code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# ASSIGNMENT 1
# +
# You all are pilots, you have to land a plane, the altitude requied for landing a plane is 1000 ft, if it is less than that tell the pilot to land the plane, or if it is more than that but less than 5000ft, ask the pilot to "come down to 1000ft", else if it is more than 5000ft, ask the pilot to "go around and try later"
# +
altitude= int(input("Current altitude "))
if altitude<=1000:
print("Safe to land")
elif ((altitude>=1000) & (altitude<=5000)):
print("Bring down to 1000")
elif altitude>5000:
print("Turn around")
# +
# ASSIGNMENT 2
# +
# Using FOR loop print all the prime numbers between 1- 200.
# -
for num in range (1, 201):
count = 0
for i in range(2, (num//2 + 1)):
if(num % i == 0):
count = count + 1
break
if (count == 0 and num!=1):
print(num)
| Python day 3 assignment.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Semana 12
#
# ## Módulo 4a: Ferramentas digitais e o ensino de história
#
# **Período**: 17/01/2022 a 21/01/2022
#
# **CH**: 2h
# ### Atividade Assíncrona 6 (AA)
#
# Tutorial 02: **Construindo mapas que contam histórias com o StoryMapJS**
# <iframe width="560" height="315" src="https://www.youtube.com/embed/86xpQQmf8kM" title="YouTube video player" frameborder="0" allow="accelerometer; autoplay; clipboard-write; encrypted-media; gyroscope; picture-in-picture" allowfullscreen></iframe>
| cclhm0069/_build/jupyter_execute/mod4a/sem12.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.7.1
# language: julia
# name: julia-1.7
# ---
using FFTW
?fft
fft(randn(5))
n = 10
θ = range(0, 2π; length=n+1)[1:n]
b = randn(n)
[exp(-im*k*θ[j]) for k=0:n-1, j=1:n] * b - fft(b)
| lectures/Untitled20.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Demo for Linking MyGene.info and MyVariant.info Using JSON-LD
# Use Case Scenario:
# Upstream analysis identified a missense variant (chr8:g.99440236C>A).
# The analyst wants to obtain the matching InterPro data in order to assess the likely functional significance of this variant.
# #### Step 1: Query MyVariant.info to retrieve the annotation object for variant chr8:g.99440236C>A
from jsonld_application import BioThings
bt = BioThings()
doc = bt.get('chr8:g.99440236C>A', 'variant')
# #### Step 2: List all available apis linked from this variant
doc.get_linked_apis()
# #### Step 3: Link to mygene.info
gene_doc = doc.get_jsondoc('gene')
# #### Step 4: Query mygene.info for Interpro
gene_doc['interpro']
| tutorial/Demo for Linking MyGene.info and MyVariant.info Using JSON-LD.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Calculators
# + [markdown] slideshow={"slide_type": "-"} tags=["remove-cell"]
# **CS1302 Introduction to Computer Programming**
# ___
# + [markdown] slideshow={"slide_type": "subslide"}
# Run the following to load additional tools required for this lab.
# In particular, the `math` library provides many useful mathematical functions and constants.
# + hide_input=false init_cell=true slideshow={"slide_type": "-"}
# %reset -f
from ipywidgets import interact
import matplotlib.pyplot as plt
import numpy as np
import math
from math import log, exp, sin, cos, tan, pi
# + [markdown] slideshow={"slide_type": "subslide"}
# The following code is a Python one-liner that creates a calculator.
# Evaluate the cell with `Ctrl+Enter`:
# + slideshow={"slide_type": "-"}
print(eval(input()))
# + [markdown] slideshow={"slide_type": "fragment"}
# Try some calculations below using this calculator:
# 1. $2^3$ by entering `2**3`;
# 1. $\frac23$ by entering `2/3`;
# 1. $\left\lceil\frac32\right\rceil$ by entering `3//2`;
# 1. $3\mod 2$ by entering `3%2`;
# 1. $\sqrt{2}$ by entering `2**0.5`; and
# 1. $\sin(\pi/6)$ by entering `sin(pi/6)`;
# + [markdown] slideshow={"slide_type": "fragment"}
# For this lab, you will create more powerful and dedicated calculators.
# We will first show you a demo. Then, it will be your turn to create the calculators.
# + [markdown] slideshow={"slide_type": "slide"}
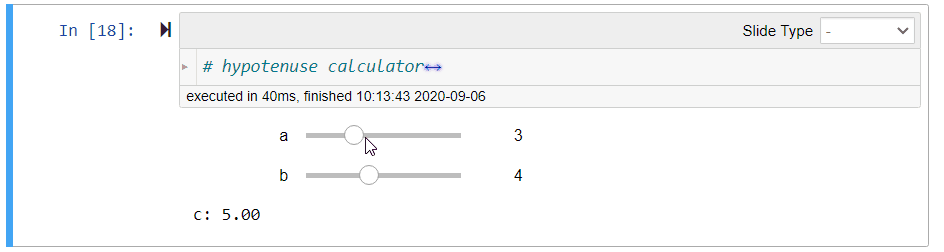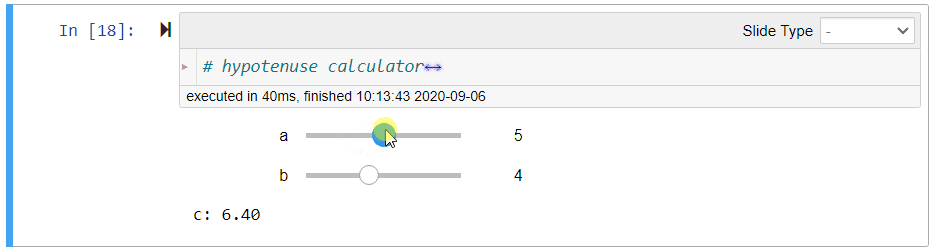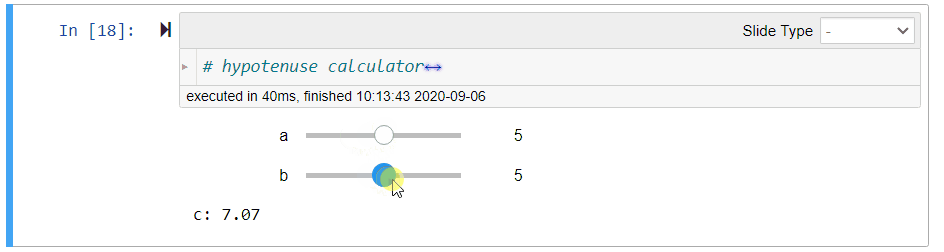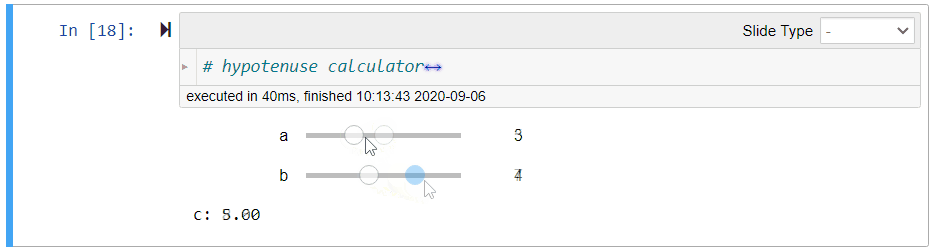
# ## Hypotenuse Calculator (Demo)
# + [markdown] slideshow={"slide_type": "-"}
# 
# + [markdown] slideshow={"slide_type": "fragment"}
# Using the Pythagoras theorem below, we can define the following function `length_of_hypotenus` to calculate the length `c` of the hypotenuse given the lengths `a` and `b` of the other sides of a right-angled triangle:
# $$c = \sqrt{a^2 + b^2}$$
# + slideshow={"slide_type": "fragment"}
def length_of_hypotenuse(a, b):
c = (a**2 + b**2)**(0.5) # Pythagoras
return c
# + [markdown] slideshow={"slide_type": "fragment"}
# - You need not understand how a function is defined, but
# - you should know how to *write the formula as a Python expression*, and
# - *assign to the variable* `c` the value of the expression (Line 2).
# + [markdown] slideshow={"slide_type": "subslide"}
# For example, you may be asked to write Line 2, while Line 1 and 3 are given to you:
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Complete the function below to return the length `c` of the hypotenuse given the lengths `a` and `b`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "9f8cd82c449d76657e17c6b1397ddeb2", "grade": false, "grade_id": "length_of_hypotenus", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def length_of_hypotenuse(a, b):
# YOUR CODE HERE
raise NotImplementedError()
return c
# + [markdown] slideshow={"slide_type": "fragment"}
# Note that indentation affects the execution of Python code. The assignment statement must be indented to indicate that it is part of the *body* of the function.
# (Try removing the indentation and see what happens.)
# + [markdown] slideshow={"slide_type": "subslide"}
# We will use widgets (`ipywidgets`) to let user interact with the calculator more easily:
# + code_folding=[0] slideshow={"slide_type": "-"}
# hypotenuse calculator
@interact(a=(0, 10, 1), b=(0, 10, 1))
def calculate_hypotenuse(a=3, b=4):
print('c: {:.2f}'.format(length_of_hypotenuse(a, b)))
# + [markdown] slideshow={"slide_type": "fragment"}
# After running the above cell, you can move the sliders to change the values of `a` and `b`. The value of `c` will be updated immediately.
# + [markdown] slideshow={"slide_type": "fragment"}
# - For this lab, you need not know how write widgets, but
# - you should know how to *format a floating point number* (Line 3).
# + [markdown] slideshow={"slide_type": "subslide"}
# You can check your code with a few cases listed in the test cell below.
# + code_folding=[0] deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "b419ad388ec1d742ab7cf99085178470", "grade": true, "grade_id": "test-length_of_hypotenus", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "-"}
# tests
def test_length_of_hypotenuse(a, b, c):
c_ = length_of_hypotenuse(a, b)
correct = math.isclose(c, c_)
if not correct:
print(f'For a={a} and b={b}, c should be {c}, not {c_}.')
assert correct
test_length_of_hypotenuse(3, 4, 5)
test_length_of_hypotenuse(0, 0, 0)
test_length_of_hypotenuse(4, 7, 8.06225774829855)
# + [markdown] slideshow={"slide_type": "slide"}
# ## Quadratic Equation
# + [markdown] slideshow={"slide_type": "subslide"}
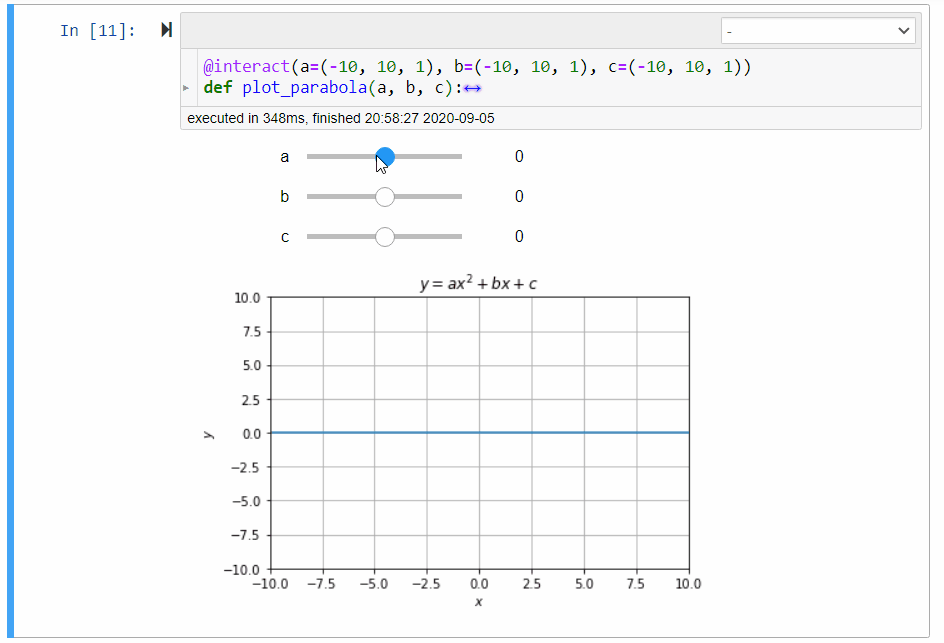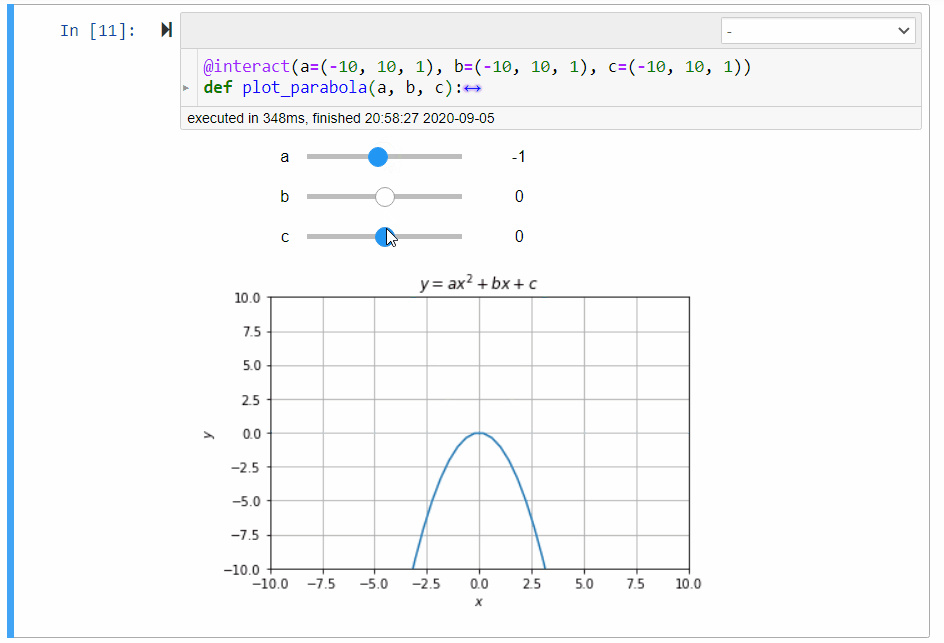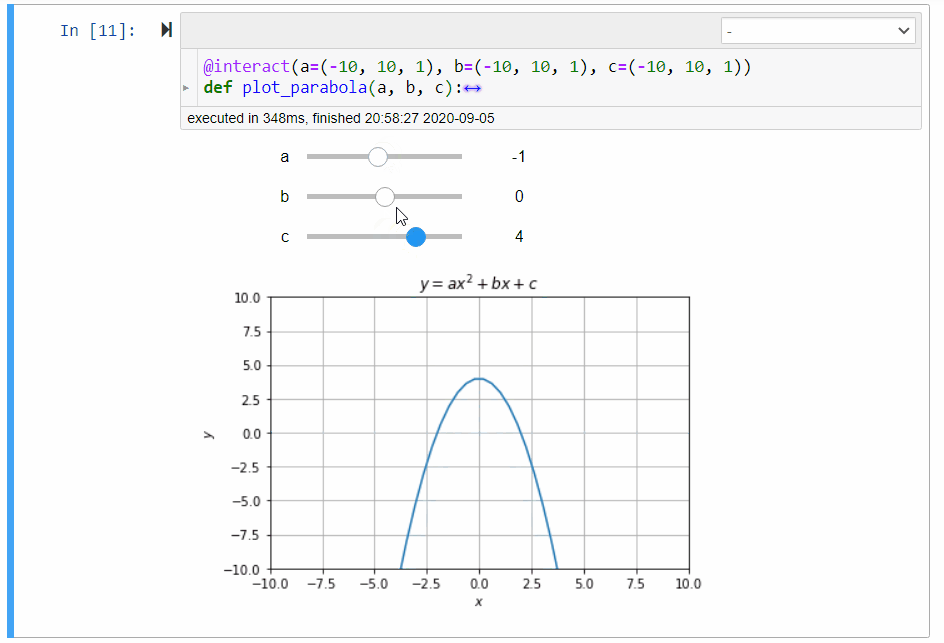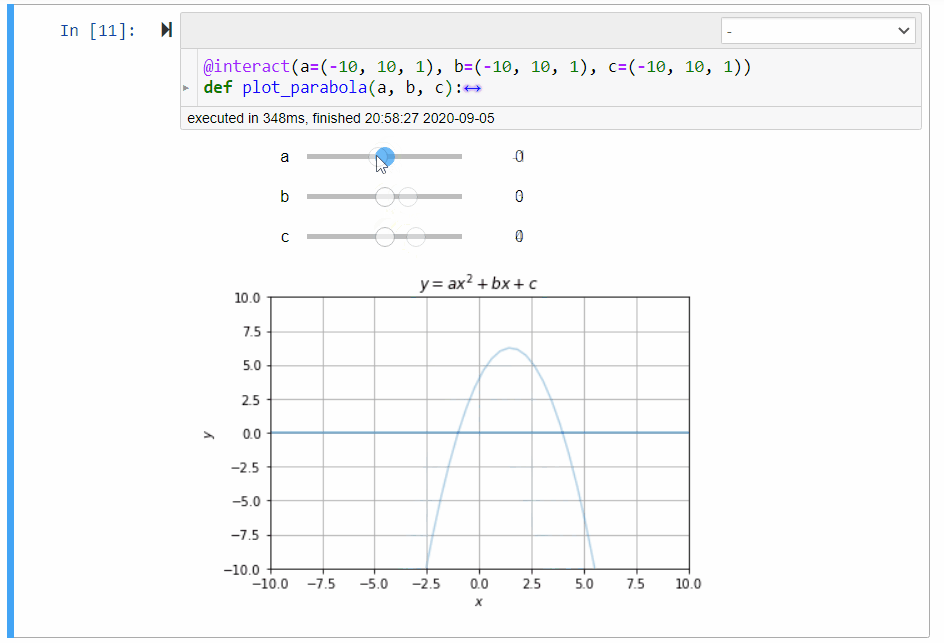
# ### Graphical Calculator for Parabola
# + [markdown] slideshow={"slide_type": "-"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# In mathematics, the collection of points $(x,y)$ satisfying the following equation forms a *parabola*:
#
# $$
# y=ax^2+bx+c
# $$
# where $a$, $b$, and $c$ are real numbers called the *coefficients*.
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Given the variables `x`, `a`, `b`, and `c` store the $x$-coordinate and the coefficients $a$, $b$, and $c$ respectively, assign to `y` the corresponding $y$-coordinate for the parabola.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "2c86fa4ce83bcbe0906f9b64b5c1a032", "grade": false, "grade_id": "get_y", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def get_y(x, a, b, c):
# YOUR CODE HERE
raise NotImplementedError()
return y
# + code_folding=[0] deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "f788b3d68b5184b680c391a373c57d46", "grade": true, "grade_id": "test-get_y", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "fragment"}
# tests
def test_get_y(y,x,a,b,c):
y_ = get_y(x,a,b,c)
correct = math.isclose(y,y_)
if not correct:
print(f'With (x, a, b, c)={x,a,b,c}, y should be {y} not {y_}.')
assert correct
test_get_y(0,0,0,0,0)
test_get_y(1,0,1,2,1)
test_get_y(2,0,2,1,2)
# + code_folding=[] slideshow={"slide_type": "subslide"}
# graphical calculator for parabola
@interact(a=(-10, 10, 1), b=(-10, 10, 1), c=(-10, 10, 1))
def plot_parabola(a, b, c):
xmin, xmax, ymin, ymax, resolution = -10, 10, -10, 10, 50
ax = plt.gca()
ax.set_title(r'$y=ax^2+bx+c$')
ax.set_xlabel(r'$x$')
ax.set_ylabel(r'$y$')
ax.set_xlim([xmin, xmax])
ax.set_ylim([ymin, ymax])
ax.grid()
x = np.linspace(xmin, xmax, resolution)
ax.plot(x, get_y(x, a, b, c))
# + [markdown] slideshow={"slide_type": "slide"}
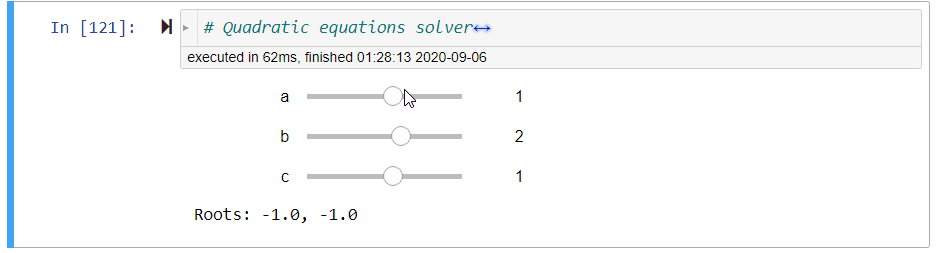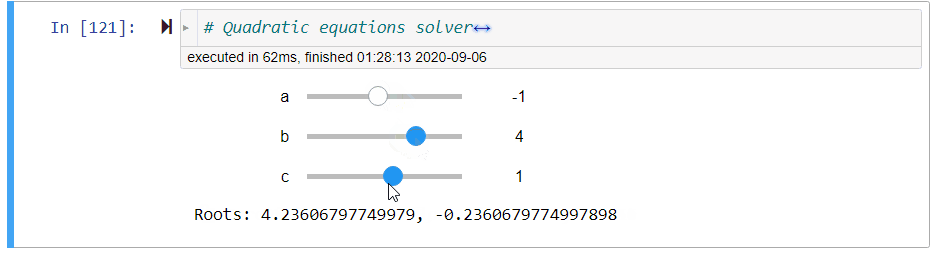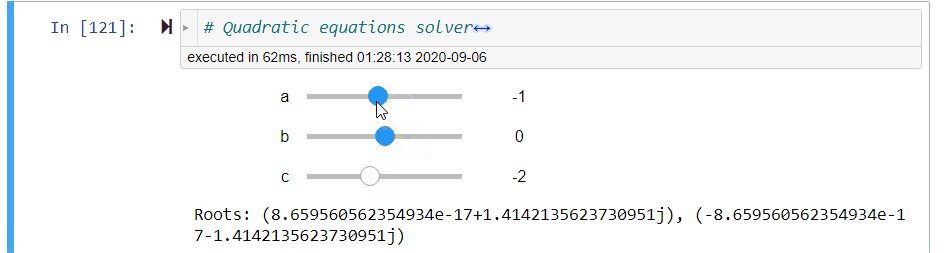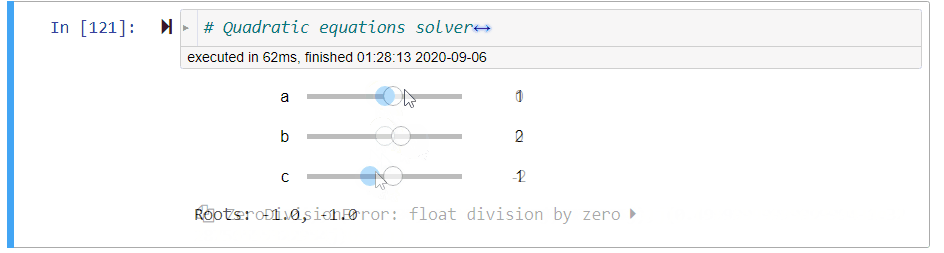
# ### Quadratic Equation Solver
# + [markdown] slideshow={"slide_type": "-"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# For the quadratic equation
#
# $$
# ax^2+bx+c=0,
# $$
# the *roots* (solutions for $x$) are give by
#
# $$
# \frac{-b-\sqrt{b^2-4ac}}{2a},\frac{-b+\sqrt{b^2-4ac}}{2a}.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Assign to `root1` and `root2` the values of the first and second roots above respectively.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "ac8d52d2f7328a894d73bce80d19dafc", "grade": false, "grade_id": "get_roots", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "fragment"}
def get_roots(a, b, c):
# YOUR CODE HERE
raise NotImplementedError()
return root1, root2
# + code_folding=[] deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "6c4b5df11d68aac1425187107b9a4788", "grade": true, "grade_id": "test-get_roots", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "fragment"}
# tests
def test_get_roots(roots, a, b, c):
roots_ = get_roots(a, b, c)
correct = all([math.isclose(roots[i], roots_[i]) for i in range(2)])
if not correct:
print(f'With (a, b, c)={a,b,c}, roots should be {roots} not {roots_}.')
assert correct
test_get_roots((-1.0, 0.0), 1, 1, 0)
test_get_roots((-1.0, -1.0), 1, 2, 1)
test_get_roots((-2.0, -1.0), 1, 3, 2)
# + code_folding=[0] slideshow={"slide_type": "fragment"}
# quadratic equations solver
@interact(a=(-10,10,1),b=(-10,10,1),c=(-10,10,1))
def quadratic_equation_solver(a=1,b=2,c=1):
print('Roots: {}, {}'.format(*get_roots(a,b,c)))
# + [markdown] slideshow={"slide_type": "slide"}
# ## Number Conversion
# + [markdown] slideshow={"slide_type": "subslide"}
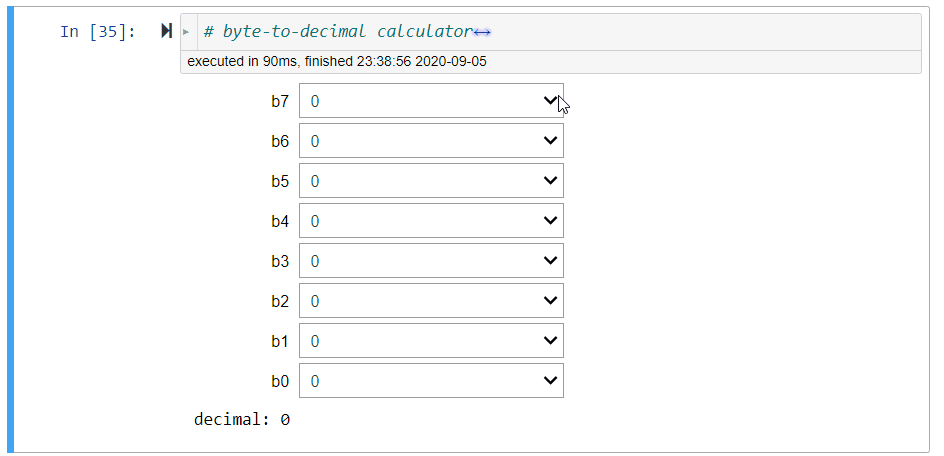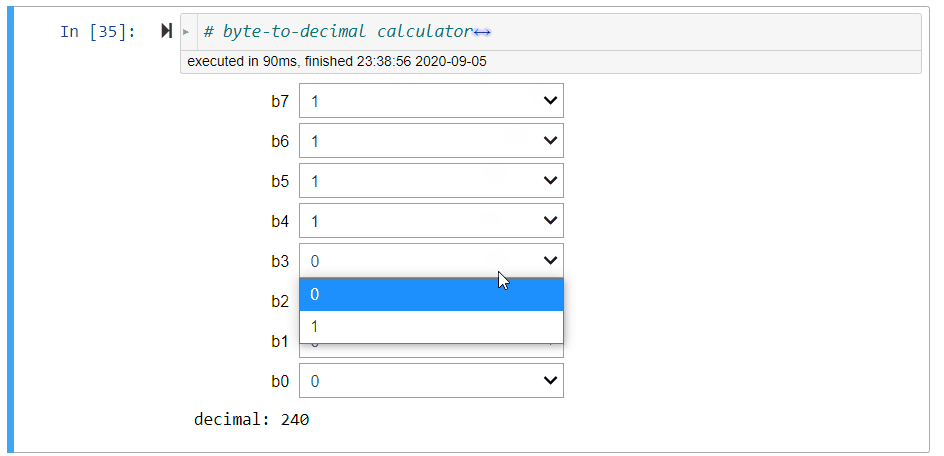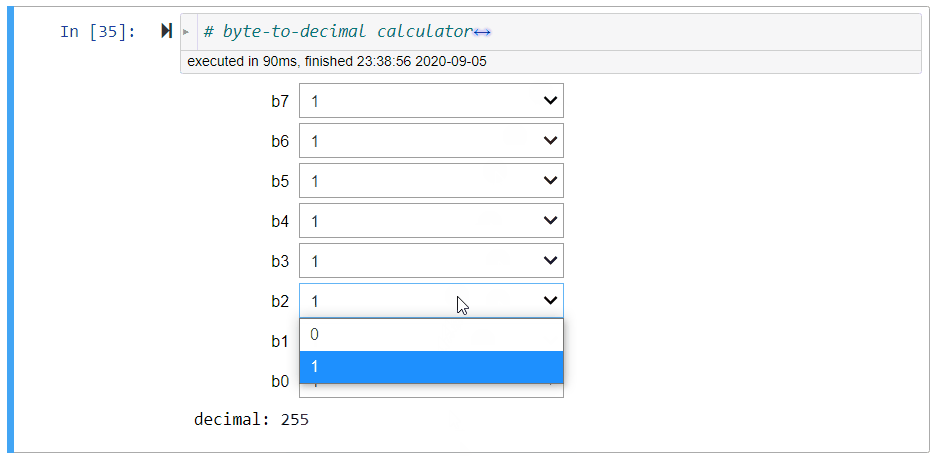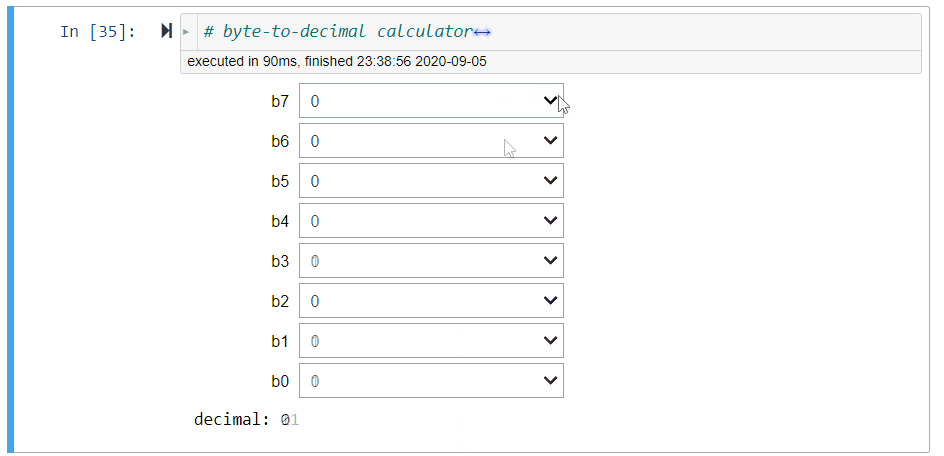
# ### Byte-to-Decimal Calculator
# + [markdown] slideshow={"slide_type": "-"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# Denote a binary number stored as a byte ($8$-bit) as
#
# $$
# b_7\circ b_6\circ b_5\circ b_4\circ b_3\circ b_2\circ b_1\circ b_0,
# $$
# where $\circ$ concatenates $b_i$'s together into a binary string.
# + [markdown] slideshow={"slide_type": "fragment"}
# The binary string can be converted to a decimal number by the formula
#
# $$
# b_7\cdot 2^7 + b_6\cdot 2^6 + b_5\cdot 2^5 + b_4\cdot 2^4 + b_3\cdot 2^3 + b_2\cdot 2^2 + b_1\cdot 2^1 + b_0\cdot 2^0.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# E.g., the binary string `'11111111'` is the largest integer represented by a byte:
#
# $$
# 2^7+2^6+2^5+2^4+2^3+2^2+2^1+2^0=255=2^8-1.
# $$
# + [markdown] slideshow={"slide_type": "fragment"}
# **Exercise** Assign to `decimal` the *integer* value represented by a byte.
# The byte is a sequence of bits assigned to the variables `b7,b6,b5,b4,b3,b2,b1,b0` as *characters*, either `'0'` or `'1'`.
# + deletable=false nbgrader={"cell_type": "code", "checksum": "771c9f92222e78d280a947cc2cd341bb", "grade": false, "grade_id": "byte_to_decimal", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "fragment"}
def byte_to_decimal(b7, b6, b5, b4, b3, b2, b1, b0):
# YOUR CODE HERE
raise NotImplementedError()
return decimal
# + code_folding=[0] deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "aba592363064888174547dbbad7818c0", "grade": true, "grade_id": "test-byte_to_decimal", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "fragment"}
# tests
def test_byte_to_decimal(decimal, b7, b6, b5, b4, b3, b2, b1, b0):
decimal_ = byte_to_decimal(b7, b6, b5, b4, b3, b2, b1, b0)
correct = decimal == decimal_ and isinstance(decimal_, int)
if not correct:
print(
f'{b7}{b6}{b5}{b4}{b3}{b2}{b1}{b0} should give {decimal} not {decimal_}.'
)
assert correct
test_byte_to_decimal(38, '0', '0', '1', '0', '0', '1', '1', '0')
test_byte_to_decimal(20, '0', '0', '0', '1', '0', '1', '0', '0')
test_byte_to_decimal(22, '0', '0', '0', '1', '0', '1', '1', '0')
# + code_folding=[0]
# byte-to-decimal calculator
bit = ['0', '1']
@interact(b7=bit, b6=bit, b5=bit, b4=bit, b3=bit, b2=bit, b1=bit, b0=bit)
def convert_byte_to_decimal(b7, b6, b5, b4, b3, b2, b1, b0):
print('decimal:', byte_to_decimal(b7, b6, b5, b4, b3, b2, b1, b0))
# + [markdown] slideshow={"slide_type": "subslide"}
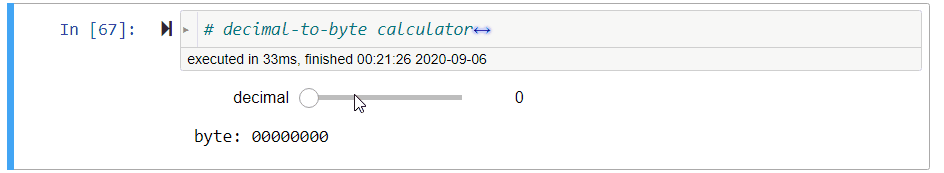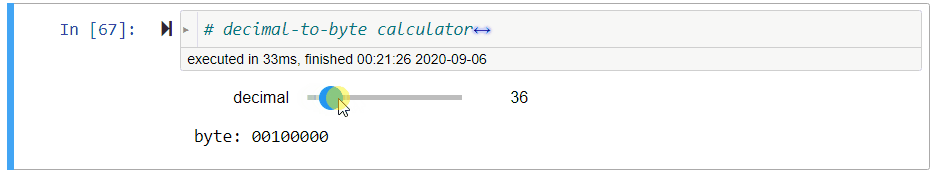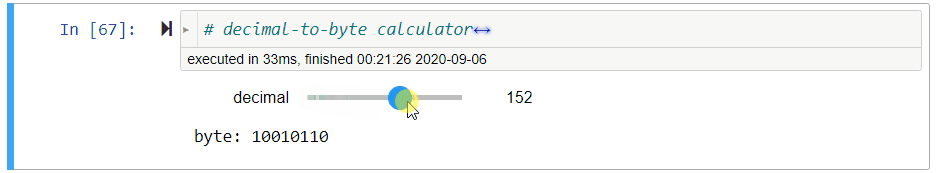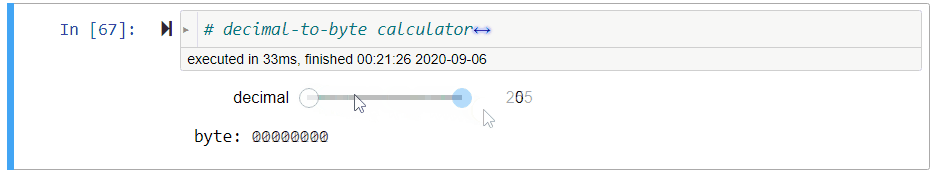
# ### Decimal-to-Byte Calculator
# + [markdown] slideshow={"slide_type": "-"}
# 
# + [markdown] slideshow={"slide_type": "subslide"}
# **Exercise** Assign to `byte` a *string of 8 bits* that represents the value of `decimal`, a non-negative decimal integer from $0$ to $2^8-1=255$.
# *Hint: Use `//` and `%`.*
# + deletable=false nbgrader={"cell_type": "code", "checksum": "9de8a3c79ad6c5af5db8a55ce33675e4", "grade": false, "grade_id": "decimal_to_byte", "locked": false, "schema_version": 3, "solution": true, "task": false} slideshow={"slide_type": "-"}
def decimal_to_byte(decimal):
# YOUR CODE HERE
raise NotImplementedError()
return byte
# + code_folding=[0] deletable=false editable=false nbgrader={"cell_type": "code", "checksum": "0c1b31ee5b25577be5ae3f3694976177", "grade": true, "grade_id": "test-decimal_to_byte", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false} slideshow={"slide_type": "fragment"}
# tests
def test_decimal_to_byte(byte,decimal):
byte_ = decimal_to_byte(decimal)
correct = byte == byte_ and isinstance(byte, str) and len(byte) == 8
if not correct:
print(
f'{decimal} should be represented as the byte {byte}, not {byte_}.'
)
assert correct
test_decimal_to_byte('01100111', 103)
test_decimal_to_byte('00000011', 3)
test_decimal_to_byte('00011100', 28)
# + code_folding=[0] slideshow={"slide_type": "fragment"}
# decimal-to-byte calculator
@interact(decimal=(0,255,1))
def convert_decimal_to_byte(decimal=0):
print('byte:', decimal_to_byte(decimal))
| Lab2/Calculators.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.7 64-bit (''transit_occupancy'': conda)'
# name: python3
# ---
import pandas as pd
import geopandas as gpd
import requests,json
from shapely import geometry, ops
import numpy as np
pd.set_option('display.max_columns', None)
def calculate_bearing(lat1, lng1, lat2, lng2):
"""
Calculate the compass bearing(s) between pairs of lat-lng points.
Vectorized function to calculate (initial) bearings between two points'
coordinates or between arrays of points' coordinates. Expects coordinates
in decimal degrees. Bearing represents angle in degrees (clockwise)
between north and the geodesic line from point 1 to point 2.
Parameters
----------
lat1 : float or numpy.array of float
first point's latitude coordinate
lng1 : float or numpy.array of float
first point's longitude coordinate
lat2 : float or numpy.array of float
second point's latitude coordinate
lng2 : float or numpy.array of float
second point's longitude coordinate
Returns
-------
bearing : float or numpy.array of float
the bearing(s) in decimal degrees
"""
# get the latitudes and the difference in longitudes, in radians
lat1 = np.radians(lat1)
lat2 = np.radians(lat2)
d_lng = np.radians(lng2 - lng1)
# calculate initial bearing from -180 degrees to +180 degrees
y = np.sin(d_lng) * np.cos(lat2)
x = np.cos(lat1) * np.sin(lat2) - np.sin(lat1) * np.cos(lat2) * np.cos(d_lng)
initial_bearing = np.degrees(np.arctan2(y, x))
# normalize to 0-360 degrees to get compass bearing
return initial_bearing % 360
df=gpd.read_file('USA_Tennessee.geojson')
davidson=df[df.County=='DAVIDSON'].copy()
osm=pd.read_csv('USA_Tennessee.csv')
davidson=davidson.merge(osm,on='XDSegID',how='left')
davidson['osm_way']=davidson.OSMWayIDs.apply(lambda x: x.split(';'))
davidson=davidson.drop(['OSMWayIDs', 'OSMWayDirections', 'WayStartOffset_m', 'WayEndOffset_m',
'WayStartOffset_percent', 'WayEndOffset_percent','AllTmcList','PrimaryTmc','RoadNumber','Country','State'],axis=1)
davidson=davidson.explode('osm_way')
davidson = davidson[['XDSegID','osm_way','Bearing','geometry']]
davidson.columns= davidson.columns.str.strip().str.lower()
davidson.to_pickle('davidson_osm_inrix.pkl')
davidson['bearings']=davidson.apply(lambda x: calculate_bearing(x.geometry.coords.xy[1][0], x.geometry.coords.xy[0][0], x.geometry.coords.xy[1][-1], x.geometry.coords.xy[0][-1]),axis=1)
davidson['osm_way']=davidson.osm_way.apply(int)
davidson=davidson.rename(columns={'osm_way':'osmid'})
edges=davidsonedges.merge(davidson,on='osmid')
edges=edges.drop(['travel_time', 'name', 'bridge', 'maxspeed',
'tunnel', 'junction', 'access', 'width', 'service', 'area',
'geometry_x','geometry_y'],axis=1)
edges
edges['bearing_diff']=edges.apply(lambda x: abs(x.bearing_x-x.bearings) >=90,axis=1)
edges
edges=edges.drop(edges[edges.bearing_diff].index)
edges=edges.rename(columns={'bearing_x':'bearing_osm','bearings':'bearing_inrix','bearing_y':'osm_direction'})
edges=edges.drop('bearing_diff',axis=1)
edges.to_parquet('nashville_osm_edges.parquet')
edges
davidsonmodel=pd.read_parquet('davidson_inrix_model_month_day_hour.parquet')
davidsonmodel['speed_mean']=davidsonmodel.speed_mean.apply(lambda x: int(x* 1.69093))
davidsonmodel
datapoints=davidsonmodel[['day','hour','month']].drop_duplicates()
datapoints
for row in datapoints.itertuples():
samplemodel=davidsonmodel[(davidsonmodel.day==row.day) & (davidsonmodel.month==row.month) & (davidsonmodel.hour==row.hour)]
edges2=edges.merge(samplemodel,on='xdsegid',how='left')
edges2['speed_kph']=edges2.apply(lambda x: x.speed_mean if not np.isnan(x.speed_mean) else x.speed_kph,axis=1)
edges2=edges2.groupby(['u','v','key'],as_index=False).agg({'speed_kph':'mean'})
edges2['speed_kph']=edges2.speed_kph.apply(int)
edges2[['u','v','speed_kph']].to_csv(f'segment_speed_data/segment_speed_data_{row.month}_{row.day}_{row.hour}.csv',index=False,header=False)
edges2
# !ls -lhrt segment_speed_data/*.csv
| tn-analysis/generatePerSegmentSpeedPerMonthPerDayTypePerHour.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import pandas as pd
import numpy as np
import math
import time
import argparse
import torch
from sklearn.model_selection import StratifiedKFold, train_test_split
from sklearn.preprocessing import StandardScaler
from torch.utils.data import DataLoader, Dataset
import torch.nn.functional as F
from torch.nn import Module
from torch import optim
import tqdm
import shutil
from torch.utils.tensorboard import SummaryWriter
import datetime
from multiprocessing import cpu_count
import matplotlib.pyplot as plt
import seaborn as sns
from sklearn.metrics import roc_curve, auc
from matplotlib.pylab import mpl
# %matplotlib qt
plt.rcParams['xtick.direction'] = 'in'
plt.rcParams['ytick.direction'] = 'in'
mpl.rcParams['axes.unicode_minus'] = False #显示负号
# + code_folding=[0, 66]
def classify_provider(features_path, label_path, n_splits, batch_size, num_workers, method='origin'):
# Amp,RiseT,Dur,Eny,RMS,Counts
with open(features_path, 'r') as f:
feature = np.array([i.split(',')[6:-4] for i in f.readlines()[1:]])
feature = feature.astype(np.float32)
with open(label_path, 'r') as f:
label = np.array([i.strip() for i in f.readlines()[1:]])
label = label.astype(np.float32).reshape(-1, 1)
label[np.where(label == 2)] = 0
ext = np.zeros([feature.shape[0], 1]).astype(np.float32)
ext[np.where(label == 0)[0].tolist()] = 1
label = np.concatenate((label, ext), axis=1)
df = pd.DataFrame(feature)
df.columns = ['Amp', 'RiseT', 'Dur', 'Eny', 'RMS', 'Counts']
df['Counts/Dur'] = df['Counts'] / df['Dur']
df['RiseT/Dur'] = df['RiseT'] / df['Dur']
df['Eny/Dur'] = df['Eny'] / df['Dur']
df['Amp*RiseT'] = df['Amp'] * df['RiseT']
if method == '10_select':
feature = df.values
elif method == '6_select':
feature = df[['Eny', 'Amp*RiseT', 'Dur', 'RMS', 'Counts/Dur', 'RiseT/Dur']].values
train_dfs = list()
val_dfs = list()
all_dfs = list()
if n_splits != 1:
skf = StratifiedKFold(n_splits=n_splits, shuffle=True, random_state=69)
for train_df_index, val_df_index in skf.split(feature, label[:, 0]):
train_dfs.append([feature[train_df_index], label[train_df_index, :]])
val_dfs.append([feature[val_df_index], label[val_df_index, :]])
else:
df_temp = train_test_split(feature, label, test_size=0.2, stratify=label, random_state=69)
train_dfs.append([df_temp[0], df_temp[2]])
val_dfs.append([df_temp[1], df_temp[3]])
all_dfs.append([np.concatenate((df_temp[0], df_temp[1]), axis=0),
np.concatenate((df_temp[2], df_temp[3]), axis=0)])
# print(len(train_dfs), len(val_dfs), len(all_dfs))
dataloaders = list()
for df_index, (train_df, val_df, all_df) in enumerate(zip(train_dfs, val_dfs, all_dfs)):
train_dataset = SteelClassDataset(train_df)
val_dataset = SteelClassDataset(val_df)
all_dataset = SteelClassDataset(all_df)
train_dataloader = DataLoader(train_dataset,
batch_size=batch_size,
num_workers=num_workers,
pin_memory=False,
shuffle=True)
val_dataloader = DataLoader(val_dataset,
batch_size=batch_size,
num_workers=num_workers,
pin_memory=False,
shuffle=False)
all_dataloader = DataLoader(all_dataset,
batch_size=batch_size,
num_workers=num_workers,
pin_memory=False,
shuffle=False)
dataloaders.append([train_dataloader, val_dataloader, all_dataloader])
return feature, label, dataloaders
class SteelClassDataset(Dataset):
def __init__(self, dataset):
super(SteelClassDataset, self).__init__()
self.feature = dataset[0]
self.label = dataset[1]
def __getitem__(self, idx):
x = self.feature[idx]
y = self.label[idx]
return x, y
def __len__(self):
return len(self.label)
# + code_folding=[0]
class Classify_model(Module):
def __init__(self, layer, method='origin', training=True):
super(Classify_model, self).__init__()
if method in ['origin', '6_select']:
self.linear1 = torch.nn.Linear(6, layer[0])
self.linear2 = torch.nn.Linear(layer[0], layer[1])
self.linear3 = torch.nn.Linear(layer[1], 2)
elif method == '10_select':
self.linear1 = torch.nn.Linear(10, layer[0])
self.linear2 = torch.nn.Linear(layer[0], layer[1])
self.linear3 = torch.nn.Linear(layer[1], layer[2])
self.linear4 = torch.nn.Linear(layer[2], 2)
self.relu = torch.nn.ReLU()
self.sigmoid = torch.nn.Sigmoid()
self.training = training
def forward(self, input, method):
if method in ['origin', '6_select']:
y = self.linear1(input)
y = self.relu(y)
y = self.linear2(y)
y = self.relu(y)
y = self.linear3(y)
elif method == '10_select':
y = self.linear1(input)
y = self.sigmoid(y)
y = F.dropout(y, 0.5, training=self.training)
y = self.linear2(y)
y = self.sigmoid(y)
y = F.dropout(y, 0.5, training=self.training)
y = self.linear3(y)
y = self.sigmoid(y)
y = F.dropout(y, 0.5, training=self.training)
y = self.linear4(y)
return y
# + code_folding=[0]
class Solver():
def __init__(self, model, method):
self.model = model
self.method = method
def forward(self, x):
outputs = self.model(x, self.method)
return outputs
def cal_loss(self, targets, predicts, criterion):
return criterion(predicts, targets)
def backword(self, optimizer, loss):
loss.backward()
optimizer.step()
optimizer.zero_grad()
def save_checkpoint(self, save_path, state, is_best):
# torch.save(state, save_path)
if is_best:
torch.save(state, save_path)
print('Saving Best Model.')
# save_best_path = save_path.replace('.pth', '_best.pth')
# shutil.copyfile(save_path, save_best_path)
def load_checkpoint(self, load_path):
if os.path.isfile(load_path):
checkpoint = torch.load(load_path, map_location='cpu')
# self.model.module.load_state_dict(checkpoint['state_dict'])
print('Successfully Loaded from %s' % (load_path))
return self.model
else:
raise FileNotFoundError(
"Can not find weight file in {}".format(load_path))
# + code_folding=[0]
if __name__ == "__main__":
method = 'origin'
features_path = './pri_database.txt'
label_path = './label.txt'
weight_path = './checkpoints/origin/2020-11-16T14-26-21-16-classify/classify_fold0_origin_0.973121.pth'
n_splits = 1
batch_size = 16
layer = [100, 80, 10]
feature, _, dataloaders = classify_provider(
features_path, label_path, n_splits, batch_size, 0, method)
model = Classify_model(layer, method, training=False)
model.eval()
model.train(False)
solver = Solver(model, method)
checkpoint = torch.load(weight_path, map_location=torch.device('cpu'))
model.load_state_dict(checkpoint['state_dict'], False)
tbar = tqdm.tqdm(dataloaders[0][2], ncols=80)
res, res_sigmoid, res_01, Label = [], [], [], []
with torch.no_grad():
for i, (x, labels) in enumerate(tbar):
labels_row = solver.forward(x)
labels_predict = torch.sigmoid(labels_row)
predict = (labels_predict > 0.5).float()
res_01.append(predict.data.cpu().numpy())
res_sigmoid.append(labels_predict.data.cpu().numpy())
res.append(labels_row.data.cpu().numpy())
Label.append(labels.data.cpu().numpy())
correct.append(sum(predict+labels == 2).data.cpu().numpy())
# + code_folding=[0]
def metric(p, t):
tp = ((p[:,1] + t[:,1]) == 2).sum(0) # True positives
tn = ((p[:,0] + t[:,0]) == 2).sum(0) # True negatives
total_pos = t[:, 1].sum(0)
total_neg = t[:, 0].sum(0)
fp = total_neg - tn # 将负类错误预测为正类数
fn = total_pos - tp # 将正类错误预测为负类数
# 各个类别预测正确的正样本、负样本数目
class_neg_accuracy = tn / t[:, 0].sum(0)
class_pos_accuracy = tp / t[:, 1].sum(0)
return class_pos_accuracy, class_neg_accuracy, tp, total_pos, tn, total_neg, fp, fn
# -
pre = np.concatenate([i for i in res_01])
label = np.concatenate([i for i in Label])
class_pos_accuracy, class_neg_accuracy, tp, total_pos, tn, total_neg, fp, fn = metric(pre, label)
print('{:.5f}, {:.5f}, {}/{:.0f}, {}/{:.0f}, {:.0f}, {:.0f}'.
format(class_pos_accuracy, class_neg_accuracy, tp, total_pos, tn, total_neg, fp, fn))
# +
_, _, tp, _, tn, _, fp, fn = metric(pre, label)
cm = np.zeros([2, 2])
cm[0][0], cm[0][1], cm[1][0], cm[1][1] = tp, fn, fp, tn
f,ax=plt.subplots(figsize=(2.5, 2))
sns.heatmap(cm,annot=True,ax=ax, fmt='.20g') #画热力图
# ax.set_title('Confusion Matrix') #标题
# ax.set_xlabel('Predicted label') #x轴
# ax.set_ylabel('True label') #y轴
ax.xaxis.set_ticks_position('top')
ax.yaxis.set_ticks_position('left')
# ax.set_xticks(['Class 1', 'Class 2'])
# ax.set_yticks(['Class 1', 'Class 2'])
ax.tick_params(bottom=False,top=False,left=False,right=False)
# -
# ## ROC curve
# +
y_pre = np.concatenate([i for i in res_sigmoid])
fpr, tpr, thersholds = roc_curve(label[: ,0], y_pre[: ,0])
roc_auc = auc(fpr, tpr)
plt.figure(figsize=[8, 6])
plt.plot(fpr, tpr, 'k--', color='navy', label='ROC (area = {0:.3f})'.format(roc_auc), lw=2)
ax = plt.gca()
ax.spines['bottom'].set_linewidth(2)
ax.spines['left'].set_linewidth(2)
ax.spines['right'].set_linewidth(2)
ax.spines['top'].set_linewidth(2)
font_legend = {'family':'DejaVu Sans','weight':'normal','size':15}
font_label = {'family': 'DejaVu Sans', 'weight': 'bold', 'size': 20}
font_title = {'family': 'DejaVu Sans', 'weight': 'bold', 'size': 30}
# 设置坐标刻度值的大小以及刻度值的字体
plt.tick_params(labelsize=16)
labels = ax.get_xticklabels() + ax.get_yticklabels()
[label.set_fontname('DejaVu Sans') for label in labels]
# plt.xlim(-5, 150)
# plt.xticks(range(0, 149, 29), labels=['1', '30', '60', '90', '120', '150'])
plt.ylabel('False Positive Rate', font_label)
plt.xlabel('True Positive Rate', font_label)
plt.legend(loc="lower right", prop=font_legend)
# -
# ## Export ML
def cal_ML(tmp, features_path, cls_1, cls_2):
tmp_1, tmp_2 = sorted(tmp[cls_1]), sorted(tmp[cls_2])
N1, N2 = len(tmp_1), len(tmp_2)
ML_y1, ML_y2 = [], []
Error_bar1, Error_bar2 = [] ,[]
for j in range(N1):
valid_x = sorted(tmp_1)[j:]
E0 = valid_x[0]
Sum = np.sum(np.log(valid_x/E0))
N_prime = N1 - j
alpha = 1 + N_prime / Sum
error_bar = (alpha - 1) / pow(N_prime, 0.5)
ML_y1.append(alpha)
Error_bar1.append(error_bar)
for j in range(N2):
valid_x = sorted(tmp_2)[j:]
E0 = valid_x[0]
Sum = np.sum(np.log(valid_x/E0))
N_prime = N2 - j
alpha = 1 + N_prime / Sum
error_bar = (alpha - 1) / pow(N_prime, 0.5)
ML_y2.append(alpha)
Error_bar2.append(error_bar)
with open(features_path[:-4] + '_1 ' + 'Energy' + '_ML.txt', 'w') as f:
f.write('Energy, ε, Error bar\n')
for j in range(len(ML_y1)):
f.write('{}, {}, {}\n'.format(sorted(tmp_1)[j], ML_y1[j], Error_bar1[j]))
with open(features_path[:-4] + '_2 ' + 'Energy' + '_ML.txt', 'w') as f:
f.write('Energy, ε, Error bar\n')
for j in range(len(ML_y2)):
f.write('{}, {}, {}\n'.format(sorted(tmp_2)[j], ML_y2[j], Error_bar2[j]))
predict = np.concatenate([i for i in res_01])
cls_1 = predict[:, 0] == 1
cls_2 = predict[:, 1] == 1
features_path = r'6_2_select.txt'
cal_ML(feature[:, 3], features_path, cls_1, cls_2)
sum(cls_1), sum(cls_2), sum(label[:, 0] == 1), sum(label[:, 1] == 1)
# ## SVM
from sklearn.svm import SVC
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
from sklearn.ensemble import RandomForestClassifier
import pandas as pd
import seaborn as sns
from sklearn.metrics import classification_report, accuracy_score, precision_score, recall_score, f1_score, cohen_kappa_score, roc_curve, auc, confusion_matrix
# + code_folding=[0, 36, 45]
def plot_norm(ax, xlabel=None, ylabel=None, zlabel=None, title=None, x_lim=[], y_lim=[], z_lim=[], legend=True, grid=False,
legend_loc='upper left', font_color='black', legendsize=11, labelsize=14, titlesize=15, ticksize=13, linewidth=2):
ax.spines['bottom'].set_linewidth(linewidth)
ax.spines['left'].set_linewidth(linewidth)
ax.spines['right'].set_linewidth(linewidth)
ax.spines['top'].set_linewidth(linewidth)
# 设置坐标刻度值的大小以及刻度值的字体 Arial
ax.tick_params(which='both', width=linewidth, labelsize=ticksize, colors=font_color)
labels = ax.get_xticklabels() + ax.get_yticklabels()
[label.set_fontname('Arial') for label in labels]
font_legend = {'family': 'Arial', 'weight': 'normal', 'size': legendsize}
font_label = {'family': 'Arial', 'weight': 'bold', 'size': labelsize, 'color':font_color}
font_title = {'family': 'Arial', 'weight': 'bold', 'size': titlesize, 'color':font_color}
if x_lim:
ax.set_xlim(x_lim[0], x_lim[1])
if y_lim:
ax.set_ylim(y_lim[0], y_lim[1])
if z_lim:
ax.set_zlim(z_lim[0], z_lim[1])
if legend:
plt.legend(loc=legend_loc, prop=font_legend)
if grid:
ax.grid(ls='-.')
if xlabel:
ax.set_xlabel(xlabel, font_label)
if ylabel:
ax.set_ylabel(ylabel, font_label)
if zlabel:
ax.set_zlabel(zlabel, font_label)
if title:
ax.set_title(title, font_title)
plt.tight_layout()
def plot_confmat(tn, fp, fn, tp):
cm = np.zeros([2, 2])
cm[0][0], cm[0][1], cm[1][0], cm[1][1] = tn, fp, fn, tp
f, ax=plt.subplots(figsize=(2.5, 2))
sns.heatmap(cm,annot=True, ax=ax, fmt='.20g') #画热力图
ax.xaxis.set_ticks_position('top')
ax.yaxis.set_ticks_position('left')
ax.tick_params(bottom=False,top=False,left=False,right=False)
def print_res(model, x_pred, y_true):
target_pred = model.predict(x_pred)
true = np.sum(target_pred == y_true)
print('预测对的结果数目为:', true)
print('预测错的的结果数目为:', y_true.shape[0]-true)
print('使用SVM预测的准确率为:',
accuracy_score(y_true, target_pred))
print('使用SVM预测的精确率为:',
precision_score(y_true, target_pred))
print('使用SVM预测的召回率为:',
recall_score(y_true, target_pred))
print('使用SVM预测的F1值为:',
f1_score(y_true, target_pred))
print('使用SVM预测b的Cohen’s Kappa系数为:',
cohen_kappa_score(y_true, target_pred))
print('使用SVM预测的分类报告为:','\n',
classification_report(y_true, target_pred))
tn, fp, fn, tp = confusion_matrix(y_true, target_pred).ravel()
plot_confmat(tn, fp, fn, tp)
return target_pred
# -
# ### Dislocation
# +
fold = r'C:\Users\jonah\Desktop\Ni_dislocation.csv'
data = pd.read_csv(fold).astype(np.float32)
feature = data.iloc[:, :-1].values
label = np.array(data.iloc[:, -1].tolist()).reshape(-1, 1)
# ext = np.zeros([label.shape[0], 1]).astype(np.float32)
# ext[np.where(label == 0)[0]] = 1
# label = np.concatenate((label, ext), axis=1)
df_temp = train_test_split(feature, label, test_size=0.2, stratify=label, random_state=69)
stdScaler = StandardScaler().fit(df_temp[0])
trainStd = stdScaler.transform(df_temp[0])
testStd = stdScaler.transform(df_temp[1])
svm = SVC(max_iter=200, random_state=100).fit(trainStd, df_temp[2].reshape(-1))
print('建立的SVM模型为:\n', svm)
rf = RandomForestClassifier(max_depth=10, random_state=100).fit(trainStd, df_temp[2].reshape(-1))
print('建立的RF模型为:\n', rf)
# -
target_pred_svm = print_res(svm, testStd, df_temp[3].reshape(-1))
# + code_folding=[]
target_pred_rf = print_res(rf, testStd, df_temp[3].reshape(-1))
# +
fpr_svm, tpr_svm, thersholds_svm = roc_curve(df_temp[3].reshape(-1), target_pred_svm)
roc_auc_svm = auc(fpr_svm, tpr_svm)
fpr_rf, tpr_rf, thersholds_rf = roc_curve(df_temp[3].reshape(-1), target_pred_rf)
roc_auc_rf = auc(fpr_rf, tpr_rf)
fig = plt.figure(figsize=[6, 3.9])
ax = plt.subplot()
ax.plot(fpr_svm, tpr_svm, 'k--', color='navy', label='SVM (area = {0:.3f})'.format(roc_auc_svm), lw=2)
ax.plot(fpr_rf, tpr_rf, 'k--', color='green', label='RF (area = {0:.3f})'.format(roc_auc_rf), lw=2)
plot_norm(ax, 'True Positive Rate', 'False Positive Rate', legend_loc='lower right')
# -
fold = r'C:\Users\jonah\Desktop\Nano_Ni_3_cnts_4.csv'
data = pd.read_csv(fold).astype(np.float32)
nano_ni = data.values
stdScaler = StandardScaler().fit(nano_ni)
trainStd = stdScaler.transform(nano_ni)
target_pred = svm.predict(trainStd)
sum(target_pred), target_pred.shape
fold = r'C:\Users\jonah\Desktop\Nano_Ni_3_cnts_4.csv'
data = pd.read_csv(fold).astype(np.float32)
nano_ni = data.values
stdScaler = StandardScaler().fit(nano_ni)
trainStd = stdScaler.transform(nano_ni)
target_pred = rf.predict(trainStd)
sum(target_pred), target_pred.shape
# ### Twinning
# +
fold = r'C:\Users\jonah\Desktop\Ni_twinning.csv'
data = pd.read_csv(fold).astype(np.float32)
feature = data.iloc[:, :-1].values
label = np.array(data.iloc[:, -1].tolist()).reshape(-1, 1)
df_temp = train_test_split(feature, label, test_size=0.2, stratify=label, random_state=69)
stdScaler = StandardScaler().fit(df_temp[0])
trainStd = stdScaler.transform(df_temp[0])
testStd = stdScaler.transform(df_temp[1])
svm = SVC(max_iter=200, random_state=100).fit(trainStd, df_temp[2].reshape(-1))
print('建立的SVM模型为:\n', svm)
rf = RandomForestClassifier(max_depth=10, random_state=100).fit(trainStd, df_temp[2].reshape(-1))
print('建立的RF模型为:\n', rf)
# -
target_pred_svm = print_res(svm, testStd, df_temp[3].reshape(-1))
p, r = 0.9889, 0.9828
2*p*r/(p+r)
# + code_folding=[]
target_pred_rf = print_res(rf, testStd, df_temp[3].reshape(-1))
# +
fpr_svm, tpr_svm, thersholds_svm = roc_curve(df_temp[3].reshape(-1), target_pred_svm)
roc_auc_svm = auc(fpr_svm, tpr_svm)
fpr_rf, tpr_rf, thersholds_rf = roc_curve(df_temp[3].reshape(-1), target_pred_rf)
roc_auc_rf = auc(fpr_rf, tpr_rf)
fig = plt.figure(figsize=[6, 3.9])
ax = plt.subplot()
ax.plot(fpr_svm, tpr_svm, 'k--', color='navy', label='SVM (area = {0:.3f})'.format(roc_auc_svm), lw=2)
ax.plot(fpr_rf, tpr_rf, 'k--', color='green', label='RF (area = {0:.3f})'.format(roc_auc_rf), lw=2)
plot_norm(ax, 'True Positive Rate', 'False Positive Rate', legend_loc='lower right')
# -
fold = r'C:\Users\jonah\Desktop\Nano_Ni_3_cnts_4_~dislocation.csv'
data = pd.read_csv(fold).astype(np.float32)
nano_ni = data.values
stdScaler = StandardScaler().fit(nano_ni)
trainStd = stdScaler.transform(nano_ni)
target_pred = rf.predict(trainStd)
sum(target_pred), target_pred.shape
# ### Crack
# +
fold = r'C:\Users\jonah\Desktop\Ni_twinning&crack.csv'
data = pd.read_csv(fold).astype(np.float32)
feature = data.iloc[:, :-1].values
label = np.array(data.iloc[:, -1].tolist()).reshape(-1, 1)
df_temp = train_test_split(feature, label, test_size=0.2, stratify=label, random_state=69)
stdScaler = StandardScaler().fit(df_temp[0])
trainStd = stdScaler.transform(df_temp[0])
testStd = stdScaler.transform(df_temp[1])
svm = SVC(max_iter=200, random_state=100).fit(trainStd, df_temp[2].reshape(-1))
print('建立的SVM模型为:\n', svm)
rf = RandomForestClassifier(max_depth=10, random_state=100).fit(trainStd, df_temp[2].reshape(-1))
print('建立的RF模型为:\n', rf)
# -
target_pred_svm = print_res(svm, testStd, df_temp[3].reshape(-1))
# + code_folding=[]
target_pred_rf = print_res(rf, testStd, df_temp[3].reshape(-1))
# +
fpr_svm, tpr_svm, thersholds_svm = roc_curve(df_temp[3].reshape(-1), target_pred_svm)
roc_auc_svm = auc(fpr_svm, tpr_svm)
fpr_rf, tpr_rf, thersholds_rf = roc_curve(df_temp[3].reshape(-1), target_pred_rf)
roc_auc_rf = auc(fpr_rf, tpr_rf)
fig = plt.figure(figsize=[6, 3.9])
ax = plt.subplot()
ax.plot(fpr_svm, tpr_svm, 'k--', color='navy', label='SVM (area = {0:.3f})'.format(roc_auc_svm), lw=2)
ax.plot(fpr_rf, tpr_rf, 'k--', color='green', label='RF (area = {0:.3f})'.format(roc_auc_rf), lw=2)
plot_norm(ax, 'True Positive Rate', 'False Positive Rate', legend_loc='lower right')
| 1/NN.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/marjpri/deep-learning-v2-pytorch/blob/master/test.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="zTLkQ6rntv1P" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="72e6e52f-fe2b-4cda-ebad-2747f5172d81"
# !pwd
# + id="TrtHHqpVug-e" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="4f243763-b6f2-45ed-8794-f205baa38cad"
# cd sample_data
# + id="aLCRuB1gupSh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="fc202768-1f71-4602-ddfc-461129e0077a"
# !ls
# + id="XxxBluDwuthL" colab_type="code" colab={}
# !git clone https://github.com/marjpri/deep-learning-v2-pytorch.git
# + id="V6F94EE7vXuO" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="7f284efa-8aee-4414-eaad-80a62a66daba"
# !pwd
# + id="h79fvLmw248A" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="8f2b3141-3ae1-4577-958c-d8c34850a13e"
# cd /content/sample_data/deep-learning-v2-pytorch/
# + id="f1POfU8i3bUW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 251} outputId="8c7578c8-2b78-4805-deeb-24038133204b"
# !ls
# + id="87bWlwmd3eEQ" colab_type="code" colab={}
# #!rm -rf attention/
# + id="hVN8QF1r34kF" colab_type="code" colab={}
from zipfile import ZipFile
file_name = 'data.zip'
#opening the zip file in read mode
with ZipFile (filename, 'r') as zip:
print ('extracting of the file now...')
zip.extractall()
print ('Done')
# + id="HQeBDt9v5KRg" colab_type="code" colab={}
#:rm -rf data.zip
# + id="YhEx5dnP5tyE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="25205c41-922e-4976-e9cb-637e422eb5c7"
# cd /
# + id="r2v-ZfVx5vPs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 133} outputId="cfc1066d-f032-4c93-ecea-50319a9d8e30"
:ls
# + id="iajFEu4V5y_H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 53} outputId="d9c0b92d-334f-46f7-a7ae-96e063c10489"
# !ls
# + id="GTity7z15-fb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="dcbbc356-c974-4f31-932d-53bbbf34cc3d"
# cd content/
# + id="17qMPdVt6R8t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="dc35e3b6-5dbb-48c5-c043-0fbd9ef737be"
# cd sample_data/
# + id="4O4-AtXl6W4E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 71} outputId="69ddb7dc-cfc2-4710-ada8-f308ecb9dfb9"
# !ls
# + id="2tMANXud6ZtH" colab_type="code" colab={}
| test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.2 64-bit
# language: python
# name: python39264bit19b4f0c4d0b341bebd274d780a05c1c2
# ---
from matplotlib import pyplot as plt
import random
from time import time
import secrets
def frequency(bit_string: str, seq: int, n: int):
data = {}
for i in range(2**seq):
x = bin(i)[2:]
if len(x) <seq:
x = '0'*(seq-len(x)) + x
data[x] = bit_string.count(x)
return data
# -----------------
# ## Usando random.getrandbits()
n = 100000000
x = bin(random.getrandbits(n))[2:] # gerando uma uma bit string de n bit
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 3))
seq = 1
for axn in (ax1, ax2):
data = frequency(x, seq, n)
axn.stem(data.keys(), data.values())
axn.set_xlabel('Subsequences')
axn.set_ylabel('Frequency')
seq+=1
# +
fig, (ax3, ax4) = plt.subplots(1, 2, figsize=(15, 3))
seq = 3
for axn in (ax3, ax4):
data = frequency(x, seq, n)
axn.stem(data.keys(), data.values())
axn.set_xlabel('Subsequences')
axn.set_ylabel('Frequency')
seq+=1
# -
# -------------------------
# ## Usando secrets.randbits()
n = 100000000
x = bin(secrets.randbits(n))[2:] # gerando uma uma bit string de n bit
# +
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 3))
seq = 1
for axn in (ax1, ax2):
data = frequency(x, seq, n)
axn.stem(data.keys(), data.values())
axn.set_xlabel('Subsequences')
axn.set_ylabel('Frequency')
seq+=1
# +
fig, (ax3, ax4) = plt.subplots(1, 2, figsize=(15, 3))
seq = 3
for axn in (ax3, ax4):
data = frequency(x, seq, n)
axn.stem(data.keys(), data.values())
axn.set_xlabel('Subsequences')
axn.set_ylabel('Frequency')
seq+=1
# +
fig, ax5 = plt.subplots(1, figsize=(15, 3))
seq = 5
data = frequency(x, seq, n)
ax5.stem(data.keys(), data.values())
ax5.set_xlabel('Subsequences')
ax5.set_ylabel('Frequency')
# -
# ___________
#
# ## "random" bit sting gerada por humano
# ### bit string length = 776
# +
x = """10101101010101000100101001001001010101010101010101001010101010101010100101010101010101100000001101011101
01010001001010010101000101010100110101011101010101001101010101010001011101001010101001011010101011010101010110
10101010101010101101010101101010110101010110101101010110101101011001001001010000000000101010110101111111101001
0101010101000100001010101001011110000000101001010101011110000100101010000001010100101101110010100111000100001111
0000011010010000001011100001011100001010100101010001010100101010101111100001100101001010100010101000001100001010
1010101101010101010010101010101001011110100100100101001001010110101000011001001011010101001110101001010001010
01010010101001010010101111010000001011100000010100101011010010100101000010101011100101000100101001000101010100010"""
n = len(x)
print(len(x)) # tamanho de x
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 3))
seq = 3
for axn in (ax1, ax2):
data = frequency(x, seq, n)
axn.stem(data.keys(), data.values())
axn.set_xlabel('Subsequences')
axn.set_ylabel('Frequency')
for k in data.keys():
print(f'data[{k}]: {data[k]}')
seq+=1
# -
# ## secrets.randbit()
# ### bit string lenght = 776
# +
n = 776
x = bin(random.getrandbits(n))[2:]
print(len(x)) # tamanho de x
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(15, 3))
seq = 3
for axn in (ax1, ax2):
data = frequency(x, seq, n)
axn.stem(data.keys(), data.values())
axn.set_xlabel('Subsequences')
axn.set_ylabel('Frequency')
for k in data.keys():
print(f'data[{k}]: {data[k]}')
seq+=1
| Frequency-Stability-Property.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <!DOCTYPE html>
# <html>
# <body>
# <div align="center">
# <h3>Prepared by <NAME></h3>
#
# <h1>Data Visualization with Matplotlib</h1>
#
# <h3>Connect with me on - <a href="https://www.linkedin.com/in/anushka-bajpai-423412ab/ ">LinkedIn</h3></div></div>
#
# </div>
# </body>
# </html>
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
# %matplotlib inline
# # Area Plot
x = np.arange(1,31)
y = np.random.normal(10,11,size=30)
y = np.square(y)
plt.figure(figsize=(16,6))
plt.plot(x,y)
plt.fill_between(x, y)
plt.show()
# #### Changing Fill Color
# +
x = np.arange(1,31)
y = np.random.normal(10,11,size=30)
y = np.square(y)
plt.figure(figsize=(16,6))
plt.fill_between( x, y, color="#baf1a1") # #Changing Fill color
plt.plot(x, y, color='#7fcd91') # Color on edges
plt.title("$ Area $ $ chart $" , fontsize = 16)
plt.xlabel("$X$" , fontsize = 16)
plt.ylabel("$Y$" , fontsize = 16)
plt.show()
# -
# #### Changing Fill Color and its transperancy
# +
x = np.arange(1,31)
y = np.random.normal(10,11,size=30)
y = np.square(y)
plt.figure(figsize=(16,6))
plt.fill_between( x, y, color="#C8D700" , alpha = 0.3) # Changing transperancy using Alpha parameter
plt.plot(x, y, color='#36BD00')
plt.title("$ Area $ $ chart $" , fontsize = 16)
plt.xlabel("$X$" , fontsize = 16)
plt.ylabel("$Y$" , fontsize = 16)
plt.show()
# +
x = np.arange(1,51)
y = np.random.normal(1,5,size=50)
y = np.square(y)
plt.figure(figsize=(16,6))
plt.fill_between( x, y, color="#5ac8fa", alpha=0.4)
plt.plot(x, y, color="blue", alpha=0.6) # Bold line on edges
plt.title("$ Area $ $ chart $" , fontsize = 16)
plt.xlabel("$X$" , fontsize = 16)
plt.ylabel("$Y$" , fontsize = 16)
plt.show()
plt.figure(figsize=(16,6))
plt.fill_between( x, y, color="#5ac8fa", alpha=0.4)
plt.plot(x, y, color="blue", alpha=0.2) # Less stronger line on edges
plt.title("$ Area $ $ chart $" , fontsize = 14)
plt.xlabel("$X$" , fontsize = 14)
plt.ylabel("$Y$" , fontsize = 14)
plt.show()
# -
# #### Stacked Area plot
# +
x=np.arange(1,6)
y1 = np.array([1,5,9,13,17])
y2 = np.array([2,6,10,14,16])
y3 = np.array([3,7,11,15,19])
y4 = np.array([4,8,12,16,20])
plt.figure(figsize=(8,6))
plt.stackplot(x,y1,y2,y3,y4, labels=['Y1','Y2','Y3','Y4'])
plt.legend(loc='upper left')
plt.show()
# -
x=np.arange(1,6)
y=[ [1,5,9,13,17], [2,6,10,14,16], [3,7,11,15,19] , [4,8,12,16,20] ]
plt.figure(figsize=(8,6))
plt.stackplot(x,y , labels=['Y1','Y2','Y3','Y4'])
plt.legend(loc='upper left')
plt.show()
x=np.arange(1,7)
y=[ [1,5,9,3,17,1], [2,6,10,4,16,2], [3,7,11,5,19,1] , [4,8,12,6,20,2] ]
plt.figure(figsize=(10,6))
plt.stackplot(x,y , labels=['Y1','Y2','Y3','Y4'])
plt.legend(loc='upper left')
plt.show()
# #### Changing Fill Color and its transperancy in Stacked Plot
# +
x=np.arange(1,7)
y=[ [1,5,9,3,17,1], [2,6,10,4,16,2], [3,7,11,5,19,1] , [4,8,12,6,20,2] ]
plt.figure(figsize=(11,6))
plt.stackplot(x,y , labels=['Y1','Y2','Y3','Y4'] , colors= ["#00b159" , "#ffc425", "#f37735", "#ff3b30"])
plt.legend(loc='upper left')
plt.show()
plt.figure(figsize=(11,6))
plt.stackplot(x,y, labels=['Y1','Y2','Y3','Y4'], colors= ["#00b159" , "#ffc425", "#f37735", "#ff3b30"], alpha=0.7 )
plt.legend(loc='upper left')
plt.show()
plt.figure(figsize=(11,6))
plt.stackplot(x,y, labels=['Y1','Y2','Y3','Y4'], colors= ["#00b159" , "#ffc425", "#f37735", "#ff3b30"], alpha=0.5 )
plt.legend(loc='upper left')
plt.show()
| Matplotlib/.ipynb_checkpoints/6. Area Chart-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # jammer
# ## 03 Adapt the Starfish `grid_tools.py` to accept the Marley grids
# ### I- Experiment
#
# <NAME>
# Friday, March 31, 2017
import numpy as np
import matplotlib.pyplot as plt
% matplotlib inline
% config InlineBackend.figure_format = 'retina'
import seaborn as sns
import pandas as pd
from os import listdir
import astropy.units as u
import astropy.constants as C
# SpeX prism library resolution is $75-200$. We'll pick 130. The resolution changes significantly from one end to the other... this could be a significant problem.
dv = C.c/130.0
dv.to(u.km/u.s)
import Starfish
from Starfish.grid_tools import MarleyGridInterface as Marley
mygrid = Marley()
my_params = np.array([525, 4.25])
flux, hdr = mygrid.load_flux(my_params)
flux
plt.plot(mygrid.wl, flux)
# Woohoo, it works!
| notebooks/jammer_03-01_Adapt_Starfish_grid_tools_to_marley_grids.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
import bayesnet as bn
np.random.seed(1234)
# -
x_train = np.linspace(-3, 3, 10)[:, None]
y_train = np.cos(x_train) + np.random.normal(0, 0.1, size=x_train.shape)
class BayesianNetwork(bn.Network):
def __init__(self, n_input, n_hidden, n_output):
super().__init__(
w1_mu=np.zeros((n_input, n_hidden)),
w1_s=np.zeros((n_input, n_hidden)),
b1_mu=np.zeros(n_hidden),
b1_s=np.zeros(n_hidden),
w2_mu=np.zeros((n_hidden, n_output)),
w2_s=np.zeros((n_hidden, n_output)),
b2_mu=np.zeros(n_output),
b2_s=np.zeros(n_output)
)
def __call__(self, x, y=None):
self.qw1 = bn.random.Gaussian(
self.w1_mu, bn.softplus(self.w1_s),
p=bn.random.Gaussian(0, 1)
)
self.qb1 = bn.random.Gaussian(
self.b1_mu, bn.softplus(self.b1_s),
p=bn.random.Gaussian(0, 1)
)
self.qw2 = bn.random.Gaussian(
self.w2_mu, bn.softplus(self.w2_s),
p=bn.random.Gaussian(0, 1)
)
self.qb2 = bn.random.Gaussian(
self.b2_mu, bn.softplus(self.b2_s),
p=bn.random.Gaussian(0, 1)
)
h = bn.tanh(x @ self.qw1.draw() + self.qb1.draw())
mu = h @ self.qw2.draw() + self.qb2.draw()
self.py = bn.random.Gaussian(mu, 0.1, data=y)
if y is None:
return self.py.draw().value
# +
model = BayesianNetwork(1, 20, 1)
optimizer = bn.optimizer.Adam(model, 0.1)
optimizer.set_decay(0.9, 100)
for _ in range(10000):
model.clear()
model(x_train, y_train)
elbo = model.elbo()
elbo.backward()
optimizer.update()
# -
x = np.linspace(-3, 3, 1000)[:, None]
plt.scatter(x_train, y_train)
y = [model(x) for _ in range(100)]
y_mean = np.mean(y, axis=0)
y_std = np.std(y, axis=0)
plt.plot(x, y_mean, c="orange")
plt.fill_between(x.ravel(), (y_mean - y_std).ravel(), (y_mean + y_std).ravel(), color="orange", alpha=0.2)
plt.show()
parameter = []
parameter.extend(list(model.w1_mu.value.ravel()))
parameter.extend(list(model.b1_mu.value.ravel()))
parameter.extend(list(model.w2_mu.value.ravel()))
parameter.extend(list(model.b2_mu.value.ravel()))
plt.plot(parameter)
class ARDNetwork(bn.Network):
def __init__(self, n_input, n_hidden, n_output):
super().__init__(
w1_mu=np.zeros((n_input, n_hidden)),
w1_s=np.zeros((n_input, n_hidden)),
shape_w1=np.ones((n_input, n_hidden)),
rate_w1=np.ones((n_input, n_hidden)),
b1_mu=np.zeros(n_hidden),
b1_s=np.zeros(n_hidden),
shape_b1=np.ones(n_hidden),
rate_b1=np.ones(n_hidden),
w2_mu=np.zeros((n_hidden, n_output)),
w2_s=np.zeros((n_hidden, n_output)),
shape_w2=np.ones((n_hidden, n_output)),
rate_w2=np.ones((n_hidden, n_output)),
b2_mu=np.zeros(n_output),
b2_s=np.zeros(n_output),
shape_b2=np.ones(n_output),
rate_b2=np.ones(n_output)
)
def __call__(self, x, y=None):
ptau_w1 = bn.random.Gamma(1., 1e-2)
self.qtau_w1 = bn.random.Gamma(bn.softplus(self.shape_w1), bn.softplus(self.rate_w1), p=ptau_w1)
pw1 = bn.random.Gaussian(0, tau=self.qtau_w1.draw())
self.qw1 = bn.random.Gaussian(self.w1_mu, bn.softplus(self.w1_s), p=pw1)
ptau_b1 = bn.random.Gamma(1., 1e-2)
self.qtau_b1 = bn.random.Gamma(bn.softplus(self.shape_b1), bn.softplus(self.rate_b1), p=ptau_b1)
pb1 = bn.random.Gaussian(0, tau=self.qtau_b1.draw())
self.qb1 = bn.random.Gaussian(self.b1_mu, bn.softplus(self.b1_s), p=pb1)
ptau_w2 = bn.random.Gamma(1., 1e-2)
self.qtau_w2 = bn.random.Gamma(bn.softplus(self.shape_w2), bn.softplus(self.rate_w2), p=ptau_w2)
pw2 = bn.random.Gaussian(0, tau=self.qtau_w2.draw())
self.qw2 = bn.random.Gaussian(self.w2_mu, bn.softplus(self.w2_s), p=pw2)
ptau_b2 = bn.random.Gamma(1., 1e-2)
self.qtau_b2 = bn.random.Gamma(bn.softplus(self.shape_b2), bn.softplus(self.rate_b2), p=ptau_b2)
pb2 = bn.random.Gaussian(0, tau=self.qtau_b2.draw())
self.qb2 = bn.random.Gaussian(self.b2_mu, bn.softplus(self.b2_s), p=pb2)
h = bn.tanh(x @ self.qw1.draw() + self.qb1.draw())
mu = h @ self.qw2.draw() + self.qb2.draw()
self.py = bn.random.Gaussian(mu, 0.1, data=y)
if y is None:
return self.py.draw().value
# +
model = ARDNetwork(1, 20, 1)
optimizer = bn.optimizer.Adam(model, 0.1)
optimizer.set_decay(0.9, 100)
for _ in range(10000):
model.clear()
model(x_train, y_train)
elbo = model.elbo()
elbo.backward()
optimizer.update()
# -
x = np.linspace(-3, 3, 1000)[:, None]
plt.scatter(x_train, y_train)
y = [model(x) for _ in range(100)]
y_mean = np.mean(y, axis=0)
y_std = np.std(y, axis=0)
plt.plot(x, y_mean, c="orange")
plt.fill_between(x.ravel(), (y_mean - y_std).ravel(), (y_mean + y_std).ravel(), color="orange", alpha=0.2)
plt.show()
parameter = []
parameter.extend(list(model.w1_mu.value.ravel()))
parameter.extend(list(model.b1_mu.value.ravel()))
parameter.extend(list(model.w2_mu.value.ravel()))
parameter.extend(list(model.b2_mu.value.ravel()))
plt.plot(parameter)
| notebook/AutomaticRelevanceDetermination.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id='top'></a>
#
# # *pythonic* package development tutorial
#
# There are great resources out there for getting your projects documented and distributed. I got started using [shablona](https://github.com/uwescience/shablona/tree/master/doc) from the eScience institute at U Washington. I would still recommend that to folks. I've found, however, that it can be helpful to start bare-bones and walk through a tutorial to build your package up, to really understand how everything is working together. So in the following, if you follow the tutorial, you'll start with a basic directory structure, and proced to add on documentation, web hosting, continuous integration, coverage, and finally deploy your package on pypi.
#
# ## Overview
#
# By the grace of open-source-dev there are several free lunches you should know of:
#
# 2. [sphinx](#sphinx)
# 1. sphinx can be a bit [finicky](https://samnicholls.net/2016/06/15/how-to-sphinx-readthedocs/). The most important feature to introduce to you to today will be
# 2. [autodocs](https://samnicholls.net/2016/06/15/how-to-sphinx-readthedocs/) where we generate documentation from just your
# 3. [docstrings](http://sphinxcontrib-napoleon.readthedocs.io/en/latest/example_google.html) super cool!
# 1. [github pages](#ghpages)
# 1. Eventually I'll add a segment on getting your documentation on read the docs. But while code is still in development I've found rtd to be overkill. GH Pages is a simple alternative that hosts your static html files and doesn't require building your site on a remote server.
# 3. [travis CI](#travis)
# 1. "CI" stands for continuous integration. These folks provide you with a free service -- up to 1 hour of CPU time on their servers to run all of your unit tests.
# 4. [coveralls](#coveralls)
# 1. how much of that passed build is covered?!
# 5. [pypi](#pypi)
# 1. You want people using your code as fast as possible, right?
# <a id='sphinx'></a>
#
# ## Sphinx
#
# [back to top](#top)
#
# ### Basic directory structure
#
# Clone this repo and cd into the main directory. Checkout the package organization:
#
# ```
# $ tree
# .
# ├── ecsdemo
# │ ├── __init__.py
# │ ├── core.py
# │ ├── data
# │ │ ├── climate_sentiment_m1.h5
# │ │ └── tweet_global_warming.csv
# │ ├── input.py
# │ ├── tests
# │ │ ├── __init__.py
# │ │ └── test_ecsdemo.py
# │ └── version.py
# ├── LICENSE
# ├── README.md
# ├── docs
# │ ├── Makefile
# │ ├── _static
# │ ├── conf.py
# │ ├── index.rst
# │ └── source
# │ ├── ecsdemo.core.rst
# │ ├── ecsdemo.rst
# │ └── ecsdemo.tests.rst
# ├── examples
# │ ├── README.ipynb
# └── setup.py
# ```
#
# We're about to find out just how busy this directory structure can be with these added open source features. But for now, the main project lives under `ecsdemo/` with `tests/` and `data/` subdirectories.
#
# Go ahead and inspect the contents of the core.py and test_ecsdemo.py files, in case you're interested. There's some common elements here in the package development world. `core.py` contains, well, the core code of the package. In a larger package you might have other modules living here such as `analysis.py` or `visualize.py`, depending on how you want to organize your code. For now, the `core.py` file contains four functions: `load_data, data_setup, baseline_model` and one class: `Benchmark`. You can learn more about pythonic naming conventions from the [pep8](https://www.python.org/dev/peps/pep-0008/) documentation.
#
# ### Makefile
#
# Time to get to Sphinx! cd over to the docs directory. In this tutorial, I've setup the appropriate rst files already. I haven't had excellent luck with using sphinx-quickstart or sphinx-autogen, personally. And so I will always start with a template such as this and modify the `.rst` files as needed. Suffice to say, if you are interested in creating your documentation from scratch I found this [source](https://samnicholls.net/2016/06/15/how-to-sphinx-readthedocs/) helpful.
#
# All you need to do is type `make html` in the `docs/` directory where your `Makefile` is sitting. and Sphinx will generate static html documents of your site.
#
# ```
# $ tree -L 2
# .
# ├── Makefile
# ├── _build
# │ ├── doctrees
# │ └── html
# ├── _static
# ├── conf.py
# ├── index.rst
# └── source
# ├── ecsdemo.core.rst
# ├── ecsdemo.rst
# └── ecsdemo.tests.rst
# ```
#
# Use your preferred browser to checkout your site: `open _build/html/index.html`. If you navigate to the API you'll see how Sphinx autmoatically formats your docstrings for you, super neat!
# <a id='ghpages'></a>
#
# ## Github Pages
#
# [back to top](#top)
#
# At this point we're ready to make our documentation live. Personally, my favorite method while I'm still developing a project is Github pages. This is because github allows us to directly host our statically generated files with Sphinx. As your code grows up, you'll want to migrate to something more robust like readthedocs.io. The downside of doing this initially is that readthedocs compiles your website as you push to github and little changes in your code structure or prerequisites can break the build. It's just easier to put this work off until you're at version 0.0.1.
#
# ### Github UI
#
# In your browser, navigate to the settings folder for your cloned github project and scroll down until you see Github Pages. Change the Source option to master branch /docs folder and hit save.
#
# ### Local changes
#
# Now in your local repo we're going to do a bit of a juggling act. We've been working in docs/ directory we'll want to move this sphinx stuff to its own home and make sure our statically generated files live here. So in your main directory this would like:
# ```
# $ mkdir sphinx
# $ mv docs/* sphinx/
# $ mv sphinx/_build/html/* docs/
# $ tree -L 1 docs/
# ├── docs
# │ ├── _sources
# │ ├── _static
# │ ├── genindex.html
# │ ├── index.html
# │ ├── objects.inv
# │ ├── py-modindex.html
# │ ├── search.html
# │ ├── searchindex.js
# │ └── source
# ```
# By default, github uses jekyll to build its' sites. We'll want to turn this feature off since our site is already built. In the `docs/` directory simply type `touch .nojekyll`. Now add/commit/push your changes. Your project is live under the url (yourgihtubname).io/ECS
# <a id='travis'></a>
#
# ## Travis CI
#
# [back to top](#top)
#
# ### Makefile
#
# We're going to use travis to run our unit tests. Good coding practice dictates that we also check our code for readability. This is done using pep8.
#
# Before we do this with travis we'll want to test our code locally. Same as with the autodocumentation, a Makefile makes this job easier for this. You'll create a new Makefile in the main directory and add the following:
# ```
# flake8:
# @if command -v flake8 > /dev/null; then \
# echo "Running flake8"; \
# flake8 flake8 --ignore N802,N806,F401 `find . -name \*.py | grep -v setup.py | grep -v /docs/ | grep -v /sphinx/`; \
# else \
# echo "flake8 not found, please install it!"; \
# exit 1; \
# fi;
# @echo "flake8 passed"
# ```
# Basically, we're asking flake8 to run some but not all, tests on some but not all, files.
#
# at the end of that file we'll also add:
# ```
# test:
# py.test
# ```
# You can now run `make flake8` and `make test` and see that your package passes your unittests, locally.
#
# ### requirements.txt
#
# Travis CI needs to know how to build the environment to run our code. We'll do this with a requirements.txt file, also in the main directory:
# ```
# keras
# tensorflow
# scikit-learn
# pandas
# ```
#
# ### .travis.yml
#
# If you haven't already, navigate over to travis-ci.org and create an account with them, then import the ECS project.
#
# To get this up and running with travis we'll need to add a .travis.yml file:
#
# ```
# language: python
# sudo: false
#
# deploy:
# provider: pypi
# user: wesleybeckner
# password:
# secure:
# on:
# tags: true
# repo: ECS
#
# env:
# global:
# - PIP_DEPS="pytest coveralls pytest-cov flake8"
#
# python:
# - '3.6'
#
#
# install:
# - travis_retry pip install $PIP_DEPS
# - travis_retry pip install numpy cython
# - travis_retry pip install -r requirements.txt
# - travis_retry pip install -e .
#
# before_script: # configure a headless display to test plot generation
# - "export DISPLAY=:99.0"
# - "sh -e /etc/init.d/xvfb start"
# - sleep 3 # give xvfb some time to start
#
# script:
# - flake8 --ignore N802,N806,W503,F401 `find . -name \*.py | grep -v setup.py | grep -v version.py | grep -v __init__.py | grep -v /docs/ | grep -v /sphinx/`
# - mkdir for_test
# - cd for_test
# - py.test --pyargs ecsdemo --cov-report term-missing --cov=ecsdemo
# ```
#
# you'll notice that the `.travis.yml` file contains the same flake8 and py.test commands. git add/commit/push and checkout your passing travis build.
