the protein and software that computes its dynamics efficiently.
# ### Procedure
#
# 1. Load initial coordinates of protein atoms (from *.pdb file)
# 2. Choose force field parameters (in potential function V from section 5).
# 3. Choose parameters of the experiment: temperature, pressure, box size, solvation, boundary conditions
# 4. Choose integrator, i.e. algorithm for solving equation of motion
# 5. Run simulation, save coordinates time to time (to *.dcd file).
# 6. Visualize the trajectory
# 7. Perform the analysis
# __NOTE__: It is better for students to gain a little understanding of how the following packages are working under the hood before continuing the notebook.
#
# * __NGLViewer__: NGL Viewer is a collection of tools for web-based molecular graphics. WebGL is employed to display molecules like proteins and DNA/RNA with a variety of representations.
#
# * __MDAnalysis__: MDAnalysis is an object-oriented Python library to analyze trajectories from molecular dynamics (MD) simulations in many popular formats. It can write most of these formats, too, together with atom selections suitable for visualization or native analysis tools.
#
# * __Openmm__: Openmm consists of two parts: One is a set of libraries that lets programmers easily add molecular simulation features to their programs and the other parts is an “application layer” that exposes those features to end users who just want to run simulations
pdb_file = 'data/villin_water.pdb'
# pdb_file = 'data/polyALA.pdb'
# pdb_file = 'data/polyGLY.pdb'
# pdb_file = 'data/polyGV.pdb'
file0 = open(pdb_file, 'r')
counter = 0
for line in file0:
if counter < 10:
print(line)
counter += 1
# +
from simtk.openmm.app import *
from simtk.openmm import *
from simtk.unit import *
import MDAnalysis as md
from MDAnalysis.tests import datafiles
import nglview as ng
from sys import stdout
u = md.Universe(datafiles.PSF, datafiles.DCD)
view = ng.show_mdanalysis(u, gui=True)
view
# -
# ---
# ### Example: MD simulation of protein folding into alpha-helix
# Run a simulation of fully extended polyalanine __polyALA.pdb__ for 400 picoseconds in a vacuo environment with T=300 K and see if it can fold to any secondary structure:
# +
### 1.loading initial coordinates
pdb_file = "data/polyGV.pdb"
pdb = PDBFile(pdb_file)
### 2.choosing a forcefield parameters
ff = ForceField('amber10.xml')
system = ff.createSystem(pdb.topology, nonbondedMethod=CutoffNonPeriodic)
### 3. Choose parameters of the experiment: temperature, pressure, box size, solvation, boundary conditions, etc
temperature = 300*kelvin
frictionCoeff = 1/picosecond
time_step = 0.002*picoseconds
total_steps = 400*picoseconds / time_step
### 4. Choose an algorithm (integrator)
integrator = LangevinIntegrator(temperature, frictionCoeff, time_step)
### 5. Run simulation, saving coordinates time to time:
### 5a. Create a simulation object
simulation = Simulation(pdb.topology, system, integrator)
simulation.context.setPositions(pdb.positions)
### 5b. Minimize energy
simulation.minimizeEnergy()
### 5c. Save coordinates to dcd file and energies to a standard output console:
simulation.reporters.append(DCDReporter('data/polyALA_traj.dcd', 1000))
simulation.reporters.append(StateDataReporter(stdout, 5000, step=True, potentialEnergy=True,\
temperature=True, progress=True, totalSteps = total_steps))
### 5d. Run!
simulation.step(total_steps)
# -
### 6. Visualization
sys = md.Universe(pdb_file, 'data/polyALA_traj.dcd')
ng.show_mdanalysis(sys, gui=True)
# ## Congrats!
#
# The notebook is available at https://github.com/Naghipourfar/molecular-biomechanics/
| proteomics/MD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib notebook
# %matplotlib inline
import numpy as np
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.font_manager as fm
myfont = fm.FontProperties(fname='C:/Windows/Fonts/STKAITI.TTF',size=14)
matplotlib.rcParams["axes.unicode_minus"] = False
def subplot_plot():
'''子图'''
# 子图列表
style_list = ['g+-','r*-','b.-','yo-'] # plot中的fmt参数[颜色][标记样式][线形]
# 依次画图
for i in range(4):
x = np.linspace(0.,2+i,num=10*(i+1))
y = np.sin((5-i)*np.pi*x)
# 子图的生成
plt.subplot(2,2,i+1)
plt.title("子图 %d" % (i+1),fontproperties=myfont)
plt.plot(x,y,style_list[i])
plt.show()
return
subplot_plot()
# 柱状图
def bar():
apple = (20,35,30,35,27)
orange = (25,32,34,20,25)
plt.title("柱状图",fontproperties=myfont)
index = np.arange(len(apple))
bar_width = 0.3
plt.bar(index,apple,width=bar_width,alpha=0.2,color='m',label='苹果')
plt.bar(index+bar_width,orange,width=bar_width,alpha=0.8,color='y',label='橘子')
plt.legend(loc='upper right',prop=myfont,shadow=True)
# 设置柱状图标示
for x,y in zip(index,apple):
plt.text(x,y+0.3,y,ha="center",va="bottom")
for x,y in zip(index,orange):
plt.text(x+bar_width,y+0.3,y,ha="center",va="bottom")
# 设置刻度范围及坐标轴名称
plt.ylim(0,45)
plt.xlabel("Group")
plt.ylabel("Scores")
plt.xticks(index+(bar_width/2),('A','B','C','D','E'))
plt.show()
return
bar()
# 带折线的柱状图
def bar_with_line():
men = np.array((20, 35, 30, 35, 27, 25, 32, 34, 20, 25))
women = np.array((25, 32, 34, 20, 25, 20, 35, 30, 35, 27))
plt.title("Bar With Line")
index = np.arange(len(men))
bar_width = 0.7
# 画柱状图
plt.bar(index,men,width=bar_width,alpha=0.4,color='m',label='men')
plt.bar(index, -women, width=bar_width,alpha=0.4,color="r",label="women")
# 画折线图
plt.plot(index,men, marker="o", linestyle="-", color="r",label='men line')
plt.plot(index, -women, marker=".", linestyle="--", color="b",label="women line")
# 图形标示
for x, y in zip(index,men):
plt.text(x, y+1, y, ha="center", va="bottom")
for x, y in zip(index,women):
plt.text(x, -y-1, y, ha="center", va="top")
# 设置y轴和图例
plt.ylim(-45,80)
plt.legend(loc='upper right',shadow=True)
plt.show()
return
bar_with_line()
# 层次柱状图
def table_plot():
# 生成测试数据
data = np.array([
[1, 4, 2, 5, 2],
[2, 1, 1, 3, 6],
[5, 3, 6, 4, 1]
])
# 设置标题
plt.title("层次柱状图", fontproperties=myfont)
# 设置相关参数
index = np.arange(len(data[0]))
color_index = ["r", "g", "b"]
# 声明底部位置
bottom = np.array([0, 0, 0, 0, 0])
# 依次画图,并更新底部位置
for i in range(len(data)):
plt.bar(index, data[i], width=0.5, color=color_index[i], bottom=bottom, alpha=0.7, label="标签 %d" % i)
bottom += data[i]
# 设置图例位置
plt.legend(loc="upper left", prop=myfont, shadow=True)
# 图形显示
plt.show()
return
table_plot()
# 饼图
def pie_plot():
# 生成测试数据
sizes = [15, 30, 45, 10]
labels = ["Frogs", "中文", "Dogs", "Logs"]
colors = ["yellowgreen", "gold", "lightskyblue", "lightcoral"]
# 设置标题
plt.title("饼图", fontproperties=myfont)
# 设置突出参数
explode = [0, 0.05, 0, 0]
# 画饼状图
patches, l_text, p_text = plt.pie(sizes, explode=explode,
labels=labels, colors=colors,
autopct="%1.1f%%", shadow=True,
startangle=90)
for text in l_text:
text.set_fontproperties(myfont)
plt.axis("equal")
# 图形显示
plt.show()
return
pie_plot()
# 散点图
def scatter_plot():
# 生成测试数据
point_count = 1000
x_index = np.random.random(point_count)
y_index = np.random.random(point_count)
# 设置标题
plt.title("散点图", fontproperties=myfont)
# 设置相关参数
color_list = np.random.random(point_count)
scale_list = np.random.random(point_count) * 100
# 画散点图
plt.scatter(x_index, y_index, s=scale_list, c=color_list, marker="o")
# 图形显示
plt.show()
return
scatter_plot()
# 填充图
def fill_plot():
# 生成测试数据
x = np.linspace(-2*np.pi, 2*np.pi, 1000, endpoint=True)
y = np.sin(x)
# 设置标题
plt.title("填充图", fontproperties=myfont)
# 画图
plt.plot(x, y, color="blue", alpha=1.00)
# 填充图形, plt.fill_between(x, y1, y2, where=None, *kwargs)
plt.fill_between(x, 0, y, where=(y > 0), color="blue", alpha=0.25)
plt.fill_between(x, 0, y, where=(y < 0), color="red", alpha=0.25)
# 图形显示
plt.show()
return
fill_plot()
| matplot/plots.ipynb |
# -*- coding: utf-8 -*-
# # Julia REPL
#
# Julia 附带了一个全功能的交互式命令行 REPL(read-eval-print loop),其内置于 `julia` 可执行文件中。它除了允许快速简便地执行 Julia 语句外,还具有可搜索的历史记录,tab 补全,许多有用的按键绑定以及专用的 help 和 shell 模式。只需不附带任何参数地调用 `julia` 或双击可执行文件即可启动 REPL:
# + attributes={"classes": ["@eval"], "id": ""}
io = IOBuffer()
Base.banner(io)
banner = String(take!(io))
import Markdown
Markdown.parse("```\n\$ julia\n\n$(banner)\njulia>\n```")
# -
# 要退出交互式会话,在空白行上键入 `^D`——control 键和 `d` 键,或者先键入 `quit()`,然后键入 return 或 enter 键。REPL 用横幅和 `julia>` 提示符欢迎你。
#
# ## 不同的提示符模式
#
# ### Julian 模式
#
# The REPL has five main modes of operation. The first and most common is the Julian prompt. It
# is the default mode of operation; each new line initially starts with `julia>`. It is here that
# you can enter Julia expressions. Hitting return or enter after a complete expression has been
# entered will evaluate the entry and show the result of the last expression.
# + attributes={"classes": ["jldoctest"], "id": ""}
julia> string(1 + 2)
"3"
# -
# 交互式运行有许多独特的实用功能。除了显示结果外,REPL 还将结果绑定到变量 `ans` 上。一行的尾随分号可用作禁止显示结果的标志。
# + attributes={"classes": ["jldoctest"], "id": ""}
julia> string(3 * 4);
julia> ans
"12"
# -
# In Julia mode, the REPL supports something called *prompt pasting*. This activates when pasting
# text that starts with `julia> ` into the REPL. In that case, only expressions starting with
# `julia> ` are parsed, others are removed. This makes it possible to paste a chunk of code
# that has been copied from a REPL session without having to scrub away prompts and outputs. This
# feature is enabled by default but can be disabled or enabled at will with `REPL.enable_promptpaste(::Bool)`.
# If it is enabled, you can try it out by pasting the code block above this paragraph straight into
# the REPL. This feature does not work on the standard Windows command prompt due to its limitation
# at detecting when a paste occurs.
#
# Objects are printed at the REPL using the [`show`](@ref) function with a specific [`IOContext`](@ref).
# In particular, the `:limit` attribute is set to `true`.
# Other attributes can receive in certain `show` methods a default value if it's not already set,
# like `:compact`.
# It's possible, as an experimental feature, to specify the attributes used by the REPL via the
# `Base.active_repl.options.iocontext` dictionary (associating values to attributes). For example:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> rand(2, 2)
2×2 Array{Float64,2}:
0.8833 0.329197
0.719708 0.59114
julia> show(IOContext(stdout, :compact => false), "text/plain", rand(2, 2))
0.43540323669187075 0.15759787870609387
0.2540832269192739 0.4597637838786053
julia> Base.active_repl.options.iocontext[:compact] = false;
julia> rand(2, 2)
2×2 Array{Float64,2}:
0.2083967319174056 0.13330606013126012
0.6244375177790158 0.9777957560761545
# -
# In order to define automatically the values of this dictionary at startup time, one can use the
# [`atreplinit`](@ref) function in the `~/.julia/config/startup.jl` file, for example:
# + attributes={"classes": ["julia"], "id": ""}
atreplinit() do repl
repl.options.iocontext[:compact] = false
end
# -
# ### 帮助模式
#
# When the cursor is at the beginning of the line, the prompt can be changed to a help mode by typing
# `?`. Julia will attempt to print help or documentation for anything entered in help mode:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> ? # upon typing ?, the prompt changes (in place) to: help?>
help?> string
search: string String Cstring Cwstring RevString randstring bytestring SubString
string(xs...)
Create a string from any values using the print function.
# -
# Macros, types and variables can also be queried:
# +
help?> @time
@time
A macro to execute an expression, printing the time it took to execute, the number of allocations,
and the total number of bytes its execution caused to be allocated, before returning the value of the
expression.
See also @timev, @timed, @elapsed, and @allocated.
help?> Int32
search: Int32 UInt32
Int32 <: Signed
32-bit signed integer type.
# -
# A string or regex literal searches all docstrings using [`apropos`](@ref):
# +
help?> "aprop"
REPL.stripmd
Base.Docs.apropos
help?> r"ap..p"
Base.:∘
Base.shell_escape_posixly
Distributed.CachingPool
REPL.stripmd
Base.Docs.apropos
# -
# Help mode can be exited by pressing backspace at the beginning of the line.
#
# ### [Shell mode](@id man-shell-mode)
#
# Just as help mode is useful for quick access to documentation, another common task is to use the
# system shell to execute system commands. Just as `?` entered help mode when at the beginning
# of the line, a semicolon (`;`) will enter the shell mode. And it can be exited by pressing backspace
# at the beginning of the line.
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> ; # upon typing ;, the prompt changes (in place) to: shell>
shell> echo hello
hello
# -
# !!! note
# For Windows users, Julia's shell mode does not expose windows shell commands.
# Hence, this will fail:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> ; # upon typing ;, the prompt changes (in place) to: shell>
shell> dir
ERROR: IOError: could not spawn `dir`: no such file or directory (ENOENT)
Stacktrace!
.......
# -
# However, you can get access to `PowerShell` like this:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> ; # upon typing ;, the prompt changes (in place) to: shell>
shell> powershell
Windows PowerShell
Copyright (C) Microsoft Corporation. All rights reserved.
PS C:\Users\elm>
# -
# ... and to `cmd.exe` like that (see the `dir` command):
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> ; # upon typing ;, the prompt changes (in place) to: shell>
shell> cmd
Microsoft Windows [version 10.0.17763.973]
(c) 2018 Microsoft Corporation. All rights reserved.
C:\Users\elm>dir
Volume in drive C has no label
Volume Serial Number is 1643-0CD7
Directory of C:\Users\elm
29/01/2020 22:15 <DIR> .
29/01/2020 22:15 <DIR> ..
02/02/2020 08:06 <DIR> .atom
# -
# ### Pkg mode
#
# The Package manager mode accepts specialized commands for loading and updating packages. It is entered
# by pressing the `]` key at the Julian REPL prompt and exited by pressing CTRL-C or pressing the backspace key
# at the beginning of the line. The prompt for this mode is `pkg>`. It supports its own help-mode, which is
# entered by pressing `?` at the beginning of the line of the `pkg>` prompt. The Package manager mode is
# documented in the Pkg manual, available at [https://julialang.github.io/Pkg.jl/v1/](https://julialang.github.io/Pkg.jl/v1/).
