code
stringlengths 38
801k
| repo_path
stringlengths 6
263
|
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# import dependencies
import pandas as pd
import os
import requests
from splinter import Browser
from bs4 import BeautifulSoup
from selenium import webdriver
import time
# # Collect the latest News Title and Paragraph Text
executable_path = {'executable_path': 'chromedriver.exe'}
browser = Browser('chrome', **executable_path, headless=False)
# URL of page to be scraped
url = "https://mars.nasa.gov/news/"
browser.visit(url)
# Create BeautifulSoup object; parse with 'html.parser'
html = browser.html
soup = BeautifulSoup(html, 'html.parser')
# getting latest news title and paragraph from nasa news url
news_title = soup.find("div",class_="content_title").text
print(f"News Title: {news_title}")
news_p = soup.find("div", class_="article_teaser_body").text
print(f"News Paragraph: {news_p}")
# # # JPL Mars Space Images - Featured Image
space_image_url = "https://www.jpl.nasa.gov/spaceimages/?search=&category=Mars"
browser.visit(space_image_url)
space_image_html = browser.html
soup = BeautifulSoup(space_image_html,"html.parser")
image = soup.find('img', class_ = 'thumb')['src']
featured_image_url = "https://www.jpl.nasa.gov" + image
print(f"Space Image URL : {featured_image_url}")
# # Mars Weather
# scrape the latest Mars weather tweet
mars_weather_url = "https://twitter.com/marswxreport?lang=en"
browser.visit(mars_weather_url)
# Create BeautifulSoup object; parse with 'html.parser'
weather_html = browser.html
soup = BeautifulSoup(weather_html, 'html.parser')
# Save the tweet text for the weather report as a variable called
mars_weather = soup.find("p", class_="TweetTextSize TweetTextSize--normal js-tweet-text tweet-text").text
print(f"Mars Weather Tweet : {mars_weather}")
# # ## Mars Facts
mars_fact_url = "https://space-facts.com/mars/"
# browser.visit(mars_fact_url)
mars_table = pd.read_html(mars_fact_url)
# print(mars_table)
df = mars_table[0]
df.columns = ['description', 'values']
df = df.set_index('description')
df.head(15)
mars_facts = df.to_html()
print(mars_facts)
# saving the html file as table.hyml
df.to_html('table.html')
# # Mars Hemispheres
hemispheres_url = "https://astrogeology.usgs.gov/search/results?q=hemisphere+enhanced&k1=target&v1=Mars"
browser.visit(hemispheres_url)
hemisphere_html = browser.html
soup = BeautifulSoup(hemisphere_html, 'html.parser')
# +
hemisphere_list = []
category = soup.find("div", class_ = "result-list" )
hemispheres =category.find_all("div", class_="item")
for hemisphere in hemispheres:
title = hemisphere.find("h3").text
title = title.strip("Enhanced")
end_link = hemisphere.find("a")["href"]
image_link = "https://astrogeology.usgs.gov/" + end_link
browser.visit(image_link)
hemisphere_html = browser.html
soup = BeautifulSoup(hemisphere_html, 'html.parser')
downloads = soup.find("div", class_="downloads")
image_url = downloads.find("a")["href"]
hemisphere_list.append({"title": title, "img_url": image_url})
hemisphere_list
# -
|
mission_to_mars.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [new2]
# language: python
# name: Python [new2]
# ---
from numpy import roots
from math import atan, fabs
def cubic (Qx, Qy, Ux, Uy):
a = Uy
b = (Ux + 2*Qy)
c = (2*Qx - Uy)
d = -Ux
det = -4*b**3*d + b**2*c**2 -4*a*c**3 + 18*a*b*c*d - 27*a**2*d**2
print det
if (det < 0):
answer = 'c'
print answer
if (det > 0):
a = roots([a, b, c, d])
a = a.real
print a
a = atan(a[0])/(fabs(atan(a[0]))) + atan(a[1])/(fabs(atan(a[1]))) + atan(a[2])/(fabs(atan(a[2])))
if (a == 3.0):
answer = 'b'
if (a == 1.0):
answer = 'b'
if (a == -1.0):
answer = 'a'
print answer
#return answer
|
outdated/0.1.1/cubic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Immediate Task: Externalize all helper functions
# # Satisfactory
from scipy.io import loadmat, savemat
import matplotlib.pyplot as plt
import numpy as np
A = loadmat('/Users/hayden/Desktop/norm/normalized_data/norm_M_552_256.mat')
A = A["M"]
plt.imshow(A)
np.mean(A)
np.std(A)
# +
# ! mkdir normalized_data
# ! mkdir normalized_data/medium
# +
from scipy.io import loadmat, savemat
import numpy as np
import glob, os
#np.std(A)
def normalize2mat(suffix = "_1024"):
prefix_lst = ["Intensity"]
pathDict = {}
TopDir = "./"
for prefix in prefix_lst:
for file in glob.glob(TopDir + size + prefix + "*.mat"):
print(file)
pathDict[prefix] = file
path_ = pathDict["Intensity"]
loaded_data = loadmat(file_)
out_path = "normalized_data/" + size + "Intensity" + suffix + ".mat"
A_new = loaded_data["M"].copy()
A_mean, A_std = np.mean(A_new), np.std(A_new)
A_new = (A_new - A_mean) / A_std
mDictOut = {"M" : A_new, "orig_mean" : A_mean, "orig_std" : A_std, "M_orig" : loaded_data["M"]}
savemat(file_name = out_path, mdict =mDictOut)
normalize2mat()
# -
tmp = loadmat("normalized_data/medium/Intensity_1024.mat")
for ke in tmp.keys():
print(ke)
# +
# %matplotlib inline
#reservoir.__file__
# %run -i '../MARIOS/PyFiles/imports.py'
# %run -i '../MARIOS/PyFiles/helpers.py'
# %run -i "../MARIOS/PyFiles/experiment.py"
experiment = EchoStateExperiment(size = "medium", target_frequency = 2000,
obs_hz = 500, target_hz = 500, verbose = False)
experiment.get_observers(method = "freq", split = 0.5, aspect = 0.9, plot_split = True)
bounds = {
'llambda' : -10,#(-12, 1),
'connectivity': (-3, 0), # 0.5888436553555889,
'n_nodes': 100,#(100, 1500),
'spectral_radius': (0.05, 0.99),
'regularization': (-10,-2)#(-12, 1)
#all are log scale except spectral radius and n_nodes
}
#example cv args:
cv_args = {
'bounds' : bounds,
'initial_samples' : 100,
'subsequence_length' : 250, #150 for 500
'eps' : 1e-5,
'cv_samples' : 4,
'max_iterations' : 1000,
'scoring_method' : 'tanh',
"n_jobs" : 8
}
#experiment.RC_CV(cv_args = cv_args, model = "uniform")
# -
# # Next add in capability for interpolation, hybrid, then debug and send to Marios. By next week have this built into your package, in fact that's likely the next step.
# +
# #!python execute.py
def load_data(file = "default"):
if file == "default":
nf = get_new_filename(exp = exp, current = True)
else:
nf = file
with open(nf) as json_file: # 'non_exp_w.txt'
datt = json.load(json_file)
#datt = non_exp_best_args["dat"]
#datt["obs_tr"], datt["obs_te"] = np.array(datt["obs_tr"]), np.array(datt["obs_te"])
#datt["resp_tr"], datt["resp_te"] = np.array(datt["resp_tr"]), np.array(datt["resp_te"])
return(datt)
#experiment.save_json(exp = False)
bp = "/Users/hayden/Desktop/DL_LAB/Reservoir/MARIOS/"
fp = bp + 'experiment_results/2k/medium/split_0.5/targetKhz:_0.01__obskHz:_0.03.txt'
hi = load_data(file = fp)
for i in hi.keys():
print(i)
if type(hi[i]) == dict:
for j in hi[i].keys():
print(" " +j)
#hi["prediction"]["exponential"]
display(hi["best arguments"])
# +
hi = {"delme" : 0}
# -
display(experiment.Train.shape)
display(experiment.Test.shape)
display(experiment.xTr.shape)
display(experiment.xTe.shape)
print(experiment.target_frequency)
def Freq2idx(val):
"""
Translates a desired target frequency into a desired index
"""
target_frequency = min(range(len(experiment.f)), key=lambda i: abs(experiment.f[i] - val))
return(target_frequency)
def hz2idx(midpoint,
#method = "all",
obs_idx = None,
obs_hz = None,
target_hz = None, silent = True):
"""
This function acts as a helper function to simple_block and get_observers
and is thus designed. It takes a desired hz amount and translates that to indices of the data.
To do one frequency use Freq2idx.
"""
# [lb, ub] stands for [lowerbound, upperbound]
def endpoints2list(lb, ub):
return list(range(int(lb), int(ub + 1)))
# spread vs total hz
obs_spread, target_spread = obs_hz / 2, target_hz / 2
# get the response range endpoints
respLb, respUb = [Freq2idx(midpoint - target_spread),
Freq2idx(midpoint + target_spread)]
# Listify it:
resp_idx_Lst = endpoints2list(respLb, respUb)
resp_Freq_Lst = [experiment.f[i] for i in resp_idx_Lst]
#get the observer range endpoints:
obs_high_Ub, obs_high_lb = respUb + Freq2idx(obs_spread) + 1, respUb + 1
obs_low_lb, obs_low_Ub = respLb - Freq2idx(obs_spread) - 1, respLb - 1
# create the equivilant observer lists
obs_idx_Lst1 = endpoints2list(obs_low_lb, obs_low_Ub)
obs_idx_Lst2 = endpoints2list(obs_high_lb, obs_high_Ub)
obs_Freq_Lst1 = [experiment.f[i] for i in obs_idx_Lst1]
obs_Freq_Lst2 = [experiment.f[i] for i in obs_idx_Lst2]
if silent != True:
print("resp_indexes : " + str(resp_idx_Lst))
print("observer frequencies upper domain: " + str(resp_Freq_Lst) +
" , range: "+ str(abs(resp_Freq_Lst[0] - resp_Freq_Lst[-1])) +" Hz\n")
print("observer indexes lower domain: " + str(obs_idx_Lst1))
print("observer frequencies lower domain: " + str(obs_Freq_Lst1) +
" , range: "+ str(abs(obs_Freq_Lst1[0] - obs_Freq_Lst1[-1])) +" Hz\n")
print("observer indexes upper domain: " + str(obs_idx_Lst2))
print("observer frequencies upper domain: " + str(obs_Freq_Lst2) +
" , range: "+ str(abs(obs_Freq_Lst2[0] - obs_Freq_Lst2[-1])) +" Hz\n")
assert obs_idx_Lst1 + resp_idx_Lst + obs_idx_Lst2 == list(range(obs_idx_Lst1[0],
obs_idx_Lst2[-1]+1))
dict2Return = {"obs_idx": obs_idx_Lst1 + obs_idx_Lst2,
"resp_idx": resp_idx_Lst,
"obs_freq" : obs_Freq_Lst1 + obs_Freq_Lst2,
"resp_freq" : resp_Freq_Lst}
return(dict2Return)
hz2idx(2000, obs_hz = 40, target_hz = 55)
# +
#fig, ax = plt.subplots(1, 1, figsize = (6,5))
#experiment.olab_display(ax)
display(experiment.dat["resp_idx"])
display(experiment.dat["obs_idx"])
# -
# # Still needs work:
# +
size, obs = "medium", "large_obs"
def simple_block(target_frequency, target_spread = None, n_obs = None): #self
ctr = experiment.key_freq_idxs[target_frequency]
if target_spread != None:
#resp_bounds is the response bounds ie the target area bounds.
resp_bounds = [ctr - target_spread, ctr + target_spread]
#resp_bounds = [[resp_bounds[0], resp_bounds[1]]]
else: #TODO does this part of the if-else statement actually do anything?
response_bounds = None
resp_bounds = [ctr, ctr]
assert n_obs != None, "if you want to have no observers then #TODO"
print("resp_bounds : " + str(resp_bounds))
obs_bounds = [[resp_bounds[0] - n_obs, resp_bounds[0]],
[resp_bounds[1], resp_bounds[1] + n_obs ]]
bounds = {"response_bounds" : resp_bounds, "observer_bounds" : obs_bounds}
return(bounds) # self.block_bounds_dict
#TODO: block needs to be redefined and self contained such that it is more simply defined.
# the current method is simpler but has a serious flaw ie it's too complex.
# the redefinition should need only 1) the target frequency, the spread, and the number of observers.
#n_obs = complex_dict[size][obs] #(on each side)
#experiment.target_freq_["spread"] = 12
#[height - 700, height - 450]
bounds_dict = simple_block(2000, target_spread = 12, n_obs = 20)
experiment.get_observers(method = "block",
missing = ctr,
split = 0.5,
#dataset = experiment.A,
observer_range = bounds_dict["observer_bounds"], #format: [[425, 525], [527,627]],
response_range = bounds_dict["response_bounds"], #format: [[525, 527]],
aspect = 1)
Train, Test, xTr, xTe = dat["obs_tr"], dat["obs_te"], dat["resp_tr"], dat["resp_te"]
# -
# # GOAL: The first step is to save the best_arguments results in some sort of easily accessed file. Also save the data: ie the predictions. We need a better file system with automatically changing variable names and nice file structure.
#
# Additionally lets make the selection of different sizes of spectogram seamless.
#
#
# # User Defined values:
file_ = "spectogram_data/"
size = "medium" #"medium"
freq = "2k"
spread = "small_spread"
obs = "small_obs"
file_ = file_ + size + "/"
file_
# #...
# +
#size_options = { "small" : 512, "medium" : 1024, "publish": 3000}
spect_files = { "publish" : "_new", "small" : "_512" , "original" : "", "medium" : "_1024"}
spect_xrange = { "publish" : range(0,3400, 500), "small" : range(0, 512, 50)}
# -
# # TODO: Hard code values of frequencies
# +
def spect_(request, size = size):
"""
This function returns a hard-coded spectogram size specific value.
"""
hi = complex_dict[size]
hi = hi[request]
return(hi)
#spect_('2k')
# -
# # i. Load Data
# +
files2import = [file_ + i + spect_files[size] for i in ("T", "f", "Intensity") ]
print(files2import)
data_lst = []
for i in files2import:
data_lst.append(loadmat(i))
print("successfully loaded " +str(files2import) )
T, f, A = data_lst
#preprocessing
T, A = T['T'], A['M']
T, A = np.transpose(T), (A - np.mean(A))/np.std(A)
A_orig = A.copy()
print("data loaded")
A_orig = np.rot90(A_orig, k = 1, axes = (0, 1))
plt.imshow(A_orig)
f = f['f'].reshape(-1,).tolist()
global max_freq; max_freq = int(np.max(f))
print("maximum frequency: " + str(max_freq))
freq_axis_len = A.shape[0]
time_axis_len = A.shape[1]
print(A.shape)
# -
# # Target Freq code, repeated above observers for sense, olab_display
def olab_display(axis, f = f, return_index = False):
oA = np.rot90(A_orig.copy().copy(), k = 3, axes = (0, 1))
#oA stands for other lab A
oA = pd.DataFrame(oA).copy()
f = [int(i) for i in f]
freq_idx = f#[ int(i / 100) * 100 for i in f.tolist()]
oA.index = freq_idx
yticks = list( range( 0, max_freq, 1000))
y_ticks = [ int(i) for i in yticks]
my_heat = sns.heatmap(oA, center=0, cmap=sns.color_palette("CMRmap"),
yticklabels = A.shape[0]//10, ax = axis)
#, cmap = sns.color_palette("RdBu_r", 7))
axis.set_ylabel('Frequency (Hz)')#,rotation=0)
axis.set_xlabel('time')
my_heat.invert_yaxis()
plt.yticks(rotation=0)
return(freq_idx)
# # FIND KEY FREQUENCIES
# ## KEY FUNCTION: IDX2FREQ
# +
key_freq_idxs = {}
for i in (2000, 4000, 8000):
height = freq_axis_len
key_freq_idxs[i] = height - idx2Freq(i)
display(key_freq_idxs)
target_freq_ = { #"center" : complex_dict[size][freq] ,
"spread" : 50 } #complex_dict[size][spread]}
# -
# # Display Data vertically with correct index
fig, ax = plt.subplots(1, 1, figsize = (6,5))
#freq_idx = olab_display(ax)
# # Horizontal display with proper index
# +
A_pd = pd.DataFrame(A_orig)
A_pd.columns = freq_idx
my_heat= sns.heatmap(A_pd, center=0, cmap=sns.color_palette("CMRmap"))
plt.xlabel('Frequency (Hz)')
plt.ylabel('time')
A = A_pd.values
# -
# # Helper Functions
#
# Consider eliminating validation set superfluous lines
# ### plot_timeseries and sub_helper function
# +
def build_pd(np_, n_series):
series_len = np_.shape[0]
for i in range(n_series):
id_np = np.zeros((series_len, 1)).reshape(-1, 1) + i
series_spec = np_[:, i].reshape(-1, 1)
t = np.array( list( range( series_len))).reshape(-1, 1)
pd_spec = np.concatenate( [ t, series_spec, id_np], axis = 1)
pd_spec = pd.DataFrame(pd_spec)
pd_spec.columns = ["t", "x", "id"]
if i == 0:
df = pd_spec
else:
df = pd.concat([df, pd_spec], axis = 0)
return(df)
def plot_timeseries(prediction_,
train,
test,
titl = "ESN ",
series2plot = 0,
method = None,
label_loc = (0., 0.)):
"""
This function makes three plots:
the prediction, the residual, the loss.
It was built for single predictions, but needs to be upgraded to deal with multiple output.
We need to show: average residual, average loss.
"""
full_dat = np.concatenate([train, test], axis = 0); full_dat_avg = np.mean(full_dat, axis = 1)
n_series, series_len = test.shape[1], test.shape[0]
assert method in ["all", "single", "avg"], "Please choose a method: avg, all, or single"
#assert method != "all", "Not yet implimented #TODO"
if method == "single":
label_loc = (0.02, 0.65)
#key indexes
trainlen, testlen, pred_shape = train.shape[0], test.shape[0], prediction_.shape[0]
if method == "single":
if n_series > 1:
print("There are " + str(n_series) + " time series, you selected time series "
+ str(series2plot + 1))
# avoid choosing all of the columns. subset by the selected time series.
train, test, prediction = train[:, series2plot], test[:, series2plot], prediction_[:, series2plot]
# set up dataframe
xTrTarg_pd = pd.DataFrame(test)
t = pd.DataFrame(list(range(len(xTrTarg_pd))))
# append time
Target_pd = pd.concat([xTrTarg_pd, t], axis = 1)
Target_pd.columns = ["x", "t"]
#calculate the residual
resid = test.reshape(-1,)[:pred_shape] - prediction.reshape(-1,) #pred_shape[0]
rmse_spec = str(round(myMSE(prediction, test), 5))
full_dat = np.concatenate([train, test], axis = 0)
elif method == "avg":
rmse_spec = str(round(myMSE(prediction_, test), 5))
prediction = prediction_.copy().copy()
def collapse(array):
return(np.mean(array, axis = 1))
vals = []
#y - yhat
resid_np = test - prediction_
for i in [train, test, prediction_, resid_np]:
vals.append(collapse(i))
train, test, prediction_avg, resid = vals
#return(prediction)
else: ##############################################################################################
#TODO make a loop and finish this, hopefully pretty colors.
rmse_spec = str(round(myMSE(prediction_, test), 5))
pd_names = ["Lines", "prediction", "resid"]
pd_datasets = [ full_dat, prediction_, test - prediction_]
rez = {}
for i in range(3):
# TODO functionalize this to streamline the other plots.
name_spec = pd_names[i]
dataset_spec = pd_datasets[i]
rez[name_spec] = build_pd(dataset_spec, n_series)
Lines_pd, resid_pd, prediction_pd = rez["Lines"], np.abs(rez["resid"]), rez["prediction"]
#display(Lines_pd) #np.zeros((4,1))
####### labels
if method in ["single"]:
plot_titles = [ titl + "__: Prediction vs Ground Truth, rmse_: " + rmse_spec,
titl + "__: Prediction Residual",
titl + "__: Prediction Loss"]
plot_labels = [
["Ground Truth","prediction"]
]
elif method == "avg":
plot_titles = [titl + "__: Avg Prediction vs Avg Ground Truth, total rmse_: " + rmse_spec,
titl + "__: Avg Prediction Residual",
titl + "__: Avg Prediction Loss"]
plot_labels = [
[ "", "Avg Ground Truth", "avg. prediction"]
]
elif method == "all":
plot_titles = [titl + "__: Visualization of Time series to Predict, rmse_: " + rmse_spec,
titl + "__: Prediction Residuals", titl + "__: Prediction Loss"
]
### [plotting]
#display(Target_pd)
fig, ax = plt.subplots(3, 1, figsize=(16,10))
i = 0 # plot marker
j = 0 # subplot line marker
######################################################################## i. (avg.) prediction plot
if method in ["single", "avg"]:
if method == "single": col, alph = "cyan", 0.5,
else: col, alph = "grey", 0.3
### ground truth
ax[i].plot(range(full_dat.shape[0]), full_dat,'k', label=plot_labels[i][j],
color = col, linewidth = 1, alpha = alph); j+=1
if method == "avg":
ax[i].plot(range(full_dat.shape[0]), full_dat_avg,'k', label=plot_labels[i][j],
color = "cyan", linewidth = 1, alpha = 0.8); j+=1
# ground truth style
ax[i].plot(range(full_dat.shape[0]), full_dat_avg,'k', color = "blue", linewidth = 0.5, alpha = 0.4)
else:
# ground truth style
ax[i].plot(range(full_dat.shape[0]), full_dat,'k', color = "blue", linewidth = 0.5, alpha = 0.4)
### prediction
#pred style, pred
if method == "single":
ax[i].plot(range(trainlen,trainlen+testlen), prediction,'k',
color = "white", linewidth = 1.75, alpha = .4)
ax[i].plot(range(trainlen,trainlen+testlen), prediction,'k',
color = "red", linewidth = 1.75, alpha = .3)
ax[i].plot(range(trainlen,trainlen+testlen),prediction,'k',
label=plot_labels[i][j], color = "magenta", linewidth = 0.5, alpha = 1); j+=1
else: #potentially apply this to the all plot as well. Maybe only have two methods.
ax[i].plot(range(trainlen,trainlen+testlen), prediction,'k',
color = "pink", linewidth = 1.75, alpha = .35)
ax[i].plot(range(trainlen,trainlen+testlen), prediction_avg,'k',
color = "red", linewidth = 1.75, alpha = .4, label = "prediction avg")
#first plot labels
ax[i].set_title(plot_titles[i])
ax[i].legend(loc=label_loc)
i+=1; j = 0
else:
sns.lineplot( x = "t", y = "x", hue = "id", ax = ax[i],
data = Lines_pd, alpha = 0.5,
palette = sns.color_palette("hls", n_series))
ax[i].set_title(plot_titles[i])
i+=1
if method in ["single", "avg"]:
######################################################################## ii. Residual plot
ax[i].plot(range(0,trainlen),np.zeros(trainlen),'k',
label="", color = "black", alpha = 0.5)
ax[i].plot(range(trainlen, trainlen + testlen), resid.reshape(-1,),'k',
color = "orange", alpha = 0.5)
# second plot labels
#ax[1].legend(loc=(0.61, 1.1))
ax[i].set_title(plot_titles[i])
i+=1
else:
resid_pd_mn = resid_pd.pivot(index = "t",
columns = "id",
values = "x"); resid_pd_mn = resid_pd_mn.mean(axis = 1)
sns.lineplot( x = "t", y = "x", hue = "id", ax = ax[i], data = resid_pd, alpha = 0.35, label = None)
for j in range(n_series):
ax[i].lines[j].set_linestyle((0, (3, 1, 1, 1, 1, 1)))#"dashdot")
sns.lineplot(ax = ax[i], data = resid_pd_mn, alpha = 0.9, color = "r",
label = "mean residual")
ax[i].set_title(plot_titles[i])
i+=1
####################################################################### iii. Loss plot
if method in ["single", "avg"]:
ax[i].plot(range(0,trainlen),np.zeros(trainlen),'k',
label="", color = "black", alpha = 0.5)
ax[i].plot(range(trainlen,trainlen+testlen),resid.reshape(-1,)**2,'k',
color = "r", alpha = 0.5)
# second plot labels
#ax[2].legend(loc=(0.61, 1.1))
ax[i].set_title(plot_titles[i])
elif method == "all":
# create the loss dataframe
loss_pd = resid_pd.copy();
vals = loss_pd['x'].copy().copy(); loss_pd['x'] = vals **2
loss_pd_mn = loss_pd.pivot(index = "t",
columns = "id",
values = "x"); loss_pd_mn = loss_pd_mn.mean(axis = 1)
sns.lineplot( x = "t", y = "x", hue = "id", ax = ax[i], data = loss_pd, alpha = 0.35, label = None)
for j in range(n_series):
ax[i].lines[j].set_linestyle((0, (3, 1, 1, 1, 1, 1)))#"dashdot")
sns.lineplot(ax = ax[i], data =loss_pd_mn, alpha = 0.9, color = "magenta",
label = "mean loss")
ax[i].set_title(plot_titles[i])
i+=1
plt.subplots_adjust(hspace=0.5)
plt.show()
# -
# ### get_observers and helper functions
# +
def diff(first, second):
second = set(second)
return [item for item in first if item not in second]
def my_range2lst(response_range):
"""
This function takes on two forms: lst and lst_of_lsts
in the lst form, it simply takes a list [a,b] where a<b ie a numerical range, and converts that into a list
of all of the values contained by the range.
The reason we have a function at all is because of the lst_of_lsts option, where it returns multiple ranges.
"""
if type(response_range[0]) != list:
response_range_lst = [response_range]
else:
response_range_lst = response_range
lst_idx = []
for i, range_ in enumerate(response_range_lst):
range_start = range_[0]
range_stop = range_[1]
lst_idx += np.sort( np.array( list( range( range_start, range_stop)))).tolist()
lst_idx = np.sort(np.array(lst_idx)).tolist()
return(lst_idx)
def myMSE(prediction,target):
return np.sqrt(np.mean((prediction.flatten() - target.flatten() )**2))
def Shape(lst):
npObj, label = lst; print(label + " shape: " + str(npObj.shape))
# validation version
def get_observers(aspect = 6,
dataset = A,
method = "random",
missing = key_freq_idxs[2000],
num_observers = 20,
observer_range = None,
plot_split = True,
response_range = None,
split = 0.2,
max_freq = max_freq,
freq_idx = f
):
"""
arguments:
aspect: affect the size of the returned plot.
dataset: obvious
method:
(+) random
(+) equal #similar to barcode, equal spacing, with k missing block. Low priority.
(+) block
(+) barcode #TODO block but with gaps between observers.
# I think this will show that you don't really need every line of the data to get similar accuracy
missing: either
(+) any integer: (standing for column of the spectogram) or
(+) "all" : which stands for all of the remaining target series.
num_observers: the number of observers that you want if you choose the "random" method.
observer_range: if you select the "block" opion
"""
n_rows = dataset.shape[0]
n_cols = dataset.shape[1]
train_len = int(n_rows * split)
val_split = dataset.shape[0]
test_len = n_rows - train_len
col_idx = list(range(n_cols))
#remove the response column which we are trying to use for inpainting
if method == "random":
col_idx.remove(missing)
obs_idx = np.random.choice(col_idx, num_observers, replace = False)
response = dataset[ : , missing].reshape(-1,1)
response_idx = [missing]
elif method == "eq":
print("equal spacing")
print("NOT YET IMPLIMENTED")
elif method == "all":
obs_idx = np.random.choice( col_idx, num_observers, replace = False)
response_idx = diff( col_idx, obs_idx.tolist())
response = dataset[ : , response_idx]
### The following is problematic because you haven't dealt with the case where they overlap.
### BLOCK
elif method == "block":
"""
This method either blocks observers and/or the response area.
"""
print("you selected the block method")
if response_range == None:
response_idx = [missing]
response = dataset[ : , missing].reshape( -1, 1)
else:
response_idx = my_range2lst(response_range)
response = dataset[ : , response_idx].reshape( -1, len( response_idx))
for resp_idx_spec in response_idx:
col_idx.remove( resp_idx_spec)
if observer_range == None:
col_idx.remove( missing)
obs_idx = np.sort( np.random.choice( col_idx,
num_observers,
replace = False))
else:
obs_idx = my_range2lst(observer_range)
# check for problems with the block method:
union_obs_resp_set = set(obs_idx) & set(response_idx)
err_msg = "Error: overlap in obs_idx and response_idx \n"
err_msg += "overlap: " + str(list(union_obs_resp_set))
assert list(union_obs_resp_set) == [], err_msg
observers = dataset[ :val_split, obs_idx]
observers_tr = observers[ :train_len, : ]
observers_te = observers[ train_len : val_split, : ]
response_tr = response[ :train_len, : ]
response_te = response[ train_len:val_split, : ]
### Visualize the train test split and the observers
if plot_split == True:
red, yellow, blue, black = [255, 0, 0], [255, 255, 0], [0, 255, 255], [0, 0, 0]
orange, green, white = [255, 165, 0], [ 0, 128, 0], [255, 255, 255]
#preprocess:
split_img = np.full(( n_rows, n_cols, 3), black)
# assign observer lines
for i in obs_idx:
split_img[ : , i] = np.full(( 1, n_rows, 3), yellow)
# assign target area
for i in response_idx:
split_img[ :train_len, i] = np.full(( 1, train_len, 3), blue)
split_img[ train_len:, i] = np.full(( 1, test_len, 3), red)
legend_elements = [Patch(facecolor='cyan', edgecolor='blue', label='Train'),
Patch(facecolor='red', edgecolor='red', label='Test'),
Patch(facecolor='yellow', edgecolor='orange', label='Observers')]
# Create the figure
fig, ax = plt.subplots( 1, 2, figsize = ( 12, 6))
ax = ax.flatten()
solid_color_np = np.transpose(split_img.T, axes = (1,2,0))
#solid_color_pd.index = freq_idx
# The legend:
#https://matplotlib.org/3.1.1/gallery/text_labels_and_annotations/custom_legends.html
##################################### START plots
# simple olab_display( sns heatmap on the right)
#++++++++++++++++++++++++++++++++++++ plot 1: sns heatmap
olab_display(ax[1])
ax[1].set_title("Spectogram Data")
# retrieve labels to share with plot 0
# We need to retrieve the labels now.
plt.sca(ax[1])
locs, labels = plt.yticks()
freq_labels = np.array([int(label.get_text()) for label in labels])
#++++++++++++++++++++++++++++++++++++ plot 0: diagram showing training, test splits and observers.
#now plot the diagram on the left:
ax[0].set_title("Dataset Split Visualization")
ax[0].imshow(solid_color_np, aspect = aspect)
### fixing labels on plot 0, involved!
# label axes, legend
ax[0].set_ylabel('Frequency (Hz)'); ax[0].set_xlabel('time')
ax[0].legend(handles=legend_elements, loc='lowerright')
#now calculate the new positions
max_idx = solid_color_np.shape[0]
#new positions
new_p = (freq_labels/max_freq) * max_idx
adjustment = max_idx - np.max(new_p); new_p += adjustment -1; new_p = np.flip(new_p)
plt.sca(ax[0]); plt.yticks(ticks = new_p, labels = freq_labels)
plt.show()
##################################### END plots
# print dimensions ect.
print_lst = [(observers_tr, "X target"), (observers_te, "X test")]
print_lst += [(response_tr, "response train"), (response_te, "response test")]
for i in print_lst:
Shape(i)
print("observer_range: " + str(observer_range))
if response_idx == None:
print("target index: " + str(missing))
else:
print("response range: " + str(response_range))
dat2be = {"obs_tr" : observers_tr,
"obs_te" : observers_te,
"resp_tr" : response_tr,
"resp_te" : response_te,
"obs_idx" : obs_idx,
"resp_idx" : response_idx}
return(dat2be)
def currTime():
now = datetime.now()
current_time = now.strftime("%H:%M:%S")
print("Current Time =", current_time)
currTime()
# -
# # Block Prediction
# +
#TODO: split this dict up or eliminate it. Let's just have an observer dict.
complex_dict = {
"small" : {
#target_frequencies
"2k" : 101,
"4k" : 206,
"8k" : 307,
#target spread sizes
"no_spread" : None,
"small_spread" : 4,
"medium_spread" : 12,
"large_spread" : 24,
#observer values
"small_obs" : 10,
"medium_obs" : 25,
"large_obs" : 50
},
"medium" : {
#target_frequencies
"2k" : 101,
"4k" : 206,
"8k" : 307,
#target spread sizes
"no_spread" : None,
"small_spread" : 4,
"medium_spread" : 12,
"large_spread" : 24,
#observer values
"small_obs" : 10,
"medium_obs" : 25,
"large_obs" : 50
},
"publish": {
"2k" : 546,
"4k" : 1089,
"8k" : 2177,
"0.5_sec" : 1371,
"0.7_sec" : 1924
}
}
# size: medium, 1k by 1k
obs = "large_obs"
n_obs = complex_dict[size][obs] #(on each side)
ctr = key_freq_idxs[2000]
target_freq_["spread"] = 12
if target_freq_["spread"] != None:
target_frequency = [ctr - target_freq_["spread"],
ctr + target_freq_["spread"]]#[height - 700, height - 450]
tf = target_frequency
resp_range_ = [[tf[0], tf[1]]]
else:
resp_range_ = None
tf = [ctr, ctr]
obs_range_ = [[tf[0] - n_obs, tf[0]], [tf[1], tf[1] + n_obs ]]
dat = get_observers(method = "block",
missing = ctr,
split = 0.5,
dataset = A,
observer_range = obs_range_, #format: [[425, 525], [527,627]],
response_range = resp_range_, #format: [[525, 527]],
aspect = 1)
Train, Test, xTr, xTe = dat["obs_tr"], dat["obs_te"], dat["resp_tr"], dat["resp_te"]
print("total observers: " + str(len(dat["obs_idx"])))
print("total targets: " + str(len(dat["resp_idx"])))
# try to make an interactive result, where you can pull up any timeseries. R?
# +
print("sanity check")
print("Does the obs_tr match the what we expect?")
orig_series, processed_series = A[0:dat["obs_tr"].shape[0], dat["obs_idx"][0]], dat["obs_tr"][:,0]
plt.plot(orig_series, color = "red")
plt.plot(processed_series, linestyle = "--", color = "cyan")
assert(np.array_equal(orig_series, processed_series))
# -
# # Data Saving Functions
# +
def count_files(path, current):
count = 0
for path in pathlib.Path(path).iterdir():
if path.is_file():
count += 1
if current:
count = count - 1
return("_" + str(count))
def get_new_filename(exp,
obs = len(dat["obs_idx"]),
target_freq = "2k",
ctr = key_freq_idxs[2000],
spread = target_freq_["spread"],
current = False
):
"""
ideally this function will serve two purposes: it will return a new filename and return
a dict of data so that we can recreate the experiment.
This should include
1) The obs and resp indices, the "best_arguments" (the optimized hyper-parameters),
and the prediction.
"""
if exp:
prefix = 'exp_w'
else:
prefix = 'non_exp_w'
obs, ctr, spread = str(obs), str(ctr), str(spread)
new_dir = "results/" + size + "/" + target_freq + "/"
count = count_files(new_dir, current = current)
new_filename = prefix + count + ".txt"
return(new_dir + new_filename )
def getData2Save(): #best_arguments, prediction = obs_prediction
"""
Save the data
current issue: how do we initialize this function properly?
"""
err_msg = "YOU NEED TO CALL THIS FUNCTION LATER "
json2be = {}
# 1) saving the structure of the data and split
json2be["basic_info"] = {"size" : size,
"freq" : freq,
"target_freq_" : target_freq_,
"n_obs" : len(dat["obs_idx"]),
"n_target" : len(dat["resp_idx"]),
"split_cutoff" : dat["resp_tr"].shape[0]}
#jsonify dat
new_dat = dat.copy().copy()
for key, item in new_dat.items():
if type(item) != list:
new_dat[key] = item.tolist()
json2be["dat"] = new_dat
# 2) saving the optimized hyper-parameters
try:
best_arguments
except NameError:
err_msg + "RC not yet trained"
else:
json2be["best_arguments"] = best_arguments
# 3) saving the prediction, mse
try:
obs_prediction
except NameError:
err_msg + "obs_prediction not yet created"
else:
json2be["prediction"] = obs_prediction.tolist()
mse = my_MSE(obs_prediction, dat["resp_te"], verbose = False)
json2be["results"] = {
"MSE" : mse,
"RMSE" : np.sqrt(mse)
}
return(json2be)
def save_json(exp):
save_spec_ = getData2Save()
new_file = get_new_filename(exp = exp)
with open(new_file, "w") as outfile:
data = json.dump(save_spec_, outfile)
def my_MSE(prediction, truth, verbose = True, label = ""):
mse_matrix = (prediction - truth)**2
mse = np.sum(mse_matrix)/(mse_matrix.shape[0]*mse_matrix.shape[1])
if verbose == True:
print(label + " MSE: " + str(mse))
return(mse)
# -
# # non exponential weights:
# +
hi, bye = {"eps" : 1e-5}, {"danger" : 12345}
hi = {**hi, **bye}
hi
# +
# %%time
currTime()
run_now = False
if run_now == True:
bounds = {
#'llambda' : (-12, 1),
'connectivity': 0.5888436553555889, #(-3, 0)
'n_nodes': (100, 1500),
'spectral_radius': (0.05, 0.99),
'regularization': (-12, 1),
}
esn_cv = EchoStateNetworkCV(bounds = bounds,
initial_samples=100,
subsequence_length=250, #150 for 500
eps=1e-5,
cv_samples=8,
max_iterations=1000,
scoring_method='tanh',
exp_weights = False,
obs_index = dat['obs_idx'],
target_index = dat["resp_idx"])
# n_jobs is the number of computer cores.
esn_cv.n_jobs = 5
#Optimize the hyper-parameters
best_arguments = esn_cv.optimize( x = Train, y = xTr)
# Save the Data
save_json(exp = False)
""" 7/26, 2k, medium, spread 12, n_obs = 50, llambda is : 8.249737488807662e-05
best_arguments = {
'connectivity': 0.5888436553555889,
'n_nodes': 100,
'spectral_radius': 0.05,
'regularization': 0.00019661357203730104}
"""
# -
def load_best_args(exp, file = "default"):
if file == "default":
nf = get_new_filename(exp = exp, current = True)
else:
nf = file
with open(nf) as json_file: # 'non_exp_w.txt'
non_exp_best_args = json.load(json_file)
best_args = non_exp_best_args["best_arguments"]
return(best_args)
def load_dat(exp, file = "default"):
if file == "default":
nf = get_new_filename(exp = exp, current = True)
else:
nf = file
with open(nf) as json_file: # 'non_exp_w.txt'
non_exp_best_args = json.load(json_file)
datt = non_exp_best_args["dat"]
datt["obs_tr"], datt["obs_te"] = np.array(datt["obs_tr"]), np.array(datt["obs_te"])
datt["resp_tr"], datt["resp_te"] = np.array(datt["resp_tr"]), np.array(datt["resp_te"])
return(datt)
file_= "/Users/hayden/Desktop/DL_LAB/Reservoir_fork/Reservoir/rc_phase2/results/medium/2k/final/7_26/0.9 split/non_exp_w_0.txt"
dat = load_dat(False, file = file_)
non_exp_best_args = load_best_args(False, file = file_)
Train, Test, xTr, xTe = dat["obs_tr"], dat["obs_te"], dat["resp_tr"], dat["resp_te"]
print("total observers: " + str(len(dat["obs_idx"])))
print("total targets: " + str(len(dat["resp_idx"])))
print(A.shape)
print(xTe.shape[0] + xTr.shape[0])
range(A.shape[0] - xTe.shape[0], A.shape[0])
# +
def runInterpolation(columnwise = False):
#2D interpolation
#observer coordinates
"""
for i, column_idx in enumerate(dat["resp_idx"]):
print(column_idx)
values += list(A[:,column_idx].reshape(-1,))
point_lst += list(zip(range(A.shape[0]), [column_idx]*A.shape[0]))
print(len(point_lst))
print(len(values))
"""
#Training points
missing_ = 60
points_to_predict = []
values = []
#visible
point_lst = []
total_zone_idx = dat["resp_idx"] + dat["obs_idx"]
#Train zone
for x in range(xTr.shape[0]):
# resonse points : train
for y in total_zone_idx:
point_lst += [(x,y)]#list(zip(range(Train.shape[0]) , [missing_]*Train.shape[0]))
values += [A[x,y]]
#Test zone
for x in range(A.shape[0] - xTe.shape[0], A.shape[0]):
# resonse points : train
for y in dat["resp_idx"]:
points_to_predict += [(x,y)]#list(zip(range(Train.shape[0]) , [missing_]*Train.shape[0]))
#values += [A[x,y]]
#observer points
for y in dat["obs_idx"]:
point_lst += [(x,y)]
values += [A[x,y]]
#just iterate through dat_idx
#print("point list length: " + str(len(point_lst)))
#print(xTe.shape)
#print(xTe.shape[0] * xTe.shape[1])
#observer points
#values += list(A[:Train.shape[0], column_idx].reshape(-1,))
#nnpoints_to_predict = list(zip(list(range(Train.shape[0], A.shape[0])), [missing_]*xTe.shape[0]))
ip2_pred = griddata(point_lst, values, points_to_predict, method='cubic')
ip2_pred = ip2_pred.reshape(xTe.shape)
ip2_resid = ip2_pred - xTe
#points we can see in the training set
###plots:
ip_res = {"prediction": ip2_pred, "resid" : ip2_resid,
"nrmse" : nrmse(pred_ = ip2_pred, truth = xTe, columnwise = columnwise) }
return(ip_res)
#sns.distplot(esn_obs.weights)
# +
# #%run -i '/Users/hayden/Desktop/DL_LAB/Reservoir/build/lib/reservoir/esn.py'
#new_file = nf
#non_exp_best_args = best_arguments#load_best_args(exp = False)#, file ="results/small/2k/exp_w_7.txt")#, file = "results/_orig/non_exp_w_small_2k_100ctr_sig12.txt")
#dat = load_dat(exp = False, file = "results/small/2k/exp_w_2.txt")
#Train, Test, xTr, xTe = dat["obs_tr"], dat["obs_te"], dat["resp_tr"], dat["resp_te"]
print("loaded!")
#non_exp_best_args['connectivity'] = [10**(-0.23)]
esn_obs = EchoStateNetwork(**non_exp_best_args, exponential = False,
resp_idx = dat["resp_idx"], obs_idx = dat['obs_idx'])
esn_obs.llambda = 10 ** (-2)
esn_obs.train(x = Train, y = xTr)
def my_predict(esn_spec, test, n_steps = None):
if not n_steps:
n_steps = test.shape[0]
return esn_spec.predict(n_steps, x = Test[:n_steps,:])
uniform_obs_prediction = my_predict(esn_obs, Test)
col_to_plot = 3
hi = plot_timeseries(uniform_obs_prediction,
train = xTr,
test = xTe,
titl = "Bayesian optimized normal RC",
method = "all",
label_loc = (0.01, 0.7))
sns.distplot(esn_obs.in_weights)
#save_json(exp = False)
display(non_exp_best_args)
# -
# # exponential Weights Optimization
# %%time
currTime()
"""'connectivity': 0.5888436553555889,
'n_nodes': 100,
'spectral_radius': 0.05,
'regularization': 0.00019661357203730104"""
run_now = False
if run_now == True:
bounds = {'llambda': (-12, 1), 'n_nodes': 1466,
'spectral_radius': 0.9436655497550964,
'regularization': np.log(0.005634420389409391)/np.log(10),
'connectivity': np.log(0.5888436553555889)/np.log(10),
}
for name, domain in bounds.items():
if type(domain) == np.float64:
print("hi")
bounds[name] = float(bounds[name])
print(type(domain))
esn_cv = EchoStateNetworkCV(bounds = bounds,
initial_samples=100,
subsequence_length= 250,
eps=1e-5,
cv_samples=5,
max_iterations=1000,
scoring_method='tanh',
exp_weights = True,
obs_index = dat['obs_idx'],
target_index = dat["resp_idx"])
### n_jobs is the number of computer cores.
esn_cv.n_jobs = 4
esn_cv.n_jobs
best_arguments = esn_cv.optimize( x = Train, y = xTr)
with open('exp_w.txt', "w") as outfile:
data = json.dump(best_arguments, outfile)
"""
best_arguments = {
'llambda': 8.249737488807662e-05,
'connectivity': 0.5888436553555889,
'n_nodes': 100,
'spectral_radius': 0.05,
'regularization': 0.00019661357203730104}
"""
# +
#exp_best_args
# -
file_= "/Users/hayden/Desktop/DL_LAB/Reservoir_fork/Reservoir/rc_phase2/results/medium/2k/final/7_26/0.9 split/exp_w_1.txt"
dat = load_dat(False, file = file_)
exp_best_args = load_best_args(False, file = file_)
Train, Test, xTr, xTe = dat["obs_tr"], dat["obs_te"], dat["resp_tr"], dat["resp_te"]
print("total observers: " + str(len(dat["obs_idx"])))
print("total targets: " + str(len(dat["resp_idx"])))
# +
# #%run -i '/Users/hayden/Desktop/DL_LAB/Reservoir/build/lib/reservoir/esn.py'
#exp_best_args = best_arguments
#exp_best_args = load_best_args(exp = True)
display(exp_best_args)
display(non_exp_best_args)
esn_obs = EchoStateNetwork(**exp_best_args, exponential = True,
resp_idx = dat["resp_idx"], obs_idx = dat['obs_idx'], plot = True)
#esn_obs.llambda = 0.01
esn_obs.train(x = Train, y = xTr)
fig,ax = plt.subplots(1,1)
sns.distplot(esn_obs.in_weights, ax = ax)
ax.set_title("Exponential Attention weights Kernel Density")
def my_predict(test, n_steps = None):
if not n_steps:
n_steps = test.shape[0]
return esn_obs.predict(n_steps, x = Test[:n_steps,:])
exp_obs_prediction = my_predict(Test)
plot_timeseries(exp_obs_prediction, method ="avg",
train = xTr, test = xTe, titl = "Bayesian optimized RC", series2plot = 0)
# -
# # Comparison / Improvement:
# +
def compare(truth,
unif_w_pred = None,
exp_w_pred = None,
columnwise = False,
verbose = False):
"""
This function provides two things, conditional on the columnwise variable.
columnwise = False: cross-model comparison of nrmse
columnwise = True: model nrmse correlary for each point.
"""
ip_res = runInterpolation(columnwise = columnwise)
if type(unif_w_pred) != type(None):
unif_nrmse = nrmse(pred_ = unif_w_pred, truth = truth, columnwise = columnwise)
if type(exp_w_pred) != type(None):
exp_nrmse = nrmse(pred_ = exp_w_pred , truth = truth, columnwise = columnwise)
assert type(columnwise) == bool, "columnwise must be a boolean"
if columnwise == False:
if verbose != False:
print("cubic spline interpolation nrmse: " + str(ip_res["nrmse"]))
print("uniform weights rc nrmse: " + str(unif_nrmse))
print("exponential weights rc nrmse: " + str(exp_nrmse))
print("creating barplot")
df = pd.DataFrame({"interpolation" : ip_res["nrmse"],
"uniform rc" : unif_nrmse,
"exponential rc" : exp_nrmse}, index = [0])
display(df)
plt.figure(num=None, figsize=(8, 6), dpi=80, facecolor='w', edgecolor='k')
sns.catplot(data = df, kind = "bar")
plt.title("model vs nrmse")
plt.ylabel("nrmse")
improvement = []
for rc_nrmse in[unif_nrmse, exp_nrmse]:
impr_spec = ((ip_res["nrmse"] - rc_nrmse)/ip_res["nrmse"]) * 100
impr_spec = [round(impr_spec,1)]
improvement += impr_spec
pct_improve_unif, pct_improve_exp = improvement
if pct_improve_unif > 0:
print("unif improvement vs interpolation: nrmse " + str(-pct_improve_unif) + "%")
else:
print("rc didn't beat interpolation: nrmse +" + str(-pct_improve_unif) + "%")
if pct_improve_exp > 0:
print("exp improvement vs interpolation: nrmse " + str(-pct_improve_exp) + "%")
else:
print("rc didn't beat interpolation: nrmse +" + str(-pct_improve_exp) + "%")
impr_rc_compare = round(((unif_nrmse - exp_nrmse)/unif_nrmse) * 100,1)
if impr_rc_compare > 0:
print("exp rc improvement vs unif rc: nrmse " + str(-impr_rc_compare) + "%")
else:
print("exp weights didn't improve rc: nrmse +" + str(-impr_rc_compare) + "%")
else:
print("creating first figure")
model_names = ["interpolation", "uniform rc", "exponential rc"]
for i, model_rmse_np in enumerate([ip_res["nrmse"], unif_nrmse, exp_nrmse]):
model_rmse_pd = pd.melt(pd.DataFrame(model_rmse_np.T))
model_rmse_pd.columns = ["t","y"]
model_rmse_pd["model"] = model_names[i]
if i == 0:
models_pd = model_rmse_pd
else:
models_pd = pd.concat([models_pd, model_rmse_pd ], axis = 0)
fig, ax = plt.subplots(1,1, figsize = (11, 6))
sns.lineplot(x = "t", y = "y", hue = "model", data = models_pd, ax = ax)
ax.set_title("model vs rmse")
ax.set_ylabel("nrmse")
ax.set_xlabel("Test idx")
#return(df)
# -
compare(truth = xTe, unif_w_pred = uniform_obs_prediction, exp_w_pred = exp_obs_prediction)
print("2k, 24 target sequences, 100 observers, 0.9 split")
hi = compare(truth = xTe,
unif_w_pred = uniform_obs_prediction,
exp_w_pred = exp_obs_prediction,
columnwise = True)
#hi = pd.melt(hi)
# +
hi.columns = ["model", "nrmse"]
idxx = list(range(hi.shape[0]//3)) + list(range(hi.shape[0]//3)) + list(range(hi.shape[0]//3))
# -
sns.lineplot(x = idxx, y = "nrmse", hue = "model", data = hi)
nrmse(pred_ = exp_obs_prediction , truth = xTe, columnwise = True)
# # Simple predictions
# +
#TODO
# -
# # Now testing multiple outputs:
# +
# %%time
#dat = get_observers("all")
#plt.plot(xTr)
Train = dat["obs_tr"]
Test = dat["obs_te"]
xTr = dat["resp_tr"]
xTe = dat["resp_te"]
# +
# %%time
bounds = {
#'input_scaling': (0, 1),
#'feedback_scaling': (0, 1),
'leaking_rate' : (0, 1),
'spectral_radius': (0.01, 1.25),
'regularization': (-12, 1),
'connectivity': (-3, 0),
'n_nodes': (100, 1000)
}
esn_cv = EchoStateNetworkCV(bounds=bounds,
initial_samples=100,
subsequence_length=250,
eps=1e-4,
cv_samples=3,
max_iterations=1000,
scoring_method='tanh',
verbose=True)
### n_jobs is the number of computer cores.
esn_cv.n_jobs = 8
esn_cv.n_jobs
best_arguments = esn_cv.optimize(x = Train, y = xTr)
# +
esn_obs = EchoStateNetwork(**best_arguments)
esn_obs.train(x = Train, y = xTr)
BO_obs_prediction = my_predict(Test)
#plot_timeseries(prediction, train = xTrTarg, test = xTeTarg )
plot_timeseries(BO_obs_prediction, train = xTr, test = xTe, titl = "Bayes", col2plot = 0)
plot_timeseries(BO_obs_prediction, train = xTr, test = xTe, titl = "Bayes", col2plot = 40)
# +
kk = 20
fig, ax = plt.subplots(1,1,figsize = (16,4))
orig_idx = list(range(0,len(xTe[:,0])-1, kk))
resids_y_vals = []
resids_x_vals = []
count = 0
x_range= np.array(list(range(len(xTe[:,1])-1)))
print(x_range)
for i in range(0,100):
count+=1
resid_i = np.abs(xTe[:,i] - BO_obs_prediction[:,i])[orig_idx]
resids_y_vals += resid_i.tolist()
print(resid_i)
break
resids_x_vals += list(range(len(x_range[orig_idx].tolist())))
plt.plot( resid_i, alpha = 0.05, color = "blue")
my_dict = {"x":resids_x_vals, "y":resids_y_vals}
my_np = pd.DataFrame(my_dict).values
sorted_array = pd.DataFrame(my_np[np.argsort(my_np[:, 0])])
sorted_array.columns = ["x", "y"]
display(sorted_array)
sns.regplot( x = "x", y = "y",data = sorted_array, color = "red", x_jitter=.1) #alpha = 0.1)
ax.set_title = "Absolute Residual: all 180 Timeseries"
#clean this up by making the reg-plot work via a dataframe. But since this is extra I'm moving on.
# -
xTe.shape
fig, ax = plt.subplots(1,1,figsize = (16,4))
for i in range(len(dat["resp_idx"])):
resid_i = (xTe[:,i]-BO_obs_prediction[:,i])**2
plt.scatter(x = range(len(resid_i)), y = resid_i, alpha = 0.1, c= resid_i)
ax.set_title = "Absolute Residual: all 180 Timeseries"
# +
loss_np = np.zeros([xTe.shape[0], xTr.shape[1]])
total_np = np.zeros([pred_.shape[0], pred_.shape[1]])
fig, ax = plt.subplots(1,1,figsize = (16,4))
for count, i in enumerate(range(len(dat["resp_idx"]))):
resid_i = abs(xTe[:,i] - BO_obs_prediction[:,i])#**2
loss_np[:,i] = resid_i
plt.plot( resid_i, alpha = 0.04, color = "blue")
for count, i in enumerate(range(len(dat["resp_idx"]))):
resid_i = (xTe[:,i] - BO_obs_prediction[:,i])**2
#print(resid_i.shape)
ax.set_title = "Absolute Residual: all 180 Timeseries"
# +
#let's get the original data
def other_lab(arr):
return(np.rot90(arr, k=1, axes=(0, 1)))
fig, ax = plt.subplots(1,3, figsize=[15,10])
ax = ax.flatten()
#plt.imshow(loss_np)
sns.heatmap(other_lab(loss_np), ax = ax[0])
ax[0].set_title("residual error")
ax[1].set_title("prediction")
sns.heatmap(other_lab(BO_obs_prediction), ax = ax[1], cmap = "CMRmap")
ax[2].set_title("original")
sns.heatmap(other_lab(xTe), ax = ax[2], cmap = "CMRmap")
plt.subplots_adjust(hspace = 0.5)
# +
orig_idx = list(range(0, 420, 20))
#reverse interpolation function
def non_ip(arr):
return(arr[orig_idx,:])
fig, ax = plt.subplots(1,3, figsize=[15,10])
ax = ax.flatten()
ax[0].set_title("original")
sns.heatmap(other_lab(non_ip(xTe)), ax = ax[0], cmap = "CMRmap")
ax[0].set_xlabel("time")
ax[1].set_title("prediction")
sns.heatmap(other_lab(non_ip(BO_obs_prediction)), ax = ax[1], cmap = "CMRmap")
ax[1].set_xlabel("time")
sns.heatmap(other_lab(non_ip(loss_np)), ax = ax[2])
ax[2].set_title("residual error")
ax[2].set_xlabel("time")
# -
# # Deleted Scenes:
# exp_weights
# +
def exp_w(llambda = 1, distance = None, verbose = False):
"""
Args:
llambda: is llambda in an exponential function.
distance: is a distance matrix.
This function calculates weights via attention ie the
distance matrix which measures the distance from the
observer sequences to the target sequences.
"""
exp_np = np.exp( - llambda * distance) #*llambda
exp_np = exp_np.sum(axis = 0).reshape( -1 )
#normalize the max weight to 1.
exp_np = (exp_np) / np.max(exp_np)
if(verbose) == True:
display(pd.DataFrame(exp_np))
print("sum = " + str(np.sum(exp_np)))
return(exp_np)
def build_distance_matrix(resp, obs, verbose = False):
"""
args:
resp is the response index (a list of integers associated with the target train/test time series
(for example individual frequencies)
obs is the same for the observation time-series.
Description:
DistsToTarg stands for distance numpy array
"""
for i, resp_seq in enumerate(resp):
DistsToTarg = abs(resp_seq - np.array(obs)).reshape(1,-1)
if i == 0:
distance_np = DistsToTarg
else:
distance_np = np.concatenate([distance_np, DistsToTarg], axis = 0)
if verbose == True:
display(pd.DataFrame(distance_np))
return(distance_np)
def get_exp_weights(resp_idx = dat["resp_idx"],
obs_idx = dat["obs_idx"],
plot = True): #change the automatic var assignments
d_mat = build_distance_matrix(resp_idx, obs_idx)
exp_weights = exp_w(llambda = 0.05, distance = d_mat)
n_temp = len(exp_weights)
sign = np.random.choice([-1,1],n_temp)
#print("exp weights shape", exp_weights.shape)
exp_weights *= sign
pd_ = pd.DataFrame({"obs_idx": obs_idx, "weight": exp_weights})
#print("max_weight: " + str(np.max(exp_weights)))
if plot == True:
fig, ax = plt.subplots(1,1, figsize = (6, 4))
sns.scatterplot(x = "obs_idx", y = "weight", data = pd_, ax = ax)
ax.set_title("Exponential Attention Weights")
return(exp_weights)
get_exp_weights()
# -
# Visualization
fig, ax = plt.subplots(1,1, figsize = (16,4))
for i in range(A.shape[0]):
ax.plot( A[:,i], alpha = 0.01)
ax.set_xlabel("Time")
ax.set_ylabel("Normalized Intensity")
# Expand
# +
from scipy.interpolate import interp1d
from scipy.interpolate import griddata
def set_Diff(set_a, set_b):
if type(set_a) == range:
set_a = list(set_a)
if type(set_b) == range:
set_b = list(set_b)
if type(set_a) == list:
set_a = set(set_a)
if type(set_b) == list:
set_b = set(set_b)
c = set_a.intersection(set_b)
set_diff = set_a.union(set_b)-c
return(list(set_diff))
def get_all_coords(shape_):
"""
args: shape: the shape of the numpy array you want all the cartesian points for.
"""
pnt_lst = []
range0 = range(shape_[0])
range1 = range(shape_[1])
for x_coord in range0:
pnt_lst += list(zip([x_coord]*len(range1), range1))
return(pnt_lst)
def expand(arr, k = 6, axis = 0):
"""
args:
k: the number of points (padding) between points
arr: the numpy array that we're trying to expand via interpolation
axis: 0 is row-wise, 1 is column wise.
"""
other_axis = (axis +1) % 2
known_pnt_lst = []
known_values = []
unknown_pnt_lst = []
x_coords = range(0, arr.shape[axis]*k, k)
full_lst = list(range(arr.shape[axis]*k))
unknown_x_coords = set_Diff(x_coords, full_lst)
#all coords
for i in range(A.shape[1]):
known_pnt_lst += list(zip(x_coords, [i]*A.shape[1]))
known_values += list(A[:,i].reshape(-1,))
unknown_pnt_lst += list(zip(unknown_x_coords, [i] * A.shape[1]))
#pnts_to_ip is points to interpolate
#return({"act_pnts" : known_pnt_lst, "pnts_to_ip" : known_pnt_lst})
#verbatim:
#for i, column_idx in enumerate(dat["tr_idx"]):
# print(column_idx)
# values += list(A[:,column_idx].reshape(-1,))
# point_lst += list(zip(range(A.shape[0]), [column_idx]*A.shape[0]))
#print(len(point_lst))
#print(len(values))
#Next in this function:
point_lst = known_pnt_lst
points_to_predict = get_all_coords((A.shape[0]*k, A.shape[1]))#unknown_pnt_lst
ip2_pred = griddata(point_lst, known_values, points_to_predict, method='cubic')
ip2_pred = ip2_pred.reshape(-1, A.shape[1])
return({"prediction" : ip2_pred, "non_ip_idx": x_coords })
71*(706)
# -
def how_many_less_than(threshold= 0.1):
"""
This function tells home many are in np array are less than a a value.
"""
threshold = 0.1
pct_less = int(100 * np.sum(exp_weights < threshold)/len(exp_weights))
print("pct of exp_weights less than " + str(threshold)+ ": " + str(pct_less)+"%")
# +
### # ii. Hackily change directory and then run esn_cv.py
#import os
#reservoir_package_dir = '/Users/hayden/Desktop/DL_LAB/Reservoir/reservoir'
#os.chdir(reservoir_package_dir)
# #%run -i './esn_cv.py'
#/Users/hayden/Desktop/DL_LAB/Reservoir/reservoir
# -
# prediction = expand(A, k = 20)
# res = prediction
# pred_, non_ip_idx = res["prediction"], res["non_ip_idx"]
#
# fig, ax = plt.subplots(1,2, figsize = (16,6))
# ax = ax.flatten()
#
# sns.heatmap(np.rot90(pred_, k=1, axes=(0, 1)), center=0, cmap=sns.color_palette("CMRmap"), ax = ax[0])
# ax[0].set_ylabel('Frequency (Hz)')
# ax[0].set_xlabel('time')
# ax[0].set_title("expanded spectogram (via interpolation)")
# #
# sns.heatmap(np.rot90(A, k=1, axes=(0, 1)), center=0, cmap=sns.color_palette("CMRmap"), ax = ax[1])#, cmap = sns.color_palette("RdBu_r", 7))
# #, cmap = sns.color_palette("RdBu_r", 7))
# ax[1].set_ylabel('Frequency (Hz)')
# ax[1].set_xlabel('time')
# ax[1].set_title("Original spectogram")
# plt.show()
# # Boundaries discussed with Zhizhuo
"""
Block:
1371 = 0.5 sec, 1924 = 0.7 sec, 1.1 end
2k = 520-570 missing 50, 100 lines missing 500 - 600
# observers: 50% of missing
Three window sizes 0.5->0.7, 0.5 -> 0.9, 0.5 -> 1.1
"""
window_end1, window_end2, window_end3 = 1371, 1924, "end"
#missing/response range
missing_range_50 = range(520,571)
missing_range_100 = range(500, 600)
display(missing_range)
width_half = 25
obs1_start_lhs, obs1_stop_lhs = range(520-width_half,520), range(571, 571 + width_half)
width_half = 50
obs1_start_lhs, obs1_stop_lhs = range(520-width_half,520), range(571, 571+width_half)
# # Example of randomly selected obs below
dat = get_observers("block",
num_observers = 40,
missing = 150,
split = 0.0,
dataset = A,
#observer_range = [[100,225], [275,400]],
#response_range = [[225,275]],
#Marios's request:
#observer_range = [[100,200], [300,400]],
response_range = [[2000,2002]],
aspect = 1)
# +
#unknown function:
# if this is set to 0 the reservoir package will be reinstalled
#assuming it is sitting in a parent directory.
### YOU NEED TO RESTART THE NOTEBOOK after doing this, it also must be stopped manually.
# #%run -i "../reinstall.sh"
# #! pip uninstall --yes reservoir
# #! yes | pip install ../Reservoir/
# -
# # old version of plot_timeseries
def plot_timeseries(prediction_, train, test, titl = "ESN ", series2plot = 0, method = None):
"""
This function makes three plots:
the prediction, the residual, the loss.
It was built for single predictions, but needs to be upgraded to deal with multiple output.
We need to show: average residual, average loss.
"""
assert method != None, "Please choose a method: avg or single"
if method == "single":
# avoid choosing all of the columns. subset by the selected time series.
train, test, prediction = train[:, series2plot], test[:, series2plot], prediction_[:, series2plot]
#key indexes
trainlen, testlen, pred_shape = len(train), len(test), len(prediction)
# set up dataframe
xTrTarg_pd = pd.DataFrame(test)
t = pd.DataFrame(list(range(len(xTrTarg_pd))))
# append time
Target_pd = pd.concat([xTrTarg_pd, t], axis = 1)
Target_pd.columns = ["x", "t"]
#calculate the residual
resid = test.reshape(-1,)[:pred_shape] - prediction.reshape(-1,) #pred_shape[0]
rmse_spec = str(round(myMSE(prediction, test), 5))
full_dat = np.concatenate([train, test], axis = 0)
#display(Target_pd)
fig, ax = plt.subplots(3,1, figsize=(16,10))
#The first plot
#sns.lineplot(x = "t", y = "x", data = Target_pd, color = "black", ax = ax[0])
ax[0].plot(range(full_dat.shape[0]), full_dat,'k', label="Ground Truth",
color = "blue", linewidth = 3, alpha = 0.4)
ax[0].plot(range(trainlen,trainlen+testlen),prediction,'k',
label="prediction", color = "r", linestyle = "--", linewidth = 2, alpha = .95)
#first plot labels
ax[0].set_title(titl + "__: Prediction vs Ground Truth, rmse_: " + rmse_spec)
ax[0].legend(loc=(0.31, 1.1))#(0.61, 1.1))
# the second plot
ax[1].plot(range(0,trainlen),np.zeros(trainlen),'k',
label="", color = "black", alpha = 0.5)
ax[1].plot(range(trainlen,trainlen+testlen),resid.reshape(-1,),'k',
color = "purple", alpha = 0.5)
# second plot labels
#ax[1].legend(loc=(0.61, 1.1))
ax[1].set_title(titl + "__: Prediction Residual")
# the second plot
ax[2].plot(range(0,trainlen),np.zeros(trainlen),'k',
label="", color = "black", alpha = 0.5)
ax[2].plot(range(trainlen,trainlen+testlen),resid.reshape(-1,)**2,'k',
color = "r", alpha = 0.5)
# second plot labels
#ax[2].legend(loc=(0.61, 1.1))
ax[2].set_title(titl + "__: Prediction Loss")
plt.subplots_adjust(hspace=0.5)
plt.show()
# # class inspection:
""" Here's how to inspect a class:
import inspect
for i in inspect.getmembers(experiment.unif_esn_cv):
#print(i)
#experiment.best_arguments = unif_cv.optimize(x = experiment.Train, y = experiment.xTr)
"""
|
MARIOS/secondary_notebooks/spectogram_data_ip.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import gzip, json, glob
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
plt.rcParams.update({
"text.usetex": False,
"font.family": "serif",
#"font.serif": ["Palatino"],
"legend.frameon": False,
"legend.fancybox": False,
'font.size': 8,
'axes.linewidth': 0.6,
'xtick.major.width': 0.6,
'ytick.major.width': 0.6,
'xtick.minor.width': 0.6,
'ytick.minor.width': 0.6,
"lines.linewidth": 0.9,
"axes.grid": True,
"grid.color": "#EEE"
})
plt.rc("text.latex", preamble=r"\usepackage{amsmath}")
# +
def load_files(pattern): # data map (example_i[, target_label]) => data
data = {}
max_memory = 0.0
for filename in glob.glob(pattern):
print("Reading file:", filename)
with gzip.open(filename, "r") as f:
lines = f.readlines()
for line in lines:
j = json.loads(line)
#print(filename, j.keys())
example_i = j["example_i"]
if "target_label" in j:
key = (example_i, j["target_label"])
else:
key = example_i
d = data.get(key, {})
data[key] = d
if "veritas_deltas" in j:
max_memory = max(max_memory, max(x["memory"][-1] for x in j["veritas_log"]))
try: column_prefix = f"veritas{j['max_time']:02d}"
except:
s0 = filename.find("time")+4
s1 = filename.find("-", s0)
max_time = int(filename[s0:s1])
#print("no max time in", filename, f"extracted '{max_time}' from filename")
column_prefix = f"veritas{max_time:02d}"
d[f"{column_prefix}_time"] = j["veritas_time"]
d[f"{column_prefix}_delta"] = j["veritas_deltas"][-1][0]
#print("deltas", j["veritas_deltas"])
if "merge_ext" in j and "max_clique" in j["merge_ext"]:
column_prefix = f"mext_T{j['merge_ext']['max_clique']}_L{j['merge_ext']['max_level']}"
d[f"{column_prefix}_time"] = j["merge_ext"]["times"][-1]
d[f"{column_prefix}_delta"] = j["merge_ext"]["deltas"][-1]
if "kantchelian" in j:
column_prefix = "kan"
d[f"{column_prefix}_time"] = j["kantchelian"]["time_p"]
d[f"{column_prefix}_delta"] = j["kantchelian_delta"]
#print(j["kantchelian"]["bounds"])
print(f"max_memory for {pattern} is: {max_memory/(1024*1024)}")
return data
def get_column_names(data):
columns = set()
for value in data.values():
columns |= value.keys()
return sorted(columns)
def to_df(data):
colnames = get_column_names(data)
columns = {}
index = pd.Series(list(data.keys()))
for c in colnames:
values = {}
for key, value in data.items():
if c in value:
values[key] = value[c]
columns[c] = values
df = pd.DataFrame(columns)
df.set_index(index)
return df
def load_to_df(pattern, dropna=True):
data = load_files(pattern)
df = to_df(data)
df.sort_index(inplace=True, axis=0)
if dropna: df = df.dropna()
return df
# -
dfs={}
dfs["covtype"] = load_to_df("/home/laurens/repos/veritas/tests/experiments/results/r1-covtype-*")
dfs["f-mnist"] = load_to_df("/home/laurens/repos/veritas/tests/experiments/results/r1-f-mnist-*")
dfs["higgs"] = load_to_df("/home/laurens/repos/veritas/tests/experiments/results/r1-higgs-*")
dfs["ijcnn1"] = load_to_df("/home/laurens/repos/veritas/tests/experiments/results/r1-ijcnn1-*")
dfs["mnist"] = load_to_df("/home/laurens/repos/veritas/tests/experiments/results/r1-mnist-*")
dfs["webspam"] = load_to_df("/home/laurens/repos/veritas/tests/experiments/results/r1-webspam-*")
dfs["mnist2v6"] = load_to_df("/home/laurens/repos/veritas/tests/experiments/results/r1-mnist2v6-*")
gridcolor="#EEEEEE"
# # Bound difference
# +
#datasets = ["covtype", "f-mnist", "higgs"]
#datasets = ["ijcnn1", "mnist", "webspam", "mnist2v6"]
datasets = ["covtype", "f-mnist", "higgs", "ijcnn1", "mnist", "webspam", "mnist2v6"]
datasets = ["webspam"]
fig, axs = plt.subplots(1, len(datasets), figsize=(len(datasets)*4.0, 1.8))
fig.subplots_adjust(left=0.15, right=0.9, top=0.85, bottom=0.25)
axs=[axs]
for d, ax in zip(datasets, axs):
df = dfs[d]
#display(d, df)
time_columns = [c for c in df.columns if c.endswith("time")]
delta_columns = [c for c in df.columns if c.endswith("delta")]
time_mean = df[time_columns].mean()
time_std = df[time_columns].std()
#div_from_opt = df[delta_columns].subtract(df["kan_delta"], axis=0).abs().mean()
div_from_opt = df[delta_columns].mean()
speedup = (1.0/df[time_columns].divide(df["kan_time"], axis=0)).mean()
scale_ = np.log10(div_from_opt.max().max()).round()
scale = 10**-scale_ * 10
print("scale", d, scale)
#div_from_opt *= scale
veritas_time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("veritas")]
veritas_delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("veritas")]
mer_time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("mext")]
mer_delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("mext")]
#ax.set_title(f"{d} (n={len(df)})")
ax.set_title(f"{d}")
ax.set_xlabel("Time")
#ax.set_ylabel("Robustness delta value")
#if scale != 1.0:
# ax.text(-0.2, 1.1, f'$\\delta \\times 10^{{{scale_:.0f}}}$', horizontalalignment='left', verticalalignment='center', transform=ax.transAxes)
#else:
ax.text(-0.1, 1.1, f'$\\delta$', horizontalalignment='left', verticalalignment='center', transform=ax.transAxes)
#ax.plot(time_mean[veritas_time_columns], div_from_opt[veritas_delta_columns], marker=".", linestyle="-", markersize=4, label="Veritas")
ax.errorbar(time_mean[veritas_time_columns], div_from_opt[veritas_delta_columns], xerr=time_std[veritas_time_columns],
capthick=1.0, elinewidth=None, capsize=2.0, marker=".", linestyle=":", markersize=4, errorevery=4, label="Veritas")
#for i, (x, y, m) in enumerate(zip(time_mean[veritas_time_columns], div_from_opt[veritas_delta_columns], speedup[veritas_time_columns])):
# ax.text(x, y-0.1, f"{m:.0f}×", horizontalalignment='right', verticalalignment='top', c="gray")
#l, = ax.plot(time_mean[mer_time_columns], div_from_opt[mer_delta_columns], marker="8", markersize=5, linestyle=":", label="Merge")
#for i, (x, y, m) in enumerate(zip(time_mean[mer_time_columns], div_from_opt[mer_delta_columns], speedup[mer_time_columns])):
# ax.text(x, y-0.1, f"{m:.0f}×", horizontalalignment='right', verticalalignment='top', c="gray")
#ax.axhline(y=div_from_opt[mer_delta_columns][0], c=l.get_color(), ls=l.get_linestyle())
ax.errorbar(time_mean[mer_time_columns], div_from_opt[mer_delta_columns], xerr=time_std[mer_time_columns],
capthick=1.0, elinewidth=None, capsize=2.0, marker="8", markersize=5, linestyle="", label="Merge")
#l, = ax.plot(time_mean[["kan_time"]], div_from_opt[["kan_delta"]], marker="*", linestyle=":", markersize=4, label="MILP")
ax.errorbar(time_mean[["kan_time"]], div_from_opt[["kan_delta"]], marker="*", linestyle="", markersize=4,
xerr=time_std[["kan_time"]],
capthick=1.0, elinewidth=0.0, capsize=2.0, barsabove=True, label="MILP")
ax.axhline(y=div_from_opt["kan_delta"], c="gray", ls=":", label="Exact")
ax.set_xscale("log")
#xlim = (0.0, 1.1*time_mean["kan_time"])
#ax.set_xlim(xlim)
#ax.set_xticks(list(np.arange(0.0,xlim[1], 10.0)))
ax.legend(fontsize="large", bbox_to_anchor=(1.0, 0.8))
plt.savefig(f"/tmp/bound_err_{datasets[0]}.pdf")
plt.show()
# -
# # Counting stats
# +
fig, axs = plt.subplots(1, len(dfs), figsize=(len(dfs)*5, 4))
better_stats = {}
worse_stats = {}
same_stats = {}
for (d, df), ax in zip(dfs.items(), axs):
time_columns = [c for c in df.columns if c.endswith("time")]
delta_columns = [c for c in df.columns if c.endswith("delta")]
time_mean = df[time_columns].mean()
veritas_time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("veritas")]
veritas_delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("veritas")]
mer_delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("mext")]
mer_time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("mext")]
div_from_opt = df[delta_columns].subtract(df["kan_delta"], axis=0).abs()
same_threshold = (df["kan_delta"].quantile(0.8) - df["kan_delta"].quantile(0.2)) / 100
print(f"same threshold {d}: {same_threshold}")
ver_better = div_from_opt[delta_columns].subtract(div_from_opt[mer_delta_columns[0]], axis=0) < -same_threshold
ver_worse = div_from_opt[delta_columns].subtract(div_from_opt[mer_delta_columns[0]], axis=0) > same_threshold
ver_same = ~ver_better & ~ver_worse
#display(ver_better.sum(), ver_worse.sum(), ver_same.sum())
n = len(df)
ax.set_title(f"{d} (n={n})")
ax.set_xlabel("Time [s]")
#ax.set_ylabel("Robustness delta value")
ax.text(-0.1, 1.04, '%', horizontalalignment='right', verticalalignment='center', transform=ax.transAxes)
ax.plot(time_mean[veritas_time_columns], ver_better.sum()[veritas_delta_columns]/n, marker="^", linestyle=":", label="Better")
ax.plot(time_mean[veritas_time_columns], ver_worse.sum()[veritas_delta_columns]/n, marker="v", linestyle=":", label="Worse")
ax.plot(time_mean[veritas_time_columns], ver_same.sum()[veritas_delta_columns]/n, marker=".", linestyle=":", label="Same")
ax.axvline(x=time_mean[mer_time_columns[0]], ls="--", color="gray", label="Merge time")
#ax.set_xscale("log")
ax.legend()
better_stats[d] = ver_better.sum()[veritas_delta_columns]/n
worse_stats[d] = ver_worse.sum()[veritas_delta_columns]/n
same_stats[d] = ver_same.sum()[veritas_delta_columns]/n
# -
better_df = (pd.DataFrame(better_stats).transpose()*100).round(1)
worse_df = (pd.DataFrame(worse_stats).transpose()*100).round(1)
same_df = (pd.DataFrame(same_stats).transpose()*100).round(1)
display(better_df, worse_df, same_df)
# # How many problems are solved in 1s, 2s, ...
# +
datasets = ["covtype", "f-mnist", "higgs", "ijcnn1", "mnist", "mnist2v6"]
fig, axs = plt.subplots(1, len(datasets), figsize=(len(datasets)*1.4, 1.8), sharey=True, sharex=True)
fig.subplots_adjust(left=0.04, bottom=0.22, right=0.99
, top=0.7, wspace=0.1, hspace=0.4)
axs = axs.flatten()
better_stats = {}
worse_stats = {}
same_stats = {}
for d, ax in zip(datasets, axs):
df = dfs[d]
time_columns = [c for c in df.columns if c.endswith("time")]
delta_columns = [c for c in df.columns if c.endswith("delta")]
time_mean = df[time_columns].mean()
veritas_time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("veritas")]
veritas_delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("veritas")]
mer_delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("mext")]
mer_time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("mext")]
times = np.linspace(0, 12, 50)
div_from_opt = df[veritas_delta_columns+mer_delta_columns].subtract(df["kan_delta"], axis=0).abs()
#q10, q50, q90 = pd.Series(div_from_opt.to_numpy().flatten()).quantile([0.25, 0.5, 1.0])
#q50 = pd.Series(div_from_opt.to_numpy().flatten()).median()
q50 = pd.Series(df[mer_delta_columns].subtract(df["kan_delta"], axis=0).abs().to_numpy().flatten()).mean()
#print(d, "quantiles:", q10, q50, q90)
#print(" how often do we see median? ", (div_from_opt==q50).sum().sum()/len(df) * 100)
#print(" how many unique error values?", len(pd.Series(div_from_opt.to_numpy().flatten()).unique()), np.prod(div_from_opt.shape))
#in_time_ver = pd.concat([df[veritas_time_columns[0]]]*len(times), axis=1).le(times, axis=1)
#in_time_ver.columns = [f"in_time{t:.2f}" for t in times]
#in_time_mer = pd.concat([df[mer_time_columns[0]]]*len(times), axis=1).le(times, axis=1)
#in_time_mer.columns = [f"in_time{t:.2f}" for t in times]
in_time_kan = pd.concat([df["kan_time"]]*len(times), axis=1).le(times, axis=1)
in_time_kan.columns = [f"in_time{t:.2f}" for t in times]
in_time_ver10 = None
in_time_ver50 = None
in_time_ver90 = None
for tcol, dcol in zip(veritas_time_columns, veritas_delta_columns):
#x10 = pd.concat([(df[tcol]<=t) & ((df[dcol]-df["kan_delta"]).abs()<=q10) for t in times], axis=1)
x50 = pd.concat([(df[tcol]<=t) & ((df[dcol]-df["kan_delta"]).abs()<=q50) for t in times], axis=1)
#x90 = pd.concat([(df[tcol]<=t) & ((df[dcol]-df["kan_delta"]).abs()<=q90) for t in times], axis=1)
#in_time_ver10 = (in_time_ver10 | x10) if in_time_ver10 is not None else x10
in_time_ver50 = (in_time_ver50 | x50) if in_time_ver50 is not None else x50
#in_time_ver90 = (in_time_ver90 | x90) if in_time_ver90 is not None else x90
#in_time_mer10 = pd.concat([(df[mer_time_columns[0]]<=t) & ((df[mer_delta_columns[0]]-df["kan_delta"]).abs()<=q10) for t in times], axis=1)
in_time_mer50 = pd.concat([(df[mer_time_columns[0]]<=t) & ((df[mer_delta_columns[0]]-df["kan_delta"]).abs()<=q50) for t in times], axis=1)
#in_time_mer90 = pd.concat([(df[mer_time_columns[0]]<=t) & ((df[mer_delta_columns[0]]-df["kan_delta"]).abs()<=q90) for t in times], axis=1)
n = len(df)
#ax.set_title(f"{d} (n={n}, m={q50:.2g})")
#ax.set_title(f"{d} (n={n})")
ax.set_title(f"{d}")
ax.set_xlabel("Time")
#ax.set_ylabel("Robustness delta value")
if d=="covtype":# or d=="ijcnn1":
ax.text(-0.1, 1.09, '%', horizontalalignment='right', verticalalignment='center', transform=ax.transAxes)
#lv, = ax.plot(times, in_time_ver.mean()*100, ls=(0, (2, 4)))
#lm, = ax.plot(times, in_time_mer.mean()*100, ls=(0, (1, 4)))
lv, = ax.plot(times, in_time_ver50.mean()*100, ls="-", label="Veritas")
#ax.fill_between(times, in_time_ver10.mean()*100, in_time_ver90.mean()*100, alpha=0.1, color=lv.get_color())
#ax.plot(times, in_time_ver10.mean()*100, ls=(0, (1, 4)), c=lv.get_color())
#ax.plot(times, in_time_ver90.mean()*100, ls=(0, (1, 4)), c=lv.get_color())
lm, = ax.plot(times, in_time_mer50.mean()*100, ls="--", label="Merge")
#ax.fill_between(times, in_time_mer10.mean()*100, in_time_mer90.mean()*100, alpha=0.1, color=lm.get_color())
#ax.plot(times, in_time_mer10.mean()*100, ls=(0, (1, 4)), c=lm.get_color())
#ax.plot(times, in_time_mer90.mean()*100, ls=(0, (1, 4)), c=lm.get_color())
ax.plot(times, in_time_kan.mean()*100, ls="-.", label="MILP")
axs[1].legend(ncol=3, bbox_to_anchor=(3.4, 1.6), fontsize="large")
#for ax in axs[3:]: ax.set_xlabel("Time")
plt.savefig("/tmp/solved_per_time.pdf")
# -
# # Tables
# +
rows = {}
def map_name(n):
if "kan" in n:
return "MIPS"
if "veritas" in n:
#return f"$\\ouralg{{}}_{{{int(n[7:9])}}}$"
return "\\ouralg{}"
if "mext" in n:
return "\\merge{}"
def which_column(d):
if d == "f-mnist":
return "veritas06"
else:
return "veritas02"
for i, (d, df) in enumerate(dfs.items()):
time_columns = [c for c in df.columns if c.endswith("time") and (not c.startswith("veritas") or c.startswith(which_column(d)))]
delta_columns = [c for c in df.columns if c.endswith("delta") and (not c.startswith("veritas") or c.startswith(which_column(d)))]
time_mean = df[time_columns].mean()
r1 = df[delta_columns].mean()
r1[r1.index[1:]] /= r1[r1.index[0]]
r1[r1.index[1:]] *= 100.0
r1[r1.index[1:]] = [f"\\SI{{{x:.3g}}}{{\percent}}" for x in r1[r1.index[1:]]]
r1.index = [map_name(n) for n in r1.index]
r2 = df[time_columns].mean()
r2.index = [map_name(n) for n in r2.index]
r3 = df[time_columns].std()
r3.index = [map_name(n) for n in r3.index]
r4 = df[time_columns].mean()
r4[r4.index[1:]] = r4[r4.index[0]] / r4[r4.index[1:]]
r4[r4.index[0]] = ""
r4[r4.index[1:]] = [f"\\SI{{{x:.0f}}}{{\times}}" for x in r4[r4.index[1:]]]
r4.index = [map_name(n) for n in r4.index]
rows[(d, "$\\delta$")] = r1
rows[(d, "$t$")] = r2
rows[(d, "$\\times$")] = r4
rows[(d, "$\\sigma_t$")] = r3
means_df = pd.DataFrame(rows)
means_df = means_df.transpose()
means_df
# -
print(means_df.to_latex(escape=False))
# +
# Counts table
rows = {}
def map_name(n):
if "veritas" in n:
#return f"$\\ouralg{{}}_{{{int(n[7:9])}}}$"
return f"Budget {int(n[7:9])}"
for i, (d, df) in enumerate(dfs.items()):
time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("ver")]
delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("ver")]
mer_time_column = [c for c in df.columns if c.endswith("time") and c.startswith("mext")][0]
mer_delta_column = [c for c in df.columns if c.endswith("delta") and c.startswith("mext")][0]
kan_delta = df["kan_delta"]
same_threshold = (kan_delta.quantile(0.6) - kan_delta.quantile(0.4)) / 1000
mer_abs_diff = (df[mer_delta_column] - kan_delta).abs()
r1 = df[delta_columns].subtract(kan_delta, axis=0).abs().lt(mer_abs_diff, axis=0).mean()
r1 *= 100.0
r1.index = [map_name(n) for n in r1.index]
r2 = df[delta_columns].subtract(kan_delta, axis=0).subtract(mer_abs_diff, axis=0).le(-same_threshold).mean()
r2 *= 100.0
r2.index = [map_name(n) for n in r2.index]
r3 = df[delta_columns].subtract(kan_delta, axis=0).subtract(mer_abs_diff, axis=0).ge(same_threshold).mean()
r3 *= 100.0
r3.index = [map_name(n) for n in r3.index]
r4 = df[delta_columns].subtract(kan_delta, axis=0).subtract(mer_abs_diff, axis=0).abs().lt(same_threshold).mean()
r4 *= 100.0
r4.index = [map_name(n) for n in r4.index]
r5 = df[time_columns].lt(df[mer_time_column], axis=0).mean()
r5 *= 100.0
r5.index = [map_name(n) for n in r5.index]
r6_a = df[time_columns].gt(df[mer_time_column], axis=0)
r6_b = df[delta_columns].subtract(kan_delta, axis=0).subtract(mer_abs_diff, axis=0).ge(same_threshold)
r6_a.columns = [map_name(n) for n in r6_a.columns]
r6_b.columns = [map_name(n) for n in r6_b.columns]
r6 = (r6_a & r6_b).mean()
r6 *= 100.0
#rows[(d, "r1")] = r1
rows[(d, "better")] = r2
rows[(d, "worse")] = r3
rows[(d, "same")] = r4
rows[(d, "faster")] = r5
rows[(d, "slower and worse")] = r6
counts_df = pd.DataFrame(rows)
counts_df = counts_df.transpose()
counts_df.round(1)
# -
formatter = lambda x: f"\\SI{{{x:.1f}}}{{\percent}}"
print(counts_df.to_latex(escape=False, formatters=[formatter] * 5))
time_columns = [c for c in df.columns if c.endswith("time")]
delta_columns = [c for c in df.columns if c.endswith("delta")]
veritas_time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("veritas")]
veritas_delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("veritas")]
mer_time_columns = [c for c in df.columns if c.endswith("time") and c.startswith("mext")]
mer_delta_columns = [c for c in df.columns if c.endswith("delta") and c.startswith("mext")]
delta_mean = df[delta_columns].mean()
time_mean = df[time_columns].mean()
df[delta_columns]
plt.errorbar(time_mean, delta_mean, marker="x", ls="", xerr=df[time_columns].std())#, yerr=df[delta_columns].std())
time_mean[veritas_time_columns]
div_from_opt = df[delta_columns].subtract(df["kan_delta"], axis=0).abs().mean()
plt.title("Mean absolute difference of delta value")
plt.plot(time_mean[veritas_time_columns], div_from_opt[veritas_delta_columns], marker=".", linestyle=":", label="Veritas")
l, = plt.plot(time_mean[mer_time_columns], div_from_opt[mer_delta_columns], marker="o", linestyle=":", label="Merge")
plt.axhline(y=div_from_opt[mer_delta_columns][0], c=l.get_color(), ls=l.get_linestyle())
l, = plt.plot(time_mean[["kan_time"]], div_from_opt[["kan_delta"]], marker="x", linestyle=":", label="MILP")
plt.axhline(y=div_from_opt["kan_delta"], c=l.get_color(), ls=l.get_linestyle())
plt.legend()
plt.title("Mean delta value")
plt.plot(time_mean[veritas_time_columns], delta_mean[veritas_delta_columns], marker=".", linestyle=":", label="Veritas")
l, = plt.plot(time_mean[mer_time_columns], delta_mean[mer_delta_columns], marker="o", linestyle=":", label="Merge")
plt.axhline(y=delta_mean[mer_delta_columns][0], c=l.get_color(), ls=l.get_linestyle())
l, = plt.plot(time_mean[["kan_time"]], delta_mean[["kan_delta"]], marker="x", linestyle=":", label="MILP")
plt.axhline(y=delta_mean["kan_delta"], c=l.get_color(), ls=l.get_linestyle())
plt.legend()
time_mean
df[delta_columns].subtract(df["kan_delta"], axis=0).describe()
df[df["kan_delta"]>20]
dfs["f-mnist"] = load_to_df("/home/laurens/repos/veritas/tests/experiments/results/r1-f-mnist-time2*")
# +
df = dfs["mnist"]
# -
|
tests/experiments/notebooks/robustness-all.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import math
import torch
import gpytorch
from matplotlib import pyplot as plt
import seaborn as sns
from sklearn.preprocessing import StandardScaler, MinMaxScaler
import pandas as pd
import numpy as np
from scipy.stats import norm
from gp_dev.core import *
from ddop.datasets import load_yaz
from pathlib import Path
import datetime
import category_encoders as ce
# %matplotlib inline
# %load_ext autoreload
# %autoreload 2
from sklearn.gaussian_process import GaussianProcessRegressor
from sklearn.gaussian_process.kernels import RBF, WhiteKernel, RationalQuadratic, ExpSineSquared
path = Path("..")
path_ds = path/'datasets'
path_res = path/'res_data'
path_plot = path/'plots'
# +
res = []
test = {}
products = ['CALAMARI', 'FISH', 'SHRIMP', 'CHICKEN', 'KOEFTE', 'LAMB', 'STEAK']
for method in ['one-hot encoding', 'target encoding']:
if method == 'one-hot encoding':
df = load_yaz(encode_date_features=True, include_prod=None, include_date=False).frame
else:
df = load_yaz(encode_date_features=False, include_prod=None, include_date=False).frame
# make train/val/test
n_train = 600
train_df, test_df = df.iloc[:n_train, :], df.iloc[n_train:, :]
n_test = len(test_df)
train_df = pd.melt(train_df,
id_vars = train_df.columns.difference(products),
value_vars= products)
test_df = pd.melt(test_df,
id_vars = test_df.columns.difference(products),
value_vars= products)
train_x_df, train_y_df = train_df.iloc[:, :-1], train_df.iloc[:, -1]
test_x_df, test_y_df = test_df.iloc[:, :-1], test_df.iloc[:, -1]
train_y = train_y_df.values
test_y = test_y_df.values
# target encoding
if method == 'target encoding':
for cat in ['WEEKDAY', 'MONTH', 'YEAR', 'ISHOLIDAY', 'WEEKEND', 'variable']:
encoder = ce.TargetEncoder()
train_x_df[cat] = encoder.fit_transform(train_x_df[cat].astype('category'), train_y_df)
test_x_df[cat] = encoder.transform(test_x_df[cat].astype('category'))
elif method == 'one-hot encoding':
for cat in ['variable']:
encoder = ce.OneHotEncoder()
train_x_df = pd.concat([train_x_df, encoder.fit_transform(train_x_df[cat].astype('category'), train_y_df)], axis=1).drop(columns = cat)
test_x_df = pd.concat([test_x_df, encoder.transform(test_x_df[cat].astype('category'))], axis=1).drop(columns = cat)
scaler = MinMaxScaler()
scaler.fit(train_x_df)
train_x = scaler.transform(train_x_df)
test_x = scaler.transform(test_x_df)
# Kernel with parameters given in GPML book
k1 = 1**2 * RBF(length_scale=0.261) # long term smooth rising trend
k2 = 2.4**2 * RBF(length_scale=90.0) \
* ExpSineSquared(length_scale=1.3, periodicity=1.0) # seasonal component
k3 = 0.66**2 \
* RationalQuadratic(length_scale=1.2, alpha=0.78) # medium term irregularity
k4 = 0.18**2 * RBF(length_scale=0.134) \
+ WhiteKernel(noise_level=1.09**2) # noise terms
if method == 'timeseries':
kernel_gpml = k1 + k2 + k3 + k4
elif method == 'one-hot encoding':
kernel_gpml = k1 + k4
elif method == 'target encoding':
kernel_gpml = k1 + k4
gp = GaussianProcessRegressor(kernel=kernel_gpml, normalize_y=True)#, alpha=1)
gp.fit(train_x, train_y)
print("\nLearned kernel: %s" % gp.kernel_)
print("Log-marginal-likelihood: %.3f"
% gp.log_marginal_likelihood(gp.kernel_.theta))
nv_means, y_std = gp.predict(test_x, return_std=True)
nv_sigma = y_std
for i, target in enumerate(products):
for c in range(5,100, 5):
cu = c/100
co = 1-cu
nv_solution = nv_means[i*n_test:(i+1)*n_test]+norm.ppf(cu/(cu+co))*nv_sigma[i*n_test:(i+1)*n_test]
cost = np.mean([nv_cost(q, y, cu, co) for q, y in zip(nv_solution, test_y[i*n_test:(i+1)*n_test])])
test[target, c, 'diff', 'test'] = nv_solution - test_y[i*n_test:(i+1)*n_test]
test[target, c, 'solution', 'test'] = nv_solution
test[target, c, 'value', 'test'] = train_y[i*n_test:(i+1)*n_test]
test[target, c, 'mean', 'test'] = nv_means[i*n_test:(i+1)*n_test]
test[target, c, 'std', 'test'] = nv_sigma[i*n_test:(i+1)*n_test]
ser_tmp=pd.Series({"cu":cu, "co":co, "cost":cost, "type":method, "target": target, "split": 'test'})
res.append(ser_tmp)
nv_means, y_std = gp.predict(train_x, return_std=True)
nv_sigma = y_std
for i, target in enumerate(products):
for c in range(5,100, 5):
cu = c/100
co = 1-cu
nv_solution = nv_means[i*n_train:(i+1)*n_train]+norm.ppf(cu/(cu+co))*nv_sigma[i*n_train:(i+1)*n_train]
cost = np.mean([nv_cost(q, y, cu, co) for q, y in zip(nv_solution, train_y[i*n_train:(i+1)*n_train])])
test[target, c, 'diff', 'train'] = nv_solution - train_y[i*n_train:(i+1)*n_train]
test[target, c, 'solution', 'train'] = nv_solution
test[target, c, 'value', 'train'] = train_y[i*n_train:(i+1)*n_train]
test[target, c, 'mean', 'train'] = nv_means[i*n_train:(i+1)*n_train]
test[target, c, 'std', 'train'] = nv_sigma[i*n_train:(i+1)*n_train]
ser_tmp=pd.Series({"cu":cu, "co":co, "cost":cost, "type":method, "target": target, "split": 'train'})
res.append(ser_tmp)
#df_res = pd.DataFrame(res)
for target in products:
method = 'saa'
df = load_yaz(encode_date_features=False, include_prod=[target], include_date=False).frame
# make train/val/test
n_train = 600
train_df, test_df = df.iloc[:n_train, :], df.iloc[n_train:, :]
train_x_df, train_y_df = train_df.iloc[:, :-1], train_df.iloc[:, -1]
test_x_df, test_y_df = test_df.iloc[:, :-1], test_df.iloc[:, -1]
train_y = train_y_df.values
test_y = test_y_df.values
for c in range(5,100, 5):
cu = c/100
co = 1-cu
nv_quantile = np.quantile(train_y, q=cu/(cu+co))
cost= np.mean([nv_cost(nv_quantile, y, cu, co) for y in test_y])
nv_means, nv_sigma = 0,0
ser_tmp=pd.Series({"cu":cu, "co":co, "cost":cost, "type":method, "target": target, "split": 'train'})
res.append(ser_tmp)
df_res = pd.DataFrame(res)
# -
for c in range(5,100, 5):
cu = c/100
co = 1-cu
print(norm.ppf(cu/(cu+co)))
# nnn= 'SOF_results_Symmetric.csv'
# df_tmp = pd.read_csv(nnn)
# #df_tmp = df_tmp.drop(columns=["Unnamed: 0"])
# df_tmp['target']="STEAK"
# df_tmp.to_csv(nnn, index=False)
test['CALAMARI', 5, 'std', 'test'].min()
test['CALAMARI', 5, 'std', 'test']
test['CALAMARI', 5, 'mean', 'test']
test['CALAMARI', 5, 'std', 'test'].min()
test['CALAMARI', 5, 'diff', 'test']
test['CALAMARI', 5, 'value', 'test']
test['CALAMARI', 5, 'solution', 'test']
y
df_plot = df_res
#df_plot = pd.read_csv('res_data/gp_all-paste.csv')
df_plot = df_plot[~(df_plot.type.isin(["rf_rf", "rf_grf", "rf_oracle"]))]
#df_plot = df_plot[~(df_plot.type.isin(["rf_approx_risk", "rf_approx_sol", "oracle"]))]
#df_plot = df_plot[~(df_plot.type.isin(["saa", "rf"]))]
sns.set(rc={'figure.figsize':(15,15)})
sns.set_style('whitegrid')
sns.relplot(data=df_plot, x="cu", y="cost",col_wrap=3,facet_kws={'sharey':False},style='split',
col="target", hue="type",kind="line", aspect=1, height=4);
|
nbs_dev/GP_YAZ_scikit_multi.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="DKR4qUAjMbTJ" colab_type="text"
# # enumerate() Fonksiyonu
# + [markdown] id="-TWAb0-jMgMj" colab_type="text"
# enumerate ingilizcede numaralandırmak anlamına gelir. dolayısıyla enumerate() fosnkiyonu bize numaralandırma işlemi yapar. Bu fonskiyon bize 2 farklı bilgi verir:
# bir öğe ve bu öğeye ait bir sıra numarası.
# + [markdown] id="oroCXBgiM7fG" colab_type="text"
# enumarete() fonksiyonu reversed fonksiyonu gibi sade bir şekilde kullanılamaz
# + id="SOzlcXHsNCEr" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="c22f0e61-25e8-4489-839e-f6494a992364"
print(enumerate("istihza")) # bu şekilde kullanılamaz
# + id="6nE5E6fHNXdm" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="d8a85bb0-6915-44d4-d55a-c505602d74cc"
print(*enumerate("istihza"))
# + id="RVWLnCk7NdLV" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="54cf87f8-3afa-4270-be66-22c3ed40bf26"
print(*enumerate("murat"))
# + [markdown] id="kRb7hsfNNnge" colab_type="text"
# for döngüsünde de kullanabiliriz
# + id="9qVypTCCNqyc" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="55785c0a-b327-4ce8-b39d-63c3e2414648"
for m in enumerate("istihza"):
print(m)
# + [markdown] id="4qgzxfVwN29T" colab_type="text"
# Önceki işlemden dir("") komutunu kullanarak metotları sıralamıştık:
# + id="927Cgx01OBZo" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 739} outputId="ef183bbc-3abf-43e2-a52c-c40d8a80bc82"
sayac = 0
for i in dir(""):
if "_" not in i:
sayac += 1
print(i,sayac)
# + [markdown] id="m9hy9R5lOPCW" colab_type="text"
# Aynı işlemi enumerate() fonksiyonu ile kolayca yazabiliriz:
# + id="DLjyYXPZOWVp" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="ea477087-5ddf-499e-a2ff-1964c3571f10"
for sıra,metot in enumerate(dir("")):
print(sıra,metot)
# + [markdown] id="Fbsm_HH2Ohp9" colab_type="text"
# metot ve sıra numaralarını yerlerini değiştirelim:
# + id="W56fo0pnOk4u" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="beac6b52-0472-40c0-ade3-dcd1369606eb"
for sıra,metot in enumerate(dir("")):
print(metot,sıra)
# + id="ITXCe213O0dE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="1aba1141-c3f7-42fe-a719-de2c736fec89"
for sıra,metot in enumerate(dir("")):
print(sıra,metot,len(metot))
# + [markdown] id="isOVqO-XPEtq" colab_type="text"
# Burada dir("") ile elde ettiğimiz metotların sırasını(sıra), metotların adlarını(metot) ve her bir metodun kaç karakterden oluştuğunu ise len(metot) yoluyla çıktı elde ettik.
# + [markdown] id="zkI4I2-KPXvA" colab_type="text"
# İstersek fonskiyonun numaralandırmaya hangi sayıdan başlayacağını seçebiliriz
# + id="s-E0uY5sPd5W" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 134} outputId="ec69c99b-c4a5-4352-f5d7-eb0533d4dbd7"
for sıra,metot in enumerate("istihza",1):
print(sıra,metot)
|
karakterDizileri/enumerateFonksiyon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Qol-ovwtLUoI" outputId="8412b848-729d-41b0-e5f5-71c659e07518"
# %cd '/content/drive/My Drive/AI'
# + id="akQKed_BKnZ_"
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from keras.utils import np_utils
from keras.models import Sequential
from keras.layers import Dense,Dropout,Activation,Flatten,LeakyReLU
from keras.optimizers import Adam
from keras.losses import categorical_crossentropy
from keras.initializers import RandomNormal
from keras.regularizers import l1
# + [markdown] id="bboUEvSNzAUX"
#
# # ساخت دیتاست جدید خطی
#
#
#
#
# + id="qJhF6o0H16ev"
new_file = open('data.csv', 'w')
for i in range(10000):
weights = np.array([-1.7, 1.35, 8.75]).reshape((1, 3))
x = np.random.randn(3) * 5
y = np.matmul(weights, x) + 17.76
new_file.write(
','.join(map(str, list(x) + list(y)))
)
new_file.write('\n')
new_file.close()
# + id="_OQ3skMDKrSu"
dataframe = pd.read_csv("data.csv")
# + id="FXBhEoYpKuZi"
nn_input = np.asarray(dataframe.iloc[:,0:dataframe.shape[1]-1])
nn_output = np.asarray(dataframe.iloc[:,-1])
# + colab={"base_uri": "https://localhost:8080/"} id="o-PAwNJfFiwd" outputId="79695ab7-2797-42c5-b597-692d87a8cb89"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] id="uogdXBLizbaE"
# # تقسیم بندی داده ها به سه دسته ی مستقل آموزشی و ارزیاب و تست
#
# + id="04LoNzphKzqR"
from sklearn.model_selection import train_test_split
x_train, x_temp, y_train, y_temp = train_test_split( nn_input, nn_output, test_size = 0.1, random_state = 55)
x_valid, x_test, y_valid, y_test = train_test_split( x_temp, y_temp, test_size = 0.1, random_state = 65)
# + colab={"base_uri": "https://localhost:8080/"} id="QO-Cg2VHYzwf" outputId="4318b95c-b900-4add-b9db-f7b9d31214a2"
x_train.shape, x_valid.shape, x_test.shape
# + [markdown] id="p-Tbm63hJ5LP"
# # <center dir="rtl"> مدل اول </center>
#
# <div dir="rtl">
#
# در این مدل شبکه ای پیاده سازی شده است که سه لایه دارد، لایه ی اول شامل ۳۰ نرون، لایه ی دوم شامل ۵۰ نرون و لایه ی آخر که خروجی است یک نرون دارد
#
#
# استفاده شده است MSE Lossدر این شبکه از
#
# بعنوان تابع بهینه ساز استفاده شده است Adamهمچنین از تابع
#
#
# :نتیجه
#
# عملکرد این مدل را بررسی MAE ,MSE با استفاده از معیارهای epochs بعد از آموزش شبکه به اندازه ی ۱۰۰
# می کنیم
#
#
# MSE = 0.668 , MAE = 0.035
# </div>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="2zdE1MmKJ4vy" outputId="8274846a-a53a-46f5-8b47-7ede19b3cb11"
model1 = Sequential()
model1.add(Dense(30, input_dim=dataframe.shape[1]-1))
model1.add(Dense(50))
model1.add(Dense(1))
model1.compile(loss='mse', optimizer='adam', metrics=['mse', 'mae', 'mape'])
history = model1.fit(x_test, y_test, epochs=100, batch_size=len(x_test), verbose=1)
predictions = model1.predict(x_test)
print("\n___________________________________________________________")
MSE = np.sum(np.sqrt(np.mean(np.square(predictions - y_test))))/y_test.shape[0]
MAE = np.sum(np.sqrt(np.abs(np.mean(predictions - y_test))))/y_test.shape[0]
print("MSE: ", MSE)
print("MAE: ", MAE)
plt.plot(history.history['loss'])
plt.plot(history.history['mse'])
plt.plot(history.history['mae'])
plt.plot(history.history['mape'])
plt.show()
# + [markdown] id="EedWq9Yz1GXD"
# # <center dir="rtl"> مدل دوم </center>
#
# <div dir="rtl">
#
# در این مدل شبکه ای پیاده سازی شده است که ۷ لایه دارد و هز لایه به ترتیب ۲۰۰، ۵۰۰، ۸۰۰، ۹۰۰، ۵۰۰،، ۱۰۰ و
# لایه ی آخر یک نرون دارد
#
# بعنوان تابع فعال ساز در همه ی لایه ها از Relu استفاده کرده ایم
#
#
# در این شبکه از تابع بهینه ساز Adagrad
# استفاده میکنیم
#
# همچنین مدل ۲۰۰ ایپاک آموزش داده شده است
#
# در این شبکه از تابع بهینه ساز MSE استفاده شده است
#
# :نتیجه
#
# MSE = 0.665 , MAE = 0.0008
#
# </div>
#
# + id="AbHnX9Q7Kzng" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="eb0d7295-d318-4b56-ed37-c073326520bd"
model2 = Sequential()
model2.add(Dense(200, input_dim=dataframe.shape[1]-1, activity_regularizer=l1(0.005)))
model2.add(Activation('relu'))
model2.add(Dense(500, activity_regularizer=l1(0.006)))
model2.add(Activation('relu'))
model2.add(Dense(800, activity_regularizer=l1(0.004)))
model2.add(Activation('relu'))
model2.add(Dense(900, activity_regularizer=l1(0.004)))
model2.add(Activation('relu'))
model2.add(Dense(500, activity_regularizer=l1(0.004)))
model2.add(Activation('relu'))
model2.add(Dense(100, activity_regularizer=l1(0.004)))
model2.add(Activation('relu'))
model2.add(Dense(1, activity_regularizer=l1(0.002)))
model2.compile(loss='mse',
optimizer='adagrad',
metrics=['mse', 'mae', 'mape'])
history = model2.fit(x_train, y_train, epochs = 200,
validation_data=(x_valid, y_valid),
batch_size = 16, verbose = 1)
predictions = model2.predict(x_test)
print("\n___________________________________________________________")
MSE = np.sum(np.sqrt(np.mean(np.square(predictions - y_test))))/y_test.shape[0]
MAE = np.sum(np.sqrt(np.abs(np.mean(predictions - y_test))))/y_test.shape[0]
print("MSE: ", MSE)
print("MAE: ", MAE)
plt.plot(history.history['loss'])
plt.plot(history.history['mse'])
plt.plot(history.history['mae'])
plt.plot(history.history['mape'])
plt.show()
# + [markdown] id="TB0CDV2q1-F5"
# ## <center dir="rtl"> تحلیل عملکرد مدل دوم روی داده های زمان آموزش و تست </center>
#
# <div dir="rtl">
#
# با بررسی معیارهای MSE و MAE
# روی داده های آموزشی میبینیم که این دو معیار بسیار به صفر نزدیک بودند و
# آنچه که در زمان تست بدست آوردیم بسیار تفاوت دارند ازین رو متوجه میشویم که مدل آورفیت شده و مشکل بیش برازش دارد
# علت این مشکل، پیچیده شدن بیش از حد شبکه است بطوری که میتوان گفت شبکه توانسته داده های آموزشی را حفظ میکند اما در زمان مواجهه با داده های جدید با شکست روبرو شده است. در واقع ما در این حالت نتوانسته این بخوبی از امکانات شبکه استفاده کنیم
#
# برای رفع این مشکل شبکه سوم را طراحی میکنیم
#
# </div>
#
# + [markdown] id="eJKuDov_2c5c"
# ## <center dir="rtl"> مدل سوم </center>
#
# <div dir="rtl">
#
# :شبکه قبلی دچار مشکل بیش برازش شده بود برای رفع این مشکل از دو راه حل استفاده کرده ایم
#
# استفاده از Drop out برای منظم سازی
#
# استفاده از تابع بهینه ساز
# rmsprop
# که برای مسئله رگرسیون تابع بهینه ساز مناسب تری است
#
# دیگر ویژگیهای این شبکه تماما مثل شبکه قبلی می باشد
#
# نتیجه:
#
# MSE = 0.558 , MAE = 0.03
#
# تغییرات اعمال شده موجب به رفع مشکل بیش برازش شده و مدل بهبود پیدا کرده است
#
#
#
#
# </div>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="QILQAxoZQi1v" outputId="a97f2224-4c2e-47af-cf84-0eefc61e3c80"
model3 = Sequential()
model3.add(Dense(200, input_dim=dataframe.shape[1]-1, activity_regularizer=l1(0.005)))
model3.add(Activation('relu'))
model3.add(Dropout(0.1))
model3.add(Dense(500, activity_regularizer=l1(0.006)))
model3.add(Activation('relu'))
model3.add(Dropout(0.2))
model3.add(Dense(800, activity_regularizer=l1(0.004)))
model3.add(Activation('relu'))
model3.add(Dropout(0.3))
model3.add(Dense(900, activity_regularizer=l1(0.004)))
model3.add(Activation('relu'))
model3.add(Dropout(0.3))
model3.add(Dense(500, activity_regularizer=l1(0.004)))
model3.add(Activation('relu'))
model3.add(Dropout(0.2))
model3.add(Dense(100, activity_regularizer=l1(0.004)))
model3.add(Activation('relu'))
model3.add(Dropout(0.4))
model3.add(Dense(1, activity_regularizer=l1(0.002)))
model3.compile(loss='mse',
optimizer='rmsprop',
metrics=['mse', 'mae', 'mape'])
history = model3.fit(x_train, y_train, epochs = 200,
validation_data=(x_valid, y_valid),
batch_size = 16, verbose = 1)
predictions = model3.predict(x_test)
print("\n___________________________________________________________")
MSE = np.sum(np.sqrt(np.mean(np.square(predictions - y_test))))/y_test.shape[0]
MAE = np.sum(np.sqrt(np.abs(np.mean(predictions - y_test))))/y_test.shape[0]
print("MSE: ", MSE)
print("MAE: ", MAE)
plt.plot(history.history['loss'])
plt.plot(history.history['mse'])
plt.plot(history.history['mae'])
plt.plot(history.history['mape'])
plt.show()
# + [markdown] id="fBck1n3-3cpj"
# ## <center dir="rtl"> مدل چهارم </center>
#
# <div dir="rtl">
#
# حالا که با افزودن
# Drop out
# به شبکه و همچنین تغییر تابع بهینه ساز توانستیم بر بیش برازش چیره شویم امتحان
# میکنیم که آیا افزایش میزان
# Drop out
# نرونها به شبکه قبلی، آیا موجب بهبود کارائی میشود یا خیر
#
#
#
# نتیجه:
#
# MSE = 0.545 , MAE = 0.042
#
# همانطور که مشخص است این تغییرات تاثیر مثبتی داشته و موجب به بهبود عملکرد مدل شده است
#
# </div>
#
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="lV3Lh-EMyoHE" outputId="e3983311-4825-423a-ef1d-d702a405ffb5"
model4 = Sequential()
model4.add(Dense(200, input_dim=dataframe.shape[1]-1, activity_regularizer=l1(0.005)))
model4.add(Activation('relu'))
model4.add(Dropout(0.2))
model4.add(Dense(500, activity_regularizer=l1(0.006)))
model4.add(Activation('relu'))
model4.add(Dropout(0.3))
model4.add(Dense(800, activity_regularizer=l1(0.004)))
model4.add(Activation('relu'))
model4.add(Dropout(0.4))
model4.add(Dense(900, activity_regularizer=l1(0.004)))
model4.add(Activation('relu'))
model4.add(Dropout(0.5))
model4.add(Dense(500, activity_regularizer=l1(0.004)))
model4.add(Activation('relu'))
model4.add(Dropout(0.4))
model4.add(Dense(100, activity_regularizer=l1(0.004)))
model4.add(Activation('relu'))
model4.add(Dropout(0.3))
model4.add(Dense(1, activity_regularizer=l1(0.002)))
model4.compile(loss='mse',
optimizer='rmsprop',
metrics=['mse', 'mae', 'mape'])
history = model4.fit(x_train, y_train, epochs = 200,
validation_data=(x_valid, y_valid),
batch_size = 16, verbose = 1)
predictions = model4.predict(x_test)
print("\n___________________________________________________________")
MSE = np.sum(np.sqrt(np.mean(np.square(predictions - y_test))))/y_test.shape[0]
MAE = np.sum(np.sqrt(np.abs(np.mean(predictions - y_test))))/y_test.shape[0]
print("MSE: ", MSE)
print("MAE: ", MAE)
plt.plot(history.history['loss'])
plt.plot(history.history['mse'])
plt.plot(history.history['mae'])
plt.plot(history.history['mape'])
plt.show()
# + [markdown] id="qx4C7bcy388T"
# ## <center dir="rtl"> مدل پنجم </center>
#
# <div dir="rtl">
# این مدل تماما مشابه با مدل قبلی ایست با این تفاوت که بجای استفاده از
# MSE
# بعنوان تابع خطا از
# MAE
# استفاده شده است
#
#
# نتیجه:
#
# MSE = 0.515 , MAE = 0.020
#
# همانگونه که انتظار میرود، استفاده از این تابع موجب بهبود معیار
# MAE
# میشود اما
# MSE
# بهبودی نداشته است
# </div>
#
# + id="5v_soV27MfcQ" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="bac0ed78-a639-4f93-f2e6-904ff0237241"
model5 = Sequential()
model5.add(Dense(200, input_dim=dataframe.shape[1]-1, activity_regularizer=l1(0.005)))
model5.add(Activation('relu'))
model5.add(Dropout(0.2))
model5.add(Dense(500, activity_regularizer=l1(0.006)))
model5.add(Activation('relu'))
model5.add(Dropout(0.3))
model5.add(Dense(800, activity_regularizer=l1(0.004)))
model5.add(Activation('relu'))
model5.add(Dropout(0.4))
model5.add(Dense(900, activity_regularizer=l1(0.004)))
model5.add(Activation('relu'))
model5.add(Dropout(0.5))
model5.add(Dense(500, activity_regularizer=l1(0.004)))
model5.add(Activation('relu'))
model5.add(Dropout(0.4))
model5.add(Dense(100, activity_regularizer=l1(0.004)))
model5.add(Activation('relu'))
model5.add(Dropout(0.3))
model5.add(Dense(1, activity_regularizer=l1(0.002)))
model5.compile(loss='mae',
optimizer='rmsprop',
metrics=['mse', 'mae', 'mape'])
history = model5.fit(x_train, y_train, epochs = 200,
validation_data=(x_valid, y_valid),
batch_size = 16, verbose = 1)
predictions = model5.predict(x_test)
print("\n___________________________________________________________")
MSE = np.sum(np.sqrt(np.mean(np.square(predictions - y_test))))/y_test.shape[0]
MAE = np.sum(np.sqrt(np.abs(np.mean(predictions - y_test))))/y_test.shape[0]
print("MSE: ", MSE)
print("MAE: ", MAE)
plt.plot(history.history['loss'])
plt.plot(history.history['mse'])
plt.plot(history.history['mae'])
plt.plot(history.history['mape'])
plt.show()
# + [markdown] id="1himI79T4crZ"
# ## <center dir="rtl"> مدل ششم </center>
#
# <div dir="rtl">
#
# ساختار اصلی شبکه در این مدل همانند شبکه قبلی است اما در آن از تابع خطای
# MAPE
# استفاده شده است. همچنین
# ایپاک ها برابر با ۱۰۰۰۰۰۰۰ ست شده است اما به کمک تکنیک
# early stopping
# تنها تا زمانی فرآیند آموزش
# ادامه می یابد که مدل دچار بیش برازش نشده است
#
# نتیجه:
#
# فرآیند آموزش تا ۲۱۷ ایپاک جلو رفت و سپس به پایان رسید
#
# MSE = 0.516 , MAE = 0.018
#
# به کمک این تغییرات مدل مجددا بهبود یافته است و
# MAE
# بدست آمده دچار کاهش شده است
# </div>
#
# + id="PjK6n529rRhR" colab={"base_uri": "https://localhost:8080/", "height": 1000} outputId="f9804604-a2c8-4b61-b64d-85484fd509a3"
from keras.callbacks import EarlyStopping
model6 = Sequential()
model6.add(Dense(200, input_dim=dataframe.shape[1]-1, activity_regularizer=l1(0.005)))
model6.add(Activation('relu'))
model6.add(Dropout(0.2))
model6.add(Dense(500, activity_regularizer=l1(0.006)))
model6.add(Activation('relu'))
model6.add(Dropout(0.3))
model6.add(Dense(800, activity_regularizer=l1(0.004)))
model6.add(Activation('relu'))
model6.add(Dropout(0.4))
model6.add(Dense(900, activity_regularizer=l1(0.004)))
model6.add(Activation('relu'))
model6.add(Dropout(0.5))
model6.add(Dense(500, activity_regularizer=l1(0.004)))
model6.add(Activation('relu'))
model6.add(Dropout(0.4))
model6.add(Dense(100, activity_regularizer=l1(0.004)))
model6.add(Activation('relu'))
model6.add(Dropout(0.3))
model6.add(Dense(1, activity_regularizer=l1(0.002)))
model3.compile(loss='mape', metrics=['mse', 'mae', 'mape'], optimizer=Adam(lr=1e-3, decay=1e-3 / 200))
es = EarlyStopping(monitor='val_loss', mode='min', verbose=1, patience=200)
history = model3.fit(x_train, y_train, validation_data=(x_valid, y_valid), epochs=10000000, batch_size=100, verbose=2, callbacks=[es])
predictions = model3.predict(x_test)
print("\n___________________________________________________________")
MSE = np.sum(np.sqrt(np.mean(np.square(predictions - y_test))))/y_test.shape[0]
MAE = np.sum(np.sqrt(np.abs(np.mean(predictions - y_test))))/y_test.shape[0]
print("MSE: ", MSE)
print("MAE: ", MAE)
plt.plot(history.history['loss'])
plt.plot(history.history['mse'])
plt.plot(history.history['mae'])
plt.plot(history.history['mape'])
plt.show()
# + [markdown] id="c-lZtSCn5LfK"
# ## <center dir="rtl"> نتیجه گیری نهائی </center>
#
# <div dir="rtl">
#
# با بررسی مدلهای گوناگون به این نتیجه رسیدیم که بهترین تابع بهینه ساز در خصوص این مسئله تابع
# rmsprop
# میباشد
#
# همچنین بسته به اینکه معیار سنجش کارایی مدل ما، چه چیزی است براساس آن تابع خطا را مشخص میکنیم که در حین آموزش همان معیار بهینه سازی شود
#
# از آنجا که نمیدانیم چه تعداد ایپاک برای آموزش شبکه مناسب است از تکنیک
# early stopping
# استفاده کرده و
# و مدل را با تعداد مناسبی ایپاک آموزش میدهیم
#
#
# اگر از شبکه ساده ای برای آموزش مدل استفاده کنیم از امکانات گسترده شبکه عصبی بی بهره بودیم و اگر از شبکه ای پیچیده به تنهایی استفاده کنیم باز هم شبکه بر روی داده هایی که قبلا مشاهده نکرده عملکرد خوبی ندارد، به همین علت شبکه ای پیچیده میسازیم و با استفاده از تکنیکهای مختلف منظم سازی از آورفیت شدن مدل جلوگیری میکنیم
# از جمله این تکنیک ها در مسئله ی رگرسیون میتوان به تکنیک
# Drop out
# اشاره کرد
#
# </div>
#
|
Regression-with-NN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Neural & Behavioral Modeling - Week 3 (Exercises)
# %config IPCompleter.greedy=True
# %matplotlib inline
from numpy import *
from matplotlib.pyplot import *
from IPython.display import *
from scipy.stats import pearsonr, zscore, ttest_ind
import time
# ## 1 Replicate exisiting findings/models (10 points)
# Based on your personal interest, please choose and replicate ONE of the following studies:
#
# 1. <a href="http://ccl.northwestern.edu/netlogo/models/FlockingVeeFormations">Flocking Vee Formations</a> in NetLogo's [Sample Models::Biology]
#
# 2. <a href="http://ccl.northwestern.edu/netlogo/models/WolfSheepPredation">Wolf-Sheep Predation</a> in NetLogo's [Sample Models::Biology]
#
# 3. <a href="https://www.meetup.com/hkexperiment/events/226024232/">MIT Matching Game</a> or <NAME>., & <NAME>. (1986). <a href="https://pdfs.semanticscholar.org/6533/189b1bc1040f59a728a75cc62e7bde401091.pdf">The matching hypothesis reexamined</a>. Journal of Personality and Social Psychology, 51(4), 673.