# <a id='coveralls'></a>
#
# ## Coveralls
#
# [back to top](#top)
#
# Other than setting up an account with coveralls and linking it to your github account, you don't have much to do here. At the end of your .travis.yml file add the following:
# ```
#
# after_success:
# - coveralls
# ```
# travis ci will now pass your build to coveralls.
# <a id='pypi'></a>
#
# ## pypi
#
# [back to top](#top)
#
# `setup.py` contains the information that will launch our project on pypi. cd to your main directory and issue the command: `python setup.py sdist upload` (you'll need to have registered an email address + account with them before hand)
#
# At the end of this tutorial, your directory structure will have grown substantially!
# ```
# $ tree -L 3
# .
# ├── ecsdemo
# │ ├── __init__.py
# │ ├── __pycache__
# │ │ ├── __init__.cpython-36.pyc
# │ │ ├── core.cpython-36.pyc
# │ │ ├── input.cpython-36.pyc
# │ │ └── version.cpython-36.pyc
# │ ├── core.py
# │ ├── data
# │ │ ├── climate_sentiment_m1.h5
# │ │ └── tweet_global_warming.csv
# │ ├── tests
# │ │ ├── __init__.py
# │ │ ├── __pycache__
# │ │ └── test_ecsdemo.py
# │ └── version.py
# ├── ecsdemo.egg-info
# │ ├── PKG-INFO
# │ ├── SOURCES.txt
# │ ├── dependency_links.txt
# │ ├── requires.txt
# │ └── top_level.txt
# ├── LICENSE
# ├── Makefile
# ├── README.md
# ├── dist
# │ └── ecsdemo-0.0.dev0.tar.gz
# ├── docs
# │ ├── _sources
# │ │ ├── index.rst.txt
# │ │ └── source
# │ ├── _static
# │ │ ├── ajax-loader.gif
# │ │ ├── basic.css
# │ │ ├── comment-bright.png
# │ │ ├── comment-close.png
# │ │ ├── comment.png
# │ │ ├── css
# │ │ ├── doctools.js
# │ │ ├── documentation_options.js
# │ │ ├── down-pressed.png
# │ │ ├── down.png
# │ │ ├── file.png
# │ │ ├── fonts
# │ │ ├── jquery-3.2.1.js
# │ │ ├── jquery.js
# │ │ ├── js
# │ │ ├── minus.png
# │ │ ├── plus.png
# │ │ ├── pygments.css
# │ │ ├── searchtools.js
# │ │ ├── underscore-1.3.1.js
# │ │ ├── underscore.js
# │ │ ├── up-pressed.png
# │ │ ├── up.png
# │ │ └── websupport.js
# │ ├── genindex.html
# │ ├── index.html
# │ ├── objects.inv
# │ ├── py-modindex.html
# │ ├── search.html
# │ ├── searchindex.js
# │ └── source
# │ ├── ecsdemo.core.html
# │ ├── ecsdemo.html
# │ └── ecsdemo.tests.html
# ├── examples
# │ ├── README.ipynb
# │ ├── README.txt
# │ ├── demo1.png
# │ └── demo2.png
# ├── requirements.txt
# ├── setup.py
# └── sphinx
# ├── Makefile
# ├── _build
# │ ├── doctrees
# │ └── html
# ├── _static
# ├── conf.py
# ├── index.rst
# └── source
# ├── ecsdemo.core.rst
# ├── ecsdemo.rst
# └── ecsdemo.tests.rst
# ```
# ## More to come
#
# 1. sdist vs bdist
# 2. Google cloud platform
# 3. Twitter API
| examples/README.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
df = pd.read_csv('/Users/jo-elle/Desktop/Data301/project/course-project-solo_305/data/raw.data/UCI_Credit_Card.csv')
df
# ## In this notebook, I spent some time getting to know my data and then drafted my method chaining in here before creating it into a function, it is a bit of a mess.... But it was really helpful in understanding my data and what needs to be changed and updated!
#I am going to attempt to find the mean age, mean bill payment, mean bill amount, and the range of different education level
#I am also going to look for correlations, and use groupby to find more specific analysis
df.mean()
df.info()
# I am going to now describe all my variables and split them up according to whether they contain catergorical variables or actualy values
#Categorical variables description, this is showing me the percent of number in these three categories that use number to classify a characterisitc
df[['SEX', 'EDUCATION', 'MARRIAGE']].describe()
# Payment delay description
df[['PAY_0', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6']].describe()
# Bill Statement description
df[['BILL_AMT1', 'BILL_AMT2', 'BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6']].describe()
df.LIMIT_BAL.describe()
# Bill Statement description
df[['BILL_AMT1', 'BILL_AMT2', 'BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6']].describe()
# Now that I know my data better I am going to clean it
# The 3 in marriage can be categorized as other, other can mean divroced, widowed etc...
df.loc[df.MARRIAGE == 3, 'MARRIAGE'] = "other"
df.MARRIAGE.value_counts()
# +
df.loc[df.EDUCATION == 5, 'EDUCATION'] = 4
df.loc[df.EDUCATION == 6, 'EDUCATION'] = 4
df.loc[df.EDUCATION == 0, 'EDUCATION'] = 4
df.EDUCATION.value_counts()
# -
# Payment delay description
df[['PAY_1', 'PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6']].describe()
# +
#Looking at my payment data, there's -2, and what is 0 if -1 is pay duly?...
#which isnt defined in my variable list in my READ.MD, so I need to combine -2,-1 and 0 into one
#should look into making a function for this, this seems counterintuitive
a = (df1.PAY_0 == -2) | (df1.PAY_0 == -1) | (df1.PAY_0 == 0)
df.loc[a, 'PAY_1'] = 0
a = (df1.PAY_2 == -2) | (df1.PAY_2 == -1) | (df1.PAY_2 == 0)
df.loc[a, 'PAY_2'] = 0
a = (df1.PAY_3 == -2) | (df1.PAY_3 == -1) | (df1.PAY_3 == 0)
df.loc[a, 'PAY_3'] = 0
a = (df1.PAY_4 == -2) | (df1.PAY_4 == -1) | (df1.PAY_4 == 0)
df.loc[a, 'PAY_4'] = 0
a = (df1.PAY_5 == -2) | (df1.PAY_5 == -1) | (df1.PAY_5 == 0)
df.loc[a, 'PAY_5'] = 0
a = (df1.PAY_6 == -2) | (df1.PAY_6 == -1) | (df1.PAY_6 == 0)
df.loc[a, 'PAY_6'] = 0
late = df1[['PAY_0','PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6']]
df1
# +
#First attempt at method chaining
#Also just renaming columns that are long to type out and that don't make much sense to me personally
#Dropping lots of columns as I am interested in seeing if default rate is at correlated to characteristics or the amount of balance limit
df9 = (df
.drop(columns={'BILL_AMT1', 'BILL_AMT2', 'BILL_AMT3', 'BILL_AMT4', 'BILL_AMT5', 'BILL_AMT6'})
.drop(columns={"PAY_AMT1",'PAY_AMT2','PAY_AMT3','PAY_AMT4','PAY_AMT5','PAY_AMT6'})
.rename(columns={"PAY_0": "PAY_1"})
.rename(columns={"default.payment.next.month":'Default'}))
df9
# -
df1 = df.rename(columns={"PAY_0": "PAY_1"})
df1
a = (df1.PAY_1 == -2) | (df1.PAY_1 == -1) | (df1.PAY_1 == 0)
df.loc[a, 'PAY_1'] = 0
a = (df1.PAY_2 == -2) | (df1.PAY_2 == -1) | (df1.PAY_2 == 0)
df.loc[a, 'PAY_2'] = 0
a = (df1.PAY_3 == -2) | (df1.PAY_3 == -1) | (df1.PAY_3 == 0)
df.loc[a, 'PAY_3'] = 0
a = (df1.PAY_4 == -2) | (df1.PAY_4 == -1) | (df1.PAY_4 == 0)
df.loc[a, 'PAY_4'] = 0
a = (df1.PAY_5 == -2) | (df1.PAY_5 == -1) | (df1.PAY_5 == 0)
df.loc[a, 'PAY_5'] = 0
a = (df1.PAY_6 == -2) | (df1.PAY_6 == -1) | (df1.PAY_6 == 0)
df.loc[a, 'PAY_6'] = 0
late = df1[['PAY_1','PAY_2', 'PAY_3', 'PAY_4', 'PAY_5', 'PAY_6']]
df1
#Now I'm going to start my initial analysis, first I'm looking at any correlation between sex and payment
df1.groupby(['SEX', 'default.payment.next.month']).size()
dfgender = df1.groupby(['SEX', 'default.payment.next.month']).size().unstack(1)
dfgender
dfgender.plot(kind='bar', stacked = True)
dfgender['perc'] = (dfgender[1]/(dfgender[0] + dfgender[1]))
dfgender
def correlation(Col1, Col2):
res = df.groupby([Col1, Col2]).size().unstack()
res['perc'] = (res[res.columns[1]]/(res[res.columns[0]] + res[res.columns[1]]))
return res
#Thus is just taking what I did above and putting it into a function so I can repeat with my other variables that will help anser my research question
correlation("EDUCATION", "default.payment.next.month")
correlation("MARRIAGE", "default.payment.next.month")
# +
subset = df1[['SEX', 'EDUCATION', 'MARRIAGE', 'PAY_1', 'PAY_2', 'PAY_3', 'PAY_4',
'PAY_5', 'PAY_6', 'default.payment.next.month']]
f, axes = plt.subplots(3, 3, figsize=(20, 15), facecolor='white')
f.suptitle('FREQUENCY OF CATEGORICAL VARIABLES (BY TARGET)')
ax1 = sns.countplot(x="SEX", hue="default.payment.next.month", data=subset, palette="Blues", ax=axes[0,0])
ax2 = sns.countplot(x="EDUCATION", hue="default.payment.next.month", data=subset, palette="Blues",ax=axes[0,1])
ax3 = sns.countplot(x="MARRIAGE", hue="default.payment.next.month", data=subset, palette="Blues",ax=axes[0,2])
ax4 = sns.countplot(x="PAY_1", hue="default.payment.next.month", data=subset, palette="Blues", ax=axes[1,0])
ax5 = sns.countplot(x="PAY_2", hue="default.payment.next.month", data=subset, palette="Blues", ax=axes[1,1])
ax6 = sns.countplot(x="PAY_3", hue="default.payment.next.month", data=subset, palette="Blues", ax=axes[1,2])
ax7 = sns.countplot(x="PAY_4", hue="default.payment.next.month", data=subset, palette="Blues", ax=axes[2,0])
ax8 = sns.countplot(x="PAY_5", hue="default.payment.next.month", data=subset, palette="Blues", ax=axes[2,1])
ax9 = sns.countplot(x="PAY_6", hue="default.payment.next.month", data=subset, palette="Blues", ax=axes[2,2]);
# -
| analysis/joelle.analysis/.ipynb_checkpoints/datawrangle_for_Milestone2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] tags=[]
# # Quick Review of scikit-learn
#
# <a href="https://colab.research.google.com/github/thomasjpfan/ml-workshop-intermediate-1-of-2/blob/master/notebooks/00-review-sklearn.ipynb"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
# -
# Install dependencies for google colab
import sys
if 'google.colab' in sys.modules:
# %pip install -r https://raw.githubusercontent.com/thomasjpfan/ml-workshop-intermediate-1-of-2/master/requirements.txt
import sklearn
assert sklearn.__version__.startswith("1.0"), "Plese install scikit-learn 1.0"
import seaborn as sns
sns.set_theme(context="notebook", font_scale=1.2,
rc={"figure.figsize": [10, 6]})
sklearn.set_config(display="diagram")
# +
from sklearn.datasets import fetch_openml
steel = fetch_openml(data_id=1504, as_frame=True)
# -
print(steel.DESCR)
_ = steel.data.hist(figsize=(30, 15), layout=(5, 8))
# ### Split Data
# +
from sklearn.model_selection import train_test_split
X, y = steel.data, steel.target
X_train, X_test, y_train, y_test = train_test_split(
X, y, random_state=42, stratify=y)
# -
# ### Train DummyClassifer
# +
from sklearn.dummy import DummyClassifier
dc = DummyClassifier(strategy='prior').fit(X_train, y_train)
dc.score(X_test, y_test)
# -
# ### Train KNN based model
# +
from sklearn.preprocessing import StandardScaler
from sklearn.pipeline import make_pipeline
from sklearn.neighbors import KNeighborsClassifier
knc = make_pipeline(
StandardScaler(),
KNeighborsClassifier()
)
knc.fit(X_train, y_train)
# -
knc.score(X_test, y_test)
# ## Exercise 1
#
# 1. Load the wisconsin breast cancer dataset from `sklearn.datasets.load_breast_cancer`.
# 2. Is the labels imbalanced? (**Hint**: `value_counts`)
# 3. Split the data into a training and test set.
# 4. Create a pipeline with a `StandardScaler` and `LogisticRegression` and fit on the training set.
# 5. Evalute the pipeline on the test set.
# 6. **Extra**: Use `sklearn.metrics.f1_score` to compute the f1 score on the test set.
from sklearn.datasets import load_breast_cancer
from sklearn.linear_model import LogisticRegression
from sklearn.metrics import f1_score
# **If you are running locally**, you can uncomment the following cell to load the solution into the cell. On **Google Colab**, [see solution here](https://github.com/thomasjpfan/ml-workshop-intermediate-1-of-2/blob/master/notebooks/solutions/00-ex01-solutions.py).
# +
# # %load solutions/00-ex01-solutions.py
| notebooks/00-review-sklearn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
x = np.arange(-5.0, 5.0, 0.1)
# Ingat persamaan Y = (a)X + (b)
y = 2*(x) + 3
y_noise = 2 * np.random.normal(size=x.size)
ydata = y + y_noise
#plt.figure(figsize=(8,6))
plt.plot(x, ydata, 'bo')
plt.plot(x,y, 'r')
plt.ylabel('Dependent Variable')
plt.xlabel('Independent Variable')
plt.show()
# +
x = np.arange(-5.0, 5.0, 0.1)
# Ingat untuk Regresi Linear Y=(a)X+(b), namun untuk persamaan non Linear berbeda!
y = 1*(x**3) + 1*(x**2) + 1*x + 3
y_noise = 20 * np.random.normal(size=x.size)
ydata = y + y_noise
plt.plot(x, ydata, 'bo')
plt.plot(x,y, 'r')
plt.ylabel('Dependent Variable')
plt.xlabel('Independent Variable')
plt.show()
# -
# Pada hasil gambar di atas dapat dilihat fungsi $x^3$ dan $x^2$ adalah independent variables. Inilah fungsi non-linear.
# ### Contoh fungsi Quadratic
# Persamaannya: Y = $X^2$
# +
x = np.arange(-5.0, 5.0, 0.1)
y = np.power(x,2)
y_noise = 2 * np.random.normal(size=x.size)
ydata = y + y_noise
plt.plot(x, ydata, 'bo')
plt.plot(x,y, 'r')
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# ### Contoh fungsi Exponential
# Fungsinya: Y = a+$bc^x$ dimana b ≠0, c > 0 , c ≠1, dan x adalah bilangan real. Basis, c, adalah nilai konstanta dan eksponensial, x, adalah sebuah variabel.
# +
X = np.arange(-5.0, 5.0, 0.1)
Y= np.exp(X)
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Independent Variable')
plt.show()
# -
# ### Contoh fungsi Logarithmic
# Persamaannya: y= log(x)
# +
X = np.arange(-5.0, 5.0, 0.1)
Y = np.log(X)
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Indepdendent Variable')
plt.show()
# -
# ### Sigmoidal/Logistic
# Persamaannya:
# Y = a + b/1+$c^X-D$
# +
X = np.arange(-5.0, 5.0, 0.1)
Y = 1-4/(1+np.power(3, X-2))
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Independent Variable')
plt.show()
# -
# ## Contoh Regresi Non-Linear
# +
import numpy as np
import pandas as pd
# ubah ke data-frame
df = pd.read_csv("china_gdp.csv")
df.tail()
# -
# ## Plotting Dataset
plt.figure(figsize=(8,5))
x_data, y_data = (df["Year"].values, df["Value"].values)
plt.plot(x_data, y_data, 'ro')
plt.ylabel('GDP')
plt.xlabel('Year')
plt.show()
# Terlihat Pertumbuhannya sangat lambat,lalu mulai 2005 bertumbuh sangat signifikan, dan kemudian naik drastis di tahun 2010.
# #### Pilih model
# Dari grafik awal dapat kita lihat bahwa fungsi logistik cocok untuk kasus ini.
# +
X = np.arange(-5.0, 5.0, 0.1)
Y = 1.0 / (1.0 + np.exp(-X))
plt.plot(X,Y)
plt.ylabel('Dependent Variable')
plt.xlabel('Independent Variable')
plt.show()
# -
# #### Membangun Model
# buat model regresi untuk kasus tersebut
def sigmoid(x, Beta_1, Beta_2):
y = 1 / (1 + np.exp(-Beta_1*(x-Beta_2)))
return y
# Coba dengan contoh fungsi sigmoid terhadap data
#
# +
beta_1 = 0.10
beta_2 = 1990.0
# fungsi logistik (sigmoid)
Y_pred = sigmoid(x_data, beta_1 , beta_2)
# Inisialisasi terhadap datapoints
plt.plot(x_data, Y_pred*15000000000000.)
plt.plot(x_data, y_data, 'ro')
# -
# Kita harus dapatkan parameter terbaik untuk model kita, oleh karenanya yang pertama harus dilakukan adalah normalisasi x dan y
# Normalisasi data
xdata =x_data/max(x_data)
ydata =y_data/max(y_data)
# Bagaimana menemukan parameter terbaik untuk fit line?
from scipy.optimize import curve_fit
popt, pcov = curve_fit(sigmoid, xdata, ydata)
# Cetak parameter terakhir
print(" beta_1 = %f, beta_2 = %f" % (popt[0], popt[1]))
# Plot-kan ke hasil model regresi
x = np.linspace(1960, 2015, 55)
x = x/max(x)
plt.figure(figsize=(8,5))
y = sigmoid(x, *popt)
plt.plot(xdata, ydata, 'ro', label='data')
plt.plot(x,y, linewidth=3.0, label='fit')
plt.legend(loc='best')
plt.ylabel('GDP')
plt.xlabel('Year')
plt.show()
# ### Hitung akurasi dari model yang dibuat
# +
# split data into train/test
msk = np.random.rand(len(df)) < 0.8
train_x = xdata[msk]
test_x = xdata[~msk]
train_y = ydata[msk]
test_y = ydata[~msk]
# build the model using train set
popt, pcov = curve_fit(sigmoid, train_x, train_y)
# predict using test set
y_hat = sigmoid(test_x, *popt)
# evaluation
print("Mean absolute error: %.2f" % np.mean(np.absolute(y_hat - test_y)))
print("Residual sum of squares (MSE): %.2f" % np.mean((y_hat - test_y) ** 2))
from sklearn.metrics import r2_score
print("R2-score: %.2f" % r2_score(y_hat , test_y) )
# -
| Pertemuan 12/Regresi non-linear.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pd
import spotipy
from spotipy.oauth2 import SpotifyClientCredentials
from env import cid, c_secret
from prepare import change_dtypes, handle_nulls, set_index
from preprocessing import create_features
# -
pd.set_option("display.max_colwidth", 10000)
client_credentials_manager = SpotifyClientCredentials(client_id=cid,
client_secret=c_secret)
sp = spotipy.Spotify(client_credentials_manager=client_credentials_manager)
sp.trace=False
def create_spotipy_client():
client_credentials_manager = SpotifyClientCredentials(client_id=cid,
client_secret=c_secret)
sp = spotipy.Spotify(client_credentials_manager=client_credentials_manager)
sp.trace=False
return sp
lz_uri = 'spotify:artist:36QJpDe2go2KgaRleHCDTp'
results = sp.artist_top_tracks(lz_uri)
sample_data = pd.DataFrame(results['tracks']).head(10)
sample_data
sample_data.to_csv('sample_data.csv', index=False)
genres = pd.DataFrame(sp.recommendation_genre_seeds())
genres
genres.to_csv('genres.csv', index=False)
recommendations = sp.recommendations(seed_genres=['folk' ,'punk'])
pd.DataFrame(recommendations['tracks'])
playlist_features_list = ["artist","album","track_name","track_id","danceability","energy","key",
"loudness","mode", "speechiness","instrumentalness","liveness","valence","tempo",
"duration_ms","time_signature"]
playlist_df = pd.DataFrame(columns = playlist_features_list)
playlist = sp.user_playlist_tracks("spotify", "33I6RpefRQcRh69xEczaKT")
playlist
def analyze_playlist(creator, playlist_id, sp_client, offset=0):
# Create empty dataframe
playlist_features_list = ["artist","album","release_date","track_name","track_id",'label',
"danceability","energy","key","loudness","mode", "speechiness","instrumentalness",
"liveness","valence","tempo", "duration_ms","time_signature"]
playlist_df = pd.DataFrame(columns = playlist_features_list)
# Loop through every track in the playlist, extract features and append the features to the playlist df
playlist = sp_client.user_playlist_tracks(creator, playlist_id, offset=offset)['items']
for track in playlist:
# Create empty dict
playlist_features = {}
# Get metadata
if track['track']['album']['artists'] == []:
continue
else:
playlist_features['artist'] = track['track']['album']['artists'][0]['name']
playlist_features["album"] = track["track"]["album"]["name"]
playlist_features["release_date"] = track["track"]["album"]["release_date"]
playlist_features["track_name"] = track["track"]["name"]
playlist_features["track_id"] = track["track"]["id"]
playlist_features['explicit'] = track['track']['explicit']
playlist_features["popularity"] = track["track"]["popularity"]
playlist_features['disc_number'] = track['track']['disc_number']
playlist_features['track_number'] = track['track']['track_number']
playlist_features['album_id'] = track['track']['album']['id']
playlist_features['album_type'] = track['track']['album']['album_type']
# Get audio features
audio_features = sp_client.audio_features(playlist_features["track_id"])
if audio_features is None:
for feature in playlist_features_list[6:]:
playlist_features[feature] = None
elif audio_features[0] is None:
for feature in playlist_features_list[6:]:
playlist_features[feature] = None
else:
for feature in playlist_features_list[6:]:
playlist_features[feature] = audio_features[0][feature]
# Get album popularity
album_features = sp_client.album(playlist_features['album_id'])
if album_features is None:
for feature in playlist_features_list[5:6]:
playlist_features[feature] = None
else:
playlist_features['label'] = album_features['label']
# Concat the dfs
track_df = pd.DataFrame(playlist_features, index = [0])
playlist_df = pd.concat([playlist_df, track_df], ignore_index = True)
return playlist_df
for offset in range(0, 6000, 100):
print(f'Making page with offset = {offset}')
playlist_df = analyze_playlist('spotify:user:afrodeezeemusic', '3P6Pr6iEqvK5fl4UkgdQ7T', sp, offset)
playlist_df.to_csv('data/playlist-offset-' + str(offset) + '.csv')
testing = analyze_playlist('spotify:user:afrodeezeemusic', '3P6Pr6iEqvK5fl4UkgdQ7T', sp)
testing
album_features = sp.album('2J1hMj78HfdcMrmL2Sk6eR')
album_features
album_features['']
data.to_csv('capstone_playlist.csv', index=False)
data.key.value_counts()
playlist = sp.user_playlist_tracks(creator, playlist_id, offset=offset)['items']
new_data
playlist = sp.user_playlist_tracks('spotify:user:afrodeezeemusic', '3P6Pr6iEqvK5fl4UkgdQ7T', limit=100)
playlist.keys()
playlist['items'][0]['track']['album']['id']
headers = {'Authorization': 'Bearer BQCbWO3XkU1v_ajHFrS2qVIasL<KEY>', 'Content-Type': 'application/json'}
url = 'https://spotify.com/v1/playlists/3P6Pr6iEqvK5fl4UkgdQ7T/tracks'
import requests
response = requests.get(url, headers=headers)
response.status_code
import logging
# make a basic logging configuration
# here we set the level of logging to DEBUG
logging.basicConfig(
level=logging.DEBUG
)
# make a debug message
logging.debug("This is a simple debug log")
df = pd.read_csv('full-playlist.csv', index_col=0)
df.head()
create_features(df)
df.head()
def prepare_df(df):
df = create_features(df)
df = handle_nulls(df)
df = change_dtypes(df)
df = set_index(df)
return df
prepare_df(df)
df = prepare_df(df)
df.head()
def concat_csv_files():
'''
Loops through each csv file of acquired data to combine into one df.
No parameters needed, only needs the files saved in the working directory.
Returns the one df.
'''
# sets initial df as file of first 700 observations
df = pd.read_csv('data/playlist-offset-0.csv', index_col=0)
# loops through 700 - 6000 by one hundreds, matching the csv file names
# as it loops, it combines the csv file to the original df
for offset in range(100, 5901, 100):
# saves next csv file as a df
add_df = pd.read_csv(f'data/playlist-offset-{offset}.csv', index_col=0)
# adds the new df to the original df
df = pd.concat([df, add_df], ignore_index=True)
# returns the csv files combined in one dataframe, should be 6_000 observations
return df
full_playlist = concat_csv_files()
full_playlist
full_playlist.to_csv('full-playlist.csv')
def get_capstone_playlist():
# Let this loop run as it gathers the tracks from the playlist
for offset in range(0, 6000, 100):
# Prints out how many pages in the loop is. Each page is 100 tracks + or - a few if nulls appear
print(f'Making page with offset = {offset}')
# Analyze the first 100 tracks past the offset
playlist_df = analyze_playlist('spotify:user:afrodeezeemusic', '3P6Pr6iEqvK5fl4UkgdQ7T', sp, offset)
# Write each dataframe of 100 tracks to a csv. If the function ends early in an error you will still have some data
playlist_df.to_csv('data/playlist-offset-' + str(offset) + '.csv')
# use the concat_csv_files function to concat all the dataframes together into one complete dataframe
df = concat_csv_files()
return df
| sandbox/mays/mays_acquire.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/BrianBehnke/competition/blob/master/BigDataBowl.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="mHsjwRAflcSj" colab_type="code" colab={}
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# + id="vp7Pia2gljB8" colab_type="code" colab={}
# Setting urls to the data from https://github.com/nfl-football-ops/Big-Data-Bowl/tree/master/Data
games_url = "https://raw.githubusercontent.com/nfl-football-ops/Big-Data-Bowl/master/Data/games.csv"
players_url = "https://raw.githubusercontent.com/nfl-football-ops/Big-Data-Bowl/master/Data/players.csv"
plays_url = "https://raw.githubusercontent.com/nfl-football-ops/Big-Data-Bowl/master/Data/plays.csv"
gamedata_url = "https://raw.githubusercontent.com/nfl-football-ops/Big-Data-Bowl/master/Data/tracking_gameId_2017091711.csv"
# + id="Q3f_e4u1llFb" colab_type="code" outputId="b117ff31-70ec-48f1-e3a7-0f9deef3d85d" colab={"base_uri": "https://localhost:8080/", "height": 389}
# Placing data into dataframe for games
df_games = pd.read_csv(games_url)
print(df_games.shape)
df_games.head()
# + id="0lcPC4jVll77" colab_type="code" outputId="0cdc75e5-a305-4e90-b7a4-6d4f4c9ea7de" colab={"base_uri": "https://localhost:8080/", "height": 221}
# Placing data into dataframe for players
df_players = pd.read_csv(players_url)
print(df_players.shape)
df_players.head()
# + id="ZdvbToE5lm1y" colab_type="code" outputId="ffb10807-eb1f-4b9b-c019-6d76be6e22b4" colab={"base_uri": "https://localhost:8080/", "height": 920}
# Placing data into dataframe for plays
df_plays = pd.read_csv(plays_url)
print(df_plays.shape)
df_plays.head().T
# + id="DINUCyeC9ubT" colab_type="code" outputId="f0cbc393-ae06-4f7d-d0cc-10aa06bfb837" colab={"base_uri": "https://localhost:8080/", "height": 686}
# Placing data into dataframe for plays
gamedata = pd.read_csv(gamedata_url)
print(gamedata.shape)
gamedata.head(20)
# + id="dKMx-2g2Bz3k" colab_type="code" outputId="21d13f1d-d287-4b59-cbce-a33812be1bf6" colab={"base_uri": "https://localhost:8080/", "height": 1969}
play36 = gamedata[gamedata['playId'] == 36]
play36
# + id="Z_3nyuHAB770" colab_type="code" colab={}
niles = play36[play36.displayName == '<NAME>']
# + id="HpDLY_GrCIQs" colab_type="code" outputId="79f4aec6-7ff3-49a9-db9d-ad7d81176bc3" colab={"base_uri": "https://localhost:8080/", "height": 351}
for row in niles:
plt.scatter(x=niles['y'], y=niles['x'])
plt.ylim(0,120)
plt.xlim(0,53.3)
| BigDataBowl.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ------------------------------------------------------------------------------------------
# Copyright (c) Microsoft Corporation. All rights reserved.
# Licensed under the MIT License (MIT). See LICENSE in the repo root for license information.
# ------------------------------------------------------------------------------------------
import sys
import os
# # GradCam visualizations
# +
import matplotlib.pyplot as plt
import numpy as np
from typing import Optional, Any
def _plot_one_map(map: np.ndarray,
image: np.ndarray,
channel: int,
slice: int,
row: int,
col: Optional[int]) -> None:
"""
Plots one visualization overlaid on one image for one
specific channel and slice at a chosen subplot.
"""
if col is not None:
current_ax = ax[row, col]
else:
current_ax = ax[row]
current_ax.imshow(image[channel, slice], cmap=plt.gray())
pos = current_ax.imshow(
map[channel, slice],
vmin=map.min(),
vmax=map.max(),
cmap=plt.jet(),
alpha=0.7
)
current_ax.set_xticks([])
current_ax.set_yticks([])
if slice == 0:
plt.colorbar(pos, plt.jet(), current_ax)
current_ax.set_anchor('W')
# + tags=["parameters"]
subject_id = "The subject id"
gradcam_dir = "Path/to/gradcam"
has_non_image_features = True
has_image_features = True
probas = ["[0], [0]"]
ground_truth_labels = ["[0], [0]"]
non_image_labels = [""]
encode_jointly = False
imaging_feature_type = "Image"
target_position = "Default"
value_image_and_segmentation = "ImageAndSegmentation"
# -
if has_image_features:
image = np.load(os.path.join(os.path.join(gradcam_dir, "image.npy")))
grad_cam = np.load(os.path.join(gradcam_dir, "gradcam.npy"))
guided_grad_cam = np.load(os.path.join(gradcam_dir, "guided_grad_cam.npy"))
if has_non_image_features:
gradcam_non_image_features = np.load(os.path.join(gradcam_dir, "non_image_pseudo_cam.npy"))
# +
if has_image_features:
channels, slices = image.shape[:2]
if imaging_feature_type == value_image_and_segmentation:
channels //= 2
else:
channels = gradcam_non_image_features.shape[0]
if has_non_image_features:
gradcam_non_image_features = gradcam_non_image_features / gradcam_non_image_features.sum() * 100
# -
# ### Model prediction
print(f"Subject ID: {subject_id}")
print(f"Target position: {target_position}")
print(f"Probability predicted by the model {probas}")
print(f"Ground truth label {ground_truth_labels}")
# ### Plot features importance
if has_non_image_features:
fig, ax = plt.subplots(figsize=(20,7))
x_data = np.arange(len(non_image_labels))
ax.bar(x_data, gradcam_non_image_features.flatten()) # type: ignore
ax.set_xticks(x_data)
ax.set_xticklabels(non_image_labels, rotation=90)
ax.set_title(f"Relative non-imaging feature importance (%)")
else:
print("This model only uses imaging features")
# ### GradCam maps
if has_image_features:
if encode_jointly:
fig, ax = plt.subplots(slices, 1, figsize=(10, 5*slices))
for i in range(slices):
_plot_one_map(grad_cam, image, 0, i, i, None)
else:
fig, ax = plt.subplots(slices, channels, figsize=(20, 2*slices))
for i in range(slices):
for channel in range(channels):
_plot_one_map(grad_cam, image, channel, i, i, channel)
else:
"This model is not using images"
# ### GuidedGradCam maps
# +
figure_title = "GuidedGrad for images" if imaging_feature_type != value_image_and_segmentation \
else "GuidedGradCam for segmentations"
if has_image_features:
fig, ax = plt.subplots(slices, channels, figsize=(20, 2*slices))
for i in range(slices):
for channel in range(channels):
_plot_one_map(guided_grad_cam, image, channel, i, i, channel)
plt.suptitle(figure_title)
else:
"This model is not using images"
# -
if imaging_feature_type == value_image_and_segmentation and has_image_features:
fig, ax = plt.subplots(slices, channels, figsize=(20, 2*slices))
for i in range(slices):
for channel in range(channels):
_plot_one_map(guided_grad_cam, image, channel + channels, i, i, channel)
plt.suptitle("GuidedGrad for Segmentations (imaging_type is ImageAndSegmentation)")
| InnerEye/ML/visualizers/gradcam_visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tfod
# language: python
# name: tfod
# ---
# + [markdown] id="QUANWN3rpfC9"
# # 0. Setup Paths
# + id="146BB11JpfDA"
import os
# + id="42hJEdo_pfDB"
CUSTOM_MODEL_NAME = 'my_ssd_mobnet'
PRETRAINED_MODEL_NAME = 'ssd_mobilenet_v2_fpnlite_320x320_coco17_tpu-8'
PRETRAINED_MODEL_URL = 'http://download.tensorflow.org/models/object_detection/tf2/20200711/ssd_mobilenet_v2_fpnlite_320x320_coco17_tpu-8.tar.gz'
TF_RECORD_SCRIPT_NAME = 'generate_tfrecord.py'
LABEL_MAP_NAME = 'label_map.pbtxt'
# + id="hbPhYVy_pfDB"
paths = {
'WORKSPACE_PATH': os.path.join('Tensorflow', 'workspace'),
'SCRIPTS_PATH': os.path.join('Tensorflow','scripts'),
'APIMODEL_PATH': os.path.join('Tensorflow','models'),
'ANNOTATION_PATH': os.path.join('Tensorflow', 'workspace','annotations'),
'IMAGE_PATH': os.path.join('Tensorflow', 'workspace','images'),
'MODEL_PATH': os.path.join('Tensorflow', 'workspace','models'),
'PRETRAINED_MODEL_PATH': os.path.join('Tensorflow', 'workspace','pre-trained-models'),
'CHECKPOINT_PATH': os.path.join('Tensorflow', 'workspace','models',CUSTOM_MODEL_NAME),
'OUTPUT_PATH': os.path.join('Tensorflow', 'workspace','models',CUSTOM_MODEL_NAME, 'export'),
'TFJS_PATH':os.path.join('Tensorflow', 'workspace','models',CUSTOM_MODEL_NAME, 'tfjsexport'),
'TFLITE_PATH':os.path.join('Tensorflow', 'workspace','models',CUSTOM_MODEL_NAME, 'tfliteexport'),
'PROTOC_PATH':os.path.join('Tensorflow','protoc')
}
# + id="LwhWZMI0pfDC"
files = {
'PIPELINE_CONFIG':os.path.join('Tensorflow', 'workspace','models', CUSTOM_MODEL_NAME, 'pipeline.config'),
'TF_RECORD_SCRIPT': os.path.join(paths['SCRIPTS_PATH'], TF_RECORD_SCRIPT_NAME),
'LABELMAP': os.path.join(paths['ANNOTATION_PATH'], LABEL_MAP_NAME)
}
# + id="HR-TfDGrpfDC"
for path in paths.values():
if not os.path.exists(path):
if os.name == 'posix':
# !mkdir -p {path}
if os.name == 'nt':
# !mkdir {path}
# + [markdown] id="OLU-rs_ipfDE"
# # 1. Download TF Models Pretrained Models from Tensorflow Model Zoo and Install TFOD
# +
# https://www.tensorflow.org/install/source_windows
# + id="K-Cmz2edpfDE"
if os.name=='nt':
# !pip install wget
import wget
# + id="iA1DIq5OpfDE"
if not os.path.exists(os.path.join(paths['APIMODEL_PATH'], 'research', 'object_detection')):
# !git clone https://github.com/tensorflow/models {paths['APIMODEL_PATH']}
# + id="rJjMHbnDs3Tv"
# Install Tensorflow Object Detection
if os.name=='posix':
# !apt-get install protobuf-compiler
# !cd Tensorflow/models/research && protoc object_detection/protos/*.proto --python_out=. && cp object_detection/packages/tf2/setup.py . && python -m pip install .
if os.name=='nt':
url="https://github.com/protocolbuffers/protobuf/releases/download/v3.15.6/protoc-3.15.6-win64.zip"
wget.download(url)
# !move protoc-3.15.6-win64.zip {paths['PROTOC_PATH']}
# !cd {paths['PROTOC_PATH']} && tar -xf protoc-3.15.6-win64.zip
os.environ['PATH'] += os.pathsep + os.path.abspath(os.path.join(paths['PROTOC_PATH'], 'bin'))
# !cd Tensorflow/models/research && protoc object_detection/protos/*.proto --python_out=. && copy object_detection\\packages\\tf2\\setup.py setup.py && python setup.py build && python setup.py install
# !cd Tensorflow/models/research/slim && pip install -e .
# -
VERIFICATION_SCRIPT = os.path.join(paths['APIMODEL_PATH'], 'research', 'object_detection', 'builders', 'model_builder_tf2_test.py')
# Verify Installation
# !python {VERIFICATION_SCRIPT}
# !pip install tensorflow --upgrade
# !pip uninstall protobuf matplotlib -y
# !pip install protobuf matplotlib==3.2
import object_detection
# !pip list
# + colab={"base_uri": "https://localhost:8080/"} id="csofht2npfDE" outputId="ff5471b2-bed2-43f2-959c-327a706527b6"
if os.name =='posix':
# !wget {PRETRAINED_MODEL_URL}
# !mv {PRETRAINED_MODEL_NAME+'.tar.gz'} {paths['PRETRAINED_MODEL_PATH']}
# !cd {paths['PRETRAINED_MODEL_PATH']} && tar -zxvf {PRETRAINED_MODEL_NAME+'.tar.gz'}
if os.name == 'nt':
wget.download(PRETRAINED_MODEL_URL)
# !move {PRETRAINED_MODEL_NAME+'.tar.gz'} {paths['PRETRAINED_MODEL_PATH']}
# !cd {paths['PRETRAINED_MODEL_PATH']} && tar -zxvf {PRETRAINED_MODEL_NAME+'.tar.gz'}
# + [markdown] id="M5KJTnkfpfDC"
# # 2. Create Label Map
# + id="p1BVDWo7pfDC"
labels = [{'name':'ThumbsUp', 'id':1}, {'name':'ThumbsDown', 'id':2}, {'name':'ThankYou', 'id':3}, {'name':'LiveLong', 'id':4}]
with open(files['LABELMAP'], 'w') as f:
for label in labels:
f.write('item { \n')
f.write('\tname:\'{}\'\n'.format(label['name']))
f.write('\tid:{}\n'.format(label['id']))
f.write('}\n')
# + [markdown] id="C88zyVELpfDC"
# # 3. Create TF records
# + colab={"base_uri": "https://localhost:8080/"} id="kvf5WccwrFGq" outputId="49902aeb-0bd7-4298-e1a0-5b4a64eb2064"
# OPTIONAL IF RUNNING ON COLAB
ARCHIVE_FILES = os.path.join(paths['IMAGE_PATH'], 'archive.tar.gz')
if os.path.exists(ARCHIVE_FILES):
# !tar -zxvf {ARCHIVE_FILES}
# + colab={"base_uri": "https://localhost:8080/"} id="KWpb_BVUpfDD" outputId="56ce2a3f-3933-4ee6-8a9d-d5ec65f7d73c"
if not os.path.exists(files['TF_RECORD_SCRIPT']):
# !git clone https://github.com/nicknochnack/GenerateTFRecord {paths['SCRIPTS_PATH']}
# + colab={"base_uri": "https://localhost:8080/"} id="UPFToGZqpfDD" outputId="0ebb456f-aadc-4a1f-96e6-fbfec1923e1c"
# !python {files['TF_RECORD_SCRIPT']} -x {os.path.join(paths['IMAGE_PATH'], 'train')} -l {files['LABELMAP']} -o {os.path.join(paths['ANNOTATION_PATH'], 'train.record')}
# !python {files['TF_RECORD_SCRIPT']} -x {os.path.join(paths['IMAGE_PATH'], 'test')} -l {files['LABELMAP']} -o {os.path.join(paths['ANNOTATION_PATH'], 'test.record')}
# + [markdown] id="qT4QU7pLpfDE"
# # 4. Copy Model Config to Training Folder
# + id="cOjuTFbwpfDF"
if os.name =='posix':
# !cp {os.path.join(paths['PRETRAINED_MODEL_PATH'], PRETRAINED_MODEL_NAME, 'pipeline.config')} {os.path.join(paths['CHECKPOINT_PATH'])}
if os.name == 'nt':
# !copy {os.path.join(paths['PRETRAINED_MODEL_PATH'], PRETRAINED_MODEL_NAME, 'pipeline.config')} {os.path.join(paths['CHECKPOINT_PATH'])}
# + [markdown] id="Ga8gpNslpfDF"
# # 5. Update Config For Transfer Learning
# + id="Z9hRrO_ppfDF"
import tensorflow as tf
from object_detection.utils import config_util
from object_detection.protos import pipeline_pb2
from google.protobuf import text_format
# + id="c2A0mn4ipfDF"
config = config_util.get_configs_from_pipeline_file(files['PIPELINE_CONFIG'])
# + colab={"base_uri": "https://localhost:8080/"} id="uQA13-afpfDF" outputId="907496a4-a39d-4b13-8c2c-e5978ecb1f10"
config
# + id="9vK5lotDpfDF"
pipeline_config = pipeline_pb2.TrainEvalPipelineConfig()
with tf.io.gfile.GFile(files['PIPELINE_CONFIG'], "r") as f:
proto_str = f.read()
text_format.Merge(proto_str, pipeline_config)
# + id="rP43Ph0JpfDG"
pipeline_config.model.ssd.num_classes = len(labels)
pipeline_config.train_config.batch_size = 4
pipeline_config.train_config.fine_tune_checkpoint = os.path.join(paths['PRETRAINED_MODEL_PATH'], PRETRAINED_MODEL_NAME, 'checkpoint', 'ckpt-0')
pipeline_config.train_config.fine_tune_checkpoint_type = "detection"
pipeline_config.train_input_reader.label_map_path= files['LABELMAP']
pipeline_config.train_input_reader.tf_record_input_reader.input_path[:] = [os.path.join(paths['ANNOTATION_PATH'], 'train.record')]
pipeline_config.eval_input_reader[0].label_map_path = files['LABELMAP']
pipeline_config.eval_input_reader[0].tf_record_input_reader.input_path[:] = [os.path.join(paths['ANNOTATION_PATH'], 'test.record')]
# + id="oJvfgwWqpfDG"
config_text = text_format.MessageToString(pipeline_config)
with tf.io.gfile.GFile(files['PIPELINE_CONFIG'], "wb") as f:
f.write(config_text)
# + [markdown] id="Zr3ON7xMpfDG"
# # 6. Train the model
# + id="B-Y2UQmQpfDG"
TRAINING_SCRIPT = os.path.join(paths['APIMODEL_PATH'], 'research', 'object_detection', 'model_main_tf2.py')
# + id="jMP2XDfQpfDH"
command = "python {} --model_dir={} --pipeline_config_path={} --num_train_steps=2000".format(TRAINING_SCRIPT, paths['CHECKPOINT_PATH'],files['PIPELINE_CONFIG'])
# + colab={"base_uri": "https://localhost:8080/"} id="A4OXXi-ApfDH" outputId="117a0e83-012b-466e-b7a6-ccaa349ac5ab"
print(command)
# + colab={"base_uri": "https://localhost:8080/"} id="i3ZsJR-qpfDH" outputId="cabec5e1-45e6-4f2f-d9cf-297d9c1d0225"
# !{command}
# + [markdown] id="4_YRZu7npfDH"
# # 7. Evaluate the Model
# + id="80L7-fdPpfDH"
command = "python {} --model_dir={} --pipeline_config_path={} --checkpoint_dir={}".format(TRAINING_SCRIPT, paths['CHECKPOINT_PATH'],files['PIPELINE_CONFIG'], paths['CHECKPOINT_PATH'])
# + colab={"base_uri": "https://localhost:8080/"} id="lYsgEPx9pfDH" outputId="8632d48b-91d2-45d9-bcb8-c1b172bf6eed"
print(command)
# + id="lqTV2jGBpfDH"
# !{command}
# + [markdown] id="orvRk02UpfDI"
# # 8. Load Train Model From Checkpoint
# + id="8TYk4_oIpfDI"
import os
import tensorflow as tf
from object_detection.utils import label_map_util
from object_detection.utils import visualization_utils as viz_utils
from object_detection.builders import model_builder
from object_detection.utils import config_util
# + id="tDnQg-cYpfDI"
# Load pipeline config and build a detection model
configs = config_util.get_configs_from_pipeline_file(files['PIPELINE_CONFIG'])
detection_model = model_builder.build(model_config=configs['model'], is_training=False)
# Restore checkpoint
ckpt = tf.compat.v2.train.Checkpoint(model=detection_model)
ckpt.restore(os.path.join(paths['CHECKPOINT_PATH'], 'ckpt-5')).expect_partial()
@tf.function
def detect_fn(image):
image, shapes = detection_model.preprocess(image)
prediction_dict = detection_model.predict(image, shapes)
detections = detection_model.postprocess(prediction_dict, shapes)
return detections
# + [markdown] id="0EmsmbBZpfDI"
# # 9. Detect from an Image
# + id="Y_MKiuZ4pfDI"
import cv2
import numpy as np
from matplotlib import pyplot as plt
# %matplotlib inline
# + id="cBDbIhNapfDI"
category_index = label_map_util.create_category_index_from_labelmap(files['LABELMAP'])
# + id="Lx3crOhOzITB"
IMAGE_PATH = os.path.join(paths['IMAGE_PATH'], 'test', 'livelong.02533422-940e-11eb-9dbd-5cf3709bbcc6.jpg')
# + colab={"base_uri": "https://localhost:8080/", "height": 269} id="Tpzn1SMry1yK" outputId="c392a2c5-10fe-4fc4-9998-a1d4c7db2bd3"
img = cv2.imread(IMAGE_PATH)
image_np = np.array(img)
input_tensor = tf.convert_to_tensor(np.expand_dims(image_np, 0), dtype=tf.float32)
detections = detect_fn(input_tensor)
num_detections = int(detections.pop('num_detections'))
detections = {key: value[0, :num_detections].numpy()
for key, value in detections.items()}
detections['num_detections'] = num_detections
# detection_classes should be ints.
detections['detection_classes'] = detections['detection_classes'].astype(np.int64)
label_id_offset = 1
image_np_with_detections = image_np.copy()
viz_utils.visualize_boxes_and_labels_on_image_array(
image_np_with_detections,
detections['detection_boxes'],
detections['detection_classes']+label_id_offset,
detections['detection_scores'],
category_index,
use_normalized_coordinates=True,
max_boxes_to_draw=5,
min_score_thresh=.8,
agnostic_mode=False)
plt.imshow(cv2.cvtColor(image_np_with_detections, cv2.COLOR_BGR2RGB))
plt.show()
# + [markdown] id="IsNAaYAo0WVL"
# # 10. Real Time Detections from your Webcam
# -
# !pip uninstall opencv-python-headless -y
# + id="o_grs6OGpfDJ"
cap = cv2.VideoCapture(0)
width = int(cap.get(cv2.CAP_PROP_FRAME_WIDTH))
height = int(cap.get(cv2.CAP_PROP_FRAME_HEIGHT))
while cap.isOpened():
ret, frame = cap.read()
image_np = np.array(frame)
input_tensor = tf.convert_to_tensor(np.expand_dims(image_np, 0), dtype=tf.float32)
detections = detect_fn(input_tensor)
num_detections = int(detections.pop('num_detections'))
detections = {key: value[0, :num_detections].numpy()
for key, value in detections.items()}
detections['num_detections'] = num_detections
# detection_classes should be ints.
detections['detection_classes'] = detections['detection_classes'].astype(np.int64)
label_id_offset = 1
image_np_with_detections = image_np.copy()
viz_utils.visualize_boxes_and_labels_on_image_array(
image_np_with_detections,
detections['detection_boxes'],
detections['detection_classes']+label_id_offset,
detections['detection_scores'],
category_index,
use_normalized_coordinates=True,
max_boxes_to_draw=5,
min_score_thresh=.8,
agnostic_mode=False)
cv2.imshow('object detection', cv2.resize(image_np_with_detections, (800, 600)))
if cv2.waitKey(10) & 0xFF == ord('q'):
cap.release()
cv2.destroyAllWindows()
break
# + [markdown] id="rzlM4jt0pfDJ"
# # 10. Freezing the Graph
# + id="n4olHB2npfDJ"
FREEZE_SCRIPT = os.path.join(paths['APIMODEL_PATH'], 'research', 'object_detection', 'exporter_main_v2.py ')
# + id="0AjO93QDpfDJ"
command = "python {} --input_type=image_tensor --pipeline_config_path={} --trained_checkpoint_dir={} --output_directory={}".format(FREEZE_SCRIPT ,files['PIPELINE_CONFIG'], paths['CHECKPOINT_PATH'], paths['OUTPUT_PATH'])
# + colab={"base_uri": "https://localhost:8080/"} id="F6Lsp3tCpfDJ" outputId="c3828529-bf06-4df5-d7f3-145890ec3edd"
print(command)
# + colab={"base_uri": "https://localhost:8080/"} id="1Sw1ULgHpfDJ" outputId="6fd441e1-9fc9-4889-d072-3395c21e40b6"
# !{command}
# + [markdown] id="wTPmdqaXpfDK"
# # 11. Conversion to TFJS
# + colab={"base_uri": "https://localhost:8080/"} id="gZ6UzY_fpfDK" outputId="0c84722e-1c2b-4002-d857-80827ade828a"
# !pip install tensorflowjs
# + id="0oxbVynHpfDK"
command = "tensorflowjs_converter --input_format=tf_saved_model --output_node_names='detection_boxes,detection_classes,detection_features,detection_multiclass_scores,detection_scores,num_detections,raw_detection_boxes,raw_detection_scores' --output_format=tfjs_graph_model --signature_name=serving_default {} {}".format(os.path.join(paths['OUTPUT_PATH'], 'saved_model'), paths['TFJS_PATH'])
# + colab={"base_uri": "https://localhost:8080/"} id="DB2AGNmJpfDK" outputId="fbc9f747-f511-47e8-df8f-5ea65cef0374"
print(command)
# + colab={"base_uri": "https://localhost:8080/"} id="K7rfT4-hpfDK" outputId="532707fd-6feb-4bc6-84a3-325b5d16303c"
# !{command}
# + id="o8_hm-itpfDK"
# Test Code: https://github.com/nicknochnack/RealTimeSignLanguageDetectionwithTFJS
# + [markdown] id="VtUw73FHpfDK"
# # 12. Conversion to TFLite
# + id="XviMtewLpfDK"
TFLITE_SCRIPT = os.path.join(paths['APIMODEL_PATH'], 'research', 'object_detection', 'export_tflite_graph_tf2.py ')
# + id="us86cjC4pfDL"
command = "python {} --pipeline_config_path={} --trained_checkpoint_dir={} --output_directory={}".format(TFLITE_SCRIPT ,files['PIPELINE_CONFIG'], paths['CHECKPOINT_PATH'], paths['TFLITE_PATH'])
# + colab={"base_uri": "https://localhost:8080/"} id="n1r5YO3rpfDL" outputId="5fcdf7a4-eee2-4365-f1ca-1751968379ea"
print(command)
# + colab={"base_uri": "https://localhost:8080/"} id="I-xWpHN8pfDL" outputId="7f6bacd8-d077-43b5-c131-5b081fba24a4"
# !{command}
# + id="iJfYMbN6pfDL"
FROZEN_TFLITE_PATH = os.path.join(paths['TFLITE_PATH'], 'saved_model')
TFLITE_MODEL = os.path.join(paths['TFLITE_PATH'], 'saved_model', 'detect.tflite')
# -
command = "tflite_convert \
--saved_model_dir={} \
--output_file={} \
--input_shapes=1,300,300,3 \
--input_arrays=normalized_input_image_tensor \
--output_arrays='TFLite_Detection_PostProcess','TFLite_Detection_PostProcess:1','TFLite_Detection_PostProcess:2','TFLite_Detection_PostProcess:3' \
--inference_type=FLOAT \
--allow_custom_ops".format(FROZEN_TFLITE_PATH, TFLITE_MODEL, )
# + colab={"base_uri": "https://localhost:8080/"} id="E8GwUeoFpfDL" outputId="fac43ea4-cc85-471b-a362-e994b06fd583"
print(command)
# + colab={"base_uri": "https://localhost:8080/"} id="Nbd7gqHMpfDL" outputId="7c8fe6d5-2415-4641-8548-39d425c202f7"
# !{command}
# + [markdown] id="5NQqZRdA21Uc"
# # 13. Zip and Export Models
# + id="tTVTGCQp2ZJJ"
# !tar -czf models.tar.gz {paths['CHECKPOINT_PATH']}
# + colab={"base_uri": "https://localhost:8080/"} id="whShhB0x3PYJ" outputId="b773201d-35c9-46a8-b893-4a76bd4d5d97"
from google.colab import drive
drive.mount('/content/drive')
| 2. Training and Detection.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# - Two other moving averages are commonly used among financial market :
# - Weighted Moving Average (WMA)
# - Exponential Moving Average (EMA)
# ## Weighted Moving Average
# - In some applications, one of the limitations of the simple moving average is that it gives equal weight to each of the daily prices included in the window. E.g., in a 10-day moving average, the most recent day receives the same weight as the first day in the window: each price receives a 10% weighting.
#
# - Compared to the Simple Moving Average, the Linearly Weighted Moving Average (or simply Weighted Moving Average, WMA), gives more weight to the most recent price and gradually less as we look back in time. On a 10-day weighted average, the price of the 10th day would be multiplied by 10, that of the 9th day by 9, the 8th day by 8 and so on. The total will then be divided by the sum of the weights (in this case: 55). In this specific example, the most recent price receives about 18.2% of the total weight, the second more recent 16.4%, and so on until the oldest price in the window that receives 0.02% of the weight.
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib as mpl
# - We apply a style for our charts. If you’re using Jupyter it’s a good idea to add the %matplotlib inline instruction (and skip plt.show() when creating charts):
plt.style.use('fivethirtyeight')
data = pd.read_csv('MV EMV.csv', index_col = 'Date')
df = pd.read_csv('cs movavg.csv', index_col = 'Date')
data.index = pd.to_datetime(data.index)
df.index = pd.to_datetime(df.index)
# We can drop the old index column:
data = data.drop(columns='Unnamed: 0')
df = df.drop(columns='Unnamed: 0')
display(data.head(15))
display(df.head(15))
(data.shape), (df.shape)
# - We are going to consider only the Price and 10-Day WMA columns for now and move to the EMA later on
weights = np.arange(1,11) #this creates an array with integers 1 to 10 included
weights
# - When it comes to linearly weighted moving averages, the pandas library does not have a ready off-the-shelf method to calculate them. It offers, however, a very powerful and flexible method: ** .apply()** This method allows us to create and pass any custom function to a rolling window: that is how we are going to calculate our Weighted Moving Average. To calculate a 10-Day WMA, we start by creating an array of weights - whole numbers from 1 to 10:
wma10 = data['Price'].rolling(10).apply(lambda prices: np.dot(prices, weights)/weights.sum(), raw=True)
wma10.head(20)
# ### Study
df['Our 10-day WMA'] = np.round(wma10, decimals=3)
df[['Price', '10-day WMA', 'Our 10-day WMA']].head(20)
# - The two WMA columns look the same. There are a few differences in the third decimal place, but we can put that down to rounding error and conclude that our implementation of the WMA is correct. In a real-life application, if we want to be more rigorous we should compute the differences between the two columns and check that they are not too large. For now, we keep things simple and we can be satisfied with the visual inspection.
# -------------
sma10 = data['Price'].rolling(10).mean()
data['10-day SMA'] = np.round(sma10, decimals=3)
data['10-day WMA'] = np.round(wma10, decimals=3)
data[['Price', '10-day SMA', '10-day WMA']].head(20)
plt.figure(figsize = (12,6))
plt.plot(data['Price'], label="Price")
plt.plot(data['10-day WMA'], label="10-Day WMA")
plt.plot(data['10-day SMA'], label="10-Day SMA")
plt.xlabel("Date")
plt.ylabel("Price")
plt.legend()
plt.show()
# - As we can see, both averages smooth out the price movement. The WMA is more reactive and follows the price closer than the SMA: we expect that since the WMA gives more weight to the most recent price observations. Also, both moving average series start on day 10: the first day with enough available data to compute the averages.
# ------------------------
# ## Exponential Moving Average
# - Similarly to the Weighted Moving Average, the Exponential Moving Average (EMA) assigns a greater weight to the most recent price observations. While it assigns lesser weight to past data, it is based on a recursive formula that includes in its calculation all the past data in our price series.
#
# - Pandas includes a method to compute the EMA moving average of any time series: .ewm().
ema10 = data['Price'].ewm(span=10).mean()
ema10.head(10)
df['Our 10-day EMA'] = np.round(ema10, decimals=3)
df[['Price', '10-day EMA', 'Our 10-day EMA']].head(20)
# - As you have already noticed, we have a problem here: the 10-day EMA that we just calculated does not correspond to the one calculated in the downloaded spreadsheet. One starts on day 10, while the other starts on day 1. Also, the values do not match exactly.
#
#
# - Is our calculation wrong? Or is the calculation in the provided spreadsheet wrong? Neither: those two series correspond to two different definitions of EMA. To be more specific, the formula used to compute the EMA is the same. What changes is just the use of the initial values.
#
#
# - If we look carefully at the definition of Exponential Moving Average on the StockCharts.com web page we can notice one important detail: they start calculating a 10-day moving average on day 10, disregarding the previous days and replacing the price on day 10 with its 10-day SMA. It’s a different definition than the one applied when we calculated the EMA using the .ewm() method directly.
#
#
# - The following lines of code create a new modified price series where the first 9 prices (when the SMA is not available) are replaced by NaN and the price on the 10th date becomes its 10-Day SMA:
modPrice = df['Price'].copy()
modPrice.iloc[0:10] = sma10[0:10]
modPrice.head(20)
# - We can use this modified price series to calculate a second version of the EWM. By looking at the documentation, we can note that the .ewm() method has an adjust parameter that defaults to True. This parameter adjusts the weights to account for the imbalance in the beginning periods (if you need more detail, see the Exponentially weighted windows section in the pandas documentation).
#
#
# - If we want to emulate the EMA as in our spreadsheet using our modified price series, we don’t need this adjustment. We then set adjust=False:
ema10alt = modPrice.ewm(span=10, adjust=False).mean()
df['Our 2nd 10-Day EMA'] = np.round(ema10alt, decimals=3)
df[['Price', '10-day EMA', 'Our 10-day EMA', 'Our 2nd 10-Day EMA']].head(20)
# - Now, we are doing much better. We have obtained an EMA series that matches the one calculated in the spreadsheet.
#
#
# - We ended up with two different versions of EMA in our hands:
#
#
# **1. ema10:** This version uses the plain .ewm() method, starts at the beginning of our price history but does not match the definition used in the spreadsheet.
#
# **2. ema10alt:** This version starts on day 10 (with an initial value equal to the 10-day SMA) and matches the definition on our spreadsheet.
#
#
# - Which one is the best to use? The answer is: it depends on what we need for our application and to build our system. If we need an EMA series that starts from day 1, then we should choose the first one. On the other hand, if we need to use our average in combination with other averages that have no values for the initial days (such as the SMA), then the second is probably the best one.
#
#
# - The second EMA is widely used among financial market analysts: if we need to implement an already existing system, we need to be careful to use the correct definition. Otherwise, the results may not be what is expected from us and may put the accuracy of all of our work into question. In any case, the numeric difference between those two averages is minimal, with an impact on our trading or investment decision system limited to the initial days.
plt.figure(figsize = (12,6))
plt.plot(df['Price'], label="Price")
plt.plot(wma10, label="10-Day WMA")
plt.plot(sma10, label="10-Day SMA")
plt.plot(ema10, label="10-Day EMA-1")
plt.plot(ema10alt, label="10-Day EMA-2")
plt.xlabel("Date")
plt.ylabel("Price")
plt.legend()
plt.show()
# - Of all the moving averages, the **WMA (Weighted Moving Average)** appears the one that is **more responsive** and tags the **price more closely**, while the SMA is the one that responds with some more lag. The two versions of the EMA tend to overlap each other, mainly in the last days.
# ---------------
| 28_Stock_Market/1_Moving_Average/2_Weighted & Exponential.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fit $k_{ij}$ interactions parameter of Hexane and Ethanol
#
# This notebook has te purpose of showing how to optimize the $k_{ij}$ for a mixture in SGTPy.
#
# First it's needed to import the necessary modules
# +
import numpy as np
from SGTPy import component, mixture, saftvrmie
from SGTPy.fit import fit_kij
# -
# Now that the functions are available it is necessary to create the mixture.
# +
ethanol = component('ethanol2C', ms = 1.7728, sigma = 3.5592 , eps = 224.50,
lambda_r = 11.319, lambda_a = 6., eAB = 3018.05, rcAB = 0.3547,
rdAB = 0.4, sites = [1,0,1], cii= 5.3141080872882285e-20)
hexane = component('hexane', ms = 1.96720036, sigma = 4.54762477, eps = 377.60127994,
lambda_r = 18.41193194, cii = 3.581510586936205e-19)
mix = mixture(hexane, ethanol)
# -
# Now the experimental equilibria data is read and a tuple is created. It includes the experimental liquid composition, vapor composition, equilibrium temperature and pressure. This is done with ```datavle = (Xexp, Yexp, Texp, Pexp)```
#
# +
# Experimental data obtained from <NAME>. Chem. Eng. Data, vol. 5, no. 3, pp. 243–247, 1960.
# Experimental temperature saturation in K
Texp = np.array([351.45, 349.15, 346.35, 340.55, 339.05, 334.95, 332.55, 331.85,
331.5 , 331.25, 331.15, 331.4 , 331.6 , 332.3 , 333.35, 336.65,
339.85, 341.85])
# Experimental pressure in Pa
Pexp = np.array([101330., 101330., 101330., 101330., 101330., 101330., 101330.,
101330., 101330., 101330., 101330., 101330., 101330., 101330.,
101330., 101330., 101330., 101330.])
# Experimental liquid composition
Xexp = np.array([[0. , 0.01 , 0.02 , 0.06 , 0.08 , 0.152, 0.245, 0.333, 0.452,
0.588, 0.67 , 0.725, 0.765, 0.898, 0.955, 0.99 , 0.994, 1. ],
[1. , 0.99 , 0.98 , 0.94 , 0.92 , 0.848, 0.755, 0.667, 0.548,
0.412, 0.33 , 0.275, 0.235, 0.102, 0.045, 0.01 , 0.006, 0. ]])
# Experimental vapor composition
Yexp = np.array([[0. , 0.095, 0.193, 0.365, 0.42 , 0.532, 0.605, 0.63 , 0.64 ,
0.65 , 0.66 , 0.67 , 0.675, 0.71 , 0.745, 0.84 , 0.935, 1. ],
[1. , 0.905, 0.807, 0.635, 0.58 , 0.468, 0.395, 0.37 , 0.36 ,
0.35 , 0.34 , 0.33 , 0.325, 0.29 , 0.255, 0.16 , 0.065, 0. ]])
datavle = (Xexp, Yexp, Texp, Pexp)
# -
# The function ```fit_kij``` optimize the $k_{ij}$. The function requires bounds for the parameter, as well as the mixture object and the equilibria data.
# bounds fors kij
kij_bounds = (-0.01, 0.01)
fit_kij(kij_bounds, mix, datavle = datavle)
# For more information just run:
# ```fit_kij?```
| SGTPy-examples/SGTPy's paper notebooks/Fit Equilibrium Hexane + Etanol.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # This notebook is for uploading models to AWS S3
import os
import boto3
# ## Get AWS keys saved in environment variables
aws_access_key = os.getenv('AWS_ACCESS_KEY_ID', 'default')
aws_secret_access_key = os.getenv('AWS_SECRET_ACCESS_KEY', 'default')
def connect_s3(bucket_name, aws_access_key, aws_secret_access_key):
# get aws credentials
client = boto3.client("s3", aws_access_key_id=aws_access_key, \
aws_secret_access_key=aws_secret_access_key)
s3 = boto3.resource("s3")
bucket = s3.Bucket(bucket_name)
file_name_list = [file.key for file in bucket.objects.all()]
return file_name_list, client
client = boto3.client("s3", aws_access_key_id=aws_access_key, \
aws_secret_access_key=aws_secret_access_key)
s3 = boto3.resource("s3")
for bucket in s3.buckets.all():
print(bucket.name)
# # Upload function
import logging
from botocore.exceptions import ClientError
def upload_file(file_name, bucket, object_name=None):
"""Upload a file to an S3 bucket
:param file_name: File to upload
:param bucket: Bucket to upload to
:param object_name: S3 object name. If not specified then file_name is used
:return: True if file was uploaded, else False
"""
# If S3 object_name was not specified, use file_name
if object_name is None:
object_name = file_name
# Upload the file
#s3_client = boto3.client('s3')
try:
response = client.upload_file(file_name, bucket, object_name)
except ClientError as e:
logging.error(e)
return False
return True
client.upload_file('../st_webapp/models/lrpipe_cg.pkl','smartrelocator','lrpipe_cg.pkl')
client.upload_file('../st_webapp/test3.py','smartrelocator','test3.py')
client.upload_file('../st_webapp/relocatortools.py','smartrelocator','relocatortools.py')
client.upload_file('../st_webapp/breedlist.csv','smartrelocator','breedlist.csv')
client.upload_file('../st_webapp/statelist.csv','smartrelocator','statelist.csv')
| src/data_acquisition/s3_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# ## Combining DataFrames
import pandas as pd
articles_df = pd.read_csv('articles.csv', dtype={'LanguageId':str})
articles_df[articles_df.LanguageId == '4']
articles_df = pd.read_csv('articles.csv', encoding='utf8')
articles_df = pd.read_csv('articles.csv',
parse_dates=[['Year', 'Month', 'Day']],
keep_date_col=True)
articles_df[['Year_Month_Day', 'Year', 'Month', 'Day']]
articles_df
articles_df = pd.read_csv('articles.csv',
parse_dates={'Date': ['Year', 'Month', 'Day']},
keep_date_col=True)
articles_df[(articles_df.Date >= '2015-07-01') & (articles_df.Date < '2015-08-01')]
articles_df = pd.read_csv('articles.csv',
parse_dates={'Date': ['Year', 'Month', 'Day']},
keep_date_col=True,
usecols=['First_Author', 'Year', 'Month', 'Day'])
articles_df.drop(['Day', 'Month', 'Year'], axis=1)
articles_df.to_csv('ournew.csv', encoding='utf8')
new_df = pd.read_csv('ournew.csv', encoding='utf8', index_col=0)
new_df
| notebooks/Episode 4.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1> Hyper-parameter tuning </h1>
#
# **Learning Objectives**
# 1. Understand various approaches to hyperparameter tuning
# 2. Automate hyperparameter tuning using AI Platform HyperTune
#
# ## Introduction
#
# In the previous notebook we achieved an RMSE of **4.13**. Let's see if we can improve upon that by tuning our hyperparameters.
#
# Hyperparameters are parameters that are set *prior* to training a model, as opposed to parameters which are learned *during* training.
#
# These include learning rate and batch size, but also model design parameters such as type of activation function and number of hidden units.
#
# Here are the four most common ways to finding the ideal hyperparameters:
# 1. Manual
# 2. Grid Search
# 3. Random Search
# 4. Bayesian Optimzation
#
# **1. Manual**
#
# Traditionally, hyperparameter tuning is a manual trial and error process. A data scientist has some intution about suitable hyperparameters which they use as a starting point, then they observe the result and use that information to try a new set of hyperparameters to try to beat the existing performance.
#
# Pros
# - Educational, builds up your intuition as a data scientist
# - Inexpensive because only one trial is conducted at a time
#
# Cons
# - Requires alot of time and patience
#
# **2. Grid Search**
#
# On the other extreme we can use grid search. Define a discrete set of values to try for each hyperparameter then try every possible combination.
#
# Pros
# - Can run hundreds of trials in parallel using the cloud
# - Gauranteed to find the best solution within the search space
#
# Cons
# - Expensive
#
# **3. Random Search**
#
# Alternatively define a range for each hyperparamter (e.g. 0-256) and sample uniformly at random from that range.
#
# Pros
# - Can run hundreds of trials in parallel using the cloud
# - Requires less trials than Grid Search to find a good solution
#
# Cons
# - Expensive (but less so than Grid Search)
#
# **4. Bayesian Optimization**
#
# Unlike Grid Search and Random Search, Bayesian Optimization takes into account information from past trials to select parameters for future trials. The details of how this is done is beyond the scope of this notebook, but if you're interested you can read how it works here [here](https://cloud.google.com/blog/products/gcp/hyperparameter-tuning-cloud-machine-learning-engine-using-bayesian-optimization).
#
# Pros
# - Picks values intelligenty based on results from past trials
# - Less expensive because requires fewer trials to get a good result
#
# Cons
# - Requires sequential trials for best results, takes longer
#
# **AI Platform HyperTune**
#
# AI Platform HyperTune, powered by [Google Vizier](https://ai.google/research/pubs/pub46180), uses Bayesian Optimization by default, but [also supports](https://cloud.google.com/ml-engine/docs/tensorflow/hyperparameter-tuning-overview#search_algorithms) Grid Search and Random Search.
#
#
# When tuning just a few hyperparameters (say less than 4), Grid Search and Random Search work well, but when tunining several hyperparameters and the search space is large Bayesian Optimization is best.
PROJECT = "qwiklabs-gcp-00-34ffb0f0dc65" # Replace with your PROJECT
BUCKET = "cloud-training-bucket" # Replace with your BUCKET
REGION = "us-central1" # Choose an available region for AI Platform
TFVERSION = "1.14" # TF version for AI Platform
import os
os.environ["PROJECT"] = PROJECT
os.environ["BUCKET"] = BUCKET
os.environ["REGION"] = REGION
os.environ["TFVERSION"] = TFVERSION
# ## Move code into python package
#
# Let's package our updated code with feature engineering so it's AI Platform compatible.
# + language="bash"
# mkdir taxifaremodel
# touch taxifaremodel/__init__.py
# -
# ## Create model.py
#
# Note that any hyperparameters we want to tune need to be exposed as command line arguments. In particular note that the number of hidden units is now a parameter.
# +
# %%writefile taxifaremodel/model.py
import tensorflow as tf
import numpy as np
import shutil
print(tf.__version__)
#1. Train and Evaluate Input Functions
CSV_COLUMN_NAMES = ["fare_amount","dayofweek","hourofday","pickuplon","pickuplat","dropofflon","dropofflat"]
CSV_DEFAULTS = [[0.0],[1],[0],[-74.0],[40.0],[-74.0],[40.7]]
def read_dataset(csv_path):
def _parse_row(row):
# Decode the CSV row into list of TF tensors
fields = tf.decode_csv(records = row, record_defaults = CSV_DEFAULTS)
# Pack the result into a dictionary
features = dict(zip(CSV_COLUMN_NAMES, fields))
# NEW: Add engineered features
features = add_engineered_features(features)
# Separate the label from the features
label = features.pop("fare_amount") # remove label from features and store
return features, label
# Create a dataset containing the text lines.
dataset = tf.data.Dataset.list_files(file_pattern = csv_path) # (i.e. data_file_*.csv)
dataset = dataset.flat_map(map_func = lambda filename:tf.data.TextLineDataset(filenames = filename).skip(count = 1))
# Parse each CSV row into correct (features,label) format for Estimator API
dataset = dataset.map(map_func = _parse_row)
return dataset
def train_input_fn(csv_path, batch_size = 128):
#1. Convert CSV into tf.data.Dataset with (features,label) format
dataset = read_dataset(csv_path)
#2. Shuffle, repeat, and batch the examples.
dataset = dataset.shuffle(buffer_size = 1000).repeat(count = None).batch(batch_size = batch_size)
return dataset
def eval_input_fn(csv_path, batch_size = 128):
#1. Convert CSV into tf.data.Dataset with (features,label) format
dataset = read_dataset(csv_path)
#2.Batch the examples.
dataset = dataset.batch(batch_size = batch_size)
return dataset
#2. Feature Engineering
# One hot encode dayofweek and hourofday
fc_dayofweek = tf.feature_column.categorical_column_with_identity(key = "dayofweek", num_buckets = 7)
fc_hourofday = tf.feature_column.categorical_column_with_identity(key = "hourofday", num_buckets = 24)
# Cross features to get combination of day and hour
fc_day_hr = tf.feature_column.crossed_column(keys = [fc_dayofweek, fc_hourofday], hash_bucket_size = 24 * 7)
# Bucketize latitudes and longitudes
NBUCKETS = 16
latbuckets = np.linspace(start = 38.0, stop = 42.0, num = NBUCKETS).tolist()
lonbuckets = np.linspace(start = -76.0, stop = -72.0, num = NBUCKETS).tolist()
fc_bucketized_plat = tf.feature_column.bucketized_column(source_column = tf.feature_column.numeric_column(key = "pickuplon"), boundaries = lonbuckets)
fc_bucketized_plon = tf.feature_column.bucketized_column(source_column = tf.feature_column.numeric_column(key = "pickuplat"), boundaries = latbuckets)
fc_bucketized_dlat = tf.feature_column.bucketized_column(source_column = tf.feature_column.numeric_column(key = "dropofflon"), boundaries = lonbuckets)
fc_bucketized_dlon = tf.feature_column.bucketized_column(source_column = tf.feature_column.numeric_column(key = "dropofflat"), boundaries = latbuckets)
def add_engineered_features(features):
features["dayofweek"] = features["dayofweek"] - 1 # subtract one since our days of week are 1-7 instead of 0-6
features["latdiff"] = features["pickuplat"] - features["dropofflat"] # East/West
features["londiff"] = features["pickuplon"] - features["dropofflon"] # North/South
features["euclidean_dist"] = tf.sqrt(features["latdiff"]**2 + features["londiff"]**2)
return features
feature_cols = [
#1. Engineered using tf.feature_column module
tf.feature_column.indicator_column(categorical_column = fc_day_hr),
fc_bucketized_plat,
fc_bucketized_plon,
fc_bucketized_dlat,
fc_bucketized_dlon,
#2. Engineered in input functions
tf.feature_column.numeric_column(key = "latdiff"),
tf.feature_column.numeric_column(key = "londiff"),
tf.feature_column.numeric_column(key = "euclidean_dist")
]
#3. Serving Input Receiver Function
def serving_input_receiver_fn():
receiver_tensors = {
'dayofweek' : tf.placeholder(dtype = tf.int32, shape = [None]), # shape is vector to allow batch of requests
'hourofday' : tf.placeholder(dtype = tf.int32, shape = [None]),
'pickuplon' : tf.placeholder(dtype = tf.float32, shape = [None]),
'pickuplat' : tf.placeholder(dtype = tf.float32, shape = [None]),
'dropofflat' : tf.placeholder(dtype = tf.float32, shape = [None]),
'dropofflon' : tf.placeholder(dtype = tf.float32, shape = [None]),
}
features = add_engineered_features(receiver_tensors) # 'features' is what is passed on to the model
return tf.estimator.export.ServingInputReceiver(features = features, receiver_tensors = receiver_tensors)
#4. Train and Evaluate
def train_and_evaluate(params):
OUTDIR = params["output_dir"]
model = tf.estimator.DNNRegressor(
hidden_units = params["hidden_units"].split(","), # NEW: paramaterize architecture
feature_columns = feature_cols,
model_dir = OUTDIR,
config = tf.estimator.RunConfig(
tf_random_seed = 1, # for reproducibility
save_checkpoints_steps = max(100, params["train_steps"] // 10) # checkpoint every N steps
)
)
# Add custom evaluation metric
def my_rmse(labels, predictions):
pred_values = tf.squeeze(input = predictions["predictions"], axis = -1)
return {"rmse": tf.metrics.root_mean_squared_error(labels = labels, predictions = pred_values)}
model = tf.contrib.estimator.add_metrics(model, my_rmse)
train_spec = tf.estimator.TrainSpec(
input_fn = lambda: train_input_fn(params["train_data_path"]),
max_steps = params["train_steps"])
exporter = tf.estimator.FinalExporter(name = "exporter", serving_input_receiver_fn = serving_input_receiver_fn) # export SavedModel once at the end of training
# Note: alternatively use tf.estimator.BestExporter to export at every checkpoint that has lower loss than the previous checkpoint
eval_spec = tf.estimator.EvalSpec(
input_fn = lambda: eval_input_fn(params["eval_data_path"]),
steps = None,
start_delay_secs = 1, # wait at least N seconds before first evaluation (default 120)
throttle_secs = 1, # wait at least N seconds before each subsequent evaluation (default 600)
exporters = exporter) # export SavedModel once at the end of training
tf.logging.set_verbosity(v = tf.logging.INFO) # so loss is printed during training
shutil.rmtree(path = OUTDIR, ignore_errors = True) # start fresh each time
tf.estimator.train_and_evaluate(model, train_spec, eval_spec)
# -
# ## Create task.py
# #### **Exercise 1**
#
# The code cell below has two TODOs for you to complete.
#
# Firstly, in model.py above we set the number of hidden units in our model to be a hyperparameter. This means `hidden_units` must be exposed as a command line argument when we submit our training job to Cloud ML Engine. Modify the code below to add an flag for `hidden_units`. Be sure to include a description for the `help` field and specify the data `type` that the model should expect to receive. You can also include a `default` value. Look to the other parser arguments to make sure you have the formatting corret.
#
# Second, when doing hyperparameter tuning we need to make sure the output directory is different for each run, otherwise successive runs will overwrite previous runs. In `task.py` below, add some code to append the trial_id to the output direcroty of the training job.
#
# **Hint**: You can use `json.loads(os.environ.get('TF_CONFIG', '{}')).get('task', {}).get('trial', '')` to extract the trial id of the training job. You will want to append this quanity to the output directory `args['output_dir']` to make sure the output directory is different for each run.
# +
# %%writefile taxifaremodel/task.py
import argparse
import json
import os
from . import model
if __name__ == "__main__":
parser = argparse.ArgumentParser()
parser.add_argument(
# TODO: Your code goes here
)
parser.add_argument(
"--train_data_path",
help = "GCS or local path to training data",
required = True
)
parser.add_argument(
"--train_steps",
help = "Steps to run the training job for (default: 1000)",
type = int,
default = 1000
)
parser.add_argument(
"--eval_data_path",
help = "GCS or local path to evaluation data",
required = True
)
parser.add_argument(
"--output_dir",
help = "GCS location to write checkpoints and export models",
required = True
)
parser.add_argument(
"--job-dir",
help="This is not used by our model, but it is required by gcloud",
)
args = parser.parse_args().__dict__
# Append trial_id to path so trials don"t overwrite each other
# This code can be removed if you are not using hyperparameter tuning
args["output_dir"] = os.path.join(
# TODO: Your code goes here
)
# Run the training job
model.train_and_evaluate(args)
# -
# ## Create hypertuning configuration
#
# We specify:
# 1. How many trials to run (`maxTrials`) and how many of those trials can be run in parrallel (`maxParallelTrials`)
# 2. Which algorithm to use (in this case `GRID_SEARCH`)
# 3. Which metric to optimize (`hyperparameterMetricTag`)
# 4. The search region in which to constrain the hyperparameter search
#
# Full specification options [here](https://cloud.google.com/ml-engine/reference/rest/v1/projects.jobs#HyperparameterSpec).
#
# Here we are just tuning one parameter, the number of hidden units, and we'll run all trials in parrallel. However more commonly you would tune multiple hyperparameters.
# #### **Exercise 2**
#
# Add some additional hidden units to the `hyperparam.yaml` file below to potetially explore during the hyperparameter job.
# %%writefile hyperparam.yaml
trainingInput:
scaleTier: BASIC
hyperparameters:
goal: MINIMIZE
maxTrials: 10
maxParallelTrials: 10
hyperparameterMetricTag: rmse
enableTrialEarlyStopping: True
algorithm: GRID_SEARCH
params:
- parameterName: hidden_units
type: CATEGORICAL
categoricalValues:
- 10,10
- 64,32
- 128,64,32
- # TODO: Your code goes here
# ## Run the training job|
#
# Same as before with the addition of `--config=hyperpam.yaml` to reference the file we just created.
#
# This will take about 20 minutes. Go to [cloud console](https://console.cloud.google.com/mlengine/jobs) and click on the job id. Once the job is completed, the choosen hyperparameters and resulting objective value (RMSE in this case) will be shown. Trials will sorted from best to worst.
# #### **Exercise 3**
#
# Submit a hyperparameter tuning job to the cloud. Fill in the missing arguments below. This is similar to the exercise you completed in the `02_tensorlfow/g_distributed` notebook. Note that one difference here is that we now specify a `config` parameter giving the location of our .yaml file.
OUTDIR="gs://{}/taxifare/trained_hp_tune".format(BUCKET)
# !gsutil -m rm -rf # TODO: Your code goes here
# !gcloud ai-platform # TODO: Your code goes here
--package-path= # TODO: Your code goes here
--module-name= # TODO: Your code goes here
--config= # TODO: Your code goes here
--job-dir= # TODO: Your code goes here
--python-version= # TODO: Your code goes here
--runtime-version= # TODO: Your code goes here
--region= # TODO: Your code goes here
-- \
--train_data_path=gs://{BUCKET}/taxifare/smallinput/taxi-train.csv \
--eval_data_path=gs://{BUCKET}/taxifare/smallinput/taxi-valid.csv \
--train_steps=5000 \
--output_dir={OUTDIR}
# ## Results
#
# The best result is RMSE **4.02** with hidden units = 128,64,32.
#
# This improvement is modest, but now that we have our hidden units tuned let's run on our larger dataset to see if it helps.
#
# Note the passing of hyperparameter values via command line
OUTDIR="gs://{}/taxifare/trained_large_tuned".format(BUCKET)
# !gsutil -m rm -rf {OUTDIR} # start fresh each time
# !gcloud ai-platform jobs submit training taxifare_large_$(date -u +%y%m%d_%H%M%S) \
# --package-path=taxifaremodel \
# --module-name=taxifaremodel.task \
# --job-dir=gs://{BUCKET}/taxifare \
# --python-version=3.5 \
# --runtime-version={TFVERSION} \
# --region={REGION} \
# --scale-tier=STANDARD_1 \
# -- \
# --train_data_path=gs://cloud-training-demos/taxifare/large/taxi-train*.csv \
# --eval_data_path=gs://cloud-training-demos/taxifare/small/taxi-valid.csv \
# --train_steps=200000 \
# --output_dir={OUTDIR} \
# --hidden_units="128,64,32"
# ## Analysis
#
# Our RMSE improved to **3.85**
# ## Challenge excercise
#
# Try to beat the current RMSE:
#
# - Try adding more engineered features or modifying existing ones
# - Try tuning additional hyperparameters
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
| courses/machine_learning/deepdive/03_model_performance/labs/d_hyperparameter_tuning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/EduardoMoraesRitter/Titanic-Machine-Learning-from-Disaster/blob/master/clusterizacao_kmean_titanic.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="XSB1BuMHmHQ8" colab_type="text"
# https://chrisalbon.com/machine_learning/clustering/k-means_clustering/
# + id="MHUHBJZPnyKw" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 735} outputId="0ac0f035-1b79-4623-9358-726edcadcbc6"
import pandas as pd
import matplotlib.pyplot as pl
import seaborn as sns
train = pd.read_csv('https://raw.githubusercontent.com/EduardoMoraesRitter/Titanic-Machine-Learning-from-Disaster/master/train.csv')
print(train.head())
train_simples = train.drop(['PassengerId', 'Name', 'Ticket', 'Cabin', 'PassengerId', 'SibSp' , 'Parch', 'Pclass', 'Embarked', 'Sex', 'Survived'], axis=1)
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(strategy="median")
imputer.fit(train_simples)
train_no_null = imputer.transform(train_simples)
train_simples = pd.DataFrame(train_no_null, columns=train_simples.columns)
print(train_simples.head())
from sklearn.cluster import KMeans
from sklearn.metrics import adjusted_rand_score
model = KMeans(n_clusters=3)
model.fit(train_simples)
print("Top terms per cluster:")
order_centroids = model.cluster_centers_.argsort()[:, ::-1]
for i in range(3):
print("\n Cluster %d:" % i),
for ind in order_centroids[i, :100]:
print(train_simples.iloc[ind].tolist())#(' %s' % terms[ind]),
print
pl.scatter(train_simples.Age, train_simples.Fare, c = model.labels_)
# + [markdown] id="IxAcs3vOnx6l" colab_type="text"
# https://www.naftaliharris.com/blog/visualizing-dbscan-clustering/
# + id="5z8J1oUKZrsG" colab_type="code" colab={}
import pandas as pd
import matplotlib.pyplot as pl
import seaborn as sns
from sklearn.cluster import KMeans
from sklearn.metrics import adjusted_rand_score
# + id="as8y87tCZeTM" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="82b690c0-0f78-44b3-dd50-6b1732db9908"
train = pd.read_csv('https://raw.githubusercontent.com/EduardoMoraesRitter/Titanic-Machine-Learning-from-Disaster/master/train.csv')
test = pd.read_csv('https://raw.githubusercontent.com/EduardoMoraesRitter/Titanic-Machine-Learning-from-Disaster/master/test.csv')
train.head()
# + id="HEWckp__Zivk" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 504} outputId="5e98ed96-d97f-4b9c-dca8-f679399c6fc8"
pl.figure(figsize=(10,8))
sns.heatmap(train.corr(), annot=True, cmap='Greens')
# + id="QzDePv1Hj9sf" colab_type="code" colab={}
train.drop(['Name', 'Ticket', 'Cabin', 'PassengerId'], axis=1, inplace=True)
test.drop(['Name', 'Ticket', 'Cabin'], axis=1, inplace=True)
# + id="WMMFbYBWj9v2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 437} outputId="9a2bf003-fe9a-40cf-fdc0-ed6232a28330"
print('antes \n', train.isnull().sum())
from sklearn.impute import SimpleImputer
imputer = SimpleImputer(strategy="median")
train_numerico = train.drop(['Sex','Embarked'], axis=1)
test_numerico = test.drop(['Sex','Embarked'], axis=1)
imputer.fit(train_numerico)
train_no_null = imputer.transform(train_numerico)
imputer.fit(test_numerico)
test_no_null = imputer.transform(test_numerico)
train_tr = pd.DataFrame(train_no_null, columns=train_numerico.columns)
test_tr = pd.DataFrame(test_no_null, columns=test_numerico.columns)
print('\n depois \n', train_tr.isnull().sum())
print(train_tr.head())
# + id="RROtiChwkEUt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="42f8dcf9-76a7-44ae-9aef-a42a0474a716"
train_cat = train[['Sex', 'Embarked']]
test_cat = test[['Sex', 'Embarked']]
train_cat_encoded = pd.get_dummies(train_cat)
test_cat_encoded = pd.get_dummies(test_cat)
train_cat_encoded.head()
train = train_tr.join(train_cat_encoded)
train = test_tr.join(test_cat_encoded)
train.head()
# + id="7A1wLaZl1tdb" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 195} outputId="5c03e2e8-2344-4cc1-ad83-365064f4c4b5"
train.drop(['Pclass', 'PassengerId'], axis=1, inplace=True)
train.head()
# + id="Vl6IS04crYlt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 614} outputId="f1a6c08a-301b-4789-9724-849d14242d57"
Nc = range(1, 10)
kmeans_list = [KMeans(n_clusters=i) for i in Nc]
print(kmeans_list)
score = [kmeans_list[i].fit(train).score(train) for i in range(len(kmeans_list))]
score
pl.plot(Nc,score)
pl.xlabel('Number of Clusters')
pl.ylabel('Score')
pl.title('Elbow Curve')
pl.show()
# + id="lU110JXN2vsI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 246} outputId="d954c3b0-b4c9-4d7d-ec22-4b9af87f1dac"
pl.figure(figsize=(12, 6))
pl.plot(range(1, 40), error, color='black', linestyle='dashed', marker='o',
markerfacecolor='grey', markersize=10)
pl.title('Error Rate K Value')
pl.xlabel('K Value')
pl.ylabel('Mean Error')
# + id="EtQbikwOZjuI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="23a4c457-1bda-4ec5-b065-8c25ddebdfd5"
model = KMeans(n_clusters=4, init='k-means++', max_iter=100, n_init=1)
model.fit(train)
# + id="VeMKHzRkaJok" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 353} outputId="6464727f-9408-4c16-cce8-65bae584e7fe"
#model.cluster_centers_.argsort()[:, ::-1]
model.labels_
#order_centroids = model.cluster_centers_.argsort()[:, ::-1]
#order_centroids
# for i in range(4):
# print("\n Cluster %d:" % i),
# for ind in order_centroids[i, :1000]:
# print(ind)#train.iloc[ind].tolist())
# print
# + id="OrEB5ZqOiP9H" colab_type="code" colab={}
import numpy as np
z = pl.scatter(train.Fare, train.Age, train.Pclass, c = model.labels_)
# + [markdown] id="KPfuNtd2tTOt" colab_type="text"
# https://scikit-learn.org/stable/auto_examples/cluster/plot_cluster_iris.html
# + id="mqoYHqUMnyaF" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="dea6d0be-9414-4728-ee9d-9b14d3f855ce"
print(__doc__)
# Code source: <NAME>
# Modified for documentation by <NAME>
# License: BSD 3 clause
import numpy as np
import matplotlib.pyplot as plt
# Though the following import is not directly being used, it is required
# for 3D projection to work
from mpl_toolkits.mplot3d import Axes3D
from sklearn.cluster import KMeans
from sklearn import datasets
np.random.seed(5)
iris = datasets.load_iris()
X = iris.data
y = iris.target
estimators = [('k_means_iris_8', KMeans(n_clusters=8)),
('k_means_iris_3', KMeans(n_clusters=3)),
('k_means_iris_bad_init', KMeans(n_clusters=3, n_init=1,
init='random'))]
fignum = 1
titles = ['8 clusters', '3 clusters', '3 clusters, bad initialization']
for name, est in estimators:
fig = plt.figure(fignum, figsize=(10, 8))
ax = Axes3D(fig, rect=[0, 0, .95, 1], elev=48, azim=134)
est.fit(X)
labels = est.labels_
ax.scatter(X[:, 3], X[:, 0], X[:, 2],
c=labels.astype(np.float), edgecolor='k')
ax.w_xaxis.set_ticklabels([])
ax.w_yaxis.set_ticklabels([])
ax.w_zaxis.set_ticklabels([])
ax.set_xlabel('Petal width')
ax.set_ylabel('Sepal length')
ax.set_zlabel('Petal length')
ax.set_title(titles[fignum - 1])
ax.dist = 12
fignum = fignum + 1
# Plot the ground truth
fig = plt.figure(fignum, figsize=(10, 8))
ax = Axes3D(fig, rect=[0, 0, .95, 1], elev=48, azim=134)
for name, label in [('Setosa', 0),
('Versicolour', 1),
('Virginica', 2)]:
ax.text3D(X[y == label, 3].mean(),
X[y == label, 0].mean(),
X[y == label, 2].mean() + 2, name,
horizontalalignment='center',
bbox=dict(alpha=.2, edgecolor='w', facecolor='w'))
# Reorder the labels to have colors matching the cluster results
y = np.choose(y, [1, 2, 0]).astype(np.float)
ax.scatter(X[:, 3], X[:, 0], X[:, 2], c=y, edgecolor='k')
ax.w_xaxis.set_ticklabels([])
ax.w_yaxis.set_ticklabels([])
ax.w_zaxis.set_ticklabels([])
ax.set_xlabel('Petal width')
ax.set_ylabel('Sepal length')
ax.set_zlabel('Petal length')
ax.set_title('Ground Truth')
ax.dist = 12
fig.show()
# + id="g3bq9k8TtT1N" colab_type="code" colab={}
| clusterizacao_kmean_titanic.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# # Introduction to Data Science
# # Activity for Lecture 10: Linear Regression 2
# *COMP 5360 / MATH 4100, University of Utah, http://datasciencecourse.net/*
#
# Name:
#
# Email:
#
# UID:
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Class exercise: analysis of the credit dataset
#
# Recall the 'Credit' dataset introduced in class and available [here](http://www-bcf.usc.edu/~gareth/ISL/data.html).
# This dataset consists of some credit card information for 400 people.
#
# First import the data and convert income to thousands.
#
# +
# imports and setup
import scipy as sc
import numpy as np
import pandas as pd
import statsmodels.formula.api as sm #Last lecture: used statsmodels.formula.api.ols() for OLS
from sklearn import linear_model #Last lecture: used sklearn.linear_model.LinearRegression() for OLS
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['figure.figsize'] = (10, 6)
from mpl_toolkits.mplot3d import Axes3D
from matplotlib import cm
# Import data from Credit.csv file
credit = pd.read_csv('Credit.csv',index_col=0) #load data
credit["Income"] = credit["Income"].map(lambda x: 1000*x)
credit
# -
# ## Activity 1: A First Regression Model
#
# **Exercise:** First regress Limit on Rating:
# $$
# \text{Limit} = \beta_0 + \beta_1 \text{Rating}.
# $$
# Since credit ratings are primarily used by banks to determine credit limits, we expect that Rating is very predictive for Limit, so this regression should be very good.
