code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 110} colab_type="code" id="3v26sVq_4wiG" outputId="7b414a67-10b0-4e63-ac0c-06891d7b65ee"
import os
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from keras.models import load_model
from keras.preprocessing import image
loaded_model = tf.keras.models.load_model('model/firesmoke_model.h5')
loaded_model.layers[0].input_shape #(None, 150, 150, 3)
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="aB8H3KOs7vF2" outputId="93ba383d-4056-4e48-febe-ff4df7d939ce"
new_model=load_model('model/firesmoke_model.h5')
fire='data/test/fire/image_0.jpg'
smoke='data/test/smoke/122.jpg'
fire=image.load_img(fire,target_size=(150,150))
smoke=image.load_img(smoke,target_size=(150,150))
fire=image.img_to_array(fire)
fire=image.img_to_array(fire)
#-------------------------------------------
fire=np.expand_dims(fire,axis=0)
smoke=np.expand_dims(smoke,axis=0)
fire=fire/255
smoke=smoke/255
print(fire.shape)
print(smoke.shape)
a=classifier.predict_classes(fire)
b=classifier.predict_classes(smoke)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="tgAgZdGR-4zY" outputId="69d05b6b-cbe1-4030-87b7-7431cc798e9a"
print(a)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="vNBWxV3b_kNc" outputId="2d5089f6-4eca-4634-d5ed-25f609d0e7ab"
print(b)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} colab_type="code" id="mbpFZw6x_l-5" outputId="1a0391ae-d613-4d1d-f2f8-dacdd48ad097"
a=classifier.predict_classes(fire)
b=classifier.predict_classes(smoke)
if a==0:
print('Fire Detect')
else:
print('Smoke Detect')
# + colab={"base_uri": "https://localhost:8080/", "height": 269} colab_type="code" id="uVxEnnGPAZZb" outputId="e6357573-557d-45b6-9389-5b667a34d841"
import cv2
img=cv2.imread('data/test/fire/image_0.jpg',1)
img=cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
_file="data/test/fire/image_0.jpg"
images=image.load_img(_file,target_size=(150,150))
images=image.img_to_array(images)
images=np.expand_dims(images,axis=0)
images=images/255
test_model=load_model('model/firesmoke_model.h5')
prediction=test_model.predict_classes(images)
if prediction==0:
img =cv2.putText(img=np.copy(img), text="Fire", org=(10,20),fontFace=2, fontScale=0.75, color=(0,0,255), thickness=1)
elif prediction==1:
img =cv2.putText(img=np.copy(img), text="Smoke", org=(10,30),fontFace=2, fontScale=0.75, color=(0,0,255), thickness=1)
plt.imshow(img)
plt.show()
# + colab={"base_uri": "https://localhost:8080/", "height": 261} colab_type="code" id="46NCHDgFAtRX" outputId="0481a6b5-8166-4805-f495-90aa50737b9b"
import cv2
img=cv2.imread('data/test/smoke/122.jpg',1)
img=cv2.cvtColor(img,cv2.COLOR_BGR2RGB)
_file="data/test/smoke/122.jpg"
images=image.load_img(_file,target_size=(150,150))
images=image.img_to_array(images)
images=np.expand_dims(images,axis=0)
images=images/255
test_model=load_model('model/firesmoke_model.h5')
prediction=test_model.predict_classes(images)
if prediction==0:
img =cv2.putText(img=np.copy(img), text="Fire", org=(10,20),fontFace=2, fontScale=0.75, color=(0,0,255), thickness=1)
elif prediction==1:
img =cv2.putText(img=np.copy(img), text="Smoke", org=(10,30),fontFace=2, fontScale=0.75, color=(0,0,255), thickness=1)
plt.imshow(img)
plt.show()
# + colab={} colab_type="code" id="pZoL_bsnBcUo"
|
Keras_model/firedetection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Loading the MNIST data
#
# This is the MNIST data obtainable at http://yann.lecun.com/exdb/mnist/
#
# The data is supplied as IDX files compressed in gzip format. The code below unzips the data, converts the IDX file to an ndarray, reshapes and one-hot encodes as necessary, scales the data and finally pickles the data for easy loading into the main script.
#
# It's worth noting that the pickled data files are not backward compatible with Python 2.X, so if you haven't yet started using Python 3.X then you should download the gzips yourself and run this script locally to generate Python 2.X compatible pickle files. YMMV.
#
# Finally, the details of the data are available on the website above. But in a nutshell, the training data contains 60 000 images, and the testing data contains 10 000 images. I randomly removed 10 000 of the training data points to set aside as a validation set.
import pickle
import gzip
import idx2numpy
import numpy as np
from sklearn.cross_validation import train_test_split
from sklearn.preprocessing import MinMaxScaler
# +
# Uncompress the gzips and convert the IDX files to ndarray
with gzip.open('data/gzips/train-images-idx3-ubyte.gz', 'rb') as f:
xtrain = idx2numpy.convert_from_file(f)
with gzip.open('data/gzips/train-labels-idx1-ubyte.gz', 'rb') as f:
ytrain = idx2numpy.convert_from_file(f)
# Reshape the images to an [nXm] array
xtrain = xtrain.reshape(len(xtrain),-1)
xtrain = MinMaxScaler().fit_transform(xtrain)
# One-hot encode the y values
ytrain = np.eye(10)[ytrain].reshape(len(ytrain),10)
# Seperate out the validation set. Note: the random_state parameter will ensure you get the same results as me.
xtrain, xval, ytrain, yval = train_test_split(xtrain, ytrain, test_size=10000, random_state=0)
# Write the pickled files for importing easily into other scripts
with open('data/pickled/xtrain.pickle', 'wb') as f:
pickle.dump(xtrain, f, pickle.HIGHEST_PROTOCOL)
with open('data/pickled/xval.pickle', 'wb') as f:
pickle.dump(xval, f, pickle.HIGHEST_PROTOCOL)
with open('data/pickled/ytrain.pickle', 'wb') as f:
pickle.dump(ytrain, f, pickle.HIGHEST_PROTOCOL)
with open('data/pickled/yval.pickle', 'wb') as f:
pickle.dump(yval, f, pickle.HIGHEST_PROTOCOL)
# +
# As above, but for the test set
with gzip.open('data/gzips/t10k-images-idx3-ubyte.gz', 'rb') as f:
xtest = idx2numpy.convert_from_file(f)
with gzip.open('data/gzips/t10k-labels-idx1-ubyte.gz', 'rb') as f:
ytest = idx2numpy.convert_from_file(f)
xtest = xtest.reshape(len(xtest),-1)
xtest = MinMaxScaler().fit_transform(xtest)
ytest = np.eye(10)[ytest].reshape(len(ytest),10)
with open('data/pickled/xtest.pickle', 'wb') as f:
pickle.dump(xtest, f, pickle.HIGHEST_PROTOCOL)
with open('data/pickled/ytest.pickle', 'wb') as f:
pickle.dump(ytest, f, pickle.HIGHEST_PROTOCOL)
|
nn-from-scratch/MNIST-loader.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# View more python learning tutorial on my Youtube and Youku channel!!!
# Youtube video tutorial: https://www.youtube.com/channel/UCdyjiB5H8Pu7aDTNVXTTpcg
# Youku video tutorial: http://i.youku.com/pythontutorial
import tkinter as tk
window = tk.Tk()
window.title('my window')
window.geometry('200x200')
l = tk.Label(window, bg='yellow', width=20, text='empty')
l.pack()
def print_selection(v):
l.config(text='you have selected ' + v)
s = tk.Scale(window, label='try me', from_=5, to=11, orient=tk.HORIZONTAL,
length=200, showvalue=0, tickinterval=2, resolution=0.01, command=print_selection)
s.pack()
window.mainloop()
# + pycharm={"name": "#%%\n"}
|
tkinterTUT/tk6_scale.py.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python (ml_ebook)
# language: python
# name: ml_ebook
# ---
# In this assignment, you are required to implement functions to calculate the listed performance measures below from scratch. **YOU CAN USE NUMPY FOR CALCULATIONS AND MATPLOTLIB FOR VISUALIZATION. DON'T USE ANY OTHER PACKAGE**.
#
# * Accuracy
# * Confusion matrix
# * Percision
# * Recall
# * F1 score
# * AUC of ROC
#
#
# You're also required to implement functions to display the following
# * Confusion matrix
# * Percision VS Recall curve
# * ROC curve
#
# After finishing the assignment, please run all cells then save the notebook as PDF and attach the PDF to the original notebook in the email.
# +
# DON'T CHANGE THIS
# DON'T
import numpy as np
np.random.seed(42)
y = np.random.randint(2, size=(100, 1)) # Generating random labels
y_preds = np.random.uniform(0, 1, size=(100, 1)) # Generating random predictions
# +
# DON'T CHANGE THIS
import matplotlib.pyplot as plt
plt.plot(y)
plt.plot(y_preds)
# -
# You can start your implementation below.
# <h4>Accuracy</h4>
# +
y_preds_rounded = y_preds.round()
def accuracy(y_preds_rounded, y):
acc = (y == y_preds_rounded).sum() / len(y)
return acc
print(f"Accuracy is: {accuracy(y_preds_rounded, y)}")
# -
# <br>
# <h4>Confusion matrix</h4>
# +
def confusion_matrix(y_preds_rounded, y):
num_classes = len(np.unique(y))
result = np.zeros((num_classes, num_classes))
for i in range(len(y)):
result[int(y[i])][int(y_preds_rounded[i])] += 1
return result
conf_matrix = confusion_matrix(y_preds_rounded, y)
print(f"Confusion Matrix: \n{conf_matrix}")
# +
def plot_confusion_matrix(conf_matrix, title='Confusion matrix'):
fig, ax = plt.subplots(figsize=(5, 5))
ax.matshow(conf_matrix, cmap=plt.cm.BuPu, alpha=0.4)
for i in range(conf_matrix.shape[0]):
for j in range(conf_matrix.shape[1]):
ax.text(x=j, y=i, s=conf_matrix[i, j],
va='center', ha='center',
size='xx-large')
plt.xlabel('Predictions', fontsize=18)
plt.ylabel('Actuals', fontsize=18)
plt.title(title, fontsize=18)
plt.show()
plot_confusion_matrix(conf_matrix)
# -
# <br>
# <h4>3. Percision</h4>
# +
# True Positive --> 1,1
# False Positive --> 1,0
def precision(y_preds_rounded, y):
TP = ((y_preds_rounded == 1) & (y == 1)).sum()
FP = ((y_preds_rounded == 1) & (y == 0)).sum()
precision = TP / (TP + FP)
return precision
precision_val = precision(y_preds_rounded, y)
print(f"Precision is: {precision_val}")
# -
# <h4>4. Recall</h4>
# +
# True Positive --> 1,1
# False Negative --> 0,1
def recall(y_preds_rounded, y):
TP = ((y_preds_rounded == 1) & (y == 1)).sum()
FN = ((y_preds_rounded == 0) & (y == 1)).sum()
recall = (TP) / (TP + FN)
return recall
recall_val = recall(y_preds_rounded, y)
print(f"Recall is: {recall_val}")
# -
# <br>
# <h5>Percision VS Recall curve </h5>
# +
def precision_recall_curve(y_preds_rounded, y):
thresholds = np.arange(0, 1, 0.1)
precisons = []
recalls = []
for t in thresholds:
y_preds_rounded = np.array([1 if i >= t else 0 for i in y_preds])
y_preds_rounded = y_preds_rounded.reshape(y.shape)
TP = ((y_preds_rounded == 1) & (y == 1)).sum()
FN = ((y_preds_rounded == 0) & (y == 1)).sum()
TN = ((y_preds_rounded == 0) & (y == 0)).sum()
FP = ((y_preds_rounded == 1) & (y == 0)).sum()
precisons.append(TP / (TP+FP))
recalls.append (TP / (TP+FN))
plt.plot(recalls, precisons, color='purple')
precision_recall_curve(y_preds_rounded, y)
# -
# <br>
# <h4>5. F1 Score</h4>
# +
def f1_score(precision, recall):
score = 2 * (precision * recall) / (precision + recall)
return score
score = f1_score(precision_val, recall_val)
print(f"F1_Score: {score}")
# -
# <h4>6. AUC of ROC</h4>
# +
def auc_roc(y_preds_rounded, y):
TP = ((y_preds_rounded == 1) & (y == 1)).sum()
FN = ((y_preds_rounded == 0) & (y == 1)).sum()
TN = ((y_preds_rounded == 0) & (y == 0)).sum()
FP = ((y_preds_rounded == 1) & (y == 0)).sum()
FPR = FP / ( FP + TN )
TPR = TP / ( TP + FN )
roc = (1+TPR-FPR) / 2
return roc
auc_roc_val = auc_roc(y_preds_rounded, y)
print(f"AUC_ROC: {auc_roc_val}")
# +
def roc_curve(y_preds_rounded, y):
thresholds = np.arange(0, 1, 0.1)
TPR_lst = []
FPR_lst = []
for t in thresholds:
y_preds_rounded = np.array([1 if i >= t else 0 for i in y_preds])
y_preds_rounded = y_preds_rounded.reshape(y.shape)
TP = ((y_preds_rounded == 1) & (y == 1)).sum()
FN = ((y_preds_rounded == 0) & (y == 1)).sum()
TN = ((y_preds_rounded == 0) & (y == 0)).sum()
FP = ((y_preds_rounded == 1) & (y == 0)).sum()
TPR_lst.append(TP / (TP+FN))
FPR_lst.append(FP / (FP+TN))
plt.plot([1,0], [1,0], linestyle='--', color='grey')
plt.plot(FPR_lst, TPR_lst)
roc_curve(y_preds_rounded, y)
|
exercises/E1 - Classification Performance Measures.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Pairs Trade Algorithm
# <NAME>
# ***
# The pairs trade is a market neutral trading strategy enabling traders to profit from virtually any market conditions: uptrend, downtrend, or sideways movement. This strategy is categorized as a statistical arbitrage and convergence trading strategy. The pair trading was pioneered by <NAME> and later led by <NAME>’s quantitative group at <NAME> in the 1980s.[1](https://en.wikipedia.org/wiki/Pairs_trade)
# This notebook will introduce my implementation of pairs trade. We will be switching gears from the tech stocks to pairs of stocks I found to have similar overall trend.
# +
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.dates import YearLocator, MonthLocator, DateFormatter
import sys
sys.path.append('./src/')
from random_walk_forcast import *
# -
# %matplotlib inline
sns.set_style("whitegrid", {'axes.edgecolor': '.6',
'axes.facecolor': '0.9',
'grid.color': '.82',
'legend.frameon': True,
'axes.labelsize': 'small'})
pairs = pd.read_csv('data/pairs_bundle.csv', index_col=0)
pairs.index = pd.to_datetime(pairs.index)
def plot_pair(series1, series2, names):
"""
Plot pair series with name subsitutions
Args: series1 (ndarray)---first stock of the pair
series2 (ndarray)---second stock of the pair
names (list)--------list of names for the stocks
Returns: (None) plots inline
"""
fig, ax = plt.subplots(figsize=(20,8));
years = YearLocator();
yearsFmt = DateFormatter('%Y');
ax.xaxis.set_major_locator(years);
ax.xaxis.set_major_formatter(yearsFmt);
ax.autoscale_view();
index = series1.index
plt.title(names[0] + ' and ' +
names[1] +' (Adj. Close)', fontsize=20);
plt.ylabel('Adj. Close', fontsize=15);
plt.xlabel('Time', fontsize=15);
ax.plot_date(index, series1, 'indianred', label=names[0]);
ax.plot_date(index, series2, 'steelblue', label=names[1]);
plt.legend(loc=2, prop={'size':15}, frameon=True);
# Notice that the long term trends for these pairs of stocks (Verizon, At&t) and (Chrevron and Exxon) are very similar. We can assume that devations are due to random market fluxuations. With this assumption that the stock prices will converge eventually we can make a long term profit.
# The high level explanation of the algorithm, as I understand it, is when stocks move away from their long term trend sell shares of the stock thats increasing (being temporarily overvalued) and use those earnings to buy shares of the stock thats decreasing (being temporarily undervalued). This is called 'opening a position'. Then when the stocks converge, close your position, sell the stock you bought at a lower price and buy the stock you sold at a higher price. This assumes you own some shares of both these stocks throughout the process.
# The following algorithm will ignore keeping track of number of stocks at any given time. Meaning there may be situations where we sell stocks we don't own. This actually is not an issue because you can short stocks (instead of selling them) if you suspect the price will go down. Without getting into much of the finance I will move forward.
plot_pair(pairs['VZ'], pairs['T'], ['Verizon','At&t'])
plot_pair(pairs['CVX'], pairs['XOM'], ['Chevron','Exxon'])
# We will be working with the ratios of these pairs:
def plot_ratio(ratio, name, deviations=[1], positions=[]):
"""
Plots the ratio of the stocks
Args: ratio (ndarray)----the ratio of the stocks in question
name (string)------the name of the ratio
devations (list)---the devations to plot
positions (list)---the positions to plot
Returns: (None) plots inplace
"""
fig = plt.subplots(figsize=(20,8));
plt.title('Ratio ' + name + ' Adjusted Close', fontsize=20);
plt.ylabel('Ratio', fontsize=15);
plt.xlabel('Time Index', fontsize=15);
plt.xlim([0,ratio.size])
plt.xticks(np.arange(0, ratio.size, 500))
plt.plot(np.arange(ratio.size), ratio, 'black', label='$Ratio$', alpha=0.5);
plt.plot([0, ratio.size], [ratio.mean(), ratio.mean()], 'steelblue', lw=2, label=r'$\hat{\mu}$');
for color, std in zip(['y','orange','salmon','red'], deviations):
latex_prep = '$' + str(std) + '$'
plt.plot([0, ratio.size], [ratio.mean()-std*ratio.std(), ratio.mean()-std*ratio.std()],
'--', lw=2, label='$\hat{\mu} \pm$' + latex_prep + '$\hat{\sigma}$', color=color);
plt.plot([0, ratio.size], [ratio.mean()+std*ratio.std(), ratio.mean()+std*ratio.std()],
'--', lw=2, color=color);
if positions:
opening_days, closing_days = [], []
opening_ratios, closing_ratios = [], []
for position in positions:
if 'open' in position.keys():
for day in position['open']:
opening_days.append(day)
opening_ratios.append(ratio.ix[day])
if 'close' in position.keys():
closing_days.append(position['close'])
closing_ratios.append(ratio.ix[position['close']])
plt.scatter(x=opening_days, y=opening_ratios, s=125, color='lime', edgecolor='black', label='$Open$ '+'$Position$')
plt.scatter(x=closing_days, y=closing_ratios, s=125, color='red', edgecolor='black', label='$Close$ '+'$Position$')
plt.legend(loc='best', prop={'size':15}, frameon=True);
# We see that for the Verizon/ATT ratio there is quite a lot of movement about their average ratio (about 1.5).
plot_ratio(pairs['VZ/T'], name='Verizon/ATT', deviations=[])
# We see that for the Chevron/Exxon ratio there is a little bit of movement about their average ratio (about 1).
plot_ratio(pairs['CVX/XOM'], name='Chevron/Exxon', deviations=[])
def identify_positions(ratio, k, start=50):
"""
Identifies all positions using a dynamic step value
Args: ratio (ndarray)---the ratio of the stocks
k (int)-----------the intial devation to open a position
start (int)-------the day to start
Results: (list of maps) the positions
"""
results = []
result = {"open":[]}
mean, sd = ratio.mean(), ratio.std()
day, size = start, ratio.size
while day < size:
if abs(ratio.ix[day] - mean) > k*sd:
result["open"].append(day)
walk = get_expected_walk(ratio[:day], 20)[1]
day += int(walk.max() / walk.min())
elif abs(ratio.ix[day] - mean) < 0.05 and len(result["open"]) > 0 :
result["close"] = day
results.append(result)
result = {"open":[]}
day += 1
else:
day += 1
if len(result['open']) > 0:
result['close'] = day-1
results.append(result)
return results
# Here we see the Verizon/At&t ratio plotted with an error line at 1 standard deviation. Where open positions are opened in green and closed in red.
positions_cell = identify_positions(pairs['VZ/T'], 1)
plot_ratio(pairs['VZ/T'], 'Verizon, At&t', deviations=[1], positions=positions_cell)
# Now it's time to back test out algorithm and calculate our return on investment if we had traded on it it for the past 25 years.
def back_trade(init_investment, numer_prices, denom_prices, ratio, positions, swap_count=50):
"""
Back trades with the given positions
Args: init_investment (int)----initial total investment
numer_prices (ndarray)---the series of the numerator stock w/ respect to the ratio
denom_prices (ndarry)----the series of the denominator stock w/ respect to the ratio
ratio (ndarray)----------the ratio of the series'
positions (list of maps)-the postions to trade on
swap_count (int)---------the number of stocks to swap at a given open position
Returns: (map) the result object
"""
cur_portfolio_value = init_investment
for position in positions:
if all(ratio[position['open']] > ratio.mean()):
openings = len(position['open'])
cur_portfolio_value += np.sum(swap_count*numer_prices[position['open']])
cur_portfolio_value -= np.sum(swap_count*denom_prices[position['open']])
cur_portfolio_value -= openings*swap_count*numer_prices[position['close']]
cur_portfolio_value += openings*swap_count*denom_prices[position['close']]
elif all(ratio[position['open']] < ratio.mean()):
openings = len(position['open'])
cur_portfolio_value -= np.sum(swap_count*numer_prices[position['open']])
cur_portfolio_value += np.sum(swap_count*denom_prices[position['open']])
cur_portfolio_value += openings*swap_count*numer_prices[position['close']]
cur_portfolio_value -= openings*swap_count*denom_prices[position['close']]
return {'init_investment': init_investment,
'net_gain': cur_portfolio_value - init_investment,
'net_gain/year': (cur_portfolio_value - init_investment) / (len(ratio) / 252) } # 252 trade days / year
# Using these positions we will back trade our algorithm. Using an initial investment of 10000 dollars and swaping 100 shares at each open position we net a total gain of about 15,500 dollars over about 25 years. Averaging out to about an 5% annual return on investment. This fails to beat the S&P500.
back_trade(10000, pairs['VZ'].values, pairs['T'].values, pairs['VZ/T'].values, positions_cell, 100)
# Here we see the Chevron/Exxon ratio plotted with an error line at 1 standard deviation. Where open positions are opened in green and closed in red.
positions_gas = identify_positions(pairs['CVX/XOM'], 1)
plot_ratio(pairs['CVX/XOM'], 'Chevron, Exxon', deviations=[1], positions=positions_gas)
# Using these positions we will back trade our algorithm. Using an initial investment of 10000 dollars and swaping 100 shares at each open position we net a total gain of about 40,000 dollars over about 25 years. Averaging out to about an 15% annual return on investment. This outpreforms the S&P. Of course we have to examine this method on many more stocks to tune our thresholds for oppening and closing positions.
back_trade(10000, pairs['CVX'].values, pairs['XOM'].values, pairs['CVX/XOM'].values, positions_gas, 100)
# ___
from IPython.display import HTML
HTML("""<style>@import "http://fonts.googleapis.com/css?family=Lato|Source+Code+Pro|Montserrat:400,700";@font-face{font-family:"Computer Modern";src:url('http://mirrors.ctan.org/fonts/cm-unicode/fonts/otf/cmunss.otf')}.rendered_html h1,h2,h3,h4,h5,h6,p{font-family:'Computer Modern'}p,ul{font-family:'Computer Modern'}div#notebook-container{-webkit-box-shadow:none;box-shadow:none}h1{font-size:70pt}h2{font-size:50pt}h3{font-size:40pt}h4{font-size:35pt}h5{font-size:30pt}h6{font-size:25pt}.rendered_html p{font-size:11pt;line-height:14pt}.CodeMirror pre{font-family:'Source Code Pro', monospace;font-size:09pt}div.input_area{border:none;background:#f5f5f5}ul{font-size:10.5pt;font-family:'Computer Modern'}ol{font-size:11pt;font-family:'Computer Modern'}.output_png img{display:block;margin-left:auto;margin-right:auto}</style>""")
|
Example-Project/04-PairsTradeAlgorithm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # citare : https://github.com/DantesLegacy/TensorFlow_AudioSet_Example/blob/master/src/get_youtube_dataset_balanced.py
#
# +
import csv, sys
import os
import wave
import contextlib
from __future__ import unicode_literals
import youtube_dl
# +
filename = os.path.abspath(os.path.join(os.getcwd(),os.pardir,os.pardir))+'/data/data_split.csv'
rownum = 0
path = os.path.abspath(os.path.join(os.getcwd(),os.pardir,os.pardir))+'/data/Audio/'
project_path = os.path.abspath(os.path.join(os.getcwd(),os.pardir,os.pardir))+'/src'
# specify the index of files that is downloaded last time (to resume downloading)
# Basically this is a simple work around for this downloader, where I sometimes accidentally close the program or sometimes it just hangs in my environment
last_processed_row = 0
# +
#TEST
id='TEST'
os.chdir(path)
ydl_opts = {
'format': 'bestaudio/best',
#'outtmpl': 'e:/python/downloadedsongs/%(title)s.%(ext)s', # <--- pay attention here
#'download_archive': 'downloaded_songs.txt',
'outtmpl': str(float(2)) + '_' + str(id)+'.%(ext)s',
'postprocessors': [{
'key': 'FFmpegExtractAudio',
'preferredcodec': 'wav',
'preferredquality': '192',
}],
#'logger': MyLogger(),
#'progress_hooks': [my_hook],
}
with youtube_dl.YoutubeDL(ydl_opts) as ydl:
ydl.download(['https://www.youtube.com/watch?v=dP15zlyra3c'])
# -
# +
def youtube_download_os_call(id, start_time, idx):#richiama i comandi del os che chiamano ffmpeg(per convertire il video), # che chaima youtube_dl
ret = os.system('ffmpeg -n -ss ' + start_time +
' -i $(youtube-dl -i -w --extract-audio ' #-w:not overwrite #-i:Continue on download errors #Convert video files to audio-only file
'--audio-format wav --audio-quality 0 '
'--get-url https://www.youtube.com/watch?v=' + id + ')'
' -t 10 ' + path + idx + '_' + id + '.wav') #-t = durata
return ret
'''
def get_wav_file_length(path, idx, id):
sample = project_path + path + idx + '_' + id + '.wav'
with contextlib.closing(wave.open(sample, 'r')) as f:
frames = f.getnframes()
rate = f.getframerate()
length = frames / float(rate)
print(length)
return length
'''
def create_error_file(id, idx):
with open(path + idx + '_' + id + '_ERROR.wav', 'a'):
os.utime(path + idx + '_' + id + '_ERROR.wav', None)
def youtube_downloader(id, start_time, idx):
ret = youtube_download_os_call(id, start_time, idx)
print('ffmpeg -n -ss ' + start_time +
' -i $(youtube-dl -i -w --extract-audio '
'--audio-format wav --audio-quality 0 '
'--get-url https://www.youtube.com/watch?v=' + id + ')'
' -t 10 AudioSet/balanced_train/' + idx + '_' + id + '.wav')
return ret
# +
with open(filename, newline='') as f:
reader = csv.reader(f)
try:
for row in reader:
if rownum <= last_processed_row:
rownum += 1
continue
# Skip the 1 line header
if rownum >= 1:
print(row)
ret = youtube_downloader(row[1], str(float(row[2].lstrip())),
str(rownum - 1))
# If there was an error downloading the file
# This sometimes happens if videos are blocked or taken down
if ret != 0:
create_error_file(row[1], str(rownum - 1))
rownum += 1
except csv.Error as e:
sys.exit('file {}, line {}: {}'.format(filename, reader.line_num, e))
# -
|
src/utils/.ipynb_checkpoints/Audio_downloader-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# - https://storage.googleapis.com/deepmind-media/dqn/DQNNaturePaper.pdf
# - https://ai.intel.com/demystifying-deep-reinforcement-learning/
# - https://danieltakeshi.github.io/2016/11/25/frame-skipping-and-preprocessing-for-deep-q-networks-on-atari-2600-games/
#
# - https://github.com/AndersonJo/dqn-pytorch/blob/master/dqn.py
# - https://github.com/hengyuan-hu/rainbow
# - https://github.com/transedward/pytorch-dqn
import matplotlib.pyplot as plt
import gym
import cv2
import random
import math
import numpy as np
import torch
import torch.nn as nn
import torch.nn.functional as F
import torch.optim as optim
from collections import deque
from collections import namedtuple
import copy
import tqdm
SEED = 1234
CAPACITY = 10_000
BATCH_SIZE = 32
GAME = 'PongNoFrameskip-v4'
N_ACTIONS = gym.make(GAME).action_space.n
LEARNING_START = CAPACITY
UPDATE_FREQ = 1
EPSILON_START = 1.0
EPSILON_END = 0.01
EPSILON_STEPS = 30_000
GAMMA = 0.99
TARGET_UPDATE = 1_000
PRINT_UPDATE = 5_000
torch.manual_seed(SEED)
torch.cuda.manual_seed(SEED)
torch.cuda.manual_seed_all(SEED)
np.random.seed(SEED)
random.seed(SEED)
torch.backends.cudnn.deterministic = True
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
class ReplayMemory:
def __init__(self, capacity, batch_size):
"""
Replay memory that holds examples in the form of (s, a, r, s')
args:
capacity (int): the size of the memory
batch_size (int): size of batches used for training model
"""
self.batch_size = batch_size
self.capacity = capacity
#the memory holds al the (s, a, r, s') pairs
#a deque is first-in-first-out, i.e. when you push an example onto the queue
#and it at maximum capacity, the oldest example is popped off the queue
self.memory = deque(maxlen=self.capacity)
#examples in the queue are saved as Transitions
#makes the code more readable and intuitive when getting examples from the memory
self.Transition = namedtuple('Transition', ('state', 'action', 'reward', 'next_state'))
def push(self, state, action, reward, next_state):
"""
Places an (s, a, r, s') example in the memory
args:
state (np.array): the observation obtained from the environment before the action
action (list[int]): the action taken
reward (list[int]): the reward received from the action taken at the current state
next_state (np.array or None): the observation obtained from the environment after the action,
is None when the state is a terminal state
"""
#convert all to tensors
state = torch.FloatTensor(state)
action = torch.LongTensor([action])
reward = torch.FloatTensor([reward])
if next_state is not None:
next_state = torch.FloatTensor(next_state)
#create a transition
transition = self.Transition(state=state, action=action, reward=reward, next_state=next_state)
#add to the memory
self.memory.append(transition)
def sample(self):
"""
Gets a random sample of n = batch_size examples from the memory
The transition returned contains n of each elements, i.e. a batch_size of 32
means this will return a tuple of (32 states, 32 actions, 32 rewards, 32 next_states)
returns:
Transitions (namedtuple): a tuple of (s, a, r, s'),
"""
#sample batch_size transitions for the memory
transitions = random.sample(self.memory, self.batch_size)
#unzip and then rezip so each element contains batch_size examples
return self.Transition(*(zip(*transitions)))
def __len__(self):
"""
Returns the length of the memory, i.e. number of examples in the memory
returns:
length (int): number of examples in the memory
"""
return len(self.memory)
# +
class NoopResetEnv(gym.Wrapper):
def __init__(self, env, noop_max=30):
"""Sample initial states by taking random number of no-ops on reset.
No-op is assumed to be action 0.
"""
gym.Wrapper.__init__(self, env)
self.noop_max = noop_max
self.override_num_noops = None
self.noop_action = 0
assert env.unwrapped.get_action_meanings()[0] == 'NOOP'
def reset(self, **kwargs):
""" Do no-op action for a number of steps in [1, noop_max]."""
self.env.reset(**kwargs)
if self.override_num_noops is not None:
noops = self.override_num_noops
else:
noops = self.unwrapped.np_random.randint(1, self.noop_max + 1)
assert noops > 0
obs = None
for _ in range(noops):
obs, _, done, _ = self.env.step(self.noop_action)
if done:
obs = self.env.reset(**kwargs)
return obs
def step(self, ac):
return self.env.step(ac)
class MaxAndSkipEnv(gym.Wrapper):
def __init__(self, env, skip=4):
"""Return only every `skip`-th frame"""
gym.Wrapper.__init__(self, env)
# most recent raw observations (for max pooling across time steps)
self._obs_buffer = np.zeros((2,)+env.observation_space.shape, dtype=np.uint8)
self._skip = skip
def reset(self):
return self.env.reset()
def step(self, action):
"""Repeat action, sum reward, and max over last observations."""
total_reward = 0.0
done = None
for i in range(self._skip):
obs, reward, done, info = self.env.step(action)
if i == self._skip - 2: self._obs_buffer[0] = obs
if i == self._skip - 1: self._obs_buffer[1] = obs
total_reward += reward
if done:
break
# Note that the observation on the done=True frame
# doesn't matter
max_frame = self._obs_buffer.max(axis=0)
return max_frame, total_reward, done, info
def reset(self, **kwargs):
return self.env.reset(**kwargs)
class FireResetEnv(gym.Wrapper):
def __init__(self, env):
"""Take action on reset for environments that are fixed until firing."""
gym.Wrapper.__init__(self, env)
assert env.unwrapped.get_action_meanings()[1] == 'FIRE'
assert len(env.unwrapped.get_action_meanings()) >= 3
def reset(self, **kwargs):
self.env.reset(**kwargs)
obs, _, done, _ = self.env.step(1)
if done:
self.env.reset(**kwargs)
obs, _, done, _ = self.env.step(2)
if done:
self.env.reset(**kwargs)
return obs
def step(self, ac):
return self.env.step(ac)
class EpisodicLifeEnv(gym.Wrapper):
def __init__(self, env):
"""Make end-of-life == end-of-episode, but only reset on true game over.
Done by DeepMind for the DQN and co. since it helps value estimation.
"""
gym.Wrapper.__init__(self, env)
self.lives = 0
self.was_real_done = True
def step(self, action):
obs, reward, done, info = self.env.step(action)
self.was_real_done = done
# check current lives, make loss of life terminal,
# then update lives to handle bonus lives
lives = self.env.unwrapped.ale.lives()
if lives < self.lives and lives > 0:
# for Qbert sometimes we stay in lives == 0 condtion for a few frames
# so its important to keep lives > 0, so that we only reset once
# the environment advertises done.
done = True
self.lives = lives
return obs, reward, done, info
def reset(self, **kwargs):
"""Reset only when lives are exhausted.
This way all states are still reachable even though lives are episodic,
and the learner need not know about any of this behind-the-scenes.
"""
if self.was_real_done:
obs = self.env.reset(**kwargs)
else:
# no-op step to advance from terminal/lost life state
obs, _, _, _ = self.env.step(0)
self.lives = self.env.unwrapped.ale.lives()
return obs
class WarpFrame(gym.ObservationWrapper):
def __init__(self, env):
"""Warp frames to 84x84 as done in the Nature paper and later work."""
gym.ObservationWrapper.__init__(self, env)
self.width = 84
self.height = 84
self.observation_space = gym.spaces.Box(low=0, high=255,
shape=(self.height, self.width, 1), dtype=np.uint8)
def observation(self, frame):
frame = cv2.cvtColor(frame, cv2.COLOR_RGB2GRAY)
frame = cv2.resize(frame, (self.width, self.height), interpolation=cv2.INTER_AREA)
return frame[:, :, None]
class ClipRewardEnv(gym.RewardWrapper):
def __init__(self, env):
gym.RewardWrapper.__init__(self, env)
def reward(self, reward):
"""Bin reward to {+1, 0, -1} by its sign."""
return np.sign(reward)
class LazyFrames(object):
def __init__(self, frames):
"""This object ensures that common frames between the observations are only stored once.
It exists purely to optimize memory usage which can be huge for DQN's 1M frames replay
buffers.
This object should only be converted to numpy array before being passed to the model.
You'd not believe how complex the previous solution was."""
self._frames = frames
self._out = None
def _force(self):
if self._out is None:
self._out = np.concatenate(self._frames, axis=2)
self._frames = None
return self._out
def __array__(self, dtype=None):
out = self._force()
if dtype is not None:
out = out.astype(dtype)
return out
def __len__(self):
return len(self._force())
def __getitem__(self, i):
return self._force()[i]
class FrameStack(gym.Wrapper):
def __init__(self, env, k):
"""Stack k last frames.
Returns lazy array, which is much more memory efficient.
See Also
--------
baselines.common.atari_wrappers.LazyFrames
"""
gym.Wrapper.__init__(self, env)
self.k = k
self.frames = deque([], maxlen=k)
shp = env.observation_space.shape
self.observation_space = gym.spaces.Box(low=0, high=255, shape=(shp[0], shp[1], shp[2] * k), dtype=np.uint8)
def reset(self):
ob = self.env.reset()
for _ in range(self.k):
self.frames.append(ob)
return self._get_ob()
def step(self, action):
ob, reward, done, info = self.env.step(action)
self.frames.append(ob)
return self._get_ob(), reward, done, info
def _get_ob(self):
assert len(self.frames) == self.k
return LazyFrames(list(self.frames))
class ImageToPyTorch(gym.ObservationWrapper):
"""
Image shape to num_channels x weight x height
"""
def __init__(self, env):
super(ImageToPyTorch, self).__init__(env)
old_shape = self.observation_space.shape
self.observation_space = gym.spaces.Box(low=0.0, high=1.0, shape=(old_shape[-1], old_shape[0], old_shape[1]), dtype=np.uint8)
def observation(self, observation):
return np.swapaxes(observation, 2, 0)
def wrap_pytorch(env):
return ImageToPyTorch(env)
def make_atari(env_id):
env = gym.make(env_id)
assert 'NoFrameskip' in env.spec.id
env = NoopResetEnv(env, noop_max=30)
env = MaxAndSkipEnv(env, skip=4)
return env
def wrap_deepmind(env, episode_life=True, clip_rewards=True, frame_stack=True, scale=False):
"""Configure environment for DeepMind-style Atari.
"""
if episode_life:
env = EpisodicLifeEnv(env)
#if 'FIRE' in env.unwrapped.get_action_meanings():
#env = FireResetEnv(env)
env = WarpFrame(env)
if scale:
env = ScaledFloatFrame(env)
if clip_rewards:
env = ClipRewardEnv(env)
if frame_stack:
env = FrameStack(env, 4)
return env
# -
class DQN(nn.Module):
def __init__(self, n_actions):
super(DQN, self).__init__()
self.conv1 = nn.Conv2d(4, 32, kernel_size=8, stride=4)
self.conv2 = nn.Conv2d(32, 64, kernel_size=4, stride=2)
self.conv3 = nn.Conv2d(64, 64, kernel_size=3, stride=1)
self.fc1 = nn.Linear(7*7*64, 512)
self.fc2 = nn.Linear(512, n_actions) #actions from from env.action_space.n
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.relu(self.conv2(x))
x = F.relu(self.conv3(x))
x = F.relu(self.fc1(x.view(x.size(0), -1))) #flattens the (N, C, H, W) to (N, C*H*W)
return self.fc2(x)
class Agent:
def __init__(self, env, mem, model, update_freq, learning_start, e_start, e_end, e_steps, gamma, target_update, print_update):
"""
An agent class that handles training the model
args:
mem (ReplayMemory): ReplayMemory object
env (Environment): Environment object
model (nn.Module): PyTorch model
update_freq (int): we only update the model every update_freq steps, 1 means update every step
learning_start (int): we only start updating the model after learning_start steps
e_start (int): initial value of epsilon
e_end (int): minimum value of epsilon
e_steps (int): controls the rate of decay from e_start to e_end
gamma (float): decay rate of rewards
target_update (int): update target model after this many parameter updates
print_update (int): print summary of performance after this many steps
"""
self.env = env
self.mem = mem
self.model = model
self.update_freq = update_freq
self.learning_start = learning_start
self.e_start = e_start
self.e_end = e_end
self.e_steps = e_steps
self.gamma = gamma
self.target_update = target_update
self.print_update = print_update
self.steps = 0 #number of steps taken
self.episodes = 0 #number of episodes
#put model on gpu if available
self.model = model.to(device)
#create target model
#set to evaluation mode to turn off batch-norm/dropout if used
self.target = copy.deepcopy(self.model)
self.target.eval()
#create optimizer
self.optimizer = optim.Adam(self.model.parameters(), lr=1e-4)
def get_epsilon(self):
"""
Calculates the value of epsilon from the current number of steps
returns:
epsilon (float): the probability of doing a random action
"""
epsilon = self.e_end + (self.e_start - self.e_end) * math.exp(-1. * self.steps / self.e_steps)
return epsilon
def get_action(self, state):
"""
Selects action to perform, with probability = epsilon chooses a random action,
else chooses the best predicted action of the model
args:
state (np.array): input state to the model
returns:
action (int): the index of the action
"""
#get value of epsilon
epsilon = self.get_epsilon()
#with probablity of epsilon, pick a random action
if random.random() < epsilon:
action = self.env.action_space.sample()
else:
#with probability of (1 - epsilon) pick predicted value
with torch.no_grad():
state = torch.FloatTensor(state).unsqueeze(0).to(device) #convert to tensor, reshape and add to gpu
Qsa = self.model(state) #pass state through model to get Qsa
action = Qsa.max(1)[1].item() #action is max Qsa value
#make sure the value is an integer
assert isinstance(action, int)
return action
def train(self):
"""
Main training loop of the model
Algorithm:
While true:
- resets environment and gets initial state
While episode is not done:
- selects which action to take
- performs action on environment and receives next state, reward and if the episode has ended
- sums rewards across episode
- pushes the (s, a, r, s') tuple onto the memory
- updates the current state to be the state receives from the environment
- increase total number of steps and steps within episode
- updates model parameters every update_freq steps and only after learning_start steps
- updates target model after target_update parameter updates and only after learning_start steps
- prints summary every print_update steps
- increase number of episodes
- update list of all total episode rewards
"""
training_done = False
reward_per_episode = []
pbar = tqdm.tqdm()
while not training_done:
episode_done = False
episode_reward = 0
episode_steps = 0
#get initial state
state = self.env.reset()
while not episode_done:
pbar.update()
#get action
action = self.get_action(state)
#apply action while skipping frames
next_state, reward, episode_done, info = self.env.step(action)
#sum rewards
episode_reward += reward
#add to memory, if episode has finished set next_state to None
mem.push(state, action, reward, None if episode_done else next_state)
#make next_state the new state
state = next_state
#increase number of steps
self.steps += 1
episode_steps += 1
#update model parameters
if self.steps % self.update_freq == 0 and self.steps > self.learning_start:
loss = self.optimize()
#update target model
if self.steps % (self.target_update*self.update_freq) == 0 and self.steps > self.learning_start:
self.target.load_state_dict(self.model.state_dict())
#print summary
if self.steps % self.print_update == 0:
avg_reward_per_episode = np.mean(reward_per_episode[-10:]) #average reward of last 10 episodes
reward_per_episode = []
print(f'Episodes: {self.episodes}, Steps: {self.steps}, Epsilon: {self.get_epsilon():.2f}, Avg. Reward per Ep: {avg_reward_per_episode:.2f}')
#increase number of episodes
self.episodes += 1
reward_per_episode.append(episode_reward)
def optimize(self):
"""
Update model parameters
Algorithm:
- get a batch of transitions
- find out which next_states are terminal states
-
"""
#get a batch
transitions = mem.sample()
#need to set the Q value of terminal states to 0
#this mask will be 1 for non-terminal next_states and 0 for terminal next_states
non_terminal_mask = torch.ByteTensor(list(map(lambda ns: ns is not None, transitions.next_state)))
#this will be 1 for terminal next_states, and 0 for non-terminal next states
terminal_mask = 1 - non_terminal_mask
#state_batch = (N*C,H,W), where N is batch_size, C is phi_length, H and W state height and width
state_batch = torch.cat(transitions.state).to(device)
#action_batch = (N, 1)
action_batch = torch.cat(transitions.action).unsqueeze(1).to(device)
#reward_batch = (N, 1)
reward_batch = torch.cat(transitions.reward).unsqueeze(1).to(device)
#next_state_batch = (M*C,H,W), where M is number of non_terminal next_state in the batch
non_terminal_next_state_batch = torch.cat([ns for ns in transitions.next_state if ns is not None]).to(device)
#reshape to (N,C,H,W)
state_batch = state_batch.view(mem.batch_size, 4, 84, 84)
#reshape to (V,C,H,W)
non_terminal_next_state_batch = non_terminal_next_state_batch.view(-1, 4, 84, 84)
#get predicted Q values from model
Q_preds = self.model(state_batch)
#get Q values of action taken, shape (N,1)
Q_vals = Q_preds.gather(1, action_batch)
#get Q values from target model
target_pred = self.target(non_terminal_next_state_batch)
#tensor for placing target values
target_vals = torch.zeros(mem.batch_size, 1).to(device)
#fill in target values for non_terminal states
#the terminal states will stay initialized as zeros
target_vals[non_terminal_mask] = target_pred.max(1)[0].unsqueeze(1)
expected_vals = reward_batch + (target_vals * self.gamma)
#calculate loss between Q values and target values
loss = F.smooth_l1_loss(Q_vals, expected_vals.detach())
#zero gradients
self.optimizer.zero_grad()
#calculate gradients
loss.backward()
#clamp gradients
for p in self.model.parameters():
p.grad.data.clamp_(-1, 1)
#update parameters
self.optimizer.step()
return loss.item()
# +
env = make_atari(GAME)
env = wrap_deepmind(env)
env = wrap_pytorch(env)
env.seed(SEED)
mem = ReplayMemory(CAPACITY, BATCH_SIZE)
model = DQN(N_ACTIONS)
agent = Agent(env, mem, model, UPDATE_FREQ, LEARNING_START, EPSILON_START, EPSILON_END, EPSILON_STEPS, GAMMA, TARGET_UPDATE, PRINT_UPDATE)
# -
agent.train()
|
1 - DQN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Jupyter Notebook Title
# ## Contents of Notebook
# Description of contents:
#
# Example Github Project
#
# Author: <NAME>
#
#
# Contact: <EMAIL>, +61 (0) 413208746
# ### Notebook magic commands
# %matplotlib inline
# ### Notebook imports
# +
# all imports should go here
import pandas as pd
import sys
import os
import subprocess
import datetime
import platform
import datetime
import matplotlib.pyplot as plt
# -
# ### Notebook version status
# + language="javascript"
# var kernel = IPython.notebook.kernel;
# var thename = window.document.getElementById("notebook_name").innerHTML;
# var command = "theNotebook = " + "'"+thename+"'";
# kernel.execute(command);
# +
# show info to support reproducability
def python_env_name():
envs = subprocess.check_output('conda env list').splitlines()
# get unicode version of binary subprocess output
envu = [x.decode('ascii') for x in envs]
active_env = list(filter(lambda s: '*' in str(s), envu))[0]
env_name = str(active_env).split()[0]
return env_name
#end python_env_name
print('python version : ' + sys.version)
print('python environment :', python_env_name())
print('pandas version : ' + pd.__version__)
print('current wkg dir: ' + os.getcwd())
print('Notebook name: ' + theNotebook)
print('Notebook run at: ' + str(datetime.datetime.now())+ ' local time')
print('Notebook run at: ' + str(datetime.datetime.utcnow()) + ' UTC')
print('Notebook run on: ' + platform.platform())
# -
# ### get current GIT identification
# +
# GIT_LOCATION is the path of the git executable
GIT_LOCATION = \
'C:\\Users\\donrc\\AppData\\Local\\GitHub\\PortableGit_f02737a78695063deace08e96d5042710d3e32db\\cmd\\git.exe'
# -
from githubid import githubid
githubid.get_repo_version(git_location = GIT_LOCATION)
# +
help(githubid)
# -
print('Hi')
print('Cell created in dev branch')
import githubid.githubid as gid
# +
# gid.get_repo_version?
# -
(v1,v2,v3,v4,v5,v6)=gid.get_repo_version(git_location = GIT_LOCATION)
v2[0:7]
v3[-7:]
v4
# !pytest
gid.get_repo_version('a')
# ### Customizations for notebook
# path to saved figures
FIGURE_PREFIX = '../figures/'
# ### Required notebooks to be run first
from IPython.display import FileLink
FileLink('../develop/a.ipynb')
# ### Display associated webpages (eg source of data)
from IPython.display import IFrame
IFrame("http://www.net-analysis.com", width = 800, height = 200)
# ### Save figures to figures directory
# +
def save_figure(figure_title='TemplateNotebookFigure', figure_prefix='../figures/'):
"""
save_figure: same the current matplotlib.pyplot figure as a jpg file
a file is generated from the supplied title, and the date and time
Inputs:
figure_title: string, incorporated into file name
figure_prefix: string giving relative (or absolute) path the save location
Returns:
True is save OK
False otherwise
Outputs:
prints error messages on exceptions
"""
try:
fname = figure_prefix+figure_title+datetime.datetime.now().strftime("%Y%m%d-%H%M%S")+'.jpg'
plt.savefig(fname)
print(fname+' saved.')
except IOError as err:
print('Unable to save figure - IO Error!')
print("IO error: {0}".format(err))
except OSError as err:
print('Unable to save figure - OS Error!')
print("OS error: {0}".format(err))
except:
print('Unable to save figure - Unexpected Error!')
print("Unexpected error:", sys.exc_info()[0])
raise
#end try
#end save_figure
x =[1,2,3,4,5,6]
y =[2,4,5,2,5,9]
plot2 = sns.barplot(x=x, y=y)
save_figure(figure_title='Test', figure_prefix=FIGURE_PREFIX)
plt.show()
# -
# ### Display images
from IPython.display import Image
Image(filename='../figures/apc.jpg', height = 100, width = 200)
# ## Conclusions / Summary
|
develop/2018-01-31-dc-Github NoteBook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd> Surface Element</dd>
# <dt>Dependencies</dt> <dd>Matplotlib</dd>
# <dt>Backends</dt> <dd><a href='../matplotlib/Surface.ipynb'>Matplotlib</a></dd> <dd><a href='./Surface.ipynb'>Plotly</a></dd>
#
# </dl>
# </div>
import numpy as np
import holoviews as hv
hv.notebook_extension('plotly')
# ``Surface`` is used for a set of gridded points whose associated value dimension represents samples from a continuous surface. ``Surface`` is equivalent to an ``Image`` type and supports all the same data formats, including simply NumPy arrays with associated ``bounds`` and other gridded data formats such as xarray.
#
# Rendering a large can often be quite expensive, using ``rstride`` and ``cstride`` we can draw a coarser surface. We can also control the ``azimuth``, ``elevation`` and ``distance`` as plot options to control the camera angle:
# %%opts Surface [width=500 height=500] (cmap='plasma')
hv.Surface(np.sin(np.linspace(0,100*np.pi*2,10000)).reshape(100,100))
# In addition to a simple surface plots, the matplotlib surface plot also supports other related ``plot_type`` modes including ``'wireframe'`` and ``'contour'`` plots:
# %%opts Surface [width=500 height=500] (cmap='fire')
xs = np.arange(-4, 4, 0.25)
ys = np.arange(-4, 4, 0.25)
X, Y = np.meshgrid(xs, ys)
R = np.sqrt(X**2 + Y**2)
Z = np.sin(R)
surface = hv.Surface((xs, ys, Z))
surface
|
examples/reference/elements/plotly/Surface.ipynb
|
# ## Overview
#
# This notebook will show you how to create and query a table or DataFrame that you uploaded to DBFS. [DBFS](https://docs.databricks.com/user-guide/dbfs-databricks-file-system.html) is a Databricks File System that allows you to store data for querying inside of Databricks. This notebook assumes that you have a file already inside of DBFS that you would like to read from.
#
# This notebook is written in **Python** so the default cell type is Python. However, you can use different languages by using the `%LANGUAGE` syntax. Python, Scala, SQL, and R are all supported.
# +
# File location and type
file_location = "/FileStore/tables/Lockdown_Join_with_normaliztion_factor-ed399.csv"
file_type = "csv"
# CSV options
infer_schema = "false"
first_row_is_header = "false"
delimiter = ","
# The applied options are for CSV files. For other file types, these will be ignored.
df = spark.read.format(file_type) \
.option("inferSchema", infer_schema) \
.option("header", first_row_is_header) \
.option("sep", delimiter) \
.load(file_location)
display(df)
# +
# Create a view or table
temp_table_name = "Lockdown_Join_with_normaliztion_factor-ed399_csv"
df.createOrReplaceTempView(temp_table_name)
# +
# %sql
/* Query the created temp table in a SQL cell */
select * from `Lockdown_Join_with_normaliztion_factor-ed399_csv`
# +
# With this registered as a temp view, it will only be available to this particular notebook. If you'd like other users to be able to query this table, you can also create a table from the DataFrame.
# Once saved, this table will persist across cluster restarts as well as allow various users across different notebooks to query this data.
# To do so, choose your table name and uncomment the bottom line.
permanent_table_name = "Lockdown_Join_with_normaliztion_factor-ed399_csv"
# df.write.format("parquet").saveAsTable(permanent_table_name)
|
Codes/Gathering_Data/2020-06-03 - DBFS Example_AN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mnassar/segfault/blob/main/SegFault_TABLE_VI.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="qiLcbwyKprHd" outputId="e2f56804-19f6-42dd-e943-99845115fff1"
# !pip install foolbox
# + id="oJ_HdoXdqDoL"
import tensorflow.compat.v2 as tf
tf.enable_v2_behavior()
from sklearn.model_selection import train_test_split
from xgboost import XGBClassifier
from sklearn.svm import SVC
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import cross_val_score
from skimage.segmentation import felzenszwalb, slic, quickshift
from skimage.segmentation import mark_boundaries
from skimage.util import img_as_float
import numpy as np
import foolbox as fb
import matplotlib.pyplot as plt
# + colab={"base_uri": "https://localhost:8080/"} id="n03S8j8CqVj0" outputId="2ddeac84-72cd-4c76-efaa-8c611be6c2c3"
# !wget --no-check-certificate 'https://docs.google.com/uc?export=download&id=1H4KEE0Vp8DFZOe_QfcxqOxEVnpun-uka' -O CIFAR10model.h5
# + colab={"base_uri": "https://localhost:8080/"} id="dOedPFqNqW78" outputId="9bbbf135-f916-40d8-99cc-0540b0edaefd"
# load the cifar classifier
from tensorflow.keras.models import load_model
pretrained_model = load_model('CIFAR10model.h5')
pretrained_model.trainable = False
pretrained_model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 818, "referenced_widgets": ["2e92b96c6620458b82e2beda61395ee2", "<KEY>", "<KEY>", "7b2a7ade2b0243bfbaed0911a57dd619", "<KEY>", "50c8d6de07ec46df8366dd192f066e1a", "<KEY>", "<KEY>", "dcaa704771c34edb8eef94936da4e161", "<KEY>", "1ce5b69bf4824722bd79c70fd6ebb1fb", "832041fe39bd4ea08829b8b58d6dcade", "<KEY>", "<KEY>", "<KEY>", "9b4ad833574c4b85b46c06a5951c18a5", "1142dbdd09154d078cea3b2bdc8a9204", "<KEY>", "78438bb10e3c4262be164c420d18afaa", "7d8e2a378aa346e1a94d318909e43793", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "891efea2ff7841fba9ab519feb45ae54", "fe476c5d55934f27a55ee1fe84ea008c", "97ec02037d3f4abe938e97cf31fe6d7d", "faea4ffa2d154d19a5873acb97817ad4", "cf00e7eb8efe4727be5b6015e43956b4", "<KEY>", "<KEY>", "<KEY>", "d44d0f0b80b84546a25b63a86da9e899", "42f7ae624d9845fb839b672b452e9ae2", "<KEY>", "<KEY>", "b70f6807a5094de1b89e6ac3d6565a11", "<KEY>", "742526288acb43dbb88552e1565ea00c", "<KEY>", "fdbc18f5102c4c0e9299f05e63843a45", "<KEY>", "<KEY>", "<KEY>", "507bd2712ce64f3a93f182509169d237", "<KEY>", "ef18715ad7864ddd9b3472a8d65d2597", "<KEY>", "<KEY>", "<KEY>", "<KEY>", "9394b9e5759e4219994db1301d941f78", "983fc366707e4669ac3d3568c8544825", "<KEY>", "<KEY>"]} id="mnPscTUZqYe7" outputId="1b8ef1eb-ba25-4216-bff6-32ea591adb41"
import tensorflow_datasets as tfds
(ds_train, ds_test), ds_info = tfds.load(
'cifar10',
split=['train', 'test'],
shuffle_files=True,
as_supervised=True,
with_info=True,
)
print (ds_info)
# + id="y11fRR1wqafL"
# normalize images
# these are the numbers used during training the model
mean = 120.70748
std = 64.150024
bound_min = (0-mean)/std
bound_max = (255-mean)/std
BATCH_SIZE=128
def normalize_img(image, label):
"""Normalizes images: `uint8` -> `float32`."""
# return tf.cast(image, tf.float32) / 255., tf.one_hot(label, 10)
return (tf.cast(image, tf.float32) - mean) / std, tf.one_hot(label, 10)
ds_train = ds_train.map(
normalize_img, num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds_train = ds_train.cache()
ds_train = ds_train.shuffle(ds_info.splits['train'].num_examples)
ds_train = ds_train.batch(BATCH_SIZE)
ds_train = ds_train.prefetch(tf.data.experimental.AUTOTUNE)
ds_test = ds_test.map(
normalize_img, num_parallel_calls=tf.data.experimental.AUTOTUNE)
ds_test = ds_test.batch(BATCH_SIZE)
ds_test = ds_test.cache()
ds_test = ds_test.prefetch(tf.data.experimental.AUTOTUNE)
# + [markdown] id="IAj7dww7eC7F"
# # Dataset Preparation
# + id="M14UT-8LsVsy"
NB_BATCHES = 30 # means that we will have NB_BATCHES radnom normal batches
BATCH_SIZE=128
# and NB_BATCHES random adversarial batches each coming from a different normal batch and a different epsilon
# + [markdown] id="oml7ClnQS_NS"
# Choose Attack
# + id="vfq1TxjsT-Y6" colab={"base_uri": "https://localhost:8080/"} outputId="0e2dea92-d329-41b6-e266-c3ad6368ca4b"
fmodel = fb.models.TensorFlowModel(model=pretrained_model, bounds=(bound_min, bound_max))
#attack = fb.attacks.PGD()
# attack = fb.attacks.L2CarliniWagnerAttack(steps=500)
#attack = fb.attacks.FGSM()
attack = fb.attacks.LinfDeepFoolAttack()
# + id="tDgvhziSrRUj"
ds_experiment = []
ds_experiment_f = []
preds = []
fpreds = []
epsilons = [0.02, 0.06, 0.1]
fmodel = fb.models.TensorFlowModel(model=pretrained_model, bounds=(bound_min, bound_max))
gen = iter(ds_train)
for b in range(NB_BATCHES):
images, labels = gen.next()
preds.append(pretrained_model.predict(images))
ds_experiment.append(images)
labels_class = tf.argmax(labels, axis=1)
# raw, fimages, is_adv = attack(fmodel, images, criterion=fb.criteria.Misclassification(labels_class), epsilons=None) # free epsilon for C&W
raw, fimages, is_adv = attack(fmodel, images, criterion=fb.criteria.Misclassification(labels_class),epsilons=epsilons[b%3])
ds_experiment_f.append(fimages)
fpreds.append(pretrained_model.predict(fimages))
# + colab={"base_uri": "https://localhost:8080/"} id="hK1RAoRKdy8A" outputId="6e8482f2-c90f-4433-c9a1-963e66882be5"
ds_experiment += ds_experiment_f
len(ds_experiment)
# + id="Y0yxiAiUd0GX"
from keras import backend as K
# pretrained_model.summary()
NB_LAYERS=20
NB_NODES_PER_LAYER=200
# + colab={"base_uri": "https://localhost:8080/"} id="0NCakIyCeBKV" outputId="9b31a90e-b0fd-46a3-afec-1a33089c4c40"
inp = pretrained_model.input
layers_= pretrained_model.layers[-NB_LAYERS:]
outputs = [lay.output for lay in layers_]
intermediate_model = K.function([inp], outputs)
# select NB_NODES_PER_LAYER random nodes from each selected layer
print ("these nodes will be used to compute the IQR-"+str(NB_LAYERS*NB_NODES_PER_LAYER))
node_indices=[]
for lay in layers_[:-1]:
# we omit the first dim (batch dim) of each layer
node_indices.append([[np.random.randint(0,d) for d in lay.output.shape[1:]] for s in range(NB_NODES_PER_LAYER)])
# print("%s:" % lay.name)
# add the last layer
node_indices.append([[x] for x in range(10)])
# print("%s:" % layers_[-1].name)
print(intermediate_model)
# + [markdown] id="Qm1tVpkjh8G3"
# # IQR calculations with segmentation
# + id="CQu8qE0-fSxq"
from skimage.segmentation import felzenszwalb, slic, quickshift
from skimage.segmentation import mark_boundaries
from skimage.util import img_as_float
# + id="AbnhY4q7gNpa"
# SLIC experiment
# number of segments
# n = 32
# n = 64
# n = 128
# QuickShift experiment s-0,1,2 m=10,20
s = 0
m = 10
# felzenszwalb experiment s=1,10,100,1000
# s = 1000
# + colab={"base_uri": "https://localhost:8080/"} id="D5slqEu1hlFS" outputId="19eb1bcf-a8fe-40b9-ab97-44c0e0cf1446"
# %%time
iqr_all = []
iqr_10_all = []
iqr_4k_all =[]
for b in range(2*NB_BATCHES):
print("batch %d:" % b)
images = ds_experiment[b]
preds = pretrained_model.predict(images)
preds_value = tf.reduce_max(preds, axis=1)
preds_idx = tf.argmax(preds, axis=1)
preds_layers = intermediate_model(images)
preds_4k = []
for u in range(NB_LAYERS): # loop through the last 10 layers
for v in node_indices[u]: # loop through the 10 random nodes for that layer
t = tuple(v)
# print((0,*t))
preds_4k.append ( preds_layers[u][(...,*t)] )
preds_4k = np.array(preds_4k).T
# print (preds_value)
# print (preds_idx)
# print ( preds_value == tf.gather_nd(preds, list(zip(range(BATCH_SIZE), preds_idx))) )
iqr = []
iqr10 = []
iqr4k = []
# choose your seg!
# segImages = [ slic(img, n_segments=n) for img in images ]
segImages = [ quickshift(img, sigma=s, max_dist=m) for img in images]
# segImages = [ felzenszwalb(img, scale=s) for img in images]
max_segs = [ segImg.max() for segImg in segImages ]
max_n_segs = max (max_segs)
# max_n_segs = max (segImg.max() for segImg in segImages)
print (max_n_segs)
# input()
# mask = np.ones((1,32,32,3))
for i in range(max_n_segs+1): # i is seg number here
mask = np.ones((BATCH_SIZE,32,32,3))
# mask the segment for all images in the batch
for j in range(BATCH_SIZE): # j is image number here
# print (segImages[j]==i)
# input()
mask[j,(segImages[j]==i),:] = 0
images_0 = images * mask
# for testing the mask
# choose a number (image in batch)
# t = 11
# print (segImages[t].max())
# plt.figure(figsize = (1,1))
# plt.axis('off')
# plt.imshow((images[t].numpy() * std + mean).astype(np.uint8))
# plt.show()
# plt.figure(figsize = (1,1))
# plt.axis('off')
# plt.imshow((images_0[t].numpy() * std + mean).astype(np.uint8))
# plt.show()
# input()
preds_0 = pretrained_model.predict(np.array(images_0))
preds_value_0 = tf.gather_nd(preds_0, list(zip(range(BATCH_SIZE), preds_idx)))
iqr.append(abs(preds_value - preds_value_0))
iqr10.append(abs(preds - preds_0))
preds_layers_0 = intermediate_model(images_0)
preds_4k_0 = []
for u in range(NB_LAYERS): # loop through the last 10 layers ]
for v in node_indices[u]: # loop through the 10 random nodes for that layer
t = tuple(v)
preds_4k_0.append ( preds_layers_0[u][(...,*t)] )
preds_4k_0 = np.array(preds_4k_0).T
iqr4k.append(abs(preds_4k - preds_4k_0))
# for some images the number of segments is less than max_n_segs
# this results in a mask of all ones, the same image will be predicted
# In this case the difference of predicted value(s) will be 0
# to ignore these 0s we transform them to NAN and we use NANpercentile function
# insted of percentile function
# replace iqr vals of 0 with nan
iqr = np.array(iqr)
iqr10 = np.array(iqr10)
iqr4k = np.array(iqr4k)
for j in range(BATCH_SIZE):
iqr[max_segs[j]+1:,j] = np.nan
iqr10[max_segs[j]+1:,j,:] = np.nan
iqr4k[max_segs[j]+1:,j,:] = np.nan
# nanpercentile is a bit slow, can be enhanced
a,b = np.nanpercentile (iqr4k, [25, 75], axis=0)
iqr_4k_vals = b - a
a,b = np.nanpercentile(iqr10, [25, 75], axis=0)
iqr_10_vals = b - a
a,b = np.nanpercentile(iqr, [25, 75], axis=0)
iqr_vals = b - a
iqr_all.append(iqr_vals)
iqr_10_all.append(iqr_10_vals)
iqr_4k_all.append(iqr_4k_vals)
# + [markdown] id="pZFvP4XWSv57"
# # classification
# + [markdown] id="7HGTw7TE4DT2"
# #1_D
# + colab={"base_uri": "https://localhost:8080/"} id="JY-L4YXcSqee" outputId="0104ef63-ab9e-43fa-cf3d-9a0a3d8a9742"
X = np.array(iqr_all).flatten()
y = np.concatenate( ( np.zeros(NB_BATCHES*BATCH_SIZE), np.ones(NB_BATCHES*BATCH_SIZE) ) )
score = cross_val_score(XGBClassifier(), X.reshape(-1,1), y, cv=2)
print (score)
# + colab={"base_uri": "https://localhost:8080/"} id="-X8rKhtdbwVD" outputId="e8cf86fe-6ce1-4694-fe67-a0d150e1182b"
import math
print (len([x for x in X if math.isnan(x)]))
print (len(X))
# + id="kRTy6OqaS2oI"
X_train, X_test, y_train, y_test = train_test_split(X.reshape(-1,1), y, test_size=0.2)
# rdm = RandomForestClassifier().fit(X_train,y_train)
# svc = SVC(probability=True).fit(X_train,y_train)
xgb = XGBClassifier().fit(X_train,y_train)
# + colab={"base_uri": "https://localhost:8080/"} id="iEKZ5TvIS6Gn" outputId="08a6d8fc-dbc0-40af-c3a1-94ac9dfcbee8"
# print(rdm.score(X_train,y_train))
# print(rdm.score(X_test,y_test))
# # print(svc.score(X_train,y_train))
# print(svc.score(X_test,y_test))
# print(xgb.score(X_train,y_train))
print(xgb.score(X_test,y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="zUgq4FmgS_iJ" outputId="3b71c9f6-8da7-41a8-9a05-915b302edcfb"
# AUC
from sklearn.metrics import roc_curve, roc_auc_score
# rdm_probs = rdm.predict_proba(X_test)[:,1]
# svc_probs = svc.predict_proba(X_test)[:,1]
xgb_probs = xgb.predict_proba(X_test)[:,1]
# rdm_auc = roc_auc_score(y_test, rdm_probs)
# svc_auc = roc_auc_score(y_test, svc_probs)
xgb_auc = roc_auc_score(y_test, xgb_probs)
# print('Random Forest: AUROC = %.3f' %(rdm_auc) )
# print('SVC: AUROC = %.3f' %(svc_auc) )
print('RaXGBClassifier: AUROC = %.3f' %(xgb_auc) )
# + [markdown] id="-aNntfBJTOw8"
# # IQR-10D
# + colab={"base_uri": "https://localhost:8080/"} id="M7Uwt6GLTOgY" outputId="b16583f0-7fe7-4793-abe4-c06fd8ce2524"
X = np.array(iqr_10_all).reshape(-1,10)
y = np.concatenate( ( np.zeros(NB_BATCHES*BATCH_SIZE), np.ones(NB_BATCHES*BATCH_SIZE) ) )
score = cross_val_score(XGBClassifier(), X, y, cv=2)
print (score)
# + colab={"base_uri": "https://localhost:8080/"} id="dkIIbeECTXJ6" outputId="084c2932-e2c4-4960-e47c-19c5ae5c7244"
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.2)
# rdm = RandomForestClassifier().fit(X_train,y_train)
# svc = SVC(probability=True).fit(X_train,y_train)
xgb = XGBClassifier().fit(X_train,y_train)
# print(rdm.score(X_test,y_test))
# print(svc.score(X_test,y_test))
print(xgb.score(X_test,y_test))
# + colab={"base_uri": "https://localhost:8080/"} id="4uHV61XpTYzH" outputId="33887157-78ba-4c93-8d38-4d37a6b323fa"
# AUC
from sklearn.metrics import roc_curve, roc_auc_score
# rdm_probs = rdm.predict_proba(X_test)[:,1]
# svc_probs = svc.predict_proba(X_test)[:,1]
xgb_probs = xgb.predict_proba(X_test)[:,1]
# rdm_auc = roc_auc_score(y_test, rdm_probs)
# svc_auc = roc_auc_score(y_test, svc_probs)
xgb_auc = roc_auc_score(y_test, xgb_probs)
# print('Random Forest: AUROC = %.3f' %(rdm_auc) )
# print('SVC: AUROC = %.3f' %(svc_auc) )
print('RaXGBClassifier: AUROC = %.3f' %(xgb_auc) )
# + [markdown] id="vc4UsEZYTebs"
# # IQR-4K-D
#
# + [markdown] id="v87YtmwWTiGW"
# # Classification
# + colab={"base_uri": "https://localhost:8080/"} id="Ixzwf7ASTfRC" outputId="4e45174b-9933-409b-8c8b-87f5c7a236e2"
# We try a very basic classification
X = np.array(iqr_4k_all).reshape(-1,3810)
y = np.concatenate( ( np.zeros(NB_BATCHES*BATCH_SIZE), np.ones(NB_BATCHES*BATCH_SIZE) ) )
score = cross_val_score(XGBClassifier(), X, y, cv=2)
print (score)
# + id="3v_krmqhTngG"
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.2)
# rdm = RandomForestClassifier().fit(X_train,y_train)
# svc = SVC(probability=True).fit(X_train,y_train)
xgb = XGBClassifier().fit(X_train,y_train)
# print(rdm.score(X_test,y_test))
# print(svc.score(X_test,y_test))
print(xgb.score(X_test,y_test))
# + id="zUSuij5_Tp0b" colab={"base_uri": "https://localhost:8080/"} outputId="9822f354-151c-4e57-9c0a-f55691619e74"
# AUC
from sklearn.metrics import roc_curve, roc_auc_score
# rdm_probs = rdm.predict_proba(X_test)[:,1]
# svc_probs = svc.predict_proba(X_test)[:,1]
xgb_probs = xgb.predict_proba(X_test)[:,1]
# rdm_auc = roc_auc_score(y_test, rdm_probs)
# svc_auc = roc_auc_score(y_test, svc_probs)
xgb_auc = roc_auc_score(y_test, xgb_probs)
# print('Random Forest: AUROC = %.3f' %(rdm_auc) )
# print('SVC: AUROC = %.3f' %(svc_auc) )
print('RaXGBClassifier: AUROC = %.3f' %(xgb_auc) )
|
SegFault_TABLE_VI.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# The Code is originally from towardsdatascience.com
# +
import skimage.data
# Reading the image
img = skimage.data.chelsea()
# Converting the image into gray. (It means, we don't want to use 3 Channels-R,G,B- for make problem easier)
img = skimage.color.rgb2gray(img)
# +
import numpy as np
l1_filter = np.zeros((2,3,3)) # 2 : num of filters, 3x3 kernel size
# -
l1_filter[0, :, :] = np.array([[[-1, 0, 1],
[-1, 0, 1],
[-1, 0, 1]]])
l1_filter[1, :, :] = np.array([[[1, 1, 1],
[0, 0, 0],
[-1, -1, -1]]])
# +
# Testing Code
print(img.shape)
print(len(img.shape))
print(img.shape[-1])
print(l1_filter.shape) # (num of filters, width of filter, height of filter)
# -
def conv(img, conv_filter):
if len(img.shape) > 2 or len(conv_filter.shape) > 3: # Check if number of image channels matches the filter depth.
if image.shape[-1] != conv_filter.shape[-1]:
print("Error : Number of channels in both image and filter must match.")
sys.exit()
if conv_filter.shape[-1] != conv_filter.shape[2]: # Check if filter dimensions are equal.
print("Error : Filter must be a square matrix. i.e. number of rows and columns must match.")
sys.exit()
if conv_filter.shape[1] % 2 == 0: # Check if filter dimensions are odd.
print("Error : Filter must have an odd size. i.e. number of rows and columns must be odd.")
sys.exit()
# An empty feature map to hold the output of convolving the filter(s) with the image.
feature_maps = numpy.zeros((img.shape[0]-conv_filter.shape[1] + 1, img.shape[1]-conv_filter.shape[1] + 1, conv_filter.shape[0]))
# Dimension of feature_maps : (Image Width(img.shape[0]) - Filter Width(conv_filter.shape[1]) + 1(Bias), Image Height(img.shape[1]) - Filter Height(conv_filter.shape[1] + 1(Bias), Num of channels(conv_filter.shape[0]))
# conv_filter.shape[1] == conv_filter.shape[2] (3 == 3)
# Convolving the image by the filter(s).
for filter_num in range(conv_filter.shape[0]): # for loop within filters(shape[0] is 2 for now)
print("Filter ", filter_num + 1) # since idx starts from 0, we add 1
curr_filter = conv_filter[filter_num, :] # getting a filter from the bank.
"""
Checking if there are multiple channels for the single filter.
If so, then each channel will convolve the image.
The result of all convolutions are summed to return a single feature map.
"""
if len(curr_filter.shape) > 2: # shape of curr_filter should be (3,3) so len will be 2(no depth)
conv_map = conv_(img[:,:,0], curr_filter[:,:,0]) # Array holding the sum of all feature maps.
for ch_num in range(1, curr_filter.shape[-1]): # Convolving each channel with the image and summing the results.
conv_map = conv_map + conv_(img[:,:,ch_num], curr_filter[:,:,ch_num])
else: # There is just a single channel in the filter
conv_map = conv_(img, curr_filter)
feature_maps[:,:, filter_num] = conv_map # Holding feature map with the current filter
return feature_maps # Returning all feature maps
def conv_(img, conv_filter): # function for convolution operation
filter_size = conv_filter.shape[1] # conv_filter : (num of filter, width, height)
result = np.zeros((img.shape)) # make tuple size of img
# Looping through the image to apply the convolution operation.
for r in np.uint16(np.arange(filter_size/2.0, img.shape[0]-filter_size/2.0 + 1)):
for c in np.uint16(np.arange(filter_size/2.0, img.shape[1]-filter_size/2.0 + 1)):
"""
Getting the current region to get multiplied with the filter.
How to loop through the image and get the region based on the image and filter sizes is the most tricky part of convolution.
"""
curr_region = img[r - np.uint16(np.floor(filter_size/2.0)):r + np.uint16(np.ceil(filter_size/2.0)),
c - np.uint16(np.floor(filter_size/2.0)):c + np.uint16(np.ceil(filter_size/2.0))]
# Element-wise multiplication between the current region and the filter.
curr_result = curr_region * conv_filter
conv_sum = np.sum(curr_result) # Summing the result of multiplication.
result[r, c] = conv_sum # Saving the summation in the convolution layer feature map.
# Clipping the outliers of the result matrix.
final_result = result[np.uint16(filter_size/2.0):result.shape[0] - np.uint16(filter_size/2.0),
np.uint16(filter_size/2.0):result.shape[1] - np.uint16(filter_size/2.0)]
return final_result
|
docs/2008/Resources/Deep Learning Coding with Python/.ipynb_checkpoints/Simple CNN coding in Numpy-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="UoQjqjBOIzfG" colab_type="text"
# # Colaboratory
# + [markdown] id="9mCeNhcsvyDn" colab_type="text"
# ## Before you start
#
# When you open a new Colab from Github (like this one), you cannot save changes. So it's usually best to store the Colab in you personal drive `"File > Save a copy in drive..."` **before** you do anything else.
# + [markdown] id="BcXbc54tbXq7" colab_type="text"
# ## Introduction
# + [markdown] id="fPP3Zw5iV2DP" colab_type="text"
#
#
# Some important links to keep open during the workshop – open these tabs **now**!:
#
# - [TF documentation](https://www.tensorflow.org/api_docs/python/tf) : Use the search box (top right) to get documentation on Tensorflow's rich API.
#
# - [solutions/](https://github.com/tensorflow/workshops/tree/master/extras/amld/notebooks/solutions) : Every notebook in the `exercises/` directory has a corresponding notebook in the `solutions/` directory.
#
# ---
#
# Colaboratory (Colab) is a Jupyter notebook environment which allows you to work with data and code in an interactive manner. You can decide where you want to run your code:
#
# * Using a hosted runtime provided by Google (**default**)
# * Locally using your own machine and resources
#
# It supports Python 3 and comes with a set of pre-installed libraries like Tensorflow and Matplotlib but also gives you the option to install more libraries on demand. The resulting notebooks can be shared in a straightforward way.
#
# Caveats:
#
# * The virtual machines used for the runtimes are **ephemeral** so make sure to safe your data in a persistent location like locally (downloading), in the Google Cloud Storage or Google Drive.
# * The service is free of use but the performance of default runtimes can be insufficient for your purposes.
# * You have the option to select a runtime with GPU or TPU support.
# * "Colaboratory is intended for interactive use. Long-running background computations, particularly on GPUs, may be stopped. [...] We encourage users who wish to run continuous or long-running computations through Colaboratory’s UI to use a local runtime." - See [Colaboratory FAQ](https://research.google.com/colaboratory/faq.html "Colaboratory FAQ")
# + [markdown] id="iDbiROodhaFQ" colab_type="text"
# **Getting started**
#
# 1. Connect to a runtime now by clicking `connect` in the top right corner if you don't already see a green checkmark there.
# 2. To get a better overview you might want to activate the *Table of contents* by clicking on the arrow on the left.
# + [markdown] id="M54Z135xV2DW" colab_type="text"
# ### Important shortcuts
#
# Action | Colab Shortcut | Jupyter Shortcut
# ---|---|---
# Executes current cell | `<CTRL-ENTER>` | `<CTRL-ENTER>`
# Executes current cell and moves to next cell | `<SHIFT-ENTER>` | `S<HIFT-ENTER>`
# Executes current selection | `<CTRL-SHIFT-ENTER>` | `N/A`
# Insert cell above | `<CTRL-M> <A>` | `<A>`
# Append cell below | `<CTRL-M> <B>` | `<B>`
# Shows searchable command palette | `<CTRL-SHIFT-P>` | `<CTRL-SHIFT-P>`
# Convert cell to code | `<CTRL-M> <Y>` | `<Y>`
# Convert cell to Markdown | `<CTRL-M> <M>` | `<M>`
# Autocomplete (on by default) | `<CTRL+SPACE>` | `<TAB>`
# Goes from edit to "command" mode | `<ESC>` | `<ESC>`
# Goes from "command" to edit mode | `<ENTER>` | `<ENTER>`
# Show keyboard shortcuts | `<CTRL-M> <H>` | `<H>`
# <p align="center"><b>Note:</b> On OS X you can use `<COMMAND>` instead of `<CTRL>`</p>
#
# Give it a try!
# + id="iJ4YliVgZNCq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5251639e-ba56-4e86-f37e-757fbbcd13b6" executionInfo={"status": "ok", "timestamp": 1582756148909, "user_tz": -60, "elapsed": 924, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# YOUR ACTION REQUIRED:
# Execute this cell first using <CTRL-ENTER> and then using <SHIFT-ENTER>.
# Note the difference in which cell is selected after execution.
print('Hello world!')
# + [markdown] id="U-B6-3lmkMwJ" colab_type="text"
# You can also only execute one single statement in a cell.
# + id="IprMQ_z8kOsA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="51dc29c4-801a-442f-d607-61b6fb851926" executionInfo={"status": "ok", "timestamp": 1582756149186, "user_tz": -60, "elapsed": 1193, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# YOUR ACTION REQUIRED:
# Execute only the first print statement by selecting the first line and pressing
# <CTRL-SHIFT-ENTER>.
print('Only print this line.')
print('Avoid printing this line.')
# + [markdown] id="_Hlf6HDER7YY" colab_type="text"
# **What to do if you get stuck**
#
# If you should get stuck and the documentation doesn't help you consider using additional help.
# + id="QIKn9TOwV2De" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="16bb1398-774a-4bca-e28d-b8c25bec7e10" executionInfo={"status": "ok", "timestamp": 1582756149187, "user_tz": -60, "elapsed": 1191, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
def xor_str(a, b):
return ''.join([chr(ord(a[i % len(a)]) ^ ord(b[i % len(b)]))
for i in range(max(len(a), len(b)))])
# YOUR ACTION REQUIRED:
# Try to find the correct value for the variable below.
workshop_secret = 'Tensorflow rocks' #workshop_secret = '(replace me!)'
xor_str(workshop_secret,
'\x03\x00\x02\x10\x00\x1f\x03L\x1b\x18\x00\x06\x07\x06K2\x19)*S;\x17\x08\x1f\x00\x05F\x1e\x00\x14K\x115\x16\x07\x10\x1cR1\x03\x1d\x1cS\x1a\x00\x13J')
# Hint: You might want to checkout the ../solutions directory
# (you should already have opened this directory in a browser tab :-)
# + [markdown] id="UPQzxvqbMXqr" colab_type="text"
# ### Importing TensorFlow
# + [markdown] id="rScwW1Q69xJ-" colab_type="text"
# We'll be using **TensorFlow 2.1.0** in this workshop. This will soon be the default, but for the time being we still need to activate it with the Colab-specific `%tensorflow_version` magic.
# + id="Ys4hO5MtkhCC" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="a89428b1-5ef5-4880-e55f-e78cfe9e4408" executionInfo={"status": "ok", "timestamp": 1582756149188, "user_tz": -60, "elapsed": 1189, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# We must call this "magic" before importing TensorFlow. We will explain
# further down what "magics" (starting with %) are.
# %tensorflow_version 2.x
# + colab_type="code" id="n_J1HPc4kpbl" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9365ea52-a036-4425-c38e-a51b09cbeef9" executionInfo={"status": "ok", "timestamp": 1582756156635, "user_tz": -60, "elapsed": 8634, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Include basic dependencies and display the tensorflow version.
import tensorflow as tf
tf.__version__
# + [markdown] id="1uKqdyHFB8NY" colab_type="text"
# ### Running shell commands
#
# You can run shell commands directly in Colab: simply prepend the command with a **!**.
# + id="FY3pksK4V2DY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="d2d7f45c-de66-4c05-872e-4ed0c05c4ee3" executionInfo={"status": "ok", "timestamp": 1582756159826, "user_tz": -60, "elapsed": 11822, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Print the current working directory and list all files in it.
# !pwd
# !ls
# + id="8BbKSuKslVaI" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 415} outputId="7673b17f-4123-4da4-cefb-f997646d5950" executionInfo={"status": "ok", "timestamp": 1582756164534, "user_tz": -60, "elapsed": 16527, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Especially useful: Installs new packages.
# !pip install qrcode
import qrcode
qrcode.make('Colab rocks!')
# + [markdown] id="ywxYXQ-lCc2_" colab_type="text"
# **Autocompletion and docstrings**
#
# Jupyter shows possible completions of partially typed
# commands.
#
# Try it for yourself by displaying all available `tf.` methods that start with `one`.
# + id="a_jmV9woV2Db" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="953d5057-ef50-4bc8-9c30-8ce95a552e51" executionInfo={"status": "ok", "timestamp": 1582756164535, "user_tz": -60, "elapsed": 16526, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# YOUR ACTION REQUIRED:
# Set the cursor to after tf.one and press <CTRL-SPACE>.
# On Mac, only <OPTION-ESCAPE> might work.
tf.one_hot #tf.one
# + [markdown] id="PuBe6L8WOWuS" colab_type="text"
# In addition, you can also display docstrings to see the function signature and possible parameters.
# + id="hPiM64eaOi2y" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3e6029ce-dc64-40c8-8a6e-444a983b81bb" executionInfo={"status": "ok", "timestamp": 1582756164535, "user_tz": -60, "elapsed": 16523, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# YOUR ACTION REQUIRED:
# Complete the command to `tf.maximum` and then add the opening bracket "(" to
# see the function documentation.
tf.maximum([1, 2, 3], [2, 2, 2]) #tf.maximu
# + [markdown] id="NK1yV3Ye0pbC" colab_type="text"
# Alternatively, you might also inspect function details with docstrings if available by appending a "?".
# + id="P-IF1l5u0MLT" colab_type="code" colab={}
# tf.maximum?
# + [markdown] id="-wBzGaDMaCg2" colab_type="text"
# **Note:** This also works for any other type of object as can be seen below.
# + id="nLXoW6IXZEvP" colab_type="code" colab={}
test_dict = {'key0': 'Tensor', 'key1': 'Flow'}
# test_dict?
# + [markdown] id="_mahgn1AHL61" colab_type="text"
# ## Runtimes
#
# As noted in the introduction above, Colab provides multiple runtimes with different hardware accelerators:
#
# * CPU (default)
# * GPU
# * TPU
#
# which can be selected by choosing `"Runtime > Change runtime type"` in the menu.
#
# Please be aware that selecting a new runtime will assign a new virtual machine (VM).
# In general, assume that any changes you make to the VM environment including data storage are **ephemeral**. Particularly, this might require to **execute previous cells again** as their content is unknown to a new runtime otherwise.