#
# ### Search modes
#
# In all of the above modes, the executed lines get saved to a history file, which can be searched.
# To initiate an incremental search through the previous history, type `^R` -- the control key
# together with the `r` key. The prompt will change to ```(reverse-i-search)`':```, and as you
# type the search query will appear in the quotes. The most recent result that matches the query
# will dynamically update to the right of the colon as more is typed. To find an older result using
# the same query, simply type `^R` again.
#
# Just as `^R` is a reverse search, `^S` is a forward search, with the prompt ```(i-search)`':```.
# The two may be used in conjunction with each other to move through the previous or next matching
# results, respectively.
#
# ## Key bindings
#
# The Julia REPL makes great use of key bindings. Several control-key bindings were already introduced
# above (`^D` to exit, `^R` and `^S` for searching), but there are many more. In addition to the
# control-key, there are also meta-key bindings. These vary more by platform, but most terminals
# default to using alt- or option- held down with a key to send the meta-key (or can be configured
# to do so), or pressing Esc and then the key.
#
# | Keybinding | Description |
# |:------------------- |:---------------------------------------------------------------------------------------------------------- |
# | **Program control** | |
# | `^D` | Exit (when buffer is empty) |
# | `^C` | Interrupt or cancel |
# | `^L` | Clear console screen |
# | Return/Enter, `^J` | New line, executing if it is complete |
# | meta-Return/Enter | Insert new line without executing it |
# | `?` or `;` | Enter help or shell mode (when at start of a line) |
# | `^R`, `^S` | Incremental history search, described above |
# | **Cursor movement** | |
# | Right arrow, `^F` | Move right one character |
# | Left arrow, `^B` | Move left one character |
# | ctrl-Right, `meta-F`| Move right one word |
# | ctrl-Left, `meta-B` | Move left one word |
# | Home, `^A` | Move to beginning of line |
# | End, `^E` | Move to end of line |
# | Up arrow, `^P` | Move up one line (or change to the previous history entry that matches the text before the cursor) |
# | Down arrow, `^N` | Move down one line (or change to the next history entry that matches the text before the cursor) |
# | Shift-Arrow Key | Move cursor according to the direction of the Arrow key, while activating the region ("shift selection") |
# | Page-up, `meta-P` | Change to the previous history entry |
# | Page-down, `meta-N` | Change to the next history entry |
# | `meta-<` | Change to the first history entry (of the current session if it is before the current position in history) |
# | `meta->` | Change to the last history entry |
# | `^-Space` | Set the "mark" in the editing region (and de-activate the region if it's active) |
# | `^-Space ^-Space` | Set the "mark" in the editing region and make the region "active", i.e. highlighted |
# | `^G` | De-activate the region (i.e. make it not highlighted) |
# | `^X^X` | Exchange the current position with the mark |
# | **Editing** | |
# | Backspace, `^H` | Delete the previous character, or the whole region when it's active |
# | Delete, `^D` | Forward delete one character (when buffer has text) |
# | meta-Backspace | Delete the previous word |
# | `meta-d` | Forward delete the next word |
# | `^W` | Delete previous text up to the nearest whitespace |
# | `meta-w` | Copy the current region in the kill ring |
# | `meta-W` | "Kill" the current region, placing the text in the kill ring |
# | `^K` | "Kill" to end of line, placing the text in the kill ring |
# | `^Y` | "Yank" insert the text from the kill ring |
# | `meta-y` | Replace a previously yanked text with an older entry from the kill ring |
# | `^T` | Transpose the characters about the cursor |
# | `meta-Up arrow` | Transpose current line with line above |
# | `meta-Down arrow` | Transpose current line with line below |
# | `meta-u` | Change the next word to uppercase |
# | `meta-c` | Change the next word to titlecase |
# | `meta-l` | Change the next word to lowercase |
# | `^/`, `^_` | Undo previous editing action |
# | `^Q` | Write a number in REPL and press `^Q` to open editor at corresponding stackframe or method |
# | `meta-Left Arrow` | indent the current line on the left |
# | `meta-Right Arrow` | indent the current line on the right |
# | `meta-.` | insert last word from previous history entry |
#
# ### Customizing keybindings
#
# Julia's REPL keybindings may be fully customized to a user's preferences by passing a dictionary
# to `REPL.setup_interface`. The keys of this dictionary may be characters or strings. The key
# `'*'` refers to the default action. Control plus character `x` bindings are indicated with `"^x"`.
# Meta plus `x` can be written `"\\M-x"` or `"\ex"`, and Control plus `x` can be written
# `"\\C-x"` or `"^x"`.
# The values of the custom keymap must be `nothing` (indicating
# that the input should be ignored) or functions that accept the signature
# `(PromptState, AbstractREPL, Char)`.
# The `REPL.setup_interface` function must be called before the REPL is initialized, by registering
# the operation with [`atreplinit`](@ref) . For example, to bind the up and down arrow keys to move through
# history without prefix search, one could put the following code in `~/.julia/config/startup.jl`:
# + attributes={"classes": ["julia"], "id": ""}
import REPL
import REPL.LineEdit
const mykeys = Dict{Any,Any}(
# Up Arrow
"\e[A" => (s,o...)->(LineEdit.edit_move_up(s) || LineEdit.history_prev(s, LineEdit.mode(s).hist)),
# Down Arrow
"\e[B" => (s,o...)->(LineEdit.edit_move_down(s) || LineEdit.history_next(s, LineEdit.mode(s).hist))
)
function customize_keys(repl)
repl.interface = REPL.setup_interface(repl; extra_repl_keymap = mykeys)
end
atreplinit(customize_keys)
# -
# Users should refer to `LineEdit.jl` to discover the available actions on key input.
#
# ## Tab completion
#
# In both the Julian and help modes of the REPL, one can enter the first few characters of a function
# or type and then press the tab key to get a list all matches:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> stri[TAB]
stride strides string strip
julia> Stri[TAB]
StridedArray StridedMatrix StridedVecOrMat StridedVector String
# -
# The tab key can also be used to substitute LaTeX math symbols with their Unicode equivalents,
# and get a list of LaTeX matches as well:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> \pi[TAB]
julia> π
π = 3.1415926535897...
julia> e\_1[TAB] = [1,0]
julia> e₁ = [1,0]
2-element Array{Int64,1}:
1
0
julia> e\^1[TAB] = [1 0]
julia> e¹ = [1 0]
1×2 Array{Int64,2}:
1 0
julia> \sqrt[TAB]2 # √ is equivalent to the sqrt function
julia> √2
1.4142135623730951
julia> \hbar[TAB](h) = h / 2\pi[TAB]
julia> ħ(h) = h / 2π
ħ (generic function with 1 method)
julia> \h[TAB]
\hat \hermitconjmatrix \hkswarow \hrectangle
\hatapprox \hexagon \hookleftarrow \hrectangleblack
\hbar \hexagonblack \hookrightarrow \hslash
\heartsuit \hksearow \house \hspace
julia> α="\alpha[TAB]" # LaTeX completion also works in strings
julia> α="α"
# -
# A full list of tab-completions can be found in the [Unicode Input](@ref) section of the manual.
#
# Completion of paths works for strings and julia's shell mode:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> path="/[TAB]"
.dockerenv .juliabox/ boot/ etc/ lib/ media/ opt/ root/ sbin/ sys/ usr/
.dockerinit bin/ dev/ home/ lib64/ mnt/ proc/ run/ srv/ tmp/ var/
shell> /[TAB]
.dockerenv .juliabox/ boot/ etc/ lib/ media/ opt/ root/ sbin/ sys/ usr/
.dockerinit bin/ dev/ home/ lib64/ mnt/ proc/ run/ srv/ tmp/ var/
# -
# Tab completion can help with investigation of the available methods matching the input arguments:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> max([TAB] # All methods are displayed, not shown here due to size of the list
julia> max([1, 2], [TAB] # All methods where `Vector{Int}` matches as first argument
max(x, y) in Base at operators.jl:215
max(a, b, c, xs...) in Base at operators.jl:281
julia> max([1, 2], max(1, 2), [TAB] # All methods matching the arguments.
max(x, y) in Base at operators.jl:215
max(a, b, c, xs...) in Base at operators.jl:281
# -
# Keywords are also displayed in the suggested methods after `;`, see below line where `limit`
# and `keepempty` are keyword arguments:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> split("1 1 1", [TAB]
split(str::AbstractString; limit, keepempty) in Base at strings/util.jl:302
split(str::T, splitter; limit, keepempty) where T<:AbstractString in Base at strings/util.jl:277
# -
# The completion of the methods uses type inference and can therefore see if the arguments match
# even if the arguments are output from functions. The function needs to be type stable for the
# completion to be able to remove non-matching methods.
#
# Tab completion can also help completing fields:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> import UUIDs
julia> UUIDs.uuid[TAB]
uuid1 uuid4 uuid_version
# -
# Fields for output from functions can also be completed:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> split("","")[1].[TAB]
lastindex offset string
# -
# The completion of fields for output from functions uses type inference, and it can only suggest
# fields if the function is type stable.
#
# Dictionary keys can also be tab completed:
# + attributes={"classes": ["julia-repl"], "id": ""}
julia> foo = Dict("qwer1"=>1, "qwer2"=>2, "asdf"=>3)
Dict{String,Int64} with 3 entries:
"qwer2" => 2
"asdf" => 3
"qwer1" => 1
julia> foo["q[TAB]
"qwer1" "qwer2"
julia> foo["qwer
# -
# ## Customizing Colors
#
# The colors used by Julia and the REPL can be customized, as well. To change the
# color of the Julia prompt you can add something like the following to your
# `~/.julia/config/startup.jl` file, which is to be placed inside your home directory:
# + attributes={"classes": ["julia"], "id": ""}
function customize_colors(repl)
repl.prompt_color = Base.text_colors[:cyan]
end
atreplinit(customize_colors)
# -
# The available color keys can be seen by typing `Base.text_colors` in the help mode of the REPL.
# In addition, the integers 0 to 255 can be used as color keys for terminals
# with 256 color support.
#
# You can also change the colors for the help and shell prompts and
# input and answer text by setting the appropriate field of `repl` in the `customize_colors` function
# above (respectively, `help_color`, `shell_color`, `input_color`, and `answer_color`). For the
# latter two, be sure that the `envcolors` field is also set to false.
#
# It is also possible to apply boldface formatting by using
# `Base.text_colors[:bold]` as a color. For instance, to print answers in
# boldface font, one can use the following as a `~/.julia/config/startup.jl`:
# + attributes={"classes": ["julia"], "id": ""}
function customize_colors(repl)
repl.envcolors = false
repl.answer_color = Base.text_colors[:bold]
end
atreplinit(customize_colors)
# -
# You can also customize the color used to render warning and informational messages by
# setting the appropriate environment variables. For instance, to render error, warning, and informational
# messages respectively in magenta, yellow, and cyan you can add the following to your
# `~/.julia/config/startup.jl` file:
# + attributes={"classes": ["julia"], "id": ""}
ENV["JULIA_ERROR_COLOR"] = :magenta
ENV["JULIA_WARN_COLOR"] = :yellow
ENV["JULIA_INFO_COLOR"] = :cyan
# -
# ## TerminalMenus
#
# TerminalMenus is a submodule of the Julia REPL and enables small, low-profile interactive menus in the terminal.
#
# ### Examples
# + attributes={"classes": ["julia"], "id": ""}
import REPL
using REPL.TerminalMenus
options = ["apple", "orange", "grape", "strawberry",
"blueberry", "peach", "lemon", "lime"]
# -
# #### RadioMenu
#
# The RadioMenu allows the user to select one option from the list. The `request`
# function displays the interactive menu and returns the index of the selected
# choice. If a user presses 'q' or `ctrl-c`, `request` will return a `-1`.
# + attributes={"classes": ["julia"], "id": ""}
# `pagesize` is the number of items to be displayed at a time.
# The UI will scroll if the number of options is greater
# than the `pagesize`
menu = RadioMenu(options, pagesize=4)
# `request` displays the menu and returns the index after the
# user has selected a choice
choice = request("Choose your favorite fruit:", menu)
if choice != -1
println("Your favorite fruit is ", options[choice], "!")
else
println("Menu canceled.")
end
# -
# Output:
Choose your favorite fruit:
^ grape
strawberry
> blueberry
v peach
Your favorite fruit is blueberry!
# #### MultiSelectMenu
#
# The MultiSelectMenu allows users to select many choices from a list.
# + attributes={"classes": ["julia"], "id": ""}
# here we use the default `pagesize` 10
menu = MultiSelectMenu(options)
# `request` returns a `Set` of selected indices
# if the menu us canceled (ctrl-c or q), return an empty set
choices = request("Select the fruits you like:", menu)
if length(choices) > 0
println("You like the following fruits:")
for i in choices
println(" - ", options[i])
end
else
println("Menu canceled.")
end
# -
# Output:
Select the fruits you like:
[press: d=done, a=all, n=none]
[ ] apple
> [X] orange
[X] grape
[ ] strawberry
[ ] blueberry
[X] peach
[ ] lemon
[ ] lime
You like the following fruits:
- orange
- grape
- peach
# ### Customization / Configuration
#
# #### ConfiguredMenu subtypes
#
# Starting with Julia 1.6, the recommended way to configure menus is via the constructor.
# For instance, the default multiple-selection menu
# +
julia> menu = MultiSelectMenu(options, pagesize=5);
julia> request(menu) # ASCII is used by default
[press: d=done, a=all, n=none]
[ ] apple
[X] orange
[ ] grape
> [X] strawberry
v [ ] blueberry
# -
# can instead be rendered with Unicode selection and navigation characters with
# + attributes={"classes": ["julia"], "id": ""}
julia> menu = MultiSelectMenu(options, pagesize=5, charset=:unicode);
julia> request(menu)
[press: d=done, a=all, n=none]
⬚ apple
✓ orange
⬚ grape
→ ✓ strawberry
↓ ⬚ blueberry
# -
# More fine-grained configuration is also possible:
# + attributes={"classes": ["julia"], "id": ""}
julia> menu = MultiSelectMenu(options, pagesize=5, charset=:unicode, checked="YEP!", unchecked="NOPE", cursor='⧐');
julia> request(menu)
julia> request(menu)
[press: d=done, a=all, n=none]