#
# 4. <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2014). <a href="http://www.mpmlab.org/groups/">The emergence of “us and them” in 80 lines of code: Modeling group genesis in homogeneous populations</a>. Psychological Science, 25(4), 982-990.
#
# 5. <NAME>., & <NAME>. (2015). <a href="http://journals.sagepub.com/doi/abs/10.1177/0956797615605798">Memory transmission in small groups and large networks: An agent-based model</a>. Psychological Science, 26(12), 1909-1917.
# The paper I decide to replicate:
#
# 4. <NAME>., <NAME>., <NAME>., <NAME>., <NAME>., & <NAME>. (2014). <a href="http://www.mpmlab.org/groups/">The emergence of “us and them” in 80 lines of code: Modeling group genesis in homogeneous populations</a>. Psychological Science, 25(4), 982-990.
# ## 1.1 Interacting Agent model
#
# ### Function: interact (without trust)
#
# #### Input:
# the interacting two people (p1,p2) <br>
# #### Model Parameters:
# (reciprocity,transistivity)<br>
# #### Output:
# (Clossness_matrix, Payoff) updated<br>
#
#
def interact(p1,p2,reciprocity,transistivity,Clossness_matrix, Payoff):
prob = [random.random() for i in range(2)]
#print(str(p1)+str(p2))
#print(prob)
# play the prison dilema
result = [-1,-1]
for i in range(len(prob)):
if prob[i] < Clossness_matrix[p1][p2]:
result[i] = 1
#else:
#pass
# Calculate and record the playoff
if sum(result) == 2: # colloborative
#print('Colaborate!')
Payoff[p1].append(1)
Payoff[p2].append(1)
Clossness_matrix[p1][p2] = 1-(1-Clossness_matrix[p1][p2])/reciprocity # reciprocity
Clossness_matrix[p2][p1] = Clossness_matrix[p1][p2]
# transistivity
for i in range(len(Clossness_matrix)):
#print(i)
if i not in [p1,p2]:
if abs(Clossness_matrix[p1][i]-0.5) > abs(Clossness_matrix[p2][i]-0.5):
if Clossness_matrix[p1][i] > 0.5:
Clossness_matrix[p2][i] = 1-(1-Clossness_matrix[p2][i])/transistivity
Clossness_matrix[i][p2] = Clossness_matrix[p2][i]
elif Clossness_matrix[p1][i] < 0.5:
Clossness_matrix[p2][i] = Clossness_matrix[p2][i]/transistivity
Clossness_matrix[i][p2] = Clossness_matrix[p2][i]
else:
pass
elif abs(Clossness_matrix[p2][i]-0.5) > abs(Clossness_matrix[p1][i]-0.5):
if Clossness_matrix[p2][i] > 0.5:
Clossness_matrix[p1][i] = 1-(1-Clossness_matrix[p1][i])/transistivity
Clossness_matrix[i][p1] = Clossness_matrix[p1][i]
elif Clossness_matrix[p2][i] < 0.5:
Clossness_matrix[p1][i] = Clossness_matrix[p1][i]/transistivity
Clossness_matrix[i][p1] = Clossness_matrix[p1][i]
else:
pass
else:
pass
else:
pass
elif sum(result) == -2:
#print('Defeat!')
Payoff[p1].append(-1)
Payoff[p2].append(-1)
Clossness_matrix[p1][p2] = Clossness_matrix[p1][p2]/reciprocity
Clossness_matrix[p2][p1] = Clossness_matrix[p1][p2]
elif result[0] == 1 and result[1] == -1:
#print('Nothing!')
Payoff[p1].append(3)
Payoff[p2].append(-3)
else:
#print('Nothing!')
Payoff[p1].append(-3)
Payoff[p2].append(3)
# Update others playoff = 0 in this round
for i in range(len(Clossness_matrix)):
if i not in [p1,p2]:
Payoff[i].append(0)
#print(result)
#Clossness_matrix = symm(Clossness_matrix)
#print(Clossness_matrix)
# ### Function: clustering coefficient
#
# #### Input:
# closeness_matrix <br>
#
# #### Output:
# clustering_coefficient<br><br>
#
#
# Reference: see the Opsahl, T., & <NAME>. (2009). <a href="https://pdfs.semanticscholar.org/6209/2baa5625985f85667c3d4a0bd8d2f8e9f736.pdf">Clustering in weighted networks.</a> Social Networks, 31, 155–163.
def Clustering_coefficient(cmatrix_ori):
cmatrix= cmatrix_ori.copy()
cmatrix = cmatrix + (random.random((len(cmatrix_ori), len(cmatrix_ori)))-0.5)/1000
fill_diagonal(cmatrix,0)
cmatrix = around(cmatrix)
n = trace(linalg.matrix_power(cmatrix,3))
d = linalg.matrix_power(cmatrix,2).sum()- trace(linalg.matrix_power(cmatrix,2))
#print(d)
if d ==0:
c_eff = 0
else:
c_eff = n/d
return c_eff
# #### Test the Group-forming Model: clustering through time
# reciprocity = 3, transistivity = 2, N = 40, round_number = 10000, iterations = 1
# +
# Model parameters:
reciprocity = 3
transistivity = 2
# How many people & rounds?
sz = 40
rounds_number = 10000
# Intialization
C_sum = []
# Drawing List
Drawing_list = arange(sz)
start = time.time()
# Starting rounds
for count in range(1):
#print(count)
C_eff = []
Clossness_matrix = zeros((sz,sz))+0.5
fill_diagonal(Clossness_matrix,0)
Payoff = [[] for i in range(sz)]
rounds=0
#print(Clossness_matrix)
#print(Payoff)
#print(rounds)
while(rounds<rounds_number):
#print(Drawing_list)
person1 = random.choice(Drawing_list)
person2 = random.choice(delete(Drawing_list,person1))
#person1 = random.choice(Drawing_list)
#person2 = random.choice(Drawing_list[0:-2])
if person2 == person1:
person2+=1
if random.random() < Clossness_matrix[person1][person2]:
#print(Clossness_matrix)
interact(person1, person2,reciprocity,transistivity,Clossness_matrix,Payoff)
rounds+=1
#print("Rounds "+str(rounds))
#print(Clossness_matrix)
C_eff.append(Clustering_coefficient(Clossness_matrix))
#if rounds % 100 == 0:
#print(rounds)
else:
#print(Clossness_matrix)
rounds+=1
C_eff.append(Clustering_coefficient(Clossness_matrix))
C_sum.append(C_eff)
#print(Clossness_matrix)
#print(Payoff)
#print(rounds)
end = time.time()
elapsed = end - start
print ("Time taken: ", elapsed, "seconds.")
#plot(range(rounds_number),C_eff)
xlim(0,5000)
ylim(0,1)
k=[]
for j in range(rounds_number):
k.append(mean([C_sum[i][j] for i in range(len(C_sum))]))
#print(k)
plot(range(1,rounds_number+1),k)
xlabel("Round")
ylabel("Clustering Coefficient")
# -
# ### Function: subgroups for calculating group number and size
#
# #### Input:
# closeness_matrix <br>
#
# #### Output:
# a list summary all the subgroups
#
# +
def Group_size(cmatrix_ori):
cmatrix = cmatrix_ori.copy()
cmatrix = around(cmatrix)
group=[]
all_people=list(range(len(cmatrix)))
for i in range(len(cmatrix)):
if i in all_people:
#print(i)
subgroup = []
subgroup.append(i)
for j in range(i,len(cmatrix)):
if cmatrix[i][j] !=0:
subgroup.append(j)
if len(subgroup)!=0:
if len(all_people) == len(subgroup):
group.append(subgroup)
all_people=[]
break
else:
for i in subgroup:
all_people.remove(i)
#print(subgroup)
group.append(subgroup)
#print(all_people)
return group
# -
# ### Replication of Fig.1 - Visulization:
#
# Use PCA to analysis clossess_matrix, visualize the two major components (z-score standardized)
from sklearn import decomposition
def visulization(cmatrix_ori, r):
cmatrix= cmatrix_ori.copy()
cmatrix = cmatrix + (random.random((len(cmatrix_ori), len(cmatrix_ori)))-0.5)/1000
fill_diagonal(cmatrix,0)
Y = cmatrix.copy()
pca = decomposition.PCA(n_components=2)
pca.fit(Y)
Y = pca.transform(Y)
clf() # clear previous figure
plot(zscore(Y[:,0]),zscore(Y[:,1]),'o')
title("Rounds: "+ str(r))
display(gcf()); clear_output(wait=True) # to allow dynamic plots
# #### Visualize the Group-forming Model: clustering through time
# reciprocity = 3, transistivity = 2, N = 40, round_number = 10000, iterations = 1
# +
# Model parameters:
reciprocity = 3
transistivity = 2
# How many people & rounds?
sz = 40
rounds_number = 10000
# Intialization
C_sum = []
# Drawing List
Drawing_list = arange(sz)
start = time.time()
# Starting rounds
for count in range(1):
#print(count)
C_eff = []
Clossness_matrix = zeros((sz,sz))+0.5
fill_diagonal(Clossness_matrix,0)
Payoff = [[] for i in range(sz)]
rounds=0
#print(Clossness_matrix)
#print(Payoff)
#print(rounds)
while(rounds<rounds_number):
#print(Drawing_list)
person1 = random.choice(Drawing_list)
person2 = random.choice(delete(Drawing_list,person1))
#person1 = random.choice(Drawing_list)
#person2 = random.choice(Drawing_list[0:-2])
if person2 == person1:
person2+=1
if random.random() < Clossness_matrix[person1][person2]:
#print(Clossness_matrix)
interact(person1, person2,reciprocity,transistivity,Clossness_matrix,Payoff)
rounds+=1
#print("Rounds "+str(rounds))
#print(Clossness_matrix)
C_eff.append(Clustering_coefficient(Clossness_matrix))
#if rounds % 100 == 0:
#print(rounds)
else:
#print(Clossness_matrix)
rounds+=1
C_eff.append(Clustering_coefficient(Clossness_matrix))
if rounds % 100 ==0:
visulization(Clossness_matrix, rounds)
C_sum.append(C_eff)
#print(Clossness_matrix)
#print(Payoff)
#print(rounds)
end = time.time()
elapsed = end - start
print ("Time taken: ", elapsed, "seconds.")
#plot(range(rounds_number),C_eff)
#xlim(0,5000)
#ylim(0,1)
#k=[]
#for j in range(rounds_number):
# k.append(mean([C_sum[i][j] for i in range(len(C_sum))]))
#print(k)
#plot(range(1,rounds_number+1),k)
#xlabel("Round")
#ylabel("Clustering Coefficient")
print('Group_number: ' +str(len(Group_size(Clossness_matrix))))
print(Group_size(Clossness_matrix))
# -
# Note that the visualization does not imply the real numbers of group:
# It looks like 4 groups in the plot, but 5 groups according to the cluster analysis.
# ## 1.2 Robust group formation
#
#
# From N = 10 to 100, Is the group formation robust?
#
#
# ### Wrap up the similuation into a function
# +
def Group_form(rc, trans, N_people, r_num, ger_times):
reciprocity = rc
transistivity = trans
# How many people & rounds?
sz = N_people
rounds_number = r_num
# Intialization
C_sum = []
#G_sum = []
# Drawing List
Drawing_list = arange(sz)
start = time.time()
# Starting rounds
for count in range(ger_times):
#print(count)
C_eff = []
Clossness_matrix = zeros((sz,sz))+0.5
fill_diagonal(Clossness_matrix,0)
Payoff = [[] for i in range(sz)]
rounds=0
#print(Clossness_matrix)
#print(Payoff)
#print(rounds)
while(rounds<rounds_number):
#print(Drawing_list)
person1 = random.choice(Drawing_list)
person2 = random.choice(delete(Drawing_list,person1))
#person1 = random.choice(Drawing_list)
#person2 = random.choice(Drawing_list[0:-2])
if person2 == person1:
person2+=1
if random.random() < Clossness_matrix[person1][person2]:
#print(Clossness_matrix)
interact(person1, person2,reciprocity,transistivity,Clossness_matrix, Payoff)
rounds+=1
#print("Rounds "+str(rounds))
#print(Clossness_matrix)
C_eff.append(Clustering_coefficient(Clossness_matrix))
#if rounds % 100 == 0:
#print(rounds)
else:
#print(Clossness_matrix)
rounds+=1
C_eff.append(Clustering_coefficient(Clossness_matrix))
C_sum.append(C_eff)
#G_sum.append(Group_size(Clossness_matrix))
end = time.time()
elapsed = end - start
print ("Time taken: ", elapsed, "seconds.")
#plot(range(rounds_number),C_eff)
xlim(0,5000)
ylim(0,1)
k=[]
for j in range(rounds_number):
k.append(mean([C_sum[i][j] for i in range(len(C_sum))]))
#print(k)
plot(range(1,rounds_number+1),k)
xlabel("Round")
ylabel("Clustering Coefficient")
#print(around(Clossness_matrix))
#summary = Group_size(Clossness_matrix)
#print(G_sum)
#return G_sum
# -
# ### Replication of Fig. 3
# reciprocity = 3, transistivity = 2, N = 10, 25, 40, 55, 70, 85, 100 <br>
# round_number = 10000, iterations = 10 (in the original paper, they use 100,000)
#
N = [10, 25, 40, 55, 70, 85, 100]
for n in N:
Group_form(3,2,n,10000,10)
#group_size = n/group_num
legend(['N = '+ str(n) for n in N])
#print(group_num)
#print(group_size)
# ## 1.3 How reciprocity and transistivity affect group number and size ?
#
# #### Wrap up the similuation customed to group-number-and-size calculations into a function
# +
def Group_form_size(rc, trans, N_people, r_num, ger_times):
reciprocity = rc
transistivity = trans
# How many people & rounds?
sz = N_people
rounds_number = r_num
# Intialization
#C_sum = []
G_sum = []
P_sum = []
# Drawing List
Drawing_list = arange(sz)
start = time.time()
# Starting rounds
for count in range(ger_times):
#print(count)
#C_eff = []
Clossness_matrix = zeros((sz,sz))+0.5
fill_diagonal(Clossness_matrix,0)
Payoff = [[] for i in range(sz)]
rounds=0
#print(Clossness_matrix)
#print(Payoff)
#print(rounds)
while(rounds<rounds_number):
#print(Drawing_list)
person1 = random.choice(Drawing_list)
person2 = random.choice(delete(Drawing_list,person1))
#person1 = random.choice(Drawing_list)
#person2 = random.choice(Drawing_list[0:-2])
if person2 == person1:
person2+=1
if random.random() < Clossness_matrix[person1][person2]:
#print(Clossness_matrix)
interact(person1, person2,reciprocity,transistivity,Clossness_matrix, Payoff)
rounds+=1
#print("Rounds "+str(rounds))
#print(Clossness_matrix)
#C_eff.append(Clustering_coefficient(Clossness_matrix))
#if rounds % 100 == 0:
#print(rounds)
else:
#print(Clossness_matrix)
rounds+=1
#C_eff.append(Clustering_coefficient(Clossness_matrix))
#C_sum.append(C_eff)
G_sum.append(len(Group_size(Clossness_matrix)))
P_sum.append(len(Clossness_matrix)/len(Group_size(Clossness_matrix)))
end = time.time()
elapsed = end - start
print ("Time taken: ", elapsed, "seconds.")
#plot(range(rounds_number),C_eff)
#xlim(0,5000)
#ylim(0,1)
#k=[]
#for j in range(rounds_number):
# k.append(mean([C_sum[i][j] for i in range(len(C_sum))]))
#print(k)
#plot(range(rounds_number),k)
#print(around(Clossness_matrix))
#summary = Group_size(Clossness_matrix)
#print(G_sum)
average_group_num = mean(G_sum)
average_group_size = mean(P_sum)
#print(G_sum)
#print(P_sum)
return average_group_num, average_group_size # 每一次generation的分組結果[[A],[B],[C]],
# -
# ### Replication of Fig. 4
#
# N = 50, round_number = 10000<br>
# low_value = 2, high_value = 8 <br>
# fixed value = 5 <br>
# iterations = 80 (higher than 10 to gain more statistical power)
# +
R = [5]
T = [5]
exam_times = 80
for r in R:
#print('r = '+str(r))
group_num_t_low, group_size_t_low = Group_form_size(r,2,50,10000,exam_times)
group_num_t_high, group_size_t_high = Group_form_size(r,10,50,10000,exam_times)
for t in T:
#print('t = '+str(t))
group_num_r_low, group_size_r_low = Group_form_size(2,t,50,10000,exam_times)
group_num_r_high, group_size_r_high =Group_form_size(10,t,50,10000,exam_times)
# -
# ### Number of Groups
# +
z = ['low','high']
plot(z, [group_num_r_low,group_num_r_high],'-o')
plot(z, [group_num_t_low,group_num_t_high],'-o')
ylim(0,8)
legend(['reciprocity', 'transistivity'])
# -
# ### Group size
z = ['low','high']
plot(z, [group_size_r_low,group_size_r_high],'-o')
plot(z, [group_size_t_low,group_size_t_high],'-o')
legend(['reciprocity', 'transistivity'])
ylim(0,14)
# ## 1.4 How number of players affect group number and size ?
# ### Replication of Fig. 5
# reciprocity = 3, transistivity = 2, N = 50,
# round_number = 30000,(more rounds to gurantee stablization)
# iterations = 30
#
# +
N = [10+5*i for i in range(19)]
print(N)
group_num_N = []
group_size_N = []
for n in N:
print('N=' +str(n))
group_num, group_size = Group_form_size(3,2,n,30000,30)
group_num_N.append(group_num)
group_size_N.append(group_size)
plot(N, group_num_N)
plot(N, group_size_N)
legend(['group_num','group_size'])
xlabel("Number of Players")
ylabel("Number")
# -
# ## 1.4 How trust affect group formation ?
#
# Modify the interation with additional parameter: A <br>
# A > 0 -> more trust between players<br>
# A < 0 -> more suspicion between players<br>
def interact_trust(p1,p2,reciprocity,transistivity,Clossness_matrix, Payoff, A):
prob = [random.random() for i in range(2)]
#print(str(p1)+str(p2))
#print(prob)
# play the prison dilema
result = [-1,-1]
for i in range(len(prob)):
if prob[i] < Clossness_matrix[p1][p2]+A:
result[i] = 1
#else:
#pass
# Calculate and record the playoff
if sum(result) == 2: # colloborative
#print('Colaborate!')
Payoff[p1].append(1)
Payoff[p2].append(1)
Clossness_matrix[p1][p2] = 1-(1-Clossness_matrix[p1][p2])/reciprocity # reciprocity
Clossness_matrix[p2][p1] = Clossness_matrix[p1][p2]
# transistivity
for i in range(len(Clossness_matrix)):
#print(i)
if i not in [p1,p2]:
if abs(Clossness_matrix[p1][i]-0.5) > abs(Clossness_matrix[p2][i]-0.5):
if Clossness_matrix[p1][i] > 0.5:
Clossness_matrix[p2][i] = 1-(1-Clossness_matrix[p2][i])/transistivity
Clossness_matrix[i][p2] = Clossness_matrix[p2][i]
elif Clossness_matrix[p1][i] < 0.5:
Clossness_matrix[p2][i] = Clossness_matrix[p2][i]/transistivity
Clossness_matrix[i][p2] = Clossness_matrix[p2][i]
else:
pass
elif abs(Clossness_matrix[p2][i]-0.5) > abs(Clossness_matrix[p1][i]-0.5):
if Clossness_matrix[p2][i] > 0.5:
Clossness_matrix[p1][i] = 1-(1-Clossness_matrix[p1][i])/transistivity
Clossness_matrix[i][p1] = Clossness_matrix[p1][i]
elif Clossness_matrix[p2][i] < 0.5:
Clossness_matrix[p1][i] = Clossness_matrix[p1][i]/transistivity
Clossness_matrix[i][p1] = Clossness_matrix[p1][i]
else:
pass
else:
pass
else:
pass
elif sum(result) == -2:
#print('Defeat!')
Payoff[p1].append(-1)
Payoff[p2].append(-1)
Clossness_matrix[p1][p2] = Clossness_matrix[p1][p2]/reciprocity
Clossness_matrix[p2][p1] = Clossness_matrix[p1][p2]
elif result[0] == 1 and result[1] == -1:
#print('Nothing!')
Payoff[p1].append(3)
Payoff[p2].append(-3)
else:
#print('Nothing!')
Payoff[p1].append(-3)
Payoff[p2].append(3)
# Update others playoff = 0 in this round
for i in range(len(Clossness_matrix)):
if i not in [p1,p2]:
Payoff[i].append(0)
#print(result)
#Clossness_matrix = symm(Clossness_matrix)
#print(Clossness_matrix)
# #### Wrap up the similuation with trust into a function
# +
def Group_form_trust(rc, trans, N_people, r_num, ger_times, A):
reciprocity = rc
transistivity = trans
# How many people & rounds?
sz = N_people
rounds_number = r_num
# Intialization
C_sum = []
#G_sum = []
# Drawing List
Drawing_list = arange(sz)
start = time.time()
# Starting rounds
for count in range(ger_times):
print('Counts= '+str(count))
C_eff = []
Clossness_matrix = zeros((sz,sz))+0.5
fill_diagonal(Clossness_matrix,0)
Payoff = [[] for i in range(sz)]
rounds=0
#print(Clossness_matrix)
#print(Payoff)
#print(rounds)
while(rounds<rounds_number):
#print(Drawing_list)
person1 = random.choice(Drawing_list)
person2 = random.choice(delete(Drawing_list,person1))
#person1 = random.choice(Drawing_list)
#person2 = random.choice(Drawing_list[0:-2])
if person2 == person1:
person2+=1
if random.random() < Clossness_matrix[person1][person2]:
#print(Clossness_matrix)
interact_trust(person1, person2,reciprocity,transistivity,Clossness_matrix, Payoff, A)
rounds+=1
#print("Rounds "+str(rounds))
#print(Clossness_matrix)
C_eff.append(Clustering_coefficient(Clossness_matrix))
else:
#print(Clossness_matrix)
rounds+=1
C_eff.append(Clustering_coefficient(Clossness_matrix))
if rounds % (r_num/100) == 0:
print('Rounds= '+str(rounds))
C_sum.append(C_eff)
#G_sum.append(Group_size(Clossness_matrix))
end = time.time()
elapsed = end - start
print ("Time taken: ", elapsed, "seconds.")
#plot(range(rounds_number),C_eff)
xlim(0,r_num)
ylim(0,1)
k=[]
for j in range(rounds_number):
k.append(mean([C_sum[i][j] for i in range(len(C_sum))]))
#print(k)
semilogx(range(1,rounds_number+1),k)
xlabel("Round")
ylabel("Clustering Coefficient")
#print(around(Clossness_matrix))
#summary = Group_size(Clossness_matrix)
#print(G_sum)
#return G_sum
# -
# ### Replication of Fig. 7
# reciprocity = 3, transistivity = 2, N = 50, <br>
# A = -0.5 ~ 0.5, interval = 0.1<br>
# round_number = 1000,000<br>
# iterations = 10<br>
#
# +
A = [round(-0.5+0.1*i,1) for i in range(11)]
#A = [0]
for a in A:
print('A= '+ str(a))
Group_form_trust(3,2,50,1000000,10,a)
legend(['A = '+ str(a) for a in A])
#print(A)
#print(group_size)
# -
|
hw04/existingModels.ipynb
|
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cs
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: .NET (C#)
// language: C#
// name: .net-csharp
// ---
// + dotnet_interactive={"language": "csharp"}
#r "./../../../../../../public/src/L4-application/BoSSSpad/bin/Release/net5.0/BoSSSpad.dll"
using System;
using ilPSP;
using ilPSP.Utils;
using BoSSS.Platform;
using BoSSS.Foundation;
using BoSSS.Foundation.XDG;
using BoSSS.Foundation.Grid;
using BoSSS.Solution;
using BoSSS.Application.XNSE_Solver;
using System.IO;
using BoSSS.Foundation.IO;
using ilPSP.Tracing;
using BoSSS.Application.BoSSSpad;
using BoSSS.Solution.Gnuplot;
using BoSSS.Solution.AdvancedSolvers;
using static BoSSS.Application.BoSSSpad.BoSSSshell;
Init();
// + dotnet_interactive={"language": "csharp"}
////================================
//// pt2 of this sheet: Process Data
////================================
// Load Tables
//string basepath = @"B:\BoSSS-smuda\public\doc\handbook\apdx-MPISolverPerformance\strongScaling\IBM_RotCube\";
string basepath = System.IO.Directory.GetCurrentDirectory()+@"\";
List<System.Data.DataTable> LoadedTabs = new List<System.Data.DataTable>();
//LoadedTabs.Add(TableExtensions.LoadFromFile(basepath+"strong_scaling_3_2021-9-24.json"));
LoadedTabs.Add(TableExtensions.LoadFromFile(basepath+"weak_scaling_5_1_p-MG_2021-10-13.json"));
//LoadedTabs.Add(TableExtensions.LoadFromFile(basepath+"strong_scaling_3_1_2021-9-24.json"));
//LoadedTabs.Add(TableExtensions.LoadFromFile(basepath+"strong_scaling_3_3_2021-9-.json"));
var LoadTab = LoadedTabs.Pick(0);
LoadedTabs.ForEach(s=>LoadTab.Merge(s)); LoadTab.Rows.Count()
// + dotnet_interactive={"language": "csharp"}
string[] ColumnNames = new string[]{
"OrthonormalizationMultigrid.AddSol",
"Schwarz.Solve",
"OrthonormalizationMultigrid.MinimizeResidual",
"OrthonormalizationMultigrid.Residual",
"OrthonormalizationMultigrid.Solve",
"overlap_scaling",
"block_solve_level",
"caching",
"nocaching",
"OrthonormalizationMultigrid.VerivyCurrentResidual",
"Restriction",
"Prolongation",
"SolverRun_time",
"LSolverIter",
"DGdegree:Velocity*",
"DOF",
"MGDepth",
"NoOfNodes",
"NoOfCores",
"DirectSolver.Solve",
};
// + dotnet_interactive={"language": "csharp"}
var SesTab = LoadTab.ExtractColumns(ColumnNames);
// + dotnet_interactive={"language": "csharp"}
// Average of Repititions ... pt1: Group Runs
var RegistryOfRuns = new Dictionary<string,List<System.Data.DataRow>>();
foreach(System.Data.DataRow Row in SesTab.Rows){
string test = "";
test += "k"+Convert.ToString(Row["DGdegree:Velocity*"]);
test += "_c"+Convert.ToString(Row["NoOfCores"]);
List<System.Data.DataRow> Reps;
if(RegistryOfRuns.TryGetValue(test,out Reps)){
Reps.Add(Row);
} else {
Reps = new List<System.Data.DataRow>();
Reps.Add(Row);
RegistryOfRuns.Add(test,Reps);
}
} RegistryOfRuns.OrderBy(s=>s.Key[0]).OrderBy(s=>s.Key[1]).Select(s=>new object[]{s.Key,s.Value.Count()})
// + dotnet_interactive={"language": "csharp"}
// Average of Repititions ... pt2: Compute Average
var tmp = SesTab.Clone();
tmp.Clear();
foreach(var kvp in RegistryOfRuns){
var newrow = tmp.NewRow();
var sample = kvp.Value.First();
newrow["DGdegree:Velocity*"]=Convert.ToInt32(sample["DGdegree:Velocity*"]);
newrow["NoOfCores"]=Convert.ToInt32(sample["NoOfCores"]);
foreach(System.Data.DataColumn col in tmp.Columns){
if(col.ColumnName=="DGdegree:Velocity*"||col.ColumnName=="NoOfCores")
continue;
double value=0.0;
foreach(System.Data.DataRow row in kvp.Value){
value+=Convert.ToDouble(row[col.ColumnName]);
}
newrow[col]=value / kvp.Value.Count();
}
tmp.Rows.Add(newrow);
}
SesTab = tmp;
SesTab.Print()
// + dotnet_interactive={"language": "csharp"}
//string thisdir = Directory.GetCurrentDirectory();
//SesTab.ToCSVFile(thisdir+@"\table.csv");
// + dotnet_interactive={"language": "csharp"}
// Minitab blaupause 4 Runtime per iteration
var Minitab_blueprint = new System.Data.DataTable();
Minitab_blueprint.Columns.Add("NoOfCores", typeof(Int32));
Minitab_blueprint.Columns.Add("runtime_per_iter", typeof(Double));
Minitab_blueprint.Columns.Add("Speedup", typeof(Double));
// + dotnet_interactive={"language": "csharp"}
// fill minitables for plots
var listofDGdegree = new List<int>();
foreach(System.Data.DataRow Row in SesTab.Rows){
listofDGdegree.Add(Convert.ToInt32(Row["DGdegree:Velocity*"]));
}
List<double> DGdegrees=listofDGdegree.GroupBy(s=>s).Select(s=>s.Average()).ToList();
string dirpath = Directory.GetCurrentDirectory();
foreach(double DG in DGdegrees){
var MiniTab = Minitab_blueprint.Clone();
foreach(System.Data.DataRow Row in SesTab.Rows){
if(Convert.ToInt32(Row["DGdegree:Velocity*"])!=DG) continue;
var newrow = MiniTab.NewRow();
newrow["NoOfCores"] = Convert.ToInt32(Row["NoOfCores"]);
double iter = Convert.ToDouble(Row["LSolverIter"]);
double runtime = Convert.ToDouble(Row["SolverRun_time"]);
newrow["runtime_per_iter"] = runtime / iter;
MiniTab.Rows.Add(newrow);
}
System.Data.DataView view = new System.Data.DataView(MiniTab);
view.Sort = "NoOfCores";
MiniTab=view.ToTable();
double reference = Convert.ToDouble(MiniTab.Rows[0]["runtime_per_iter"]);
foreach(System.Data.DataRow Row in MiniTab.Rows){
Row["Speedup"]=reference/Convert.ToDouble(Row["runtime_per_iter"]);
}
MiniTab.ToCSVFile(dirpath+@"\strong_k"+DG+".dat");
} dirpath
// + dotnet_interactive={"language": "csharp"}
// Generate rnd Formatting for massive multiplots
var theDict = new List<Tuple<string,DashTypes,LineColors>>();
public static Tuple<string,DashTypes,LineColors>[] AofFormats;
var dashes = new DashTypes[]{DashTypes.Dashed,DashTypes.DotDashed,DashTypes.DotDotDashed};
var points= new PointTypes[]{PointTypes.OpenCircle,PointTypes.OpenCircle,PointTypes.OpenDiamond,PointTypes.OpenUpperTriangle};
var colors = new LineColors[]{LineColors.Red, LineColors.Green, LineColors.Blue, LineColors.Magenta, LineColors.Yellow, LineColors.Black, LineColors.Orange, LineColors.Grey};
int iSweep=0;
foreach(string Name in SesTab.GetColumnNames()){
//if(Name!="OrthonormalizationMultigrid.MinimizeResidual"&&Name!="overlap_scaling")
// continue;
if(Name=="NoOfCores"||Name=="LSolverIter"||Name=="DGdegree:Velocity*"||Name=="NoOfNodes"||Name=="MGDepth"||Name=="DOF")
continue;
int one=iSweep%(dashes.Length-1);
int two=iSweep/(dashes.Length-1);
theDict.Add(new Tuple<string,DashTypes,LineColors>(Name,dashes[one],colors[two]));
iSweep++;
}
AofFormats = theDict.ToArray()
// + dotnet_interactive={"language": "csharp"}
public static class DoWhatUWant{
public static PlotRowSelectorEx Solver_DG_Proj(int DG){
return delegate (int iSweep, int iTabRow,
IDictionary<string, object> Row,
out string Nmn,
out PlotFormat Fmt,
out double xValue, out double yValue) {
int k = Convert.ToInt32(Row["DGdegree:Velocity*"]);
Nmn = AofFormats[iSweep].Item1;
if(k != DG ) {
// degree does not match -> not in this plot
Nmn = null;
Fmt = null;
xValue = 0;
yValue = 0;
return;
}
Fmt = new PlotFormat();
Fmt.PointSize = 0.5;
Fmt.Style = Styles.LinesPoints;
Fmt.LineWidth = 3;
Fmt.DashType = AofFormats[iSweep].Item2;
Fmt.LineColor = AofFormats[iSweep].Item3;
double iter = Convert.ToDouble(Row["LSolverIter"]);
yValue = Convert.ToDouble(Row[Nmn])/iter;
//yValue = Convert.ToDouble(Row[Nmn]);
xValue = Convert.ToDouble(Row["NoOfCores"]);
};
}
public static PlotRowSelectorEx Solver_DG_Proj(){
return delegate (int iSweep, int iTabRow,
IDictionary<string, object> Row,
out string Nmn,
out PlotFormat Fmt,
out double xValue, out double yValue) {
int k = Convert.ToInt32(Row["DGdegree:Velocity*"]);
Fmt = new PlotFormat();
Fmt.PointSize = 0.5;
Fmt.Style = Styles.LinesPoints;
Fmt.LineWidth = 3;
switch(k){
case 2:
Nmn="k2";
Fmt.LineColor = LineColors.Red;
break;
case 3:
Nmn="k3";
Fmt.LineColor = LineColors.Orange;
break;
case 4:
Nmn="k4";
Fmt.LineColor = LineColors.Blue;
break;
default:
throw new Exception("vergiss es");
}
double time = Convert.ToDouble(Row["SolverRun_time"]);
double iter = Convert.ToDouble(Row["LSolverIter"]);
double timeSchwarz = Convert.ToDouble(Row["Schwarz.Solve"]);
//yValue = Convert.ToDouble(Row["NoOfNodes"]);
yValue = time/iter;
//yValue = Convert.ToDouble(Row["MGDepth"]);
//yValue=Convert.ToDouble(Row["LSolverIter"]);
//yValue = iter;
//yValue = time - timeSchwarz;
xValue = Convert.ToDouble(Row["NoOfCores"]);
};
}
}
// + dotnet_interactive={"language": "csharp"}
int k = 4;
int xMin = 8;
int xMax = 256;
// + dotnet_interactive={"language": "csharp"}
Plot2Ddata[,] multiplots = new Plot2Ddata[1,1];
//multiplots[0,0] = SesTab.ToPlot(DoWhatUWant.Solver_DG_Proj(),AofFormats.Length);
multiplots[0,0] = SesTab.ToPlot(DoWhatUWant.Solver_DG_Proj(k),AofFormats.Length);
multiplots[0,0].LogX = true;
multiplots[0,0].LogY = false;
multiplots[0,0].tmargin = 0;
multiplots[0,0].bmargin = 5;
multiplots[0,0].XrangeMin=xMin;
multiplots[0,0].XrangeMax=xMax;
multiplots[0,0].YrangeMin=0;
multiplots[0,0].YrangeMax=2;
multiplots[0,0].Ylabel = "iter";
multiplots[0,0].Xlabel = "cores";
multiplots[0,0].LegendAlignment=new string[]{"i","t","r"};
multiplots.PlotNow()
// + dotnet_interactive={"language": "csharp"}
// + dotnet_interactive={"language": "csharp"}
|
doc/handbook/apdx-MPISolverPerformance/strongScaling/IBM_RotSphere/Part2_0-scaling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 align="center">
# <a href="https://github.com/ml-tooling/ml-workspace" title="ML Workspace Home">
# <img width=50% alt="" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/ml-workspace-logo.png"></a>
# </h1>
#
# <div align="center">
# <strong>All-in-one web-based development environment for machine learning</strong>
# </div>
# <div align="center">
# <a href="#Features">Features & Screenshots</a> •
# <a href="#Support">Support</a> •
# <a href="https://github.com/ml-tooling/ml-workspace/issues/new?labels=bug&template=01_bug-report.md">Report a Bug</a> •
# <a href="https://github.com/ml-tooling/ml-workspace">Github Repo</a> •
# <a href="https://github.com/ml-tooling/ml-workspace#faq">FAQ</a> •
# <a href="https://github.com/ml-tooling/ml-workspace#known-issues">Known Issues</a> •
# <a href="#Next-Steps">Next Steps</a>
# </div>
#
# The ML workspace is an all-in-one web-based IDE specialized for machine learning and data science. It is simple to deploy and gets you started within minutes to productively built ML solutions on your own machines. This workspace is the ultimate tool for developers preloaded with a variety of popular data science libraries (e.g., Tensorflow, PyTorch, Keras, Sklearn) and dev tools (e.g., Jupyter, VS Code, Tensorboard) perfectly configured, optimized, and integrated.
#
# ## Highlights
#
# - 💫 Jupyter, JupyterLab, and Visual Studio Code web-based IDEs.
# - 🗃 Pre-installed with many popular data science libraries & tools.
# - 🖥 Full Linux desktop GUI accessible via web browser.
# - 🔀 Seamless Git integration optimized for notebooks.
# - 📈 Integrated hardware & training monitoring via Tensorboard & Netdata.
# - 🚪 Access from anywhere via Web, SSH, or VNC under a single port.
# - 🎛 Usable as remote kernel (Jupyter) or remote machine (VS Code) via SSH.
# - 🐳 Easy to deploy on Mac, Linux, and Windows via Docker.
# ## Connect to Lab
# The workspace has a python lab-client library pre-installed which provides easy access to the Lab API and capabilities to download/upload data, and run experiments. Below is a sneak peak into how you can use this library. To be able to execute this code, please replace `LAB_PROJECT` with one of your projects on the connected Lab instance and upload [**this dataset**](/docs/walkthrough/data/download-dataset.html) to the connected project from dataset section of the Lab UI.
#
# <div class="alert alert-info">
# To run this code you need to have created a project and uploaded the provided dataset file to the Lab instance!
# </div>
# +
from lab_client import Environment
# Initialize environment
env = Environment(project="LAB_PROJECT", # Lab project you want to work on
# Only required in stand-alone workspace deployments
# lab_endpoint="LAB_ENDPOINT",
# lab_api_token="LAB_TOKEN"
)
# Show environment information
env.print_info()
# -
# Run an experiment and sync experiment metadata to Lab:
# +
# Create experiment
exp = env.create_experiment('Welcome Tutorial')
# Get file (make sure that you have uploaded the dataset in the connected project)
text_corpus_path = env.get_file('datasets/news-categorized.csv')
# Define experiment
import random
def shuffle_lines(exp, params, artifacts):
# for this example we will just shuffle all lines in a file
random.seed(params["seed"])
lines = open(text_corpus_path).readlines()
random.shuffle(lines)
shuffeled_file_path = exp.create_file_path("shuffled_corpus.csv")
open(shuffeled_file_path, 'w').writelines(lines)
# log metadata
exp.log_metric("lines", len(lines))
# upload data
env.upload_file(shuffeled_file_path, "dataset")
# Define parameter configuration for experiment run
params = {
'seed': 1
}
# Run experiment and automatically sync all metadata with Lab
exp.run_exp(shuffle_lines, params)
# -
# More information about the lab-client library are available in the [**Lab Client Tutorial**](./tutorials/lab-client-tutorial.ipynb).
# ## /workspace Folder
# The default work directory is `/workspace` which is also the root directory of this Jupyter instance. Everything in that directory is persisted on the underlying host and backed up to the configured Lab instance every day.
#
# <div class="alert alert-warning">
# Use the <b>/workspace</b> directory for all your important work artifacts! Data within other directories might get lost.
# </div>
# ## Support
# The ML Workspace project is maintained by [<NAME>](https://twitter.com/LukasMasuch)
# and [<NAME>](https://twitter.com/raethlein). Please understand that we won't be able
# to provide individual support via email. We also believe that help is much more
# valuable if it's shared publicly so that more people can benefit from it.
#
#
# <table align="left" style="">
# <thead>
# <th style="text-align: left;">Type</th>
# <th style="text-align: left; min-width: 300px;">Channel</th>
# </thead>
# <tr>
# <td style="text-align: left;">🚨 Bug Reports</td>
# <td style="text-align: left;"><a href="https://github.com/ml-tooling/ml-workspace/issues?utf8=%E2%9C%93&q=is%3Aopen+is%3Aissue+label%3Abug+sort%3Areactions-%2B1-desc+" title="Open Bug Report"><img align="left" src="https://img.shields.io/github/issues/ml-tooling/ml-workspace/bug.svg"></a></td>
# </tr>
# <tr>
# <td style="text-align: left;">🎁 Feature Requests</td>
# <td style="text-align: left;"><a href="https://github.com/ml-tooling/ml-workspace/issues?q=is%3Aopen+is%3Aissue+label%3Afeature-request+sort%3Areactions-%2B1-desc" title="Open Feature Request"><img align="left" src="https://img.shields.io/github/issues/ml-tooling/ml-workspace/feature-request.svg?label=feature%20requests"></a> </td>
# </tr>
# <tr>
# <td style="text-align: left;">👩💻 Usage Questions</td>
# <td style="text-align: left;"><a href="https://stackoverflow.com/questions/tagged/ml-tooling" title="Open Question on Stackoverflow"><img align="left" src="https://img.shields.io/badge/stackoverflow-ml--tooling-orange.svg"></a> <a href="https://gitter.im/ml-tooling/ml-workspace" title="Chat on Gitter"><img align="left" style="margin-left: 5px;" src="https://badges.gitter.im/ml-tooling/ml-workspace.svg"></a></td>
# </tr>
# <tr>
# <td style="text-align: left;">🗯 General Discussion</td>
# <td style="text-align: left;"><div align="left"><a href="https://gitter.im/ml-tooling/ml-workspace" title="Chat on Gitter"><img align="left" src="https://badges.gitter.im/ml-tooling/ml-workspace.svg"></a><a href="https://twitter.com/mltooling" title="ML Tooling on Twitter"><img align="left" style="margin-left: 5px;" src="https://img.shields.io/twitter/follow/mltooling.svg?style=social"></a></div></td>
# </tr>
# </table>
# ## Features
# <div align="center">
# <a href="#Jupyter">Jupyter</a> •
# <a href="#Desktop-GUI">Desktop GUI</a> •
# <a href="#Visual-Studio-Code">VS Code</a> •
# <a href="#JupyterLab">JupyterLab</a> •
# <a href="#Git-Integration">Git Integration</a> •
# <a href="#File-Sharing">File Sharing</a> •
# <a href="#Access-Ports">Access Ports</a> •
# <a href="#Tensorboard">Tensorboard</a> •
# <a href="#Extensibility">Extensibility</a> •
# <a href="#Hardware-Monitoring">Hardware Monitoring</a> •
# <a href="#SSH-Access">SSH Access</a> •
# <a href="#Remote-Development">Remote Development</a> •
# <a href="#Run-as-a-job">Job Execution</a>
# </div>
#
# The workspace is equipped with a selection of best-in-class open-source development tools to help with the machine learning workflow. Many of these tools can be started from the `Open Tool` menu from Jupyter (the main application of the workspace):
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/open-tools.png"/>
#
# ### Install Anything
#
# Within your workspace you have **full root & sudo privileges** to install any library or tool you need via terminal (e.g., `pip`, `apt-get`, `conda`, or `npm`). You can find more ways to extend the workspace within the [Extensibility](#Extensibility) section.
# !pip install matplotlib-venn
# <div class="alert alert-success">
# <b>Install Dependencies in Notebooks:</b> It’s a good idea to include cells which install and load any custom libraries or files (which are not pre-installed in the workspace) that your notebook needs.
# </div>
# ### Jupyter
#
# [Jupyter Notebook](https://jupyter.org/) is a web-based interactive environment for writing and running code. The main building blocks of Jupyter are the file-browser, the notebook editor, and kernels. The file-browser provides an interactive file manager for all notebooks, files, and folders in the `/workspace` directory.
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/jupyter-tree.png"/>
#
# A new notebook can be created by clicking on the `New` drop-down button at the top of the list and selecting the desired language kernel.
#
# <div class="alert alert-info">
# You can spawn interactive <b>terminal</b> instances as well by selecting <code>New -> Terminal</code> in the file-browser.
# </div>
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/jupyter-notebook.png"/>
#
# The notebook editor enables users to author documents that include live code, markdown text, shell commands, LaTeX equations, interactive widgets, plots, and images. These notebook documents provide a complete and self-contained record of a computation that can be converted to various formats and shared with others.
#
# <div class="alert alert-info">
# This workspace has a variety of <b>third-party Jupyter extensions</b> activated. You can configure these extensions in the nbextensions configurator: <b>nbextensions</b> tab on the file browser
# </div>
#
# The Notebook allows code to be run in a range of different programming languages. For each notebook document that a user opens, the web application starts a **kernel** that runs the code for that notebook and returns output. This workspace has a Python 3 and Python 2 kernel pre-installed. Additional Kernels can be installed to get access to other languages (e.g., R, Scala, Go) or additional computing resources (e.g., GPUs, CPUs, Memory).
#
# <div class="alert alert-info">
# <b>Python 2</b> support is deprecated and not fully supported. Please only use Python 2 if necessary!
# </div>
#
# ### Desktop GUI
#
# This workspace provides an HTTP-based VNC access to the workspace via [noVNC](https://github.com/novnc/noVNC). Thereby, you can access and work within the workspace with a fully-featured desktop GUI. To access this desktop GUI, go to `Open Tool`, select `VNC`, and click the `Connect` button. In the case you are asked for a password, use `<PASSWORD>`.
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/desktop-vnc.png"/>
#
# Once you are connected, you will see a desktop GUI that allows you to install and use full-fledged web-browsers or any other tool that is available for Ubuntu. Within the `Tools` folder on the desktop, you will find a collection of install scripts that makes it straightforward to install some of the most commonly used development tools, such as Atom, PyCharm, R-Runtime, R-Studio, or Postman (just double-click on the script).
#
# **Clipboard:** If you want to share the clipboard between your machine and the workspace, you can use the copy-paste functionality as described below:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/desktop-vnc-clipboard.png"/>
#
# <div class="alert alert-success">
# <b>Long-running tasks:</b> Use the desktop GUI for long-running Jupyter executions. By running notebooks from the browser of your workspace desktop GUI, all output will be synchronized to the notebook even if you have disconnected your browser from the notebook.
# </div>
#
# ### Visual Studio Code
#
# [Visual Studio Code](https://github.com/microsoft/vscode) (`Open Tool -> VS Code`) is an open-source lightweight but powerful code editor with built-in support for a variety of languages and a rich ecosystem of extensions. It combines the simplicity of a source code editor with powerful developer tooling, like IntelliSense code completion and debugging. The workspace integrates VS Code as a web-based application accessible through the browser-based on the awesome [code-server](https://github.com/cdr/code-server) project. It allows you to customize every feature to your liking and install any number of third-party extensions.
#
# <p align="center"><img src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/vs-code.png"/></p>
#
# The workspace also provides a VS Code integration into Jupyter allowing you to open a VS Code instance for any selected folder, as shown below:
#
# <p align="center"><img src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/vs-code-open.png"/></p>
#
# ### JupyterLab
#
# [JupyterLab](https://github.com/jupyterlab/jupyterlab) (`Open Tool -> JupyterLab`) is the next-generation user interface for Project Jupyter. It offers all the familiar building blocks of the classic Jupyter Notebook (notebook, terminal, text editor, file browser, rich outputs, etc.) in a flexible and powerful user interface. This JupyterLab instance comes pre-installed with a few helpful extensions such as a the [jupyterlab-toc](https://github.com/jupyterlab/jupyterlab-toc), [jupyterlab-git](https://github.com/jupyterlab/jupyterlab-git), and [juptyterlab-tensorboard](https://github.com/chaoleili/jupyterlab_tensorboard).
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/jupyterlab.png"/>
#
# ### Git Integration
#
# Version control is a crucial aspect of productive collaboration. To make this process as smooth as possible, we have integrated a custom-made Jupyter extension specialized on pushing single notebooks, a full-fledged web-based Git client ([ungit](https://github.com/FredrikNoren/ungit)), a tool to open and edit plain text documents (e.g., `.py`, `.md`) as notebooks ([jupytext](https://github.com/mwouts/jupytext)), as well as a notebook merging tool ([nbdime](https://github.com/jupyter/nbdime)). Additionally, JupyterLab and VS Code also provide GUI-based Git clients.
#
# #### Clone Repository
#
# For cloning repositories via `https`, we recommend to navigate to the desired root folder and to click on the `git` button as shown below:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/git-open.png"/>
#
# This might ask for some required settings and, subsequently, opens [ungit](https://github.com/FredrikNoren/ungit), a web-based Git client with a clean and intuitive UI that makes it convenient to sync your code artifacts. Within ungit, you can clone any repository. If authentication is required, you will get asked for your credentials.
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/git-ungit-credentials.png"/>
#
# #### Push, Pull, Merge, and Other Git Actions
#
# To commit and push a single notebook to a remote Git repository, we recommend to use the Git plugin integrated into Jupyter, as shown below:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/git-push-notebook.png"/>
#
# For more advanced Git operations, we recommend to use [ungit](https://github.com/FredrikNoren/ungit). With ungit, you can do most of the common git actions such as push, pull, merge, branch, tag, checkout, and many more.
#
# #### Diffing and Merging Notebooks
#
# Jupyter notebooks are great, but they often are huge files, with a very specific JSON file format. To enable seamless diffing and merging via Git this workspace is pre-installed with [nbdime](https://github.com/jupyter/nbdime). Nbdime understands the structure of notebook documents and, therefore, automatically makes intelligent decisions when diffing and merging notebooks. In the case you have merge conflicts, nbdime will make sure that the notebook is still readable by Jupyter, as shown below:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/git-nbdime-merging.png"/>
#
# Furthermore, the workspace comes pre-installed with [jupytext](https://github.com/mwouts/jupytext), a Jupyter plugin that reads and writes notebooks as plain text files. This allows you to open, edit, and run scripts or markdown files (e.g., `.py`, `.md`) as notebooks within Jupyter. In the following screenshot, we have opened a markdown file via Jupyter:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/git-jupytext.png"/>
#
# In combination with Git, jupytext enables a clear diff history and easy merging of version conflicts. With both of those tools, collaborating on Jupyter notebooks with Git becomes straightforward.
#
# ### File Sharing
#
# The workspace has a feature to share any file or folder with anyone via a token-protected link. To share data via a link, select any file or folder from the Jupyter directory tree and click on the share button as shown in the following screenshot:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/file-sharing-open.png"/>
#
# This will generate a unique link protected via a token that gives anyone with the link access to view and download the selected data via the [Filebrowser](https://github.com/filebrowser/filebrowser) UI:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/file-sharing-filebrowser.png"/>
#
# To deactivate or manage (e.g., provide edit permissions) shared links, open the Filebrowser via `Open Tool -> Filebrowser` and select `Settings->User Management`.
#
# ### Access Ports
#
# It is possible to securely access any workspace internal port by selecting `Open Tool -> Access Port`. With this feature, you are able to access a REST API or web application running inside the workspace directly with your browser. The feature enables developers to build, run, test, and debug REST APIs or web applications directly from the workspace.
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/access-port.png"/>
#
# If you want to use an HTTP client or share access to a given port, you can select the `Get shareable link` option. This generates a token-secured link that anyone with access to the link can use to access the specified port.
#
# <div class="alert alert-info">
# The HTTP app requires to be resolved from a relative URL path or configure a base path (<code>/tools/PORT/</code>).
# </div>
#
# <details>
#
# <summary>Example (click to expand...)</summary>
#
# 1. Start an HTTP server on port `1234` by running this command in a terminal within the workspace: `python -m http.server 1234`
# 2. Select `Open Tool -> Access Port`, input port `1234`, and select the `Get shareable link` option.
# 3. Click `Access`, and you will see the content provided by Python's `http.server`.
# 4. The opened link can also be shared to other people or called from external applications (e.g., try with Incognito Mode in Chrome).
#
# </details>
#
# ### SSH Access
#
# SSH provides a powerful set of features that enables you to be more productive with your development tasks. You can easily set up a secure and passwordless SSH connection to a workspace by selecting `Open Tool -> SSH`. This will generate a secure setup command that can be run on any Linux or Mac machine to configure a passwordless & secure SSH connection to the workspace. Alternatively, you can also download the setup script and run it (instead of using the command).
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/ssh-access.png"/>
#
# <div class="alert alert-info">
# The setup script only runs on Mac and Linux. Windows is currently not supported.
# </div>
#
# Just run the setup command or script on the machine from where you want to setup a connection to the workspace and input a name for the connection (e.g., `my-workspace`). You might also get asked for some additional input during the process, e.g. to install a remote kernel if `remote_ikernel` is installed. Once the passwordless SSH connection is successfully setup and tested, you can securely connect to the workspace by simply executing `ssh my-workspace`.
#
# Besides the ability to execute commands on a remote machine, SSH also provides a variety of other features that can improve your development workflow as described in the following sections.
#
# <details>
# <summary><b>Tunnel Ports</b> (click to expand...)</summary>
#
# An SSH connection can be used for tunneling application ports from the remote machine to the local machine, or vice versa. For example, you can expose the workspace internal port `5901` (VNC Server) to the local machine on port `5000` by executing:
#
# ```bash
# ssh -nNT -L 5000:localhost:5901 my-workspace
# ```
#
# <div class="alert alert-info">
# To expose an application port from your local machine to a workspace, use the <code>-R</code> option (instead of <code>-L</code>).
# </div>
#
# After the tunnel is established, you can use your favorite VNC viewer on your local machine and connect to `vnc://localhost:5000` (default password: `<PASSWORD>`). To make the tunnel connection more resistant and reliable, we recommend to use [autossh](https://www.harding.motd.ca/autossh/) to automatically restart SSH tunnels in the case that the connection dies:
#
# ```bash
# autossh -M 0 -f -nNT -L 5000:localhost:5901 my-workspace
# ```
#
# Port tunneling is quite useful when you have started any server-based tool within the workspace that you like to make accessible for another machine. In its default setting, the workspace has a variety of tools already running on different ports, such as:
#
# - `8080`: Main workspace port with access to all integrated tools.
# - `8090`: Jupyter server.
# - `8054`: VS Code server.
# - `5901`: VNC server.
# - `3389`: RDP server.
# - `22`: SSH server.
#
# You can find port information on all the tools in the [supervisor configuration](https://github.com/ml-tooling/ml-workspace/blob/master/resources/config/supervisord.conf).
#
# <div class="alert alert-info">
# 📖 For more information about port tunneling/forwarding, we recommend <a href="https://www.everythingcli.org/ssh-tunnelling-for-fun-and-profit-local-vs-remote">this guide</a>.
# </div>
# <br>
# </details>
#
# <details>
# <summary><b>Copy Data via SCP</b> (click to expand...)</summary>
#
# [SCP](https://linux.die.net/man/1/scp) allows files and directories to be securely copied to, from, or between different machines via SSH connections. For example, to copy a local file (`./local-file.txt`) into the `/workspace` folder inside the workspace, execute:
#
# ```bash
# scp ./local-file.txt my-workspace:/workspace
# ```
#
# To copy the `/workspace` directory from `my-workspace` to the working directory of the local machine, execute:
#
# ```bash
# scp -r my-workspace:/workspace .
# ```
#
# <div class="alert alert-info">
# 📖 For more information about scp, we recommend <a href="https://www.garron.me/en/articles/scp.html">this guide</a>.
# </div>
# <br>
# </details>
#
# <details>
# <summary><b>Sync Data via Rsync</b> (click to expand...)</summary>
#
# [Rsync](https://linux.die.net/man/1/rsync) is a utility for efficiently transferring and synchronizing files between different machines (e.g., via SSH connections) by comparing the modification times and sizes of files. The rsync command will determine which files need to be updated each time it is run, which is far more efficient and convenient than using something like scp or sftp. For example, to sync all content of a local folder (`./local-project-folder/`) into the `/workspace/remote-project-folder/` folder inside the workspace, execute:
#
# ```bash
# rsync -rlptzvP --delete --exclude=".git" "./local-project-folder/" "my-workspace:/workspace/remote-project-folder/"
# ```
#
# If you have some changes inside the folder on the workspace, you can sync those changes back to the local folder by changing the source and destination arguments:
#
# ```bash
# rsync -rlptzvP --delete --exclude=".git" "my-workspace:/workspace/remote-project-folder/" "./local-project-folder/"
# ```
#
# You can rerun these commands each time you want to synchronize the latest copy of your files. Rsync will make sure that only updates will be transferred.
#
# <div class="alert alert-info">
# 📖 You can find more information about rsync on <a href="https://linux.die.net/man/1/rsync">this man page</a>.
# </div>
# <br>
# </details>
#
# <details>
# <summary><b>Mount Folders via SSHFS</b> (click to expand...)</summary>
#
# Besides copying and syncing data, an SSH connection can also be used to mount directories from a remote machine into the local filesystem via [SSHFS](https://github.com/libfuse/sshfs).
# For example, to mount the `/workspace` directory of `my-workspace` into a local path (e.g. `/local/folder/path`), execute:
#
# ```bash
# sshfs -o reconnect my-workspace:/workspace /local/folder/path
# ```
#
# Once the remote directory is mounted, you can interact with the remote file system the same way as with any local directory and file.
#
# <div class="alert alert-info">
# 📖 For more information about sshfs, we recommend <a href="https://www.digitalocean.com/community/tutorials/how-to-use-sshfs-to-mount-remote-file-systems-over-ssh">this guide</a>.
# </div>
# <br>
# </details>
#
# ### Remote Development
#
# The workspace can be integrated and used as a remote runtime (also known as remote kernel/machine/interpreter) for a variety of popular development tools and IDEs, such as Jupyter, VS Code, PyCharm, Colab, or Atom Hydrogen. Thereby, you can connect your favorite development tool running on your local machine to a remote machine for code execution. This enables a **local-quality development experience with remote-hosted compute resources**.
#
# These integrations usually require a passwordless SSH connection from the local machine to the workspace. To set up an SSH connection, please follow the steps explained in the [SSH Access](#ssh-access) section.
#
# <details>
# <summary><b>Jupyter - Remote Kernel</b> (click to expand...)</summary>
#
# The workspace can be added to a Jupyter instance as a remote kernel by using the [remote_ikernel](https://bitbucket.org/tdaff/remote_ikernel/) tool. If you have installed remote_ikernel (`pip install remote_ikernel`) on your local machine, the SSH setup script of the workspace will automatically offer you the option to setup a remote kernel connection.
#
# <div class="alert alert-info">
# When running kernels on remote machines, the notebooks themselves will be saved onto the local filesystem, but the kernel will only have access to the filesystem of the remote machine running the kernel. If you need to sync data, you can make use of rsync, scp, or sshfs as explained in the <a href="#SSH-Access">SSH Access</a> section.
# </div>
#
# In case you want to manually setup and manage remote kernels, use the [remote_ikernel](https://bitbucket.org/tdaff/remote_ikernel/src/default/README.rst) command-line tool, as shown below:
#
# ```bash
# # Change my-workspace with the name of a workspace SSH connection
# remote_ikernel manage --add \
# --interface=ssh \
# --kernel_cmd="ipython kernel -f {connection_file}" \
# --name="ml-server Py 3.6" \
# --host="my-workspace"
# ```
#
# You can use the remote_ikernel command line functionality to list (`remote_ikernel manage --show`) or delete (`remote_ikernel manage --delete <REMOTE_KERNEL_NAME>`) remote kernel connections.
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/remote-dev-jupyter-kernel.png"/>
#
# </details>
#
# <details>
# <summary><b>VS Code - Remote Machine</b> (click to expand...)</summary>
#
# The Visual Studio Code [Remote - SSH](https://marketplace.visualstudio.com/items?itemName=ms-vscode-remote.remote-ssh) extension allows you to open a remote folder on any remote machine with SSH access and work with it just as you would if the folder were on your own machine. Once connected to a remote machine, you can interact with files and folders anywhere on the remote filesystem and take full advantage of VS Code's feature set (IntelliSense, debugging, and extension support). The discovers and works out-of-the-box with passwordless SSH connections as configured by the workspace SSH setup script. To enable your local VS Code application to connect to a workspace:
#
# 1. Install [Remote - SSH](https://marketplace.visualstudio.com/items?itemName=ms-vscode-remote.remote-ssh) extension inside your local VS Code.
# 2. Run the SSH setup script of a selected workspace as explained in the [SSH Access](#ssh-access) section.
# 3. Open the Remote-SSH panel in your local VS Code. All configured SSH connections should be automatically discovered. Just select any configured workspace connection you like to connect to as shown below:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/remote-dev-vscode.gif"/>
#
# <div class="alert alert-info">
# 📖 You can find additional features and information about the Remote SSH extension in <a href="https://code.visualstudio.com/docs/remote/ssh">this guide</a>.
# </div>
# <br>
# </details>
#
# ### Tensorboard
#
# [Tensorboard](https://www.tensorflow.org/tensorboard) provides a suite of visualization tools to make it easier to understand, debug, and optimize your experiment runs. It includes logging features for scalar, histogram, model structure, embeddings, and text & image visualization. The workspace comes pre-installed with [jupyter_tensorboard extension](https://github.com/lspvic/jupyter_tensorboard) that integrates Tensorboard into the Jupyter interface with functionalities to start, manage, and stop instances. You can open a new instance for a valid logs directory, as shown below:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/tensorboard-open.png" />
#
# If you have opened a Tensorboard instance in a valid log directory, you will see the visualizations of your logged data:
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/tensorboard-dashboard.png" />
#
# <div class="alert alert-info">
# Tensorboard can be used in combination with many other ML frameworks besides Tensorflow. By using the <a href="https://github.com/lanpa/tensorboardX">tensorboardX</a> library you can log basically from any python based library. Also, PyTorch has a direct Tensorboard integration as described <a href="https://pytorch.org/docs/stable/tensorboard.html">here</a>.
# </div>
#
# If you prefer to see the tensorboard directly within your notebook, you can make use of following **Jupyter magic**:
# %load_ext tensorboard.notebook
# %tensorboard --logdir /workspace/path/to/logs
# <div class="alert alert-success">
# <b>Experiment Tracking Integration:</b> The experiment tracking from the lab client library has tensorboard logging capabilities integrated for any machine learning framework (see the Environment Tutorial).
# </div>
# ### Hardware Monitoring
#
# The workspace provides two pre-installed web-based tools to help developers during model training and other experimentation tasks to get insights into everything happening on the system and figure out performance bottlenecks.
#
# [Netdata](https://github.com/netdata/netdata) (`Open Tool -> Netdata`) is a real-time hardware and performance monitoring dashboard that visualize the processes and services on your Linux systems. It monitors metrics about CPU, GPU, memory, disks, networks, processes, and more.
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/hardware-monitoring-netdata.png" />
#
# [Glances](https://github.com/nicolargo/glances) (`Open Tool -> Glances`) is a web-based hardware monitoring dashboard as well and can be used as an alternative to Netdata.
#
# <img style="width: 100%" src="https://github.com/ml-tooling/ml-workspace/raw/master/docs/images/features/hardware-monitoring-glances.png"/>
#
# <div class="alert alert-info">
# Netdata and Glances will show you the hardware statistics for the entire machine on which the workspace container is running.
# </div>
#
# ### Run as a job
#
# <div class="alert alert-info">
# A job is defined as any computational task that runs for a certain time to completion, such as a model training or a data pipeline.
# </div>
#
# The workspace image can also be used to execute arbitrary Python code without starting any of the pre-installed tools. This provides a seamless way to productize your ML projects since the code that has been developed interactively within the workspace will have the same environment and configuration when run as a job via the same workspace image.
#
# <details>
# <summary><b>Run Python code as a job via the workspace image</b> (click to expand...)</summary>
#
# To run Python code as a job, you need to provide a path or URL to a code directory (or script) via `EXECUTE_CODE`. The code can be either already mounted into the workspace container or downloaded from a version control system (e.g., git or svn) as described in the following sections. The selected code path needs to be python executable. In case the selected code is a directory (e.g., whenever you download the code from a VCS) you need to put a `__main__.py` file at the root of this directory. The `__main__.py` needs to contain the code that starts your job.
#
# #### Run code from version control system
#
# You can execute code directly from Git, Mercurial, Subversion, or Bazaar by using the pip-vcs format as described in [this guide](https://pip.pypa.io/en/stable/reference/pip_install/#vcs-support). For example, to execute code from a [subdirectory](https://github.com/ml-tooling/ml-workspace/tree/master/resources/tests/ml-job) of a git repository, just run:
#
# ```bash
# docker run --env EXECUTE_CODE="git+https://github.com/ml-tooling/ml-workspace.git#subdirectory=resources/tests/ml-job" mltooling/ml-workspace:latest
# ```
#
# <div class="alert alert-info">
# 📖 For additional information on how to specify branches, commits, or tags please refer to <a href="https://pip.pypa.io/en/stable/reference/pip_install/#vcs-support">this guide</a>.
# </div>
#
# #### Run code mounted into the workspace
#
# In the following example, we mount and execute the current working directory (expected to contain our code) into the `/workspace/ml-job/` directory of the workspace:
#
# ```bash
# docker run -v "${PWD}:/workspace/ml-job/" --env EXECUTE_CODE="/workspace/ml-job/" mltooling/ml-workspace:latest
# ```
#
# #### Install Dependencies
#
# In the case that the pre-installed workspace libraries are not compatible with your code, you can install or change dependencies by just adding one or multiple of the following files to your code directory:
#
# - `requirements.txt`: [pip requirements format](https://pip.pypa.io/en/stable/user_guide/#requirements-files) for pip-installable dependencies.
# - `environment.yml`: [conda environment file](https://docs.conda.io/projects/conda/en/latest/user-guide/tasks/manage-environments.html?highlight=environment.yml#creating-an-environment-file-manually) to create a separate Python environment.
# - `setup.sh`: A shell script executed via `/bin/bash`.
#
# The execution order is 1. `environment.yml` -> 2. `setup.sh` -> 3. `requirements.txt`
#
# #### Test job in interactive mode
#
# You can test your job code within the workspace (started normally with interactive tools) by executing the following python script:
#
# ```bash
# python /resources/scripts/execute_code.py /path/to/your/job
# ```
#
# #### Build a custom job image
#
# It is also possible to embed your code directly into a custom job image, as shown below:
#
# ```dockerfile
# FROM mltooling/ml-workspace:latest
#
# # Add job code to image
# COPY ml-job /workspace/ml-job
# ENV EXECUTE_CODE=/workspace/ml-job
#
# # Install requirements only
# RUN python /resources/scripts/execute_code.py --requirements-only
#
# # Execute only the code at container startup
# CMD ["python", "/resources/docker-entrypoint.py", "--code-only"]
# ```
#
# </details>
#
# ### Pre-installed Libraries and Interpreters
#
# The workspace is pre-installed with many popular interpreters, data science libraries, and ubuntu packages:
#
# - **Interpreter:** Miniconda 3 (Python 3.6), Java 8, NodeJS 11
# - **Python libraries:** Tensorflow, Keras, Pytorch, Sklearn, CNTK, XGBoost, Theano, Fastai, and [many more](https://github.com/ml-tooling/ml-workspace/tree/master/resources/libraries)
#
# The full list of installed tools can be found within the [Dockerfile](https://github.com/ml-tooling/ml-workspace/blob/master/Dockerfile).
#
# <div class="alert alert-info">
# For every minor version release, we run vulnerability, virus, and security checks within the workspace using <a href="https://vuls.io/">vuls</a>, <a href="https://pyup.io/safety/">safety</a>, and <a href="https://www.clamav.net/">clamav</a> to make sure that the workspace environment is as secure as possible.
# </div>
#
# ### Extensibility
#
# The workspace provides a high degree of extensibility. Within the workspace, you have **full root & sudo privileges** to install any library or tool you need via terminal (e.g., `pip`, `apt-get`, `conda`, or `npm`). You can open a terminal by one of the following ways:
#
# - **Jupyter:** `New -> Terminal`
# - **Desktop VNC:** `Applications -> Terminal Emulator`
# - **JupyterLab:** `File -> New -> Terminal`
# - **VS Code:** `Terminal -> New Terminal`
#
# Additionally, pre-installed tools such as Jupyter, JupyterLab, and Visual Studio Code each provide their own rich ecosystem of extensions. The workspace also contains a [collection of installer scripts](../tree/tools) for many commonly used development tools or libraries (e.g., `PyCharm`, `Zeppelin`, `RStudio`, `Starspace`). Those scripts can be either executed from the Desktop VNC (double-click on the script within the `Tools` folder on the Desktop) or from a terminal (execute any tool script from the `/resources/tools/` folder).
#
# <details>
# <summary>Example (click to expand...)</summary>
#
# For example, to install the [Apache Zeppelin](https://zeppelin.apache.org/) notebook server, simply execute:
#
# ```bash
# /resources/tools/zeppelin.sh --port=1234
# ```
#
# After installation, refresh the Jupyter website and the Zeppelin tool will be available under `Open Tool -> Zeppelin`. Other tools might only be available within the Desktop VNC (e.g., `atom` or `pycharm`) or do not provide any UI (e.g., `starspace`, `docker-client`).
# </details>
#
# As an alternative to extending the workspace at runtime, you can also customize the workspace Docker image to create your own flavor as explained in the [FAQ](https://github.com/ml-tooling/ml-workspace#faq) section.
#
# ### Deploy Locally
#
# If you want to deploy the workspace stand-alone on your computer, just execute the following command from within a directory which you like to mount into the `/workspace` directory of the workspace container:
#
#
# ``` bash
# docker run -d -v $(PWD):/workspace -p 8080:8080 mltooling/ml-workspace:latest
# ```
# ## Next Steps
# It is recommended to go through the tutorials in the given order:
# - [Lab Client Tutorial](./tutorials/lab-client-tutorial.ipynb): Learn how to connect to ML Lab and run your experiments.
# - [Unified Model Tutorial](./tutorials/unified-model-tutorial.ipynb): Package your model logic, requirements, and artifacts into a single self-contained & executable file.
# - [Visualization in Jupyter](./tutorials/visualization-tutorial.ipynb): Introduction to data visualization with various common charting tools.
# - [Experiment Template](./templates/experiment-template.ipynb): Start your own high-quality reusable experiment notebook with this template.
|
services/lab-workspace/docker-res/tutorials/welcome.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Deploy Object Detection Model Use ModelCI
#
# MMDetction is a well-known open source object detection toolbox based on PyTorch. You can refer to <https://arxiv.org/abs/1906.07155> for more details.
#
# By walking through this tutorial, you will be able to:
#
# - Load pretained MMDetction model
# - Convert MMDetction model into ONNX format
# - Register and retrieve models by ModelHub
#
# ## 1. Prequisities
#
# ### 1.1 Installation of MMDetction
#
# Firstly you have to install MMDetction according to official instructions : <https://mmdetection.readthedocs.io/en/latest/get_started.html#installation>
# + tags=[]
# !pip install https://download.openmmlab.com/mmcv/dist/latest/torch1.7.0/cu102/mmcv_full-latest%2Btorch1.7.0%2Bcu102-cp37-cp37m-manylinux1_x86_64.whl
# !git clone https://github.com/open-mmlab/mmdetection.git
# !cd mmdetection && pip install -q -r requirements/build.txt && pip install -q -v -e .
# -
# ### 1.2 Start ModelCI Service
# Then we can start our ModelCI service, you should at least set mongodb password before starting. You can refer to [last notebook](https://github.com/cap-ntu/ML-Model-CI/blob/master/example/notebook/image_classification_model_deployment.ipynb) for more details.
# %env MONGO_PASSWORD=<PASSWORD>
# !conda activate modelci && modelci start
# ## 2. Build MMdetection Model
# ### 2.1 Imports
# We should import the following functions:
# - preprocess_example_input: for generating tensor and meta info from example image file
# - build_model_from_cfg: for building model form config file and checkpoint file
from mmdet.core import preprocess_example_input, build_model_from_cfg
# ### 2.2 Model Config
#
# We should either use a dict or config file for configuration of MMDetection model, to make things simple, we use a config file provided by MMDetection.
#
# Notice:
#
# - You may need to manually download pretrained model checkpoints from [MMDetection models zoo](https://github.com/open-mmlab/mmdetection/blob/master/docs/model_zoo.md).
# - Only a few MMdet models are able to converted into ONNX format, you can refer to [documentation](https://mmdetection.readthedocs.io/en/latest/tutorials/pytorch2onnx.html#list-of-supported-models-exportable-to-onnx) for more detail.
config_file = 'mmdetection/configs/retinanet/retinanet_r50_fpn_1x_coco.py'
checkpoint_file = 'retinanet_r50_fpn_1x_coco_20200130-c2398f9e.pth'
# ### 2.3 Build Model
# Then we can build our MMdetection model based on the configuration above and the checkpoint file we already download.
model = build_model_from_cfg(config_file, checkpoint_file)
# Before conversion, we need to modify forward function to provide the necessary **kwargs parameters such as img_metas.
#
# In order to obtain valid bbox data during the onnx tracing process, we also need to use a tensor generated from image file as model input instead of random tensors.
input_config = {
'input_shape': (1,3,224,224),
'input_path': 'mmdetection/demo/demo.jpg',
'normalize_cfg': {
'mean': (123.675, 116.28, 103.53),
'std': (58.395, 57.12, 57.375)
}
}
one_img, one_meta = preprocess_example_input(input_config)
one_img.shape
from functools import partial
model.forward = partial(model.forward, img_metas=[[one_meta]], return_loss=False)
# ## 3. Register Model
# We can convert the pytorch model above into optimized formats, such as ONNX through modelci
#
# ### 3.1 Imports
# - modelci.hub.manager: for registering model into ModelHub
# - modelci.types.bo: for constructing model inputs paramenters
# - modelci.types.trtis_objects: for specifying model input shape format
from modelci.hub.manager import register_model
from modelci.types.bo import IOShape, Framework, Task, Engine, Metric, ModelVersion
from modelci.types.trtis_objects import ModelInputFormat
# ### 3.2 Sepcify Input and Output
#
# Here are some parameters need to be specified before model conversion.
# - inputs: The model inputs info
# - outputs: The model outputs info
#
inputs = [IOShape([-1, 3, 204, 204], dtype=float, name='IMAGE', format=ModelInputFormat.FORMAT_NCHW)]
outputs = [
IOShape([-1, 100, 5], dtype=float, name='BBOX'),
IOShape([-1, 100], dtype=float, name='SCORE')
]
# ### 3.3 Register
#
# In this step, the inputs and outputs variable generated before are reused.
register_model(
model,
dataset='COCO',
task=Task.OBJECT_DETECTION,
metric={Metric.MAP: 0.365},
outputs=outputs,
inputs=inputs,
architecture='RetinaNet',
framework=Framework.PYTORCH,
version=ModelVersion('1'),
model_input=[one_img],
profile=False
)
# As we could see, MLModelCI support auto conversion of PyTorch models into both torchscript and ONNX format, as a result.
#
# However, this model cannot be transformed into torchscript format, but supportive of ONNX format conversion, there could be serveral factors contributing to model conversion failture such as the model structure and code format.
# ## 4. Retrieve Model
#
# The following steps will retrieve the model we just registered.
from modelci.hub.manager import retrieve_model
retrieved_models = retrieve_model(
architecture_name = 'RetinaNet',
framework = Framework.PYTORCH,
version=ModelVersion('1')
)
retrieved_models
# It's no wonder we get two model objects here cause there is an addition ONNX format model created automatically during previous registering process.
retrieved_models[0].__dict__
retrieved_models[1].__dict__
|
example/notebook/object_detection_model_deployment.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Understanding RNN structure
# - Distinguished from feedforward nets, RNNs are structures that can well handle data with "sequential" format by preserving previous "state"
# - Thus, grasping concepts of **"sequences"** and (hidden) **"states"** in RNNs is crucial
#
# <br>
# <img src="http://karpathy.github.io/assets/rnn/charseq.jpeg" style="width: 500px"/>
import numpy as np
from keras.models import Model, Sequential
from keras.layers import *
# ## 1. SimpleRNN
# Input shape of SimpleRNN should be 3D tensor => (batch_size, timesteps, input_dim)
# - **batch_size**: ommitted when creating RNN instance (== None). Usually designated when fitting model.
# - **timesteps**: number of input sequence per batch
# - **input_dim**: dimensionality of input sequence
# for instance, consider below array
x = np.array([[
[1, # => input_dim 1
2, # => input_dim 2
3], # => input_dim 3 # => timestep 1
[4, 5, 6] # => timestep 2
], # => batch 1
[[7, 8, 9], [10, 11, 12]], # => batch 2
[[13, 14, 15], [16, 17, 18]] # => batch 3
])
print('(Batch size, timesteps, input_dim) = ',x.shape)
# rnn = SimpleRNN(50)(Input(shape = (10,))) => error
# rnn = SimpleRNN(50)(Input(shape = (10, 30, 40))) => error
rnn = SimpleRNN(50)(Input(shape = (10, 30)))
# **return_state** = **return_sequences** = **False** ====> output_shape = **(batch_size = None, num_units)**
rnn = SimpleRNN(50)(Input(shape = (10, 30)))
print(rnn.shape)
# **return_sequences = True** ====> output_shape = **(batch_size, timesteps, num_units)**
rnn = SimpleRNN(50, return_sequences = True)(Input(shape = (10, 30)))
print(rnn.shape)
# return_state = True ===> outputs list of tensor: **[output, state]**
# - if return_sequences == False =>> output_shape = (batch_size, num_units)
# - if return_sequences == True =>> output_shape = (batch_size, timesteps, num_units)
rnn = SimpleRNN(50, return_sequences = False, return_state = True)(Input(shape = (10, 30)))
print(rnn[0].shape) # shape of output
print(rnn[1].shape) # shape of last state
rnn = SimpleRNN(50, return_sequences = True, return_state = True)(Input(shape = (10, 30)))
print(rnn[0].shape) # shape of output
print(rnn[1].shape) # shape of last state
# Current output and state can be unpacked as below
output, state = SimpleRNN(50, return_sequences = True, return_state = True)(Input(shape = (10, 30)))
print(output.shape)
print(state.shape)
# ## 2. LSTM
# - Outputs of LSTM are quite similar to those of RNNs, but there exist subtle differences
# - If you compare two diagrams below, there is one more type of "state" that is preserved to next module
#
# <br>
# <img src="http://colah.github.io/posts/2015-08-Understanding-LSTMs/img/LSTM3-SimpleRNN.png" style="width: 500px"/>
#
# <center> Standard RNN </center>
#
# <br>
# <img src="http://colah.github.io/posts/2015-08-Understanding-LSTMs/img/LSTM3-chain.png" style="width: 500px"/>
#
# <center> LSTM </center>
# In addition to "hidden state (ht)" in RNN, there exist "cell state (Ct)" in LSTM structure
#
# <br>
# <img src="http://colah.github.io/posts/2015-08-Understanding-LSTMs/img/LSTM3-focus-o.png" style="width: 500px"/>
#
# <center> Hidden State </center>
#
# <br>
# <img src="http://colah.github.io/posts/2015-08-Understanding-LSTMs/img/LSTM3-focus-C.png" style="width: 500px"/>
#
# <center> Cell State </center>
lstm = LSTM(50)(Input(shape = (10, 30)))
print(lstm.shape)
lstm = LSTM(50, return_sequences = False, return_state = True)(Input(shape = (10, 30)))
print(lstm[0].shape) # shape of output
print(lstm[1].shape) # shape of hidden state
print(lstm[2].shape) # shape of cell state
lstm = LSTM(50, return_sequences = True, return_state = True)(Input(shape = (10, 30)))
print(lstm[0].shape) # shape of output
print(lstm[1].shape) # shape of hidden state
print(lstm[2].shape) # shape of cell state
output, hidden_state, cell_state = LSTM(50, return_sequences = True, return_state = True)(Input(shape = (10, 30)))
print(output.shape)
print(hidden_state.shape)
print(cell_state.shape)
# ## 3. GRU
# - GRU, Popular variant of LSTM, does not have cell state
# - Hence, it has only hidden state, as simple RNN
gru = GRU(50)(Input(shape = (10, 30)))
print(gru.shape)
gru = GRU(50, return_sequences = False, return_state = True)(Input(shape = (10, 30)))
print(gru[0].shape) # shape of output
print(gru[1].shape) # shape of hidden state
gru = GRU(50, return_sequences = True, return_state = True)(Input(shape = (10, 30)))
print(gru[0].shape) # shape of output
print(gru[1].shape) # shape of hidden state
output, hidden_state = GRU(50, return_sequences = True, return_state = True)(Input(shape = (10, 30)))
print(output.shape)
print(hidden_state.shape)
|
NLP/1-Basic-RNN/0-understanding-rnn-structure.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: commons
# language: python
# name: commons
# ---
# Load data
import pandas as pd
pd.set_option('display.max_columns', 100)
train = pd.read_csv("../data/interim/train.csv")
dev = pd.read_csv("../data/interim/dev.csv")
test = pd.read_csv("../data/interim/test.csv")
df = pd.read_csv("../data/raw/WA_Fn-UseC_-Telco-Customer-Churn.csv")
assert df.shape[0] == train.shape[0] + dev.shape[0] + test.shape[0]
train.shape, dev.shape, test.shape, df.shape
# Same ratio of target values
print(" ", train['churn'].value_counts(normalize=True).values, "\n ", \
dev['churn'].value_counts(normalize=True).values, "\n ", \
test['churn'].value_counts(normalize=True).values)
train.head()
for column in train.columns:
print("-------", column, "-------\n", train[column].value_counts(), "\n", "-"*(len(column)+15), "\n")
train.info()
dev['senior_citizen'].value_counts()
train.columns
|
notebooks/2.exploration.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] pycharm={}
# # Making pairwise alignments with clustalo
# + pycharm={}
import os
import glob
import Bio
import pandas as pd
import numpy as np
from Bio import SeqIO
from Bio import AlignIO
# + pycharm={}
os.chdir("../data/families_seq/") # newfiles4 - version of working directory
# + [markdown] pycharm={}
# ##### Here we prepare our sample for Clustal
# + pycharm={}
# Here you can check how much sequences contain your sample-file
dictt = SeqIO.to_dict(SeqIO.parse("sample.fasta", "fasta"))
fname_list = list(dictt.keys())
len(fname_list)
# + [markdown] pycharm={}
# ##### Now we are going to make a lot of PAIRWISE-ALIGNMENT files
# + pycharm={}
i = 0
# # !mkdir clustalo
seq1, seq2 = fname_list[0], fname_list[2]
for num1 in range(i, len(fname_list)): #(i, len(fname_list)): (len(fname_list))
for num2 in range(i + 1, len(fname_list)): #(i + 1, len(fname_list)): (len(fname_list))
seq1, seq2 = fname_list[num1], fname_list[num2]
out = open('../data/depo.fasta', 'w')
out.write(">" + str(dictt[seq1].id) + '\n' + str(dictt[seq1].seq) + '\n' + ">" + str(dictt[seq2].id) + '\n' + str(dictt[seq2].seq) + '\n')
out.close()
# ! clustalo -i ./depo.fasta -o ./'{seq1}_{seq2}'.fasta
i += 1
# + pycharm={}
# To check how much pairwise-alignment files in your working directory
sum(list(range(62)))
# + pycharm={}
# ! rm depo.fasta
# + [markdown] pycharm={}
# # Making matrices
# + [markdown] pycharm={}
# ##### Firstly, we'll check, if there're the number of files we need in our 'clustalo' directory
# + pycharm={}
len(fname_list)**2 == len(glob.glob('*.fasta'))
# + pycharm={}
#os.mkdir('./matrices')
os.chdir('./matrices/')
os.getcwd()
# + [markdown] pycharm={}
# ##### Making matrices
# + pycharm={}
'''
This function returns you one matrix with shape == (1, 1200, 10)
'''
def make_matrix(fname):
# input file with pairwise alignment
alignment = AlignIO.read(fname, "fasta")
out_name = str(fname)[:-6]
# making file for matrix
out = open('mat_' + str(out_name) + '.txt', 'a')
for record in alignment:
out.write(str(record.seq) + '\n')
alignment_len = len(str(record.seq).strip())
out.close()
# making dataset 2 x alignment_len
fname = open('mat_' + str(out_name) + '.txt', 'r')
#print(fname)
seq1 = list(fname.readline().strip())
seq2 = list(fname.readline().strip())
# print(len(seq1), len(seq2))
fname.close()
data = pd.DataFrame(np.array((seq1, seq2)))
# making ZERO-matrix and dataset 10 x alignment_len
matrix = np.zeros((10, 1200)) #((10, 1200)) # or ((10, 1200))
mat_data = pd.DataFrame(matrix)
# [A1 T1 G1 C1 GAP1][A2 T2 G2 C2 GAP2]
# [00 01 02 03 0004][05 06 07 08 0009]
# print(alignment_len)
for num in range(alignment_len):
sym1 = data[num][0]
sym2 = data[num][1]
if sym1: #== 'A' or sym1 == 'T' or sym1 == 'G' or sym1 == 'C' or sym1 == '-':
if sym1 == 'A': mat_data[num][0] = 1
elif sym1 == 'T': mat_data[num][1] = 1
else:
if sym1 == 'G': mat_data[num][2] = 1
elif sym1 == 'C': mat_data[num][3] = 1
else: mat_data[num][4] = 1
if sym2: #== 'A' or sym2 == 'T' or sym2 == 'G' or sym2 == 'C' or sym2 == '-':
if sym2 == 'A': mat_data[num][5] = 1
elif sym2 == 'T': mat_data[num][6] = 1
else:
if sym2 == 'G': mat_data[num][7] = 1
elif sym2 == 'C': mat_data[num][8] = 1
else: mat_data[num][9] = 1
matrix1 = mat_data.astype(int)
# print(matrix1)
# Matrix reshaping
matrix = np.swapaxes(matrix, 1, 0).reshape(1, 1200, 10)
return matrix
# Saving binary file in .npy format
#print(matrix.shape)
os.remove('mat_' + str(out_name) + '.txt')
#np.save(str(out_name) + '.npy', matrix)
# + [markdown] pycharm={}
# ##### OLD VARIANT; SEE NEW VARIANT BELOW
# + pycharm={}
MakeMatrix = make_matrix('./')
# + pycharm={}
'''
I check if our sequences have the same length, don't look at this cell
'''
for file in glob.glob("mat*.txt"):
with open(file, 'r') as fname:
seq1 = fname.readline().strip()
seq2 = fname.readline().strip()
if (len(seq1) == len(seq2)) != True:
print(fname)
# + pycharm={}
# We shoul copy all files with pairwise alignments into the our working directory
mat = int(0)
sp = []
for file in glob.glob("*.fasta"):
print(file)
sp.append(file)
if type(mat) == int:
mat = make_matrix(file)
else:
nwm = make_matrix(file)
mat = np.concatenate((mat, nwm), axis=0)
mat.shape
print(len(sp))
# mat_RF02272_JH835498.1,4_RF03115_URS0000D65EA6_682634,5.txt
# + pycharm={}
np.save('./ncRNApair_datav5.npy', mat)
# + [markdown] pycharm={}
# ##### Getting binary vector (2500,)
# + pycharm={}
pre_vec = []
for pair in sp:
# print((int(pair.split(",")[1][0]), int(pair.split(",")[2][0])))
check = (int(pair.split(",")[1][0]) == int(pair.split(",")[2][0]))
if check == True:
pre_vec.append(1)
else:
pre_vec.append(0)
bin_vec = np.array(pre_vec)
# + pycharm={}
np.save('./ncRNApair_labelv5.npy', bin_vec)
# + [markdown] pycharm={}
# ##### NEW VARIANT; with more adequate order (in accordance with famlables.txt)
# + pycharm={}
'''
Here we make a list of ordered file-names
We got *fam70_test.txt* earlier using Creating_sample.ipynb
'''
new_sp = []
fnames = []
order = []
with open("fam70_test.txt", 'r') as fname:
for string in fname:
new_sp.append(string.strip()[:-2])
for file in glob.glob("*.fasta"):
fnames.append(file)
for i in new_sp:
for j in fnames:
if i in j.split(",")[0]:
order.append(j)
# + pycharm={}
# Checking the length of our list
len(order)
# + pycharm={}
# We should copy all files with pairwise alignments into the our working directory
mat = int(0)
for file in order: # glob.glob("*.fasta"):
if type(mat) == int:
mat = make_matrix(file)
else:
mat = np.concatenate((mat, make_matrix(file)), axis=0)
mat.shape
# + pycharm={}
# Save your matrix into file to use it later for neural network (input)
np.save('./ncRNApair_datav7_test.npy', mat)
# + pycharm={}
# Getting a binary vector (are the sequences belong to the same family)
pre_vec = []
for pair in order:
check = (int(pair.split(",")[1][0]) == int(pair.split(",")[2][0]))
if check == True:
pre_vec.append(0) # old version: pre_vec.append(1)
else:
pre_vec.append(1) # old version: pre_vec.append(0)
bin_vec = np.array(pre_vec)
# + pycharm={}
# Save your vector into file to use it later for neural network (input)
np.save('./ncRNApair_labelv7_test.npy', bin_vec)
|
notebooks/PairwiseClustalo_GettingMatrices.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.9.4 64-bit (''test'': conda)'
# name: python3710jvsc74a57bd04374d16277cd59720eda5e9a892d33ee7e53ac8b7c0031fbe42f60839aa8916a
# ---
# # Sensitivity estimate example
# +
import numpy as np
import matplotlib.pyplot as pl
# %pylab inline
import sys
sys.path.append('../../scripts')
from linear_regression import linreg
from IPython.core.display import Image
Image(filename='../../img/sensitivity_error_example.png',width=400)
# -
x = np.array([0.5, 1.0, 2.0, 5.0, 10.0, 20.0, 50.0, 100.0])
y = np.array([0.4, 1.0, 2.3, 6.9, 15.8, 36.4, 110.1, 253.2])
pl.plot(x,y,'--o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$y$ [V]')
pl.title('Calibration curve')
# Sensitivity, $K$ is:
#
# $ K_i = \left( \frac{\partial y}{\partial x} \right)_{x_i} $
K = np.diff(y)/np.diff(x)
print (K)
pl.plot(x[1:],K,'--o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$K$ [V/cm]')
pl.title('Sensitivity')
# Instead of working with non-linear curve of sensitivity we can use the usual trick: the logarithmic scale
pl.loglog(x,y,'--o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$y$ [V]')
pl.title('Logarithmic scale')
logK = np.diff(np.log10(y))/np.diff(np.log10(x))
print( logK)
pl.plot(x[1:],logK,'--o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$K$ [V/cm]')
pl.title('Logarithmic sensitivity')
pl.plot([x[1],x[-1]],[1.2,1.2],'r--')
pl.loglog(x,y,'o',x,x**(1.2))
pl.xlabel('$x$ [cm]')
pl.ylabel('$y$ [V]')
pl.title('Logarithmic scale')
pl.legend(('$y$','$x^{1.2}$'),loc='best')
pl.plot(x,y-x**(1.2),'o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$y - y_c$ [V]')
pl.title('Deviation plot')
# pl.legend(('$y$','$x^{1.2}$'),loc='best')
# ## Regression analysis
# Following the recipe of http://www.answermysearches.com/how-to-do-a-simple-linear-regression-in-python/124/
print (linreg(np.log10(x),np.log10(y)))
pl.loglog(x,y,'o',x,x**(1.21)-0.01252)
pl.xlabel('$x$ [cm]')
pl.ylabel('$y$ [V]')
pl.title('Logarithmic scale')
pl.legend(('$y$','$x^{1.2}$'),loc='best')
pl.plot(x,y-(x**(1.21)-0.01252),'o')
pl.xlabel('$x$ [cm]')
pl.ylabel('$y - y_c$ [V]')
pl.title('Deviation plot');
# pl.legend(('$y$','$x^{1.2}$'),loc='best')
|
notebooks/calibration/sensitivity_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from pymongo import MongoClient
# from auth import username, password
import urllib
from pprint import pprint
from tqdm import tqdm
import tensorflow as tf
import multiprocessing
from transformers import AutoTokenizer, TFAutoModel
import pickle
import numpy as np
import os
import preprocessor as p
import collections
import random
seed = 1111
# +
from transformers import (
AutoConfig,
AutoTokenizer,
TFAutoModelForSequenceClassification,
AdamW,
glue_convert_examples_to_features
)
# Choose model
# @markdown >The default model is <i><b>COVID-Twitter-BERT</b></i>. You can however choose <i><b>BERT Base</i></b> or <i><b>BERT Large</i></b> to compare these models to the <i><b>COVID-Twitter-BERT</i></b>. All these three models will be initiated with a random classification layer. If you go directly to the Predict-cell after having compiled the model, you will see that it still runs the predition. However the output will be random. The training steps below will finetune this for the specific task. <br /><br />
model_name = 'digitalepidemiologylab/covid-twitter-bert' #@param ["digitalepidemiologylab/covid-twitter-bert", "bert-large-uncased", "bert-base-uncased"]
# Initialise tokenizer
tokenizer = AutoTokenizer.from_pretrained(model_name)
# -
data_dir = '../src/data'
standardized_suffix = 'standardized'
# +
def create_int_feature(values):
feature = tf.train.Feature(int64_list=tf.train.Int64List(value=list(values)))
return feature
def create_float_feature(values):
feature = tf.train.Feature(float_list=tf.train.FloatList(value=list(values)))
return feature
# +
def generate_onehot_labels():
conspiracies = set()
for filename in os.listdir('../src/data'):
if 'standardized' in filename:
conspiracies.add(filename.split('-')[0])
return {consp:i for i, consp in enumerate(conspiracies)}
# generate_onehot_labels()
# -
def create_masked_lm_predictions(tokens, masked_lm_prob,
max_predictions_per_seq, vocab_words, rng):
"""Creates the predictions for the masked LM objective."""
cand_indexes = []
for (i, token) in enumerate(tokens):
if token == vocab["[CLS]"] or token == vocab["[SEP]"]:
continue
# Whole Word Masking means that if we mask all of the wordpieces
# corresponding to an original word. When a word has been split into
# WordPieces, the first token does not have any marker and any subsequence
# tokens are prefixed with ##. So whenever we see the ## token, we
# append it to the previous set of word indexes.
#
# Note that Whole Word Masking does *not* change the training code
# at all -- we still predict each WordPiece independently, softmaxed
# over the entire vocabulary.
if (FLAGS.do_whole_word_mask and len(cand_indexes) >= 1 and
token.startswith("##")):
cand_indexes[-1].append(i)
else:
cand_indexes.append([i])
rng.shuffle(cand_indexes)
output_tokens = list(tokens)
num_to_predict = min(max_predictions_per_seq,
max(1, int(round(len(tokens) * masked_lm_prob))))
masked_lms = []
covered_indexes = set()
for index_set in cand_indexes:
if len(masked_lms) >= num_to_predict:
break
# If adding a whole-word mask would exceed the maximum number of
# predictions, then just skip this candidate.
if len(masked_lms) + len(index_set) > num_to_predict:
continue
is_any_index_covered = False
for index in index_set:
if index in covered_indexes:
is_any_index_covered = True
break
if is_any_index_covered:
continue
for index in index_set:
covered_indexes.add(index)
masked_token = None
# 80% of the time, replace with [MASK]
if rng.random() < 0.8:
masked_token = "[MASK]"
else:
# 10% of the time, keep original
if rng.random() < 0.5:
masked_token = tokens[index]
# 10% of the time, replace with random word
else:
masked_token = vocab_words[rng.randint(0, len(vocab_words) - 1)]
output_tokens[index] = masked_token
masked_lms.append(MaskedLmInstance(index=index, label=tokens[index]))
assert len(masked_lms) <= num_to_predict
masked_lms = sorted(masked_lms, key=lambda x: x.index)
masked_lm_positions = []
masked_lm_labels = []
for p in masked_lms:
masked_lm_positions.append(p.index)
masked_lm_labels.append(p.label)
return (output_tokens, masked_lm_positions, masked_lm_labels)
def convert_instance_to_examples(X, label, tokenizer):
input_ids = X['input_ids']
token_type_ids = X['token_type_ids']
attention_mask = X['attention_mask']
rng = random.Random(seed)
masked_lm_prob = 0.1
vocab = tokenizer.get_vocab()
reverse_vocab = {v:k for k,v in vocab.items()}
for i in tqdm(range(input_ids.shape[0])):
output_tokens, masked_lm_positions, masked_lm_labels = create_masked_lm_predictions(input_ids[i], masked_lm_prob, max_predictions_per_seq, vocab_words, rng)
features = collections.OrderedDict()
features["input_ids"] = create_int_feature(np.array(input_ids[i]))
features["input_mask"] = create_int_feature(np.array(token_type_ids[i]))
features["segment_ids"] = create_int_feature(np.array(attention_mask[i]))
features["masked_lm_positions"] = create_int_feature(np.array(attention_mask[i]))
features["masked_lm_ids"] = create_float_feature(np.array(attention_mask[i]))
features["masked_lm_weights"] = create_int_feature(np.array(attention_mask[i]))
features["next_sentence_labels"] = create_int_feature(np.array(label))
break
# +
labels = generate_onehot_labels()
for filename in os.listdir('../src/data'):
if 'standardized' in filename:
with open('../src/data/'+filename, 'rb') as f:
X = pickle.load(f)
y = np.zeros(len(labels.keys()))
y[labels[filename.split('-')[0]]] = 1
y = tf.convert_to_tensor(y, dtype=tf.int64)
convert_instance_to_examples(X, y, tokenizer)
# print(y)
break
# -
|
scripts/Convert to TFRecords.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Case Study by: <NAME>
# ## 1.Read the three csv files which contains the score of same students in term1 for each Subject
import pandas as pd
df_math = pd.read_csv(r"C:\Users\saqla\Desktop\Python Certification for Data Science\Modules\Module 4\Dataset\MathScoreTerm1.csv", index_col='ID')
df_ds = pd.read_csv(r"C:\Users\saqla\Desktop\Python Certification for Data Science\Modules\Module 4\Dataset\DSScoreTerm1.csv", index_col='ID')
df_physics = pd.read_csv(r"C:\Users\saqla\Desktop\Python Certification for Data Science\Modules\Module 4\Dataset\PhysicsScoreTerm1.csv", index_col='ID')
df_ds
# ## 2.Remove the name and ethnicity column (to ensure confidentiality
df_math.drop(columns=['Name','Ethinicity'], inplace=True)
df_ds.drop(columns=['Name','Ethinicity'], inplace=True)
df_physics.drop(columns=['Name','Ethinicity'], inplace=True)
df_physics
# ## 3.Fill missing score data with zero
df_math.fillna(0,inplace=True)
df_ds.fillna(0,inplace=True)
df_physics.fillna(0,inplace=True)
df_physics.isna().sum()
# ## 4.Merge the three files
merged_df = df_math.merge(df_ds, on='ID', suffixes=('_math','_ds')).merge(df_physics,on='ID', suffixes=('_ds','_physics'))
merged_df
merged_df.drop(columns=['Age_math','Age_ds','Sex_math','Sex_ds'], inplace=True)
merged_df
merged_df.rename(columns={'Score':'Score_physics','Subject':'Subject_physics'}, inplace=True)
merged_df
# ## 5.Change Sex(M/F) Columnto 1/2 for further analysis
merged_df['Sex'].replace({'M':1,'F':2},inplace=True)
merged_df
# ## 6.Store the data in new file –ScoreFinal.csv
merged_df.to_csv(r"C:\Users\saqla\Desktop\Python Certification for Data Science\Modules\Module 4\Dataset\ScoreFinal.csv")
|
Python Practice/3. Numpy, Pandas and Matplotlib Practice/Module 4 - Case Study 2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
from collections import defaultdict
import re
# +
data_df = pd.read_csv("../data/Sharktankpitchesdeals.csv").drop(['Season_Epi_code', 'Pitched_Business_Identifier', 'Deal_Shark'], axis = 1)
# -
data_df.head()
data_df.iloc[0]['Pitched_Business_Desc']
# +
train, validate, test = np.split(data_df, [int(.7 * len(data_df)), int(.8 * len(data_df))])
# Get all data and labels in particular dataframes
train_data = train.drop(columns='Deal_Status')
train_labels = train['Deal_Status']
validate_data = validate.drop(columns='Deal_Status')
validate_labels = validate['Deal_Status']
test_data = test.drop(columns='Deal_Status')
test_labels = test['Deal_Status']
# -
train_data.shape
validate_data.shape
test_data.shape
type(train_data)
train_data.columns[0]
data = "Menu is 123absolutely perfect,loved it! "
re.sub('[^a-z\s]+',' ',data,flags=re.IGNORECASE)
re.sub('(\s+)',' ',data)
# +
arr = defaultdict(lambda:0)
arr[1]
arr[2]
list(arr.items())
arr[1]+=1
list(arr.items())
# -
train_data.iloc[0][0]
arr = np.array(train_data)
arr.shape
arr2 = arr.reshape(arr.shape[0],)
arr2.shape
arr2[0]
# <img src="../images/naive_bayes_formula.png"/>
# <img src="../images/likelihood_prob.png"/>
def preprocess_string(str_arg):
cleaned_str=re.sub('[^a-z\s]+',' ',str_arg,flags=re.IGNORECASE) #every char except alphabets is replaced
cleaned_str=re.sub('(\s+)',' ',cleaned_str) #multiple spaces are replaced by single space
cleaned_str=cleaned_str.lower() #converting the cleaned string to lower case
return cleaned_str # eturning the preprocessed string in tokenized form
# +
class CategoryInfo:
def __init__(self, bow_dict, prob_class, denom):
self.bow_dict = bow_dict
self.prob_class = prob_class
self.denom = denom
# -
class NaiveBayes:
def __init__(self,unique_classes):
self.classes=unique_classes # Constructor is sinply passed with unique number of classes of the training set
def addToBow(self, example, dict_index):
if isinstance(example,np.ndarray): example=example[0]
for token_word in example.split(): #for every word in preprocessed example
self.bow_dicts[dict_index][token_word]+=1 #increment in its count
def train(self, dataset, labels):
self.examples=dataset
self.labels=labels
self.bow_dicts=np.array([defaultdict(lambda:0) for index in range(self.classes.shape[0])])
#only convert to numpy arrays if initially not passed as numpy arrays - else its a useless recomputation
if not isinstance(self.examples,np.ndarray): self.examples=np.array(self.examples).reshape(self.examples.shape[0],)
if not isinstance(self.labels,np.ndarray): self.labels=np.array(self.labels).reshape(self.labels.shape[0],)
#constructing BoW for each category
for cat_index, cat in enumerate(self.classes):
all_cat_examples=self.examples[self.labels==cat] #filter all examples of category == cat
#get examples preprocessed
cleaned_examples=[preprocess_string(cat_example) for cat_example in all_cat_examples]
cleaned_examples=pd.DataFrame(data=cleaned_examples)
#now costruct BoW of this particular category
np.apply_along_axis(self.addToBow, 1 , cleaned_examples, cat_index)
###################################################################################################
prob_classes=np.empty(self.classes.shape[0])
all_words=[]
cat_word_counts=np.empty(self.classes.shape[0])
for cat_index,cat in enumerate(self.classes):
#Calculating prior probability p(c) for each class
prob_classes[cat_index]=np.sum(self.labels==cat)/float(self.labels.shape[0])
#Calculating total counts of all the words of each class
count=list(self.bow_dicts[cat_index].values())
cat_word_counts[cat_index]=np.sum(np.array(list(self.bow_dicts[cat_index].values())))+1 # |v| is remaining to be added
#get all words of this category
all_words+=self.bow_dicts[cat_index].keys()
#combine all words of every category & make them unique to get vocabulary -V- of entire training set
self.vocab=np.unique(np.array(all_words))
self.vocab_length=self.vocab.shape[0]
#computing denominator value
denoms=np.array([cat_word_counts[cat_index] + self.vocab_length + 1 for cat_index,cat in enumerate(self.classes)])
self.cats_info=[CategoryInfo(self.bow_dicts[cat_index], prob_classes[cat_index], denoms[cat_index]) for cat_index,cat in enumerate(self.classes)]
self.cats_info=np.array(self.cats_info)
def getExampleProb(self,test_example):
likelihood_prob=np.zeros(self.classes.shape[0]) #to store probability w.r.t each class
#finding probability w.r.t each class of the given test example
for cat_index,cat in enumerate(self.classes):
for test_token in test_example.split(): #split the test example and get p of each test word
####################################################################################
#This loop computes : for each word w [ count(w|c)+1 ] / [ count(c) + |V| + 1 ]
####################################################################################
#get total count of this test token from it's respective training dict to get numerator value
test_token_counts=self.cats_info[cat_index].bow_dict.get(test_token, 0) + 1
#now get likelihood of this test_token word
test_token_prob=test_token_counts/float(self.cats_info[cat_index].denom)
#remember why taking log? To prevent underflow!
likelihood_prob[cat_index] += np.log(test_token_prob)
# we have likelihood estimate of the given example against every class but we need posterior probility
post_prob=np.empty(self.classes.shape[0])
for cat_index,cat in enumerate(self.classes):
post_prob[cat_index]=likelihood_prob[cat_index] + np.log(self.cats_info[cat_index].prob_class)
return post_prob
def test(self,test_set):
predictions=[] #to store prediction of each test example
for example in test_set:
#preprocess the test example the same way we did for training set exampels
cleaned_example=preprocess_string(example)
#simply get the posterior probability of every example
post_prob=self.getExampleProb(cleaned_example) #get prob of this example for both classes
#simply pick the max value and map against self.classes!
predictions.append(self.classes[np.argmax(post_prob)])
return np.array(predictions)
nb=NaiveBayes(np.unique(train_labels))
# +
print ("---------------- Training In Progress --------------------")
nb.train(train_data, train_labels) #start tarining by calling the train function
print ('----------------- Training Completed ---------------------')
# +
validate_examples=np.array(validate_data).reshape(validate_data.shape[0],)
validatePclasses=nb.test(validate_examples)
#check how many predcitions actually match original test labels
validate_acc=np.sum(validatePclasses==validate_labels)/float(validate_labels.shape[0])
print ("Validate Set Examples: ",validate_labels.shape[0])
print ("Validate Set Accuracy: ",validate_acc*100,"%")
# +
test_examples=np.array(test_data).reshape(test_data.shape[0],)
pclasses=nb.test(test_examples)
#check how many predcitions actually match original test labels
test_acc=np.sum(pclasses==test_labels)/float(test_labels.shape[0])
print ("Test Set Examples: ",test_labels.shape[0])
print ("Test Set Accuracy: ",test_acc*100,"%")
# +
train_examples=np.array(train_data).reshape(train_data.shape[0],)
# For Fun, this should be close to 100%.
pclasses=nb.test(train_examples)
#check how many predcitions actually match original test labels
train_acc=np.sum(pclasses==train_labels)/float(train_labels.shape[0])
print ("Train Set Examples: ",train_labels.shape[0])
print ("Train Set Accuracy: ",train_acc*100,"%")
# -
|
Assignments/Assignment_NaiveBayes_SharkTank.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Coletando Dados
#
# <p>Existem vários formatos para um conjunto de dados: .csv, .json, .xlsx etc.<br>
# E esse conjunto pode estar armazenado em diferentes lugares, localmente ou online.
#
# Neste notebook, você aprenderá como carregar um conjunto de dados em formato csv (comma separated values), salvo localmente. Iremos utilizar o Automobile Dataset (uma fonte gratuita disponível <a href="https://archive.ics.uci.edu/ml/machine-learning-databases/autos/imports-85.data" target="_blank">aqui</a>)
#
#
# Usaremos também este arquivo como um exemplo para praticar formas de tratamento e leitura dos dados.
#
#
# importando as bibliotecas que iremos utilizar
# Pandas e Numpy
import pandas as pd
import numpy as np
# ### Lendo o arquivo
#
# Usamos a função <b>pandas.read_csv()</b> para ler o arquivo csv.
#
# Como parametro dessa função, enviaremos uma string com o caminho do arquivo que queremos ler (pode ser uma URL ou o endereço do arquivo local). Se verificarmos o arquivo, iremos notar que não existe cabeçalho, nesse caso, podemos definir também o parametro <i>headers=None</i> para que o pandas não defina automaticamente a primeira linha como cabeçalho (iremos criar cabeçalhos em seguida).
caminho_arquivo = "https://archive.ics.uci.edu/ml/machine-learning-databases/autos/imports-85.data"
df = pd.read_csv(caminho_arquivo, header=None)
# Com o nosso dataframe (df) criado, podemos dar uma breve olhada nos dados. Para isso, podemos utilizar o método <i>dataframe.head(n)</i> que exibe as primeiras n linhas do dataframe ou <i>dataframe.tail(n)</i> que exibe as últimas n linhas.
df.head() # se você não definir um valor de n, o padrão será 5
# ### Adicionando cabeçalhos
# <p> Como vimos, o arquivo que utilizamos não possui um cabeçalho (a primeira linha não descreve os nomes das colunas). É importante que tenhamos cabeçalhos definidos para que possamos trabalhar com os dados e compreender qual informação cada coluna representa.
#
# Iremos utilizar a descrição das colunas do dataset que estamos utilizando, disponível <a href="https://archive.ics.uci.edu/ml/datasets/Automobile" target="_blank">aqui</a> para adicionarmos o nosso cabeçalho manualmente.
# +
#primeiro criamos uma lista de "cabeçalhos" que inclui todos os nomes de colunas em ordem.
headers = ["symboling","normalized-losses","make","fuel-type","aspiration", "num-of-doors","body-style",
"drive-wheels","engine-location","wheel-base", "length","width","height","curb-weight","engine-type",
"num-of-cylinders", "engine-size","fuel-system","bore","stroke","compression-ratio","horsepower",
"peak-rpm","city-mpg","highway-mpg","price"]
#em seguida, usamos dataframe.columns = headers para substituir os cabeçalhos pela lista que criamos.
df.columns = headers
df.head(10)
# -
# Repare os dados da coluna <i>normalized-losses</i>. Algumas linhas exibem o caracter ? no lugar do valor que deveriam apresentar. Esse tipo de erro é comum e necessita de uma intervenção antes de declarar que o dataframe está pronto para análise.