#
# Use the 'ols' function from the statsmodels python library. What is the $R^2$ value? What are $H_0$ and $H_A$ for the associated hypothesis test and what is the $p$-value?
#
# your code goes here
# **Your answer goes here:**
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activity 2: Predicting Limit without Rating
#
# Since Rating and Limit are almost the same variable, next we'll forget about Rating and just try to predict Limit from the real-valued variables (non-categorical variables): Income, Cards, Age, Education, Balance.
#
# **Exercise:** Develop a multilinear regression model to predict Rating. Interpret the results.
#
# For now, just focus on the real-valued variables (Income, Cards, Age, Education, Balance)
# and ignore the categorical variables (Gender, Student, Married, Ethnicity).
#
#
# + slideshow={"slide_type": "-"}
# your code goes here
# + [markdown] slideshow={"slide_type": "-"}
# Which independent variables are good/bad predictors? What is the best overall model?
#
# **Your observations:**
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Activity 3: Incorporating Categorical Variables Into Regression Models
#
# Now consider the binary categorical variables which we mapped to integer 0, 1 values in class.
# + slideshow={"slide_type": "-"}
credit["Gender_num"] = credit["Gender"].map({' Male':0, 'Female':1})
credit["Student_num"] = credit["Student"].map({'Yes':1, 'No':0})
credit["Married_num"] = credit["Married"].map({'Yes':1, 'No':0})
# + [markdown] slideshow={"slide_type": "-"}
# Can you improve the model you developed in Activity 2 by incorporating one or more of these variables?
#
# + slideshow={"slide_type": "-"}
# your code here
# -
# **Your answer goes here:**
| 10-LinearRegression2/10-LinearRegression2_Activity.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IHA2 - Catching Pokemon
# 
# In this home assignment, you'll apply roughly the same principles we used when doing logistic regression on the Iris dataset, but on a new and very interesting dataset. We'll use the [Predict'em All](https://www.kaggle.com/semioniy/predictemall) dataset from Kaggle (download the dataset directly from them). This dataset consists of roughly 293,000 [pokemon](http://www.pokemongo.com/) sightings (historical appearances of Pokemon in the Pokemon Go game), having coordinates, time, weather, population density, distance to pokestops/ gyms etc. as features. A comprehensive list of all the features is available at [the dataset's homepage](https://www.kaggle.com/semioniy/predictemall)
#
# The context is simple: you are a Pokemon hunter, and there are only three Pokemon left for you to complete your collection. You'll do anything to capture them, including changing where you'll spend your next holidays! You know that some Pokemon only spawn in certain places of the world. Since you like machine learning so much, you figure it would be a great idea to train a classifier that, based on a location's latitude and longitude, can tell us which Pokemon is more likely to appear there.
#
# The assignment is broken down into six steps.
#
# 1. Loading the data and extracting the desired subset of it
# 2. Visualization of the dataset
# 3. Preprocessing
# 4. Training
# 5. Evaluation
# 6. Exploration
#
#
# Feel free to add cells wherever you see fit, and play around with this notebook as much as you want when developing the solutions. However, the solution you upload to ping-pong must have the exact format shown here, with only the cells present here.
#
# Don't restrict yourself only to what was taught so far. Some of the tasks might require you to search for new information. [The python docs](https://docs.python.org/3/), [keras docs](https://keras.io/), [stackoverflow](https://stackoverflow.com/), and Google are your friends!
# ## 0. Imports
# Import any necessary modules here.
# ## 1. Loading and extracting subset
# Load the `'300k.csv'` file using pandas.
# Create a new `DataFrame` with only the columns `latitude`, `longitude`, and `class`.
# Note that the `class` column specifies which pokemon it is. However, it only has the numerical id of the pokemon. For convenience, use the following dictionary to convert between ids and names.
name_dict={1: 'Bulbasaur', 2: 'Ivysaur', 3: 'Venusaur', 4: 'Charmander', 5: 'Charmeleon', 6: 'Charizard', 7: 'Squirtle', 8:
'Wartortle', 9: 'Blastoise', 10: 'Caterpie', 11: 'Metapod', 12: 'Butterfree', 13: 'Weedle', 14: 'Kakuna',
15: 'Beedrill', 16: 'Pidgey', 17: 'Pidgeotto', 18: 'Pidgeot', 19: 'Rattata', 20: 'Raticate', 21: 'Spearow',
22: 'Fearow', 23: 'Ekans', 24: 'Arbok', 25: 'Pikachu', 26: 'Raichu', 27: 'Sandshrew', 28: 'Sandslash',
29: '<NAME>', 30: 'Nidorina', 31: 'Nidoqueen',32: '<NAME>', 33: 'Nidorino', 34: 'Nidoking', 35: 'Clefairy',
36: 'Clefable', 37: 'Vulpix', 38: 'Ninetales', 39: 'Jigglypuff', 40: 'Wigglytuff', 41: 'Zubat', 42: 'Golbat',
43: 'Oddish', 44: 'Gloom', 45: 'Vileplume', 46: 'Paras', 47: 'Parasect', 48: 'Venonat', 49: 'Venomoth',
50: 'Diglett', 51: 'Dugtrio', 52: 'Meowth', 53: 'Persian', 54: 'Psyduck',55: 'Golduck', 56: 'Mankey',
57: 'Primeape', 58: 'Growlithe', 59: 'Arcanine', 60: 'Poliwag', 61: 'Poliwhirl', 62: 'Poliwrath',
63: 'Abra', 64: 'Kadabra', 65: 'Alakazam', 66: 'Machop', 67: 'Machoke', 68: 'Machamp', 69: 'Bellsprout',
70: 'Weepinbell', 71: 'Victreebel', 72: 'Tentacool', 73: 'Tentacruel', 74: 'Geodude', 75: 'Graveler',
76: 'Golem', 77: 'Ponyta', 78: 'Rapidash', 79: 'Slowpoke', 80: 'Slowbro', 81: 'Magnemite', 82: 'Magneton',
83: "Farfetch'd", 84: 'Doduo', 85: 'Dodrio', 86: 'Seel', 87: 'Dewgong', 88: 'Grimer', 89: 'Muk',
90: 'Shellder', 91: 'Cloyster', 92: 'Gastly', 93: 'Haunter', 94: 'Gengar', 95: 'Onix', 96: 'Drowzee',
97: 'Hypno', 98: 'Krabby', 99: 'Kingler', 100: 'Voltorb', 101: 'Electrode', 102: 'Exeggcute', 103: 'Exeggutor',
104: 'Cubone', 105: 'Marowak', 106: 'Hitmonlee', 107: 'Hitmonchan', 108: 'Lickitung', 109: 'Koffing',
110: 'Weezing', 111: 'Rhyhorn', 112: 'Rhydon', 113: 'Chansey', 114: 'Tangela', 115: 'Kangaskhan', 116: 'Horsea',
117: 'Seadra', 118: 'Goldeen', 119: 'Seaking', 120: 'Staryu', 121: 'Starmie', 122: 'Mr. Mime', 123: 'Scyther',
124: 'Jynx', 125: 'Electabuzz', 126: 'Magmar', 127: 'Pinsir', 128: 'Tauros', 129: 'Magikarp', 130: 'Gyarados',
131: 'Lapras', 132: 'Ditto', 133: 'Eevee', 134: 'Vaporeon', 135: 'Jolteon', 136: 'Flareon', 137: 'Porygon',
138: 'Omanyte', 139: 'Omastar', 140: 'Kabuto', 141: 'Kabutops', 142: 'Aerodactyl', 143: 'Snorlax', 144: 'Articuno',
145: 'Zapdos', 146: 'Moltres', 147: 'Dratini', 148: 'Dragonair', 149: 'Dragonite', 150: 'Mewtwo', 'Bulbasaur': 1, 'Ivysaur': 2, 'Venusaur': 3, 'Charmander': 4, 'Charmeleon': 5, 'Charizard': 6, 'Squirtle': 7, 'Wartortle': 8, 'Blastoise': 9, 'Caterpie': 10, 'Metapod': 11, 'Butterfree': 12, 'Weedle': 13, 'Kakuna': 14, 'Beedrill': 15, 'Pidgey': 16, 'Pidgeotto': 17, 'Pidgeot': 18, 'Rattata': 19, 'Raticate': 20, 'Spearow': 21, 'Fearow': 22, 'Ekans': 23, 'Arbok': 24, 'Pikachu': 25, 'Raichu': 26, 'Sandshrew': 27, 'Sandslash': 28, '<NAME>': 29, 'Nidorina': 30, 'Nidoqueen': 31, '<NAME>': 32, 'Nidorino': 33, 'Nidoking': 34, 'Clefairy': 35, 'Clefable': 36, 'Vulpix': 37, 'Ninetales': 38, 'Jigglypuff': 39, 'Wigglytuff': 40, 'Zubat': 41, 'Golbat': 42, 'Oddish': 43, 'Gloom': 44, 'Vileplume': 45, 'Paras': 46, 'Parasect': 47, 'Venonat': 48, 'Venomoth': 49, 'Diglett': 50, 'Dugtrio': 51, 'Meowth': 52, 'Persian': 53, 'Psyduck': 54, 'Golduck': 55, 'Mankey': 56, 'Primeape': 57, 'Growlithe': 58, 'Arcanine': 59, 'Poliwag': 60, 'Poliwhirl': 61, 'Poliwrath': 62, 'Abra': 63, 'Kadabra': 64, 'Alakazam': 65, 'Machop': 66, 'Machoke': 67, 'Machamp': 68, 'Bellsprout': 69, 'Weepinbell': 70, 'Victreebel': 71, 'Tentacool': 72, 'Tentacruel': 73, 'Geodude': 74, 'Graveler': 75, 'Golem': 76, 'Ponyta': 77, 'Rapidash': 78, 'Slowpoke': 79, 'Slowbro': 80, 'Magnemite': 81, 'Magneton': 82, 'Farfetch\'d': 83, 'Doduo': 84, 'Dodrio': 85, 'Seel': 86, 'Dewgong': 87, 'Grimer': 88, 'Muk': 89, 'Shellder': 90, 'Cloyster': 91, 'Gastly': 92, 'Haunter': 93, 'Gengar': 94, 'Onix': 95, 'Drowzee': 96, 'Hypno': 97, 'Krabby': 98, 'Kingler': 99, 'Voltorb': 100, 'Electrode': 101, 'Exeggcute': 102, 'Exeggutor': 103, 'Cubone': 104, 'Marowak': 105, 'Hitmonlee': 106, 'Hitmonchan': 107, 'Lickitung': 108, 'Koffing': 109, 'Weezing': 110, 'Rhyhorn': 111, 'Rhydon': 112, 'Chansey': 113, 'Tangela': 114, 'Kangaskhan': 115, 'Horsea': 116, 'Seadra': 117, 'Goldeen': 118, 'Seaking': 119, 'Staryu': 120, 'Starmie': 121, 'Mr. Mime': 122, 'Scyther': 123, 'Jynx': 124, 'Electabuzz': 125, 'Magmar': 126, 'Pinsir': 127, 'Tauros': 128, 'Magikarp': 129, 'Gyarados': 130, 'Lapras': 131, 'Ditto': 132, 'Eevee': 133, 'Vaporeon': 134, 'Jolteon': 135, 'Flareon': 136, 'Porygon': 137, 'Omanyte': 138, 'Omastar': 139, 'Kabuto': 140, 'Kabutops': 141, 'Aerodactyl': 142, 'Snorlax': 143, 'Articuno': 144, 'Zapdos': 145, 'Moltres': 146, 'Dratini': 147, 'Dragonair': 148, 'Dragonite': 149, 'Mewtwo': 150}
# example usage (you can index either by name or id)
print(name_dict['Gengar'])
print(name_dict[94])
# We are only interested in three specific pokemon: Diglett, Seel, and Tauros.
# <table style="width:100%">
# <tr>
# <th> <center>Diglett</center> </th>
# <th> <center>Seel</center> </th>
# <th> <center>Tauros</center> </th>
# </tr>
# <tr>
# <td></td>
# <td></td>
# <td></td>
# </tr>
# </table>
#
#
# Filter the dataset to contain only these pokemon.
# ## 2. Visualization of the dataset
# Plot histogram of the number of occurrences of each class.
# Is the dataset balanced?
#
# **Your answer**: (fill in here)
# Plot a scatter plot where the first dimension is latitude, the second is longitude, and each point is a Pokemon. Further, the color of each point should represent which Pokemon it is. Lastly, the marker at each point should be an `'x'`. Make sure to label each axis.
#
# Hints:
#
# - The `scatter` method from `matplotlib` accepts an argument called `c`.
# - The `scatter` method also accepts an argument called `marker`.
# Is there any other visualization you think would be useful? If so, insert it here.
# How hard do you think the problem is? Which classes can/cannot be easily separated?
#
# **Your answer**: (fill in here)
#
# Which accuracy do you expect to achieve?
#
# **Your answer**: (fill in here)
# ## 3. Preprocessing
# Prepare input and output vectors.
# Separate your data into training and test sets. 20% of the data should be in the test set.
# ## 4. Training
# Choose an architecture for your network.
# Train the network. When training, separate 25% of your training data into a validation set.
# For you to pass this assignment, you must obtain an accuracy on the validation set greater than 50%. It may be necessary to search for a good architecture by trying several different ones. If you want a challenge, try getting an accuracy greater than 63%.
# ## 5. Evaluation
# Once you achieved at least 50% accuracy in the validation set, we are done with training. Now we'll evaluate the performance of your classifier on the test set.
# Compute the accuracy on the test set.
# Compute the confusion matrix of your predictions on the test set.
# ## 6. Exploration
# You have now trained and evaluated a neural network for this particular classification task. Can you provide a brief explanation as to how you could use it to decide where to travel, if you're interested in capturing the aforementioned Pokemons?
#
# **Answer**: (fill in here)
# Is(are) there any other feature(s) from the original dataset (e.g. hour of the day, pressure, wind speed, population density, etc.) which you think would be valuable to add as an input feature to your classifier to improve its performance?
#
# **Your answer**: (fill in here)
# To investigate your hypothesis, plot a histogram of the selected feature(s) for each one of the pokemons we're interested in. For example, if you think pressure and population density are valuable for prediction, plot 6 histograms. 3 of them will be the pressure histograms for each class ('Diglett', 'Seel' and 'Tauros'), and the other 3 will be the population density for each class.
# What does(do) this(ese) histogram(s) show you? Could it be beneficial to add this(ese) new feature(s) as input? Explain why/why not.
#
# **Your answer**: (fill in here)
# ## 7. (optional)
# Assuming you found useful new features in the last part of this assignment, train a new classifier that uses these featues as well. Did the accuracy on the validation set improve? What's the highest accuracy you can achieve?
| Home Assignments/IHA2/IHA2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "-"}
# @IMPORT-MERGE
import numpy as np
import pandas as pd
from plaster.tools.plots import plots
from plaster.run.run import RunResult
from plaster.run.job import JobResult
from plaster.tools.ipynb_helpers.displays import hd
from plaster.tools.zplots import zplots
# +
# @REMOVE-FROM-TEMPLATE
from plumbum import local
print(local.cwd)
job = JobResult("../../../jobs_folder/tau8_local_bg03_merge/")
z = zplots.setup()
# -
# # Precision-Recall - All Runs Combined
plots.plot_pr_for_job( job, _size=600, classifier="nn_v2" )
# # Dataframe summary of runs + CSV export
# @REMOVE-FROM-TEMPLATE
PGEN_report_precisions = (0.9,0.8,0.7)
pepstrs = run.prep.pepstrs()
display(pepstrs)
len(''.join(pepstrs.seqstr.values))
# +
# This is to control the ordering of the columns in the csv
cols = ['run_i', 'run_name', 'pro_i', 'pro_id', 'pep_i', 'pep_start', 'pep_stop', 'at_prec', 'recall_at_prec', 'score_at_prec', 'ptms', 'P2', 'seqstr', 'seqlen', 'flustr', 'flu_pros', 'false_i', 'false_type', 'false_pro_i', 'false_pep_i','false_flustr', 'false_weight']
# This is to control the sorting
sort = ['run_i','pro_i','pep_start', 'at_prec', 'recall_at_prec', 'pep_i', 'false_weight' ]
ascend = [True,True,True,False,False,True,False]
proteins_of_interest = job.get_pros_of_interest().drop_duplicates('pro_id')
include_poi_only = len(proteins_of_interest) > 0 # poi only if there are some specified
precisions = PGEN_report_precisions
n_falses = 2
pep_false_df = pd.concat([
job.all_dfs(lambda run:run.nn_v2_call_bag().false_rates_all_peps__ptm_info(prec, n_falses, protein_of_interest_only=include_poi_only))
for prec in precisions
]).sort_values(by=sort,ascending=ascend).reset_index()[cols]
display(pep_false_df)
pep_false_df.to_csv('./runs_pr_with_falses.csv',index=False,float_format="%g")
| plaster/gen/nb_templates/train_and_test_epilog_template.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv
# language: python
# name: venv
# ---
# # Quora Insincere Questions Classification
# ## Detect toxic content to improve online conversations
#
#
# +
# !wget https://github.com/ravi-ilango/odsc2020_nlp/blob/main/lab2/quora_data.zip?raw=true -O quora_data.zip
# !unzip quora_data.zip
# +
import random
import os
from datetime import datetime
#deal with tensors
import torch
#handling text data
from torchtext import data
#for attention LSTM
from torch.autograd import Variable
# +
#Reproducing same results
SEED = 2315
#Torch
torch.manual_seed(SEED)
#Cuda algorithms
torch.backends.cudnn.deterministic = True
# -
# ### Load custom dataset using torchtext.data.TabularDataset
# +
#loading custom dataset
def tokenizer(text): # create a tokenizer function
return text.split(' ')
TEXT = data.Field(tokenize=tokenizer,batch_first=True,include_lengths=True)
LABEL = data.LabelField(dtype = torch.float, batch_first=True)
training_data=data.TabularDataset(path = 'quora_data/train.csv',
format = 'csv',
fields = [
(None, None),
('text', TEXT),
('label', LABEL)
],
skip_header = True)
#print preprocessed text
print(vars(training_data.examples[0]))
# -
# ### Split into training and validation datasets
train_data, test_data = training_data.split(split_ratio=0.5, random_state = random.seed(SEED))
train_data, valid_data = train_data.split(split_ratio=0.4, random_state = random.seed(SEED))
# ### Prepare input sequence
# This step takes around ~5 min
#Build vocab dictionary
TEXT.build_vocab(train_data, min_freq=3, vectors = "glove.6B.100d")
LABEL.build_vocab(train_data)
# +
print("Size of TEXT vocabulary: {}\n".format(len(TEXT.vocab)))
print("Size of LABEL vocabulary: {}\n".format(len(LABEL.vocab)))
print("Commonly used words: {}\n".format(TEXT.vocab.freqs.most_common(10)))
#Word dictionary
#TEXT.vocab.stoi
# -
# ### Prepare training data generators
# +
#check whether cuda is available
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
#set batch size
BATCH_SIZE = 64
#Load an iterator
train_iterator, valid_iterator = data.BucketIterator.splits(
(train_data, valid_data),
batch_size = BATCH_SIZE,
sort_key = lambda x: len(x.text),
sort_within_batch=True,
device = device)
# -
# ### LSTM Classifier
# +
import torch.nn as nn
class classifier(nn.Module):
#define all the layers used in model
def __init__(self, vocab_size, embedding_dim, hidden_dim, output_dim, n_layers,
bidirectional, dropout):
#Constructor
super().__init__()
#embedding layer
self.embedding = nn.Embedding(vocab_size, embedding_dim)
#lstm layer
self.lstm = nn.LSTM(embedding_dim,
hidden_dim,
num_layers=n_layers,
bidirectional=bidirectional,
dropout=dropout,
batch_first=True)
#dense layer
self.fc = nn.Linear(hidden_dim * 2, output_dim)
#activation function
self.act = nn.Sigmoid()
def forward(self, text, text_lengths):
#text = [batch size,sent_length]
embedded = self.embedding(text)
#embedded = [batch size, sent_len, emb dim]
#packed sequence
packed_embedded = nn.utils.rnn.pack_padded_sequence(embedded, text_lengths,batch_first=True)
packed_output, (hidden, cell) = self.lstm(packed_embedded)
#hidden = [batch size, num layers * num directions,hid dim]
#cell = [batch size, num layers * num directions,hid dim]
#concat the final forward and backward hidden state
hidden = torch.cat((hidden[-2,:,:], hidden[-1,:,:]), dim = 1)
#hidden = [batch size, hid dim * num directions]
dense_outputs=self.fc(hidden)
#Final activation function
outputs=self.act(dense_outputs)
return outputs
# -
# #### Instantiate a LSTM Classifier model
# +
#define hyperparameters
size_of_vocab = len(TEXT.vocab)
embedding_dim = 100
num_hidden_nodes = 32
num_output_nodes = 1
num_layers = 2
bidirection = True
dropout = 0.2
#instantiate the model
model = classifier(size_of_vocab, embedding_dim, num_hidden_nodes,num_output_nodes, num_layers,
bidirectional = True, dropout = dropout)
# -
# +
#architecture
print(model)
#No. of trainable parameters
def count_parameters(model):
return sum(p.numel() for p in model.parameters() if p.requires_grad)
print(f'The model has {count_parameters(model):,} trainable parameters')
#Initialize the pretrained embedding ()
pretrained_embeddings = TEXT.vocab.vectors
# model.embedding.weight.data.copy_(pretrained_embeddings) # TODO PLEASE USE THIS FOR LSTM
print(pretrained_embeddings.shape)
# +
import torch.optim as optim
#define optimizer and loss
optimizer = optim.Adam(model.parameters())
criterion = nn.BCELoss()
#define metric
def binary_accuracy(preds, y):
#round predictions to the closest integer
rounded_preds = torch.round(preds)
correct = (rounded_preds == y).float()
acc = correct.sum() / len(correct)
return acc
#push to cuda if available
model = model.to(device)
criterion = criterion.to(device)
# -
device
# ### Model Train function
def train(model, iterator, optimizer, criterion):
#initialize every epoch
epoch_loss = 0
epoch_acc = 0
#set the model in training phase
model.train()
for batch in iterator:
#resets the gradients after every batch
optimizer.zero_grad()
#retrieve text and no. of words
text, text_lengths = batch.text
#convert to 1D tensor
predictions = model(text, text_lengths).squeeze()
#compute the loss
loss = criterion(predictions, batch.label)
#compute the binary accuracy
acc = binary_accuracy(predictions, batch.label)
#backpropage the loss and compute the gradients
loss.backward()
#update the weights
optimizer.step()
#loss and accuracy
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# ### Model Evaluate function
def evaluate(model, iterator, criterion):
#initialize every epoch
epoch_loss = 0
epoch_acc = 0
#deactivating dropout layers
model.eval()
#deactivates autograd
with torch.no_grad():
for batch in iterator:
#retrieve text and no. of words
text, text_lengths = batch.text
#convert to 1d tensor
predictions = model(text, text_lengths).squeeze()
#compute loss and accuracy
loss = criterion(predictions, batch.label)
acc = binary_accuracy(predictions, batch.label)
#keep track of loss and accuracy
epoch_loss += loss.item()
epoch_acc += acc.item()
return epoch_loss / len(iterator), epoch_acc / len(iterator)
# ### Check model's forward pass
for batch in train_iterator:
#retrieve text and no. of words
text, text_lengths = batch.text
print ("text.shape: ", text.shape)
print ("text_lengths.shape: ", text_lengths.shape)
#convert to 1D tensor
predictions = model(text, text_lengths)
print ("predictions.shape: ", predictions.shape)
break
model_path = 'saved_weights.pt'
# ### Train the model
#
# This step takes around ~4 min
# +
N_EPOCHS = 2
best_valid_loss = float('inf')
for epoch in range(N_EPOCHS):
#train the model
train_loss, train_acc = train(model, train_iterator, optimizer, criterion)
#evaluate the model
valid_loss, valid_acc = evaluate(model, valid_iterator, criterion)
#save the best model
if valid_loss < best_valid_loss:
best_valid_loss = valid_loss
torch.save(model.state_dict(), model_path)
ts_string = datetime.now().strftime("%m/%d/%Y %H:%M:%S")
print(f'\n {ts_string} Epoch: {epoch}')
print(f'\t Train Loss: {train_loss:.3f} | Train Acc: {train_acc*100:.2f}%')
print(f'\t Val. Loss: {valid_loss:.3f} | Val. Acc: {valid_acc*100:.2f}%')
# -
# ### Predict
# +
#load weights
model.load_state_dict(torch.load(model_path));
model.eval();
def prepare_text(sentence):
# Tokenize
tokenized = [tok for tok in tokenizer(sentence)]
# Replace tokens by index from dictionary
indexed = [TEXT.vocab.stoi[t] for t in tokenized]
length = [len(indexed)]
# Convert to tensors
tensor = torch.LongTensor(indexed).to(device)
tensor = tensor.unsqueeze(1).T
length = torch.LongTensor(length)
return tensor, length
def predict(model, sentence):
tensor, length = prepare_text(sentence)
prediction = model(tensor, length) #prediction
return prediction.item()
# -
sentence = "What is your favorite person in history?"
tokenized = [tok for tok in tokenizer(sentence)]
indexed = [TEXT.vocab.stoi[t] for t in tokenized]
[len(indexed)]
def insincere_or_not(pred):
return 'Insincere Question' if pred > .5 else 'Normal Question'
#sincere question
pred = predict(model, "What is your favorite person in history?")
print (insincere_or_not(pred))
#insincere question
pred = predict(model, "Why Indian girls go crazy about marrying <NAME>?")
print (insincere_or_not(pred))
# ### Note
#
# This notebook used data and code from a blog in https://www.analyticsvidhya.com
| lab2/quora_classifier_lstm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Using a ttf font file in Matplotlib
#
#
# Although it is usually not a good idea to explicitly point to a single ttf file
# for a font instance, you can do so using the `.font_manager.FontProperties`
# *fname* argument.
#
# Here, we use the Computer Modern roman font (``cmr10``) shipped with
# Matplotlib.
#
# For a more flexible solution, see
# :doc:`/gallery/text_labels_and_annotations/font_family_rc_sgskip` and
# :doc:`/gallery/text_labels_and_annotations/fonts_demo`.
#
# +
from pathlib import Path
import matplotlib as mpl
from matplotlib import font_manager as fm
import matplotlib.pyplot as plt
fig, ax = plt.subplots()
fpath = Path(mpl.get_data_path(), "fonts/ttf/cmr10.ttf")
prop = fm.FontProperties(fname=fpath)
ax.set_title(f'This is a special font: {fpath.name}', fontproperties=prop)
ax.set_xlabel('This is the default font')
plt.show()
# -
# ------------
#
# References
# """"""""""
#
# The use of the following functions, methods, classes and modules is shown
# in this example:
#
#
import matplotlib
matplotlib.font_manager.FontProperties
matplotlib.axes.Axes.set_title
| matplotlib/gallery_jupyter/text_labels_and_annotations/font_file.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Resolver
#
# This notebook is used to normalize dataset provided from Kaggle.
import os
import math
import numpy as np
import pandas as pd
import sys
import matplotlib.pyplot as plt
train_data = pd.read_table("data/training_set.tsv")
# ### Essay set 1
# +
set1 = train_data[train_data['essay_set'] == 1]
rating_columns = set1.columns.values[3:].tolist()
set1['avg_score']=set1[rating_columns].mean(axis=1)
set1['avg_score']=((set1[rating_columns]-set1[rating_columns].min())/(set1[rating_columns].max()-set1[rating_columns].min())).mean(axis=1)
set1[['essay_id', 'essay', 'avg_score']]
set1.hist(column='avg_score', bins=24, ax= plt.figure(figsize = (12,5)).gca())
set1.max()
# -
# ### Essay set 2
# +
set2 = train_data[train_data['essay_set'] == 2]
rating_columns = set2.columns.values[3:].tolist()
set2['avg_score']=set2[rating_columns].mean(axis=1)
set2['avg_score']=((set2[rating_columns]-set2[rating_columns].min())/(set2[rating_columns].max()-set2[rating_columns].min())).mean(axis=1)
set2[['essay_id', 'essay', 'avg_score']]
set2.hist(column='avg_score', bins=24, ax= plt.figure(figsize = (12,5)).gca())
set2.max()
# -
# ### Essay set 3
# +
set3 = train_data[train_data['essay_set'] == 3]
rating_columns = set3.columns.values[3:].tolist()
set3['avg_score']=set3[rating_columns].mean(axis=1)
set3['avg_score']=((set3[rating_columns]-set3[rating_columns].min())/(set3[rating_columns].max()-set3[rating_columns].min())).mean(axis=1)
set3[['essay_id', 'essay', 'avg_score']]
set3.hist(column='avg_score', bins=24, ax= plt.figure(figsize = (12,5)).gca())
set3.max()
# -
# ### Essay set 4
# +
set4 = train_data[train_data['essay_set'] == 4]
rating_columns = set4.columns.values[3:].tolist()
set4['avg_score']=set4[rating_columns].mean(axis=1)
set4['avg_score']=((set4[rating_columns]-set4[rating_columns].min())/(set4[rating_columns].max()-set4[rating_columns].min())).mean(axis=1)
set4[['essay_id', 'essay', 'avg_score']]
set4.hist(column='avg_score', bins=24, ax= plt.figure(figsize = (12,5)).gca())
set4.max()
# -
# ### Essay set 5
# +
set5 = train_data[train_data['essay_set'] == 5]
rating_columns = set5.columns.values[3:].tolist()
set5['avg_score']=set5[rating_columns].mean(axis=1)
set5['avg_score']=((set5[rating_columns]-set5[rating_columns].min())/(set5[rating_columns].max()-set5[rating_columns].min())).mean(axis=1)
set5[['essay_id', 'essay', 'avg_score']]
set5.hist(column='avg_score', bins=24, ax= plt.figure(figsize = (12,5)).gca())
set5.max()
# -
# ### Essay set 6
# +
set6 = train_data[train_data['essay_set'] == 6]
rating_columns = set6.columns.values[3:].tolist()
set6['avg_score']=set6[rating_columns].mean(axis=1)
set6['avg_score']=((set6[rating_columns]-set6[rating_columns].min())/(set6[rating_columns].max()-set6[rating_columns].min())).mean(axis=1)
set6[['essay_id', 'essay', 'avg_score']]
set6.hist(column='avg_score', bins=24, ax= plt.figure(figsize = (12,5)).gca())
set6.max()
# -
# ### Essay set 7
# +
set7 = train_data[train_data['essay_set'] == 7]
rating_columns = set7.columns.values[3:].tolist()
set7['avg_score']=set7[rating_columns].mean(axis=1)
set7['avg_score']=((set7[rating_columns]-set7[rating_columns].min())/(set7[rating_columns].max()-set7[rating_columns].min())).mean(axis=1)
set7[['essay_id', 'essay', 'avg_score']]
set7.hist(column='avg_score', bins=24, ax= plt.figure(figsize = (12,5)).gca())
set7.max()
# -
# ### Essay set 8
# +
set8 = train_data[train_data['essay_set'] == 8]
rating_columns = set8.columns.values[3:].tolist()
set8['avg_score']=set8[rating_columns].mean(axis=1)
set8['avg_score']=((set8[rating_columns]-set8[rating_columns].min())/(set8[rating_columns].max()-set8[rating_columns].min())).mean(axis=1)
set8[['essay_id', 'essay', 'avg_score']]
set8.hist(column='avg_score', bins=24, ax= plt.figure(figsize = (12,5)).gca())
set8.max()
| DataSorter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .ps1
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: .NET (PowerShell)
# language: PowerShell
# name: .net-powershell
# ---
# # T1546.005 - Event Triggered Execution: Trap
# Adversaries may establish persistence by executing malicious content triggered by an interrupt signal. The <code>trap</code> command allows programs and shells to specify commands that will be executed upon receiving interrupt signals. A common situation is a script allowing for graceful termination and handling of common keyboard interrupts like <code>ctrl+c</code> and <code>ctrl+d</code>.
#
# Adversaries can use this to register code to be executed when the shell encounters specific interrupts as a persistence mechanism. Trap commands are of the following format <code>trap 'command list' signals</code> where "command list" will be executed when "signals" are received.(Citation: Trap Manual)(Citation: Cyberciti Trap Statements)
# ## Atomic Tests
#Import the Module before running the tests.
# Checkout Jupyter Notebook at https://github.com/cyb3rbuff/TheAtomicPlaybook to run PS scripts.
Import-Module /Users/0x6c/AtomicRedTeam/atomics/invoke-atomicredteam/Invoke-AtomicRedTeam.psd1 - Force
# ### Atomic Test #1 - Trap
# After exiting the shell, the script will download and execute.
# After sending a keyboard interrupt (CTRL+C) the script will download and execute.
#
# **Supported Platforms:** macos, linux
# #### Attack Commands: Run with `sh`
# ```sh
# trap "nohup sh $PathToAtomicsFolder/T1546.005/src/echo-art-fish.sh | bash" EXIT
# exit
# trap "nohup sh $PathToAtomicsFolder/T1546.005/src/echo-art-fish.sh | bash" SIGINt
# ```
Invoke-AtomicTest T1546.005 -TestNumbers 1
# ## Detection
# Trap commands must be registered for the shell or programs, so they appear in files. Monitoring files for suspicious or overly broad trap commands can narrow down suspicious behavior during an investigation. Monitor for suspicious processes executed through trap interrupts.
| playbook/tactics/privilege-escalation/T1546.005.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.8 64-bit (''base'': conda)'
# language: python
# name: python3
# ---
# # Contrastive Learning For Normal Distribution
# The normal distribution, for a given mean and variance, specifies the probability density to be:
#
# $$f(x) = \frac{1}{\sigma \sqrt{2 \pi}} \cdot e^{-\frac{1}{2}(\frac{x - \mu}{\sigma})^2}$$
#
# The function is a normalized statistical model, meaning that the probability desnity funcion integrates to $1$ regardless of what values of $\sigma$ and $\pi$ are used. However, suppose instead we now we have
#
# $$f(x) = e^{-\frac{1}{2}(\frac{x - \mu}{\sigma})^2 + c}$$
#
# Now the function no longer itegrates to 1, which poses a problem for MLE estimation of the parameters. Below we use noise contrastive estimation to calculate $\mu, \sigma$, and $c$ for a distribution when we are only given samples of the distribution.
# +
import matplotlib as mpl
from matplotlib import cm
import matplotlib.pyplot as plt
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
torch.set_default_dtype(torch.float64)
import matplotlib.pyplot as plt
import matplotlib as mpl
from matplotlib import cm
from sys import exit
import scipy.integrate as integrate
from scipy import optimize
from torch.distributions.normal import Normal
from torch.distributions.uniform import Uniform
from torch.nn import functional as F
# +
true_dist = Normal(0.7, 1.5)
true_samples = true_dist.sample((100000,))
noise_dist = Normal(-1, 2)
noise_samples = noise_dist.sample((100000,))
# -
# Now we can plot the histograms of the samples to visual the distributions.
true_samples_np = true_samples.cpu().detach().numpy()
noise_samples_np = noise_samples.cpu().detach().numpy()
plt.hist(true_samples_np, density = True, bins = 100)
plt.show()
plt.hist(noise_samples_np, density = True, bins = 100)
plt.show()
# ## Learn Constants $\mu, \sigma, c$
# We have a sample of random vectors $x \in \mathbf{R}^d$ which follows an unknown probability density function $p_d(x)$. It is possible to model $p_d(x) = p_m(x; \theta)$ where $\theta$ is a vector of parameters. There exist some parameters $\theta'$ such that $p_d(x) = p_m(x;\theta')$.
#
# However, $\int_{}^{} p_m(x;\theta) dx = 1$.
#
# Even though the contraint can be fufilled in principle by redefining $$p_m(x; \theta) = \frac{p(x;\theta)}{\int_{}^{} p(x;\theta) dx}$$
#
# Here, $\int_{}^{} p(x;\theta) dx$ is called the normalizing constant in statistics or partition function in thermodynamics. In pratice, this integral is rarely tractable.
#
# Noise contrastive estimations defines an estimator that can be estimated by maximizing the objective function. For observed samples $X = (x_1, x_2, ..., x_T)$. We also have $Y = (y_1, y_2, ..., y_T)$ that come from a noise distribution $p_n(x)$. A note below is that we can denote the sigmoid function as $S(x) = \frac{1}{1+e^{-x}}$, such that $h(x; \theta) = S(G(x; \theta))$. Here, the objective function is:
#
# $$J_T(\theta) = \frac{1}{2T} \sum_{t=1}^{T} \ln[ h(x_t; \theta) ] + \ln[1-h(y_t; \theta)] $$
#
# $$h(x; \theta) = \frac{1}{1+e^{-G(x, \theta)}}$$
#
# $$G(x, \theta) = \ln p_m(x; \theta) - \ln p_n(x)$$
#
# Below we can code up all of the functions.
# +
def ln_p_n(x):
return noise_dist.log_prob(x)
def ln_p_m(x, theta):
mu, sigma, c = theta[0], theta[1], theta[2]
ln_p_m = -(x-mu)**2/(2*sigma**2) + c #Normal(mu, sigma).log_prob(x)
return ln_p_m #+ 0.5*(np.log(2*np.pi) + np.log(sigma)) - c
def G_x_theta(x, theta):
return ln_p_m(x, theta) - ln_p_n(x)
def h_x_theta(x, theta):
return torch.sigmoid(G_x_theta(x, theta))
def J_T(theta, X_true, Y_true):
T = X_true.size()[0] + Y_true.size()[0]
J_T = torch.sum(torch.log(h_x_theta(X_true, theta)) + torch.log(1 - h_x_theta(Y_true, theta)))
return -(1/(2*T))*J_T
#print(J_T([0, 1, 1], true_samples, noise_samples))
#print(J_T([0, 0.5, 1], true_samples, noise_samples))
print(optimize.minimize(J_T, x0=[2, 2, 2], args=(true_samples, noise_samples)))
print(np.log(1/(1.5*np.sqrt(2*np.pi))))
# -
# Above, we see that using NCE allows us to properly learn the values of $\mu, \sigma, c$.
# # Learning a General Potential Energy Function
# Above, we defined a normal distribtion to be of the form
#
# $$p(x) = e^{-\frac{(x-\mu)^2}{2\sigma^2}+c}$$
#
# Here, contrastive learning can find the parameters, $\mu, \sigma, c$. However, what if we instead have a neural network parameterise $U_\theta(x)$, such that we have $p(x) = e^{-U_\theta(x)}$?
# +
class NCE(nn.Module):
def __init__(self):
super(NCE, self).__init__()
self.U_x = nn.Sequential(
nn.Linear(1, 20),
nn.Tanh(),
nn.Linear(20, 100),
nn.Tanh(),
nn.Linear(100, 20),
nn.Tanh(),
nn.Linear(20, 1),
)
def forward(self, x):
return -self.U_x(x)
def ln_p_m(self, x): #this is U(x)
return -self.U_x(x)
def ln_p_n(self, x):
return noise_dist.log_prob(x)
def G_x_theta(self, x):
return self.ln_p_m(x) - self.ln_p_n(x)
def h_x_theta(self, x):
return torch.sigmoid(self.G_x_theta(x))
def loss(self, X_true, Y_true):
T = X_true.size()[0] + Y_true.size()[0]
J_T = torch.sum(torch.log(self.h_x_theta(X_true)) + torch.log(1 - self.h_x_theta(Y_true)))
return -(1/(2*T))*J_T
device = torch.device("cpu")
model = NCE().to(device)
optimizer = optim.Adam(model.parameters(), lr=1e-3)
# +
def train(epoch, true_samples, noise_samples):
model.train()
train_loss = 0
#true_samples = true_samples.to(device)
#noise_samples = noise_samples.to(device)
t = true_dist.sample((1000,1))
n = noise_dist.sample((1000,1))
optimizer.zero_grad()
loss = model.loss(t, n)
loss.backward()
train_loss += loss.item()
optimizer.step()
print(loss)
for epoch in range(1, 2000):
if epoch % 200 == 0:
print("Epoch", epoch)
train(epoch, true_samples, noise_samples)
# +
def true_U_x(x, mu, sigma, c):
return (-(x-mu)**2)/(2*sigma**2) + c
lines = np.linspace(-3.3, 4.7, 1000)
mu = 0.7
sigma = 1.5
c = -1.324403641312837
U_x_curve_true = [true_U_x(l, mu, sigma, c) for l in lines]
U_x_curve = [(model(torch.tensor(l).reshape(1,1))).detach().numpy()[0][0] for l in lines]
plt.plot(lines, U_x_curve_true)
plt.plot(lines, U_x_curve)
# +
def U_learned_x(x):
return np.exp((model(torch.tensor(x).reshape(1,1).to(device))).cpu().detach().numpy()[0][0])
integrate.quad(U_learned_x, -4, 4)[0]
| my_experiments/contrastive_learning_normal_distribtuion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="P9bJCDjdlgG6" colab_type="text"
# # **Spit some [tensor] flow**
#
# We need to learn the intricacies of tensorflow to master deep learning
#
# `Let's get this over with`
#
# + id="aQwc0re5mFld" colab_type="code" outputId="9cc81a4b-fffe-4508-9972-0388f5b6cfb7" colab={"base_uri": "https://localhost:8080/", "height": 35}
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import tensorflow as tf
print(tf.__version__)
# + [markdown] id="GMzqdmCYuI26" colab_type="text"
# ## Reference MachineLearningMastery.com
# + id="snz824BefAC-" colab_type="code" colab={}
from tensorflow.keras.layers import Input, Dense, Dropout, Flatten, Conv2D
from tensorflow.keras.models import Model
from tensorflow.keras.optimizers import SGD, Adam
from glob import glob
import sys, os
import cv2
# + id="qyw8HvOuBEZm" colab_type="code" outputId="fee9faec-4031-4989-c216-6fc724aaeb88" colab={"base_uri": "https://localhost:8080/", "height": 208}
# !wget https://www.theluxecafe.com/wp-content/uploads/2014/07/ferrari-spider-indian-theluxecafe.jpg
# + id="DXPaWBYPZlJj" colab_type="code" outputId="09ba0c8b-d922-4241-f076-5e66a06234ee" colab={"base_uri": "https://localhost:8080/", "height": 35}
# !ls
# + id="lpuwJ23ka6Ia" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 259} outputId="bb53ec9f-6db8-4655-a084-271dc1f66813"
X = cv2.imread('ferrari-spider-indian-theluxecafe.jpg')
X = cv2.cvtColor(X, cv2.COLOR_BGR2RGB)
plt.imshow(X)
# + id="l4oJJRlna--w" colab_type="code" outputId="3d8c3356-e95a-46f6-a0b2-01bc5ad573f1" colab={"base_uri": "https://localhost:8080/", "height": 35}
print(X.shape)
# + id="BTYoTUirr-EH" colab_type="code" colab={}
IMAGE_SIZE = X.shape
# + id="hriiTY7Frw34" colab_type="code" colab={}
X = np.expand_dims(X, axis=0)
# + id="hKbC-up0ru3O" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b3a02106-3f21-43a5-ce01-e6a16ec479ff"
print(X.shape)
# + id="E4OgFqA6ezQH" colab_type="code" outputId="cd5b63ca-da6a-4a2f-eff7-df76504307c7" colab={"base_uri": "https://localhost:8080/", "height": 35}
y = np.ndarray([1])
print(y.shape)
# + id="2PGHMmndhxfa" colab_type="code" colab={}
i_layer = Input(shape = IMAGE_SIZE)
h_layer = Conv2D(8, (3,3), strides = 1, activation='relu', padding='same')(i_layer)
h_layer = Flatten()(h_layer)
o_layer = Dense(1, activation='sigmoid')(h_layer)
model = Model(i_layer, o_layer)
# + id="rsDKDP41nW2Z" colab_type="code" outputId="766eca3e-8425-4b96-de35-018420ac5923" colab={"base_uri": "https://localhost:8080/", "height": 295}
model.summary()
# + id="bYip2sQaNiNI" colab_type="code" colab={}
model.compile(
optimizer='adam',
loss='binary_crossentropy',
metrics=['accuracy'])
# + id="tEsZnuikNwv3" colab_type="code" outputId="3bd5a414-1eda-4b83-99a2-a856e4b80e36" colab={"base_uri": "https://localhost:8080/", "height": 364}
report = model.fit(X, y, epochs = 10)
# + id="lfkBS6LttPXC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 87} outputId="10da7b96-6870-4ea2-b857-0d44de0df355"
model.layers
# + id="IaFsU6TEtWZs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="0bfa7783-d302-48d6-c0e1-23b9ada80962"
conv_layer = model.layers[1]
print(conv_layer)
# + id="ydxtsAxHtboo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="b8ae000f-c503-4b8c-b62d-49d39fb017f8"
filters, biases = conv_layer.get_weights()
print(conv_layer.name, filters.shape)
# + id="nywZL9y-t-cS" colab_type="code" colab={}
f_min, f_max = filters.min(), filters.max()
filters = (filters - f_min) / (f_max - f_min)
# + id="JodK4kinuFpg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 578} outputId="bae7b7f5-65f3-49ce-a7aa-56fd5267b41d"
plt.figure(figsize=(20,10))
n_filters, idx = 8, 1
for i in range(n_filters):
# get filter
f = filters[:, :, :, i]
for j in range(3):
ax = plt.subplot(n_filters, 3, idx)
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(f[:, :, j], cmap='gray')
idx += 1
plt.show()
# + id="-1fJo8_Zu4Rw" colab_type="code" colab={}
model_visual = Model(inputs=model.inputs, outputs=conv_layer.output)
# + id="IKrVb7QGvBAQ" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 225} outputId="eb0da9f7-3250-4d31-ec2c-693fc8b2683c"
model_visual.summary()
# + id="eUX4AkITvEIK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="7526fb19-595e-49c3-ccaa-2426f3af2e6d"
maps = model_visual(X)
print(maps.shape)
# + id="jQUsrdFGvD8f" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 294} outputId="1725ae27-4085-4acc-8652-2800b90e0309"
plt.figure(figsize=(20,10))
square = 4
idx = 1
for _ in range(square):
for _ in range(square):
if (idx > square * 2):
break
# specify subplot and turn of axis
ax = plt.subplot(square, square, idx)
ax.set_xticks([])
ax.set_yticks([])
plt.imshow(maps[0, :, :, idx-1], cmap='gray')
idx += 1
plt.show()
# + id="-43_M5dlzGAq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3b63c87f-83c1-4536-eeb6-4c33899eb58a"
maps.shape[3]
# + id="8AEYi9xCwyxR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ef26fc27-9b36-410d-c694-f4ca52c53dd7"
for i in range(maps.shape[3]):
ax = plt.subplot()
plt.imshow(maps[0, :, :, i], cmap='gray')
ax.set_xticks([])
ax.set_yticks([])
plt.show()
# + id="uqO39u3WzRCb" colab_type="code" colab={}
| Tensorflow_2X_Notebooks/Demo123_Convolution_Visualization.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### DECLARING |0> AND |1>
import numpy as np
q0=np.array([[1],[0]])
q1=np.array([[0],[1]])
print("\nq0=\n",q0 )
print("\nq1=\n",q1)
# ### DECLARING UNITARY MATRIX AND HERMITIAN MATRIX
X = np.array([[0,1],[1, 0]])
print("X = \n",X)
hermitianMatrix = np.conj(X).transpose()
unitaryMatrix = np.matmul(X, hermitianMatrix)
print("unitaryMatrix: XX* =\n ",unitaryMatrix)
print("hermitianMatrix: if A* = A i.e,\nA =\n",X,"\nand A* =\n",hermitianMatrix,"\ntherefore we can conclude A=A*, therefore A is a Hermitian Matrix.")
# ### INNER PRODUCT, OUTER PRODUCT AND TENSOR PRODUCT
print("INNER PRODUCT")
F = np.array([2, 1-2j])
V = np.array([[1, 2]])
print("Inner product of F and V is\n",np.dot(V,np.conj(F).transpose()))
print("Outer Product of F and V is\n",np.outer(F,V))
print("Tensor Product of \nV=\n", V,"\nand F =\n",F,"\nis =\n",np.tensordot(V,F,0))
# ### QUANTUM GATES: PAULI-X(NOT), PAULI-Z, HADAMARD, CNOT, TOFFOLI
print("PAULI-X Gate output when applied on state 0: \n",np.matmul(X, q0))
print("PAULI-X Gate output when applied on state 1: \n",np.matmul(X, q1))
print("PAULI Z-Gate")
Z = np.array([[1,0],[0, -1]])
print("When acted on state 0: \n|0>:\n",np.matmul(Z,q0))
print("When acted on state 1: \n|1>:\n",np.matmul(Z,q1))
print("Z- gate leaves |0 unchanged, and flips the sign of |1 to give −|1")
import math
print("HADAMARD GATE")
X1 = np.array([[1,0],[0,1]])
H = np.dot(1/math.sqrt(2),np.array([[1,1],[1,-1]]))
print("Output of the hadamard gate when applied to |0>:\n",np.matmul(H,q0))
print("Output of the hadamard gate when applied to |1>:\n",np.matmul(H,q1))
print("CNOT GATE")
q00 = np.tensordot(q0,q0,0)
q01 = np.tensordot(q0,q1,0)
q10 = np.tensordot(q1,q0,0)
q11 = np.tensordot(q1,q1,0)
cnot = np.array([[1,0,0,0],[0,1,0,0],[0,0,0,1],[0,0,1,0]])
q10 = np.tensordot(q1,q0,0)
cnot.resize(4,4)
q10.resize(4,1)
q11.resize(4,1)
q00.resize(4,1)
q01.resize(4,1)
print("CNOT |00> =\n", np.matmul(cnot,q00))
print("CNOT |01> =\n",np.matmul(cnot,q01))
print("CNOT |10> =\n",np.matmul(cnot,q10))
print("CNOT |11> =\n",np.matmul(cnot,q11))
q000=np.tensordot(q00,q0,0)
q001=np.tensordot(q00,q1,0)
q010=np.tensordot(q01,q0,0)
q011=np.tensordot(q01,q1,0)
q100=np.tensordot(q10,q0,0)
q101=np.tensordot(q10,q1,0)
q110=np.tensordot(q11,q0,0)
q111=np.tensordot(q11,q1,0)
Toffoli = np.array([
[1,0,0,0,0,0,0,0],
[0,1,0,0,0,0,0,0],
[0,0,1,0,0,0,0,0],
[0,0,0,1,0,0,0,0],
[0,0,0,0,1,0,0,0],
[0,0,0,0,0,1,0,0],
[0,0,0,0,0,0,0,1],
[0,0,0,0,0,0,1,0]])
q000.resize(8,1)
q001.resize(8,1)
q010.resize(8,1)
q011.resize(8,1)
q100.resize(8,1)
q101.resize(8,1)
q110.resize(8,1)
q111.resize(8,1)
a = np.array([q000,q001,q010,q011,q100,q101,q110,q111])
av = np.array(["000","001","010","011","100","101","110","111"])
for i in range(0,8):
print("Toffoli |",av[i],"> =\n",np.matmul(Toffoli,a[i]))
| QUBITOPERATIONSANDGATES.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
x=np.array([1,2,3])
x
y=x*3+1
y
def GD(x,y,epoch):
w=1
rate=0.003
b=1
for i in range(epoch):
loss=y-(w*x+b)
wd=-(2/len(x))*sum(x*(loss))
bd=-(2/len(x))*sum(loss)
w-=wd*rate
b-=bd*rate
if i%(100)==0:
print('epoch : ',i,'\tw : ',w,'\t\tb : ',b,'\t\tloss : ',sum(loss**2))
return w,b
w,b=GD(x,y,3000)
print(w*x+b)
| gradient_descent.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/oonid/growth-hacking-with-nlp-sentiment-analysis/blob/master/create_dictionary_based_sentiment_analyzer.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="fv0gaMFmHwra" colab_type="text"
# # Dictionary Based Sentiment Analyzer
#
# * Word tokenization
# * Sentence tokenization
# * Scoring of the reviews
# * Comparison of the scores with the reviews in plots
# * Measuring the distribution
# * Handling negation
# * Adjusting your dictionary-based sentiment analyzer
# * Checking your results
# + id="yo6ViHkBKc6x" colab_type="code" outputId="d670ca5e-a91a-4263-d686-02c4bd63baef" colab={"base_uri": "https://localhost:8080/", "height": 107}
# all imports and related
# %matplotlib inline
import pandas as pd
import numpy as np
import altair as alt
from nltk import download as nltk_download
from nltk.tokenize import word_tokenize, sent_tokenize
from nltk.sentiment.util import mark_negation
nltk_download('punkt') # required by word_tokenize
from collections import Counter
# + [markdown] id="81cswx9RIFFt" colab_type="text"
# ### load the small_corpus CSV
#
# run process from
# [create_dataset.ipynb](https://github.com/oonid/growth-hacking-with-nlp-sentiment-analysis/blob/master/create_dataset.ipynb)
#
# # # copy file **small_corpus.csv** to this Google Colab Files (via file upload or mount drive).
#
# + id="LxyLiywrHXBE" colab_type="code" outputId="b9630401-04ea-438b-f406-40d0b484ea93" colab={"base_uri": "https://localhost:8080/", "height": 415}
df = pd.read_csv('small_corpus.csv')
df
# + id="PFxrdXW-OLvp" colab_type="code" outputId="8b801f8b-79ac-401a-ada1-6d141934fe12" colab={"base_uri": "https://localhost:8080/", "height": 69}
# check if any columns has null, and yes the reviews column has
df.isnull().any()
# + id="BEXN07u-Osnm" colab_type="code" outputId="afb85f37-2ae5-4c00-ab10-99a9ce1575ca" colab={"base_uri": "https://localhost:8080/", "height": 69}
# repair null in column reviews with empty string ''
df.reviews = df.reviews.fillna('')
# test again
df.isnull().any()
# + id="M9c65mzkHDDE" colab_type="code" outputId="834024a0-cbc8-4c75-eaa4-c21132ddb4d3" colab={"base_uri": "https://localhost:8080/", "height": 575}
rating_list = list(df['ratings'])
review_list = list(df['reviews'])
print(rating_list[:5])
for r in review_list[:5]:
print('--\n{}'.format(r))
# + [markdown] id="47i2oNUgQU-F" colab_type="text"
# ### tokenize the sentences and words of the reviews
# + id="lufWRnRULCbQ" colab_type="code" outputId="22a04d20-28b6-4240-bada-4577b17c6d36" colab={"base_uri": "https://localhost:8080/", "height": 225}
word_tokenized = df['reviews'].apply(word_tokenize)
word_tokenized
# + id="KWe_2rN2LmmK" colab_type="code" outputId="f11a386e-bf0f-4de4-8ace-ec101201d160" colab={"base_uri": "https://localhost:8080/", "height": 225}
sent_tokenized = df['reviews'].apply(sent_tokenize)
sent_tokenized
# + [markdown] id="9J4INBoDQgIi" colab_type="text"
# ### download the opinion lexicon of NLTK
#
# use it with reference to it source:
#
# https://www.nltk.org/_modules/nltk/corpus/reader/opinion_lexicon.html
#
#
# + id="B5a72gM7Pl6N" colab_type="code" outputId="da5efd57-7787-4bc1-96ab-f0e8bfbd4295" colab={"base_uri": "https://localhost:8080/", "height": 52}
# imports and related
nltk_download('opinion_lexicon')
from nltk.corpus import opinion_lexicon
# + id="aE_AwlMwQxB6" colab_type="code" outputId="7976a0ec-1367-4a89-f888-99b09cb496d9" colab={"base_uri": "https://localhost:8080/", "height": 193}
print('total lexicon words: {}'.format(len(opinion_lexicon.words())))
print('total lexicon negatives: {}'.format(len(opinion_lexicon.negative())))
print('total lexicon positives: {}'.format(len(opinion_lexicon.positive())))
print('sample of lexicon words (first 10, by id):')
print(opinion_lexicon.words()[:10]) # print first 10 sorted by file id
print('sample of lexicon words (first 10, by alphabet):')
print(sorted(opinion_lexicon.words())[:10]) # print first 10 sorted alphabet
positive_set = set(opinion_lexicon.positive())
negative_set = set(opinion_lexicon.negative())
print(len(positive_set))
print(len(negative_set))
# + id="3X92TU6UGb6Y" colab_type="code" outputId="c4ce426a-6769-415e-e38a-7b4a7f092fc8" colab={"base_uri": "https://localhost:8080/", "height": 104}
def simple_opinion_test(words):
if words not in opinion_lexicon.words():
print('{} not covered on opinion_lexicon'.format(words))
else:
if words in opinion_lexicon.negative():
print('{} is negative'.format(words))
if words in opinion_lexicon.positive():
print('{} is positive'.format(words))
simple_opinion_test('awful')
simple_opinion_test('beautiful')
simple_opinion_test('useless')
simple_opinion_test('Great') # must be lower case
simple_opinion_test('warming')
# + [markdown] id="eZlGLTnEQRjX" colab_type="text"
# ### classify each review in a scale of -1 to +1
# + id="SNP52NhPKqGH" colab_type="code" colab={}
# the process to score review:
# * tokenize review (from multiple sentence) become sentences
# * so sentence score will be build from it words
def score_sentence(sentence):
"""sentence (input) are words that tokenize from sentence.
return score between -1 and 1
if the total positive greater than total negative then return 0 to 1
if the total negative greater than total positive then return -1 to 0
"""
# opinion lexicon not contains any symbol character, and must be set lower
selective_words = [w.lower() for w in sentence if w.isalnum()]
total_selective_words = len(selective_words)
# count total words that categorized as positive from opinion lexicon
total_positive = len([w for w in selective_words if w in positive_set])
# count total words that categorized as negative from opinion lexicon
total_negative = len([w for w in selective_words if w in negative_set])
if total_selective_words > 0: # has at least 1 word to categorize
return (total_positive - total_negative) / total_selective_words
else: # no selective words
return 0
def score_review(review):
"""review (input) is single review, could be multiple sentences.
tokenize review become sentences.
tokenize sentence become words.
collect sentence scores as list, called sentiment scores.
score of review = sum of all sentence scores / total of all sentence scores
return score of review
"""
sentiment_scores = []
sentences = sent_tokenize(review)
# process per sentence
for sentence in sentences:
# tokenize sentence become words
words = word_tokenize(sentence)
# calculate score per sentence, passing tokenized words as input
sentence_score = score_sentence(words)
# add to list of sentiment scores
sentiment_scores.append(sentence_score)
# mean value = sum of all sentiment scores / total of sentiment scores
if sentiment_scores: # has at least 1 sentence score
return sum(sentiment_scores) / len(sentiment_scores)
else: # return 0 if no sentiment_scores, avoid division by zero
return 0
# + id="TDLK8lhORwxI" colab_type="code" outputId="80ab5e55-b2bd-403c-e2b5-a8082ca30e1c" colab={"base_uri": "https://localhost:8080/", "height": 35}
review_sentiments = [score_review(r) for r in review_list]
print(review_sentiments[:5])
# + id="LEJRmJhJWitq" colab_type="code" outputId="a8e45411-75ca-444b-d634-63b6350dd21d" colab={"base_uri": "https://localhost:8080/", "height": 592}
print(rating_list[:5])
print(review_sentiments[:5])
for r in review_list[:5]:
print('--\n{}'.format(r))
# + id="fkDNwlnrXLWE" colab_type="code" outputId="a9d3d16e-3375-4629-d117-1c2dbb7216ff" colab={"base_uri": "https://localhost:8080/", "height": 415}
df = pd.DataFrame({
"rating": rating_list,
"review": review_list,
"review dictionary based sentiment": review_sentiments,
})
df
# + id="THjjD3F8XjI_" colab_type="code" colab={}
df.to_csv('dictionary_based_sentiment.csv', index=False)
# + [markdown] id="NjOPECvRhWNp" colab_type="text"
# # Compare the scores of the product reviews with the product ratings using a plot
# + id="yKMJV8K3YUxV" colab_type="code" outputId="a6ea62a0-3ff5-41bf-ef28-0e2640035779" colab={"base_uri": "https://localhost:8080/", "height": 35}
rating_counts = Counter(rating_list)
print('distribution of rating as dictionary: {}'.format(rating_counts))
# + [markdown] id="rw3_s1u5h6vk" colab_type="text"
# ### a plot of the distribution of the ratings
# + id="ds84rsVuaUOY" colab_type="code" outputId="02e4c5c7-08d7-4791-fbf4-726bc733f501" colab={"base_uri": "https://localhost:8080/", "height": 202}
# ratings as str will be different with ratings as int from keys()
dfrc = pd.DataFrame({
"ratings": [str(k) for k in rating_counts.keys()],
"counts": list(rating_counts.values())
})
dfrc
# + id="GR6Sy7iqbNw4" colab_type="code" outputId="4da7dc5d-957e-48ed-eb28-be636b9259f1" colab={"base_uri": "https://localhost:8080/", "height": 364}
rating_counts_chart = alt.Chart(dfrc).mark_bar().encode(x="ratings", y="counts")
rating_counts_chart
# + [markdown] id="BR8nkkyLiBPN" colab_type="text"
# ### a plot of the distribution of the sentiment scores
# + id="MVcar_wGbeu_" colab_type="code" outputId="1dee8837-2147-4823-cfe0-a2b16ae6785b" colab={"base_uri": "https://localhost:8080/", "height": 159}
# get histogram value
# with the value of the probability density function at the bin,
# normalized such that the integral over the range is 1
hist, bin_edges = np.histogram(review_sentiments, density=True)
print('histogram value: {}'.format(hist))
print('bin_edges value: {}'.format(bin_edges)) # from -1 to 1
print()
labels = [(str(l[0]), str(l[1])) for l in zip(bin_edges, bin_edges[1:])]
print('labels: {}'.format(labels))
labels = [" ".join(label) for label in labels]
print('labels: {}'.format(labels))
# + id="LZgUfH6edTxk" colab_type="code" outputId="8a8b5aa1-6a00-42ad-f992-a5177e2c082a" colab={"base_uri": "https://localhost:8080/", "height": 355}
dfsc = pd.DataFrame({
"sentiment scores": labels,
"counts": hist,
})
dfsc
# + id="efGd6CaCeues" colab_type="code" outputId="1bf5cf41-31e6-452e-c84b-4e8519dc37d9" colab={"base_uri": "https://localhost:8080/", "height": 536}
# sentiment_counts_chart = alt.Chart(dfsc).mark_bar() \
# .encode(x="sentiment scores", y="counts")
sentiment_counts_chart = alt.Chart(dfsc).mark_bar() \
.encode(x=alt.X("sentiment scores", sort=labels), y="counts")
sentiment_counts_chart
# + [markdown] id="DnyGwN9MiPYm" colab_type="text"
# ### a plot about the relation of the sentiment scores and product ratings
# + id="yDujLuF7fPBq" colab_type="code" outputId="74bf1dc3-169d-44a4-f51c-0a82787fb004" colab={"base_uri": "https://localhost:8080/", "height": 415}
# explore if there's relationship between ratings and sentiments
dfrs = pd.DataFrame({
"ratings": [str(r) for r in rating_list],
"sentiments": review_sentiments,
})
dfrs
# + id="X_wMFf9kgI9y" colab_type="code" outputId="0282a799-49ba-4dda-99e5-0a8168de9837" colab={"base_uri": "https://localhost:8080/", "height": 364}
rating_sentiments_chart = alt.Chart(dfrs).mark_bar()\
.encode(x="ratings", y="sentiments", color="ratings", \
tooltip=["ratings", "sentiments"])\
.interactive()
rating_sentiments_chart
# + [markdown] id="QS2n43Hc7HWX" colab_type="text"
# # Measure the correlation of the sentiment scores and product ratings
#
# article from [machinelearningmastery](https://machinelearningmastery.com/how-to-use-correlation-to-understand-the-relationship-between-variables/) about how to use correlation to understand the relationship between variable.
#
# * Covariance. Variables can be related by a linear relationship.
# * Pearson's Correlation. Pearson correlation coefficient can be used to summarize the strength of the linear relationship between two data samples.
# * Spearman's Correlation. Two variables may be related by a non-linear relationship, such that the relationship is stronger or weaker across the distribution of the variables.
#
# import pearsonr and spearmanr from package scipy.stats
#
# + id="Rs470PVsgXxY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="4b09da10-521e-46b3-9ec0-07aa6052b80c"
from scipy.stats import pearsonr, spearmanr
pearson_correlation, _ = pearsonr(rating_list, review_sentiments)
print('pearson correlation: {}'.format(pearson_correlation))
spearman_correlation, _ = spearmanr(rating_list, review_sentiments)
print('spearman correlation: {}'.format(spearman_correlation))
# Spearman rank correlation value said that there's weak correlation
# between rating and review score (sentiments)
# + [markdown] id="y4kRPqiP_wuW" colab_type="text"
# # Improve your sentiment analyzer in order to reduce contradictory cases
# + [markdown] id="cjzP6L3L_0y6" colab_type="text"
# ### need to handle negation, since mostly those cases are contradictory when there is negation in the sentence (e.g., no problem)
# + id="miokDL3M93cP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 277} outputId="5a2c9f02-dbac-493e-c985-7f245360e6cb"
for idx, review in enumerate(review_list):
r = rating_list[idx]
s = review_sentiments[idx]
if r == 5 and s < -0.2:
# rating 5 but sentiment negative below -0.2
print('({}, {}): {}'.format(r, s, review))
if r == 1 and s > 0.3:
# rating 1 but got sentiment positive more than 0.3
print('({}, {}): {}'.format(r, s, review))
# + [markdown] id="G_E9iKIeBvVx" colab_type="text"
# ### use the mark_negation function to handle negation
# + id="s1rzdrEiBW05" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 89} outputId="baf504d7-61db-426f-a7b9-aa797e38e95b"
test_sentence = 'Does not work correctly with xbox'
print(mark_negation(test_sentence.split()))
# not detected on "No problems."
test_sentence = 'Would buy again. No problems.'
print(mark_negation(test_sentence.split()))
# sentence from sample solution works to detect "no problems."
test_sentence = "I received these on time and no problems. No damages battlfield never fails"
print(mark_negation(test_sentence.split()))
# + id="ugEPQQ6JCSyQ" colab_type="code" colab={}
| create_dictionary_based_sentiment_analyzer.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **Imports:**
# +
import numpy as np
import matplotlib.pyplot as plt
plt.style.use('seaborn-whitegrid')
prop_cycle = plt.rcParams['axes.prop_cycle']
colors = prop_cycle.by_key()['color']
import ipywidgets as widgets
# -
# # Plotting function
# $$
# \mathcal{B}(p_1,p_2,t,\overline{x}_1,m)=\left\{ (x_{1},x_{2})\in\mathbb{R}_{+}^{2}\,|\,\begin{cases}
# p_{1}x_{1}+p_{2}x_{2}=m & \text{hvis }x_{1}\leq\overline{x}_{1}\\
# p_{1}\overline{x}_{1}+(p_{1}+t)(x_{1}-\overline{x}_{1})+p_{2}x_{2}=m & \text{hvis }x_{1}>\overline{x}_{1}
# \end{cases}\right\}
# $$
def plot_budgetset(p1,p2,t,x1bar,m,x1_max=10,x2_max=10,name=None):
# a. figure
fig = plt.figure(figsize=(6,6),dpi=100)
ax = fig.add_subplot(1,1,1)
# fill
x = [0,0,x1bar,x1bar+(m-p1*x1bar)/(p1+t)]
y = [0,m/p2,(m-p1*x1bar)/p2,0]
ax.fill(x,y,alpha=0.80)
# line
x = [0,x1bar,x1bar+(m-p1*x1bar)/(p1+t)]
y = [m/p2,(m-p1*x1bar)/p2,0]
ax.plot(x,y,color='black')
# details
ax.set_xlabel('$x_1$')
ax.set_ylabel('$x_2$')
ax.set_xlim([0,x1_max])
ax.set_ylim([0,x2_max])
# save
if not name is None: fig.savefig(name)
# # Static
plot_budgetset(p1=1.0,p2=1.0,t=1.0,x1bar=2.0,m=6.0)
# # Interactive
widgets.interact(lambda p1,p2,t,x1bar,m: plot_budgetset(p1=p1,p2=p2,t=t,x1bar=x1bar,m=m),
p1=widgets.FloatSlider(description=r'p1',min=0.1,max=5.0,step=0.05,value=1.0),
p2=widgets.FloatSlider(description=r'p2',min=0.1,max=5.0,step=0.05,value=1.0),
t=widgets.FloatSlider(description=r't',min=-1.0,max=5.0,step=0.05,value=1.0),
x1bar=widgets.FloatSlider(description=r'x1bar',min=0.1,max=5.0,step=0.05,value=2.0),
m=widgets.FloatSlider(description=r'm',min=0.1, max=10.0,step=0.05,value=6.0)
);
| Lec2_BudgetSetKink.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Gaussian Mixture Model(Application)
#
# 在写代码之前我们需要注意一下几个点:
#
# - [多元正态分布](https://zh.wikipedia.org/wiki/%E5%A4%9A%E5%85%83%E6%AD%A3%E6%80%81%E5%88%86%E5%B8%83)
#
# - 在理论上多元正态分布的参数$\Sigma$是需要半正定和[非奇异](https://www.zhihu.com/question/35318893)的,但是由于EM是迭代算法,所以在迭代的过程中可能会产生一些奇异的矩阵,所以我们在使用[multivariate_normal](https://docs.scipy.org/doc/numpy-1.15.1/reference/generated/numpy.random.multivariate_normal.html)计算多元正态分布的时候我们需要调节参数
# - allow_singular=True
# - 参数$\Sigma$必须要是[半正定](https://blog.csdn.net/you1314520me/article/details/78856322#_63)的.
# ### 1 Import package
import numpy as np
import pandas as pd
from sklearn.datasets import load_iris
import matplotlib.pyplot as plt
from sklearn import mixture
from scipy.stats import multivariate_normal
# ### 2 Load Data
#
# 这里的数据集使用的是来自于sklearn的鸢尾花数据集数据集的futures我们选择:
# - sepal length,sepal width
#
# 另外由于GMM是非监督学习,在训练过程中我们不需要使用label,但是我们依然要拿出label以方便于后面正确率的比较
def create_data():
iris = load_iris()
df = pd.DataFrame(iris.data, columns=iris.feature_names)
df['label'] = iris.target
df.columns = ['sepal length', 'sepal width', 'petal length', 'petal width', 'label']
data = np.array(df.iloc[:100, [0, 1, -1]])
return data[:,:2], data[:,-1]
X, y = create_data()
# 绘制源数据样本
plt.scatter(X[:,0],X[:,1],c=y)
# 可以看出鸢尾花数据集有两类,一类是上方的紫色,一类是下方的黄色,所以我们如果要对此数据集进行GMM训练,那么C是2.
#
# 其中C代表高斯模型的数量
# ### 2 各个参数的维度
#
#
# 如果存在C个高斯的混合模型,且源数据X的形状是(m,d):
# - m: Number of sample,d:Dimension or features.
#
#
# $\alpha_l$: 每一个高斯模型都含有一个权重$\alpha$,所以$\alpha$的形状是(C,),初始值我们一般都选择$\frac{1}{C}$是比较合理的.
#
# $\mu_l$: 每一个高斯模型都含有一个$\mu$,且在多元正态分布中单个样本的各个特征都有一个对应的$\mu_i$,所以$\mu$的形状是(C,d)
#
# $\Sigma_l$: 每一个高斯模型都含有一个协方差矩阵$\Sigma$,且每一个协方差矩阵中的形状是(d,d),所以$\Sigma$的形状是(C,d,d)
#
# $p(Z|X,\theta) = \gamma$:每一个高斯模型都能够给予一个样本点在自身高斯模型下的概率,所以$\gamma$的形状应该是(m,C)
# d:查看特征数量
d = X.shape[1]
# 定义初始化的$\alpha$:
#
# $\alpha=\frac{1}{C}$
C = 2
alpha = np.ones(C,dtype=np.float32) * (1.0 / C)
alpha
# 初始化的$\mu$:
#
# shape of mu = (C,d)
#
# 这里初始化的值最好选用服从standard normal的数值
mu = np.random.randn(C,d)
mu
# 初始化$\Sigma$:
#
# 在初始化协方差$\Sigma$的时候我们需要先建立三维数组(C,d,d)以方便后面代码中的循环,保证每次循环出来的是满足非正定的方正(对称矩阵)
#
# 
#
#
Sigma_ = np.random.randn(C,d,d)
Sigma_
Sigma = np.array([np.dot(A,A.T) for A in Sigma_])
Sigma
# ### 3 Model
#
# #### 3.1 建立初始化函数,里面包括了$\alpha,\mu,\Sigma$的初始化方式
def initial_parameters(C,d):
"""
initialization parameters.
Parameters:
----------
C: Number of Gaussian model.
d:dimension
Returns:
-------
parameter of Gaussian model.
alpha: shape = (C,)
mu:shape=(C,d)
Sigma:(C,d,d)
"""
np.random.seed(1)
alpha = np.ones(C,dtype=np.float32) * (1.0 / C)
mu = np.random.randn(C,d)
Sigma_ = np.random.randn(C,d,d)
Sigma = np.array([np.dot(A,A.T) for A in Sigma_])
return alpha,mu,Sigma
d = X.shape[1]
C = 2
alpha,mu,Sigma = initial_parameters(C=C,d = d)
print('alpha:{}\nmu:{},\nSigma:{}'.format(alpha,mu,Sigma))
# #### 3.2 Start E-step
#
# E-step:
# - 我们需要直接从后验概率$P(z|x,\theta)$计算
#
# $P(z_{ik}|x_i,\theta_k)=\frac{\alpha_k N(x_i|\theta_k)}{\sum_{k}^{K}\alpha_k N(x_i|\theta_k)}$
#
# **Ps:**
# - 使用multivariate_normal.pdf来计算多元正态的值
# - allow_singular=True:允许奇异矩阵
# - gamma:响应度,也就是说它能给出某一个样本点属于某一个高斯模型的概率
# +
def E_step(X,alpha,mu,Sigma):
"""
Implementation E step.
The gamma equal posterior probability.
Parameters:
----------
X: input data, training set.
parameters of Gaussian model:
1.alpha
2.mu
3.Sigma
Return:
------
gamma:response probability.
"""
m,d = X.shape
C = alpha.shape[0]
gamma = np.zeros(shape=(m,C))
for i in range(C):
gamma[:,i] = multivariate_normal.pdf(X,mean=mu[i],cov=Sigma[i],allow_singular=True)
gamma *= alpha[i]
gamma = gamma / np.sum(gamma,axis=1,keepdims= True)
return gamma
# -
gamma = E_step(X,alpha,mu,Sigma)
print('gamma:{}'.format(gamma[:10]))
# 可以看出在测试迭代中,gamma的第一行代表第一个样本属于两个高斯的概率:
#
# 可以看出第一个样本属于第一个高斯的概率为1,这不一定是正确的,因为只是初次迭代.
# #### 3.3 M-step:
#
# M-step负责更新参数,需要注意的是:
#
# 在更新$\Sigma$的时候需要使用外积[np.outer](https://docs.scipy.org/doc/numpy-1.14.0/reference/generated/numpy.outer.html)计算,这样的出来的结果才是对称的,另外不能保证$\Sigma$非奇异,原因开始的时候已经说明.
#
# 在取出gamma中的值计算的时候,由于Numpy的影响,我们需要增加一个维度np.newaxis来使得两个向量可以计算.
#
# $\alpha=\frac{\sum_{i=1}^{N}P(z_{ik}|x_i,\theta_k)}{N}$
#
# $\mu=\frac{\sum_{i=1}^{N}P(z_{ik}|x_i,\theta_k)x_i}{\sum_{i=1}^{N}P(z_{ik}|x_i,\theta_k)}$
#
# $\Sigma=\frac{\sum_{i=1}^{N}[x_i-\mu_k][x_i-\mu_k]^T P(z_{ik}|x_i,\theta_k)}{\sum_{i=1}^{N}P(z_{ik}|x_i,\theta_k)}$
def M_step(X,gamma):
"""
Update parameters.
Parameters:
---------
X: training set.
gamma: response probability.
Return:
------
parameters of Gaussian mixture model.
1.alpha
2.mu
3.Sigma
"""
m,d = X.shape
C = gamma.shape[1]
alpha = np.zeros(C)
mu = np.zeros((C,d))
Sigma = np.zeros((C,d,d))
for c in range(C):
# gamma[:,c][:,np.newaxis]: add axis in gamma[:,c]'s column
alpha[c] = np.sum(gamma[:,c]) / m
mu[c,:]= np.sum(gamma[:,c][:,np.newaxis] * X,axis=0) / np.sum(gamma[:,c])
# Sigma:singular matrix maybe.
Sigma[c,:] = np.sum([gamma[i,c] * np.outer(X[i] - mu[c], X[i] - mu[c]) for i in range(m)], axis=0) / np.sum(gamma[:,c])
return alpha,mu,Sigma
alpha,mu,Sigma = M_step(X,gamma)
print('alpha:{}\nmu:{},\nSigma:{}'.format(alpha,mu,Sigma))
# #### 3.4 Build GMM
#
# 现在可以开始建立模型GMM
#
# 这里我们会设置一个threshold,来使得在迭代过程中如果满足threshold,则退出迭代.因为在threshold内如果保证了gamma的收敛,我们就没有必要继续迭代了.
def GMM(X,C,iter_,threshold):
"""
Build Gaussian Mixture Model.
Parameters:
----------
X: training data (m,d)
iter_: number of iteration.
threshold: threshold value,stop condition.
Return:
------
The best parameters:
1.alpha
2.mu
3.Sigma
"""
d = X.shape[1]
alpha,mu,Sigma = initial_parameters(C=C,d = d)
for Iter in range(iter_):
Sigma_prev = Sigma
gamma = E_step(X,alpha,mu,Sigma)
alpha,mu,Sigma = M_step(X,gamma)
if (Sigma - Sigma_prev).all() <= threshold: # stop condition.
break
return alpha,mu,Sigma
best_alpha,best_mu,best_Sigma = GMM(X,2,100,1e-3)
print(best_alpha)
print(best_mu)
print(best_Sigma)
# 现在,已经得到了当前最优的参数模型,那么我们将其带入E-step中求出后验概率,并且后验概率中每一组预测值中最大的索引作为分类标签.
gamma = E_step(X,alpha=best_alpha,mu = best_mu,Sigma=best_Sigma)
labels = gamma.argmax(axis=1) # axis=1,按照列之间来比较,取出概率最大的一个应对于某一个样本点属于某一个高斯.
plt.scatter(X[:,0],X[:,1],c=labels)
plt.title('Predict Result.')
plt.show()
# 可以看出预测的结果是可以的,只有貌似左下角的点分类错误
#
# 对于这种简单的样本GMM模型是可以的,由于是非监督算法,那么正确率是肯定会低于监督算法的.
# ### 4. plot Gaussian Mixture Model
#
# 下面我们来动态的看看整个训练过程中GMM的变化.
#
# 在Jupyter中,我们可以使用:
# - %matplotlib inline
# - from IPython import display
# - display.clear_output(wait=True)
#
# 来绘制动态过程.
#
# 在绘制过程中,我们需要使用[clabel](https://matplotlib.org/gallery/images_contours_and_fields/contour_demo.html#sphx-glr-gallery-images-contours-and-fields-contour-demo-py)来绘制等高线.
def plotGaussian(X,mu,Sigma):
# %matplotlib inline
from IPython import display
# meshgraid
x = np.linspace(0., 8.,num=100)
y = np.linspace(0., 5.,num=100)
xx, yy = np.meshgrid(x, y)
meshgrid_X = np.array([xx.ravel(), yy.ravel()]).T
fig, ax = plt.subplots()
C = X.shape[1]
# plot two Gaussian model.
for c in range(C):
Z = multivariate_normal.pdf(meshgrid_X,mean=mu[c],cov=Sigma[c],allow_singular=True)
Z = Z.reshape(xx.shape)
CS = ax.contour(xx, yy, Z)
ax.clabel(CS, inline=False, fontsize=False)
ax.scatter(X[:,0],X[:,1],c=labels)
plt.xlim(X[:,0].min(),X[:,0].max())
plt.ylim(X[:,1].min(),X[:,1].max())
plt.xticks(()) # hide x,y ticks
plt.yticks(())
plt.title('Gaussian Mixture Model')
plt.pause(0.1)
plt.show()
display.clear_output(wait=True)
plotGaussian(X,best_mu,best_Sigma)
# #### 4.1 Gaussian fitting animation
def Gaussian_Fitting_Animation(X,C,iter_,threshold):
d = X.shape[1]
alpha,mu,Sigma = initial_parameters(C=C,d = d)
for Iter in range(iter_):
Sigma_prev = Sigma
gamma = E_step(X,alpha,mu,Sigma)
alpha,mu,Sigma = M_step(X,gamma)
plotGaussian(X,mu=mu,Sigma=Sigma)
if (np.abs((Sigma - Sigma_prev))<=threshold).all(): # 迭代停止条件
break
Gaussian_Fitting_Animation(X=X,C=2,iter_=100,threshold=1e-3)
# ### 5 Scikit-learn
#
# 我们也可以使用scikit-learn来快速实现GMM
# +
from sklearn import mixture
gmm = mixture.GaussianMixture(n_components=2,covariance_type='full')
gmm.fit(X)
labels = gmm.predict(X)
plt.scatter(X[:,0],X[:,1],c=labels)
plt.title('Predict Result.')
plt.show()
# -
# 可以看出结果与我们自身编写的GMM是类似的.
# ### 6. Summary
#
# - 在编写GMM的过程中,很显然理论上的方法和代码中可能不一样,典型的就是$\Sigma$参数.
# - 这里没有定义损失函数.损失函数有很多有[常见的分类和回归的损失函数](https://www.cnblogs.com/massquantity/p/8964029.html)你也可以自己选择正确的使用,当然你也可以定义之前我们的损失函数KL.
# ### Homework:
#
# 将data_set文件中的GMM_data数据集使用GMM模型进行拟合.
#
# Good Luck~~
| 7-3Gaussian Mixture Model(Application).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 目的
# - 正味重量分布を推定する
# - 独立に推定するのではなく、種類を合わせてモデルを立てる
# +
import os, sys
import numpy as np
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
import pymc3 as pm
import ChocoUtils as cu
# -
sns.set()
# ## データの取得と確認
data = cu.get_data(db_file='../../data/choco-ball.db', filter_str="taste in (0, 1, 4, 11, 12)")
grp = data.groupby(['taste'])
print(grp.count()[['net_weight']])
# 仕様の重量
spec = {0:28.0, 1:25.0, 4:22.0, 7:26.0, 10:22.0, 11:22.0, 12:22.0}
taste_name = {0:"peanut", 1:"strawberry", 4:"ama-zake",
7:"pachi-pachi", 10:"pineapple",
11:"milk-caramel", 12:"choco-banana"}
# +
fig = plt.figure(figsize=(16, 4))
ax = fig.subplots(1, 2)
bins = np.linspace(0, 5.0, 30)
for key, value in grp:
sns.distplot(value["net_weight"], bins=np.linspace(21, 33, 50), hist=True, label=taste_name[key], ax=ax[0])
sns.distplot((value["net_weight"] - spec[key]), bins=bins, hist=True, label=taste_name[key], ax=ax[1])
ax[0].set_xlabel("NetWeight [g]")
ax[1].set_xlabel("(NetWeight - Spec) [g]")
ax[0].legend()
ax[1].legend()
fig.savefig("weight_histogram.png")
# -
# ## 重量パラメータの推定
# - 重量は正規分布に従うと仮定 ←この仮定はあとで見直し
# - 独立にパラメータを予測した場合と、階層モデルで予測した場合を比較する
taste = data['taste'].values
taste_idx = pd.Categorical(data['taste']).codes
spec_lst = np.array([spec[i] for i in set(taste)])
print(spec_lst)
print(set(taste))
print(set(taste_idx))
# ### 独立にパラメータを予測
with pm.Model() as comparing_weight:
alpha = pm.Normal('alpha', mu=0, sd=10, shape=len(set(taste_idx)))
sds = pm.HalfNormal('sds', sd=10, shape=len(set(taste_idx)))
mu = pm.Deterministic('mu', spec_lst[taste_idx]+alpha[taste_idx])
weights = pm.Normal('weights', mu=mu, sd=sds[taste_idx], observed=data['net_weight'].values)
trace = pm.sample(5000, chains=1)
pm.traceplot(trace)
# ### 階層モデルとして推定
with pm.Model() as comparing_weight_h:
# 階層事前分布
am = pm.Normal('am', mu=0, sd=10)
asd = pm.HalfNormal('asd', sd=10)
# 事前分布
alpha = pm.Normal('alpha', mu=am, sd=asd, shape=len(set(taste_idx)))
mu = pm.Deterministic('mu', spec_lst[taste_idx]+alpha[taste_idx])
sds = pm.HalfNormal('sds', sd=10, shape=len(set(taste_idx)))
# 重量モデル
weights = pm.Normal('weights', mu=mu, sd=sds[taste_idx], observed=data['net_weight'].values)
trace = pm.sample(5000, chains=1)
pm.traceplot(trace)
| analysis/jupyter/estimate-net-weight-multi.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Question 1
# +
# Create a bank account class that has two attributes
class bank:
def __init__(self,ownername,balance):
self.ownername = ownername
self.balance = balance
def deposit(self,amount):
self.balance +=amount
print("your updated balance is :", self.balance)
def withdraw(self,amount):
if(self.balance>amount):
self.balance-=amount
print("your updated balance is :", self.balance)
else:
print("you don't have enough cradit in your account, see you have only",self.balance)
# -
Prani = bank("kolanoor",20000)
Prani.deposit(10000)
Prani.withdraw(20000)
Prani.deposit(1000)
Prani.withdraw(15000)
# # Question 2
# +
# Create a cone class that has two attributes
import math
class cone:
def __init__(self,radius,height):
self.radius=radius
self.height=height
def volume(self):
vol = math.pi * (self.radius**2) * (self.height/3)
print("Volume of this cone is : ",vol)
def surfaceArea(self):
area = math.pi* self.radius *(self.radius+(math.sqrt((self.radius**2)+(self.height**2))))
print("Surface area of this cone is ",area)
# -
con = cone(4,6)
con.volume()
con.surfaceArea()
| Assignment 5 Day-6.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Fitting Models
# **Learning Objectives:** learn to fit models to data using linear and non-linear regression.
#
# This material is licensed under the MIT license and was developed by <NAME>. It was adapted from material from <NAME> and <NAME>.
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
from scipy import optimize as opt
from IPython.html.widgets import interact
# ## Introduction
# In Data Science it is common to start with data and develop a *model* of that data. Such models can help to explain the data and make predictions about future observations. In fields like Physics, these models are often given in the form of differential equations, whose solutions explain and predict the data. In most other fields, such differential equations are not known. Often, models have to include sources of uncertainty and randomness. Given a set of data, *fitting* a model to the data is the process of tuning the parameters of the model to *best* explain the data.