#
# Let's take a closer look at one of such provided VMs.
#
# + [markdown] id="whv1zmERgI5v" colab_type="text"
# Once we have been assigned a runtime we can inspect it further.
# + id="AwLSimGKgHFd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="3de5cff2-659f-4c4b-d868-dda688f3fcc8" executionInfo={"status": "ok", "timestamp": 1582756166562, "user_tz": -60, "elapsed": 18542, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Display how long the system has been running.
# Note : this shows "0 users" because no user is logged in via SSH.
# !uptime
# + [markdown] id="f69GlgeggRED" colab_type="text"
# As can be seen, the machine has been allocated just very recently for our purposes.
#
# **VM specifications**
# + colab_type="code" id="AAXhttijkq8P" colab={"base_uri": "https://localhost:8080/", "height": 570} outputId="a0111ac7-d47d-4f50-c471-46a300dce4a6" executionInfo={"status": "ok", "timestamp": 1582756171179, "user_tz": -60, "elapsed": 23157, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Display available and used memory.
# !free -h
print("-"*70)
# Display the CPU specification.
# !lscpu
print("-"*70)
# Display the GPU specification (if available).
!(nvidia-smi | grep -q "has failed") && echo "No GPU found!" || nvidia-smi
# + [markdown] id="bqCA9-VDIjsu" colab_type="text"
# ## Plotting
#
# The notebook environment also provides options to visualize and interact with data.
#
# We'll take a short look at the plotting/visualization libraries Matplotlib and Altair.
# + [markdown] id="1iJfoQ86AsfO" colab_type="text"
# ### Matplotlib
#
# Matplotlib is one of the most famous Python plotting libraries and can be used to plot results within a cell's output (see [Matplotlib Introduction](https://matplotlib.org/users/intro.html "Matplotlib Introduction")).
#
# Let's try to plot something with it.
# + id="gyk4SEk_V2DW" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 282} outputId="aec3e96a-82b4-4867-b681-e123c6d1168d" executionInfo={"status": "ok", "timestamp": 1582756171181, "user_tz": -60, "elapsed": 23154, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFS<KEY>sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Display the Matplotlib outputs within a cell's output.
# %matplotlib inline
import numpy as np
from matplotlib import pyplot
# Create a randomized scatterplot using matplotlib.
x = np.random.rand(100).astype(np.float32)
noise = np.random.normal(scale=0.3, size=len(x))
y = np.sin(x * 7) + noise
pyplot.scatter(x, y)
# + [markdown] id="lhUudwcUafIm" colab_type="text"
# ### Altair
#
# Another declarative visualization library for Python is Altair (see [Altair: Declarative Visualization in Python](https://altair-viz.github.io/)).
#
# Try to zoom in/out and to hover over individual data points in the resulting plot below.
# + id="uO3MmbhbIw_t" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 367} outputId="aee12f31-b3f5-4dfc-f612-3925f90781e0" executionInfo={"status": "ok", "timestamp": 1582756171502, "user_tz": -60, "elapsed": 23472, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Load an example dataset.
from vega_datasets import data
cars = data.cars()
# Plot the dataset, referencing dataframe column names.
import altair as alt
alt.Chart(cars).mark_point().encode(
x='Horsepower',
y='Miles_per_Gallon',
color='Origin',
tooltip=['Name', 'Origin', 'Horsepower', 'Miles_per_Gallon']
).interactive()
# + [markdown] id="cBUraBMAHE0J" colab_type="text"
# ## Notebook Magics
#
# The IPython and Colab environment support built-in magic commands called magics (see: [IPython - Magics](https://ipython.readthedocs.io/en/stable/interactive/magics.html)).
#
# In addition to default Python, these commands might be handy for example when it comes to interacting directly with the VM or the Notebook itself.
#
#
# + [markdown] id="KO1K1UnIfU0u" colab_type="text"
# ### Cell magics
#
# Cell magics define a mode for a complete cell and are prefixed with **%%**.
#
# Examples include:
#
# * **%%bash** or **%%sh**
# * **%%html**
# * **%%javascript**
#
# + id="2qT4YGbOQ9FG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 214} outputId="10447421-6020-4c33-d5da-c6c25d4c85f6" executionInfo={"status": "ok", "timestamp": 1582756171503, "user_tz": -60, "elapsed": 23471, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}} language="sh"
# echo "This is a shell script!"
# # List all running VM processes.
# ps -ef
# echo "Done"
# + id="qUSc9sT2JskX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 36} outputId="5b4390a5-fcc5-497d-be3f-6d9ca37b4e60" executionInfo={"status": "ok", "timestamp": 1582756171504, "user_tz": -60, "elapsed": 23469, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Embed custom HTML directly into a cell's output.
# %%html
<marquee>HTML rocks</marquee>
# + [markdown] id="Uy0j4iJ5d8v3" colab_type="text"
# ### Line magics
#
# You can also make use of line magics which can be inserted anywhere at the beginning of a line inside a cell and need to be prefixed with **%**.
#
# Examples include:
#
#
# * **%time** - display the required time to execute the current line
# * **%cd** - change the current working directory
# * **%pdb** - invoke an interactive Python debugger
# * **%lsmagic** - list all available line magic and cell magic functions
#
#
# + [markdown] id="mrdhMGsyN2Gl" colab_type="text"
# For example, if you want to find out how long one specific line requires to be executed you can just prepend **%time**.
#
# + id="Kz0d1QoINx95" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="aac123b1-5b0f-4b16-99cf-f068e727494b" executionInfo={"status": "ok", "timestamp": 1582756171504, "user_tz": -60, "elapsed": 23466, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
n = 1000000
# %time list1 = [i for i in range(n)]
print("")
# %time list2 = [i for i in range(int(n/2))]
# + [markdown] id="Y_Q3lleVPMVJ" colab_type="text"
# **Note:** Some line magics like **%time** can also be used for complete cells by writing **%%time**.
# + id="LCIrCKs9a1eu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="ae5ae154-275a-4763-e045-e21af2cf66b2" executionInfo={"status": "ok", "timestamp": 1582756171785, "user_tz": -60, "elapsed": 23745, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# %%time
n = 1000000
list1 = [i for i in range(n)]
list2 = [i for i in range(int(n/2))]
# + [markdown] id="BGurTn7rVH_I" colab_type="text"
# ## Data handling
#
# There are multiple ways to provide data to a Colabs's VM environment.
# > **Note:** This section only applies to Colab.
# > Jupyter has a file explorer and other options for data handling.
#
# The options include:
# * Uploading files from the local file system.
# * Connecting to Google Cloud Storage (explained below).
# * Connecting to Google Drive (see: [Snippets: Drive](https://colab.sandbox.google.com/notebooks/snippets/drive.ipynb); will be used in the next Colabs).
# + [markdown] id="Zdb5RpNmxoJl" colab_type="text"
# **Uploading files from the local file system**
#
# If you need to manually upload files to the VM, you can use the files tab on the left. The files tab also allows you to browse the contents of the VM and when you double click on a file you'll see a small text editor on the right.
# + [markdown] id="swVo-jxCW6KK" colab_type="text"
# **Connecting to Google Cloud Storage**
#
# [Google Cloud Storage](https://cloud.google.com/storage/?hl=de) (GCS) is a cloud file storage service with a RESTful API.
#
# We can utilize it to store our own data or to access data provided by the following identifier:
#
#
# ```
# gs://[BUCKET_NAME]/[OBJECT_NAME]
# ```
#
# We'll use the data provided in **gs://amld-datasets/zoo_img** as can be seen below.
#
# Before we can interact with the cloud environment, we need to grant permissions accordingly (also see [External data: Cloud Storage](https://colab.research.google.com/notebooks/io.ipynb#scrollTo=S7c8WYyQdh5i)).
# + id="1aJ1e7JFVIsQ" colab_type="code" colab={}
from google.colab import auth
auth.authenticate_user()
# + [markdown] id="f6_4yqiOfWKt" colab_type="text"
# List a subset of the contained files using the [gsutil tool](https://cloud.google.com/storage/docs/gsutil?hl=en).
# + id="wJAutwkEWWxg" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 194} outputId="c23ef39f-9f2e-45e4-a000-3472446cc5f4" executionInfo={"status": "ok", "timestamp": 1582756212187, "user_tz": -60, "elapsed": 64142, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# !gsutil ls gs://amld-datasets/zoo_img | head
# + [markdown] id="d10y2KZpfsed" colab_type="text"
# Conveniently, TensorFlow natively supports multiple file systems such as:
#
# * GCS - Google Cloud Storage
# * HDFS - Hadoop
# * S3 - Amazon Simple Storage
#
# An example for the GCS filesystem can be seen below.
# + id="MDUnLXGXWnO8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 194} outputId="6567ad09-600d-44ba-919b-c00bfd2b7b18" executionInfo={"status": "ok", "timestamp": 1582756213170, "user_tz": -60, "elapsed": 65123, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Note: This cell hangs if you forget to call auth.authenticate_user() above.
tf.io.gfile.glob('gs://amld-datasets/zoo_img/*')[:10]
# + [markdown] id="AHhyyTLRHDlH" colab_type="text"
# ## Snippets
#
# Finally, we can take a look at the snippets support in Colab.
# > If you're using Jupyter please see [Jupyter contrib nbextensions - Snippets menu](https://jupyter-contrib-nbextensions.readthedocs.io/en/latest/nbextensions/snippets_menu/readme.html) as this is not natively supported.
#
# Snippets are a way to quickly "bookmark" pieces of code or text that you might want to insert into specific cells.
#
#
#
#
#
# + id="4Dph99hAr61H" colab_type="code" colab={}
# YOUR ACTION REQUIRED:
# Explore existing snippets by going to the `Code snippets` section.
# Click on the <> button on the left sidebar to open the snippets.
# Alternatively, you can press `<CTRL><ALT><P>` (or `<COMMAND><OPTION><P>` for
# OS X).
# + [markdown] id="KIwzrAszTiTA" colab_type="text"
# We have created some default snippets for this workshop in:
# > https://colab.research.google.com/drive/1OFSjEmqC-UC66xs-LR7-xmgkvxYTrAcN
#
# In order to use these snippets, you can:
#
# 1. Click on "Tools > Settings".
# 2. Copy the above url into "Custom snippet notebook URL" and press enter.
#
# As soon as you update the settings, the snippets will then become available in every Colab. Search for "amld" to quickly find them.
#
# Alternatively, you can also add snippets via the API (but this needs to be done for every Colab/kernel):
# + id="_rO9y-43U_lL" colab_type="code" colab={}
from google.colab import snippets
# snippets.register('https://colab.research.google.com/drive/1OFSjEmqC-UC66xs-LR7-xmgkvxYTrAcN')
# + [markdown] id="57T5LlVEpfp2" colab_type="text"
# **Pro tip** : Maybe this is a good moment to create your own snippets and register them in settings. You can then start collecting often-used code and have it ready when you need it... In this Colab you'll need to have text cells with titles (like `### snippet name`) preceeding the code cells.
# + [markdown] id="grr29b3-gCd-" colab_type="text"
# # ----- Optional part -----
# + [markdown] id="W0Uq0G0LrtjG" colab_type="text"
# ## Custom line magic
#
# You can also define your own line/cell magic in the following way.
# + id="Y8bjXSjBJ2oP" colab_type="code" colab={}
from IPython.core.magic import register_line_cell_magic
@register_line_cell_magic
def mymagic(line_content, cell_content=None):
print('line_content="%s" cell_content="%s"' % (line_content, cell_content))
# + id="lu7Z7VTEKPWh" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="7b61ac7c-1fc1-4e01-9950-9e4c4809841c" executionInfo={"status": "ok", "timestamp": 1582756213173, "user_tz": -60, "elapsed": 65116, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# %mymagic Howdy Alice!
# + id="GGBRfLm6LTvU" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="9858547e-3152-4031-e6f9-49b55b5777e3" executionInfo={"status": "ok", "timestamp": 1582756213173, "user_tz": -60, "elapsed": 65113, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# %%mymagic simple question
Howdy Alice!
how are you?
# + [markdown] id="hZUSUu5A1mAD" colab_type="text"
# ## Forms
#
# You can simplify cells by hiding their code and displaying a form instead.
#
#
# **Note:** You can display or hide the code by double clicking the form which might be on the right side.
# + id="2EkpzaQA1u7y" colab_type="code" cellView="form" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="cbaf62d7-4f5f-4b82-8d9d-7571be17003d" executionInfo={"status": "ok", "timestamp": 1582756213174, "user_tz": -60, "elapsed": 65110, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
#@title Execute me
# Hidden cell content.
print("Double click the cell to see its content.")
# + id="adKaHZNZ2jZo" colab_type="code" cellView="both" colab={"base_uri": "https://localhost:8080/", "height": 52} outputId="b4a88299-727b-46b4-8fe3-c56d42a809a7" executionInfo={"status": "ok", "timestamp": 1582756213175, "user_tz": -60, "elapsed": 65108, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# Form example mostly taken from "Adding form fields" Snippet.
#@title Example form
#@markdown Specify some test data and execute this cell.
string_type = 'test_string' #@param {type: "string"}
slider_value = 145 #@param {type: "slider", min: 100, max: 200}
number = 1339 #@param {type: "number"}
date = '2019-01-26' #@param {type: "date"}
pick_me = "a" #@param ['a', 'b', 'c']
#@markdown ---
print("Submitted data:")
print(string_type, slider_value, number, date, pick_me)
# + [markdown] id="kx_ETyD5r0vJ" colab_type="text"
# ## Interactive debugging
# + [markdown] id="8rXJALYPr-Eg" colab_type="text"
# An example of an IPython tool that you can utilize is the interactive debugger
# provided inside an IPython environment like Colab.
#
# For instance, by using **%pdb on**, you can automatically trigger the debugger on exceptions to further analyze the state.
#
# Some useful debugger commands are:
#
# Description | Command
# ---|---
# **h**(elp) | Display available commands
# **p**(rint) `x` | Show content of object `x`
# **w**(here) | Show current instruction pointer position
# **q**(uit) | Leave the debugger
# + id="sMGORnaE2yJ-" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="9b8f3cb3-dc3a-4ad2-9a3a-9de4fc7ad1c7" executionInfo={"status": "ok", "timestamp": 1582756213175, "user_tz": -60, "elapsed": 65105, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAYx167H2vNmFSKlsQkQY-bjbJ-3sPGymaG0kXO=s64", "userId": "08860260976100898876"}}
# YOUR ACTION REQUIRED:
# Execute this cell, print the variable contents of a, b and exit the debugger.
# %pdb on
a = 67069 / 47 - 0x5a
b = a - 0x539
#c = a / b # Will throw an exception.
# + [markdown] id="eHQDeKx06WPq" colab_type="text"
# We'll not dive further into debugging but it's useful to know that this option exists.
#
# Please see [Python Docs - pdb The Python Debugger](https://docs.python.org/2/library/pdb.html) for more information.
# + [markdown] id="kPD8cAscKCes" colab_type="text"
# ## A Word of Warning
#
# While notebook environments like Colab/Jupyter provide many benefits, they also come with some caveats that you should be aware of.
# One example is that you might quickly execute cells in a wrong order leading to unexpected behavior.
#
# If you're interested in more examples feel free to take a look at:
#
# [Youtube - I don't like notebooks by <NAME>](https://www.youtube.com/watch?v=7jiPeIFXb6U) (duration ~56 minutes)
|
extras/amld/notebooks/solutions/0_colab.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Image classification with MNIST data, solutions to exercises
#
# ## Loading and visualizing the data
#
# First, let's load the dataset using keras helpers and visualize some images using pyplot
# + jupyter={"outputs_hidden": false}
#The pylab inline below is something you may need to make images and plots visible in Jupyter, depending on your Anaconda setup
# %pylab inline
import numpy as np
import matplotlib.pyplot as pp
import os
os.environ["CUDA_VISIBLE_DEVICES"]="-1" #disable Tensorflow GPU usage, a simple example like this runs faster on CPU
import tensorflow as tf
from tensorflow import keras
#load the MNIST dataset
mnist = keras.datasets.mnist
(x_train, y_train),(x_test, y_test) = mnist.load_data()
#Scale the pixel intensity values to 0...1 from 0...255
#Fortunately, we don't a StandardScaler here
x_train, x_test = x_train / 255.0, x_test / 255.0
#check the shape: you should see that x_train is a 3D tensor,
#with 60000 instances of 2D tensors 28x28 pixels
print("shape: ",x_train.shape)
#because the keras layers we will use need explicitly defined pixel channel count as the fourth dimension,
#we reshape:
x_train=np.reshape(x_train,[x_train.shape[0],x_train.shape[1],x_train.shape[2],1])
print("new shape: ",x_train.shape)
#do the same for test data
x_test=np.reshape(x_test,[x_test.shape[0],x_test.shape[1],x_test.shape[2],1])
#visualize some of the images
pp.figure(1)
for i in range(8):
pp.subplot(1,8,1+i)
#imshow expects a 2d tensor, thus we pick the i:th image, full width and height, and the first and only color channel
pp.imshow(x_train[i,:,:,0])
# -
# # Exercise 1: Adding more layers to the fully connected network
# The first exercise was to simply add more layers. We only need one line of code per layer
# + jupyter={"outputs_hidden": false}
#Let's import the layer types we need
from tensorflow.keras.layers import Dense #fully connected layer
from tensorflow.keras.layers import Flatten #converts images to vectors of numbers
#As before, we use a simply sequential, i.e., multilayer architecture
model = keras.models.Sequential()
#Flatten converts a batch of multidimensional data into a batch of 1D data.
#This is what the fully connected layers expect.
#For example, the rows of an image are simply stacked after each other.
#If the data was not images, we would not need this.
model.add(Flatten())
#This is the extra layer. You can try modifying the neuron and layer counts and see how the neuron weights and classification accuracy change
model.add(Dense(64, activation='relu'))
#The output layer is fully connected, with 1 neuron for each 10 classes.
#For classification, one should use the softmax activation.
#This means that each output neuron can be thought as the probability of a class.
model.add(Dense(10, activation='softmax'))
#Compile the model. We use sparse_categorical_crossentropy loss instead of categorical_crossentropy,
#because the label data contains indices instead of one-hot vectors
model.compile(loss=keras.losses.sparse_categorical_crossentropy,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
#Train the network
model.fit(x_train, y_train,
batch_size=32,
epochs=5,
verbose=1,
validation_data=(x_test, y_test))
# -
# We can visualize the first layer weights similar to before. Note that for the second layer, we will have as many inputs as the previous layer has neurons, and the input tensors can no longer be interpreted as images. You could, however, try investigating those weights by synthesizing (optimizing) an input image that maximally excites a neuron. This is one of the exercises in the [Adversarial MNIST](AdversarialMNIST.ipynb) tutorial.
# + jupyter={"outputs_hidden": false}
#Visualize some of the first layer neuron weights
#First, query the weights. We use index 1 because index 0 is the flatten layer
weights=model.layers[1].get_weights()[0]
#Create a figure with appropriate size
nNeuronsToVisualize=10
pp.figure(1,figsize=[nNeuronsToVisualize*2,2])
#Loop over the neurons
for i in range(nNeuronsToVisualize):
#Weights is a 2D tensor where the first dimension indexes over data variables, second over neurons
image=weights[:,i]
#We must reshape back to an image
image=np.reshape(image,[28,28])
#Now we can display
pp.subplot(1,nNeuronsToVisualize,1+i)
pp.title("Neuron {}".format(i))
pp.imshow(image)
# -
# ## A convolutional neural network
#
# For the rest of the exercises, we again train a convolutional neural network, which gives better classification accuracy.
# + jupyter={"outputs_hidden": false}
#Let's import the layer types we need
from tensorflow.keras.layers import Dense #fully connected layer
from tensorflow.keras.layers import Conv2D #convolutional layer with 2D filters (for audio you would use 1D)
from tensorflow.keras.layers import Dropout #this mitigates overfitting
#As before, we use a simply sequential, i.e., multilayer architecture
model = keras.models.Sequential()
#Instead of using fully connected layers like before, we use convolutional ones.
#We use 5x5 pixel features, and use strides of 2x2 to drop resolution by a factor of 2 after each layer
model.add(Conv2D(16, kernel_size=(5, 5), strides=[2,2],
activation='relu',
input_shape=(28,28,1,)))
model.add(Conv2D(32, (5, 5), activation='relu', strides=[2,2]))
#After the previous two layers, we are at 7x7 pixel resolution instead of the original 28x28 pixels.
#Thus, 5x5 filters would not be meaningful, as they would encompass almost the whole images
model.add(Conv2D(32, (3, 3), activation='relu', strides=[2,2]))
#Now, we are at 3x3 pixel resolution and there's no point in doing convolutions anymore.
#Instead, we'll just add a small fully connected layer just like above
#Again, we first need to flatten from a batch of images to a batch of 1D tensors
model.add(Flatten())
#Some regularization
model.add(Dropout(0.5))
#One fully connected
model.add(Dense(32, activation='relu'))
#More regularization
model.add(Dropout(0.5))
#Last fully connected layer, with softmax activation, which is what one needs for classification.
#Softmax means that each output neuron can be thought as the probability of a class.
#We use 10 neurons because MNIST has 10 classes.
model.add(Dense(10, activation='softmax'))
#Compile the model. We use sparse_categorical_crossentropy loss instead of categorica_crossentropy,
#because the label data contains indices instead of one-hot vectors
model.compile(loss=keras.losses.sparse_categorical_crossentropy,
optimizer=keras.optimizers.Adam(),
metrics=['accuracy'])
#Train the network
model.fit(x_train, y_train,
batch_size=32,
epochs=5,
verbose=1,
validation_data=(x_test, y_test))
# -
# Let's test the classifier with an images.
# + jupyter={"outputs_hidden": false}
#this is the test image
testIdx=0
#Show the image
print("Testing with image:")
pp.imshow(x_test[testIdx,:,:,0])
pp.show()
#We index by testIdx:testIdx+1 to pass a batch of one image to the network instead of just one image
classProbabilities=model.predict(x_test[testIdx:testIdx+1])
print("Predicted class probabilities: ",classProbabilities)
#np.argmax returns the index of the largest value in a Numpy tensor.
#np.max returns the largest value
print("Most probable class is {}, with probability {}".format(np.argmax(classProbabilities),np.max(classProbabilities)))
# -
# # Exercise 2: Test the classifier with synthetic images
# The second exercise was to create an artificial image, to practice a bit more Numpy tensor manipulation. You could also consider using a Python library for drawing lines etc. into Numpy arrays, e.g., skimage
#
# Here, we simply use np.zeros() to create images with all pixels zero, and then use tensor indexing to set some pixels to 1.
# + jupyter={"outputs_hidden": false}
#MNIST images are 28x28 pixels
image1=np.zeros([28,28])
#Draw a vertical bar. Note: The usual convention expected by Numpy and Tensorflow is that
#in 2D tensors representing images, the first dimension denotes vertical position and second denotes horizontal
image1[5:22,14:16]=1
#Visualize
pp.imshow(image1)
#Test classification.
#Note: we reshape the single image into a batch. (Try running the code without to see what error you get!)
#Reshaping does not change Tensor contents,
#it just changes the way contents are indexed
classProbabilities=model.predict(np.reshape(image1,[1,28,28,1]))
print("Predicted class probabilities: ",classProbabilities)
#np.argmax returns the index of the largest value in a Numpy tensor.
#np.max returns the largest value
print("Most probable class is {}, with probability {}".format(np.argmax(classProbabilities),np.max(classProbabilities)))
# -
# **Try changing the position of the number vertically or horizontally and running the code again!** Does it affect the classification?
#
# Theory says that a fully connected network is more sensitive to the position, but a convolutional neural network should care less about it.
#
# You can change what network you use by re-running one of the network building and training cells above.
#
# Also, **try drawing different numbers or patterns.** Can you fool the network with something that doesn't look like a number? This is called an adversarial image, and neural networks are quite prone to them, if trained with too little data that does not contain all the possible types of images and variations the network will be tested with. See also the [Adversarial MNIST tutorial](AdversarialMNIST.ipynb) for an example of how to optimize images to fool the network.
# # Exercise 3: Visualize the images with lowest correct class probabilities
# This is a good way to gain insights into both 1) the quality of your dataset, and 2) what the network learns.
#
# There is a few ways to do this. First, we show how to extract the correct class probabilities from the probability distributions predicted by the network.
#
# An alternative is to query the loss function values from the network. The cross-entropy loss is a distance metric between the real class probability distribution and the distribution output by the network. In case of fully known real classes, the real probability distributions are one-hot, i.e., the probability of the correct class is 1 and the other probabilities are zero. This means that finding the image with largest loss is equal to finding the image with lowest correct class probability (for the mathematically inclined, can you figure out why this is the case?). However, getting the loss values out from the network requires either [overriding some Keras callbacks](https://stackoverflow.com/questions/48118111/get-loss-values-for-each-training-instance-keras) or building the network a bit differently, as shown at the bottom of this notebook.
# + jupyter={"outputs_hidden": false}
#We begin by compiling a 2D array that includes both image indices and correct classes
#We can get an 1D tensor of indices using np.arange
nImages = y_test.shape[0]
indices=np.arange(nImages)
#Now we can stack both the 1D tensor of indices and the 1D tensor of correct classes
#The axis=1 defines the dimension along which to stack
classesAndIndices=np.stack([indices,y_test],axis=1)
print("Classes and indices",classesAndIndices)
#Next, pass the whole test data through the network
#This will result in a 2D tensor containing a 1D tensor of class probabilities for each image
predictedProbabilities=model.predict(x_test)
print("Predicted probabilities: ",predictedProbabilities)
#Now, we can use the classes and indices to index the probabilities
correctClassProbabilities=predictedProbabilities[classesAndIndices[:,0],classesAndIndices[:,1]]
print("Correct class probabilities: ",correctClassProbabilities)
# -
# We can now find the minimum and display the corresponding image.
# + jupyter={"outputs_hidden": false}
index=np.argmin(correctClassProbabilities)
print("Correct class {}, probability {}".format(y_test[index],correctClassProbabilities[index]))
pp.imshow(x_test[index,:,:,0])
# -
# Now, if we want to show multiple images, we can use np.argsort() to pick the indices
# + jupyter={"outputs_hidden": false}
sortedIndices=np.argsort(correctClassProbabilities)
nImages=8
pp.figure(1,figsize=[nImages*3,3])
for i in range(nImages):
pp.subplot(1,nImages,1+i)
index=sortedIndices[i]
pp.imshow(x_test[index,:,:,0])
pp.title("{}, prob. {:1.5f}".format(y_test[index],correctClassProbabilities[index]))
# -
# Just out of curiosity, let's also show the images with high correct class probabilities
# + jupyter={"outputs_hidden": false}
sortedIndices=np.argsort(correctClassProbabilities)
nImages=8
pp.figure(1,figsize=[nImages*3,3])
for i in range(nImages):
pp.subplot(1,nImages,1+i)
#we can use negative indexing to index the last elements. Since i starts from 0, we also need to have the "-1"
index=sortedIndices[-i-1]
pp.imshow(x_test[index,:,:,0])
pp.title("{}, prob. {:1.5f}".format(y_test[index],correctClassProbabilities[index]))
# -
|
Code/Jupyter/MNIST_solutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nbgrader={}
# # Matplotlib Exercise 1
# + [markdown] nbgrader={}
# ## Imports
# + nbgrader={}
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
# + [markdown] nbgrader={}
# ## Line plot of sunspot data
# + [markdown] nbgrader={}
# Download the `.txt` data for the "Yearly mean total sunspot number [1700 - now]" from the [SILSO](http://www.sidc.be/silso/datafiles) website. Upload the file to the same directory as this notebook.
# + deletable=false nbgrader={"checksum": "7f8ea13f251ef02c216ed08cad6516a7", "grade": true, "grade_id": "matplotlibex01a", "points": 1}
import os
assert os.path.isfile('yearssn.dat')
# + [markdown] nbgrader={}
# Use `np.loadtxt` to read the data into a NumPy array called `data`. Then create two new 1d NumPy arrays named `years` and `ssc` that have the sequence of year and sunspot counts.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
data = np.loadtxt('yearssn.dat')
year = data[:,0]
ssc = data[:,1]
# + deletable=false nbgrader={"checksum": "487fbe3f8889876c782a18756175d727", "grade": true, "grade_id": "matplotlibex01b", "points": 1}
assert len(year)==315
assert year.dtype==np.dtype(float)
assert len(ssc)==315
assert ssc.dtype==np.dtype(float)
# + [markdown] nbgrader={}
# Make a line plot showing the sunspot count as a function of year.
#
# * Customize your plot to follow Tufte's principles of visualizations.
# * Adjust the aspect ratio/size so that the steepest slope in your plot is *approximately* 1.
# * Customize the box, grid, spines and ticks to match the requirements of this data.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
plt.figure(figsize=(50,5))
plt.plot(year, ssc)
plt.grid(True)
plt.yticks([50,100,150,200], [50,100,150,200])
plt.xticks([1700,1750,1800,1850,1900,1950,2000], [1700,1750,1800,1850,1900,1950,2000])
plt.xlabel('Year')
plt.ylabel('Sun Spot Count')
plt.title('Sun Spot Counts per Year 1700-Now')
# + deletable=false nbgrader={"checksum": "d7cdb9758e069eb5f0d1c1b4c4f56668", "grade": true, "grade_id": "matplotlibex01c", "points": 3}
assert True # leave for grading
# + [markdown] nbgrader={}
# Describe the choices you have made in building this visualization and how they make it effective.
# + [markdown] deletable=false nbgrader={"checksum": "89c49052b770b981791536f5c2b07e13", "grade": true, "grade_id": "matplotlibex01d", "points": 1, "solution": true}
# YOUR ANSWER HERE
#
# First I streched the graph way out horizontally to lessen the slope, making the data easier to read.
# I then labeled each axis and gave the graph a title, because not doing so is crazy talk.
# Then I altered what 'ticks' show up, as only certain values are important to what this graph is trying to show.
# + [markdown] nbgrader={}
# Now make 4 subplots, one for each century in the data set. This approach works well for this dataset as it allows you to maintain mild slopes while limiting the overall width of the visualization. Perform similar customizations as above:
#
# * Customize your plot to follow Tufte's principles of visualizations.
# * Adjust the aspect ratio/size so that the steepest slope in your plot is *approximately* 1.
# * Customize the box, grid, spines and ticks to match the requirements of this data.
# + deletable=false nbgrader={"checksum": "6cff4e8e53b15273846c3aecaea84a3d", "solution": true}
year
year_1st=year[0:100]
year_2nd=year[101:200]
year_3rd=year[201:300]
year_4th=year[301:400]
ssc_1st=ssc[0:100]
ssc_2nd=ssc[101:200]
ssc_3rd=ssc[201:300]
ssc_4th=ssc[301:400]
plt.figure(figsize=(30,10))
plt.subplot(4,1,1)
plt.title('Sun Spot Counts per Year')
plt.plot(year_1st, ssc_1st)
plt.tight_layout()
plt.yticks([50,100,150,200], [50,100,150,200])
plt.ylabel('Sun Spot Count')
plt.subplot(4,1,2)
plt.plot(year_2nd, ssc_2nd)
plt.tight_layout()
plt.yticks([50,100,150,200], [50,100,150,200])
plt.ylabel('Sun Spot Count')
plt.subplot(4,1,3)
plt.plot(year_3rd, ssc_3rd)
plt.tight_layout()
plt.yticks([50,100,150,200], [50,100,150,200])
plt.ylabel('Sun Spot Count')
plt.subplot(4,1,4)
plt.plot(year_4th, ssc_4th)
plt.tight_layout()
plt.yticks([50,100,150,200], [50,100,150,200])
plt.xlabel('Year')
plt.ylabel('Sun Spot Count')
# + deletable=false nbgrader={"checksum": "332b489afbabd6c48e3456fb8db4ee88", "grade": true, "grade_id": "matplotlibex01e", "points": 4}
assert True # leave for grading
# -
|
assignments/assignment04/MatplotlibEx01.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Reversed Linked List
# [link](https://www.algoexpert.io/questions/Reverse%20Linked%20List)
# ## My Solution
def reverseLinkedList(head):
# Write your code here.
prv, cur = None, head
while cur is not None:
nxt = cur.next
cur.next = prv
prv = cur
cur = nxt
return prv
# +
def reverseLinkedList(head):
# Write your code here.
return reverseListHelper(None, head, head.next)
# O(n) time | O(n) space
def reverseListHelper(prevNode, currentNode, nextNode):
currentNode.next = prevNode
if nextNode is None:
return currentNode
return reverseListHelper(currentNode, nextNode, nextNode.next)
# -
# ## Expert Solution
# O(n) time | O(1) space - where n is the number of nodes in the Linked List
def reverseLinkedList(head):
previousNode, currentNode = None, head
while currentNode is not None:
nextNode = currentNode.next
currentNode.next = previousNode
previousNode = currentNode
currentNode = nextNode
return previousNode
# ## Thoughts
# ### my solution 2
# this is not a space efficient solution as the optimal solution. the O(n) space comes from the call stack of the recursive function. but this solution is very clear to show the prev, current and next Nodes shifting process.
|
algoExpert/reverse_linked_list/solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Using MediaEmailer Module
# #### 1.First create an folder named Attachment where you are using the script Insert Required Attachments to be sent in Attachments folder
# #### 2.Import MediaEmailer
from MediaEmailer import MediaEmailer
# #### 3. Create email details
# +
#enter user's gmail account name
sendersEmail = "<EMAIL>"
#enter user's gmail password
sendersPassword = "<PASSWORD>"
#enter in recipients' email address
recipients = ["<EMAIL>","<EMAIL>","<EMAIL>"]
#enter in baby's status
childName = "<NAME>"
date = "January 15, 2016"
time = "11:00 AM"
weight = "8 pounds, 7 ounce"
height = "11 inches"
# -
# #### 4. Use MediaEmailer.send_birth_email_with_attachments function
#
MediaEmailer.send_birth_email_with_attachments(sendersEmail,sendersPassword,recipients,childName,date,time,weight,height)
|
UsingMediaEmailer.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import matplotlib.pyplot as plt
plt.figure(figsize = (5,5))
case="01"
testcase=case+".txt"
with open(testcase, "r") as f:
s = f.readlines()
for line in s:
xleft,ybottom,xright,ytop = list(map(float, line.split()))
points = [(xleft,ybottom), (xleft,ytop), (xright, ytop), (xright, ybottom), (xleft,ybottom)]
points = list(zip(*points))
plt.plot(points[0], points[1], color = 'red')
plt.savefig(case+".png")
# -
(43-6)*(42-11)
|
Assignment-1/Documentation/test/plot_show.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # IF Else
# +
num1 = int(input("Enter Number1: "))
num2 = int(input("Enter Number2: "))
if (num1 > num2):
print("num1 '",num1, "' is greater than num2' ", num2, "'")
elif (num2 > num1):
print("num2 '",num2, "' is greater than num1' ", num1, "'")
else:
print("num1 '",num1, "' is equal to num2' ", num2, "'")
# -
# # Loops
#
# ## While Loop
# +
#While loop does the exactly same thing what "if statement" does, but instead of running the code block once,
#they jump back to the point where it began the code and repeats the whole process again.
#===============================================================
count = 0
while (count<3):
count += 1
print("I am called ", count, " time")
print("\n")
# Combining while with else
count = 0
while (count<3):
count += 1
print("I am called ", count, " time")
else:
print("While Loop Exited")
print("\n")
#--- when we dont know how many loops do our program needs to process in advance, we use while instead of for loop
#--- in the below example we might not know in advance how many loops do we need hence using while loop
#---do the sum of positive numbers in the list until first negative number met
list1 = [1,2,3,4,-4,-5]
i=0
sum1=0
while (list1[i]>0):
sum1 += list1[i]
i +=1;
print(sum1)
# -
# ## For Loop
# +
#In Python, "for loops" are called iterators.
#Just like while loop, "For Loop" is also used to repeat the program.
#But unlike while loop which depends on condition true or false.
#"For Loop" depends on the elements it has to iterate.
#===============================================================
# with numbers
for x in range (10,20,2):
print(x)
print("\n")
#with strings
Months = ["Jan","Feb","Mar","April","May","June"]
for month in Months:
print(month)
print("\n")
print("use the break statement")
# use the break statement
for x in range (10,20,1):
if (x==16):
break #terminate for loop at 16
elif (x%2 == 0):
print(x)
print("\n")
print("use the continue statement")
# use the continue statement
for x in range (10,20,1):
if (x == 14):
continue #skip '14'
print(x)
|
02_IfElse_and_Loops.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import json
import re
from flask import Flask, request
import jsonpickle
from config_src import config
# -
def get_snippets():
document = json.loads(request.json, encoding="utf-8")
#exclude cases when only one-words query!!!!!
documents = document["documents"]
terms = []
for term in document["terms"]:
terms.append(term["inverted_index"][0]["pos"][0])
result_lists = []
for doc in documents:
'''
if terms[0] < 20:
snippet = doc["text"][0:min(240, len(doc["text"]) - 1)]
elif terms[0] > 20:
snippet = doc["text"][terms[0]:min(terms[0]+240, len(doc["text"]) - 1)]
'''
snippet = doc["text"][0:len(doc["text"] - 1)]
doc["snippet"] = doc["text"]
search_res = dict()
search_res["results"] = documents
return json.dumps(search_res, ensure_ascii=False)
# +
app = Flask(__name__)
@app.route('/snippets', methods=["POST"])
def get_snippets():
"""
:param dict params: Like {"documents": [list of Documents],
"terms": list of dicts [{"term": "word1",
"inverted_index": [dict1, dict2, ...]}]}
:return list of Documents documents: With updated snippet attributes
"""
params = jsonpickle.decode(request.json)
documents = params["documents"]
query = params["query"]
#query = " ".join([i["term"] for i in search_terms])
for doc in documents:
doc.snippet = doc.text[:min(240, len(doc.text))]
return jsonpickle.encode(documents)
# -
if __name__ == "__main__":
app.run(host=config.SNIPPETS_HOST, port=config.SNIPPETS_PORT)
|
src/server_snippets.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Text classification
#
# Some use cases -
# * Understanding audience sentiment from social media,
# * Detection of spam and non-spam emails,
# * Auto tagging of customer queries, and
# * Categorization of news articles into defined topics.
#
# An end to end text classification pipeline is composed of three main components
# * Step 1: Dataset preparation:
# - Loading, preprocessing and splitting into train & test
#
# * Step 2: Feature engineering:
# - Raw dataset is transformed into features which can be used by ML models. Creating new features.
#
# * Step 3: Model training:
# - Training on a labelled dataset
#
# * Step 4: Improve performance of text classifier
#
# ----
# #### 1. Dataset preparation
# <b> 1.1 Loading the libraries & dataset </b>
# +
from sklearn import model_selection, preprocessing, linear_model, naive_bayes, metrics, svm
from sklearn.feature_extraction.text import TfidfVectorizer, CountVectorizer
from sklearn import decomposition, ensemble
import pandas, xgboost, numpy, textblob, string
# +
# load the dataset
data = open('dataset/corpus', encoding="utf8").read()
labels, texts = [], []
for i, line in enumerate(data.split("\n")):
content = line.split()
labels.append(content[0])
texts.append(" ".join(content[1:]))
# create a dataframe using texts and lables
trainDF = pandas.DataFrame()
trainDF['text'] = texts
trainDF['label'] = labels
# -
# <b> 1.2 Split the dataset into training and testing </b>
# +
# split the dataset into training and validation datasets
train_x, valid_x, train_y, valid_y = model_selection.train_test_split(trainDF['text'], trainDF['label'])
# label encode the target variable
encoder = preprocessing.LabelEncoder()
train_y = encoder.fit_transform(train_y)
valid_y = encoder.fit_transform(valid_y)
# -
# ---
# #### 2. Feature Engineering
#
# In this step, raw text data will be transformed into feature vectors and new features will be created using the existing dataset.
#
# <b> 2.1 Count Vectors as features </b>
# +
# create a count vectorizer object
count_vect = CountVectorizer(analyzer='word', token_pattern=r'\w{1,}')
count_vect.fit(trainDF['text'])
# transform the training and validation data using count vectorizer object
xtrain_count = count_vect.transform(train_x)
xvalid_count = count_vect.transform(valid_x)
# -
# <b> 2.2 TF-IDF Vectors as features
#
# TF-IDF Vectors can be generated at different levels of input tokens (words, characters, n-grams)
#
# a. Word Level TF-IDF : Matrix representing tf-idf scores of every term in different documents <br>
# b. N-gram Level TF-IDF : N-grams are the combination of N terms together. This Matrix representing tf-idf scores of N-grams <br>
# c. Character Level TF-IDF : Matrix representing tf-idf scores of character level n-grams in the corpus
# +
# word level tf-idf
tfidf_vect = TfidfVectorizer(analyzer='word', token_pattern=r'\w{1,}', max_features=5000)
tfidf_vect.fit(trainDF['text'])
xtrain_tfidf = tfidf_vect.transform(train_x)
xvalid_tfidf = tfidf_vect.transform(valid_x)
# ngram level tf-idf
tfidf_vect_ngram = TfidfVectorizer(analyzer='word', token_pattern=r'\w{1,}', ngram_range=(2,3), max_features=5000)
tfidf_vect_ngram.fit(trainDF['text'])
xtrain_tfidf_ngram = tfidf_vect_ngram.transform(train_x)
xvalid_tfidf_ngram = tfidf_vect_ngram.transform(valid_x)
# characters level tf-idf
tfidf_vect_ngram_chars = TfidfVectorizer(analyzer='char', token_pattern=r'\w{1,}', ngram_range=(2,3), max_features=5000)
tfidf_vect_ngram_chars.fit(trainDF['text'])
xtrain_tfidf_ngram_chars = tfidf_vect_ngram_chars.transform(train_x)
xvalid_tfidf_ngram_chars = tfidf_vect_ngram_chars.transform(valid_x)
# -
# <b> 2.3 Topic Modeling as features </b>
# +
# train a LDA Model
lda_model = decomposition.LatentDirichletAllocation(n_components=20, learning_method='online', max_iter=20)
X_topics = lda_model.fit_transform(xtrain_count)
topic_word = lda_model.components_
vocab = count_vect.get_feature_names()
# view the topic models
n_top_words = 10
topic_summaries = []
for i, topic_dist in enumerate(topic_word):
topic_words = numpy.array(vocab)[numpy.argsort(topic_dist)][:-(n_top_words+1):-1]
topic_summaries.append(' '.join(topic_words))
# -
# -------
# #### 3. Model Building
# The final step in the text classification framework is to train a classifier using the features created in the previous step
#
# * Naives Bayes Classifier
def train_model(classifier, feature_vector_train, label, feature_vector_valid, is_neural_net=False):
# fit the training dataset on the classifier
classifier.fit(feature_vector_train, label)
# predict the labels on validation dataset
predictions = classifier.predict(feature_vector_valid)
if is_neural_net:
predictions = predictions.argmax(axis=-1)
return metrics.accuracy_score(predictions, valid_y)
# +
# Naive Bayes on Count Vectors
accuracy = train_model(naive_bayes.MultinomialNB(), xtrain_count, train_y, xvalid_count)
print("NB, Count Vectors: ", accuracy)
# Naive Bayes on Word Level TF IDF Vectors
accuracy = train_model(naive_bayes.MultinomialNB(), xtrain_tfidf, train_y, xvalid_tfidf)
print("NB, WordLevel TF-IDF: ", accuracy)
# Naive Bayes on Ngram Level TF IDF Vectors
accuracy = train_model(naive_bayes.MultinomialNB(), xtrain_tfidf_ngram, train_y, xvalid_tfidf_ngram)
print("NB, N-Gram Vectors: ", accuracy)
# Naive Bayes on Character Level TF IDF Vectors
accuracy = train_model(naive_bayes.MultinomialNB(), xtrain_tfidf_ngram_chars, train_y, xvalid_tfidf_ngram_chars)
print("NB, CharLevel Vectors: ", accuracy)
# -
# * Linear Classifier
# +
# Linear Classifier on Count Vectors
accuracy = train_model(linear_model.LogisticRegression(), xtrain_count, train_y, xvalid_count)
print("LR, Count Vectors: ", accuracy)
# Linear Classifier on Word Level TF IDF Vectors
accuracy = train_model(linear_model.LogisticRegression(), xtrain_tfidf, train_y, xvalid_tfidf)
print("LR, WordLevel TF-IDF: ", accuracy)
# Linear Classifier on Ngram Level TF IDF Vectors
accuracy = train_model(linear_model.LogisticRegression(), xtrain_tfidf_ngram, train_y, xvalid_tfidf_ngram)
print("LR, N-Gram Vectors: ", accuracy)
# Linear Classifier on Character Level TF IDF Vectors
accuracy = train_model(linear_model.LogisticRegression(), xtrain_tfidf_ngram_chars, train_y, xvalid_tfidf_ngram_chars)
print("LR, CharLevel Vectors: ", accuracy)
# -
# * SVM Model
# SVM on Ngram Level TF IDF Vectors
accuracy = train_model(svm.SVC(), xtrain_tfidf_ngram, train_y, xvalid_tfidf_ngram)
print("SVM, N-Gram Vectors: ", accuracy)
# * Bagging Model
# +
# RF on Count Vectors
accuracy = train_model(ensemble.RandomForestClassifier(), xtrain_count, train_y, xvalid_count)
print("RF, Count Vectors: ", accuracy)
# RF on Word Level TF IDF Vectors
accuracy = train_model(ensemble.RandomForestClassifier(), xtrain_tfidf, train_y, xvalid_tfidf)
print("RF, WordLevel TF-IDF: ", accuracy)
# -
# * Boosting Model
# +
# Extereme Gradient Boosting on Count Vectors
accuracy = train_model(xgboost.XGBClassifier(), xtrain_count.tocsc(), train_y, xvalid_count.tocsc())
print("Xgb, Count Vectors: ", accuracy)
# Extereme Gradient Boosting on Word Level TF IDF Vectors
accuracy = train_model(xgboost.XGBClassifier(), xtrain_tfidf.tocsc(), train_y, xvalid_tfidf.tocsc())
print("Xgb, WordLevel TF-IDF: ", accuracy)
# Extereme Gradient Boosting on Character Level TF IDF Vectors
accuracy = train_model(xgboost.XGBClassifier(), xtrain_tfidf_ngram_chars.tocsc(), train_y, xvalid_tfidf_ngram_chars.tocsc())
print("Xgb, CharLevel Vectors: ", accuracy)
# -
# #### 4. Improving Text Classification Models
#
# While the above framework can be applied to a number of text classification problems, but to achieve a good accuracy some improvements can be done in the overall framework. For example, following are some tips to improve the performance of text classification models and this framework.
#
# 1. Text Cleaning : text cleaning can help to reducue the noise present in text data in the form of stopwords, punctuations marks, suffix variations etc. This article can help to understand how to implement text classification in detail.
#
# 2. Hstacking Text / NLP features with text feature vectors : In the feature engineering section, we generated a number of different feature vectros, combining them together can help to improve the accuracy of the classifier.
#
# 3. Hyperparamter Tuning in modelling : Tuning the paramters is an important step, a number of parameters such as tree length, leafs, network paramters etc can be fine tuned to get a best fit model.
#
# 4. Ensemble Models : Stacking different models and blending their outputs can help to further improve the results. Read more about ensemble models here
#
# -------
|
notebooks/NLP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %pylab inline
largestPrimeFactor = 2
def largest_prime_factor(num):
global largestPrimeFactor
for i in range(2, num+1):
if num%i == 0:
if largestPrimeFactor < i:
largestPrimeFactor = i
num = largest_prime_factor(int(num/i))
break
def largest_prime_factor_of(num):
global largestPrimeFactor
largestPrimeFactor = 1
largest_prime_factor(num)
print(largestPrimeFactor)
# %time largest_prime_factor_of(600851475143)
|
Lokesh Tiwari/.ipynb_checkpoints/Project_Euler_problem_3-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Load
#
# In this exercise, you'll load data into different formats: a csv file, a json file, and a SQLite database.
#
# You'll work with the GDP, population, and projects data. Run the code cell below to read in and clean the World Bank population and gdp data. This code creates a dataframe called df_indicator with both the gdp and population data.
# +
# run this code cell - there is nothing for you to do in this code cell
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# read in the projects data set and do basic wrangling
gdp = pd.read_csv('../data/gdp_data.csv', skiprows=4)
gdp.drop(['Unnamed: 62', 'Indicator Name', 'Indicator Code'], inplace=True, axis=1)
population = pd.read_csv('../data/population_data.csv', skiprows=4)
population.drop(['Unnamed: 62', 'Indicator Name', 'Indicator Code'], inplace=True, axis=1)
# Reshape the data sets so that they are in long format
gdp_melt = gdp.melt(id_vars=['Country Name', 'Country Code'],
var_name='year',
value_name='gdp')
# Use back fill and forward fill to fill in missing gdp values
gdp_melt['gdp'] = gdp_melt.sort_values('year').groupby(['Country Name', 'Country Code'])['gdp'].fillna(method='ffill').fillna(method='bfill')
population_melt = population.melt(id_vars=['Country Name', 'Country Code'],
var_name='year',
value_name='population')
# Use back fill and forward fill to fill in missing population values
population_melt['population'] = population_melt.sort_values('year').groupby('Country Name')['population'].fillna(method='ffill').fillna(method='bfill')
# merge the population and gdp data together into one data frame
df_indicator = gdp_melt.merge(population_melt, on=('Country Name', 'Country Code', 'year'))
# filter out values that are not countries
non_countries = ['World',
'High income',
'OECD members',
'Post-demographic dividend',
'IDA & IBRD total',
'Low & middle income',
'Middle income',
'IBRD only',
'East Asia & Pacific',
'Europe & Central Asia',
'North America',
'Upper middle income',
'Late-demographic dividend',
'European Union',
'East Asia & Pacific (excluding high income)',
'East Asia & Pacific (IDA & IBRD countries)',
'Euro area',
'Early-demographic dividend',
'Lower middle income',
'Latin America & Caribbean',
'Latin America & the Caribbean (IDA & IBRD countries)',
'Latin America & Caribbean (excluding high income)',
'Europe & Central Asia (IDA & IBRD countries)',
'Middle East & North Africa',
'Europe & Central Asia (excluding high income)',
'South Asia (IDA & IBRD)',
'South Asia',
'Arab World',
'IDA total',
'Sub-Saharan Africa',
'Sub-Saharan Africa (IDA & IBRD countries)',
'Sub-Saharan Africa (excluding high income)',
'Middle East & North Africa (excluding high income)',
'Middle East & North Africa (IDA & IBRD countries)',
'Central Europe and the Baltics',
'Pre-demographic dividend',
'IDA only',
'Least developed countries: UN classification',
'IDA blend',
'Fragile and conflict affected situations',
'Heavily indebted poor countries (HIPC)',
'Low income',
'Small states',
'Other small states',
'Not classified',
'Caribbean small states',
'Pacific island small states']
# remove non countries from the data
df_indicator = df_indicator[~df_indicator['Country Name'].isin(non_countries)]
df_indicator.reset_index(inplace=True, drop=True)
df_indicator.columns = ['countryname', 'countrycode', 'year', 'gdp', 'population']
# output the first few rows of the data frame
df_indicator.head()
# -
# Run this code cell to read in the countries data set. This will create a data frame called df_projects containing the World Bank projects data. The data frame only has the 'id', 'countryname', 'countrycode', 'totalamt', and 'year' columns.
# +
# run this code cell - there is nothing for you to do here
# !pip install pycountry
from pycountry import countries
# read in the projects data set with all columns type string
df_projects = pd.read_csv('../data/projects_data.csv', dtype=str)
df_projects.drop(['Unnamed: 56'], axis=1, inplace=True)
df_projects['countryname'] = df_projects['countryname'].str.split(';').str.get(0)
# set up the libraries and variables
from collections import defaultdict
country_not_found = [] # stores countries not found in the pycountry library
project_country_abbrev_dict = defaultdict(str) # set up an empty dictionary of string values
# TODO: iterate through the country names in df_projects.
# Create a dictionary mapping the country name to the alpha_3 ISO code
for country in df_projects['countryname'].drop_duplicates().sort_values():
try:
# TODO: look up the country name in the pycountry library
# store the country name as the dictionary key and the ISO-3 code as the value
project_country_abbrev_dict[country] = countries.lookup(country).alpha_3
except:
# If the country name is not in the pycountry library, then print out the country name
# And store the results in the country_not_found list
country_not_found.append(country)
# run this code cell to load the dictionary
country_not_found_mapping = {'Co-operative Republic of Guyana': 'GUY',
'Commonwealth of Australia':'AUS',
'Democratic Republic of Sao Tome and Prin':'STP',
'Democratic Republic of the Congo':'COD',
'Democratic Socialist Republic of Sri Lan':'LKA',
'East Asia and Pacific':'EAS',
'Europe and Central Asia': 'ECS',
'Islamic Republic of Afghanistan':'AFG',
'Latin America':'LCN',
'Caribbean':'LCN',
'Macedonia':'MKD',
'Middle East and North Africa':'MEA',
'Oriental Republic of Uruguay':'URY',
'Republic of Congo':'COG',
"Republic of Cote d'Ivoire":'CIV',
'Republic of Korea':'KOR',
'Republic of Niger':'NER',
'Republic of Kosovo':'XKX',
'Republic of Rwanda':'RWA',
'Republic of The Gambia':'GMB',
'Republic of Togo':'TGO',
'Republic of the Union of Myanmar':'MMR',
'Republica Bolivariana de Venezuela':'VEN',
'Sint Maarten':'SXM',
"Socialist People's Libyan Arab Jamahiriy":'LBY',
'Socialist Republic of Vietnam':'VNM',
'Somali Democratic Republic':'SOM',
'South Asia':'SAS',
'St. Kitts and Nevis':'KNA',
'St. Lucia':'LCA',
'St. Vincent and the Grenadines':'VCT',
'State of Eritrea':'ERI',
'The Independent State of Papua New Guine':'PNG',
'West Bank and Gaza':'PSE',
'World':'WLD'}
project_country_abbrev_dict.update(country_not_found_mapping)
df_projects['countrycode'] = df_projects['countryname'].apply(lambda x: project_country_abbrev_dict[x])
df_projects['boardapprovaldate'] = pd.to_datetime(df_projects['boardapprovaldate'])
df_projects['year'] = df_projects['boardapprovaldate'].dt.year.astype(str).str.slice(stop=4)
df_projects['totalamt'] = pd.to_numeric(df_projects['totalamt'].str.replace(',',""))
df_projects = df_projects[['id', 'countryname', 'countrycode', 'totalamt', 'year']]
df_projects.head()
# -
# # Exercise Part 1
#
# The first few cells in this workbook loaded and cleaned the World Bank Data. You now have two data frames:
# * df_projects, which contain data from the projects data set
# * df_indicator, which contain population and gdp data for various years
#
# They both have country code variables. Note, however, that there could be countries represented in the projects data set that are not in the indicator data set and vice versus.
#
# In this first exercise, merge the two data sets together using country code and year as common keys. When joining the data sets, keep all of the data in the df_projects dataframe even if there is no indicator data for that country code.
# +
# TODO: merge the projects and indicator data frames together using countrycode and year as common keys
# Use a left join so that all projects are returned even if the country/year combination does not have
# indicator data
df_merged = df_projects.merge(df_indicator, on=['countrycode', 'year'], how='left')
# df_merged.shape
# -
# If you look at the first few rows of the merged data set, you'll see NaN's for the indicator data. That is because the indicator data does not contain values for 2018. If you run the code cell below, you should get the following results:
#
# |id | countryname_x| countrycode|totalamt|year|countryname_y|gdp| population|
# |:-:|:-:|:-:|:-:|:-:|:-:|:-:|:-:|
# |256|P161982|Hashemite Kingdom of Jordan|JOR|0|2017|Jordan|4.006831e+10|9702353.0|
# |301|P162407|Hashemite Kingdom of Jordan|JOR|147700000|2017|Jordan|4.006831e+10|9702353.0|
# |318|P160103|Hashemite Kingdom of Jordan|JOR|0|2017|Jordan|4.006831e+10|9702353.0|
# |464|P161905|Hashemite Kingdom of Jordan|JOR|50000000|2017|Jordan|4.006831e+10|9702353.0|
# |495|P163387|Hashemite Kingdom of Jordan|JOR|36100000|2017|Jordan|4.006831e+10|9702353.0|
# |515|P163241|Hashemite Kingdom of Jordan|JOR|0|2017|Jordan|4.006831e+10|9702353.0|
# Run this code to check your work
df_merged[(df_merged['year'] == '2017') & (df_merged['countryname_y'] == 'Jordan')]
# # Exercise Part 2
#
# Output the df_merged dataframe as a json file. You can use the pandas [to_json() method](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.to_json.html).
# +
import os
os.getcwd()
# -
os.listdir()
# +
# TODO: Output the df_merged data frame as a json file
# HINT: Pandas has a to_json() method
# HINT: use orient='records' to get one of the more common json formats
# HINT: be sure to specify the name of the json file you want to create as the first input into to_json
df_merged.to_json('countrydata.json', orient='records')
# -
# If you go to File->Open and then open the 17_load_exercise folder, you can see the json file you created.
#
# The records should look like this (note that the order of the records might be different, but the format should look similar):
# ```
# [{"id":"P162228","countryname_x":"World","countrycode":"WLD","totalamt":0,"year":"2018","countryname_y":null,"gdp":null,"population":null},{"id":"P163962","countryname_x":"Democratic Republic of the Congo","countrycode":"COD","totalamt":200000000,"year":"2018","countryname_y":null,"gdp":null,"population":null},{"id":"P167672","countryname_x":"People's Republic of Bangladesh","countrycode":"BGD","totalamt":58330000,"year":"2018","countryname_y":null,"gdp":null,"population":null},{"id":"P158768","countryname_x":"Islamic Republic of Afghanistan","countrycode":"AFG","totalamt":20000000,"year":"2018","countryname_y":null,"gdp":null,"population":null},{"id":"P161364","countryname_x":"Federal Republic of Nigeria","countrycode":"NGA","totalamt":100000000,"year":"2018","countryname_y":null,"gdp":null,"population":null},{"id":"P161483","countryname_x":"Republic of Tunisia","countrycode":"TUN","totalamt":500000000,"year":"2018","countryname_y":null,"gdp":null,"population":null}
# ```
#
#
# Check
pd.read_json('countrydata.json')
# # Exercise Part 3
#
# Output the df_merged dataframe as a csv file. You can use the pandas [to_csv() method](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.to_csv.html).
# +
# TODO: Output the df_merged data frame as a csv file
# HINT: The to_csv() method is similar to the to_json() method.
# HINT: If you do not want the data frame indices in your result, use index=False
df_merged.to_csv('countrydata.csv', index=False)
# -
# If you go to File->Open and then open the 17_load_exercise folder, you can see the csv file you created.
#
# The records should look something like this:
# ```
# id,countryname_x,countrycode,totalamt,year,countryname_y,gdp,population
# P162228,World,WLD,0,2018,,,
# P163962,Democratic Republic of the Congo,COD,200000000,2018,,,
# P167672,People's Republic of Bangladesh,BGD,58330000,2018,,,
# P158768,Islamic Republic of Afghanistan,AFG,20000000,2018,,,
# P161364,Federal Republic of Nigeria,NGA,100000000,2018,,,
# P161483,Republic of Tunisia,TUN,500000000,2018,,,
# P161885,Federal Republic of Nigeria,NGA,350000000,2018,,,
# P162069,Federal Republic of Nigeria,NGA,225000000,2018,,,
# P163540,Federal Republic of Nigeria,NGA,125000000,2018,,,
# P163576,Lebanese Republic,LBN,329900000,2018,,,
# P163721,Democratic Socialist Republic of Sri Lan,LKA,200000000,2018,,,
# P164082,Federal Republic of Nigeria,NGA,400000000,2018,,,
# P164783,Nepal,NPL,0,2018,,,
# ```
#
#
# Check
pd.read_csv('countrydata.csv')
# # Exercise Part 4
#
# Output the df_merged dataframe as a sqlite database file. For this exercise, you can put all of the data as one table. In the next exercise, you'll create a database with multiple tables.
# +
import sqlite3
# connect to the database
# the database file will be worldbank.db
# note that sqlite3 will create this database file if it does not exist already
conn = sqlite3.connect('worldbank.db')
# TODO: output the df_merged dataframe to a SQL table called 'merged'.
# HINT: Use the to_sql() method
# HINT: Use the conn variable for the connection parameter
# HINT: You can use the if_exists parameter like if_exists='replace' to replace a table if it already exists
df_merged.to_sql(name='merged', con=conn, if_exists='replace', index=False)
# -
# Run the code cell below to make sure that the worldbank.db file was created and the merged table loaded. You should get an output that is formatted like this:
#
# |id|countryname_x|countrycode|totalamt|year|countryname_y|gdp|population
# |-|-|-|-|-|-|-|-|
# |P162033|'Federative Republic of Brazil'|BRA|125000000|2017|Brazil|2.055506e+12|209288278.0|
pd.read_sql('SELECT * FROM merged WHERE year = "2017" AND countrycode = "BRA"', con = conn).head()
conn.close()
# # Exercise Part 5
#
# Output the data to a SQL database like in the previous exercise; however, this time, put the df_indicator data in one table and the df_projects data in another table. Call the df_indicator table 'indicator' and the df_projects table 'projects'.
# +
import sqlite3
# connect to the database
# the database file will be worldbank.db
# note that sqlite3 will create this database file if it does not exist already
conn = sqlite3.connect('worldbank.db')
# TODO: output the df_merged dataframe to a SQL table called 'merged'.
# HINT: Use the to_sql() method
# HINT: Use the conn variable for the connection parameter
# HINT: You can use the if_exists parameter like if_exists='replace' to replace a table if it already exists
df_indicator.to_sql(name='indicator', con=conn, if_exists='replace', index=False)
df_projects.to_sql(name='projects', con=conn, if_exists='replace', index=False)
# -
# Run the code cell below to see if your code is working properly. The code cell below runs a SQL query against the worldbank.db database joining the indicator table with the projects table.
# +
'''
pd.read_sql('SELECT * FROM projects LEFT JOIN indicator ON \
projects.countrycode = indicator.countrycode AND \
projects.year = indicator.year WHERE \
projects.year = "2017" AND projects.countrycode = "BRA"', con = conn).head()
'''
sql = """
SELECT * FROM projects
LEFT JOIN indicator
ON projects.countrycode = indicator.countrycode
AND projects.year = indicator.year
WHERE projects.year = "2017"
AND projects.countrycode = "BRA";
"""
pd.read_sql(sql, conn).head()
# -
# commit any changes to the database and close the connection to the database
conn.commit()
conn.close()
# # Exercise Part 4 & 5 (sqlalchemy)
# + tags=[]
from sqlalchemy import create_engine
# connect to the database
# the database file will be worldbank.db
# note that sqlite3 will create this database file if it does not exist already
engine = create_engine('sqlite:///worldbank.db')
conn = engine.raw_connection()
# TODO: output the df_merged dataframe to a SQL table called 'merged'.
# HINT: Use the to_sql() method
# HINT: Use the conn variable for the connection parameter
# HINT: You can use the if_exists parameter like if_exists='replace' to replace a table if it already exists
df_merged.to_sql(name='merged', con=conn, if_exists='replace', index=False)
# -
pd.read_sql('SELECT * FROM merged WHERE year = "2017" AND countrycode = "BRA"', con = conn).head()
df_indicator.to_sql(name='indicator', con=conn, if_exists='replace', index=False)
df_projects.to_sql(name='projects', con=conn, if_exists='replace', index=False)
# +
sql = """
SELECT * FROM projects
LEFT JOIN indicator
ON projects.countrycode = indicator.countrycode
AND projects.year = indicator.year
WHERE projects.year = "2017"
AND projects.countrycode = "BRA";
"""
pd.read_sql(sql, conn).head()
# -
# commit any changes to the database and close the connection to the database
conn.commit()
conn.close()
# # Exercise Part 6 (Challenge)
#
# SQLite, as its name would suggest, is somewhat limited in its functionality. For example, the Alter Table command only allows you to change a table name or to add a new column to a table. You can't, for example, add a primary key to a table once the table is already created.
#
# If you want more control over a sqlite3 database, it's better to use the sqlite3 library directly. Here is an example of how to use the sqlite3 library to create a table in the database, insert a value, and then run a SQL query on the database. Run the code cells below to see the example.
#
# ### Demo
# +
# connect to the data base
conn = sqlite3.connect('worldbank.db')
# get a cursor
cur = conn.cursor()
# drop the test table in case it already exists
cur.execute("DROP TABLE IF EXISTS test")
# create the test table including project_id as a primary key
cur.execute("CREATE TABLE test (project_id TEXT PRIMARY KEY, countryname TEXT, countrycode TEXT, totalamt REAL, year INTEGER);")
# insert a value into the test table
cur.execute("INSERT INTO test (project_id, countryname, countrycode, totalamt, year) VALUES ('a', 'Brazil', 'BRA', '100,000', 1970);")
# commit changes made to the database
conn.commit()
# select all from the test table
cur.execute("SELECT * FROM test")
cur.fetchall()
# -
# commit any changes and close the data base
conn.close()
# ### Exercise
#
# Now, it's your turn. Use the sqlite3 library to connect to the worldbank.db database. Then:
# * Create a table, called projects, for the projects data where the primary key is the id of each project.
# * Create another table, called gdp, that contains the gdp data.
# * And create another table, called population, that contains the population data.
#
# Here is the schema for each table.
# ##### projects
#
# * project_id text
# * countryname text
# * countrycode text
# * totalamt real
# * year integer
#
# project_id is the primary key
#
# ##### gdp
# * countryname text
# * countrycode text
# * year integer
# * gdp real
#
# (countrycode, year) is the primary key
#
# ##### population
# * countryname text
# * countrycode text
# * year integer
# * population integer
#
# (countrycode, year) is the primary key
#
# After setting up the tables, write code that inserts the data into each table. (Note that this database is not normalized. For example, countryname and countrycode are in all three tables. You could make another table with countrycode and countryname and then create a foreign key constraint in the projects, gdp, and population tables. If you'd like an extra challenge, create a country table with countryname and countrycode. Then create the other tables with foreign key constraints).
#
# Follow the TODOs in the next few code cells to finish the exercise.
# +
# connect to the data base
conn = sqlite3.connect('worldbank.db')
# get a cursor
cur = conn.cursor()
# drop tables created previously to start fresh
cur.execute("DROP TABLE IF EXISTS test")
cur.execute("DROP TABLE IF EXISTS indicator")
cur.execute("DROP TABLE IF EXISTS projects")
cur.execute("DROP TABLE IF EXISTS gdp")
cur.execute("DROP TABLE IF EXISTS population")
# TODO create the projects table including project_id as a primary key
# HINT: Use cur.execute("SQL Query")
sql = """
CREATE TABLE IF NOT EXISTS projects (
project_id TEXT PRIMARY KEY,
countryname TEXT,
countrycode TEXT,
totalamt REAL,
year INTEGER
);"""
cur.execute(sql)
# TODO: create the gdp table including (countrycode, year) as primary key
# HINT: To create a primary key on multiple columns, you can do this:
# CREATE TABLE tablename (columna datatype, columnb datatype, columnc dataype, PRIMARY KEY (columna, columnb));
sql = """
CREATE TABLE IF NOT EXISTS gdp (
countryname TEXT,
countrycode TEXT,
year INTEGER,
gdp REAL,
PRIMARY KEY (countrycode, year)
);"""
cur.execute(sql)
# TODO: create the population table including (countrycode, year) as primary key
sql = """
CREATE TABLE IF NOT EXISTS population (
countryname TEXT,
countrycode TEXT,
year INTEGER,
population INTEGER,
PRIMARY KEY (countrycode, year)
);"""
cur.execute(sql)
# commit changes to the database. Do this whenever modifying a database
conn.commit()
# +
# %%time
# TODO:insert project values into the projects table
# HINT: Use a for loop with the pandas iterrows() method
# HINT: The iterrows() method returns two values: an index for each row and a tuple of values
# HINT: Some of the values for totalamt and year are NaN. Because you've defined
# year and totalamt as numbers, you cannot insert NaN as a value into those columns.
# When totalamt or year equal NaN, you'll need to change the value to something numeric
# like, for example, zero
df = df_projects.copy()
# NaN in df_projects is 'nan'
# df[df.id == 'P000031'].year[15894] == 'nan'
df.replace({'nan': 'NULL'}, inplace=True)
for idx, row in df.iterrows():
project_id, countryname, countrycode, totalamt, year = row
sql = f"""
INSERT INTO projects (project_id, countryname, countrycode, totalamt, year)
VALUES ("{project_id}", "{countryname}", "{countrycode}", {totalamt}, {year});
"""
cur.execute(sql)
# commit changes to the dataset after any changes are made
conn.commit()
# +
# Check number of rows
assert len(pd.read_sql('SELECT * FROM projects;', conn)) == len(df_projects), \
'One or more rows were inserted into projects failed'
pd.read_sql('SELECT * FROM projects WHERE year is NULL', conn)
# +
# %%time
# TODO: insert gdp values into the gdp table
df = df_indicator.drop(columns=['population']).copy()
df.replace({'nan': 'NULL'}, inplace=True)
for idx, row in df.iterrows():
countryname, countrycode, year, gdp = row
sql = f"""
INSERT INTO gdp (countryname, countrycode, year, gdp)
VALUES ("{countryname}", "{countrycode}", {year}, {gdp});
"""
cur.execute(sql)
# commit changes to the dataset after any changes are made
conn.commit()
# +
# Check number of rows
assert len(pd.read_sql('SELECT * FROM gdp;', conn)) == len(df_indicator), \
'One or more rows were inserted into gdp failed'
pd.read_sql('SELECT * FROM gdp', conn)
# +
# %%time
# TODO: insert population values into the population table
df = df_indicator.drop(columns=['gdp']).copy()
df.replace({'nan': 'NULL'}, inplace=True)
for idx, row in df.iterrows():
countryname, countrycode, year, population = row
sql = f"""
INSERT INTO population (countryname, countrycode, year, population)
VALUES ("{countryname}", "{countrycode}", {year}, {population});
"""
cur.execute(sql)
# commit changes to the dataset after any changes are made
conn.commit()
# +
# Check number of rows
assert len(pd.read_sql('SELECT * FROM population;', conn)) == len(df_indicator), \
'One or more rows were inserted into population failed'
pd.read_sql('SELECT * FROM population;', conn)
# -
# run this command to see if your tables were loaded as expected
sql = """
SELECT * FROM projects
JOIN gdp
ON projects.countrycode = gdp.countrycode
AND projects.year = gdp.year
JOIN population
ON projects.countrycode = population.countrycode
AND projects.year = population.year;
"""
result = pd.read_sql(sql, con=conn)
result.shape
# If everything went well, the code above should output (15537, 13) as the shape of the resulting data frame.
# commit any changes and close the database
conn.commit()
conn.close()
# # Conclusion
#
# Once you have extracted data and transformed it, the final step is to load the data into storage. In this exercise, you stored results into a csv file, a json file, and a SQLite database.
|
lessons/ETLPipelines/17_load_exercise/17_load_exercise.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
# # Plot memory fingerprint
#
#
# This example plots a fingerprint. Briefly, a fingerprint
# can be described as a summary of how a subject organizes information with
# respect to the multiple features of the stimuli. In addition to presentation
# and recall data, a features object is passed to the Egg class. It is comprised
# of a dictionary for each presented stimulus that contains feature dimensions and
# values for each stimulus.
#
#
#
# +
# Code source: <NAME>
# License: MIT
#import
import quail
#load data
egg = quail.load('example')
# analyze and plot
egg.analyze('fingerprint', listgroup=['average']*8, features=['temporal']).plot(title='Memory Fingerprint')
|
docs/auto_examples/plot_fingerprint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Import pandas
import pandas as pd
# Load dataset
cc_apps = pd.read_csv("cc_approvals.data", header = None)
# Inspect data
cc_apps.head()
# +
# Print summary statistics
cc_apps_description = cc_apps.describe()
print(cc_apps_description)
print('\n')
# Print DataFrame information
cc_apps_info = cc_apps.info()
print(cc_apps_info)
print('\n')
# Inspect missing values in the dataset
cc_apps.isnull().sum()
# +
# Import train_test_split
from sklearn.model_selection import train_test_split
# Drop the features 11 and 13
cc_apps = cc_apps.drop([11, 13], axis = 1)
# Split into train and test sets
cc_apps_train, cc_apps_test = train_test_split(cc_apps, test_size=0.33, random_state=42)
# +
# Import numpy
import numpy as np
# Replace the '?'s with NaN in the train and test sets
cc_apps_train = cc_apps_train.replace("?",np.NaN)
cc_apps_test = cc_apps_test.replace("?",np.NaN)
# +
# Impute the missing values with mean imputation
cc_apps_train.fillna(cc_apps_train.mean(), inplace=True)
cc_apps_test.fillna(cc_apps_test.mean(), inplace=True)
# Count the number of NaNs in the datasets and print the counts to verify
print(cc_apps_train.isnull().sum())
print(cc_apps_test.isnull().sum())
# +
# Iterate over each column of cc_apps_train
for col in cc_apps_train:
# Check if the column is of object type
if cc_apps_train[col].dtypes == 'object':
# Impute with the most frequent value
cc_apps_train = cc_apps_train.fillna(cc_apps_train[col].value_counts().index[0])
cc_apps_test = cc_apps_test.fillna(cc_apps_test[col].value_counts().index[0])
# Count the number of NaNs in the dataset and print the counts to verify
print(cc_apps_train.isnull().sum())
print(cc_apps_test.isnull().sum())
print(cc_apps_train)
print(cc_apps_test)
# -
# Convert the categorical features in the train and test sets independently
cc_apps_train = pd.get_dummies(cc_apps_train)
cc_apps_test = pd.get_dummies(cc_apps_test)
# Reindex the columns of the test set aligning with the train set
cc_apps_test = cc_apps_test.reindex(columns=cc_apps_train.columns, fill_value=0)
# +
# Import MinMaxScaler
from sklearn.preprocessing import MinMaxScaler
# Segregate features and labels into separate variables
X_train, y_train = cc_apps_train.iloc[:,:-1].values, cc_apps_train.iloc[:,-1].values
X_test, y_test = cc_apps_test.iloc[:,:-1].values, cc_apps_test.iloc[:,-1].values
# Instantiate MinMaxScaler and use it to rescale X_train and X_test
scaler = MinMaxScaler(feature_range=(0,1))
rescaledX_train = scaler.fit_transform(X_train)
rescaledX_test = scaler.transform(X_test)
# +
# Import LogisticRegression
from sklearn.linear_model import LogisticRegression
# Instantiate a LogisticRegression classifier with default parameter values
logreg = LogisticRegression()
# Fit logreg to the train set
logreg.fit(rescaledX_train, y_train)
# +
# Import confusion_matrix
from sklearn.metrics import confusion_matrix
# Use logreg to predict instances from the test set and store it
y_pred = logreg.predict(rescaledX_test)
# Get the accuracy score of logreg model and print it
print("Accuracy of logistic regression classifier: ", logreg.score(rescaledX_test,y_test))
# Print the confusion matrix of the logreg model
confusion_matrix(y_test,y_pred)
# +
# Import GridSearchCV
from sklearn.model_selection import GridSearchCV
# Define the grid of values for tol and max_iter
tol = [0.01, 0.001 ,0.0001]
max_iter = [100, 150, 200]
# Create a dictionary where tol and max_iter are keys and the lists of their values are the corresponding values
param_grid = dict(tol=tol, max_iter=max_iter)
param_grid
# +
# Instantiate GridSearchCV with the required parameters
grid_model = GridSearchCV(estimator=logreg, param_grid=param_grid, cv=5)
# Fit grid_model to the data
grid_model_result = grid_model.fit(rescaledX_train,y_train)
# Summarize results
best_score, best_params = grid_model_result.best_score_, grid_model_result.best_params_
print("Best: %f using %s" % (best_score, best_params))
# Extract the best model and evaluate it on the test set
best_model = grid_model_result.best_estimator_
print("Accuracy of logistic regression classifier: ", best_model.score(rescaledX_test,y_test))
grid_model.predict(rescaledX_train)
# -
from sklearn.metrics import mean_absolute_error
# define mean absolute percentage error function for further calculations.
def MAPE(Y_actual,Y_Predicted):
mape = np.mean(np.abs((Y_actual - Y_Predicted)/Y_actual))*100
return mape
# +
print(f"Accuracy of training set:", best_model.score(X_train,y_train))
print(f"Accuracy of testing set: ", best_model.score(rescaledX_test,y_test))
# +
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=2)
knn.fit(X_train, y_train)
knn.score(X_train, y_train), knn.score(X_test, y_test)
y_knn_pred_test = knn.predict(X_test)
print(f"Accuracy of training set:", knn.score(X_train, y_train))
print(f"Accuracy of testing set: ", knn.score(X_test, y_test))
# +
from sklearn.linear_model import LogisticRegression
from sklearn.tree import DecisionTreeRegressor
dtree = DecisionTreeRegressor(random_state=1)
# Fit model
dtree.fit(X_train, y_train)
y_tree_pred = dtree.predict(X_test)
print(f"Accuracy of training set:", dtree.score(X_train, y_train))
print(f"Accuracy of testing set: ", dtree.score(X_test, y_test))
# +
from sklearn.ensemble import RandomForestRegressor
forest_model = RandomForestRegressor(random_state=1)
forest_model.fit(X_train, y_train)
y_forest_pred = forest_model.predict(X_test)
print(f"Accuracy of training set:", forest_model.score(X_train, y_train))
print(f"Accuracy of testing set: ", forest_model.score(X_test, y_test))
|
Credit-Card-Approvals-Utku-Sokat.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # VIPERS SHAM Project
#
# This notebook is part of the VIPERS-SHAM project:
# http://arxiv.org/abs/xxxxxxx
#
# Copyright 2019 by <NAME>, <EMAIL>
# All rights reserved.
# This file is released under the "MIT License Agreement". Please see the LICENSE
# file that should have been included as part of this package.
# +
# %matplotlib inline
import sys
import os
import logging
from matplotlib import pyplot as plt
plt.style.use("small.style")
from matplotlib.ticker import FormatStrFormatter,ScalarFormatter
from matplotlib import colors,cm, ticker
import logging
logging.basicConfig(level=logging.INFO)
import numpy as np
from scipy import interpolate, integrate
import emcee
import progressbar
import growthcalc
import load
import utils
import emulator
# -
nsteps = 1000
samples = ['sdss','L1','L2','L3','L4']
redshifts = {'sdss':.06, 'L1':0.6, 'L2':0.7, 'L3':0.8, 'L4':0.9}
rmin = 1
def decomp(cmat, thresh=.1):
u,s,v = np.linalg.svd(cmat)
cut = np.abs(s).max()*thresh
o = np.abs(s)>cut
s = s[o]
v = v[o]
return s, v
# +
r_sdss,wp_sdss,cov_sdss = load.load_sdss()
sel = r_sdss > rmin
r_sdss = r_sdss[sel]
wp_sdss = wp_sdss[sel]
cov_sdss = cov_sdss[sel,:][:,sel]
s,v = decomp(cov_sdss)
data = [(r_sdss, wp_sdss, s, v)]
for sample in samples[1:]:
r,wp = np.loadtxt('../data/vipers/wp_sM{sample}.txt'.format(sample=sample[1]), unpack=True)
cmat = np.loadtxt('../data/vipers/cov_{sample}.txt'.format(sample=sample))
sel = r > rmin
r = r[sel]
wp = wp[sel]
cmat = cmat[sel,:][:,sel]
s, v = decomp(cmat)
data.append((r, wp, s, v))
# -
shamdata = {}
for sample in ['sdss','L1','L2','L3','L4']:
sham = load.load_sham(sample=sample, template="../data/sham400/nz_{sample}/wp_snap{snapshot:7.5f}.txt")
snapshots = sham.keys()
snapshots.sort()
for key in snapshots:
r, wp = sham[key]
sel = r > 1
r = r[sel]
wp = wp[sel]
if not sample in shamdata:
shamdata[sample] = []
shamdata[sample].append((key, r, wp))
a_samples = []
interpolators = []
for key in samples:
y = []
x = []
for a,r,w in shamdata[key]:
sel = r > 1
r = r[sel]
y.append(w[sel])
x.append(a)
y = np.array(y)
x = np.array(x)
f = emulator.WpInterpolator(x, r, y)
interpolators.append(f)
a_samples.append(1./(1+redshifts[key]))
a_samples = np.array(a_samples)
emu = emulator.Emulator(a_samples, interpolators)
def conf_interval(x, y, p=0.68, alpha = 0.99):
""" """
best = y.argmax()
x0 = x[best]
ybest = y[best]
h = ybest
t = np.sum(y)*p
while np.sum(y[y>h]) < t:
h *= alpha
s = x[np.where(y>h)]
low = s[0]
high = s[-1]
return x0, x0-low, high-x0
# +
def loglike(p, emu, data):
""" """
sig8, gamma = p
if gamma < -5:
return -np.infty
if gamma > 5:
return -np.infty
if sig8 < 0:
return -np.infty
try:
model = emu(gamma, sig8)
except ValueError:
return -np.infty
assert len(model) == len(data)
chi2 = 0
for i in range(len(data)):
r,wp,s,v = data[i]
rsham, wsham = model[i]
d = np.dot(v, wp - wsham)
chi2 += np.sum(d**2 / s)
return -chi2/2.
def loglike_gamma(p, emu, data):
""" """
gamma = p
if gamma < -5:
return -np.infty
if gamma > 5:
return -np.infty
try:
model = emu(gamma, None)
except ValueError:
return -np.infty
assert len(model) == len(data)
chi2 = 0
for i in range(len(data)):
r,wp,s,v = data[i]
rsham, wsham = model[i]
d = np.dot(v, wp - wsham)
chi2 += np.sum(d**2 / s)
return -chi2/2.
def loglike_s8(p, emu, data):
""" """
s8 = p
try:
model = emu(None, s8)
except ValueError:
return -np.infty
assert len(model) == len(data)
chi2 = 0
for i in range(len(data)):
r,wp,s,v = data[i]
rsham, wsham = model[i]
d = np.dot(v, wp - wsham)
chi2 += np.sum(d**2 / s)
return -chi2/2.
