code stringlengths 38 801k | repo_path stringlengths 6 263 |
|---|---|
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="7dRi_BDWErNf" colab_type="text"
# 
#
# Sponsored by the [BYU PCCL Lab](https://).
#
# # AI Dungeon 2 is currently down due to high download costs.
# # <a href="https://twitter.com/nickwalton00?ref_src=twsrc%5Etfw" class="twitter-follow-button" data-show-count="false">Follow @nickwalton00</a> on twitter for updates on when it will be available again.
#
# ## About
# * While you wait you can [read adventures others have had](https://aidungeon.io/)
# * [Read more](https://pcc.cs.byu.edu/2019/11/21/ai-dungeon-2-creating-infinitely-generated-text-adventures-with-deep-learning-language-models/) about how AI Dungeon 2 is made.
#
# * Please [support AI Dungeon 2](https://www.patreon.com/join/AIDungeon/) to help get it back up.
# + id="FKqlSCrpS9dH" colab_type="code" colab={}
# !git clone --depth 1 --branch master https://github.com/samchristenoliphant/AIDungeon/
# %cd AIDungeon
# !./install.sh
from IPython.display import clear_output
clear_output()
print("Download Complete!")
# + id="YjArwbWh6XwN" colab_type="code" colab={}
from IPython.display import Javascript
display(Javascript('''google.colab.output.setIframeHeight(0, true, {maxHeight: 5000})'''))
# !python play.py
| AIDungeon_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="J1wRG8laa8Pm"
# ## Arno's Engram keyboard layout
#
# Engram is a key layout optimized for comfortable and efficient touch typing in English
# created by [<NAME>](https://binarybottle.com),
# with [open source code](https://github.com/binarybottle/engram) to create other optimized key layouts.
# You can install the Engram layout on [Windows, macOS, and Linux](https://keyman.com/keyboards/engram)
# or [try it out online](https://keymanweb.com/#en,Keyboard_engram).
# An article is under review (see the [preprint](https://www.preprints.org/manuscript/202103.0287/v1) for an earlier (and superceded) version with description).
#
# Letters are optimally arranged according to ergonomics factors that promote reduction of lateral finger movements and more efficient typing of high-frequency letter pairs. The most common punctuation marks are logically grouped together in the middle columns and numbers are paired with mathematical and logic symbols (shown as pairs of default and Shift-key-accessed characters):
#
# [{ 1| 2= 3~ 4+ 5< 6> 7^ 8& 9% 0* ]} /\
# bB yY oO uU '( ") lL dD wW vV zZ #$ @`
# cC iI eE aA ,; .: hH tT sS nN qQ
# gG xX jJ kK -_ ?! rR mM fF pP
#
# Letter frequencies (Norvig, 2012), showing that the Engram layout emphasizes keys in the home row:
#
# B Y O U L D W V Z
# C I E A H T S N Q
# G X J K R M F P
#
# 53 59 272 97 145 136 60 38 3
# 119 270 445 287 180 331 232 258 4
# 67 8 6 19 224 90 86 76
#
# See below for a full description and comparisons with other key layouts.
#
# ### Standard diagonal keyboard (default and Shift-key layers)
# 
#
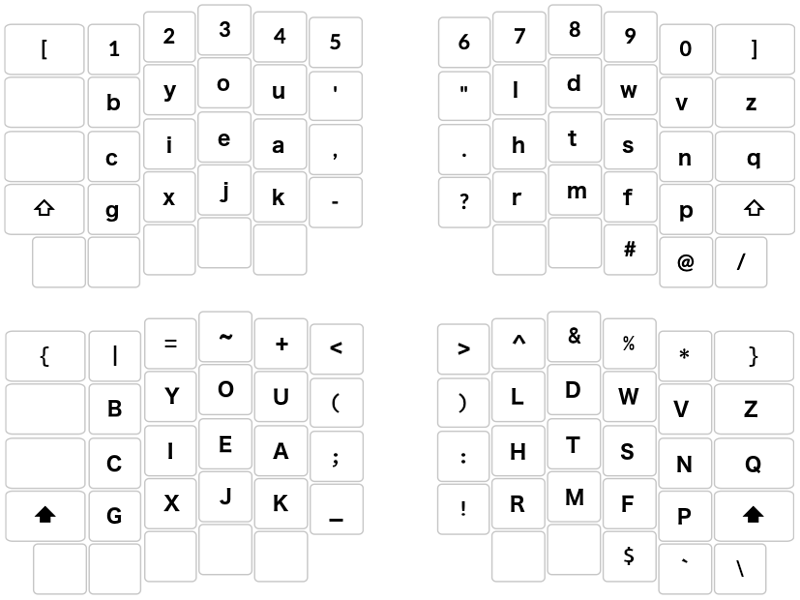
# ### "Ergonomic" orthonormal keyboard (default and Shift-key layers)
# 
#
# (c) 2021 <NAME>, MIT license
#
# ----------------
# + [markdown] colab_type="text" id="awscg4wBa8Po"
# # Contents
# 1. [Why a new keyboard layout?](#why)
# 2. [How does Engram compare with other key layouts?](#scores)
# 3. [Guiding criteria](#criteria)
# 4. Setup:
# - [Dependencies and functions](#import)
# - [Speed matrix](#speed)
# - [Strength matrix](#strength)
# - [Flow matrix and Engram scoring model](#flow)
# 5. Steps:
# - [Step 1: Define the shape of the key layout to minimize lateral finger movements](#step1)
# - [Step 2: Arrange the most frequent letters based on comfort and bigram frequencies](#step2)
# - [Step 3: Optimize assignment of the remaining letters](#step3)
# - [Step 4: Evaluate winning layout](#step4)
# - [Step 5: Arrange non-letter characters in easy-to-remember places](#step5)
# + [markdown] colab_type="text" id="SSdE4O9Wa8Pp"
# ## Why a new keyboard layout? <a name="why">
#
# **Personal history** <br>
# In the future, I hope to include an engaging rationale for why I took on this challenge.
# Suffice to say I love solving problems, and I have battled repetitive strain injury
# ever since I worked on an old DEC workstation at the MIT Media Lab while composing
# my thesis back in the 1990s.
# I have experimented with a wide variety of human interface technologies over the years --
# voice dictation, one-handed keyboard, keyless keyboard, foot mouse, and ergonomic keyboards
# like the Kinesis Advantage and [Ergodox](https://configure.ergodox-ez.com/ergodox-ez/layouts/APXBR/latest/0) keyboards with different key switches.
# While these technologies can significantly improve comfort and reduce strain,
# if you have to type on a keyboard, it can only help to use a key layout optimized according to sound ergonomics principles.
#
# I have used different key layouts (Qwerty, Dvorak, Colemak, etc.)
# for communications and for writing and programming projects,
# and have primarily relied on Colemak for the last 10 years.
# **I find that most to all of these key layouts:**
#
# - Demand too much strain on tendons
# - *strenuous lateral extension of the index and little fingers*
# - Ignore the ergonomics of the human hand
# - *different finger strengths*
# - *different finger lengths*
# - *natural roundedness of the hand*
# - *easier for shorter fingers to reach below than above longer fingers*
# - *easier for longer fingers to reach above than below shorter fingers*
# - *ease of little-to-index finger rolls vs. reverse*
# - Over-emphasize alternation between hands and under-emphasize same-hand, different-finger transitions
# - *same-row, adjacent finger transitions are easy and comfortable*
# - *little-to-index finger rolls are easy and comfortable*
#
# While I used ergonomics principles outlined below and the accompanying code to help generate the Engram layout,
# I also relied on massive bigram frequency data for the English language.
# if one were to follow the procedure below and use a different set of bigram frequencies for another language or text corpus,
# they could create a variant of the Engram layout, say "Engram-French", better suited to the French language.
#
# **Why "Engram"?** <br>
# The name is a pun, referring both to "n-gram", letter permutations and their frequencies that are used to compute the Engram layout, and "engram", or memory trace, the postulated change in neural tissue to account for the persistence of memory, as a nod to my attempt to make this layout easy to remember.
# + [markdown] colab_type="text" id="vkv2v3gla8Pt"
# ## How does Engram compare with other key layouts? <a name="scores">
#
# Below we compare the Engram layout with different prominent key layouts (Colemak, Dvorak, QWERTY, etc.) for some large, representative, publicly available data (all text sources are listed below and available on [GitHub](https://github.com/binarybottle/text_data)).
#
# #### Engram Scoring Model scores (x100) for layouts, based on publicly available text data
#
# Engram scores higher for all text and software sources than all other layouts according to its own scoring model (higher scores are better):
#
# | Layout | Google bigrams | Alice | Memento | Tweets_100K | Tweets_20K | Tweets_MASC | Spoken_MASC | COCA_blogs | iweb | Monkey | Coder | Rosetta |
# | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- |
# | Engram | 62.48 | 61.67 | 62.30 | 63.03 | 60.28 | 62.49 | 61.56 | 62.19 | 62.38 | 62.23 | 62.51 | 62.48 |
# | Halmak | 62.40 | 61.60 | 62.23 | 62.93 | 60.26 | 62.43 | 61.51 | 62.13 | 62.31 | 62.16 | 62.46 | 62.40 |
# | Hieamtsrn | 62.39 | 61.64 | 62.27 | 62.99 | 60.27 | 62.47 | 61.53 | 62.16 | 62.35 | 62.20 | 62.49 | 62.39 |
# | Norman | 62.35 | 61.57 | 62.20 | 62.86 | 60.21 | 62.39 | 61.47 | 62.08 | 62.27 | 62.12 | 62.40 | 62.35 |
# | Workman | 62.37 | 61.59 | 62.22 | 62.91 | 60.23 | 62.41 | 61.49 | 62.10 | 62.29 | 62.14 | 62.43 | 62.37 |
# | MTGap 2.0 | 62.32 | 61.59 | 62.21 | 62.88 | 60.22 | 62.39 | 61.49 | 62.09 | 62.28 | 62.13 | 62.42 | 62.32 |
# | QGMLWB | 62.31 | 61.58 | 62.21 | 62.90 | 60.25 | 62.40 | 61.49 | 62.10 | 62.29 | 62.14 | 62.43 | 62.31 |
# | Colemak Mod-DH | 62.36 | 61.60 | 62.22 | 62.90 | 60.26 | 62.41 | 61.49 | 62.12 | 62.30 | 62.16 | 62.44 | 62.36 |
# | Colemak | 62.36 | 61.58 | 62.20 | 62.89 | 60.25 | 62.40 | 61.48 | 62.10 | 62.29 | 62.14 | 62.43 | 62.36 |
# | Asset | 62.34 | 61.56 | 62.18 | 62.86 | 60.25 | 62.37 | 61.46 | 62.07 | 62.25 | 62.10 | 62.39 | 62.34 |
# | Capewell-Dvorak | 62.29 | 61.56 | 62.17 | 62.86 | 60.20 | 62.36 | 61.47 | 62.06 | 62.24 | 62.10 | 62.37 | 62.29 |
# | Klausler | 62.34 | 61.58 | 62.20 | 62.89 | 60.25 | 62.39 | 61.48 | 62.09 | 62.27 | 62.12 | 62.41 | 62.34 |
# | Dvorak | 62.31 | 61.56 | 62.17 | 62.85 | 60.23 | 62.35 | 61.46 | 62.06 | 62.24 | 62.09 | 62.35 | 62.31 |
# | QWERTY | 62.19 | 61.49 | 62.08 | 62.72 | 60.17 | 62.25 | 61.39 | 61.96 | 62.13 | 61.99 | 62.25 | 62.19 |
#
# ---
#
# [Keyboard Layout Analyzer](http://patorjk.com/keyboard-layout-analyzer/) (KLA) scores for the same text sources
#
# > The optimal layout score is based on a weighted calculation that factors in the distance your fingers moved (33%), how often you use particular fingers (33%), and how often you switch fingers and hands while typing (34%).
#
# Engram scores highest for 7 of the 9 and second highest for 2 of the 9 text sources; Engram scores third and fourth highest for the two software sources, "Coder" and "Rosetta" (higher scores are better):
#
# | Layout | Alice in Wonderland | Memento screenplay | 100K tweets | 20K tweets | MASC tweets | MASC spoken | COCA blogs | iweb | Monkey | Coder | Rosetta |
# | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- |
# | Engram | 70.13 | 57.16 | 64.64 | 58.58 | 60.24 | 64.39 | 69.66 | 68.25 | 67.66 | 46.81 | 47.69 |
# | Halmak | 66.25 | 55.03 | 60.86 | 55.53 | 57.13 | 62.32 | 67.29 | 65.50 | 64.75 | 45.68 | 47.60 |
# | Hieamtsrn | 69.43 | 56.75 | 64.40 | 58.95 | 60.47 | 64.33 | 69.93 | 69.15 | 68.30 | 46.01 | 46.48 |
# | Colemak Mod-DH | 65.74 | 54.91 | 60.75 | 54.94 | 57.15 | 61.29 | 67.12 | 65.98 | 64.85 | 47.35 | 48.50 |
# | Norman | 62.76 | 52.33 | 57.43 | 53.24 | 53.90 | 59.97 | 62.80 | 60.90 | 59.82 | 43.76 | 46.01 |
# | Workman | 64.78 | 54.29 | 59.98 | 55.81 | 56.25 | 61.34 | 65.27 | 63.76 | 62.90 | 45.33 | 47.76 |
# | MTGAP 2.0 | 66.13 | 53.78 | 59.87 | 55.30 | 55.81 | 60.32 | 65.68 | 63.81 | 62.74 | 45.38 | 44.34 |
# | QGMLWB | 65.45 | 54.07 | 60.51 | 56.05 | 56.90 | 62.23 | 66.26 | 64.76 | 63.91 | 46.38 | 45.72 |
# | Colemak | 65.83 | 54.94 | 60.67 | 54.97 | 57.04 | 61.36 | 67.14 | 66.01 | 64.91 | 47.30 | 48.65 |
# | Asset | 64.60 | 53.84 | 58.66 | 54.72 | 55.35 | 60.81 | 64.71 | 63.17 | 62.44 | 45.54 | 47.52 |
# | Capewell-Dvorak | 66.94 | 55.66 | 62.14 | 56.85 | 57.99 | 62.83 | 66.95 | 65.23 | 64.70 | 45.30 | 45.62 |
# | Klausler | 68.24 | 59.91 | 62.57 | 56.45 | 58.34 | 64.04 | 68.34 | 66.89 | 66.31 | 46.83 | 45.66 |
# | Dvorak | 65.86 | 58.18 | 60.93 | 55.56 | 56.59 | 62.75 | 66.64 | 64.87 | 64.26 | 45.46 | 45.55 |
# | QWERTY | 53.06 | 43.74 | 48.28 | 44.99 | 44.59 | 51.79 | 52.31 | 50.19 | 49.18 | 38.46 | 39.89 |
#
# ---
#
# #### Keyboard Layout Analyzer consecutive same-finger key presses
#
# KLA (and other) distance measures may not accurately reflect natural typing, so below is a more reliable measure of one source of effort and strain -- the tally of consecutive key presses with the same finger for different keys. Engram scores lowest for 6 of the 11 texts, second lowest for two texts, and third or fifth lowest for three texts, two of which are software text sources (lower scores are better):
#
# KLA (and other) distance measures may not accurately reflect natural typing, so below is a more reliable measure of one source of effort and strain -- the tally of consecutive key presses with the same finger for different keys. Engram scores lowest for 6 of the 9 and second or third lowest for 3 of the 9 text sources, and third or fifth lowest for the two software text sources (lower scores are better):
#
# | Layout | Alice | Memento | Tweets_100K | Tweets_20K | Tweets_MASC | Spoken_MASC | COCA_blogs | iweb | Monkey | Coder | Rosetta |
# | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- | --- |
# | Engram | 216 | 11476 | 320406 | 120286 | 7728 | 3514 | 137290 | 1064640 | 37534 | 125798 | 5822 |
# | Halmak | 498 | 13640 | 484702 | 170064 | 11456 | 5742 | 268246 | 2029634 | 68858 | 144790 | 5392 |
# | Hieamtsrn | 244 | 12096 | 311000 | 119490 | 8316 | 3192 | 155674 | 1100116 | 40882 | 158698 | 7324 |
# | Norman | 938 | 20012 | 721602 | 213890 | 16014 | 9022 | 595168 | 3885282 | 135844 | 179752 | 7402 |
# | Workman | 550 | 13086 | 451280 | 136692 | 10698 | 6156 | 287622 | 1975564 | 71150 | 132526 | 5550 |
# | MTGap 2.0 | 226 | 14550 | 397690 | 139130 | 10386 | 6252 | 176724 | 1532844 | 58144 | 138484 | 7272 |
# | QGMLWB | 812 | 17820 | 637788 | 189700 | 14364 | 7838 | 456442 | 3027530 | 100750 | 149366 | 8062 |
# | Colemak Mod-DH | 362 | 10960 | 352578 | 151736 | 9298 | 4644 | 153984 | 1233770 | 47438 | 117842 | 5328 |
# | Colemak | 362 | 10960 | 352578 | 151736 | 9298 | 4644 | 153984 | 1233770 | 47438 | 117842 | 5328 |
# | Asset | 520 | 12519 | 519018 | 155246 | 11802 | 5664 | 332860 | 2269342 | 77406 | 140886 | 6020 |
# | Capewell-Dvorak | 556 | 14226 | 501178 | 163878 | 12214 | 6816 | 335056 | 2391416 | 78152 | 151194 | 9008 |
# | Klausler | 408 | 14734 | 455658 | 174998 | 11410 | 5212 | 257878 | 1794604 | 59566 | 135782 | 7444 |
# | Dvorak | 516 | 13970 | 492604 | 171488 | 12208 | 5912 | 263018 | 1993346 | 64994 | 142084 | 6484 |
#
# ---
#
# #### Symmetry, switching, and roll measures
#
# The measures of hand symmetry, hand switching, finger switching, and hand runs without row jumps are from the [Carpalx](http://mkweb.bcgsc.ca/carpalx/?keyboard_layouts) website and are based on literature from the Gutenberg Project. Engram ties for highest score on two of the measures, third highest for hand switching (since it emphasizes hand rolls), and has a median value for hand runs (higher absolute scores are considered better).
#
# The roll measures are the number of bigrams (in billions of instances from Norvig's analysis of Google data) that engage inward rolls (little-to-index sequences), within the four columns of one hand, or any column across two hands. Engram scores second highest for the 32-keys and highest for the 24-keys, where the latter ensures that we are comparing Engram's letters with letters in other layouts (higher scores are better):
#
# | Layout | hand symmetry (%, right<0) | hand switching (%) | finger switching (%) | hand runs without row jumps (%) | inward rolls, billions (32 keys) | inward rolls, billions (24 keys) |
# | --- | --- | --- | --- | --- | --- | --- |
# | Engram | -99 | 61 | 93 | 82 | 4.64 | 4.51 |
# | Hieamtsrn | -96 | 59 | 93 | 85 | 4.69 | 4.16 |
# | Halmak | 99 | 63 | 93 | 81 | 4.59 | 4.25 |
# | Norman | 95 | 52 | 90 | 77 | 3.99 | 3.61 |
# | Workman | 95 | 52 | 93 | 79 | 4.16 | 3.63 |
# | MTGAP 2.0 | 98 | 48 | 93 | 76 | 3.96 | 3.58 |
# | QGMLWB | -97 | 57 | 91 | 84 | 4.36 | 2.81 |
# | Colemak Mod-DH | -94 | 52 | 93 | 78 | 4.15 | 3.51 |
# | Colemak | -94 | 52 | 93 | 83 | 4.17 | 3.16 |
# | Asset | 96 | 52 | 91 | 82 | 4.03 | 3.05 |
# | Capewell-Dvorak | -91 | 59 | 92 | 82 | 4.39 | 3.66 |
# | Klausler | -94 | 62 | 93 | 86 | 4.42 | 3.52 |
# | Dvorak | -86 | 62 | 93 | 84 | 4.40 | 3.20 |
# | QWERTY | 85 | 51 | 89 | 68 | 3.62 | 2.13 |
#
# ---
#
# | Layout | Year | Website |
# | --- | --- | --- |
# | Engram | 2021 | https://engram.dev |
# | [Halmak 2.2](https://keyboard-design.com/letterlayout.html?layout=halmak-2-2.en.ansi) | 2016 | https://github.com/MadRabbit/halmak |
# | [Hieamtsrn](https://www.keyboard-design.com/letterlayout.html?layout=hieamtsrn.en.ansi) | 2014 | https://mathematicalmulticore.wordpress.com/the-keyboard-layout-project/#comment-4976 |
# | [Colemak Mod-DH](https://keyboard-design.com/letterlayout.html?layout=colemak-mod-DH-full.en.ansi) | 2014 | https://colemakmods.github.io/mod-dh/ |
# | [Norman](https://keyboard-design.com/letterlayout.html?layout=norman.en.ansi) | 2013 | https://normanlayout.info/ |
# | [Workman](https://keyboard-design.com/letterlayout.html?layout=workman.en.ansi) | 2010 | https://workmanlayout.org/ |
# | [MTGAP 2.0](https://www.keyboard-design.com/letterlayout.html?layout=mtgap-2-0.en.ansi) | 2010 | https://mathematicalmulticore.wordpress.com/2010/06/21/mtgaps-keyboard-layout-2-0/ |
# | [QGMLWB](https://keyboard-design.com/letterlayout.html?layout=qgmlwb.en.ansi) | 2009 | http://mkweb.bcgsc.ca/carpalx/?full_optimization |
# | [Colemak](https://keyboard-design.com/letterlayout.html?layout=colemak.en.ansi) | 2006 | https://colemak.com/ |
# | [Asset](https://keyboard-design.com/letterlayout.html?layout=asset.en.ansi) | 2006 | http://millikeys.sourceforge.net/asset/ |
# | Capewell-Dvorak | 2004 | http://michaelcapewell.com/projects/keyboard/layout_capewell-dvorak.htm |
# | [Klausler](https://www.keyboard-design.com/letterlayout.html?layout=klausler.en.ansi) | 2002 | https://web.archive.org/web/20031001163722/http://klausler.com/evolved.html |
# | [Dvorak](https://keyboard-design.com/letterlayout.html?layout=dvorak.en.ansi) | 1936 | https://en.wikipedia.org/wiki/Dvorak_keyboard_layout |
# | [QWERTY](https://keyboard-design.com/letterlayout.html?layout=qwerty.en.ansi) | 1873 | https://en.wikipedia.org/wiki/QWERTY |
#
# ---
#
# | Text source | Information |
# | --- | --- |
# | "Alice in Wonderland" | Alice in Wonderland (Ch.1) |
# | "Memento screenplay" | [Memento screenplay](https://www.dailyscript.com/scripts/memento.html) |
# | "100K tweets" | 100,000 tweets from: [Sentiment140 dataset](https://data.world/data-society/twitter-user-data) training data |
# | "20K tweets" | 20,000 tweets from [Gender Classifier Data](https://www.kaggle.com/crowdflower/twitter-user-gender-classification) |
# | "MASC tweets" | [MASC](http://www.anc.org/data/masc/corpus/) tweets (cleaned of html markup) |
# | "MASC spoken" | [MASC](http://www.anc.org/data/masc/corpus/) spoken transcripts (phone and face-to-face: 25,783 words) |
# | "COCA blogs" | [Corpus of Contemporary American English](https://www.english-corpora.org/coca/) [blog samples](https://www.corpusdata.org/) |
# | "Rosetta" | "Tower of Hanoi" (programming languages A-Z from [Rosetta Code](https://rosettacode.org/wiki/Towers_of_Hanoi)) |
# | "Monkey text" | Ian Douglas's English-generated [monkey0-7.txt corpus](https://zenodo.org/record/4642460) |
# | "Coder text" | Ian Douglas's software-generated [coder0-7.txt corpus](https://zenodo.org/record/4642460) |
# | "iweb cleaned corpus" | First 150,000 lines of Shai Coleman's [iweb-corpus-samples-cleaned.txt](https://colemak.com/pub/corpus/iweb-corpus-samples-cleaned.txt.xz) |
#
# Reference for Monkey and Coder texts:
# <NAME>. (2021, March 28). Keyboard Layout Analysis: Creating the Corpus, Bigram Chains, and Shakespeare's Monkeys (Version 1.0.0). Zenodo. http://doi.org/10.5281/zenodo.4642460
# + [markdown] colab_type="text" id="wm3T-hmja8Ps"
# ## Guiding criteria <a name="criteria">
#
# 1. Assign letters to keys that don't require lateral finger movements.
# 2. Promote alternating between hands over uncomfortable same-hand transitions.
# 3. Assign the most common letters to the most comfortable keys.
# 4. Arrange letters so that more frequent bigrams are easier to type.
# 5. Promote little-to-index-finger roll-ins over index-to-little-finger roll-outs.
# 6. Balance finger loads according to their relative strength.
# 7. Avoid stretching shorter fingers up and longer fingers down.
# 8. Avoid using the same finger.
# 9. Avoid skipping over the home row.
# 10. Assign the most common punctuation to keys in the middle of the keyboard.
# 11. Assign easy-to-remember symbols to the Shift-number keys.
#
# ### Factors used to compute the Engram layout <a name="factors">
# - **N-gram letter frequencies** <br>
#
# [Peter Norvig's analysis](http://www.norvig.com/mayzner.html) of data from Google's book scanning project
# - **Flow factors** (transitions between ordered key pairs) <br>
# These factors are influenced by Dvorak's 11 criteria (1936).
# + [markdown] colab_type="text" id="2eTQ4jxPa8Pv"
# ### Import dependencies and functions <a name="import">
# +
# # %load code/engram_variables.py
# Print .png figures and .txt text files
print_output = False # True
# Apply strength data
apply_strength = True
min_strength_factor = 0.9
letters24 = ['E','T','A','O','I','N','S','R','H','L','D','C','U','M','F','P','G','W','Y','B','V','K','X','J']
keys24 = [1,2,3,4, 5,6,7,8, 9,10,11,12, 13,14,15,16, 17,18,19,20, 21,22,23,24]
instances24 = [4.45155E+11,3.30535E+11,2.86527E+11,2.72277E+11,2.69732E+11,2.57771E+11,
2.32083E+11,2.23768E+11,1.80075E+11,1.44999E+11,1.36018E+11,1.19156E+11,
97273082907,89506734085,85635440629,76112599849,66615316232,59712390260,
59331661972,52905544693,37532682260,19261229433,8369138754,5657910830]
max_frequency = 4.45155E+11 #1.00273E+11
instances_denominator = 1000000000000
# Establish which layouts are within a small difference of the top-scoring layout
# (the smallest difference between two penalties, 0.9^8 - 0.9^9, in one of 24^2 key pairs):
delta = 0.9**8 - 0.9**9
factor24 = ((24**2 - 1) + (1-delta)) / (24**2)
factor32 = ((32**2 - 1) + (1-delta)) / (32**2)
# Establish which layouts are within a small difference of each other when using the speed matrix.
# We define an epsilon equal to 13.158 ms for a single bigram (of the 32^2 possible bigrams),
# where 13.158 ms is one tenth of 131.58 ms, the fastest measured digraph tapping speed (30,000/228 = 131.58 ms)
# recorded in the study: "Estimation of digraph costs for keyboard layout optimization",
# A Iseri, <NAME>, International Journal of Industrial Ergonomics, 48, 127-138, 2015.
#data_matrix_speed = Speed32x32
#time_range = 243 # milliseconds
#norm_range = np.max(data_matrix_speed) - np.min(data_matrix_speed) # 0.6535662299854439
#ms_norm = norm_range / time_range # 0.0026895729629030614
#epsilon = 131.58/10 * ms_norm / (32**2)
epsilon = 0.00003549615849447514
# + colab={"base_uri": "https://localhost:8080/", "height": 71} colab_type="code" id="q1wNgX_FDzRH" outputId="7c14cebc-a4b7-4a77-d14f-26cbc7690c28"
# # %load code/engram_functions.py
# Import dependencies
import xlrd
import numpy as np
from sympy.utilities.iterables import multiset_permutations
import matplotlib
import matplotlib.pyplot as plt
import seaborn as sns
def permute_optimize_keys(fixed_letters, fixed_letter_indices, open_letter_indices,
all_letters, keys, data_matrix, bigrams, bigram_frequencies,
min_score=0, verbose=False):
"""
Find all permutations of letters, optimize layout, and generate output.
"""
matrix_selected = select_keys(data_matrix, keys, verbose=False)
unassigned_letters = []
for all_letter in all_letters:
if all_letter not in fixed_letters:
unassigned_letters.append(all_letter)
if len(unassigned_letters) == len(open_letter_indices):
break
letter_permutations = permute_letters(unassigned_letters, verbose)
if verbose:
print("{0} permutations".format(len(letter_permutations)))
top_permutation, top_score = optimize_layout(np.array([]), matrix_selected, bigrams, bigram_frequencies,
letter_permutations, open_letter_indices,
fixed_letters, fixed_letter_indices, min_score, verbose)
return top_permutation, top_score, letter_permutations
def permute_optimize(starting_permutation, letters, all_letters, all_keys,
data_matrix, bigrams, bigram_frequencies, min_score=0, verbose=False):
"""
Find all permutations of letters, optimize layout, and generate output.
"""
matrix_selected = select_keys(data_matrix, all_keys, verbose=False)
open_positions = []
fixed_positions = []
open_letters = []
fixed_letters = []
assigned_letters = []
for iletter, letter in enumerate(letters):
if letter.strip() == "":
open_positions.append(iletter)
for all_letter in all_letters:
if all_letter not in letters and all_letter not in assigned_letters:
open_letters.append(all_letter)
assigned_letters.append(all_letter)
break
else:
fixed_positions.append(iletter)
fixed_letters.append(letter)
letter_permutations = permute_letters(open_letters, verbose)
if verbose:
print("{0} permutations".format(len(letter_permutations)))
top_permutation, top_score = optimize_layout(starting_permutation, matrix_selected, bigrams,
bigram_frequencies, letter_permutations, open_positions,
fixed_letters, fixed_positions, min_score, verbose)
return top_permutation, top_score
def select_keys(data_matrix, keys, verbose=False):
"""
Select keys to quantify pairwise relationships.
"""
# Extract pairwise entries for the keys:
nkeys = len(keys)
Select = np.zeros((nkeys, nkeys))
u = 0
for i in keys:
u += 1
v = 0
for j in keys:
v += 1
Select[u-1,v-1] = data_matrix[i-1,j-1]
# Normalize matrix with min-max scaling to a range with max 1:
newMin = np.min(Select) / np.max(Select)
newMax = 1.0
Select = newMin + (Select - np.min(Select)) * (newMax - newMin) / (np.max(Select) - np.min(Select))
if verbose:
# Heatmap of array
heatmap(data=Select, title="Matrix heatmap", xlabel="Key 1", ylabel="Key 2", print_output=False); plt.show()
return Select
def permute_letters(letters, verbose=False):
"""
Find all permutations of a given set of letters (max: 8-10 letters).
"""
letter_permutations = []
for p in multiset_permutations(letters):
letter_permutations.append(p)
letter_permutations = np.array(letter_permutations)
return letter_permutations
def score_layout(data_matrix, letters, bigrams, bigram_frequencies, verbose=False):
"""
Compute the score for a given letter-key layout (NOTE normalization step).
"""
# Create a matrix of bigram frequencies:
nletters = len(letters)
F2 = np.zeros((nletters, nletters))
# Find the bigram frequency for each ordered pair of letters in the permutation:
for i1 in range(nletters):
for i2 in range(nletters):
bigram = letters[i1] + letters[i2]
i2gram = np.where(bigrams == bigram)
if np.size(i2gram) > 0:
F2[i1, i2] = bigram_frequencies[i2gram][0]
# Normalize matrices with min-max scaling to a range with max 1:
newMax = 1
minF2 = np.min(F2)
maxF2 = np.max(F2)
newMin2 = minF2 / maxF2
F2 = newMin + (F2 - minF2) * (newMax - newMin2) / (maxF2 - minF2)
# Compute the score for this permutation:
score = np.average(data_matrix * F2)
if verbose:
print("Score for letter permutation {0}: {1}".format(letters, score))
return score
def tally_bigrams(input_text, bigrams, normalize=True, verbose=False):
"""
Compute the score for a given letter-key layout (NOTE normalization step).
"""
# Find the bigram frequency for each ordered pair of letters in the input text
#input_text = [str.upper(str(x)) for x in input_text]
input_text = [str.upper(x) for x in input_text]
nchars = len(input_text)
F = np.zeros(len(bigrams))
for ichar in range(0, nchars-1):
bigram = input_text[ichar] + input_text[ichar + 1]
i2gram = np.where(bigrams == bigram)
if np.size(i2gram) > 0:
F[i2gram] += 1
# Normalize matrix with min-max scaling to a range with max 1:
if normalize:
newMax = 1
newMin = np.min(F) / np.max(F)
F = newMin + (F - np.min(F)) * (newMax - newMin) / (np.max(F) - np.min(F))
bigram_frequencies_for_input = F
if verbose:
print("Bigram frequencies for input: {0}".format(bigram_frequencies_for_input))
return bigram_frequencies_for_input
def tally_layout_samefinger_bigrams(layout, bigrams, bigram_frequencies, nkeys=32, verbose=False):
"""
Tally the number of same-finger bigrams within (a list of 24 letters representing) a layout:
['P','Y','O','U','C','I','E','A','G','K','J','X','M','D','L','B','R','T','N','S','H','V','W','F']
"""
if nkeys == 32:
# Left: Right:
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
same_finger_keys = [[1,5],[5,9],[1,9], [2,6],[6,10],[2,10],
[3,7],[7,11],[3,11], [4,8],[8,12],[4,12],
[25,26],[26,27],[25,27], [28,29],[29,30],[28,30], [31,32],
[4,25],[4,26],[4,27], [8,25],[8,26],[8,27], [12,25],[12,26],[12,27],
[13,28],[13,29],[13,30], [17,28],[17,29],[17,30], [21,28],[21,29],[21,30],
[31,16],[31,20],[31,24], [32,16],[32,20],[32,24],
[13,17],[17,21],[13,21], [14,18],[18,22],[14,22],
[15,19],[19,23],[15,23], [16,20],[20,24],[16,24]]
elif nkeys == 24:
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
same_finger_keys = [[1,5],[5,9],[1,9], [2,6],[6,10],[2,10],
[3,7],[7,11],[3,11], [4,8],[8,12],[4,12],
[13,17],[17,21],[13,21], [14,18],[18,22],[14,22],
[15,19],[19,23],[15,23], [16,20],[20,24],[16,24]]
layout = [str.upper(x) for x in layout]
max_frequency = 1.00273E+11
samefinger_bigrams = []
samefinger_bigram_counts = []
for bigram_keys in same_finger_keys:
bigram1 = layout[bigram_keys[0]-1] + layout[bigram_keys[1]-1]
bigram2 = layout[bigram_keys[1]-1] + layout[bigram_keys[0]-1]
i2gram1 = np.where(bigrams == bigram1)
i2gram2 = np.where(bigrams == bigram2)
if np.size(i2gram1) > 0:
samefinger_bigrams.append(bigram1)
samefinger_bigram_counts.append(max_frequency * bigram_frequencies[i2gram1] / np.max(bigram_frequencies))
if np.size(i2gram2) > 0:
samefinger_bigrams.append(bigram2)
samefinger_bigram_counts.append(max_frequency * bigram_frequencies[i2gram2] / np.max(bigram_frequencies))
samefinger_bigrams_total = np.sum([x[0] for x in samefinger_bigram_counts])
if verbose:
print(" Total same-finger bigram frequencies: {0:15.0f}".format(samefinger_bigrams_total))
return samefinger_bigrams, samefinger_bigram_counts, samefinger_bigrams_total
def tally_layout_bigram_rolls(layout, bigrams, bigram_frequencies, nkeys=32, verbose=False):
"""
Tally the number of bigrams that engage little-to-index finger inward rolls
for (a list of 24 or 32 letters representing) a layout,
within the four columns of one hand, or any column across two hands.
layout = ['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','B','V','N','T','R','S','H','M','W','F']
bigram_rolls, bigram_roll_counts, bigram_rolls_total = tally_layout_bigram_rolls(layout, bigrams, bigram_frequencies, nkeys=24, verbose=True)
"""
if nkeys == 32:
# Left: Right:
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
roll_keys = [[1,2],[2,3],[3,4], [5,6],[6,7],[7,8], [9,10],[10,11],[11,12],
[16,15],[15,14],[14,13], [20,19],[19,18],[18,17], [24,23],[23,22],[22,21],
[1,3],[2,4],[1,4], [5,7],[6,8],[5,8], [9,11],[10,12],[9,12],
[16,14],[15,13],[16,13], [20,18],[19,17],[20,17], [24,22],[23,21],[24,21],
[1,6],[1,7],[1,8],[2,7],[2,8],[3,8],
[5,2],[5,3],[5,4],[6,3],[6,4],[7,4],
[5,10],[5,11],[5,12],[6,11],[6,12],[7,12],
[9,6],[9,7],[9,8],[10,7],[10,8],[11,8],
[16,19],[16,18],[16,17],[15,18],[15,17],[14,17],
[20,15],[20,14],[20,13],[19,14],[19,13],[18,13],
[20,23],[20,22],[20,21],[19,22],[19,21],[18,21],
[24,19],[24,18],[24,17],[23,18],[23,17],[22,17],
[1,10],[1,11],[1,12],[2,11],[2,12],[3,12],
[9,2],[9,3],[9,4],[10,3],[10,4],[11,4],
[16,23],[16,22],[16,21],[15,22],[15,21],[14,21],
[24,15],[24,14],[24,13],[23,14],[23,13],[22,13]]
for i in [1,2,3,4,5,6,7,8,9,10,11,12, 25,26,27]:
for j in [13,14,15,16,17,18,19,20,21,22,23,24, 28,29,30,31,32]:
roll_keys.append([i,j])
for i in [13,14,15,16,17,18,19,20,21,22,23,24, 28,29,30,31,32]:
for j in [1,2,3,4,5,6,7,8,9,10,11,12, 25,26,27]:
roll_keys.append([i,j])
elif nkeys == 24:
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
roll_keys = [[1,2],[2,3],[3,4], [5,6],[6,7],[7,8], [9,10],[10,11],[11,12],
[16,15],[15,14],[14,13], [20,19],[19,18],[18,17], [24,23],[23,22],[22,21],
[1,3],[2,4],[1,4], [5,7],[6,8],[5,8], [9,11],[10,12],[9,12],
[16,14],[15,13],[16,13], [20,18],[19,17],[20,17], [24,22],[23,21],[24,21],
[1,6],[1,7],[1,8],[2,7],[2,8],[3,8], [5,2],[5,3],[5,4],[6,3],[6,4],[7,4],
[5,10],[5,11],[5,12],[6,11],[6,12],[7,12], [9,6],[9,7],[9,8],[10,7],[10,8],[11,8],
[16,19],[16,18],[16,17],[15,18],[15,17],[14,17], [20,15],[20,14],[20,13],[19,14],[19,13],[18,13],
[20,23],[20,22],[20,21],[19,22],[19,21],[18,21], [24,19],[24,18],[24,17],[23,18],[23,17],[22,17],
[1,10],[1,11],[1,12],[2,11],[2,12],[3,12], [9,2],[9,3],[9,4],[10,3],[10,4],[11,4],
[16,23],[16,22],[16,21],[15,22],[15,21],[14,21], [24,15],[24,14],[24,13],[23,14],[23,13],[22,13]]
for i in range(0,12):
for j in range(12,24):
roll_keys.append([i,j])
for i in range(12,24):
for j in range(0,12):
roll_keys.append([i,j])
layout = [str.upper(x) for x in layout]
max_frequency = 1.00273E+11
bigram_rolls = []
bigram_roll_counts = []
for bigram_keys in roll_keys:
bigram1 = layout[bigram_keys[0]-1] + layout[bigram_keys[1]-1]
bigram2 = layout[bigram_keys[1]-1] + layout[bigram_keys[0]-1]
i2gram1 = np.where(bigrams == bigram1)
i2gram2 = np.where(bigrams == bigram2)
if np.size(i2gram1) > 0:
bigram_rolls.append(bigram1)
bigram_roll_counts.append(max_frequency * bigram_frequencies[i2gram1] / np.max(bigram_frequencies))
if np.size(i2gram2) > 0:
bigram_rolls.append(bigram2)
bigram_roll_counts.append(max_frequency * bigram_frequencies[i2gram2] / np.max(bigram_frequencies))
bigram_rolls_total = np.sum([x[0] for x in bigram_roll_counts])
if verbose:
print(" Total bigram inward roll frequencies: {0:15.0f}".format(bigram_rolls_total))
return bigram_rolls, bigram_roll_counts, bigram_rolls_total
def optimize_layout(starting_permutation, data_matrix, bigrams, bigram_frequencies, letter_permutations,
open_positions, fixed_letters, fixed_positions=[], min_score=0, verbose=False):
"""
Compute scores for all letter-key layouts.
"""
top_permutation = starting_permutation
top_score = min_score
use_score_function = False
nletters = len(open_positions) + len(fixed_positions)
F2 = np.zeros((nletters, nletters))
# Loop through the permutations of the selected letters:
for p in letter_permutations:
letters = np.array(['W' for x in range(nletters)]) # KEEP to initialize!
for imove, open_position in enumerate(open_positions):
letters[open_position] = p[imove]
for ifixed, fixed_position in enumerate(fixed_positions):
letters[fixed_position] = fixed_letters[ifixed]
# Compute the score for this permutation:
if use_score_function:
score = score_layout(data_matrix, letters, bigrams, bigram_frequencies, verbose=False)
else:
# Find the bigram frequency for each ordered pair of letters in the permutation:
for i1 in range(nletters):
for i2 in range(nletters):
bigram = letters[i1] + letters[i2]
i2gram = np.where(bigrams == bigram)
if np.size(i2gram) > 0:
F2[i1, i2] = bigram_frequencies[i2gram][0]
# Normalize matrices with min-max scaling to a range with max 1:
newMax = 1
minF2 = np.min(F2)
maxF2 = np.max(F2)
newMin2 = minF2 / maxF2
F = newMin + (F2 - minF2) * (newMax - newMin2) / (maxF2 - minF2)
# Compute the score for this permutation:
score = np.average(data_matrix * F)
if score > top_score:
top_score = score
top_permutation = letters
if verbose:
if top_score == min_score:
print("top_score = min_score")
print("{0:0.8f}".format(top_score))
print(*top_permutation)
return top_permutation, top_score
def exchange_letters(letters, fixed_letter_indices, all_letters, all_keys, data_matrix,
bigrams, bigram_frequencies, verbose=True):
"""
Exchange letters, 8 keys at a time (8! = 40,320) selected twice in 14 different ways:
Indices:
0 1 2 3 12 13 14 15
4 5 6 7 16 17 18 19
8 9 10 11 20 21 22 23
1. Top rows
0 1 2 3 12 13 14 15
2. Bottom rows
8 9 10 11 20 21 22 23
3. Top and bottom rows on the right side
12 13 14 15
20 21 22 23
4. Top and bottom rows on the left side
0 1 2 3
8 9 10 11
5. Top right and bottom left rows
12 13 14 15
8 9 10 11
6. Top left and bottom right rows
0 1 2 3
20 21 22 23
7. Center of the top and bottom rows on both sides
1 2 13 14
9 10 21 22
8. The eight corners
0 3 12 15
8 11 20 23
9. Left half of the top and bottom rows on both sides
0 1 12 13
8 9 20 21
10. Right half of the top and bottom rows on both sides
2 3 14 15
10 11 22 23
11. Left half of non-home rows on the left and right half of the same rows on the right
0 1 14 15
8 9 22 23
12. Right half of non-home rows on the left and left half of the same rows on the right
2 3 12 13
10 11 20 21
13. Top center and lower sides
1 2 13 14
8 11 20 23
14. Top sides and lower center
0 3 12 15
9 10 21 22
15. Repeat 1-14
"""
top_score = score_layout(data_matrix, letters, bigrams, bigram_frequencies, verbose=False)
print('Initial score: {0}'.format(top_score))
print(*letters)
top_permutation = letters
lists_of_open_indices = [
[0,1,2,3,12,13,14,15],
[8,9,10,11,20,21,22,23],
[12,13,14,15,20,21,22,23],
[0,1,2,3,8,9,10,11],
[12,13,14,15,8,9,10,11],
[0,1,2,3,20,21,22,23],
[1,2,13,14,9,10,21,22],
[0,3,12,15,8,11,20,23],
[0,1,12,13,8,9,20,21],
[2,3,14,15,10,11,22,23],
[0,1,14,15,8,9,22,23],
[2,3,12,13,10,11,20,21],
[1,2,8,11,13,14,20,23],
[0,3,9,10,12,15,21,22]
]
lists_of_print_statements = [
'1. Top rows',
'2. Bottom rows',
'3. Top and bottom rows on the right side',
'4. Top and bottom rows on the left side',
'5. Top right and bottom left rows',
'6. Top left and bottom right rows',
'7. Center of the top and bottom rows on both sides',
'8. The eight corners',
'9. Left half of the top and bottom rows on both sides',
'10. Right half of the top and bottom rows on both sides',
'11. Left half of non-home rows on the left and right half of the same rows on the right',
'12. Right half of non-home rows on the left and left half of the same rows on the right',
'13. Top center and lower sides',
'14. Top sides and lower center'
]
for istep in [1,2]:
if istep == 1:
s = "Set 1: 14 letter exchanges: "
elif istep == 2:
s = "Set 2: 14 letter exchanges: "
for ilist, open_indices in enumerate(lists_of_open_indices):
print_statement = lists_of_print_statements[ilist]
if verbose:
print('{0} {1}'.format(s, print_statement))
starting_permutation = top_permutation.copy()
for open_index in open_indices:
if open_index not in fixed_letter_indices:
top_permutation[open_index] = ''
top_permutation, top_score = permute_optimize(starting_permutation, top_permutation, letters24,
keys24, data_matrix, bigrams, bigram_frequencies,
min_score=top_score, verbose=True)
if verbose:
print('')
print(' -------- DONE --------')
print('')
return top_permutation, top_score
def rank_within_epsilon(numbers, epsilon, factor=False, verbose=True):
"""
numbers = np.array([10,9,8,7,6])
epsilon = 1
rank_within_epsilon(numbers, epsilon, factor=False, verbose=True)
>>> array([1., 1., 2., 2., 3.])
numbers = np.array([0.798900824, 0.79899900824, 0.79900824])
epsilon = 0.9**8 - 0.9**9
factor24 = ((24**2 - 1) + (1-epsilon)) / (24**2) # 0.999925266109375
rank_within_epsilon(numbers, factor24, factor=True, verbose=True)
>>> array([2., 1., 1.])