# primeiro, iremos subistutir o caracter '?' por NaN (not a number)
# crio um dataframe temporário (df1) que vai armazenar o resultado dessa operação
df1=df.replace('?',np.NaN)
# +
# Agora utilizamos a função dropna que vai excluir todas as linhas do dataframe que contenham
# o valor NaN na coluna preço:
df=df1.dropna(subset=["price"], axis=0)
df.head(20)
# -
# # É isso!
#
# ### Este é apenas um exemplo de coleta e tratamento básico de dados.
#
# ### Muito obrigado pela sua leitura!
#
# ## Autor:
#
# <h4><NAME></h4>
#
# Você pode encontrar mais conteúdo no meu Medium<br> ou então entrar em contato comigo :D
#
# <a href="https://www.linkedin.com/in/anderson-cordeiro-26986430/" target="_blank">[LinkedIn]</a>
# <a href="https://medium.com/@andcordeiro" target="_blank">[Medium]</a>
#
|
pandas-lendo-csv.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # First Words
# Finding your very own best first word with data analytics, Jupyter Notebooks, Python, and Matplotlib
# Getting all the allowed guesses
with open("allowed_words.txt") as f:
all_words = f.read().split("\n")
print(all_words[:50])
print(len(all_words))
# # Vowels and consonants
# People love to use well en-voweled starting words!
#
# Meanwhile, there is an opposing shool of thought about using sonorant consonants (y, w, l, r, m, n, and ng).
# +
# the most en-voweled words
from collections import defaultdict
vowel_count = defaultdict(int)
for word in all_words:
for letter in "aeiou":
if letter in word:
vowel_count[word] += 1
sorted_vowel_count = sorted(list(vowel_count.items()), key=lambda x:x[1], reverse=True)
print(sorted_vowel_count[:20])
# +
# A demo on sorting tuples
my_dict = {"a": 1, "b": 11, "v": 12, "j": 4}
new_list = sorted(list(my_dict.items()), key=lambda x:x[1], reverse=True)
print(new_list)
# +
# Sonorant consonants galore *y, w, l, r, m, n, and ng*
# the most en-voweled words
from collections import defaultdict
cons_count = defaultdict(int)
for word in all_words:
for letter in ["y", "w", "l", "r", "m", "n"]:
if letter in word:
cons_count[word] += 1
sorted_cons_count = sorted(list(cons_count.items()), key=lambda x:x[1], reverse=True)
print(sorted_cons_count[:20])
# -
# ## Letter Frequency
# What are the most abundant letters in 5 letter words?
# +
# Counting up all the letters in all the words
from collections import defaultdict
total_count = defaultdict(int)
for word in all_words:
for letter in word:
total_count[letter] += 1
sorted(total_count.items(), key=lambda x:x[1], reverse=True)
print(total_count)
# +
# Graphing the letter distribution\
import matplotlib.pyplot as plt
data = sorted(total_count.items())
x = [i[0] for i in data]
y = [i[1] for i in data]
plt.bar(x, y)
print(total_count)
# +
# Adding this to our scoring system - what is the scale of the score?
total_words = len(all_words)
print(total_count["e"]/total_words)
# -
# ## Letter Placement
# Does where we put out letters matter much?
# +
# Counting up letter's places
from collections import defaultdict
position_count = defaultdict(int)
for word in all_words:
for i, letter in enumerate(word):
position_count[(i,letter)] += 1
print(position_count)
# +
# Graphing each position's best letters
import string
from matplotlib.pyplot import figure
alphabet = string.ascii_lowercase
width = 1.2
group_gap = 3
start = 1
num_groups = 5
group_size = len(alphabet)
group_starts = [start]
for i in range(1, num_groups):
group_starts.append(group_starts[-1] + group_size * width + group_gap)
figure(figsize=(25, 6))
count = 0
for i, group_start in enumerate(group_starts):
spots = [group_start + width * i for i in range(group_size)]
y = [position_count[(i, letter)] for letter in alphabet]
plt.bar(spots, y)
count += 1
label_text = []
label_spots = []
# label_spots = [group_start + group_size * width /2 for i in range(group_size)]
label_spots = []
for i, start in enumerate(group_starts):
label_spots += [start + width * i for i in range(group_size)]
label_text = list(alphabet*5)
plt.xticks(label_spots, label_text)
# +
# Graphing each letter's best position
from matplotlib.pyplot import figure
width = 1
group_gap = 2
start = 1
num_groups = len(alphabet)
group_size = 5
group_starts = [start]
for i in range(1, num_groups):
group_starts.append(group_starts[-1] + group_size * width + group_gap)
figure(figsize=(25, 6))
label_spots = []
label_text = []
for i, group_start in enumerate(group_starts):
letter = alphabet[i]
spots = [group_start + width * i for i in range(group_size)]
y = [position_count[(i, letter)] for i in range(group_size)]
plt.bar(spots, y)
# label_spots = [group_start + group_size * width /2 for i in range(group_size)]
label_spots = [group_start + (group_size - 1) * width/2 for group_start in group_starts]
label_text = list(alphabet)
plt.xticks(label_spots, label_text)
# +
# Scoring based on position - what is the best score?
position_count[(4,"s")]/len(all_words)
# -
# ## Brining it all together - A scoring system
# +
# # Brining it all together
from collections import defaultdict
def score_possibilites(word_selection, total_words=None):
score_count = defaultdict(int)
if total_words == None:
total_words = len(word_selection)
for word in word_selection:
if len(set(list(word))) < len(word):
continue
# for letter in ["y", "w", "l", "r", "m", "n", "ng"]:
# if letter in word:
# score_count[word] += 0.5
# for letter in "aeiou":
# if letter in word:
# score_count[word] += 0.5
for letter in set(list(word)):
score_count[word] += total_count[letter]/total_words
for i, letter in enumerate(word):
position_score = position_count[(i, letter)]/total_words
score_count[word] += 1.1 * position_score
sorted_score_count = sorted(list(score_count.items()), key=lambda x:x[1], reverse=True)
return sorted_score_count
sorted_score_count = score_possibilites(all_words, len(all_words))
for word, score in sorted_score_count[0:40]:
print(word, score)
# -
# ## Are we ready for turn 2?
# Have we set ourselvse up for success? Or taken all the useful letters in one go?
#
# How our scoring system looks going into turn 2
# (if you want to try a disjoin word, or you're palying hard mode and your first word is a dud!)
# +
sorted_score_count = score_possibilites(all_words, len(all_words))
def no_incorect_letters(word, incorrect):
for letter in word:
if letter in incorrect:
return False
return True
def eliminate_first_letters(word_selection, first_word):
remaining_words = []
for word in word_selection:
if no_incorect_letters(word, first_word):
remaining_words.append(word)
return remaining_words
total_words = len(all_words)
double_scores = []
for first_word, score in sorted_score_count[0:500]:
remaining_words = eliminate_first_letters(all_words, first_word)
second_words = score_possibilites(remaining_words, total_words=total_words)
best_second, second_score = second_words[0]
double_score = (f'{first_word} ({round(score, 2)}) -> {best_second} ({round(second_score,2)})', score + second_score)
double_scores.append(double_score)
# print(f'{first_word} ({round(score, 2)}) -> {best_second} ({round(second_score,2)}) = {round(score + second_score, 2)}')
sorted_doubles = sorted(double_scores, key=lambda x: x[1], reverse=True)
for content, score in sorted_doubles[:20]:
print(content + " = " + str(round(score, 2)))
|
Episode 2 - FIRST WORDS/first_words.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
#
#
# Working with sensor locations
# =============================
#
# This tutorial describes how to read and plot sensor locations, and how
# the physical location of sensors is handled in MNE-Python.
# :depth: 2
#
# As usual we'll start by importing the modules we need and loading some
# `example data <sample-dataset>`:
#
# +
import os
import numpy as np
import matplotlib.pyplot as plt
from mpl_toolkits.mplot3d import Axes3D # noqa
import mne
sample_data_folder = mne.datasets.sample.data_path()
sample_data_raw_file = os.path.join(sample_data_folder, 'MEG', 'sample',
'sample_audvis_raw.fif')
raw = mne.io.read_raw_fif(sample_data_raw_file, preload=True, verbose=False)
# -
# About montages and layouts
# ^^^^^^^^^^^^^^^^^^^^^^^^^^
#
# MNE-Python comes pre-loaded with information about the sensor positions of
# many MEG and EEG systems. This information is stored in *layout files* and
# *montages*. :class:`Layouts <mne.channels.Layout>` give sensor positions in 2
# dimensions (defined by ``x``, ``y``, ``width``, and ``height`` values for
# each sensor), and are primarily used for illustrative purposes (i.e., making
# diagrams of approximate sensor positions in top-down diagrams of the head).
# In contrast, :class:`montages <mne.channels.DigMontage>` contain sensor
# positions in 3D (``x``, ``y``, ``z``, in meters). Many layout and montage
# files are included during MNE-Python installation, and are stored in your
# ``mne-python`` directory, in the :file:`mne/channels/data/layouts` and
# :file:`mne/channels/data/montages` folders, respectively:
#
#
data_dir = os.path.join(os.path.dirname(mne.__file__), 'channels', 'data')
for subfolder in ['layouts', 'montages']:
print('\nBUILT-IN {} FILES'.format(subfolder[:-1].upper()))
print('======================')
print(sorted(os.listdir(os.path.join(data_dir, subfolder))))
# .. sidebar:: Computing sensor locations
#
# If you are interested in how standard ("idealized") EEG sensor positions
# are computed on a spherical head model, the `eeg_positions`_ repository
# provides code and documentation to this end.
#
# As you may be able to tell from the filenames shown above, the included
# montage files are all for EEG systems. These are *idealized* sensor positions
# based on a spherical head model. Montage files for MEG systems are not
# provided because the 3D coordinates of MEG sensors are included in the raw
# recordings from MEG systems, and are automatically stored in the ``info``
# attribute of the :class:`~mne.io.Raw` file upon loading. In contrast, layout
# files *are* included for MEG systems (to facilitate easy plotting of MEG
# sensor location diagrams).
#
# You may also have noticed that the file formats and filename extensions of
# layout and montage files vary considerably. This reflects different
# manufacturers' conventions; to simplify this, the montage and layout loading
# functions in MNE-Python take the filename *without its extension* so you
# don't have to keep track of which file format is used by which manufacturer.
# Examples of this can be seen in the following sections.
#
# If you have digitized the locations of EEG sensors on the scalp during your
# recording session (e.g., with a Polhemus Fastrak digitizer), these can be
# loaded in MNE-Python as :class:`~mne.channels.DigMontage` objects; see
# `reading-dig-montages` (below).
#
#
# Working with layout files
# ~~~~~~~~~~~~~~~~~~~~~~~~~
#
# To load a layout file, use the :func:`mne.channels.read_layout`
# function, and provide the filename *without* its file extension. You can then
# visualize the layout using its :meth:`~mne.channels.Layout.plot` method, or
# (equivalently) by passing it to :func:`mne.viz.plot_layout`:
#
#
biosemi_layout = mne.channels.read_layout('biosemi')
biosemi_layout.plot() # same result as: mne.viz.plot_layout(biosemi_layout)
# Similar to the ``picks`` argument for selecting channels from
# :class:`~mne.io.Raw` objects, the :meth:`~mne.channels.Layout.plot` method of
# :class:`~mne.channels.Layout` objects also has a ``picks`` argument. However,
# because layouts only contain information about sensor name and location (not
# sensor type), the :meth:`~mne.channels.Layout.plot` method only allows
# picking channels by index (not by name or by type). Here we find the indices
# we want using :func:`numpy.where`; selection by name or type is possible via
# :func:`mne.pick_channels` or :func:`mne.pick_types`.
#
#
midline = np.where([name.endswith('z') for name in biosemi_layout.names])[0]
biosemi_layout.plot(picks=midline)
# If you're working with a :class:`~mne.io.Raw` object that already has sensor
# positions incorporated, you can create a :class:`~mne.channels.Layout` object
# with either the :func:`mne.channels.make_eeg_layout` function or
# (equivalently) the :func:`mne.channels.find_layout` function.
#
#
layout_from_raw = mne.channels.make_eeg_layout(raw.info)
# same result as: mne.channels.find_layout(raw.info, ch_type='eeg')
layout_from_raw.plot()
# <div class="alert alert-info"><h4>Note</h4><p>There is no corresponding ``make_meg_layout`` function because sensor
# locations are fixed in a MEG system (unlike in EEG, where the sensor caps
# deform to fit each subject's head). Thus MEG layouts are consistent for a
# given system and you can simply load them with
# :func:`mne.channels.read_layout`, or use :func:`mne.channels.find_layout`
# with the ``ch_type`` parameter, as shown above for EEG.</p></div>
#
# All :class:`~mne.channels.Layout` objects have a
# :meth:`~mne.channels.Layout.save` method that allows writing layouts to disk,
# in either :file:`.lout` or :file:`.lay` format (which format gets written is
# inferred from the file extension you pass to the method's ``fname``
# parameter). The choice between :file:`.lout` and :file:`.lay` format only
# matters if you need to load the layout file in some other software
# (MNE-Python can read either format equally well).
#
#
# Working with montage files
# ~~~~~~~~~~~~~~~~~~~~~~~~~~
#
# Built-in montages are loaded and plotted in a very similar way to layouts.
# However, the :meth:`~mne.channels.Montage.plot` method of
# :class:`~mne.channels.Montage` objects has some additional parameters, such
# as whether to display channel names or just points (the ``show_names``
# parameter) and whether to display sensor positions in 3D or as a 2D topomap
# (the ``kind`` parameter):
#
#
ten_twenty_montage = mne.channels.make_standard_montage('standard_1020')
ten_twenty_montage.plot(show_names=False)
fig = ten_twenty_montage.plot(kind='3d')
fig.gca().view_init(azim=70, elev=15)
# Similar functionality is also available with the
# :meth:`~mne.io.Raw.plot_sensors` method of :class:`~mne.io.Raw` objects,
# again with the option to plot in either 2D or 3D.
# :meth:`~mne.io.Raw.plot_sensors` also allows channel selection by type, can
# color-code channels in various ways (by default, channels listed in
# ``raw.info['bads']`` will be plotted in red), and allows drawing into an
# existing matplotlib ``axes`` object (so the channel positions can easily be
# made as a subplot in a multi-panel figure):
#
#
fig = plt.figure()
ax2d = fig.add_subplot(121)
ax3d = fig.add_subplot(122, projection='3d')
raw.plot_sensors(ch_type='eeg', axes=ax2d)
raw.plot_sensors(ch_type='eeg', axes=ax3d, kind='3d')
ax3d.view_init(azim=70, elev=15)
#
# Reading sensor digitization files
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
#
# It's probably evident from the 2D topomap above that there is some
# irregularity in the EEG sensor positions in the `sample dataset
# <sample-dataset>` — this is because the sensor positions in that dataset are
# digitizations of the sensor positions on an actual subject's head. Depending
# on what system was used to scan the positions one can use different
# reading functions (:func:`mne.channels.read_dig_captrack` for
# a CapTrak Brain Products system, :func:`mne.channels.read_dig_egi`
# for an EGI system, :func:`mne.channels.read_dig_polhemus_isotrak` for
# Polhemus ISOTRAK, :func:`mne.channels.read_dig_fif` to read from
# a `.fif` file or :func:`mne.channels.read_dig_hpts` to read MNE `.hpts`
# files. The read :class:`montage <mne.channels.DigMontage>` can then be added
# to :class:`~mne.io.Raw` objects with the :meth:`~mne.io.Raw.set_montage`
# method; in the sample data this was done prior to saving the
# :class:`~mne.io.Raw` object to disk, so the sensor positions are already
# incorporated into the ``info`` attribute of the :class:`~mne.io.Raw` object.
# See the documentation of the reading functions and
# :meth:`~mne.io.Raw.set_montage` for further details. Once loaded,
# locations can be plotted with :meth:`~mne.channels.DigMontage.plot` and
# saved with :meth:`~mne.channels.DigMontage.save`, like when working
# with a standard montage.
#
# The possibilities to read in digitized montage files are summarized
# in `dig-formats`.
#
# <div class="alert alert-info"><h4>Note</h4><p>When setting a montage with :meth:`~mne.io.Raw.set_montage`
# the measurement info is updated at two places (the `chs`
# and `dig` entries are updated). See `tut-info-class`.
# `dig` will potentially contain more than channel locations,
# such HPI, head shape points or fiducials 3D coordinates.</p></div>
#
# Rendering sensor position with mayavi
# ^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^^
#
# It is also possible to render an image of a MEG sensor helmet in 3D, using
# mayavi instead of matplotlib, by calling the :func:`mne.viz.plot_alignment`
# function:
#
#
fig = mne.viz.plot_alignment(raw.info, trans=None, dig=False, eeg=False,
surfaces=[], meg=['helmet', 'sensors'],
coord_frame='meg')
mne.viz.set_3d_view(fig, azimuth=50, elevation=90, distance=0.5)
# :func:`~mne.viz.plot_alignment` requires an :class:`~mne.Info` object, and
# can also render MRI surfaces of the scalp, skull, and brain (by passing
# keywords like ``'head'``, ``'outer_skull'``, or ``'brain'`` to the
# ``surfaces`` parameter) making it useful for `assessing coordinate frame
# transformations <plot_source_alignment>`. For examples of various uses of
# :func:`~mne.viz.plot_alignment`, see
# :doc:`../../auto_examples/visualization/plot_montage`,
# :doc:`../../auto_examples/visualization/plot_eeg_on_scalp`, and
# :doc:`../../auto_examples/visualization/plot_meg_sensors`.
#
# .. LINKS
#
#
#
|
dev/_downloads/59a29cf7eb53c7ab95857dfb2e3b31ba/plot_40_sensor_locations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
conda activate alpacaenv
# +
# Import Libraries
import os
import requests
import pandas as pd
from dotenv import load_dotenv
import json
# %matplotlib inline
# -
# Laod .env environment
load_dotenv()
# Pull in API Key
api_key = os.getenv("glassnode_api")
type(api_key)
# Define crypto currencies to pull
crypto_list = ["BTC", "BCH", "ETH", "LTC", "USDT", "DOGE"]
# Define URLs
price_url = 'https://api.glassnode.com/v1/metrics/market/price_usd'
volume_url = 'https://api.glassnode.com/v1/metrics/transactions/transfers_volume_sum'
mkt_cap_url = 'https://api.glassnode.com/v1/metrics/market/marketcap_usd'
mining_url = 'https://api.glassnode.com/v1/metrics/mining/volume_mined_sum'
exchange_fee_url = 'https://api.glassnode.com/v1/metrics/fees/exchanges_sum'
# ## BTC API Data Pull
# +
# Price API Request
btc_price_res = requests.get(price_url,
params={'a': 'BTC',
'i': '24h',
'api_key': api_key})
# Convert price to Pandas Dataframe, set index to time and clean up file
btc_price_df = pd.read_json(btc_price_res.text, convert_dates=['t'])
btc_price_df.columns = ['Date', 'Price']
btc_price_df.set_index('Date', inplace=True)
# Volume API Request
btc_volume_res = requests.get(volume_url,
params={'a': 'BTC',
'i': '24h',
'api_key': api_key})
# Convert volume to Pandas Dataframe, set index to time and clean up file
btc_volume_df = pd.read_json(btc_volume_res.text, convert_dates=['t'])
btc_volume_df.columns = ['Date', 'Volume']
btc_volume_df.set_index('Date', inplace=True)
# Market Cap API Request
btc_mkt_cap_res = requests.get(mkt_cap_url,
params={'a': 'BTC',
'i': '24h',
'api_key': api_key})
# Convert Market Cap to Pandas Dataframe, set index to time and clean up file
btc_mkt_cap_df = pd.read_json(btc_mkt_cap_res.text, convert_dates=['t'])
btc_mkt_cap_df.columns = ['Date', 'Market Cap']
btc_mkt_cap_df.set_index('Date', inplace=True)
# Mining API Request
btc_mining_res = requests.get(mining_url,
params={'a': 'BTC',
'i': '24h',
'api_key': api_key})
# Convert Mining to Pandas Dataframe, set index to time and clean up file
btc_mining_df = pd.read_json(btc_mining_res.text, convert_dates=['t'])
btc_mining_df.columns = ['Date', 'Blocks Mined']
btc_mining_df.set_index('Date', inplace=True)
# -
btc_price_df
btc_volume_df
btc_mkt_cap_df
btc_mining_df
# ## BTC Data Aggregating & Cleaning
# +
# Define all the different data frames into a list
btc_frames = [btc_price_df, btc_volume_df, btc_mkt_cap_df, btc_mining_df]
# Concatenate all the dataframes into one
btc_data = pd.concat(btc_frames, axis=1, join="outer", ignore_index=False)
btc_data
# -
|
.ipynb_checkpoints/Get_Glen_a_condo-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Batch processing with Kubeflow Pipelines
# In this notebook we will dive into how you can run batch processing with Kubeflow Pipelines and Seldon Core.
#
# Dependencies:
# * Seldon core installed as per the docs with [Istio Ingress](https://docs.seldon.io/projects/seldon-core/en/latest/workflow/install.html#install-seldon-core-with-helm)
# * Kubeflow Pipelines installed (installation instructions in this notebook)
#
# 
#
# ### Kubeflow Pipelines Setup
#
# Setup the pipeline in your current cluster:
# + language="bash"
# export PIPELINE_VERSION=0.5.1
# kubectl apply -k github.com/kubeflow/pipelines/manifests/kustomize/cluster-scoped-resources?ref=$PIPELINE_VERSION
# kubectl wait --for condition=established --timeout=60s crd/applications.app.k8s.io
# kubectl apply -k github.com/kubeflow/pipelines/manifests/kustomize/env/dev?ref=$PIPELINE_VERSION
# -
# We also install the Python Library so we can create our pipeline:
pip install kfp==0.5.1
# ### Add Batch Data
#
# In order to run our batch job we will need to create some batch data that can be used to process.
#
# This batch dataset will be pushed to a minio instance so it can be downloaded from Minio (which we need to install first)
#
# #### Install Minio
# + language="bash"
# helm install minio stable/minio \
# --set accessKey=minioadmin \
# --set secretKey=minioadmin \
# --set image.tag=RELEASE.2020-04-15T19-42-18Z
# -
# #### Forward the Minio port so you can access it
#
# You can do this by runnning the following command in your terminal:
# ```
# kubectl port-forward svc/minio 9000:9000
# ```
#
# #### Configure local minio client
# !mc config host add minio-local http://localhost:9000 minioadmin minioadmin
# #### Create some input for our model
#
# We will create a file that will contain the inputs that will be sent to our model
with open("assets/input-data.txt", "w") as f:
for i in range(10000):
f.write("[[1, 2, 3, 4]]\n")
# Check the contents of the file
# !wc -l assets/input-data.txt
# !head assets/input-data.txt
# #### Upload the file to our minio
# !mc mb minio-local/data
# !mc cp assets/input-data.txt minio-local/data/
# ### Create Kubeflow Pipeline
#
# We are now able to create a kubeflow pipeline that will allow us to enter the batch parameters through the UI.
#
# We will also be able to add extra steps that will download the data from a Minio client.
# mkdir -p assets/
# We use the pipeline syntax to create the kubeflow pipeline, as outlined below:
# +
# %%writefile assets/seldon-batch-pipeline.py
import kfp.dsl as dsl
import yaml
from kubernetes import client as k8s
@dsl.pipeline(
name='SeldonBatch',
description='A batch processing pipeline for seldon models'
)
def nlp_pipeline(
namespace="kubeflow",
seldon_server="SKLEARN_SERVER",
model_path="gs://seldon-models/v1.13.0-dev/sklearn/iris",
gateway_endpoint="istio-ingressgateway.istio-system.svc.cluster.local",
retries=3,
replicas=10,
workers=100,
input_path="data/input-data.txt",
output_path="data/output-data.txt"):
"""
Pipeline
"""
vop = dsl.VolumeOp(
name='seldon-batch-pvc',
resource_name="seldon-batch-pvc",
modes=dsl.VOLUME_MODE_RWO,
size="2Mi"
)
seldon_deployment_yaml = f"""
apiVersion: machinelearning.seldon.io/v1
kind: SeldonDeployment
metadata:
name: "{{{{workflow.name}}}}"
namespace: "{namespace}"
spec:
name: "{{{{workflow.name}}}}"
predictors:
- graph:
children: []
implementation: "{seldon_server}"
modelUri: "{model_path}"
name: classifier
name: default
"""
deploy_step = dsl.ResourceOp(
name="deploy_seldon",
action="create",
k8s_resource=yaml.safe_load(seldon_deployment_yaml))
scale_and_wait = dsl.ContainerOp(
name="scale_and_wait_seldon",
image="bitnami/kubectl:1.17",
command="bash",
arguments=[
"-c",
f"sleep 10 && kubectl scale --namespace {namespace} --replicas={replicas} sdep/{{{{workflow.name}}}} && sleep 2 && kubectl rollout status deploy/$(kubectl get deploy -l seldon-deployment-id={{{{workflow.name}}}} -o jsonpath='{{.items[0].metadata.name'}})"
])
download_from_object_store = dsl.ContainerOp(
name="download-from-object-store",
image="minio/mc:RELEASE.2020-04-17T08-55-48Z",
command="sh",
arguments=[
"-c",
f"mc config host add minio-local http://minio.default.svc.cluster.local:9000 minioadmin minioadmin && mc cp minio-local/{input_path} /assets/input-data.txt"
],
pvolumes={ "/assets": vop.volume })
batch_process_step = dsl.ContainerOp(
name='data_downloader',
image='seldonio/seldon-core-s2i-python37:1.13.0-dev command="seldon-batch-processor",
arguments=[
"--deployment-name", "{{workflow.name}}",
"--namespace", namespace,
"--host", gateway_endpoint,
"--retries", retries,
"--input-data-path", "/assets/input-data.txt",
"--output-data-path", "/assets/output-data.txt",
"--benchmark"
],
pvolumes={ "/assets": vop.volume }
)
upload_to_object_store = dsl.ContainerOp(
name="upload-to-object-store",
image="minio/mc:RELEASE.2020-04-17T08-55-48Z",
command="sh",
arguments=[
"-c",
f"mc config host add minio-local http://minio.default.svc.cluster.local:9000 minioadmin minioadmin && mc cp /assets/output-data.txt minio-local/{output_path}"
],
pvolumes={ "/assets": vop.volume })
delete_step = dsl.ResourceOp(
name="delete_seldon",
action="delete",
k8s_resource=yaml.safe_load(seldon_deployment_yaml))
scale_and_wait.after(deploy_step)
download_from_object_store.after(scale_and_wait)
batch_process_step.after(download_from_object_store)
upload_to_object_store.after(batch_process_step)
delete_step.after(upload_to_object_store)
if __name__ == '__main__':
import kfp.compiler as compiler
compiler.Compiler().compile(nlp_pipeline, __file__ + '.tar.gz')
# -
# ### Trigger the creation
# We will run the python file which triggers the creation of the pipeline that we can the upload on the UI:
# !python assets/seldon-batch-pipeline.py
# Check the pipeline has been created:
# !ls assets/
# ### Open the Kubeflow Pipelines UI
#
# We can now open the UI by port forwarding the UI with the following command:
#
# ```
# kubectl port-forward svc/ml-pipeline-ui -n kubeflow 8000:80
# ```
#
# And we can open it locally in our browser via [http://localhost:8000](http://localhost:8000)
#
# Now we can follow the standard steps to create and deploy the kubeflow pipline
#
# 
|
examples/batch/kubeflow-pipelines-batch/README.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Introduction
#
# For this project, you will act as a data researcher for the **I**nternational **H**eadquarter of **E**mpathy **A**nd **L**ogic (**IHEAL**) . You will be helping them analyze data on GDP and life expectancy from the World Health Organization and the World Bank to support their case that there is a correlation or pattern between the GDP and life expectancy of a country.
#
# To quote the Vice President of Intuition and Systems at **IHEAL:**
#
# > "We know in our hearts and minds that there is an unjust connection between the wealth of a nation, and the life of its people, but we can't get buy in from the people in power without the data to support this."
#
# During this project, you will analyze, prepare, and plot data, and seek to answer questions in a meaningful way.
#
# After you perform analysis, you'll be creating a blog post to share on the **IHEAL** website.
#
# **BIG Question**: Is there a correlation between GDP and life expectancy of a country?
#
# GDP Source:[World Bank](https://data.worldbank.org/indicator/NY.GDP.MKTP.CD)national accounts data, and OECD National Accounts data files.
#
# Life expectancy Data Source: [World Health Organization](vhttp://apps.who.int/gho/data/node.main.688)
#
# ## Step 1 Import Python Modules
# Import the modules that you'll be using in this project:
# - `from matplotlib import pyplot as plt`
# - `import pandas as pd`
# - `import seaborn as sns`
# ## Step 2 Ingest The Data
# To look for connections between GDP and life expectancy you will need to load the datasets into DataFrames so that they can be visualized.
#
# Load **gdp_data_washed.csv** into a DataFrame called `gdp`. Then, quickly inspect the DataFrame using `.head()`.
#
# Hint:<font color=white> Use pd.read_csv()</font>
#
# Load **life_expectancy_data_washed.csv** into a DataFrame called `life`. Then, quickly inspect the DataFrame using `.head()`.
# ## Step 3 Examine The Data
# The datasets are large and it may be easier to view the entire dataset locally on your computer. You can open the CSV files directly from the folder you downloaded for this project.
#
# Let's learn more about our data:
# - GDP stands for **G**ross **D**omestic **P**roduct. GDP is a monetary measure of the market value of all final goods and services produced in a time period.
# - The GDP values are in current US dollars.
# Answer the following questions by inspecting the data in the file **gdp_data_washed.csv**.
# What five countries are represented in the data?
# What years are represented in the data?
# Answer the following questions by inspecting the data in the file **life_expectancy_data_washed.csv**.
# Check that the same five countries are represented in the data.
#
# Which of the five countries represented in the data, do you think would win in a soccer (fútbol) tournament?
# How many rows are there for each country (careful, this one can be a bit tricky)?
# What determines the order that each country's row entries are in. (What order are the 'China' entries in?)
# ## Step 4 Optimize The DataFrame
#
# Look at the column names of the DataFrame `life` using `.head()`.
# What do you notice? The first two column names are one word each, and the third is five whole words long! `Life expectancy at birth (years)` is descriptive, which will be good for labeling the axis, but a little difficult to wrangle for coding the plot itself.
#
# **Optimize The DataFrame Part A:**
#
# Use Pandas to change the name of the last column to `LEABY`.
#
# Hint: Use `.rename()`. [You can read the documentation here.](https://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.rename.html)). </font>
# Run `life.head()` again to check your new column name worked.
# ---
# **Optimize The DataFrame Part B:**
#
# Call `.head()` on the DataFrame `gdp`.
# If you look at the two DataFrames, you can see that the `Year` column is in a different position in each one. Not including the index, `Year` is the second column in `life` and the third column in `gdp`. It will be easier to work with these DataFrames if they are set up as similarly as possible.
#
# Using Pandas rearrange the columns so that `Year` is the second column in `gdp`. Then check your work with `.head()`.
# ## Step 5 Bar Charts To Compare Average
# To take a first high level look at both datasets, create a bar chart for each DataFrame:
#
# A) Creae a bar chart from the data in `gdp` using `Country` on the x-axis and `GDP` on the y-axis.
# B) Create a bar chart using the data in `life` with `Country` on the x-axis and `LEABY` on the y-axis.
# What do you notice about the two bar charts? Do they look similar?
# ## Step 6 Box Plots To Compare Distributions
# Another way to compare two datasets is to visualize the distributions of each and to look for patterns in the shapes.
#
# We have added the code to instantiate a figure with two subplots below. Create a box plot for each DataFrame and set the `ax` argument for one equal to `ax` and the other equal to `ax2`.
# f, (ax, ax2) = plt.subplots(2,1, figsize=(15, 10))
# What do you notice about the two sets of distributions? Do the box plots resemble each other?
# ## Step 7 Point Plots GDP
#
# We want to compare the GDPs of the countries over time, in order to eventually look for correlation between GDP and life expectancy.
#
# Plot the `gdp` DataFrame on a Seaborn `.pointplot()`
#
# Start by setting the variables `f, ax` equal to `plt.subplots()`. This will instantiate a figure and give us access to the axes through the variable name `ax`.
#
# Then, set the size of the figure to 15x6 by passing `figsize=(15, 6)` to `plt.subplots()`.
#
# The syntax for a Seaborn point plot is:
# ```python
# sns.pointplot(x="", y="", hue = "", data=)
# ```
# Create a point plot from the DataFrame `gdp`, using the "Year" column for the `x` argument, the "GDP" column for the `y` argument, and the "Country" column for the `hue` argument. Use the variable `ax` for your plot, like this:
# ```python
# ax = sns.pointplot()
# ```
#
f, ax = plt.subplots(figsize=(15, 6))
ax = sns.pointplot(x="Year", y="GDP", hue = "Country", data=gdp)
# The years across the x-axes are difficult to read because there are so many values. One way to address this issue is to rotate the tick labels. Rotate the x-axes tick labels in your plot using `plt.xticks(rotation=70)`
#
# Also set the label of the y-axis using the following line of code:
# ax.set(ylabel="GDP in Trillions of U.S. Dollars")
# +
f, ax = plt.subplots(figsize=(15, 6))
ax = sns.pointplot(x="Year", y="GDP", hue = "Country", data=gdp)
plt.xticks(rotation=70)
ax.set(ylabel="GDP in Trillions of U.S. Dollars")
# -
# Because the values of GDP are so high, the tick labels on the y-axis can be a little confusing. You can reformat the values to be in trillions using the code we've put in the cell for you below. Run the code to see the difference.
# +
from matplotlib.ticker import FuncFormatter
def trillions(x, pos):
'The two args are the value and tick position'
return '$%1.1fT' % (x*1e-12)
formatter = FuncFormatter(trillions)
f, ax = plt.subplots(figsize=(15, 6))
ax = sns.pointplot(x="Year", y="GDP", hue = "Country", data=gdp)
plt.xticks(rotation=70)
ax.yaxis.set_major_formatter(formatter)
ax.set(ylabel="GDP in Trillions of U.S. Dollars")
# -
# What are your first impressions looking at the visualized data?
#
# - Which countries' line changes the most?
# - What years are there the biggest changes in the data?
# - Which country has had the least change in GDP over time?
# - Can you think of any reasons that the data looks like this for particular countries?
# ## Step 8 Point Plots Life Expectancy
# To compare GDP and life expectancy over time we also need to plot the life expectancy data.
# Start again by setting the variables `f, ax ` equal to `plt.subplots()`. Set the size of the figure to 12x6.
#
# Create a point plot from the DataFrame `life`, using the "Year" column for the `x` argument, the "LEABY" column for the `y` argument, and the "Country" column for the `hue` argument. Use the variable `ax` for your plot, like this:
# ```python
# ax = sns.pointplot()
# ```
#
# Set the y-axis label back to "Life expectancy at birth (years)" using `ax.set()`.
f, ax = plt.subplots(figsize=(12, 6))
ax = sns.pointplot(x="Year", y="LEABY", hue = "Country", data=life)
ax.set(ylabel="Life expectancy at birth (years)")
# What are your first impressions looking at the visualized data?
#
# - Which countries' line changes the most?
# - What years are there the biggest changes in the data?
# - Which country has had the least change in life expectancy over time?
# - Can you think of any reasons that the data looks like this for particular countries?
# ## Step 9 Combined Data
# To create a visualization that will make it easier to see the possible correlation between GDP and life expectancy, you can plot each set of data on its own subplot, on a shared figure.
#
# To make this easier, we have created a CSV file that has columns for both GDP and life expectancy for the years 2000-2015.
#
# Use `pd.read_csv()` to import the file **combined_data.csv** to a variable named `combined_data`. Then, check the new DataFrame using `.head()`.
#
# read in combined csv
combined_data = pd.read_csv("combined_data.csv")
# head
combined_data.head()
# ## Step 10 Comparison Over Time
# Create a figure with two subplots, divided into 2 rows and 1 column.
#
# - We have set up the code for you on line 1 in the cell below. Complete the figure by passing the following arguments to `plt.subplots()`
# - `2`-- the number of rows for the subplots
# - `1` -- the number of columns for the subplots
# - `figsize=(15, 15)` -- the size of the figure
#
#
# Use the `combined_data` DataFrame to create point plots for the GDP and Life expectancy over the same 16 years.
#
# The code for `pointplot()` will be the same as you have previously used in this project, with the addition of the `ax` argument which can be added after the `data` argument.
# - Set the `ax` argument for one `pointplot()` to `ax1`, and the other to `ax2`.
# +
from matplotlib.ticker import FuncFormatter
def trillions(x, pos):
'The two args are the value and tick position'
return '$%1.1fT' % (x*1e-12)
formatter = FuncFormatter(trillions)
sns.set_style("dark")
f, (ax1, ax2) = plt.subplots(2,1,figsize=(15, 15) )
sns.set_palette("Dark2")
ax1 = sns.pointplot(x="Year", y="LEABY", hue = "Country", data=combined_data, ax=ax1)
ax2 = sns.pointplot(x="Year", y="GDP", hue = "Country", data=combined_data, ax=ax2)
ax2.yaxis.set_major_formatter(formatter)
ax1.set(ylabel="Life expectancy at birth (years)")
ax2.set(ylabel="GDP in Trillions of U.S. Dollars")
ax2.set_title("GDP in Trillions of Dollars Over Time")
ax1.set_title("Life Expectancy In Years Over Time")
# -
# You may have to look closely to see the GDP difference between Zimbabwe and the Dominican Republic, as they are very similar over this time period.
# Which countries have the highest and lowest GDP?
# Which countries have the highest and lowest life expectancy?
# ## Step 11 Researching Data Context
# Based on the visualization, choose one part the data to research a little further so you can add some real world context to the visualization. You can choose anything you like, or use the example question below.
#
# What happened in China between 1991-2016 that increased the GDP so drastically?
# ## Step 12 Create Blog Post
# Use the content you have created in this Jupyter notebook to create a blog post on this data.
# ## Bonus/Optional Probably Delete
x = combined_data["LEABY"]
z = combined_data["GDP"]
y = combined_data["Year"]
# sns.lvplot(data=combined_data, scale="linear", palette="mako")
# ax = sns.kdeplot(z, y, shade=True, cmap="mako")
# ax2 = sns.kdeplot(z, shade=True, color="b")
|
Notebooks/DV-Portfolio-Life-Expectancy-and-GDP/.ipynb_checkpoints/global_data_capstone-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Test Recurrent Policy with Extreme Parameter Variation
# +
import numpy as np
import os,sys
sys.path.append('../../../../RL_lib/Agents')
sys.path.append('../../../../RL_lib/Policies/PPO')
sys.path.append('../../../../RL_lib/Policies/Common')
sys.path.append('../../../../RL_lib/Utils')
sys.path.append('../../../Env')
sys.path.append('../../../Imaging')
# %load_ext autoreload
# %load_ext autoreload
# %autoreload 2
# %matplotlib nbagg
import os
print(os.getcwd())
# + language="html"
# <style>
# .output_wrapper, .output {
# height:auto !important;
# max-height:1000px; /* your desired max-height here */
# }
# .output_scroll {
# box-shadow:none !important;
# webkit-box-shadow:none !important;
# }
# </style>
# -
# # Optimize Policy
# +
from lander_model import Lander_model
from ic_gen import Landing_icgen
import rl_utils
logger = rl_utils.Logger()
# -
from render_traj_seeker import render_traj
traj = np.load('traj_1.npy').item()
render_traj(traj)
import plot_rl_stats2
fname = "optimize_WATTVW_FOV-AR=5_history"
rl_stats = plot_rl_stats2.RL_stats(None,logger)
rl_stats.load_history(fname)
rl_stats.plot_rewards()
rl_stats.plot_rf()
rl_stats.plot_vf()
print(rl_stats.history.keys())
print(env.rl_stats.history.keys())
|
Experiments/Extended/Test_HF/plot_history.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
import struct
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import keras
from keras.models import Sequential
from keras.layers import Dense, Dropout, Flatten
from keras.layers import Conv2D, MaxPooling2D
def load_mnist(dataset="training", path="."):
if dataset is "training":
fname_img = os.path.join(path, 'train-images.idx3-ubyte')
fname_lbl = os.path.join(path, 'train-labels.idx1-ubyte')
elif dataset is "testing":
fname_img = os.path.join(path, 't10k-images.idx3-ubyte')
fname_lbl = os.path.join(path, 't10k-labels.idx1-ubyte')
else:
raise ValueError("dataset must be 'testing' or 'training'")
# Load everything in some numpy arrays
with open(fname_lbl, 'rb') as flbl:
magic, num = struct.unpack(">II", flbl.read(8))
lbl = keras.utils.to_categorical(np.fromfile(flbl, dtype=np.int8), 10)
with open(fname_img, 'rb') as fimg:
magic, num, rows, cols = struct.unpack(">IIII", fimg.read(16))
#img = np.fromfile(fimg, dtype=np.uint8).reshape(len(lbl), rows, cols).astype('float32') / 255
img = np.fromfile(fimg, dtype=np.uint8).reshape(len(lbl), rows*cols).astype('float32') / 255
return img, lbl
def simple_dnn(batch_size, epochs):
np.random.seed(7)
data_path = os.path.join('.', 'data')
x_train, y_train = load_mnist(path=data_path)
x_test, y_test = load_mnist(dataset='testing', path=data_path)
model = Sequential()
model.add(Dense(512, activation='relu', input_shape=(784,)))
model.add(Dropout(0.2))
model.add(Dense(512, activation='relu'))
model.add(Dropout(0.5))
model.add(Dense(10, activation='softmax'))
#model.summary()
model.compile(loss=keras.losses.categorical_crossentropy,
optimizer=keras.optimizers.Adadelta(),
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
simple_dnn(128, 10)
def cnn_dnn(batch_size, epochs):
np.random.seed(7)
data_path = os.path.join('.', 'data')
x_train, y_train = load_mnist(path=data_path)
x_test, y_test = load_mnist(dataset='testing', path=data_path)
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1)
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
model = Sequential()
model.add(Conv2D(32, kernel_size=(5, 5),
activation='relu',
input_shape=(28,28,1)))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(10, activation='softmax'))
#model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
cnn_dnn(128, 5)
def cnn_deeper(batch_size, epochs):
np.random.seed(7)
data_path = os.path.join('.', 'data')
x_train, y_train = load_mnist(path=data_path)
x_test, y_test = load_mnist(dataset='testing', path=data_path)
x_train = x_train.reshape(x_train.shape[0], 28, 28, 1)
x_test = x_test.reshape(x_test.shape[0], 28, 28, 1)
model = Sequential()
model.add(Conv2D(32, kernel_size=(5, 5),
activation='relu',
input_shape=(28,28,1)))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(64, (3, 3), activation='relu'))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Dropout(0.2))
model.add(Flatten())
model.add(Dense(128, activation='relu'))
model.add(Dropout(0.2))
model.add(Dense(64, activation='relu'))
model.add(Dense(10, activation='softmax'))
#model.summary()
model.compile(loss='categorical_crossentropy', optimizer='adam',
metrics=['accuracy'])
model.fit(x_train, y_train,
batch_size=batch_size,
epochs=epochs,
verbose=1,
validation_data=(x_test, y_test))
score = model.evaluate(x_test, y_test, verbose=0)
print('Test loss:', score[0])
print('Test accuracy:', score[1])
cnn_deeper(128,5)
|
ML/kerasTest.ipynb
|
# + [markdown] colab_type="text" id="9TV7IYeqifSv"
# ##### Copyright 2018 The TensorFlow Authors. [Licensed under the Apache License, Version 2.0](#scrollTo=ByZjmtFgB_Y5).
# + colab={} colab_type="code" id="tRIJp_4m_Afz"
// #@title Licensed under the Apache License, Version 2.0 (the "License"); { display-mode: "form" }
// Licensed under the Apache License, Version 2.0 (the "License");
// you may not use this file except in compliance with the License.
// You may obtain a copy of the License at
//
// https://www.apache.org/licenses/LICENSE-2.0
//
// Unless required by applicable law or agreed to in writing, software
// distributed under the License is distributed on an "AS IS" BASIS,
// WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
// See the License for the specific language governing permissions and
// limitations under the License.
# + [markdown] colab_type="text" id="sI1ZtrdiA4aY"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/swift/tutorials/python_interoperability"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/swift/blob/master/docs/site/tutorials/python_interoperability.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/swift/blob/master/docs/site/tutorials/python_interoperability.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="8sa42_NblqRE"
# # Python interoperability
#
# Swift For TensorFlow supports Python interoperability.
#
# You can import Python modules from Swift, call Python functions, and convert values between Swift and Python.
# + colab={} colab_type="code" id="kZRlD4utdPuX"
import Python
print(Python.version)
# + [markdown] colab_type="text" id="W7MpNcIwIIy8"
# ## Setting the Python version
# + [markdown] colab_type="text" id="lM9dRji7IIy8"
# By default, when you `import Python`, Swift searches system library paths for the newest version of Python installed.
# To use a specific Python installation, set the `PYTHON_LIBRARY` environment variable to the `libpython` shared library provided by the installation. For example:
#
# `export PYTHON_LIBRARY="~/anaconda3/lib/libpython3.7m.so"`
#
# The exact filename will differ across Python environments and platforms.
# + [markdown] colab_type="text" id="eoyLeSQVIIy9"
# Alternatively, you can set the `PYTHON_VERSION` environment variable, which instructs Swift to search system library paths for a matching Python version. Note that `PYTHON_LIBRARY` takes precedence over `PYTHON_VERSION`.
#
# In code, you can also call the `PythonLibrary.useVersion` function, which is equivalent to setting `PYTHON_VERSION`.
# + colab={} colab_type="code" id="FCMWR11NIIy-"
// PythonLibrary.useVersion(2)
// PythonLibrary.useVersion(3, 7)
# + [markdown] colab_type="text" id="HrlMNOinIIy_"
# __Note: you should run `PythonLibrary.useVersion` right after `import Python`, before calling any Python code. It cannot be used to dynamically switch Python versions.__
# + [markdown] colab_type="text" id="mIbIOW0HIIzA"
# Set `PYTHON_LOADER_LOGGING=1` to see [debug output for Python library loading](https://github.com/apple/swift/pull/20674#discussion_r235207008).
# + [markdown] colab_type="text" id="rU0WY_sJodio"
# ## Basics
#
# In Swift, `PythonObject` represents an object from Python.
# All Python APIs use and return `PythonObject` instances.
#
# Basic types in Swift (like numbers and arrays) are convertible to `PythonObject`. In some cases (for literals and functions taking `PythonConvertible` arguments), conversion happens implicitly. To explicitly cast a Swift value to `PythonObject`, use the `PythonObject` initializer.
#
# `PythonObject` defines many standard operations, including numeric operations, indexing, and iteration.
# + colab={} colab_type="code" id="kqXILiXhq-iM"
// Convert standard Swift types to Python.
let pythonInt: PythonObject = 1
let pythonFloat: PythonObject = 3.0
let pythonString: PythonObject = "Hello Python!"
let pythonRange: PythonObject = PythonObject(5..<10)
let pythonArray: PythonObject = [1, 2, 3, 4]
let pythonDict: PythonObject = ["foo": [0], "bar": [1, 2, 3]]
// Perform standard operations on Python objects.
print(pythonInt + pythonFloat)
print(pythonString[0..<6])
print(pythonRange)
print(pythonArray[2])
print(pythonDict["bar"])
# + colab={} colab_type="code" id="fEAEyUExXT3I"
// Convert Python objects back to Swift.
let int = Int(pythonInt)!
let float = Float(pythonFloat)!
let string = String(pythonString)!
let range = Range<Int>(pythonRange)!
let array: [Int] = Array(pythonArray)!
let dict: [String: [Int]] = Dictionary(pythonDict)!
// Perform standard operations.
// Outputs are the same as Python!
print(Float(int) + float)
print(string.prefix(6))
print(range)
print(array[2])
print(dict["bar"]!)
# + [markdown] colab_type="text" id="1pMewsl0VgnJ"
# `PythonObject` defines conformances to many standard Swift protocols:
# * `Equatable`
# * `Comparable`
# * `Hashable`
# * `SignedNumeric`
# * `Strideable`
# * `MutableCollection`
# * All of the `ExpressibleBy_Literal` protocols
#
# Note that these conformances are not type-safe: crashes will occur if you attempt to use protocol functionality from an incompatible `PythonObject` instance.
# + colab={} colab_type="code" id="W9bUsiOxVf_v"
let one: PythonObject = 1
print(one == one)
print(one < one)
print(one + one)
let array: PythonObject = [1, 2, 3]
for (i, x) in array.enumerated() {
print(i, x)
}
# + [markdown] colab_type="text" id="w3lmTRCWT5sS"
# To convert tuples from Python to Swift, you must statically know the arity of the tuple.
#
# Call one of the following instance methods:
# - `PythonObject.tuple2`
# - `PythonObject.tuple3`
# - `PythonObject.tuple4`
# + colab={} colab_type="code" id="fQ0HEX89T4mW"
let pythonTuple = Python.tuple([1, 2, 3])
print(pythonTuple, Python.len(pythonTuple))
// Convert to Swift.
let tuple = pythonTuple.tuple3
print(tuple)
# + [markdown] colab_type="text" id="Te7sNNx9c_am"
# ## Python builtins
#
# Access Python builtins via the global `Python` interface.
# + colab={} colab_type="code" id="jpcOByipc75O"
// `Python.builtins` is a dictionary of all Python builtins.
_ = Python.builtins
// Try some Python builtins.
print(Python.type(1))
print(Python.len([1, 2, 3]))
print(Python.sum([1, 2, 3]))
# + [markdown] colab_type="text" id="H2wwUL1tY3JX"
# ## Importing Python modules
#
# Use `Python.import` to import a Python module. It works like the `import` keyword in `Python`.
# + colab={} colab_type="code" id="XrZee8n3Y17_"
let np = Python.import("numpy")
print(np)
let zeros = np.ones([2, 3])
print(zeros)
# + [markdown] colab_type="text" id="hQvza3dUXlr0"
# Use the throwing function `Python.attemptImport` to perform safe importing.
# + colab={} colab_type="code" id="QD-uQGuaXhrM"
let maybeModule = try? Python.attemptImport("nonexistent_module")
print(maybeModule)
# + [markdown] colab_type="text" id="Qej_Z6V3mZnG"
# ## Conversion with `numpy.ndarray`
#
# The following Swift types can be converted to and from `numpy.ndarray`:
# - `Array<Element>`
# - `ShapedArray<Scalar>`
# - `Tensor<Scalar>`
#
# Conversion succeeds only if the `dtype` of the `numpy.ndarray` is compatible with the `Element` or `Scalar` generic parameter type.
#
# For `Array`, conversion from `numpy` succeeds only if the `numpy.ndarray` is 1-D.
# + colab={} colab_type="code" id="hPvKgZBeDQ1p"
import TensorFlow
let numpyArray = np.ones([4], dtype: np.float32)
print("Swift type:", type(of: numpyArray))
print("Python type:", Python.type(numpyArray))
print(numpyArray.shape)
# + colab={} colab_type="code" id="ZuDgZ5cBS3Uk"
// Examples of converting `numpy.ndarray` to Swift types.
let array: [Float] = Array(numpy: numpyArray)!
let shapedArray = ShapedArray<Float>(numpy: numpyArray)!
let tensor = Tensor<Float>(numpy: numpyArray)!
// Examples of converting Swift types to `numpy.ndarray`.
print(array.makeNumpyArray())
print(shapedArray.makeNumpyArray())
print(tensor.makeNumpyArray())
// Examples with different dtypes.
let doubleArray: [Double] = Array(numpy: np.ones([3], dtype: np.float))!
let intTensor = Tensor<Int32>(numpy: np.ones([2, 3], dtype: np.int32))!
# + [markdown] colab_type="text" id="8EQFZZ5iafwh"
# ## Displaying images
#
# You can display images in-line using `matplotlib`, just like in Python notebooks.
# + colab={} colab_type="code" id="vjQ7Rd3_IXuX"
// This cell is here to display plots inside a Jupyter Notebook.
// Do not copy it into another environment.
%include "EnableIPythonDisplay.swift"
IPythonDisplay.shell.enable_matplotlib("inline")
# + colab={} colab_type="code" id="jUzsa2cxafQV"
let np = Python.import("numpy")
let plt = Python.import("matplotlib.pyplot")
let time = np.arange(0, 10, 0.01)
let amplitude = np.exp(-0.1 * time)
let position = amplitude * np.sin(3 * time)
plt.figure(figsize: [15, 10])
plt.plot(time, position)
plt.plot(time, amplitude)
plt.plot(time, -amplitude)
plt.xlabel("Time (s)")
plt.ylabel("Position (m)")
plt.title("Oscillations")
plt.show()
|
docs/site/tutorials/python_interoperability.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="Tb8Dlc3cvb-g"
# # wav2vec-u CV-sv - prepare text
#
# > "Running prepare_text.sh for wav2vec-u on Common Voice Swedish"
#
# - toc: false
# - branch: master
# - badges: false
# - hidden: true
# - categories: [kaggle, wav2vec-u]
# + [markdown] id="v8L8t1YLvm0n"
# Original [here](https://www.kaggle.com/jimregan/wav2vec-u-cv-swedish-text-prep)
# + _cell_guid="b1076dfc-b9ad-4769-8c92-a6c4dae69d19" _uuid="8f2839f25d086af736a60e9eeb907d3b93b6e0e5" papermill={"duration": 0.032818, "end_time": "2021-05-30T15:46:53.236930", "exception": false, "start_time": "2021-05-30T15:46:53.204112", "status": "completed"} tags=[] id="welcome-offer" outputId="5692778d-9602-46d7-e136-71f1bd1cfaee"
# %cd /opt
# + papermill={"duration": 38.419926, "end_time": "2021-05-30T15:47:31.683409", "exception": false, "start_time": "2021-05-30T15:46:53.263483", "status": "completed"} tags=[] id="painted-roman"
# %%capture
# !tar xvf /kaggle/input/extract-prebuilt-kaldi-from-docker/kaldi.tar
# + papermill={"duration": 0.026387, "end_time": "2021-05-30T15:47:31.727544", "exception": false, "start_time": "2021-05-30T15:47:31.701157", "status": "completed"} tags=[] id="danish-cooking" outputId="a0b689f8-97dc-460f-a420-f2de6ec16135"
# %cd /tmp
# + papermill={"duration": 8.062336, "end_time": "2021-05-30T15:47:39.808700", "exception": false, "start_time": "2021-05-30T15:47:31.746364", "status": "completed"} tags=[] id="driven-comparative"
# !git clone https://github.com/pytorch/fairseq/
# + papermill={"duration": 11.517856, "end_time": "2021-05-30T15:47:51.349544", "exception": false, "start_time": "2021-05-30T15:47:39.831688", "status": "completed"} tags=[] id="copyrighted-diameter"
# %%capture
# !pip install phonemizer
# + papermill={"duration": 47.179551, "end_time": "2021-05-30T15:48:38.552240", "exception": false, "start_time": "2021-05-30T15:47:51.372689", "status": "completed"} tags=[] id="described-retirement"
# %%capture
# !pip install git+https://github.com/pytorch/fairseq/
# + papermill={"duration": 4.993642, "end_time": "2021-05-30T15:48:43.580968", "exception": false, "start_time": "2021-05-30T15:48:38.587326", "status": "completed"} tags=[] id="robust-soldier"
# %%capture
# !apt-get -y install espeak
# + papermill={"duration": 1.949371, "end_time": "2021-05-30T15:48:45.565530", "exception": false, "start_time": "2021-05-30T15:48:43.616159", "status": "completed"} tags=[] id="auburn-penetration"
# !git clone https://github.com/kpu/kenlm
# + papermill={"duration": 3.278477, "end_time": "2021-05-30T15:48:48.872968", "exception": false, "start_time": "2021-05-30T15:48:45.594491", "status": "completed"} tags=[] id="equal-portfolio"
# %%capture
# !apt-get -y install libeigen3-dev liblzma-dev zlib1g-dev libbz2-dev
# + papermill={"duration": 53.050345, "end_time": "2021-05-30T15:49:41.951862", "exception": false, "start_time": "2021-05-30T15:48:48.901517", "status": "completed"} tags=[] id="cooperative-louis"
# %%capture
# %cd kenlm
# !mkdir build
# %cd build
# !cmake ..
# !make -j 4
# %cd /tmp
# + papermill={"duration": 0.049347, "end_time": "2021-05-30T15:49:42.044261", "exception": false, "start_time": "2021-05-30T15:49:41.994914", "status": "completed"} tags=[] id="announced-wealth"
import os
os.environ['PATH'] = f"{os.environ['PATH']}:/tmp/kenlm/build/bin/"
os.environ['FAIRSEQ_ROOT'] = '/tmp/fairseq'
# + papermill={"duration": 0.36141, "end_time": "2021-05-30T15:49:42.434694", "exception": false, "start_time": "2021-05-30T15:49:42.073284", "status": "completed"} tags=[] id="southwest-nelson"
# !cat /kaggle/input/wav2vec-u-cv-swedish-audio/*.wrd | grep -v '^$' | sort| uniq > /kaggle/working/sentences.txt
# + papermill={"duration": 0.038432, "end_time": "2021-05-30T15:49:42.502078", "exception": false, "start_time": "2021-05-30T15:49:42.463646", "status": "completed"} tags=[] id="searching-bleeding" outputId="1d569a3c-baa2-4e4c-e801-e36c2c78d88f"
# %cd fairseq/examples/wav2vec/unsupervised
# + papermill={"duration": 3.536067, "end_time": "2021-05-30T15:49:46.081855", "exception": false, "start_time": "2021-05-30T15:49:42.545788", "status": "completed"} tags=[] id="traditional-thirty"
# %%capture
# !apt-get -y install zsh
# + papermill={"duration": 0.3284, "end_time": "2021-05-30T15:49:46.453999", "exception": false, "start_time": "2021-05-30T15:49:46.125599", "status": "completed"} tags=[] id="judicial-yugoslavia"
# !mkdir /kaggle/working/preppedtext
# + papermill={"duration": 0.037329, "end_time": "2021-05-30T15:49:46.520482", "exception": false, "start_time": "2021-05-30T15:49:46.483153", "status": "completed"} tags=[] id="engaging-princeton" outputId="42ba9f3b-a6ee-4e73-dc07-6ad878441534"
# %cd scripts
# + [markdown] papermill={"duration": 0.042909, "end_time": "2021-05-30T15:49:46.606603", "exception": false, "start_time": "2021-05-30T15:49:46.563694", "status": "completed"} tags=[] id="incorrect-grill"
# The next part requires a FastText language id model; I don't know where the 187 language model comes from, but there is a model for 176 languages [here](https://fasttext.cc/docs/en/language-identification.html#content)
# + papermill={"duration": 13.015984, "end_time": "2021-05-30T15:49:59.665834", "exception": false, "start_time": "2021-05-30T15:49:46.649850", "status": "completed"} tags=[] id="discrete-columbia"
# !wget https://dl.fbaipublicfiles.com/fasttext/supervised-models/lid.176.bin
# + papermill={"duration": 0.585651, "end_time": "2021-05-30T15:50:00.292077", "exception": false, "start_time": "2021-05-30T15:49:59.706426", "status": "completed"} tags=[] id="alleged-willow"
# !cat normalize_and_filter_text.py|sed -e 's/187/176/' > tmp
# !mv tmp normalize_and_filter_text.py
# + papermill={"duration": 0.048158, "end_time": "2021-05-30T15:50:00.380148", "exception": false, "start_time": "2021-05-30T15:50:00.331990", "status": "completed"} tags=[] id="dedicated-alarm"
# Needed to see what's going wrong
os.environ['HYDRA_FULL_ERROR'] = '1'
# + papermill={"duration": 0.045137, "end_time": "2021-05-30T15:50:00.464563", "exception": false, "start_time": "2021-05-30T15:50:00.419426", "status": "completed"} tags=[] id="strategic-trading"
import os
os.environ['LD_LIBRARY_PATH'] = '/opt/conda/lib:/opt/kaldi/tools/openfst-1.6.7/lib:/opt/kaldi/src/lib'
# + [markdown] papermill={"duration": 0.040367, "end_time": "2021-05-30T15:50:00.545473", "exception": false, "start_time": "2021-05-30T15:50:00.505106", "status": "completed"} tags=[] id="boolean-aging"
# There are two lines with missing variables in `prepare_text.sh` - [pull request](https://github.com/pytorch/fairseq/pull/3569) - so replace the file.
#
# While I'm replacing the file: most of the first part of the script is unneeded, as I already have a phonetic dictionary, so I'm using that instead.
#
# With the calls of the `preprocess.py` script, make sure to check the threshold: there's a divide by zero if the threshold is set too high.
# + [markdown] papermill={"duration": 0.04037, "end_time": "2021-05-30T15:50:00.627225", "exception": false, "start_time": "2021-05-30T15:50:00.586855", "status": "completed"} tags=[] id="sound-manner"
# Config options for `kaldi_initializer.py`
#
# - `in_labels`: a naming component, for the Kaldi lexicons/fsts (required)
# - `wav2letter_lexicon`: path to wav2letter lexicon
# - `out_labels`: a naming component, for the Kaldi lexicons/fsts: set to `in_label` if missing
# - `kaldi_root`: path to Kaldi: `/opt/kaldi` for my kaggle image
# - `fst_dir`: path where generated fsts will be saved
# - `data_dir`: path to phones data
# - `lm_arpa`: path to the lm in ARPA format
# - `blank_symbol`: CTC blank symbol (`<s>` here)
# - `silence_symbol`: Kaldi symbol for silence (`<SIL>` is set for two of the scripts)
#
# A config file needs to exist for this, even though the options set in it seem to be ignored.
# + papermill={"duration": 0.32394, "end_time": "2021-05-30T15:50:00.990647", "exception": false, "start_time": "2021-05-30T15:50:00.666707", "status": "completed"} tags=[] id="adaptive-calculation"
# !mkdir /tmp/fairseq/examples/speech_recognition/kaldi/config/
# + papermill={"duration": 0.050151, "end_time": "2021-05-30T15:50:01.081969", "exception": false, "start_time": "2021-05-30T15:50:01.031818", "status": "completed"} tags=[] id="straight-morocco" outputId="c4ccae41-fa1d-4c5c-a4b6-959d88e71b93"
# %%writefile /tmp/fairseq/examples/speech_recognition/kaldi/config/config.yaml
kaldi_root: "/opt/kaldi"
# + _kg_hide-input=true papermill={"duration": 0.052733, "end_time": "2021-05-30T15:50:01.174168", "exception": false, "start_time": "2021-05-30T15:50:01.121435", "status": "completed"} tags=[] id="organizational-membership" outputId="d1b311a6-a9d7-4e48-c811-4ee84ef83d89"
# %%writefile prepare_text.sh
# #!/usr/bin/env zsh
# Copyright (c) Facebook, Inc. and its affiliates.
#
# This source code is licensed under the MIT license found in the
# LICENSE file in the root directory of this source tree.
lg=$1
text_path=$2
target_dir=$3
#ph_lg=${lg:l}
#if test "$lg" = 'fr'; then
# ph_lg='fr-fr'
#elif test "$lg" = 'en'; then
# ph_lg='en-us'
#elif test "$lg" = 'pt'; then
# ph_lg='pt-br'
#fi
ph_lg="sv"
# echo $lg
# echo $ph_lg
# echo $text_path
# echo $target_dir
# mkdir -p $target_dir
#python normalize_and_filter_text.py --lang $lg < $text_path | grep -v '\-\-\-' >! $target_dir/lm.upper.lid.txt
#python $FAIRSEQ_ROOT/fairseq_cli/preprocess.py --dataset-impl mmap --trainpref $target_dir/lm.upper.lid.txt --only-source --destdir $target_dir --thresholdsrc 2 --padding-factor 1 --dict-only
#cut -f1 -d' ' $target_dir/dict.txt | grep -v -x '[[:punct:]]*' | grep -Pv '\d\d\d\d\d+' >! $target_dir/words.txt
# cp /kaggle/input/wav2vec-u-cv-swedish-audio/train.wrd $target_dir/lm.upper.lid.txt
cut -f1 -d' ' /kaggle/input/wav2vec-u-cv-swedish-audio/dict.train >! $target_dir/words.txt
#one=$(echo "1" | PHONEMIZER_ESPEAK_PATH=$(which espeak) phonemize -p ' ' -w '' -l $ph_lg --language-switch remove-flags)
#sed 's/$/ 1/' $target_dir/words.txt | PHONEMIZER_ESPEAK_PATH=$(which espeak) phonemize -o $target_dir/phones.txt -p ' ' -w '' -l $ph_lg -j 70 --language-switch remove-flags
cut -f2- -d' ' /kaggle/input/wav2vec-u-cv-swedish-audio/dict.train >! $target_dir/phones.txt
# #echo "one is ${one}"
#sed -i "s/${one}$//" $target_dir/phones.txt
#paste $target_dir/words.txt $target_dir/phones.txt >! $target_dir/lexicon.lst
# cp /kaggle/input/wav2vec-u-cv-swedish-audio/dict.train $target_dir/lexicon.lst
#python $FAIRSEQ_ROOT/fairseq_cli/preprocess.py --dataset-impl mmap --trainpref $target_dir/phones.txt --only-source --destdir $target_dir/phones --thresholdsrc 1000 --padding-factor 1 --dict-only
python $FAIRSEQ_ROOT/fairseq_cli/preprocess.py --dataset-impl mmap --trainpref $target_dir/phones.txt --only-source --destdir $target_dir/phones --thresholdsrc 2 --padding-factor 1 --dict-only
python filter_lexicon.py -d $target_dir/phones/dict.txt < $target_dir/lexicon.lst >! $target_dir/lexicon_filtered.lst
python phonemize_with_sil.py -s 0.25 --surround --lexicon $target_dir/lexicon_filtered.lst < $target_dir/lm.upper.lid.txt >! $target_dir/phones/lm.phones.filtered.txt
# cp $target_dir/phones/dict.txt $target_dir/phones/dict.phn.txt
# echo "<SIL> 0" >> $target_dir/phones/dict.phn.txt
python $FAIRSEQ_ROOT/fairseq_cli/preprocess.py --dataset-impl mmap --trainpref $target_dir/phones/lm.phones.filtered.txt --workers 70 --only-source --destdir $target_dir/phones --srcdict $target_dir/phones/dict.phn.txt
lmplz -o 4 < $target_dir/lm.upper.lid.txt --discount_fallback --prune 0 0 0 3 >! $target_dir/kenlm.wrd.o40003.arpa
build_binary $target_dir/kenlm.wrd.o40003.arpa $target_dir/kenlm.wrd.o40003.bin
lg=$lg python $FAIRSEQ_ROOT/examples/speech_recognition/kaldi/kaldi_initializer.py fst_dir=$target_dir/fst/phn_to_words_sil lm_arpa=$target_dir/kenlm.wrd.o40003.arpa wav2letter_lexicon=$target_dir/lexicon_filtered.lst data_dir=$target_dir/phones "blank_symbol='<SIL>'" "in_labels='phn'" "kaldi_root='/opt/kaldi'"
lg=$lg python $FAIRSEQ_ROOT/examples/speech_recognition/kaldi/kaldi_initializer.py fst_dir=$target_dir/fst/phn_to_words lm_arpa=$target_dir/kenlm.wrd.o40003.arpa wav2letter_lexicon=$target_dir/lexicon_filtered.lst data_dir=$target_dir/phones "in_labels='phn'" "kaldi_root='/opt/kaldi'"
lmplz -o 4 < $target_dir/phones/lm.phones.filtered.txt --discount_fallback >! $target_dir/phones/lm.phones.filtered.04.arpa
build_binary -s $target_dir/phones/lm.phones.filtered.04.arpa $target_dir/phones/lm.phones.filtered.04.bin
lmplz -o 6 < $target_dir/phones/lm.phones.filtered.txt --discount_fallback >! $target_dir/phones/lm.phones.filtered.06.arpa
build_binary -s $target_dir/phones/lm.phones.filtered.06.arpa $target_dir/phones/lm.phones.filtered.06.bin
lg=$lg python $FAIRSEQ_ROOT/examples/speech_recognition/kaldi/kaldi_initializer.py fst_dir=$target_dir/fst/phn_to_phn_sil lm_arpa=$target_dir/phones/lm.phones.filtered.06.arpa data_dir=$target_dir/phones "blank_symbol='<SIL>'" "in_labels='phn'" "kaldi_root='/opt/kaldi'"
# + [markdown] papermill={"duration": 0.040205, "end_time": "2021-05-30T15:50:01.255805", "exception": false, "start_time": "2021-05-30T15:50:01.215600", "status": "completed"} tags=[] id="dynamic-walker"
# `add-self-loop-simple.cc` attempts to use `std::endl` with `KALDI_LOG`, which doesn't work, so rewrite that (I'm not sure if this actually prevents anything from working, but it is really distracting).
# + _kg_hide-input=true papermill={"duration": 0.048429, "end_time": "2021-05-30T15:50:01.344658", "exception": false, "start_time": "2021-05-30T15:50:01.296229", "status": "completed"} tags=[] id="whole-upper" outputId="<PASSWORD>"
# %%writefile /tmp/fairseq/examples/speech_recognition/kaldi/add-self-loop-simple.cc
/*
* Copyright (c) Facebook, Inc. and its affiliates.
*
* This source code is licensed under the MIT license found in the
* LICENSE file in the root directory of this source tree.
*/
#include <iostream>
#include "fstext/fstext-lib.h" // @manual
#include "util/common-utils.h" // @manual
/*
* This program is to modify a FST without self-loop by:
* for each incoming arc with non-eps input symbol, add a self-loop arc
* with that non-eps symbol as input and eps as output.
*
* This is to make sure the resultant FST can do deduplication for repeated
* symbols, which is very common in acoustic model
*
*/
namespace {
int32 AddSelfLoopsSimple(fst::StdVectorFst* fst) {
typedef fst::MutableArcIterator<fst::StdVectorFst> IterType;
int32 num_states_before = fst->NumStates();
fst::MakePrecedingInputSymbolsSame(false, fst);
int32 num_states_after = fst->NumStates();
KALDI_LOG << "There are " << num_states_before
<< " states in the original FST; "
<< " after MakePrecedingInputSymbolsSame, there are "
<< num_states_after << " states ";
auto weight_one = fst::StdArc::Weight::One();
int32 num_arc_added = 0;
fst::StdArc self_loop_arc;
self_loop_arc.weight = weight_one;
int32 num_states = fst->NumStates();
std::vector<std::set<int32>> incoming_non_eps_label_per_state(num_states);
for (int32 state = 0; state < num_states; state++) {
for (IterType aiter(fst, state); !aiter.Done(); aiter.Next()) {
fst::StdArc arc(aiter.Value());
if (arc.ilabel != 0) {
incoming_non_eps_label_per_state[arc.nextstate].insert(arc.ilabel);
}
}
}
for (int32 state = 0; state < num_states; state++) {
if (!incoming_non_eps_label_per_state[state].empty()) {
auto& ilabel_set = incoming_non_eps_label_per_state[state];
for (auto it = ilabel_set.begin(); it != ilabel_set.end(); it++) {
self_loop_arc.ilabel = *it;
self_loop_arc.olabel = 0;
self_loop_arc.nextstate = state;
fst->AddArc(state, self_loop_arc);
num_arc_added++;
}
}
}
return num_arc_added;
}
void print_usage() {
std::cout << "add-self-loop-simple usage:\n"
"\tadd-self-loop-simple <in-fst> <out-fst> \n";
}
} // namespace
int main(int argc, char** argv) {
if (argc != 3) {
print_usage();
exit(1);
}
auto input = argv[1];
auto output = argv[2];
auto fst = fst::ReadFstKaldi(input);
auto num_states = fst->NumStates();
KALDI_LOG << "Loading FST from " << input << " with " << num_states
<< " states.";
int32 num_arc_added = AddSelfLoopsSimple(fst);
KALDI_LOG << "Adding " << num_arc_added << " self-loop arcs ";
fst::WriteFstKaldi(*fst, std::string(output));
KALDI_LOG << "Writing FST to " << output;
delete fst;
}
# + papermill={"duration": 40.409853, "end_time": "2021-05-30T15:50:41.796279", "exception": false, "start_time": "2021-05-30T15:50:01.386426", "status": "completed"} tags=[] id="partial-quarter" outputId="ee4f7e43-5a09-4bcf-b7b3-2701df035e7d"
# !zsh prepare_text.sh sv /kaggle/working/sentences.txt /kaggle/working/preppedtext
|
_notebooks/2021-05-26-wav2vec-u-cv-swedish-text-prep.ipynb
|
# ---
# jupyter:
# jupytext:
# cell_metadata_filter: -run_control,-deletable,-editable,-jupyter,-slideshow
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <p><font size="6"><b> Pandas: Methods for data cleaning</b></font></p>
#
#
# > *© 2021, <NAME> and <NAME> (<mailto:<EMAIL>>, <mailto:<EMAIL>>). Licensed under [CC BY 4.0 Creative Commons](http://creativecommons.org/licenses/by/4.0/)*
#
# ---
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
# -
# A number of Pandas functions are useful when cleaning up raw data and converting it to a data set ready for analysis and visualisation. In this notebook a selection of methods are introduced:
#
# - `drop`
# - `rename`
# - `replace`
# - `explode`
# - `drop_duplicates`/`duplicates`
# - `astype`
# - `unique`
# - `.str.`-methods
#
# __Note:__ Working with _missing values_ is tackled in a dedicated notebook [pandas_08_missing_values](./pandas_08_missing_values.ipynb).
# We showcase using a _dirty_ example data:
# + tags=[]
countries = pd.DataFrame({'county name': ['Belgium', 'Flance', 'Germany', 'Netherlands', ['United Kingdom', 'Germany']],
'population': [11.3, 64.3, 81.3, 16.9, 64.9],
'area': [30510, 671308, 357050, 41526, [244820, np.nan]],
'capital': ['Brussels', ' Paris ', 'Barlin', 'Amsterdam', 'London']})
countries
# -
# ## `drop`
# Drop columns (or rows) by name (this can also be achieved by selecting the columns you want to keep, but if you only want to drop a few columns, `drop()` is easier). Specify a list of column names to drop:
countries.drop(columns=["area", "capital"])
# ## `rename`
# Use a `dict` with the dictionary keys the old column/index name and the dictionary values the new column/index name:
countries = countries.rename(columns={"county name": "country"})
# ## `replace`
# Replace values in a column. Different inputs can be used. The most basic one is providing a value `to_replace` and a new `value`:
countries["capital"].replace("Barlin", "Berlin")
# Similar to `rename`, one can use a `dict` with the dictionary keys the old data and the dictionary values the new data:
countries = countries.replace({"Barlin": "Berlin", "Flance": "France"})
countries
# ## `explode`
# [`explode`](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.explode.html) multiple values in a cell to individual records (rows). Not regularly required, but very powerful when in case:
countries_exploded = countries.explode(["country", "area"])
countries_exploded
# ## `drop_duplicates`
# Checking duplicate values in a data set with [`duplicated`](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.duplicated.html) or remove duplicate values with [`drop_duplicates`](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.drop_duplicates.html):
countries_exploded.duplicated(subset=["country"])
countries_exploded = countries_exploded.drop_duplicates(subset=["country"], keep="first").copy() # More on this copy later
countries_exploded
# ## `astype`
# Pandas read functions might not always use the most appropriate data type for each of the columns. Converting them to a different data type can also improve the memory usage of the DataFrame (e.g. `int16` versus `float64`). The [`astype` ](https://pandas.pydata.org/docs/reference/api/pandas.DataFrame.astype.html) method supports data type conversion to both [Numpy data types](https://numpy.org/doc/stable/user/basics.types.html) as well as [Pandas specific data types](https://pandas.pydata.org/docs/user_guide/basics.html#dtypes).
countries_exploded.dtypes
countries_exploded["area"] = countries_exploded["area"].astype(int)
countries_exploded.dtypes
# ## `unique`
# Working with larger data sets, knowing which values are in a column:
countries_exploded["capital"].unique()
# ## `.str.`-methods
# Noticed the redundant spaces in the capital column?
#
# Whereas `replace` could work for this specific case (it also accepts _regular expressions_):
#
# ```python
# countries_exploded["capital"].replace(r"^\s+|\s+", "", regex=True)
# ```
#
# Pandas provides a set of convenient __string__ methods to handle these (element-wise) cleaning of strings, each of them accessible with the `.str.` accessor (e.g. `str.split`, `str.startswith`, `removeprefix`):
countries_exploded["capital"] = countries_exploded["capital"].str.strip()
countries_exploded["capital"].unique()
# <div class="alert alert-info">
#
# __INFO__
#
# For an overview of the available `.str.`-methods, see https://pandas.pydata.org/docs/user_guide/text.html#method-summary
#
# </div>
# ## Exercises: application on a real dataset
# For these exercises, we will use data of road casualties in Belgium in 2020 [made available by statbel](https://statbel.fgov.be/en/open-data/road-casualties-2020). The [metadata](https://statbel.fgov.be/sites/default/files/files/opendata/Verkeersslachtoffers/TF_ACCIDENTS_VICTIMS_META.xlsx) is available as well as a reference. The data contains the number of victims due to road causalities:
#
# - `MS_VCT`: Number of victims
# - `MS_VIC_OK`: Number of uninjured
# - `MS_SLY_INJ`: Number of slightly injured
# - `MS_SERLY_INJ`: Number of severely injured
# - `MS_MORY_INJ`: Number of mortally injured
# - `MS_DEAD`: Number of dead
# - `MS_DEAD_30_DAYS`: Number of dead after 30 days
#
# Together with metadata about date and time, the victim and road type, light conditions, location,...
#
# Pandas can load the data directly from the `zip`-file :
casualties_raw = pd.read_csv("./data/TF_ACCIDENTS_VICTIMS_2020.zip",
compression='zip',
sep="|",
low_memory=False)
casualties_raw.head()
# <div class="alert alert-info">
#
# __INTERMEZZO - display options__
#
# Pandas provides a number of configurable settings to display data, for example `display.max_rows`, `display.precision` and `display.max_columns`. When exploring a new data set, adjusting the `display.max_columns` setting is of particular interest to be able to scroll the full data set.
#
# See https://pandas.pydata.org/docs/user_guide/options.html#options-and-settings for the documentation and an [overview of the available settings](https://pandas.pydata.org/docs/user_guide/options.html#available-options)
#
# </div>
pd.options.display.max_columns = 45
casualties_raw.head()
# Whereas the data is already well organised and structured, some adjustments are required to support further analysis:
#
# - Combine the day and hour into a single datetime-aware data type.
# - Clean up the column names.
# - Metadata is always provided both in Dutch and French.
# - ...
#
# Let's apply the cleaning methods to clean up the data in the next set of exercises.
# <div class="alert alert-success">
#
# **EXERCISE 1**
#
# Remove all the `_FR` metadata columns from the `casualties_raw` data set and assign the result to a new variable `casualties_nl`. Access the column names using the `df.columns` attribute and use your standard Python [list comprehension](https://docs.python.org/3/tutorial/datastructures.html#list-comprehensions) skills.
#
# <details><summary>Hints</summary>
#
# - Instead of enlisting the column names manually, a [list comprehension](https://docs.python.org/3/tutorial/datastructures.html#list-comprehensions) - a _feature of standard Python_ - can be used to select the columns names ending on `_FR`.
# - Within the list comprehension, the [`endswith()`](https://docs.python.org/3/library/stdtypes.html#str.endswith) standard method can be used to check if a column name ends on `_FR`.
# - ! Pandas also provides the `.str.endswith()` method, but this is for the data values inside a DataFrame. In this exercise we want to adjust the column names itself.
# - Remove columns with the `drop()` method.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning1.py
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning2.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 2**
#
# A number of the remaining metadata columns names have the `TX_` and the `_DESCR_NL` in the column name. Clean up these column names by removing the `TX_` at the start and the `_DESCR_NL` at the end of the column names. Update the `casualties_nl` variable, and assign the result to `casualties`.
#
# <details><summary>Hints</summary>
#
# - Use the `rename` method and apply the mapping on the `columns`.
# - Manually writing a mapping dictionary, e.g. `{"TX_DAY_OF_WEEK_DESCR_NL": "WEEK_DAY"}`, is fine. However, [dict comprehensions](https://docs.python.org/3/tutorial/datastructures.html#dictionaries) can be used as well to setup the mapping between old and new values.
# - `removeprefix()` and `removesuffix()` are [Python string methods](https://docs.python.org/3/library/stdtypes.html#string-methods) to remove start/trailing characters if present.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning3.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 3**
#
# Check the unique values of the `SEX` column.
#
# Based on the the values, create a mapping to replace the values with the english version (`"male", "female"`). Use `None` for the unknown values (`Onbekend` in Dutch). Apply the mapping to overwrite the values in the `SEX` column with the new value.
#
# <details><summary>Hints</summary>
#
# - Create the mapping by hand and define a `dict`.
# - Use the `replace()` method to update the values of the `SEX` column.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning4.py
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning5.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 4**
#
# Check the unique values of the `DT_HOUR` column. Which of the data values is used as _not a number_ (not known)? Verify the amount of records that with the `DT_HOUR` not known.
#
# A check with the data provider confirmed that the record(s) with value 99 did actually happen at 9 AM and are a typo instead of _not a number_ replacement value. Replace the 99 values with the real hour of the day in the `DT_HOUR` column.
#
# <details><summary>Hints</summary>
#
# - The number `99` is not a valid hour of the day and used as not a number data point.
# - Only one data record has an unknown hour of the day.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning6.py
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning7.py
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning8.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 5**
#
# In the `AGE_CLS` column, the age is formatted as `X tot Y jaar` (i.e. _x till y year_). Remove the Dutch description and convert the data into a format `Y - Y` to define the age classes.
#
# Use the string methods as much as possible. The `Onbekend`, ` ` (empty string) and `75 jaar en meer` data values can be done by direct replacement into `None`, `None` and `> 75` respectively.
#
# <details><summary>Hints</summary>
#
# - Use the `.str.replace()` (note the difference with the Pandas `replace()` method) and the `str.removesuffix()` methods to convert the data format.
# - Add an additional `str.strip` to get rid of the spaces and the 'unknown' number of spaces in the empty string case.
# - Using the `replace()` method with a dictionary just works for the remaining two values: `{"Onbekend": None, "75 jaar en meer": ">75"}`. It will leave other values (not specified in the dictionary) as is.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning9.py
# -
# verify outcome
casualties["AGE_CLS"].unique()
# <div class="alert alert-success">
#
# **EXERCISE 6**
#
# The day (`DT_DAY`) and hour (`DT_HOUR`) are two separate columns instead of a single `datetime` data type column.
#
# - Check the data types of the `DT_DAY` and `DT_HOUR` columns.
# - Combine the two columns into a single column (using _string concatenation_) and use the `pd.to_datetime` function to convert the combined column (call the column `"datetime"`).
#
# <details><summary>Hints</summary>
#
# - The data type of columns is available as the `dtypes` attribute.
# - String concatenation is done element-wise in Pandas using the `+` operator. Do not forget to convert the `DT_HOUR` column into a `str` column using `astype`.
# - Without adding a minute level, the datetime conversion of `pd.to_datetime` won't work. Add `":00"` to provide minutes as well.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning10.py
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning11.py
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning12.py
# -
# <div class="alert alert-success">
#
# **EXERCISE 7**
#
# For columns consistiong of a limited number of categories with (_ordinal data_) or without a logical order, Pandas has a specific data type: `Categorical`. An example in the data set is the `DAY_OF_WEEK` (from `Monday` -> `Sunday`).
#
# For this conversion, the `astype` is not sufficient. Use the `pd.Categorical` function (check the documentation) to create a new column `week_day` with the week days defined as a Categorical variable. Use Monday as the first day of the week and make sure the categories are ordered.
#
# <details><summary>Hints</summary>
#
# - See [Pandas categorical info](https://pandas.pydata.org/pandas-docs/stable/user_guide/categorical.html#object-creation) for more information
# - Use `ordered=True` to define ordered data.
#
# </details>
#
# </div>
casualties["DAY_OF_WEEK"] = casualties["DAY_OF_WEEK"].replace({"maandag": "Monday", "dinsdag": "Tuesday", "woensdag": "Wednesday",
"donderdag": "Thursday", "vrijdag": "Friday", "zaterdag": "Saturday",
"zondag": "Sunday"})
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning13.py
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning14.py
# -
# <div class="alert alert-success">
#
# **(OPTIONAL) EXERCISE 8**
#
# The data set contains the number of victims. The link with the individual accidents is not available in the current data set and multiple records/rows of the data set can belong to a single accident.
#
# We can expect that records with the same day, hour, municipality , light condition, road type and build up area are probably linked to the same accident. Try to estimate the number of accidents.
#
# <details><summary>Hints</summary>
#
# - This exercise is a special case of the `drop_duplicates` method. When we drop duplicate records when `"DT_DAY", "DT_HOUR", "CD_MUNTY_REFNIS", "BUILD_UP_AREA","LIGHT_COND", "ROAD_TYPE"` are all the same, we have an estimate on the number of accidents.
# - Use the `subset` parameter to define a specific set of column names.
#
# </details>
#
# </div>
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning15.py
# + tags=["nbtutor-solution"]
# # %load _solutions/pandas_09_data_cleaning16.py
|
notebooks/pandas_09_data_cleaning.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib
import matplotlib.pyplot as plt
import numpy as np
x = np.linspace(-10,10,300)
sin_y = np.sin(x)
cos_y = np.cos(x)
plt.figure(figsize=(15, 5))
plt.plot(x,sin_y,color='red', marker='', linestyle='dashed', linewidth=1, markersize=1)
plt.gray()
plt.ylabel("my_y")
plt.plot()
gist_x = np.random.normal(0.9,1,1000)
gist_x2 = np.random.normal(2,2,1000)
plt.figure(figsize=(15, 5))
plt.hist(gist_x2)
plt.hist(gist_x)
# +
noise = np.random.normal(0,1,300)
x = np.random.uniform(0, 100, 300)
y3 = np.random.uniform(0, 100, 300)
y1 = 3 * x + noise
y2 = 10 * x + noise
plt.plot(x, y1)
plt.plot(x, y2)
# -
plt.scatter(y1,y3)
# <center>
# <img src="https://github.com/hse-econ-data-science/dap_2021_spring/blob/main/sem10_visual/images/visual.png?raw=true" height="400" width="700">
# </center>
# # <center> Визуализация данных в python </center>
# Визуализация занимает важную часть в анализе данных. Представляя информацию в графическом виде, вы облегчаете процесс ее восприятия, что дает возможность выделять дополнительные закономерности, оценивать соотношения величин, быстрее доносить ключевые аспекты в данных.
#
# Начнем с небольшой "памятки", о которой всегда нужно помнить при создании любых графиков.
# ## <center> Как визуализировать данные и заставить всех тебя ненавидеть </center>
# 1. Заголовок графика для слабаков. По графику всегда понятно, какие данные и явления он описывает.
# 2. Ни в коем случае не подписывай ни одной оси у графика. Пусть смотрящий развивает свою интуицую!
# 3. Единицы измерения совсем не обязательны. Какая разница, в чем измеряли количество - в людях или в литрах!
# 4. Чем меньше шрифт на графике, тем острее зрение смотрящего.
# 5. На одном графике нужно стараться уместить всю информацию, которая у тебя есть в датасете. С полными названиями, расшифровками, сносками. Чем больше текста - тем информативнее!
# 6. При любой возможности используйте 3D и спецэффекты, пусть знают, что ты — прирожденный дизайнер. К тому же, так будет меньше визуальных искажений.
# Если серьезно, то обязательно посмотрите список рекомендованных материалов в конце ноутбука по правилам оформления графиков и работе с библиотеками для визуализации данных в Python.
#
# Основные библиотеки для визуализации в Python - это `matplotlib`, `seaborn`, `plotly`. Сегодня познакомимся с первыми двумя
# +
import numpy as np # библиотека для матриц и математики
import pandas as pd # библиотека для работы с табличками
# библиотеки для визуализации
import matplotlib.pyplot as plt
import seaborn as sns
# plt.style.use('ggplot') # стиль графиков
# %matplotlib inline
# -
# # Продажи и оценки видеоигр
# Работаем с датасетом по продажам и оценкам видео-игр. Датасет взят с [Кеггла](https://www.kaggle.com/rush4ratio/video-game-sales-with-ratings).
#
# __Описание колонок:__
# * `Name` $-$ название видеоигры
# * `Platform` $-$ платформа, на которой игра была запущена
# * `Year_of_Release` $-$ год релиза
# * `Genre` $-$ жанр
# * `Publisher` $-$ издатель
# * `NA_Sales` $-$ объем продаж в Северной Америке (в млн штук)
# * `EU_Sales` $-$ объем продаж в Евросоюзе (в млн штук)
# * `JP_Sales` $-$ объем продаж в Японии (в млн штук)
# * `Other_Sales` $-$ объем продаж в остальном мире (в млн штук)
# * `Global_Sales` $-$ общий объем продаж (в млн штук)
# * `Critic_Score` $-$ совокупный балл, составленный сотрудниками Metacritic
# * `Critic_Count` $-$ кол-во критиков, оцениваемых игру
# * `User_Score` $-$ совокупный балл, составленный подписчиками Metacritic (пользователями)
# * `User_Count` $-$ кол-во пользователей, оцениваемых игру
# * `Developer` $-$ ответственный за создание игры
# * `Rating` $-$ рейтинг (Everyone, Teen, Adults Only и тд)
df = pd.read_csv('data/video_games_sales.csv')
df.sort_values(by="Year_of_Release")
# В данных много пропусков, поэтому давайте выкинем все пропущенные наблюдения. Также видим, что некоторые колонки pandas привел не к тому типу. Исправим это:
# +
df = df.dropna()
df['User_Score'] = df.User_Score.astype('float64')
df['Year_of_Release'] = df.Year_of_Release.astype('int64')
df['User_Count'] = df.User_Count.astype('int64')
df['Critic_Count'] = df.Critic_Count.astype('int64')
# -
df.shape
# ## 1. Изучаем основы matplotlib на примере линейных графиков
# Самый простой вариант создания графика в matplotlib - функция `plt.plot()`, которой мы передаем два аргумента - что положить на ось _x_, а что на _y_. Если у вас переменные в числовом формате, то без проблем получите линейный график (line plot)
# `plt.plot(x = ___, y = ___)`
# **Посмотрим на динамику продаж игр в мире по годам:**
gb.values
gb
gb["NA_Sales"]
# +
# агрегируем данные за год
gb = df.groupby('Year_of_Release')["NA_Sales","EU_Sales","JP_Sales"].sum()
# строим график
plt.plot(gb.index, gb["NA_Sales"], label="NA_Sales")
plt.plot(gb.index, gb["EU_Sales"], label="EU_Sales")
plt.plot(gb.index, gb["JP_Sales"], label="JP_Sales")
plt.legend()
# -
pd.DataFrame(gb)
# ### Задача №1:
#
# Постройте один график, на котором будут отображаться сразу все показатели продаж (NA_Sales, EU_Sales, JP_Sales, Other_Sales, Global_Sales)
# +
## Ваш код
# -
# В идеальной вселенной мы создаем графики функцией `subplots`, которая генерирует наш график в 2 переменные (обычно их называют fig и ax):
#
# * `fig` отвечает за график в целом. Воспринимайте ее как файл, который хранит график как картинку.
# * `ax` $-$ это ось координат, на которой мы собственно строим график. Все элементы графика хранятся как раз в ней.
fig, ax = plt.subplots(2,2, figsize=(7,5)) # создали полоино для графика из 1 ряда и 1 колонки (1 график)
# figsize -это размер нашего прямоугольника в неочевидных единицах.
# Какой размер удачный? экспериментируйте!
# **Посмотрим на динамику продаж игр в мире по годам, но теперь используем `subplots`**
# +
# строим график
gb = df.groupby('Year_of_Release')["Global_Sales", "NA_Sales","EU_Sales","JP_Sales"].sum()
fig, ax = plt.subplots(2,2, figsize=(10,5))
# строим график
ax[0, 0].plot(gb.index, gb["NA_Sales"], label="NA_Sales")
ax[1, 0].plot(gb.index, gb["EU_Sales"], label="EU_Sales")
ax[0, 1].plot(gb.index, gb["JP_Sales"], label="JP_Sales")
ax[1, 1].plot(gb.index, gb["Global_Sales"], label="Global_Sales")
# -
# В переменной _ax_ на самом деле лежит куча методов, которые позволяют сделать график более приятным. Посмотрим на несколько из них:
# +
fig, ax = plt.subplots(1,1, figsize=(10,5))
# параметры самого графика (цвет линии, стиль и т.д.) определяем как параметры в методе plot()
# меняем цвет и стиль линии на пунктир. Matplotlib знает некоторые стандартные цвета, и их можно задать прямо словом
# так же можно передать hex цвет. Например, #8c92ac
ax.plot(gb.index, gb.values, color = 'grey', ls = ':')
# если вы обратили внимание, то в нашем самом первом графике шкала с годами сломамлась и стала float. Matplotlib принудительно
# делает x непрерывной переменной для линейного графика. Мы хотим оставить шкалу год в целых числах.
ax.locator_params(integer=True)
# называем шкалы x и y, выбираем размер шрифта.
ax.set_xlabel('Year of Release', fontsize=12)
ax.set_ylabel('Global Sales', fontsize=12)
# делаем правую и верхнюю границу графика невидимыми
ax.spines['right'].set_visible(False)
ax.spines['top'].set_visible(False)
# делаем засечки на шкалах x и y потоньше
ax.xaxis.set_tick_params(width=0.2)
ax.yaxis.set_tick_params(width=0.2)
# уменьшаем толщину оставших границ графика с помощью цикла
# (можно и без цикла отдельной строкой для каждой границы, как делали выше)
for spine in ['bottom','left']:
ax.spines[spine].set_linewidth(0.2)
# -
# ### Создаем систему графиков
fig, ax = plt.subplots(2,2, figsize=(14,9)) # увеличиваем количество объектов до двух рядов и двух колонок.
# Теперь у нас есть четыре графика!
#
# Объект _ax_ в данном случае **становится матрицой**. И чтобы обратиться к каждому графику, нужно обратиться к нему по индексу из _ax_.
# +
fig, ax = plt.subplots(2,2, figsize=(14,9))
ax[0][0].plot(gb.index, gb.values);
# -
# Ниже посмотрим как с помощью цикла for можно применить какую-то модификацию ко всем графикам системы.
# +
fig, ax= plt.subplots(2, 2, figsize=(14,9))
for x in range(2):
for y in range(2):
ax[x][y].set_xlabel('X')
# -
# ### Задача №2:
#
# Создайте систему графиков, у которой:
# * на 1ом графике изображены продажи в Северной Америке (`NA_Sales`);
# * на 2ом продажи в Европейском Союзе (`EU_Sales`);
# * на 3ем продажи в Японии (`JP_Sales`)
# * на 4ом графике все 3 линии вместе;
# * Каждый график должен быть разного цвета. В четвертом графике - каждый график должен быть такого же цвета, как и в своей ячейке + 4ый график должен иметь легенду
# * В заголовке к каждому графику напишите то, что стоит у него по оси _y_
# +
## Ваш код
# -
# ## 2. Графики для категориальных переменных
# **Посмотрим на кол-во игр за каждый год с помощью столбчатой диаграммы:**
df.groupby('Year_of_Release').Name.count()
df.groupby('Year_of_Release').Name.count().plot(kind='bar');
# **Посмотрим отдельно на кол-во игр жанра Sports и Action за последние 5 лет. Сделаем двойную диаграмму и развернем ее горизонтально**
# +
fig, ax = plt.subplots(nrows=1, ncols=2, figsize=(10, 8))
gb_sports = df[df.Genre=='Sports'].groupby('Year_of_Release').Name.count().tail(5).sort_index(ascending=False)
gb_sports.plot(kind='barh', ax=ax[0])
ax[0].set_title('Sports')
gb_act = df[df.Genre=='Action'].groupby('Year_of_Release').Name.count().tail(5).sort_index(ascending=False)
gb_act.plot(kind='barh', ax=ax[1])
ax[1].set_title('Action')
ax[1].yaxis.set_ticks_position('right')
for i in range(2):
ax[i].set_ylabel('', visible=False) # убираем подпись к шкале, которая генерируется автоматически
# -
# Подумайте, чем плох такой график?
# ### Задача №3:
#
# С помощью двух вертикальных столбчатых диаграмм выведите среднюю оценку критиков и пользователей игр различных рейтингов.
# +
## Ваш код
# -
# ### Задача №4:
#
# В виде горизонтальной столбчатой диаграммы выведите топ-5 девелоперов с наибольшей средней оценкой от критиков. Рядом выведите диаграмму их самой успешной игры по общим продажам.
# +
## Ваш код
# -
# ### Задача №5:
#
# Сравните оценки игр от критиков для топ-5 крупнейших игровых платформ (по кол-ву игр) с помощью boxplot из пакета seaborn.
# +
## Ваш код
# -
# ## 3*. Мультивариативный график рассеяния
#
# Сейчас будем работать с наборам данных, который содержит информацию о количестве преступлений в штатах США в 2005 году.
crimes = pd.read_csv('data/crimeRatesByState2005.tsv', sep='\t')
crimes.head()
# График рассеяния (scatter plot) $-$ это такой график, у которого по оси _x_ и _y_ отложены непрерывные переменные. График состоит из точек, каждая из которых отвечает за свое наблюдение.
# Посмотрим на график зависимостей убийств от краж со взломом (burglary)
fig, ax = plt.subplots()
ax.scatter(crimes['murder'], crimes['motor_vehicle_theft']);
# Здесь каждая точка отвечает за свой штат. По оси _x_ мы видим сколько в штате было убийств, а по оси _y_ сколько краж со взломом
# Попробуем на график добавить дополнительную информацию, например, в качестве размера точки обозначим кол-во населения
# +
fig, ax = plt.subplots()
# добавляем параметр s (size) и говорим, какая переменная будет за него отвечать
ax.scatter(crimes['murder'], crimes['burglary'], s = crimes['population']);
# -
# Размер населения такой большой, что точка захватила всю область координат. Давайте попробуем нашу переменную масштабировать - нам же важны относительные размеры штатов относительно друг друга, а не абсолютные значения. Значения маштабирования тоже выбираем экспериментально: то, что лучше выглядит и более информативно.
fig, ax = plt.subplots()
ax.scatter(crimes['murder'], crimes['burglary'], s = crimes['population']/35000);
# Отлично, однако следующая проблема - слияние точек. Давайте добавим параметр прозрачности, чтобы было видно, где они накладываются друг на друга.
fig, ax = plt.subplots()
ax.scatter(crimes['murder'], crimes['burglary'], s = crimes['population']/35000, alpha = 0.5);
# Добавим теперь еще какую-нибудь переменную (например, robbery) и засунем ее в параметр цвета
fig, ax = plt.subplots()
ax.scatter(crimes['murder'], crimes['burglary'], s = crimes['population']/35000, alpha = 0.5,
c = crimes['Robbery']); # задаем новый аргумент c (color) и присваиваем ему значение переменной
# Осталось узнать, что значит какой цвет. Для этого нужно сохранить график в переменную и передать ее как аргумент функции `colorbar()`. Также можем поменять цветовую шкалу с помощью аргумента cmap.
fig, ax = plt.subplots()
color_graph = ax.scatter(crimes['murder'], crimes['burglary'], s = crimes['population']/35000, alpha = 0.5, cmap = 'inferno',
c = crimes['Robbery'])
plt.colorbar(color_graph);
# Последнее что тут сделаем - это подпишем штаты
# +
fig, ax = plt.subplots(figsize = (22,10))
color_graph = ax.scatter(crimes['murder'], crimes['burglary'], s = crimes['population']/35000,
c = crimes['Robbery'], cmap = 'inferno', alpha = 0.5, linewidth = 0)
ax.yaxis.set_ticks_position('left')
ax.xaxis.set_ticks_position('bottom')
ax.set_xlabel('Murder', fontsize = 10)
ax.set_ylabel('Bulglarly', fontsize = 10)
for i, state in enumerate(crimes['state']): # с помощью enumerate создаем из колонок с названиями штатов объект кортежей вида индекс - название штата.
ax.annotate(state, (crimes['murder'][i], crimes['burglary'][i]), fontsize = 10) # используем метод annotate, которому говорим, что нужно брать имя штата
# из кортежа, который создали с помощью enumerate, а координаты подписи задаем через
# значения наших переменных x и y для нужного индекса из объекта enumerate (обращается к нужному
# ряду в датафрейме)
plt.colorbar(color_graph);
# -
|
sem10_visual/sem10_205.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Assignment 2
# ## Data
#
# This is a historical dataset on the modern Olympic Games, including all the Games from Athens 1896 to Rio 2016. The data was taken from Kaggle. The `athlete_events` Dataset contains $271,116$ rows and $15$ columns.
#
# **Source:**
#
# <NAME> (2018) 120 years of Olympic history: athletes and results, athlete_events, Found at: https://www.kaggle.com/heesoo37/120-years-of-olympic-history-athletes-and-results#athlete_events.csv
#
# **ATTRIBUTES:**
#
# **athlete_events.csv**
#
# | Column Name | Data Type | Description/Notes |
# |:----:|:----:|:----|
# | ID | integer | Unique number for each athlete |
# | Name | string | Athlete’s name |
# | Sex | string | M or F |
# | Age | integer | |
# | Height | integer | In centimeters |
# | Weight | integer | In kilograms |
# | Team | string | Team name |
# | NOC | string | National Olympic Committee, 3 letter code (Matches with `NOC` from noc_regions.csv) |
# | Games | string | Year and season |
# | Year | integer | |
# | Season | string | Summer or Winter |
# | City | string | Host city |
# | Sport | string | |
# | Event | string | |
# | Medal | string | Gold, Silver, Bronze, or NA |
# Download athlete_events.csv from the link above and load it into a DataFrame called `athlete_events`:
# Your answer goes here
# Use `info()` method on this DataFrame to get a sense of the data:
# Your answer goes here
# ## Question 1
# Clearly not every athlete receives a medal in the olympics. How many records have a non-null value for `Medal` field? In other words, how many medals were given according to this dataset?
# Your answer goes here
# Use the `head()` method to inspect the first few rows of the data:
# Your answer goes here
# ## Question 2
#
# What is the minimum and maximum `Year`?
# Your answer goes here
# Your answer goes here
# ## Question 3
#
# Is the following statement True or False?
#
# > Average age of female athletes who attended the olympic games after 1990 has raised when compared to the era before then.
# Your answer goes here
# Your answer goes here
# ## Question 4
#
# How many Gold medals were given to men from 1970 to 2000 (including both years)?
# Your answer goes here
# ## Question 5
#
# How many athletes attended the olympic games in 1968?
#
# Hint: You can use the method `.unique()` on any pandas Series to get the unique values. Any column subset of a DataFrame returns a Series object.
# Your answer goes here
# ## Question 6
#
# Who won the event "Swimming Men's 100 metres Breaststroke" in 2004? Please note that there is a typo in the Event description!
# Your answer goes here
|
05-Data-Transformation/Assignment-2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + language="javascript"
# require(["codemirror/keymap/sublime", "notebook/js/cell", "base/js/namespace"],
# function(sublime_keymap, cell, IPython) {
# cell.Cell.options_default.cm_config.keyMap = 'sublime';
# var cells = IPython.notebook.get_cells();
# for(var cl=0; cl< cells.length ; cl++){
# cells[cl].code_mirror.setOption('keyMap', 'sublime');
# }
# }
# );
# -
# change the cell width
from IPython.core.display import display, HTML
display(HTML("<style>.container { width:85% !important; }</style>"))
# %load_ext autoreload
# %autoreload 2
# %matplotlib inline
# %config InlineBackend.print_figure_kwargs={'facecolor' : "w"}
import warnings
warnings.filterwarnings('ignore')
from more_itertools import chunked
from pathlib import Path
# import all samples as dictionary ...
from samples import *
import os
os.environ["MKL_NUM_THREADS"] = "1"
os.environ["NUMEXPR_NUM_THREADS"] = "1"
os.environ["OMP_NUM_THREADS"] = "1"
# # %matplotlib inline
import matplotlib.pyplot as plt
import matplotlib as mpl
mpl.style.use('seaborn-white')
import seaborn as sns
import multiprocess as mp
import numpy as np
import pandas as pd
import bioframe
import cooltools
import cooler
from matplotlib.gridspec import GridSpec
import bbi
from cooltools import snipping
from matplotlib.colors import LogNorm
# https://stackoverflow.com/questions/48625475/python-shifted-logarithmic-colorbar-white-color-offset-to-center
class MidPointLogNorm(LogNorm):
def __init__(self, vmin=None, vmax=None, midpoint=None, clip=False):
LogNorm.__init__(self, vmin=vmin, vmax=vmax, clip=clip)
self.midpoint=midpoint
def __call__(self, value, clip=None):
result, is_scalar = self.process_value(value)
x, y = [np.log(self.vmin), np.log(self.midpoint), np.log(self.vmax)], [0, 0.5, 1]
return np.ma.array(np.interp(np.log(value), x, y), mask=result.mask, copy=False)
# +
# Use bioframe to fetch the genomic features from the UCSC.
hg19_chromsizes = bioframe.fetch_chromsizes('hg19', as_bed=True)
hg19_cens = bioframe.fetch_centromeres('hg19')
hg19_arms = bioframe.split(hg19_chromsizes, hg19_cens, cols_points=['chrom', 'mid'])
# Select only chromosomes that are present in the cooler.
# This step is typically not required! we call it only because the test data are reduced.
hg19_chromsizes = hg19_chromsizes.set_index("chrom").loc[autosomal_chroms].reset_index()
hg19_arms = hg19_arms.set_index("chrom").loc[autosomal_chroms].reset_index()
# call this to automaticly assign names to chromosomal arms:
hg19_arms = bioframe.parse_regions(hg19_arms)
hg19_arms["name"] = [f"{chrom}{arm}" for chrom in autosomal_chroms for arm in list('pq')]
hg19_arms.to_csv("hg19_arms.bed",sep="\t",index=False,header=False)
# +
conditions = ['442-NT','442-IAA']
binsize = 2000
binsize_human = f"{int(binsize/1_000)}kb"
cooler_paths = {
'442-NT' : f'CkoCT442-NT-pool.mcool::resolutions/{binsize}',
'442-IAA' : f'CkoCT442-IAA-pool.mcool::resolutions/{binsize}',
}
long_names = {
'442-NT' : 'CTCFdegron-442-nontreated-pool',
'442-IAA': 'CTCFdegron-442-auxin-pool',
}
pal = sns.color_palette('colorblind')
colors = {
'442-NT': "#e74c3c",
'442-IAA': pal[2],
}
clrs = {
cond: cooler.Cooler(cooler_paths[cond]) for cond in conditions
}
# -
# iterate over samples to calculate insulation on:
for cond in conditions:
lname,*_ = cooler_paths[cond].split(".mcool")
target_exp_file = Path(f"{lname}.{binsize_human}.cis.exp.tsv")
if target_exp_file.is_file():
print("already exist !")
print(target_exp_file)
continue
else:
print("working on ...")
print(cond,lname)
######################################
# !cooltools compute-expected \
# --nproc 4 \
# --output {target_exp_file} \
# --regions hg19_arms.bed \
# --contact-type cis \
# --balance \
# --weight-name weight \
# --ignore-diags 2 \
# {cooler_paths[cond]}
######################################
expected_paths = {
'442-NT' : f'CkoCT442-NT-pool.{binsize_human}.cis.exp.tsv',
'442-IAA' : f'CkoCT442-IAA-pool.{binsize_human}.cis.exp.tsv',
}
expecteds = {
cond: pd.read_csv(expected_paths[cond], sep="\t") for cond in conditions
}
# +
# read a BEDPE file with interactions chr1,start1, ...end2 - with optional annotations
# that would enable one to split pileups by category ...
sites = pd.read_table("nearestCTCF_TSS.bedpe", delimiter='\t')
# sites = pd.read_table("allCTCF_TSS.bedpe", delimiter='\t')
sites['mid1'] = (sites['start1'] + sites['end1'])/2
sites['mid2'] = (sites['start2'] + sites['end2'])/2
print(len(sites))
sites.head()
# +
snippet_flank = 50_000
windows1 = snipping.make_bin_aligned_windows(
binsize,
sites['chrom1'],
sites['mid1'],
flank_bp=snippet_flank)
windows2 = snipping.make_bin_aligned_windows(
binsize,
sites['chrom2'],
sites['mid2'],
flank_bp=snippet_flank)
windows = pd.merge(windows1, windows2, left_index=True, right_index=True, suffixes=('1', '2'))
windows = snipping.assign_regions(windows, hg19_arms) # function "assign_regions" might move
windows = windows.dropna() # do we need this ?
# let's filter sites here as well ...
arm_sites = sites.loc[windows.index,:].reset_index(drop=True)
windows = windows.reset_index(drop=True)
#
windows.head()
# -
# the most time consuming step !!!
stacks = {}
piles = {}
for cond in conditions:
snipper = snipping.ObsExpSnipper(clrs[cond], expecteds[cond], regions=hg19_arms)
stack = snipping.pileup(windows, snipper.select, snipper.snip)
stacks[cond] = stack
# flipping can be done here ...
piles[cond] = np.nanmean(stack, axis=2)
# +
_piles = {}
_mask = (arm_sites["distance"]>50_000) & (arm_sites["distance"]<500_000)
for cond in conditions:
_substack = stacks[cond][:,:,_mask]
_piles[cond] = np.nanmean(_substack, axis=2)
gs = plt.GridSpec(nrows=1, ncols=len(conditions) + 1, width_ratios=[20] * len(conditions) + [1])
fig = plt.figure(figsize=(6 * len(conditions), 6))
opts = dict(
norm = MidPointLogNorm(vmin=0.8,vmax=1.5,midpoint=1),
extent=[-snippet_flank//1000, snippet_flank//1000, -snippet_flank//1000, snippet_flank//1000],
# cmap='RdBu_r',
cmap='coolwarm',
interpolation="nearest"
)
for i, cond in enumerate(conditions):
ax = plt.subplot(gs[i])
img = ax.matshow(
_piles[cond],
**opts)
ax.xaxis.tick_bottom()
if i > 0:
ax.yaxis.set_visible(False)
plt.title(long_names[cond])
ax = plt.subplot(gs[len(conditions)])
plt.colorbar(img, cax=ax)
plt.suptitle(f'{snippet_flank//1000}kb flank\n'
f'Hi-C resolution = {binsize//1000}kb; # of pairs = {_mask.sum()}')
# -
# ### Now we can start subsetting and splitting our `sites` dataframe (table with interactions) into subgroups by annotation
# at the end of each subsetting - the goal is to get indices corresponding to the subsetted conditions and extract snippets corresponding to those indices from `stacks[cond]` -> that would be the pileup of the subset of interactions ...
# here is our criteria to select a subgroup of snippets from the stack ....
upstream_pCTCF_pTSS_mask = (arm_sites["type1"]=="ctcf") & \
(arm_sites["strand1"]=="+") & \
(arm_sites["type2"]=="tss") & \
(arm_sites["strand2"]=="+") & \
(arm_sites["distance"]>50_000) & \
(arm_sites["distance"]<500_000)
#
upstream_pCTCF_pTSS_piles = {}
for cond in conditions:
# extract only desired snippets from the stack
# stacks are 3D arrays, with first 2 coords along the heatmap
# and the last coord (index) is along snippets (individual interactions from BEDPE) ...
_substack = stacks[cond][:,:,upstream_pCTCF_pTSS_mask]
upstream_pCTCF_pTSS_piles[cond] = np.nanmean(_substack, axis=2)
# +
gs = plt.GridSpec(nrows=1, ncols=len(conditions) + 1, width_ratios=[20] * len(conditions) + [1])
fig = plt.figure(figsize=(6 * len(conditions), 6))
opts = dict(
norm = MidPointLogNorm(vmin=0.8,vmax=1.5,midpoint=1),
extent=[-snippet_flank//1000, snippet_flank//1000, -snippet_flank//1000, snippet_flank//1000],
# cmap='RdBu_r',
cmap='coolwarm',
interpolation="nearest"
)
for i, cond in enumerate(conditions):
ax = plt.subplot(gs[i])
img = ax.matshow(
upstream_pCTCF_pTSS_piles[cond],
**opts)
ax.xaxis.tick_bottom()
if i > 0:
ax.yaxis.set_visible(False)
plt.title(long_names[cond])
ax = plt.subplot(gs[len(conditions)])
plt.colorbar(img, cax=ax)
plt.suptitle(f'{snippet_flank//1000}kb flank\n'
f'Hi-C resolution = {binsize//1000}kb; # of pairs = {upstream_pCTCF_pTSS_mask.sum()}')
# -
# #### here is the selection we used for _ALL-PAIRWISE_ interactions - right before we turned to neigbours ...
# +
# here is our criteria to select a subgroup of snippets from the stack ....
upstreamCTCF_pTSS_mask = (sites["type1"]=="ctcf") & (sites["type2"]=="tss") & (sites["strand2"]=="+")
upstreamCTCF_pTSS_piles = {}
for cond in conditions:
# extract only desired snippets from the stack
# stacks are 3D arrays, with first 2 coords along the heatmap
# and the last coord (index) is along snippets (individual interactions from BEDPE) ...
_substack = stacks[cond][:,:,upstreamCTCF_pTSS_mask]
# flipping can be done here ...
upstreamCTCF_pTSS_piles[cond] = np.nanmean(_substack, axis=2)
# more conditions to check ...
# (sites["type1"]=="ctcf") & (sites["type2"]=="tss") & (sites["strand2"]=="+")
# (sites["type2"]=="ctcf") & (sites["type1"]=="tss") & (sites["strand1"]=="-")
# (sites["type1"]=="ctcf") & (sites["type2"]=="tss") & (sites["strand2"]=="-")
# (sites["type2"]=="ctcf") & (sites["type1"]=="tss") & (sites["strand1"]=="+")
# +
gs = plt.GridSpec(nrows=1, ncols=len(conditions) + 1, width_ratios=[20] * len(conditions) + [1])
fig = plt.figure(figsize=(6 * len(conditions), 6))
opts = dict(
norm = MidPointLogNorm(vmin=0.82,vmax=1.22,midpoint=1),
extent=[-snippet_flank//1000, snippet_flank//1000, -snippet_flank//1000, snippet_flank//1000],
# cmap='RdBu_r',
cmap='coolwarm',
interpolation="nearest"
)
for i, cond in enumerate(conditions):
ax = plt.subplot(gs[i])
img = ax.matshow(
upstreamCTCF_pTSS_piles[cond],
**opts)
ax.xaxis.tick_bottom()
if i > 0:
ax.yaxis.set_visible(False)
plt.title(long_names[cond])
ax = plt.subplot(gs[len(conditions)])
plt.colorbar(img, cax=ax)
plt.suptitle(f'{snippet_flank//1000}kb flank\n'
f'Hi-C resolution = {binsize//1000}kb; # of pairs = {upstreamCTCF_pTSS_mask.sum()}')
# -
# ### we learned how to flip part of the stackup ...
#
# #### extract stacks for a coupleof relevant conditions that could be combined by flipping - and combine them !!!
# +
# here is our criteria to select a subgroup of snippets from the stack ....
downstream_mCTCF_mTSS_mask = (arm_sites["type1"]=="tss") & \
(arm_sites["strand1"]=="-") & \
(arm_sites["type2"]=="ctcf") & \
(arm_sites["strand2"]=="-") & \
(arm_sites["distance"]>50_000) & \
(arm_sites["distance"]<500_000)
upstream_pCTCF_pTSS_mask = (arm_sites["type1"]=="ctcf") & \
(arm_sites["strand1"]=="+") & \
(arm_sites["type2"]=="tss") & \
(arm_sites["strand2"]=="+") & \
(arm_sites["distance"]>50_000) & \
(arm_sites["distance"]<500_000)
# when selecting multiple conditions - it is reasonable to assume
# that these conditions are mutually exclusive
# check that the following way:
print("checking if conditions are mutually exclusive ...")
print(f"number of sites that satisfy either condition: {(upstream_pCTCF_pTSS_mask|downstream_mCTCF_mTSS_mask).sum()} ")
print(f"number of sites that satisfy condition1: {upstream_pCTCF_pTSS_mask.sum()} ")
print(f"number of sites that satisfy condition2: {downstream_mCTCF_mTSS_mask.sum()} ")
print(f"the sum {upstream_pCTCF_pTSS_mask.sum()+downstream_mCTCF_mTSS_mask.sum()} of the two latter ones is supposed to be equal to the former one")
combined_piles = {}
for cond in conditions:
# extract only desired snippets from the stack
# stacks are 3D arrays, with first 2 coords along the heatmap
# and the last coord (index) is along snippets (individual interactions from BEDPE) ...
_substack1 = stacks[cond][:,:,upstream_pCTCF_pTSS_mask]
_substack2 = stacks[cond][:,:,downstream_mCTCF_mTSS_mask]
# try to flip _substack2 !
# this is going to be specific in each and every case:
_substack2 = np.rot90(_substack2,1)[:,::-1,:]
# flipping can be done here ...
combined_piles[cond] = np.nanmean(
np.concatenate([_substack1,_substack2],axis=2),
axis=2
)
# +
gs = plt.GridSpec(nrows=1, ncols=len(conditions) + 1, width_ratios=[20] * len(conditions) + [1])
fig = plt.figure(figsize=(6 * len(conditions), 6))
opts = dict(
norm = MidPointLogNorm(vmin=0.8,vmax=1.5,midpoint=1),
extent=[-snippet_flank//1000, snippet_flank//1000, -snippet_flank//1000, snippet_flank//1000],
# cmap='RdBu_r',
cmap='coolwarm',
interpolation="nearest"
)
for i, cond in enumerate(conditions):
ax = plt.subplot(gs[i])
img = ax.matshow(
combined_piles[cond],
**opts)
ax.xaxis.tick_bottom()
if i > 0:
ax.yaxis.set_visible(False)
plt.title(long_names[cond])
ax = plt.subplot(gs[len(conditions)])
plt.colorbar(img, cax=ax)
plt.suptitle(f'{snippet_flank//1000}kb flank\n'
f'Hi-C resolution = {binsize//1000}kb; # of pairs = {(upstream_pCTCF_pTSS_mask|downstream_mCTCF_mTSS_mask).sum()}')
# -
combined_piles1 = combined_piles
# ### other combination !...
# +
# here is our criteria to select a subgroup of snippets from the stack ....
downstream_mCTCF_pTSS_mask = (arm_sites["type1"]=="tss") & \
(arm_sites["strand1"]=="+") & \
(arm_sites["type2"]=="ctcf") & \
(arm_sites["strand2"]=="-") & \
(arm_sites["distance"]>50_000) & \
(arm_sites["distance"]<500_000)
upstream_pCTCF_mTSS_mask = (arm_sites["type1"]=="ctcf") & \
(arm_sites["strand1"]=="+") & \
(arm_sites["type2"]=="tss") & \
(arm_sites["strand2"]=="-") & \
(arm_sites["distance"]>50_000) & \
(arm_sites["distance"]<500_000)
# when selecting multiple conditions - it is reasonable to assume
# that these conditions are mutually exclusive
# check that the following way:
print("checking if conditions are mutually exclusive ...")
print(f"number of sites that satisfy either condition: {(upstream_pCTCF_mTSS_mask|downstream_mCTCF_pTSS_mask).sum()} ")
print(f"number of sites that satisfy condition1: {upstream_pCTCF_mTSS_mask.sum()} ")
print(f"number of sites that satisfy condition2: {downstream_mCTCF_pTSS_mask.sum()} ")
print(f"the sum {upstream_pCTCF_mTSS_mask.sum()+downstream_mCTCF_pTSS_mask.sum()} of the two latter ones is supposed to be equal to the former one")
combined_piles = {}
for cond in conditions:
# extract only desired snippets from the stack
# stacks are 3D arrays, with first 2 coords along the heatmap
# and the last coord (index) is along snippets (individual interactions from BEDPE) ...
_substack1 = stacks[cond][:,:,upstream_pCTCF_mTSS_mask]
_substack2 = stacks[cond][:,:,downstream_mCTCF_pTSS_mask]
# try to flip _substack2 !
# this is going to be specific in each and every case:
_substack2 = np.rot90(_substack2,1)[:,::-1,:]
# flipping can be done here ...
combined_piles[cond] = np.nanmean(
np.concatenate([_substack1,_substack2],axis=2),
axis=2
)
# +
gs = plt.GridSpec(nrows=1, ncols=len(conditions) + 1, width_ratios=[20] * len(conditions) + [1])
fig = plt.figure(figsize=(6 * len(conditions), 6))
opts = dict(
norm = MidPointLogNorm(vmin=0.8,vmax=1.5,midpoint=1),
extent=[-snippet_flank//1000, snippet_flank//1000, -snippet_flank//1000, snippet_flank//1000],
# cmap='RdBu_r',
cmap='coolwarm',
interpolation="nearest"
)
for i, cond in enumerate(conditions):
ax = plt.subplot(gs[i])
img = ax.matshow(
combined_piles[cond],
**opts)
ax.xaxis.tick_bottom()
if i > 0:
ax.yaxis.set_visible(False)
plt.title(long_names[cond])
ax = plt.subplot(gs[len(conditions)])
plt.colorbar(img, cax=ax)
plt.suptitle(f'{snippet_flank//1000}kb flank\n'
f'Hi-C resolution = {binsize//1000}kb; # of pairs = {(upstream_pCTCF_mTSS_mask|downstream_mCTCF_pTSS_mask).sum()}')
# +
# Quantifying the middle line here ...
# plt.plot(np.nanmean(combined_piles1[conditions[1]][25-2:25+3],axis=0))
# plt.plot(np.nanmean(combined_piles2[conditions[1]][25-2:25+3],axis=0))
# ax = plt.gca()
# ax.set_ylim((1.,1.5))
|
Submission1_notebooks/Dot-pileup-template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# <small><i>This notebook was put together by [<NAME>](http://www.vanderplas.com) for PyCon 2015. Source and license info is on [GitHub](https://github.com/jakevdp/sklearn_pycon2015/).</i></small>
# # 介绍Scikit-Learn:用Python进行机器学习
# 这部分将覆盖基本Scikit-Learn(下面简称sklearn),它是一种Python实现的机器学习工具集。详见:http://scikit-learn.org
# ## 概要
# **主要目标**:介绍机器学习核心概念,和如何使用sklearn包。
# - 机器学习的定义
# - sklearn中的数据表示
# - sklearn API介绍
# ## 关于Scikit-Learn
# [sklearn](http://scikit-learn.org)用Python实现了许多众所周知的机器学习算法,并且提供清晰和成熟的API接口。全世界达数百人参与贡献了sklearn的代码,它在工业和学术界被大量应用。
# sklearn基于[NumPy](www.numpy.org)和[SciPy](http://scipy.org),这两个库在数组处理和科学计算方面很强大。此外sklearn不适合大数据集处理,尽管在这方面已经做了一些[工作](https://github.com/ogrisel/parallel_ml_tutorial)。
# # 什么是机器学习?
#
# 在这部分我们将开始探索基本的机器学习规则。机器学习通过调整参数来学习已知的数据,从而建立模型来预测新的数据。机器学习作为人工智能的一个领域,通过某种程度的泛化使得计算机更加智能。
#
# 这里我们将看一下两个非常简单的例子。第一个是**分类**,图片显示一个二维数据的集合,不同的颜色表示不同的分类。一个分类算法可以划分两种不同颜色的点集:
#
# +
# %matplotlib inline
# set seaborn plot defaults.
# This can be safely commented out
import seaborn; seaborn.set()
# eliminate warnings
def warn(*args, **kwargs):
pass
import warnings
warnings.warn = warn
# -
# Import the example plot from the figures directory
from fig_code import plot_sgd_separator
plot_sgd_separator()
# 这貌似像一个简单的任务,但大道至简。通过画出分割线形成模型,我们可以让这个模型泛化到新的数据集,它可以用来预测分割新的点集(红色和蓝色)。
#
# 如果你想要查看源码,可以使用`%load`命令。
# +
# # %load fig_code/sgd_separator.py
import numpy as np
import matplotlib.pyplot as plt
from sklearn.linear_model import SGDClassifier
from sklearn.datasets.samples_generator import make_blobs
def plot_sgd_separator():
# we create 50 separable points
X, Y = make_blobs(n_samples=50, centers=2,
random_state=0, cluster_std=0.60)
# fit the model
clf = SGDClassifier(loss="hinge", alpha=0.01,
n_iter=200, fit_intercept=True)
clf.fit(X, Y)
# plot the line, the points, and the nearest vectors to the plane
xx = np.linspace(-1, 5, 10)
yy = np.linspace(-1, 5, 10)
X1, X2 = np.meshgrid(xx, yy)
Z = np.empty(X1.shape)
for (i, j), val in np.ndenumerate(X1):
x1 = val
x2 = X2[i, j]
p = clf.decision_function(np.array([[x1, x2]]))
Z[i, j] = p[0]
levels = [-1.0, 0.0, 1.0]
linestyles = ['dashed', 'solid', 'dashed']
colors = 'k'
ax = plt.axes()
ax.contour(X1, X2, Z, levels, colors=colors, linestyles=linestyles)
ax.scatter(X[:, 0], X[:, 1], c=Y, cmap=plt.cm.Paired)
ax.axis('tight')
if __name__ == '__main__':
plot_sgd_separator()
plt.show()
# -
# 下一个我们看下**回归**的例子,数据集的最佳拟合线
from fig_code import plot_linear_regression
plot_linear_regression()
# 我们又看到了一个拟合数据的模型,这个模型也可以泛化到新的数据。这个模型通过学习训练数据,从而预测测试集的结果:给它x值,它可以预测y值。
# ## 数据在Scikit-learn中的表示
# 机器学习通过数据建立模型,基于这点,我们开始讨论,数据如何表示才能让计算机可以理解。我们将举一些matplotlib的例子并且可视化它们。
#
# 在sklearn中的大多数机器学习算法的数据存储在二维的数组或者矩阵中。有的采用``numpy arrays``,还有的采用``scipy.sparse``矩阵。数组的大小设定为`[n_samples, n_features]`
#
# - **n_samples:** 样本的个数,每个样本是待处理的一项数据。它可以是一篇文档,一张图片,一段音频,一段视频,数据库或CSV文件中的某一行,甚至一个你能描述其特征的可量化的集合。
# - **n_features** 特征的数目,用一种可以量化的方式来描述每一个样本的特征数目。它一般是实数,在一些场合下可以为布尔值或者离散的数值。
#
# 特征的数量必须是提前确定的。它可能有很高的维度(例如:上百万的维度),其中一些样本中大部分的特征值为零。在这种场景下scipy.sparse矩阵就很有用了,它比numpy arrays更加节约内存。
#
# ## 一个简单的例子:Iris(鸢尾花)数据集
# 在这个例子中,我们将看一下存储在sklearn中的iris数据。这个数据集包含三种不同种类的iris测量值。让我们看下下图:
# +
from IPython.core.display import Image, display
display(Image(filename='images/iris_setosa.jpg'))
print("Iris Setosa\n")
display(Image(filename='images/iris_versicolor.jpg'))
print("Iris Versicolor\n")
display(Image(filename='images/iris_virginica.jpg'))
print("Iris Virginica")
# -
# ### 问题:
# **如果我们需要一个算法来识别不同种类的iris(鸢尾花),需要哪些数据?**
#
# 我们需要一个`[n_samples x n_features]`的二维数组
#
# - `n_samples`表示什么?
# - `n_features`表示什么?
#
# 每一个样本必须有确定数量的特征,每个特征都是样本的某种量化值。
#
# ### 从sklearn中加载iris数据集
#
#
# sklearn中包含了这些种类的iris数据集。这数据集包含下面的特征值:
#
# - 数据集中的特征:
# 1. 花萼长度(单位cm)
# 2. 花萼宽度(单位cm)
# 3. 花瓣长度(单位cm)
# 4. 花瓣宽度(单位cm)
#
# - 预测的目标分类:
# 1. 山鸢尾(Iris Setosa)
# 2. 杂色鸢尾(Iris Versicolour)
# 3. 维吉尼亚鸢尾(Iris Virginica)
#
# ``sklearn``包含iris的CSV文件包括加载到numpy arrays的函数:
from sklearn.datasets import load_iris
iris = load_iris()
iris.keys()
n_samples, n_features = iris.data.shape
print((n_samples, n_features))
print(iris.data[0])
print(iris.data.shape)
print(iris.target.shape)
print(iris.target)
# 这个数据集是四维的,但是我们可以用scatter plot每次展示两维:
# +
import numpy as np
import matplotlib.pyplot as plt
x_index = 0
y_index = 1
# this formatter will label the colorbar with the correct target names
formatter = plt.FuncFormatter(lambda i, *args: iris.target_names[int(i)])
plt.scatter(iris.data[:, x_index], iris.data[:, y_index],
c=iris.target, cmap=plt.cm.get_cmap('RdYlBu', 3))
plt.colorbar(ticks=[0, 1, 2], format=formatter)
plt.clim(-0.5, 2.5)
plt.xlabel(iris.feature_names[x_index])
plt.ylabel(iris.feature_names[y_index]);
# -
# ### 练习
#
# 改变上面代码中`x_index`和`y_index`值,找出一个点可以最大化的分割这三类
#
# 这是个**降维**的练习,我们稍后会见到。
# ## 其他可用的数据
#
# 有以下的三种形式:
#
# - **sklearn包数据** 在安装的时候,这些小的数据集被打包在sklearn中,通过``sklearn.datasets.load_*``函数能够被加载
# - **下载的数据** 这些大的数据集可以通过sklearn提供的``sklearn.datasets.fetch_*``函数从网上下载
# - **生成的数据** 这些数据可以通过sklearn提供的``sklearn.datasets.make_*``函数,基于一个随机种子从模型中产生
#
# 你能通过IPython的tab补全功能,浏览这些函数。在从sklearn中导入``datasets``子模块后,你可以
# 输入datasets.load_ + TAB或datasets.fetch_ + TAB或datasets.make_ + TAB来浏览这些函数列表。
from sklearn import datasets
# +
# Type datasets.fetch_<TAB> or datasets.load_<TAB> in IPython to see all possibilities
# datasets.fetch_
# +
# datasets.load_
# -
# 在下一章,我们将使用这些数据集,并且学习机器学习的基本方法。
|
notebooks/02.1-Machine-Learning-Intro-zh.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Modelos de reapertura modular
# ### Paquetes
import networkx as nx
import matplotlib.pyplot as plt
import EoN
from matplotlib import rc
import scipy
import numpy as np
import random
import pandas as pd
import numpy as np
from collections import defaultdict
from pathlib import Path
# ### Definir parámetros epidemiológicos
# ##### Leer las redes para tener el tamaño
g = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_cdmx_infomap.graphml")) #red cdmx
g_susana = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_susana.graphml"))
# ### Número de casos
# +
recuperados = 34849
infectados = 10596
poblacion_real = 9000000
poblacion_escalada = len(g.nodes())
recuperados_escalados = round(recuperados * poblacion_escalada / poblacion_real)
infectados_escalados = round(infectados * poblacion_escalada / poblacion_real)
print(recuperados_escalados)
print(infectados_escalados)
#infectados_totales = round(100*infectados_escalados/10)
#recuperados_totales = round(100*recuperados_escalados/10)
infectados_totales = 3*infectados_escalados
recuperados_totales = 3*recuperados_escalados
#infectados_totales = infectados_escalados
#recuperados_totales = recuperados_escalados
print(infectados_totales)
print(recuperados_totales)
# -
# #### Función para preparar la red para modelo epidemiológico
def funcion_preparadora(nw):
nw = nw.copy()
#Stochastic transition E-I
ei_node_attribute = {node: random.uniform(a=0.5, b=2) for node in nw.nodes()}
#Stochastic transition I-R
#will multiply recovery rate for some factor so it is between 14 and 28 days
ir_node_attribute = {node: random.randint(a=1, b=2) for node in nw.nodes()}
#Transmission weight - varies for each pair of interactions (some contacts are very transmisive, others not so)
edge_attribute_dict = {edge: 0.5+random.random() for edge in nw.edges()}
nx.set_node_attributes(nw, values=ei_node_attribute, name='expose2infect_weight')
nx.set_node_attributes(nw, values=ir_node_attribute, name='infect2recove_weight')
nx.set_edge_attributes(nw, values=edge_attribute_dict, name='transmission_weight')
return(nw)
# ##### Definir los parámetros de transmisión y recuperación
# +
### rates
ei_rate = 1/1.5 # so that minimum rate is 12 hours, maximum is 72
ir_rate = 1/(14) # so minimum time is 14 days, and maximum time is 28 days
#transmission_rate = 0.155 #old SEIRSplus
#transmission_rate = 0.030
#transmission_rate = 0.00125
#transmission_rate = 0.0025
#transmission_rate = 0.0040
#this represents the baseline transmission. Let's use the one we used with SEIRSplus
#SIGMA = 1/5.2
#GAMMA = 1/10
GAMMA = 1/14
#MU_I = 0.002
R0 = 2.5
transmission_rate = 1/(1/GAMMA) * R0
Rt = 0.995055947438355
transmission_rate = 1/(1/GAMMA) * Rt
#transmission_rate = 0.125/2
print(transmission_rate)
# -
# ##### Definir la red de transiciones - describe la dinámica epidémica
# +
#the node intrinsic transitions
H = nx.DiGraph()
H.add_node('S') #This line is actually unnecessary since 'S' does not change status intrinsically
#
H.add_edge('E', 'I', rate = ei_rate, weight_label='expose2infect_weight')
#
H.add_edge('I', 'R', rate = ir_rate, weight_label ='infect2recove_weight')
#interaction depending transitions
J = nx.DiGraph()
J.add_edge(('I', 'S'), ('I', 'E'), rate = transmission_rate, weight_label='transmission_weight')
# The line above states that an 'I' individual will cause an 'S' individual
# to transition to 'E' with rate equal to 0.1 times the partnership's attribute
# 'transmission_weight'.
# -
# ###### Define initial states
# +
IC = defaultdict(lambda: 'S')
for node in range(infectados_totales-1):
IC[node] = 'I'
for node in range(infectados_totales, recuperados_totales):
IC[node] = 'R'
return_statuses = ('S', 'E', 'I', 'R')
# -
# ### Modelo con Reapertura Total
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g)
nomen = "reapertura_total"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# ### Modelo Con Parámetros de JNSD
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_susana)
nomen = "jnsd"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# ## Evaluar Reapertura aleatoria, diferentes niveles de población reintegrándose
g_05 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reactivate_05percent.graphml"))
g_10 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reactivate_10percent.graphml"))
g_15 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reactivate_15percent.graphml"))
g_20 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reactivate_20percent.graphml"))
g_25 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reactivate_25percent.graphml"))
g_50 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reactivate_50percent.graphml"))
# #### 5%
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_05)
nomen = "random_05"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# #### 10%
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_10)
nomen = "random_10"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# #### 15%
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_15)
nomen = "random_15"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# #### 20%
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_20)
nomen = "random_20"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# #### 25%
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_25)
nomen = "random_25"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# #### 50%
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_50)
nomen = "random_50"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# ## Evaluar reapertura modular
g_m1 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_m1.graphml")) #2mods
g_m2 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_m2.graphml")) #3mods
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_m1)
nomen = "m1"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_m2)
nomen = "m2"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# ### Reapertura modular con mayor carga
g_m2x = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_m3.graphml")) #2X
g_m3x = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_m4.graphml")) #3X
g_m4x = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_m6.graphml")) #4X
g_m5x = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_m5.graphml")) #4X
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_m2x)
nomen = "m2X"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_m3x)
nomen = "m3X"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_m4x)
nomen = "m4X"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_m5x)
nomen = "m5X"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# ### Smart module selection
# Here, we pick small modules...
#20% population using a smart module selection
g_smartmod_01 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_smartmod_01.graphml")) #2X
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_smartmod_01)
nomen = "smartmod_01"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
# ### Comparar reapertura smartmod vs not smart mod al 50% de la población
#20% population using a smart module selection
g_yessmartmod_50 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_yessmart_mod50.graphml"))
g_notsmartmod_50 = nx.convert_node_labels_to_integers(nx.read_graphml("results/redes_eon/g_reapertura_nonsmart_mod50.graphml"))
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_yessmartmod_50)
nomen = "smartmod_50"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# +
np.random.seed(725)
random.seed(725)
prep_g = funcion_preparadora(g_notsmartmod_50)
nomen = "m50"
pathnomen = "results/" + nomen
Path(pathnomen).mkdir(parents=True, exist_ok=True)
for i in range(0, 100):
print(i)
np.random.seed(i)
random.seed(i)
outnam = pathnomen + "/" + nomen + "_" + str(i) + ".csv"
print(outnam)
mod_nuloAbs = EoN.Gillespie_simple_contagion(G=prep_g,
spontaneous_transition_graph=H,
nbr_induced_transition_graph=J,
IC=IC,
return_statuses=return_statuses,
return_full_data=True,
tmax=1000
)
my_result = mod_nuloAbs
my_result.summary()[0]
predata = my_result.summary()[1]
predata["t"] = my_result.summary()[0]
df = pd.DataFrame(predata)
df = df.assign(I_pc=100*df['I']/poblacion_escalada)
df = df.assign(R_pc=100*df['R']/poblacion_escalada)
df.to_csv(outnam)
# -
|
ReaperturaModular_V2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Задание 5.2 - Word2Vec with Negative Sampling
#
# В этом задании мы натренируем свои версию word vectors с negative sampling на том же небольшом датасете.
#
#
# Несмотря на то, что основная причина использования Negative Sampling - улучшение скорости тренировки word2vec, в нашем игрушечном примере мы **не требуем** улучшения производительности. Мы используем negative sampling просто как дополнительное упражнение для знакомства с PyTorch.