# -
def run_chain(sampler, p0, nsteps=1000):
""" """
bar = progressbar.ProgressBar(max_value=nsteps)
for i, result in enumerate(sampler.sample(p0, iterations=nsteps)):
bar.update(i)
return result
# +
ndim, nwalkers = 2, 100
p0 = np.random.normal(0, 0.01, (nwalkers,ndim)) + np.array([0.8, 0.55])
sampler = emcee.EnsembleSampler(nwalkers, ndim, loglike, args=[emu, data])
pos, prob, state = run_chain(sampler, p0, 100)
sampler.reset()
pos, prob, state = run_chain(sampler, pos, nsteps)
# -
gamma = np.linspace(-1,2.5,2000)
ll = np.zeros(len(gamma))
bar = progressbar.ProgressBar()
for i in bar(range(len(gamma))):
ll[i] = loglike_gamma(gamma[i], emu, data)
ll = np.exp(np.array(ll))
ll = ll / np.sum(ll) / (gamma[1]-gamma[0])
a,b,c = conf_interval(gamma, ll)
print a,b,c
print a, np.sqrt((b**2+c**2)/2.)
sig8 = np.linspace(0.5,1.2, 2000)
ll_s8 = np.zeros(len(sig8))
bar = progressbar.ProgressBar()
for i in bar(range(len(sig8))):
ll_s8[i] = loglike_s8(sig8[i], emu, data)
ll_s8 = np.exp(np.array(ll_s8))
ll_s8 = ll_s8 / np.sum(ll_s8) / (sig8[1]-sig8[0])
a,b,c = conf_interval(sig8, ll_s8)
print a,b,c
print a, np.sqrt((b**2+c**2)/2.)
def plot_lim(ax, t, g, s):
t = np.array(t,dtype='d')
xx = []
yy = []
for i,c in enumerate(t):
k = c.searchsorted(1)
if k > 0 and k < len(s):
yy.append(s[k])
xx.append(g[i])
ax.plot(xx,yy, c='k', dashes=[1,1,1,1])
# +
from matplotlib import gridspec
bins = (
np.linspace(0.7, 1.02, 20),
np.linspace(-0.5, 1.5, 20)
)
ext = [bins[1][0],bins[1][-1],bins[0][0],bins[0][-1]]
h, ex, ey = np.histogram2d(*sampler.flatchain.T, bins=bins)
h = h.T
h_sig,e_s = np.histogram(sampler.flatchain.T[0], bins=bins[0], density=True)
h_gam,e_g = np.histogram(sampler.flatchain.T[1], bins=bins[1], density=True)
levels = utils.lowerwater(h, (0.866, 0.395))
a = (ey[1]-ey[0])/(ex[1]-ex[0])
fig = plt.figure(figsize=(4.2,4))
gs = gridspec.GridSpec(2, 2)
gs.update(right=0.9, top=0.9, hspace=0,wspace=0)
ax1 = plt.subplot(gs[1,0])
ax2 = plt.subplot(gs[0,0])
ax3 = plt.subplot(gs[1,1])
g = np.linspace(-0.5,1.5,200)
s = np.linspace(0.5,1.2,200)
zmin,zmax = emu.check_bounds(g,s)
plot_lim(ax1, zmin<-0.3, g, s)
plot_lim(ax1, zmax<1.5, g, s)
ax1.text(-0.2,0.78,"$z>1.5$", rotation=-65)
ax1.text(0.8,0.99,"$z<-0.3$", rotation=-5)
ax1.contour(h.T, levels=levels, colors='k', extent=ext, origin='lower')
ax1.axvline(0.55, dashes=[3,1,1,1], c='firebrick')
ax1.axhline(0.82, dashes=[4,1], c='navy')
ax1.set_xlim(ext[0],ext[1])
ax1.set_ylim(ext[2],ext[3])
ax2.plot((e_g[1:]+e_g[:-1])/2., h_gam, c='k')
ax2.plot(gamma, ll, c='navy', dashes=[4,1])
print "gamma", conf_interval(gamma, ll)
ax2.set_xlim(ext[0],ext[1])
ax3.plot(h_sig, (e_s[1:]+e_s[:-1])/2., c='k')
ax3.plot(ll_s8, sig8, c='firebrick', dashes=[3,1,1,1])
print "sig8", conf_interval(sig8, ll_s8)
ax3.set_ylim(ext[2],ext[3])
plt.setp(ax2.get_xticklabels(), visible=False)
plt.setp(ax2.get_yticklabels(), visible=False)
plt.setp(ax3.get_xticklabels(), visible=False)
ax2.xaxis.tick_top()
ax3.yaxis.tick_right()
ax1.set_ylabel("$\sigma_8$")
ax1.set_xlabel("$\gamma$")
ax1.text(0.55, 0.7, "0.55", rotation=90, va='bottom', ha='right', color='firebrick')
ax1.text(1.5,0.82, "0.82", va='bottom', ha='right', color='navy')
ax1.yaxis.set_minor_locator(ticker.MultipleLocator(0.01))
ax3.yaxis.set_minor_locator(ticker.MultipleLocator(0.01))
ax1.xaxis.set_minor_locator(ticker.MultipleLocator(0.2))
ax2.xaxis.set_minor_locator(ticker.MultipleLocator(0.2))
plt.savefig("../figs/fig9.pdf")
# -
|
analysis/fig9-likelihood.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="lGfOJRsTWKAU"
# # How to Build a Transformer for Supervised Classification Using Keras and TensorFlow
#
# [](https://colab.research.google.com/github/scaleapi/dev-blog/blob/main/exchange-transformer/transformer_example.ipynb)
#
# https://exchange.scale.com/home/blogs/transformers-what-they-are-and-why-they-matter
#
# **Subtitle**: Here's how to implement a transformer in Python for supervised document classification
#
# **Summary**: You can apply a transformer to solve supervised classification problems. Here are some foundational concepts to get you started building your own transformer model
#
# **Byline**: <NAME>
#
# In [Transformers: What They Are and Why They Matter](https://exchange.scale.com/home/blogs/transformers-what-they-are-and-why-they-matter), I discussed the theory and the mathematical details behind how transformers work. This time I'm going to show you how to build a simple transformer model for supervised classification tasks in Python using the APIs and objects from the Keras and TensorFlow libraries. We'll work with the famous [20 newsgroup text dataset](https://scikit-learn.org/0.19/datasets/twenty_newsgroups.html), which consists of around 18,000 newsgroup posts classified into 20 topics.
#
# Your goal will be to learn which newsgroup belongs to which category/topic.
#
# My goal is to show you how to build your own transformer model for classification, so we won't perform any sophisticated or complex preprocessing or fine tuning of the model to beat the performance of other methods on the same task. It is also important to note that the output shown below will not match the output that you get at your end. Due to the stochastic nature of the algorithms involved, the code output will be different with every run of the program.
#
# # A Conceptual Diagram of the Transformer
#
# The figure below shows a conceptual diagram of the transformer we are about to build. This is a simplified version of the transformer model discussed in the seminal paper [Attention is All You Need](https://arxiv.org/abs/1706.03762). The paper describes a model based on self attention for sequence-to-sequence learning. In this article, we'll use the same model for supervised classification. While the input to the model is a sequence of words, the output is not a sequence, as it represents a class of the document. Hence, this transformer model consists of only an encoder layer, followed by fully connected feedforward layers for classification.
#
# 
#
# *Figure 1: The overall transformer model (left). The encoder layer’s details are shown in the dashed pink box (right). Source: Mehreen Saeed*
#
# # The Import Section
# The first step is to import the following libraries into your code.
# + id="VkzmoCetWKAY"
# Different layers
from tensorflow.keras.layers import MultiHeadAttention, Input, Dense
from tensorflow.keras.layers import LayerNormalization, Layer
from tensorflow.keras.layers import TextVectorization, Embedding, GlobalAveragePooling1D
# For miscellaneous functions
from tensorflow.data import Dataset
from tensorflow import convert_to_tensor, string, float32, shape, range, reshape
from tensorflow.keras import utils
# Keras models
from tensorflow.keras import Model, Sequential
# For datasets
from sklearn.datasets import fetch_20newsgroups
# For evaluation
from sklearn import metrics
from sklearn.metrics import confusion_matrix, ConfusionMatrixDisplay
# For math/arrays
import numpy as np
# For plotting
import matplotlib.pyplot as plt
# + [markdown] id="tn6KWFXiWKAa"
# # Load the Dataset
# The scikit-learn library includes the 20-newsgroup dataset with an option to load the training or test data. The code below reads both train and test subsets using `fetch_20newsgroups`, converts the labels to categorical labels, and prints some statistics.
# + id="9Qnz_3jHWKAa" outputId="193fbe31-1916-4186-f0cc-65e04ffe8114"
# Load the training dataset while removing headers, footers and quotes
train_dataset = fetch_20newsgroups(subset='train', random_state=0,
remove=("headers", "footers", "quotes"))
train_X, train_Y = (train_dataset.data, train_dataset.target)
# Test dataset
test_dataset = fetch_20newsgroups(subset='test', random_state=0,
remove=("headers", "footers", "quotes"))
test_X, test_Y = (test_dataset.data, test_dataset.target)
# Target classes
newsgroup_names = train_dataset.target_names
# Total classes
n_classes = len(train_dataset.target_names)
# Convert to binary vectors to represent categories
train_Y_categorical = utils.to_categorical(train_Y)
test_Y_categorical = utils.to_categorical(test_Y)
#Print statistics
print("Total training sequences: ", len(train_X))
print("Total test sequences: ", len(test_X))
print("Target categories are: ", newsgroup_names)
# + [markdown] id="gZVsR9HbWKAc"
# From the above output, we can see that the 20 categories overlap. For example, `talk.politics.mideast` and `talk.politics.misc` have many newsgroups in common. Strictly speaking, this is more of a multi-labeling task, where one newsgroup can be assigned to multiple classes. However, we'll perform multi-class classification, where one example newsgroup is assigned to only one category.
#
# An example newsgroup/data point is shown below. You can change the value of `example_index` to see more newsgroup texts and their categories.
# + id="5Lf9MdmSWKAd" outputId="dac7186f-9e92-4319-eb86-d234564cb0ea"
example_index = 10
print('Category: ', newsgroup_names[train_Y[example_index]])
print('Corresponding text: ', train_X[example_index])
# + [markdown] id="8TwGC6tbWKAe"
# # Text Vectorization: Converting Words to Numbers
#
# The training data we just read is a series of English sentences. We need to convert the textual data to numbers so that it can be input to the transformer model. One possible solution is to use the `TextVectorization` layer from the Keras library. This layer can be trained to learn a vocabulary consisting of all the unique words in a corpus using the `adapt` method. You can then use the trained vectorized layer to replace each word in a sentence with its corresponding dictionary index.
#
# Here is a small toy example to help you understand the working of this layer. We have two simple sentences that are converted to fixed-size numeric vectors of length 8. The learned dictionary and the corresponding vectorized sentences are printed at the end of the code. To help distinguish between different variables in this article, all the variable names related to the toy examples are preceded by `toy_`.
# + id="wKnZYUDKWKAe" outputId="858b95a0-8e37-41f3-e310-c33e41e7ad1c"
toy_sentences = [["I am happy today"], ["today weather is awesome"]]
# Create the TextVectorization layer
toy_vectorize_layer = TextVectorization(
output_sequence_length=8,
max_tokens=15)
# Learn a dictionary
toy_vectorize_layer.adapt(Dataset.from_tensor_slices(toy_sentences))
# Use the trained TextVectorization to replace each word by its
# dictionary index
toy_vectorized_words = toy_vectorize_layer(convert_to_tensor(toy_sentences, dtype=string))
print("Dictionary: ", toy_vectorize_layer.get_vocabulary())
print("Vectorized words: ", toy_vectorized_words)
# + [markdown] id="K0dRznJ4WKAf"
# ## Vectorize the Training and Test Data
#
# Let's apply text vectorization to our training and test samples from the newsgroup dataset.
# + id="_e9_tM9PWKAf"
# The total distinct words to use
vocab_size = 25000
# Specify the maximum charancters to consider in each newsgroup
sequence_length = 300
train_X_tensor = Dataset.from_tensor_slices(train_X)
# TextVectorization layer
vectorize_layer = TextVectorization(
output_sequence_length=sequence_length,
max_tokens=vocab_size)
# Adapt method trains the TextVectorization layer and
# creates a dictionary
vectorize_layer.adapt(train_X_tensor)
# Convert all newsgroups in train_X to vectorized tensors
train_X_tensors = convert_to_tensor(train_X, dtype=string)
train_X_vectorized = vectorize_layer(train_X_tensors)
# Convert all newsgroups in test_X to vectorized tensors
test_X_tensors = convert_to_tensor(test_X, dtype=string)
test_X_vectorized = vectorize_layer(test_X_tensors)
# + [markdown] id="YkVivw9zWKAg"
# # The Embedding Layer: Positional Encoding of Words and Indices
#
# In all natural language processing tasks, the order of words within text is important. Changing the position of words in a sentence can change its entire meaning. The transformer model does not use any convolutions or recurrences, and hence, the positional information of words has to be explicitly added to the input data before being processed by the transformer. The authors of [Attention is All You Need](https://arxiv.org/abs/1706.03762) recommend using a sum of word embeddings and positional encodings as the input to the encoder. They also proposed their own scheme that uses sinusoids for mapping words and positions to the embedded vectors.
#
# To keep things simple, instead of implementing embedding using sinusoids, we'll use Keras' `Embedding` layer that initializes all embeddings to random numbers and later learns the embedding during the training phase. Here is an example on the smaller toy sentences to help you understand what the embedding layer does.
# + id="anyc4eMKWKAg"
# Embedding for words
toy_word_embedding_layer = Embedding(input_dim=15, output_dim=4)
toy_embedded_words = toy_word_embedding_layer(toy_vectorized_words)
# Embedding for positions
toy_position_embedding_layer = Embedding(input_dim=8, output_dim=4)
toy_positions = range(start=0, limit=8, delta=1)
toy_embedded_positions = toy_position_embedding_layer(toy_positions)
# + [markdown] id="SKFfNl5rWKAg"
# The `toy_embedded_words` are the word embeddings corresponding to each sentence. Each word in a sentence is represented by a vector. As each vectorized toy sentence has a maximum length of eight, there are eight corresponding positions in a sentence. The `embedded_toy_positions` are the corresponding embeddings for the word positions. They are the same for all sentences. The two embeddings are summed together to form the output of the text preprocessing stage. Both word and position embeddings for this toy example are shown in the figure below.
#
# 
#
# *Figure 2: The word and position embeddings. The position embeddings are added to the word embeddings of each sentence. Source: Mehreen Saeed.*
#
# + [markdown] id="ne3n2RTVWKAh"
# ## Writing a Customized Embedding Layer
#
# While the Keras' `Embedding` layer implements the basic functionality for initializing and learning the encoding, there is no layer that adds the word and position embeddings to produce the final output. We have to write our own custom layer to implement this step. Below is our own `EmbeddingLayer` that has two `Embedding` layers:
# - `word_embedding`: Maps words to their corresponding encodings
# - `position_embedding`: Maps positions/word indices to their corresponding encoding
#
# The `call()` method uses the Keras' `Embedding` layer to compute the words, position embeddings, and return their sum.
# + id="adh3HMhPWKAh"
class EmbeddingLayer(Layer):
def __init__(self, sequence_length, vocab_size, embed_dim):
super(EmbeddingLayer, self).__init__()
self.word_embedding = Embedding(input_dim=vocab_size, output_dim=embed_dim)
self.position_embedding = Embedding(input_dim=sequence_length, output_dim=embed_dim)
def call(self, tokens):
sequence_length = shape(tokens)[-1]
all_positions = range(start=0, limit=sequence_length, delta=1)
positions_encoding = self.position_embedding(all_positions)
words_encoding = self.word_embedding(tokens)
return positions_encoding + words_encoding
# + [markdown] id="d9ETWebGWKAh"
# # The Encoder Layer
#
# There is no direct implementation of the transformer encoder layer in Keras. However, we can write our own custom layer by making use of the [`MultiHeadAttention` layer](https://keras.io/api/layers/attention_layers/multi_head_attention/) and the [`Dense` layer](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense).
#
# ### The Multihead Attention Layer
# In my tutorial on [transformers](https://exchange.scale.com/home/blogs/transformers-what-they-are-and-why-they-matter), I explained the multihead layer in detail. The computations taking place in this layer appear in the figure below.
#
# 
#
# *Figure 3: The multihead attention layer. Source: Mehreen Saeed.*
#
#
# Fortunately, the Keras library includes an implementation of `MultiHeadAttention` layer. We'll use this layer to build the encoder layer.
#
# If you are interested in the nitty gritty details of the Keras\' `MultiHeadAttention` layer, here is a toy example to help you understand. The `MultiHeadAttention` attention layer encapsulates a list of eight weights, which are:
# - Index 0: Weights for linear projection of query
# - Index 1: Bias corresponding to linear projection of query
# - Index 2: Weights for linear projection of key
# - Index 3: Bias corresponding to linear projection of key
# - Index 4: Weights for linear projection of values
# - Index 5: Bias corresponding to linear projection of values
# - Index 6: Weights for linear projection of concatenated heads
# - Index 7: Bias corresponding to linear projection of concatenated heads
# + id="Xjt0qCpNWKAi" outputId="8fd44903-d1a2-482e-c376-ea868fa6128f"
toy_multihead = MultiHeadAttention(num_heads=1, key_dim=3)
toy_x = np.array([[[1, 2, 3]]])
toy_x_tensor = convert_to_tensor(toy_x, dtype=float32)
toy_attn_output, toy_attn_wts = toy_multihead(toy_x_tensor, toy_x_tensor, return_attention_scores=True)
print('Multihead layer output: \n', toy_attn_output)
print('\nMultihead attention wts: \n', toy_attn_wts)
print('\nTotal Layer weights: ', len(toy_multihead.get_weights()))
# + [markdown] id="ZHWrvfA2WKAi"
# ## Implementing the Encoder Layer
# The code implementing the custom `EncoderLayer` is shown below. The `call()` method implements the basic operations that take place in the encoder layer.
# + id="5vzTWfGqWKAi"
class EncoderLayer(Layer):
def __init__(self, total_heads, total_dense_units, embed_dim):
super(EncoderLayer, self).__init__()
# Multihead attention layer
self.multihead = MultiHeadAttention(num_heads=total_heads, key_dim=embed_dim)
# Feed forward network layer
self.nnw = Sequential(
[Dense(total_dense_units, activation="relu"),
Dense(embed_dim)])
# Normalization
self.normalize_layer = LayerNormalization()
def call(self, inputs):
attn_output = self.multihead(inputs, inputs)
normalize_attn = self.normalize_layer(inputs + attn_output)
nnw_output = self.nnw(normalize_attn)
final_output = self.normalize_layer(normalize_attn + nnw_output)
return final_output
# + [markdown] id="qjjMXhqrWKAj"
# To help you understand the computations taking place in the `call()` method, the figure below shows the encoder layer along with the corresponding code that implements it.
#
# 
#
# *Figure 4: The conceptual diagram of the encoder layer (left). The corresponding code (right).*
#
# + [markdown] id="Wu8ExO4wWKAj"
# # Powering Up the Transformer
#
# It's time to construct our final transformer model from the `EmbeddingLayer` and `EncoderLayer`. We also need to add the `GlobalAveragePooling1D` layer followed by a `Dense` layer. The final output of the transformer is produced by a softmax layer, where each unit of the layer corresponds to a category of the text documents.
#
# The following code constructs a transformer model for supervised classification and prints its summary.
# + id="1H5Wk08rWKAj" outputId="7073387e-aa5e-491a-b392-fcabd03367b7"
embed_dim = 64
num_heads = 2
total_dense_units = 60
# Our two custom layers
embedding_layer = EmbeddingLayer(sequence_length, vocab_size, embed_dim)
encoder_layer = EncoderLayer(num_heads, total_dense_units, embed_dim)
# Start connecting the layers together
inputs = Input(shape=(sequence_length, ))
emb = embedding_layer(inputs)
enc = encoder_layer(emb)
pool = GlobalAveragePooling1D()(enc)
d = Dense(total_dense_units, activation="relu")(pool)
outputs = Dense(n_classes, activation="softmax")(d)
# Construct the transformer model
transformer_model = Model(inputs=inputs, outputs=outputs)
transformer_model.compile(optimizer="adam", loss="categorical_crossentropy", metrics=['accuracy', 'Precision', 'Recall'])
transformer_model.summary()
# + [markdown] id="L6OGCEWWWKAj"
# ## Training the Transformer
# The code below trains the transformer model using a 33% split for the validation set.
# + id="iE-44B75WKAk" outputId="d0173705-f67b-438e-b4a3-863e8f49e688"
history = transformer_model.fit(
train_X_vectorized, train_Y_categorical, batch_size=32, epochs=4, validation_split=0.33)
# + [markdown] id="Xu58M-tRWKAk"
# The `history` object stores the learning history of our model. We have the following keys in the `history.history` object returned by the `fit()` method.
# + id="OX7JbirgWKAk" outputId="5066bc3f-50b0-41d3-ef72-5a249536b46c"
print(history.history.keys())
# + [markdown] id="cuOT7UzPWKAk"
# The keys preceded by `val_` indicate the metric corresponding to the validation set. The four metrics that are being recorded are:
#
# - _Loss_: The [categorical cross entropy](https://keras.io/api/losses/probabilistic_losses/#categoricalcrossentropy-class) loss function
# - _Accuracy_: Percentage of correctly classified examples
# - *Precision*: Total true positives divided by the total instances labeled as positive. For multi-class classification, precision is computed for all classes and then averaged.
# - *Recall*: Total true positives divided by the total number of examples that belong to the positive class. For multi-class classification, recall is calculated for all classes and then averaged.
#
# Let's visualize the learning history by plotting various metrics.
# + colab={"base_uri": "https://localhost:8080/", "height": 256} id="PF3U8ZfjWKAl" outputId="b20eeda0-55de-4e81-d5d3-b43b4a60aa93"
fig = plt.figure(figsize=(18,4))
metric = ['loss', 'accuracy', 'precision', 'recall']
validation_metric = ['val_loss', 'val_accuracy', 'val_precision', 'val_recall']
for i,j,k in zip(metric, validation_metric, np.arange(len(metric))):
fig.add_subplot(141+k)
plt.plot(history.history[i])
plt.plot(history.history[j])
plt.legend(['Training', 'Validation'])
plt.title('Training and validation ' + i)
plt.xlabel('Epoch Number')
plt.ylabel('Accuracy')
plt.show()
# + [markdown] id="N7IaqkGPWKAl"
# # Viewing the Learned Word and Position Embeddings
# Keras' `Embedding` layer starts with random encodings and learns them during the training phase, and you can visualize these embeddings by rendering them as an image.
#
# The code below creates two models, i.e., a `random_embeddings_model` that uses an initialized untrained embedding layer and a `learned_embeddings_model` that is built from the learned embedding layer of our transformer model. Once the models are created, the first training example is passed through both models and its output rendered as an image. A comparison of both renderings show how the embeddings start with completely random values, which take on a more smooth and ordered form after training. This learned embeddings layer, therefore, assigns each newsgroup example a unique signature that distinguishes it from from other newsgroups.
# + id="2dBpVt2LWKAl" outputId="0bf0f545-5854-4e07-e433-82f4f7682935"
# Get the random embeddings
random_embedding_layer = EmbeddingLayer(sequence_length, vocab_size, embed_dim)
random_emb = random_embedding_layer(inputs)
random_embeddings_model = Model(inputs=inputs, outputs=random_emb)
random_embedding = random_embeddings_model.predict(train_X_vectorized[0:1,])
random_matrix = reshape(random_embedding[0, :, :], (sequence_length, embed_dim))
# Get the learned embeddings
learned_embeddings_model = Model(inputs=inputs, outputs=emb)
learned_embedding = learned_embeddings_model.predict(train_X_vectorized[0:1,])
learned_matrix = reshape(learned_embedding[0, :, :], (sequence_length, embed_dim))
# Render random embeddings
fig = plt.figure(figsize=(15, 8))
ax = plt.subplot(1, 2, 1)
cax = ax.matshow(random_matrix)
plt.gcf().colorbar(cax)
plt.title('Random embeddings', y=1)
# Render learned embeddings
ax = plt.subplot(1, 2, 2)
cax = ax.matshow(learned_matrix)
plt.gcf().colorbar(cax)
plt.title('Learned embeddings', y=1)
plt.show()
# + [markdown] id="gjPbd2y-WKAl"
# # Evaluating the Transformer's Classification Performance
# We can evaluate our trained model on the test set using different methods, discussed below.
#
# ## The Accuracy, Precision, and Recall Metrics
# The accuracy, precision, and recall metrics were all recorded during training. The `evaluate()` method of the `transformer_model` object returns these metrics in an array. Let's print them for both the training and test sets.
# + id="OL9oHeeZWKAl" outputId="be72bde5-d444-4aba-f197-f04ab952a2b6"
train_metrics = transformer_model.evaluate(train_X_vectorized, train_Y_categorical)
test_metrics = transformer_model.evaluate(test_X_vectorized, test_Y_categorical)
print('Training set evaluation - Accuracy:\n', train_metrics[1],
' Precision: ', train_metrics[2], ' Recall: ', train_metrics[3])
print('Test set evaluation - Accuracy:\n', test_metrics[1],
' Precision: ', test_metrics[2], ' Recall: ', test_metrics[3], '\n')
# + [markdown] id="9M4GnkPQWKAm"
# ## The Confusion Matrix
# The code below computes the confusion matrix and displays it using the functions provided in the scikit-learn library.
# + id="f35vgsHnWKAm" outputId="6a23c46f-7f40-4871-e07a-a323c2393f0f"
# For confusion matrix
test_predict = transformer_model.predict(test_X_vectorized)
test_predict_labels = np.argmax(test_predict, axis=1)
fig, ax = plt.subplots(figsize=(15, 15))
# Create and display the confusion matrix
test_confusion_matrix = confusion_matrix(test_Y, test_predict_labels)
cm = ConfusionMatrixDisplay(confusion_matrix=test_confusion_matrix,
display_labels=newsgroup_names)
cm.plot(xticks_rotation="vertical", ax=ax)
plt.title('Confusion Matrix of the Test Set')
plt.show()
print('Correct classification: ', np.sum(np.diagonal(test_confusion_matrix)), '/', len(test_predict_labels))
# + [markdown] id="fYkBoeHmWKAm"
# At a first glance, the various scores on the test set don't look very good. However, we can understand the results by looking at the confusion matrix and observing where the errors are taking place. Because many categories/topics in the newsgroup dataset are similar and can contain overlapping content, a newsgroup can belong to multiple classes at the same time. We can see from the confusion matrix that a majority of errors have occurred in similar classes. For example, when the true label is `comp.windows.x`, many examples are classified into `comp.graphics` or `comp.os.ms-windows.misc`. This is also the case for `sci.religion.christian` and `talk.religion.misc`.
# + [markdown] id="vkR9sow4WKAm"
# # Next Steps
#
# Now you know how to develop a simple transformer model for solving supervised classification problems. The encoder layer is the basic ingredient required for classification tasks, and you can now easily add a decoder layer to this model and experiment with sequence to sequence tasks such as language translation, sentence paraphrasing, and even document summarization.
#
# You can also add more encoder layers to the transformer model we just built and experiment with different values of sequence lengths, embedding dimensions, and number of dense units in the feed forward layer. The concept of [self attention](https://exchange.scale.com/public/blogs/attention-models-what-they-are-and-why-they-matter) , along with its implementation in transformers, creates a powerful model that is already being used to solve many real-world problems. Transformers are likely to [make their way into more and more machine learning tasks in the near future](https://exchange.scale.com/public/blogs/state-of-ai-report-2021-transformers-taking-ai-world-by-storm-nathan-benaich).
# + [markdown] id="rUaPsAtyaxcT"
# # Learn More
#
#
#
# * [How to Build a Fully Connected Feedforward Neural Network Using Keras and TensorFlow](https://exchange.scale.com/home/blogs/how-to-build-a-fully-connected-feedforward-neural-network-using-keras-and-tensorflow)
#
# * [How to Build an Autoencoder Using Keras and TensorFlow](https://exchange.scale.com/home/blogs/how-to-build-an-autoencoder-using-keras-tensorflow)
#
# * [Data Labeling: What It Is and Why It Matters](https://exchange.scale.com/public/blogs/data-labeling-what-it-is-and-why-it-matters)
#
#
# * [State of AI Report: Transformers Are Taking the AI World by Storm](https://exchange.scale.com/public/blogs/state-of-ai-report-2021-transformers-taking-ai-world-by-storm-nathan-benaich)
#
# * [Attention Models: What They Are and Why They Matter](https://exchange.scale.com/public/blogs/attention-models-what-they-are-and-why-they-matter)
#
# * [How to Use TensorFlow.js to Create JavaScript-Based ML](
# https://exchange.scale.com/public/blogs/how-to-build-next-generation-web-based-ml-apps-using-tensorflowjs-with-google)
#
#
#
|
exchange-transformer/transformer_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/pcsilcan/dm/blob/master/20202/dm_20202_0402_spam_detector.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="pv_mSKg-FP0D" colab_type="code" colab={}
from scipy.io import arff
import pandas as pd
import numpy as np
import tensorflow as tf
from tensorflow import keras
# + id="SEl9jQAPFY8D" colab_type="code" colab={}
repo='https://raw.githubusercontent.com/pcsilcan/dm/master/20202/data/'
filename = 'spambase.arff'
url = '%s%s'%(repo, filename)
# + id="6w7kO02-FqkX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 67} outputId="a86974d9-2109-4f74-e3ab-9d8be727b7e8"
# !curl -O $url
# + id="RmFZj_XvFvG2" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 215} outputId="94b9db83-542d-438d-c8f0-a534903e8491"
data = arff.loadarff(filename)
df = pd.DataFrame(data[0])
df['class'] = df['class'].apply(int)
df.head()
# + id="PV1e9FyRF4Rs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 151} outputId="ec0b7057-c186-4d17-c1a0-542eec2ca5af"
p = [ 1, 2, 3, 4, 2, 3, 1, 0 ]
hp = np.zeros((len(p), 5))
hp[np.arange(len(p)), p] = 1
hp
# + id="W7B9000FJrgK" colab_type="code" colab={}
X_train = df.iloc[:3664, :-1].to_numpy()
temp = df.iloc[:3664, -1].to_numpy()
Y_train = np.zeros((len(X_train), 2))
Y_train[np.arange(len(X_train)), temp] = 1
X_validation = df.iloc[3664:4580, :-1].to_numpy()
temp = df.iloc[3664:4580, -1].to_numpy()
Y_validation = np.zeros((len(X_validation), 2))
Y_validation[np.arange(len(X_validation)), temp] = 1
X_test = df.iloc[4580:, :-1].to_numpy()
temp = df.iloc[4580:, -1].to_numpy()
Y_test = np.zeros((len(X_test), 2))
Y_test[np.arange(len(X_test)), temp] = 1
# + id="IVnKJocvRPNN" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 202} outputId="81b4012c-44ab-4e5b-9699-fe0c448d74c8"
print(X_train.shape, Y_train.shape)
Y_train[:10]
# + id="pVUY-n9EJ7T4" colab_type="code" colab={}
model = keras.Sequential()
model.add(keras.layers.Dense(16, input_dim=57, activation='relu'))
model.add(keras.layers.Dense(8, activation='relu'))
model.add(keras.layers.Dense(4, activation='relu'))
model.add(keras.layers.Dense(2, activation='softmax'))
# + id="BCnXtyERP1cB" colab_type="code" colab={}
model.compile(loss='binary_crossentropy', optimizer='adam', metrics=['accuracy'])
# + id="Bji7n2d4Qcmc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 706} outputId="88768fa7-fe0f-432f-8859-2532715ab002"
model.fit(X_train, Y_train, epochs=20, batch_size=10)
# + id="DfKkl7cmQsHX" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="219e2e38-bea6-4533-f1ad-0cbe94119cd7"
model.evaluate(X_validation, Y_validation)
# + id="aA67UlfzTdNj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 50} outputId="ecf5f000-690a-485b-860d-9e9249e3b266"
model.evaluate(X_test, Y_test)
# + id="oJauzqiIU1wj" colab_type="code" colab={}
|
20202/dm_20202_0402_spam_detector.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
# +
def linear(a,b,x):
return a*x + b
lin = lambda x: linear(1,3,x)
def sigmoid(f):
return lambda x: 1/(1+np.exp(f(-x)))
sigmoid_lin = sigmoid(lin)
# -
x=np.linspace(-10,10,10)
y=np.linspace(-10,10,100)
plt.plot(x,sigmoid_lin(x),'r', label='linspace(-10,10,10)')
plt.plot(y,sigmoid_lin(y),'b', label='linspace(-10,10,100)')
plt.grid()
plt.xlabel('X Axis')
plt.ylabel('Y Axis')
plt.title('Sigmoid Function')
plt.suptitle('Sigmoid')
|
basics/loss_function.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: python3.7.4
# language: python
# name: python3.7.4
# ---
# # MNISTでセグメンテーションに挑戦
#
#
# +
import os
import shutil
import numpy as np
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
import torch
import torchvision
import torchvision.transforms as transforms
import binarybrain as bb
# -
# ## 初期設定
# +
# configuration
net_name = 'MnistSegmentation'
data_path = os.path.join('./data/', net_name)
rtl_sim_path = '../../verilog/mnist'
rtl_module_name = 'MnistSegmentation'
output_velilog_file = os.path.join(data_path, rtl_module_name + '.v')
sim_velilog_file = os.path.join(rtl_sim_path, rtl_module_name + '.v')
bin_mode = True
frame_modulation_size = 1
epochs = 32
mini_batch_size = 32
rows = 3
cols = 3
depth = 28
margen = 0
# -
# ## データセット準備
#
# データセットを自作する
# +
# dataset
dataset_path = './data/'
dataset_train = torchvision.datasets.MNIST(root=dataset_path, train=True, transform=transforms.ToTensor(), download=True)
dataset_test = torchvision.datasets.MNIST(root=dataset_path, train=False, transform=transforms.ToTensor(), download=True)
# 面積の比率で重みを作っておく
areas = np.zeros((11))
for img, label in dataset_train:
img = img.numpy()
areas[label] += np.mean(img)
areas[10] += np.mean(1.0-img)
areas /= len(dataset_train)
wight = 1 / areas
wight /= np.max(wight)
def make_teacher_image(gen, rows, cols, margin=0):
source_img = np.zeros((1, rows*28, cols*28), dtype=np.float32)
teaching_img = np.zeros((11, rows*28, cols*28), dtype=np.float32)
for row in range(rows):
for col in range(cols):
x = col*28
y = row*28
img, label = gen.__next__()
source_img[0,y:y+28,x:x+28] = img
teaching_img[label,y:y+28,x:x+28] = img
teaching_img[10,y:y+28,x:x+28] = 1.0-img
msk1 = teaching_img > 0.5
msk0 = teaching_img <= 0.5
teaching_img[msk1] = 1.0
teaching_img[msk0] = 0.0
# for i in range(11):
# teaching_img[i] *= wight[i]
teaching_img[10] *= 0.1
if margin > 0:
return source_img, teaching_img[:,margin:-margin,margin:-margin]
else:
return source_img, teaching_img
def transform_data(dataset, n, rows, cols, margin):
def data_gen():
l = len(dataset)
i = 0
while True:
yield dataset[i%l]
i +=1
gen = data_gen()
source_imgs = []
teaching_imgs = []
for _ in range(n):
x, t = make_teacher_image(gen, rows, cols, margin)
source_imgs.append(x)
teaching_imgs.append(t)
return source_imgs, teaching_imgs
class MyDatasets(torch.utils.data.Dataset):
def __init__(self, source_imgs, teaching_imgs, transforms=None):
self.transforms = transforms
self.source_imgs = source_imgs
self.teaching_imgs = teaching_imgs
def __len__(self):
return len(self.source_imgs)
def __getitem__(self, index):
source_img = self.source_imgs[index]
teaching_img = self.teaching_imgs[index]
if self.transforms:
source_img, teaching_img = self.transforms(source_img, teaching_img)
return source_img, teaching_img
source_imgs_train, teaching_imgs_train = transform_data(dataset_train, mini_batch_size*10, rows, cols, margen)
my_dataset_train = MyDatasets(source_imgs_train, teaching_imgs_train)
source_imgs_test, teaching_imgs_test = transform_data(dataset_test, mini_batch_size*5, rows, cols, margen)
my_dataset_test = MyDatasets(source_imgs_test, teaching_imgs_test)
loader_train = torch.utils.data.DataLoader(dataset=my_dataset_train, batch_size=mini_batch_size, shuffle=True)
loader_test = torch.utils.data.DataLoader(dataset=my_dataset_test, batch_size=mini_batch_size, shuffle=False)
# -
i = 0
for source_imgs, teaching_imgs in loader_test:
print(source_imgs[0].shape)
print(teaching_imgs[0].shape)
plt.figure(figsize=(12,6))
plt.subplot(1,12,1)
plt.imshow(source_imgs[0][0], 'gray')
for j in range(11):
plt.subplot(1,12,2+j)
plt.imshow(teaching_imgs[0][j], 'gray')
plt.show()
i += 1
if i > 5:
break
# ## ネットワーク構築
import random
class BinaryConvLayer(bb.Sequential):
def __init__(self, hidden_ch, output_ch, *, padding='same', bin_dtype=bb.DType.FP32, input_shape=None, name=None):
self.layers = [
# input(pointwise)
bb.Convolution2d(
bb.Sequential([
bb.DifferentiableLut([hidden_ch*6, 1, 1], bin_dtype=bin_dtype),
bb.DifferentiableLut([hidden_ch, 1, 1], connection='serial', bin_dtype=bin_dtype),
]),
filter_size=(1, 1),
fw_dtype=bin_dtype),
# hidden(depthwise)
bb.Convolution2d(
bb.Sequential([
bb.DifferentiableLut([hidden_ch, 1, 1], connection='depthwise', bin_dtype=bin_dtype),
]),
filter_size=(3, 3), padding=padding,
fw_dtype=bin_dtype),
# output(pointwise)
bb.Convolution2d(
bb.Sequential([
bb.DifferentiableLut([output_ch*6, 1, 1], connection='serial', bin_dtype=bin_dtype),
bb.DifferentiableLut([output_ch, 1, 1], connection='serial', bin_dtype=bin_dtype),
]),
filter_size=(1, 1),
fw_dtype=bin_dtype),
]
super(BinaryConvLayer, self).__init__(self.layers, input_shape=input_shape, name=name)
def forward(self, x_buf, train=True):
self.bypass = random.random() > 0.5
if not train and self.bypass:
return x_buf
return super(BinaryConvLayer, self).forward(x_buf, train=train)
def backward(self, dy_buf):
if self.bypass:
return dy_buf
return super(BinaryConvLayer, self).backward(dy_buf)
# 重み付きの損失関数を自作する
class MyLoss(bb.LossFunction):
def __init__(self, weight):
super(MyLoss, self).__init__(self)
self.weight = weight
self.clear()
def clear(self):
self.loss = 0.
self.n = 0
def get(self):
if self.n == 0:
return 0
return self.loss / self.n
def calculate(self, y_buf, t_buf):
grad = y_buf.numpy() - t_buf.numpy();
grad *= self.weight
err = grad * grad
self.loss += np.mean(err) * y_buf.get_frame_size()
self.n += y_buf.get_frame_size()
return bb.FrameBuffer.from_numpy(grad)
# +
# バイナリ時は BIT型を使えばメモリ削減可能
bin_dtype = bb.DType.BIT if bin_mode else bb.DType.FP32
def make_conv_layer(hidden_ch, output_ch, padding='same', bin_dtype=bb.DType.BIT):
return bb.Sequential([
# input(pointwise)
bb.Convolution2d(
bb.Sequential([
bb.DifferentiableLut([hidden_ch*6, 1, 1], bin_dtype=bin_dtype),
bb.DifferentiableLut([hidden_ch, 1, 1], connection='serial', bin_dtype=bin_dtype),
]),
filter_size=(1, 1),
fw_dtype=bin_dtype),
# hidden(depthwise)
bb.Convolution2d(
bb.Sequential([
bb.DifferentiableLut([hidden_ch, 1, 1], connection='depthwise', bin_dtype=bin_dtype),
]),
filter_size=(3, 3), padding=padding,
fw_dtype=bin_dtype),
# output(pointwise)
bb.Convolution2d(
bb.Sequential([
bb.DifferentiableLut([output_ch*6, 1, 1], connection='serial', bin_dtype=bin_dtype),
bb.DifferentiableLut([output_ch, 1, 1], connection='serial', bin_dtype=bin_dtype),
]),
filter_size=(1, 1),
fw_dtype=bin_dtype),
])
sub_net = bb.Sequential()
sub_net.append(make_conv_layer(72, 36, bin_dtype=bin_dtype))
for i in range(depth-2):
sub_net.append(BinaryConvLayer(72, 36, bin_dtype=bin_dtype))
sub_net.append(
bb.Convolution2d(
bb.Sequential([
bb.DifferentiableLut([512], connection='serial', batch_norm=True, bin_dtype=bin_dtype),
bb.DifferentiableLut([11*6*6], connection='random', batch_norm=False, bin_dtype=bin_dtype),
bb.DifferentiableLut([11*6], connection='serial', batch_norm=False, bin_dtype=bin_dtype),
bb.DifferentiableLut([11], connection='serial', batch_norm=False, bin_dtype=bin_dtype),
]),
padding='same',
filter_size=(3, 3),
fw_dtype=bin_dtype))
# define network
net = bb.Sequential([
bb.RealToBinary(frame_modulation_size=frame_modulation_size, bin_dtype=bin_dtype),
sub_net,
bb.BinaryToReal(frame_integration_size=frame_modulation_size, bin_dtype=bin_dtype)
])
net.set_input_shape([1, rows*28, cols*28])
if bin_mode:
net.send_command("binary true")
print(net.get_info(2))
# -
# ## 学習実施
#
# 学習を行います
# +
# bb.load_networks(data_path, net)
# learning
loss = bb.LossSoftmaxCrossEntropy() # MyLoss(loss_weight) # bb.LossMeanSquaredError()
metrics = bb.MetricsCategoricalAccuracy()
optimizer = bb.OptimizerAdam()
optimizer.set_variables(net.get_parameters(), net.get_gradients())
for epoch in range(epochs):
# View
i = 0
for x_imgs, t_imgs in loader_test:
x_buf = bb.FrameBuffer.from_numpy(np.array(x_imgs).astype(np.float32))
t_buf = bb.FrameBuffer.from_numpy(np.array(t_imgs).astype(np.float32))
y_buf = net.forward(x_buf, train=False)
result_imgs = y_buf.numpy()
plt.figure(figsize=(16,8))
plt.subplot(1,12,1)
plt.imshow(x_imgs[0][0], 'gray')
for j in range(11):
plt.subplot(1,12,2+j)
plt.imshow(result_imgs[0][j], 'gray')
plt.show()
i += 1
if i > 5:
break
# learning
loss.clear()
metrics.clear()
with tqdm(loader_train) as tqdm_loadr:
for x_imgs, t_imgs in tqdm_loadr:
x_buf = bb.FrameBuffer.from_numpy(np.array(x_imgs).astype(np.float32))
t_buf = bb.FrameBuffer.from_numpy(np.array(t_imgs).astype(np.float32))
y_buf = net.forward(x_buf, train=True)
dy_buf = loss.calculate(y_buf, t_buf)
metrics.calculate(y_buf, t_buf)
net.backward(dy_buf)
optimizer.update()
tqdm_loadr.set_postfix(loss=loss.get(), acc=metrics.get())
# test
loss.clear()
metrics.clear()
for x_imgs, t_imgs in loader_test:
x_buf = bb.FrameBuffer.from_numpy(np.array(x_imgs).astype(np.float32))
t_buf = bb.FrameBuffer.from_numpy(np.array(t_imgs).astype(np.float32))
y_buf = net.forward(x_buf, train=False)
loss.calculate(y_buf, t_buf)
metrics.calculate(y_buf, t_buf)
bb.save_networks(data_path, net)
print('epoch[%d] : loss=%f acc=%f' % (epoch, loss.get(), metrics.get()))
# -
plt.figure(figsize=(16,8))
for source_imgs, teaching_imgs in loader_test:
x_buf = bb.FrameBuffer.from_numpy(np.array(x_imgs).astype(np.float32))
t_buf = bb.FrameBuffer.from_numpy(np.array(t_imgs).astype(np.float32))
y_buf = net.forward(x_buf, train=False)
result_imgs = y_buf.numpy()
plt.subplot(1,12,1)
plt.imshow(source_imgs[0][0], 'gray')
for i in range(11):
plt.subplot(1,12,2+i)
plt.imshow(result_imgs[0][i], 'gray')
break
# ## RTL(Verilog)変換
#
# FPGA化するために Verilog に変換します。インターフェースはXilinx社のAXI4 Stream Video 仕様(フレームスタートでtuserが立つ)となります。
# MaxPooling の単位で画像サイズが縮小されてしまうので、現状、この単位でしか変換できないため3つに分けて出力しています。
# +
# export verilog
with open(output_velilog_file, 'w') as f:
f.write('`timescale 1ns / 1ps\n\n')
bb.dump_verilog_lut_cnv_layers(f, rtl_module_name, sub_net)
# Simulation用ファイルに上書きコピー
shutil.copyfile(output_velilog_file, sim_velilog_file)
|
tests/python/mnist/MnistSegmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from urllib.request import urlopen
from bs4 import BeautifulSoup
#url = "https://www.grubhub.com/restaurant/cast-iron-waffles-9604-longstone-ln-charlotte/1444707"
#html = urlopen(url)
import codecs
htmlfile = './Cast Iron Waffles Delivery - 9604 Longstone Ln Charlotte _ Order Online With Grubhub (12_09_2020 03_06_15).html'
file = codecs.open(htmlfile, "r", "utf-8")
soup = BeautifulSoup(file.read(),'lxml')
# -
section= soup.find_all('section',{'class':'cb-expansion-panel'})
data = []
for main in section:
header = main.find('div', {'class': 'menuSection-headerTitle u-flex u-flex-justify-xs--between'}).getText()
item_div = main.find_all('div',{'class':'s-card-wrapper'})
for menu in item_div:
div = menu.find('div',{'class':'u-inset-squished-3'})
name = div.find('a',{'class':'menuItem-name'}).getText()
descp = div.find('p',{'class':'u-text-secondary'}).getText()
p = soup.find('span', {'itemprop':'price'}).getText()
data.append([header,name,descp,p])
data[20]
import pandas as pd
df = pd.DataFrame(data,columns=['categories','menu', 'description', 'price'])
df
|
scripts/Site Scrapping.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
x = np.arange(10)
y = np.random.randint(10,20,10)
x
y
plt.plot(x,y,color = 'red',label = "firstplot")
plt.plot(y,x,color = 'blue', label = "second plot")
plt.xlabel("array x value")
plt.ylabel("array y value")
plt.title("scatter plot")
plt.legend(loc = "lower left")
plt.show()
# +
plt.subplot(2,2,1)
plt.plot(x,y,color = 'red',label = "firstplot")
plt.xlabel("array x1 value")
plt.ylabel("array y1 value")
plt.title("scatter plot")
plt.legend(loc = "lower left")
plt.subplot(2,2,2)
plt.scatter(x,y,color = 'red',label = "firstplot")
plt.xlabel("array x1 value")
plt.ylabel("array y1 value")
plt.title("scatter plot")
plt.legend(loc = "lower left")
plt.subplot(2,2,3)
plt.scatter(x,y,color = 'red',label = "firstplot")
plt.xlabel("array x1 value")
plt.ylabel("array y1 value")
plt.title("scatter plot")
plt.legend(loc = "lower left")
plt.subplot(2,2,4)
plt.scatter(y,x, marker ="*",color = 'blue', label = "second plot")
plt.xlabel("array x2 value")
plt.ylabel("array y2value")
plt.title("scatter plot")
plt.legend(loc = "lower left")
plt.show()
# -
a = np.random.randint(30,40,10)
b = np.random.randint(20,30,10)
plt.subplot(2,2,1)
plt.bar(a,b)
plt.xlabel("a")
plt.ylabel("y")
plt.title("bar graph")
plt.subplot(2,2,2)
plt.plot(a,b)
plt.xlabel("a")
plt.ylabel("y")
plt.title("line graph")
plt.subplot(2,2,3)
plt.scatter(a,b)
plt.xlabel("a")
plt.ylabel("y")
plt.title("scatter graph")
languages = ["c","c++","java","python","php"]
students = [23,17,35,29,12]
plt.pie(students,labels = languages,autopct="%3f%%")
plt.show()
fig = plt.figure()
ax = fig.add_axes([0,0,1,1])
ax.pie(students,labels = languages,autopct="%1.2f%%")
plt.show()
import seaborn as sns
iris = sns.load_dataset("tips")
iris
iris.isnull().sum()
plt.subplot(2,2,1)
plt.scatter(iris['sepal_length'],iris["sepal_width"],marker = "*",color = "pink")
plt.subplot(2,2,2)
plt.scatter(iris['sepal_length'],iris["petal_length"],marker = "+",color = "Yellow")
plt.show()
sns.pairplot(iris,hue='species',palette = "coolwarm")
iris
x = [1,2,3,4,5]
y = [1,2,3,4,5]
plt.bar(x,y,color = "red")
plt.show()
sns.heatmap(iris.corr(),annot = True)
sns.countplot(x="species", data =iris)
iris["species"].value_counts()
sns.boxplot(x = "species",y = "sepal_length",data = iris,palette = "rainbow")
sns.violinplot(x = "species",y = "sepal_length",data = iris,palette = "rainbow")
# + active=""
# importing the libraries
# reading the dataset
# checking the null values
# replacing the null values
# converting categorical data into numerical data
# split the data in too input variables and output variable
# converting numerially converted data to binary format
# splitting the data in to test and trian
# scaling the data-optional
#
# -
import pandas as pd
import numpy as np
dataset = pd.read_csv(r"D:\pythongriet\Churn_Modelling.csv")
dataset.head(10)
dataset.isnull().any()
dataset.isnull().sum()
dataset.fillna(dataset['CreditScore'].mean(),inplace = True)
dataset.isnull().any()
# + active=""
# convesion will take place according to alphabets
# France - 0
# Germany 1
# Spain - 4
# india - 2
# japan - 3
# -
from sklearn.preprocessing import LabelEncoder
le = LabelEncoder()
dataset['Geography'] = le.fit_transform(dataset['Geography'])
dataset['Gender'] = le.fit_transform(dataset['Gender'])
dataset.head(10)
x = dataset.iloc[:,3:13]
y = dataset.iloc[:,13]
type(y)
x = dataset.iloc[:,3:13].values
y = dataset.iloc[:,13].values
type(x)
x
y
from sklearn.preprocessing import OneHotEncoder
one = OneHotEncoder(categorical_features = [1])
x = one.fit_transform(x).toarray()
x
from sklearn.preprocessing import OneHotEncoder
one = OneHotEncoder()
z = one.fit_transform(x[:,1:2]).toayyay()
x = np.delete(x,1,axis = 1)
x = np.concatenate((z,x),axis = 1)
|
matplotlib and seaborn lib.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from use_translator import *
query = {
"message": {
"query_graph": {
"edges": {
"e00": {
"subject": "n00",
"object": "n01",
"predicates": [
"biolink:related_to"
]
#,"provided_by": {"allowlist": []}
}
},
"nodes": {
"n00": {
"categories": [
"biolink:SmallMolecule"
],
"ids": [
"CHEBI:3380"
]
},
"n01": {
"categories": [
"biolink:Disease"
],
"ids": [
"MONDO:0005301"
]
}
}
}
}
}
sr = strider(query)
kps = requests.get('https://kp-registry.renci.org/kps').json().keys()
print('x')
for kp in kps:
print(kp)
query['message']['query_graph']['edges']['e00']['provided_by']['allowlist'] = [kp]
#printjson(query)
s = strider(query)
print('----')
ar['message']['query_graph']
sr
|
minihackathons/WF C.3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # The Viterbi Algorithm for Hidden Markov Models
#
# This is a notebook written by <NAME> ([@luisguiserrano](https://github.com/luisguiserrano)) of Udacity.
#
# This is a simple implementation of the Viterbi Algorithm for training HMMs.
#
# This notebook is a supplement for this video.
# https://www.youtube.com/watch?v=kqSzLo9fenk
#
# In here, we use this notation:
# - `s` stands for Sunny
# - `r` stands for Rainy
# - `h` stands for Happy
# - `g` stands for Grumpy
#
# And the goal is, given a sequence of moods, we find the most likely sequence of types of weathers that caused that sequence of moods.
# + deletable=true editable=true
from numpy import random
# + deletable=true editable=true
# Transition Probabilities
p_ss = 0.8
p_sr = 0.2
p_rs = 0.4
p_rr = 0.6
# Initial Probabilities
p_s = 2/3
p_r = 1/3
# Emission Probabilities
p_sh = 0.8
p_sg = 0.2
p_rh = 0.4
p_rg = 0.6
moods = ['H', 'H', 'G', 'G', 'G', 'H']
probabilities = []
weather = []
if moods[0] == 'H':
probabilities.append((p_s*p_sh, p_r*p_rh))
else:
probabilities.append((p_s*p_sg, p_r*p_rg))
for i in range(1,len(moods)):
yesterday_sunny, yesterday_rainy = probabilities[-1]
if moods[i] == 'H':
today_sunny = max(yesterday_sunny*p_ss*p_sh, yesterday_rainy*p_rs*p_sh)
today_rainy = max(yesterday_sunny*p_sr*p_rh, yesterday_rainy*p_rr*p_rh)
probabilities.append((today_sunny, today_rainy))
else:
today_sunny = max(yesterday_sunny*p_ss*p_sg, yesterday_rainy*p_rs*p_sg)
today_rainy = max(yesterday_sunny*p_sr*p_rg, yesterday_rainy*p_rr*p_rg)
probabilities.append((today_sunny, today_rainy))
for p in probabilities:
if p[0] > p[1]:
weather.append('S')
else:
weather.append('R')
weather
# + deletable=true editable=true
probabilities
|
hmm/simple-hmm.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from astropy.time import Time
pd.plotting.register_matplotlib_converters()
# +
#user-defined parameters
quantile_thresh = 0.995 # the percentile to consider [0.995]
time_res='1H' #the time resoltuion at which to perfrom the analysis ['1H']
n_categories = 4 # number of categories for V, B [4]. 4 = quartiles, etc
n_cl_bins = 100 # number of cost/loss bins for plotting [100]
# +
#Load and process data
######################
def load_omni():
columns = ['date', 'time', 'hgi_lat', 'hgi_lon', 'br', 'bt', 'bn', 'b', 'v', 'v_lat', 'v_lon', 'density', 'temperature']
omni = pd.read_csv('OMNI_COHO1HR_MERGED_MAG_PLASMA_199207.txt', delim_whitespace=True, skiprows=240, skipfooter=3, names=columns, engine='python')
# Combine date and time into timeindex
omni['time'] = pd.to_datetime(omni['date']+'T'+omni['time'], format='%d-%m-%YT%H:%M:%S.%f')
omni.drop(columns='date', inplace=True)
omni.set_index('time', inplace=True)
# Set invalid numbers to NaN
id_bad = omni == -1e31
omni[id_bad] = np.NaN
return omni
def load_icme():
"""
Function to load in the Richardson and Cane ICME list.
"""
columns = ['shock', 'leading_edge', 'trailing_edge', 'type']
icme = pd.read_csv('ICMEs.dat', delim_whitespace=True, names=columns)
# Convert MJD to datetimes.
icme['shock'] = pd.to_datetime(Time(icme['shock'], format='mjd').datetime)
icme['leading_edge'] = pd.to_datetime(Time(icme['leading_edge'], format='mjd').datetime)
icme['trailing_edge'] = pd.to_datetime(Time(icme['trailing_edge'], format='mjd').datetime)
return icme
def cumdf(data, bin_edges):
hist, bins = np.histogram(data, bins=bin_edges)
cdf = np.cumsum(hist) / np.sum(hist)
return cdf
omni = load_omni()
icme = load_icme()
#resample the omni data at the required resolution
omni=omni.resample(time_res).mean()
# Remove rows from omni with any bad speed, density or imf magnitude data (all needed for G calculaiton)
#omni.dropna(how='any', subset=['density', 'v', 'b'], inplace=True)
# compute geoeffectivness parameter
alpha=0.5;
# IMF clock angle
theta=np.arctan2(-omni['bt'],omni['bn']);
# Equation 1 in Owens et al. 2017
d_exp = 2.0/3.0 - alpha
b_exp = 2.0*alpha
v_exp = 7.0/3.0 - 2.0*alpha
omni['g'] = (omni['density']**d_exp) * (omni['b']**b_exp) * (omni['v']**v_exp) * (np.sin(theta/2.0)**4.0) * 1e-6
g_thresh = np.nanquantile(omni['g'], quantile_thresh)
print("{:3.1f}th percentile of G: {:3.4f}".format(quantile_thresh*100.0,g_thresh))
plt.plot(omni['g'], 'k-', zorder=0)
plt.hlines(g_thresh, omni.index.min(), omni.index.max(), colors='r', linestyles='--', zorder=1)
plt.xlim(omni.index.min(), omni.index.max())
plt.ylim(0, omni['g'].max())
plt.xlabel('Time')
plt.ylabel('Geoeffectiveness')
# +
# Compute the quantiles of the CME speed distribution
n_bins = 10000 # number of bins for CDF [10000]
# Define G bins for computing CDFs
g_min = omni['g'].min()
g_max = omni['g'].max()
dg = (g_max-g_min)/n_bins
g_bin_edges = np.arange(g_min, g_max+dg, dg)
g_bin_centres = (g_bin_edges[0:-1] + g_bin_edges[1:]) / 2.0
# Loop through the ICMEs and compute the average CME properties, and also mask the solar wind time series.
# Add in keys to icme and omni for the average solar wind properties and cme properties.
for key in ['v', 'b']:
icme[key] = np.NaN*np.zeros(icme.shape[0])
for key in ['cme_v', 'cme_b', 'region', 'type']:
if key not in ['region', 'type']:
omni[key] = np.NaN*np.zeros(omni.shape[0])
else:
omni[key] = np.zeros(omni.shape[0])
for i, row in icme.iterrows():
# Find solar wind period between cme shock and trailing edge
id_cme = (omni.index >= row['shock']) & (omni.index <= row['trailing_edge'])
if np.any(id_cme):
# Update ICME with solar wind parameters
icme.loc[i, 'v'] = omni.loc[id_cme, 'v'].mean(skipna=True)
icme.loc[i, 'b'] = omni.loc[id_cme, 'b'].mean(skipna=True)
# Update solar wind paramters with average CME properties and type
omni.loc[id_cme, 'cme_v'] = icme.loc[i, 'v']
omni.loc[id_cme, 'cme_b'] = icme.loc[i, 'b']
omni.loc[id_cme, 'region'] = 1 # Flag for being in sheath, will update CME regions after.
omni.loc[id_cme, 'type'] = icme.loc[i, 'type']
# Update region flag if in CME rather than sheath
id_cme = (omni.index >= row['leading_edge']) & (omni.index <= row['trailing_edge'])
if np.any(id_cme):
omni.loc[id_cme, 'region'] = 2
# +
#Compute the G values for each CME quantile
quantiles = np.arange(1, n_categories, 1) / n_categories
v_quantiles = icme['v'].quantile(quantiles)
b_quantiles = icme['b'].quantile(quantiles)
print('V quantile boundaries are: ',v_quantiles)
print('B quantile boundaries are: ',b_quantiles)
# Find the indices of SW parameters for the different v and b quantiles and sw classifications (cme, no cme)
groups = {}
groups['all'] = np.nonzero(omni['g'].to_numpy())[0]
groups['no_cme'] = np.nonzero(omni['region'].to_numpy() == 0)[0]
groups['cme'] = np.nonzero(omni['region'].to_numpy() > 0)[0]
cme_v = omni['cme_v'].to_numpy()
cme_v_id = np.nonzero(np.isfinite(cme_v))[0]
cme_v = cme_v[cme_v_id]
cme_b = omni['cme_b'].to_numpy()
cme_b_id = np.nonzero(np.isfinite(cme_b))[0]
cme_b = cme_b[cme_b_id]
for i in range(v_quantiles.size + 1):
v_key = "v_{:02d}".format(i)
b_key = "b_{:02d}".format(i)
if i == 0:
id_group = cme_v <= v_quantiles.values[i] # do nans need to be exlucded here?
which_v = np.nonzero(id_group)[0]
groups[v_key] = cme_v_id[which_v]
id_group = cme_b <= b_quantiles.values[i]
which_b = np.nonzero(id_group)[0]
groups[b_key] = cme_b_id[which_b]
elif (i > 0) & (i < v_quantiles.size):
id_group = (cme_v > v_quantiles.values[i-1]) & (cme_v <= v_quantiles.values[i])
which_v = np.nonzero(id_group)[0]
groups[v_key] = cme_v_id[which_v]
id_group = (cme_b > b_quantiles.values[i-1]) & (cme_b <= b_quantiles.values[i])
which_b = np.nonzero(id_group)[0]
groups[b_key] = cme_b_id[which_b]
elif i == v_quantiles.size:
id_group = cme_v > v_quantiles.values[i-1]
which_v = np.nonzero(id_group)[0]
groups[v_key] = cme_v_id[which_v]
id_group = cme_b > b_quantiles.values[i-1]
which_b = np.nonzero(id_group)[0]
groups[b_key] = cme_b_id[which_b]
# Now the combined V and B groups
for i in range(v_quantiles.size + 1):
v_key = "v_{:02d}".format(i)
for j in range(b_quantiles.size + 1):
b_key = "b_{:02d}".format(j)
vb_key = v_key + '_' + b_key
# Also get the intersection of the matched quantiles for the combined v-b category
groups[vb_key] = np.intersect1d(groups[v_key], groups[b_key])
# -
cme_v[np.isfinite(cme_v)]
# +
# Compute the exceedance probability and numbers above and below threshold for each grouping of the data.
prob = {}
number = {}
# Find g_bin closest to threshold from below.
pos = np.nonzero(g_bin_centres <= g_thresh)[0] # should this be <=?
id_exceed = pos[-1]
for key, index in groups.items():
g_sub = omni.iloc[index]['g']
cdf = cumdf(g_sub, g_bin_edges)
prob[key] = 1.0 - cdf[id_exceed]
n_above = np.sum(g_sub > g_thresh)
n_below = np.sum(g_sub <= g_thresh)
n_all = np.sum(np.isfinite(g_sub))
number[key] = {'above': n_above, 'below': n_below, 'all': n_all}
# -
# Ouput key statistics
print("Ncme = {}, Ncme' = {}".format(number['cme']['all'], number['cme']['above']))
print("Nsw = {}, Nsw' = {}".format(number['no_cme']['all'], number['no_cme']['above']))
print("p'sw = {:3.4f}".format(number['no_cme']['above']/number['no_cme']['all']))
print("p'cme = {:3.4f}".format(number['cme']['above']/number['cme']['all']))
# +
#Perform cost/loss analysis
costs = np.geomspace(0.001, 1, num=n_cl_bins)
# DataFrame to store all cost calculations
costloss = pd.DataFrame({'cost':costs, 'perfect': 0, 'climatology': 0, 'cmes': 0, 'v': 0, 'b': 0, 'vb': 0})
loss = 1.0
for i, cost in enumerate(costs):
# cost of a perfect deterministic forecast is the number of times the
# threshold is exceeded, multiplied by the cost
costloss.loc[i, 'perfect'] = cost * number['all']['above']
# cost of climatology is different if the climatological probability is
# above or below the c/l ratio
if (cost <= prob['all']): # always take action
# cost is the clratio at all times
costloss.loc[i, 'climatology'] = cost * number['all']['all']
elif (cost > prob['all']): #never take action
# cost is simply all the missed events
costloss.loc[i, 'climatology'] = loss * number['all']['above']
##########################
# Update the costs due to no_cme solar wind
if cost <= prob['no_cme']:
costloss.loc[i, 'cmes'] = cost * number['no_cme']['all']
costloss.loc[i, 'v'] = cost * number['no_cme']['all']
costloss.loc[i, 'b'] = cost * number['no_cme']['all']
costloss.loc[i, 'vb'] = cost * number['no_cme']['all']
elif cost > prob['no_cme']:
costloss.loc[i, 'cmes'] = loss * number['no_cme']['above']
costloss.loc[i, 'v'] = loss * number['no_cme']['above']
costloss.loc[i, 'b'] = loss * number['no_cme']['above']
costloss.loc[i, 'vb'] = loss * number['no_cme']['above']
##########################
# Update the costs due to icme arrivals
if cost <= prob['cme']:
costloss.loc[i, 'cmes'] += cost * number['cme']['all']
elif cost > prob['cme']:
costloss.loc[i, 'cmes'] += loss * number['cme']['above']
##########################
# Update the costs due to v & b categories
for j in range(v_quantiles.size + 1):
v_key = "v_{:02d}".format(j)
b_key = "b_{:02d}".format(j)
if cost <= prob[v_key]:
costloss.loc[i, 'v'] += cost * number[v_key]['all']
elif cost > prob[v_key]:
costloss.loc[i, 'v'] += loss * number[v_key]['above']
if cost <= prob[b_key]:
costloss.loc[i, 'b'] += cost * number[b_key]['all']
elif cost > prob[v_key]:
costloss.loc[i, 'b'] += loss * number[b_key]['above']
##########################
# Update the costs due to combined v & b categories
for j in range(v_quantiles.size + 1):
v_key = "v_{:02d}".format(j)
for k in range(b_quantiles.size + 1):
b_key = "b_{:02d}".format(k)
vb_key = v_key + '_' + b_key
if cost <= prob[vb_key]:
costloss.loc[i, 'vb'] += cost * number[vb_key]['all']
elif cost > prob[vb_key]:
costloss.loc[i, 'vb'] += loss * number[vb_key]['above']
# Compute the relative forecast value (clim - forecast)/(clim - perfect)
for key in ['cmes', 'v', 'b', 'vb']:
new_key = key+"_relative"
costloss[new_key] = 100 * (costloss['climatology'] - costloss[key]) / (costloss['climatology'] - costloss['perfect'])
# +
#plot the cost-loss summary
fig, ax = plt.subplots(figsize=(8,6))
ax.fill_between(costloss['cost'], 0, costloss['cmes_relative'], color=[0.7, 0.7, 0.7], label='ICME Arrival time only')
ax.plot(costloss['cost'], costloss['v_relative'], 'b-', label='+ ICME V')
ax.plot(costloss['cost'], costloss['b_relative'], 'r-', label='+ ICME B')
ax.plot(costloss['cost'], costloss['vb_relative'], 'k-', label='+ ICME V & B')
ax.set_xscale('log')
#ax.set_ylim(0, 70)
ax.set_xticks((0.001,0.01,0.1,1))
ax.set_xticklabels(('0.001','0.01','0.1','1'))
ax.set_xlim(0.001, 1.0)
#legend
handles, labels = plt.gca().get_legend_handles_labels()
legendorder = [3,0,1,2]
plt.legend([handles[idx] for idx in legendorder],[labels[idx] for idx in legendorder])
ax.grid(b=True, which='both', axis='both')
ax.set_ylabel('Forecast value (%)')
ax.set_xlabel('C/L: Relative cost of taking mitigating action')
ax.annotate('', xy=(0.75, 1.1), xycoords='axes fraction', xytext=(1, 1.1),
arrowprops=dict(arrowstyle="<-", color='k'))
ax.annotate('False alarms\ncannot be tolerated', xy=(0.8, 1.1), xycoords='axes fraction', xytext=(0.52, 1.04),ha='left' )
ax.annotate('', xy=(0.225, 1.1), xycoords='axes fraction', xytext=(0, 1.1),
arrowprops=dict(arrowstyle="<-", color='k'))
ax.annotate('False alarms\ncan be tolerated', xy=(0.8, 1.1), xycoords='axes fraction', xytext=(0.45, 1.04),ha='right' )
|
CostLoss.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture-05-2 Fresnel Equation
# * author : <NAME>
# * Data : 2019/12/13
# %matplotlib inline
import matplotlib.pyplot as plt
import numpy as np
def SnellLaw(theta_i, ni, nt=1.0):
# theta_i : incident angle
# ni : the refractive index of the incident material
# nt : the refractive index of the transmissive material
theta_i = np.array( theta_i, dtype=np.complex_ )
# theta_t : refractive angle
theta_t = np.arcsin( (ni/nt) * np.sin(theta_i) )
return theta_t
def cal_r_tau_TE(theta_i, ni, nt=1.0):
# theta_i : incident angle
# ni : the refractive index of the incident material
# nt : the refractive index of the transmissive material
# theta_t : refractive angle
theta_t = SnellLaw( theta_i, ni, nt)
# r_TE : reflection coefficients defined on the electric field
# tau_TE : transmission coefficients defined on the electric field
r_TE = ( ni*np.cos(theta_i)-nt*np.cos(theta_t) ) / ( ni*np.cos(theta_i)+nt*np.cos(theta_t) )
tau_TE = ( 2*ni*np.cos(theta_i) ) / ( ni*np.cos(theta_i)+nt*np.cos(theta_t) )
return r_TE, tau_TE
def cal_r_tau_TM(theta_i, ni, nt=1.0):
# theta_i : incident angle
# ni : the refractive index of the incident material
# nt : the refractive index of the transmissive material
# theta_t : refractive angle
theta_t = SnellLaw( theta_i, ni, nt)
# r_TM : reflection coefficients defined on the electric field
# tau_TM : transmission coefficients defined on the electric field
r_TM = ( nt*np.cos(theta_i)-ni*np.cos(theta_t) ) / ( ni*np.cos(theta_t)+nt*np.cos(theta_i) )
tau_TM = ( 2*ni*np.cos(theta_i) ) / ( ni*np.cos(theta_t)+nt*np.cos(theta_i) )
return r_TM, tau_TM
def CriticalAngle( ni, nt):
# ni : the refractive index of the incident material
# nt : the refractive index of the transmissive material
return np.arcsin( nt/ni ) if nt<=ni else np.pi/2
def BrewsterAngle(ni, nt=1.0):
# ni : the refractive index of the incident material
# nt : the refractive index of the transmissive material
theta_B = np.arctan( nt/ni )
return theta_B
def cal_kz(k, kx, ky):
# n : refractive index
# ko : wave number in vacuum
# kx : wave number along x-axis
# ky : wave number along y-axis
kz = np.sqrt( k**2 - kx**2 - ky**2, dtype=np.complex_ )
return kz
# ### Now we are going to show you how to prove the Fresnel equation by rigorous electromagnetic wave equations.
# ### what we have learned in Lecture 1 (Plane Wave)
def wave_equation(A0, x, y, z, t=0.0, w=1.0, kx=1.0, ky=1.0, kz=1.0):
# A0 : the complex amplitude of the wave
# x, y, z : positions
# t : time
# w : angular frequency
# kx, ky, kz : wave number components
return A0 * np.exp( 1j * ( kx*x + ky*y + kz*z - w*t) )
def cal_k(kx, ky, kz):
# kx, ky, kz : wave number components
return np.sqrt(kx**2 + ky**2 + kz**2)
def cal_normalize_direction(kx, ky, kz):
# kx, ky, kz : wave number components
k = cal_k(kx, ky, kz)
return kx/k, ky/k, kz/k
# ### what we have learned in Lecture 3 (Boundary Conditions)
def cal_epsilon_r_from_n(n=1.0):
# n : refractive index
return n**2
def cal_D_from_E(E, epsilon=1.0):
# E : electric field
# epsilon : permittivity (=epsilon0 * epsilonr)
return E * epsilon
def cal_Dz2(Dz1, sigma_s = 0.0):
# Dz1 : z component of the displacement vector field in material 1
# sigma_s : surface charge density
return Dz1 + sigma_s
def cal_Bz2(Bz1):
# Bz1 : z component of the magnetic field in material 1
return Bz1
def cal_B_from_H(H, mu=1.0):
# H : magnetic intensity field
# mu : permeability (= mu0 * mur)
return H * mu
def cal_Ex2(Ex1):
# Ex1 : x component of the electric field in material 1
return Ex1
def cal_Ey2(Ey1):
# Ey1 : y component of the electric field in material 1
return Ey1
def cal_Hx2(Hx1, Jsy = 0.0):
# Hx1 : x component of the magnetic intensity field in material 1
# Jsy : y component of the surface current density
return Hx1 + Jsy
def cal_Hy2(Hy1, Jsx = 0.0):
# Hy1 : y component of the magnetic intensity field in material 1
# Jsx : x component of the surface current density
return Hy1 - Jsx
# ### what we have learned in Lecture 4 (TE mode and TM mode)
# TE set
def cal_Hx_from_Ey(Ey, kz, w, mu0 = 1.0, mur=1.0):
# Ey : y component of the electic field
# kz : z component of the wave vector
# w : angular frequency
# mu0 : permeability in vacuum
# mur : relative permeability
return -(kz/w/mu0/mur) * Ey
def cal_Hz_from_Ey(Ey, kx, w, mu0 = 1.0, mur=1.0):
# Ey : t component of the electric field
# kx : x component of the wave vector
# w : angular frequency
# mu0 : permeability in vacuum
# mur : relative permeability
return (kx/w/mu0/mur) * Ey
# TM set
def cal_Ex_from_Hy(Hy, kz, w, e0=1.0, er=1.0):
# Hy : y component of the magnetic intensity field
# kz : z component of the wave vector
# w : angular frequency
# e0 : permittivity in vacuum
# er : relative permittivity
return (kz/w/e0/er)*Hy
def cal_Ez_from_Hy(Hy, kx, w, e0=1.0, er=1.0):
# Hy : y component of the magnetic intensity field
# kx : x component of the wave vector
# w : angular frequency
# e0 : permittivity in vacuum
# er : relative permittivity
return -(kx/w/e0/er)*Hy
# ## Incidence, Reflection and Transmission Definitions
#
# From the experiments, as shown in **Fig. 1**, when there is an incident ray in material 1 (on xz plane, $k_y = 0$), there would be one reflected ray also in material 1 and a transmissive ray in material 2. Now we are going to calculate the reflection coeffcients, $\gamma_{TE,TM}$, between the reflected ray and the incident ray and the transmission coefficients, $\tau_{TE,TM}$, between the transmissive ray and the incident ray.
#
# <img src="Lecture-5-Material/Fig-1.jpg" width="700">
# $$Fig. 1. $$
#
# ========================================================================================================================
# #### Supporting 1
# The field toward +z could be represented:
#
# <font size="4">$$F^+(x,y,z) = F_+exp(i(k_xx+k_yy+k_zz)) - (S1)$$</font>
#
# and the field toward -z could be represented:
#
# <font size="4">$$F^-(x,y,z) = F_-exp(i(k_xx+k_yy-k_zz)) - (S2)$$</font>
#
# where $k_z = \sqrt{k^2 - k_x^2 - k_y^2}$, which is a constant and $F_+$ and $F_-$ are also complex constants.
#
# ========================================================================================================================
#
# Suppose the total electric field and total the magnetic field in material 1 are $E_{x,1}, E_{y,1}, E_{z,1}, H_{x,1}, H_{y,1}, H_{z,1}$ and the electric field and the magnetic field in material 2 are $E_{x,2}, E_{y,2}, E_{z,2}, H_{x,2}, H_{y,2}, H_{z,2}$.
#
# ### TE mode incidence (**Fig. 1(a)**)
# Because the incident wave (toward +z) and the reflected wave (toward -z) are both in material 1, then the total field in material 1 would be
#
# <font size="4"> for $z<0$ </font>
# <font size="4">$$E_{y,1}(\textbf{r}) = E_{y,1}^+(\textbf{r}) + E_{y,1}^-(\textbf{r}) - (1)$$</font>
# <font size="4">$$H_{x,1}(\textbf{r}) = H_{x,1}^+(\textbf{r}) + H_{x,1}^-(\textbf{r}) - (2)$$</font>
# <font size="4">$$H_{z,1}(\textbf{r}) = H_{z,1}^+(\textbf{r}) + H_{z,1}^-(\textbf{r}) - (3)$$</font>
#
# Only transmissive wave is in material 2 (toward +z).
#
# <font size="4"> for $z>0$ </font>
# <font size="4">$$E_{y,2}(\textbf{r}) = E_{y,2}^+(\textbf{r}) - (4)$$</font>
# <font size="4">$$H_{x,2}(\textbf{r}) = H_{x,2}^+(\textbf{r}) - (5)$$</font>
# <font size="4">$$H_{z,2}(\textbf{r}) = H_{z,2}^+(\textbf{r}) - (6)$$</font>
#
# Similarly, for TM mode incidence,
#
# ### TM mode incidence (**Fig. 1(b)**)
#
# <font size="4"> for $z<0$ </font>
# <font size="4">$$H_{y,1}(\textbf{r}) = H_{y,1}^+(\textbf{r}) + H_{y,1}^-(\textbf{r}) - (7)$$</font>
# <font size="4">$$E_{x,1}(\textbf{r}) = E_{x,1}^+(\textbf{r}) + E_{x,1}^-(\textbf{r}) - (8)$$</font>
# <font size="4">$$E_{z,1}(\textbf{r}) = E_{z,1}^+(\textbf{r}) + E_{z,1}^-(\textbf{r}) - (9)$$</font>
# <font size="4"> for $z>0$ </font>
# <font size="4">$$H_{y,2}(\textbf{r}) = H_{y,2}^+(\textbf{r}) - (10)$$</font>
# <font size="4">$$E_{x,2}(\textbf{r}) = E_{x,2}^+(\textbf{r}) - (11)$$</font>
# <font size="4">$$E_{z,2}(\textbf{r}) = E_{z,2}^+(\textbf{r}) - (12)$$</font>
#
def cal_total_field(F_plus, F_minus=0.0):
return F_plus + F_minus
# ## Match Boundary Conditions
#
# Applied with the boundary conditions
#
# <font size="4">$$\hat{\textbf{a}}_n\cdot(\textbf{D}_1-\textbf{D}_2) = \rho_S = 0- (13)$$</font>
# <font size="4">$$\hat{\textbf{a}}_n\cdot(\textbf{B}_1-\textbf{B}_2) = 0 - (14)$$</font>
# <font size="4">$$\hat{\textbf{a}}_n\times(\textbf{E}_1-\textbf{E}_2) = \textbf{0} - (15)$$</font>
# <font size="4">$$\hat{\textbf{a}}_n\times(\textbf{H}_1-\textbf{H}_2) = \textbf{J}_S = \textbf{0}- (16)$$</font>
#
# and no charge and current would accumulate on the interface; as a result, the tangential components of $\textbf{E}$ and $\textbf{H}$ should be continuous on the interface and the normal components of $\textbf{D}$ and $\textbf{B}$ should also be continuous.
#
# For similicity, in the following section, we only consider non-magnetic material. (i.e. $\mu_r = 1.0$)
#
# ### TE mode Incidence
# <font size="4">$$E_{y,1}(z=0^-,t) = E_{y,2}(z=0^+,t) - (17)$$</font>
# <font size="4">$$H_{x,1}(z=0^-,t) = H_{x,2}(z=0^+,t) - (18)$$</font>
# <font size="4">$$H_{z,1}(z=0^-,t) = H_{z,2}(z=0^+,t) - (19)$$</font>
#
# then insert **Eq. (1-6)** into **Eq. (17-19)**,
#
# <font size="4">$$E_{y,1}^+(z=0^-,t) + E_{y,1}^-(z=0^-,t) = E_{y,2}^+(z=0^+,t) - (20)$$</font>
# <font size="4">$$H_{x,1}^+(z=0^-,t) + H_{x,1}^-(z=0^-,t) = H_{x,2}^+(z=0^+,t) - (21)$$</font>
# <font size="4">$$H_{z,1}^+(z=0^-,t) + H_{z,1}^-(z=0^-,t) = H_{z,2}^+(z=0^+,t) - (22)$$</font>
#
# Insert **Eq. (S1-S2)** into **Eq. (20-22)**
#
# <font size="4">$$E_{y,1,+}exp(ik_{x,1}x) + E_{y,1,-}exp(ik_{x,1}x) = E_{y,2,+}exp(ik_{x,2}x) - (23)$$</font>
# <font size="4">$$H_{x,1,+}exp(ik_{x,1}x) + H_{x,1,-}exp(ik_{x,1}x) = H_{x,2,+}exp(ik_{x,2}x) - (24)$$</font>
# <font size="4">$$H_{z,1,+}exp(ik_{x,1}x) + H_{z,1,-}exp(ik_{x,1}x) = H_{z,2,+}exp(ik_{x,2}x) - (25)$$</font>
#
# Because **Eq. (23-25)** hold for all x, then it is obvious that $k_{x,1}$ and $k_{x,2}$ should be the same, which is the same as what we have learned in **Lecture 2 (Snell's Law)**. We showed that because the wave front should be continuous on the interface, the tangential components of the wave vector should be the same and thus $k_{x,1} = k_{x,2}$. As a result, the Snell's Law must be followed, indicating that when you calculate the EM wave properties by Maxwell's equations, you do not need to consider Snell's Law because the Snell's Law is already contained in Maxwell's equations.
#
# And in **Lecture 4**, we have derived the relation between $H_x$, $H_z$, and $E_y$,
#
# <font size="4">$$H_x = -\frac{k_z}{\omega\mu_0\mu_r}E_y - (26)$$</font>
# <font size="4">$$H_z = +\frac{k_x}{\omega\mu_0\mu_r}E_y - (27)$$</font>
# <font size="4">$$E_y(\textbf{r},t) = E_{y,0}exp(i (\textbf{k}\cdot\textbf{r}-\omega t)) - (28)$$</font>
#
# **Eq. (23-25)** would become:
#
# <font size="4">$$E_{y,1,+} + E_{y,1,-} = E_{y,2,+} - (29)$$</font>
# <font size="4">$$-\frac{k_{z,1}}{\omega\mu_0}E_{y,1,+} + -\frac{-k_{z,1}}{\omega\mu_0}E_{y,1,-} = -\frac{k_{z,2}}{\omega\mu_0}E_{y,2,+} - (30)$$</font>
# <font size="4">$$+\frac{k_{x,1}}{\omega\mu_0}E_{y,1,+} + \frac{k_{x,1}}{\omega\mu_0}E_{y,1,-} = +\frac{k_{x,2}}{\omega\mu_0}E_{y,2,+} - (31)$$</font>
#
# Interestingly, because $k_{x,1} = k_{x,2}$, then **Eq. (29)** and **Eq.(31)** are the same and only one equation should be considered. (As discussed in **Lecture 1**, there exists a relation between the charge density and the current density.)
# ### Reflection Coefficients and Transmission Coefficients of TE mode
#
# With the definition of the reflection coefficient, $\gamma$, and the transmission coefficient, $\tau$,
#
# <font size="4">$$\gamma_{s,E} = \frac{E_{y,1,-}}{E_{y,1,+}} - (32)$$</font>
# <font size="4">$$\tau_{s,E} = \frac{E_{y,2,+}}{E_{y,1,+}} - (33)$$</font>
#
# Then, **Eq. (29-31)** can be further elaborated as:
#
# <font size="4">$$ 1 + \gamma_{s,E} = \tau_{s,E} - (34)$$</font>
# <font size="4">$$ k_{z,1}( 1 - \gamma_{s,E}) = k_{z,2}\tau_{s,E} - (35)$$</font>
#
# then,
#
# <font size="4">$$ \tau_{s,E} - \gamma_{s,E} = 1 - (36)$$</font>
# <font size="4">$$ k_{z,2}\tau_{s,E} + k_{z,1}\gamma_{s,E} = k_{z,1} - (37)$$</font>
#
# In order to solve the equations, **Eq. (36-37)** can be represented by the matrix representation:
#
# <font size="4">$$\left( \begin{array}{cc} 1 & -1 \\ k_{z,2} & k_{z,1} \end{array} \right)\left( \begin{array}{c} \tau_{s,E} \\ \gamma_{s,E} \end{array} \right) = \left( \begin{array}{c} 1 \\ k_{z,1} \end{array} \right) - (38)$$</font>
#
# Then,
#
# <font size="4">$$\left( \begin{array}{c} \tau_{s,E} \\ \gamma_{s,E} \end{array} \right) = \frac{1}{k_{z,1}+k_{z,2}} \left( \begin{array}{cc} k_{z,1} & 1 \\ -k_{z,2} & 1 \end{array} \right)\left( \begin{array}{c} 1 \\ k_{z,1} \end{array} \right) - (39)$$</font>
#
# so
#
# <font size="4">$$\left( \begin{array}{c} \tau_{s,E} \\ \gamma_{s,E} \end{array} \right) = \left( \begin{array}{c} \frac{2k_{z,1}}{k_{z,1}+k_{z,2}} \\ \frac{k_{z,1}-k_{z,2}}{k_{z,1}+k_{z,2}} \end{array} \right) - (40)$$</font>
#
# Besides, $k_z = nk_ocos\theta$, then **Eq. (40)** would become
#
# <font size="4">$$\left( \begin{array}{c} \tau_{s,E} \\ \gamma_{s,E} \end{array} \right) = \left( \begin{array}{c} \frac{2n_1k_ocos\theta_1}{n_1k_ocos\theta_1+n_2k_ocos\theta_2} \\ \frac{n_1k_ocos\theta_1-n_2k_ocos\theta_2}{n_1k_ocos\theta_1+n_2k_ocos\theta_2} \end{array} \right) - (41)$$</font>
#
# is the same as the equations in [wikipedia (Chinese)](https://zh.wikipedia.org/wiki/%E8%8F%B2%E6%B6%85%E8%80%B3%E6%96%B9%E7%A8%8B) [wikipedia (English)](https://en.wikipedia.org/wiki/Fresnel_equations).