"""
numbers = np.array(numbers)
Isort = np.argsort(-numbers)
numbers_sorted = numbers[Isort]
count = 1
ranks = np.zeros(np.size(numbers))
for i, num in enumerate(numbers_sorted):
if ranks[i] == 0:
if factor:
lower_bound = num * epsilon
else:
lower_bound = num - epsilon
bounded_nums1 = num >= numbers_sorted
bounded_nums2 = numbers_sorted >= lower_bound
bounded_nums = bounded_nums1 * bounded_nums2
count += 1
for ibounded, bounded_num in enumerate(bounded_nums):
if bounded_num == True:
ranks[ibounded] = count
uranks = np.unique(ranks)
nranks = np.size(uranks)
new_ranks = ranks.copy()
new_count = 0
for rank in uranks:
new_count += 1
same_ranks = ranks == rank
for isame, same_rank in enumerate(same_ranks):
if same_rank == True:
new_ranks[isame] = new_count
#ranks_sorted = new_ranks[Isort]
ranks_sorted = [np.int(x) for x in new_ranks]
if verbose:
for i, num in enumerate(numbers_sorted):
print(" ({0}) {1}".format(np.int(ranks_sorted[i]), num))
return numbers_sorted, ranks_sorted, Isort
def print_matrix_info(matrix_data, matrix_label, nkeys, nlines=10):
"""
Print matrix output.
"""
print("{0} min = {1}, max = {2}".format(matrix_label, np.min(matrix_data), np.max(matrix_data)))
matrix_flat = matrix_data.flatten()
argsort = np.argsort(matrix_flat)
print("{0} key number pairs with minimum values:".format(matrix_label))
for x in argsort[0:nlines]:
if x % nkeys == 0:
min_row = np.int(np.ceil(x / nkeys)) + 1
min_col = 1
else:
min_row = np.int(np.ceil(x / nkeys))
min_col = x - nkeys * (min_row-1) + 1
print(" {0} -> {1} ({2})".format(min_row, min_col, matrix_flat[x]))
print("{0} key number pairs with maximum values:".format(matrix_label))
max_sort = argsort[-nlines::]
for x in max_sort[::-1]:
if x % nkeys == 0:
max_row = np.int(np.ceil(x / nkeys)) + 1
max_col = 1
else:
max_row = np.int(np.ceil(x / nkeys))
max_col = x - nkeys * (max_row-1) + 1
print(" {0} -> {1} ({2})".format(max_row, max_col, matrix_flat[x]))
def heatmap(data, title="", xlabel="", ylabel="", x_axis_labels=[], y_axis_labels=[], print_output=True):
"""
Plot heatmap of matrix.
"""
# use heatmap function, set the color as viridis and
# make each cell seperate using linewidth parameter
plt.figure()
sns_plot = sns.heatmap(data, xticklabels=x_axis_labels, yticklabels=y_axis_labels, linewidths=1,
cmap="viridis", square=True, vmin=np.min(data), vmax=np.max(data))
plt.title(title)
plt.xlabel(xlabel)
plt.ylabel(ylabel)
sns_plot.set_xticklabels(x_axis_labels) #, rotation=75)
sns_plot.set_yticklabels(y_axis_labels)
if print_output:
sns_plot.figure.savefig("{0}_heatmap.png".format(title))
def histmap(data, title="", print_output=True):
"""
Plot histogram.
"""
sns.distplot(data)
plt.title(title)
if print_output:
sns_plot.figure.savefig("{0}_histogram.png".format(title))
def print_layout24(layout):
"""
Print layout.
"""
print(' {0} {1}'.format(' '.join(layout[0:4]),
' '.join(layout[12:16])))
print(' {0} {1}'.format(' '.join(layout[4:8]),
' '.join(layout[16:20])))
print(' {0} {1}'.format(' '.join(layout[8:12]),
' '.join(layout[20:24])))
def print_layout24_instances(layout, letters24, instances24, bigrams, bigram_frequencies):
"""
Print billions of instances per letter (not Z or Q) in layout form.
layout = ['P','Y','O','U','C','I','E','A','G','K','J','X','M','D','L','B','R','T','N','S','H','V','W','F']
print_layout24_instances(layout, letters24, instances24, bigrams, bigram_frequencies)
"""
layout_instances = []
layout_instances_strings = []
for letter in layout:
index = letters24.index(letter)
layout_instances.append(instances24[index])
layout_instances_strings.append('{0:3.0f}'.format(instances24[index]/instances_denominator))
print(' {0} {1}'.format(' '.join(layout_instances_strings[0:4]),
' '.join(layout_instances_strings[12:16])))
print(' {0} {1}'.format(' '.join(layout_instances_strings[4:8]),
' '.join(layout_instances_strings[16:20])))
print(' {0} {1}'.format(' '.join(layout_instances_strings[8:12]),
' '.join(layout_instances_strings[20:24])))
left_sum = np.sum(layout_instances[0:12])
right_sum = np.sum(layout_instances[12:24])
pL = ''
pR = ''
if left_sum > right_sum:
pL = ' ({0:3.2f}%)'.format(100 * (left_sum - right_sum) / right_sum)
elif right_sum > left_sum:
pR = ' ({0:3.2f}%)'.format(100 * (right_sum - left_sum) / left_sum)
print('\n left: {0}{1} right: {2}{3}'.format(left_sum, pL, right_sum, pR))
tally_layout_samefinger_bigrams(layout, bigrams, bigram_frequencies, nkeys=24, verbose=True)
tally_layout_bigram_rolls(layout, bigrams, bigram_frequencies, nkeys=24, verbose=True)
def print_bigram_frequency(input_pair, bigrams, bigram_frequencies):
"""
>>> print_bigram_frequency(['t','h'], bigrams, bigram_frequencies)
"""
# Find the bigram frequency
input_text = [str.upper(str(x)) for x in input_pair]
nchars = len(input_text)
for ichar in range(0, nchars-1):
bigram1 = input_text[ichar] + input_text[ichar + 1]
bigram2 = input_text[ichar + 1] + input_text[ichar]
i2gram1 = np.where(bigrams == bigram1)
i2gram2 = np.where(bigrams == bigram2)
if np.size(i2gram1) > 0:
freq1 = bigram_frequencies[i2gram1[0][0]]
print("{0}: {1:3.2f}".format(bigram1, freq1))
if np.size(i2gram2) > 0:
freq2 = bigram_frequencies[i2gram2[0][0]]
print("{0}: {1:3.2f}".format(bigram2, freq2))
# + [markdown] colab_type="text" id="rFiySi8rDzRN"
# ### Bigram frequencies <a name="ngrams">
#
# [<NAME>'s ngrams table](http://www.norvig.com/mayzner.html](http://www.norvig.com/mayzner.html)
#
# [NOTE: If you want to compute an optimized layout for another language, or based on another corpus, you can run the tally_bigrams() function above and replace bigram_frequencies below before running the rest of the code.]
# + colab={} colab_type="code" id="K68F0fkqDzRO"
# # %load code/load_bigram_frequencies.py
load_original_ngram_files = False
if load_original_ngram_files:
ngrams_table = "data/bigrams-trigrams-frequencies.xlsx"
wb = xlrd.open_workbook(ngrams_table)
ngrams_sheet = wb.sheet_by_index(0)
# 1-grams and frequencies
onegrams = np.array(())
onegram_frequencies = np.array(())
i = 0
start1 = 0
stop1 = 0
while stop1 == 0:
if ngrams_sheet.cell_value(i, 0) == "2-gram":
stop1 = 1
elif ngrams_sheet.cell_value(i, 0) == "1-gram":
start1 = 1
elif start1 == 1:
onegrams = np.append(onegrams, ngrams_sheet.cell_value(i, 0))
onegram_frequencies = np.append(onegram_frequencies, ngrams_sheet.cell_value(i, 1))
i += 1
onegram_frequencies = onegram_frequencies / np.sum(onegram_frequencies)
# 2-grams and frequencies
bigrams = np.array(())
bigram_frequencies = np.array(())
i = 0
start1 = 0
stop1 = 0
while stop1 == 0:
if ngrams_sheet.cell_value(i, 0) == "3-gram":
stop1 = 1
elif ngrams_sheet.cell_value(i, 0) == "2-gram":
start1 = 1
elif start1 == 1:
bigrams = np.append(bigrams, ngrams_sheet.cell_value(i, 0))
bigram_frequencies = np.append(bigram_frequencies, ngrams_sheet.cell_value(i, 1))
i += 1
bigram_frequencies = bigram_frequencies / np.sum(bigram_frequencies)
# Save:
if print_output:
file = open("onegrams.txt", "w+")
file.write(str(onegrams))
file.close()
file = open("onegram_frequencies.txt", "w+")
file.write(str(onegram_frequencies))
file.close()
file = open("bigrams.txt", "w+")
file.write(str(bigrams))
file.close()
file = open("bigram_frequencies.txt", "w+")
file.write(str(bigram_frequencies))
file.close()
# Print:
print(repr(onegrams))
print(repr(onegram_frequencies))
print(repr(bigrams))
print(repr(bigram_frequencies))
else:
onegrams = np.array(['E', 'T', 'A', 'O', 'I', 'N', 'S', 'R', 'H', 'L', 'D', 'C', 'U',
'M', 'F', 'P', 'G', 'W', 'Y', 'B', 'V', 'K', 'X', 'J', 'Q', 'Z'],
dtype='<U32')
onegram_frequencies = np.array([0.12492063, 0.09275565, 0.08040605, 0.07640693, 0.07569278,
0.07233629, 0.06512767, 0.06279421, 0.05053301, 0.04068986,
0.03816958, 0.03343774, 0.02729702, 0.02511761, 0.02403123,
0.02135891, 0.01869376, 0.01675664, 0.0166498 , 0.01484649,
0.01053252, 0.00540513, 0.00234857, 0.00158774, 0.00120469,
0.00089951])
bigrams = np.array(['TH', 'HE', 'IN', 'ER', 'AN', 'RE', 'ON', 'AT', 'EN', 'ND', 'TI',
'ES', 'OR', 'TE', 'OF', 'ED', 'IS', 'IT', 'AL', 'AR', 'ST', 'TO',
'NT', 'NG', 'SE', 'HA', 'AS', 'OU', 'IO', 'LE', 'VE', 'CO', 'ME',
'DE', 'HI', 'RI', 'RO', 'IC', 'NE', 'EA', 'RA', 'CE', 'LI', 'CH',
'LL', 'BE', 'MA', 'SI', 'OM', 'UR', 'CA', 'EL', 'TA', 'LA', 'NS',
'DI', 'FO', 'HO', 'PE', 'EC', 'PR', 'NO', 'CT', 'US', 'AC', 'OT',
'IL', 'TR', 'LY', 'NC', 'ET', 'UT', 'SS', 'SO', 'RS', 'UN', 'LO',
'WA', 'GE', 'IE', 'WH', 'EE', 'WI', 'EM', 'AD', 'OL', 'RT', 'PO',
'WE', 'NA', 'UL', 'NI', 'TS', 'MO', 'OW', 'PA', 'IM', 'MI', 'AI',
'SH', 'IR', 'SU', 'ID', 'OS', 'IV', 'IA', 'AM', 'FI', 'CI', 'VI',
'PL', 'IG', 'TU', 'EV', 'LD', 'RY', 'MP', 'FE', 'BL', 'AB', 'GH',
'TY', 'OP', 'WO', 'SA', 'AY', 'EX', 'KE', 'FR', 'OO', 'AV', 'AG',
'IF', 'AP', 'GR', 'OD', 'BO', 'SP', 'RD', 'DO', 'UC', 'BU', 'EI',
'OV', 'BY', 'RM', 'EP', 'TT', 'OC', 'FA', 'EF', 'CU', 'RN', 'SC',
'GI', 'DA', 'YO', 'CR', 'CL', 'DU', 'GA', 'QU', 'UE', 'FF', 'BA',
'EY', 'LS', 'VA', 'UM', 'PP', 'UA', 'UP', 'LU', 'GO', 'HT', 'RU',
'UG', 'DS', 'LT', 'PI', 'RC', 'RR', 'EG', 'AU', 'CK', 'EW', 'MU',
'BR', 'BI', 'PT', 'AK', 'PU', 'UI', 'RG', 'IB', 'TL', 'NY', 'KI',
'RK', 'YS', 'OB', 'MM', 'FU', 'PH', 'OG', 'MS', 'YE', 'UD', 'MB',
'IP', 'UB', 'OI', 'RL', 'GU', 'DR', 'HR', 'CC', 'TW', 'FT', 'WN',
'NU', 'AF', 'HU', 'NN', 'EO', 'VO', 'RV', 'NF', 'XP', 'GN', 'SM',
'FL', 'IZ', 'OK', 'NL', 'MY', 'GL', 'AW', 'JU', 'OA', 'EQ', 'SY',
'SL', 'PS', 'JO', 'LF', 'NV', 'JE', 'NK', 'KN', 'GS', 'DY', 'HY',
'ZE', 'KS', 'XT', 'BS', 'IK', 'DD', 'CY', 'RP', 'SK', 'XI', 'OE',
'OY', 'WS', 'LV', 'DL', 'RF', 'EU', 'DG', 'WR', 'XA', 'YI', 'NM',
'EB', 'RB', 'TM', 'XC', 'EH', 'TC', 'GY', 'JA', 'HN', 'YP', 'ZA',
'GG', 'YM', 'SW', 'BJ', 'LM', 'CS', 'II', 'IX', 'XE', 'OH', 'LK',
'DV', 'LP', 'AX', 'OX', 'UF', 'DM', 'IU', 'SF', 'BT', 'KA', 'YT',
'EK', 'PM', 'YA', 'GT', 'WL', 'RH', 'YL', 'HS', 'AH', 'YC', 'YN',
'RW', 'HM', 'LW', 'HL', 'AE', 'ZI', 'AZ', 'LC', 'PY', 'AJ', 'IQ',
'NJ', 'BB', 'NH', 'UO', 'KL', 'LR', 'TN', 'GM', 'SN', 'NR', 'FY',
'MN', 'DW', 'SB', 'YR', 'DN', 'SQ', 'ZO', 'OJ', 'YD', 'LB', 'WT',
'LG', 'KO', 'NP', 'SR', 'NQ', 'KY', 'LN', 'NW', 'TF', 'FS', 'CQ',
'DH', 'SD', 'VY', 'DJ', 'HW', 'XU', 'AO', 'ML', 'UK', 'UY', 'EJ',
'EZ', 'HB', 'NZ', 'NB', 'MC', 'YB', 'TP', 'XH', 'UX', 'TZ', 'BV',
'MF', 'WD', 'OZ', 'YW', 'KH', 'GD', 'BM', 'MR', 'KU', 'UV', 'DT',
'HD', 'AA', 'XX', 'DF', 'DB', 'JI', 'KR', 'XO', 'CM', 'ZZ', 'NX',
'YG', 'XY', 'KG', 'TB', 'DC', 'BD', 'SG', 'WY', 'ZY', 'AQ', 'HF',
'CD', 'VU', 'KW', 'ZU', 'BN', 'IH', 'TG', 'XV', 'UZ', 'BC', 'XF',
'YZ', 'KM', 'DP', 'LH', 'WF', 'KF', 'PF', 'CF', 'MT', 'YU', 'CP',
'PB', 'TD', 'ZL', 'SV', 'HC', 'MG', 'PW', 'GF', 'PD', 'PN', 'PC',
'RX', 'TV', 'IJ', 'WM', 'UH', 'WK', 'WB', 'BH', 'OQ', 'KT', 'RQ',
'KB', 'CG', 'VR', 'CN', 'PK', 'UU', 'YF', 'WP', 'CZ', 'KP', 'DQ',
'WU', 'FM', 'WC', 'MD', 'KD', 'ZH', 'GW', 'RZ', 'CB', 'IW', 'XL',
'HP', 'MW', 'VS', 'FC', 'RJ', 'BP', 'MH', 'HH', 'YH', 'UJ', 'FG',
'FD', 'GB', 'PG', 'TK', 'KK', 'HQ', 'FN', 'LZ', 'VL', 'GP', 'HZ',
'DK', 'YK', 'QI', 'LX', 'VD', 'ZS', 'BW', 'XQ', 'MV', 'UW', 'HG',
'FB', 'SJ', 'WW', 'GK', 'UQ', 'BG', 'SZ', 'JR', 'QL', 'ZT', 'HK',
'VC', 'XM', 'GC', 'FW', 'PZ', 'KC', 'HV', 'XW', 'ZW', 'FP', 'IY',
'PV', 'VT', 'JP', 'CV', 'ZB', 'VP', 'ZR', 'FH', 'YV', 'ZG', 'ZM',
'ZV', 'QS', 'KV', 'VN', 'ZN', 'QA', 'YX', 'JN', 'BF', 'MK', 'CW',
'JM', 'LQ', 'JH', 'KJ', 'JC', 'GZ', 'JS', 'TX', 'FK', 'JL', 'VM',
'LJ', 'TJ', 'JJ', 'CJ', 'VG', 'MJ', 'JT', 'PJ', 'WG', 'VH', 'BK',
'VV', 'JD', 'TQ', 'VB', 'JF', 'DZ', 'XB', 'JB', 'ZC', 'FJ', 'YY',
'QN', 'XS', 'QR', 'JK', 'JV', 'QQ', 'XN', 'VF', 'PX', 'ZD', 'QT',
'ZP', 'QO', 'DX', 'HJ', 'GV', 'JW', 'QC', 'JY', 'GJ', 'QB', 'PQ',
'JG', 'BZ', 'MX', 'QM', 'MZ', 'QF', 'WJ', 'ZQ', 'XR', 'ZK', 'CX',
'FX', 'FV', 'BX', 'VW', 'VJ', 'MQ', 'QV', 'ZF', 'QE', 'YJ', 'GX',
'KX', 'XG', 'QD', 'XJ', 'SX', 'VZ', 'VX', 'WV', 'YQ', 'BQ', 'GQ',
'VK', 'ZJ', 'XK', 'QP', 'HX', 'FZ', 'QH', 'QJ', 'JZ', 'VQ', 'KQ',
'XD', 'QW', 'JX', 'QX', 'KZ', 'WX', 'FQ', 'XZ', 'ZX'], dtype='<U32')
bigram_frequencies = np.array([3.55620339e-02, 3.07474124e-02, 2.43274529e-02, 2.04826481e-02,
1.98515108e-02, 1.85432319e-02, 1.75804642e-02, 1.48673230e-02,
1.45424846e-02, 1.35228145e-02, 1.34257882e-02, 1.33939375e-02,
1.27653906e-02, 1.20486963e-02, 1.17497528e-02, 1.16812337e-02,
1.12842988e-02, 1.12327374e-02, 1.08744953e-02, 1.07489847e-02,
1.05347566e-02, 1.04126653e-02, 1.04125115e-02, 9.53014842e-03,
9.32114579e-03, 9.25763559e-03, 8.71095073e-03, 8.70002319e-03,
8.34931851e-03, 8.29254235e-03, 8.25280566e-03, 7.93859725e-03,
7.93006486e-03, 7.64818391e-03, 7.63241814e-03, 7.27618866e-03,
7.26724441e-03, 6.98707488e-03, 6.91722265e-03, 6.88165290e-03,
6.85633031e-03, 6.51417363e-03, 6.24352184e-03, 5.97765978e-03,
5.76571076e-03, 5.76283716e-03, 5.65269345e-03, 5.50057242e-03,
5.46256885e-03, 5.42747781e-03, 5.38164098e-03, 5.30301559e-03,
5.29886071e-03, 5.27529444e-03, 5.08937452e-03, 4.92966405e-03,
4.87753568e-03, 4.84902069e-03, 4.77989185e-03, 4.77282719e-03,
4.74470916e-03, 4.64574958e-03, 4.60971757e-03, 4.54257059e-03,
4.47772200e-03, 4.42103298e-03, 4.31534618e-03, 4.25820178e-03,
4.25013516e-03, 4.15745843e-03, 4.12608242e-03, 4.05151268e-03,
4.05075209e-03, 3.97732158e-03, 3.96527277e-03, 3.94413046e-03,
3.86884200e-03, 3.85337077e-03, 3.85189513e-03, 3.84646388e-03,
3.78793431e-03, 3.77605408e-03, 3.74420703e-03, 3.73663638e-03,
3.67956418e-03, 3.65492648e-03, 3.61676413e-03, 3.61373182e-03,
3.60899233e-03, 3.47234973e-03, 3.45829494e-03, 3.39212478e-03,
3.37488213e-03, 3.36877623e-03, 3.30478042e-03, 3.23572471e-03,
3.17759946e-03, 3.17691369e-03, 3.16447752e-03, 3.15240004e-03,
3.15172398e-03, 3.11176534e-03, 2.95503911e-03, 2.89966768e-03,
2.87848219e-03, 2.86282435e-03, 2.84865969e-03, 2.84585627e-03,
2.81484803e-03, 2.69544349e-03, 2.62987083e-03, 2.54961380e-03,
2.54906719e-03, 2.54783715e-03, 2.52606379e-03, 2.47740122e-03,
2.39175226e-03, 2.36573195e-03, 2.33400171e-03, 2.29786417e-03,
2.27503360e-03, 2.27277101e-03, 2.23911052e-03, 2.21754315e-03,
2.18017446e-03, 2.17360835e-03, 2.14044590e-03, 2.13767970e-03,
2.13188615e-03, 2.10259217e-03, 2.04932647e-03, 2.04724906e-03,
2.03256516e-03, 2.02845908e-03, 1.96777866e-03, 1.95449429e-03,
1.95410531e-03, 1.91254221e-03, 1.89316385e-03, 1.88234971e-03,
1.87652262e-03, 1.84944194e-03, 1.83351654e-03, 1.78086545e-03,
1.76468430e-03, 1.75132925e-03, 1.71573739e-03, 1.70683303e-03,
1.66405086e-03, 1.63999785e-03, 1.62732115e-03, 1.62613977e-03,
1.60361051e-03, 1.54749379e-03, 1.51636562e-03, 1.51067364e-03,
1.49901610e-03, 1.49455831e-03, 1.49011351e-03, 1.48460771e-03,
1.48077067e-03, 1.47541326e-03, 1.47480347e-03, 1.46316579e-03,
1.46204465e-03, 1.43745726e-03, 1.41513491e-03, 1.39980075e-03,
1.38382616e-03, 1.36545598e-03, 1.36333253e-03, 1.36012483e-03,
1.35189358e-03, 1.32127808e-03, 1.30185876e-03, 1.28328757e-03,
1.27907576e-03, 1.26260675e-03, 1.23637099e-03, 1.23094105e-03,
1.21386641e-03, 1.20743055e-03, 1.19536134e-03, 1.19032774e-03,
1.17626124e-03, 1.16805780e-03, 1.14618533e-03, 1.11559852e-03,
1.06597119e-03, 1.05782134e-03, 1.04699320e-03, 1.04540205e-03,
1.01153313e-03, 9.97734501e-04, 9.86028683e-04, 9.84491816e-04,
9.79174450e-04, 9.78784303e-04, 9.70343472e-04, 9.68322624e-04,
9.66708177e-04, 9.60690121e-04, 9.59749105e-04, 9.43900197e-04,
9.40242103e-04, 9.28331656e-04, 9.26685761e-04, 9.14014864e-04,
9.02555222e-04, 8.92112065e-04, 8.85803335e-04, 8.77507468e-04,
8.62646840e-04, 8.57695087e-04, 8.54499050e-04, 8.43925356e-04,
8.31382851e-04, 8.23722323e-04, 8.16643644e-04, 7.89875969e-04,
7.86444549e-04, 7.42072946e-04, 7.36927617e-04, 7.27646949e-04,
7.25004577e-04, 7.11071849e-04, 6.92833068e-04, 6.71807283e-04,
6.68638321e-04, 6.56391013e-04, 6.51990243e-04, 6.49048818e-04,
6.43397537e-04, 6.43118050e-04, 6.37839069e-04, 6.21864133e-04,
6.06367626e-04, 5.99162639e-04, 5.87024289e-04, 5.74860663e-04,
5.72519573e-04, 5.68447140e-04, 5.58806800e-04, 5.45711864e-04,
5.37896691e-04, 5.34768852e-04, 5.20071483e-04, 5.18874875e-04,
5.16054649e-04, 5.14388309e-04, 5.11931727e-04, 5.04227393e-04,
5.00890900e-04, 4.97325634e-04, 4.75088970e-04, 4.66605249e-04,
4.58324041e-04, 4.29127437e-04, 4.27514542e-04, 4.17186146e-04,
4.16199437e-04, 3.94646924e-04, 3.94183167e-04, 3.86306652e-04,
3.61812839e-04, 3.50841120e-04, 3.49059129e-04, 3.23402665e-04,
3.22604151e-04, 3.11527347e-04, 3.10032877e-04, 3.07611603e-04,
2.96010489e-04, 2.88197255e-04, 2.77494857e-04, 2.70735751e-04,
2.67122244e-04, 2.64790886e-04, 2.64597695e-04, 2.63237166e-04,
2.61362824e-04, 2.59399816e-04, 2.58614910e-04, 2.57579773e-04,
2.49143242e-04, 2.49036616e-04, 2.47547306e-04, 2.36748821e-04,
2.35282013e-04, 2.32245156e-04, 2.30209194e-04, 2.28229670e-04,
2.27822992e-04, 2.20319919e-04, 2.17945603e-04, 2.13543715e-04,
1.97145202e-04, 1.90526970e-04, 1.90304866e-04, 1.88393786e-04,
1.85754127e-04, 1.85322815e-04, 1.81767370e-04, 1.74089940e-04,
1.71644610e-04, 1.71039222e-04, 1.69557657e-04, 1.66839046e-04,
1.64718022e-04, 1.59561636e-04, 1.57658164e-04, 1.54026397e-04,
1.52211752e-04, 1.51115808e-04, 1.47564559e-04, 1.46841709e-04,
1.36432949e-04, 1.35005671e-04, 1.32141796e-04, 1.27573620e-04,
1.27432415e-04, 1.26388914e-04, 1.25919175e-04, 1.23965197e-04,
1.21174483e-04, 1.18691292e-04, 1.18219114e-04, 1.17637524e-04,
1.17526303e-04, 1.13037594e-04, 1.10863960e-04, 1.09331046e-04,
1.08837112e-04, 1.06567401e-04, 1.05698197e-04, 1.00512685e-04,
1.00106518e-04, 9.85814937e-05, 9.17495595e-05, 9.15174736e-05,
9.09807382e-05, 8.79007001e-05, 8.16240791e-05, 7.91627682e-05,
7.79158645e-05, 7.56940333e-05, 7.44394656e-05, 7.18101849e-05,
6.97589276e-05, 6.81802488e-05, 6.69029567e-05, 6.54143249e-05,
6.08786925e-05, 6.07607969e-05, 6.03570614e-05, 5.98994801e-05,
5.95001291e-05, 5.94970869e-05, 5.86983574e-05, 5.79700512e-05,
5.66119466e-05, 5.50952209e-05, 5.47453912e-05, 5.43839597e-05,
5.25861529e-05, 4.89722417e-05, 4.78187439e-05, 4.77415865e-05,
4.77107257e-05, 4.62616737e-05, 4.60653783e-05, 4.60409299e-05,
4.56730211e-05, 4.54645078e-05, 4.52324283e-05, 4.38982745e-05,
4.36906610e-05, 4.33593810e-05, 4.31226640e-05, 4.29912118e-05,
4.29446346e-05, 4.17137339e-05, 3.93478837e-05, 3.84895449e-05,
3.84390172e-05, 3.81834469e-05, 3.53827628e-05, 3.47222349e-05,
3.37168917e-05, 3.18518637e-05, 3.15951703e-05, 3.12905207e-05,
3.10605585e-05, 3.02567524e-05, 2.91709879e-05, 2.89567711e-05,
2.85652293e-05, 2.82994071e-05, 2.80417376e-05, 2.77861205e-05,
2.77303518e-05, 2.76273746e-05, 2.72172235e-05, 2.69880432e-05,
2.66503046e-05, 2.66033916e-05, 2.62086568e-05, 2.59259584e-05,
2.57640153e-05, 2.56299050e-05, 2.54449453e-05, 2.51909823e-05,
2.47409597e-05, 2.46797892e-05, 2.42472084e-05, 2.35748710e-05,
2.24438116e-05, 2.24317329e-05, 2.23097275e-05, 2.21249597e-05,
2.17815183e-05, 2.15248592e-05, 2.09465192e-05, 2.09125513e-05,
1.96913177e-05, 1.95330853e-05, 1.91064697e-05, 1.88952009e-05,
1.85746459e-05, 1.81220081e-05, 1.78919334e-05, 1.73267658e-05,
1.61874055e-05, 1.60765855e-05, 1.58740992e-05, 1.45486411e-05,
1.40812264e-05, 1.36678429e-05, 1.32768479e-05, 1.31460479e-05,
1.30872012e-05, 1.29588223e-05, 1.25748548e-05, 1.24146066e-05,
1.22821602e-05, 1.22486357e-05, 1.20714645e-05, 1.20448925e-05,
1.19866728e-05, 1.18936663e-05, 1.17590888e-05, 1.17001978e-05,
1.16346360e-05, 1.11092945e-05, 1.08992577e-05, 1.06740258e-05,
1.06735218e-05, 1.06144296e-05, 1.05679067e-05, 1.03656570e-05,
1.03317955e-05, 9.98437559e-06, 9.01036943e-06, 8.85768061e-06,
8.76035160e-06, 8.60019167e-06, 8.19227801e-06, 7.80479658e-06,
7.53516931e-06, 7.44150882e-06, 7.30644125e-06, 7.26777599e-06,
7.06747616e-06, 6.95177332e-06, 6.85925126e-06, 6.74132156e-06,
6.71322068e-06, 6.70106994e-06, 6.66133186e-06, 6.47626505e-06,
6.38130476e-06, 6.29576510e-06, 6.24612583e-06, 5.93271496e-06,
5.92132104e-06, 5.83947722e-06, 5.76779879e-06, 5.76465728e-06,
5.53187023e-06, 5.47131015e-06, 5.33180695e-06, 5.22417954e-06,
5.20732008e-06, 5.15949060e-06, 5.11569104e-06, 4.95336950e-06,
4.94557425e-06, 4.73636484e-06, 4.63955858e-06, 4.53340156e-06,
4.22935422e-06, 4.19307790e-06, 4.17347414e-06, 4.12142146e-06,
4.11855764e-06, 3.80541311e-06, 3.36707879e-06, 3.29563656e-06,
3.17577578e-06, 3.05442971e-06, 2.98983688e-06, 2.97762691e-06,
2.95066092e-06, 2.91720550e-06, 2.89840858e-06, 2.77497857e-06,
2.76265227e-06, 2.74176112e-06, 2.70310579e-06, 2.61648976e-06,
2.60275585e-06, 2.56616744e-06, 2.55465117e-06, 2.49712549e-06,
2.42815484e-06, 2.37933375e-06, 2.35040476e-06, 2.33914845e-06,
2.33036549e-06, 2.32978989e-06, 2.28930419e-06, 2.28804340e-06,
2.26346210e-06, 2.24353844e-06, 2.23182640e-06, 2.23165865e-06,
2.22696341e-06, 2.22115030e-06, 2.21572164e-06, 2.20668084e-06,
2.19243658e-06, 2.17382266e-06, 2.08159887e-06, 2.07762818e-06,
1.95415065e-06, 1.88693410e-06, 1.83219245e-06, 1.81431726e-06,
1.67631850e-06, 1.67169206e-06, 1.63803449e-06, 1.57770706e-06,
1.56577585e-06, 1.53130790e-06, 1.52519015e-06, 1.52439998e-06,
1.49350905e-06, 1.47212210e-06, 1.45715861e-06, 1.40331777e-06,
1.38641504e-06, 1.29786439e-06, 1.27069447e-06, 1.25613209e-06,
1.23105569e-06, 1.22268909e-06, 1.21688094e-06, 1.18065108e-06,
1.18060143e-06, 1.16794389e-06, 1.13216621e-06, 1.12716419e-06,
1.12418866e-06, 1.12412659e-06, 1.05684621e-06, 1.05049722e-06,
1.04986594e-06, 1.03676402e-06, 1.03482230e-06, 9.96847192e-07,
9.75926251e-07, 9.54397081e-07, 9.36101632e-07, 9.30100914e-07,
9.27467975e-07, 8.92801774e-07, 8.85217179e-07, 8.58891337e-07,
7.80484800e-07, 7.67724409e-07, 7.54031637e-07, 7.45052550e-07,
7.32511689e-07, 7.06828122e-07, 6.59585949e-07, 6.40055245e-07,
6.18628925e-07, 6.17142222e-07, 6.09904832e-07, 6.07242457e-07,
5.72270900e-07, 5.49823535e-07, 5.22568859e-07, 5.01838721e-07,
4.91372576e-07, 4.82981856e-07, 4.69688423e-07, 4.59727658e-07,
4.54795508e-07, 4.22875379e-07, 4.13494116e-07, 3.99834682e-07,
3.97288987e-07, 3.87644926e-07, 3.84245584e-07, 3.81268632e-07,
3.67029696e-07, 3.57267536e-07, 3.52642869e-07, 3.51058992e-07,
3.44112772e-07, 3.36167495e-07, 3.24215712e-07, 3.23810344e-07,
3.21814716e-07, 3.21505459e-07, 3.10936465e-07, 2.88018831e-07,
2.86309762e-07, 2.76140106e-07, 2.63218703e-07, 2.56899508e-07,
2.51244222e-07, 2.25386521e-07, 2.15766576e-07, 2.03018243e-07,
1.99078411e-07, 1.97551987e-07, 1.96981706e-07, 1.92415912e-07,
1.84391194e-07, 1.81253585e-07, 1.78663913e-07, 1.77747846e-07,
1.59541769e-07, 1.38003378e-07, 1.36499298e-07, 1.22889160e-07,
1.22576357e-07, 1.19711121e-07, 1.09597855e-07, 9.97477409e-08,
9.65292710e-08, 9.36271510e-08, 9.35785637e-08, 9.34540807e-08,
8.40270671e-08, 7.82629028e-08, 7.54898762e-08, 6.64058115e-08,
5.96748649e-08, 5.79118882e-08, 5.73650143e-08, 5.65688198e-08,
5.34673852e-08, 5.34237630e-08, 5.29956976e-08, 4.84174907e-08,
3.83818937e-08])
# + [markdown] colab_type="text" id="46wIL5xzDzRS"
# ## Speed matrix <a name="speed">
# ### 24x24 relative Speed matrix between key pair (averaged for left/right symmetry)
#
# - does not take into account order of key pairs (see Flow24x24 matrix)
# - the original version was constructed with data from right-handed people
# - 24 keys that don't require extending index or little fingers ("home block keys")
#
# ### Home block keys
#
# Left: Right:
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
#
# Interkey stroke times in milliseconds from Table 3 of <br>
# "Estimation of digraph costs for keyboard layout optimization", <br>
# A Iseri, Ma Eksioglu, International Journal of Industrial Ergonomics, 48, 127-138, 2015. <br>
# Key numbering in article and in spreadsheet:
#
# Left: Right:
# 1 4 7 10 13 16 19 22 25 28 31
# 2 5 8 11 14 17 20 23 26 29 32
# 3 6 9 12 15 18 21 24 27 30
#
# ### Load table of interkey speeds
# + colab={} colab_type="code" id="095yG4iPDzRT"
# # %load data/Time24x24.py
# code/load_original_interkey_speeds.py
# Left: Right:
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
Time24x24 = np.array([
[196,225,204,164,266,258,231,166,357,325,263,186,169,176,178,186,156,156,158,163,171,175,177,189],
[225,181,182,147,239,245,196,150,289,296,229,167,162,169,170,178,148,148,150,155,163,167,169,182],
[204,182,170,149,196,194,232,155,237,214,263,166,157,164,165,173,143,143,145,150,158,163,164,177],
[164,147,149,169,160,161,157,226,165,185,234,257,154,162,163,171,141,141,143,148,156,160,162,175],
[266,239,196,160,196,240,208,166,271,267,208,169,143,150,151,160,129,129,132,137,145,149,151,163],
[258,245,194,161,240,181,183,149,245,256,184,150,138,145,146,154,124,124,126,131,139,144,145,158],
[231,196,232,157,208,183,170,149,201,215,239,151,134,141,142,150,120,120,122,127,135,140,141,154],
[166,150,155,226,166,149,149,169,160,147,170,221,133,140,141,150,119,119,122,126,135,139,141,153],
[357,289,237,165,271,245,201,160,196,236,194,161,171,178,179,188,157,157,160,164,173,177,179,191],
[325,296,214,185,267,256,215,147,236,181,184,157,166,173,174,182,152,152,154,159,167,172,173,186],
[263,229,263,234,208,184,239,170,194,184,170,150,159,166,167,176,145,145,148,153,161,165,167,179],
[186,167,166,257,169,150,151,221,161,157,150,169,153,160,161,169,139,139,141,146,154,159,160,173],
[169,162,157,154,143,138,134,133,171,166,159,153,151,147,141,145,188,151,142,164,213,204,162,149],
[176,169,164,162,150,145,141,140,178,173,166,160,147,151,189,209,137,207,191,206,149,227,208,197],
[178,170,165,163,151,146,142,141,179,174,167,161,141,189,157,253,136,188,210,231,155,226,239,276],
[186,178,173,171,160,154,150,150,188,182,176,169,145,209,253,170,147,206,251,233,164,268,362,271],
[156,148,143,141,129,124,120,119,157,152,145,139,188,137,136,147,151,133,138,152,192,149,139,144],
[156,148,143,141,129,124,120,119,157,152,145,139,151,207,188,206,133,151,179,183,145,204,183,201],
[158,150,145,143,132,126,122,122,160,154,148,141,142,191,210,251,138,179,157,240,145,185,208,229],
[163,155,150,148,137,131,127,126,164,159,153,146,164,206,231,233,152,183,240,170,160,220,293,242],
[171,163,158,156,145,139,135,135,173,167,161,154,213,149,155,164,192,145,145,160,151,140,142,145],
[175,167,163,160,149,144,140,139,177,172,165,159,204,227,226,268,149,204,185,220,140,151,175,188],
[177,169,164,162,151,145,141,141,179,173,167,160,162,208,239,362,139,183,208,293,142,175,157,230],
[189,182,177,175,163,158,154,153,191,186,179,173,149,197,276,271,144,201,229,242,145,188,230,170]])
# +
# # %load code/load_interkey_speeds24x24.py
# Left/right symmetric version of the Time24x24 matrix
# (The original version was constructed with data from right-handed people.)
# <NAME>, <NAME> / International Journal of Industrial Ergonomics 48 (2015) 127e138
# Left: Right:
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
I = [ 1, 2, 3, 4, 5, 6, 7, 8, 9,10,11,12, 16,15,14,13, 20,19,18,17, 24,23,22,21]
J = [16,15,14,13, 20,19,18,17, 24,23,22,21, 1, 2, 3, 4, 5, 6, 7, 8, 9,10,11,12]
TimeSymmetric24x24 = np.ones((24,24))
for i1, I1 in enumerate(I):
for i2, I2 in enumerate(I):
J1 = J[i1] - 1
J2 = J[i2] - 1
avgvalue = (Time24x24[I1-1,I2-1] + Time24x24[J1,J2]) / 2
#print(Time24x24[I1-1,I2-1], Time24x24[J1,J2], avgvalue)
TimeSymmetric24x24[I1-1,I2-1] = avgvalue
TimeSymmetric24x24[J1,J2] = avgvalue
# Normalize matrix with min-max scaling to a range with maximum = 1:
newMin = np.min(Time24x24) / np.max(Time24x24)
newMax = 1.0
Time24x24 = newMin + (Time24x24 - np.min(Time24x24)) * (newMax - newMin) / (np.max(Time24x24) - np.min(Time24x24))
# Convert relative interkey stroke times to relative speeds by subtracting from 1:
Speed24x24 = 1 - Time24x24 + np.min(Time24x24)
# Normalize matrix with min-max scaling to a range with maximum = 1:
newMin = np.min(TimeSymmetric24x24) / np.max(TimeSymmetric24x24)
newMax = 1.0
TimeSymmetric24x24 = newMin + (TimeSymmetric24x24 - np.min(TimeSymmetric24x24)) * (newMax - newMin) / (np.max(TimeSymmetric24x24) - np.min(TimeSymmetric24x24))
# Convert relative interkey stroke times to relative speeds by subtracting from 1:
SpeedSymmetric24x24 = 1 - TimeSymmetric24x24 + np.min(TimeSymmetric24x24)
# Print:
#print_matrix_info(matrix_data=Speed24x24, matrix_label="Speed24x24", nkeys=24, nlines=50)
#heatmap(data=Speed24x24, title="Speed24x24", xlabel="Key 1", ylabel="Key 2", print_output=print_output)
# + [markdown] colab_type="text" id="tFfuA8zMDzRg"
# ## Strength matrix <a name="strength">
#
# ### Relative finger position STRENGTH matrix
#
# Finger strengths are based on peak keyboard reaction forces (in newtons) from Table 4 of <br>
# "Keyboard Reaction Force and Finger Flexor Electromyograms during Computer Keyboard Work" <br>
# BJ Martin, TJ Armstrong, <NAME>, S Natarajan, Human Factors,1996,38(4),654-664:
#
# middle 2.36
# index 2.26
# ring 2.02
# little 1.84
#
# index/middle: 0.9576271186440678
# ring/middle: 0.8559322033898306
# little/middle: 0.7796610169491526
#
# For reference, Table 1 of "Ergonomic keyboard layout designed for the Filipino language", 2016 (doi: 10.1007/978-3-319-41694-6_41) presents "average finger strength of Filipinos [n=30, ages 16-36] measured in pounds":
#
# L R
# little 3.77 4.27
# ring 4.54 5.08
# middle 5.65 6.37
# index 6.09 6.57
#
# 6.57/4.27 = 1.54
# 6.09/3.77 = 1.62
# 6.37/5.08 = 1.25
# 5.65/4.54 = 1.24
#
# We won't use these results as I don't feel they represent relative strength relevant for typing: "Respondents were asked to sit in upright position, with their wrists resting on a flat surface. A pinch gauge was placed within each finger's reach. The respondents were asked to exert maximum pressure on the device."
#
# The following does not take into account order of key pairs (see Flow matrix).
#
# +
# # %load code/load_strength_data.py
# Normalize by the highest peak force (middle finger):
middle_force = 2.36
index_force = 2.26
ring_force = 2.02
little_force = 1.84
middle_norm = 1.0
index_norm = index_force / middle_force
ring_norm = ring_force / middle_force
little_norm = little_force / middle_force
print('index/middle: {0}'.format(index_norm))
print('ring/middle: {0}'.format(ring_norm))
print('little/middle: {0}'.format(little_norm))
# Relative left/right hand strength (assume equal):
lf = 1.0
rf = 1.0
strengths24 = np.array((
lf * little_norm, lf * ring_norm, lf * middle_norm, lf * index_norm,
lf * little_norm, lf * ring_norm, lf * middle_norm, lf * index_norm,
lf * little_norm, lf * ring_norm, lf * middle_norm, lf * index_norm,
rf * index_norm, rf * middle_norm, rf * ring_norm, rf * little_norm,
rf * index_norm, rf * middle_norm, rf * ring_norm, rf * little_norm,
rf * index_norm, rf * middle_norm, rf * ring_norm, rf * little_norm))
# Create a finger-pair position strength matrix by adding pairs of strength values:
Strength24x24 = np.zeros((24, 24))
for i in range(24):
Strength24x24[i,:] = strengths24
Strength24x24 = (Strength24x24 + Strength24x24.transpose())
# Normalize matrix with min-max scaling to a range with maximum = 1:
#newMin = strength_factor
newMin = min_strength_factor # np.min(Strength24x24) / np.max(Strength24x24)
newMax = 1.0
Strength24x24 = newMin + (Strength24x24 - np.min(Strength24x24)) * (newMax - newMin) / (np.max(Strength24x24) - np.min(Strength24x24))
# Print:
print_matrix_info(matrix_data=Strength24x24, matrix_label="Strength24x24", nkeys=24, nlines=10)
heatmap(data=Strength24x24, title="Strength24x24", xlabel="Key 1", ylabel="Key 2", print_output=print_output)
# Save:
if print_output:
file = open("Strength24x24.txt", "w+")
file.write(str(Strength24x24))
file.close()
penalty = 1.0 # Penalty for lateral (index, little) finger placement (1 = no penalty)
strengths32 = np.array((lf * little_norm, lf * ring_norm, lf * middle_norm, lf * index_norm,
lf * little_norm, lf * ring_norm, lf * middle_norm, lf * index_norm,
lf * little_norm, lf * ring_norm, lf * middle_norm, lf * index_norm,
rf * index_norm, rf * middle_norm, rf * ring_norm, rf * little_norm,
rf * index_norm, rf * middle_norm, rf * ring_norm, rf * little_norm,
rf * index_norm, rf * middle_norm, rf * ring_norm, rf * little_norm,
lf * index_norm * penalty, lf * index_norm * penalty, lf * index_norm * penalty,
rf * index_norm * penalty, rf * index_norm * penalty, rf * index_norm * penalty,
rf * little_norm * penalty, rf * little_norm * penalty))
# Create a finger-pair position strength matrix by adding pairs of strength values:
Strength32x32 = np.zeros((32, 32))
for i in range(32):
Strength32x32[i,:] = strengths32
Strength32x32 = (Strength32x32 + Strength32x32.transpose())
# Normalize matrix with min-max scaling to a range with maximum = 1:
newMin = np.min(Strength32x32) / np.max(Strength32x32)
newMax = 1.0
Strength32x32 = newMin + (Strength32x32 - np.min(Strength32x32)) * (newMax - newMin) / (np.max(Strength32x32) - np.min(Strength32x32))
# Print:
print_matrix_info(matrix_data=Strength32x32, matrix_label="Strength32x32", nkeys=32, nlines=10)
heatmap(data=Strength32x32, title="Strength32x32", xlabel="Key 1", ylabel="Key 2", print_output=print_output)
# Save:
if print_output:
file = open("Strength32x32.txt", "w+")
file.write(str(Strength32x32))
file.close()
# + [markdown] colab_type="text" id="Dnn5-8S5DzRy"
# ## Flow matrix and Engram scoring model <a name="flow">
#
# The Flow24x24 matrix takes into account ease of transition between ordered pairs of keys.
#
# Our optimization algorithm finds every permutation of a given set of letters, maps these letter permutations to a set of keys, and ranks these letter-key mappings according to a score reflecting ease of typing key pairs and frequency of letter pairs (bigrams). The score is the average of the scores for all possible bigrams in this arrangement. The score for each bigram is a product of the frequency of occurrence of that bigram, the frequency of each of the bigram’s characters, and flow, strength (and optional speed) factors for the key pair.