NOPE apple
YEP! orange
NOPE grape
⧐ YEP! strawberry
↓ NOPE blueberry
# -
# Aside from the overall `charset` option, for `RadioMenu` the configurable options are:
#
# - `cursor::Char='>'|'→'`: character to use for cursor
# - `up_arrow::Char='^'|'↑'`: character to use for up arrow
# - `down_arrow::Char='v'|'↓'`: character to use for down arrow
# - `updown_arrow::Char='I'|'↕'`: character to use for up/down arrow in one-line page
# - `scroll_wrap::Bool=false`: optionally wrap-around at the beginning/end of a menu
# - `ctrl_c_interrupt::Bool=true`: If `false`, return empty on ^C, if `true` throw InterruptException() on ^C
#
# `MultiSelectMenu` adds:
#
# - `checked::String="[X]"|"✓"`: string to use for checked
# - `unchecked::String="[ ]"|"⬚")`: string to use for unchecked
#
# You can create new menu types of your own.
# Types that are derived from `TerminalMenus.ConfiguredMenu` configure the menu options at construction time.
#
# #### Legacy interface
#
# Prior to Julia 1.6, and still supported throughout Julia 1.x, one can also configure menus by calling
# `TerminalMenus.config()`.
#
# ## References
#
# ### REPL
# + attributes={"classes": ["@docs"], "id": ""}
Base.atreplinit
# -
# ### TerminalMenus
#
# #### Configuration
# + attributes={"classes": ["@docs"], "id": ""}
REPL.TerminalMenus.Config
REPL.TerminalMenus.MultiSelectConfig
REPL.TerminalMenus.config
# -
# #### User interaction
# + attributes={"classes": ["@docs"], "id": ""}
REPL.TerminalMenus.request
# -
# #### AbstractMenu extension interface
#
# Any subtype of `AbstractMenu` must be mutable, and must contain the fields `pagesize::Int` and
# `pageoffset::Int`.
# Any subtype must also implement the following functions:
# + attributes={"classes": ["@docs"], "id": ""}
REPL.TerminalMenus.pick
REPL.TerminalMenus.cancel
REPL.TerminalMenus.writeline
# -
# It must also implement either `options` or `numoptions`:
# + attributes={"classes": ["@docs"], "id": ""}
REPL.TerminalMenus.options
REPL.TerminalMenus.numoptions
# -
# If the subtype does not have a field named `selected`, it must also implement
# + attributes={"classes": ["@docs"], "id": ""}
REPL.TerminalMenus.selected
# -
# The following are optional but can allow additional customization:
# + attributes={"classes": ["@docs"], "id": ""}
REPL.TerminalMenus.header
REPL.TerminalMenus.keypress
| zh_CN_Jupyter_Learn/stdlib/REPL/docs/src/index.md.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sproboticworks/ml-course/blob/master/NLP%20Song%20Lyric%20Generation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="tZBzjOrhvJPo" colab_type="text"
# # Import Packages
# + id="ibbqXFpLvCdQ" colab_type="code" colab={}
import tensorflow as tf
from tensorflow.keras.preprocessing.sequence import pad_sequences
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.models import Sequential
from tensorflow.keras.optimizers import Adam
import numpy as np
# + id="o1KY4GQBuN3X" colab_type="code" colab={}
# !wget --no-check-certificate \
# https://storage.googleapis.com/sproboticworks/master/assets/datasets/irish-lyrics-eof.txt \
# -O /tmp/irish-lyrics-eof.txt
# + [markdown] id="BD7xUHw8vMvT" colab_type="text"
# # Tokenize Data
# + id="W-hsFCjAvTQM" colab_type="code" colab={}
tokenizer = Tokenizer()
data = open('/tmp/irish-lyrics-eof.txt').read()
corpus = data.lower().split("\n")
tokenizer.fit_on_texts(corpus)
total_words = len(tokenizer.word_index) + 1
print(tokenizer.word_index)
print('Total Words: {}'.format(total_words))
# + [markdown] id="YpH9dNcmvXRW" colab_type="text"
# # Prepare Data
# + id="ZayyDMJFvZUk" colab_type="code" colab={}
input_sequences = []
for index, line in enumerate(corpus):
token_list = tokenizer.texts_to_sequences([line])[0]
for i in range(1, len(token_list)):
n_gram_sequence = token_list[:i+1]
input_sequences.append(n_gram_sequence)
if index == 0:
print("'"+line+"' => {}".format(token_list))
print("Input Sequences :")
print('\n'.join(map(str, input_sequences)))
# + id="BzU4pSJP4W4k" colab_type="code" colab={}
# pad sequences
max_sequence_len = max([len(x) for x in input_sequences])
input_sequences = np.array(pad_sequences(input_sequences, maxlen=max_sequence_len, padding='pre'))
first_line_len = len(corpus[0].split())
print("Padded Input Sequences for the first line :")
print(input_sequences[:first_line_len-1])
# + id="jcC-yHOs4Yx6" colab_type="code" colab={}
xs = input_sequences[:,:-1]
labels = input_sequences[:,-1]
ys = tf.keras.utils.to_categorical(labels, num_classes=total_words)
# + id="q-2SO23hv9g4" colab_type="code" colab={}
#Consider first sentence
print("Sentence : {}".format(corpus[0]))
print("Tokens : {}".format(input_sequences[first_line_len-2]))
print("X : {}".format(xs[first_line_len-2]))
print("Label : {}".format(labels[first_line_len-2]))
print("Y : {}".format(ys[first_line_len-2]))
# + [markdown] id="nf332STewG8s" colab_type="text"
# #Build Model
# + id="M6zioxu1wIY8" colab_type="code" colab={}
embedding_dim = 100
model = tf.keras.Sequential([
tf.keras.layers.Embedding(total_words, embedding_dim, input_length=max_sequence_len-1),
tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(150)),
tf.keras.layers.Dense(total_words, activation='softmax')
])
# + id="fLA0enWGwock" colab_type="code" colab={}
model.compile(loss='categorical_crossentropy', optimizer=tf.keras.optimizers.Adam(lr=0.01), metrics=['accuracy'])
# + [markdown] id="mjNT2-3two8f" colab_type="text"
# # Train Model
# + id="UlLfTvISwrlk" colab_type="code" colab={}
history = model.fit(xs, ys, epochs=100)
# + [markdown] id="R6AD8p2wwuMT" colab_type="text"
# # Visualize Training Results
# + id="KF1LbAz5wwEz" colab_type="code" colab={}
import matplotlib.pyplot as plt
def plot_graphs(history, string):
plt.plot(history.history[string])
plt.xlabel("Epochs")
plt.ylabel(string)
plt.show()
# + id="lAkGjD6jwx1T" colab_type="code" colab={}
plot_graphs(history, 'accuracy')
# + [markdown] id="eDdWSAHdw0DE" colab_type="text"
# #Generate Text
# + id="FRgFlKHUw2L8" colab_type="code" colab={}
seed_text = "With great power comes great responsibility"
next_words = 100
for _ in range(next_words):
token_list = tokenizer.texts_to_sequences([seed_text])[0]
token_list = pad_sequences([token_list], maxlen=max_sequence_len-1, padding='pre')
predicted_labels = model.predict(token_list, verbose=0)
predicted_index = np.argmax(predicted_labels, axis=-1)
output_word = ""
for word, index in tokenizer.word_index.items():
if index == predicted_index:
output_word = word
break
seed_text += " " + output_word
print(seed_text)
| NLP Song Lyric Generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.7 64-bit (''caco3'': conda)'
# name: python3
# ---
# Include more cores (LV28-44-3, LV29-114-3, and SO178-12-3). Basically they have higher TOC and lower Carbonates measurements. S0178-12-3 only has TOC measurements.
# +
import numpy as np
import pandas as pd
import glob
import matplotlib.pyplot as plt
#plt.style.use('ggplot')
plt.style.use('seaborn-colorblind')
plt.style.use('dark_background')
plt.rcParams['figure.dpi'] = 300
plt.rcParams['savefig.dpi'] = 300
plt.rcParams['savefig.bbox'] = 'tight'
plt.rcParams['savefig.transparent'] = True
# %matplotlib inline
import datetime
date = datetime.datetime.now().strftime('%Y%m%d')
# -
# # Read and build spectral datasets
# ## The cores having same spe format as the previous cores
# + jupyter={"source_hidden": true} tags=[]
file_name = []
spe_all = []
depth_all = []
cps_all = []
core_all = []
s_depth_all = []
# do it core by core
for core in ['data/LV29-114-3', 'data/SO178-12-3']:
# only read the 10kV which having better signal to the light elements
spe_dir = glob.glob('{}/Run 1 at 10kV/*.spe'.format(core))
# make sure the order follows the depthes in filename
spe_dir.sort()
for spe in spe_dir:
check_depth = spe.split()[3].split('_')[-1]
# there are some inconsistencies in nameing...as usual
# 5 and 6 digis means in mm
if len(check_depth) >= 5:
start_depth = int(check_depth)
# 3 and 4 digit means in cm, needs to be multipled to be mm
elif len(check_depth) >= 3:
start_depth = int(check_depth) * 10
file_name.append(spe.split('/')[-1])
with open(spe, 'r') as f:
content = []
lines = f.readlines()
for line in lines[49:]:
content = np.hstack((content, line.split()))
section_depth = int(lines[13][:-3])
spe_all.append(content.astype(int))
cps_all.append(int(lines[28]))
core_all.append(core[5:])
s_depth_all.append(section_depth)
depth_all.append(section_depth + start_depth)
print('core {} is done.'.format(core))
# -
# ## LV28-44-3
# The spe format of the core LV28-44-3 is different to the previous cores so the codes to catch information need to be modified.<br>
# 1. No X_Position: this value was used for section depth. I adopt the value from the file name instead.
# 1. No TotalCPS: I simply use 0 as the value. In future, if we need cps (so far we don't use it), the values from this core should be detected since no CPS should be 0.
# 1. The channels' values start from line 22 instead of 49.
# +
core = 'data/LV28-44-3'
# only read the 10kV which having better signal to the light elements
spe_dir = glob.glob('{}/Run 1 at 10kV/*.spe'.format(core))
# make sure the order follows the depthes in filename
spe_dir.sort()
for spe in spe_dir:
check_depth = spe.split()[3].split('_')[-1]
# there are some inconsistencies in nameing...as usual
# 5 and 6 digis means in mm
if len(check_depth) >= 5:
start_depth = int(check_depth)
# 3 and 4 digit means in cm, needs to be multipled to be mm
elif len(check_depth) >= 3:
start_depth = int(check_depth) * 10
file_name.append(spe.split('/')[-1])
with open(spe, 'r') as f:
content = []
lines = f.readlines()
for line in lines[22:]:
content = np.hstack((content, line.split()))
section_depth = round(float(spe.split()[4][:-2]))
spe_all.append(content.astype(int))
cps_all.append(0)
core_all.append(core[5:])
s_depth_all.append(section_depth)
depth_all.append(section_depth + start_depth)
# -
spe_df = pd.DataFrame(spe_all, columns = [str(_) for _ in range(2048)])
spe_df['cps'] = cps_all
spe_df['core'] = core_all
spe_df['composite_depth_mm'] = depth_all
spe_df['section_depth_mm'] = s_depth_all
spe_df['filename'] = file_name
spe_df
spe_df[spe_df.isnull().any(axis=1)]
# ## Build composite_id
spe_df.composite_depth_mm.max()
# +
composite_id = []
for core, depth in zip(spe_df.core, spe_df.composite_depth_mm):
composite_id.append('{}_{:05}'.format(core, depth))
spe_df['composite_id'] = composite_id
# -
# ## Drop duplicates
clean_df = spe_df.drop_duplicates('composite_id', keep = 'last')
len(clean_df)
# ### Check those duplicates
spe_df.loc[spe_df.composite_id.duplicated(keep = 'last'), spe_df.columns[-6:]]
# Just some overlaps at section edges. I would simply delete them.
# ## Build section
clean_df = clean_df.set_index('composite_id')
clean_df[clean_df.section_depth_mm == 0]
# No section depth start from 0.
# +
section_all = []
# make sure the order follows the core and composite depth
clean_df.sort_values(by = 'composite_id', axis = 0, inplace = True)
for core in np.unique(clean_df.core):
# I assume every core scanned from section 0 so the first section in the core is marked as section 0
# the deeper the larger number
section = 0
X = clean_df.loc[clean_df.core == core, 'section_depth_mm']
for i in range(len(X)):
section_all.append(section)
try:
# when section changes, the section depth should be rest to smaller number
if X[i] > X[i + 1]:
section += 1
except IndexError:
print('bottom of the core {}'.format(core))
clean_df['section'] = section_all
# -
clean_df
clean_df.to_csv('data/spe_dataset_{}.csv'.format(date))
# ## Read bulk chemistry
bulk_28_df = pd.read_excel('data/Bulk chem/LV28-44-3_TCN.xlsx')
bulk_28_df
bulk_28_df.isna().any()
bulk_29_df = pd.read_excel('data/Bulk chem/LV29 114-3_TOC%_CaCO3%LMax.xls')
bulk_29_df
bulk_29_df = bulk_29_df.dropna(axis=0)
bulk_29_df
# Drop out 16 rows having null values.
bulk_178_df = pd.read_table('data/Bulk chem/SO178-12-3_TOC.txt', header=0, usecols=range(3))
bulk_178_df
bulk_178_df.isna().any()
# ## Merge three cores' bulk chemistry
# The depths in LV29-114-3 are all integer instead of XX.5 like previous cores which use mid depth. I assume it's a mistake so add 0.5 to the depths.
depth = np.hstack((bulk_28_df['Depth'].values, bulk_29_df['Depth (cm)']+.5, bulk_178_df['Teufe(cm)']))
toc = np.hstack((bulk_28_df['TOC (wt. %)'].values, bulk_29_df['TOC(%)'], bulk_178_df['Kohlenstoff(%)']))
# SO178-12-3 doesn't have CaCO3 so I simply asign np.NaN
caco3 = np.hstack((bulk_28_df['CaCO3 (wt. %)'].values, bulk_29_df['CaCO3 (%)'], [np.NaN for _ in range(len(bulk_178_df))]))
core = np.hstack((['LV28-44-3' for _ in range(len(bulk_28_df))], ['LV29-114-3' for _ in range(len(bulk_29_df))], ['SO178-12-3' for _ in range(len(bulk_178_df))]))
print(len(depth), len(toc), len(caco3), len(core))
bulk_df = pd.DataFrame({'mid_depth_mm': depth*10,
'TOC%': toc,
'CaCO3%': caco3,
'core': core
})
bulk_df
bulk_df.to_csv('data/bulk_dataset_{}.csv'.format(date))
# # Combine the dataset to the previous datasets
# The new cores all lack TC and core SO178-12-3 lacks CaCO3 also.
bulk_df = pd.read_csv('data/bulk_dataset_20201215.csv', index_col=0)
clean_df = pd.read_csv('data/spe_dataset_20201215.csv', index_col=0)
bulk_p_df = pd.read_csv('data/bulk_dataset_20201007.csv', index_col=0)
spe_p_df = pd.read_csv('data/spe_dataset_20201008.csv', index_col=0)
print(bulk_df.shape, clean_df.shape)
print(bulk_p_df.shape, spe_p_df.shape)
bulk_c_df = pd.concat([bulk_p_df, bulk_df], axis=0, join='outer')
spe_c_df = pd.concat([spe_p_df, clean_df], axis=0, join='outer')
print(bulk_c_df.shape, spe_c_df.shape)
bulk_c_df
# # Merge spe and bulk datasets
# +
mask_c = spe_c_df.columns[:2048] # only the channels
merge_df = pd.DataFrame()
for index, row in bulk_c_df.iterrows():
mid = row['mid_depth_mm']
core = row['core']
# get the spe in 10 mm interval
mask_r = (spe_c_df.composite_depth_mm >= (mid-5)) & (spe_c_df.composite_depth_mm <= (mid+5)) & (spe_c_df.core == core)
merge_df = pd.concat(
[merge_df, spe_c_df.loc[mask_r, mask_c].apply(np.mean, axis = 0).append(row)],
axis = 1
)
merge_df = merge_df.T.reset_index(drop = True)
# -
merge_df
# ### Check rows having nan in any column
merge_df[merge_df.isnull().any(axis = 1)]
# ### Check rows having nan in spetra
# They mean the data points has bulk measurements but without XRF measurement.
merge_df[merge_df.iloc[:, :2048].isnull().any(axis = 1)]
merge_df[~merge_df.iloc[:, :2048].isnull().any(axis = 1)]
# Comparing to the previous merged dataset (382), the updated dataset has 317 more data points.
# ## Export dataset
# This dataset combines the preivous and updated merged datasets. The data points having no TC or CaCO3 measurements are still kept.