#
# Then up to now, we have solved the reflection coefficients and the transmission coefficient of TE mode.
#
def solve_r_tau_s_E(kz1, kz2):
# kz1 : the z component of the wave vector in material 1
# kz2 : the z component of the wave vector in material 2
# according to Eq. (40)
tau_s_E = 2*kz1 / (kz1 + kz2)
r_s_E = (kz1 - kz2) / (kz1 + kz2)
return tau_s_E, r_s_E
# ### TM mode Incidence
# Follow the same procedure in TE mode incidence.
#
# <font size="4">$$H_{y,1}(z=0^-,t)=H_{y,2}(z=0^+,t) - (42)$$</font>
# <font size="4">$$E_{x,1}(z=0^-,t)=E_{x,2}(z=0^+,t) - (43)$$</font>
# <font size="4">$$E_{z,1}(z=0^-,t)=E_{z,2}(z=0^+,t) - (44)$$</font>
#
# then insert **Eq. (7-12)** into **Eq. (42-44)**,
#
# <font size="4">$$H_{y,1}^+(z=0^-,t) + H_{y,1}^-(z=0^-,t) = H_{y,2}^+(z=0^+,t) - (45)$$</font>
# <font size="4">$$E_{x,1}^+(z=0^-,t) + E_{x,1}^-(z=0^-,t) = E_{x,2}^+(z=0^+,t) - (46)$$</font>
# <font size="4">$$E_{z,1}^+(z=0^-,t) + E_{z,1}^-(z=0^-,t) = E_{z,2}^+(z=0^+,t) - (47)$$</font>
#
# Insert **Eq. (S1-S2)** into **Eq. (45-47)** and $k_{x,1} = k_{x,2}$
#
# <font size="4">$$H_{y,1,+} + H_{y,1,-} = H_{y,2,+} - (48)$$</font>
# <font size="4">$$E_{x,1,+} + E_{x,1,-} = E_{x,2,+} - (49)$$</font>
# <font size="4">$$E_{z,1,+} + E_{z,1,-} = E_{z,2,+} - (50)$$</font>
#
# In **Lecture 4**
#
# <font size="4">$$E_x = + \frac{k_z}{\omega\epsilon_0\epsilon_r}H_y - (51)$$</font>
# <font size="4">$$E_z = - \frac{k_x}{\omega\epsilon_0\epsilon_r}H_y - (52)$$</font>
#
# With **Eq. (51-52)**, **Eq. (48-50)** would become:
#
# <font size="4">$$H_{y,1,+} + H_{y,1,-} = H_{y,2,+} - (53)$$</font>
# <font size="4">$$\frac{k_{z,1}}{\omega\epsilon_0\epsilon_{r,1}}H_{y,1,+} + \frac{-k_{z,1}}{\omega\epsilon_0\epsilon_{r,1}}H_{y,1,-} = \frac{k_{z,2}}{\omega\epsilon_0\epsilon_{r,2}}H_{y,2,+} - (54)$$</font>
#
#
# Define $\gamma_{p,H} = H_{y,1,-}/H_{y,1,+}$ and $\tau_{p,H} = H_{y,2,+}/H_{y,1,+}$
#
# <font size="4">$$ 1 + \gamma_{p,H} = \tau_{p,H} - (54)$$</font>
# <font size="4">$$ \frac{k_{z,1}}{\epsilon_{r,1}}( 1 - \gamma_{p,H}) = \frac{k_{z,2}}{\epsilon_{r,2}}\tau_{p,H} - (55)$$</font>
#
# Then the matrix representation would be:
#
# <font size="4">$$\left( \begin{array}{cc} 1 & -1 \\ \frac{k_{z,2}}{\epsilon_{r,2}} & \frac{k_{z,1}}{\epsilon_{r,1}} \end{array} \right)\left( \begin{array}{c} \tau_{p,H} \\ \gamma_{p,H} \end{array} \right) = \left( \begin{array}{c} 1 \\ \frac{k_{z,1}}{\epsilon_{r,1}} \end{array} \right) - (56)$$</font>
#
# Then,
#
# <font size="4">$$\left( \begin{array}{c} \tau_{p,H} \\ \gamma_{p,H} \end{array} \right) = \frac{1}{\frac{k_{z,1}}{\epsilon_{r,1}}+\frac{k_{z,2}}{\epsilon_{r,2}}} \left( \begin{array}{cc} \frac{k_{z,1}}{\epsilon_{r,1}} & 1 \\ \frac{-k_{z,2}}{\epsilon_{r,2}} & 1 \end{array} \right)\left( \begin{array}{c} 1 \\ \frac{k_{z,1}}{\epsilon_{r,1}} \end{array} \right) - (57)$$</font>
#
# so
#
# <font size="4">$$\left( \begin{array}{c} \tau_{p,H} \\ \gamma_{p,H} \end{array} \right) = \frac{1}{\frac{k_{z,1}}{\epsilon_{r,1}}+\frac{k_{z,2}}{\epsilon_{r,2}}}\left( \begin{array}{c} \frac{2k_{z,1}}{\epsilon_{r,1}} \\ \frac{k_{z,1}}{\epsilon_{r,1}}-\frac{k_{z,2}}{\epsilon_{r,2}} \end{array} \right) = \left( \begin{array}{c} \frac{2\epsilon_{r,2}k_{z,1}}{\epsilon_{r,2}k_{z,1}+\epsilon_{r,1}k_{z,2}} \\ \frac{\epsilon_{r,2}k_{z,1}-\epsilon_{r,1}k_{z,2}}{\epsilon_{r,2}k_{z,1}+\epsilon_{r,1}k_{z,2}} \end{array} \right) - (58)$$</font>
#
# Interestingly the reflection coefficient and the transmission coefficient in **Eq. (58)** is different from the equations in [wikipedia (Chinese)](https://zh.wikipedia.org/wiki/%E8%8F%B2%E6%B6%85%E8%80%B3%E6%96%B9%E7%A8%8B) [wikipedia (English)](https://en.wikipedia.org/wiki/Fresnel_equations) because the reflection coefficient and the transmission coefficient in wiki is defined based on the total electric field of TM mode.
#
# In order to compare the reuslts with wiki, the total electric field of TM mode calculated according to
#
# <font size="4">$$\textbf{k}\times\textbf{H}= -\omega(\epsilon_0\epsilon_r\textbf{E}) - (59)$$</font>
#
# in **Lecture 4**. Then
#
# <font size="4">$$\textbf{E} = \frac{1}{\omega\epsilon_0\epsilon_r} \textbf{k}\times\textbf{H} - (60)$$</font>
#
# Then
#
# <font size="4">$$\left( \begin{array}{c} E_x \\ E_y \\ E_z \end{array} \right) = \frac{1}{\omega\epsilon_0\epsilon_r} \left( \begin{array}{c} k_yH_z-k_zH_y \\ k_zH_x-k_xH_z\\ k_xH_y-k_yH_x \end{array} \right) = \frac{1}{\omega\epsilon_0\epsilon_r} \left( \begin{array}{c} -k_zH_y \\ 0\\ k_xH_y \end{array} \right) - (61)$$</font>
#
# and
#
# <font size="4">$$E = \sqrt{E_x^2+E_y^2+E_z^2} = \frac{k}{\omega\epsilon_0\epsilon_r}H_y = \frac{n\omega\sqrt{\epsilon_0\mu_0}}{\omega\epsilon_0n^2}H_y = \frac{1}{n}\sqrt{\frac{\mu_0}{\epsilon_o}}H_y - (62)$$</font>
#
# As a result,
#
# <font size="4">$$\gamma_{p,E} = \frac{E_{1,-}}{E_{1,+}} = \frac{H_{y,1,-}}{H_{y,1,+}} = \gamma_{p,H} = \frac{\epsilon_{r,2}k_{z,1}-\epsilon_{r,1}k_{z,2}}{\epsilon_{r,2}k_{z,1}+\epsilon_{r,1}k_{z,2}} - (63)$$</font>
# <font size="4">$$\tau_{p,E} = \frac{E_{2,+}}{E_{1,+}} = \frac{n_1H_{y,2,+}}{n_2H_{y,1,+}} = \frac{n_1}{n_2}\tau_{p,H} = \frac{n_1}{n_2} \frac{2\epsilon_{r,2}k_{z,1}}{\epsilon_{r,2}k_{z,1}+\epsilon_{r,1}k_{z,2}} - (64)$$</font>
#
# With $k_z = nk_ocos\theta$, **Eq. (63-64)** is the same as in [wikipedia (Chinese)](https://zh.wikipedia.org/wiki/%E8%8F%B2%E6%B6%85%E8%80%B3%E6%96%B9%E7%A8%8B) [wikipedia (English)](https://en.wikipedia.org/wiki/Fresnel_equations).
def solve_r_tau_p_H(kz1, kz2, n1, n2):
# kz1 : the z component of the wave vector in material 1
# kz2 : the z component of the wave vector in material 2
# n1 : the reflective index of material 1
# n2 : the reflective index of material 2
er1, er2 = n1**2, n2**2
# according to Eq. (58)
tau_p_H = (2*er2*kz1) / (er2*kz1 + er1*kz2)
r_p_H = (er2*kz1 - er1*kz2) / (er2*kz1 + er1*kz2)
return tau_p_H, r_p_H
def solve_r_tau_p_E(kz1, kz2, n1, n2):
# kz1 : the z component of the wave vector in material 1
# kz2 : the z component of the wave vector in material 2
# n1 : the reflective index of material 1
# n2 : the reflective index of material 2
er1, er2 = n1**2, n2**2
# according to Eq. (64)
tau_p_E = (n1/n2) * (2*er2*kz1) / (er2*kz1 + er1*kz2)
r_p_E = (er2*kz1 - er1*kz2) / (er2*kz1 + er1*kz2)
return tau_p_E, r_p_E
# ## Example
# +
n1, n2 = 1.5, 1.0
theta1 = np.linspace( 0, np.pi/2, 100)
wavelength = 520
e0, mu0 = 1.0, 1.0
theta2 = SnellLaw(theta1, n1, n2)
c = 1/np.sqrt(e0*mu0)
ko = 2*np.pi/wavelength
w = c * ko
k1, k2 = n1*ko, n2*ko
kx1 = k1 * np.sin(theta1)
kz1 = k1 * np.cos(theta1)
ky1 = np.zeros( kx1.shape )
kx2 = k2 * np.sin(theta2)
kz2 = k2 * np.cos(theta2)
ky2 = np.zeros( kx2.shape )
# -
# calculation from Fresnel Equations (wiki)
r_TE_FE, tau_TE_FE = cal_r_tau_TE(theta1, n1, n2)
r_TM_FE, tau_TM_FE = cal_r_tau_TM(theta1, n1, n2)
# calculation from the matrix
tau_s_E, r_s_E = solve_r_tau_s_E(kz1, kz2)
tau_p_E, r_p_E = solve_r_tau_p_E(kz1, kz2, n1, n2)
tau_p_H, r_p_H = solve_r_tau_p_H(kz1, kz2, n1, n2)
# +
## plot TE data
fig, axes = plt.subplots(1,2)
plt.rcParams['figure.figsize'] = [15, 10]
axes[0].plot( theta1*180/np.pi, np.real( r_TE_FE ), 'r-', linewidth=1.0)
axes[0].plot( theta1*180/np.pi, np.imag( r_TE_FE ), 'b-', linewidth=1.0)
axes[0].plot( theta1*180/np.pi, np.real( r_s_E ), 'ro', linewidth=2.0)
axes[0].plot( theta1*180/np.pi, np.imag( r_s_E ), 'bo', linewidth=2.0)
axes[0].set_xlabel(r'$\theta_i$ (deg.)', fontsize=20)
axes[0].set_ylabel(r'$\gamma_{TE}$', fontsize=20)
axes[0].legend(['real FE', 'imag. FE', 'real MAT', 'imag. MAT'], fontsize=10)
plt.xlim( 0, 90);
axes[1].plot( theta1*180/np.pi, np.real( tau_TE_FE ), 'r-', linewidth=1.0)
axes[1].plot( theta1*180/np.pi, np.imag( tau_TE_FE ), 'b-', linewidth=1.0)
axes[1].plot( theta1*180/np.pi, np.real( tau_s_E ), 'ro', linewidth=2.0)
axes[1].plot( theta1*180/np.pi, np.imag( tau_s_E ), 'bo', linewidth=2.0)
axes[1].set_xlabel(r'$\theta_i$ (deg.)', fontsize=20)
axes[1].set_ylabel(r'$\tau_{TE}$', fontsize=20)
axes[1].legend(['real FE', 'imag. FE', 'real MAT', 'imag. MAT'], fontsize=10)
plt.xlim( 0, 90);
# +
## plot TE data
fig, axes = plt.subplots(1,2)
plt.rcParams['figure.figsize'] = [15, 10]
axes[0].plot( theta1*180/np.pi, np.real( r_TM_FE ), 'r-', linewidth=1.0)
axes[0].plot( theta1*180/np.pi, np.imag( r_TM_FE ), 'b-', linewidth=1.0)
axes[0].plot( theta1*180/np.pi, np.real( r_p_E ), 'ro', linewidth=2.0)
axes[0].plot( theta1*180/np.pi, np.imag( r_p_E ), 'bo', linewidth=2.0)
axes[0].set_xlabel(r'$\theta_i$ (deg.)', fontsize=20)
axes[0].set_ylabel(r'$\gamma_{TM}$', fontsize=20)
axes[0].legend(['real FE', 'imag. FE', 'real MAT', 'imag. MAT'], fontsize=10)
plt.xlim( 0, 90);
axes[1].plot( theta1*180/np.pi, np.real( tau_TM_FE ), 'r-', linewidth=1.0)
axes[1].plot( theta1*180/np.pi, np.imag( tau_TM_FE ), 'b-', linewidth=1.0)
axes[1].plot( theta1*180/np.pi, np.real( tau_p_E ), 'ro', linewidth=2.0)
axes[1].plot( theta1*180/np.pi, np.imag( tau_p_E ), 'bo', linewidth=2.0)
axes[1].set_xlabel(r'$\theta_i$ (deg.)', fontsize=20)
axes[1].set_ylabel(r'$\tau_{TM}$', fontsize=20)
axes[1].legend(['real FE', 'imag. FE', 'real MAT', 'imag. MAT'], fontsize=10)
plt.xlim( 0, 90);
# -
# The above figures show that the equations in [wikipedia (Chinese)](https://zh.wikipedia.org/wiki/%E8%8F%B2%E6%B6%85%E8%80%B3%E6%96%B9%E7%A8%8B) [wikipedia (English)](https://en.wikipedia.org/wiki/Fresnel_equations) is the same what we have derived. However, in our derivation, we can calculate all the field quantities.
# +
# TE mode
Ey1p = np.zeros( theta1.shape, dtype=np.complex ) # incidence
Hx1p = cal_Hx_from_Ey(Ey1p, kz1, w, mu0 = 1.0, mur=1.0)
Hz1p = cal_Hz_from_Ey(Ey1p, kx1, w, mu0 = 1.0, mur=1.0)
Ey1n = r_s_E * Ey1p
Hx1n = cal_Hx_from_Ey(Ey1n,-kz1, w, mu0 = 1.0, mur=1.0)
Hz1n = cal_Hz_from_Ey(Ey1n, kx1, w, mu0 = 1.0, mur=1.0)
Ey2p = tau_s_E * Ey1p
Hx2p = cal_Hx_from_Ey(Ey2p, kz2, w, mu0 = 1.0, mur=1.0)
Hz2p = cal_Hz_from_Ey(Ey2p, kx2, w, mu0 = 1.0, mur=1.0)
# +
# check boundary
dEy = Ey1p + Ey1n - Ey2p
print( np.sum( np.abs(dEy) ) == 0.0 )
dHx = Hx1p + Hx1n - Hx2p
print( np.sum( np.abs(dHx) ) == 0.0 )
dHz = Hz1p + Hz1n - Hz2p
print( np.sum( np.abs(dHz) ) == 0.0 )
# +
# TM mode
Hy1p = np.zeros( theta1.shape, dtype=np.complex ) # incidence
Ex1p = cal_Hx_from_Ey(Ey1p, kz1, w, mu0 = mu0, mur=1.0)
Ez1p = cal_Hz_from_Ey(Ey1p, kx1, w, mu0 = mu0, mur=1.0)
Hy1n = r_p_H * Hy1p
Ex1n = cal_Ex_from_Hy(Hy1n,-kz1, w, e0 = e0, er=n1**2)
Ez1n = cal_Ez_from_Hy(Hy1n, kx1, w, e0 = e0, er=n1**2)
Hy2p = tau_p_H * Hy1p
Ex2p = cal_Ex_from_Hy(Hy2p, kz2, w, e0 = e0, er=n2**2)
Ez2p = cal_Ez_from_Hy(Hy2p, kx2, w, e0 = e0, er=n2**2)
# +
# check boundary
dHy = Hy1p + Hy1n - Hy2p
print( np.sum( np.abs(dHy) ) == 0.0 )
dEx = Ex1p + Ex1n - Ex2p
print( np.sum( np.abs(dEx) ) == 0.0 )
dEz = Ez1p + Ez1n - Ez2p
print( np.sum( np.abs(dEz) ) == 0.0 )
# -
# From the above calculation, we found that all the boundary conditions holds. However, in the above calculation, we suppose $k_y = 0.0$, material 1 and material 2 are non-magnetic ($\mu_r = 1.0$), and only consider 1 interface. In the following section. We'll going to introduce more powerful method, called transfer matrix method (TMM), to solve this questions.
# ## The results are wrong when $\theta_i > \theta_c$ and we will deal with this in Lecture 6-2.
|
Lecture-05-2 Fresnel Equation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.10.2 64-bit
# language: python
# name: python3
# ---
# +
# taking input from year
number = int(input("Enter any number: "))
number=67
#prime number is always greater than 1
if number > 1:
for i in range(2,number):
if (number % i) == 0:
print (number,"is not a prime number")
break
else:
print(number,"is not a prime number")
#if the entered number is les than or equal to 1
# then it is not prime number
|
Know Your Code/Python/Python Notes/Programs/Program 2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7 (pytorch)
# language: python
# name: pytorch
# ---
# # importing libraries
import os
import numpy as np
import glob
import torch
import torch.nn as nn
import torch.optim as optim
from torchvision.transforms import transforms
from torch.utils.data import DataLoader
from torch.utils.data import Dataset
import pathlib
import torchvision
import matplotlib.pyplot as plt
import torch.nn.functional as F
from pprint import pprint
import torchinfo
from torch.autograd import Variable
from pytorch_model_summary import summary
from tqdm import tqdm
import random
from torchvision.utils import make_grid
import torchvision.transforms.functional as TF
import cv2
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
print(device)
# transformation
transformer = transforms.Compose([
transforms.Grayscale(),
transforms.ToTensor(),
transforms.Normalize([0.5], [0.5]),
])
# # path to dataset
# +
train_path = r'C:/Users/utkar/Desktop/ML/pytorch/presentData/captcha_dataset/train_dataset'
test_path = r'C:/Users/utkar/Desktop/ML/pytorch/presentData/captcha_dataset/test_dataset'
train_loader = DataLoader(
torchvision.datasets.ImageFolder(train_path, transform=transformer),
batch_size = 16,
shuffle = True,
num_workers=4,
pin_memory=True,
)
test_loader = DataLoader(
torchvision.datasets.ImageFolder(test_path, transform=transformer),
shuffle = False,
num_workers=4,
pin_memory=True
)
# -
# # label dictionary
labels_map = {
0: "2", 1: "3", 2: "4", 3: "5", 4: "6", 5: "7", 6: "8", 7: "9", 8: "A", 9: "B", 10: "C", 11: "D", 12: "E", 13: "F",
14: "G", 15: "H", 16: "J", 17: "K", 18: "L", 19: "M", 20: "P", 21: "Q", 22: "R", 23: "S", 24: "T", 25: "U", 26: "V",
27: "W",28: "X", 29: "Y", 30: "a", 31: "b", 32: "c", 33: "d", 34: "e", 35: "f", 36: "h", 37: "j", 38: "k", 39: "m",
40: "n", 41: "p", 42: "q", 43: "r", 44: "s", 45: "t", 46: "u", 47: "v", 48: "w", 49: "x", 50: "y",
}
# +
a = 0
fig = plt.figure(figsize=(10, 10))
for i in range(4):
b = np.random.randint(150, 3400)
img, _ = train_loader.dataset[b]
fig.add_subplot(2, 2, a+1)
plt.imshow(img.permute(1, 2, 0), cmap="gray")
a += 1
if a == 4:
a = 0
break
plt.show()
# -
# classes
root = pathlib.Path(train_path)
classes = sorted([j.name.split('/')[-1] for j in root.iterdir()])
print(classes)
print(len(classes))
class CNNModel(nn.Module):
def __init__(self):
super(CNNModel, self).__init__()
self.network = nn.Sequential(
nn.Conv2d(1, 64, kernel_size=3, stride=1, padding=1),
nn.MaxPool2d(2, 2),
nn.Dropout(0.4),
nn.ReLU(),
nn.Conv2d(64, 64, kernel_size=3, stride=1, padding=1),
nn.MaxPool2d(2, 2),
nn.Dropout(0.4),
nn.ReLU(),
nn.Conv2d(64, 128, kernel_size=3, stride=1, padding=1),
nn.Dropout(0.4),
nn.ReLU(),
nn.Flatten(),
nn.Linear(8192, 128),
nn.Dropout(0.4),
nn.ReLU(),
nn.Linear(128, 256),
nn.Dropout(0.4),
nn.ReLU(),
nn.Linear(256, 51),
nn.LogSoftmax(dim=1)
)
def forward(self, x):
x = self.network(x)
return x
model = CNNModel().to(device)
print(model)
pprint(torchinfo.summary(model, input_size=(1, 1, 35, 35)))
optimizer = optim.Adam(model.parameters(), lr=0.001)
loss_fn = nn.CrossEntropyLoss()
train_count = len(glob.glob(train_path + '/**/*.png'))
test_count = len(glob.glob(test_path + '/**/*.png'))
print(train_count)
print(test_count)
print(train_loader.dataset)
# # training model
# +
epochs = 10
acc = 0.0
for epoch in range(epochs):
model.train()
train_acc = 0.0
train_loss = 0.0
for i, (images, labels) in enumerate(train_loader):
images = Variable(images.cuda())
labels = Variable(labels.cuda())
optimizer.zero_grad()
outputs = model(images)
loss = loss_fn(outputs, labels)
loss.backward()
optimizer.step()
train_loss += loss.cpu().data*images.size(0)
_, prediction = torch.max(outputs.data, 1)
train_acc += int(torch.sum(prediction == labels.data))
train_acc = train_acc/train_count
train_loss = train_loss/train_count
model.eval()
test_acc = 0
print('|----------------------------------------------------------------------------------------|')
for i, (images, labels)in enumerate(test_loader):
images = Variable(images.cuda())
labels = Variable(labels.cuda())
outputs = model(images)
_, prediction = torch.max(outputs.data, 1)
test_acc += int(torch.sum(prediction == labels.data))
test_acc = test_acc/test_count
print(f'epoch {epoch+1} training loss {train_loss} training accuracy {train_acc}')
print(f'testing accuracy {test_acc}')
# -
print(train_acc)
print(train_loss)
# +
a = 0
fig = plt.figure(figsize=(5, 5))
b = np.random.randint(90, 900)
b1 = np.random.randint(50, 100)
for i in range(4):
img, _ = test_loader.dataset[i*b1]
img1 = img.unsqueeze(0)
print(img1.shape)
img1 = Variable(img1.cuda())
outputs = model(img1)
_, prediction = torch.max(outputs.data, 1)
fig.add_subplot(2, 2, a+1)
plt.imshow(img.permute(1, 2, 0), cmap="gray")
plt.title(labels_map[prediction.item()])
a += 1
if a == 4:
a = 0
break
plt.show()
# -
# # saving the model
torch.save(model.state_dict(), 'captcha_model.pth')
|
model for captcha training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Torch Core
# This module contains all the basic functions we need in other modules of the fastai library (split with [`core`](/core.html#core) that contains the ones not requiring pytorch). Its documentation can easily be skipped at a first read, unless you want to know what a given fuction does.
# + hide_input=true
from fastai.gen_doc.nbdoc import *
from fastai.torch_core import *
# -
# ## Global constants
# `AdamW = partial(optim.Adam, betas=(0.9,0.99))` <div style="text-align: right"><a href="https://github.com/fastai/fastai/blob/master/fastai/torch_core.py#L43">[source]</a></div>
# `bn_types = (nn.BatchNorm1d, nn.BatchNorm2d, nn.BatchNorm3d)` <div style="text-align: right"><a href="https://github.com/fastai/fastai/blob/master/fastai/torch_core.py#L41">[source]</a></div>
# `default_device = torch.device('cuda') if torch.cuda.is_available() else torch.device('cpu')` <div style="text-align: right"><a href="https://github.com/fastai/fastai/blob/master/fastai/torch_core.py#L42">[source]</a></div>
# ## Functions that operate conversions
# + hide_input=true
show_doc(flatten_model, full_name='flatten')
# -
# Flattens all the layers of `m`.
# + hide_input=true
show_doc(model2half)
# + hide_input=true
show_doc(np2model_tensor)
# + hide_input=true
show_doc(requires_grad)
# + hide_input=true
show_doc(tensor)
# + hide_input=true
show_doc(to_cpu)
# + hide_input=true
show_doc(to_data)
# + hide_input=true
show_doc(to_detach)
# + hide_input=true
show_doc(to_device)
# + hide_input=true
show_doc(to_half)
# + hide_input=true
show_doc(to_np)
# + hide_input=true
show_doc(try_int)
# -
# ## Functions to deal with model initialization
# + hide_input=true
show_doc(apply_init)
# + hide_input=true
show_doc(apply_leaf)
# + hide_input=true
show_doc(cond_init)
# + hide_input=true
show_doc(in_channels)
# + hide_input=true
show_doc(init_default)
# -
# ## Functions to get information of a model
# + hide_input=true
show_doc(children)
# + hide_input=true
show_doc(first_layer)
# + hide_input=true
show_doc(last_layer)
# + hide_input=true
show_doc(num_children)
# + hide_input=true
show_doc(one_param)
# + hide_input=true
show_doc(range_children)
# + hide_input=true
show_doc(trainable_params)
# -
# ## Functions to deal with BatchNorm layers
# + hide_input=true
show_doc(bn2float)
# + hide_input=true
show_doc(set_bn_eval)
# + hide_input=true
show_doc(split_bn_bias)
# -
# ## Functions to get random tensors
# + hide_input=true
show_doc(log_uniform)
# -
log_uniform(0.5,2,(8,))
# + hide_input=true
show_doc(rand_bool)
# -
rand_bool(0.5, 8)
# + hide_input=true
show_doc(uniform)
# -
uniform(0,1,(8,))
# + hide_input=true
show_doc(uniform_int)
# -
uniform_int(0,2,(8,))
# ## Other functions
# + hide_input=true
show_doc(FloatItem, title_level=3)
# + hide_input=true
show_doc(calc_loss)
# + hide_input=true
show_doc(data_collate)
# + hide_input=true
show_doc(grab_idx)
# + hide_input=true
show_doc(logit)
# + hide_input=true
show_doc(logit_)
# + hide_input=true
show_doc(model_type)
# + hide_input=true
show_doc(np_address)
# + hide_input=true
show_doc(split_model)
# -
# If `splits` are layers, the model is split at those (not included) sequentially. If `want_idxs` is True, the corresponding indexes are returned. If `splits` are lists of layers, the model is split according to those.
# + hide_input=true
show_doc(split_model_idx)
# + hide_input=true
show_doc(trange_of)
# -
# ## Undocumented Methods - Methods moved below this line will intentionally be hidden
show_doc(tensor__array__)
# ## New Methods - Please document or move to the undocumented section
|
docs_src/torch_core.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + slideshow={"slide_type": "slide"}
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
import matplotlib.dates as mdates
from matplotlib.collections import LineCollection
import plotly
import plotly.graph_objects as go
# +
file = r'Pistachio_Low.csv'
df = pd.read_csv(file)
df['datetime'] = pd.to_datetime(df['DATE'])
#df
df2 = df.set_index(df.datetime)
#df2 = df2.drop(['datetime'], axis=1)
df2 = df2.drop(['DATE'], axis=1)
df2 = df2.sort_index() # For the monthly file, 1/1/16-7/1/16 were misplaced. Have to rank this for plotting
df_nan = df2[np.isnan(df2['ET'])]
# Use linear interpolation to simulate gap data that match the interval
filled = df2.interpolate(limit=2, method='linear')
df_filled = df_nan.merge(filled, left_index=True, right_index=True, how='left')
df_filled.drop(df_filled.columns[[0, 1, 2]], axis = 1, inplace = True)
xmin = min(df2.index)
xmax = max(df2.index)
# +
# Creating masks for periods with gaps
s1mask = np.isfinite(df2.ET)
s2mask = np.isfinite(df2.ET_corr)
# Creating masks for models
SIMSmask = np.isfinite(df2.SIMS_3x3)
# DAmask = np.isfinite(df2.DISALEXI_3x3)
EEmask = np.isfinite(df2.EEMETRIC_3x3)
PTmask = np.isfinite(df2.PTJPL_3x3)
BOPmask = np.isfinite(df2.SSEBOP_3x3)
ENmask = np.isfinite(df2.ensemble_mean_3x3)
# +
fig = go.Figure()
# ----------------
# <<< --- |Adjustement Area| --- >>>
# ----------------
# Trace 1 -> Unclosed flux
fig.add_trace(go.Scatter(
x=df2.index,
y=df2.ET,
name = 'Unclosed ETa',
fill= 'none',
mode='lines',
line_color='dimgrey',
line_width=0,
showlegend=False))
# Trace 2 -> Closed flux for establish band
fig.add_trace(go.Scatter(
x=df2.index,
y=df2.ET_corr,
name = 'Energy Imbalance Band',
fill= 'tonexty',
mode='lines',
line_color='dimgrey',
line_width=0))
# ---------------
# <<< --- |Model Variables| --- >>>
# ---------------
# Trace 3 -> SIMS
fig.add_trace(go.Scatter(
x=df2.index[SIMSmask],
y=df2.SIMS_3x3[SIMSmask],
name="SIMS_ETc",
mode='lines+markers',
line_color='royalblue',
opacity=0.8,
marker = dict(color='royalblue', size = 8, symbol = 0, line=dict(width=1,
color='black'))))
# Trace 4 -> DISALEXI
# fig.add_trace(go.Scatter(
# x=df2.index[DAmask],
# y=df2.DISALEXI_3x3[DAmask],
# name="DisALEXI_ETc",
# mode='markers', #'lines+markers'
# #line_color='BurlyWood',
# opacity=0.8,
# marker = dict(color='BurlyWood', size = 8, symbol = 2, line=dict(width=1,
# color='black'))))
# Trace 5 -> EEMETRIC
fig.add_trace(go.Scatter(
x=df2.index[EEmask],
y=df2.EEMETRIC_3x3[EEmask],
name="EEMETRIC_ETc",
mode='lines+markers',
line_color='coral',
opacity=0.8,
marker = dict(color='coral', size = 8, symbol = 1, line=dict(width=1,
color='black'))))
# Trace 6 -> PTJPL
fig.add_trace(go.Scatter(
x=df2.index[PTmask],
y=df2.PTJPL_3x3[PTmask],
name="PTJPL_ETc",
mode='lines+markers',
line_color='MediumSpringGreen',
opacity=0.8,
marker = dict(color='MediumSpringGreen', size = 8, symbol = 5, line=dict(width=1,
color='black'))))
# Trace 7 -> SSEBOP
fig.add_trace(go.Scatter(
x=df2.index[BOPmask],
y=df2.SSEBOP_3x3[BOPmask],
name="SSEBOP_ETc",
mode='lines+markers',
line_color='orchid',
opacity=0.8,
marker = dict(color='orchid', size = 8, symbol = 14, line=dict(width=1,
color='black'))))
# Trace 8 -> Ensemble
fig.add_trace(go.Scatter(
x=df2.index[ENmask],
y=df2.ensemble_mean_3x3[ENmask],
name="Ensemble_ETc",
mode='lines+markers',
line_color='red',
opacity=0.7,
marker = dict(color='red', size = 9, symbol = 17, line=dict(width=1,
color='black'))))
# ---------
# <<< ---- |Flux Data| ---- >>>
# ---------
# Trace 9 -> Unclosed flux for reference
fig.add_trace(go.Scatter(
x=df2.index,
y=df2.ET,
name = 'Unclosed ETa',
mode='none',
showlegend=False,
marker=dict(color='rgba(0, 0, 0, 0)',
size=4,
line=dict(color='red',width=1))))
# Trace 10 -> Closed flux for reference
fig.add_trace(go.Scatter(
x=df2.index,
y=df2.ET_corr,
name = 'Closed ETa',
mode='none',
showlegend=False,
marker=dict(color='rgba(0, 0, 0, 0)',
size=4,
line=dict(color='black',width=1))))
#fig.update_xaxes(showgrid=True, gridwidth=1, gridcolor='darkgray')
fig.update_layout(
autosize=True,
title={'text':"Intercomparson of Monthly ET (mm) Pistachio Low Salinity [3x3 Grid] <br> EBC = (H+LE)/(Rn-G) = 0.71; Correction Factor = 1/EBC = 1.41",
'x':0.145,
'y':0.93,
'xanchor': 'left',
'yanchor': 'top'},
xaxis_title= "Date Slider",
yaxis_title= "Evapotranspiration (mm)",
xaxis_rangeslider_visible=True,
plot_bgcolor='rgba(0,0,0,0)',
titlefont=dict(
family="Times New Roman",
size=22,
color='black'),
font=dict(
family="Times New Roman",
size=18,
color='black'))
# Add Dropdown Menu Option
# For visible: all traces are ranked sequentially as listed above (e.g. Trace#)
fig.update_layout(
updatemenus=[
dict(
active=0,
buttons=list([
dict(label="All",
method="update",
args=[{"visible": [True]},
{"title": "Intercomparson of Monthly ET (mm) Pistachio Low Salinity [3x3 Grid] <br> EBC = (H+LE)/(Rn-G) = 0.71; Correction Factor = 1/EBC = 1.41",
}]),
dict(label="SIMS",
method="update",
args=[{"visible": [True, True, True, False, False, False, True, True, True]},
{"title": "SIMS Monthly Evapotranspiration (mm)",
}]),
dict(label="EEMETRIC",
method="update",
args=[{"visible": [True, True, False, True, False, False, True, True, True]},
{"title": "EEMETRIC Monthly Evapotranspiration (mm)",
}]),
dict(label="PT-JPL",
method="update",
args=[{"visible": [True, True, False, False, True, False, True, True, True]},
{"title": "PT-JPL Monthly Evapotranspiration (mm)",
}]),
dict(label="SSEBop",
method="update",
args=[{"visible": [True, True, False, False, False, True, True, True, True]},
{"title": "SSEBop Monthly Evapotranspiration (mm)",
}]),
dict(label="Ensemble",
method="update",
args=[{"visible": [True, True, False, False, False, False, True, True, True]},
{"title": "Ensemble Monthly Evapotranspiration (mm)",
}]),
]),
direction="down",
pad={"r": 10, "t": 10},
showactive=True,
x=-0.23,
xanchor="left",
y=1.08,
yanchor="top")],
annotations=[
dict(text="<b>Traces</b>", x=-0.23, xref="paper", y=1.12, yref="paper",
align="left", showarrow=False)])
fig.update_xaxes(showline=True,
linewidth=1,
linecolor='black',
mirror=True)
#showgrid=True,
#gridwidth=1,
#gridcolor='Gainsboro')
fig.update_yaxes(showline=True, linewidth=1, linecolor='black', mirror=True)
fig.show(config={'scrollZoom': True})
# -
# The plot above isn't well sized, when exporting, the .html will autosize
plotly.offline.plot(fig, config={'scrollZoom': True}, filename = 'Monthly_Pist_Low_INT_3x3.html', auto_open=False)
|
Timeseries/Monthly/Monthly_Intercomparison.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:packt]
# language: python
# name: conda-env-packt-py
# ---
# # One Hot Encoding
# ## Manual One-Hot Encoding
def onehot_word(word):
lookup = {v[1]: v[0] for v in enumerate(set(word))}
word_vector = []
for c in word:
one_hot_vector = [0] * len(lookup)
one_hot_vector[lookup[c]] = 1
word_vector.append(one_hot_vector)
return word_vector
onehot_word('data')
# ## One Hot Encoding Using Keras
import keras
keras.preprocessing.text.one_hot('dawn of man', n=5)
import keras
from keras.preprocessing.text import Tokenizer
from keras.utils import to_categorical
from keras.preprocessing.text import one_hot
import pandas as pd
# +
with open('../data/100lines.txt') as lines_file:
movie_lines = [ line.strip() for line in lines_file.readlines()]
movie_lines[:10]
# -
tokenizer = Tokenizer(char_level=True)
tokenizer.fit_on_texts(movie_lines)
lines_as_integers = tokenizer.texts_to_sequences(movie_lines)
lines_as_integers[:4]
vocabulary = set(int_value for line_values in lines_as_integers for int_value in line_values)
vocabulary_size = len(vocabulary)
movie_lines_one_hot = [keras.utils.to_categorical(line, vocabulary_size+1) for line in lines_as_integers]
movie_lines = pd.read_csv('../data/100lines.txt', sep='\t', header=None)
movie_lines.columns = ['line']
movie_lines_one_hot[0:4]
tokenizer.fit_on_texts(movie_lines)
int_sequence = tokenizer.texts_to_sequences(movie_lines)
int_sequence
vocabulary = set(int_value for line_values in int_sequence for int_value in line_values)
vocabulary_size = len(vocabulary)
keras.utils.to_categorical([1,2,3], vocabulary_size)
to_categorical([1,2,3,1, 0], 5)
import numpy as np
np.array(int_sequence)
movie_lines
# ### Character Level Encoding Using Keras
text = '<NAME> for man'
from keras.preprocessing.text import Tokenizer
import numpy as np
char_tokenizer = Tokenizer(char_level=True)
char_tokenizer.fit_on_texts(text)
char_tokenizer.texts_to_sequences(text)
char_tokenizer.index_word
char_tokenizer.word_index
char_vectors = char_tokenizer.texts_to_matrix(text)
char_vectors
char_vectors.shape
char_tokenizer.index_word[np.argmax(char_vectors[0])]
# ## One Hot Encoding Words
# +
with open('../data/100lines.txt') as lines_file:
movie_lines = [ line.strip() for line in lines_file.readlines()]
movie_lines[:10]
# -
lines_array = np.array(movie_lines)
lines_array.reshape(-1,1)
lines_array.shape
from sklearn import preprocessing
wordOneHotEncoder = preprocessing.OneHotEncoder()
labelEncoder = preprocessing.LabelEncoder()
movie_labels = labelEncoder.fit_transform(lines_array)
movie_labels
movie_labels.reshape(-1,1)
movie_onehot = wordOneHotEncoder.fit_transform(movie_labels.reshape(-1,1))
movie_onehot.toarray()
|
Lesson7/completed_notebooks/Exercise57.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# ### Exercice 1:
# Write a Python class named square constructed by a length and two methods which will compute the area and the perimeter of the square.
class square():
def __init__(self,length):
self.length = length
print("its here")
def s_area(self):
return self.length * self.length
def r_perimeter(self):
return 4 * self.length
#a = square(3)
#print(a.r_perimeter())
#print(a.s_area())
# ### Exercice 2:
# Write a python class rectangle that inherits from the square class.
# +
class rectangle(square):
#__init__ method only
def __init__(self,largeur, lenght):
self.largeur = largeur
super(rectangle, self).__init__(lenght)
pass
r = rectangle(20, 50)
print(r.r_perimeter())
print(r.s_area())
# -
# ### Exercice 3:
# +
class SampleClass:
def __init__(self, a):
self.a = a
x = SampleClass(3)
print(x.a)
x.a = 23
print(x.a)
# -
# Use python decorators to make the above code works
|
exercices/part4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # HM2: Numerical Optimization for Logistic Regression.
#
# ### Name: <NAME>
#
# ## 0. You will do the following:
#
# 1. Read the lecture note: [click here](https://github.com/wangshusen/DeepLearning/blob/master/LectureNotes/Logistic/paper/logistic.pdf)
#
# 2. Read, complete, and run my code.
#
# 3. **Implement mini-batch SGD** and evaluate the performance.
#
# 4. Convert the .IPYNB file to .HTML file.
#
# * The HTML file must contain **the code** and **the output after execution**.
#
# * Missing **the output after execution** will not be graded.
#
# 5. Upload this .HTML file to your Google Drive, Dropbox, or your Github repo. (If you submit the file to Google Drive or Dropbox, you must make the file "open-access". The delay caused by "deny of access" may result in late penalty.)
#
# 6. Submit the link to this .HTML file to Canvas.
#
# * Example: https://github.com/wangshusen/CS583-2020S/blob/master/homework/HM2/HM2.html
#
#
# ## Grading criteria:
#
# 1. When computing the ```gradient``` and ```objective function value``` using a batch of samples, use **matrix-vector multiplication** rather than a FOR LOOP of **vector-vector multiplications**.
#
# 2. Plot ```objective function value``` against ```epochs```. In the plot, compare GD, SGD, and MB-SGD (with $b=8$ and $b=64$). The plot must look reasonable.
# # 1. Data processing
#
# - Download the Diabete dataset from https://www.csie.ntu.edu.tw/~cjlin/libsvmtools/datasets/binary/diabetes
# - Load the data using sklearn.
# - Preprocess the data.
# ## 1.1. Load the data
# +
from sklearn import datasets
import numpy
x_sparse, y = datasets.load_svmlight_file('diabetes')
x = x_sparse.todense()
print('Shape of x: ' + str(x.shape))
print('Shape of y: ' + str(y.shape))
# -
# ## 1.2. Partition to training and test sets
# +
# partition the data to training and test sets
n = x.shape[0]
n_train = 640
n_test = n - n_train
rand_indices = numpy.random.permutation(n)
train_indices = rand_indices[0:n_train]
test_indices = rand_indices[n_train:n]
x_train = x[train_indices, :]
x_test = x[test_indices, :]
y_train = y[train_indices].reshape(n_train, 1)
y_test = y[test_indices].reshape(n_test, 1)
print('Shape of x_train: ' + str(x_train.shape))
print('Shape of x_test: ' + str(x_test.shape))
print('Shape of y_train: ' + str(y_train.shape))
print('Shape of y_test: ' + str(y_test.shape))
# -
# ## 1.3. Feature scaling
# Use the standardization to trainsform both training and test features
# +
# Standardization
import numpy
# calculate mu and sig using the training set
d = x_train.shape[1]
mu = numpy.mean(x_train, axis=0).reshape(1, d)
sig = numpy.std(x_train, axis=0).reshape(1, d)
# transform the training features
x_train = (x_train - mu) / (sig + 1E-6)
# transform the test features
x_test = (x_test - mu) / (sig + 1E-6)
print('test mean = ')
print(numpy.mean(x_test, axis=0))
print('test std = ')
print(numpy.std(x_test, axis=0))
# -
# ## 1.4. Add a dimension of all ones
# +
n_train, d = x_train.shape
x_train = numpy.concatenate((x_train, numpy.ones((n_train, 1))), axis=1)
n_test, d = x_test.shape
x_test = numpy.concatenate((x_test, numpy.ones((n_test, 1))), axis=1)
print('Shape of x_train: ' + str(x_train.shape))
print('Shape of x_test: ' + str(x_test.shape))
# -
# # 2. Logistic regression model
#
# The objective function is $Q (w; X, y) = \frac{1}{n} \sum_{i=1}^n \log \Big( 1 + \exp \big( - y_i x_i^T w \big) \Big) + \frac{\lambda}{2} \| w \|_2^2 $.
# Calculate the objective function value
# Inputs:
# w: d-by-1 matrix
# x: n-by-d matrix
# y: n-by-1 matrix
# lam: scalar, the regularization parameter
# Return:
# objective function value (scalar)
def objective(w, x, y, lam):
n, d = x.shape
yx = numpy.multiply(y, x) # n-by-d matrix
yxw = numpy.dot(yx, w) # n-by-1 matrix
vec1 = numpy.exp(-yxw) # n-by-1 matrix
vec2 = numpy.log(1 + vec1) # n-by-1 matrix
loss = numpy.mean(vec2) # scalar
reg = lam / 2 * numpy.sum(w * w) # scalar
return loss + reg
# +
# initialize w
d = x_train.shape[1]
w = numpy.zeros((d, 1))
# evaluate the objective function value at w
lam = 1E-6
objval0 = objective(w, x_train, y_train, lam)
print('Initial objective function value = ' + str(objval0))
# -
# # 3. Numerical optimization
# ## 3.1. Gradient descent
#
# The gradient at $w$ is $g = - \frac{1}{n} \sum_{i=1}^n \frac{y_i x_i }{1 + \exp ( y_i x_i^T w)} + \lambda w$
# Calculate the gradient
# Inputs:
# w: d-by-1 matrix
# x: n-by-d matrix
# y: n-by-1 matrix
# lam: scalar, the regularization parameter
# Return:
# g: g: d-by-1 matrix, full gradient
def gradient(w, x, y, lam):
n, d = x.shape
yx = numpy.multiply(y, x) # n-by-d matrix
yxw = numpy.dot(yx, w) # n-by-1 matrix
vec1 = numpy.exp(yxw) # n-by-1 matrix
vec2 = numpy.divide(yx, 1+vec1) # n-by-d matrix
vec3 = -numpy.mean(vec2, axis=0).reshape(d, 1) # d-by-1 matrix
g = vec3 + lam * w
return g
# Gradient descent for solving logistic regression
# Inputs:
# x: n-by-d matrix
# y: n-by-1 matrix
# lam: scalar, the regularization parameter
# stepsize: scalar
# max_iter: integer, the maximal iterations
# w: d-by-1 matrix, initialization of w
# Return:
# w: d-by-1 matrix, the solution
# objvals: a record of each iteration's objective value
def grad_descent(x, y, lam, stepsize, max_iter=100, w=None):
n, d = x.shape
objvals = numpy.zeros(max_iter) # store the objective values
if w is None:
w = numpy.zeros((d, 1)) # zero initialization
for t in range(max_iter):
objval = objective(w, x, y, lam)
objvals[t] = objval
print('Objective value at t=' + str(t) + ' is ' + str(objval))
g = gradient(w, x, y, lam)
w -= stepsize * g
return w, objvals
# Run gradient descent.
lam = 1E-6
stepsize = 1.0
w, objvals_gd = grad_descent(x_train, y_train, lam, stepsize)
# ## 3.2. Stochastic gradient descent (SGD)
#
# Define $Q_i (w) = \log \Big( 1 + \exp \big( - y_i x_i^T w \big) \Big) + \frac{\lambda}{2} \| w \|_2^2 $.
#
# The stochastic gradient at $w$ is $g_i = \frac{\partial Q_i }{ \partial w} = -\frac{y_i x_i }{1 + \exp ( y_i x_i^T w)} + \lambda w$.
# Calculate the objective Q_i and the gradient of Q_i
# Inputs:
# w: d-by-1 matrix
# xi: 1-by-d matrix
# yi: scalar
# lam: scalar, the regularization parameter
# Return:
# obj: scalar, the objective Q_i
# g: d-by-1 matrix, gradient of Q_i
def stochastic_objective_gradient(w, xi, yi, lam):
d = xi.shape[0]
yx = yi * xi # 1-by-d matrix
yxw = float(numpy.dot(yx, w)) # scalar
# calculate objective function Q_i
loss = numpy.log(1 + numpy.exp(-yxw)) # scalar
reg = lam / 2 * numpy.sum(w * w) # scalar
obj = loss + reg
# calculate stochastic gradient
g_loss = -yx.T / (1 + numpy.exp(yxw)) # d-by-1 matrix
g = g_loss + lam * w # d-by-1 matrix
return obj, g
# SGD for solving logistic regression
# Inputs:
# x: n-by-d matrix
# y: n-by-1 matrix
# lam: scalar, the regularization parameter
# stepsize: scalar
# max_epoch: integer, the maximal epochs
# w: d-by-1 matrix, initialization of w
# Return:
# w: the solution
# objvals: record of each iteration's objective value
def sgd(x, y, lam, stepsize, max_epoch=100, w=None):
n, d = x.shape
objvals = numpy.zeros(max_epoch) # store the objective values
if w is None:
w = numpy.zeros((d, 1)) # zero initialization
for t in range(max_epoch):
# randomly shuffle the samples
rand_indices = numpy.random.permutation(n)
x_rand = x[rand_indices, :]
y_rand = y[rand_indices, :]
objval = 0 # accumulate the objective values
for i in range(n):
xi = x_rand[i, :] # 1-by-d matrix
yi = float(y_rand[i, :]) # scalar
obj, g = stochastic_objective_gradient(w, xi, yi, lam)
objval += obj
w -= stepsize * g
stepsize *= 0.9 # decrease step size
objval /= n
objvals[t] = objval
print('Objective value at epoch t=' + str(t) + ' is ' + str(objval))
return w, objvals
# Run SGD.
lam = 1E-6
stepsize = 0.1
w, objvals_sgd = sgd(x_train, y_train, lam, stepsize)
# # 4. Compare GD with SGD
#
# Plot objective function values against epochs.
# +
import matplotlib.pyplot as plt
# %matplotlib inline
fig = plt.figure(figsize=(6, 4))
epochs_gd = range(len(objvals_gd))
epochs_sgd = range(len(objvals_sgd))
line0, = plt.plot(epochs_gd, objvals_gd, '--b', LineWidth=4)
line1, = plt.plot(epochs_sgd, objvals_sgd, '-r', LineWidth=2)
plt.xlabel('Epochs', FontSize=20)
plt.ylabel('Objective Value', FontSize=20)
plt.xticks(FontSize=16)
plt.yticks(FontSize=16)
plt.legend([line0, line1], ['GD', 'SGD'], fontsize=20)
plt.tight_layout()
plt.show()
fig.savefig('compare_gd_sgd.pdf', format='pdf', dpi=1200)
# -
# # 5. Prediction
# Predict class label
# Inputs:
# w: d-by-1 matrix
# X: m-by-d matrix
# Return:
# f: m-by-1 matrix, the predictions
def predict(w, X):
xw = numpy.dot(X, w)
f = numpy.sign(xw)
return f
# evaluate training error
f_train = predict(w, x_train)
diff = numpy.abs(f_train - y_train) / 2
error_train = numpy.mean(diff)
print('Training classification error is ' + str(error_train))
# evaluate test error
f_test = predict(w, x_test)
diff = numpy.abs(f_test - y_test) / 2
error_test = numpy.mean(diff)
print('Test classification error is ' + str(error_test))
# # 6. Mini-batch SGD (fill the code)
#
#
# ?]## 6.1. Compute the objective $Q_I$ and its gradient using a batch of samples
#
# Define $Q_I (w) = \frac{1}{b} \sum_{i \in I} \log \Big( 1 + \exp \big( - y_i x_i^T w \big) \Big) + \frac{\lambda}{2} \| w \|_2^2 $, where $I$ is a set containing $b$ indices randomly drawn from $\{ 1, \cdots , n \}$ without replacement.
#
# The stochastic gradient at $w$ is $g_I = \frac{\partial Q_I }{ \partial w} = \frac{1}{b} \sum_{i \in I} \frac{- y_i x_i }{1 + \exp ( y_i x_i^T w)} + \lambda w$.
# Calculate the objective Q_I and the gradient of Q_I
# Inputs:
# w: d-by-1 matrix
# xi: b-by-d matrix
# yi: b-by-1 matrix
# lam: scalar, the regularization parameter
# b: integer, the batch size
# Return:
# obj: scalar, the objective Q_i
# g: d-by-1 matrix, gradient of Q_i
def mb_stochastic_objective_gradient(w, xi, yi, lam, b):
# Fill the function
# Follow the implementation of stochastic_objective_gradient
# Use matrix-vector multiplication; do not use FOR LOOP of vector-vector multiplications
yx = numpy.multiply(yi, xi)
yxw = numpy.dot(yx, w) # b-by-1 matrix
# calculate the objective function value
elem_loss = numpy.log(1 + numpy.exp(-yxw))
loss = numpy.sum(elem_loss, axis=0)
obj = float((1/b) * loss + (lam/2) * numpy.sum(w*w))
# calculate the gradient
elem_grad_loss = -yx / (1 + numpy.exp(yxw))
grad_loss = (1/b) * numpy.sum(elem_grad_loss, axis=0)
g = grad_loss.T + lam*w
return obj, g
# ## 6.2. Implement mini-batch SGD
#
# Hints:
# 1. In every epoch, randomly permute the $n$ samples (just like SGD).
# 2. Each epoch has $\frac{n}{b}$ iterations. In every iteration, use $b$ samples, and compute the gradient and objective using the ``mb_stochastic_objective_gradient`` function. In the next iteration, use the next $b$ samples, and so on.
#
# Mini-Batch SGD for solving logistic regression
# Inumpyuts:
# x: n-by-d matrix
# y: n-by-1 matrix
# lam: scalar, the regularization parameter
# b: integer, the batch size
# stepsize: scalar
# max_epoch: integer, the maximal epochs
# w: d-by-1 matrix, initialization of w
# Return:
# w: the solution
# objvals: record of each iteration's objective value
def mb_sgd(x, y, lam, b, stepsize, max_epoch=100, w=None):
# Fill the function
# Follow the implementation of sgd
# Record one objective value per epoch (not per iteration!)
n, d = x.shape
objvals: numpy.ndarray = numpy.zeros((max_epoch))
if n % b != 0:
raise ValueError("Dataset must be evenly divisible into batches. Invalid batch length")
if w is None:
w = numpy.zeros((d, 1)) # zero initialization
num_batches: int = n // b
for e in range(max_epoch):
rand_indices: numpy.ndarray = numpy.random.permutation(n)
x_rand: numpy.ndarray = x[rand_indices, ...]
y_rand: numpy.ndarray = y[rand_indices, ...]
x_batch: numpy.ndarray = numpy.split(x_rand, num_batches)
y_batch: numpy.ndarray = numpy.split(y_rand, num_batches)
objval = 0 # accumulate objective values
for i in range(num_batches):
obj, g = mb_stochastic_objective_gradient(w, x_batch[i], y_batch[i], lam, b)
objval += obj
w -= stepsize*g
stepsize *= 0.9 # decrease step size
objval /= num_batches
objvals[e] = objval
print(f'Objective value at epoch e={str(e)} is {str(numpy.round(objval, 5))}')
return w, objvals
# ## 6.3. Run MB-SGD
# MB-SGD with batch size b=8
lam = 1E-6 # do not change
b = 8 # do not change
stepsize = 0.1 # you must tune this parameter
# stepsize ~ 4 for .48338 convergence for batch size of 640
w, objvals_mbsgd8 = mb_sgd(x_train, y_train, lam, b, stepsize)
f_train = predict(w, x_train)
diff = numpy.abs(f_train - y_train) / 2
error_train = numpy.mean(diff)
print('Training classification error is ' + str(error_train))
# +
# MB-SGD with batch size b=64
lam = 1E-6 # do not change
b = 64 # do not change
stepsize = 0.6 # you must tune this parameter
w, objvals_mbsgd64 = mb_sgd(x_train, y_train, lam, b, stepsize)
# -
f_train = predict(w, x_train)
diff = numpy.abs(f_train - y_train) / 2
error_train = numpy.mean(diff)
print('Training classification error is ' + str(error_train))
# # 7. Plot and compare GD, SGD, and MB-SGD
# You are required to compare the following algorithms:
#
# - Gradient descent (GD)
#
# - SGD
#
# - MB-SGD with b=8
#
# - MB-SGD with b=64
#
# Follow the code in Section 4 to plot ```objective function value``` against ```epochs```. There should be four curves in the plot; each curve corresponds to one algorithm.
# Hint: Logistic regression with $\ell_2$-norm regularization is a strongly convex optimization problem. All the algorithms will converge to the same solution. **In the end, the ``objective function value`` of the 4 algorithms will be the same. If not the same, your implementation must be wrong. Do NOT submit wrong code and wrong result!**
# +
# plot the 4 curves:
# +
fig = plt.figure(figsize=(6, 4))
epochs_mbsgd8 = range(len(objvals_mbsgd8))
epochs_mbsgd64 = range(len(objvals_mbsgd64))
line0, = plt.plot(epochs_mbsgd8, objvals_mbsgd8, '--b', LineWidth=4)
line1, = plt.plot(epochs_mbsgd64, objvals_mbsgd64, '-r', LineWidth=2)
line2, = plt.plot(epochs_sgd, objvals_sgd, '--m', LineWidth=3)
line3, = plt.plot(epochs_gd, objvals_gd, '-g', LineWidth=1)
plt.xlabel('Epochs', FontSize=20)
plt.ylabel('Objective Value', FontSize=20)
plt.xticks(FontSize=16)
plt.yticks(FontSize=16)
plt.legend([line0, line1, line2, line3], ["MBSGD8", "MBSGD64", "SGD", "GD"], fontsize=20)
plt.tight_layout()
plt.show()
fig.savefig('compare_mbsgd_gd_sgd.pdf', format='pdf', dpi=1200)
|
assignments/HM2/HM2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="DweYe9FcbMK_"
# ##### Copyright 2018 The TensorFlow Authors.
#
#
# + cellView="form" colab={} colab_type="code" id="AVV2e0XKbJeX"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="sUtoed20cRJJ"
# # Load text
# + [markdown] colab_type="text" id="1ap_W4aQcgNT"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/load_data/text"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/load_data/text.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/load_data/text.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/load_data/text.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="NWeQAo0Ec_BL"
# This tutorial provides an example of how to use `tf.data.TextLineDataset` to load examples from text files. `TextLineDataset` is designed to create a dataset from a text file, in which each example is a line of text from the original file. This is potentially useful for any text data that is primarily line-based (for example, poetry or error logs).
#
# In this tutorial, we'll use three different English translations of the same work, Homer's Illiad, and train a model to identify the translator given a single line of text.
# + [markdown] colab_type="text" id="fgZ9gjmPfSnK"
# ## Setup
# + colab={} colab_type="code" id="baYFZMW_bJHh"
from __future__ import absolute_import, division, print_function, unicode_literals
try:
# # %tensorflow_version only exists in Colab.
# !pip install tf-nightly
except Exception:
pass
import tensorflow as tf
import tensorflow_datasets as tfds
import os
# + [markdown] colab_type="text" id="YWVWjyIkffau"
# The texts of the three translations are by:
#
# - [<NAME>](https://en.wikipedia.org/wiki/William_Cowper) — [text](https://storage.googleapis.com/download.tensorflow.org/data/illiad/cowper.txt)
#
# - [<NAME>](https://en.wikipedia.org/wiki/Edward_Smith-Stanley,_14th_Earl_of_Derby) — [text](https://storage.googleapis.com/download.tensorflow.org/data/illiad/derby.txt)
#
# - [<NAME>](https://en.wikipedia.org/wiki/Samuel_Butler_%28novelist%29) — [text](https://storage.googleapis.com/download.tensorflow.org/data/illiad/butler.txt)
#
# The text files used in this tutorial have undergone some typical preprocessing tasks, mostly removing stuff — document header and footer, line numbers, chapter titles. Download these lightly munged files locally.
# + colab={} colab_type="code" id="4YlKQthEYlFw"
DIRECTORY_URL = 'https://storage.googleapis.com/download.tensorflow.org/data/illiad/'
FILE_NAMES = ['cowper.txt', 'derby.txt', 'butler.txt']
for name in FILE_NAMES:
text_dir = tf.keras.utils.get_file(name, origin=DIRECTORY_URL+name)
parent_dir = os.path.dirname(text_dir)
parent_dir
# + [markdown] colab_type="text" id="q3sDy6nuXoNp"
# ## Load text into datasets
#
# Iterate through the files, loading each one into its own dataset.
#
# Each example needs to be individually labeled, so use `tf.data.Dataset.map` to apply a labeler function to each one. This will iterate over every example in the dataset, returning (`example, label`) pairs.
# + colab={} colab_type="code" id="K0BjCOpOh7Ch"
def labeler(example, index):
return example, tf.cast(index, tf.int64)
labeled_data_sets = []
for i, file_name in enumerate(FILE_NAMES):
lines_dataset = tf.data.TextLineDataset(os.path.join(parent_dir, file_name))
labeled_dataset = lines_dataset.map(lambda ex: labeler(ex, i))
labeled_data_sets.append(labeled_dataset)
# + [markdown] colab_type="text" id="M8PHK5J_cXE5"
# Combine these labeled datasets into a single dataset, and shuffle it.
#
# + colab={} colab_type="code" id="6jAeYkTIi9-2"
BUFFER_SIZE = 50000
BATCH_SIZE = 64
TAKE_SIZE = 5000
# + colab={} colab_type="code" id="Qd544E-Sh63L"
all_labeled_data = labeled_data_sets[0]
for labeled_dataset in labeled_data_sets[1:]:
all_labeled_data = all_labeled_data.concatenate(labeled_dataset)
all_labeled_data = all_labeled_data.shuffle(
BUFFER_SIZE, reshuffle_each_iteration=False)
# + [markdown] colab_type="text" id="r4JEHrJXeG5k"
# You can use `tf.data.Dataset.take` and `print` to see what the `(example, label)` pairs look like. The `numpy` property shows each Tensor's value.
# + colab={} colab_type="code" id="gywKlN0xh6u5"
for ex in all_labeled_data.take(5):
print(ex)
# + [markdown] colab_type="text" id="5rrpU2_sfDh0"
# ## Encode text lines as numbers
#
# Machine learning models work on numbers, not words, so the string values need to be converted into lists of numbers. To do that, map each unique word to a unique integer.
#
# ### Build vocabulary
#
# First, build a vocabulary by tokenizing the text into a collection of individual unique words. There are a few ways to do this in both TensorFlow and Python. For this tutorial:
#
# 1. Iterate over each example's `numpy` value.
# 2. Use `tfds.features.text.Tokenizer` to split it into tokens.
# 3. Collect these tokens into a Python set, to remove duplicates.
# 4. Get the size of the vocabulary for later use.
# + colab={} colab_type="code" id="YkHtbGnDh6mg"
tokenizer = tfds.features.text.Tokenizer()
vocabulary_set = set()
for text_tensor, _ in all_labeled_data:
some_tokens = tokenizer.tokenize(text_tensor.numpy())
vocabulary_set.update(some_tokens)
vocab_size = len(vocabulary_set)
vocab_size
# + [markdown] colab_type="text" id="0W35VJqAh9zs"
# ### Encode examples
#
# Create an encoder by passing the `vocabulary_set` to `tfds.features.text.TokenTextEncoder`. The encoder's `encode` method takes in a string of text and returns a list of integers.
# + colab={} colab_type="code" id="gkxJIVAth6j0"
encoder = tfds.features.text.TokenTextEncoder(vocabulary_set)
# + [markdown] colab_type="text" id="v6S5Qyabi-vo"
# You can try this on a single line to see what the output looks like.
# + colab={} colab_type="code" id="jgxPZaxUuTbk"
example_text = next(iter(all_labeled_data))[0].numpy()
print(example_text)
# + colab={} colab_type="code" id="XoVpKR3qj5yb"
encoded_example = encoder.encode(example_text)
print(encoded_example)
# + [markdown] colab_type="text" id="p9qHM0v8k_Mg"
# Now run the encoder on the dataset by wrapping it in `tf.py_function` and passing that to the dataset's `map` method.
# + colab={} colab_type="code" id="HcIQ7LOTh6eT"
def encode(text_tensor, label):
encoded_text = encoder.encode(text_tensor.numpy())
return encoded_text, label
# + [markdown] colab_type="text" id="eES_Z1ia-Om-"
# You want to use `Dataset.map` to apply this function to each element of the dataset. `Dataset.map` runs in graph mode.
#
# * Graph tensors do not have a value.
# * In graph mode you can only use TensorFlow Ops and functions.
#
# So you can't `.map` this function directly: You need to wrap it in a `tf.py_function`. The `tf.py_function` will pass regular tensors (with a value and a `.numpy()` method to access it), to the wrapped python function.
# + colab={} colab_type="code" id="KmQVsAgJ-RM0"
def encode_map_fn(text, label):
# py_func doesn't set the shape of the returned tensors.
encoded_text, label = tf.py_function(encode,
inp=[text, label],
Tout=(tf.int64, tf.int64))
# `tf.data.Datasets` work best if all components have a shape set
# so set the shapes manually:
encoded_text.set_shape([None])
label.set_shape([])
return encoded_text, label
all_encoded_data = all_labeled_data.map(encode_map_fn)
# + [markdown] colab_type="text" id="_YZToSXSm0qr"
# ## Split the dataset into test and train batches
#
# Use `tf.data.Dataset.take` and `tf.data.Dataset.skip` to create a small test dataset and a larger training set.
#
# Before being passed into the model, the datasets need to be batched. Typically, the examples inside of a batch need to be the same size and shape. But, the examples in these datasets are not all the same size — each line of text had a different number of words. So use `tf.data.Dataset.padded_batch` (instead of `batch`) to pad the examples to the same size.
# + colab={} colab_type="code" id="r-rmbijQh6bf"
train_data = all_encoded_data.skip(TAKE_SIZE).shuffle(BUFFER_SIZE)
train_data = train_data.padded_batch(BATCH_SIZE)
test_data = all_encoded_data.take(TAKE_SIZE)
test_data = test_data.padded_batch(BATCH_SIZE)
# + [markdown] colab_type="text" id="Xdz7SVwmqi1l"
# Now, `test_data` and `train_data` are not collections of (`example, label`) pairs, but collections of batches. Each batch is a pair of (*many examples*, *many labels*) represented as arrays.
#
# To illustrate:
# + colab={} colab_type="code" id="kMslWfuwoqpB"
sample_text, sample_labels = next(iter(test_data))
sample_text[0], sample_labels[0]
# + [markdown] colab_type="text" id="UI4I6_Sa0vWu"
# Since we have introduced a new token encoding (the zero used for padding), the vocabulary size has increased by one.
# + colab={} colab_type="code" id="IlD1Lli91vuc"
vocab_size += 1
# + [markdown] colab_type="text" id="K8SUhGFNsmRi"
# ## Build the model
#
#
# + colab={} colab_type="code" id="QJgI1pow2YR9"
model = tf.keras.Sequential()
# + [markdown] colab_type="text" id="wi0iiKLTKdoF"
# The first layer converts integer representations to dense vector embeddings. See the [word embeddings tutorial](../text/word_embeddings.ipynb) or more details.
# + colab={} colab_type="code" id="DR6-ctbY638P"
model.add(tf.keras.layers.Embedding(vocab_size, 64))
# + [markdown] colab_type="text" id="_8OJOPohKh1q"
# The next layer is a [Long Short-Term Memory](http://colah.github.io/posts/2015-08-Understanding-LSTMs/) layer, which lets the model understand words in their context with other words. A bidirectional wrapper on the LSTM helps it to learn about the datapoints in relationship to the datapoints that came before it and after it.
# + colab={} colab_type="code" id="x6rnq6DN_WUs"
model.add(tf.keras.layers.Bidirectional(tf.keras.layers.LSTM(64)))
# + [markdown] colab_type="text" id="cdffbMr5LF1g"
# Finally we'll have a series of one or more densely connected layers, with the last one being the output layer. The output layer produces a probability for all the labels. The one with the highest probability is the models prediction of an example's label.
# + colab={} colab_type="code" id="QTEaNSnLCsv5"
# One or more dense layers.
# Edit the list in the `for` line to experiment with layer sizes.
for units in [64, 64]:
model.add(tf.keras.layers.Dense(units, activation='relu'))
# Output layer. The first argument is the number of labels.
model.add(tf.keras.layers.Dense(3))
# + [markdown] colab_type="text" id="zLHPU8q5DLi_"
# Finally, compile the model. For a softmax categorization model, use `sparse_categorical_crossentropy` as the loss function. You can try other optimizers, but `adam` is very common.
# + colab={} colab_type="code" id="pkTBUVO4h6Y5"
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
# + [markdown] colab_type="text" id="DM-HLo5NDhql"
# ## Train the model
#
# This model running on this data produces decent results (about 83%).
# + colab={} colab_type="code" id="aLtO33tNh6V8"
model.fit(train_data, epochs=3, validation_data=test_data)
# + colab={} colab_type="code" id="KTPCYf_Jh6TH"
eval_loss, eval_acc = model.evaluate(test_data)
print('\nEval loss: {:.3f}, Eval accuracy: {:.3f}'.format(eval_loss, eval_acc))
|
site/en/tutorials/load_data/text.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="sx9e_pXlCuti"
# %load_ext autoreload
# %autoreload 2
# + [markdown] colab_type="text" id="UMMut8UVCutt"
# # Experiments
# We'll go through learning feature embeddings using different loss functions on MNIST dataset. This is just for visualization purposes, thus we'll be using 2-dimensional embeddings which isn't the best choice in practice.
#
# For every experiment the same embedding network is used (32 conv 5x5 -> PReLU -> MaxPool 2x2 -> 64 conv 5x5 -> PReLU -> MaxPool 2x2 -> Dense 256 -> PReLU -> Dense 256 -> PReLU -> Dense 2) and we don't do any hyperparameter search.
# + [markdown] colab_type="text" id="BcmGBqXeCutw"
# # Prepare dataset
# We'll be working on MNIST dataset
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 119} colab_type="code" executionInfo={"elapsed": 3691, "status": "ok", "timestamp": 1528587668602, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="AQSHPH9P0BNx" outputId="023d99ab-56d0-4116-e5be-3827035a70dd"
import torch
from torchvision.datasets import FashionMNIST
from torchvision import transforms
mean, std = 0.28604059698879553, 0.35302424451492237
batch_size = 256
train_dataset = FashionMNIST('./data/FashionMNIST', train=True, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((mean,), (std,))
]))
test_dataset = FashionMNIST('./data/FashionMNIST', train=False, download=True,
transform=transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((mean,), (std,))
]))
cuda = torch.cuda.is_available()
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size, shuffle=True, **kwargs)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
n_classes = 10
# + [markdown] colab_type="text" id="TcZTFRnjCut3"
# ## Common setup
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="Dz2xh66UCut5"
import torch
from torch.optim import lr_scheduler
import torch.optim as optim
from torch.autograd import Variable
from trainer import fit
import numpy as np
cuda = torch.cuda.is_available()
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
fashion_mnist_classes = ['T-shirt/top', 'Trouser', 'Pullover', 'Dress', 'Coat',
'Sandal', 'Shirt', 'Sneaker', 'Bag', 'Ankle boot']
colors = ['#1f77b4', '#ff7f0e', '#2ca02c', '#d62728',
'#9467bd', '#8c564b', '#e377c2', '#7f7f7f',
'#bcbd22', '#17becf']
mnist_classes = fashion_mnist_classes
def plot_embeddings(embeddings, targets, xlim=None, ylim=None):
plt.figure(figsize=(10,10))
for i in range(10):
inds = np.where(targets==i)[0]
plt.scatter(embeddings[inds,0], embeddings[inds,1], alpha=0.5, color=colors[i])
if xlim:
plt.xlim(xlim[0], xlim[1])
if ylim:
plt.ylim(ylim[0], ylim[1])
plt.legend(mnist_classes)
def extract_embeddings(dataloader, model):
with torch.no_grad():
model.eval()
embeddings = np.zeros((len(dataloader.dataset), 2))
labels = np.zeros(len(dataloader.dataset))
k = 0
for images, target in dataloader:
if cuda:
images = images.cuda()
embeddings[k:k+len(images)] = model.get_embedding(images).data.cpu().numpy()
labels[k:k+len(images)] = target.numpy()
k += len(images)
return embeddings, labels
# + [markdown] colab_type="text" id="rxkRm-RO7Bji"
# # Baseline: Classification with softmax
# We'll train the model for classification and use outputs of penultimate layer as embeddings
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="y29hMdwo7Bjl"
# Set up data loaders
batch_size = 256
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
train_loader = torch.utils.data.DataLoader(train_dataset, batch_size=batch_size, shuffle=True, **kwargs)
test_loader = torch.utils.data.DataLoader(test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
# Set up the network and training parameters
from networks import EmbeddingNet, ClassificationNet
from metrics import AccumulatedAccuracyMetric
embedding_net = EmbeddingNet()
model = ClassificationNet(embedding_net, n_classes=n_classes)
if cuda:
model.cuda()
loss_fn = torch.nn.NLLLoss()
lr = 1e-2
optimizer = optim.Adam(model.parameters(), lr=lr)
scheduler = lr_scheduler.StepLR(optimizer, 8, gamma=0.1, last_epoch=-1)
n_epochs = 20
log_interval = 50
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 2397} colab_type="code" executionInfo={"elapsed": 438517, "status": "ok", "timestamp": 1528588135301, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="J5ntvNU77Bjo" outputId="bf6814a5-8cce-45c4-df08-0cb764dafefe"
fit(train_loader, test_loader, model, loss_fn, optimizer, scheduler, n_epochs, cuda, log_interval, metrics=[AccumulatedAccuracyMetric()])
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1167} colab_type="code" executionInfo={"elapsed": 23634, "status": "ok", "timestamp": 1528588158972, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="REAGu3f-7Bjt" outputId="7b20e0c5-cb5b-4b38-ddb1-78692c9c7852"
train_embeddings_baseline, train_labels_baseline = extract_embeddings(train_loader, model)
plot_embeddings(train_embeddings_baseline, train_labels_baseline)
val_embeddings_baseline, val_labels_baseline = extract_embeddings(test_loader, model)
plot_embeddings(val_embeddings_baseline, val_labels_baseline)
# + [markdown] colab_type="text" id="j9FhHE-tCuuc"
# # Siamese network
# We'll train a siamese network that takes a pair of images and trains the embeddings so that the distance between them is minimized if their from the same class or greater than some margin value if they represent different classes.
# We'll minimize a contrastive loss function*:
# $$L_{contrastive}(x_0, x_1, y) = \frac{1}{2} y \lVert f(x_0)-f(x_1)\rVert_2^2 + \frac{1}{2}(1-y)\{max(0, m-\lVert f(x_0)-f(x_1)\rVert_2)\}^2$$
#
# *<NAME>, <NAME>, <NAME>, [Dimensionality reduction by learning an invariant mapping](http://yann.lecun.com/exdb/publis/pdf/hadsell-chopra-lecun-06.pdf), CVPR 2006*
#
# ## Steps
# 1. Create a dataset returning pairs - **SiameseMNIST** class from *datasets.py*, wrapper for MNIST-like classes.
# 2. Define **embedding** *(mapping)* network $f(x)$ - **EmbeddingNet** from *networks.py*
# 3. Define **siamese** network processing pairs of inputs - **SiameseNet** wrapping *EmbeddingNet*
# 4. Train the network with **ContrastiveLoss** - *losses.py*
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="-qpIq-TzCuue"
# Set up data loaders
from datasets import SiameseMNIST
# Step 1
siamese_train_dataset = SiameseMNIST(train_dataset) # Returns pairs of images and target same/different
siamese_test_dataset = SiameseMNIST(test_dataset)
batch_size = 128
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
siamese_train_loader = torch.utils.data.DataLoader(siamese_train_dataset, batch_size=batch_size, shuffle=True, **kwargs)
siamese_test_loader = torch.utils.data.DataLoader(siamese_test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
# Set up the network and training parameters
from networks import EmbeddingNet, SiameseNet
from losses import ContrastiveLoss
# Step 2
embedding_net = EmbeddingNet()
# Step 3
model = SiameseNet(embedding_net)
if cuda:
model.cuda()
# Step 4
margin = 1.
loss_fn = ContrastiveLoss(margin)
lr = 1e-3
optimizer = optim.Adam(model.parameters(), lr=lr)
scheduler = lr_scheduler.StepLR(optimizer, 8, gamma=0.1, last_epoch=-1)
n_epochs = 20
log_interval = 500
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1037} colab_type="code" executionInfo={"elapsed": 930771, "status": "ok", "timestamp": 1528589092774, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="_IqiBATeCuuh" outputId="8110dc3e-337c-417b-8e88-ac0ceaa3337b"
fit(siamese_train_loader, siamese_test_loader, model, loss_fn, optimizer, scheduler, n_epochs, cuda, log_interval)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1167} colab_type="code" executionInfo={"elapsed": 22170, "status": "ok", "timestamp": 1528589115036, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="rBBBuPjiCuup" outputId="6f426277-9dda-48e6-9eef-e4c54349d33a"
train_embeddings_cl, train_labels_cl = extract_embeddings(train_loader, model)
plot_embeddings(train_embeddings_cl, train_labels_cl)
val_embeddings_cl, val_labels_cl = extract_embeddings(test_loader, model)
plot_embeddings(val_embeddings_cl, val_labels_cl)
# + [markdown] colab_type="text" id="MbKXy6yQCuuu"
# # Triplet network
# We'll train a triplet network, that takes an anchor, positive (same class as anchor) and negative (different class than anchor) examples. The objective is to learn embeddings such that the anchor is closer to the positive example than it is to the negative example by some margin value.
#
# 
# Source: [2] *Schroff, Florian, <NAME>, and <NAME>. [Facenet: A unified embedding for face recognition and clustering.](https://arxiv.org/abs/1503.03832) CVPR 2015.*
#
# **Triplet loss**: $L_{triplet}(x_a, x_p, x_n) = max(0, m + \lVert f(x_a)-f(x_p)\rVert_2^2 - \lVert f(x_a)-f(x_n)\rVert_2^2$\)
#
# ## Steps
# 1. Create a dataset returning triplets - **TripletMNIST** class from *datasets.py*, wrapper for MNIST-like classes
# 2. Define **embedding** *(mapping)* network $f(x)$ - **EmbeddingNet** from *networks.py*
# 3. Define **triplet** network processing triplets - **TripletNet** wrapping *EmbeddingNet*
# 4. Train the network with **TripletLoss** - *losses.py*
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="jv4DvFucCuuu"
# Set up data loaders
from datasets import TripletMNIST
triplet_train_dataset = TripletMNIST(train_dataset) # Returns triplets of images
triplet_test_dataset = TripletMNIST(test_dataset)
batch_size = 128
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
triplet_train_loader = torch.utils.data.DataLoader(triplet_train_dataset, batch_size=batch_size, shuffle=True, **kwargs)
triplet_test_loader = torch.utils.data.DataLoader(triplet_test_dataset, batch_size=batch_size, shuffle=False, **kwargs)
# Set up the network and training parameters
from networks import EmbeddingNet, TripletNet
from losses import TripletLoss
margin = 1.
embedding_net = EmbeddingNet()
model = TripletNet(embedding_net)
if cuda:
model.cuda()
loss_fn = TripletLoss(margin)
lr = 1e-3
optimizer = optim.Adam(model.parameters(), lr=lr)
scheduler = lr_scheduler.StepLR(optimizer, 8, gamma=0.1, last_epoch=-1)
n_epochs = 20
log_interval = 500
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1037} colab_type="code" executionInfo={"elapsed": 1305892, "status": "ok", "timestamp": 1528590422778, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="Dj9AoYpsCuuz" outputId="4f81c75f-1d16-4c9c-e05c-b26d5f39fa84"
fit(triplet_train_loader, triplet_test_loader, model, loss_fn, optimizer, scheduler, n_epochs, cuda, log_interval)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1167} colab_type="code" executionInfo={"elapsed": 22976, "status": "ok", "timestamp": 1528590445794, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="ysh4Ry7ZCuu_" outputId="2ea247e2-eb96-405a-aa5a-b3c0b565d2fd"
train_embeddings_tl, train_labels_tl = extract_embeddings(train_loader, model)
plot_embeddings(train_embeddings_tl, train_labels_tl)
val_embeddings_tl, val_labels_tl = extract_embeddings(test_loader, model)
plot_embeddings(val_embeddings_tl, val_labels_tl)
# + [markdown] colab_type="text" id="x7C9H1_nCuvJ"
# # Online pair/triplet selection - negative mining
# There are couple of problems with siamese and triplet networks.
# 1. The number of possible pairs/triplets grows **quadratically/cubically** with the number of examples. It's infeasible to process them all
# 2. We generate pairs/triplets randomly. As the training continues, more and more pairs/triplets are easy to deal with (their loss value is very small or even 0), preventing the network from training. We need to provide the network with **hard examples**.
# 3. Each image that is fed to the network is used only for computation of contrastive/triplet loss for only one pair/triplet. The computation is somewhat wasted; once the embedding is computed, it could be reused for many pairs/triplets.
#
# To deal with that efficiently, we'll feed a network with standard mini-batches as we did for classification. The loss function will be responsible for selection of hard pairs and triplets within mini-batch. In these case, if we feed the network with 16 images per 10 classes, we can process up to $159*160/2 = 12720$ pairs and $10*16*15/2*(9*16) = 172800$ triplets, compared to 80 pairs and 53 triplets in previous implementation.