#
# #### Dvorak et al. (1936) defined eleven criteria for the design and evaluation of keyboard layouts:
# 1. Deviation from the balance of hand and finger loads should be as low as possible.
# 2. Percentage of tapping with the same fingers should be as low as possible.
# 3. Percentage of tapping that includes top row should be as low as possible.
# 4. Percentage of tapping that includes bottom row should be as low as possible.
# 5. Percentage of tapping in the home row should be as high as possible.
# 6. Percentage of tapping by alternating hands should be as high as possible.
# 7. Percentage of hurdles with the same finger should be as low as possible.
# 8. Percentage of hurdles with adjacent offset fingers should be as low as possible.
# 9. Percentage of hurdles with remote fingers should be as low as possible.
# 10. Percentage of reach with the same finger should be as low as possible.
# 11. Percentage of reach with adjacent offset fingers should be as low as possible.
#
# #### Synopsis of above criteria for pairwise key presses when touch typing:
# 1. Alternate between hands.
# 2. Balance finger loads, and avoid using the same finger.
# 3. Avoid the upper and lower rows, and avoid skipping over the home row.
# 4. Avoid tapping adjacent offset rows with the same or adjacent offset fingers.
#
# ### Factors to penalize strenuous key transitions
#
# Direction:
#
# - outward = 0.9: outward roll of fingers from the index to little finger (same hand)
#
# Dexterity:
#
# - side_above_3away = 0.9
# - index and little finger type two keys, one or more rows apart (same hand)
# - side_above_2away = 0.9^2 = 0.81
# - index finger types key a row or two above ring finger key, or
# - little finger types key a row or two above middle finger key (same hand)
# - side_above_1away = 0.9^3 = 0.729
# - index finger types key a row or two above middle finger key, or
# - little finger types key a row or two above ring finger key (same hand)
# - middle_above_ring = 0.9
# - middle finger types key a row or two above ring finger key (same hand)
# - ring_above_middle = 0.9^3 = 0.729
# - ring finger types key a row or two above middle finger key (same hand)
# - lateral = 0.9
# - lateral movement of (index or little) finger outside of 8 vertical columns
#
# Distance:
#
# - skip_row_3away = 0.9
# - index and little fingers type two keys that skip over home row (same hand)
# - (e.g., one on bottom row, the other on top row)
# - skip_row_2away = 0.9^3 = 0.729
# - little and middle or index and ring fingers type two keys that skip over home row (same hand)
# - skip_row_1away = 0.9^5 = 0.59049
# - little and ring or middle and index fingers type two keys that skip over home row (same hand)
#
# Repetition:
#
# - skip_row_0away = 0.9^4 = 0.6561
# - same finger types two keys that skip over home row
# - same_finger = 0.9^5 = 0.59049
# - use same finger again for a different key
# - cannot accompany outward, side_above, or adjacent_shorter_above
#
# Strength: Accounted for by the strength matrix (minimum value for the little finger = 0.9)
# -
# ### Example flow values for left side home block
#
# No penalty (for same hand, both keys in the same row in an inward roll or repeating the same key):
#
# 2=>2, 2=>3, 3=>4, 2=>4, 1=>4
#
# 1 2 3 4
# 5 6 7 8
# 9 10 11 12
#
# Penalty = 0.9:
#
# outward: 2=>1, 3=>1, 3=>2, 4=>1, 4=>2, 4=>3, 6=>5, 7=>6, 7=>5, 8=>7, 8=>6, 8=>5,...
# middle_above_ring: 6=>3, 10=>7
# side_above_3away: 1=>8, 5=>4, 5=>12, 9=>8
# index_above: 1=>4, 2=>4, 3=>4, 4=>4
#
# Penalty = 0.9^2:
#
# middle_above_ring * outward: 3=>6, 7=>10
# side_above_3away * outward: 8=>1, 4=>5, 12=>5, 8=>9
# side_above_2away: 1=>7, 6=>4, 5=>11, 10=>8
# skip_row_3away * side_above_3away: 1=>12, 9=>4
# skip_row_2away: 2=>12, 9=>3
# ring_above_middle 2=>7, 6=>11
# side_above_2away * outward: 7=>1, 4=>6, 11=>5, 8=>10
# side_above_1away: 1=>6, 7=>4, 5=>10, 11=>8
#
# Penalty = 0.9^3:
#
# skip_row_3away * side_above_3away * outward: 12=>1, 4=>9
#
# Penalty = 0.9^4:
#
# ring_above_middle * outward: 7=>2, 11=>6
# side_above_1away * outward: 4=>7, 6=>1, 10=>5, 4=>7
#
# Penalty = 0.9^5:
#
# same_finger: 4=>8, 8=>4, 1=>5, 5=>1, 5=>9, 9=>5, 2=>6, 6=>2,...
# skip_row_2away * side_above_2away: 10=>4, 1=>11
# skip_row_1away: 1=>10, 9=>2, 3=>12
#
# Penalty = 0.9^6:
#
# skip_row_2away * side_above_2away * outward: 4=>10, 11=>1
# skip_row_1away * outward: 10=>1, 2=>9, 12=>3
#
# Penalty = 0.9^8
#
# skip_row_1away * ring_above_middle: 2=>11
# skip_row_1away * side_above_1away: 1=>10, 11=>4
#
# Penalty = 0.9^9
#
# skip_row_1away * ring_above_middle * outward: 11=>2
# skip_row_0away * same_finger: 1=>9, 9=>1, 4=>12, 12=>4, 2=>10, 10=>2, 3=>11, 11=>3
# skip_row_1away * side_above_1away * outward: 10=>1, 4=>11
# +
# # %load code/load_flow_matrices.py
# Penalizing factors for 24 keys (1 = no penalty; set to less than 1 to penalize):
# Dexterity
side_above_3away = 0.9 # index and little finger type two keys, one or more rows apart (same hand)
side_above_2away = 0.81 # index finger types key a row or two above ring finger key, or
# little finger types key a row or two above middle finger key (same hand)
side_above_1away = 0.729 # index finger types key a row or two above middle finger key, or
# little finger types key a row or two above ring finger key (same hand)
middle_above_ring = 0.9 # middle finger types key a row or two above ring finger key (same hand)
ring_above_middle = 0.729 # ring finger types key a row or two above middle finger key (same hand)
lateral = 0.9 # lateral movement of (index or little) finger outside of 8 vertical columns
# Direction
outward = 0.9 # outward roll of fingers from the index to little finger (same hand)
# Distance
skip_row_3away = 0.9 # index and little fingers type two keys that skip over home row (same hand)
# (e.g., one on bottom row, the other on top row)
skip_row_2away = 0.729 # little and middle or index and ring fingers type two keys that skip over home row (same hand)
skip_row_1away = 0.59049 # little and ring or middle and index fingers type two keys that skip over home row (same hand)
# Repetition
skip_row_0away = 0.6561 # same finger types two keys that skip over home row
same_finger = 0.59049 # use same finger again for a different key
# Unused or redundant parameters
same_hand = 1.0 # (addressed by splitting up the most frequent letters across left/right sides above)
not_home_row = 1.0 # at least one key not on home row
side_top = 1.0 # index or little finger types top corner key
shorter_above = 1.0 # (taken care of by side_above_[1,2,3]away parameters)
adjacent_offset = 1.0 # (taken care of by side_above_1away, middle_above_ring, ring_above_middle parameters)
inside_top = 1.0 # index finger types top corner key (taken care of by side_above_1away parameter)
index_above = 1.0 # index finger types top corner key (unless other bigram key is in the top row for the same hand)
# (taken care of by side_above_[1,2,3]away parameters)
def create_24x24_flow_matrix(not_home_row, side_top, side_above_3away, side_above_2away, side_above_1away,
middle_above_ring, ring_above_middle, outward, skip_row_3away,
skip_row_2away, skip_row_1away, skip_row_0away, same_finger, lateral,
same_hand, shorter_above, adjacent_offset, inside_top, index_above):
all_24_keys = [1,2,3,4, 5,6,7,8, 9,10,11,12, 13,14,15,16, 17,18,19,20, 21,22,23,24]
# Create a matrix and multiply by flow factors that promote easy interkey transitions:
T = np.ones((24, 24))
# 7. Promote alternating between hands over uncomfortable transitions with the same hand.
if same_hand < 1.0:
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
for i in range(0,12):
for j in range(0,12):
T[i,j] *= same_hand
for i in range(12,24):
for j in range(12,24):
T[i,j] *= same_hand
# 8. Promote little-to-index-finger roll-ins over index-to-little-finger outwards.
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
if outward < 1.0:
# same-row roll-outs:
roll_ins = [[1,2],[2,3],[3,4], [5,6],[6,7],[7,8], [9,10],[10,11],[11,12],
[16,15],[15,14],[14,13], [20,19],[19,18],[18,17], [24,23],[23,22],[22,21]]
for x in roll_ins:
T[x[1]-1, x[0]-1] *= outward
# same-row roll-outs, skipping keys:
roll_ins_skip_keys = [[1,3],[2,4],[1,4], [5,7],[6,8],[5,8], [9,11],[10,12],[9,12],
[16,14],[15,13],[16,13], [20,18],[19,17],[20,17], [24,22],[23,21],[24,21]]
for x in roll_ins_skip_keys:
T[x[1]-1, x[0]-1] *= outward
# adjacent-row roll-outs:
roll_ins_adj_rows = [[1,6],[1,7],[1,8],[2,7],[2,8],[3,8], [5,2],[5,3],[5,4],[6,3],[6,4],[7,4],
[5,10],[5,11],[5,12],[6,11],[6,12],[7,12], [9,6],[9,7],[9,8],[10,7],[10,8],[11,8],
[16,19],[16,18],[16,17],[15,18],[15,17],[14,17], [20,15],[20,14],[20,13],[19,14],[19,13],[18,13],
[20,23],[20,22],[20,21],[19,22],[19,21],[18,21], [24,19],[24,18],[24,17],[23,18],[23,17],[22,17]]
for x in roll_ins_adj_rows:
T[x[1]-1, x[0]-1] *= outward
# upper<->lower row roll-outs:
roll_ins_skip_home = [[1,10],[1,11],[1,12],[2,11],[2,12],[3,12], [9,2],[9,3],[9,4],[10,3],[10,4],[11,4],
[16,23],[16,22],[16,21],[15,22],[15,21],[14,21], [24,15],[24,14],[24,13],[23,14],[23,13],[22,13]]
for x in roll_ins_skip_home:
T[x[1]-1, x[0]-1] *= outward
# 9. Avoid stretching shorter fingers up and longer fingers down.
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
if index_above < 1.0:
for x in [4]:
for y in [4,5,6,7,8,9,10,11,12,13,14,15,16,17,18,19,20,21,22,23,24]:
T[x-1, y-1] *= index_above
T[y-1, x-1] *= index_above
for x in [13]:
for y in [1,2,3,4,5,6,7,8,9,10,11,12,13,17,18,19,20,21,22,23,24]:
T[x-1, y-1] *= index_above
T[y-1, x-1] *= index_above
if inside_top < 1.0:
for x in [4,13]:
for j in range(0,24):
T[x-1, j] *= inside_top
T[j, x-1] *= inside_top
if side_top < 1.0:
for x in [1,4,13,16]:
for j in range(0,24):
T[x-1, j] *= side_top
T[j, x-1] *= side_top
if side_above_1away < 1.0:
for x in [1]:
for y in [6,10]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [5]:
for y in [10]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [4]:
for y in [7,11]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [8]:
for y in [11]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [13]:
for y in [18,22]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [17]:
for y in [22]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [16]:
for y in [19,23]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [20]:
for y in [23]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
if side_above_2away < 1.0:
for x in [1]:
for y in [7,11]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [5]:
for y in [11]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [4]:
for y in [6,10]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [8]:
for y in [10]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [13]:
for y in [19,23]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [17]:
for y in [23]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [16]:
for y in [18,22]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [20]:
for y in [22]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
if side_above_3away < 1.0:
for x in [1]:
for y in [8,12]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [5]:
for y in [12]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [4]:
for y in [5,9]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [8]:
for y in [9]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [13]:
for y in [20,24]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [17]:
for y in [24]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [16]:
for y in [17,21]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [20]:
for y in [21]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
if shorter_above < 1.0:
for x in [1]:
for y in [6,7,8,10,11,12]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [2]:
for y in [7,11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [4]:
for y in [6,7,10,11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [5]:
for y in [10,11,12]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [6]:
for y in [11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [8]:
for y in [10,11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [16]:
for y in [17,18,19,21,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [15]:
for y in [18,22]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [13]:
for y in [18,19,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [20]:
for y in [21,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [19]:
for y in [22]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [17]:
for y in [22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
if ring_above_middle < 1.0:
ring_above_middles = [[2,7],[6,11],[2,11],
[15,18],[19,22],[15,22]]
for x in ring_above_middles:
T[x[0]-1, x[1]-1] *= ring_above_middle
T[x[1]-1, x[0]-1] *= ring_above_middle
if middle_above_ring < 1.0:
middle_above_rings = [[6,3],[10,7],[10,3],
[19,14],[23,18],[23,14]]
for x in middle_above_rings:
T[x[0]-1, x[1]-1] *= middle_above_ring
T[x[1]-1, x[0]-1] *= middle_above_ring
# 10. Avoid using the same finger.
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
if same_finger < 1.0:
same_fingers = [[1,5],[5,9],[1,9], [2,6],[6,10],[2,10], [3,7],[7,11],[3,11], [4,8],[8,12],[4,12],
[13,17],[17,21],[13,21], [14,18],[18,22],[14,22], [15,19],[19,23],[15,23], [16,20],[20,24],[16,24]]
for x in same_fingers:
T[x[0]-1, x[1]-1] *= same_finger
T[x[1]-1, x[0]-1] *= same_finger
# 11. Avoid the upper and lower rows.
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
if not_home_row < 1.0:
not_home_row_keys = [1,2,3,4, 9,10,11,12, 13,14,15,16, 21,22,23,24]
for x in not_home_row_keys:
for j in range(0,23):
T[x-1, j] *= not_home_row
T[j, x-1] *= not_home_row
# 12. Avoid skipping over the home row.
# 1 2 3 4 13 14 15 16
# 5 6 7 8 17 18 19 20
# 9 10 11 12 21 22 23 24
if skip_row_0away < 1.0:
skip_top = [1, 2, 3, 4, 13,14,15,16]
skip_bot = [9,10,11,12, 21,22,23,24]
for ix, x in enumerate(skip_top):
y = skip_bot[ix]
T[x-1, y-1] *= skip_row_0away
T[y-1, x-1] *= skip_row_0away
if skip_row_1away < 1.0:
skip_top = [1, 2, 2, 3, 3, 4, 13,14,14,15,15,16]
skip_bot = [10,9,11,10,12,11, 22,21,23,22,24,23]
for ix, x in enumerate(skip_top):
y = skip_bot[ix]
T[x-1, y-1] *= skip_row_1away
T[y-1, x-1] *= skip_row_1away
if skip_row_2away < 1.0:
skip_top = [1, 2,3, 4, 13,14,15,16]
skip_bot = [11,12,9,10, 23,24,21,22]
for ix, x in enumerate(skip_top):
y = skip_bot[ix]
T[x-1, y-1] *= skip_row_2away
T[y-1, x-1] *= skip_row_2away
if skip_row_3away < 1.0:
skip_top = [1, 4, 13,16]
skip_bot = [12,9, 24,21]
for ix, x in enumerate(skip_top):
y = skip_bot[ix]
T[x-1, y-1] *= skip_row_3away
T[y-1, x-1] *= skip_row_3away
Flow24x24 = T
# Normalize matrix with min-max scaling to a range with maximum = 1:
newMin = np.min(Flow24x24) / np.max(Flow24x24)
newMax = 1.0
Flow24x24 = newMin + (Flow24x24 - np.min(Flow24x24)) * (newMax - newMin) / (np.max(Flow24x24) - np.min(Flow24x24))
return Flow24x24
Flow24x24 = create_24x24_flow_matrix(not_home_row, side_top,
side_above_3away, side_above_2away, side_above_1away, middle_above_ring, ring_above_middle, outward,
skip_row_3away, skip_row_2away, skip_row_1away, skip_row_0away, same_finger, lateral, same_hand,
shorter_above, adjacent_offset, inside_top, index_above)
# Print:
print_matrix_info(matrix_data=Flow24x24, matrix_label="Flow24x24", nkeys=24, nlines=30)
heatmap(data=Flow24x24, title="Flow24x24", xlabel="Key 1", ylabel="Key 2", print_output=print_output)
def create_32x32_flow_matrix(not_home_row, side_top, side_above_3away, side_above_2away, side_above_1away,
middle_above_ring, ring_above_middle, outward, skip_row_3away,
skip_row_2away, skip_row_1away, skip_row_0away, same_finger, lateral,
same_hand, shorter_above, adjacent_offset, inside_top, index_above):
all_32_keys = [1,2,3,4, 5,6,7,8, 9,10,11,12, 13,14,15,16, 17,18,19,20, 21,22,23,24,
25,26,27, 28,29,30, 31,32]
# Create a matrix and multiply by flow factors that promote easy interkey transitions:
T = np.ones((32, 32))
if lateral < 1.0:
for x in all_32_keys:
for y in [25,26,27, 28,29,30, 31,32]:
T[x-1, y-1] *= lateral
T[y-1, x-1] *= lateral
# 7. Promote alternating between hands over uncomfortable transitions with the same hand.
if same_hand < 1.0:
for i in [1,2,3,4,5,6,7,8,9,10,11,12, 25,26,27]:
for j in [1,2,3,4,5,6,7,8,9,10,11,12, 25,26,27]:
T[i-1,j-1] *= same_hand
for i in [13,14,15,16,17,18,19,20,21,22,23,24, 28,29,30,31,32]:
for j in [13,14,15,16,17,18,19,20,21,22,23,24, 28,29,30,31,32]:
T[i-1,j-1] *= same_hand
# 8. Promote little-to-index-finger roll-ins over index-to-little-finger outsward rolls.
# Penalize (index, little) finger lateral movements:
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
if outward < 1.0:
# same-row roll-outs:
roll_ins = [[1,2],[2,3],[3,4], [5,6],[6,7],[7,8], [9,10],[10,11],[11,12],
[16,15],[15,14],[14,13], [20,19],[19,18],[18,17], [24,23],[23,22],[22,21]]
for x in roll_ins:
T[x[1]-1, x[0]-1] *= outward
# same-row roll-outs, skipping keys:
roll_ins_skip_keys = [[1,3],[2,4],[1,4], [5,7],[6,8],[5,8], [9,11],[10,12],[9,12],
[16,14],[15,13],[16,13], [20,18],[19,17],[20,17], [24,22],[23,21],[24,21]]
#[1,25],[2,25],[3,25],
#[5,26],[6,26],[7,26],
#[9,27],[10,27],[11,27],
#[16,28],[15,28],[14,28],
#[20,29],[19,29],[18,29],
#[24,30],[23,30],[22,30],
#[31,15],[31,14],[31,13],[31,28],
#[32,19],[32,18],[32,17],[32,29]]
for x in roll_ins_skip_keys:
T[x[1]-1, x[0]-1] *= outward
# adjacent-row roll-outs:
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
roll_ins_adj_rows = [[1,6],[1,7],[1,8],[2,7],[2,8],[3,8],
[5,2],[5,3],[5,4],[6,3],[6,4],[7,4],
[5,10],[5,11],[5,12],[6,11],[6,12],[7,12],
[9,6],[9,7],[9,8],[10,7],[10,8],[11,8],
[16,19],[16,18],[16,17],[15,18],[15,17],[14,17],
[20,15],[20,14],[20,13],[19,14],[19,13],[18,13],
[20,23],[20,22],[20,21],[19,22],[19,21],[18,21],
[24,19],[24,18],[24,17],[23,18],[23,17],[22,17]]
#[5,25],[6,25],[7,25],[8,25],
#[5,27],[6,27],[7,27],[8,27],
#[1,26],[2,26],[3,26],[4,26],
#[9,26],[10,26],[11,26],[12,26],
#[16,29],[15,29],[14,29],[13,29],
#[24,29],[23,29],[22,29],[21,29],
#[20,28],[19,28],[18,28],[17,28],
#[20,30],[19,30],[18,30],[17,30],
#[31,20],[31,19],[31,18],[31,17],[31,29],
#[32,16],[32,15],[32,14],[32,13],[32,28],
#[32,24],[32,23],[32,22],[32,21],[32,30]]
for x in roll_ins_adj_rows:
T[x[1]-1, x[0]-1] *= outward
# upper<->lower row roll-outs:
roll_ins_skip_home = [[1,10],[1,11],[1,12],[2,11],[2,12],[3,12],
[9,2],[9,3],[9,4],[10,3],[10,4],[11,4],
[16,23],[16,22],[16,21],[15,22],[15,21],[14,21],
[24,15],[24,14],[24,13],[23,14],[23,13],[22,13]]
#[16,30],[15,30],[14,30],[13,30],
#[9,25],[10,25],[11,25],[12,25],
#[24,28],[23,28],[22,28],[21,28],
#[1,27],[2,27],[3,27],[4,27],
#[31,24],[31,23],[31,22],[31,21],[31,30]]
for x in roll_ins_skip_home:
T[x[1]-1, x[0]-1] *= outward
# 9. Avoid stretching shorter fingers up and longer fingers down.
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
if index_above < 1.0:
for x in [4]:
for y in [4,5,6,7,8,26,9,10,11,12,27,28,13,14,15,16,31,29,17,18,19,20,32,30,21,22,23,24]:
T[x-1, y-1] *= index_above
T[y-1, x-1] *= index_above
for x in [25]:
for y in [25,5,6,7,8,26,9,10,11,12,27,28,13,14,15,16,31,29,17,18,19,20,32,30,21,22,23,24]:
T[x-1, y-1] *= index_above
T[y-1, x-1] *= index_above
for x in [13]:
for y in [1,2,3,4,25,5,6,7,8,26,9,10,11,12,27,13,29,17,18,19,20,32,30,21,22,23,24]:
T[x-1, y-1] *= index_above
T[y-1, x-1] *= index_above
for x in [28]:
for y in [1,2,3,4,25,5,6,7,8,26,9,10,11,12,27,28,29,17,18,19,20,32,30,21,22,23,24]:
T[x-1, y-1] *= index_above
T[y-1, x-1] *= index_above
if inside_top < 1.0:
for x in [4,25,28,13]:
for j in range(0,32):
T[x-1, j] *= inside_top
T[j, x-1] *= inside_top
if side_top < 1.0:
for x in [1,4,25,28,13,16,31]:
for j in range(0,32):
T[x-1, j] *= side_top
T[j, x-1] *= side_top
if side_above_1away < 1.0:
for x in [1]:
for y in [6,10]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [5]:
for y in [10]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [4,25]:
for y in [7,11]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [8,26]:
for y in [11]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [13,28]:
for y in [18,22]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [17,29]:
for y in [22]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [16,31]:
for y in [19,23]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
for x in [20,32]:
for y in [23]:
T[x-1, y-1] *= side_above_1away
T[y-1, x-1] *= side_above_1away
if side_above_2away < 1.0:
for x in [1]:
for y in [7,11]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [5]:
for y in [11]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [4,25]:
for y in [6,10]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [8,26]:
for y in [10]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [13,28]:
for y in [19,23]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [17,29]:
for y in [23]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [16,31]:
for y in [18,22]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
for x in [20,32]:
for y in [22]:
T[x-1, y-1] *= side_above_2away
T[y-1, x-1] *= side_above_2away
if side_above_3away < 1.0:
for x in [1]:
for y in [8,12,26,27]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [5]:
for y in [12,27]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [4,25]:
for y in [5,9]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [8,26]:
for y in [9]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [13,28]:
for y in [20,24,32]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [17,29]:
for y in [24]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [16,31]:
for y in [17,21,29,30]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
for x in [20,32]:
for y in [21,30]:
T[x-1, y-1] *= side_above_3away
T[y-1, x-1] *= side_above_3away
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
if shorter_above < 1.0:
for x in [1]:
for y in [6,7,8,26,10,11,12,27]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [2]:
for y in [7,11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [4]:
for y in [6,7,10,11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [25]:
for y in [6,7,10,11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [5]:
for y in [10,11,12,27]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [6]:
for y in [11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [8]:
for y in [10,11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [26]:
for y in [10,11]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [16]:
for y in [29,17,18,19,30,21,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [31]:
for y in [29,17,18,19,30,21,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [15]:
for y in [18,22]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [13]:
for y in [18,19,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [28]:
for y in [18,19,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [20]:
for y in [30,21,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [32]:
for y in [30,21,22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [19]:
for y in [22]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [17]:
for y in [22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
for x in [29]:
for y in [22,23]:
T[x-1, y-1] *= shorter_above
T[y-1, x-1] *= shorter_above
if ring_above_middle < 1.0:
ring_above_middles = [[2,7],[6,11],[2,11],
[15,18],[19,22],[15,22]]
for x in ring_above_middles:
T[x[0]-1, x[1]-1] *= ring_above_middle
T[x[1]-1, x[0]-1] *= ring_above_middle
if middle_above_ring < 1.0:
middle_above_rings = [[6,3],[10,7],[10,3],
[19,14],[23,18],[23,14]]
for x in middle_above_rings:
T[x[0]-1, x[1]-1] *= middle_above_ring
T[x[1]-1, x[0]-1] *= middle_above_ring
# 10. Avoid using the same finger.
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
if same_finger < 1.0:
same_fingers = [[1,5],[5,9],[1,9], [2,6],[6,10],[2,10],
[3,7],[7,11],[3,11], [4,8],[8,12],[4,12],
[25,26],[26,27],[25,27], [28,29],[29,30],[28,30], [31,32],
[4,25],[4,26],[4,27], [8,25],[8,26],[8,27], [12,25],[12,26],[12,27],
[13,28],[13,29],[13,30], [17,28],[17,29],[17,30], [21,28],[21,29],[21,30],
[31,16],[31,20],[31,24], [32,16],[32,20],[32,24],
[13,17],[17,21],[13,21], [14,18],[18,22],[14,22],
[15,19],[19,23],[15,23], [16,20],[20,24],[16,24]]
for x in same_fingers:
T[x[0]-1, x[1]-1] *= same_finger
T[x[1]-1, x[0]-1] *= same_finger
# 11. Avoid the upper and lower rows.
if not_home_row < 1.0:
not_home_row_keys = [1,2,3,4,25, 9,10,11,12,27, 28,13,14,15,16,31, 30,21,22,23,24]
for x in not_home_row_keys:
for j in range(0,32):
T[x-1, j] *= not_home_row
T[j, x-1] *= not_home_row
# 12. Avoid skipping over the home row.
# 1 2 3 4 25 28 13 14 15 16 31
# 5 6 7 8 26 29 17 18 19 20 32
# 9 10 11 12 27 30 21 22 23 24
if skip_row_0away < 1.0:
skip_top = [1, 2, 3, 4, 4,25,25, 28,28,13,13,14,15,16,31]
skip_bot = [9,10,11,12,27,12,27, 30,21,30,21,22,23,24,24]
for ix, x in enumerate(skip_top):
y = skip_bot[ix]
T[x-1, y-1] *= skip_row_0away
T[y-1, x-1] *= skip_row_0away
if skip_row_1away < 1.0:
skip_top = [1, 2, 2, 3, 3, 4, 4,25, 28,13,13,14,14,15,15,16,31]
skip_bot = [10,9,11,10,12,11,27,11, 22,30,22,21,23,22,24,23,23]
for ix, x in enumerate(skip_top):
y = skip_bot[ix]
T[x-1, y-1] *= skip_row_1away
T[y-1, x-1] *= skip_row_1away
if skip_row_2away < 1.0:
skip_top = [1, 2,3, 4,25, 28,13,14,15,16,31]
skip_bot = [11,12,9,10,10, 23,23,24,21,22,22]
for ix, x in enumerate(skip_top):
y = skip_bot[ix]
T[x-1, y-1] *= skip_row_2away
T[y-1, x-1] *= skip_row_2away
if skip_row_3away < 1.0:
skip_top = [1, 4,25, 28,13,16,16,31,31]
skip_bot = [12,9, 9, 24,24,21,30,21,30]
for ix, x in enumerate(skip_top):
y = skip_bot[ix]
T[x-1, y-1] *= skip_row_3away
T[y-1, x-1] *= skip_row_3away
Flow32x32 = T
# Normalize matrix with min-max scaling to a range with maximum = 1:
newMin = np.min(Flow32x32) / np.max(Flow32x32)
newMax = 1.0
Flow32x32 = newMin + (Flow32x32 - np.min(Flow32x32)) * (newMax - newMin) / (np.max(Flow32x32) - np.min(Flow32x32))
return Flow32x32
Flow32x32 = create_32x32_flow_matrix(not_home_row, side_top,
side_above_3away, side_above_2away, side_above_1away, middle_above_ring, ring_above_middle, outward,
skip_row_3away, skip_row_2away, skip_row_1away, skip_row_0away, same_finger, lateral, same_hand,
shorter_above, adjacent_offset, inside_top, index_above)
# Print:
print_matrix_info(matrix_data=Flow32x32, matrix_label="Flow32x32", nkeys=32, nlines=30)
heatmap(data=Flow32x32, title="Flow32x32", xlabel="Key 1", ylabel="Key 2", print_output=print_output)
# + [markdown] colab_type="text" id="WMvP493uDzSU"
# ## Combine Strength and Flow matrices <a name="strengthflow">
# + colab={"base_uri": "https://localhost:8080/", "height": 695} colab_type="code" id="UP7FUBR2DzSX" outputId="5dc11788-2c69-4f69-ab60-a07ac17e092f"
# # %load code/combine_scoring_matrices.py
# 24 keys:
Factors24x24 = Flow24x24
if apply_strength:
Factors24x24 = Strength24x24 * Factors24x24
# Print:
print_matrix_info(matrix_data=Factors24x24, matrix_label="Factors24x24", nkeys=24, nlines=30)
heatmap(data=Factors24x24, title="Factors24x24", xlabel="Key 1", ylabel="Key 2", print_output=print_output)
# Save:
if print_output:
file = open("Factors24x24.txt", "w+")
file.write(str(Factors24x24))
file.close()
# 32 keys:
Factors32x32 = Flow32x32
if apply_strength:
Factors32x32 = Strength32x32 * Factors32x32
# Print:
print_matrix_info(matrix_data=Factors32x32, matrix_label="Factors32x32", nkeys=32, nlines=30)
heatmap(data=Factors32x32, title="Factors32x32", xlabel="Key 1", ylabel="Key 2", print_output=print_output)
# Save:
if print_output:
file = open("Factors32x32.txt", "w+")
file.write(str(Factors32x32))
file.close()
# -
# ## Four steps
#
# We will assign letters to keys by choosing the arrangement with the highest score according to our scoring model. However, there are over four hundred septillion, or four hundred trillion trillion (26! = 403,291,461,126,605,635,584,000,000, or 4.032914611 E+26) possible arrangements of 26 letters (24! = 6.204484017 E+23), so we will arrange the letters in four steps, based on ergonomics principles. These consist of (Step 1) assigning the eight most frequent letters to different keys, optimizing assignment of the remaining (Step 2) eight most frequent letters, and (Step 3) eight least frequent letters (besides Z and Q), and (Step 4) exchanging letters.
#
# ## Step 1: Define the shape of the key layout to minimize lateral finger movements<a name="step1">
#
# We will assign 24 letters to 8 columns of keys separated by two middle columns reserved for punctuation. These 8 columns require no lateral finger movements when touch typing, since there is one column per finger. The most comfortable keys include the left and right home rows (keys 5-8 and 17-20), the top-center keys (2,3 and 14,15) that allow the longer middle and ring fingers to uncurl upwards, as well as the bottom corner keys (9,12 and 21,24) that allow the shorter fingers to curl downwards. We will assign the two least frequent letters, Z and Q (or J), to the two hardest-to-reach keys lying outside the 24-key columns in the upper right (25 and 26):
#
# Left: Right:
# 1 2 3 4 13 14 15 16 25
# 5 6 7 8 17 18 19 20 26
# 9 10 11 12 21 22 23 24
#
# We will consider the most comfortable keys to be those typed by either hand on the home row, by the ring and middle finger above the home row, and by the index and little finger below the home row, with a preference for the strongest (index and middle) fingers:
#
# - 2 3 - - 14 15 -
# 5 6 7 8 17 18 19 20
# 9 - - 12 21 - - 24
# + [markdown] colab_type="text" id="REInHU9tdYLP"
# ## Step 2: Arrange the most frequent letters based on comfort and bigram frequencies <a name="step2">
#
# In prior experiments using the methods below, all vowels consistently automatically clustered together. Below, we will arrange vowels on one side and the most frequent consonants to the other side to encourage balance and alternation across hands. Since aside from the letters Z and Q there is symmetry across left and right sides, we will decide later which side the vowels and which side the most frequent consonants should go.
#
# ### Vowels
#
# **E**, T, **A, O, I**, N, S, R, H, L, D, C, U, M, F, P, G, W, Y, B, V, K, X, J, Q, Z
#
# The highest frequency bigrams that contain two vowels are listed below in bold, with more than 10 billion instances in <NAME>vig's analysis of Google data:
#
# **OU, IO, EA, IE**, AI, IA, EI, UE, UA, AU, UI, OI, EO, OA, OE
#
# OU 24,531,132,241
# IO 23,542,263,265
# EA 19,403,941,063
# IE 10,845,731,320
# AI 8,922,759,715
# IA 8,072,199,471
# EI 5,169,898,489
# UE 4,158,448,570
# UA 3,844,138,094
# AU 3,356,322,923
# UI 2,852,182,384
# OI 2,474,275,212
# EO 2,044,268,477
# OA 1,620,913,259
# OE 1,089,254,517
#
# We will assign the most frequent vowels with over 100 billion instances in Norvig's analysis (E=445,A=331,O=272,I=270) to four of the six most comfortable keys on the left side of the keyboard (keys 2,3,5,6,7,8). We will assign the letter E, the most frequent in the English language, to either of the strongest (index and middle) fingers on the home row, and assign the other three vowels such that (1) the home row keys typed by the index and middle fingers are not left vacant, and any top-frequency bigram (more than 10 billion instances in Norvig's analysis) (2) does not use the same finger and (3) reads from left to right (ex: EA, not AE) for ease of typing (inward roll from little to index finger vs. outward roll from index to little finger). These constraints lead to three arrangements of the four vowels:
#
# - - O - - - O - - - - -
# - I E A I - E A I O E A
# - - - - - - - - - - - -
#
# ### Consonants
#
# On the right side of the keyboard, we will assign four of the five most frequent consonants (with over 5% or 150 billion instances in Norvig's analysis: T=331, N=258, S=232, R=224, and H=180) to the four home row keys. We will assign the letter T, the most frequent consonant in the English language, to either of the strongest (index and middle) fingers on the home row. As with the left side, letters are placed so that top-frequency bigrams read from right to left (ex: HT, not TH) for ease of typing. The top-frequency bigrams (more than 10 billion instances in Norvig's analysis) include: TH, ND, ST, NT, CH, NS, CT, TR, RS, NC, and RT (below 10 billion instances these bigrams start to occur in reverse, such as RT and TS):
#
# TH 100,272,945,963 3.56%
# ND 38,129,777,631 1.35%
# ST 29,704,461,829 1.05%
# NT 29,359,771,944 1.04%
# CH 16,854,985,236 0.60%
# NS 14,350,320,288
# CT 12,997,849,406
# TR 12,006,693,396
# RS 11,180,732,354
# NC 11,722,631,112
# RT 10,198,055,461
#
# The above constraints lead to five arrangements of the consonants:
#
# - - - - - - - - - - - - - - - - - - - -
# R T S N H T S N H T S R H T N R T S N R
# - - - - - - - - - - - - - - - - - - - -
#
# We will assign the fifth consonant to a vacant key on the left home row if there is a vacancy, otherwise to the key below the right index finger (any other assignment requires the same finger to type a high-frequency bigram). The resulting 19 initial layouts, each with 15 unassigned keys, are represented below with the three rows on the left and right side of the keyboard as a linear string of letters, with unassigned keys denoted by “-”.
#
# --O- HIEA ---- ---- RTSN ----
# --O- RIEA ---- ---- HTSN ----
# --O- NIEA ---- ---- HTSR ----
# --O- SIEA ---- ---- HTNR ----
# --O- IHEA ---- ---- RTSN ----
# --O- IREA ---- ---- HTSN ----
# --O- INEA ---- ---- HTSR ----
# --O- ISEA ---- ---- HTNR ----
# --O- -IEA ---- ---- RTSN H---
# --O- -IEA ---- ---- HTSN R---
# --O- -IEA ---- ---- HTSR N---
# --O- I-EA ---- ---- RTSN H---
# --O- I-EA ---- ---- HTSN R---
# --O- I-EA ---- ---- HTSR N---
# ---- IOEA ---- ---- RTSN H---
# ---- IOEA ---- ---- HTSN R---
# ---- IOEA ---- ---- HTSR N---
# --O- HIEA ---- ---- TSNR ----
# --O- IHEA ---- ---- TSNR ----
# -
# ## Step 3: Optimize assignment of the remaining letters <a name="step3">
#
# We want to assign letters to the 17 unassigned keys in each of the above 19 layouts based on our scoring model. That would mean scoring all possible arrangements for each layout and choosing the arrangement with the highest score, but since there are over 355 trillion (17!) possible ways of arranging 17 letters, we will break up the assignment into two stages for the most frequent and least frequent remaining letters.
#
# ### Most frequent letters
# We will compute scores for every possible arrangement of the seven most frequent of the remaining letters (in bold below) assigned to vacancies among the most comfortable sixteen keys.
#
# E, T, A, O, I, N, S, R, H, **L, D, C, U, M, F, P**, G, W, Y, B, V, K, X, J, Q, Z
#
# Left: Right:
# - 2 3 - - 14 15 -
# 5 6 7 8 17 18 19 20
# 9 - - 12 21 - - 24
#
# Since there are 5,040 (7!) possible combinations of eight letters for each of the 19 layouts, we need to score and evaluate 95,760 layouts. To score each arrangement of letters, we construct a frequency matrix where we multiply a matrix containing the frequency of each ordered pair of letters (bigram) by our flow and strength matrices to compute a score.
#
# ### Least frequent letters
# Next we will compute scores for every possible (40,320 = 8!) arrangement of the least frequent eight letters (in bold below, besides Z and Q) in the remaining keys, after substituting in the 19 results of the above for an additional 766,080 layouts:
#
# E, T, A, O, I, N, S, R, H, L, D, C, U, M, F, P, **G, W, Y, B, V, K, X, J**, Q, Z
#
# Left: Right:
# 1 - - 4 13 - - 16
# - - - - - - - -
# - 10 11 - - 22 23 -
#
# ### Further optimize layouts by exchanging more letters
#
# If we relax the above fixed initializations and permit further exchange of letters, then we can search for even higher-scoring layouts. As a final optimization step we exchange letters, eight keys at a time (8! = 40,320) selected twice in 14 different ways, in each of the above 19 layouts, to score a total of 21,450,240 more combinations. We allow the following keys to exchange letters:
#
# 1. Top rows
# 2. Bottom rows
# 3. Top and bottom rows on the right side
# 4. Top and bottom rows on the left side
# 5. Top right and bottom left rows
# 6. Top left and bottom right rows
# 7. Center of the top and bottom rows on both sides
# 8. The eight corners
# 9. Left half of the top and bottom rows on both sides
# 10. Right half of the top and bottom rows on both sides
# 11. Left half of non-home rows on the left and right half of the same rows on the right
# 12. Right half of non-home rows on the left and left half of the same rows on the right
# 13. Top center and lower sides
# 14. Top sides and lower center
# 15. Repeat 1-14
# +
"""
NOTE: This procedure takes hours to run.