#
# Перед запуском нужно запустить скрипт `download_data.sh`, чтобы скачать данные.
#
# Датасет и модель очень небольшие, поэтому это задание можно выполнить и без GPU.
#
#
# +
import torch
import torch.nn as nn
import torch.optim as optim
from torch.utils.data import Dataset
from torchvision import transforms
import numpy as np
import matplotlib.pyplot as plt
# We'll use Principal Component Analysis (PCA) to visualize word vectors,
# so make sure you install dependencies from requirements.txt!
from sklearn.decomposition import PCA
# %matplotlib inline
# +
import os
import numpy as np
class StanfordTreeBank:
'''
Wrapper for accessing Stanford Tree Bank Dataset
https://nlp.stanford.edu/sentiment/treebank.html
Parses dataset, gives each token and index and provides lookups
from string token to index and back
Allows to generate random context with sampling strategy described in
word2vec paper:
https://papers.nips.cc/paper/5021-distributed-representations-of-words-and-phrases-and-their-compositionality.pdf
'''
def __init__(self):
self.index_by_token = {} # map of string -> token index
self.token_by_index = []
self.sentences = []
self.token_freq = {}
self.token_reject_by_index = None
def load_dataset(self, folder):
filename = os.path.join(folder, "datasetSentences.txt")
with open(filename, "r", encoding="latin1") as f:
l = f.readline() # skip the first line
for l in f:
splitted_line = l.strip().split()
words = [w.lower() for w in splitted_line[1:]] # First one is a number
self.sentences.append(words)
for word in words:
if word in self.token_freq:
self.token_freq[word] +=1
else:
index = len(self.token_by_index)
self.token_freq[word] = 1
self.index_by_token[word] = index
self.token_by_index.append(word)
self.compute_token_prob()
def compute_token_prob(self):
words_count = np.array([self.token_freq[token] for token in self.token_by_index])
words_freq = words_count / np.sum(words_count)
# Following sampling strategy from word2vec paper
self.token_reject_by_index = 1- np.sqrt(1e-5/words_freq)
def check_reject(self, word):
return np.random.rand() > self.token_reject_by_index[self.index_by_token[word]]
def get_random_context(self, context_length=5):
"""
Returns tuple of center word and list of context words
"""
sentence_sampled = []
while len(sentence_sampled) <= 2:
sentence_index = np.random.randint(len(self.sentences))
sentence = self.sentences[sentence_index]
sentence_sampled = [word for word in sentence if self.check_reject(word)]
center_word_index = np.random.randint(len(sentence_sampled))
words_before = sentence_sampled[max(center_word_index - context_length//2,0):center_word_index]
words_after = sentence_sampled[center_word_index+1: center_word_index+1+context_length//2]
return sentence_sampled[center_word_index], words_before+words_after
def num_tokens(self):
return len(self.token_by_index)
data = StanfordTreeBank()
data.load_dataset("./stanfordSentimentTreebank/")
print("Num tokens:", data.num_tokens())
for i in range(5):
center_word, other_words = data.get_random_context(5)
print(center_word, other_words)
# -
# # Dataset для Negative Sampling должен быть немного другим
#
# Как и прежде, Dataset должен сгенерировать много случайных контекстов и превратить их в сэмплы для тренировки.
#
# Здесь мы реализуем прямой проход модели сами, поэтому выдавать данные можно в удобном нам виде.
# Напоминаем, что в случае negative sampling каждым сэмплом является:
# - вход: слово в one-hot представлении
# - выход: набор из одного целевого слова и K других случайных слов из словаря.
# Вместо softmax + cross-entropy loss, сеть обучается через binary cross-entropy loss - то есть, предсказывает набор бинарных переменных, для каждой из которых функция ошибки считается независимо.
#
# Для целевого слова бинарное предсказание должно быть позитивным, а для K случайных слов - негативным.
#
# Из набора слово-контекст создается N сэмплов (где N - количество слов в контексте), в каждом из них K+1 целевых слов, для только одного из которых предсказание должно быть позитивным.
# Например, для K=2:
#
# Слово: `orders` и контекст: `['love', 'nicest', 'to', '50-year']` создадут 4 сэмпла:
# - input: `orders`, target: `[love: 1, any: 0, rose: 0]`
# - input: `orders`, target: `[nicest: 1, fool: 0, grass: 0]`
# - input: `orders`, target: `[to: 1, -: 0, the: 0]`
# - input: `orders`, target: `[50-year: 1, ?: 0, door: 0]`
#
# Все слова на входе и на выходе закодированы через one-hot encoding, с размером вектора равным количеству токенов.
# +
num_negative_samples = 10
class Word2VecNegativeSampling(Dataset):
'''
PyTorch Dataset for Word2Vec with Negative Sampling.
Accepts StanfordTreebank as data and is able to generate dataset based on
a number of random contexts
'''
def __init__(self, data, num_negative_samples, num_contexts=30000):
'''
Initializes Word2VecNegativeSampling, but doesn't generate the samples yet
(for that, use generate_dataset)
Arguments:
data - StanfordTreebank instace
num_negative_samples - number of negative samples to generate in addition to a positive one
num_contexts - number of random contexts to use when generating a dataset
'''
self.data = data
self.num_negative_samples = num_negative_samples
self.num_contexts = num_contexts
self.samples = []
self.num_tokens = data.num_tokens()
self.word_to_index = {}
self.index_to_word = {}
self.number = 0
# TODO: Implement what you need for other methods!
def generate_dataset(self):
'''
Generates dataset samples from random contexts
Note: there will be more samples than contexts because every context
can generate more than one sample
'''
self.samples.clear()
for i in range(self.num_contexts):
center_w, other_w = self.data.get_random_context()
if center_w not in self.word_to_index:
self.word_to_index[center_w] = int(self.number)
self.index_to_word[self.number] = center_w
self.number += 1
for word in other_w:
if word not in self.word_to_index:
self.word_to_index[word] = int(self.number)
self.index_to_word[self.number] = word
self.number += 1
self.samples.append((self.word_to_index[center_w], self.word_to_index[word]))
# TODO: Implement generating the dataset
# You should sample num_contexts contexts from the data and turn them into samples
# Note you will have several samples from one context
def __len__(self):
'''
Returns total number of samples
'''
return len(self.samples)
# TODO: Return the number of samples
def __getitem__(self, index):
'''
Returns i-th sample
Return values:
input_vector - index of the input word (not torch.Tensor!)
output_indices - torch.Tensor of indices of the target words. Should be 1+num_negative_samples.
output_target - torch.Tensor with float targets for the training. Should be the same size as output_indices
and have 1 for the context word and 0 everywhere else
'''
item = self.samples[index]
input_vector = item[0]
neg_samples = np.random.choice(self.num_tokens, self.num_negative_samples, replace=False)
while item[1] in neg_samples:
neg_samples = np.random.choice(self.num_tokens, self.num_negative_samples, replace=False)
output_indices = torch.cat((torch.Tensor(neg_samples), torch.Tensor([item[1]])), 0)
output_target = torch.zeros_like(output_indices)
output_indices = output_indices.type(torch.LongTensor)
output_target = output_target.type(torch.FloatTensor)
output_target[-1] = 1
return input_vector, output_indices, output_target
# TODO: Generate tuple of 3 return arguments for i-th sample
dataset = Word2VecNegativeSampling(data, num_negative_samples, 10)
input_vector, output_indices, output_target = dataset[0]
print("Sample - input: %s, output indices: %s, output target: %s" % (int(input_vector), output_indices, output_target)) # target should be able to convert to int
assert isinstance(output_indices, torch.Tensor)
assert output_indices.shape[0] == num_negative_samples+1
assert isinstance(output_target, torch.Tensor)
assert output_target.shape[0] == num_negative_samples+1
assert torch.sum(output_target) == 1.0
# -
# # Создаем модель
#
# Для нашей задачи нам придется реализовать свою собственную PyTorch модель.
# Эта модель реализует свой собственный прямой проход (forward pass), который получает на вход индекс входного слова и набор индексов для выходных слов.
#
# Как всегда, на вход приходит не один сэмпл, а целый batch.
# Напомним, что цели улучшить скорость тренировки у нас нет, достаточно чтобы она сходилась.
# +
# Create the usual PyTorch structures
dataset = Word2VecNegativeSampling(data, num_negative_samples, 30000)
dataset.generate_dataset()
# As before, we'll be training very small word vectors!
wordvec_dim = 10
class Word2VecNegativeSamples(nn.Module):
def __init__(self, num_tokens):
super(Word2VecNegativeSamples, self).__init__()
self.input = nn.Linear(num_tokens, 10, bias=False)
self.output = nn.Linear(10, num_tokens, bias=False)
def forward(self, input_index_batch, output_indices_batch):
'''
Implements forward pass with negative sampling
Arguments:
input_index_batch - Tensor of ints, shape: (batch_size, ), indices of input words in the batch
output_indices_batch - Tensor if ints, shape: (batch_size, num_negative_samples+1),
indices of the target words for every sample
Returns:
predictions - Tensor of floats, shape: (batch_size, um_negative_samples+1)
'''
results = []
for i in range(input_index_batch.shape[0]):
in_w = self.input.weight[:, input_index_batch[i]]
out_w = self.output.weight[output_indices_batch[i]].T
results.append(in_w @ out_w)
return torch.stack(results)
# TODO Implement forward pass
# Hint: You can use for loop to go over all samples on the batch,
# run every sample indivisually and then use
# torch.stack or torch.cat to produce the final result
nn_model = Word2VecNegativeSamples(data.num_tokens())
nn_model.type(torch.FloatTensor)
# +
def extract_word_vectors(nn_model):
'''
Extracts word vectors from the model
Returns:
input_vectors: torch.Tensor with dimensions (num_tokens, num_dimensions)
output_vectors: torch.Tensor with dimensions (num_tokens, num_dimensions)
'''
# TODO: Implement extracting word vectors from param weights
# return tuple of input vectors and output vectos
return nn_model.input.weight.clone().T, nn_model.output.weight.clone()
untrained_input_vectors, untrained_output_vectors = extract_word_vectors(nn_model)
assert untrained_input_vectors.shape == (data.num_tokens(), wordvec_dim)
assert untrained_output_vectors.shape == (data.num_tokens(), wordvec_dim)
# -
def train_neg_sample(model, dataset, train_loader, optimizer, scheduler, num_epochs):
'''
Trains word2vec with negative samples on and regenerating dataset every epoch
Returns:
loss_history, train_history
'''
loss = nn.BCEWithLogitsLoss().type(torch.FloatTensor)
loss_history = []
train_history = []
for epoch in range(num_epochs):
model.train() # Enter train mode
dataset.generate_dataset()
loss_accum = 0
correct_samples = 0
total_samples = 0
for i_step, (x, y, z) in enumerate(train_loader):
prediction = model(x, y)
loss_value = loss(prediction, z)
optimizer.zero_grad()
loss_value.backward()
optimizer.step()
correct_samples += torch.sum(((prediction > 0) == z).all(dim=1))
total_samples += z.shape[0]
loss_accum += loss_value
ave_loss = loss_accum / i_step
train_accuracy = float(correct_samples) / total_samples
scheduler.step()
loss_history.append(float(ave_loss))
train_history.append(train_accuracy)
# TODO: Implement training using negative samples
# You can estimate accuracy by comparing prediction values with 0
# And don't forget to step the scheduler!
print("Average loss: %f, Train accuracy: %f" % (ave_loss, train_accuracy))
return loss_history, train_history
# # Ну и наконец тренировка!
#
# Добейтесь значения ошибки меньше **0.25**.
# +
# Finally, let's train the model!
# TODO: We use placeholder values for hyperparameters - you will need to find better values!
optimizer = optim.Adam(nn_model.parameters(), lr=1e-2, weight_decay=0)
scheduler = optim.lr_scheduler.CosineAnnealingWarmRestarts(optimizer, 5, T_mult=2)
train_loader = torch.utils.data.DataLoader(dataset, batch_size=60, drop_last=True)
loss_history, train_history = train_neg_sample(nn_model, dataset, train_loader, optimizer, scheduler, 15)
# -
# Visualize training graphs
plt.subplot(211)
plt.plot(train_history)
plt.subplot(212)
plt.plot(loss_history)
# # Визуализируем вектора для разного вида слов до и после тренировки
#
# Как и ранее, в случае успешной тренировки вы должны увидеть как вектора слов разных типов (например, знаков препинания, предлогов и остальных)
# +
trained_input_vectors, trained_output_vectors = extract_word_vectors(nn_model)
assert trained_input_vectors.shape == (data.num_tokens(), wordvec_dim)
assert trained_output_vectors.shape == (data.num_tokens(), wordvec_dim)
def visualize_vectors(input_vectors, output_vectors, title=''):
full_vectors = torch.cat((input_vectors, output_vectors), 0)
wordvec_embedding = PCA(n_components=2).fit_transform(full_vectors.detach().numpy())
# Helpful words form CS244D example
# http://cs224d.stanford.edu/assignment1/index.html
visualize_words = {'green': ["the", "a", "an"],
'blue': [",", ".", "?", "!", "``", "''", "--"],
'brown': ["good", "great", "cool", "brilliant", "wonderful",
"well", "amazing", "worth", "sweet", "enjoyable"],
'orange': ["boring", "bad", "waste", "dumb", "annoying", "stupid"],
'red': ['tell', 'told', 'said', 'say', 'says', 'tells', 'goes', 'go', 'went']
}
plt.figure(figsize=(7,7))
plt.suptitle(title)
for color, words in visualize_words.items():
points = np.array([wordvec_embedding[data.index_by_token[w]] for w in words])
for i, word in enumerate(words):
plt.text(points[i, 0], points[i, 1], word, color=color,horizontalalignment='center')
plt.scatter(points[:, 0], points[:, 1], c=color, alpha=0.3, s=0.5)
visualize_vectors(untrained_input_vectors, untrained_output_vectors, "Untrained word vectors")
visualize_vectors(trained_input_vectors, trained_output_vectors, "Trained word vectors")
# -
|
assignments/assignment5/NegativeSampling.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from __future__ import print_function, division
import numpy as np
import pandas as pd
import sys
from PIL import Image
import torch
import torch.nn as nn
import torch.nn.functional as F
from torch.autograd import Variable
import torch.utils.data
import torchvision
import spatial_vae.models as models
####data loading and making coordinates#####
mnist_test = np.load('D:/project/CMU/toytask/mnist_rotated/images_test.npy')
# img = Image.fromarray(mnist_test[55], 'L')
# img.save('my.png')
# img.show()
mnist_test = torch.from_numpy(mnist_test).float()/255 #normalized
n=m=28
xgrid = np.linspace(-1, 1, m)
#gives a list that contains m=28 evenly spaced values between -1 and 1
ygrid = np.linspace(1, -1, n)
x0,x1 = np.meshgrid(xgrid, ygrid)
#each point in y is paired with a point in x
x_coord = np.stack([x0.ravel(), x1.ravel()], 1)# xo.ravel rolls the 2d array x0
x_coord = torch.from_numpy(x_coord).float()#784 points each represented by an x,y pair
#tensor of x and y coordinates each corresponding to one of the 784 pixel values
print(x_coord.shape)
# -
# 2 list of evenly spaced numbers are created. Each number in 1list is associated with all the other numbers in another list
# A coordinate frame is created, it has 784 points intended for 784 pixel values. Each point has 2 coordinate values
y_test = mnist_test.view(-1, n*m)#image pixel values for 10k flattened images
print(y_test.shape)
use_cuda = torch.cuda.is_available()#check gpu availability and move data to gpu
if use_cuda:
# y_train = y_train.cuda()
y_test = y_test.cuda()
x_coord = x_coord.cuda()
# x_coord is our cartesian frame having 2 coordinates for 784 points per image
# and y_test is our image data having 784 pixel values per image
# +
data_test = torch.utils.data.TensorDataset(y_test)
#alternative to this would be a customized dataset class
import sys
import torch.nn as nn
#latent value for
z_dim = 2
print('# training with z-dim:', z_dim, file=sys.stderr)#latent values
num_layers = 2
hidden_dim = 500
activation = nn.Tanh
print('# using the spatial generator architecture', file=sys.stderr)
# -
# Inside the model arch z_dim is increased to 5 if rotational and translation inferences are to be done
# Translation has 2 latent dims associated with it, possibly for accounting it in x0 and x1 directions
# Rotation has only 1 because angular rotation can happen only in x0 x1 plane
# +
ckpt_path1 = 'D:\\project\\CMU\\toytask\\spatial-VAE-master\\saved_models_generator_epoch2.sav'
ckpt_path2 = 'D:\\project\\CMU\\toytask\\spatial-VAE-master\\saved_models_inference_epoch2.sav'
#load the models
#pnet is the spatial vae and qnet is the inference network
p_net = torch.load(ckpt_path1)
q_net=torch.load(ckpt_path2)
#loaded the models including the model class structure etc because it was saved in the original script using torch.save
#good practice for saving and loading is to just save and load the state dicts because it doesnt deal with hard coded info
#and is flexible
if use_cuda:
p_net.cuda()
q_net.cuda()
dx_scale = 0.1
theta_prior = np.pi/4
#prior assumption about the values of rotation and translation
print('# using priors: theta={}, dx={}'.format(theta_prior, dx_scale), file=sys.stderr)
# -
# priors are our assumptions about how much rotation or translation is expected
# +
N = len(mnist_test)
params = list(p_net.parameters()) + list(q_net.parameters())
lr = 1e-4
optim = torch.optim.Adam(params, lr=lr)
#define optimizer
minibatch_size = 100
test_iterator = torch.utils.data.DataLoader(data_test, batch_size=minibatch_size)
#data loader for dynamic loading
# -
# To be read in the order 1. eval_model 2. eval_minibatch. In eval_minibatch, we first use the inference network to get the latent distribution of the image using its higher representations. Then we do rotation and translation operations on our pre fixed coordinate/cartesian system by using samples from the inferenced latent distribution as the other operand in respective tasks. This establishes a relationship of dependence between transformed cartesian coordinates and latent distribution, which in my opinion explicitly reflects changes in one into the other. Using the transformed coordinates we reconstruct our image rotated and translated using the spatial vae model. The spatial vae model has learned a function that directly takes coordinate values and latent values to form the desired image.
#
def eval_minibatch(x, y, p_net, q_net, rotate=True, translate=True, dx_scale=0.1, theta_prior=np.pi, use_cuda=False):
#y is a batch of images , x is the set of coordinates in cartesian system
b = y.size(0)#batch size
x = x.expand(b, x.size(0), x.size(1))#784pixels,2coords for each image in a batch of 100
#does some replication of values to give a new view of tensor although the original tensor remains same
# first do inference on the latent variables
if use_cuda:
y = y.cuda()
z_mu,z_logstd = q_net(y)
#get an estimate about the mean and std dev, and thus distriution,of latent vars, establishes a relation between latent variables and images
z_std = torch.exp(z_logstd) #exponential the log
z_dim = z_mu.size(1)
#z=5, 2 are unstructured,1 for translational, 2 for rotational ??idk why maybe they require 2 for mean n std
# draw samples from variational posterior to calculate
# E[p(x|z)]
r = Variable(x.data.new(b,z_dim).normal_())#find the difference between x.new and x.data.new
#creates an autograd tensor from a gaussian distr with mean 0 and std 1 havin the same data type as X and
#shape as X's view in 100,5
# 100,5 tensor initialized with normal distribution; autograd tensor
#each point his sampled from this random distribution and projected to the latent distribution
z = z_std*r + z_mu
#latent vector
kl_div = 0
if rotate:
# z[0] is the rotation
theta_mu = z_mu[:,0]#theta_mu and other theta variables are added to the computational graph
theta_std = z_std[:,0]
theta_logstd = z_logstd[:,0]
theta = z[:,0]
z = z[:,1:]#isolate the latent variable of rotation from others
z_mu = z_mu[:,1:]
z_std = z_std[:,1:]
z_logstd = z_logstd[:,1:]
# calculate rotation matrix
rot = Variable(theta.data.new(b,2,2).zero_())#get zero tensors having same type as theta and size as mentioned
rot[:,0,0] = torch.cos(theta)
rot[:,0,1] = torch.sin(theta)
rot[:,1,0] = -torch.sin(theta)
rot[:,1,1] = torch.cos(theta)
#to do the coordinate transformation by rotation
x = torch.bmm(x, rot) # rotate coordinates by theta
# calculate the KL divergence term
sigma = theta_prior
#loss term for inference on latent
kl_div = -theta_logstd + np.log(sigma) + (theta_std**2 + theta_mu**2)/2/sigma**2 - 0.5
if translate:
# z[0,1] are the translations
dx_mu = z_mu[:,:2]
dx_std = z_std[:,:2]
dx_logstd = z_logstd[:,:2]
dx = z[:,:2]*dx_scale # scale dx by standard deviation
dx = dx.unsqueeze(1)
z = z[:,2:]
x = x + dx # translate coordinates
# reconstruct the image by making it depend on 784 rotated and translated cartesian coordinates + latent variables
#784coords+5latent_vars input gives 784 pixel values
y_hat = p_net(x.contiguous(), z)
y_hat = y_hat.view(b, -1)
# y_hat1 = y_hat.reshape(b,28,28).cpu().detach().numpy()
# img = Image.fromarray(y_hat1[55], 'L')
# img.save('my.png')
# img.show()
size = y.size(1)
log_p_x_g_z = -F.binary_cross_entropy_with_logits(y_hat, y)*size
z_kl = -z_logstd + 0.5*z_std**2 + 0.5*z_mu**2 - 0.5
kl_div = kl_div + torch.sum(z_kl, 1)
kl_div = kl_div.mean()
elbo = log_p_x_g_z - kl_div
return elbo, log_p_x_g_z, kl_div
def eval_model(iterator, x_coord, p_net, q_net, rotate=True, translate=True
, dx_scale=0.1, theta_prior=np.pi, use_cuda=False):
p_net.eval()
q_net.eval()
#iterator batch of 10k images 784 pixels each, x_coord is cartesian system 784 points with xandy cords
c = 0
gen_loss_accum = 0
kl_loss_accum = 0
elbo_accum = 0
for y, in iterator:
b = y.size(0)
x = Variable(x_coord)# 784 points with 2 coordinates each
y = Variable(y) #batchsize,100 images 784 pixel values each
# print(x.shape,y.shape)
elbo, log_p_x_g_z, kl_div = eval_minibatch(x, y, p_net, q_net, rotate=rotate, translate=translate
, dx_scale=dx_scale, theta_prior=theta_prior
, use_cuda=use_cuda)
elbo = elbo.item() #detaches tensors/losses from the computational graph so that they dont burden the computational processes
gen_loss = -log_p_x_g_z.item()
kl_loss = kl_div.item()
c += b
delta = b*(gen_loss - gen_loss_accum)
gen_loss_accum += delta/c
delta = b*(elbo - elbo_accum)
elbo_accum += delta/c
delta = b*(kl_loss - kl_loss_accum)
kl_loss_accum += delta/c
return elbo_accum, gen_loss_accum, kl_loss_accum
# +
output = sys.stdout
print('\t'.join(['Epoch', 'Split', 'ELBO', 'Error', 'KL']), file=output)
elbo_accum,gen_loss_accum,kl_loss_accum = eval_model(test_iterator, x_coord, p_net,q_net, rotate=True, translate=True,dx_scale=dx_scale, theta_prior=theta_prior,use_cuda=use_cuda)
line = '\t'.join(['test', str(elbo_accum), str(gen_loss_accum), str(kl_loss_accum)])
print(line, file=output)
output.flush()
# -
|
test_notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# name: python3
# ---
# # Australian Men's Basketball Tokyo 2020
#
# Comparison of historical efficiency with latest value in Tokyo 2020 Olympics.
#
# + id="O8htI4mYkWzO" executionInfo={"status": "ok", "timestamp": 1614030628784, "user_tz": -660, "elapsed": 2244, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}}
import matplotlib.pyplot as plt
import pandas as pd
import seaborn as sns
# -
def compute_epg(da):
# missing TO
# per-game & percentages available only
da['EFF.Calc'] = da['Pts'] + da['Reb'] + da['Ast'] + da['Stl'] + da['Blk'] + da['FG%']/100.0 + da['3%']/100.0 + da['1%']/100
return(da)
# ## Historical stats
# + colab={"base_uri": "https://localhost:8080/"} id="OKKe2js8h62G" executionInfo={"status": "ok", "timestamp": 1614030673842, "user_tz": -660, "elapsed": 25792, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}} outputId="afb9dc89-5b52-4a51-c661-2530cdead667"
# authenticate
# from google.colab import drive
# drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="G__slQ02iHa8" executionInfo={"status": "ok", "timestamp": 1614030679348, "user_tz": -660, "elapsed": 1009, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}} outputId="a20364f3-d688-4423-8e27-51e587d41965"
# df_players = pd.read_excel('/content/drive/My Drive/AusBoomers/aus_boomers_players.xlsx')
# df_players.head()
# +
df_players = pd.read_excel('../Data/aus_boomers_players.xlsx')
# Filter players to final team selection
df_players = df_players.loc[df_players['Selected'] == 'Y']
# Select first 7 rows
df_players = df_players.iloc[:, :7]
df_players.head()
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="CdzQFDQ2iSnN" executionInfo={"status": "ok", "timestamp": 1614030691569, "user_tz": -660, "elapsed": 1008, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}} outputId="3ae56ef7-36ca-4f49-a658-3b9d2f5359ee"
# df_stats = pd.read_excel('/content/drive/My Drive/AusBoomers/aus_boomers_proballer_stats.xlsx')
# df_stats.head()
# -
df_stats = pd.read_excel('../Data/aus_boomers_proballer_stats.xlsx')
df_stats.head()
df_stats.info()
# + id="6z3qq62pze1J" executionInfo={"status": "ok", "timestamp": 1614030699545, "user_tz": -660, "elapsed": 743, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}}
# how does published Eff compare with ad-hoc EFF.Calc value?
da = compute_epg(df_stats)
# + id="MZc3GPFhMBF2" executionInfo={"status": "ok", "timestamp": 1614030710650, "user_tz": -660, "elapsed": 748, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}}
# construct year from season label
da['Year'] = "20" + da['Season'].str[:2]
da['Year'] = pd.to_numeric(da['Year'])
# + colab={"base_uri": "https://localhost:8080/", "height": 289} id="CLG7o1fo0kAQ" executionInfo={"status": "ok", "timestamp": 1614030713668, "user_tz": -660, "elapsed": 790, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}} outputId="45479130-fedd-4894-c025-e0183f662e72"
da.head()
# + colab={"base_uri": "https://localhost:8080/"} id="rLDTVnjr4EcZ" executionInfo={"status": "ok", "timestamp": 1614030723629, "user_tz": -660, "elapsed": 731, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}} outputId="f2faa7ac-3fc1-49f0-8b62-a0081f30f746"
dh = da[da['Source']=="list-regular"].Name.unique()
len(dh)
# + id="XGxTYwkc2lZK" executionInfo={"status": "ok", "timestamp": 1614030735411, "user_tz": -660, "elapsed": 741, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}}
# current regular season stats
df1 = da[da['Source']=="list-regular"]
df2 = df1.groupby(["Name"]).apply(lambda x: x.sort_values(["Year"], ascending = False)).reset_index(drop=True)
dc = df2.groupby('Name').head(1)
# + id="p5pMZTDO8aC7" executionInfo={"status": "ok", "timestamp": 1614030742672, "user_tz": -660, "elapsed": 819, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}}
dt = dc[['Season','Age','League','Team','Name']]
# + id="CnBYw1Lqn-t8" executionInfo={"status": "ok", "timestamp": 1614030750707, "user_tz": -660, "elapsed": 747, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}}
# concatenate current team with player
dh = da[['Year','Season','Name','Eff']]
dh = dh.merge(dt,on='Name')
dp = df_players[['Name','Position','Height']]
dh = dh.merge(dp,on='Name')
dh['Player-Team'] = dh['Name'] + '\n(' + dh['Team'] + ') ' + '\n Pos: ' + dh['Position'] + ' '+ ' Height: ' + dh['Height'].astype('str') + ' '
# + id="w284tB0dJpej" executionInfo={"status": "ok", "timestamp": 1614030756951, "user_tz": -660, "elapsed": 740, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}}
# calculate mean for given year
dh = dh.groupby(['Name','Year','Player-Team','Position'],as_index=False).mean()
# + colab={"base_uri": "https://localhost:8080/"} id="aDx0zHkA99G-" executionInfo={"status": "ok", "timestamp": 1614030760044, "user_tz": -660, "elapsed": 737, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}} outputId="4146540c-b5c8-4bef-eae5-dc9eb8461a9b"
dh.Position.unique()
# + id="KDnNR3XRoG9-" executionInfo={"status": "ok", "timestamp": 1614030763462, "user_tz": -660, "elapsed": 749, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}}
# order player positions
t = pd.CategoricalDtype(categories=['PG', 'SG', 'SF', 'PF', 'C'], ordered=True)
dh['sort']=pd.Series(dh.Position, dtype=t)
dh.sort_values(by=['sort','Name','Height'],ascending=[True,True,True],ignore_index=True,inplace=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} id="E6eU_H6mEzNK" executionInfo={"status": "ok", "timestamp": 1614030766571, "user_tz": -660, "elapsed": 761, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}} outputId="eb9d0d85-d888-4b6a-f149-293e5b04b99e"
dh.head()
# -
# ## Olympic stats
dt2020_stats = pd.read_excel('../Data/tokyo2020_proballer_stats.xlsx')
# +
dt2020_stats = compute_epg(dt2020_stats)
dt2020_stats['Diff'] = dt2020_stats['EFF.Calc'] - dt2020_stats['Eff']
dt2020_stats['Ratio'] = dt2020_stats['EFF.Calc'] / dt2020_stats['Eff']
dt2020_stats
# -
# sort facets by position and height
# sort data by year
dh = dh.sort_values(by=['sort','Height','Year'])
# +
# get the list of ordered names from the dataframe (collapsing duplicates)
ordered_names = pd.Series(dh['Name']).unique()
# reformat as dataframe (so we can do a join later)
dy = pd.DataFrame(ordered_names,columns=['Player'])
# -
# subset of Olympic stats
dz = dt2020_stats[['Player','EFF.Calc']]
dz.head()
# sort efficiency values using our sorted names
dy = dy.join(dz.set_index('Player'),on='Player')[['Player','EFF.Calc']]
dy.head()
# + id="bFcHDs5AoNhX" colab={"base_uri": "https://localhost:8080/", "height": 750} executionInfo={"status": "ok", "timestamp": 1614030785310, "user_tz": -660, "elapsed": 12082, "user": {"displayName": "<NAME>", "photoUrl": "", "userId": "07937251379700638090"}} outputId="7c87013b-364c-4128-ae36-c53088980218"
# Create a grid : initialize it
g = sns.FacetGrid(dh, col='Player-Team', hue='Position', col_wrap=4)
# Add the line over the area with the plot function
g = g.map(plt.plot, 'Year', 'Eff')
# Fill the area with fill_between
g = g.map(plt.fill_between, 'Year', 'Eff', alpha=0.2).set_titles("{col_name} Player-Team")
# Control the title of each facet
g = g.set_titles("{col_name}")
axes = g.axes.flatten()
for i, ax in enumerate(axes):
ax.axhline(dy.iloc[i]['EFF.Calc'], ls='--', c='purple')
# Add a title for the whole plot
plt.subplots_adjust(top=0.85)
sns.set(rc={'figure.figsize':(12,8)})
g = g.fig.suptitle('Australian Men\'s Olympic Basketball Performance\nHistorical Effciency vs Final Tokyo 2020 Olympic Value')
plt.text(2020,-18,"linkedin.com/in/evassiliadis",ha='right',alpha=0.5)
# plt.text(2020,-16,"github.com/vass1138/wnbl",ha='right',alpha=0.5)
plt.savefig('boomers_tokyo2020_eff_history.png')
plt.show()
|
Scripts/proballer_analyse_selected.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
import pandas as pd
import os
from pprint import pprint as pp
os.getcwd()
df = pd.read_csv(r"D:\Python\pycharmWorkspace\xinsheng\data\u.data",
sep='\t',
# nrows=100,
names=['user_id', 'item_id', 'rating', 'timestamp'])
d = dict()
for _, row in df.iterrows():
user_id = str(row['user_id'])
item_id = str(row['item_id'])
rating = str(row['rating'])
if user_id not in d:
d[user_id] = {item_id: rating}
else:
d[user_id][item_id] = rating
# d
(d['196'])
pp(d['186'])
pp(set(d['196']) & set(d['186']))
w = dict()
# +
for u in d:
for v in d:
if u not in w:
w[u] = dict()
if u == v:
continue
w[u][v] = len(set(d[u]) & set(d[v]))
w[u][v] = 2 * w[u][v] / (len(d[u]) + len(d[v]))
pp(w[10])
# -
|
xinsheng/test.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
from image_processing import *
# %matplotlib inline
# -
# ## 2.1
# +
#from skimage.viewer import ImageViewer as IV
#viewer = IV(wombats)
#viewer.show()
# -
imshow(wombats)
# ## 2.2
backyard.shape
backyard[99, 199, :]
# ## 2.3
emu.shape
emu[108, 60, :]
np.set_printoptions(precision=4)
emu[108, 60, :].astype(float) / 255
# ## 2.4
# !exiftool read/cassowary.png
a = 23
b = uint8(a)
# %whos int uint8
# ## 2.5
imsave("cameraman.png", cameraman)
|
Chapter/Chapter 2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <table>
# <tr>
# <td style="background-color:#ffffff;"><a href="https://qsoftware.lu.lv/index.php/qworld/" target="_blank"><img src="..\images\qworld.jpg" width="70%" align="left"></a></td>
# <td style="background-color:#ffffff;" width="*"></td>
# <td style="background-color:#ffffff;vertical-align:text-top;"><a href="https://qsoftware.lu.lv" target="_blank"><img src="..\images\logo.jpg" width="25%" align="right"></a></td>
# </tr>
# <tr><td colspan="3" align="right" style="color:#777777;background-color:#ffffff;font-size:12px;">
# prepared by <a href="http://abu.lu.lv" target="_blank"><NAME></a>
# </td></tr>
# <tr><td colspan="3" align="right" style="color:#bbbbbb;background-color:#ffffff;font-size:11px;font-style:italic;">
# This cell contains some macros. If there is a problem with displaying mathematical formulas, please run this cell to load these macros.
# </td></tr>
# </table>
# $ \newcommand{\bra}[1]{\langle #1|} $
# $ \newcommand{\ket}[1]{|#1\rangle} $
# $ \newcommand{\braket}[2]{\langle #1|#2\rangle} $
# $ \newcommand{\dot}[2]{ #1 \cdot #2} $
# $ \newcommand{\biginner}[2]{\left\langle #1,#2\right\rangle} $
# $ \newcommand{\mymatrix}[2]{\left( \begin{array}{#1} #2\end{array} \right)} $
# $ \newcommand{\myvector}[1]{\mymatrix{c}{#1}} $
# $ \newcommand{\myrvector}[1]{\mymatrix{r}{#1}} $
# $ \newcommand{\mypar}[1]{\left( #1 \right)} $
# $ \newcommand{\mybigpar}[1]{ \Big( #1 \Big)} $
# $ \newcommand{\sqrttwo}{\frac{1}{\sqrt{2}}} $
# $ \newcommand{\dsqrttwo}{\dfrac{1}{\sqrt{2}}} $
# $ \newcommand{\onehalf}{\frac{1}{2}} $
# $ \newcommand{\donehalf}{\dfrac{1}{2}} $
# $ \newcommand{\hadamard}{ \mymatrix{rr}{ \sqrttwo & \sqrttwo \\ \sqrttwo & -\sqrttwo }} $
# $ \newcommand{\vzero}{\myvector{1\\0}} $
# $ \newcommand{\vone}{\myvector{0\\1}} $
# $ \newcommand{\vhadamardzero}{\myvector{ \sqrttwo \\ \sqrttwo } } $
# $ \newcommand{\vhadamardone}{ \myrvector{ \sqrttwo \\ -\sqrttwo } } $
# $ \newcommand{\myarray}[2]{ \begin{array}{#1}#2\end{array}} $
# $ \newcommand{\X}{ \mymatrix{cc}{0 & 1 \\ 1 & 0} } $
# $ \newcommand{\Z}{ \mymatrix{rr}{1 & 0 \\ 0 & -1} } $
# $ \newcommand{\Htwo}{ \mymatrix{rrrr}{ \frac{1}{2} & \frac{1}{2} & \frac{1}{2} & \frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & \frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} \\ \frac{1}{2} & -\frac{1}{2} & -\frac{1}{2} & \frac{1}{2} } } $
# $ \newcommand{\CNOT}{ \mymatrix{cccc}{1 & 0 & 0 & 0 \\ 0 & 1 & 0 & 0 \\ 0 & 0 & 0 & 1 \\ 0 & 0 & 1 & 0} } $
# $ \newcommand{\norm}[1]{ \left\lVert #1 \right\rVert } $
# <h2>Multiple Rotations</h2>
# The trivial way of implementing more than one rotation in parallel is to use a separate qubit for each rotation.
#
# If we have $ t $ different rotations with angles $ \theta_1,\ldots,\theta_t $, then we can use $ t $ qubits.
#
# Alternatively, we can use $ \log_2 (t) + 1 $ qubits (assuming that $t$ is a power of 2) that implement the following unitary matrix:
# $$
# R(\theta_1,\ldots,\theta_t) =
# \mymatrix{rr|rr|cc|rr}{
# \cos \theta_1 & -\sin \theta_1 & 0 & 0 & \cdots & \cdots & 0 & 0 \\
# \sin \theta_1 & \cos \theta_1 & 0 & 0 & \cdots & \cdots & 0 & 0 \\ \hline
# 0 & 0 & \cos \theta_2 & -\sin \theta_2 & \cdots & \cdots & 0 & 0 \\
# 0 & 0 & \sin \theta_2 & \cos \theta_2 & \cdots & \cdots & 0 & 0 \\ \hline
# \vdots & \vdots & \vdots & \vdots & \ddots & & \vdots & \vdots \\
# \vdots & \vdots & \vdots & \vdots & & \ddots & \vdots & \vdots \\ \hline
# 0 & 0 & 0 & 0 & \cdots & \cdots & \cos \theta_t & -\sin \theta_t \\
# 0 & 0 & 0 & 0 & \cdots & \cdots & \sin \theta_t & \cos \theta_t \\
# } .
# $$
# We can use this idea to solve the problem $\sf MOD_p$ (see <a href="B72_Rotation_Automata.ipynb" target="_blank">Rotation Automata</a>).
#
# We implement $ t $ rotation automata in this way.
#
# At the beginning of the computation, we apply Hadamard operator in each qubit. Then, we apply the operator $ R(\theta_1,\ldots,\theta_t) $ for each symbol from the stream. Once the stream is finished, we apply Hadamard operator in each qubit again.
#
# If we observe only state 0 in each qubit, then we consider the stream having the length of a multiple of $\sf p$. Otherwise, we consider the stream having the length of not a multiple of $\sf p$.
# <h3> Constructing $ R(\theta_1,\theta_2) $ </h3>
# When $t=2$, $ \log_2 (2) + 1 = 2 $. So, both implementations use the same number of qubits.
#
# But, it is a good starting point to construct the following unitary operator:
# $$
# R(\theta_1,\theta_2) =
# \mymatrix{rrrr}{
# \cos \theta_1 & -\sin \theta_1 & 0 & 0 \\
# \sin \theta_1 & \cos \theta_1 & 0 & 0 \\
# 0 & 0 & \cos \theta_2 & -\sin \theta_2 \\
# 0 & 0 & \sin \theta_2 & \cos \theta_2 \\
# } .
# $$
# <div style="background-color:#f8f8f8;">
# <b> Technical Remark:</b>
#
# When two qubits are combined (tensored) in qiskit, say $ qreg[0] $ and $ qreg[1] $, they are ordered as $ qreg[1] \otimes qreg[0] $.
#
# If there are $n$ qubits, say $ qreg[0],\ldots,qreg[n-1] $ to be combined, they are ordered in qiskit as
#
# $$ qreg[n-1] \otimes \cdots \otimes qreg[0] . $$
# </div>
# We use a controlled rotation gate $ cu3 $ in qiskit.
#
# <b> Gate $u3$: </b>
#
# The gate $ u3 $ is a generic one-qubit gate for rotation on Bloch sphere. It takes three parameters, and if we pass zeros as the second and third parameters, we implement our rotation gate $ ry $:
#
# u3(2*theta,0,0,qubit)
# is equivalent to
#
# ry(2*theta,qubit)
# Both make a rotation with angle $\theta$ in the real-valued qubit in counter-clockwise direction.
# <b> Gate $cu3$: </b>
#
# The two-qubit gate $ cu3 $ takes five parameters. We use it as follows:
#
# cu3(2*theta,0,0,control_qubit,target_qubit)
# If the control qubit is in state $ \ket{1} $, then the rotation
#
# ry(2*theta,target_qubit)
# is applied (to the target qubit).
# The base states of two qubits are ordered as $ \myarray{c}{00 \\ 01 \\ 10 \\ 11 } $
# or equivalently grouped as $ \myarray{c}{ 0 \otimes \myvector{0 \\ 1} \\ \hline 1 \otimes \myvector{0 \\ 1} } $.
#
# We can apply a rotation to the first qubit controlled by the second qubit.
#
# To construct $ R(\theta_1,\theta_2) $:
# <ol>
# <li> When the second qubit is in state $ \ket{0} $, we can apply the rotation with angle $ \theta_1 $. </li>
# <li> When the second qubit is in state $ \ket{1} $, we can apply the rotation with angle $ \theta_2 $. </li>
# </ol>
#
# Now, we implement this by also printing the constructed unitary matrix.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
from math import pi
# the angles of rotations
theta1 = pi/4
theta2 = pi/6
# the circuit with two qubits
qreg = QuantumRegister(2)
creg = ClassicalRegister(2)
mycircuit = QuantumCircuit(qreg,creg)
# when the second qubit is in |0>, the first qubit is rotated by theta1
mycircuit.x(qreg[1])
mycircuit.cu3(2*theta1,0,0,qreg[1],qreg[0])
mycircuit.x(qreg[1])
# when the second qubit is in |1>, the first qubit is rotated by theta2
mycircuit.cu3(2*theta2,0,0,qreg[1],qreg[0])
# we read the unitary matrix
job = execute(mycircuit,Aer.get_backend('unitary_simulator'),optimization_level=0)
u=job.result().get_unitary(mycircuit,decimals=3)
# we print the unitary matrix in nice format
for i in range(len(u)):
s=""
for j in range(len(u)):
val = str(u[i][j].real)
while(len(val)<8): val = " "+val
s = s + val
print(s)
# -
# <h3>Task 1</h3>
#
# Verify that the printed matrix is $ R(\pi/4,\pi/6) $.
#
# your code is here
#
# <a href="B76_Multiple_Rotations_Solutions.ipynb#task1">click for our solution</a>
# <h3> Constructing $ R(\theta_1,\theta_2,\theta_3,\theta_4) $ </h3>
# We can use $ \log_2(4) + 1 = 3 $ qubits to construct $ R(\theta_1,\theta_2,\theta_3,\theta_4) $.
# The base states of three qubits are ordered as $ \myarray{c}{000 \\ 001 \\ 010 \\ 011 \\ 100 \\ 101 \\ 110 \\ 111 } $
# or equivalently grouped as $
# \myarray{c}{
# 00 \otimes \myvector{0 \\ 1} \\ \hline
# 01 \otimes \myvector{0 \\ 1} \\ \hline
# 10 \otimes \myvector{0 \\ 1} \\ \hline
# 11 \otimes \myvector{0 \\ 1}
# } $.
# By using a rotation gate controlled by two qubits, we can easily implement our unitary operator.
#
# But, if we have a rotation gate controlled by only one qubit, then we use additional tricks (and qubits) and controlled CNOT gate by two qubits (also called Toffoli gate):
#
# circuit.ccx(control-qubit1,control-qubit2,target-qubit)
# <div style="background-color:#f9f9f9;">
# In general, if $ t = 2^n $, then we can construct $ R(\theta_1,\ldots,\theta_t) $ by using no more than $ 2\log_2(t) $ qubits (instead of $t$ qubits).
# </div>
# <h3> Pseudo construction </h3>
#
# We start with a construction using three angles.
# <h3> Task 2</h3>
#
# Consider a quantum circuit with 3 qubits.
#
# When the third qubit is in state $ \ket{1} $, apply the gate
#
# cu3(2*theta1,0,0,qreg[2],qreg[0])
#
# When the second qubit is in state $ \ket{1} $, apply the gate
#
# cu3(2*theta2,0,0,qreg[1],qreg[0])
#
# When the third qubit is in state $ \ket{0} $, apply the gate
#
# cu3(2*theta3,0,0,qreg[2],qreg[0])
#
# Guess the corresponding unitary matrix, which should be of the form:
#
# $$
# \mymatrix{rr|rr|rr|rr}{
# \cos a_1 & -\sin a_1 & 0 & 0 & 0 & 0 & 0 & 0 \\
# \sin a_1 & \cos a_1 & 0 & 0 & 0 & 0 & 0 & 0 \\ \hline
# 0 & 0 & \cos a_2 & -\sin a_2 & 0 & 0 & 0 & 0 \\
# 0 & 0 & \sin a_2 & \cos a_2 & 0 & 0 & 0 & 0 \\ \hline
# 0 & 0 & 0 & 0 & \cos a_3 & -\sin a_3 & 0 & 0 \\
# 0 & 0 & 0 & 0 & \sin a_3 & \cos a_3 & 0 & 0 \\ \hline
# 0 & 0 & 0 & 0 & 0 & 0 & \cos a_4 & -\sin a_4 \\
# 0 & 0 & 0 & 0 & 0 & 0 & \sin a_4 & \cos a_4 \\
# }
# $$
#
# In other words, find $a_1$, $ a_2 $, $a_3$, and $a_4$ in terms of $ \theta_1 $, $\theta_2$, and $ \theta_3 $.
# <a href="B76_Multiple_Rotations_Solutions.ipynb#task2">click for our solution</a>
# <h3>Task 3</h3>
#
# Implement Task 2 by picking three angles, and verify the constructed matrix.
#
# your solution is here
#
# <a href="B76_Multiple_Rotations_Solutions.ipynb#task3">click for our solution</a>
# <h3>Task 4</h3>
#
# Create a circuit for solving problem $ \sf MOD_{31} $ by using the implementation in Task 3.
#
# Pick $ \theta_1 $, $ \theta_2 $, and $ \theta_3 $ randomly.
#
# At the beginning of the stream and after reading the stream, apply Hadamard operators to each qubit.
#
# Execute your quantum program on the streams of lengths from 1 to 31.
#
# your solution is here
#
# <a href="B76_Multiple_Rotations_Solutions.ipynb#task4">click for our solution</a>
# <h3>Task 5 (optional)</h3>
#
# Based on Task 4, design your own solution for problem $ \sf MOD_{91} $ by using four qubits.
#
# Remark that up to 8 different rotations can be implemented by using four qubits.
#
# your solution is here
#
# <h3> Main construction </h3>
# To implement an operator controlled by two qubits, we use an auxiliary qubit.
#
# Depending on the desired values of two qubits, the auxiliary qubit is flipped to $ \ket{1} $ and then the operation is implemented controlled by the auxiliary qubit.
#
# Here we describe the case when the control qubits are in state $ \ket{01} $.
#
# We also draw the circuit.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
from math import pi
# initialize the circuit
qreg = QuantumRegister(4)
circuit = QuantumCircuit(qreg)
# we use the fourth qubit as the auxiliary
# apply a rotation to the first qubit when the third and second qubits are in states |0> and |1>
# change the state of the third qubit to |1>
circuit.x(qreg[2])
# if both the third and second qubits are in states |1>, the state of auxiliary qubit is changed to |1>
circuit.ccx(qreg[2],qreg[1],qreg[3])
# the rotation is applied to the first qubit if the state of auxiliary qubit is |1>
circuit.cu3(2*pi/6,0,0,qreg[3],qreg[0])
# reverse the effects
circuit.ccx(qreg[2],qreg[1],qreg[3])
circuit.x(qreg[2])
circuit.draw()
# -
# Based on this idea, different rotation operators are applied to the first qubit when the third and second qubits are in $ \ket{00} $, $ \ket{01} $, $ \ket{10} $, and $ \ket{11} $.
#
# We present how to construct $ R(\pi/10,2\pi/10,3\pi/10,4\pi/10) $.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
from math import pi,sin
# the angles of rotations
theta1 = pi/10
theta2 = 2*pi/10
theta3 = 3*pi/10
theta4 = 4*pi/10
# for verification, print sin(theta)'s
print("sin(theta1) = ",round(sin(theta1),3))
print("sin(theta2) = ",round(sin(theta2),3))
print("sin(theta3) = ",round(sin(theta3),3))
print("sin(theta4) = ",round(sin(theta4),3))
print()
qreg = QuantumRegister(4)
circuit = QuantumCircuit(qreg)
# the third qubit is in |0>
# the second qubit is in |0>
circuit.x(qreg[2])
circuit.x(qreg[1])
circuit.ccx(qreg[2],qreg[1],qreg[3])
circuit.cu3(2*theta1,0,0,qreg[3],qreg[0])
# reverse the effects
circuit.ccx(qreg[2],qreg[1],qreg[3])
circuit.x(qreg[1])
circuit.x(qreg[2])
# the third qubit is in |0>
# the second qubit is in |1>
circuit.x(qreg[2])
circuit.ccx(qreg[2],qreg[1],qreg[3])
circuit.cu3(2*theta2,0,0,qreg[3],qreg[0])
# reverse the effects
circuit.ccx(qreg[2],qreg[1],qreg[3])
circuit.x(qreg[2])
# the third qubit is in |1>
# the second qubit is in |0>
circuit.x(qreg[1])
circuit.ccx(qreg[2],qreg[1],qreg[3])
circuit.cu3(2*theta3,0,0,qreg[3],qreg[0])
# reverse the effects
circuit.ccx(qreg[2],qreg[1],qreg[3])
circuit.x(qreg[1])
# the third qubit is in |1>
# the second qubit is in |1>
circuit.ccx(qreg[2],qreg[1],qreg[3])
circuit.cu3(2*theta4,0,0,qreg[3],qreg[0])
# reverse the effects
circuit.ccx(qreg[2],qreg[1],qreg[3])
# read the corresponding unitary matrix
job = execute(circuit,Aer.get_backend('unitary_simulator'),optimization_level=0)
unitary_matrix=job.result().get_unitary(circuit,decimals=3)
for i in range(len(unitary_matrix)):
s=""
for j in range(len(unitary_matrix)):
val = str(unitary_matrix[i][j].real)
while(len(val)<7): val = " "+val
s = s + val
print(s)
# -
# <b>Remarks:</b>
#
# The constructed matrix is bigger than our main matrix because of the auxiliary qubit.
#
# Our main matrix appears at the top-left quarter of the constructed matrix.
#
# The rest of the constructed matrix does not affect our computation unless the auxiliary qubit is set to state $ \ket{1} $ (except the auxiliary operations).
# <h3>Task 6 (optional)</h3>
#
# Assume that $\theta_1=\frac{\pi}{11}$, $\theta_2=2\frac{\pi}{11}$, $\theta_3=4\frac{\pi}{11}$, and $\theta_4=8\frac{\pi}{11}$ are the given angles in the above construction.
#
# Calculate (by hand or in your mind) the angles of the rotations in the bottom-left quarter of the constructed matrix by following the construction steps.
# <h3>Task 7</h3>
#
# Create a circuit for solving problem $ \sf MOD_{61} $ by using the above implementation.
#
# Pick $ \theta_1 $, $ \theta_2 $, $ \theta_3 $, and $ \theta_4 $ randomly.
#
# At the beginning of the stream and after reading the stream, apply Hadamard operators to each qubit.
#
# Execute your quantum program on the streams of lengths 1, 11, 21, 31, 41, 51, and 61.
#
# your solution is here
#
# <a href="B76_Multiple_Rotations_Solutions.ipynb#task7">click for our solution</a>
# <h3>Task 8</h3>
#
# How many qubits we use to implement the main construction having 16 rotations in parallel?
#
# Please specify the number of control qubits and auxiliary qubits.
# <a href="B76_Multiple_Rotations_Solutions.ipynb#task8">click for our solution</a>
# <h3>Bonus (saving some qubits)</h3>
#
# We can use additional trick to save some qubits in our implementation. The idea relies on the following fact: if you apply a rotation between two NOT gates, the rotation will happen in the opposite direction.
#
# We can use this idea to implement a rotation by $\theta$ in the following way:
# <ul>
# <li>Rotate in the qubit by $\frac{\theta}{2}$;</li>
# <li>Apply NOT to the qubit;</li>
# <li>Rotate in the qubit by $-\frac{\theta}{2}$;</li>
# <li>Apply NOT to the qubit.</li>
# </ul>
#
# As a result we will rotate in the qubit by $\theta$. We can control NOT and rotation operations and perform a rotation only when all control qubits are in state 1, relying on the following simple facts:
# <ul>
# <li>Two NOT gates result into identity operation;</li>
# <li>Rotations by $\frac{\theta}{2}$ and $-\frac{\theta}{2}$ result into identity operation.</li>
# </ul>
#
# Below you can see the code that shows how can we use the discussed ideas to control rotations on one qubit by three qubits. If the state of at least one of the control qubits is 0, then the identity will be applied to the controlled qubit.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
from math import pi
qreg11 = QuantumRegister(4)
creg11 = ClassicalRegister(4)
theta = pi/4
# define our quantum circuit
mycircuit11 = QuantumCircuit(qreg11,creg11)
def ccc_ry(angle,q1,q2,q3,q4):
mycircuit11.cu3(angle/2,0,0,q3,q4)
mycircuit11.ccx(q1,q2,q4)
mycircuit11.cu3(-angle/2,0,0,q3,q4)
mycircuit11.ccx(q1,q2,q4)
ccc_ry(2*theta,qreg11[3],qreg11[2],qreg11[1],qreg11[0])
mycircuit11.draw(output='mpl')
# -
# The code below demonstrates the implementation of 8 rotations with total 4 qubits, one of which is controlled by others.
# +
from qiskit import QuantumRegister, ClassicalRegister, QuantumCircuit, execute, Aer
from math import pi
qreg12 = QuantumRegister(4)
creg12 = ClassicalRegister(4)
theta1 = pi/16
theta2 = 2*pi/16
theta3 = 3*pi/16
theta4 = 4*pi/16
theta5 = 5*pi/16
theta6 = 6*pi/16
theta7 = 7*pi/16
theta8 = 8*pi/16
# define our quantum circuit
mycircuit12 = QuantumCircuit(qreg12,creg12)
def ccc_ry(angle,q1,q2,q3,q4):
mycircuit12.cu3(angle/2,0,0,q3,q4)
mycircuit12.ccx(q1,q2,q4)
mycircuit12.cu3(-angle/2,0,0,q3,q4)
mycircuit12.ccx(q1,q2,q4)
mycircuit12.x(qreg12[3])
mycircuit12.x(qreg12[2])
mycircuit12.x(qreg12[1])
ccc_ry(2*theta1,qreg12[3],qreg12[2],qreg12[1],qreg12[0])
mycircuit12.x(qreg12[1])
mycircuit12.x(qreg12[2])
mycircuit12.x(qreg12[3])
mycircuit12.x(qreg12[3])
mycircuit12.x(qreg12[2])
#mycircuit12.x(qreg12[1])
ccc_ry(2*theta2,qreg12[3],qreg12[2],qreg12[1],qreg12[0])
#mycircuit12.x(qreg12[1])
mycircuit12.x(qreg12[2])
mycircuit12.x(qreg12[3])
mycircuit12.x(qreg12[3])
#mycircuit12.x(qreg12[2])
mycircuit12.x(qreg12[1])
ccc_ry(2*theta3,qreg12[3],qreg12[2],qreg12[1],qreg12[0])
mycircuit12.x(qreg12[1])
#mycircuit12.x(qreg12[2])
mycircuit12.x(qreg12[3])
mycircuit12.x(qreg12[3])
#mycircuit12.x(qreg12[2])
#mycircuit12.x(qreg12[1])
ccc_ry(2*theta4,qreg12[3],qreg12[2],qreg12[1],qreg12[0])
#mycircuit12.x(qreg12[1])
#mycircuit12.x(qreg12[2])
mycircuit12.x(qreg12[3])
#mycircuit12.x(qreg12[3])
mycircuit12.x(qreg12[2])
mycircuit12.x(qreg12[1])
ccc_ry(2*theta5,qreg12[3],qreg12[2],qreg12[1],qreg12[0])
mycircuit12.x(qreg12[1])
mycircuit12.x(qreg12[2])
#mycircuit12.x(qreg12[3])
#mycircuit12.x(qreg12[3])
mycircuit12.x(qreg12[2])
#mycircuit12.x(qreg12[1])
ccc_ry(2*theta6,qreg12[3],qreg12[2],qreg12[1],qreg12[0])
#mycircuit12.x(qreg12[1])
mycircuit12.x(qreg12[2])
#mycircuit12.x(qreg12[3])
#mycircuit12.x(qreg12[3])
#mycircuit12.x(qreg12[2])
mycircuit12.x(qreg12[1])
ccc_ry(2*theta7,qreg12[3],qreg12[2],qreg12[1],qreg12[0])
mycircuit12.x(qreg12[1])
#mycircuit12.x(qreg12[2])
#mycircuit12.x(qreg12[3])
#mycircuit12.x(qreg12[3])
#mycircuit12.x(qreg12[2])
#mycircuit12.x(qreg12[1])
ccc_ry(2*theta8,qreg12[3],qreg12[2],qreg12[1],qreg12[0])
#mycircuit12.x(qreg12[1])
#mycircuit12.x(qreg12[2])
#mycircuit12.x(qreg12[3])
job = execute(mycircuit12,Aer.get_backend('unitary_simulator'),optimization_level=0)
u=job.result().get_unitary(mycircuit12,decimals=3)
for i in range(len(u)):
s=""
for j in range(len(u)):
val = str(u[i][j].real)
while(len(val)<7): val = " "+val
s = s + val
print(s)
# -
# <h3>Task 9</h3>
#
# By using the discussed ideas, how many qubits can we have to implement 16 rotations in parallel?
#
# Please specify the number of control qubits and auxiliary qubits.
# <a href="B76_Multiple_Rotations_Solutions.ipynb#task9">click for our solution</a>
|
bronze/B76_Multiple_Rotations.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Titanic Dataset
#
# ## Imports
#
# Start with importing the necessary libraries.
# +
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import matplotlib.gridspec as gridspec
from scipy.stats import gaussian_kde
# -
# ## Data
#
# Load in the data. <br>
# Source: https://www.kaggle.com/c/titanic
df = pd.read_csv('data/train.csv')
print('Number of Rows:', len(df))
print('Columns:', list(df.columns))
# ## Data Exploration
#
# Get a sense of the data. Look for patterns to measure statistically.
df.head(10)
# ### General Trends
#
# Simple relationships between survival and other variables.
base_survival = ((df.Survived.value_counts() * 100)/ len(df)).round(1).astype(str) + '%'
base_survival.index = ['Died', 'Survived']
base_survival
pt = df.pivot_table(index='Survived')
pt.index = pd.Series(['No', 'Yes'], name='Survived?')
pt
# A few things of notice:
# <ul>
# <li>Here we see that survivors tended to be younger than non-survivors.</li>
# <li>They also tended to have paid a higher fare and stayed in a higher class (indicated by a lower number) than non-survivors. </li>
# <li>In general, having a greater number of parents or children on board is correlated with a greater survival rate, but having <i>fewer</i> siblings/spouses was related to a higher rate of survival.</li>
# </ul>
# ### Sex
#
# Look at differences in survival rates between the sexes:
print('Passenger sexual demographics:')
df.Sex.value_counts()
print('Survival rates by sex:')
df.groupby('Sex').agg(np.mean)['Survived']
df.pivot_table(index='Sex',
columns='Survived',
aggfunc=np.mean)
# So far we can see that female passengers were much more likely to survive the ship's sinking than men. We can also see that female survivors tended to be older than female passengers who did not survive, but that the opposite relationship holds for men.
# ### Age
print('Min Age:', min(df.Age))
print('Max Age:', max(df.Age))
# +
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
died = df[df.Survived == 0]['Age'].dropna()
survived = df[df.Survived == 1]['Age'].dropna()
died_density = gaussian_kde(died)
xs = np.linspace(0,80,200)
died_density.covariance_factor = lambda : .25
died_density._compute_covariance()
plt.plot(xs,died_density(xs), color='#cc2000', label='Died')
survived_density = gaussian_kde(survived)
xs = np.linspace(0,80,200)
survived_density.covariance_factor = lambda : .25
survived_density._compute_covariance()
plt.plot(xs,survived_density(xs), color='#228b22', label='Survived')
plt.yticks([])
plt.tick_params(bottom='off')
for spine in ax.spines.values():
spine.set_visible(False)
plt.legend()
plt.xlabel('Age in Years')
plt.title('Density of Age by Survival Status', fontsize=15)
plt.show()
# -
# Looking at the plot above, we can see that the age trends for people in both survival groups are quite similar. We can see that survivors had a somewhat wider range of ages, while those who died were more concentrated at around 20-30 years old. Also there is a large concentration of very young children (less than 5-7 years old) that survived.
# ### Socio-Economics
print('Min Fare:', min(df.Fare))
print('Max Fare:', max(df.Fare))
# +
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
died = df[df.Survived == 0]['Fare'].dropna()
survived = df[df.Survived == 1]['Fare'].dropna()
died_density = gaussian_kde(died)
xs = np.linspace(0,520,200)
died_density.covariance_factor = lambda : .25
died_density._compute_covariance()
plt.plot(xs,died_density(xs), color='#cc2000', label='Died')
survived_density = gaussian_kde(survived)
xs = np.linspace(0,520,200)
survived_density.covariance_factor = lambda : .25
survived_density._compute_covariance()
plt.plot(xs,survived_density(xs), color='#228b22', label='Survived')
plt.yticks([])
plt.tick_params(bottom='off')
for spine in ax.spines.values():
spine.set_visible(False)
plt.legend()
plt.xlabel('Fare Price')
plt.title('Density of Fare Price by Survival Status', fontsize=15)
plt.show()
# -
# From the above plot we can see that most of the passengers who died paid quite a low fair price. While those who lived tended to have paid higher prices to board.
# Now we'll explore the relationship between cabin class and survival.
# +
fig = plt.figure(figsize=(12,10))
gspec = gridspec.GridSpec(2,2)
died = df[df.Survived == 0]['Pclass'].value_counts().sort_index()
survived = df[df.Survived == 1]['Pclass'].value_counts().sort_index()
class_labels = ['1st Class', '2nd Class', '3rd Class']
class_colors = ['#228b22', '#00688b', '#dd2000']
ax = plt.subplot(gspec[0,0])
_, _, autotexts = plt.pie(died, labels=class_labels, colors=class_colors, autopct='%1.1f%%')
for autotext in autotexts:
autotext.set_color('white')
autotext.set_size(12)
plt.title('Died', fontsize=15)
ax = plt.subplot(gspec[0,1])
_, _, autotexts = plt.pie(survived, labels=class_labels, colors=class_colors, autopct='%1.1f%%')
for autotext in autotexts:
autotext.set_color('white')
autotext.set_size(12)
plt.title('Survived', fontsize=15)
ax = plt.subplot(gspec[1,:])
class_survival_rates = list(map(lambda class_num: len(df[(df.Pclass == class_num) & (df.Survived == 1)])/len(df[df.Pclass == class_num]),
sorted(df.Pclass.unique())))
#survivors
x_pos = np.arange(len(df.Pclass.unique()))
plt.bar(x_pos, class_survival_rates, width=0.5, color=class_colors)
for pat in ax.patches:
ax.annotate(str(round(pat.get_height(), 3) * 100) + '%', (pat.get_x() + pat.get_width()/2, pat.get_height() - 0.05), horizontalalignment='center', color='white')
plt.yticks([])
plt.xticks(x_pos, class_labels)
plt.xlim(-2,4)
plt.tick_params(bottom='off')
for spine in ax.spines.values():
spine.set_visible(False)
plt.title('Survival Rates by Class', fontsize=15)
plt.show()
# -
((df.Pclass.value_counts()*100 / len(df)).round(1).astype(str) + '%').sort_index()
# ## Constructing a Model
# +
import sklearn
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.preprocessing import LabelEncoder
from sklearn.model_selection import train_test_split
from sklearn.metrics import roc_auc_score
# -
df.dtypes
#Train/Test Split and creating Dummy Variables
feature_cols = ['Pclass', 'Sex', 'Age', 'SibSp', 'Parch', 'Fare', 'Embarked']
df = df[feature_cols + ['Survived']].dropna()
df['Sex'] = LabelEncoder().fit_transform(df.Sex)
df['Embarked'] = LabelEncoder().fit_transform(df.Embarked)
X_train, X_test, y_train, y_test = train_test_split(df[feature_cols], df.Survived, random_state=0)
# ### Baseline
# +
tree = DecisionTreeClassifier(random_state=0)
tree.fit(X_train, y_train)
print('Accuracy on training set:',tree.score(X_train, y_train))
print('Accuracy on test set:',tree.score(X_test, y_test))
print('Area under the ROC curve score on test set:',roc_auc_score(y_test, tree.predict_proba(X_test)[:, 1]))
# -
# ### More Advanced
# +
forest = GradientBoostingClassifier(learning_rate=0.1, max_depth=4, random_state=0)
forest.fit(X_train, y_train)
print('Accuracy on training set:',forest.score(X_train, y_train))
print('Accuracy on test set:',forest.score(X_test, y_test))
print('Area under the ROC curve score on test set:',roc_auc_score(y_test, forest.predict_proba(X_test)[:, 1]))
# +
feature_importances = dict(zip(feature_cols, forest.feature_importances_))
sorted_importances = sorted(list(feature_importances.items()), key=lambda x: x[1], reverse=True)
print("Feature\t\tImportance")
print("-------\t\t----------")
for feature, importance in sorted_importances:
padding = "\t\t" if len(feature) <= 6 else "\t"
print(feature, padding, importance)
# -
def add_percent_labels(ax):
for pat in ax.patches:
if pat.get_height() > 0.05:
height_offset = -0.02
color = "white"
else:
height_offset = 0.01
color = "#1f77b4"
rounded_importance_label = str((int(round(pat.get_height(), 3) * 1000)) / 10) + '%'
ax.annotate(
rounded_importance_label, (pat.get_x() + pat.get_width()/2, pat.get_height() + height_offset),
horizontalalignment='center',
color=color
)
# +
fig = plt.figure(figsize=(8,6))
ax = fig.add_subplot(111)
features = [x[0] for x in sorted_importances]
importances = [x[1] for x in sorted_importances]
x_pos = np.arange(len(sorted_importances))
plt.bar(x_pos, importances)
plt.yticks([])
plt.xticks(x_pos, features)
plt.tick_params(bottom='off')
for spine in ax.spines.values():
spine.set_visible(False)
add_percent_labels(ax)
plt.title('Feature Importances')
plt.show()
# -
# ### A Little Further
# +
male = df[df.Sex == 1]
female = df[df.Sex == 0]
feature_cols.remove('Sex')
X_train_male, X_test_male, y_train_male, y_test_male = train_test_split(male[feature_cols], male.Survived, random_state=0)
X_train_female, X_test_female, y_train_female, y_test_female = train_test_split(female[feature_cols], female.Survived, random_state=0)
male_forest = GradientBoostingClassifier(learning_rate=0.1, max_depth=4, random_state=0)
male_forest.fit(X_train_male, y_train_male)
female_forest = GradientBoostingClassifier(learning_rate=0.1, max_depth=4, random_state=0)
female_forest.fit(X_train_female, y_train_female)
print('Male')
print('Accuracy on training set:',male_forest.score(X_train_male, y_train_male))
print('Accuracy on test set:',male_forest.score(X_test_male, y_test_male))
print('Area under the ROC curve score on test set:',roc_auc_score(y_test_male, male_forest.predict_proba(X_test_male)[:, 1]))
print('\nFemale')
print('Accuracy on training set:',female_forest.score(X_train_female, y_train_female))
print('Accuracy on test set:',female_forest.score(X_test_female, y_test_female))
print('Area under the ROC curve score on test set:',roc_auc_score(y_test_female, female_forest.predict_proba(X_test_female)[:, 1]))
# +
# sort using original sorting (above)
sorted_features = features
male_feature_importances = dict(zip(feature_cols, men_forest.feature_importances_))
sorted_male_importances = sorted(list(male_feature_importances.items()), key=lambda x: sorted_features.index(x[0]))
female_feature_importances = dict(zip(feature_cols, women_forest.feature_importances_))
sorted_female_importances = sorted(list(female_feature_importances.items()), key=lambda x: sorted_features.index(x[0]))
# +
fig = plt.figure(figsize=(12,8))
ax = fig.add_subplot(121)
male_features = [x[0] for x in sorted_male_importances]
male_importances = [x[1] for x in sorted_male_importances]
x_pos = np.arange(len(sorted_male_importances))
plt.bar(x_pos, male_importances)
plt.yticks([])
plt.xticks(x_pos, male_features)
plt.tick_params(bottom='off')
for spine in ax.spines.values():
spine.set_visible(False)
add_percent_labels(ax)
plt.title('Male Feature Importances')
ax = fig.add_subplot(122)
female_features = [x[0] for x in sorted_female_importances]
female_importances = [x[1] for x in sorted_female_importances]
x_pos = np.arange(len(sorted_female_importances))
plt.bar(x_pos, female_importances)
plt.yticks([])
plt.xticks(x_pos, female_features)
plt.tick_params(bottom='off')
for spine in ax.spines.values():
spine.set_visible(False)
add_percent_labels(ax)
plt.title('Female Feature Importances')
plt.show()
plt.show()
# -
# Of notice in the above charts:
# <ul>
# <li>Age becomes very important with Sex removed</li>
# <li>Age was more important than Fare for men but the reverse was true for women</li>
# <li>Pclass was much more important for women than men</li>
# <li>SibSp was much more important for men than for women</li>
# </ul>
|
titanic.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] colab_type="text" deletable=true editable=true id="kR-4eNdK6lYS"
# Deep Learning
# =============
#
# Assignment 3
# ------------
#
# Previously in `2_fullyconnected.ipynb`, you trained a logistic regression and a neural network model.
#
# The goal of this assignment is to explore regularization techniques.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" deletable=true editable=true id="JLpLa8Jt7Vu4"
# These are all the modules we'll be using later. Make sure you can import them
# before proceeding further.
from __future__ import print_function
import numpy as np
import tensorflow as tf
from six.moves import cPickle as pickle
# + [markdown] colab_type="text" deletable=true editable=true id="1HrCK6e17WzV"
# First reload the data we generated in `1_notmnist.ipynb`.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 11777, "status": "ok", "timestamp": 1449849322348, "user": {"color": "", "displayName": "", "isAnonymous": false, "isMe": true, "permissionId": "", "photoUrl": "", "sessionId": "0", "userId": ""}, "user_tz": 480} id="y3-cj1bpmuxc" outputId="e03576f1-ebbe-4838-c388-f1777bcc9873"
pickle_file = 'notMNIST.pickle'
with open(pickle_file, 'rb') as f:
save = pickle.load(f)
train_dataset = save['train_dataset']
train_labels = save['train_labels']
valid_dataset = save['valid_dataset']
valid_labels = save['valid_labels']
test_dataset = save['test_dataset']
test_labels = save['test_labels']
del save # hint to help gc free up memory
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
# + [markdown] colab_type="text" deletable=true editable=true id="L7aHrm6nGDMB"
# Reformat into a shape that's more adapted to the models we're going to train:
# - data as a flat matrix,
# - labels as float 1-hot encodings.
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}, "output_extras": [{"item_id": 1}]} colab_type="code" deletable=true editable=true executionInfo={"elapsed": 11728, "status": "ok", "timestamp": 1449849322356, "user": {"color": "", "displayName": "", "isAnonymous": false, "isMe": true, "permissionId": "", "photoUrl": "", "sessionId": "0", "userId": ""}, "user_tz": 480} id="IRSyYiIIGIzS" outputId="3f8996ee-3574-4f44-c953-5c8a04636582"
image_size = 28
num_labels = 10
def reformat(dataset, labels):
dataset = dataset.reshape((-1, image_size * image_size)).astype(np.float32)
# Map 1 to [0.0, 1.0, 0.0 ...], 2 to [0.0, 0.0, 1.0 ...]
labels = (np.arange(num_labels) == labels[:,None]).astype(np.float32)
return dataset, labels
train_dataset, train_labels = reformat(train_dataset, train_labels)
valid_dataset, valid_labels = reformat(valid_dataset, valid_labels)
test_dataset, test_labels = reformat(test_dataset, test_labels)
print('Training set', train_dataset.shape, train_labels.shape)
print('Validation set', valid_dataset.shape, valid_labels.shape)
print('Test set', test_dataset.shape, test_labels.shape)
# + cellView="both" colab={"autoexec": {"startup": false, "wait_interval": 0}} colab_type="code" deletable=true editable=true id="RajPLaL_ZW6w"
def accuracy(predictions, labels):
return (100.0 * np.sum(np.argmax(predictions, 1) == np.argmax(labels, 1))
/ predictions.shape[0])
# + [markdown] colab_type="text" deletable=true editable=true id="sgLbUAQ1CW-1"
# ---
# Problem 1
# ---------
#
# Introduce and tune L2 regularization for both logistic and neural network models. Remember that L2 amounts to adding a penalty on the norm of the weights to the loss. In TensorFlow, you can compute the L2 loss for a tensor `t` using `nn.l2_loss(t)`. The right amount of regularization should improve your validation / test accuracy.
#
# ---
# + [markdown] deletable=true editable=true
# ---
# Logistic Model
# ----
# + deletable=true editable=true
batch_size = 128
beta = 0.005
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(batch_size, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
weights = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases = tf.Variable(tf.zeros([num_labels]))
# Training computation.
logits = tf.matmul(tf_train_dataset, weights) + biases
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=tf_train_labels, logits=logits)
+ beta*tf.nn.l2_loss(weights)
)
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits)
valid_prediction = tf.nn.softmax(
tf.matmul(tf_valid_dataset, weights) + biases)
test_prediction = tf.nn.softmax(tf.matmul(tf_test_dataset, weights) + biases)
# + deletable=true editable=true
num_steps = 3001
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
# + [markdown] deletable=true editable=true
# * beta = 0.01 - Test accuracy: 88.5%
# * beta = 0.5 - Test accuracy: 55.5%
# * beta = 0.005 - Test accuracy: 88.7%
# * beta = 0.001 - Test accuracy: 88.7%
# * beta = 0.0001 - Test accuracy: 86.2%
#
# + [markdown] deletable=true editable=true
# ---
# Hidden Layer Model
# ---
# + deletable=true editable=true
batch_size = 128
beta=0.005
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(batch_size, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
weights_layer_1 = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases_layer_1 = tf.Variable(tf.zeros([num_labels]))
# Layer 2 weights have an input dimension = output of first layer
weights_layer_2 = tf.Variable(
tf.truncated_normal([num_labels, num_labels]))
biases_layer_2 = tf.Variable(tf.zeros([num_labels]))
# Training computation.
logits_layer_1 = tf.matmul(tf_train_dataset, weights_layer_1) + biases_layer_1
relu_output = tf.nn.relu(logits_layer_1)
logits_layer_2 = tf.matmul(relu_output, weights_layer_2) + biases_layer_2
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=tf_train_labels, logits=logits_layer_2)
+ beta*tf.nn.l2_loss(weights_layer_1)
+ beta*tf.nn.l2_loss(weights_layer_2)
)
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits_layer_2)
logits_l_1_valid = tf.matmul(tf_valid_dataset, weights_layer_1) + biases_layer_1
relu_valid = tf.nn.relu(logits_l_1_valid)
logits_l_2_valid = tf.matmul(relu_valid, weights_layer_2) + biases_layer_2
valid_prediction = tf.nn.softmax(logits_l_2_valid)
logits_l_1_test = tf.matmul(tf_test_dataset, weights_layer_1) + biases_layer_1
relu_test = tf.nn.relu(logits_l_1_test)
logits_l_2_test = tf.matmul(relu_test, weights_layer_2) + biases_layer_2
test_prediction = tf.nn.softmax(logits_l_2_test)
# + deletable=true editable=true
num_steps = 3001
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
# + [markdown] deletable=true editable=true
# beta = 0.01 - Test accuracy: 88.7%
# beta = 0.5 - Test accuracy: 45.9% (and slow)
# beta = 0.005 - Test accuracy: 89.2%
# beta = 0.001 - Test accuracy: 89.2%
# beta = 0.0001 - Test accuracy: 85.7%
# + [markdown] colab_type="text" deletable=true editable=true id="na8xX2yHZzNF"
# ---
# Problem 2
# ---------
# Let's demonstrate an extreme case of overfitting. Restrict your training data to just a few batches. What happens?
#
# ---
# + deletable=true editable=true
num_steps = 3001
#Restrict training data
reduced_train_dataset = train_dataset[:640, :]
reduced_train_labels = train_labels[:640, :]
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (reduced_train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = reduced_train_dataset[offset:(offset + batch_size), :]
batch_labels = reduced_train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
# + [markdown] deletable=true editable=true
# Restricted to 1000 samples in each batch - we get quick convergence and 100% accuracy on the mini-batch but poor performance on the validation dataset and poorer performance on the unseen test dataset.
# * Minibatch loss at step 3000: 0.268746
# * Minibatch accuracy: 100.0%
# * Validation accuracy: 78.4%
# * Test accuracy: 85.1%
# On 640 samples:
# * Minibatch loss at step 3000: 0.196770
# * Minibatch accuracy: 100.0%
# * Validation accuracy: 76.6%
# * Test accuracy: 83.6%
# + [markdown] colab_type="text" deletable=true editable=true id="ww3SCBUdlkRc"
# ---
# Problem 3
# ---------
# Introduce Dropout on the hidden layer of the neural network. Remember: Dropout should only be introduced during training, not evaluation, otherwise your evaluation results would be stochastic as well. TensorFlow provides `nn.dropout()` for that, but you have to make sure it's only inserted during training.
#
# What happens to our extreme overfitting case?
#
# ---
# + deletable=true editable=true
batch_size = 128
beta=0.005
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(batch_size, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
weights_layer_1 = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases_layer_1 = tf.Variable(tf.zeros([num_labels]))
# Layer 2 weights have an input dimension = output of first layer
weights_layer_2 = tf.Variable(
tf.truncated_normal([num_labels, num_labels]))
biases_layer_2 = tf.Variable(tf.zeros([num_labels]))
# Training computation.
logits_layer_1 = tf.matmul(tf_train_dataset, weights_layer_1) + biases_layer_1
relu_output = tf.nn.relu(logits_layer_1)
# Introduce dropout - probability feature is kept is passed as a variable
keep_probability = tf.placeholder(tf.float32)
dropout_output = tf.nn.dropout(relu_output, keep_probability)
logits_layer_2 = tf.matmul(dropout_output, weights_layer_2) + biases_layer_2
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=tf_train_labels, logits=logits_layer_2)
+ beta*tf.nn.l2_loss(weights_layer_1)
+ beta*tf.nn.l2_loss(weights_layer_2)
)
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits_layer_2)
logits_l_1_valid = tf.matmul(tf_valid_dataset, weights_layer_1) + biases_layer_1
relu_valid = tf.nn.relu(logits_l_1_valid)
logits_l_2_valid = tf.matmul(relu_valid, weights_layer_2) + biases_layer_2
valid_prediction = tf.nn.softmax(logits_l_2_valid)
logits_l_1_test = tf.matmul(tf_test_dataset, weights_layer_1) + biases_layer_1
relu_test = tf.nn.relu(logits_l_1_test)
logits_l_2_test = tf.matmul(relu_test, weights_layer_2) + biases_layer_2
test_prediction = tf.nn.softmax(logits_l_2_test)
# + deletable=true editable=true
num_steps = 3001
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels, keep_probability: 0.5}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
# + [markdown] deletable=true editable=true
# Dropout doesn't improve performance for me - maybe I'm applying it wrong - getting test accuracy of 80%.
#
# Try on reduced batch size data below:
# + deletable=true editable=true
num_steps = 3001
#Restrict training data
reduced_train_dataset = train_dataset[:640, :]
reduced_train_labels = train_labels[:640, :]
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (reduced_train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = reduced_train_dataset[offset:(offset + batch_size), :]
batch_labels = reduced_train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels, keep_probability: 0.5}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
# + [markdown] deletable=true editable=true
# Does reduce overfitting but does not increase accuracy.
# + [markdown] colab_type="text" deletable=true editable=true id="-b1hTz3VWZjw"
# ---
# Problem 4
# ---------
#
# Try to get the best performance you can using a multi-layer model! The best reported test accuracy using a deep network is [97.1%](http://yaroslavvb.blogspot.com/2011/09/notmnist-dataset.html?showComment=1391023266211#c8758720086795711595).
#
# One avenue you can explore is to add multiple layers.
#
# Another one is to use learning rate decay:
#
# global_step = tf.Variable(0) # count the number of steps taken.
# learning_rate = tf.train.exponential_decay(0.5, global_step, ...)
# optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss, global_step=global_step)
#
# ---
#
# + [markdown] deletable=true editable=true
# Try adding an additional layer:
# + deletable=true editable=true
batch_size = 128
beta=0.005
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(None, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(None, num_labels))
# Variables.
weights_layer_1 = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases_layer_1 = tf.Variable(tf.zeros([num_labels]))
# Layer 2 weights have an input dimension = output of first layer
weights_layer_2 = tf.Variable(
tf.truncated_normal([num_labels, num_labels]))
biases_layer_2 = tf.Variable(tf.zeros([num_labels]))
# Layer 3
weights_layer_3 = tf.Variable(
tf.truncated_normal([num_labels, num_labels]))
biases_layer_3 = tf.Variable(tf.zeros([num_labels]))
# Training computation.
# Compute layer 1
logits_layer_1 = tf.matmul(tf_train_dataset, weights_layer_1) + biases_layer_1
relu_output_1 = tf.nn.relu(logits_layer_1)
# Introduce dropout - probability feature is kept is passed as a variable
keep_probability = tf.placeholder(tf.float32)
dropout_output_1 = tf.nn.dropout(relu_output_1, keep_probability)
# Compute layer 2
logits_layer_2 = tf.matmul(dropout_output_1, weights_layer_2) + biases_layer_2
relu_output_2 = tf.nn.relu(logits_layer_2)
dropout_output_2 = tf.nn.dropout(relu_output_2, keep_probability)
# Computer layer 3
logits_layer_3 = tf.matmul(dropout_output_2, weights_layer_3) + biases_layer_3
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=tf_train_labels, logits=logits_layer_3)
+ beta*tf.nn.l2_loss(weights_layer_1)
+ beta*tf.nn.l2_loss(weights_layer_2)
+ beta*tf.nn.l2_loss(weights_layer_3)
)
# Optimizer.
optimizer = tf.train.GradientDescentOptimizer(0.5).minimize(loss)
# Predictions for the data.
train_prediction = tf.nn.softmax(logits_layer_3)
# Determine accuracy
correct_prediction = tf.equal(tf.argmax(train_prediction,1), tf.argmax(tf_train_labels,1))
accuracy_calc = tf.reduce_mean(tf.cast(correct_prediction, tf.float32))*100
# + deletable=true editable=true
num_steps = 3001
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels, keep_probability: 1.0}
_, l, accuracy = session.run(
[optimizer, loss, accuracy_calc], feed_dict=feed_dict
)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy)
valid_feed_dict = {tf_train_dataset : valid_dataset, tf_train_labels : valid_labels, keep_probability: 1.0}
print("Validation accuracy: %.1f%%" % accuracy_calc.eval(feed_dict=valid_feed_dict))
test_feed_dict = {tf_train_dataset : test_dataset, tf_train_labels : test_labels, keep_probability: 1.0}
print("Test accuracy: %.1f%%" % accuracy_calc.eval(feed_dict=test_feed_dict))
# + [markdown] deletable=true editable=true
# 87.2 with 3 layers and no dropout
# Dies at 10% accuracy with 0.5 dropout - is it basically destroying all the information?
# Yes even with 0.9 keep probability - only get 25%
#
# My code may be wrong somehow.
# + [markdown] deletable=true editable=true
# ---
# Learning rate decay
# ---
# + deletable=true editable=true
batch_size = 128
beta=0.005
graph = tf.Graph()
with graph.as_default():
# Input data. For the training data, we use a placeholder that will be fed
# at run time with a training minibatch.
tf_train_dataset = tf.placeholder(tf.float32,
shape=(batch_size, image_size * image_size))
tf_train_labels = tf.placeholder(tf.float32, shape=(batch_size, num_labels))
tf_valid_dataset = tf.constant(valid_dataset)
tf_test_dataset = tf.constant(test_dataset)
# Variables.
weights_layer_1 = tf.Variable(
tf.truncated_normal([image_size * image_size, num_labels]))
biases_layer_1 = tf.Variable(tf.zeros([num_labels]))
# Layer 2 weights have an input dimension = output of first layer
weights_layer_2 = tf.Variable(
tf.truncated_normal([num_labels, num_labels]))
biases_layer_2 = tf.Variable(tf.zeros([num_labels]))
# Training computation.
logits_layer_1 = tf.matmul(tf_train_dataset, weights_layer_1) + biases_layer_1
relu_output = tf.nn.relu(logits_layer_1)
logits_layer_2 = tf.matmul(relu_output, weights_layer_2) + biases_layer_2
loss = tf.reduce_mean(
tf.nn.softmax_cross_entropy_with_logits(labels=tf_train_labels, logits=logits_layer_2)
+ beta*tf.nn.l2_loss(weights_layer_1)
+ beta*tf.nn.l2_loss(weights_layer_2)
)
# Optimizer.
global_step = tf.Variable(0) # count the number of steps taken.
learning_rate = tf.train.exponential_decay(0.5, global_step, 100, 0.96)
optimizer = tf.train.GradientDescentOptimizer(learning_rate).minimize(loss, global_step=global_step)
# Predictions for the training, validation, and test data.
train_prediction = tf.nn.softmax(logits_layer_2)
logits_l_1_valid = tf.matmul(tf_valid_dataset, weights_layer_1) + biases_layer_1
relu_valid = tf.nn.relu(logits_l_1_valid)
logits_l_2_valid = tf.matmul(relu_valid, weights_layer_2) + biases_layer_2
valid_prediction = tf.nn.softmax(logits_l_2_valid)
logits_l_1_test = tf.matmul(tf_test_dataset, weights_layer_1) + biases_layer_1
relu_test = tf.nn.relu(logits_l_1_test)
logits_l_2_test = tf.matmul(relu_test, weights_layer_2) + biases_layer_2
test_prediction = tf.nn.softmax(logits_l_2_test)
# + deletable=true editable=true
num_steps = 3001
with tf.Session(graph=graph) as session:
tf.global_variables_initializer().run()
print("Initialized")
for step in range(num_steps):
# Pick an offset within the training data, which has been randomized.
# Note: we could use better randomization across epochs.
offset = (step * batch_size) % (train_labels.shape[0] - batch_size)
# Generate a minibatch.
batch_data = train_dataset[offset:(offset + batch_size), :]
batch_labels = train_labels[offset:(offset + batch_size), :]
# Prepare a dictionary telling the session where to feed the minibatch.
# The key of the dictionary is the placeholder node of the graph to be fed,
# and the value is the numpy array to feed to it.
feed_dict = {tf_train_dataset : batch_data, tf_train_labels : batch_labels}
_, l, predictions = session.run(
[optimizer, loss, train_prediction], feed_dict=feed_dict)
if (step % 500 == 0):
print("Minibatch loss at step %d: %f" % (step, l))
print("Minibatch accuracy: %.1f%%" % accuracy(predictions, batch_labels))
print("Validation accuracy: %.1f%%" % accuracy(
valid_prediction.eval(), valid_labels))
print("Test accuracy: %.1f%%" % accuracy(test_prediction.eval(), test_labels))
# + [markdown] deletable=true editable=true
# Got 89.9% - with rate decay every 500 steps
# + deletable=true editable=true
|
3_regularization.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import pandas as pd
import grblas
import dimsum as ds
from dimsum import Dimension, CalendarDimension, Schema, NULL, Flat, Pivot
from dimsum import alignment
size = Dimension('size', ['S', 'M', 'L', 'XL'])
shape = Dimension('shape', ['Circle', 'Square', 'Hexagon', 'Triangle', 'Pentagon', 'Octagon'])
color = Dimension('color', ['Red', 'Blue', 'Yellow', 'Green'])
quality = Dimension('quality', ['Bad', 'Okay', 'Good'])
# Calendar Dimensions
quarters = CalendarDimension('Q', pd.period_range('2020-Q1', '2021-Q4', freq='Q'), format='%Y-Q%q')
months = CalendarDimension('M', pd.period_range('2020-01', '2021-12', freq='M'))
years = CalendarDimension('Y', pd.period_range('2020', periods=5, freq='A'))
days = CalendarDimension('D', pd.period_range('2020-01-01', '2020-03-03', freq='D'))
schema = Schema([size, shape, color, quality, years, quarters, months, days])
list(schema.calendar)
schema.calendar['Q->Y']
# ### A
vals = [1, 2, 3, 4, 5]
codes = [
{'size': 'S', 'color': 'Green', 'shape': 'Circle'},
{'size': 'M', 'color': 'Green', 'shape': 'Triangle'},
{'size': 'L', 'color': 'Red', 'shape': 'Triangle'},
{'size': 'M', 'color': 'Blue', 'shape': 'Circle'},
{'size': 'S', 'color': 'Yellow', 'shape': 'Circle'},
]
df = pd.DataFrame(codes)
df['value'] = vals
s = df.set_index(['size', 'color', 'shape'])['value']
# Load from Series
a = schema.load(s)
a
# ### A_alt
vals = [10, 20, 30, 40, 50]
codes = [
{'size': 'S', 'color': 'Green', 'shape': 'Hexagon'},
{'size': 'M', 'color': 'Green', 'shape': 'Triangle'},
{'size': 'XL', 'color': 'Yellow', 'shape': 'Triangle'},
{'size': 'M', 'color': 'Blue', 'shape': 'Circle'},
{'size': 'S', 'color': 'Yellow', 'shape': 'Circle'},
]
df = pd.DataFrame(codes)
df['value'] = vals
# Load from DataFrame with value_column
a_alt = schema.load(df, dims=['size', 'color', 'shape'], value_column='value')
a_alt
# ### A_sub
vals = [1, 2, 3, 4, 5]
codes = [
{'size': 'S', 'color': 'Green'},
{'size': 'M', 'color': 'Green'},
{'size': 'L', 'color': 'Red'},
{'size': 'M', 'color': 'Blue'},
{'size': 'S', 'color': 'Yellow'},
]
df = pd.DataFrame(codes)
df['value'] = vals
# Load from DataFrame
a_sub = schema.load(df, ['size', 'color'])
a_sub
# ### B_sub
d = {
('S', 'Circle'): 10,
('M', 'Circle'): 20,
('L', 'Triangle'): 30,
('XL', 'Triangle'): 40,
}
# Load from dict
b_sub = schema.load(d, ['size', 'shape'])
b_sub
# ### B
data = [
['S', 'Circle', 'Bad', 5],
['S', 'Circle', 'Okay', 10],
['M', 'Circle', 'Okay', 20],
['L', 'Triangle', 'Good', 30],
['L', 'Triangle', 'Bad', 35],
['XL', 'Triangle', 'Bad', 40],
]
# Load from list of lists
b = schema.load(data, ['size', 'shape', 'quality'])
b
# ### C
data = {'Circle': 5, 'Square': 10, 'Hexagon': 20, 'Triangle': 30, 'Pentagon': 35, 'Octagon': 40}
# Load from dict with single dimension
c = schema.load(data, 'shape')
c
# ### D
df = pd.DataFrame({
'size': ['S', 'S', 'M', 'L', 'L', 'XL', NULL],
'quality': ['Bad', 'Okay', 'Okay', 'Good', 'Bad', NULL, None],
'vals': [1.1, 2.2, 3.3, 5.5, 4.4, 6.6, 7.7],
})
d = schema.load(df, ['quality', 'size'])
d
# ### Already aligned
x, y = ds.align(a.X[a_alt.X], a_alt.X[-1])
a
a_alt
x
y
a.X[a_alt.X] + a_alt.X[-100]
# ### Subset alignment
x, y = ds.align(a, b_sub)
a
b_sub
x
y
a + b_sub
a.X[-100] + b_sub
a + b_sub.X[-100]
# ### Partial disjoint alignment
x, y = ds.align(a, b)
a.pivot(left={'size', 'shape'})
b.pivot(left={'size', 'shape'})
x
y
a + b
a.X[-100] + b
a + b.X[-100]
# ### Fully disjoint alignment
x, y = ds.align(d, c)
c
d
x
y
d + c
# ### Pushdown
a
qual_pushdown = schema.load({'Bad': 0.05, 'Okay': 0.75, 'Good': 0.2}, 'quality')
qual_pushdown
# Verify that pushdown values sum to 1.0
qual_pushdown.reduce()
pushdown = a * qual_pushdown
pushdown
# ### Time Aggregation
csv = """
size,color,M,value
S,Red,2020-01,100.5
S,Blue,2020-01,174.2
M,Red,2020-01,77.5
M,Blue,2020-01,28.6
M,Green,2020-01,198.0
S,Red,2020-02,100.5
S,Blue,2020-02,174.2
M,Red,2020-02,71.5
M,Blue,2020-02,29.7
M,Green,2020-02,201.0
S,Red,2020-03,99.3
S,Blue,2020-03,177.1
M,Red,2020-03,62.1
M,Blue,2020-03,21.9
M,Green,2020-03,205.1
S,Red,2020-04,88.8
S,Blue,2020-04,173.3
M,Red,2020-04,72.5
M,Blue,2020-04,19.8
M,Green,2020-04,222.0
S,Red,2020-05,67.7
S,Blue,2020-05,171.1
M,Red,2020-05,77.7
M,Blue,2020-05,31.8
M,Green,2020-05,222.2
S,Red,2020-06,55.3
S,Blue,2020-06,164.6
M,Red,2020-06,79.3
M,Blue,2020-06,23.3
M,Green,2020-06,192.7
S,Red,2020-07,111.9
S,Blue,2020-07,177.1
M,Red,2020-07,66.6
M,Blue,2020-07,29.2
M,Green,2020-07,199.9
"""
import io
df = pd.read_csv(io.StringIO(csv))
data = schema.load(df.set_index(['size', 'color', 'M'])['value'])
data.pivot(top='M')
q2m = schema.calendar['M->Q']
q2m
x = data * q2m
x
xm = x.reduce_columns()
xm.pivot(top='Q')
data.pivot(top='M')
# # Random examples
a.pivot(top='color')
b
a + 2
a - b.pivot(top='quality').reduce_rows()
a.X[1] * b.X[1]
a.X[b]
b.X[a]
np.sin(a.pivot(left='size'))
np.arctan2(a, b.X)
np.arctan2(a, b.X[5.5])
np.sin(a)
a == 5
a << 4
a > 2
cond = schema.load(pd.DataFrame({'size': ['L', 'S', 'M'], 'shape': ['Circle', 'Triangle', 'Circle'], 'value': [True, False, False]}), ['size', 'shape'], 'value')
cond
a.filter(cond.X[True])
ds.where(cond.X[False], 5, b)
a.X[99].filter(cond)
b.filter(~cond)
x = schema.load({'S': 5, 'M': 0, 'L': 12.9, NULL: 44.4}, 'size')
x
y = schema.load({'XL': 14, 'L': 8}, 'size')
y
shifter = schema.load({'M': 1, NULL: 1}, 'size')
shifter
x.shift('size', -shifter)
shifter
shifter = schema.load({'M': 1}, 'size')
shifter
x.shift('size', shifter.X[0])
a
a.X+b
a.X[22].align(b)
a.X[b]
b.pivot(left={'size', 'shape'})
b.X[a]
a.pivot(left={'size', 'shape'})
c
a
c.cross_align(a)
a.cross_align(c)
a.align(c)
shifter = schema.load({'M': 1}, 'size')
shifter
a.shift('size', shifter.X)
a.codes('shape')
schema.dimension_enums('shape')
a.pivot(left='shape').reduce_rows()
a.filter(a.match(size={'M', 'S'}, color={'Yellow', 'Blue'}))
schema.encode(size='M')
a.filter(a.codes('size') > schema.encode(size='S'))
ds.where(a.match(size='S', color='Yellow'), -99, a)
|
notebooks/Playground.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/daquarti/AI/blob/main/segmentation.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="HOBX63US90d5"
# ## Image segmentation with CamVid
# + id="ICCdaQm7OpBb"
# + colab={"base_uri": "https://localhost:8080/"} id="7OPBSzL62Pmb" outputId="5496e03b-ba83-4a5c-9f79-c0178c8983d9"
# Load the Drive helper and mount
from google.colab import drive
# This will prompt for authorization.menos de 48hs de evolución
drive.mount('/content/drive')
# + id="y54ZuVr12qgP"
path = "/content/drive/My Drive/computer_vision/segmentation/train"
# + id="mjWRuZCc90d_"
# %reload_ext autoreload
# %autoreload 2
# %matplotlib inline
# + id="QiIinCWh90eA"
from fastai.vision import *
from fastai.callbacks.hooks import *
from fastai.utils.mem import *
import fastai
# fastai.torch_core.defaults.device = 'cpu'
# + id="wa9DDNyz90eA"
insulators = Path(path)
#codes = np.loadtxt(insulators/"codes.txt", dtype=str); codes
label = insulators/"masks"
image = insulators/"images"
get_y = lambda x: label/f"{x.stem}.png"
# + [markdown] id="Eygzsfrv90eB"
# ## Data
# + colab={"base_uri": "https://localhost:8080/"} id="ldLDetk090eB" outputId="a84f8dc3-44c6-4b88-da34-df56ada97b6e"
fnames = get_image_files(image)
fnames[:3]
# + colab={"base_uri": "https://localhost:8080/"} id="aKI7PUGt90eC" outputId="78082849-31b8-4468-cc5c-be9b6576a0c6"
lbl_names = get_image_files(label)
lbl_names[:3]
# + colab={"base_uri": "https://localhost:8080/", "height": 302} id="feLJzIjd90eC" outputId="ac4247fd-102a-466b-8fc5-33a5605b2b45"
img_f = fnames[2]
img = open_image(img_f)
img.show(figsize=(5,5))
# + id="QBWPzvWF90eD"
get_y_fn = lambda x: label/f"{x.stem}.png"
# + colab={"base_uri": "https://localhost:8080/", "height": 302} id="OgQmVKiD90eD" outputId="efc94811-1a30-4995-9016-d910a69bb061"
mask = open_mask(get_y_fn(img_f))
mask.show(figsize=(5,5), alpha=1)
# + colab={"base_uri": "https://localhost:8080/"} id="1ojDYQOd90eD" outputId="7f3994da-8559-4174-d3ca-625e2f30fb64"
src_size = np.array(mask.shape[1:])
src_size,mask.data
# + id="QruogNpA90eD"
#codes = np.loadtxt(insulators/'codes.txt', dtype=str); codes
# + id="JGd01wnmQmDk" colab={"base_uri": "https://localhost:8080/"} outputId="8e8ad37e-cdc9-4e70-d76d-b46743c3b6f5"
codes = np.array(['fondo', 'riñon'], dtype='<U17'); codes
# + [markdown] id="0iyffiM090eE"
# ## Datasets
# + colab={"base_uri": "https://localhost:8080/"} id="0aJlTQBj90eE" outputId="7cbd5397-039a-4724-fda8-1b9a9740caef"
size = (256,256)#src_size//2
print(size)
bs=2
# free = gpu_mem_get_free_no_cache()
# # the max size of bs depends on the available GPU RAM
# if free > 8200: bs=8
# else: bs=4
# print(f"using bs={bs}, have {free}MB of GPU RAM free")
# + id="1IUR172A90eE"
class SegLabelListCustom(SegmentationLabelList):
def open(self, fn): return open_mask(fn, div=True)
class SegItemListCustom(ImageList):
_label_cls = SegLabelListCustom
src = (SegItemListCustom.from_folder(image)
.split_by_rand_pct(0.2)
.label_from_func(get_y_fn, classes=codes) )
# + id="DS6htAs2rPe6" colab={"base_uri": "https://localhost:8080/", "height": 117} outputId="c5801e8f-e0aa-459d-855a-f91d2fd3fa67"
open_mask(fnames[2])
# + id="4DEY6GW2pe9C" colab={"base_uri": "https://localhost:8080/", "height": 117} outputId="882dec8c-b189-4ba4-e13e-c2825a88e0f4"
open_mask(get_y_fn(fnames[2]))
# + id="h-nmLvZ6rOJr"
# + id="Jq6nuvYQ90eE"
data = (src.transform(get_transforms(), size=size, tfm_y=True )
.databunch(bs=bs)
.normalize(imagenet_stats))
# + colab={"base_uri": "https://localhost:8080/", "height": 512} id="-T5ZFAkB90eF" outputId="a6918937-0113-4920-8523-2b28302f739c"
data.show_batch(5, figsize=(10,7))
# + colab={"base_uri": "https://localhost:8080/", "height": 512} id="J00PqUgF90eF" outputId="e827cb62-f6c5-425f-b45b-8d2cf3745122"
data.show_batch(5, figsize=(10,7), ds_type=DatasetType.Valid)
# + [markdown] id="qn66-w6V90eF"
# ## Model
# + colab={"base_uri": "https://localhost:8080/"} id="4u1Ybjye90eF" outputId="e483a2da-7643-4c42-8dfa-fab925cd3845"
name2id = {v:k for k,v in enumerate(codes)}
print(name2id)
# void_code = name2id['Void']
def acc_camvid(input, target):
target = target.squeeze(1)
# mask = target != void_code
return (input.argmax(dim=1)==target).float().mean()
# + id="tXo9kr_390eG"
metrics=acc_camvid
# metrics=accuracy
# + id="6kUUa6xU90eG"
wd=1e-2
# + colab={"base_uri": "https://localhost:8080/", "height": 81, "referenced_widgets": ["01a0baef0f4c49c5b6593bcb719860b7", "1221d4c2e20b403eb48a2d8536278fbb", "17ab90da1672445e9ae1d486e3872bab", "202d4e28aa7a4742937e75317918e916", "56b49622f02442c490e4d051ad5e4cd1", "be76ecdc3f7a40a28b47507c955709be", "b817fe1f02254059aa3f57fe3bfff955", "05082a431dd54cd1a60f08592075c277"]} id="294KAiMz90eG" outputId="c3da9b63-2497-4f3e-e13e-1e04a250956c"
learn = unet_learner(data, models.resnet34, metrics=metrics, wd=wd)
# + colab={"base_uri": "https://localhost:8080/"} id="eEB6Z43w90eG" outputId="b55129bd-f80e-49aa-e2b5-51ac643b3ae0"
data.classes
# + colab={"base_uri": "https://localhost:8080/", "height": 373} id="9adacsve90eG" outputId="cb80b26d-a7d7-40ff-c69c-d7f1fb6f31f1"
lr_find(learn)
learn.recorder.plot()
# + id="WnBUmc2u90eH"
lr=1e-5
# + colab={"base_uri": "https://localhost:8080/", "height": 335} id="rDIb0DLE90eH" outputId="91688463-93ee-4e62-a608-93244dd7fb1e"
learn.fit_one_cycle(10, slice(lr), pct_start=0.9)
# + id="-1IzrpNK90eH"
learn.save(path + 'stage-1')
# + id="ylmRCu4G90eH"
learn.load(path + 'stage-1');
# + colab={"base_uri": "https://localhost:8080/", "height": 117} id="aD_OwJ0K90eI" outputId="b7686eaa-016a-49ae-b893-2f41991ea15d"
img = open_image(fnames[2])
# + id="mc3-ztIF90eI"
prediction = learn.predict(img)
# + id="8LQhIpz3ewfz" colab={"base_uri": "https://localhost:8080/", "height": 272} outputId="9e4b8b62-9d6e-4f92-b8f7-7ac6ecf9adc1"
prediction[0]
# + colab={"base_uri": "https://localhost:8080/", "height": 522} id="r4r88q8r90eI" outputId="f3aa9e4c-75c9-4921-b12e-18ce299acac7"
learn.show_results(rows=3, figsize=(8,9))
# + id="b7kF_IoZ90eI"
learn.unfreeze()
# + colab={"base_uri": "https://localhost:8080/", "height": 371} id="O4POtw2O90eI" outputId="2776e3b3-99a8-4c22-b8d9-b20b18edeef8"
lr_find(learn)
learn.recorder.plot()
# + id="EvnPQ2kL90eJ"
lr = 1e-5
lrs = slice(lr/400,lr/4)
# + colab={"base_uri": "https://localhost:8080/", "height": 335} id="TctSDJr890eJ" outputId="18c0e5a3-8fb7-4519-87d4-e9e548672a6e"
learn.fit_one_cycle(10, lrs, pct_start=0.8)
# + id="j1olkLTH90eJ"
learn.save(path + 'stage-2');
# + [markdown] id="ebtelBcu90eJ"
# ## Go big
# + [markdown] id="LCI-vtuw90eJ"
# You may have to restart your kernel and come back to this stage if you run out of memory, and may also need to decrease `bs`.
# + id="iC63zo9x90eK"
#learn.destroy() # uncomment once 1.0.46 is out
size = (512, 512)
free = gpu_mem_get_free_no_cache()
# the max size of bs depends on the available GPU RAM
if free > 8200: bs=3
else: bs=1
print(f"using bs={bs}, have {free}MB of GPU RAM free")
# + id="GWkty52M90eK"
data = (src.transform(get_transforms(), size=size, tfm_y=True)
.databunch(bs=bs)
.normalize(imagenet_stats))
# + id="2aTmMtxJ90eK"
learn = unet_learner(data, models.resnet34, metrics=metrics, wd=wd)
# + id="xbmdGhH990eK"
learn.load('stage-2');
# + id="TaoQWm-g90eL"
lr_find(learn)
learn.recorder.plot()
# + id="_OeZsgX390eL"
lr=1e-6
# + id="bPY-IdMP90eL"
learn.fit_one_cycle(10, slice(lr), pct_start=0.8)
# + id="UodjK_8x90eL"
learn.save('stage-1-big')
# + id="r82eV0Wc90eL"
learn.load('stage-1-big');
# + id="hHTIV8jE90eM"
learn.unfreeze()
# + id="HVDzbJvC90eM"
lr_find(learn)
learn.recorder.plot()
# + id="6otDzsLM90eM"
lrs = slice(1e-5,lr/10)
# + id="F60E3bB990eM"
learn.fit_one_cycle(10, lrs)
# + id="RKCiUxR190eN"
learn.save('stage-2-big')
# + id="aylWOJZd90eN"
learn.load('stage-2-big');
# + id="_akXtVTq90eN"
img = open_image('/home/drones/fastai/caravan/image/4df1536de792_11.jpg')
# + id="nBo13ZYM90eN"
prediction = learn.predict(img)[0]
# + id="8lJEryjU90eN"
prediction
# + id="ujfVB4mk90eN"
|
segmentation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Tutorial Overview
# This set of five tutorials (installation, package setup, data setup, running, analyzing) will explain the UncertaintyForest class. After following the steps below, you should have the ability to run the code on your own machine and interpret the results.
#
# If you haven't seen it already, take a look at the first and second parts of this set of tutorials called `UncertaintyForest_Tutorials_1-Installation` and `UncertaintyForest_Tutorial_2-Package-Setup`
#
# # 3: Data Setup
# ## *Goal: Understand the data and the parameters that will be passed to the UncertaintyForest instance*
# ### First, we have to import some modules to have everything we need.
# The top two sections are standard packages, the third block is just specifying where to look for the packages listed below, the fourth block is another standard package, and the final block is for importing the actual UncertaintyForest class.
# +
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.ensemble import RandomForestClassifier
from sklearn.calibration import CalibratedClassifierCV
from sklearn.model_selection import train_test_split
from sklearn.ensemble import BaggingClassifier
from sklearn.tree import DecisionTreeClassifier
from tqdm.notebook import tqdm
from joblib import Parallel, delayed
from proglearn.forest import UncertaintyForest
# -
# ### Now, we create the function that will make data that we'll train on.
# Here, we use randomized data because if the learner can learn that, then it can learn most anything.
def generate_data(n, d, var):
'''
Parameters
---
n : int
The number of data to be generated
d : int
The number of features to generate for each data point
var : double
The variance in the data
'''
# create the mean matrix for the data (here it's just a mean of 1)
means = [np.ones(d) * -1, np.ones(d)]
# create the data with the given parameters (variance)
X = np.concatenate([np.random.multivariate_normal(mean, var * np.eye(len(mean)),
size=int(n / 2)) for mean in means])
# create the labels for the data
y = np.concatenate([np.ones(int(n / 2)) * mean_idx for mean_idx in range(len(means))])
return X, y
# ### Lastly, the parameters of the uncertainty forest are defined.
# Real Params.
n_train = 50
n_test = 10000
d = 100
var = 0.25
num_trials = 10
n_estimators = 100
# It will be important to understand each of these parameters, so we'll go into more depth on what they mean:
# * `n_train` is the number of training data that will be used to train the learner
# * `n_test` is the number of test data that will be used to assess how well the learner is at classifying
# * `d` is the dimensionality of the input space (i.e. how many features the data has)
# * `var` is the variance of the data
# * `num_trials` is the number of times we'll generate data, train, and test to make sure our results are not outliers
# * `num_estimators` is the number of trees in the forest
# ### You're done with part 3 of the tutorial!
#
# ### Move on to part 4 (called "UncertaintyForest_Tutorial_4-Running")
|
tutorials/UncertaintyForestTutorials/UncertaintyForest_Tutorial_3-Data-Setup.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Dcrawf/Deep-Dive-Final/blob/master/DeepDiveFinal.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="fax_AbV6wI6b" colab_type="text"
# # Welcome to Deep Dive
#
# ---
#
# + [markdown] id="jx3adb2Ewqao" colab_type="text"
# Deep Dive is the product of my exploration of Google's Deep Dream program. It is built off the existing Deep Dream code by <NAME>, and uses the Inception trained model. Each code cell must be run in order. You can either run them all by hand, by pressing the play button on the top left corner of each code cell, or you may press ctrl+f9 to run them all.
#
# + [markdown] id="5PgZRSpkZx23" colab_type="text"
# Download the Inception model from Google.
# + id="8cMJYa69bhp9" colab_type="code" outputId="cd06039a-d873-44c4-9e45-18c08ab57b68" colab={"base_uri": "https://localhost:8080/", "height": 561}
# !wget --no-check-certificate https://storage.googleapis.com/download.tensorflow.org/models/inception5h.zip && unzip -n inception5h.zip
# !wget https://github.com/tensorflow/tensorflow/raw/master/tensorflow/examples/tutorials/deepdream/pilatus800.jpg
with open('pilatus800.jpg','rb') as f:
file_contents = f.read()
print("Inception model downloaded.")