merge_df[~merge_df.iloc[:, :2048].isnull().any(axis = 1)].to_csv('data/spe+bulk_dataset_{}.csv'.format(date))
# The datasets should include previous data, not only the data compiled this time
bulk_c_df.to_csv('data/bulk_dataset_20201215.csv')
spe_c_df.to_csv('data/spe_dataset_20201215.csv')
| build_database_05.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Pandas Data Structures
#
# * Series
# * Dataframe
# * Panel
import pandas as pd
import numpy as np
# ## Series
# ### Series Creation
np.random.seed(100)
data = np.random.rand(7)
ser = pd.Series(data)
ser
# #### Create a Series Structure of first 5 months of the year with a specified index of month names:
import calendar as cal
monthName = [cal.month_name[i] for i in np.arange(1,13)]
months = pd.Series(np.arange(1,13),index = monthName)
months
# #### Index of Series
#
months.index
# #### Series using Python Dictionary
currDict = {'US' :'dollar','UK' : 'pound','Germany': 'euro',
'Mexico' : 'peso','Nigeria':'Naira','China':'yuan','Japan':'yen'}
currDict
currSeries =pd.Series(currDict)
currSeries
# ### Operations on Series
currDict['China']
# #### Assignment Operation
currDict['China']='Yuan'
currDict
currDict.get('UK')
# #### Slicing
currSeries[:2]
currVal = {'US' :73,'UK' : 103,'Germany': 80,
'Mexico' : 4,'Nigeria':0.003,'China':12,'Japan':0.3}
currVal = pd.Series(currVal)
currVal
np.mean(currVal)
np.std(currVal)
currVal*currVal
np.sqrt(currVal)
# ### Slicing in Series
currVal[1:]
currVal[currVal>100]
currVal[1:]+currVal[:-2]
# ## DateFrame 2-D labeled array
# ### Using Dictionaries of Series
stockSummaries = {'AMZN':pd.Series([346.15,0.59,459,0.52,589.8,158.88],index =[
'Closing Price','EPS','Shares Outstanding(M)','Beta','P/E','Market Cap(B)']),
'GOOG': pd.Series([1133.43,36.05,335.83,0.87,31.44,380.64],
index=['Closing price','EPS','Shares Outstanding(M)',
'Beta','P/E','Market Cap(B)']),'FB': pd.Series([61.48,0.59,2450,104.93,150.92],
index=['Closing price','EPS','Shares Outstanding(M)',
'P/E', 'Market Cap(B)']),
'YHOO': pd.Series([34.90,1.27,1010,27.48,0.66,35.36],
index=['Closing price','EPS','Shares Outstanding(M)',
'P/E','Beta', 'Market Cap(B)']),
'TWTR':pd.Series([65.25,-0.3,555.2,36.23],
index=['Closing price','EPS','Shares Outstanding(M)',
'Market Cap(B)']),
'AAPL':pd.Series([501.53,40.32,892.45,12.44,447.59,0.84],
index=['Closing price','EPS','Shares Outstanding(M)','P/E',
'Market Cap(B)','Beta'])}
stockSummaries
stockDF =pd.DataFrame(stockSummaries)
stockDF
stockDF =pd.DataFrame(stockSummaries,index =['Closing price','EPS','Shares Outstanding (M)',
'P/E','Market Cap(B)','Beta'])
stockDF
stockDF_1 =pd.DataFrame(stockSummaries,index =['Closing price','EPS','Shares Outstanding (M)',
'P/E','Market Cap(B)','Beta'],columns =['FB','TWTR','SCNW'])
stockDF_1
stockDF.index
stockDF.columns
# ### Using a Dictionary of ndarrays/lists
algos={'search':['DFS','BFS','Binary Search',
'Linear','ShortestPath (Djikstra)'],
'sorting': ['Quicksort','Mergesort', 'Heapsort',
'Bubble Sort', 'Insertion Sort'],
'machine learning':['RandomForest',
'K Nearest Neighbor',
'Logistic Regression',
'K-Means Clustering',
'Linear Regression']}
algoDF =pd.DataFrame(algos)
algoDF
# #### Defining Index
pd.DataFrame(algos,index =['algo_1','algo_2','algo_3','algo_4','algo_5'])
# ### From a Sturctured array
# +
memberData = np.zeros((4,),dtype=[('Name','<U15'), ('Age','i4'), ('Weight','f2')])
memberData[:] = [('Sanjeev',37,162.4),('Yingluck',45,137.8),
('Emeka',28,153.2),
('Amy',67,101.3)]
memberDF =pd.DataFrame(memberData)
memberDF
# -
pd.DataFrame(memberData,index =['a','b','c','d'])
currSeries.name ='currency'
pd.DataFrame(currSeries)
# ### Operations on DataFrame
memberDF
# #### Assignment Operation
memberDF['Height']=60
memberDF
# #### Deletion Operation
del memberDF['isSenior'];
memberDF
# #### Column Inserted at the end; to insert a column at a specific location
memberDF.insert(2,'isSenior',memberDF['Age']>60)
memberDF
# #### Alignment
ore1DF=pd.DataFrame(np.array([[20,35,25,20],
[11,28,32,29]]),
columns=['iron','magnesium',
'copper','silver'])
ore2DF=pd.DataFrame(np.array([[14,34,26,26],
[33,19,25,23]]),
columns=['iron','magnesium',
'gold','silver'])
ore1DF
ore2DF
ore2DF + ore1DF
# #### Other mathematical operations
np.sqrt(ore2DF),np.sqrt(ore1DF)
| Pandas_Data_Structures_Notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
from sklearn.model_selection import train_test_split
from sklearn.linear_model import LinearRegression, Ridge, Lasso
import seaborn as sns
import matplotlib as plt
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.base import BaseEstimator, TransformerMixin
from sklearn.pipeline import Pipeline, make_union
from sklearn.metrics import mean_squared_error
import math
# Regularization is a technique which makes a model generalize better therefore preventing overfitting.
#
# * L1 regularization = Lasso Regression
# * L2 regularization = Ridge Regression
#
# Unlike L2 that reduces the coefficient to a value close to but not equal to 0, L1 shrinks the less important feature’s coefficient to 0 thus, removing some features. So, this works well for feature selection in case we have a huge number of features.
data = pd.read_csv('Weather.csv')
final_data = data.fillna(0)
yVar = final_data['MaxTemp'].values.reshape(-1,1)
xVar = final_data['MinTemp'].values.reshape(-1,1)
X_train, X_test, y_train, y_test = train_test_split(xVar, yVar, test_size=0.2, random_state=0)
linReg = LinearRegression()
linReg.fit(X_train, y_train)
# +
y_pred = linReg.predict(X_test)
mse = mean_squared_error(y_test, y_pred)
rmse = math.sqrt(mse)
print('RMSE: {}'.format(rmse))
print('Test score: {}'.format(linReg.score(X_test, y_test)))
# +
steps = [
('model', Ridge(alpha=10, fit_intercept=True))
]
ridge_pipe = Pipeline(steps)
ridge_pipe.fit(X_train, y_train)
print('Test Score: {}'.format(ridge_pipe.score(X_test, y_test)))
y_pred_ridge = ridge_pipe.predict(X_test)
mse = mean_squared_error(y_test, y_pred_ridge)
rmse = math.sqrt(mse)
print('RMSE: {}'.format(rmse))
# +
steps = [
('model', Lasso(alpha=0.9, fit_intercept=True))
]
lasso_pipe = Pipeline(steps)
lasso_pipe.fit(X_train, y_train)
print('Test score: {}'.format(lasso_pipe.score(X_test, y_test)))
y_pred_lasso = lasso_pipe.predict(X_test)
mse = mean_squared_error(y_test, y_pred_lasso)
rmse = math.sqrt(mse)
print('RMSE: {}'.format(rmse))
# -
| Section 7/7.5.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import xgboost as xgb
import sklearn.datasets
import sklearn.metrics
import sklearn.model_selection
from bayeso import bo
from bayeso.wrappers import wrappers_bo
from bayeso.utils import utils_plotting
from bayeso.utils import utils_bo
# +
digits = sklearn.datasets.load_digits()
data_digits = digits.images
data_digits = np.reshape(data_digits,
(data_digits.shape[0], data_digits.shape[1] * data_digits.shape[2]))
labels_digits = digits.target
data_train, data_test, labels_train, labels_test = sklearn.model_selection.train_test_split(
data_digits, labels_digits, test_size=0.3, stratify=labels_digits)
# -
def fun_target(bx):
model_xgb = xgb.XGBClassifier(
max_depth=int(bx[0]),
n_estimators=int(bx[1]),
use_label_encoder=False
)
model_xgb.fit(data_train, labels_train, eval_metric='mlogloss')
preds_test = model_xgb.predict(data_test)
return 1.0 - sklearn.metrics.accuracy_score(labels_test, preds_test)
# +
str_fun = 'xgboost'
# (max_depth, n_estimators)
bounds = np.array([[1, 10], [100, 500]])
num_bo = 5
num_iter = 25
num_init = 1
# +
model_bo = bo.BO(bounds, debug=False)
list_X = []
list_Y = []
list_time = []
for ind_bo in range(0, num_bo):
print('BO Round:', ind_bo + 1)
X_final, Y_final, time_final, _, _ = wrappers_bo.run_single_round(
model_bo, fun_target, num_init, num_iter,
seed=42 * ind_bo)
list_X.append(X_final)
list_Y.append(Y_final)
list_time.append(time_final)
arr_X = np.array(list_X)
arr_Y = np.array(list_Y)
arr_time = np.array(list_time)
arr_Y = np.expand_dims(np.squeeze(arr_Y), axis=0)
arr_time = np.expand_dims(arr_time, axis=0)
# -
for ind_bo in range(0, num_bo):
bx_best, y_best = utils_bo.get_best_acquisition_by_history(arr_X[ind_bo],
arr_Y[0, ind_bo][..., np.newaxis])
print('BO Round', ind_bo + 1)
print(bx_best, y_best)
utils_plotting.plot_minimum_vs_iter(arr_Y, [str_fun], num_init, True,
str_x_axis='Iteration',
str_y_axis='1 - Accuracy')
utils_plotting.plot_minimum_vs_time(arr_time, arr_Y, [str_fun], num_init, True,
str_x_axis='Time (sec.)',
str_y_axis='1 - Accuracy')
| src/example_hpo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.10 64-bit (''parcels-dev'': conda)'
# name: python3
# ---
# ### Salish Sea Oil Spill Scenarios
#
# This notebook describes oil particle tracking for spill scenarios in the Salish Sea developed in conjunction with the Canadian Department of Fisheries and Oceans (DFO). The code is for translating particle tracking from *ocean parcels* using the *Salish Sea Cast* grid into input forcing files for *Atlantis* and resulting in oil dispersal on the *Salish Sea Atlantis Model* box grid.
import sys
import os
import math
import xarray as xr
import geopandas as gpd
import pandas as pd
import matplotlib.cm as cm
import matplotlib.pyplot as plt
import matplotlib.colors as colors
from IPython.display import Image
from netCDF4 import Dataset
from shapely.geometry import Point
from pathlib import Path
from pprint import pprint
from parcels import AdvectionRK4, VectorField, Variable
from parcels import FieldSet, plotTrajectoriesFile, Variable, ScipyParticle, Field
import numpy as np
from datetime import timedelta
sys.path.append('/ocean/rlovindeer/Atlantis/ssam_oceanparcels/Parcels_Utils/particle_tracking/parcels/')
from util.seed_particles import get_particles, get_release_times
# Data Paths
currents = Path('/ocean/rlovindeer/Atlantis/Physics/Raw_Transport_Data/')
winds = Path('/ocean/rlovindeer/Atlantis/Physics/Wind/')
sea_grid = Path('/ocean/rlovindeer/Atlantis/Physics/Grids/ubcSSnBathymetryV17-02_a29d_efc9_4047.nc')
air_grid = Path('/ocean/rlovindeer/Atlantis/Physics/Grids/ubcSSaAtmosphereGridV1_0f03_6268_df4b.nc')
fraser_discharge = Path('/data/dlatorne/SOG-projects/SOG-forcing/ECget/Fraser_flow')
tides = '/ocean/rlovindeer/MOAD/analysis-raisha/notebooks/contaminant-dispersal/results/Tides/'
# ### Spill Scenarios
# Scenarios are based on discussion with DFO & Transport Canada. See [this documented](https://docs.google.com/spreadsheets/d/17HgaXoKG5b0zkigri6Vdw7fNviDKVzzTVrbpVVRrCjk/edit?usp=sharing) for full scenario descriptions, including spill locations, and type and volume of spill.
# Spill release times
release_start = '2020-07-01' ## winter starts in December, Summer in Jul - Aug
release_end = '2020-07-02'
release_YYYYMM = '2020-07'
spill_volume = 2e6 # L
# +
# Oil type properties & spill location selection
bitumen = {
"Weight": 1021.9, # g/L
"Naphthalene": 0.024, # mg/g oil
"Phenanthrene": 0.017,
"Pyrene": 0.010,
"Benzo": 0.003,
}
BunkerC = {
"Weight": 1017.8,
"Naphthalene": 0.680,
"Phenanthrene": 0.796,
"Pyrene": 0.266,
"Benzo": 0.056,
}
Diesel = {
"Weight": 841.6,
"Naphthalene": 3.664,
"Phenanthrene": 1.000,
"Pyrene": 0.000,
"Benzo": 0.000,
}
Crude = {
"Weight": 933.6,
"Naphthalene": 0.654,
"Phenanthrene": 0.327,
"Pyrene": 0.013,
"Benzo": 0.002,
}
fuel_type = {
"bitumen" : bitumen,
"BunkerC" : BunkerC,
"Diesel" : Diesel,
"Crude" : Crude,
}
file_id = int(input( ))
scenario = {1 : "5b_Turn_Point_Diluted_bitumen",
2 : "6a_VancouverHarbour_BunkerC",
3 : "7a_JohnsonStrait_BunkerC",
4 : "4a_ActivePass_Diesel",}
print("\nScenario running :", scenario[file_id], sep = " ")
# -
# ## Creating particle movement through Ocean Parcels
# +
#Kernels
def WindAdvectionRK4(particle, fieldset, time):
"""Advection of particles using fourth-order Runge-Kutta integration.