--O- HIEA ---- ---- RTSN ----
--O- RIEA ---- ---- HTSN ----
--O- NIEA ---- ---- HTSR ----
--O- SIEA ---- ---- HTNR ----
--O- IHEA ---- ---- RTSN ----
--O- IREA ---- ---- HTSN ----
--O- INEA ---- ---- HTSR ----
--O- ISEA ---- ---- HTNR ----
--O- -IEA ---- ---- RTSN H---
--O- -IEA ---- ---- HTSN R---
--O- -IEA ---- ---- HTSR N---
--O- I-EA ---- ---- RTSN H---
--O- I-EA ---- ---- HTSN R---
--O- I-EA ---- ---- HTSR N---
---- IOEA ---- ---- RTSN H---
---- IOEA ---- ---- HTSN R---
---- IOEA ---- ---- HTSR N---
--O- HIEA ---- ---- TSNR ----
--O- IHEA ---- ---- TSNR ----
"""
fixed_letter_lists1 = [
['O','H','I','E','A','R','T','S','N'],
['O','R','I','E','A','H','T','S','N'],
['O','N','I','E','A','H','T','S','R'],
['O','S','I','E','A','H','T','N','R'],
['O','I','H','E','A','R','T','S','N'],
['O','I','R','E','A','H','T','S','N'],
['O','I','N','E','A','H','T','S','R'],
['O','I','S','E','A','H','T','N','R'],
['O','I','E','A','R','T','S','N','H'],
['O','I','E','A','H','T','S','N','R'],
['O','I','E','A','H','T','S','R','N'],
['O','I','E','A','R','T','S','N','H'],
['O','I','E','A','H','T','S','N','R'],
['O','I','E','A','H','T','S','R','N'],
['I','O','E','A','R','T','S','N','H'],
['I','O','E','A','H','T','S','N','R'],
['I','O','E','A','H','T','S','R','N'],
['O','H','I','E','A','T','S','N','R'],
['O','I','H','E','A','T','S','N','R']]
# Keys for step 1:
# - 2 3 - - 14 15 -
# 5 6 7 8 17 18 19 20
# 9 - - 12 21 - - 24
keys1 = [2,3, 5,6,7,8, 9,12, 14,15, 17,18,19,20, 21,24]
# Indices for step 1:
# - 0 1 - - 8 9 -
# 2 3 4 5 10 11 12 13
# 6 - - 7 14 - - 15
fixed_letter_index_lists1 = [[1, 2,3,4,5, 10,11,12,13],
[1, 2,3,4,5, 10,11,12,13],
[1, 2,3,4,5, 10,11,12,13],
[1, 2,3,4,5, 10,11,12,13],
[1, 2,3,4,5, 10,11,12,13],
[1, 2,3,4,5, 10,11,12,13],
[1, 2,3,4,5, 10,11,12,13],
[1, 2,3,4,5, 10,11,12,13],
[1, 3,4,5, 10,11,12,13, 14],
[1, 3,4,5, 10,11,12,13, 14],
[1, 3,4,5, 10,11,12,13, 14],
[1, 2, 4,5, 10,11,12,13, 14],
[1, 2, 4,5, 10,11,12,13, 14],
[1, 2, 4,5, 10,11,12,13, 14],
[ 2,3,4,5, 10,11,12,13, 14],
[ 2,3,4,5, 10,11,12,13, 14],
[ 2,3,4,5, 10,11,12,13, 14],
[1, 2,3,4,5, 10,11,12,13],
[1, 2,3,4,5, 10,11,12,13]]
open_letter_index_lists1 = [[0, 6,7, 8,9, 14,15],
[0, 6,7, 8,9, 14,15],
[0, 6,7, 8,9, 14,15],
[0, 6,7, 8,9, 14,15],
[0, 6,7, 8,9, 14,15],
[0, 6,7, 8,9, 14,15],
[0, 6,7, 8,9, 14,15],
[0, 6,7, 8,9, 14,15],
[0, 2, 6,7, 8,9, 15],
[0, 2, 6,7, 8,9, 15],
[0, 2, 6,7, 8,9, 15],
[0, 3, 6,7, 8,9, 15],
[0, 3, 6,7, 8,9, 15],
[0, 3, 6,7, 8,9, 15],
[0,1, 6,7, 8,9, 15],
[0,1, 6,7, 8,9, 15],
[0,1, 6,7, 8,9, 15],
[0, 6,7, 8,9, 14,15],
[0, 6,7, 8,9, 14,15]]
# All 24 key indices:
# 0 1 2 3 12 13 14 15
# 4 5 6 7 16 17 18 19
# 8 9 10 11 20 21 22 23
# Open indices:
# 0 - - 3 12 - - 15
# - - - - - - - -
# - 9 10 - - 21 22 -
fixed_letter_indices2 = [1,2, 4,5,6,7, 8,11, 13,14, 16,17,18,19, 20,23]
open_letter_indices2 = [0,3, 9,10, 12,15, 21,22]
fixed_letter_index_lists3 = [[2, 4,5,6,7, 16,17,18,19],
[2, 4,5,6,7, 16,17,18,19],
[2, 4,5,6,7, 16,17,18,19],
[2, 4,5,6,7, 16,17,18,19],
[2, 4,5,6,7, 16,17,18,19],
[2, 4,5,6,7, 16,17,18,19],
[2, 4,5,6,7, 16,17,18,19],
[2, 4,5,6,7, 16,17,18,19],
[2, 5,6,7, 16,17,18,19, 20],
[2, 5,6,7, 16,17,18,19, 20],
[2, 5,6,7, 16,17,18,19, 20],
[2, 4, 6,7, 16,17,18,19, 20],
[2, 4, 6,7, 16,17,18,19, 20],
[2, 4, 6,7, 16,17,18,19, 20],
[ 4,5,6,7, 16,17,18,19, 20],
[ 4,5,6,7, 16,17,18,19, 20],
[ 4,5,6,7, 16,17,18,19, 20],
[2, 4,5,6,7, 16,17,18,19],
[2, 4,5,6,7, 16,17,18,19]]
# Loop through initialized layouts with assigned vowels and consonants
top_layouts = []
nlists = len(fixed_letter_lists1)
for ilist, fixed_letters1 in enumerate(fixed_letter_lists1):
fixed_letter_indices1 = fixed_letter_index_lists1[ilist]
fixed_letter_indices3 = fixed_letter_index_lists3[ilist]
open_letter_indices1 = open_letter_index_lists1[ilist]
print('Layout {0}'.format(ilist+1))
print(*fixed_letters1)
print("Most frequent letters")
top_permutation1, top_score1, letter_permutations1 = permute_optimize_keys(fixed_letters1, fixed_letter_indices1,
open_letter_indices1, letters24, keys1, Factors24x24,
bigrams, bigram_frequencies, min_score=0, verbose=False)
fixed_letters2 = top_permutation1
print("Least frequent remaining letters")
top_permutation2, top_score2, letter_permutations2 = permute_optimize_keys(fixed_letters2, fixed_letter_indices2,
open_letter_indices2, letters24, keys24, Factors24x24,
bigrams, bigram_frequencies, min_score=0, verbose=False)
fixed_letters3 = top_permutation2
print("Further optimize layouts by exchanging sets of letters")
top_permutation3, top_score3 = exchange_letters(fixed_letters3, fixed_letter_indices3, letters24, keys24,
Factors24x24, bigrams, bigram_frequencies, verbose=True)
top_layouts.append(top_permutation3)
# -
#
#
#
# #### Optimized layouts (outcome of above)
load_top_layouts = True
print_layouts = False
if load_top_layouts:
top_layouts = [
['B','Y','O','U','H','I','E','A','V','K','J','X','L','D','G','F','R','T','S','N','C','M','W','P'],
['W','Y','O','U','R','I','E','A','G','X','J','K','L','D','C','B','H','T','S','N','M','F','V','P'],
['J','P','O','U','N','I','E','A','B','K','Y','X','M','C','G','V','H','T','S','R','L','D','F','W'],
['J','P','O','U','S','I','E','A','G','K','Y','X','M','C','W','V','H','T','N','R','D','L','F','B'],
['J','P','O','U','I','H','E','A','B','K','Y','X','L','D','G','F','R','T','S','N','C','M','V','W'],
['J','W','O','U','I','R','E','A','G','X','K','Y','L','D','C','B','H','T','S','N','M','F','V','P'],
['J','P','O','U','I','N','E','A','B','X','K','Y','M','C','G','V','H','T','S','R','L','D','F','W'],
['J','G','O','U','I','S','E','A','P','X','K','Y','M','C','W','V','H','T','N','R','D','L','F','B'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','F','B','R','T','S','N','H','M','V','W'],
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','F','P'],
['P','Y','O','U','C','I','E','A','G','K','J','X','M','L','F','B','H','T','S','R','N','D','V','W'],
['J','G','O','U','I','C','E','A','B','X','Y','K','L','D','F','V','R','T','S','N','H','M','W','P'],
['J','G','O','U','I','C','E','A','B','X','Y','K','L','D','W','V','H','T','S','N','R','M','F','P'],
['P','G','O','U','I','C','E','A','K','X','J','Y','M','L','F','B','H','T','S','R','N','D','V','W'],
['J','G','U','K','I','O','E','A','P','X','Y','F','L','D','V','B','R','T','S','N','H','M','C','W'],
['J','G','U','X','I','O','E','A','W','K','Y','F','L','D','C','B','H','T','S','N','R','M','V','P'],
['J','G','U','K','I','O','E','A','P','X','Y','F','M','L','B','W','H','T','S','R','N','D','V','C'],
['J','P','O','U','H','I','E','A','G','K','Y','X','M','C','F','V','T','S','N','R','D','L','B','W'],
['J','P','O','U','I','H','E','A','G','K','Y','X','M','C','F','V','T','S','N','R','D','L','B','W']]
if print_layouts:
print('Layouts:\n')
for layout in top_layouts:
print(layout)
# ### Rank optimized layouts
# +
# # %load code/rank_layouts.py
layout_strings = []
scores = []
for layout in top_layouts:
layout_string = ' '.join(layout)
score = score_layout(Factors24x24, layout, bigrams, bigram_frequencies, verbose=False)
#print(' {0} {1}'.format(layout_string, score))
layout_strings.append(layout_string)
scores.append(score)
# Establish which layouts are within a small difference of the top-scoring layout
scores_sorted, ranks_sorted, Isort = rank_within_epsilon(scores, factor24, factor=True, verbose=False)
layouts_sorted = []
layout_strings_sorted = []
for i in Isort:
layouts_sorted.append(top_layouts[i])
layout_strings_sorted.append(layout_strings[i])
print('\n (#) Rank Score')
for i, rank in enumerate(ranks_sorted):
print(' ({0}) {1}: {2} {3}'.format(i+1, rank, layout_strings_sorted[i], scores_sorted[i]))
print('\nLayouts tied for first place, with relative letter frequencies:\n')
#print(' Rank Score')
first_ranks = []
first_layouts = []
first_layout_strings = []
first_scores = []
for i, rank in enumerate(ranks_sorted):
if rank == 1:
first_ranks.append(rank)
first_layouts.append(layout_strings_sorted[i])
first_layout_strings.append(layouts_sorted[i])
first_scores.append(scores_sorted[i])
Isort2 = np.argsort([-x for x in first_scores])
first_ranks_sorted = []
first_layouts_sorted = []
first_layout_strings_sorted = []
first_scores_sorted = []
for i in Isort2:
first_ranks_sorted.append(first_ranks[i])
first_layouts_sorted.append(first_layouts[i])
first_layout_strings_sorted.append(first_layout_strings[i])
first_scores_sorted.append(first_scores[i])
#for i, first_layout in enumerate(first_layouts):
# print(' {0}: {1} {2}'.format(first_ranks_sorted[i],
# first_layout, # first_layout_strings_sorted[i],
# first_scores_sorted[i]))
# Print layouts:
for i, layout_string in enumerate(first_layout_strings_sorted):
layout = first_layouts_sorted[i]
print(' Layout {0}:\n'.format(Isort2[i] + 1))
print_layout24(layout_string)
print('')
print_layout24_instances(layout_string, letters24, instances24, bigrams, bigram_frequencies)
print('')
# -
# #### Ranked, optimized layouts
#
# We will select the second layout tied for first place as our candidate winner, so that the most frequent bigram (TH, over 100 billion) is on the home row and easier to type.
#
# Rank Score
# 1: P Y O U C I E A G K J X L D F B R T S N H M V W 0.7079134589554652
# 1: B Y O U C I E A G X J K L D W V H T S N R M F P 0.7078676989043136
# 2: J G O U I C E A B X Y K L D F V R T S N H M W P 0.7078208372363046
# 2: B Y O U H I E A V K J X L D G F R T S N C M W P 0.7078164910125013
# 2: J P O U H I E A G K Y X M C F V T S N R D L B W 0.707806617890607
# 2: J G O U I C E A B X Y K L D W V H T S N R M F P 0.7077802597858632
# 3: P Y O U C I E A G K J X M L F B H T S R N D V W 0.707765513186795
# 3: J P O U I H E A G K Y X M C F V T S N R D L B W 0.7077455939244159
# 3: J P O U I H E A B K Y X L D G F R T S N C M V W 0.7077426951024633
# 4: P G O U I C E A K X J Y M L F B H T S R N D V W 0.7076779754232723
# 5: J P O U S I E A G K Y X M C W V H T N R D L F B 0.707608035505442
# 5: J G U K I O E A P X Y F L D V B R T S N H M C W 0.707560090465515
# 5: W Y O U R I E A G X J K L D C B H T S N M F V P 0.7075589351593826
# 6: J G O U I S E A P X K Y M C W V H T N R D L F B 0.707549787929756
# 6: J G U X I O E A W K Y F L D C B H T S N R M V P 0.7075212659110061
# 7: J W O U I R E A G X K Y L D C B H T S N M F V P 0.7074562433695609
# 7: J P O U I N E A B X K Y M C G V H T S R L D F W 0.7074435243752765
# 7: J P O U N I E A B K Y X M C G V H T S R L D F W 0.707432984110794
# 7: J G U K I O E A P X Y F M L B W H T S R N D V C 0.7074108195944783
#
# Above layouts that tied for first place, with letter frequencies (2nd layout identical to Engram v2.0):
#
# P Y O U L D F B 76 59 272 97 145 136 86 53
# C I E A R T S N 119 270 445 287 224 331 232 258
# G K J X H M V W 67 19 6 8 180 90 38 60
#
# left: 1.725T right: 1.831T (6.09%)
# Total same-finger bigram frequencies: 31002467582
# Total bigram inward roll frequencies: 4595272424809
#
#
# B Y O U L D W V 53 59 272 97 145 136 60 38
# C I E A H T S N 119 270 445 287 180 331 232 258
# G X <NAME> R M F P 67 8 6 19 224 90 86 76
#
# left: 1.702T right: 1.854T (8.90%)
# Total same-finger bigram frequencies: 31422990907
# Total bigram inward roll frequencies: 4595756397870
# ### Optional: rank variations of top-scoring layouts
#
# As an alternative to simply choosing the candidate winner layout, we can generate variations of this layout and find those variants within a small difference of one another and select from among these variants. For this, we select keys to vary, compute scores for every combination of the letters assigned to these keys, and select among those that are tied for first place. Below we vary those keys with different letters in the two layouts tied for first place, except we fix H above R (as in the second layout, our candidate winner) so that the most frequent bigram (TH, over 100 billion) is easy to type.
score_variants = True
if score_variants:
# Candidate winner above:
#
# B Y O U L D W V
# C I E A H T S N
# G X J K R M F P
# - Y O U L D - -
# C I E A H T S N
# G - J - R M - -
fixed_letters = ['Y','O','U', 'C','I','E','A', 'G','J', 'L','D', 'H','T','S','N', 'R','M']
fixed_letter_indices = [1,2,3, 4,5,6,7, 8,10, 12,13, 16,17,18,19, 20,21]
open_letter_indices = [0, 9,11, 14,15, 22,23]
top_variant_permutation, top_variant_score, variant_letter_permutations = permute_optimize_keys(fixed_letters,
fixed_letter_indices, open_letter_indices, letters24, keys24, Factors24x24,
bigrams, bigram_frequencies, min_score=0, verbose=False)
print(top_variant_permutation)
print(top_variant_score)
if score_variants:
variant_scores = []
nletters = len(fixed_letter_indices) + len(open_letter_indices)
layout_variant_strings = []
for ipermutation, letter_permutation in enumerate(variant_letter_permutations):
variant_letters = np.array(['W' for x in range(nletters)]) # KEEP to initialize!
for imove, open_letter_index in enumerate(open_letter_indices):
variant_letters[open_letter_index] = letter_permutation[imove]
for ifixed, fixed_letter_index in enumerate(fixed_letter_indices):
variant_letters[fixed_letter_index] = fixed_letters[ifixed]
layout_variant_strings.append(variant_letters)
# Compute the score for this permutation:
variant_score = score_layout(Factors24x24, variant_letters,
bigrams, bigram_frequencies, verbose=False)
variant_scores.append(variant_score)
layout_variants = []
for layout_string in layout_variant_strings:
layout = ' '.join(layout_string)
layout_variants.append(layout)
variant_scores_sorted, variant_ranks_sorted, Isort_variants = rank_within_epsilon(variant_scores,
factor24, factor=True, verbose=False)
layout_variants_sorted = []
layout_variant_strings_sorted = []
for i in Isort_variants:
layout_variants_sorted.append(layout_variants[i])
layout_variant_strings_sorted.append(layout_variant_strings[i])
print(' (#) Rank: Layout Score')
for i, rank in enumerate(variant_ranks_sorted):
if rank == 1:
print(' ({0}) {1}: {2} {3}'.format(i + 1, rank,
layout_variants_sorted[i],
variant_scores_sorted[i]))
# Print layouts:
Ifirst_place = []
layout_variants_first_place = []
layout_variant_strings_first_place = []
for i, rank in enumerate(variant_ranks_sorted):
if rank == 1:
layout_string = layout_variant_strings_sorted[i]
layout = layout_variants_sorted[i]
print('\n Layout {0}:\n'.format(i + 1))
print_layout24(layout_string)
print('')
print_layout24_instances(layout_string, letters24, instances24,
bigrams, bigram_frequencies)
Ifirst_place.append(i)
layout_variants_first_place.append(layout)
layout_variant_strings_first_place.append(layout_string)
# Our candidate winner scored highest among its (7! = 5,040) variants. The 42 variants tied for first place are listed below:
#
# (#) Rank: Layout Score
# (1) 1: B Y O U C I E A G X J K L D W V H T S N R M F P 0.7078676989043137
# (2) 1: B Y O U C I E A G K J X L D W V H T S N R M F P 0.7078625576908392
# (3) 1: W Y O U C I E A G X J K L D V B H T S N R M F P 0.7078577061845288
# (4) 1: P Y O U C I E A G K J X L D W V H T S N R M F B 0.7078565092277237
# (5) 1: W Y O U C I E A G K J X L D V B H T S N R M F P 0.7078522283063508
# (6) 1: B Y O U C I E A G X J K L D V W H T S N R M F P 0.7078519616931854
# (7) 1: P Y O U C I E A G X J K L D W V H T S N R M F B 0.7078517296463457
# (8) 1: B Y O U C I E A G X J K L D W F H T S N R M V P 0.7078490260211918
# (9) 1: B Y O U C I E A G K J X L D V W H T S N R M F P 0.707846820479711
# (10) 1: P Y O U C I E A G K J X L D W B H T S N R M V F 0.7078454560742882
# (11) 1: B Y O U C I E A G K J X L D W F H T S N R M V P 0.7078438848077173
# (12) 1: P Y O U C I E A G K J X L D W B H T S N R M F V 0.7078431094974508
# (13) 1: P Y O U C I E A G K J X L D V B H T S N R M F W 0.7078419742548276
# (14) 1: P Y O U C I E A G K J X L D V W H T S N R M F B 0.7078411358167733
# (15) 1: P Y O U C I E A G X J K L D W B H T S N R M V F 0.70784067649291
# (16) 1: W Y O U C I E A G X J K L D F B H T S N R M V P 0.7078403744444377
# (17) 1: P Y O U C I E A G K J X L D W F H T S N R M V B 0.7078391282354274
# (18) 1: P Y O U C I E A G X J K L D W B H T S N R M F V 0.7078383299160728
# (19) 1: P Y O U C I E A G X J K L D V B H T S N R M F W 0.7078371946734496
# (20) 1: W Y O U C I E A G X J K L D B V H T S N R M F P 0.7078371584583636
# (21) 1: P Y O U C I E A G X J K L D V W H T S N R M F B 0.7078363562353953
# (22) 1: B Y O U C I E A G X J K L D F W H T S N R M V P 0.7078359835497579
# (23) 1: W Y O U C I E A G K J X L D F B H T S N R M V P 0.7078348965662598
# (24) 1: P Y O U C I E A G X J K L D W F H T S N R M V B 0.7078343486540493
# (25) 1: W Y O U C I E A G K J X L D B V H T S N R M F P 0.7078316805801855
# (26) 1: B Y O U C I E A G K J X L D F W H T S N R M V P 0.7078308423362834
# (27) 1: P Y O U C I E A G K J X L D W V H T S N R M B F 0.7078260494151115
# (28) 1: P Y O U C I E A G K J X L D F W H T S N R M V B 0.7078260359767987
# (29) 1: P Y O U C I E A G K J X L D F B H T S N R M V W 0.7078245475443425
# (30) 1: W Y O U C I E A G X J K L D F V H T S N R M B P 0.7078214911264225
# (31) 1: P Y O U C I E A G K J X L D B V H T S N R M F W 0.7078214181411706
# (32) 1: P Y O U C I E A G X J K L D W V H T S N R M B F 0.7078212698337334
# (33) 1: P Y O U C I E A G X J K L D F W H T S N R M V B 0.7078212563954208
# (34) 1: B Y O U C I E A G X J K L D F V H T S N R M W P 0.7078210837714037
# (35) 1: P Y O U C I E A G X J K L D F B H T S N R M V W 0.7078197679629645
# (36) 1: W Y O U C I E A G X J K L D V F H T S N R M B P 0.70781880339861
# (37) 1: B Y O U C I E A G X J K L D V F H T S N R M W P 0.7078184238466051
# (38) 1: W Y O U C I E A G X J K L D B F H T S N R M V P 0.7078172387197521
# (39) 1: P Y O U C I E A G X J K L D B V H T S N R M F W 0.7078166385597925
# (40) 1: W Y O U C I E A G K J X L D F V H T S N R M B P 0.7078160132482443
# (41) 1: P Y O U C I E A G K J X L D V B H T S N R M W F 0.7078159646633373
# (42) 1: B Y O U C I E A G K J X L D F V H T S N R M W P 0.7078159425579292
#
# Letters shared across all layout variants tied for first place:
#
# - Y O U L D - -
# C I E A H T S N
# G - J - R M - -
#
# If we list only those layouts in descending order by score that have progressively lower same-finger bigram counts, then we end up with the candidate winner (Variant 1) and Variant 3:
#
# Variant 1 = Layout 2 above:
#
# B Y O U L D W V
# C I E A H T S N
# G X J K R M F P
#
# 53 59 272 97 145 136 60 38
# 119 270 445 287 180 331 232 258
# 67 8 6 19 224 90 86 76
#
# left: 1.702T right: 1.854T (8.90%)
# Total same-finger bigram frequencies: 31422990907
# Total bigram inward roll frequencies: 4595756397870
#
# Variant 3:
#
# W Y O U L D V B
# C I E A H T S N
# G X J K R M F P
#
# 60 59 272 97 145 136 38 53
# 119 270 445 287 180 331 232 258
# 67 8 6 19 224 90 86 76
#
# left: 1.709T right: 1.847T (8.07%)
# Total same-finger bigram frequencies: 28475089052
# Total bigram inward roll frequencies: 4605502028148
if score_variants:
layout_variant_strings_first_place = [
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','F','P'],
['B','Y','O','U','C','I','E','A','G','K','J','X','L','D','W','V','H','T','S','N','R','M','F','P'],
['W','Y','O','U','C','I','E','A','G','X','J','K','L','D','V','B','H','T','S','N','R','M','F','P'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','W','V','H','T','S','N','R','M','F','B'],
['W','Y','O','U','C','I','E','A','G','K','J','X','L','D','V','B','H','T','S','N','R','M','F','P'],
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','V','W','H','T','S','N','R','M','F','P'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','F','B'],
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','F','H','T','S','N','R','M','V','P'],
['B','Y','O','U','C','I','E','A','G','K','J','X','L','D','V','W','H','T','S','N','R','M','F','P'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','W','B','H','T','S','N','R','M','V','F'],
['B','Y','O','U','C','I','E','A','G','K','J','X','L','D','W','F','H','T','S','N','R','M','V','P'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','W','B','H','T','S','N','R','M','F','V'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','V','B','H','T','S','N','R','M','F','W'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','V','W','H','T','S','N','R','M','F','B'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','B','H','T','S','N','R','M','V','F'],
['W','Y','O','U','C','I','E','A','G','X','J','K','L','D','F','B','H','T','S','N','R','M','V','P'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','W','F','H','T','S','N','R','M','V','B'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','B','H','T','S','N','R','M','F','V'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','V','B','H','T','S','N','R','M','F','W'],
['W','Y','O','U','C','I','E','A','G','X','J','K','L','D','B','V','H','T','S','N','R','M','F','P'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','V','W','H','T','S','N','R','M','F','B'],
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','F','W','H','T','S','N','R','M','V','P'],
['W','Y','O','U','C','I','E','A','G','K','J','X','L','D','F','B','H','T','S','N','R','M','V','P'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','F','H','T','S','N','R','M','V','B'],
['W','Y','O','U','C','I','E','A','G','K','J','X','L','D','B','V','H','T','S','N','R','M','F','P'],
['B','Y','O','U','C','I','E','A','G','K','J','X','L','D','F','W','H','T','S','N','R','M','V','P'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','W','V','H','T','S','N','R','M','B','F'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','F','W','H','T','S','N','R','M','V','B'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','F','B','H','T','S','N','R','M','V','W'],
['W','Y','O','U','C','I','E','A','G','X','J','K','L','D','F','V','H','T','S','N','R','M','B','P'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','B','V','H','T','S','N','R','M','F','W'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','B','F'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','F','W','H','T','S','N','R','M','V','B'],
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','F','V','H','T','S','N','R','M','W','P'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','F','B','H','T','S','N','R','M','V','W'],
['W','Y','O','U','C','I','E','A','G','X','J','K','L','D','V','F','H','T','S','N','R','M','B','P'],
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','V','F','H','T','S','N','R','M','W','P'],
['W','Y','O','U','C','I','E','A','G','X','J','K','L','D','B','F','H','T','S','N','R','M','V','P'],
['P','Y','O','U','C','I','E','A','G','X','J','K','L','D','B','V','H','T','S','N','R','M','F','W'],
['W','Y','O','U','C','I','E','A','G','K','J','X','L','D','F','V','H','T','S','N','R','M','B','P'],
['P','Y','O','U','C','I','E','A','G','K','J','X','L','D','V','B','H','T','S','N','R','M','W','F'],
['B','Y','O','U','C','I','E','A','G','K','J','X','L','D','F','V','H','T','S','N','R','M','W','P']]
# ## Step 4: Evaluate winning layout <a name="step4">
#
# We evaluate the candidate winner with tests:
#
# 1. Evaluate optimized layouts using interkey speed estimates
# 2. Evaluate variants of the candidate winner using interkey speed estimates
# 3. Evaluate sensitivity of the variants to the scoring parameters
# 4. Search for higher-scoring layouts by rearranging letters
# 5. Compare with alternate layout based solely on interkey speed estimates
# ### Test 1. Evaluate optimized layouts using interkey speed estimates
# Below we rescore all of the 20 top-scoring layouts optimized from the 20 initialized layouts, and replace the factor matrix with the inter-key speed matrix. The same two layouts that tied for first place do so again.
#
# *Note:*
#
# The speed matrix contains normalized interkey stroke times derived from a published study ("Estimation of digraph costs for keyboard layout optimization", A Iseri, Ma Eksioglu, International Journal of Industrial Ergonomics, 48, 127-138, 2015). To establish which layouts are within a small difference of each other when using the speed matrix, we define an epsilon equal to 131.58 ms for a single bigram (of the 32^2 possible bigrams), where 131.58 ms is the fastest measured digraph tapping speed (30,000/228 = 131.58 ms) recorded in the above study.
#
# "Digraph-tapping rate changes dramatically across the digraph types. The range is between 82 and 228 taps per 30 s. The difference is nearly three times between the slowest and the fastest digraphs. From this result it can be concluded that the assignment of letter pairs on the correct digraph keys on the keyboard can have a high impact on the typing speed."
test_layout_strings = first_layout_strings_sorted
# +
# # %load code/test/score_speed_of_layouts.py
data_matrix_speed = Speed24x24 # SpeedSymmetric24x24
speed_scores = []
for letters in test_layout_strings:
score = score_layout(data_matrix_speed, letters, bigrams, bigram_frequencies, verbose = False)
speed_scores.append(score)
speed_scores_sorted, speed_ranks_sorted, Isort_speed = rank_within_epsilon(speed_scores,
epsilon, factor=False, verbose=False)
speed_layouts_sorted = []
speed_layout_strings_sorted = []
for i in Isort_speed:
speed_layouts_sorted.append(' '.join(test_layout_strings[i]))
speed_layout_strings_sorted.append(test_layout_strings[i])
count = 0
print(' (#) Layout Speed score')
for i, isort_speed in enumerate(Isort_speed):
if speed_ranks_sorted[isort_speed] == 1:
count += 1
if isort_speed < 9:
s = ' '
else:
s = ' '
print(' ({0}) {1}{2} {3}'.format(isort_speed+1, s,
speed_layouts_sorted[i],
speed_scores_sorted[i]))
print('\n {0} of {1} layouts tied for first place'.format(count, len(test_layout_strings)))
# -
# ### Test 2. Evaluate variants of the candidate winner using interkey speed estimates
# Below we rescore all of the 5,040 variants of the candidate winner that are tied for first place, replacing the factor matrix with the inter-key speed matrix. The candidate winner scores highest.
test_layout_strings = layout_variant_strings_first_place
# +
# # %load code/test/score_speed_of_layouts.py
data_matrix_speed = Speed24x24 # SpeedSymmetric24x24
speed_scores = []
for letters in test_layout_strings:
score = score_layout(data_matrix_speed, letters, bigrams, bigram_frequencies, verbose = False)
speed_scores.append(score)
speed_scores_sorted, speed_ranks_sorted, Isort_speed = rank_within_epsilon(speed_scores,
epsilon, factor=False, verbose=False)
speed_layouts_sorted = []
speed_layout_strings_sorted = []
for i in Isort_speed:
speed_layouts_sorted.append(' '.join(test_layout_strings[i]))
speed_layout_strings_sorted.append(test_layout_strings[i])
count = 0
print(' Layout Speed score')
for i, isort_speed in enumerate(Isort_speed):
if speed_ranks_sorted[isort_speed] == 1:
count += 1
if isort_speed < 9:
s = ' '
else:
s = ' '
print(' ({0}){1}{2} {3}'.format(isort_speed+1, s,
speed_layouts_sorted[i],
speed_scores_sorted[i]))
print(' {0} of {1} layouts tied for first place'.format(count, len(test_layout_strings)))
# -
# Variant 1 (the candidate winner above) scores highest:
#
# Layout Speed score
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7023756439425117
# (30) W Y O U C I E A G X J K L D F V H T S N R M B P 0.7023734892525684
# (20) W Y O U C I E A G X J K L D B V H T S N R M F P 0.7023700909720256
# (6) B Y O U C I E A G X J K L D V W H T S N R M F P 0.7023688377122477
# (22) B Y O U C I E A G X J K L D F W H T S N R M V P 0.702367226885074
# (3) W Y O U C I E A G X J K L D V B H T S N R M F P 0.7023627643568422
# (7) P Y O U C I E A G X J K L D W V H T S N R M F B 0.7023607516204574
# (16) W Y O U C I E A G X J K L D F B H T S N R M V P 0.7023603659811735
# (8) B Y O U C I E A G X J K L D W F H T S N R M V P 0.7023583852103916
# (21) P Y O U C I E A G X J K L D V W H T S N R M F B 0.7023538733148424
# (2) B Y O U C I E A G K J X L D W V H T S N R M F P 0.7023520610893563
# (4) P Y O U C I E A G K J X L D W V H T S N R M F B 0.7023484279427685
# (18) P Y O U C I E A G X J K L D W B H T S N R M F V 0.7023464351081202
# (25) W Y O U C I E A G K J X L D B V H T S N R M F P 0.7023461467370498
# (9) B Y O U C I E A G K J X L D V W H T S N R M F P 0.7023452548590922
# (19) P Y O U C I E A G X J K L D V B H T S N R M F W 0.7023449431149574
# (24) P Y O U C I E A G X J K L D W F H T S N R M V B 0.7023436988861739
# (26) B Y O U C I E A G K J X L D F W H T S N R M V P 0.7023436440319186
# (14) P Y O U C I E A G K J X L D V W H T S N R M F B 0.7023415496371536
# (28) P Y O U C I E A G K J X L D F W H T S N R M V B 0.7023402284944377
# (5) W Y O U C I E A G K J X L D V B H T S N R M F P 0.7023388201218663
# (23) W Y O U C I E A G K J X L D F B H T S N R M V P 0.7023364217461976
# (11) B Y O U C I E A G K J X L D W F H T S N R M V P 0.7023348023572361
# (12) P Y O U C I E A G K J X L D W B H T S N R M F V 0.7023341114304313
# (13) P Y O U C I E A G K J X L D V B H T S N R M F W 0.7023326194372687
# (17) P Y O U C I E A G K J X L D W F H T S N R M V B 0.7023313752084851
# (27) P Y O U C I E A G K J X L D W V H T S N R M B F 0.7023309175507675
# (29) P Y O U C I E A G K J X L D F B H T S N R M V W 0.7023301885671278
# (15) P Y O U C I E A G X J K L D W B H T S N R M V F 0.7023301589694194
# (10) P Y O U C I E A G K J X L D W B H T S N R M V F 0.7023178352917306
#
# 30 of 42 layouts tied for first place
# ### Test 3. Evaluate sensitivity of the variants to the scoring parameters
#
# We run a test below on the variants of the candidate winner layout to see how robust they are to removal of scoring parameters. We removed each of the 11 scoring parameters one by one and ranked the new scores for the variants above. Variant 1 (the candidate winner) scores highest for 8 of the 11 cases, and second highest for two other cases, demonstrating that this layout is not sensitive to individual parameters.
# +
# # %load code/test/remove_parameters_rescore.py
params0 = [side_above_3away, side_above_2away, side_above_1away, middle_above_ring, ring_above_middle,
outward, skip_row_3away, skip_row_2away, skip_row_1away, skip_row_0away, same_finger]
param_names = ['side_above_3away', 'side_above_2away', 'side_above_1away',
'middle_above_ring', 'ring_above_middle', 'outward', 'skip_row_3away',
'skip_row_2away', 'skip_row_1away', 'skip_row_0away', 'same_finger']
params_lists = []
for i in range(len(params0)):
params_list = params0.copy()
params_list[i] = 1.0
params_lists.append(params_list)
for iparam, P in enumerate(params_lists):
print(' Remove parameter {0}:'.format(param_names[iparam]))
data_matrix_param = create_24x24_flow_matrix(not_home_row, side_top,
P[0],P[1],P[2],P[3],P[4],P[5],P[6],P[7],P[8],P[9],P[10],
1,1,1,1,1,1)
if apply_strength:
data_matrix_param = Strength24x24 * data_matrix_param
param_scores = []
for letters in test_layout_strings:
score = score_layout(data_matrix_param, letters, bigrams, bigram_frequencies, verbose=False);
param_scores.append(score)
param_scores_sorted, param_ranks_sorted, Isort_param = rank_within_epsilon(param_scores, factor24, factor=True, verbose=False)
param_layouts_sorted = []
param_layout_strings_sorted = []
for i in Isort_param:
param_layouts_sorted.append(' '.join(test_layout_strings[i]))
param_layout_strings_sorted.append(test_layout_strings[i])
print(' Variant Score')
count = 0
for i, isort_param in enumerate(Isort_param):
count += 1
if param_ranks_sorted[isort_param] == 1:
if isort_param < 9:
s = ' '
else:
s = ' '
print(' ({0}){1}{2} {3}'.format(isort_param+1, s,
param_layouts_sorted[i],
param_scores_sorted[i]))
print(' {0} of {1} layouts tied for first place'.format(count, len(test_layout_strings)))
# + active=""
# Remove parameter side_above_3away:
# (6) B Y O U C I E A G X J K L D V W H T S N R M F P 0.7107633027019034
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7107623334764219
# Remove parameter side_above_2away:
# (2) B Y O U C I E A G K J X L D W V H T S N R M F P 0.7130518654000207
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7130513395263505
# Remove parameter side_above_1away:
# (5) W Y O U C I E A G K J X L D V B H T S N R M F P 0.7148772594313253
# (3) W Y O U C I E A G X J K L D V B H T S N R M F P 0.7148711293283665
# (2) B Y O U C I E A G K J X L D W V H T S N R M F P 0.7148593915832421
# (23) W Y O U C I E A G K J X L D F B H T S N R M V P 0.7148583101988224
# (4) P Y O U C I E A G K J X L D W V H T S N R M F B 0.7148543601588774
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7148530691183211
# Remove parameter middle_above_ring:
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7092201999241033
# Remove parameter ring_above_middle:
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7114189279608791
# Remove parameter outward:
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7201947803218552
# Remove parameter skip_row_3away:
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7087608738602452
# Remove parameter skip_row_2away:
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7127292945043059
# Remove parameter skip_row_1away:
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7182207100993533
# Remove parameter skip_row_0away:
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.712081162928148
# Remove parameter same_finger:
# (1) B Y O U C I E A G X J K L D W V H T S N R M F P 0.7305410820225844
# -
# ### Test 4. Search for higher-scoring layouts by rearranging letters
#
# The following test is to see if allowing random sets of eight letters to rearrange in every possible combination improves the score of the winning layout. After randomly selecting eight letters from (13 of the letters in) the top-scoring layout, creating layouts from every permutation of these letters, and computing their scores, we get identical results as the original layout. We repeated this test over a thousand times (40,320,000 layouts).
winner24 = ['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','F','P']
run_stability_test = True
if run_stability_test:
original_score = score_layout(Factors24x24, winner24, bigrams, bigram_frequencies, verbose=False)
top_score_test4 = original_score
nunber_of_tests = 1000
size_random_set = 8
indices = [0,1, 8,9,10,11, 12,13,14,15, 21,22,23]
# B Y O U L D W V
# C I E A H T S N
# G X J K R M F P
# 0 1 - - 12 13 14 15
# - - - - - - - -
# 8 9 10 11 - 21 22 23
print(original_score)
for i in range(nunber_of_tests):
print(i)
letters_copy = winner24.copy()
random_indices = []
while np.size(random_indices) < size_random_set:
random_index = indices[np.int( np.round( (np.size(indices) - 1) * np.random.random(1) )[0])]
if random_index not in random_indices:
random_indices.append(random_index)
for irand in random_indices:
letters_copy[np.int(irand)] = ''
top_permutation_test4, top_score_test4 = permute_optimize(winner24, letters_copy, letters24, keys24,
Factors24x24, bigrams, bigram_frequencies,
min_score=top_score_test4, verbose=False)
if ''.join(top_permutation_test4) != ''.join(winner24) and top_score_test4 > original_score:
print(top_score_test4)
print(*top_permutation_test4)
# ### Test 5. Compare with alternate layout based solely on interkey speed estimates
#
# Since we use interkey speed estimates to independently corroborate the practical utility of our top-scoring initialized layouts and variants generated from our candidate winner, the question arises whether a better layout could be generated using the above procedure and based solely on interkey speed estimates. To do this, we simply set apply_strength=False and Factors24x24=Speed24x24 and ran Steps 1 through 3 above. The resulting layouts have two to three times higher same-finger bigram frequencies, which is not a good sign of the ease with which they can be typed. This indirectly demonstrates that fast-to-type layouts do not necessarily translate to less strenuous layouts.
#
# (#) Rank Score
# (0) 1: J Y U G I O E A X K W D L C F V R T S N H M P B 0.7028248210994403
# (1) 1: J Y O F U I E A X K G D L C B V R T S N H M P W 0.7028092866027337
# (2) 1: J B U P I O E A X K Y F L D C V H T S R N M W G 0.7027885065002167
# (3) 1: J P O F U I E A X Y K G L D C V H T S R N M W B 0.7027774348054611
# (4) 1: J Y U G I O E A X K W D L C F V H T S N R M P B 0.7027766978615982
# (5) 2: J Y O F U I E A X K G D L C W V H T S N R M P B 0.7027604410329258
# (6) 3: J Y O F I U E A X K G D L C B V R T S N H M P W 0.7027015337086406
# (7) 3: J P O F I U E A X Y K G L D C V H T S R N M W B 0.7026779438898121
# (8) 3: J Y O F I U E A X K G D L C W V H T S N R M P B 0.7026531181501796
# (9) 4: J U O F I H E A X Y K G D L C V T S N R P M W B 0.7026052409973239
# (10) 4: J U O F H I E A X Y K G D L C V T S N R P M W B 0.7025798551167619
# (11) 5: J U O G I H E A X Y K D C F W V R T S N L M P B 0.7025168489505383
# (12) 5: J U O G H I E A X Y K D C F W V R T S N L M P B 0.7025072606193864
# (13) 6: J G O F I S E A X Y K U D L C V H T N R P M W B 0.7024132916102113
# (14) 6: J Y O F S I E A X K G U D L C V H T N R P M W B 0.7023840624087121
# (15) 7: J W O U I R E A X K Y G L C F V H T S N D M P B 0.7021673985385113
# (16) 7: J P O F I N E A X Y K U M G C V H T S R L D W B 0.7021345744708818
# (17) 8: J Y O F R I E A X K G U L C W V H T S N D M P B 0.7020921733913089
# (18) 8: J P O F N I E A X Y K U M G C V H T S R L D W B 0.7020744010726611
#
# Layouts tied for first place, with letter frequencies:
#
# Layout 1:
#
# J Y U G L C F V
# I O E A R T S N
# X K W D H M P B
#
# 6 59 97 67 145 119 86 38
# 270 272 445 287 224 331 232 258
# 8 19 60 136 180 90 76 53
#
# left: 1.726T right: 1.830T (6.03%)
# Total same-finger bigram frequencies: 83350937269
# Total bigram inward roll frequencies: 4619080035315
#
# Layout 2:
#
# J Y O F L C B V
# U I E A R T S N
# X K G D H M P W
#
# 6 59 272 86 145 119 53 38
# 97 270 445 287 224 331 232 258
# 8 19 67 136 180 90 76 60
#
# left: 1.752T right: 1.804T (2.99%)
# Total same-finger bigram frequencies: 85067873377
# Total bigram inward roll frequencies: 4595756638318
#
# Layout 3:
#
# J B U P L D C V
# I O E A H T S R
# X K Y F N M W G
#
# 6 53 97 76 145 136 119 38
# 270 272 445 287 180 331 232 224
# 8 19 59 86 258 90 60 67
#
# left: 1.678T right: 1.878T (11.89%)
# Total same-finger bigram frequencies: 67426732036
# Total bigram inward roll frequencies: 4698191302186
#
# Layout 4:
#
# J P O F L D C V
# U I E A H T S R
# X Y K G N M W B
#
# 6 76 272 86 145 136 119 38
# 97 270 445 287 180 331 232 224
# 8 59 19 67 258 90 60 53
#
# left: 1.692T right: 1.864T (10.17%)
# Total same-finger bigram frequencies: 55581492895
# Total bigram inward roll frequencies: 4538464009444
#
# Layout 5:
#
# J Y U G L C F V
# I O E A H T S N
# X K W D R M P B
#
# 6 59 97 67 145 119 86 38
# 270 272 445 287 180 331 232 258
# 8 19 60 136 224 90 76 53
#
# left: 1.726T right: 1.830T (6.03%)
# Total same-finger bigram frequencies: 83350937269
# Total bigram inward roll frequencies: 4619080035315
# ### Assign letters Z and Q and test left/right swap
#
# Test to see if equal or higher scores are obtained for the following:
#
# 1. Assign Z and either Q or J to keys 112 and 113
# 2. Swap left and right sides
# +
layouts_26letters = [
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','F','P', '-','-','-', '-','-','-', 'Z','Q'],
['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','F','P', '-','-','-', '-','-','-', 'Q','Z'],
['V','W','D','L','N','S','T','H','P','F','M','R','U','O','Y','B','A','E','I','C','K','J','X','G', '-','-','-', '-','-','-', 'Z','Q'],
['V','W','D','L','N','S','T','H','P','F','M','R','U','O','Y','B','A','E','I','C','K','J','X','G', '-','-','-', '-','-','-', 'Q','Z']]
data_matrix = Factors32x32
scores_26letters = []
for layout_26letters in layouts_26letters:
scores_26letters.append(score_layout(data_matrix, layout_26letters, bigrams, bigram_frequencies, verbose=False))
scores_26letters_sorted, ranks_26letters_sorted, Isort_26letters = rank_within_epsilon(scores_26letters,
factor32, factor=True, verbose=False)
print('\n Rank Score')
for i, rank in enumerate(ranks_26letters_sorted):
layout_string = layouts_26letters[Isort_26letters[i]]
layout = ' '.join(layout_string)
print(' {0}: {1} {2}'.format(rank, layout, scores_26letters_sorted[i]))
print('')
print_layout24(layouts_26letters[0])
#bigram_strings = [['f','l'],['f','r'],['p','l'],['p','r'],['w','r'],['w','l']]
#for bigram_string in bigram_strings:
# print_bigram_frequency(bigram_string, bigrams, bigram_frequencies)
# -
# Z above Q received the highest score:
#
# Rank Score
# 1: B Y O U C I E A G X J K L D W V H T S N R M F P - - - - - - Z Q 0.621987268013091
# 1: B Y O U C I E A G X J K L D W V H T S N R M F P - - - - - - Q Z 0.6219870422703005
# 1: V W D L N S T H P F M R U O Y B A E I C K J X G - - - - - - Q Z 0.6219847143830128
# 1: V W D L N S T H P F M R U O Y B A E I C K J X G - - - - - - Z Q 0.6219774708803041
#
# The letters of the Engram layout:
#
# B Y O U L D W V Z
# C I E A H T S N Q
# G X J K R M F P
winner24 = ['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','F','P']
winner32 = ['B','Y','O','U','C','I','E','A','G','X','J','K','L','D','W','V','H','T','S','N','R','M','F','P', '-','-','-', '-','-','-', 'Z','Q']
# ### Optional stability test
#
# The following test is used to compare the score of the winning layout after rearranging random letters.
run_stability_test = True
if run_stability_test:
original_score = score_layout(Factors24x24, winner24, bigrams, bigram_frequencies, verbose=False)
top_score_test5 = original_score
nunber_of_tests = 1000
size_random_set = 8
indices = [0,1, 8,9,10,11, 12,13,14,15, 21,22,23]
# B Y O U L D W V
# C I E A H T S N
# G X J K R M F P
# 0 1 - - 12 13 14 15
# - - - - - - - -
# 8 9 10 11 - 21 22 23
print(original_score)
for i in range(nunber_of_tests):
print(i)
letters_copy = winner24.copy()
random_indices = []
while np.size(random_indices) < size_random_set:
random_index = indices[np.int( np.round( (np.size(indices) - 1) * np.random.random(1) )[0])]
if random_index not in random_indices:
random_indices.append(random_index)
for irand in random_indices:
letters_copy[np.int(irand)] = ''
top_permutation_test5, top_score_test5 = permute_optimize(winner24, letters_copy, letters24, keys24,
Factors24x24, bigrams, bigram_frequencies,
min_score=top_score_test5, verbose=False)
if ''.join(top_permutation_test5) != ''.join(winner24) and top_score_test5 > original_score:
print(top_score_test5)
print(*top_permutation_test5)
# + [markdown] colab_type="text" id="TPW3wZw2DzT7"
# ## Step 5: Arrange non-letter characters in easy-to-remember places <a name="step5">
#
# Now that we have all 26 letters accounted for, we turn our attention to non-letter characters, taking into account frequency of punctuation and ease of recall.
# + [markdown] colab_type="raw" id="ul_j8VsZDzT7"
# ### Frequency of punctuation marks
#
# - Statistical values of punctuation frequency in 20 English-speaking countries (Table 1): <br>
# <NAME> & <NAME>. (2018). Frequency Distributions of Punctuation Marks in English: Evidence from Large-scale Corpora. English Today. 10.1017/S0266078418000512. <br>
# https://www.researchgate.net/publication/328512136_Frequency_Distributions_of_Punctuation_Marks_in_English_Evidence_from_Large-scale_Corpora
# <br>"frequency of punctuation marks attested for twenty English-speaking countries and regions... The data were acquired through GloWbE."
# "The corpus of GloWbE (2013) is a large English corpus collecting international English from the internet, containing about 1.9 billion words of text from twenty different countries. For further information on the corpora used, see https://corpus.byu.edu/."