#
# When a model has a linear dependence on its parameters, such as $a x^2 + b x + c$, this process is known as *linear regression*. When a model has a non-linear dependence on its parameters, such as $ a e^{bx} $, this process in known as non-linear regression. Thus, fitting data to a straight line model of $m x + b $ is linear regression, because of its linear dependence on $m$ and $b$ (rather than $x$).
# ## Fitting a straight line
# A classical example of fitting a model is finding the slope and intercept of a straight line that goes through a set of data points $\{x_i,y_i\}$. For a straight line the model is:
#
# $$
# y_{model}(x) = mx + b
# $$
#
# Given this model, we can define a metric, or *cost function*, that quantifies the error the model makes. One commonly used metric is $\chi^2$, which depends on the deviation of the model from each data point ($y_i - y_{model}(x_i)$) and the measured uncertainty of each data point $ \sigma_i$:
#
# $$
# \chi^2 = \sum_{i=1}^N \left(\frac{y_i - y_{model}(x)}{\sigma_i}\right)^2
# $$
#
# When $\chi^2$ is small, the model's predictions will be close the data points. Likewise, when $\chi^2$ is large, the model's predictions will be far from the data points. Given this, our task is to minimize $\chi^2$ with respect to the model parameters $\theta = [m, b]$ in order to find the best fit.
#
# To illustrate linear regression, let's create a synthetic data set with a known slope and intercept, but random noise that is additive and normally distributed.
# +
N = 50
m_true = 2
b_true = -1
dy = 2.0 # uncertainty of each point
np.random.seed(0)
xdata = 10 * np.random.random(N) # don't use regularly spaced data
ydata = b_true + m_true * xdata + np.random.normal(0.0, dy, size=N) # our errors are additive
plt.errorbar(xdata, ydata, dy,fmt='.k', ecolor='lightgray')
plt.xlabel('x')
plt.ylabel('y');
# -
# ### Fitting by hand
# It is useful to see visually how changing the model parameters changes the value of $\chi^2$. By using IPython's `interact` function, we can create a user interface that allows us to pick a slope and intercept interactively and see the resulting line and $\chi^2$ value.
#
# Here is the function we want to minimize. Note how we have combined the two parameters into a single parameters vector $\theta = [m, b]$, which is the first argument of the function:
def chi2(theta, x, y, dy):
# theta = [b, m]
return np.sum(((y - theta[0] - theta[1] * x) / dy) ** 2)
def manual_fit(b, m):
modely = m*xdata + b
plt.plot(xdata, modely)
plt.errorbar(xdata, ydata, dy,fmt='.k', ecolor='lightgray')
plt.xlabel('x')
plt.ylabel('y')
plt.text(1, 15, 'b={0:.2f}'.format(b))
plt.text(1, 12.5, 'm={0:.2f}'.format(m))
plt.text(1, 10.0, '$\chi^2$={0:.2f}'.format(chi2([b,m],xdata,ydata, dy)))
interact(manual_fit, b=(-3.0,3.0,0.01), m=(0.0,4.0,0.01));
# Go ahead and play with the sliders and try to:
#
# * Find the lowest value of $\chi^2$
# * Find the "best" line through the data points.
#
# You should see that these two conditions coincide.
# ### Minimize $\chi^2$ using `scipy.optimize.minimize`
# Now that we have seen how minimizing $\chi^2$ gives the best parameters in a model, let's perform this minimization numerically using `scipy.optimize.minimize`. We have already defined the function we want to minimize, `chi2`, so we only have to pass it to `minimize` along with an initial guess and the additional arguments (the raw data):
theta_guess = [0.0,1.0]
result = opt.minimize(chi2, theta_guess, args=(xdata,ydata,dy))
# Here are the values of $b$ and $m$ that minimize $\chi^2$:
theta_best = result.x
print(theta_best)
# These values are close to the true values of $b=-1$ and $m=2$. The reason our values are different is that our data set has a limited number of points. In general, we expect that as the number of points in our data set increases, the model parameters will converge to the true values. But having a limited number of data points is not a problem - it is a reality of most data collection processes.
#
# We can plot the raw data and the best fit line:
# +
xfit = np.linspace(0,10.0)
yfit = theta_best[1]*xfit + theta_best[0]
plt.plot(xfit, yfit)
plt.errorbar(xdata, ydata, dy,
fmt='.k', ecolor='lightgray')
plt.xlabel('x')
plt.ylabel('y');
# -
# ### Minimize $\chi^2$ using `scipy.optimize.leastsq`
# Performing regression by minimizing $\chi^2$ is known as *least squares* regression, because we are minimizing the sum of squares of the deviations. The linear version of this is known as *linear least squares*. For this case, SciPy provides a purpose built function, `scipy.optimize.leastsq`. Instead of taking the $\chi^2$ function to minimize, `leastsq` takes a function that computes the deviations:
# +
def deviations(theta, x, y, dy):
return (y - theta[0] - theta[1] * x) / dy
result = opt.leastsq(deviations, theta_guess, args=(xdata, ydata, dy), full_output=True)
# -
# Here we have passed the `full_output=True` option. When this is passed the [covariance matrix](http://en.wikipedia.org/wiki/Covariance_matrix) $\Sigma_{ij}$ of the model parameters is also returned. The uncertainties (as standard deviations) in the parameters are the square roots of the diagonal elements of the covariance matrix:
#
# $$ \sigma_i = \sqrt{\Sigma_{ii}} $$
#
# A proof of this is beyond the scope of the current notebook.
theta_best = result[0]
theta_cov = result[1]
print('b = {0:.3f} +/- {1:.3f}'.format(theta_best[0], np.sqrt(theta_cov[0,0])))
print('m = {0:.3f} +/- {1:.3f}'.format(theta_best[1], np.sqrt(theta_cov[1,1])))
# We can again plot the raw data and best fit line:
# +
yfit = theta_best[0] + theta_best[1] * xfit
plt.errorbar(xdata, ydata, dy,
fmt='.k', ecolor='lightgray');
plt.plot(xfit, yfit, '-b');
# -
# ### Fitting using `scipy.optimize.curve_fit`
# SciPy also provides a general curve fitting function, `curve_fit`, that can handle both linear and non-linear models. This function:
#
# * Allows you to directly specify the model as a function, rather than the cost function (it assumes $\chi^2$).
# * Returns the covariance matrix for the parameters that provides estimates of the errors in each of the parameters.
#
# Let's apply `curve_fit` to the above data. First we define a model function. The first argument should be the independent variable of the model.
def model(x, b, m):
return m*x+b
# Then call `curve_fit` passing the model function and the raw data. The uncertainties of each data point are provided with the `sigma` keyword argument. If there are no uncertainties, this can be omitted. By default the uncertainties are treated as relative. To treat them as absolute, pass the `absolute_sigma=True` argument.
theta_best, theta_cov = opt.curve_fit(model, xdata, ydata, sigma=dy)
# Again, display the optimal values of $b$ and $m$ along with their uncertainties:
print('b = {0:.3f} +/- {1:.3f}'.format(theta_best[0], np.sqrt(theta_cov[0,0])))
print('m = {0:.3f} +/- {1:.3f}'.format(theta_best[1], np.sqrt(theta_cov[1,1])))
# We can again plot the raw data and best fit line:
# +
xfit = np.linspace(0,10.0)
yfit = theta_best[1]*xfit + theta_best[0]
plt.plot(xfit, yfit)
plt.errorbar(xdata, ydata, dy,
fmt='.k', ecolor='lightgray')
plt.xlabel('x')
plt.ylabel('y');
# -
# ## Non-linear models
# So far we have been using a linear model $y_{model}(x) = m x +b$. Remember this model was linear, not because of its dependence on $x$, but on $b$ and $m$. A non-linear model will have a non-linear dependece on the model parameters. Examples are $A e^{B x}$, $A \cos{B x}$, etc. In this section we will generate data for the following non-linear model:
#
# $$y_{model}(x) = Ae^{Bx}$$
#
# and fit that data using `curve_fit`. Let's start out by using this model to generate a data set to use for our fitting:
npoints = 20
Atrue = 10.0
Btrue = -0.2
xdata = np.linspace(0.0, 20.0, npoints)
dy = np.random.normal(0.0, 0.1, size=npoints)
ydata = Atrue*np.exp(Btrue*xdata) + dy
# Plot the raw data:
plt.plot(xdata, ydata, 'k.')
plt.xlabel('x')
plt.ylabel('y');
# Let's see if we can use non-linear regression to recover the true values of our model parameters. First define the model:
def exp_model(x, A, B):
return A*np.exp(x*B)
# Then use `curve_fit` to fit the model:
theta_best, theta_cov = opt.curve_fit(exp_model, xdata, ydata)
# Our optimized parameters are close to the true values of $A=10$ and $B=-0.2$:
print('A = {0:.3f} +/- {1:.3f}'.format(theta_best[0], np.sqrt(theta_cov[0,0])))
print('B = {0:.3f} +/- {1:.3f}'.format(theta_best[1], np.sqrt(theta_cov[1,1])))
# Plot the raw data and fitted model:
xfit = np.linspace(0,20)
yfit = exp_model(xfit, theta_best[0], theta_best[1])
plt.plot(xfit, yfit)
plt.plot(xdata, ydata, 'k.')
plt.xlabel('x')
plt.ylabel('y');
# ### A note about transforming to a linear model
#
# Another approach to dealing with non-linear models is to linearize them with a transformation. For example, the exponential model used above,
#
# $$y_{model}(x) = Ae^{Bx},$$
#
# can be linearized by taking the natural log of both sides:
#
# $$ ln(y) = ln(A) + B x $$
#
# This model is linear in the parameters $ln(A)$ and $B$ and can be treated as a standard linear regression problem. This approach is used in most introductory physics laboratories. **However, in most cases, transforming to a linear model will give a poor fit. The reasons for this are a bit subtle, but here is the basic idea:
#
# * Least squares regression assumes that errors are symmetric, additive and normally distributed. This assumption has been present throughout this notebook, when we generated data by *adding* a small amount of randomness to our data using `np.random.normal`.
# * Transforming the data with a non-linear transformation, such as the square root, exponential or logarithm will not lead to errors that follow this assumption.
# * However, in the rare case that there are no (minimal) random errors in the original data set, the transformation approach will give the same result as the non-linear regression on the original model.
#
# Here is a [nice discussion](http://www.mathworks.com/help/stats/examples/pitfalls-in-fitting-nonlinear-models-by-transforming-to-linearity.html) of this in the Matlab documentation.
# ## Model selection
# In all of the examples in this notebook, we started with a model and used that model to generate data. This was done to make it easy to check the predicted model parameters with the true values used to create the data set. However, in the real world, you almost never know the model underlying the data. Because of this, there is an additional step called *model selection* where you have to figure out a way to pick a good model. This is a notoriously difficult problem, especially when the randomness in the data is large.
#
# * Pick the simplest possible model. In general picking a more complex model will give a better fit. However, it won't be a useful model and will make poor predictions about future data. This is known as [overfitting](http://en.wikipedia.org/wiki/Overfitting).
# * Whenever possible, pick a model that has a underlying theoretical foundation or motivation. For example, in Physics, most of our models come from well tested differential equations.
# * There are more advanced methods (AIC,BIC) that can assist in this model selection process. A good discussion can be found in [this notebook](https://github.com/jakevdp/2014_fall_ASTR599/blob/master/notebooks/14_Optimization.ipynb) by <NAME>.
| Fitting/FittingModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: action_prediction
# language: python
# name: action_prediction
# ---
# +
# load libraries
import pandas as pd
import numpy as np
import seaborn as sns
import os
import matplotlib.pyplot as plt
from action_prediction import constants as const
from action_prediction.data import DataSet
from action_prediction import run_models
# %load_ext autoreload
# %autoreload 2
import warnings
warnings.filterwarnings("ignore")
# +
# initialize class
data = DataSet(task='social_prediction')
# load behavior
df_behav = data.load_behav()
# load eyetracking
df_eye = data.load_eye(data_type='events')
# merge eyetracking with behav
df_merged = data.merge_behav_eye(dataframe_behav=df_behav, dataframe_eye=df_eye)
# +
# run models
# NOTE:
# model functions have been saved in modeling.py and the high-level model routine has been saved in run_models.py
# to modify the models, go to get_model_features in modelling.py and hardcode in new model features
models = run_models.run(dataframe = df_merged, model_names= ['context', 'eye-tracking', 'eye-tracking + context'])
models
# -
| notebooks/1.1-mk-modelling.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 1.1.0 Collect Altmetrics
import pandas as pd
import requests
from glob import glob
import json
from clustergrammer2 import net
all_files = glob('../markdown_files/*.md')
len(all_files)
# ### Get the Latest Papers
url = 'https://connect.biorxiv.org/relate/collection_json.php?grp=181'
r = requests.get(url)
req_dict = json.loads(r.text)
# ### Update Altmetric Scores and Links
altmetric_base_url = 'http://api.altmetric.com/v1/doi/'
altmetric_scores = {}
altmetric_details_url = {}
doi_words = {}
all_words = []
arr_papers = req_dict['rels']
print('number of papers', len(arr_papers))
for inst_paper in arr_papers:
inst_doi = inst_paper['rel_doi']
r = requests.get(altmetric_base_url + inst_doi)
if r.text != 'Not Found':
alt_req_dict = json.loads(r.text)
altmetric_scores[inst_doi] = alt_req_dict['score']
altmetric_details_url[inst_doi] = alt_req_dict['details_url']
r.text
# ### Save Scores and Links to JSON Document
# !mkdir ../altmetric_data/
net.save_dict_to_json(altmetric_scores, '../altmetric_data/altmetric_scores.json')
net.save_dict_to_json(altmetric_scores, '../altmetric_data/altmetric_details_url.json')
| notebooks/1.1.0_Collect_Altmetrics.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
movies = pd.read_csv('tmdb_5000_movies.csv')
credits = pd.read_csv('tmdb_5000_credits.csv')
movies.head(2) #Gives info about the first two movies,in the dataset
movies.shape #gives the dimensions of the matrix
credits.head(2) #Gives info about the first two movie credits,in the dataset
credits.shape #gives the dimensions of the matrix
"""
Shows all the attributes and values associated with the
first movie's crew (in our case its avatar)
"""
credits.head(1)['crew'].values
movies = movies.merge(credits,on = 'title') #Merging the two datasets using the common column title(can also be joined as per id)
movies.shape #gives the dimensions of the merged dataset
movies.head(1) #columns have now increased, news column are added at the end
"""
Useful Recommending attributes
genre :Theme based recommending
keywords :Sometimes when You cant really name the movie, but you have words to describe
id :For handling purposes
language :(but in our dataset 4500+ are in english so, not very useful)
title :Exact titles can result in highly accurate results
overview :Similiar stories help in recommending them to the user
popularity :Very important factor, though our approach currently avoids the numeric data
Release data :For example some would like to watch movies from the 90s or 80s
cast :Recommendation based on actors,actresses
crew :Recommendation based on directors
"""
#Keeping attributes necessary for creating tags for our data
movies = movies[['movie_id','title','genres','overview','keywords','cast','crew']]
# +
#Preprocessing of data begins
#STEP 1 : Checking for Any missing data
movies.isnull().sum() #Gives the summary of rows which have null values
# -
movies.dropna(inplace=True) #dropna() function is used to remove rows and columns with Null/NaN values.
# +
#STEP 2: Checking for any duplicated data
movies.duplicated().sum()
# +
#STEP 3: Customising and Refining data to get our tags for every movie in the dataset
import ast
# +
#Three Helper Functions to ease the task of customising and refining data
def convert(data): #The data set is in string format
List = [] #We need a list
for i in ast.literal_eval(data): #Converting the string data into a list of tags
List.append(i['name'])
return List
# -
"""
The method for converting the string data to list of tags, is same as that used for keywords and genres
But in the case for cast column, the idea is to give priority to the top 4 leading acters/actresses for recommendation
This will increase the efficiency and readability of the code(as well as the working matrix)
This is done to get the recommendation as per the first thought that the use gets when he/she hears the name of a movie
For example : If the user hears the name Iron Man, the first acter that will pop up in the user's mind will be
'<NAME>'
"""
def top_four_people(data): #The data set is in string format
List = []
counter = 0 # Counter to get the top 4 cast of the movie
for i in ast.literal_eval(data):
if counter < 4:
List.append(i['name'])
counter += 1
return List
"""
For the case of Crew column the only need is to get the name of the director of the movie.
People usually don't remember who was the VFX expert, or who did the final editing, or who designed the sets
But people Do remember The Director in many cases
For Example, the momemnt User hears the name Justice League, the first is Snyder's Cut, Which actually gives the name
<NAME>, the director of the Snyder cut
Proving Point : What was name of the head of the vfx team?
: Like its mentioned, people dont remember :)
"""
def get_me_the_director(data):
List = []
for i in ast.literal_eval(data):
if i['job'] == 'Director':
List.append(i['name'])
return List
# +
#Refining our data with the help of helper functions
movies['genres']= movies['genres'].apply(convert) #Provides the list of raw tags for all the movies containing
#the name-values in the genre column of the movies
movies['keywords']= movies['keywords'].apply(convert) #Provides the list of raw tags for all the movies containing
#the name-values in the keywords column of the movies
movies['cast']= movies['cast'].apply(top_four_people) #Provides the list of top four actors/actressses
#for all the movies
movies['crew']= movies['crew'].apply(get_me_the_director) #Provides the list of directors for all the movies
movies['overview'] = movies['overview'].apply(lambda x:x.split()) #Converts the overview string for each movie to a list
#containg all the words in the string
# -
movies.head() #Displaying all the changes done to refine our data, conversion to a list of tags
# +
"""
STEP 4 : Transformation of data
Before creating the tags that can be used in the recommender system, the spaces between the raw tags has to be removed
So that no ambiguity arises when a raw tag which consists of a few words, is converted into usable tags where every word of
the raw-tag becomes and individual tag.
For Example : The system wants 'Science Fiction' as a one word tag 'ScienceFiction' and not as 'Science' and 'Fiction'
seperately.
: This space is problem as now 'science' and 'fiction' tags will be used even when there might not be a need.
: Like in case of a real autobiography of a scientist, science will be a tag, but the user who watched a
science-fiction movie might not be interested in real stories. And still will receive the recommendation of
every movie which may or may not be fiction but related to science, just because 'science' and 'fiction' tags
were not dealt with properly.
Similair problem can occur when two actors sharing the first name are treated as an individual entity
Below is the code implementation to do the task
The lambda function used, removes the space between each element(if any) of the list supplied
"""
movies['cast'] = movies['cast'].apply(lambda x:[i.replace(' ','') for i in x])
movies['crew'] = movies['crew'].apply(lambda x:[i.replace(' ','') for i in x])
movies['genres'] = movies['genres'].apply(lambda x:[i.replace(' ','') for i in x])
movies['keywords'] = movies['keywords'].apply(lambda x:[i.replace(' ','') for i in x])
# -
movies.head() #Displaying data with the changes done to its spacing
# +
"""
STEP 5 : Providing tags to our data
Now that the missing-data problem, duplicate data problem and space problems have been resolved, its time to get the refined
tags, which will be used by our data.
For this a new column containing the tags will be created and the five columns that were refined previously will be merged
into it.
Following this step the five columns will be dropped(Not necessary but must be done), so as to improve the space utilisation
"""
movies['tags'] = movies['overview'] + movies['genres'] + movies['keywords'] + movies['cast'] + movies['crew']
#A new database with only three columns: movie_id,title and tags
Movie = movies.drop(columns=['overview','genres','keywords','cast','crew'])
# -
Movie.head() #Displaying data with the changes done to its spacing
# +
#Now converting the merged lists in tags into one string data i.e. a group of tags
Movie['tags'] = Movie['tags'].apply(lambda x: " ".join(x))
#The mergeed lists are now combined into one group,below mentioned code displays the tags for first movie
Movie['tags'][0]
# -
"""
The Natural Language Toolkit (NLTK) is used for processing the tags, to remove redundancy and to save on the space occupied by almost identical tags
PorterStemmer is used for stemming the tags
"""
import nltk
from nltk import PorterStemmer
ps = PorterStemmer()
# +
#Function to stem the tags and return the list of tags
def stem(data):
List = []
for i in data.split():
List.append(ps.stem(i))
return " ".join(List)
# -
Movie['tags'] = Movie['tags'].apply(stem) #Stemming the tags
# +
"""
Now that tags are ready Vectorisation of data can be done.
The main task at hand is to find the similarities between the movies so that the recommender can act accordingly and
can provide the appropriate.
To do this the entire dataset has to be vectorised where each movie reprents a point on a 2d graph
And the recommender system will suggest the closest 'n' points, from a given point.
To convert the text to vectors, our system will be using "Bag of Words Technique".
There are other advanced techniques but because it is our first project so we chose to use a simpler yet efficient technique.
While vectorisation, the stop words are to omitted, eg: words like is,are,to etc.
For this task Scikit-learn library will be used
"""
from sklearn.feature_extraction.text import CountVectorizer
cv = CountVectorizer(max_features=5000,stop_words='english')
vector = cv.fit_transform(Movie['tags']).toarray()
vector.shape
# -
"""
For getting the cosine similarity
The definition of similarity between two vectors u and v is, in fact, the ratio between their dot product and the product of their magnitudes.
By applying the definition of similarity, this will be in fact equal to 1 if the two vectors are identical,
and it will be 0 if the two are orthogonal.
In other words, the similarity is a number bounded between 0 and 1 that tells us how much the two vectors are similar.
"""
from sklearn.metrics.pairwise import cosine_similarity
similarity = cosine_similarity(vector)
similarity[0] #Shows The Similarity between the tags
def recommend(movie):
index = Movie[Movie['title'] == movie].index[0]
distances = sorted(list(enumerate(similarity[index])),reverse=True,key = lambda x: x[1])
for i in distances[1:7]:
print(Movie.iloc[i[0]].title)
recommend('Ramanujan')
| .ipynb_checkpoints/Presentation-ready-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Moments: Mean, Variance, Skew, Kurtosis
# Create a roughly normal-distributed random set of data:
# +
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
vals = np.random.normal(10, 0.5, 10000)
plt.hist(vals, 50)
plt.show()
# -
# The first moment is the mean; this data should average out to about 0:
np.mean(vals)
# The second moment is the variance:
np.var(vals)
# The third moment is skew - since our data is nicely centered around 0, it should be almost 0:
import scipy.stats as sp
sp.skew(vals)
# The fourth moment is "kurtosis", which describes the shape of the tail. For a normal distribution, this is 0:
sp.kurtosis(vals)
# ## Activity
# Understanding skew: change the normal distribution to be centered around 10 instead of 0, and see what effect that has on the moments.
#
# The skew is still near zero; skew is associated with the shape of the distribution, not its actual offset in X.
| mlcourse/Moments.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: ipykernel_py2
# ---
def count(numbers):
total = 0
for x in numbers:
if x < 20:
total += 1
return total
list_1 = [1,3,7,15,23,43,56,98]
count(list_1)
list_2 = [1,3,7,15,23,43,56,98,17]
count(list_2)
| course_2/course_material/Part_4_Python/S29_L175/Python 2/All In - Lecture_Py2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: venv-datascience
# language: python
# name: venv-datascience
# ---
import pandas as pd
delays_df = pd.read_csv("Data/Lots_of_flight_data.csv")
delays_df.head()
# # 1) break down data into X (features), y(target/label) data set
# - x: input
# - y: predit or output
X = delays_df.loc[:, ["DISTANCE", "CRS_ELAPSED_TIME"]]
X.head()
y = delays_df.loc[:, ["ARR_DELAY"]]
y.head()
# # 2) Split into Training and Testing Data
# +
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(
X,
y,
test_size = 0.3,
random_state = 42
)
# -
X_train.shape
X_test.shape
y_train.shape
y_test.shape
y_train.head()
X_train.head()
| More Python Data Tools - Microsoft/06.Splitting test and training data with scikit-learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Table of Contents
# 1. What is tuple?
# Advantages of Tuple over List
# 2. Creating a tuple
# 3. Accessing Elements in a Tuple
# 1. Indexing
# 2. Negative Indexing
# 3. Slicing
# 4. Changing a tuple
# 5. Deleting a Tuple
# 6. Python Tuple Methods
# 7. Other Tuple Operations
# 1. Tuple Membership Test
# 2. Iterating Through a Tuple
# 3. Built-in Functions with Tuple
# # What is tuple?
# In Python programming, a tuple is similar to a list. The difference between the two is that we cannot change the elements of a tuple once it is assigned whereas in a list, elements can be changed.
# # Advantages of Tuple over List
# Since, tuples are quite similiar to lists, both of them are used in similar situations as well.
#
# However, there are certain advantages of implementing a tuple over a list. Below listed are some of the main advantages:
#
# We generally use tuple for heterogeneous (different) datatypes and list for homogeneous (similar) datatypes.
# Since tuple are immutable, iterating through tuple is faster than with list. So there is a slight performance boost.
# Tuples that contain immutable elements can be used as key for a dictionary. With list, this is not possible.
# If you have data that doesn't change, implementing it as tuple will guarantee that it remains write-protected.
# # Creating a Tuple
# A tuple is created by placing all the items (elements) inside a parentheses (), separated by comma. The parentheses are optional but is a good practice to write it.
#
# A tuple can have any number of items and they may be of different types (integer, float, list, string etc.).
# +
# empty tuple
# Output: ()
my_tuple = ()
print(my_tuple)
# tuple having integers
# Output: (1, 2, 3)
my_tuple = (1, 2, 3)
print(my_tuple)
# tuple with mixed datatypes
# Output: (1, "Hello", 3.4)
my_tuple = (1, "Hello", 3.4)
print(my_tuple)
# nested tuple
# Output: ("mouse", [8, 4, 6], (1, 2, 3))
my_tuple = ("mouse", [8, 4, 6], (1, 2, 3))
print(my_tuple)
# tuple can be created without parentheses
# also called tuple packing
# Output: 3, 4.6, "dog"
my_tuple = 3, 4.6, "dog"
print(my_tuple)
# tuple unpacking is also possible
# Output:
# 3
# 4.6
# dog
a, b, c = my_tuple
print(a)
print(b)
print(c)
# -
# Creating a tuple with one element is a bit tricky.
#
# Having one element within parentheses is not enough. We will need a trailing comma to indicate that it is in fact a tuple.
# +
# only parentheses is not enough
# Output: <class 'str'>
my_tuple = ("hello")
print(type(my_tuple))
# need a comma at the end
# Output: <class 'tuple'>
my_tuple = ("hello",)
print(type(my_tuple))
# parentheses is optional
# Output: <class 'tuple'>
my_tuple = "hello",
print(type(my_tuple))
# -
# # Accessing Elements in a Tuple
# There are various ways in which we can access the elements of a tuple.
#
# 1. Indexing
# We can use the index operator [] to access an item in a tuple where the index starts from 0.
#
# So, a tuple having 6 elements will have index from 0 to 5. Trying to access an element other that (6, 7,...) will raise an IndexError.
#
# The index must be an integer, so we cannot use float or other types. This will result into TypeError.
#
# Likewise, nested tuple are accessed using nested indexing, as shown in the example below.
# +
my_tuple = ('p','e','r','m','i','t')
# Output: 'p'
print(my_tuple[0])
# Output: 't'
print(my_tuple[5])
# index must be in range
# If you uncomment line 14,
# you will get an error.
# IndexError: list index out of range
#print(my_tuple[6])
# index must be an integer
# If you uncomment line 21,
# you will get an error.
# TypeError: list indices must be integers, not float
#my_tuple[2.0]
# nested tuple
n_tuple = ("mouse", [8, 4, 6], (1, 2, 3))
# nested index
# Output: 's'
print(n_tuple[0][3])
# nested index
# Output: 4
print(n_tuple[1][1])
# -
# # Negative Indexing
# Python allows negative indexing for its sequences.
#
# The index of -1 refers to the last item, -2 to the second last item and so on.
# +
my_tuple = ('p','e','r','m','i','t')
# Output: 't'
print(my_tuple[-1])
# Output: 'p'
print(my_tuple[-6])
# -
# # 3. Slicing
# We can access a range of items in a tuple by using the slicing operator - colon ":".
# +
my_tuple = ('p','r','o','g','r','a','m','i','z')
# elements 2nd to 4th
# Output: ('r', 'o', 'g')
print(my_tuple[1:4])
# elements beginning to 2nd
# Output: ('p', 'r')
print(my_tuple[:-7])
# elements 8th to end
# Output: ('i', 'z')
print(my_tuple[7:])
# elements beginning to end
# Output: ('p', 'r', 'o', 'g', 'r', 'a', 'm', 'i', 'z')
print(my_tuple[:])
# -
# # Changing a Tuple
# Unlike lists, tuples are immutable.
#
# This means that elements of a tuple cannot be changed once it has been assigned. But, if the element is itself a mutable datatype like list, its nested items can be changed.
#
# We can also assign a tuple to different values (reassignment).
#
# +
my_tuple = (4, 2, 3, [6, 5])
# we cannot change an element
# If you uncomment line 8
# you will get an error:
# TypeError: 'tuple' object does not support item assignment
#my_tuple[1] = 9
# but item of mutable element can be changed
# Output: (4, 2, 3, [9, 5])
my_tuple[3][0] = 9
print(my_tuple)
# tuples can be reassigned
# Output: ('p', 'r', 'o', 'g', 'r', 'a', 'm', 'i', 'z')
my_tuple = ('p','r','o','g','r','a','m','i','z')
print(my_tuple)
# -
# We can use + operator to combine two tuples. This is also called concatenation.
#
# We can also repeat the elements in a tuple for a given number of times using the * operator.
#
# Both + and * operations result into a new tuple.
# +
# Concatenation
# Output: (1, 2, 3, 4, 5, 6)
print((1, 2, 3) + (4, 5, 6))
# Repeat
# Output: ('Repeat', 'Repeat', 'Repeat')
print(("Repeat",) * 3)
# -
# # Deleting a Tuple
# As discussed above, we cannot change the elements in a tuple. That also means we cannot delete or remove items from a tuple.
#
# But deleting a tuple entirely is possible using the keyword del.
# +
my_tuple = ('p','r','o','g','r','a','m','i','z')
# can't delete items
# if you uncomment line 8,
# you will get an error:
# TypeError: 'tuple' object doesn't support item deletion
#del my_tuple[3]
# can delete entire tuple
# NameError: name 'my_tuple' is not defined
del my_tuple
my_tuple
# -
# # Python Tuple Methods
# Methods that add items or remove items are not available with tuple. Only the following two methods are available.
# |Method |Description |
# -------------------------------------------------------------------
# |**count(x)** |Return the number of items that is equal to x|
# |**index(x)** |Return index of first item that is equal to x|
# +
my_tuple = ('a','p','p','l','e',)
# Count
# Output: 2
print(my_tuple.count('p'))
# Index
# Output: 3
print(my_tuple.index('l'))
# -
# # Other Tuple Operations
# # Tuple Membership Test
# We can test if an item exists in a tuple or not, using the keyword in.
# +
my_tuple = ('a','p','p','l','e',)
# In operation
# Output: True
print('a' in my_tuple)
# Output: False
print('b' in my_tuple)
# Not in operation
# Output: True
print('g' not in my_tuple)
# -
# # Iterating Through a Tuple
# Using a for loop we can iterate though each item in a tuple.
# Output:
# Hello John
# Hello Kate
for name in ('John','Kate'):
print("Hello",name)
# # Built-in Functions with Tuple
# Built-in functions like all(), any(), enumerate(), len(), max(), min(), sorted(), tuple() etc. are commonly used with tuple to perform different tasks.
from IPython.display import Image
Image(filename='img/Tuple Built Funcation.jpg')
| Data structure/Python Tuple.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Mark and Recapture
# + [markdown] tags=[]
# Think Bayes, Second Edition
#
# Copyright 2020 <NAME>
#
# License: [Attribution-NonCommercial-ShareAlike 4.0 International (CC BY-NC-SA 4.0)](https://creativecommons.org/licenses/by-nc-sa/4.0/)
# + tags=[]
# If we're running on Colab, install empiricaldist
# https://pypi.org/project/empiricaldist/
import sys
IN_COLAB = 'google.colab' in sys.modules
if IN_COLAB:
# !pip install empiricaldist
# + tags=[]
# Get utils.py
from os.path import basename, exists
def download(url):
filename = basename(url)
if not exists(filename):
from urllib.request import urlretrieve
local, _ = urlretrieve(url, filename)
print('Downloaded ' + local)
download('https://github.com/AllenDowney/ThinkBayes2/raw/master/soln/utils.py')
# + tags=[]
from utils import set_pyplot_params
set_pyplot_params()
# -
# This chapter introduces "mark and recapture" experiments, in which we sample individuals from a population, mark them somehow, and then take a second sample from the same population. Seeing how many individuals in the second sample are marked, we can estimate the size of the population.
#
# Experiments like this were originally used in ecology, but turn out to be useful in many other fields. Examples in this chapter include software engineering and epidemiology.
#
# Also, in this chapter we'll work with models that have three parameters, so we'll extend the joint distributions we've been using to three dimensions.
#
# But first, grizzly bears.
# ## The Grizzly Bear Problem
#
# In 1996 and 1997 researchers deployed bear traps in locations in British Columbia and Alberta, Canada, in an effort to estimate the population of grizzly bears. They describe the experiment in [this article](https://www.researchgate.net/publication/229195465_Estimating_Population_Size_of_Grizzly_Bears_Using_Hair_Capture_DNA_Profiling_and_Mark-Recapture_Analysis).
#
# The "trap" consists of a lure and several strands of barbed wire intended to capture samples of hair from bears that visit the lure. Using the hair samples, the researchers use DNA analysis to identify individual bears.
#
# During the first session, the researchers deployed traps at 76 sites. Returning 10 days later, they obtained 1043 hair samples and identified 23 different bears. During a second 10-day session they obtained 1191 samples from 19 different bears, where 4 of the 19 were from bears they had identified in the first batch.
#
# To estimate the population of bears from this data, we need a model for the probability that each bear will be observed during each session. As a starting place, we'll make the simplest assumption, that every bear in the population has the same (unknown) probability of being sampled during each session.
# With these assumptions we can compute the probability of the data for a range of possible populations.
#
# As an example, let's suppose that the actual population of bears is 100.
#
# After the first session, 23 of the 100 bears have been identified.
# During the second session, if we choose 19 bears at random, what is the probability that 4 of them were previously identified?
# I'll define
#
# * $N$: actual population size, 100.
#
# * $K$: number of bears identified in the first session, 23.
#
# * $n$: number of bears observed in the second session, 19 in the example.
#
# * $k$: number of bears in the second session that were previously identified, 4.
#
# For given values of $N$, $K$, and $n$, the probability of finding $k$ previously-identified bears is given by the [hypergeometric distribution](https://en.wikipedia.org/wiki/Hypergeometric_distribution):
#
# $$\binom{K}{k} \binom{N-K}{n-k}/ \binom{N}{n}$$
#
# where the [binomial coefficient](https://en.wikipedia.org/wiki/Binomial_coefficient), $\binom{K}{k}$, is the number of subsets of size $k$ we can choose from a population of size $K$.
# To understand why, consider:
#
# * The denominator, $\binom{N}{n}$, is the number of subsets of $n$ we could choose from a population of $N$ bears.
#
# * The numerator is the number of subsets that contain $k$ bears from the previously identified $K$ and $n-k$ from the previously unseen $N-K$.
#
# SciPy provides `hypergeom`, which we can use to compute this probability for a range of values of $k$.
# +
import numpy as np
from scipy.stats import hypergeom
N = 100
K = 23
n = 19
ks = np.arange(12)
ps = hypergeom(N, K, n).pmf(ks)
# -
# The result is the distribution of $k$ with given parameters $N$, $K$, and $n$.
# Here's what it looks like.
# + tags=[]
import matplotlib.pyplot as plt
from utils import decorate
plt.bar(ks, ps)
decorate(xlabel='Number of bears observed twice',
ylabel='PMF',
title='Hypergeometric distribution of k (known population 100)')
# -
# The most likely value of $k$ is 4, which is the value actually observed in the experiment.
# That suggests that $N=100$ is a reasonable estimate of the population, given this data.
#
# We've computed the distribution of $k$ given $N$, $K$, and $n$.
# Now let's go the other way: given $K$, $n$, and $k$, how can we estimate the total population, $N$?
# ## The Update
#
# As a starting place, let's suppose that, prior to this study, an expert estimates that the local bear population is between 50 and 500, and equally likely to be any value in that range.
#
# I'll use `make_uniform` to make a uniform distribution of integers in this range.
# +
import numpy as np
from utils import make_uniform
qs = np.arange(50, 501)
prior_N = make_uniform(qs, name='N')
prior_N.shape
# -
# So that's our prior.
#
# To compute the likelihood of the data, we can use `hypergeom` with constants `K` and `n`, and a range of values of `N`.
# +
Ns = prior_N.qs
K = 23
n = 19
k = 4
likelihood = hypergeom(Ns, K, n).pmf(k)
# -
# We can compute the posterior in the usual way.
posterior_N = prior_N * likelihood
posterior_N.normalize()
# And here's what it looks like.
# + tags=[]
posterior_N.plot(color='C4')
decorate(xlabel='Population of bears (N)',
ylabel='PDF',
title='Posterior distribution of N')
# -
# The most likely value is 109.
posterior_N.max_prob()
# But the distribution is skewed to the right, so the posterior mean is substantially higher.
posterior_N.mean()
# And the credible interval is quite wide.
posterior_N.credible_interval(0.9)
# This solution is relatively simple, but it turns out we can do a little better if we model the unknown probability of observing a bear explicitly.
# ## Two-Parameter Model
#
# Next we'll try a model with two parameters: the number of bears, `N`, and the probability of observing a bear, `p`.
#
# We'll assume that the probability is the same in both rounds, which is probably reasonable in this case because it is the same kind of trap in the same place.
#
# We'll also assume that the probabilities are independent; that is, the probability a bear is observed in the second round does not depend on whether it was observed in the first round. This assumption might be less reasonable, but for now it is a necessary simplification.
#
# Here are the counts again:
K = 23
n = 19
k = 4
# For this model, I'll express the data in a notation that will make it easier to generalize to more than two rounds:
#
# * `k10` is the number of bears observed in the first round but not the second,
#
# * `k01` is the number of bears observed in the second round but not the first, and
#
# * `k11` is the number of bears observed in both rounds.
#
# Here are their values.
k10 = 23 - 4
k01 = 19 - 4
k11 = 4
# Suppose we know the actual values of `N` and `p`. We can use them to compute the likelihood of this data.
#
# For example, suppose we know that `N=100` and `p=0.2`.
# We can use `N` to compute `k00`, which is the number of unobserved bears.
# +
N = 100
observed = k01 + k10 + k11
k00 = N - observed
k00
# -
# For the update, it will be convenient to store the data as a list that represents the number of bears in each category.
x = [k00, k01, k10, k11]
x
# Now, if we know `p=0.2`, we can compute the probability a bear falls in each category. For example, the probability of being observed in both rounds is `p*p`, and the probability of being unobserved in both rounds is `q*q` (where `q=1-p`).
p = 0.2
q = 1-p
y = [q*q, q*p, p*q, p*p]
y
# Now the probability of the data is given by the [multinomial distribution](https://en.wikipedia.org/wiki/Multinomial_distribution):
#
# $$\frac{N!}{\prod x_i!} \prod y_i^{x_i}$$
#
# where $N$ is actual population, $x$ is a sequence with the counts in each category, and $y$ is a sequence of probabilities for each category.
#
# SciPy provides `multinomial`, which provides `pmf`, which computes this probability.
# Here is the probability of the data for these values of `N` and `p`.
# +
from scipy.stats import multinomial
likelihood = multinomial.pmf(x, N, y)
likelihood
# -
# That's the likelihood if we know `N` and `p`, but of course we don't. So we'll choose prior distributions for `N` and `p`, and use the likelihoods to update it.
# ## The Prior
#
# We'll use `prior_N` again for the prior distribution of `N`, and a uniform prior for the probability of observing a bear, `p`:
qs = np.linspace(0, 0.99, num=100)
prior_p = make_uniform(qs, name='p')
# We can make a joint distribution in the usual way.
# +
from utils import make_joint
joint_prior = make_joint(prior_p, prior_N)
joint_prior.shape
# -
# The result is a Pandas `DataFrame` with values of `N` down the rows and values of `p` across the columns.
# However, for this problem it will be convenient to represent the prior distribution as a 1-D `Series` rather than a 2-D `DataFrame`.
# We can convert from one format to the other using `stack`.
# +
from empiricaldist import Pmf
joint_pmf = Pmf(joint_prior.stack())
joint_pmf.head(3)
# + tags=[]
type(joint_pmf)
# + tags=[]
type(joint_pmf.index)
# + tags=[]
joint_pmf.shape
# -
# The result is a `Pmf` whose index is a `MultiIndex`.
# A `MultiIndex` can have more than one column; in this example, the first column contains values of `N` and the second column contains values of `p`.
#
# The `Pmf` has one row (and one prior probability) for each possible pair of parameters `N` and `p`.
# So the total number of rows is the product of the lengths of `prior_N` and `prior_p`.
#
# Now we have to compute the likelihood of the data for each pair of parameters.
# ## The Update
#
# To allocate space for the likelihoods, it is convenient to make a copy of `joint_pmf`:
likelihood = joint_pmf.copy()
# As we loop through the pairs of parameters, we compute the likelihood of the data as in the previous section, and then store the result as an element of `likelihood`.
# +
observed = k01 + k10 + k11
for N, p in joint_pmf.index:
k00 = N - observed
x = [k00, k01, k10, k11]
q = 1-p
y = [q*q, q*p, p*q, p*p]
likelihood[N, p] = multinomial.pmf(x, N, y)
# -
# Now we can compute the posterior in the usual way.
# + tags=[]
posterior_pmf = joint_pmf * likelihood
posterior_pmf.normalize()
# -
# We'll use `plot_contour` again to visualize the joint posterior distribution.
# But remember that the posterior distribution we just computed is represented as a `Pmf`, which is a `Series`, and `plot_contour` expects a `DataFrame`.
#
# Since we used `stack` to convert from a `DataFrame` to a `Series`, we can use `unstack` to go the other way.
joint_posterior = posterior_pmf.unstack()
# And here's what the result looks like.
# + tags=[]
from utils import plot_contour
plot_contour(joint_posterior)
decorate(title='Joint posterior distribution of N and p')
# -
# The most likely values of `N` are near 100, as in the previous model. The most likely values of `p` are near 0.2.
#
# The shape of this contour indicates that these parameters are correlated. If `p` is near the low end of the range, the most likely values of `N` are higher; if `p` is near the high end of the range, `N` is lower.
#
# Now that we have a posterior `DataFrame`, we can extract the marginal distributions in the usual way.
# +
from utils import marginal
posterior2_p = marginal(joint_posterior, 0)
posterior2_N = marginal(joint_posterior, 1)
# + [markdown] tags=[]
# Here's the posterior distribution for `p`:
# + tags=[]
posterior2_p.plot(color='C1')
decorate(xlabel='Probability of observing a bear',
ylabel='PDF',
title='Posterior marginal distribution of p')
# + [markdown] tags=[]
# The most likely values are near 0.2.
# -
# Here's the posterior distribution for `N` based on the two-parameter model, along with the posterior we got using the one-parameter (hypergeometric) model.
# +
posterior_N.plot(label='one-parameter model', color='C4')
posterior2_N.plot(label='two-parameter model', color='C1')
decorate(xlabel='Population of bears (N)',
ylabel='PDF',
title='Posterior marginal distribution of N')
# -
# With the two-parameter model, the mean is a little lower and the 90% credible interval is a little narrower.
# + tags=[]
print(posterior_N.mean(),
posterior_N.credible_interval(0.9))
# + tags=[]
print(posterior2_N.mean(),
posterior2_N.credible_interval(0.9))
# + [markdown] tags=[]
# The two-parameter model yields a narrower posterior distribution for `N`, compared to the one-parameter model, because it takes advantage of an additional source of information: the consistency of the two observations.
#
# To see how this helps, consider a scenario where `N` is relatively low, like 138 (the posterior mean of the two-parameter model).
# + tags=[]
N1 = 138
# + [markdown] tags=[]
# Given that we saw 23 bears during the first trial and 19 during the second, we can estimate the corresponding value of `p`.
# + tags=[]
mean = (23 + 19) / 2
p = mean/N1
p
# + [markdown] tags=[]
# With these parameters, how much variability do you expect in the number of bears from one trial to the next? We can quantify that by computing the standard deviation of the binomial distribution with these parameters.
# + tags=[]
from scipy.stats import binom
binom(N1, p).std()
# + [markdown] tags=[]
# Now let's consider a second scenario where `N` is 173, the posterior mean of the one-parameter model. The corresponding value of `p` is lower.
# + tags=[]
N2 = 173
p = mean/N2
p
# + [markdown] tags=[]
# In this scenario, the variation we expect to see from one trial to the next is higher.
# + tags=[]
binom(N2, p).std()
# + [markdown] tags=[]
# So if the number of bears we observe is the same in both trials, that would be evidence for lower values of `N`, where we expect more consistency.
# If the number of bears is substantially different between the two trials, that would be evidence for higher values of `N`.
#
# In the actual data, the difference between the two trials is low, which is why the posterior mean of the two-parameter model is lower.
# The two-parameter model takes advantage of additional information, which is why the credible interval is narrower.
# + [markdown] tags=[]
# ## Joint and Marginal Distributions
#
# Marginal distributions are called "marginal" because in a common visualization they appear in the margins of the plot.
#
# Seaborn provides a class called `JointGrid` that creates this visualization.
# The following function uses it to show the joint and marginal distributions in a single plot.
# + tags=[]
import pandas as pd
from seaborn import JointGrid
def joint_plot(joint, **options):
"""Show joint and marginal distributions.
joint: DataFrame that represents a joint distribution
options: passed to JointGrid
"""
# get the names of the parameters
x = joint.columns.name
x = 'x' if x is None else x
y = joint.index.name
y = 'y' if y is None else y
# make a JointGrid with minimal data
data = pd.DataFrame({x:[0], y:[0]})
g = JointGrid(x=x, y=y, data=data, **options)
# replace the contour plot
g.ax_joint.contour(joint.columns,
joint.index,
joint,
cmap='viridis')
# replace the marginals
marginal_x = marginal(joint, 0)
g.ax_marg_x.plot(marginal_x.qs, marginal_x.ps)
marginal_y = marginal(joint, 1)
g.ax_marg_y.plot(marginal_y.ps, marginal_y.qs)
# + tags=[]
joint_plot(joint_posterior)
# + [markdown] tags=[]
# A `JointGrid` is a concise way to represent the joint and marginal distributions visually.
# -
# ## The Lincoln Index Problem
#
# In [an excellent blog post](http://www.johndcook.com/blog/2010/07/13/lincoln-index/), <NAME> wrote about the Lincoln index, which is a way to estimate the
# number of errors in a document (or program) by comparing results from
# two independent testers.
# Here's his presentation of the problem:
#
# > "Suppose you have a tester who finds 20 bugs in your program. You
# > want to estimate how many bugs are really in the program. You know
# > there are at least 20 bugs, and if you have supreme confidence in your
# > tester, you may suppose there are around 20 bugs. But maybe your
# > tester isn't very good. Maybe there are hundreds of bugs. How can you
# > have any idea how many bugs there are? There's no way to know with one
# > tester. But if you have two testers, you can get a good idea, even if
# > you don't know how skilled the testers are."
#
# Suppose the first tester finds 20 bugs, the second finds 15, and they
# find 3 in common; how can we estimate the number of bugs?
#
# This problem is similar to the Grizzly Bear problem, so I'll represent the data in the same way.
k10 = 20 - 3
k01 = 15 - 3
k11 = 3
# But in this case it is probably not reasonable to assume that the testers have the same probability of finding a bug.
# So I'll define two parameters, `p0` for the probability that the first tester finds a bug, and `p1` for the probability that the second tester finds a bug.
#
# I will continue to assume that the probabilities are independent, which is like assuming that all bugs are equally easy to find. That might not be a good assumption, but let's stick with it for now.
#
# As an example, suppose we know that the probabilities are 0.2 and 0.15.
p0, p1 = 0.2, 0.15
# We can compute the array of probabilities, `y`, like this:
def compute_probs(p0, p1):
"""Computes the probability for each of 4 categories."""
q0 = 1-p0
q1 = 1-p1
return [q0*q1, q0*p1, p0*q1, p0*p1]
y = compute_probs(p0, p1)
y
# With these probabilities, there is a
# 68% chance that neither tester finds the bug and a
# 3% chance that both do.
#
# Pretending that these probabilities are known, we can compute the posterior distribution for `N`.
# Here's a prior distribution that's uniform from 32 to 350 bugs.
qs = np.arange(32, 350, step=5)
prior_N = make_uniform(qs, name='N')
prior_N.head(3)
# I'll put the data in an array, with 0 as a place-keeper for the unknown value `k00`.
data = np.array([0, k01, k10, k11])
# And here are the likelihoods for each value of `N`, with `ps` as a constant.
# +
likelihood = prior_N.copy()
observed = data.sum()
x = data.copy()
for N in prior_N.qs:
x[0] = N - observed
likelihood[N] = multinomial.pmf(x, N, y)
# -
# We can compute the posterior in the usual way.
posterior_N = prior_N * likelihood
posterior_N.normalize()
# And here's what it looks like.
# + tags=[]
posterior_N.plot(color='C4')
decorate(xlabel='Number of bugs (N)',
ylabel='PMF',
title='Posterior marginal distribution of n with known p1, p2')
# + tags=[]
print(posterior_N.mean(),
posterior_N.credible_interval(0.9))
# -
# With the assumption that `p0` and `p1` are known to be `0.2` and `0.15`, the posterior mean is 102 with 90% credible interval (77, 127).
# But this result is based on the assumption that we know the probabilities, and we don't.
# ## Three-Parameter Model
#
# What we need is a model with three parameters: `N`, `p0`, and `p1`.
# We'll use `prior_N` again for the prior distribution of `N`, and here are the priors for `p0` and `p1`:
qs = np.linspace(0, 1, num=51)
prior_p0 = make_uniform(qs, name='p0')
prior_p1 = make_uniform(qs, name='p1')
# Now we have to assemble them into a joint prior with three dimensions.
# I'll start by putting the first two into a `DataFrame`.
joint2 = make_joint(prior_p0, prior_N)
joint2.shape
# Now I'll stack them, as in the previous example, and put the result in a `Pmf`.
joint2_pmf = Pmf(joint2.stack())
joint2_pmf.head(3)
# We can use `make_joint` again to add in the third parameter.
joint3 = make_joint(prior_p1, joint2_pmf)
joint3.shape
# The result is a `DataFrame` with values of `N` and `p0` in a `MultiIndex` that goes down the rows and values of `p1` in an index that goes across the columns.
# + tags=[]
joint3.head(3)
# -
# Now I'll apply `stack` again:
joint3_pmf = Pmf(joint3.stack())
joint3_pmf.head(3)
# The result is a `Pmf` with a three-column `MultiIndex` containing all possible triplets of parameters.
#
# The number of rows is the product of the number of values in all three priors, which is almost 170,000.
joint3_pmf.shape
# That's still small enough to be practical, but it will take longer to compute the likelihoods than in the previous examples.
#
# Here's the loop that computes the likelihoods; it's similar to the one in the previous section:
# +
likelihood = joint3_pmf.copy()
observed = data.sum()
x = data.copy()
for N, p0, p1 in joint3_pmf.index:
x[0] = N - observed
y = compute_probs(p0, p1)
likelihood[N, p0, p1] = multinomial.pmf(x, N, y)
# -
# We can compute the posterior in the usual way.
posterior_pmf = joint3_pmf * likelihood
posterior_pmf.normalize()
# Now, to extract the marginal distributions, we could unstack the joint posterior as we did in the previous section.
# But `Pmf` provides a version of `marginal` that works with a `Pmf` rather than a `DataFrame`.
# Here's how we use it to get the posterior distribution for `N`.
posterior_N = posterior_pmf.marginal(0)
# And here's what it looks like.
# + tags=[]
posterior_N.plot(color='C4')
decorate(xlabel='Number of bugs (N)',
ylabel='PDF',
title='Posterior marginal distributions of N')
# + tags=[]
posterior_N.mean()
# -
# The posterior mean is 105 bugs, which suggests that there are still many bugs the testers have not found.
#
# Here are the posteriors for `p0` and `p1`.
# + tags=[]
posterior_p1 = posterior_pmf.marginal(1)
posterior_p2 = posterior_pmf.marginal(2)
posterior_p1.plot(label='p1')
posterior_p2.plot(label='p2')
decorate(xlabel='Probability of finding a bug',
ylabel='PDF',
title='Posterior marginal distributions of p1 and p2')
# + tags=[]
posterior_p1.mean(), posterior_p1.credible_interval(0.9)
# + tags=[]
posterior_p2.mean(), posterior_p2.credible_interval(0.9)
# -
# Comparing the posterior distributions, the tester who found more bugs probably has a higher probability of finding bugs. The posterior means are about 23% and 18%. But the distributions overlap, so we should not be too sure.
# This is the first example we've seen with three parameters.
# As the number of parameters increases, the number of combinations increases quickly.
# The method we've been using so far, enumerating all possible combinations, becomes impractical if the number of parameters is more than 3 or 4.
#
# However there are other methods that can handle models with many more parameters, as we'll see in <<_MCMC>>.
# ## Summary
#
# The problems in this chapter are examples of [mark and recapture](https://en.wikipedia.org/wiki/Mark_and_recapture) experiments, which are used in ecology to estimate animal populations. They also have applications in engineering, as in the Lincoln index problem. And in the exercises you'll see that they are used in epidemiology, too.
#
# This chapter introduces two new probability distributions:
#
# * The hypergeometric distribution is a variation of the binomial distribution in which samples are drawn from the population without replacement.
#
# * The multinomial distribution is a generalization of the binomial distribution where there are more than two possible outcomes.
#
# Also in this chapter, we saw the first example of a model with three parameters. We'll see more in subsequent chapters.
# ## Exercises
# **Exercise:** [In an excellent paper](http://chao.stat.nthu.edu.tw/wordpress/paper/110.pdf), Anne Chao explains how mark and recapture experiments are used in epidemiology to estimate the prevalence of a disease in a human population based on multiple incomplete lists of cases.
#
# One of the examples in that paper is a study "to estimate the number of people who were infected by hepatitis in an outbreak that occurred in and around a college in northern Taiwan from April to July 1995."
#
# Three lists of cases were available:
#
# 1. 135 cases identified using a serum test.
#
# 2. 122 cases reported by local hospitals.
#
# 3. 126 cases reported on questionnaires collected by epidemiologists.
#
# In this exercise, we'll use only the first two lists; in the next exercise we'll bring in the third list.
#
# Make a joint prior and update it using this data, then compute the posterior mean of `N` and a 90% credible interval.
# + [markdown] tags=[]
# The following array contains 0 as a place-holder for the unknown value of `k00`, followed by known values of `k01`, `k10`, and `k11`.
# + tags=[]
data2 = np.array([0, 73, 86, 49])
# + [markdown] tags=[]
# These data indicate that there are 73 cases on the second list that are not on the first, 86 cases on the first list that are not on the second, and 49 cases on both lists.
#
# To keep things simple, we'll assume that each case has the same probability of appearing on each list. So we'll use a two-parameter model where `N` is the total number of cases and `p` is the probability that any case appears on any list.
#
# Here are priors you can start with (but feel free to modify them).
# + tags=[]
qs = np.arange(200, 500, step=5)
prior_N = make_uniform(qs, name='N')
prior_N.head(3)
# + tags=[]
qs = np.linspace(0, 0.98, num=50)
prior_p = make_uniform(qs, name='p')
prior_p.head(3)
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
# **Exercise:** Now let's do the version of the problem with all three lists. Here's the data from Chou's paper:
#
# ```
# Hepatitis A virus list
# P Q E Data
# 1 1 1 k111 =28
# 1 1 0 k110 =21
# 1 0 1 k101 =17
# 1 0 0 k100 =69
# 0 1 1 k011 =18
# 0 1 0 k010 =55
# 0 0 1 k001 =63
# 0 0 0 k000 =??
# ```
#
# Write a loop that computes the likelihood of the data for each pair of parameters, then update the prior and compute the posterior mean of `N`. How does it compare to the results using only the first two lists?
# + [markdown] tags=[]
# Here's the data in a NumPy array (in reverse order).
# + tags=[]
data3 = np.array([0, 63, 55, 18, 69, 17, 21, 28])
# + [markdown] tags=[]
# Again, the first value is a place-keeper for the unknown `k000`. The second value is `k001`, which means there are 63 cases that appear on the third list but not the first two. And the last value is `k111`, which means there are 28 cases that appear on all three lists.
#
# In the two-list version of the problem we computed `ps` by enumerating the combinations of `p` and `q`.
# + tags=[]
q = 1-p
ps = [q*q, q*p, p*q, p*p]
# + [markdown] tags=[]
# We could do the same thing for the three-list version, computing the probability for each of the eight categories. But we can generalize it by recognizing that we are computing the cartesian product of `p` and `q`, repeated once for each list.
#
# And we can use the following function (based on [this StackOverflow answer](https://stackoverflow.com/questions/58242078/cartesian-product-of-arbitrary-lists-in-pandas/58242079#58242079)) to compute Cartesian products:
# + tags=[]
def cartesian_product(*args, **options):
"""Cartesian product of sequences.
args: any number of sequences
options: passes to `MultiIndex.from_product`
returns: DataFrame with one column per sequence
"""
index = pd.MultiIndex.from_product(args, **options)
return pd.DataFrame(index=index).reset_index()
# + [markdown] tags=[]
# Here's an example with `p=0.2`:
# + tags=[]
p = 0.2
t = (1-p, p)
df = cartesian_product(t, t, t)
df
# + [markdown] tags=[]
# To compute the probability for each category, we take the product across the columns:
# + tags=[]
y = df.prod(axis=1)
y
# + [markdown] tags=[]
# Now you finish it off from there.
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# +
# Solution goes here
# -
| notebooks/chap15.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sampath11/DS-Unit-1-Sprint-1-Dealing-With-Data/blob/master/module1-afirstlookatdata/LS_DSPT3_111_A_First_Look_at_Data.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="Okfr_uhwhS1X" colab_type="text"
# # Lambda School Data Science - A First Look at Data
#
#
# + [markdown] id="9dtJETFRhnOG" colab_type="text"
# ## Lecture - let's explore Python DS libraries and examples!
#
# The Python Data Science ecosystem is huge. You've seen some of the big pieces - pandas, scikit-learn, matplotlib. What parts do you want to see more of?
# + id="WiBkgmPJhmhE" colab_type="code" colab={}
## Trying out different Python libraries
import numpy as np
# + id="mS9wu2_U5E0v" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="be7b1fbb-2876-42c2-ef5b-d256d33f36f4"
np.random.randint(0, 10, size=10)
# + id="ytNnHx4_5R3G" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="7dd53993-01d6-405d-fe61-105a2f1c3b2e"
## Change vars
np.random.randint ( 0, 100, size=10)
# + id="srrpvro95Zhf" colab_type="code" colab={}
## Second Library
import matplotlib.pyplot as plt
# + id="8D5xY93j5oSm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="f423a2a2-625c-48ee-9777-508b199a86e4"
x = [1, 2, 3, 4, 5]
y=[2, 4, 6, 8, 10]
print ( x, y)
# + id="2NS_p_h053iH" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="b75cbae4-48ef-483c-c87f-5646f5e0a28e"
# now plot
plt.scatter ( x, y)
# + id="Vte3M6WS59GG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="6386271f-ccfb-43ee-d7c2-d119c482cd11"
plt.scatter ( x, y, color='r')
# + id="jsTrPS7V6ILa" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 287} outputId="d9b516a5-1b09-46f1-e992-0b68a50a19d2"
# Now plot the same thing
plt.plot (x, y, color='r')
# + [markdown] id="lOqaPds9huME" colab_type="text"
# ## Assignment - now it's your turn
#
# Pick at least one Python DS library, and using documentation/examples reproduce in this notebook something cool. It's OK if you don't fully understand it or get it 100% working, but do put in effort and look things up.
# + id="TGUS79cOhPWj" colab_type="code" colab={}
# TODO - your code here
# Use what we did live in lecture as an example
import pandas as pd
# + id="JODJJNoV7xbo" colab_type="code" colab={}
df = pd.DataFrame ( {'first_col':x , 'second_col':y})
# + id="bu3Rg8Ai8H8H" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 203} outputId="4bba49ce-93f1-4eda-b974-6212fb84b698"
df
# + id="QvjaeXNX9FmL" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 123} outputId="8b304c47-84dd-4ef4-d659-084d5090c224"
df['second_col']
# + id="5PXcapX89T8U" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="9c0066e1-032d-4383-9c84-01d342b3a03d"
df.shape
# + id="rqPUkva79aLx" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="832c41a6-5270-44d0-a8a6-d418e1568581"
type (df)
# + id="ZJRcJtBq9kgc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="a0140640-6a25-46e5-84f9-0194e3622d95"
type (df['second_col'])
# + id="1YPUF7-Z_iZr" colab_type="code" colab={}
z= ['a', 'b', 'c', 'd', 'e']
df['new_col'] = z
# + id="Xwl_-fkuAira" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 203} outputId="4d537996-c015-4561-f201-c9bc7aca5706"
df
# + id="vlZN6z0OAm5b" colab_type="code" colab={}
df['calc_col'] = df['second_col'] > 5
# + id="rJMGgsNSA7Xi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 203} outputId="ff5935a9-3551-4b7c-ff01-db05cf74cfb1"
df
# + id="YoNpYdxjBJT0" colab_type="code" colab={}
# + [markdown] id="7-yMpG-wBUH-" colab_type="text"
# ## Assignment Answers
# 1. Describe in a paragraph of text what you did and why, as if you were writing an email to somebody interested but nontechnical.
#
# ## Used example provided in class and tried to create additional columns in array but with data type of char in addition to int and boolean
#
# 2. What was the most challenging part of what you did?
#
# ## It was not that challenging
#
# 3. What was the most interesting thing you learned?
#
# ## Python libraries are powerful, syntax may be simple but usage and value from them may be very powerful
#
# 4. What area would you like to explore with more time?
#
# ## continue to dig deep into pandas, numpy, pyplot in addition to what was mentioned in class - plotly and bokeh
# + [markdown] id="Y_Y-lvrWBKbQ" colab_type="text"
# ## Assignment Answers
# + [markdown] id="BT9gdS7viJZa" colab_type="text"
# ### Assignment questions
#
# After you've worked on some code, answer the following questions in this text block:
#
# 1. Describe in a paragraph of text what you did and why, as if you were writing an email to somebody interested but nontechnical.
#
# 2. What was the most challenging part of what you did?
#
# 3. What was the most interesting thing you learned?
#
# 4. What area would you like to explore with more time?
#
#
#
# + [markdown] id="_XXg2crAipwP" colab_type="text"
# ## Stretch goals and resources
#
# Following are *optional* things for you to take a look at. Focus on the above assignment first, and make sure to commit and push your changes to GitHub (and since this is the first assignment of the sprint, open a PR as well).
#
# - [pandas documentation](https://pandas.pydata.org/pandas-docs/stable/)
# - [scikit-learn documentation](http://scikit-learn.org/stable/documentation.html)
# - [matplotlib documentation](https://matplotlib.org/contents.html)
# - [Awesome Data Science](https://github.com/bulutyazilim/awesome-datascience) - a list of many types of DS resources
#
# Stretch goals:
#
# - Find and read blogs, walkthroughs, and other examples of people working through cool things with data science - and share with your classmates!
# - Write a blog post (Medium is a popular place to publish) introducing yourself as somebody learning data science, and talking about what you've learned already and what you're excited to learn more about.
| module1-afirstlookatdata/LS_DSPT3_111_A_First_Look_at_Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pyspark import SparkConf, SparkContext
conf = SparkConf().setMaster("yarn")
sc = SparkContext(conf=conf)
sc
try:
text
except NameError:
text = sc.textFile("hdfs://name:9000/csv/student.csv")
first = text.first()
first
data = text.filter(lambda x: x != first)
data.first()
data = data.map(lambda x: x.split(","))
data.take(1)
data.map(lambda x: x[0:3] + x[3:5] + x[19:]).take(3)
first
def to_int(x):
for i, j in enumerate(x[3:-2]):
x[i + 3] = int(j)
return x
def student_filter(school=None, div=None, level=None, grade=None, gender=None, loc=None, year=None):
filter_data = data
if school:
filter_data = filter_data.filter(lambda x: x[0] == school)
if div:
filter_data = filter_data.filter(lambda x: x[1] == div)
if level:
filter_data = filter_data.filter(lambda x: x[2] == level)
if loc:
filter_data = filter_data.filter(lambda x: x[19] == loc)
if year:
filter_data = filter_data.filter(lambda x: x[20] == year)
if grade:
if grade == "一年級":
filter_data = filter_data.map(lambda x: x[0:5] + x[19:])
elif grade == "二年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[5:7] + x[19:])
elif grade == "三年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[7:9] + x[19:])
elif grade == "四年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[9:11] + x[19:])
elif grade == "五年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[11:13] + x[19:])
elif grade == "六年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[13:15] + x[19:])
elif grade == "七年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[15:17] + x[19:])
elif grade == "延修生":
filter_data = filter_data.map(lambda x: x[0:3] + x[17:])
if gender == "男生":
filter_data = filter_data.map(lambda x: x[0:4] + x[5:])
elif gender == "女生":
filter_data = filter_data.map(lambda x: x[0:3] + x[4:])
if grade == None and gender:
if gender == "男生":
filter_data = filter_data.map(lambda x: x[0:3] + x[3:18:2] + x[19:])
elif gender == "女生":
filter_data = filter_data.map(lambda x: x[0:3] + x[4:19:2] + x[19:])
filter_data = filter_data.map(to_int)
return filter_data.map(lambda x: x[0:3] + [sum(x[3:-2])] + x[-2:]).collect()
def initializaion():
textRDD = sc.textFile("hdfs://name:9000/csv/student.csv")
text1 = textRDD.collect()
textList = []
for data1 in text1:
textList.append(data1.split(','))
tableRDD = sc.parallelize(textList)
return tableRDD
def filtering(oriRDD, schoolName=None, division=None, level=None, grade=None, gender=None, county=None, year=None):
filteredRDD = oriRDD
if schoolName:
filteredRDD = filteredRDD.filter(lambda x: x[0] == schoolName or x[0] == "學校名稱")
if division:
filteredRDD = filteredRDD.filter(lambda x: x[1] == division or x[1] == "日間∕進修別")
if level:
filteredRDD = filteredRDD.filter(lambda x: x[2] == level or x[2] == "等級別")
if county:
filteredRDD = filteredRDD.filter(lambda x: x[19] == county or x[19] == "縣市名稱")
if year:
filteredRDD = filteredRDD.filter(lambda x: x[20] == year or x[20] == "學年度")
tList = filteredRDD.collect()
if level and grade:
for t in tList:
if grade == '一年級':
del t[5:19]
elif grade == '二年級':
del t[3:5]
del t[5:17]
elif grade == '三年級':
del t[3:7]
del t[5:15]
elif grade == '四年級':
del t[3:9]
del t[5:13]
elif grade == '五年級':
del t[3:11]
del t[5:11]
elif grade == '六年級':
del t[3:13]
del t[5:9]
elif grade == '七年級':
del t[3:15]
del t[5:7]
elif grade == '延修生':
del t[3:17]
if gender == '男生':
del t[4]
elif gender == '女生':
del t[3]
elif level and grade == None and gender:
for t in tList:
if gender == '男生':
del t[4:19:2]
elif gender == '女生':
del t[3:18:2]
for t in tList[1:]:
sumN = 0
for n in t[3:-2]:
sumN = sumN + int(n)
t[3] = str(sumN)
del t[4:-2]
tList[0][3] = '學生人數'
del tList[0][4:-2]
sumN = 0
for t in tList[1:]:
sumN += int(t[3])
return {'total': sumN, 'data': tList}
dic = filtering(initializaion(), schoolName='國立臺灣大學', level='碩士', year='105')
for s in dic['data']:
print(s)
print(dic['total'])
dic['data']
student_filter(school='國立臺灣大學', gender='男生')
student_filter(school='國立臺灣大學', grade='一年級', gender='男生')
l = ['2', '3', '4']
sum(int(l[0:2]))
l1 = ["1", "2", "3"]
year = 123
for i in [year, year, year]:
if i == 123:
i = 0
print(year)
def student_filter_v1(school=None, div=None, level=None, grade=None, gender=None, loc=None, year=None):
filter_data = data
if school:
filter_data = filter_data.filter(lambda x: x[0] == school)
if div:
filter_data = filter_data.filter(lambda x: x[1] == div)
if level:
filter_data = filter_data.filter(lambda x: x[2] == level)
if loc:
filter_data = filter_data.filter(lambda x: x[19] == loc)
if year:
filter_data = filter_data.filter(lambda x: x[20] == year)
if level and grade:
if grade == "一年級":
filter_data = filter_data.map(lambda x: x[0:5] + x[19:])
elif grade == "二年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[5:7] + x[19:])
elif grade == "三年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[7:9] + x[19:])
elif grade == "四年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[9:11] + x[19:])
elif grade == "五年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[11:13] + x[19:])
elif grade == "六年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[13:15] + x[19:])
elif grade == "七年級":
filter_data = filter_data.map(lambda x: x[0:3] + x[15:17] + x[19:])
elif grade == "延修生":
filter_data = filter_data.map(lambda x: x[0:3] + x[17:])
if gender == "男生":
filter_data = filter_data.map(lambda x: x[0:4] + x[5:])
elif gender == "女生":
filter_data = filter_data.map(lambda x: x[0:3] + x[4:])
elif level and grade == None and gender:
if gender == "男生":
filter_data = filter_data.map(lambda x: x[0:3] + x[3:18:2] + x[19:])
elif gender == "女生":
filter_data = filter_data.map(lambda x: x[0:3] + x[4:19:2] + x[19:])
filter_data = filter_data.map(to_int)
return filter_data.map(lambda x: x[0:3] + [sum(x[3:-2])] + x[-2:]).collect()
| notebook/.ipynb_checkpoints/spark-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Code to generate Figure 2
# +
import pandas as pd
import numpy as np
from sklearn.model_selection import train_test_split
from sklearn import preprocessing
from utils.svm import LinearSupportVectorMachine_noOffset
from utils.RandFourier import RandomFourier
from utils.functions import *
import matplotlib
import matplotlib.pyplot as plt
figwidth = 4
figheight = 4
# -
# ## Generate data
# +
SS = 250
n=2*SS
sigma = .1
mean1 = [1,1]
mean2 = [0,0]
cov1 = [[sigma,0],[0,sigma]]
cov2 = [[sigma,0],[0,sigma]]
s1 = []
s12 = []
s2 = []
s22 = []
import matplotlib.pyplot as plt
np.random.seed(seed=1)
x1, y1 = np.random.multivariate_normal(mean1, cov1, SS).T
x2, y2 = np.random.multivariate_normal(mean2, cov2, SS).T
s1.append([x1,y1])
s2.append([x2,y2])
X = []
Y = []
for i in range(SS):
X.append([s1[0][0][i], s1[0][1][i]])
Y.append(-1)
X.append([s2[0][0][i], s2[0][1][i]])
Y.append(1)
X=np.array(X)
X_train, X_test, ytrain, ytest = train_test_split(X, Y,test_size=0.3, random_state=40)
y_train = np.asarray(ytrain)
y_test = np.asarray(ytest)
K = np.outer(X_train,X_train)
kappa = np.sqrt(K.max())
# Plot
fig, ax = plt.subplots(figsize=[figwidth, figheight], dpi=100)
ax.scatter(x=X_train[y_train==1,0], y=X_train[y_train==1,1], s=8, c='tab:blue', marker='o')
ax.scatter(x=X_train[y_train==-1,0], y=X_train[y_train==-1,1], s=5, c='tab:red', marker='s')
ax.set(xlabel='', ylabel='', title='')
ax.set(xlabel='$x_1$', ylabel='$x_2$', title='')
ax.grid()
plt.show()
# -
# ## Find a prototype for each class
# +
# prototype[0]: class 1
# prototype[1]: class -1
prototype = [np.mean(X_train[y_train == 1], axis=0), np.mean(X_train[y_train == -1], axis=0)]
fig, ax = plt.subplots(figsize=[figwidth, figheight], dpi=100)
ax.scatter(x=X_train[y_train==1,0], y=X_train[y_train==1,1], s=8, c='tab:blue', marker='o')
ax.scatter(x=X_train[y_train==-1,0], y=X_train[y_train==-1,1], s=5, c='tab:red', marker='s')
ax.plot(prototype[0][0], prototype[0][1],'kd')
ax.plot(prototype[1][0], prototype[1][1],'kd')
ax.set(xlabel='', ylabel='', title='')
ax.set(xlabel='$x_1$', ylabel='$x_2$', title='')
ax.grid()
plt.show()
# -
# ## Train SVM Classifier
# +
C = np.sqrt(n)
X_train_b = add_bias(X_train)
X_test_b = add_bias(X_test)
SVM = LinearSupportVectorMachine_noOffset(C=C)
lagr_multipliers, idx_support_vectors, support_vectors, support_vector_labels = SVM.fit(X_train_b, y_train)
# weight vector
w = np.dot(lagr_multipliers*support_vector_labels,support_vectors)
evaluation = np.sign(SVM.predict(X_test_b).flatten()) == y_test
print("Classification accuracy:", sum(evaluation)/len(evaluation))
# +
F = 2
proba = 0.9
beta = 5
# calculate noise scale: lambda
noise_lambda = 4*C*kappa*np.sqrt(F+1)/(beta*n)
np.random.seed(seed=10)
mu = np.random.laplace(loc=0.0, scale=noise_lambda, size=(1,F+1))
w_tilde = w + mu.flatten()
# +
x_min, x_max, y_min, y_max = -1,2,-1,2
res = 500j
XX, YY = np.mgrid[x_min:x_max:res, y_min:y_max:res]
X_grid = np.c_[XX.ravel(), YY.ravel()]
X_grid_b = add_bias(X_grid)
Z = SVM.predict(X_grid_b)
Z = Z.reshape(XX.shape)
Z_p = SVM.predict(X_grid_b, mu)
Z_p = Z_p.reshape(XX.shape)
# +
# Plot
fig, ax = plt.subplots(figsize=[figwidth, figheight], dpi=100)
ax.scatter(x=X_train[y_train==1,0], y=X_train[y_train==1,1], s=6, c='tab:blue', marker='o')
ax.scatter(x=X_train[y_train==-1,0], y=X_train[y_train==-1,1], s=6, c='tab:red', marker='s')
plt.contour(XX, YY, Z, colors=['k'], linestyles=['-'], levels=[0])
plt.contour(XX, YY, Z_p, colors=['k'], linestyles=['--'], levels=[0])
ax.plot(prototype[0][0], prototype[0][1],'kd')
ax.plot(prototype[1][0], prototype[1][1],'kd')
ax.set(xlabel='', ylabel='', title='')
ax.grid()
plt.show()
# -
# ## Select instance and prototype
#
# * Select test data instance whose classification we want to explain
# * Select prototype of opposite class label than that of the selected instance
# +
#np.random.seed(seed=1)
#idx_selected = np.random.randint(0,len(X_test))
idx_selected = 1
instance = [X_test[idx_selected]]
instance_b = [X_test_b[idx_selected]]
prediction_instance = np.sign(SVM.predict(instance_b, noise=0).flatten())
print("Index of selected test data:", idx_selected, "-- Predicted class:", prediction_instance, "-- True class label:", y_test[idx_selected])
# choose prototype of different class than instance
selected_prototype = prototype[np.where(prediction_instance != [1,-1])[0][0]]
# -
# ## Compute explanations
# +
b_robust = prediction_instance*w_tilde / (noise_lambda*np.sqrt(2)*np.log(2*(1-proba)))
b_non_robust = prediction_instance*w_tilde
b_opt = prediction_instance*w
explanation_robust = socp_opt(instance, F, b_robust)
explanation_non_robust = counterfactual_explanation_linear(instance, F, b_non_robust)
explanation_opt = counterfactual_explanation_linear(instance, F, b_opt)
# +
# Plot
fig, ax = plt.subplots(figsize=[figwidth, figwidth], dpi=100)
ax.scatter(x=X_train[y_train==1,0], y=X_train[y_train==1,1], s=15, edgecolors='tab:green', marker=".", facecolors='tab:green')
ax.scatter(x=X_train[y_train==-1,0], y=X_train[y_train==-1,1], s=30, edgecolors='tab:grey', marker='+', facecolors='tab:grey')
plt.contour(XX, YY, Z, colors=['k'], linestyles=['-'], levels=[0], linewidths=1.5)
plt.contour(XX, YY, Z_p, colors=['k'], linestyles=['--'], levels=[0], linewidths=1.5)
ax.plot(instance[0][0],instance[0][1], marker='o', markersize=6, ls="", c='k',mfc="k", label="Instance")
ax.plot(explanation_robust[0],explanation_robust[1],markersize=8, ls="", marker='*', c='tab:red', mfc="tab:red", label="Robust explanation")
ax.plot(explanation_non_robust[0],explanation_non_robust[1], markersize=6, ls="", marker='d', c='k', mfc="w",label="Non-robust explanation")
ax.plot(explanation_opt[0],explanation_opt[1], markersize=6, ls="", marker='s', c='k', mfc="w",label="Optimal explanation")
ax.set(xlabel='$x_1$', ylabel='$x_2$', title='')
ax.grid()
ti = np.arange(-.6,1.3,step=0.2)
ax.set_xticks(ti)
ax.set_xlim(-.6,1.2)
ax.set_ylim(-.4,1.4)
ax.legend(loc='lower right',fontsize=8)
plt.show()
# -
| Illustration Linear SVM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
pwd
# Dependencies and Setup
import pandas as pd
import pathlib
import csv
import scipy.stats as st
import numpy as np
import os
# Create a path
path = pathlib.Path("../raw_data/zillow_dataset")
# Loop over the files example of path: os.listdir(../raw_data/zillow_dataset)
files = [path/file for file in os.listdir(path)]
files
# Read each csv
dfs = [pd.read_csv(file).assign(houseTypes=str(file).split("_")[-2]) for file in files if ".csv" in str(file)]
# Combine each csv as one
z_housetypes_df = pd.concat(dfs)
z_housetypes_df
# remove all states that are not CA (California)
only_CA_df = z_housetypes_df.loc[(z_housetypes_df['State'] == 'CA' )]
only_CA_df
# Delete extraneous columns
delete_extraneous_columns = only_CA_df.drop(columns= [
"SizeRank"
, "RegionID"
, "RegionType"
, "StateName"
, "State"
, "City"
, "Metro"
])
delete_extraneous_columns.head()
# Remove the county from each name
delete_extraneous_columns["CountyName"] = delete_extraneous_columns["CountyName"].str.replace(" County", "")
delete_extraneous_columns
# Use melt() function to make the dates as columns
melt_z_df = delete_extraneous_columns.melt(var_name = "dates", value_name="prices", id_vars=['RegionName', 'CountyName', 'houseTypes']).astype({"dates":"datetime64"})
melt_z_df
# Delete extraneous columns
d_melt_z_df = melt_z_df.drop(columns= [
"dates"
])
d_melt_z_df
# Rename the columns
rename_columns = d_melt_z_df.rename(columns={"RegionName":"zipCode", "CountyName": "countyName"})
rename_columns
# Delete NaN
delete_nan = rename_columns.dropna(inplace = False)
pd.DataFrame(delete_nan)
# +
# assert [f'Number is {"NaN"}' for numbers in delete_nan["prices"]]
# +
# assert [f for numbers in {delete_nan["prices"] != "NaN"}]
# -
# Create final variable for last table
final_z_df = delete_nan
final_z_df
# Delete extraneous columns
final_df = final_z_df.drop(columns= [
"zipCode"
])
final_df
# Final cleaned csv to be used
final_df.to_csv("../cleaned_data/final_zillow_df.csv")
# Group by County
by_county_final_df = final_df.groupby(['countyName', 'houseTypes']).mean() #astype({"countyName":str,"hospitalOverallRating":int})
pd.DataFrame(by_county_final_df)
# Final cleaned csv to be used
by_county_final_df.to_csv("../cleaned_data/by_county_zillow_pricing_df.csv")
| Data/cleaning_data_wrbk/zillow_merged_cleaning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ## Examples of querying the database
#
# Most users will access teh oil datbase via the WebAPI or via PYGNOME.
#
# However, one can work with the database directly via its python interacea s well.
#
# Here are some examples.
import oil_library
from pprint import pprint
# getting an oil object by name:
#oil_library.get_oil('jp-8')
oil = oil_library.get_oil('ABU SAFAH, ARAMCO')
print oil
print type(oil)
# An oil object has all sorts of properties you might be interested in
print "API is:", oil.api
print "This oil's name is: {}\nADIOS ID: {}\nIt has and API of {}".format(
oil.name, oil.adios_oil_id, oil.api)
print "Here are all its attributes:"
pprint(vars(oil))
# +
#Getting an oil by the ADIOS ID:
oil = oil_library.get_oil('AD00010')
# -
oil
# what if there are duplicate names?
# you get an exeption with a nice error message
try:
oil = oil_library.get_oil('JP-8')
except Exception as err:
print err
oil = oil_library.get_oil('AD02434')
print oil
# ## Working with the DB directly
#
# There are currently on a few simiple querries built in to the oil_libary API. But you can work with the DB directly to do any querying you might want
#
# The system is built on the SQAlchemy Object-relational mapper:
#
# https://www.sqlalchemy.org/
#
# ### Accessing the database object
#
# You can get the database session object with:
#
# `session = oil_library._get_db_session()`
#
# Then you can use the session object to query the datbase:
#
#
#
#
#
# get the datbase session
session = oil_library._get_db_session()
# get the "oil" table -- where most of the work is done..
Oil = oil_library.models.Oil
# do a query:
result = session.query(Oil).filter(Oil.name == 'JP-8')
# see all the results
result.all()
# iterate through the results
[oil.adios_oil_id for oil in result]
oil_library.get_oil(' ad02433 ')
| documentation/notebooks/QueryingTheDatabase.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
x = np.linspace(0., 3., 30)
y = np.linspace(0., 1., 20)
z = np.linspace(0., .5, 10)
xx, yy, zz = np.meshgrid(x,y,z, indexing = 'xy')
print('x shape = ',x.shape)
print('xx shape = ',xx.shape)
x1 = xx.shape[0]
x2 = xx.shape[1]
x3 = xx.shape[2]
print(x1,x2,x3)
xflat = np.reshape(xx,-1)
print(xflat.shape)
newxx = np.reshape(xflat,(x1,x2,x3))
print(newxx.shape)
(xx==newxx).all()
| docs/notebooks/example_array_reshaping.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## see test_query.ipynb
import os
import sys
import subprocess
ROOT_PATH=os.getcwd()
manage = os.path.join(ROOT_PATH , "manage.py")
print(subprocess.check_output([sys.executable, manage, "makemigrations","--noinput"]))
print(subprocess.check_output([sys.executable, manage, "migrate","--noinput"]))
# +
import pydot
import time
timestr = time.strftime("%y%m%d_%H%M%S")
dotstr=subprocess.check_output([sys.executable, manage, "graph_models", "-a"])
# with open("models.dot","w") as f:
# f.write(dotstr)
(graph,) = pydot.graph_from_dot_file('models.dot')
graph = pydot.graph_from_dot_data(dotstr)
graph.write_png(timestr + '.png')
#open the png
# -
from IPython.display import display, Markdown, Latex
md='''<img src="%s",width=60,height=60>'''%(timestr + '.png')
display(Markdown(md))
| beta_python3/05_gen_models.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
arr = np.array([[1,2,3], [4,5,6]], dtype=np.int64)
# ### inspect general information of an array
print(np.info(arr))
# ### inspect the data type of an array
print(arr.dtype)
# ### inspect the dimension of an array
print(arr.shape)
# ### inspect length of an array
print(len(arr))
# ### inspect the number of dimensions of an array
print(arr.ndim)
# ### inspect the number of elements in an array
print(arr.size)
# ### inspect the number of bytes of each element in an array
print(arr.itemsize)
# ### inspect the memory size of an array (in byte)
# arr.nbytes = arr.size * arr.itemsize
print(arr.nbytes)
| 2. Inspect an Array.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Exploring
import pandas
from pandas import DataFrame
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
data = pandas.read_csv('cost_revenue_clean.csv')
data.describe()
X = DataFrame(data, columns = ['production_budget_usd'])
y = DataFrame(data, columns = ['worldwide_gross_usd'])
X.describe()
y.describe()
# # Data Vizualisation
plt.figure(figsize=(10,6))
plt.scatter(X, y, alpha=0.3)
plt.title('Film Cost vs Global Revenue')
plt.xlabel('Production Budget $')
plt.ylabel('Worldwide Gross $')
plt.ylim(0, 3000000000)
plt.xlim(0, 450000000)
plt.show()
# # Model Training
regression = LinearRegression()
regression.fit(X,y)
print('Slope coefficient:')
regression.coef_
print('Intercept :')
regression.intercept_
# +
plt.figure(figsize=(10,6))
plt.scatter(X, y, alpha=0.3)
# Adding the regression line here:
plt.plot(X, regression.predict(X), color = 'red', linewidth = 3)
plt.title('Film Cost vs Global Revenue')
plt.xlabel('Production Budget $')
plt.ylabel('Worldwide Gross $')
plt.ylim(0, 3000000000)
plt.xlim(0, 450000000)
plt.show()
# -
# ## Model Score
#Getting r square from Regression
regression.score(X, y)
| movie-box-office-revenue-prediction.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] deletable=true editable=true
# # BigQuery
# + [markdown] deletable=true editable=true
# ## 사전작업
# 1. [Google 로그인](https://accounts.google.com/Login)
# 2. [Project 생성 및 선택](https://console.cloud.google.com/project?_ga=1.105140352.267165872.1487136809)
#
# 단, 처음에 체험판 등록하면 300$ 제공
#
# 3. [BigQuery API 등록](https://console.cloud.google.com/flows/enableapi?apiid=bigquery&_ga=1.139222832.267165872.1487136809)
#
# + [markdown] deletable=true editable=true
# 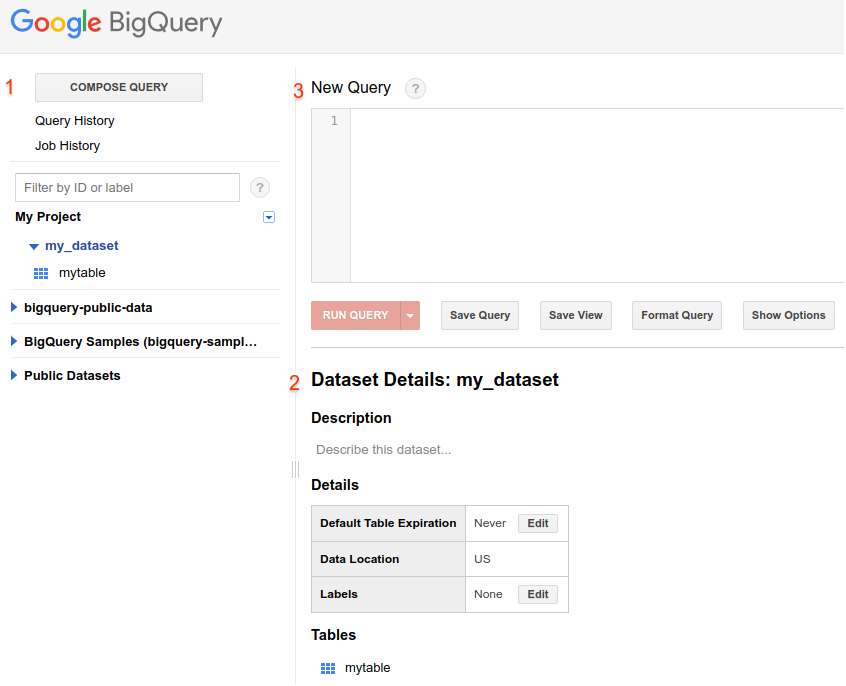
# + [markdown] deletable=true editable=true
# ## Left Side( the navigation bar )
# - COMPOSE QUERY : 쿼리 생성
# - Query History
# - Job History
# * 의문점 : Job은 뭐라고 정의해야할까-
# - project명
# - Public Dataset : 샘플 데이터로 실험 가능
#
# ## Right Side( Query Editor )
# - SQL Query 문 작성
# - format Query를 누르면 조금 더 형식을 맞춤
#
#
# ## Keyboard shortcuts
# | Windows/Linux | Mac | Action |
# | :---: | :---: | :---: |
# | Ctrl + Space | Ctrl + Space | If no query is open: compose new query. If query editor is open: autocomplete current word. |
# | Ctrl + Enter | Cmd + Enter | Run current query. |
# | Tab | Tab | Autocomplete current word. |
# | Ctrl | Cmd | Highlight table names. |
# | Ctrl + click on table name | Cmd + click on table name | Open table schema. |
# | Ctrl + E | Cmd + E | Run query from selection. |
# | Ctrl + / | Cmd + / | Comment current or selected line(s). |
# | Ctrl + Shift + F | Cmd + Shift + F | Format query. |
# + [markdown] deletable=true editable=true
# # BigQuery 설명
#
# - 관계형, noSQL도 아님..! 가까운 것을 찾으라고 하면 NoSQL과 유사하다고 보면 됨
# - 맵리듀스도 아님
# - 오픈소스도 아니라는 점-!
# - 데이터셋 - 테이블 - 스키마 구조
#
# + [markdown] deletable=true editable=true
# # 본격 BigQuery
#
# - [BigQuery 설명 페이지](https://developers.google.com/bigquery/docs/query-reference?hl=ko#having-)
# - [BigQuery Web UI](https://bigquery.cloud.google.com/welcome)
#
# - COMPOSE QUERY 클릭
#
# ~~~
# #standardSQL
# SELECT
# weight_pounds, state, year, gestation_weeks
# FROM
# `bigquery-public-data.samples.natality`
# ORDER BY weight_pounds DESC LIMIT 10;
# ~~~
#
#
# - Public Datasets을 클릭하면 각종 데이터를 볼 수 있음
# + [markdown] deletable=true editable=true
# ## Data load(in local)
# [baby names data](http://www.ssa.gov/OACT/babynames/names.zip)
#
# - in Web UI, create new dataset 클릭
# - 이름을 설정한 후, create new table 클릭
# - 나머지 설정!
# ~~~
# SELECT
# name, count
# FROM
# babynames.names_2014
# WHERE
# gender = 'M'
# ORDER BY count DESC LIMIT 5;
# ~~~
# + [markdown] deletable=true editable=true
# # Reference
# [Quickstart Using the Web UI](https://cloud.google.com/bigquery/quickstart-web-ui)
#
# [빅쿼리 공식문서](https://developers.google.com/bigquery/docs/query-reference?hl=ko#having-)
# + [markdown] deletable=true editable=true
# # Word
# - incurrying : 초래하는
# + deletable=true editable=true
| Google_Cloud_Platform/02. BigQuery.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="a1g6vLUtt9I9"
import numpy as np
import numba
from numba import jit , njit
# + id="8N0j1DSGt9I_"
# Global Config Variables
n0 = 1000 # number of p=0 points in metric space
V = n0 # Threshold for p=0
K = 10# No of clusters
A= 5# No of attributes
iterations = 80 # maximum iteration in clustering
runs =120
# + id="l4AI4R6Nt9JA"
import numpy as np
import os
import sys
import pandas
import random
import pandas as pd
import numpy as np
from matplotlib import pyplot as plt
from math import log2
from sklearn import preprocessing
import sys
import timeit
# + id="kqb5faS9t9JA"
import datetime
import json
import random
from collections import defaultdict
import matplotlib.patches as mpatches
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
from matplotlib import cm
#from pyclustering.cluster.kmedians import kmedians
from scipy.spatial.distance import pdist, squareform
from sklearn.cluster import KMeans
from sklearn.preprocessing import StandardScaler
from sklearn import preprocessing
import os
# + id="rlaUUedgt9JB" outputId="787eb8a2-89ec-4014-f32b-e37db527ce82"
from scipy.stats import norm
from sklearn.cluster import KMeans
# !pip install scikit-learn-extra
from sklearn.metrics import pairwise_distances
from itertools import permutations
# + id="MqgZi60jt9JD" outputId="12da7f02-5af4-42b3-e3fd-15010b3444ff"
print(os.getcwd())
def load_Adult(data_dir=''):
data_dir = data_dir
_path = 'adult_p.csv'
data_path = os.path.join(data_dir, _path)
K = 10
df = pandas.read_csv(data_path, sep=',')
#print(df.head())
#print(len(df))
return df
load_Adult()
# + id="4GF-oe8vt9JD" outputId="1d0a599f-af46-4d6b-a99d-0e21c9f2df45"
df=load_Adult()
df= df.round(decimals=5)
print(len(df))
df = df.dropna()
print(len(df))
#df['type'] = df['type']-1
typ = df['gender'].values
#print(len(typ))
#print(df.head(10))
c1 = np.count_nonzero(typ == 0)
c2 = np.count_nonzero(typ == 1)
print(c1/(c1+c2))
print(c2/(c1+c2))
print(c1)
print(c2)
dfDropped = df.drop(columns=['gender'])
# + id="0nVD6Ciot9JF"
def dual_print(f,*args,**kwargs):
#print(*args,**kwargs)
print(*args,**kwargs,file=f)
def load_dataset(csv_name):
# read the dataset from csv_name and return as pandas dataframe
df = pd.read_csv(csv_name, header=None)
return df
def k_random_index(df,K):
# return k random indexes in range of dataframe
return random.sample(range(0, len(df)), K)
def find_k_initial_centroid(df,K):
centroids = [] # make of form [ [x1,y1]....]
rnd_idx = k_random_index(df,K)
#print(rnd_idx)
for i in rnd_idx:
coordinates =[]
for a in range(0,A):
coordinates.append(df.loc[i][a])
centroids.append(coordinates) #df is X,Y,....., Type
return centroids
#nOt using
def calc_distance(x1, y1, x2, y2):
# returns the euclidean distance between two points
return ((x1 - x2) ** 2 + (y1 - y2) ** 2) ** 0.5
def calc_distance_a(centroid, point):
#print('çalculating distance\n')
sum_ = 0
for i in range(0, len(centroid)):
sum_ = sum_ + (centroid[i]-point[i])**2
return sum_ #**0.5
@njit(parallel=False)
def find_distances_fast(k_centroids, df):
dist = np.zeros((len(k_centroids),len(df),A+2),np.float64)
Kcnt = 0
for c in k_centroids: #K-centroid is of form [ c1=[x1,y1.....z1], c2=[x2,y2....z2].....]
l = np.zeros((len(df),A+2),np.float64)
index = 0
for row in df: # row is now x,y,z......type
# append all coordinates to point
dis = np.sum((c- row[:A])**2)#calc_distance_a(c, point)
#Processing the vector for list
row_list = np.array([dis])
#append distance or l norm
row_list = np.append(row_list,row[:A+1])
#append all coordinates #append type of this row
l[index] = row_list
index = index + 1
#[dist, X, Y,....Z , type]
# l contains list of type [dist,X,Y.....,Z,type] for each points in metric space
dist[Kcnt]= l
Kcnt = Kcnt + 1
# return dist which contains distances of all points from every centroid
return dist
def find_distances(k_centroids, df):
dist = []
for c in k_centroids: #K-centroid is of form [ c1=[x1,y1.....z1], c2=[x2,y2....z2].....]
l = []
# for row in df:
for index, row in df.iterrows(): # row is now x,y,z......type
point =[]
for a in range(0, A):
point.append(row.iloc[a]) # append all coordinates
dis = calc_distance_a(c, point)
#Processing the vector for list
row_list = [dis]
#append distance or l norm
for a in range(0, A):
row_list.append(row.iloc[a]) #append all coordinates
#print(row.iloc[a+1])
row_list.append(row.iloc[a+1]) #append type of this row
l.append(row_list)
#l.append([calc_distance(c[0], c[1], row[0], row[1]), row[0], row[1], row[2]]) # [dist, X, Y,....Z , type]
# l contains list of type [dist,X,Y.....,Z,type] for each points in metric space
dist.append(l)
# return dist which contains distances of all points from every centroid
return dist
def sort_and_valuation(dist):
sorted_val = []
for each_centroid_list in dist:
each_centroid_list_sorted = sorted(each_centroid_list, key=lambda x: (x[A+1], x[0])) # A+1 is index of type , 0 is dist
sorted_val.append(each_centroid_list_sorted)
# sort on basis of type & then dist.
# Now all whites are towards start and all black are after white as they have additional V added to their valuation
# Among the whites, the most closest is at start of list as it has more valuation.
# Similarly sort the black points among them based on distance as did with white
return sorted_val
def clustering(sorted_valuation, hashmap_points,K):
n = len(hashmap_points.keys()) # total number of points in metric space
cluster_assign = []
for i in range(0, K):
cluster_assign.append([]) # initially all clusters are empty
map_index_cluster = []
for i in range(0,K+2):
map_index_cluster.append(0)
#initially check all sorted evaluation from 0th index
number_of_point_alloc = 0
curr_cluster = 0
# until all points are allocated
while number_of_point_alloc != n: # As convergence is guaranteed that all points will be allocated to some cluster set
start_inde = map_index_cluster[curr_cluster % K]
for inde in range(start_inde,len(sorted_valuation[curr_cluster % K])):
each = sorted_valuation[curr_cluster % K][inde]
# each is (dist,X,Y,....Z,type)
if hashmap_points[tuple(each[1: -1])] == 0: # each is (dist, X,Y,....Z, type)
cluster_assign[curr_cluster].append(each)
hashmap_points[tuple(each[1: -1])] = 1
number_of_point_alloc += 1
map_index_cluster[curr_cluster % K] = inde #next time start from here as isse prev all allocated
break
curr_cluster = (curr_cluster + 1) % K
return cluster_assign
def update_centroids_median(cluster_assign,K):
new_centroids = []
for k in range(0, K):
cAk = np.array(cluster_assign[k])
cAk = np.delete(cAk,[0,-1],axis=1)
if len(cAk) %2 ==0 and len(cAk)>0:
cc = [np.median(np.array(cAk[:-1])[:,cl]) for cl in range(0,cAk.shape[1])]
new_centroids.append(cc)
elif len(cAk) %2 !=0 and len(cAk)>0:
cc = [np.median(np.array(cAk)[:,cl]) for cl in range(0,cAk.shape[1])]
new_centroids.append(cc)
elif len(cAk)==0:
print("Error: No centroid found updation error")
return new_centroids
def update_centroids(cluster_assign,K):
new_centroids = []
for k in range(0, K):
sum_a = []
for i in range(0, A):
sum_a.append(0)
for each in cluster_assign[k]:
sum_a = [sum(x) for x in zip(sum_a, each[1:-1])]
#each is (dist,X,Y,.....Z,type)
new_coordinates = []
for a in range(0, A):
new_coordinates.append(sum_a[a] / len(cluster_assign[k]))
new_centroids.append(new_coordinates)
k=k+1
return new_centroids
def calc_clustering_objective(k_centroid, cluster_assign,K):
cost = 0
for k in range(0, K):
for each in cluster_assign[k]: #each is (dist, X,Y,....,Z,type)
dd = calc_distance_a(k_centroid[k], each[1:-1])
cost = cost + (dd)
return cost
def calc_fairness_error(df, cluster_assign,K):
U = [] # distribution of each type in original target dataset for each J = 0 , 1....