Function needs to be converted to Kernel object before execution"""
if particle.beached == 0:
wp = fieldset.wind_percentage ## this needs to be added to the fieldset
if wp > 0:
(u1, v1) = fieldset.UV[time, particle.depth, particle.lat, particle.lon]
u1 = u1 * wp
v1 = v1 * wp
lon1, lat1 = (particle.lon + u1*.5*particle.dt, particle.lat + v1*.5*particle.dt)
(u2, v2) = fieldset.UVwind[time + .5 * particle.dt, particle.depth, lat1, lon1]
u2 = u2 * wp
v2 = v2 * wp
lon2, lat2 = (particle.lon + u2*.5*particle.dt, particle.lat + v2*.5*particle.dt)
(u3, v3) = fieldset.UVwind[time + .5 * particle.dt, particle.depth, lat2, lon2]
u3 = u3 * wp
v3 = v3 * wp
lon3, lat3 = (particle.lon + u3*particle.dt, particle.lat + v3*particle.dt)
(u4, v4) = fieldset.UVwind[time + particle.dt, particle.depth, lat3, lon3]
u4 = u4 * wp
v4 = v4 * wp
u_wind = (u1 + 2*u2 + 2*u3 + u4) / 6. * particle.dt
v_wind = (v1 + 2*v2 + 2*v3 + v4) / 6. * particle.dt
particle.lon += (u1 + 2*u2 + 2*u3 + u4) / 6. * particle.dt
particle.lat += (v1 + 2*v2 + 2*v3 + v4) / 6. * particle.dt
particle.beached = 2
def BeachTesting(particle, fieldset, time):
""" Testing if particles are on land. if 'yes' particle will be removed"""
if particle.beached == 2:
(u, v) = fieldset.UV[time, particle.depth, particle.lat, particle.lon]
#print(u, v)
if u == 0 and v == 0:
particle.beached = 1
else:
particle.beached = 0
def DeleteParticle(particle, fieldset, time):
particle.delete()
def DecayParticle(particle, fieldset, time):
dt = particle.dt
field_decay_value = fieldset.decay
decay = math.exp(0 * dt/field_decay_value) #math.exp(-1.0 * dt/field_decay_value)
particle.decay_value = particle.decay_value * decay
# +
# Salish Sea NEMO Model Grid, Geo-location and Bathymetry, v17-02
# Currents
u_current = sorted([p for p in currents.glob(str(release_YYYYMM) + '*URaw_variables.nc')])
v_current = sorted([p for p in currents.glob(str(release_YYYYMM) + '*VRaw_variables.nc')])
filenames = {
'U': {'lon': sea_grid,'lat': sea_grid,'data': u_current},
'V': {'lon': sea_grid,'lat': sea_grid,'data': v_current}
}
variables = {'U': 'uVelocity','V': 'vVelocity'}
dimensions = {'lon': 'longitude', 'lat': 'latitude', 'time': 'time'}
fieldset = FieldSet.from_nemo(filenames, variables, dimensions, allow_time_extrapolation=True);
print('fieldset created from_nemo')
fieldset.add_constant('decay', 1.0 * 3600.0);
print('decay constant added')
# +
# HRDPS, Salish Sea, Atmospheric Forcing Grid, Geo-location, v1"
wind_paths = sorted([p for p in winds.glob(str(release_YYYYMM) + '*Wind_variables.nc')])
wind_filenames = {'lon': os.fspath(air_grid),'lat': os.fspath(air_grid),'data': wind_paths}
wind_dimensions = {'lon': 'longitude', 'lat': 'latitude', 'time': 'time'}
pprint(wind_filenames)
# +
Uwind_field = Field.from_netcdf(wind_filenames, ('U_wind', 'u_wind'),
wind_dimensions,
fieldtype='U',
allow_time_extrapolation=True,
transpose=False,
deferred_load=False)
Vwind_field = Field.from_netcdf(wind_filenames, ('V_wind', 'v_wind'),
wind_dimensions,
fieldtype='V',
allow_time_extrapolation=True,
transpose=False,
deferred_load=False)
print('wind data loaded')
# +
# change longitude for the wind field
Uwind_field.grid.lon = Uwind_field.grid.lon - 360
Vwind_field.grid.lon = Vwind_field.grid.lon - 360
[x_min, x_max, y_min, y_max] = Uwind_field.grid.lonlat_minmax
Uwind_field.grid.lonlat_minmax = [x_min - 360, x_max - 360, y_min, y_max]
Vwind_field.grid.lonlat_minmax = [x_min - 360, x_max - 360, y_min, y_max]
## adding the wind field to the fieldset object
fieldset.add_field(Uwind_field)
fieldset.add_field(Vwind_field)
wind_field = VectorField('UVwind', Uwind_field, Vwind_field)
fieldset.add_vector_field(wind_field)
# -
# wind_percentage
# We need to do a sensitivity analysis of the percetage of wind to be used here
wind_percentage = 1
fieldset.add_constant('wind_percentage', wind_percentage/100.0)
# + active=""
# Just in case we want to add a maximum age
# # fieldset_sum.add_constant('max_age', dispersal_length)
# +
class MyParticle(ScipyParticle):
initial_time = -100
decay_value = Variable('decay_value', dtype=np.float32, initial=1.0)
beached = Variable('beached', dtype=np.int32, initial=0.)
age = Variable('age', dtype=np.int32, initial=0.)
# Particle Features
num_particles_per_day = 100
feature_release_index = 0
input_shapefile_name = "/ocean/rlovindeer/Atlantis/ssam_oceanparcels/SalishSea/Shape_Scenarios/" + scenario[file_id] + ".shp"
release_depth = -0.1
release_start_time = np.datetime64(release_start)
release_end_time = np.datetime64(release_end)
time_origin = fieldset.U.grid.time_origin.time_origin
print('setting up particles')
# -
# ### Salish Sea Conditions during the scenario run
# Tide data is taken from [DFO-Pacific website](https://www.pac.dfo-mpo.gc.ca/science/charts-cartes/obs-app/observed-eng.aspx?StationID=07735) at the nearest tide gauge.
# Surface winds are taken from hourly atmospheric field values from the Environment Canada High Resolution Deterministic Prediction System (HRDPS) atmospheric forcing model, and used to force the surface winds for Ocean Parcels.
# Discharge from the Fraser River (m^3) is acquired from Salish Sea Cast.
# Tides
Tide_location = 'SandyCove_'
tide_filename = tides + 'Tides_' + Tide_location + str(release_start) +'.png'
Image(filename= tide_filename)
# # Winds
# place = 'Sand Heads'
# #lat_lon = places.PLACES[place]['GEM2.5 grid ji']
# #lat_lon
#
# surface_wind = winds.glob(str(release_start) + '_Wind_variables.nc')
# print(surface_wind)
#
# wind_velocity = xr.open_dataset(surface_wind)
#
# uvelocity = wind_velocity.u_wind.isel(gridX=135, gridY=151).data
# vvelocity = wind_velocity.v_wind.isel(gridX=135, gridY=151).data
# wind_speed, winds = wind_tools.wind_speed_dir(uvelocity, vvelocity)
#
# fig, ax = plt.subplots(1,1, figsize = (14,6))
#
# ax.plot(wind_velocity.time,wind_speed, color = 'darkblue', linewidth = 2)
# ax.set_title('wind speed at '+place+' (m/s)', fontsize = 12)
# ax.tick_params(labelsize=12)
# +
# Fraser River Discharge
# format is YYYY MM DD m^3
df = pd.read_csv(fraser_discharge, skiprows=35000, names=['date_flow'])
df = pd.DataFrame(df.date_flow.str.split(' ',3).tolist(),columns = ['Y','M','D','flow_m^3'])
df1 = df.loc[df['Y'] == '2020']
df2 = df1.loc[df['M'] == '07']
df3 = df2.loc[df['D']== '01']
df3
# +
[release_times, p, num_particles] = get_release_times(time_origin, num_particles_per_day, release_start_time, release_end_time);
pset = get_particles(fieldset, num_particles, input_shapefile_name, MyParticle, feature_release_index, release_times, release_depth);
#print(pset)
# Building the kernels
decay_kernel = pset.Kernel(DecayParticle);
beaching_kernel = pset.Kernel(BeachTesting);
ForcingWind_kernel = pset.Kernel(WindAdvectionRK4);
# Adding to the main kernel
my_kernel = AdvectionRK4 + decay_kernel + ForcingWind_kernel + beaching_kernel;
output_file_name = scenario[file_id] + str(release_start_time) + '_OP.nc'
print(output_file_name)
# +
try:
os.system('rm ' + output_file_name)
except:
pass
print('executing particle kernel')
# +
## Output properties
output_file = pset.ParticleFile(name= output_file_name, outputdt = timedelta(minutes = 60))
pset.execute(my_kernel, # the kernel (which defines how particles move)
runtime=timedelta(hours = 24*6), # total length of the run
dt = timedelta(minutes = 60), # timestep of the kernel
output_file = output_file) # file name and the time step of the outputs
output_file.close()
plotTrajectoriesFile(output_file_name);
print('particle trajectories completed')
# -
# ## Parcing Ocean Parcels output into Atlantis input files
#
SalishSea_shapefile = "/ocean/rlovindeer/Atlantis/ssam_oceanparcels/SalishSea/SalishSea_July172019_2/SalishSea_July172019.shp"
data_df = gpd.read_file(SalishSea_shapefile)
# +
numLayers = 7;
numSites = data_df.shape[0]
numTargetSites = numSites
outputDT = 60*60*12 #12 hours
stepsPerDay = int(86400.0/ outputDT);
numStepsPerDT = stepsPerDay;
numStepsPerDT = int(outputDT/3600.0)
debug = False
inputFileName = output_file_name
pfile = xr.open_dataset(str(inputFileName), decode_cf=True)
# +
lon = np.ma.filled(pfile.variables['lon'], np.nan)
lat = np.ma.filled(pfile.variables['lat'], np.nan)
time = np.ma.filled(pfile.variables['time'], np.nan)
z = np.ma.filled(pfile.variables['z'], np.nan)
probs = np.ma.filled(pfile.variables['decay_value'], np.nan)
numParticles = lon.shape[0]
trackDates = [];
for i in range(0,numParticles):
#print(time[i][0])
trackDates.append(time[i][0]);
RDiff = max(trackDates) - min(trackDates);
minDate = np.datetime64(release_start+"T00:30:00");
ts = pd.to_datetime(str(minDate));
d = ts.strftime('%Y-%m-%d %H:%M:%S');
print(d)
# +
numReleaseDays = 1;
numReleaseSteps = numReleaseDays * stepsPerDay;
trackLength = len(lon[0]);
print('trackLength = ' + str(trackLength))
print('numStepsPerDT = ' + str(numStepsPerDT))
numSteps = int(trackLength / numStepsPerDT);
# +
# Create the netcdf output file for Atlantis
netcdfFileName = "Atlantis_" + scenario[file_id] + str(release_start_time) + ".nc"
try:
os.remove(netcdfFileName)
except:
pass
ncfile = Dataset(netcdfFileName, "w", format="NETCDF4", clobber=True)
# Dimensions
time = ncfile.createDimension("t", None)
b = ncfile.createDimension("b", numTargetSites)
z = ncfile.createDimension("z", numLayers)
# Variables
times = ncfile.createVariable("time","f4",("t",))
oil = ncfile.createVariable("oil","f4",("t", "b", "z"))
Naphthalene = ncfile.createVariable("Naphthalene","f4",("t", "b", "z"))
Phenanthrene = ncfile.createVariable("Phenanthrene","f4",("t", "b", "z"))
Pyrene = ncfile.createVariable("Pyrene","f4",("t", "b", "z"))
Benzo = ncfile.createVariable("Benzo","f4",("t", "b", "z"))
# Attributes
times.units = "seconds since 1950-01-01 00:00:00 +10"
times.dt = outputDT
times.long_name = "time"
oil.units = "gOIL/m^3"
oil.long_name = "Concentration of oil"
Naphthalene.units = "mgPAH/m^3"
Naphthalene.long_name = "Naphthalene"
Naphthalene.missing_value = 0.0
Naphthalene.valid_min = 0.0
Naphthalene.valid_max = 10000000000.0
Phenanthrene.units = "mgPAH/m^3"
Phenanthrene.long_name = "Phenanthrene"
Phenanthrene.missing_value = 0.0
Phenanthrene.valid_min = 0.0
Phenanthrene.valid_max = 10000000000.0
Pyrene.units = "mgPAH/m^3"
Pyrene.long_name = "Pyrene"
Pyrene.missing_value = 0.0
Pyrene.valid_min = 0.0
Pyrene.valid_max = 10000000000.0
Benzo.units = "mgPAH/m^3"
Benzo.long_name = "Benzo(a)pyrene"
Benzo.missing_value = 0.0
Benzo.valid_min = 0.0
Benzo.valid_max = 10000000000.0
# Populate variables with data
timeData = np.arange(0,(numSteps + numReleaseSteps)*outputDT,outputDT)
times[:] = timeData;
boxDispersal = np.zeros((numSteps + numReleaseSteps, numTargetSites));
# +
for partIndex in range(0, numParticles):
trackDateDiff = trackDates[partIndex] - minDate;
trackDateDiff = trackDateDiff/ np.timedelta64(1, 's')
timeOffset = int(abs((trackDateDiff /outputDT)));
for stepIndex in range(0, numSteps):
timeValue = stepIndex + timeOffset
partLon = lon[partIndex][stepIndex * numStepsPerDT];
partLat = lat[partIndex][stepIndex * numStepsPerDT];
partProb = probs[partIndex][stepIndex * numStepsPerDT];
matchFound = 0;
for targetIndex in range(0, numTargetSites):
path = data_df.iloc[targetIndex].geometry
checks = path.contains(Point(partLon, partLat));
if checks:
boxDispersal[timeValue][targetIndex] = boxDispersal[timeValue][targetIndex] + partProb;