#
# - Google N-grams and Twitter analysis: <br>
# "Punctuation Input on Touchscreen Keyboards: Analyzing Frequency of Use and Costs" <br>
# <NAME>, <NAME> - College Park: The Human-Computer Interaction Lab. 2013 <br>
# https://www.cs.umd.edu/sites/default/files/scholarly_papers/Malik.pdf <br>
# "the Twitter corpora included substantially higher punctuation use than the Google corpus, <br>
# comprising 7.5% of characters in the mobile tweets and 7.6% in desktop versus only 4.4%... <br>
# With the Google corpus,only 6 punctuation symbols (. -’ ( ) “) appeared more frequently than [q]"
#
# - "Frequencies for English Punctuation Marks" by <NAME> <br>
# http://www.viviancook.uk/Punctuation/PunctFigs.htm <br>
# "Based on a writing system corpus some 459 thousand words long. <br>
# This includes three novels of different types (276 thousand words), <br>
# selections of articles from two newspapers (55 thousand), <br>
# one bureaucratic report (94 thousand), and assorted academic papers <br>
# on language topics (34 thousand). More information is in <br>
# Cook, V.J. (2013) ‘Standard punctuation and the punctuation of the street’ <br>
# in <NAME> and <NAME> (eds.), Essential Topics in Applied Linguistics and Multilingualism, <br>
# Springer International Publishing Switzerland (2013), 267-290"
#
# - "A Statistical Study of Current Usage in Punctuation": <br>
# <NAME>., & <NAME>. (1924). A Statistical Study of Current Usage in Punctuation. The English Journal, 13(5), 325-331. doi:10.2307/802253
#
# - "Computer Languages Character Frequency"
# by <NAME>. <br>
# Date: 2013-05-23. Last updated: 2020-06-29. <br>
# http://xahlee.info/comp/computer_language_char_distribution.html <br>
# NOTE: biased toward C (19.8%) and Py (18.5%), which have high use of "_".
#
# Frequency:
#
# Sun: Malik: Ruhlen: Cook: Xah:
# /1M N-gram % /10,000 /1,000 All% JS% Py%
#
# . 42840.02 1.151 535 65.3 6.6 9.4 10.3
# , 44189.96 556 61.6 5.8 8.9 7.5
# " 2.284 44 26.7 3.9 1.6 6.2
# ' 2980.35 0.200 40 24.3 4.4 4.0 8.6
# - 9529.78 0.217 21 15.3 4.1 1.9 3.0
# () 4500.81 0.140 7 7.4 9.8 8.1
# ; 1355.22 0.096 22 3.2 3.8 8.6
# z 0.09 - -
# : 3221.82 0.087 11 3.4 3.5 2.8 4.7
# ? 4154.78 0.032 14 5.6 0.3
# / 0.019 4.0 4.9 1.1
# ! 2057.22 0.013 3 3.3 0.4
# _ 0.001 11.0 2.9 10.5
# = 4.4 10.7 5.4
# * 3.6 2.1
# > 3.0 1.4
# $ 2.7 1.6
# # 2.2 3.2
# {} 1.9 4.2
# < 1.3
# & 1.3
# \ 1.2 1.1
# [] 0.9 1.9 1.2
# @ 0.8
# | 0.6
# + 0.6 1.9
# % 0.4
# + [markdown] colab_type="text" id="sdl3lLOfDzT8"
# ### Add punctuation keys and number keys
#
# We will assign the most frequent punctuation according to Sun, et al (2018) to the six keys in the middle two columns: . , " ' - ? ; : () ! _
#
# B Y O U ' " L D W V Z
# C I E A , . H T S N Q
# G X J K - ? R M F P
#
# We will use the Shift key to group similar punctuation marks (separating and joining marks in the left middle column and closing marks in the right middle column):
#
# B Y O U '( ") L D W V Z
# C I E A ,; .: H T S N Q
# G X J K -_ ?! R M F P
#
# **Separating marks (left)**: The comma separates text in lists; the semicolon can be used in place of the comma to separate items in a list (especially if these items contain commas); open parenthesis sets off an explanatory word, phrase, or sentence.
#
# **Joining marks (left)**: The apostrophe joins words as contractions; the hyphen joins words as compounds; the underscore joins words in cases where whitespace characters are not permitted (such as in variables or file names).
#
# **Closing marks (right)**: A sentence usually ends with a period, question mark, or exclamation mark. The colon ends one statement but precedes the following: an explanation, quotation, list, etc. Double quotes and close parenthesis closes a word, clause, or sentence separated by an open parenthesis.
#
# **Number keys**:
# The numbers are flanked to the left and right by [square brackets], and {curly brackets} accessed by the Shift key. Each of the numbers is paired with a mathematical or logic symbol accessed by the Shift key:
#
# { | = ~ + < > ^ & % * } \
# [ 1 2 3 4 5 6 7 8 9 0 ] /
#
# 1: | (vertical bar or "pipe" represents the logical OR operator: 1 stroke, looks like the number one)
# 2: = (equal: 2 strokes, like the Chinese character for "2")
# 3: ~ (tilde: "almost equal", often written with 3 strokes, like the Chinese character for "3")
# 4: + (plus: has four quadrants; resembles "4")
# 5 & 6: < > ("less/greater than"; these angle brackets are directly above the other bracket keys)
# 7: ^ (caret for logical XOR operator as well as exponentiation; resembles "7")
# 8: & (ampersand: logical AND operator; resembles "8")
# 9: % (percent: related to division; resembles "9")
# 0: * (asterisk: for multiplication; resembles "0")
#
# The three remaining keys in many common keyboards (flanking the upper right hand corner Backspace key) are displaced in special keyboards, such as the Kinesis Advantage and Ergodox. For the top right key, we will assign the forward slash and backslash: / \\. For the remaining two keys, we will assign two symbols that in modern usage have significance in social media: the hash/pound sign and the "at sign". The hash or hashtag identifies digital content on a specific topic (the Shift key accesses the dollar sign). The "at sign" identifies a location or affiliation (such as in email addresses) and acts as a "handle" to identify users in popular social media platforms and online forums.
#
# The resulting Engram layout:
#
# { | = ~ + < > ^ & % * } \
# [ 1 2 3 4 5 6 7 8 9 0 ] /
#
# B Y O U '( ") L D W V Z #$ @`
# C I E A ,; .: H T S N Q
# G X J K -_ ?! R M F P
#
# -
| engram_layout_v2.0.ipynb |
# +
"""
Cat class
"""
# Implement a class called "Cat" with the following properties:
# name
# breed
# age
# also, implement a method called "speak" that should print out "purr"
class Cat:
def __init__(self, name, breed, age):
self.name = name
self.breed = breed
self.age = age
def speak(self):
print('purr')
c = Cat("test", "test", 5)
c.speak()
| pset_classes/class_basics/solutions/nb/p2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
df = pd.read_csv('data/src/sample_pandas_normal.csv', index_col=0)
print(df)
print(df.drop('Charlie', axis=0))
print(df.drop('Charlie'))
print(df.drop(index='Charlie'))
print(df.drop(['Bob', 'Dave', 'Frank']))
print(df.drop(index=['Bob', 'Dave', 'Frank']))
df_org = df.copy()
df_org.drop(index=['Bob', 'Dave', 'Frank'], inplace=True)
print(df_org)
print(df.index[[1, 3, 5]])
print(df.drop(df.index[[1, 3, 5]]))
print(df.drop(index=df.index[[1, 3, 5]]))
df_noindex = pd.read_csv('data/src/sample_pandas_normal.csv')
print(df_noindex)
print(df_noindex.index)
print(df_noindex.drop([1, 3, 5]))
print(df_noindex.drop(df_noindex.index[[1, 3, 5]]))
df_noindex_sort = df_noindex.sort_values('state')
print(df_noindex_sort)
print(df_noindex_sort.index)
print(df_noindex_sort.drop([1, 3, 5]))
print(df_noindex_sort.drop(df_noindex_sort.index[[1, 3, 5]]))
print(df.drop('state', axis=1))
print(df.drop(columns='state'))
print(df.drop(['state', 'point'], axis=1))
print(df.drop(columns=['state', 'point']))
df_org = df.copy()
df_org.drop(columns=['state', 'point'], inplace=True)
print(df_org)
print(df.columns[[1, 2]])
print(df.drop(df.columns[[1, 2]], axis=1))
print(df.drop(columns=df.columns[[1, 2]]))
print(df.drop(index=['Bob', 'Dave', 'Frank'],
columns=['state', 'point']))
print(df.drop(index=df.index[[1, 3, 5]],
columns=df.columns[[1, 2]]))
df_org = df.copy()
df_org.drop(index=['Bob', 'Dave', 'Frank'],
columns=['state', 'point'], inplace=True)
print(df_org)
| notebook/pandas_drop.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# + slideshow={"slide_type": "skip"}
# %matplotlib notebook
import numpy as np
import matplotlib.pyplot as plt
# + [markdown] slideshow={"slide_type": "slide"}
# # Vocabulary
#
# In the previous parts, you learned how matplotlib organizes plot-making by figures and axes. We broke down the components of a basic figure and learned how to create them. You also learned how to add one or more axes to a figure, and how to tie them together. You even learned how to change some of the basic appearances of the axes. Finally, we went over some of the many plotting methods that matplotlib has to draw on those axes. With all that knowledge, you should be off making great and wonderful figures.
#
# Of course! While the previous sections may have taught you some of the structure and syntax of matplotlib, it did not describe much of the substance and vocabulary of the library. This section will go over many of the properties that are used throughout the library. Note that while many of the examples in this section may show one way of setting a particular property, that property may be applicible elsewhere in completely different context. This is the "language" of matplotlib.
# + [markdown] slideshow={"slide_type": "slide"}
# # Colors
#
# This is, perhaps, the most important piece of vocabulary in matplotlib. Given that matplotlib is a plotting library, colors are associated with everything that is plotted in your figures. Matplotlib supports a [very robust language](http://matplotlib.org/api/colors_api.html#module-matplotlib.colors) for specifying colors that should be familiar to a wide variety of users.
#
# ### Colornames
# First, colors can be given as strings. For very basic colors, you can even get away with just a single letter:
#
# - b: blue
# - g: green
# - r: red
# - c: cyan
# - m: magenta
# - y: yellow
# - k: black
# - w: white
# + [markdown] slideshow={"slide_type": "slide"}
# Other colornames that are allowed are the HTML/CSS colornames such as "burlywood" and "chartreuse" are valid. See the [full list](http://www.w3schools.com/html/html_colornames.asp) of the 147 colornames. They allow "grey" where-ever "gray" appears in that list of colornames. All of these colornames are case-insensitive.
#
# ### Hex values
# Colors can also be specified by supplying an HTML/CSS hex string, such as `'#0000FF'` for blue.
#
# ### 256 Shades of Gray
# A gray level can be given instead of a color by passing a string representation of a number between 0 and 1, inclusive. `'0.0'` is black, while `'1.0'` is white. `'0.75'` would be a lighter shade of gray.
# + [markdown] slideshow={"slide_type": "slide"}
# ### RGB[A] tuples
#
# You may come upon instances where the previous ways of specifying colors do not work. This can sometimes happen in some of the deeper, stranger levels of the code. When all else fails, the universal language of colors for matplotlib is the RGB[A] tuple. This is the "Red", "Green", "Blue", and sometimes "Alpha" tuple of floats in the range of [0, 1]. One means full saturation of that channel, so a red RGBA tuple would be `(1.0, 0.0, 0.0, 1.0)`, whereas a partly transparent green RGBA tuple would be `(0.0, 1.0, 0.0, 0.75)`. The documentation will usually specify whether it accepts RGB or RGBA tuples. Sometimes, a list of tuples would be required for multiple colors, and you can even supply a Nx3 or Nx4 numpy array in such cases.
#
# In functions such as `plot()` and `scatter()`, while it may appear that they can take a color specification, what they really need is a "format specification", which includes color as part of the format. Unfortunately, such specifications are string only and so RGB[A] tuples are not supported for such arguments (but you can still pass an RGB[A] tuple for a "color" argument).
#
# Note, oftentimes there is a separate argument for "alpha" where-ever you can specify a color. The value for "alpha" will usually take precedence over the alpha value in the RGBA tuple. There is no easy way around this problem.
# + slideshow={"slide_type": "slide"}
# # %load exercises/3.1-colors.py
t = np.arange(0.0, 5.0, 0.2)
plt.plot(t, t, , t, t**2, , t, t**3, )
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Markers
# [Markers](http://matplotlib.org/api/markers_api.html) are commonly used in [`plot()`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot) and [`scatter()`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.scatter) plots, but also show up elsewhere. There is a wide set of markers available, and custom markers can even be specified.
#
# marker | description ||marker | description ||marker | description ||marker | description
# :----------|:--------------||:---------|:--------------||:---------|:--------------||:---------|:--------------
# "." | point ||"+" | plus ||"," | pixel ||"x" | cross
# "o" | circle ||"D" | diamond ||"d" | thin_diamond || |
# "8" | octagon ||"s" | square ||"p" | pentagon ||"\*" | star
# "|" | vertical line||"\_" | horizontal line ||"h" | hexagon1 ||"H" | hexagon2
# 0 | tickleft ||4 | caretleft ||"<" | triangle_left ||"3" | tri_left
# 1 | tickright ||5 | caretright ||">" | triangle_right||"4" | tri_right
# 2 | tickup ||6 | caretup ||"^" | triangle_up ||"2" | tri_up
# 3 | tickdown ||7 | caretdown ||"v" | triangle_down ||"1" | tri_down
# "None" | nothing ||`None` | nothing ||" " | nothing ||"" | nothing
# + slideshow={"slide_type": "slide"}
xs, ys = np.mgrid[:4, 9:0:-1]
markers = [".", "+", ",", "x", "o", "D", "d", "", "8", "s", "p", "*", "|", "_", "h", "H", 0, 4, "<", "3",
1, 5, ">", "4", 2, 6, "^", "2", 3, 7, "v", "1", "None", None, " ", ""]
descripts = ["point", "plus", "pixel", "cross", "circle", "diamond", "thin diamond", "",
"octagon", "square", "pentagon", "star", "vertical bar", "horizontal bar", "hexagon 1", "hexagon 2",
"tick left", "caret left", "triangle left", "tri left", "tick right", "caret right", "triangle right", "tri right",
"tick up", "caret up", "triangle up", "tri up", "tick down", "caret down", "triangle down", "tri down",
"Nothing", "Nothing", "Nothing", "Nothing"]
fig, ax = plt.subplots(1, 1, figsize=(14, 4))
for x, y, m, d in zip(xs.T.flat, ys.T.flat, markers, descripts):
ax.scatter(x, y, marker=m, s=100)
ax.text(x + 0.1, y - 0.1, d, size=14)
ax.set_axis_off()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercise 3.2
# Try out some different markers and colors
# + slideshow={"slide_type": "slide"}
# # %load exercises/3.2-markers.py
t = np.arange(0.0, 5.0, 0.2)
plt.plot(t, t, , t, t**2, , t, t**3, )
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Linestyles
# Line styles are about as commonly used as colors. There are a few predefined linestyles available to use. Note that there are some advanced techniques to specify some custom line styles. [Here](http://matplotlib.org/1.3.0/examples/lines_bars_and_markers/line_demo_dash_control.html) is an example of a custom dash pattern.
#
# linestyle | description
# -------------------|------------------------------
# '-' | solid
# '--' | dashed
# '-.' | dashdot
# ':' | dotted
# 'None' | draw nothing
# ' ' | draw nothing
# '' | draw nothing
#
# Also, don't mix up ".-" (line with dot markers) and "-." (dash-dot line) when using the ``plot`` function!
# + slideshow={"slide_type": "slide"}
fig = plt.figure()
t = np.arange(0.0, 5.0, 0.2)
plt.plot(t, t, '-', t, t**2, '--', t, t**3, '-.', t, -t, ':')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# It is a bit confusing, but the line styles mentioned above are only valid for lines. Whenever you are dealing with the linestyles of the edges of "Patch" objects, you will need to use words instead of the symbols. So "solid" instead of "-", and "dashdot" instead of "-.". This issue is be fixed for the v2.1 release and allow these specifications to be used interchangably.
# -
fig, ax = plt.subplots(1, 1)
ax.bar([1, 2, 3, 4], [10, 20, 15, 13], ls='dashed', ec='r', lw=5)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Plot attributes
# With just about any plot you can make, there are many attributes that can be modified to make the lines and markers suit your needs. Note that for many plotting functions, matplotlib will cycle the colors for each dataset you plot. However, you are free to explicitly state which colors you want used for which plots. For the [`plt.plot()`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.plot) and [`plt.scatter()`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.scatter) functions, you can mix the specification for the colors, linestyles, and markers in a single string.
# + slideshow={"slide_type": "slide"}
fig = plt.figure()
t = np.arange(0., 5., 0.2)
# red dashes, blue squares and green triangles
plt.plot(t, t, 'r--', t, t**2, 'bs', t, t**3, 'g^')
plt.show()
# -
# | Property | Value Type
# |------------------------|-------------------------------------------------
# |alpha | float
# |color or c | any matplotlib color
# |dash_capstyle | ['butt', 'round' 'projecting']
# |dash_joinstyle | ['miter' 'round' 'bevel']
# |dashes | sequence of on/off ink in points
# |drawstyle | [ ‘default’ ‘steps’ ‘steps-pre’
# | | ‘steps-mid’ ‘steps-post’ ]
# |linestyle or ls | [ '-' '--' '-.' ':' 'None' ' ' '']
# | | and any drawstyle in combination with a
# | | linestyle, e.g. 'steps--'.
# |linewidth or lw | float value in points
# |marker | [ 0 1 2 3 4 5 6 7 'o' 'd' 'D' 'h' 'H'
# | | '' 'None' ' ' `None` '8' 'p' ','
# | | '+' 'x' '.' 's' '\*' '\_' '|'
# | | '1' '2' '3' '4' 'v' '<' '>' '^' ]
# |markeredgecolor or mec | any matplotlib color
# |markeredgewidth or mew | float value in points
# |markerfacecolor or mfc | any matplotlib color
# |markersize or ms | float
# |solid_capstyle | ['butt' 'round' 'projecting']
# |solid_joinstyle | ['miter' 'round' 'bevel']
# |visible | [`True` `False`]
# |zorder | any number
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercise 3.3
#
# Make a plot that has a dotted red line, with large yellow diamond markers that have a green edge
# + slideshow={"slide_type": "slide"}
# # %load exercises/3.3-properties.py
t = np.arange(0.0, 5.0, 0.1)
a = np.exp(-t) * np.cos(2*np.pi*t)
plt.plot(t, a, )
plt.show()
# -
# # Colormaps
#
# Another very important property of many figures is the colormap. The job of a colormap is to relate a scalar value to a color. In addition to the regular portion of the colormap, an "over", "under" and "bad" color can be optionally defined as well. NaNs will trigger the "bad" part of the colormap.
#
# As we all know, we create figures in order to convey information visually to our readers. There is much care and consideration that have gone into the design of these colormaps. Your choice in which colormap to use depends on what you are displaying. In mpl, the "jet" colormap has historically been used by default, but it will often not be the colormap you would want to use.
# +
# # %load http://matplotlib.org/mpl_examples/color/colormaps_reference.py # For those with v1.2 or higher
"""
Reference for colormaps included with Matplotlib.
This reference example shows all colormaps included with Matplotlib. Note that
any colormap listed here can be reversed by appending "_r" (e.g., "pink_r").
These colormaps are divided into the following categories:
Sequential:
These colormaps are approximately monochromatic colormaps varying smoothly
between two color tones---usually from low saturation (e.g. white) to high
saturation (e.g. a bright blue). Sequential colormaps are ideal for
representing most scientific data since they show a clear progression from
low-to-high values.
Diverging:
These colormaps have a median value (usually light in color) and vary
smoothly to two different color tones at high and low values. Diverging
colormaps are ideal when your data has a median value that is significant
(e.g. 0, such that positive and negative values are represented by
different colors of the colormap).
Qualitative:
These colormaps vary rapidly in color. Qualitative colormaps are useful for
choosing a set of discrete colors. For example::
color_list = plt.cm.Set3(np.linspace(0, 1, 12))
gives a list of RGB colors that are good for plotting a series of lines on
a dark background.
Miscellaneous:
Colormaps that don't fit into the categories above.
"""
import numpy as np
import matplotlib.pyplot as plt
# Have colormaps separated into categories:
# http://matplotlib.org/examples/color/colormaps_reference.html
cmaps = [('Perceptually Uniform Sequential',
['viridis', 'inferno', 'plasma', 'magma']),
('Sequential', ['Blues', 'BuGn', 'BuPu',
'GnBu', 'Greens', 'Greys', 'Oranges', 'OrRd',
'PuBu', 'PuBuGn', 'PuRd', 'Purples', 'RdPu',
'Reds', 'YlGn', 'YlGnBu', 'YlOrBr', 'YlOrRd']),
('Sequential (2)', ['afmhot', 'autumn', 'bone', 'cool',
'copper', 'gist_heat', 'gray', 'hot',
'pink', 'spring', 'summer', 'winter']),
('Diverging', ['BrBG', 'bwr', 'coolwarm', 'PiYG', 'PRGn', 'PuOr',
'RdBu', 'RdGy', 'RdYlBu', 'RdYlGn', 'Spectral',
'seismic']),
('Qualitative', ['Accent', 'Dark2', 'Paired', 'Pastel1',
'Pastel2', 'Set1', 'Set2', 'Set3']),
('Miscellaneous', ['gist_earth', 'terrain', 'ocean', 'gist_stern',
'brg', 'CMRmap', 'cubehelix',
'gnuplot', 'gnuplot2', 'gist_ncar',
'nipy_spectral', 'jet', 'rainbow',
'gist_rainbow', 'hsv', 'flag', 'prism'])]
nrows = max(len(cmap_list) for cmap_category, cmap_list in cmaps)
gradient = np.linspace(0, 1, 256)
gradient = np.vstack((gradient, gradient))
def plot_color_gradients(cmap_category, cmap_list):
fig, axes = plt.subplots(nrows=nrows)
fig.subplots_adjust(top=0.95, bottom=0.01, left=0.2, right=0.99)
axes[0].set_title(cmap_category + ' colormaps', fontsize=14)
for ax, name in zip(axes, cmap_list):
ax.imshow(gradient, aspect='auto', cmap=plt.get_cmap(name))
pos = list(ax.get_position().bounds)
x_text = pos[0] - 0.01
y_text = pos[1] + pos[3]/2.
fig.text(x_text, y_text, name, va='center', ha='right', fontsize=10)
# Turn off *all* ticks & spines, not just the ones with colormaps.
for ax in axes:
ax.set_axis_off()
for cmap_category, cmap_list in cmaps:
plot_color_gradients(cmap_category, cmap_list)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# When colormaps are created in mpl, they get "registered" with a name. This allows one to specify a colormap to use by name.
# + slideshow={"slide_type": "slide"}
fig, (ax1, ax2) = plt.subplots(1, 2)
z = np.random.random((10, 10))
ax1.imshow(z, interpolation='none', cmap='gray')
ax2.imshow(z, interpolation='none', cmap='coolwarm')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Mathtext
# Oftentimes, you just simply need that superscript or some other math text in your labels. Matplotlib provides a very easy way to do this for those familiar with LaTeX. Any text that is surrounded by dollar signs will be treated as "[mathtext](http://matplotlib.org/users/mathtext.html#mathtext-tutorial)". Do note that because backslashes are prevelent in LaTeX, it is often a good idea to prepend an `r` to your string literal so that Python will not treat the backslashes as escape characters.
# + slideshow={"slide_type": "slide"}
fig = plt.figure()
plt.scatter([1, 2, 3, 4], [4, 3, 2, 1])
plt.title(r'$\sigma_{i=15}$', fontsize=20)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Annotations and Arrows
# There are two ways one can place arbitrary text anywhere they want on a plot. The first is a simple [`text()`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.text). Then there is the fancier [`annotate()`](http://matplotlib.org/api/pyplot_api.html#matplotlib.pyplot.annotate) function that can help "point out" what you want to annotate.
# + slideshow={"slide_type": "slide"}
fig = plt.figure()
t = np.arange(0.0, 5.0, 0.01)
s = np.cos(2*np.pi*t)
plt.plot(t, s, lw=2)
plt.annotate('local max', xy=(2, 1), xytext=(4, 1.5),
arrowprops=dict(facecolor='black', shrink=0.0))
plt.ylim(-2, 2)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# There are all sorts of boxes for your text, and arrows you can use, and there are many different ways to connect the text to the point that you want to annotate. For a complete tutorial on this topic, go to the [Annotation Guide](http://matplotlib.org/users/annotations_guide.html). In the meantime, here is a table of the kinds of arrows that can be drawn
# + slideshow={"slide_type": "slide"}
import matplotlib.patches as mpatches
styles = mpatches.ArrowStyle.get_styles()
ncol = 2
nrow = (len(styles)+1) // ncol
figheight = (nrow+0.5)
fig = plt.figure(figsize=(4.0*ncol/0.85, figheight/0.85))
fontsize = 0.4 * 70
ax = fig.add_axes([0, 0, 1, 1])
ax.set_xlim(0, 4*ncol)
ax.set_ylim(0, figheight)
def to_texstring(s):
s = s.replace("<", r"$<$")
s = s.replace(">", r"$>$")
s = s.replace("|", r"$|$")
return s
for i, (stylename, styleclass) in enumerate(sorted(styles.items())):
x = 3.2 + (i//nrow)*4
y = (figheight - 0.7 - i%nrow)
p = mpatches.Circle((x, y), 0.2, fc="w")
ax.add_patch(p)
ax.annotate(to_texstring(stylename), (x, y),
(x-1.2, y),
ha="right", va="center",
size=fontsize,
arrowprops=dict(arrowstyle=stylename,
patchB=p,
shrinkA=50,
shrinkB=5,
fc="w", ec="k",
connectionstyle="arc3,rad=-0.25",
),
bbox=dict(boxstyle="square", fc="w"))
ax.set_axis_off()
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# ## Exercise 3.4
# Point out a local minimum with a fancy red arrow.
# + slideshow={"slide_type": "slide"}
# # %load exercises/3.4-arrows.py
t = np.arange(0.0, 5.0, 0.01)
s = np.cos(2*np.pi*t)
plt.plot(t, s, lw=2)
plt.annotate('local max', xy=(2, 1), xytext=(3, 1.5),
arrowprops=dict())
plt.ylim(-2, 2)
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# # Hatches
# A Patch object can have a hatching defined for it.
#
# * / - diagonal hatching
# * \ - back diagonal
# * | - vertical
# * \- - horizontal
# * \+ - crossed
# * x - crossed diagonal
# * o - small circle
# * O - large circle (upper-case 'o')
# * . - dots
# * \* - stars
#
# Letters can be combined, in which case all the specified
# hatchings are done. If same letter repeats, it increases the
# density of hatching of that pattern.
#
# ## Ugly tie contest!
# + slideshow={"slide_type": "slide"}
fig = plt.figure()
bars = plt.bar([1, 2, 3, 4], [10, 12, 15, 17])
plt.setp(bars[0], hatch='x', facecolor='w')
plt.setp(bars[1], hatch='xx-', facecolor='orange')
plt.setp(bars[2], hatch='+O.', facecolor='c')
plt.setp(bars[3], hatch='*', facecolor='y')
plt.show()
# + [markdown] slideshow={"slide_type": "slide"}
# To learn more, please see this guide on [customizing matplotlib](http://matplotlib.org/users/customizing.html).
| course/matplotlib/3. Vocabulary.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# <img src="./intro_images/MIE.PNG" width="100%" align="left" />
# <table style="float:right;">
# <tr>
# <td>
# <div style="text-align: right"><a href="https://alandavies.netlify.com" target="_blank">Dr <NAME></a></div>
# <div style="text-align: right">Senior Lecturer Health Data Science</div>
# <div style="text-align: right">University of Manchester</div>
# </td>
# <td>
# <img src="./intro_images/alan.PNG" width="30%" />
# </td>
# </tr>
# </table>
# # 1.0 Introduction to programming with Python
# ****
# #### About this Notebook
# This notebook introduces the <code>Python</code> programming language and the <code>Jupyter</code> notebook environment.
# <div class="alert alert-block alert-warning"><b>Learning Objectives:</b>
# <br/> At the end of this notebook you will be able to:
#
# - Run code in the notebook environment
#
# - Know where to go to find answers to coding questions
#
# </div>
# <a id="top"></a>
#
# <b>Table of contents</b><br>
#
# 1.0 [Jupyter notebooks](#jupyter)<br>
# 2.0 [Learning to code](#learning)<br>
# 3.0 [Getting help](#gettinghelp)
# <a id="jupyter"></a>
# #### 1.0 Jupyter notebooks
# This series of notebooks contains details about how to program with Python. Take your time to read through them and have a go at answering the questions/tasks. The best way to learn to code is to learn by doing.
# We use several conventions used in the notebooks. Code snippets and keywords/new terms are displayed using <code>this</code> format. Each notebook is numbered and contains sub-headings that can be jumped to by clicking on the heading in the table of contents. Each notebook also starts by outlining the purpose of the book.
# <div class="alert alert-block alert-warning">
# Learning objectives are displayed in the yellow box.
# </div>
# <div class="alert alert-success">
# Green boxes are for notes and extra information. Something that might be interesting or provide additional context/information about a topic.
# </div>
# <div class="alert alert-danger">
# Red boxes are for important points that you should pay attention to.
# </div>
# <div class="alert alert-block alert-info">
# Blue boxes are for tasks. Most tasks are followed by an empty code cell or cells for you to attempt a solution to the task. Many tasks also have a <b>show solution</b> button that shows and hides the model answer. Don't worry if your solution is not exactly the same as ours. There are usually multiple ways to solve the same problem.
# </div>
# <a id="learning"></a>
# #### 2.0 Learning to code
# Learning to code is an emotional journey. There will be ups and downs. It is also important that you have realistic expectations. I will share 2 analogies with you to give some context. Learning to code is like learning to play an instrument like the guitar. We can show you what a guitar looks like, what notes the strings represent and teach you a few chords. By the end you could play a few simple tunes. You will certainly not be <a href="https://en.wikipedia.org/wiki/Jimi_Hendrix" target="_blank">Jimi Hendrix</a>. You would need to spend a lot of time practicing and developing these skills in your own time outside of lessons. Another analogy is something like carpentry. We can show you the tool kit (hammer, saw, plane etc.), we can explain how the tools work and show you with examples some of the things you can do with those tools. But again we can't show you all the possible combinations of things you could make with them. This is something you will experience over time and by working on different projects. Programming is as much an art as it is a science.
# Below is a short video that explains the <code>Jupyter notebook</code> environment and the conventions we use in the notebooks to convey certain information such as tasks, points of interest and how to execute (run) Python code in the notebooks.
# + [markdown] cell_style="center"
# The notebook lets us present you with text, images, videos and other interactive elements all in one place. Green boxes contain additional notes or extra information. The code cells allow you to input and run Python code. The blue boxes represent exercises. You can click on the <code>Show solution</code> button under an exercise to show or hide the suggested solution. To run a cell hold the <code>shift</code> key and press the <code>enter</code> button at the same time <code>shift + enter</code>.
# -
# ****
# + [markdown] cell_style="split"
# Some information about Python:
# <ul>
# <li>Developed by <NAME> in 1991</li>
# <li>Named after Monty Python (English comedians)</li>
# <li>Supports multiple programming paradigms</li>
# <li>Is open source (free)</li>
# <li>Programs are platform independent</li>
# <li>Often referred to as a ‘glue’ language</li>
# <li>One of the most popular languages for data science</li>
# </ul>
# + [markdown] cell_style="split"
# <img src="./intro_images/Guido.jpg" width="100" />
# <br>
# <div style="text-align:center"><NAME></div>
# + [markdown] cell_style="center"
# <div class="alert alert-block alert-info">
# <b>Task 1:</b>
# <br>
# Let's run our very first Python program to display the classic <code>Hello world</code> message on the screen. To do this select (click on) the cell below and hold the <code>SHIFT</code> key and press the <code>ENTER</code> key at the same time. Alternatively click on the <code>run cell</code> button on the menu above. You should see <code>Hello world</code> displayed under the cell.
# </div>
# -
print("Hello world")
# <div class="alert alert-danger">
# <b>Note:</b> You should run every cell in the notebook. Sometimes a cell will require that a previous cell or cells have been run first.
# </div>
# + [markdown] solution2="hidden" solution2_first=true
# <div class="alert alert-block alert-info">
# <b>Task 2:</b>
# <br>
# 1. Write the line of code that outputs "Hello world" above again in the cell below but this time change it to say <code>Hello Python</code><br />
# 2. Run the cell and view the output.<br />
# 3. Click on <code>Show solution</code> and compare your answer.
# </div>
# + solution2="hidden"
print("Hello Python")
# -
# <a id="gettinghelp"></a>
# #### 3.0 Getting help
# When learning to code you may run into problems when trying to solve problems. A lot of coding (programming) involves problem solving. There are often many different ways of solving a problem. Even seasoned software engineers have to look things up all the time. One way to do this is to do an internet search for the problem you are trying to solve. One useful resource is <code>Stack overflow</code> that answers many programming questions. You can post your own questions too but usually someone has already asked a similar question before. People vote on the answers.
# + [markdown] cell_style="split"
# <img src="./intro_images/overflow.PNG" width="100%" />
# + [markdown] cell_style="split"
# <img src="./intro_images/overflow1.PNG" width="100%" />
# -
# One of the most difficult aspects of this is knowing what to ask. Over time you will learn the best way to phrase your questions. You can also find answers to questions very similar to your own that you can modify for your own purposes. You can find Stack Overflow here <a href="https://stackoverflow.com/" target="_blank">https://stackoverflow.com/</a>. Other useful resources include <a href="https://realpython.com/python-first-steps/" target="_blank">Real Python</a> and Python's official website <a href="https://www.python.org/about/help/" target="_blank">Python.org</a>.
# You have completed the first notebook. Go to the next book in the sequence to learn about <code>variables</code>.
# ### Notebook details
# <br>
# <i>Notebook created by <strong>Dr. <NAME></strong>.
# <br>
# © <NAME> 2021
# ## Notes:
# + cell_style="center"
| Intro to Python/Intro to Python Book 1 (introduction).ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Transfer Training and Prediction Results
# +
from __future__ import print_function, division
import sys
import platform
import time
import os
import copy
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import json
from pprint import pprint
import torch
import torch.nn as nn
import torch.optim as optim
from torch.optim import lr_scheduler
import torch.backends.cudnn as cudnn
import torchvision
from torchvision import datasets, models, transforms
# this is necessary to use the common functions
# assumes directory structure was maintained
sys.path.insert(0, '../common/')
# from common.torch_utils import train_model,get_device
# from torch_utils import (train_model,
# mnist_dataloader,
# dataset_preview)
from torch_utils import *
# print some versions
print(f'Python Version: {platform.python_version()}')
print(f'PyTorch Version: {torch.__version__}')
print(f'Torchvision Version: {torchvision.__version__}')
print(f'CUDA Version: {torch.version.cuda}')
# get device (defaults to GPU if available)
device = get_device()
# -
# ## Data Preprocessing for Limited Dataset
# We need to collect the MNIST data and create the dataloaders for PyTorch. To make a clean notebook, we have created a helper function to do most of the work (refer to `/src/common/torch_utils.py`). For training the base model, we will use a batch size of 32.
# +
################################
# prepare the pre-trained model:
# Note the following considerations given our dataset for ResNet
# -> MNIST data are 1-channel (grascale) of size and has 10 output classes
# -> ResNet model expects 3-channel (RGB) images of size 224x224 as input
# and has 1000 output classes
# == We must changet the last fully connected layer to match 10 classes
################################
PRETRAINED = True # <-- we will be finetunning
OUTPUT_DIR='output'
BATCH_SIZE = 32
NUM_CLASSES = 10
LIMITED_DATASET_SIZE = 650
NUM_EPOCHS = 100
data_transforms = {
'train': transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5), (0.5))
]),
'val': transforms.Compose([
transforms.ToTensor(),
transforms.Normalize((0.5), (0.5))
]),
}
# use helper to create the dataloaders
tmp = mnist_dataloader(data_transforms,
batch_size=BATCH_SIZE,
sample_size=LIMITED_DATASET_SIZE,
pred_size=0.05)
dataloaders, dataset_sizes, class_names = tmp
print(f"Dataset sizes: {dataset_sizes}")
print(f"Class names: {class_names}")
# define the set to use for testing the model
inputs, labels = next(iter(dataloaders['pred']))
inputs = inputs.cuda()
# preview the dataset
dataset_preview(dataloaders['train'],'Sample MNIST Training Set')
dataset_preview(dataloaders['val'], 'Sample MNIST Validation Set')
# -
# ## Train ResNet18 on the Limited Dataset
# +
model_name = 'ResNet18'
model = models.resnet18(pretrained=PRETRAINED)
# modify the network to work with MNIST 28x28x1 with 10 outputs
num_features = model.fc.in_features # keep features unchanged
model.fc = nn.Linear(num_features, NUM_CLASSES) # output layer, 10 classes
model.conv1 = torch.nn.Conv1d(1, 64, (7, 7), (2, 2), (3, 3), bias=False) # first layer, single channel images
# use helper function to train the model (outputs model and Pandas DF)
print('\n'+'*'*40)
print(f'Training model \'{model_name}\' with limited dataset size of {LIMITED_DATASET_SIZE}...')
cudnn.benchmark = True
model, resnet18_df,_ = train_model(device, model, dataloaders, dataset_sizes,
output_dir=OUTPUT_DIR, status=10,
num_epochs=NUM_EPOCHS)
# save the data for others to use
# <-- train_model should have created this dir
results_file = f'{model_name}_size-{LIMITED_DATASET_SIZE}_results_{time.strftime("%Y-%m-%dT%H%M%S")}.csv'
resnet18_df.to_csv(os.path.join(OUTPUT_DIR,results_file),
columns=resnet18_df.columns)
print(f'> Saved results to \'{results_file}\'.')
# -
# ### Let's check Resnet18 with 2x & 10x the data
# +
# use helper to create the dataloaders
tmp = mnist_dataloader(data_transforms,
batch_size=BATCH_SIZE,
sample_size=LIMITED_DATASET_SIZE*2,
pred_size=0.05)
resnet18_1k, resnet18_1k_sizes, _ = tmp
print(f"Dataset sizes: {resnet18_1k_sizes}")
# Let's run with 10x the data!
model_name = 'ResNet18-1k'
model_1k = models.resnet18(pretrained=PRETRAINED)
# modify the network to work with MNIST 28x28x1 with 10 outputs
num_features = model_1k.fc.in_features # keep features unchanged
model_1k.fc = nn.Linear(num_features, NUM_CLASSES) # output layer, 10 classes
model_1k.conv1 = torch.nn.Conv1d(1, 64, (7, 7), (2, 2), (3, 3), bias=False) # first layer, single channel images
# use helper function to train the model (outputs model and Pandas DF)
print('\n'+'*'*40)
print(f'Training model \'{model_name}\' with limited dataset size of {LIMITED_DATASET_SIZE}...')
cudnn.benchmark = True
model_1k, resnet18_1k_df,_ = train_model(device, model_1k, resnet18_1k, resnet18_1k_sizes,
output_dir=OUTPUT_DIR, status=10,
num_epochs=NUM_EPOCHS)
# save the data for others to use
# <-- train_model should have created this dir
results_file = f'{model_name}_size-6500_results_{time.strftime("%Y-%m-%dT%H%M%S")}.csv'
resnet18_1k_df.to_csv(os.path.join(OUTPUT_DIR,results_file),
columns=resnet18_1k_df.columns)
print(f'> Saved results to \'{results_file}\'.')
# +
# use helper to create the dataloaders
tmp = mnist_dataloader(data_transforms,
batch_size=BATCH_SIZE,
sample_size=6500,
pred_size=0.05)
resnet18_6k, resnet18_6k_sizes, _ = tmp
print(f"Dataset sizes: {resnet18_6k_sizes}")
# Let's run with 10x the data!
model_name = 'ResNet18-6k'
model_6k = models.resnet18(pretrained=PRETRAINED)
# modify the network to work with MNIST 28x28x1 with 10 outputs
num_features = model_6k.fc.in_features # keep features unchanged
model_6k.fc = nn.Linear(num_features, NUM_CLASSES) # output layer, 10 classes
model_6k.conv1 = torch.nn.Conv1d(1, 64, (7, 7), (2, 2), (3, 3), bias=False) # first layer, single channel images
# use helper function to train the model (outputs model and Pandas DF)
print('\n'+'*'*40)
print(f'Training model \'{model_name}\' with limited dataset size of {LIMITED_DATASET_SIZE}...')
cudnn.benchmark = True
model_6k, resnet18_6k_df,_ = train_model(device, model_6k, resnet18_6k, resnet18_6k_sizes,
output_dir=OUTPUT_DIR, status=10,
num_epochs=NUM_EPOCHS)
# save the data for others to use
# <-- train_model should have created this dir
results_file = f'{model_name}_size-6500_results_{time.strftime("%Y-%m-%dT%H%M%S")}.csv'
resnet18_6k_df.to_csv(os.path.join(OUTPUT_DIR,results_file),
columns=resnet18_6k_df.columns)
print(f'> Saved results to \'{results_file}\'.')
# +
## Compare all of the training/validation times
# collect all model outputs
model_dict = {'resnet18': resnet18_df, 'resnet18_1k': resnet18_1k_df,'resnet18_6k': resnet18_6k_df}
dfs = []
for m in model_dict:
# do some cleanup and make a model column
df = model_dict[m].set_index('epoch').copy()
df['model'] = m
dfs.append(df)
# plot performance curves
loss_df = pd.concat(dfs).groupby(['model'])[["average_training_loss", "average_validation_loss"]]
acc_df = pd.concat(dfs).groupby(['model'])[["training_acc", "validaton_acc"]]
fig, axes = plt.subplots(nrows=3, ncols=2,figsize=(14,14))
resnet18_df.plot(y=['average_training_loss','average_validation_loss'], ax=axes[0,0])
resnet18_1k_df.plot(y=['average_training_loss','average_validation_loss'], ax=axes[1,0])
resnet18_6k_df.plot(y=['average_training_loss','average_validation_loss'], ax=axes[2,0])
axes[0,0].set_ylim([0,3])
axes[1,0].set_ylim([0,3])
axes[2,0].set_ylim([0,3])
resnet18_df.plot(y=['training_acc','validaton_acc'],ax=axes[0,1])
resnet18_1k_df.plot(y=['training_acc','validaton_acc'],ax=axes[1,1])
resnet18_6k_df.plot(y=['training_acc','validaton_acc'],ax=axes[2,1]).get_figure()
# titles and such
axes[0,0].set_title('Average Loss per Epoch')
axes[0,1].set_title('Training Accuracy per Epoch')
axes[0,0].set_ylabel('Resnet18 w/ 650 Samples (1x)')
axes[1,0].set_ylabel('Resnet18 w/ 1300 Samples (2x)')
axes[2,0].set_ylabel('Resnet18 w/ 6500 Samples (10x)')
# Save the plots
img_file = f'loss_transfer-mnist_{time.strftime("%Y-%m-%dT%H%M%S")}.png'
fig.savefig(os.path.join(OUTPUT_DIR,img_file))
# concatenetate all model results and sum up the times
time_df = pd.concat(dfs) \
.groupby(['model'])[["training_time", "validation_time"]] \
.apply(lambda x : x.astype(float).sum())
# show the times
print(time_df)
print()
# plot the times
fig2 = time_df.plot.bar(figsize=(10,8)).get_figure()
plt.title('Showing Training/Validation Time for Each Model')
# other it'll show up as 'dummy'
plt.ylabel('Time (s)')
plt.xlabel('')
plt.xticks(rotation=45)
plt.show()
# Save the plots
img_file = f'time_transfer-mnist_{time.strftime("%Y-%m-%dT%H%M%S")}.png'
fig2.savefig(os.path.join(OUTPUT_DIR,img_file))
# -
# ## Evaluate ResNet18s
# +
# Evaluate the model using the prediction set held-off
model.eval()
# make predictions an plot the results
fig = plot_classes_preds(model,inputs,labels,class_names)
plt.plot()
model_6k.eval()
# make predictions an plot the results
fig = plot_classes_preds(model_6k,inputs,labels,class_names)
plt.plot()
# -
# ### Preliminary look at training/validation time
# +
## Compare all of the training/validation times
df = resnet18_df.set_index('epoch').copy()
df['model'] = 'resnet18'
# concatenetate all model results and sum up the times
time_df = df.groupby(['model'])[["training_time", "validation_time"]] \
.apply(lambda x : x.astype(float).sum())
# show the times
print(time_df)
print()
# plot the times
fig = time_df.plot.bar(figsize=(8,6))
plt.title('Showing Total Training/Validation Time for Each Model')
# other it'll show up as 'dummy'
plt.ylabel('Time (s)')
plt.xlabel('')
plt.xticks(rotation=45)
plt.show()
# -
# ## Train ResNet50 on the Limited Dataset
# +
model_name = 'ResNet50'
model = models.resnet50(pretrained=PRETRAINED)
# modify the network to work with MNIST 28x28x1 with 10 outputs
num_features = model.fc.in_features
model.fc = nn.Linear(num_features, NUM_CLASSES)
model.conv1 = torch.nn.Conv1d(1, 64, (7, 7), (2, 2), (3, 3), bias=False)
# use helper function to train the model (outputs model and Pandas DF)
print('\n'+'*'*40)
print(f'Training model \'{model_name}\' with limited dataset size of {LIMITED_DATASET_SIZE}...')
cudnn.benchmark = True
model, resnet50_df,_ = train_model(device, model, dataloaders, dataset_sizes,
output_dir=OUTPUT_DIR, status=10,
num_epochs=NUM_EPOCHS)
# save the data for others to use
# <-- train_model should have created this dir
results_file = f'{model_name}_size-{LIMITED_DATASET_SIZE}_results_{time.strftime("%Y-%m-%dT%H%M%S")}.csv'
resnet50_df.to_csv(os.path.join(OUTPUT_DIR,results_file),
columns=resnet50_df.columns)
print(f'> Saved results to \'{results_file}\'.')