P_k_sum_over_j = [] # distribution in kth cluster sum_k( sum_j( Uj * j wale/total_in_cluster ) )
f_error = 0
cnt_j_0 = 0
cnt_j_1 = 0
# cnt_j_2 = 0
cnt = 0
for index, row in df.iterrows():
if row.iloc[-1] == 1:
cnt_j_1 += 1
elif row.iloc[-1] == 0:
cnt_j_0 += 1
# elif row.iloc[-1] == 2:
# cnt_j_2 += 1
cnt += 1
U.append(cnt_j_0 / cnt)
U.append(cnt_j_1 / cnt)
#U.append(cnt_j_2 / cnt)
for k in range(0, K): # for each cluster
for j in range(0, len(U)): #for each demographic group
cnt_j_cluster = 0
cnt_total = 0
for each in cluster_assign[k]:
if int(each[-1]) == j: #each is (dist,X, Y.....,Z,type)
cnt_j_cluster += 1
cnt_total += 1
if cnt_j_cluster !=0 and cnt_total != 0:
P_k_sum_over_j.append(-U[j] * np.log((cnt_j_cluster / cnt_total)/U[j]))
else:
P_k_sum_over_j.append(0) #log(0)=0 considered
for each in P_k_sum_over_j:
f_error += each
return f_error
def calc_balance(cluster_assign,K):
S_k = [] # balance of each k cluster
balance = 0 # min (S_k)
for k in range(0, K):
cnt_j_0 = 0
cnt_j_1 = 0
# cnt_j_2 = 0
cnt = 0
for each in cluster_assign[k]:
if int(each[-1]) == 1:
cnt_j_1 += 1
elif int(each[-1]) == 0:
cnt_j_0 += 1
# elif int(each[-1]) == 2:
# cnt_j_2 += 1
cnt += 1
if cnt_j_0 != 0 and cnt_j_1 != 0 :#and cnt_j_2!= 0:
S_k.append(min([cnt_j_0 / cnt_j_1, cnt_j_1 / cnt_j_0 ]))#, cnt_j_1 / cnt_j_2 , cnt_j_2 / cnt_j_1 , cnt_j_0 / cnt_j_2, cnt_j_2 / cnt_j_0 ]))
elif cnt_j_0 == 0 or cnt_j_1 ==0 :#or cnt_j_2==0:
S_k.append(0)
balance = min(S_k)
return balance
# + id="uBwSXAMnt9JG"
def main():
# Step1 : Load the dataset
list_fair_K=[]
list_obj_K =[]
list_balance_K=[]
os.makedirs('Adult_kmeans_permu')
for kk in [10]:#2,5,10,15,20,30,40]:
K = kk
print(" K=="+str(K)+" ")
list_fair_run=[]
list_obj_run =[]
list_balance_run=[]
seeds = [0,100,200,300,400,500,600,700,800,900,1000,1100]
for run in range(0,runs):
np.random.seed(seeds[run])
random.seed(seeds[run])
f = open('Adult_kmeans_permu/K_'+str(K)+'_run_'+str(run)+'_output.txt', 'a')
print("+"*100)
print(' RUN : '+ str(run))
list_fair_iter=[]
list_obj_iter =[]
list_balance_iter=[]
# Step2 : Find initial K random centroids using k_random_index(df) & find_k_initial_centroid(df)
k_centroid= find_k_initial_centroid(df,kk)
k_centroid_permu = list(permutations(k_centroid))
random.shuffle(k_centroid_permu)
k_centroid = k_centroid_permu[0]
permu_index = 0
max_permu_len =100 #len(k_centroid_permu)
print("Number of Permutations : "+str(max_permu_len))
prev_assignment =[]
cluster_assignment = []
for i in range(0, K):
cluster_assignment.append([]) # initially all clusters are empty
sum_time = 0
curr_itr = 0
prev_objective_cost=-1
objective_cost = 0
# Step3 : Find distances from the centroids using find_distances() with list of [ [x1,y1,z1..] , [x2,y2,z2..]....] centroids format list
while True:# and prev_objective_cost != objective_cost:
start = time.process_time()#timeit.default_timer()
dual_print(f,'Calulating distance for iteration : '+ str(curr_itr)+'\n')
df1 = df.values
k_centroids1= np.array(k_centroid)
dist = find_distances_fast(k_centroids1, df1)
dual_print(f,'Finished calc distance for iteration : '+ str(curr_itr)+'\n')
# Step4 : Find Valuation matrix for all centroids using sort_and_valuation()
dual_print(f,'Calulating Valuation for iteration : '+ str(curr_itr)+'\n')
valuation = sort_and_valuation(dist)
dual_print(f,'Finished Valuation for iteration : '+ str(curr_itr)+'\n')
#Step5 : Perform clustering using valuation matrix & hashmap of all points in metric
hash_map = {}
for index, row in df.iterrows():
temp = tuple(row[:-1])
hash_map.update({tuple(row[:-1]): 0}) #dict is of form { (x,y): 0 , ....}
dual_print(f,'Finding clusters for iteration : '+ str(curr_itr)+'\n')
prev_assignment = cluster_assignment
cluster_assignment = clustering(valuation, hash_map,K)
dual_print(f,'Finished finding cluster for iteration : '+ str(curr_itr)+'\n')
# print("Finding balance ")
balance = calc_balance(cluster_assignment,K)
f_error = calc_fairness_error(df, cluster_assignment,K)
clustering_cost = calc_clustering_objective(k_centroid,cluster_assignment,K)
objective_cost = np.round(clustering_cost,3)
list_balance_iter.append(str(balance))
list_obj_iter.append(str(objective_cost))
list_fair_iter.append(str(f_error))
dual_print(f,'balance : ' + str(balance) + '\n')
dual_print(f,'Fairness Error : ' + str(f_error) + '\n')
dual_print(f,'Clustering Objective/Cost ' + str(clustering_cost) + '\n')
#Step6 : Print the cluster assignments
#Step7 : Find new centroids using mean of all points in current assignment
stopFlag =0
if permu_index < max_permu_len-1:
permu_index += 1
k_centroid = k_centroid_permu[permu_index] #update_centroids(cluster_assignment,K)
else:
stopFlag =1
dual_print(f,'Finished centroid updation for iteration : '+ str(curr_itr)+'\n')
dual_print(f,'Iteration No: '+str(curr_itr)+' : updated centroid are : '+ str(k_centroid))
#Step8 : Repeat from Step3 until clusters are same or iterations reach upper limit
stop = time.process_time()#timeit.default_timer()
sum_time += (stop - start)
dual_print(f,'Time for iteration : ' + str(curr_itr) + ' is ' + str(stop - start) + '\n')
curr_itr += 1
if stopFlag==1:
break
dual_print(f,'-----------------------------Finished-----------------------------------------------\n')
print('Total time taken to converge '+ str(sum_time)+'\n')
print('Iterations total taken for convergence : '+str(curr_itr)+'\n')
dual_print(f,'Total time taken is '+ str(sum_time)+'\n')
dual_print(f,'Iterations total : '+str(curr_itr-1))
#Step 10 : Find balance , fairness error , and clustering objective or cost
balance_converged = calc_balance(cluster_assignment,K)
f_error_converged = calc_fairness_error(df, cluster_assignment,K)
clustering_cost_converged = calc_clustering_objective(k_centroid,cluster_assignment,K)
print("\nCost variation over iterations")
print(list_obj_iter)
print("\nBalance variation over iterations")
print(list_balance_iter)
print("\nFairness error over iterations")
print(list_fair_iter)
print('\n')
print('Final converged balance : ' + str(balance_converged) + '\n')
print('Final Converged Fairness Error : ' + str(f_error_converged) + '\n')
print('Final converged Clustering Objective/Cost ' + str(clustering_cost_converged) + '\n')
dual_print(f,'Converged balance : ' + str(balance_converged) + '\n')
dual_print(f,'Converged Fairness Error : ' + str(f_error_converged) + '\n')
dual_print(f,'Converged Clustering Objective/Cost ' + str(clustering_cost_converged) + '\n')
f.close()
run = run +1
list_obj_run.append(clustering_cost_converged)
list_fair_run.append(f_error_converged)
list_balance_run.append(balance_converged)
print("@"*70)
print("Cost variations over run")
print(str(list_obj_run))
print("balance variations over run")
print(str(list_balance_run))
print("fairness error over run")
print(str(list_fair_run))
print("#"*30)
print("Mean Cost variations over run")
print(str(np.mean(np.array(list_obj_run))))
print("Std Dev Cost variations over run")
print(str(np.std(np.array(list_obj_run))))
print("#"*30)
list_obj_K.append(np.mean(np.array(list_obj_run)))
list_fair_K.append(np.mean(np.array(list_fair_run)))
list_balance_K.append(np.mean(np.array(list_balance_run)))
print("%"*70)
print("Cost variations over K")
print(str(list_obj_K))
print("balance variations over K")
print(str(list_balance_K))
print("fairness error over K")
print(str(list_fair_K))
print("#"*30)
# + id="yQGBVfEIt9JH" outputId="4074e1ca-f909-4a76-9aff-3b964700c279"
import time
if __name__ == "__main__":
main()
| Ablation Study/Impact of center order/FRAC_OE_adult_KmeansPermu.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from mpl_toolkits.mplot3d import Axes3D
import matplotlib.pyplot as plt
# +
fig = plt.figure(figsize=(10,10))
ax = fig.add_subplot(111, projection='3d')
xs =[1,35,70,104,138,173,207,241,275,309,343,378,412,447,481,515,548]
ys =[1,5,5311,70768,332424,1118206,3044940,5903932,8088851,9534964,10374932,11000000,11409831,15000000,26752447,30000000,31528114]
zs =[1.9,1.95,1.91,1.33,1.2,1.25,1.08,0.97,0.89,0.9,0.95,0.96,1.34,1.41,0.79,0.77,1.01]
ax.scatter(xs, ys, zs, c='r', marker='^')
ax.set_xlabel('Days')
ax.set_ylabel('Total Cases in Millions')
ax.set_zlabel('Reproduction Rate')
plt.show()
# -
| Codes/3D_model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jonfisik/Projects/blob/master/3EscalonamentoPython.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="d-483GRS13wO"
import numpy as np
# + id="SdPMT1Ri2Kvd"
M = np.array([[1,4,3,1],[2,5,4,4],[1,-3,-2,5]])
print(M)
# + id="LZHwvlXk2j9y"
M[1,:] = -2*M[0,:] + M[1,:]
print(M)
# + id="NdicNfyN2z55"
M[2,:] = -1*M[0,:] + M[2,:]
print(M)
# + id="LoU11Kbk3Byp"
M[1,:] = -1/3*M[1,:]
print(M)
# + id="AP_Zb2VA3N-Q"
M[0,:] = -4*M[1,:] + M[0,:]
print(M)
# + id="g6TZETQj3Z1p"
M[2,:] = 7*M[1,:] + M[2,:]
print(M)
# + id="TPZnLJ014CRP"
M[2,:] = -3*M[2,:]
print(M)
# + id="mlSDcZzt4sDE"
M[1,:] = -1/3*M[2,:] + M[1,:]
print(M)
# + id="I6tJlMKD6sVZ"
M[0,:] = -1/3*M[2,:] + M[0,:]
print(M)
# + id="o8UIyrCs67DY"
M[2,:] = M[2,:]/3
print(M)
# + id="YQe6H-P46_n3"
M[1,:] = 5*M[2,:] + M[1,:]
print(M)
# + id="mxcRoYP1pWsS"
M[0,:] = 2*M[2,:] + M[0,:]
print(M)
# + id="NQZtRik7pm_B"
| python/AlgebraLinearPython/3EscalonamentoPython.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import os
os.environ['PYTHONHASHSEED']=str(0)
import random
random.seed(0)
import numpy as np
np.random.seed(0)
# the libraries above are used to set seeds for replication purposes
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
import math
import pickle
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler,MinMaxScaler
from workalendar.europe import Greece
cal = Greece()
# libraries for preprocessing
import tensorflow as tf
import keras
from keras.preprocessing.sequence import TimeseriesGenerator
from keras.models import Sequential,Model
from keras.models import load_model
from keras.layers import Dropout
from keras.layers import Input,Dense,LSTM,GRU,RNN
from sklearn.metrics import mean_squared_error
from keras.callbacks import ModelCheckpoint
from keras.regularizers import l1
from keras.regularizers import l2
#tensorflow and keras
from IPython.display import Image
from IPython.core.display import HTML
# -
data=pd.read_excel('all_data.xlsx')
validation_set=data.tail(5424)
data=data.head(21600)
data
# +
def working_days(df_date): # function to flag national holidays and weekends in Greece
if cal.is_working_day(df_date):
return 1
else:
return 0
data.index=data.Date
data['Month'] = data.index.month
data['Holiday_weekend']=data['Date'].apply(working_days)
data=data.reset_index(drop=True)
data
# -
# # Scale Data
# +
input_data=data[['Load','Month','Temperature_Athens','Wind_Athens','Humidity_Athens',
'Temperature_Thessaloniki','Wind_Thessaloniki','Humidity_Thessaloniki']] # this is what we are going to use as input
output_data=data['Load'] # we only want to predict the load as output
input_scaler=MinMaxScaler() # scale all input data to [0,1] range
input_scaled=input_scaler.fit_transform(input_data)
input_scaled=pd.DataFrame(input_scaled)
input_scaled['Holiday_weekend']=data['Holiday_weekend']
input_scaled.columns=['Load','Month','Temperature_Athens','Wind_Athens','Humidity_Athens',
'Temperature_Thessaloniki','Wind_Thessaloniki','Humidity_Thessaloniki','Holiday_weekend']
output_scaler=MinMaxScaler() # scale the output load to [0,1] range as well
output_scaled=output_scaler.fit_transform(np.array(output_data).reshape(-1,1))
output_scaled=pd.DataFrame(output_scaled)
output_scaled.columns=['Load']
filename1 = 'input_scaler.sav' # save the fitted scalers in a file
filename2 = 'output_scaler.sav'
pickle.dump(input_scaler, open(filename1, 'wb'))
pickle.dump(output_scaler, open(filename2, 'wb'))
# -
input_scaled.head(50)
# # Timeseries Forecasting with Sliding Window
# In forecasting, we are interested in predicting future values of the timeseries based on historical data. In order to forecast with supervised learning methods such as neural networks, we should transform our data in input features and target or output features. In our case the target features should be the future values of electrical load, while the input features are the n past values of electrical load and weather data.
#
# We can use the functions below to transform our dataset in this format. Each observation in our transformed data will be an array of n historical values while the corresponding target feature will be an array of 24 future values.
#
# Via this transformation, we are performing a **sliding window forecast**, in this case of size 24h. So every observation in our sample is shifted 24h later than the previous observation. This is called sliding window because every observation in our sample is **slided** by 24h forward.
# 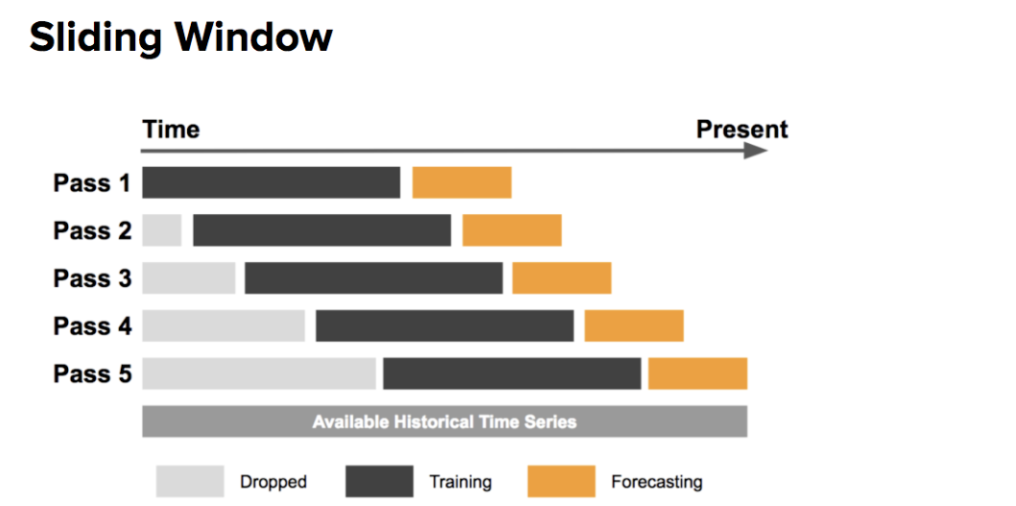
# Source: https://eng.uber.com/forecasting-introduction/
# 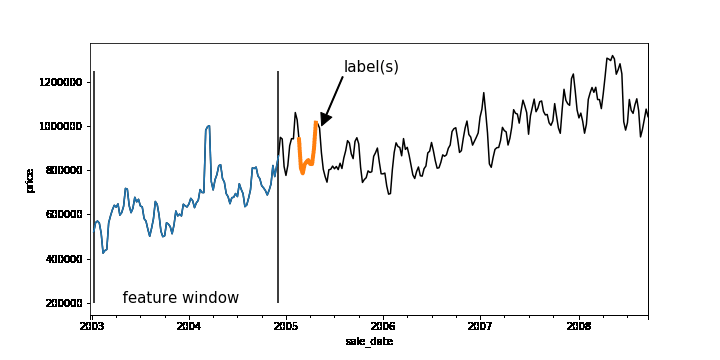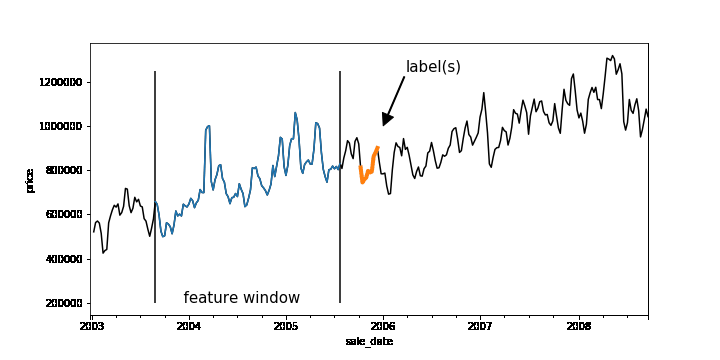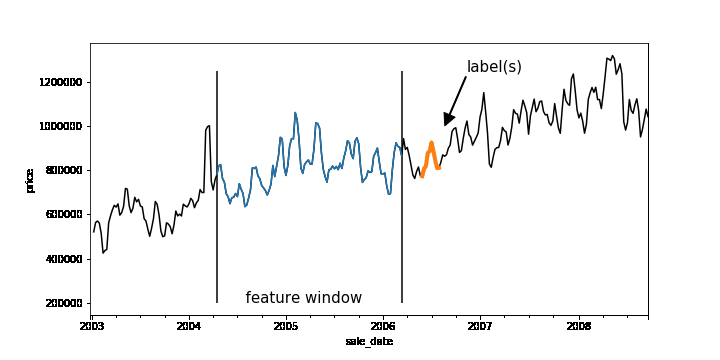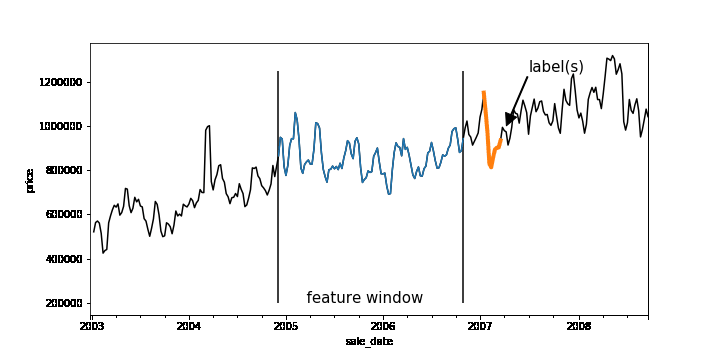
# In this run, we use 96 past hours as the input size, 24h which is the sliding window size and 24 h as the desired output size.
# +
# every split holds 96 hours (input size) that is 24h (sliding window size) later than the previous split
def create_x(x,y):
generator = TimeseriesGenerator(x,y,length=96, batch_size=24)
input_x=[]
output_y=[]
for i in range(len(generator)):
input_x.append(generator[i][0][0])
output_y.append(generator[i][1][0])
return (input_x)
def create_y(x,y): # we create y (output data) splitted every 24 hours (length of output we want to predict)
generator = TimeseriesGenerator(x,y,length=24, batch_size=24)
input_x=[]
output_y=[]
for i in range(len(generator)):
input_x.append(generator[i][0][0])
output_y.append(generator[i][1][0])
return (input_x)
# +
x=create_x(input_scaled.values,output_scaled.values)
y=create_y(output_scaled.values,input_scaled.values)
x=np.array(x)
y=np.array(y)
y=y[4:] #first 4 days (96 h) should be only in the input, so that the output contains future values of the correspondent input features
# -
# x is in 3d shape, since LSTM layers require data to be in **[samples,timesteps,features]** shape.
x=x[:y.shape[0]]
y=y.reshape(y.shape[0],y.shape[1])
# We are going to use a total of 895 samples, where each sample is represented by 9 features and we use 96 values for each feature.
x.shape
y.shape
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.2,random_state=0)
# +
tf.keras.backend.clear_session()
tf.random.set_seed(0)
model = Sequential()
model.add(LSTM(100,input_shape=(96, 9),return_sequences=True))
model.add(LSTM(100,return_sequences=True))
model.add(LSTM(50,return_sequences=False))
# model.add(Dropout(0.2))
model.add(Dense(24,activation='sigmoid')) # sigmoid since we have scaled to [0,1]
model.summary()
# +
#batch is the number of samples that is shown to the network before a weight update is performed
n_epochs = 1000 # don't care much about epochs since we use early stopping
batch=32
model.compile(optimizer='adam', loss='mse')
checkpointer = ModelCheckpoint(filepath="LSTM_model_96h.h5",
verbose=1,
save_best_only=True)
es_callback = keras.callbacks.EarlyStopping(monitor='val_loss',patience=5) # early stopping
history = model.fit(x_train, y_train,
epochs=n_epochs,
batch_size=batch,
shuffle=True,
validation_split=0.20,
verbose=0,
callbacks=[checkpointer,es_callback])
df_loss = pd.DataFrame(history.history)
df_loss[['loss','val_loss']].plot()
# +
#reasonable learning curve
# +
model = load_model('LSTM_model_96h.h5')
input_scaler= pickle.load(open("input_scaler.sav", "rb"))
output_scaler= pickle.load(open("output_scaler.sav", "rb"))
predictions_train = model.predict(x_train)
predictions_test = model.predict(x_test)
predictions_train = output_scaler.inverse_transform(predictions_train)
y_train=output_scaler.inverse_transform(y_train)
predictions_test = output_scaler.inverse_transform(predictions_test)
y_test=output_scaler.inverse_transform(y_test)
trainScore = math.sqrt(mean_squared_error(y_train, predictions_train))
print('Train Score: %.2f RMSE' % (trainScore))
testScore = math.sqrt(mean_squared_error(y_test, predictions_test))
print('Test Score: %.2f RMSE' % (testScore))
# +
# probably test size should be bigger
# +
predictions=pd.DataFrame(predictions_test)
predictions['Day']=predictions.index +1
actual=pd.DataFrame(y_test)
actual['Day']=actual.index +1
actual=pd.melt(actual,id_vars=['Day'],var_name='Hour').sort_values(by=['Day','Hour']).reset_index(drop=True)
predicted=pd.melt(predictions,id_vars=['Day'],var_name='Hour').sort_values(by=['Day','Hour']).reset_index(drop=True)
joined=pd.DataFrame()
joined['Actual']=actual['value']
joined['Predicted']=predicted['value']
plot_data=joined[['Actual','Predicted']]
plot_data.head(24).plot()
# -
plot_data.iloc[48:96].plot()
plot_data.tail(72).plot()
plot_data.iloc[1800:2000].plot()
plot_data.iloc[1900:1950].plot()
plot_data.iloc[2950:3010].plot()
# With this model we can predict 24 future values of the electrical load using the past 96 values of load and weather data. We still face the issue of the forecasting horizon (since the model needs the exact n past values in order to predict the next 24) as in the feed forward models, but we can see that LSTM layers make better use of sequence data such as timeseries.
# # Grid Search Parameters
# We are interested in finding the optimal parameter values that will result in the lowest training error. After identifying those parameter, we can introduce reguralization (dropout, l1,l2 norm) to reduce overfitting.
# +
# use the same functions as previously but use input length as a variable as well
def create_x(x,y,x_length):
generator = TimeseriesGenerator(x,y,length=x_length, batch_size=24)
input_x=[]
output_y=[]
for i in range(len(generator)):
input_x.append(generator[i][0][0])
output_y.append(generator[i][1][0])
return (input_x)
def create_y(x,y): # keep the sliding window of size 24 and output size 24
generator = TimeseriesGenerator(x,y,length=24, batch_size=24)
input_x=[]
output_y=[]
for i in range(len(generator)):
input_x.append(generator[i][0][0])
output_y.append(generator[i][1][0])
return (input_x)
def grid_search(scaler,input_size,batch,layer_type):
############### follow the same preprocessing pipeline ###################################
input_data=data[['Load','Month','Temperature_Athens','Wind_Athens','Humidity_Athens',
'Temperature_Thessaloniki','Wind_Thessaloniki','Humidity_Thessaloniki']]
output_data=data['Load']
input_scaler=scaler
input_scaled=input_scaler.fit_transform(input_data)
input_scaled=pd.DataFrame(input_scaled)
input_scaled['Holiday_weekend']=data['Holiday_weekend']
input_scaled.columns=['Load','Month','Temperature_Athens','Wind_Athens','Humidity_Athens',
'Temperature_Thessaloniki','Wind_Thessaloniki','Humidity_Thessaloniki','Holiday_weekend']
output_scaler=scaler
output_scaled=output_scaler.fit_transform(np.array(output_data).reshape(-1,1))
output_scaled=pd.DataFrame(output_scaled)
output_scaled.columns=['Load']
filename1 = 'input_scaler.sav'
filename2 = 'output_scaler.sav'
pickle.dump(input_scaler, open(filename1, 'wb'))
pickle.dump(output_scaler, open(filename2, 'wb'))
x=create_x(input_scaled.values,output_scaled.values,input_size)
y=create_y(output_scaled.values,input_scaled.values)
x=np.array(x)
y=np.array(y)
flag_point=int(input_size/24)
y=y[flag_point:]
x=x[:y.shape[0]]
y=y.reshape(y.shape[0],y.shape[1])
################### Train & Test Split ###########################################
temp_train_scores=[]
temp_test_scores=[]
finish_point=0
while finish_point < 5: #for every model try 5 different train test splits and average results from all splits
x_train,x_test,y_train,y_test = train_test_split(x,y,test_size=0.2,random_state=finish_point)
#################### Build Model #################################################
tf.keras.backend.clear_session()
tf.random.set_seed(0)
model = Sequential()
model.add(layer_type(100,input_shape=(input_size, 9),return_sequences=True))
model.add(layer_type(100,return_sequences=False))
model.add(Dense(24,activation='sigmoid'))
n_epochs = 1000
batch=batch
model.compile(optimizer='adam', loss='mse')
checkpointer = ModelCheckpoint(filepath="load_model.h5",
verbose=0,
save_best_only=True)
es_callback = keras.callbacks.EarlyStopping(monitor='val_loss',patience=3) # early stopping
history = model.fit(x_train, y_train,
epochs=n_epochs,
batch_size=batch,
shuffle=True,
validation_split=0.20,
verbose=0,
callbacks=[checkpointer,es_callback])
################# load trained model and make predictions ################################
model = load_model('load_model.h5')
input_scaler= pickle.load(open("input_scaler.sav", "rb"))
output_scaler= pickle.load(open("output_scaler.sav", "rb"))
predictions_train = model.predict(x_train)
predictions_test = model.predict(x_test)
predictions_train = output_scaler.inverse_transform(predictions_train)
y_train=output_scaler.inverse_transform(y_train)
predictions_test = output_scaler.inverse_transform(predictions_test)
y_test=output_scaler.inverse_transform(y_test)
trainScore = math.sqrt(mean_squared_error(y_train, predictions_train))
testScore = math.sqrt(mean_squared_error(y_test, predictions_test))
temp_train_scores.append(trainScore)
temp_test_scores.append(testScore)
finish_point += 1
print('Train Score: %.2f RMSE' % (sum(temp_train_scores)/len(temp_train_scores)))
print('Test Score: %.2f RMSE' % (sum(temp_test_scores)/len(temp_test_scores)))
return(sum(temp_train_scores)/len(temp_train_scores),sum(temp_test_scores)/len(temp_test_scores))
# +
import warnings
warnings.filterwarnings('ignore')
layer_types=[keras.layers.LSTM, keras.layers.GRU, keras.layers.RNN]
input_sizes=[24,48,72,96] # transform data with all these input sizes
batch_sizes=[None,15,32,64] # try these batch sizes
train_scores=[]
test_scores=[]
inputs=[]
batches=[]
nn_layers=[]
for k in layer_types:
for i in input_sizes:
for j in batch_sizes:
print('Input Size:' + str(i))
print('Batch Size:' + str(j))
print('Layer Type:' + str(k))
print('|---------------------| :')
tr,te=grid_search(MinMaxScaler(),i,j,k)
train_scores.append(tr)
test_scores.append(te)
inputs.append(i)
batches.append(j)
nn_layers.append(k)
# -
my_dict = {'Input_Size': inputs,'Batch_Size': batches, 'Train_Score': train_scores, 'Test_Score': test_scores}
df = pd.DataFrame(my_dict)
df
df.loc[df['Train_Score'].idxmin()]
df.loc[df['Test_Score'].idxmin()]
# # Trying Best Model on Validation set
# We are now going to train a model with all data we used previously for training and testing and validate the forecasting accuracy on the small validation set we kept at the beginning. We are going to build a model with **LSTM layers** of input size **72 past hours** **without setting a batch size**.
# +
input_data=data[['Load','Month','Temperature_Athens','Wind_Athens','Humidity_Athens',
'Temperature_Thessaloniki','Wind_Thessaloniki','Humidity_Thessaloniki']]
output_data=data['Load']
input_scaler=MinMaxScaler() # scale all input data to [0,1] range
input_scaled=input_scaler.fit_transform(input_data)
input_scaled=pd.DataFrame(input_scaled)
input_scaled['Holiday_weekend']=data['Holiday_weekend']
input_scaled.columns=['Load','Month','Temperature_Athens','Wind_Athens','Humidity_Athens',
'Temperature_Thessaloniki','Wind_Thessaloniki','Humidity_Thessaloniki','Holiday_weekend']
output_scaler=MinMaxScaler() # scale the output load to [0,1] range as well
output_scaled=output_scaler.fit_transform(np.array(output_data).reshape(-1,1))
output_scaled=pd.DataFrame(output_scaled)
output_scaled.columns=['Load']
filename1 = 'input_scaler.sav' # save the fitted scalers in a file
filename2 = 'output_scaler.sav'
pickle.dump(input_scaler, open(filename1, 'wb'))
pickle.dump(output_scaler, open(filename2, 'wb'))
# +
def create_x(x,y):
generator = TimeseriesGenerator(x,y,length=72, batch_size=24)
input_x=[]
output_y=[]
for i in range(len(generator)):
input_x.append(generator[i][0][0])
output_y.append(generator[i][1][0])
return (input_x)
def create_y(x,y): # we create y (output data) splitted every 24 hours (length of output we want to predict)
generator = TimeseriesGenerator(x,y,length=24, batch_size=24)
input_x=[]
output_y=[]
for i in range(len(generator)):
input_x.append(generator[i][0][0])
output_y.append(generator[i][1][0])
return (input_x)
x=create_x(input_scaled.values,output_scaled.values)
y=create_y(output_scaled.values,input_scaled.values)
x=np.array(x)
y=np.array(y)
y=y[3:] #first 3 days (72 h) should be only in the input, so that the output contains future values of the correspondent input features
# -
x=x[:y.shape[0]]
y=y.reshape(y.shape[0],y.shape[1])
print(x.shape)
print(y.shape)
# +
tf.keras.backend.clear_session()
tf.random.set_seed(0)
model = Sequential()
model.add(LSTM(100,input_shape=(72, 9),return_sequences=True))
model.add(Dropout(0.2)) # introduce dropout between layers, to reduce overfitting
model.add(LSTM(100,return_sequences=False))
model.add(Dense(24,activation='sigmoid'))
n_epochs = 1000
model.compile(optimizer='adam', loss='mse', metrics=['mape']) # also report the Mean Absolute Percentage Error at each epoch
checkpointer = ModelCheckpoint(filepath="best_load_model_sliding_window_24.h5",
verbose=1,
save_best_only=True)
es_callback = keras.callbacks.EarlyStopping(monitor='val_loss',patience=5) # early stopping
history = model.fit(x, y, # now use the whole dataset (without splitting to train and test)
epochs=n_epochs,
shuffle=True,
validation_split=0.20,
verbose=1,
callbacks=[checkpointer,es_callback])
# -
model.save_weights("best_load_model_sliding_window_24_weights")
df_loss = pd.DataFrame(history.history)
df_loss[['loss','val_loss']].plot()
validation_set.index=validation_set.Date
validation_set['Month'] = validation_set.index.month
validation_set['Holiday_weekend']=validation_set['Date'].apply(working_days)
validation_set=validation_set.reset_index(drop=True)
validation_set.head(20)
# +
input_data=validation_set[['Load','Month','Temperature_Athens','Wind_Athens','Humidity_Athens',
'Temperature_Thessaloniki','Wind_Thessaloniki','Humidity_Thessaloniki']]
output_data=validation_set['Load']
input_scaled=input_scaler.transform(input_data)
input_scaled=pd.DataFrame(input_scaled)
input_scaled['Holiday_weekend']=data['Holiday_weekend']
input_scaled.columns=['Load','Month','Temperature_Athens','Wind_Athens','Humidity_Athens',
'Temperature_Thessaloniki','Wind_Thessaloniki','Humidity_Thessaloniki','Holiday_weekend']
output_scaled=output_scaler.transform(np.array(output_data).reshape(-1,1))
output_scaled=pd.DataFrame(output_scaled)
output_scaled.columns=['Load']
# +
x=create_x(input_scaled.values,output_scaled.values)
y=create_y(output_scaled.values,input_scaled.values)
x=np.array(x)
y=np.array(y)
y=y[3:]
x=x[:y.shape[0]]
y=y.reshape(y.shape[0],y.shape[1])
print(x.shape)
print(y.shape)
# +
model = load_model('best_load_model_sliding_window_24.h5')
predictions_validation_set = model.predict(x)
predictions_validation_set = output_scaler.inverse_transform(predictions_validation_set)
y_actual=output_scaler.inverse_transform(y)
val_score = math.sqrt(mean_squared_error(y_actual, predictions_validation_set))
print('Validation Set Score: %.2f RMSE' % (val_score))
# +
predictions=pd.DataFrame(predictions_validation_set)
predictions['Day']=predictions.index +1
actual=pd.DataFrame(y_actual)
actual['Day']=actual.index +1
actual=pd.melt(actual,id_vars=['Day'],var_name='Hour').sort_values(by=['Day','Hour']).reset_index(drop=True)
predicted=pd.melt(predictions,id_vars=['Day'],var_name='Hour').sort_values(by=['Day','Hour']).reset_index(drop=True)
joined=pd.DataFrame()
joined['Actual']=actual['value']
joined['Predicted']=predicted['value']
plot_data=joined[['Actual','Predicted']]
plot_data.head(24).plot()
# -
plot_data.tail(24).plot()
plot_data.iloc[500:600].plot()
plot_data['Absolute Percentage Error']=(abs(plot_data['Actual']-plot_data['Predicted'])/plot_data['Actual'])*100
plot_data['Absolute Error']=abs(plot_data['Actual']-plot_data['Predicted'])
plot_data['Residual']=plot_data['Actual']-plot_data['Predicted']
plot_data
plot_data['Absolute Percentage Error'].mean()
plot_data['Absolute Error'].mean()
plot_data['Residual'].plot(figsize=(20,8))
plot_data['Residual'].hist(figsize=(15,8))
# The residuals appear to be normaly distributed centered around 0. These are the results we would expect our model to have in a real case scenario.
| Models/LSTM Models with Sliding Window size_24.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <div style="text-align: center; line-height: 0; padding-top: 2px;">
# <img src="https://www.quantiaconsulting.com/logos/quantia_logo_orizz.png" alt="Quantia Consulting" style="width: 600px; height: 250px">
# </div>
# # Final Challenge A
# ---
#
# ## Using the KDDCup dataset, find the best SML model
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
# The KDDCup intrusion detection data stream records intrusions simulated in a military network environment. The task is to classify network traffic into **normal** (80.31% of the cases) or some kind of **intrusion** (19.69% of the cases) described by *41 features*, of which *34 numeric* and *7 nominal*. The problem of temporal dependence is particularly evident here. Inspecting the raw stream confirms that there are time periods of intrusions rather than single instances of intrusions.
# The *7 nominal* attributes are:
# - protocol_type
# - service
# - flag
# - land
# - logged_in
# - is_host_login
# - is_guest_login
#
# **Class:** `class` | 0: normal, 1: intrusion
#
# **Samples:** 494021
#
df = pd.read_csv("../datasets/KDDCup.csv")#.iloc[:10000,:]
features = df.columns[:-1]
df.dtypes
# ## Plot data distribution
# ---
# Encode the protocol_type, service and flag attributes and apply the **PCA**
# +
from sklearn.preprocessing import OrdinalEncoder
from sklearn.preprocessing import StandardScaler
from sklearn.decomposition import PCA
encoder = OrdinalEncoder()
encDf = df.copy()
encDf[["protocol_type","service","flag"]] = encoder.fit_transform(encDf[["protocol_type","service","flag"]])
features = encDf.drop(["class"],axis=1)
label = encDf["class"]
scaled_features = StandardScaler().fit_transform(features)
num_components = 2
pca = PCA(n_components=num_components)
pca_data = pca.fit_transform(scaled_features)
pca_data.shape
pc_df = pd.DataFrame(data = pca_data,columns = ["pc1", "pc2"])
pc_df["class"] = label
normalities = pc_df[pc_df["class"] == 0]
intrusions = pc_df[pc_df["class"] == 1]
fig = plt.figure()
ax1 = fig.add_subplot(111)
ax1.scatter(x=normalities["pc1"], y=normalities["pc2"], marker="s", label='Normalities: # ' + str(normalities.shape[0]))
ax1.scatter(x=intrusions["pc1"], y=intrusions["pc2"], marker="o", label='Intrusions: # ' + str(intrusions.shape[0]))
plt.legend(loc='upper left');
fig.tight_layout()
plt.show()
# -
# ## Find the best SML model!
# ---
# ### Hint
# Since the stream is imbalances, the Accuracy is not a reliable metric. Try using [BalancedAccuracy](https://riverml.xyz/latest/api/metrics/BalancedAccuracy/) and [GeometricMean](https://riverml.xyz/latest/api/metrics/GeometricMean/) instead. To combine them, use [Metrics](https://riverml.xyz/latest/api/metrics/Metrics/)
from river.stream import iter_pandas
from river.evaluate import progressive_val_score
from river.metrics import Metrics
from river.metrics import BalancedAccuracy
from river.metrics import GeometricMean
from river import compose
| lab/4.0_Final_Challenge_A.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 0.6.2
# language: julia
# name: julia-0.6
# ---
# First, a few functions to gather the data from the Nektar output
# +
using DataFrames, Base.Test, Glob, Query
function benchmark_info(filename::String)
regex = match(r"(gcp|cosma)/(cardiac|naca65)/\2_(\d+)_(\d+)/out", filename)
platform = Symbol(regex.captures[1])
bench = Symbol(regex.captures[2])
nodes = parse(Int64, regex.captures[3])
ppn = parse(Int64, regex.captures[4])
platform, bench, nodes, ppn
end
function benchmark_data(filename::String)
@assert isfile(filename)
regex = r"Steps: (\d+)\s+Time:\s+(\S*)\s+CPU Time: (\S*)s"
steps, tstep, time = Int64[], Float64[], Float64[]
open(filename) do file
for line in eachline(file)
capture = match(regex, line)
capture == nothing && continue
push!(steps, parse(Int64, capture.captures[1]))
push!(tstep, parse(Float64, capture.captures[2]))
push!(time, parse(Float64, capture.captures[3]))
end
end
platform, bench, nnodes, nprocs = benchmark_info(filename)
DataFrame(:step => steps, :tstep => tstep, :time => time,
:nodes => nnodes, :ppn => nprocs, :n => nnodes * nprocs,
:benchmark => bench, :platform => platform)
end
function gather_data(files)
result = benchmark_data(files[1])
for filename in files[2:end]
result = [result; benchmark_data(filename)]
end
categorical!(result, :benchmark)
categorical!(result, :platform)
result
end
@testset "opening file" begin
filename = "gcp/cardiac/cardiac_1_8/out"
@test benchmark_info(filename)[1] == :gcp
@test benchmark_info(filename)[2] == :cardiac
@test benchmark_info(filename)[3] == 1
@test benchmark_info(filename)[4] == 8
regex = r"Steps: (\d+)\s+Time:\s+(\S*)\s+CPU Time: (\S*)s"
line = "Steps: 579 Time: 11.58 CPU Time: 11.6915s"
@test match(regex, line) != nothing
data = benchmark_data(filename)
@test nrow(data) == 4285
@test all(data[:nodes] .== 1)
@test all(data[:ppn] .== 8)
@test all(data[:benchmark] .== :cardiac)
@test all(data[1:6, :step] .== 1:6)
@test all(data[1:6, :tstep] .≈ 0.02 .* collect(1:6))
@test all(data[1:2, :time] .≈ [7.4155, 1.64536])
end
# -
# Then we actually gather the data, and compute the speedup vs the code running on Cosma with a single process.
data = gather_data(glob([r"(gcp|cosma)", r"(cardiac|naca65)", r"(cardiac|naca65)_\d+_\d+", "out"]));
means = @from r in data begin
@where r.step != 1
@group r by (r.n, r.nodes, r.ppn, r.platform, r.benchmark) into g
@select {n=g.key[1], nodes=g.key[2],
ppn=g.key[3], platform=g.key[4],
benchmark=g.key[5], mean=mean(g..time)}
@collect DataFrame
end
serial = means[(means[:n] .== 1) .& (means[:platform] .== :cosma), :]
speedup = @from r in means begin
@join s in serial on r.benchmark equals s.benchmark
@select {r.benchmark, r.platform, r.n, r.nodes, r.ppn, r.mean, speedup=s.mean/r.mean}
@collect DataFrame
end;
# We create a plot for each benchmark
# +
using StatPlots, GR
gr()
cardiac = speedup |>
@filter(_.benchmark == :cardiac && _.ppn < 20) |>
@map({_.n, _.platform, _.speedup}) |>
@df StatPlots.scatter(
:n, :speedup, group=:platform,
markershape=ifelse.(:platform .== :gcp, :cross, :circle))
title!(cardiac, "Cardiac electro-physiology")
xlabel!(cardiac, "Number of processes")
ylabel!(cardiac, "Speedup vs one process on Cosma");
# -
naca = speedup |>
@filter(_.benchmark == :naca65 && _.ppn < 20) |>
@map({_.n, _.platform, _.speedup}) |>
@df StatPlots.scatter(
:n, :speedup, group=:platform,
markershape=ifelse.(:platform .== :gcp, :cross, :circle))
title!(naca, "Wing cross-section")
xlabel!(naca, "Number of processes")
ylabel!(naca, "Speedup vs one process on Cosma");
# Then plot and save
plots = Plots.plot(cardiac, naca, legend=(0.8, 0.2))
Plots.savefig("benchmarks.png")
plots
speedup[speedup[:n] .== 1, :]
0.0124915 / 0.0190376
| nektar/Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="eQ7K9blSUKTs" colab_type="text"
# Copyright 2019 Google LLC
#
# Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
#
# # (Attentive) Neural Processes for 1D regression
#
# Regression is usually cast as modelling the distribution of output **y** given input **x** via a deterministic function, such as a neural network, taking **x** as input. In this setting, the model is trained on a dataset of input-output pairs, and predictions of the outputs are independent of each other given the inputs. An alternative approach to regression involves using the training data to compute a distribution over functions that map inputs to outputs, and using draws from that distribution to make predictions on test inputs. This approach allows for reasoning about multiple functions consistent with the data, and can capture the co-variability in outputs given inputs. In the Bayesian machine learning literature, non-parametric models such as Gaussian Processes (GPs) are popular choices of this approach.
#
# [Neural Processes](https://arxiv.org/abs/1807.01622) (NPs) also approach regression by modelling a distribution over regression functions. Each function models the distribution of the output given an input, conditioning on some observed input-output pairs, which we call the context. Modelling this distribution over functions was made possible by incorporating a latent variable to the [Conditional Neural Process](https://arxiv.org/abs/1807.01613) (CNP).
#
# However NPs suffer from underfitting, giving inaccurate predictions at the inputs of the observed data they condition on. Hence [Attentive Neural Processes](https://arxiv.org/abs/1901.05761) (ANPs) were introduced, addressing this issue by incorporating attention into NPs. We share the implementation of ANPs (and NPs, which is a special case of ANP with uniform attention) for a 1D regression task where (A)NPs are trained on random 1D functions.
# + [markdown] id="UMJxjfsTa08h" colab_type="text"
# ## Imports
# + id="Ncb7M0FpNfix" colab_type="code" colab={}
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
import collections
from google.colab import files
# + [markdown] id="ARGYAmEsa5K7" colab_type="text"
# ## Data generator
# Instead of training using observations from a single function (as in classic regression tasks), we would like to train on a dataset that comes from multiple functions with shared characteristics. Hence for training, we use data that comes from a Gaussian Process (GP) with randomly varying kernel parameters. At each training iteration, we sample a batch of random kernel parameters, and for each parameter setting we sample a curve (a realisation) from the corresponding GP. We select random points on each curve to be the targets and a subset to be the contexts for optimising the training loss. The data generation is almost the same as for the [implementation of CNPs](https://github.com/deepmind/conditional-neural-process/blob/master/conditional_neural_process.ipynb), but with kernel parameters varying randomly at each iteration.
# + id="Px-atGEfNnWT" colab_type="code" cellView="form" colab={}
#@title
# The (A)NP takes as input a `NPRegressionDescription` namedtuple with fields:
# `query`: a tuple containing ((context_x, context_y), target_x)
# `target_y`: a tensor containing the ground truth for the targets to be
# predicted
# `num_total_points`: A vector containing a scalar that describes the total
# number of datapoints used (context + target)
# `num_context_points`: A vector containing a scalar that describes the number
# of datapoints used as context
# The GPCurvesReader returns the newly sampled data in this format at each
# iteration
NPRegressionDescription = collections.namedtuple(
"NPRegressionDescription",
("query", "target_y", "num_total_points", "num_context_points"))
class GPCurvesReader(object):
"""Generates curves using a Gaussian Process (GP).
Supports vector inputs (x) and vector outputs (y). Kernel is
mean-squared exponential, using the x-value l2 coordinate distance scaled by
some factor chosen randomly in a range. Outputs are independent gaussian
processes.
"""
def __init__(self,
batch_size,
max_num_context,
x_size=1,
y_size=1,
l1_scale=0.6,
sigma_scale=1.0,
random_kernel_parameters=True,
testing=False):
"""Creates a regression dataset of functions sampled from a GP.
Args:
batch_size: An integer.
max_num_context: The max number of observations in the context.
x_size: Integer >= 1 for length of "x values" vector.
y_size: Integer >= 1 for length of "y values" vector.
l1_scale: Float; typical scale for kernel distance function.
sigma_scale: Float; typical scale for variance.
random_kernel_parameters: If `True`, the kernel parameters (l1 and sigma)
will be sampled uniformly within [0.1, l1_scale] and [0.1, sigma_scale].
testing: Boolean that indicates whether we are testing. If so there are
more targets for visualization.
"""
self._batch_size = batch_size
self._max_num_context = max_num_context
self._x_size = x_size
self._y_size = y_size
self._l1_scale = l1_scale
self._sigma_scale = sigma_scale
self._random_kernel_parameters = random_kernel_parameters
self._testing = testing
def _gaussian_kernel(self, xdata, l1, sigma_f, sigma_noise=2e-2):
"""Applies the Gaussian kernel to generate curve data.
Args:
xdata: Tensor of shape [B, num_total_points, x_size] with
the values of the x-axis data.
l1: Tensor of shape [B, y_size, x_size], the scale
parameter of the Gaussian kernel.
sigma_f: Tensor of shape [B, y_size], the magnitude
of the std.
sigma_noise: Float, std of the noise that we add for stability.
Returns:
The kernel, a float tensor of shape
[B, y_size, num_total_points, num_total_points].
"""
num_total_points = tf.shape(xdata)[1]
# Expand and take the difference
xdata1 = tf.expand_dims(xdata, axis=1) # [B, 1, num_total_points, x_size]
xdata2 = tf.expand_dims(xdata, axis=2) # [B, num_total_points, 1, x_size]
diff = xdata1 - xdata2 # [B, num_total_points, num_total_points, x_size]
# [B, y_size, num_total_points, num_total_points, x_size]
norm = tf.square(diff[:, None, :, :, :] / l1[:, :, None, None, :])
norm = tf.reduce_sum(
norm, -1) # [B, data_size, num_total_points, num_total_points]
# [B, y_size, num_total_points, num_total_points]
kernel = tf.square(sigma_f)[:, :, None, None] * tf.exp(-0.5 * norm)
# Add some noise to the diagonal to make the cholesky work.
kernel += (sigma_noise**2) * tf.eye(num_total_points)
return kernel
def generate_curves(self):
"""Builds the op delivering the data.
Generated functions are `float32` with x values between -2 and 2.
Returns:
A `CNPRegressionDescription` namedtuple.
"""
num_context = tf.random_uniform(
shape=[], minval=3, maxval=self._max_num_context, dtype=tf.int32)
# If we are testing we want to have more targets and have them evenly
# distributed in order to plot the function.
if self._testing:
num_target = 400
num_total_points = num_target
x_values = tf.tile(
tf.expand_dims(tf.range(-2., 2., 1. / 100, dtype=tf.float32), axis=0),
[self._batch_size, 1])
x_values = tf.expand_dims(x_values, axis=-1)
# During training the number of target points and their x-positions are
# selected at random
else:
num_target = tf.random_uniform(shape=(), minval=0,
maxval=self._max_num_context - num_context,
dtype=tf.int32)
num_total_points = num_context + num_target
x_values = tf.random_uniform(
[self._batch_size, num_total_points, self._x_size], -2, 2)
# Set kernel parameters
# Either choose a set of random parameters for the mini-batch
if self._random_kernel_parameters:
l1 = tf.random_uniform([self._batch_size, self._y_size,
self._x_size], 0.1, self._l1_scale)
sigma_f = tf.random_uniform([self._batch_size, self._y_size],
0.1, self._sigma_scale)
# Or use the same fixed parameters for all mini-batches
else:
l1 = tf.ones(shape=[self._batch_size, self._y_size,
self._x_size]) * self._l1_scale
sigma_f = tf.ones(shape=[self._batch_size,
self._y_size]) * self._sigma_scale
# Pass the x_values through the Gaussian kernel
# [batch_size, y_size, num_total_points, num_total_points]
kernel = self._gaussian_kernel(x_values, l1, sigma_f)
# Calculate Cholesky, using double precision for better stability:
cholesky = tf.cast(tf.cholesky(tf.cast(kernel, tf.float64)), tf.float32)
# Sample a curve
# [batch_size, y_size, num_total_points, 1]
y_values = tf.matmul(
cholesky,
tf.random_normal([self._batch_size, self._y_size, num_total_points, 1]))
# [batch_size, num_total_points, y_size]
y_values = tf.transpose(tf.squeeze(y_values, 3), [0, 2, 1])
if self._testing:
# Select the targets
target_x = x_values
target_y = y_values
# Select the observations
idx = tf.random_shuffle(tf.range(num_target))
context_x = tf.gather(x_values, idx[:num_context], axis=1)
context_y = tf.gather(y_values, idx[:num_context], axis=1)
else:
# Select the targets which will consist of the context points as well as
# some new target points
target_x = x_values[:, :num_target + num_context, :]
target_y = y_values[:, :num_target + num_context, :]
# Select the observations
context_x = x_values[:, :num_context, :]
context_y = y_values[:, :num_context, :]
query = ((context_x, context_y), target_x)
return NPRegressionDescription(
query=query,
target_y=target_y,
num_total_points=tf.shape(target_x)[1],
num_context_points=num_context)
# + id="kRTKMWLHVBMq" colab_type="code" colab={}
# periodic kernel
#https://www.cs.toronto.edu/~duvenaud/cookbook/
NPRegressionDescription = collections.namedtuple(
"NPRegressionDescription",
("query", "target_y", "num_total_points", "num_context_points"))
class GPSinCurvesReader(object):
"""Generates curves using a Gaussian Process (GP).
Supports vector inputs (x) and vector outputs (y). Kernel is
mean-squared exponential, using the x-value l2 coordinate distance scaled by
some factor chosen randomly in a range. Outputs are independent gaussian
processes.
"""
def __init__(self,
batch_size,
max_num_context,
x_size=1,
y_size=1,
l1_scale=0.6,
sigma_scale=1.0,
random_kernel_parameters=True,
testing=False):
"""Creates a regression dataset of functions sampled from a GP.
Args:
batch_size: An integer.
max_num_context: The max number of observations in the context.
x_size: Integer >= 1 for length of "x values" vector.
y_size: Integer >= 1 for length of "y values" vector.
l1_scale: Float; typical scale for kernel distance function.
sigma_scale: Float; typical scale for variance.
random_kernel_parameters: If `True`, the kernel parameters (l1 and sigma)
will be sampled uniformly within [0.1, l1_scale] and [0.1, sigma_scale].
testing: Boolean that indicates whether we are testing. If so there are
more targets for visualization.
"""
self._batch_size = batch_size
self._max_num_context = max_num_context
self._x_size = x_size
self._y_size = y_size
self._l1_scale = l1_scale
self._sigma_scale = sigma_scale
self._random_kernel_parameters = random_kernel_parameters
self._testing = testing
def _gaussian_kernel(self, xdata, l1, sigma_f, sigma_noise=2e-2):
"""Applies the Gaussian kernel to generate curve data.
Args:
xdata: Tensor of shape [B, num_total_points, x_size] with
the values of the x-axis data.
l1: Tensor of shape [B, y_size, x_size], the scale
parameter of the Gaussian kernel.
sigma_f: Tensor of shape [B, y_size], the magnitude
of the std.
sigma_noise: Float, std of the noise that we add for stability.
Returns:
The kernel, a float tensor of shape
[B, y_size, num_total_points, num_total_points].
"""
num_total_points = tf.shape(xdata)[1]
# Expand and take the difference
xdata1 = tf.expand_dims(xdata, axis=1) # [B, 1, num_total_points, x_size]
xdata2 = tf.expand_dims(xdata, axis=2) # [B, num_total_points, 1, x_size]
diff = xdata1 - xdata2 # [B, num_total_points, num_total_points, x_size]
# [B, y_size, num_total_points, num_total_points, x_size]
norm = 2*tf.square(tf.math.sin(3.14*diff[:, None, :, :, :])) / l1[:, :, None, None, :]
norm = tf.reduce_sum(
norm, -1) # [B, data_size, num_total_points, num_total_points]
# [B, y_size, num_total_points, num_total_points]
kernel = tf.square(sigma_f)[:, :, None, None] * tf.exp(-norm)
# Add some noise to the diagonal to make the cholesky work.
kernel += (sigma_noise**2) * tf.eye(num_total_points)
return kernel
def generate_curves(self):
"""Builds the op delivering the data.
Generated functions are `float32` with x values between -2 and 2.
Returns:
A `CNPRegressionDescription` namedtuple.
"""
num_context = tf.random_uniform(
shape=[], minval=3, maxval=self._max_num_context, dtype=tf.int32)
# If we are testing we want to have more targets and have them evenly
# distributed in order to plot the function.
if self._testing:
num_target = 400
num_total_points = num_target
x_values = tf.tile(
tf.expand_dims(tf.range(-2., 2., 1. / 100, dtype=tf.float32), axis=0),
[self._batch_size, 1])
x_values = tf.expand_dims(x_values, axis=-1)
# During training the number of target points and their x-positions are
# selected at random
else:
num_target = tf.random_uniform(shape=(), minval=0,
maxval=self._max_num_context - num_context,
dtype=tf.int32)
num_total_points = num_context + num_target
x_values = tf.random_uniform(
[self._batch_size, num_total_points, self._x_size], -2, 2)
# Set kernel parameters
# Either choose a set of random parameters for the mini-batch
if self._random_kernel_parameters:
l1 = tf.random_uniform([self._batch_size, self._y_size,
self._x_size], 0.1, self._l1_scale)
sigma_f = tf.random_uniform([self._batch_size, self._y_size],
0.1, self._sigma_scale)
# Or use the same fixed parameters for all mini-batches
else:
l1 = tf.ones(shape=[self._batch_size, self._y_size,
self._x_size]) * self._l1_scale
sigma_f = tf.ones(shape=[self._batch_size,
self._y_size]) * self._sigma_scale
# Pass the x_values through the Gaussian kernel
# [batch_size, y_size, num_total_points, num_total_points]
kernel = self._gaussian_kernel(x_values, l1, sigma_f)
# Calculate Cholesky, using double precision for better stability:
cholesky = tf.cast(tf.cholesky(tf.cast(kernel, tf.float64)), tf.float32)
# Sample a curve
# [batch_size, y_size, num_total_points, 1]
y_values = tf.matmul(
cholesky,
tf.random_normal([self._batch_size, self._y_size, num_total_points, 1]))
# [batch_size, num_total_points, y_size]
y_values = tf.transpose(tf.squeeze(y_values, 3), [0, 2, 1])
if self._testing:
# Select the targets
target_x = x_values
target_y = y_values
# Select the observations
idx = tf.random_shuffle(tf.range(num_target))
context_x = tf.gather(x_values, idx[:num_context], axis=1)
context_y = tf.gather(y_values, idx[:num_context], axis=1)
else:
# Select the targets which will consist of the context points as well as
# some new target points
target_x = x_values[:, :num_target + num_context, :]
target_y = y_values[:, :num_target + num_context, :]
# Select the observations
context_x = x_values[:, :num_context, :]
context_y = y_values[:, :num_context, :]
query = ((context_x, context_y), target_x)
return NPRegressionDescription(
query=query,
target_y=target_y,
num_total_points=tf.shape(target_x)[1],
num_context_points=num_context)
# + [markdown] id="GhMwui0VfmNn" colab_type="text"
# ## Attentive Neural Processes: a short introduction
#
# Here below are the model diagrams for the **NP** (left) and **ANP** (right).
#
# 
#
# In the NP, the **context points** $(x_C,y_C)=(x_i, y_i)_{i \in C}$ are passed through the encoder that consists of two paths, a **deterministic path** and and a **latent path**.
#
# In the **deterministic path**, each context pair $(x_i,y_i)$ is passed through an MLP (shared parameters across the contexts) to produce representation $r_i$. These are aggregated by taking the mean to produce the deterministic code $r_C$.
#
# In the **latent path**, a code $s_C$ is computed in a similar manner from the representations $s_i$, and is used to parameterise the distribution of the latent variable $z$, giving the latent distribution $q(z|s_C)$.
#
# In the decoder, the $r_C$ and $z$ are concatenated alongside $x_*$ and passed through an MLP to produce the parameters of the distribution $p(y_*|x_*,r_C,z)$.
#
# The motivation for having a global latent is to model different realisations of the data generating stochastic process - each sample of $z$ would correspond to one realisation of the stochastic process. One can define the model using just the deterministic path, just the latent path, or both.
#
# One problem of the NP is that the **mean-aggregation step in the encoder acts as a bottleneck**: since taking the mean across context representations gives the same weight to each context point, it is difficult for the decoder to learn which context points provide relevant information for a given target prediction.
#
# This is addressed by ANPs, where the mean-aggregation is replaced by a **cross-attention mechanism** - the target query $x_*$ attends to the key-value pairs $(x_i,r_i)_{i \in C}$ and assigns weights $w_i$ to each pair to form a query-specific representation $r_*=\sum_i w_i r_i$. This is precisely where the model allows each query to attend more closely to the context points that it deems relevant for the prediction. Note that if we use uniform attention (all $w_i$ equal), then we revert to the NP.
#
# Another change is the **self-attention mechanism** that replaces the MLPs in the encoder, used in order to model interactions between the context points. However for the 1D regression task here, we do not use self-attention and resort to the MLP setting as it is shown to be sufficient, and just use cross-attention.
#
# Learning for both the NP and ANP is done by optimising the ELBO to the log predictive likelihood:
#
# $$\log p(y_T|x_T,x_C,y_C) \geq
# \mathbb{E}_{q(z|s_T)} [\log p(y_T|x_T,r_C,z)] - KL ( q(z|s_T) \Vert q(z|s_C) )$$
# where $C$ represents contexts and $T$ represents targets.
# + id="ps97odopnvkv" colab_type="code" colab={}
# utility methods
def batch_mlp(input, output_sizes, variable_scope):
"""Apply MLP to the final axis of a 3D tensor (reusing already defined MLPs).
Args:
input: input tensor of shape [B,n,d_in].
output_sizes: An iterable containing the output sizes of the MLP as defined
in `basic.Linear`.
variable_scope: String giving the name of the variable scope. If this is set
to be the same as a previously defined MLP, then the weights are reused.
Returns:
tensor of shape [B,n,d_out] where d_out=output_sizes[-1]
"""
# Get the shapes of the input and reshape to parallelise across observations
batch_size, _, filter_size = input.shape.as_list()
output = tf.reshape(input, (-1, filter_size))
output.set_shape((None, filter_size))
# Pass through MLP
with tf.variable_scope(variable_scope, reuse=tf.AUTO_REUSE):
for i, size in enumerate(output_sizes[:-1]):
output = tf.nn.relu(
tf.layers.dense(output, size, name="layer_{}".format(i)))
# Last layer without a ReLu
output = tf.layers.dense(
output, output_sizes[-1], name="layer_{}".format(i + 1))
# Bring back into original shape
output = tf.reshape(output, (batch_size, -1, output_sizes[-1]))
return output
# + [markdown] id="kli9cfXqt2Jf" colab_type="text"
# ## Encoder: Deterministic Path
#
# The encoder in the deterministic path is shared between all context pairs and consists of an MLP and an attention module. Each context $x_i$ and $y_i$ are concatenated and passed through the MLP (with relu non-linearities) to output a representation $r_i$. These $(r_i)_{i \in C}$ and $x_i$ are fed into the cross-attention module, along with query $x_*$ to output a query-specific representation $r_*$. The MLP architecture is given by the `output_sizes` argument, a list of hidden layer sizes, and the `attention` argument which is the cross-attention module defined later on.
# + id="yXdB68gkPwy2" colab_type="code" colab={}
class DeterministicEncoder(object):
"""The Deterministic Encoder."""
def __init__(self, output_sizes, attention, use_self_attention = False):
"""(A)NP deterministic encoder.
Args:
output_sizes: An iterable containing the output sizes of the encoding MLP.
attention: The attention module.
"""
self._output_sizes = output_sizes
self._attention = attention
self._use_self_attention = use_self_attention
def __call__(self, context_x, context_y, target_x):
"""Encodes the inputs into one representation.
Args:
context_x: Tensor of shape [B,observations,d_x]. For this 1D regression
task this corresponds to the x-values.
context_y: Tensor of shape [B,observations,d_y]. For this 1D regression
task this corresponds to the y-values.
target_x: Tensor of shape [B,target_observations,d_x].
For this 1D regression task this corresponds to the x-values.
Returns:
The encoded representation. Tensor of shape [B,target_observations,d]
"""
# Concatenate x and y along the filter axes
encoder_input = tf.concat([context_x, context_y], axis=-1)
if self._use_self_attention:
print('Uaing self attention in the deterministic encoder')
encoder_input = batch_mlp(encoder_input, [2,128,128],"deterministic_encoder_self")
with tf.variable_scope("deterministic_encoder_self", reuse=tf.AUTO_REUSE):
encoder_input = self._attention(encoder_input,encoder_input,encoder_input)
# Pass final axis through MLP
hidden = batch_mlp(encoder_input, self._output_sizes,
"deterministic_encoder")
# Apply attention
with tf.variable_scope("deterministic_encoder", reuse=tf.AUTO_REUSE):
hidden = self._attention(context_x, target_x, hidden)
return hidden
# + [markdown] id="1944scGst6fS" colab_type="text"
# ## Encoder: Latent Path
#
# The encoder in the latent path is again shared by all context pairs and consists of an MLP. After the MLP (with relu non-linearities) is applied the resulting representation is aggregated by taking the mean across the contexts and a further MLP is applied to compute the mean and variance of the Gaussian $q(z|s_C)$. Again the initial MLP's architecture is given by the `output_sizes` argument, and the `num_latent` argument sets the latent dimensionality.
# + id="gufhxfGSRCs9" colab_type="code" colab={}
class LatentEncoder(object):
"""The Latent Encoder."""
def __init__(self, output_sizes, num_latents, attention, use_self_attention):
"""(A)NP latent encoder.
Args:
output_sizes: An iterable containing the output sizes of the encoding MLP.
num_latents: The latent dimensionality.
"""
self._output_sizes = output_sizes
self._num_latents = num_latents
self._attention = attention
self._use_self_attention = use_self_attention
def __call__(self, x, y):
"""Encodes the inputs into one representation.
Args:
x: Tensor of shape [B,observations,d_x]. For this 1D regression
task this corresponds to the x-values.
y: Tensor of shape [B,observations,d_y]. For this 1D regression
task this corresponds to the y-values.
Returns:
A normal distribution over tensors of shape [B, num_latents]
"""
# Concatenate x and y along the filter axes
encoder_input = tf.concat([x, y], axis=-1)
if self._use_self_attention:
print('Using self attention in the latent encoder')
encoder_input = batch_mlp(encoder_input, [2,128,128],"latent_encoder_self")
with tf.variable_scope("latent_encoder_self", reuse=tf.AUTO_REUSE):
encoder_input = self._attention(encoder_input,encoder_input,encoder_input)
# Pass final axis through MLP
hidden = batch_mlp(encoder_input, self._output_sizes, "latent_encoder")
# Aggregator: take the mean over all points
hidden = tf.reduce_mean(hidden, axis=1)
# Have further MLP layers that map to the parameters of the Gaussian latent
with tf.variable_scope("latent_encoder", reuse=tf.AUTO_REUSE):
# First apply intermediate relu layer
hidden = tf.nn.relu(
tf.layers.dense(hidden,
(self._output_sizes[-1] + self._num_latents)/2,
name="penultimate_layer"))
# Then apply further linear layers to output latent mu and log sigma
mu = tf.layers.dense(hidden, self._num_latents, name="mean_layer")
log_sigma = tf.layers.dense(hidden, self._num_latents, name="std_layer")
# Compute sigma
sigma = 0.1 + 0.9 * tf.sigmoid(log_sigma)
return tf.contrib.distributions.Normal(loc=mu, scale=sigma)
# + id="of3idijgOnpG" colab_type="code" colab={}
# TODO: add self-attention as an option
class LatentEncoder_cross(object):
"""The Latent Encoder."""
def __init__(self, output_sizes, num_latents,attention,use_self_attention):
"""(A)NP latent encoder.
Args:
output_sizes: An iterable containing the output sizes of the encoding MLP.
num_latents: The latent dimensionality.
"""
self._output_sizes = output_sizes
print("Using cross attention in the latent encoder ")
self._num_latents = num_latents
self._attention = attention
self._use_self_attention = use_self_attention
def __call__(self, x_cont, y_cont, x_targ):
"""Encodes the inputs into one representation.
Args:
x: Tensor of shape [B,observations,d_x]. For this 1D regression
task this corresponds to the x-values.
y: Tensor of shape [B,observations,d_y]. For this 1D regression
task this corresponds to the y-values.
Returns:
A normal distribution over tensors of shape [B, num_latents]
"""
# Concatenate x and y along the filter axes
encoder_input = tf.concat([x_cont, y_cont], axis=-1)
if self._use_self_attention:
print('Using self attention in the latent cross encoder')
encoder_input = batch_mlp(encoder_input, [2,128,128],"latent_encoder_cross_self")
with tf.variable_scope("latent_encoder_cross_self", reuse=tf.AUTO_REUSE):
encoder_input = self._attention(encoder_input,encoder_input,encoder_input)
# Pass final axis through MLP
hidden_mean = batch_mlp(encoder_input, [2,self._num_latents], "latent_encode_cross_mean")
hidden_var = batch_mlp(encoder_input, [2,self._num_latents], "latent_encode_cross_var")
sigma = 0.1 + 0.9 * tf.sigmoid(hidden_var)
z_samps = tf.contrib.distributions.Normal(loc = hidden_mean, scale = sigma).sample()
with tf.variable_scope("latent_cross", reuse=tf.AUTO_REUSE):
z_star = self._attention(x_cont,x_targ,z_samps)
return z_star, z_samps
# + [markdown] id="roGKUH3Nt9Xg" colab_type="text"
# ## Decoder
#
# The context representation (either $z$ or $z$ and $r_C$ concatenated) and the target inputs $x_T$ are fed into the decoder. First they are concatenated and passed through an MLP, whose architecture is given by the `output_sizes` argument. The MLP outputs the mean and variance of the Gaussian $p(y_T|x_T,r_C,z)$.
# + id="RLhx-J8ALyST" colab_type="code" cellView="both" colab={}
#@title
class Decoder(object):
"""The Decoder."""
def __init__(self, output_sizes, apply_attention=True, attention = None):
"""(A)NP decoder.
Args:
output_sizes: An iterable containing the output sizes of the decoder MLP
as defined in `basic.Linear`.
"""
self._output_sizes = output_sizes
self._apply_attention = apply_attention
if self._apply_attention:
self._attention = attention
print('Decoder Attention is ', self._apply_attention)
def __call__(self, representation, target_x):
"""Decodes the individual targets.
Args:
representation: The representation of the context for target predictions.
Tensor of shape [B,target_observations,?].
target_x: The x locations for the target query.
Tensor of shape [B,target_observations,d_x].
Returns:
dist: A multivariate Gaussian over the target points. A distribution over
tensors of shape [B,target_observations,d_y].
mu: The mean of the multivariate Gaussian.
Tensor of shape [B,target_observations,d_x].
sigma: The standard deviation of the multivariate Gaussian.
Tensor of shape [B,target_observations,d_x].
"""
if self._apply_attention:
print('Using self attention in the decoder')
hidden_decoder = batch_mlp(target_x, [1,128,128],"hidden_decoder")
print("Input Hidden ", hidden_decoder)
with tf.variable_scope("hidden_decoder", reuse=tf.AUTO_REUSE):
hidden_decoder = self._attention(hidden_decoder ,hidden_decoder,hidden_decoder)
print("Attention Hidden ", hidden_decoder)
hidden = tf.concat([representation, hidden_decoder], axis=-1)
else:
hidden = tf.concat([representation, target_x], axis=-1)
# Pass final axis through MLP
hidden = batch_mlp(hidden, self._output_sizes, "decoder")
print("Output hidden ", hidden)
# Get the mean an the variance
mu, log_sigma = tf.split(hidden, 2, axis=-1)
# Bound the variance
sigma = 0.1 + 0.9 * tf.nn.softplus(log_sigma)
# Get the distribution
dist = tf.contrib.distributions.MultivariateNormalDiag(
loc=mu, scale_diag=sigma)
return dist, mu, sigma
# + [markdown] id="bOfbRLHpt_KK" colab_type="text"
# ## Model
#
# We can put the encoders and the decoder together to form the ANP model.
# + id="n5XJ6gr7ZDPl" colab_type="code" colab={}
class LatentModel(object):
"""The (A)NP model."""
def __init__(self, latent_encoder_output_sizes, num_latents,
decoder_output_sizes, use_deterministic_path=True,
deterministic_encoder_output_sizes=None, attention=None,
use_attention_decoder=False,
use_encoder_determ_self_attention = False,
use_encoder_latent_self_attention = False,
use_encoder_latent_cross_attention = False):
"""Initialises the model.
Args:
latent_encoder_output_sizes: An iterable containing the sizes of hidden
layers of the latent encoder.
num_latents: The latent dimensionality.
decoder_output_sizes: An iterable containing the sizes of hidden layers of
the decoder. The last element should correspond to d_y * 2
(it encodes both mean and variance concatenated)
use_deterministic_path: a boolean that indicates whether the deterministic
encoder is used or not.
deterministic_encoder_output_sizes: An iterable containing the sizes of
hidden layers of the deterministic encoder. The last one is the size
of the deterministic representation r.
attention: The attention module used in the deterministic encoder.
Only relevant when use_deterministic_path=True.
"""
self._latent_encoder = LatentEncoder(latent_encoder_output_sizes,
num_latents,attention, use_encoder_latent_self_attention)
self._use_encoder_latent_cross_attention = use_encoder_latent_cross_attention
if use_encoder_latent_cross_attention:
self._latent_encoder_cross = LatentEncoder_cross(latent_encoder_output_sizes,
num_latents,attention,use_encoder_latent_self_attention)
self._decoder = Decoder(decoder_output_sizes, apply_attention = use_attention_decoder, attention = attention)
self._use_deterministic_path = use_deterministic_path
if use_deterministic_path:
self._deterministic_encoder = DeterministicEncoder(
deterministic_encoder_output_sizes, attention, use_encoder_determ_self_attention)
def __call__(self, query, num_targets, target_y=None):
"""Returns the predicted mean and variance at the target points.
Args:
query: Array containing ((context_x, context_y), target_x) where:
context_x: Tensor of shape [B,num_contexts,d_x].
Contains the x values of the context points.
context_y: Tensor of shape [B,num_contexts,d_y].
Contains the y values of the context points.
target_x: Tensor of shape [B,num_targets,d_x].
Contains the x values of the target points.
num_targets: Number of target points.
target_y: The ground truth y values of the target y.
Tensor of shape [B,num_targets,d_y].
Returns:
log_p: The log_probability of the target_y given the predicted
distribution. Tensor of shape [B,num_targets].
mu: The mean of the predicted distribution.
Tensor of shape [B,num_targets,d_y].
sigma: The variance of the predicted distribution.
Tensor of shape [B,num_targets,d_y].
"""
(context_x, context_y), target_x = query
# Pass query through the encoder and the decoder
prior = self._latent_encoder(context_x, context_y)
if self._use_encoder_latent_cross_attention:
latent_cross,z_samps_prior = self._latent_encoder_cross(context_x, context_y, target_x)
# For training, when target_y is available, use targets for latent encoder.
# Note that targets contain contexts by design.
if target_y is None:
latent_rep = prior.sample()
# For testing, when target_y unavailable, use contexts for latent encoder.
else:
posterior = self._latent_encoder(target_x, target_y)
latent_rep = posterior.sample()
if self._use_encoder_latent_cross_attention:
latent_cross,z_samps_posterior = self._latent_encoder_cross(target_x, target_y, target_x)
latent_rep = tf.tile(tf.expand_dims(latent_rep, axis=1),
[1, num_targets, 1])
if self._use_deterministic_path:
deterministic_rep = self._deterministic_encoder(context_x, context_y,
target_x)
if self._use_encoder_latent_cross_attention:
representation = tf.concat([deterministic_rep, latent_cross, latent_rep], axis=-1)
else:
representation = tf.concat([deterministic_rep,latent_rep],axis=-1)
else:
if self._use_encoder_latent_cross_attention:
representation = tf.concat([latent_cross, latent_rep], axis=-1)
else:
representation = latent_rep
dist, mu, sigma = self._decoder(representation, target_x)
# If we want to calculate the log_prob for training we will make use of the
# target_y. At test time the target_y is not available so we return None.
if target_y is not None:
log_p = dist.log_prob(target_y)
posterior = self._latent_encoder(target_x, target_y)
kl = tf.reduce_sum(
tf.contrib.distributions.kl_divergence(posterior, prior),
axis=-1, keepdims=True)
kl = tf.tile(kl, [1, num_targets])
# # TODO: KL for cross_attention part of latent encoder
# kl_cross = tf.reduce_sum(
# tf.contrib.distributions.kl_divergence(z_samps_posterior, z_samps_prior),
# axis=-1, keepdims=True)
# kl_cross = tf.tile(kl_cross, [1, num_targets])
loss = - tf.reduce_mean(log_p - kl / tf.cast(num_targets, tf.float32))
else:
log_p = None
kl = None
loss = None
return mu, sigma, log_p, kl, loss
# + [markdown] id="ULaa6hZEuHvU" colab_type="text"
# ## Cross-Attention Module
# Given a set of key-value pairs $(k_i,v_i)_{i \in I}$ and query $q$, an attention module computes weights for each key and aggregates the values with these weights to form the value corresponding to the query.
#
#
# **`rep`** determines whether the raw inputs to the module will be used as the keys and queries, or whether you will pass them through an MLP and use the output instead. One of 'identity', 'mlp'.
#
# **`output_sizes`** determines the architecture of the MLP used to obtain the keys/queries if `rep` is 'mlp'.
#
# **`att_type`** is a string argument that determines the type of attention used. Valid choices of attention are: uniform, laplace, dot product, multihead.
#
# * **Uniform** $((k_i,v_i)_{i\in I}, q)= \frac{1}{|I|} \sum_i v_i$
#
# * **Laplace** $((k_i,v_i)_{i\in I}, q)= \sum_i w_i v_i, \hspace{2mm} w_i \propto \exp(-\frac{||q - k_i||_1}{l})$
#
# * **DotProduct** $((k_i,v_i)_{i\in I}, q)= \sum_i w_i v_i, \hspace{2mm} w_i \propto \exp(q^\top k_i / \sqrt{d_k})$ where $k_i \in \mathbb{R}^{d_k}$.
#
# * **Multihead** $((k_i,v_i)_{i\in I}, q)= \mathcal{L}^O(\text{concat}(\text{head}_1, \ldots, \text{head}_H))$, $\text{head}_h = \text{DotProduct}((\mathcal{L}^K_h(k_i),\mathcal{L}^V_h(v_i))_{i \in I}, \mathcal{L}^Q_h(q))$
#
# where $\mathcal{L}$ are linear maps with trainable parameters.
#
# **`scale`**: length scale $l$ in Laplace attention.
#
# **`normalise`**: whether to use a softmax so that weights sum to 1 or not.
#
# **`num_heads`**: $H$, the number of heads for multihead attention.
# + id="DImJP8HfhmmM" colab_type="code" cellView="form" colab={}
#@title
def uniform_attention(q, v):
"""Uniform attention. Equivalent to np.
Args:
q: queries. tensor of shape [B,m,d_k].
v: values. tensor of shape [B,n,d_v].
Returns:
tensor of shape [B,m,d_v].
"""
total_points = tf.shape(q)[1]
rep = tf.reduce_mean(v, axis=1, keepdims=True) # [B,1,d_v]
rep = tf.tile(rep, [1, total_points, 1])
return rep
def laplace_attention(q, k, v, scale, normalise):
"""Computes laplace exponential attention.
Args:
q: queries. tensor of shape [B,m,d_k].
k: keys. tensor of shape [B,n,d_k].
v: values. tensor of shape [B,n,d_v].
scale: float that scales the L1 distance.
normalise: Boolean that determines whether weights sum to 1.
Returns:
tensor of shape [B,m,d_v].
"""
k = tf.expand_dims(k, axis=1) # [B,1,n,d_k]
q = tf.expand_dims(q, axis=2) # [B,m,1,d_k]
unnorm_weights = - tf.abs((k - q) / scale) # [B,m,n,d_k]
unnorm_weights = tf.reduce_sum(unnorm_weights, axis=-1) # [B,m,n]
if normalise:
weight_fn = tf.nn.softmax
else:
weight_fn = lambda x: 1 + tf.tanh(x)
weights = weight_fn(unnorm_weights) # [B,m,n]
rep = tf.einsum('bik,bkj->bij', weights, v) # [B,m,d_v]
return rep
def dot_product_attention(q, k, v, normalise):
"""Computes dot product attention.
Args:
q: queries. tensor of shape [B,m,d_k].
k: keys. tensor of shape [B,n,d_k].
v: values. tensor of shape [B,n,d_v].
normalise: Boolean that determines whether weights sum to 1.
Returns:
tensor of shape [B,m,d_v].
"""
#print('I hate it')
d_k = tf.shape(q)[-1]
scale = tf.sqrt(tf.cast(d_k, tf.float32))
unnorm_weights = tf.einsum('bjk,bik->bij', k, q) / scale # [B,m,n]
if normalise:
weight_fn = tf.nn.softmax
else:
weight_fn = tf.sigmoid
weights = weight_fn(unnorm_weights) # [B,m,n]
rep = tf.einsum('bik,bkj->bij', weights, v) # [B,m,d_v]
return rep
def multihead_attention(q, k, v, num_heads=8):
"""Computes multi-head attention.
Args:
q: queries. tensor of shape [B,m,d_k].
k: keys. tensor of shape [B,n,d_k].
v: values. tensor of shape [B,n,d_v].
num_heads: number of heads. Should divide d_v.
Returns:
tensor of shape [B,m,d_v].
"""
d_k = q.get_shape().as_list()[-1]
d_v = v.get_shape().as_list()[-1]
head_size = d_v / num_heads
key_initializer = tf.random_normal_initializer(stddev=d_k**-0.5)
value_initializer = tf.random_normal_initializer(stddev=d_v**-0.5)
rep = tf.constant(0.0)
for h in range(num_heads):
o = dot_product_attention(
tf.layers.Conv1D(head_size, 1, kernel_initializer=key_initializer,
name='wq%d' % h, use_bias=False, padding='VALID')(q),
tf.layers.Conv1D(head_size, 1, kernel_initializer=key_initializer,
name='wk%d' % h, use_bias=False, padding='VALID')(k),
tf.layers.Conv1D(head_size, 1, kernel_initializer=key_initializer,
name='wv%d' % h, use_bias=False, padding='VALID')(v),
normalise=True)
rep += tf.layers.Conv1D(d_v, 1, kernel_initializer=value_initializer,
name='wo%d' % h, use_bias=False, padding='VALID')(o)
return rep
class Attention(object):
"""The Attention module."""
def __init__(self, rep, output_sizes, att_type, scale=1., normalise=True,
num_heads=8):
"""Create attention module.