# + id="63tM5etcczm6" colab_type="code" colab={}
from io import BytesIO
from IPython.display import clear_output, Image, display
import numpy as np
import PIL.Image
import tensorflow as tf
from __future__ import print_function
# https://github.com/Dcrawf/Deep-Dive-Assets
# https://github.com/ProGamerGov/Protobuf-Dreamer
# https://github.com/ProGamerGov/Protobuf-Dreamer/wiki/Interesting-Layers-And-Channels
# http://storage.googleapis.com/deepdream/visualz/tensorflow_inception/index.html
# start with a gray image with a little noise
img_noise = np.random.uniform(size=(224,224,3)) + 100.0
# Create tensorflow session
model_fn = 'tensorflow_inception_graph.pb'
# creating TensorFlow session and loading the model
# Then initialize graph using graph() function of TensorFlow
graph = tf.Graph()
# Initialize a session using the graph
sess = tf.InteractiveSession(graph=graph)
# Open existing saved in-session graph
with tf.gfile.GFile(model_fn, 'rb') as f:
# Once we open it, we can read the graph and parse it
graph_def = tf.GraphDef()
# Using the ParseFromString() method of TensorFlow
graph_def.ParseFromString(f.read())
# We need to define out input so we create an input tensor using the placeholder() method
# Called input with the size of 32 bits
t_input = tf.placeholder(np.float32, name='input')
# Then we define image net mean value of pixels in an image as 117
imagenet_mean = 117.0
# We subtract it from the input tensor and store it in the preprocessed variable
t_preprocessed = tf.expand_dims(t_input-imagenet_mean, 0)
# Then load the graph_def variable we initialized as the newly processed tensor
tf.import_graph_def(graph_def, {'input':t_preprocessed})
def T(layer):
'''Helper for getting layer output tensor'''
return graph.get_tensor_by_name("import/%s:0"%layer)
# + [markdown] id="uwwRKVx_JRwV" colab_type="text"
# **Please only upload .jpg files for now**
# + id="8q4tY63WcuGl" colab_type="code" outputId="acaeadc6-a608-4541-f7d8-d2f59b6c7939" colab={"resources": {"http://localhost:8080/nbextensions/google.colab/files.js": {"data": "<KEY> "ok": true, "headers": [["content-type", "application/javascript"]], "status": 200, "status_text": ""}}, "base_uri": "https://localhost:8080/", "height": 74}
from google.colab import files
uploaded = files.upload()
# + id="X_nNQ6Jduc-2" colab_type="code" colab={}
if type(uploaded) is not dict: uploaded = uploaded.files ## Deal with filedit versions
file_contents = uploaded[list(uploaded.keys())[0]]
# + [markdown] id="niMe7Xu-I_-s" colab_type="text"
# # Show the Image Before the DeepDream
# + id="zS04LJnEdWr-" colab_type="code" outputId="edc98d05-d63d-4717-bf85-8d2417aec163" colab={"base_uri": "https://localhost:8080/", "height": 868}
def showarray(a, fmt="jpeg"):
# Ensure the pixel-values are between 0 and 255
a = np.uint8(np.clip(a, 0, 255))
f = BytesIO()
PIL.Image.fromarray(a).save(f, fmt)
display(Image(data=f.getvalue()))
img0 = sess.run(tf.image.decode_image(file_contents))
showarray(img0)
# + id="zyfMQcrZbjPQ" colab_type="code" colab={}
# These parameters let us control the strenth of the deepdream.
octave_n = 4
octave_scale = 1.4
iter_n = 10
strength = 200
# Helper function that uses TensorFlow to resize an image
def resize(img, new_size):
return sess.run(tf.image.resize_bilinear(img[np.newaxis,:], new_size))[0]
# Apply gradients to an image in a series of tiles
def calc_grad_tiled(img, t_grad, tile_size=256):
'''Random shifts are applied to the image to blur tile boundaries over
multiple iterations.'''
h, w = img.shape[:2]
sx, sy = np.random.randint(tile_size, size=2)
# We randomly roll the image in x and y to avoid seams between tiles.
img_shift = np.roll(np.roll(img, sx, 1), sy, 0)
grad = np.zeros_like(img)
for y in range(0, max(h-tile_size//2, tile_size),tile_size):
for x in range(0, max(w-tile_size//2, tile_size),tile_size):
sub = img_shift[y:y+tile_size,x:x+tile_size]
g = sess.run(t_grad, {t_input:sub})
grad[y:y+tile_size,x:x+tile_size] = g
imggrad = np.roll(np.roll(grad, -sx, 1), -sy, 0)
# Add the image gradient to the image and return the result
return img + imggrad*(strength * 0.01 / (np.abs(imggrad).mean()+1e-7))
# Applies deepdream at multiple scales
def render_deepdream(t_obj, input_img, show_steps = True):
# Collapse the optimization objective to a single number (the loss)
t_score = tf.reduce_mean(t_obj)
# We need the gradient of the image with respect to the objective
t_grad = tf.gradients(t_score, t_input)[0]
# split the image into a number of octaves (laplacian pyramid)
img = input_img
octaves = []
for i in range(octave_n-1):
lo = resize(img, np.int32(np.float32(img.shape[:2])/octave_scale))
octaves.append(img-resize(lo, img.shape[:2]))
img = lo
# generate details octave by octave
for octave in range(octave_n):
if octave>0:
hi = octaves[-octave]
img = resize(img, hi.shape[:2])+hi
for i in range(iter_n):
img = calc_grad_tiled(img, t_grad)
if show_steps:
clear_output()
showarray(img)
return img
# + [markdown] id="Az9wbYd0J4rK" colab_type="text"
# # Output Image
#
# ---
#
# + [markdown] id="xcAoR92tLJWk" colab_type="text"
# * Octave: The input image is downscaled, and gradient ascent is applied to all the images, and then the result is merged into the final image
# * Octave scale: Relative scale the ascent is applied to
# * Iteration: How many images are in the feedback loop
# * Strength: How strongly the DeepDream is applied to the image
# * Layer: Reference to a tensor that will be maximized
#
#
#
# **After making changes to the slider, please press the play button again to apply changes to your image.**
#
#
#
# + id="r3vN7FJlw-VW" colab_type="code" outputId="9d0203a8-05e9-48de-d825-0bc347c9ac31" colab={"base_uri": "https://localhost:8080/", "height": 797}
octave_n = 4 #@param {type:"slider", max: 10}
octave_scale = 2 #@param {type:"number"}
iter_n = 14 #@param {type:"slider", max: 50}
strength = 252 #@param {type:"slider", max: 1000}
layer = "mixed3b" #@param ["mixed3a", "mixed3b", "mixed4a", "mixed4c", "mixed5a"]
final = render_deepdream(tf.square(T(layer)), img0)
|
DeepDiveFinal.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/novoforce/Exploring-Tensorflow/blob/main/Edurekha_tf_course/Assignment_3_q3.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + colab={"base_uri": "https://localhost:8080/"} id="PZcGXb8-yxEH" outputId="3c43c032-1083-4b74-8bfe-297844415894"
import tensorflow as tf
from tensorflow import keras
from keras.layers import Dense,Input
from keras.models import Model
import matplotlib.pyplot as plt
print(tf.__version__)
# + [markdown] id="274-x57DzDXE"
# # Data Exploration
# + colab={"base_uri": "https://localhost:8080/"} id="KmX3fqFqy79Y" outputId="93c4adcc-3397-47e9-dd9c-8175aa7b16f1"
(x_train,y_train), (x_test, y_test) = keras.datasets.fashion_mnist.load_data()
print("Shape of the datasets:> ",x_train.shape,x_test.shape)
# + colab={"base_uri": "https://localhost:8080/"} id="CHsPsvkk0z-s" outputId="8ba99598-f03b-486f-90a7-20a47afaaa1e"
from collections import Counter
items_train = Counter(y_train).keys()
items_test = Counter(y_test).keys()
print("No of unique items in the dataset are:", len(items_train),len(items_test))
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="zJKJbpnYzgNC" outputId="eb678ebc-1820-4feb-88fd-a2c8efdfc6b8"
plt.figure(figsize=(12, 12))
for i in range(0,9):
plt.subplot(5, 3, i+1)
plt.imshow(x_train[i], cmap="gray") #'image_list' is the list of images
plt.tight_layout()
plt.show()
# + [markdown] id="r15T7vQF2DJU"
# # Data Pre-processing:
# + id="N0NSoJUy0PId"
#Normalize the dataset between 0 & 1
x_train= x_train.astype('float32') / 255
x_test= x_test.astype('float32') / 255
# + id="bX49bR_p8EFw"
#Adding Salt and Pepper noise in the images
import numpy as np
from skimage.util import random_noise
for i in range(len(x_train)):
x_train[i]= random_noise(x_train[i],mode='gaussian',var=0.01) #salt and pepper noise
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="IiQtT5kZ2wW5" outputId="989c5658-febf-47e8-c9e5-a861887c639c"
plt.figure(figsize=(12, 12))
for i in range(0,9):
plt.subplot(5, 3, i+1)
plt.imshow(x_train[i], cmap="gray") #'image_list' is the list of images
plt.tight_layout()
plt.show()
# + id="ZuYFueXvBEmR"
x_train= x_train.reshape((len(x_train), np.prod(x_train.shape[1:])))
x_test= x_test.reshape((len(x_test), np.prod(x_test.shape[1:])))
# + [markdown] id="29zd_nmQALU0"
# # Autoencoder
# + colab={"base_uri": "https://localhost:8080/"} id="3b3ALtRp2wzE" outputId="e20ba64d-42ee-421d-e941-4a41631c57c8"
# 784-> 32-> 784
encoding_dim= 32 # compress the image to the factor of 784/32 == 24.5
input_img= Input(shape=(784,))
encoded= Dense(encoding_dim,activation="relu")(input_img) #encoded representation
decoded= Dense(784,activation="sigmoid")(encoded) #reconstruction from the encoded form
#The reason behind usage of sigmoid is so as to get the image in the range of 0-1(normalized)
auto_encoder= Model(input_img,decoded)
auto_encoder.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="2mHgKs6VARPx" outputId="562eeb9c-3b51-47bd-ef17-5dd0f8944960"
encoder= Model(input_img,encoded)
encoder.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="rOWnATo2ATWA" outputId="d3ccfbf0-ebb8-4f7a-b7a4-323ce53b7890"
# Build decoder
encoded_input= Input(shape=(encoding_dim,)) # flexibility of giving our own input
decoder_layer= auto_encoder.layers[-1] (encoded_input) #retreiving the last layer of the autoencoder(to keep the trained weights)
decoder= Model(encoded_input,decoder_layer)
decoder.summary()
# + colab={"base_uri": "https://localhost:8080/"} id="zyN9d8-zAVW-" outputId="50cde150-bf3c-4869-d419-c48ad91b6d29"
auto_encoder.compile(optimizer='adam',loss='binary_crossentropy',metrics=['accuracy'])
history= auto_encoder.fit(x_train,x_train,epochs=10,batch_size=256,shuffle=True,validation_data=(x_test,x_test))
# + [markdown] id="JUJvdIUQFyWE"
# # Evaluating the model performance
# + id="5NJyDLs8A8ga"
encoded_img= encoder.predict(x_test)
decoded_img= decoder.predict(encoded_img)
# + [markdown] id="bNnBm47XF3Nt"
# # Visualize the trained model output
# + colab={"base_uri": "https://localhost:8080/", "height": 542} id="3w03k3ngDUX5" outputId="9189066b-8770-4722-b8ce-42e7c55b1d5d"
plt.figure(figsize=(12, 12))
for i in range(0,9):
plt.subplot(5, 3, i+1)
plt.imshow(decoded_img[i].reshape(28,28), cmap="gray") #'image_list' is the list of images
# plt.xlabel(CLASSES[labels[i]]) # 'labels' is the list of labels
plt.tight_layout()
plt.show()
# + [markdown] id="U7BsT-CJGC6G"
# # Performance metrics plotting
# + colab={"base_uri": "https://localhost:8080/", "height": 591} id="tT3UcGQYDbF-" outputId="7c36c508-eba0-4d61-9077-a945b10a7bc1"
# summarize training for accuracy
print(history.history.keys())
plt.plot(history.history['accuracy']) # training is the variable from the fit method
plt.plot(history.history['val_accuracy'])
plt.title('model accuracy')
plt.ylabel('accuracy')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# summarize traning for loss
plt.plot(history.history['loss'])
plt.plot(history.history['val_loss'])
plt.title('model loss')
plt.ylabel('loss')
plt.xlabel('epoch')
plt.legend(['train', 'test'], loc='upper left')
plt.show()
# + [markdown] id="S4BFDQ703Kvv"
# # Summary
#
# In this project we created an Auto-encoder which de-noises the image from Fashion-MNIST dataset.
#
# **Architectural design**
#
# An auto-encoder consists of a Encoder and a Decoder model combined together to form a Auto-encoder.
#
# As the name suggests:
#
# Encoder will encode the image/data and the decoder will decode(reconstructs) the image/data.
#
# Main point to note is that it's an Unsupervised way of training the model. So here in this experiment setting the (X,Y) both are the images.
#
# **Experiment design**
#
# **Encoder**
#
# We have designed an Encoder with layers as:
#
# 784 --> 32
#
# Here a 28x28(784 vector) image is scaled down to mare 32 vector representation.
# This compression forces the neural network to virtually discard the noise and only learn the important representation.
#
# **Decoder**
#
# 32 --> 784
#
# Here the 32 vector is scaled up to the actual 784 vector representation.
# This de-compression forces the neural network to re-construct the image back.
# Since the compressed representation does not contain the noise, So the de-compressed representation will not contain the noise.
#
# Thus achieving the de-noising capability.
#
# + id="J7mtJlFlEvQz"
|
Edurekha_tf_course/Assignment_3/Assignment_3_q3.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# LIL' GRAPHICS
#
# Use Excel, Illustrator or something like https://infogr.am/ to make a graphic about the Lil's, or the Lil's vs. the Biggies. Just a simple bar graph of their various popularities sounds good to me.
#
# SEARCH ENGINE
#
# Make a non-IPython Notebook that automates browsing for top tracks
# Prompts for an artist
# you put it in, displays the results, asks which one you want (numbered)
# you enter a number
# It displays their top tracks, then their MOST popular album and their least popular album. if they only have one album it says that they only have one album.
|
05/Spotify-API-2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import sys
sys.path.append('../scripts/')
from dp_policy_agent import *
from dynamic_programming import *
class QmdpAgent(DpPolicyAgent): ###qmdp2
def __init__(self, time_interval, estimator, goal, puddles, sampling_num=10, widths=np.array([0.2, 0.2, math.pi/18]).T, \
puddle_coef=100.0, lowerleft=np.array([-4, -4]).T, upperright=np.array([4, 4]).T):
super().__init__(time_interval, estimator, goal, puddle_coef, widths, lowerleft, upperright)
self.dp = DynamicProgramming(widths, goal, puddles, time_interval, sampling_num)
self.dp.value_function = self.init_value()
self.evaluations = np.array([0.0, 0.0, 0.0]) #Q_MDP値を入れる。描画用
self.current_value = 0.0 #現在の状態価値の平均値を入れる。描画用
def init_value(self):
tmp = np.zeros(self.dp.index_nums)
for line in open("value.txt", "r"):
d = line.split()
tmp[int(d[0]), int(d[1]), int(d[2])] = float(d[3])
return tmp
def evaluation(self, action, indexes): #これ以降追加
return sum([self.dp.action_value(action, i, out_penalty=False) for i in indexes])/len(indexes) #パーティクルの重みの正規化が前提
def policy(self, pose, goal=None):
indexes = [self.to_index(p.pose, self.pose_min, self.index_nums, self.widths) for p in self.estimator.particles]
self.current_value = sum([self.dp.value_function[i] for i in indexes])/len(indexes) #描画用に計算
self.evaluations = [self.evaluation(a, indexes) for a in self.dp.actions]
return self.dp.actions[np.argmax(self.evaluations)]
def draw(self, ax, elems):
super().draw(ax, elems)
elems.append(ax.text(-4.5, -4.6, "{:.3} => [{:.3}, {:.3}, {:.3}]".format(self.current_value, *self.evaluations), fontsize=8))
def trial(animation):
time_interval = 0.1
world = PuddleWorld(30, time_interval, debug=not animation)
##ランドマークの追加(意地悪な位置に)##
m = Map()
for ln in [(1,4), (4,1), (-4,-4)]: m.append_landmark(Landmark(*ln))
world.append(m)
##ゴール・水たまりの追加(これは特に変更なし)##
goal = Goal(-3,-3)
puddles = [Puddle((-2, 0), (0, 2), 0.1), Puddle((-0.5, -2), (2.5, 1), 0.1)]
world.append(goal)
world.append(puddles[0])
world.append(puddles[1])
##ロボットを作る##
init_pose = np.array([2.5, 2.5, 0]).T
pf = Mcl(m, init_pose, 100)
a = QmdpAgent(time_interval, pf, goal, puddles)
r = Robot(init_pose, sensor=Camera(m), agent=a, color="red")
world.append(r)
world.draw()
return a
a = trial(True)
|
section_pomdp/qmdp2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
import cranet
from cranet import nn, optim
from cranet.nn import functional as F
from cranet.data import Dataset, DataLoader
import numpy as np
from matplotlib import pyplot as plt
from sklearn.metrics import confusion_matrix
import itertools
import os
print(cranet.__version__)
# -
class MnistDataset(Dataset):
train_img = 'train-images-idx3-ubyte'
train_lab = 'train-labels-idx1-ubyte'
test_img = 't10k-images-idx3-ubyte'
test_lab = 't10k-labels-idx1-ubyte'
def __init__(self, root, mode, transform=None, transform_target=None):
self.mode = mode
self.transform = transform
self.transform_target = transform_target
self.images = []
self.labels = []
self._load_data(os.path.join(root, 'MNIST', 'raw'))
def _load_data(self, data_dir):
if self.mode == 'train':
image_file = os.path.join(data_dir, self.train_img)
label_file = os.path.join(data_dir, self.train_lab)
elif self.mode == 'test':
image_file = os.path.join(data_dir, self.test_img)
label_file = os.path.join(data_dir, self.test_lab)
else:
raise RuntimeError('mode must be train or test')
with open(image_file, 'rb') as f:
f.read(4) # magic
self.size = int.from_bytes(f.read(4), "big")
r = int.from_bytes(f.read(4), "big")
c = int.from_bytes(f.read(4), "big")
for _ in range(self.size):
mat = []
for i in range(r):
mat.append([])
for j in range(c):
mat[i].append(int.from_bytes(f.read(1), "big"))
self.images.append(np.array(mat))
with open(label_file, 'rb') as f:
f.read(4) # magic
sz = int.from_bytes(f.read(4), "big") # size
assert self.size == sz
for _ in range(self.size):
lab = np.array(int.from_bytes(f.read(1), "big"))
self.labels.append(lab)
def __len__(self):
return self.size
def __getitem__(self, idx):
img = self.images[idx]
lab = self.labels[idx]
if self.transform is not None:
img = self.transform(img)
if self.transform_target is not None:
lab = self.transform_target(lab)
return img, lab
# +
def trans(x):
x = x.reshape(28*28)
return cranet.as_tensor(x)
def trans_lab(x):
return cranet.as_tensor(x)
train_ds = MnistDataset('data', 'train', trans, trans_lab)
test_ds = MnistDataset('data', 'test', trans, trans_lab)
# -
train_ld = DataLoader(train_ds, 64)
test_ld = DataLoader(test_ds, 1000)
sample_image_batch, sample_label_batch = next(iter(train_ld))
sample_image = sample_image_batch.numpy()[0]
sample_label = sample_label_batch.numpy()[0]
print(sample_image_batch.shape)
print(sample_label_batch.shape)
plt.imshow(sample_image.reshape(28, 28))
print(sample_label)
class Model(nn.Module):
def __init__(self):
super().__init__()
self.l0 = nn.Linear(28 * 28, 64)
self.l1 = nn.Linear(64, 64)
self.l2 = nn.Linear(64, 32)
self.final = nn.Linear(32, 10)
def forward(self, x: cranet.Tensor) -> cranet.Tensor:
x = self.l0(x)
x = F.sigmoid(x)
x = self.l1(x)
x = F.sigmoid(x)
x = self.l2(x)
x = F.sigmoid(x)
x = self.final(x)
x = F.log_softmax(x, dim=1)
return x
model = Model()
print(model)
optm = optim.Adam(model.parameters())
def train(model, train_loader, optimizer, epoch):
model.train()
for batch_idx, (inp, lab) in enumerate(train_loader):
optimizer.zero_grad()
out = model(inp)
loss = F.nll_loss(out, lab)
loss.backward()
optimizer.step()
loss_v = loss.item()
if batch_idx % 100 == 0:
print('Train Epoch: {} [{}/{} ({:.0f}%)]\tLoss: {:.6f}'.format(
epoch, batch_idx * 64, len(train_loader.dataset),
100. * batch_idx / len(train_loader), loss_v))
def test(model, loader, label=''):
model.eval()
loss = 0
correct = 0
with cranet.no_grad():
for inp, lab in loader:
out = model(inp)
loss += F.nll_loss(out, lab, reduction='sum').item()
pre = out.numpy().argmax(axis=1)
correct += (pre == lab.numpy()).sum().item()
data_size = len(loader.dataset)
loss /= data_size
accu = correct / data_size
print('\n{} set: Average loss: {:.4f}, Accuracy: {}/{} ({:.0f}%)\n'.format(
label, loss, correct, data_size, accu*100.))
return accu, loss
train_loss = []
train_accu = []
test_loss = []
test_accu = []
for epoch in range(10):
train(model, train_ld, optm, epoch)
accu, loss = test(model, train_ld, 'Train')
train_loss.append(loss)
train_accu.append(accu)
accu, loss = test(model, test_ld, 'Test')
test_loss.append(loss)
test_accu.append(accu)
plt.figure()
plt.title('loss')
plt.plot(train_loss, label='train loss')
plt.plot(test_loss, label='test loss')
plt.legend()
plt.show()
plt.figure()
plt.title("accuracy")
plt.plot(train_accu, label='train_accu')
plt.plot(test_accu, label='test_accu')
plt.legend()
plt.show()
def eval(model, loader):
pre_arr = []
tar_arr = []
for inp, tar in loader:
out = model(inp)
pre = out.numpy().argmax(axis=1)
pre_arr.append(pre)
tar_arr.append(tar.numpy())
pre_arr = np.concatenate(pre_arr)
tar_arr = np.concatenate(tar_arr)
return confusion_matrix(tar_arr, pre_arr)
cm = eval(model, test_ld)
def plot_confusion_matrix(cm,
target_names,
title='Confusion matrix',
cmap=None,
normalize=True):
"""
given a sklearn confusion matrix (cm), make a nice plot
Arguments
---------
cm: confusion matrix from sklearn.metrics.confusion_matrix
target_names: given classification classes such as [0, 1, 2]
the class names, for example: ['high', 'medium', 'low']
title: the text to display at the top of the matrix
cmap: the gradient of the values displayed from matplotlib.pyplot.cm
see http://matplotlib.org/examples/color/colormaps_reference.html
plt.get_cmap('jet') or plt.cm.Blues
normalize: If False, plot the raw numbers
If True, plot the proportions
Usage
-----
plot_confusion_matrix(cm = cm, # confusion matrix created by
# sklearn.metrics.confusion_matrix
normalize = True, # show proportions
target_names = y_labels_vals, # list of names of the classes
title = best_estimator_name) # title of graph
Citiation
---------
http://scikit-learn.org/stable/auto_examples/model_selection/plot_confusion_matrix.html
"""
accuracy = np.trace(cm) / float(np.sum(cm))
misclass = 1 - accuracy
if cmap is None:
cmap = plt.get_cmap('Blues')
plt.figure(figsize=(8, 6))
plt.imshow(cm, interpolation='nearest', cmap=cmap)
plt.title(title)
plt.colorbar()
if target_names is not None:
tick_marks = np.arange(len(target_names))
plt.xticks(tick_marks, target_names, rotation=45)
plt.yticks(tick_marks, target_names)
if normalize:
cm = cm.astype('float') / cm.sum(axis=1)[:, np.newaxis]
thresh = cm.max() / 1.5 if normalize else cm.max() / 2
for i, j in itertools.product(range(cm.shape[0]), range(cm.shape[1])):
if normalize:
plt.text(j, i, "{:0.4f}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
else:
plt.text(j, i, "{:,}".format(cm[i, j]),
horizontalalignment="center",
color="white" if cm[i, j] > thresh else "black")
plt.tight_layout()
plt.ylabel('True label')
plt.xlabel('Predicted label\naccuracy={:0.4f}; misclass={:0.4f}'.format(
accuracy, misclass))
plt.show()
plot_confusion_matrix(cm, [0, 1, 2, 3, 4, 5, 6, 7, 8, 9])
|
examples/mnist/mnist.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.6 (tensorflow)
# language: python
# name: rga
# ---
# + [markdown] colab_type="text" id="LSIM-PITWYFa"
# <a href="https://colab.research.google.com/github/jeffheaton/t81_558_deep_learning/blob/master/t81_558_class_06_3_resnet.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] colab_type="text" id="YDTXd8-Lmp8Q"
# # T81-558: Applications of Deep Neural Networks
# **Module 6: Convolutional Neural Networks (CNN) for Computer Vision**
# * Instructor: [<NAME>](https://sites.wustl.edu/jeffheaton/), McKelvey School of Engineering, [Washington University in St. Louis](https://engineering.wustl.edu/Programs/Pages/default.aspx)
# * For more information visit the [class website](https://sites.wustl.edu/jeffheaton/t81-558/).
# + [markdown] colab_type="text" id="ncNrAEpzmp8S"
# # Module 6 Material
#
# * Part 6.1: Image Processing in Python [[Video]](https://www.youtube.com/watch?v=4Bh3gqHkIgc&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_1_python_images.ipynb)
# * Part 6.2: Keras Neural Networks for Digits and Fashion MNIST [[Video]](https://www.youtube.com/watch?v=-SA8BmGvWYE&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_2_cnn.ipynb)
# * **Part 6.3: Implementing a ResNet in Keras** [[Video]](https://www.youtube.com/watch?v=qMFKsMeE6fM&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_3_resnet.ipynb)
# * Part 6.4: Using Your Own Images with Keras [[Video]](https://www.youtube.com/watch?v=VcFja1fUNSk&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_4_keras_images.ipynb)
# * Part 6.5: Recognizing Multiple Images with YOLO Darknet [[Video]](https://www.youtube.com/watch?v=oQcAKvBFli8&list=PLjy4p-07OYzulelvJ5KVaT2pDlxivl_BN) [[Notebook]](t81_558_class_06_5_yolo.ipynb)
# -
# # Google CoLab Instructions
#
# The following code ensures that Google CoLab is running the correct version of TensorFlow.
# + colab={"base_uri": "https://localhost:8080/", "height": 51} colab_type="code" id="fU9UhAxTmp8S" outputId="e6641ea1-be85-45a9-bfbc-b12b18669d74"
# Detect Colab if present
try:
from google.colab import drive
COLAB = True
print("Note: using Google CoLab")
# %tensorflow_version 2.x
except:
print("Note: not using Google CoLab")
COLAB = False
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return f"{h}:{m:>02}:{s:>05.2f}"
# + [markdown] colab_type="text" id="Q09yMGGcmp9N"
# # Part 6.3: Implementing a ResNet in Keras
#
# Deeper neural networks are more difficult to train. Residual learning was introduced to ease the training of networks that are substantially deeper than those used previously. ResNet explicitly reformulates the layers as learning residual functions with reference to the layer inputs, instead of learning unreferenced functions. On the ImageNet dataset this method was evaluated with residual nets with a depth of up to 152 layers---8x deeper than VGG nets but still having lower complexity. An ensemble of these residual nets achieves 3.57% error on the ImageNet test set. This result won the 1st place on the ILSVRC 2015 classification task. This technique can also be applied to the CIFAR-10 with 100 and 1000 layers.
#
# ResNet was introduced in the following paper:
#
# * <NAME>, <NAME>, <NAME>, and <NAME>. [Deep residual learning for image recognition](https://arxiv.org/abs/1512.03385). arXiv preprint arXiv:1512.03385,2015.
#
# What is a residual?
#
# * [Residual](https://www.merriam-webster.com/dictionary/residual): an internal aftereffect of experience or activity that influences later behavior
#
# To implement a ResNet we need to give Keras the notion of a residual block. This is essentially two dense layers with a "skip connection" (or residual connection). A residual block is shown in Figure 6.SKIP.
#
#
# **Figure 6.SKIP: Skip Layers**
# 
#
# Residual blocks are typically used with convolutional neural networks (CNNs). This allows very deep neural networks of CNNs to be created. Figure 6.RES shows several different ResNets.
#
# **Figure 6.RES: ResNets**
# 
#
#
# ### Keras Sequence vs Functional Model API
#
# Most of the neural networks create in this course have made use of the Keras sequence object. You might have noticed that we briefly made use of another type of neural network object for the ResNet, the Model. These are the [two major means](https://keras.io/getting-started/functional-api-guide/) of constructing a neural network in Keras:
#
# * [Sequential](https://keras.io/getting-started/sequential-model-guide/) - Simplified interface to Keras that supports most models where the flow of information is a simple sequence from input to output.
# * [Keras Functional API](https://keras.io/getting-started/functional-api-guide/) - More complex interface that allows neural networks to be constructed of reused layers, multiple input layers, and supports building your own recurrent connections.
#
# It is important to point out that these are not two specific types of neural network. Rather, they are two means of constructing neural networks in Keras. Some types of neural network can be implemented in either, such as dense feedforward neural networks (like we used for the Iris and MPG datasets). However, other types of neural network, like ResNet and GANs can only be used in the Functional Model API.
#
# ### CIFAR Dataset
#
# The [CIFAR-10 and CIFAR-100](https://www.cs.toronto.edu/~kriz/cifar.html) datasets are also frequently used by the neural network research community. These datasets were originally part of a competition.
#
# The CIFAR-10 data set contains low-res images that are divided into 10 classes. The CIFAR-100 data set contains 100 classes in a hierarchy.
#
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="gx-DMPysmp9N" outputId="787950ea-00e9-4590-8d61-480d5656f581"
from __future__ import print_function
import tensorflow.keras
from tensorflow.keras.layers import Dense, Conv2D
from tensorflow.keras.layers import BatchNormalization, Activation
from tensorflow.keras.layers import AveragePooling2D, Input, Flatten
from tensorflow.keras.optimizers import Adam
from tensorflow.keras.callbacks import ModelCheckpoint
from tensorflow.keras.callbacks import LearningRateScheduler
from tensorflow.keras.callbacks import ReduceLROnPlateau
from tensorflow.keras.preprocessing.image import ImageDataGenerator
from tensorflow.keras.regularizers import l2
from tensorflow.keras import backend as K
from tensorflow.keras.models import Model
from tensorflow.keras.datasets import cifar10
import numpy as np
import os
# Load the CIFAR10 data.
(x_train, y_train), (x_test, y_test) = cifar10.load_data()
# + [markdown] colab_type="text" id="372-21BTmp9P"
# Samples from the loaded CIFAR dataset can be displayed using the following code.
# + colab={"base_uri": "https://localhost:8080/", "height": 574} colab_type="code" id="ZyJdYNTDmp9P" outputId="179831b7-a7fc-40c8-8020-520ebdd6bda7"
# %matplotlib inline
import numpy as np
import matplotlib.pyplot as plt
from six.moves import cPickle
ROWS = 10
x = x_train.astype("uint8")
fig, axes1 = plt.subplots(ROWS,ROWS,figsize=(10,10))
for j in range(ROWS):
for k in range(ROWS):
i = np.random.choice(range(len(x)))
axes1[j][k].set_axis_off()
axes1[j][k].imshow(x[i:i+1][0])
# + [markdown] colab_type="text" id="AKQDnk2Gmp9R"
# We will construct a ResNet and train it on the CIFAR-10 dataset. The following block of code defines some constant values that define how the network is constructed.
# + colab={} colab_type="code" id="E60qRcFzmp9R"
# Training parameters
BATCH_SIZE = 32 # orig paper trained all networks with batch_size=128
EPOCHS = 200 # 200
USE_AUGMENTATION = True
NUM_CLASSES = np.unique(y_train).shape[0] # 10
COLORS = x_train.shape[3]
# Subtracting pixel mean improves accuracy
SUBTRACT_PIXEL_MEAN = True
# Model version
# Orig paper: version = 1 (ResNet v1),
# Improved ResNet: version = 2
# (ResNet v2)
VERSION = 1
# Computed depth from supplied model parameter n
if VERSION == 1:
DEPTH = COLORS * 6 + 2
elif version == 2:
DEPTH = COLORS * 9 + 2
# + [markdown] colab_type="text" id="I91EXi2hmp9T"
# The following function implements a learning rate decay schedule.
# + colab={} colab_type="code" id="pHUm2jSUmp9T"
def lr_schedule(epoch):
"""Learning Rate Schedule
Learning rate is scheduled to be reduced after 80, 120, 160, 180 epochs.
Called automatically every epoch as part of callbacks during training.
# Arguments
epoch (int): The number of epochs
# Returns
lr (float32): learning rate
"""
lr = 1e-3
if epoch > 180:
lr *= 0.5e-3
elif epoch > 160:
lr *= 1e-3
elif epoch > 120:
lr *= 1e-2
elif epoch > 80:
lr *= 1e-1
print('Learning rate: ', lr)
return lr
# + [markdown] colab_type="text" id="7-du9R2rmp9V"
# The following code implements a ResNet block. This includes two convolutional layers with a skip connection. Both V1 and V2 of ResNet make use of this type of layer.
# + colab={} colab_type="code" id="DPXtreZ6mp9V"
def resnet_layer(inputs,
num_filters=16,
kernel_size=3,
strides=1,
activation='relu',
batch_normalization=True,
conv_first=True):
"""2D Convolution-Batch Normalization-Activation stack builder
# Arguments
inputs (tensor): input tensor from input image or previous layer
num_filters (int): Conv2D number of filters
kernel_size (int): Conv2D square kernel dimensions
strides (int): Conv2D square stride dimensions
activation (string): activation name
batch_normalization (bool): whether to include batch normalization
conv_first (bool): conv-bn-activation (True) or
bn-activation-conv (False)
# Returns
x (tensor): tensor as input to the next layer
"""
conv = Conv2D(num_filters,
kernel_size=kernel_size,
strides=strides,
padding='same',
kernel_initializer='he_normal',
kernel_regularizer=l2(1e-4))
x = inputs
if conv_first:
x = conv(x)
if batch_normalization:
x = BatchNormalization()(x)
if activation is not None:
x = Activation(activation)(x)
else:
if batch_normalization:
x = BatchNormalization()(x)
if activation is not None:
x = Activation(activation)(x)
x = conv(x)
return x
# + [markdown] colab_type="text" id="FDk1ekrxmp9X"
# ### ResNet V1
#
# * <NAME>, <NAME>, <NAME>, and <NAME>. [Deep residual learning for image recognition](https://arxiv.org/abs/1512.03385). arXiv preprint arXiv:1512.03385,2015.
# + colab={} colab_type="code" id="YLQe8BE-mp9X"
def resnet_v1(input_shape, depth, num_classes=10):
"""ResNet Version 1 Model builder [a]
Stacks of 2 x (3 x 3) Conv2D-BN-ReLU
Last ReLU is after the shortcut connection.
At the beginning of each stage, the feature
map size is halved (downsampled)
by a convolutional layer with strides=2, while the number of
filters is
doubled. Within each stage, the layers have the same number
filters and the same number of filters.
Features maps sizes:
stage 0: 32x32, 16
stage 1: 16x16, 32
stage 2: 8x8, 64
The Number of parameters is approx the same as Table 6 of [a]:
ResNet20 0.27M
ResNet32 0.46M
ResNet44 0.66M
ResNet56 0.85M
ResNet110 1.7M
# Arguments
input_shape (tensor): shape of input image tensor
depth (int): number of core convolutional layers
num_classes (int): number of classes (CIFAR10 has 10)
# Returns
model (Model): Keras model instance
"""
if (depth - 2) % 6 != 0:
raise ValueError('depth should be 6n+2 (eg 20, 32, 44 in [a])')
# Start model definition.
num_filters = 16
num_res_blocks = int((depth - 2) / 6)
inputs = Input(shape=input_shape)
x = resnet_layer(inputs=inputs)
# Instantiate the stack of residual units
for stack in range(3):
for res_block in range(num_res_blocks):
strides = 1
# first layer but not first stack
if stack > 0 and res_block == 0:
strides = 2 # downsample
y = resnet_layer(inputs=x,
num_filters=num_filters,
strides=strides)
y = resnet_layer(inputs=y,
num_filters=num_filters,
activation=None)
# first layer but not first stack
if stack > 0 and res_block == 0:
# linear projection residual shortcut connection to match
# changed dims
x = resnet_layer(inputs=x,
num_filters=num_filters,
kernel_size=1,
strides=strides,
activation=None,
batch_normalization=False)
x = tensorflow.keras.layers.add([x, y])
x = Activation('relu')(x)
num_filters *= 2
# Add classifier on top.
# v1 does not use BN after last shortcut connection-ReLU
x = AveragePooling2D(pool_size=8)(x)
y = Flatten()(x)
outputs = Dense(num_classes,
activation='softmax',
kernel_initializer='he_normal')(y)
# Instantiate model.
model = Model(inputs=inputs, outputs=outputs)
return model
# + [markdown] colab_type="text" id="SzNJPXncmp9a"
# ### ResNet V2
#
# A second version of ResNet was introduced in the following paper. This form of ResNet is commonly referred to as ResNet V2.
#
# * <NAME>., <NAME>., <NAME>., & <NAME>. (2016, October). [Identity mappings in deep residual networks](https://arxiv.org/abs/1603.05027). In European conference on computer vision (pp. 630-645). Springer, Cham.
#
# The following code constructs a ResNet V2 network. The primary difference of the full preactivation 'v2' variant compared to the 'v1' variant is the use of [batch normalization](https://arxiv.org/abs/1502.03167) before every weight layer.
# + colab={} colab_type="code" id="8WfihOPhmp9a"
def resnet_v2(input_shape, depth, num_classes=10):
"""ResNet Version 2 Model builder [b]
Stacks of (1 x 1)-(3 x 3)-(1 x 1) BN-ReLU-Conv2D or also known as
bottleneck layer
First shortcut connection per layer is 1 x 1 Conv2D.
Second and onwards shortcut connection is identity.
At the beginning of each stage, the feature map size is
halved (downsampled) by a convolutional layer with
strides=2, while the number of filter maps is
doubled. Within each stage, the layers have the same
number filters and the same filter map sizes.
Features maps sizes:
conv1 : 32x32, 16
stage 0: 32x32, 64
stage 1: 16x16, 128
stage 2: 8x8, 256
# Arguments
input_shape (tensor): shape of input image tensor
depth (int): number of core convolutional layers
num_classes (int): number of classes (CIFAR10 has 10)
# Returns
model (Model): Keras model instance
"""
if (depth - 2) % 9 != 0:
raise ValueError('depth should be 9n+2 (eg 56 or 110 in [b])')
# Start model definition.
num_filters_in = 16
num_res_blocks = int((depth - 2) / 9)
inputs = Input(shape=input_shape)
# v2 performs Conv2D with BN-ReLU on input before splitting into 2 paths
x = resnet_layer(inputs=inputs,
num_filters=num_filters_in,
conv_first=True)
# Instantiate the stack of residual units
for stage in range(3):
for res_block in range(num_res_blocks):
activation = 'relu'
batch_normalization = True
strides = 1
if stage == 0:
num_filters_out = num_filters_in * 4
if res_block == 0: # first layer and first stage
activation = None
batch_normalization = False
else:
num_filters_out = num_filters_in * 2
if res_block == 0: # first layer but not first stage
strides = 2 # downsample
# bottleneck residual unit
y = resnet_layer(inputs=x,
num_filters=num_filters_in,
kernel_size=1,
strides=strides,
activation=activation,
batch_normalization=batch_normalization,
conv_first=False)
y = resnet_layer(inputs=y,
num_filters=num_filters_in,
conv_first=False)
y = resnet_layer(inputs=y,
num_filters=num_filters_out,
kernel_size=1,
conv_first=False)
if res_block == 0:
# linear projection residual shortcut connection to match
# changed dims
x = resnet_layer(inputs=x,
num_filters=num_filters_out,
kernel_size=1,
strides=strides,
activation=None,
batch_normalization=False)
x = tensorflow.keras.layers.add([x, y])
num_filters_in = num_filters_out
# Add classifier on top.
# v2 has BN-ReLU before Pooling
x = BatchNormalization()(x)
x = Activation('relu')(x)
x = AveragePooling2D(pool_size=8)(x)
y = Flatten()(x)
outputs = Dense(num_classes,
activation='softmax',
kernel_initializer='he_normal')(y)
# Instantiate model.
model = Model(inputs=inputs, outputs=outputs)
return model
# + [markdown] colab_type="text" id="-CzYSlxSmp9c"
# With all of this defined, we can run the ResNet.
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="Jg7cCBZnmp9c" outputId="0a911f28-a64b-4989-c803-a1cc88a56cb1"
# Input image dimensions.
input_shape = x_train.shape[1:]
# Normalize data.
x_train = x_train.astype('float32') / 255
x_test = x_test.astype('float32') / 255
# If subtract pixel mean is enabled
if SUBTRACT_PIXEL_MEAN:
x_train_mean = np.mean(x_train, axis=0)
x_train -= x_train_mean
x_test -= x_train_mean
print('x_train shape:', x_train.shape)
print(x_train.shape[0], 'train samples')
print(x_test.shape[0], 'test samples')
print('y_train shape:', y_train.shape)
# Convert class vectors to binary class matrices.
y_train = tensorflow.keras.utils.to_categorical(y_train, NUM_CLASSES)
y_test = tensorflow.keras.utils.to_categorical(y_test, NUM_CLASSES)
# Create the neural network
if VERSION == 2:
model = resnet_v2(input_shape=input_shape, depth=DEPTH)
else:
model = resnet_v1(input_shape=input_shape, depth=DEPTH)
model.compile(loss='categorical_crossentropy',
optimizer=Adam(lr=lr_schedule(0)),
metrics=['accuracy'])
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} colab_type="code" id="CC1x1NIvmp9f" outputId="0ab4b67e-2eb0-40e9-c3b8-34d09c19ccd8"
import time
start_time = time.time()
# Prepare callbacks for model saving and for learning rate adjustment.
lr_scheduler = LearningRateScheduler(lr_schedule)
lr_reducer = ReduceLROnPlateau(factor=np.sqrt(0.1),
cooldown=0,
patience=5,
min_lr=0.5e-6)
callbacks = [lr_reducer, lr_scheduler]
# Run training, with or without data augmentation.
if not USE_AUGMENTATION:
print('Not using data augmentation.')
model.fit(x_train, y_train,
batch_size=BATCH_SIZE,
epochs=EPOCHS,
validation_data=(x_test, y_test),
shuffle=True,
callbacks=callbacks)
else:
print('Using real-time data augmentation.')
# This will do preprocessing and realtime data augmentation:
datagen = ImageDataGenerator(
# set input mean to 0 over the dataset
featurewise_center=False,
# set each sample mean to 0
samplewise_center=False,
# divide inputs by std of dataset
featurewise_std_normalization=False,
# divide each input by its std
samplewise_std_normalization=False,
# apply ZCA whitening
zca_whitening=False,
# epsilon for ZCA whitening
zca_epsilon=1e-06,
# randomly rotate images in the range (deg 0 to 180)
rotation_range=0,
# randomly shift images horizontally
width_shift_range=0.1,
# randomly shift images vertically
height_shift_range=0.1,
# set range for random shear
shear_range=0.,
# set range for random zoom
zoom_range=0.,
# set range for random channel shifts
channel_shift_range=0.,
# set mode for filling points outside the input boundaries
fill_mode='nearest',
# value used for fill_mode = "constant"
cval=0.,
# randomly flip images
horizontal_flip=True,
# randomly flip images
vertical_flip=False,
# set rescaling factor (applied before any other transformation)
rescale=None,
# set function that will be applied on each input
preprocessing_function=None,
# image data format, either "channels_first" or "channels_last"
data_format=None,
# fraction of images reserved for validation
# (strictly between 0 and 1)
validation_split=0.0)
# Compute quantities required for featurewise normalization
# (std, mean, and principal components if ZCA whitening is applied).
datagen.fit(x_train)
# Fit the model on the batches generated by datagen.flow().
model.fit_generator(datagen.flow(x_train, y_train,
batch_size=BATCH_SIZE),
validation_data=(x_test, y_test),
epochs=EPOCHS, verbose=0, workers=1,
callbacks=callbacks,
use_multiprocessing=False)
elapsed_time = time.time() - start_time
print("Elapsed time: {}".format(hms_string(elapsed_time)))
# + [markdown] colab_type="text" id="SpndV2K_mp9g"
# The trained neural network can now be evaluated.
# + colab={"base_uri": "https://localhost:8080/", "height": 88} colab_type="code" id="Rcllz7JOmp9h" outputId="8cb6283d-3627-4532-9fa1-ad7ff7392698"
# Score trained model.
scores = model.evaluate(x_test, y_test, verbose=1)
print('Test loss:', scores[0])
print('Test accuracy:', scores[1])
|
t81_558_class_06_3_resnet.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <a id="introduction"></a>
# ## Introduction to Dask
# #### By <NAME>
# -------
#
# In this notebook, we will show how to get started with Dask using basic Python primitives like integers and strings.
#
# **Table of Contents**
#
# * [Introduction to Dask](#introduction)
# * [Setup](#setup)
# * [Introduction to Dask](#dask)
# * [Conclusion](#conclusion)
# <a id="setup"></a>
# ## Setup
#
# This notebook was tested using the following Docker containers:
#
# * `rapidsai/rapidsai:0.6-cuda10.0-devel-ubuntu18.04-gcc7-py3.7` from [DockerHub](https://hub.docker.com/r/rapidsai/rapidsai)
# * `rapidsai/rapidsai-nightly:0.6-cuda10.0-devel-ubuntu18.04-gcc7-py3.7` from [DockerHub](https://hub.docker.com/r/rapidsai/rapidsai-nightly)
#
# This notebook was run on the NVIDIA Tesla V100 GPU. Please be aware that your system may be different and you may need to modify the code or install packages to run the below examples.
#
# If you think you have found a bug or an error, please file an issue here: https://github.com/rapidsai/notebooks/issues
#
# Before we begin, let's check out our hardware setup by running the `nvidia-smi` command.
# !nvidia-smi
# Next, let's see what CUDA version we have:
# !nvcc --version
# !apt update
# !apt install -y graphviz
# !conda install graphviz
# !conda install python-graphviz
# <a id="dask"></a>
# ## Introduction to Dask
#
# Dask is a library that allows for parallelized computing. Written in Python, it allows one to compose complex workflows using large data structures like those found in NumPy, Pandas, and cuDF. In the following examples and notebooks, we'll show how to use Dask with cuDF to accelerate common ETL tasks as well as build and train machine learning models like Linear Regression and XGBoost.
#
# To learn more about Dask, check out the documentation here: http://docs.dask.org/en/latest/
#
# #### Client/Workers
#
# Dask operates by creating a cluster composed of a "client" and multiple "workers". The client is responsible for scheduling work; the workers are responsible for actually executing that work.
#
# Typically, we set the number of workers to be equal to the number of computing resources we have available to us. For CPU based workflows, this might be the number of cores or threads on that particlular machine. For example, we might set `n_workers = 8` if we have 8 CPU cores or threads on our machine that can each operate in parallel. This allows us to take advantage of all of our computing resources and enjoy the most benefits from parallelization.
#
# On a system with one or more GPUs, we usually set the number of workers equal to the number of GPUs available to us. Dask is a first class citizen in the world of General Purpose GPU computing and the RAPIDS ecosystem makes it very easy to use Dask with cuDF and XGBoost.
#
# Before we get started with Dask, we need to setup a Local Cluster of workers to execute our work and a Client to coordinate and schedule work for that cluster. As we see below, we can inititate a `cluster` and `client` using only few lines of code.
# +
import dask; print('Dask Version:', dask.__version__)
from dask.distributed import Client, LocalCluster
import subprocess
# parse the hostname IP address
cmd = "hostname --all-ip-addresses"
process = subprocess.Popen(cmd.split(), stdout=subprocess.PIPE)
output, error = process.communicate()
ip_address = str(output.decode()).split()[0]
# create a local cluster with 4 workers
n_workers = 4
cluster = LocalCluster(ip=ip_address, n_workers=n_workers)
client = Client(cluster)
# -
# Let's inspect the `client` object to view our current Dask status. We should see the IP Address for our Scheduler as well as the the number of workers in our Cluster.
# show current Dask status
client
# You can also see the status and more information at the Dashboard, found at `http://<ip_address>/status`. You can ignore this for now, we'll dive into this in subsequent tutorials.
#
# With our client and workers setup, it's time to execute our first program in parallel. We'll define a function that takes some value `x` and adds 5 to it.
def add_5_to_x(x):
return x + 5
# Next, we'll iterate through our `n_workers` and create an execution graph, where each worker is responsible for taking its ID and passing it to the function `add_5_to_x`. For example, the worker with ID 2 will take its ID and add 5, resulting in the value 7.
# +
from dask import delayed
addition_operations = [delayed(add_5_to_x)(i) for i in range(n_workers)]
addition_operations
# -
# The above output shows a list of several `Delayed` objects. An important thing to note is that the workers aren't actually executing these results - we're just defining the execution graph for our client to execute later. The `delayed` function wraps our function `add_5_to_x` and returns a `Delayed` object. This ensures that this computation is in fact "delayed" - or lazily evaluated - and not executed on the spot i.e. when we define it.
#
# Next, let's sum each one of these intermediate results. We can accomplish this by wrapping Python's built-in `sum` function using our `delayed` function and storing this in a variable called `total`.
total = delayed(sum)(addition_operations)
total
# Using the `graphviz` library, we can use the `visualize` method of a `Delayed` object to visualize our current graph.
total.visualize()
# As we mentioned before, none of these results - intermediate or final - have actually been compute. We can compute them using the `compute` method of our `client`.
# +
import time
addition_futures = client.compute(addition_operations, optimize_graph=False, fifo_timeout="0ms")
total_future = client.compute(total, optimize_graph=False, fifo_timeout="0ms")
time.sleep(1) # this will give Dask time to execute each worker
# -
# Let's inspect the output of each call to `client.compute`:
addition_futures
# We can see from the above output that our `addition_futures` variable is a list of `Future` objects - not the "actual results" of adding 5 to each of `[0, 1, 2, 3]`. These `Future` objects are a promise that at one point a computation will take place and we will be left with a result. Dask is responsible for ensuring that promise by delegating that task to the appropriate Dask worker and collecting the result.
#
# Let's take a look at our `total_future` object:
print(total_future)
print(type(total_future))
# Again, we see that this is an object of type `Future` as well as metadata about the status of the request (i.e. whether it has finished or not), the type of the result, and a key associated with that operation. To collect and print the result of each of these `Future` objects, we can call the `result()` method.
addition_results = [future.result() for future in addition_futures]
print('Addition Results:', addition_results)
# Now we see the results that we want from our addition operations. We can also use the simpler syntax of the `client.gather` method to collect our results.
addition_results = client.gather(addition_futures)
total_result = client.gather(total_future)
print('Addition Results:', addition_results)
print('Total Result:', total_result)
# Awesome! We just wrote our first distributed workflow.
#
# To confirm that Dask is truly executing in parallel, let's define a function that sleeps for 1 second and returns the string "Success!". In serial, this function should take our 4 workers around 4 seconds to execute.
def sleep_1():
time.sleep(1)
return 'Success!'
# +
# %%time
for _ in range(n_workers):
sleep_1()
# -
# As expected, our process takes about 4 seconds to run. Now let's execute this same workflow in parallel using Dask.
# +
# %%time
# define delayed execution graph
sleep_operations = [delayed(sleep_1)() for _ in range(n_workers)]
# use client to perform computations using execution graph
sleep_futures = client.compute(sleep_operations, optimize_graph=False, fifo_timeout="0ms")
# collect and print results
sleep_results = client.gather(sleep_futures)
print(sleep_results)
# -
# Using Dask, we see that this whole process takes a little over a second - each worker is executing in parallel!
# <a id="conclusion"></a>
# ## Conclusion
#
# In this tutorial, we learned how to use Dask with basic Python primitives like integers and strings.
#
# To learn more about RAPIDS, be sure to check out:
#
# * [Open Source Website](http://rapids.ai)
# * [GitHub](https://github.com/rapidsai/)
# * [Press Release](https://nvidianews.nvidia.com/news/nvidia-introduces-rapids-open-source-gpu-acceleration-platform-for-large-scale-data-analytics-and-machine-learning)
# * [NVIDIA Blog](https://blogs.nvidia.com/blog/2018/10/10/rapids-data-science-open-source-community/)
# * [Developer Blog](https://devblogs.nvidia.com/gpu-accelerated-analytics-rapids/)
# * [NVIDIA Data Science Webpage](https://www.nvidia.com/en-us/deep-learning-ai/solutions/data-science/)
|
getting_started_notebooks/basics/Getting_Started_with_Dask.ipynb
|
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# formats: ipynb,py:light
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # To determine the classification of Game results using Decision trees & Random Forests
import numpy as npL
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
# %matplotlib inline
df = pd.read_csv('../../data/processed/LOLOracleData_ChampStats.csv')
df.head()
tagcols = ['BToptags','BJngtags','BMidtags','BAdctags','BSuptags','RToptags','RJngtags','RMidtags','RAdctags','RSuptags']
tag_dummies = pd.get_dummies(df[tagcols])
tag_dummies.head(2)
df2 = pd.concat([df,tag_dummies],axis=1)
df2.head(2)
df2.drop(tagcols,axis=1,inplace=True)
df2.head(2)
# ### Lets perform Decision Trees and Random Forests
X = df2.drop('Winner',axis=1)
y = df2['Winner']
X.head(2)
X.shape
from sklearn.model_selection import train_test_split
X_train,X_test,y_train,y_test = train_test_split(X,y,test_size=0.3,random_state=101)
from sklearn.tree import DecisionTreeClassifier
dtc = DecisionTreeClassifier()
dtc.fit(X_train,y_train)
predict_dtc = dtc.predict(X_test)
from sklearn.metrics import classification_report,confusion_matrix
print (confusion_matrix(y_test,predict_dtc))
print('\n')
print(classification_report(y_test,predict_dtc))
from sklearn.ensemble import RandomForestClassifier
rfc = RandomForestClassifier(n_estimators=300)
rfc.fit(X_train,y_train)
predict_rfc = rfc.predict(X_test)
print (confusion_matrix(y_test,predict_rfc))
print('\n')
print(classification_report(y_test,predict_rfc))
|
models/level2/DTRFMethod-Complex.ipynb
|
# <img src="data:image/svg+xml;base64,PHN2ZyBpZD0iTGF5ZXJfMSIgeG1sbnM9Imh0dHA6Ly93d3cudzMub3JnLzIwMDAvc3ZnIiB4bWxuczp4bGluaz0iaHR0cDovL3d3dy53My5vcmcvMTk5OS94bGluayIgeD0iMHB4IiB5PSIwcHgiDQoJIHZpZXdCb3g9Ii0yMjYgMzc3LjEgMTU4LjUgMzkuOSIgc3R5bGU9ImVuYWJsZS1iYWNrZ3JvdW5kOm5ldyAtMjI2IDM3Ny4xIDE1OC41IDM5Ljk7IiB4bWw6c3BhY2U9InByZXNlcnZlIj4NCgkgPHRpdGxlPkZ1bGwgU3RpdGNoIExvZ288L3RpdGxlPg0KCTxnIGlkPSJ3b3JkbWFyayI+DQoJCTxwYXRoIGQ9Ik0tMTU2LjcsMzkwLjFsLTAuNC0zLjljMCwwLTIuMS0wLjgtNC4zLTAuOGMtNCwwLTUuOSwxLjctNS45LDQuNWMwLDMuMiwyLjgsNC4xLDUuOSw1LjENCgkJCWMzLjksMS4yLDguMiwyLjUsOC4yLDguMmMwLDYtNC43LDguMS0xMC4yLDguMWMtMy41LDAtNi45LTAuOS04LjEtMS41di02LjNoMi42bDAuNSw0LjFjMC42LDAuMiwzLjEsMC44LDUuMywwLjgNCgkJCWMzLjksMCw2LjMtMS41LDYuMy00LjljMC0zLjQtMi43LTQuNC01LjctNS4zYy0zLjktMS4yLTguNC0yLjItOC40LThjMC01LjUsNC4zLTcuOSw5LjctNy45YzMuMiwwLDUuOSwwLjgsNy4yLDEuM3Y2LjUNCgkJCUwtMTU2LjcsMzkwLjF6Ii8+DQoJCTxwYXRoIGQ9Ik0tMTEwLjIsNDEwLjFjLTEuNSwwLjYtMy40LDEuMi01LjQsMS4yYy0zLjcsMC01LjQtMS43LTUuNC01LjJ2LTEyYzAtMC4yLDAtMC40LTAuMy0wLjRoLTIuNHYtMi4yDQoJCQljMy41LDAsMy45LTQuNywzLjktNC43aDIuM3YzLjljMCwwLjIsMCwwLjQsMC4zLDAuNGg1Ljd2Mi43aC02djExLjRjMCwyLjQsMC42LDMuNCwyLjgsMy40YzEuMiwwLDIuNS0wLjMsMy42LTAuOEwtMTEwLjIsNDEwLjF6Ii8+DQoJCTxwYXRoIGQ9Ik0tMTM3LjYsNDEwLjFjLTEuNSwwLjYtMy40LDEuMi01LjQsMS4yYy0zLjcsMC01LjQtMS43LTUuNC01LjJ2LTEyYzAtMC4yLDAtMC40LTAuMy0wLjRoLTIuNHYtMi4yDQoJCQljMy41LDAsMy45LTQuNywzLjktNC43aDIuM3YzLjljMCwwLjIsMCwwLjQsMC4zLDAuNGg1Ljd2Mi43aC02djExLjRjMCwyLjQsMC42LDMuNCwyLjgsMy40YzEuMiwwLDIuNS0wLjMsMy42LTAuOEwtMTM3LjYsNDEwLjF6Ii8+DQoJCTxwYXRoIGQ9Ik0tMTMwLjQsMzg2LjhjLTEuNiwwLTIuNS0wLjktMi41LTIuNWMwLTEuNSwwLjktMi40LDIuNS0yLjRjMS42LDAsMi41LDAuOSwyLjUsMi40DQoJCQlDLTEyOCwzODUuOS0xMjguOCwzODYuOC0xMzAuNCwzODYuOCBNLTEzNC44LDQxMC44di0yLjFsMy4yLTAuM3YtMTQuOWwtMi43LTAuNGwwLjMtMmg1Ljl2MTcuNGwzLjEsMC4zdjIuMUgtMTM0Ljh6Ii8+DQoJCTxwYXRoIGQ9Ik0tOTIuMyw0MDkuN2MtMS41LDEtNCwxLjYtNi42LDEuNmMtNi42LDAtOS40LTQtOS40LTEwLjFjMC03LjMsNC4yLTEwLjUsMTAuMS0xMC41DQoJCQljMi44LDAsNSwwLjgsNS45LDEuMnY1LjFsLTIuNi0wLjFsLTAuNC0zLjFjLTAuNS0wLjItMi0wLjMtMy4xLTAuM2MtMy40LDAtNi4yLDItNi4yLDcuNWMwLDUuNCwyLjUsNy41LDYuMyw3LjUNCgkJCWMxLjksMCwzLjktMC42LDQuOS0xLjNMLTkyLjMsNDA5Ljd6Ii8+DQoJCTxwYXRoIGQ9Ik0tNzYuOCw0MTAuOHYtMi4xbDIuNS0wLjJ2LTkuOGMwLTIuOS0wLjUtNS4xLTMuMi01LjFjLTIuMywwLTQuNiwxLjMtNi4xLDIuNXYxMi40bDIuOCwwLjJ2Mi4xaC04LjgNCgkJCXYtMi4xbDIuNS0wLjJ2LTI0LjNsLTIuOC0wLjRsMC4zLTJoNS45djExLjRjMS44LTEuMyw0LTIuNSw2LjktMi41YzQuMSwwLDUuOSwyLjcsNS45LDd2MTAuOWwyLjgsMC4ydjIuMUgtNzYuOHoiLz4NCgk8L2c+DQoJPHBhdGggaWQ9ImxvZ29tYXJrIiBkPSJNLTIxMy4zLDM5NS43di0yLjhoMjcuMnYtMTUuOEgtMjI2djIxLjRoMjguNHYyLjhILTIyNlY0MTdoMzkuOXYtMjEuNEgtMjEzLjN6IE0tMjA1LjcsMzgxLjINCgkJYzEuMSwwLDIsMC45LDIsMmMwLDEuMS0wLjksMi0yLDJjLTEuMSwwLTItMC45LTItMkMtMjA3LjYsMzgyLjEtMjA2LjcsMzgxLjItMjA1LjcsMzgxLjJ6IE0tMjE0LjIsMzg3LjNoMTcuN2wtMC44LDIuOGgtMTYuMQ0KCQlMLTIxNC4yLDM4Ny4zeiBNLTIwNS43LDQxMi45Yy0xLjEsMC0yLTAuOS0yLTJjMC0xLjEsMC45LTIsMi0yYzEuMSwwLDIsMC45LDIsMkMtMjAzLjcsNDEyLTIwNC42LDQxMi45LTIwNS43LDQxMi45eiBNLTIxNC4yLDQwNi44DQoJCWwwLjctMi44aDE2LjJsMC44LDIuOEgtMjE0LjJ6Ii8+DQo8L3N2Zz4NCg==" width=200 style="float:left" />
# <br />
# <br />
#
# <hr />
#
# ## Your variables
#
# `stitch_context` is an object exposing access to connections and table load information from your Stitch account.
#
# `stitch_context.connections` references the integrations and destination currently configured in Stitch that are accessible by Scripts.
#
# ```python
# # explore the list of Stitch connections available to Scripts
# [name for name in stitch_context.connections]
# ```
#
# Connect to the destination you configured in Stitch:
#
# ```python
# # run a query directly on your data warehouse
# import psycopg2.extras
# warehouse = stitch_context.connections['Default Warehouse']['client']
# cur = warehouse.cursor(cursor_factory=psycopg2.extras.RealDictCursor)
# query = 'SELECT NOW()'
# cur.execute(query)
# results = cur.fetchall()
# print(results[0]['now'])
# ```
#
# `stitch_context.tables_loaded` is an object containing the names of tables Stitch loaded since the last invocation of this Script. Here, in the interactive notebook context, the `stitch_context.tables_loaded` object contains dummy data to illustrate the structure. When the Script is executed on a schedule, `stitch_context.tables_loaded` will contain the actual integrations and tables that were loaded. Note: this variable is only relevant for post-load Scripts.
#
# ```python
# {'integration_0': ['table_0', 'table_1'],
# 'integration_1': ['table_0', 'table_1', 'table_2']}
# ```
#
# ## Available Python packages
#
#
# ### Tools
#
# - [`requests`](http://docs.python-requests.org/en/master/) - simple HTTP library
# - [`pandas`](http://pandas.pydata.org/) - data analysis
# - [`numpy`](http://www.numpy.org/) - numerical / technical computing
# - [`scipy`](https://www.scipy.org/about.html) - scientific computing
# - [`scikit-learn`](http://scikit-learn.org/stable/index.html) - machine learning
# - [`matplotlib`](https://matplotlib.org/) - 2D plotting
# - [`snowflake-sqlalchemy`](https://github.com/snowflakedb/snowflake-sqlalchemy) - [SQLAlchemy ORM](http://www.sqlalchemy.org/) with Snowflake dialect
# - [`lifetimes`](https://github.com/CamDavidsonPilon/lifetimes) - customer lifetime value
#
#
# ### Connection clients
#
# - [`psycopg2`](http://initd.org/psycopg/) - Redshift, PostgreSQL
# - [`pymysql`](https://pymysql.readthedocs.io/en/latest/) - MySQL
# - [`google-cloud-bigquery`](https://github.com/GoogleCloudPlatform/google-cloud-python) - BigQuery
# - [`snowflake`](https://github.com/snowflakedb/snowflake-connector-python) - Snowflake
# - [`marketorestpython`](https://github.com/jepcastelein/marketo-rest-python) - Marketo
# - [`simple-salesforce`](https://github.com/simple-salesforce/simple-salesforce) - Salesforce.com
# - For Autopilot and HubSpot connections, Scripts provide thin wrappers to the [Autopilot API](https://autopilot.docs.apiary.io/) and [HubSpot API](https://developers.hubspot.com/docs/overview)s respectively, e.g.:
#
# ```python
# autopilot = stitch_context.connections['my_autopilot_conn']['client']
# contacts = autopilot.get('/contacts')
# import pprint as pp
# pp.pprint(contacts.json())
# autopilot.post('/contact', json=<contact obj>)
# ```
#
#
# ```python
# hubspot = stitch_context.connections['my_hubspot_conn']['client']
# # Query and body params can be passed directly to the client. See the HubSpot API docs for more info.
# # hubspot.get(url, params={"<param0>": "<value0>"}, json={"<body_param0>": "<value0>"})
# contacts = hubspot.get('https://api.hubapi.com/contacts/v1/lists/all/contacts/all', params={'count': 2})
# ```
#
# ### Logging
#
# Output from Script runs can be displayed in the Stitch Scripts UI via the `log` object. This is useful for debugging and to provide insight into Script execution. Read more about logging [here](https://docs.python.org/3.5/library/logging.html).
#
# ```python
# log.info("This log line will be available in the Stitch Scripts UI")
# ```
|
templates/bare/template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src="../img/logo_white_bkg_small.png" align="right" />
#
# # PyDrill Demonstration
# This notebook demonstrates how to use the PyDrill module to connect to Apache Drill and query data. The complete documentation for PyDrill can be found at http://pydrill.readthedocs.io
#
# The essential steps are:
# 1. Import the module
# 2. Open a connection to Drill
# 3. Execute a query
# 4. Do something with the results.
#
# You will first need to install PyDrill. This can be done by opening a terminal and typing:
# ```python
# pip install pydrill
# ```
# ## Step 1: Import the PyDrill module
# After you've done this, you will be able to import the PyDrill module.
from pydrill.client import PyDrill
import pandas as pd
import matplotlib.pyplot as plt
# %matplotlib inline
plt.style.use('ggplot')
# ## Step 2: Open a connection to Drill
# The next step is to open a connection to Drill. Once you've opened the connection, you will want to verify that the connection was successfully opened before executing any queries. PyDrill includes an `is_active()` method for this purpose.
# +
#Open a connection to Drill
drill = PyDrill(host='localhost', port=8047)
#Verify the connection is active, throw an error if not.
if not drill.is_active():
raise ImproperlyConfigured('Please run Drill first')
# -
# ## Step 3: Execute a query and get the results
# The next and final step is to execute a query in Drill. When you call the `.query()` method, PyDrill returns an iterable object from which you can extract the rows of your results. You can also get PyDrill to return a pandas DataFrame.
# +
#Execute query in Drill
query_result = drill.query('''
SELECT JobTitle,
AVG( TO_NUMBER( AnnualSalary, '¤' )) AS avg_salary,
COUNT( DISTINCT `EmpName` ) AS number
FROM dfs.drillclass.`baltimore_salaries_2016.csvh`
GROUP BY JobTitle
Order BY avg_salary DESC
LIMIT 50
''')
#Iterate through the rows.
for row in query_result:
print( row )
# -
# ### Retrieving a DataFrame
# You can also get PyDrill to directly return a DataFrame by using the `.to_dataframe()` method of the results object.
df = query_result.to_dataframe()
df.head()
# ## In Class Exercise:
# Using the data in the `dailybots.csv` file use Drill to:
# 1. Query the file to produce a summary of infections by day.
# 2. Store this data in a dataframe using the `to_dataframe()` method.
# 3. Create a line plot of this data by calling the .plot() method on the dataframe
#
# In order to render the plot, you will need to convert the column containing the numeric field into a numeric data type. The command below converts a column called `bot_count` to a floating point number which can be rendered in a line chart.
# ```python
# df['host_count'] = df['host_count'].astype(int)
# ```
#
# If you are unfamiliar with pandas and the corresponding plotting libraries, the documentation is available here: http://pandas.pydata.org/pandas-docs/stable/generated/pandas.DataFrame.plot.html.
# +
query_result = drill.query( """SELECT `date`, SUM( CAST(`hosts` AS INT) ) AS host_count
FROM dfs.drillclass.`dailybots.csvh`
GROUP BY `date`
ORDER BY `date`""")
#Get the dataframe
df = query_result.to_dataframe()
#Convert the host count to an integer field
df['host_count'] = df['host_count'].astype(int)
#Sort by the date column
df.sort_values('date', inplace=True, ascending=True)
# -
df.head()
#Plot the data
df.plot(figsize=(14,8))
# ## Building a Report using Drill and Superset
# In this example, we will use a HTTPD logfile and create a report of suspicious activity. The file `hackers-access.httpd` is a web server log.
#
# You can read more about this file here: https://github.com/nielsbasjes/logparser/tree/master/examples/demolog
#
# The path `/join_form` is a path which should be unknown to anyone except people trying to break into the site. Using Drill and Superset, create visualizations which answer the following questions:
# 1. What time of day are people trying to access this page?
# 2. What browsers are they using?
# 3. Where are they coming from?
|
answers/Worksheet 8 - Working with Drill - Answers.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
from decision_tree.decision_tree_model import ClassificationTree
class RandomForest():
"""Random Forest classifier. Uses a collection of classification trees that
trains on random subsets of the data using a random subsets of the features.
Parameters:
-----------
n_estimators: int
The number of classification trees that are used.
max_features: int
The maximum number of features that the classification trees are allowed to
use.
min_samples_split: int
The minimum number of samples needed to make a split when building a tree.
min_gain: float
The minimum impurity required to split the tree further.
max_depth: int
The maximum depth of a tree.
"""
def __init__(self, n_estimators=100, min_samples_split=2, min_gain=0,
max_depth=float("inf"), max_features=None):
self.n_estimators = n_estimators
self.min_samples_split = min_samples_split
self.min_gain = min_gain
self.max_depth = max_depth
self.max_features = max_features
self.trees = []
# Bulid forest
for _ in range(self.n_estimators):
tree = ClassificationTree(min_samples_split=self.min_samples_split, min_impurity=self.min_gain,
max_depth=self.max_depth)
self.trees.append(tree)
def fit(self, X, Y):
# Every tree use random data set(bootstrap) and random feature
sub_sets = self.get_bootstrap_data(X, Y)
n_features = X.shape[1]
if self.max_features == None:
self.max_features = int(np.sqrt(n_features))
for i in range(self.n_estimators):
# Get random features
sub_X, sub_Y = sub_sets[i]
idx = np.random.choice(n_features, self.max_features, replace=True)
sub_X = sub_X[:, idx]
self.trees[i].fit(sub_X, sub_Y)
self.trees[i].feature_indices = idx
print("tree", i, "fit complete")
def predict(self, X):
y_preds = []
for i in range(self.n_estimators):
idx = self.trees[i].feature_indices
sub_X = X[:, idx]
y_pre = self.trees[i].predict(sub_X)
y_preds.append(y_pre)
y_preds = np.array(y_preds).T
y_pred = []
for y_p in y_preds:
# np.bincount() counts the frequencies each index appears
# np.argmax()returns the maximun index in the arr
# cheak np.bincount() and np.argmax() in numpy Docs
y_pred.append(np.bincount(y_p.astype('int')).argmax())
return y_pred
def get_bootstrap_data(self, X, Y):
# Get int(n_estimators) data by bootstrap
m = X.shape[0]
Y = Y.reshape(m, 1)
# Combine X and Y for bootstrap
X_Y = np.hstack((X, Y))
np.random.shuffle(X_Y)
data_sets = []
for _ in range(self.n_estimators):
idm = np.random.choice(m, m, replace=True)
bootstrap_X_Y = X_Y[idm, :]
bootstrap_X = bootstrap_X_Y[:, :-1]
bootstrap_Y = bootstrap_X_Y[:, -1:]
data_sets.append([bootstrap_X, bootstrap_Y])
return data_sets
|
Random Forest.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Lecture 1 - September 27, 2021
# # Synthetic Aperture Radar (SAR)
# # Part 1: Range Resolution
# +
# --- To get started, we set the ipympl backend, which makes matplotlib plots interactive.
# --- We do this using a magic command, starting with %.
# %matplotlib widget
# --- import libraries
import sys
sys.path.append('/projects/src/')
import numpy as np
import matplotlib.pyplot as plt
from numpy.fft import fft, fftfreq, fftshift, ifft
# --- import rat class
from ste_io import *
# --- Useful functions
def rect(t, T0):
# Implements y = rect(t/T0) = 1 if abs(t/T0) <= 0.5, 0 otherwise
# All input / outputs are Numpy arrays
N = t.shape[0]
y = np.zeros(N, 'float32')
y[np.abs(t/T0) <= 0.5] = 1
return y
# --- Constants
# speed of light
c0 = 2.9972190e+08
# -
# ## Exercise 1
#
# The typical P-band chirp transmitted by the F-SAR system is characterized by the following input parameters:
# +
# --- Input parameters
# chirp duration, s
T = 1.0016000e-05
# chirp bandwidth, Hz
Wr = 50.0e6
# -
# In the focusing process, the following parameters are chosen for ther ange axis:
# number of samples in range
Ns = 3772
# range sampling, m
rs = 1.1988876
# 1. Calculate and plot the chirp signal;
# 2. Generate the impulse responde of the matched filter;
# 3. Calculate and plot the output (convolution) of the matched filter with the chirp as input.
# ### Solution
# +
# create time axis
range_axis = np.linspace(0, Ns-1, Ns) * rs
range_axis = range_axis - np.max(range_axis)/2
# from range to time
time = range_axis * 2/c0
# generation of chirp
kr = Wr / T
s0_t = np.exp(1j * np.pi * kr * time**2) * rect(time, T)
# plot chirp
plt.figure()
plt.plot(time, np.real(s0_t))
plt.xlabel('Time (s)')
plt.ylabel('Real part')
plt.title('Chirp')
plt.grid()
# generation of matched filter
h0_t = np.conj(s0_t[::-1])
# calculate the output
u0_t = np.convolve(s0_t, h0_t, 'same')
# plot chirp
plt.figure()
plt.plot(time, np.real(u0_t))
plt.xlabel('Time (s)')
plt.ylabel('Real part')
plt.title('Output of matched filter')
plt.grid()
# -
# ## Exercise 2
# The files */projects/data/01-sar/signal1_rc.npy* and */projects/data/01-sar/signal2_rc.npy* contain the received signals along two range lines originated by the same range chirp and sampled with the same parameters as in Exercise 1.
#
# 1. Open the files (use np.load);
# 2. Plot the real part of the signals;
# 3. Focus in range by using a matched filter;
# 4. ... how many scatterers do you see in the two cases? and at which range?
# ### Solution
# ## Exercise 3
# We are ready now to focus real F-SAR P-band data !
#
# 1. Open the raw data image in the file */projects/data/01-sar/raw-img.rat*, and visualize it (only the amplitude).
# 2. Compress it in range using the same chirp as in the Exercise 1, this time using the Fourier transform. What do you see?
#
# Tip:
# Use the provided rat class to open the data - Example: img = rrat(*filename.rat*)
# ### Solution
|
notebooks-us-solution/2021_09_27_Lecture1_SAR_Part1_Mon.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="KdUFcDsdzRyw"
# # Clonamos el repositorio para obtener los dataSet
# + colab={"base_uri": "https://localhost:8080/", "height": 34} colab_type="code" id="mHReFf3_y9ms" outputId="c17545fd-c7dd-42c2-e3ad-4f55db21611f"
# !git clone https://github.com/joanby/machinelearning-az.git
# + [markdown] colab_type="text" id="vNKZXgtKzU2x"
# # Damos acceso a nuestro Drive
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="5gu7KWnzzUQ0" outputId="abe602b4-3a59-470e-d508-037c6966002b"
from google.colab import drive
drive.mount('/content/drive')
# + [markdown] colab_type="text" id="1gUxIkHWzfHV"
# # Test it
# + colab={} colab_type="code" id="mIQt3jBMzYRE"
# !ls '/content/drive/My Drive'
# + [markdown] colab_type="text" id="mHsK36uN0XB-"
# # Google colab tools
# + colab={} colab_type="code" id="kTzwfUPWzrm4"
from google.colab import files # Para manejar los archivos y, por ejemplo, exportar a su navegador
import glob # Para manejar los archivos y, por ejemplo, exportar a su navegador
from google.colab import drive # Montar tu Google drive
# + [markdown] colab_type="text" id="uab9OAbV8hYN"
# # Instalar dependendias
# + colab={"base_uri": "https://localhost:8080/", "height": 122} colab_type="code" id="qukjDgj98kE4" outputId="95b5f2b5-7149-436a-b1fb-ad567cc783bd"
# !pip install sklearn
# + [markdown] colab_type="text" id="3yFpBwmNz70v"
# # Regresión Logística
#
# + [markdown] colab_type="text" id="v8OxSXXSz-OP"
# # Cómo importar las librerías
#
# + colab={} colab_type="code" id="edZX51YLzs59"
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
# + [markdown] colab_type="text" id="8XfXlqtF0B58"
# # Importar el data set
#
# + colab={} colab_type="code" id="-nnozsHsz_-N"
dataset = pd.read_csv('/content/machinelearning-az/datasets/Part 3 - Classification/Section 14 - Logistic Regression/Social_Network_Ads.csv')
X = dataset.iloc[:, [2,3]].values
y = dataset.iloc[:, 4].values
# + [markdown] colab_type="text" id="SsVEdPzf4XmV"
# # Dividir el data set en conjunto de entrenamiento y conjunto de testing
#
# + colab={} colab_type="code" id="v9CtwK834bjy"
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size = 0.25, random_state = 0)
# + [markdown] colab_type="text" id="5AH_uCEz68rb"
# # Escalado de variables
# + colab={"base_uri": "https://localhost:8080/", "height": 54} colab_type="code" id="oeuAy8LI69vi" outputId="10346439-d6ac-4abd-b5bb-033e9a284716"
from sklearn.preprocessing import StandardScaler
sc_X = StandardScaler()
X_train = sc_X.fit_transform(X_train)
X_test = sc_X.transform(X_test)
# -
# # Ajustar el clasificador en el Conjunto de Entrenamiento
# # Crear el modelo de clasificación aquí
# + [markdown] colab_type="text" id="je3kcRlG7JV5"
# # Predicción de los resultados con el Conjunto de Testing
#
# + colab={} colab_type="code" id="HS-M9s587Kj3"
y_pred = classifier.predict(X_test)
# -
# # Elaborar una matriz de confusión
from sklearn.metrics import confusion_matrix
cm = confusion_matrix(y_test, y_pred)
# + [markdown] colab_type="text" id="JnMLSqzW8NH7"
# # Representación gráfica de los resultados del algoritmo en el Conjunto de Entrenamiento
# -
from matplotlib.colors import ListedColormap
# + colab={} colab_type="code" id="1qZ3wRR08Oar"
X_set, y_set = X_train, y_train
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Clasificador (Conjunto de Entrenamiento)')
plt.xlabel('Edad')
plt.ylabel('Sueldo Estimado')
plt.legend()
plt.show()
# -
# # Representación gráfica de los resultados del algoritmo en el Conjunto de Testing
X_set, y_set = X_test, y_test
X1, X2 = np.meshgrid(np.arange(start = X_set[:, 0].min() - 1, stop = X_set[:, 0].max() + 1, step = 0.01),
np.arange(start = X_set[:, 1].min() - 1, stop = X_set[:, 1].max() + 1, step = 0.01))
plt.contourf(X1, X2, classifier.predict(np.array([X1.ravel(), X2.ravel()]).T).reshape(X1.shape),
alpha = 0.75, cmap = ListedColormap(('red', 'green')))
plt.xlim(X1.min(), X1.max())
plt.ylim(X2.min(), X2.max())
for i, j in enumerate(np.unique(y_set)):
plt.scatter(X_set[y_set == j, 0], X_set[y_set == j, 1],
c = ListedColormap(('red', 'green'))(i), label = j)
plt.title('Clasificador (Conjunto de Test)')
plt.xlabel('Edad')
plt.ylabel('Sueldo Estimado')
plt.legend()
plt.show()
|
datasets/Part 3 - Classification/Section 14 - Logistic Regression/classification_template.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import os
import matplotlib.pyplot as plt
import joblib
plt.style.use("fivethirtyeight")
# !pip install joblib
class Perceptron:
def __init__(self, eta, epochs):
self.weights = np.random.randn(3) * 1e-4
self.eta = eta # learning rate
self.epochs = epochs # iterations
def _z_outcome(self, inputs, weights):
return np.dot(inputs, weights)
def activation_function(self, z):
return np.where(z > 0, 1, 0)
def fit(self, X, y):
self.X = X
self.y = y
X_with_bias = np.c_[self.X, -np.ones((len(self.X), 1))]
print(f"X with bias: \n{X_with_bias}")
for epoch in range(self.epochs):
print("--"*10)
print(f"for epoch >> {epoch + 1}")
print("--"*10)
z = self._z_outcome(X_with_bias, self.weights)
y_hat = self.activation_function(z)
print(f"predicted value after forward pass: \n{y_hat}")
self.error = self.y - y_hat
print(f"error: \n{self.error}")
self.weights = self.weights + self.eta * np.dot(X_with_bias.T, self.error)
print(f"updated weights after epoch: {epoch + 1}/{self.epochs}: \n{self.weights}")
print(f"##"*10)
def predict(self, X):
X_with_bias = np.c_[X, -np.ones((len(X), 1))]
z = self._z_outcome(X_with_bias, self.weights)
return self.activation_function(z)
# +
OR={
"x1":[0,0,1,1],
"x2":[0,1,0,0],
"y": [0,1,1,1]
}
df_OR=pd.DataFrame(OR)
# -
df_OR
def prepare_data(df,target_col="y"):
X=df.drop(target_col,axis=1)
y=df[target_col]
return X,y
# +
X,y =prepare_data(df_OR)
ETA=0.1
EPOCHS=20
model_or=Perceptron(eta=ETA,epochs=EPOCHS)
model_or.fit(X,y)
# -
model_or.predict(X=[[1,1]])
|
research_env/perceptron implementation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
Let's teach the Robots to distinguish words and numbers.
You are given a string with words and numbers separated by whitespaces (one space).
The words contains only letters. You should check if the string contains three words in succession.
For example, the string "start 5 one two three 7 end" contains three words in succession.
Input: A string with words.
Output: The answer as a boolean.
Example:
checkio("Hello World hello") == True
checkio("He is 123 man") == False
checkio("1 2 3 4") == False
checkio("bla bla bla bla") == True
checkio("Hi") == False
How it is used: This teaches you how to work with strings and introduces some useful functions.
Precondition: The input contains words and/or numbers. There are no mixed words (letters and digits combined).
0 < len(words) < 100
让我们教机器人区分单词和数字。
给你一个字符串,用空白(一个空格)分隔单词和数字。 单词只包含字母。
您应该检查字符串是否包含三个字连续。 例如,字符串“start 5 one two three 7 end”连续包含三个单词。
输入:包含文字的字符串。
输出:作为布尔值的答案。
# +
import re
def checkio(words: str) -> bool:
# demo 1
# c = 0
# for i in words.split(): # 将用split方法分割的字符串for循环遍历
# if i.isalpha(): # 如果是字母的话则c自增1
# c += 1
# else: # 关键点在这,如果其中有一次不是字母,都将c=0
# c = 0
# if c >= 3: # 如果连续三次都是字母返回True
# return True
# else:
# return False
# demo 2
# if re.search('\s?([a-zA-Z]+\s){2}[a-zA-Z]+\s?', words):
# return True
# else:
# return False
# demo 3
# return bool(re.search(r'[a-zA-Z]+\s[a-zA-Z]+\s[a-zA-Z]+', words, flags=re.I))
return bool(re.findall('\D+\s\D+\s\D+', words))
# These "asserts" using only for self-checking and not necessary for auto-testing
if __name__ == '__main__':
assert checkio("Hello World hello") == True, "Hello"
assert checkio("He is 123 man") == False, "123 man"
assert checkio("1 2 3 4") == False, "Digits"
assert checkio("bla bla bla bla") == True, "Bla Bla"
assert checkio("Hi") == False, "Hi"
assert checkio("one two 3 four five six 7 eight 9 ten eleven 12") == True, "four five six"
print("Coding complete? Click 'Check' to review your tests and earn cool rewards!")
|
three-words.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import os
os.environ['CUDA_VISIBLE_DEVICES'] = '1'
from albert import modeling
from albert import optimization
from albert import tokenization
import tensorflow as tf
import numpy as np
import json
tokenizer = tokenization.FullTokenizer(
vocab_file='albert-base-2020-04-10/sp10m.cased.v10.vocab', do_lower_case=False,
spm_model_file='albert-base-2020-04-10/sp10m.cased.v10.model')
# +
import pickle
with open('train_X.pkl', 'rb') as fopen:
train_X, train_Y, train_depends = pickle.load(fopen)
with open('test_X.pkl', 'rb') as fopen:
test_X, test_Y, test_depends = pickle.load(fopen)
with open('tags.pkl', 'rb') as fopen:
idx2tag, tag2idx = pickle.load(fopen)
# -
tag2idx
albert_config = modeling.AlbertConfig.from_json_file('albert-base-2020-04-10/config.json')
albert_config
BERT_INIT_CHKPNT = 'albert-base-2020-04-10/model.ckpt-400000'
epoch = 3
batch_size = 32
warmup_proportion = 0.1
num_train_steps = int(len(train_X) / batch_size * epoch)
num_warmup_steps = int(num_train_steps * warmup_proportion)
class BiAAttention:
def __init__(self, input_size_encoder, input_size_decoder, num_labels):
self.input_size_encoder = input_size_encoder
self.input_size_decoder = input_size_decoder
self.num_labels = num_labels
self.W_d = tf.get_variable("W_d", shape=[self.num_labels, self.input_size_decoder],
initializer=tf.contrib.layers.xavier_initializer())
self.W_e = tf.get_variable("W_e", shape=[self.num_labels, self.input_size_encoder],
initializer=tf.contrib.layers.xavier_initializer())
self.U = tf.get_variable("U", shape=[self.num_labels, self.input_size_decoder, self.input_size_encoder],
initializer=tf.contrib.layers.xavier_initializer())
def forward(self, input_d, input_e, mask_d=None, mask_e=None):
batch = tf.shape(input_d)[0]
length_decoder = tf.shape(input_d)[1]
length_encoder = tf.shape(input_e)[1]
out_d = tf.expand_dims(tf.matmul(self.W_d, tf.transpose(input_d, [0, 2, 1])), 3)
out_e = tf.expand_dims(tf.matmul(self.W_e, tf.transpose(input_e, [0, 2, 1])), 2)
output = tf.matmul(tf.expand_dims(input_d, 1), self.U)
output = tf.matmul(output, tf.transpose(tf.expand_dims(input_e, 1), [0, 1, 3, 2]))
output = output + out_d + out_e
if mask_d is not None:
d = tf.expand_dims(tf.expand_dims(mask_d, 1), 3)
e = tf.expand_dims(tf.expand_dims(mask_e, 1), 2)
output = output * d * e
return output
class BiLinear:
def __init__(self, left_features, right_features, out_features):
self.left_features = left_features
self.right_features = right_features
self.out_features = out_features
self.U = tf.get_variable("U-bi", shape=[out_features, left_features, right_features],
initializer=tf.contrib.layers.xavier_initializer())
self.W_l = tf.get_variable("Wl", shape=[out_features, left_features],
initializer=tf.contrib.layers.xavier_initializer())
self.W_r = tf.get_variable("Wr", shape=[out_features, right_features],
initializer=tf.contrib.layers.xavier_initializer())
def forward(self, input_left, input_right):
left_size = tf.shape(input_left)
output_shape = tf.concat([left_size[:-1], [self.out_features]], axis = 0)
batch = tf.cast(tf.reduce_prod(left_size[:-1]), tf.int32)
input_left = tf.reshape(input_left, (batch, self.left_features))
input_right = tf.reshape(input_right, (batch, self.right_features))
tiled = tf.tile(tf.expand_dims(input_left, axis = 0), (self.out_features,1,1))
output = tf.transpose(tf.reduce_sum(tf.matmul(tiled, self.U), axis = 2))
output = output + tf.matmul(input_left, tf.transpose(self.W_l))\
+ tf.matmul(input_right, tf.transpose(self.W_r))
return tf.reshape(output, output_shape)
# +
_NEG_INF = -1e9
class Model:
def __init__(
self,
learning_rate,
hidden_size_word,
training = True,
cov = 0.0):
self.words = tf.placeholder(tf.int32, (None, None))
self.heads = tf.placeholder(tf.int32, (None, None))
self.types = tf.placeholder(tf.int32, (None, None))
self.switch = tf.placeholder(tf.bool, None)
self.mask = tf.cast(tf.math.not_equal(self.words, 0), tf.float32)
self.maxlen = tf.shape(self.words)[1]
self.lengths = tf.count_nonzero(self.words, 1)
mask = self.mask
heads = self.heads
types = self.types
self.arc_h = tf.layers.Dense(hidden_size_word)
self.arc_c = tf.layers.Dense(hidden_size_word)
self.attention = BiAAttention(hidden_size_word, hidden_size_word, 1)
self.type_h = tf.layers.Dense(hidden_size_word)
self.type_c = tf.layers.Dense(hidden_size_word)
self.bilinear = BiLinear(hidden_size_word, hidden_size_word, len(tag2idx))
model = modeling.AlbertModel(
config=albert_config,
is_training=training,
input_ids=self.words,
input_mask=self.mask,
use_one_hot_embeddings=False)
output_layer = model.get_sequence_output()
arc_h = tf.nn.elu(self.arc_h(output_layer))
arc_c = tf.nn.elu(self.arc_c(output_layer))
self._arc_h = arc_h
self._arc_c = arc_c
type_h = tf.nn.elu(self.type_h(output_layer))
type_c = tf.nn.elu(self.type_c(output_layer))
self._type_h = type_h
self._type_c = type_c
out_arc = tf.squeeze(self.attention.forward(arc_h, arc_c, mask_d=self.mask,
mask_e=self.mask), axis = 1)
self.out_arc = out_arc
batch = tf.shape(out_arc)[0]
max_len = tf.shape(out_arc)[1]
sec_max_len = tf.shape(out_arc)[2]
batch_index = tf.range(0, batch)
decode_arc = out_arc + tf.linalg.diag(tf.fill([max_len], -np.inf))
minus_mask = tf.expand_dims(tf.cast(1 - mask, tf.bool), axis = 2)
minus_mask = tf.tile(minus_mask, [1, 1, sec_max_len])
decode_arc = tf.where(minus_mask, tf.fill(tf.shape(decode_arc), -np.inf), decode_arc)
self.decode_arc = decode_arc
self.heads_seq = tf.argmax(decode_arc, axis = 1)
self.heads_seq = tf.identity(self.heads_seq, name = 'heads_seq')
# self.decode_arc_t = tf.transpose(decode_arc, (0, 2, 1))
# sequence_loss_depends = tf.contrib.seq2seq.sequence_loss(logits = self.decode_arc_t,
# targets = self.heads,
# weights = mask)
t = tf.cast(tf.transpose(self.heads_seq), tf.int32)
broadcasted = tf.broadcast_to(batch_index, tf.shape(t))
concatenated = tf.transpose(tf.concat([tf.expand_dims(broadcasted, axis = 0),
tf.expand_dims(t, axis = 0)], axis = 0))
type_h = tf.gather_nd(type_h, concatenated)
out_type = self.bilinear.forward(type_h, type_c)
self.tags_seq = tf.argmax(out_type, axis = 2)
self.tags_seq = tf.identity(self.tags_seq, name = 'tags_seq')
log_likelihood, transition_params = tf.contrib.crf.crf_log_likelihood(
out_type, self.types, self.lengths
)
crf_loss = tf.reduce_mean(-log_likelihood)
self.logits, _ = tf.contrib.crf.crf_decode(
out_type, transition_params, self.lengths
)
self.logits = tf.identity(self.logits, name = 'logits')
batch = tf.shape(out_arc)[0]
max_len = tf.shape(out_arc)[1]
batch_index = tf.range(0, batch)
t = tf.transpose(heads)
broadcasted = tf.broadcast_to(batch_index, tf.shape(t))
concatenated = tf.transpose(tf.concat([tf.expand_dims(broadcasted, axis = 0),
tf.expand_dims(t, axis = 0)], axis = 0))
type_h = tf.gather_nd(type_h, concatenated)
out_type = self.bilinear.forward(type_h, type_c)
minus_inf = -1e8
minus_mask = (1 - mask) * minus_inf
out_arc = out_arc + tf.expand_dims(minus_mask, axis = 2) + tf.expand_dims(minus_mask, axis = 1)
loss_arc = tf.nn.log_softmax(out_arc, dim=1)
loss_type = tf.nn.log_softmax(out_type, dim=2)
loss_arc = loss_arc * tf.expand_dims(mask, axis = 2) * tf.expand_dims(mask, axis = 1)
loss_type = loss_type * tf.expand_dims(mask, axis = 2)
num = tf.reduce_sum(mask) - tf.cast(batch, tf.float32)
child_index = tf.tile(tf.expand_dims(tf.range(0, max_len), 1), [1, batch])
t = tf.transpose(heads)
broadcasted = tf.broadcast_to(batch_index, tf.shape(t))
concatenated = tf.transpose(tf.concat([tf.expand_dims(broadcasted, axis = 0),
tf.expand_dims(t, axis = 0),
tf.expand_dims(child_index, axis = 0)], axis = 0))
loss_arc = tf.gather_nd(loss_arc, concatenated)
loss_arc = tf.transpose(loss_arc, [1, 0])
t = tf.transpose(types)
broadcasted = tf.broadcast_to(batch_index, tf.shape(t))
concatenated = tf.transpose(tf.concat([tf.expand_dims(broadcasted, axis = 0),
tf.expand_dims(child_index, axis = 0),
tf.expand_dims(t, axis = 0)], axis = 0))
loss_type = tf.gather_nd(loss_type, concatenated)
loss_type = tf.transpose(loss_type, [1, 0])
cost = (tf.reduce_sum(-loss_arc) / num) + (tf.reduce_sum(-loss_type) / num)
self.cost = tf.cond(self.switch, lambda: cost + crf_loss, lambda: cost)
self.optimizer = optimization.create_optimizer(self.cost, learning_rate,
num_train_steps, num_warmup_steps, False)
mask = tf.sequence_mask(self.lengths, maxlen = self.maxlen)
self.prediction = tf.boolean_mask(self.logits, mask)
mask_label = tf.boolean_mask(self.types, mask)
correct_pred = tf.equal(tf.cast(self.prediction, tf.int32), mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
self.prediction = tf.cast(tf.boolean_mask(self.heads_seq, mask), tf.int32)
mask_label = tf.boolean_mask(self.heads, mask)
correct_pred = tf.equal(self.prediction, mask_label)
correct_index = tf.cast(correct_pred, tf.float32)
self.accuracy_depends = tf.reduce_mean(tf.cast(correct_pred, tf.float32))
# +
tf.reset_default_graph()
sess = tf.InteractiveSession()
learning_rate = 2e-5
hidden_size_word = 256
model = Model(learning_rate, hidden_size_word)
sess.run(tf.global_variables_initializer())
# -
var_lists = tf.get_collection(tf.GraphKeys.TRAINABLE_VARIABLES, scope = 'bert')
saver = tf.train.Saver(var_list = var_lists)
saver.restore(sess, BERT_INIT_CHKPNT)
# +
from tensorflow.keras.preprocessing.sequence import pad_sequences
batch_x = train_X[:5]
batch_x = pad_sequences(batch_x,padding='post')
batch_y = train_Y[:5]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = train_depends[:5]
batch_depends = pad_sequences(batch_depends,padding='post')
# -
sess.run([model.accuracy, model.accuracy_depends, model.cost],
feed_dict = {model.words: batch_x,
model.types: batch_y,
model.heads: batch_depends,
model.switch: False})
sess.run([model.accuracy, model.accuracy_depends, model.cost],
feed_dict = {model.words: batch_x,
model.types: batch_y,
model.heads: batch_depends,
model.switch: True})
# +
from tqdm import tqdm
epoch = 2
for e in range(epoch):
train_acc, train_loss = [], []
test_acc, test_loss = [], []
train_acc_depends, test_acc_depends = [], []
pbar = tqdm(
range(0, len(train_X), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(train_X))
batch_x = train_X[i: index]
batch_x = pad_sequences(batch_x,padding='post')
batch_y = train_Y[i: index]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = train_depends[i: index]
batch_depends = pad_sequences(batch_depends,padding='post')
if batch_x.shape == batch_y.shape:
acc_depends, acc, cost, _ = sess.run(
[model.accuracy_depends, model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.words: batch_x,
model.types: batch_y,
model.heads: batch_depends,
model.switch: False
},
)
train_loss.append(cost)
train_acc.append(acc)
train_acc_depends.append(acc_depends)
pbar.set_postfix(cost = cost, accuracy = acc, accuracy_depends = acc_depends)
pbar = tqdm(
range(0, len(test_X), batch_size), desc = 'test minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(test_X))
batch_x = test_X[i: index]
batch_x = pad_sequences(batch_x,padding='post')
batch_y = test_Y[i: index]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = test_depends[i: index]
batch_depends = pad_sequences(batch_depends,padding='post')
if batch_x.shape == batch_y.shape:
acc_depends, acc, cost = sess.run(
[model.accuracy_depends, model.accuracy, model.cost],
feed_dict = {
model.words: batch_x,
model.types: batch_y,
model.heads: batch_depends,
model.switch: False
},
)
test_loss.append(cost)
test_acc.append(acc)
test_acc_depends.append(acc_depends)
pbar.set_postfix(cost = cost, accuracy = acc, accuracy_depends = acc_depends)
print(
'epoch: %d, training loss: %f, training acc: %f, training depends: %f, valid loss: %f, valid acc: %f, valid depends: %f\n'
% (e, np.mean(train_loss),
np.mean(train_acc),
np.mean(train_acc_depends),
np.mean(test_loss),
np.mean(test_acc),
np.mean(test_acc_depends)
))
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'albert-base-dependency/model.ckpt')
# +
epoch = 1
for e in range(epoch):
train_acc, train_loss = [], []
test_acc, test_loss = [], []
train_acc_depends, test_acc_depends = [], []
pbar = tqdm(
range(0, len(train_X), batch_size), desc = 'train minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(train_X))
batch_x = train_X[i: index]
batch_x = pad_sequences(batch_x,padding='post')
batch_y = train_Y[i: index]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = train_depends[i: index]
batch_depends = pad_sequences(batch_depends,padding='post')
if batch_x.shape == batch_y.shape:
acc_depends, acc, cost, _ = sess.run(
[model.accuracy_depends, model.accuracy, model.cost, model.optimizer],
feed_dict = {
model.words: batch_x,
model.types: batch_y,
model.heads: batch_depends,
model.switch: True
},
)
train_loss.append(cost)
train_acc.append(acc)
train_acc_depends.append(acc_depends)
pbar.set_postfix(cost = cost, accuracy = acc, accuracy_depends = acc_depends)
pbar = tqdm(
range(0, len(test_X), batch_size), desc = 'test minibatch loop'
)
for i in pbar:
index = min(i + batch_size, len(test_X))
batch_x = test_X[i: index]
batch_x = pad_sequences(batch_x,padding='post')
batch_y = test_Y[i: index]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = test_depends[i: index]
batch_depends = pad_sequences(batch_depends,padding='post')
if batch_x.shape == batch_y.shape:
acc_depends, acc, cost = sess.run(
[model.accuracy_depends, model.accuracy, model.cost],
feed_dict = {
model.words: batch_x,
model.types: batch_y,
model.heads: batch_depends,
model.switch: True
},
)
test_loss.append(cost)
test_acc.append(acc)
test_acc_depends.append(acc_depends)
pbar.set_postfix(cost = cost, accuracy = acc, accuracy_depends = acc_depends)
print(
'epoch: %d, training loss: %f, training acc: %f, training depends: %f, valid loss: %f, valid acc: %f, valid depends: %f\n'
% (e, np.mean(train_loss),
np.mean(train_acc),
np.mean(train_acc_depends),
np.mean(test_loss),
np.mean(test_acc),
np.mean(test_acc_depends)
))
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'albert-base-dependency/model.ckpt')
# -
def merge_sentencepiece_tokens_tagging(x, y):
new_paired_tokens = []
n_tokens = len(x)
rejected = ['[CLS]', '[SEP]']
i = 0
while i < n_tokens:
current_token, current_label = x[i], y[i]
if not current_token.startswith('▁') and current_token not in rejected:
previous_token, previous_label = new_paired_tokens.pop()
merged_token = previous_token
merged_label = [previous_label]
while (
not current_token.startswith('▁')
and current_token not in rejected
):
merged_token = merged_token + current_token.replace('▁', '')
merged_label.append(current_label)
i = i + 1
current_token, current_label = x[i], y[i]
merged_label = merged_label[0]
new_paired_tokens.append((merged_token, merged_label))
else:
new_paired_tokens.append((current_token, current_label))
i = i + 1
words = [
i[0].replace('▁', '')
for i in new_paired_tokens
if i[0] not in rejected
]
labels = [i[1] for i in new_paired_tokens if i[0] not in rejected]
return words, labels
# +
import re
from unidecode import unidecode
from malaya.function.parse_dependency import DependencyGraph
PUNCTUATION = '!"#$%&\'()*+,./:;<=>?@[\]^_`{|}~'
def transformer_textcleaning(string):
"""
use by any transformer model before tokenization
"""
string = unidecode(string)
string = re.sub('\\(dot\\)', '.', string)
string = (
re.sub(re.findall(r'\<a(.*?)\>', string)[0], '', string)
if (len(re.findall(r'\<a (.*?)\>', string)) > 0)
and ('href' in re.findall(r'\<a (.*?)\>', string)[0])
else string
)
string = re.sub(
r'\w+:\/{2}[\d\w-]+(\.[\d\w-]+)*(?:(?:\/[^\s/]*))*', ' ', string
)
string = re.sub(r'[ ]+', ' ', string).strip().split()
string = [w for w in string if w[0] != '@']
string = ' '.join(string)
string = re.sub(f'([{PUNCTUATION}])', r' \1 ', string)
string = re.sub('\s{2,}', ' ', string)
original_string = string.split()
string = [
(original_string[no], word.title() if word.isupper() else word)
for no, word in enumerate(string.split())
if len(word)
]
return [s[0] for s in string], [s[1] for s in string]
def parse_X(left):
bert_tokens = ['[CLS]']
for no, orig_token in enumerate(left):
t = tokenizer.tokenize(orig_token)
bert_tokens.extend(t)
bert_tokens.append("[SEP]")
t = tokenizer.convert_tokens_to_ids(bert_tokens)
return t, bert_tokens, [1] * len(t)
# -
def dependency_graph(tagging, indexing):
"""
Return helper object for dependency parser results. Only accept tagging and indexing outputs from dependency models.
"""
result = []
for i in range(len(tagging)):
result.append(
'%d\t%s\t_\t_\t_\t_\t%d\t%s\t_\t_'
% (i + 1, tagging[i][0], int(indexing[i][1]), tagging[i][1])
)
return DependencyGraph('\n'.join(result), top_relation_label='root')
string = '<NAME>'
sequence = transformer_textcleaning(string)[1]
parsed_sequence, bert_sequence, mask = parse_X(sequence)
h, t = sess.run([model.heads_seq, model.tags_seq],
feed_dict = {
model.words: [parsed_sequence],
},
)
h = h[0] - 2
t = [idx2tag[d] for d in t[0]]
merged_h = merge_sentencepiece_tokens_tagging(bert_sequence, h)
merged_t = merge_sentencepiece_tokens_tagging(bert_sequence, t)
tagging = list(zip(merged_t[0], merged_t[1]))
indexing = list(zip(merged_h[0], merged_h[1]))
dep = dependency_graph(tagging, indexing)
dep.to_graphvis()
string = 'KUALA LUMPUR: Ketua Penerangan BERSATU, Datuk Wan Saiful Wan Jan membidas kenyataan Datuk Seri Najib Razak dan Ketua Pemuda UMNO, Datuk Dr <NAME> Dusuki yang mempertikaikan tindakan kerajaan melaksanakan sekatan pergerakan penuh. Beliau berkata, Najib dan <NAME> sengaja memetik kenyataan Perdana Menteri, Tan Sri Muhyiddin Yassin yang tidak lengkap untuk mengelirukan rakyat. Wan Saiful berkata, beliau sudah menjangka ada kenyataan balas daripada Najib mengenai tulisan beliau berhubung kesan positif sekatan pergerakan penuh.'
sequence = transformer_textcleaning(string)[1]
parsed_sequence, bert_sequence, mask = parse_X(sequence)
h, t = sess.run([model.heads_seq, model.tags_seq],
feed_dict = {
model.words: [parsed_sequence],
},
)
h = h[0] - 2
t = [idx2tag[d] for d in t[0]]
merged_h = merge_sentencepiece_tokens_tagging(bert_sequence, h)
merged_t = merge_sentencepiece_tokens_tagging(bert_sequence, t)
tagging = list(zip(merged_t[0], merged_t[1]))
indexing = list(zip(merged_h[0], merged_h[1]))
dep = dependency_graph(tagging, indexing)
dep.to_graphvis()
saver = tf.train.Saver(tf.trainable_variables())
saver.save(sess, 'albert-base-dependency/model.ckpt')
# +
tf.reset_default_graph()
sess = tf.InteractiveSession()
learning_rate = 2e-5
hidden_size_word = 256
model = Model(learning_rate, hidden_size_word, training = False)
sess.run(tf.global_variables_initializer())
saver = tf.train.Saver(tf.trainable_variables())
saver.restore(sess, 'albert-base-dependency/model.ckpt')
# -
def pred2label(pred):
out = []
for pred_i in pred:
out_i = []
for p in pred_i:
out_i.append(idx2tag[p])
out.append(out_i)
return out
def evaluate(heads_pred, types_pred, heads, types, lengths,
symbolic_root=False, symbolic_end=False):
batch_size, _ = heads_pred.shape
ucorr = 0.
lcorr = 0.
total = 0.
ucomplete_match = 0.
lcomplete_match = 0.
corr_root = 0.
total_root = 0.
start = 1 if symbolic_root else 0
end = 1 if symbolic_end else 0
for i in range(batch_size):
ucm = 1.
lcm = 1.
for j in range(start, lengths[i] - end):
total += 1
if heads[i, j] == heads_pred[i, j]:
ucorr += 1
if types[i, j] == types_pred[i, j]:
lcorr += 1
else:
lcm = 0
else:
ucm = 0
lcm = 0
if heads[i, j] == 0:
total_root += 1
corr_root += 1 if heads_pred[i, j] == 0 else 0
ucomplete_match += ucm
lcomplete_match += lcm
return ucorr / total, lcorr / total, corr_root / total_root
# +
arcs, types, roots = [], [], []
real_Y, predict_Y = [], []
for i in tqdm(range(0, len(test_X), batch_size)):
index = min(i + batch_size, len(test_X))
batch_x = test_X[i: index]
batch_x = pad_sequences(batch_x,padding='post')
batch_y = test_Y[i: index]
batch_y = pad_sequences(batch_y,padding='post')
batch_depends = test_depends[i: index]
batch_depends = pad_sequences(batch_depends,padding='post')
tags_seq, heads = sess.run(
[model.logits, model.heads_seq],
feed_dict = {
model.words: batch_x,
},
)
arc_accuracy, type_accuracy, root_accuracy = evaluate(heads - 1, tags_seq, batch_depends - 1, batch_y,
np.count_nonzero(batch_x, axis = 1))
arcs.append(arc_accuracy)
types.append(type_accuracy)
roots.append(root_accuracy)
predicted = pred2label(tags_seq)
real = pred2label(batch_y)
predict_Y.extend(predicted)
real_Y.extend(real)
# +
temp_real_Y = []
for r in real_Y:
temp_real_Y.extend(r)
temp_predict_Y = []
for r in predict_Y:
temp_predict_Y.extend(r)
# -
from sklearn.metrics import classification_report
print(classification_report(temp_real_Y, temp_predict_Y, digits = 5))
print('arc accuracy:', np.mean(arcs))
print('types accuracy:', np.mean(types))
print('root accuracy:', np.mean(roots))
strings = ','.join(
[
n.name
for n in tf.get_default_graph().as_graph_def().node
if ('Variable' in n.op
or 'Placeholder' in n.name
or '_seq' in n.name
or 'alphas' in n.name
or 'logits' in n.name
or 'self/Softmax' in n.name)
and 'Adam' not in n.name
and 'beta' not in n.name
and 'global_step' not in n.name
and 'adam' not in n.name
and 'gradients/bert' not in n.name
]
)
def freeze_graph(model_dir, output_node_names):
if not tf.gfile.Exists(model_dir):
raise AssertionError(
"Export directory doesn't exists. Please specify an export "
'directory: %s' % model_dir
)
checkpoint = tf.train.get_checkpoint_state(model_dir)
input_checkpoint = checkpoint.model_checkpoint_path
absolute_model_dir = '/'.join(input_checkpoint.split('/')[:-1])
output_graph = absolute_model_dir + '/frozen_model.pb'
clear_devices = True
with tf.Session(graph = tf.Graph()) as sess:
saver = tf.train.import_meta_graph(
input_checkpoint + '.meta', clear_devices = clear_devices
)
saver.restore(sess, input_checkpoint)
output_graph_def = tf.graph_util.convert_variables_to_constants(
sess,
tf.get_default_graph().as_graph_def(),
output_node_names.split(','),
)
with tf.gfile.GFile(output_graph, 'wb') as f:
f.write(output_graph_def.SerializeToString())
print('%d ops in the final graph.' % len(output_graph_def.node))
freeze_graph('albert-base-dependency', strings)
transforms = ['add_default_attributes',
'remove_nodes(op=Identity, op=CheckNumerics, op=Dropout)',
'fold_batch_norms',
'fold_old_batch_norms',
'quantize_weights(fallback_min=-10, fallback_max=10)',
'strip_unused_nodes',
'sort_by_execution_order']
# +
from tensorflow.tools.graph_transforms import TransformGraph
tf.set_random_seed(0)
pb = 'albert-base-dependency/frozen_model.pb'
input_graph_def = tf.GraphDef()
with tf.gfile.FastGFile(pb, 'rb') as f:
input_graph_def.ParseFromString(f.read())
if 'bert' in pb:
inputs = ['Placeholder']
a = ['dense/BiasAdd']
if 'xlnet' in pb:
inputs = ['Placeholder', 'Placeholder_1', 'Placeholder_2']
a = ['transpose_3']
transformed_graph_def = TransformGraph(input_graph_def,
inputs,
['logits', 'heads_seq'] + a, transforms)
with tf.gfile.GFile(f'{pb}.quantized', 'wb') as f:
f.write(transformed_graph_def.SerializeToString())
|
session/dependency/albert-base.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Machine Learning for Finance Freestyle
#
# In this lab you'll be given the opportunity to apply everything you have learned to build a trading strategy for SP500 stocks. First, let's introduce the dataset you'll be using.
# ## The Data
#
# Use BigQuery's magic function to pull data as follows:
#
# Dataset Name: ml4f
# Table Name: percent_change_sp500
#
# The following query will pull 10 rows of data from the table:
# %%bigquery df
SELECT
*
FROM
`cloud-training-prod-bucket.ml4f.percent_change_sp500`
LIMIT
10000
df.head()
# As you can see, the table contains daily open and close data for SP500 stocks. The table also contains some features that have been generated for you using [navigation functions](https://cloud.google.com/bigquery/docs/reference/standard-sql/navigation_functions) and [analytic functions](https://cloud.google.com/bigquery/docs/reference/standard-sql/analytic-function-concepts). Let's dig into the schema a bit more.
# %%bigquery
SELECT
* EXCEPT(is_generated, generation_expression, is_stored, is_updatable)
FROM
`cloud-training-prod-bucket.ml4f`.INFORMATION_SCHEMA.COLUMNS
WHERE
table_name = "percent_change_sp500"
# Most of the features, like `open` and `close` are pretty straightforward. The features generated using analytic functions, such as `close_MIN_prior_5_days` are best described using an example. Let's take the 6 most recent rows of data for IBM and reproduce the `close_MIN_prior_5_days` column.
# %%bigquery
SELECT
*
FROM
`cloud-training-prod-bucket.ml4f.percent_change_sp500`
WHERE
symbol = 'IBM'
ORDER BY
Date DESC
LIMIT 6
# For `Date = 2013-02-01` how did we arrive at `close_MIN_prior_5_days = 0.989716`? The minimum close over the past five days was `203.07`. This is normalized by the current day's close of `205.18` to get `close_MIN_prior_5_days = 203.07 / 205.18 = 0.989716`. The other features utilizing analytic functions were generated in a similar way. Here are explanations for some of the other features:
#
# * __scaled_change__: `tomo_close_m_close / close`
# * __s_p_scaled_change__: This value is calculated the same way as `scaled_change` but for the S&P 500 index.
# * __normalized_change__: `scaled_change - s_p_scaled_change` The normalization using the S&P index fund helps ensure that the future price of a stock is not due to larger market effects. Normalization helps us isolate the factors contributing to the performance of a stock_market.
# * __direction__: This is the target variable we're trying to predict. The logic for this variable is as follows:
#
# ```sql
# CASE
# WHEN normalized_change < -0.01 THEN 'DOWN'
# WHEN normalized_change > 0.01 THEN 'UP'
# ELSE 'STAY'
# END AS direction
# ```
# ## Create classification model for `direction`
#
# In this example, your job is to create a classification model to predict the `direction` of each stock. Be creative! You can do this in any number of ways. For example, you can use BigQuery, Scikit-Learn, or AutoML. Feel free to add additional features, or use time series models.
#
#
# ### Establish a Simple Benchmark
#
# One way to assess the performance of a model is to compare it to a simple benchmark. We can do this by seeing what kind of accuracy we would get using the naive strategy of just predicting the majority class. Across the entire dataset, the majority class is 'STAY'. Using the following query we can see how this naive strategy would perform.
# %%bigquery
WITH subset as (
SELECT
Direction
FROM
`cloud-training-prod-bucket.ml4f.percent_change_sp500`
WHERE
tomorrow_close IS NOT NULL
)
SELECT
Direction,
100.0 * COUNT(*) / (SELECT COUNT(*) FROM subset) as percentage
FROM
subset
GROUP BY
Direction
# So, the naive strategy of just guessing the majority class would have accuracy of around 54% across the entire dataset. See if you can improve on this.
# ### Train Your Own Model
# +
# %matplotlib inline
import pandas as pd
import matplotlib.pyplot as plt
import numpy as np
from sklearn import linear_model
from sklearn.multiclass import OneVsRestClassifier
from sklearn.svm import SVC
from sklearn.metrics import mean_squared_error
from sklearn.metrics import r2_score
plt.rc('figure', figsize=(12, 8.0))
# -
df.head()
df['Date'] = pd.to_datetime(df['Date'])
# +
# TODO: Write code to build a model to predict Direction
# -
df.info()
print(type(df))
df.dropna(inplace=True)
df.sort_values(by=['Date'], inplace=True, ascending=True)
df.head()
df.columns
df.plot(x='Date', y='Close');
df.shape
def encode(arg):
switcher = {
'STAY': 0,
'UP': 1,
'DOWN': -1
}
return switcher.get(arg, -5)
encode('UP')
df['direction_encoded'] = df.apply(lambda x: encode(x['direction']), axis=1)
# +
features = [
'close_MIN_prior_5_days',
'close_MIN_prior_20_days',
'close_MIN_prior_260_days',
'close_MAX_prior_5_days',
'close_MAX_prior_20_days',
'close_MAX_prior_260_days',
'close_AVG_prior_5_days',
'close_AVG_prior_20_days',
'close_AVG_prior_260_days',
# 'symbol',
'days_on_market',
'scaled_change',
's_p_scaled_change',
'normalized_change'
]
target = 'direction_encoded'
X_train, X_test = df.loc[:8000, features], df.loc[8000:, features]
Y_train, Y_test = df.loc[:8000, target], df.loc[8000:, target]
# clf = OneVsRestClassifier(SVC()).fit(X_train, Y_train)
regr = linear_model.LinearRegression(fit_intercept=False)
regr.fit(X_train, Y_train)
y_pred = regr.predict(X_test)
# y_pred = clf.predict(X_test)
print('RMSE: {0:.2f}'.format(np.sqrt(mean_squared_error(Y_test, y_pred))))
# -
print('Variance Score: {0:.2f}'.format(r2_score(Y_test, y_pred)))
plt.scatter(Y_test, y_pred)
plt.xlabel('Actual')
plt.ylabel('Predicted')
plt.legend();
|
freestyle.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp read_data
# -
# # read_data
#
# > read and preprocess netcdf data
from nbdev import *
#export
import xarray as xr
from glob import glob
#export
def transform_calendar(ds,
timedim="time",
calendarname="proleptic_gregorin"):
"""Transforms calendar of time index in xarray dataset"""
ds[timedim].attrs['calendar'] = calendarname
return ds
show_doc(transform_calendar)
# Use in combination with `xr.open_dataset([...], decode_time=False)`
#
# Args:
# - ds (object): xarray dataset
#
# Kwargs:
#
# - timedim (string): name of time dim
# - calendarname (string): calendar name
#
# Returns:
# - ds (xarray dataset)
#export
def read_netcdfs(files, dim, transform_func, transform_calendar=None, cftime = True):
"""Reads multiples netcdfs files. Should be used when open_mfdatasets is to slow."""
def process_one_path(path):
if transform_calendar is not None:
calendar = False
else:
calendar = True
with xr.open_dataset(path, decode_times = calendar, use_cftime = cftime) as ds:
if transform_calendar is not None:
ds[dim].attrs['calendar'] = transform_calendar
ds = xr.decode_cf(ds, use_cftime = cftime)
if transform_func is not None:
ds = transform_func(ds)
ds.load()
return ds
paths = sorted(glob(files))
datasets = [process_one_path(p) for p in paths]
combined = xr.concat(datasets, dim)
return combined
show_doc(read_netcdfs)
# Args:
# - files : Path to files (str)
# - dim : dimension to concat files (if transform_calendar=T, concat along time)
# - transform_func : additional preprocessing option
# - transform_calendar : name of calendar (see function transform_calendar())
#
# Returns:
# - combined netcdf files as xarray object
#
# Example:
#
#
# combined = read_netcdfs("*.nc", dim = "TIME",
# transform_func=lambda ds:ds.salt,
# transform_calendar="proleptic_gregorian")
|
07_read_data.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: PyCharm (mit_model_code)
# language: python
# name: pycharm-43a0cb91
# ---
# # Import packages
import datetime
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.metrics import f1_score, recall_score, precision_score
# # Define some constants
DEMOGRAPHICS_PATH = "./data/processed/demographics.csv"
RESPONDENTS_PATH = "./data/processed/all_respondents.csv"
SAVE_PLOT_PATH = "./plots/"
SAVE_PLOT = True
# # Read in the processed data sets
demographics = pd.read_csv(DEMOGRAPHICS_PATH, parse_dates=["start_date", "end_date"], infer_datetime_format=True, dtype={"respondent_id": "object"})
all_respondents = pd.read_csv(RESPONDENTS_PATH, dtype={"respondent_id": "object"})
# # Define some helper functions
# +
def plot_accuracy_by_group(group_category, group_names, df_input=demographics, font_size=24):
"""Automatically create an boxplot to show the accuracy distribution by the group_category"""
# determine the post-processing step based on group_category
if group_category in ("highest_academic_degree_held", "current_position"):
df_pivot = df_input[["respondent_id", "accuracy", group_category]]
xticklabels = group_names
else:
df_pivot = df_input.melt(id_vars=["respondent_id", "accuracy"],
var_name=group_category,
value_vars=group_names).dropna().drop(columns="value")
xticklabels = [name.replace("_", " ").capitalize() for name in group_names]
# count the number of people within each subgroup
count_by_group = df_pivot.value_counts(subset=group_category)
# generate a summary table for the accuracy distribution
summary_by_group = df_pivot.groupby(by=group_category).describe()
# create the boxplot
plt.figure(figsize=(12, 9))
plot_by_group = sns.boxplot(data = df_pivot, x=group_category, y="accuracy", order=group_names)
plot_by_group.set_xticklabels(xticklabels)
plot_by_group.set_xlabel(group_category.replace("_", " ").capitalize(), fontsize=24)
plot_by_group.set_ylabel("Accuracy", fontsize=font_size)
plot_by_group.tick_params(labelsize=font_size)
# annotate the number of people within each subgroup
x_bottom, _ = plot_by_group.get_xlim()
_, y_top = plot_by_group.get_ylim()
plot_by_group.annotate("# of\npeople", xy=(x_bottom, y_top),
ha="right", va="bottom", fontsize=font_size, color="black")
for index, name in enumerate(group_names):
plot_by_group.annotate(str(count_by_group.loc[name]), xy=(index, y_top),
ha="center", va="bottom", fontsize=24, color="black")
return summary_by_group, plot_by_group
def plot_descriptor_usage(which, df_input, num_compounds=5, font_size=15):
"""A function to create the usage plot."""
# determine whether to plot the top or bottom 5 compounds in terms of classification accuracy
if which == "top":
compound_descriptors = (
descriptor_usage_ranked.head(n=num_compounds).apply(np.sum, axis=0) / (53 * num_compounds) * 100
).sort_values(ascending=False)
title = "Descriptor Usage of {} Most Accurate Compounds".format(num_compounds)
elif which == "bottom":
compound_descriptors = (
descriptor_usage_ranked.tail(n=num_compounds).apply(np.sum, axis=0) / (53 * num_compounds) * 100
).sort_values(ascending=False)
title = "Descriptor Usage of {} Least Accurate Compounds".format(num_compounds)
else:
raise KeyError('Invalid input: please use either "top" or "bottom"')
# create the plot
plt.figure(figsize=(8, 6))
compound_descriptor_plot = sns.barplot(x=compound_descriptors.index, y=compound_descriptors.values, color="#1f77b4",
saturation=1)
compound_descriptor_plot.set_ylabel("Relative frequency (%)", fontsize=font_size)
compound_descriptor_plot.set_title(title, fontsize=15)
_ = plt.setp(compound_descriptor_plot.get_xticklabels(), rotation=-45, ha="left", rotation_mode="anchor")
compound_descriptor_plot.tick_params(labelsize=font_size)
return compound_descriptor_plot
def eval_performance(df_input, pos_class, groupby_col="respondent_id"):
"""Create a dataframe that contains the human performance for a given binary classification task."""
# binarize the label
replace_dict = {"metal": 0, "insulator": 0, "mit": 0}
replace_dict[pos_class] = 1
df_eval = df_input.replace({"true_label": replace_dict, "predicted_label": replace_dict})
# a convenient function to evaluate performance with the functions in score_funcs
def custom_score(df_subgroup, score_funcs, avg_method="weighted"):
true_values = df_subgroup["true_label"]
predicted_value = df_subgroup["predicted_label"]
return [score_func(true_values, predicted_value, average=avg_method) for score_func in score_funcs]
# evaluate precision, recall, f1_score within each binary classification task
df_eval_grouped = df_eval.groupby(by=[groupby_col]).apply(func=custom_score, score_funcs=[precision_score, recall_score, f1_score])
df_performance = pd.DataFrame(df_eval_grouped.to_list(), columns=["precision_weighted", "recall_weighted", "f1_weighted"])
df_performance[groupby_col] = df_eval_grouped.index
df_performance = df_performance[[groupby_col, "precision_weighted", "recall_weighted", "f1_weighted"]]
df_result = df_performance.melt(id_vars="respondent_id",
value_vars=["precision_weighted", "recall_weighted", "f1_weighted"],
var_name="metric_name", value_name="raw_metric")
df_result["model_type"] = "Human"
df_result["positive_class"] = pos_class.capitalize() if pos_class!= "mit" else pos_class.upper()
return df_result[["model_type", "positive_class", "metric_name", "raw_metric"]]
# -
# ## Demographics dataset
#
# The `demographics` dataset, which as the name suggests, contains the demographics information of all 53 respondents who completed all the questions on the survey.