# -
# ## Evaluate ResNet50
# +
# Evaluate the model using the prediction set held-off
model.eval()
# make predictions an plot the results
fig = plot_classes_preds(model,inputs,labels,class_names)
plt.plot()
# plot performance curves
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(16,8))
resnet50_df.plot(x='epoch', y=['average_training_loss','average_validation_loss'],ax=axes[0])
axes[0].set_title('Average Loss per Epoch')
axes[0].set_ylim([0,3])
resnet50_df.plot(x='epoch', y=['training_acc','validaton_acc'],ax=axes[1])
axes[1].set_title('Training Accuract per Epoch')
img_file = f'loss_{model_name}_{time.strftime("%Y-%m-%dT%H%M%S")}.png'
fig.savefig(os.path.join(OUTPUT_DIR,img_file))
# +
## Compare all of the training/validation times
df = resnet50_df.set_index('epoch').copy()
df['model'] = 'resnet50'
# concatenetate all model results and sum up the times
time_df = df.groupby(['model'])[["training_time", "validation_time"]] \
.apply(lambda x : x.astype(float).sum())
# show the times
print(time_df)
print()
# plot the times
fig = time_df.plot.bar(figsize=(8,6))
plt.title('Showing Total Training/Validation Time for Each Model')
# other it'll show up as 'dummy'
plt.ylabel('Time (s)')
plt.xlabel('')
plt.xticks(rotation=45)
plt.show()
# -
# ## Train VGG11 on the Limited Dataset
# +
model_name = 'VGG11'
model = models.vgg11(pretrained=PRETRAINED)
# modify the network to work with MNIST 28x28x1 with 10 outputs
num_features = model.classifier[6].in_features
model.features[0] = torch.nn.Conv2d(1, 64, 3, 1, 1)
model.features = torch.nn.Sequential(*[model.features[ii] for ii in range(15)])
model.classifier = torch.nn.Sequential(*[model.classifier[jj] for jj in range(4)])
model.classifier[-1] = torch.nn.Linear(num_features, NUM_CLASSES)
# use helper function to train the model (outputs model and Pandas DF)
print('\n'+'*'*40)
print(f'Training model \'{model_name}\' with limited dataset size of {LIMITED_DATASET_SIZE}...')
cudnn.benchmark = True
model, vgg11_df,_ = train_model(device, model, dataloaders, dataset_sizes,
output_dir=OUTPUT_DIR, status=10,
num_epochs=NUM_EPOCHS)
# save the data for others to use
# <-- train_model should have created this dir
results_file = f'{model_name}_size-{LIMITED_DATASET_SIZE}_results_{time.strftime("%Y-%m-%dT%H%M%S")}.csv'
vgg11_df.to_csv(os.path.join(OUTPUT_DIR,results_file),
columns=vgg11_df.columns)
print(f'> Saved results to \'{results_file}\'.')
# -
# ## Evaluate VGG11
# +
# Evaluate the model using the prediction set held-off
model.eval()
# make predictions an plot the results
fig = plot_classes_preds(model,inputs,labels,class_names)
plt.plot()
# plot performance curves
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(16,8))
vgg11_df.plot(x='epoch', y=['average_training_loss','average_validation_loss'],ax=axes[0])
axes[0].set_title('Average Loss per Epoch')
axes[0].set_ylim([0,3])
vgg11_df.plot(x='epoch', y=['training_acc','validaton_acc'],ax=axes[1])
axes[1].set_title('Training Accuract per Epoch')
img_file = f'loss_{model_name}_{time.strftime("%Y-%m-%dT%H%M%S")}.png'
fig.savefig(os.path.join(OUTPUT_DIR,img_file))
# -
# ## Train VGG16 on the Limited Dataset
# +
model_name = 'VGG16'
model = models.vgg16(pretrained=PRETRAINED)
# modify the network to work with MNIST 28x28x1 with 10 outputs
num_features = model.classifier[6].in_features
model.features[0] = torch.nn.Conv2d(1, 64, 3, 1, 1)
model.features = torch.nn.Sequential(*[model.features[ii] for ii in range(23)])
model.classifier = torch.nn.Sequential(*[model.classifier[jj] for jj in range(4)])
model.classifier[-1] = torch.nn.Linear(num_features, NUM_CLASSES)
# use helper function to train the model (outputs model and Pandas DF)
print('\n'+'*'*40)
print(f'Training model \'{model_name}\' with limited dataset size of {LIMITED_DATASET_SIZE}...')
cudnn.benchmark = True
model, vgg16_df,_ = train_model(device, model, dataloaders, dataset_sizes,
output_dir=OUTPUT_DIR, status=10,
num_epochs=NUM_EPOCHS)
# save the data for others to use
# <-- train_model should have created this dir
results_file = f'{model_name}_size-{LIMITED_DATASET_SIZE}_results_{time.strftime("%Y-%m-%dT%H%M%S")}.csv'
vgg16_df.to_csv(os.path.join(OUTPUT_DIR,results_file),
columns=vgg16_df.columns)
print(f'> Saved results to \'{results_file}\'.')
# -
# ## Evaluate VGG16
# +
# Evaluate the model using the prediction set held-off
model.eval()
# make predictions an plot the results
fig = plot_classes_preds(model,inputs,labels,class_names)
plt.plot()
# plot performance curves
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(16,8))
vgg16_df.plot(x='epoch', y=['average_training_loss','average_validation_loss'],ax=axes[0])
axes[0].set_title('Average Loss per Epoch')
axes[0].set_ylim([0,3])
vgg16_df.plot(x='epoch', y=['training_acc','validaton_acc'],ax=axes[1])
axes[1].set_title('Training Accuract per Epoch')
img_file = f'loss_{model_name}_{time.strftime("%Y-%m-%dT%H%M%S")}.png'
fig.savefig(os.path.join(OUTPUT_DIR,img_file))
# -
# ## Train DenseNet161 on the Limited Dataset
# +
model_name = 'DenseNet161'
model = models.densenet161(pretrained=PRETRAINED)
# modify the network to work with MNIST 28x28x1 with 10 outputs
model.features.conv0 = torch.nn.Conv2d(1, 96, kernel_size=(7, 7), stride=(2, 2), padding=(3, 3), bias=False)
model.features.pool0 = torch.nn.Identity()
model.classifier = torch.nn.Linear(2208, NUM_CLASSES, bias=True)
# use helper function to train the model (outputs model and Pandas DF)
print('\n'+'*'*40)
print(f'Training model \'{model_name}\' with limited dataset size of {LIMITED_DATASET_SIZE}...')
cudnn.benchmark = True
model, densnet161_df,_ = train_model(device, model, dataloaders, dataset_sizes,
output_dir=OUTPUT_DIR, status=10,
num_epochs=NUM_EPOCHS)
# save the data for others to use
# <-- train_model should have created this dir
results_file = f'{model_name}_size-{LIMITED_DATASET_SIZE}_results_{time.strftime("%Y-%m-%dT%H%M%S")}.csv'
densnet161_df.to_csv(os.path.join(OUTPUT_DIR,results_file),
columns=densnet161_df.columns)
print(f'> Saved results to \'{results_file}\'.')
# -
# ## Evaluate DesneNet161
# +
# Evaluate the model using the prediction set held-off
model.eval()
# make predictions an plot the results
fig = plot_classes_preds(model,inputs,labels,class_names)
plt.plot()
# plot performance curves
fig, axes = plt.subplots(nrows=1, ncols=2,figsize=(16,8))
densnet161_df.plot(x='epoch', y=['average_training_loss','average_validation_loss'],ax=axes[0])
axes[0].set_title('Average Loss per Epoch')
axes[0].set_ylim([0,3])
densnet161_df.plot(x='epoch', y=['training_acc','validaton_acc'],ax=axes[1])
axes[1].set_title('Training Accuract per Epoch')
img_file = f'loss_{model_name}_{time.strftime("%Y-%m-%dT%H%M%S")}.png'
fig.savefig(os.path.join(OUTPUT_DIR,img_file))
# -
# ## Compare times from all models
# +
## Compare all of the training/validation times
# collect all model outputs
model_dict = {'resnet18': resnet18_df, 'resnet50': resnet50_df,'vgg11':vgg11_df,'vgg16':vgg16_df}
dfs = []
for model in models:
# do some cleanup and make a model column
df = model_dict[model].set_index('epoch').copy()
df['model'] = model
dfs.append(df)
# concatenetate all model results and sum up the times
time_df = pd.concat(dfs) \
.groupby(['model'])[["training_time", "validation_time"]] \
.apply(lambda x : x.astype(float).sum())
# show the times
print(time_df)
print()
# plot the times
time_df.plot.bar(figsize=(10,8))
plt.title('Showing Training/Validation Time for Each Model')
# other it'll show up as 'dummy'
plt.ylabel('Time (s)')
plt.xlabel('')
plt.xticks(rotation=45)
plt.show()
# -
| src/transfer-learning/transfer-learning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# #### Steps used:
#
# 1> Data cleansing and Wrangling
# 2> Define the metrics for which model is getting optimized.
# 3> Feature Engineering
# 4> Data Pre-processing
# 5> Feature Selection
# 6> Split the data into training and test data sets.
# 7> Model Selection
# 8> Model Validation
# 9> Interpret the result
# 10> save Model
# 11> reload model for prediction of test .csv
# 12>do data cleaning for test.csv
# 13> predict Fees
# ## Importing Dataset
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
from sklearn.linear_model import LinearRegression
from sklearn.metrics import mean_squared_error
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
data = pd.read_csv('Final_Train.csv')
data.head()
# ### This problem is Linear Regression Problem
data.shape
data.dtypes
# ## EDA Process
data.describe()
# Min - Max: As there is alot of diffrence between min-max one need to do is scaling of data.
# ### Checking null values in dataset
# +
data.isnull().sum()
# -
print("Number of Categories: ")
for ColName in data[['Qualification','Experience','Profile','Rating','Miscellaneous_Info','Place','Fees']]:
print("{} = {}".format(ColName,len(data[ColName].unique())))
# Observation: Rating, Place and Miscellaneous_Info is having null values
# ### Handling null values
data
#1. Function to replace NAN values with mode value this both rows are categorical,
#not numeric based with datatype of float or int
def impute_nan_most_frequent_category(data,ColName):
# .mode()[0] - gives first category name
most_frequent_category=data[ColName].mode()[0]
# replace nan values with most occured category
#data[ColName + "_Imputed"] = data[ColName]
#data[ColName + "_Imputed"].fillna(most_frequent_category,inplace=True)
data[ColName] = data[ColName]
data[ColName].fillna(most_frequent_category,inplace=True)
# +
#2. Call function to impute most occured category
for Columns in ['Rating','Miscellaneous_Info','Place']:
impute_nan_most_frequent_category(data,Columns)
# Display imputed result
data[['Rating','Miscellaneous_Info','Place']].head(10)
# -
data
# +
#3. Drop actual columns if we have renamed the columns
#data = data.drop(['Rating','Miscellaneous_Info','Place'], axis = 1)
# -
data
#Rechecking null values in dataset
data.isnull().sum()
# Observation: Null values are handled
# # Data Cleaning
# ## Skewness
data.skew()
#checking for outliers
data.iloc[:,:].boxplot(figsize=[20,8])
plt.show()
# Here, Fees is target variable so no need to handle skewness or outliers
data.dtypes
# Observation: Here all the columns are categorical type and target value is int but as there are all feature variable as categorical data , there is no need to hangle skewness .
# # Feature Engineering
# Can be done with Miscellaneous_Info but, no need as it seems minimum(negative) correlation in further steps
# # Data Encoding
# As the data are categorical, so need to encode the data.
from sklearn.preprocessing import OrdinalEncoder
enc = OrdinalEncoder()
for i in data.columns:
if data[i].dtypes == "object":
data[i]=enc.fit_transform(data[i].values.reshape(-1,1))
data
# # Visualization of the Data:
# For categorical data we can use: counterplot and for numerical data we can use: distplot
data.columns
import seaborn as sns
alpha = sns.countplot(x="Qualification",data=data)
print(data["Qualification"].value_counts())
import seaborn as sns
alpha = sns.countplot(x="Experience",data=data)
print(data["Experience"].value_counts())
import seaborn as sns
alpha = sns.countplot(x="Profile",data=data)
print(data["Profile"].value_counts())
import seaborn as sns
alpha = sns.countplot(x="Rating",data=data)
print(data["Rating"].value_counts())
import seaborn as sns
alpha = sns.countplot(x="Place",data=data)
print(data["Place"].value_counts())
# ### Correlation
corr_matrix_hmap=data.corr()
plt.figure(figsize=(22,20))
sns.heatmap(corr_matrix_hmap,annot=True,linewidths=0.1,fmt="0.2f")
plt.show()
corr_matrix_hmap["Fees"].sort_values(ascending=False)
# Correlations are checked on the basis of target variable, i.e Fees
# max corelated is Qualification then, Place the, Rating and so on in decresing manner
plt.figure(figsize=(10,5))
data.corr()['Fees'].sort_values(ascending=False).drop(['Fees']).plot(kind='bar',color='c')
plt.xlabel('Feature',fontsize=14)
plt.ylabel('Column with Target Name',fontsize=14)
plt.title('correlation',fontsize=18)
plt.show()
# Observation : Maximum correlation with fees is based on Qualification.
# Negative correlation with fees is Miscellaneous, Place
# Even, Experience is negatively correlated
# # Seperating Independent variable and Target Variable
# x= independent variable
x = data.iloc[:,0:-1]
x.head()
#y = target variable = Fees
y = data.iloc[:,-1]
y.head()
data.head()
x.shape
y.shape
# ## SCALING the data using Min-Max Scaler
data.describe()
# As there is alot of diffrence between MIN-MAX there need to be data to be SCALED
from sklearn.preprocessing import MinMaxScaler
mms = MinMaxScaler()
from sklearn.linear_model import LinearRegression
lr=LinearRegression()
from sklearn.metrics import r2_score
from sklearn.model_selection import train_test_split
import warnings
warnings.filterwarnings('ignore')
data=mms.fit_transform(data)
data
# # Model Training
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.30,random_state = 42)
x_train.shape
y_train.shape
x_test.shape
y_test.shape
data.shape
# ### Finding Best Random State
# +
from sklearn.linear_model import LinearRegression
maxAccu=0
maxRS=0
for i in range(1,200):
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=.30,random_state = i)
LR = LinearRegression()
LR.fit(x_train,y_train)
predrf = LR.predict(x_test)
mse = r2_score(y_test,predrf)
if mse > maxAccu:
maxAccu = mse
maxRS = i
print("Best score is: ",maxAccu,"on Random_state",maxRS)
# -
x_train,x_test,y_train,y_test=train_test_split(x,y,test_size=0.30,random_state = 25)
LR = LinearRegression()
LR.fit(x_train,y_train)
predrf = LR.predict(x_test)
print('r2 Score:',r2_score(y_test,predrf))
# +
pred_train = LR.predict(x_train)
pred_test =LR.predict(x_test)
Train_accuracy = r2_score(y_train,pred_train)
Test_accuracy = r2_score(y_test,pred_test)
maxAccu=0
maxRS=0
from sklearn.model_selection import cross_val_score
for j in range(2,16):
cv_score=cross_val_score(LR,x,y,cv=j)
cv_mean = cv_score.mean()
if cv_mean > maxAccu:
maxAccu = cv_mean
maxRS = j
print(f"At cross fold {j} cv score is {cv_mean} and accuracy score training is {Train_accuracy} and accuracy for the testing is {Test_accuracy}")
print("\n")
# -
# #### Observation: As At Fold 6, the diffrence between cross validation score and accuracy is least, will choose fold 6
# ## Regularization
# To mitigate the problem of overfitting and underfitting Regularization Methods are used: Lasso, Ridge or ElasticNet .
from sklearn.model_selection import cross_val_score
import warnings
warnings.filterwarnings('ignore')
# +
#Lasso tries to ommit coefficient value (the value which dont affect y)
from sklearn.model_selection import GridSearchCV
from sklearn.linear_model import Lasso
parameters = {'alpha':[.0001,.001,.01,.1,1,10],'random_state':list(range(0,10))}
ls=Lasso()
clf=GridSearchCV(ls,parameters)
clf.fit(x_train,y_train)
print(clf.best_params_)
# +
ls = Lasso(alpha=0.0001,random_state=0)
ls.fit(x_train,y_train)
ls.score(x_train,y_train)
pred_ls=ls.predict(x_test)
lss=r2_score(y_test,pred_ls)
lss
# +
#cross_validation_mean = cv_mean
#cross_validation_score= cv_score
cross_validation_score = cross_val_score(ls,x,y,cv=6)
cross_validation_mean = cross_validation_score.mean()
cross_validation_mean
# -
# ## Ensemble Technique
# +
from sklearn.model_selection import GridSearchCV
from sklearn.ensemble import RandomForestRegressor
parameters = {'criterion':['mse','mae'],'max_features':["auto","sqrt","log2"]}
rf = RandomForestRegressor()
clf=GridSearchCV(rf,parameters)
clf.fit(x_train,y_train)
print(clf.best_params_)
# +
rf=RandomForestRegressor(criterion="mae",max_features="sqrt")
rf.fit(x_train,y_train)
rf.score(x_train,y_train)
pred_decision=rf.predict(x_test)
rfs = r2_score(y_test,pred_decision)
print('R2 Score:',rfs*100)
rfscore=cross_val_score(rf,x,y,cv=6)
rfc=rfscore.mean()
print("Cross Validation Score:",rfc*100)
# -
# # Saving Model
import pickle
filename = "Doctor_Fees.pkl"
pickle.dump(rf,open(filename,"wb"))
loaded_model=pickle.load(open('Doctor_Fees.pkl','rb'))
result=loaded_model.score(x_test,y_test)
print(result)
conclusion = pd.DataFrame([loaded_model.predict(x_test)[:],pred_decision[:]],index=["Predicted","Original"])
conclusion
# # TEST DATASET
df = pd.read_csv('Final_Test.csv')
df
df.describe()
df.shape
df.dtypes
# # Data Cleaning
print("Number of Categories: ")
for ColName in df[['Qualification','Experience','Profile','Rating','Miscellaneous_Info','Place']]:
print("{} = {}".format(ColName,len(df[ColName].unique())))
#checking null values in dataset
df.isnull().sum()
#1. Function to replace NAN values with mode value this both rows are categorical,
#not numeric based with datatype of float or int
def impute_nan_most_frequent_category(df,ColName):
# .mode()[0] - gives first category name
most_frequent_category=df[ColName].mode()[0]
# replace nan values with most occured category
#data[ColName + "_Imputed"] = data[ColName]
#data[ColName + "_Imputed"].fillna(most_frequent_category,inplace=True)
df[ColName] = df[ColName]
df[ColName].fillna(most_frequent_category,inplace=True)
# +
#2. Call function to impute most occured category
for Columns in ['Rating','Miscellaneous_Info','Place']:
impute_nan_most_frequent_category(df,Columns)
# Display imputed result
df[['Rating','Miscellaneous_Info','Place']].head(10)
# -
df
#Rechecking null values in dataset
df.isnull().sum()
from sklearn.preprocessing import OrdinalEncoder
enc = OrdinalEncoder()
for i in df.columns:
if df[i].dtypes == "object":
df[i]=enc.fit_transform(df[i].values.reshape(-1,1))
df
corr_matrix_hmap=df.corr()
plt.figure(figsize=(22,20))
sns.heatmap(corr_matrix_hmap,annot=True,linewidths=0.1,fmt="0.2f")
plt.show()
test_data = df.iloc[:,0:]
test_data.head()
# # Load saved model
loaded_model=pickle.load(open('Doctor_Fees.pkl','rb'))
result=loaded_model.score(x_test,y_test)
print(result)
conclusion = pd.DataFrame([loaded_model.predict(test_data)[:],pred_decision[:]],index=["Predicted","Original"])
conclusion
fees = pd.DataFrame([loaded_model.predict(test_data)[:]])
fees
df.columns
df.shape
df.insert(6, " Fees", fees.values.reshape(-1,1))
df
# #### so Fees in the above df are the predicted values for the test data set
| Doctor_Fees .ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="Q25FKmGohQgP"
# #### Problem 1:
# -
# In this task you will do some basic datafile manipulation to learn how to import and work with .csv files. Please have a quick look at the "solar_temp_dataset.csv" to see the format of the file, it contains the history of temperature and sunlight radiation in London over the past two weeks.
# Import `pandas` as ```pd``` and use the function ```pd.read_csv()``` with the appropriate settings for `delimiter`, `parse_dates`, and `skiprows` to import the datafile and save it in a variable named "dataset". If you are not familiar with these settings, read the pandas online documentation which you can find by searching the command on google.
# + colab={} colab_type="code" id="VDqUM5oxswW0"
# Write your solution here
# -
# Print out the `.head()` function on your variable to see if the format of the imported file suits your needs. Google how to pick out separate columns from the pandas dataframe object, and see if they behave as numpy arrays.
# +
# Write your solution here
# -
# In one plot, plot the temperature and radiation as a function of time. Is there an obvious correlation between these two variables, would you expect it to be there?
# +
# Write your solution here
# -
# Plot a scale for the temperature on the left vertical axis of the plot, and a scale for the radiation on the right vertical axis of the plot. Make sure to also include vertical labels, appropriately labeled horizontal time steps, and a legend. Try to make the plot as publishable as possible.
# + [markdown] colab_type="text" id="HG0NNPZhhQgR"
# #### Problem 2:
# -
# Use the temperature data from the previous problem, and the `boxplot` function included in the `matplotlib` library to plot the temperature distribution. Interpret the various lines in the boxplot. Plot the same data as a histogram in another subplot, how do the respective features of these two plots correspond, when do you think it's useful to use boxplots instead of histograms?
# + colab={} colab_type="code" id="f_LnTfuDswW6"
# Write your solution here
# + [markdown] colab_type="text" id="bmQBIM1xswW9"
# #### Problem 3
# -
# Create a numpy array with 200 linearly rising points, that follow the linear equation $y = 3.14x + 2.718$. Add gaussian noise with `mean = 0` and `std = 100` to the data. Plot the resulting $y$ data as a function of $x$.
#
# _Hint:_
# Look at `np.linspace` and `np.random`
# + colab={} colab_type="code" id="k8QaYfaNswW_"
# Write your solution here
| notebooks/workshop-1/probleem_sheet.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
import matplotlib.pyplot as plt
import pandas as pd
import matplotlib
import urllib.request # request library for downloading a url
import os.path
# -
# %matplotlib inline
matplotlib.rcParams['figure.figsize'] = (10, 8) # set default figure size, 8in by 6in
# # Example Download of UCI Data Set
#
# Here is an example of downloading a file from an internet URL address, then loading it into
# a pandas dataframe.
# create a report hook function, so that the urlretrieve() can display
# a status report while downloading
def urlretrieve_reporthook(block_number, read_size, total_file_size):
if block_number % 100 == 0:
print("\rReading %d / %d complete" % (read_size * block_number, total_file_size), end="")
# +
# the UCI datasets have been pre-divided into test and training sets
test_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/00421/aps_failure_test_set.csv'
test_file = './data/aps-failure-test-set.csv'
train_url = 'https://archive.ics.uci.edu/ml/machine-learning-databases/00421/aps_failure_training_set.csv'
train_file = './data/aps-failure-train-set.csv'
# download the training data csv (comma separated values) file into our data folder
# I picked a relatively large dataset file/example here (45 MB), so this may take a bit of time to
# download on a slow connection.
# always good to check and only download if we don't already have the file, so we can more easily
# rerun all cells without causing a long download to be done every time
if not os.path.exists(train_file):
print('Beginning file download with urllib2...')
urllib.request.urlretrieve(train_url, train_file, reporthook=urlretrieve_reporthook)
# -
# load the csv file into a pandas dataframe
# the train file we receive has 20 lines of copyright/header information we need to skip over
# also the csv file uses na to represent missing data, which is not interpreted as a missing by
# pandas by default. By specifying this as a na_values, all of the columnes are interpreted
# as numberic types and NaN are the numeric values given to the missing data.
train = pd.read_csv(train_file, skiprows=20, na_values=['na'])
# show some information about the data
num_samples, num_features = train.shape
print("Number of features:", num_features)
print("number of training samples:", num_samples)
# +
# If we correctly interpret 'na' as missing data, we get 170 of the 171 columns interpreted as numeric.
print(train.dtypes)
# The count for describe shows the total present values for each feature out of the 60000 samples for each one.
train.describe()
# -
# the first column is actually the label/target we would want to use if we were to build a classifier.
# The values are 'neg' and 'pos'
np.unique(train['class'])
# we would want to remove this column from the training data, and create a y (training labels), that uses 0 for
# the 'neg' class, and 1 for the 'pos' class, for scikit-learn training.
# slice columns 1 to 171, which are the training features
X_train = train.iloc[:,1:]
print(X_train.shape)
# +
# slice column 0 which are the training labels
y_train = train.iloc[:,0]
print(y_train.shape)
print(y_train.dtype)
# make numeric by translating 'neg' to 0 and 'pos' to 1
mapping = {'neg': 0, 'pos': 1}
y_train.replace(mapping, inplace=True)
print(y_train)
| assignments/Assg-07-Data-Analytics-Project.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/Francisco-Dan/daa_2021_/blob/master/02_Octubre.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="OV9jygaVigto"
# ## Meteorología en México
# En Sistema meteorológico nacional lleva el registro de la lluvias desde el año 1985 y lo pone a disposición de la población por medio de la pagina datos.gob.mx.
#
# En la siguiente liga se encuentran 2 archivos separados por comas CSV correspondientes a los registros de lluvias
# mensuales y anuales de los años 2017 y 2018. En los columnas se encuentran 13, correspondientes al promedio mensual y el promedio anual.
# En los renglones se encuentran 33, correspondientes a cada uno de los 32 estados y a nivel nacional.
#
# https://drive.google.com/file/d/1lamkxgq2AsXRu81Y4JTNXLVld4og7nxt/view?usp=sharing
#
#
# ## Planteamiento del problema
# Diseñar un algoritmo y programarlo para que:
# 1. Solicite por teclado el año, el estado y el mes, en base a esa información:
# - muestre en pantalla el promedio de ese mes en ese estado en el año seleccionado.
# - muestre en pantalla el promedio anual del estado seleccionado.
# - muestre la suma de los 12 meses de ese estado en el año seleccionado.
#
# 2. Busque el mes que mas llovió en todos los estados durante esos dos años. Imprimir año, estado y mes.
# 3. Busque el mes que menos llovió en los dos. Imprimir año, estado y mes.
# + id="GY48t22wiMHc"
from 2017.py import Pre2017
| 02_Octubre.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Microstructure Tutorial
#
# by <NAME>, School of Earth Sciences and Byrd Polar & Climate Research Center <EMAIL>
#
# ## Learning Objectives
#
# At the end of this tutorial you should be able to...
#
# * Explain why microstructure is important for remote sensing
# * Define measures of microstructure, especially specific surface area
# * Access and visualize tree different microstructure measurements from SnowEx Grand Mesa 2020
#
# ## Acknowledgments
#
# Contributions from: <NAME>, <NAME>, <NAME>, <NAME>, <NAME>, <NAME>. This relies heavily on the snowexsqul database and example scripts created by <NAME>.
#
# ## Caveats
#
# The integrating sphere and the SMP data are published at NSIDC; you can read the pages there for documentation etc. However the microCT data are not yet published; please contact Lauren Farnsworth (<EMAIL>) for questions on the CT data.
#
#
# ## Fun Facts About Snow Microstructure
#
# Snow microstructure plays a super important role in snow physics and snow remote sensing, so a lot of effort went towards measuring it in SnowEx 2020!
#
# There are several different quantities that are used to measure snow microstructure, including "grain size". Grain size measurements are challenging to make in a repeatable way, and are also challenging to relate to the physical quantities that control remote sensing measurements. In the last ~15 years or so, a lot of effort has gone into more objective ways to measure microstructure.
#
# Snow microstructure governs response of remote sensing to snow cover for visible, near-infrared and high-frequency microwave wavelengths. See Figure 1, below, and read {cite:p}`Dozier2009`, for more information.
#
# <img src="images/dozier2009_fig2.jpg" alt="dozier-figure" width="400px">
#
# <b>Snow microstructure governs visible and near-infrared reflectance. This is figure 2 from {cite:p}`Dozier2009`</b>
#
# Radar measurements such as those made by the Ku-band SWEARR instrument are also very sensitive to snow microstructure.
#
# <img src="images/modeled-swe-L-response.png" alt="radar-figure" width="400px">
#
# <b>Modeled response of radar backscatter to SWE and single-scatter albedo (which in turn is a function of snow microstructure), based on a simple model suggested by {cite:p}`Ulaby2014`</b>
#
# Snow microstructure is super important to efforts to launch a Ku-band SAR to measure global snow water equivalent (SWE). An important area of research right now is exploring how to use estimates of microstructure (e.g. from snowpack evolution models) to improve SWE retrievals.
#
# Snow microstructure evolves through the season, and varies a lot with depth. Snow microstructure evolution is controlled by other snow properties, such as snow temperature, snow height and snow liquid water content. A really great resource on snow microstructure is <NAME>'s recent talks:
# * [Snow Metamorphism](https://boisestate.hosted.panopto.com/Panopto/Pages/Viewer.aspx?id=9a051fe4-db09-4f1d-92db-acd1003cc5cc)
# * [Snow Grain Identification](https://boisestate.hosted.panopto.com/Panopto/Pages/Embed.aspx?id=136f7580-c1e2-4c55-ab33-acca003c23d6)
# ## SnowEx Microstructure Measurement Background
#
# ### Basic Microstructure Definitions
#
# Microstructure definitions take a bit of getting used to. It's very easy to get confused. Specific surface area (SSA) is one of the most important quantity used to measure snow microstructure, so that's the focus here. Note that SSA is not the be-all and end-all, so there's a short table describing how to relate SSA to other quantities just below. A couple of good reads on all of this is {cite:p}`Matzler2002` and {cite:p}`Matzl2010`.
#
#
# <img src="images/lowe_etal_2011_v2.png" alt="lowe-figure" width="400px">
#
# <b>Coarse and fine snow microstructure revealed by microCT. The microCT snow renderings on the left are Figure 2 from {cite:p}`Lowe2011`. The colorbars indicate that fine-grained snow (a) has high SSA and low D<sub>eq</sub>, whereas coarse-grained snow (b) has low SSA and high D<sub>eq</sub>.</b>
#
# Use Figure 3 above to ground these definitions: SSA is the surface area of the ice-air interface, normalized in some way. Confusingly, SSA is defined in a couple of different ways in the literature: sometimes, surface area within a particular volume of interest (VOI) is normalized by the mass of the ice in the VOI. As defined in this way, SSA has units of length squared per mass, usually expressed as m<sup>2</sup>/kg. Instead of normalizing by mass, SSA is sometimes defined by normalizing by the volume of the VOI (this is SSA<sub>v</sub> in {cite:p}`Matzl2010`), and sometimes by normalizing by the volume of the ice in the VOI (this is SSA<sub>i</sub> in {cite:p}`Matzl2010`, and q in {cite:p}`Matzler2002`). Here let's just go with the first definition I mentioned:
#
# $$
# SSA = \frac{\text{Surface area of ice-air interface}}{\text{Mass of ice}} \quad
# $$
#
# SSA tends to take values between 5 and 150 m<sup>2</sup>/kg: fresh, fine-grained snow has high SSA, and coarse snow has low SSA. Because it takes a little while for SSA values to become intuitive, a useful derived metric is the equivalent grain diameter (D<sub>eq</sub>; note that this is identical to D<sub>q</sub> in {cite:p}`Matzler2002`), which by definition is the diameter that a sphere would have if it had a particular value of SSA. This is a one-to-one relationship, so there are no assumptions involved.
#
# $$
# D_{eq} = \frac{6}{SSA \rho_i} \quad
# $$
# Relationships of specific surface area to other metrics are given in this list if you're curious but otherwise just skip past this bit
# * Sometimes people refer to the "optical grain diameter", which is the same as D<sub>eq</sub>. The "optical" refers to {cite:p}`Grenfell1999`, who showed that any snow with a particular SSA had similar (not identical) radiative transfer properties regardless of particle shape in the visible and near-infrared parts of the spectrum. But note the same is not true in the microwave spectrum.
# * Autocorrelation length is usually one of two metrics that summarize the two-point microstructure autocorelation function of the three-dimensional ice-air matrix. Think of the probability that you change media (from ice to air or vice versa) as you move a certain distance within the snow microstructure. The length that defines the likelihood that you'll change media is (an approximation of the correlation length). SSA is by definition (with almost no assumptions) equal to the slope of the autocorrelation function at the origin. But microwave scattering is controlled by correlations at longer lags. For more check out {cite:p}`Matzler2002`. The lack of closing the loop between SSA and correlation length is a significant issue when we have measurements of SSA and microwaves as we do in SnowEx.
# * Geometric grain size is what we usually measure when we measure with a hand lens. You can try to relate it to SSA or corelation length, but it is not always possible, and will change with different observers.
#
# Time to stop this list but there are many other metrics as well.
# ### Microstructure Instruments
#
# Now that we know what we're trying to measure (SSA, or correlation length) how do we actually measure? let's talk just about three techniques used in SnowEx 2020 Grand Mesa.
#
# <img src="images/microCT.png" alt="ct-figure" width="800px">
#
# <b>Left: <NAME> transports microCT samples from field sites back to Grand Mesa Lodge in a cold storage container. Right: the microCT machine in the lab at CRREL.</b>
#
# Micro-computed tomography (microCT) is the only laboratory-based method used here, and it is the gold standard, although it does still come with caveats. The idea of microCT is to remove a sample of snow from a snow pit face, and either cast it with a compound such as diethyl pthalate that is still a liquid at 0° C, or preserve the same at a very cold temperature. Then the sample is sent back to the laboratory, and bombared with x-rays, similar to how you get x-rays to see if a bone is broken at the doctor. For much more on microCT, check out {cite:p}`Heggli2011`. microCT can be used to extract a ton of information about snow microstructure, including SSA, correlation length and many others.
#
# <img src="images/IntegratingSpheres.png" alt="ct-figure" width="800px">
#
# <b>Left: <NAME> operates an IceCube unit at the Grand Mesa Lodge intercomparison snowpit. Top right: schematic showing the integrating sphere measurement principle, from {cite:p}`Gallet2009`. Bottom right: snow in the IceCube sampling container, from {cite:p}`Leppanen2018`. </b>
#
# Integrating spheres are field-based and you make the measurements on samples extracted from the snowpit face. The principle of the measurement is based on firing a laser at the snow sample, within a special reflective hollow sphere, where one side is filled by the snow sample, and measuring how much of the laser is reflected at a sensor at a known geometry. For more information, check out {cite:p}`Gallet2009`. Most integrating sphere measurements are either made by a commercial firm (A2 Photonics) known as the [IceCube](https://a2photonicsensors.com/icecube-ssa/) or a version constructed at the University of Sherbrooke known as the IRIS {cite:p}`Montpetit2012`. These approaches are set up to measure SSA only. There were three of these at Grand Mesa - one of the Sherbrooke IRIS units, and two IceCubes, one from Finnish Meteorological Institute, and one from Ohio State University.
#
# <img src="images/SMP.png" alt="ct-figure" width="800px">
#
# <b>Left: Megan Mason operates the SMP. Right: Closeup of the SMP sensor tip.</b>
#
# Snow micropenetrometers are also a field-based approach, but they do not require a snowpit, enabling far more observations to be made. Instead, an automated motor pushes a probe vertically downwards into the snowpack. The probe measures the force required to break snow microstructure, yielding a wealth of information. Snow density, specific surface area and correlation length can be retrieved; for background see {cite:p}`Lowe2012` and {cite:p}`Proksch2015`. The micropen effort at Grand Mesa was led by Boise State University. A key thing to be aware of is that differences in the various SMP instruments mean that the empirical relationship of {cite:p}`Proksch2015` will give quite poor results for the particular instrument used in SnowEx, as fully explained in {cite:p}`Calonne2020`.
#
# These methods are not the only ways to measure microstructure! There are several others not mentioned here, but not used at Grand Mesa 2020. Ask if interested.
#
# ## SnowEx Microstructure Measurement Data Overview
#
# Of the three methods described above, microCT is by far the most expensive and most time consuming. Samples have to be transported back to the laboratory and the processing time requires a microCT machine. Thus the fewest CT sapmles are taken.
#
# The integrating spheres require a snowpit to be dug, so we have an intermediate number of them: ~100.
#
# The micropen measurements are by far the fastest to make, so a cross pattern of SMP measurements was made on orthogonal directions intersecting at the snowpit. There are thousands of SMP profiles from Grand Mesa 2020.
# ## Working with the data
#
# We're going to do two things! First, we'll intercompare the three different integrating sphere instruments at four different pits where we had multiple instruments operating. We'd expect these data to be fairly self-consistent. Second, we'll compare all three methods (integrating sphere, SMP and microCT) at a single pit where we had all of these measurements present. Here especially with the SMP we would expect to need to intercalibrate the data to match local conditions; so far SSA has only been fit to SMP force measurements in one study, and we should assume we'll need a local calibration to get a tight fit.
# ### 0. Load needed modules
# +
# Modules needed to access snowexsql: SnowEx field data database
from snowexsql.db import get_db
from snowexsql.data import LayerData, PointData
from snowexsql.conversions import points_to_geopandas, query_to_geopandas
# Modules needed to work with data
import geoalchemy2.functions as gfunc
import geopandas as gpd
import matplotlib.pyplot as plt
import pandas as pd
import numpy as np
import warnings
warnings.filterwarnings('ignore')
#note - this cell does not return any output
# -
# ### 1. Intercompare Integrating Sphere Datasets
# There were three integrating spheres. The IRIS unit from University of Sherbrooke was operated by <NAME>. The IceCube unit from the Finnish Meteorological Institute was operated by <NAME>. And the IceCube unit from Ohio State was operated by <NAME> and <NAME>. Carefully read the [documentation page](https://nsidc.org/data/SNEX20_SSA/versions/1) at NSIDC if you are interested in the data. If you are using the data for a project, please contact the authors and mention what you're doing - they'll appreciate it! Contact for SSA is <NAME> (<EMAIL>).
#
# See Micah's tutorial on datasets for more on this! Won't explain too much here
# +
db_name = 'snow:hackweek@192.168.127.124/snowex'
engine, session = get_db(db_name)
# Grab all the equivalent diameter profiles
q = session.query(LayerData).filter(LayerData.type == 'specific_surface_area')
df = query_to_geopandas(q, engine)
# End our database session to avoid hanging transactions
session.close()
df.head() #check out the results of the query
# -
# Since we want to intercompare integrating spheres, we need to isolate only the sites that actually had multiple integrating spheres measuring the same snow.
# +
# Grab all the sites with equivalent diameter data (set reduces a list to only its unique entries)
sites = df['site_id'].unique()
# Store all site names that have mulitple SSA instruments
multi_instr_sites = []
instruments = []
for site in sites:
# Grab all the layers associated to this site
site_data = df.loc[df['site_id'] == site]
# Do a set on all the instruments used here
instruments_used = site_data['instrument'].unique()
if len(instruments_used) > 1:
multi_instr_sites.append(site)
# Get a unqique list of SSA instruments that were colocated
instruments = df['instrument'].unique()
instruments #check out the list of instruments. note that the IceCube values are displayed as serial numbers
# -
# Finally, plot all Integrating Sphere SSA profiles at all Multi-Integrating Sphere Sites
# +
# Setup the subplot for each site for each instrument
fig, axes = plt.subplots(1, len(multi_instr_sites), figsize=(4*len(multi_instr_sites), 8))
# Establish plot colors unique to the instrument
c = ['k', 'm', 'c']
colors = {inst:c[i] for i,inst in enumerate(instruments)}
# Loop over all the multi-instrument sites
for i, site in enumerate(multi_instr_sites):
# Grab the plot for this site
ax = axes[i]
# Loop over all the instruments at this site
for instr in instruments:
# Grab our profile by site and instrument
ind = df['site_id'] == site
ind2 = df['instrument'] == instr
profile = df.loc[ind & ind2].copy()
# Don't plot it unless there is data
if len(profile.index) > 0:
# Sort by depth so samples that are take out of order won't mess up the plot
profile = profile.sort_values(by='depth')
# Layer profiles are always stored as strings.
profile['value'] = profile['value'].astype(float)
# Plot our profile
ax.plot(profile['value'], profile['depth'], colors[instr], label=instr)
# Labeling and plot style choices
ax.legend()
ax.set_xlabel('SSA [m^2/kg]')
ax.set_ylabel('Height above snow-soil interface [cm]')
ax.set_title('Site {}'.format(site.upper()))
# Set the x limits to show more detail
ax.set_xlim((8, 75))
plt.tight_layout()
plt.show()
# -
# ### 2. Pull the snowmicropenetrometer data and compute SSA
#
# The next step is to grab some SMP data to compare to. We're going to get the SMP at site 2N13, where we have a copule of SSA profiles from interating spheres (as well as microCT data, to be looked at in the next step!).
#
# The SMP measurements for SnowEx 2020 GrandMesa were all made by <NAME>. If you're interested in working with the SMP data, please carefully read the NSIDC [documentation page](https://nsidc.org/data/SNEX20_SMP/versions/1). If you're planning to work with the data, please reach out to the author; the contact is (<NAME> <EMAIL>). If you use a profile, consider checking out the comments which are described in the [Excel sheet linked from the Technical References part of the NSIDC documentation](https://nsidc.org/sites/nsidc.org/files/technical-references/SNEX20_SMP_FieldNotes.xlsx), where there are some really useful comments.
#
# There are a few steps here, and one reason for that is just that the SMP data is quite large, and so the full-resolution SMP could not be included in Micah's database. The full resolution profile from SMP is resolved ever 1.25 mm! Instead, the SMP data in Micah's database is sampled to only every 100th datapoint, so it's every 12.5 cm. But the database is still very useful! What we'll do is use the database to find the right profile, then go and download that full resolution dataset from the NSIDC. Easy-peasey!
#
# As mentioned above, {cite:p}`Calonne2020` tested applying the relationship of {cite:p}`Proksch2015` and got quite poor results, explained them by the difference in hardware between generations of SMP instruments. We were unaware of that when designing the tutorial, and so set up use of the so-called official SMP processing repository, linked below, which has not yet been updated with the latest relationship. This would make a great project, as mentioned later!
#
# First up, we'll visualize the location of the SMP profiles, along with the snowpit location.
# +
site = '2N13'
engine_smp, session_smp = get_db(db_name)
q_smp = session_smp.query(LayerData).filter(LayerData.type == 'force').filter(LayerData.site_id.contains(site) )
df_smp = query_to_geopandas(q_smp, engine_smp)
q_pit=session_smp.query(LayerData).filter(LayerData.type == 'hand_hardness').filter(LayerData.site_id.contains(site) )
df_pit = query_to_geopandas(q_pit, engine_smp)
session_smp.close()
# Plot SMP profile locations with colored by the time they were taken using upside down triangles
ax = df_smp.plot(column='time', cmap='jet', marker='v', label='SMP', figsize=(5,5), markersize=100, edgecolor='black')
ax.plot(df_pit.easting, df_pit.northing, color='black', marker='s', markersize=15, label='Pit ({})'.format(site))
# Add important labels
ax.set_xlabel('Easting [m]')
ax.set_ylabel('Northing [m]')
plt.suptitle('SMP Locations at Site {} Showing Acquisition Order'.format(site), fontsize=16)
# Avoid using Scientific notation for coords.
ax.ticklabel_format(style='plain', useOffset=False)
ax.legend()
# plt.tight_layout()
plt.show()
# -
# Next up, let's find the closest SMP profile to the snowpit, and then find the profile ID of that profile, which is in the comments in the database.
# +
# find closest SMP profile to the pit
# No profile is taken at the same time, so we grab all the unique times and sort them
times = sorted(df_smp['time'].unique())
nprofiles=len(times)
ids=np.empty(nprofiles)
p=0
for t in times:
ind = df_smp['time'] == t
data = df_smp.loc[ind].copy()
ids[p]=data.iloc[0].id
p+=1
i_dists=df_smp['id'].isin(ids)
df_smp_dists=df_smp.loc[i_dists]
df_smp_dists=df_smp_dists.assign(dists=-1)
df_smp_dists['dists']=np.sqrt((df_smp_dists['easting']-df_pit.iloc[0].easting)**2+(df_smp_dists['northing']-df_pit.iloc[0].northing)**2)
df_smp_dists.sort_values(by='dists')[['comments','dists']].head() #check out the list of profiles sorted by distance to pit
# -
# So the id of the closest SMP profile is S19M1174. I went to the [SMP page on NSIDC](https://nsidc.org/data/SNEX20_SMP/versions/1), and went to "Download" and searched for this ID, downloaded the profile, and then re-uploaded to my home directory here in the Jupyter hub.
#
# Ok next up, we have to compute SSA from the SMP data. For this, we'll use the "snowmicropyn" modules created by the Swiss SLF. You can read more about them [at this site](https://snowmicropyn.readthedocs.io/en/latest/). The software is a little out of date on Python versions; just ignore any warnings that pop up below! Also, the use of the {cite:p}`Proksch2015` relationship is also out-of-date, as mentioned above. Getting it updated for use with this tutorial would make a perfect project!
#
# The next cell pulls in the needed modules, and then plots the profile of force measurements needed to break through the snow microstructure.
from snowmicropyn import Profile
from snowmicropyn import proksch2015
# Pull in some tutorial datasets
# !aws s3 sync --quiet s3://snowex-data/tutorial-data/microstructure/ /tmp/microstructure
p = Profile.load('/tmp/microstructure/SMP/SNEX20_SMP_S19M1174_2N13_20200206.PNT',)
plt.plot(p.samples.distance, p.samples.force)
# Prettify our plot a bit
plt.title(p.name)
plt.ylabel('Force [N]')
plt.xlabel('Depth [mm]')
plt.show()
# The next step is the actual calculation of SSA from the force data. It then displays the data and lets you see that there is now a column called SSA! Note that this function is "proksch2015". You can read about how it works in Martin Proksch's paper {cite:p}`Proksch2015`.
# the window size is a parameter you can play with. default is 2.5 mm setting to 10 mm does some smoothing
p2015 = proksch2015.calc(p.samples,window=10)
p2015.head() #check out the first few values of SSA
# ### 3. Read microCT data, and compare intergrating sphere, SMP and CT data
# The microCT samples were extracted in the field and processed at CRREL by <NAME>, and is not yet published at NSIDC. Please contact her with questions (<EMAIL>)!
#
# This module reads in microCT datafiles which are stored as text. Some additional data are available, showing the computer generated slices through the ice-air interface: contact Mike (<EMAIL>) if you want to look at a subset of these data that Lauren has shared.