# uncomment line below to ignore particle decay during debugging.
#boxDispersal[timeValue][targetIndex] = boxDispersal[timeValue][targetIndex] + 1.0
matchFound = 1
if debug:
print('At time ' + str(timeValue) + ' Particle (' + str(partIndex) + ') in box ' + str(data_df.iloc[targetIndex].BOX_ID))
break;
if matchFound == 0:
if debug:
print('No match for particle')
print(partLon, partLat)
#break
oil[:, :, 5] = boxDispersal * fuel_type[scenario[file_id].split(sep = '_')[-1]]["Weight"] * spill_volume;
Naphthalene[:, :, :] = oil[:, :, :] * fuel_type[scenario[file_id].split(sep = '_')[-1]]["Naphthalene"];
Phenanthrene[:, :, :] = oil[:, :, :] * fuel_type[scenario[file_id].split(sep = '_')[-1]]["Phenanthrene"];
Pyrene[:, :, :] = oil[:, :, :] * fuel_type[scenario[file_id].split(sep = '_')[-1]]["Pyrene"];
Benzo[:, :, :] = oil[:, :, :] * fuel_type[scenario[file_id].split(sep = '_')[-1]]["Benzo"];
ncfile.close()
# -
# ## Animating Oil Dispersal Scenario in the Salish Sea Atlantis Model
# +
boxes = data_df['BOTZ']
land_boxes = boxes==0
land_boxes = data_df.index[land_boxes]
numReleaseDays = RDiff
print('numReleaseDays = ' + str(numReleaseDays))
numReleaseDTS = int(abs(numReleaseDays/np.timedelta64(1, 'h')));
totalNumOfTS = int(numReleaseDTS + trackLength);
print('totalNumOfTS = ' + str(totalNumOfTS))
print('trackLength = ' + str(trackLength))
print(numParticles)
# +
trackLonsPadded = np.zeros((int(numParticles), totalNumOfTS));
trackLatsPadded = np.zeros((int(numParticles), totalNumOfTS));
particlesAge = np.zeros((int(numParticles), totalNumOfTS));
for trackIndex in range(0,numParticles):
#print(trackDates[trackIndex])
#print(minDate)
trackDateDiff = trackDates[trackIndex] - minDate
#print(trackDateDiff/np.timedelta64(1, 'h'))
trackNumsToPad = int(trackDateDiff/np.timedelta64(1, 'h'))
#print(trackNumsToPad)
trackLonsPadded[trackIndex][0:trackNumsToPad] = 0;
trackLatsPadded[trackIndex][0:trackNumsToPad] = 0;
trackLonsPadded[trackIndex][trackNumsToPad:trackNumsToPad + trackLength] = lon[:][trackIndex];
trackLatsPadded[trackIndex][trackNumsToPad:trackNumsToPad + trackLength] = lat[:][trackIndex];
numSteps = int(trackLength / numStepsPerDT);
# +
savefile_prefix = 'boxes'
pfile = xr.open_dataset(str(netcdfFileName), decode_cf=True)
print(pfile)
time = np.ma.filled(pfile.variables['time'], np.nan)
oil = np.ma.filled(pfile.variables['Naphthalene'], np.nan)
num_steps = time.shape[0]
#print(num_steps)
# +
# Creating statis figures with a log scale
_cmap = cm.coolwarm
file_names = []
land_df = data_df.loc[land_boxes]
for time_index in range(0, num_steps):
plon = trackLonsPadded[:, time_index]
plat = trackLatsPadded[:, time_index]
plon = plon[plon<0]
plat = plat[plat>0]
time_oil = oil[time_index]
data_df['oil'] = time_oil
data_df.loc[land_boxes, 'oil'] = 1
ax = data_df.plot(figsize=(9, 15), column = 'oil', cmap = _cmap, norm=colors.SymLogNorm(
linthresh=0.000001, linscale=0.000001,
vmin=-0.0001, vmax=1e10, base=10),
legend=True, legend_kwds={'label': "Relative Oil Concentration"
},)
land_df.plot(ax=ax, color='white')
ax.scatter(plon, plat, s=0.0001, color='lightgrey', zorder=20)
ax.set_title(time[time_index])
plot_name = savefile_prefix + '_time_' + str(time_index).zfill(3) + '.png'
plt.savefig(plot_name)
file_names.append(plot_name)
plt.close()
# +
from PIL import Image
import glob
# Create the frames
frames = []
imgs = glob.glob("*.png")
imgs.sort()
for i in imgs:
new_frame = Image.open(i)
frames.append(new_frame)
# Save into loop
anim_name = 'Oil_Scenario_' + scenario[file_id] + str(release_start_time) +'c.gif'
frames[0].save(anim_name, format='GIF',
append_images=frames[1:],
save_all=True,
duration=300, loop=0)
file_name_str = ' '.join(file_names);
os.system('rm ' + file_name_str);
# -
from IPython.display import Image
with open(anim_name,'rb') as anim:
display(Image(anim.read()))
| notebooks/contaminant-dispersal/Spill_Scenarios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # "Updated_map - Indian States"
# > "Test page created using Fast pages and Altair to check functionality and interactivity"
#
# - toc: false
# - branch: master
# - badges: false
# - comments: true
# - categories: [fastpages, jupyter]
# - image: images/some_folder/your_image.png
# - hide: false
# - search_exclude: true
# - metadata_key1: metadata_value1
# - metadata_key2: metadata_value2
#hide
import geopandas as gpd
import json
import altair as alt
#import itables.interactive
import pandas as pd
# +
#hide
from IPython.core.display import HTML
display(HTML("""
<style>
.output {
display: flex;
align-items: center;
text-align: center;
}
</style>
"""))
# -
#hide
gdf = gpd.read_file('states_india.shp')
gdf
#hide
gdf = gdf[['geometry','st_nm']]
gdf.columns = ['geometry','state']
gdf['count'] = 0
gdf['count'][14]=1
gdf['count'][15]=2
gdf['count'][17]=2
gdf['count'][0]=4
gdf['count'][29]=4
gdf['count'][11]=4
gdf['count'][18]=6
gdf['count'][25]=6
#hide
gdf["x"] = gdf.centroid.x
gdf["y"] = gdf.centroid.y
gdf
#hide
gdf['Authority'] = 'NA'
gdf['FRT_system'] = 'NA'
gdf['Place'] = 'NA'
#hide
gdf['Authority'][0] = 'Hyderabad Police'
gdf['FRT_system'][0] = 'TSCOP + CCTNS'
gdf['Place'][0] = 'Hyderabad'
gdf['Authority'][25] = 'Chennai Police'
gdf['FRT_system'][25] = 'FaceTagr'
gdf['Place'][25] = 'Chennai'
gdf['Authority'][35] = 'Delhi Police'
gdf['FRT_system'][35] = 'AFRS'
gdf['Place'][35] = 'Delhi'
gdf['Authority'][27] = 'UP Police'
gdf['FRT_system'][27] = 'Trinetra'
gdf['Place'][27] = 'UP/Lucknow'
# +
#hide_input
multi = alt.selection_multi(fields=['count','state'], bind='legend')
color = alt.condition(multi,
alt.Color('count', type='ordinal',
scale=alt.Scale(scheme='yellowgreenblue')),
alt.value('lightgray'))
choro = alt.Chart(gdf).mark_geoshape(
stroke='black'
).encode(
color=color,
tooltip=['state','count']
).add_selection(
multi
).properties(
width=300,
height=400
)
c1 = alt.layer(choro).configure_legend(
orient = 'bottom-right',
direction = 'horizontal',
padding = 10,
rowPadding = 15
)
labels = alt.Chart(gdf).mark_text().encode(
longitude='x',
latitude='y',
text='count',
size=alt.value(8),
opacity=alt.value(0.6)
)
# Base chart for data tables
ranked_text = alt.Chart(gdf).mark_text(align='left').encode(
y=alt.Y('row_number:O',axis=None)
).transform_window(
row_number='row_number()'
).transform_filter(
multi
).transform_window(
rank='rank(row_number)'
).transform_filter(
alt.datum.rank<36
).properties(
width=15
)
# Data Tables
state = ranked_text.encode(text='state').properties(title='State')
a = ranked_text.encode(text='Authority').properties(title='Authority')
b = ranked_text.encode(text='FRT_system').properties(title='FRT_system')
c = ranked_text.encode(text='Place').properties(title='Place')
text = alt.hconcat(state,a,b,c) # Combine data tables
# Build chart
# alt.vconcat(
# choro+labels,
# text
# ).configure_legend(
# orient = 'left',
# padding = 10,
# rowPadding = 15
# ).configure_view(strokeWidth=0)
(c1+labels).configure_view(strokeWidth=0)
# -
#hide
import pandas as pd
frt = pd.read_csv('frt.csv')
# ### FRT Systems Deployed in India
#hide_input
frt
# +
#hide
# # Base chart for data tables
# ranked_text = alt.Chart(gdf).mark_text().encode(
# y=alt.Y('row_number:O',axis=None)
# ).transform_window(
# row_number='row_number()'
# ).transform_filter(
# multi
# ).transform_window(
# rank='rank(row_number)'
# ).transform_filter(
# alt.datum.rank<20
# )
# # Data Tables
# state = ranked_text.encode(text='state').properties(title='State')
# a = ranked_text.encode(text='Authority').properties(title='Authority')
# b = ranked_text.encode(text='FRT_system').properties(title='FRT_system')
# c = ranked_text.encode(text='Place').properties(title='Place')
# text = alt.hconcat(state,a,b,c) # Combine data tables
# # Build chart
# alt.hconcat(
# choro+labels,
# text
# ).configure_legend(
# orient = 'left',
# padding = 10,
# rowPadding = 15
# )
| _notebooks/2020-06-20-India_test_altair.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.5 64-bit
# name: python37564bit0b633b15d70b48adb30f596c0f66ec29
# ---
# + tags=[]
from cy_components.helpers.formatter import DateFormatter as df
from datetime import datetime
import pytz
now = datetime.now()
ts = df.convert_local_date_to_timestamp(now)
date_utc = df.convert_timestamp_to_local_date(ts, tz=pytz.utc)
date_local = df.convert_timestamp_to_local_date(ts)
date_local_with_tz = df.convert_timestamp_to_local_date(ts, tz=pytz.timezone("Asia/Shanghai"))
print(now, ts)
print(date_utc, date_utc.astimezone(tz=pytz.timezone("Asia/Shanghai")))
print(date_local, date_local.tzinfo, date_local.astimezone(tz=pytz.utc), date_local.replace(tzinfo=pytz.utc))
print(date_local_with_tz)
# + tags=[]
from cy_components.helpers.formatter import DateFormatter as df
from datetime import datetime
import pytz
date_string = '2020-07-07 22:23:00'
date = df.convert_string_to_local_date(date_string)
print(date.tzinfo, date)
date_replace = date.replace(tzinfo=pytz.timezone('America/Nome'))
date_as = date.astimezone(tz=pytz.timezone('America/Nome'))
print(date_replace)
print(date_as)
print(df.convert_local_date_to_timestamp(date_replace))
print(df.convert_local_date_to_timestamp(date_as))
date_replace = date.replace(tzinfo=pytz.utc)
date_as = date.astimezone(tz=pytz.utc)
print(date_replace)
print(date_as)
print(df.convert_local_date_to_timestamp(date_replace))
print(df.convert_local_date_to_timestamp(date_as))
# -
| tests/test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Numpy - operações que reduzem uma dimensão
#
# Uma das categorias de operações matriciais bastante útil do Numpy é a de operações
# que reduzem uma dimensão da matriz.
#
# Por exemplo, se eu quero calcular o perfil de cinza médio das linhas de uma imagem, é
# necessário fazer, para cada coluna, uma média entre todos os pixels daquela coluna.
#
# Leia com atenção o tutorial:
#
# - [Redução de eixo](../master/tutorial_numpy_1_5a.ipynb)
#
# O Numpy possui a facilidade de redução de eixo em muitas operações tais como `mean`, `max`, `min`, `all`, `any`, `sum` e
# tantas outras.
#
# Estas funções permitem passar como parâmetro a dimensão em que o `ndarray` será reduzido.
#
# Vejamos o exemplo a seguir, agora usando a redução implícita sem a necessidade de fazer o laço anterior:
# %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib.image as mpimg
import numpy as np
f = mpimg.imread('../data/cameraman.tif')[:6,:10] # slice das 6 primeiras linhas e 10 primeiras colunas
print('f=\n', f)
g = f.mean(axis=0)
print('g=\n', g.round(1))
print('f.shape=',f.shape)
print('f.mean(0).shape=',f.mean(axis=0).shape)
# Veja o exemplo de cálculo de valor mínimo da uma imagem das teclas da calculadora na página `master:tutorial_numpy_1_5a Redução de eixo`.
# Repita a seguir o mesmo exemplo, porém utilizando o perfil máximo no lugar do perfil mínimo:
f = mpimg.imread('../data/keyb.tif')
plt.imshow(f,cmap='gray')
m = f.mean()
print('f.mean', m)
hmax = f.max(axis=0)
plt.plot(f[75,:])
plt.plot(hmax)
#,ylabel='intensidade máxima',xlabel='coluna'),'f.max(0), valor máximo de cada coluna')
# ## Teste de autoavaliação
#
# Faça o teste múltipla escolha a seguir para verificar os conhecimentos adquiridos com esta atividade.
# O teste é para autoestudo e pode ser repetido várias vezes:
#
# - `http://adessowiki.fee.unicamp.br/adesso-1/q/ae2-2/ Teste de autoestudo - Redução de eixo`
#
| deliver/Atividade_2_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %run BBDTconvert.py
# # Print feature bins for each formula
import os
for _, _, files in os.walk('bbdt_run2/'):
for filename in files:
if not filename.endswith('.mx'):
continue
with open("bbdt_run2/{}".format(filename), "r") as f:
print "FORMULA:", filename
hlt = unpack_formula(f)
print "\n\n\n"
# # Convert each formula to BBDT format
import cPickle
with open('models/bbdt_thresholds.pkl', 'r') as f:
thresholds_hlt2 = cPickle.load(f)
with open('models/bbdt_thresholds_hlt1.pkl', 'r') as f:
thresholds_hlt1 = cPickle.load(f)
thresholds = dict(thresholds_hlt1.items() + thresholds_hlt2.items())
for _, _, files in os.walk('bbdt_run2/'):
for filename in files:
if not filename.endswith('.mx'):
continue
with open("bbdt_run2/{}".format(filename), "r") as f:
print "FORMULA:", filename
write_formula("bbdt_run2/{}".format(filename), "bbdt_run2/{}.bbdt".format(filename), thresholds[filename])
| BBDT-prepare.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # One curve
# ## plot a simple curve and play with it
#
# - $x=[0, pi]$
#
# - $y=e^x$
#
# - see documentation: http://www.silx.org/doc/silx/dev/modules/gui/plot/plotwindow.html#silx.gui.plot.PlotWindow.Plot1D
#
# - see tutorial: http://www.silx.org/doc/silx/dev/modules/gui/plot/getting_started.html
#
# - use Plot1D and Plot1D.addCurve
# - legend is used as the ID of the curve. So if a new curve is setted with an existing id it will erase the first curve
#
# 
#
# play with the interface:
# - log scale
# - grid
# - display points
# - ...
import numpy
from silx.gui.plot import Plot1D
# %gui qt
import numpy
x=numpy.linspace(0, numpy.pi, 1000)
y=numpy.exp(x)
...
# ## Shift the curve
# get back the curve and add an offset in y axis
#
# - $y=y+100.0$
# - get all needed data from the 'Plot1D' object
# - use getCurve([curveID]) function. Return :
# - x
# - y
# - legend
# - info (if some informations has been added)
# - params (color, linewidth...)
#
# 
...
# # Many curves
# ## plot the following functionin the same plot window
# - $y=sin(x)$
#
# - $y=cos(x)$
#
# - $y=x $
#
# - play with the curve selection from options->legend
#
# 
...
# ## remove one curve by the id
#
# - using the 'Plot1D' function 'remove([curveID])'
...
# ## shift curves by 30 in the x axis
# - by using the functions of the 'Plot1D' object
# - getAllCurves
# - addCurve
# - keep at least the color of the curve
# - Result should be close to
#
# 
...
# # ROI
#
# ## load data from data/spectrum.dat
import silx.io
sf = silx.io.open("data/spectrum.dat")
x_data=sf['1.1/measurement/channel']
y_data=sf['1.1/measurement/counts']
# ## Plot the data
plot=Plot1D()
x=numpy.linspace(0.0, numpy.pi)
y=numpy.sin(x)
plot.addCurve(x_data, y_data)
plot.setYAxisLogarithmic(True)
plot.show()
# options -> ROI -> add ROI -> select min and max limits.
# estimate integral between lower and upper limits
# - Raw counts
# 
# - Net counts
# 
| silx/plot/Plot1DExercise.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# Dependencies
import requests
import json
# URL for GET requests to retrieve vehicle data
url = "https://api.spacexdata.com/v2/launchpads"
# +
# Print the response object to the console
# +
# Retrieving data and converting it into JSON
# +
# Pretty Print the output of the JSON
| 1/Activities/01-Ins_RequestsIntro/Unsolved/Ins_Requests_Demo.ipynb |
/ -*- coding: utf-8 -*-
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: SQL
/ language: sql
/ name: SQL
/ ---
/ + [markdown] azdata_cell_guid="b65ecc14-f5b1-469e-ad46-41d63783f8fc"
/ # Introduction to SQL for Excel Users – Part 14: More Dates
/
/ [Original post](https://www.daveondata.com/blog/introduction-to-sql-for-excel-users-part-14-more-dates/)
/ + [markdown] azdata_cell_guid="31d5ed53-2900-4541-a260-a0b820a08cae"
/ ## RFM Analysis
/
/ I cover RFM analysis more extensively in Part 11 of the series, I will summarize the work so far for convenience.
/
/ RFM analysis is a simple, but wildly useful, technique from the world of direct marketing.
/
/ RFM is primarily used as means of quantifying the value of customers along three vectors:
/
/ - **R**ecency – How recently has a customer made a purchase?
/ - **F**requency – How often does a customer make a purchase?
/ - **M**onetary – How much does a customer spend on purchases?
/
/ The analysis consists of ranking each customers **R**, **F**, and **M** with a score ranging from 1 to 10 – where 10 is the best score.
/
/ In the previous post I skipped Recency as I had not covered dates as of yet.
/
/ The following query was the end result of the previous post that provided **F** and **M** scores via the NTILE window function:
/ + azdata_cell_guid="29d1d9eb-93a5-489d-84c0-57914536e61a"
WITH CustomerSalesOrders AS
(
SELECT FIS.CustomerKey
,FIS.SalesOrderNumber
,SUM(SalesAmount) AS SalesAmount
FROM FactInternetSales FIS
GROUP BY FIS.CustomerKey, FIS.SalesOrderNumber
),
CustomerSalesOrderHistory AS
(
SELECT CSO.CustomerKey
,COUNT(*) AS SalesOrderCount
,SUM(CSO.SalesAmount) AS SalesAmount
FROM CustomerSalesOrders CSO
GROUP BY CSO.CustomerKey
)
SELECT CSOH.CustomerKey
,NTILE(10) OVER (ORDER BY CSOH.SalesOrderCount ASC) AS FrequencyScore
,NTILE(10) OVER (ORDER BY CSOH.SalesAmount ASC) AS MonetaryScore
FROM CustomerSalesOrderHistory CSOH
ORDER BY CSOH.CustomerKey;
/ + [markdown] azdata_cell_guid="dbd83073-516c-470a-9573-79db7134450f"
/ ## Adding Sales Order Dates
/
/ To assign a Recency score to customers I need sales order dates.