#
# **Disclaimer**: The demographic-related questions were designed to anonymize each individual response, as **no** names, emails or institution affiliations were collected. The 4 questions on demographics are about
#
# 1. Field of study
# * Materials Science
# * Physics
# * Chemistry
# 2. Highest academic degree held
# * BA/BS
# * MS
# * PhD
# 3. Research type
# * Experimental
# * Computational
# * Theoretical
# 4. Current position
# * Graduate student
# * Postdoc
# * Faculty
# * Staff scientist
demographics.head()
demographics.info()
# ## All respondents dataset
#
# The `all_respondent` dataset contains the individual classification on the included 18 compounds, as well as the descriptors important to each respondent when classifying each compound.
all_respondents.head()
all_respondents.info()
# # Demographics analysis
#
# There are 53 respondents in total.
demographics.shape[0]
# We can quickly check the accuracy distribution of all 53 respondents.
accuracy_dist = sns.histplot(data=demographics, x="accuracy", bins=np.linspace(0, 1, 11))
accuracy_dist.set_xlabel("Accuracy")
# Here are some statistics for the accuracy.
demographics.accuracy.describe()
# We can also examine the relation between accuracy and the time taken to complete the survey.
time_vs_accuracy = sns.scatterplot(x=(demographics.end_date - demographics.start_date) / datetime.timedelta(minutes=1),
y=demographics.accuracy)
time_vs_accuracy.set_xlabel("Completion time (minute)")
time_vs_accuracy.set_ylabel("Accuracy")
# ## Accuracy by field of study
field_of_study_summary, field_of_study_plot= plot_accuracy_by_group("field_of_study",
["materials_science", "physics", "chemistry"])
if SAVE_PLOT: field_of_study_plot.figure.savefig(SAVE_PLOT_PATH + "field_of_study.pdf", dpi=300, bbox_inches="tight")
field_of_study_summary
# ## Accuracy by highest academic degree held
academic_degree_summary, academic_degree_plot= plot_accuracy_by_group("highest_academic_degree_held",
["BA/BS", "MS", "PhD"])
if SAVE_PLOT: academic_degree_plot.figure.savefig(SAVE_PLOT_PATH + "academic_degree.pdf", dpi=300, bbox_inches="tight")
academic_degree_summary
# ## Accuracy by research type
research_type_summary, research_type_plot= plot_accuracy_by_group("research_type",
["experimental", "computational", "theoretical"])
if SAVE_PLOT: research_type_plot.figure.savefig(SAVE_PLOT_PATH + "research_type.pdf", dpi=300, bbox_inches="tight")
research_type_summary
# ## Accuracy by current position
current_position_summary, current_position_plot = plot_accuracy_by_group("current_position",
["Graduate Student", "Postdoc", "Faculty", "Staff Scientist"])
if SAVE_PLOT: current_position_plot.figure.savefig(SAVE_PLOT_PATH + "current_position.pdf", dpi=300, bbox_inches="tight")
current_position_summary
# # Descriptor usage analysis
#
# Let's first find the classification accuracy for each compound.
accuracy_by_compound = all_respondents.groupby(by="formula").apply(func=lambda df: np.mean(df.true_label == df.predicted_label)).to_frame()
accuracy_by_compound = accuracy_by_compound.rename(columns={0: "accuracy"})
accuracy_by_compound = accuracy_by_compound.sort_values(by="accuracy", ascending=False)
accuracy_by_compound
# Then, we need to figure for each compound, how many times each of the 11 descriptors are used.
descriptor_usage = all_respondents.drop(columns=["respondent_id", "true_label",
"predicted_label", "other_please_specify"]).groupby("formula").apply(
func=lambda df: np.sum(~df.isna())
)
descriptor_usage
# We need to join the previous 2 dataframes together.
descriptor_usage_ranked = accuracy_by_compound.join(descriptor_usage).drop(columns=["accuracy", "formula"])
descriptor_usage_ranked = descriptor_usage_ranked.rename(columns={"stoichiometry": "Stoichiometry",
"crystal_structure": "Crystal_structure",
"average_metal_oxygen_bond_distance": "Average_MO_distance",
"total_number_of_valence_electrons": "Valence_electron",
"mass_density": "Mass_density",
"mean_electronegativity_of_elements_in_formula": "Mean_electronegativity",
"polarizability_of_the_compound": "Polarizability",
"standard_deviation_of_average_ionic_radius_of_elements": "Std_of_ionic_radius",
"crystal_field_splitting_energy": "Crystal_field_splitting",
"electronic_correlation": "Electronic_correlation"})
descriptor_usage_ranked
# Plot the descriptor usage for the 5 compounds with the highest accuracy.
five_most_accurate_plot = plot_descriptor_usage("top", descriptor_usage_ranked)
if SAVE_PLOT: five_most_accurate_plot.figure.savefig(SAVE_PLOT_PATH + "five_most_accurate_plot.pdf", dpi=300, bbox_inches="tight")
# Plot the descriptor usage for the 5 compounds with the lowest accuracy.
five_least_accurate_plot = plot_descriptor_usage("bottom", descriptor_usage_ranked)
if SAVE_PLOT: five_least_accurate_plot.figure.savefig(SAVE_PLOT_PATH + "five_least_accurate_plot.pdf", dpi=300, bbox_inches="tight")
# # Evaluate human performance in terms of binary classification tasks
#
# The 3 binary classification tasks are as follows.
#
# 1. Metal vs. non-Metal
# 2. Insulator vs. non-Insulator
# 3. MIT vs. non-MIT
# initialize an empty list to store the results
human_results = []
for positive_class in ["metal", "insulator", "mit"]:
human_results.append(eval_performance(all_respondents, positive_class))
# concatenate the results into a single dataframe
df_human_results = pd.concat(human_results, ignore_index=True)
df_human_results
# Save the results table.
df_human_results.to_csv("./data/processed/human_binary_classification_results.csv", index=False)
# # Check what other descriptors are suggested for each compound
pd.set_option('max_colwidth', 200)
all_respondents[~all_respondents.other_please_specify.isna()][["formula", "other_please_specify"]].sort_values("formula")
|
mit_classification_survey_analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Generate Sine binary radian lookup table
# Binary radians. 0 to 256 = 0 to 2pi
# Output is -127 to + 127
import matplotlib.pyplot as plt
import math
x = range(0,256)
y = []
for x_ in x:
y.append(int(round(math.sin(x_/256.0*(2*math.pi))*127)))
plt.rcParams['figure.figsize'] = [25, 15]
plt.plot(x,y, drawstyle='steps-post')
# +
f = open('sin_lut.dat', 'w')
for x_, y_ in zip(x,y):
s = '{:02x} '.format(struct.pack('b',y_)[0])
print(s)
f.write(s)
f.close()
|
Sim/.ipynb_checkpoints/trig_lut-checkpoint.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Principal Components Regression
#
# Similar to forward stepwise, but use principal components instead of QR decomposition.
import csv
import numpy as np
norm = np.linalg.norm
# +
def dot(a, b):
return (a*b).sum(axis=-2)
def angle(a, b):
# https://stackoverflow.com/a/13849249
return np.arccos(np.clip(dot(a/norm(a), b/norm(b)), -1.0, 1.0))
def single_variate_regression(x, y):
# Assumes x and y are column vectors
# Returns beta
return dot(x, y) / dot(x, x)
def compute_residual(x, z_basis):
gammas = [single_variate_regression(z, x) for z in z_basis]
return x - sum([gamma * z for gamma, z in zip(gammas, z_basis)]), gammas
def prepend_1(x):
# prepend 1 to every row in x
return np.concatenate((np.ones((x.shape[0], 1)), x), axis=1)
# +
with open('../datasets/winequality/winequality-white.csv') as f:
reader = csv.reader(f, delimiter=';')
header = next(reader)
body = np.asarray([[float(value) for value in row] for row in reader])
x = body[:, :-1]
y = body[:, -1:]
N = 1000
train = (x[:N], y[:N])
test = (x[N:], y[N:])
def subset_regression(k):
assert k >= 1
x, y = train
x = prepend_1(x)
# Principal component decomposition
U, d, V = np.linalg.svd(x, full_matrices=False)
V = V.T
ordering = np.argsort(-d) # Highest to lowest variance
idxs = ordering[:k]
y_hat = U[:, idxs] @ U[:, idxs].T @ y
# Compute MSE
train_mse = np.power(y - y_hat, 2).mean()
print('train mse', train_mse)
# Forward stepwise subset selection.
for k in range(1, x.shape[1]):
# Compute regression with different subset sizes.
# Take best fitting features in each subset.
subset_regression(k)
|
chapter03/3.5_Principal_components_regression.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python2
# ---
# +
# %matplotlib inline
import pickle, os
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style("whitegrid",{'axes.grid' : False})
from BayesianNeuralNetwork import *
from HiPMDP import HiPMDP
from __future__ import print_function
from ExperienceReplay import ExperienceReplay
from multiprocessing import Pool
if not os.path.isdir('./results'):
os.mkdir('results')
# -
# ## Generate batch of transition data
# * Toy domain
# * 2 different task instances
# * Agent learning model-free using e-greedy policy based on DQN
domain = 'grid'
run_type = 'modelfree'
num_batch_instances = 2
preset_hidden_params = [{'latent_code':1},{'latent_code':2}]
ddqn_learning_rate = 0.0005
episode_count = 500
bnn_hidden_layer_size = 25
bnn_num_hidden_layers = 3
bnn_network_weights = None
eps_min = 0.15
test_inst = None
create_exp_batch = True
state_diffs = True
grid_beta = 0.1
batch_generator_hipmdp = HiPMDP(domain,preset_hidden_params,
ddqn_learning_rate=ddqn_learning_rate,
episode_count=episode_count,
run_type=run_type, eps_min=eps_min,
create_exp_batch=create_exp_batch,
num_batch_instances=num_batch_instances,
grid_beta=grid_beta,
print_output=True)
# #### *The authors firmly suggest that those walking through this example for the first time test and evaluate the HiP-MDP with the "grid" domain as its runs quicker. The remainder of this notebook is therefore built to support investigation into this simple toy domain*
#
# However, we provide example code of how to intialize a batch for use in either the Acrobot or HIV domains in the cell below.
#
# This example is meant to replace the code in lines 1-4 in the cell above. Testing the HiP-MDP entails selecting any additional parameter setting beyond the batch used here to train the initial BNN and latent weights.
#
# Pay particular attention to other parameters used for training the BNN below. They are hard coded here for demonstration purposes. You may want to reference HiPMDP.py (included in this repo) for domain specific parameters
# +
## Example code of how to initialize a batch generator for HIV or Acrobot
# domain = 'hiv' # 'acrobot'
# with open('preset_parameters/'+domain+'_preset_hidden_params','r') as f:
# preset_parameters = pickle.load(f)
# run_type = 'modelfree'
# num_batch_instances = 5
# preset_hidden_params = preset_parameters[:num_batch_instances]
# -
# This next cell will take a fair amount of time to run, it is generating several examples from the two different task instances to train the BNN and Q-network on.
(exp_buffer, networkweights, rewards, avg_rwd_per_ep, full_task_weights,
sys_param_set, mean_episode_errors, std_episode_errors) = batch_generator_hipmdp.run_experiment()
with open('results/{}_exp_buffer'.format(domain),'w') as f:
pickle.dump(exp_buffer,f)
# +
# with open('results/{}_exp_buffer'.format(domain),'r') as f:
# exp_buffer = pickle.load(f)
# -
# #### Reformat Data
# Create numpy array
exp_buffer_np = np.vstack(exp_buffer)
# Collect the instances that each transition came from
inst_indices = exp_buffer_np[:,4]
inst_indices = inst_indices.astype(int)
# Group experiences by instance
# Create dictionary where keys are instance indexes and values are np.arrays experiences
exp_dict = {}
for idx in xrange(batch_generator_hipmdp.instance_count):
exp_dict[idx] = exp_buffer_np[inst_indices == idx]
X = np.array([np.hstack([exp_buffer_np[tt,0],exp_buffer_np[tt,1]]) for tt in range(exp_buffer_np.shape[0])])
y = np.array([exp_buffer_np[tt,3] for tt in range(exp_buffer_np.shape[0])])
num_dims = 2
num_actions = 4
num_wb = 5
if state_diffs:
# subtract previous state
y -= X[:,:num_dims]
# ## Train BNN and learn latent weights using batch data
# #### Set up BNN and latent weights
relu = lambda x: np.maximum(x, 0.)
param_set = {
'bnn_layer_sizes': [num_dims+num_actions+num_wb]+[bnn_hidden_layer_size]*bnn_num_hidden_layers+[num_dims],
'weight_count': num_wb,
'num_state_dims': num_dims,
'bnn_num_samples': 50,
'bnn_batch_size': 32,
'num_strata_samples': 5,
'bnn_training_epochs': 1,
'bnn_v_prior': 1.0,
'bnn_learning_rate': 0.00005,
'bnn_alpha':0.5,
'wb_num_epochs':1,
'wb_learning_rate':0.0005
}
# Initialize latent weights for each instance
full_task_weights = np.random.normal(0.,0.1,(batch_generator_hipmdp.instance_count,num_wb))
# Initialize BNN
network = BayesianNeuralNetwork(param_set, nonlinearity=relu)
# Compute error before training
l2_errors = network.get_td_error(np.hstack((X,full_task_weights[inst_indices])), y, location=0.0, scale=1.0, by_dim=False)
print ("Before training: Mean Error: {}, Std Error: {}".format(np.mean(l2_errors),np.std(l2_errors)))
np.mean(l2_errors),np.std(l2_errors)
print ("L2 Difference in latent weights between instances: {}".format(np.sum((full_task_weights[0]-full_task_weights[1])**2)))
# #### Oscillate between training BNN and latent weights
def get_random_sample(start,stop,size):
indices_set = set()
while len(indices_set) < size:
indices_set.add(np.random.randint(start,stop))
return np.array(list(indices_set))
# We slowly train the BNN and latent weights $w_b$ so as to avoid prematurely fitting to the separate modes of input. We found that without taking this approach, the BNN associated the latent parameters as noise and ignored their contribution to the performance of the predictive model.
# size of sample to compute error on
sample_size = 10000
for i in xrange(40):
# Update BNN network weights
network.fit_network(exp_buffer_np, full_task_weights, 0, state_diffs=state_diffs,
use_all_exp=True)
print('finished BNN update '+str(i))
if i % 4 == 0:
#get random sample of indices
sample_indices = get_random_sample(0,X.shape[0],sample_size)
l2_errors = network.get_td_error(np.hstack((X[sample_indices],full_task_weights[inst_indices[sample_indices]])), y[sample_indices], location=0.0, scale=1.0, by_dim=False)
print ("After BNN update: iter: {}, Mean Error: {}, Std Error: {}".format(i,np.mean(l2_errors),np.std(l2_errors)))
# Update latent weights
for inst in np.random.permutation(batch_generator_hipmdp.instance_count):
full_task_weights[inst,:] = network.optimize_latent_weighting_stochastic(
exp_dict[inst],np.atleast_2d(full_task_weights[inst,:]),0,state_diffs=state_diffs,use_all_exp=True)
print ('finished wb update '+str(i))
# Compute error on sample of transitions
if i % 4 == 0:
#get random sample of indices
sample_indices = get_random_sample(0,X.shape[0],sample_size)
l2_errors = network.get_td_error(np.hstack((X[sample_indices],full_task_weights[inst_indices[sample_indices]])), y[sample_indices], location=0.0, scale=1.0, by_dim=False)
print ("After Latent update: iter: {}, Mean Error: {}, Std Error: {}".format(i,np.mean(l2_errors),np.std(l2_errors)))
# We check to see if the latent updates are sufficiently different so as to avoid fitting [erroneously] to the same dynamics
print ("L2 Difference in latent weights between instances: {}".format(np.sum((full_task_weights[0]-full_task_weights[1])**2)))
network_weights = network.weights
with open('results/{}_network_weights'.format(domain), 'w') as f:
pickle.dump(network.weights, f)
# with open('results/{}_network_weights'.format(domain), 'r') as f:
# network_weights = pickle.load(f)
# ## Learn dynamics and policy for new instance using HiP-MDP with embedded latent weights
results = {}
run_type = 'full'
create_exp_batch = False
episode_count = 20 # reduce episode count for demonstration since HiPMDP learns policy quickly
for run in xrange(5):
for test_inst in [0,1]:
test_hipmdp = HiPMDP(domain,preset_hidden_params,
ddqn_learning_rate=ddqn_learning_rate,
episode_count=episode_count,
run_type=run_type,
bnn_hidden_layer_size=bnn_hidden_layer_size,
bnn_num_hidden_layers=bnn_num_hidden_layers,
bnn_network_weights=network_weights, test_inst=test_inst,
eps_min=eps_min, create_exp_batch=create_exp_batch,grid_beta=grid_beta,print_output=True)
results[(test_inst,run)] = test_hipmdp.run_experiment()
with open('results/{}_results'.format(domain),'w') as f:
pickle.dump(results,f)
# +
# with open('results/{}_results'.format(domain),'r') as f:
# results = pickle.load(f)
# -
# ## Plot Results
# Group rewards, errors by instance
reward_key = 'Reward'
error_key = 'BNN Error'
clean_results = {(0,reward_key):[],(0,error_key):[],(1,reward_key):[],(1,error_key):[]}
for test_inst in [0,1]:
for run in xrange(5):
clean_results[(test_inst,reward_key)].append(results[(test_inst,run)][2])
clean_results[(test_inst,error_key)].append(results[(test_inst,run)][6])
clean_results[(test_inst,reward_key)] = np.vstack(clean_results[(test_inst,reward_key)])
clean_results[(test_inst,error_key)] = np.vstack(clean_results[(test_inst,error_key)])
def plot_results(clean_results, test_inst):
f, ax_array = plt.subplots(2,figsize=(7,7))
result_names = [reward_key,error_key]
for result_idx in xrange(2):
result = result_names[result_idx]
mean_result = np.mean(clean_results[(test_inst,result)], axis=0)
std_result = np.std(clean_results[(test_inst,result)], axis=0)
ax_array[result_idx].errorbar(x=np.arange(len(mean_result)),y=mean_result,yerr=std_result)
_ = ax_array[result_idx].set_ylim((np.min(mean_result)-0.01,np.max(mean_result)+0.01))
ax_array[result_idx].set_ylabel(result)
ax_array[1].set_xlabel("Episode")
f.suptitle("Full HiP-MDP Training Results Instance {}".format(test_inst),fontsize=12)
plot_results(clean_results, 0)
plot_results(clean_results, 1)
|
toy_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] colab_type="text" id="KQqCmLY16SYq"
# Licensed under the Apache License, Version 2.0.
# + colab={} colab_type="code" id="AgUs80V4_sPl"
from edward2.experimental.attentive_uncertainty.contextual_bandits import run_offline_contextual_bandits_gnp
# + colab={"height": 1000} colab_type="code" executionInfo={"elapsed": 3101784, "status": "ok", "timestamp": 1568773809643, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAa2HvmIe1GgREzEkl3p3FRszir9TfbhGC4qNCS=s64", "userId": "07983129814092358079"}, "user_tz": 420} id="fWax0oL8AAbg" outputId="4d349fa8-8ff7-4aea-a025-189ed261b101"
run_offline_contextual_bandits_gnp.benchmark()
# + colab={"height": 1000} colab_type="code" executionInfo={"elapsed": 2488557, "status": "ok", "timestamp": 1568186650015, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAa2HvmIe1GgREzEkl3p3FRszir9TfbhGC4qNCS=s64", "userId": "07983129814092358079"}, "user_tz": 420} id="PDCCBoOn_2qX" outputId="5cac4beb-dd5e-48e7-f04b-7e66864975ac"
run_offline_contextual_bandits_gnp.benchmark()
# + colab={} colab_type="code" id="K7ose2lU_7kA"
from edward2.experimental.attentive_uncertainty.contextual_bandits import benchmark_gnp
# + colab={"height": 1000} colab_type="code" executionInfo={"elapsed": 313748, "status": "error", "timestamp": 1568187818009, "user": {"displayName": "<NAME>", "photoUrl": "https://lh3.googleusercontent.com/a-/AAuE7mAa2HvmIe1GgREzEkl3p3FRszir9TfbhGC4qNCS=s64", "userId": "07983129814092358079"}, "user_tz": 420} id="N_7jgWQqP05n" outputId="bb42690e-b76c-45bf-ac0d-13e57ced4524"
benchmark_gnp.run_trial(0, 0.5, ['gnp_cnp_offline'])
# + colab={} colab_type="code" id="qnfwGbAEQVzu"
|
experimental/attentive_uncertainty/colabs/2019_09_10_gnp_benchmark.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
import sagemaker
BUCKET = sagemaker.session.Session().default_bucket()
# ## Amazon Polly
# ### synthesize_speech
import boto3
polly = boto3.client('polly')
response = polly.synthesize_speech(
# 作成する音声の文
Text = '''
はじめて Amazon Polly で音声を作成しました。
''',
OutputFormat = "mp3", # 出力形式を選択
VoiceId = 'Takumi', # 声を選択
)
# 音声データをファイルに出力する
with open('./first_voice.mp3', "wb") as f:
f.write(response['AudioStream'].read())
# ### SSML
response = polly.synthesize_speech(
Text = '''
<speak>
<amazon:breath duration="medium" volume="x-loud"/>
私は <break time="1s"/>
<emphasis>SSML</emphasis>
で音声を作成しました
</speak>
''',
OutputFormat = "mp3",
TextType='ssml', # SSML であることを明示する
VoiceId = 'Takumi', # 声を選択
)
with open('./ssml_voice.mp3', "wb") as f:
f.write(response['AudioStream'].read())
# ### Lexicon
LEXICON_NAME = 'emacs'
CONTENT = '''<?xml version="1.0" encoding="UTF-8"?>
<lexicon version="1.0"
xmlns="http://www.w3.org/2005/01/pronunciation-lexicon"
xmlns:xsi="http://www.w3.org/2001/XMLSchema-instance"
xsi:schemaLocation="http://www.w3.org/2005/01/pronunciation-lexicon
http://www.w3.org/TR/2007/CR-pronunciation-lexicon-20071212/pls.xsd"
alphabet="ipa"
xml:lang="ja-JP">
<lexeme><grapheme>テキストエディタ</grapheme><phoneme>ˈiːˌmæks</phoneme></lexeme>
</lexicon>'''
response = polly.put_lexicon(Content=CONTENT,Name=LEXICON_NAME)
# +
# polly.delete_lexicon(Name=LEXICON_NAME)
# +
response = polly.synthesize_speech(
Text = 'テキストエディタを開きましょう',
OutputFormat = "mp3",
VoiceId = 'Takumi',
LexiconNames=[LEXICON_NAME]
)
with open('./emacs.mp3', "wb") as f:
f.write(response['AudioStream'].read())
response = polly.synthesize_speech(
Text = 'テキストエディタを開きましょう',
OutputFormat = "mp3",
VoiceId = 'Takumi',
)
with open('./text_editor.mp3', "wb") as f:
f.write(response['AudioStream'].read())
# -
# ## Amazon Transcribe
# ### start_transcription_job
# +
# transcribe = boto3.client('transcribe')
# JOBNAME = 'my_first_transcription_job'
# transcribe.delete_transcription_job(TranscriptionJobName=JOBNAME)
# +
# 事前に音声ファイルを S3 にアップロード
s3 = boto3.client('s3')
AUDIO_FILE='emacs.mp3'
AUDIO_KEY = f'transcribe/input/{AUDIO_FILE}'
s3.upload_file(f'./{AUDIO_FILE}', BUCKET, AUDIO_KEY)
# transcription job の起動
transcribe = boto3.client('transcribe')
JOBNAME = 'my_first_transcription_job'
OUTPUT_KEY='transcribe/output/'
response = transcribe.start_transcription_job(
TranscriptionJobName=JOBNAME,
LanguageCode='ja-JP',
Media={ 'MediaFileUri': f's3://{BUCKET}/{AUDIO_KEY}'},
OutputBucketName=BUCKET,
OutputKey=OUTPUT_KEY,
)
# -
# Job が完了するまで待つ
from time import sleep
while True:
if transcribe.get_transcription_job(TranscriptionJobName=JOBNAME)['TranscriptionJob']['TranscriptionJobStatus']=='COMPLETED':break
else:sleep(5)
# +
# 結果が格納されている URI の取得
# transcript_result_uri = transcribe.get_transcription_job(
# TranscriptionJobName=JOBNAME
# )['TranscriptionJob']['Transcript']['TranscriptFileUri']
# S3 から手元の環境にダウンロード
boto3.client('s3').download_file(BUCKET, f'{OUTPUT_KEY}{JOBNAME}.json', f'./{JOBNAME}.json')
# 結果の表示
with open(f'./{JOBNAME}.json','r') as f:print(f.read())
# -
# ### create_vocabulary
# +
# 語彙 tsv の作成
CUSTOM_VOCABULARY = 'Phrase\tIPA\tSoundsLike\tDisplayAs\nemacs\t\tiimakkusu\tEmacs'
CUSTOM_VOCABULARY_CSV = './custom_vocabulary.txt'
with open(CUSTOM_VOCABULARY_CSV,'w') as f:f.write(CUSTOM_VOCABULARY)
# 語彙の登録
VOCABULARY_NAME = 'emacs'
VOCABULARY_KEY = f'transcribe/custom_vocaburary/{CUSTOM_VOCABULARY_CSV}'
s3.upload_file(f'./{CUSTOM_VOCABULARY_CSV}', BUCKET, VOCABULARY_KEY)
transcribe.create_vocabulary(
VocabularyName=VOCABULARY_NAME,
LanguageCode='ja-JP',
VocabularyFileUri=f's3://{BUCKET}/{VOCABULARY_KEY}'
)
# +
# transcribe.delete_vocabulary(VocabularyName=VOCABULARY_NAME)
# +
# 登録した語彙を利用した transcription job の起動
JOBNAME = 'custom_vocabulary_transcription_job'
response = transcribe.start_transcription_job(
TranscriptionJobName=JOBNAME,
LanguageCode='ja-JP',
Settings={'VocabularyName': VOCABULARY_NAME},
Media={'MediaFileUri': f's3://{BUCKET}/{AUDIO_KEY}'},
OutputBucketName=BUCKET,
OutputKey=OUTPUT_KEY
)
# 結果の確認 (省略)
# -
# 終わるまで待機
while True:
if transcribe.get_transcription_job(TranscriptionJobName=JOBNAME)['TranscriptionJob']['TranscriptionJobStatus']=='COMPLETED':break
else:sleep(5)
transcript_result_uri = transcribe.get_transcription_job(TranscriptionJobName=JOBNAME)['TranscriptionJob']['Transcript']['TranscriptFileUri']
boto3.client('s3').download_file(BUCKET, f'{OUTPUT_KEY}{JOBNAME}.json', f'./{JOBNAME}.json')
with open(f'./{JOBNAME}.json','r') as f:print(f.read())
# ## Amazon Translate
# ### translate_text
# +
translate = boto3.client('translate')
# 翻訳実行
response = translate.translate_text(
# 翻訳させたい文字列
Text='I use white vaseline daily to protect my skin.',
# 翻訳元の言語
SourceLanguageCode='en',
# 翻訳したい言語
TargetLanguageCode='ja',
# 好ましくない用語をマスクする機能
# Settings={
# 'Profanity': 'MASK'
# }
)
# 結果出力
print(response['TranslatedText'])
# -
# ### import_terminologies
# +
# terminologies の登録
TERMINOLOGIES = 'en,ja\nwhite vaseline,白色ワセリン'.encode()
TERMINOLOGIES_NAME = 'white_vaseline'
translate.import_terminology(
Name=TERMINOLOGIES_NAME,
MergeStrategy='OVERWRITE',
TerminologyData={
'File': TERMINOLOGIES,
'Format': 'CSV',
}
)
# 翻訳実行
response = translate.translate_text(
Text='I use white vaseline daily to protect my skin.',
# terminology の指定
TerminologyNames=[TERMINOLOGIES_NAME],
SourceLanguageCode='en',
TargetLanguageCode='ja',
)
# 結果出力
print(response['TranslatedText'])
# -
# ## Amazon Comprehend
# ### detect_dominant_language
# +
comprehend = boto3.client('comprehend')
# 使用するテキスト
TEXT = 'Toutes les grandes personnes ont d’abord été des enfants, mais peu d’entre elles s’en souviennent.'
# 未知の言語のテキストを翻訳する
response = translate.translate_text(
Text=TEXT,
# 未知の言語を detect_dominant_language API で特定する
SourceLanguageCode=comprehend.detect_dominant_language(Text=TEXT)['Languages'][0]['LanguageCode'],
TargetLanguageCode='ja',
)
# 翻訳結果の表示
print(response['TranslatedText'])
# -
# ### detect_entities
# +
# 使用するテキスト
TEXT = '今日、太郎はラーメンを食べました。おいしかったそうです。'
# エンティティの検出
response = comprehend.detect_entities(
Text=TEXT,
LanguageCode='ja',
)
print(response['Entities'])
# キーフレーズの検出
response = comprehend.detect_key_phrases(
Text=TEXT,
LanguageCode='ja',
)
print(response['KeyPhrases'])
# 感情の検出
response = comprehend.detect_sentiment(
Text=TEXT,
LanguageCode='ja',
)
print(response['SentimentScore'])
# -
|
innovate_code_2022.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <a href="https://www.cognitiveclass.ai"><img src = "https://cognitiveclass.ai/wp-content/themes/bdu3.0/static/images/cc-logo.png" align = left></a>
#
# <br>
# <br>
#
# --------------------
# # Signal to Binary Files (Train&Test)
#
# In this notebook we read the Basic 4 dataset through Spark, and convert signals into a binary file.
# + button=false new_sheet=false run_control={"read_only": false}
import sys
sys.path.insert(0, "/opt/DL/tensorflow/lib/python2.7/site-packages/")
import requests
import json
# #!pip install ibmseti
import ibmseti
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
import tensorflow as tf
import os
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Get file list
#
# I have a list of simulated files stored in an OpenStack Object Storage container that is world-readable.
#
# Download that file, split by lines and parallelize it into an RDD
# + button=false new_sheet=false run_control={"read_only": false}
r = requests.get('https://dal.objectstorage.open.softlayer.com/v1/AUTH_cdbef52bdf7a449c96936e1071f0a46b/simsignals_files/public_list_basic_v2_26may_2017.csv')
# -
filelist_txt = r.text
# + button=false new_sheet=false run_control={"read_only": false}
from pyspark import SparkConf, SparkContext
conf = SparkConf().set("spark.ui.showConsoleProgress", "false")
sc = SparkContext(appName="PythonStatusAPIDemo", conf=conf)
filelist_txt = r.text
fl_rdd = sc.parallelize(filelist_txt.split('\n')[1:-1], 20)
fl_rdd.count()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### Quick Inspection.
#
# Take a look at the first row. Each line is CSV with 'uuid' and 'signal_classification'. We'll convert each line to JSON.
# + button=false new_sheet=false run_control={"read_only": false}
def csvtojson(row):
uuid, sigclass = row.split(',')
return {'file_name':uuid+'.dat', 'uuid':uuid, 'signal_classification':sigclass}
fl_rdd2 = fl_rdd.map(csvtojson)
fl_rdd2.cache()
# + button=false new_sheet=false run_control={"read_only": false}
print fl_rdd2.first()
fl_rdd2.cache()
# + button=false new_sheet=false run_control={"read_only": false}
base_url = 'https://dal.objectstorage.open.softlayer.com/v1/AUTH_cdbef52bdf7a449c96936e1071f0a46b'
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Here is the different "signal_classifications" we have.
# + button=false new_sheet=false run_control={"read_only": false}
container = 'simsignals_basic_v2'
rdd_fname_lb = fl_rdd2.map(lambda row: (row['signal_classification'],row['file_name']))
classes = rdd_fname_lb.map(lambda row: row[0]).distinct().collect()
dictClass = dict(zip(classes, np.arange(4)))
dictClass
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Send request for data, and convert signals to spectogram
# + button=false new_sheet=false run_control={"read_only": false}
def get_spectrogram(fname,h,w,lengthRatio=1.0):
r = requests.get('{}/{}/{}'.format(base_url, container, fname), timeout=4.0)
if r.status_code != 200:
print 'Failed retrieving {}'.format(fname)
print r
return None
else:
aca = ibmseti.compamp.SimCompamp(r.content)
com_data = aca.complex_data()
ratio = int(np.sqrt(len(com_data) *lengthRatio / (h*w)))
if ratio == 0:
raise ValueError, "The selected lenght of signal is less than (Height x Width), select bigger ratio"
elif ratio == 1:
sig_data = com_data[:h*w].reshape(h,w)
spec = np.abs( np.fft.fftshift( np.fft.fft(sig_data), 1) )**2
spec = np.log(spec) # Convert to float (0-255)
image = 255*(spec/np.max(spec))
elif ratio > 1: # resize using IPL image
sig_data = com_data[:h*ratio*w*ratio].reshape(h*ratio,w*ratio)
spec = np.abs( np.fft.fftshift( np.fft.fft(sig_data), 1) )**2
spec = np.log(spec) # Convert to float (0-255)
image = 255*(spec/np.max(spec))
img = Image.fromarray(image)
img = img.resize((int(w), int(h)), Image.ANTIALIAS)
image = np.asarray(img) # float (0-255)
# convert to grayscale: int(0-255)
image = np.uint8(image)
return image
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# h and w are the hight and width of the images, and lengthRatio is the length of signal in ratio.
# + button=false new_sheet=false run_control={"read_only": false}
h = 128 # The hight of output image (bins)
w = 256 # The witdh of output image
lengthRatio = 1.0 # the length-ration of signal to be read. The higher reatio, the better resolution. E.g. 0.5 means half of time sereis.
rdd_gray_spec = rdd_fname_lb.map(lambda row: (row[1], dictClass[row[0]], get_spectrogram(row[1],h,w,lengthRatio)))
# + button=false new_sheet=false run_control={"read_only": false}
rdd_gray_spec.cache()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### verify
# + button=false new_sheet=false run_control={"read_only": false}
z=5
y= rdd_gray_spec.take(z)
for i in range(z):
img = Image.fromarray(np.float32(y[i][2]))
fig, ax = plt.subplots(figsize=(10, 5))
ax.imshow(img)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Train/Test
# + button=false new_sheet=false run_control={"read_only": false}
test, train = rdd_gray_spec.randomSplit(weights=[0.3, 0.7], seed=1)
# + button=false new_sheet=false run_control={"read_only": false}
train_data = train.map(lambda row: row[2]).collect()
train_img_data = np.array(train_data)
train_lbl = train.map(lambda row: row[1]).collect()
# + button=false new_sheet=false run_control={"read_only": false}
test_data = test.map(lambda row: row[2]).collect()
test_img_data = np.array(test_data)
test_lbl = test.map(lambda row: row[1]).collect()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Write to Binary file
# This binary file is same as famouse __mnist__ dataset format to be read by different image processing algorithms, learning techniques and pattern recognition methods.
#
# There are 4 files:
#
# - train-images-idx3-ubyte: training set images
# - train-labels-idx1-ubyte: training set labels
# - test-images-idx3-ubyte: test set images
# - test-labels-idx1-ubyte: test set labels
#
#
# + button=false new_sheet=false run_control={"read_only": false}
from array import *
def wrtieToBinary(ds_directory, name , imgData , lblData,h,w):
n = imgData.shape[0]
imgData = imgData.reshape(-1,)
data_image = array('B')
data_label = array('B')
data_image.extend(imgData)
# number of files in HEX
hexval = "{0:#0{1}x}".format(n,6)
# header for label array
data_label.extend(lblData)
header = array('B')
header.extend([0,0,8,1,0,0])
header.append(int('0x'+hexval[2:][:2],16))
header.append(int('0x'+hexval[2:][2:],16))
data_label = header + data_label
print ('Label header:' )
print(header)
# additional header for images array
if max([w,h]) <= 255:
header.extend([0,0,0,h,0,0,0,w])
else:
hex_h = "{0:#0{1}x}".format(h,6)
header.extend([0,0])
header.append(int('0x'+hex_h[2:][:2],16))
header.append(int('0x'+hex_h[2:][2:],16))
hex_w = "{0:#0{1}x}".format(w,6)
header.extend([0,0])
header.append(int('0x'+hex_w[2:][:2],16))
header.append(int('0x'+hex_w[2:][2:],16))
#raise ValueError('Image exceeds maximum size: 256x256 pixels');
header[3] = 3 # Changing MSB for image data (0x00000803)
if not os.path.exists(ds_directory):
os.makedirs(ds_directory)
print ('Image header:' )
print(header)
data_image = header + data_image
output_file = open(ds_directory + name+'-images-idx3-ubyte', 'wb')
data_image.tofile(output_file)
output_file.close()
output_file = open(ds_directory+ name+'-labels-idx1-ubyte', 'wb')
data_label.tofile(output_file)
output_file.close()
# gzip resulting files
os.system('gzip '+ ds_directory + name +'-images-idx3-ubyte '+ name +'-images-idx3-ubyte.gz')
os.system('gzip '+ ds_directory + name +'-labels-idx1-ubyte ')
# + button=false new_sheet=false run_control={"read_only": false}
ds_directory = 'SETI/SETI_ds_128x256/' # The dataset directory to write the binary files
os.system('rm '+ds_directory+'*')
print os.popen("ls -lrt "+ ds_directory).read()
# + button=false new_sheet=false run_control={"read_only": false}
wrtieToBinary(ds_directory, 'train' , train_img_data , train_lbl, h, w)
# + button=false new_sheet=false run_control={"read_only": false}
wrtieToBinary(ds_directory, 'test' , test_img_data , test_lbl, h, w)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Verify the binary files
# Lets read the binary file and plot an image
# + button=false new_sheet=false run_control={"read_only": false}
print os.popen("ls -lrt "+ ds_directory).read()
# + button=false new_sheet=false run_control={"read_only": false}
import numpy as np
import gzip
def _read32(bytestream):
dt = np.dtype(np.uint32).newbyteorder('>')
return np.frombuffer(bytestream.read(4), dtype=dt)[0]
with open(ds_directory+'train-images-idx3-ubyte.gz', 'rb') as f:
with gzip.GzipFile(fileobj=f ) as bytestream:
magic = _read32(bytestream)
if magic != 2051:
raise ValueError('Invalid magic number %d in MNIST image file: %s' %(magic, f.name))
num_images = _read32(bytestream)
rows = _read32(bytestream)
cols = _read32(bytestream)
buf = bytestream.read(rows * cols * num_images)
print(magic,num_images,rows,cols,)
data = np.frombuffer(buf, dtype=np.uint8)
data = data.reshape(num_images, rows, cols, 1)
# magic, num, rows, cols = struct.unpack(">IIII", bytestream.read(16))
# + button=false new_sheet=false run_control={"read_only": false}
gray_y = data[0].reshape(h,w)
img = Image.fromarray(np.float32(gray_y))
print (img.mode)
fig, ax = plt.subplots(figsize=(10, 5))
ax.imshow(img)
# + button=false new_sheet=false run_control={"read_only": false}
gray_y
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Verify the binary files by reader class
# __SETI.py__ is a helper class, identical to mnist dataset reader, to easily read dataset, one-hot coding, and read images as batch.
# + button=false new_sheet=false run_control={"read_only": false}
# !wget -q --output-document SETI.zip https://ibm.box.com/shared/static/jhqdhcblhua5dx2t7ixwm88okitjrl6l.zip
# !unzip -o SETI.zip
import SETI
# + button=false new_sheet=false run_control={"read_only": false}
SETIds = SETI.read_data_sets(ds_directory, one_hot=True, validation_size=0)
SETIds.train.num_examples
# + button=false new_sheet=false run_control={"read_only": false}
gray_y = SETIds.train.images[0].reshape(h,w)
img = Image.fromarray(gray_y*255)
fig, ax = plt.subplots(figsize=(10, 5))
ax.imshow(img)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Export files to object storage
# You have to move the binary file to __object storage__ if you need to download the binary file.
# + button=false new_sheet=false run_control={"read_only": false}
print os.popen("ls -lrt "+ ds_directory).read()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# You need your credentials to move your files to object storage.
# #### How to get the credentials for object storage?
#
# Check this [link](https://github.com/ibm-cds-labs/seti_at_ibm#object-storage-credentials).
# + button=false new_sheet=false run_control={"read_only": false}
#
credentials_1 = {
'auth_url':'https://identity.open.softlayer.com',
'project':'xx',
'project_id':'xx',
'region':'dallas',
'user_id':'xx',
'domain_id':'xx',
'domain_name':'819515',
'username':'xx',
'password':"xx",
'container':'x',
'tenantId':'undefined',
'filename':'SETI.py'
}
# + button=false new_sheet=false run_control={"read_only": false}
# #!pip install --user --upgrade python-swiftclient
import swiftclient.client as swiftclient
conn = swiftclient.Connection(
key=credentials_1['password'],
authurl=credentials_1['auth_url']+"/v3",
auth_version='3',
os_options={
"project_id": credentials_1['project_id'],
"user_id": credentials_1['user_id'],
"region_name": credentials_1['region']})
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Export the binary files as a single file to object storage
# + button=false new_sheet=false run_control={"read_only": false}
import gzip
local_file = 'SETI128x256.tar.gz'
os.system('tar -zcvf '+local_file+' '+ds_directory)
print ('Moving '+ local_file + '...')
with open(local_file, 'rb') as f:
with gzip.GzipFile(fileobj=f) as bytestream:
etag = conn.put_object(credentials_1['container'], local_file , f.read() )
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Export the binary files as multiple files into object storage
# + button=false new_sheet=false run_control={"read_only": false}
files = ['train-images-idx3-ubyte.gz','train-labels-idx1-ubyte.gz','test-images-idx3-ubyte.gz','test-labels-idx1-ubyte.gz']
for local_file in files:
print ('Moving '+ local_file + '...')
with open(ds_directory + local_file, 'rb') as f:
with gzip.GzipFile(fileobj=f) as bytestream:
etag = conn.put_object(credentials_1['container'], local_file , f.read() )
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Authors
#
# <div class="teacher-image" style=" float: left;
# width: 115px;
# height: 115px;
# margin-right: 10px;
# margin-bottom: 10px;
# border: 1px solid #CCC;
# padding: 3px;
# border-radius: 3px;
# text-align: center;"><img class="alignnone wp-image-2258 " src="https://ibm.box.com/shared/static/tyd41rlrnmfrrk78jx521eb73fljwvv0.jpg" alt="<NAME>" width="178" height="178" /></div>
# #### <NAME>
#
# [<NAME>](https://ca.linkedin.com/in/saeedaghabozorgi), PhD is Sr. Data Scientist in IBM with a track record of developing enterprise level applications that substantially increases clients’ ability to turn data into actionable knowledge. He is a researcher in data mining field and expert in developing advanced analytic methods like machine learning and statistical modelling on large datasets.</p>
#
#
|
tutorials/Step_5c_Convert_TS_to_unit8Dataset_DSX.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# ### Open using Jupyter Notebook. It holds the code and visualizations for developing the different classification algorithms (LibSVM, RBF SVM, Naive Bayes, Random Forest, Gradient Boosting) on the chosen subset of important features.
# +
import pandas as pd
import numpy as np
from numpy import sort
from sklearn.metrics import matthews_corrcoef, accuracy_score,confusion_matrix
from sklearn.feature_selection import SelectFromModel
from matplotlib import pyplot
import pylab as pl
from sklearn import svm
# %matplotlib inline
# +
SEED = 1234
## Selected set of most important features
featureSet=['L3_S31_F3846','L1_S24_F1578','L3_S33_F3857','L1_S24_F1406','L3_S29_F3348','L3_S33_F3863',
'L3_S29_F3427','L3_S37_F3950','L0_S9_F170', 'L3_S29_F3321','L1_S24_F1346','L3_S32_F3850',
'L3_S30_F3514','L1_S24_F1366','L2_S26_F3036']
train_x = pd.read_csv("../data/train_numeric.csv", usecols=featureSet)
train_y = pd.read_csv("../data/train_numeric.csv", usecols=['Response'])
# -
test_x = pd.read_csv("../data/test_numeric.csv", usecols=featureSet)
# +
train_x = train_x.fillna(9999999)
msk = np.random.rand(len(train_x)) < 0.7 # creating Training and validation set
X_train = train_x[msk]
Y_train = train_y.Response.ravel()[msk]
X_valid = train_x[~msk]
Y_valid = train_y.Response.ravel()[~msk]
# -
def showconfusionmatrix(cm, typeModel):
pl.matshow(cm)
pl.title('Confusion matrix for '+typeModel)
pl.colorbar()
pl.show()
# +
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
C=4
lin_svc = svm.LinearSVC(C=C).fit(X_train, Y_train)
print "LibSVM fitted"
# +
title = 'LinearSVC (linear kernel)'
predicted = lin_svc.predict(X_valid)
mcc= matthews_corrcoef(Y_valid, predicted)
print "MCC Score \t +"+title+str(mcc)
cm = confusion_matrix(predicted, Y_valid)
showconfusionmatrix(cm, title)
print "Confusion Matrix"
print (cm)
# +
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
C=4
rbf_svc = svm.SVC(kernel='rbf', gamma=0.7, C=C).fit(X_train, Y_train)
print "RBF fitted"
title = 'SVC with RBF kernel'
predicted = rbf_svc.predict(X_valid)
mcc= matthews_corrcoef(Y_valid, predicted)
print "MCC Score \t +"+title+str(mcc)
cm = confusion_matrix(predicted, Y_valid)
showconfusionmatrix(cm, title)
print "Confusion Matrix"
print (cm)
# +
from sklearn.naive_bayes import GaussianNB
gnb = GaussianNB()
clf = gnb.fit(X_train,Y_train)
print "Naive Bayes Fitted"
title = 'Naive Bayes'
predicted = clf.predict(X_valid)
mcc= matthews_corrcoef(Y_valid, predicted)
print "MCC Score \t +"+title+str(mcc)
cm = confusion_matrix(predicted, Y_valid)
showconfusionmatrix(cm, title)
print "Confusion Matrix"
print (cm)
# -
from sklearn.ensemble import RandomForestClassifier, GradientBoostingClassifier
from sklearn.cross_validation import cross_val_score
from sklearn.model_selection import GridSearchCV
rf = RandomForestClassifier(n_estimators=20, n_jobs=2)
param_grid = {
'n_estimators': [5, 10, 15, 20],
'max_depth': [2, 5, 7, 9]
}
grid_rf = GridSearchCV(rf, param_grid, cv=10)
rf_model=grid_rf.fit(X_train, Y_train)
# +
print "RF fitted"
titles = 'Random Forest'
predicted = rf_model.predict(X_valid)
mcc= matthews_corrcoef(Y_valid, predicted)
print "MCC Score \t +"+titles[0]+str(mcc)
cm = confusion_matrix(predicted, Y_valid)
showconfusionmatrix(cm, titles[0])
# -
gb = GradientBoostingClassifier(learning_rate=0.5)
param_grid = {
'n_estimators': [5, 10, 15, 20],
'max_depth': [2, 5, 7, 9]
}
grid_gb = GridSearchCV(gb, param_grid, cv=10)
gb_model=grid_gb.fit(X_train, Y_train)
# +
print "GB fitted"
title = 'Gradient Boosting'
predicted = gb_model.predict(X_valid)
mcc= matthews_corrcoef(Y_valid, predicted)
print "MCC Score \t +"+title+str(mcc)
cm = confusion_matrix(predicted, Y_valid)
showconfusionmatrix(cm, title)
|
scripts/train.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# # Xarray-spatial
# ### User Guide: Getting Set-up
# ### Installation
#
# The package can be easily installed via conda or pip.
#
# #### To install with conda run:
# conda install -c conda-forge xarray-spatial
#
# #### To install with pip run:
# pip install xarray-spatial
#
# #### To verify whether the installation was successful, open a Python session and import the package:
# import xrspatial
|
examples/user_guide/0_Getting_Setup.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] _uuid="51b9b35792a863134d05653832ddcfc9b9fc5d55"
# # Census income classification with Keras
#
# To download a copy of this notebook visit [github](https://github.com/slundberg/shap/tree/master/notebooks).
# + _uuid="6348460e1429f5690a9a674bb424d0e3222c2180"
from sklearn.model_selection import train_test_split
from keras.layers import Input, Dense, Flatten, Concatenate, concatenate, Dropout, Lambda
from keras.models import Model
from keras.layers.embeddings import Embedding
from tqdm import tqdm
import shap
# print the JS visualization code to the notebook
shap.initjs()
# + [markdown] _uuid="b0341c8c8a042cfdb8025a6cf3085680a5f29bb8"
# ## Load dataset
# + _uuid="a2dfb6995f1861cb752180e0c0510f2a850f0077"
import pandas as pd
# + _uuid="ea7795557158ad2e532c37ea1be9ac63c6a6922d"
df = pd.read_csv('../input/adult.csv')
# + _uuid="2e19963a553fbd4999320513c7692300cb82f458"
df.head()
# + _uuid="0bbc6f358f02f1532e5f51948c7a17acc5a867a3"
df.dtypes
# + _uuid="b0337930f39dcc955467584298d284c9d372f627"
X_display = df.drop('income',axis=1)
y_display = df['income']
# + _uuid="63d119fdcc100b22fef39e98ca4493b14c5c52b8"
int_columns = df.select_dtypes(['int64']).columns
df[int_columns] = df[int_columns].astype('float32')
cat_columns = df.select_dtypes(['object']).columns
df[cat_columns] = df[cat_columns].astype('category')
df[cat_columns] = df[cat_columns].apply(lambda x: x.cat.codes)
# + _uuid="1daf6acd6e8e7e9d3eb751472475934a3f99ee90"
X = df.drop('income',axis=1)
y = df['income']
# + _uuid="1c51d6e50a753206296a2660ffc14f4f4817c4fd"
#X,y = shap.datasets.adult()
#X_display,y_display = shap.datasets.adult(display=True)
# normalize data (this is important for model convergence)
dtypes = list(zip(X.dtypes.index, map(str, X.dtypes)))
for k,dtype in dtypes:
if dtype == "float32":
X[k] -= X[k].mean()
X[k] /= X[k].std()
X_train, X_valid, y_train, y_valid = train_test_split(X, y, test_size=0.2, random_state=7)
# + _uuid="33985f54eb3c6b51b856319b23e994bdfe65b9e0"
X.head()
# + [markdown] _uuid="306fd19253fc1d632983032997cbc1912b56f1ec"
# ## Train Keras model
# + _uuid="87f3b8d1dec4511815b14254e653128cdc251d2f"
# build model
input_els = []
encoded_els = []
for k,dtype in dtypes:
input_els.append(Input(shape=(1,)))
if dtype == "int8":
e = Flatten()(Embedding(X_train[k].max()+1, 1)(input_els[-1]))
else:
e = input_els[-1]
encoded_els.append(e)
encoded_els = concatenate(encoded_els)
layer1 = Dropout(0.5)(Dense(100, activation="relu")(encoded_els))
out = Dense(1)(layer1)
# train model
regression = Model(inputs=input_els, outputs=[out])
regression.compile(optimizer="adam", loss='binary_crossentropy')
regression.fit(
[X_train[k].values for k,t in dtypes],
y_train,
epochs=50,
batch_size=512,
shuffle=True,
validation_data=([X_valid[k].values for k,t in dtypes], y_valid)
)
# + [markdown] _uuid="0b40c91ff32d2827687563dc4acf4dbac96632ec"
# ## Explain predictions
#
# Here we take the Keras model trained above and explain why it makes different predictions for different individuals. SHAP expects model functions to take a 2D numpy array as input, so we define a wrapper function around the original Keras predict function.
# + _uuid="e34ed91bdcff6a302d6bd63e25865562b5312c3b"
def f(X):
return regression.predict([X[:,i] for i in range(X.shape[1])]).flatten()
# + [markdown] _uuid="294a0b684b28ac377ea5149261e304a66653c91b"
# ### Explain a single prediction
#
# Here we use a selection of 50 samples from the dataset to represent "typical" feature values, and then use 500 perterbation samples to estimate the SHAP values for a given prediction. Note that this requires 500 * 50 evaluations of the model.
# + _uuid="9a064f9e15039c5701e70c91a6d660338d1dd037"
explainer = shap.KernelExplainer(f, X.iloc[:100,:])
shap_values = explainer.shap_values(X.iloc[350,:], nsamples=500)
shap.force_plot(shap_values, X_display.iloc[350,:])
# + _uuid="1fda8fb62140b0cf9364db4effbe65553c2450ea"
shap_values = explainer.shap_values(X.iloc[167,:], nsamples=500)
shap.force_plot(shap_values, X_display.iloc[167,:])
# + [markdown] _uuid="655321b925d8c4f728552145a026d4b380c3b88a"
# ### Explain many predictions
#
# Here we repeat the above explanation process for 50 individuals. Since we are using a sampling based approximation each explanation can take a couple seconds depending on your machine setup.
# + _uuid="b7a22507b6192e8b57e414711fdd66bdcf07fd0c"
shap_values = explainer.shap_values(X.iloc[100:330,:], nsamples=500)
# + _uuid="a1bd82f19f873a46faf276d4cfaf2324ae8d275b"
shap.force_plot(shap_values, X_display.iloc[100:330,:])
# + _uuid="7835b6813b830531c6ef075d5a8fabe4650bf4ba"
shap.summary_plot(shap_values50, X.iloc[100:330,:])
# + _uuid="9e8c0b215a7420d5cc19a66be46657f1664a9250"
shap.dependence_plot("marital.status",
shap_values,
X.iloc[100:330,:],
display_features=X_display.iloc[100:330,:])
# + _uuid="0c69511164b8753e8cb5e34c7dcbd9d23c246e4d"
|
9.3_SHAP.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Notebook
#store the next available id for all new notes
import datetime
last_id = 0
# ## Note Class
class Note:
'''Represents a note in a notebook. Match against a string of searches
and store tags for each note'''
def __init__(self, memo, tags=''):
'''iniitialize a note with memo and optional space-separated tags.
Automatically set the note's creation date and unique id.'''
self.memo = memo
self.tags = tags
self.creation_date = datetime.date.today()
global last_id
last_id += 1
self.id = last_id
def match(self, filter):
'''Determine if this note matches the filter
text. Return True if it matches, False otherwise
Search is case sensitive and matches both text and tags.
'''
return filter in self.memo or filter in self.tags
# ## Notebook Class
class Notebook:
'''Represents a collection of notes that can be tagged, modified, and searched'''
def __init__(self):
'''Initialize a notebook with an empty list.'''
self.notes = []
def new_note(self, memo, tags=''):
'''Create a new note and add it to the list.'''
self.notes.append(Note(memo, tags))
def _find_note(self, note_id):
'''Locate the note with the given id.'''
for note in self.notes:
if note.id == note_id:
return note
return None
def modify_memo(self, note_id, memo):
'''Find the note with the given id and change its memo to the given value'''
self._find_note(note_id).memo = memo
def modify_tags(self, note_id, tags):
'''find the note with the given id and change its tags to the given value'''
for note in self.notes:
if note.id == note_id:
note.tags = tags
break
def search(self, filter):
'''Find all notes that match the given filter string.'''
return [note for note in self.notes if note.match(filter)]
# ### Convert to Python Executable
# +
# #!ipython nbconvert --to=python notebook.ipynb
# -
|
case_studies/notebook/notebook.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# +
# coding: utf-8
import numpy as np
def OR(x1, x2):
x = np.array([x1, x2])
w = np.array([0.5, 0.5])
b = -0.2
tmp = np.sum(w*x) + b
if tmp <= 0:
return 0
else:
return 1
if __name__ == '__main__':
for xs in [(0, 0), (1, 0), (0, 1), (1, 1)]:
y = OR(xs[0], xs[1])
print(str(xs) + " -> " + str(y))
|
ch02/or_gate.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# **NOTĂ**
# Când utilizăm [Binder](https://mybinder.org/), comanda ``python -m pip install --user numpy matplotlib`` trebuie rulată în terminal înainte de pornirea **notebook**-ului!
# # Reprezentarea traiectoriilor în planul fazelor
# Avem nevoie de următoarele [module Python 3](https://docs.python.org/3/reference/import.html):
import numpy as np
import matplotlib.pyplot as plt
# Se consideră sistemul diferenţial liniar şi omogen
# $$
# \left\{
# \begin{array}{l}
# x^{\prime}=a_{11}\cdot x+a_{12}\cdot y,\\
# y^{\prime}=a_{21}\cdot x+a_{22}\cdot y,
# \end{array}
# \right.
# \quad\mbox{unde }a_{ij}\in\mathbb{R},\,i,j\in\{1,2\},
# $$
# cu necunoscutele $x=x(t),\,y=y(t)\in C^{1}(\mathbb{R},\mathbb{R})$.
# Introducem **coeficienţii** ecuaţiilor din sistem sub forma şirului
# $$a_{11},\,a_{12},\,a_{21},\,a_{22}$$
# +
r = range(2)
ma = np.array([0, 1, -4, 0], dtype=float).reshape(2, 2)
print("Verific\u0103ri:")
for i in r:
ii = i + 1
for j in r:
print("a[{0},{1}] = {2:.6f}".format(ii, j + 1, ma[i, j]))
# -
# Sistemului îi asociem **data Cauchy**
# $$
# \left\{
# \begin{array}{l}
# x(0)=x_{0},\\
# y(0)=y_{0},
# \end{array}
# \right.
# \quad\mbox{unde }x_{0},y_{0}\in\mathbb{R}.
# $$
# +
dc = np.array([0, 1], dtype=float)
print("Verific\u0103ri:")
for i in r:
print("dc[{0}] = {1:.6f}".format(i, dc[i]))
# -
# Folosim metoda [RK4](https://en.wikipedia.org/wiki/Runge%E2%80%93Kutta_methods#The_Runge%E2%80%93Kutta_method) de **discretizare** a sistemului diferenţial. Pentru aceasta, trebuie precizate:
# * un **pas** $h$;
# * un **număr de iteraţii** $n$.
# Pasul $h$ este:
# +
h = 0.1
print("Verific\u0103ri:"
"h = {0:.6f}".format(h))
# -
# Numărul $n$ de iteraţii este:
# +
n = 1_000
print("Verific\u0103ri:"
"n = {0:d}".format(n))
# -
# **Formulele ecuaţiilor** din sistemul diferenţial sunt:
# $$
# \left\{
# \begin{array}{l}
# f(x,y,t)=a_{11}\cdot x+a_{12}\cdot y,\\
# g(x,y,t)=a_{21}\cdot x+a_{22}\cdot y.
# \end{array}
# \right.
# $$
f = lambda v1, v2, v3: ma[0, 0] * v1 + ma[0, 1] * v2
g = lambda v1, v2, v3: ma[1, 0] * v1 + ma[1, 1] * v2
# **Relaţiile de recurenţă** ale metodei **RK4** (conform <NAME>, *An Introduction to Dynamical Systems: Continuous and Discrete*, Pearson Education, Inc., 2004, **pag. 81**) definesc:
# * elemente principale ($x_{n},\,y_{n},\,t_{n}$):
# $$
# \left\{
# \begin{array}{l}
# x_{n+1} = x_{n} + kx_{RK4,n},\\
# y_{n+1} = y_{n} + ky_{RK4,n},\\
# t_{n+1} = t_{n} + h;
# \end{array}
# \right.
# $$
# * elemente intermediare ($kx_{1,n},\,ky_{1,n},\,zx_{1,n},\,zy_{1,n},kx_{2,n},\,ky_{2,n},\,zx_{2,n},\,zy_{2,n},\,kx_{3,n},\,ky_{3,n},\,zx_{3,n},\,zy_{3,n},\,kx_{4,n},\,ky_{4,n},\,kx_{RK4,n},\,ky_{RK4,n}$):
# $$
# \left\{
# \begin{array}{ll}
# kx_{1,n} = f(x_{n},y_{n},t_{n}),&[\mbox{pasul }\mathbf{1}]\\
# ky_{1,n} = g(x_{n},y_{n},t_{n}),&\\
# zx_{1,n} = x_{n} + \frac{h}{2}\cdot kx_{1,n},&\\
# zy_{1,n} = y_{n} + \frac{h}{2}\cdot ky_{1,n},&\\
# &\\
# kx_{2,n} = f\left(zx_{1,n},zy_{1,n},t_{n} + \frac{h}{2}\right),&[\mbox{pasul }\mathbf{2}]\\
# ky_{2,n} = g\left(zx_{1,n},zy_{1,n},t_{n} + \frac{h}{2}\right),&\\
# zx_{2,n} = x_{n} + \frac{h}{2}\cdot kx_{2,n},&\\
# zy_{2,n} = y_{n} + \frac{h}{2}\cdot ky_{2,n},&\\
# &\\
# kx_{3,n} = f\left(zx_{2,n},zy_{2,n},t_{n} + \frac{h}{2}\right),&[\mbox{pasul }\mathbf{3}]\\
# ky_{3,n} = g\left(zx_{2,n},zy_{2,n},t_{n} + \frac{h}{2}\right),&\\
# zx_{3,n} = x_{n} + h\cdot kx_{3,n},&\\
# zy_{3,n} = y_{n} + h\cdot ky_{3,n},&\\
# &\\
# kx_{4,n} = f(zx_{3,n},zy_{3,n},t_{n} + h),&[\mbox{pasul }\mathbf{4}]\\
# ky_{4,n} = g(zx_{3,n},zy_{3,n},t_{n} + h),&\\
# &\\
# kx_{RK4,n} = \frac{h}{6}\cdot(kx_{1,n} + 2\cdot kx_{2,n} + 2\cdot kx_{3,n} + kx_{4,n}),&[\mbox{pasul }\mathbf{5}]\\
# ky_{RK4,n} = \frac{h}{6}\cdot(ky_{1,n} + 2\cdot ky_{2,n} + 2\cdot ky_{3,n} + ky_{4,n}).&
# \end{array}
# \right.
# $$
# +
r2 = range(n)
r3 = range(n + 1)
x = np.array(r3, dtype=float)
y = np.array(r3, dtype=float)
t = np.array(r3, dtype=float)
print("Verific\u0103ri:\n"
"Obiectul x = {0:s},"
"\nObiectul y = {1:s},"
"\nObiectul t = {2:s}.".format(str(id(x)), str(id(y)), str(id(t))))
kx1 = np.array(r2, dtype=float)
ky1 = np.array(r2, dtype=float)
zx1 = np.array(r2, dtype=float)
zy1 = np.array(r2, dtype=float)
print("Obiectul kx1 = {0:s},"
"\nObiectul ky1 = {1:s},"
"\nObiectul zx1 = {2:s},"
"\nObiectul zy1 = {3:s}.".format(str(id(kx1)), str(id(ky1)), str(id(zx1)), str(id(zy1))))
kx2 = np.array(r2, dtype=float)
ky2 = np.array(r2, dtype=float)
zx2 = np.array(r2, dtype=float)
zy2 = np.array(r2, dtype=float)
print("Obiectul kx2 = {0:s},"
"\nObiectul ky2 = {1:s},"
"\nObiectul zx2 = {2:s},"
"\nObiectul zy2 = {3:s}.".format(str(id(kx2)), str(id(ky2)), str(id(zx2)), str(id(zy2))))
kx3 = np.array(r2, dtype=float)
ky3 = np.array(r2, dtype=float)
zx3 = np.array(r2, dtype=float)
zy3 = np.array(r2, dtype=float)
print("Obiectul kx3 = {0:s},"
"\nObiectul ky3 = {1:s},"
"\nObiectul zx3 = {2:s},"
"\nObiectul zy3 = {3:s}.".format(str(id(kx3)), str(id(ky3)), str(id(zx3)), str(id(zy3))))
kx4 = np.array(r2, dtype=float)
ky4 = np.array(r2, dtype=float)
print("Obiectul kx4 = {0:s},"
"\nObiectul ky4 = {1:s}.".format(str(id(kx4)), str(id(ky4))))
kxRK4 = np.array(r2, dtype=float)
kyRK4 = np.array(r2, dtype=float)
print("Obiectul kxRK4 = {0:s},"
"\nObiectul kyRK4 = {1:s}.".format(str(id(kxRK4)), str(id(kyRK4))))
x[0] = dc[0]
y[0] = dc[1]
t[0] = 0.0
h2 = 0.5 * h
h3 = h / 6.0
print("h = {0:.6f}, h/2 = {1:.6f}, h/6 = {2:.6f}".format(h, h2, h3))
# +
print("Afi\u015Farea calculului:\n"
"x(0) = {0:.6f}, y(0) = {1:.6f}, "
"t(0) = {2:.6f}".format(x[0], y[0], t[0]))
for i in r2:
i2 = i + 1
ti = t[i]
ti2 = ti + h2
ti3 = ti + h
t[i2] = ti3 # t[i+1] = t[i] + h
kx1[i] = f(x[i], y[i], ti) # [pasul 1]
ky1[i] = g(x[i], y[i], ti)
zx1[i] = x[i] + h2 * kx1[i]
zy1[i] = y[i] + h2 * ky1[i]
kx2[i] = f(zx1[i], zy1[i], ti2) # [pasul 2]
ky2[i] = g(zx1[i], zy1[i], ti2)
zx2[i] = x[i] + h2 * kx2[i]
zy2[i] = y[i] + h2 * ky2[i]
kx3[i] = f(zx2[i], zy2[i], ti2) # [pasul 3]
ky3[i] = g(zx2[i], zy2[i], ti2)
zx3[i] = x[i] + h * kx3[i]
zy3[i] = y[i] + h * ky3[i]
kx4[i] = f(zx3[i], zy3[i], ti3) # [pasul 4]
ky4[i] = g(zx3[i], zy3[i], ti3)
kxRK4[i] = h3 * (kx1[i] + 2 * kx2[i] + 2 * kx3[i] + kx4[i]) # [pasul 5]
kyRK4[i] = h3 * (ky1[i] + 2 * ky2[i] + 2 * ky3[i] + ky4[i])
x[i2] = x[i] + kxRK4[i]
y[i2] = y[i] + kyRK4[i]
print("+++\nItera\u0163ia {0:d}:\n"
"Elemente principale:".format(i2))
print("x({0}) = {1:.6f}, "
"y({2}) = {3:.6f}, "
"t({4}) = {5:.6f}".format(i2, x[i2], i2, y[i2], i2, t[i2]))
print("Elemente intermediare:\n"
"[Pasul 1]\n"
"kx1({0}) = {1:.6f}, "
"ky1({2}) = {3:.6f}, "
"zx1({4}) = {5:.6f}, "
"zy1({6}) = {7:.6f}".format(i, kx1[i], i, ky1[i], i, zx1[i], i, zy1[i]))
print("[Pasul 2]\n"
"kx2({0}) = {1:.6f}, "
"ky2({2}) = {3:.6f}, "
"zx2({4}) = {5:.6f}, "
"zy2({6}) = {7:.6f}".format(i, kx2[i], i, ky2[i], i, zx2[i], i, zy2[i]))
print("[Pasul 3]\n"
"kx3({0}) = {1:.6f}, "
"ky3({2}) = {3:.6f}, "
"zx3({4}) = {5:.6f}, "
"zy3({6}) = {7:.6f}".format(i, kx3[i], i, ky3[i], i, zx3[i], i, zy3[i]))
print("[Pasul 4]\n"
"kx4({0}) = {1:.6f}, "
"ky4({2}) = {3:.6f}".format(i, kx4[i], i, ky4[i]))
print("[Pasul 5]\n"
"kxRK4({0}) = {1:.6f}, "
"kyRK4({2}) = {3:.6f}".format(i, kxRK4[i] , i, kyRK4[i]))
# +
ca = np.linspace(-1.5, 1.5,
num=9, endpoint=True,
retstep=False, dtype=float) # Calculul axelor.
plt.figure()
plt.axis([-3, 3, -3, 3])
plt.xlabel('x')
plt.ylabel('y')
plt.title('Planul fazelor')
plt.plot(ca, np.zeros(ca.shape), 'b--') # Axa orizontala.
plt.plot(np.zeros(ca.shape), ca, 'b--') # Axa verticala.
plt.plot(0, 0, 'ro') # Intersectia axelor.
plt.text(.1, -.4, r'$O$')
plt.plot(x, y, 'g.')
plt.text(-2.5,2,
"$a_{11}$" + " = {0:.6f}, ".format(ma[0, 0])
+ "$a_{12}$" + " = {0:.6f},\n".format(ma[0, 1])
+ "$a_{21}$" + " = {0:.6f}, ".format(ma[1, 0])
+ "$a_{22}$" + " = {0:.6f}".format(ma[1, 1]) )
plt.text(-2.5, -2.5,
"$x(0)={0:.6f},\,y(0)={1:.6f}$"
"\nNum\u0103rul de itera\u0163ii: "
"$n={2:d}$. Pasul: $h={3:.6f}$.".format(x[0], y[0], n, h))
plt.show()
|
intro_SD_2019_laborator1_orbite_2D_sisteme_liniare.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Seq2Vec Sentiment Modeling in Tensorflow
# ## 1.0 - Import Packages
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras import layers
from tensorflow.keras import losses
from tensorflow.keras import regularizers
import matplotlib.pyplot as plt
import tensorflow_hub as hub
import pandas as pd
from sklearn.metrics import mean_absolute_error
from keras import backend as K
from keras.layers import Dropout
import os
import numpy as np
import seaborn as sns
import sys
module_path = os.path.abspath(os.path.join('..'))
if module_path not in sys.path:
sys.path.append(module_path)
from src.utils import *
CUR_DIR = os.path.abspath(os.curdir)
ROOT_DIR = os.path.dirname(CUR_DIR)
IMAGES_DIR = os.path.join(ROOT_DIR, "images")
DATA_DIR = os.path.join(ROOT_DIR, "data")
MODELS_DIR = os.path.join(ROOT_DIR, "models")
EVAL_DIR = os.path.join(ROOT_DIR, "evaluation")
MODEL_PERF_DIR = os.path.join(EVAL_DIR, "model_performance")
GRAPHS_DIR = os.path.join(EVAL_DIR, "graphs")
writepath = os.path.join(MODEL_PERF_DIR, "performance.csv")
# + [markdown] tags=[]
# ## 1.1 Import Data
# +
validation_df = pd.read_csv(os.path.join(DATA_DIR,'raw','validation.csv'))
training_df = pd.read_csv(os.path.join(DATA_DIR,'raw','training.csv'))
test_df = pd.read_csv(os.path.join(DATA_DIR,'raw','test.csv'))
X_train = training_df['review']
y_train = training_df['star']
X_val = validation_df['review']
y_val = validation_df['star']
X_test = test_df['review']
y_test = test_df['star']
# Convert to tensorflow datasets
train_ds = tf.data.Dataset.from_tensor_slices((X_train,y_train)).shuffle(buffer_size=1024).batch(128)
test_ds = tf.data.Dataset.from_tensor_slices((X_test,y_test)).shuffle(buffer_size=1024).batch(128)
val_ds = tf.data.Dataset.from_tensor_slices((X_val,y_val)).shuffle(buffer_size=1024).batch(128)
# -
# ## 2.0 Create embedding layer & Build simple model
# +
handle = 'https://tfhub.dev/google/tf2-preview/gnews-swivel-20dim-with-oov/1'
emb_layer = hub.KerasLayer(handle = handle, output_shape=(20,1),
input_shape=(None,),dtype=tf.string,trainable=True, name='embed')
callback = tf.keras.callbacks.EarlyStopping(monitor='val_mae', patience=3)
# -
embed = hub.load(handle)
embeddings = embed(["man","woman"])
embeddings
# ## Baseline
# +
epochs = 100
base_model = keras.Sequential([
layers.Input(shape=(), name="Input", dtype=tf.string),
emb_layer,
layers.Dense(10,activation='relu', dtype=tf.float32),
layers.Dropout(0.2),
layers.Dense(1,activation=relu_advanced)],name='2.1-Base-Reg-Swivel')
base_model.compile(loss=tf.keras.losses.MeanSquaredError(),
optimizer=keras.optimizers.Adam(lr=0.0003,decay=1e-6),
metrics=['mse','mae'])
base_history = base_model.fit(train_ds,
epochs = epochs,
validation_data=val_ds,
callbacks=[callback],
verbose=1)
# -
plot_loss(base_history,base_model)
performance_evaluation(X_test, y_test, base_model)
plot_model_path = os.path.join(IMAGES_DIR, f'plot_model_{base_model.name}.png')
tf.keras.utils.plot_model(base_model, to_file=plot_model_path ,show_shapes=True, show_dtype=True,
show_layer_names=True, rankdir='TB', expand_nested=True, dpi=96)
base_model.save(os.path.join(MODELS_DIR,base_model.name))
# + [markdown] tags=[]
# ## RNN
# -
embed
embed(X_train).shape
# +
rnn_model = tf.keras.Sequential([
layers.Input(shape=(), name="Input", dtype=tf.string),
emb_layer,
layers.Reshape(target_shape= (20,1)),
layers.Bidirectional(layers.SimpleRNN(10, activation='tanh',return_sequences=False)),
# layers.SimpleRNN(5, activation='tanh',return_sequences=False),
layers.Dense(5, activation='relu'),
# layers.Dropout(0.3),
# layers.Dense(10, activation='relu'),
# layers.Dropout(0.3),
# layers.Dense(5, activation='tanh'),
layers.Dropout(0.05),
layers.Dense(1,activation=relu_advanced)
],name='2.1-RNN-Reg-Bidirect-Swivel')
rnn_model.compile(loss=tf.keras.losses.MeanSquaredError(),
optimizer=tf.keras.optimizers.Adam(learning_rate=0.0005),
metrics=['mse','mae'])
num_epochs=100
rnn_history=rnn_model.fit(train_ds,
epochs=num_epochs,
validation_data = val_ds,
callbacks=[callback],
verbose=1)
# -
rnn_model.summary()
plot_loss(rnn_history,rnn_model)
performance_evaluation(X_test, y_test, rnn_model)
plot_model_path = os.path.join(IMAGES_DIR, f'plot_model_{rnn_model.name}.png')
tf.keras.utils.plot_model(rnn_model, to_file=plot_model_path ,show_shapes=True, show_dtype=True,
show_layer_names=True, rankdir='TB', expand_nested=True, dpi=96)
rnn_model.save(os.path.join(MODELS_DIR,rnn_model.name))
# +
# max(y_pred)
# +
gru_model = tf.keras.Sequential([
layers.Input(shape=(), name="Input", dtype=tf.string),
emb_layer,
layers.Reshape(target_shape= (20,1)),
layers.Bidirectional(layers.GRU(10, activation='tanh',return_sequences=True)),
layers.GRU(4, activation='tanh',return_sequences=False),
layers.Dropout(0.2),
# layers.Dense(5,activation='relu'),
layers.Dense(1,activation=relu_advanced)
],name='2.1-GRU-Reg-Bidirect-Swivel')
gru_model.compile(loss='mse',
optimizer=tf.keras.optimizers.Adam(learning_rate=0.001),
metrics=['mse','mae'])
num_epochs=100
gru_history=gru_model.fit(train_ds,
epochs=num_epochs,
validation_data = val_ds,
callbacks=[callback],
verbose=1)
# -
gru_model.save(os.path.join(MODELS_DIR,gru_model.name))
plot_loss(gru_history,gru_model)
performance_evaluation(X_test, y_test, gru_model)
plot_model_path = os.path.join(IMAGES_DIR, f'plot_model_{gru_model.name}.png')
tf.keras.utils.plot_model(gru_model, to_file=plot_model_path ,show_shapes=True, show_dtype=True,
show_layer_names=True, rankdir='TB', expand_nested=True, dpi=96)
# ## LSTM
# +
lstm_model = tf.keras.Sequential([
layers.Input(shape=(), name="Input", dtype=tf.string),
emb_layer,
layers.Reshape(target_shape= (20,1)),
layers.Bidirectional(layers.LSTM(10, activation='tanh',return_sequences=True)),
layers.LSTM(10, activation='tanh',return_sequences=False),
# layers.Dense(5,activation='tanh'),
layers.Dropout(0.2),
layers.Dense(1,activation=relu_advanced)
],name='2.1-LSTM-Reg-Bidirect-Swivel')
lstm_model.compile(loss='mse',
optimizer=tf.keras.optimizers.Adam(learning_rate=0.001),
metrics=['mse','mae'])
num_epochs=100
lstm_history=lstm_model.fit(train_ds,
epochs=num_epochs,
validation_data = val_ds,
callbacks=[callback],
verbose=1)
# -
performance_evaluation(X_test, y_test, lstm_model)
plot_loss(lstm_history,lstm_model)
plot_model_path = os.path.join(IMAGES_DIR, f'plot_model_{lstm_model.name}.png')
tf.keras.utils.plot_model(lstm_model, to_file=plot_model_path ,show_shapes=True, show_dtype=True,
show_layer_names=True, rankdir='TB', expand_nested=True, dpi=96)
lstm_model.save(os.path.join(MODELS_DIR,lstm_model.name))
|
notebooks/2.1-Official-Regularized-DL.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
import matplotlib.pyplot as plt
import ipywidgets as widgets
import math
from jupyterthemes import jtplot
jtplot.style()
# ## Finite case
# $x\in[M]$
M = 1000
# ### True function
_d = 10
param = np.random.uniform(size=_d) - 1/2
fmap = np.array([[(x/M)**y for y in range(_d)] for x in range(M)])
#fmap = np.random.uniform(size = (M, _d))
_f = np.matmul(fmap, param)
f = lambda x: _f[x]
plt.plot(range(M), _f)
ylim = [min(_f)-0.1, max(_f)+0.1]
# ### Approximation with d random features (no noise)
def rfmap(d):
return np.random.normal(size=(M,d))
def ols(X, y, reg=0):
A = np.matmul(X.T,X) + reg * np.eye(d)
b = np.matmul(X.T, y)
theta = np.linalg.solve(A,b)
return np.matmul(X, theta), theta
RF = rfmap(M)
err = np.zeros(M)
perr = np.zeros(M)
fhats = np.zeros((M,M))
for d in range(1, M+1):
X = RF[:, :d]
y = _f
fhat, theta = ols(X, y)
fhats[d-1] = fhat
err[d-1] = np.linalg.norm(fhat - _f, ord=1) / M
plt.plot(range(M), err)
plt.plot(range(M), 0.2/np.sqrt(range(1,M+1)))
# +
xx = range(M)
def update(d = 1):
plt.plot(xx, _f, xx, fhats[d-1])
axes = plt.gca()
axes.set_ylim(ylim)
slider = widgets.interact(update, d=widgets.IntSlider(min=1, max=M, step=1, value=1))
# -
# ### With noise
N = 900
noise = 0.001
reg = 1
S = np.random.choice(M, size=N, replace=True)
y = _f[S] + noise * np.random.normal(size=N)
RF = rfmap(M)
err = np.zeros(M)
fhats = np.zeros((M,M))
for d in range(1, M+1):
X = RF[S, :d]
fhat, theta = ols(X, y, reg)
err[d-1] = np.linalg.norm(np.matmul(RF[:, :d], theta) - _f, ord=1) / N
fhats[d-1] = np.matmul(RF[:, :d], theta)
plt.plot(range(M), err)
# +
xx = range(M)
def update(d = 1):
plt.scatter(S,y)
plt.plot(xx, fhats[d-1])
axes = plt.gca()
axes.set_ylim(ylim)
slider = widgets.interact(update, d=widgets.IntSlider(min=1, max=M, step=1, value=M))
# -
# ## Linear case
#
# $x\in\mathbb{R}^m$
def make_fun(m):
param = 2 * np.random.uniform(size=m) - 1
return lambda x : np.matmul(x, param)
#This represents functions of the form <x,c> + d
def make_rfm(d, m):
A = np.random.normal(size=(d, m))
b = np.random.normal(size=d)
return lambda X: np.matmul(A, X.T).T + b
def ls(A, b, reg=0):
return np.linalg.solve(np.matmul(A.T, A) + reg * np.eye(A.shape[-1]), np.matmul(A.T, b))
def rf_approx(X, y, d, reg=0):
rfm = make_rfm(d, X.shape[-1])
Phi = rfm(X) #Nxd
theta = ls(Phi, y, reg)
yhat = np.matmul(Phi, theta)
return yhat, rfm, theta
def gen_rank(n, m, r):
A = 2 * np.random.uniform(size=(n, r)) - 1
B = 2 * np.random.uniform(size=(r, r)) - 1
C = 2 * np.random.uniform(size=(r, m)) - 1
M = np.matmul(np.matmul(A, B), C)
assert np.linalg.matrix_rank(M) == r
return M
# ### Scalar example
# +
m = 1
N = 100
noise = 0
d = 2
f = make_fun(m)
X = 2 * np.random.uniform(size=(N, m)) - 1
y = f(X) + noise * np.random.normal(size=N)
yhat, _, _ = rf_approx(X, y, d)
xx = np.linspace(-1,1,100)
plt.scatter(X, y)
plt.plot(X, yhat)
ax = plt.gca()
ax.set_ylim([-1,1])
ax.set_xlim([-1,1])
# -
# ### Full-rank case
# +
m = 100
N = 500
noise = 0.1
reg = 0
norm_ord = 1
max_d = 2 * m
f = make_fun(m)
X = 2 * np.random.uniform(size=(N, m)) - 1
y = f(X) + noise * np.random.normal(size=N)
ls_param = ls(X, y, reg)
yls = np.matmul(X, ls_param)
lserr = np.linalg.norm(yls - y, ord=norm_ord)**norm_ord / N
lsrank = np.linalg.matrix_rank(X)
X_test = 2 * np.random.uniform(size=(N, m)) - 1
y_test = f(X_test) + noise * np.random.normal(size=N)
yls_test = np.matmul(X_test, ls_param)
test_lserr = np.linalg.norm(yls_test - y_test, ord=norm_ord)**norm_ord / N
err = []
test_err = []
ranks = []
for d in range(1, max_d+1):
yhat, rfm, theta = rf_approx(X, y, d, reg)
err.append(np.linalg.norm(y - yhat, ord=norm_ord)**norm_ord / N)
ranks.append(np.linalg.matrix_rank(rfm(X)))
#Test
yhat_test = np.matmul(rfm(X_test), theta)
test_err.append(np.linalg.norm(y_test - yhat_test, ord=norm_ord)**norm_ord / N)
xx = range(1, max_d+1)
plt.plot(xx, err)
plt.plot(xx, test_err)
plt.plot([xx[0], xx[-1]], [lserr, lserr])
plt.plot([xx[0], xx[-1]], [test_lserr, test_lserr])
plt.xlabel('d')
plt.legend(('Training error', 'Test error', 'LS', 'LS test'))
# -
xx = range(1, max_d+1)
plt.plot(xx, err)
plt.plot(xx, test_err)
plt.plot([xx[0], xx[-1]], [lserr, lserr])
plt.plot([xx[0], xx[-1]], [test_lserr, test_lserr])
plt.xlabel('d')
plt.legend(('Training error', 'Test error', 'LS', 'LS test'))
plt.gca().set_xlim((m-3, m+3))
plt.gca().set_ylim((0,err[m-4]))
print(lsrank, np.max(ranks))
plt.plot(xx, ranks)
plt.plot((xx[0], xx[-1]),(lsrank, lsrank))
plt.legend(('rank after rf', 'original rank'))
for i in range(max_d): print(i+1, err[i], test_err[i], '\n')
# ### The rank-deficient case
# +
m = 100
N = 1000
noise = 0.1
reg = 0.1
norm_ord = 1
max_d = 3*m
rank = m - 1
f = make_fun(m)
#Contexts do not span R^d
X = gen_rank(N, m, rank)
lsrank = np.linalg.matrix_rank(X)
y = f(X) + noise * np.random.normal(size=N)
ls_param = ls(X, y, reg)
yls = np.matmul(X, ls_param)
lserr = np.linalg.norm(yls - y, ord=norm_ord)**norm_ord / N
X_test = gen_rank(N, m, rank)
y_test = f(X_test) + noise * np.random.normal(size=N)
yls_test = np.matmul(X_test, ls_param)
test_lserr = np.linalg.norm(yls_test - y_test, ord=norm_ord)**norm_ord / N
err = []
test_err = []
ranks = []
for d in range(1, max_d+1):
yhat, rfm, theta = rf_approx(X, y, d, reg)
err.append(np.linalg.norm(y - yhat, ord=norm_ord)**norm_ord / N)
#print(rfm(X).shape)
ranks.append(np.linalg.matrix_rank(rfm(X)))
#Test
yhat_test = np.matmul(rfm(X_test), theta)
test_err.append(np.linalg.norm(y_test - yhat_test, ord=norm_ord)**norm_ord / N)
xx = range(1, max_d+1)
plt.plot(xx, err)
plt.plot(xx, test_err)
plt.plot([xx[0], xx[-1]], [lserr, lserr])
plt.plot([xx[0], xx[-1]], [test_lserr, test_lserr])
plt.xlabel('d')
plt.legend(('Training error', 'Test error', 'LS', 'LS test'))
np.argmax(test_err)
# -
print(lsrank, np.max(ranks))
plt.plot(xx, ranks)
plt.plot((xx[0], xx[-1]),(lsrank, lsrank))
plt.legend(('rank after rf', 'original rank'))
plt.xlabel('d')
# ## Playing with perturbed matrices
# +
n = 2000
m = 1000
r = 1
A = gen_rank(n,m,r)
std = 1
B = A + np.random.normal(size=(n, m), scale=std)
print(np.linalg.matrix_rank(B))
C = A + np.random.normal(size=m, scale=std)
print(np.linalg.matrix_rank(C))
print('RFN:')
D = A
rank = np.linalg.matrix_rank(D)
ranks = [rank]
print('%d/%d' % (rank, min(m, n)))
max_layers = 100
activation = np.cos
#activation = lambda X: np.maximum(X, 0)
#activation = lambda X: X**2 / np.linalg.norm(X**2, 2)
#activation = lambda X: 0.01 * X**2
while(np.linalg.matrix_rank(D) < min(n, m) and len(ranks) < max_layers):
W = np.random.normal(size=(n,n), scale=std)
b = np.random.normal(size=m, scale=std)
D = np.matmul(W, D) + b
D = activation(D)
rank = np.linalg.matrix_rank(D)
ranks.append(rank)
print('%d/%d' % (rank, min(m, n)))
plt.plot(range(len(ranks)), ranks)
# -
# ## Fourier random features
# ### Gaussian kernel
#This represents functions in the RKHS of the Gaussian kernel k(x,x') = exp(-gamma||x-x'||^2)
def make_fourier_rfm(d, m):
W = np.random.normal(size=(m, d))
b = np.random.uniform(low=0, high=2*math.pi, size=d)
return lambda X: np.cos(np.matmul(X, W) + b)
# +
def ls(A, b, reg=0):
return np.linalg.solve(np.matmul(A.T, A) + reg * np.eye(A.shape[-1]), np.matmul(A.T, b))
def rf_fit(X, y, rfm, reg=0):
Phi = rfm(X) #Nxd
theta = ls(Phi, y, reg)
yhat = np.matmul(Phi, theta)
return yhat, theta, lambda Z: np.matmul(rfm(np.expand_dims(Z, 1)), theta)
# -
# The Gaussian kernel is universal, which basically means we can represent all continuous functions (on compact sets)
#
# Let's try with polynomials (of scalar input)
# +
from numpy.polynomial.polynomial import Polynomial as Poly
domain = [-1, 1]
def make_poly(order=5):
coeff = 2 * np.random.uniform(size=order+1) - 1
poly = Poly(coeff, domain)
return lambda x: poly(x).squeeze()
# -
f = make_poly()
xx = np.linspace(domain[0], domain[1], num=1000)
plt.plot(xx, f(xx))
# +
N = 100
noise = 0.1
X = 2 * np.random.uniform(size=(N, 1)) - 1
y = f(X) + noise * np.random.normal(size=N)
# -
dmax = 20
reg = 0.1
results = []
ranks = []
for d in range(1, dmax+1):
rfm = make_fourier_rfm(d, 1)
results.append(rf_fit(X, y, rfm, reg))
ranks.append(np.linalg.matrix_rank(rfm(X)))
# +
xx = np.linspace(domain[0], domain[1], num=1000)
def update(d):
plt.scatter(X.squeeze(), y)
plt.plot(xx, results[d-1][2](xx), 'orange')
axes = plt.gca()
axes.set_xlim(domain)
axes.set_ylim(min(y) - 0.1, max(y) + 0.1)
slider = widgets.interact(update, d=widgets.IntSlider(min=1, max=dmax, step=1, value=1))
# -
# Rank of design matrix as a function of dimension
plt.plot(range(1, dmax+1), ranks)
# ### Extreme machines
def make_fourier_network(widths, input_dim):
layers = []
d_prev = input_dim
for d in widths:
W = np.random.normal(size=(d_prev, d))
b = np.random.uniform(low=0, high=2*math.pi, size=d)
layers.append({'W': W, 'b': b})
d_prev = d
def fourier_network(X):
P = X
for lay in layers:
P = np.cos(np.matmul(P, lay['W']) + lay['b'])
return P
return fourier_network
layers = 2
dmax = 20
reg = 0.1
results = []
ranks = []
for d in range(1, dmax+1):
rfm = make_fourier_network([d]*layers, 1)
results.append(rf_fit(X, y, rfm, reg))
ranks.append(np.linalg.matrix_rank(rfm(X)))
# +
xx = np.linspace(domain[0], domain[1], num=1000)
def update(d):
plt.scatter(X.squeeze(), y)
plt.plot(xx, results[d-1][2](xx), 'orange')
axes = plt.gca()
axes.set_xlim(domain)
axes.set_ylim(min(y) - 0.1, max(y) + 0.1)
slider = widgets.interact(update, d=widgets.IntSlider(min=1, max=dmax, step=1, value=1))
# -
# Rank of design matrix as a function of width
plt.plot(range(1, dmax+1), ranks)
# ### Gaussian RF for the Linear Rank-Deficient Scenario
n = 2000
d = 100
r = 1
noise = 0.1
reg = 0.1
def gen_data(n, d, r):
assert r <= d
feats = np.random.normal(size=(n, d))
proj = np.matmul(np.random.normal(size=(d,r)), np.random.normal(size=(r,d)))
feats = np.matmul(feats, proj)
feats = feats / np.max(feats)
return feats
param = 2 * np.random.uniform(size=d) - 1
X = gen_data(n, d, r)
rank = np.linalg.matrix_rank(X)
y = np.matmul(X, param) + noise * np.random.normal(size=n)
param_ls = ls(X, y, reg)
paramerr = np.linalg.norm(param_ls - param)
yhat = np.matmul(X, param_ls)
mse_train = np.linalg.norm(yhat - y, ord=2)**2 / n
X_test = gen_data(n, d, r)
y_test = np.matmul(X_test, param) + noise * np.random.normal(size=n)
yhat_test = np.matmul(X_test, param_ls)
mse_test = np.linalg.norm(yhat_test - y_test, ord=2)**2 / n
rank, mse_train, mse_test, paramerr
# +
rfn = make_fourier_network([100], 100)
yhat, param_rf, _ = rf_fit(X, y, rfn, reg)
mse_train = np.linalg.norm(yhat - y, ord=2)**2 / n
X_test = gen_data(n, d, r)
y_test = np.matmul(X_test, param) + noise * np.random.normal(size=n)
yhat_test = np.matmul(rfn(X_test), param_rf)
mse_test = np.linalg.norm(yhat_test - y_test, ord=2)**2 / n
rank = np.linalg.matrix_rank(rfn(X))
paramerr = None
rank, mse_train, mse_test, paramerr
# -
|
notebooks/Random Features.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Migrating from Spark to BigQuery via Dataproc -- Part 4
#
# * [Part 1](01_spark.ipynb): The original Spark code, now running on Dataproc (lift-and-shift).
# * [Part 2](02_gcs.ipynb): Replace HDFS by Google Cloud Storage. This enables job-specific-clusters. (cloud-native)
# * [Part 3](03_automate.ipynb): Automate everything, so that we can run in a job-specific cluster. (cloud-optimized)
# * [Part 4](04_bigquery.ipynb): Load CSV into BigQuery, use BigQuery. (modernize)
# * [Part 5](05_functions.ipynb): Using Cloud Functions, launch analysis every time there is a new file in the bucket. (serverless)
#
# ### Catch-up cell
# Catch-up cell. Run if you did not do previous notebooks of this sequence
# !wget http://kdd.ics.uci.edu/databases/kddcup99/kddcup.data_10_percent.gz
BUCKET='cloud-training-demos-ml' # CHANGE
# !gsutil cp kdd* gs://$BUCKET/
# ### Load data into BigQuery
# !bq mk sparktobq
BUCKET='cloud-training-demos-ml' # CHANGE
# !bq --location=US load --autodetect --source_format=CSV sparktobq.kdd_cup_raw gs://$BUCKET/kddcup.data_10_percent.gz
# ### BigQuery queries
#
# We can replace much of the initial exploratory code by SQL statements.
# %%bigquery
SELECT * FROM sparktobq.kdd_cup_raw LIMIT 5
# Ooops. There are no column headers. Let's fix this.
# +
# %%bigquery
CREATE OR REPLACE TABLE sparktobq.kdd_cup AS
SELECT
int64_field_0 AS duration,
string_field_1 AS protocol_type,
string_field_2 AS service,
string_field_3 AS flag,
int64_field_4 AS src_bytes,
int64_field_5 AS dst_bytes,
int64_field_6 AS wrong_fragment,
int64_field_7 AS urgent,
int64_field_8 AS hot,
int64_field_9 AS num_failed_logins,
int64_field_11 AS num_compromised,
int64_field_13 AS su_attempted,
int64_field_14 AS num_root,
int64_field_15 AS num_file_creations,
string_field_41 AS label
FROM
sparktobq.kdd_cup_raw
# -
# %%bigquery
SELECT * FROM sparktobq.kdd_cup LIMIT 5
# ### Spark analysis
#
# Replace Spark analysis by BigQuery SQL
# %%bigquery connections_by_protocol
SELECT COUNT(*) AS count
FROM sparktobq.kdd_cup
GROUP BY protocol_type
ORDER by count ASC
connections_by_protocol
# ### Spark SQL to BigQuery
#
# Pretty clean translation
# %%bigquery attack_stats
SELECT
protocol_type,
CASE label
WHEN 'normal.' THEN 'no attack'
ELSE 'attack'
END AS state,
COUNT(*) as total_freq,
ROUND(AVG(src_bytes), 2) as mean_src_bytes,
ROUND(AVG(dst_bytes), 2) as mean_dst_bytes,
ROUND(AVG(duration), 2) as mean_duration,
SUM(num_failed_logins) as total_failed_logins,
SUM(num_compromised) as total_compromised,
SUM(num_file_creations) as total_file_creations,
SUM(su_attempted) as total_root_attempts,
SUM(num_root) as total_root_acceses
FROM sparktobq.kdd_cup
GROUP BY protocol_type, state
ORDER BY 3 DESC
# %matplotlib inline
ax = attack_stats.plot.bar(x='protocol_type', subplots=True, figsize=(10,25))
# ### Write out report
#
# Copy the output to GCS so that we can safely delete the AI Platform Notebooks instance.
# +
import google.cloud.storage as gcs
# save locally
ax[0].get_figure().savefig('report.png');
connections_by_protocol.to_csv("connections_by_protocol.csv")
# upload to GCS
bucket = gcs.Client().get_bucket(BUCKET)
for blob in bucket.list_blobs(prefix='sparktobq/'):
blob.delete()
for fname in ['report.png', 'connections_by_protocol.csv']:
bucket.blob('sparktobq/{}'.format(fname)).upload_from_filename(fname)
# -
# Copyright 2019 Google Inc. Licensed under the Apache License, Version 2.0 (the "License"); you may not use this file except in compliance with the License. You may obtain a copy of the License at http://www.apache.org/licenses/LICENSE-2.0 Unless required by applicable law or agreed to in writing, software distributed under the License is distributed on an "AS IS" BASIS, WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied. See the License for the specific language governing permissions and limitations under the License.
|
quests/sparktobq/04_bigquery.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="PNMwXOr3CWjm"
from __future__ import print_function
import keras
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dense,Dropout,Activation,Flatten,BatchNormalization
from keras.layers import Conv2D,MaxPool2D
import tensorflow as tf
import os
import random
from keras import regularizers
import numpy as np
from tqdm import tqdm
import nibabel as nib
from skimage.io import imread, imshow
from skimage.transform import resize
import matplotlib.pyplot as plt
from __future__ import print_function
import keras
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dense,Dropout,Activation,Flatten,BatchNormalization
from keras.layers import Conv2D,MaxPooling2D
from tensorflow.keras import layers
# + id="LcKnoaQhJuji"
num_classes = 7
img_rows,img_cols = 48,48
batch_size = 64
# + id="mrEh8x4NmaeQ"
train_data_dir = '/content/drive/MyDrive/train'
validation_data_dir = '/content/drive/MyDrive/validation'
test_data_dir = '/content/drive/MyDrive/test (1)'
# + id="ExROJxPTqwIK"
'''train_datagen = ImageDataGenerator(
horizontal_flip=True,
fill_mode='nearest')'''
train_datagen = ImageDataGenerator(#rotation_range = 180,
width_shift_range = 0.1,
height_shift_range = 0.1,
horizontal_flip = True,
rescale = 1./255,
#zoom_range = 0.2,
validation_split = 0.2
)
validation_datagen = ImageDataGenerator(rescale = 1./255,
validation_split = 0.2)
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="FGXQyyveq1T5" outputId="30bcd907-7736-4a8f-99b6-6bec1222edfc"
'''validation_datagen = ImageDataGenerator(rescale=1./255)'''
# + colab={"base_uri": "https://localhost:8080/"} id="snNSfOIuq3rm" outputId="1866066a-420e-4828-b1dd-66113b17ef29"
'''train_generator = train_datagen.flow_from_directory(
train_data_dir,
color_mode='grayscale',
target_size=(img_rows,img_cols),
batch_size=batch_size,
class_mode='categorical',
shuffle=True,
subset='training')'''
train_generator = train_datagen.flow_from_directory(directory = train_data_dir,
target_size = (img_rows,img_cols),
batch_size = 64,
color_mode = "grayscale",
class_mode = "categorical",
subset = "training"
)
# + colab={"base_uri": "https://localhost:8080/"} id="9oV-UCEAq5qX" outputId="2efe1ee7-bef4-4096-9548-cb243f11ff1d"
'''validation_generator = validation_datagen.flow_from_directory(
validation_data_dir,
color_mode='grayscale',
target_size=(img_rows,img_cols),
batch_size=batch_size,
class_mode='categorical',
shuffle=True)'''
validation_generator = validation_datagen.flow_from_directory( directory = test_data_dir,
target_size = (img_rows,img_cols),
batch_size = 64,
color_mode = "grayscale",
class_mode = "categorical",
subset = "validation"
)
# + colab={"base_uri": "https://localhost:8080/"} id="wABs-BuVMX5j" outputId="0906fab5-c94e-45b8-c200-2eaaafb75f24"
from keras.optimizers import RMSprop,SGD,Adam
from __future__ import print_function
import keras
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dense,Dropout,Activation,Flatten,BatchNormalization
from keras.layers import Conv2D,MaxPooling2D
from tensorflow.keras import layers
model = Sequential()
model.add(Conv2D(32, kernel_size=(3, 3), padding='same', activation='relu', input_shape=(48, 48,1)))
model.add(Conv2D(64,(3,3), padding='same', activation='relu' ))
model.add(BatchNormalization())
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(128,(5,5), padding='same', activation='relu'))
model.add(BatchNormalization())
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(512,(3,3), padding='same', activation='relu', kernel_regularizer=regularizers.l2(0.01)))
model.add(BatchNormalization())
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Conv2D(512,(3,3), padding='same', activation='relu', kernel_regularizer=regularizers.l2(0.01)))
model.add(BatchNormalization())
model.add(MaxPool2D(pool_size=(2, 2)))
model.add(Dropout(0.25))
model.add(Flatten())
model.add(Dense(256,activation = 'relu'))
model.add(BatchNormalization())
model.add(Dropout(0.25))
model.add(Dense(512,activation = 'relu'))
model.add(BatchNormalization())
model.add(Dropout(0.25))
model.add(Dense(7, activation='softmax'))
model.compile(
optimizer = Adam(lr=0.0001),
loss='categorical_crossentropy',
metrics=['accuracy']
)
model.summary()
# + colab={"base_uri": "https://localhost:8080/", "height": 139} id="KFSgdNL0MC3Q" outputId="685c0ac4-db09-4316-81d3-554955de1250"
'''from __future__ import print_function
import keras
from keras.preprocessing.image import ImageDataGenerator
from keras.models import Sequential
from keras.layers import Dense,Dropout,Activation,Flatten,BatchNormalization
from keras.layers import Conv2D,MaxPooling2D
from tensorflow.keras import layers
model = keras.Sequential()
model.add(layers.Conv2D(filters=6, kernel_size=(3, 3), activation='relu', input_shape=(32,32,1)))
model.add(layers.AveragePooling2D())
model.add(layers.Conv2D(filters=16, kernel_size=(3, 3), activation='relu'))
model.add(layers.AveragePooling2D())
model.add(layers.Flatten())
model.add(layers.Dense(units=120, activation='relu'))
model.add(layers.Dense(units=84, activation='relu'))
model.add(layers.Dense(num_classes, activation = 'softmax'))
model.add(Flatten())
model.add(Dense(units=4096,activation="relu"))
model.add(Dense(units=4096,activation="relu"))
model.add(Dense(units=7, activation="sigmoid"))
model.summary()'''
# + id="JzNrRPg3rhDr"
from keras.optimizers import RMSprop,SGD,Adam
from keras.callbacks import ModelCheckpoint, EarlyStopping, ReduceLROnPlateau
# + id="E1S_-1Ggrlnt"
checkpoint = ModelCheckpoint('Emotion_weights.h5',
monitor='val_loss',
mode='min',
save_best_only=True,
verbose=1)
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="G0-O-Akyst7o" outputId="e96496b6-c387-4dc5-9ae2-4668eacdf332"
'''earlystop = EarlyStopping(monitor='val_loss',
min_delta=0,
patience=3,
verbose=1,
restore_best_weights=True
)'''
# + colab={"base_uri": "https://localhost:8080/", "height": 53} id="_tT6K9crsxI7" outputId="026aeb70-c0dc-4b32-bdc6-5f9e2b5296eb"
'''reduce_lr = ReduceLROnPlateau(monitor='val_loss',
factor=0.2,
patience=3,
verbose=1,
min_delta=0.0001)'''
# + id="K2OtDgmoszBm"
'''callbacks = [earlystop,checkpoint,reduce_lr]'''
callbacks = [checkpoint]
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="6RZKmLkUs1Tj" outputId="be922047-ed13-4770-82fc-00bd57ddd215"
'''model.compile(loss='categorical_crossentropy',
optimizer = Adam(lr=0.001),
metrics=['accuracy'])'''
# + id="DZwdTtrjs3NU"
epochs=100
# + colab={"base_uri": "https://localhost:8080/"} id="CGXXuuHes498" outputId="0e5d36f6-c80c-43b0-da11-7e385cd7dcd2"
history = model.fit(x=train_generator,epochs = epochs,validation_data = validation_generator)
# + id="uw6hW5p_Lwji"
test_datagen = ImageDataGenerator(rescale = 1./255)
# + colab={"base_uri": "https://localhost:8080/"} id="_QfOy0KtL4p0" outputId="20d64a12-bcba-4397-d4f4-eaa50cd23b6e"
test_generator = test_datagen.flow_from_directory( directory = test_data_dir,
target_size = (img_rows,img_cols),
batch_size = 64,
color_mode = "grayscale",
class_mode = "categorical",
)
# + colab={"base_uri": "https://localhost:8080/"} id="fLIcT9pRTHF9" outputId="7f9fc1e5-9d52-4d49-91f9-83b5ff378c11"
model.evaluate(test_generator)
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="zCtQtw87TJ2S" outputId="f1625b34-66d2-4d44-ebbe-54d0a00cf9fa"
import cv2,matplotlib.pyplot as plt
img=cv2.imread('/content/drive/MyDrive/test (1)/happy/im285.png')
plt.imshow(img)
# + colab={"base_uri": "https://localhost:8080/", "height": 285} id="ZXLWHC08-JSF" outputId="4eafa5bf-13c6-4c1f-fb40-d47b7aeb518c"
rgb_weights = [0.2989, 0.5870, 0.1140]
grayscale_image = np.dot(img[0:224,0:224,:3], rgb_weights)
plt.imshow(grayscale_image, cmap=plt.get_cmap("gray"))
# + colab={"base_uri": "https://localhost:8080/"} id="7HInwpMO-ZfI" outputId="a1555528-34a6-4a1c-ab12-6a493a5fc996"
print(img.shape)
print(type(img))
# + colab={"base_uri": "https://localhost:8080/"} id="qWc_YLwIjD6f" outputId="ffc147ab-7e15-47fb-a267-1988ef31c06d"
train_generator.class_indices
# + colab={"base_uri": "https://localhost:8080/"} id="Q76tJr-IP1Pj" outputId="c1489492-129a-4a10-833c-5a01e107c10e"
import cv2
def resize_fn(fp: str) -> np.ndarray:
""" Resize an image maintaining its proportions
Args:
fp (str): Path argument to image file
scale (Union[float, int]): Percent as whole number of original image. eg. 53
Returns:
image (np.ndarray): Scaled image
"""
_scale = lambda dim, s: int(dim * s / 100)
im: np.ndarray = cv2.imread(fp)
#plt.imshow(im)
width, height, channels = im.shape
scale_1=(48*100)/width
scale_2=(48*100)/height
print("{},{},{}".format(width,height,channels))
new_width: int = _scale(width, scale_1)
new_height: int = _scale(height, scale_2)
new_dim: tuple = (new_width, new_height)
return cv2.resize(src=im, dsize=new_dim, interpolation=cv2.INTER_LINEAR)
order=['angry','disgust','fear','happy','neutral','sad','surprised']
resized = resize_fn('/content/drive/MyDrive/test (1)/happy/im1724.png')
print(resized.shape)
rgb_weights = [0.2989, 0.5870, 0.1140]
grayscale_image = np.dot(resized[0:48,0:48,:3], rgb_weights)
grayscale_image = grayscale_image.reshape((48,48,1))
X = np.zeros((1,48, 48, 1), dtype=np.float32)
X[0]=grayscale_image
z=model.predict(X)
t=[x for x in range(0,7) if z[0][x]==max(max(model.predict(X)))]
print("Emotion of the person in the image is: {}".format(order[t[0]]))
emotion_detected=order[t[0]]
# + id="FVoQ6xmvTeqK"
import pathlib
for path in pathlib.Path('/content/drive/MyDrive/test (1)/happy').iterdir():
order=['angry','disgust','fear','happy','neutral','sad','surprised']
resized = resize_fn(str(path))
rgb_weights = [0.2989, 0.5870, 0.1140]
grayscale_image = np.dot(resized[0:48,0:48,:3], rgb_weights)
grayscale_image = grayscale_image.reshape((48,48,1))
X = np.zeros((1,48, 48, 1), dtype=np.float32)
X[0]=grayscale_image
z=model.predict(X)
t=[x for x in range(0,7) if z[0][x]==max(max(model.predict(X)))]
if order[t[0]]=='happy':
print("Emotion of the person in the image is: {}".format(order[t[0]]))
print(path)
break
# + colab={"base_uri": "https://localhost:8080/"} id="aPXkjCWnkpOf" outputId="0e642825-4469-4566-a4f7-13641fd14e0f"
print(emotion_detected.upper())
# + colab={"base_uri": "https://localhost:8080/"} id="LwkOmidHliYW" outputId="ce5eee7b-7147-42ab-eb65-c236ff8b5787"
# !pip install gtts
# + colab={"base_uri": "https://localhost:8080/", "height": 35} id="lSEv0VhLm2_i" outputId="edad7c35-028c-45b2-d631-7990761de36c"
df.iloc[0]['SONG']
# + colab={"base_uri": "https://localhost:8080/"} id="N68ZibxalMQr" outputId="01d7dbbf-78e9-4fbc-a50f-0b566fc30c69"
import pandas as pd
df=pd.read_csv(emotion_detected.upper()+'.csv')
print(df)
from gtts import gTTS
from IPython.display import Audio
tts = gTTS('Mood detected is: '+emotion_detected.upper()+'and, the list of suggested songs is:')
length_db=len(df)
a=[]
for i in range(0,length_db):
b=gTTS(df.iloc[i]['SONG']+' by '+df.iloc[i]['ARTIST'])
b.save('a'+str(i)+'.mp3')
sound_file = 'a'+str(i)+'.mp3'
Audio(sound_file, autoplay=True)
tts.save('audio_file.mp3')
sound_file = 'audio_file.mp3'
#Audio(sound_file, autoplay=True)
# + colab={"base_uri": "https://localhost:8080/"} id="F3690Td0poZH" outputId="b53c58c2-4f73-4f9a-e80e-ffbf7b0faaab"
# !pip install pydub
# + id="V7CPS0T6pacp"
from pydub import AudioSegment
sound=[]
sound.append(AudioSegment.from_file("/content/audio_file.mp3", format="mp3"))
for i in range(0,length_db):
a=AudioSegment.from_file("/content/a"+str(i)+".mp3", format="mp3")
sound.append(a)
# sound1, with sound2 appended (use louder instead of sound1 to append the louder version)
combined=0
for i in sound:
combined=combined+i
# simple export
file_handle = combined.export("/content/output.mp3", format="mp3")
# + [markdown] id="Tor5VndbsQhh"
# ##FOR HAPPY
# + colab={"base_uri": "https://localhost:8080/", "height": 75} id="Gj6K4uSdqz98" outputId="cec7d4c7-05e4-4416-bbc1-4c4d6066aa4a"
Audio('output.mp3', autoplay=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="EAEULfN0rizz" outputId="2ed9af0b-610b-4d7a-d651-912afcbb9b42"
df.head()
# + colab={"base_uri": "https://localhost:8080/"} id="a4r9cm1qr1Jd" outputId="ee5a6458-9aad-40eb-e57e-ebd3b4e48182"
import cv2
def resize_fn(fp: str) -> np.ndarray:
""" Resize an image maintaining its proportions
Args:
fp (str): Path argument to image file
scale (Union[float, int]): Percent as whole number of original image. eg. 53
Returns:
image (np.ndarray): Scaled image
"""
_scale = lambda dim, s: int(dim * s / 100)
im: np.ndarray = cv2.imread(fp)
#plt.imshow(im)
width, height, channels = im.shape
scale_1=(48*100)/width
scale_2=(48*100)/height
print("{},{},{}".format(width,height,channels))
new_width: int = _scale(width, scale_1)
new_height: int = _scale(height, scale_2)
new_dim: tuple = (new_width, new_height)
return cv2.resize(src=im, dsize=new_dim, interpolation=cv2.INTER_LINEAR)
order=['angry','disgust','fear','happy','neutral','sad','surprised']
resized = resize_fn('/content/drive/MyDrive/test (1)/angry/im0.png')
print(resized.shape)
rgb_weights = [0.2989, 0.5870, 0.1140]
grayscale_image = np.dot(resized[0:48,0:48,:3], rgb_weights)
grayscale_image = grayscale_image.reshape((48,48,1))
X = np.zeros((1,48, 48, 1), dtype=np.float32)
X[0]=grayscale_image
z=model.predict(X)
t=[x for x in range(0,7) if z[0][x]==max(max(model.predict(X)))]
print("Emotion of the person in the image is: {}".format(order[t[0]]))
emotion_detected=order[t[0]]
# + colab={"base_uri": "https://localhost:8080/"} id="Kb9wbKy-sBtP" outputId="792ffd72-273c-47e5-d38b-c4526dc7ed71"
import pandas as pd
df=pd.read_csv(emotion_detected.upper()+'.csv')
print(df)
from gtts import gTTS
from IPython.display import Audio
tts = gTTS('Mood detected is: '+emotion_detected.upper()+'and, the list of suggested songs is:')
length_db=len(df)
a=[]
for i in range(0,length_db):
b=gTTS(df.iloc[i]['SONG']+' by '+df.iloc[i]['ARTIST'])
b.save('a'+str(i)+'.mp3')
sound_file = 'a'+str(i)+'.mp3'
Audio(sound_file, autoplay=True)
tts.save('audio_file.mp3')
sound_file = 'audio_file.mp3'
#Audio(sound_file, autoplay=True)
# + id="d0sI8X4DsFnR"
from pydub import AudioSegment
sound=[]
sound.append(AudioSegment.from_file("/content/audio_file.mp3", format="mp3"))
for i in range(0,length_db):
a=AudioSegment.from_file("/content/a"+str(i)+".mp3", format="mp3")
sound.append(a)
# sound1, with sound2 appended (use louder instead of sound1 to append the louder version)
combined=0
for i in sound:
combined=combined+i
# simple export
file_handle = combined.export("/content/output.mp3", format="mp3")
# + [markdown] id="H4JqaRZKsTVe"
# ##FOR ANGRY. SIMILARLY, CAN DO FOR ALL THE OTHER EMOTIONS.
# + colab={"base_uri": "https://localhost:8080/", "height": 75} id="GIapjRBfsJH3" outputId="06bd41c4-28a8-45ad-a87a-0ec8e49c02b8"
Audio('output.mp3', autoplay=True)
# + colab={"base_uri": "https://localhost:8080/", "height": 206} id="vBrlh-0MsL69" outputId="ed53a8f7-c889-46fd-f00a-2c6ab4d50bce"
df.head()
|
AIprojFaceRecogCompleted.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Response Themes for "What new features or changes would you like to see in Jupyter Notebook? (Please list anything that comes to mind that helps you in your workflow, big or small.)"