#
# Equivalent grain size is a useful quantity to compare: because it's proportional to 1/SSA, and because after a point as you increase SSA more and more, all fine-grained snow acts more-or-less the same (converging to e.g. the "fine-grained" curve in Figure 1, above), we'll look at equivalent diameter instead of SSA in this comparison.
from read_CT_txt_files import read_CT_txt_files
# +
# read micro CT for 2N13
data_dir='/tmp/microstructure/microCT/txt/'
[SSA_CT,height_min,height_max]=read_CT_txt_files(data_dir)
SSA_CT #chck out the SSA values read in from MicroCT
# +
# get data integrating sphere data for 2N13 and plot it
site='2N13'
engine_is, session_is = get_db(db_name)
q_is = session_is.query(LayerData).filter(LayerData.type == 'specific_surface_area').filter(LayerData.site_id.contains(site) )
df_is = query_to_geopandas(q_is, engine_is)
instruments_site = df_is['instrument'].unique()
# Loop over all the integrating sphere instruments at this site. plot equivalent diameter
fig,ax = plt.subplots()
for instr in instruments_site:
# Grab our profile by site and instrument
ind = df['site_id'] == site
ind2 = df['instrument'] == instr
profile = df.loc[ind & ind2].copy()
# Don't plot it unless there is data
if len(profile.index) > 0:
# Sort by depth so samples that are take out of order won't mess up the plot
profile = profile.sort_values(by='depth')
# Layer profiles are always stored as strings.
profile['value'] = 6/917/profile['value'].astype(float)*1000
# Plot our profile
ax.plot(profile['value'], profile['depth'], colors[instr], label=instr)
#All that's left to do is plot the CT and the SMP and label the plot!
ax.plot(6/917/SSA_CT*1000,height_min,label='microCT') #CT data
ax.plot(6/917/p2015.P2015_ssa*1000,(max(p2015.distance)-p2015.distance)/10,label='SMP') #SMP data
# Labeling and plot style choices
ax.legend()
ax.set_xlabel('Equivalent diameter, mm')
ax.set_ylabel('Height above snow-soil interface [cm]')
ax.set_title('Site {}'.format(site.upper()))
plt.tight_layout()
plt.show()
# -
# Wow, so the datasets are so very different, with the SMP being by far the most different. Comparing with {cite:p}`Calonne2020` shows that the SMP is off in the same direction as diagnosed in that paper. Thus, the difference is most likely due to the difference in SMP instruments. Dr. <NAME> of Northumbria University has a github branch of the SMP software SnowMicropyn that has the newer fit relationship integrated in the software. It would be a nice project to loop in Mel's branch with this notebook and see how well things compare to SnowEx data. I'd be happy to help anyone interested get rolling on that!
#
# There's also significant differences between the microCT and the two integrating spheres. This is science - sometimes when we start intercomparing these quantities, we do not get a perfect match. This would also be a fascinating thing to explore in a Hackweek project.
# Some of the ways that you could imagine connecting microstructure measurements to other quantities would be with the SWESARR radar data. Although the radar data does seem to have some orthorectification issues that haven't been fully worked out, I can imagine these being worked around by careful choice of places you match up the microstructure to the radar. Note that places that are shallower tend to have larger D<sub>eq</sub> and vice versa, and the spatial variability in SSA was fairly low in general in Grand Mesa 2020, so looking at multiple SSA vs radar samples might not yield a great correlation. But you never know, could be fun to try! Generaly speaking, we don't expect a ton of impact of the microstructure on L-band (UAVSAR), but it would be interesting to explore that.
#
# One thing that could be of great value is to calibrate the SMP estimates of SSA to the integrating spheres. If you're interested in doing that, do reach out first. This could be a really interesting thing to explore!
#
# It might also be interesting to compare the data to hand hardness measured in the snowpit, and to traditional hand lens measurements.
| book/tutorials/microstructure/microstructure-tutorial.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# default_exp classification
# -
# # Auto ML Classidfication
#
# > Automatically trains Models for Classification
#hide
from nbdev.showdoc import *
# +
#export
import streamlit as st
import streamlit.components.v1 as components
from pdpbox import pdp
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
import shap
# load JS visualization code to notebook
shap.initjs()
import base64
from sklearn.model_selection import train_test_split
from sklearn.datasets import load_iris, load_digits
#Simple Classifiers
from sklearn.naive_bayes import GaussianNB
from sklearn.linear_model import LogisticRegression
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
#Tree based Classifier
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier
from sklearn.ensemble import GradientBoostingClassifier
from sklearn.ensemble import ExtraTreesClassifier
from xgboost import XGBClassifier
#Gradient Based Classifier
from sklearn.linear_model import SGDClassifier
from sklearn.neural_network import MLPClassifier
#Preprocessing packages
from sklearn.compose import ColumnTransformer
from sklearn.pipeline import Pipeline
from sklearn.impute import SimpleImputer
from sklearn.preprocessing import *
from sklearn.decomposition import PCA
#Metrics
from sklearn import metrics
from sklearn.metrics import *
from sklearn.model_selection import GridSearchCV
import random
from sklearn.inspection import plot_partial_dependence
import os
import base64
from io import BytesIO
def convert_str(a):
a = str(a)
return a
def scaler(scaling_scheme='standard_scaler'):
if scaling_scheme == 'max_abs_scaler':
scal = MaxAbsScaler()
elif scaling_scheme == 'min_max_scaler':
scal = MinMaxScaler()
elif scaling_scheme == 'normalizer':
scal = Normalizer()
elif scaling_scheme == 'quantile_transformer':
scal = QuantileTransformer()
elif scaling_scheme == 'robust_scaler':
scal = RobustScaler()
elif scaling_scheme == 'power_transformer':
scal = PowerTransformer()
elif scaling_scheme == 'standard_scaler':
scal = StandardScaler()
return scal
def comb(X, pairwise_linear=False, pairwise_product=False):
from itertools import combinations
X_copy = X.copy()
columns = [str(i) for i in X.columns]
X.columns = columns
comb = combinations(columns, 2)
# Print the obtained combinations
if pairwise_linear:
for i in list(comb):
a = i[0]
b = i[1]
col_name_add = a+'+'+b
X_copy[col_name_add] = X[a]+X[b]
col_name_sub = a+'-'+b
X_copy[col_name_sub] = X[a]-X[b]
if pairwise_product:
comb = combinations(columns, 2)
# Print the obtained combinations
for i in list(comb):
a = i[0]
b = i[1]
col_name = a+'*'+b
X_copy[col_name] = X[a]*X[b]
return X_copy
def rf_colselector(X_train, y_train, no_of_cols, n_estimators=100):
rf = RandomForestClassifier(n_estimators=n_estimators)
rf.fit(X_train, y_train)
importance = rf.feature_importances_
df_importance = pd.DataFrame(importance, index = X_train.columns, columns = ['importance'])
importance_sorted = df_importance.sort_values(by=['importance'], ascending=False)
selected_columns = importance_sorted[:no_of_cols].index
return selected_columns
def corr_colselector(X_train, y_train, threshold):
d = pd.concat([X_train, y_train.reset_index(drop=True)], axis=1)
columns = d.corr().iloc[:, -1][np.logical_or((d.corr().iloc[:, -1] > threshold), (d.corr().iloc[:, -1] < -threshold))].index
return columns[:-1], d.corr()
class ColProcessor():
def __init__(self, cardinality, rf_col=False, corr_col=False, label_enc=False, interaction_only=False, poly_feat=False):
self.rf_col = rf_col
self.corr_col = corr_col
self.label_enc = label_enc
self.interaction_only = interaction_only
self.poly_feat = poly_feat
self.cardinality = cardinality
def fit(self, X, y=None):
categorical_cols = [cname for cname in X.columns if X[cname].nunique() < self.cardinality and
X[cname].dtype == "object"]
numerical_cols = [cname for cname in X.columns if X[cname].dtype in ['int64', 'float64']]
my_cols = categorical_cols + numerical_cols
self.categorical_cols = categorical_cols
self.numerical_cols = numerical_cols
self.my_cols = my_cols
X = X[my_cols].copy()
imputer_num = SimpleImputer(strategy='constant')
X_dum = imputer_num.fit_transform(X[self.numerical_cols])
self.imputer_num = imputer_num
if self.categorical_cols:
imputer_cat = SimpleImputer(strategy='most_frequent')
X_cat = imputer_cat.fit_transform(X[self.categorical_cols])
self.imputer_cat = imputer_cat
if not self.label_enc:
Ohe = OneHotEncoder(handle_unknown='ignore')
Ohe.fit(X_cat)
self.Ohe = Ohe
else:
OrdEnc = OrdinalEncoder(handle_unknown='ignore')
X_cat = OrdEnc.fit(X_cat)
self.OrdEnc = OrdEnc
return self
def transform(self, X, y=None):
X_num = pd.DataFrame(data=self.imputer_num.transform(X[self.numerical_cols]), columns=self.numerical_cols)
if self.categorical_cols:
if not self.label_enc:
X_cat = pd.DataFrame(data=self.Ohe.transform(self.imputer_cat.transform(X[self.categorical_cols])).toarray(),
columns=self.Ohe.get_feature_names(input_features=self.categorical_cols))
data = pd.concat([X_cat, X_num], axis = 1)
else:
X_cat = pd.DataFrame(self.OrdEnc.transform(self.imputer_cat.transform(X[self.categorical_cols])), columns=self.categorical_cols)
data = pd.concat([X_cat.reset_index(drop=True), X_num], axis = 1)
else:
data = X_num
return data, X_num
def interaction_feats(X):
interaction = PolynomialFeatures(2, interaction_only=True)
interaction.fit(X)
X_interaction = pd.DataFrame(data=interaction.transform(X), columns=interaction.get_feature_names(X.columns))
return X_interaction
def poly_feats(X):
poly = PolynomialFeatures(2)
poly.fit(X)
X_poly = pd.DataFrame(data=poly.transform(X), columns=poly.get_feature_names(X.columns))
return X_poly
def pca_feats(X, n_comp):
pca = PCA(n_components=n_comp)
pca.fit(X)
X_pca = pd.DataFrame(data=pca.transform(X))
return X_pca
def clubbed_feats(X, polynomial_features, interaction_only, pca_on):
if polynomial_features:
X = poly_feats(X)
elif interaction_only:
X = interaction_feats(X)
if pca_on:
X = pca_feats(X, 100)
return X
def preprocess(X_train,
y_train,
X_valid,
X_test=None,
rf_col_selection=False,
rf_no_of_cols=20,
rf_n_estimators=100,
corr_col_selection=False,
corr_threshold=0.01,
pairwise_linear=False,
pairwise_product=False):
X_train = comb(X=X_train, pairwise_linear=pairwise_linear, pairwise_product=pairwise_product)
X_valid = comb(X=X_valid, pairwise_linear=pairwise_linear, pairwise_product=pairwise_product)
if type(X_test)!=type(None):
X_test = comb(X=X_test, pairwise_linear=pairwise_linear, pairwise_product=pairwise_product)
return X_train, X_valid, X_test
def final_preprocessor(X_train,
y_train,
X_valid,
X_test=None,
rf_col_selection=False,
rf_no_of_cols=20,
rf_n_estimators=100,
corr_col_selection=False,
corr_threshold=0.01,
pairwise_linear=False,
pairwise_product=False,
cardinality=100,
polynomial_features=False,
interaction_only=False,
pca_on=False,
label_enc=False
):
col = ColProcessor(cardinality=100, label_enc=label_enc)
col.fit(X_train)
data_train, X_train_num = col.transform(X_train)
data_valid, X_valid_num = col.transform(X_valid)
if type(X_test)!=type(None):
data_test, X_test_num = col.transform(X_test)
else:
X_test_num = None
X_train_num = clubbed_feats(X_train_num,
polynomial_features=polynomial_features,
interaction_only=interaction_only,
pca_on=pca_on)
X_valid_num = clubbed_feats(X_valid_num,
polynomial_features=polynomial_features,
interaction_only=interaction_only,
pca_on=pca_on)
if type(X_test)!=type(None):
X_test_num = clubbed_feats(X_test_num,
polynomial_features=polynomial_features,
interaction_only=interaction_only,
pca_on=pca_on)
train, valid, test = preprocess(X_train_num,
y_train,
X_valid_num,
X_test_num,
rf_col_selection=rf_col_selection,
rf_no_of_cols=rf_no_of_cols,
rf_n_estimators=rf_n_estimators,
corr_col_selection=corr_col_selection,
corr_threshold=corr_threshold,
pairwise_linear=pairwise_linear,
pairwise_product=pairwise_product
)
if col.categorical_cols:
if not label_enc:
Ohe_cat_cols = col.Ohe.get_feature_names(col.categorical_cols)
train = pd.concat([train, data_train[Ohe_cat_cols]], axis=1)
valid = pd.concat([valid, data_valid[Ohe_cat_cols]], axis=1)
if type(X_test)!=type(None):
test = pd.concat([test, data_test[Ohe_cat_cols]], axis=1)
else:
train = data_train
valid = data_valid
if type(X_test)!=type(None):
test = data_test
if rf_col_selection:
columns_selected = rf_colselector(train,
y_train,
no_of_cols=rf_no_of_cols,
n_estimators=rf_n_estimators)
train = train[columns_selected]
valid = valid[columns_selected]
if type(X_test)!=type(None):
test = test[columns_selected]
if corr_col_selection:
corr_cols, df = corr_colselector(train, y_train, threshold=corr_threshold)
train = train[corr_cols]
valid = valid[corr_cols]
if type(X_test)!=type(None):
test = test[corr_cols]
return train, valid, test, col
def combined_metrics(X_test, y_test, clf):
#to be used in combined metrics function.
# enc = LabelEncoder()
# trans_y_train = enc.fit_transform(y_train)
ohe = OneHotEncoder(handle_unknown='ignore')
ohe.fit(y_train.values.reshape(-1, 1))
y_test_ohe = ohe.transform(y_test.values.reshape(-1, 1)).toarray()
metrics_list = [[accuracy_score(y_test,
clf.predict(X_test))],
[precision_score(y_test, clf.predict(X_test), average = 'micro')],
[recall_score(y_test, clf.predict(X_test), average = 'micro')],
[f1_score(y_test, clf.predict(X_test), average = 'micro')],
[roc_auc_score(y_test_ohe, ohe.transform(clf.predict(X_test).reshape(-1, 1)).toarray(), multi_class='ovr')],
[hamming_loss(y_test, clf.predict(X_test))],
[log_loss(y_test_ohe, ohe.transform(clf.predict(X_test).reshape(-1, 1)).toarray())]
]
index = ['Accuracy', 'Precision', 'Recall', 'F1 Score', 'ROC AUC', 'Hamming Loss', 'Log Loss']
# index = ['Accuracy', 'Precision', 'Recall', 'F1 Score', 'Hamming Loss', 'Log Loss']
# index = ['Accuracy', 'Precision', 'Recall', 'F1 Score', 'ROC AUC', 'Hamming Loss']
df_metric = pd.DataFrame(metrics_list, index = index, columns = ['Value'])
return df_metric
def confusion_matrix_plot(cm, class_names, title = 'Confusion Matrix Plot'):
plt.clf()
plt.imshow(cm, interpolation='nearest', cmap = 'Blues_r')
plt.title(title)
plt.ylabel('True')
plt.xlabel('Predicted')
tick_marks = np.arange(len(class_names))
plt.xticks(tick_marks, class_names, rotation=45)
plt.yticks(tick_marks, class_names)
s = [['TN','FP'], ['FN', 'TP']]
for i in range(len(class_names)):
for j in range(len(class_names)):
plt.text(j,i, str(cm[i][j]))
plt.show()
def to_excel(df):
output = BytesIO()
writer = pd.ExcelWriter(output, engine='xlsxwriter')
df.to_excel(writer, index = False, sheet_name='Sheet1')
workbook = writer.book
worksheet = writer.sheets['Sheet1']
format1 = workbook.add_format({'num_format': '0.00'}) # Tried with '0%' and '#,##0.00' also.
worksheet.set_column('A:A', None, format1) # Say Data are in column A
writer.save()
processed_data = output.getvalue()
return processed_data
def get_table_download_link(df):
"""Generates a link allowing the data in a given panda dataframe to be downloaded
in: dataframe
out: href string
"""
val = to_excel(df)
b64 = base64.b64encode(val) # val looks like b'...'
return f'<a href="data:application/octet-stream;base64,{b64.decode()}" download="Your_File.xlsx">Download output file</a>' # decode b'abc' => abc
def GNB():
gnb_params = {'clf__estimator':[GaussianNB()]
}
return gnb_params
def LogisticReg():
lr_params = {'clf__estimator': [LogisticRegression()]
}
st.subheader('Logistic Regression')
penalty = st.multiselect('Penalty', ['l1', 'l2'], ['l2'])
reg = st.multiselect('C', [0.1, 1.0, 2.0], [1.0])
solver = st.multiselect('Solver', ['liblinear', 'newton-cg', 'lbfgs', 'sag', 'saga'], ['liblinear'])
lr_params['clf__estimator__penalty'] = penalty
lr_params['clf__estimator__C'] = reg
lr_params['clf__estimator__solver'] = solver
return lr_params
def KNN():
knn_params = {'clf__estimator': [KNeighborsClassifier()]
}
st.subheader('KNN')
n_neighbors = st.multiselect('Neighbors', list(range(1,30)), [5])
leaf_size = st.multiselect('Leaf Size', list(range(1,50)), [30])
p_distance = st.multiselect('Distance Metric', [1,2], [2])
knn_params['clf__estimator__n_neighbors'] = n_neighbors
knn_params['clf__estimator__leaf_size'] = leaf_size
knn_params['clf__estimator__p'] = p_distance
return knn_params
def SVM():
svm_params = {'clf__estimator': [SVC(probability=True)]
}
st.subheader('Support Vector Machines')
c = st.multiselect('C', [0.1, 1, 10, 100, 1000], [1])
gamma = st.multiselect('Gamma', ['scale', 'auto'], ['scale'])
kernel = st.multiselect('Kernel', ['linear', 'rbf', 'poly', 'sigmoid'], ['rbf'])
svm_params['clf__estimator__C'] = c
svm_params['clf__estimator__gamma'] = gamma
svm_params['clf__estimator__kernel'] = kernel
return svm_params
def DT():
dt_params = {'clf__estimator': [DecisionTreeClassifier()]}
st.subheader('Decision Tree')
criterion = st.multiselect('Criterion', ["gini", "entropy"], ['gini'])
min_samp_split = st.multiselect('Min Samples Split', [2, 10], [2])
max_depth = st.multiselect('Max Depth', [2, 5, 10], [10])
dt_params['clf__estimator__criterion'] = criterion
dt_params['clf__estimator__min_samples_leaf'] = min_samp_split
dt_params['clf__estimator__max_depth'] = max_depth
return dt_params
def RF():
rf_params = {'clf__estimator': [RandomForestClassifier()]
}
st.subheader('Random Forest')
n_estimators = st.multiselect('Number of Trees', [100, 200, 500], [100])
max_features = st.multiselect('Max Features', [2, 10, 'auto', 'sqrt', 'log2'], ['auto'])
max_depth = st.multiselect('Max Depth', [4,5,6,7,8, None], [None])
criterion = st.multiselect('Criteria', ['gini', 'entropy'], ['gini'])
rf_params['clf__estimator__n_estimators'] = n_estimators
rf_params['clf__estimator__max_features'] = max_features
rf_params['clf__estimator__max_depth'] = max_depth
rf_params['clf__estimator__criterion'] = criterion
return rf_params
def GB():
gb_params = {'clf__estimator': [GradientBoostingClassifier()]
}
st.subheader('Gradient Booster')
loss = st.multiselect('Loss Function', ['deviance', 'exponential'], ['deviance'])
learning_rate = st.multiselect('Learning Rate', [0.001, 0.01, 0.1], [0.1])
min_samples_split = st.multiselect('Min Samples Split', list(range(1, 10)), [2])
min_samples_leaf = st.multiselect('Min Samples Leaf', list(range(1, 10)), [1])
max_depth = st.multiselect('Max Depth', [1, 2, 3, 4, 5, 6], [3])
max_features = st.multiselect('Max Features', ['auto', 'log2', 'sqrt', None], [None])
criterion = st.multiselect('Criterion', ['friedman_mse', 'mse', 'mae'], ['friedman_mse'])
subsample = st.multiselect('Subsample', [0.5, 0.618, 0.8, 0.85, 0.9, 0.95, 1.0], [1.0])
n_estimators = st.multiselect('Number of Trees', [50, 100, 150, 200, 250], [100])
gb_params['clf__estimator__loss'] = loss
gb_params['clf__estimator__learning_rate'] = learning_rate
gb_params['clf__estimator__min_samples_split'] = min_samples_split
gb_params['clf__estimator__min_samples_leaf'] = min_samples_leaf
gb_params['clf__estimator__max_depth'] = max_depth
gb_params['clf__estimator__max_features'] = max_features
gb_params['clf__estimator__criterion'] = criterion
gb_params['clf__estimator__subsample'] = subsample
gb_params['clf__estimator__n_estimators'] = n_estimators
return gb_params
def ERT():
ert_params = {'clf__estimator': [ExtraTreesClassifier()]
}
st.subheader('Extra Random Trees')
n_estimators = st.multiselect('Number of Trees', [100, 200, 500, 1000], [100]) #fix
max_depth = st.multiselect('Max Depth', [None, 4, 5, 6, 7, 8, 9], [None]) #fix
min_samples_leaf = st.multiselect('Min Sample per Leaf', [1, 2, 3, 4, 5], [1])
n_jobs = st.selectbox('Parallelism', [1, 2, 3, 4, -1], 4)
ert_params['clf__estimator__n_estimators'] = n_estimators
ert_params['clf__estimator__max_depth'] = max_depth
ert_params['clf__estimator__min_samples_leaf'] = min_samples_leaf
ert_params['clf__estimator__n_jobs'] = [n_jobs]
return ert_params
def XGB():
xgb_params ={'clf__estimator':[XGBClassifier()]
}
st.subheader('XGBoost')
n_estimators = st.multiselect('Number of Trees', list(range(50, 1000, 50)), [50]) #fix
max_depth = st.multiselect('Max Depth', list(range(1, 20)), [6]) #fix
min_child_weight = st.multiselect('Min Child Weight', list(range(1, 10, 1)), [1])
gamma = st.multiselect('Gamma', list(range(0, 10)), [1])
learning_rate = st.multiselect('Learning Rate', [0.01, 0.05, 0.1, 0.2, 0.3], [0.3])
subsample = st.multiselect('Subsample', list(np.divide(range(5, 11), 10)), [1.0])
booster = st.multiselect('Booster', ['gbtree', 'gblinear'], ['gbtree'])
xgb_params['clf__estimator__n_estimators'] = n_estimators
xgb_params['clf__estimator__max_depth'] = max_depth
xgb_params['clf__estimator__min_child_weight'] = min_child_weight
xgb_params['clf__estimator__gamma'] = gamma
xgb_params['clf__estimator__learning_rate'] = learning_rate
xgb_params['clf__estimator__subsample'] = subsample
xgb_params['clf__estimator__booster'] = booster
return xgb_params
def SGD():
sgd_params = {'clf__estimator': [SGDClassifier()]
}
st.subheader('SGD')
loss = st.multiselect('Loss Function', ['hinge', 'log', 'modified_huber', 'squared_hinge', 'perceptron'], ['hinge']) #fix
max_iter = st.multiselect('Max Iterations', list(np.multiply(range(5, 16), 100)), [1000]) #fix
tol = st.multiselect('Tolerance', [0.0001, 0.001, 0.05, 0.1], [0.0001])
penalty = st.multiselect('Penalty', ['l2', 'l1', 'elasticnet'], ['l2'])
alpha = st.multiselect('Alpha', [0.0001, 0.001, 0.05, 0.1, 0.2, 0.3], [0.0001])
n_jobs = st.selectbox('Parallelization', [1, 2, 3, 4, -1], 4)
sgd_params['clf__estimator__loss'] = loss
sgd_params['clf__estimator__max_iter'] = max_iter
sgd_params['clf__estimator__tol'] = tol
sgd_params['clf__estimator__penalty'] = penalty
sgd_params['clf__estimator__alpha'] = alpha
sgd_params['clf__estimator__n_jobs'] = [n_jobs]
return sgd_params
def NN():
nn_params = {'clf__estimator': [MLPClassifier()]
}
st.subheader('Neural Network')
solver = st.multiselect('Solver', ['lbfgs', 'sgd', 'adam'], ['adam'])
max_iter = st.multiselect('Max Iterations', [1000,1100,1200,1300,1400], [1000])
alpha = st.multiselect('Alpha', list(10.0 ** -np.arange(1, 10)), [0.0001])
hidden_layer_sizes = st.multiselect('Hidden Layer Sizes', list(range(50, 500, 50)), [100])
# hidden_layer_sizes = st.multiselect('Hidden Layer Sizes', [50, 100, 150, 200, 250, 300, 350, 400, 450, 500] , [100])
nn_params['clf__estimator__solver'] = solver
nn_params['clf__estimator__max_iter'] = max_iter
nn_params['clf__estimator__alpha'] = alpha
nn_params['clf__estimator__hidden_layer_sizes'] = hidden_layer_sizes
return nn_params
data = st.file_uploader('Upload a csv')
test_data = st.file_uploader('Upload a csv for prediction:')
if (data != None) & (test_data != None):
df = pd.read_csv(data)
df_test = pd.read_csv(test_data)
# df = random.shuffle(data)
target_col =st.selectbox('Choose target variable', df.columns)
X = df.drop(target_col, axis = 1)
y = df[target_col]
test_ratio = st.number_input('Enter test split ratio, 0 < ratio < 1', min_value = 0.0,
max_value = 1.0, value = 0.2)
if test_ratio:
X_train_full, X_valid_full, y_train, y_valid = train_test_split(X, y,
test_size=test_ratio,
random_state = 0)
selected_models = st.sidebar.multiselect(
'Choose Algorithms:',(
'Gaussian NB',
'Logistic Regression',
'KNN',
'Support Vector Machines',
'Decision Tree',
'Random Forest',
'Gradient Boosting',
'Extra Random Trees',
'XGBoost',
'Stochastic Gradient Descent',
'Neural Network'), ['KNN', 'Support Vector Machines', 'Decision Tree'])
if selected_models:
func_dict = {'Gaussian NB': GNB(),
'Logistic Regression':LogisticReg(),
'KNN': KNN(),
'Support Vector Machines': SVM(),
'Decision Tree': DT(),
'Random Forest': RF(),
'Gradient Boosting': GB(),
'Extra Random Trees': ERT(),
'XGBoost': XGB(),
'Stochastic Gradient Descent': SGD(),
'Neural Network': NN()
}
param_dict = {}
for i in selected_models:
param_dict[i] = func_dict[i]
from sklearn.base import BaseEstimator, ClassifierMixin
class MyClassifier(BaseEstimator, ClassifierMixin):
def __init__(
self,
estimator = XGBClassifier(),
):
"""
A Custom BaseEstimator that can switch between classifiers.
:param estimator: sklearn object - The classifier
"""
self.estimator = estimator
def fit(self, X, y=None, **kwargs):
self.estimator.fit(X, y)
return self
def predict(self, X, y=None):
return self.estimator.predict(X)
def predict_proba(self, X):
return self.estimator.predict_proba(X)
def score(self, X, y):
return self.estimator.score(X, y)
@property
def classes_(self):
return self.estimator.classes_
X_train, X_valid, df_test, col = final_preprocessor(X_train_full,
y_train,
X_valid_full,
df_test,
rf_col_selection=True,
rf_no_of_cols=20,
rf_n_estimators=100,
corr_col_selection=True,
corr_threshold=0.2,
pairwise_linear=False,
pairwise_product=False,
cardinality=100,
polynomial_features=False,
interaction_only=False,
pca_on=False
)
data_valid = pd.concat([X_valid, y_valid.reset_index(drop=True)], axis = 1)
my_pipeline = Pipeline([('scaler', scaler(scaling_scheme='power_transformer')),
('clf', MyClassifier())
])
parameters = []
for i in selected_models:
parameters.append(param_dict[i])
st.write(parameters)
train = st.button('Train Model')
if train:
with st.spinner('Training Model...'):
from sklearn.model_selection import GridSearchCV
gscv = GridSearchCV(my_pipeline, parameters, cv=3, n_jobs=-1, return_train_score=False, verbose=3)
gscv.fit(X_train, y_train)
st.text('Best Parameters')
st.write(gscv.best_params_)
st.text('Best Score')
st.write(gscv.best_score_)
st.text('Fit vs Time vs HyperParameters')
data = gscv.cv_results_.values()
columns = gscv.cv_results_.keys()
df_fit = pd.DataFrame(data, columns).T
df_fit['param_clf__estimator'] = df_fit['param_clf__estimator'].apply(convert_str)
st.write(df_fit)
st.text('Prediction on Validation Data')
data_valid['Predicted'] = gscv.predict(X_valid)
st.write(data_valid)
st.text('Confusion Matrix')
cm = confusion_matrix(y_valid, gscv.predict(X_valid))
fig1, ax1 = plt.subplots()
class_names = y_valid.unique()
confusion_matrix_plot(cm, class_names)
st.pyplot(fig1)
st.text('Performance Metrics')
st.write(combined_metrics(X_valid, y_valid, gscv))
st.text('Partial Dependence Plot')
features = [0, 1, (0, 1)]
fig, ax = plt.subplots(1,3, figsize = (15,9))
plot_partial_dependence(gscv, X_valid, features=features, target=0, ax=ax)
plt.tight_layout()
st.pyplot(fig)
st.text('ICE Plot')
features = [0, 1]
fig, ax = plt.subplots(figsize=(7, 6))
plot_partial_dependence(gscv, X_valid, features, kind='both', target=0, ax=ax)
plt.tight_layout()
st.pyplot(fig)
st.text('Prediction on Test file')
df_test['Predicted'] = gscv.predict(df_test)
st.write(df_test)
st.text('Shapley Explainer')
# X_test = df_test.drop('Predicted', axis = 1)
explainer = shap.KernelExplainer(gscv.predict_proba, X_valid)
shap_values = explainer.shap_values(X_valid.iloc[2,:])
st.pyplot(shap.force_plot(explainer.expected_value[0], shap_values[0], X_valid.iloc[2,:], matplotlib=True, text_rotation=8))
st.text('Shapley Explainer WaterFall Plot')
f = lambda x: gscv.predict_proba(x)[:,1]
med = X_train.median().values.reshape((1,X_train.shape[1]))
explainer = shap.Explainer(f, med)
shap_values = explainer(X_train.iloc[0:100,:])
st.pyplot(shap.plots.waterfall(shap_values[2], max_display=7))
st.text('Partial Dependence Plot from pdp_box')
pdp_ = pdp.pdp_isolate(model=gscv, dataset=X_valid,
model_features=X_valid.columns,
feature=X_train.columns[0])
fig, axes = pdp.pdp_plot(pdp_isolate_out=pdp_, feature_name=X_valid.columns[0], center = True, ncols=1, figsize = (15, 10))
st.pyplot(fig)
# -
from nbdev.export import notebook2script; notebook2script()
| 00_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
import altair as alt
from collections import namedtuple
import pandas as pd
alt.data_transformers.disable_max_rows()
# JS snippet to extract table data
#
# ```javascript
# copy(
# Array.from($0.querySelectorAll('tr'))
# .map(arr => Array.from(arr.querySelectorAll('td')))
# .map(arr => arr.map(el => el.innerText))
# )
# ```
# +
def load_data():
df = pd.read_json("/lab/data/sensex.json")
df.columns = ['date', 'open', 'high', 'low', 'close', 'adj_close', 'volume']
df = df[['date', 'close']]
df = df[~df['close'].isin(['-'])]
df.date = pd.to_datetime(df.date, format='%b %d, %Y')
df.close = df.close.str.replace(',', '').astype(float)
df = df.sort_values(by='date')
df = df.reset_index(drop=True)
# Resample to weeks
# df['weeknum'] = df.date.dt.isocalendar().year * 100 + df.date.dt.isocalendar().week
# df = df.groupby('weeknum')[['date', 'close']].nth(-1).reset_index(drop=True)
return df
df = load_data()
# -
df
# +
def equity_curve(df, period):
starting_capital = 1_00_000
equity_curve = []
EquityCurve = namedtuple("EquityCurve", ["date", "cash", "units", "price", "ma", "pf_value"])
df['ma'] = df.close.rolling(window=period).mean().round(2)
df = df.dropna()
init = df.iloc[0]
tail = df[1:].reset_index(drop=True)
# calc first entry
if init.close > init.ma:
entry = EquityCurve(
date=init.date,
cash=0,
units=round(starting_capital/init.close, 4),
price=init.close,
ma=init.ma,
pf_value=starting_capital
)
else:
entry = EquityCurve(
date=init.date,
cash=starting_capital,
units=0,
price=init.close,
ma=init.ma,
pf_value=starting_capital
)
equity_curve.append(entry)
# LOOP THROUGH EVERY DATE
for index, row in tail.iterrows():
prev = equity_curve[index]
is_deployed = prev.units != 0
# update holdings value
if is_deployed and row.close > row.ma:
entry = EquityCurve(
date=row.date,
cash=0,
units=prev.units,
price=row.close,
ma=row.ma,
pf_value=round(prev.units * row.close, 2)
)
# move to cash
if is_deployed and row.close <= row.ma:
entry = EquityCurve(
date=row.date,
cash=round(prev.units * row.close, 2),
units=0,
price=row.close,
ma=row.ma,
pf_value=round(prev.units * row.close, 2)
)
# deploy cash
if not is_deployed and row.close > row.ma:
entry = EquityCurve(
date=row.date,
cash=0,
units=round(prev.pf_value/row.close, 4),
price=row.close,
ma=row.ma,
pf_value=prev.pf_value
)
# continue with previous day's value
if not is_deployed and row.close <= row.ma:
entry = EquityCurve(
date=row.date,
cash=prev.cash,
units=0,
price=row.close,
ma=row.ma,
pf_value=prev.pf_value
)
equity_curve.append(entry)
return equity_curve
eq_curve = equity_curve(df, 100)
pf = pd.DataFrame(eq_curve)
# -
pf
# +
pf.price = (pf.price.pct_change() + 1).fillna(100000).cumprod().round(2)
pf = pd.melt(pf, id_vars=['date'], value_vars=['price', 'pf_value'])
pf
# -
alt.Chart(pf).mark_line().encode(
x='date',
y='value',
color='variable',
tooltip=['date', 'value', 'variable']
).properties(
title="Sensex 100DMA Strategy",
width=800,
height=600
)
| src/notebooks/sensex_100dma.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Dependencies matplotlib's pyplot, pandas, scripy's stats with sem
# +
# Load data as diaper_data
# Display the first 10 rows of diaper_data
# +
# Create separate dataframes of each diaper type.
# Name the dataframes diaper_type_a, diaper_type_b, diaper_type_c
# Display the first 5 rows of diaper_type_a
# +
# Set samples as a list containing the list of happiness from diaper_type_a, diaper_type_b, and diaper_type_c
# Display samples
# +
# Create a variable means and set it to a list of calculated means
# Display the means
# +
# Create a variable sem and set it to a list of calculated standard error on means
# Display the standard error on means (sem)
# +
# Plot sample means with error bars
# Create the fig as ax subplot
# Add errorbars to ax with the following:
# Label the x-axis as ["A", "B", "C"],
# Add the means
# y error as sem
# Add a label to the data points
# Organize the x axis
# Update the x and y axis labels
# Show the legend.
# Show plot.
# -
| Matplotlib_Code-Drills/day-03/01/day-03_01.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## 2.4.1 넘파이
# ### 넘파이 배열
import numpy as np
# +
a = np.array([[1,2,3], [1,5,9], [3,5,7]])
print(a.ndim)
# 2
print(a.shape)
# (3,3)
print(a.size)
# 9
print(a.dtype)
# dtype('int32')
# +
a = np.zeros((2,3)) #원소가 모두 0인 배열 생성
print(a)
# array([[0., 0., 0.],
# [0., 0., 0.]])
b = np.ones((2,1)) #원소가 모두 1인 배열 생성
print(b)
# array([[1.],
# [1.]])
c = np.empty((2,2)) #원소값을 초기화하지 않은 배열 생성
print(c)
# array([[1.96088859e+243, 5.22864540e-067],
# [1.47173270e-052, 1.47214053e-052]])
d = np.arange(10, 30, 5) # 10부터 30전 까지 5단위로 배열 생성
print(d)
# array([10, 15, 20, 25])
e = np.full((2, 2), 4) #원소가 모두 4인 배열 생성
print(e)
# array([[4, 4],
# [4, 4]])
f = np.eye(3) # 3x3 크기의 단위행렬 생성
print(f)
# array([[1., 0., 0.],
# [0., 1., 0.]
# [0., 0., 1.]))
g = np.random.random((2, 2)) # 임의값을 가지는 배열 생성
print(g)
# array([[0.94039485, 0.18284953],
# [0.59283818, 0.48592838]])
# -
# ### 넘파이 기본연산
# +
a = np.array([1, 2, 3])
b = np.array([10, 20, 30])
print(a+b)
# array([11, 22, 33])
print(np.add(a,b)) # 위의 연산과 같다.
# array([11, 22, 33])
print(b-a)
# array([9, 18, 27])
print(np.subtract(b,a)) # 위의 연산과 같다.
# array([9, 18, 27])
print(a**2)
# array([1, 4, 9])
print(b<15)
# array([True, False, False])
C = np.array([[1, 2],
[3, 4]])
D = np.array([[10, 20],
[30, 10]])
print(C*D) # 원소별 곱셈
# array([[10, 40],
# [90, 40]])
print(np.dot(C,D)) # 내적(dot product) 계산
# array([[ 70, 40],
# [150, 100]])
print(C.dot(D)) #내적의 또 다른 사용법
# array([[ 70, 40],
# [150, 100]])
# +
a = np.array([[ 1, 2, 3, 4],
[ 5, 6, 7, 8],
[ 1, 3, 5, 7]])
print(a.sum(axis=0)) # 0축, 즉 열을 기준으로한 덧셈
# array([7, 11, 15, 19])
print(a.sum(axis=1)) # 1축, 즉 행을 기준으로한 덧셈
# array([10, 26, 16])
print(a.max(axis=1)) # 각 행에서의 최대값
# array([4, 8, 7])
# -
# ### 넘파이 배열 인덱싱, 슬라이싱
# +
a = np.array([1, 2, 3, 4, 5, 6, 7])
print(a[3])
# 4
print(a[-1]) # 마지막 원소
# 7
print(a[2: 5]) # 인덱스값 5를 가지는 원소 전까지
# array([3, 4, 5])
print(a[2: ])
# array([3, 4, 5, 6, 7])
print(a[ :4])
# array([1, 2, 3, 4])
# +
a = np.array([[ 1, 2, 3],
[ 4, 5, 6],
[ 7, 8, 9]])
print(a[1, 2])
# 6
print(a[ : , 1]) # 1열의 모든 원소
# array([2, 5, 8])
print(a[-1]) # 마지막 행
# array([7, 8, 9])
# -
# ### 넘파이 배열 형태 변환
# +
a = np.array([[1, 2, 3, 4],
[5, 6, 7, 8],
[9,10,11,12]])
print(a.ravel()) # 1차원 배열(벡터)로 만들어준다.
# array([1, 2, 3, 4, 5, 6, 7, 8, 9, 10, 11, 12])
print(a.reshape(2, 6))
# array([[ 1, 2, 3, 4, 5, 6],
# [ 7, 8, 9, 10, 11, 12]])
print(a.T)
# array([[ 1, 5, 9],
# [ 2, 6, 10],
# [ 3, 7, 11],
# [ 4, 8, 12]])
# -
print(a.reshape(3, -1)) # 3행만 지정해주고 열에 -1을 넣으면 자동으로 배열을 reshape한다.
# array([[ 1, 2, 3, 4],
# [ 5, 6, 7, 8],
# [ 9, 10, 11, 12]])
# ### 넘파이 브로드캐스팅
# +
a = np.array([[1,2,3], [4,5,6], [7,8,9]])
b = np.array([1,0,1])
y = np.empty_like(a) # 배열 a와 크기가 같은 원소가 비어있는 배열 생성
# 배열 b를 a의 각 행에 더해주기 위해 반복문을 사용한다.
for i in range(3):
y[i, : ] = a[i, : ] + b
print(y)
# array([[ 2, 2, 4],
# [ 5, 5, 7],
# [ 8, 8, 10]])
# +
a = np.array([[1,2,3], [4,5,6], [7,8,9]])
b = np.array([1,0,1])
c = a + b # c를 따로 선언 할 필요 없이 만들 수 있다.
print(c)
# array([[ 2, 2, 4],
# [ 5, 5, 7],
# [ 8, 8, 10]])
| 2.NLP_PREP/2.4.1.numpy.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
# +
import os
import glob
import random
import matplotlib.pyplot as plt
import numpy as np
# -
import mr
from mrcnn import model as modellib
from mrcnn import visualize
def get_ax(rows=1, cols=1, size=8):
_, ax = plt.subplots(rows, cols, figsize=(size*cols, size*rows))
return ax
config = mr.TrainConfig()
config.display()
# +
work_path = work_path = os.path.join("E:", os.sep, f"RCNN{mr.CLASS_NAME}{mr.IMAGE_DIM}Train")
os.chdir(work_path)
tif_glob = glob.glob(os.path.join("*", "images", "*.tif"))
n = int(len(tif_glob) * .8)
ds_train = mr.MRDataset()
ds_train.load_glob(tif_glob[:n])
ds_train.prepare()
ds_valid = mr.MRDataset()
ds_valid.load_glob(tif_glob[n:])
ds_valid.prepare()
# -
len(tif_glob)
image_ids = np.random.choice(ds_train.image_ids,4)
for image_id in image_ids:
image = ds_train.load_image(image_id)
mask, class_ids = ds_train.load_mask(image_id)
visualize.display_top_masks(image,mask,class_ids,ds_train.class_names)
model = modellib.MaskRCNN(mode="training",
config=config,
model_dir="logs")
# +
# Which weights to start with?
init_with = "imagenet" # imagenet, coco, or last
if init_with == "imagenet":
model.load_weights(model.get_imagenet_weights(), by_name=True)
elif init_with == "coco":
# Load weights trained on MS COCO, but skip layers that
# are different due to the different number of classes
# See README for instructions to download the COCO weights
model.load_weights(mr.COCO_MODEL_PATH, by_name=True,
exclude=["mrcnn_class_logits",
"mrcnn_bbox_fc",
"mrcnn_bbox",
"mrcnn_mask"])
elif init_with == "last":
# Load the last model you trained and continue training
model_path = model.find_last()
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
# -
model.train(ds_train, ds_valid,
learning_rate=config.LEARNING_RATE,
epochs=50,
layers='heads')
model.train(ds_train, ds_valid,
learning_rate=config.LEARNING_RATE / 10.0,
epochs=100,
layers="4+")
model.train(ds_train, ds_valid,
learning_rate=config.LEARNING_RATE / 20.0,
epochs=300,
layers="all")
# +
inference_config = mr.InferenceConfig()
# Recreate the model in inference mode
model = modellib.MaskRCNN(mode="inference",
config=inference_config,
model_dir="logs")
# Get path to saved weights
# Either set a specific path or find last trained weights
# model_path = os.path.join(ROOT_DIR, ".h5 file name here")
model_path = model.find_last()
print("Loading weights from ", model_path)
model.load_weights(model_path, by_name=True)
# +
dataset = ds_valid
image_id = random.choice(dataset.image_ids)
original_image, image_meta, gt_class_id, gt_bbox, gt_mask =\
modellib.load_image_gt(dataset, inference_config,
image_id, use_mini_mask=False)
results = model.detect([original_image], verbose=0)
r = results[0]
ax1, ax2 = get_ax(1,2)
visualize.display_instances(original_image, gt_bbox, gt_mask, gt_class_id,
dataset.class_names, ax=ax1)
visualize.display_instances(original_image, r['rois'], r['masks'], r['class_ids'],
dataset.class_names, r['scores'], ax=ax2)
| train.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: S2S Challenge
# language: python
# name: s2s
# ---
import climetlab as cml
import climetlab_s2s_ai_challenge
print(f'Climetlab version : {cml.__version__}')
print(f'Climetlab-s2s-ai-challenge plugin version : {climetlab_s2s_ai_challenge.__version__}')
from crims2s import util
cml.settings.get('cache-directory')
# + jupyter={"outputs_hidden": true} tags=[]
for f in util.ECMWF_FORECASTS:
print(f)
# -
cmlds = cml.load_dataset("s2s-ai-challenge-training-input", parameter='t2m', origin='ncep')
ncep = cmlds.to_xarray()
ncep
| notebooks/landryda/redownload.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] slideshow={"slide_type": "slide"}
# Here we will test parameter recovery and model comparison for Rescorla-Wagner (RW), Hierarchical Gaussian Filters (HGF), and Switching Gaussian Filters (SGF) models of the social influence task.
# + slideshow={"slide_type": "skip"}
import numpy as np
from scipy import io
import matplotlib.pyplot as plt
import seaborn as sns
import pandas as pd
sns.set(style = 'white', color_codes = True)
# %matplotlib inline
import sys
import os
import os
cwd = os.getcwd()
sys.path.append(cwd[:-len('befit/examples/social_influence')])
# + [markdown] slideshow={"slide_type": "subslide"}
# Lets start by generating some behavioral data from the social influence task. Here green advice/choice is encoded as 0 and the blue advice/choice is encoded as 1.