#
# We can find some strategies on how to select triplets in [2] and [3] *<NAME>, <NAME>, <NAME>, [In Defense of the Triplet Loss for Person Re-Identification](https://arxiv.org/pdf/1703.07737), 2017*
# + [markdown] colab_type="text" id="k806qej9CuvL"
# ## Online pair selection
# ## Steps
# 1. Create **BalancedBatchSampler** - samples $N$ classes and $M$ samples *datasets.py*
# 2. Create data loaders with the batch sampler
# 3. Define **embedding** *(mapping)* network $f(x)$ - **EmbeddingNet** from *networks.py*
# 4. Define a **PairSelector** that takes embeddings and original labels and returns valid pairs within a minibatch
# 5. Define **OnlineContrastiveLoss** that will use a *PairSelector* and compute *ContrastiveLoss* on such pairs
# 6. Train the network!
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="goVi1_-PCuvL"
from datasets import BalancedBatchSampler
# We'll create mini batches by sampling labels that will be present in the mini batch and number of examples from each class
train_batch_sampler = BalancedBatchSampler(train_dataset.train_labels, n_classes=10, n_samples=25)
test_batch_sampler = BalancedBatchSampler(test_dataset.test_labels, n_classes=10, n_samples=25)
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
online_train_loader = torch.utils.data.DataLoader(train_dataset, batch_sampler=train_batch_sampler, **kwargs)
online_test_loader = torch.utils.data.DataLoader(test_dataset, batch_sampler=test_batch_sampler, **kwargs)
# Set up the network and training parameters
from networks import EmbeddingNet
from losses import OnlineContrastiveLoss
from utils import AllPositivePairSelector, HardNegativePairSelector # Strategies for selecting pairs within a minibatch
margin = 1.
embedding_net = EmbeddingNet()
model = embedding_net
if cuda:
model.cuda()
loss_fn = OnlineContrastiveLoss(margin, HardNegativePairSelector())
lr = 1e-3
optimizer = optim.Adam(model.parameters(), lr=lr)
scheduler = lr_scheduler.StepLR(optimizer, 8, gamma=0.1, last_epoch=-1)
n_epochs = 20
log_interval = 250
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1037} colab_type="code" executionInfo={"elapsed": 567133, "status": "ok", "timestamp": 1528591015945, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="SacMTqN6CuvO" outputId="87a2c414-a641-4cde-ac89-96e7c10cef02"
all_embeddings = fit(online_train_loader, online_test_loader, model, loss_fn, optimizer, scheduler, n_epochs, cuda, log_interval)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1167} colab_type="code" executionInfo={"elapsed": 15155, "status": "ok", "timestamp": 1528591031137, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="ZCvF0AlCCuvX" outputId="27de240c-eb68-4de0-b57b-f386ccb23ebb"
train_embeddings_ocl, train_labels_ocl = extract_embeddings(train_loader, model)
plot_embeddings(train_embeddings_ocl, train_labels_ocl)
val_embeddings_ocl, val_labels_ocl = extract_embeddings(test_loader, model)
plot_embeddings(val_embeddings_ocl, val_labels_ocl)
# + [markdown] colab_type="text" id="UgIGiMwICuvn"
# ## Online triplet selection
# ## Steps
# 1. Create **BalancedBatchSampler** - samples $N$ classes and $M$ samples *datasets.py*
# 2. Create data loaders with the batch sampler
# 3. Define **embedding** *(mapping)* network $f(x)$ - **EmbeddingNet** from *networks.py*
# 4. Define a **TripletSelector** that takes embeddings and original labels and returns valid triplets within a minibatch
# 5. Define **OnlineTripletLoss** that will use a *TripletSelector* and compute *TripletLoss* on such pairs
# 6. Train the network!
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="JzpYzMUuCuvp"
from datasets import BalancedBatchSampler
# We'll create mini batches by sampling labels that will be present in the mini batch and number of examples from each class
train_batch_sampler = BalancedBatchSampler(train_dataset.train_labels, n_classes=10, n_samples=25)
test_batch_sampler = BalancedBatchSampler(test_dataset.test_labels, n_classes=10, n_samples=25)
kwargs = {'num_workers': 1, 'pin_memory': True} if cuda else {}
online_train_loader = torch.utils.data.DataLoader(train_dataset, batch_sampler=train_batch_sampler, **kwargs)
online_test_loader = torch.utils.data.DataLoader(test_dataset, batch_sampler=test_batch_sampler, **kwargs)
# Set up the network and training parameters
from networks import EmbeddingNet
from losses import OnlineTripletLoss
from utils import AllTripletSelector,HardestNegativeTripletSelector, RandomNegativeTripletSelector, SemihardNegativeTripletSelector # Strategies for selecting triplets within a minibatch
from metrics import AverageNonzeroTripletsMetric
margin = 1.
embedding_net = EmbeddingNet()
model = embedding_net
if cuda:
model.cuda()
loss_fn = OnlineTripletLoss(margin, RandomNegativeTripletSelector(margin))
lr = 1e-3
optimizer = optim.Adam(model.parameters(), lr=lr, weight_decay=1e-4)
scheduler = lr_scheduler.StepLR(optimizer, 8, gamma=0.1, last_epoch=-1)
n_epochs = 20
log_interval = 150
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1377} colab_type="code" executionInfo={"elapsed": 2163515, "status": "ok", "timestamp": 1528593197388, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="W-bDxqVJCuvs" outputId="1ef0d7ce-6c6a-4dc0-e879-348e081f1bd3"
fit(online_train_loader, online_test_loader, model, loss_fn, optimizer, scheduler, n_epochs, cuda, log_interval, metrics=[AverageNonzeroTripletsMetric()])
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1167} colab_type="code" executionInfo={"elapsed": 15139, "status": "ok", "timestamp": 1528593212580, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="dNrbA2hCCuvw" outputId="e2b0d141-2b88-459a-af17-f4433b5564b9"
train_embeddings_otl, train_labels_otl = extract_embeddings(train_loader, model)
plot_embeddings(train_embeddings_otl, train_labels_otl)
val_embeddings_otl, val_labels_otl = extract_embeddings(test_loader, model)
plot_embeddings(val_embeddings_otl, val_labels_otl)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1167} colab_type="code" executionInfo={"elapsed": 3897, "status": "ok", "timestamp": 1528593217032, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="aDeV-K4o7Bk2" outputId="a4c8c20b-8d41-4c4d-d1a0-4258a4fa5b30"
display_emb_online, display_emb, display_label_online, display_label = train_embeddings_ocl, train_embeddings_cl, train_labels_ocl, train_labels_cl
# display_emb_online, display_emb, display_label_online, display_label = val_embeddings_ocl, val_embeddings_cl, val_labels_ocl, val_labels_cl
x_lim = (np.min(display_emb_online[:,0]), np.max(display_emb_online[:,0]))
y_lim = (np.min(display_emb_online[:,1]), np.max(display_emb_online[:,1]))
x_lim = (min(x_lim[0], np.min(display_emb[:,0])), max(x_lim[1], np.max(display_emb[:,0])))
y_lim = (min(y_lim[0], np.min(display_emb[:,1])), max(y_lim[1], np.max(display_emb[:,1])))
plot_embeddings(display_emb, display_label, x_lim, y_lim)
plot_embeddings(display_emb_online, display_label_online, x_lim, y_lim)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}, "base_uri": "https://localhost:8080/", "height": 1167} colab_type="code" executionInfo={"elapsed": 3858, "status": "ok", "timestamp": 1528593220933, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a/default-user=s128", "userId": "113360808941199578430"}, "user_tz": -120} id="tNY7NReM0zsq" outputId="eede3036-9fa4-4e78-c7bc-e53f991074b1"
display_emb_online, display_emb, display_label_online, display_label = train_embeddings_otl, train_embeddings_tl, train_labels_otl, train_labels_tl
# display_emb_online, display_emb, display_label_online, display_label = val_embeddings_otl, val_embeddings_tl, val_labels_otl, val_labels_tl
x_lim = (np.min(display_emb_online[:,0]), np.max(display_emb_online[:,0]))
y_lim = (np.min(display_emb_online[:,1]), np.max(display_emb_online[:,1]))
x_lim = (min(x_lim[0], np.min(display_emb[:,0])), max(x_lim[1], np.max(display_emb[:,0])))
y_lim = (min(y_lim[0], np.min(display_emb[:,1])), max(y_lim[1], np.max(display_emb[:,1])))
plot_embeddings(display_emb, display_label, x_lim, y_lim)
plot_embeddings(display_emb_online, display_label_online, x_lim, y_lim)
# + colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" id="9AOfKw7W7VwT"
|
siamese-triplet/Experiments_FashionMNIST.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python TA
# language: python
# name: ta-v2
# ---
# +
import pandas as pd
import numpy as np
from utils.data_helper import get_markable_dataframe, get_embedding_variables
from model_builders.antecedentless_classifier import AntecedentlessClassifierModelBuilder
from functools import reduce
from tensorflow.keras.preprocessing.sequence import pad_sequences
# +
embedding_indexes_file_path = 'helper_files/embedding/embedding_indexes.txt'
indexed_embedding_file_path = 'helper_files/embedding/indexed_embedding.txt'
word_vector, embedding_matrix, idx_by_word, word_by_idx = get_embedding_variables(embedding_indexes_file_path, indexed_embedding_file_path)
# +
data = get_markable_dataframe("data/training/markables.csv", word_vector, idx_by_word)
data.head()
# +
max_text_length = 10
max_prev_words_length = max(map(lambda x: len(x), data.all_previous_words))
data_text = pad_sequences(data.text, maxlen=max_text_length, padding='post')
data_all_previous_words = pad_sequences(data.all_previous_words.map(lambda seq: seq[(-1*max_prev_words_length):]), maxlen=max_prev_words_length, padding='pre')
data_syntactic = data[['is_pronoun', 'entity_type', 'is_proper_name', 'is_first_person']]
data_syntactic = np.array(list(map(lambda p: reduce(lambda x,y: x + y, [i if type(i) is list else [i] for i in p]), data_syntactic.values)))
label = np.vstack(data.is_antecedentless)
# -
# # Build Model
# ## Words + Context + Numeric
words_context_numeric_model_builder = AntecedentlessClassifierModelBuilder(
use_words_feature=True,
use_context_feature=True,
use_syntactic_feature=True,
embedding_matrix=embedding_matrix,
syntactic_features_num=data_syntactic.shape[1]
)
words_context_numeric_model = words_context_numeric_model_builder.create_model()
words_context_numeric_model.fit([data_text, data_all_previous_words, data_syntactic], label, epochs=5)
|
antecedentless classifier training.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# Using a neural network to predict stock prices, using only basic data
# +
# %matplotlib inline
from matplotlib import pyplot as plt
import datetime
import pandas_datareader.data as web
import pandas as pd
import numpy as np
import seaborn as sns
from sklearn.preprocessing import StandardScaler
from sklearn.cross_validation import train_test_split
from sklearn.metrics import r2_score
# increase default figure size for matplotlib
from pylab import rcParams
rcParams['figure.figsize'] = 20, 10
from collections import defaultdict
# -
start_date = datetime.datetime(2014,1,1)
end_date = datetime.datetime(2016,8,31)
symbol = "PETR4.SA"
df = web.DataReader(symbol, 'yahoo', start_date, end_date)
# +
# df.ix[:,'Adj Close'].plot()
# -
# first I have to add a signal, either up or down, to each row
df['Delta'] = (df['Adj Close'] / df['Adj Close'].shift(1))-1
df['Signal'] = df['Delta']>0
df = df.dropna()
# df.head()
# done. now I need to do backward filling, then I'm ready to predict
backward = 30
for column in ['Delta', 'Volume']:
for i in range(1,backward+1):
new_column = "{} -d{}".format(column, i)
for row in range(backward, df.shape[0]):
df.ix[row, new_column] = df.ix[row-i, column]
df = df.dropna()
len(df.columns)
# +
# deprecated: switch to 2 labels - buy or don't buy
# look back for features, ahead for labeling
forward = 5
# boundaries
soft = .05
for row in range(df.shape[0]-forward):
# first - construct forward
count_signals = 0
max_uptick = 0
min_downtick = 0
for i in range(1,forward+1):
# capture signals
count_signals += df.ix[row+i, 'Signal']
delta = (df.ix[row+i, 'Adj Close'] / df.ix[row, 'Adj Close'])-1
if delta > max_uptick:
max_uptick = delta
if delta < min_downtick:
min_downtick = delta
# convert to label
signals = count_signals*1.0/forward
# up
if max_uptick >= soft:
df.ix[row,'Label'] = 1
else:
df.ix[row,'Label'] = 0
# -
df['Label'].value_counts(normalize=True, sort=False)
df = df.dropna()
# now to classification
# it is far from a minimal necessary
# but I will see what values can I get
# start with a simple tree
X = df.drop('Label', axis=1)
y = df['Label']
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.2, random_state=42)
y_pred_random = np.random.random_integers(low=0, high=1, size=len(y_test))
# +
from sklearn.tree import DecisionTreeClassifier as Tree
from sklearn.ensemble import GradientBoostingClassifier as GBC
from sklearn.cross_validation import train_test_split
from sklearn.svm import SVC
from sklearn.naive_bayes import GaussianNB
from sklearn.neural_network import MLPClassifier as NN
from sklearn.neighbors import KNeighborsClassifier as kNN
from sklearn.metrics import confusion_matrix
from sklearn.metrics import accuracy_score
from sklearn.metrics import f1_score
import warnings
warnings.filterwarnings("ignore", category=DeprecationWarning)
# np.random.seed(33)
classifiers = [GBC(), Tree(), SVC(), GaussianNB(), NN(), kNN()]
print accuracy_score(y_test, y_pred_random), f1_score(y_test, y_pred_random)
for clf in classifiers:
clf.fit(X_train, y_train)
y_pred = clf.predict(X_test)
print clf.__class__
print accuracy_score(y_test, y_pred), f1_score(y_test, y_pred)
# +
# kNN and Gradient Boosting are good performers
# what happens if I follow this straight
# I will buy it. sell it when it values .05
# I will be right 60% of the time
# so I lose on 40%, win on 60%
# when I win, I win 0.05. when I lose, I loose 0.05
# it can't variate more than .05 down as well
# +
# deprecated: switch to 2 labels - buy or don't buy
# look back for features, ahead for labeling
forward = 10
# boundaries
soft = df['Delta'].std()
hard = soft*2
for row in range(df.shape[0]-forward):
# first - construct forward
count_signals = 0
max_uptick = 0
min_downtick = 0
for i in range(1,forward+1):
# capture signals
count_signals += df.ix[row+i, 'Signal']
delta = (df.ix[row+i, 'Adj Close'] / df.ix[row, 'Adj Close'])-1
if delta > max_uptick:
max_uptick = delta
if delta < min_downtick:
min_downtick = delta
# convert to label
signals = count_signals*1.0/forward
# test
if row%123 == 0:
print "count_signals: {}, signals: {:.1f}, max_uptick: {:.3f}, min_downtick: {:.3f}".format(
count_signals, signals, max_uptick, min_downtick)
# up
if signals >= .8 and max_uptick >= hard:
df.ix[row,'Label'] = 5
elif (signals >= .7 and max_uptick >= soft) or max_uptick >= hard or signals >=.8:
df.ix[row,'Label'] = 4
# down
elif signals <= .2 and min_downtick <= -hard:
df.ix[row,'Label'] = 1
elif (signals <= .3 and min_downtick <= -soft) or min_downtick <= -hard or signals <=.2:
df.ix[row,'Label'] = 2
# neutral
else:
df.ix[row,'Label'] = 3
|
.ipynb_checkpoints/Predictorv3-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.7.7 64-bit
# name: python3
# ---
# + [markdown] colab_type="text" id="view-in-github"
# <a href="https://colab.research.google.com/github/gowtham1997/indicTrans-1/blob/main/indicTrans_Finetuning.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="rE4MO-8bDtwD" outputId="e54447b4-2b04-44c4-96a2-a79e7ed014ae"
# create a seperate folder to store everything
# !mkdir finetuning
# %cd finetuning
# + colab={"base_uri": "https://localhost:8080/"} id="-2Rs6_WkD_gF" outputId="95d19041-0e73-406c-a3c2-c7bddbfda916"
# clone the repo for running finetuning
# !git clone https://github.com/AI4Bharat/indicTrans.git
# %cd indicTrans
# clone requirements repositories
# !git clone https://github.com/anoopkunchukuttan/indic_nlp_library.git
# !git clone https://github.com/anoopkunchukuttan/indic_nlp_resources.git
# !git clone https://github.com/rsennrich/subword-nmt.git
# %cd ..
# + colab={"base_uri": "https://localhost:8080/"} id="duwTvJ9xEBJ1" outputId="98445af3-041d-415d-97f3-a322939260e4"
# ! sudo apt install tree
# Install the necessary libraries
# !pip install sacremoses pandas mock sacrebleu tensorboardX pyarrow indic-nlp-library
# Install fairseq from source
# !git clone https://github.com/pytorch/fairseq.git
# %cd fairseq
# # !git checkout da9eaba12d82b9bfc1442f0e2c6fc1b895f4d35d
# !pip install --editable ./
# %cd ..
# + colab={"base_uri": "https://localhost:8080/"} id="oD2EHQdqEH70" outputId="0b988dde-9da3-487c-a393-510fbcae92f3"
# download the indictrans model
# downloading the en-indic model
# this will contain:
# en-indic/
# ├── final_bin # contains fairseq dictionaries (we will use this to binarize the new finetuning data)
# │ ├── dict.SRC.txt
# │ └── dict.TGT.txt
# ├── model # contains model checkpoint(s)
# │ └── checkpoint_best.pt
# └── vocab # contains bpes for src and tgt (since we train seperate vocabularies) generated with subword_nmt (we will use this bpes to convert finetuning data to subwords)
# ├── bpe_codes.32k.SRC
# ├── bpe_codes.32k.TGT
# ├── vocab.SRC
# └── vocab.TGT
# !wget https://storage.googleapis.com/samanantar-public/V0.2/models/indic-en.zip
# !unzip indic-en.zip
# if you want to finetune indic-en models, use the link below
# # !wget https://storage.googleapis.com/samanantar-public/V0.2/models/en-indic.zip
# # !unzip en-indic.zip
# if you want to finetune indic-indic models, use the link below
# # !wget https://storage.googleapis.com/samanantar-public/V0.3/models/m2m.zip
# # !unzip m2m.zip
# + colab={"base_uri": "https://localhost:8080/"} id="lj7XNBuwE0OV" outputId="98b3a156-c205-4f1b-de79-f1d640555349"
# In this example, we will finetuning on cvit-pib corpus which is part of the WAT2021 training dataset.
# Lets first download the full wat2021 training data (cvit-pib is a part of this big training set)
# ***Note***: See the next section to mine for mining indic to indic data from english centric WAT data. This dataset can be used to finetune indic2indic model
# !wget http://lotus.kuee.kyoto-u.ac.jp/WAT/indic-multilingual/indic_wat_2021.tar.gz
# !tar -xzvf indic_wat_2021.tar.gz
# all train sets will now be in wat2021/train
# !mv finalrepo wat2021
# + colab={"base_uri": "https://localhost:8080/"} id="BSoZDR3fHpUk" outputId="11bd057b-d1b0-45b8-feac-85b3e900104e"
# this cell is for mining indic to indic data from a english centric corpus. This data can then be used to our finetune indic2indic model
# Mining Indic to Indic pairs from english centric corpus
# The `extract_non_english_pairs` in `scripts/extract_non_english_pairs.py` can be used to mine indic to indic pairs from english centric corpus.
# As described in the paper (section 2.5) , we use a very strict deduplication criterion to avoid the creation of very similar parallel sentences.
# For example, if an en sentence is aligned to M hi sentences and N ta sentences, then we would get MN hi-ta pairs. However, these pairs would be very similar and not contribute much to the training process.
# Hence, we retain only 1 randomly chosen pair out of these MN pairs.
# !mkdir wat2021-indic2indic
from indicTrans.scripts.extract_non_english_pairs import extract_non_english_pairs
"""
extract_non_english_pairs(indir, outdir, LANGS)
Extracts non-english pair parallel corpora
indir: contains english centric data in the following form:
- directory named en-xx for language xx
- each directory contains a train.en and train.xx
outdir: output directory to store mined data for each pair.
One directory is created for each pair.
LANGS: list of languages in the corpus (other than English).
The language codes must correspond to the ones used in the
files and directories in indir. Prefarably, sort the languages
in this list in alphabetic order. outdir will contain data for xx-yy,
but not for yy-xx, so it will be convenient to have this list in sorted order.
"""
# here we are using three langs to mine bn-hi, hi-gu and gu-bn pairs from wat2021/cvit-pib en-X data
# you should see the following files after running the code below
# wat2021-indic2indic
# ├── bn-gu
# │ ├── train.bn
# │ └── train.gu
# ├── bn-hi
# │ ├── train.bn
# │ └── train.hi
# └── hi-gu
# ├── train.gu
# └── train.hi
# NOTE: Make sure to dedup the output text files and remove any overlaps with test sets before finetuning
# Both of the above are implemented in scripts/remove_train_devtest_overlaps.py -> remove_train_devtest_overlaps(train_dir, devtest_dir, many2many=True)
extract_non_english_pairs('wat2021/train/cvit-pib', 'wat2021-indic2indic', ['bn', 'hi', 'gu'])
# + colab={"base_uri": "https://localhost:8080/"} id="ys_QURP3Sx7G" outputId="d41f5baa-e700-4e07-93cd-b23b08122dc5"
# wat2021
# ├── dev # contains Wat2021 dev data
# │ ├── dev.bn
# │ ├── dev.en
# │ ├── dev.gu
# │ ├── dev.hi
# │ ├── dev.kn
# │ ├── dev.ml
# │ ├── dev.mr
# │ ├── dev.or
# │ ├── dev.pa
# │ ├── dev.ta
# │ └── dev.te
# ├── README
# ├── test # contains Wat2021 test data
# │ ├── test.bn
# │ ├── test.en
# │ ├── test.gu
# │ ├── test.hi
# │ ├── test.kn
# │ ├── test.ml
# │ ├── test.mr
# │ ├── test.or
# │ ├── test.pa
# │ ├── test.ta
# │ └── test.te
# └── train # contains WAT2021 train data which has lot of corpuses (alt, bible, Jw300, etc)
# ├── alt/
# ├── bibleuedin/
# ├── iitb/
# ├── jw/
# ├── mtenglish2odia/
# ├── nlpc/
# ├── odiencorp/
# ├── opensubtitles/
# ├── pmi/
# ├── tanzil/
# ├── ted2020/
# ├── ufal/
# ├── urst/
# ├── wikimatrix/
# ├── wikititles/
# └── cvit-pib
# ├── en-bn # within a train corpus folder the files are arranged in {src_lang}-{tgt_lang}/train.{src_lang}, train.{tgt_lang}
# │ ├── train.bn
# │ └── train.en
# ├── en-gu
# │ ├── train.en
# │ └── train.gu
# ├── en-hi
# │ ├── train.en
# │ └── train.hi
# ├── en-ml
# │ ├── train.en
# │ └── train.ml
# ├── en-mr
# │ ├── train.en
# │ └── train.mr
# ├── en-or
# │ ├── train.en
# │ └── train.or
# ├── en-pa
# │ ├── train.en
# │ └── train.pa
# ├── en-ta
# │ ├── train.en
# │ └── train.ta
# └── en-te
# ├── train.en
# └── train.te
# instead of using all the data for this example, we will mainly use the cvit-pib corpus from wat2021 train set
# for dev and test set, we will use the dev and test provided by wat2021
# In case, you want to finetune on all these corpuses, you would need to merge all the training data into one folder and remove duplicate train sentence pairs.
# To do this, refer to this gist: https://gist.github.com/gowtham1997/2524f8e9559cff586d1f935e621fc598
# # copy everything to a dataset folder
# !mkdir -p dataset/train
# ! cp -r wat2021/train/cvit-pib/* dataset/train
# ! cp -r wat2021/dev dataset
# ! cp -r wat2021/test dataset
# lets cd to indicTrans
# %cd indicTrans
# + colab={"base_uri": "https://localhost:8080/"} id="8yPTbM_clKfI" outputId="d4459da6-3e0b-45c8-f291-d6761e536284"
# %%shell
exp_dir=../dataset
src_lang=en
tgt_lang=indic
# change this to indic-en, if you have downloaded the indic-en dir or m2m if you have downloaded the indic2indic model
download_dir=../en-indic
train_data_dir=$exp_dir/train
dev_data_dir=$exp_dir/dev
test_data_dir=$exp_dir/test
# echo $exp_dir
# + colab={"base_uri": "https://localhost:8080/"} id="NhwUXyYVXrOY" outputId="9ddb06dd-3fcc-4d4c-a4ec-131a9f4ea220"
# all the data preparation happens in this cell
# %%shell
exp_dir=../dataset
src_lang=en
tgt_lang=indic
# change this to indic-en, if you have downloaded the indic-en dir or m2m if you have downloaded the indic2indic model
download_dir=../en-indic
train_data_dir=$exp_dir/train
dev_data_dir=$exp_dir/dev
test_data_dir=$exp_dir/test
# echo "Running experiment ${exp_dir} on ${src_lang} to ${tgt_lang}"
train_processed_dir=$exp_dir/data
devtest_processed_dir=$exp_dir/data
out_data_dir=$exp_dir/final_bin
# mkdir -p $train_processed_dir
# mkdir -p $devtest_processed_dir
# mkdir -p $out_data_dir
# indic languages.
# cvit-pib corpus does not have as (assamese) and kn (kannada), hence its not part of this list
langs=(bn hi gu ml mr or pa ta te)
for lang in ${langs[@]};do
if [ $src_lang == en ]; then
tgt_lang=$lang
else
src_lang=$lang
fi
train_norm_dir=$exp_dir/norm/$src_lang-$tgt_lang
devtest_norm_dir=$exp_dir/norm/$src_lang-$tgt_lang
mkdir -p $train_norm_dir
mkdir -p $devtest_norm_dir
# preprocessing pretokenizes the input (we use moses tokenizer for en and indicnlp lib for indic languages)
# after pretokenization, we use indicnlp to transliterate all the indic data to devnagiri script
# train preprocessing
train_infname_src=$train_data_dir/en-${lang}/train.$src_lang
train_infname_tgt=$train_data_dir/en-${lang}/train.$tgt_lang
train_outfname_src=$train_norm_dir/train.$src_lang
train_outfname_tgt=$train_norm_dir/train.$tgt_lang
echo "Applying normalization and script conversion for train $lang"
input_size=`python scripts/preprocess_translate.py $train_infname_src $train_outfname_src $src_lang true`
input_size=`python scripts/preprocess_translate.py $train_infname_tgt $train_outfname_tgt $tgt_lang true`
echo "Number of sentences in train $lang: $input_size"
# dev preprocessing
dev_infname_src=$dev_data_dir/dev.$src_lang
dev_infname_tgt=$dev_data_dir/dev.$tgt_lang
dev_outfname_src=$devtest_norm_dir/dev.$src_lang
dev_outfname_tgt=$devtest_norm_dir/dev.$tgt_lang
echo "Applying normalization and script conversion for dev $lang"
input_size=`python scripts/preprocess_translate.py $dev_infname_src $dev_outfname_src $src_lang true`
input_size=`python scripts/preprocess_translate.py $dev_infname_tgt $dev_outfname_tgt $tgt_lang true`
echo "Number of sentences in dev $lang: $input_size"
# test preprocessing
test_infname_src=$test_data_dir/test.$src_lang
test_infname_tgt=$test_data_dir/test.$tgt_lang
test_outfname_src=$devtest_norm_dir/test.$src_lang
test_outfname_tgt=$devtest_norm_dir/test.$tgt_lang
echo "Applying normalization and script conversion for test $lang"
input_size=`python scripts/preprocess_translate.py $test_infname_src $test_outfname_src $src_lang true`
input_size=`python scripts/preprocess_translate.py $test_infname_tgt $test_outfname_tgt $tgt_lang true`
echo "Number of sentences in test $lang: $input_size"
done
# Now that we have preprocessed all the data, we can now merge these different text files into one
# ie. for en-as, we have train.en and corresponding train.as, similarly for en-bn, we have train.en and corresponding train.bn
# now we will concatenate all this into en-X where train.SRC will have all the en (src) training data and train.TGT will have all the concatenated indic lang data
python scripts/concat_joint_data.py $exp_dir/norm $exp_dir/data $src_lang $tgt_lang 'train'
python scripts/concat_joint_data.py $exp_dir/norm $exp_dir/data $src_lang $tgt_lang 'dev'
python scripts/concat_joint_data.py $exp_dir/norm $exp_dir/data $src_lang $tgt_lang 'test'
# use the vocab from downloaded dir
# cp -r $download_dir/vocab $exp_dir
# echo "Applying bpe to the new finetuning data"
bash apply_single_bpe_traindevtest_notag.sh $exp_dir
# mkdir -p $exp_dir/final
# We also add special tags to indicate the source and target language in the inputs
# Eg: to translate a sentence from english to hindi , the input would be __src__en__ __tgt__hi__ <en bpe tokens>
# echo "Adding language tags"
python scripts/add_joint_tags_translate.py $exp_dir 'train'
python scripts/add_joint_tags_translate.py $exp_dir 'dev'
python scripts/add_joint_tags_translate.py $exp_dir 'test'
data_dir=$exp_dir/final
out_data_dir=$exp_dir/final_bin
# rm -rf $out_data_dir
# binarizing the new data (train, dev and test) using dictionary from the download dir
num_workers=`python -c "import multiprocessing; print(multiprocessing.cpu_count())"`
data_dir=$exp_dir/final
out_data_dir=$exp_dir/final_bin
# # rm -rf $out_data_dir
# echo "Binarizing data. This will take some time depending on the size of finetuning data"
fairseq-preprocess --source-lang SRC --target-lang TGT \
--trainpref $data_dir/train --validpref $data_dir/dev --testpref $data_dir/test \
--destdir $out_data_dir --workers $num_workers \
--srcdict $download_dir/final_bin/dict.SRC.txt --tgtdict $download_dir/final_bin/dict.TGT.txt --thresholdtgt 5 --thresholdsrc 5
# + colab={"base_uri": "https://localhost:8080/"} id="iz6tzbe2tcs7" outputId="6705e2d6-b5cb-4810-c833-6a1370d3fce4"
# Finetuning the model
# pls refer to fairseq documentaion to know more about each of these options (https://fairseq.readthedocs.io/en/latest/command_line_tools.html)
# some notable args:
# --max-update=1000 -> for this example, to demonstrate how to finetune we are only training for 1000 steps. You should increase this when finetuning
# --arch=transformer_4x -> we use a custom transformer model and name it transformer_4x (4 times the parameter size of transformer base)
# --user_dir -> we define the custom transformer arch in model_configs folder and pass it as an argument to user_dir for fairseq to register this architechture
# --lr -> learning rate. From our limited experiments, we find that lower learning rates like 3e-5 works best for finetuning.
# --restore-file -> reload the pretrained checkpoint and start training from here (change this path for indic-en. Currently its is set to en-indic)
# --reset-* -> reset and not use lr scheduler, dataloader, optimizer etc of the older checkpoint
# --max_tokns -> this is max tokens per batch
!( fairseq-train ../dataset/final_bin \
--max-source-positions=210 \
--max-target-positions=210 \
--max-update=1000 \
--save-interval=1 \
--arch=transformer_4x \
--criterion=label_smoothed_cross_entropy \
--source-lang=SRC \
--lr-scheduler=inverse_sqrt \
--target-lang=TGT \
--label-smoothing=0.1 \
--optimizer adam \
--adam-betas "(0.9, 0.98)" \
--clip-norm 1.0 \
--warmup-init-lr 1e-07 \
--warmup-updates 4000 \
--dropout 0.2 \
--tensorboard-logdir ../dataset/tensorboard-wandb \
--save-dir ../dataset/model \
--keep-last-epochs 5 \
--patience 5 \
--skip-invalid-size-inputs-valid-test \
--fp16 \
--user-dir model_configs \
--update-freq=2 \
--distributed-world-size 1 \
--max-tokens 256 \
--lr 3e-5 \
--restore-file ../en-indic/model/checkpoint_best.pt \
--reset-lr-scheduler \
--reset-meters \
--reset-dataloader \
--reset-optimizer)
# + id="tpPsT1e7vuO9"
# To test the models after training, you can use joint_translate.sh
# joint_translate takes src_file, output_fname, src_lang, tgt_lang, model_folder as inputs
# src_file -> input text file to be translated
# output_fname -> name of the output file (will get created) containing the model predictions
# src_lang -> source lang code of the input text ( in this case we are using en-indic model and hence src_lang would be 'en')
# tgt_lang -> target lang code of the input text ( tgt lang for en-indic model would be any of the 11 indic langs we trained on:
# as, bn, hi, gu, kn, ml, mr, or, pa, ta, te)
# supported languages are:
# as - assamese, bn - bengali, gu - gujarathi, hi - hindi, kn - kannada,
# ml - malayalam, mr - marathi, or - oriya, pa - punjabi, ta - tamil, te - telugu
# model_dir -> the directory containing the model and the vocab files
# Note: if the translation is taking a lot of time, please tune the buffer_size and batch_size parameter for fairseq-interactive defined inside this joint_translate script
# here we are translating the english sentences to hindi
# !bash joint_translate.sh $exp_dir/test/test.en en_hi_outputs.txt 'en' 'hi' $exp_dir
# + id="bPqneByPxilN"
# to compute bleu scores for the predicitions with a reference file, use the following command
# arguments:
# pred_fname: file that contains model predictions
# ref_fname: file that contains references
# src_lang and tgt_lang : the source and target language
bash compute_bleu.sh en_hi_outputs.txt $exp_dir/test/test.hi 'en' 'hi'
|
indicTrans_Finetuning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import gym
import tensorflow as tf
import numpy as np
env = gym.make("CartPole-v0")
obs_dim = env.observation_space.shape[0]
n_acts = env.action_space.n
print("obs = {}, n_acts = {}".format(obs_dim, n_acts))
obs = env.reset()
obs
def mpl(x, sizes, activation = tf.tanh, output_activation=None):
for size in sizes:
x = tf.layers.dense(x,units = size, activation = activation)
return tf.layers.dense(x, units = sizes[-1], activation = output_activation)
#first layer of NN
obs_ph = tf.placeholder(shape=(None, obs_dim), dtype=tf.float32)
#creating NN
logit = mpl(obs_ph, sizes = [32,64]+[n_acts])
#array([[0.01685937, 0.00622761]], dtype=float32)
max_action = tf.multinomial(logits=logit,num_samples=1)
#array([[0]])
action = tf.squeeze(max_action, axis=1)
#array([0])
# +
#loss
weights_ph = tf.placeholder(shape=(None,), dtype = tf.float32)
act_ph = tf.placeholder(shape=(None,), dtype = tf.int32)
#one_hot taken actions
action_masks = tf.one_hot(act_ph, n_acts)
#array([[0., 1.]...], dtype=float32)
#reward that will get from this point
baseline_ph = tf.placeholder(shape=(None,), dtype = tf.float32)
# -
baseline = baseline_ph - 1
#logprob(a|s)
log_probs = tf.reduce_sum(action_masks * tf.nn.log_softmax(logit), axis=1)
#array([-0.6952652,...], dtype=float32)
loss = -tf.reduce_mean(weights_ph* log_probs* baseline)
#9.812662
#optimizer
train_op = tf.train.AdamOptimizer(learning_rate = 1e-2).minimize(loss)
#loss minimizes :D
#variables for initializing and using
sess = tf.InteractiveSession()
sess.run(tf.global_variables_initializer())
max_action = sess.run(action, {obs_ph: obs.reshape(1,-1)})[0]
#get element from array 0 or 1
def reward_to_go(rews):
n = len(rews)
rtgs = np.zeros_like(rews)
for i in reversed(range(n)):
rtgs[i] = rews[i] + (rtgs[i+1] if i+1 < n else 0)
#returns [200. 199. ... 1.]
return rtgs
#LMAO I DONT KNOW HOW IT WORKS, BUT IT WORKS
def reward_after(w):
n = len(w)
list2 = w.copy()
for i in range(n):
prev_el = w[i]
su = 0
for j in reversed(range(i)):
if prev_el >= w[j]:
break
su+=1
prev_el = w[j]
list2[i]=su if su != 0 else 1
return list2
#w[i]+=j WORK
#w[i]=200-w[i] NOT WORKING
# +
a = []
b = []
for i in reversed(range(200)):
a.append(i)
a = np.array(a)
a = a.reshape(len(a))
for i in reversed(range(100)):
b.append(i)
b = np.array(b)
b = b.reshape(len(b))
ab = np.concatenate((a,b),axis=0)
print("ab-conc;:::\n",ab)
reward_after(ab)
# -
def train_one_epoch(batch_size = 5000):
batch_obs = []
batch_acts = []
batch_rets = []
batch_lens = []
batch_weights = [] #[200.0, 199.0, ... 1.0, 200.0, ... 1.0]
obs = env.reset()
done = False
ep_rews = []
rendering_epoch = True
while True:
if rendering_epoch == True:
env.render()
batch_obs.append(obs.copy())
act = sess.run(action, {obs_ph: obs.reshape(1,-1)})[0]
obs, reward, done, _ = env.step(act)
batch_acts.append(act)
ep_rews.append(reward)
if done:
rendering_epoch = False
#recording everything!
ep_ret, ep_len = sum(ep_rews), len(ep_rews)
batch_rets.append(ep_ret)
batch_lens.append(ep_len)
# the weight for each logprob(a_t|s_t) is reward-to-go from t
batch_weights += list(reward_to_go(ep_rews))
#reset vatiables
obs, done, ep_rews = env.reset(), False, []
if len(batch_obs)>batch_size:
break
if np.mean(batch_lens)<200:
baseline = reward_after(batch_weights)
batch_loss, _ = sess.run([loss, train_op], {
obs_ph: batch_obs,
act_ph: batch_acts,
weights_ph: batch_weights,
baseline_ph: baseline
})
else:
batch_loss = 0
return batch_loss, batch_rets, batch_lens
for i in range(500):
batch_loss, batch_rets, batch_lens = train_one_epoch()
print("#%i, batch_loss: %.3f, batch_rets: %.3f, batch_lens: %.3f" \
%(i, batch_loss, np.mean(batch_rets), np.mean(batch_lens)))
if batch_loss == 0:
break
def play_episode():
rewards = 0
obs = env.reset()
while True:
env.render()
act = sess.run(action, {obs_ph: obs.reshape(1,-1)})[0]
obs, reward, is_done, _ = env.step(act)
rewards += reward
#if is_done or rewards >200:
# break
return rewards
print(play_episode())
env.close()
sum = 8
for i in range(8):
print(i)
sum+=i
sum
|
Reward to get Baseline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Training Set Expansion
# ## Get Datasets
# +
from sklearn.datasets import fetch_openml
mnist = fetch_openml('mnist_784', version=1)
# print shape of the data
X, y = mnist['data'], mnist['target']
print("X shape = ", X.shape)
print("y shape = ", y.shape)
# +
# make Test sets and training sets
X_train, X_test, y_train, y_test = X[:60000], X[60000:], y[:60000], y[60000:]
# Shuffle data
import numpy as np
shuffle_index = np.random.permutation(60000)
X_train, y_train = X_train[shuffle_index], y_train[shuffle_index]
# -
# ## Expand Datasets
# +
from scipy.ndimage.interpolation import shift
def shift_image(image, dx, dy):
image = image.reshape((28, 28))
shifted_image = shift(image, [dy, dx], cval=0, mode="constant")
return shifted_image.reshape([-1])
# +
# test if this methods works
# %matplotlib inline
import matplotlib
import matplotlib.pyplot as plt
image = X_train[1000]
shifted_image_left = shift_image(image, 0, -1)
shifted_image_right = shift_image(image, 0, 1)
shifted_image_up = shift_image(image, 1, 0)
shifted_image_down = shift_image(image, -1, 0)
# plot datasets
plt.figure()
plt.subplot(221)
plt.title("Shifted up", fontsize=14)
plt.imshow(shifted_image_up.reshape(28, 28), interpolation="nearest", cmap="Greys")
plt.subplot(222)
plt.title("Shifted down", fontsize=14)
plt.imshow(shifted_image_down.reshape(28, 28), interpolation="nearest", cmap="Greys")
plt.show()
plt.subplot(223)
plt.title("Shifted right", fontsize=14)
plt.imshow(shifted_image_right.reshape(28, 28), interpolation="nearest", cmap="Greys")
plt.subplot(224)
plt.title("Shifted left", fontsize=14)
plt.imshow(shifted_image_left.reshape(28, 28), interpolation="nearest", cmap="Greys")
plt.show()
# +
X_train_expanded = [image for image in X_train]
y_train_expanded = [image for image in y_train]
for dx, dy in ((1, 0), (-1, 0), (0, 1), (0, -1)):
for image, label in zip(X_train, y_train):
X_train_expanded.append(shift_image(image, dx, dy))
y_train_expanded.append(label)
X_train_expanded = np.array(X_train_expanded)
y_train_expanded = np.array(y_train_expanded)
# -
# shuffle train set
random_idx = np.random.permutation(len(X_train_expanded))
X_train_expanded = X_train_expanded[random_idx]
y_train_expanded = y_train_expanded[random_idx]
# ## Train & Test model (Scaled Random Forest Classification)
# +
from sklearn.ensemble import RandomForestClassifier
from sklearn.preprocessing import StandardScaler
# Scale Training set & Test set
scaler = StandardScaler()
X_train_scale = scaler.fit_transform(X_train_expanded)
X_test_scale = scaler.fit_transform(X_test)
# -
clf = RandomForestClassifier()
clf.fit(X_train_scale, y_train_expanded)
# +
# Cross Validation
from sklearn.model_selection import cross_val_score
cross_val_score(clf, X_train_scale, y_train_expanded, cv=3, scoring="accuracy")
# +
# Test Score
prediction = clf.predict(X_test_scale)
result = (prediction == y_test).mean()
print(result)
# -
|
MNIST/.ipynb_checkpoints/MNIST_TSE-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## NumPy
# +
# # !pip install numpy
# -
import numpy as np
# ### np.array()
l = [1, 2, 3, 4, 5]
npa = np.array(l)
type(l), type(npa)
npa.shape
l.append('string')
npa = np.array(l)
npa
l1 = [1, 4, 9, 16]
l
l + l1
np.array(l + l1)
l + [20]
l = [[1, 2], [3, 4], [3, 4]]
npa2d = np.array(l)
npa2d
npa2d.shape
npa2d = np.array(l, dtype = 'float')
npa2d
npa2d.astype('int')
npa2d = npa2d.astype('str')
npa2d[0][0] + str(1)
# ### `np.nan` and `np.inf`
np.nan == np.nan
None == None
np.inf == np.inf
npa2d = npa2d.astype('float')
npa2d = npa2d.astype('int')
npa2d
npa2d = npa2d.astype('float')
npa2d[0][0] = np.nan
npa2d[1][1] = np.inf
npa2d
np.isnan(npa2d)
np.isinf(npa2d)
flag = np.isnan(npa2d) | np.isinf(npa2d)
flag
npa2d[flag]
npa2d[flag] = 0
npa2d
# ### Statistical Operations
# +
# mean, var, std, med
# -
npa2d
npa2d.mean()
npa2d.min()
npa2d.max()
npa2d.var()
npa2d.std()
np.median(npa2d)
# ### Shape(), Reshape(), Ravel()
npa2d
npa2d.shape
npa2d.reshape(2, 3)
npa2d.reshape(1, 6)
npa2d.reshape(1, 6).ravel()
npa2d.flatten()
# ### sequence, repetitions, and random numbers
# #### arange(), linspace(), random(), zeros(), and ones()
np.arange(1, 5, dtype = 'float')
np.arange(1, 20, 3)
np.linspace(1, 20, 5)
np.logspace(1, 50, 10)
np.zeros([1, 5])
np.zeros([1, 2, 3])
np.ones([1, 2])
np.tile([1, 2], 3)
np.repeat([1, 2], 3)
np.random.rand(3, 3)
np.random.randint(0, 100, [3, 3])
np.random.seed(1)
np.random.randint(0, 100, [3, 3])
np.random.randint(0, 100, [3, 3])
np.unique(npa2d)
npa2d
np.unique(npa2d, return_counts= True)
# ### Where()
arr = np.array([8,94,8,56,1,3,4,5,7])
arr
np.where(arr>10)
index = np.where(arr>10)
arr[index]
index = np.where(arr<10)
arr[index]
index = np.where(arr==10)
arr[index]
arr>10
arr[arr>10]
arr.max()
arr.argmax()
arr.argmin()
arr[arr.argmin()]
# ### File Read and Write
npa2d
np.savetxt('npa2d.csv', npa2d, delimiter=',')
np.loadtxt('npa2d.csv', delimiter=',')
np.save('data.npy', npa2d)
np.load('data.npy')
np.savez('both.npz', arr, npa2d)
d = np.load('both.npz')
d.files
d[d.files[1]]
# ### Concatenate and Sorting
npa2d
np.concatenate([npa2d, npa2d], axis = 0)
np.concatenate([npa2d, npa2d], axis = 1)
np.concatenate([npa2d, arr], axis = 1)
np.vstack([npa2d, npa2d])
npa = np.hstack([npa2d, npa2d])
npa
npa.sort()
npa
npa.sort(axis = 1)
npa
# ### Working with Dates
d = np.datetime64('2020-12-01 23:34:23')
d
d + 10
oneday = np.timedelta64(1, 'D')
oneday
d + oneday
oneminute = np.timedelta64(1, 'm')
d + oneminute
|
Numpy.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# reload packages
# %load_ext autoreload
# %autoreload 2
# ### Choose GPU (this may not be needed on your computer)
# %env CUDA_DEVICE_ORDER=PCI_BUS_ID
# %env CUDA_VISIBLE_DEVICES=0
import tensorflow as tf
gpu_devices = tf.config.experimental.list_physical_devices('GPU')
if len(gpu_devices)>0:
tf.config.experimental.set_memory_growth(gpu_devices[0], True)
print(gpu_devices)
# ### load packages
from tfumap.umap import tfUMAP
import tensorflow as tf
import numpy as np
import matplotlib.pyplot as plt
from tqdm.autonotebook import tqdm
import umap
import pandas as pd
# ### Load dataset
dataset = 'cassins_dtw'
dims = (32,32,1)
from tfumap.paths import ensure_dir, MODEL_DIR, DATA_DIR
# +
syllable_df = pd.read_pickle(DATA_DIR/'cassins'/ 'cassins.pickle')
#syllable_df= syllable_df[:1000]
syllable_df[:3]
top_labels = (
pd.DataFrame(
{i: [np.sum(syllable_df.labels.values == i)] for i in syllable_df.labels.unique()}
)
.T.sort_values(by=0, ascending=False)[:20]
.T
)
top_labels
syllable_df = syllable_df[syllable_df.labels.isin(top_labels.columns)]
syllable_df[:3]
syllable_df = syllable_df.reset_index()
syllable_df['subset'] = 'train'
syllable_df.loc[:1000, 'subset'] = 'valid'
syllable_df.loc[1000:1999, 'subset'] = 'test'
#syllable_df.loc[:100, 'subset'] = 'valid'
#syllable_df.loc[100:199, 'subset'] = 'test'
specs = np.array(list(syllable_df.spectrogram.values))
specs = np.array([np.concatenate([np.zeros((32,1)), i], axis=1) for i in tqdm(specs)])
specs.shape
syllable_df['spectrogram'] = syllable_df['spectrogram'].astype('object')
syllable_df['spectrogram'] = list(specs)
np.shape(syllable_df['spectrogram'].values[0])
len(syllable_df)
Y_train = np.array(list(syllable_df.labels.values[syllable_df.subset == 'train']))
Y_valid = np.array(list(syllable_df.labels.values[syllable_df.subset == 'valid']))
Y_test = np.array(list(syllable_df.labels.values[syllable_df.subset == 'test']))
X_train = np.array(list(syllable_df.spectrogram.values[syllable_df.subset == 'train'])) #/ 255.
X_valid = np.array(list(syllable_df.spectrogram.values[syllable_df.subset == 'valid']))# / 255.
X_test = np.array(list(syllable_df.spectrogram.values[syllable_df.subset == 'test'])) #/ 255.
X_train_flat = X_train.reshape((len(X_train), np.product(np.shape(X_train)[1:])))
X_test_flat = X_test.reshape((len(X_test), np.product(np.shape(X_test)[1:])))
from sklearn.preprocessing import OrdinalEncoder
enc = OrdinalEncoder()
Y_train = enc.fit_transform([[i] for i in Y_train]).astype('int').flatten()
Y_test = enc.fit_transform([[i] for i in Y_test]).astype('int').flatten()
plt.matshow(X_train[10])
# -
# ### Create model and train
# +
from tensorflow.keras.layers import (
Conv2D,
Reshape,
Bidirectional,
Dense,
RepeatVector,
TimeDistributed,
LSTM
)
n_components=2
#shape_final = (8,2,128)
encoder = tf.keras.Sequential([
tf.keras.layers.InputLayer(input_shape=dims),
Conv2D(
filters=32, kernel_size=3, strides=(2, 2), activation=tf.nn.leaky_relu, padding="same"
),
Conv2D(
filters=64, kernel_size=3, strides=(2, 2), activation=tf.nn.leaky_relu, padding="same"
),
Conv2D(
filters=128, kernel_size=3, strides=(2, 1), activation=tf.nn.leaky_relu, padding="same"
),
Conv2D(
filters=128, kernel_size=3, strides=(2, 1), activation=tf.nn.leaky_relu, padding="same"
),
Reshape(target_shape=(8, 2*128)),
Bidirectional(LSTM(units=100, activation="relu")),
Dense(units=512),
Dense(units=n_components),
])
# -
batch_size = 5000
from tfumap.paths import ensure_dir, MODEL_DIR, DATA_DIR
from tfumap.parametric_tsne import compute_joint_probabilities, tsne_loss
save_loc = DATA_DIR/ 'parametric_tsne'/ dataset / 'P.npy'
if save_loc.exists():
P = np.load(save_loc)
else:
P = compute_joint_probabilities(X_train_flat, batch_size=batch_size, perplexity=30, verbose=2)
ensure_dir(save_loc)
np.save(save_loc, P)
X_train_subset = X_train[:np.product(P.shape[:2])]
# Joint probabilities of data
Y_train_tsne = P.reshape(X_train_subset.shape[0], -1)
X_train.shape[0]
opt = tf.keras.optimizers.Adam(lr=0.01)
n_components = 2
encoder = tf.keras.Sequential()
encoder.add(tf.keras.layers.InputLayer(input_shape=dims))
encoder.add(tf.keras.layers.Flatten())
encoder.add(tf.keras.layers.Dense(units=100, activation="relu"))
encoder.add(tf.keras.layers.Dense(units=100, activation="relu"))
encoder.add(tf.keras.layers.Dense(units=100, activation="relu"))
encoder.add(
tf.keras.layers.Dense(units=n_components, name="z")
)
encoder.compile(loss=tsne_loss(d=n_components, batch_size=batch_size), optimizer=opt)
X_train_subset = np.reshape(X_train_subset, ([len(X_train_subset)]+ list(dims)))
X_test = np.reshape(X_test, ([len(X_test)]+ list(dims)))
# because shuffle == False, the same batches are used each time...
history = encoder.fit(X_train_subset, Y_train_tsne, batch_size=batch_size, shuffle=False, nb_epoch=1000)
# ### get z for training and test
X_train = np.reshape(X_train, ([len(X_train)]+ list(dims)))
z = encoder.predict(X_train)
z_test = encoder.predict(X_test)
# ### Test plot
fig, axs = plt.subplots(ncols = 2, figsize=(10, 5))
axs[0].scatter(z[:, 0], z[:, 1], s=0.1, alpha=0.5, c=Y_train[:len(z)], cmap=plt.cm.tab10)
axs[1].scatter(z_test[:, 0], z_test[:, 1], s=1, alpha=0.5, c=Y_test, cmap=plt.cm.tab10)
# ### Save models + projections
import os
output_dir = MODEL_DIR/'projections'/ dataset / 'parametric-tsne'
encoder.save(os.path.join(output_dir, "encoder"))
np.save(output_dir / 'z.npy', z)
np.save(output_dir / 'z_test.npy', z_test)
# ### compute metrics
# #### silhouette
from tfumap.silhouette import silhouette_score_block
ss, sil_samp = silhouette_score_block(z, Y_train, n_jobs = -1)
ss
ss_test, sil_samp_test = silhouette_score_block(z_test, Y_test, n_jobs = -1)
ss_test
fig, axs = plt.subplots(ncols = 2, figsize=(10, 5))
axs[0].scatter(z[:, 0], z[:, 1], s=0.1, alpha=0.5, c=sil_samp, cmap=plt.cm.viridis)
axs[1].scatter(z_test[:, 0], z_test[:, 1], s=1, alpha=0.5, c=sil_samp_test, cmap=plt.cm.viridis)
# #### KNN
from sklearn.neighbors import KNeighborsClassifier
neigh5 = KNeighborsClassifier(n_neighbors=5)
neigh5.fit(z, Y_train)
score_5nn = neigh5.score(z_test, Y_test)
score_5nn
neigh1 = KNeighborsClassifier(n_neighbors=1)
neigh1.fit(z, Y_train)
score_1nn = neigh1.score(z_test, Y_test)
score_1nn
# #### Trustworthiness
from sklearn.manifold import trustworthiness
tw = trustworthiness(X_train_flat[:10000], z[:10000])
tw_test = trustworthiness(X_test_flat[:10000], z_test[:10000])
tw, tw_test
# #### save output metrics
metrics_df = pd.DataFrame(
columns=[
"dataset",
"class_",
"dim",
"trustworthiness",
"silhouette_score",
"silhouette_samples",
]
)
metrics_df.loc[len(metrics_df)] = [dataset, 'parametric-tsne', n_components, tw, ss, sil_samp]
metrics_df
save_loc = DATA_DIR / 'projection_metrics' / 'train' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
metrics_df.to_pickle(save_loc)
metrics_df_test = pd.DataFrame(
columns=[
"dataset",
"class_",
"dim",
"trustworthiness",
"silhouette_score",
"silhouette_samples",
]
)
metrics_df_test.loc[len(metrics_df)] = [dataset, 'parametric-tsne', n_components, tw_test, ss_test, sil_samp_test]
metrics_df_test
save_loc = DATA_DIR / 'projection_metrics' / 'test' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
metrics_df.to_pickle(save_loc)
nn_acc_df = pd.DataFrame(columns = ["method_","dimensions","dataset","1NN_acc","5NN_acc"])
nn_acc_df.loc[len(nn_acc_df)] = ['parametric-tsne', n_components, dataset, score_1nn, score_5nn]
nn_acc_df
save_loc = DATA_DIR / 'knn_classifier' / str(n_components) / (dataset + '.pickle')
ensure_dir(save_loc)
nn_acc_df.to_pickle(save_loc)
|
notebooks/dataset-projections/cassins/cassins-parametric-tsne.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="4hqMglFDu3tv"
# #**Pandas**
#
# It's a library that is used for big data analysis and the name Pandas comes from "Python Data Analysis Library" so first import pandas
# + id="yg5VX28Au1sS"
import pandas as pd
# + [markdown] id="WddZT_rdve5V"
# ##Series
#
# Firstly, we need to talk about some of the building blocks in Pandas and the first one discussed is Series
#
# Series are like arrays, 1 dimensional data that normally represents a column in a dataset
# + colab={"base_uri": "https://localhost:8080/"} id="O95-SxzEzWhI" outputId="1125e96c-a71b-4651-b5b5-2add8198d272"
#Making a basic series
#this is just like np.array([some array])
a = pd.Series([1,2,3],dtype='int8')
print(a)
print(a[0])
# + [markdown] id="aK7KDHUUz2hZ"
# As seen above, just like with normal arrays the data above or the series above has indices which by default start at 0, just like arrays or lists.
#
# However the index for the Series can be customized and we can specify that with the index = parameter in pd.Series as such...
# + colab={"base_uri": "https://localhost:8080/"} id="2GyxwB130Ojs" outputId="98bd658f-94aa-4643-cf80-fa699012cab9"
a = pd.Series([1,45,73,20],dtype='int8',index=['a','b','c','d'],name="my first")
print(a)
# + [markdown] id="5w9nd5qP0b4Y"
# As seen above the index parameter is another list that MUST be of the same size as the Series, but each individual index can be the same or different.
#
# Now we can access the values as if we were accessing a dictionary!!
#
# ---
#
#
# + colab={"base_uri": "https://localhost:8080/"} id="MOYPuE9f0x45" outputId="3fc82c62-3547-4956-f3bb-edc497a7fd1f"
print(a[0])
##Or print the specific index name to get the value
print(a['b'])
print(f'I am printing the name of the array a as: {a.name}')
# + [markdown] id="sL-oJRx-02Vv"
# But dont worry, we can still access the values using the classic numerical indices and we can access the name of the series with the: name method
# + [markdown] id="SFNfaLys1jTv"
# ##Parameters
#
# * name = 'Name of array given'
# * dtype = 'int8' - size and type stored
# * index = [list of index names]
# * copy = boolean, by default false
# * data - can be a list or a dictionray with the keys being the indices and the values, the value at each index
# + colab={"base_uri": "https://localhost:8080/"} id="9DssstIE2KlM" outputId="f0a74408-acc4-4b99-83a9-a44507d16d8e"
a = {'red':3,'blue':5,'green':9}
a = pd.Series(a)
print(a)
# + colab={"base_uri": "https://localhost:8080/"} id="_yAxeRVJ2bHl" outputId="9a0ae0cb-70b2-4b2f-d34e-87f7220f17fd"
#We could limit the numbers of rows by setting
#indices that purposely exclude certain rows
a = pd.Series(a,index=['red'])
print(a)
# + [markdown] id="CYba43Hx2ufi"
# ##DataFrames
#
# DataFrames are literally like tables of data, except that they are made from dictionaries.
# + colab={"base_uri": "https://localhost:8080/"} id="d_q0nqaP2tsv" outputId="b5a08054-b509-4209-fdfb-a5e2d36c6372"
#courtesy of W3Schools
data = {
"calories": [420, 380, 390],
"duration": [50, 40, 45]
}
dframe = pd.DataFrame(data,index=[1,2,3])
print(dframe)
# + [markdown] id="k6PaBrhG3LmW"
# As seen above the 2-D dataframe is printed and the column heading are the dictionary keys and the values are the data in the table. As usual, since indices are not specified. Each row is represented as a number starting at 0
# + [markdown] id="R24tPr4q3fP2"
# ##Parameters
#
# * index = [list of indices]
# * dtype = 'int32' - by default
#
# Some others but aren't too practical
# + [markdown] id="pzINb0fNM0DS"
# ###Column Prints
#
# We can print out specific columns by saying:
#
# df['some column'] with the print function
# + [markdown] id="PotQ5WZB4tys"
# ##Loading CSV data
#
# So we might want to actually work with some data so we have to read the data in and turn it into a good format so lets try that out with the red_csv method
# + id="vEJHaGkE45Nb"
try:
dataframe = pd.read_csv('data.csv') #this will read data from a CSV file
dataframe.to_string() #this will print the entire dataset
except Exception:
print("I/O Exception for sure")
# + [markdown] id="Ol1g0jB45mxE"
# ##Loading JSON data
# + [markdown] id="W2_GTPXo5tMU"
# Sometimes data is actually found in a json file instead so we can use the same technique like before but with the json files
#
# I would like to remind the reader that we can actually use a dictionary of dictionaries to use as data since JSOn files are actually python dictionaries in disguise
# + colab={"base_uri": "https://localhost:8080/"} id="IW0dzpts6Lz5" outputId="de465b8c-e029-4b7c-8b67-6118be3abaf7"
try:
dataframe = pd.read_csv('data.json') #this will read data from a json file
dataframe.to_string() #this will print the entire dataset
except Exception:
print("I/O Exception for sure")
data = {
"Duration":{
"0":60,
"1":60,
"2":60,
"3":45,
"4":45,
"5":60
},
"Pulse":{
"0":110,
"1":117,
"2":103,
"3":109,
"4":117,
"5":102
},
"Maxpulse":{
"0":130,
"1":145,
"2":135,
"3":175,
"4":148,
"5":127
},
"Calories":{
"0":409,
"1":479,
"2":340,
"3":282,
"4":406,
"5":300
}
}
a = pd.DataFrame(data)
#print(a.to_latex()) ##does it latex style
print(a.to_string())
# + [markdown] id="eUsoIYKr7ir5"
# ##Viewing Data
#
# There are many ways to view the whole dataset or pieces of it but we can use the to_string() method or the head(rows) method
#
# data.head(#rows to view) - view n rows or by default the first 5 rows
# + [markdown] id="qVlSD6_5-2nR"
# ###Head method
# + colab={"base_uri": "https://localhost:8080/", "height": 234} id="QqgvSh0c78xh" outputId="30a1e62e-9cfd-40e1-e7ec-9b3fbf07355d"
import numpy
d = pd.DataFrame(a)
d.head(6) #whole set or a n number of rows
#if you exceed the total number of rows it will print the whole thing and ignore the input extra
# + [markdown] id="9xPj2Ekt-68n"
# ###Tail Method
# + [markdown] id="r2VG4yo88-Yz"
# We can view the first n rows with d.head(5)
#
# We can view the LAST n rows with d.tail()
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="i-2xvxfa9HCs" outputId="c8e45ffd-b390-494e-f83d-cbcb7eef36dd"
d.tail(3)
# + [markdown] id="Ppm3rFF1--ye"
# ###Count Method
# + colab={"base_uri": "https://localhost:8080/"} id="NKDh3X8O-uBF" outputId="bf4d4565-ea74-4173-a321-e35b594ab020"
d.count() #basically we count the number of rows for all columns
# + [markdown] id="I_atybFe_PtZ"
# ###Info on Data
#
# Try the .info() method on any created DataFrame
# + [markdown] id="3s90cjGw_cdc"
# ##Cleaning Data
#
# So now we have to learn to clean our data because data is never clean or perfect when it is first given to us and a solid chunk of the job for a data scientist is the clean up of the data.
#
# Some of the ways data comes in dirty is:
#
# * empty cells
# * data is formatted incorrectly
# * Wrong data
# * Duplicates
# + [markdown] id="-VSvhTyTG1ub"
# Now I will explain how to deal with the 4 types of situations in order
#
# Firstly, lets import this data to play with it
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="VW1-qgdJI2s8" outputId="0f02eb84-77ed-4ad2-86c9-a90d07d4cd27"
df = pd.read_csv('soccerteam.csv')
df.head(15)
# + [markdown] id="f0LmQ1twG9T_"
# ###Empty Cells
#
# To deal with empty cells we are looking for rows that dont have values in all column names and normally empty values look like: NaN
#
# We can use a specific function that looks for NaN values:
#
# datafr.dropna()
# + [markdown] id="iIVL5IW-MI8W"
# ####Dropna method
# + [markdown] id="3hfhqNjUHX4O"
# Parameters:
#
# inplace = boolean value, by default it is false so the method returns the new dataframe modified but if we dont want that and change the original datafram directly we can say:
#
# inplace = true
# + colab={"base_uri": "https://localhost:8080/"} id="80_nfZovHktQ" outputId="8d613380-c958-458d-9f79-0cedef995058"
df_copy = df.dropna(inplace=False)
df_copy
# + [markdown] id="Z1LuXvAvJ9Vb"
# For now I set the implace to false so that we get a new dataframe with the NaN values all dropped.
# + [markdown] id="WsMhcWaiMMzO"
# ####Fillna Method
# + [markdown] id="qKkfoozeKIZK"
# We can instead fill in the values with a number with the fillna method:
#
# fillna(0,inplace=true)
#
# Lets try that out for fun with a copy frame:
# + colab={"base_uri": "https://localhost:8080/"} id="vtxriK9QKjOZ" outputId="495d6b93-6f44-4876-9d44-d5bde3e3fe8d"
df_copy2 = df.fillna(0.0,inplace=False)
df_copy2
# + [markdown] id="UbSsVUzcLs5y"
# Now lets actually fill in the values
# + id="8hCL7bqRN4RG"
# + colab={"base_uri": "https://localhost:8080/"} id="iUJGir6zLsZa" outputId="8cde1f0e-b7e8-4b78-e062-404d4ddf6220"
df.fillna(0.0,inplace=True)
df
# + [markdown] id="eZYKrxQ0MQOC"
# ####Fillna Column-spec
# + [markdown] id="2yWAT112L79I"
# We can also replace a specific column like the score column
#
# df['score'].fillna(122,inplace=True)
# + [markdown] id="vDM7qZfOMWfy"
# ####Fill in with Mean
# + colab={"base_uri": "https://localhost:8080/"} id="_nWYQcITMcjT" outputId="1f075171-b00a-4c40-eef8-26ee6a439236"
df2 = pd.read_csv('soccerteam.csv')
#df2['Score'] I can print just the score
mean = df2['Score'].mean()
df2['Score'].fillna(mean,inplace=True)
print(df2)
# + [markdown] id="h2aCfx16OUHY"
# ####Fill in with Median
#
# med = df2['score'].median()
# df2['Score'].fillna(median)
# + [markdown] id="-7tvdFI5OiRD"
# Fill in with Mode
#
#
# mode = df['Score'].mode()
# + [markdown] id="HxQX5ibuJf7y"
# ###Wrong Format
# + [markdown] id="F2cLowTEL2xN"
# So we can do two things: Let us covert some of the bad values to good ones with a pd.to_?(the column) or we can also drop the values if the converstion fails
# + id="Lg37UvY1Pi63"
#df2['Division'] = pd.to_numeric(df2['Division'])
# + [markdown] id="G-iTfWYbQA4y"
# ###Replacing Values
#
# Lets say we want to replace the row value for index 5 of the Score column with 10 instead of 12, we just need to make that change:
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="D_U6Dhz6QdyW" outputId="eac92f1e-e2d6-42ed-b595-184a8d90e9d3"
df2.loc[5,'Score'] = 10
df2
#if the elements in a certain row of a specific
#column are smaller than n we can replace with something else
#with a loop
# + [markdown] id="SuXspyVSRw3b"
# We also change the value of certain elements in a row if the values are less that a certain value with a for loop that will touch each index in a list of indices given by dataframe.index
# + [markdown] id="-qVmTQhwS6OJ"
# ####Changing Rows
# + colab={"base_uri": "https://localhost:8080/", "height": 511} id="K0Q5udLMRCrE" outputId="d38cf3d2-8c86-45e2-949a-5819b38984f2"
for index in df2.index:
if df2.loc[index,'Score'] < 2:
df2.loc[index,'Score'] = 100
df2
# + [markdown] id="zI7YGeftR7GS"
# ####Removing Rows
#
#
# We can also just remove the row if the condition is true
# + colab={"base_uri": "https://localhost:8080/", "height": 480} id="eVL6P7nLSCc6" outputId="81a0a6a0-454b-4ef7-8287-df03e71bad29"
for index in df2.index:
if df2.loc[index,'Score'] == 100:
df2.drop(index,inplace = True)
df2
# + [markdown] id="JTyQmWuES_e5"
# ###Removing Duplicates
# + [markdown] id="Ka2nYjR2TD6Z"
# We can also remove duplicate valeus directly, but first we might want to detect them first right?
#
# print(df2.duplicated())
#
# The method returns true for every method that is duplicated
#
# ---
#
#
# + colab={"base_uri": "https://localhost:8080/", "height": 702} id="FQ0NRH4WTaCp" outputId="285d2e77-7c9f-4899-b5bb-18b9e45efd02"
print(df2.duplicated())
df2.drop_duplicates(inplace=True)
df2
# + [markdown] id="LzpfrcfiUT5d"
# ##Correlation
#
# We can find the correlation between all rows of data, ignoring the ones that have text or non numeric data in them with the corr method
# + colab={"base_uri": "https://localhost:8080/", "height": 110} id="zPkzFbg0Uh-Y" outputId="70a85a72-5b5f-4494-cbf5-479bb794aa57"
df2.corr()
# + [markdown] id="Ag1dN4PWU-Ml"
# ##Scatterplot
#
# Pandas also has a scatter plot method for plots so we can plot all columns with the
#
# df.plot()
# + [markdown] id="UaeIivP0VJW5"
# ###Parameters
#
# df.plot(kind=, x = 'some column', y='some other column')
#
# Leave all blank and the whole dataset gets printed!
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="c272R3c-WDGV" outputId="cf56c75f-d066-468c-f9fc-ac58719b7092"
df = pd.read_csv('data.csv')
#df - WE print the first 5 and last 5 rows
df.plot()
# + [markdown] id="30lh8bXmXOrY"
# We can also plot specific columns as part of the x,y axis
# + colab={"base_uri": "https://localhost:8080/", "height": 296} id="loNOdT9SWL__" outputId="56223365-049d-43d1-872f-ebd3436aa169"
df.plot(kind ='scatter',x ='Pulse',y ='Calories')
# + [markdown] id="Gxbjz3SDXZ36"
# ###Histograms
#
# We can even plot histograms of a specific column such as the following for Duration field
# + colab={"base_uri": "https://localhost:8080/", "height": 282} id="UwjfkJ6SXiNq" outputId="d241407f-eccf-4b83-deb8-35a54da36bff"
df['Duration'].plot(kind='hist')
# + [markdown] id="hl3-lf4MPR4H"
# ##Statistics
#
# So we are finally at the part where we calculate common statistical parameter for use in modeling so we can start by getting the mean and mode and median in the set
# + [markdown] id="uXdpe6p3PheQ"
# ###Mean,Median,Mode
# + id="-cm1rBHDPl7l"
import pandas as pd
import numpy as np
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="L0pde6klPzz_" outputId="9e9756eb-1703-4064-ac03-854c22c31b40"
df = pd.read_csv('data.csv')
df2 = pd.read_csv('soccerteam.csv')
df
# + colab={"base_uri": "https://localhost:8080/"} id="cHnnZM5SRKlt" outputId="d3755cf7-afe6-4d31-d385-53174d2f1887"
mean = df['Duration'].mean()
print(mean)
#We can print all the column names
print(df.columns)
mode = df['Duration'].mode()
print(mode)
print(f'The mode of the duration column is {mode}')
median = df['Pulse'].median()
print(f'The mode of the duration column is {median}')
print("--------------Printing column statistics---------")
for column in df.columns:
median = df[column].median()
print(f'The mode of the {column} column is {median}')
# + [markdown] id="56MPx5qUTZp6"
# ###Describe Method
#
# We can use this method when we want to desribe all the data or a portion of the columns in terms of the count, mean, std, min, max and the quartiles
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="Rj2BZJGNTww5" outputId="f696e8b4-66d2-41fc-adeb-aaf167bfb121"
#We can test all columns
df.describe()
# + colab={"base_uri": "https://localhost:8080/", "height": 295} id="wfDBqtIJT1g6" outputId="8662d0f6-74b0-4120-9382-8b557a457307"
#We can also test a specific column or columns in a list format
df[['Duration','Pulse']].describe()
# + [markdown] id="LYRBy0zuUH8N"
# ###Aggregate Method
#
# We can use this one when we dont want all the stats in the describe but only a couple and just like before we can use a list with the specific stats we are looking for and a dictionary where the keys are the column names we want
#
# + colab={"base_uri": "https://localhost:8080/", "height": 142} id="CAyv_bpjUdrT" outputId="f4e7b51b-0225-4a07-80a5-5d0f809e403d"
#Here the agg takes as an argument a dictionary made up of the column
#names along with sample statistics we want to find
df.agg(
{
'Duration': ['max','min','mean'],
'Maxpulse': ['max','min','mean']
}
)
# + [markdown] id="pFcMqewAVYnG"
# ###Cumulative Sum
#
# We can get the whole dataset but counting the values before it, almost like a fibonacci sequence
# + colab={"base_uri": "https://localhost:8080/", "height": 417} id="b3WwVoPsViqz" outputId="82cd738c-290e-427e-e870-67c93125cd05"
#We can get the whole dataset with its cum sum
df.cumsum()
#Or we can get a specific column
#df['Calories'].cumsum()
|
Pandas.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Comunicados de Política Monetaria
# ## Parte 2.
#
# En este notebook analizaremos por medio de expresiones regulares los anunciones de las decisiones de política monetaria de Banxico.
import os
import requests
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from tqdm.notebook import tqdm
from unidecode import unidecode
from textract import process
from multiprocessing import pool
# %config InlineBackend.figure_format = "retina"
plt.rcParams["figure.figsize"] = (12, 4)
comunicados = pd.read_pickle("comunicados-banxico.pkl")
comunicados.head()
# Creamos una nueva carpeta llamada 'comunicados' si esta no existes
wrkdir = "comunicados"
if not os.path.exists(wrkdir):
os.makedirs(wrkdir)
# Descarga cada uno de los comunicados de Banxico y guárdalos como .pdf dentro de la carpeta `comunicados`. Para cada archivo, guarda el comunicado con formato `yyyymmdd.pdf`
def download_statements(statements, workdir):
for date, vals in tqdm(statements.iterrows()):
filename = f"{date.strftime('%Y%m%d')}.pdf"
filename = os.path.join(workdir, filename)
url = vals["url"]
r = requests.get(url)
with open(filename, "wb") as f:
f.write(r.content)
if len(os.listdir(wrkdir)) == 0:
download_statements(comunicados, wrkdir)
else:
print("Statements downloaded...")
files_minutas = os.listdir(wrkdir)
len(files_minutas)
from sklearn.model_selection import train_test_split
from sklearn.metrics import precision_score, recall_score, confusion_matrix
from nltk.tokenize import RegexpTokenizer
from nltk.corpus import stopwords
from collections import defaultdict
void_words = stopwords.words("spanish")
tokenizer = RegexpTokenizer("(?:\w+|[0-9]*\.[0-9]%)+")
# +
def lee_minuta(path):
encoding = "iso-8859-1"
return process(path, encoding=encoding).decode(encoding)
def bag_words(path, stopwords):
minuta = lee_minuta(path)
minuta = unidecode(minuta.lower())
words = [word for word in tokenizer.tokenize(minuta) if word not in stopwords]
return words
def parse_minuta(path, stopwords):
words = bag_words(path, stopwords)
return pd.Series(words).value_counts()
# -
def categorize_minutas(comunicados, working_dir=".", stop_words=[], elements=None):
elements = slice(None) if elements is None else elements
comunicados = comunicados.loc[elements]
minutas_info = defaultdict(list)
category_count = defaultdict(int)
for release_date, info in comunicados.iterrows():
release_date_file = release_date.strftime("%Y%m%d.pdf")
path = os.path.join(working_dir, release_date_file)
words = bag_words(path, stop_words)
category = info["category"]
minutas_info[category].extend(words)
category_count[category] += 1
return minutas_info, category_count
train = comunicados.assign(category=comunicados.category.apply(map_key))
train, test = train_test_split(train, test_size=0.2, random_state=314)
train = train.sort_index()
# +
npartitions = 6
delta = (comunicados.index[-1] - comunicados.index[0]) / (npartitions - 1)
delta = delta.days
init_date = comunicados.index[0]
time_slices = [slice(init_date + pd.Timedelta(days=i * delta), init_date + pd.Timedelta(days=(i + 1) * delta)) for i in range(npartitions)]
time_slices
# +
def part(slices):
return categorize_minutas(train, wrkdir, void_words, slices)
pool = Pool(processes=npartitions)
res = []
for element in tqdm(pool.imap_unordered(part, time_slices), total=npartitions):
res.append(element)
pool.close()
# -
def map_key(decision):
if decision == "disminuye" or decision == "incrementa":
return "cambio"
else:
return decision
minutas_info = defaultdict(list)
category_count = defaultdict(int)
for info, cat in res:
for key, vals in cat.items():
category_count[key] += vals
for key, vals in info.items():
minutas_info[key].extend(vals)
categories = category_count.keys()
categories
priors = pd.DataFrame.from_dict(category_count, orient="index").T
priors = priors / priors.values.sum()
priors
log_priors = np.log(priors)
minutas_df = pd.concat([pd.Series(minutas_info[cat]).value_counts() for cat in categories],
axis=1, sort=True, keys=categories)
minutas_df["mantiene"].dropna().sort_values(ascending=False)
# Removing most repeated word among all clases
topw = minutas_df.assign(total_count=minutas_df.sum(axis=1)).sort_values("total_count", ascending=False)
topw = topw.head(20).index
minutas_df.loc[topw] = np.nan
nb = minutas_df.fillna(0) + 1
nb = nb / nb.sum(axis=0)
log_nb = np.log(nb)
# ## Testing the Model
log_priors.filter([0], axis=0)
def naive_estimate(log_priors, log_probs, bag_words):
Ck = log_priors + log_probs.filter(bag_words, axis=0).sum(axis=0)
return Ck
y = test["category"].values
# +
yhat = np.empty_like(y)
for ix, (release_date, _) in enumerate(test.iterrows()):
release_date_file = release_date.strftime("%Y%m%d.pdf")
path = os.path.join(wrkdir, release_date_file)
print(path, end="\r")
trgt_minuta = parse_minuta(path, void_words)
Ck = naive_estimate(log_priors.loc[0], log_nb, trgt_minuta.index)
yhat[ix] = Ck.idxmax()
# -
from sklearn.metrics import recall_score, precision_score
confusion_matrix(y, yhat)
precision_score(y, yhat, pos_label="cambio")
recall_score(y, yhat, pos_label="cambio")
|
notebooks/.ipynb_checkpoints/naive-bayes-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _cell_guid="d78c53b6-3887-4fcc-bcc4-5b6c6683bcf1" _execution_state="idle" _uuid="184466e49a15d048e7b11f138e0fa7ab69578ab7"
# # High level insight on genetic variations
# Note: As this is my first published Kernel, am open to suggestions. If this helped you, some upvotes would be very much appreciated.
#
# ### Library and Settings
# Import required library and define constants
# + _cell_guid="fbe8a2a6-cca2-4a3d-b599-45798365819f" _execution_state="idle" _uuid="1804e866346ea1337e4149cc27267a632dee5cf8"
import os
import math
import numpy as np
import pandas as pd
import seaborn as sns
# %matplotlib inline
import matplotlib.pyplot as plt
from sklearn.feature_extraction.text import TfidfVectorizer
# + [markdown] _cell_guid="c82e43c7-3cf6-45a9-8d24-7196dcf5a27c" _execution_state="idle" _uuid="490d6e3bb8b7a0d482bf2a6f63cd53e2b24b18a9"
# ### Files
# + [markdown] _cell_guid="f644847a-b4be-40e2-99cb-5891fa6c392c" _execution_state="idle" _uuid="263fa404d856c0ccb07905d2230056e35f2039dc"
# Training data size is smaller than testing counterpart.
# + [markdown] _cell_guid="ca304d7a-624f-46f8-9dff-35bae62ceffd" _execution_state="idle" _uuid="096ec85c508ecba82d9e98fc6d25ba8df08aea57"
# ### Sneak Peak of data
# Load training and testing data. Have a quick look at columns, its shape and values
# + _cell_guid="d37bf0dc-f697-4db9-b7a8-f40f75548f66" _execution_state="idle" _uuid="1b24f2cc5d7e3946c0d66c96959bfd0bcd19055c"
data_path = "/home/robin/datatsets/msk-redefining-cancer-treatment"
train_variants_df = pd.read_csv(os.path.join(data_path, "training_variants"))
test_variants_df = pd.read_csv(os.path.join(data_path, "test_variants"))
train_text_df = pd.read_csv(os.path.join(data_path, "training_text"), sep="\|\|", engine="python", skiprows=1, names=["ID", "Text"])
test_text_df = pd.read_csv(os.path.join(data_path, "test_text"), sep="\|\|", engine="python", skiprows=1, names=["ID", "Text"])
print("Train Variant", train_variants_df.shape)
print("Train Text", train_text_df.shape)
print("Test Variant", test_variants_df.shape)
print("Test Text", test_text_df.shape)
# + [markdown] _cell_guid="16a6a5f1-9337-4538-9f9b-68b006a47cac" _execution_state="idle" _uuid="16d66eec01bb8395852e079193ce8b9e7c65bf34"
# We have more samples of test data than training data. As mentioned in data introduction, some of the test data is machine-generated to prevent hand labelling.
# + _cell_guid="10ca22a7-f3c4-473b-aa90-179712726be9" _execution_state="idle" _uuid="0a1eaea43a20487b51319f22dbeb12178c8bfe15"
train_variants_df.head()
# -
train_text_df = train_text_df.dropna()
train_text_df.shape
# + _cell_guid="95edade0-0dc9-4691-887f-36e8433f9b21" _execution_state="idle" _uuid="080a37daa067098451c9f78753d21f5bd266fb5d"
print("For training data, there are a total of")
print(len(train_variants_df.ID.unique()), "IDs")
print(len(train_variants_df.Gene.unique()), "unique genes,")
print(len(train_variants_df.Variation.unique()), "unique variations")
print(len(train_variants_df.Class.unique()), "classes")
# + [markdown] _cell_guid="0234ca37-0f15-4689-80a6-fd159982aaf0" _execution_state="idle" _uuid="865b48ac55ca3fe7cc0c46094f1d35df89bf86bd"
# There are 9 classes into which data has to be classified. Lets get the frequency of each class.
# + _cell_guid="7b66a33a-2864-4cd0-b777-d81eb1e19077" _execution_state="idle" _uuid="104a9eb9b34d366b07926b4f8a10709a30c33380"
plt.figure(figsize=(12,8))
sns.countplot(x="Class", data=train_variants_df, palette="Blues_d")
plt.ylabel('Frequency', fontsize=14)
plt.xlabel('Class', fontsize=14)
plt.title("Distribution of genetic mutation classes", fontsize=18)
plt.show()
# + _cell_guid="beb4ed25-db32-4d0d-9fd7-16a9cc5407e1" _execution_state="idle" _uuid="f920514912d3c25458964e08a16d191608eb9caa"
gene_group = train_variants_df.groupby("Gene")['Gene'].count()
minimal_occ_genes = gene_group.sort_values(ascending=True)[:10]
print("Genes with maximal occurences\n", gene_group.sort_values(ascending=False)[:10])
print("\nGenes with minimal occurences\n", minimal_occ_genes)
# + [markdown] _cell_guid="1d191531-304d-4b1f-9a1c-cccb1699f5f0" _execution_state="idle" _uuid="2fcbfc714f373f44a603cafabe98512500af434a"
# Lets have a look at some genes that has highest number of occurrences in each class.
# + _cell_guid="c5634307-e735-4874-af71-ccb91dba9ebc" _execution_state="idle" _uuid="96c09433647696d09fd915c2395e5922afdfafd8"
fig, axs = plt.subplots(ncols=3, nrows=3, figsize=(15,15))
for i in range(3):
for j in range(3):
gene_count_grp = train_variants_df[train_variants_df["Class"]==((i*3+j)+1)].groupby('Gene')["ID"].size().reset_index(name = "count")
sorted_gene_group = gene_count_grp.sort_values('count', ascending=False)
sorted_gene_group_top_7 = gene_count_grp[:7]
sns.barplot(x="Gene", y="count", data=sorted_gene_group_top_7, ax=axs[i][j])
# +
fig, axs = plt.subplots(ncols=3, nrows=3, figsize=(15,15))
for i in range(3):
for j in range(3):
gene_count_grp = train_variants_df[train_variants_df["Class"]==((i*3+j)+1)].groupby('Gene')["ID"].count().reset_index()
sorted_gene_group = gene_count_grp.sort_values('ID', ascending=False)
sorted_gene_group_top_7 = sorted_gene_group[:7]
sns.barplot(x="Gene", y="ID", data=sorted_gene_group_top_7, ax=axs[i][j])
# + [markdown] _cell_guid="e160a091-e198-4b0e-86d8-9aa65f930b98" _execution_state="idle" _uuid="2cabc4835c6aa91c8b74f9dcdf68dc8a9a4c78b0"
# Some points we can conclude from these graphs:
# 1. BRCA1 is highly dominating Class 5
# 2. SF3B1 is highly dominating Class 9
# 3. BRCA1 and BRCA2 are dominating Class 6
# + [markdown] _cell_guid="3aecc202-40fb-4e6e-866d-6580edda4889" _execution_state="idle" _uuid="b36522fd4ddb9e00c4e55985f739ca1bc8afb8a2"
# ## Lets get some insight on text data
# + _cell_guid="aba42894-024d-447c-8e75-ca4fe9f9e031" _execution_state="idle" _uuid="d450d971890e778f823d1fa68ccfe018ea979768"
train_text_df.head()
# -
train_text_df.shape
train_text_df = train_text_df.dropna()
train_text_df.shape
# +
# def func_for(df):
# for index in df.index:
# if type(df.loc[index, 'Text']) == str:
# df.loc[index, 'Text_count'] = len(df.loc[index, 'Text'].split())
# else:
# df.loc[index, 'Text_count'] = 1.0
# return df
# +
# train_text_df = func_for(train_text_df)
# -
train_text_df.loc[:, 'Text_count'] = train_text_df["Text"].apply(lambda x: len(x.split()))
train_text_df.head()
# + [markdown] _cell_guid="c10423e5-5383-4933-a5f9-76333eaba127" _execution_state="idle" _uuid="39d8d88672fa5e982bf272a68d7d0cd0f8221c25"
# Let us combine both dataframes for further use
# + _cell_guid="4fc0a329-92e8-4b0e-b901-0dc7fa0e557d" _execution_state="idle" _uuid="6b18e7c4d6b35908c15f8d8c3c23deb0ff35e6d8"
train_full = train_variants_df.merge(train_text_df, how="inner", left_on="ID", right_on="ID")
train_full[train_full["Class"]==1].head()
# + [markdown] _cell_guid="96cf9f23-8224-4375-a0ec-918842146e67" _execution_state="idle" _uuid="618b5f8429c6aa57dad6abc61cdeb4d134b2db62"
# There are multiple rows with similar texts let us check how many of them are unique and whether all similar texts belongs to same class
# + _cell_guid="1d93418f-aaee-481b-a12b-0b83a34ccc1a" _execution_state="idle" _uuid="ea58c91a84340f226f09d667a2d2650af4d88ff7"
count_grp = train_full.groupby('Class')["Text_count"]
count_grp.describe()
# + [markdown] _cell_guid="806a7ca6-a868-4542-9dfa-830f627d6250" _execution_state="idle" _uuid="268ebca98daa07c533fe0960dc625374e583079a"
# We can see there are some entries with text count of 1. Lets have a look at those entries
# + _cell_guid="827a1683-f308-4c59-afee-f3dc953a62e3" _execution_state="idle" _uuid="f8a18eb037c58d51cc15d3efbd7f3491664df3ba"
train_full[train_full["Text_count"]==1]
# + _execution_state="idle" _uuid="7459c94305e99f364ec33c2d90dae31e9b9f4caf"
train_full[train_full["Text_count"]<500.0]
# + [markdown] _cell_guid="d659f9d9-cf5e-4008-aef3-748c2c3df7bd" _execution_state="idle" _uuid="c54d8d66c69af25e06fd5304f0cba6cce6f11efe"
# As we can see there are some entries without any text data.
# Now let us get distribution of text count for each class
# + _cell_guid="c7949532-ae85-4b1d-b2d1-eab21be15919" _execution_state="idle" _uuid="df106eca64977d5c793a6b6ad51c75b7828994c1"
plt.figure(figsize=(12,8))
gene_count_grp = train_full.groupby('Gene')["Text_count"].sum().reset_index()
sns.violinplot(x="Class", y="Text_count", data=train_full, inner=None)
sns.swarmplot(x="Class", y="Text_count", data=train_full, color="w", alpha=.5);
plt.ylabel('Text Count', fontsize=14)
plt.xlabel('Class', fontsize=14)
plt.title("Text length distribution", fontsize=18)
plt.show()
# + [markdown] _cell_guid="3ab294f0-39d2-49bb-8149-85d218dea8b6" _execution_state="idle" _uuid="cafe66044c78625102c872cf89f72457db5415ed"
# Distribution looks quite interesting and now I am in love with violin plots.
# All classes have most counts in between 0 to 20000. Just as expected.
# There should be some
# + _cell_guid="b3054047-9d4f-4576-bf8f-4a8070676c08" _execution_state="idle" _uuid="01d85f3ee18a543a408b21dae31cf6db7a0eeeb1"
fig, axs = plt.subplots(ncols=3, nrows=3, figsize=(15,15))
for i in range(3):
for j in range(3):
gene_count_grp = train_full[train_full["Class"]==((i*3+j)+1)].groupby('Gene')["Text_count"].mean().reset_index()
sorted_gene_group = gene_count_grp.sort_values('Text_count', ascending=False)
sorted_gene_group_top_7 = sorted_gene_group[:7]
sns.barplot(x="Gene", y="Text_count", data=sorted_gene_group_top_7, ax=axs[i][j])
# + [markdown] _cell_guid="90964677-abc3-4496-b59d-8361b16bd595" _execution_state="idle" _uuid="f55d430e29d76a3d9827b8503a849c5dc279a365"
# Frequently occurring terms for each class
# + [markdown] _cell_guid="15a89b04-4c3f-492a-877b-7b98fdfbc2af" _execution_state="idle" _uuid="97946a1787e6ffefe6f65b0ab85bc32b56accf77"
# We need to know more about text. Tf-idf is known as one good technique to use for text transformation and get good features out of text for training our machine learning model. [Here][1] you can find more details about tf-idf and some useful code snippets.