#
# * Goal: Extract theme keywords from `features_changes` responses.
# * Data: Output from 2_clean_survey.ipynb notebook (`survey_short_columns.csv`)
# * Process from [1_ux_survey_review.ipynb](1_ux_survey_review.ipynb):
#
# > Moving forward, here's a semi-automatic procedure we can follow for identifying themes across questions:
# > 1. Take a random sample of question responses
# > 2. Write down common theme keywords
# > 3. Search back through the responses using the theme keywords
# > 4. Expand the set of keywords with other words seen in the search results
# > 5. Repeat for all themes and questions
# > Later, we can use a fully automated topic modeling approach to validate our manually generated themes.
import warnings
warnings.filterwarnings("ignore")
# %matplotlib inline
import pandas as pd
import numpy as np
# Make sure the samples come up the same for anyone that re-runs this.
rs = np.random.RandomState(123)
pd.set_option('max_colwidth', 1000)
df = pd.read_csv('survey_short_columns.csv')
def show(series):
'''Make random samples easier to read.'''
for i, value in enumerate(series):
print('{}) {}'.format(i, value), end='\n\n')
responses = pd.concat([df.features_changes_1, df.features_changes_2, df.features_changes_3])
assert len(responses) == len(df) * 3
responses.isnull().value_counts()
responses = responses.dropna()
# ## Initial Samples
#
# I ran the sampling code below multiple times and manually built up the initial set of keywords seen commonly across them. I formed groups of conceptually related keywords. Then I tried to assign a simple label to each group.
show(responses.sample(20, random_state=rs))
themes = {
'ux': ['drag', 'drop', 'drag(/|&)drop', '(multiple|groups of|individual) cells', 'sections',
'click(ing)?', 'paste', 'undo', 'all cells', 'copy/paste',
'todo', '(styles?|stylesheets?|CSS|theme|skin|font|colo(u)?r)', 'scrol(ling)?',
'fold(ing)?(.*cells)?', 'collaps(e|ing|ible)', 'hid(e|ing|den)',
'navigat(e|ion|ing)',
'(menus?|icons?|buttons?|tooltips?)', 'g?ui($|\W)',
'mobile', 'progress bar',
'(window|pane|sidebar|screen|width|height)', '(moving|reordering) cells', 'toggle',
'spell(\s|-)?check(er|ing)?', 'dialog', ],
'version_control': ['(source|version) (control|tracking|management)', 'git(hub)?',
'control version', 'not versioned', 'version-controlled',
'version(ing)?(\snotebooks)?', '((version|editing) history)|revisions', 'revisions/code',
'change track(er|ing)', ],
'dissemination': ['hid(e|ing) (code|cells|input)', 'slides?', 'deployment', 'nbconvert', 'nbviewer',
'dashboards?',
'report(ing)?', 'interact(ion|ive|ity)', 'presentations?', 'knitr',
'export', 'slides(how)?', 'sharing', 'publish(ing)?', 'PDF($|\W)', 'download', 'reus(e|able)', ],
'documentation': ['doc(s|umentation)?($|\W)', 'tutorials?', 'instructions?', 'how(\s|-)?tos?' ],
'ide': ['workspace|ide($|\W)|rstudio', 'variable (explorer|viewer)', 'debug', 'profil(er|ing)', 'breakpoints?',
'inspector', 'variables', 'pdb', ],
'hosting': ['cloud', 'cluster', 'jupyter(\s)?hub', 'tmpnb', 'desktop', 'S3', 'backup', 'docker',
'install', 'hosting', 'users', 'multi(ple|-)?\s?user', ],
'content_management': ['(folders?|director(y|ies))', 'file(\s)?system', '(toc|table of contents)', 'search',
'tagging', 'file (manager|browser)', 'hierarchy', 'find\s?(\/|and\s)replace',
'import(able)? (from)? notebooks?', 'import', 'refactor(ing)?', ],
'collab': ['collaborat(ion|ive|ing)', ],
'editor': ['lint(ing|er)s?', '(better|powerful|external) edit(or|ing)',
'multi(ple\s|-)cursor', '(text|code) edit(or|ing)', 'syntax', 'diffs?($|\W)',
'curser', 'vim?($|\W)|sublime|atom|emacs',
'(tab|code|auto)-?\s?complet(e|ion)', '(hot)?key(\s|-)?(mappings|bindings)', 'keyboard|short-?cuts?',
'parenthes(e|i)s', ],
'integration': ['integrat(e|ion)', '(apache\s)?spark', 'pydb', 'matplotlib', 'django',
'pandoc', 'wordpress', 'pyflakes', 'third-party', 'packages', ],
'visualization': ['d3', 'widgets', 'visualization', 'plotting', 'graphics', ],
'cranky': ['dial down the hype', ],
'performance': ['performance', 'more speed', 'cach(e|ing)', ],
'compatibility': ['communication of changes', 'package updates', 'compatib(ility|le)', ],
'language': ['python', 'language', 'clojure', 'fortran', 'javascript', 'R($|\W)', 'C\/C', ],
'cli': ['cli($|\W)', 'terminal', 'command line', 'console', ],
}
# ## Coverage Improvement
#
# I next ran the code below to associate the theme labels with the responses. I then iterated on running the code below to find reponses without labels. I expanded the list of keywords and themes above in order to improve coverage.
import re
def keywords_or(text, keywords):
for keyword in keywords:
if re.search('(^|\W+){}'.format(keyword), text, re.IGNORECASE):
return True
return False
def tag_themes(responses, themes):
tagged = responses.to_frame()
tagged['themes'] = ''
for theme, keywords in themes.items():
results = responses.map(lambda text: keywords_or(text, keywords))
tagged.loc[results, 'themes'] += theme + ','
print(theme, results.sum())
return tagged
tagged = tag_themes(responses, themes)
tagged.themes.str.count(',').value_counts()
tagged[tagged.themes.str.len() == 0].sample(20, random_state=rs)
# ## Precision Check
#
# I then studied a sample of responses for each theme to see if there major inaccuracies in their application (e.g., string matches that are too fuzzy).
tagged = tag_themes(responses, themes)
tagged.themes.str.count(',').value_counts()
from IPython.display import display, clear_output
# +
# for key in themes:
# clear_output()
# size = min([10, len(tagged[tagged.themes.str.contains(key)])])
# display(tagged[tagged.themes.str.contains(key)].sample(size))
# if input('Showing `{}`. Type Enter to continue, "q" to stop.'.format(key)) == 'q':
# break
# -
# I also looked at responses with multiple themes to fine tune them, and catch duplication of words in multiple themes.
tagged[tagged.themes.str.count(',') == 2].sample(20, random_state=rs)
# ## Keyword Frequencies
import matplotlib
import seaborn
counts = {}
for theme, keywords in themes.items():
for keyword in keywords:
hits = responses.map(lambda text: keywords_or(text, [keyword]))
counts[keyword] = hits.sum()
hist = pd.Series(counts).sort_values()
ax = hist[-30:].plot.barh(figsize=(8, 8))
_ = ax.set_xlabel('Mentions')
# ## Persist
#
# I save off the themes and keywords to a DataFrame with the same index as the original so that the entries can be tagged.
column = 'features_changes'
themes_df = tagged.themes.to_frame()
themes_df = themes_df.rename(columns={'themes' : column+'_themes'})
themes_df[column+'_keywords'] = ''
for theme, keywords in themes.items():
for keyword in keywords:
results = responses.map(lambda text: keywords_or(text, [keyword]))
themes_df.loc[results, column+'_keywords'] += keyword + ','
themes_df[column+'_themes'] = themes_df[column+'_themes'].str.rstrip(',')
themes_df[column+'_keywords'] = themes_df[column+'_keywords'].str.rstrip(',')
# Up above, I merged the three response fields for the question into one common pool which means we can have duplicate index value in the themes DataFrame. We need to squash these down and remove duplicates.
def union(group_df):
'''Gets the set union of themes and keywords for a given DataFrame.'''
themes = group_df[column+'_themes'].str.cat(sep=',')
themes = list(set(themes.split(',')))
themes = ','.join(theme for theme in themes if theme)
keywords = group_df[column+'_keywords'].str.cat(sep=',')
keywords = list(set(keywords.split(',')))
keywords = ','.join(keyword for keyword in keywords if keyword)
return pd.Series([themes, keywords], index=[column+'_themes', column+'_keywords'])
# We group by the index and union the themes and keywords.
themes_df = themes_df.groupby(themes_df.index).apply(union)
themes_df.head()
# The themes DataFrame should have as many rows as there are non-null responses in the original DataFrame.
assert len(themes_df) == len(df[[column+'_1', column+'_2', column+'_3']].dropna(how='all'))
themes_df.to_csv(column + '_themes.csv', sep=';')
|
surveys/2015-12-notebook-ux/analysis/prep/3g_features_changes_themes.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Visuals for LDIC2022 Paper
#
# Here, the visuals for the publication are created.
# Import standard libraries.
# +
# Python standard libraries
import os
import string
import pathlib
import datetime
import random
# scientific standard libraries
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
from matplotlib import gridspec
from matplotlib import patches as mpatches
from matplotlib import dates as mdates
# Jupyter notebook-related libraries
from IPython.display import Markdown
from IPython.display import Image
# -
# Import conflowgen
from conflowgen import ContainerFlowGenerationManager
from conflowgen import ModeOfTransport
from conflowgen import PortCallManager
from conflowgen import ExportFileFormat
from conflowgen import ExportContainerFlowManager
from conflowgen import DatabaseChooser
from conflowgen import setup_logger
from conflowgen import InboundAndOutboundVehicleCapacityPreviewReport
from conflowgen import ContainerFlowByVehicleTypePreviewReport
from conflowgen import VehicleCapacityExceededPreviewReport
from conflowgen import ModalSplitPreviewReport
from conflowgen import InboundAndOutboundVehicleCapacityAnalysisReport
from conflowgen import ContainerFlowByVehicleTypeAnalysisReport
from conflowgen import ModalSplitAnalysisReport
from conflowgen import ContainerFlowAdjustmentByVehicleTypeAnalysisReport
from conflowgen import ContainerFlowAdjustmentByVehicleTypeAnalysisSummaryReport
from conflowgen import ContainerLengthDistributionManager
from conflowgen import ContainerLength
from conflowgen import ModeOfTransportDistributionManager
# ## Generate data
#
# Now, conflowgen is initialized with the schedules.
# This is based on the script `demo_DEHAM_CTA.py` but is shortened.
# Some logging messages and some comments have been removed for brevity.
# For further information, turn to `demo_DEHAM_CTA.py`.
# + tags=[]
import conflowgen
seeded_random = random.Random(x=1)
import_root_dir = os.path.abspath(
os.path.join(
os.path.dirname(conflowgen.__path__[0]),
"demo",
"data"
)
)
import_deham_dir = os.path.join(
import_root_dir,
"DEHAM",
"CT Altenwerder"
)
df_deep_sea_vessels = pd.read_csv(
os.path.join(
import_deham_dir,
"deep_sea_vessel_input.csv"
),
index_col=[0]
)
df_feeders = pd.read_csv(
os.path.join(
import_deham_dir,
"feeder_input.csv"
),
index_col=[0]
)
df_barges = pd.read_csv(
os.path.join(
import_deham_dir,
"barge_input.csv"
),
index_col=[0]
)
df_trains = pd.read_csv(
os.path.join(
import_deham_dir,
"train_input.csv"
),
index_col=[0]
)
logger = setup_logger()
database_chooser = DatabaseChooser()
def initialize_conflowgen(figure_name: str, force_reload=False) -> ContainerFlowGenerationManager:
demo_file_name = f"demo_deham_cta_visual_{figure_name}.sqlite"
if demo_file_name in database_chooser.list_all_sqlite_databases():
database_chooser.load_existing_sqlite_database(demo_file_name)
if not force_reload:
container_flow_generation_manager = ContainerFlowGenerationManager()
return container_flow_generation_manager
else:
database_chooser.create_new_sqlite_database(demo_file_name)
container_flow_generation_manager = ContainerFlowGenerationManager()
container_flow_start_date = datetime.date(year=2021, month=7, day=1)
container_flow_end_date = datetime.date(year=2021, month=7, day=31)
container_flow_generation_manager.set_properties(
name="Demo DEHAM CTA Visual 1a",
start_date=container_flow_start_date,
end_date=container_flow_end_date
)
port_call_manager = PortCallManager()
for i, row in df_feeders.iterrows():
feeder_vehicle_name = row["vehicle_name"] + "-unique"
capacity = row["capacity"]
vessel_arrives_at_as_pandas_type = row["arrival (planned)"]
vessel_arrives_at_as_datetime_type = pd.to_datetime(vessel_arrives_at_as_pandas_type)
if vessel_arrives_at_as_datetime_type.date() < container_flow_start_date:
logger.info(f"Skipping feeder '{feeder_vehicle_name}' because it arrives before the start")
continue
if vessel_arrives_at_as_datetime_type.date() > container_flow_end_date:
logger.info(f"Skipping feeder '{feeder_vehicle_name}' because it arrives after the end")
continue
if port_call_manager.get_schedule(feeder_vehicle_name, vehicle_type=ModeOfTransport.feeder):
logger.info(f"Skipping feeder '{feeder_vehicle_name}' because it already exists")
continue
logger.info(f"Add feeder '{feeder_vehicle_name}' to database")
moved_capacity = int(round(capacity * seeded_random.uniform(0.3, 0.8) / 2))
port_call_manager.add_large_scheduled_vehicle(
vehicle_type=ModeOfTransport.feeder,
service_name=feeder_vehicle_name,
vehicle_arrives_at=vessel_arrives_at_as_datetime_type.date(),
vehicle_arrives_at_time=vessel_arrives_at_as_datetime_type.time(),
average_vehicle_capacity=capacity,
average_moved_capacity=moved_capacity,
vehicle_arrives_every_k_days=-1
)
logger.info("Start importing deep sea vessels...")
for i, row in df_deep_sea_vessels.iterrows():
deep_sea_vessel_vehicle_name = row["vehicle_name"] + "-unique"
capacity = row["capacity"]
vessel_arrives_at_as_pandas_type = row["arrival (planned)"]
vessel_arrives_at_as_datetime_type = pd.to_datetime(vessel_arrives_at_as_pandas_type)
if vessel_arrives_at_as_datetime_type.date() < container_flow_start_date:
logger.info(f"Skipping deep sea vessel '{deep_sea_vessel_vehicle_name}' because it arrives before the start")
continue
if vessel_arrives_at_as_datetime_type.date() > container_flow_end_date:
logger.info(f"Skipping deep sea vessel '{deep_sea_vessel_vehicle_name}' because it arrives after the end")
continue
if port_call_manager.get_schedule(deep_sea_vessel_vehicle_name, vehicle_type=ModeOfTransport.deep_sea_vessel):
logger.info(f"Skipping deep sea service '{deep_sea_vessel_vehicle_name}' because it already exists")
continue
logger.info(f"Add deep sea vessel '{deep_sea_vessel_vehicle_name}' to database")
moved_capacity = int(round(capacity * seeded_random.uniform(0.25, 0.5) / 2))
port_call_manager.add_large_scheduled_vehicle(
vehicle_type=ModeOfTransport.deep_sea_vessel,
service_name=deep_sea_vessel_vehicle_name,
vehicle_arrives_at=vessel_arrives_at_as_datetime_type.date(),
vehicle_arrives_at_time=vessel_arrives_at_as_datetime_type.time(),
average_vehicle_capacity=capacity,
average_moved_capacity=moved_capacity,
vehicle_arrives_every_k_days=-1
)
logger.info("Start importing barges...")
for i, row in df_barges.iterrows():
barge_vehicle_name = row["vehicle_name"] + "-unique"
capacity = row["capacity"]
vessel_arrives_at_as_pandas_type = row["arrival (planned)"]
vessel_arrives_at_as_datetime_type = pd.to_datetime(vessel_arrives_at_as_pandas_type)
if vessel_arrives_at_as_datetime_type.date() < container_flow_start_date:
logger.info(f"Skipping barge '{barge_vehicle_name}' because it arrives before the start")
continue
if vessel_arrives_at_as_datetime_type.date() > container_flow_end_date:
logger.info(f"Skipping barge '{barge_vehicle_name}' because it arrives after the end")
continue
if port_call_manager.get_schedule(barge_vehicle_name, vehicle_type=ModeOfTransport.barge):
logger.info(f"Skipping barge '{barge_vehicle_name}' because it already exists")
continue
logger.info(f"Add barge '{barge_vehicle_name}' to database")
moved_capacity = int(round(capacity * seeded_random.uniform(0.3, 0.6)))
port_call_manager.add_large_scheduled_vehicle(
vehicle_type=ModeOfTransport.barge,
service_name=barge_vehicle_name,
vehicle_arrives_at=vessel_arrives_at_as_datetime_type.date(),
vehicle_arrives_at_time=vessel_arrives_at_as_datetime_type.time(),
average_vehicle_capacity=capacity,
average_moved_capacity=moved_capacity,
vehicle_arrives_every_k_days=-1
)
logger.info("Start importing trains...")
for i, row in df_trains.iterrows():
train_vehicle_name = row["vehicle_name"]
vessel_arrives_at_as_pandas_type = row["arrival_day"]
vessel_arrives_at_as_datetime_type = pd.to_datetime(vessel_arrives_at_as_pandas_type)
if port_call_manager.get_schedule(train_vehicle_name, vehicle_type=ModeOfTransport.train):
logger.info(f"Train service '{train_vehicle_name}' already exists")
continue
capacity = 96 # in TEU, see https://www.intermodal-info.com/verkehrstraeger/
earliest_time = datetime.time(hour=1, minute=0)
earliest_time_as_delta = datetime.timedelta(hours=earliest_time.hour, minutes=earliest_time.minute)
latest_time = datetime.time(hour=5, minute=30)
latest_time_as_delta = datetime.timedelta(hours=latest_time.hour, minutes=latest_time.minute)
number_slots_minus_one = int((latest_time_as_delta - earliest_time_as_delta) / datetime.timedelta(minutes=30))
drawn_slot = seeded_random.randint(0, number_slots_minus_one)
vehicle_arrives_at_time_as_delta = earliest_time_as_delta + datetime.timedelta(hours=0.5 * drawn_slot)
vehicle_arrives_at_time = (datetime.datetime.min + vehicle_arrives_at_time_as_delta).time()
logger.info(f"Add train '{train_vehicle_name}' to database")
port_call_manager.add_large_scheduled_vehicle(
vehicle_type=ModeOfTransport.train,
service_name=train_vehicle_name,
vehicle_arrives_at=vessel_arrives_at_as_datetime_type.date(),
vehicle_arrives_at_time=vehicle_arrives_at_time,
average_vehicle_capacity=capacity,
average_moved_capacity=capacity,
vehicle_arrives_every_k_days=7
)
return container_flow_generation_manager
# -
mode_of_transport_distribution_manager = ModeOfTransportDistributionManager()
# + tags=[]
# In this scenario, we move traffic away from feeders to deep sea vessels
container_flow_generation_manager = initialize_conflowgen("1a")
diff = 0.1
mode_of_transport_distribution_1a = {
ModeOfTransport.truck: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6) - diff,
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6) + diff
},
ModeOfTransport.train: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6) - diff,
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6) + diff
},
ModeOfTransport.barge: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6) - diff,
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6) + diff
},
ModeOfTransport.feeder: {
ModeOfTransport.truck: 0.8 / (0.8 + 1.9) * 0.502,
ModeOfTransport.train: 0.8 / (0.8 + 1.9) * 0.47,
ModeOfTransport.barge: 0.8 / (0.8 + 1.9) * 0.0028,
ModeOfTransport.feeder: 0,
ModeOfTransport.deep_sea_vessel: 1.9 / (0.8 + 1.9)
},
ModeOfTransport.deep_sea_vessel: {
ModeOfTransport.truck: 4.6 / (4.6 + 1.9) * 0.502,
ModeOfTransport.train: 4.6 / (4.6 + 1.9) * 0.47,
ModeOfTransport.barge: 4.6 / (4.6 + 1.9) * 0.0028,
ModeOfTransport.feeder: 1.9 / (4.6 + 1.9),
ModeOfTransport.deep_sea_vessel: 0
}
}
mode_of_transport_distribution_manager.set_mode_of_transport_distribution(mode_of_transport_distribution_1a)
container_flow_generation_manager.generate()
export_container_flow_manager = ExportContainerFlowManager()
folder_data_visual_1a = "demo-DEHAM-visual-1a-0-1--" + str(datetime.datetime.now()).replace(":", "-").replace(" ", "--").split(".")[0]
export_container_flow_manager.export(
folder_name=folder_data_visual_1a,
file_format=ExportFileFormat.csv
)
database_chooser.close_current_connection()
# + tags=[]
# In this scenario, we keep things as they are
container_flow_generation_manager = initialize_conflowgen("1b")
mode_of_transport_distribution_1b = {
ModeOfTransport.truck: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6),
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6)
},
ModeOfTransport.train: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6),
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6)
},
ModeOfTransport.barge: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6),
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6)
},
ModeOfTransport.feeder: {
ModeOfTransport.truck: 0.8 / (0.8 + 1.9) * 0.502,
ModeOfTransport.train: 0.8 / (0.8 + 1.9) * 0.47,
ModeOfTransport.barge: 0.8 / (0.8 + 1.9) * 0.0028,
ModeOfTransport.feeder: 0,
ModeOfTransport.deep_sea_vessel: 1.9 / (0.8 + 1.9)
},
ModeOfTransport.deep_sea_vessel: {
ModeOfTransport.truck: 4.6 / (4.6 + 1.9) * 0.502,
ModeOfTransport.train: 4.6 / (4.6 + 1.9) * 0.47,
ModeOfTransport.barge: 4.6 / (4.6 + 1.9) * 0.0028,
ModeOfTransport.feeder: 1.9 / (4.6 + 1.9),
ModeOfTransport.deep_sea_vessel: 0
}
}
mode_of_transport_distribution_manager.set_mode_of_transport_distribution(mode_of_transport_distribution_1b)
container_flow_generation_manager.generate()
export_container_flow_manager = ExportContainerFlowManager()
folder_data_visual_1b = "demo-DEHAM-visual-1b-0-1--" + str(datetime.datetime.now()).replace(":", "-").replace(" ", "--").split(".")[0]
export_container_flow_manager.export(
folder_name=folder_data_visual_1b,
file_format=ExportFileFormat.csv
)
database_chooser.close_current_connection()
# + tags=[]
container_flow_generation_manager = initialize_conflowgen("1c")
# In this scenario, we move traffic away from deep sea vessels to feeders
mode_of_transport_distribution_1c = {
ModeOfTransport.truck: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6) + diff,
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6) - diff
},
ModeOfTransport.train: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6) + diff,
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6) - diff
},
ModeOfTransport.barge: {
ModeOfTransport.truck: 0,
ModeOfTransport.train: 0,
ModeOfTransport.barge: 0,
ModeOfTransport.feeder: 0.8 / (0.8 + 4.6) + diff,
ModeOfTransport.deep_sea_vessel: 4.6 / (0.8 + 4.6) - diff
},
ModeOfTransport.feeder: {
ModeOfTransport.truck: 0.8 / (0.8 + 1.9) * 0.502,
ModeOfTransport.train: 0.8 / (0.8 + 1.9) * 0.47,
ModeOfTransport.barge: 0.8 / (0.8 + 1.9) * 0.0028,
ModeOfTransport.feeder: 0,
ModeOfTransport.deep_sea_vessel: 1.9 / (0.8 + 1.9)
},
ModeOfTransport.deep_sea_vessel: {
ModeOfTransport.truck: 4.6 / (4.6 + 1.9) * 0.502,
ModeOfTransport.train: 4.6 / (4.6 + 1.9) * 0.47,
ModeOfTransport.barge: 4.6 / (4.6 + 1.9) * 0.0028,
ModeOfTransport.feeder: 1.9 / (4.6 + 1.9),
ModeOfTransport.deep_sea_vessel: 0
}
}
mode_of_transport_distribution_manager.set_mode_of_transport_distribution(mode_of_transport_distribution_1c)
container_flow_generation_manager.generate()
export_container_flow_manager = ExportContainerFlowManager()
folder_data_visual_1c = "demo-DEHAM-visual-1c-0-1--" + str(datetime.datetime.now()).replace(":", "-").replace(" ", "--").split(".")[0]
export_container_flow_manager.export(
folder_name=folder_data_visual_1c,
file_format=ExportFileFormat.csv
)
database_chooser.close_current_connection()
# -
# ## Prepare visuals
#
# Later we will convert SVG files to EMF files.
# While Microsoft does not support SVG files in its Office suite, EMF is one option how to include vector graphics into Word or PowerPoint.
# One way to convert from SVG to EMF is by using the open source software Inkscape.
# +
if os.name == 'nt':
path_to_inkscape_executable = r"C:\Program Files\Inkscape\bin\inkscape.exe"
else:
raise RuntimeException("Add your path to your inkscape executable here")
def convert_to_emf(input_file):
output_file = "".join(input_file.split(".")[0:-1]) + ".emf"
!"{path_to_inkscape_executable}" "{input_file}" --export-filename "{output_file}"
# -
# ## Load data
# +
export_folder = os.path.abspath(
os.path.join(
os.path.dirname(conflowgen.__path__[0]),
"conflowgen",
"data",
"exports"
)
)
path_to_folder_data_visual_1a = os.path.join(
export_folder,
folder_data_visual_1a
)
path_to_folder_data_visual_1b = os.path.join(
export_folder,
folder_data_visual_1b
)
path_to_folder_data_visual_1c = os.path.join(
export_folder,
folder_data_visual_1c
)
print("Working with: ", (path_to_folder_data_visual_1a, path_to_folder_data_visual_1b, path_to_folder_data_visual_1c))
# -
# ### Load containers
# +
def load_containers(path_to_folder_data: str):
path_to_containers = os.path.join(
path_to_folder_data,
"containers.csv"
)
df_containers = pd.read_csv(path_to_containers, index_col="id", dtype={
"delivered_by_truck": "Int64",
"picked_up_by_truck": "Int64",
"delivered_by_large_scheduled_vehicle": "Int64",
"picked_up_by_large_scheduled_vehicle": "Int64"
})
return df_containers
df_containers_1a = load_containers(path_to_folder_data_visual_1a)
df_containers_1b = load_containers(path_to_folder_data_visual_1b)
df_containers_1c = load_containers(path_to_folder_data_visual_1c)
# -
df_containers_1b.groupby(by="delivered_by_large_scheduled_vehicle").count()
# ### Load vehicles adhering to a schedule
# + tags=[]
def get_scheduled_vehicle_file_paths(path_to_folder_data_visual: str):
path_to_deep_sea_vessels = os.path.join(
path_to_folder_data_visual,
"deep_sea_vessels.csv"
)
path_to_feeders = os.path.join(
path_to_folder_data_visual,
"feeders.csv"
)
path_to_barges = os.path.join(
path_to_folder_data_visual,
"barges.csv"
)
path_to_trains = os.path.join(
path_to_folder_data_visual,
"trains.csv"
)
scheduled_vehicle_file_paths = {
"deep_sea_vessels": path_to_deep_sea_vessels,
"feeders": path_to_feeders,
"barges": path_to_barges,
"trains": path_to_trains
}
for name, path in scheduled_vehicle_file_paths.items():
print("Check file exists for vehicle " + name + f" in folder {path_to_folder_data_visual}.")
assert os.path.isfile(path)
return scheduled_vehicle_file_paths
scheduled_vehicle_file_paths_1a = get_scheduled_vehicle_file_paths(path_to_folder_data_visual_1a)
scheduled_vehicle_file_paths_1b = get_scheduled_vehicle_file_paths(path_to_folder_data_visual_1b)
scheduled_vehicle_file_paths_1c = get_scheduled_vehicle_file_paths(path_to_folder_data_visual_1c)
# + tags=[]
for scheduled_vehicle_file_paths in (scheduled_vehicle_file_paths_1a, scheduled_vehicle_file_paths_1b, scheduled_vehicle_file_paths_1c):
for name, path in list(scheduled_vehicle_file_paths.items()):
print("Check file size for vehicle " + name + f" in folder {path}")
size_in_bytes = os.path.getsize(path)
if size_in_bytes <= 4:
print(" This file is empty, ignoring it in the analysis from now on")
del scheduled_vehicle_file_paths_1b[name]
else:
print("Everything is ok")
# + tags=[]
def get_scheduled_vehicle_dfs(scheduled_vehicle_file_paths, figure_name: str):
scheduled_vehicle_dfs = {
name: pd.read_csv(path, index_col=0, parse_dates=["scheduled_arrival"])
for name, path in scheduled_vehicle_file_paths.items()
}
for name, df in scheduled_vehicle_dfs.items():
display(Markdown("#### " + name.replace("_", " ").capitalize() + " " + figure_name))
scheduled_vehicle_dfs[name]["vehicle_type"] = name
display(scheduled_vehicle_dfs[name].sort_values(by="scheduled_arrival"))
return scheduled_vehicle_dfs
scheduled_vehicle_dfs_1a = get_scheduled_vehicle_dfs(scheduled_vehicle_file_paths_1a, "1a")
scheduled_vehicle_dfs_1b = get_scheduled_vehicle_dfs(scheduled_vehicle_file_paths_1b, "1b")
scheduled_vehicle_dfs_1c = get_scheduled_vehicle_dfs(scheduled_vehicle_file_paths_1c, "1c")
# + tags=[]
def prepare_df_large_scheduled_vehicle(scheduled_vehicle_dfs):
df_large_scheduled_vehicle = pd.concat(
scheduled_vehicle_dfs.values()
)
df_large_scheduled_vehicle.sort_index(inplace=True)
df_large_scheduled_vehicle.info()
return df_large_scheduled_vehicle
df_large_scheduled_vehicle_1a = prepare_df_large_scheduled_vehicle(scheduled_vehicle_dfs_1a)
df_large_scheduled_vehicle_1b = prepare_df_large_scheduled_vehicle(scheduled_vehicle_dfs_1b)
df_large_scheduled_vehicle_1c = prepare_df_large_scheduled_vehicle(scheduled_vehicle_dfs_1c)
df_large_scheduled_vehicle_1b
# -
# ### Load trucks
path_to_trucks_1a = os.path.join(
path_to_folder_data_visual_1a,
"trucks.csv"
)
assert os.path.isfile(path_to_trucks_1a)
path_to_trucks_1b = os.path.join(
path_to_folder_data_visual_1b,
"trucks.csv"
)
assert os.path.isfile(path_to_trucks_1b)
path_to_trucks_1c = os.path.join(
path_to_folder_data_visual_1c,
"trucks.csv"
)
assert os.path.isfile(path_to_trucks_1c)
# +
def load_trucks(path: str):
return pd.read_csv(
path_to_trucks_1b, index_col=0,
parse_dates=[
# Pickup
"planned_container_pickup_time_prior_berthing",
"realized_container_pickup_time",
# Delivery
"planned_container_delivery_time_at_window_start",
"realized_container_delivery_time"
])
df_truck_1a = load_trucks(path_to_trucks_1a)
df_truck_1b = load_trucks(path_to_trucks_1b)
df_truck_1c = load_trucks(path_to_trucks_1c)
df_truck_1b
# + [markdown] tags=[]
# ## Plot arrival distribution over time
# -
# Prepare data
# + tags=[]
x, y, z = [], [], []
y_axis = []
y_scaling_factor = 2
for i, (name, df) in enumerate(scheduled_vehicle_dfs_1b.items()):
y_axis.append((i/y_scaling_factor, name))
if len(df) == 0:
continue
for _, row in df.iterrows():
if row["vehicle_type"] == "trains":
continue
event = row["scheduled_arrival"]
moved_capacity = row["moved_capacity"]
x.append(event)
y.append(i / y_scaling_factor)
z.append(moved_capacity)
arrivals_and_capacity_1b = pd.DataFrame({"x": x, "y": y, "z": z})
display(arrivals_and_capacity_1b)
arrivals_and_capacity_1b.y.unique()
# -
container_deliveries_by_truck_1b = df_truck_1b.groupby(
pd.Grouper(key='realized_container_delivery_time', freq='H')
).count().fillna(0)
# + [markdown] tags=[]
# Create figure that shows the arrivals of different vehicles next to each other.
# This creates a first impression of the ramp-up and ramp-down phase.
# +
fig = plt.figure(figsize=(10, 5))
gs = gridspec.GridSpec(2, 1, height_ratios=[2, 1])
ax1 = plt.subplot(gs[0]) # the upper subplot
ax1.plot(container_deliveries_by_truck_1b["delivers_container"], color="k", linewidth=.7)
ax1.set_ylabel("Truck arrivals per hour")
ax1.set_xlabel("")
ax1.set_xlim([pd.Timestamp("2021-06-27"), pd.Timestamp(pd.Timestamp("2021-08-02"))])
ax2 = plt.subplot(gs[1], sharex=ax1) # the lower subplot
color_capacity_dots = 'dimgray'
# x-axis: time, y-axis: dummy variable for lower or upper row
moved_teu_scale_factor = 10
scatterplot = ax2.scatter(
x, y,
s=np.array(z)/moved_teu_scale_factor,
marker="o", color=color_capacity_dots, edgecolor="k"
)
myFmt = mdates.DateFormatter('%d.%m.')
ax1.xaxis.set_major_formatter(myFmt)
ax2.xaxis.set_major_formatter(myFmt)
ax2.set_yticks([0.0, .5])
ax2.set_yticklabels(["Deep sea\nvessels", "Feeders"])
ax2.set_ylim([-0.3, 0.7])
handles, labels = scatterplot.legend_elements(prop="sizes", num=4)
adjusted_labels = []
for handle, label in zip(handles, labels):
label_digits_only = "".join([c for c in label if c in string.digits])
number = int(label_digits_only)
number *= moved_teu_scale_factor
formatted_label = "$\\mathdefault{" + str(number) + "}$"
adjusted_labels.append(formatted_label)
handle.set_markerfacecolor(color_capacity_dots)
ax2.legend(
handles,
adjusted_labels,
borderpad=1.2,
labelspacing=1.2,
loc=(1.02, 0),
title="Moved TEU"
)
# Hide first date (it is in June) as it distorts the plot
ax2.set_xticks(ax2.get_xticks()[1:])
# just comment this in in case you want to save the figure
plt.savefig("relationship_truck_deliveries_and_vessel_departures.svg", bbox_inches='tight')
plt.show()
# -
# just comment this in in case you want to convert the figure from svg to emf
convert_to_emf("relationship_truck_deliveries_and_vessel_departures.svg")
# ## Plot capacity for inbound and outbound movements
# Prepare data
# +
def add_delivery_and_pickup(df_containers: pd.DataFrame, df_large_scheduled_vehicle: pd.DataFrame):
vehicle_to_teu_to_deliver = {}
vehicle_to_teu_to_pickup = {}
for i, container in df_containers.iterrows():
teu = container["length"] / 20
assert 1 <= teu <= 2.5
if container["delivered_by"] != "truck":
vehicle = container["delivered_by_large_scheduled_vehicle"]
if vehicle not in vehicle_to_teu_to_deliver.keys():
vehicle_to_teu_to_deliver[vehicle] = 0
vehicle_to_teu_to_deliver[vehicle] += teu
if container["picked_up_by"] != "truck":
vehicle = container["picked_up_by_large_scheduled_vehicle"]
if vehicle not in vehicle_to_teu_to_pickup.keys():
vehicle_to_teu_to_pickup[vehicle] = 0
vehicle_to_teu_to_pickup[vehicle] += teu
s_delivery = pd.Series(vehicle_to_teu_to_deliver)
s_pickup = pd.Series(vehicle_to_teu_to_pickup)
df_large_scheduled_vehicle["capacity_delivery"] = s_delivery
df_large_scheduled_vehicle["capacity_pickup"] = s_pickup
add_delivery_and_pickup(df_containers_1a, df_large_scheduled_vehicle_1a)
add_delivery_and_pickup(df_containers_1b, df_large_scheduled_vehicle_1b)
add_delivery_and_pickup(df_containers_1c, df_large_scheduled_vehicle_1c)
df_large_scheduled_vehicle_1b
# -
# Compare 1a, 1b, and 1c with each other.
# This is an example of how parameters need to be tuned.
# +
fig, axs = plt.subplots(
nrows=1,
ncols=3,
figsize=(10, 3.3),
sharey=True
)
colors = {
'feeders': 'navy',
'deep_sea_vessels': 'olive'
}
for title, ax, df_large_scheduled_vehicle in zip("abc", axs, (
df_large_scheduled_vehicle_1a,
df_large_scheduled_vehicle_1b,
df_large_scheduled_vehicle_1c
)):
df_large_scheduled_vehicle = df_large_scheduled_vehicle[
~df_large_scheduled_vehicle['vehicle_type'].isin(
[
"trains",
"barges"
]
)
]
ax.scatter(
df_large_scheduled_vehicle['capacity_delivery'],
df_large_scheduled_vehicle['capacity_pickup'],
c=list(df_large_scheduled_vehicle['vehicle_type'].map(colors)),
marker="."
)
ax.set_title(title)
ax.axline((0, 0), slope=1.2, color='dimgray', label='Ratio of 1:1.2 (20% buffer)')
ax.axline((0, 0), slope=1, color='lightgray', label='Ratio of 1:1')
ax.set_xlim([0, 3500])
ax.set_ylim([0, 3500])
ax.set_aspect('equal', adjustable='box')
ax.grid(color='lightgray', linestyle=':', linewidth=.5)
plt.setp(axs[:], xlabel="TEU delivered by vessel")
axs[0].set_ylabel("TEU picked up by vessel")
handles, labels = ax.get_legend_handles_labels()
for vehicle_type, color in colors.items():
vehicle_type_legend_name = vehicle_type.replace("_", " ").capitalize()
patch = plt.Line2D([], [], color=color, marker="o", linewidth=0, label=vehicle_type_legend_name)
handles.append(patch)
plt.legend(
handles=handles,
loc='lower left',
bbox_to_anchor=(-2.2, -0.4),
fancybox=True,
ncol=4
)
# just comment this in in case you want to save the figure
plt.savefig("ratio_delivered_and_picked_up_containers_comparison.svg", bbox_inches='tight')
plt.show()
# -
# just comment this in in case you want to convert the figure from svg to emf
convert_to_emf("ratio_delivered_and_picked_up_containers_comparison.svg")
#
|
example-usage/Jupyter Notebook/output data inspection/visuals_for_ldic2022_paper.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="LheK0rCt1Dyr" colab_type="text"
# # Artificial Intelligence Nanodegree
# ## Machine Translation Project
# In this notebook, sections that end with **'(IMPLEMENTATION)'** in the header indicate that the following blocks of code will require additional functionality which you must provide. Please be sure to read the instructions carefully!
#
# ## Introduction
# In this notebook, you will build a deep neural network that functions as part of an end-to-end machine translation pipeline. Your completed pipeline will accept English text as input and return the French translation.
#
# - **Preprocess** - You'll convert text to sequence of integers.
# - **Models** Create models which accepts a sequence of integers as input and returns a probability distribution over possible translations. After learning about the basic types of neural networks that are often used for machine translation, you will engage in your own investigations, to design your own model!
# - **Prediction** Run the model on English text.
# + [markdown] id="qapTBq4GPB1S" colab_type="text"
# ## Environment Preparation
# + id="uThOET4Zo4Ys" colab_type="code" colab={}
NUM_EPOCHS = 50
# + [markdown] id="A5qsbpbw5vsB" colab_type="text"
# ### Visualization
# + [markdown] id="6d7eucauK3X7" colab_type="text"
# #### Function: Plot Training History
# + id="G2dv7vdvK16w" colab_type="code" colab={}
import matplotlib.pyplot as plt
import numpy
import seaborn as sns
sns.set_style('whitegrid')
def plot_training_history(history, title=None):
fig, axes = plt.subplots(1, 2, figsize=(16,6))
fig.suptitle(title)
no = 0
axes[no].plot(history.history['loss'])
axes[no].plot(history.history['val_loss'])
axes[no].set_title('Loss')
axes[no].set_ylabel('accuracy')
axes[no].set_xlabel('epoch')
#axes[no].legend(['train', 'test'], loc='upper right')
xlim = axes[no].get_xlim()
axes[no].set_xlim(0,xlim[1])
#ylim = axes[no].set_ylim()
#axes[no].set_ylim(0,ylim[1])
axes[no].set_ylim(0,3)
no = 1
axes[no].plot(history.history['acc'])
axes[no].plot(history.history['val_acc'])
axes[no].set_title('Accuracy')
axes[no].set_ylabel('accuracy')
axes[no].set_xlabel('epoch')
axes[no].set_ylim((0,1))
xlim = axes[no].get_xlim()
axes[no].set_xlim(0,xlim[1])
axes[no].legend(['train', 'test'], loc='upper left')
plt.show()
# + [markdown] id="TYeATQh4h5km" colab_type="text"
# ### Setup to run on Google Colab
# This project is run on Google Colab. This is the procedure:
# * First download `machine_translation.ipynb` file into your Google drive and access it using [Google Colab](https://colab.research.google.com/)
# * Clone the [capstone project](https://github.com/udacity/aind2-nlp-capstone.git) from github
# * Add `ain2-nlp-capstone` directory into system path so that python can import helper files
# * Install `keras`
# + id="0JMaiEmL1IHD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 85} outputId="fd34cb91-2e08-42d9-d0db-35f1623e6310"
import os
capstone_path = "/content/aind2-nlp-capstone"
if not os.path.isdir(capstone_path):
# !git clone https://github.com/udacity/aind2-nlp-capstone.git
else:
print('Repository already cloned. Skipping...')
# + [markdown] id="ezZjRw3u4OjV" colab_type="text"
# Add the capstone directory into system path so that we can import modules (e.g. help.py) from that directory.
# + id="wqPf7SvD1yGt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 221} outputId="d9bd030d-06b7-4fc1-f58e-b915996843dc"
import sys
if not capstone_path in sys.path:
print('Adding capstone path')
sys.path.insert(0, capstone_path)
else:
print('Capstone path already exists...')
print("PATH:")
print("\n".join(sys.path))
# + id="iD5G65j12Gxv" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 119} outputId="3bbeee05-9f59-42e8-9b78-a113e2073798"
# !pip install keras
# + [markdown] id="3d2lcyyQibVP" colab_type="text"
# Colab requires `%aimport` to run on separate lines for each file to be imported
# + id="vmqhrLon1Dyt" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="6c0e2516-dc0f-4499-d1d5-4808c410a2f2"
# %load_ext autoreload
# %aimport helper
# %aimport tests
# %autoreload 1
print('Import completed')
# + [markdown] id="487Y7N3pi11P" colab_type="text"
# ### Import modules
# + id="F33Kn_kz1Dy3" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="bced0aed-0ee8-4bd7-e5d9-e991f55e7c43"
import collections
import helper
import numpy as np
import project_tests as tests
from keras.preprocessing.text import Tokenizer
from keras.preprocessing.sequence import pad_sequences
from keras.models import Model, Sequential
from keras.layers import SimpleRNN, GRU, Dense, TimeDistributed, Bidirectional, ZeroPadding1D
from keras.layers.embeddings import Embedding
from keras.optimizers import Adam
from keras.losses import sparse_categorical_crossentropy
print('Modules imported!')
# + [markdown] id="tWlP1cWz1DzI" colab_type="text"
# ### Verify access to the GPU
# The following test applies only if you expect to be using a GPU, e.g., while running in a Udacity Workspace or using an AWS instance with GPU support. Run the next cell, and verify that the device_type is "GPU".
# - If the device is not GPU & you are running from a Udacity Workspace, then save your workspace with the icon at the top, then click "enable" at the bottom of the workspace.
# - If the device is not GPU & you are running from an AWS instance, then refer to the cloud computing instructions in the classroom to verify your setup steps.
# + id="wJmcqlBw1DzK" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 306} outputId="86d0ef76-d743-4a3f-fdde-a63727de9871"
from tensorflow.python.client import device_lib
print(device_lib.list_local_devices())
# + [markdown] id="Hlm9S2e-1DzQ" colab_type="text"
# ## Dataset
# We begin by investigating the dataset that will be used to train and evaluate your pipeline. The most common datasets used for machine translation are from [WMT](http://www.statmt.org/). However, that will take a long time to train a neural network on. We'll be using a dataset we created for this project that contains a small vocabulary. You'll be able to train your model in a reasonable time with this dataset.
# ### Load Data
# The data is located in `data/small_vocab_en` and `data/small_vocab_fr`. The `small_vocab_en` file contains English sentences with their French translations in the `small_vocab_fr` file. Load the English and French data from these files from running the cell below.
# + id="eiz6IXCk1DzR" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="5c4564d9-ef68-4ded-965e-8885d62e609e"
import os
homedir = capstone_path # = '.'
# Load English data
english_sentences = helper.load_data(os.path.join(homedir, 'data/small_vocab_en'))
# Load French data
french_sentences = helper.load_data(os.path.join(homedir, 'data/small_vocab_fr'))
print('Dataset Loaded')
# + [markdown] id="L5RYMqgz1Dzc" colab_type="text"
# ### Files
# Each line in `small_vocab_en` contains an English sentence with the respective translation in each line of `small_vocab_fr`. View the first two lines from each file.
# + id="4TGwCqRU1Dzd" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 105} outputId="37bbe5ae-7986-4e9d-c215-f45520c548e0"
for sample_i in range(2):
print('small_vocab_en Line {}: {}'.format(sample_i + 1, english_sentences[sample_i]))
print('small_vocab_fr Line {}: {}'.format(sample_i + 1, french_sentences[sample_i]))
# + [markdown] id="Rs7EfT0F1Dzp" colab_type="text"
# From looking at the sentences, you can see they have been preprocessed already. The puncuations have been delimited using spaces. All the text have been converted to lowercase. This should save you some time, but the text requires more preprocessing.
# ### Vocabulary
# The complexity of the problem is determined by the complexity of the vocabulary. A more complex vocabulary is a more complex problem. Let's look at the complexity of the dataset we'll be working with.
# + id="gaGNtW_b1Dzq" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="bea9a4cc-782d-454c-b0ca-af35599767ab"
english_words_counter = collections.Counter([word for sentence in english_sentences for word in sentence.split()])
french_words_counter = collections.Counter([word for sentence in french_sentences for word in sentence.split()])
print('{:,} English words.'.format(len([word for sentence in english_sentences for word in sentence.split()])))
print('{:,} unique English words.'.format(len(english_words_counter)))
print('10 Most common words in the English dataset:')
print('"' + '" "'.join(list(zip(*english_words_counter.most_common(10)))[0]) + '"')
print()
print('{:,} French words.'.format(len([word for sentence in french_sentences for word in sentence.split()])))
print('{:,} unique French words.'.format(len(french_words_counter)))
print('10 Most common words in the French dataset:')
print('"' + '" "'.join(list(zip(*french_words_counter.most_common(10)))[0]) + '"')
# + [markdown] id="lkU9KA8O1Dzs" colab_type="text"
# For comparison, _Alice's Adventures in Wonderland_ contains 2,766 unique words of a total of 15,500 words.
# ## Preprocess
# For this project, you won't use text data as input to your model. Instead, you'll convert the text into sequences of integers using the following preprocess methods:
# 1. Tokenize the words into ids
# 2. Add padding to make all the sequences the same length.
#
# Time to start preprocessing the data...
# ### Tokenize (IMPLEMENTATION)
# For a neural network to predict on text data, it first has to be turned into data it can understand. Text data like "dog" is a sequence of ASCII character encodings. Since a neural network is a series of multiplication and addition operations, the input data needs to be number(s).
#
# We can turn each character into a number or each word into a number. These are called character and word ids, respectively. Character ids are used for character level models that generate text predictions for each character. A word level model uses word ids that generate text predictions for each word. Word level models tend to learn better, since they are lower in complexity, so we'll use those.
#
# Turn each sentence into a sequence of words ids using Keras's [`Tokenizer`](https://keras.io/preprocessing/text/#tokenizer) function. Use this function to tokenize `english_sentences` and `french_sentences` in the cell below.
#
# Running the cell will run `tokenize` on sample data and show output for debugging.
# + id="Lt4HWcJ11Dzs" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 224} outputId="96abbfd3-8c59-450a-de5a-ad2fc83f672b"
from keras.preprocessing.text import Tokenizer
def tokenize(x):
"""
Tokenize x
:param x: List of sentences/strings to be tokenized
:return: Tuple of (tokenized x data, tokenizer used to tokenize x)
"""
# TODO: Implement
tokenizer = Tokenizer()
tokenizer.fit_on_texts(x)
return tokenizer.texts_to_sequences(x), tokenizer
tests.test_tokenize(tokenize)
# Tokenize Example output
text_sentences = [
'The quick brown fox jumps over the lazy dog .',
'By Jove , my quick study of lexicography won a prize .',
'This is a short sentence .']
text_tokenized, text_tokenizer = tokenize(text_sentences)
print(text_tokenizer.word_index)
print()
for sample_i, (sent, token_sent) in enumerate(zip(text_sentences, text_tokenized)):
print('Sequence {} in x'.format(sample_i + 1))
print(' Input: {}'.format(sent))
print(' Output: {}'.format(token_sent))
# + [markdown] id="0vHR5Pfg1Dz3" colab_type="text"
# ### Padding (IMPLEMENTATION)
# When batching the sequence of word ids together, each sequence needs to be the same length. Since sentences are dynamic in length, we can add padding to the end of the sequences to make them the same length.
#
# Make sure all the English sequences have the same length and all the French sequences have the same length by adding padding to the **end** of each sequence using Keras's [`pad_sequences`](https://keras.io/preprocessing/sequence/#pad_sequences) function.
# + id="jQUMUPab1Dz4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 170} outputId="6603e51f-f715-46f7-e04d-9e6b3e4175a3"
from keras.preprocessing.sequence import pad_sequences
def pad(x, length=None):
"""
Pad x
:param x: List of sequences.
:param length: Length to pad the sequence to. If None, use length of longest sequence in x.
:return: Padded numpy array of sequences
"""
# TODO: Implement
if not length:
length = max(len(seq) for seq in x)
return pad_sequences(x, maxlen=length, dtype='int32', padding='post',
truncating='post', value=0.0)
tests.test_pad(pad)
# Pad Tokenized output
test_pad = pad(text_tokenized)
for sample_i, (token_sent, pad_sent) in enumerate(zip(text_tokenized, test_pad)):
print('Sequence {} in x'.format(sample_i + 1))
print(' Input: {}'.format(np.array(token_sent)))
print(' Output: {}'.format(pad_sent))
# + [markdown] id="V3tbQUz91D0B" colab_type="text"
# ### Preprocess Pipeline
# Your focus for this project is to build neural network architecture, so we won't ask you to create a preprocess pipeline. Instead, we've provided you with the implementation of the `preprocess` function.
# + id="E45XEn6a1D0E" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 102} outputId="a9aceb41-e5d5-46d3-fbc2-03d774400261"
def preprocess(x, y):
"""
Preprocess x and y
:param x: Feature List of sentences
:param y: Label List of sentences
:return: Tuple of (Preprocessed x, Preprocessed y, x tokenizer, y tokenizer)
"""
preprocess_x, x_tk = tokenize(x)
preprocess_y, y_tk = tokenize(y)
preprocess_x = pad(preprocess_x)
preprocess_y = pad(preprocess_y)
# Keras's sparse_categorical_crossentropy function requires the labels to be in 3 dimensions
preprocess_y = preprocess_y.reshape(*preprocess_y.shape, 1)
return preprocess_x, preprocess_y, x_tk, y_tk
preproc_english_sentences, preproc_french_sentences, english_tokenizer, french_tokenizer =\
preprocess(english_sentences, french_sentences)
max_english_sequence_length = preproc_english_sentences.shape[1]
max_french_sequence_length = preproc_french_sentences.shape[1]
english_vocab_size = len(english_tokenizer.word_index)
french_vocab_size = len(french_tokenizer.word_index)
print('Data Preprocessed')
print("Max English sentence length:", max_english_sequence_length)
print("Max French sentence length:", max_french_sequence_length)
print("English vocabulary size:", english_vocab_size)
print("French vocabulary size:", french_vocab_size)
# + [markdown] id="dAxUYNX71D0M" colab_type="text"
# ## Models
# In this section, you will experiment with various neural network architectures.
# You will begin by training four relatively simple architectures.
# - Model 1 is a simple RNN
# - Model 2 is a RNN with Embedding
# - Model 3 is a Bidirectional RNN
# - Model 4 is an optional Encoder-Decoder RNN
#
# After experimenting with the four simple architectures, you will construct a deeper architecture that is designed to outperform all four models.
# ### Ids Back to Text
# The neural network will be translating the input to words ids, which isn't the final form we want. We want the French translation. The function `logits_to_text` will bridge the gab between the logits from the neural network to the French translation. You'll be using this function to better understand the output of the neural network.
# + id="qh_CNt7a1D0M" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="fae9f9f4-a3ac-4dc5-a5f0-b6e162455e41"
def logits_to_text(logits, tokenizer):
"""
Turn logits from a neural network into text using the tokenizer
:param logits: Logits from a neural network
:param tokenizer: Keras Tokenizer fit on the labels
:return: String that represents the text of the logits
"""
index_to_words = {id: word for word, id in tokenizer.word_index.items()}
index_to_words[0] = '<PAD>'
return ' '.join([index_to_words[prediction] for prediction in np.argmax(logits, 1)])
print('`logits_to_text` function loaded.')
# + [markdown] id="1Zbb4N9M1D0N" colab_type="text"
# ### Model 1: RNN (IMPLEMENTATION)
# 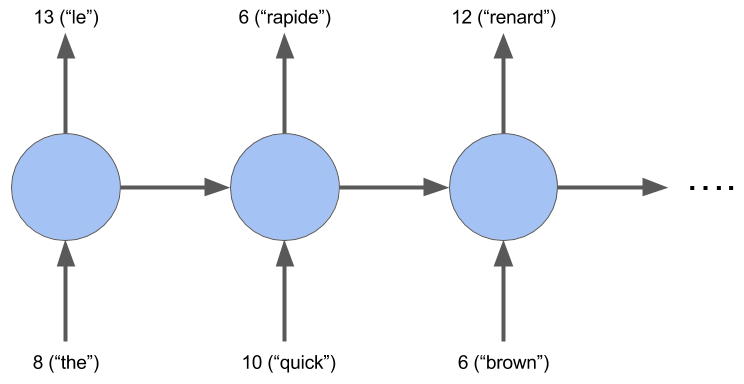
#
# A basic RNN model is a good baseline for sequence data. In this model, you'll build a RNN that translates English to French.
# + [markdown] id="25JilFo6j8nP" colab_type="text"
# #### GRU
# + id="3oRCoenckJrj" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1992} outputId="7460cb7a-aec2-4d55-cc31-1cf8a7263ef6"
def simple_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a basic RNN on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Build the layers
model = Sequential()
model.add(GRU(200, input_shape=input_shape[1:], activation="tanh",
return_sequences=True))
model.add(TimeDistributed(Dense(french_vocab_size, activation="softmax")))
learning_rate = 0.001
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(learning_rate),
metrics=['accuracy'])
return model
tests.test_simple_model(simple_model)
# Reshaping the input to work with a basic RNN
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# Train the neural network
simple_rnn_model = simple_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
print(simple_rnn_model.summary())
h = simple_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024,
epochs=NUM_EPOCHS, validation_split=0.2)
# Print prediction(s)
print(english_sentences[0])
print(logits_to_text(simple_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
# + id="q-5_Kw9Y6ijA" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="8ade540c-e66c-4ed2-fbd9-b060ac08c139"
plot_training_history(h,'Model 1: GRU')
# + [markdown] id="ByuDfriQkEYA" colab_type="text"
# #### SimpleRNN
# + id="j1h8I2Hm1D0N" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1992} outputId="f6c4af98-c649-4e07-c350-ef8492ca7af3"
def simple_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a basic RNN on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Build the layers
model = Sequential()
model.add(SimpleRNN(300, input_shape=input_shape[1:], activation = "tanh",
#dropout=0.15, recurrent_dropout=0.1,
return_sequences=True))
model.add(TimeDistributed(Dense(french_vocab_size, activation = "softmax")))
learning_rate = 0.001
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(learning_rate),
metrics=['accuracy'])
return model
tests.test_simple_model(simple_model)
# Reshaping the input to work with a basic RNN
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# Train the neural network
simple_rnn_model = simple_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
print(simple_rnn_model.summary())
h = simple_rnn_model.fit(tmp_x, preproc_french_sentences, batch_size=1024,
epochs=NUM_EPOCHS, validation_split=0.2)
# Print prediction(s)
print(english_sentences[0])
print(logits_to_text(simple_rnn_model.predict(tmp_x[:1])[0], french_tokenizer))
# + id="Np4DmRRuobxy" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="7f8811a3-4137-406a-f87b-686ee0528488"
plot_training_history(h,'Model 1: SimpleRNN')
# + [markdown] id="_PsIIEgi1D0O" colab_type="text"
# ### Model 2: Embedding (IMPLEMENTATION)
# 
#
# You've turned the words into ids, but there's a better representation of a word. This is called word embeddings. An embedding is a vector representation of the word that is close to similar words in n-dimensional space, where the n represents the size of the embedding vectors.
#
# In this model, you'll create a RNN model using embedding.
# + [markdown] id="Qh-JzHhHRw31" colab_type="text"
# #### Output Dim = 10, GRU(200)
# + id="b6DWq8ut1D0Q" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2026} outputId="563388f1-4e58-4671-e593-f8aaa303770d"
def embed_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a RNN model using word embedding on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
model = Sequential()
output_dim = 10
model.add(Embedding(input_dim=len(english_words_counter),
output_dim=output_dim,
input_length=input_shape[1]))
model.add(GRU(200, activation="tanh",
# dropout=0.2, recurrent_dropout=0.2,
return_sequences=True))
model.add(TimeDistributed(Dense(french_vocab_size, activation = "softmax")))
learning_rate = 0.001
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(learning_rate),
metrics=['accuracy'])
return model
tests.test_embed_model(embed_model)
# TODO: Reshape the input
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
#tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# TODO: Train the neural network
my_embed_model = embed_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
print(my_embed_model.summary())
h = my_embed_model.fit(tmp_x, preproc_french_sentences, batch_size=1024,
epochs=NUM_EPOCHS, validation_split=0.2)
# TODO: Print prediction(s)
print(english_sentences[0])
print(logits_to_text(my_embed_model.predict(tmp_x[:1])[0], french_tokenizer))
# + id="CC9OzaQVojE4" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="f5054566-36f2-43c1-e6f2-00c0f0e18486"
plot_training_history(h,'Model 2: Embedding & GRU')
# + [markdown] id="an9u3W2gRihh" colab_type="text"
# #### Output Dim = 100, GRU(155)
# + id="cxDSHisVOkF5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2026} outputId="948e52eb-3fc3-43f1-d0c9-b55301e71b2c"
def embed_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a RNN model using word embedding on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
model = Sequential()
output_dim = 100
model.add(Embedding(input_dim=len(english_words_counter),
output_dim=output_dim,
input_length=input_shape[1]))
model.add(GRU(155, activation="tanh",
# dropout=0.2, recurrent_dropout=0.2,
return_sequences=True))
model.add(TimeDistributed(Dense(french_vocab_size, activation = "softmax")))
learning_rate = 0.001
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(learning_rate),
metrics=['accuracy'])
return model
tests.test_embed_model(embed_model)
# TODO: Reshape the input
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
#tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# TODO: Train the neural network
my_embed_model = embed_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
print(my_embed_model.summary())
h = my_embed_model.fit(tmp_x, preproc_french_sentences, batch_size=1024,
epochs=NUM_EPOCHS, validation_split=0.2)
# TODO: Print prediction(s)
print(english_sentences[0])
print(logits_to_text(my_embed_model.predict(tmp_x[:1])[0], french_tokenizer))
# + id="eZJADVe6RQGT" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="1665f4cb-b824-4d62-964a-ec4c78e69208"
plot_training_history(h,'Model 2: Embedding & GRU - Output Dim = 100')
# + [markdown] id="LeJt-C-h1D0T" colab_type="text"
# ### Model 3: Bidirectional RNNs (IMPLEMENTATION)
# 
# One restriction of a RNN is that it can't see the future input, only the past. This is where bidirectional recurrent neural networks come in. They are able to see the future data.
# + id="W_q9twiE1D0T" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 1992} outputId="14d96a4a-0ad2-48a1-a345-66c480bc3cad"
def bd_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a bidirectional RNN model on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
model = Sequential()
model.add(Bidirectional(GRU(130, activation = "tanh", return_sequences=True),
input_shape=input_shape[1:]))
model.add(TimeDistributed(Dense(french_vocab_size, activation = "softmax")))
learning_rate = 0.001
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(learning_rate),
metrics=['accuracy'])
return model
tests.test_simple_model(bd_model)
# Reshaping the input to work with a basic RNN
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# TODO: Train and Print prediction(s)
# Train the neural network
my_bd_model = bd_model(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
print(my_bd_model.summary())
h = my_bd_model.fit(tmp_x, preproc_french_sentences, batch_size=1024,
epochs=NUM_EPOCHS, validation_split=0.2)
# Print prediction(s)
print(english_sentences[0])
print(logits_to_text(my_bd_model.predict(tmp_x[:1])[0], french_tokenizer))
# + id="srnqI0Elp0YD" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="9ad622db-d9a5-42e3-db7c-26dfdb20dbc1"
plot_training_history(h,'Model 3: Bidirectional RNN')
# + [markdown] id="9Jt3o_n81D0U" colab_type="text"
# ### Model 4: Encoder-Decoder (OPTIONAL)
# Time to look at encoder-decoder models. This model is made up of an encoder and decoder. The encoder creates a matrix representation of the sentence. The decoder takes this matrix as input and predicts the translation as output.
#
# Create an encoder-decoder model in the cell below.
# + id="lZMIKHal1D0V" colab_type="code" colab={}
def encdec_model(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train an encoder-decoder model on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# OPTIONAL: Implement
return None
tests.test_encdec_model(encdec_model)
# OPTIONAL: Train and Print prediction(s)
# + [markdown] id="lg167Giw1D0X" colab_type="text"
# ### Model 5: Custom (IMPLEMENTATION)
# Use everything you learned from the previous models to create a model that incorporates embedding and a bidirectional rnn into one model.
# + id="FrP9jw1o1D0Z" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2094} outputId="438da292-aee0-480b-aa1c-364223e62419"
def model_final(input_shape, output_sequence_length, english_vocab_size, french_vocab_size):
"""
Build and train a model that incorporates embedding, encoder-decoder, and bidirectional RNN on x and y
:param input_shape: Tuple of input shape
:param output_sequence_length: Length of output sequence
:param english_vocab_size: Number of unique English words in the dataset
:param french_vocab_size: Number of unique French words in the dataset
:return: Keras model built, but not trained
"""
# TODO: Implement
model = Sequential()
output_dim = 10
model.add(Embedding(input_dim=len(english_words_counter),
output_dim=output_dim,
input_length=input_shape[1]
))
# The final prediction problem requires different input/output sequence lengths.
# Input sequence is pad with respect to English (15 in this case) but output
# French sentence needs to be of length 21. We would not have a problem
# when we use encoder-decoder implementation but for our case we need to
# ensure equal input-output sequence length by zero padding appropriately.
model.add(ZeroPadding1D((0,output_sequence_length-input_shape[1])))
model.add(Bidirectional(GRU(125, activation="tanh",
return_sequences=True)))
model.add(TimeDistributed(Dense(french_vocab_size, activation = "softmax")))
learning_rate = 0.001
model.compile(loss=sparse_categorical_crossentropy,
optimizer=Adam(learning_rate),
metrics=['accuracy'])
return model
tests.test_embed_model(model_final)
# TODO: Reshape the input
tmp_x = pad(preproc_english_sentences, max_french_sequence_length)
#tmp_x = tmp_x.reshape((-1, preproc_french_sentences.shape[-2], 1))
# TODO: Train the neural network
my_model_final = model_final(
tmp_x.shape,
max_french_sequence_length,
english_vocab_size,
french_vocab_size)
print('Final Model Loaded')
print(my_model_final.summary())
print('Input shape = {}'.format(tmp_x.shape))
# TODO: Train the final model
h = my_model_final.fit(tmp_x, preproc_french_sentences, batch_size=1024,
epochs=NUM_EPOCHS, validation_split=0.2)
# Print prediction(s)
print(english_sentences[0])
print(logits_to_text(my_model_final.predict(tmp_x[:1])[0], french_tokenizer))
# + id="3XMgXG2Jp9Bu" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="fc70c8fa-d1ee-4e78-c051-1de1797e4bfe"
plot_training_history(h,'Model 5: Custom (Embedded & Bidirectional RNN)')
# + [markdown] id="-8y24uGj1D0a" colab_type="text"
# ## Prediction (IMPLEMENTATION)
# + id="4ZLPoJgJ1D0b" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 2128} outputId="0d37bd67-9c94-4621-9bd4-dde80e930261"
def final_predictions(x, y, x_tk, y_tk):
"""
Gets predictions using the final model
:param x: Preprocessed English data
:param y: Preprocessed French data
:param x_tk: English tokenizer
:param y_tk: French tokenizer
"""
# TODO: Train neural network using model_final
model = model_final(x.shape, y.shape[1],
len(x_tk.word_index),
len(y_tk.word_index))
print(model.summary())
h = model.fit(x, y, batch_size=1024, epochs=NUM_EPOCHS, validation_split=0.2)
## DON'T EDIT ANYTHING BELOW THIS LINE
y_id_to_word = {value: key for key, value in y_tk.word_index.items()}
y_id_to_word[0] = '<PAD>'
sentence = 'he saw a old yellow truck'
sentence = [x_tk.word_index[word] for word in sentence.split()]
sentence = pad_sequences([sentence], maxlen=x.shape[-1], padding='post')
sentences = np.array([sentence[0], x[0]])
predictions = model.predict(sentences, len(sentences))
print('Sample 1:')
print(' '.join([y_id_to_word[np.argmax(x)] for x in predictions[0]]))
print('Il a vu un vieux camion jaune')
print('Sample 2:')
print(' '.join([y_id_to_word[np.argmax(x)] for x in predictions[1]]))
print(' '.join([y_id_to_word[np.max(x)] for x in y[0]]))
## Returning history for visualization
return h
h = final_predictions(preproc_english_sentences, preproc_french_sentences, english_tokenizer, french_tokenizer)
# + id="ans2kXqAAxn5" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 431} outputId="bfe10b65-48da-4f9a-8a52-7c5a3231222b"
plot_training_history(h,'Prediction')
# + [markdown] id="Fq2pXOGn1D0h" colab_type="text"
# ## Submission
# When you're ready to submit, complete the following steps:
# 1. Review the [rubric](https://review.udacity.com/#!/rubrics/1004/view) to ensure your submission meets all requirements to pass
# 2. Generate an HTML version of this notebook
#
# - Run the next cell to attempt automatic generation (this is the recommended method in Workspaces)
# - Navigate to **FILE -> Download as -> HTML (.html)**
# - Manually generate a copy using `nbconvert` from your shell terminal
# ```
# $ pip install nbconvert
# $ python -m nbconvert machine_translation.ipynb
# ```
#
# 3. Submit the project
#
# - If you are in a Workspace, simply click the "Submit Project" button (bottom towards the right)
#
# - Otherwise, add the following files into a zip archive and submit them
# - `helper.py`
# - `machine_translation.ipynb`
# - `machine_translation.html`
# - You can export the notebook by navigating to **File -> Download as -> HTML (.html)**.
# + id="8NJMPtB91D0h" colab_type="code" colab={}
!!python -m nbconvert *.ipynb
# + [markdown] id="E2k6KYOy1D0j" colab_type="text"
# ## Optional Enhancements
#
# This project focuses on learning various network architectures for machine translation, but we don't evaluate the models according to best practices by splitting the data into separate test & training sets -- so the model accuracy is overstated. Use the [`sklearn.model_selection.train_test_split()`](http://scikit-learn.org/stable/modules/generated/sklearn.model_selection.train_test_split.html) function to create separate training & test datasets, then retrain each of the models using only the training set and evaluate the prediction accuracy using the hold out test set. Does the "best" model change?
# + [markdown] id="6Ib6Elk5hwaB" colab_type="text"
# **RESPONSE:** I already ran the model fit function using 20% validation set. The train and test sets accuracy are similar, which indicates a good generalization of the model. All model hyper-parameters are chosen such that number of total trainable parameters are similar (around 190,000) and all models ran over 50 epochs.
#
# Best model (Model 5: Custom) uses embeddings with an output dimension of 10 and bidrectional RNN (specifically GRU). Its overall accuracy is close to 96%. Even though I have not checked it out with detail some of the erronous translations might still be synonyms and hence not exactly an error. However that type of analysis requires digging down the error cases with a good understanding of French language.
|
machine_translation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Generating counterfactual explanations with any ML model
# The goal of this notebook is to show how to generate CFs for ML models using frameworks other than TensorFlow or PyTorch. This is a work in progress and here we show a method to generate diverse CFs by three methods:
# 1. Independent random sampling of features
# 2. Genetic algorithm
# 3. Querying a KD tree
#
# We use scikit-learn models for demonstration.
# # 1. Independent random sampling of features
# %load_ext autoreload
# %autoreload 2
# +
# import DiCE
import dice_ml
from dice_ml.utils import helpers # helper functions
import numpy as np
import pandas as pd
from sklearn.neural_network import MLPClassifier
from sklearn.metrics import classification_report, accuracy_score
from sklearn.pipeline import Pipeline
from sklearn.preprocessing import StandardScaler, OneHotEncoder
from sklearn.ensemble import RandomForestClassifier
# -
# ## Loading dataset
# We use the "adult" income dataset from UCI Machine Learning Repository (https://archive.ics.uci.edu/ml/datasets/adult). For demonstration purposes, we transform the data as described in dice_ml.utils.helpers module.
dataset = helpers.load_adult_income_dataset()
dataset.head()
d = dice_ml.Data(dataframe=dataset, continuous_features=['age', 'hours_per_week'], outcome_name='income')
# ## Training a custom ML model
# Below, we build an ML model using scikit-learn to demonstrate how our methods can work with any sklearn model.
# +
target = dataset["income"]
# Split data into train and test
from sklearn.model_selection import train_test_split
datasetX = dataset.drop("income", axis=1)
x_train, x_test, y_train, y_test = train_test_split(datasetX,
target,
test_size = 0.2,
random_state=0,
stratify=target)
numerical=["age", "hours_per_week"]
categorical = x_train.columns.difference(numerical)
from sklearn.compose import ColumnTransformer
# We create the preprocessing pipelines for both numeric and categorical data.
numeric_transformer = Pipeline(steps=[
('scaler', StandardScaler())])
categorical_transformer = Pipeline(steps=[
('onehot', OneHotEncoder(handle_unknown='ignore'))])
transformations = ColumnTransformer(
transformers=[
('num', numeric_transformer, numerical),
('cat', categorical_transformer, categorical)])
# Append classifier to preprocessing pipeline.
# Now we have a full prediction pipeline.
clf = Pipeline(steps=[('preprocessor', transformations),
('classifier', RandomForestClassifier())])
model = clf.fit(x_train, y_train)
# -
# provide the trained ML model to DiCE's model object
backend = 'sklearn'
m = dice_ml.Model(model=model, backend=backend)
# ## Generate diverse counterfactuals
# initiate DiCE
exp_random = dice_ml.Dice(d, m, method="random")
query_instances = x_train[4:6]
# generate counterfactuals
dice_exp_random = exp_random.generate_counterfactuals(query_instances, total_CFs=2, desired_class="opposite", verbose=False)
dice_exp_random.visualize_as_dataframe(show_only_changes=True)
# It can be observed that the random sampling method produces less sparse CFs in contrast to current DiCE's implementation. The sparsity issue with random sampling worsens with increasing *total_CFs*
# Further, different sets of counterfactuals can be generated with different random seeds.
# generate counterfactuals
dice_exp_random = exp_random.generate_counterfactuals(query_instances, total_CFs=4, desired_class="opposite", random_seed=9) # default ranomd see is 17
dice_exp_random.visualize_as_dataframe(show_only_changes=True)
# ### Selecting the features to vary
# Here, you can ensure that DiCE varies only features that it makes sense to vary.
# generate counterfactuals
dice_exp_random = exp_random.generate_counterfactuals(query_instances, total_CFs=4, desired_class="opposite",
features_to_vary=['workclass','education','occupation','hours_per_week'])
dice_exp_random.visualize_as_dataframe(show_only_changes=True)
# ### Choosing feature ranges
# Since the features are sampled randomly, they can freely vary across their range. In the below example, we show how range of continuous features can be controlled using *permitted_range* parameter that can now be passed during CF generation.
# generate counterfactuals
dice_exp_random = exp_random.generate_counterfactuals(query_instances, total_CFs=4, desired_class="opposite",
permitted_range={'age':[22,50],'hours_per_week':[40,60]})
dice_exp_random.visualize_as_dataframe(show_only_changes=True)
# # 2. Genetic Algorithm
# Here, we show how to use DiCE can be used to generate CFs for any ML model by using the genetic algorithm to find the best counterfactuals close to the query point. The genetic algorithm converges quickly, and promotes diverse counterfactuals.
# ## Training a custom ML model
# Currently, the genetic algorithm method works with scikit-learn models. We will use the same model as shown previously in the notebook. Support for Tensorflow 1&2 and Pytorch will be implemented soon.
# ## Generate diverse counterfactuals
# initiate DiceGenetic
exp_genetic = dice_ml.Dice(d, m, method='genetic')
# generate counterfactuals
dice_exp_genetic = exp_genetic.generate_counterfactuals(query_instances, total_CFs=4, desired_class=0, verbose=True)
dice_exp_genetic.visualize_as_dataframe(show_only_changes=True)
# We can also ensure that the genetic algorithm also only varies the features that you wish to vary
# generate counterfactuals
dice_exp_genetic = exp_genetic.generate_counterfactuals(query_instances, total_CFs=2, desired_class=0,
features_to_vary=['workclass','education','occupation','hours_per_week'])
dice_exp_genetic.visualize_as_dataframe(show_only_changes=True)
# You can also constrain the features to vary only within the permitted range
# generate counterfactuals
dice_exp_genetic = exp_genetic.generate_counterfactuals(query_instances, total_CFs=2, desired_class=0,
permitted_range={'age':[22,50],'hours_per_week':[40,60]})
dice_exp_genetic.visualize_as_dataframe(show_only_changes=True)
# # 3. Querying a KD Tree
# Here, we show how to use DiCE can be used to generate CFs for any ML model by finding the closest points in the dataset that give the output as the desired class. We do this efficiently by building KD trees for each class, and querying the KD tree of the desired class to find the k closest counterfactuals from the dataset. The idea behind finding the closest points from the training data itself is to ensure that the counterfactuals displayed are feasible.
# ## Training a custom ML model
# Currently, the KD tree algorithm method works with scikit-learn models. Again, we will use the same model as shown previously in the notebook. Support for Tensorflow 1&2 and Pytorch will be implemented soon.
# ## Generate diverse counterfactuals
# initiate DiceKD
exp_KD = dice_ml.Dice(d, m, method='kdtree')
# generate counterfactuals
dice_exp_KD = exp_KD.generate_counterfactuals(query_instances, total_CFs=4, desired_class="opposite")
dice_exp_KD.visualize_as_dataframe(show_only_changes=True)
# ### Selecting the features to vary
# Here, again, you can vary only features that you wish to vary. Please note that the output counterfactuals are only from the training data. If you want other counterfactuals, please use the random or genetic method.
# generate counterfactuals
dice_exp_KD = exp_KD.generate_counterfactuals(query_instances, total_CFs=4, desired_class="opposite",
features_to_vary=['age', 'workclass','education','occupation','hours_per_week'])
dice_exp_KD.visualize_as_dataframe(show_only_changes=True)
# ### Selecting the feature ranges
# Here, you can control the ranges of continuous features.
# generate counterfactuals
dice_exp_KD = exp_KD.generate_counterfactuals(query_instances, total_CFs=5, desired_class="opposite",
permitted_range={'age':[30,50],'hours_per_week':[40,60]})
dice_exp_KD.visualize_as_dataframe(show_only_changes=True)
|
docs/source/notebooks/DiCE_model_agnostic_CFs.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="rwxGnsA92emp"
# ##### Copyright 2018 The TensorFlow Authors.
# + cellView="form" colab_type="code" id="CPII1rGR2rF9" colab={}
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="JtEZ1pCPn--z"
# # 自定义训练: 演示
# + [markdown] colab_type="text" id="GV1F7tVTN3Dn"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://tensorflow.google.cn/tutorials/customization/custom_training_walkthrough"><img src="https://tensorflow.google.cn/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/zh-cn/tutorials/customization/custom_training_walkthrough.ipynb"><img src="https://tensorflow.google.cn/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/zh-cn/tutorials/customization/custom_training_walkthrough.ipynb"><img src="https://tensorflow.google.cn/images/GitHub-Mark-32px.png" />View source on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/zh-cn/tutorials/customization/custom_training_walkthrough.ipynb"><img src="https://tensorflow.google.cn/images/download_logo_32px.png" />Download notebook</a>
# </td>
# </table>
# + [markdown] colab_type="text" id="LDrzLFXE8T1l"
# 这个教程将利用机器学习的手段来对鸢尾花按照物种进行分类。本教程将利用 TensorFlow 来进行以下操作:
# 1. 构建一个模型,
# 2. 用样例数据集对模型进行训练,以及
# 3. 利用该模型对未知数据进行预测。
#
# ## TensorFlow 编程
#
# 本指南采用了以下高级 TensorFlow 概念:
#
# * 使用 TensorFlow 默认的 [eager execution](https://www.tensorflow.org/guide/eager) 开发环境,
# * 使用 [Datasets API](https://www.tensorflow.org/guide/datasets) 导入数据,
# * 使用 TensorFlow 的 [Keras API](https://keras.io/getting-started/sequential-model-guide/) 来构建各层以及整个模型。
#
# 本教程的结构同很多 TensorFlow 程序相似:
#
# 1. 数据集的导入与解析
# 2. 选择模型类型
# 3. 对模型进行训练
# 4. 评估模型效果
# 5. 使用训练过的模型进行预测
# + [markdown] colab_type="text" id="yNr7H-AIoLOR"
# ## 环境的搭建
# + [markdown] colab_type="text" id="1J3AuPBT9gyR"
# ### 配置导入
#
# 导入 TensorFlow 以及其他需要的 Python 库。 默认情况下,TensorFlow 用 [eager execution](https://www.tensorflow.org/guide/eager) 来实时评估操作, 返回具体值而不是建立一个稍后执行的[计算图](https://www.tensorflow.org/guide/graphs)。 如果您习惯使用 REPL 或 python 交互控制台, 对此您会感觉得心应手。
# + colab_type="code" id="jElLULrDhQZR" colab={}
from __future__ import absolute_import, division, print_function, unicode_literals
import os
import matplotlib.pyplot as plt
# + colab_type="code" id="bfV2Dai0Ow2o" colab={}
try:
# Colab only
# %tensorflow_version 2.x
except Exception:
pass
import tensorflow as tf
# + colab_type="code" id="g4Wzg69bnwK2" colab={}
print("TensorFlow version: {}".format(tf.__version__))
print("Eager execution: {}".format(tf.executing_eagerly()))
# + [markdown] colab_type="text" id="Zx7wc0LuuxaJ"
# ## 鸢尾花分类问题
#
# 想象一下,您是一名植物学家,正在寻找一种能够对所发现的每株鸢尾花进行自动归类的方法。机器学习可提供多种从统计学上分类花卉的算法。例如,一个复杂的机器学习程序可以根据照片对花卉进行分类。我们的要求并不高 - 我们将根据鸢尾花花萼和花瓣的长度和宽度对其进行分类。
#
# 鸢尾属约有 300 个品种,但我们的程序将仅对下列三个品种进行分类:
#
# * 山鸢尾
# * 维吉尼亚鸢尾
# * 变色鸢尾
#
# <table>
# <tr><td>
# <img src="https://www.tensorflow.org/images/iris_three_species.jpg"
# alt="Petal geometry compared for three iris species: Iris setosa, Iris virginica, and Iris versicolor">
# </td></tr>
# <tr><td align="center">
# <b>Figure 1.</b> <a href="https://commons.wikimedia.org/w/index.php?curid=170298">山鸢尾</a> (by <a href="https://commons.wikimedia.org/wiki/User:Radomil">Radomil</a>, CC BY-SA 3.0), <a href="https://commons.wikimedia.org/w/index.php?curid=248095">变色鸢尾</a>, (by <a href="https://commons.wikimedia.org/wiki/User:Dlanglois">Dlanglois</a>, CC BY-SA 3.0), and <a href="https://www.flickr.com/photos/33397993@N05/3352169862">维吉尼亚鸢尾</a> (by <a href="https://www.flickr.com/photos/33397993@N05"><NAME></a>, CC BY-SA 2.0).<br/>
# </td></tr>
# </table>
#
# 幸运的是,有人已经创建了一个包含有花萼和花瓣的测量值的[120 株鸢尾花的数据集](https://en.wikipedia.org/wiki/Iris_flower_data_set)。这是一个在入门级机器学习分类问题中经常使用的经典数据集。
# + [markdown] colab_type="text" id="3Px6KAg0Jowz"
# ## 导入和解析训练数据集
#
# 下载数据集文件并将其转换为可供此 Python 程序使用的结构。
#
# ### 下载数据集
#
# 使用 [tf.keras.utils.get_file](https://www.tensorflow.org/api_docs/python/tf/keras/utils/get_file) 函数下载训练数据集文件。该函数会返回下载文件的文件路径:
# + colab_type="code" id="J6c7uEU9rjRM" colab={}
train_dataset_url = "https://storage.googleapis.com/download.tensorflow.org/data/iris_training.csv"
train_dataset_fp = tf.keras.utils.get_file(fname=os.path.basename(train_dataset_url),
origin=train_dataset_url)
print("Local copy of the dataset file: {}".format(train_dataset_fp))
# + [markdown] colab_type="text" id="qnX1-aLors4S"
# ### 检查数据
#
# 数据集 `iris_training.csv` 是一个纯文本文件,其中存储了逗号分隔值 (CSV) 格式的表格式数据.请使用 `head -n5` 命令查看前 5 个条目:
# + colab_type="code" id="FQvb_JYdrpPm" colab={}
# !head -n5 {train_dataset_fp}
# + [markdown] colab_type="text" id="kQhzD6P-uBoq"
# 我们可以从该数据集视图中注意到以下信息:
# 1. 第一行是表头,其中包含数据集信息:
#
# * 共有 120 个样本。每个样本都有四个特征和一个标签名称,标签名称有三种可能。
# 2. 后面的行是数据记录,每个[样本](https://developers.google.com/machine-learning/glossary/#example)各占一行,其中:
# * 前四个字段是[特征](https://developers.google.com/machine-learning/glossary/#feature): 这四个字段代表的是样本的特点。在此数据集中,这些字段存储的是代表花卉测量值的浮点数。
# * 最后一列是[标签](https://developers.google.com/machine-learning/glossary/#label):即我们想要预测的值。对于此数据集,该值为 0、1 或 2 中的某个整数值(每个值分别对应一个花卉名称)。
#
# 我们用代码表示出来:
# + colab_type="code" id="9Edhevw7exl6" colab={}
# CSV文件中列的顺序
column_names = ['sepal_length', 'sepal_width', 'petal_length', 'petal_width', 'species']
feature_names = column_names[:-1]
label_name = column_names[-1]
print("Features: {}".format(feature_names))
print("Label: {}".format(label_name))
# + [markdown] colab_type="text" id="CCtwLoJhhDNc"
# 每个标签都分别与一个字符串名称(例如 “setosa” )相关联,但机器学习通常依赖于数字值。标签编号会映射到一个指定的表示法,例如:
#
# * `0` : 山鸢尾
# * `1` : 变色鸢尾
# * `2` : 维吉尼亚鸢尾
#
# 如需详细了解特征和标签,请参阅 [《机器学习速成课程》的“机器学习术语”部分](https://developers.google.com/machine-learning/crash-course/framing/ml-terminology).
# + colab_type="code" id="sVNlJlUOhkoX" colab={}
class_names = ['Iris setosa', 'Iris versicolor', 'Iris virginica']
# + [markdown] colab_type="text" id="dqPkQExM2Pwt"
# ### 创建一个 `tf.data.Dataset`
#
# TensorFlow的 [Dataset API](https://www.tensorflow.org/guide/datasets) 可处理在向模型加载数据时遇到的许多常见情况。这是一种高阶 API ,用于读取数据并将其转换为可供训练使用的格式。如需了解详情,请参阅[数据集快速入门指南](https://www.tensorflow.org/get_started/datasets_quickstart)
#
#
# 由于数据集是 CSV 格式的文本文件,请使用 [make_csv_dataset](https://www.tensorflow.org/api_docs/python/tf/data/experimental/make_csv_dataset) 函数将数据解析为合适的格式。由于此函数为训练模型生成数据,默认行为是对数据进行随机处理 (`shuffle=True, shuffle_buffer_size=10000`),并且无限期重复数据集(`num_epochs=None`)。 我们还设置了 [batch_size](https://developers.google.com/machine-learning/glossary/#batch_size) 参数:
# + colab_type="code" id="WsxHnz1ebJ2S" colab={}
batch_size = 32
train_dataset = tf.data.experimental.make_csv_dataset(
train_dataset_fp,
batch_size,
column_names=column_names,
label_name=label_name,
num_epochs=1)
# + [markdown] colab_type="text" id="gB_RSn62c-3G"
# `make_csv_dataset` 返回一个`(features, label)` 对构建的 `tf.data.Dataset` ,其中 `features` 是一个字典: `{'feature_name': value}`
#
# 这些 `Dataset` 对象是可迭代的。 我们来看看下面的一些特征:
# + colab_type="code" id="iDuG94H-C122" colab={}
features, labels = next(iter(train_dataset))
print(features)
# + [markdown] colab_type="text" id="E63mArnQaAGz"
# 注意到具有相似特征的样本会归为一组,即分为一批。更改 `batch_size` 可以设置存储在这些特征数组中的样本数。
#
# 绘制该批次中的几个特征后,就会开始看到一些集群现象:
# + colab_type="code" id="me5Wn-9FcyyO" colab={}
plt.scatter(features['petal_length'],
features['sepal_length'],
c=labels,
cmap='viridis')
plt.xlabel("Petal length")
plt.ylabel("Sepal length")
plt.show()
# + [markdown] colab_type="text" id="YlxpSyHlhT6M"
# 要简化模型构建步骤,请创建一个函数以将特征字典重新打包为形状为 `(batch_size, num_features)` 的单个数组。
#
# 此函数使用 [tf.stack](https://www.tensorflow.org/api_docs/python/tf/stack) 方法,该方法从张量列表中获取值,并创建指定维度的组合张量:
# + colab_type="code" id="jm932WINcaGU" colab={}
def pack_features_vector(features, labels):
"""将特征打包到一个数组中"""
features = tf.stack(list(features.values()), axis=1)
return features, labels
# + [markdown] colab_type="text" id="V1Vuph_eDl8x"
# 然后使用 [tf.data.Dataset.map](https://www.tensorflow.org/api_docs/python/tf/data/dataset/map) 方法将每个 `(features,label)` 对中的 `features` 打包到训练数据集中:
# + colab_type="code" id="ZbDkzGZIkpXf" colab={}
train_dataset = train_dataset.map(pack_features_vector)
# + [markdown] colab_type="text" id="NLy0Q1xCldVO"
# `Dataset` 的特征元素被构成了形如 `(batch_size, num_features)` 的数组。我们来看看前几个样本:
# + colab_type="code" id="kex9ibEek6Tr" colab={}
features, labels = next(iter(train_dataset))
print(features[:5])
# + [markdown] colab_type="text" id="LsaVrtNM3Tx5"
# ## 选择模型类型
#
# ### 为何要使用模型?
#
# [模型](https://developers.google.com/machine-learning/crash-course/glossary#model)是指特征与标签之间的关系。对于鸢尾花分类问题,模型定义了花萼和花瓣测量值与预测的鸢尾花品种之间的关系。一些简单的模型可以用几行代数进行描述,但复杂的机器学习模型拥有大量难以汇总的参数。
#
#
# 您能否在不使用机器学习的情况下确定四个特征与鸢尾花品种之间的关系?也就是说,您能否使用传统编程技巧(例如大量条件语句)创建模型?也许能,前提是反复分析该数据集,并最终确定花瓣和花萼测量值与特定品种的关系。对于更复杂的数据集来说,这会变得非常困难,或许根本就做不到。一个好的机器学习方法可为您确定模型。如果您将足够多的代表性样本馈送到正确类型的机器学习模型中,该程序便会为您找出相应的关系。
#
# ### 选择模型
#
# 我们需要选择要进行训练的模型类型。模型具有许多类型,挑选合适的类型需要一定的经验。本教程使用神经网络来解决鸢尾花分类问题。[神经网络](https://developers.google.com/machine-learning/glossary/#neural_network)可以发现特征与标签之间的复杂关系。神经网络是一个高度结构化的图,其中包含一个或多个[隐含层](https://developers.google.com/machine-learning/glossary/#hidden_layer)。每个隐含层都包含一个或多个[神经元](https://developers.google.com/machine-learning/glossary/#neuron)。 神经网络有多种类别,该程序使用的是密集型神经网络,也称为[全连接神经网络](https://developers.google.com/machine-learning/glossary/#fully_connected_layer) : 一个层中的神经元将从上一层中的每个神经元获取输入连接。例如,图 2 显示了一个密集型神经网络,其中包含 1 个输入层、2 个隐藏层以及 1 个输出层:
#
# <table>
# <tr><td>
# <img src="https://www.tensorflow.org/images/custom_estimators/full_network.png"
# alt="网络结构示意图: 输入层, 2 隐含层, 输出层">
# </td></tr>
# <tr><td align="center">
# <b>图 2.</b> 包含特征、隐藏层和预测的神经网络<br/>
# </td></tr>
# </table>
#
# 当图 2 中的模型经过训练并获得无标签样本后,它会产生 3 个预测结果:相应鸢尾花属于指定品种的可能性。这种预测称为[推理](https://developers.google.com/machine-learning/crash-course/glossary#inference)。对于该示例,输出预测结果的总和是 1.0。在图 2 中,该预测结果分解如下:山鸢尾为 0.02,变色鸢尾为 0.95,维吉尼亚鸢尾为 0.03。这意味着该模型预测某个无标签鸢尾花样本是变色鸢尾的概率为 95%。
# + [markdown] colab_type="text" id="W23DIMVPQEBt"
# ### 使用 Keras 创建模型
#
# TensorFlow [tf.keras](https://www.tensorflow.org/api_docs/python/tf/keras) API 是创建模型和层的首选方式。通过该 API,您可以轻松地构建模型并进行实验,而将所有部分连接在一起的复杂工作则由 Keras 处理。
#
# [tf.keras.Sequential](https://www.tensorflow.org/api_docs/python/tf/keras/Sequential) 模型是层的线性堆叠。该模型的构造函数会采用一系列层实例;在本示例中,采用的是 2 个[密集层](https://www.tensorflow.org/api_docs/python/tf/keras/layers/Dense)(各自包含10个节点),以及 1 个输出层(包含 3 个代表标签预测的节点。第一个层的 `input_shape` 参数对应该数据集中的特征数量,它是一项必需参数:
# + colab_type="code" id="2fZ6oL2ig3ZK" colab={}
model = tf.keras.Sequential([
tf.keras.layers.Dense(10, activation=tf.nn.relu, input_shape=(4,)), # 需要给出输入的形式
tf.keras.layers.Dense(10, activation=tf.nn.relu),
tf.keras.layers.Dense(3)
])
# + [markdown] colab_type="text" id="FHcbEzMpxbHL"
# [激活函数](https://developers.google.com/machine-learning/crash-course/glossary#activation_function)可决定层中每个节点的输出形式。 这些非线性关系很重要,如果没有它们,模型将等同于单个层。[激活函数](https://www.tensorflow.org/api_docs/python/tf/keras/activations)有很多种,但隐藏层通常使用 [ReLU](https://developers.google.com/machine-learning/crash-course/glossary#ReLU)。
#
# 隐藏层和神经元的理想数量取决于问题和数据集。与机器学习的多个方面一样,选择最佳的神经网络形状需要一定的知识水平和实验基础。一般来说,增加隐藏层和神经元的数量通常会产生更强大的模型,而这需要更多数据才能有效地进行训练。
# + [markdown] colab_type="text" id="2wFKnhWCpDSS"
# ### 使用模型
#
# 我们快速了解一下此模型如何处理一批特征:
#
# + colab_type="code" id="xe6SQ5NrpB-I" colab={}
predictions = model(features)
predictions[:5]
# + [markdown] colab_type="text" id="wxyXOhwVr5S3"
# 在此示例中,每个样本针对每个类别返回一个 [logit](https://developers.google.com/machine-learning/crash-course/glossary#logits)。
#
# 要将这些对数转换为每个类别的概率,请使用 [softmax](https://developers.google.com/machine-learning/crash-course/glossary#softmax) 函数:
# + colab_type="code" id="_tRwHZmTNTX2" colab={}
tf.nn.softmax(predictions[:5])
# + [markdown] colab_type="text" id="uRZmchElo481"
# 对每个类别执行 `tf.argmax` 运算可得出预测的类别索引。不过,该模型尚未接受训练,因此这些预测并不理想。
# + colab_type="code" id="-Jzm_GoErz8B" colab={}
print("Prediction: {}".format(tf.argmax(predictions, axis=1)))
print(" Labels: {}".format(labels))
# + [markdown] colab_type="text" id="Vzq2E5J2QMtw"
# ## 训练模型
#
# [训练](https://developers.google.com/machine-learning/crash-course/glossary#training) 是一个机器学习阶段,在此阶段中,模型会逐渐得到优化,也就是说,模型会了解数据集。目标是充分了解训练数据集的结构,以便对未见过的数据进行预测。如果您从训练数据集中获得了过多的信息,预测便会仅适用于模型见过的数据,但是无法泛化。此问题被称之为[过拟合](https://developers.google.com/machine-learning/crash-course/glossary#overfitting)—就好比将答案死记硬背下来,而不去理解问题的解决方式。
#
# 鸢尾花分类问题是[监督式机器学习](https://developers.google.com/machine-learning/glossary/#supervised_machine_learning)的一个示例: 模型通过包含标签的样本加以训练。 而在[非监督式机器学习](https://developers.google.com/machine-learning/glossary/#unsupervised_machine_learning)中,样本不包含标签。相反,模型通常会在特征中发现一些规律。
# + [markdown] colab_type="text" id="RaKp8aEjKX6B"
# ### 定义损失和梯度函数
#
# 在训练和评估阶段,我们都需要计算模型的[损失](https://developers.google.com/machine-learning/crash-course/glossary#loss)。 这样可以衡量模型的预测结果与预期标签有多大偏差,也就是说,模型的效果有多差。我们希望尽可能减小或优化这个值。
#
# 我们的模型会使用 `tf.keras.losses.SparseCategoricalCrossentropy` 函数计算其损失,此函数会接受模型的类别概率预测结果和预期标签,然后返回样本的平均损失。
# + colab_type="code" id="QOsi6b-1CXIn" colab={}
loss_object = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
# + colab_type="code" id="tMAT4DcMPwI-" colab={}
def loss(model, x, y):
y_ = model(x)
return loss_object(y_true=y, y_pred=y_)
l = loss(model, features, labels)
print("Loss test: {}".format(l))
# + [markdown] colab_type="text" id="3IcPqA24QM6B"
# 使用 [tf.GradientTape](https://www.tensorflow.org/api_docs/python/tf/GradientTape) 的前后关系来计算[梯度](https://developers.google.com/machine-learning/crash-course/glossary#gradient)以优化你的模型:
# + colab_type="code" id="x57HcKWhKkei" colab={}
def grad(model, inputs, targets):
with tf.GradientTape() as tape:
loss_value = loss(model, inputs, targets)
return loss_value, tape.gradient(loss_value, model.trainable_variables)
# + [markdown] colab_type="text" id="lOxFimtlKruu"
# ### 创建优化器
#
# [优化器](https://developers.google.com/machine-learning/crash-course/glossary#optimizer) 会将计算出的梯度应用于模型的变量,以使 `loss` 函数最小化。您可以将损失函数想象为一个曲面(见图 3),我们希望通过到处走动找到该曲面的最低点。梯度指向最高速上升的方向,因此我们将沿相反的方向向下移动。我们以迭代方式计算每个批次的损失和梯度,以在训练过程中调整模型。模型会逐渐找到权重和偏差的最佳组合,从而将损失降至最低。损失越低,模型的预测效果就越好。
#
# <table>
# <tr><td>
# <img src="https://cs231n.github.io/assets/nn3/opt1.gif" width="70%"
# alt="Optimization algorithms visualized over time in 3D space.">
# </td></tr>
# <tr><td align="center">
# <b>图 3.</b> 优化算法在三维空间中随时间推移而变化的可视化效果。<br/>(来源: <a href="http://cs231n.github.io/neural-networks-3/">斯坦福大学 CS231n 课程</a>,MIT 许可证,Image credit: <a href="https://twitter.com/alecrad"><NAME></a>)
# </td></tr>
# </table>
#
# TensorFlow有许多可用于训练的[优化算法](https://www.tensorflow.org/api_guides/python/train)。此模型使用的是 [tf.train.GradientDescentOptimizer](https://www.tensorflow.org/api_docs/python/tf/train/GradientDescentOptimizer) , 它可以实现[随机梯度下降法](https://developers.google.com/machine-learning/crash-course/glossary#gradient_descent)(SGD)。`learning_rate` 被用于设置每次迭代(向下行走)的步长。 这是一个 *超参数* ,您通常需要调整此参数以获得更好的结果。
# + [markdown] colab_type="text" id="XkUd6UiZa_dF"
# 我们来设置优化器:
# + colab_type="code" id="8xxi2NNGKwG_" colab={}
optimizer = tf.keras.optimizers.Adam(learning_rate=0.01)
# + [markdown] colab_type="text" id="jYgezhPMhaNE"
# 我们将使用它来计算单个优化步骤:
# + colab_type="code" id="rxRNTFVe56RG" colab={}
loss_value, grads = grad(model, features, labels)
print("Step: {}, Initial Loss: {}".format(optimizer.iterations.numpy(),
loss_value.numpy()))
optimizer.apply_gradients(zip(grads, model.trainable_variables))
print("Step: {}, Loss: {}".format(optimizer.iterations.numpy(),
loss(model, features, labels).numpy()))
# + [markdown] colab_type="text" id="7Y2VSELvwAvW"
# ### 训练循环
#
#
# 一切准备就绪后,就可以开始训练模型了!训练循环会将数据集样本馈送到模型中,以帮助模型做出更好的预测。以下代码块可设置这些训练步骤:
#
# 1. 迭代每个周期。通过一次数据集即为一个周期。
# 2. 在一个周期中,遍历训练 `Dataset` 中的每个样本,并获取样本的*特征*(`x`)和*标签*(`y`)。
# 3. 根据样本的特征进行预测,并比较预测结果和标签。衡量预测结果的不准确性,并使用所得的值计算模型的损失和梯度。
# 4. 使用 `optimizer` 更新模型的变量。
# 5. 跟踪一些统计信息以进行可视化。
# 6. 对每个周期重复执行以上步骤。
#
#
# `num_epochs` 变量是遍历数据集集合的次数。与直觉恰恰相反的是,训练模型的时间越长,并不能保证模型就越好。`num_epochs` 是一个可以调整的[超参数](https://developers.google.com/machine-learning/glossary/#hyperparameter)。选择正确的次数通常需要一定的经验和实验基础。
# + colab_type="code" id="AIgulGRUhpto" colab={}
## Note: 使用相同的模型变量重新运行此单元
# 保留结果用于绘制
train_loss_results = []
train_accuracy_results = []
num_epochs = 201
for epoch in range(num_epochs):
epoch_loss_avg = tf.keras.metrics.Mean()
epoch_accuracy = tf.keras.metrics.SparseCategoricalAccuracy()
# Training loop - using batches of 32
for x, y in train_dataset:
# 优化模型
loss_value, grads = grad(model, x, y)
optimizer.apply_gradients(zip(grads, model.trainable_variables))
# 追踪进度
epoch_loss_avg(loss_value) # 添加当前的 batch loss
# 比较预测标签与真实标签
epoch_accuracy(y, model(x))
# 循环结束
train_loss_results.append(epoch_loss_avg.result())
train_accuracy_results.append(epoch_accuracy.result())
if epoch % 50 == 0:
print("Epoch {:03d}: Loss: {:.3f}, Accuracy: {:.3%}".format(epoch,
epoch_loss_avg.result(),
epoch_accuracy.result()))
# + [markdown] colab_type="text" id="2FQHVUnm_rjw"
# ### 可视化损失函数随时间推移而变化的情况
# + [markdown] colab_type="text" id="j3wdbmtLVTyr"
# 虽然输出模型的训练过程有帮助,但查看这一过程往往*更有帮助*。 [TensorBoard](https://www.tensorflow.org/guide/summaries_and_tensorboard) 是与 TensorFlow 封装在一起的出色可视化工具,不过我们可以使用 `matplotlib` 模块创建基本图表。
#
# 解读这些图表需要一定的经验,不过您确实希望看到*损失*下降且*准确率*上升。
# + colab_type="code" id="agjvNd2iUGFn" colab={}
fig, axes = plt.subplots(2, sharex=True, figsize=(12, 8))
fig.suptitle('Training Metrics')
axes[0].set_ylabel("Loss", fontsize=14)
axes[0].plot(train_loss_results)
axes[1].set_ylabel("Accuracy", fontsize=14)
axes[1].set_xlabel("Epoch", fontsize=14)
axes[1].plot(train_accuracy_results)
plt.show()
# + [markdown] colab_type="text" id="Zg8GoMZhLpGH"
# ## 评估模型的效果
#
# 模型已经过训练,现在我们可以获取一些关于其效果的统计信息了。
#
# *评估* 指的是确定模型做出预测的效果。要确定模型在鸢尾花分类方面的效果,请将一些花萼和花瓣测量值传递给模型,并要求模型预测它们所代表的鸢尾花品种。然后,将模型的预测结果与实际标签进行比较。例如,如果模型对一半输入样本的品种预测正确,则 [准确率](https://developers.google.com/machine-learning/glossary/#accuracy) 为 `0.5` 。 图 4 显示的是一个效果更好一些的模型,该模型做出 5 次预测,其中有 4 次正确,准确率为 80%:
#
# <table cellpadding="8" border="0">
# <colgroup>
# <col span="4" >
# <col span="1" bgcolor="lightblue">
# <col span="1" bgcolor="lightgreen">
# </colgroup>
# <tr bgcolor="lightgray">
# <th colspan="4">样本特征</th>
# <th colspan="1">标签</th>
# <th colspan="1" >模型预测</th>
# </tr>
# <tr>
# <td>5.9</td><td>3.0</td><td>4.3</td><td>1.5</td><td align="center">1</td><td align="center">1</td>
# </tr>
# <tr>
# <td>6.9</td><td>3.1</td><td>5.4</td><td>2.1</td><td align="center">2</td><td align="center">2</td>
# </tr>
# <tr>
# <td>5.1</td><td>3.3</td><td>1.7</td><td>0.5</td><td align="center">0</td><td align="center">0</td>
# </tr>
# <tr>
# <td>6.0</td> <td>3.4</td> <td>4.5</td> <td>1.6</td> <td align="center">1</td><td align="center" bgcolor="red">2</td>
# </tr>
# <tr>
# <td>5.5</td><td>2.5</td><td>4.0</td><td>1.3</td><td align="center">1</td><td align="center">1</td>
# </tr>
# <tr><td align="center" colspan="6">
# <b>图 4.</b> 准确率为 80% 的鸢尾花分类器<br/>
# </td></tr>
# </table>
# + [markdown] colab_type="text" id="z-EvK7hGL0d8"
# ### 建立测试数据集
#
# 评估模型与训练模型相似。最大的区别在于,样本来自一个单独的[测试集](https://developers.google.com/machine-learning/crash-course/glossary#test_set),而不是训练集。为了公正地评估模型的效果,用于评估模型的样本务必与用于训练模型的样本不同。
#
# 测试 `Dataset` 的建立与训练 `Dataset` 相似。下载 CSV 文本文件并解析相应的值,然后对数据稍加随机化处理:
# + colab_type="code" id="Ps3_9dJ3Lodk" colab={}
test_url = "https://storage.googleapis.com/download.tensorflow.org/data/iris_test.csv"
test_fp = tf.keras.utils.get_file(fname=os.path.basename(test_url),
origin=test_url)
# + colab_type="code" id="SRMWCu30bnxH" colab={}
test_dataset = tf.data.experimental.make_csv_dataset(
test_fp,
batch_size,
column_names=column_names,
label_name='species',
num_epochs=1,
shuffle=False)
test_dataset = test_dataset.map(pack_features_vector)
# + [markdown] colab_type="text" id="HFuOKXJdMAdm"
# ### 根据测试数据集评估模型
#
# 与训练阶段不同,模型仅评估测试数据的一个[周期](https://developers.google.com/machine-learning/glossary/#epoch)。在以下代码单元格中,我们会遍历测试集中的每个样本,然后将模型的预测结果与实际标签进行比较。这是为了衡量模型在整个测试集中的准确率。
# + colab_type="code" id="Tw03-MK1cYId" colab={}
test_accuracy = tf.keras.metrics.Accuracy()
for (x, y) in test_dataset:
logits = model(x)
prediction = tf.argmax(logits, axis=1, output_type=tf.int32)
test_accuracy(prediction, y)
print("Test set accuracy: {:.3%}".format(test_accuracy.result()))
# + [markdown] colab_type="text" id="HcKEZMtCOeK-"
# 例如,我们可以看到对于最后一批数据,该模型通常预测正确:
# + colab_type="code" id="uNwt2eMeOane" colab={}
tf.stack([y,prediction],axis=1)
# + [markdown] colab_type="text" id="7Li2r1tYvW7S"
# ## 使用经过训练的模型进行预测
#
# 我们已经训练了一个模型并“证明”它是有效的,但在对鸢尾花品种进行分类方面,这还不够。现在,我们使用经过训练的模型对 [无标签样本](https://developers.google.com/machine-learning/glossary/#unlabeled_example)(即包含特征但不包含标签的样本)进行一些预测。
#
# 在现实生活中,无标签样本可能来自很多不同的来源,包括应用、CSV 文件和数据 Feed。暂时我们将手动提供三个无标签样本以预测其标签。回想一下,标签编号会映射到一个指定的表示法:
#
# * `0`: 山鸢尾
# * `1`: 变色鸢尾
# * `2`: 维吉尼亚鸢尾
# + colab_type="code" id="kesTS5Lzv-M2" colab={}
predict_dataset = tf.convert_to_tensor([
[5.1, 3.3, 1.7, 0.5,],
[5.9, 3.0, 4.2, 1.5,],
[6.9, 3.1, 5.4, 2.1]
])
predictions = model(predict_dataset)
for i, logits in enumerate(predictions):
class_idx = tf.argmax(logits).numpy()
p = tf.nn.softmax(logits)[class_idx]
name = class_names[class_idx]
print("Example {} prediction: {} ({:4.1f}%)".format(i, name, 100*p))
|
site/zh-cn/tutorials/customization/custom_training_walkthrough.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="B89FZT360MG8" colab_type="text"
# ## Part 2: Determine if a number is a power of two.
#
# * [Constraints](#Constraints)
# * [Test Cases](#Test-Cases)
# * [Algorithm](#Algorithm)
# * [Code](#Code)
# * [Unit Test](#Unit-Test)
# * [Solution Notebook](#Solution-Notebook)
# + [markdown] id="vC5R9S460MG9" colab_type="text"
# ## Constraints
#
# * Is the input number an int?
# * Yes
# * Can we assume the inputs are valid?
# * No
# * Is the output a boolean?
# * Yes
# * Can we assume this fits memory?
# * Yes
# + [markdown] id="sx1STn0h0MG-" colab_type="text"
# ## Test Cases
#
# * None -> TypeError
# * 0 -> False
# * 1 -> True
# * 2 -> True
# * 15 -> False
# * 16 -> True
# + [markdown] id="ZckQHKjT0MG_" colab_type="text"
# ## Algorithm
#
# We can use bit manipulation to determine if a number is a power of two.
#
# For a number to be a power of two, there must only be one bit that is a 1.
#
# We can use the following bit manipulation trick to determine this:
#
# n & (n - 1)
#
# Here's an example why:
#
# 0000 1000 = n
# 0000 0001 = 1
# 0000 0111 = n-1
#
# 0000 1000 = n
# 0000 0111 = n-1
# 0000 0000 = n & n-1, result = 0
# + [markdown] id="wOupTir-0MG_" colab_type="text"
# ## Code
# + id="8-cs-nga0MHA" colab_type="code" colab={}
class Solution(object):
def is_power_of_two(self, val):
# TODO: Implement me
# return val & val-1
return val // 1<< (val.bit_length()-1)
# + [markdown] id="ltVnKjb80MHD" colab_type="text"
# ## Unit Test
# + [markdown] id="llVg-okI0MHD" colab_type="text"
# **The following unit test is expected to fail until you solve the challenge.**
# + id="qwkNtZAe0MHE" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 307} outputId="f8549f74-78f0-4867-c9ce-ece3426f23cd"
# # %load test_is_power_of_two.py
from nose.tools import assert_equal, assert_raises
class TestSolution(object):
def test_is_power_of_two(self):
solution = Solution()
assert_raises(TypeError, solution.is_power_of_two, None)
assert_equal(solution.is_power_of_two(0), False)
assert_equal(solution.is_power_of_two(1), True)
assert_equal(solution.is_power_of_two(2), True)
assert_equal(solution.is_power_of_two(15), False)
assert_equal(solution.is_power_of_two(16), True)
assert_equal(solution.is_power_of_two(31), False)
assert_equal(solution.is_power_of_two(64), True)
assert_equal(solution.is_power_of_two(128), True)
assert_equal(solution.is_power_of_two(203), False)
print('Success: test_is_power_of_two')
def main():
test = TestSolution()
test.test_is_power_of_two()
if __name__ == '__main__':
main()
# + id="E6okIbfd0MHG" colab_type="code" colab={"base_uri": "https://localhost:8080/", "height": 34} outputId="4fd9218e-9b4f-4997-b3b1-a136b7855f21"
pip install nose
|
Exam2Part2.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <h1 style="text-align:center;">MiA <span style="text-align:center;font-size: 0.5em;">0.9.1</span></h1>
# <h2 style="text-align:center;">Mito Hacker Toolkit <i style="font-size: 0.5em;">0.7.1</i></h2>
# <h3 style="text-align:center;">Kashatus Lab @ UVA</h3>
# # Welcome to MiA (Mitochondrial Analyzer)
# #### MiA is part of Mito Hacker toolkit that enables you to quantify the segmented miotchondrial netwrk generated by Mito Miner
#
# This Jupyter notebook provides you with step-by-step directions to quantify your mitochondria.
# ## 1) Import necessary libraries
# ##### Just run the following block of code, you are not expected to enter any data here.
# +
#Base Libraries
import os
import shutil
#Core Functions
import cemia55s as cemia
# -
# ## 2) Locating your files
#
# <br>
# <details>
# <summary><span style="font-size:16px;font-weight: bold; color:red">What should I do next? (click here to expand)</span></summary>
#
# #### <span style="color:red;">You need to interact with the next cell: </span> Please run the next cell, a box will appear. Enter the relative/absolute address of the folder that contains your images, then press enter.
#
#
# #### <span style="color:red;">Examples: </span>
# * Relative Address
# * Use . if the images are in the same folder as this file
# * If your folder of the images (my_folder_of_images) is located in the same directory as this file, you should enter: my_folder_of_images
# * Absolute Address
# *If your images are located on Desktop
# * Mac: you should enter: /Users/username/Desktop/my_folder_of_images
# * Windows: you should enter: C:\Users\username\Desktop\my_folder_of_images
#
# #### <span style="color:red;">Note: </span>
# * It is preferred to have the folder of your images in the same folder as the current file that you are running
# * After you entered the desired value in the box, you should press the enter key.
# * If you have previously used Cell Catcher and Mito Miner to isolate and segment individual cells, and you want to analyze those images, you should enter the address where your original images (Multi-cell images) are located. This is the same address you used in Cell Catcher, and Mito Miner.
# * Example: If you have previously used Cell Catcher to isolate individual cells in a folder named "test", you should enter "test" as the address to your files.
# </details>
address,file_list = cemia.address()
# ### 3) Checking requirements
# +
try:
os.makedirs(os.path.join(address, 'output', 'processed','single_cells_binary'))
except:
pass
cell_list = os.listdir(os.path.join(address, 'output', 'processed','single_cells_binary'))
if len(cell_list) <= 1:
print('It seems you dont have any images to analyze!')
else:
print('You are all set!')
# -
# ## 4) Name your output data filename
# ##### What do you want to name your output file? This is the file that has your measurements results.
output_filename = input('What is the name of output csv file?\nJust enter the name without .csv extension\nWe Will add the current date to the beginning of the file name!\n\n')
# ## 5) Quantifying all the files
# #### Just run this block of code, and then go and enjoy your time. Running this block of code may take few hours depending on the number of images you have.
cemia.measurement(address,cell_list,output_filename)
# # The End!
|
MiA.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from pandas_plink import read_plink
(bim, fam, bed) = read_plink('maf1_allchr')
bim.head()
fam.head()
bed
chrom1 = bim.loc[('1', ), :]
chrom1.head()
# +
window = 5000
left = 870153 - window
right = 870153 + window
chrom1 = chrom1.reset_index()
i = chrom1[(chrom1['pos'] > left) & (chrom1['pos'] < right)].i
# -
X = bed[i,:].compute()
X
|
1000G_example.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import json
from PIL import Image
# +
qas_file_names = ['questions_answers/qas_synonyms-pg04-mask-2.json', 'questions_answers/qas_ms_2_pg04.json']
current_experiment = qas_file_names[0]
print(current_experiment)
qas_json = json.load(open(current_experiment))
# +
list_names = ['wtr', 'rtw']
current_list = list_names[0]
print(current_list)
questions_json = qas_json[current_list]
# -
# # Iterate through
qids_iter = iter(questions_json.keys())
# ## iterate until qid
input_q = '66669011'
qids_iter = iter(questions_json.keys())
qid = next(qids_iter)
while qid != input_q:
qid = next(qids_iter)
qid = next(qids_iter)
q = questions_json[qid]
print(qid)
print("==== BASELINE ====")
print("Q: {}".format(q['bl_q']))
print("A: {}".format(q['bl_ans']))
if current_experiment == 'questions_answers/qas_ms_2_pg04.json':
print("==== MASKED ====")
print("Q: {}".format(q['mexp_q']))
print("A: {}".format(q['mexp_ans']))
print("==== EXPERIMENT ====")
print("Q: {}".format(q['exp_q']))
print("A: {}".format(q['exp_ans']))
Image.open("vqa_images/{}".format(q['img_fname']))
# ### Get by QID
qid = ''
q = question_json[qid]
print(qid)
print("==== BASELINE ====")
print("Q: {}".format(q['bl_q']))
print("A: {}".format(q['bl_ans']))
print("==== EXPERIMENT ====")
print("Q: {}".format(q['exp_q']))
print("A: {}".format(q['exp_ans']))
Image.open("vqa_images/{}".format(q['img_fname']))
|
Analysis.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import pyspark
# ERRORS EVERYWHERE >.<
from pyspark.sql import SparkSession
session = SparkSession.builder.getOrCreate()
rdd = session.sparkContext.parallelize([1,2,3])
rdd.take(num=2)
rdd.count()
rdd.collect()
df = session.createDataFrame(
[[1,2,3], [4,5,6]], ['column1', 'column2', 'column3']
)
df.show(n=3)
a = list(range(10))
a.append(11)
st = 'my string'
st += 'is pretty'
my_rdd = my_rdd.map(lambda x: x*100)
import pyspark.sql.functions as funcs
import math
import pyspark.sql.types as types
def multiply_by_ten(number):
return number*10.0
multiply_udf = funcs.udf(multiply_by_ten, types.DoubleType())
transformed_df = df.withColumn(
'multiplied', multiply_udf('column1')
)
transformed_df.show()
import pyspark.sql.types as types
def take_log_in_all_columns(row: types.Row):
old_row = row.asDict()
new_row = {f'log({column_name})': math.log(value)
for column_name, value in old_row.items()}
return types.Row(**new_row)
logarithmic_dataframe = df.rdd.map(take_log_in_all_columns).toDF()
df.select('column1', 'column2')
df.where('column1 = 3')
df.join(df1, ['column1'], how='inner')
df2 = session.sql("SELECT column1 AS f1, column2 as f2 from table1")
df3 = df.withColumn(
'derived_column', df['column1'] + df['column2'] * df['column3']
)
ADULT_COLUMN_NAMES = [
"age",
"workclass",
"fnlwgt",
"education",
"education_num",
"marital_status",
"occupation",
"relationship",
"race",
"sex",
"capital_gain",
"capital_loss",
"hours_per_week",
"native_country",
"income"
]
csv_df = session.read.csv(
'data/adult.data.csv', header=False, inferSchema=True
)
for new_col, old_col in zip(ADULT_COLUMN_NAMES, csv_df.columns):
csv_df = csv_df.withColumnRenamed(old_col, new_col)
csv_df.describe().show()
work_hours_df = csv_df.groupBy(
'age'
).agg(
funcs.avg('hours_per_week'),
funcs.stddev_samp('hours_per_week')
).sort('age')
session = SparkSession.builder.config(
'spark.jars', 'bin/postgresql-42.2.16.jar'
).config(
'spark.driver.extraClassPath', 'bin/postgresql-42.2.16.jar'
).getOrCreate()
url = f"jdbc:postgresql://your_host_ip:5432/your_database"
properties = {'user': 'your_user', 'password': '<PASSWORD>'}
# read from a table into a dataframe
df = session.read.jdbc(
url=url, table='your_table_name', properties=properties
)
transformed_df.write.jdbc(
url=url, table='new_table', mode='append', properties=properties
)
|
w7/w7d3/docker/Pyspark_data_manipulation.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="58MWWgq75lMh"
# ##### Copyright 2020 The TensorFlow Hub Authors.
# + cellView="form" id="jM3hCI1UUzar"
#@title Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] id="4_NEJlxKKjyI"
# <table class="tfo-notebook-buttons" align="left">
# <td>
# <a target="_blank" href="https://www.tensorflow.org/tutorials/text/solve_glue_tasks_using_bert_on_tpu"><img src="https://www.tensorflow.org/images/tf_logo_32px.png" />View on TensorFlow.org</a>
# </td>
# <td>
# <a target="_blank" href="https://colab.research.google.com/github/tensorflow/docs/blob/master/site/en/tutorials/text/solve_glue_tasks_using_bert_on_tpu.ipynb"><img src="https://www.tensorflow.org/images/colab_logo_32px.png" />Run in Google Colab</a>
# </td>
# <td>
# <a target="_blank" href="https://github.com/tensorflow/docs/blob/master/site/en/tutorials/text/solve_glue_tasks_using_bert_on_tpu.ipynb"><img src="https://www.tensorflow.org/images/GitHub-Mark-32px.png" />View on GitHub</a>
# </td>
# <td>
# <a href="https://storage.googleapis.com/tensorflow_docs/docs/site/en/tutorials/text/solve_glue_tasks_using_bert_on_tpu.ipynb"><img src="https://www.tensorflow.org/images/download_logo_32px.png" />Download notebook</a>
# </td>
# <td>
# <a href="https://tfhub.dev/google/collections/bert/1"><img src="https://www.tensorflow.org/images/hub_logo_32px.png" />See TF Hub model</a>
# </td>
# </table>
# + [markdown] id="U5POcTVNB_dv"
# # Solve GLUE tasks using BERT on TPU
#
# BERT can be used to solve many problems in natural language processing. You will learn how to fine-tune BERT for many tasks from the [GLUE benchmark](https://gluebenchmark.com/):
#
# 1. [CoLA](https://nyu-mll.github.io/CoLA/) (Corpus of Linguistic Acceptability): Is the sentence grammatically correct?
#
# 1. [SST-2](https://nlp.stanford.edu/sentiment/index.html) (Stanford Sentiment Treebank): The task is to predict the sentiment of a given sentence.
#
# 1. [MRPC](https://www.microsoft.com/en-us/download/details.aspx?id=52398) (Microsoft Research Paraphrase Corpus): Determine whether a pair of sentences are semantically equivalent.
#
# 1. [QQP](https://data.quora.com/First-Quora-Dataset-Release-Question-Pairs) (Quora Question Pairs2): Determine whether a pair of questions are semantically equivalent.
#
# 1. [MNLI](http://www.nyu.edu/projects/bowman/multinli/) (Multi-Genre Natural Language Inference): Given a premise sentence and a hypothesis sentence, the task is to predict whether the premise entails the hypothesis (entailment), contradicts the hypothesis (contradiction), or neither (neutral).
#
# 1. [QNLI](https://rajpurkar.github.io/SQuAD-explorer/)(Question-answering Natural Language Inference): The task is to determine whether the context sentence contains the answer to the question.
#
# 1. [RTE](https://aclweb.org/aclwiki/Recognizing_Textual_Entailment)(Recognizing Textual Entailment): Determine if a sentence entails a given hypothesis or not.
#
# 1. [WNLI](https://cs.nyu.edu/faculty/davise/papers/WinogradSchemas/WS.html)(Winograd Natural Language Inference): The task is to predict if the sentence with the pronoun substituted is entailed by the original sentence.
#
# This tutorial contains complete end-to-end code to train these models on a TPU. You can also run this notebook on a GPU, by changing one line (described below).
#
# In this notebook, you will:
#
# - Load a BERT model from TensorFlow Hub
# - Choose one of GLUE tasks and download the dataset
# - Preprocess the text
# - Fine-tune BERT (examples are given for single-sentence and multi-sentence datasets)
# - Save the trained model and use it
#
# Key point: The model you develop will be end-to-end. The preprocessing logic will be included in the model itself, making it capable of accepting raw strings as input.
#
# Note: This notebook should be run using a TPU. In Colab, choose **Runtime -> Change runtime type** and verify that a **TPU** is selected.
#
# + [markdown] id="SCjmX4zTCkRK"
# ## Setup
#
# You will use a separate model to preprocess text before using it to fine-tune BERT. This model depends on [tensorflow/text](https://github.com/tensorflow/text), which you will install below.
# + id="q-YbjCkzw0yU"
# !pip install -q -U tensorflow-text
# + [markdown] id="WMaudPO1a2Hx"
# You will use the AdamW optimizer from [tensorflow/models](https://github.com/tensorflow/models) to fine-tune BERT, which you will install as well.
# + id="5zwJyopqa3uH"
# !pip install -q -U tf-models-official
# + id="_XgTpm9ZxoN9"
import os
import tensorflow as tf
import tensorflow_hub as hub
import tensorflow_datasets as tfds
import tensorflow_text as text # A dependency of the preprocessing model
import tensorflow_addons as tfa
from official.nlp import optimization
import numpy as np
tf.get_logger().setLevel('ERROR')
# + [markdown] id="sv7A19G32Kfw"
# Next, configure TFHub to read checkpoints directly from TFHub's Cloud Storage buckets. This is only recomended when running TFHub models on TPU.
#
# Without this setting TFHub would download the compressed file and extract the checkpoint locally. Attempting to load from these local files will fail with following Error:
#
# ```
# InvalidArgumentError: Unimplemented: File system scheme '[local]' not implemented
# ```
#
# This is because the [TPU can only read directly from Cloud Storage buckets](https://cloud.google.com/tpu/docs/troubleshooting#cannot_use_local_filesystem).
#
# Note: This setting is automatic in Colab.
# + id="Sz6P5pK3ldxQ"
os.environ["TFHUB_MODEL_LOAD_FORMAT"]="UNCOMPRESSED"
# + [markdown] id="__Uqe2vNETAu"
# ### Connect to a TPU
#
# To use a TPU, you will define a distribution strategy to wrap the training logic.
# See this [guide](https://www.tensorflow.org/guide/tpu) for more information.
#
# Note: You can verify that you are connected to a TPU by going to **Runtime** → **Change runtime type** to make sure that **TPU** is selected
# + id="cpHWNs1nV0Zn"
import os
if os.environ['COLAB_TPU_ADDR']:
cluster_resolver = tf.distribute.cluster_resolver.TPUClusterResolver(tpu='grpc://' + os.environ['COLAB_TPU_ADDR'])
tf.config.experimental_connect_to_cluster(cluster_resolver)
tf.tpu.experimental.initialize_tpu_system(cluster_resolver)
strategy = tf.distribute.TPUStrategy(cluster_resolver)
print('Using TPU')
elif tf.test.is_gpu_available():
strategy = tf.distribute.MirroredStrategy()
print('Using GPU')
else:
raise ValueError('Running on CPU is not recomended.')
# + [markdown] id="UVtEyxDFpKE1"
# ## Loading models from TensorFlow Hub
#
# Here you can choose which BERT model you will load from TensorFlow Hub and fine-tune.
# There are multiple BERT models available to choose from.
#
# - [BERT-Base](https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3), [Uncased](https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3) and [seven more models](https://tfhub.dev/google/collections/bert/1) with trained weights released by the original BERT authors.
# - [Small BERTs](https://tfhub.dev/google/collections/bert/1) have the same general architecture but fewer and/or smaller Transformer blocks, which lets you explore tradeoffs between speed, size and quality.
# - [ALBERT](https://tfhub.dev/google/collections/albert/1): four different sizes of "A Lite BERT" that reduces model size (but not computation time) by sharing parameters between layers.
# - [BERT Experts](https://tfhub.dev/google/collections/experts/bert/1): eight models that all have the BERT-base architecture but offer a choice between different pre-training domains, to align more closely with the target task.
# - [Electra](https://tfhub.dev/google/collections/electra/1) has the same architecture as BERT (in three different sizes), but gets pre-trained as a discriminator in a set-up that resembles a Generative Adversarial Network (GAN).
# - BERT with Talking-Heads Attention and Gated GELU [[base](https://tfhub.dev/tensorflow/talkheads_ggelu_bert_en_base/1), [large](https://tfhub.dev/tensorflow/talkheads_ggelu_bert_en_large/1)] has two improvements to the core of the Transformer architecture.
#
# See the model documentation linked above for more details.
#
# In this tutorial, you will start with BERT-base. You can use larger and more recent models for higher accuracy, or smaller models for faster training times. To change the model, you only need to switch a single line of code (shown below). All of the differences are encapsulated in the SavedModel you will download from TensorFlow Hub.