Takes in context inputs, target inputs and
representations of each context input/output pair
to output an aggregated representation of the context data.
Args:
rep: transformation to apply to contexts before computing attention.
One of: ['identity','mlp'].
output_sizes: list of number of hidden units per layer of mlp.
Used only if rep == 'mlp'.
att_type: type of attention. One of the following:
['uniform','laplace','dot_product','multihead']
scale: scale of attention.
normalise: Boolean determining whether to:
1. apply softmax to weights so that they sum to 1 across context pts or
2. apply custom transformation to have weights in [0,1].
num_heads: number of heads for multihead.
"""
self._rep = rep
self._output_sizes = output_sizes
self._type = att_type
self._scale = scale
self._normalise = normalise
if self._type == 'multihead':
self._num_heads = num_heads
def __call__(self, x1, x2, r):
"""Apply attention to create aggregated representation of r.
Args:
x1: tensor of shape [B,n1,d_x].
x2: tensor of shape [B,n2,d_x].
r: tensor of shape [B,n1,d].
Returns:
tensor of shape [B,n2,d]
Raises:
NameError: The argument for rep/type was invalid.
"""
if self._rep == 'identity':
k, q = (x1, x2)
elif self._rep == 'mlp':
# Pass through MLP
k = batch_mlp(x1, self._output_sizes, "attention")
q = batch_mlp(x2, self._output_sizes, "attention")
else:
raise NameError("'rep' not among ['identity','mlp']")
if self._type == 'uniform':
rep = uniform_attention(q, r)
elif self._type == 'laplace':
rep = laplace_attention(q, k, r, self._scale, self._normalise)
elif self._type == 'dot_product':
rep = dot_product_attention(q, k, r, self._normalise)
elif self._type == 'multihead':
rep = multihead_attention(q, k, r, self._num_heads)
else:
raise NameError(("'att_type' not among ['uniform','laplace','dot_product'"
",'multihead']"))
return rep
# + [markdown] id="zYcTQDLltFmA" colab_type="text"
# ## Ploting function
#
# Same plotting function as for the [implementation of CNPs](https://github.com/deepmind/conditional-neural-process/blob/master/conditional_neural_process.ipynb) that plots the intermediate predictions every so often during training.
# + id="AFlT3lJQTM_m" colab_type="code" cellView="both" colab={}
#@title
def plot_functions(target_x, target_y, context_x, context_y, pred_y, std):
"""Plots the predicted mean and variance and the context points.
Args:
target_x: An array of shape [B,num_targets,1] that contains the
x values of the target points.
target_y: An array of shape [B,num_targets,1] that contains the
y values of the target points.
context_x: An array of shape [B,num_contexts,1] that contains
the x values of the context points.
context_y: An array of shape [B,num_contexts,1] that contains
the y values of the context points.
pred_y: An array of shape [B,num_targets,1] that contains the
predicted means of the y values at the target points in target_x.
std: An array of shape [B,num_targets,1] that contains the
predicted std dev of the y values at the target points in target_x.
"""
# Plot everything
plt.plot(target_x[0], pred_y[0], 'b', linewidth=2)
plt.plot(target_x[0], target_y[0], 'k:', linewidth=2)
plt.plot(context_x[0], context_y[0], 'ko', markersize=10)
plt.fill_between(
target_x[0, :, 0],
pred_y[0, :, 0] - std[0, :, 0],
pred_y[0, :, 0] + std[0, :, 0],
alpha=0.2,
facecolor='#65c9f7',
interpolate=True)
# Make the plot pretty
plt.yticks([-2, 0, 2], fontsize=16)
plt.xticks([-2, 0, 2], fontsize=16)
plt.ylim([-2, 2])
plt.grid('off')
ax = plt.gca()
plt.show()
# + [markdown] id="Z69Ham1nteeD" colab_type="text"
# ## Training the (A)NP
#
# We can now start training. First we need to define some variables:
#
# **`TRAINING_ITERATIONS`**: Number of iterations used for trianing. At each iteration we sample a new batch of sample curves from GPs and pick a random set of points on each curve to be the target and a subset to be the context. We optimise the ELBO on the log predictive likelihood of the target given context.
#
# **`MAX_CONTEXT_POINTS`**: Maximum number of context points used during training. This is also set to be the upper bound on the number of target points.
#
# **`PLOT_AFTER`**: The number of iterations between the intermediate plots.
#
# **`HIDDEN_SIZE`**: Master parameter that governs the hidden layer size of all MLPs in the model and also the latent dimensionality.
#
# **`MODEL_TYPE`**: 'NP' or 'ANP'.
#
# **`ATTENTION_TYPE`**: The type of attention used for ANP. One of `uniform`, `laplace` `dot_product` or `multihead`
#
# **`random_kernel_parameters`**: Boolean to determine whether the GP kernel parameters are sample randomly for each iteration or fixed.
#
# + [markdown] id="yHqnv8FP4Rtu" colab_type="text"
# ### NP training
# First we train the NP. Notice from the plots that the predictions after 1e5 iterations do not go through the context points perfectly - i.e. underfits.
# + id="c6FcSLfnLD9_" colab_type="code" outputId="ba763990-e51f-4acc-bfda-818ba55589f9" colab={"base_uri": "https://localhost:8080/", "height": 54}
'''TRAINING_ITERATIONS = 100000 #@param {type:"number"}
MAX_CONTEXT_POINTS = 50 #@param {type:"number"}
PLOT_AFTER = 10000 #@param {type:"number"}
HIDDEN_SIZE = 128 #@param {type:"number"}
MODEL_TYPE = 'NP' #@param ['NP','ANP']
ATTENTION_TYPE = 'dot_product' #@param ['uniform','laplace','dot_product','multihead']
random_kernel_parameters=True #@param {type:"boolean"}
tf.reset_default_graph()
# Train dataset
dataset_train = GPCurvesReader(
batch_size=16, max_num_context=MAX_CONTEXT_POINTS, random_kernel_parameters=random_kernel_parameters)
data_train = dataset_train.generate_curves()
# Test dataset
dataset_test = GPCurvesReader(
batch_size=1, max_num_context=MAX_CONTEXT_POINTS, testing=True, random_kernel_parameters=random_kernel_parameters)
data_test = dataset_test.generate_curves()
# Sizes of the layers of the MLPs for the encoders and decoder
# The final output layer of the decoder outputs two values, one for the mean and
# one for the variance of the prediction at the target location
latent_encoder_output_sizes = [HIDDEN_SIZE]*4
num_latents = HIDDEN_SIZE
deterministic_encoder_output_sizes= [HIDDEN_SIZE]*4
decoder_output_sizes = [HIDDEN_SIZE]*2 + [2]
use_deterministic_path = True
# ANP with multihead attention
if MODEL_TYPE == 'ANP':
attention = Attention(rep='mlp', output_sizes=[HIDDEN_SIZE]*2,
att_type=ATTENTION_TYPE)
# NP - equivalent to uniform attention
elif MODEL_TYPE == 'NP':
attention = Attention(rep='identity', output_sizes=None, att_type='uniform')
else:
raise NameError("MODEL_TYPE not among ['ANP,'NP']")
# Define the model
model = LatentModel(latent_encoder_output_sizes, num_latents,
decoder_output_sizes, use_deterministic_path,
deterministic_encoder_output_sizes, attention)
# Define the loss
_, _, log_prob, _, loss = model(data_train.query, data_train.num_total_points,
data_train.target_y)
# Get the predicted mean and variance at the target points for the testing set
mu, sigma, _, _, _ = model(data_test.query, data_test.num_total_points)
# Set up the optimizer and train step
optimizer = tf.train.AdamOptimizer(1e-4)
train_step = optimizer.minimize(loss)
init = tf.initialize_all_variables()
# Train and plot
with tf.Session() as sess:
sess.run(init)
for it in range(TRAINING_ITERATIONS):
sess.run([train_step])
# Plot the predictions in `PLOT_AFTER` intervals
if it % PLOT_AFTER == 0:
loss_value, pred_y, std_y, target_y, whole_query = sess.run(
[loss, mu, sigma, data_test.target_y,
data_test.query])
(context_x, context_y), target_x = whole_query
print('Iteration: {}, loss: {}'.format(it, loss_value))
# Plot the prediction and the context
plot_functions(target_x, target_y, context_x, context_y, pred_y, std_y)'''
# + [markdown] id="pudKobiUJbbT" colab_type="text"
# ### ANP training
#
# Next, we train the ANP with multihead attention, using the same hyperparameter setting as NPs. Note that the predictions are now much more accurate for the observed context data.
# + id="q6wQwAwjTEc_" colab_type="code" outputId="b6fddf7b-a55a-4454-e0d9-ebfd22f5e07a" colab={"base_uri": "https://localhost:8080/", "height": 2322}
TRAINING_ITERATIONS = 30000 #@param {type:"number"}
MAX_CONTEXT_POINTS = 50 #@param {type:"number"}
PLOT_AFTER = 10000 #@param {type:"number"}
HIDDEN_SIZE = 128 #@param {type:"number"}
MODEL_TYPE = 'ANP' #@param ['NP','ANP']
ATTENTION_TYPE = 'dot_product' #@param ['uniform','laplace','dot_product','multihead']
random_kernel_parameters=True #@param {type:"boolean"}
use_decoder_self_attention = True #@param {type:"boolean"}
use_encoder_determ_self_attention = True #@param {type:"boolean"}
use_encoder_latent_self_attention = True #@param {type:"boolean"}
use_encoder_latent_cross_attention = True #@param {type:"boolean"}
filename = 'loss_arr_{}_{}_{}_{}.npy'.format(use_decoder_self_attention,
use_encoder_determ_self_attention,
use_encoder_latent_self_attention,
use_encoder_latent_cross_attention)
tf.reset_default_graph()
# Train dataset
dataset_train = GPSinCurvesReader(
batch_size=16, max_num_context=MAX_CONTEXT_POINTS)
data_train = dataset_train.generate_curves()
# Test dataset
dataset_test = GPSinCurvesReader(
batch_size=1, max_num_context=MAX_CONTEXT_POINTS, testing=True)
data_test = dataset_test.generate_curves()
# Sizes of the layers of the MLPs for the encoders and decoder
# The final output layer of the decoder outputs two values, one for the mean and
# one for the variance of the prediction at the target location
latent_encoder_output_sizes = [HIDDEN_SIZE]*4
num_latents = HIDDEN_SIZE
deterministic_encoder_output_sizes= [HIDDEN_SIZE]*4
decoder_output_sizes = [HIDDEN_SIZE]*2 + [2]
use_deterministic_path = True
# ANP with multihead attention
if MODEL_TYPE == 'ANP':
attention = Attention(rep='mlp', output_sizes=[HIDDEN_SIZE]*2,
att_type='multihead')
# NP - equivalent to uniform attention
elif MODEL_TYPE == 'NP':
attention = Attention(rep='identity', output_sizes=None, att_type='uniform')
else:
raise NameError("MODEL_TYPE not among ['ANP,'NP']")
# Define the model
model = LatentModel(latent_encoder_output_sizes, num_latents,
decoder_output_sizes, use_deterministic_path,
deterministic_encoder_output_sizes, attention,
use_attention_decoder = use_decoder_self_attention,
use_encoder_determ_self_attention = use_encoder_determ_self_attention,
use_encoder_latent_self_attention = use_encoder_latent_self_attention,
use_encoder_latent_cross_attention = use_encoder_latent_cross_attention)
# Define the loss
_, _, log_prob, _, loss = model(data_train.query, data_train.num_total_points,
data_train.target_y)
# Get the predicted mean and variance at the target points for the testing set
mu, sigma, _, _, _ = model(data_test.query, data_test.num_total_points)
# Set up the optimizer and train step
optimizer = tf.train.AdamOptimizer(1e-4)
train_step = optimizer.minimize(loss)
init = tf.initialize_all_variables()
# set up loss array
loss_arr = np.zeros(TRAINING_ITERATIONS)
# Train and plot
with tf.train.MonitoredSession() as sess:
sess.run(init)
for it in range(TRAINING_ITERATIONS):
sess.run([train_step])
# Plot the predictions in `PLOT_AFTER` intervals
if it % PLOT_AFTER == 0:
loss_value, pred_y, std_y, target_y, whole_query = sess.run(
[loss, mu, sigma, data_test.target_y,
data_test.query])
(context_x, context_y), target_x = whole_query
print('Iteration: {}, loss: {}'.format(it, loss_value))
# Plot the prediction and the context
plot_functions(target_x, target_y, context_x, context_y, pred_y, std_y)
if it%PLOT_AFTER == 0:
np.save(filename,loss_arr)
loss_arr[it] = loss_value
# + id="XLbGNC_bXaUx" colab_type="code" outputId="aa7ce5a2-cf01-4234-c542-f420faf9edcd" colab={"base_uri": "https://localhost:8080/", "height": 558}
loss_arr[29999]
np.save('loss_sin_changes',loss_arr)
files.download('loss_sin_changes.npy')
# + id="rz-TD-ss9gWx" colab_type="code" outputId="ac2c086e-9d31-42a6-f1f9-b9e6bff07fe7" colab={"base_uri": "https://localhost:8080/", "height": 34}
len(loss_arr)
| notebooks/ANP_workingversion.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TP N°3 - Potencia
# __UNLZ - Facultad de Ingeniería__
# __Electrotecnia__
# __Alumno:__ <NAME>
# <a href="https://colab.research.google.com/github/daniel-lorenzo/Electrotecnia/blob/master/Ejercitacion/TP4entrega.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
# ## Ejercicio N°1
# Calcular las potencias activa (P), reactiva (Q) y aparente (S) del siguiente circuito y dibujar el fasorial de potencias.
# <img src="img/tp4ep1.png">
# ### Solución
# __Datos:__
# $\left\{
# \begin{array}{l}
# Z_1 = 10 + j10 \, \Omega \\
# Z_2 = 0 + j10 \, \Omega \\
# Z_3 = 100 + j0 \, \Omega \\
# U_\mathrm{rms} = 220 \, \mathrm{V}
# \end{array}
# \right.$
# Cálculo de $Z_{eq} = (Z_2 + Z_3) \, || \, Z_1$
# $$ Z_{eq} = {Z_1 \cdot (Z_2 + Z_3) \over Z_1 + Z_2 + Z_3 } = 12,71 \angle 40,41^\circ \, \Omega = 9,68 + j8,24 \, \Omega $$
# Cálculo de $I_\mathrm{rms}$
# $$ I_\mathrm{rms} = {U_\mathrm{rms} \over Z_{eq} } = 17,31 \angle -40,41^\circ \, \mathrm{A} $$
# Cálculo de $S$
# $$ S = U_\mathrm{rms} I_\mathrm{rms}^* = 3807,37 \angle 40,41^\circ \, \mathrm{VA} $$
# $$ S = 2899,21 + j2467,92 \, \mathrm{VA} $$
# Entonces
# $$ P = 2899,21 \, \mathrm{W} $$
# $$ Q = 2467,92 \, \mathrm{VAr} $$
import math, cmath
# Datos:
Z1 = 10 + 10j # Ohm
Z2 = 0 + 10j # Ohm
Z3 = 100 + 0j # Ohm
Urms = 220 # V
# Defino una función que calcula elementos en paralelo
def prl(x,y):
return (x*y)/(x + y)
# (Z2 + Z3) || Z1
Zeq = prl(Z2 + Z3, Z1)
Irms = Urms/Zeq
# Potencia aparente (S):
S = Urms*( Irms.conjugate() )
phi = cmath.phase(S)
# Potencia activa (P):
P = S.real
# Potencia reactiva (Q):
Q = S.imag
print('Resultados:')
print('---------------------------------------')
print('Zeq = (%.2f < %.2f°) Ohm'%(abs(Zeq) , math.degrees( cmath.phase(Zeq) ) ) )
print('Zeq = {:.2f} Ohm'.format(Zeq))
print('Irms = (%.2f < %.2f°) A'%(abs(Irms) , math.degrees( cmath.phase(Irms) ) ))
print('---------------------------------------')
print('S = (%.2f < %.2f°) VA'%(abs(S) , math.degrees( phi ) ))
print('S = {:.2f} VA'.format(S))
print('P = %.2f W'%P)
print('Q = %.2f VAr'%abs(Q) )
print('---------------------------------------')
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.figure(figsize=(7,7))
ax = plt.gca()
ax.quiver(0,0,P,Q,angles='xy',scale_units='xy',scale=1,color='red')
ax.quiver(0,0,P,0,angles='xy',scale_units='xy',scale=1,color='blue')
ax.quiver(P,0,0,Q,angles='xy',scale_units='xy',scale=1,color='green')
plt.text(P - 250, Q, r'$\vec S$', fontsize=18, color='red', fontweight='bold')
plt.text(P - 200, 100, r'$\vec P$', fontsize=18, color='blue', fontweight='bold')
plt.text(P + 100, Q, r'$\vec Q$', fontsize=18, color='green', fontweight='bold')
plt.text(500, 200, r'$\varphi =$ %.2f°'%( math.degrees(phi) ), fontsize=14)
theta = np.linspace(0, phi, 100)
x1 = 500 * np.cos(theta)
x2 = 500 * np.sin(theta)
plt.plot(x1, x2, color='red')
ax.set_xlim([0,3500])
ax.set_ylim([-500,3000])
ax.set_aspect('equal')
plt.title('Triángulo de potencias', fontsize=18)
plt.xlabel('Re (Eje real)', fontsize=16)
plt.ylabel('Im (Eje imaginario)', fontsize=16)
plt.grid(linestyle=":")
ax.set_axisbelow(True)
plt.draw()
plt.show()
# %reset -s -f
# ## Ejercicio N°2
# Calcular el valor de la capacidad para que $\cos \varphi = 0,9$
# <img src="img/tp4ep2.png">
# ### Solución
# __Datos:__
# $\left\{
# \begin{array}{l}
# Z_1 = 10 + j5 \, \Omega \\
# Z_2 = 10 + j100 \, \Omega \\
# U = 100 \, \mathrm{V \; (rms)} \\
# \mathrm{fp} = 0,9 \\
# f = 50 \, \mathrm{Hz}
# \end{array}
# \right.
# $
# Cálculo de $Z_{eq1} = Z_1 || Z_2$
# $$ Z_{eq1} = { Z_1 \times Z_2 \over Z_1 + Z_2 } = 10,51 < 31.64^\circ \, \Omega = 8,95 + j5,51 \, \Omega $$
# Cálculo de $I_1$
# $$ I_1 = {U \over Z_{eq1}} = 9,51 \angle -31,54^\circ \, \mathrm{A} $$
# Cálculo de $S_1$
# $$ S_1 = U \times I_1^* = 951,29 \angle 31,64^\circ \, \mathrm{VA}$$
# $$ S_1 = 809,90 + j499,01 \, \mathrm{VA} $$
# Entonces:
# $$ P_1 = 809,90 \, \mathrm{W} $$
# $$ Q_1 = 499,01 \, \mathrm{VAr} $$
# $$\begin{array}{c}
# P_1 = P_2 \\
# |U| |I_1| \cos \varphi_1 = |U| |I_2| \cos \varphi_2 \\
# |I_1| \cos \varphi_1 = |I_2| \cos \varphi_2
# \end{array}$$
# Entonces
# $$\begin{array}{l}
# | I_2 | &= \displaystyle | I_1 | {\cos \varphi_1 \over \cos \varphi_2} \\
# | I_2 | &= \displaystyle | I_1 | {\cos \varphi_1 \over \mathrm{fp} } \\
# |I_2| &= 9,00 \, \mathrm{A}
# \end{array}$$
# Cálculo de $\varphi_2$
# $$ \mathrm{fp} = \cos \varphi_2 = 0,9 $$
# $$ \varphi_2 = \arccos 0,9 = 25,84^\circ $$
# Cálculo de $S_2$
# $$ S_2 = U I_2 \cos (\varphi_2) = 899,89 \angle 25,84^\circ \, \mathrm{VA} $$
# $$ S_2 = 809,90 + j392,25 \, \mathrm{VA} $$
# Luego
# $$ Q_2 = 392,25 \, \mathrm{VAr} $$
# __Cálculo de $Q_C$__
# $$ Q_C = Q_1 - Q_2 = 106,76 \, \mathrm{VAr} $$
# $$ Q_C = \frac{U^2}{X_C} = \frac{U^2}{ 1 \over \omega C } $$
# __Entonces__
# $$ C = \frac{Q_C}{2 \pi f U^2} = 33,98 \, \mu \mathrm{F} $$
import math, cmath
# Datos:
Z1 = 10 + 5j # Ohm
Z2 = 10 + 100j # Ohm
U = 100 # V
fp = 0.9
f = 50 # Hz
def prl(x,y):
return (x*y)/(x + y)
#Cálculo de Zeq
Zeq1 = prl(Z1,Z2)
# Cálculo de I1
I1 = U/Zeq1
S1 = U*I1.conjugate()
P1 = S1.real
Q1 = S1.imag
P2 = P1
phi1 = cmath.phase(S1)
# P1 = P2 --> U*I1*cos(phi1) = U*I2*cos(phi2)
# y tenemos que fp = cos(phi2)
# Cálculo de phi2
phi2 = math.acos(fp)
I2 = cmath.rect( abs(I1)*math.cos(phi1)/fp , -phi2)
S2 = U*I2.conjugate()
Q2 = S2.imag
# Cálculo de Qc
Qc = Q1 - Q2
# Cálculo de C
C = Qc/(2*math.pi*f*U**2)
# + jupyter={"source_hidden": true}
print('Resultados:')
print('---------------------------------------')
print('Zeq1 = (%.2f < %.2f°) Ohm'%(abs(Zeq1) , math.degrees( cmath.phase(Zeq1) ) ))
print('Zeq1 = {:.2f} Ohm'.format(Zeq1))
print('I1 = (%.2f < %.2f°) A'%(abs(I1), math.degrees( cmath.phase(I1) )))
print('I2 = (%.2f < %.2f) A'%(abs(I2) , math.degrees( cmath.phase(I2) ) ))
print('---------------------------------------')
print('S1 = (%.2f < %.2f°) VA'%(abs(S1) , math.degrees( cmath.phase(S1) ) ))
print('S1 = {:.2f} VA'.format(S1))
print('P1 = %.2f W'%P1)
print('Q1 = %.2f VAr'%Q1)
print('---------------------------------------')
print('S2 = (%.2f < %.2f°) VA'%(abs(S2), math.degrees( cmath.phase(S2) ) ))
print('S2 = {:.2f} VA'.format(S2) )
print('P2 = %.2f W'%P2)
print('Q2 = %.2f VAr'%(abs(Q2)))
print('---------------------------------------')
print('Qc = %.2f VAr'%Qc)
print('C = %.2f uF'%(C*1e6) )
print('---------------------------------------')
# -
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.figure(figsize=(7,7))
ax = plt.gca()
ax.quiver(0,0,P1,Q1,angles='xy',scale_units='xy',scale=1,color='red')
ax.quiver(0,0,P1,0,angles='xy',scale_units='xy',scale=1,color='blue')
ax.quiver(P1,0,0,Q1,angles='xy',scale_units='xy',scale=1,color='green')
ax.quiver(P2 + 10,0,0,Q2,angles='xy',scale_units='xy',scale=1,color='orange')
ax.quiver(0,0,P2,Q2,angles='xy',scale_units='xy',scale=1,color='orange')
plt.text(P1 - 100, Q1, r'$\vec S_1$', fontsize=18, color='red', fontweight='bold')
plt.text(P1 - 90, 30, r'$\vec P_1$', fontsize=18, color='blue', fontweight='bold')
plt.text(P1 + 20, Q1, r'$\vec Q_1$', fontsize=18, color='green', fontweight='bold')
plt.text(P2 - 80, Q2, r'$\vec S_2$', fontsize=18, color='orange', fontweight='bold')
plt.text(P2 + 20, Q2, r'$\vec Q_2$', fontsize=18, color='orange', fontweight='bold')
plt.text(200, 50, r'$\varphi_1 =$ %.2f°'%( math.degrees(phi1) ), fontsize=14)
plt.text(500, 120, r'$\varphi_2 =$ %.2f°'%( math.degrees(phi2) ), fontsize=14)
theta1 = np.linspace(0, phi1, 100)
x1 = 200 * np.cos(theta1)
x2 = 200 * np.sin(theta1)
plt.plot(x1, x2, color='red', linestyle="--")
theta2 = np.linspace(0, phi2, 100)
x3 = 500 * np.cos(theta2)
x4 = 500 * np.sin(theta2)
plt.plot(x3, x4, color='orange', linestyle="--")
ax.set_xlim([0,1000])
ax.set_ylim([-100,800])
ax.set_aspect('equal')
plt.title('Triángulo de potencias', fontsize=18)
plt.xlabel('Re (Eje real)', fontsize=14)
plt.ylabel('Im (Eje imaginario)', fontsize=14)
plt.grid(linestyle=":")
ax.set_axisbelow(True)
plt.draw()
plt.show()
# ----------
#
# <a href="https://colab.research.google.com/github/daniel-lorenzo/Electrotecnia/blob/master/Ejercitacion/TP4entrega.ipynb"><img align="left" src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open in Colab" title="Open and Execute in Google Colaboratory"></a>
| Ejercitacion/TP4entrega.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
originalDataframe = pd.read_csv("Categorical.csv")
dataframe = originalDataframe.copy()
dataframe = dataframe.loc[:, ["Longitude", "Latitude"]]
dataframe.head()
# Select number of clusters
k = 4
# Specify list of colors
colors = ["black", "red", "yellow", "green", "blue", "purple", "pink", "olive", "darkoragne", "cyan"]
def distance(a, b):
d = 0
for x1, x2 in zip(a, b):
d += (x1 - x2)**2
d = d**(1/2)
return d
def KMeans(df, k, means):
clusters = {}
for i in range(k):
clusterName = "c"+ str(i)
clusters[clusterName] = list()
values = df.values.tolist()
# Step 1: calculate distance between data points and the mean. Then assign each point to a cluster
for row in values:
distances = []
for c in range(k):
distances.append(distance(row, means[c]))
cluster = np.argmin(distances)
chosenClusterName = "c"+str(cluster)
clusters[chosenClusterName].append(row)
return clusters
# +
# Random means
means = dataframe.sample(k).values.tolist()
for i in range(4):
print("Iteration number: {}".format(i+1))
print("Clusters' means: {}".format(means))
clusters = KMeans(dataframe, k, means)
newMeans= []
for c in range(k):
newMeans.append(np.array(clusters["c"+str(c)]).mean(axis=0))
means = newMeans
plt.figure(figsize=(15, 10))
for c in range(k):
for p in clusters['c'+str(c)]:
plt.scatter(p[0], p[1], color=colors[c])
plt.show()
| Assignment 4/KMeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # 02 :: Parse meta information
#
# **Objectives:**
#
# * Read the meta data into a dataframe, save to parquet
# * Properly turn columns into
# +
import pandas as pd
import numpy as np
import os
import matplotlib as mpl
import matplotlib.pyplot as plt
import seaborn as sns
sns.set(style="whitegrid")
# -
raw_RData_path = '/media/tmo/data/work/datasets/02_ST/raw/raw_filtered.RData'
raw_parquet = '/media/tmo/data/work/datasets/02_ST/raw/raw.parquet'
lcpm_parquet = '/media/tmo/data/work/datasets/02_ST/lcpm/lcpm.parquet'
meta_parquet = '/media/tmo/data/work/datasets/02_ST/meta/meta.parquet'
wd = '/media/tmo/data/work/datasets/02_ST/ST_structure/'
slides_meta_file = wd + 'spatial_transcriptomics_slide.tsv'
# +
slide_IDs = [
'B02_D1', 'B02_E1', 'B03_C2', 'B03_D2', 'B04_D1',
'B04_E1', 'B05_D2', 'B05_E2', 'B06_E1', 'B07_C2',
'N02_C1', 'N02_D1', 'N03_C2', 'N03_D2', 'N04_D1',
'N04_E1', 'N05_C2', 'N05_D2', 'N06_D2', 'N07_C1']
def add_slide_ID(df):
df['slide_ID'] = df['spot_UID'].apply(lambda x: str(x).split('__')[0])
return df
def to_slide(slide_or_idx):
if isinstance(slide_or_idx, int):
return slide_IDs[slide_or_idx]
else:
return slide_or_idx
def read_slide_meta(slide_or_idx, wd=wd):
"""
Read the metadata of 1 slide, specified by slide name or index.
"""
file = '{0}{1}/metadata/{1}.meta.tsv'.format(wd, to_slide(slide_or_idx))
df = pd.read_csv(file, sep='\t', index_col=0)
df.index.name='spot_UID'
df = df.reset_index()
# Add slide ID
df = add_slide_ID(df)
# Region categorical
df['region'] = df['Region_predict'].astype('category')
df.drop('Region_predict', axis=1)
return df
def read_slides_meta(file=slides_meta_file):
"""
Read the .csv file with meta data about the slides.
"""
COLUMNS = ['Sample id', 'GenotypeShort', 'Age (day)', 'Age (month)']
df = pd.read_csv(file, sep='\t')[COLUMNS]
df.columns = ['sampleID', 'GT', 'age_days', 'age_months']
df['GT'] = df['GT'].astype('category')
# Add age column (young, old)
df['age'] = np.where(df['age_months'] < 10, 'young', 'old')
# Add combined column age_GT.
df['age_GT'] = df[['age', 'GT']].apply(lambda x: '_'.join(x), axis=1)
df['slide_ID'] = df['sampleID']
df.drop('sampleID', axis=1)
return pd.DataFrame(slide_IDs, columns=['slide_ID']).merge(df, how='inner', on='slide_ID')
# -
# ## Parse all meta data, combine and write to Parquet
slides_meta_df = read_slides_meta()
# +
acc = []
for idx in range(0, 20):
acc.append(read_slide_meta(idx))
meta_df = pd.concat(acc)
# -
all_meta_df = meta_df.merge(slides_meta_df, on=['slide_ID'])
all_meta_df.info()
# %%time
all_meta_df.to_parquet(meta_parquet, engine='pyarrow', compression='snappy')
| notebooks/02_from_raw/02_parse_meta.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="3lDkywn1NeyV"
# # STag Demonstration
#
# The following iPython Jupyter notebook gives a step-by-step demonstration of how to use STag to get the tag probabilities and the predicted class for a spectra.
#
# # Setup
#
# The first step is to read in the beta values for each of the tags as well as an example spectrum (this can be modified to read in an appropriate spectrum of your choice).
#
# + colab={"base_uri": "https://localhost:8080/", "height": 375} id="Ta4ahtHDK9DV" outputId="72ae8a41-2f2d-4a4f-f037-7758b52989fe"
import beta_reader
import numpy as np
import os
path = os.getcwd()
beta = beta_reader.beta_reader(path)
spectra = '%s/DES15C2aty_C2_combined_150917_v03_b00.fits' % path
name = 'DES15C2aty'
z = 0.149
# + [markdown] id="o2UPzpBoQ2Jh"
# # Pre-processing
#
# In order to use STag, spectra need to be pre-processed appropriately. This involves filtering, de-redshifting, binning, continuum removal, apodisation, and scaling.
#
# All of these steps are handled by the spectra_preprocessing package, which largely uses methods made for the software [DASH](https://github.com/daniel-muthukrishna/astrodash).
# + id="aQoFeWAvQ2bV"
import spectra_preprocessing as sp
from astropy.io import fits
#Read in the fits file of the spectra and extract the flux and wavelength
fits_file = spectra
table = fits.open(fits_file)
flux = table[0].data
w0 = table[0].header['CRVAL1']
dw = table[0].header['CDELT1']
p0 = table[0].header['CRPIX1']
nlam = len(flux)
wave = w0+dw*(np.arange(nlam, dtype='d')-p0)
table.close()
full = np.column_stack((wave, flux))
#Initialise for pre-processing
preProcess = sp.PreProcessing(full, 2500, 10000, 1024)
#Do the pre-processing steps
sfWave, sfFlux, minInd, maxInd, sfZ, sfArea = preProcess.two_column_data(z, smooth=6, minWave=2500, maxWave=10000)
#Do scaling
flux_pro = sfFlux/sfArea
# + [markdown] id="L-sUbK8ZkgT2"
# # Cutting the Spectra
#
# Many of the tags use specific wavelength ranges of the spectrum rather than the whole thing and so we create multiple instances of the original spectrum cut at the corresponding wavelengths for each tag.
# + id="N1EcHncfl0qO"
class feature_data(object):
"""a class for holding the wavelength and flux for a specific tag."""
def __init__(self, label):
self.label = label
self.wavelength = []
self.flux = []
cuts = np.genfromtxt('%s/cuts.txt' % path, dtype=int)
# silicon
si_tag = feature_data('Si')
si_tag.wavelength = cuts[0]
si_tag.flux = flux_pro[si_tag.wavelength[0]:si_tag.wavelength[1]]
#helium
he_tag = feature_data('He')
he_tag.wavelength = cuts[1]
he_tag.flux = flux_pro[he_tag.wavelength[0]:he_tag.wavelength[1]]
#calcium
ca_tag = feature_data('Ca')
ca_tag.wavelength = cuts[2]
ca_tag.flux = flux_pro[ca_tag.wavelength[0]:ca_tag.wavelength[1]]
#Helium double peak
dp_tag = feature_data('He double peak')
dp_tag.wavelength = cuts[3]
dp_tag.flux = flux_pro[dp_tag.wavelength[0]:dp_tag.wavelength[1]]
#Iron
fe_tag = feature_data('Fe')
fe_tag.wavelength = cuts[4]
fe_tag.flux = flux_pro[fe_tag.wavelength[0]:fe_tag.wavelength[1]]
#Sulphur
s_tag = feature_data('S')
s_tag.wavelength = cuts[5]
s_tag.flux = flux_pro[s_tag.wavelength[0]:s_tag.wavelength[1]]
# + [markdown] id="buC5DrRJnuj4"
# # Tagging
#
# With spectra pre-processed and the necessary cuts made, we can now get the tag probabilities of the spectra and add them to an array ready to be given to the trained classifier.
# + id="-ilC7PlVoKtO"
from tagging import log_reg_two
final = np.zeros([1,10])
#Get Hydrogen tag probabilities
H_result = log_reg_two(flux_pro, beta[0])
final[0][0] = H_result
#Get Silicon tag probabilities
Si_result = log_reg_two(si_tag.flux, beta[1])
final[0][1] = Si_result
#Get Helium emission tag probabilities
He_emi_result = log_reg_two(he_tag.flux, beta[2])
final[0][2] = He_emi_result
#Get Helium p cygni tag probabilities
He_cyg_result = log_reg_two(he_tag.flux, beta[3])
final[0][3] = He_cyg_result
#Get Helium absorption tag probabilities
He_abs_result = log_reg_two(he_tag.flux, beta[4])
final[0][4] = He_abs_result
#Get Hydrogen alpha tag probabilities
H_alp_result = log_reg_two(flux_pro, beta[5])
final[0][5] = H_alp_result
#Get Calcium tag probabilities
Ca_result = log_reg_two(ca_tag.flux, beta[6])
final[0][6] = Ca_result
#Get Helium double peak tag probabilities
dp_result = log_reg_two(dp_tag.flux, beta[7])
final[0][7] = dp_result
#Get Fe tag probabilities
fe_result = log_reg_two(fe_tag.flux, beta[8])
final[0][8] = fe_result
#Get S tag probabilities
s_result = log_reg_two(s_tag.flux, beta[9])
final[0][9] = s_result
# + [markdown] id="5I3z-TRIlGUs"
# # Tag Probabilities
#
# One of the key features of STag is that all of the tags have probabilties, which can be accessed on demand.
# -
tag_names = ['H ','Si ','He emi ','He P-cyg','He abs ','H-alpha ','Ca ','He 6678','Fe ','S ']
for i in range(0,len(tag_names)):
print("{0:s} {1:5.3f}".format(tag_names[i],final[0][i]))
# + [markdown] id="n0dH9swhqlFe"
# # Classifying
#
# We can now make our predictions for the class of the supernova by using the trained model. Since we are using softmax, we use 'np.argmax' to select the class with the highest probability, though one can see the probabilities of all the classes by printing 'class_prob'.
#
# The predicted class is given a number, which corresponds to one of the 5 possible classes:
#
# 0 = Type Ia
#
# 1 = Type II
#
# 2 = Type Ib
#
# 3 = Type Ic
#
# 4 = Type IIb
#
# + id="v5vONcaOqjuY"
import keras
v = keras.__version__
from packaging import version
if version.parse(v) < version.parse('2.5.0'):
print("You may need to update Keras")
#Load in the trained model
model = keras.models.load_model('%s/Classifier Model V2.h5' % path)
#Make classification prediction
class_prob = model.predict(final)
preds = np.argmax(class_prob, axis=-1)
print("SN %s (with redshift %.3f) predicted class is %d with a %.3f probability " % (name,z,preds,class_prob[0][preds]))
# + [markdown] id="Vgy6qy7Zm461"
# # Closing Remarks
#
# One can use STag by following the steps outlined in this notebook, and with slight modifications one can adapt this code to run on multiple spectra rather than one at a time.
#
# Note that the classifying model used has only been trained on the 10 tags shown in this notebook, if one wishes to add additional tags then the model will need to be trained again. A more detailed description of how the tags have been made and how the model was built can be found in our paper: https://arxiv.org/abs/2108.10497
| .ipynb_checkpoints/STag_Demonstration-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
##from the vscode file... data_fix_season_cut_down ...
import pandas as pd
import numpy as np
##import all the files
##file paths
Kaggle_path = "/Users/joejohns/data_bootcamp/Final_Project_NHL_prediction/Data/Kaggle_Data_Ellis/"
mp_path = "/Users/joejohns/data_bootcamp/Final_Project_NHL_prediction/Data/Money_Puck_Data/"
betting_path = "/Users/joejohns/data_bootcamp/Final_Project_NHL_prediction/Data/Betting_Data/"
##Kaggle files
df_game = pd.read_csv(Kaggle_path+'game.csv')
df_game_team_stats = pd.read_csv(Kaggle_path+'game_teams_stats.csv')
df_game_skater_stats = pd.read_csv(Kaggle_path+'game_skater_stats.csv')
df_game_goalie_stats = pd.read_csv(Kaggle_path+'game_goalie_stats.csv')
##more subtle Kaggle features:
df_game_scratches = pd.read_csv(Kaggle_path+'game_scratches.csv')
df_game_officials = pd.read_csv(Kaggle_path+'game_officials.csv')
df_team_info = pd.read_csv(Kaggle_path+'team_info.csv')
## grab all the moneypuck data
df_mp_teams = pd.read_csv(mp_path+'all_teams.csv')
## grab all betting data
df1 = pd.read_excel(io = betting_path+'nhl odds 2007-08.xlsx')
df2 = pd.read_excel(io = betting_path+'nhl odds 2008-09.xlsx')
df3 = pd.read_excel(io = betting_path+'nhl odds 2009-10.xlsx')
df4 = pd.read_excel(io = betting_path+'nhl odds 2010-11.xlsx')
df5 = pd.read_excel(io = betting_path+'nhl odds 2011-12.xlsx')
df6 = pd.read_excel(io = betting_path+'nhl odds 2012-13.xlsx')
df7 = pd.read_excel(io = betting_path+'nhl odds 2013-14.xlsx')
df8 = pd.read_excel(io = betting_path+'nhl odds 2014-15.xlsx')
df9 = pd.read_excel(io = betting_path+'nhl odds 2015-16.xlsx')
df10 = pd.read_excel(io = betting_path+'nhl odds 2016-17.xlsx')
df11 = pd.read_excel(io = betting_path+'nhl odds 2017-18.xlsx')
df12 = pd.read_excel(io = betting_path+'nhl odds 2018-19.xlsx')
df13 = pd.read_excel(io = betting_path+'nhl odds 2019-20.xlsx')
df1['season'] = 20072008
df2['season'] = 20082009
df3['season'] = 20092010
df4['season'] = 20102011
df5['season'] = 20112012
df6['season'] = 20122013
df7['season'] = 20132014
df8['season'] = 20142015
df9['season'] = 20152016
df10['season'] = 20162017
df11['season'] = 20172018
df12['season'] = 20182019
df13['season'] = 20192020
df_betting = pd.concat([df1, df2, df3, df4, df5, df6, df7, df8, df9, df10, df11, df12, df13])
##### restrict data sets
## restrict data sets
df_betting = df_betting.loc[:, ['Date', 'season','VH', 'Team', 'Open']].copy()
df_mp_teams.rename(columns={"teamId": "team_id"}, inplace = True)
df_mp_teams_all = df_mp_teams.loc[df_mp_teams['situation'] == 'all', :].copy()
##### restrict data sets
df_betting = df_betting.loc[:, ['Date', 'season','VH', 'Team', 'Open']].copy()
df_mp_teams_all = df_mp_teams.loc[df_mp_teams['situation'] == 'all', :].copy()
##drop duplicates and one column had some NaN;
##note there are more nan values in df_game_skaters/team/goalies but I think df_mp gets those.
df_game.drop_duplicates(inplace = True)
df_game.drop(columns = ['home_rink_side_start'], inplace = True)
## fix seasons in df_mp (other 2 already have 20082009 format)
def fix_mp_season(n):
return int(str(n)+str(n+1))
#test
#fix_mp_season(2010)
df_mp_teams['season'] = df_mp_teams['season'].map(fix_mp_season)
df_mp_teams_all['season'] = df_mp_teams_all['season'].map(fix_mp_season)
##restrict seasons; 20082009 to 20192020 is the range common to all 3 df's
seasons = []
for n in range(2008,2020):
seasons.append(int(str(n)+str(n+1)))
#check seasons look ok
print(seasons)
#restrict seasons:
##note: In notebook I checked that value counts of the seasons did not change after this
#restriction
df_betting = df_betting.loc[df_game['season'].isin(seasons), :].copy()
df_game = df_game.loc[df_game['season'].isin(seasons), :].copy()
df_mp_teams = df_mp_teams.loc[df_mp_teams['season'].isin(seasons), :].copy()
df_mp_teams_all = df_mp_teams_all.loc[df_mp_teams_all['season'].isin(seasons), :].copy()
##here is a count of how many games in each df ... approx the same ... so looks likely there
##should be close to full overlap in game_id's
## the index is no longer consecutive so we reset:
df_betting.reset_index(drop = True, inplace = True)
df_game.reset_index(drop = True, inplace = True)
df_mp_teams.reset_index(drop = True, inplace = True)
df_mp_teams_all.reset_index(drop = True, inplace = True)
for seas in seasons:
print(seas, len(df_mp_teams_all.loc[df_mp_teams['season']==seas])/2, len(df_game.loc[df_game['season']==seas]), len(df_betting.loc[df_betting['season']==seas])/2)
# -
df_game['date_time_GMT'][0]
# +
## check that df_mp and df_game have similar games ...
# +
## next investigate game overlaps
len(set(df_mp_teams_all['gameId']).symmetric_difference(set(df_game['game_id'])))
# -
len(set(df_mp_teams_all['gameId']).intersection(set(df_game['game_id'])))
# +
## restrict later when we do joins ...
##ok we want to assign game_id to df_mp ... this is the key step, then we can merge all dfs on game_id
##to get game_id, we need to know team, date ... so we need to fix those two things.
## start with date for df_game and df_mp ... that is easier
# +
##fix date for df_game ... we will use df_mp format ... 20080904 ymd
# +
#the function appends 0 in front of any day or month with one digit to follow mp format
def to_string(n):
s = str(n)
if len(s) ==1:
return "0"+s
else:
return s
#takes in date_time date and returns yyyymmdd eg 20080904
def game_add_date(date_time):
d = date_time.day
m = date_time.month
y = date_time.year
dd = to_string(d)
mm = to_string(m)
yyyy = str(y)
return(int(yyyy+mm+dd))
## do mp style date for df_games
df_game['date_time_GMT'] = pd.to_datetime(df_game['date_time_GMT'])
test = df_game['date_time_GMT'][105]
test
game_add_date(test)
df_game['mp_date'] = df_game['date_time_GMT'].apply(game_add_date)
# -
df_game.loc[15000:15100, ['mp_date', 'date_time_GMT']]
print(len(set(df_game['mp_date'])), len(set(df_game['date_time_GMT'])))
# +
##ok, dates look correct for df_game ... now for df_betting
# -
df_betting.head(20)
#restrict seasons: #betting is done already
def bet_mp_date(date, season):
d = str(date)
s = str(season)
if len(d) == 3:
d = "0"+d
if (900 < date):
y= s[:4]
else:
y = s[4:]
return int(y+d)
bet_mp_date(826, 20112012)
df_betting['mp_date'] = np.vectorize(bet_mp_date)(df_betting['Date'], df_betting['season'])
df_betting.shape
df_betting.loc[20000:20020, ['Date', 'season', 'mp_date' ]]
| NHL_data_shape_notebooks/Approach_2_cumul_1seas_Pischedda_data_process/Fixing date, season, team names/Tools for fixing dates, team names, seasons/Fix_dates.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="O6qb81h2NYHb"
# <h2 style = "color : Brown"> Case Study - Cricket Tournament </h2>
#
# A panel wants to select players for an upcoming league match based on their fitness. Players from all significant cricket clubs have participated in a practice match, and their data is collected. Let us now explore NumPy features using the player's data.</font>
#
# <h4 style = "color : Sky blue"> Example - 1</h4>
#
# #### Heights of the players is stored as a regular Python list: height_in. The height is expressed in inches. Can you make a numpy array out of it ?
# + colab={} colab_type="code" id="syM7L-HrNYHe"
# Define list
height_in = [74, 74, 72, 72, 73, 69, 69, 71, 76, 71, 73, 73, 74, 74, 69, 70, 73, 75, 78, 79, 76, 74, 76, 72, 71, 75, 77, 74, 73, 74, 78, 73, 75, 73, 75, 75, 74, 69, 71, 74, 73, 73, 76, 74, 74, 70, 72, 77, 74, 70, 73, 75, 76, 76, 78, 74, 74, 76, 77, 81, 78, 75, 77, 75, 76, 74, 72, 72, 75, 73, 73, 73, 70, 70, 70, 76, 68, 71, 72, 75, 75, 75, 75, 68, 74, 78, 71, 73, 76, 74, 74, 79, 75, 73, 76, 74, 74, 73, 72, 74, 73, 74, 72, 73, 69, 72, 73, 75, 75, 73, 72, 72, 76, 74, 72, 77, 74, 77, 75, 76, 80, 74, 74, 75, 78, 73, 73, 74, 75, 76, 71, 73, 74, 76, 76, 74, 73, 74, 70, 72, 73, 73, 73, 73, 71, 74, 74, 72, 74, 71, 74, 73, 75, 75, 79, 73, 75, 76, 74, 76, 78, 74, 76, 72, 74, 76, 74, 75, 78, 75, 72, 74, 72, 74, 70, 71, 70, 75, 71, 71, 73, 72, 71, 73, 72, 75, 74, 74, 75, 73, 77, 73, 76, 75, 74, 76, 75, 73, 71, 76, 75, 72, 71, 77, 73, 74, 71, 72, 74, 75, 73, 72, 75, 75, 74, 72, 74, 71, 70, 74, 77, 77, 75, 75, 78, 75, 76, 73, 75, 75, 79, 77, 76, 71, 75, 74, 69, 71, 76, 72, 72, 70, 72, 73, 71, 72, 71, 73, 72, 73, 74, 74, 72, 75, 74, 74, 77, 75, 73, 72, 71, 74, 77, 75, 75, 75, 78, 78, 74, 76, 78, 76, 70, 72, 80, 74, 74, 71, 70, 72, 71, 74, 71, 72, 71, 74, 69, 76, 75, 75, 76, 73, 76, 73, 77, 73, 72, 72, 77, 77, 71, 74, 74, 73, 78, 75, 73, 70, 74, 72, 73, 73, 75, 75, 74, 76, 73, 74, 75, 75, 72, 73, 73, 72, 74, 78, 76, 73, 74, 75, 70, 75, 71, 72, 78, 75, 73, 73, 71, 75, 77, 72, 69, 73, 74, 72, 70, 75, 70, 72, 72, 74, 73, 74, 76, 75, 80, 72, 75, 73, 74, 74, 73, 75, 75, 71, 73, 75, 74, 74, 72, 74, 74, 74, 73, 76, 75, 72, 73, 73, 73, 72, 72, 72, 72, 71, 75, 75, 74, 73, 75, 79, 74, 76, 73, 74, 74, 72, 74, 74, 75, 78, 74, 74, 74, 77, 70, 73, 74, 73, 71, 75, 71, 72, 77, 74, 70, 77, 73, 72, 76, 71, 76, 78, 75, 73, 78, 74, 79, 75, 76, 72, 75, 75, 70, 72, 70, 74, 71, 76, 73, 76, 71, 69, 72, 72, 69, 73, 69, 73, 74, 74, 72, 71, 72, 72, 76, 76, 76, 74, 76, 75, 71, 72, 71, 73, 75, 76, 75, 71, 75, 74, 72, 73, 73, 73, 73, 76, 72, 76, 73, 73, 73, 75, 75, 77, 73, 72, 75, 70, 74, 72, 80, 71, 71, 74, 74, 73, 75, 76, 73, 77, 72, 73, 77, 76, 71, 75, 73, 74, 77, 71, 72, 73, 69, 73, 70, 74, 76, 73, 73, 75, 73, 79, 74, 73, 74, 77, 75, 74, 73, 77, 73, 77, 74, 74, 73, 77, 74, 77, 75, 77, 75, 71, 74, 70, 79, 72, 72, 70, 74, 74, 72, 73, 72, 74, 74, 76, 82, 74, 74, 70, 73, 73, 74, 77, 72, 76, 73, 73, 72, 74, 74, 71, 72, 75, 74, 74, 77, 70, 71, 73, 76, 71, 75, 74, 72, 76, 79, 76, 73, 76, 78, 75, 76, 72, 72, 73, 73, 75, 71, 76, 70, 75, 74, 75, 73, 71, 71, 72, 73, 73, 72, 69, 73, 78, 71, 73, 75, 76, 70, 74, 77, 75, 79, 72, 77, 73, 75, 75, 75, 73, 73, 76, 77, 75, 70, 71, 71, 75, 74, 69, 70, 75, 72, 75, 73, 72, 72, 72, 76, 75, 74, 69, 73, 72, 72, 75, 77, 76, 80, 77, 76, 79, 71, 75, 73, 76, 77, 73, 76, 70, 75, 73, 75, 70, 69, 71, 72, 72, 73, 70, 70, 73, 76, 75, 72, 73, 79, 71, 72, 74, 74, 74, 72, 76, 76, 72, 72, 71, 72, 72, 70, 77, 74, 72, 76, 71, 76, 71, 73, 70, 73, 73, 72, 71, 71, 71, 72, 72, 74, 74, 74, 71, 72, 75, 72, 71, 72, 72, 72, 72, 74, 74, 77, 75, 73, 75, 73, 76, 72, 77, 75, 72, 71, 71, 75, 72, 73, 73, 71, 70, 75, 71, 76, 73, 68, 71, 72, 74, 77, 72, 76, 78, 81, 72, 73, 76, 72, 72, 74, 76, 73, 76, 75, 70, 71, 74, 72, 73, 76, 76, 73, 71, 68, 71, 71, 74, 77, 69, 72, 76, 75, 76, 75, 76, 72, 74, 76, 74, 72, 75, 78, 77, 70, 72, 79, 74, 71, 68, 77, 75, 71, 72, 70, 72, 72, 73, 72, 74, 72, 72, 75, 72, 73, 74, 72, 78, 75, 72, 74, 75, 75, 76, 74, 74, 73, 74, 71, 74, 75, 76, 74, 76, 76, 73, 75, 75, 74, 68, 72, 75, 71, 70, 72, 73, 72, 75, 74, 70, 76, 71, 82, 72, 73, 74, 71, 75, 77, 72, 74, 72, 73, 78, 77, 73, 73, 73, 73, 73, 76, 75, 70, 73, 72, 73, 75, 74, 73, 73, 76, 73, 75, 70, 77, 72, 77, 74, 75, 75, 75, 75, 72, 74, 71, 76, 71, 75, 76, 83, 75, 74, 76, 72, 72, 75, 75, 72, 77, 73, 72, 70, 74, 72, 74, 72, 71, 70, 71, 76, 74, 76, 74, 74, 74, 75, 75, 71, 71, 74, 77, 71, 74, 75, 77, 76, 74, 76, 72, 71, 72, 75, 73, 68, 72, 69, 73, 73, 75, 70, 70, 74, 75, 74, 74, 73, 74, 75, 77, 73, 74, 76, 74, 75, 73, 76, 78, 75, 73, 77, 74, 72, 74, 72, 71, 73, 75, 73, 67, 67, 76, 74, 73, 70, 75, 70, 72, 77, 79, 78, 74, 75, 75, 78, 76, 75, 69, 75, 72, 75, 73, 74, 75, 75, 73]
# + colab={} colab_type="code" id="kuS3VSD-NYHr"
import numpy as np
heights = np.array(height_in)
# + colab={} colab_type="code" id="K2l7qL67NYH2" outputId="a69b3b52-e221-4970-82f8-43ba3074d2ea"
heights
# + colab={} colab_type="code" id="-__JqvMaNYIC" outputId="2d122803-98a9-4e9c-f401-1aca4f0df8c8"
type(heights)
# + [markdown] colab_type="text" id="1ANNW-OJNYIM"
# <h4 style = "color : Sky blue"> Example - 2</h4>
#
# #### Count the number of pariticipants
# + colab={} colab_type="code" id="BPtkkx_kNYIO" outputId="0501f98b-3038-4764-99e2-e3e7b01145d9"
len(heights)
# + colab={} colab_type="code" id="utC2FQZnNYIb" outputId="01d007f7-71d4-48d4-ecf6-58c7186ba93c"
heights.size
# + colab={} colab_type="code" id="CHlC4G1_NYIk" outputId="bfb7e252-0792-41c6-c85d-56a748136ca3"
heights.shape
# + [markdown] colab_type="text" id="H_ykFKOrNYIv"
# <h4 style = "color : Sky blue"> Example - 3</h4>
#
# #### Convert the heights from inches to meters
# + colab={} colab_type="code" id="tFh-kO3jNYIx" outputId="8553f4db-3f0e-4a21-ff9f-0c9a61325975"
heights_m = heights * 0.0254
heights_m
# + [markdown] colab_type="text" id="P5BbAFY1NYI6"
# <h4 style = "color : Sky blue"> Example - 4</h4>
#
# #### A list of weights (in lbs) of the players is provided. Convert it to kg and calculate BMI
# + colab={} colab_type="code" id="WB52h6LbNYI7"
weights_lb = [180, 215, 210, 210, 188, 176, 209, 200, 231, 180, 188, 180, 185, 160, 180, 185, 189, 185, 219, 230, 205, 230, 195, 180, 192, 225, 203, 195, 182, 188, 200, 180, 200, 200, 245, 240, 215, 185, 175, 199, 200, 215, 200, 205, 206, 186, 188, 220, 210, 195, 200, 200, 212, 224, 210, 205, 220, 195, 200, 260, 228, 270, 200, 210, 190, 220, 180, 205, 210, 220, 211, 200, 180, 190, 170, 230, 155, 185, 185, 200, 225, 225, 220, 160, 205, 235, 250, 210, 190, 160, 200, 205, 222, 195, 205, 220, 220, 170, 185, 195, 220, 230, 180, 220, 180, 180, 170, 210, 215, 200, 213, 180, 192, 235, 185, 235, 210, 222, 210, 230, 220, 180, 190, 200, 210, 194, 180, 190, 240, 200, 198, 200, 195, 210, 220, 190, 210, 225, 180, 185, 170, 185, 185, 180, 178, 175, 200, 204, 211, 190, 210, 190, 190, 185, 290, 175, 185, 200, 220, 170, 220, 190, 220, 205, 200, 250, 225, 215, 210, 215, 195, 200, 194, 220, 180, 180, 170, 195, 180, 170, 206, 205, 200, 225, 201, 225, 233, 180, 225, 180, 220, 180, 237, 215, 190, 235, 190, 180, 165, 195, 200, 190, 190, 185, 185, 205, 190, 205, 206, 220, 208, 170, 195, 210, 190, 211, 230, 170, 185, 185, 241, 225, 210, 175, 230, 200, 215, 198, 226, 278, 215, 230, 240, 184, 219, 170, 218, 190, 225, 220, 176, 190, 197, 204, 167, 180, 195, 220, 215, 185, 190, 205, 205, 200, 210, 215, 200, 205, 211, 190, 208, 200, 210, 232, 230, 210, 220, 210, 202, 212, 225, 170, 190, 200, 237, 220, 170, 193, 190, 150, 220, 200, 190, 185, 185, 200, 172, 220, 225, 190, 195, 219, 190, 197, 200, 195, 210, 177, 220, 235, 180, 195, 195, 190, 230, 190, 200, 190, 190, 200, 200, 184, 200, 180, 219, 187, 200, 220, 205, 190, 170, 160, 215, 175, 205, 200, 214, 200, 190, 180, 205, 220, 190, 215, 235, 191, 200, 181, 200, 210, 240, 185, 165, 190, 185, 175, 155, 210, 170, 175, 220, 210, 205, 200, 205, 195, 240, 150, 200, 215, 202, 200, 190, 205, 190, 160, 215, 185, 200, 190, 210, 185, 220, 190, 202, 205, 220, 175, 160, 190, 200, 229, 206, 220, 180, 195, 175, 188, 230, 190, 200, 190, 219, 235, 180, 180, 180, 200, 234, 185, 220, 223, 200, 210, 200, 210, 190, 177, 227, 180, 195, 199, 175, 185, 240, 210, 180, 194, 225, 180, 205, 193, 230, 230, 220, 200, 249, 190, 208, 245, 250, 160, 192, 220, 170, 197, 155, 190, 200, 220, 210, 228, 190, 160, 184, 180, 180, 200, 176, 160, 222, 211, 195, 200, 175, 206, 240, 185, 260, 185, 221, 205, 200, 170, 201, 205, 185, 205, 245, 220, 210, 220, 185, 175, 170, 180, 200, 210, 175, 220, 206, 180, 210, 195, 200, 200, 164, 180, 220, 195, 205, 170, 240, 210, 195, 200, 205, 192, 190, 170, 240, 200, 205, 175, 250, 220, 224, 210, 195, 180, 245, 175, 180, 215, 175, 180, 195, 230, 230, 205, 215, 195, 180, 205, 180, 190, 180, 190, 190, 220, 210, 255, 190, 230, 200, 205, 210, 225, 215, 220, 205, 200, 220, 197, 225, 187, 245, 185, 185, 175, 200, 180, 188, 225, 200, 210, 245, 213, 231, 165, 228, 210, 250, 191, 190, 200, 215, 254, 232, 180, 215, 220, 180, 200, 170, 195, 210, 200, 220, 165, 180, 200, 200, 170, 224, 220, 180, 198, 240, 239, 185, 210, 220, 200, 195, 220, 230, 170, 220, 230, 165, 205, 192, 210, 205, 200, 210, 185, 195, 202, 205, 195, 180, 200, 185, 240, 185, 220, 205, 205, 180, 201, 190, 208, 240, 180, 230, 195, 215, 190, 195, 215, 215, 220, 220, 230, 195, 190, 195, 209, 204, 170, 185, 205, 175, 210, 190, 180, 180, 160, 235, 200, 210, 180, 190, 197, 203, 205, 170, 200, 250, 200, 220, 200, 190, 170, 190, 220, 215, 206, 215, 185, 235, 188, 230, 195, 168, 190, 160, 200, 200, 189, 180, 190, 200, 220, 187, 240, 190, 180, 185, 210, 220, 219, 190, 193, 175, 180, 215, 210, 200, 190, 185, 220, 170, 195, 205, 195, 210, 190, 190, 180, 220, 190, 186, 185, 190, 180, 190, 170, 210, 240, 220, 180, 210, 210, 195, 160, 180, 205, 200, 185, 245, 190, 210, 200, 200, 222, 215, 240, 170, 220, 156, 190, 202, 221, 200, 190, 210, 190, 200, 165, 190, 185, 230, 208, 209, 175, 180, 200, 205, 200, 250, 210, 230, 244, 202, 240, 200, 215, 177, 210, 170, 215, 217, 198, 200, 220, 170, 200, 230, 231, 183, 192, 167, 190, 180, 180, 215, 160, 205, 223, 175, 170, 190, 240, 175, 230, 223, 196, 167, 195, 190, 250, 190, 190, 190, 170, 160, 150, 225, 220, 209, 210, 176, 260, 195, 190, 184, 180, 195, 195, 219, 225, 212, 202, 185, 200, 209, 200, 195, 228, 210, 190, 212, 190, 218, 220, 190, 235, 210, 200, 188, 210, 235, 188, 215, 216, 220, 180, 185, 200, 210, 220, 185, 231, 210, 195, 200, 205, 200, 190, 250, 185, 180, 170, 180, 208, 235, 215, 244, 220, 185, 230, 190, 200, 180, 190, 196, 180, 230, 224, 160, 178, 205, 185, 210, 180, 190, 200, 257, 190, 220, 165, 205, 200, 208, 185, 215, 170, 235, 210, 170, 180, 170, 190, 150, 230, 203, 260, 246, 186, 210, 198, 210, 215, 180, 200, 245, 200, 192, 192, 200, 192, 205, 190, 186, 170, 197, 219, 200, 220, 207, 225, 207, 212, 225, 170, 190, 210, 230, 210, 200, 238, 234, 222, 200, 190, 170, 220, 223, 210, 215, 196, 175, 175, 189, 205, 210, 180, 180, 197, 220, 228, 190, 204, 165, 216, 220, 208, 210, 215, 195, 200, 215, 229, 240, 207, 205, 208, 185, 190, 170, 208, 225, 190, 225, 185, 180, 165, 240, 220, 212, 163, 215, 175, 205, 210, 205, 208, 215, 180, 200, 230, 211, 230, 190, 220, 180, 205, 190, 180, 205, 190, 195]
# + colab={} colab_type="code" id="b_qxn7VfNYJE" outputId="50804a29-64c2-4fbf-b5c7-44209056d23a"
# Converting weights in lbs to kg
weights_kg = np.array(weights_lb) * 0.453592
weights_kg
# + colab={} colab_type="code" id="Hqst4kWvNYJM" outputId="db0ce3e1-4f4f-428f-9471-58cc1e62ef4d"
# Calculate the BMI: bmi
bmi = weights_kg / (heights_m ** 2)
bmi
# + [markdown] colab_type="text" id="1hgOQonCNYJX"
# <h4 style = "color : Sky blue"> Sub-Setting Arrays</h4>
#
# ##### Fetch the first element from the bmi array
# + colab={} colab_type="code" id="bxC_4zPqNYJZ" outputId="7428318b-84be-4e24-b73f-8baf6f9dfc6b"
bmi[0]
# + [markdown] colab_type="text" id="IJw_fIF_NYJh"
# ##### Fetch the last element from the bmi array
# + colab={} colab_type="code" id="pMjBir0PNYJj" outputId="242da47a-5af9-4ac0-a875-dcf33ca53cb8"
bmi[-1]
# + [markdown] colab_type="text" id="JLHhFzmONYJr"
# ##### Fetch the first 5 elements from the bmi array
# + colab={} colab_type="code" id="qinJdYrONYJt" outputId="b7feaa6a-c71a-49e9-87e7-c72e8767b66c"
bmi[0:5]
# + [markdown] colab_type="text" id="AfBVk-mXNYJ5"
# ##### Fetch the last 5 elements from the bmi array
# + colab={} colab_type="code" id="ZTUtXBuZNYJ7" outputId="9d9120a0-6caf-4da7-a73b-0b88346ebf38"
bmi[-5:]
# + [markdown] colab_type="text" id="UwWgOMiLNYKE"
# <h4 style = "color : Sky blue"> Conditional Sub-Setting Arrays</h4>
#
# ##### Count the number of pariticipants who are underweight i.e. bmi < 21
#
# + colab={} colab_type="code" id="JT-qmbkPNYKG" outputId="7347c038-039e-4907-c5bf-61fade4b2fd6"
bmi < 21
# + colab={} colab_type="code" id="A9NNHZ8wNYKO" outputId="336085a3-7911-4904-99f3-afd0d23e5bac"
bmi [ bmi<21]
# + colab={} colab_type="code" id="sUejHbbONYKX" outputId="8f9d6cc9-445e-4737-e729-2cdd0e81a830"
underweight_players = bmi [ bmi<21]
underweight_players
# + colab={} colab_type="code" id="w_eRWB_YNYKf" outputId="e65c1510-27a3-4ea6-da73-93b1c7db62ec"
underweight_players.size
# + [markdown] colab_type="text" id="UoLIRApcNYKn"
# <h4 style = "color : Sky blue"> NumPy Functions</h4>
# + [markdown] colab_type="text" id="2D3fJra3NYKo"
# ##### Find the largest BMI value
# + colab={} colab_type="code" id="bbRRch9INYKq" outputId="5171b37a-7b12-4b8f-e469-68dd54267909"
max(bmi)
# + colab={} colab_type="code" id="fumC4BybNYKy" outputId="26d6ce11-fbe5-4dd3-f4b7-1c4140407be7"
bmi.max()
# + [markdown] colab_type="text" id="BGQBQcs9NYK6"
# ##### Find lowest BMI value
# + colab={} colab_type="code" id="ZHkzhfynNYK8" outputId="44eecd88-a98c-4342-c114-a279e6af3a89"
bmi.min()
# + [markdown] colab_type="text" id="S_wKFbq0NYLG"
# ##### Find average BMI value
# + colab={} colab_type="code" id="xC4IdjdHNYLJ" outputId="99485c01-7c47-4b7e-e1e6-45074a4e5938"
bmi.mean()
| Course_1-PreLaunch_Preparatory_Content/Module_2-Python_Libraries/1-NumPy/Session2_Operations_over_1D_Arrays.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # K-Means and Hierarchical clustering
# ### Libraries
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib as mpl
import matplotlib.pyplot as plt
import scipy.cluster.hierarchy as sch
from sklearn.cluster import KMeans
# -
# ## Data
#
# Data set: **`Animals with Attributes`** contains information about 50 animals.
# For each, it has 85 real-valued features that capture various properties of the animal: where it lives, what it eats, and so on.
# You can download the data set from: http://attributes.kyb.tuebingen.mpg.de
# !find ../../_data | grep -1 classes.txt
# !find ../../_data | grep -1 predicate-matrix-continuous.txt
# Load in the data set. The file `'classes.txt'` contains the names of the 50 animals. The file `'predicate-matrix-continuous.txt'` contains the data itself: 85 attributes per animal.
# !head -2 ../../_data/Animals_with_Attributes/predicate-matrix-continuous.txt
# ### Load txt matrix
X = np.loadtxt('../../_data/Animals_with_Attributes/predicate-matrix-continuous.txt')
X.shape
# !head '../../_data/Animals_with_Attributes/classes.txt'
df = pd.read_csv('../../_data/Animals_with_Attributes/classes.txt', header=None, index_col=0, delimiter='\t')
df_classes = df[1]
df_classes.sample(5)
# ## K-means clustering
# We now run Lloyd's algorithm to obtain a flat clustering of the data. In the code below, we ask for k=15 clusters, but you should experiment with other choices.
#
# We ask for random initialization, which means that different runs of the algorithm will potentially have different outcomes. It is worth running the algorithm several times to see how the results change.
# ### Train K-means model
k = 8
kmeans = KMeans(n_clusters=k, init='random').fit(X)
# ### Cluster distribution and labels
from collections import Counter
kmeans.labels_
Counter(kmeans.labels_)
# ### Clusters by dictionary - cluster:[values,...]
clusters = {cluster:[] for cluster in kmeans.labels_}
_ = [clusters[cluster].append(label) for cluster, label in zip(kmeans.labels_, df_classes)]
clusters
# ### Sum of distance to closest cluster center
# +
plt.figure(figsize=(6,6))
mpl.rc('axes.spines', left=True, top=False, right=False, bottom=True) # hide axis/spines
mpl.rc('xtick', color='k') # hide xticks
dist_cluster = []
for k in range(1, 18):
kmeans = KMeans(n_clusters=k, init='random').fit(X)
dist_cluster.append(kmeans.inertia_)
_ = plt.plot(range(1, 18), dist_cluster, '-o');
# -
km6 = KMeans(n_clusters=6, init='random').fit(X)
km14 = KMeans(n_clusters=14, init='random').fit(X)
# +
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(18, 6))
ax1.scatter(X[:,0], X[:,1], s=40, c=km6.labels_, cmap=plt.cm.prism)
ax1.set_title('K-Means Clustering Results with K=6')
ax1.scatter(km6.cluster_centers_[:, 0], km6.cluster_centers_[:, 1], marker='+', s=100, c='k', linewidth=2)
ax2.scatter(X[:, 0], X[:, 1], s=40, c=km14.labels_, cmap=plt.cm.prism)
ax2.set_title('K-Means Clustering Results with K=14')
ax2.scatter(km14.cluster_centers_[:, 0], km14.cluster_centers_[:, 1], marker='+', s=100, c='k', linewidth=2);
# -
# ### Kmeans using PCA
from sklearn.preprocessing import scale
from sklearn.decomposition import PCA
pca = PCA(n_components=2)
pca.fit(scale(X))
pca.fit_transform(scale(X));
df = pd.DataFrame(pca.fit_transform(X) , columns=['PC1', 'PC2'])
df.sample(5)
# ### Vector Loadings - Eigen Vectors
pca.components_[0]
pca_loadings = pd.DataFrame(pca.components_.T, columns=['V1', 'V2'])
pca_loadings.head()
# +
plt.figure(figsize=(6,6))
mpl.rc('axes.spines', left=True, top=False, right=False, bottom=True) # hide axis/spines
mpl.rc('xtick', color='k') # hide xticks
dist_cluster = []
for k in range(1, 18):
kmeans = KMeans(n_clusters=k, init='random').fit(df)
dist_cluster.append(kmeans.inertia_)
_ = plt.plot(range(1, 18), dist_cluster, '-o');
# -
km4 = KMeans(n_clusters=4, init='random').fit(df)
km8 = KMeans(n_clusters=8, init='random').fit(df)
# +
import seaborn as sns
# https://seaborn.pydata.org/tutorial/color_palettes.html
colors = ["#67E568","#257F27","#08420D","#FFF000","#FFB62B","#E56124","#E53E30","#7F2353","#F911FF","#9F8CA6"]
cpal = sns.color_palette(colors)
sns.palplot(cpal, 1)
cmap_mpl = mpl.colors.ListedColormap(cpal.as_hex()) # discrete
# +
fig, (ax1, ax2) = plt.subplots(1,2, figsize=(18, 6))
ax1.scatter(df['PC1'], df['PC2'], s=40, c=km4.labels_, cmap=cmap_mpl)
ax1.set_title('K-Means Clustering Results with K=4')
ax1.scatter(km4.cluster_centers_[:, 0], km4.cluster_centers_[:, 1], marker='+', s=100, c='k', linewidth=2)
ax2.scatter(df['PC1'], df['PC2'], s=40, c=km8.labels_, cmap=cmap_mpl)
ax2.set_title('K-Means Clustering Results with K=8')
ax2.scatter(km8.cluster_centers_[:, 0], km8.cluster_centers_[:, 1], marker='+', s=100, c='k', linewidth=2);
# -
# ### Compare clusters with and without PCA
clusters_pca = {cluster:[] for cluster in km8.labels_}
_ = [clusters_pca[cluster].append(label) for cluster, label in zip(km8.labels_, df_classes)]
clusters_pca
clusters
# +
from matplotlib.pyplot import rcParams
mpl.rc('axes.spines', left=False, top=False, right=False, bottom=False)
mpl.rc('xtick', color='w') # hide xticks
mpl.rc('ytick', color='w') # hide xticks
# +
fig , ax1 = plt.subplots(figsize=(12, 12))
colors = ["#67E568","#257F27","#08420D","#FFF000","#FFB62B","#E56124","#E53E30","#7F2353","#F911FF","#9F8CA6"]
_ = ax1.set_xlim(-300, 200)
_ = ax1.set_ylim(-150, 150)
# Plot Principal Components 1 and 2
for i in df.index:
label = km8.labels_[i]
_ = ax1.annotate(df_classes.iloc[i], (df.PC1.loc[i], df.PC2.loc[i]), ha='center',
color=colors[label], size=14, alpha=.9)
ax1.set_xlabel('Principal Component 1', size=14)
ax1.set_ylabel('Principal Component 2', size=14)
# Plot Eigen Vectors
mp = 1000
ax1.arrow(0, 0, pca_loadings.V1[0]*mp, pca_loadings.V2[0]*mp, color='blue')
ax1.arrow(0, 0, pca_loadings.V1[1]*mp, pca_loadings.V2[1]*mp, color='blue');
# -
# ## Hierarchical clustering
# We use the built-in hierarchical clustering module of `scipy` to apply **Ward's method** to our data.
#
# Lloyd's algorithm potentially returns a different solution each time it is run.
z = sch.linkage(X, method='ward')
# ### Show dendogram
# ### Set defaults dendogram
# +
from matplotlib.pyplot import rcParams
# plt.rcParams.find_all
mpl.rc('figure', figsize=[10., 12.])
mpl.rc('axes.spines', left=False, top=False, right=False, bottom=False) # hide axis/spines
mpl.rc('xtick', color='w') # hide xticks
# +
_ = plt.figure(figsize=(10, 12))
# Display dendrogram
info = sch.dendrogram(z, orientation='left', labels=df_classes.values, leaf_font_size=12)
leaves_in_reverse = info['ivl']
# -
# ### Caveats and questions regarding clustering
# Here are some things to think about:
#
# **Multiple runs of Lloyd's algorithm**
# Lloyd's algorithm potentially returns a different solution each time it is run.
# Is there any reason to run it more than once?
# For instance, is there a sensible way of combining the information from several runs,
# of interpreting the similarities and differences?
#
# **Sensitivity to the choice of features**
# Both clustering methods are highly sensitive to the choice of features.
# How would you feel if the results changed dramatically when just one or two features were dropped?
#
# **Criteria for success**
# This is clearly an application in which we are hoping that clustering will discover 'natural groups' in the data.
# To what extent do the algorithms succeed at this? Are the clusters mostly reasonable?
# Can we, in general, hope that tha clustering will perfectly capture what we want?
# Under what conditions would we be pleased with the clustering?
| clustering/clustering_kmeans_hierarchical.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Time series in pandas
# ## 1. Creating and using a DatetimeIndex
# The pandas Index is a powerful way to handle time series data, so it is valuable to know how to build one yourself. Pandas provides the pd.to_datetime() function for just this task. For example, if passed the list of strings ['2015-01-01 091234','2015-01-01 091234'] and a format specification variable, such as format='%Y-%m-%d %H%M%S, pandas will parse the string into the proper datetime elements and build the datetime objects.
#
# In this exercise, a list of temperature data and a list of date strings has been pre-loaded for you as temperature_list and date_list respectively. Your job is to use the .to_datetime() method to build a DatetimeIndex out of the list of date strings, and to then use it along with the list of temperature data to build a pandas Series.
# Importing required packages
import pandas as pd
# +
# Prepare a format string: time_format
time_format = '%Y-%m-%d %H:%M'
# Convert date_list into a datetime object: my_datetimes
my_datetimes = pd.to_datetime(date_list, format=time_format)
my_datetimes
# -
# Construct a pandas Series using temperature_list and my_datetimes: time_series
time_series = pd.Series(temperature_list, index=my_datetimes)
time_series.tail()
# Awesome job! Next to DataFrames, Series are another important object that pandas allows us to create, and they're very convenient for time series data
# ## 2. Partial string indexing and slicing
# Pandas time series support "partial string" indexing. What this means is that even when passed only a portion of the datetime, such as the date but not the time, pandas is remarkably good at doing what one would expect. Pandas datetime indexing also supports a wide variety of commonly used datetime string formats, even when mixed.
#
# In this exercise, a time series that contains hourly weather data has been pre-loaded for you. This data was read using the parse_dates=True option in read_csv() with index_col="Dates" so that the Index is indeed a DatetimeIndex.
#
# Extract all data from the 'Temperature' column into the variable ts0. Your job is to use a variety of natural date strings to extract one or more values from ts0.
#
# After you are done, you will have three new variables - ts1, ts2, and ts3. You can slice these further to extract only the first and last entries of each. Try doing this after your submission for more practice.
# Import in weather data
ts0 = pd.read_csv("datasets/weather_data_austin_2010.csv", parse_dates= True, index_col = "Date")
ts0.tail()
# Extract the hour from 9pm to 10pm on '2010-10-11': ts1
ts1 = ts0.loc['2010-10-11 21:00:00':'2010-10-11 22:00:00']
ts1.info()
# Extract '2010-07-04' from ts0: ts2
ts2 = ts0.loc['2010-07-04']
ts2.info()
# Extract data from '2010-12-15' to '2010-12-31': ts3
ts3 = ts0.loc['2010-12-15':'2010-12-31']
ts3.info()
# ## 3. Reindexing the Index
# Reindexing is useful in preparation for adding or otherwise combining two time series data sets. To reindex the data, we provide a new index and ask pandas to try and match the old data to the new index. If data is unavailable for one of the new index dates or times, you must tell pandas how to fill it in. Otherwise, pandas will fill with NaN by default.
#
# In this exercise, load the two time series data sets containing daily data, each indexed by dates. The first, ts1, includes weekends, but the second, ts2, does not. The goal is to combine the two data sets in a sensible way. Your job is to reindex the second data set so that it has weekends as well, and then add it to the first. When you are done, it would be informative to inspect your results.
# Reindex without fill method: ts3
ts3 = ts2.reindex(ts1.index)
ts3
# Reindex with fill method, using forward fill: ts4
ts4 = ts2.reindex(ts1.index, method='ffill')
ts4
# Combine ts1 + ts2: sum12
sum12 = ts1+ts2
sum12.tail()
# Combine ts1 + ts3: sum13
sum13 = ts1+ts3
sum13
# Combine ts1 + ts4: sum14
sum14 = ts1+ts4
sum14
# Wonderful work! Understanding how indexing and reindexing works is a valuable skill.
# ## 4. Resampling and frequency
# Pandas provides methods for resampling time series data. When downsampling or upsampling, the syntax is similar, but the methods called are different. Both use the concept of 'method chaining' - df.method1().method2().method3() - to direct the output from one method call to the input of the next, and so on, as a sequence of operations, one feeding into the next.
#
# For example, if you have hourly data, and just need daily data, pandas will not guess how to throw out the 23 of 24 points. You must specify this in the method. One approach, for instance, could be to take the mean, as in df.resample('D').mean().
#
# In this exercise, load the data set containing hourly temperature data. Your job is to resample the data using a variety of aggregation methods to answer a few questions.
# Copying ts0 to df dataframe
df = ts0.copy()
# Downsample to 6 hour data and aggregate by mean: df1
df1 = df['Temperature'].resample('6h').mean()
df1.tail()
# Downsample to daily data and count the number of data points: df2
df2 = df['Temperature'].resample('D').count()
df2.tail()
# Excellent job! You'll get a lot more practice with resampling in the coming exercises!
# ## 5. Separating and resampling
# With pandas, you can resample in different ways on different subsets of your data. For example, resampling different months of data with different aggregations. In this exercise, the data set containing hourly temperature data from the last exercise has been pre-loaded.
#
# Your job is to resample the data using a variety of aggregation methods. You will be working with the 'Temperature' column.
# Extract temperature data for August: august
august = df['Temperature']['2010-08']
august.tail()
# Downsample to obtain only the daily highest temperatures in August: august_highs
august_highs = august.resample('D').max()
august_highs.tail()
# Extract temperature data for February: february
february = df['Temperature']['2010-02']
february.tail()
# Downsample to obtain the daily lowest temperatures in February: february_lows
february_lows = february.resample('D').min()
february_lows.tail()
# ## 6. Rolling mean and frequency
# In this exercise, some hourly weather data is pre-loaded for you. You will continue to practice resampling, this time using rolling means.
#
# Rolling means (or moving averages) are generally used to smooth out short-term fluctuations in time series data and highlight long-term trends. You can read more about them [here](https://en.wikipedia.org/wiki/Moving_average).
#
# To use the .rolling() method, you must always use method chaining, first calling .rolling() and then chaining an aggregation method after it. For example, with a Series hourly_data, hourly_data.rolling(window=24).mean() would compute new values for each hourly point, based on a 24-hour window stretching out behind each point. The frequency of the output data is the same: it is still hourly. Such an operation is useful for smoothing time series data.
#
# Your job is to resample the data using the combination of .rolling() and .mean().
# Extract data from 2010-Aug-01 to 2010-Aug-15: unsmoothed
unsmoothed = df['Temperature']['2010-Aug-01':'2010-Aug-15']
# Apply a rolling mean with a 24 hour window: smoothed
smoothed = unsmoothed.rolling(window = 24).mean()
# Create a new DataFrame with columns smoothed and unsmoothed: august
august = pd.DataFrame({'smoothed':smoothed, 'unsmoothed':unsmoothed})
august.tail()
# %matplotlib inline
# Plot both smoothed and unsmoothed data using august.plot().
august.plot(figsize=(18, 5));
# ## 7. Resample and roll with it
# As of pandas version 0.18.0, the interface for applying rolling transformations to time series has become more consistent and flexible, and feels somewhat like a groupby (If you do not know what a groupby is, don't worry, you will learn about it in the next course!).
#
# You can now flexibly chain together resampling and rolling operations. In this exercise, the same weather data from the previous exercises has been pre-loaded for you. Your job is to extract one month of data, resample to find the daily high temperatures, and then use a rolling and aggregation operation to smooth the data.
# Extract the August 2010 data: august
august = df['Temperature']['2010-08']
august.tail()
# Resample to daily data, aggregating by max: daily_highs
daily_highs = august.resample('D').max()
daily_highs
# Use a rolling 7-day window with method chaining to smooth the daily high temperatures in August
daily_highs_smoothed = daily_highs.rolling(window = 7).mean()
daily_highs_smoothed
# ## 8. Method chaining and filtering
# We've seen that pandas supports method chaining. This technique can be very powerful when cleaning and filtering data.
#
# In this exercise, load the DataFrame containing flight departure data for a single airline and a single airport for the month of July 2015. Your job is to use .str() filtering and method chaining to generate summary statistics on flight delays each day to Dallas.
flight = pd.read_csv("datasets/austin_airport_departure_data_2015_july.csv", header= 10,
parse_dates= True, index_col= "Date (MM/DD/YYYY)")
flight.info()
flight.columns
# Strip extra whitespace from the column names: df.columns
flight.columns = flight.columns.str.strip()
flight.columns
# Extract data for which the destination airport is Dallas: dallas
dallas = flight['Destination Airport'].str.contains('DAL')
dallas.tail()
# Compute the total number of Dallas departures each day: daily_departures
daily_departures = dallas.resample("D").sum()
daily_departures.tail()
# Generate the summary statistics for daily Dallas departures: stats
stats = daily_departures.describe()
stats
# Great work! You'll return to this dataset later in this chapter.
#
#
# ## 9. Missing values and interpolation
# One common application of interpolation in data analysis is to fill in missing data.
#
# In this exercise, noisy measured data that has some dropped or otherwise missing values has been loaded. The goal is to compare two time series, and then look at summary statistics of the differences. The problem is that one of the data sets is missing data at some of the times.
#
# Your job is to first interpolate to fill in the data for all days. Then, compute the differences between the two data sets, now that they both have full support for all times. Finally, generate the summary statistics that describe the distribution of differences.
# Chopping the dataframe to emulate missing values
ts5 = ts2.iloc[0: 3, :]
# Reindexing based on ts2 to show how the last entries chopped off are now missing values
ts5.reindex(ts2.index).head()
# Reset the index of ts5 to ts2, and then use linear interpolation to fill in the NaNs: ts5_interp
ts5_interp = ts5.reindex(ts2.index).interpolate(how="linear")
ts5_interp.head()
# +
# Import numpy
import numpy as np
# Compute the absolute difference of ts1 and ts5_interp: differences
differences = np.abs(ts2-ts5_interp)
# Generate and print summary statistics of the differences
differences.describe()
# -
# ## 10. Time zones and conversion
# Time zone handling with pandas typically assumes that you are handling the Index of the Series. In this exercise, you will learn how to handle timezones that are associated with datetimes in the column data, and not just the Index.
#
# You will work with the flight departure dataset again, and this time you will select Los Angeles ('LAX') as the destination airport.
#
# Here we will use a mask to ensure that we only compute on data we actually want. To learn more about Boolean masks, click [here](https://docs.scipy.org/doc/numpy/reference/maskedarray.generic.html)!
# +
# Build a Boolean mask to filter for the 'LAX' departure flights: mask
mask = flight['Destination Airport'] == "LAX"
# Use the mask to subset the data and reset the index of the flight dataframe: la
la = flight[mask].reset_index()
la.info()
# -
# Combine two columns of data to create a datetime series: times_tz_none
times_tz_none = pd.to_datetime( la['Date (MM/DD/YYYY)'].astype(str) + ' ' + la['Wheels-off Time'] )
times_tz_none.tail()
# Localize the time to US/Central: times_tz_central
times_tz_central = times_tz_none.dt.tz_localize("US/CENTRAL")
times_tz_central.tail()
# Convert the datetimes from US/Central to US/Pacific
times_tz_pacific = times_tz_central.dt.tz_convert("US/PACIFIC")
times_tz_pacific.tail()
# ## 11. Plotting time series, datetime indexing
# Pandas handles datetimes not only in your data, but also in your plotting.
#
# In this exercise, some time series data has been pre-loaded. However, we have not parsed the date-like columns nor set the index, as we have done for you in the past!
#
# The plot displayed is how pandas renders data with the default integer/positional index. Your job is to convert the 'Date' column from a collection of strings into a collection of datetime objects. Then, you will use this converted 'Date' column as your new index, and re-plot the data, noting the improved datetime awareness. After you are done, you can cycle between the two plots you generated by clicking on the 'Previous Plot' and 'Next Plot' buttons.
#
# Before proceeding, look at the plot shown and observe how pandas handles data with the default integer index.
weather = pd.read_csv("datasets/weather_data_austin_2010.csv")
weather.tail()
df = weather.loc[0:750, ["Temperature", "Date"]]
df.tail()
# Plot the raw data before setting the datetime index
df.plot(figsize=(18, 5));
# +
# Convert the 'Date' column into a collection of datetime objects: df.Date
df.Date = pd.to_datetime(df.Date)
# Set the index to be the converted 'Date' column
df.set_index("Date", inplace=True)
# Re-plot the DataFrame to see that the axis is now datetime aware!
df.plot(figsize=(18, 5));
# -
# ## 12. Plotting date ranges, partial indexing
# Now that you have set the DatetimeIndex in your DataFrame, you have a much more powerful and flexible set of tools to use when plotting your time series data. Of these, one of the most convenient is partial string indexing and slicing. In this exercise, we've pre-loaded a full year of Austin 2010 weather data, with the index set to be the datetime parsed 'Date' column as shown in the previous exercise.
#
# Your job is to use partial string indexing of the dates, in a variety of datetime string formats, to plot all the summer data and just one week of data together.