/
/ More specifically, I need the most recent sales order date for each customer.
/
/ As discussed in Part 11, I need to account for two aspects of the data:
/
/ Every sales order consists of multiple rows of data – one row for each sales order line
/ There can be multiple sales orders per customer
/ No worries!
/
/ Nothing the MAX aggregate function can’t handle!
/
/ First up, I need to change the CustomerSalesOrders CTE to return the most recent sales order (i.e., MAX) date for each sales order:
/ + azdata_cell_guid="44d0e11a-e871-4ac6-b428-dbdff8f5ac80"
WITH CustomerSalesOrders AS
(
SELECT FIS.CustomerKey
,FIS.SalesOrderNumber
,SUM(SalesAmount) AS SalesAmount
,MAX(OrderDate) AS OrderDate
FROM FactInternetSales FIS
GROUP BY FIS.CustomerKey, FIS.SalesOrderNumber
),
CustomerSalesOrderHistory AS
(
SELECT CSO.CustomerKey
,COUNT(*) AS SalesOrderCount
,SUM(CSO.SalesAmount) AS SalesAmount
FROM CustomerSalesOrders CSO
GROUP BY CSO.CustomerKey
)
SELECT CSOH.CustomerKey
,NTILE(10) OVER (ORDER BY CSOH.SalesOrderCount ASC) AS FrequencyScore
,NTILE(10) OVER (ORDER BY CSOH.SalesAmount ASC) AS MonetaryScore
FROM CustomerSalesOrderHistory CSOH
ORDER BY CSOH.CustomerKey;
/ + [markdown] azdata_cell_guid="782dc1b2-7fd1-4120-8325-3b9131b910de"
/ By using MAX I’ve made sure that every sales order has the single, most recent order date.
/
/ Moving on, I need to modify the CustomerSalesOrderHistory CTE to have the most recent order date for each customer.
/
/ Once again, MAX to the rescue:
/ + azdata_cell_guid="1c3d6a36-341a-4fa7-b39b-c4f1fe90ddb4"
WITH CustomerSalesOrders AS
(
SELECT FIS.CustomerKey
,FIS.SalesOrderNumber
,SUM(SalesAmount) AS SalesAmount
,MAX(OrderDate) AS OrderDate
FROM FactInternetSales FIS
GROUP BY FIS.CustomerKey, FIS.SalesOrderNumber
),
CustomerSalesOrderHistory AS
(
SELECT CSO.CustomerKey
,COUNT(*) AS SalesOrderCount
,SUM(CSO.SalesAmount) AS SalesAmount
,MAX(CSO.OrderDate) AS MostRecentOrderDate
FROM CustomerSalesOrders CSO
GROUP BY CSO.CustomerKey
)
SELECT CSOH.CustomerKey
,NTILE(10) OVER (ORDER BY CSOH.SalesOrderCount ASC) AS FrequencyScore
,NTILE(10) OVER (ORDER BY CSOH.SalesAmount ASC) AS MonetaryScore
FROM CustomerSalesOrderHistory CSOH
ORDER BY CSOH.CustomerKey;
/ + [markdown] azdata_cell_guid="657b7937-fa50-4043-8f0f-72588a8edb30"
/ With MostRecentOrderDate added to CustomerSalesOrderHistory I’ve got all the raw materials I need to calculate Recency.
/ + [markdown] azdata_cell_guid="a61b3881-3bc4-412a-950f-e065c7fa375c"
/ ## Date Differences in Excel
/
/ It shoud come as no surprise that Excel has a collection of date and time functions.
/
/ One of the most useful of these functions is DATEDIF.
/
/ The mighty DATEDIF allows for the calculation of elapsed time between two dates using different timescales.
/
/ For example, how many days have elapsed between two dates:
/
/ 
/
/ 
/
/ I calculated elapsed months and years by using m or y instead of d as the last DATEDIF parameter:
/
/ 
/
/ Like chocolate and peanut butter, DATEDIF is even better with the NOW function:
/
/ 
/
/ 
/
/ Awesome!
/
/ Given the pattern, it ain’t surprising that working with dates in SQL is basically the same. 😁
/ + [markdown] azdata_cell_guid="f5f9da9e-ba53-4e78-97a6-3d66ede3f43b"
/ ## Date Differences T-SQL
/
/ It shouldn’t surprise you to know that SQL Server has a number of date and time functions.
/
/ As with Excel, one of the most useful T-SQL date functions is the mighty DATEDIFF.
/
/ Conceptually, T-SQL’s DATEDIFF works just like Excel’s DATEDIF, but the parameters are in a different order:
/
/ ```
/ DATEDIFF(<timescale>, <start date>, <end date>)
/ ```
/ + azdata_cell_guid="bd9239e7-8253-4bf9-9086-d894b906183f"
SELECT DATEDIFF(DAY,'2018-01-01 00:00:00', '2020-01-01 00:00:00') AS DiffInDays
,DATEDIFF(MONTH,'2018-01-01 00:00:00', '2020-01-01 00:00:00') AS DiffInMonths
,DATEDIFF(YEAR,'2018-01-01 00:00:00', '2020-01-01 00:00:00') AS DiffInYears
/ + [markdown] azdata_cell_guid="4834e9d7-289f-4d57-acb1-5402938cc6b3"
/ Also, just like in Excel, I can get the chocolate and peanut butter effect by combining DATEDIFF with CURRENT_TIMESTAMP:
/ + azdata_cell_guid="be790a3a-99de-4a9a-87d2-39daf4985792"
SELECT DATEDIFF(DAY,'2018-01-01 00:00:00', CURRENT_TIMESTAMP) AS DiffInDays
,DATEDIFF(MONTH,'2018-01-01 00:00:00', CURRENT_TIMESTAMP) AS DiffInMonths
,DATEDIFF(YEAR,'2018-01-01 00:00:00', CURRENT_TIMESTAMP) AS DiffInYears
/ + [markdown] azdata_cell_guid="d5ee0901-82f3-42b8-8005-5161eaef86a3"
/ Most excellent.
/
/ With the ability to calculate elapsed times, I can now finish up the RFM analysis.
/ + [markdown] azdata_cell_guid="78b94bc9-54e7-490c-883a-367e27cebc65"
/ Using DATEDIFF and CURRENT_TIMESTAMP I can modify the CustomerSalesOrderHistory CTE to calculate the elapsed days from the most recent sales order:
/ + azdata_cell_guid="45196fd0-1b93-42ff-976c-d9c0e959a607"
WITH CustomerSalesOrders AS
(
SELECT FIS.CustomerKey
,FIS.SalesOrderNumber
,SUM(SalesAmount) AS SalesAmount
,MAX(OrderDate) AS OrderDate
FROM FactInternetSales FIS
GROUP BY FIS.CustomerKey, FIS.SalesOrderNumber
),
CustomerSalesOrderHistory AS
(
SELECT CSO.CustomerKey
,COUNT(*) AS SalesOrderCount
,SUM(CSO.SalesAmount) AS SalesAmount
,DATEDIFF(DAY, MAX(CSO.OrderDate), CURRENT_TIMESTAMP) AS ElapsedDaysToMostRecentOrder
FROM CustomerSalesOrders CSO
GROUP BY CSO.CustomerKey
)
SELECT CSOH.CustomerKey
,NTILE(10) OVER (ORDER BY CSOH.SalesOrderCount ASC) AS FrequencyScore
,NTILE(10) OVER (ORDER BY CSOH.SalesAmount ASC) AS MonetaryScore
FROM CustomerSalesOrderHistory CSOH
ORDER BY CSOH.CustomerKey;
/ + [markdown] azdata_cell_guid="fee65149-6220-40f3-abac-183a9a576ffc"
/ Next up, I need to modify the outer query to add the Recency score:
/ + azdata_cell_guid="931869a2-8b03-4cd4-8be1-64ebc62a5ee0"
WITH CustomerSalesOrders AS
(
SELECT FIS.CustomerKey
,FIS.SalesOrderNumber
,SUM(SalesAmount) AS SalesAmount
,MAX(OrderDate) AS OrderDate
FROM FactInternetSales FIS
GROUP BY FIS.CustomerKey, FIS.SalesOrderNumber
),
CustomerSalesOrderHistory AS
(
SELECT CSO.CustomerKey
,COUNT(*) AS SalesOrderCount
,SUM(CSO.SalesAmount) AS SalesAmount
,DATEDIFF(DAY, MAX(CSO.OrderDate), CURRENT_TIMESTAMP) AS ElapsedDaysToMostRecentOrder
FROM CustomerSalesOrders CSO
GROUP BY CSO.CustomerKey
)
SELECT CSOH.CustomerKey
,NTILE(10) OVER (ORDER BY CSOH.ElapsedDaysToMostRecentOrder DESC) AS RecencyScore
,NTILE(10) OVER (ORDER BY CSOH.SalesOrderCount ASC) AS FrequencyScore
,NTILE(10) OVER (ORDER BY CSOH.SalesAmount ASC) AS MonetaryScore
FROM CustomerSalesOrderHistory CSOH
ORDER BY CSOH.CustomerKey;
/ + [markdown] azdata_cell_guid="267813f6-4e0f-42bb-8baa-3018e7921aa3"
/ Notice that the RecencyScore is calculated with NTILE using CSOH.ElapsedDaysToMostRecentOrder in descending order.
/
/ This is because I want the smallest value to recieve a score of 10 (i.e., less elapsed days are better).
/
/ Alrighty, then!
/
/ Lastly, if I wanted to see just my 10-10-10 customers:
/ + azdata_cell_guid="ff85bc1f-c324-4888-8341-ba8fd2ff680c"
WITH CustomerSalesOrders AS
(
SELECT FIS.CustomerKey
,FIS.SalesOrderNumber
,SUM(SalesAmount) AS SalesAmount
,MAX(OrderDate) AS OrderDate
FROM FactInternetSales FIS
GROUP BY FIS.CustomerKey, FIS.SalesOrderNumber
),
CustomerSalesOrderHistory AS
(
SELECT CSO.CustomerKey
,COUNT(*) AS SalesOrderCount
,SUM(CSO.SalesAmount) AS SalesAmount
,DATEDIFF(DAY, MAX(CSO.OrderDate), CURRENT_TIMESTAMP) AS ElapsedDaysToMostRecentOrder
FROM CustomerSalesOrders CSO
GROUP BY CSO.CustomerKey
),
RFMAnalysis AS
(
SELECT CSOH.CustomerKey
,NTILE(10) OVER (ORDER BY CSOH.ElapsedDaysToMostRecentOrder DESC) AS RecencyScore
,NTILE(10) OVER (ORDER BY CSOH.SalesOrderCount ASC) AS FrequencyScore
,NTILE(10) OVER (ORDER BY CSOH.SalesAmount ASC) AS MonetaryScore
FROM CustomerSalesOrderHistory CSOH
)
SELECT RFM.CustomerKey
,RFM.RecencyScore
,RFM.FrequencyScore
,RFM.MonetaryScore
FROM RFMAnalysis RFM
WHERE RFM.RecencyScore = 10 AND
RFM.FrequencyScore = 10 AND
RFM.MonetaryScore = 10
ORDER BY RFM.CustomerKey ASC;
/ + [markdown] azdata_cell_guid="4a063244-9566-4a7b-9fd5-bb388bb86b22"
/ There you have it.
/
/ RFM analysis is a wildly simple and useful technique.
/
/ I’ve personally used the ideas of RFM, for example, to rank US zip codes in terms of desirability for marketing efforts.
/
/ Now you can use RFM with your own business data.
/ + [markdown] azdata_cell_guid="47f0d43b-8a1f-4acc-8ad3-35ccde949e5d"
/ ## The Learning Arc
/
/ This won’t be the last time I cover working with time, but the series will be moving on.
/
/ Next up is coverage of working with more than 1 table of data at a time.
/
/ Yes, it is time to cover JOINs.