#
#
# [1]: https://buhrmann.github.io/tfidf-analysis.html
# + _cell_guid="9cb69fbc-af5e-42cf-8110-f8e8c299a423" _execution_state="idle" _uuid="a1b8483f3d1f8e4b1ea8c43856d3f0da09c99328"
def top_tfidf_feats(row, features, top_n=10):
topn_ids = np.argsort(row)[::-1][:top_n]
top_feats = [(features[i], row[i]) for i in topn_ids]
df = pd.DataFrame(top_feats)
df.columns = ['feature', 'tfidf']
return df
def top_feats_in_doc(Xtr, features, row_id, top_n=10):
row = np.squeeze(Xtr[row_id].toarray())
return top_tfidf_feats(row, features, top_n)
def top_mean_feats(Xtr, features, grp_ids=None, min_tfidf=0.1, top_n=10):
if grp_ids:
D = Xtr[grp_ids].toarray()
else:
D = Xtr.toarray()
D[D < min_tfidf] = 0
tfidf_means = np.mean(D, axis=0)
return top_tfidf_feats(tfidf_means, features, top_n)
def top_feats_by_class(Xtr, y, features, min_tfidf=0.1, top_n=10):
dfs = []
labels = np.unique(y)
for label in labels:
ids = np.where(y==label)
feats_df = top_mean_feats(Xtr, features, ids, min_tfidf=min_tfidf, top_n=top_n)
feats_df.label = label
dfs.append(feats_df)
return dfs
def plot_tfidf_classfeats_h(dfs):
fig = plt.figure(figsize=(12, 100), facecolor="w")
x = np.arange(len(dfs[0]))
for i, df in enumerate(dfs):
#z = int(str(int(i/3)+1) + str((i%3)+1))
ax = fig.add_subplot(9, 1, i+1)
ax.spines["top"].set_visible(False)
ax.spines["right"].set_visible(False)
ax.set_frame_on(False)
ax.get_xaxis().tick_bottom()
ax.get_yaxis().tick_left()
ax.set_xlabel("Mean Tf-Idf Score", labelpad=16, fontsize=16)
ax.set_ylabel("Gene", labelpad=16, fontsize=16)
ax.set_title("Class = " + str(df.label), fontsize=18)
ax.ticklabel_format(axis='x', style='sci', scilimits=(-2,2))
ax.barh(x, df.tfidf, align='center')
ax.set_yticks(x)
ax.set_ylim([-1, x[-1]+1])
yticks = ax.set_yticklabels(df.feature)
plt.subplots_adjust(bottom=0.09, right=0.97, left=0.15, top=0.95, wspace=0.52)
plt.show()
# + [markdown] _cell_guid="4f884aee-b395-47b4-81e9-34a1f42a50b9" _execution_state="idle" _uuid="203d234937611553cca1090d8fe2125d5587f565"
# Lets plot out some top features we got using Tf-Idf for each class
# + _cell_guid="f415354e-4459-409d-b803-d137ebf0b64a" _execution_state="idle" _uuid="9b4bd847d8b258cb97754c38a3836fce701cef11"
tfidf = TfidfVectorizer(
min_df=5, max_features=16000, strip_accents='unicode',lowercase =True,
analyzer='word', token_pattern=r'\w+', use_idf=True,
smooth_idf=True, sublinear_tf=True, stop_words = 'english').fit(train_full["Text"])
Xtr = tfidf.fit_transform(train_full["Text"])
y = train_full["Class"]
features = tfidf.get_feature_names()
top_dfs = top_feats_by_class(Xtr, y, features)
# + _cell_guid="e15a6894-6c4b-49cb-af1e-2bef22b99399" _execution_state="idle" _uuid="69a2cfadb1e0f7e6da544ccfc25d1bdd5234eee3"
plot_tfidf_classfeats_h(top_dfs)
# + [markdown] _cell_guid="b355216f-301d-401c-a3ac-3c1456d9f52b" _execution_state="busy" _uuid="c775d954b90c6b3877128137b37d09d54bc0a301"
#
# To be continued...
|
docs/data-maining/brief-insight-on-genetic-variations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Bias-correction for weather forecasts
#
# We will use IBM PAIRS to study and correct biases in the GFS forecast. In detail, we will compare the temperature forecast made by NOAA's GFS with the ERA5 reanalysis. Aggregating 5 years worth of data will give evidence of a systematic yet location-dependent bias in the GFS data. We will then use PAIRS to remove these biases from the GFS forecast, leading to an MAE reduction of roughly 50%.
#
# For details regarding both data layers, consult the PAIRS data explorer at https://ibmpairs.mybluemix.net/data-explorer. The layer IDs are
# - 49423 (ERA5)
# - 50195 (GFS)
from datetime import datetime, timedelta
import os, numpy as np, pandas as pd
from matplotlib import pyplot as plt
from ibmpairs import paw
# To start, we set some global variables. I.e. the name of your user account et cetera. Note that this assumes the existence of a file `ibmpairspass.txt` in your home directory containing access credentials for IBM PAIRS.
PAIRS_USER = '<username>'
PAIRS_SERVER = 'https://pairs.res.ibm.com'
BASE_URI = '/'
PAIRS_PASSWORD = paw.get_pairs_api_password(PAIRS_SERVER, PAIRS_USER, passFile=os.path.expanduser('~/ibmpairspass.txt'))
PAIRS_CREDENTIALS = (PAIRS_USER, PAIRS_PASSWORD)
# The following helps when converting `datetime` objects to strings in ISO 8601-compliant format.
iso8601 = '%Y-%m-%dT%H:%M:%SZ'
# ## Step 1: Temporal joins between the GFS and ERA5 data
#
# Since the GFS data is a forecast, it does not only depend on a single timestamp. Indeed, a forecast is characterized both by its *valid time* and *issue time*. I.e. the time the forecast is for and the time it was issued. The difference between these is known as the *lead time* or *horizon*. I.e. `horizon = valid time - issue time`. Most forecasts in PAIRS are stored by issue time, simplifying the comparison with observations. The horizon is then an additional dimension, usually in hours. (For details consult the data explorer.)
#
# To keep things simple, we will only consider the GFS data at horizon 6 h. I.e. predictions which were made 6 hours into the future. (Since the GFS predicts up to 15 days ahead, the maximum horizon is 360 hours.) The forecast is issued daily, which means that as long as we keep the horizon fixed at 6 we have one value every 24 hours. (To obtain data at higher frequencies, we could query additional horizons.)
#
# The ERA5 data on the other hand is hourly. To join these two, we have to ensure that we only request timestamps for which both are defined. (The situation would be a bit different for a parameter which is accumulcated over a certain amount of time such as precipitation. Since temperature is generally considered an instantaneous quantity, we can simply join identical timestamps. Details on whether a parameter is instantaneous or defined over a certain measurement interval can be found in the data explorer.
#
# Thus, we make a point query to both data layers to find all timestamps during the years 2014 to 2018.
pointQueryJson = {
'layers' : [
{
'type' : 'raster', 'id' : '50195', 'dimensions' : [{'name' : 'horizon', 'value' : '6'}]
},
{
'type' : 'raster', 'id' : '49423'
}
],
'spatial' : {'type' : 'point', 'coordinates' : ['40', '-100']},
'temporal' : {'intervals' : [{
'start' : (datetime(2014, 1, 1) - timedelta(seconds = 1)).strftime(iso8601),
'end' : datetime(2019, 1, 1).strftime(iso8601)
}]}
}
pointQuery = paw.PAIRSQuery(pointQueryJson, PAIRS_SERVER, PAIRS_CREDENTIALS)
pointQuery.submit()
pointQuery.vdf['value'] = pd.to_numeric(pointQuery.vdf['value'])
# Some simple *pandas* transformations allow us to identify timestamps for which both layers are defined.
#
# **Note** we can use this simple point query since both the forecast and reanalysis data is defined for all locations in the world at the same time. I.e. if we find data at a single time and point, we know that there will be data everywhere else in the world at that timestamp. This would not be the case were we to compare satellite data from e.g. Sentinel or Landsat satellites.
pointQuery.vdf.pivot_table(index = 'timestamp', columns = 'layerId', values = 'value').dropna().head()
completeDates = pointQuery.vdf.pivot_table(index = 'timestamp', columns = 'layerId', values = 'value').dropna().index.to_series(keep_tz = True)
# ## Step 2a: Identifying the overall bias
#
# To proceed, we use the timestamps stored in `completeDates` to compare the GFS and ERA5 data at those timestamps. Since taking the mean and subtraction are interchangable operations, we take the mean for each dataset before calculating the bias. (Simply taking the average of all values in 2014-2018 would have led to wildly inconsistent results. At fixed horizon, the GFS layer only contains values for 0:00 UTC while the ERA5 one contains hourly data. Thus we would have compared aggregates over 0:00 UTC with those over all hours of the day.)
biasQueryJson = {
'layers' : [
{
'alias' : 'gfs',
'type' : 'raster', 'id' : '50195', 'dimensions' : [{'name' : 'horizon', 'value' : '6'}],
'aggregation' : 'Mean',
'temporal' : {'intervals' : [{'snapshot' : ts.strftime(iso8601)} for _, ts in completeDates.iteritems()]},
'output' : False
},
{
'alias' : 'era5',
'type' : 'raster', 'id' : '49423', 'aggregation' : 'Mean',
'temporal' : {'intervals' : [{'snapshot' : ts.strftime(iso8601)} for _, ts in completeDates.iteritems()]},
'output' : False
},
{
'alias' : 'bias',
'expression' : '$gfs - $era5'
}
],
'spatial' : {'type' : 'square', 'coordinates' : ['-90', '-170', '90', '170']},
'temporal' : {'intervals' : [{
'start' : (datetime(2015, 1, 1) - timedelta(seconds = 1)).strftime(iso8601),
'end' : datetime(2016, 1, 1).strftime(iso8601)
}]}
}
biasQuery = paw.PAIRSQuery(biasQueryJson, PAIRS_SERVER, PAIRS_CREDENTIALS)
biasQuery.submit()
# Note: The queries in this notebook are somewhat substantial and should take 5-10 minutes to complete. Calling `.poll_till_finished()` will thus block the notebook for an extended time. It's worth while to check the status of the query by calling `.poll()` and checking the `.queryStatus` object:
biasQuery.poll()
biasQuery.queryStatus.json()['status']
biasQuery.poll_till_finished()
biasQuery.download()
biasQuery.create_layers()
# At first sight, we find fairly clear evidence of systematic bias. Temperatures in North- and South-America are generally predicted as too low while those in North and South-Africa appear to be too high.
plt.figure(figsize = (20, 8))
plt.imshow(
biasQuery.data['Expression-bias[bias]-Exp'],
vmin = -5, vmax = 5, cmap = 'seismic',
extent = [biasQuery.metadata['Expression-bias[bias]-Exp']['details']['boundingBox'][l] for l in ['minLongitude', 'maxLongitude', 'minLatitude', 'maxLatitude']]
)
plt.colorbar(label = 'Error [K]')
plt.title('Mean bias in GFS data 2014-2018')
plt.savefig('MeanBiasInGFS2014-2018.png', dpi = 60, bbox_inches = 'tight')
plt.show()
# Having said that, it is instructive to look at the overall error distribution. Accumulating spatially leads to what is an essentially neglegible bias.
pd.Series(biasQuery.data['Expression-bias[bias]-Exp'].reshape(-1)).dropna().hist(bins = 25, log = True)
plt.show()
pd.Series(biasQuery.data['Expression-bias[bias]-Exp'].reshape(-1)).dropna().describe()
# Finally, we note an mean absolute error (MAE) of about 0.86 degrees Kelvin.
pd.Series(biasQuery.data['Expression-bias[bias]-Exp'].reshape(-1)).dropna().abs().mean()
# ## Step 2b: Temporal dependence of the biases
#
# One might ask whether the bias we identified in the previous query is a systematic bias or an artifact of the aggregation. So let us take a further look. The following query calculates the bias for each year in the 5-year period independently.
# +
annualLayers = [
[
{
'alias' : 'gfs_{}'.format(year),
'type' : 'raster', 'id' : '50195', 'dimensions' : [{'name' : 'horizon', 'value' : '6'}],
'aggregation' : 'Mean',
'temporal' : {'intervals' : [{'snapshot' : ts.strftime(iso8601)} for _, ts in completeDates.iteritems() if ts.year == year]},
'output' : False
},
{
'alias' : 'era5_{}'.format(year),
'type' : 'raster', 'id' : '49423', 'aggregation' : 'Mean',
'temporal' : {'intervals' : [{'snapshot' : ts.strftime(iso8601)} for _, ts in completeDates.iteritems() if ts.year == year]},
'output' : False
},
{
'alias' : 'bias_{}'.format(year),
'expression' : '$gfs_{year} - $era5_{year}'.format(year = year),
}
] for year in range(2014, 2019)
]
annualLayers = [ll for l in annualLayers for ll in l]
annualBiasQueryJson = {
'layers' : annualLayers,
'spatial' : {'type' : 'square', 'coordinates' : ['-90', '-170', '90', '170']},
'temporal' : {'intervals' : [{
'start' : (datetime(2015, 1, 1) - timedelta(seconds = 1)).strftime(iso8601),
'end' : datetime(2016, 1, 1).strftime(iso8601)
}]}
}
# -
annualBiasQuery = paw.PAIRSQuery(annualBiasQueryJson, PAIRS_SERVER, PAIRS_CREDENTIALS)
annualBiasQuery.submit()
annualBiasQuery.poll()
annualBiasQuery.queryStatus.json()['status']
annualBiasQuery.poll_till_finished()
annualBiasQuery.download()
annualBiasQuery.create_layers()
# To keep the plot simple we only show data for 2014 - 2017. The situation is essentially the same for 2018.
extent = extent = [annualBiasQuery.metadata['Expression-bias_2014[bias_2014]-Exp']['details']['boundingBox'][l] for l in ['minLongitude', 'maxLongitude', 'minLatitude', 'maxLatitude']]
fig, ax = plt.subplots(2, 2, figsize = (30, 16), sharex = True, sharey = True)
ax[0][0].imshow(annualBiasQuery.data['Expression-bias_2014[bias_2014]-Exp'], vmin = -5, vmax = 5, cmap = 'seismic', extent = extent)
ax[0][1].imshow(annualBiasQuery.data['Expression-bias_2015[bias_2015]-Exp'], vmin = -5, vmax = 5, cmap = 'seismic', extent = extent)
ax[1][0].imshow(annualBiasQuery.data['Expression-bias_2016[bias_2016]-Exp'], vmin = -5, vmax = 5, cmap = 'seismic', extent = extent)
ax[1][1].imshow(annualBiasQuery.data['Expression-bias_2017[bias_2017]-Exp'], vmin = -5, vmax = 5, cmap = 'seismic', extent = extent)
ax[0][0].set_title('2014')
ax[0][1].set_title('2015')
ax[1][0].set_title('2016')
ax[1][1].set_title('2017')
plt.tight_layout()
plt.savefig('TemporalDependenceOfBias.png', dpi = 60, bbox_inches = 'tight')
plt.show()
# As we can see, the biases are fairly consistent with time. We see differences in the details between the above aggregation periods, but overall structures are the same. I.e. too low predictions in the Americas, Europe and at the North Pole, too high predictions in North and South Africa, Northern and Central Asia as well as Antarctica.
# ## Step 3: A bias corrected forecast
#
# Having confirmed the existence of a bias, we can now issue a bias-corrected forecast for 2019. I.e. we use the bias measured during the previous 5-year period to shift the predictions for 2019. To start, we again make a point query to facilitat the temporal join.
pointQuery2019Json = {
'layers' : [
{
'type' : 'raster', 'id' : '50195', 'dimensions' : [{'name' : 'horizon', 'value' : '6'}]
},
{
'type' : 'raster', 'id' : '49423'
}
],
'spatial' : {'type' : 'point', 'coordinates' : ['40', '-100']},
'temporal' : {'intervals' : [{
'start' : (datetime(2019, 1, 1) - timedelta(seconds = 1)).strftime(iso8601),
'end' : datetime(2019, 7, 1).strftime(iso8601)
}]}
}
pointQuery2019 = paw.PAIRSQuery(pointQuery2019Json, PAIRS_SERVER, PAIRS_CREDENTIALS)
pointQuery2019.submit()
pointQuery2019.vdf['value'] = pd.to_numeric(pointQuery2019.vdf['value'])
completeDates2019 = pointQuery2019.vdf.pivot_table(index = 'timestamp', columns = 'layerId', values = 'value').dropna().index.to_series(keep_tz = True)
# The following query again calculates the forecast bias of the years 2014-2018 and applies it to forecasts for the first half of 2019. Subsequently we calculate the bias of both the raw and bias-corrected 2019 forecasts.
biasCorrectedQueryJson = {
'layers' : [
{
'alias' : 'historic_gfs',
'type' : 'raster', 'id' : '50195', 'dimensions' : [{'name' : 'horizon', 'value' : '6'}],
'aggregation' : 'Mean',
'temporal' : {'intervals' : [{'snapshot' : ts.strftime(iso8601)} for _, ts in completeDates.iteritems()]},
'output' : False
},
{
'alias' : 'historic_era5',
'type' : 'raster', 'id' : '49423', 'aggregation' : 'Mean',
'temporal' : {'intervals' : [{'snapshot' : ts.strftime(iso8601)} for _, ts in completeDates.iteritems()]},
'output' : False
},
{
'alias' : 'gfs',
'type' : 'raster', 'id' : '50195', 'dimensions' : [{'name' : 'horizon', 'value' : '6'}],
'aggregation' : 'Mean',
'temporal' : {'intervals' : [{'snapshot' : ts.strftime(iso8601)} for _, ts in completeDates2019.iteritems()]},
'output' : False
},
{
'alias' : 'era5',
'type' : 'raster', 'id' : '49423', 'aggregation' : 'Mean',
'temporal' : {'intervals' : [{'snapshot' : ts.strftime(iso8601)} for _, ts in completeDates2019.iteritems()]},
'output' : False
},
{
'alias' : 'bias',
'expression' : '$gfs - $era5',
'output' : True
},
{
'alias' : 'bias_corrected_forecast',
'expression' : '$gfs - $historic_gfs + $historic_era5',
'output' : True
},
{
'alias' : 'bias_of_bias_corrected_forecast',
'expression' : '$gfs - $historic_gfs + $historic_era5 - $era5',
'output' : True
}
],
'spatial' : {'type' : 'square', 'coordinates' : ['-90', '-170', '90', '170']},
'temporal' : {'intervals' : [{
'start' : (datetime(2015, 1, 1) - timedelta(seconds = 1)).strftime(iso8601),
'end' : datetime(2016, 1, 1).strftime(iso8601)
}]}
}
biasCorrectedQuery = paw.PAIRSQuery(biasCorrectedQueryJson, PAIRS_SERVER, PAIRS_CREDENTIALS)
biasCorrectedQuery.submit()
biasCorrectedQuery.poll_till_finished()
biasCorrectedQuery.download()
biasCorrectedQuery.create_layers()
# ## Step 4: Results
#
# We can finally analyze the impact of the bias correction. To do so we both plot the spatial distribution of biases but also calculate a number of global metrics. To start, let's take a look at the spatial distribution.
extent = extent = [biasCorrectedQuery.metadata['Expression-bias[bias]-Exp']['details']['boundingBox'][l] for l in ['minLongitude', 'maxLongitude', 'minLatitude', 'maxLatitude']]
fig, ax = plt.subplots(2, 1, figsize = (30, 16), sharex = True, sharey = True)
ax[0].imshow(
biasCorrectedQuery.data['Expression-bias[bias]-Exp'], vmin = -5, vmax = 5, cmap = 'seismic', extent = extent
)
ax[1].imshow(
biasCorrectedQuery.data['Expression-bias_of_bias_corrected_forecast[bias_of_bias_corrected_forecast]-Exp'], vmin = -5, vmax = 5, cmap = 'seismic', extent = extent
)
ax[0].set_title('Bias of raw forecast')
ax[1].set_title('Bias of bias-corrected forecast')
plt.tight_layout()
plt.savefig('PerformanceOfBiasCorrectedForecast.png', dpi = 60, bbox_inches = 'tight')
plt.show()
# Since the two plots above are on the same color scale (see the use of `vmin` and `vmax`), we can make direct comparisons from the colors. Clearly, the bias corrected forecast shows an overall improvement. Note that the underprediction in the Americas, the Indian subcontinent and Europe has turned into an overprediction. The situation is somewhat more complicated in the rest of the world though.
#
# To complement this impression we calculate some general metrics. Note that the mean absolute error (MAE) has improved quite significantly.
pd.DataFrame({
'Raw forecast' : pd.Series(biasCorrectedQuery.data['Expression-bias[bias]-Exp'].reshape(-1)).dropna().describe().append(pd.Series({'MAE' : pd.Series(biasCorrectedQuery.data['Expression-bias[bias]-Exp'].reshape(-1)).dropna().abs().mean()})),
'Bias-corrected forecast' : pd.Series(biasCorrectedQuery.data['Expression-bias_of_bias_corrected_forecast[bias_of_bias_corrected_forecast]-Exp'].reshape(-1)).dropna().describe().append(pd.Series({'MAE' : pd.Series(biasCorrectedQuery.data['Expression-bias_of_bias_corrected_forecast[bias_of_bias_corrected_forecast]-Exp'].reshape(-1)).dropna().abs().mean()}))
}).round(2)
|
examples/BiasCorrectionOfWeatherForecasts/biasCorrection.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# TSG035 - Spark History logs
# ===========================
#
# Description
# -----------
#
# Steps
# -----
#
# ### Parameters
# + tags=["parameters"]
import re
tail_lines = 2000
pod = None # All
container='hadoop-livy-sparkhistory'
log_files = [ "/var/log/supervisor/log/sparkhistory*" ]
expressions_to_analyze = [
re.compile(".{23} WARN "),
re.compile(".{23} ERROR ")
]
# -
# ### Instantiate Kubernetes client
# + tags=["hide_input"]
# Instantiate the Python Kubernetes client into 'api' variable
import os
try:
from kubernetes import client, config
from kubernetes.stream import stream
if "KUBERNETES_SERVICE_PORT" in os.environ and "KUBERNETES_SERVICE_HOST" in os.environ:
config.load_incluster_config()
else:
config.load_kube_config()
api = client.CoreV1Api()
print('Kubernetes client instantiated')
except ImportError:
from IPython.display import Markdown
display(Markdown(f'SUGGEST: Use [SOP059 - Install Kubernetes Python module](../install/sop059-install-kubernetes-module.ipynb) to resolve this issue.'))
raise
# -
# ### Get the namespace for the big data cluster
#
# Get the namespace of the big data cluster from the Kuberenetes API.
#
# NOTE: If there is more than one big data cluster in the target
# Kubernetes cluster, then set \[0\] to the correct value for the big data
# cluster.
# + tags=["hide_input"]
# Place Kubernetes namespace name for BDC into 'namespace' variable
try:
namespace = api.list_namespace(label_selector='MSSQL_CLUSTER').items[0].metadata.name
except IndexError:
from IPython.display import Markdown
display(Markdown(f'SUGGEST: Use [TSG081 - Get namespaces (Kubernetes)](../monitor-k8s/tsg081-get-kubernetes-namespaces.ipynb) to resolve this issue.'))
display(Markdown(f'SUGGEST: Use [TSG010 - Get configuration contexts](../monitor-k8s/tsg010-get-kubernetes-contexts.ipynb) to resolve this issue.'))
display(Markdown(f'SUGGEST: Use [SOP011 - Set kubernetes configuration context](../common/sop011-set-kubernetes-context.ipynb) to resolve this issue.'))
raise
print('The kubernetes namespace for your big data cluster is: ' + namespace)
# -
# ### Get tail for log
# + tags=["hide_input"]
# Display the last 'tail_lines' of files in 'log_files' list
pods = api.list_namespaced_pod(namespace)
entries_for_analysis = []
for p in pods.items:
if pod is None or p.metadata.name == pod:
for c in p.spec.containers:
if container is None or c.name == container:
for log_file in log_files:
print (f"- LOGS: '{log_file}' for CONTAINER: '{c.name}' in POD: '{p.metadata.name}'")
try:
output = stream(api.connect_get_namespaced_pod_exec, p.metadata.name, namespace, command=['/bin/sh', '-c', f'tail -n {tail_lines} {log_file}'], container=c.name, stderr=True, stdout=True)
except Exception:
print (f"FAILED to get LOGS for CONTAINER: {c.name} in POD: {p.metadata.name}")
else:
for line in output.split('\n'):
for expression in expressions_to_analyze:
if expression.match(line):
entries_for_analysis.append(line)
print(line)
print("")
print(f"{len(entries_for_analysis)} log entries found for further analysis.")
# -
# ### Analyze log entries and suggest relevant Troubleshooting Guides
# + tags=["hide_input"]
# Analyze log entries and suggest further relevant troubleshooting guides
from IPython.display import Markdown
tsgs = []
suggestions = 0
for entry in entries_for_analysis:
print (entry)
for tsg in tsgs:
if entry.find(tsg[0]) != -1:
display(Markdown(f'SUGGEST: Use [{tsg[2]}](tsg[1]) to resolve this issue.'))
suggestions = suggestions + 1
print("")
print(f"{len(entries_for_analysis)} log entries analyzed. {suggestions} further troubleshooting suggestions made inline.")
# -
print('Notebook execution complete.')
|
Big-Data-Clusters/GDR1/public/content/log-analyzers/tsg035-get-sparkhistory-logs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pathlib import Path
from netaddr import IPNetwork, IPRange, IPAddress
import pandas
jpnic = []
for l in pandas.read_csv('data/jpnic-ipv4_20200121.csv').itertuples():
jpnic.extend(IPRange(l.start, l.end).cidrs())
iana = pandas.read_csv('data/iana-ipv4_20200120.csv')
apnic = [IPNetwork(_.replace('/', '.0.0.0/')) for _ in iana[iana['Designation'] == 'APNIC']['Prefix']]
ipaddr = [IPAddress(_) for _ in Path('tests/test_address').read_text().splitlines()]
# -
ipaddr[:10]
# +
import ClustIPy
cidrs = ClustIPy.agglomerative(ipaddr, 30)
cidrs[:10]
# +
import numpy as np
from netaddr import IPNetwork
from matplotlib.pyplot import imsave
coords = np.load('data/hilvert_p12_n2.npz')['arr_0']
img = np.zeros((4096, 4096))
for net in apnic:
try:
for s in net.subnet(24):
pos = int(int(s.ip)/256)
x, y = coords[pos, :]
img[x, y] = 32
except:
pass
for net in jpnic:
try:
for s in net.subnet(24):
pos = int(int(s.ip)/256)
x, y = coords[pos, :]
img[x, y] = 64
except:
pass
for net in cidrs:
try:
for s in net.subnet(24):
pos = int(int(s.ip)/256)
x, y = coords[pos, :]
img[x, y] = 128 + 64
except:
pass
for ip in ipaddr:
pos = int(int(IPNetwork(f'{ip}/24').network)/256)
x, y = coords[pos, :]
img[x, y] = 255
imsave('viz_aws_gip.pdf', img)
# +
from matplotlib.pyplot import subplots, get_cmap
fig, ax = subplots(figsize=(16, 16))
ax.set_xticks([])
ax.set_yticks([])
ax.text(0, -10, 'APNIC', color=get_cmap('viridis')(31), weight='heavy', size=15)
ax.text(400, -10, 'JPNIC', color=get_cmap('viridis')(63), weight='heavy', size=15)
ax.text(800, -10, 'Cluster', color=get_cmap('viridis')(192), weight='heavy', size=15)
ax.imshow(img)
|
viz_aws_gip.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="v1CUZ0dkOo_F"
# ##### Copyright 2019 The TensorFlow Authors.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + cellView="form" id="qmkj-80IHxnd"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="_xnMOsbqHz61"
# # Pix2Pix
# + [markdown] id="Ds4o1h4WHz9U"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/generative/pix2pix"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />Смотрите на TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs-l10n/blob/master/site/ru/tutorials/generative/pix2pix.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Запустите в Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs-l10n/blob/master/site/ru/tutorials/generative/pix2pix.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />Изучайте код на GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs-l10n/site/ru/tutorials/generative/pix2pix.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Скачайте ноутбук</a>
# </td>
# </table>
# + [markdown] id="8a2322303a3f"
# Note: Вся информация в этом разделе переведена с помощью русскоговорящего Tensorflow сообщества на общественных началах. Поскольку этот перевод не является официальным, мы не гарантируем что он на 100% аккуратен и соответствует [официальной документации на английском языке](https://www.tensorflow.org/?hl=en). Если у вас есть предложение как исправить этот перевод, мы будем очень рады увидеть pull request в [tensorflow/docs](https://github.com/tensorflow/docs) репозиторий GitHub. Если вы хотите помочь сделать документацию по Tensorflow лучше (сделать сам перевод или проверить перевод подготовленный кем-то другим), напишите нам на [<EMAIL> list](https://groups.google.com/a/tensorflow.org/forum/#!forum/docs-ru).
# + [markdown] id="ITZuApL56Mny"
# Это руководство демонстрирует преобразование изображение в изображение с использованием условных GAN, как описано в [Преобразование изображения в изображение с использованием условных состязательных сетей](https://arxiv.org/abs/1611.07004). Используя эту технику, мы можем раскрасить черно-белые фотографии, преобразовать карты Google в Google Earth и т. д. В этом руководстве мы преобразовываем фасады зданий в настоящие здания.
#
# В примере мы будем использовать [База данных фасадов CMP](http://cmp.felk.cvut.cz/~tylecr1/facade/), любезно предоставленная [Центром машинного восприятия](http://cmp.felk .cvut.cz/) в [Чешском техническом университете в Праге](https://www.cvut.cz/). Чтобы наш пример был кратким, мы будем использовать предварительно обработанную [копию](https://people.eecs.berkeley.edu/~tinghuiz/projects/pix2pix/datasets/) этого набора данных, созданную авторами [paper](https://arxiv.org/abs/1611.07004) выше.
#
# Каждая эпоха занимает около 15 секунд на одном графическом процессоре V100.
#
# Ниже приведен результат, полученный после обучения модели для 200 эпох.
#
# 
# 
# + [markdown] id="e1_Y75QXJS6h"
# ## Импорт TensorFlow и других библиотек
# + id="YfIk2es3hJEd"
import tensorflow as tf
import os
import time
from matplotlib import pyplot as plt
from IPython import display
# + id="wifwThPoEj7e"
# !pip install -U tensorboard
# + [markdown] id="iYn4MdZnKCey"
# ## Загрузка датасета
#
# Вы можете загрузить этот набор данных и аналогичные ему [здесь](https://people.eecs.berkeley.edu/~tinghuiz/projects/pix2pix/datasets). Как упоминалось в [статье](https://arxiv.org/abs/1611.07004), мы применяем случайную рябь и зеркальное отображение к набору обучающих данных.
#
# * При случайной ряби размер изображения изменяется до `286 x 286`, а затем случайным образом обрезается до `256 x 256`.
# * При случайном зеркальном отображении изображение переворачивается по горизонтали, т.е. слева направо.
# + id="Kn-k8kTXuAlv"
_URL = 'https://people.eecs.berkeley.edu/~tinghuiz/projects/pix2pix/datasets/facades.tar.gz'
path_to_zip = tf.keras.utils.get_file('facades.tar.gz',
origin=_URL,
extract=True)
PATH = os.path.join(os.path.dirname(path_to_zip), 'facades/')
# + id="2CbTEt448b4R"
BUFFER_SIZE = 400
BATCH_SIZE = 1
IMG_WIDTH = 256
IMG_HEIGHT = 256
# + id="aO9ZAGH5K3SY"
def load(image_file):
image = tf.io.read_file(image_file)
image = tf.image.decode_jpeg(image)
w = tf.shape(image)[1]
w = w // 2
real_image = image[:, :w, :]
input_image = image[:, w:, :]
input_image = tf.cast(input_image, tf.float32)
real_image = tf.cast(real_image, tf.float32)
return input_image, real_image
# + id="4OLHMpsQ5aOv"
inp, re = load(PATH+'train/100.jpg')
# преобразуем в int чтобы matplotlib мог показать изображение
plt.figure()
plt.imshow(inp/255.0)
plt.figure()
plt.imshow(re/255.0)
# + id="rwwYQpu9FzDu"
def resize(input_image, real_image, height, width):
input_image = tf.image.resize(input_image, [height, width],
method=tf.image.ResizeMethod.NEAREST_NEIGHBOR)
real_image = tf.image.resize(real_image, [height, width],
method=tf.image.ResizeMethod.NEAREST_NEIGHBOR)
return input_image, real_image
# + id="Yn3IwqhiIszt"
def random_crop(input_image, real_image):
stacked_image = tf.stack([input_image, real_image], axis=0)
cropped_image = tf.image.random_crop(
stacked_image, size=[2, IMG_HEIGHT, IMG_WIDTH, 3])
return cropped_image[0], cropped_image[1]
# + id="muhR2cgbLKWW"
# нормализуем изображение до [-1, 1]
def normalize(input_image, real_image):
input_image = (input_image / 127.5) - 1
real_image = (real_image / 127.5) - 1
return input_image, real_image
# + id="fVQOjcPVLrUc"
@tf.function()
def random_jitter(input_image, real_image):
# ресайз до 286 x 286 x 3
input_image, real_image = resize(input_image, real_image, 286, 286)
# случайная обреска до 256 x 256 x 3
input_image, real_image = random_crop(input_image, real_image)
if tf.random.uniform(()) > 0.5:
# случайное зеркальное отображение
input_image = tf.image.flip_left_right(input_image)
real_image = tf.image.flip_left_right(real_image)
return input_image, real_image
# + [markdown] id="wfAQbzy799UV"
# Как вы можете видеть на изображениях ниже,- они проходят через случайное зашумливание
# Случайное зашумливание, описанное в документе, должно
#
# 1. Изменить размер изображения на больший.
# 2. Произвольно обрезать до нужного размера.
# 3. Произвольно перевернуть изображение по горизонтали.
# + id="n0OGdi6D92kM"
plt.figure(figsize=(6, 6))
for i in range(4):
rj_inp, rj_re = random_jitter(inp, re)
plt.subplot(2, 2, i+1)
plt.imshow(rj_inp/255.0)
plt.axis('off')
plt.show()
# + id="tyaP4hLJ8b4W"
def load_image_train(image_file):
input_image, real_image = load(image_file)
input_image, real_image = random_jitter(input_image, real_image)
input_image, real_image = normalize(input_image, real_image)
return input_image, real_image
# + id="VB3Z6D_zKSru"
def load_image_test(image_file):
input_image, real_image = load(image_file)
input_image, real_image = resize(input_image, real_image,
IMG_HEIGHT, IMG_WIDTH)
input_image, real_image = normalize(input_image, real_image)
return input_image, real_image
# + [markdown] id="PIGN6ouoQxt3"
# ## Входной конвейер
# + id="SQHmYSmk8b4b"
train_dataset = tf.data.Dataset.list_files(PATH+'train/*.jpg')
train_dataset = train_dataset.map(load_image_train,
num_parallel_calls=tf.data.experimental.AUTOTUNE)
train_dataset = train_dataset.shuffle(BUFFER_SIZE)
train_dataset = train_dataset.batch(BATCH_SIZE)
# + id="MS9J0yA58b4g"
test_dataset = tf.data.Dataset.list_files(PATH+'test/*.jpg')
test_dataset = test_dataset.map(load_image_test)
test_dataset = test_dataset.batch(BATCH_SIZE)
# + [markdown] id="THY-sZMiQ4UV"
# ## Создание генератора
# * Архитектура генератора - это модифицированная U-Net.
# * Каждый блок в энкодере(Conv -> Batchnorm -> Leaky ReLU)
# * Каждый блок в декодере(Transposed Conv -> Batchnorm -> Dropout (применяется к первым трем блокам) -> ReLU)
# * Между энкодером и декодером есть пропускаемые соединения(как в U-Net).
#
# + id="tqqvWxlw8b4l"
OUTPUT_CHANNELS = 3
# + id="3R09ATE_SH9P"
def downsample(filters, size, apply_batchnorm=True):
initializer = tf.random_normal_initializer(0., 0.02)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2D(filters, size, strides=2, padding='same',
kernel_initializer=initializer, use_bias=False))
if apply_batchnorm:
result.add(tf.keras.layers.BatchNormalization())
result.add(tf.keras.layers.LeakyReLU())
return result
# + id="a6_uCZCppTh7"
down_model = downsample(3, 4)
down_result = down_model(tf.expand_dims(inp, 0))
print (down_result.shape)
# + id="nhgDsHClSQzP"
def upsample(filters, size, apply_dropout=False):
initializer = tf.random_normal_initializer(0., 0.02)
result = tf.keras.Sequential()
result.add(
tf.keras.layers.Conv2DTranspose(filters, size, strides=2,
padding='same',
kernel_initializer=initializer,
use_bias=False))
result.add(tf.keras.layers.BatchNormalization())
if apply_dropout:
result.add(tf.keras.layers.Dropout(0.5))
result.add(tf.keras.layers.ReLU())
return result
# + id="mz-ahSdsq0Oc"
up_model = upsample(3, 4)
up_result = up_model(down_result)
print (up_result.shape)
# + id="lFPI4Nu-8b4q"
def Generator():
inputs = tf.keras.layers.Input(shape=[256,256,3])
down_stack = [
downsample(64, 4, apply_batchnorm=False), # (bs, 128, 128, 64)
downsample(128, 4), # (bs, 64, 64, 128)
downsample(256, 4), # (bs, 32, 32, 256)
downsample(512, 4), # (bs, 16, 16, 512)
downsample(512, 4), # (bs, 8, 8, 512)
downsample(512, 4), # (bs, 4, 4, 512)
downsample(512, 4), # (bs, 2, 2, 512)
downsample(512, 4), # (bs, 1, 1, 512)
]
up_stack = [
upsample(512, 4, apply_dropout=True), # (bs, 2, 2, 1024)
upsample(512, 4, apply_dropout=True), # (bs, 4, 4, 1024)
upsample(512, 4, apply_dropout=True), # (bs, 8, 8, 1024)
upsample(512, 4), # (bs, 16, 16, 1024)
upsample(256, 4), # (bs, 32, 32, 512)
upsample(128, 4), # (bs, 64, 64, 256)
upsample(64, 4), # (bs, 128, 128, 128)
]
initializer = tf.random_normal_initializer(0., 0.02)
last = tf.keras.layers.Conv2DTranspose(OUTPUT_CHANNELS, 4,
strides=2,
padding='same',
kernel_initializer=initializer,
activation='tanh') # (bs, 256, 256, 3)
x = inputs
# понижение размерности
skips = []
for down in down_stack:
x = down(x)
skips.append(x)
skips = reversed(skips[:-1])
# Повышение размерности и установление пропуска соединений
for up, skip in zip(up_stack, skips):
x = up(x)
x = tf.keras.layers.Concatenate()([x, skip])
x = last(x)
return tf.keras.Model(inputs=inputs, outputs=x)
# + id="dIbRPFzjmV85"
generator = Generator()
tf.keras.utils.plot_model(generator, show_shapes=True, dpi=64)
# + id="U1N1_obwtdQH"
gen_output = generator(inp[tf.newaxis,...], training=False)
plt.imshow(gen_output[0,...])
# + [markdown] id="dpDPEQXIAiQO"
# * **Расчет потерь в генераторе**
# * Это сигмоидная кросс-энтропия сгенерированных изображений и **массива из них**.
# * [Статья](https://arxiv.org/abs/1611.07004) также включает потерю L1, которая представляет собой MAE(средняя абсолютная ошибка) между сгенерированным изображением и целевым изображением.
# * Это позволяет сгенерированному изображению стать структурно похожим на целевое изображение.
# * Формула для расчета общих потерь генератора loss = gan_loss + LAMBDA * l1_loss, где LAMBDA = 100. Это значение было определено авторами [статьи](https://arxiv.org/abs/1611.07004).
# + [markdown] id="fSZbDgESHIV6"
# Процедура обучения генератора показана ниже:
# + id="cyhxTuvJyIHV"
LAMBDA = 100
# + id="90BIcCKcDMxz"
def generator_loss(disc_generated_output, gen_output, target):
gan_loss = loss_object(tf.ones_like(disc_generated_output), disc_generated_output)
# mean absolute error
l1_loss = tf.reduce_mean(tf.abs(target - gen_output))
total_gen_loss = gan_loss + (LAMBDA * l1_loss)
return total_gen_loss, gan_loss, l1_loss
# + [markdown] id="TlB-XMY5Awj9"
# 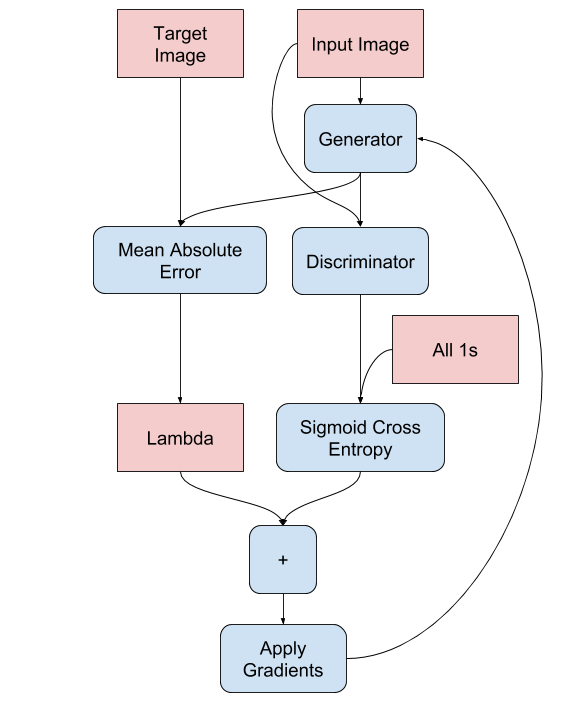
#
# + [markdown] id="ZTKZfoaoEF22"
# ## Создание дискриминатора
# * Дискриминатор - это PatchGAN.
# * Каждый блок в дискриминаторе это (Conv -> BatchNorm -> Leaky ReLU)
# * Размерность вывода после последнего слоя: (batch_size, 30, 30, 1)
# * Каждый патч 30x30 на выходе классифицирует часть входного изображения размером 70x70 (такая архитектура называется PatchGAN).
# * Дискриминатор получает 2 входа.
# * Входное изображение и целевое изображение, которое следует классифицировать как реальное.
# * Входное изображение и сгенерированное изображение(вывод генератора), которое следует классифицировать как подделку.
# * Мы объединяем эти 2 ввода вместе в коде (`tf.concat ([inp, tar], axis = -1)`)
# + id="ll6aNeQx8b4v"
def Discriminator():
initializer = tf.random_normal_initializer(0., 0.02)
inp = tf.keras.layers.Input(shape=[256, 256, 3], name='input_image')
tar = tf.keras.layers.Input(shape=[256, 256, 3], name='target_image')
x = tf.keras.layers.concatenate([inp, tar]) # (bs, 256, 256, channels*2)
down1 = downsample(64, 4, False)(x) # (bs, 128, 128, 64)
down2 = downsample(128, 4)(down1) # (bs, 64, 64, 128)
down3 = downsample(256, 4)(down2) # (bs, 32, 32, 256)
zero_pad1 = tf.keras.layers.ZeroPadding2D()(down3) # (bs, 34, 34, 256)
conv = tf.keras.layers.Conv2D(512, 4, strides=1,
kernel_initializer=initializer,
use_bias=False)(zero_pad1) # (bs, 31, 31, 512)
batchnorm1 = tf.keras.layers.BatchNormalization()(conv)
leaky_relu = tf.keras.layers.LeakyReLU()(batchnorm1)
zero_pad2 = tf.keras.layers.ZeroPadding2D()(leaky_relu) # (bs, 33, 33, 512)
last = tf.keras.layers.Conv2D(1, 4, strides=1,
kernel_initializer=initializer)(zero_pad2) # (bs, 30, 30, 1)
return tf.keras.Model(inputs=[inp, tar], outputs=last)
# + id="YHoUui4om-Ev"
discriminator = Discriminator()
tf.keras.utils.plot_model(discriminator, show_shapes=True, dpi=64)
# + id="gDkA05NE6QMs"
disc_out = discriminator([inp[tf.newaxis,...], gen_output], training=False)
plt.imshow(disc_out[0,...,-1], vmin=-20, vmax=20, cmap='RdBu_r')
plt.colorbar()
# + [markdown] id="AOqg1dhUAWoD"
# **Расчет потерь дискриминатора**
# * Функция потерь дискриминатора принимает 2 входа: **[реальные изображения, сгенерированные изображения]**
# * real_loss - это сигмоидная кросс-энтропия **реальных изображений** и **массива единиц(поскольку это настоящие изображения)**
# * generated_loss - сигмоидная кросс-энтропия **сгенерированных изображений** и **массива нулей(поскольку это поддельные изображения)**
# * В результате **total_loss** - это сумма real_loss и generated_loss
# + id="Q1Xbz5OaLj5C"
loss_object = tf.keras.losses.BinaryCrossentropy(from_logits=True)
# + id="wkMNfBWlT-PV"
def discriminator_loss(disc_real_output, disc_generated_output):
real_loss = loss_object(tf.ones_like(disc_real_output), disc_real_output)
generated_loss = loss_object(tf.zeros_like(disc_generated_output), disc_generated_output)
total_disc_loss = real_loss + generated_loss
return total_disc_loss
# + [markdown] id="-ede4p2YELFa"
# Процедура обучения дискриминатора показана ниже.
#
# Чтобы узнать больше об архитектуре и гиперпараметрах, вы можете обратиться к [статье](https://arxiv.org/abs/1611.07004).
# + [markdown] id="IS9sHa-1BoAF"
# 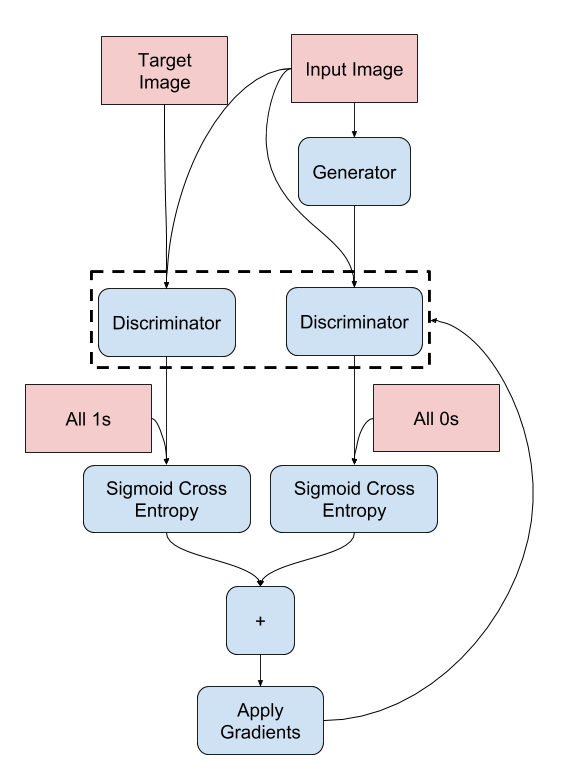
#
# + [markdown] id="0FMYgY_mPfTi"
# ## Определение оптимайзера и сохранения чекпойнтов
#
# + id="lbHFNexF0x6O"
generator_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
discriminator_optimizer = tf.keras.optimizers.Adam(2e-4, beta_1=0.5)
# + id="WJnftd5sQsv6"
checkpoint_dir = './training_checkpoints'
checkpoint_prefix = os.path.join(checkpoint_dir, "ckpt")
checkpoint = tf.train.Checkpoint(generator_optimizer=generator_optimizer,
discriminator_optimizer=discriminator_optimizer,
generator=generator,
discriminator=discriminator)
# + [markdown] id="Rw1fkAczTQYh"
# ## Генерация изображения
#
# Напишите функцию для построения изображений во время обучения.
#
# * Мы передаем изображения из тестового датасета в генератор.
# * Генератор преобразует входное изображение в выходное.
# * Последний шаг - построить прогнозы и **вуаля!**
# + [markdown] id="Rb0QQFHF-JfS"
# Примечание: аргумент `training` установлен в `True` намеренно, так как нам нужна пакетная статистика при запуске модели на тестовом наборе данных.
# Если мы используем `training = False`, мы получим накопленную статистику для всего набора данных(чего мы не хотим).
# + id="RmdVsmvhPxyy"
def generate_images(model, test_input, tar):
prediction = model(test_input, training=True)
plt.figure(figsize=(15,15))
display_list = [test_input[0], tar[0], prediction[0]]
title = ['Input Image', 'Ground Truth', 'Predicted Image']
for i in range(3):
plt.subplot(1, 3, i+1)
plt.title(title[i])
plt.imshow(display_list[i] * 0.5 + 0.5)
plt.axis('off')
plt.show()
# + id="8Fc4NzT-DgEx"
for example_input, example_target in test_dataset.take(1):
generate_images(generator, example_input, example_target)
# + [markdown] id="NLKOG55MErD0"
# ## Обучение
#
# * Для каждого входного изображения генерируем выходное изображение.
# * Дискриминатор получает входное и сгенерированное изображение в качестве первого входа. Второй вход - это входное и целевое изображения.
# * Далее рассчитываем потери генератора и дискриминатора.
# * Затем вычисляем градиенты потерь как для генератора, так и для переменных дискриминатора(входных данных) и применяем их к оптимизатору.
# * Затем логируем потери в TensorBoard
# + id="NS2GWywBbAWo"
EPOCHS = 150
# + id="xNNMDBNH12q-"
import datetime
log_dir="logs/"
summary_writer = tf.summary.create_file_writer(
log_dir + "fit/" + datetime.datetime.now().strftime("%Y%m%d-%H%M%S"))
# + id="KBKUV2sKXDbY"
@tf.function
def train_step(input_image, target, epoch):
with tf.GradientTape() as gen_tape, tf.GradientTape() as disc_tape:
gen_output = generator(input_image, training=True)
disc_real_output = discriminator([input_image, target], training=True)
disc_generated_output = discriminator([input_image, gen_output], training=True)
gen_total_loss, gen_gan_loss, gen_l1_loss = generator_loss(disc_generated_output, gen_output, target)
disc_loss = discriminator_loss(disc_real_output, disc_generated_output)
generator_gradients = gen_tape.gradient(gen_total_loss,
generator.trainable_variables)
discriminator_gradients = disc_tape.gradient(disc_loss,
discriminator.trainable_variables)
generator_optimizer.apply_gradients(zip(generator_gradients,
generator.trainable_variables))
discriminator_optimizer.apply_gradients(zip(discriminator_gradients,
discriminator.trainable_variables))
with summary_writer.as_default():
tf.summary.scalar('gen_total_loss', gen_total_loss, step=epoch)
tf.summary.scalar('gen_gan_loss', gen_gan_loss, step=epoch)
tf.summary.scalar('gen_l1_loss', gen_l1_loss, step=epoch)
tf.summary.scalar('disc_loss', disc_loss, step=epoch)
# + [markdown] id="hx7s-vBHFKdh"
# Фактический цикл обучения:
#
# * Итерирует по количеству эпох.
# * В каждую эпоху очищает дисплей и запускает `generate_images`, чтобы показать прогресс.
# * В каждую эпоху выполняет итерацию по набору тренировочных данных, печатая '.' для каждого примера.
# * Сохраняет контрольную точку каждые 20 эпох.
# + id="2M7LmLtGEMQJ"
def fit(train_ds, epochs, test_ds):
for epoch in range(epochs):
start = time.time()
display.clear_output(wait=True)
for example_input, example_target in test_ds.take(1):
generate_images(generator, example_input, example_target)
print("Epoch: ", epoch)
# Тренировка
for n, (input_image, target) in train_ds.enumerate():
print('.', end='')
if (n+1) % 100 == 0:
print()
train_step(input_image, target, epoch)
print()
# сохраняем чекпойнт каждые 20 эпох
if (epoch + 1) % 20 == 0:
checkpoint.save(file_prefix = checkpoint_prefix)
print ('Time taken for epoch {} is {} sec\n'.format(epoch + 1,
time.time()-start))
checkpoint.save(file_prefix = checkpoint_prefix)
# + [markdown] id="wozqyTh2wmCu"
# Этот цикл обучения сохраняет логи, которые вы можете просматривать в TensorBoard для отслеживания прогресса обучения. Работая локально, вы запускаете отдельный процесс TensorBoard. В ноутбуке, если вы хотите контролировать процесс с помощью TensorBoard, проще всего запустить программу просмотра перед началом обучения.
#
# Чтобы запустить программу просмотра, вставьте в ячейку кода следующее:
# + id="Ot22ujrlLhOd"
#docs_infra: no_execute
# %load_ext tensorboard
# %tensorboard --logdir {log_dir}
# + [markdown] id="Pe0-8Bzg22ox"
# Теперь запустите цикл обучения:
# + id="a1zZmKmvOH85"
fit(train_dataset, EPOCHS, test_dataset)
# + [markdown] id="oeq9sByu86-B"
# Если вы хотите опубликовать результаты TensorBoard _публично_, вы можете загрузить журналы в [TensorBoard.dev](https://tensorboard.dev/), скопировав и выполнив следующее в ячейку кода.
#
# Примечание. Для этого требуется учетная запись Google.
#
# ```
# # # !tensorboard dev upload --logdir {log_dir}
# ```
# + [markdown] id="l-kT7WHRKz-E"
# Внимание! Эта команда не завершается. Он предназначен для постоянной загрузки результатов длительных экспериментов. После того, как ваши данные загружены, вам необходимо остановить выполнение команды, выполнив "interrupt execution" в вашем инструменте для работы с ноутбуком(jupyter, colab, etc).
# + [markdown] id="-lGhS_LfwQoL"
# Вы можете просмотреть [результаты предыдущего запуска](https://tensorboard.dev/experiment/lZ0C6FONROaUMfjYkVyJqw) этого ноутбука на [TensorBoard.dev](https://tensorboard.dev/).
#
# TensorBoard.dev - это управляемый интерфейс для размещения, отслеживания и обмена экспериментами машинного обучения.
#
# Он также может быть включен с помощью `<iframe>`:
# + id="8IS4c93guQ8E"
display.IFrame(
src="https://tensorboard.dev/experiment/lZ0C6FONROaUMfjYkVyJqw",
width="100%",
height="1000px")
# + [markdown] id="DMTm4peo3cem"
# Интерпретация логов из GAN сложнее, чем интерпретация простой классификациии или регрессионной модели. На что следует обратить внимание:
#
# * Убедитесь, что ни одна из моделей не «выиграла». Если либо `gen_gan_loss`, либо `disc_loss` становятся очень низким, это показатель того, что одна модель доминирует над другой, и вы не обучаете комбинированную модель успешно.
# * Значение `log(2) = 0.69` является хорошим показателем для этих потерь, поскольку это значение указывает на неуверенность модели: дискриминатор в среднем одинаково неуверен в обоих вариантах.
# * Для `disc_loss` значение ниже `0.69` означает, что дискриминатор работает лучше, чем случайно, на объединенном наборе реальных + сгенерированных изображений.
# * Для `gen_gan_loss` значение ниже `0.69` означает, что генератор работает лучше, чем случайно, обманывая дескриминатор.
# * По мере обучения значение `gen_l1_loss` должно уменьшаться.
# + [markdown] id="kz80bY3aQ1VZ"
# ## Восстановление последнего чекпойнта и тестирование
# + id="HSSm4kfvJiqv"
# !ls {checkpoint_dir}
# + id="4t4x69adQ5xb"
# восстанавливаем последний чекпойнт из checkpoint_dir
checkpoint.restore(tf.train.latest_checkpoint(checkpoint_dir))
# + [markdown] id="1RGysMU_BZhx"
# ## Генерация с использованием тестового датасета
# + id="KUgSnmy2nqSP"
# Запускаем обученную модель на нескольких примерах из тестового набора данных
for inp, tar in test_dataset.take(5):
generate_images(generator, inp, tar)
|
site/ru/tutorials/generative/pix2pix.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:.conda-PythonData] *
# language: python
# name: conda-env-.conda-PythonData-py
# ---
import os
import pandas as pd
df = pd.read_csv("cities.csv")
df
df.to_html()
|
WebVisualizations/.ipynb_checkpoints/df_to_HTML-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
# Connect to Database
import os
database_url = os.environ["database_url2"]
verbose = True
from sqlalchemy import create_engine
from sqlalchemy.orm import sessionmaker
session = sessionmaker(bind=create_engine(database_url, echo=False), autoflush=False)()
# +
# Select Daten aus der stunden Datenbank
from sqlalchemy import func
ereignisse = ["K", "COR"]
zeitpunkt = "Monat Oktober 2021"
from kostunden.database.models import Mitarbeiterereignis, Mitarbeiter, Kostenstelle
q = session.query(Kostenstelle.kennung, func.count(Kostenstelle.kennung)).select_from(Mitarbeiterereignis).join(Mitarbeiter).join(Kostenstelle, Kostenstelle.id == Mitarbeiter.kostenstelle_id) \
.filter(Mitarbeiterereignis.datum >= '2021-10-01', Mitarbeiterereignis.datum <= '2021-10-31')\
.filter(Mitarbeiterereignis.ganztaegig.in_(ereignisse))\
.order_by(Kostenstelle.kennung)\
.group_by(Kostenstelle.kennung)
kostenstelle_ausfalltage = {d[0]: d[1] for d in list(q)}
if verbose:
print(kostenstelle_ausfalltage)
# +
# Select Daten aus der stunden Datenbank
from sqlalchemy import func
from kostunden.database.models import Mitarbeiterstunden
q = session.query(Kostenstelle.kennung, func.count(Kostenstelle.kennung)).select_from(Mitarbeiterstunden).join(Kostenstelle) \
.filter(Mitarbeiterstunden.datum >= '2021-10-01', Mitarbeiterstunden.datum <= '2021-10-31')\
.order_by(Kostenstelle.kennung) \
.group_by(Kostenstelle.kennung)
kostenstelle_ausfuehrtage = {d[0]: d[1] for d in list(q)}
if verbose:
print(kostenstelle_ausfuehrtage)
# +
# merge Listen Ausfalltage und Ausfühtage
q = session.query(Kostenstelle.kennung).order_by("kennung").all()
alle_kostenstellen = [k[0] for k in list(q)]
data = list()
for kostenstelle in alle_kostenstellen:
# data.append({"kostenstelle": kostenstelle, "ausfuehrtage": kostenstelle_ausfuehrtage.get(kostenstelle, 0), "ausfalltage": kostenstelle_ausfalltage.get(kostenstelle, 0)})
data.append({"Kostenstelle": kostenstelle, "Typ": "Ausführtage", "Anzahl": kostenstelle_ausfuehrtage.get(kostenstelle, 0)})
data.append({"Kostenstelle": kostenstelle, "Typ": "Ausfalltage", "Anzahl": kostenstelle_ausfalltage.get(kostenstelle, 0)})
if verbose:
print(data)
# +
# DataFrame erstellen
import pandas as pd
data_frame = pd.DataFrame(data)
if verbose:
print(data_frame)
# +
import numpy as np
def show_values(axs, orient="v", space=.01):
def _single(ax):
if orient == "v":
for p in ax.patches:
_x = p.get_x() + p.get_width() / 2
_y = p.get_y() + p.get_height() + (p.get_height()*0.01)
value = '{:.1f}'.format(p.get_height())
ax.text(_x, _y, value, ha="center")
elif orient == "h":
for p in ax.patches:
_x = p.get_x() + p.get_width() + float(space)
_y = p.get_y() + p.get_height() - (p.get_height()*0.5)
value = '{:.1f}'.format(p.get_width())
ax.text(_x, _y, value, ha="left")
if isinstance(axs, np.ndarray):
for idx, ax in np.ndenumerate(axs):
_single(ax)
else:
_single(axs)
# +
# Diagramm erstellen
import matplotlib.pyplot as plt
import seaborn as sn
sn.set(font_scale=1.0)
fig, ax = plt.subplots(figsize=(20,7))
palette = ['tab:green','tab:red']
p = sn.barplot(data = data_frame
,x = 'Kostenstelle'
,y = 'Anzahl'
,hue = 'Typ'
,palette=palette
)
ax.set_title(f"Ausfalltage im {zeitpunkt} wegen Krankheit ({', '.join(ereignisse)})", y=1, fontsize = 16)
show_values(p)
# -
|
notebooks/ausfalltage_wegen_krankheit.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 122} colab_type="code" id="VGneF6wedinh" outputId="293f24c8-3ff1-43f5-cac5-b8353c78db96"
import sys
sys.path[0] = '/tensorflow-2.1.0/python3.6'
from google.colab import drive
drive.mount('/drive')
# + colab={} colab_type="code" id="y1xKcH__dinq"
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers, Sequential, losses, optimizers, datasets
from tensorflow.keras.utils import to_categorical
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="vTHHiW0Ddinu" outputId="6863adda-22fd-429c-e9f5-1a22959c102f"
tf.__version__
# + colab={"base_uri": "https://localhost:8080/", "height": 85} colab_type="code" id="f2GOo6Apdinz" outputId="adcec94c-c38f-4263-e67f-0912c9f66001"
(x, y), (x_test, y_test) = datasets.cifar100.load_data()
print(x.shape,y.shape,x_test.shape,y_test.shape)
print(x.dtype,y.dtype)
# + colab={} colab_type="code" id="VTPgQhmrdin2"
def preprocess(x, y):
x = tf.cast(x, dtype=tf.float32) / 255.
y = tf.cast(to_categorical(tf.squeeze(tf.cast(y, dtype=tf.int32), axis=1), num_classes=100), dtype=tf.int32)
return x,y
# + colab={} colab_type="code" id="HAbJ7PgVdin5"
class BasicBlock(layers.Layer):
def __init__(self, filter_num, stride=1):
super(BasicBlock, self).__init__()
self.conv1 = layers.Conv2D(filter_num, (3, 3), strides=stride, padding='same')
self.bn1 = layers.BatchNormalization()
self.relu = layers.Activation('relu')
self.drop1 = layers.Dropout(0.5)
self.conv2 = layers.Conv2D(filter_num, (3, 3), strides=1, padding='same')
self.bn2 = layers.BatchNormalization()
self.drop2 = layers.Dropout(0.5)
if stride != 1:
self.downsample = layers.Conv2D(filter_num, (1, 1), strides=(stride, stride), padding='valid')
else:
self.downsample = lambda x:x
def call(self, inputs, training=None):
out = self.conv1(inputs)
out = self.bn1(out)
out = self.relu(out)
out = self.drop1(out)
out = self.conv2(out)
out = self.bn2(out)
out = self.drop2(out)
identity = self.downsample(inputs)
output = layers.add([identity, out])
output = tf.nn.relu(output)
return output
# + colab={} colab_type="code" id="uAm4qiA8din6"
class ResNet(keras.Model):
def __init__(self, layer_dims, num_classes=100):
super(ResNet, self).__init__()
self.stem = Sequential([layers.Conv2D(64, (3, 3), strides=(1, 1), padding='same'),
layers.BatchNormalization(),
layers.Activation('relu'),
layers.MaxPool2D(pool_size=(2, 2), strides=(1, 1), padding='same')
])
self.block1 = self.build_resblock(64, layer_dims[0])
self.block2 = self.build_resblock(128, layer_dims[1], stride=2)
self.block3 = self.build_resblock(256, layer_dims[2], stride=2)
self.block4 = self.build_resblock(512, layer_dims[3], stride=2)
self.avgpool = layers.GlobalAveragePooling2D()
self.fc = layers.Dense(num_classes, activation=tf.nn.softmax)
def call(self, inputs, training=None):
x = self.stem(inputs)
x = self.block1(x)
x = self.block2(x)
x = self.block3(x)
x = self.block4(x)
x = self.avgpool(x)
x = self.fc(x)
return x
def build_resblock(self, filter_num, blocks, stride=1):
res_blocks = Sequential()
res_blocks.add(BasicBlock(filter_num, stride))
for _ in range(1, blocks):
res_blocks.add(BasicBlock(filter_num, stride=1))
return res_blocks
# + colab={} colab_type="code" id="8RN82lU8din8"
def main(x, y, x_test, y_test):
epochs = 100
model = ResNet([2, 2, 2, 2])
model.build(input_shape=(None, 32, 32, 3))
model.summary()
save_best = keras.callbacks.ModelCheckpoint('/drive/My Drive/Github/CNN/ResNet_best_model.h5', monitor='val_loss', verbose=1, save_best_only=True, mode='min')
early_stop = keras.callbacks.EarlyStopping(monitor='val_loss', verbose=1, min_delta=0, patience=100, mode='auto')
callbacks_list = [early_stop, save_best]
model.compile(optimizer=optimizers.Adam(lr=1e-2),
loss=losses.categorical_crossentropy,
metrics=['accuracy'])
x, y = preprocess(x, y)
x_test, y_test = preprocess(x_test, y_test)
history = model.fit(x=x, y=y, epochs=epochs, batch_size=512, validation_data=(x_test, y_test), verbose=1, callbacks=callbacks_list)
return history
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="ESxyjSOwdin-" outputId="c81e4cb1-2e81-46f8-b30f-5eb5739e423d"
main(x, y, x_test, y_test)
# + colab={} colab_type="code" id="j0BGg9gfdioA"
|
Image Classification/ResNet18.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Extracting Titanic Disaster Data From Kaggle
# !pip install python-dotenv
from dotenv import load_dotenv, find_dotenv
# find .env automatically by walking up directories until it's found
dotenv_path = find_dotenv()
print(dotenv_path)
# load up the entries as environment variables
load_dotenv(dotenv_path, override=True)
# extracting environment variable using os.environ.get
import os
os.environ.get('KAGGLE_USERNAME')
# KAGGLE_USERNAME = os.environ.get("KAGGLE_PASSWORD")
# print(KAGGLE_USERNAME)
# imports
import requests
from requests import session
import os
from dotenv import load_dotenv, find_dotenv
# +
payload = {
'action': 'login',
'username': os.environ.get("KAGGLE_USERNAME"),
'password': <PASSWORD>("<PASSWORD>")
}
# url for train file (get the link from Kaggle website)
url = 'https://www.kaggle.com/c/titanic/download/train.csv'
# setup session
with session() as c:
# post request
c.post('https://www.kaggle.com/account/login', data=payload)
# get request
response = c.get(url)
# print response text
print(response.text)
# +
from requests import session
# payload
payload = {
'action': 'login',
'username': os.environ.get("KAGGLE_USERNAME"),
'password': <PASSWORD>("<PASSWORD>")
}
def extract_data(url, file_path):
'''
extract data from kaggle
'''
# setup session
with session() as c:
c.post('https://www.kaggle.com/account/login', data=payload)
# oppen file to write
with open(file_path, 'w') as handle:
response = c.get(url)
handle.write(response.text)
# +
# urls
train_url = 'https://www.kaggle.com/c/titanic/download/train.csv'
test_url = 'https://www.kaggle.com/c/titanic/download/test.csv'
# file paths
raw_data_path = os.path.join(os.path.pardir,'data','raw')
train_data_path = os.path.join(raw_data_path, 'train.csv')
test_data_path = os.path.join(raw_data_path, 'test.csv')
# extract data
extract_data(train_url,train_data_path)
extract_data(test_url,test_data_path)
# -
# !ls -l ../data/raw
# ### Builiding the file script
get_raw_data_script_file = os.path.join(os.path.pardir,'src','data','get_raw_data.py')
# +
# %%writefile $get_raw_data_script_file
# -*- coding: utf-8 -*-
import os
from dotenv import find_dotenv, load_dotenv
from requests import session
import logging
# payload for login to kaggle
payload = {
'action': 'login',
'username': os.environ.get("KAGGLE_USERNAME"),
'password': <PASSWORD>("<PASSWORD>")
}
def extract_data(url, file_path):
'''
method to extract data
'''
with session() as c:
c.post('https://www.kaggle.com/account/login', data=payload)
with open(file_path, 'w') as handle:
response = c.get(url, stream=True)
for block in response.iter_content(1024):
handle.write(block)
def main(project_dir):
'''
main method
'''
# get logger
logger = logging.getLogger(__name__)
logger.info('getting raw data')
# urls
train_url = 'https://www.kaggle.com/c/titanic/download/train.csv'
test_url = 'https://www.kaggle.com/c/titanic/download/test.csv'
# file paths
raw_data_path = os.path.join(project_dir,'data','raw')
train_data_path = os.path.join(raw_data_path, 'train.csv')
test_data_path = os.path.join(raw_data_path, 'test.csv')
# extract data
extract_data(train_url,train_data_path)
extract_data(test_url,test_data_path)
logger.info('downloaded raw training and test data')
if __name__ == '__main__':
# getting root directory
project_dir = os.path.join(os.path.dirname(__file__), os.pardir, os.pardir)
# setup logger
log_fmt = '%(asctime)s - %(name)s - %(levelname)s - %(message)s'
logging.basicConfig(level=logging.INFO, format=log_fmt)
# find .env automatically by walking up directories until it's found
dotenv_path = find_dotenv()
# load up the entries as environment variables
load_dotenv(dotenv_path)
# call the main
main(project_dir)
# -
# !python $get_raw_data_script_file
|
notebooks/1.0-ak-extract-titanic-data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <small><small><i>
# All the IPython Notebooks in Python Mini-Projects series by Dr. <NAME> are available @ **[GitHub](https://github.com/milaan9/91_Python_Mini_Projects)**
# </i></small></small>
# # Convert .py to .exe file format
#
# A **`.py`** to **`.exe`** converter using a simple graphical interface and **[PyInstaller](https://www.pyinstaller.org/)** in Python.
# ## Pre-requisites
#
# **Python : 3.5-3.9**
#
# To have the interface displayed in the images, you will need chrome. If chrome is not installed or --no-chrome is supplied, the default browser will be used.
#
# >**Note:** As of **[PyInstaller 4.0](https://github.com/pyinstaller/pyinstaller/releases/tag/v4.0)**, Python 2.7 is no longer supported. Read **[Python 2.7 Support](https://pypi.org/project/auto-py-to-exe/#python-27-support)** below for steps on how to use this tool with Python 2.7.
# ## Open your Anaconda Command Promt and type:
#
# **`pip install auto-py-to-exe`**
#
# Then to run it, execute the following in the terminal:
#
# **`auto-py-to-exe`**
# ## Using the Application
#
# 1. At first you will see empty console
# <br>
# <br>
# <div>
# <img src="img/py2exe1.png" width="500"/>
# </div>
# <br>
# <br>
# 1. Select your script location (paste in or use a file explorer)
# - Outline will become blue when file exists
# <br>
# <br>
# <div>
# <img src="img/py2exe2.png" width="500"/>
# </div>
# <br>
# <br>
# 2. Select other options and add things like an other files and output location
# <br>
# <br>
# <div>
# <img src="img/py2exe3.png" width="500"/>
# </div>
# <br>
# <br>
# 3. Click the big blue button at the bottom to convert
# <br>
# <br>
# <div>
# <img src="img/py2exe4.png" width="500"/>
# </div>
# <br>
# <br>
# 4. Find your converted files in /output when completed
# <br>
# <br>
# <div>
# <img src="img/py2exe5.png" width="500"/>
# </div>
|
003_Convert_.py_to_.exe/003_Convert_.py_to_.exe.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# this document is used to explain how to calculate document similarity using tf-idf method
#
#
documents = [
"we are doing just fine fine fine",
"we are doing just fine ",
]
# +
# from tfidf import tf_idf
# -
def tokenize(documents):
result = []
for document in documents:
result.append(document.split(" "))
print(result)
return result
tokens = tokenize(documents)
terms = []
for document in tokens:
for word in document:
if word not in terms:
terms.append(word)
terms
def calc_tf(terms,documents,method):
tf_vector = np.zeros((len(documents),len(terms)))
for d, document in enumerate(documents):
for t, term in enumerate(terms):
tf = method(term,document)
tf_vector.itemset((d,t),tf)
return tf_vector
# in `tf_raw` operation, we use `raw count` to calculate `tf`
# so that ${tf}_{(t,d)} = f_{(t,d)}$
def tf_raw(term, document):
tf = 0
for word in document:
if word == term:
tf += 1
return tf
# `tf_binary` tf is 1 if the term exist and 0 if term nonexistent on the document
def tf_binary(term,document):
return 1 if term in document else 0
# `tf_termfrequency`
#
# $\mathrm{tf} = \frac{f_{(t,d)}}{\displaystyle\sum_{{w\in d}}{f_{(w,d)}}}$
def tf_termfrequency (term,document):
tf = 0
for word in document:
if term == word:
tf += 1
return tf/len(document)
# after choosing which tf method to use, you can calculate tf this way:
def calc_idf (terms,documents,method):
idf_vector = np.zeros((1,len(terms)))
for t, term in enumerate(terms):
idf = method(term, documents)
idf_vector.itemset((0,t),idf)
return idf_vector
# $\mathrm{idf}_{(t,D)} = \log_{10}\frac{N}{\mathit{df}} $
def idf_norm(term,documents):
N = len(documents)
df = 0
for document in documents:
if term in document:
df += 1
print(f"np.log10({N}/{np.abs(df)})")
return np.log10(N/np.abs(df))
# there's some chance the term (`term in terms`) is not in the corpus (`documents`) so you adjust the equation to avoid division by zero as such
#
# $\mathrm{idf}_{(t,D)} = \log_{10}\frac{N}{\mathit{df}+1} $
def idf_smooth(term,documents):
df = 0
for document in documents:
if term in document:
df += 1
return np.log((len(documents)+1)/(np.abs(df)+1))
tf_vector = calc_tf(terms,tokens,tf_termfrequency)
print(tf_vector)
idf_vector = calc_idf(terms,documents,idf_norm)
print(idf_vector)
def calculate_tfidf(tf_vector, idf_vector):
tfidf_vector = np.ones(tf_vector.shape)
for i in range(tf_vector.shape[0]):
for j in range(tf_vector.shape[1]):
tfidf = tf_vector.item((i,j)) * idf_vector.item((0,j))
tfidf_vector.itemset((i,j),tfidf)
return tfidf_vector
tfidf_v = calculate_tfidf(tf_vector,idf_vector)
np.dot(tfidf_v,tfidf_v.T)
np.log10(100)
|
fltools/tfidf_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
x = np.array([1,2,3,4,5])
y = np.array([3,4,5,6,7])
x_ = x.mean()
y_ = y.mean()
print('mean x:',x_)
print('mean y:',y_)
tu_so = (x-x_)*(y-y_)
print(tu_so)
tu_so = tu_so.sum()
print('tu so:',tu_so)
mau_so = np.sqrt() ((x-x_)*(x-x_)).sum()*((y-y_)*(y-y_)).sum()
|
Notebook/Ex_Jupyter/he_so_tuong_quan/Untitled.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from sklearn.metrics import confusion_matrix, classification_report, accuracy_score
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.optim import Adam
from torch.utils.data import DataLoader,random_split
from torchvision import transforms
from torchvision.datasets import ImageFolder
# +
transform = transforms.Compose(
[transforms.Resize((50,50)),
transforms.ToTensor(),
transforms.Normalize((0.5, 0.5, 0.5), (0.5, 0.5, 0.5))])
def load_dataset_from_folder(batch_size,num_workers,shuffle=True):
path = 'data/train/'
all_data = ImageFolder(
root = path,
transform = transform
)
train_size = int(0.9 * len(all_data))
validation_size = len(all_data) - train_size
train_dataset, test_dataset = torch.utils.data.random_split(all_data, [train_size, validation_size])
training_data_loader = DataLoader(
train_dataset,
batch_size=batch_size,
num_workers=num_workers,
shuffle=shuffle
)
validation_dataset_loader = DataLoader(
test_dataset,
batch_size=batch_size,
num_workers=num_workers,
shuffle=shuffle
)
return training_data_loader,validation_dataset_loader
class Net(nn.Module):
fin_x = 0
fin_y = 0
image_x = 50 #36
image_y = 50 #41
k_size = 3
stride = 2
pool_size = 2
def __init__(self):
super(Net, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(3, 32, self.k_size, self.stride),
nn.ReLU(),
nn.MaxPool2d(self.pool_size, self.pool_size))
self.layer2 = nn.Sequential(
nn.Conv2d(32, 64, self.k_size, self.stride),
nn.ReLU(),
nn.MaxPool2d(self.pool_size, self.pool_size))
self.drop_out = nn.Dropout()
tmpx = int((int((self.image_x-self.k_size)/self.stride)+1)/self.pool_size)
tmpy = int((int((self.image_y-self.k_size)/self.stride)+1)/self.pool_size)
self.fin_x = int((int((tmpx-self.k_size)/self.stride)+1)/self.pool_size)
self.fin_y = int((int((tmpy-self.k_size)/self.stride)+1)/self.pool_size)
self.fc1 = nn.Sequential(
nn.Linear(64 * self.fin_x * self.fin_y, 2048),
nn.ReLU()
)
self.fc2 = nn.Sequential(
nn.Linear(2048, 4),
nn.ReLU()
)
self.sof = nn.LogSoftmax()
def forward(self, x):
x = self.layer1(x)
x = self.layer2(x)
x = x.view(-1, 64 * self.fin_x * self.fin_y)
x = self.drop_out(x)
x = self.fc1(x)
x = self.fc2(x)
x = self.sof(x)
return x
# net = Net()
# net = net.cuda()
# +
model = Net()
# defining the optimizer
optimizer = Adam(model.parameters(), lr=0.07)
# defining the loss function
criterion = nn.CrossEntropyLoss()
# checking if GPU is available
if torch.cuda.is_available():
print('Found Cuda')
model = model.cuda()
criterion = criterion.cuda()
print(model)
# criterion = nn.NLLLoss()
# optimizer = optim.Adam(net.parameters(), lr=0.00001)
# +
batch_size = 100
shuffle = True
num_workers = 4
train_generator,test_generator = load_dataset_from_folder(batch_size,num_workers,shuffle)
# train_set = my_dataloader('test_x_0_100_rgb.npy', 'test_y_0_100_rgb.npy')
# train_generator = DataLoader(train_set, batch_size=batch_size, shuffle=True, num_workers=4)
# -
print(len(train_generator.dataset))
print(len(test_generator.dataset))
for epoch in range(10):
running_loss = 0.0
validation_loss = 0.0
for step,(x,y) in enumerate(train_generator):
# print(x.shape)
optimizer.zero_grad()
outputs = model(x)
loss = criterion(outputs, y)
loss.backward()
optimizer.step()
running_loss += loss.item()
print('[%d] Training loss: %.3f' %(epoch + 1, running_loss))
with torch.no_grad():
for step,(x,y) in enumerate(test_generator):
outputs = model(x)
loss = criterion(outputs, y)
validation_loss += loss.item()
print('[%d] Validation loss: %.3f' %(epoch + 1, validation_loss))
# +
# print(train_set[0])
for x,y in train_generator:
print(x.shape)
preds_Test = torch.argmax(model(x),1)
test_examples = x.numpy()
gold = y.numpy()
predictions = preds_Test.numpy()
break
# print(preds_Test)
# -
# test_result_np=test_result.numpy()
# print(preds_Test.numpy())
print("----------Test Accuracy-----------")
print(accuracy_score(gold, predictions))
print(gold,predictions)
print(confusion_matrix(gold, predictions))
print(classification_report(gold, predictions))
# +
# with torch.no_grad():
# output = model(train_x)
# softmax = torch.exp(output).cpu()
# prob = list(softmax.numpy())
# predictions = np.argmax(prob, axis=1)
# # accuracy on training set
# accuracy_score(train_y, predictions)
# +
# # prediction for validation set
# with torch.no_grad():
# output = model(val_x)
# softmax = torch.exp(output).cpu()
# prob = list(softmax.numpy())
# predictions = np.argmax(prob, axis=1)
# # accuracy on validation set
# accuracy_score(val_y, predictions)
# +
# # loading test images
# test_img = []
# for img_name in tqdm(test['id']):
# # defining the image path
# image_path = 'test_ScVgIM0/test/' + str(img_name) + '.png'
# # reading the image
# img = imread(image_path, as_gray=True)
# # normalizing the pixel values
# img /= 255.0
# # converting the type of pixel to float 32
# img = img.astype('float32')
# # appending the image into the list
# test_img.append(img)
# # converting the list to numpy array
# test_x = np.array(test_img)
# test_x.shape
# +
# # converting training images into torch format
# test_x = test_x.reshape(10000, 1, 28, 28)
# test_x = torch.from_numpy(test_x)
# test_x.shape
# +
# # generating predictions for test set
# with torch.no_grad():
# output = model(test_x)
# softmax = torch.exp(output).cpu()
# prob = list(softmax.numpy())
# predictions = np.argmax(prob, axis=1)
# -
|
A4/old_code/ConvNet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + inputHidden=false outputHidden=false
# !pip install --upgrade altair vega_datasets
# +
## jakevdp recommended install https://jakevdp.github.io/blog/2017/12/05/installing-python-packages-from-jupyter/
# import sys
# # !{sys.executable} -m pip install --upgrade altair vega_datasets
# -
# + inputHidden=false outputHidden=false
import altair as alt
from vega_datasets import data
cars = data.cars()
# -
# # Faceted Scatter Plot with Linked Brushing
#
# This is an example of using an interval selection to control the color of points across multiple facets.
# +
brush = alt.selection(type='interval', resolve='global')
base = alt.Chart(cars).mark_point().encode(
y='Miles_per_Gallon',
color=alt.condition(brush, 'Origin', alt.ColorValue('gray'))
).add_selection(
brush
).properties(
width=250,
height=250
)
print("Select a region in the chart below to try this out!")
base.encode(x='Horsepower') | base.encode(x='Acceleration')
# + inputHidden=false outputHidden=false
|
python/altair.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# <div class="contentcontainer med left" style="margin-left: -50px;">
# <dl class="dl-horizontal">
# <dt>Title</dt> <dd>DynamicMap Container</dd>
# <dt>Dependencies</dt> <dd>Bokeh</dd>
# <dt>Backends</dt> <dd><a href='./DynamicMap.ipynb'>Bokeh</a></dd> <dd><a href='../matplotlib/DynamicMap.ipynb'>Matplotlib</a></dd> <dd><a href='../plotly/DynamicMap.ipynb'>Plotly</a></dd>
# </dl>
# </div>
import numpy as np
import holoviews as hv
hv.extension('bokeh')
# A DynamicMap is an explorable multi-dimensional wrapper around a callable that returns HoloViews objects. A ``DynamicMap`` callable cannot return ``Layouts``, ``NdLayouts``, ``GridSpaces`` or other ``DynamicMaps`` or ``HoloMaps`` but can contain any other HoloViews object. See the [Building Composite Objects](../../../user_guide/06-Building_Composite_Objects.ipynb) user guide for details on how to compose containers and for the user-guide describing ``DynamicMap`` in more detail, see the [Live Data](../../../user_guide/07-Live_Data.ipynb) user guide.
#
#
# <p><center><div class="alert alert-info" role="alert"><b>Note: </b>To work with live data, you need a live Python server, not a static web site, which is why the outputs shown below are GIF animations. If you run this notebook yourself, you will be able to try out your own interactions and compare them to the displayed GIF animations.</div></center></p>
# ### ``DynamicMap`` holds callables
# Although a ``DynamicMap`` holds a user supplied callable, this can be seen as as a generalization of [``HoloMap``](./HoloMap.ipynb) which holds dictionaries of elements: the key is then conceptually the arguments to the callable and the value is the object the callable returns. This conceptual model assume the callable is a true function where a set of arguments always maps to the same output, no matter how many times it is called.
#
# For [``HoloMap``](./HoloMap.ipynb), we used the ``sine_curve`` function below to generate a dictionary of ``Curve`` objects. With ``DynamicMap``, we can use it directly:
# +
def sine_curve(phase, freq):
xvals = [0.1* i for i in range(100)]
return hv.Curve((xvals, [np.sin(phase+freq*x) for x in xvals]))
# When run live, this cell's output should match the behavior of the GIF below
dmap = hv.DynamicMap(sine_curve, kdims=['phase', 'frequency'])
dmap.redim.range(phase=(0.5,1), frequency=(0.5,1.25))
# -
# <img src='https://s3-eu-west-1.amazonaws.com/assets.holoviews.org/gifs/examples/containers/bokeh/DynamicMap.gif'>
# Unlike a ``HoloMap`` which is limited by the static number of items in the supplied dictionary (which must all exist in memory at once), this ``DynamicMap`` lets you pick any phase or frequency within the supplied range.
#
# Although ``DynamicMap`` is designed as the dynamic counterpart of [``HoloMap``](./HoloMap.ipynb), the fact that it accepts a code specification as opposed to data opens up a large set of new possibilities. The [Live Data](../../../user_guide/07-Live_Data.ipynb) user guide is dedicated to exploring what can be done with ``DynamicMap``.
|
examples/reference/containers/bokeh/DynamicMap.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import packages
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# %matplotlib inline
# set up variables
delta_x = 25/335 #(nm)
delta_t = 0.016667 #(s)
length = 25 #(nm)
finaltime = 0.016667*335 #(s)
# import data
df = pd.read_excel('D.xlsx', sheet_name='2')
df.loc[0]
# position we choose
points_number = int(length/delta_x+1)
position = np.linspace(0,length,points_number)
D_point_new = []
for i in range(1,len(df)-1):
new_D = (df.loc[i+1]-df.loc[i])/(df.loc[i+1]+df.loc[i-1]-2*df.loc[i])*(delta_x**2)/delta_t
D_point_new.append(new_D)
D_point_new
|
ode_pve/Calculate D.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # for Mac OS
import os
os.environ['KMP_DUPLICATE_LIB_OK']='True'
# +
import math
import random
import gym
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import torch.nn.functional as F
from torch.distributions import Categorical
# -
from IPython.display import clear_output
import matplotlib.pyplot as plt
# %matplotlib inline
use_cuda = torch.cuda.is_available()
device = torch.device("cuda" if use_cuda else "cpu")
use_cuda
# ## Create Environments
# +
from multiprocessing_env import SubprocVecEnv
num_envs = 12
env_name = "CartPole-v0"
def make_env():
def _thunk():
env = gym.make(env_name)
return env
return _thunk
envs = [make_env() for i in range(num_envs)]
envs = SubprocVecEnv(envs)
env = gym.make(env_name)
# +
def plot(frame_idx, rewards):
clear_output(True)
plt.figure(figsize=(20,5))
plt.subplot(131)
plt.title('frame %s. reward: %s' % (frame_idx, rewards[-1]))
plt.plot(rewards)
plt.show()
def test_env(vis=False):
state = env.reset()
if vis: env.render()
done = False
total_reward = 0
while not done:
state = torch.FloatTensor(state).unsqueeze(0).to(device)
dist, _ = model(state)
next_state, reward, done, _ = env.step(dist.sample().cpu().numpy()[0])
state = next_state
if vis: env.render()
total_reward += reward
env.close()
return total_reward
# -
# ## Neural Network
class ActorCritic(nn.Module):
def __init__(self, num_inputs, num_outputs, hidden_size, std=0.0):
super(ActorCritic, self).__init__()
self.critic = nn.Sequential(
nn.Linear(num_inputs, hidden_size),
nn.ReLU(),
nn.Linear(hidden_size, 1)
)
self.actor = nn.Sequential(
nn.Linear(num_inputs, hidden_size),
nn.ReLU(),
nn.Linear(hidden_size, num_outputs),
nn.Softmax(dim=1),
)
def forward(self, x):
#x = torch.FloatTensor(x).unsqueeze(0)
value = self.critic(x)
#x = torch.FloatTensor(x).unsqueeze(0)
probs = self.actor(x)
dist = Categorical(probs)
dist.sample()
return dist, value
# ## A2C: Synchronous Advantage Actor Critic
#
# [OpenAI Blog:]("https://blog.openai.com/baselines-acktr-a2c/#a2canda3c\")
#
# The Asynchronous Advantage Actor Critic method (A3C) has been very influential since the paper was published. The algorithm combines a few key ideas:
#
# - An updating scheme that operates on fixed-length segments of experience (say, 20 timesteps) and uses these segments to compute estimators of the returns and advantage function.