#
# First, remind yourself how to extract one month of temperature data using 'May 2010' as a key into df.Temperature[], and call head() to inspect the result: df.Temperature['May 2010'].head().
df = pd.read_csv("datasets/weather_data_austin_2010.csv", parse_dates = True, index_col = "Date")
# Plot the summer data
df.Temperature['2010-Jun':'2010-Aug'].plot(figsize=(18, 5),style="g-");
# Plot the one week data
df.Temperature['2010-06-10':'2010-06-17'].plot(figsize=(18, 5), style="r+:")
# Plotting one week of temperature and pressure Dewpoint in one diagram with custom styles
df[["Temperature", "DewPoint"]]['2010-06-10':'2010-06-17'].plot(figsize=(18, 10), style=":s" , subplots= True);
| Datacamp Assignments/Data Science Track/Pandas Foundations/3. Time series in pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="DweYe9FcbMK_"
# ##### Copyright 2019 The TensorFlow Authors.
#
#
# + cellView="form" colab_type="code" id="AVV2e0XKbJeX" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="sUtoed20cRJJ"
# # 用 tf.data 加载 CSV 数据
# + [markdown] colab_type="text" id="1ap_W4aQcgNT"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://tensorflow.google.cn/tutorials/load_data/csv"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />在 Tensorflow.org 上查看</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/load_data/csv.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />在 Google Colab 运行</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/zh-cn/tutorials/load_data/csv.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />在 Github 上查看源代码</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/site/zh-cn/tutorials/load_data/csv.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />下载此 notebook</a>
# </td>
# </table>
# + [markdown] id="Z4x9SYyev8QZ" colab_type="text"
# Note: 我们的 TensorFlow 社区翻译了这些文档。因为社区翻译是尽力而为, 所以无法保证它们是最准确的,并且反映了最新的
# [官方英文文档](https://www.tensorflow.org/?hl=en)。如果您有改进此翻译的建议, 请提交 pull request 到
# [tensorflow/docs](https://github.com/tensorflow/docs) GitHub 仓库。要志愿地撰写或者审核译文,请加入
# [<EMAIL> Google Group](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-zh-cn)。
# + [markdown] colab_type="text" id="C-3Xbt0FfGfs"
# 这篇教程通过一个示例展示了怎样将 CSV 格式的数据加载进 `tf.data.Dataset`。
#
# 这篇教程使用的是泰坦尼克号乘客的数据。模型会根据乘客的年龄、性别、票务舱和是否独自旅行等特征来预测乘客生还的可能性。
# + [markdown] colab_type="text" id="fgZ9gjmPfSnK"
# ## 设置
# + colab_type="code" id="I4dwMQVQMQWD" colab={}
try:
# Colab only
# %tensorflow_version 2.x
except Exception:
pass
# + colab_type="code" id="baYFZMW_bJHh" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
import functools
import numpy as np
import tensorflow as tf
import tensorflow_datasets as tfds
# + colab_type="code" id="Ncf5t6tgL5ZI" colab={}
TRAIN_DATA_URL = "https://storage.googleapis.com/tf-datasets/titanic/train.csv"
TEST_DATA_URL = "https://storage.googleapis.com/tf-datasets/titanic/eval.csv"
train_file_path = tf.keras.utils.get_file("train.csv", TRAIN_DATA_URL)
test_file_path = tf.keras.utils.get_file("eval.csv", TEST_DATA_URL)
# + colab_type="code" id="4ONE94qulk6S" colab={}
# 让 numpy 数据更易读。
np.set_printoptions(precision=3, suppress=True)
# + [markdown] colab_type="text" id="Wuqj601Qw0Ml"
# ## 加载数据
#
# 开始的时候,我们通过打印 CSV 文件的前几行来了解文件的格式。
# + colab_type="code" id="54Dv7mCrf9Yw" colab={}
# !head {train_file_path}
# + [markdown] colab_type="text" id="YOYKQKmMj3D6"
# 正如你看到的那样,CSV 文件的每列都会有一个列名。dataset 的构造函数会自动识别这些列名。如果你使用的文件的第一行不包含列名,那么需要将列名通过字符串列表传给 `make_csv_dataset` 函数的 `column_names` 参数。
# + [markdown] colab_type="text" id="ZS-bt1LvWn2x"
#
#
#
#
# ```python
#
# CSV_COLUMNS = ['survived', 'sex', 'age', 'n_siblings_spouses', 'parch', 'fare', 'class', 'deck', 'embark_town', 'alone']
#
# dataset = tf.data.experimental.make_csv_dataset(
# ...,
# column_names=CSV_COLUMNS,
# ...)
#
# ```
#
# + [markdown] colab_type="text" id="gZfhoX7bR9u4"
# 这个示例使用了所有的列。如果你需要忽略数据集中的某些列,创建一个包含你需要使用的列的列表,然后传给构造器的(可选)参数 `select_columns`。
#
# ```python
#
# dataset = tf.data.experimental.make_csv_dataset(
# ...,
# select_columns = columns_to_use,
# ...)
#
# ```
# + [markdown] colab_type="text" id="67mfwr4v-mN_"
# 对于包含模型需要预测的值的列是你需要显式指定的。
# + colab_type="code" id="iXROZm5f3V4E" colab={}
LABEL_COLUMN = 'survived'
LABELS = [0, 1]
# + [markdown] colab_type="text" id="t4N-plO4tDXd"
# 现在从文件中读取 CSV 数据并且创建 dataset。
#
# (完整的文档,参考 `tf.data.experimental.make_csv_dataset`)
#
# + colab_type="code" id="Co7UJ7gpNADC" colab={}
def get_dataset(file_path):
dataset = tf.data.experimental.make_csv_dataset(
file_path,
batch_size=12, # 为了示例更容易展示,手动设置较小的值
label_name=LABEL_COLUMN,
na_value="?",
num_epochs=1,
ignore_errors=True)
return dataset
raw_train_data = get_dataset(train_file_path)
raw_test_data = get_dataset(test_file_path)
# + [markdown] colab_type="text" id="vHUQFKoQI6G7"
# dataset 中的每个条目都是一个批次,用一个元组(*多个样本*,*多个标签*)表示。样本中的数据组织形式是以列为主的张量(而不是以行为主的张量),每条数据中包含的元素个数就是批次大小(这个示例中是 12)。
#
# 阅读下面的示例有助于你的理解。
# + colab_type="code" id="qWtFYtwXIeuj" colab={}
examples, labels = next(iter(raw_train_data)) # 第一个批次
print("EXAMPLES: \n", examples, "\n")
print("LABELS: \n", labels)
# + [markdown] colab_type="text" id="9cryz31lxs3e"
# ## 数据预处理
# + [markdown] colab_type="text" id="tSyrkSQwYHKi"
# ### 分类数据
#
# CSV 数据中的有些列是分类的列。也就是说,这些列只能在有限的集合中取值。
#
# 使用 `tf.feature_column` API 创建一个 `tf.feature_column.indicator_column` 集合,每个 `tf.feature_column.indicator_column` 对应一个分类的列。
#
# + colab_type="code" id="mWDniduKMw-C" colab={}
CATEGORIES = {
'sex': ['male', 'female'],
'class' : ['First', 'Second', 'Third'],
'deck' : ['A', 'B', 'C', 'D', 'E', 'F', 'G', 'H', 'I', 'J'],
'embark_town' : ['Cherbourg', 'Southhampton', 'Queenstown'],
'alone' : ['y', 'n']
}
# + colab_type="code" id="kkxLdrsLwHPT" colab={}
categorical_columns = []
for feature, vocab in CATEGORIES.items():
cat_col = tf.feature_column.categorical_column_with_vocabulary_list(
key=feature, vocabulary_list=vocab)
categorical_columns.append(tf.feature_column.indicator_column(cat_col))
# + colab_type="code" id="H18CxpHY_Nma" colab={}
# 你刚才创建的内容
categorical_columns
# + [markdown] colab_type="text" id="R7-1QG99_1sN"
# 这将是后续构建模型时处理输入数据的一部分。
# + [markdown] colab_type="text" id="9AsbaFmCeJtF"
# ### 连续数据
# + [markdown] colab_type="text" id="o2maE8d2ijsq"
# 连续数据需要标准化。
#
# 写一个函数标准化这些值,然后将这些值改造成 2 维的张量。
#
# + colab_type="code" id="REKqO_xHPNx0" colab={}
def process_continuous_data(mean, data):
# 标准化数据
data = tf.cast(data, tf.float32) * 1/(2*mean)
return tf.reshape(data, [-1, 1])
# + [markdown] colab_type="text" id="VPsoMUgRCpUM"
# 现在创建一个数值列的集合。`tf.feature_columns.numeric_column` API 会使用 `normalizer_fn` 参数。在传参的时候使用 [`functools.partial`](https://docs.python.org/3/library/functools.html#functools.partial),`functools.partial` 由使用每个列的均值进行标准化的函数构成。
# + colab_type="code" id="WKT1ASWpwH46" colab={}
MEANS = {
'age' : 29.631308,
'n_siblings_spouses' : 0.545455,
'parch' : 0.379585,
'fare' : 34.385399
}
numerical_columns = []
for feature in MEANS.keys():
num_col = tf.feature_column.numeric_column(feature, normalizer_fn=functools.partial(process_continuous_data, MEANS[feature]))
numerical_columns.append(num_col)
# + colab_type="code" id="Bw0I35xRS57V" colab={}
# 你刚才创建的内容。
numerical_columns
# + [markdown] colab_type="text" id="M37oD2VcCO4R"
# 这里使用标准化的方法需要提前知道每列的均值。如果需要计算连续的数据流的标准化的值可以使用 [TensorFlow Transform](https://www.tensorflow.org/tfx/transform/get_started)。
# + [markdown] colab_type="text" id="kPWkC4_1l3IG"
# ### 创建预处理层
# + [markdown] colab_type="text" id="R3QAjo1qD4p9"
# 将这两个特征列的集合相加,并且传给 `tf.keras.layers.DenseFeatures` 从而创建一个进行预处理的输入层。
# + colab_type="code" id="3-OYK7GnaH0r" colab={}
preprocessing_layer = tf.keras.layers.DenseFeatures(categorical_columns+numerical_columns)
# + [markdown] colab_type="text" id="DlF_omQqtnOP"
# ## 构建模型
# + [markdown] colab_type="text" id="lQoFh16LxtT_"
# 从 `preprocessing_layer` 开始构建 `tf.keras.Sequential`。
# + colab_type="code" id="3mSGsHTFPvFo" colab={}
model = tf.keras.Sequential([
preprocessing_layer,
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(128, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid'),
])
model.compile(
loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
# + [markdown] colab_type="text" id="hPdtI2ie0lEZ"
# ## 训练、评估和预测
# + [markdown] colab_type="text" id="8gvw1RE9zXkD"
# 现在可以实例化和训练模型。
# + colab_type="code" id="sW-4XlLeEQ2B" colab={}
train_data = raw_train_data.shuffle(500)
test_data = raw_test_data
# + colab_type="code" id="Q_nm28IzNDTO" colab={}
model.fit(train_data, epochs=20)
# + [markdown] colab_type="text" id="QyDMgBurzqQo"
# 当模型训练完成的时候,你可以在测试集 `test_data` 上检查准确性。
# + colab_type="code" id="eB3R3ViVONOp" colab={}
test_loss, test_accuracy = model.evaluate(test_data)
print('\n\nTest Loss {}, Test Accuracy {}'.format(test_loss, test_accuracy))
# + [markdown] colab_type="text" id="sTrn_pD90gdJ"
# 使用 `tf.keras.Model.predict` 推断一个批次或多个批次的标签。
# + colab_type="code" id="Qwcx74F3ojqe" colab={}
predictions = model.predict(test_data)
# 显示部分结果
for prediction, survived in zip(predictions[:10], list(test_data)[0][1][:10]):
print("Predicted survival: {:.2%}".format(prediction[0]),
" | Actual outcome: ",
("SURVIVED" if bool(survived) else "DIED"))
| site/zh-cn/tutorials/load_data/csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={} colab_type="code" id="QL-ROwF131XU"
import numpy as np
import seaborn as sns
from math import sqrt
import matplotlib
import matplotlib.pyplot as plt
import matplotlib.ticker
# %matplotlib inline
import seaborn as sns
sns.set_context("talk")
# + [markdown] colab_type="text" id="xg7SLxza31XY"
# ### Метод Гаусса
# + colab={} colab_type="code" id="XT8_-XlZ31XZ"
def gauss(A):
m = len(A)
assert all([len(row) == m + 1 for row in A[1:]]), "Некорректная матрица! Строки разной длины!"
n = m + 1
for k in range(m):
pivots = [abs(A[i][k]) for i in range(k, m)]
i_max = pivots.index(max(pivots)) + k
assert A[i_max][k] != 0, "Матрица вырождена!"
A[k], A[i_max] = A[i_max], A[k]
for i in range(k + 1, m):
f = A[i][k] / A[k][k]
for j in range(k + 1, n):
A[i][j] -= A[k][j] * f
A[i][k] = 0
# Решаем уравнение Ax=b для верхне-треугольной матрицы A
x = list()
for i in range(m - 1, -1, -1):
x.insert(0, A[i][m] / A[i][i])
for k in range(i - 1, -1, -1):
A[k][m] -= A[k][i] * x[0]
return x
# + [markdown] colab_type="text" id="IY4PxVzY31Xd"
# ### Метод Зейделя
# + colab={} colab_type="code" id="v8tUQwHJ31Xe"
def seidel(A, b, optimal, discrepancy=False, eps=1e-8, x_0=None, max_iteration=100, verbose=False):
n = len(A)
x_1 = list()
iterator = 0
x = [.0 for i in range(n)] if x_0 is None else x_0
converge = False
while not converge:
x_new = np.copy(x)
for i in range(n):
s1 = sum(A[i][j] * x_new[j] for j in range(i))
s2 = sum(A[i][j] * x[j] for j in range(i + 1, n))
x_new[i] = (b[i] - s1 - s2) / A[i][i]
# оптимальный критерий останова
if optimal and len(x_1) != 0:
converge = norm_R**iterator / (1 - norm_R) * sqrt(sum((x_1[i]) ** 2 for i in range(n))) <= eps
if converge and verbose:
print(f'Оптимальный критерий останова сработал на {iterator+1} шаге')
elif discrepancy and len(x_1) != 0:
converge = sqrt(sum([x**2 for x in error(A, x_new, b)])) <= eps
if converge and verbose:
print(f'Стандартный критерий останова по невязке сработал на {iterator+1} шаге')
else:
converge = sqrt(sum((x_new[i] - x[i]) ** 2 for i in range(n))) <= eps
if converge and verbose:
print(f'Стандартный критерий останова по изменению значения за итерацию сработал на {iterator+1} шаге')
x = x_new
if iterator == 0:
x_1 = x_new
iterator += 1
if iterator > max_iteration:
raise ValueError("Не сходится за отведенное число итераций")
return x
# + [markdown] colab_type="text" id="YzEo0gW331Xi"
# ### Функция вычисления невязки
# + colab={} colab_type="code" id="8Tbx7EOi31Xj"
def error(A, x, b):
y = [x[0] for x in (np.matrix(A) * np.matrix(x).transpose()).tolist()]
return [abs(b[i]-y[i]) for i in range(len(b))]
# + colab={"base_uri": "https://localhost:8080/", "height": 136} colab_type="code" id="Ck0IzYpB6Op7" outputId="49aad85a-630e-43d7-94c8-7c695a84a91c"
# составляем тестовую расширенную матрицу А
n = 99
a = [1] * n
b = [10] * n
c = [1] * n
p = [1] * (n+1)
f = [1+x for x in range(n+1)]
test_1 = [[b[0], c[0]] + [0]*(n-1) + [f[0]]]
for i in range(n-1):
temp = [0]*(n+1)
temp[i:i+2] = a[i+1], b[i+1], c[i+1]
temp[-1] = f[i+1]
test_1.append(temp)
test_1.append(p+[f[n]])
# матрица А и столбец ответов b
A = [row[:-1] for row in test_1]
b = [row[-1] for row in test_1]
print(np.matrix(test_1))
# + [markdown] colab_type="text" id="wabgzDIy4iCh"
# ### Описание матрицы
# + colab={"base_uri": "https://localhost:8080/", "height": 68} colab_type="code" id="WrA78DA14gJu" outputId="d4bd25f5-06c7-4f34-b257-2338a3c158c1"
# вычисляем min и max собственные значения матрицы А
eigenvalues = np.linalg.eig(A)[0]
lambda_max = max(eigenvalues)
lambda_min = min(eigenvalues)
# вычисляем число обусловленности матрицы А
cond = np.linalg.cond(A)
print('lambda_min(A) = %f\nlambda_max(A) = %f\nчисл. обусловленности = ||A||*||A^(-1)|| = %f'\
%(lambda_min, lambda_max, cond))
# + colab={} colab_type="code" id="3p18J5PrM3Ln"
D = np.diag(np.diag(A))
L = np.tril(A, k = -1)
U = np.triu(A, k = 1)
R = - (np.matrix(np.linalg.inv(L+D)) * U)
norm_R = max([sum([abs(x_i) for x_i in x]) for x in np.transpose(R).tolist()])
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="7fRjg9-6OHFt" outputId="ee0175e1-ef20-4cc1-a7bf-3fa4e52a6825"
# вычисляем min и max собственные значения матрицы R
eigenvalues = np.linalg.eig(R)[0]
abs_lambda_max = max([max(eigenvalues), -min(eigenvalues)])
print('max(|lambda(R)|) = %0.3f\nnorm(R) = %0.3e' %(abs_lambda_max.real, norm_R))
# + [markdown] colab_type="text" id="X-dtsnpkP2T4"
# $\|B\| =$ 2.020e-01$< 1$
#
# $\Rightarrow$ выполнено достаточное условие сходимости итерационного метода, эквивалентного методу Зейделя
#
# **Оптимальный критерий останова:** $\|x^*-x_k\| \le \frac{\| B \|^k}{1- \|B\|} \| x_1 - x_0\| = \frac{\| B \|^k}{1- \|B\|} \| x_1\|$
# + [markdown] colab_type="text" id="gnYSSCat31Xm"
# ### Тест функций на пункте (a)
# + colab={} colab_type="code" id="SsDlgtzF31Xp"
# применяем метод Гаусса
gauss_res = gauss(test_1)
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="zrhOI8_K452f" outputId="4eb24ebf-c95b-4237-bb06-4692d720b9ab"
# применяем метод Зейделя
#@title Параметры метода Зейделя
max_iteration = 100 #@param {type:"slider", min:10, max:100, step:5}
eps = 1e-8 #@param {type:"number"}
optimal = True #@param {type:"boolean"}
discrepancy = False #@param {type:"boolean"}
seidel_res = seidel(A, b, optimal, discrepancy=discrepancy, max_iteration=max_iteration, eps=eps, verbose=True)
print(f'MAE(невязка) = {np.mean([abs(x) for x in error(A, seidel_res, b)])}')
# + [markdown] colab_type="text" id="ghiL5gpGbOPx"
# **Выводы:**
# 1. Применение оптимального критерия останова позволяет добиться сходимости на 17 шаге с невязкой *3.98e-15*
#
# 2. Применение стандартного критерия для $\varepsilon < $ *1e-15* позволяет методу сойтись за меньшее число итераций при меньшей точности невязки
# + [markdown] colab_type="text" id="et_61Zf531Xu"
# ### Инфографика
# + colab={"base_uri": "https://localhost:8080/", "height": 480} colab_type="code" id="Ht747nP_kSW1" outputId="7aafafd6-786f-4482-b629-a528546934b5"
xs = np.linspace(1e-10, 0.001, 200)
ys = [np.mean(error(A, seidel(A, b, False, max_iteration=max_iteration, eps=x), b)) for x in xs]
plt.title("График зависимости $MAE$ невязки от $\epsilon$")
plt.xlabel('$\epsilon$')
plt.ylabel('MAE')
plt.ylim((-0.1*max(ys), 1.1*max(ys)))
plt.xlim((-0.00005, 0.00105))
plt.scatter(xs, ys, color='darkslateblue', s=5);
# + [markdown] colab_type="text" id="oFJef4yUdS8I"
# **Комментарий:** сходимость имеет выраженный ступенчатый вид
# + [markdown] colab_type="text" id="RXmcZCHm63X8"
# ### Mетод SVD
# + [markdown] colab_type="text" id="nM-CxpUo7BN8"
# $A \, x = b$ -- СЛАУ
#
# $A = U\, S\, V^T$ -- сингулярное разложение матрицы $A$
#
# $x = V \, diag(\frac{1}{s_{ii}}) \, U^T\, b$ -- решение СЛАУ
# + colab={} colab_type="code" id="sW2XQ24b_CS-"
def diag_frac(S):
for i in range(len(S)):
S[i][i] = 1.0/S[i][i]
return S
# + colab={} colab_type="code" id="0BzkxNqt7AAG"
U, S, V_T = np.linalg.svd(A, full_matrices=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="sWXeTCe58V-l" outputId="1b6ea309-3da5-4d62-afd8-b2d3e65d79cf"
svd_res = [x[0] for x in (np.transpose(V_T) @ diag_frac(np.diag(S)) @ np.transpose(U) @ np.transpose(np.matrix(b))).tolist()]
print('MAE(м.SVD) = %.4e' %np.mean(error(A, svd_res, b)))
# + [markdown] colab_type="text" id="BT7r-8Xdn1CK"
# ### Метод верхней релаксации
# + colab={} colab_type="code" id="KDwUbwwHfRoj"
def SOR(A, b, eps, max_iteration, verbose=False):
D = np.diag(np.diag(A))
L = np.tril(A, k = -1)
U = np.triu(A, k = 1)
Q = np.linalg.inv(D + w * L)
K = np.dot(Q,((1.0 - w) * D - w * U))
F = w*np.dot(Q, b)
x = np.zeros_like(b)
for j in range(1, max_iteration):
x_new = np.dot(K, x) + F
converge = sqrt(sum((x_new[i] - x[i]) ** 2 for i in range(n))) <= eps
if converge:
if verbose:
print(f'Потребовалось {j} итераций')
return x_new
x = x_new
raise ValueError("Не сходится за отведенное число итераций")
# + colab={} colab_type="code" id="RBoAZ6kymu8W"
def optimal_w(A):
D = np.diag(np.diag(A))
L = np.tril(A, k = -1)
U = np.triu(A, k = 1)
R = - (np.matrix(np.linalg.inv(L+D)) * U)
spect_radius = max(map(abs, np.linalg.eig(R)[0]))
w_opt = 2 / (1 + sqrt(1-spect_radius**2))
return w_opt
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="lSKUryTxffBi" outputId="4c4f4586-608e-4aef-bfac-1ab5753fdc04"
w = 1 #@param {type:"slider", min:0, max:2, step:0.05}
max_iteration = 100 #@param {type:"slider", min:10, max:100, step:5}
eps = 1e-13 #@param {type:"number"}
w_optimal = True #@param {type:"boolean"}
if w_optimal:
w = optimal_w(A)
sor_res = SOR(A, b, eps, max_iteration, verbose=True)
print(f'MAE(невязка) = {np.mean([abs(x) for x in error(A, sor_res, b)])}')
# + [markdown] colab_type="text" id="TogR01ovnf-o"
# **Вывод:** применение оптимального веса $w$ позволяет добиться *MAE(невязки)* $\approx$*6.68e-15* на *17* шаге
# + [markdown] colab_type="text" id="IFR2tGbY31X2"
# ### Результаты
# _Оценим невязку используя, используя_ ***MAE*** _(средняя абсолютная ошибка) к вектору истинных ответов $b$ и ответам $b_0$, полученным с помощью выбранного метода._
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="aTikwzIS31X3" outputId="2d13722d-7457-4cab-f941-ec7d187ee88c"
gaus_mae = np.mean([abs(x) for x in error(A, gauss_res, b)])
seid_mae = np.mean([abs(x) for x in error(A, seidel(A, b, True, max_iteration=max_iteration, eps=eps, verbose=False), b)])
svd_mae = np.mean([abs(x) for x in error(A, svd_res, b)])
sor_mae = np.mean([abs(x) for x in error(A, sor_res, b)])
print('MAE(м.Гаусса) = %.4e\nMAE(м.Зейделя) = %0.4e, iters = 19\nMAE(м.SVD) = %0.4e\nMAE(м.SOR) = %0.4e, iters = 17' %(gaus_mae, seid_mae, svd_mae, sor_mae))
# + colab={} colab_type="code" id="N92e_bTR6dJ6"
#### ДОДЕЛАТЬ ГРАФИЧЕСКУЮ ЗАВИСИМОСТЬ КРИТЕРИЕВ ОСТАНОВА -- ЧИСЛО ВЕРНЫХ ЗНАКОВ ПОСЛЕ ЗАПЯТОЙ ДЛЯ РАЗНЫХ КРИТЕРИЕВ ОСТАНОВА (ОПТИМАЛЬНЫЙ, ПО ДЕЛЬТЕ, ПО НЕВЯЗКЕ)
#### ПОСТОРИТЬ ГРАФИК
#### ОБЩЕЕ ИСЛЕДОВАНИЕ НЕВЯЗКИ
# -
# ### Анализ взаимосвязей критериев останова
#
# * оптимальный: $\frac{\| B \|^k}{1- \|B\|} \| x_1 - x_0\| < \varepsilon$
# * по изменению значения за итерацию: $\| \Delta x_{k+1} \|_E = \sqrt{\sum \limits_{j=1}^n (x_{k+1}[j]-x_k[j])^2} < \varepsilon$
# * по невязке: $\| f_k - f\|_E = \sqrt{\sum \limits_{j=1}^n (f_{k}[j]-f[j])^2} < \varepsilon$, где $f_k = A \cdot x_k$ - невязка на $k-$ом шаге
# Рассмотрим число верных цифр в результате работы каждого из этих методов при различных значения параметра $\varepsilon$. Для интерпретации результата рассматриваются среднее значение числа верных цифр после запятой в векторе ответов $x$ и среднеквадратическое отклонение этой величины.
# + colab={} colab_type="code" id="xWhSeuTOGQfg"
# вычисление числа верных цифр после запятой
def right_digits(x, true_x):
max_by_element = [np.max([0]+[i for i in range(1, 16) if abs(x[j]-true_x[j])<10**(-i)]) for j in range(len(x))]
return np.mean(max_by_element), np.std(max_by_element)
# + colab={} colab_type="code" id="danuUxH6Do8d"
true_x = np.linalg.solve(A, b)
# + colab={} colab_type="code" id="Ss2X5uKbSs7B"
epss = [10**(-i) for i in np.linspace(0, 12, 120)]
# optimal
optimal, discrepancy = True, False
opt_coincides = list()
for eps in epss:
res = seidel(A, b, optimal, discrepancy=discrepancy,\
max_iteration=100, eps=eps, verbose=False)
opt_coincides.append(right_digits(res, true_x))
# delta
optimal, discrepancy = False, False
delta_coincides = list()
for eps in epss:
res = seidel(A, b, optimal, discrepancy=discrepancy,\
max_iteration=100, eps=eps, verbose=False)
delta_coincides.append(right_digits(res, true_x))
# discrepancy
coin_epss = epss[:]
optimal, discrepancy = False, True
coin_coincides = list()
for eps in coin_epss:
res = seidel(A, b, optimal, discrepancy=discrepancy,\
max_iteration=100, eps=eps, verbose=False)
coin_coincides.append(right_digits(res, true_x))
# +
xs = np.linspace(0, 12, 120)
ys = [x[0] for x in opt_coincides]
top_border = [x[0] + x[1] for x in opt_coincides]
low_border = list(reversed([x[0] - x[1] for x in opt_coincides]))
plt.figure(figsize=(12, 8))
plt.ylim((2, 16))
plt.xlim((-0.25, 12.25))
plt.xlabel('$i$: $\epsilon=10^{-i}$')
plt.ylabel('Число правильных цифр')
plt.title('''График зависимости числа правильных цифр после запятой от $\epsilon=10^{-i}$
для оптимального параметра останова''')
plt.plot(xs, ys, linestyle='--', label='среднее значение');
plt.plot([-1, 13], [15, 15], ':', c='red', lw=1.5)
plt.fill(list(xs)+list(reversed(xs)), top_border+low_border, alpha=0.3, label='интервал std');
plt.legend(loc=4);
# +
xs = np.linspace(0, 12, 120)
ys = [x[0] for x in delta_coincides]
top_border = [x[0] + x[1] for x in delta_coincides]
low_border = list(reversed([x[0] - x[1] for x in delta_coincides]))
plt.figure(figsize=(12, 8))
plt.ylim((2, 16))
plt.xlim((-0.25, 12.25))
plt.xlabel('$i$: $\epsilon=10^{-i}$')
plt.ylabel('Число правильных цифр')
plt.title('''График зависимости числа правильных цифр после запятой от $\epsilon=10^{-i}$
для остановки по изменению значения за итерацию''')
plt.plot(xs, ys, linestyle='--', label='среднее значение');
plt.plot([-1, 13], [15, 15], ':', c='red', lw=1.5)
plt.fill(list(xs)+list(reversed(xs)), top_border+low_border, alpha=0.3, label='интервал std');
plt.legend();
# +
xs = np.linspace(0, 12, 120)
ys = [x[0] for x in coin_coincides]
top_border = [x[0] + x[1] for x in coin_coincides]
low_border = list(reversed([x[0] - x[1] for x in coin_coincides]))
plt.figure(figsize=(12, 8))
plt.ylim((2, 16))
plt.xlim((-0.25, 12.25))
plt.xlabel('$i$: $\epsilon=10^{-i}$')
plt.ylabel('Число правильных цифр')
plt.title('''График зависимости числа правильных цифр после запятой от $\epsilon=10^{-i}$
для остановки по невязке''')
plt.plot(xs, ys, linestyle='--', label='среднее значение');
plt.plot([-1, 13], [15, 15], ':', c='red', lw=1.5)
plt.fill(list(xs)+list(reversed(xs)), top_border+low_border, alpha=0.3, label='интервал std');
plt.legend();
# -
eps = 1e-5
print('epsilon = 1e-5')
_ = seidel(A, b, True, discrepancy=False,
max_iteration=100, eps=eps, verbose=True)
_ = seidel(A, b, False, discrepancy=False,
max_iteration=100, eps=eps, verbose=True)
_ = seidel(A, b, False, discrepancy=True,
max_iteration=100, eps=eps, verbose=True)
eps = 1e-10
print('epsilon = 1e-10')
_ = seidel(A, b, True, discrepancy=False,
max_iteration=100, eps=eps, verbose=True)
_ = seidel(A, b, False, discrepancy=False,
max_iteration=100, eps=eps, verbose=True)
_ = seidel(A, b, False, discrepancy=True,
max_iteration=100, eps=eps, verbose=True)
eps = 1e-12
print('epsilon = 1e-5')
_ = seidel(A, b, True, discrepancy=False,
max_iteration=100, eps=eps, verbose=True)
_ = seidel(A, b, False, discrepancy=False,
max_iteration=100, eps=eps, verbose=True)
_ = seidel(A, b, False, discrepancy=True,
max_iteration=100, eps=eps, verbose=True)
# Не смотря на полученный результат второй и третий критерии не эквивалентны и совпадают лишь в рамках этой задачи. Что связано с устройством матрицы и установлена путем рассмотрения пошаговых значений.
| task_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import torch
from omegaconf import OmegaConf
from deepnote import MusicRepr
from importlib import reload
from pytorch_lightning import Trainer, seed_everything
from pytorch_lightning.loggers import TensorBoardLogger
from pytorch_lightning.callbacks import ModelCheckpoint, LearningRateMonitor
seed_everything(42)
# -
# ## Config
conf = OmegaConf.load('conf.yaml')
# ## Dataset
# +
from midi_transformer import LMDataset, get_dataloaders
dataset = LMDataset(
**conf['data']
)
# -
len(dataset)
train_loader, val_loader = get_dataloaders(dataset, batch_size=20, n_jobs=4, val_frac=0.1)
x, y = dataset[0]
x.shape, y.shape
for b in val_loader:
for k in b:
print(k, b[k].shape)
break
# ## Model
# +
from midi_transformer import CPTransformer
model = CPTransformer(conf['model'])
print('model has', model.count_parameters(), 'parameters.')
# -
model.step(b)
# ## Trainer
# +
name = '-'.join(conf['data']['instruments'])
print('model name:',name)
logger = TensorBoardLogger(save_dir='logs/', name=name)
lr_logger = LearningRateMonitor(logging_interval='step')
checkpoint = ModelCheckpoint(
dirpath=f'weights/{name}/',
filename='{epoch}-{val_loss:.2f}',
monitor='val_loss',
save_top_k=1,
period=1
)
trainer = Trainer(
benchmark=True,
gpus=1,
# reload_dataloaders_every_epoch=True,
# gradient_clip_val=0.5,
accumulate_grad_batches=1,
logger=logger,
max_epochs=conf['model']['max_epochs'],
callbacks=[checkpoint, lr_logger]
)
# -
trainer.fit(model, train_loader, val_loader)
trainer.save_checkpoint(f'weights/{name}/last.ckpt')
# ## generation
gen = model.generate(
prompt=None,
max_len=100,
temperatures={
'ttype' : 2,
'position': 0.9,
'tempo': 0.5,
'chord': 0.8,
'instrument': 3.,
'pitch': 2,
'duration': 0.8,
'velocity': 0.8
}
)
seq = MusicRepr.from_cp(gen)
seq.to_midi('gen.mid')
| examples/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Homework 1.2 - Pandas
# ### NAME:
# ### STUDENT ID:
# Load required modules
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
# ## Pandas Introduction
#
# ## 1. Reading File
# #### 1.1) Read the CSV file called 'data3.csv' into a dataframe called df.
# #### Data description
# * Data source: http://www.fao.org/nr/water/aquastat/data/query/index.html
# * Data, units:
# * GDP, current USD (CPI adjusted)
# * NRI, mm/yr
# * Population density, inhab/km^2
# * Total area of the country, 1000 ha = 10km^2
# * Total Population, unit 1000 inhabitants
# your code here
# #### 1.2) Display the first 10 rows of the dataframe.
# your code here
# #### 1.3) Display the column names.
# your code here
# #### 1.4) Use iloc to display the first 3 rows and first 4 columns.
# your code here
# ## 2. Data Preprocessing
#
# #### 2.1) Find all the rows that have 'NaN' in the 'Symbol' column. Display first 5 rows.
#
# ##### Hint : You might have to use a mask
# your code here
# #### 2.2) Now, we will try to get rid of the NaN valued rows and columns. First, drop the column 'Other' which only has 'NaN' values. Then drop all other rows that have any column with a value 'NaN'. Store the result in place. Then display the last 5 rows of the dataframe.
# your code here
# #### 2.3) For our analysis we do not want all the columns in our dataframe. Lets drop all the redundant columns/ features.
# #### **Drop columns**: **Area Id, Variable Id, Symbol**. Save the new dataframe as df1. Display the first 5 rows of the new dataframe.
# your code here
# #### 2.4) Display all the unique values in your new dataframe for each of the columns: Area, Variable Name, Year.
# your code here
# #### 2.5) Convert the 'Year' column float values to pandas datetime objects, where each year is represented as the first day of that year. Also display the first 5 values of the Year column after conversion.
#
# ##### For eg: 1962.0 will be represented as 1962-01-01
# your code here
# ## 3. Plot
# #### 3.1) Plot a bar graph showing the count for each unique value in the column 'Area'. Give it a title.
# your code here
# ## 4. Extract statistics from the data
#
# #### 4.1) Create a dataframe 'dftemp' to store rows where Area is 'Iceland'. Display the dataframe.
# your code here
# #### 4.2) Print the years (with the same format as 2.5) when the National Rainfall Index (NRI) was greater than 900 and less than 950 in Iceland. Use the dataframe you created in the previous question 'dftemp'.
# your code here
# ## 5. US statistics
# #### 5.1) Create a new DataFrame called **`df_usa`** that only contains values where 'Area' is equal to 'United States of America'. Set the indices to be the 'Year' column (Use .set_index( ), set inplace=True ). Display the dataframe head.
# your code here
# #### 5.2) Pivot the DataFrame so that the unique values in the column 'Variable Name' becomes the columns. The DataFrame values should be the ones in the the 'Value' column. Save it in df_usa. Display the dataframe head.
# your code here
# #### 5.3) Rename new columns to ['GDP','NRI','PD','Area','Population'] and display the head.
# your code here
# #### 5.4) Replace all 'Nan' values in df_usa with 0. Display the head of the dataframe.
# your code here
# ## 6. Use df_usa
#
# #### 6.1) Multiply the 'Area' column for all rows by 10 (so instead of 1000 ha, the unit becomes 100 ha = 1km^2). Display the dataframe head.
# your code here
# #### 6.2) Create a new column in df_usa called 'GDP/capita' and populate it with the calculated GDP per capita. Round the results to two decimal points. Display the dataframe head.
# GDP per capita = (GDP / Population) * 1000
# your code here
# #### 6.3) Find the maximum value of the 'NRI' column in the US (using pandas methods). What year does the max value occur? Display the values.
# your code here
# ### Congratulations on completing hw 1.2! Don't forget to click Kernel -> Restart & Run All, save your file, download or print as pdf, and submit pdf to Gradescope.
| x-archive-temp/m120-pandas/hw1.2_pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Task
# # Data
import numpy as np
import csv
from sklearn.preprocessing import MultiLabelBinarizer
from scipy.sparse import hstack
import pickle as pkl
from utils.tokenizer import tokenize_corpus
def getNames(data):
names = []
if not data:
return names
parsedData = eval(data)
if not parsedData:
return names
for pieceOfInfo in parsedData:
name = pieceOfInfo['name']
names.append(name)
return np.array(names)
with open('./data/links.csv', 'r', encoding='utf-8', newline='') as f:
reader = csv.reader(f)
next(reader, None)
id_to_movieId = dict()
for line in reader:
try:
id_to_movieId[int(line[2])] = int(line[0])
except:
pass
with open('./data/movies_metadata.csv', encoding= 'utf-8') as csvFile:
reader = csv.DictReader(csvFile)
i = 0
for row in reader:
dataEmbeded[i, 0] = row['overview']
try:
dataEmbeded[i, 1] = id_to_movieId[int(row['id'])]
except:
pass
dataEmbeded[i, 2] = row['adult'] == 1
dataEmbeded[i, 3] = row['budget']
dataEmbeded[i, 4] = getNames(row['genres'])
dataEmbeded[i, 5] = row['popularity']
dataEmbeded[i, 6] = getNames(row['production_companies'])
dataEmbeded[i, 7] = row['production_countries'] == "[{'iso_3166_1': 'US', 'name': 'United States of America'}]"
dataEmbeded[i, 8] = row['revenue']
dataEmbeded[i, 9] = getNames(row['spoken_languages'])
i += 1
one_hot = MultiLabelBinarizer(sparse_output=True)
genres = one_hot.fit_transform(dataEmbeded[:,4])
production_companies = one_hot.fit_transform(dataEmbeded[:,6])
spoken_languages = one_hot.fit_transform(dataEmbeded[:,9])
BoW = tokenize_corpus(dataEmbeded[:,0], stop_words = False, BoW = True)
data = hstack([BoW, genres, spoken_languages])
with open('./data/data.npy', 'wb') as pikeler:
data = {'ids':dataEmbeded[:, 1], 'data':data}
pkl.dump(data, pikeler)
# # Model
# ## Explication of base models
# ### Colaborative Deep Learning
#
# The first model on which we based ourselves is Hao Wang's model based on a Stacked Denoising Auto Encoder (SDAE), in charge of the item-based part. The principle this network is as follows:
# * We have a MLP neural network that is given a vector input and has to reproduce it as output.
# * A noise is applied to the input to make the network more robust
# * This network applies transformations to this vector until having a vector of small size compared to the input.
# * Then on a second part of the network, it reapplies transformations to this vector of small size until finding a vector of the same size as the entry. The loss is given by the difference between the input vector and the output vector in order to push the network to apply a reversible transformation within it.
# * In this way our network can be cut in half. A part that is an encoder that, given a large vector, encode a smaller, denser vector supposed to represent it. And a second part, able to decode this vector to find the original vector.
#
# This type of network is particularly interesting with bag of words approach because it gives at first a vector often very sparse with the size of the vocabulary, unusable without size reduction.
#
# <img src="./images/SDAE.png" width=300px>
#
# On the other hand, for the collaborative part, embeddings are created for the users and items. Embeddings are widely used in other filed of domain (notably NLP), but are particularly adapted for this application. Indeed, embeddings are dense vectors representing an entity, the closer entities are, the closer their embeddings will be.
#
# After that, the item embedding and the dense vector created by the SDAE are concatenated making the full item embedding.
# Once this is done, the user and full item embedding are multiplied to form the ratings predictions.
#
# <img src="./images/MF.png" width=600px>
#
# The full architected is as follow:
#
# <img src="./images/CDL.png" width=400px>
# ### Neural Collaborative Filter
#
# The second model is based on the first one, however <NAME> et al. that the matrix multiplication is suboptimal and doesn't have enough capacity to represent the non-linear relations between users, items and ratings. It is therefore proposed to replace the multiplication by a neural network.
#
# <img src="./images/NCF_1.png" width=400px>
#
# The intuition behind this is that matrix multiplication is a special case of the MLP. Indeed, with the right weights (identity), a network can simply give the result of a matrix multiplication. Like so:
#
# <img src="./images/NCF_3.png" width=200px>
#
# <img src="./images/NCF_2.png" width=400px>
#
# However, empirical results showed that keeping the matrix multiplication still yield better results. The model they propose is then the following:
# <img src="./images/NCF_4.png" width=400px>
# ### Our model: Neural Hybrid Recommender
#
# We kept the main ideas proposed earlier but added a couple of improvements:
# * Addition of regularization layers (Batch-norm and Dropout)
# * Concatation of the SDAE to the Neural Collaborative Filter
# * Use of Adam optimizer
#
# The batch-norm improves the Convergence speed and Dropout prevents over-fitting. Adam optimizer adds Momentum en Nesterov Momentum and has proven to fasten the optimization.
#
# The model is then:
#
# <img src="./images/NHR.png" width=400px>
# # Results
| Project/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys
import os.path
import json
# sys.path.append(
# os.path.abspath(os.path.join(os.path.dirname(__file__), os.path.pardir)))
# sys.path.append('../')
# import models.models as database
# from sqlalchemy.exc import IntegrityError
# from config.config import env
import numpy
from sklearn import cross_validation
from sklearn.feature_extraction.text import TfidfVectorizer
from sklearn.feature_selection import SelectPercentile, f_classif
def process_text(word_file, profile_file):
sp_stop_words = import_sp_stop_words('../local_data/stop_words_es.json')
## FEATURES AS PARAGRAPHS OF TEXT
with open(word_file, 'r') as words:
word_data = [x for x in words]
## LABELS WITH PEOPLE PROFILES
with open(profile_file, 'r') as profiles:
profile_data = [y for y in profiles]
# print(word_data, profile_data)
assert len(word_data) == len(profile_data)
print(len(word_data) == len(profile_data))
features_train, features_test, labels_train, labels_test = cross_validation.train_test_split(word_data, profile_data, test_size=0.1, random_state=42)
### text vectorization--go from strings to lists of numbers
vectorizer = TfidfVectorizer(sublinear_tf=True, max_df=0.5,
stop_words=sp_stop_words)
# print vectorizer
features_train_transformed = vectorizer.fit_transform(features_train)
features_test_transformed = vectorizer.transform(features_test)
selector = SelectPercentile(f_classif, percentile=1)
selector.fit(features_train_transformed, labels_train)
features_train_transformed = selector.transform(features_train_transformed).toarray()
features_test_transformed = selector.transform(features_test_transformed).toarray()
print(features_train_transformed)
print(features_test_transformed)
return features_train_transformed, features_test_transformed, labels_train, labels_test
process_text('../local_data/texto_perfiles.csv', '../local_data/labels.csv')
# -
import json
def import_sp_stop_words(jsonfile):
with open(jsonfile, 'r') as file:
jsondict = json.load(file)
return jsondict['stop_words_es']
| bot/.ipynb_checkpoints/text_process-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
from PIL import Image
import numpy as np
# +
first_class_train = 'data/1class_train/'
second_class_train = 'data/2class_train/'
third_class_train = 'data/3class_train/'
first_class_test = 'data/1class_test/'
second_class_test = 'data/2class_test/'
third_class_test = 'data/3class_test/'
# +
def norm_image(img):
"""
Normalize PIL image
Normalizes luminance to (mean,std)=(0,1), and applies a [1%, 99%] contrast stretch
"""
img_y, img_b, img_r = img.convert('YCbCr').split()
img_y_np = np.asarray(img_y).astype(float)
img_y_np /= 255
img_y_np -= img_y_np.mean()
img_y_np /= img_y_np.std()
scale = np.max([np.abs(np.percentile(img_y_np, 1.0)),
np.abs(np.percentile(img_y_np, 99.0))])
img_y_np = img_y_np / scale
img_y_np = np.clip(img_y_np, -1.0, 1.0)
img_y_np = (img_y_np + 1.0) / 2.0
img_y_np = (img_y_np * 255 + 0.5).astype(np.uint8)
img_y = Image.fromarray(img_y_np)
img_ybr = Image.merge('YCbCr', (img_y, img_b, img_r))
img_nrm = img_ybr.convert('RGB')
return img_nrm
def norm_image_(img):
print(type(img))
return (img - 127) / 255
# -
def imgToArray(image):
x=image.convert('L')
y=np.asarray(x.getdata(),dtype=np.float64).reshape((x.size[1],x.size[0]))
y=np.asarray(y,dtype=np.uint8)
return (y - 127) / 255
def resize_image(img, size):
n_x, n_y = img.size
if n_y > n_x:
n_y_new = size
n_x_new = int(size * n_x / n_y + 0.5)
else:
n_x_new = size
n_y_new = int(size * n_y / n_x + 0.5)
img_res = img.resize((n_x_new, n_y_new), resample=Image.BICUBIC)
img_pad = Image.new('RGB', (size, size), (128, 128, 128))
ulc = ((size - n_x_new) // 2, (size - n_y_new) // 2)
img_pad.paste(img_res, ulc)
return img_pad
# +
new_size = 224
def getImages_wo_resize(path):
names = os.listdir(path)
img_arr = []
for name in names:
p = path + name
img = Image.open(p)
img_arr.append(img)
return img_arr
def getImages(path):
names = os.listdir(path)
img_arr = []
for name in names:
p = path + name
try:
img = Image.open(p)
img_arr.append(imgToArray(resize_image(norm_image(img), new_size)))
except OSError:
print('remove ' + p)
os.remove(p)
print('remove ' + 'data/images/' + name)
os.remove('data/images/' + name)
return img_arr
def resize(img, size):
#wpercent = basewidth / img.size[0]
#hsize = int((float(img.size[1]) * float(wpercent)))
#return img.resize((basewidth,hsize), Image.ANTIALIAS)
return img.resize((size, size), Image.ANTIALIAS)
# +
first_class_train_img = getImages(first_class_train)
second_class_train_img = getImages(second_class_train)
third_class_train_img = getImages(third_class_train)
first_class_test_img = getImages(first_class_test)
second_class_test_img = getImages(second_class_test)
third_class_test_img = getImages(third_class_test)
# +
from keras.models import Sequential
from keras.layers import Dense, Dropout, Activation, Flatten
from keras.layers.convolutional import Convolution2D, MaxPooling2D
from keras.utils import np_utils
img_rows, img_cols = 224, 224
# +
# make (X_train, y_train), (X_test, y_test)
import numpy as np
X_train = np.array(first_class_train_img + second_class_train_img +third_class_train_img)
y_train = np.array([np.repeat(0, len(first_class_train_img))] +
[np.repeat(1, len(second_class_train_img))] +
[np.repeat(2, len(second_class_train_img))])
X_test = np.array(first_class_test_img+ second_class_test_img+ third_class_test_img)
y_test = np.array([np.repeat(0, len(first_class_test_img))] +
[np.repeat(1, len(second_class_test_img))] +
[np.repeat(2, len(second_class_test_img))])
len_train = len(first_class_train_img) + len(second_class_train_img) + len(third_class_train_img)
print(len(first_class_train_img))
print(X_train.shape)
print(len_train)
#X_train = X_train.reshape(len_train)
y_train = y_train.reshape(len_train)
len_test = len(first_class_test_img) + len(second_class_test_img) + len(third_class_test_img)
#X_test = X_test.reshape(len_test)
y_test = y_test.reshape(len_test)
y_test.shape
# -
img_rows, img_cols = 224, 224
# +
#theano
X_train = X_train.reshape(X_train.shape[0], 1, img_rows, img_cols)
X_test = X_test.reshape(X_test.shape[0], 1, img_rows, img_cols)
input_shape = (1, img_rows, img_cols)
X_train = X_train.astype('float32')
X_test = X_test.astype('float32')
#X_train /= 255
#X_test /= 255
Y_train = np_utils.to_categorical(y_train, 3)
Y_test = np_utils.to_categorical(y_test, 3)
model = Sequential()
model.add(Convolution2D(20, 5, 5, border_mode='valid', input_shape=input_shape))
model.add(Activation('relu'))
model.add(Convolution2D(20, 5, 5))
model.add(Activation('relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(128))
model.add(Activation('relu'))
model.add(Dropout(0.5))
model.add(Dense(3))
model.add(Activation('softmax'))
model.compile(loss='categorical_crossentropy', optimizer='adam', metrics=['accuracy'])
# +
nb_epoch = 12
res = model.fit(X_train, Y_train, batch_size=128, nb_epoch=nb_epoch, verbose=1, validation_data=(X_test, Y_test), shuffle=True)
score = model.evaluate(X_test, Y_test, verbose=0)
# +
# %matplotlib inline
import matplotlib.pyplot as plt
x = range(nb_epoch)
plt.plot(x, res.history['acc'], label="train acc")
plt.plot(x, res.history['val_acc'], label="val acc")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
plt.show()
plt.plot(x, res.history['loss'], label="train loss")
plt.plot(x, res.history['val_loss'], label="val loss")
plt.legend(loc='center left', bbox_to_anchor=(1, 0.5))
| Pics5000.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Presentation Visualization
import matplotlib
import matplotlib.animation
# %matplotlib inline
import matplotlib.pyplot as plt
plt.ioff()
from IPython.display import HTML
# +
import math
import mantrap
import torch
env = mantrap.environment.Trajectron(ego_position=torch.tensor([0.0, -1.0]))
env.add_ado(position=torch.tensor([3, 2]), velocity=torch.tensor([0, -1.0]))
env.add_ado(position=torch.tensor([-3, 7]), velocity=torch.tensor([1.0, -1.0]))
env.add_ado(position=torch.tensor([-4, -3]), velocity=torch.tensor([1.0, 1.0]))
# -
# ### Why is the socially-aware navigation difficult ?
# +
fig, ax = plt.subplots(1, 1, figsize=(20, 9))
N = 20
def update(n: int):
plt.axis("off")
ax.cla()
ego_controls = torch.stack([torch.tensor([1, 0])] * 10)
ego_controls[0, 1] = - n / N * 4
ego_controls[5:, 1] = n / N * 4
env.visualize_prediction_w_controls(ego_controls, ax=ax, display_wo=True)
return ax
anim = matplotlib.animation.FuncAnimation(fig, update, frames=N + 1, interval=300)
anim.save(f"/Users/sele/Desktop/difficult.gif", dpi=60, writer='imagemagick')
# -
# ### Goal-Based Trajectory Solver
env = mantrap.environment.SocialForcesEnvironment(ego_position=torch.tensor([0.0, -1.0]))
env.add_ado(position=torch.tensor([3, 2]), velocity=torch.tensor([0, -1.0]))
env.add_ado(position=torch.tensor([-3, 7]), velocity=torch.tensor([1.0, -1.0]))
env.add_ado(position=torch.tensor([-4, -3]), velocity=torch.tensor([1.0, 1.0]))
# +
solver = mantrap.solver.IPOPTSolver(env=env, goal=torch.tensor([8, 0]), is_logging=True,
modules=[mantrap.modules.GoalNormModule, mantrap.modules.SpeedLimitModule],
eval_env=mantrap.environment.Trajectron)
_, _, = solver.solve(time_steps=30)
solver.visualize_scenes(save=True, figsize=(20, 9))
# -
# ### Interactive Trajectory Solver
# +
solver = mantrap.solver.IPOPTSolver(env=env, goal=torch.tensor([8, 0]), is_logging=True,
eval_env=mantrap.environment.Trajectron)
_, _, = solver.solve(time_steps=30)
solver.visualize_scenes(save=True, figsize=(20, 9))
# +
env = mantrap.environment.PotentialFieldEnvironment(ego_position=torch.tensor([0.0, -1.0]))
env.add_ado(position=torch.tensor([3, 2]), velocity=torch.tensor([0, -1.0]))
env.add_ado(position=torch.tensor([-3, 7]), velocity=torch.tensor([1.0, -1.0]))
env.add_ado(position=torch.tensor([-4, -3]), velocity=torch.tensor([1.0, 1.0]))
solver = mantrap.solver.IPOPTSolver(env=env, goal=torch.tensor([8, 0]), is_logging=True,
eval_env=mantrap.environment.Trajectron)
_, _, = solver.solve(time_steps=30)
solver.visualize_scenes(save=True, figsize=(20, 9))
# +
env = mantrap.environment.SocialForcesEnvironment(ego_position=torch.tensor([0.0, -1.0]))
env.add_ado(position=torch.tensor([7, -3]), velocity=torch.tensor([0, -1.0]))
env.add_ado(position=torch.tensor([-5, 2]), velocity=torch.tensor([1.0, 0.2]))
env.add_ado(position=torch.tensor([5, 2]), velocity=torch.tensor([1.0, 0.2]))
env.add_ado(position=torch.tensor([-2, 2]), velocity=torch.tensor([1.0, -1.5]))
solver = mantrap.solver.IPOPTSolver(env=env, goal=torch.tensor([8, 0]), is_logging=True,
eval_env=mantrap.environment.Trajectron)
_, _, = solver.solve(time_steps=30)
solver.visualize_scenes(save=True, figsize=(20, 9))
# +
env = mantrap.environment.Trajectron(ego_position=torch.tensor([-5.0, -1.0]),
ego_type=mantrap.agents.IntegratorDTAgent)
env.add_ado(position=torch.tensor([3, 2]), velocity=torch.tensor([1.0, -0.1]))
env.add_ado(position=torch.tensor([-3, 2]), velocity=torch.tensor([0.2, -0.8]))
env.add_ado(position=torch.tensor([0, 0]), velocity=torch.tensor([-0.2, 0.6]))
env.add_ado(position=torch.tensor([4, -3]), velocity=torch.tensor([0.0, -1.0]))
env.add_ado(position=torch.tensor([-5, -3]), velocity=torch.tensor([1.4, -0.2]))
ego_controls = torch.zeros((10, 2))
ego_controls[:10, 0] = torch.ones(10) * 3.0
ego_controls[:10, 1] = - torch.tensor([2 * math.cos(math.pi / 5 * k) for k in range(10)])
env.visualize_prediction_w_controls(ego_controls, grid=False)
plt.savefig("/Users/sele/Desktop/title.png", dpi=300)
# +
env = mantrap.environment.SocialForcesEnvironment(ego_position=torch.tensor([0.0, -1.0]))
for _ in range(6):
position_random = torch.rand(2) * 16.0 - 8.0
velocity_random = torch.rand(2) * 2.0 - 1.0
env.add_ado(position=position_random, velocity=velocity_random)
solver = mantrap.solver.IPOPTSolver(env=env, goal=torch.tensor([8, 0]), is_logging=True,
eval_env=mantrap.environment.Trajectron)
_, _, = solver.solve(time_steps=30)
solver.visualize_scenes(save=True, figsize=(20, 9))
# -
# ### ORCA Failures
# +
import numpy as np
orca_results = np.loadtxt("/Users/sele/Desktop/videos/orca.txt")
# +
solver = mantrap.solver.baselines.ORCASolver(env=env, goal=torch.tensor([8, 0]), is_logging=True,
eval_env=mantrap.environment.Trajectron)
_, _, = solver.solve(time_steps=30)
solver.visualize_scenes(save=True, figsize=(20, 9))
# -
# ### IPOPT Failures
# +
env = mantrap.environment.Trajectron(ego_position=torch.tensor([0.0, 0.0]))
env.add_ado(position=torch.tensor([1, 0]), velocity=torch.tensor([-1.0, 0.0]))
solver = mantrap.solver.IPOPTSolver(env=env, goal=torch.tensor([8, 0]), is_logging=True)
_, _, = solver.solve(time_steps=20)
solver.visualize_scenes(save=True, figsize=(20, 9))
# -
# ### Gradient Map
# +
import numpy as np
N = 40
env = mantrap.environment.Trajectron(ego_position=torch.tensor([0.0, -1.0]))
env.add_ado(position=torch.tensor([3, 1]), velocity=torch.tensor([0, -1.0]))
solver = mantrap.solver.IPOPTSolver(env=env, goal=torch.tensor([8, 0]), t_planning=1)
heatmap = np.zeros((N, N, 2))
for ix, x in enumerate(np.linspace(-2, 2, num=N)):
for iy, y in enumerate(np.linspace(-2, 2, num=N)):
heatmap[ix, iy, :] = solver.gradient(z=np.zeros((1, 2)))
magmap = np.linalg.norm(heatmap, axis=-1)
plt.cla()
plt.imshow(magmap)
plt.axis("off")
plt.savefig("/Users/sele/Desktop/gradmap.png", dpi=100)
# -
env.visualize_prediction_w_controls(torch.zeros(10, 2))
plt.savefig("/Users/sele/Desktop/gradmap_scene.png", dpi=100)
# ### Multi-Modal Probability Distribution
# +
import math
import numpy as np
def gaussian(x, mean: float, std: float) -> float:
return 1.0 / (std * math.sqrt(2 * math.pi)) * np.exp(-0.5 * ((x - mean) / std) ** 2)
x = np.linspace(0, 4, num=100)
gaussians = []
for mean, std in zip([2.0, 1.0, 1.3], [0.6, 0.4, 0.8]):
g = gaussian(x, mean=mean, std=std)
gaussians.append(g)
fig, ax = plt.subplots(1, 1)
for g in gaussians:
plt.plot(x, g, "-", color="g")
plt.xticks([])
plt.yticks([])
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
ax.spines['left'].set_linewidth(3)
ax.spines['left'].set_color("k")
ax.spines['bottom'].set_linewidth(3)
ax.spines['bottom'].set_color("k")
def arrowed_spines(ax=None, arrowLength=20, labels=('X', 'Y'), arrowStyle='<|-'):
xlabel, ylabel = labels
for i, spine in enumerate(['left', 'bottom']):
# Set up the annotation parameters
t = ax.spines[spine].get_transform()
xy, xycoords = [1, 0], ('axes fraction', t)
xytext, textcoords = [arrowLength, 0], ('offset points', t)
# create arrowprops
arrowprops = dict( arrowstyle=arrowStyle,
facecolor=ax.spines[spine].get_facecolor(),
linewidth=ax.spines[spine].get_linewidth(),
alpha = ax.spines[spine].get_alpha(),
zorder=ax.spines[spine].get_zorder(),
linestyle = ax.spines[spine].get_linestyle() )
if spine is 'bottom':
ha, va = 'left', 'center'
xarrow = ax.annotate(xlabel, xy, xycoords=xycoords, xytext=xytext,
textcoords=textcoords, ha=ha, va='center', fontsize=20,
arrowprops=arrowprops)
else:
ha, va = 'center', 'bottom'
yarrow = ax.annotate(ylabel, xy[::-1], xycoords=xycoords[::-1],
xytext=xytext[::-1], textcoords=textcoords[::-1],
ha='center', va=va, arrowprops=arrowprops, fontsize=20)
return xarrow, yarrow
arrowed_spines(ax=ax, labels=("x", "p(x)"))
plt.savefig("/Users/sele/Desktop/gmm.png")
| examples/presentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 安裝
# ! pip install .
# ## 查看主程式幫助
# ! cvtwd -h
# ## 更新匯率表
# ! cvtwd update -h
# ! cvtwd update
# ## 查看支援的貨幣信息
# ! cvtwd info -h
# ! cvtwd info
# ## 查看匯率
# ! cvtwd lookup -h
# ! cvtwd lookup
# ! cvtwd lookup -c EUR AUD SGD
# #### 防呆機制
# ! cvtwd lookup -c USB RMB
# ## 換算匯率
# ! cvtwd convert -h
# ! cvtwd convert 300 USD
# ! cvtwd convert 300 USD spot
# ! cvtwd convert 300 USD spot JPY
# ! cvtwd convert 300 USD spot JPY spot
# #### 防呆機制
# ! cvtwd convert 300 JPN
#
| Example_commands.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/rodrigowe1988/Desafio-de-Data-Science/blob/main/DSNP_3_0_Lista_de_Exerc%C3%ADcios.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="vvplGyW2N3rX"
# # Lista de Exercícios
#
# A lista de exercícios tem por objetivo colocar a mão na massa e relembrar os conceitos que foram passados nas aulas anteriores.
#
# <center><img src="https://images.unsplash.com/photo-1542903660-eedba2cda473?ixid=MnwxMjA3fDB8MHxwaG90by1wYWdlfHx8fGVufDB8fHx8&ixlib=rb-1.2.1&auto=format&fit=crop&w=1050&q=80" height="400px"></center>
#
# Optei por não necessariamente seguir a sequencia das aulas, colocando os exercícios
# + [markdown] id="9GQoXiddIpr9"
# ## Exercício 1
#
# * Importar o pacote Pandas
# * importar os dados de ocorrências da aviação civil neste link
# * Ver qual o tamanho (linhas x colunas) do *DataFrame*
# * Verificar as 5 primeiras
# + [markdown] id="hdRXuNktKw9p"
# **Importe o Pandas e importe os dados de ocorrências da aviação civil que se encontram em `csv` [neste link](https://raw.githubusercontent.com/carlosfab/dsnp2/master/datasets/ocorrencias_aviacao.csv).**
# + id="8TpD5o8MLMaV"
import pandas as pd
# + [markdown] id="Tf8ScLP9LRjq"
# **Veja qual o tamanho (linhas e colunas) do conjunto de dados importado.**
# + id="vGsqOw52LnKZ"
df = pd.read_csv("https://raw.githubusercontent.com/carlosfab/dsnp2/master/datasets/ocorrencias_aviacao.csv")
# + [markdown] id="OvvxJlZXL-BH"
# **Identifique o tipo de cada coluna (inteiro, float, object).**
# + id="MjAmdNplMOMJ" outputId="29ec3937-25eb-4579-99dc-e5b307463f3e" colab={"base_uri": "https://localhost:8080/"}
print(f"O DataFrame possui {df.shape[0]} linhas e {df.shape[1]} colunas.")
# + [markdown] id="ew5bo-eZMTCZ"
# **Veja as 3 primeiras entradas do conjunto de dados.**
# + id="e1xx_cXFMaM2" outputId="8c26e302-f98c-416e-d3ba-e395de920d25" colab={"base_uri": "https://localhost:8080/", "height": 247}
df.head(3)
# + [markdown] id="p_2rj-KqMbtA"
# **Veja quantos valores únicos existem para a coluna `ocorrencia_classificacao`**
# + id="DVtxiIREMnnV" outputId="dacbb125-5617-433b-bd33-e774b0411523" colab={"base_uri": "https://localhost:8080/"}
df.ocorrencia_classificacao.unique()
# + [markdown] id="ZoUq8qxPMqPj"
# **Descubra qual Estado possui o maior número de ocorrências com aeronaves civis.**
# + id="Fdin50sWM29y" outputId="0ec97daf-f812-4329-aa9d-9c19beb91199" colab={"base_uri": "https://localhost:8080/"}
df.ocorrencia_uf.value_counts().head(1)
# + [markdown] id="jaE4lKoMNDNG"
# **Quantos por cento das ocorrências são do tipo `FALHA DO MOTOR EM VOO`?**
# + id="MZNDxepbNqFr" outputId="6054855d-7572-4a81-dcc6-25054a1efe0a" colab={"base_uri": "https://localhost:8080/"}
porcentagem_falha = df.loc[df.ocorrencia_tipo == "FALHA DO MOTOR EM VOO"].shape[0] / df.shape[0] * 100
print(f"A porcentagem de acidente com FALHA DO MOTOR é de {porcentagem_falha:.2f}%.")
# + [markdown] id="BZCNVc1NOVeb"
# ## Exercício 2
#
# Você irá trabalhar agora com os dados da ação BBAS3, do Banco do Brasil na BOVESPA.
#
# Na variável `df_bbas3`, foi importado [este arquivo `csv`](https://raw.githubusercontent.com/carlosfab/dsnp2/master/datasets/BBAS3.SA.csv), onde o *index* do *DataFrame* representa os valores da ação para determinado dia. Responda as perguntas abaixo.
# + id="4XZI_FjDPFsR"
# importar pandas
import pandas as pd
# importar o csv com dados da BBAS3
df_bbas3 = pd.read_csv("https://raw.githubusercontent.com/carlosfab/dsnp2/master/datasets/BBAS3.SA.csv",
index_col="Date")
# + [markdown] id="j5gGguY3P67Q"
# **Veja as 5 primeiras entradas da variável `df_bbas3`.**
# + id="MlGjWTeXQEB3" outputId="605b2885-5ebb-46bd-fc09-4a68cdba580f" colab={"base_uri": "https://localhost:8080/", "height": 235}
df_bbas3.head()
# + [markdown] id="O3tTLDfTQTGZ"
# **Quais são os valores da ação para o dia 04 de outubro de 2019?**
# + id="kPSOenAuQj81" outputId="6c877429-62ec-42f4-ddf3-bf3960a9e1ba" colab={"base_uri": "https://localhost:8080/"}
df_bbas3.loc["2019-10-04"].Close
# + [markdown] id="PRZtV761Qr0-"
# **Qual a média do Volume financeiro movimentado em todo o período?**
# + id="n-vZleQPEGey" outputId="ec3fd3b2-a0fb-48b5-e13b-8e65b06e17e3" colab={"base_uri": "https://localhost:8080/"}
df_bbas3.Volume.mean()
# + [markdown] id="_pO0tCV1EdD0"
# ## Exercício 3
#
# Você irá trabalhar agora com os dados do projeto +BIKE, usado em aulas passadas.
#
# Na variável df_bike, foi importado [este arquivo csv](http://dl.dropboxusercontent.com/s/yyfeoxqw61o3iel/df_rides.csv).
#
#
# + id="YFvJfTuVE0da" outputId="96fe6214-3e2e-4872-ab2c-d8ed09d02a85" colab={"base_uri": "https://localhost:8080/", "height": 272}
# importar pandas
import pandas as pd
# importar o csv com dados da BBAS3
df_bike = pd.read_csv("http://dl.dropboxusercontent.com/s/yyfeoxqw61o3iel/df_rides.csv")
# ver as primeiras entradas
df_bike.head()
# + [markdown] id="sj3lwZctE7KP"
# **Qual a porcentagem de valores ausentes para a coluna `user_gender`?**
#
# + id="g8GJ7kgLFEbr" outputId="1eda399e-44e4-4d06-de67-434bbe131187" colab={"base_uri": "https://localhost:8080/"}
gender_null = df_bike.user_gender.isnull().sum()
print(f"A coluna user_gender possui {gender_null} linhas nulas.")
# + [markdown] id="oDVOu-XpFZYI"
# **Preencha a coluna `user_gender` com o valor mais frequente.**
# + id="7f9WWlD6FZnJ" outputId="0ce23dff-b7ea-46d9-ff12-2c67a2de55cd" colab={"base_uri": "https://localhost:8080/"}
#descobrindo qual o valor mais frequente (moda)
df_bike.user_gender.value_counts()
# + id="VlxhUHDi-6vD"
df_bike.fillna('M', inplace=True)
# + id="yBec6K0A_OH_" outputId="a6855181-3d66-4709-f3aa-e8844b771218" colab={"base_uri": "https://localhost:8080/"}
#após o preenchimento dos valores nulos
df_bike.user_gender.value_counts()
# + [markdown] id="ckDXwzIzF7XT"
# **Plote um boxplot para a coluna `ride_duration`.**
# + id="xIluDO2wGGE6" outputId="c29b442d-b3cc-4163-dcc0-5f2066bbd422" colab={"base_uri": "https://localhost:8080/", "height": 283}
df_bike.ride_duration.plot(kind="box")
| DSNP_3_0_Lista_de_Exercícios.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.9.7 64-bit
# language: python
# name: python3
# ---
# +
from tensorflow.keras.utils import to_categorical
import pandas as pd
import os
import pandas as pd
import soundfile as sf
import numpy as np
from sklearn.utils import shuffle
from sys import platform
from sklearn.model_selection import train_test_split, GroupShuffleSplit
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
seed=42
n_class="3class"
class_dist = {0:2462,1:3000,2:3000}
# +
cols=['video_id','start_time','mid_ts','label','audio','vggish']
d=np.load(f'./resources/working_data/vocal_only_data_with_vggish.npy',allow_pickle=True)
df = pd.DataFrame(d,columns=cols)
lut = pd.read_csv(f'../dataset/lookup.csv')
df=df.merge(lut[['video_id','band_name']],on='video_id')
feature_df=df[['label','audio','band_name','video_id']]
mapping=[]
for index,row in feature_df.iterrows():
if row['label'] == 'clean':
mapping.append(0)
if row['label'] == 'highfry':
mapping.append(1)
if row['label'] == 'layered':
mapping.append(1)
if row['label'] == 'lowfry':
mapping.append(1)
if row['label'] == 'midfry':
mapping.append(1)
if row['label'] == 'no_vocals':
mapping.append(2)
feature_df.insert(3,'label_mapped',mapping)
feature_df.groupby('label_mapped')['audio'].count()
# +
cols=['video_id','start_time','mid_ts','label','audio','vggish']
d=np.load(f'./resources/working_data/vocal_only_data_with_vggish.npy',allow_pickle=True)
df = pd.DataFrame(d,columns=cols)
lut = pd.read_csv(f'../dataset/lookup.csv')
df=df.merge(lut[['video_id','band_name']],on='video_id')
feature_df=df[['label','audio','band_name','video_id']]
mapping=[]
for index,row in feature_df.iterrows():
if row['label'] == 'clean':
mapping.append(0)
if row['label'] == 'highfry':
mapping.append(1)
if row['label'] == 'layered':
mapping.append(1)
if row['label'] == 'lowfry':
mapping.append(1)
if row['label'] == 'midfry':
mapping.append(1)
if row['label'] == 'no_vocals':
mapping.append(2)
feature_df.insert(3,'label_mapped',mapping)
from imblearn.under_sampling import RandomUnderSampler
undersample = RandomUnderSampler(sampling_strategy=class_dist,random_state=seed)
X = feature_df[['audio','band_name','video_id']].to_numpy()
y=le.fit_transform(feature_df['label_mapped'].to_numpy())
X_under, y_under = undersample.fit_resample(X, y)
video_ids=X_under[:,2]
band_names = X_under[:,1]
# X_under=X_under[:,0]#.reshape(-1,1).flatten()
y_under=y_under
gss = GroupShuffleSplit(n_splits=5, train_size=.7, random_state=seed)
train,test = next(gss.split(X_under, y_under, band_names))
X_train = X_under[train]
X_test1 = X_under[test]
y_train = y_under[train]
y_test1 = y_under[test]
X_test,X_valid,y_test,y_valid = train_test_split(X_test1, y_test1,test_size=0.5,random_state=seed)
# train_video_ids=video_ids[train]
# test_video_ids=video_ids[test]
X_train = X_under[train]
X_test1 = X_under[test]
y_train = y_under[train]
y_test1 = y_under[test]
X_test,X_valid,y_test,y_valid = train_test_split(X_test1, y_test1,test_size=0.5,random_state=seed)
train_bands = X_train[:,1]
test_bands = X_test[:,1]
valid_bands = X_valid[:,1]
train_songs = X_train[:,2]
test_songs = X_test[:,2]
valid_songs = X_valid[:,2]
X_train = X_train[:,0]
X_test = X_test[:,0]
X_valid = X_valid[:,0]
d=pd.DataFrame()
d['y_train'] = y_train
d['blah'] = 1
print('TRAIN')
print(d.groupby('y_train')['blah'].count())
train = d['blah'].sum()
d=pd.DataFrame()
d['y_test'] = y_test
d['blah'] = 1
print('TEST')
print(d.groupby('y_test')['blah'].count())
test = d['blah'].sum()
d=pd.DataFrame()
d['y_valid'] = y_valid
d['blah'] = 1
print('VALID')
print(d.groupby('y_valid')['blah'].count())
valid = d['blah'].sum()
print(f"Train:Test:Validation - {train}:{test}:{valid}")
y_train_hot = to_categorical(y_train)
y_test_hot = to_categorical(y_test)
y_valid_hot = to_categorical(y_valid)
X_train,y_train=shuffle(X_train,y_train_hot, random_state=seed)
X_test,y_test=shuffle(X_test,y_test_hot, random_state=seed)
X_valid,y_valid=shuffle(X_valid,y_valid_hot, random_state=seed)
np.save(f'./resources/working_data/{n_class}_x_train-rawaudio.npy', X_train)
np.save(f'./resources/working_data/{n_class}_x_test-rawaudio.npy', X_test)
np.save(f'./resources/working_data/{n_class}_x_valid-rawaudio.npy', X_valid)
np.save(f'./resources/working_data/{n_class}_y_train-rawaudio.npy', y_train)
np.save(f'./resources/working_data/{n_class}_y_test-rawaudio.npy', y_test)
np.save(f'./resources/working_data/{n_class}_y_valid-rawaudio.npy', y_valid)
# -
np.intersect1d(test_songs,train_songs)
np.intersect1d(train_songs,valid_songs)
np.intersect1d(test_songs,valid_songs)
# # VGGish
# +
d=np.load(f'./resources/working_data/vocal_only_data_with_vggish.npy',allow_pickle=True)
feature_df=df[['label','vggish','band_name']]
mapping=[]
for index,row in feature_df.iterrows():
if row['label'] == 'clean':
mapping.append(0)
if row['label'] == 'highfry':
mapping.append(1)
if row['label'] == 'layered':
mapping.append(1)
if row['label'] == 'lowfry':
mapping.append(1)
if row['label'] == 'midfry':
mapping.append(1)
if row['label'] == 'no_vocals':
mapping.append(2)
feature_df.insert(3,'label_mapped',mapping)
from imblearn.under_sampling import RandomUnderSampler
undersample = RandomUnderSampler(sampling_strategy=class_dist,random_state=seed)
X = feature_df[['vggish','band_name']].to_numpy()
y=le.fit_transform(feature_df['label_mapped'].to_numpy())
X_under, y_under = undersample.fit_resample(X, y)
band_names = X_under[:,1]
X_under=X_under[:,0]#.reshape(-1,1).flatten()
X_under=np.concatenate(X_under).reshape(X_under.shape[0],128)
y_under=y_under
gss = GroupShuffleSplit(n_splits=5, train_size=.7, random_state=seed)
train,test = next(gss.split(X_under, y_under, band_names))
# for train_idx,test_idx in gss.split(X_under, y_under, band_names):
# print(train_idx,test_idx)
X_train = X_under[train]
X_test1 = X_under[test]
y_train = y_under[train]
y_test1 = y_under[test]
X_test,X_valid,y_test,y_valid = train_test_split(X_test1, y_test1,test_size=0.5,random_state=seed)
y_train_hot = to_categorical(y_train)
y_test_hot = to_categorical(y_test)
y_valid_hot = to_categorical(y_valid)
X_train,y_train=shuffle(X_train,y_train_hot, random_state=seed)
X_test,y_test=shuffle(X_test,y_test_hot, random_state=seed)
X_valid,y_valid=shuffle(X_valid,y_valid_hot, random_state=seed)
np.save(f'./resources/working_data/{n_class}_x_train-vggish.npy', X_train)
np.save(f'./resources/working_data/{n_class}_x_test-vggish.npy', X_test)
np.save(f'./resources/working_data/{n_class}_x_valid-vggish.npy', X_valid)
np.save(f'./resources/working_data/{n_class}_y_train-vggish.npy', y_train)
np.save(f'./resources/working_data/{n_class}_y_test-vggish.npy', y_test)
np.save(f'./resources/working_data/{n_class}_y_valid-vggish.npy', y_valid)
# -
# # Features
# +
cols=['video_id', 'start_time', 'mid_ts', 'label', 'average_zcr',
'zcr_stddev', 'mfcc1_mean', 'mfcc2_mean', 'mfcc3_mean',
'mfcc4_mean', 'mfcc5_mean', 'mfcc6_mean', 'mfcc7_mean', 'mfcc8_mean',
'mfcc9_mean', 'mfcc10_mean', 'mfcc11_mean', 'mfcc12_mean',
'mfcc13_mean', 'mfcc1_std', 'mfcc2_std', 'mfcc3_std', 'mfcc4_std',
'mfcc5_std', 'mfcc6_std', 'mfcc7_std', 'mfcc8_std', 'mfcc9_std',
'mfcc10_std', 'mfcc11_std', 'mfcc12_std', 'mfcc13_std',
'delta_mfcc1_mean', 'delta_mfcc2_mean', 'delta_mfcc3_mean',
'delta_mfcc4_mean', 'delta_mfcc5_mean', 'delta_mfcc6_mean',
'delta_mfcc7_mean', 'delta_mfcc8_mean', 'delta_mfcc9_mean',
'delta_mfcc10_mean', 'delta_mfcc11_mean', 'delta_mfcc12_mean',
'delta_mfcc13_mean', 'delta_mfcc1_std', 'delta_mfcc2_std',
'delta_mfcc3_std', 'delta_mfcc4_std', 'delta_mfcc5_std',
'delta_mfcc6_std', 'delta_mfcc7_std', 'delta_mfcc8_std',
'delta_mfcc9_std', 'delta_mfcc10_std', 'delta_mfcc11_std',
'delta_mfcc12_std', 'delta_mfcc13_std',
'centroid_mean','centroid_std',
'contrast_mean','contrast_std',
'flatness_mean','flatness_std',
'rolloff_mean','rolloff_std','rms_mean','rms_std','vggish']
d=np.load(f'./resources/working_data/vocal_only_features.npy',allow_pickle=True)
df = pd.DataFrame(d,columns=cols)
lut = pd.read_csv(f'../dataset/lookup.csv')
df=df.merge(lut[['video_id','band_name']],on='video_id')
df
feature_df=df[['label', 'band_name', 'average_zcr',
'zcr_stddev', 'mfcc1_mean', 'mfcc2_mean', 'mfcc3_mean',
'mfcc4_mean', 'mfcc5_mean', 'mfcc6_mean', 'mfcc7_mean', 'mfcc8_mean',
'mfcc9_mean', 'mfcc10_mean', 'mfcc11_mean', 'mfcc12_mean',
'mfcc13_mean', 'mfcc1_std', 'mfcc2_std', 'mfcc3_std', 'mfcc4_std',
'mfcc5_std', 'mfcc6_std', 'mfcc7_std', 'mfcc8_std', 'mfcc9_std',
'mfcc10_std', 'mfcc11_std', 'mfcc12_std', 'mfcc13_std',
'delta_mfcc1_mean', 'delta_mfcc2_mean', 'delta_mfcc3_mean',
'delta_mfcc4_mean', 'delta_mfcc5_mean', 'delta_mfcc6_mean',
'delta_mfcc7_mean', 'delta_mfcc8_mean', 'delta_mfcc9_mean',
'delta_mfcc10_mean', 'delta_mfcc11_mean', 'delta_mfcc12_mean',
'delta_mfcc13_mean', 'delta_mfcc1_std', 'delta_mfcc2_std',
'delta_mfcc3_std', 'delta_mfcc4_std', 'delta_mfcc5_std',
'delta_mfcc6_std', 'delta_mfcc7_std', 'delta_mfcc8_std',
'delta_mfcc9_std', 'delta_mfcc10_std', 'delta_mfcc11_std',
'delta_mfcc12_std', 'delta_mfcc13_std',
'centroid_mean','centroid_std',
'contrast_mean','contrast_std',
'flatness_mean','flatness_std',
'rolloff_mean','rolloff_std','rms_mean','rms_std']]
mapping=[]
for index,row in feature_df.iterrows():
if row['label'] == 'clean':
mapping.append(0)
if row['label'] == 'highfry':
mapping.append(1)
if row['label'] == 'layered':
mapping.append(1)
if row['label'] == 'lowfry':
mapping.append(1)
if row['label'] == 'midfry':
mapping.append(1)
if row['label'] == 'no_vocals':
mapping.append(2)
feature_df.insert(3,'label_mapped',mapping)
from imblearn.under_sampling import RandomUnderSampler
undersample = RandomUnderSampler(sampling_strategy=class_dist,random_state=seed)
X = feature_df[['average_zcr',
'zcr_stddev', 'mfcc1_mean', 'mfcc2_mean', 'mfcc3_mean',
'mfcc4_mean', 'mfcc5_mean', 'mfcc6_mean', 'mfcc7_mean', 'mfcc8_mean',
'mfcc9_mean', 'mfcc10_mean', 'mfcc11_mean', 'mfcc12_mean',
'mfcc13_mean', 'mfcc1_std', 'mfcc2_std', 'mfcc3_std', 'mfcc4_std',
'mfcc5_std', 'mfcc6_std', 'mfcc7_std', 'mfcc8_std', 'mfcc9_std',
'mfcc10_std', 'mfcc11_std', 'mfcc12_std', 'mfcc13_std',
'delta_mfcc1_mean', 'delta_mfcc2_mean', 'delta_mfcc3_mean',
'delta_mfcc4_mean', 'delta_mfcc5_mean', 'delta_mfcc6_mean',
'delta_mfcc7_mean', 'delta_mfcc8_mean', 'delta_mfcc9_mean',
'delta_mfcc10_mean', 'delta_mfcc11_mean', 'delta_mfcc12_mean',
'delta_mfcc13_mean', 'delta_mfcc1_std', 'delta_mfcc2_std',
'delta_mfcc3_std', 'delta_mfcc4_std', 'delta_mfcc5_std',
'delta_mfcc6_std', 'delta_mfcc7_std', 'delta_mfcc8_std',
'delta_mfcc9_std', 'delta_mfcc10_std', 'delta_mfcc11_std',
'delta_mfcc12_std', 'delta_mfcc13_std',
'centroid_mean','centroid_std',
'contrast_mean','contrast_std',
'flatness_mean','flatness_std',
'rolloff_mean','rolloff_std','rms_mean','rms_std','band_name']].to_numpy()
y=le.fit_transform(feature_df['label_mapped'].to_numpy())
X_under, y_under = undersample.fit_resample(X, y)
band_names = X_under[:,-1]
X_under=X_under[:,:-1]#.reshape(-1,1).flatten()
y_under=y_under
gss = GroupShuffleSplit(n_splits=5, train_size=.7, random_state=seed)
train,test = next(gss.split(X_under, y_under, band_names))
# for train_idx,test_idx in gss.split(X_under, y_under, band_names):
# print(train_idx,test_idx)
X_train = X_under[train]
X_test1 = X_under[test]
y_train = y_under[train]
y_test1 = y_under[test]
X_test,X_valid,y_test,y_valid = train_test_split(X_test1, y_test1,test_size=0.5,random_state=seed)
y_train_hot = to_categorical(y_train)
y_test_hot = to_categorical(y_test)
y_valid_hot = to_categorical(y_valid)
X_train,y_train=shuffle(X_train,y_train_hot, random_state=seed)
X_test,y_test=shuffle(X_test,y_test_hot, random_state=seed)
X_valid,y_valid=shuffle(X_valid,y_valid_hot, random_state=seed)
np.save(f'./resources/working_data/{n_class}_x_train-features_unnormalized.npy', X_train)
np.save(f'./resources/working_data/{n_class}_x_test-features_unnormalized.npy', X_test)
np.save(f'./resources/working_data/{n_class}_x_valid-features_unnormalized.npy', X_valid)
np.save(f'./resources/working_data/{n_class}_y_train-features_unnormalized.npy', y_train)
np.save(f'./resources/working_data/{n_class}_y_test-features_unnormalized.npy', y_test)
np.save(f'./resources/working_data/{n_class}_y_valid-features_unnormalized.npy', y_valid)
# -
# # MFCCs only
# +
cols=['video_id', 'start_time', 'mid_ts', 'label', 'average_zcr',
'zcr_stddev', 'mfcc1_mean', 'mfcc2_mean', 'mfcc3_mean',
'mfcc4_mean', 'mfcc5_mean', 'mfcc6_mean', 'mfcc7_mean', 'mfcc8_mean',
'mfcc9_mean', 'mfcc10_mean', 'mfcc11_mean', 'mfcc12_mean',
'mfcc13_mean', 'mfcc1_std', 'mfcc2_std', 'mfcc3_std', 'mfcc4_std',
'mfcc5_std', 'mfcc6_std', 'mfcc7_std', 'mfcc8_std', 'mfcc9_std',
'mfcc10_std', 'mfcc11_std', 'mfcc12_std', 'mfcc13_std',
'delta_mfcc1_mean', 'delta_mfcc2_mean', 'delta_mfcc3_mean',
'delta_mfcc4_mean', 'delta_mfcc5_mean', 'delta_mfcc6_mean',
'delta_mfcc7_mean', 'delta_mfcc8_mean', 'delta_mfcc9_mean',
'delta_mfcc10_mean', 'delta_mfcc11_mean', 'delta_mfcc12_mean',
'delta_mfcc13_mean', 'delta_mfcc1_std', 'delta_mfcc2_std',
'delta_mfcc3_std', 'delta_mfcc4_std', 'delta_mfcc5_std',
'delta_mfcc6_std', 'delta_mfcc7_std', 'delta_mfcc8_std',
'delta_mfcc9_std', 'delta_mfcc10_std', 'delta_mfcc11_std',
'delta_mfcc12_std', 'delta_mfcc13_std',
'centroid_mean','centroid_std',
'contrast_mean','contrast_std',
'flatness_mean','flatness_std',
'rolloff_mean','rolloff_std','rms_mean','rms_std','vggish']
d=np.load(f'./resources/working_data/vocal_only_features.npy',allow_pickle=True)
df = pd.DataFrame(d,columns=cols)
lut = pd.read_csv(f'../dataset/lookup.csv')
df=df.merge(lut[['video_id','band_name']],on='video_id')
df
feature_df=df[['label', 'band_name', 'mfcc1_mean', 'mfcc2_mean', 'mfcc3_mean',
'mfcc4_mean', 'mfcc5_mean', 'mfcc6_mean', 'mfcc7_mean', 'mfcc8_mean',
'mfcc9_mean', 'mfcc10_mean', 'mfcc11_mean', 'mfcc12_mean',
'mfcc13_mean', 'mfcc1_std', 'mfcc2_std', 'mfcc3_std', 'mfcc4_std',
'mfcc5_std', 'mfcc6_std', 'mfcc7_std', 'mfcc8_std', 'mfcc9_std',
'mfcc10_std', 'mfcc11_std', 'mfcc12_std', 'mfcc13_std',
'delta_mfcc1_mean', 'delta_mfcc2_mean', 'delta_mfcc3_mean',
'delta_mfcc4_mean', 'delta_mfcc5_mean', 'delta_mfcc6_mean',
'delta_mfcc7_mean', 'delta_mfcc8_mean', 'delta_mfcc9_mean',
'delta_mfcc10_mean', 'delta_mfcc11_mean', 'delta_mfcc12_mean',
'delta_mfcc13_mean', 'delta_mfcc1_std', 'delta_mfcc2_std',
'delta_mfcc3_std', 'delta_mfcc4_std', 'delta_mfcc5_std',
'delta_mfcc6_std', 'delta_mfcc7_std', 'delta_mfcc8_std',
'delta_mfcc9_std', 'delta_mfcc10_std', 'delta_mfcc11_std',
'delta_mfcc12_std', 'delta_mfcc13_std']]
mapping=[]
for index,row in feature_df.iterrows():
if row['label'] == 'clean':
mapping.append(0)
if row['label'] == 'highfry':
mapping.append(1)
if row['label'] == 'layered':
mapping.append(1)
if row['label'] == 'lowfry':
mapping.append(1)
if row['label'] == 'midfry':
mapping.append(1)
if row['label'] == 'no_vocals':
mapping.append(2)
feature_df.insert(3,'label_mapped',mapping)
from imblearn.under_sampling import RandomUnderSampler
undersample = RandomUnderSampler(sampling_strategy=class_dist,random_state=seed)
X = feature_df[['mfcc1_mean', 'mfcc2_mean', 'mfcc3_mean',
'mfcc4_mean', 'mfcc5_mean', 'mfcc6_mean', 'mfcc7_mean', 'mfcc8_mean',
'mfcc9_mean', 'mfcc10_mean', 'mfcc11_mean', 'mfcc12_mean',
'mfcc13_mean', 'mfcc1_std', 'mfcc2_std', 'mfcc3_std', 'mfcc4_std',
'mfcc5_std', 'mfcc6_std', 'mfcc7_std', 'mfcc8_std', 'mfcc9_std',
'mfcc10_std', 'mfcc11_std', 'mfcc12_std', 'mfcc13_std',
'delta_mfcc1_mean', 'delta_mfcc2_mean', 'delta_mfcc3_mean',
'delta_mfcc4_mean', 'delta_mfcc5_mean', 'delta_mfcc6_mean',
'delta_mfcc7_mean', 'delta_mfcc8_mean', 'delta_mfcc9_mean',
'delta_mfcc10_mean', 'delta_mfcc11_mean', 'delta_mfcc12_mean',
'delta_mfcc13_mean', 'delta_mfcc1_std', 'delta_mfcc2_std',
'delta_mfcc3_std', 'delta_mfcc4_std', 'delta_mfcc5_std',
'delta_mfcc6_std', 'delta_mfcc7_std', 'delta_mfcc8_std',
'delta_mfcc9_std', 'delta_mfcc10_std', 'delta_mfcc11_std',
'delta_mfcc12_std', 'delta_mfcc13_std','band_name']].to_numpy()
y=le.fit_transform(feature_df['label_mapped'].to_numpy())
X_under, y_under = undersample.fit_resample(X, y)
band_names = X_under[:,-1]
X_under=X_under[:,:-1]#.reshape(-1,1).flatten()
y_under=y_under
gss = GroupShuffleSplit(n_splits=5, train_size=.7, random_state=seed)
train,test = next(gss.split(X_under, y_under, band_names))
# for train_idx,test_idx in gss.split(X_under, y_under, band_names):
# print(train_idx,test_idx)
X_train = X_under[train]
X_test1 = X_under[test]
y_train = y_under[train]
y_test1 = y_under[test]
X_test,X_valid,y_test,y_valid = train_test_split(X_test1, y_test1,test_size=0.5,random_state=seed)
y_train_hot = to_categorical(y_train)
y_test_hot = to_categorical(y_test)
y_valid_hot = to_categorical(y_valid)
X_train,y_train=shuffle(X_train,y_train_hot, random_state=seed)
X_test,y_test=shuffle(X_test,y_test_hot, random_state=seed)
X_valid,y_valid=shuffle(X_valid,y_valid_hot, random_state=seed)
np.save(f'./resources/working_data/{n_class}_x_train-mfcc_only_unnormalized.npy', X_train)
np.save(f'./resources/working_data/{n_class}_x_test-mfcc_only_unnormalized.npy', X_test)
np.save(f'./resources/working_data/{n_class}_x_valid-mfcc_only_unnormalized.npy', X_valid)
np.save(f'./resources/working_data/{n_class}_y_train-mfcc_only_unnormalized.npy', y_train)
np.save(f'./resources/working_data/{n_class}_y_test-mfcc_only_unnormalized.npy', y_test)
np.save(f'./resources/working_data/{n_class}_y_valid-mfcc_only_unnormalized.npy', y_valid)
# -
# # Features Only
# +
cols=['video_id', 'start_time', 'mid_ts', 'label', 'average_zcr',
'zcr_stddev', 'mfcc1_mean', 'mfcc2_mean', 'mfcc3_mean',
'mfcc4_mean', 'mfcc5_mean', 'mfcc6_mean', 'mfcc7_mean', 'mfcc8_mean',
'mfcc9_mean', 'mfcc10_mean', 'mfcc11_mean', 'mfcc12_mean',
'mfcc13_mean', 'mfcc1_std', 'mfcc2_std', 'mfcc3_std', 'mfcc4_std',
'mfcc5_std', 'mfcc6_std', 'mfcc7_std', 'mfcc8_std', 'mfcc9_std',
'mfcc10_std', 'mfcc11_std', 'mfcc12_std', 'mfcc13_std',
'delta_mfcc1_mean', 'delta_mfcc2_mean', 'delta_mfcc3_mean',
'delta_mfcc4_mean', 'delta_mfcc5_mean', 'delta_mfcc6_mean',
'delta_mfcc7_mean', 'delta_mfcc8_mean', 'delta_mfcc9_mean',
'delta_mfcc10_mean', 'delta_mfcc11_mean', 'delta_mfcc12_mean',
'delta_mfcc13_mean', 'delta_mfcc1_std', 'delta_mfcc2_std',
'delta_mfcc3_std', 'delta_mfcc4_std', 'delta_mfcc5_std',
'delta_mfcc6_std', 'delta_mfcc7_std', 'delta_mfcc8_std',
'delta_mfcc9_std', 'delta_mfcc10_std', 'delta_mfcc11_std',
'delta_mfcc12_std', 'delta_mfcc13_std',
'centroid_mean','centroid_std',
'contrast_mean','contrast_std',
'flatness_mean','flatness_std',
'rolloff_mean','rolloff_std','rms_mean','rms_std','vggish']
d=np.load(f'./resources/working_data/vocal_only_features.npy',allow_pickle=True)
df = pd.DataFrame(d,columns=cols)
lut = pd.read_csv(f'../dataset/lookup_new.csv')
df=df.merge(lut[['video_id','band_name']],on='video_id')
df
feature_df=df[['label', 'band_name', 'average_zcr',
'zcr_stddev','centroid_mean','centroid_std',
'contrast_mean','contrast_std',
'flatness_mean','flatness_std',
'rolloff_mean','rolloff_std','rms_mean','rms_std']]
mapping=[]
for index,row in feature_df.iterrows():
if row['label'] == 'clean':
mapping.append(0)
if row['label'] == 'highfry':
mapping.append(1)
if row['label'] == 'layered':
mapping.append(1)
if row['label'] == 'lowfry':
mapping.append(1)
if row['label'] == 'midfry':
mapping.append(1)
if row['label'] == 'no_vocals':
mapping.append(2)
feature_df.insert(3,'label_mapped',mapping)
from imblearn.under_sampling import RandomUnderSampler
undersample = RandomUnderSampler(sampling_strategy=class_dist,random_state=seed)
X = feature_df[['average_zcr',
'zcr_stddev','centroid_mean','centroid_std',
'contrast_mean','contrast_std',
'flatness_mean','flatness_std',
'rolloff_mean','rolloff_std','rms_mean','rms_std','band_name']].to_numpy()
y=le.fit_transform(feature_df['label_mapped'].to_numpy())
X_under, y_under = undersample.fit_resample(X, y)
band_names = X_under[:,-1]
X_under=X_under[:,:-1]#.reshape(-1,1).flatten()
y_under=y_under
gss = GroupShuffleSplit(n_splits=5, train_size=.7, random_state=seed)
train,test = next(gss.split(X_under, y_under, band_names))
# for train_idx,test_idx in gss.split(X_under, y_under, band_names):
# print(train_idx,test_idx)
X_train = X_under[train]
X_test1 = X_under[test]
y_train = y_under[train]
y_test1 = y_under[test]
X_test,X_valid,y_test,y_valid = train_test_split(X_test1, y_test1,test_size=0.5,random_state=seed)
y_train_hot = to_categorical(y_train)
y_test_hot = to_categorical(y_test)
y_valid_hot = to_categorical(y_valid)
X_train,y_train=shuffle(X_train,y_train_hot, random_state=seed)
X_test,y_test=shuffle(X_test,y_test_hot, random_state=seed)
X_valid,y_valid=shuffle(X_valid,y_valid_hot, random_state=seed)
np.save(f'./resources/working_data/{n_class}_x_train-features_only_unnormalized.npy', X_train)
np.save(f'./resources/working_data/{n_class}_x_test-features_only_unnormalized.npy', X_test)
np.save(f'./resources/working_data/{n_class}_x_valid-features_only_unnormalized.npy', X_valid)
np.save(f'./resources/working_data/{n_class}_y_train-features_only_unnormalized.npy', y_train)
np.save(f'./resources/working_data/{n_class}_y_test-features_only_unnormalized.npy', y_test)
np.save(f'./resources/working_data/{n_class}_y_valid-features_only_unnormalized.npy', y_valid)
# -
| benchmark-results/3class_results/train_test_split_3class.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Notebook Examples for Chapter 1
import warnings
# these are innocuous but irritating
warnings.filterwarnings("ignore", message="numpy.dtype size changed")
warnings.filterwarnings("ignore", message="numpy.ufunc size changed")
# %matplotlib inline
# +
import IPython.display as disp
import ee
ee.Initialize()
minlon = 6.31
minlat = 50.83
maxlon = 6.58
maxlat = 50.95
rect = ee.Geometry.Rectangle([minlon,minlat,maxlon,maxlat])
collection = ee.ImageCollection('COPERNICUS/S1_GRD') \
.filterBounds(rect) \
.filterDate(ee.Date('2017-05-01'), ee.Date('2017-06-01')) \
.filter(ee.Filter.eq('transmitterReceiverPolarisation', ['VV','VH'])) \
.filter(ee.Filter.eq('resolution_meters', 10)) \
.filter(ee.Filter.eq('instrumentMode', 'IW'))
image = ee.Image(collection.first()).clip(rect)
url = image.select('VV').getThumbURL({'min':-20,'max':0})
disp.Image(url=url)
# -
run scripts/ex1_1 imagery/AST_20070501 3
# ### Covariance matrix of a multispectral image
# +
from osgeo import gdal
from osgeo.gdalconst import GA_ReadOnly
gdal.AllRegister()
infile = 'imagery/AST_20070501'
inDataset = gdal.Open(infile,GA_ReadOnly)
cols = inDataset.RasterXSize
rows = inDataset.RasterYSize
# data matrix
G = np.zeros((rows*cols,3))
for b in range(3):
band = inDataset.GetRasterBand(b+1)
tmp = band.ReadAsArray(0,0,cols,rows).ravel()
G[:,b] = tmp - np.mean(tmp)
# covariance matrix
C = np.mat(G).T*np.mat(G)/(cols*rows-1)
print C
# -
# ### Eigenvalues and eigenvectors of the covariance matrix
eigenvalues, eigenvectors = np.linalg.eigh(C)
print eigenvalues
print eigenvectors
U = eigenvectors
print U.T*U
# ### Singular value decomposition
import numpy as np
b = np.mat([1,2,3])
# an almost singular matrix
A = b.T*b + np.random.rand(3,3)*0.001
# a symmetric almost singular matrix
A = A + A.T
print 'determinant: %f'%np.linalg.det(A)
# singular value decomposition
U,Lambda,V = np.linalg.svd(A)
print 'Lambda = %s'%str(Lambda)
print 'U = %s'%str(U)
print 'V = %s'%str(V)
# ### Principal components analysis
run scripts/ex1_2 imagery/AST_20070501 imagery/pca.tif
run scripts/dispms -f 'imagery/pca.tif' -p [1,2,3] -e 4
| src/.ipynb_checkpoints/Chapter1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # K-Means Example