/
/ Stay healthy and happy data sleuthing
| content/dod_sql_14.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Building predictive models classifier partition by cases no dates
# ## Generic library methods
# +
import pandas as pd
import os
import numpy as np
from sklearn.metrics import accuracy_score, confusion_matrix, precision_score, recall_score, plot_confusion_matrix
from sklearn.utils.multiclass import unique_labels
from sklearn.model_selection import train_test_split
import matplotlib.pyplot as plt
import warnings
SEED = 17
def get_model_prediction(model, X_test):
return model.predict(X_test)
def evaluate_model(model, X_test, y_test, y_predicted, model_name):
merged_df = matrixes_to_dataframe(X_test, y_test)
df_predicted = pd.DataFrame({ 'Label' : y_predicted, 'Index' : merged_df.index} )
df_results = pd.DataFrame({ 'Label' : merged_df.Label, 'Index' : merged_df.index} )
print_model_metrics(model, X_test, y_test, y_predicted, model_name)
compare_prediction_visual(df_predicted , df_results, model_name)
def matrixes_to_dataframe(X_test, y_test):
y_test_df = pd.DataFrame(y_test)
y_test_df.columns = ['Label']
X_test_df = pd.DataFrame(X_test)
X_test_df.columns = ['Yesterday_Open', 'Yesterday_Close', 'Yesterday_Volume',
'Yesterday_Low', 'Yesterday_High', 'Open', 'High', 'Low', 'Close',
'Volume', 'OpenInt', 'Average_High_Low', 'Average_Day',
'Diff_Close_Open', 'Diff_Today_Open', 'Diff_Today_Close',
'Diff_Today_High', 'Diff_Today_Low', 'Month', 'Year', 'Day',
'Yesterday_Month', 'Yesterday_Year', 'Yesterday_Day']
df = pd.concat([X_test_df, y_test_df], axis=1, sort=False)
return df
def compare_prediction_visual(df_predicted, df_real, model_name='Unknown model'):
fig, axs = plt.subplots(1, 2, figsize=(15, 6), sharey=True, sharex=True, facecolor='w', edgecolor='k', dpi=80)
title = model_name + ' ' + 'classification results'
df_predicted.plot(kind='scatter', x='Index',y='Label', ax=axs[0], title='Prediction results')
df_real.plot(kind='scatter', x='Index',y='Label', ax=axs[1] ,title='Real classification results')
save_plt_report(model_name)
def print_model_metrics(model, X_test, y_test, y_predicted, model_name='Unknown model'):
print ('Metrics for model: {}'.format(model_name))
warnings.filterwarnings('ignore')
result = accuracy_score(y_test, y_predicted)
print ('\tAccuracy: {0:.3f}'.format(result))
print ('\tPrecision: {0:.3f}'.format(precision_score(y_test, y_predicted)))
print ('\tRecall: {0:.3f}'.format(recall_score(y_test, y_predicted)))
print ('\tPredicts {0:.3f}% time the stock price goes down'.format(result*100))
cm = confusion_matrix(y_test, y_predicted)
print( '\tConfusion matrix:')
print(cm)
title ="{}: Confusion matrix".format(model_name)
file = '{}_confision_matrix.png'.format(model_name)
disp = plot_confusion_matrix(model, X_test, y_test,
display_labels=["Don't increase", "Increase"],
cmap=plt.cm.Blues, normalize=None, values_format="3.0f")
disp.ax_.set_title(title)
save_plt_report(file)
plt.show()
def save_plt_report(file):
visualization_path = os.path.join(os.path.pardir, 'reports', 'figures', 'classifier-balanced', file)
plt.savefig(visualization_path);
def describe_arrays(y, array_name):
## results descriptions
print ('')
print ('')
print ('mean amount of increases in {0} : {1:.3f}'.format(array_name, np.mean(y)))
print ('total amount of increases in {0} : {1}'.format(array_name, np.sum(y)))
print ('total amount of registers in {0} : {1}'.format(array_name, y.size))
print ('total amount of decresses in {0} : {1}'.format(array_name, y.size - np.sum(y)))
def split_train_test_dataset(dataset):
clean_dataset, result_df = generate_train_test_datasets(dataset)
X = clean_dataset.values.astype('float')
y = result_df['Label'].ravel()
X_train, X_test, y_train, y_test = train_test_split(X, y,random_state=SEED)
return X_train, X_test, y_train, y_test
def generate_train_test_datasets(dataset):
result_df = pd.DataFrame()
result_df['Label'] = dataset.Label
result_df['Index'] = dataset.index
result_df['Date'] = dataset.Date
clean_dataset = dataset.drop(columns=['Tomorrow_Date', 'Tomorrow_Open','Label','Diff_Tomorrow_Open','Date','Yesterday_Date'])
print(clean_dataset.columns)
return clean_dataset, result_df
def load_dataset(subfolder='', file='aapl.us.txt', data_type='raw', index_column=0):
data_path = os.path.join(os.path.pardir, 'data', data_type, subfolder, file)
print('Opening file ', data_path)
df = pd.read_csv(data_path, index_col=index_column)
print('%d missing values found' % df.isnull().sum().sum())
return df
# -
dataset = load_dataset(file='dataset_feature_vector.csv', data_type='processed')
X_train, X_test, y_train, y_test = split_train_test_dataset(dataset )
describe_arrays(y_train, "y_train")
describe_arrays(y_test, "y_test")
# ## Generate models
# +
from sklearn.linear_model import LogisticRegression
from sklearn.dummy import DummyClassifier
def generate_simple_logistic_model(X_train, y_train):
model = LogisticRegression(random_state=0, solver='liblinear')
model.fit(X_train,y_train)
return model
def generate_dummy_model(X_train, y_train):
model = DummyClassifier(strategy='most_frequent', random_state=0)
model.fit(X_train, y_train)
return model
# -
linear_model = generate_dummy_model(X_train, y_train)
y_predicted = get_model_prediction(linear_model, X_test)
evaluate_model(linear_model, X_test, y_test, y_predicted, "Linear model")
logistic_model = generate_simple_logistic_model(X_train, y_train)
y_predicted = get_model_prediction(logistic_model, X_test)
evaluate_model(logistic_model, X_test, y_test, y_predicted, "Logistic model")
# +
from sklearn.model_selection import GridSearchCV
from sklearn.utils.testing import ignore_warnings
from sklearn.exceptions import ConvergenceWarning
def generate_GS_logistic_model(X_train, y_train):
logistic_model = generate_simple_logistic_model(X_train, y_train)
parameters = {'C':[1.0, 10.0, 50.0, 100.0, 1000.0], 'penalty' : ['l1','l2'],'class_weight' : ['balanced',None], 'solver' : ['liblinear','saga'], 'max_iter' : [100,1000,10000]}
model = GridSearchCV(logistic_model, param_grid=parameters, cv=3)
with ignore_warnings(category=ConvergenceWarning):
model.fit(X_train, y_train)
print ('Best parameters for logistic model grid {}'.format(model.best_params_))
print ('Best score : {0:.2f}'.format(model.best_score_))
return model
# -
logistic_GS_model = generate_GS_logistic_model(X_train, y_train)
y_predicted = get_model_prediction(logistic_GS_model, X_test)
evaluate_model(logistic_model, X_test, y_test, y_predicted, "Logistic model grid search")
# ## Feature standarization
def generate_FS_GS_logistic_model(X_train, y_train):
logistic_model = generate_simple_logistic_model(X_train, y_train)
parameters = {'C':[1.0, 10.0, 50.0, 100.0, 1000.0], 'penalty' : ['l1','l2'],'class_weight' : ['balanced',None], 'solver' : ['liblinear','saga'], 'max_iter' : [100,1000,10000]}
model = GridSearchCV(logistic_model, param_grid=parameters, cv=3)
with ignore_warnings(category=ConvergenceWarning):
model.fit(X_train, y_train)
print ('Best parameters for logistic model grid {}'.format(model.best_params_))
print ('Best score : {0:.2f}'.format(model.best_score_))
return model
from sklearn.preprocessing import MinMaxScaler, StandardScaler
scaler = StandardScaler()
X_train_scaled = scaler.fit_transform(X_train)
X_test_scaled = scaler.transform(X_test)
logistic_FS_GS_model = generate_FS_GS_logistic_model(X_train_scaled, y_train)
y_predicted = get_model_prediction(logistic_FS_GS_model, X_test_scaled)
evaluate_model(logistic_FS_GS_model, X_test_scaled, y_test, y_predicted, "Logistic model with FS and grid search")
# ## Random forest
from sklearn.ensemble import RandomForestClassifier
def generate_random_forest_model(X_train, y_train):
parameters = {
'n_estimators':[50, 100, 200,1000],
'min_samples_leaf':[1, 5,10,50],
'max_features' : ('auto','sqrt','log2'),
}
rf = RandomForestClassifier(random_state=0, oob_score=True)
model = GridSearchCV(rf, parameters, cv=5)
with ignore_warnings(category=ConvergenceWarning):
model.fit(X_train, y_train)
print ('Best parameters for random forest model grid {}'.format(model.best_params_))
print ('Best score : {0:.2f}'.format(model.best_score_))
return model
random_forest_GS_model = generate_random_forest_model(X_train, y_train)
y_predicted = get_model_prediction(random_forest_GS_model, X_test)
evaluate_model(random_forest_GS_model, X_test, y_test, y_predicted, "Random Forest with grid search")
| notebooks/Classifier 2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit
# name: python3
# ---
# # Quantitative Value Strategy
# ## Imports
import sys
# !{sys.executable} -m pip install numpy
# !{sys.executable} -m pip install pandas
# !{sys.executable} -m pip install python-dotenv
# !{sys.executable} -m pip install requests
# !{sys.executable} -m pip install xlsxwriter
from dotenv import load_dotenv
import math
import numpy
import os
import pandas
import requests
from scipy import stats
import xlsxwriter
# ## Import List of Stocks
stocks = pandas.read_csv('sp_500_stocks.csv')
stocks
# ## Acquiring API Token
key = os.getenv('ACCESS_KEY')
key
# ## Making a First API Call
symbol = 'AAPL'
url = f'https://sandbox.iexapis.com/stable/stock/{symbol}/quote?token={key}'
data = requests.get(url).json()
data
# ### Parsing API Call
price = data['latestPrice']
peRatio = data['peRatio']
# ## Executing Batch API Call
def chuncks(lst, n):
for i in range(0, len(lst), n):
yield lst[i:i + n]
symbol_groups = list(chuncks(stocks['Ticker'], 100))
symbol_groups
symbol_strings = []
for i in range(0, len(symbol_groups)):
symbol_strings.append(','.join(symbol_groups[i]))
symbol_strings
columns = ['Ticker', 'Price', 'Price-to-Earnings Ratio', 'Number of Shares to Buy']
columns
final_dataframe = pandas.DataFrame(columns=columns)
final_dataframe
# +
for symbol_string in symbol_strings:
url = f'https://sandbox.iexapis.com/stable/stock/market/batch?symbols={symbol_string}&types=quote&token={key}'
data = requests.get(url).json()
for symbol in symbol_string.split(','):
final_dataframe = final_dataframe.append(
pandas.Series([
symbol,
data[symbol]['quote']['latestPrice'],
data[symbol]['quote']['peRatio'],
'N/A'
], index= columns),
ignore_index=True
)
final_dataframe
# -
# ## Removing Glamour Stocks
final_dataframe.sort_values('Price-to-Earnings Ratio', ascending=True, inplace=True)
final_dataframe
final_dataframe = final_dataframe[final_dataframe['Price-to-Earnings Ratio'] > 0]
final_dataframe
final_dataframe = final_dataframe[:50]
final_dataframe
final_dataframe.reset_index(inplace=True)
final_dataframe
final_dataframe.drop('index', axis=1, inplace=True)
final_dataframe
# ### Calculating Shares to Buy
def portfolio_input():
global portfolio_size
portfolio_size = input('Enter the Portfolio size')
try:
val = float(portfolio_size)
except:
print('Try again. Enter the Portfolio size')
portfolio_size = input('Enter the Portfolio size')
portfolio_input()
portfolio_size
position_size = float(portfolio_size) / len(final_dataframe.index)
position_size
# +
for i in final_dataframe.index:
price = final_dataframe.loc[i, 'Price']
final_dataframe.loc[i, 'Number of Shares to Buy'] = math.floor(position_size / price)
final_dataframe
| quantitative_value.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from genesis import parsers
# +
# Change this
test_dir = '/Users/chrisonian/Code/genesis/examples/outfiles/'
out_fname = test_dir +'genesis.out'
dfl_fname = test_dir +'genesis.out.dfl'
fld_fname = test_dir +'genesis.out.fld'
# Get parameters from .out file
odat = parsers.parse_genesis_out(out_fname)
params = odat['param']
my_ncar = params['ncar']
my_dgrid = params['dgrid']
my_nz = 1
# +
my_dfl = parsers.parse_genesis_dfl(dfl_fname, nx=my_ncar)
my_dfl
# -
my_fld = parsers.parse_genesis_fld(fld_fname, my_ncar, my_nz)
my_fld[-1]
# # Plot
import matplotlib.pyplot as plt
# Field phase at end, slice 0
def plot_field(dat, dgrid):
ndat = np.angle(dat)
plt.imshow(ndat, extent = [1000*dgrid*i for i in [-1,1,-1,1]])
plt.xlabel('x (mm)')
plt.ylabel('y (mm)')
plt.show()
plot_field(my_dfl[:, :, 0], my_dgrid )
# Field phrase from history file, slice 0
plot_field(my_fld[:, :, 0, -1], my_dgrid )
| examples/example_parsing_genesis_field.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Eexamples of calculating interpretation quality metrics.
# ## Typically, you might need to obtain whole dataset metrics, then please use ready script *metrics.py* (which will do all calculations under the hood and return dataset-aggregated metrics and per-molecule metrics too, if needed.This notebook does the same job.
# ## However, there is a possibility to use functions from the script individually. The notebook shows how to use them.
# ### If you are interested in the details about metrics calculations, the notebook explains them step-by-step.
# ### Notebook may be convinient if you are interested in analysis of individual molecules.
#
#
import metrics as mt
from rdkit import Chem
from rdkit.Chem.Draw import SimilarityMaps
import numpy as np
import pandas as pd
# ### We will use as an example N dataset, where activity is defined as the count of Nitrogen.
# ### Load file with contributions
contribs = mt.read_contrib("example_notebook_data/N_contrib_per_atom_dc.txt")
contribs
# ### Load ground thruth labels
lbls = mt.read_lbls_from_sdf(
"example_notebook_data/N_train_lbl_small.sdf", lbls_field_name="lbls", sep=",")
lbls
# ### Merge contributions with ground truth. Note: the merge is *inner*, only molecules present in both will remain
merged = mt.merge_lbls_contribs(contribs, lbls)
merged
# ### Now we are ready to calculate metrics
# +
## Let's calculate AUC. which_lbls="positive" indicates to calculate AUC for positively contributing atoms. Specifying "negative"
## you switch to negatively contributing atoms, which in this dataset are absent.
# -
auc = mt.calc_auc(merged, which_lbls="positive", contrib_col_name="contribution")
auc
# +
## Lets compute another metric - top-n score, with variable n (indicated by n_list parameter="infinity"), and also with fixed n=3,5
# -
top_n = mt.calc_top_n(merged, n_list=[np.inf,3,5], contrib_col_name="contribution")
top_n.keys() # top-n is a dictionary with bunch of dataframes
# ### We will need only 'variable_top_n', 'top3' and 'top5' dataframes,
# ### In each dataframe "top_score" column contains scores (per molecule).
top_n["variable_top_n"]
top_n["top3"].top_score
top_n["top5"].top_score
# ## Now we can aggregate metrics per dataset
summary = mt.summarize([auc, top_n])
summary
# ## We can look at individual molecules, if we are interested
# ### Let's have a look at first molecule:
mols = Chem.SDMolSupplier("example_notebook_data/N_train_lbl_small.sdf")
m = mols[0]
nm = m.GetProp("_Name")
nm
m
auc["auc_pos"][nm] # look at AUC for this molecule, it's not perfect (<1)
top_n['top3']['top_score'][nm] # look at top-n score for this molecule, it's only 0.5
# ## Let's visualize contributions of atoms. Nitrogen has ground truth label "1". So contributions for nitrogens should be the highest
# ### We will use RDKit's Similarity Maps for highlighting atoms:
wt = {}
for n,atom in enumerate(Chem.rdmolfiles.CanonicalRankAtoms(m)):
wt[atom] = contribs.loc[contribs.molecule==nm,"contribution"].iloc[n]
sim_map = SimilarityMaps.GetSimilarityMapFromWeights(m,wt)
# ## As we can see, 1 Nitrogen atom has the highest contribution (brightest green), but second highest contribution belongs to Carbon (not second Nitrogen), so AUC is not perfect and top-n is 0.5.
| example_notebook.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## CIFAR-10 UAP - Evaluation
# We show examples of the following:
# 1. Evaluation on clean datasets (train, test)
# 2. Evaluation with chess noise pattern
# 3. Loading and evaluation on pre-computed UAP
# +
import matplotlib.pyplot as plt
import numpy as np
import os
import sys
import torch
sys.path.append(os.path.realpath('..'))
from utils import loader_cifar, model_cifar, evaluate
dir_data = '/data/cifar10'
dir_uap = '../uaps/cifar10/'
testloader = loader_cifar(dir_data = dir_data, train = False)
trainloader = loader_cifar(dir_data = dir_data, train = True)
# -
# load model
model, best_acc = model_cifar('resnet18', ckpt_path = '../resnet18.pth')
print(best_acc)
# ## 1. Clean
_, _, _, _, outputs, labels = evaluate(model, trainloader, uap = None)
print('Train accuracy:', sum(outputs == labels) / len(labels))
_, _, _, _, outputs, labels = evaluate(model, testloader, uap = None)
print('Test accuracy:', sum(outputs == labels) / len(labels))
# ## 2. Chessboard pattern
# load pattern
uap = torch.load(dir_uap + 'chess.pth')
# visualize chessboard
plt.imshow(np.transpose(uap, (1, 2, 0)))
# evaluate
eps = 10 / 255
_, _, _, _, outputs, labels = evaluate(model, testloader, uap = uap * eps)
print('Accuracy:', sum(outputs == labels) / len(labels))
plt.title('Clean test set distribution')
plt.hist(labels)
# plot histogram
plt.title('Chessboard test set distribution')
plt.hist(outputs)
# ## 3. Pre-computed UAP
# load UAP targeting class 4 with eps = 10
y_target = 4
eps = 10
uap = torch.load(dir_uap + 'sgd-tgt%i-eps%i.pth' % (y_target, eps))
# visualize UAP
uap_max = torch.max(uap)
plt.imshow(np.transpose(((uap / uap_max) + 1) / 2, (1, 2, 0)))
# evaluate
_, _, _, _, outputs, labels = evaluate(model, testloader, uap = uap)
print('Accuracy:', sum(outputs == labels) / len(labels))
print('Targeted success rate:', sum(outputs == y_target) / len(labels))
# plot histogram
plt.title('UAP test set distribution')
plt.hist(outputs)
| notebooks/cifar10_eval.ipynb |