# - Architectures that share layers between the policy and value function.
# - Asynchronous updates.
#
# After reading the paper, AI researchers wondered whether the asynchrony led to improved performance (e.g. “perhaps the added noise would provide some regularization or exploration?“), or if it was just an implementation detail that allowed for faster training with a CPU-based implementation.
#
# As an alternative to the asynchronous implementation, researchers found you can write a synchronous, deterministic implementation that waits for each actor to finish its segment of experience before performing an update, averaging over all of the actors. One advantage of this method is that it can more effectively use of GPUs, which perform best with large batch sizes. This algorithm is naturally called A2C, short for advantage actor critic. (This term has been used in several papers.)
#
#
def compute_returns(next_value, rewards, masks, gamma=0.99):
R = next_value
returns = []
for step in reversed(range(len(rewards))):
R = rewards[step] + gamma * R * masks[step]
returns.insert(0, R)
return returns
# +
num_inputs = envs.observation_space.shape[0]
num_outputs = envs.action_space.n
#Hyper params:
hidden_size = 256 # "hidden_size = 256"
lr = 3e-4
num_steps = 5
model = ActorCritic(num_inputs, num_outputs, hidden_size).to(device)
optimizer = optim.Adam(model.parameters())
# -
max_frames = 20000
frame_idx = 0
test_rewards = []
# +
state = envs.reset()
# for episode in range(num_episodes):
# done = False
# #state = env.reset()
# state = envs.reset()
# epi_reward = 0.
while frame_idx < max_frames:
log_probs = []
values = []
rewards = []
masks = []
entropy = 0
for _ in range(num_steps):
state = torch.FloatTensor(state).to(device)
dist, value = model(state)
action = dist.sample()
next_state, reward, done, _ = envs.step(action.cpu().numpy())
#action = dist.sample().data.numpy()[0]
#next_state, reward, done, _ = env.step(action)
log_prob = dist.log_prob(action)
entropy += dist.entropy().mean()
log_probs.append(log_prob)
values.append(value)
rewards.append(torch.FloatTensor(reward).unsqueeze(1).to(device))
masks.append(torch.FloatTensor(1 - done).unsqueeze(1).to(device))
state = next_state
frame_idx += 1
if frame_idx % 1000 == 0:
test_rewards.append(np.mean([test_env() for _ in range(10)]))
plot(frame_idx, test_rewards)
next_state = torch.FloatTensor(next_state).to(device)
_, next_value = model(next_state)
returns = compute_returns(next_value, rewards, masks)
log_probs = torch.cat(log_probs)
returns = torch.cat(returns).detach()
values = torch.cat(values)
advantage = returns - values
actor_loss = -(log_probs * advantage.detach()).mean()
critic_loss = advantage.pow(2).mean()
loss = actor_loss + 0.5 * critic_loss - 0.001 * entropy
optimizer.zero_grad()
loss.backward()
optimizer.step()
# -
print(test_env(True))
|
a2c-cartpolev0/a2c-cartpolev0.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# This is a slap-dash attempt at reviewing all of CMPSC131 in a single notebook. Much will be skipped, but that is ok. We can add it later! Let me know what is missing and it will be ammended.
#
# The list of topics covered in CMPSC131 include:
# 1. Variables and Datatypes
# 2. Lists
# 3. Conditional Statements
# 4. Dictionaries
# 5. User Input and Loops
# 6. Functions
# 7. Classes
# 8. Files and Exceptions
# # 1. Variables and Datatypes
msg = "What a review notebook!"
print(msg)
# Python is a duck typed language.This means that the Python interpretter finds the suitable datatype for the variable you define based on its use.
#
# Some rules about variables are:
# * names can contain only letters, numbers, and underscores. They can not start with a number
# * no spaces in variable names
# * do not use reserved words (i.e. for, in, not, and ...)
# * use short and descriptive names
#
# What is the datatype of the variable 'msg'?
#
# HINT: type(msg)
type(msg)
# The string sata type has several built in utilities that can prove useful.
print(msg.title())
print(msg.upper())
print(msg.lower())
print(msg.split())
print(msg.rjust(57))
# Python has several data types for handling numerical values.
# * $\mathbb{C}$ Complex
# * $\mathbb{R}$ Real $\rightarrow$ called floating point
# * $\mathbb{Z}$ Integers
a = 4 #integer
b = 3. #real
print(a + b) #you can add
print(a - b) #or subtract
print(a / b) #divide
print(a // b) #integer division
print(a % b) #or get the modulus (remainder)
# ### All of above output was type cast to floating point. Python determined that the operation should be all floating point, because b was a float. You can type cast variables back to integers.
a = -2
int(a/b)
# # 2. LISTS
#
# A list is a collection of items in a particular order. You can access specific members of a list with an index.
#
# Initialization of a list is done with \[\] square brackets.
coffeeTypes = ['espresso', 'drip', 'pour over']
coffeeTypes[-1] #Start counting at zero
# Items can be added to or remove from a list also.
coffeeTypes.append('aeropress')
coffeeTypes.insert(2,'cold brew')
[print(coffee.ljust(15) +str(' index = %1d' %index).rjust(12)) for index,coffee in enumerate(coffeeTypes)]
print(*coffeeTypes)
# Items can be removed from a list in a few ways.
# * Delete by index
# * remove method - built in
del coffeeTypes[1] #let us remove drip
coffeeTypes.remove('aeropress')
print(*coffeeTypes) #or dump contents of the list this way
popCoffee = coffeeTypes.pop()
print(popCoffee)
# The pop command pops the last item from the list according to index, not insertion order. This is not a LIFO stack.
#
# You can also pop on an index. _.pop(1)
#
# The list can also be sorted. Permanently with the built in sort method or temporarily with sorted method.
socks = ['tube', 'no show', 'crew', 'weird toe socks', 'ankle', ]
print(socks)
print(sorted(socks))
print(socks)
socks.sort()
print(socks)
socks.reverse()
print(socks)
print(socks[-1::-1])
n = len(socks)
print(n)
socks[n] #why is this?
print(socks[-1]) #access last member
print(socks[-1::-1]) #start at end and then move to front
print(socks[-1::-2]) #start at end and move to front by 2
print(socks[2:]) #start at 3rd item
# The range function has a ton of functionality for making lists of values between values.
#
# range(start, stop, stride)
#
# Because of zero-indexing the range will actually stop at stop -1
for x in range(0,4,1):
print(x)
print('--------')
#Equivalent Statement
for x in range(4): #default start is zero and stride is 1
print(x)
#Type cast a range to a list
evens = list(range(2,11,2))
print(evens)
squareRoots = []
for x in range(7):
squareRoots.append(x**0.5)
sRs = [x**0.5 for x in range(7)] #bad variable name, not descriptive
print(squareRoots)
print(sRs)
# # Conditional Statements
#
# The standards are all here:
# * if
# * if else
# * if else if (elif) else
#
# Below is a simple primality test for the some number n
n = 11
isPrime = True
if n <= 3:
isPrime = n > 1
elif n % 2 == 0 or n % 3 == 0:
isPrime = False
i = 5
while i ** 2 <= n:
if n % i == 0 or n % (i + 2) == 0:
isPrime = False
i += 6
print(isPrime)
# The above is ok, but we can easily turn it into a function to test many values.
#
# Let us make a dictionary of values and list if they are a prime or not.
def is_prime(n: int) -> str:
"""Primality test using 6k+-1 optimization."""
if n == 1:
return 'not prime'
elif n==2 or n==3:
return 'Prime!'
elif n % 2 == 0 or n % 3 == 0:
return 'not prime'
i = 5
while i ** 2 <= n:
if n % i == 0 or n % (i + 2) == 0:
return 'not prime'
i += 6
return 'Prime!'
primeDict = {}
for x in range(1,1000000):
primeDict[is_prime(x)]=x
primeDict #our dictionary isn't well defined is it, only saves the last value
# Wow! What a terrible example for a dictionary. For each key we have a single value. A different data structure would be better here. I can think of a whole bunch, but let use think of a better set of data to represent as a dictionary.
sandwiches = {} #define an empty dictionary with braces
# define a dictionary with key:value format and delimit entries with commas
sandwiches = {'hoagie':'long roll',
'hamburger':'Martin\'s Potato Roll',
'hotdog':'not a sandwich'}
# Failure to define a dictionary
webstersDict = {(1, 2.0): 'tuples can be keys',
1: 'ints can be keys',
'run': 'strings can be keys',
['sock', 1, 2.0]: 'lists can NOT be keys'}
a = 3
print(a*3)
# # nate
#
# thats my name
#
# $H_0^1(kr) $
#
# 
# + active=""
#
|
CMPSC131 Review.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# +
import os
import h5py
import subprocess
import numpy as np
from astropy.io import fits
from pydl.pydlutils.spheregroup import spherematch
from feasibgs import util as UT
from feasibgs import catalogs as Cat
# -
import matplotlib as mpl
import matplotlib.pyplot as pl
mpl.rcParams['text.usetex'] = True
mpl.rcParams['font.family'] = 'serif'
mpl.rcParams['axes.linewidth'] = 1.5
mpl.rcParams['axes.xmargin'] = 1
mpl.rcParams['xtick.labelsize'] = 'x-large'
mpl.rcParams['xtick.major.size'] = 5
mpl.rcParams['xtick.major.width'] = 1.5
mpl.rcParams['ytick.labelsize'] = 'x-large'
mpl.rcParams['ytick.major.size'] = 5
mpl.rcParams['ytick.major.width'] = 1.5
mpl.rcParams['legend.frameon'] = False
# %matplotlib inline
# Read in GAMA-Legacy catalog
# read in GAMA-Legacy catalog
cata = Cat.GamaLegacy()
gleg = cata.Read()
cataid = gleg['gama-photo']['cataid'] # GAMA catalog id of each object
ngal = len(cataid)
print('%i galaxies with GAMA-Legacy catalog' % ngal)
#
f_gama_sdss = fits.open(''.join([UT.dat_dir(), 'gama/ExternalSpecAll.fits']))
fdata = f_gama_sdss[1].data
fdata.names
print fdata.field('SPECID')[fdata.field("SURVEY") == 'SDSS'][:10]
print fdata.field('RA')[fdata.field("SURVEY") == 'SDSS'][:10]
print fdata.field('Dec')[fdata.field("SURVEY") == 'SDSS'][:10]
print fdata.field('Z')[fdata.field("SURVEY") == 'SDSS'][:10]
# Read in SDSS specObj
f_sdss = h5py.File(''.join([UT.dat_dir(), 'sdss/specObj-dr8.hdf5']), 'r')
ra_sdss = f_sdss['plug_ra'].value
dec_sdss = f_sdss['plug_dec'].value
print('%i galaxies with DR8 specObj' % len(ra_sdss))
f_sdss['z'].value.min(), f_sdss['z'].value.max()
m_sdss, m_gleg, d_match = spherematch(ra_sdss, dec_sdss,
gleg['gama-photo']['ra'], gleg['gama-photo']['dec'], 3*0.000277778)
print d_match[:10]
print('%i matching galaxies' % len(m_sdss))
fig = plt.figure(figsize=(10,10))
sub = fig.add_subplot(111)
sub.scatter(ra_sdss, dec_sdss, c='k', s=5, label='SDSS DR8')
sub.scatter(gleg['gama-photo']['ra'], gleg['gama-photo']['dec'], c='C1', s=4, label='GAMA')
sub.set_xlabel('RA', fontsize=30)
sub.set_xlim([179., 181.])
sub.set_ylabel('Dec', fontsize=30)
sub.set_ylim([-1., 1.])
sub.legend(loc='upper left', markerscale=5, prop={'size':20})
|
notebook/notes_gama_sdss1and2_emline.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Train A Smartcab to Drive
# <NAME>
#
# 
#
# This project forks [Udacity's Machine Learning Nanodegree Smartcab project](https://github.com/udacity/machine-learning/tree/master/projects/smartcab) with my solution, modifying/adding `smartcab/agent.py` and `smartcab/notebookhelpers.py` as well as this `README`.
# ### Overall summary of the final agent learning algorithm:
# In order to build a reinforcement learning agent to solve this problem, I ended up implementing $Q$ learning from the transitions. In class, we covered $\epsilon$-greedy exploration, where we selected the optimal action based on $Q$ with some probability 1 - $\epsilon$ and randomly otherwise. This obviously puts more weight on the current optimal strategy, but I wanted to put more or less weight on more or less suboptimal strategies as well. I did this by sampling actions in a simualted annealing fashion, assigning actions softmax probabilities of being sampled using the current $Q$ value with a decaying temperature. Further, each $Q(s, a_i)$ value is updated based on it's own exponentially decaying learning rate: $\alpha(s, a_i)$. The current temperature, $T(s)$, is defined as the mean of the decaying $\alpha(s, a)$ over all actions such that:
#
# $$T(s) = \frac{1}{n}\sum_{i=0}^{n}{\alpha(s', a_j')}$$
# $$P(a_i|Q,s) = \frac{e^{Q(s, a_i) / T(s)}}{\sum_{i=0}^{n}{e^{Q(s, a_i) / T(s)}}}$$
#
# Once the action for exploration, $a_i$, is sampled, the algorithm realizes a reward, $R(s, a_i)$, and new state, $s'$. I then update $Q$ using the action that maximizes Q for the new state. The update equations for $Q$ and $\alpha(s, a_i)$ are below:
# $$Q_{t+1}(s, a_i) = (1 - \alpha_t(s, a_i))Q_t(s, a_i) + \alpha_t(s, a_i)[R(s, a_i) + 0.05 \max_{a'}{Q_t(s', a')}]$$
# $$\alpha_t(s, a_i) = 0.5(\alpha(s, a_i) - 0.05) + 0.05$$
#
# and initially:
# $$Q_{0}(s, a_i) = 0$$
# $$\alpha_{0}(s, a_i) = 1.0$$
#
# Note that while $\alpha(s, a_i)$ is decaying at each update, it hits a minimum of 0.05 (thus it never quits learning fully). Also, I chose a very low $\gamma=0.05$ here to discount the next maximum $Q$.
#
# In terms of my state space, I use the following:
# - waypoint: {left, right, forward}
# - light: {green, red}
# - oncoming: {None, left, right, forward}
# - left: {True, False}
# - right: {True, False}
# ### Before implementing Q-Learning, did the smartcab eventually make it to the target location?
#
# When randomly selecting actions, it's very literally acting out a random walk. It's worthing noting that on a 2D lattice, it's been proven that a random-walking agent will almost surely reach any point as the number of steps approaches infinity (McCrea Whipple, 1940). In other words, it will almost surely make it to the target location, especially because this 2D grid also has a finite number of points.
# ### Justification behind the state space, and how it models the agent and environment.
#
# I picked the state space mentioned above based on features I believed mattered to the optimal solution. The waypoint effectively proxies the shortest path, and the light generally signals whether None is the right action. These two features alone should be sufficient to get a fairly good accuracy, though I did not test it. Further, I added traffic because this information can help optimize certain actions. For example, you can turn right on red conditional on no traffic from the left. You can turn left on green conditional on no oncoming traffic.
#
# I did not include the deadline here because we are incentivized to either follow the waypoint or stop to avoid something illegal. If we were learning our own waypoint based on the header, the deadline may be useful as a boolean feature once we’re close. Perhaps this would signal whether or not it would be efficient to take a right turn on red. Again, the deadline doesn’t help much given the game rewards.
#
# I also compressed left/right which previously could be {None, left, right, forward} based on the other agents signals. Now they are True/False based on whether or not cars existed left/right. You could also likely compress the state space conditional on a red light, where only traffic on the left matters. I strayed from this approach as it involved too much hard coding for rules the Reinforcement Learner could learn with sufficient exploration.
#
# There are only 96 unique states. Assuming each trial runs at least 5 steps, 100 trials views at least 500 states. Estimating the probability that each state will be seen here is tough since each state has a different probability of being picked based on the unknown true state distribution. Assuming the chance a state is picked is uniform, this becomes the Coupon Collector’s problem, where the expected number of trials, $T$, until $k$ coupons are collected out of a total of $n$ is:
#
# $$E[T_{n,k}] = n \sum_{i=n-k}^{n}\frac{1}{i}$$
#
# We can see below that assuming states are drawn uniformly, we’d expect to see all of the states after about 500 runs, and about 90% after only 250 runs:
# +
import numpy as np
import pandas as pd
import seaborn as sns
import pylab
# %matplotlib inline
def expected_trials(total_states):
n_drawn = np.arange(1, total_states)
return pd.Series(
total_states * np.cumsum(1. / n_drawn[::-1]),
n_drawn
)
expected_trials(96).plot(
title='Expected number of trials until $k$ distinct states are seen',
figsize=(15, 10))
_ = pylab.xlabel('$k$ (# of states seen)')
# -
# Obviously, states are not drawn uniformly, but rather based on the simulated distribution with 3 dummy cars. Thus we’re more likely to have sampled the most likely states, and the missing states are less likely to be encountered later than if we had drawn states uniformly. In a production environment, I would make sure I run this until every possible state has been seen a sufficient number of times (potentially through stratification). For this project, I think seeing around 500 states is sufficient, and thus 100 trials should train a fairly reasonable agent.
# ### Changes in agent behavior after implementing Q-Learning
#
# Initially after training the agent, it would consistently approach the destination, but would take very odd paths. For example, on a red light, it would commonly take a right turn, perhaps optimistic the next intersection would allow for a left turn on green, despite the penalty for disobeying the waypoint. I found this was due to my gamma being extremely high (0.95). Overall I was ultimately weighting the future rewards much more than the current penalties and rewards for taking each correct turn. The resulting agent somewhat ignored the waypoint and took it’s own optimal course, likely based on the fact that right turns on red just tend to be optimal. I think it’s reasonable that over time, the agent would have learned to follow the waypoint, assuming it’s the most efficient way to the destination, and perhaps the high gamma was causing slow convergence. It’s also possible the agent, weighting the final outcomes higher, found a more optimal waypoint to the end goal (ignoring illegal and waypoint penalties), but I think this is unlikely.
#
# During training, the agent would occasionally pick a suboptimal action (based on Q). Usually this was akin to taking a legal right turn on red, when the waypoint wanted to wait and go forward. This was done to ensure the agent sufficiently explored the state space. If I simply picked the action corresponding to the maximum $Q$ value, the agent would likely get stuck in a local optima. Instead the randomness allows it to eventually converge to a global optima.
#
# To visualize, the success rate while the initial $Q$-Learning model is shown below, followed by that same agent (now learned) using the optimal policy only:
# +
from smartcab.notebookhelpers import generated_sim_stats
def plot_cumulative_success_rate(stats, sim_type, ax=None):
columns = [
'always_reached_destination',
'reached_destination',
'missed_destination',
]
stats[columns].cumsum().plot(
ax=ax, kind='area', stacked=False,
title='%s Success Rate Over Trials: %.2f%%' % (
sim_type, stats.reached_destination.mean()*100
)
)
pylab.xlabel('Trial#')
def train_test_plots(train_stats, test_stats, plot):
_, (top, bottom) = pylab.subplots(
2, sharex=True, sharey=True, figsize=(15, 12))
plot(train_stats, 'Train', ax=top)
plot(test_stats, 'Test', ax=bottom)
# Generate training, and test simulations
learned_agent_env, train_stats = generated_sim_stats(
n_trials=100,
gamma=0.95,
alpha_span=100,
min_alpha=0.05,
initial_alpha=0.2,
)
_, test_stats = generated_sim_stats(
agent_env=learned_agent_env, n_trials=100)
train_test_plots(train_stats, test_stats,
plot_cumulative_success_rate)
# -
# What I noticed here was that the train performance was very similiar to the test performance in terms of the success rate. My intuition is that this is mainly due to the high gamma, which results in Q values that are slow to converge. Finally my $\alpha$ were decaying fairly slowly due to a span of 100, this caused my temperatures to stay high and randomly sample many suboptimal actions. Combined, this exacerbated bad estimates of Q values, which caused the test run to fail to significantly improve the overall success rate.
#
# However, I did find that the test run was much safer after taking a look at the cumulative trips with crimes, thus it was learning:
# +
def plot_cumulative_crimes(stats, sim_type, ax=None):
(stats['crimes'.split()] > 0).cumsum().plot(
ax=ax, kind='area', stacked=False, figsize=(15, 8),
title='Cumulative %s Trials With Any Crimes: %.0f%%' % (
sim_type, (stats.crimes > 0).mean()*100
)
)
pylab.ylabel('# of Crimes')
pylab.xlabel('Trial#')
train_test_plots(train_stats, test_stats,
plot_cumulative_crimes)
# -
# ### Updates to the final agent and final performance
# Generate training, and test simulations
learned_agent_env, train_stats = generated_sim_stats(
n_trials=100,
gamma=0.05,
initial_alpha=1.0,
min_alpha=0.05,
alpha_span=2.0,
)
_, test_stats = generated_sim_stats(
agent_env=learned_agent_env, n_trials=100)
# I made two major changes (as shown in the code above), based on my observations of the initial agent. First, I reduced the $\gamma$ all the way to 0.05 from 0.95. This caused my agent to pay much more attention to the current correct turn, and less on the final goal. This also means I can set a much larger initial $\alpha$ value since a majority of the new value is now deterministic (the reward).
#
# Another key observation I made was that optimal moves were deterministic based on the state. In order to exploit this in the learner, I considered the following cases:
#
# 1. Reward of 12:
# - This is assigned when the car makes the correct move to the destination (not illegal or suboptimal).
# 2. Reward of 9.5:
# - This is assigned when the car makes an incorrect move to the destination (perhaps teleporting from one side to the other)
# - I map this to -0.5
# 3. Reward of 9:
# - This is assigned when the car makes an illegal move to the destination
# - I map this to -1
# 4. Reward of 2:
# - This is assigned when the car legally follows the waypoint
# 5. Reward of 0:
# - This is assigned when the car stops
# 6. Reward of -0.5:
# - This is assigned when the car makes a suboptimal but legal move (doesn't follow waypoint)
# 7. Reward of -1:
# - This is assigned when the car makes an illegal move
#
#
# Now, any action with a positive reward is now an optimal action, and any action with a negative reward is suboptimal. Therefore, if I can get a positive reward, a good learner should not bother looking at any other actions, pruning the rest. If I encounter a negative reward, a good learner should never try that action again. The only uncertainty comes into play when the reward is 0 (stopping). In this case, we must try each action until we either find a positive rewarding action or rule them all out (as < 0). An optimal explorer then, will assign a zero probability to negative rewards, 1 probability to positive rewards, and a non-zero probability to 0 rewards. It follows that the initial value of Q should be 0 here. Naturally then, the explorer will do best as my temperature, $T$, for the softmax (action sampling) probabilities approaches 0. Since the temperature is modeled as the average $\alpha$, I greatly reduced the span of $\alpha$, from 200 to 2, promoting quick convergence for $\alpha \to 0.05$ and thus $T \to 0.05$. I then increased the initial value of $\alpha$ to 1.0 in order to learn $Q$ values much quicker (with higher magnitudes), knowing the $\alpha$ values themselves, will still decay to their minimum value of 0.05 quickly.
#
# The final performance can be seen below:
train_test_plots(train_stats, test_stats,
plot_cumulative_success_rate)
train_test_plots(train_stats, test_stats,
plot_cumulative_crimes)
# ### Optimality of the final policy
#
# The agent effectively either took the waypoint or sat if it was illegal. That, to me, is optimal. Something I also looked into was learning my own waypoint by giving relative headings to the destination [up-left, up, up-right, left, right, down-left, down, down-right]. Obviously the environment is rewarding the wrong rewards for this scenario (tuned to the given waypoint), and I did not want to tamper with the environment so I wasn’t able to test this sufficiently.
#
# To get a formal measure of optimality, for each trial, I counted the number of steps, $t$, as well as the number of suboptimal steps (legal but not following waypoint) $t_s$ and crime (illegal) steps $t_c$. Optimality, $\theta$, on each trial is then 1 minus the ratio of non-optimal steps:
#
# $$\theta = 1 - \frac{t_n + t_c}{t}$$
#
# This is shown below for each trial:
# +
def plot_cumulative_optimality(stats, sim_type, ax=None):
(1. - (stats['suboptimals'] + stats['crimes']) /
stats['n_turns']).plot(
ax=ax, kind='area', stacked=False, figsize=(15, 8),
title='%s Optimality in Each Trial' % (
sim_type
)
)
pylab.ylabel('% of Optimal Moves')
pylab.xlabel('Trial#')
train_test_plots(train_stats, test_stats,
plot_cumulative_optimality)
# -
# Overall, the training model seems to learn the optimal path pretty quickly. Once exploration is turned off, the agent is near optimal.
|
README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# ## New Fitting Tools (May/June 2017)
#
# The development branch provides a rewrite of the fitting tools with the goal of defining a single redshift fitting algorithm whether or not you're fitting the `multifit.MultiBeam` or `stack.StackFitter` objects. Fitting the drizzled spectra is generally the fastest, and is *much* faster when many individual beam extractions are available for a given object. For example, sources in the HUDF area can contain as many as 180 individual beams from FIGS, 3D-HST and archival observations, and fitting on the `multifit.MultiBeam` object can take minutes or even hours per source. However, the full WCS information is only preserved for the `multifit.MultiBeam` objects, so these must be used to drizzle the continuum-subtracted, rectified emission line maps.
#
# Also implemented are now more compact data formats for storing all of the outputs of a given source that can be easily distributed without having to provide all of the original FLT files. The `multifit.MultiBeam` object now knows how to read/write a single file (`*beams.fits`) that contains all of the information necessary for performing the fitting analysis.
#
# #### The `fitting.GroupFitter` object can also incorporate broad-band photometry in the redshift fit, though this will be documented at a later time.
# %matplotlib inline
# +
import glob
import time
import os
import numpy as np
import matplotlib.pyplot as plt
import astropy.io.fits as pyfits
import drizzlepac
import grizli
import grizli.stack
# -
# Initialize the GroupFLT object we computed with WFC3IR_Reduction. When loaded from save files
# doesn't much matter what `ref_file` is, since that will be read from the GrismFLT files rather
# than regenerated.
grp = grizli.multifit.GroupFLT(grism_files=glob.glob('*.0?.GrismFLT.fits'), direct_files=[],
ref_file='../Catalog/ERS_goodss_3dhst.v4.0.F160W_orig_sci.fits',
seg_file='../Catalog/ERS_GOODS-S_IR.seg.fits',
catalog='../Catalog/ERS_GOODS-S_IR.cat',
cpu_count=8)
# +
# Extract an object
TARGET = 'ers-grism'
fcontam = 0.2
id = 40776 # emission line object from the other notebook
# +
beams = grp.get_beams(id, size=32)
mb = grizli.multifit.MultiBeam(beams, fcontam=fcontam, group_name=TARGET)
# Make drizzled spectra
hdu, fig = mb.drizzle_grisms_and_PAs(fcontam=fcontam, flambda=False, size=32,
scale=1., kernel='point')
fig.savefig('{0}_{1:05d}.stack.png'.format(TARGET, id))
hdu.writeto('{0}_{1:05d}.stack.fits'.format(TARGET, id), clobber=True)
# Save beam extractions
mb.write_master_fits()
# +
# The continuum model used above is just the flat f-lambda spectrum.
# Fit a polynomial continuum to make it a bit cleaner
wave = np.linspace(3000,2.e4,1000)
tpoly = grizli.utils.polynomial_templates(wave, order=3)
pfit = mb.template_at_z(z=0, templates=tpoly, fitter='lstsq', get_uncertainties=False)
# Re-drizzle with the fit outputs (don't have to do the first drizzle in practice)
hdu, fig = mb.drizzle_grisms_and_PAs(fcontam=fcontam, flambda=False, size=32,
scale=1., kernel='point', zfit=pfit)
fig.savefig('{0}_{1:05d}.stack.png'.format(TARGET, id))
hdu.writeto('{0}_{1:05d}.stack.fits'.format(TARGET, id), clobber=True)
# +
### Initialize templates
# Line complexes for redshift fits.
# Need to have artificially high line FWHM so don't get aliasing in the
# redshift fit.
t0 = grizli.utils.load_templates(fwhm=1200, line_complexes=True, fsps_templates=True)
# Continuum + individual line templates. Can use unresolved line width here
t1 = grizli.utils.load_templates(fwhm=120, line_complexes=False, fsps_templates=True)
# Line drizzle parameters
pline = {'kernel': 'point', 'pixfrac': 0.2, 'pixscale': 0.1, 'size': 8, 'wcs': None}
# -
# ### Fit the individual extractions
# +
# Have generated `mb` above directly from the `beams` extracted from `grp`,
# but here show how it can be generated from the `beams.fits` file saved above.
mb = grizli.multifit.MultiBeam('{0}_{1:05d}.beams.fits'.format(TARGET, id),
group_name=TARGET, fcontam=fcontam)
# Redshift fit
m0 = time.time()
fit = mb.xfit_redshift(templates=t0, zr=[0.1, 2.4], dz=[0.004, 0.0005],
prior=None, fitter='nnls', verbose=False)
m1 = time.time()
# Best-fit template
tfit = mb.template_at_z(z=fit.meta['z_risk'][0], templates=t1,
fit_background=True, fitter='nnls')
m2 = time.time()
fig = mb.xmake_fit_plot(fit, tfit)
print('Redshift fit: {0:.1f} s, Template fit: {1:.1f} s'.format(m1-m0, m2-m1))
# -
# ### Fit the drizzled spectrum
# +
st = grizli.stack.StackFitter(files='{0}_{1:05d}.stack.fits'.format(TARGET, id),
group_name=TARGET, sys_err=0.02, mask_min=0.1,
fit_stacks=False, fcontam=fcontam, pas=None, extensions=None,
min_ivar=0.01, overlap_threshold=3, verbose=True)
# Available extensions. With `fit_stacks=False`, will be each GRISM,PA combination available. Otherwise, will
# be stacks of all PAs for each grism
print('Extensions: ', st.ext)
# +
# Redshift fit, code is identical
m0 = time.time()
fit = st.xfit_redshift(templates=t0, zr=[0.1, 2.4], dz=[0.004, 0.0005],
prior=None, fitter='nnls', verbose=False)
m1 = time.time()
# Best-fit template
tfit = st.template_at_z(z=fit.meta['z_risk'][0], templates=t1,
fit_background=True, fitter='nnls')
m2 = time.time()
fig = st.xmake_fit_plot(fit, tfit)
print('Redshift fit: {0:.1f} s, Template fit: {1:.1f} s'.format(m1-m0, m2-m1))
# -
# ### Wrapper script
#
# Fit the drizzled spectra first (fast) and then the individual spectra near the best redshift. Note that `fitting.run_all` loads the `stack.fits` and `beams.fits` files for a given `id` and `root`name.
# +
### Wrapper script to fit the drizzle spectra first (fast) and then the individual spectra near the best redshift
m0 = time.time()
pline['pixscale'] = 0.1
out = grizli.fitting.run_all(id, t0=t0, t1=t1, fwhm=1200, zr=[0.1, 2.4], dz=[0.004, 0.0005],
fitter='nnls', group_name=TARGET, fit_stacks=False, prior=None,
fcontam=0.2, pline=pline, mask_sn_limit=3, fit_beams=True,
root=TARGET, fit_trace_shift=False, phot=None, verbose=False)
mb, st, fit, tfit, line_hdu = out
m1 = time.time()
print('Run time: {0:.1f} s'.format(m1-m0))
# -
# Output files
ls_str = 'ls -lth {0}_{1:05d}* > lsr; cat lsr'.format(TARGET, id)
os.system(ls_str)
# !cat lsr
# Redrizzle with the full continuum model fit
hdu, fig = mb.drizzle_grisms_and_PAs(fcontam=fcontam, flambda=False, size=32,
scale=1., kernel='point', zfit=tfit)
fig.savefig('{0}_{1:05d}.stack.png'.format(TARGET, id))
hdu.writeto('{0}_{1:05d}.stack.fits'.format(TARGET, id), clobber=True)
# ### Output file
#
# Everything related to the fit is stored in the `full.fits` file.
out = pyfits.open('{0}_{1:05d}.full.fits'.format(TARGET, id)) # The same as line_hdu
out.info()
# PrimaryHDU
h = out[0].header
for k in h:
print('{0:10} = {1} / {2}'.format(k, h[k], h.comments[k]))
# Show the drizzled lines and direct image cutout, which are extensions `DSCI`, `LINE`, etc.
from imp import reload
reload(grizli.fitting)
fig = grizli.fitting.show_drizzled_lines(out, size_arcsec=1.6, cmap='plasma_r')
fig.savefig('{0}_{1:05d}.line.png'.format(TARGET, id))
# +
# Information about the redshift fit is stored in the `ZFIT` extensions.
# The `ZFIT_STACK` extension is the full grid fit with the stacked spectra, and
# `ZFIT_BEAM` is the zoom in with the beam cutout spectra.
#
# Potential fit quality parameters might be (some combination of):
# BIC_TEMP < BIC_POLY >> Bayesian information criteria for the template+z vs. simple polynomial
# fit. If this is False, then the template fit doesn't provide a
# significant improvement (e.g., featureless continuum)
#
# CHIMIN/DOF > xx >> Large reduced chi-squared
#
# MIN_RISK > xx >> "Risk" parameter from Tanaka et al. (2017), from 0 to 1.
# This value is large for relatively flat or multi-modal PDF(z).
zfit_st = grizli.utils.GTable.read(out['ZFIT_STACK'])
zfit_mb = grizli.utils.GTable.read(out['ZFIT_BEAM'])
h = out['ZFIT_STACK'].header
for k in out['ZFIT_STACK'].header:
print('{0:10} = {1} / {2}'.format(k, h[k], h.comments[k]))
# -
print(zfit_st.colnames)
plt.plot(zfit_st['zgrid'], np.log(zfit_st['pdf']))
plt.plot(zfit_mb['zgrid'], np.log(zfit_mb['pdf']))
plt.xlabel('z'); plt.ylabel('PDF(z)')
## Best-fit templates are in `TEMPL`. Full FSPS templates through far-IR
templ = grizli.utils.GTable.read(out['TEMPL'])
templ.info()
# +
import astropy.units as u
fig = plt.figure(figsize=[8,4])
ax1 = fig.add_subplot(121); ax2 = fig.add_subplot(122)
for ax in [ax1, ax2]:
# Continuum only
ax.plot(templ['wave'].to(u.micron), templ['continuum'], color='k', alpha=0.5)
# With lines
ax.plot(templ['wave'].to(u.micron), templ['full'], color='r', alpha=0.5)
ax.set_xlabel(r'$\lambda\ /\ \mu\mathrm{m}$')
ax1.set_ylim(1.e-19, 1.e-17)
ax1.set_xlim(0.7, 1.8) # zoom around spectrum
ax1.semilogy()
ax1.set_ylabel(r'$f_\lambda$; '+templ['full'].unit.__str__())
ax2.set_ylim(1.e-22, 1.e-17)
ax2.set_xlim(0.2, 200)
ax2.loglog()
# -
# Emission line information stored in `COVAR` extension, the data of which is the
# fit covariance at best redshift that is used to compute the parameter uncertainties
h = out['COVAR'].header
for k in h:
print('{0:10} = {1} / {2}'.format(k, h[k], h.comments[k]))
|
examples/Fitting-tools.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Debugging, conditional statements, functions
#
# In this lesson, we will focus on finding and correcting errors in code; making decisions; and using functions. By the end of the session, you should be able to:
# 1. read Python error messages and fix the code.
# 2. use if/elsif/else statements in Python to make decisions.
# 3. create and call your own functions to do calculations.
#
# ## Debugging - finding and correcting errors in your code
#
# It will happen (especially at the beginning) that we will have errors in our code and it will not run. It is important to develop skills to be able to identify where the errors are and fix them. Python will indicate errors in a number of ways. Consider the code below.
# +
# A simple programme with errors
print('Hello there)
name = Input('What is your name? ')
print('Hello there ' + Name ', how are you?\n')
# -
# Copy the code above into a script and run it. In the shell, you should get the following:
# <img src="./img/debug_error.PNG" alt="error messages" align='center' style="width: 800px;"/>
# The ouput in the shell is pointing to a syntax error in line 2. In Thonny, on the right hand side an assistant window also opens up, which provides additional information on the error.
# <img src="./img/assistant_window.PNG" alt="error messages" align='center' style="width: 600px;"/>
# The assistant is saying that the closing quotation mark is missing in the `print` statement.
# When a program does not run read the messages that Thonny is giving you to help fix the code.
#
# ## Exercises
# 1. Correct the remaining errors in the script above.
# 2. Copy the following code into a script and correct it. Read the messages Thonny is telling you to find the errors.
# +
This programme asks user for radius and tells them the volume of the sphere
strain = inpt('What is the strain? ')
E = input("What is the Young's modulus? ")
stress = strain * e
print('If the strain is ' + strain + ' and the Young\'s modulus is ' + E + ' then the stress is ' + stress + '.\n')
# -
# ## Conditional statements: using if/elseif/else statements to make decisions
#
# Conditional statements allow you to execute certain sections of code when you run the program. If a condition is true then a section of code will be executed. If the condition is false then that section of code is skipped.
# <img src="./img/if.png" alt="error messages" align='center' style="width: 400px;"/>
# The simplest conditional statement is the if statement. Consider the following code
# +
# A few lines of code demonstrating the if statement
number = int(input('What is your integer? ')) # User asked for an integer number
if number == 5: # Compare the number to 5
print('You guessed the number!\n') # if they are equal then this line is executed; if not then the code is skipped
print('The guessing game has ended!')
# -
# Note the structure or syntax of the `if` statement. The `if` is followed by a condition (`number == 5`) and then a colon `:`. If the condition is true then the indented lines underneath the `if` statement are exeecuted. In this case there is only one line of code indented after the `if` statement. The indentation is very important - if it is not there the code will not run properly.
#
# == is called a relational operator or comparison operator. It checks whether the two objects either side of the == are equal. More information on comparison operators can be found [here](https://www.tutorialspoint.com/python/python_basic_operators.htm) or on page 20 of your notes.
#
# We can extend the above programme to tell the user if they did not guess the correct number. This is done using an if/else block. In this case if the condition is false then another section of code is executed.
# <img src="./img/if_else.png" alt="error messages" align='center' style="width: 300px;"/>
# +
# A simple programme demonstrating if/else statements
number = int(input('What is the number?'))
if number == 5:
print('You guessed the number!')
else:
print('Sorry. Wrong number.') # This time if the condition number == 5 is false then this line is executed.
print('The guessing game has ended')
# -
# Note the spacing of the code in the above example. The `else` statement is aligned with its respective `if` statement. The code after the `if` and `else` statements are indented 4 spaces. This is a requirement in Python.
#
# You can programme for more choices using the if/elif/else block.
# <img src="./img/if_elseif_else.png" alt="error messages" align='center' style="width: 500px;"/>
# +
# A simple programme demonstrating if/elif/else statements. Checks whether a number is positive, negative or zero
number = float(input('What is the number to two places of decimal? '))
if number < 0:
print(f'The number {number:5.2f} is negative.')
elif number > 0:
print(f'The number {number:5.2f} is positive.')
else:
print('The number is zero.')
print('The guessing game has ended')
# -
# ### Aside - formatting numbers to fixed number of decimal places
# Note how the number is printed to the shell. It is different to how we did it before. Previously we did something like:
number = 5.68791
print('The number ' + str(number) + ' is positive.')
# The method above prints out the number exactly as it is stored. The method in the guessing game script allows us to control the format of the number. This method is known as f-strings. F-strings are very powerful and more information on using them can be found [here](https://realpython.com/python-f-strings/#f-strings-a-new-and-improved-way-to-format-strings-in-python). We are only focussing here on how to output decimal numbers to a specified number of decimal places.
#
# using the specifier `number:5.2f}`. This is a specifier for fixed point notation. This gives more control over the format of numbers we output. The `f` after the `5.2` denotes fixed point notation. The number before the decimal point specifies the minimum width allowed for printing the number. The number after the decimal point specifies the number of decimal places.
#
# #### Quick exercise
# Consider a component with a measured stress of 12.443 MPa under a strain of 0.0148. Try outputting the line *The maximum stress in the component is 12.44 MPa when the strain is 0.015.* to the shell or the cell below.
print(f'The maximum stress in the component is {12.44:5.3f} MPa when the strain is {0.015:5.3f}.')
# The following script demonstrates the use of f-strings to output a number in different formats.
# +
# Programme to demonstrate the use of %f
random_number = 1234.056789; # Just picking a random number
print(f'{random_number:10.1f}\n{random_number:9.3f}\n{random_number:12.6f}\n{random_number:15.5f}\n')
# -
# Note the effect of the number before the decimal point. This will be useful when we create tables of numbers and we want them to be nicely aligned.
# ## Exercises
# Create a Python script to do the following:
# 3. Ask the user for any number between 0 and 100. If they input a number outside this range, tell them it is outside the range. Otherwise tell them if it is less than 50 or greater than 50. Format the output so that the number is to three places of decimal.
#
# ## Creating and using functions
# A function is a block of code that performs an action and can be reused. Functions can accept input and output arguments. This makes them different from the scripts we have written already, which do not accept any arguments.
# An overview of the structure and use of functions in Python can be found [here](https://www.tutorialspoint.com/python/python_functions.htm).
#
# Consider the following basic function, which adds two numbers.
# +
# This is a function that adds the two numbers passed into it
def add_numbers(a, b):
result = a + b
return result
# This script calls the function add_numbers, which is defined below.
total_of_numbers = add_numbers(5,6) # The values 5 and 6 are passed
# into the function and the function returns the sum of them.
print(f'The sum of the numbers 5 and 6 is {total_of_numbers}.')
# -
# Note the structure of a function above. It must start with `def` followed by the name of the function (in this case `add_numbers`), followed by a list of input arguments in brackets (in this case called `a` and `b`). The code within the function must be indented. The last line of the function returns any output arguments if there are any (in this case the variable `result` is returned). It must adhere to this format.
#
# Functions can be defined in the script file before they are called. They can also be located in another file but in this case they must be imported. For now, we will define the functions in the same file as the script in which they are called.
#
# ## Exercises
# Create Python scripts and functions to do the following:
# 4. Create a function that calculates the stress of a material. The input arguments for the function are the strain and Young's modulus; the output argument is the stress. Call the function from a script where the user is asked for the strain and Young's modulus. The script also prints the stress to the output window.
# 5. Create a function that calculates the Reynold's number for flow in a pipe. The input arguments are the velocity, diameter, and kinematic viscosity; the output argument is the Reynold's number. Call the function from a script where the user is asked for the velocity, diameter, and viscosity.
# 6. Modify the script in the previous problem to tell the user whether the flow is laminar (Re < 2000), transitional (2000 < Re < 2500), or turbulent (Re > 2500).
#
#
#
|
topic_02_debugging_if_functions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
#Creates a loop to run through all states in country_codes.csv, running
#mobius.py code to grab the csv file of that state's data, storing in data
import pandas as pd
import os
#move to the mobility report folder as the new working directory
home_dir = '/Users/samismalling/Documents/mobility-report-data-extractor-master'
os.chdir(home_dir)
#import country_codes.csv file:
codes = pd.read_csv('/Users/samismalling/Documents/mobility-report-data-extractor-master/CovidMobile/data/us_state_codes.csv', header = None)
#path names
mobius = './mobius.py'
output_folder = home_dir+'/CovidMobile/data'
dates_file = home_dir+'/config/dates_lookup_2020_04_05.csv'
for state in codes.iloc[23:,2]:
#download pdf and svg files
os.system('python {} download {} 2020-04-05'.format(mobius, state))
#change everything to new state
input_pdf = home_dir+'/pdfs/{}_2020-04-05.pdf'.format(state)
input_svg = home_dir+'/svgs/{}_2020-04-05.svg'.format(state)
#extract csv file and put into data folder in CovidMobile
os.system('python {} full {} {} {} {}'.format(mobius, input_pdf, input_svg, output_folder, dates_file))
|
notebooks/Archive/grab_csvs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="CYNAmoj4zSXl"
# Crie um algoritmo em Python que peça 2 números e imprima o maior deles.
# + id="ViAfpy3GzPN-" colab={"base_uri": "https://localhost:8080/", "height": 68} executionInfo={"status": "ok", "timestamp": 1588878381912, "user_tz": 180, "elapsed": 7521, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEtguPXQdA29TQHTsmcffYdzT6jgW0T8l9pmw2=s64", "userId": "03736159141047480973"}} outputId="092be1e7-d26f-4002-84c9-5606b8c3d705"
a = float(input("Digite um valor"))
b = float(input("Digite um valor"))
if a > b:
print("O valor {0} é maior que {1}".format(a,b))
else:
print("O valor {0} é maior que {1}".format(b,a))
# + [markdown] id="gdMis92izSmQ"
# Crie um algoritmo em Python que peça um valor e mostre na tela se o valor é positivo ou negativo.
# + id="BFpWkdCBz_zh"
a = float(input("Digite um valor"))
if a > 0:
print("O valor {0} é positivo".format(a))
else:
print("O valor {0} é negativo".format(a)
# + [markdown] id="E_0l_Vx6zSo2"
# Crie um algoritmo em Python que verifique se uma letra digitada é vogal ou consoante.
# + id="KwJf7XZS0AZW" colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"status": "ok", "timestamp": 1588974138038, "user_tz": 180, "elapsed": 3384, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEtguPXQdA29TQHTsmcffYdzT6jgW0T8l9pmw2=s64", "userId": "03736159141047480973"}} outputId="6ac51b9f-0e44-462e-ad24-ee764e0bb60f"
a = str(input("Digite uma letra "))
if a == "a" or a == "e" or a == "i" or a == "o"or a == "u":
print("Sua letra é uma vogal")
else:
print("Sua letra é uma consoante")
# + [markdown] id="1i4VCiiczSrL"
# Faça um programa para a leitura de duas notas parciais de um aluno. O programa deve calcular a média alcançada por aluno e apresentar:
# * "Aprovado", se a média alcançada for maior ou igual a sete;
# * "Reprovado", se a média for menor do que sete;
# * "Aprovado com Distinção", se a média for igual a 10.
# + id="yoW67hjA0BHy"
# + [markdown] id="4ep7kezvzStO"
# Crie um algoritmo em Python que leia três números e mostre o maior deles.
# + id="6K6IHWdr0Blt" colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"status": "ok", "timestamp": 1588974394016, "user_tz": 180, "elapsed": 3953, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEtguPXQdA29TQHTsmcffYdzT6jgW0T8l9pmw2=s64", "userId": "03736159141047480973"}} outputId="bf77ffe0-6919-47ad-9f4e-94c4d5e48fc9"
a = int(input("Digite um número "))
b = int(input("Digite um número "))
c = int(input("Digite um número "))
if a > b and a > c:
print("Seu numero é o maior {0}".format(a))
elif b > c:
print("Seu numero é o maior {0}".format(b))
else:
print("Seu numero é o maior {0}".format(c))
# + [markdown] id="lIlfG-ATzgOr"
# Faça um programa que pergunte o preço de 5 produtos e informe qual produto você deve comprar, sabendo que a decisão é sempre pelo mais barato.
# + id="-y4pB6br0DD_" colab={"base_uri": "https://localhost:8080/", "height": 119} executionInfo={"status": "ok", "timestamp": 1589104218107, "user_tz": 180, "elapsed": 8726, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEtguPXQdA29TQHTsmcffYdzT6jgW0T8l9pmw2=s64", "userId": "03736159141047480973"}} outputId="d573d128-a9eb-424f-b03a-0571e4aec1aa"
lista = []
for i in range(5):
lista.insert(i, int(input("Digite o valor do produto")))
item = lista[0]
for j in range(5):
if (lista[j] <= item):
item = lista[j]
print("O produto mais barato é {0}".format(item))
# + [markdown] id="Vj4OXsJQziu_"
# Crie um algoritmo em Python que leia três números e mostre-os em ordem decrescente.
# + id="RmApGpg30D00" colab={"base_uri": "https://localhost:8080/", "height": 85} executionInfo={"status": "ok", "timestamp": 1589105026771, "user_tz": 180, "elapsed": 11240, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEtguPXQdA29TQHTsmcffYdzT6jgW0T8l9pmw2=s64", "userId": "03736159141047480973"}} outputId="6909a630-a1ea-48df-eeb6-e979ebbf3a1b"
lista = []
for i in range (3):
lista.insert(i,int(input("Digite um valor ")))
lista.reverse()
print("Sua lista em ordem decrescente é {0}".format(str(lista)))
# + [markdown] id="e7id8Ewgzlow"
# As Organizações Mendéz resolveram dar um aumento de salário aos seus colaboradores e lhe contraram para desenvolver o programa que calculará os reajustes em Python. Faça um programa que receba o salário de um colaborador e o reajuste segundo o seguinte critério, baseado no salário atual:
#
# * salários entre 680,00 e 800,00 : aumento de 15%<br>
# * salários entre 800,00 e 2500,00 : aumento de 10%<br>
# * salários de 2500,00 em diante : aumento de 5% após o aumento ser realizado.
#
# Informe na tela:
# * o salário antes do reajuste;
# * o percentual de aumento aplicado;
# * o valor do aumento;
# * o novo salário, após o aumento.
# + id="Kl0ainsv0Eif" colab={"base_uri": "https://localhost:8080/", "height": 51} executionInfo={"status": "ok", "timestamp": 1589104850563, "user_tz": 180, "elapsed": 3826, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14GiEtguPXQdA29TQHTsmcffYdzT6jgW0T8l9pmw2=s64", "userId": "03736159141047480973"}} outputId="123096dc-7950-4211-f698-cf19604bf910"
sal_antigo = float(input("Digite seu salário "))
if 680.00 <= sal_antigo >= 800.00:
aum = sal_antigo * 0.15
sal_novo = sal_antigo + aum
print("Seu salario agora será {0}, você teve um aumento de 15%, o valor do aumento é {1} e seu salario antigo era {2}".format(sal_novo,aum,sal_antigo))
elif 800.00 < sal_antigo >= 2500.00:
aum = sal_antigo * 0.10
sal_novo = sal_antigo + aum
print("Seu salario agora será {0}, você teve um aumento de 10%, o valor do aumento é {1} e seu salario antigo era {2}".format(sal_novo,aum,sal_antigo))
elif 2500.00 < sal_antigo:
aum = sal_antigo * 0.05
sal_novo = sal_antigo + aum
print("Seu salario agora será {0}, você teve um aumento de 5%, o valor do aumento é {1} e seu salario antigo era {2}".format(sal_novo,aum,sal_antigo))
else:
print("algo errado")
|
Exercícios básicos - 03 - Feitos.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:anaconda2-aparent_proj]
# language: python
# name: conda-env-anaconda2-aparent_proj-py
# ---
# +
#trying to train regression model to classify a site
import numpy as np
import sklearn as sk
from sklearn import metrics
form sklearn import linear_model
import matplotlib.pyplot as plt
import math
import pandas as pd
#
|
logRegressionV1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.5.2
# language: julia
# name: julia-1.5
# ---
diff_forward(f, x; h=sqrt(eps(Float64))) = (f(x+h) - f(x))/h
diff_central(f, x; h=sqrt(eps(Float64))) = (f(x+h/2) - f(x-h/2))/h
diff_backward(f, x; h=sqrt(eps(Float64))) = (f(x) - f(x-h))/h
f1(x) = x^3 - 2x
diff_backward(f1, 4.6)
diff_central(f1, 4.6)
diff_forward(f1, 4.6)
|
Code/Chapter 2 - Derivatives and Gradients/2.4.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Multi-Spectral Registration
#
# Copyright (c) 2020, <NAME>, <NAME>, University of Nevada, Reno.
#
# All rights reserved.
#
# This is an IPython Notebook demonstrating usage of the multispectral registration package.
# + tags=[]
# %matplotlib inline
import time
from multi_spect_tools import multi_spect_common
from multi_spect_tools import multi_spect_image_io
from multi_spect_tools import multi_spect_plotting_utils
from multi_spect_tools import sitk_multi_spect_registration
# -
# ## Create Registration Object
# First, we will create a registration object, passing it a path to a configuration file.
# + tags=[]
# Create a registration object
sitk_reg_obj = sitk_multi_spect_registration.sitk_registration("cfg/reg_config.ini")
# -
# ## Load an image from the dataset
# Next, we will load an image from the dataset listed in the configuration file. If the configuration file does not list a dataset, this snippet will fail. Image paths are indexed by their ID, interpretted by their file name. For more information, please see the README section on dataset processing. Note that the paths for dataset processing are not required for image alignment; images may be loaded independently, and simply fed to the alignment function demonstrated in the next code block.
# + tags=[]
# Choose an image to load
img_id = 28
# Load the paths to this image
image_paths = sitk_reg_obj.config.get_img_paths(img_id)
# Load the image
input_image = multi_spect_image_io.load_image_from_path_list(image_paths)
# Display the image
multi_spect_plotting_utils.show_multi_ch_image(input_image, sitk_reg_obj.config.ordered_channel_names)
# -
# ## Run Alignment
# Next, we will run the alignment of the loaded image by calling the _align_ function. We pass the input image, and receive the aligned output image, as well as a results object. The results object includes a number of metrics regarding the alignment, as well as the transformation objects applied to retrieve the aligned image. This transformation will be used in subsequent cells as the initial transform for the next image's alignment.
# + tags=[]
output_image, results = sitk_reg_obj.align(input_image,init_transforms=None, print_output=True)
# -
# ## Show the Aligned Image
# Here we use the plotting utility's function _show_merged_ to display the aligned image.
# The function will plot the image as well as rectangles showing the relative position of each channel.
#
# The function takes:
#
# 1. The aligned image
# 2. The indicies of the desired RGB channels
# 3. A channel to mix with the RGB
# 4. The corner points of each of the aligned channels
# 5. The color of each box to draw
#
# In the line below, the red channel is given to exist at the 2nd index of the image, the green at the 1st, and the blue at the 0th. These three channels will be blended (averaged) with the 3rd channel, which in this case is NIR.
# + tags=[]
frame_colors = {'Red':'r', 'Blue':'b', 'Green':'g', 'NIR':'m'}
multi_spect_plotting_utils.show_merged(output_image, [2,1,0], blend_ch=3, image_bounds=results.corner_points, colors=frame_colors)
# -
# ## Show Convergence Metrics for Each Channel
# Below, we plot the convergence values per iteration for each channel.
multi_spect_plotting_utils.plot_metric_vals(results.metric_logs, ['b', 'g', 'r'], sitk_reg_obj.config.moving_channel_names, 3)
# ## Load a 2nd Image
# To demonstrate the process of providing an initial transform, we will load and display a 2nd image.
# Choose an image to load
img_id = 24
# Load the paths to this image
image_paths = sitk_reg_obj.config.get_img_paths(img_id)
# Load the image
input_image = multi_spect_image_io.load_image_from_path_list(image_paths)
# ## Align the image, using the previous result's transformation
# Here we use the *results* object to acquire the previous alignment's final transform and feed it to the alignment of the next image.
# + tags=[]
output_image, results = sitk_reg_obj.align(input_image, init_transforms=results.alignment_transform, print_output=True)
# -
# ## Show the Aligned Image
multi_spect_plotting_utils.show_merged(output_image, [2,1,0], 3, results.corner_points, frame_colors)
|
Sitk_Demo_Registration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# In the following example, **title**, **x label** and **y label** are added to the [barplot](http://python-graph-gallery.com/barplot/) using the `title()`, `xlabel()`, and `ylabel()` functions of the [matplotlib](http://python-graph-gallery.com/matplotlib/) library.
#
# Those functions are applied to a barplot in the example, but the same method would work for other chart types.
# +
# libraries
import numpy as np
import matplotlib.pyplot as plt
# create dataset
height = [3, 12, 5, 18, 45]
bars = ('A', 'B', 'C', 'D', 'E')
x_pos = np.arange(len(bars))
# Create bars and choose color
plt.bar(x_pos, height, color = (0.5,0.1,0.5,0.6))
# Add title and axis names
plt.title('My title')
plt.xlabel('categories')
plt.ylabel('values')
# Create names on the x axis
plt.xticks(x_pos, bars)
# Show graph
plt.show()
# -
# >Note: the [matplotlib section](https://www.python-graph-gallery.com/matplotlib/) provides a lot of tips and tricks on how to customize a matplotlib chart!
|
src/notebooks/4-add-title-and-axis-label.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.5 64-bit (''env'': venv)'
# language: python
# name: python3
# ---
# +
# Load Libraries
import os
import pandas as pd
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
# Set up paths
PATH = os.getcwd()[:-13]
DATA_PATH = PATH + "Dataset\superstore.csv"
df = pd.read_csv(DATA_PATH)
# -
# **Check data loaded properly**
df.head(5)
# **Drop sales since we don't want to "peek" at profit**
df = df.drop("Sales",axis=1)
# **Check for nulls and dtypes**
df.info()
# Look for strong correlations among numerical data
sns.heatmap(df.corr())
# # Brainstorm
#
# Types of data we have:
# - Geographic
# - Time
# - Descriptive
# - Numerical
#
# GEOGRAPHIC:
# - These products are being shipped
# - Distance from shipping site to location
# - Try to determine shipping location
#
# TIME:
# - Is there a time of year that makes more?
# - How many years of data do we have?
# - How does the profit change over time?
# - Are are there locations that buy more during a time of year?
# - Grab days of the week/ months/ years
#
# DESCRIPTIVE:
# - Do certain areas like certain catagories more?
# - What product sells the most/least?
# - Who is buying?
# - Who buys more Corporate/Consumer? (Quantity and profit)
# - What kind of shipping is preferred? By whom?
#
# NUMERICAL:
# - Who gets most/least discounts? Why?
# - When do discounts grow?
# - Where gets the most/least discounts? Why?
#
# THOUGHTS:
# There are many more questions to be asked and areas that could be explored, but
# this should suffice for a glipmse into what is happening with our website.
#
# ACTION:
# Get the days of the week/ months/ year and then go top to bottom exploring each
# thought. (Subject to revision)
#
#
# Extract months/years/dayofweek and convert to pandas datetime object
def fix_time(data):
# Convert to pandas datetime object
data['Order Date'] = pd.to_datetime(data["Order Date"])
data['Ship Date'] = pd.to_datetime(data["Ship Date"])
# Get the day of the week
data['Order_Day'] = data['Order Date'].dt.day_name()
data['Ship_Day'] = data['Ship Date'].dt.day_name()
# Get the month
data['Order_Month'] = data['Order Date'].dt.month_name()
data['Ship_Month'] = data['Ship Date'].dt.month_name()
# Get the year
data['Order_Year'] = data['Order Date'].dt.year.astype('object')
data['Ship_Year'] = data['Ship Date'].dt.year.astype('object')
return data
eda = fix_time(df)
eda
# **Use Tableau to visualize geographic data**
# TIME:
# - Is there a time of year that makes more?
# - How many years of data do we have?
# - How does the profit change over time?
# - Are are there locations that buy more during a time of year?
# - Grab days of the week/ months/ years
# 2015.5 was a good year, what happened?
# We have Ship but not Order
sns.lineplot(data=eda, x='Ship_Year', y='Profit')
sns.lineplot(data=eda, x='Order_Year', y='Profit')
eda.Ship_Year.sort_values().value_counts()
eda.Order_Year.sort_values().value_counts()
sns.barplot(data=eda, x='Order_Day', y='Profit')
sns.barplot(data=eda, x='Ship_Day', y='Profit')
sns.scatterplot(data=eda, x='State', y='Profit')
eda.info()
drop_lst = (['Row ID',
'Order Date', 'Ship Date',
'Order ID', 'Product Name',
'Customer Name','Country',
'Ship_Day','Ship_Year',
'Ship_Month', 'Customer ID'])
eda.drop(drop_lst, axis=1, inplace=True)
eda.to_csv('sales EDA.csv')
|
Data Analysis/sales_EDA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import os
import os.path as path
from sklearn.model_selection import train_test_split
index_list = [
[0,2],
[3,58],
[59,63],
[64,182],
[183,185],
[186,190]
]
y_list = ['bed', 'bird', 'cat', 'dog', 'house', 'tree']
base_path = 'data'
# +
# Load npz
savepath = path.join(base_path, '172_bbc.npz')
l = np.load(savepath)
bbc = l['bbc']
bbclabel = l['bbclabel']
savepath = path.join(base_path, '172_dht.npz')
l = np.load(savepath)
dht = l['dht']
dhtlabel = l['dhtlabel']
# -
savepath = path.join(base_path, '172_val.npz')
l = np.load(savepath)
valimg = l['valimg']
vallabel = l['vallabel']
img_shape = (172, 172, 3)
# +
x_train = np.zeros((len(bbc)+len(dht), *img_shape))
for i in range(len(bbc)):
x_train[i] = bbc[i].reshape(img_shape)
for i in range(len(dht)):
x_train[len(bbc)+i] = dht[i].reshape(img_shape)
# +
a = np.argmax(bbclabel, axis=1)
b = np.argmax(dhtlabel, axis=1)
y_train = np.hstack((a,b))
for i, [begin, end] in enumerate(index_list):
y_train[(y_train >= begin) & (y_train <= end)] = i
# +
valimg = l['valimg']
vallabel = l['vallabel']
x_val = np.zeros((len(valimg), *img_shape))
for i in range(len(valimg)):
x_val[i] = valimg[i].reshape(img_shape)
y_val = np.argmax(vallabel, axis=1)
for i, [begin, end] in enumerate(index_list):
y_val[(y_val >= begin) & (y_val <= end)] = i
# + active=""
# y_label = np.argmax(y_data, axis=1)
# y_text = ['bed', 'bird', 'cat', 'dog', 'house', 'tree']
# y_table = {i:text for i, text in enumerate(y_text)}
# y_table_array = np.array([(i, text) for i, text in enumerate(y_text)])
# + active=""
# x_train_temp, x_test, y_train_temp, y_test = train_test_split(
# x_2d_data, y_label, test_size=0.2, random_state=42, stratify=y_label)
#
# x_train, x_val, y_train, y_val = train_test_split(
# x_train_temp, y_train_temp, test_size=0.25, random_state=42, stratify=y_train_temp)
#
# x_train.shape, y_train.shape, x_val.shape, y_val.shape, x_test.shape, y_test.shape
# -
np.savez_compressed(path.join(base_path, 'imagenet_6_class_172_train_data.npz'),
x_data=x_train, y_data=y_train, y_list=y_list)
np.savez_compressed(path.join(base_path, 'imagenet_6_class_172_val_data.npz'),
x_data=x_val, y_data=y_val, y_list=y_list)
|
make_172_imagenet_6_class_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/sergejhorvat/TensorFlow-in-Practice---Coursera/blob/master/Natural%20Language%20Processing%20in%20TensorFlow/Week%202%20-%20Lesson%201.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="P-AhVYeBWgQ3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 680} outputId="65331d82-ee26-4d8e-9407-68502670358f"
# NOTE: PLEASE MAKE SURE YOU ARE RUNNING THIS IN A PYTHON3 ENVIRONMENT
# !pip install tensorflow-gpu==2.0.0
import tensorflow as tf
print(tf.__version__)
# This is needed for the iterator over the data
# But not necessary if you have TF 2.0 installed
# #!pip install tensorflow==2.0.0-beta0
# Only needed if TF != 2.0.0
#tf.enable_eager_execution()
# !pip install -q tensorflow-datasets
# + id="_IoM4VFxWpMR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 276} outputId="91f24c3b-22ee-4275-8332-b865b6536583"
import tensorflow_datasets as tfds
imdb, info = tfds.load("imdb_reviews", with_info=True, as_supervised=True)
# + id="wHQ2Ko0zl7M4" colab_type="code" colab={}
import numpy as np
train_data, test_data = imdb['train'], imdb['test']
training_sentences = []
training_labels = []
testing_sentences = []
testing_labels = []
# str(s.tonumpy()) is needed in Python3 instead of just s.numpy()
for s,l in train_data:
training_sentences.append(str(s.numpy()))
training_labels.append(l.numpy())
for s,l in test_data:
testing_sentences.append(str(s.numpy()))
testing_labels.append(l.numpy())
training_labels_final = np.array(training_labels)
testing_labels_final = np.array(testing_labels)
# + id="P8TlrYW-4GW8" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="540ec3dd-aef7-4fec-a2f9-7d5c43eab629"
# Explore datasets
training_sentences[1]
# + id="7n15yyMdmoH1" colab_type="code" colab={}
vocab_size = 10000
embedding_dim = 16
max_length = 120
trunc_type='post'
oov_tok = "<OOV>"
from tensorflow.keras.preprocessing.text import Tokenizer
from tensorflow.keras.preprocessing.sequence import pad_sequences
tokenizer = Tokenizer(num_words = vocab_size, oov_token=oov_tok)
tokenizer.fit_on_texts(training_sentences)
word_index = tokenizer.word_index
sequences = tokenizer.texts_to_sequences(training_sentences)
padded = pad_sequences(sequences,maxlen=max_length, truncating=trunc_type)
testing_sequences = tokenizer.texts_to_sequences(testing_sentences)
testing_padded = pad_sequences(testing_sequences,maxlen=max_length)
# + colab_type="code" id="9axf0uIXVMhO" colab={"base_uri": "https://localhost:8080/", "height": 74} outputId="54e5f166-5321-48a2-eec1-610c201bbec7"
reverse_word_index = dict([(value, key) for (key, value) in word_index.items()])
def decode_review(text):
return ' '.join([reverse_word_index.get(i, '?') for i in text])
print(decode_review(padded[1]))
print(training_sentences[1])
# + id="5NEpdhb8AxID" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 311} outputId="053877f8-e6dd-4260-8d78-6d92abf244d8"
model = tf.keras.Sequential([
tf.keras.layers.Embedding(vocab_size, embedding_dim, input_length=max_length),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(6, activation='relu'),
tf.keras.layers.Dense(1, activation='sigmoid')
])
model.compile(loss='binary_crossentropy',optimizer='adam',metrics=['accuracy'])
model.summary()
# + id="V5LLrXC-uNX6" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 441} outputId="47e7befc-b4e3-4b32-85be-29fb8724471c"
num_epochs = 10
model.fit(padded,
training_labels_final,
epochs=num_epochs,
validation_data=(testing_padded, testing_labels_final)
)
# + id="yAmjJqEyCOF_" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 35} outputId="3ac5af07-602a-4820-b276-0bbaf1f55824"
e = model.layers[0]
weights = e.get_weights()[0]
print(weights.shape) # shape: (vocab_size, embedding_dim)
# + id="jmB0Uxk0ycP6" colab_type="code" colab={}
import io
out_v = io.open('vecs.tsv', 'w', encoding='utf-8')
out_m = io.open('meta.tsv', 'w', encoding='utf-8')
for word_num in range(1, vocab_size):
word = reverse_word_index[word_num]
embeddings = weights[word_num]
out_m.write(word + "\n")
out_v.write('\t'.join([str(x) for x in embeddings]) + "\n")
out_v.close()
out_m.close()
# + id="VDeqpOCVydtq" colab_type="code" colab={}
try:
from google.colab import files
except ImportError:
pass
else:
files.download('vecs.tsv')
files.download('meta.tsv')
# + id="YRxoxc2apscY" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 55} outputId="03a2d00d-626b-47d1-b3b5-75da90686744"
sentence = "I really think this is amazing. honest."
sequence = tokenizer.texts_to_sequences(sentence)
print(sequence)
# + id="BESx5mC_CI-U" colab_type="code" colab={}
|
Natural Language Processing in TensorFlow/Week 2 - Lesson 1.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import sys
sys.path.append('./slidescore/')
from slidescore_utils import APIClient
import os
import json
import numpy as np
import PIL
from sklearn import metrics
df_h = pd.read_csv('/project/schirris/basisscripts/step_4/basis_slidescore_answers_summary.csv')
all_labels = pd.read_csv('/project/schirris/basisscripts/step_3/data_basis_brca_with_labels.csv').groupby('case').mean()
df_ai = pd.read_csv('/home/yonis/SimCLR-1/logs/eval/1342/regression_output_epoch_20_2020-10-09-16-24-40.csv').set_index('patient')
im_path = '/home/yonis/SimCLR-1/logs/eval/1342/roc_curve_epoch_20.png'
# +
# display(Image.open(im_path))
# -
df_h['patient']=df_h.apply(lambda x: x['path'].split('/')[-1].split('_')[1].split('-')[0], axis=1)
# df_h['WSI']=df.apply(lambda x: x['path'].split('/')[-1][7:-5], axis=1)
# df_h['label'] = df.apply(lambda x: all_labels.loc[x['WSI'], 'HRD'], axis=1)
# df_h['label'] = df_h.apply(lambda x: all_labels.loc[x['patient'], 'median_HRD'], axis=1)
df_h_test = df_h[df_h['patient'].isin(df_ai.index)]
df_h_test = df_h_test[df_h_test['hrd'].isin(['No', 'Yes'])]
len(df_h_test)
df_h_test['h_label'] = df_h_test.apply(lambda x: 1 if x['hrd'] == "Yes" else 0, axis=1)
df_h_test['real_label'] = df_h_test.apply(lambda x: all_labels.at[x['patient'], 'median_HRD'], axis=1)
df_h_test['correct'] = df_h_test.apply(lambda x: x['h_label']==x['real_label'], axis=1)
df_h_test['ai_label'] = df_h_test.apply(lambda x: df_ai.at[x['patient'], 'preds'], axis=1)
df_h_test['correct']
|
notebooks/.ipynb_checkpoints/13_hugo_vs_ai-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ___
#
# ___
# # K Nearest Neighbors with Python
#
# You've been given a classified data set from a company! They've hidden the feature column names but have given you the data and the target classes.
#
# We'll try to use KNN to create a model that directly predicts a class for a new data point based off of the features.
#
# Let's grab it and use it!
# ## Import Libraries
#
#
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import numpy as np
# %matplotlib inline
# ## Get the Data
#
# Set index_col=0 to use the first column as the index.
df = pd.read_csv("Classified Data",index_col=0)
df.head()
# ## Standardize the Variables
#
# Because the KNN classifier predicts the class of a given test observation by identifying the observations that are nearest to it, the scale of the variables matters. Any variables that are on a large scale will have a much larger effect on the distance between the observations, and hence on the KNN classifier, than variables that are on a small scale.
from sklearn.preprocessing import StandardScaler
scaler = StandardScaler()
scaler.fit(df.drop('TARGET CLASS',axis=1))
scaled_features = scaler.transform(df.drop('TARGET CLASS',axis=1))
df_feat = pd.DataFrame(scaled_features,columns=df.columns[:-1])
df_feat.head()
# ## Train Test Split
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(scaled_features,df['TARGET CLASS'],
test_size=0.30)
# ## Using KNN
#
# Remember that we are trying to come up with a model to predict whether someone will TARGET CLASS or not. We'll start with k=1.
from sklearn.neighbors import KNeighborsClassifier
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
# ## Predictions and Evaluations
#
# Let's evaluate our KNN model!
from sklearn.metrics import classification_report,confusion_matrix
print(confusion_matrix(y_test,pred))
print(classification_report(y_test,pred))
# ## Choosing a K Value
#
# Let's go ahead and use the elbow method to pick a good K Value:
# +
error_rate = []
# Will take some time
for i in range(1,40):
knn = KNeighborsClassifier(n_neighbors=i)
knn.fit(X_train,y_train)
pred_i = knn.predict(X_test)
error_rate.append(np.mean(pred_i != y_test))
# -
plt.figure(figsize=(10,6))
plt.plot(range(1,40),error_rate,color='blue', linestyle='dashed', marker='o',
markerfacecolor='red', markersize=10)
plt.title('Error Rate vs. K Value')
plt.xlabel('K')
plt.ylabel('Error Rate')
# Here we can see that that after arouns K>23 the error rate just tends to hover around 0.06-0.05 Let's retrain the model with that and check the classification report!
# +
# FIRST A QUICK COMPARISON TO OUR ORIGINAL K=1
knn = KNeighborsClassifier(n_neighbors=1)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
print('WITH K=1')
print('\n')
print(confusion_matrix(y_test,pred))
print('\n')
print(classification_report(y_test,pred))
# +
# NOW WITH K=23
knn = KNeighborsClassifier(n_neighbors=23)
knn.fit(X_train,y_train)
pred = knn.predict(X_test)
print('WITH K=23')
print('\n')
print(confusion_matrix(y_test,pred))
print('\n')
print(classification_report(y_test,pred))
# -
# # Great job!
#
# We were able to squeeze some more performance out of our model by tuning to a better K value!
|
Topics_Master/14-K-Nearest-Neighbors/01-K Nearest Neighbors with Python.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
import numpy as np
import pandas as pd
import tensorflow as tf
import matplotlib.pyplot as plt
from tensorflow.contrib import learn
from sklearn.metrics import mean_squared_error
from lstm import generate_data, lstm_model, load_csvdata
import dateutil.parser
import datetime
import matplotlib.dates as mdates
LOG_DIR = './ops_logs/lstm_weather'
TIMESTEPS = 10
RNN_LAYERS = [{'num_units': 5}]
DENSE_LAYERS = [10, 10]
TRAINING_STEPS = 100000
BATCH_SIZE = 100
PRINT_STEPS = TRAINING_STEPS / 100
# # Transform QCLCD Data Function
# downloaded weather data from http://www.ncdc.noaa.gov/qclcd/QCLCD
def load_weather_frame(filename):
#load the weather data and make a date
data_raw = pd.read_csv(filename, dtype={'Time': str, 'Date': str})
data_raw['WetBulbCelsius'] = data_raw['WetBulbCelsius'].astype(float)
times = []
for index, row in data_raw.iterrows():
_t = datetime.time(int(row['Time'][:2]), int(row['Time'][:-2]), 0) #2153
_d = datetime.datetime.strptime( row['Date'], "%Y%m%d" ) #20150905
times.append(datetime.datetime.combine(_d, _t))
data_raw['_time'] = pd.Series(times, index=data_raw.index)
df = pd.DataFrame(data_raw, columns=['_time','WetBulbCelsius'])
return df.set_index('_time')
# # Load The Data as CSV
# This is QCLCD data for PDX. It is waht will be used to train the model.
# scale values to reasonable values and convert to float
data_weather = load_weather_frame("data/QCLCD_PDX_20150901.csv")
X, y = load_csvdata(data_weather, TIMESTEPS, seperate=False)
# # Run The Model and Fit Predictions
regressor = learn.Estimator(model_fn=lstm_model(TIMESTEPS, RNN_LAYERS, DENSE_LAYERS),
model_dir=LOG_DIR)
# +
# create a lstm instance and validation monitor
validation_monitor = learn.monitors.ValidationMonitor(X['val'], y['val'],
every_n_steps=PRINT_STEPS,
early_stopping_rounds=1000)
regressor.fit(X['train'], y['train'],
monitors=[validation_monitor],
batch_size=BATCH_SIZE,
steps=TRAINING_STEPS)
predicted = regressor.predict(X['test'])
# +
#not used in this example but used for seeing deviations
rmse = np.sqrt(((predicted - y['test']) ** 2).mean(axis=0))
score = mean_squared_error(predicted, y['test'])
print ("MSE: %f" % score)
# +
# plot the data
all_dates = data_weather.index.get_values()
fig, ax = plt.subplots(1)
fig.autofmt_xdate()
predicted_values = predicted.flatten() #already subset
predicted_dates = all_dates[len(all_dates)-len(predicted_values):len(all_dates)]
predicted_series = pd.Series(predicted_values, index=predicted_dates)
plot_predicted, = ax.plot(predicted_series, label='predicted (c)')
test_values = y['test'].flatten()
test_dates = all_dates[len(all_dates)-len(test_values):len(all_dates)]
test_series = pd.Series(test_values, index=test_dates)
plot_test, = ax.plot(test_series, label='2015 (c)')
xfmt = mdates.DateFormatter('%b %d %H')
ax.xaxis.set_major_formatter(xfmt)
# ax.fmt_xdata = mdates.DateFormatter('%Y-%m-%d %H')
plt.title('PDX Weather Predictions for 2016 vs 2015')
plt.legend(handles=[plot_predicted, plot_test])
plt.show()
|
lstm_weather.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import re
import pandas as pd
from nltk.tokenize import WordPunctTokenizer
from bs4 import BeautifulSoup
from src.tweet_cleaner import TweetCleaner
cols = ['sentiment','id','date','query_string','user','text']
df = pd.read_csv("data/source/train.csv", header=None, names=cols, encoding="ISO-8859-1")
df.head()
df.drop(['id','date','query_string','user'],axis=1,inplace=True)
df.head()
df['sentiment'] = df['sentiment'].map({0: 0, 4: 1})
df.sentiment.value_counts()
clean_tweets = []
tweetCleaner = TweetCleaner()
count=-1
for tweet in df['text']:
count+=1
if( (count)%100000 == 0):
print(count, 'tweets processed')
clean_tweets.append(tweetCleaner.clean(tweet))
print('done')
clean_df = pd.DataFrame(clean_tweets, columns=['text'])
clean_df['sentiment'] = df.sentiment
clean_df.head()
clean_df.text = clean_df.text[clean_df.text != '']
clean_df.dropna(inplace=True)
clean_df.reset_index(drop=True,inplace=True)
clean_df.info()
clean_df.to_csv('data/clean_tweets_test.csv',encoding='utf-8')
|
data/data_preprocessing.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # TensorRTに変換
# Pytorchのresnet18モデルをTensorRTモデルに変換します。
#
# PytorchモデルをTensorRTモデルに変換するには、[torch2trt](https://github.com/NVIDIA-AI-IOT/torch2trt) を利用します。
# モデルのベンチマーク速度は以下のように向上します。
#
# |Model|Nano (PyTorch)|Nano (TensorRT)|Xavier (PyTorch)|Xavier (TensorRT)|
# |-------------|-------------|-------------|-------------|-------------|
# |alexnet|46.4|69.9|250|580|
# |resnet18|29.4|90.2|140|712|
#
# アプリケーション全体のFPSはモデル単体のベンチマーク結果よりも遅くなります。
# +
import torch
import torchvision
device = torch.device('cuda')
model = torchvision.models.resnet18(pretrained=False)
model.fc = torch.nn.Linear(512, 2)
model = model.cuda().eval().half()
# -
# 学習済みモデルを読込みます。
model.load_state_dict(torch.load('road_following_model.pth'))
# TensorRTに変換します。
# +
from torch2trt import torch2trt
data = torch.zeros((1, 3, 224, 224)).cuda().half()
model_trt = torch2trt(model, [data], fp16_mode=True)
# -
# TernsorRTモデルを保存します。
torch.save(model_trt.state_dict(), 'road_following_model_trt.pth')
# ## 次の作業
#
# 04.road_following.ipynb を実行し、TensorRTに変換した学習済みモデルで、実際の走行にトライします。
|
notebooks/03.convert_to_trt.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="4vqepaR4bZzJ"
# # Tutorial on ASR Finetuning with CTC model
# Let's finetune a pretrained ASR model!
#
# Here we provide pre-trained speech recognition model with CTC loss that is trained on many open-sourced datasets. Details can be found in [Rethinking Evaluation in ASR: Are Our Models Robust Enough?](https://arxiv.org/abs/2010.11745)
# + [markdown] id="AbTYA-j_vk-7"
# ## Step 1: Install `Flashlight`
# First we install `Flashlight` and its dependencies. Flashlight is built from source with either CPU/CUDA backend and installation takes **~16 minutes**.
#
# For installation out of colab notebook please use [link](https://github.com/fairinternal/flashlight#building).
#
#
#
# + id="oDJo0P8DRL3N"
# First, choose backend to build with
backend = 'CUDA' #@param ["CPU", "CUDA"]
# Clone Flashlight
# !git clone https://github.com/flashlight/flashlight.git
# install all dependencies for colab notebook
# !source flashlight/scripts/colab/colab_install_deps.sh
# + [markdown] id="3yXKe3wUWiWV"
# Build CPU/CUDA Backend of `Flashlight`:
# - Build from current master.
# - Builds the ASR app.
# - Resulting binaries in `/content/flashlight/build/bin/asr`.
#
# If using a GPU Colab runtime, build the CUDA backend; else build the CPU backend.
# + id="x5QFOur3AcB1"
# export necessary env variables
# %env MKLROOT=/opt/intel/mkl
# %env ArrayFire_DIR=/opt/arrayfire/share/ArrayFire/cmake
# %env DNNL_DIR=/opt/dnnl/dnnl_lnx_2.0.0_cpu_iomp/lib/cmake/dnnl
if backend == "CUDA":
# Total time: ~13 minutes
# !cd flashlight && mkdir -p build && cd build && \
# cmake .. -DCMAKE_BUILD_TYPE=Release \
# -DFL_BUILD_TESTS=OFF \
# -DFL_BUILD_EXAMPLES=OFF \
# -DFL_BUILD_APP_ASR=ON && \
# make -j$(nproc)
elif backend == "CPU":
# Total time: ~14 minutes
# !cd flashlight && mkdir -p build && cd build && \
# cmake .. -DFL_BACKEND=CPU \
# -DCMAKE_BUILD_TYPE=Release \
# -DFL_BUILD_TESTS=OFF \
# -DFL_BUILD_EXAMPLES=OFF \
# -DFL_BUILD_APP_ASR=ON && \
# make -j$(nproc)
else:
raise ValueError(f"Unknown backend {backend}")
# + [markdown] id="e-V0U-Dow-vs"
# Let's take a look around.
# + colab={"base_uri": "https://localhost:8080/"} id="k6tRnX1iHCoX" outputId="e7a0fb5f-3507-4c7e-801c-9308ea65637e"
# Binaries are located in
# !ls flashlight/build/bin/asr
# + [markdown] id="__56IY0xcgMv"
# ## Step 2: Setup Finetuning
# + [markdown] id="T9078zTzcdS3"
# #### Downloading the model files
#
# First, let's download the pretrained models for finetuning.
#
# For acoustic model, you can choose from
#
# >Architecture | # Params | Criterion | Model Name | Arch Name
# >---|---|:---|:---:|:---:
# > Transformer|70Mil|CTC|am_transformer_ctc_stride3_letters_70Mparams.bin |am_transformer_ctc_stride3_letters_70Mparams.arch
# > Transformer|300Mil|CTC|am_transformer_ctc_stride3_letters_300Mparams.bin | am_transformer_ctc_stride3_letters_300Mparams.arch
# > Conformer|25Mil|CTC|am_conformer_ctc_stride3_letters_25Mparams.bin|am_conformer_ctc_stride3_letters_25Mparams.arch
# > Conformer|87Mil|CTC|am_conformer_ctc_stride3_letters_87Mparams.bin|am_conformer_ctc_stride3_letters_87Mparams.arch
# > Conformer|300Mil|CTC|am_conformer_ctc_stride3_letters_300Mparams.bin| am_conformer_ctc_stride3_letters_300Mparams.arch
#
# For demonstration, we will use the model in first row and download the model and its arch file.
# + id="Xn8XH01Chilw"
# !wget -nv --continue -o /dev/null https://dl.fbaipublicfiles.com/wav2letter/rasr/tutorial/am_transformer_ctc_stride3_letters_70Mparams.bin -O model.bin # acoustic model
# !wget -nv --continue -o /dev/null https://dl.fbaipublicfiles.com/wav2letter/rasr/tutorial/am_transformer_ctc_stride3_letters_70Mparams.arch -O arch.txt # model architecture file
# + [markdown] id="AlArtmiEhitM"
# Along with the acoustic model, we will also download the tokens file, lexicon file
# + id="RPDEovLVc46Z"
# !wget -nv --continue -o /dev/null https://dl.fbaipublicfiles.com/wav2letter/rasr/tutorial/tokens.txt -O tokens.txt # tokens (defines predicted tokens)
# !wget -nv --continue -o /dev/null https://dl.fbaipublicfiles.com/wav2letter/rasr/tutorial/lexicon.txt -O lexicon.txt # lexicon files (defines mapping between words)
# + [markdown] id="UKZpiqt9bM4a"
# #### Downloading the dataset
#
# For finetuning the model, we provide a limited supervision dataset based on [AMI Corpus](http://groups.inf.ed.ac.uk/ami/corpus/). It consists of 10m, 1hr and 10hr subsets organized as follows.