#
# Implement K-Means algorithm with TensorFlow, and apply it to classify
# handwritten digit images. This example is using the MNIST database of
# handwritten digits as training samples (http://yann.lecun.com/exdb/mnist/).
#
#
#
#
# +
from __future__ import print_function
import numpy as np
import tensorflow as tf
from tensorflow.contrib.factorization import KMeans
# Ignore all GPUs, tf random forest does not benefit from it.
import os
os.environ["CUDA_VISIBLE_DEVICES"] = ""
# -
# Import MNIST data
from tensorflow.examples.tutorials.mnist import input_data
mnist = input_data.read_data_sets("/tmp/data/", one_hot=True)
full_data_x = mnist.train.images
# +
# Parameters
num_steps = 50 # Total steps to train
batch_size = 1024 # The number of samples per batch
k = 25 # The number of clusters
num_classes = 10 # The 10 digits
num_features = 784 # Each image is 28x28 pixels
# Input images
X = tf.placeholder(tf.float32, shape=[None, num_features])
# Labels (for assigning a label to a centroid and testing)
Y = tf.placeholder(tf.float32, shape=[None, num_classes])
# K-Means Parameters
kmeans = KMeans(inputs=X, num_clusters=k, distance_metric='cosine',
use_mini_batch=True)
# +
# Build KMeans graph
(all_scores, cluster_idx, scores, cluster_centers_initialized,
cluster_centers_vars,init_op,train_op) = kmeans.training_graph()
cluster_idx = cluster_idx[0] # fix for cluster_idx being a tuple
avg_distance = tf.reduce_mean(scores)
# Initialize the variables (i.e. assign their default value)
init_vars = tf.global_variables_initializer()
# +
# Start TensorFlow session
sess = tf.Session()
# Run the initializer
sess.run(init_vars, feed_dict={X: full_data_x})
sess.run(init_op, feed_dict={X: full_data_x})
# Training
for i in range(1, num_steps + 1):
_, d, idx = sess.run([train_op, avg_distance, cluster_idx],
feed_dict={X: full_data_x})
if i % 10 == 0 or i == 1:
print("Step %i, Avg Distance: %f" % (i, d))
# +
# Assign a label to each centroid
# Count total number of labels per centroid, using the label of each training
# sample to their closest centroid (given by 'idx')
counts = np.zeros(shape=(k, num_classes))
for i in range(len(idx)):
counts[idx[i]] += mnist.train.labels[i]
# Assign the most frequent label to the centroid
labels_map = [np.argmax(c) for c in counts]
labels_map = tf.convert_to_tensor(labels_map)
# Evaluation ops
# Lookup: centroid_id -> label
cluster_label = tf.nn.embedding_lookup(labels_map, cluster_idx)
# Compute accuracy
correct_prediction = tf.equal(cluster_label, tf.cast(tf.argmax(Y, 1), tf.int32))
accuracy_op = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))
# Test Model
test_x, test_y = mnist.test.images, mnist.test.labels
print("Test Accuracy:", sess.run(accuracy_op, feed_dict={X: test_x, Y: test_y}))
| kmeans.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: kaggle_microsoft-malware-prediction
# kernelspec:
# display_name: Kaggle_Microsoft-Malware-Prediction
# language: python
# name: kaggle_microsoft-malware-prediction
# ---
# + outputHidden=false inputHidden=false
import pandas
from sklearn.decomposition import PCA
from sklearn import preprocessing
# + outputHidden=false inputHidden=false
sample_data = pandas.read_csv("../data/train.csv",nrows=2000)
# + outputHidden=false inputHidden=false
sample_data
# + outputHidden=false inputHidden=false
pca = PCA().fit(sample_data)
plt.plot(np.cumsum(pca.explained_variance_ratio_))
plt.xlabel('number of components')
plt.ylabel('cumulative explained variance');
| notebooks/exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + _uuid="40fff571d215af00b45ec5faf36c8c77bf08e423"
import numpy as np
import pandas
import os
from urllib.request import urlretrieve
import zipfile
# + _uuid="543e41f967b6cd873f4ef10b0d21af2b53b55652"
import os
import csv
import gzip
import collections
import re
import io
import json
import xml.etree.ElementTree as ET
import untangle
import requests
import pandas as pd
# + _uuid="a7f2a49ab9c59fae771266bc063e1d483844f942"
zip_ref = zipfile.ZipFile(filename, 'r')
zip_ref.extractall()
zip_ref.close()
# -
filename="full database.xml"
obj=untangle.parse(filename)
df_drugbank_sm=pd.DataFrame(columns=["drugbank_id","name","cas","smiles","logP ALOGPS", "logP ChemAxon", "solubility ALOGPS", "pKa (strongest acidic)", "pKa (strongest basic)"])
df_drugbank_sm
i=-1
#iterate over drug entries to extract information
for drug in obj.drugbank.drug:
drug_type= str(drug["type"])
# select for small molecule drugs
if drug_type in ["small molecule", "Small Molecule", "Small molecule"]:
i=i+1
#Get drugbank_id
for id in drug.drugbank_id:
if str(id["primary"])=="true":
df_drugbank_sm.loc[i, "drugbank_id"]=id.cdata
#Drug name
df_drugbank_sm.loc[i,"name"]=drug.name.cdata
#Drug CAS
df_drugbank_sm.loc[i, "cas"]=drug.cas_number.cdata
#Get SMILES, logP, Solubility
#Skip drugs with no structure. ("DB00386","DB00407","DB00702","DB00785","DB00840",
# "DB00893","DB00930","DB00965", "DB01109","DB01266",
# "DB01323", "DB01341"...)
if len(drug.calculated_properties.cdata)==0: #If there is no calculated properties
continue
else:
for property in drug.calculated_properties.property:
if property.kind.cdata == "SMILES":
df_drugbank_sm.loc[i, "smiles"]=property.value.cdata
if property.kind.cdata == "logP":
if property.source.cdata == "ALOGPS":
df_drugbank_sm.loc[i, "logP ALOGPS"]=property.value.cdata
if property.source.cdata == "ChemAxon":
df_drugbank_sm.loc[i, "logP ChemAxon"]=property.value.cdata
if property.kind.cdata == "Water Solubility":
df_drugbank_sm.loc[i, "solubility ALOGPS"]=property.value.cdata
if property.kind.cdata == "pKa (strongest acidic)":
df_drugbank_sm.loc[i, "pKa (strongest acidic)"]=property.value.cdata
if property.kind.cdata == "pKa (strongest basic)":
df_drugbank_sm.loc[i, "pKa (strongest basic)"]=property.value.cdata
df_drugbank_sm.head()
#Drop drugs without SMILES from the dataframe
df_drugbank_smiles = df_drugbank_sm.dropna()
df_drugbank_smiles= df_drugbank_smiles.reset_index(drop=True)
print(df_drugbank_smiles.shape)
df_drugbank_smiles.to_csv('drugbank_smiles_df.csv',index=False)
| Scripts/choderalab-drugbank.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import numpy as np
print(np.__version__)
import matplotlib.pyplot as plt
# %run 4.3.2-common.ipynb
# +
# Let us now try a similar experiment with
# uncorrelated data
N = 100
x_0 = np.random.normal(0, 100, N)
x_1 = np.random.normal(0, 100, N)
X = np.column_stack((x_0, x_1))
# Perform PCA
# Note that principal values are close to
# each other - the spread is comparable in both
# directions.
principal_values, principal_components = pca(X)
print("Principal values are {}".format(principal_values))
# Find the index with highest principal value
major_index = np.argmax(principal_values)
minor_index = np.argmin(principal_values)
# Plot
plt.figure()
plt.scatter(X[:, 0], X[: , 1],
color="green")
plt.title('Uncorrelated dataset')
plt.xlabel('x0')
plt.ylabel('x1')
# Let us plot the principal components
draw_line(principal_components[:, major_index],
min_x=-200, max_x=200)
draw_line(principal_components[:, minor_index],
min_x=-150, max_x=150, color="red")
plt.show()
first_principal_vec = principal_components[:,
major_index]
first_principal_vec = first_principal_vec.reshape((-1,
1))
X_proj = np.dot(X, first_principal_vec)
# Information lost due to dimensionality reduction.
# (here we've lost real info)
X_back_proj = np.dot(X_proj,
np.linalg.pinv(first_principal_vec))
info_loss = np.sqrt(np.mean((X_back_proj - X)**2))
print("Loss in Information due to\ndimensionality"
" reduction: {}\n(high - we're "
"losing real info)".format(info_loss))
| python/ch4/4.3.2-pca-uncorrelated-numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # The Convex Hull Problem
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Pound a bunch of nails into a board, then stretch a rubber band around them and let the rubber band snap taut, like this:
#
# <img src="http://www.personal.kent.edu/~rmuhamma/Compgeometry/MyCG/Gifs-CompGeometry/ch2.gif">
#
# The rubber band has traced out the *convex hull* of the set of nails. It turns out this is an important problem with applications in computer graphics, robot motion planning, geographical information systems, ethology, and other areas.
# More formally, we say that:
#
# *Given a finite set, **P**, of points in a plane, the convex hull of **P** is a polygon, **H**, such that:*
#
# - *Every point in **P** lies either on or inside of **H**.*
# - *Every vertex of **H** is a point in **P**.*
# - **H** *is convex: a line segment joining any two vertexes of **H** either is an edge of **H** or lies inside **H**.*
#
#
# In this notebook we develop an algorithm to find the convex hull (and show examples of how to use `matplotlib` plotting). The first thing to do is decide how we will represent the objects of interest:
#
# - **Point**: We'll define a class such that `Point(3, 4)` is a point where `p.x` is 3 and `p.y` is 4.
# - **Set of Points**: We'll use a Python set: `{Point(0,0), Point(3,4), ...}`
# - **Polygon**: We'll represent a polygon as an ordered list of vertex points.
#
# First, get the necessary imports done:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
from __future__ import division, print_function
# %matplotlib inline
import matplotlib.pyplot as plt
import collections
import random
import math
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # Points and Sets of Points
#
# I'll define the class `Point` as a named tuple of `x` and `y` coordinates, and `Points(n)` as a function that creates a set of *n* random points.
#
# There are two complications to the function `Points(n)`:
# 1. A second optional argument is used to set the random seed. This way, the same call to `Points` will return the same result each time. That makes it easier to reproduce tests. If you want different sets of points, just pass in different values for the seed.
# 2. Since `matplotlib` plots on a 3×2 rectangle by default, the points will be uniformly sampled from a 3×2 box (with a small border of 0.05 on each edge to prevent the points from bumping up against the edge of the box).
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
Point = collections.namedtuple('Point', 'x, y')
def Points(n, seed=42):
"Generate n random points within a 3 x 2 box."
random.seed((n, seed))
b = 0.05 # border
return {Point(random.uniform(b, 3-b), random.uniform(b, 2-b))
for _ in range(n)}
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
Points(3)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # Visualizing Points and Line Segments
#
#
# Now let's see how to visualize points; I'll define a function `plot_points`. We will want to be able to see:
# - The **points** themselves.
# - Optionally, **line segments** between points. An optional `style` parameter allows you to specify whether you want lines or not, and what color they should be. This parameter uses the standard [style format](http://matplotlib.org/1.3.1/api/pyplot_api.html#matplotlib.pyplot.plot) defined by matplotlib; for example, `'r.'` means red colored dots with no lines, `'bs-'` means blue colored squares with lines between them, and `'go:'` means green colored circles with dotted lines between them. The lines go from point to point in order; if you want the lines to close
# back from the last point to the first (to form a complete polygon), specify `closed=True`. (For that to work,
# the collection of points must be a list; with `closed=False` the collection can be any collection.)
# - Optionally, **labels** on the points that let us distinguish one from another. You get
# labels (integers from 0 to *n*) if you specify `labels=True`.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
def plot_points(points, style='r.', labels=False, closed=False):
"""Plot a collection of points. Optionally change the line style, label points with numbers,
and/or form a closed polygon by closing the line from the last point to the first."""
if labels:
for (i, (x, y)) in enumerate(points):
plt.text(x, y, ' '+str(i))
if closed:
points = points + [points[0]]
plt.plot([p.x for p in points], [p.y for p in points], style, linewidth=2.5)
plt.axis('scaled'); plt.axis('off')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Here's an example:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_points(Points(200))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # Convexity
#
#
# We want to make a *convex* hull, so we better have some way of determining whether a polygon is *convex*. Let's examine one that is:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
octagon = [Point(-10, 0), Point(-7, -7), Point(0, -10), Point(+7, -7),
Point(+10, 0), Point(+7, +7), Point(0, +10), Point(-7, 7)]
plot_points(octagon, 'bs-', labels=True, closed=True)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# If you start at point 0 at the left and proceed in order counterclockwise around the octagon, following edges from point to point, you can see that at every vertex you are making a **left** turn.
#
# Now let's consider a non-convex polygon:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
pacman = octagon[:4] + [Point(0, 0)] + octagon[5:]
plot_points(pacman, 'ys-', labels=True, closed=True)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# The `pacman` polygon is non-convex; you can see that a line from point 3 to point 5 passes *outside* the polygon. You can also see that as you move counterclockwise from 3 to 4 to 5 you turn **right** at 4. That leads to the idea: **a polygon is convex if there are no right turns** as we go around the polygon counterclockwise.
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # Turn Directions
#
#
# Now how do we determine if a turn from point A to B to C is a left turn at B or a right turn (or straight)? Consider this diagram:
#
# <img src="http://norvig.com/convexhull.jpg">
#
# It is a left turn at B if angle β is bigger than angle α; in other words, if β's opposite-over-adjacent ratio is bigger than α's:
#
# (C.y - B.y) / (C.x - B.x) > (B.y - A.y) / (B.x - A.x)
#
# But if we did that computation, we'd need special cases for when each denominator is zero. So multiply each side by the denominators:
#
# (B.x - A.x) * (C.y - B.y) > (B.y - A.y) * (C.x - B.x)
#
# (*Note:* This step should make you very nervous! In general, multiplying both sides of an inequality by a negative number reverses the inequality, and here the denominators might be negative. In this case it works out; basically because we are doing two multiplications so that negatives cancel out, but [the math proof](https://en.wikipedia.org/wiki/Cross_product) is tricky, involving some concepts in vector algebra, so I won't duplicate it here; instead I will provide good test coverage below.)
#
# That leads to the function definition:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
def turn(A, B, C):
"Is the turn from A->B->C a 'right', 'left', or 'straight' turn?"
diff = (B.x - A.x) * (C.y - B.y) - (B.y - A.y) * (C.x - B.x)
return ('right' if diff < 0 else
'left' if diff > 0 else
'straight')
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # Sketch of Convex Hull Algorithm
#
#
# Now we have the first part of a strategy to find the convex hull:
#
# > *Travel a path along the points in some order. (It is not yet clear exactly what order.) Any point along the way that does not mark a left-hand turn is not part of the hull.*
#
# What's a good order? Let's see what happens if we start at the leftmost point and work our way to the rightmost. We can achieve that ordering by calling the built-in function `sorted` on the points (since points are tuples, `sorted` sorts them lexicographically: first by their first component, `x`, and if there are ties, next by their `y` component). We start with 11 random points, and I will define a function to help plot the partial hull as we go:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
def plot_partial_hull(points, hull_indexes=()):
"Plot the points, labeled, with a blue line for the points named by indexes."
plot_points(points, labels=True)
plot_points([points[i] for i in hull_indexes], 'bs-')
# + [markdown] run_control={}
# Here are the points without any hull:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Now I will start building up the hull by following the points in order from point 0 to 1 to 2 to 3:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [0, 1, 2, 3])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# We see that we made a valid left turn at point 1, but a right turn at 2. So we remove point 2 from the hull:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [0, 1, 3])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# We move on to points 4 and 5:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [0, 1, 3, 4, 5])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Point 4 is a right turn, so we remove it:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [0, 1, 3, 5])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# But now we see point 3 is also a right turn. The addition of one new point (5) can remove multiple points (4 and 3) from the hull. We remove 3 and move on to 6, 7, and 8:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [0, 1, 5, 6, 7, 8])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Point 7 is a right turn so we remove 7 and move on to 9:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [0, 1, 5, 6, 8, 9])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Point 8 is a right turn, so we remove 8. But then 6 and 5 are also right turns, so they too are removed. We proceed on to 10:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [0, 1, 9, 10])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Now what do we do? We got all the way to the end of our set of 11 points, but we only got half the hull (the lower half). Well, if looking at all the points in left-to-right order gives us the lower half of the hull, maybe looking at all the points in right-to-left order will give us the upper half. Let's try.
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [10, 9, 8])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Point 9 is a right turn; remove it and move on:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [10, 8, 7, 6, 5])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Adding 5 reveals 6, and then 7, to be right turns; remove them and move on to 4:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [10, 8, 5, 4])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Remove 5 and continue on to 3 and then 2:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [10, 8, 4, 3, 2])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Now 3 is a right turn; remove it and continue on to 1 and finally 0:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [10, 8, 4, 2, 1 ,0])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Adding 0 makes, 1, and then 2 be right turns, so they are removed:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [10, 8, 4, 0])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Let's bring back the lower hull and concatenate it with the upper hull:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_partial_hull(sorted(Points(11)), [0, 1, 9, 10] + [10, 8, 4, 0])
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# That's all there is to the basic idea of the algorithm, but there are a few edge cases to worry about:
#
# * **Degenerate polygons**: What happens when there are only 1 or 2 (or zero) points? Such a set of points should be considered convex because there is no way to draw a line segment that goes outside the points.
#
# * **Colinear points:** if three or more points are colinear, we should keep only the two "outside" ones. The rationale for not keeping them all is that we want the convex hull to be the minimal possible set of points. We need to keep the outside ones because they mark true corners in the hull. We can achieve this by rejecting a point when it is a "straight" turn as well as when it is a "right" turn.
#
# * **First and last points:** An astute reader might have noticed that our algorithm only rejects the middle point, point B, in the A->B->C turn. That means that the first and last point in sorted order will never be a candidate for rejection, and thus will always end up on the hull. Is that correct? Yes it is. The first point is the leftmost point, the one with lowest `x` value (and if there are ties, it is the lowest-leftmost point). That is an extreme corner, so it should always be on the hull. A similar argument holds for the last point in sorted order.
#
#
#
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # Implementation of Convex Hull Algorithm
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
def convex_hull(points):
"Find the convex hull of a set of points."
if len(points) <= 3:
return points
# Find the two half-hulls and append them, but don't repeat first and last points
upper = half_hull(sorted(points))
lower = half_hull(reversed(sorted(points)))
return upper + lower[1:-1]
def half_hull(sorted_points):
"Return the half-hull from following points in sorted order."
# Add each point C in order; remove previous point B if A->B-C is not a left turn.
hull = []
for C in sorted_points:
# if A->B->C is not a left turn ...
while len(hull) >= 2 and turn(hull[-2], hull[-1], C) != 'left':
hull.pop() # ... then remove B from hull.
hull.append(C)
return hull
# -
# We can try it out on our 11 random points, but it is not easy to tell at a glance whether the answer is correct:
convex_hull(Points(11))
# + [markdown] run_control={}
# # Visualization of Results
#
# To visualize the results of the algorithm, I'll define a function to call `convex_hull` and plot the results:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
def plot_convex_hull(points):
"Find the convex hull of these points, and show a plot."
hull = convex_hull(points)
plot_points(points)
plot_points(hull, 'bs-', closed=True)
print(len(hull), 'of', len(points), 'points on hull')
# -
plot_convex_hull(Points(11))
# Now the octagon and pacman shapes:
plot_convex_hull(octagon)
plot_convex_hull(pacman)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# How about 100 random points?
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_convex_hull(Points(100))
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# Will 10,000 points be slow?
# +
P10K = Points(10000)
# %timeit convex_hull(P10K)
# -
# No problem! Still well under a second! Here's what it looks like:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
plot_convex_hull(P10K)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# How about a non-random set? Here is a set of coordinates of 80 US cities:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
P = Point
USA = {
P(-621, 289), P(-614, 297), P(-613, 319), P(-613, 342), P(-612, 263),
P(-612, 332), P(-603, 247), P(-599, 277), P(-592, 238), P(-591, 323),
P(-586, 229), P(-581, 289), P(-581, 305), P(-576, 253), P(-568, 260),
P(-563, 322), P(-560, 234), P(-560, 285), P(-559, 292), P(-558, 246),
P(-557, 259), P(-555, 225), P(-549, 271), P(-543, 321), P(-535, 313),
P(-530, 249), P(-524, 278), P(-524, 288), P(-515, 308), P(-505, 206),
P(-504, 327), P(-492, 207), P(-488, 194), P(-488, 248), P(-487, 264),
P(-484, 305), P(-484, 328), P(-482, 297), P(-480, 289), P(-477, 210),
P(-470, 319), P(-468, 291), P(-462, 247), P(-461, 328), P(-452, 271),
P(-450, 210), P(-450, 226), P(-450, 245), P(-441, 311), P(-440, 301),
P(-438, 233), P(-438, 293), P(-431, 278), P(-425, 266), P(-423, 273),
P(-422, 213), P(-422, 236), P(-420, 251), P(-415, 297), P(-413, 196),
P(-409, 214), P(-409, 290), P(-401, 181), P(-401, 253), P(-400, 230),
P(-400, 282), P(-394, 251), P(-394, 301), P(-387, 263), P(-385, 272),
P(-371, 285), P(-370, 285), P(-369, 299), P(-363, 309), P(-357, 292),
P(-355, 297), P(-352, 306), P(-344, 314), P(-340, 328), P(-608, 270)
}
plot_convex_hull(USA)
# + [markdown] run_control={}
# A decidedly non-random set of points:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
grid = {Point(x+0.5, y+0.5)
for x in range(10) for y in range(10)}
plot_convex_hull(grid)
# + [markdown] run_control={}
# A variant with some noise thrown in:
# +
def noisy(points, d=0.3, seed=42):
"Add some uniform noise to each of the points."
random.seed(seed)
def noise(): return random.uniform(-d, +d)
return {Point(x + noise(), y + noise())
for (x, y) in points}
plot_convex_hull(noisy(grid))
# -
# Circles and donuts:
square = {Point(random.uniform(-1, 1), random.uniform(-1, 1)) for _ in range(1000)}
circle = {p for p in square if p.x ** 2 + p.y ** 2 < 1}
plot_convex_hull(circle)
donut = {p for p in square if 0.2 < (p.x ** 2 + p.y ** 2) < 1}
plot_convex_hull(donut)
# +
def rad(degrees): return degrees * math.pi / 180.0
sine = {Point(rad(d), 5 * math.sin(rad(d))) for d in range(720)}
plot_convex_hull(noisy(sine))
# +
donut2 = noisy({Point(5 * math.sin(rad(d)), 5 * math.cos(rad(d))) for d in range(360)})
plot_convex_hull(donut2)
# -
# # Tests
#
# So far, everything looks good! But I would gain even more confidence if we could pass a test suite:
# +
def tests():
# Tests of `turn`
assert turn(octagon[0], octagon[1], octagon[2]) == 'left'
assert turn(octagon[2], octagon[3], octagon[4]) == 'left'
assert turn(octagon[1], octagon[0], octagon[7]) == 'right'
assert turn(octagon[5], octagon[6], octagon[7]) == 'left'
assert turn(octagon[2], octagon[1], octagon[0]) == 'right'
assert turn(pacman[1], pacman[2], pacman[3]) == 'left'
assert turn(pacman[3], pacman[4], pacman[5]) == 'right'
assert turn(Point(0, 0), Point(0, 1), Point(0, 2)) == 'straight'
assert turn(Point(2, 1), Point(3, 1), Point(4, 1)) == 'straight'
assert turn(Point(2, 1), Point(4, 1), Point(3, 1)) == 'straight'
assert turn(Point(0, 0), Point(1, 1), Point(2, 2)) == 'straight'
assert turn(Point(0, 0), Point(-1, -1), Point(2, 2)) == 'straight'
# More tests of `turn`, covering negative denominator
A, B = Point(-2, -2), Point(0, 0)
assert turn(A, B, Point(1, 3)) == 'left'
assert turn(A, B, Point(2, 2)) == 'straight'
assert turn(A, B, Point(3, 1)) == 'right'
assert turn(A, B, Point(-1, 1)) == 'left'
assert turn(A, B, Point(-1, -4)) == 'right'
assert turn(A, B, Point(-1, -1)) == 'straight'
assert turn(B, A, Point(-3, -4)) == 'left'
assert turn(B, A, Point(-4, -3)) == 'right'
assert turn(B, A, Point(-1, -1)) == 'straight'
assert turn(B, A, Point(-3, -3)) == 'straight'
# Tests of convex_hull
assert convex_hull(octagon)== octagon
assert convex_hull(circle) == convex_hull(donut)
assert convex_hull(circle) == convex_hull(convex_hull(circle))
for n in (0, 1, 2, 3):
assert convex_hull(Points(n)) == Points(n)
collinear = {Point(x, 0) for x in range(100)}
assert convex_hull(collinear) == [min(collinear), max(collinear)]
P = Point(5, 5)
assert convex_hull(collinear | {P}) == [min(collinear), max(collinear), P]
grid1 = {Point(x, y) for x in range(10) for y in range(10)}
assert convex_hull(grid1) == [Point(0, 0), Point(9, 0), Point(9, 9), Point(0, 9)]
return 'tests pass'
tests()
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# ## How Many Points on the Hull?
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# The number of points on the hull for `Points(N)` seems to increase slowly as `N` increases.
# How slowly? Let's try to find out. We'll average the number of points on the hull for `Points(N)` over, say, 60 random trials:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
def average_hull_size(N, trials=60):
"""Compute the average hull size of N random points
(averaged over the given number of random trials)."""
return sum(len(convex_hull(Points(N, seed=trials+i)))
for i in range(trials)) / trials
# -
# We'll do this for several values of *N*, taken as powers of 2:
# +
hull_sizes = [average_hull_size(2**e)
for e in range(14)]
print(' N Hull Size')
for e in range(14):
print('{:4}: {:4.1f}'.format(2**e, hull_sizes[e]))
# -
# Then we'll plot the results, with *N* on a log<sub>2</sub> scale:
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
def plot_hull_sizes(hull_sizes):
plt.plot(hull_sizes, 'bo-')
plt.ylabel('Hull size')
plt.xlabel('log_2(number of points)')
plot_hull_sizes(hull_sizes)
# -
# That sure looks like a straight line!
#
# That means we can define `estimated_hull_size` by computing a slope and intercept of the line. (I won't bother doing linear regression; I'll just draw a straight line from the first to the last point in `hull_sizes`.)
# + button=false deletable=true new_sheet=false run_control={"read_only": false}
def estimated_hull_size(N):
"Estimated hull size for N random points, (inter/extra)polating from hull_sizes."
slope = (hull_sizes[-1] - hull_sizes[0]) / (len(hull_sizes) - 1)
return hull_sizes[0] + slope * math.log(N, 2)
# Plot actual average hull sizes in blue, and estimated hull sizes in red
plot_hull_sizes(hull_sizes)
plt.plot([estimated_hull_size(2**e)
for e in range(len(hull_sizes))],
'r--');
# -
# Here's an estimate of the number of points on the hull of a quadrillion random points:
estimated_hull_size(10**15)
# + [markdown] button=false deletable=true new_sheet=false run_control={"read_only": false}
# # Concluding Remarks and Further Reading
#
# The convex hull problem is an interesting exercise in algorithm design.
# The algorithm covered here is called [Andrew's Monotone Chain](https://en.wikibooks.org/wiki/Algorithm_Implementation/Geometry/Convex_hull/Monotone_chain).
# It is a variant of the [Graham Scan](https://en.wikipedia.org/wiki/Graham_scan).
# You can read more from [Tamassia](http://cs.brown.edu/courses/cs016/docs/old_lectures/ConvexHull-Notes.pdf) or [Wikipedia](https://en.wikipedia.org/wiki/Convex_hull).
| ipynb/Convex Hull.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
s = pd.Series(list('asdadeadesdasesda'))
s
s.unique()
s.value_counts()
dados = pd.read_csv('aluguel.csv', sep = ';')
dados
dados.Tipo.unique()
dados.Tipo.value_counts()
# ### Testando moedas
m1 = 'CCcCCccCCCccCcCccCcCcCCCcCCcccCCcCcCcCcccCCcCcccCc'
m2 = 'CCCCCccCccCcCCCCccCccccCccCccCCcCccCcCcCCcCccCccCc'
m3 = 'CccCCccCcCCCCCCCCCCcccCccCCCCCCccCCCcccCCCcCCcccCC'
m4 = 'cCCccCCccCCccCCccccCcCcCcCcCcCcCCCCccccCCCcCCcCCCC'
m5 = 'CCCcCcCcCcCCCcCCcCcCCccCcCCcccCccCCcCcCcCcCcccccCc'
# c = Cara
# C = Coroa
# +
# s1 = pd.Series(list(m1))
# s2 = pd.Series(list(m2))
# s3 = pd.Series(list(m3))
# s4 = pd.Series(list(m4))
# s5 = pd.Series(list(m5))
# +
# s1.value_counts()
# -
eventos = {'m1': list(m1),
'm2': list(m2),
'm3': list(m3),
'm4': list(m4),
'm5': list(m5)}
moedas = pd.DataFrame(eventos)
df = pd.DataFrame(data = ['Cara', 'Coroa'],
index = ['c', 'C'],
columns = ['Faces'])
for item in moedas:
df = pd.concat([df, moedas[item].value_counts()],
axis = 1)
df
moedas
| DS_02_Python_Pandas_Tratando_e_Analisando_Dados/extra_aula7_contadores.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# https://github.com/facebookresearch/ParlAI/blob/master/parlai/tasks/wizard_of_wikipedia/build.py
# +
import json
with open('train.json') as fopen:
data = json.load(fopen)
# +
informations = []
for row in data:
topic = row['chosen_topic']
persona = row['persona']
for k in row['chosen_topic_passage']:
text = f'{topic} <> {persona} <> {k}'
v = {'topic': topic, 'persona': persona, 'passage': k, 'text': text}
informations.append(v)
len(informations)
# -
informations[:2]
passages = []
for row in data:
topic = row['chosen_topic']
persona = row['persona']
for k in row['dialog']:
for z in k['retrieved_passages']:
for key, v in z.items():
for r in v:
text = f'{key} <> {persona} <> {r}'
v = {'topic': topic, 'persona': persona, 'text': text, 'key': key}
passages.append(v)
len(passages)
passages[:2]
# +
dialogs = []
for row in data:
topic = row['chosen_topic']
persona = row['persona']
for k in range(1, len(row['dialog'])):
before = row['dialog'][k - 1]['text']
now = row['dialog'][k]['text']
text = f'{persona} <> {before} <> {now}'
v = {'topic': topic, 'persona': persona, 'before': before, 'now': now, 'text': text}
dialogs.append(v)
# -
len(dialogs)
dialogs[:2]
with open('valid_random_split.json') as fopen:
valid = json.load(fopen)
# +
for row in valid:
topic = row['chosen_topic']
persona = row['persona']
for k in row['chosen_topic_passage']:
text = f'{topic} <> {persona} <> {k}'
v = {'topic': topic, 'persona': persona, 'passage': k, 'text': text}
informations.append(v)
len(informations)
# -
for row in valid:
topic = row['chosen_topic']
persona = row['persona']
for k in row['dialog']:
for z in k['retrieved_passages']:
for key, v in z.items():
for r in v:
text = f'{key} <> {persona} <> {r}'
v = {'topic': topic, 'persona': persona, 'text': text, 'key': key}
passages.append(v)
for row in valid:
topic = row['chosen_topic']
persona = row['persona']
for k in range(1, len(row['dialog'])):
before = row['dialog'][k - 1]['text']
now = row['dialog'][k]['text']
text = f'{persona} <> {before} <> {now}'
v = {'topic': topic, 'persona': persona, 'before': before, 'now': now, 'text': text}
dialogs.append(v)
with open('valid_topic_split.json') as fopen:
valid = json.load(fopen)
# +
for row in valid:
topic = row['chosen_topic']
persona = row['persona']
for k in row['chosen_topic_passage']:
text = f'{topic} <> {persona} <> {k}'
v = {'topic': topic, 'persona': persona, 'passage': k, 'text': text}
informations.append(v)
len(informations)
# +
for row in valid:
topic = row['chosen_topic']
persona = row['persona']
for k in row['dialog']:
for z in k['retrieved_passages']:
for key, v in z.items():
for r in v:
text = f'{key} <> {persona} <> {r}'
v = {'topic': topic, 'persona': persona, 'text': text, 'key': key}
passages.append(v)
len(passages)
# +
for row in valid:
topic = row['chosen_topic']
persona = row['persona']
for k in range(1, len(row['dialog'])):
before = row['dialog'][k - 1]['text']
now = row['dialog'][k]['text']
text = f'{persona} <> {before} <> {now}'
v = {'topic': topic, 'persona': persona, 'before': before, 'now': now, 'text': text}
dialogs.append(v)
len(dialogs)
# -
with open('test_topic_split.json') as fopen:
test = json.load(fopen)
# +
for row in test:
topic = row['chosen_topic']
persona = row['persona']
for k in row['chosen_topic_passage']:
text = f'{topic} <> {persona} <> {k}'
v = {'topic': topic, 'persona': persona, 'passage': k, 'text': text}
informations.append(v)
len(informations)
# +
for row in test:
topic = row['chosen_topic']
persona = row['persona']
for k in row['dialog']:
for z in k['retrieved_passages']:
for key, v in z.items():
for r in v:
text = f'{key} <> {persona} <> {r}'
v = {'topic': topic, 'persona': persona, 'text': text, 'key': key}
passages.append(v)
len(passages)
# +
for row in test:
topic = row['chosen_topic']
persona = row['persona']
for k in range(1, len(row['dialog'])):
before = row['dialog'][k - 1]['text']
now = row['dialog'][k]['text']
text = f'{persona} <> {before} <> {now}'
v = {'topic': topic, 'persona': persona, 'before': before, 'now': now, 'text': text}
dialogs.append(v)
len(dialogs)
# -
with open('test_random_split.json') as fopen:
test = json.load(fopen)
# +
for row in test:
topic = row['chosen_topic']
persona = row['persona']
for k in row['chosen_topic_passage']:
text = f'{topic} <> {persona} <> {k}'
v = {'topic': topic, 'persona': persona, 'passage': k, 'text': text}
informations.append(v)
len(informations)
# +
for row in test:
topic = row['chosen_topic']
persona = row['persona']
for k in row['dialog']:
for z in k['retrieved_passages']:
for key, v in z.items():
for r in v:
text = f'{key} <> {persona} <> {r}'
v = {'topic': topic, 'persona': persona, 'text': text, 'key': key}
passages.append(v)
len(passages)
# +
for row in test:
topic = row['chosen_topic']
persona = row['persona']
for k in range(1, len(row['dialog'])):
before = row['dialog'][k - 1]['text']
now = row['dialog'][k]['text']
text = f'{persona} <> {before} <> {now}'
v = {'topic': topic, 'persona': persona, 'before': before, 'now': now, 'text': text}
dialogs.append(v)
len(dialogs)
# -
dialogs[-10:]
with open('dialogs.json', 'w') as fopen:
json.dump(dialogs, fopen)
with open('passages.json', 'w') as fopen:
json.dump(passages, fopen)
with open('informations.json', 'w') as fopen:
json.dump(informations, fopen)
| chatbot/wiki-wizard/parse.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
''' testing importance sampling and convergence rates'''
# cd ..
from models.vae import VAE
import torch
import matplotlib.pyplot as plt
from PIL import Image
from scipy.misc import imresize as resize
import numpy as np
from torchvision import transforms
# +
vae = VAE(3, 32, conditional=True)
best_filename = 'exp_dir/vae/best.tar'
logger_filename = 'exp_dir/vae/'+ 'logger.json'
state = torch.load(best_filename, map_location={'cuda:0': 'cpu'})
print("Reloading vae at epoch {}"
", with test error {}".format(
state['epoch'],
state['precision']))
vae.load_state_dict(state['state_dict'])
# -
obs_file = 'datasets/downloads_from_server/trimmed_rollout0.npz'
# +
transform = transforms.Compose([
transforms.ToPILImage(),
transforms.Resize((64, 64)),
transforms.ToTensor()
])
data = np.load(obs_file)
observations = data['observations']
rewards = data['rewards']
print(observations.shape)
ind = 600
obs = transform(observations[ind])
rew = rewards[ind]
plt.figure()
plt.imshow(obs.permute(1,2,0))
plt.show()
# -
obs.shape
# ## Generating Reconstructions of the Image
from torchvision.utils import save_image
IMAGE_RESIZE_DIM=64
with torch.no_grad():
last_test_observations = obs.unsqueeze(0)
last_test_rewards = torch.Tensor([rew]).unsqueeze(0)
encoder_mu, encoder_logsigma, latent_s, decoder_mu, decoder_logsigma = vae(last_test_observations, last_test_rewards)
recon_batch = decoder_mu + (decoder_logsigma.exp() * torch.randn_like(decoder_mu))
recon_batch = recon_batch.view(recon_batch.shape[0], 3, IMAGE_RESIZE_DIM, IMAGE_RESIZE_DIM)
#sample = torch.randn(IMAGE_RESIZE_DIM, LATENT_SIZE).to(device) # random point in the latent space.
# image reduced size by the latent size. 64 x 32. is this a batch of 64 then??
#sample = vae.decoder(sample).cpu()
decoder_mu = decoder_mu.view(decoder_mu.shape[0], 3, IMAGE_RESIZE_DIM, IMAGE_RESIZE_DIM)
to_save = torch.cat([last_test_observations.cpu(), recon_batch.cpu(), decoder_mu.cpu()], dim=0)
print('to save shape', to_save.shape)
save_image(to_save,
'notebooks/test_vae_recon' + '.png')
# ## Looking at the learnt Sigmas
plt.hist(decoder_logsigma.exp().cpu())
sigmas = decoder_logsigma.exp().view(recon_batch.shape[0], 3,
IMAGE_RESIZE_DIM, IMAGE_RESIZE_DIM).squeeze().cpu()
sigmas.shape
import seaborn as sns
for i in range(3):
plt.figure()
ax = sns.heatmap(sigmas[i])
plt.show()
plt.figure()
ax = sns.heatmap(sigmas.mean(dim=0))
plt.show()
# ## Evaluating estimator efficiency
#
# Using importance sampling. $E_{q(z|x)}[ p(x|z)p(z) / q(z|x) ]$
last_test_observations.shape
last_test_rewards.shape
# +
#delta = torch.Tensor([0.000001])
max_samps = 200
rand_inds = np.random.randint(0,1000, 10)
for rand_ind in rand_inds:
obs = transform(observations[rand_ind])
rew = rewards[rand_ind]
last_test_observations = obs.unsqueeze(0)
last_test_rewards = torch.Tensor([rew]).unsqueeze(0)
with torch.no_grad():
p_o = [0.0]
p_o_rew = [0.0]
encoder_mu, encoder_logsigma = vae.encoder(last_test_observations, last_test_rewards)
real_obs = last_test_observations.view(last_test_observations.size(0), -1) # flattening all but the batch.
log_p_r = torch.distributions.Normal(3.5, 0.1).log_prob(rew)
def importance_sample( ):
z = encoder_mu + (encoder_logsigma.exp() * torch.randn_like(encoder_mu))
decoder_mu, decoder_logsigma = vae.decoder(z, last_test_rewards)
log_P_OBS_GIVEN_S = torch.distributions.Normal(decoder_mu, decoder_logsigma.exp()).log_prob(real_obs)
log_P_OBS_GIVEN_S = log_P_OBS_GIVEN_S.sum(dim=-1) #multiply the probabilities within the batch.
#log_P_OBS_GIVEN_S = log_P_OBS_GIVEN_S+torch.log(delta)
log_P_S = torch.distributions.Normal(0.0, 1.0).log_prob(z).sum(dim=-1)
log_Q_S_GIVEN_X = torch.distributions.Normal(encoder_mu, encoder_logsigma.exp()).log_prob(z).sum(dim=-1)
#print(log_P_OBS_GIVEN_S, log_P_S, log_Q_S_GIVEN_X)
return log_P_OBS_GIVEN_S+log_P_S - log_Q_S_GIVEN_X
for i in range(max_samps):
next_prob = importance_sample()
p_o.append(p_o[-1]+next_prob)
p_o_rew.append(p_o_rew[-1]+next_prob+log_p_r)
p_o = p_o[1:]
p_o_rew = p_o_rew[1:]
p_o = np.array(p_o) / (np.arange(max_samps)+1)
p_o_rew = np.array(p_o_rew) / (np.arange(max_samps)+1)
plt.figure()
plt.plot(np.arange(max_samps)+1, p_o, label='p_o')
plt.plot(np.arange(max_samps)+1, p_o_rew, label = 'rew')
plt.xlabel('number of samples')
plt.ylabel('log_prob')
plt.legend()
plt.show()
plt.figure()
plt.imshow(obs.permute(1,2,0))
plt.show()
print('='*10)
# +
# full loss function:
# -
expand_shape = list(encoder_mu.shape)
expand_shape.append(3)
#expand_shape = torch.Tensor(expand_shape)
expand_shape
torch.randn(expand_shape).shape
(encoder_logsigma.exp().unsqueeze(-1) * torch.randn(expand_shape)).shape
# ## Conditional VAE
with torch.no_grad():
images = []
for _ in range(10):
# sample Z from standard normal and condition on different r's.
p_z = torch.distributions.Normal(0.0, 1.0)
z = p_z.sample((1,32))
cond_reward_high = torch.Tensor([[3.1]])
cond_reward_mid = torch.Tensor([[1.1]])
cond_reward_low = torch.Tensor([[-0.1]])
for c_rew in [3.1, 1.1, -0.1]:
cond_reward = torch.Tensor([[c_rew]])
decoder_mu, decoder_logsigma = vae.decoder(z, cond_reward)
recon_batch = decoder_mu + (decoder_logsigma.exp() * torch.randn_like(decoder_mu))
recon_batch = recon_batch.view(recon_batch.shape[0], 3, IMAGE_RESIZE_DIM, IMAGE_RESIZE_DIM)
decoder_mu = decoder_mu.view(decoder_mu.shape[0], 3, IMAGE_RESIZE_DIM, IMAGE_RESIZE_DIM)
images.append(decoder_mu.squeeze())
plt.figure()
plt.title('Mu '+str(c_rew))
plt.imshow(decoder_mu.squeeze().permute(1,2,0))
plt.show()
'''plt.figure()
plt.title('sample '+str(c_rew))
plt.imshow(recon_batch.squeeze().permute(1,2,0))
plt.show()'''
print('='*10)
to_save = torch.stack(images) #torch.cat([last_test_observations.cpu(), recon_batch.cpu(), decoder_mu.cpu()], dim=0)
print('to save shape', to_save.shape)
save_image(to_save,
'notebooks/rew_conds' + '.png')
recon_batch
p_z = torch.distributions.Normal(0.0, 1.0)
| notebooks/log_p_obs.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # OOP Exercise - Part 1
#
#
# # Shirt Class 만들어 보기
#
# 클래스가 어떻게 구성되어야 하는지와 객체를 인스턴스화하는 방법을 보았습니다.
#
# 이제 Shirt 클래스를 인스턴스화 하는 코드를 작성해 보겠습니다.
class Shirt:
def __init__(self, shirt_color, shirt_size, shirt_style, shirt_price):
self.color = shirt_color
self.size = shirt_size
self.style = shirt_style
self.price = shirt_price
def change_price(self, new_price):
self.price = new_price
def discount(self, discount):
return self.price * (1 - discount)
# +
# color red, size S, style long-sleeve, price 25의 특성을 가진 shirt class를 인스턴스화 해보겠습니다.
shirt_one = Shirt(shirt_color = "red", shirt_size = "S", shirt_style = "long-sleeve", shirt_price = 25)
# +
# 제대로 인스턴스화가 되었는지 살펴보기 위해 한 속성을 살펴봅시다.
print(shirt_one.price)
# 메서드는 잘 작동할까요?
shirt_one.change_price(10)
print(shirt_one.price)
print(shirt_one.discount(.12))
# +
# 이번에는 color orange, size L, style short-sleeve, price 10을 속성으로 가지는 클래스를 만들어 보겠습니다.
shirt_two = Shirt(shirt_color = "orange", shirt_size = "L", shirt_style = "short-sleeve", shirt_price = 10)
# +
# 두 인스턴스의 price를 가지고 total cost를 계산해 볼 수도 있습니다.
total = shirt_one.price + shirt_two.price
# +
# discount 매서드를 사용한 뒤에 값을 계산하는 것 역시 가능합니다.
total_discount = shirt_one.discount(.14) + shirt_two.discount(.06)
| _posts/.ipynb_checkpoints/2020-08-01-aws-08_shirt_exercise-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Best Hour of the Day to Post on Hacker News: Analyzing the Posts' Data
#
# This project aims to analyze a dataset of submissions to the popular technology site Hacker News. According to [Wikipedia](https://en.wikipedia.org/wiki/Hacker_News), Hacker News is a social news website focused on computer science and entrepreneurship and it is run by Y Combinator, <NAME>ham's investment fund and startup incubator.
#
# The dataset we'll use is available [here](https://www.kaggle.com/hacker-news/hacker-news-posts) and has almost 300,000 rows, each row representing a post. It includes the following columns:
#
# * title: title of the post (self explanatory)
#
# * url: the url of the item being linked to
#
# * num_points: the number of upvotes the post received
#
# * num_comments: the number of comments the post received
#
# * author: the name of the account that made the post
#
# * created_at: the date and time the post was made (the time zone is Eastern Time in the US)
#
# For this project, we are particulary interested in posts whose titles begin with *Ask HN* and *Show HN*. The first one is used to aks the community a question while the second one is used to show the community something, it could be a project, a product, or just something the author finds interesting enough to share. Our goal is to determine if a post created in a particular moment of the day is more interacted with than posts cretated in other moments. In another words, we are interesd in answering the question: is there a best moment of the day to post on Hacker News?
#
#
# 
# ## Exploring the Data
#
# We'll begin by importing the libraries we'll use and reading the dataset into a dataframe. Then we'll display the first five rows.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
hn_data = pd.read_csv('HN_posts_year_to_Sep_26_2016.csv', encoding='utf8')
hn_data.head()
# -
# The dataframe contains seven columns, but we are most interested in the `num_points`, `num_comments` and `created_at` columns to answer hour question.
#
# Let's see more information about the data.
hn_data.describe(include='all')
# We can see that the posts receive, in average, 15 points and 6.5 comments.
# As we are looking for Ask HN and Show HN posts, we will now create three new dataframes:
#
# * A dataframe for the Ask Hn posts;
# * A dataframe for the Show HN posts;
# * A dataframe for the other posts.
#
# The we'll see how many posts each dataframe contains.
# +
# Creating the dataframes
ask = hn_data[hn_data['title'].str.lower().str.startswith('ask hn')].copy()
show = hn_data[hn_data['title'].str.lower().str.startswith('show hn')].copy()
other = hn_data[~((hn_data['title'].str.lower().str.startswith('ask hn')) | (hn_data['title'].str.lower().str.startswith('show hn')))].copy()
# Printing the number of rows
print('Number of Ask posts: ', ask.shape[0])
print('Number of Show posts: ', show.shape[0])
print('Number of Other posts: ', other.shape[0])
# -
# We can see that the majority of posts are neither Ask HN nor Show HN. This does not affect our goal, though.
#
# Let's do some quickly exploring in these dataframes.
ask.describe(include='all')
show.describe(include='all')
other.describe(include='all')
# We can see the that Ask HN posts receive about twice as much commentaries than the Show HN posts, but less points, in average. Users seem to be more interested in answer people's questions than interacting with what they want to show. The Other posts receive more than 15 points and 6 comments in average.
# ## Working with Dates
#
# The `created_at` column contains data about the moment each post was created. As our goal in this project is to determine the best moment of the day to creat a post, we need to extract the value of the hour from this column. For that, in each dataframe, we will:
#
# * Convert the values in the `created_at` column to datetime objects using the `pandas.to_datetime()` function;
# * Create the `hour` column and fill it with the values for the hour using the `Series.dt.hour` method;
# * Create a series containing the hour column sorted by index, so it's easier to plot.
# +
# Converting the values to datetime
ask['created_at'] = pd.to_datetime(ask['created_at'])
show['created_at'] = pd.to_datetime(show['created_at'])
other['created_at'] = pd.to_datetime(other['created_at'])
# Creating the 'hour' column
ask['hour'] = ask['created_at'].dt.hour.astype(int)
show['hour'] = show['created_at'].dt.hour.astype(int)
other['hour'] = other['created_at'].dt.hour.astype(int)
# Creating series sorted by index
ask_posts = ask['hour'].value_counts().sort_index()
show_posts = show['hour'].value_counts().sort_index()
other_posts = other['hour'].value_counts().sort_index()
# -
# Now let's plot these series.
# +
fig, ax = plt.subplots(figsize=(15,5))
ax.plot(ask_posts.index, ask_posts, linewidth=3, color=(109/255, 204/255, 218/255), label='Ask HN')
ax.plot(show_posts.index, show_posts, linewidth=3, color=(205/255, 204/255, 93/255), label='Show HN')
ax.tick_params(bottom=False, top=False, left=False, right=False)
ax.set_yticks([300, 600, 900])
ax.set_xticks(ask_posts.index)
ax.set_title('Posts Created per Hour')
ax.legend(loc='upper left')
for kew, spine in ax.spines.items():
spine.set_visible(False)
plt.show()
# +
fig, ax = plt.subplots(figsize=(15,5))
ax.plot(other_posts.index, other_posts, linewidth=3, color=(255/255, 158/255, 74/255), label='Other')
ax.tick_params(bottom=False, top=False, left=False, right=False)
ax.set_yticks([10000, 20000])
ax.set_xticks(other_posts.index)
ax.set_title('Posts Created per Hour')
ax.legend(loc='upper left')
for kew, spine in ax.spines.items():
spine.set_visible(False)
plt.show()
# -
# We can see that the majority of posts are created after 12 o'clock, especially from 14 to 18 o'clock, reaching the summit around from 15 to 16 o´clock.
#
# We had to plot the Other posts separately because the number of posts would compromise the visualization of the Show HN and Ask HN posts.
# ## Analyzing the Number of Comments and Points
#
# We will now use the `DataFrame.groupby()` method to calculate the average number of points and comments per post created in each hour of the day. We'll do this for each dataframe and asign the results to variables
# +
ask_com = ask.groupby('hour')['num_comments'].mean().sort_index()
ask_pts = ask.groupby('hour')['num_points'].mean().sort_index()
show_com = show.groupby('hour')['num_comments'].mean().sort_index()
show_pts = show.groupby('hour')['num_points'].mean().sort_index()
other_com = other.groupby('hour')['num_comments'].mean().sort_index()
other_pts = other.groupby('hour')['num_points'].mean().sort_index()
# -
# And now let's visualize this data and see what we can conclude from it.
# +
fig, (ax1, ax2) = plt.subplots(2, 1, figsize=(15,10))
ax1.plot(ask_com.index, ask_com, linewidth=4, color=(109/255, 204/255, 218/255), label='Ask HN')
ax1.plot(show_com.index, show_com, linewidth=4, color=(205/255, 204/255, 93/255), label='Show HN')
ax1.plot(other_com.index, other_com, linewidth=2, color=(214/255, 39/255, 40/255), label='Other')
ax1.tick_params(bottom=False, top=False, left=False, right=False, labelbottom=False)
ax1.set_xticks(ask_com.index)
ax1.set_yticks([10, 20, 30])
ax1.set_title('Average comments per post created in each hour of the day')
ax1.legend(loc='upper right')
for kew, spine in ax1.spines.items():
spine.set_visible(False)
ax2.plot(ask_pts.index, ask_pts, linewidth=4, color=(109/255, 204/255, 218/255))
ax2.plot(show_pts.index, show_pts, linewidth=4, color=(205/255, 204/255, 93/255))
ax2.plot(other_pts.index, other_pts, linewidth=2, color=(214/255, 39/255, 40/255))
ax2.tick_params(bottom=False, top=False, left=False, right=False)
ax2.set_xticks(ask_pts.index)
ax2.set_yticks([10, 20, 25])
ax2.set_title('Average points per post created in each hour of the day')
for kew, spine in ax2.spines.items():
spine.set_visible(False)
plt.tight_layout()
plt.show()
# -
# Aparently, only for Ask HN there's a great difference in the number of comments if the post is created in one specific hour of day. If you have question to the Hacker News community, the best moment by far to submit it is between 15 and 16 o'clock as posts created in this period of time receive almost 30 comments in average. Subimiting a question between 12 and 14 o'clock is also good as these time periods are the second and third more commentend upon.These periods are also the best to creat your Ask HN posts if want to get more points.
#
# If the goal of a submission is to show the community something usign Show HN, the period of time in which the post is created does not make such a great differenece in the numer of comments. However, Show HN posts receive more points when they are created between 10 and 14 o'clock. If we assume that a post with more points is also more visualized, then that is the best period of time to show your ideias to the Hacker News community. Show HN posts created between 12 and 13 o'clock rceive, in average, almost 21 points.
#
# If your post is neither a Ask HN nor a Show HN post, the hour of day in which it is created makes even less difference as the range of average comments goes only from 7.59 comments per post at the best moment for creating a post and 5.84 comments per post at the worst moment to do it. The range of average points only goes from 16.71 to 13.79 points, so the moment you creat this kind of post is not important.
# # Conclusion
#
# In this project we went through a dataset containing data from almost 300,000 Hacker News submissions. Our goal was to determine if there's a particular moment of the day in which creating posts would draw more attention to the post.
#
# We are now able to conclude that for Ask HN posts there are definitely better moments of the day to submit a post. As for the Show HN posts, we still can consider that there is at least one best moment of the day to submit something, but the difference is not as big as in the Ask HN posts. For the other posts the difference between posting in a determined moment or in any moment of the day is so small that we should not take it into consideration.
#
# Finally, this data can be useful if you're looking to show a product or a project and wants it to draw some attetion or even if you need to ask a question and it is important that your question receive a lot of answers. For those scenarios you should definitely consider that there are better moments of the day to create your post.
#
| The Best Hour of the Day to Post on Hacker News - Analyzing the Posts' Data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from utilities_namespace import *
# from helpers import developer_mode
from signature_scoring.scoring_functions import gsea
# from importlib import reload; reload(gsea)
pd.options.mode.chained_assignment = None
# ## Load data & methods definitions
from helpers.notebooks import notebooks_importer
# %%capture
import Breast_cancer_data as data_brca
# %%capture
from Benchmarking_setup import standard_benchmark
from functools import partial
brca_standard_benchmark = partial(
standard_benchmark,
# force the pipline to show full progress and use
# my custom multiprocessing (which is more reliable)
per_test_progress=True
)
# %%capture
from Selected_functions import (
selected_single_sample_functions,
selected_multi_sample_functions
)
# %%capture
import Published_BRCA_stratifications
from Published_BRCA_stratifications import (
# this gets us the participant-cluster relation for different clustering methods
all_stratifications,
# this is one of them; different as based only on expression,
# thus able to explor tumour heterogenity
pam50_brca
)
# %store -r validation_perturbations
from data_sources.tcga.stratification import get_subtype_by_sample, group_by_subtype
pam50_subtype_sample_df = get_subtype_by_sample(data_brca.brca_expression, pam50_brca, subtype_column='subtype_selected')
pam50_subtype_sample_df.head()
pam50_samples_by_type = group_by_subtype(pam50_subtype_sample_df, subtype_column='subtype_selected')
# ## Permutations generation
from signature_scoring.evaluation.subtypes import random_subtypes_benchmark
from signature_scoring.evaluation import permutations
def generate_permutations(funcs, stratifications, n, packages, single_sample, comment=''):
prefix = 'ss' if single_sample else 'ms'
for name, samples_by_type in samples_by_statification_and_subtype.items():
print(name)
generate_permutations_package = partial(
permutations.generate,
random_subtypes_benchmark,
data_brca.brca_expression,
samples_by_type,
benchmark_partial=partial(
brca_standard_benchmark,
indications_signatures=data_brca.indications_singatures,
contraindications_signatures=data_brca.contraindications_singatures,
unassigned_signatures=validation_perturbations
),
)
for package in range(packages):
print(package)
pickle_name = f'{prefix}_{name}_{comment}_{package}'.replace('__', '_')
generate_permutations_package(
funcs=funcs,
n=n, pickle_name=pickle_name,
single_sample=single_sample,
multi_sample=not single_sample
)
# ### The odd case - PAM50:
# PAM50 has subtypes assigned to samples, not participants thus has data generated separately:
# %store -r samples_by_statification_and_subtype
samples_by_statification_and_subtype['pam50'] = pam50_samples_by_type
# ### Single sample stratifications
# Using 10 * 20 = 200 permutations
generate_permutations(
selected_single_sample_functions,
samples_by_statification_and_subtype,
n=10, packages=20, single_sample=True
)
# ### cudaGSEA functions
#
# had much less permutations (only 50) in the thesis as those can be only calculated on GPU and there was only one GPU-enabled computer available:
cuda_funcs = {
gsea_score_phenotypes_cuda_hallmarks,
gsea_score_phenotypes_cuda_reactome
}
generate_permutations(
list(cuda_funcs),
samples_by_statification_and_subtype,
n=5, packages=10, single_sample=False,
comment='cuda_only'
)
# ### All the other multi-sample functions have the full 200 permutations
generate_permutations(
list(set(selected_multi_sample_functions) - cuda_funcs),
samples_by_statification_and_subtype,
n=10, packages=20, single_sample=False,
comment='cuda_only'
)
| notebooks/Signature-based drug-disease associations/Permutation_generation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Solving a New Keynesian model with Python
#
# This notebook is part of a computational appendix that accompanies the paper.
#
# > MATLAB, Python, Julia: What to Choose in Economics?
# >> Coleman, Lyon, Maliar, and Maliar (2017)
#
# In order to run the codes in this notebook you will need to install and configure a few Python packages. We recommend following the instructions on [quantecon.org](https://lectures.quantecon.org/jl/getting_started.html) for getting a base python installation set up. Then to acquire additional packages used in this notebook, uncomment the lines in the cell below (delete the `#` and space at the beginning of the line) and then run the cell:
# +
# # !pip install git+https://github.com/EconForge/interpolation.py.git
# # !pip install git+https://github.com/naught101/sobol_seq.git
# + [markdown] outputExpanded=false
# ## Python Code
#
# The Python version of our algorithm is implemented as a few methods defined on
# a core class named `Model`. This class is itself composed of instances of three
# different classes that hold the model parameters, steady state, and grids
# needed to describe the numerical model. Before we get to the classes, we need
# to bring in some dependencies:
# + outputExpanded=false
import math
from math import sqrt
import time as time
from collections import namedtuple
import numpy as np
from numpy import exp
from interpolation.complete_poly import (_complete_poly_impl_vec,
_complete_poly_impl,
complete_polynomial)
import sobol_seq
# set seed on random number generator to make results reproducible
np.random.seed(42)
# + [markdown] outputExpanded=false
# We will also need the following two functions, which use monomial rules to
# compute quadrature nodes and weights:
# + outputExpanded=false
def qnwmonomial1(vcv):
n = vcv.shape[0]
n_nodes = 2*n
z1 = np.zeros((n_nodes, n))
# In each node, random variable i takes value either 1 or -1, and
# all other variables take value 0. For example, for N = 2,
# z1 = [1 0; -1 0; 0 1; 0 -1]
for i in range(n):
z1[2*i:2*(i+1), i] = [1, -1]
sqrt_vcv = np.linalg.cholesky(vcv)
R = np.sqrt(n)*sqrt_vcv
ϵj = z1 @ R
ωj = np.ones(n_nodes) / n_nodes
return ϵj, ωj
def qnwmonomial2(vcv):
n = vcv.shape[0]
assert n == vcv.shape[1], "Variance covariance matrix must be square"
z0 = np.zeros((1, n))
z1 = np.zeros((2*n, n))
# In each node, random variable i takes value either 1 or -1, and
# all other variables take value 0. For example, for N = 2,
# z1 = [1 0; -1 0; 0 1; 0 -1]
for i in range(n):
z1[2*i:2*(i+1), i] = [1, -1]
z2 = np.zeros((2*n*(n-1), n))
i = 0
# In each node, a pair of random variables (p,q) takes either values
# (1,1) or (1,-1) or (-1,1) or (-1,-1), and all other variables take
# value 0. For example, for N = 2, `z2 = [1 1; 1 -1; -1 1; -1 1]`
for p in range(n-1):
for q in range(p+1, n):
z2[4*i:4*(i+1), p] = [1, -1, 1, -1]
z2[4*i:4*(i+1), q] = [1, 1, -1, -1]
i += 1
sqrt_vcv = np.linalg.cholesky(vcv)
R = np.sqrt(n+2)*sqrt_vcv
S = np.sqrt((n+2)/2)*sqrt_vcv
ϵj = np.row_stack([z0, z1 @ R, z2 @ S])
ωj = np.concatenate([2/(n+2) * np.ones(z0.shape[0]),
(4-n)/(2*(n+2)**2) * np.ones(z1.shape[0]),
1/(n+2)**2 * np.ones(z2.shape[0])])
return ϵj, ωj
# + [markdown] outputExpanded=false
# ## Classes
#
# First we have the `Params` class, which holds all the model parameters as well
# as the paramters that drive the algorithm.
# + outputExpanded=false
SteadyState = namedtuple("SteadyState",
["Yn", "Y", "π", "δ", "L", "C", "F", "S", "R", "w"])
class Params(object):
def __init__(self, zlb=True, γ=1, β=0.99, ϑ=2.09, ϵ=4.45, ϕ_y=0.07,
ϕ_π=2.21, μ=0.82, Θ=0.83, πstar=1, gbar=0.23,
ρηR=0.0, ρηa=0.95, ρηL=0.25, ρηu=0.92, ρηB=0.0, ρηG=0.95,
σηR=0.0028, σηa=0.0045, σηL=0.0500, σηu=0.0054, σηB=0.0010,
σηG=0.0038, degree=2):
self.zlb = zlb # whether or not the zlb should be imposed
self.γ = γ # Utility-function parameter
self.β = β # Discount factor
self.ϑ = ϑ # Utility-function parameter
self.ϵ = ϵ # Parameter in the Dixit-Stiglitz aggregator
self.ϕ_y = ϕ_y # Parameter of the Taylor rule
self.ϕ_π = ϕ_π # Parameter of the Taylor rule
self.μ = μ # Parameter of the Taylor rule
self.Θ = Θ # Share of non-reoptimizing firms (Calvo's pricing)
self.πstar = πstar # Target (gross) inflation rate
self.gbar = gbar # Steady-state share of government spending in output
# autocorrelation coefficients
self.ρηR = ρηR # See process (28) in MM (2015)
self.ρηa = ρηa # See process (22) in MM (2015)
self.ρηL = ρηL # See process (16) in MM (2015)
self.ρηu = ρηu # See process (15) in MM (2015)
self.ρηB = ρηB # See process (17) in MM (2015)
self.ρηG = ρηG # See process (26) in MM (2015)
# standard deviations
self.σηR = σηR # See process (28) in MM (2015)
self.σηa = σηa # See process (22) in MM (2015)
self.σηL = σηL # See process (16) in MM (2015)
self.σηu = σηu # See process (15) in MM (2015)
self.σηB = σηB # See process (17) in MM (2015)
self.σηG = σηG # See process (26) in MM (2015)
self.degree = degree
@property
def vcov(self):
return np.diag([self.σηR**2, self.σηa**2, self.σηL**2,
self.σηu**2, self.σηB**2, self.σηG**2])
@property
def steady_state(self):
Yn_ss = exp(self.gbar)**(self.γ/(self.ϑ+self.γ))
Y_ss = Yn_ss
π_ss = 1.0
δ_ss = 1.0
L_ss = Y_ss/δ_ss
C_ss = (1-self.gbar)*Y_ss
F_ss = C_ss**(-self.γ)*Y_ss/(1-self.β*self.Θ*π_ss**(self.ϵ-1))
S_ss = L_ss**self.ϑ*Y_ss/(1-self.β*self.Θ*π_ss**self.ϵ)
R_ss = π_ss/self.β
w_ss = (L_ss**self.ϑ)*(C_ss**self.γ)
return SteadyState(Yn_ss, Y_ss, π_ss, δ_ss, L_ss, C_ss, F_ss, S_ss, R_ss, w_ss)
# + [markdown] outputExpanded=false
# Notice that we have a namedtuple to hold the steady state of the model. Using
# the namedtuple infrastructure allows us to have convenient "dot-style" access
# to the steady state, without defining a full class.
#
# Given an instance of `Params` class, we can construct the grid on which we will
# solve the model.
#
# The `Grids` class holds this grid as well as matrices used to compute
# expectations.
# + outputExpanded=false
class Grids(object):
def __init__(self, p, m=200, kind="rands"):
if kind == "sobol":
ub = np.array([
2 * p.σηR / sqrt(1 - p.ρηR**2),
2 * p.σηa / sqrt(1 - p.ρηa**2),
2 * p.σηL / sqrt(1 - p.ρηL**2),
2 * p.σηu / sqrt(1 - p.ρηu**2),
2 * p.σηB / sqrt(1 - p.ρηB**2),
2 * p.σηG / sqrt(1 - p.ρηG**2),
1.05, # R
1.0 # δ
])
lb = -ub
lb[[6, 7]] = [1.0, 0.95] # adjust lower bound for R and δ
s = sobol_seq.i4_sobol_generate(8, m)
s *= (ub - lb)
s += lb
ηR = s[:, 0]
ηa = s[:, 1]
ηL = s[:, 2]
ηu = s[:, 3]
ηB = s[:, 4]
ηG = s[:, 5]
R = s[:, 6]
δ = s[:, 7]
else:
# Values of exogenous state variables are distributed uniformly
# in the interval +/- std/sqrt(1-rho_nu**2)
ηR = (-2*p.σηR + 4*p.σηR*np.random.rand(m)) / sqrt(1-p.ρηR**2)
ηa = (-2*p.σηa + 4*p.σηa*np.random.rand(m)) / sqrt(1-p.ρηa**2)
ηL = (-2*p.σηL + 4*p.σηL*np.random.rand(m)) / sqrt(1-p.ρηL**2)
ηu = (-2*p.σηu + 4*p.σηu*np.random.rand(m)) / sqrt(1-p.ρηu**2)
ηB = (-2*p.σηB + 4*p.σηB*np.random.rand(m)) / sqrt(1-p.ρηB**2)
ηG = (-2*p.σηG + 4*p.σηG*np.random.rand(m)) / sqrt(1-p.ρηG**2)
# Values of endogenous state variables are distributed uniformly
# in the intervals [1 1.05] and [0.95 1], respectively
R = 1 + 0.05*np.random.rand(m)
δ = 0.95 + 0.05*np.random.rand(m)
self.ηR = ηR
self.ηa = ηa
self.ηL = ηL
self.ηu = ηu
self.ηB = ηB
self.ηG = ηG
self.R = R
self.δ = δ
# shape (8, m)
self.X = np.vstack([np.log(R), np.log(δ), ηR, ηa, ηL, ηu, ηB, ηG])
# shape (n_complete(8, 2), m)
self.X0_G = {
1: complete_polynomial(self.X, 1),
p.degree: complete_polynomial(self.X, p.degree)
}
# shape (2*n=12, n=6)
self.ϵ_nodes, self.ω_nodes = qnwmonomial1(p.vcov)
# all shape (len(ϵ_nodes), m)
self.ηR1 = p.ρηR * ηR[None, :] + self.ϵ_nodes[:, None, 0]
self.ηa1 = p.ρηa * ηa[None, :] + self.ϵ_nodes[:, None, 1]
self.ηL1 = p.ρηL * ηL[None, :] + self.ϵ_nodes[:, None, 2]
self.ηu1 = p.ρηu * ηu[None, :] + self.ϵ_nodes[:, None, 3]
self.ηB1 = p.ρηB * ηB[None, :] + self.ϵ_nodes[:, None, 4]
self.ηG1 = p.ρηG * ηG[None, :] + self.ϵ_nodes[:, None, 5]
# + [markdown] outputExpanded=false
# Finally, we construct the Model class, which has an instance of Params,
# SteadyState and Grids as its three attributes.
#
# This block of code will be longer than the others because we also include
# routines to solve and simulate the model as methods on the Model class. These
# methods will be clearly marked and commented.
# + outputExpanded=false
class Model(object):
def __init__(self, p=Params(), g=None):
if g is None:
g = Grids(p)
self.p = p
self.g = g
self.s = self.p.steady_state
def init_coefs(self):
"Iniital guess for coefs. We evaluate interpoland as coefs @ basis_mat"
npol = self.g.X0_G[1].shape[0]
coefs = np.full((3, npol), 1e-5)
coefs[:, 0] = [self.s.S, self.s.F, self.s.C**(-self.p.γ)]
return coefs
def step(self, S, F, C, δ0, R0, ηG, ηa, ηL, ηR):
# simplify notation
Θ, ϵ, gbar, ϑ, γ = self.p.Θ, self.p.ϵ, self.p.gbar, self.p.ϑ, self.p.γ
β, μ, ϕ_π, ϕ_y, πs = self.p.β, self.p.μ, self.p.ϕ_π, self.p.ϕ_y, self.s.π
# Compute pie(t) from condition (35) in MM (2015)
π0 = ((1-(1-Θ)*(S/F)**(1-ϵ))/Θ)**(1/(ϵ-1))
# Compute delta(t) from condition (36) in MM (2015)
δ1 = ((1-Θ)*((1-Θ*π0**(ϵ-1))/(1-Θ))**(ϵ/(ϵ-1))+Θ*π0**ϵ/δ0)**(-1)
# Compute Y(t) from condition (38) in MM (2015)
Y0 = C/(1-gbar/exp(ηG))
# Compute L(t) from condition (37) in MM (2015)
L0 = Y0/exp(ηa)/δ1
# Compute Yn(t) from condition (31) in MM (2015)
Yn0 = (exp(ηa)**(1+ϑ)*(1-gbar/exp(ηG))**(-γ)/exp(ηL))**(1/(ϑ+γ))
# Compute R(t) from conditions (27), (39) in MM (2015) -- Taylor rule
R1 = πs/β*(R0*β/πs)**μ*((π0/πs)**ϕ_π * (Y0/Yn0)**ϕ_y)**(1-μ)*exp(ηR)
return π0, δ1, Y0, L0, Yn0, R1
def solve(self, damp=0.1, tol=1e-7):
# rename self to m to make code below readable
m = self
n = len(m.g.ηR)
n_nodes = len(m.g.ω_nodes)
## allocate memory
# euler equations
e = np.zeros((3, n))
# previous iteration S, F, C
S0_old_G = np.ones(n)
F0_old_G = np.ones(n)
C0_old_G = np.ones(n)
# current iteration S, F, C
S0_new_G = np.ones(n)
F0_new_G = np.ones(n)
C0_new_G = np.ones(n)
# future S, F, C
S1 = np.zeros((n_nodes, n))
F1 = np.zeros((n_nodes, n))
C1 = np.zeros((n_nodes, n))
for deg in [1, self.p.degree]:
# housekeeping
err = 1.0
X0_G = m.g.X0_G[deg]
if deg > 1:
# compute degree coefs using degree 1 coefs as guess
coefs = np.linalg.lstsq(X0_G.T, e.T)[0].T
else:
coefs = self.init_coefs()
while err > tol:
# Current choices (at t)
# ------------------------------
SFC0 = coefs @ X0_G
S0 = SFC0[0, :] # Compute S(t) using coefs
F0 = SFC0[1, :] # Compute F(t) using coefs
C0 = (SFC0[2, :])**(-1/m.p.γ) # Compute C(t) using coefs
π0, δ1, Y0, L0, Yn0, R1 = self.step(S0, F0, C0, m.g.δ, m.g.R,
m.g.ηG, m.g.ηa, m.g.ηL, m.g.ηR)
if self.p.zlb:
R1 = np.maximum(R1, 1.0)
for u in range(n_nodes):
# Form complete polynomial of degree "Degree" (at t+1) on future state
grid1 = [np.log(R1), np.log(δ1), m.g.ηR1[u, :], m.g.ηa1[u, :],
m.g.ηL1[u, :], m.g.ηu1[u, :], m.g.ηB1[u, :], m.g.ηG1[u, :]]
X1 = complete_polynomial(grid1, deg)
S1[u, :] = coefs[0, :] @ X1 # Compute S(t+1)
F1[u, :] = coefs[1, :] @ X1 # Compute F(t+1)
C1[u, :] = (coefs[2, :] @ X1)**(-1/m.p.γ) # Compute C(t+1)
# Compute next-period π using condition
# (35) in MM (2015)
π1 = ((1-(1-m.p.Θ)*(S1/F1)**(1-m.p.ϵ))/m.p.Θ)**(1/(m.p.ϵ-1))
# Evaluate conditional expectations in the Euler equations
#---------------------------------------------------------
e[0, :] = exp(m.g.ηu)*exp(m.g.ηL)*L0**m.p.ϑ*Y0/exp(m.g.ηa) + m.g.ω_nodes @ (m.p.β*m.p.Θ*π1**m.p.ϵ*S1)
e[1, :] = exp(m.g.ηu)*C0**(-m.p.γ)*Y0 + m.g.ω_nodes @ (m.p.β*m.p.Θ*π1**(m.p.ϵ-1)*F1)
e[2, :] = m.p.β*exp(m.g.ηB)/exp(m.g.ηu)*R1 * (m.g.ω_nodes @ ((exp(m.g.ηu1)*C1**(-m.p.γ)/π1)))
# Variables of the current iteration
#-----------------------------------
np.copyto(S0_new_G, S0)
np.copyto(F0_new_G, F0)
np.copyto(C0_new_G, C0)
# Compute and update the coefficients of the decision functions
# -------------------------------------------------------------
coefs_hat = np.linalg.lstsq(X0_G.T, e.T)[0].T
# Update the coefficients using damping
coefs = damp*coefs_hat + (1-damp)*coefs
# Evaluate the percentage (unit-free) difference between the values
# on the grid from the previous and current iterations
# -----------------------------------------------------------------
# The convergence criterion is adjusted to the damping parameters
err = (np.mean(abs(1-S0_new_G/S0_old_G)) +
np.mean(abs(1-F0_new_G/F0_old_G)) +
np.mean(abs(1-C0_new_G/C0_old_G)))
# Store the obtained values for S(t), F(t), C(t) on the grid to
# be used on the subsequent iteration in Section 10.2.6
#-----------------------------------------------------------------------
np.copyto(S0_old_G, S0_new_G)
np.copyto(F0_old_G, F0_new_G)
np.copyto(C0_old_G, C0_new_G)
return coefs
def simulate(self, coefs=None, capT=10201):
if coefs is None:
coefs = self.solve()
# rename self to m to make code below readable
m = self
# create namedtuple to hold simulation results in an organized container
Simulation = namedtuple("Simulation",
["nuR", "nua", "nuL", "nuu", "nuB", "nuG",
"δ", "R", "S", "F", "C", "π", "Y", "L", "Yn", "w"])
# 11. Simualating a time-series solution
#---------------------------------------
# Initialize the values of 6 exogenous shocks and draw innovations
#-----------------------------------------------------------------
nuR = np.zeros(capT)
nua = np.zeros(capT)
nuL = np.zeros(capT)
nuu = np.zeros(capT)
nuB = np.zeros(capT)
nuG = np.zeros(capT)
# Generate the series for shocks
#-------------------------------
rands = np.random.randn(capT-1, 6)
for t in range(capT-1):
nuR[t+1] = self.p.ρηR*nuR[t] + self.p.σηR*rands[t, 0]
nua[t+1] = self.p.ρηa*nua[t] + self.p.σηa*rands[t, 1]
nuL[t+1] = self.p.ρηL*nuL[t] + self.p.σηL*rands[t, 2]
nuu[t+1] = self.p.ρηu*nuu[t] + self.p.σηu*rands[t, 3]
nuB[t+1] = self.p.ρηB*nuB[t] + self.p.σηB*rands[t, 4]
nuG[t+1] = self.p.ρηG*nuG[t] + self.p.σηG*rands[t, 5]
δ = np.ones(capT+1) # Allocate memory for the time series of delta(t)
R = np.ones(capT+1) # Allocate memory for the time series of R(t)
S = np.ones(capT) # Allocate memory for the time series of S(t)
F = np.ones(capT) # Allocate memory for the time series of F(t)
C = np.ones(capT) # Allocate memory for the time series of C(t)
π = np.ones(capT) # Allocate memory for the time series of π(t)
Y = np.ones(capT) # Allocate memory for the time series of Y(t)
L = np.ones(capT) # Allocate memory for the time series of L(t)
Yn = np.ones(capT) # Allocate memory for the time series of Yn(t)
w = np.ones(capT)
pol_bases = np.empty(coefs.shape[1])
states = np.empty(8)
for t in range(capT):
states[0] = math.log(R[t]); states[1] = math.log(δ[t])
states[2] = nuR[t]; states[3] = nua[t]
states[4] = nuL[t]; states[5] = nuu[t]
states[6] = nuB[t]; states[7] = nuG[t]
_complete_poly_impl_vec(states, self.p.degree, pol_bases)
vals = coefs @ pol_bases
S[t] = vals[0]
F[t] = vals[1]
C[t] = (vals[2])**(-1/m.p.γ)
π[t], δ[t+1], Y[t], L[t], Yn[t], R[t+1] = self.step(S[t], F[t], C[t],
δ[t], R[t], nuG[t],
nua[t], nuL[t],
nuR[t])
# Compute real wage
w[t] = exp(nuL[t])*(L[t]**m.p.ϑ)*(C[t]**m.p.γ)
# If ZLB is imposed, set R(t)=1 if ZLB binds
if self.p.zlb:
R[t+1] = max(R[t+1], 1.0)
return Simulation(nuR, nua, nuL, nuu, nuB, nuG, δ, R, S, F, C, π, Y, L, Yn, w)
def residuals(self, coefs, sim, burn=200):
m = self # rename self to m so the rest of this code is more readable
capT = len(sim.w)
resids = np.zeros((capT, 9))
# Integration method for evaluating accuracy
# ------------------------------------------
# Monomial integration rule with 2N**2+1 nodes
ϵ_nodes, ω_nodes = qnwmonomial2(m.p.vcov)
n_nodes = len(ω_nodes)
# Allocate for arrays needed in the loop
basis_mat = np.empty((8, n_nodes))
X1 = np.empty((coefs.shape[1], n_nodes))
nuR1 = np.empty(n_nodes)
nua1 = np.empty(n_nodes)
nuL1 = np.empty(n_nodes)
nuu1 = np.empty(n_nodes)
nuB1 = np.empty(n_nodes)
nuG1 = np.empty(n_nodes)
for t in range(capT): # For each given point,
# Take the corresponding value for shocks at t
#---------------------------------------------
nuR0 = sim.nuR[t] # nuR(t)
nua0 = sim.nua[t] # nua(t)
nuL0 = sim.nuL[t] # nuL(t)
nuu0 = sim.nuu[t] # nuu(t)
nuB0 = sim.nuB[t] # nuB(t)
nuG0 = sim.nuG[t] # nuG(t)
# Exctract time t values for all other variables (and t+1 for R, δ)
#------------------------------------------------------------------
R0 = sim.R[t] # R(t-1)
δ0 = sim.δ[t] # δ(t-1)
R1 = sim.R[t+1] # R(t)
δ1 = sim.δ[t+1] # δ(t)
L0 = sim.L[t] # L(t)
Y0 = sim.Y[t] # Y(t)
Yn0 = sim.Yn[t] # Yn(t)
π0 = sim.π[t] # π(t)
S0 = sim.S[t] # S(t)
F0 = sim.F[t] # F(t)
C0 = sim.C[t] # C(t)
# Fill basis matrix with R1, δ1 and shocks
#-----------------------------------------
# Note that we do not premultiply by standard deviations as ϵ_nodes
# already include them. All these variables are vectors of length n_nodes
nuR1[:] = nuR0*m.p.ρηR + ϵ_nodes[:, 0]
nua1[:] = nua0*m.p.ρηa + ϵ_nodes[:, 1]
nuL1[:] = nuL0*m.p.ρηL + ϵ_nodes[:, 2]
nuu1[:] = nuu0*m.p.ρηu + ϵ_nodes[:, 3]
nuB1[:] = nuB0*m.p.ρηB + ϵ_nodes[:, 4]
nuG1[:] = nuG0*m.p.ρηG + ϵ_nodes[:, 5]
basis_mat[0, :] = np.log(R1)
basis_mat[1, :] = np.log(δ1)
basis_mat[2, :] = nuR1
basis_mat[3, :] = nua1
basis_mat[4, :] = nuL1
basis_mat[5, :] = nuu1
basis_mat[6, :] = nuB1
basis_mat[7, :] = nuG1
# Future choices at t+1
#----------------------
# Form a complete polynomial of degree "Degree" (at t+1) on future state
# variables; n_nodes-by-npol
_complete_poly_impl(basis_mat, self.p.degree, X1)
# Compute S(t+1), F(t+1) and C(t+1) in all nodes using coefs
S1 = coefs[0, :] @ X1
F1 = coefs[1, :] @ X1
C1 = (coefs[2, :] @ X1)**(-1/m.p.γ)
# Compute π(t+1) using condition (35) in MM (2015)
π1 = ((1-(1-m.p.Θ)*(S1/F1)**(1-m.p.ϵ))/m.p.Θ)**(1/(m.p.ϵ-1))
# Compute residuals for each of the 9 equilibrium conditions
#-----------------------------------------------------------
resids[t, 0] = 1-(ω_nodes @
(exp(nuu0)*exp(nuL0)*L0**m.p.ϑ*Y0/exp(nua0) +
m.p.β*m.p.Θ*π1**m.p.ϵ*S1)/S0
)
resids[t, 1] = 1 - (ω_nodes @
(exp(nuu0)*C0**(-m.p.γ)*Y0 + m.p.β*m.p.Θ*π1**(m.p.ϵ-1)*F1)/F0
)
resids[t, 2] = 1.0 -(ω_nodes @
(m.p.β*exp(nuB0)/exp(nuu0)*R1*exp(nuu1)*C1**(-m.p.γ)/π1)/C0**(-m.p.γ)
)
resids[t, 3] = 1-((1-m.p.Θ*π0**(m.p.ϵ-1))/(1-m.p.Θ))**(1/(1-m.p.ϵ))*F0/S0
resids[t, 4] = 1-((1-m.p.Θ)*((1-m.p.Θ*π0**(m.p.ϵ-1))/(1-m.p.Θ))**(m.p.ϵ/(m.p.ϵ-1)) + m.p.Θ*π0**m.p.ϵ/δ0)**(-1)/δ1
resids[t, 5] = 1-exp(nua0)*L0*δ1/Y0
resids[t, 6] = 1-(1-m.p.gbar/exp(nuG0))*Y0/C0
resids[t, 7] = 1-(exp(nua0)**(1+m.p.ϑ)*(1-m.p.gbar/exp(nuG0))**(-m.p.γ)/exp(nuL0))**(1/(m.p.ϑ+m.p.γ))/Yn0
resids[t, 8] = 1-m.s.π/m.p.β*(R0*m.p.β/m.s.π)**m.p.μ*((π0/m.s.π)**m.p.ϕ_π * (Y0/Yn0)**m.p.ϕ_y)**(1-m.p.μ)*exp(nuR0)/R1 # Taylor rule
# If the ZLB is imposed and R>1, the residuals in the Taylor rule (the
# 9th equation) are zero
if m.p.zlb and R1 <= 1:
resids[t, 8] = 0.0
return resids[burn:, :]
# + [markdown] outputExpanded=false
# ## Running the code
#
# Now that we've done all the hard work to define the model, its solution and
# simulation, and accuracy checks, let's put things together and run the code!
# + outputExpanded=false
def main(m=Model(), file=None):
if file is None:
mprint = print
else:
def mprint(*x):
print(*x, file=file)
# solve the model
t1 = time.time()
coefs = m.solve()
solve_time = time.time() - t1
# simulate the model
t1 = time.time()
sim = m.simulate(coefs)
sim_time = time.time() - t1
# check accuracy
t1 = time.time()
resids = m.residuals(coefs, sim)
resids_time = time.time() - t1
tot_time = solve_time + sim_time + resids_time
mean_err = np.log10(abs(resids).mean())
max_err = np.log10(abs(resids).max())
max_err_eqn = np.log10(abs(resids).max(1) + 1e-16)
l1 = np.log10(abs(resids).max(0).sum())
mprint("Solver time (in seconds): ", solve_time)
mprint("Simulation time (in seconds): ", sim_time)
mprint("Residuals time (in seconds): ", resids_time)
mprint("Total time (in seconds): ", tot_time)
mprint("\nAPPROXIMATION ERRORS (log10):")
mprint("\ta) mean error in the model equations: {:0.3f}".format(mean_err))
mprint("\tb) sum of max error per equation: {:0.3f}".format(l1));
mprint("\tc) max error in the model equations: {:0.3f}".format(max_err))
mprint("\td) max error by equation: ", max_err_eqn)
mprint("tex row:", "{:.2f} & {:.2f} & {:.2f}".format(l1, max_err, tot_time))
return solve_time, sim_time, resids_time, coefs, sim, resids
def build_paper_table():
with open("output.log", "w") as f:
for params in (dict(πstar=1.0, σηL=0.1821, zlb=False),
dict(πstar=1.0, σηL=0.4054, zlb=False),
dict(πstar=1.0, σηL=0.1821, zlb=True)):
for grid_kind in ["sobol", "random"]:
p = Params(**params)
g = Grids(p, kind=grid_kind)
m = Model(p, g)
print("working with params:", params, file=f)
print("And grid type:", grid_kind, file=f)
main(m, f)
print("\n"*5, file=f)
# -
build_paper_table()
main();
| quanteconomics/NKModel_CLMM_python.ipynb |