# + cellView="form" id="y8_ctG55-uTX"
#@title Choose a BERT model to fine-tune
bert_model_name = 'bert_en_uncased_L-12_H-768_A-12' #@param ["bert_en_uncased_L-12_H-768_A-12", "bert_en_uncased_L-24_H-1024_A-16", "bert_en_wwm_uncased_L-24_H-1024_A-16", "bert_en_cased_L-12_H-768_A-12", "bert_en_cased_L-24_H-1024_A-16", "bert_en_wwm_cased_L-24_H-1024_A-16", "bert_multi_cased_L-12_H-768_A-12", "small_bert/bert_en_uncased_L-2_H-128_A-2", "small_bert/bert_en_uncased_L-2_H-256_A-4", "small_bert/bert_en_uncased_L-2_H-512_A-8", "small_bert/bert_en_uncased_L-2_H-768_A-12", "small_bert/bert_en_uncased_L-4_H-128_A-2", "small_bert/bert_en_uncased_L-4_H-256_A-4", "small_bert/bert_en_uncased_L-4_H-512_A-8", "small_bert/bert_en_uncased_L-4_H-768_A-12", "small_bert/bert_en_uncased_L-6_H-128_A-2", "small_bert/bert_en_uncased_L-6_H-256_A-4", "small_bert/bert_en_uncased_L-6_H-512_A-8", "small_bert/bert_en_uncased_L-6_H-768_A-12", "small_bert/bert_en_uncased_L-8_H-128_A-2", "small_bert/bert_en_uncased_L-8_H-256_A-4", "small_bert/bert_en_uncased_L-8_H-512_A-8", "small_bert/bert_en_uncased_L-8_H-768_A-12", "small_bert/bert_en_uncased_L-10_H-128_A-2", "small_bert/bert_en_uncased_L-10_H-256_A-4", "small_bert/bert_en_uncased_L-10_H-512_A-8", "small_bert/bert_en_uncased_L-10_H-768_A-12", "small_bert/bert_en_uncased_L-12_H-128_A-2", "small_bert/bert_en_uncased_L-12_H-256_A-4", "small_bert/bert_en_uncased_L-12_H-512_A-8", "small_bert/bert_en_uncased_L-12_H-768_A-12", "albert_en_base", "albert_en_large", "albert_en_xlarge", "albert_en_xxlarge", "electra_small", "electra_base", "experts_pubmed", "experts_wiki_books", "talking-heads_base", "talking-heads_large"]
map_name_to_handle = {
'bert_en_uncased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_L-12_H-768_A-12/3',
'bert_en_uncased_L-24_H-1024_A-16':
'https://tfhub.dev/tensorflow/bert_en_uncased_L-24_H-1024_A-16/3',
'bert_en_wwm_uncased_L-24_H-1024_A-16':
'https://tfhub.dev/tensorflow/bert_en_wwm_uncased_L-24_H-1024_A-16/3',
'bert_en_cased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_cased_L-12_H-768_A-12/3',
'bert_en_cased_L-24_H-1024_A-16':
'https://tfhub.dev/tensorflow/bert_en_cased_L-24_H-1024_A-16/3',
'bert_en_wwm_cased_L-24_H-1024_A-16':
'https://tfhub.dev/tensorflow/bert_en_wwm_cased_L-24_H-1024_A-16/3',
'bert_multi_cased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_multi_cased_L-12_H-768_A-12/3',
'small_bert/bert_en_uncased_L-2_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-128_A-2/1',
'small_bert/bert_en_uncased_L-2_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-256_A-4/1',
'small_bert/bert_en_uncased_L-2_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-512_A-8/1',
'small_bert/bert_en_uncased_L-2_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-2_H-768_A-12/1',
'small_bert/bert_en_uncased_L-4_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-4_H-128_A-2/1',
'small_bert/bert_en_uncased_L-4_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-4_H-256_A-4/1',
'small_bert/bert_en_uncased_L-4_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-4_H-512_A-8/1',
'small_bert/bert_en_uncased_L-4_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-4_H-768_A-12/1',
'small_bert/bert_en_uncased_L-6_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-6_H-128_A-2/1',
'small_bert/bert_en_uncased_L-6_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-6_H-256_A-4/1',
'small_bert/bert_en_uncased_L-6_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-6_H-512_A-8/1',
'small_bert/bert_en_uncased_L-6_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-6_H-768_A-12/1',
'small_bert/bert_en_uncased_L-8_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-8_H-128_A-2/1',
'small_bert/bert_en_uncased_L-8_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-8_H-256_A-4/1',
'small_bert/bert_en_uncased_L-8_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-8_H-512_A-8/1',
'small_bert/bert_en_uncased_L-8_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-8_H-768_A-12/1',
'small_bert/bert_en_uncased_L-10_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-10_H-128_A-2/1',
'small_bert/bert_en_uncased_L-10_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-10_H-256_A-4/1',
'small_bert/bert_en_uncased_L-10_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-10_H-512_A-8/1',
'small_bert/bert_en_uncased_L-10_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-10_H-768_A-12/1',
'small_bert/bert_en_uncased_L-12_H-128_A-2':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-12_H-128_A-2/1',
'small_bert/bert_en_uncased_L-12_H-256_A-4':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-12_H-256_A-4/1',
'small_bert/bert_en_uncased_L-12_H-512_A-8':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-12_H-512_A-8/1',
'small_bert/bert_en_uncased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/small_bert/bert_en_uncased_L-12_H-768_A-12/1',
'albert_en_base':
'https://tfhub.dev/tensorflow/albert_en_base/2',
'albert_en_large':
'https://tfhub.dev/tensorflow/albert_en_large/2',
'albert_en_xlarge':
'https://tfhub.dev/tensorflow/albert_en_xlarge/2',
'albert_en_xxlarge':
'https://tfhub.dev/tensorflow/albert_en_xxlarge/2',
'electra_small':
'https://tfhub.dev/google/electra_small/2',
'electra_base':
'https://tfhub.dev/google/electra_base/2',
'experts_pubmed':
'https://tfhub.dev/google/experts/bert/pubmed/2',
'experts_wiki_books':
'https://tfhub.dev/google/experts/bert/wiki_books/2',
'talking-heads_base':
'https://tfhub.dev/tensorflow/talkheads_ggelu_bert_en_base/1',
'talking-heads_large':
'https://tfhub.dev/tensorflow/talkheads_ggelu_bert_en_large/1',
}
map_model_to_preprocess = {
'bert_en_uncased_L-24_H-1024_A-16':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'bert_en_uncased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'bert_en_wwm_cased_L-24_H-1024_A-16':
'https://tfhub.dev/tensorflow/bert_en_cased_preprocess/1',
'bert_en_cased_L-24_H-1024_A-16':
'https://tfhub.dev/tensorflow/bert_en_cased_preprocess/1',
'bert_en_cased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_cased_preprocess/1',
'bert_en_wwm_uncased_L-24_H-1024_A-16':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-2_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-2_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-2_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-2_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-4_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-4_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-4_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-4_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-6_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-6_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-6_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-6_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-8_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-8_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-8_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-8_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-10_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-10_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-10_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-10_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-12_H-128_A-2':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-12_H-256_A-4':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-12_H-512_A-8':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'small_bert/bert_en_uncased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'bert_multi_cased_L-12_H-768_A-12':
'https://tfhub.dev/tensorflow/bert_multi_cased_preprocess/1',
'albert_en_base':
'https://tfhub.dev/tensorflow/albert_en_preprocess/1',
'albert_en_large':
'https://tfhub.dev/tensorflow/albert_en_preprocess/1',
'albert_en_xlarge':
'https://tfhub.dev/tensorflow/albert_en_preprocess/1',
'albert_en_xxlarge':
'https://tfhub.dev/tensorflow/albert_en_preprocess/1',
'electra_small':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'electra_base':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'experts_pubmed':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'experts_wiki_books':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'talking-heads_base':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
'talking-heads_large':
'https://tfhub.dev/tensorflow/bert_en_uncased_preprocess/1',
}
tfhub_handle_encoder = map_name_to_handle[bert_model_name]
tfhub_handle_preprocess = map_model_to_preprocess[bert_model_name]
print(f'BERT model selected : {tfhub_handle_encoder}')
print(f'Preprocessing model auto-selected: {tfhub_handle_preprocess}')
# + [markdown] id="7WrcxxTRDdHi"
# ## Preprocess the text
#
# On the [Classify text with BERT colab](https://www.tensorflow.org/tutorials/text/classify_text_with_bert) the preprocessing model is used directly embedded with the BERT encoder.
#
# This tutorial demonstrates how to do preprocessing as part of your input pipeline for training, using Dataset.map, and then merge it into the model that gets exported for inference. That way, both training and inference can work from raw text inputs, although the TPU itself requires numeric inputs.
#
# TPU requirements aside, it can help performance have preprocessing done asynchronously in an input pipeline (you can learn more in the [tf.data performance guide](https://www.tensorflow.org/guide/data_performance)).
#
# This tutorial also demonstrates how to build multi-input models, and how to adjust the sequence length of the inputs to BERT.
#
# Let's demonstrate the preprocessing model.
# + id="-ePjboKOPmv4"
bert_preprocess = hub.load(tfhub_handle_preprocess)
tok = bert_preprocess.tokenize(tf.constant(['Hello TensorFlow!']))
print(tok)
# + [markdown] id="tRMCuruaQb5X"
# Each preprocessing model also provides a method,`.bert_pack_inputs(tensors, seq_length)`, which takes a list of tokens (like `tok` above) and a sequence length argument. This packs the inputs to create a dictionary of tensors in the format expected by the BERT model.
# + id="lraoc4csP0lY"
text_preprocessed = bert_preprocess.bert_pack_inputs([tok, tok], tf.constant(20))
print('Shape Word Ids : ', text_preprocessed['input_word_ids'].shape)
print('Word Ids : ', text_preprocessed['input_word_ids'][0, :16])
print('Shape Mask : ', text_preprocessed['input_mask'].shape)
print('Input Mask : ', text_preprocessed['input_mask'][0, :16])
print('Shape Type Ids : ', text_preprocessed['input_type_ids'].shape)
print('Type Ids : ', text_preprocessed['input_type_ids'][0, :16])
# + [markdown] id="KyBsEhoa0_7r"
# Here are some details to pay attention to:
# - `input_mask` The mask allows the model to cleanly differentiate between the content and the padding. The mask has the same shape as the `input_word_ids`, and contains a 1 anywhere the `input_word_ids` is not padding.
# - `input_type_ids` has the same shape of `input_mask`, but inside the non-padded region, contains a 0 or a 1 indicating which sentence the token is a part of.
# + [markdown] id="H63KFuKcRwjO"
# Next, you will create a preprocessing model that encapsulates all this logic. Your model will take strings as input, and return appropriately formatted objects which can be passed to BERT.
#
# Each BERT model has a specific preprocessing model, make sure to use the proper one described on the BERT's model documentation.
#
# Note: BERT adds a "position embedding" to the token embedding of each input, and these come from a fixed-size lookup table. That imposes a max seq length of 512 (which is also a practical limit, due to the quadratic growth of attention computation). For this colab 128 is good enough.
# + id="KeHEYKXGqjAZ"
def make_bert_preprocess_model(sentence_features, seq_length=128):
"""Returns Model mapping string features to BERT inputs.
Args:
sentence_features: a list with the names of string-valued features.
seq_length: an integer that defines the sequence length of BERT inputs.
Returns:
A Keras Model that can be called on a list or dict of string Tensors
(with the order or names, resp., given by sentence_features) and
returns a dict of tensors for input to BERT.
"""
input_segments = [
tf.keras.layers.Input(shape=(), dtype=tf.string, name=ft)
for ft in sentence_features]
# Tokenize the text to word pieces.
bert_preprocess = hub.load(tfhub_handle_preprocess)
tokenizer = hub.KerasLayer(bert_preprocess.tokenize, name='tokenizer')
segments = [tokenizer(s) for s in input_segments]
# Optional: Trim segments in a smart way to fit seq_length.
# Simple cases (like this example) can skip this step and let
# the next step apply a default truncation to approximately equal lengths.
truncated_segments = segments
# Pack inputs. The details (start/end token ids, dict of output tensors)
# are model-dependent, so this gets loaded from the SavedModel.
packer = hub.KerasLayer(bert_preprocess.bert_pack_inputs,
arguments=dict(seq_length=seq_length),
name='packer')
model_inputs = packer(truncated_segments)
return tf.keras.Model(input_segments, model_inputs)
# + [markdown] id="kk5SS1bStmfP"
# Let's demonstrate the preprocessing model. You will create a test with two sentences input (input1 and input2). The output is what a BERT model would expect as input: `input_word_ids`, `input_masks` and `input_type_ids`.
# + id="BehJu3wLtAg-"
test_preprocess_model = make_bert_preprocess_model(['my_input1', 'my_input2'])
test_text = [np.array(['some random test sentence']),
np.array(['another sentence'])]
text_preprocessed = test_preprocess_model(test_text)
print('Keys : ', list(text_preprocessed.keys()))
print('Shape Word Ids : ', text_preprocessed['input_word_ids'].shape)
print('Word Ids : ', text_preprocessed['input_word_ids'][0, :16])
print('Shape Mask : ', text_preprocessed['input_mask'].shape)
print('Input Mask : ', text_preprocessed['input_mask'][0, :16])
print('Shape Type Ids : ', text_preprocessed['input_type_ids'].shape)
print('Type Ids : ', text_preprocessed['input_type_ids'][0, :16])
# + [markdown] id="qXU6bQWmNfhp"
# Let's take a look at the model's structure, paying attention to the two inputs you just defined.
# + id="a2_XrcVPFiz_"
tf.keras.utils.plot_model(test_preprocess_model)
# + [markdown] id="GRVVol5G9i0b"
# To apply the preprocessing in all the inputs from the dataset, you will use the `map` function from the dataset. The result is then cached for [performance](https://www.tensorflow.org/guide/data_performance#top_of_page).
# + id="1zhR-SVwx4_J"
AUTOTUNE = tf.data.experimental.AUTOTUNE
def load_dataset_from_tfds(in_memory_ds, info, split, batch_size,
bert_preprocess_model):
is_training = split.startswith('train')
dataset = tf.data.Dataset.from_tensor_slices(in_memory_ds[split])
num_examples = info.splits[split].num_examples
if is_training:
dataset = dataset.shuffle(num_examples)
dataset = dataset.repeat()
dataset = dataset.batch(batch_size)
dataset = dataset.map(lambda ex: (bert_preprocess_model(ex), ex['label']))
dataset = dataset.cache().prefetch(buffer_size=AUTOTUNE)
return dataset, num_examples
# + [markdown] id="pDNKfAXbDnJH"
# ## Define your model
#
# You are now ready to define your model for sentence or sentence pair classification by feeding the preprocessed inputs through the BERT encoder and putting a linear classifier on top (or other arrangement of layers as you prefer), and using dropout for regularization.
#
# Note: Here the model will be defined using the [Keras functional API](https://www.tensorflow.org/guide/keras/functional)
#
# + id="aksj743St9ga"
def build_classifier_model(num_classes):
inputs = dict(
input_word_ids=tf.keras.layers.Input(shape=(None,), dtype=tf.int32),
input_mask=tf.keras.layers.Input(shape=(None,), dtype=tf.int32),
input_type_ids=tf.keras.layers.Input(shape=(None,), dtype=tf.int32),
)
encoder = hub.KerasLayer(tfhub_handle_encoder, trainable=True, name='encoder')
net = encoder(inputs)['pooled_output']
net = tf.keras.layers.Dropout(rate=0.1)(net)
net = tf.keras.layers.Dense(num_classes, activation=None, name='classifier')(net)
return tf.keras.Model(inputs, net, name='prediction')
# + [markdown] id="TTa5VZssizDm"
# Let's try running the model on some preprocessed inputs.
# + id="e6mJ3WXhwUa8"
test_classifier_model = build_classifier_model(2)
bert_raw_result = test_classifier_model(text_preprocessed)
print(tf.sigmoid(bert_raw_result))
# + [markdown] id="RT7wuJ8HQv1l"
# Let's take a look at the model's structure. You can see the three BERT expected inputs.
# + id="4hfcmt7tNR_X"
tf.keras.utils.plot_model(test_classifier_model)
# + [markdown] id="1s0xpHS-XQcP"
# ## Choose a task from GLUE
#
# You are going to use a TensorFlow DataSet from the [GLUE](https://www.tensorflow.org/datasets/catalog/glue) benchmark suite.
#
# Colab lets you download these small datasets to the local filesystem, and the code below reads them entirely into memory, because the separate TPU worker host cannot access the local filesystem of the colab runtime.
#
# For bigger datasets, you'll need to create your own [Google Cloud Storage](https://cloud.google.com/storage) bucket and have the TPU worker read the data from there. You can learn more in the [TPU guide](https://www.tensorflow.org/guide/tpu#input_datasets).
#
# It's recommended to start with the CoLa dataset (for single sentence) or MRPC (for multi sentence) since these are small and don't take long to fine tune.
# + cellView="form" id="RhL__V2mwRNH"
tfds_name = 'glue/cola' #@param ['glue/cola', 'glue/sst2', 'glue/mrpc', 'glue/qqp', 'glue/mnli', 'glue/qnli', 'glue/rte', 'glue/wnli']
tfds_info = tfds.builder(tfds_name).info
sentence_features = list(tfds_info.features.keys())
sentence_features.remove('idx')
sentence_features.remove('label')
available_splits = list(tfds_info.splits.keys())
train_split = 'train'
validation_split = 'validation'
test_split = 'test'
if tfds_name == 'glue/mnli':
validation_split = 'validation_matched'
test_split = 'test_matched'
num_classes = tfds_info.features['label'].num_classes
num_examples = tfds_info.splits.total_num_examples
print(f'Using {tfds_name} from TFDS')
print(f'This dataset has {num_examples} examples')
print(f'Number of classes: {num_classes}')
print(f'Features {sentence_features}')
print(f'Splits {available_splits}')
with tf.device('/job:localhost'):
# batch_size=-1 is a way to load the dataset into memory
in_memory_ds = tfds.load(tfds_name, batch_size=-1, shuffle_files=True)
# The code below is just to show some samples from the selected dataset
print(f'Here are some sample rows from {tfds_name} dataset')
sample_dataset = tf.data.Dataset.from_tensor_slices(in_memory_ds[train_split])
labels_names = tfds_info.features['label'].names
print(labels_names)
print()
sample_i = 1
for sample_row in sample_dataset.take(5):
samples = [sample_row[feature] for feature in sentence_features]
print(f'sample row {sample_i}')
for sample in samples:
print(sample.numpy())
sample_label = sample_row['label']
print(f'label: {sample_label} ({labels_names[sample_label]})')
print()
sample_i += 1
# + [markdown] id="lFhjoYtsoVNF"
# The dataset also determines the problem type (classification or regression) and the appropriate loss function for training.
# + id="OWPOZE-L3AgE"
def get_configuration(glue_task):
loss = tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True)
if glue_task is 'glue/cola':
metrics = tfa.metrics.MatthewsCorrelationCoefficient()
else:
metrics = tf.keras.metrics.SparseCategoricalAccuracy(
'accuracy', dtype=tf.float32)
return metrics, loss
# + [markdown] id="EgJoTvo2DsWq"
# ## Train your model
#
# Finally, you can train the model end-to-end on the dataset you chose.
#
# First, create the preprocessing model with `make_bert_preprocess_model` to use when loading the dataset (train and evaluation splits) with TFDS (`load_dataset_from_tfds`). This auxiliary model runs on the host CPU, not the TPU, so it gets built and the SavedModel for preprocessing gets loaded outside the distribution strategy scope.
#
# The model fine-tuning goes inside the `strategy.scope` block so it can take advantage of the TPU.
#
# For the learning rate (`init_lr`), we use the same schedule as BERT pre-training: linear decay of a notional initial learning rate, prefixed with a linear warm-up phase over the first 10% of training steps (`num_warmup_steps`). In line with the BERT paper, the initial learning rate is smaller for fine-tuning (best of 5e-5, 3e-5, 2e-5).
# + id="AiU5_ioh_fEr"
epochs = 3
batch_size = 32
init_lr = 2e-5
print(f'Fine tuning {tfhub_handle_encoder} model')
bert_preprocess_model = make_bert_preprocess_model(sentence_features)
with strategy.scope():
# metric have to be created inside the strategy scope
metrics, loss = get_configuration(tfds_name)
train_dataset, train_data_size = load_dataset_from_tfds(
in_memory_ds, tfds_info, train_split, batch_size, bert_preprocess_model)
steps_per_epoch = train_data_size // batch_size
num_train_steps = steps_per_epoch * epochs
num_warmup_steps = num_train_steps // 10
validation_dataset, validation_data_size = load_dataset_from_tfds(
in_memory_ds, tfds_info, validation_split, batch_size,
bert_preprocess_model)
validation_steps = validation_data_size // batch_size
classifier_model = build_classifier_model(num_classes)
optimizer = optimization.create_optimizer(
init_lr=init_lr,
num_train_steps=num_train_steps,
num_warmup_steps=num_warmup_steps,
optimizer_type='adamw')
classifier_model.compile(optimizer=optimizer, loss=loss, metrics=[metrics])
classifier_model.fit(
x=train_dataset,
validation_data=validation_dataset,
steps_per_epoch=steps_per_epoch,
epochs=epochs,
validation_steps=validation_steps)
# + [markdown] id="Rtn7jewb6dg4"
# ## Export for inference
#
# You will create a final model that has the preprocessing part and the fine-tuned BERT we've just created.
#
# At inference time, preprocessing needs to be part of the model (because there is no longer a separate input queue as for training data that does it). Preprocessing is not just computation; it has its own resources (the vocab table) that must be attached to the Keras Model that is saved for export.
# This final assembly is what will be saved.
#
# You are going to save the model on colab and later you can download to keep it for the future (**View -> Table of contents -> Files**).
#
# + id="ShcvqJAgVera"
main_save_path = './my_models'
bert_type = tfhub_handle_encoder.split('/')[-2]
saved_model_name = f'{tfds_name.replace("/", "_")}_{bert_type}'
saved_model_path = os.path.join(main_save_path, saved_model_name)
preprocess_inputs = bert_preprocess_model.inputs
bert_encoder_inputs = bert_preprocess_model(preprocess_inputs)
bert_outputs = classifier_model(bert_encoder_inputs)
model_for_export = tf.keras.Model(preprocess_inputs, bert_outputs)
print(f'Saving {saved_model_path}')
# Save everything on the Colab host (even the variables from TPU memory)
save_options = tf.saved_model.SaveOptions(experimental_io_device='/job:localhost')
model_for_export.save(saved_model_path, include_optimizer=False, options=save_options)
# + [markdown] id="C2qyM9Q9z12v"
# ## Test the model
#
# The final step is testing the results of your exported model.
#
# Just to make some comparison, let's reload the model and test it using some inputs from the test split from the dataset.
# + id="BhI0_W0kbXji"
load_options = tf.saved_model.LoadOptions(experimental_io_device='/job:localhost')
reloaded_model = tf.saved_model.load(saved_model_path, options=load_options)
# + id="4yl-CEcDDXzX"
#@title Utility methods
def prepare(record):
model_inputs = [[record[ft]] for ft in sentence_features]
return model_inputs
def prepare_serving(record):
model_inputs = {ft: record[ft] for ft in sentence_features}
return model_inputs
def print_bert_results(test, bert_result, dataset_name):
bert_result_class = tf.argmax(bert_result, axis=1)[0]
if dataset_name == 'glue/cola':
print(f'sentence: {test[0].numpy()}')
if bert_result_class == 1:
print(f'This sentence is acceptable')
else:
print(f'This sentence is unacceptable')
elif dataset_name == 'glue/sst2':
print(f'sentence: {test[0]}')
if bert_result_class == 1:
print(f'This sentence has POSITIVE sentiment')
else:
print(f'This sentence has NEGATIVE sentiment')
elif dataset_name == 'glue/mrpc':
print(f'sentence1: {test[0]}')
print(f'sentence2: {test[1]}')
if bert_result_class == 1:
print(f'Are a paraphrase')
else:
print(f'Are NOT a paraphrase')
elif dataset_name == 'glue/qqp':
print(f'question1: {test[0]}')
print(f'question2: {test[1]}')
if bert_result_class == 1:
print(f'Questions are similar')
else:
print(f'Questions are NOT similar')
elif dataset_name == 'glue/mnli':
print(f'premise : {test[0]}')
print(f'hypothesis: {test[1]}')
if bert_result_class == 1:
print(f'This premise is NEUTRAL to the hypothesis')
elif bert_result_class == 2:
print(f'This premise CONTRADICTS the hypothesis')
else:
print(f'This premise ENTAILS the hypothesis')
elif dataset_name == 'glue/qnli':
print(f'question: {test[0]}')
print(f'sentence: {test[1]}')
if bert_result_class == 1:
print(f'The question is NOT answerable by the sentence')
else:
print(f'The question is answerable by the sentence')
elif dataset_name == 'glue/rte':
print(f'sentence1: {test[0]}')
print(f'sentence2: {test[1]}')
if bert_result_class == 1:
print(f'Sentence1 DOES NOT entails sentence2')
else:
print(f'Sentence1 entails sentence2')
elif dataset_name == 'glue/wnli':
print(f'sentence1: {test[0]}')
print(f'sentence2: {test[1]}')
if bert_result_class == 1:
print(f'Sentence1 DOES NOT entails sentence2')
else:
print(f'Sentence1 entails sentence2')
print(f'Bert raw results:{bert_result[0]}')
print()
# + [markdown] id="12VA4BcKuR7n"
# ### Test
# + id="dt-O94gcwbIi"
test_dataset = tf.data.Dataset.from_tensor_slices(in_memory_ds[test_split])
for test_row in test_dataset.shuffle(1000).map(prepare).take(5):
if len(sentence_features) == 1:
result = reloaded_model(test_row[0])
else:
result = reloaded_model(list(test_row))
print_bert_results(test_row, result, tfds_name)
# + [markdown] id="3cOmih754Y_M"
# If you want to use your model on [TF Serving](https://www.tensorflow.org/tfx/guide/serving), remember that it will call your SavedModel through one of its named signatures. Notice there are some small differences in the input. In Python, you can test them as follows:
# + id="b0vTQAXKN_K0"
serving_model = reloaded_model.signatures['serving_default']
for test_row in test_dataset.shuffle(1000).map(prepare_serving).take(5):
result = serving_model(**test_row)
# the 'prediction' key is the classifier's defined model name
print_bert_results(list(test_row.values()), result['prediction'], tfds_name)
# + [markdown] id="GOA5bX2g3wCW"
# You did it! Your saved model could be used for serving or simple inference in a process, with a simpler api with less code and easier to maintain.
#
# ## Next Steps
#
# Now that you've tried one of the base BERT models, you can try other ones to achieve more accuracy or maybe with smaller model versions.
#
# You can also try in other datasets.
|
site/en/tutorials/text/solve_glue_tasks_using_bert_on_tpu.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + id="WDXck-2aGkEz" colab_type="code" outputId="3d975313-11b6-4b25-c2a4-664a1ce66575" colab={"base_uri": "https://localhost:8080/", "height": 530}
# !pip install torch # framework
# !pip install --upgrade reedsolo
# !python --version
# !pip install librosa
# !pip install torchvision
# + id="hZ7QF4YbN1Pl" colab_type="code" outputId="ffec10f7-1bac-4f4c-df23-e6c4365d04cb" colab={"base_uri": "https://localhost:8080/", "height": 139}
from google.colab import drive
drive.mount('/content/drive')
# %cd /content/drive/My\ Drive/
# + id="yCY2rWEiOqDA" colab_type="code" colab={}
import numpy as np
import librosa
import librosa.display
import datetime
import matplotlib.pyplot as plt
from torch.nn.functional import binary_cross_entropy_with_logits, mse_loss
from torchvision import datasets, transforms
from IPython.display import clear_output
import torchvision
from torchvision.datasets.vision import VisionDataset
from torch.optim import Adam
from tqdm import notebook
import torch
import os
import os.path
import gc
import sys
from PIL import ImageFile, Image
ImageFile.LOAD_TRUNCATED_IMAGES = True
# + id="9_dX58S6hOoq" colab_type="code" colab={}
epochs = 4
data_depth = 4
hidden_size = 32
device = torch.device('cuda' if torch.cuda.is_available() else 'cpu')
LOAD_MODEL=False
PATH='/content/drive/My Drive/results/model/DenseEncoder_DenseDecoder_-0.001_2020-03-08_15:45:59.dat'
AUD_EXTENSIONS = ('.flac', '.wav', '.mp3', '.mp4')
audio = True
# + id="1S0bTwfpK3YG" colab_type="code" colab={}
import torch
import torch.nn.functional as F
from torch.autograd import Variable
import numpy as np
from math import exp
# -*- coding: utf-8 -*-
import zlib
from math import exp
import torch
from reedsolo import RSCodec
from torch.nn.functional import conv2d
rs = RSCodec(250)
def text_to_bits(text):
"""Convert text to a list of ints in {0, 1}"""
return bytearray_to_bits(text_to_bytearray(text))
def bits_to_text(bits):
"""Convert a list of ints in {0, 1} to text"""
return bytearray_to_text(bits_to_bytearray(bits))
def bytearray_to_bits(x):
"""Convert bytearray to a list of bits"""
result = []
for i in x:
bits = bin(i)[2:]
bits = '00000000'[len(bits):] + bits
result.extend([int(b) for b in bits])
return result
def bits_to_bytearray(bits):
"""Convert a list of bits to a bytearray"""
ints = []
for b in range(len(bits) // 8):
byte = bits[b * 8:(b + 1) * 8]
ints.append(int(''.join([str(bit) for bit in byte]), 2))
return bytearray(ints)
def text_to_bytearray(text):
"""Compress and add error correction"""
assert isinstance(text, str), "expected a string"
x = zlib.compress(text.encode("utf-8"))
x = rs.encode(bytearray(x))
return x
def bytearray_to_text(x):
"""Apply error correction and decompress"""
try:
text = rs.decode(x)
text = zlib.decompress(text)
return text.decode("utf-8")
except BaseException:
return False
def gaussian(window_size, sigma):
gauss = torch.Tensor([exp(-(x - window_size//2)**2/float(2*sigma**2)) for x in range(window_size)])
return gauss/gauss.sum()
def create_window(window_size, channel):
_1D_window = gaussian(window_size, 1.5).unsqueeze(1)
_2D_window = _1D_window.mm(_1D_window.t()).float().unsqueeze(0).unsqueeze(0)
window = Variable(_2D_window.expand(channel, 1, window_size, window_size).contiguous())
return window
def _ssim(img1, img2, window, window_size, channel, size_average = True):
mu1 = F.conv2d(img1, window, padding = window_size//2, groups = channel)
mu2 = F.conv2d(img2, window, padding = window_size//2, groups = channel)
mu1_sq = mu1.pow(2)
mu2_sq = mu2.pow(2)
mu1_mu2 = mu1*mu2
sigma1_sq = F.conv2d(img1*img1, window, padding = window_size//2, groups = channel) - mu1_sq
sigma2_sq = F.conv2d(img2*img2, window, padding = window_size//2, groups = channel) - mu2_sq
sigma12 = F.conv2d(img1*img2, window, padding = window_size//2, groups = channel) - mu1_mu2
C1 = 0.01**2
C2 = 0.03**2
ssim_map = ((2*mu1_mu2 + C1)*(2*sigma12 + C2))/((mu1_sq + mu2_sq + C1)*(sigma1_sq + sigma2_sq + C2))
if size_average:
return ssim_map.mean()
else:
return ssim_map.mean(1).mean(1).mean(1)
class SSIM(torch.nn.Module):
def __init__(self, window_size = 11, size_average = True):
super(SSIM, self).__init__()
self.window_size = window_size
self.size_average = size_average
self.channel = 1
self.window = create_window(window_size, self.channel)
def forward(self, img1, img2):
(_, channel, _, _) = img1.size()
if channel == self.channel and self.window.data.type() == img1.data.type():
window = self.window
else:
window = create_window(self.window_size, channel)
if img1.is_cuda:
window = window.cuda(img1.get_device())
window = window.type_as(img1)
self.window = window
self.channel = channel
return _ssim(img1, img2, window, self.window_size, channel, self.size_average)
def ssim(img1, img2, window_size = 11, size_average = True):
(_, channel, _, _) = img1.size()
window = create_window(window_size, channel)
if img1.is_cuda:
window = window.cuda(img1.get_device())
window = window.type_as(img1)
return _ssim(img1, img2, window, window_size, channel, size_average)
# + id="UO7Y-DuVKj38" colab_type="code" colab={}
import torch
from torch import nn
import numpy
class BasicEncoder(nn.Module):
"""
The BasicEncoder module takes an cover image and a data tensor and combines
them into a steganographic image.
"""
def _name(self):
return "BasicEncoder"
def _conv2d(self, in_channels, out_channels):
return nn.Conv2d(
in_channels=in_channels,
out_channels=out_channels,
kernel_size=3,
padding=1
)
def _build_models(self):
self.conv1 = nn.Sequential(
self._conv2d(3, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv2 = nn.Sequential(
self._conv2d(self.hidden_size + self.data_depth, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv3 = nn.Sequential(
self._conv2d(self.hidden_size, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv4 = nn.Sequential(
self._conv2d(self.hidden_size, 3),
)
return self.conv1, self.conv2, self.conv3, self.conv4
def __init__(self, data_depth, hidden_size):
super().__init__()
self.data_depth = data_depth
self.hidden_size = hidden_size
self._models = self._build_models()
self.name = self._name()
def forward(self, image, data):
x = self._models[0](image)
x_1 = self._models[1](torch.cat([x] + [data], dim=1))
x_2 = self._models[2](x_1)
x_3 = self._models[3](x_2)
return x_3
class ResidualEncoder(BasicEncoder):
def _name(self):
return "ResidualEncoder"
def forward(self, image, data):
return image + super().forward(self, image, data)
class DenseEncoder(BasicEncoder):
def _name(self):
return "DenseEncoder"
def _build_models(self):
self.conv1 = super()._build_models()[0]
self.conv2 = super()._build_models()[1]
self.conv3 = nn.Sequential(
self._conv2d(self.hidden_size * 2 +
self.data_depth, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv4 = nn.Sequential(
self._conv2d(self.hidden_size * 3 + self.data_depth, 3)
)
return self.conv1, self.conv2, self.conv3, self.conv4
def forward(self, image, data):
x = self._models[0](image)
x_list = [x]
x_1 = self._models[1](torch.cat(x_list+[data], dim=1))
x_list.append(x_1)
x_2 = self._models[2](torch.cat(x_list+[data], dim=1))
x_list.append(x_2)
x_3 = self._models[3](torch.cat(x_list+[data], dim=1))
x_list.append(x_3)
return image + x_3
# + id="gPc3PmGwKoZe" colab_type="code" colab={}
import torch
from torch import nn
#from torch.nn import Sigmoid
#from torch.distributions import Bernoulli
class BasicDecoder(nn.Module):
"""
The BasicDecoder module takes an steganographic image and attempts to decode
the embedded data tensor.
Input: (N, 3, H, W)
Output: (N, D, H, W)
"""
def _name(self):
return "BasicDecoder"
def _conv2d(self, in_channels, out_channels):
return nn.Conv2d(
in_channels=in_channels,
out_channels=out_channels,
kernel_size=3,
padding=1
)
def _build_models(self):
self.conv1 = nn.Sequential(
self._conv2d(3, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv2 = nn.Sequential(
self._conv2d(self.hidden_size, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv3 = nn.Sequential(
self._conv2d(self.hidden_size, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv4 = nn.Sequential(
self._conv2d(self.hidden_size, self.data_depth),
#nn.Sigmoid(),
)
return self.conv1, self.conv2, self.conv3, self.conv4
def forward(self, image):
x = self._models[0](image)
x_1 = self._models[1](x)
x_2 = self._models[2](x_1)
x_3 = self._models[3](x_2)
#x_4 = Bernoulli(x_3).sample()
return x_3
def __init__(self, data_depth, hidden_size):
super().__init__()
self.data_depth = data_depth
self.hidden_size = hidden_size
self._models = self._build_models()
self.name = self._name()
class DenseDecoder(BasicDecoder):
def _name(self):
return "DenseDecoder"
def _build_models(self):
self.conv1 = super()._build_models()[0]
self.conv2 = super()._build_models()[1]
self.conv3 = nn.Sequential(
self._conv2d(self.hidden_size * 2, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size)
)
self.conv4 = nn.Sequential(
self._conv2d(self.hidden_size * 3, self.data_depth),
#nn.Sigmoid(),
)
return self.conv1, self.conv2, self.conv3, self.conv4
def forward(self, image):
x = self._models[0](image)
x_list = [x]
x_1 = self._models[1](torch.cat(x_list, dim=1))
x_list.append(x_1)
x_2 = self._models[2](torch.cat(x_list, dim=1))
x_list.append(x_2)
x_3 = self._models[3](torch.cat(x_list, dim=1))
x_list.append(x_3)
return x_3
# + id="he1SeqWRKtb8" colab_type="code" colab={}
import torch
from torch import nn
class BasicCritic(nn.Module):
"""
The BasicCritic module takes an image and predicts whether it is a cover
image or a steganographic image (N, 1).
Input: (N, 3, H, W)
Output: (N, 1)
"""
def _name(self):
return "BasicCritic"
def _conv2d(self, in_channels, out_channels):
return nn.Conv2d(
in_channels=in_channels,
out_channels=out_channels,
kernel_size=3
)
def _build_models(self):
self.conv1 = nn.Sequential(
self._conv2d(3, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv2 = nn.Sequential(
self._conv2d(self.hidden_size, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv3 = nn.Sequential(
self._conv2d(self.hidden_size, self.hidden_size),
nn.LeakyReLU(inplace=True),
nn.BatchNorm2d(self.hidden_size),
)
self.conv4 = nn.Sequential(
self._conv2d(self.hidden_size, 1)
)
return self.conv1,self.conv2,self.conv3,self.conv4
def __init__(self, hidden_size):
super().__init__()
self.hidden_size = hidden_size
self._models = self._build_models()
self.name = self._name()
def forward(self, image):
x = self._models[0](image)
x_1 = self._models[1](x)
x_2 = self._models[2](x_1)
x_3 = self._models[3](x_2)
return torch.mean(x_3.view(x_3.size(0), -1), dim=1)
# + id="YVLWLMetCrGe" colab_type="code" colab={}
def plot(name, train_epoch, values, path, save):
clear_output(wait=True)
plt.close('all')
fig = plt.figure()
fig = plt.ion()
fig = plt.subplot(1, 1, 1)
fig = plt.title('epoch: %s -> %s: %s' % (train_epoch, name, values[-1]))
fig = plt.ylabel(name)
fig = plt.xlabel('validation_set')
fig = plt.plot(values)
fig = plt.grid()
get_fig = plt.gcf()
fig = plt.draw() # draw the plot
fig = plt.pause(1) # show it for 1 second
if save:
now = datetime.datetime.now()
get_fig.savefig('%s/%s_%.3f_%d_%s.png' %
(path, name, train_epoch, values[-1], now.strftime("%Y-%m-%d_%H:%M:%S")))
# + id="mZCf5oX12gWH" colab_type="code" colab={}
def test(encoder,decoder,data_depth,train_epoch,cover,payload):
# %matplotlib inline
generated = encoder.forward(cover, payload)
decoded = decoder.forward(generated)
decoder_loss = binary_cross_entropy_with_logits(decoded, payload)
decoder_acc = (decoded >= 0.0).eq(
payload >= 0.5).sum().float() / payload.numel() # .numel() calculate the number of element in a tensor
print("Decoder loss: %.3f"% decoder_loss.item())
print("Decoder acc: %.3f"% decoder_acc.item())
f, ax = plt.subplots(1, 2)
plt.title("%s_%s"%(encoder.name,decoder.name))
cover=np.transpose(np.squeeze(cover.cpu()), (1, 2, 0))
ax[0].imshow(cover)
ax[0].axis('off')
generated=np.transpose(np.squeeze((generated.cpu()).detach().numpy()), (1, 2, 0))
ax[1].imshow(generated)
ax[1].axis('off')
now = datetime.datetime.now()
print("payload :")
print(payload)
print("decoded :")
decoded[decoded<0]=0
decoded[decoded>0]=1
print(decoded)
now = datetime.datetime.now()
plt.savefig('results/samples/%s_%s_%d_%.3f_%d_%s.png' %
(encoder.name,decoder.name, data_depth,decoder_acc, train_epoch, now.strftime("%Y-%m-%d_%H:%M:%S")))
# + id="zHGbk-t3imrL" colab_type="code" colab={}
def save_model(encoder,decoder,critic,en_de_optimizer,cr_optimizer,metrics,ep):
now = datetime.datetime.now()
cover_score = metrics['val.cover_score'][-1]
name = "%s_%s_%+.3f_%s.dat" % (encoder.name,decoder.name,cover_score,
now.strftime("%Y-%m-%d_%H:%M:%S"))
fname = os.path.join('.', 'results/model', name)
states = {
'state_dict_critic': critic.state_dict(),
'state_dict_encoder': encoder.state_dict(),
'state_dict_decoder': decoder.state_dict(),
'en_de_optimizer': en_de_optimizer.state_dict(),
'cr_optimizer': cr_optimizer.state_dict(),
'metrics': metrics,
'train_epoch': ep,
'date': now.strftime("%Y-%m-%d_%H:%M:%S"),
}
torch.save(states, fname)
path='results/plots/train_%s_%s_%s'% (encoder.name,decoder.name,now.strftime("%Y-%m-%d_%H:%M:%S"))
try:
os.mkdir(os.path.join('.', path))
except Exception as error:
print(error)
plot('encoder_mse', ep, metrics['val.encoder_mse'], path, True)
plot('decoder_loss', ep, metrics['val.decoder_loss'], path, True)
plot('decoder_acc', ep, metrics['val.decoder_acc'], path, True)
plot('cover_score', ep, metrics['val.cover_score'], path, True)
plot('generated_score', ep, metrics['val.generated_score'], path, True)
plot('ssim', ep, metrics['val.ssim'], path, True)
plot('psnr', ep, metrics['val.psnr'], path, True)
plot('bpp', ep, metrics['val.bpp'], path, True)
# + id="UMqZ9W6Ug6dR" colab_type="code" colab={}
def fit_gan(encoder,decoder,critic,en_de_optimizer,cr_optimizer,metrics,train_loader,valid_loader):
for ep in range(epochs):
print("Epoch %d" %(ep+1))
for cover, _ in notebook.tqdm(train_loader):
gc.collect()
cover = cover.to(device)
N, _, H, W = cover.size()
# sampled from the discrete uniform distribution over 0 to 2
payload = torch.zeros((N, data_depth, H, W),
device=device).random_(0, 2)
generated = encoder.forward(cover, payload)
cover_score = torch.mean(critic.forward(cover))
generated_score = torch.mean(critic.forward(generated))
cr_optimizer.zero_grad()
(cover_score - generated_score).backward(retain_graph=False)
cr_optimizer.step()
for p in critic.parameters():
p.data.clamp_(-0.1, 0.1)
metrics['train.cover_score'].append(cover_score.item())
metrics['train.generated_score'].append(generated_score.item())
for cover, _ in notebook.tqdm(train_loader):
gc.collect()
cover = cover.to(device)
N, _, H, W = cover.size()
# sampled from the discrete uniform distribution over 0 to 2
payload = torch.zeros((N, data_depth, H, W),
device=device).random_(0, 2)
generated = encoder.forward(cover, payload)
decoded = decoder.forward(generated)
encoder_mse = mse_loss(generated, cover)
decoder_loss = binary_cross_entropy_with_logits(decoded, payload)
decoder_acc = (decoded >= 0.0).eq(
payload >= 0.5).sum().float() / payload.numel()
generated_score = torch.mean(critic.forward(generated))
en_de_optimizer.zero_grad()
(100 * encoder_mse + decoder_loss +
generated_score).backward() # Why 100?
en_de_optimizer.step()
metrics['train.encoder_mse'].append(encoder_mse.item())
metrics['train.decoder_loss'].append(decoder_loss.item())
metrics['train.decoder_acc'].append(decoder_acc.item())
for cover, _ in notebook.tqdm(valid_loader):
gc.collect()
cover = cover.to(device)
N, _, H, W = cover.size()
# sampled from the discrete uniform distribution over 0 to 2
payload = torch.zeros((N, data_depth, H, W),
device=device).random_(0, 2)
generated = encoder.forward(cover, payload)
decoded = decoder.forward(generated)
encoder_mse = mse_loss(generated, cover)
decoder_loss = binary_cross_entropy_with_logits(decoded, payload)
decoder_acc = (decoded >= 0.0).eq(
payload >= 0.5).sum().float() / payload.numel()
generated_score = torch.mean(critic.forward(generated))
cover_score = torch.mean(critic.forward(cover))
metrics['val.encoder_mse'].append(encoder_mse.item())
metrics['val.decoder_loss'].append(decoder_loss.item())
metrics['val.decoder_acc'].append(decoder_acc.item())
metrics['val.cover_score'].append(cover_score.item())
metrics['val.generated_score'].append(generated_score.item())
metrics['val.ssim'].append(
ssim(cover, generated).item())
metrics['val.psnr'].append(
10 * torch.log10(4 / encoder_mse).item())
metrics['val.bpp'].append(
data_depth * (2 * decoder_acc.item() - 1))
print('encoder_mse: %.3f - decoder_loss: %.3f - decoder_acc: %.3f - cover_score: %.3f - generated_score: %.3f - ssim: %.3f - psnr: %.3f - bpp: %.3f'
%(encoder_mse.item(),decoder_loss.item(),decoder_acc.item(),cover_score.item(),generated_score.item(), ssim(cover, generated).item(),10 * torch.log10(4 / encoder_mse).item(),data_depth * (2 * decoder_acc.item() - 1)))
save_model(encoder,decoder,critic,en_de_optimizer,cr_optimizer,metrics,ep)
# + id="JnpEj9wjN-G-" colab_type="code" colab={}
def fig2data ( fig ):
"""
@brief Convert a Matplotlib figure to a 4D numpy array with RGB channels and return it
@param fig a matplotlib figure
@return a numpy 3D array of RGB values
"""
# draw the renderer
fig.canvas.draw ( )
# Get the RGB buffer from the figure
w,h = fig.canvas.get_width_height()
buf = np.frombuffer ( fig.canvas.tostring_rgb(), dtype=np.uint8 )
buf.shape = ( w, h,3 )
return buf
def has_file_allowed_extension(filename, extensions):
"""Checks if a file is an allowed extension.
Args:
filename (string): path to a file
extensions (tuple of strings): extensions to consider (lowercase)
Returns:
bool: True if the filename ends with one of given extensions
"""
return filename.lower().endswith(extensions)
def is_image_file(filename):
"""Checks if a file is an allowed image extension.
Args:
filename (string): path to a file
Returns:
bool: True if the filename ends with a known image extension
"""
return has_file_allowed_extension(filename, AUD_EXTENSIONS)
def make_dataset(dir, class_to_idx, extensions=None, is_valid_file=None):
images = []
dir = os.path.expanduser(dir)
if not ((extensions is None) ^ (is_valid_file is None)):
raise ValueError("Both extensions and is_valid_file cannot be None or not None at the same time")
if extensions is not None:
def is_valid_file(x):
return has_file_allowed_extension(x, extensions)
for target in sorted(class_to_idx.keys()):
d = os.path.join(dir, target)
if not os.path.isdir(d):
continue
for root, _, fnames in sorted(os.walk(d, followlinks=True)):
for fname in sorted(fnames):
path = os.path.join(root, fname)
if is_valid_file(path):
item = (path, class_to_idx[target])
images.append(item)
return images
def pil_loader(path):
# open path as file to avoid ResourceWarning (https://github.com/python-pillow/Pillow/issues/835)
with open(path, 'rb') as f:
y, sr = librosa.load(f.name) # your file
S = librosa.feature.melspectrogram(y=y, sr=sr, n_mels=128, fmax=8000)
img = librosa.display.specshow(librosa.power_to_db(S, ref=np.max), fmax=8000).get_figure()
f=fig2data(img)
w, h, d = f.shape
img=Image.frombytes( "RGB", ( w ,h ), f.tostring( ) )
return img.convert('RGB')
def accimage_loader(path):
import accimage
try:
return accimage.Image(path)
except IOError:
# Potentially a decoding problem, fall back to PIL.Image
return pil_loader(path)
def default_loader(path):
from torchvision import get_image_backend
if get_image_backend() == 'accimage':
return accimage_loader(path)
else:
return pil_loader(path)
class AudToImageFolder(datasets.DatasetFolder):
"""A generic data loader where the images are arranged in this way: ::
root/dog/xxx.png
root/dog/xxy.png
root/dog/xxz.png
root/cat/123.png
root/cat/nsdf3.png
root/cat/asd932_.png
Args:
root (string): Root directory path.
transform (callable, optional): A function/transform that takes in an PIL image
and returns a transformed version. E.g, ``transforms.RandomCrop``
target_transform (callable, optional): A function/transform that takes in the
target and transforms it.
loader (callable, optional): A function to load an image given its path.
is_valid_file (callable, optional): A function that takes path of an Image file
and check if the file is a valid file (used to check of corrupt files)
Attributes:
classes (list): List of the class names.
class_to_idx (dict): Dict with items (class_name, class_index).
imgs (list): List of (image path, class_index) tuples
"""
def __init__(self, root, transform=None, target_transform=None,
loader=default_loader, is_valid_file=None):
super(AudToImageFolder, self).__init__(root, loader, AUD_EXTENSIONS if is_valid_file is None else None,
transform=transform,
target_transform=target_transform,
is_valid_file=is_valid_file)
self.imgs = self.samples
# + colab_type="code" outputId="9fe79d2b-36d9-44f0-f076-b72be4603cac" id="cQLj2IDbTvOS" colab={"base_uri": "https://localhost:8080/", "height": 117, "referenced_widgets": ["6bea0c9de09640e3993372d36775e552", "67baca1aec9f452e8f3e4ff08896a4f4", "787da502c8df4c78bd7b804e700671f0", "4cb603612a52484cac4af3803ef18b46", "<KEY>", "b3a441bc8a6b422085735ec55507f283", "82540accf51f4368adf429f87e8ab6e2", "89f6cf8573844db59a71bbf53a7befa6"]}
if __name__ == '__main__':
for func in [
lambda: os.mkdir(os.path.join('.', 'results')),
lambda: os.mkdir(os.path.join('.', 'results/model')),
lambda: os.mkdir(os.path.join('.', 'results/plots'))]: # create directories
try:
func()
except Exception as error:
print(error)
continue
METRIC_FIELDS = [
'val.encoder_mse',
'val.decoder_loss',
'val.decoder_acc',
'val.cover_score',
'val.generated_score',
'val.ssim',
'val.psnr',
'val.bpp',
'train.encoder_mse',
'train.decoder_loss',
'train.decoder_acc',
'train.cover_score',
'train.generated_score',
]
mu = [.5, .5, .5]
sigma = [.5, .5, .5]
transform = transforms.Compose([transforms.RandomHorizontalFlip(),
transforms.RandomCrop(
360, pad_if_needed=True),
transforms.ToTensor(),
transforms.Normalize(mu, sigma)])
if audio:
data_dir="LibriSpeech/devclean" # directory to audio
train_set = AudToImageFolder(os.path.join(
data_dir, "train/"), transform=transform)
train_loader = torch.utils.data.DataLoader(
train_set, batch_size=4, shuffle=True)
valid_set = AudToImageFolder(os.path.join(
data_dir, "val/"), transform=transform)
valid_loader = torch.utils.data.DataLoader(
valid_set, batch_size=4, shuffle=False)
else:
data_dir = 'div2k'
train_set = datasets.ImageFolder(os.path.join(
data_dir, "train/"), transform=transform)
train_loader = torch.utils.data.DataLoader(
train_set, batch_size=4, shuffle=True)
valid_set = datasets.ImageFolder(os.path.join(
data_dir, "val/"), transform=transform)
valid_loader = torch.utils.data.DataLoader(
valid_set, batch_size=4, shuffle=False)
encoder = DenseEncoder(data_depth, hidden_size).to(device)
decoder = DenseDecoder(data_depth, hidden_size).to(device)
critic = BasicCritic(hidden_size).to(device)
cr_optimizer = Adam(critic.parameters(), lr=1e-4)
en_de_optimizer = Adam(list(decoder.parameters()) +
list(encoder.parameters()), lr=1e-4)
metrics = {field: list() for field in METRIC_FIELDS}
if LOAD_MODEL:
if torch.cuda.is_available():
checkpoint = torch.load(PATH)
else:
checkpoint = torch.load(PATH, map_location=lambda storage, loc: storage)
critic.load_state_dict(checkpoint['state_dict_critic'])
encoder.load_state_dict(checkpoint['state_dict_encoder'])
decoder.load_state_dict(checkpoint['state_dict_decoder'])
en_de_optimizer.load_state_dict(checkpoint['en_de_optimizer'])
cr_optimizer.load_state_dict(checkpoint['cr_optimizer'])
metrics=checkpoint['metrics']
ep=checkpoint['train_epoch']
date=checkpoint['date']
critic.train()
encoder.train()
decoder.train()
print('GAN loaded')
print(critic)
print(encoder)
print(decoder)
print(en_de_optimizer)
print(cr_optimizer)
print(date)
else:
fit_gan(encoder,decoder,critic,en_de_optimizer,cr_optimizer,metrics,train_loader,valid_loader)
# + id="NFFfYzJVK-I6" colab_type="code" colab={}
cover, _ = next(iter(valid_set))
_, H, W = cover.size()
cover = cover[None].to(device)
payload = torch.zeros((1, data_depth, H, W),device=device).random_(0, 2)
test(encoder,decoder,data_depth,epochs,cover,payload)
# + id="SjwtoNP4ed9x" colab_type="code" colab={}
|
S_GAN.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: rl
# language: python
# name: rl
# ---
# +
# %matplotlib inline
import gym
import matplotlib
import numpy as np
import sys
from collections import defaultdict
if "../" not in sys.path:
sys.path.append("../")
from lib.envs.blackjack import BlackjackEnv
from lib import plotting
matplotlib.style.use('ggplot')
# -
env = BlackjackEnv()
def mc_prediction(policy, env, num_episodes, discount_factor=1.0):
"""
Monte Carlo prediction algorithm. Calculates the value function
for a given policy using sampling.
Args:
policy: A function that maps an observation to action probabilities.
env: OpenAI gym environment.
num_episodes: Number of episodes to sample.
discount_factor: Gamma discount factor.
Returns:
A dictionary that maps from state -> value.
The state is a tuple and the value is a float.
"""
# Keeps track of sum and count of returns for each state
# to calculate an average. We could use an array to save all
# returns (like in the book) but that's memory inefficient.
returns_sum = defaultdict(float)
returns_count = defaultdict(float)
# The final value function
V = defaultdict(float)
# Implement this!
for i_episode in range(1,num_episodes+1):
if i_episode % 1000 == 0:
print("\rEpisode {}/{}.".format(i_episode, num_episodes), end="")
sys.stdout.flush()
episodes=[]
state = env.reset()
for t in range(100):
action = sample_policy(state)
next_state, reward, done, _ = env.step(action)
episodes.append((state,action,reward))
if done:
break
state=next_state
G=0.0
episodes.reverse()
"""
for ind,ep in enumerate(episodes[:-2]):
G=discount_factor*G+ep[2]
b_ep=[i[0] for i in episodes[(ind+1):]]
if ep[0] not in b_ep:
returns_sum[ep[0]]+=G
returns_count[ep[0]]+=1.0
ep=episodes[-1]
G=discount_factor*G+ep[2]
returns_sum[ep[0]]+=G
returns_count[ep[0]]+=1.0
"""
for ind,ep in enumerate(episodes):
G=discount_factor*G+ep[2]
returns_sum[ep[0]]+=G
returns_count[ep[0]]+=1.0
for v in returns_sum:
V[v]=returns_sum[v]/returns_count[v]
return V
def standard_mc_prediction(policy, env, num_episodes, discount_factor=1.0):
"""
Monte Carlo prediction algorithm. Calculates the value function
for a given policy using sampling.
Args:
policy: A function that maps an observation to action probabilities.
env: OpenAI gym environment.
num_episodes: Number of episodes to sample.
discount_factor: Gamma discount factor.
Returns:
A dictionary that maps from state -> value.
The state is a tuple and the value is a float.
"""
# Keeps track of sum and count of returns for each state
# to calculate an average. We could use an array to save all
# returns (like in the book) but that's memory inefficient.
returns_sum = defaultdict(float)
returns_count = defaultdict(float)
# The final value function
V = defaultdict(float)
for i_episode in range(1, num_episodes + 1):
# Print out which episode we're on, useful for debugging.
if i_episode % 1000 == 0:
print("\rEpisode {}/{}.".format(i_episode, num_episodes), end="")
sys.stdout.flush()
# Generate an episode.
# An episode is an array of (state, action, reward) tuples
episode = []
state = env.reset()
for t in range(100):
action = policy(state)
next_state, reward, done, _ = env.step(action)
episode.append((state, action, reward))
if done:
break
state = next_state
# Find all states the we've visited in this episode
# We convert each state to a tuple so that we can use it as a dict key
states_in_episode = set([tuple(x[0]) for x in episode])
for state in states_in_episode:
# Find the first occurance of the state in the episode
first_occurence_idx = next(i for i,x in enumerate(episode) if x[0] == state)
# Sum up all rewards since the first occurance
G = sum([x[2]*(discount_factor**i) for i,x in enumerate(episode[first_occurence_idx:])])
# Calculate average return for this state over all sampled episodes
returns_sum[state] += G
returns_count[state] += 1.0
V[state] = returns_sum[state] / returns_count[state]
return V
def sample_policy(observation):
"""
A policy that sticks if the player score is > 20 and hits otherwise.
"""
score, dealer_score, usable_ace = observation
return 0 if score >= 20 else 1
# +
V_10k= mc_prediction(sample_policy, env, num_episodes=10000)
plotting.plot_value_function(V_10k, title="10,000 Steps")
V_500k = mc_prediction(sample_policy, env, num_episodes=500000)
plotting.plot_value_function(V_500k, title="500,000 Steps")
# -
|
MC/MC Prediction.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <font style="font-size:96px; font-weight:bolder; color:#0040a0"><img src="http://montage.ipac.caltech.edu/docs/M51_logo.png" alt="M" style="float: left; padding: 25px 30px 25px 0px;" /></font>
#
# <i><b>Montage</b> Montage is an astronomical image toolkit with components for reprojection, background matching, coaddition and visualization of FITS files. It can be used as a set of command-line tools (Linux, OS X and Windows), C library calls (Linux and OS X) and as Python binary extension modules.
#
# The Montage source is written in ANSI-C and code can be downloaded from GitHub ( https://github.com/Caltech-IPAC/Montage ). The Python package can be installed from PyPI ("</i>pip install MontagePy<i>"). The package has no external dependencies. See http://montage.ipac.caltech.edu/ for details on the design and applications of Montage.
#
# # MontagePy.main modules: mFitExec
#
# The Montage modules are generally used as steps in a workflow to create a mosaic of a set of input images. These steps are: determine the geometry of the mosaic on the sky, reproject the images to a common frame and spatial sampling; rectify the backgrounds to a common level, and coadd the images into a mosaic. This page illustrates the use of one Montage module, mFitExec, which fits planes to a set of images (most often overlap differences).
#
# Visit <a href="Mosaic.ipynb">Building a Mosaic with Montage</a> to see how mFitExec is used as part of a workflow to creage a mosaic (or the <a href="Mosaic_oneshot.ipynb"> one shot </a> version if you just want to see the commands). See the complete list of Montage Notebooks <a href="http://montage.ipac.caltech.edu/MontageNotebooks">here</a>.
#
# +
from MontagePy.main import mFitExec, mViewer
help(mFitExec)
# -
# ## mFitExec Example
#
# mFitExec is a general utility for fitting a set of image differences with a plane. It's most common use is in the process of evaluating overlap area in a set of images as part of background rectification. That process consists of taking the output of the overlaps calculation (mOverlaps) and for each overlap pair doing the difference (mDiff), fitting it to determine offset between those two images (mFitplane).
#
# There are two ways to do this in bulk. We can either generate all the differences first, then fit them all or we can loop over the differences, generating and fitting that difference before moving on to the next. This first approach uses more disks space as all the differences have to be stored in between steps. The second can clean up as it goes.
#
# Obviously the second approach is preferable unless you want to keep the differences (possibly for quality evaluation).
#
# mFitExec is part of the first approach. It runs mFitplane on difference file from the input list and stores the result in a table. This table will usually be handed to mBgModel to determine what corrections to make to each image to minimize background differences.
#
# At this point in the processing, we already have a set of "like" images (same projection), have made a list of all the overlaps (normally using mOverlaps) and generated difference images. mFitExec loops over these images and fits a plane to each one.
# +
rtn = mFitExec('M17/diffs.tbl',
'work/M17/fits.tbl',
'M17/diffs')
print(rtn)
# -
# The output is a set of planes representing the difference between two original images. This set will be sent to mBgModel for modelling into a set of corrections to the original images to bring the backgrounds in line.
# +
import os
import numpy as np
import pandas as pd
from astropy.io import ascii
ipactable = ascii.read('work/M17/fits.tbl').to_pandas()
ipactable
# -
# <p/>
#
# ## Error Handling
#
# If mFitExec encounters an error, the return structure will just have two elements: a status of 1 ("error") and a message string that tries to diagnose the reason for the error.
#
# For instance, if the user specifies a table that doesn't exist:
#
rtn = mFitExec('M17/unknown.tbl',
'work/M17/fits.tbl',
'M17/diffs')
print(rtn)
#
#
# ## Classic Montage: mFitExec as a Stand-Alone Program
#
# ### mFitExec Unix/Windows Command-line Arguments
#
# <p>mFitExec can also be run as a command-line tool in Linux, OS X, and Windows:</p>
#
# <p><tt>
# <b>Usage:</b> mFitExec [-d] [-l(evel-only)] [-s statusfile] diffs.tbl fits.tbl diffdir
# </tt></p>
# <p> </p>
# <p>If you are writing in C/C++, mFitExec can be accessed as a library function:</p>
#
# <pre>
# /*-*****************************************************************/
# /* */
# /* mFitExec */
# /* */
# /* After mDiffExec has been run using the table of overlaps found */
# /* by mOverlaps, use this executive to run mFitplane on each of */
# /* the differences. Write the fits to a file to be used by */
# /* mBModel. */
# /* */
# /* char *tblfile Table file list of images to fit. */
# /* char *fitfile Table file for output difference fits info. */
# /* char *diffdir Directory for temporary output diff files. */
# /* int levelOnly Flag to fit level of diff only, not slopes. */
# /* int debug Debug flag. */
# /* */
# /*******************************************************************/
#
# struct mFitExecReturn *mFitExec(char *tblfile, char *fitfile, char *diffdir, int levelOnly, int debugin)
# </pre>
# <p><b>Return Structure</b></p>
# <pre>
# struct mFitExecReturn
# {
# int status; // Return status (0: OK, 1:ERROR)
# char msg [1024]; // Return message (for error return)
# char json[4096]; // Return parameters as JSON string
# int count; // Number of differences
# int failed; // Number of fits to differences that failed
# int warning; // Number of fits to differences that produced warnings
# int missing; // Number of missing difference images
# };
# </pre>
|
mFitExec.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="KrGn7nFRNEuv" outputId="69cc38a0-7e73-425b-b268-de6933757e21"
import pandas as pd
import numpy as np
import pylab as plt
from sklearn import metrics
from sklearn import cluster
from sklearn import model_selection
from sklearn.preprocessing import StandardScaler, PowerTransformer
from sklearn.compose import TransformedTargetRegressor
from sklearn import pipeline
from sklearn.compose import ColumnTransformer
from datetime import datetime as dt , time , date
import datetime
import seaborn as sns
from dateutil.parser import parse
import scipy.stats as stats
from sklearn.cluster import KMeans , MiniBatchKMeans
import lightgbm as lgb
# %matplotlib inline
SEED = 70
# + [markdown] colab_type="text" id="mLsLLmI1NEu7"
# # Load data
# + colab={"base_uri": "https://localhost:8080/", "height": 122} colab_type="code" id="NdDdJBY4NKQj" outputId="0c6c2051-7df5-42fa-b93e-0db5ebf21f14"
from google.colab import drive
drive.mount('/content/drive')
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="1OZ2XY8ANIls" outputId="6dc0afaa-552d-417b-96fb-1d94aaf22735"
# !unzip -P trrfd "./drive/My Drive/UmojaHackYassir.zip"
# + colab={"base_uri": "https://localhost:8080/", "height": 235} colab_type="code" id="7B7mMjyINEu7" outputId="cf20ac14-305c-4925-cf2d-dfe30a0d2e73"
# load training
training = pd.read_csv('UmojaHack25July/Train.csv').set_index('ID')
training['Timestamp'] = pd.to_datetime(training['Timestamp'])
# testing data
testing = pd.read_csv('UmojaHack25July/Test.csv').set_index('ID')
testing['Timestamp'] = pd.to_datetime(testing['Timestamp'])
# weather data
weather_df = pd.read_csv('UmojaHack25July/Weather.csv')
weather_df['date'] = pd.to_datetime(weather_df['date'])
# + [markdown] colab_type="text" id="GiRa3Ez0Ka7h"
# # Adding datetime features
# + colab={} colab_type="code" id="W5oT-R8IQn9P"
def add_date_features(df,timestamp_id):
df.loc[:, 'pickup_weekday'] = df[timestamp_id].dt.weekday
df.loc[:, 'pickup_hour_weekofyear'] = df[timestamp_id].dt.weekofyear
df.loc[:, 'pickup_hour'] = df[timestamp_id].dt.hour
df.loc[:, 'pickup_minute'] = df[timestamp_id].dt.minute
df.loc[:, 'pickup_dt'] = (df[timestamp_id] - df[timestamp_id].min()).dt.total_seconds()
df.loc[:, 'pickup_week_hour'] = df['pickup_weekday'] * 24 + df['pickup_hour']
def get_season(now):
Y=2000
seasons = [(1, (date(Y, 1, 1), date(Y, 3, 20))),
(2, (date(Y, 3, 21), date(Y, 6, 20))),
(3, (date(Y, 6, 21), date(Y, 9, 22))),
(4, (date(Y, 9, 23), date(Y, 12, 20))),
(1, (date(Y, 12, 21), date(Y, 12, 31)))]
if isinstance(now, datetime.datetime):
now = now.date()
now = now.replace(year=Y)
return next(season for season, (start, end) in seasons
if start <= now <= end)
df['day_of_week']=df[timestamp_id].dt.dayofweek
df['day_of_week']= df['day_of_week'].astype('category')
df['is_month_end']=df[timestamp_id].dt.is_month_end
df['is_month_end']= df['is_month_end'].astype('category')
df['is_month_start']=df[timestamp_id].dt.is_month_start
df['is_month_start']= df['is_month_start'].astype('category')
df['day_of_year']=df[timestamp_id].dt.dayofyear
df['day_of_year']= df['day_of_year'].astype('category')
df['is_quarter_end']=df[timestamp_id].dt.is_quarter_end
df['is_quarter_end']= df['is_quarter_end'].astype('category')
df['is_quarter_start']=df[timestamp_id].dt.is_quarter_start
df['is_quarter_start']= df['is_quarter_start'].astype('category')
df['season']=df[timestamp_id].apply(get_season)
df['season']= df['season'].astype('category')
# + colab={} colab_type="code" id="P6Za9CAdQvZZ"
#mergin test and train data to process them together (adding features ...etc)
testing['test']=True
training['test']=False
data=training.append(testing)
#adding date features
add_date_features(data,'Timestamp')
# + colab={} colab_type="code" id="36wP47rJ2-j6"
# + [markdown] colab_type="text" id="yO69eFwwiX6x"
# # Cleaning eta and distance
# + [markdown] colab_type="text" id="Tkhex0PV2RcB"
# the dataset contained some anomalies that needed to be removed , this code summarizes the steps token to clean it
# + colab={} colab_type="code" id="kzzCCnMHVUvx"
#creating speed feature to clean our dataset
data["speed"]=data.apply(lambda x : (x["Trip_distance"] / x['ETA'] )* (3.6) if x['test']==False else None,axis="columns")
# + colab={} colab_type="code" id="KND3I2i0dqbY"
#dropping anomalies in the train data (which is why i added 'data.test==False' condition)
filter = (data.test==False) & ( (data.ETA<10 ) | (data.speed>120) | ( data.Trip_distance<1 | ( (data.Trip_distance<1 ) & (data.ETA < 60) ) ) )
cleaned = data.drop(data[filter].index)
# + [markdown] colab_type="text" id="42lk9n79uqz2"
# # Clustering and gps processing
# + [markdown] colab_type="text" id="FoP_JYIB3xXJ"
# in order to add more context to our gps coordinates I produced two kind of clustering; trip clustering and pickup , dropoff clustering
# + colab={} colab_type="code" id="3GbxKkMdgVtX"
# + colab={} colab_type="code" id="ZmANE7_y5l4-"
#trip clustering
kmean=KMeans(n_clusters=10,random_state=SEED)
coords = cleaned[['Origin_lat' ,'Origin_lon' ,'Destination_lat' ,'Destination_lon']]
cleaned["cluster_id"]=kmean.fit_predict(coords)
cleaned["cluster_id"]=cleaned["cluster_id"].astype('category')
# + colab={} colab_type="code" id="O5tsI4ISWUtE"
#pickup and dropoff clustering separately
minikmean=MiniBatchKMeans(n_clusters=12,batch_size=1000,random_state=SEED)
cl_pickup = minikmean.fit_predict(cleaned[['Origin_lat' ,'Origin_lon']])
cl_dropoff = minikmean.fit_predict(cleaned[['Destination_lat' ,'Destination_lon']])
cleaned["cluster_Dest"] = cl_dropoff
cleaned["cluster_Orig"] = cl_pickup
cleaned["cluster_Dest"]=cleaned["cluster_Dest"].astype('category')
cleaned["cluster_Orig"]=cleaned["cluster_Orig"].astype('category')
# + [markdown] colab_type="text" id="DPkYdAqnK9kg"
# # PCA for gps coordinates
# + [markdown] colab_type="text" id="TEZBpvNV4k3g"
# here i tried to apply pca to see the impact on the predictive model ,I was inspired by this [kernel](https://www.kaggle.com/gaborfodor/from-eda-to-the-top-lb-0-367)
# + colab={} colab_type="code" id="wi9Abcr7Kj4V"
from sklearn.decomposition import PCA
coords =np.vstack((cleaned[['Origin_lat' ,'Origin_lon']],
cleaned[['Destination_lat' ,'Destination_lon']]))
pca = PCA(random_state=SEED).fit(coords) # define 2 main axis
cleaned['Origin_pca0'] = pca.transform(cleaned[['Origin_lon', 'Origin_lat']])[:,0]
cleaned['Origin_pca1'] = pca.transform(cleaned[['Origin_lon', 'Origin_lat']])[:,1]
cleaned['Destination_pca0'] = pca.transform(cleaned[['Destination_lon', 'Destination_lat']])[:,0]
cleaned['Destination_pca1'] = pca.transform(cleaned[['Destination_lon', 'Destination_lat']])[:,1]
cleaned['distance_pca0'] = np.abs(cleaned.Origin_pca0-cleaned.Destination_pca0)
cleaned['distance_pca1'] = np.abs(cleaned.Origin_pca1-cleaned.Destination_pca1)
# + [markdown] colab_type="text" id="mLVQQKrM8tlu"
# # Adding weather
# + [markdown] colab_type="text" id="WOGjurM0LHos"
# first we need to fill missing values for December 31th 2019 , we will need it later in the test set since it contains trips on that date
# + colab={} colab_type="code" id="ezxtuulxKvKd"
#I opted for the simplest solution by simply imputing missing values by averaging values from the previous 2 months
res=weather_df.mean()
res["date"]=parse("2019-12-31")
# add the new row to our weather dataframe
weather_df=weather_df.append(res,ignore_index=True)
#merging data weather
cleaned['date'] = cleaned.Timestamp.dt.date
cleaned['date'] = pd.to_datetime(cleaned['date'])
cleaned=cleaned.reset_index().merge(weather_df,how="left",on="date").set_index('ID')
# + colab={} colab_type="code" id="_89PcdVF9ObZ"
#create a boolean field is_rainy , == True if the precipitation measure is different than zero
cleaned["is_rainy"]=cleaned['total_precipitation'].apply(lambda x : True if (x>0.0 or x<0.0) else False )
cleaned["is_rainy"]=cleaned["is_rainy"].astype('category')
# + [markdown] colab_type="text" id="v4DUYLtUihg-"
# # Target analysis
# + [markdown] colab_type="text" id="-inFdniRLolr"
# before transformation
# + colab={"base_uri": "https://localhost:8080/", "height": 296} colab_type="code" id="fikxMJyOiuno" outputId="de251bc7-c543-4a4f-fe98-b9e2629b9e07"
target = cleaned[cleaned.test==False]["ETA"]
sns.distplot(target)
print("skewness : ",stats.skew(target.values))
# + [markdown] colab_type="text" id="b3jJM-OL6okF"
# we can see that the target ETA is positively skewed and don't follow a normal distribution , let's try the boxcox transform to fix that
#
# <h2>Box-Cox transformation</h2>
#
# Box-cox transform belongs to the family of power transformation techniques, which are useful to stabilize variance and make the data more gaussian-like distribution. Box-Cox transform are most common since it allows you to try various exponens and choose the best $\lambda$ value instead of picking one manually.
#
# $$
# y_i^{(\lambda)} =
# \begin{cases}
# \dfrac{y_i^\lambda - 1}{\lambda} & \text{if } \lambda \neq 0, \\
# \ln y_i & \text{if } \lambda = 0,
# \end{cases}
# $$
#
# to know more about Transforming Variables techniques , read the following <a href="https://heartbeat.fritz.ai/hands-on-with-feature-engineering-techniques-transforming-variables-acea03472e24" >medium article</a>
#
# + colab={} colab_type="code" id="a3gWyYkMjDdk"
from sklearn.preprocessing import PowerTransformer
boxcox_transformer = PowerTransformer(method='box-cox', standardize=False)
# + colab={"base_uri": "https://localhost:8080/", "height": 282} colab_type="code" id="B40e8aSGijSr" outputId="f9500e6f-5f84-47bf-9b1b-e83a6448c4af"
target = cleaned[cleaned.test==False]["ETA"]
target=boxcox_transformer.fit_transform(target.values.reshape(-1,1))
sns.distplot(target)
print("skewness : ",stats.skew(target))
# + [markdown] colab_type="text" id="-AXCWs7a6537"
# the curve now is way less skewed than the origin
# + [markdown] colab_type="text" id="yzH-i6ugjnDG"
# # Training
# + colab={} colab_type="code" id="bns4_6T5juM9"
# + colab={} colab_type="code" id="MXU_6-wzkovk"
import math
#creating rmse adapted to our transformation
def rmse_boxcox(y_hat, data):
y_true = data.get_label()
y_true = boxcox_transformer.inverse_transform(y_true.reshape(-1,1))
y_hat = boxcox_transformer.inverse_transform(y_hat.reshape(-1,1))
return 'rmse_boxcox', math.sqrt(metrics.mean_squared_error(y_true,y_hat)) , False
# + colab={} colab_type="code" id="gPPpRuTogWlB"
drop_cols=["ETA",'Timestamp',"speed","date",'test']
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="ByrXYiJilKr-" outputId="f307eff5-b0d1-4b82-fba7-b1ad7920a5f1"
#running cross validation on our train set using Lightgbm
split = model_selection.TimeSeriesSplit(n_splits=5)
training = cleaned[cleaned.test==False]
X_train = training.drop(drop_cols,axis=1)
Y_train = target.squeeze()
train_set=lgb.Dataset(X_train,Y_train)
params={"objective":"regression",
'n_estimators':5000}
eval_hist=lgb.cv(params=params,feval=rmse_boxcox,train_set=train_set,folds=split,
early_stopping_rounds=50,seed=SEED)
# + colab={"base_uri": "https://localhost:8080/", "height": 204} colab_type="code" id="LN83ovz6whaP" outputId="c434687d-8937-4dce-ad7f-6aeacb84d929"
pd.DataFrame(eval_hist).tail()
# + [markdown] colab_type="text" id="IZ1Fa1M179KW"
# from the table above , the index represent : num_tress - 1 , we will pick up that value since it performed well on our cv
# in our case , it's 2864
# + colab={"base_uri": "https://localhost:8080/", "height": 119} colab_type="code" id="hlXsdc59p4DP" outputId="bcce19a3-c5a7-4f68-a29e-489ddece2c4b"
#training our final model
model=lgb.LGBMRegressor(n_estimators=2864,random_state=SEED)
model.fit(X_train,Y_train)
# + colab={"base_uri": "https://localhost:8080/", "height": 837} colab_type="code" id="0t_jKb3hJnop" outputId="ea7c61d9-1ab3-49ba-b921-f184e6db2dfc"
lgb.plot_importance(model,figsize=(15,15))
# + [markdown] colab_type="text" id="fJ4SbqJnyqav"
# # submission
#
# + colab={"base_uri": "https://localhost:8080/", "height": 224} colab_type="code" id="9Gq6ZBFPq2wY" outputId="5d145bcd-22a3-4752-db54-7c308eda81d7"
testing = cleaned[cleaned.test==True]
testing.drop(drop_cols,axis=1,inplace=True)
testing["ETA"]=boxcox_transformer.inverse_transform(model.predict(testing).reshape(-1,1))
testing[['ETA']].to_csv('submit.csv')
# + [markdown] colab_type="text" id="8yXz7Ppfy3-0"
# # saving models (optional)
# + colab={} colab_type="code" id="Br6c8vYAtMeG"
import pickle
pkl_filename = "./drive/My Drive/kmean.pkl"
with open(pkl_filename, 'wb') as file:
pickle.dump(kmean, file)
# + colab={} colab_type="code" id="rwKX-Ic0tig_"
pkl_filename = "./drive/My Drive/Minibatchkmean.pkl"
with open(pkl_filename, 'wb') as file:
pickle.dump(minikmean, file)
# + colab={} colab_type="code" id="DH-JOD3ZuQ-C"
pkl_filename = "./drive/My Drive/pca.pkl"
with open(pkl_filename, 'wb') as file:
pickle.dump(pca, file)
# + colab={} colab_type="code" id="0hnYgyqeuWWy"
pkl_filename = "./drive/My Drive/lgbRegressor.pkl"
with open(pkl_filename, 'wb') as file:
pickle.dump(model, file)
|
Yassir_ETA_prediction_solution.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <img src='./IMG/head_a.png' alt='Logo Head' align='center' width='100%'></img>
# <br>
# <img src='./IMG/header_1.png' alt='Logo UNSPIDER' align='left' width='50%'></img>
# <br>
# <a href="./03_USO_DEL_PAQUETE_GEEMAP.ipynb">**<< 03 - USO DEL PAQUETE GEEMAP**</a><span style="float:right;"><a href="./05_MAPEO_INTERACTIVO_USANDO_API_PYTHON_DE_GEE_GEEMAP.ipynb">**05 - MAPA INTERACTIVO USANDO API Python de geemap >>**</a>
# <hr>
# # 4. CONVERSIÓN AUTOMÁTICA DE GEE - Javascripts a Python y Jupyter
# Los siguientes ejemplos requieren el paquete **geemap**, que puede instalarse utilizando `pip install geemap`. Consulte la sección de <a href="./02_INSTALACIÓN_DEL PAQUETE_GEEMAP.ipynb"> Instalación para más información</a>.
#
# Inicie un cuaderno interactivo con **Google Colab**. Tenga en cuenta que la conversión no siempre funciona a la perfección. Es posible que aún se necesiten cambios manuales adicionales. ui y chart no son compatibles. El código fuente de este módulo de conversión automatizado se puede encontrar en github del Profesor Wu, Q en [conversion.py.](https://github.com/giswqs/geemap/blob/master/geemap/conversion.py).
# <a href="https://colab.research.google.com/github/Alexanderariza/FOREST-FIRES-ON-GEE-GEEMAP/blob/master/05_MAPEO_INTERACTIVO_USANDO_API_PYTHON_DE_GEE_GEEMAP.ipynb"><img src='./IMG/COLAB.svg' alt='Logo CO' align='left' width='10%'></img>
# <br>
#
# <a href="https://nbviewer.jupyter.org/github/Alexanderariza/FOREST-FIRES-ON-GEE-GEEMAP/blob/master/05_MAPEO_INTERACTIVO_USANDO_API_PYTHON_DE_GEE_GEEMAP.ipynb"><img src='./IMG/NT_vie.svg' alt='NT_VW' align='left' width='10%'></img>
# <br>
# * Instalar Earth Engine API y geemap:
# +
import subprocess
try:
import geemap
except ImportError:
print('geemap package not installed. Installing ...')
subprocess.check_call(["python", '-m', 'pip', 'install', 'geemap'])
# -
# * Convierta los JavaScripts de Earth Engine en scripts de Python:
# +
import os
from geemap.conversion import *
# Create a temporary working directory
work_dir = os.path.join(os.path.expanduser('~'), 'geemap')
# Get Earth Engine JavaScript examples. There are five examples in the geemap package folder.
# Change js_dir to your own folder containing your Earth Engine JavaScripts,
# such as js_dir = '/path/to/your/js/folder'
js_dir = get_js_examples(out_dir=work_dir)
# Convert all Earth Engine JavaScripts in a folder recursively to Python scripts.
js_to_python_dir(in_dir=js_dir, out_dir=js_dir, use_qgis=True)
print("Python scripts saved at: {}".format(js_dir))
# -
# * Convierta scripts de Python de Earth Engine en cuadernos Jupyter:
# +
# Convert all Earth Engine Python scripts in a folder recursively to Jupyter notebooks.
nb_template = get_nb_template() # Get the notebook template from the package folder.
py_to_ipynb_dir(js_dir, nb_template)
# Execute all Jupyter notebooks in a folder recursively and save the output cells.
execute_notebook_dir(in_dir=js_dir)
# -
# <a href="./03_USO_DEL_PAQUETE_GEEMAP.ipynb">**<< 03 - USO DEL PAQUETE GEEMAP**</a><span style="float:right;"><a href="./05_MAPEO_INTERACTIVO_USANDO_API_PYTHON_DE_GEE_GEEMAP.ipynb">**05 - MAPA INTERACTIVO USANDO API Python de geemap >>**</a>
# <hr>
# <img src='./IMG/UNicon.png' alt='Logo down' align='right' width='8%'></img>
# <i><p style="text-align:right;">Material organizado por el [**<NAME>**](https://www.researchgate.net/profile/Alexander_Ariza2), como apoyo al portal de conocimiento de la oficina de [**UNSPIDER**](https://www.un-spider.org), (2020) .
|
04_GEE_JavaScripts_2_Python_NB.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: MindSpore-1.0.1
# language: python
# name: mindspore-1.0.1
# ---
# # 深度概率编程CVAE
#
# ## 概述
#
# 本例采用MindSpore的深度概率编程方法应用于条件变分自编码器(CVAE)模型训练。
# 整体流程如下:
#
# 1. 数据集准备
# 2. 定义条件变分自编码器网络;
# 3. 定义损失函数和优化器;
# 4. 训练生成模型。
# 5. 生成新样本或重构输入样本。
#
# > 本例适用于GPU和Ascend环境。
# ## 数据准备
#
# ### 下载数据集
#
# 本例使用MNIST_Data数据集,在Jupyter Notebook中执行如下命令进行下载并解压到对应位置:
# !mkdir -p ./datasets/MNIST_Data/train ./datasets/MNIST_Data/test
# !wget -NP ./datasets/MNIST_Data/train https://mindspore-website.obs.myhuaweicloud.com/notebook/datasets/mnist/train-labels-idx1-ubyte --no-check-certificate
# !wget -NP ./datasets/MNIST_Data/train https://mindspore-website.obs.myhuaweicloud.com/notebook/datasets/mnist/train-images-idx3-ubyte --no-check-certificate
# !wget -NP ./datasets/MNIST_Data/test https://mindspore-website.obs.myhuaweicloud.com/notebook/datasets/mnist/t10k-labels-idx1-ubyte --no-check-certificate
# !wget -NP ./datasets/MNIST_Data/test https://mindspore-website.obs.myhuaweicloud.com/notebook/datasets/mnist/t10k-images-idx3-ubyte --no-check-certificate
# !tree ./datasets/MNIST_Data
# ### 数据增强
#
# 将数据集增强为适应CVAE网络训练要求的数据,本例主要是将原始图片像素大小由$28\times28$增强为$32\times32$,同时将多张图片组成1个`batch`来加速训练。
# +
import mindspore.common.dtype as mstype
import mindspore.dataset as ds
import mindspore.dataset.vision.c_transforms as CV
def create_dataset(data_path, batch_size=32, repeat_size=1,
num_parallel_workers=1):
"""
create dataset for train or test
"""
# define dataset
mnist_ds = ds.MnistDataset(data_path)
resize_height, resize_width = 32, 32
rescale = 1.0 / 255.0
shift = 0.0
# define map operations
resize_op = CV.Resize((resize_height, resize_width)) # Bilinear mode
rescale_op = CV.Rescale(rescale, shift)
hwc2chw_op = CV.HWC2CHW()
# apply map operations on images
mnist_ds = mnist_ds.map(operations=resize_op, input_columns="image", num_parallel_workers=num_parallel_workers)
mnist_ds = mnist_ds.map(operations=rescale_op, input_columns="image", num_parallel_workers=num_parallel_workers)
mnist_ds = mnist_ds.map(operations=hwc2chw_op, input_columns="image", num_parallel_workers=num_parallel_workers)
# apply DatasetOps
mnist_ds = mnist_ds.batch(batch_size)
mnist_ds = mnist_ds.repeat(repeat_size)
return mnist_ds
# -
# ## 定义条件变分自编码器网络
#
# 变分自编码器的构成主要分为三个部分,编码器,解码器和隐空间。
# 其中:
# 编码器(Encoder)主要作用是将训练数据进行降维,压缩,提取特征,形成特征向量,存储在隐空间中。
# 解码器(Decoder)主要作用是将训练数据因空间分布的参数进行解码,还原生成出新的图像。
# 隐空间主要作用是将模型的特征按照某种分布特性进行存储,属于编码器和解码器中间的桥梁。
# 本例中条件变分自编码器(CVAE)是在变分自编码器的基础上增添标签训练,在后续随机采样生成图片的过程中,可以施加标签指定生成该条件的图片。
# +
import os
import mindspore.nn as nn
from mindspore import context, Tensor
import mindspore.ops as ops
context.set_context(mode=context.GRAPH_MODE,device_target="GPU")
IMAGE_SHAPE=(-1,1,32,32)
image_path = os.path.join("./datasets/MNIST_Data","train")
class Encoder(nn.Cell):
def __init__(self, num_classes):
super(Encoder, self).__init__()
self.fc1 = nn.Dense(1024 + num_classes, 400)
self.relu = nn.ReLU()
self.flatten = nn.Flatten()
self.concat = ops.Concat(axis=1)
self.one_hot = nn.OneHot(depth=num_classes)
def construct(self, x, y):
x = self.flatten(x)
y = self.one_hot(y)
input_x = self.concat((x, y))
input_x = self.fc1(input_x)
input_x = self.relu(input_x)
return input_x
class Decoder(nn.Cell):
def __init__(self):
super(Decoder, self).__init__()
self.fc2 = nn.Dense(400, 1024)
self.sigmoid = nn.Sigmoid()
self.reshape = ops.Reshape()
def construct(self, z):
z = self.fc2(z)
z = self.reshape(z, IMAGE_SHAPE)
z = self.sigmoid(z)
return z
# -
# ## 定义优化器和损失函数
#
# 定义条件变分自编码器的损失函数,将图像与label关联。
# 损失函数采用ELBO函数,此函数用于计算解码图像和原图像的差值,并通过对比两个图像的差值,以及图像分布的均值之差来计算两图的损失情况。
# 优化器采用`nn.Adam`来最小化损失值。
# +
from mindspore.nn.probability.dpn import ConditionalVAE
from mindspore.nn.probability.infer import ELBO, SVI
class CVAEWithLossCell(nn.WithLossCell):
"""
Rewrite WithLossCell for CVAE
"""
def construct(self, data, label):
out = self._backbone(data, label)
return self._loss_fn(out, label)
# define the encoder and decoder
encoder = Encoder(num_classes=10)
decoder = Decoder()
# define the vae model
cvae = ConditionalVAE(encoder, decoder, hidden_size=400, latent_size=20,num_classes=10)
# define the loss function
net_loss = ELBO(latent_prior='Normal', output_prior='Normal')
# define the optimizer
optimizer = nn.Adam(params=cvae.trainable_params(), learning_rate=0.001)
net_with_loss = CVAEWithLossCell(cvae,net_loss)
vi = SVI(net_with_loss=net_with_loss,optimizer=optimizer)
# -
# 参数解释:
#
# - num_classes:类别数量,本例中为0-9个数字,共计10个种类。
# - ConditionalVAE:条件自编码器模型,将编码器,解码器,压缩大小,隐空间维度和类别数量等变分自编码器网络初始化。
# - `encoder`:编码器网络。
# - `decoder`:解码器网络。
# - `hiddend_size`:数据压缩后的大小,本例为400。
# - `latent_size`:隐空间的向量维度,向量维度越大,分别的特征维度越多,图像特征越清晰,本例中可调节维度大小为20。
# - `num_classes`:类别数量。
# - ELBO:变分自编码器的损失函数。
# - `latent_prior`:隐空间初始化分布,本例中隐空间的参数遵循正态分布。
# - `output_prior`:输出权重的初始化分布,本例中其权重参数初始化分布遵循正态分布。
# - nn.Adam:优化器。
# - CVAEWithLossCell:本例重建了`nn.WithlossCell`函数,使得生成的数据,附带标签(label)。
# - SVI:模型函数,类似MindSpore中的Model,此函数为变分自编码器专用模型函数。
# ## 训练生成模型
#
# 生成训练数据,将调用上述代码中`vi`的训练模式,对模型进行训练,训练完成后打印出模型的loss值。
# define the training dataset
ds_train = create_dataset(image_path, 32, 1)
# run the vi to return the trained network.
cvae = vi.run(train_dataset=ds_train, epochs=5)
# get the trained loss
trained_loss = vi.get_train_loss()
print(trained_loss)
# ### 样本重建
#
# 先定义可视化绘图函数`plot_image`,用于样本重建和条件采样生成数据的可视化。
#
# 使用训练好的模型,查看重建数据的能力如何,这里取一组原始数据进行重建,执行如下代码:
# +
import matplotlib.pyplot as plt
import numpy as np
def plot_image(sample_data,col_num=4,row_num=8,count=0):
for i in sample_data:
plt.subplot(col_num,row_num,count+1)
plt.imshow(np.squeeze(i.asnumpy()))
plt.axis("off")
count += 1
plt.show()
sample = next(ds_train.create_dict_iterator(output_numpy=True, num_epochs=1))
sample_x = Tensor(sample['image'], dtype=mstype.float32)
sample_y = Tensor(sample['label'], dtype=mstype.int32)
reconstructed_sample = cvae.reconstruct_sample(sample_x, sample_y)
print('The shape of the reconstructed sample is ', reconstructed_sample.shape)
print("\n=============The Original Images=============")
plot_image(sample_x)
print("\n============The Reconstruct Images=============")
plot_image(reconstructed_sample)
# -
# 对比原图片,CVAE生成的图片能明显对应上原始图片,但还稍显模糊。说明训练效果已经达到但还有提升空间。
# ### 条件样本采样
#
# 在隐空间中进行条件采样,本例使用条件为`(0,1)`,对应生成`(0,1)`的图像数据,同时将采样生成的数据进行可视化。
# test function: generate_sample
sample_label = Tensor([i for i in range(0,2)]*16, dtype=mstype.int32)
# test function: generate_sample
generated_sample = cvae.generate_sample(sample_label, 32, IMAGE_SHAPE)
# test function: reconstruct_sample
print('The shape of the generated sample is ', generated_sample.shape)
plot_image(generated_sample,4,8)
# 在条件为`(0,1)`特征采样中,生成的图片有的看起来像其他的数字,说明图像在特征分布中,其他数字的部分特征与`(0,1)`的特征出现了交叉,而随机采样正好采样到了这些交叉特征,导致`(0,1)`图片出现了其他数字的特征。
|
docs/notebook/apply_deep_probability_programming/apply_deep_probability_programming_cvae.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Data Preprocessing
# We split the ground truth masks into 4 binary masks :
# * `gt1` : Left Ventricule / Endocardium.
# * `gt2` : Myocardium.
# * `gt3` : Left Atrium.
# * `gt4` : Epicardium (Union of Myocarium with Endocardium).
# +
import SimpleITK as sitk
import os
import numpy as np
# Pixel spacing in mm as set by the original dataset
spacing = [0.308,0.154]
# Path to parent directory of dataset (Either test or train/val)
PATH="/home/mourad/TDSI/challenge/testmourad/"
for r,d,f in os.walk(PATH):
for file in f:
if ('ED_gt.mhd' in file) or ('ES_gt.mhd' in file):
image = sitk.GetArrayFromImage(sitk.ReadImage(os.path.join(r,file)))
# We eliminate the first axis to get a 2D image
image = image.squeeze()
lv = np.where (image == 1, 1, 0)
myo = np.where (image == 2,1 ,0)
la = np.where (image == 3, 1 , 0)
epi = myo + lv
# Encoding in Char as the original format
lv = lv.astype('uint8')
epi = epi.astype('uint8')
la = la.astype('uint8')
myo = myo.astype('uint8')
lvImage = sitk.GetImageFromArray(lv,isVector=False)
lvImage.SetSpacing(spacing)
sitk.WriteImage(lvImage,os.path.join(r,file)[:-4]+"1.mhd",False)
epiImage = sitk.GetImageFromArray(epi,isVector=False)
epiImage.SetSpacing(spacing)
sitk.WriteImage(epiImage,os.path.join(r,file)[:-4]+"4.mhd",False)
laImage = sitk.GetImageFromArray(la,isVector=False)
laImage.SetSpacing(spacing)
sitk.WriteImage(laImage,os.path.join(r,file)[:-4]+"3.mhd",False)
myoImage = sitk.GetImageFromArray(myo,isVector=False)
myoImage.SetSpacing(spacing)
sitk.WriteImage(myoImage,os.path.join(r,file)[:-4]+"2.mhd",False)
# +
import SimpleITK as sitk
import os
import numpy as np
PATH="/home/mourad/TDSI/challenge/data/camus_separated"
for r,d,f in os.walk(PATH):
for file in f:
for i in range (3)
if ('ED_gt' + str(i) + '.mhd' in file) or ('ES_gt' + str (i) + '.mhd' in file):
image = sitk.GetArrayFromImage(sitk.ReadImage(os.path.join(r,file)))
plt.imshow(image)
|
samples/CAMUS/dataset_preprocess.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # imports
# %matplotlib inline
import matplotlib.pyplot as plt
from scipy.ndimage import correlate
import numpy as np
from skimage import data
from skimage.color import rgb2gray
from skimage.transform import rescale,resize
# # original image input
# +
im = rgb2gray(data.coffee())
im = resize(im, (64,64))
print(im.shape)
plt.axis('off')
plt.imshow(im, cmap = 'gray');
# -
# # horizontal edge filter
# +
filter1 = np.array([
[ 1, 1, 1],
[ 0, 0, 0],
[-1, -1, -1]
])
new_image = np.zeros(im.shape)
im_pad = np.pad(im, 1, 'constant')
for i in range(im.shape[0]):
for j in range(im.shape[1]):
try:
new_image[i,j] = \
im_pad[i-1,j-1] * filter1[0,0] + \
im_pad[i-1,j] * filter1[0,1] + \
im_pad[i-1,j+1] * filter1[0,2] + \
im_pad[i,j-1] * filter1[1,0] + \
im_pad[i,j] * filter1[1,1] + \
im_pad[i,j+1] * filter1[1,2] +\
im_pad[i+1,j-1] * filter1[2,0] + \
im_pad[i+1,j] * filter1[2,1] + \
im_pad[i+1,j+1] * filter1[2,2]
except:
pass
plt.axis('off')
plt.imshow(new_image, cmap='Greys');
# -
# # vertical edge filter
# +
filter2 = np.array([
[ -1, 0, 1],
[ -1, 0, 1],
[ -1, 0, 1]
])
new_image = np.zeros(im.shape)
im_pad = np.pad(im,1, 'constant')
for i in range(im.shape[0]):
for j in range(im.shape[1]):
try:
new_image[i,j] = \
im_pad[i-1,j-1] * filter2[0,0] + \
im_pad[i-1,j] * filter2[0,1] + \
im_pad[i-1,j+1] * filter2[0,2] + \
im_pad[i,j-1] * filter2[1,0] + \
im_pad[i,j] * filter2[1,1] + \
im_pad[i,j+1] * filter2[1,2] +\
im_pad[i+1,j-1] * filter2[2,0] + \
im_pad[i+1,j] * filter2[2,1] + \
im_pad[i+1,j+1] * filter2[2,2]
except:
pass
plt.axis('off')
plt.imshow(new_image, cmap='Greys');
# -
# # horizontal edge filter with stride 2
# +
filter1 = np.array([
[ 1, 1, 1],
[ 0, 0, 0],
[-1, -1, -1]
])
stride = 2
new_image = np.zeros((int(im.shape[0] / stride), int(im.shape[1] / stride)))
im_pad = np.pad(im,1, 'constant')
for i in range(0,im.shape[0],stride):
for j in range(0,im.shape[1],stride):
try:
new_image[int(i/stride),int(j/stride)] = \
im_pad[i-1,j-1] * filter1[0,0] + \
im_pad[i-1,j] * filter1[0,1] + \
im_pad[i-1,j+1] * filter1[0,2] + \
im_pad[i,j-1] * filter1[1,0] + \
im_pad[i,j] * filter1[1,1] + \
im_pad[i,j+1] * filter1[1,2] +\
im_pad[i+1,j-1] * filter1[2,0] + \
im_pad[i+1,j] * filter1[2,1] + \
im_pad[i+1,j+1] * filter1[2,2]
except:
pass
plt.axis('off')
plt.imshow(new_image, cmap='Greys');
# -
# # vertical edge filter with stride 2
# +
filter2 = np.array([
[ -1, 0, 1],
[ -1, 0, 1],
[ -1, 0, 1]
])
stride = 2
new_image = np.zeros((int(im.shape[0] / stride), int(im.shape[1] / stride)))
im_pad = np.pad(im,1, 'constant')
for i in range(0,im.shape[0],stride):
for j in range(0,im.shape[1],stride):
try:
new_image[int(i/stride),int(j/stride)] = \
im_pad[i-1,j-1] * filter2[0,0] + \
im_pad[i-1,j] * filter2[0,1] + \
im_pad[i-1,j+1] * filter2[0,2] + \
im_pad[i,j-1] * filter2[1,0] + \
im_pad[i,j] * filter2[1,1] + \
im_pad[i,j+1] * filter2[1,2] +\
im_pad[i+1,j-1] * filter2[2,0] + \
im_pad[i+1,j] * filter2[2,1] + \
im_pad[i+1,j+1] * filter2[2,2]
except:
pass
plt.axis('off')
plt.imshow(new_image, cmap='Greys');
# -
|
GAN/02_02_deep_learning_convolutions.ipynb
|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Project 3: Implement SLAM
#
# ---
#
# ## Project Overview
#
# In this project, you'll implement SLAM for robot that moves and senses in a 2 dimensional, grid world!
#
# SLAM gives us a way to both localize a robot and build up a map of its environment as a robot moves and senses in real-time. This is an active area of research in the fields of robotics and autonomous systems. Since this localization and map-building relies on the visual sensing of landmarks, this is a computer vision problem.
#
# Using what you've learned about robot motion, representations of uncertainty in motion and sensing, and localization techniques, you will be tasked with defining a function, `slam`, which takes in six parameters as input and returns the vector `mu`.
# > `mu` contains the (x,y) coordinate locations of the robot as it moves, and the positions of landmarks that it senses in the world
#
# You can implement helper functions as you see fit, but your function must return `mu`. The vector, `mu`, should have (x, y) coordinates interlaced, for example, if there were 2 poses and 2 landmarks, `mu` will look like the following, where `P` is the robot position and `L` the landmark position:
# ```
# mu = matrix([[Px0],
# [Py0],
# [Px1],
# [Py1],
# [Lx0],
# [Ly0],
# [Lx1],
# [Ly1]])
# ```
#
# You can see that `mu` holds the poses first `(x0, y0), (x1, y1), ...,` then the landmark locations at the end of the matrix; we consider a `nx1` matrix to be a vector.
#
# ## Generating an environment
#
# In a real SLAM problem, you may be given a map that contains information about landmark locations, and in this example, we will make our own data using the `make_data` function, which generates a world grid with landmarks in it and then generates data by placing a robot in that world and moving and sensing over some numer of time steps. The `make_data` function relies on a correct implementation of robot move/sense functions, which, at this point, should be complete and in the `robot_class.py` file. The data is collected as an instantiated robot moves and senses in a world. Your SLAM function will take in this data as input. So, let's first create this data and explore how it represents the movement and sensor measurements that our robot takes.
#
# ---
# ## Create the world
#
# Use the code below to generate a world of a specified size with randomly generated landmark locations. You can change these parameters and see how your implementation of SLAM responds!
#
# `data` holds the sensors measurements and motion of your robot over time. It stores the measurements as `data[i][0]` and the motion as `data[i][1]`.
#
# #### Helper functions
#
# You will be working with the `robot` class that may look familiar from the first notebook,
#
# In fact, in the `helpers.py` file, you can read the details of how data is made with the `make_data` function. It should look very similar to the robot move/sense cycle you've seen in the first notebook.
# +
import numpy as np
from helpers import make_data
# your implementation of slam should work with the following inputs
# feel free to change these input values and see how it responds!
# world parameters
num_landmarks = 5 # number of landmarks
N = 20 # time steps
world_size = 100.0 # size of world (square)
# robot parameters
measurement_range = 50.0 # range at which we can sense landmarks
motion_noise = 2.0 # noise in robot motion
measurement_noise = 2.0 # noise in the measurements
distance = 20.0 # distance by which robot (intends to) move each iteratation
# make_data instantiates a robot, AND generates random landmarks for a given world size and number of landmarks
data = make_data(N, num_landmarks, world_size, measurement_range, motion_noise, measurement_noise, distance)
# -
# ### A note on `make_data`
#
# The function above, `make_data`, takes in so many world and robot motion/sensor parameters because it is responsible for:
# 1. Instantiating a robot (using the robot class)
# 2. Creating a grid world with landmarks in it
#
# **This function also prints out the true location of landmarks and the *final* robot location, which you should refer back to when you test your implementation of SLAM.**
#
# The `data` this returns is an array that holds information about **robot sensor measurements** and **robot motion** `(dx, dy)` that is collected over a number of time steps, `N`. You will have to use *only* these readings about motion and measurements to track a robot over time and find the determine the location of the landmarks using SLAM. We only print out the true landmark locations for comparison, later.
#
#
# In `data` the measurement and motion data can be accessed from the first and second index in the columns of the data array. See the following code for an example, where `i` is the time step:
# ```
# measurement = data[i][0]
# motion = data[i][1]
# ```
#
# +
# print out some stats about the data
time_step = 0
print('Example measurements: \n', data[time_step][0])
print('\n')
print('Example motion: \n', data[time_step][1])
# -
# Try changing the value of `time_step`, you should see that the list of measurements varies based on what in the world the robot sees after it moves. As you know from the first notebook, the robot can only sense so far and with a certain amount of accuracy in the measure of distance between its location and the location of landmarks. The motion of the robot always is a vector with two values: one for x and one for y displacement. This structure will be useful to keep in mind as you traverse this data in your implementation of slam.
# ## Initialize Constraints
#
# One of the most challenging tasks here will be to create and modify the constraint matrix and vector: omega and xi. In the second notebook, you saw an example of how omega and xi could hold all the values the define the relationships between robot poses `xi` and landmark positions `Li` in a 1D world, as seen below, where omega is the blue matrix and xi is the pink vector.
#
# <img src='images/motion_constraint.png' width=50% height=50% />
#
#
# In *this* project, you are tasked with implementing constraints for a 2D world. We are referring to robot poses as `Px, Py` and landmark positions as `Lx, Ly`, and one way to approach this challenge is to add *both* x and y locations in the constraint matrices.
#
# <img src='images/constraints2D.png' width=50% height=50% />
#
# You may also choose to create two of each omega and xi (one for x and one for y positions).
# ### TODO: Write a function that initializes omega and xi
#
# Complete the function `initialize_constraints` so that it returns `omega` and `xi` constraints for the starting position of the robot. Any values that we do not yet know should be initialized with the value `0`. You may assume that our robot starts out in exactly the middle of the world with 100% confidence (no motion or measurement noise at this point). The inputs `N` time steps, `num_landmarks`, and `world_size` should give you all the information you need to construct intial constraints of the correct size and starting values.
#
# *Depending on your approach you may choose to return one omega and one xi that hold all (x,y) positions *or* two of each (one for x values and one for y); choose whichever makes most sense to you!*
def initialize_constraints(N, num_landmarks, world_size):
''' This function takes in a number of time steps N, number of landmarks, and a world_size,
and returns initialized constraint matrices, omega and xi.'''
## Recommended: Define and store the size (rows/cols) of the constraint matrix in a variable
rows = (2 * N) + (2 * num_landmarks)
cols = (2 * N) + (2 * num_landmarks)
## TODO: Define the constraint matrix, Omega, with two initial "strength" values
## for the initial x, y location of our robot
omega = np.zeros(shape=(rows, cols))
Px_init = 0
Py_init = 1
for coordinate in [Px_init, Py_init]:
omega[coordinate][coordinate] = 1
## TODO: Define the constraint *vector*, xi
## you can assume that the robot starts out in the middle of the world with 100% confidence
xi = np.zeros(shape=(rows, 1))
for coordinate in [Px_init, Py_init]:
xi[coordinate] = world_size / 2
# return output
return omega, xi
# ### Test as you go
#
# It's good practice to test out your code, as you go. Since `slam` relies on creating and updating constraint matrices, `omega` and `xi` to account for robot sensor measurements and motion, let's check that they initialize as expected for any given parameters.
#
# Below, you'll find some test code that allows you to visualize the results of your function `initialize_constraints`. We are using the [seaborn](https://seaborn.pydata.org/) library for visualization.
#
# **Please change the test values of N, landmarks, and world_size and see the results**. Be careful not to use these values as input into your final smal function.
#
# This code assumes that you have created one of each constraint: `omega` and `xi`, but you can change and add to this code, accordingly. The constraints should vary in size with the number of time steps and landmarks as these values affect the number of poses a robot will take `(Px0,Py0,...Pxn,Pyn)` and landmark locations `(Lx0,Ly0,...Lxn,Lyn)` whose relationships should be tracked in the constraint matrices. Recall that `omega` holds the weights of each variable and `xi` holds the value of the sum of these variables, as seen in Notebook 2. You'll need the `world_size` to determine the starting pose of the robot in the world and fill in the initial values for `xi`.
# import data viz resources
import matplotlib.pyplot as plt
from pandas import DataFrame
import seaborn as sns
# %matplotlib inline
# +
# define a small N and world_size (small for ease of visualization)
N_test = 5
num_landmarks_test = 2
small_world = 10
# initialize the constraints
initial_omega, initial_xi = initialize_constraints(N_test, num_landmarks_test, small_world)
# +
# define figure size
plt.rcParams["figure.figsize"] = (10,7)
# display omega
sns.heatmap(DataFrame(initial_omega), cmap='Blues', annot=True, linewidths=.5);
# +
# define figure size
plt.rcParams["figure.figsize"] = (1,7)
# display xi
sns.heatmap(DataFrame(initial_xi), cmap='Oranges', annot=True, linewidths=.5);
# -
# ---
# ## SLAM inputs
#
# In addition to `data`, your slam function will also take in:
# * N - The number of time steps that a robot will be moving and sensing
# * num_landmarks - The number of landmarks in the world
# * world_size - The size (w/h) of your world
# * motion_noise - The noise associated with motion; the update confidence for motion should be `1.0/motion_noise`
# * measurement_noise - The noise associated with measurement/sensing; the update weight for measurement should be `1.0/measurement_noise`
#
# #### A note on noise
#
# Recall that `omega` holds the relative "strengths" or weights for each position variable, and you can update these weights by accessing the correct index in omega `omega[row][col]` and *adding/subtracting* `1.0/noise` where `noise` is measurement or motion noise. `Xi` holds actual position values, and so to update `xi` you'll do a similar addition process only using the actual value of a motion or measurement. So for a vector index `xi[row][0]` you will end up adding/subtracting one measurement or motion divided by their respective `noise`.
#
# ### TODO: Implement Graph SLAM
#
# Follow the TODO's below to help you complete this slam implementation (these TODO's are in the recommended order), then test out your implementation!
#
# #### Updating with motion and measurements
#
# With a 2D omega and xi structure as shown above (in earlier cells), you'll have to be mindful about how you update the values in these constraint matrices to account for motion and measurement constraints in the x and y directions. Recall that the solution to these matrices (which holds all values for robot poses `P` and landmark locations `L`) is the vector, `mu`, which can be computed at the end of the construction of omega and xi as the inverse of omega times xi: $\mu = \Omega^{-1}\xi$
#
# **You may also choose to return the values of `omega` and `xi` if you want to visualize their final state!**
# +
## TODO: Complete the code to implement SLAM
## slam takes in 6 arguments and returns mu,
## mu is the entire path traversed by a robot (all x,y poses) *and* all landmarks locations
def slam(data, N, num_landmarks, world_size, motion_noise, measurement_noise):
## TODO: Use your initilization to create constraint matrices, omega and xi
omega, xi = initialize_constraints(N, num_landmarks, world_size)
## TODO: Iterate through each time step in the data
## get all the motion and measurement data as you iterate
for timestep in range(N-1):
measurement = data[timestep][0]
motion = data[timestep][1]
dx = motion[0]
dy = motion[1]
## TODO: update the constraint matrix/vector to account for all *measurements*
## this should be a series of additions that take into account the measurement noise
x0 = (timestep * 2)
x1 = x0 + 2
y0 = x0 + 1
y1 = y0 + 2
mea_n = measurement_noise
mot_n = motion_noise
for landmark in measurement:
dx_lm = landmark[1]
dy_lm = landmark[2]
x0_lm = (N * 2) + (landmark[0] * 2)
y0_lm = x0_lm + 1
# update omega values for measurement noise
omega[x0][x0] += 1 / mea_n
omega[x0_lm][x0_lm] += 1 / mea_n
omega[x0][x0_lm] -= 1 / mea_n
omega[x0_lm][x0] -= 1 / mea_n
omega[y0][y0] += 1.0 / mea_n
omega[y0_lm][y0_lm] += 1.0 / mea_n
omega[y0][y0_lm] -= 1.0 / mea_n
omega[y0_lm][y0] -= 1.0 / mea_n
# update xi values for measurement noise
xi[x0] -= dx_lm / mea_n
xi[x0_lm] += dx_lm / mea_n
xi[y0] -= dy_lm / mea_n
xi[y0_lm] += dy_lm / mea_n
## TODO: update the constraint matrix/vector to account for all *motion* and motion noise
# update omega values for motion noise
omega[x0][x0] += 1 / mot_n
omega[x1][x1] += 1 / mot_n
omega[x0][x1] -= 1 / mot_n
omega[x1][x0] -= 1 / mot_n
omega[y0][y0] += 1 / mot_n
omega[y1][y1] += 1 / mot_n
omega[y0][y1] -= 1 / mot_n
omega[y1][y0] -= 1 / mot_n
# update xi values for motion noise
xi[x0] -= dx / mot_n
xi[y0] -= dy / mot_n
xi[x1] += dx / mot_n
xi[y1] += dy / mot_n
## TODO: After iterating through all the data
## Compute the best estimate of poses and landmark positions
## using the formula, omega_inverse * Xi
omega_inv = np.linalg.inv(np.matrix(omega))
mu = omega_inv * xi
# return `mu`
return mu
# -
# ## Helper functions
#
# To check that your implementation of SLAM works for various inputs, we have provided two helper functions that will help display the estimated pose and landmark locations that your function has produced. First, given a result `mu` and number of time steps, `N`, we define a function that extracts the poses and landmarks locations and returns those as their own, separate lists.
#
# Then, we define a function that nicely print out these lists; both of these we will call, in the next step.
#
# a helper function that creates a list of poses and of landmarks for ease of printing
# this only works for the suggested constraint architecture of interlaced x,y poses
def get_poses_landmarks(mu, N):
# create a list of poses
poses = []
for i in range(N):
poses.append((mu[2*i].item(), mu[2*i+1].item()))
# create a list of landmarks
landmarks = []
for i in range(num_landmarks):
landmarks.append((mu[2*(N+i)].item(), mu[2*(N+i)+1].item()))
# return completed lists
return poses, landmarks
def print_all(poses, landmarks):
print('\n')
print('Estimated Poses:')
for i in range(len(poses)):
print('['+', '.join('%.3f'%p for p in poses[i])+']')
print('\n')
print('Estimated Landmarks:')
for i in range(len(landmarks)):
print('['+', '.join('%.3f'%l for l in landmarks[i])+']')
# ## Run SLAM
#
# Once you've completed your implementation of `slam`, see what `mu` it returns for different world sizes and different landmarks!
#
# ### What to Expect
#
# The `data` that is generated is random, but you did specify the number, `N`, or time steps that the robot was expected to move and the `num_landmarks` in the world (which your implementation of `slam` should see and estimate a position for. Your robot should also start with an estimated pose in the very center of your square world, whose size is defined by `world_size`.
#
# With these values in mind, you should expect to see a result that displays two lists:
# 1. **Estimated poses**, a list of (x, y) pairs that is exactly `N` in length since this is how many motions your robot has taken. The very first pose should be the center of your world, i.e. `[50.000, 50.000]` for a world that is 100.0 in square size.
# 2. **Estimated landmarks**, a list of landmark positions (x, y) that is exactly `num_landmarks` in length.
#
# #### Landmark Locations
#
# If you refer back to the printout of *exact* landmark locations when this data was created, you should see values that are very similar to those coordinates, but not quite (since `slam` must account for noise in motion and measurement).
# +
# call your implementation of slam, passing in the necessary parameters
mu = slam(data, N, num_landmarks, world_size, motion_noise, measurement_noise)
# print out the resulting landmarks and poses
if(mu is not None):
# get the lists of poses and landmarks
# and print them out
poses, landmarks = get_poses_landmarks(mu, N)
print_all(poses, landmarks)
# -
# ## Visualize the constructed world
#
# Finally, using the `display_world` code from the `helpers.py` file (which was also used in the first notebook), we can actually visualize what you have coded with `slam`: the final position of the robot and the positon of landmarks, created from only motion and measurement data!
#
# **Note that these should be very similar to the printed *true* landmark locations and final pose from our call to `make_data` early in this notebook.**
# +
# import the helper function
from helpers import display_world
# Display the final world!
# define figure size
plt.rcParams["figure.figsize"] = (20,20)
# check if poses has been created
if 'poses' in locals():
# print out the last pose
print('Last pose: ', poses[-1])
# display the last position of the robot *and* the landmark positions
display_world(int(world_size), poses[-1], landmarks)
# -
# ### Question: How far away is your final pose (as estimated by `slam`) compared to the *true* final pose? Why do you think these poses are different?
#
# You can find the true value of the final pose in one of the first cells where `make_data` was called. You may also want to look at the true landmark locations and compare them to those that were estimated by `slam`. Ask yourself: what do you think would happen if we moved and sensed more (increased N)? Or if we had lower/higher noise parameters.
# **Answer**:
# * Movement and measurement noise both affect the estimation. A higher noise will increase variance of the data make the prediction more uncertain. Therefore, the noise should be kept as low as possible that predicted target value(s) and true target value(s) are as close as possible.
# ## Testing
#
# To confirm that your slam code works before submitting your project, it is suggested that you run it on some test data and cases. A few such cases have been provided for you, in the cells below. When you are ready, uncomment the test cases in the next cells (there are two test cases, total); your output should be **close-to or exactly** identical to the given results. If there are minor discrepancies it could be a matter of floating point accuracy or in the calculation of the inverse matrix.
#
# ### Submit your project
#
# If you pass these tests, it is a good indication that your project will pass all the specifications in the project rubric. Follow the submission instructions to officially submit!
# +
# Here is the data and estimated outputs for test case 1
test_data1 = [[[[1, 19.457599255548065, 23.8387362100849], [2, -13.195807561967236, 11.708840328458608], [3, -30.0954905279171, 15.387879242505843]], [-12.2607279422326, -15.801093326936487]], [[[2, -0.4659930049620491, 28.088559771215664], [4, -17.866382374890936, -16.384904503932]], [-12.2607279422326, -15.801093326936487]], [[[4, -6.202512900833806, -1.823403210274639]], [-12.2607279422326, -15.801093326936487]], [[[4, 7.412136480918645, 15.388585962142429]], [14.008259661173426, 14.274756084260822]], [[[4, -7.526138813444998, -0.4563942429717849]], [14.008259661173426, 14.274756084260822]], [[[2, -6.299793150150058, 29.047830407717623], [4, -21.93551130411791, -13.21956810989039]], [14.008259661173426, 14.274756084260822]], [[[1, 15.796300959032276, 30.65769689694247], [2, -18.64370821983482, 17.380022987031367]], [14.008259661173426, 14.274756084260822]], [[[1, 0.40311325410337906, 14.169429532679855], [2, -35.069349468466235, 2.4945558982439957]], [14.008259661173426, 14.274756084260822]], [[[1, -16.71340983241936, -2.777000269543834]], [-11.006096015782283, 16.699276945166858]], [[[1, -3.611096830835776, -17.954019226763958]], [-19.693482634035977, 3.488085684573048]], [[[1, 18.398273354362416, -22.705102332550947]], [-19.693482634035977, 3.488085684573048]], [[[2, 2.789312482883833, -39.73720193121324]], [12.849049222879723, -15.326510824972983]], [[[1, 21.26897046581808, -10.121029799040915], [2, -11.917698965880655, -23.17711662602097], [3, -31.81167947898398, -16.7985673023331]], [12.849049222879723, -15.326510824972983]], [[[1, 10.48157743234859, 5.692957082575485], [2, -22.31488473554935, -5.389184118551409], [3, -40.81803984305378, -2.4703329790238118]], [12.849049222879723, -15.326510824972983]], [[[0, 10.591050242096598, -39.2051798967113], [1, -3.5675572049297553, 22.849456408289125], [2, -38.39251065320351, 7.288990306029511]], [12.849049222879723, -15.326510824972983]], [[[0, -3.6225556479370766, -25.58006865235512]], [-7.8874682868419965, -18.379005523261092]], [[[0, 1.9784503557879374, -6.5025974151499]], [-7.8874682868419965, -18.379005523261092]], [[[0, 10.050665232782423, 11.026385307998742]], [-17.82919359778298, 9.062000642947142]], [[[0, 26.526838150174818, -0.22563393232425621], [4, -33.70303936886652, 2.880339841013677]], [-17.82919359778298, 9.062000642947142]]]
## Test Case 1
##
# Estimated Pose(s):
# [50.000, 50.000]
# [37.858, 33.921]
# [25.905, 18.268]
# [13.524, 2.224]
# [27.912, 16.886]
# [42.250, 30.994]
# [55.992, 44.886]
# [70.749, 59.867]
# [85.371, 75.230]
# [73.831, 92.354]
# [53.406, 96.465]
# [34.370, 100.134]
# [48.346, 83.952]
# [60.494, 68.338]
# [73.648, 53.082]
# [86.733, 38.197]
# [79.983, 20.324]
# [72.515, 2.837]
# [54.993, 13.221]
# [37.164, 22.283]
# Estimated Landmarks:
# [82.679, 13.435]
# [70.417, 74.203]
# [36.688, 61.431]
# [18.705, 66.136]
# [20.437, 16.983]
### Uncomment the following three lines for test case 1 and compare the output to the values above ###
mu_1 = slam(test_data1, 20, 5, 100.0, 2.0, 2.0)
poses, landmarks = get_poses_landmarks(mu_1, 20)
print_all(poses, landmarks)
# +
# Here is the data and estimated outputs for test case 2
test_data2 = [[[[0, 26.543274387283322, -6.262538160312672], [3, 9.937396825799755, -9.128540360867689]], [18.92765331253674, -6.460955043986683]], [[[0, 7.706544739722961, -3.758467215445748], [1, 17.03954411948937, 31.705489938553438], [3, -11.61731288777497, -6.64964096716416]], [18.92765331253674, -6.460955043986683]], [[[0, -12.35130507136378, 2.585119104239249], [1, -2.563534536165313, 38.22159657838369], [3, -26.961236804740935, -0.4802312626141525]], [-11.167066095509824, 16.592065417497455]], [[[0, 1.4138633151721272, -13.912454837810632], [1, 8.087721200818589, 20.51845934354381], [3, -17.091723454402302, -16.521500551709707], [4, -7.414211721400232, 38.09191602674439]], [-11.167066095509824, 16.592065417497455]], [[[0, 12.886743222179561, -28.703968411636318], [1, 21.660953298391387, 3.4912891084614914], [3, -6.401401414569506, -32.321583037341625], [4, 5.034079343639034, 23.102207946092893]], [-11.167066095509824, 16.592065417497455]], [[[1, 31.126317672358578, -10.036784369535214], [2, -38.70878528420893, 7.4987265861424595], [4, 17.977218575473767, 6.150889254289742]], [-6.595520680493778, -18.88118393939265]], [[[1, 41.82460922922086, 7.847527392202475], [3, 15.711709540417502, -30.34633659912818]], [-6.595520680493778, -18.88118393939265]], [[[0, 40.18454208294434, -6.710999804403755], [3, 23.019508919299156, -10.12110867290604]], [-6.595520680493778, -18.88118393939265]], [[[3, 27.18579315312821, 8.067219022708391]], [-6.595520680493778, -18.88118393939265]], [[], [11.492663265706092, 16.36822198838621]], [[[3, 24.57154567653098, 13.461499960708197]], [11.492663265706092, 16.36822198838621]], [[[0, 31.61945290413707, 0.4272295085799329], [3, 16.97392299158991, -5.274596836133088]], [11.492663265706092, 16.36822198838621]], [[[0, 22.407381798735177, -18.03500068379259], [1, 29.642444125196995, 17.3794951934614], [3, 4.7969752441371645, -21.07505361639969], [4, 14.726069092569372, 32.75999422300078]], [11.492663265706092, 16.36822198838621]], [[[0, 10.705527984670137, -34.589764174299596], [1, 18.58772336795603, -0.20109708164787765], [3, -4.839806195049413, -39.92208742305105], [4, 4.18824810165454, 14.146847823548889]], [11.492663265706092, 16.36822198838621]], [[[1, 5.878492140223764, -19.955352450942357], [4, -7.059505455306587, -0.9740849280550585]], [19.628527845173146, 3.83678180657467]], [[[1, -11.150789592446378, -22.736641053247872], [4, -28.832815721158255, -3.9462962046291388]], [-19.841703647091965, 2.5113335861604362]], [[[1, 8.64427397916182, -20.286336970889053], [4, -5.036917727942285, -6.311739993868336]], [-5.946642674882207, -19.09548221169787]], [[[0, 7.151866679283043, -39.56103232616369], [1, 16.01535401373368, -3.780995345194027], [4, -3.04801331832137, 13.697362774960865]], [-5.946642674882207, -19.09548221169787]], [[[0, 12.872879480504395, -19.707592098123207], [1, 22.236710716903136, 16.331770792606406], [3, -4.841206109583004, -21.24604435851242], [4, 4.27111163223552, 32.25309748614184]], [-5.946642674882207, -19.09548221169787]]]
## Test Case 2
##
# Estimated Pose(s):
# [50.000, 50.000]
# [69.035, 45.061]
# [87.655, 38.971]
# [76.084, 55.541]
# [64.283, 71.684]
# [52.396, 87.887]
# [44.674, 68.948]
# [37.532, 49.680]
# [31.392, 30.893]
# [24.796, 12.012]
# [33.641, 26.440]
# [43.858, 43.560]
# [54.735, 60.659]
# [65.884, 77.791]
# [77.413, 94.554]
# [96.740, 98.020]
# [76.149, 99.586]
# [70.211, 80.580]
# [64.130, 61.270]
# [58.183, 42.175]
# Estimated Landmarks:
# [76.777, 42.415]
# [85.109, 76.850]
# [13.687, 95.386]
# [59.488, 39.149]
# [69.283, 93.654]
### Uncomment the following three lines for test case 2 and compare to the values above ###
mu_2 = slam(test_data2, 20, 5, 100.0, 2.0, 2.0)
poses, landmarks = get_poses_landmarks(mu_2, 20)
print_all(poses, landmarks)
|
Project3_Landmark Detection/3. Landmark Detection and Tracking.ipynb
|