#
# ```
# dev.lst # development set
# test.lst # test set
# train_10min_0.lst # first 10 min fold
# train_10min_1.lst
# train_10min_2.lst
# train_10min_3.lst
# train_10min_4.lst
# train_10min_5.lst
# train_9hr.lst # remaining data of the 10h split (10h=1h+9h)
# ```
# The 10h split is created by combining the data from the 9h split and the 1h split. The 1h split is itself made of 6 folds of 10 min splits.
#
# The recipe used for preparing this corpus can be found [here](https://github.com/flashlight/wav2letter/tree/master/data/ami).
#
# **You can also use your own dataset to finetune the model instead of AMI Corpus.**
# + colab={"base_uri": "https://localhost:8080/"} id="naYUr0cOjF1Q" outputId="cb9ff351-a9ad-4232-d4ab-d72f34796098"
# !rm ami_limited_supervision.tar.gz
# !wget -nv --continue -o /dev/null https://dl.fbaipublicfiles.com/wav2letter/rasr/tutorial/ami_limited_supervision.tar.gz -O ami_limited_supervision.tar.gz
# !tar -xf ami_limited_supervision.tar.gz
# !ls ami_limited_supervision
# + [markdown] id="Mj32IH3HvjZo"
# ### Get baseline WER before finetuning
#
# Before proceeding to finetuning, let's test (viterbi) WER on AMI dataset to we have something to compare results after finetuning.
# + id="ppGOZZWR4t_y" colab={"base_uri": "https://localhost:8080/"} outputId="ecfb6def-2e4f-4482-ddc4-a710e6e5cc5f"
# ! ./flashlight/build/bin/asr/fl_asr_test --am model.bin --datadir '' --emission_dir '' --uselexicon false \
# --test ami_limited_supervision/test.lst --tokens tokens.txt --lexicon lexicon.txt --show
# + [markdown] id="0jcXZGQmm-Rs"
# We can see that the viterbi WER is 26.6% before finetuning.
# + [markdown] id="Ryc17nPTds7Q"
# ## Step 3: Run Finetuning
#
#
# Now, let's run finetuning with the AMI Corpus to see if we can improve the WER.
#
# Important parameters for `fl_asr_finetune_ctc`:
#
# `--train`, `--valid` - list files for training and validation sets respectively. Use comma to separate multiple files
#
# `--datadir` - [optional] base path to be used for `--train`, `--valid` flags
#
# `--lr` - learning rate for SGD
#
# `--momentum` - SGD momentum
#
# `--lr_decay` - epoch at which learning decay starts
#
# `--lr_decay_step` - learning rate halves after this epoch interval starting from epoch given by `lr_decay`
#
# `--arch` - architecture file. Tune droupout if necessary.
#
# `--tokens` - tokens file
#
# `--batchsize` - batchsize per process
#
# `--lexicon` - lexicon file
#
# `--rundir` - path to store checkpoint logs
#
# `--reportiters` - Number of updates after which we will run evaluation on validation data and save model, if 0 we only do this at end of each epoch
#
#
# >Amount of train data | Config to use
# >---|---|
# > 10 min| --train train_10min_0.lst
# > 1 hr| --train train_10min_0.lst,train_10min_1.lst,train_10min_2.lst,train_10min_3.lst,train_10min_4.lst,train_10min_5.lst
# > 10 hr| --train train_10min_0.lst,train_10min_1.lst,train_10min_2.lst,train_10min_3.lst,train_10min_4.lst,train_10min_5.lst,train_9hr.lst
#
# Let's run finetuning with 10hr AMI data (**~7min** for 1000 updates with evaluation on dev set)
#
# + colab={"base_uri": "https://localhost:8080/"} id="1RscueA4dlz0" outputId="4906a93a-28c4-4078-c750-1bbc39f4e3e6"
# ! ./flashlight/build/bin/asr/fl_asr_tutorial_finetune_ctc model.bin \
# --datadir ami_limited_supervision \
# --train train_10min_0.lst,train_10min_1.lst,train_10min_2.lst,train_10min_3.lst,train_10min_4.lst,train_10min_5.lst,train_9hr.lst \
# --valid dev:dev.lst \
# --arch arch.txt \
# --tokens tokens.txt \
# --lexicon lexicon.txt \
# --rundir checkpoint \
# --lr 0.025 \
# --netoptim sgd \
# --momentum 0.8 \
# --reportiters 1000 \
# --lr_decay 100 \
# --lr_decay_step 50 \
# --iter 25000 \
# --batchsize 4 \
# --warmup 0
# + [markdown] id="4BCed5Whw5YQ"
# ## Step 4: Run Decoding
# + [markdown] id="e3pHr97mLp8W"
# #### Viterbi decoding
#
# + id="OrrthfLYAfB_" colab={"base_uri": "https://localhost:8080/"} outputId="d5cea038-9d0a-40ff-dcd2-fe998eec39d7"
# ! ./flashlight/build/bin/asr/fl_asr_test --am checkpoint/001_model_dev.bin --datadir '' --emission_dir '' --uselexicon false \
# --test ami_limited_supervision/test.lst --tokens tokens.txt --lexicon lexicon.txt --show
# + [markdown] id="6oz870UrAkJu"
# Viterbi WER improved from 26.6% to 19.5% after 1 epoch with finetuning...
# + [markdown] id="d3iJXpsQLwW0"
# #### Beam Search decoding with a language model
#
# To do this, download the finetuned model and use the [Inference CTC tutorial](https://colab.research.google.com/github/flashlight/flashlight/blob/master/flashlight/app/asr/tutorial/notebooks/InferenceAndAlignmentCTC.ipynb)
# + [markdown] id="EPmZlgbRNmh1"
# ## Step 5: Running with your own data
#
# To finetune on your own data, create `train`, `dev` and `test` list files and run the finetuning step.
#
# Each list file consists of multiple lines with each line describing one sample in the following format :
# ```
# <sample_id> <path_to_audio_file> <duration> <transcript>
# ```
#
# For example, let's take a look at the `dev.lst` file from AMI corpus.
# + colab={"base_uri": "https://localhost:8080/"} id="MKW6ktM7NxWV" outputId="19020539-f010-4670-cd81-60fa92f79593"
# ! head ami_limited_supervision/dev.lst
# + [markdown] id="_Ag_BKtdPedy"
# #### Recording your own audio
#
# For example, you can record your own audio and finetune the model...
#
# Installing a few packages first...
# + id="Z9XKBvzpPqm4"
# !apt-get install sox
# !pip install ffmpeg-python sox
# + id="LpPIM81ZLGnn"
from flashlight.scripts.colab.record import record_audio
# + [markdown] id="WaxI4DAaK6gM"
# **Let's record now the following sentences:**
#
# **1:** A flashlight or torch is a small, portable spotlight.
#
# + id="mEFlB90VOtaF" colab={"base_uri": "https://localhost:8080/", "height": 136} outputId="a9b1f261-117f-4ffb-9212-1ab022d2e8e2"
record_audio("recorded_audio_1")
# + [markdown] id="x0DEuxuwLLOc"
# **2:** Its function is a beam of light which helps to see and it usually requires batteries.
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="N84pjDEsKs6S" outputId="f546e80e-671f-43c6-8822-0db9f7271946"
record_audio("recorded_audio_2")
# + [markdown] id="4fIrr7DrLb_L"
# **3:** In 1896, the first dry cell battery was invented.
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="qGn8e_cALcZO" outputId="4accd936-7a4d-4fcb-a3ee-dd30ee5823f1"
record_audio("recorded_audio_3")
# + [markdown] id="7PLh3W-8LPTo"
# **4:** Unlike previous batteries, it used a paste electrolyte instead of a liquid.
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="TF813JDeKt6i" outputId="586b8f61-21a7-418d-bf21-802eeea3f0b1"
record_audio("recorded_audio_4")
# + [markdown] id="sl3xY-w_LWpF"
# **5** This was the first battery suitable for portable electrical devices, as it did not spill or break easily and worked in any orientation.
# + colab={"base_uri": "https://localhost:8080/", "height": 136} id="Zpp6K6JJLZDW" outputId="4e953fe0-6f72-4589-89e6-2e71ac47a833"
record_audio("recorded_audio_5")
# + [markdown] id="vyfYgfHnLqxf"
# ### Create now new training/dev lists:
#
# (yes, you need to edit transcriptions below to your recordings)
# + id="vLK6F1cyLtGp"
import sox
transcriptions = [
"a flashlight or torch is a small portable spotlight",
"its function is a beam of light which helps to see and it usually requires batteries",
"in eighteen ninthy six the first dry cell battery was invented",
"unlike previous batteries it used a paste electrolyte instead of a liquid",
"this was the first battery suitable for portable electrical devices, as it did not spill or break easily and worked in any orientation"
]
with open("own_train.lst", "w") as f_train, open("own_dev.lst", "w") as f_dev:
for index, transcription in enumerate(transcriptions):
fname = "recorded_audio_" + str(index + 1) + ".wav"
duration_ms = sox.file_info.duration(fname) * 1000
if index % 2 == 0:
f_train.write("{}\t{}\t{}\t{}\n".format(
index + 1, fname, duration_ms, transcription))
else:
f_dev.write("{}\t{}\t{}\t{}\n".format(
index + 1, fname, duration_ms, transcription))
# + [markdown] id="y8ou2LnDQONk"
# ### Check at first model quality on dev before finetuning
# + id="Fq4g9tCAQFw8"
# ! ./flashlight/build/bin/asr/fl_asr_test --am model.bin --datadir '' --emission_dir '' --uselexicon false \
# --test own_dev.lst --tokens tokens.txt --lexicon lexicon.txt --show
# + [markdown] id="VUy16oZRQjT1"
# ### Finetune on recorded audio samples
#
# Play with parameters if needed.
# + id="YAtnwKDXMtZ2"
# ! ./flashlight/build/bin/asr/fl_asr_tutorial_finetune_ctc model.bin \
# --datadir= \
# --train own_train.lst \
# --valid dev:own_dev.lst \
# --arch arch.txt \
# --tokens tokens.txt \
# --lexicon lexicon.txt \
# --rundir own_checkpoint \
# --lr 0.025 \
# --netoptim sgd \
# --momentum 0.8 \
# --reportiters 1000 \
# --lr_decay 100 \
# --lr_decay_step 50 \
# --iter 25000 \
# --batchsize 4 \
# --warmup 0
# + [markdown] id="3dWOwSVgQ3bL"
# ### Test finetuned model
#
# (unlikely you get significant improvement with just five phrases, but let's check!)
# + id="j4uG8XYVTqQu"
# ! ./flashlight/build/bin/asr/fl_asr_test --am own_checkpoint/001_model_dev.bin --datadir '' --emission_dir '' --uselexicon false \
# --test own_dev.lst --tokens tokens.txt --lexicon lexicon.txt --show
|
flashlight/app/asr/tutorial/notebooks/FinetuneCTC.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import tensorflow as tf
print(tf.math.add(1, 2))
print(tf.math.add([1, 2,3], [3, 4,5]))
print(tf.math.square(5))
print(tf.math.reduce_sum([1, 2, 3]))
|
Chapter 01/tensors.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="WesiGGnUJehP" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 17} outputId="801cf8fd-b56b-409e-a09d-bdd0a432f5ca"
# http://pytorch.org/
from os import path
from wheel.pep425tags import get_abbr_impl, get_impl_ver, get_abi_tag
platform = '{}{}-{}'.format(get_abbr_impl(), get_impl_ver(), get_abi_tag())
accelerator = 'cu80' if path.exists('/opt/bin/nvidia-smi') else 'cpu'
# !pip install -q http://download.pytorch.org/whl/{accelerator}/torch-0.3.0.post4-{platform}-linux_x86_64.whl torchvision
import torch
# + id="hjCa95mEBkxd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 153} outputId="cba25ba5-10df-44f2-fb08-9fb0cb361dc8"
import os
import torch
import torch.nn as nn
from torch.autograd import Variable
import torchvision.datasets as dset
import torchvision.transforms as transforms
import torch.nn.functional as F
import torch.optim as optim
## load mnist dataset
use_cuda = torch.cuda.is_available()
root = './data'
if not os.path.exists(root):
os.mkdir(root)
trans = transforms.Compose([transforms.ToTensor(), transforms.Normalize((0.5,), (1.0,))])
# if not exist, download mnist dataset
train_set = dset.MNIST(root=root, train=True, transform=trans, download=True)
test_set = dset.MNIST(root=root, train=False, transform=trans, download=True)
batch_size = 100
train_loader = torch.utils.data.DataLoader(
dataset=train_set,
batch_size=batch_size,
shuffle=True)
test_loader = torch.utils.data.DataLoader(
dataset=test_set,
batch_size=batch_size,
shuffle=False)
print('==>>> total trainning batch number: {}'.format(len(train_loader)))
print('==>>> total testing batch number: {}'.format(len(test_loader)))
## network
class MLPNet(nn.Module):
def __init__(self):
super(MLPNet, self).__init__()
self.fc1 = nn.Linear(28*28, 500)
self.fc2 = nn.Linear(500, 256)
self.fc3 = nn.Linear(256, 10)
def forward(self, x):
x = x.view(-1, 28*28)
x = F.relu(self.fc1(x))
x = F.relu(self.fc2(x))
x = self.fc3(x)
return x
def name(self):
return "MLP"
class LeNet(nn.Module):
def __init__(self):
super(LeNet, self).__init__()
self.conv1 = nn.Conv2d(1, 20, 5, 1)
self.conv2 = nn.Conv2d(20, 50, 5, 1)
self.fc1 = nn.Linear(4*4*50, 500)
self.fc2 = nn.Linear(500, 10)
def forward(self, x):
x = F.relu(self.conv1(x))
x = F.max_pool2d(x, 2, 2)
x = F.relu(self.conv2(x))
x = F.max_pool2d(x, 2, 2)
x = x.view(-1, 4*4*50)
x = F.relu(self.fc1(x))
x = self.fc2(x)
return x
def name(self):
return "LeNet"
# + id="05fV-EfKBkxi" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1207} outputId="75af2347-502e-483f-aba3-e97b02fa6ba5"
## training
model = LeNet()
if use_cuda:
model = model.cuda()
optimizer = optim.SGD(model.parameters(), lr=0.01, momentum=0.9)
criterion = nn.CrossEntropyLoss()
for epoch in range(10):
# trainning
ave_loss = 0
for batch_idx, (x, target) in enumerate(train_loader):
optimizer.zero_grad()
if use_cuda:
x, target = x.cuda(), target.cuda()
x, target = Variable(x), Variable(target)
out = model(x)
loss = criterion(out, target)
ave_loss = ave_loss * 0.9 + loss.data[0] * 0.1
loss.backward()
optimizer.step()
if (batch_idx+1) % 100 == 0 or (batch_idx+1) == len(train_loader):
print('==>>> epoch: {}, batch index: {}, train loss: {:.6f}'.format(
epoch, batch_idx+1, ave_loss))
# testing
correct_cnt, ave_loss = 0, 0
total_cnt = 0
for batch_idx, (x, target) in enumerate(test_loader):
if use_cuda:
x, target = x.cuda(), target.cuda()
x, target = Variable(x, volatile=True), Variable(target, volatile=True)
out = model(x)
loss = criterion(out, target)
_, pred_label = torch.max(out.data, 1)
total_cnt += x.data.size()[0]
correct_cnt += (pred_label == target.data).sum()
# smooth average
ave_loss = ave_loss * 0.9 + loss.data[0] * 0.1
if(batch_idx+1) % 100 == 0 or (batch_idx+1) == len(test_loader):
print('==>>> epoch: {}, batch index: {}, test loss: {:.6f}, acc: {:.3f}'.format(
epoch, batch_idx+1, ave_loss, correct_cnt * 1.0 / total_cnt))
torch.save(model.state_dict(), model.name())
# + id="ierGZaPfBkxm" colab_type="code" colab={}
the_model = LeNet()
the_model.load_state_dict(torch.load("LeNet")) #读取网络参数
|
minst.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import tensorflow as tf
import tensorflow_probability as tfp
import numpy as np
import matplotlib.pyplot as plt
tfd = tfp.distributions
tfb = tfp.bijectors
from tensorflow.keras.models import Model
from tensorflow.keras.layers import Dense
# -
# # Discretized Logistic Mixture Distribution
# ## Single Logistic Distribution
vals = np.linspace(-10, 10, 1000)
len(vals)
dist1 = tfd.Logistic(loc=1, scale=1)
dist1
dist1.mean()
dist1.stddev()
dist2 = tfd.Logistic(loc=1, scale=0.25)
dist3 = tfd.Logistic(loc=1, scale=2)
# ### Plot distributions by different sufficient statistics
# +
plt.figure(figsize=(14,5))
plt.subplot(1,2,1)
plt.plot(vals, dist1.prob(vals))
plt.plot(vals, dist2.prob(vals), ls='-.')
plt.plot(vals, dist3.prob(vals), ls='--')
plt.legend(['scale=1', 'scale=0.25', 'scale=2'], loc='upper left')
plt.xlabel('x', fontsize=14)
plt.ylabel('probability density function', fontsize=14)
plt.grid()
plt.subplot(1,2,2)
plt.plot(vals, dist1.cdf(vals))
plt.plot(vals, dist2.cdf(vals),ls='-.')
plt.plot(vals, dist3.cdf(vals),ls='--')
plt.legend(['scale=1', 'scale=0.25','scale=2'], loc='upper left')
plt.xlabel('x',fontsize=14)
plt.ylabel('cumulative probability density function',fontsize=14)
plt.grid()
plt.show()
# -
# ## Discretization
# TF document: https://www.tensorflow.org/probability/api_docs/python/tfp/distributions/QuantizedDistribution
bits = 4
n_vals = np.linspace(0, 15, 16)
n_vals
logi = tfd.Logistic(loc=1, scale=0.25)
discretized_logi = tfd.QuantizedDistribution(logi, low=0., high=2**bits-1.)
discretized_logi.prob(n_vals)
plt.stem(n_vals, discretized_logi.prob(n_vals))
vals = np.linspace(-10, 10, 1000)
# +
plt.figure(figsize=(14,5))
plt.subplot(1,2,1)
plt.plot(vals, logi.prob(vals))
plt.stem(n_vals, discretized_logi.prob(n_vals), use_line_collection=True, markerfmt='C1o',linefmt='C1-.')
plt.legend(['Inner distribution (pdf)','Quantized version (pmf)'], loc='upper left')
plt.legend(['Inner distribution','Quantized version'], loc='upper right')
plt.xlabel('x',fontsize=14)
plt.ylabel('probability density function',fontsize=14)
plt.subplot(1,2,2)
plt.plot(vals, logi.cdf(vals))
plt.plot(vals, discretized_logi.cdf(vals))
plt.legend(['Inner distribution','Quantized version'], loc='lower right')
plt.xlabel('x',fontsize=14)
plt.ylabel('cumulative probability density function',fontsize=14)
plt.show()
# -
discretized_logi.prob(1)
t_vals = np.linspace(0,5,10)
plt.plot(logi.prob(t_vals).numpy())
plt.plot(discretized_logi.prob(t_vals).numpy())
print(logi.sample(10))
print(discretized_logi.sample(10))
# Discretized version should have summation of probabilities as 1.
np.sum(discretized_logi.prob(n_vals))
# Because of ceiling, stem does not match with its original distribution, so it needs to be shifted by 0.5 to left.
# +
bits = 4
logi = tfd.Logistic(loc=1, scale=0.25)
logi_shift = tfd.TransformedDistribution(distribution=logi, bijector=tfb.Shift(shift=-0.5))
discretized_logi = tfd.QuantizedDistribution(logi_shift, low=0., high=2**bits-1.)
# -
vals = np.linspace(-10,10,1000)
# +
plt.figure(figsize=(14,5))
plt.subplot(1,2,1)
plt.plot(vals, logi.prob(vals))
plt.plot(vals, logi_shift.prob(vals))
plt.stem(n_vals, discretized_logi.prob(n_vals), use_line_collection=True, markerfmt='C1o',linefmt='C1-.')
plt.legend(['Inner distribution','Shifted','Quantized version'], loc='upper right')
plt.xlabel('x',fontsize=14)
plt.ylabel('probability density function',fontsize=14)
plt.subplot(1,2,2)
plt.plot(vals, logi.cdf(vals))
plt.plot(vals, logi_shift.cdf(vals))
plt.plot(vals, discretized_logi.cdf(vals))
plt.legend(['Inner distribution','Shifted','Quantized version'], loc='lower right')
plt.xlabel('x',fontsize=14)
plt.ylabel('cumulative probability density function',fontsize=14)
plt.show()
# -
# ### Mixture of logistic distributions and quantized version of the mixtures
locs = (4.0, 10.0)
scales = (0.25, 0.5)
probs = (0.8, 0.2)
dists = tfd.Logistic(loc=locs, scale=scales)
dists_tf = tfd.TransformedDistribution(dists, bijector=tfb.Shift(shift=-0.5))
quant = tfd.QuantizedDistribution(dists_tf, 0, high=2**4-1.)
# +
mixture = tfd.MixtureSameFamily(
mixture_distribution=tfd.Categorical(probs=probs),
components_distribution=dists)
quant_mixture = tfd.MixtureSameFamily(
mixture_distribution=tfd.Categorical(probs=probs),
components_distribution=quant)
# -
vals = np.linspace(0,15.1,1000)
# +
plt.figure(figsize=(14,5))
plt.subplot(1,2,1)
plt.plot(vals, mixture.prob(vals))
plt.stem(n_vals, quant_mixture.prob(n_vals), use_line_collection=True, markerfmt='C1o',linefmt='C1-.')
plt.legend(['Inner distribution (pdf)','Quantized version (pmf)'], loc='upper right')
plt.xlabel('x',fontsize=14)
plt.ylabel('probability',fontsize=14)
plt.subplot(1,2,2)
plt.plot(vals, mixture.cdf(vals))
plt.plot(vals, quant_mixture.cdf(vals),ls='-.')
plt.legend(['Inner distribution','Quantized version'], loc='lower right')
plt.xlabel('x',fontsize=14)
plt.ylabel('cumulative probability density function',fontsize=14)
plt.show()
# -
np.sum(quant_mixture.prob(n_vals))
np.mean(dists.sample(100), axis=0)
np.mean(quant.sample(100), axis=0)
mixture.sample(10).numpy()
quant_mixture.sample(10).numpy()
# ### Using the mixture of logistics in a neural network
def quant_mixture_logistic(out, bits=8, num=3):
loc, un_scale, logits = tf.split(out,
num_or_size_splits=num,
axis=-1)
print('out: ', out)
print('LOC: ', loc)
print('un_scale: ', un_scale)
print('logits: ', logits)
scale = tf.nn.softplus(un_scale)
discretized_logistic_dist = tfd.QuantizedDistribution(
distribution=tfd.TransformedDistribution(
distribution=tfd.Logistic(loc=loc, scale=scale),
bijector=tfb.AffineScalar(shift=-0.5)),
low=0.,
high=2**bits - 1.)
mixture_dist = tfd.MixtureSameFamily(
mixture_distribution=tfd.Categorical(logits=logits),
components_distribution=discretized_logistic_dist)
return mixture_dist
inputs = tf.keras.layers.Input(shape=(100,))
h1 = Dense(10, activation='tanh')(inputs)
out = Dense(6)(h1)
p_y = tfp.layers.DistributionLambda(quant_mixture_logistic)(out)
model = Model(inputs=inputs, outputs=p_y)
model.summary()
|
Notebooks/04_Probabilistic_DL_in_the_Wild.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# * Given a stock data, which has open and close prices for each trading day, we want to create an array of (n_samples, n_prices).
# > n_prices = 10
#
# > n_samples = np.floor(stock_data's length/n_prices)
#
#
# +
import pandas as pd
import numpy as np
dict_tickers = {
'Apple': 'AAPL',
'Microsoft': 'MSFT',
'Google': 'GOOG',
'Bitcoin': 'BTC-USD',
'Facebook': 'FB'
}
# -
def stockDataTransformer(filepath):
df = pd.read_csv(filepath)
df.set_index('Date', inplace=True)
df1 = df[['Open', 'Close']].copy()
data = df1.values
n_samples = data.shape[0]//10*10
reshape_number = data.shape[0]*data.shape[1]//10
data1 = data[:n_samples].reshape((reshape_number, 10))
return data1
tickerName = 'Apple'
filepath = f"raw-stock-data/{tickerName}.csv"
cols = [[f"open{i}", f"close{i}"] for i in range(1, 6)]
cols = [item for sublist in cols for item in sublist]
cols
pd.DataFrame(stockDataTransformer(filepath), columns = cols)
|
simple-data-generator/stockDataTransformer_nb.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# Text Processing and Machine Learning
# ====================================
#
# - pre-processing and tokenization (splitting text into words)
# - n-grams, vectorization and word embeddings
# - train and evaluate a text classifier
# - a short look into [Hugging Face's transformers library](https://huggingface.co/transformers/)
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Natural Language Processing
#
# [Natural language processing (NLP)](https://en.wikipedia.org/wiki/Natural_language_processing) is about programming computers to process and analyze natural language data (text and speech).
#
# During the text classification training we touch only some aspects of NLP, namely
#
# - tokenization or splitting a text into words (aka. tokens)
# - the representation of words in a vector space (word embeddings)
#
# NLP modules for Python:
#
# - [spaCy](https://spacy.io/) or [spaCy on pypi](https://pypi.org/project/spacy/)
# - [NLTK](https://www.nltk.org/) or [NLTK on pypi](https://pypi.org/project/nltk/)
#
# + [markdown] slideshow={"slide_type": "slide"}
# ## Machine Learning
#
# The field of machine learning is too broad to be introduced here. Please, see [Google's machine learning crash course](https://developers.google.com/machine-learning/crash-course/ml-intro).
# + [markdown] slideshow={"slide_type": "slide"}
# ## fastText
#
# [fastText](https://fasttext.cc/) is a software library for text
# classification and word representation learning. See the fastText
# tutorials for
#
# - [text classification](https://fasttext.cc/docs/en/supervised-tutorial.html)
# - [word representation learning](https://fasttext.cc/docs/en/unsupervised-tutorial.html)
#
# We will now follow the [fastText text
# classification](https://fasttext.cc/docs/en/supervised-tutorial.html)
# tutorial (cf. documentation of the [Python module
# "fasttext"](https://pypi.org/project/fasttext/)) to train and apply
# a text classifier.
#
# + [markdown] slideshow={"slide_type": "subslide"}
# The fastText tutorial uses the StackExchange cooking data set. We will use the [Kaggle Toxic Comment Classification Challenge](https://www.kaggle.com/c/jigsaw-toxic-comment-classification-challenge/overview) data set. In order to download the data set, you need to register at [Kaggle.com](https://www.kaggle.com/).
#
# After the data set is downloaded and unpacked into the folder `data/kaggle-jigsaw-toxic`, you should see the tree files `train.csv`, `test.csv` and `test_labels.csv` in the mentioned folder.
# + slideshow={"slide_type": "subslide"}
import pandas as pd
df_train = pd.read_csv('data/kaggle-jigsaw-toxic/train.csv')
#df.head()
# + slideshow={"slide_type": "subslide"}
labels = ['toxic', 'severe_toxic', 'obscene', 'threat', 'insult', 'identity_hate']
df_train[labels].mean()
# + [markdown] slideshow={"slide_type": "subslide"}
# Only 10% of the comments are toxic. What does it mean for building a classifier?
# + slideshow={"slide_type": "subslide"}
# tokenize the comments
import string
from nltk.tokenize import TweetTokenizer
tweet_tokenizer = TweetTokenizer(reduce_len=True)
def tokenize(text):
global tweet_tokenizer
words = tweet_tokenizer.tokenize(text)
words = filter(lambda w: w != ''
and w not in string.punctuation, words)
words = map(lambda w: w.lower(), words)
return ' '.join(words)
tokenize("You're a hero! http://example.com/index.html")
# + slideshow={"slide_type": "subslide"}
# write data to fastText train file
train_file = 'data/kaggle-jigsaw-toxic/train.txt'
def write_line_fasttext(fp, row):
global labels
line = ''
for label in labels:
if row[label] == 1:
if line:
line += ' '
line += '__label__' + label
if line:
line += ' '
else:
line += '__label__none '
line += tokenize(row['comment_text'])
fp.write(line)
fp.write('\n')
with open(train_file, 'w') as fp:
df_train.apply(lambda row: write_line_fasttext(fp, row), axis=1)
# -
# !pip install fasttext
# + slideshow={"slide_type": "subslide"}
# train a model
import fasttext
model = fasttext.train_supervised(input=train_file, wordNgrams=2, minCount=2)
# + slideshow={"slide_type": "subslide"}
model.predict(tokenize("This is a well-written article."))
# model.predict(tokenize("Fuck you!"), k=5)
# + slideshow={"slide_type": "subslide"}
# looking into the underlying word embeddings
model.get_nearest_neighbors('idiot', k=20)
# + slideshow={"slide_type": "subslide"}
# save the model
model_file = 'data/kaggle-jigsaw-toxic/model.bin'
model.save_model(model_file)
# + slideshow={"slide_type": "subslide"}
df_test = pd.read_csv('data/kaggle-jigsaw-toxic/test.csv')
df_test_labels = pd.read_csv('data/kaggle-jigsaw-toxic/test_labels.csv')
# join both tables
df_test = df_test.merge(df_test_labels, on='id')
# skip rows not labelled / not used
df_test = df_test[df_test['toxic'] != -1]
test_file = 'data/kaggle-jigsaw-toxic/test.txt'
# write test set for fastText
with open(test_file, 'w') as fp:
df_test.apply(lambda row: write_line_fasttext(fp, row), axis=1)
# + [markdown] slideshow={"slide_type": "subslide"}
# ### Model Validation
#
# See also: [precision and recall](https://en.wikipedia.org/wiki/Precision_and_recall)
# + slideshow={"slide_type": "subslide"}
model.test(test_file)
# + slideshow={"slide_type": "subslide"}
res_per_label = model.test_label(test_file)
for label in res_per_label.items():
print(label)
# + slideshow={"slide_type": "subslide"}
# in case the fastText command-line tool is installed: it has a nice output formatter
# !fasttext test-label \
# data/kaggle-jigsaw-toxic/model.bin \
# data/kaggle-jigsaw-toxic/test.txt
# + [markdown] slideshow={"slide_type": "slide"}
# ## Transformers
#
# - https://en.wikipedia.org/wiki/Transformer_(machine_learning_model)
# - [Hugging Face's transformers library](https://huggingface.co/transformers/): unique interface and provisioning of various transformer language models
# - see https://huggingface.co/course
# + slideshow={"slide_type": "subslide"}
# !pip install transformers
# !pip install tensorflow
# !pip install "transformers[sentencepiece]"
# + slideshow={"slide_type": "subslide"}
from transformers import pipeline
p = pipeline('fill-mask', model='bert-base-german-cased')
# + slideshow={"slide_type": "subslide"}
for s in p("Er arbeitet als [MASK]."): print(s)
# + slideshow={"slide_type": "subslide"}
pipeline_fill_mask = pipeline('fill-mask', model='bert-base-german-cased')
def fill_mask(cloze):
global pipeline_fill_mask
for s in pipeline_fill_mask(cloze):
print('%-20s\t%.5f' % (s['token_str'], s['score']))
# + slideshow={"slide_type": "subslide"}
fill_mask("Er arbeitet als [MASK] in einer Klinik.")
# + slideshow={"slide_type": "subslide"}
fill_mask("Er arbeitet als [MASK] in einer Lungenklinik.")
# + slideshow={"slide_type": "subslide"}
fill_mask("Er arbeitet als [MASK] bei BMW.")
# + slideshow={"slide_type": "subslide"}
fill_mask("Er arbeitet als [MASK] an der Universität Konstanz.")
# + slideshow={"slide_type": "subslide"}
fill_mask("Sie arbeitet als [MASK] an der Universität Konstanz.")
# + slideshow={"slide_type": "subslide"}
fill_mask("Sie ist wirklich [MASK].")
# + slideshow={"slide_type": "subslide"}
fill_mask("Er ist wirklich [MASK].")
# + slideshow={"slide_type": "subslide"}
help(pipeline)
# + slideshow={"slide_type": "subslide"}
p = pipeline('sentiment-analysis')
p("I'm happy.")
# + slideshow={"slide_type": "subslide"}
p("I'm sad.")
# + slideshow={"slide_type": "subslide"}
p("I'm not happy.")
# + slideshow={"slide_type": "subslide"}
import transformers
p = pipeline('ner', aggregation_strategy=transformers.pipelines.AggregationStrategy.SIMPLE)
p("""We would like to belatedly welcome Ulrich Glassmann of the Europa-Universität
Flensburg (#EUF), who is currently a guest at the Cluster. Ulrich has just decided
to extend his stay until the end of June, welcome news indeed!""")
# + slideshow={"slide_type": "subslide"}
p = pipeline('translation', model='facebook/wmt19-de-en')
p("""Nicht nur unterschiedliche Berechnungen bereiten Kopfzerbrechen.
Bei der Eigenwahrnehmung zeigt sich: In Deutschland gibt es massive
Missverständnisse über Ausmaß und Art von Ungleichheit.""")
# + slideshow={"slide_type": "subslide"}
p = pipeline('translation', model='facebook/wmt19-en-de')
p("""We would like to belatedly welcome <NAME> of the Europa-Universität
Flensburg (#EUF), who is currently a guest at the Cluster. Ulrich has just decided
to extend his stay until the end of June, welcome news indeed!""")
# + slideshow={"slide_type": "subslide"}
p = pipeline('text-generation')
p("In Germany there are massive misunderstandings about the extent and type of inequality.")
# + slideshow={"slide_type": "subslide"}
p("some in Germany feel they have reached greater levels of economic equality without having")
# + [markdown] slideshow={"slide_type": "subslide"}
# Transformers can be "fine-tuned" to a specific task, see [training of transformers](https://huggingface.co/transformers/training.html). Adding a task-specific head to a transformer pre-trained on large amounts of training data (usually 100 GBs or even TBs of text) saves resources spent for training and can overcome the problem of not enough training data. Manually labelling training data is expensive and naturally puts a limit on the amount of training data. But even if the vocabulary in the training data is limited, there's a good chance that the pre-trained transformer has seen the unknown words in the huge data used for pre-training.
|
5_nlp_ml.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + id="imgZBevn_klO"
# # %tensorflow_version 1.x
# # !pip install --upgrade opencv-python==3.4.2.17
import numpy as np
import tensorflow as tf
import tensorflow.keras
import tensorflow.keras.backend as K
# import os
from tensorflow.keras.datasets import fashion_mnist,mnist,cifar10
# import keras.backend as K
from tensorflow.keras.layers import Conv2D,Activation,BatchNormalization,UpSampling2D,Embedding,ZeroPadding2D, Input, Flatten, Dense, Reshape, LeakyReLU, Dropout,MaxPooling2D
from tensorflow.keras.models import Sequential, Model
from tensorflow.keras.optimizers import Adam, SGD, RMSprop
from tensorflow.keras import regularizers
from tensorflow.keras.utils import Progbar
from keras.initializers import RandomNormal
import random
from sklearn.model_selection import train_test_split
# from keras.utils import np_utils
from tensorflow.keras import utils as np_utils
# + id="v0-DYwB8kiFk"
nb_classes = 10
batch_size = 128
maxepoches = 250
learning_rate = 0.1
lr_decay = 1e-6
lr_drop = 20
def lr_scheduler(epoch):
return learning_rate * (0.5 ** (epoch // lr_drop))
reduce_lr = tf.keras.callbacks.LearningRateScheduler(lr_scheduler)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 5338, "status": "ok", "timestamp": 1615396128353, "user": {"displayName": "Mus<NAME>.", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggo6kK28_a_Swx27ReDE7W6SlcMcsOsRyiTC_xFvA=s64", "userId": "11675938905762231877"}, "user_tz": -300} id="e04yi6rW_qJg" outputId="4674c6eb-b8bf-45ef-c180-42e9e51b71f2"
#Loading and splitting the dataset into train, validation and test
(X_Train, y_Train), (X_test, y_test) = cifar10.load_data()
X_train, X_val, y_train, y_val = train_test_split(X_Train, y_Train, test_size=0.20)
# convert y_train and y_test to categorical binary values
Y_train = np_utils.to_categorical(y_train, nb_classes)
Y_val = np_utils.to_categorical(y_val, nb_classes)
y_test = np_utils.to_categorical(y_test, nb_classes)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 3607, "status": "ok", "timestamp": 1615396128355, "user": {"displayName": "<NAME>.", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggo6kK28_a_Swx27ReDE7W6SlcMcsOsRyiTC_xFvA=s64", "userId": "11675938905762231877"}, "user_tz": -300} id="Lj_XM_dfmqnV" outputId="ac5a0bf9-7d8f-4f00-960b-8deb84052b90"
X_Train.shape
# + id="443_UL2p_qyQ"
# Reshape them to batch_size, width,height,#channels
X_train = X_train.reshape(40000, 32, 32, 3)
X_val = X_val.reshape(10000, 32, 32, 3)
X_test = X_test.reshape(10000, 32, 32, 3)
X_train = X_train.astype('float32')
X_val = X_val.astype('float32')
X_test = X_test.astype('float32')
# Normalize the values
X_train /= 255
X_val /= 255
X_test /= 255
# + id="5aQn2hUFNVDY"
init=RandomNormal(mean=0,stddev=0.02)
input_shape = (32, 32, 3) # Input shape of each image
weight_decay = 0.0005
def build_model():
# Build the network of vgg for 10 classes with massive dropout and weight decay as described in the paper.
model = Sequential()
model.add(Conv2D(64, (3, 3), padding='same',
input_shape=input_shape,kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.3))
model.add(Conv2D(64, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
model.add(Conv2D(128, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Conv2D(128, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
model.add(Conv2D(256, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Conv2D(256, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Conv2D(256, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
model.add(Conv2D(512, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Conv2D(512, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Conv2D(512, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
model.add(Conv2D(512, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Conv2D(512, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.4))
model.add(Conv2D(512, (3, 3), padding='same',kernel_regularizer=regularizers.l2(weight_decay)))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(MaxPooling2D(pool_size=(2, 2),padding='same'))
model.add(Dropout(0.5))
model.add(Flatten())
model.add(Dense(256,kernel_regularizer=regularizers.l2(weight_decay), name='dense_1'))
model.add(Activation('relu'))
model.add(BatchNormalization())
model.add(Dropout(0.5))
model.add(Dense(10, name='dense_2'))
model.add(Activation('softmax'))
return model
teacher = build_model()
sgd = SGD(lr=learning_rate, decay=lr_decay, momentum=0.9, nesterov=True)
teacher.compile(loss='categorical_crossentropy',optimizer=sgd, metrics=['accuracy'])
# + id="mp4il80HMRFn"
# teacher.fit(X_train,Y_train,batch_size=128,epochs=150,verbose=1,callbacks=[reduce_lr],validation_data=(X_val,Y_val))
# + id="JU8uZIWuQWKA"
teacher.load_weights("Cifar10_Teacher.h5")
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 37294, "status": "ok", "timestamp": 1615396192908, "user": {"displayName": "<NAME>.", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggo6kK28_a_Swx27ReDE7W6SlcMcsOsRyiTC_xFvA=s64", "userId": "11675938905762231877"}, "user_tz": -300} id="zBZO-MXLmml_" outputId="e50939c1-acc5-49e5-c358-b3ac99fdb8d5"
# Y_test = np_utils.to_categorical(y_test, nb_classes)
loss, acc =teacher.evaluate(X_test, y_test, verbose=1)
loss, acc
# + id="bPVxVj6L_sCz"
#Collect the dense vector from the previous layer output and store it in a different model
teacher_WO_Softmax = Model(teacher.input, teacher.get_layer('dense_1').output)
# + id="GhcEQ0Z-_scF"
#Extracting dense representation from the teacher network
train_dense = teacher_WO_Softmax.predict(X_train)
val_dense = teacher_WO_Softmax.predict(X_val)
# + id="XG0WGCpM_suF"
#Splitting the training dense vector among N students(in this case 2)
s1Train=train_dense[:,:64]
s2Train=train_dense[:,64:128]
s3Train=train_dense[:,128:192]
s4Train=train_dense[:,192:256]
s1Val=val_dense[:,:64]
s2Val=val_dense[:,64:128]
s3Val=val_dense[:,128:192]
s4Val=val_dense[:,192:256]
# + id="MKx8VcoJxwH0"
def define_model(name):
model = Sequential()
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same', input_shape=(32, 32, 3), name=name))
model.add(Conv2D(32, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.2))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(Conv2D(64, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.3))
model.add(Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(Conv2D(128, (3, 3), activation='relu', kernel_initializer='he_uniform', padding='same'))
model.add(MaxPooling2D((2, 2)))
model.add(Dropout(0.4))
model.add(Flatten())
model.add(Dense(16, activation='relu', kernel_initializer='he_uniform'))
model.add(Dropout(0.2))
model.add(Dense(64, activation='relu', kernel_initializer='he_uniform',name='req'+name))
model.compile(optimizer='nadam', loss='mse', metrics=['accuracy'])
return model
student1 = define_model('s1')
student1.summary()
# + [markdown] id="bY_42UFKHaZO"
# TCN
#
# +
# Testing with LR
s1=define_model("s1")
s2=define_model("s2")
s3=define_model("s3")
s4=define_model("s4")
opt=Adam(lr=0.0002, beta_1=0.9, beta_2=0.999, amsgrad=False)
# +
s1.compile(loss='mse', optimizer=opt)
s2.compile(loss='mse', optimizer=opt)
s1.fit(X_train,s1Train,
batch_size=256,
epochs=80,
verbose=1,
validation_data=(X_val,s1Val))
s2.fit(X_train,s2Train,
batch_size=256,
epochs=60,
verbose=1,
validation_data=(X_val,s2Val))
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 76533, "status": "ok", "timestamp": 1615397271851, "user": {"displayName": "<NAME>.", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggo6kK28_a_Swx27ReDE7W6SlcMcsOsRyiTC_xFvA=s64", "userId": "11675938905762231877"}, "user_tz": -300} id="tbiMHpxnHbd_" outputId="e017a372-0165-417b-fce6-f77671de9dfa"
s3.compile(loss='mse', optimizer=opt)
s4.compile(loss='mse', optimizer=opt)
s3.fit(X_train,s3Train,
batch_size=256,
epochs=70,
verbose=1,
validation_data=(X_val,s3Val))
s4.fit(X_train,s4Train,
batch_size=256,
epochs=60,
verbose=1,
validation_data=(X_val,s4Val))
# + id="mMTpeLnwINEn"
o1=s1.get_layer("reqs1").output
o2=s2.get_layer("reqs2").output
o3=s3.get_layer("reqs3").output
o4=s4.get_layer("reqs4").output
output=tensorflow.keras.layers.concatenate([o1,o2,o3,o4])
output=Activation('relu')(output)
output2=Dropout(0.5)(output) # For reguralization
output3=Dense(10,activation="softmax", name="d1")(output2)
mm4=Model([s1.get_layer("s1").input,s2.get_layer("s2").input,
s3.get_layer("s3").input,s4.get_layer("s4").input], output3)
my_weights=teacher.get_layer('dense_2').get_weights()
mm4.get_layer('d1').set_weights(my_weights)
# + colab={"base_uri": "https://localhost:8080/"} executionInfo={"elapsed": 102421, "status": "ok", "timestamp": 1615397435107, "user": {"displayName": "<NAME>.", "photoUrl": "https://lh3.googleusercontent.com/a-/AOh14Ggo6kK28_a_Swx27ReDE7W6SlcMcsOsRyiTC_xFvA=s64", "userId": "11675938905762231877"}, "user_tz": -300} id="q8bMaMg4LSpu" outputId="f7d52f3f-412d-4927-fd34-69341ee200a4"
i=0
for l in mm4.layers[:len(mm4.layers)-2]:
l.trainable=False
# print(l)
mm4.compile(loss='categorical_crossentropy',
optimizer=Adam(learning_rate=0.0002),
metrics=['accuracy'])
# Without finetune
batch_size = 256
mm4_history=mm4.fit([X_train,X_train,X_train,X_train], Y_train,
batch_size=batch_size,
epochs=50,
verbose=1,
validation_data=([X_val,X_val,X_val,X_val], Y_val))
|
CIFAR-10/Feed Forward Files/[4] Students on CIFAR10 Using FF.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# <h1> Text Classification using TensorFlow on Cloud ML Engine (using pretrained embedding) </h1>
#
# This notebook illustrates:
# <ol>
# <li> Creating datasets for Machine Learning using BigQuery
# <li> Creating a text classification model using the high-level Estimator API and a pre-trained embedding. (this is the difference vs <a href="txtcls1.ipyb"> txtcls1.ipynb </a>)
# <li> Training on Cloud ML Engine
# <li> Deploying model
# <li> Predicting with model
# </ol>
# change these to try this notebook out
BUCKET = 'cloud-training-demos-ml'
PROJECT = 'cloud-training-demos'
REGION = 'us-central1'
import os
os.environ['BUCKET'] = BUCKET
os.environ['PROJECT'] = PROJECT
os.environ['REGION'] = REGION
# !gcloud config set project $PROJECT
import tensorflow as tf
print tf.__version__
# The idea is to look at the title of a newspaper article and figure out whether the article came from the New York Times or from TechCrunch. Look at <a href="txtcls1.ipyb"> txtcls1.ipynb </a> for a solution that learns words embeddings as part of the problem itself. In this notebook, I will show how to use a pretrained embedding instead.
#
# <h2> Data exploration and preprocessing in BigQuery </h2>
# <p>
# See <a href="txtcls1.ipyb"> txtcls1.ipynb </a> for an explanation. Here, I simply repeat the key steps to create the dataset.
# +
import google.datalab.bigquery as bq
query="""
SELECT source, LOWER(REGEXP_REPLACE(title, '[^a-zA-Z0-9 $.-]', ' ')) AS title FROM
(SELECT
ARRAY_REVERSE(SPLIT(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.'))[OFFSET(1)] AS source,
title
FROM
`bigquery-public-data.hacker_news.stories`
WHERE
REGEXP_CONTAINS(REGEXP_EXTRACT(url, '.*://(.[^/]+)/'), '.com$')
AND LENGTH(title) > 10
)
WHERE (source = 'github' OR source = 'nytimes' OR source = 'techcrunch')
"""
traindf = bq.Query(query + " AND MOD(ABS(FARM_FINGERPRINT(title)),4) > 0").execute().result().to_dataframe()
evaldf = bq.Query(query + " AND MOD(ABS(FARM_FINGERPRINT(title)),4) = 0").execute().result().to_dataframe()
import os, shutil
DATADIR='data/txtcls2'
shutil.rmtree(DATADIR, ignore_errors=True)
os.makedirs(DATADIR)
traindf.to_csv( os.path.join(DATADIR,'train.csv'), header=False, index=False, encoding='utf-8', sep='\t')
evaldf.to_csv( os.path.join(DATADIR,'eval.csv'), header=False, index=False, encoding='utf-8', sep='\t')
# -
# %bash
gsutil cp data/txtcls2/*.csv gs://${BUCKET}/txtcls2/
# ## Pre-trained embedding
#
# To provide words as inputs to a neural network, we have to convert words to numbers. Ideally, we want related words to have numbers that are close to each other. This is what an embedding (such as word2vec) does. Here, I'll use the <a href="https://nlp.stanford.edu/projects/glove/">GloVe</a> embedding from Stanford just because, at 160MB, it is smaller than <a href="https://code.google.com/archive/p/word2vec/">word2vec</a> from Google (1.5 GB).
# <p>
# For testing purposes, I will also create a smaller file, consisting of the 1000 most common words.
# %bash
wget http://nlp.stanford.edu/data/glove.6B.zip
# +
# %bash
unzip -p glove.6B.zip glove.6B.50d.txt | gzip > pretrained_embedding.txt.gz
# #rm glove.6B.zip
# rm subset_embedding.txt*
zcat pretrained_embedding.txt.gz | head -1000 > subset_embedding.txt
gzip subset_embedding.txt
# -
# %bash
zcat subset_embedding.txt.gz | head -1
gsutil cp *_embedding.txt.gz gs://${BUCKET}/txtcls2/
# %bash
gsutil ls -l gs://${BUCKET}/txtcls2/*.txt.gz
# +
PADWORD = 'ZYXW'
from tensorflow.python.lib.io import file_io
class Word2Vec:
'''
vocab, embeddings
'''
def vocab_size(self):
return len(self.vocab)
def embed_dim(self):
return len(self.embeddings[0])
def __init__(self, filename):
import gzip, StringIO
import numpy as np
self.vocab = [PADWORD]
self.embeddings = [0]
with file_io.FileIO(filename, mode='rb') as f:
compressedFile = StringIO.StringIO(f.read())
decompressedFile = gzip.GzipFile(fileobj=compressedFile)
for line in decompressedFile:
pieces = line.split()
self.vocab.append(pieces[0])
self.embeddings.append(np.asarray(pieces[1:], dtype='float32'))
self.embeddings[0] = np.zeros_like(self.embeddings[1])
self.vocab.append('') # for out-of-value words
self.embeddings.append(np.ones_like(self.embeddings[1]))
self.embeddings = np.array(self.embeddings)
print('Loaded {}D vectors for {} words from {}'.format(self.embed_dim(), self.vocab_size(), filename))
# -
#wv = Word2Vec('gs://{}/txtcls2/pretrained_embedding.txt.gz'.format(BUCKET))
wv = Word2Vec('subset_embedding.txt.gz'.format(BUCKET))
print wv.embeddings.shape
# <h2> TensorFlow code </h2>
#
# Please explore the code in this <a href="txtcls2/trainer">directory</a> -- <a href="txtcls2/trainer/model.py">model.py</a> contains the key TensorFlow model and <a href="txtcls2/trainer/task.py">task.py</a> has a main() that launches off the training job.
#
# The following cells should give you an idea of what the model code does. The idea is to load up the embedding file and get vectors corresponding to the words in that file's vocabulary. For example "the" might be mapped to a 50D vector. Now, whenever we see "the" in the input document, we need to replace it by the 50D vector.
# The method that does this:
# <pre>
# tf.nn.embedding_lookup
# </pre>
# requires the *index* of the word "the" in the original file (perhaps that index=1). To find the index, we will use VocabularyProcessor.
# +
import tensorflow as tf
from tensorflow.contrib import lookup
from tensorflow.python.platform import gfile
import numpy as np
print tf.__version__
MAX_DOCUMENT_LENGTH = 5
# raw input
lines = ['Some title', 'A longer title', 'An even longer title', 'This is longer than doc length']
lines = [line.lower() for line in lines]
#lines = tf.constant(lines) # vocabprocessor doesn't work
# we first create word-ids for each of the words in the glove embedding file
vocab_processor = tf.contrib.learn.preprocessing.VocabularyProcessor(MAX_DOCUMENT_LENGTH)
vocab_processor.fit(wv.vocab) # word to word-id
wordid_to_embed = tf.convert_to_tensor(wv.embeddings) # word-id to embedding
# take lines of input and find word-ids; then lookup the embedding for each word-id
tensorids = np.array(list(vocab_processor.transform(lines)))
numbers = tf.nn.embedding_lookup(wordid_to_embed, tensorids)
with tf.Session() as sess:
print "numbers=", numbers.eval()[0], numbers.shape
# -
# However, [as pointed out by <NAME>](https://stackoverflow.com/questions/35687678/using-a-pre-trained-word-embedding-word2vec-or-glove-in-tensorflow), tf.constants are not memory efficient. To avoid storing multiple copies of the wordid_to_embed tensor, we should use a Variable.
# <p>
# Also, although the VocabularyProcessor has that convenient transform() method, it is pure Python and can not handle Tensors. Our "lines" will actually be a tensor in real-life. So, we have to use index_table and do a lookup using that ... This code also differs in how we handle "out-of-bucket" words -- we use ones (because PADWORD is mapped to zeros) whereas vocab processor uses zeros.
# +
import tensorflow as tf
from tensorflow.contrib import lookup
from tensorflow.python.platform import gfile
import numpy as np
print tf.__version__
MAX_DOCUMENT_LENGTH = 5
# raw input
lines = ['Some title', 'A longer title', 'An even longer title', 'This is longer than doc length']
lines = [line.lower() for line in lines]
lines = tf.constant(lines)
wordid_to_embed = tf.Variable(tf.constant(0.0, shape=[wv.vocab_size(), wv.embed_dim()]), trainable=False, name="embedding")
embedding_placeholder = tf.placeholder(tf.float32, [wv.vocab_size(), wv.embed_dim()])
embedding_init = wordid_to_embed.assign(embedding_placeholder)
# take lines of input and find word-ids; then lookup the embedding for each word-id
table = tf.contrib.lookup.index_table_from_tensor(tf.convert_to_tensor(wv.vocab[:-1]), num_oov_buckets=1)
words = tf.string_split(lines)
densewords = tf.sparse_tensor_to_dense(words, default_value=PADWORD)
numbers = table.lookup(densewords)
padding = tf.constant([[0,0],[0,MAX_DOCUMENT_LENGTH]])
padded = tf.pad(numbers, padding)
sliced = tf.slice(padded, [0,0], [-1, MAX_DOCUMENT_LENGTH])
embeds = tf.nn.embedding_lookup(wordid_to_embed, sliced)
with tf.Session() as sess:
tf.tables_initializer().run()
tf.get_default_session().run(embedding_init, feed_dict={embedding_placeholder: wv.embeddings})
print "embeds=", embeds.eval()[0], embeds.shape
# -
# %bash
grep -E "def |class " txtcls1/trainer/model.py
# Let's make sure the code works locally on a small dataset for a few steps. Because of the size of the graph, though, this will take a *long* time and may crash on smaller machines (it has to evaluate the graph five times and write out 5 checkpoints).
# %bash
# echo "bucket=${BUCKET}"
# rm -rf outputdir
export PYTHONPATH=${PYTHONPATH}:${PWD}/txtcls1
python -m trainer.task \
--bucket=${BUCKET} \
--output_dir=outputdir \
--glove_embedding=gs://${BUCKET}/txtcls2/subset_embedding.txt.gz \
--job-dir=./tmp --train_steps=200
# When I ran it, I got a 37% accuracy after a few steps. Once the code works in standalone mode, you can run it on Cloud ML Engine. You can monitor the job from the GCP console in the Cloud Machine Learning Engine section. Since we have 72,000 examples and batchsize=32, train_steps=36,000 essentially means 16 epochs.
# %bash
OUTDIR=gs://${BUCKET}/txtcls2/trained_model
JOBNAME=txtcls_$(date -u +%y%m%d_%H%M%S)
# echo $OUTDIR $REGION $JOBNAME
gsutil -m rm -rf $OUTDIR
gsutil cp txtcls1/trainer/*.py $OUTDIR
gcloud ml-engine jobs submit training $JOBNAME \
--region=$REGION \
--module-name=trainer.task \
--package-path=$(pwd)/txtcls1/trainer \
--job-dir=$OUTDIR \
--staging-bucket=gs://$BUCKET \
--scale-tier=BASIC_GPU \
--runtime-version=1.4 \
-- \
--bucket=${BUCKET} \
--output_dir=${OUTDIR} \
--glove_embedding=gs://${BUCKET}/txtcls2/pretrained_embedding.txt.gz \
--train_steps=36000
# Training finished with an accuracy of 54.5%.
# <h2> Deploy trained model </h2>
# <p>
# Deploying the trained model to act as a REST web service is a simple gcloud call.
# %bash
gsutil ls gs://${BUCKET}/txtcls2/trained_model/export/Servo/
# %bash
MODEL_NAME="txtcls"
MODEL_VERSION="v2"
MODEL_LOCATION=$(gsutil ls gs://${BUCKET}/txtcls1/trained_model/export/Servo/ | tail -1)
# echo "Deleting and deploying $MODEL_NAME $MODEL_VERSION from $MODEL_LOCATION ... this will take a few minutes"
#gcloud ml-engine versions delete ${MODEL_VERSION} --model ${MODEL_NAME}
#gcloud ml-engine models delete ${MODEL_NAME}
#gcloud ml-engine models create ${MODEL_NAME} --regions $REGION
gcloud ml-engine versions create ${MODEL_VERSION} --model ${MODEL_NAME} --origin ${MODEL_LOCATION}
# <h2> Use model to predict </h2>
# <p>
# Send a JSON request to the endpoint of the service to make it predict which publication the article is more likely to run in. These are actual titles of articles in the New York Times, github, and TechCrunch on June 19. These titles were not part of the training or evaluation datasets.
# +
from googleapiclient import discovery
from oauth2client.client import GoogleCredentials
import json
credentials = GoogleCredentials.get_application_default()
api = discovery.build('ml', 'v1', credentials=credentials,
discoveryServiceUrl='https://storage.googleapis.com/cloud-ml/discovery/ml_v1_discovery.json')
request_data = {'instances':
[
{
'title': 'Supreme Court to Hear Major Case on Partisan Districts'.lower()
},
{
'title': 'Furan -- build and push Docker images from GitHub to target'.lower()
},
{
'title': 'Time Warner will spend $100M on Snapchat original shows and ads'.lower()
},
]
}
parent = 'projects/%s/models/%s/versions/%s' % (PROJECT, 'txtcls', 'v2')
response = api.projects().predict(body=request_data, name=parent).execute()
print "response={0}".format(response)
# -
# Copyright 2017 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License
|
courses/machine_learning/deepdive/09_sequence/txtcls2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/jamestheengineer/data-science-from-scratch-Python/blob/master/Chapter_19.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="0a7a8s3fAKso"
# Only do this once per VM, otherwise you'll get multiple clones and nested directories
# #!git clone https://github.com/jamestheengineer/data-science-from-scratch-Python.git
# #%cd data-science-from-scratch-Python/
# #!pip install import-ipynb
#import import_ipynb
# + id="Bll3cxBdIujf"
# Deep learning chapter
Tensor = list
from typing import List
def shape(tensor: Tensor) -> List[int]:
sizes: List[int] = []
while isinstance(tensor, list):
sizes.append(len(tensor))
tensor = tensor[0]
return sizes
assert shape([1, 2, 3]) == [3]
assert shape([[1, 2], [3, 4], [5, 6]]) == [3, 2]
# + id="2In7rs_dJwae"
def is_1d(tensor: Tensor) -> bool:
"""
If tensor[0] is a list, it's a higher-order tensor.
Otherwise, tensor is 1-dimensional (that is, a vector)
"""
return not isinstance(tensor[0], list)
assert is_1d([1,2,3])
assert not is_1d([[1,2],[3,4]])
# + id="NiAIHUcUu8ep"
def tensor_sum(tensor: Tensor) -> float:
"""Sums up all the values in the tensor"""
if is_1d(tensor):
return sum(tensor) # just a list of floats, use Python sum
else:
return sum(tensor_sum(tensor_i) # Call tensor_sum on each row
for tensor_i in tensor) # and sum up those results
assert tensor_sum([1,2,3]) == 6
assert tensor_sum([[1,2],[3,4]]) == 10
# + id="mF97Cb4SvYXi"
from typing import Callable
def tensor_apply(f: Callable[[float], float], tensor: Tensor) -> Tensor:
"""Applies f elementwise"""
if is_1d(tensor):
return [f(x) for x in tensor]
else:
return [tensor_apply(f, tensor_i) for tensor_i in tensor]
assert tensor_apply(lambda x: x+1, [1, 2, 3]) == [2,3,4]
assert tensor_apply(lambda x: 2 * x, [[1,2],[3,4]]) == [[2,4],[6,8]]
# + id="IFSLmJ4JwifU"
def zeros_like(tensor: Tensor) -> Tensor:
return tensor_apply(lambda _: 0.0, tensor)
assert zeros_like([1,2,3]) == [0,0,0]
assert zeros_like([[1,2],[3,4]]) == [[0,0],[0,0]]
# + id="fvN1x2Gow_5v"
def tensor_combine(f: Callable[[float, float], float],
t1: Tensor,
t2: Tensor) -> Tensor:
"""Applies f to corresponding elements of t1 and t2"""
if is_1d(t1):
return [f(x,y) for x, y in zip(t1,t2)]
else:
return [tensor_combine(f, t1_i, t2_i)
for t1_i, t2_i in zip(t1, t2)]
import operator
assert tensor_combine(operator.add, [1,2,3],[4,5,6]) == [5,7,9]
assert tensor_combine(operator.mul, [1,2,3], [4,5,6]) == [4,10,18]
# + id="2sH1ExZIyHFP"
from typing import Iterable, Tuple
class Layer:
"""
Our neural networks will be composed of layers, each of which
knows how to do some computation on its inputs in the "forward"
direction and propagate gradients in the "backward" direction.
"""
def backward(self, gradient):
"""
Similarly, we're not going to be prescriptive about what the
gradient looks like. It's up to you the user to make sure
that you're doing things sensibly.
"""
raise NotImplementedError
def params(self) -> Iterable[Tensor]:
"""
Returns the parameters of this layer. The default implementation
returns nothing, so that if you have a layer with no parameters
you don't have to implement this.
"""
return ()
def grads(self) -> Iterable[Tensor]:
"""
Returns the gradients, in the smae order as params().
"""
return ()
# + id="z3Uh-B4REbEM"
from Chapter_18 import sigmoid
class Sigmoid(Layer):
def forward(self, input: Tensor) -> Tensor:
"""
Apply sigmoid to each element of the input tensor,
and save the results to use in backpropagation.
"""
self.sigmoids = tensor_apply(sigmoid, input)
return self.sigmoids
def backward(self, gradient: Tensor) -> Tensor:
return tensor_combine(lambda sig, grad: sig * (1 - sig) * grad,
self.sigmoids,
gradient)
# + id="aywa2pqdAwUP"
# Functions to randomly generate our weight tensors
import random
from Chapter_06 import inverse_normal_cdf
def random_uniform(*dims: int) -> Tensor:
if len(dims) == 1:
return [random.random() for _ in range(dims[0])]
else:
return [random_uniform(*dims[1:]) for _ in range(dims[0])]
def random_normal(*dims: int,
mean: float = 1.0,
variance: float = 1.0) -> Tensor:
if len(dims) == 1:
return [mean + variance * inverse_normal_cdf(random.random())
for _ in range(dims[0])]
else:
return [random_normal(*dims[1:], mean=mean, variance=variance)
for _ in range(dims[0])]
assert shape(random_uniform(2,3,4)) == [2,3,4]
assert shape(random_normal(5,6,mean=10)) == [5,6]
# + id="irGD2UmzWmW3"
# Wrap them all in a random_tensor function
def random_tensor(*dims: int, init: str = 'normal') -> Tensor:
if init == 'normal':
return random_normal(*dims)
elif init == 'uniform':
return random_uniform(*dims)
elif init == 'xavier':
variance = len(dims) / sum(dims)
return random_normal(*dims, variance=variance)
else:
raise ValueError(f"unknown init: {init}")
# + id="SxHZdBlSXq1U"
# Define the linear layer
from Chapter_04 import dot
class Linear(Layer):
def __init__(self, input_dim: int, output_dim: int, init: str = 'xavier') -> None:
"""
A layer of output_dim neurons, each with input_dim weights
(and a bias).
"""
self.input_dim = input_dim
self.output_dim = output_dim
# self.w[o] is the weights for the o-th neuron
self.w = random_tensor(output_dim, input_dim, init=init)
# self.b[o] is the bias term for the o-th neuron
self.b = random_tensor(output_dim, init=init)
def forward(self, input: Tensor) -> Tensor:
# Save the input to use in the backward pass.
self.input = input
# Return the vector of neuron outputs.
return [dot(input, self.w[o]) + self.b[o]
for o in range(self.output_dim)]
def backward(self, gradient: Tensor) -> Tensor:
# Each b[o] gets added to output[o], which means
# the gradient of b is the same as the output gradient.
self.b_grad = gradient
# Each w[o][i] multiplies input[i] and gets added to output[o].
# So its gradient is input[i] * gradient[o].
self.w_grad = [[self.input[i] * gradient[o]
for i in range(self.input_dim)]
for o in range(self.output_dim)]
# Each input[i] multiplies every w[o][i] and gets added to every
# output[o]. So its gradient is the sum of w[o][i] * gradient[o]
# across all the outputs.
return [sum(self.w[o][i] * gradient[o] for o in range(self.output_dim))
for i in range(self.input_dim)]
def params(self) -> Iterable[Tensor]:
return [self.w, self.b]
def grads(self) -> Iterable[Tensor]:
return [self.w_grad, self.b_grad]
# + id="meJVjeykOOrR"
# Define a sequence of layers
from typing import List
class Sequential(Layer):
"""
A layer consisting of a sequence of other layers.
It's up to you to make sure that the output of each layer
makes sense as the input to the next layer.
"""
def __init__(self, layers: List[Layer]) -> None:
self.layers = layers
def forward(self, input):
"""Just forward the input through the layers in order."""
for layer in self.layers:
input = layer.forward(input)
return input
def backward(self, gradient):
"""Just backpropagate the gradient through the layers in reverse."""
for layer in reversed(self.layers):
gradient = layer.backward(gradient)
return gradient
def params(self) -> Iterable[Tensor]:
"""Just return the params from each layer."""
return (param for layer in self.layers for param in layer.params())
def grads(self) -> Iterable[Tensor]:
"""Just return the grads from each layer."""
return (grad for layer in self.layers for grad in layer.grads())
# + id="YJF7Z1enxQzp"
# But we still need to train it, so more classes to write
class Loss:
def loss(self, predicted: Tensor, actual: Tensor) -> float:
"""How good are out predictions? (Larger numbers are worse.)"""
raise NotImplementedError
def gradient(self, predicted: Tensor, actual: Tensor) -> Tensor:
"""How does the loss change as the predictions change?"""
raise NotImplementedError
class SSE(Loss):
"""Loss function that computes the sum of the squared errors."""
def loss(self, predicted: Tensor, actual: Tensor) -> float:
# Compute the tensor of squared differences
squared_errors = tensor_combine(
lambda predicted, actual: (predicted - actual) ** 2,
predicted,
actual)
# And just add them up
return tensor_sum(squared_errors)
def gradient(self, predicted: Tensor, actual: Tensor) -> Tensor:
return tensor_combine(
lambda predicted, actual: 2 * (predicted - actual),
predicted,
actual)
sse_loss = SSE()
assert sse_loss.loss([1, 2, 3], [10, 20, 30]) == 9 ** 2 + 18 ** 2 + 27 ** 2
assert sse_loss.gradient([1, 2, 3], [10, 20, 30]) == [-18, -36, -54]
# + id="wcm3xqmz3QKM"
# Need an absract optimizer so we can create different variants of gradient descent
class Optimizer:
"""
An optimizer updates the weights of a layer (in place) using information
known by either the layer or the optimizer (or by both).
"""
def step(self, layer: Layer) -> None:
raise NotImplementedError
class GradientDescent(Optimizer):
def __init__(self, learning_rate: float = 0.1) -> None:
self.lr = learning_rate
def step(self, layer: Layer) -> None:
for param, grad in zip(layer.params(), layer.grads()):
# Update param using a gradient step
param[:] = tensor_combine(
lambda param, grad: param - grad * self.lr,
param,
grad)
# Note the slice operator. You have to use this if you want to affect the original list. For example
tensor = [[1,2],[3,4]]
for row in tensor:
row = [0,0]
assert tensor == [[1,2],[3,4]], "assignment doesn't update a list"
for row in tensor:
row[:] = [0,0]
assert tensor == [[0,0], [0,0]], "but slice assignment does"
# + id="hjm1GougwwY_"
# Let's show the flexibility of this design by implmenting one more optimizer
class Momentum(Optimizer):
def __init__(self,
learning_rate: float,
momentum: float = 0.9) -> None:
self.lr = learning_rate
self.mo = momentum
self.updates: List[Tensor] = [] # running average
def step(self, layer: Layer) -> None:
# If we have no previous updates, start with all zeros.
if not self.updates:
self.updates = [zeros_like(grad) for grad in layer.grads()]
for update, param, grad in zip(self.updates,
layer.params(),
layer.grads()):
# Apply momentum
update[:] = tensor_combine(
lambda u, g: self.mo * u + (1 - self.mo) * g,
update,
grad)
# Then take a gradient step
param[:] = tensor_combine(
lambda p, u: p - self.lr * u,
param,
update)
# + id="FIfK4VQe2Cpq"
# Let's revisit XOR
# training data
xs = [[0., 0], [0., 1], [1., 0], [1., 1]]
ys = [[0.], [1.], [1.], [0.]]
random.seed(0)
net = Sequential([
Linear(input_dim=2, output_dim=2),
Sigmoid(),
Linear(input_dim=2, output_dim=1)
])
import tqdm
optimizer = GradientDescent(learning_rate=0.1)
loss = SSE()
#with tqdm.trange(3000) as t:
# for epoch in t:
# epoch_loss = 0.0
# for x, y in zip(xs, ys):
# predicted = net.forward(x)
# epoch_loss += loss.loss(predicted, y)
# gradient = loss.gradient(predicted, y)
# net.backward(gradient)
# optimizer.step(net)
# t.set_description(f"xor loss {epoch_loss:.3f}")
#for param in net.params():
# print(param)
# + id="LvAHjZQD-cJC"
# One replacement for sigmoid is tanh
import math
def tanh(x: float) -> float:
# If x is very large or very small, tanh is (essentially) 1 or -1.
# We check for this because, e.g., math.exp(1000) raises an error.
if x < -100: return -1
elif x > 100: return 1
em2x = math.exp(-2 * x)
return (1 - em2x) / (1 + em2x)
class Tanh(Layer):
def forward(self, input: Tensor) -> Tensor:
# Save tanh output to use in backward pass.
self.tanh = tensor_apply(tanh, input)
return self.tanh
def backward(self, gradient: Tensor) -> Tensor:
return tensor_combine(
lambda tanh, grad: (1 - tanh ** 2) * grad,
self.tanh,
gradient)
# + id="bDVcd2KfbEht"
# Another popular one is relu
class Relu(Layer):
def forward(self, input: Tensor) -> Tensor:
self.input = input
return tensor_apply(lambda x: max(x, 0), input)
def backward(self, gradient: Tensor) -> Tensor:
return tensor_combine(lambda x, grad: grad if x > 0 else 0,
self.input,
gradient)
# + id="CfGlOtBo87mk"
# Fizzbuzz revisted
from Chapter_18 import binary_encode, fizz_buzz_encode, argmax
xs = [binary_encode(n) for n in range(101, 1024)]
ys = [fizz_buzz_encode(n) for n in range(101, 1024)]
NUM_HIDDEN = 25
random.seed(0)
def fizzbuzz_accuracy(low: int, hi: int, net: Layer) -> float:
num_correct = 0
for n in range(low, hi):
x = binary_encode(n)
predicted = argmax(net.forward(x))
actual = argmax(fizz_buzz_encode(n))
if predicted == actual:
num_correct += 1
return num_correct / (hi - low)
# + id="mTx8Ldwjbi9l"
net = Sequential([
Linear(input_dim=10, output_dim=NUM_HIDDEN, init='uniform'),
Tanh(),
Linear(input_dim=NUM_HIDDEN, output_dim=4, init="uniform"),
Sigmoid()
])
optimizer = Momentum(learning_rate=0.1, momentum=0.9)
loss = SSE()
#with tqdm.trange(1000) as t:
# for epoch in t:
# epoch_loss = 0.0
# for x, y in zip(xs, ys):
# predicted = net.forward(x)
# epoch_loss += loss.loss(predicted, y)
# gradient = loss.gradient(predicted, y)
# net.backward(gradient)
# optimizer.step(net)
# accuracy = fizzbuzz_accuracy(101, 1024, net)
# t.set_description(f"fb loss: {epoch_loss:.2f} acc: {accuracy:.2f}")
# Now check results on the test set
#print("test results", fizzbuzz_accuracy(1, 101, net))
# + id="OIRaAo01iSBc"
def softmax(tensor: Tensor) -> Tensor:
"""Softmad along the last dimension"""
if is_1d(tensor):
# Subtract the largest value for numerical stability.
largest = max(tensor)
exps = [math.exp(x - largest) for x in tensor]
sum_of_exps = sum(exps) # This is the total "weight"
return [exp_i / sum_of_exps # Probability is the fraction
for exp_i in exps] # of the total weight
else:
return [softmax(tensor_i) for tensor_i in tensor]
class SoftmaxCrossEntropy(Loss):
"""
This is the negative-log-likelihood of the ovserved values, given the
neural net model. So if we choose weights ot minimize it, out model will
be maximinzing the likelihood of the observed data.
"""
def loss(self, predicted: Tensor, actual: Tensor) -> float:
# Apply softmax to get probabilities
probabilities = softmax(predicted)
# This will be log p_i for the actual class i and 0 for the other
# classes. We add a tiny amount to p to avoid taking log(0).
likelihoods = tensor_combine(lambda p, act: math.log(p + 1e-30) * act,
probabilities,
actual)
# And then we just sum up the negatives.
return -tensor_sum(likelihoods)
def gradient(self, predicted: Tensor, actual: Tensor) -> Tensor:
probabilities = softmax(predicted)
# Isn't this a pleasant equation:
return tensor_combine(lambda p, actual: p - actual,
probabilities,
actual)
# + id="gJln8vzcj4bC"
net = Sequential([
Linear(input_dim=10, output_dim=NUM_HIDDEN, init='uniform'),
Tanh(),
Linear(input_dim=NUM_HIDDEN, output_dim=4, init="uniform"),
# No final Sigmoid layer now
])
optimizer = Momentum(learning_rate=0.1, momentum=0.9)
loss = SoftmaxCrossEntropy()
#with tqdm.trange(1000) as t:
# for epoch in t:
# epoch_loss = 0.0
# for x, y in zip(xs, ys):
# predicted = net.forward(x)
# epoch_loss += loss.loss(predicted, y)
# gradient = loss.gradient(predicted, y)
# net.backward(gradient)
# optimizer.step(net)
# accuracy = fizzbuzz_accuracy(101, 1024, net)
# t.set_description(f"fb loss: {epoch_loss:.2f} acc: {accuracy:.2f}")
# Now check results on the test set
#print("test results", fizzbuzz_accuracy(1, 101, net))
# + id="NduyXxAnKmG2"
# Dropout is another method, like regularization, to avoid over fitting data
class Dropout(Layer):
def __init__(self, p: float) -> None:
self.p = p
self.train = True
def forward(self, input: Tensor) -> Tensor:
if self.train:
# Create a mask of 0s and 1s shaped like the input
# using the specified proabability
self.mask = tensor_apply(
lambda _: 0 if random.random() < self.p else 1,
input)
# Mulitple by the mask to dropout inputs
return tensor_combine(operator.mul, input, self.mask)
else:
# During evaluation just scale down the outputs uniformly.
return tensor_apply(lambda x: x * (1 - selp.p), input)
def backward(self, gradient: Tensor) -> Tensor:
if self.train:
# Only propagate the gradients where mask == 1
return tensor_combine(operator.mul, gradient, self.mask)
else:
raise RunTimeError("don't call backward when not in train mode")
# + id="HFJodByp9pRt"
# MNIST example
# !pip install mnist
# + id="fejjNsD-9zmo"
import mnist
# This will download the data; change this to where you want it.
# Yes, it's a 0-argument function, that's what the library expects.
# Yes, I'm assigning a lambda to a variable, like I said never to do
# !pwd
# + id="5-_P8cQE-qu2"
mnist.temporary_dir = lambda: '/content/data-science-from-scratch-Python'
# Each of these functions first downloads the data and returns a numpy array
# We call .tolist() because our tensors are just lists
train_images = mnist.train_images().tolist()
train_labels = mnist.train_labels().tolist()
assert shape(train_images) == [60000, 28, 28]
assert shape(train_labels) == [60000]
# + id="oH7etgER_Shh"
import matplotlib.pyplot as plt
fig, ax = plt.subplots(10,10)
for i in range(10):
for j in range(10):
# Plot each image in black and white and hide the axes.
ax[i][j].imshow(train_images[10 * i + j], cmap='Greys')
ax[i][j].xaxis.set_visible(False)
ax[i][j].yaxis.set_visible(False)
plt.show()
# + id="GWbjzvhXACSM"
# Load the test images
test_images = mnist.test_images().tolist()
test_labels = mnist.test_labels().tolist()
assert shape(test_images) == [10000, 28, 28]
assert shape(test_labels) == [10000]
# + id="tlx3JyiFA_K5"
# We need to flatten, normalize, and re-center our data
# Compute the average pixel value
avg = tensor_sum(train_images) / 60000 / 28 / 28
# Recenter, rescale, and flatten
train_images = [[(pixel - avg) / 256 for row in image for pixel in row]
for image in train_images]
test_images = [[(pixel - avg) / 256 for row in image for pixel in row]
for image in test_images]
assert shape(train_images) == [60000, 784], "images should be flattened"
assert shape(test_images) == [10000, 784], "images should be flattened"
# After centering, average pixel should be very close to zero
assert -0.0001 < tensor_sum(train_images) < 0.0001
# + id="f4izS9AgCMXc"
# We also want to one hot encode the targets
def one_hot_encode(i: int, num_labels: int = 10) -> List[float]:
return [1.0 if j == i else 0.0 for j in range(num_labels)]
assert one_hot_encode(3) == [0,0,0,1,0,0,0,0,0,0]
assert one_hot_encode(2, num_labels=5) == [0,0,1,0,0]
train_labels = [one_hot_encode(label) for label in train_labels]
test_labels = [one_hot_encode(label) for label in test_labels]
assert shape(train_labels) == [60000, 10]
assert shape(test_labels) == [10000, 10]
# + id="5e_Xr0yEFpML"
import tqdm
def loop(model: Layer,
images: List[Tensor],
labels: List[Tensor],
loss: Loss,
optimizer: Optimizer = None) -> None:
correct = 0 # Track number of correct predictions.
total_loss = 0.0 # Track total loss
# with tqdm.trange(len(images)) as t:
# for i in t:
# predicted = model.forward(images[i]) # Predict
# if argmax(predicted) == argmax(labels[i]): # Check for
# correct += 1 # correctness
# total_loss += loss.loss(predicted, labels[i]) # Compute loss
# If we are training, backpropagate gradient and update weights
# if optimizer is not None:
# gradient = loss.gradient(predicted, labels[i])
# model.backward(gradient)
# optimizer.step(model)
# And update our metrics in the progress bar
# avg_loss = total_loss / (i + 1)
# acc = correct / (i + 1)
# t.set_description(f"mnist loss: {avg_loss:.3f} acc: {acc:.3f}")
# + id="yzb7yajqH0cp"
random.seed(0)
# Logistic regression is just a linear layer followed by softmax
model = Linear(784, 10)
loss = SoftmaxCrossEntropy()
# This optimizer seems to work
optimizer = Momentum(learning_rate=0.01, momentum=0.99)
# Train on the trianing data
loop(model, train_images, train_labels, loss, optimizer)
# Test on the test data (no optimizer means just evaluate)
loop(model, test_images, test_labels, loss)
# + id="nmjIv7cYIbU2"
# Let's see if we can do better with a DNN
random.seed(0)
# Name them so we can turn trian on and off
dropout1 = Dropout(0.1)
dropout2 = Dropout(0.1)
model = Sequential([
Linear(784, 30), # Hidden layer 1: size 30
dropout1,
Tanh(),
Linear(30, 10), # Hidden layer 2: size 10
dropout2,
Tanh(),
Linear(10,10) # Output layer
])
# And we can use the same training loop
optimizer = Momentum(learning_rate=0.01, momentum=0.99)
loss = SoftmaxCrossEntropy()
# Enable dropout and train (takes > 20 min on my laptop)
dropout1.train = dropout2.train = True
loop(model, train_images, train_labels, loss, optimizer)
# Disable dropout and evaluate
dropout1 = dropout2 = False
loop(model, test_images, test_labels, loss)
# + id="Aur2fDG1J-HM"
# Saving and loading models
import json
def save_weights(model: Layer, filename: str) -> None:
weights = list(model.params())
with open(filename, 'w') as f:
json.dump(weights, f)
def load_weights(model: Layer, filename: str) -> None:
with open(filename) as f:
weights = json.load(f)
# Check for consistency
assert all(shape(param) == shape(weight)
for param, weight in zip(model.params(), weights))
# Then load using slice assignment
for param, weight in zip(model.params(), weights):
param[:] = weight
# + id="46s5Cj8eK95w"
|
Chapter_19.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generating adversarial examples on MNIST using C&W and DDN
#
# In this notebook we will generate adversarial examples on MNIST using two methods:
#
# * Carlini and Wagner (C&W) L2 attack (https://arxiv.org/abs/1608.04644)
# * Decoupled Direction and Norm (DDN) (https://arxiv.org/abs/1811.09600)
#
# We will attack a robust model trained with DDN - the noise required to attack an image is quite noticeable for this model.
#
# Note: this example requires matplotlib (pip install matplotlib)
# +
import argparse
import torch
import time
from torch.utils import data
from torchvision import datasets, transforms
from torchvision.utils import save_image, make_grid
from fast_adv.models.mnist import SmallCNN
from fast_adv.attacks import DDN, CarliniWagnerL2
from fast_adv.utils import requires_grad_, l2_norm
import matplotlib.pyplot as plt
import os
# %matplotlib inline
torch.manual_seed(42)
device = torch.device('cuda:0' if torch.cuda.is_available() else 'cpu')
data_path = 'data/mnist' # Change this if you already downloaded MNIST elsewhere
model_url = 'https://www.dropbox.com/s/9onr3jfsuc3b4dh/mnist.pth?dl=1'
model_path = 'mnist.pth'
# +
# Loading the data
dataset = datasets.MNIST(data_path, train=False,
transform=transforms.ToTensor(),
download=True)
loader = data.DataLoader(dataset, shuffle=False, batch_size=16)
x, y = next(iter(loader))
x = x.to(device)
y = y.to(device)
plt.imshow(make_grid(x, nrow=16).permute(1,2,0))
plt.axis('off');
# +
print('Loading model')
if not os.path.exists(model_path):
import urllib
print('Downloading model')
urllib.request.urlretrieve(model_url, model_path)
model = SmallCNN()
model.load_state_dict(torch.load(model_path))
model.eval().to(device)
requires_grad_(model, False)
# +
print('Running DDN 100 attack')
attacker = DDN(steps=100, device=device)
start = time.time()
ddn_atk = attacker.attack(model, x, labels=y, targeted=False)
ddn_time = time.time() - start
print('Completed in {:.2f}s'.format(ddn_time))
plt.imshow(make_grid(ddn_atk, nrow=16).permute(1,2,0))
plt.axis('off');
# +
print('Running C&W 4 x 25 attack (limited to 100 iterations)')
cwattacker100 = CarliniWagnerL2(device=device,
image_constraints=(0, 1),
num_classes=10,
search_steps=4,
max_iterations=25,
learning_rate=0.5,
initial_const=1.0)
start = time.time()
cw100_atk = cwattacker100.attack(model, x, labels=y, targeted=False)
cw100_time = time.time() - start
print('Completed in {:.2f}s'.format(cw100_time))
plt.imshow(make_grid(cw100_atk, nrow=16).permute(1,2,0))
plt.axis('off');
# +
print('Running C&W 9 x 10000 attack')
cwattacker = CarliniWagnerL2(device=device,
image_constraints=(0, 1),
num_classes=10)
start = time.time()
cw_atk = cwattacker.attack(model, x, labels=y, targeted=False)
cw_time = time.time() - start
print('Completed in {:.2f}s'.format(cw_time))
plt.imshow(make_grid(cw_atk, nrow=16).permute(1,2,0))
plt.axis('off');
# +
all_imgs = torch.cat((x, cw100_atk, cw_atk, ddn_atk))
img_grid = make_grid(all_imgs, nrow=16, pad_value=0)
plt.imshow(img_grid.permute(1,2,0))
plt.axis('off')
# Print metrics
pred_orig = model(x).argmax(dim=1).cpu()
pred_cw = model(cw_atk).argmax(dim=1).cpu()
pred_cw100 = model(cw100_atk).argmax(dim=1).cpu()
pred_ddn = model(ddn_atk).argmax(dim=1).cpu()
print('C&W 4 x 25 done in {:.1f}s: Success: {:.2f}%, Mean L2: {:.4f}.'.format(
cw100_time,
(pred_cw100 != y.cpu()).float().mean().item() * 100,
l2_norm(cw100_atk - x).mean().item()
))
print('C&W 9 x 10000 done in {:.1f}s: Success: {:.2f}%, Mean L2: {:.4f}.'.format(
cw_time,
(pred_cw != y.cpu()).float().mean().item() * 100,
l2_norm(cw_atk - x).mean().item()
))
print('DDN 100 done in {:.1f}s: Success: {:.2f}%, Mean L2: {:.4f}.'.format(
ddn_time,
(pred_ddn != y.cpu()).float().mean().item() * 100,
l2_norm(ddn_atk - x).mean().item()
))
print()
print('Figure: top row: original images; 2nd: C&W 4x25 atk; 3rd: C&W 9x10000 atk; 4th: DDN 100 atk')
|
examples/mnist_noteboook_example.ipynb
|