# + slideshow={"slide_type": "subslide"}
import torch
from torch import ones, zeros, tensor
torch.manual_seed(1234)
nsub = 50 #number of subjects
trials = 120 #number of samples
from befit.tasks import SocialInfluence
from befit.simulate import Simulator
from befit.inference import Horseshoe, Normal
from befit.agents import RLSocInf, HGFSocInf, SGFSocInf
# load stimuli (trial offers, advices, and reliability of advices)
reliability = torch.from_numpy(np.load('advice_reliability.npy')).float()
reliability = reliability.reshape(trials, -1, 1).repeat(1, 1, nsub).reshape(trials, -1).unsqueeze(0)
offers = torch.from_numpy(np.load('offers.npy')).reshape(trials, -1, 1).repeat(1, 1, nsub)\
.reshape(trials, -1).unsqueeze(0)
stimuli = {'offers': offers,
'reliability': reliability}
socinfl = SocialInfluence(stimuli, nsub=nsub)
# RL agent
rl_agent = RLSocInf(runs=2*nsub, trials=trials)
trans_pars1 = torch.arange(-.5,.5,1/(2*nsub)).reshape(-1, 1) + tensor([[-2., 4., 0., 0.]])
rl_agent.set_parameters(trans_pars1)
sim1 = Simulator(socinfl, rl_agent, runs=2*nsub, trials=trials)
sim1.simulate_experiment()
# HGF agent
hgf_agent = HGFSocInf(runs=2*nsub, trials=trials)
trans_pars2 = torch.arange(-.5, .5, 1/(2*nsub)).reshape(-1, 1) + tensor([[2., 0., 4., 0., 0.]])
hgf_agent.set_parameters(trans_pars2)
sim2 = Simulator(socinfl, hgf_agent, runs=2*nsub, trials=trials)
sim2.simulate_experiment()
# SGF agent
sgf_agent = SGFSocInf(runs=2*nsub, trials=trials)
trans_pars3 = torch.arange(-.5, .5, 1/(2*nsub)).reshape(-1, 1) + tensor([[-2., -1., 4., 0., 0.]])
sgf_agent.set_parameters(trans_pars3)
sim3 = Simulator(socinfl, sgf_agent, runs=2*nsub, trials=trials)
sim3.simulate_experiment();
def posterior_accuracy(labels, df, vals):
for i, lbl in enumerate(labels):
std = df.loc[df['parameter'] == lbl].groupby(by='subject').std()
mean = df.loc[df['parameter'] == lbl].groupby(by='subject').mean()
print(lbl, np.sum(((mean+2*std).values[:, 0] > vals[i])*((mean-2*std).values[:, 0] < vals[i]))/(2*nsub))
# -
# plot performance of different agents in different blocks
# +
def compute_mean_performance(outcomes, responses):
cc1 = (outcomes * responses > 0.).float() # accept reliable offer
cc2 = (outcomes * (1 - responses) < 0.).float() # reject unreliable offer
return torch.einsum('ijk->k', cc1 + cc2)/trials
perf1 = compute_mean_performance(sim1.stimulus['outcomes'][..., 0],
sim1.responses.float()).numpy().reshape(2, -1)
print('RL agent: ', np.median(perf1, axis = -1))
fig, ax = plt.subplots(1,2, sharex = True, sharey = True)
ax[0].hist(perf1[0]);
ax[1].hist(perf1[1]);
fig.suptitle('RL agent', fontsize = 20);
ax[0].set_ylim([0, 20]);
ax[0].set_xlim([.5, 1.]);
perf2 = compute_mean_performance(sim2.stimulus['outcomes'][..., 0],
sim2.responses.float()).numpy().reshape(2, -1)
print('HGF agent: ', np.median(perf2, axis = -1))
fig, ax = plt.subplots(1,2, sharex = True, sharey = True)
ax[0].hist(perf2[0]);
ax[1].hist(perf2[1]);
fig.suptitle('HGF agent', fontsize = 20);
ax[0].set_ylim([0, 20]);
ax[0].set_xlim([.5, 1.]);
perf3 = compute_mean_performance(sim3.stimulus['outcomes'][..., 0],
sim3.responses.float()).numpy().reshape(2, -1)
print('SGF agent: ', np.median(perf3, axis = -1))
fig, ax = plt.subplots(1,2, sharex = True, sharey = True)
ax[0].hist(perf3[0]);
ax[1].hist(perf3[1]);
fig.suptitle('SGF agent', fontsize = 20);
ax[0].set_ylim([0, 20]);
ax[0].set_xlim([.5, 1.]);
# + [markdown] slideshow={"slide_type": "slide"}
# Fit simulated behavior
# +
stimulus = sim1.stimulus
stimulus['mask'] = torch.ones(1, 120, 100)
rl_infer = Horseshoe(rl_agent, stimulus, sim1.responses)
rl_infer.infer_posterior(iter_steps=200)
labels = [r'$\alpha$', r'$\zeta$', r'$\beta$', r'$\theta$']
tp_df = rl_infer.sample_posterior(labels, n_samples=1000)
# -
sim1.responses.dtype
# Compute fit quality and plot posterior estimates from a hierarchical parameteric model
# +
labels = [r'$\alpha$', r'$\zeta$', r'$\beta$', r'$\theta$']
trans_pars_rl = tp_df.melt(id_vars='subject', var_name='parameter')
vals = [trans_pars1[:,0].numpy(), trans_pars1[:, 1].numpy(), trans_pars1[:, 2].numpy(), trans_pars1[:, 3].numpy()]
posterior_accuracy(labels, trans_pars_rl, vals)
# +
plt.figure()
#plot convergence of stochasitc ELBO estimates (log-model evidence)
plt.plot(rl_infer2.loss[-400:])
g = sns.FacetGrid(trans_pars_rl, col="parameter", height=3, sharey=False);
g = (g.map(sns.lineplot, 'subject', 'value', ci='sd'));
labels = [r'$\alpha$', r'$\zeta$', r'$\beta$', r'bias']
for i in range(len(labels)):
g.axes[0,i].plot(np.arange(2*nsub), trans_pars1[:,i].numpy(),'ro', zorder = 0);
# -
# fit HGF agent to simulated data
# +
stimulus = sim2.stimulus
stimulus['mask'] = torch.ones(1, 120, 100)
hgf_infer = Horseshoe(hgf_agent, stimulus, sim2.responses)
hgf_infer.infer_posterior(iter_steps=200)
labels = [r'$\mu_0^2$', r'$\eta$', r'$\zeta$', r'$\beta$', r'$\theta$']
hgf_tp_df, hgf_mu_df, hgf_sigma_df = hgf_infer.sample_posterior(labels, n_samples=1000)
# +
labels = [r'$\mu_0^2$', r'$\eta$', r'$\zeta$', r'$\beta$', r'$\theta$']
trans_pars_hgf = hgf_tp_df.melt(id_vars='subject', var_name='parameter')
vals = [trans_pars2[:, i].numpy() for i in range(len(labels))]
posterior_accuracy(labels, trans_pars_hgf, vals)
# -
# Plot posterior estimates from simulated data for the HGF agent
# +
plt.figure()
#plot convergence of stochasitc ELBO estimates (log-model evidence)
plt.plot(hgf_infer.loss[-400:])
g = sns.FacetGrid(trans_pars_hgf, col="parameter", height=3, sharey=False);
g = (g.map(sns.lineplot, 'subject', 'value', ci='sd'));
for i in range(len(labels)):
g.axes[0,i].plot(np.arange(2*nsub), trans_pars2[:,i].numpy(),'ro', zorder = 0);
# +
stimulus = sim3.stimulus
stimulus['mask'] = torch.ones(1, 120, 100)
sgf_infer = Horseshoe(sgf_agent, stimulus, sim3.responses)
sgf_infer.infer_posterior(iter_steps=200)
labels = [r'$\rho_1$', r'$h$', r'$\zeta$', r'$\beta$', r'$\theta$']
sgf_tp_df, sgf_mu_df, sgf_sigma_df = sgf_infer.sample_posterior(labels, n_samples=1000)
# +
labels = [r'$\rho_1$', r'$h$', r'$\zeta$', r'$\beta$', r'$\theta$']
trans_pars_sgf = sgf_tp_df.melt(id_vars='subject', var_name='parameter')
vals = [trans_pars3[:, i].numpy() for i in range(len(labels))]
posterior_accuracy(labels, trans_pars_sgf, vals)
# +
plt.figure()
#plot convergence of stochasitc ELBO estimates (log-model evidence)
plt.plot(sgf_infer.loss[-400:])
g = sns.FacetGrid(trans_pars_sgf, col="parameter", height=3, sharey=False);
g = (g.map(sns.lineplot, 'subject', 'value', ci='sd'));
for i in range(len(labels)):
g.axes[0,i].plot(np.arange(2*nsub), trans_pars3[:,i].numpy(),'ro', zorder = 0);
# +
g = sns.PairGrid(sgf_mu_df)
g = g.map_diag(sns.kdeplot)
g = g.map_offdiag(plt.scatter)
g = sns.PairGrid(sgf_sigma_df)
g = g.map_diag(sns.kdeplot)
g = g.map_offdiag(plt.scatter)
# -
#plt.plot(rl_infer.loss[-400:]);
plt.plot(hgf_infer.loss[-400:]);
plt.plot(sgf_infer.loss[-400:]);
# Test model comparison
# +
stimulus = sim1.stimulus
stimulus['mask'] = torch.ones(1, 120, 100)
rl_infer = [Horseshoe(rl_agent, stimulus, sim1.responses),
Horseshoe(rl_agent, stimulus, sim2.responses),
Horseshoe(rl_agent, stimulus, sim3.responses)]
evidences = torch.zeros(3, 3, 2*nsub)
for i in range(3):
rl_infer[i].infer_posterior(iter_steps = 500)
evidences[0, i] = rl_infer[i].get_log_evidence_per_subject()
hgf_infer = [Horseshoe(hgf_agent, stimulus, sim1.responses),
Horseshoe(hgf_agent, stimulus, sim2.responses),
Horseshoe(hgf_agent, stimulus, sim3.responses)]
for i in range(3):
hgf_infer[i].infer_posterior(iter_steps = 500)
evidences[1, i] = hgf_infer[i].get_log_evidence_per_subject()
sgf_infer = [Horseshoe(sgf_agent, stimulus, sim1.responses),
Horseshoe(sgf_agent, stimulus, sim2.responses),
Horseshoe(sgf_agent, stimulus, sim3.responses)]
for i in range(3):
sgf_infer[i].infer_posterior(iter_steps = 500)
evidences[2, i] = sgf_infer[i].get_log_evidence_per_subject()
# -
print((evidences[:, 0].argmax(dim=0) == 0).sum().float()/(2*nsub))
print((evidences[:, 1].argmax(dim=0) == 1).sum().float()/(2*nsub))
print((evidences[:, 2].argmax(dim=0) == 2).sum().float()/(2*nsub))
evidences.sum(-1)
# The diagonal elements in the above matrix are not always the lowest values for the corresponding column, which shows that we cannot accuretly infer the correct model over population, and probably not per subject. More detailed analysis of the possible parameteric models is required.
| examples/social_influence/test_inference_for_rw_hgf_sgf.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3-azureml
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Initial Configurations
# + [markdown] nteract={"transient": {"deleting": false}}
# #### Import Libraries
# + gather={"logged": 1618680718916} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#Import required Libraries
import os
import numpy as np # linear algebra
import pandas as pd # data processing, CSV file I/O (e.g. pd.read_csv)
import seaborn as sns
import matplotlib.pyplot as plt
from matplotlib.image import imread
import cv2
# %matplotlib inline
import warnings
warnings.filterwarnings("ignore")
import azureml.core
import azureml.automl
#from azureml.core.experiment import Experiment
from azureml.core import Workspace, Dataset, Datastore
#Import Model specific libraries
from tensorflow.keras.preprocessing.image import ImageDataGenerator , load_img ,img_to_array
from tensorflow.keras.models import Sequential
from tensorflow.keras.layers import Activation, Dropout, Flatten, Dense, Conv2D, MaxPooling2D
from tensorflow.keras.callbacks import EarlyStopping
from sklearn.metrics import classification_report,confusion_matrix
# + [markdown] nteract={"transient": {"deleting": false}}
# #### Autheticate the AML Workspace
# + gather={"logged": 1618676030870} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#Autheticate the AML Workspace
workspace = Workspace.from_config()
output = {}
output['Subscription ID'] = workspace.subscription_id
output['Workspace Name'] = workspace.name
output['Resource Group'] = workspace.resource_group
output['Location'] = workspace.location
pd.set_option('display.max_colwidth', -1)
outputDf = pd.DataFrame(data = output, index = [''])
outputDf.T
# + gather={"logged": 1618679228079} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#data_folder = os.path.join(os.getcwd(), 'data')
#os.path.join(data_folder,"**/train-images-idx3-ubyte.gz"
#print (os.path.join(data_folder,"castingdata/casting_data/train/def_front/cast_def_0_1001.jpeg"))
image_path = os.path.join(data_folder,"castingdata/casting_data/casting_data/train/ok_front/cast_ok_0_1001.jpeg")
image_path
#os.path.isfile(image_path)
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Get Data
# + [markdown] nteract={"transient": {"deleting": false}}
# #### Upload folders
# + gather={"logged": 1618676641703} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
data_folder = os.path.join(os.getcwd(), 'data')
#/mnt/batch/tasks/shared/LS_root/mounts/clusters/compute-cpu-ds12-v2/code/Users/rabiswas/Manufacturing_Casting_Classification/Notebooks/data
#Create the data directory
os.makedirs(data_folder, exist_ok=True)
#Manually upload the folders from Kaggle
# + [markdown] nteract={"transient": {"deleting": false}}
# #### Visualize the image files
# + gather={"logged": 1618678341856} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# view some casting images which are ok
image_path = os.path.join(data_folder,"castingdata/casting_data/casting_data/train/ok_front/cast_ok_0_1001.jpeg")
img = plt.imread(image_path)
plt.figure(figsize=(12,8))
plt.imshow(img,cmap='gray')
# + gather={"logged": 1618678348706} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# view some casting images which are defective
image_path = os.path.join(data_folder,"castingdata/casting_data/casting_data/train/def_front/cast_def_0_1001.jpeg")
img = plt.imread(image_path)
plt.figure(figsize=(12,8))
plt.imshow(img,cmap='gray')
# + gather={"logged": 1618679355934} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# view some casting images which are ok
image=cv2.imread("/mnt/batch/tasks/shared/LS_root/mounts/clusters/compute-cpu-ds12-v2/code/Users/rabiswas/Manufacturing_Casting_Classification/Notebooks/data/castingdata/casting_data/casting_data/train/ok_front/cast_ok_0_1005.jpeg")
plt.title("OK IMAGE")
cv2.putText(image, "OK_FRONT", (10, 30), cv2.FONT_HERSHEY_SIMPLEX, 0.6, (0, 255, 0), 2)
plt.imshow(image)
# + gather={"logged": 1618679425256} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
# view some casting images which are defective
image=cv2.imread("/mnt/batch/tasks/shared/LS_root/mounts/clusters/compute-cpu-ds12-v2/code/Users/rabiswas/Manufacturing_Casting_Classification/Notebooks/data/castingdata/casting_data/casting_data/train/def_front/cast_def_0_1007.jpeg")
plt.title("DEFECTIVE IMAGE")
cv2.putText(image, "DEFECT_FRONT", (10, 30), cv2.FONT_HERSHEY_SIMPLEX, 0.6, (0, 255, 0), 2)
plt.imshow(image)
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Data Preparation
# + [markdown] nteract={"transient": {"deleting": false}}
# Image Augmentation is a way of applying different types of transformation techniques on actual images, thus producing copies of the same image with alterations. This helps to train deep learning models on more image variations than what is present in the actual dataset.
#
# This is especially useful when we do not have enough images to train the model on, hence we can use augmented images to enlarge the training set and provide more images to the model.
#
# The ImageDataGenerator class in Keras is used for implementing image augmentation. The major advantage of the Keras ImageDataGenerator class is its ability to produce real-time image augmentation. This simply means it can generate augmented images dynamically during the training of the model making the overall mode more robust and accurate.
#
#
# + gather={"logged": 1618680744540} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
image_gen = ImageDataGenerator(rescale=1/255) # Rescale the image by normalizing it
# + gather={"logged": 1618680931477} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
casting_data_dir = os.path.join(data_folder,"castingdata/casting_data/casting_data/")
train_path = casting_data_dir + 'train/'
test_path = casting_data_dir + 'test/'
image_shape = (300,300,1)
batch_size = 32
# + [markdown] nteract={"transient": {"deleting": false}}
# #### flow_from_directory -> Takes the path to a directory & generates batches of augmented data.
#
# #### Arguments used
# ##### directory
# string, path to the target directory. It should contain one subdirectory per class. Any PNG, JPG, BMP, PPM or TIF images inside each of the subdirectories directory tree will be included in the generator. See this script for more details.
# #### target_size
# Tuple of integers (height, width), defaults to (256,256). The dimensions to which all images found will be resized.
# #### color_mode
# One of "grayscale", "rgb", "rgba". Default: "rgb". Whether the images will be converted to have 1, 3, or 4 channels.
# #### classes
# Optional list of class subdirectories (e.g. ['dogs', 'cats']). Default: None. If not provided, the list of classes will be automatically inferred from the subdirectory names/structure under directory, where each subdirectory will be treated as a different class (and the order of the classes, which will map to the label indices, will be alphanumeric). The dictionary containing the mapping from class names to class indices can be obtained via the attribute class_indices.
# #### class_mode
# One of "categorical", "binary", "sparse", "input", or None. Default: "categorical". Determines the type of label arrays that are returned: - "categorical" will be 2D one-hot encoded labels, - "binary" will be 1D binary labels, "sparse" will be 1D integer labels, - "input" will be images identical to input images (mainly used to work with autoencoders). - If None, no labels are returned (the generator will only yield batches of image data, which is useful to use with model.predict()). Please note that in case of class_mode None, the data still needs to reside in a subdirectory of directory for it to work correctly.
# #### batch_size
# Size of the batches of data (default: 32).
# #### shuffle
# Whether to shuffle the data (default: True) If set to False, sorts the data in alphanumeric order.
#
# + gather={"logged": 1618680933727} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#we using keras inbuild function to ImageDataGenerator so we do not need to label all images into 0 and 1 it automatic create it and batch also during trainng
train_set = image_gen.flow_from_directory(train_path,
target_size=image_shape[:2],
color_mode="grayscale",
batch_size=batch_size,
class_mode='binary',shuffle=True)
test_set = image_gen.flow_from_directory(test_path,
target_size=image_shape[:2],
color_mode="grayscale",
batch_size=batch_size,
class_mode='binary',shuffle=False)
# + gather={"logged": 1618680988058} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
train_set.class_indices
# + gather={"logged": 1618681050464} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
test_set.class_indices
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Convolutional model creation
# https://www.tensorflow.org/api_docs/python/tf/keras
#
# #### Keras Sequential Model
# The first way of creating neural networks is with the help of the Keras Sequential Model. The basic idea behind this API is to just arrange the Keras layers in sequential order, this is the reason why this API is called Sequential Model. Even in most of the simple artificial neural networks, layers are put in sequential order, the flow of data takes place between layers in one direction.
#
# Keras sequential model API is useful to create simple neural network architectures without much hassle. The only disadvantage of using the Sequential API is that it doesn’t allow us to build Keras models with multiple inputs or outputs. Instead, it is limited to just 1 input tensor and 1 output tensor.
#
# This flow chart shown below depicts the functioning of Sequential API.
# For adding layers to a sequential model, we can create different types of layers first and then use the add() function for adding them.
#
# #### Keras Convolution Layer
# Keras provides many ready-to-use layer API and Keras convolution layer is just one of them. It is used for creating convolutions over an image in the CNN model.
# For two-dimensional inputs, such as images, they are represented by keras.layers.Conv2D: the Conv2D layer.
#
# ##### Attributes
# __Filters__ represents the number of filters that should be learnt by the convolutional layer. From the schematic drawing above, you should understand that each filter slides over the input image, generating a “feature map” as output.
# The __kernel size__ represents the number of pixels in height and width that should be summarized, i.e. the two-dimensional width and height of the filter.
# The __stride__ tells us how the kernel jumps over the input image. If the stride is 1, it slides pixel by pixel. If it’s two, it jumps one pixel. It jumps two with a stride of 3, and so on.
# The __padding__ tells us what happens when the kernels/filters don’t fit, for example because the input image has a width and height that do not match with the combination of kernel size and stride.
# Depending on the backend you’re using Keras with, the channels (each image has image channels, e.g. 3 channels with Red-Green-Blue or RGB) are in the first dimension or the last. Hence, the __data format__ represents whether it’s a channels first or channels last approach. With recent versions of Keras, which support TensorFlow only, this is no longer a concern.
# If you’re using dilated convolutions, the __dilation rate__ can be specified as well.
# The __activation function__ to which the linear output of the Conv2D layer is fed to make it nonlinear can be specified too.
# A __bias value__ can be added to each layer in order to scale the learnt function vertically. This possibly improves training results. It can be configured here, especially if you don’t want to use biases. By default, it’s enabled.
# The __initializer__ for the kernels, the biases can be configured too, as well as __regularizers__ and __constraints__.
#
# #### Model compilation and fitting the data
# Keras allows you to do so quite easily: with model.compile and model.fit. The compile call allows you to specify the loss function, the optimizer and additional metrics, of which we use accuracy, as it’s intuitive to humans. Then, with fit, we can fit the input_train and target_train (i.e. the inputs and targets of our training set) to the model, actually starting the training process. We do so based on the options that we configured earlier, i.e. batch size, number of epochs, verbosity mode and validation split.
#
# + gather={"logged": 1618681205551} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#Creating model
model = Sequential()
model.add(Conv2D(filters=8, kernel_size=(3,3),input_shape=image_shape, activation='relu',))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(filters=16, kernel_size=(3,3),input_shape=image_shape, activation='relu',))
model.add(MaxPooling2D(pool_size=(2, 2)))
model.add(Conv2D(filters=16, kernel_size=(3,3),input_shape=image_shape, activation='relu',))
model.add(MaxPooling2D(pool_size=(2, 2)))
# + [markdown] nteract={"transient": {"deleting": false}}
# #### Keras Layers
# Keras layers are the building blocks of the Keras library that can be stacked together just like legos for creating neural network models. This ease of creating neural networks is what makes Keras the preferred deep learning framework by many. There are different types of Keras layers available for different purposes while designing your neural network architecture.
#
# At a high-level Keras gives you two choices to create layers by using Keras Layers API and Keras Custom Layers.
#
# 1) Kera Layers API
# Keras provides plenty of pre-built layers for different neural network architectures and purposes via its Keras Layers API.
# These available layers are normally sufficient for creating most of the deep learning models with considerable flexibility.
#
# Below are some of the popular Keras layers –
#
# - Dense Layer
# - Flattened Layer
# - Dropout Layer
# - Reshape Layer
# - Permute Layer
# - RepeatVector Layer
# - Lambda Layer
# - Pooling Layer
# - Locally Connected Layer
#
# 2) Custom Keras Layers
#
# Although Keras Layer API covers a wide range of possibilities it does not cover all types of use-cases. This is why Keras also provides flexibility to create our own custom layer to tailor-make it
# as per our needs.
#
# Below are the ones we used:
# 1. Dense Layer
# Dense Layer is a widely used Keras layer for creating a deeply connected layer in the neural network where each of the neurons of the dense layers receives input from all neurons of the previous layer. At its core, it performs dot product of all the input values along with the weights for obtaining the output.
# The dense layer’s output shape is altered by changing the number of neurons/units specified in the layer. Each and every layer has its own batch size as its first dimension.
# 2. Flatten Layer
# As its name suggests, Flatten Layers is used for flattening of the input. For example, if we have an input shape as (batch_size, 3,3), after applying the flatten layer, the output shape is changed to (batch_size,9).
#
# Flatten function has one argument as follows –
#
# data_format – An optional argument, it mainly helps in preserving weight ordering when data formats are switched.
# 3. Pooling Layer
# The pooling layer is used for applying max pooling operations on temporal data.
#
# The syntax of the pooling layer function is shown below –
#
# The pool_size refers the max pooling windows.
# strides refer the factors for downscale.
#
#
# +
model.add(Flatten())
model.add(Dense(224))
model.add(Activation('relu'))
# Last layer
model.add(Dense(1))
model.add(Activation('sigmoid'))
model.compile(loss='binary_crossentropy',
optimizer='adam',
metrics=['accuracy'])
early_stop = EarlyStopping(monitor='val_loss',patience=2)
# + gather={"logged": 1618689082856} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
results = model.fit_generator(train_set,epochs=20,
validation_data=test_set,
callbacks=[early_stop])
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Analizing model performance
# + gather={"logged": 1618689083523} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
losses = pd.DataFrame(model.history.history)
# + gather={"logged": 1618689084211} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
losses[['loss','val_loss']].plot()
# + gather={"logged": 1618689085196} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
losses[['accuracy','val_accuracy']].plot()
# + gather={"logged": 1618689085750} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#first we will find predict probability
pred_probability = model.predict_generator(test_set)
# + gather={"logged": 1618689086325} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#here it's true label for test set
test_set.classes
# + gather={"logged": 1618689086891} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
predictions = pred_probability > 0.5
#if model predict greater than 0.5 it conveted to 1 means ok_front
# + gather={"logged": 1618689087235} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
print(classification_report(test_set.classes,predictions))
# + gather={"logged": 1618689087691} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
plt.figure(figsize=(10,6))
sns.heatmap(confusion_matrix(test_set.classes,predictions),annot=True)
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Predict on some test images
# + gather={"logged": 1618790606202} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#img = cv2.imread('/mnt/batch/tasks/shared/LS_root/mounts/clusters/compute-cpu-ds12-v2/code/Users/rabiswas/Manufacturing_Casting_Classification/Notebooks/data/castingdata/casting_data/casting_data/test/ok_front/cast_ok_0_1020.jpeg')
img = cv2.imread('/mnt/batch/tasks/shared/LS_root/mounts/clusters/compute-cpu-ds12-v2/code/Users/rabiswas/Manufacturing_Casting_Classification/Notebooks/data/castingdata/casting_data/casting_data/test/ok_front/cast_ok_0_10.jpeg')
img = img/255 #rescaling
pred_img =img.copy()
# + gather={"logged": 1618790609023} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
plt.figure(figsize=(12,8))
plt.imshow(img,cmap='gray')
# + gather={"logged": 1618790776416} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#prediction = model.predict(img.reshape(-1,300,300,1))
prediction = model.predict_classes(img.reshape(-1,300,300,1))
if (prediction.max() < 0.5):
print("def_front")
cv2.putText(pred_img, "def_front", (10, 30), cv2.FONT_HERSHEY_SIMPLEX, 0.6, (0, 255, 0), 2)
else:
print("ok_front")
cv2.putText(pred_img, "ok_front", (10, 30), cv2.FONT_HERSHEY_SIMPLEX, 0.6, (0, 255, 0), 2)
plt.imshow(pred_img,cmap='gray')
plt.axis('off')
plt.show()
# + gather={"logged": 1618790818588} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#img1 = cv2.imread('/mnt/batch/tasks/shared/LS_root/mounts/clusters/compute-cpu-ds12-v2/code/Users/rabiswas/Manufacturing_Casting_Classification/Notebooks/data/castingdata/casting_data/casting_data/test/def_front/cast_def_0_1134.jpeg')
img1 = cv2.imread('/mnt/batch/tasks/shared/LS_root/mounts/clusters/compute-cpu-ds12-v2/code/Users/rabiswas/Manufacturing_Casting_Classification/Notebooks/data/castingdata/casting_data/casting_data/test/def_front/cast_def_0_1059.jpeg')
img1 = img1/255
pred_img1 =img1.copy()
# + gather={"logged": 1618790789342} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
plt.figure(figsize=(12,8))
plt.imshow(img1,cmap='gray')
# + gather={"logged": 1618790829556} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#prediction = model.predict(img1.reshape(-1,300,300,1))
prediction = model.predict_classes(img1.reshape(-1,300,300,1))
if (prediction.max()<0.5):
print("def_front")
cv2.putText(pred_img1, "def_front", (10, 30), cv2.FONT_HERSHEY_SIMPLEX, 0.6, (0, 255, 0), 2)
else:
print("ok_front")
cv2.putText(pred_img1, "ok_front", (10, 30), cv2.FONT_HERSHEY_SIMPLEX, 0.6, (0, 255, 0), 2)
plt.imshow(pred_img1,cmap='gray')
plt.axis('off')
plt.show()
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Saving the model
# + gather={"logged": 1618759431983} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
model.save('inspection_of_casting_products.h5')
# + [markdown] nteract={"transient": {"deleting": false}}
# ### Model evaluation
# The final step is to evaluate our model after we performed training. Keras allows you to do so with model.evaluate. We are using the unseen test datset. This way, we can be sure that we test the model with data that it hasn’t seen before during training, evaluating its power to generalize to new data. Evaluation is done in a non-verbose way, and the results are printed on screen.
# + gather={"logged": 1618760074920} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
def display_eval_metrics(e_data):
msg='Model Metrics after Training'
#print_in_color(msg, (255,255,0), (55,65,80))
msg='{0:^24s}{1:^24s}'.format('Metric', 'Value')
#print_in_color(msg, (255,255,0), (55,65,80))
for key,value in e_data.items():
print (f'{key:^24s}{value:^24.5f}')
acc=e_data['accuracy']* 100
return acc
# + gather={"logged": 1618760276268} jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
#Accuracy on the test set
e_dict=model.evaluate( test_set, batch_size=batch_size, verbose=1, steps=None, return_dict=True)
acc=display_eval_metrics(e_dict)
print('accuracy of model on the test set is %5.2f' %acc)
# + jupyter={"outputs_hidden": false, "source_hidden": false} nteract={"transient": {"deleting": false}}
| notebook/Casting_Product_Classification.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import unittest
import numpy as np
import pandas as pd
import numpy.testing as np_testing
import pandas.testing as pd_testing
import os
import sys
import import_ipynb
from sklearn.cluster import KMeans
class Test(unittest.TestCase):
def _dirname_if_file(self, filename):
if os.path.isdir(filename):
return filename
else:
return os.path.dirname(os.path.abspath(filename))
def setUp(self):
import Exercise2_02
self.exercise = Exercise2_02
self.data = pd.read_csv('circles.csv')
def test_input_frames(self):
pd_testing.assert_frame_equal(self.exercise.data, self.data)
def test_kmeans(self):
est_kmeans = KMeans(n_clusters=5, random_state=0)
est_kmeans.fit(self.data)
pred_kmeans = est_kmeans.predict(self.data)
np_testing.assert_equal(pred_kmeans, self.exercise.pred_kmeans)
if __name__ == '__main__':
unittest.main(argv=['first-arg-is-ignored'], exit=False)
| Chapter02/Exercise2.02/.ipynb_checkpoints/test_exercise2_02-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import numpy as np
# fix random seed for reproducibility
seed = 7
np.random.seed(seed)
# +
from keras.layers import Input, Dense
from keras.models import Model
from keras import regularizers
def build_model(data):
# this is the size of our encoded representations
encoding_dim = 32 # 32 floats -> compression of factor 24.5, assuming the input is 784 floats
# this is our input placeholder
original_dim=data.shape[1]
input_img = Input(shape=(original_dim,))
# "encoded" is the encoded representation of the input
encoded = Dense(256, activation='tanh',
activity_regularizer=regularizers.l1(2*10e-5))(input_img)
encoded = Dense(128, activation='tanh',
activity_regularizer=regularizers.l1(2*10e-5))(encoded)
encoded = Dense(32, activation='tanh',
activity_regularizer=regularizers.l1(2*10e-5))(encoded)
decoded = Dense(128, activation='tanh')(encoded)
decoded = Dense(256, activation='tanh')(decoded)
decoded = Dense(original_dim, activation='tanh')(decoded)
# this model maps an input to its reconstruction
autoencoder = Model(input_img, decoded)
# this model maps an input to its encoded representation
encoder = Model(input_img, encoded)
autoencoder.compile(optimizer='adadelta', loss='binary_crossentropy')
autoencoder.fit(data, data,
epochs=10,
batch_size=100,
shuffle=True)
return encoder
# -
from nilearn.decomposition import CanICA
def prepare_data(func_filenames):
canica = CanICA(n_components=20, smoothing_fwhm=6.,
memory="nilearn_cache", memory_level=2,
threshold=3., verbose=10, random_state=0)
data=canica.prepare_data(func_filenames)
return data
# +
from nilearn.connectome import ConnectivityMeasure
def corr(all_time_series):
connectivity_biomarkers = {}
conn_measure = ConnectivityMeasure(kind='correlation', vectorize=True)
connectivity_biomarkers = conn_measure.fit_transform(all_time_series)
return connectivity_biomarkers
# +
from sklearn.neural_network import MLPClassifier
from sklearn.neighbors import KNeighborsClassifier
from sklearn.svm import SVC
from sklearn.gaussian_process import GaussianProcessClassifier
from sklearn.gaussian_process.kernels import RBF
from sklearn.tree import DecisionTreeClassifier
from sklearn.ensemble import RandomForestClassifier, AdaBoostClassifier
from sklearn.naive_bayes import GaussianNB
from sklearn.discriminant_analysis import QuadraticDiscriminantAnalysis
def classify(train_X,train_Y, test_X, test_Y):
names = ["Nearest Neighbors", "Linear SVM", "RBF SVM", "Gaussian Process",
"Decision Tree", "Random Forest", "Neural Net", "AdaBoost",
"Naive Bayes", "QDA"]
classifiers = [
KNeighborsClassifier(3),
SVC(kernel="linear", C=0.025),
SVC(gamma=2, C=1),
GaussianProcessClassifier(1.0 * RBF(1.0)),
DecisionTreeClassifier(max_depth=5),
RandomForestClassifier(max_depth=5, n_estimators=10, max_features=1),
MLPClassifier(alpha=1),
AdaBoostClassifier(),
GaussianNB(),
QuadraticDiscriminantAnalysis()]
scores = []
for name, clf in zip(names, classifiers):
clf.fit(train_X, train_Y)
score=clf.score(test_X,test_Y)
scores.append(score)
return scores
# -
from sklearn.model_selection import StratifiedKFold
from keras import backend as K
def CV(X,Y):
n_subjects=range(0,len(Y))
# define 10-fold cross validation test harness
kfold = StratifiedKFold(n_splits=5, shuffle=True, random_state=seed)
cvscores = []
for train,test in kfold.split(n_subjects,Y):
train_X = [X[i*20:i*20+19] for i in train]
train_Y = [Y[i] for i in train]
test_Y = [Y[i] for i in test]
model=build_model(np.vstack(train_X))
train_D = [model.predict(X[i*20:i*20+19]) for i in train]
test_D = [model.predict(X[i*20:i*20+19]) for i in test]
#Release GPU memory after model is used
K.clear_session()
train_FC=corr(train_D)
test_FC=corr(test_D)
score=classify(train_FC,train_Y, test_FC, test_Y)
cvscores.append(score)
return cvscores
# +
from nilearn import datasets
adhd_dataset = datasets.fetch_adhd(n_subjects=40,data_dir='/home/share/TmpData/Qinglin/nilearn_data/')
X = prepare_data(adhd_dataset.func) # list of 4D nifti files for each subject
Y = adhd_dataset.phenotypic['adhd']
cvscores=CV(X,Y)
# -
cvscores
| experiments/.ipynb_checkpoints/CV-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # 'open-visualizations' for repeated measures in Python - #2
#
# *<NAME>*
# *<EMAIL>*
# *19-03-2020*
# ## Background
#
# This tutorial is a follow up on my ['Open-visualizations tutorial for repeated measures in R'](https://github.com/jorvlan/open-visualizations/tree/master/R) and contributes to a GitHub repository called ['open-visualizations'](https://github.com/jorvlan/open-visualizations).
#
# Next to this notebook, I have also created another tutorial in Python with a slightly different approach which includes R-like behavior with `plotnine`.
# See ['open-visualizations'](https://github.com/jorvlan/open-visualizations) to view that tutorial.
#
# If you have any questions, or suggestions for improvement, please open an issue in the GitHub repository [open-visualizations](https://github.com/jorvlan/open-visualizations).
#
# If you use my repository for your research, please reference it.
#
#
#
#
# ## Load libraries
import numpy as np
import pandas as pd
import seaborn as sns
import matplotlib.pyplot as plt
# ## Activate folder where figures are stored
#
# - By default, figures will not be saved. If you want to save figures, set savefig to `True`.
# +
savefig = False
if savefig:
#Load libraries
import os
from os.path import isdir
#Get current working directory, but you can specify your own directory of course.
cwd = os.getcwd()
if os.path.exists(cwd + "/repmes_tutorial_2_python/figs"):
print("Directory already exists")
#Assign the existing directory to a variable
fig_dir = cwd + "/repmes_tutorial_2_python/figs"
elif not os.path.exists(cwd + "/repmes_tutorial_2_python/figs"):
print("Directory does not exist and will be created ......")
os.makedirs(cwd + "/repmes_tutorial_2_python/figs")
if isdir(cwd + "/repmes_tutorial_2_python/figs"):
print('Directory was created succesfully')
#Assign the created directory to a variable
fig_dir = cwd + "/repmes_tutorial_2_python/figs"
else:
print("Something went wrong")
# -
# ## Initialize a dataset
# +
# Create a dummy dataset
N=30
np.random.seed(3)
data = np.random.normal(size=(N,))
#Create the dataframe in a wide format with 'Before' and 'After ' as columns
df = pd.DataFrame({'Before': data,
'After': data+1})
#Set the amount of jitter and create a dataframe containing the jittered x-axis values
jitter_1 = 0
np.random.seed(3)
df_jitter_1 = pd.DataFrame(np.random.normal(loc=0, scale=jitter_1, size=df.values.shape), columns=df.columns)
#Update the dataframe with adding a number based on the length on the columns. Otherwise all datapoints would be at the same x-axis location.
df_jitter_1 += np.arange(len(df.columns))
#Inspect the created dataframe
pd.options.display.float_format = '{:.3f}'.format
print("The dataframe with 2 variables ")
print(df[['Before', 'After']])
# -
# ## Figure 1
#
# - In Figure 1, we only display the individual datapoints.
# Define pre-settings
w = 6
h = 6
title_size = 20
xlab_size = 15
ylab_size = 20
labels = ['Before', 'After']
# +
# Create empty figure and plot the individual datapoints
fig, ax = plt.subplots(figsize=(15,9))
for col in df:
ax.plot(df_jitter_1[col], df[col], 'o', alpha=.6, zorder=2, ms=10, mew=1.5)
#Additional settings
ax.set_xticks(range(len(df.columns)))
ax.set_xticklabels((labels), size= xlab_size)
ax.set_xlim(-1, len(df.columns))
ax.set_ylabel('Value', size = ylab_size)
ax.set_title('Figure 1: individual datapoints', size = title_size)
sns.despine()
if savefig:
plt.savefig(fig_dir + "/figure1.png", width = w, height = h)
# -
# ## Figure 2
# +
# Create empty figure and plot the individual datapoints
fig, ax = plt.subplots(figsize=(15,9))
for col in df:
ax.plot(df_jitter_1[col], df[col], 'o', alpha=.6, zorder=2, ms=10, mew=1.5)
for idx in df.index:
ax.plot(df_jitter_1.loc[idx,['Before','After']], df.loc[idx,['Before','After']], color = 'gray', linewidth = 2, linestyle = '--',alpha = .3)
#Additonal settings
ax.set_xticks(range(len(df.columns)))
ax.set_xticklabels((labels), size= xlab_size)
ax.set_xlim(-1, len(df.columns))
ax.set_ylabel('Value', size = ylab_size)
ax.set_title('Figure 2: individual datapoints with lines', size = title_size)
sns.despine()
if savefig:
plt.savefig(fig_dir + "/figure2.png", width = w, height = h)
# -
# ## Figure 3
# +
#Set the amount of jitter and create a dataframe containing the jittered x-axis values
jitter_2 = 0.05
np.random.seed(3)
df_jitter_2 = pd.DataFrame(np.random.normal(loc=0, scale=jitter_2, size=df.values.shape), columns=df.columns)
#Update the dataframe with adding a number based on the length on the columns. Otherwise all datapoints would be at the same x-axis location.
df_jitter_2 += np.arange(len(df.columns))
# Create empty figure and plot the individual datapoints
fig, ax = plt.subplots(figsize=(15,9))
for col in df:
ax.plot(df_jitter_2[col], df[col], 'o', alpha=.6, zorder=2, ms=10, mew=1.5)
for idx in df.index:
ax.plot(df_jitter_2.loc[idx,['Before','After']], df.loc[idx,['Before','After']], color = 'gray', linewidth = 2, linestyle = '--',alpha = .3)
#Additonal settings
ax.set_xticks(range(len(df.columns)))
ax.set_xticklabels((labels), size= xlab_size)
ax.set_xlim(-1, len(df.columns))
ax.set_ylabel('Value', size = ylab_size)
ax.set_title('Figure 3: individual datapoints with lines and jitter', size = title_size)
sns.despine()
if savefig:
plt.savefig(fig_dir + "/figure3.png", width = w, height = h)
# -
# ## Figure 4
# +
#Merge dataframe from wide to long for sns.pointplot
df_long = pd.melt(df, value_vars=['Before','After'])
# Create empty figure and plot the individual datapoints
fig, ax = plt.subplots(figsize=(15,9))
for col in df:
ax.plot(df_jitter_2[col], df[col], 'o', alpha=.6, zorder=2, ms=10, mew=1.5)
for idx in df.index:
ax.plot(df_jitter_2.loc[idx,['Before','After']], df.loc[idx,['Before','After']], color = 'gray', linewidth = 2, linestyle = '--',alpha = .3)
sns.pointplot(x='variable', y='value', ci=95, data=df_long, join=False, scale=1.5, color = 'black', capsize = .03) #palette = 'Paired'
#Additonal settings
ax.set_xticks(range(len(df.columns)))
ax.set_xticklabels((labels), size= xlab_size)
ax.set_xlim(-1, len(df.columns))
ax.set_ylabel('Value', size = ylab_size)
ax.set_title('Figure 4: individual datapoints with lines, jitter and statistics', size = title_size)
sns.despine()
if savefig:
plt.savefig(fig_dir + "/figure4.png", width = w, height = h)
# -
# ## Figure 5
# +
# Create empty figure and plot the individual datapoints
fig, ax = plt.subplots(figsize=(15,9))
for col in df:
ax.plot(df_jitter_2[col], df[col], 'o', alpha=1, zorder=2, ms=10, mew=1.5)
for idx in df.index:
ax.plot(df_jitter_2.loc[idx,['Before','After']], df.loc[idx,['Before','After']], color = 'gray', linewidth = 2, linestyle = '--',alpha = .3)
sns.pointplot(x='variable', y='value', ci=95, data=df_long, join=False, scale=0.01, color = 'black', capsize = .03)
sns.violinplot(x='variable', y='value', data=df_long, hue = 'variable', split = True, inner = 'quartile', cut=1)
#Additonal settings
ax.set_xticks(range(len(df.columns)))
ax.set_xticklabels((labels), size= xlab_size)
ax.set_xlim(-1, len(df.columns))
ax.set_ylabel('Value', size = ylab_size)
ax.set_title('Figure 5: individual datapoints, lines, jitter, statistics, violins', size = title_size)
ax.legend_.remove()
sns.despine()
plt.setp(ax.collections, alpha=.02)
if savefig:
plt.savefig(fig_dir + "/figure5.png", width = w, height = h)
# -
# ## Figure 6
# +
#Create a dataframe do display 4 conditions
df_2 = pd.DataFrame({'Before': data,
'After': data+1,
'Before1': data,
'After1': data-1})
df_jitter_3 = pd.DataFrame(np.random.normal(loc=0, scale=jitter_2, size=df_2.values.shape), columns=df_2.columns)
df_jitter_3
#Do an additional step to create a jittered values for the 4 columns.. i.e., jitter values around condition 1 and 2 + jitter values for condition 3 and 4.
df_jitter_3 += np.arange(len(df_2.columns))
df_jitter_3
# Create empty figure and plot the individual datapoints
fig, ax = plt.subplots(figsize=(15,9))
for col in df_2:
ax.plot(df_jitter_3[col], df_2[col], 'o', alpha=.6, zorder=2, ms=10, mew=1.5)
for idx in df_2.index:
ax.plot(df_jitter_3.loc[idx,['Before','After']], df_2.loc[idx,['Before','After']], color = 'gray', linewidth = 2, linestyle = '--',alpha = .3)
ax.plot(df_jitter_3.loc[idx,['Before1','After1']], df_2.loc[idx,['Before1','After1']], color = 'gray', linewidth = 2, linestyle = '--', alpha =.3)
#Additonal settings
ax.set_xticks(range(len(df.columns)))
ax.set_xticklabels((['Before', 'After', 'Before', 'After']), size= xlab_size)
ax.set_xlim(-1, len(df.columns))
ax.set_ylabel('Value', size = ylab_size)
ax.set_title('Figure 6: individual datapoints with lines, jitter: 4 conditions', size = title_size)
sns.despine()
plt.setp(ax.collections, alpha=.02)
plt.setp(ax, xticks=[0, 1, 2, 3, 4])
if savefig:
plt.savefig(fig_dir + "/figure6.png", width = w, height = h)
# -
# ## Figure 7
# +
#Merge dataframe from wide to long for sns.pointplot
df_long_2 = pd.melt(df_2, value_vars=['Before','After', 'Before1', 'After1'])
# Create empty figure and plot the individual datapoints
fig, ax = plt.subplots(figsize=(15,9))
for col in df_2:
ax.plot(df_jitter_3[col], df_2[col], 'o', alpha=.6, zorder=2, ms=10, mew=1.5)
for idx in df_2.index:
ax.plot(df_jitter_3.loc[idx,['Before','After']], df_2.loc[idx,['Before','After']], color = 'gray', linewidth = 2, linestyle = '--',alpha = .3)
ax.plot(df_jitter_3.loc[idx,['Before1','After1']], df_2.loc[idx,['Before1','After1']], color = 'gray', linewidth = 2, linestyle = '--', alpha = .3)
sns.pointplot(x='variable', y='value', ci=95, data=df_long_2, join=False, scale=1.5, color = 'black', capsize = .03)
#Additonal settings
ax.set_xticks(range(len(df.columns)))
ax.set_xticklabels((['Before', 'After', 'Before', 'After']), size= xlab_size)
ax.set_xlim(-1, len(df.columns))
ax.set_ylabel('Value', size = ylab_size)
ax.set_title('Figure 7: individual datapoints with lines, jitter, statistics: 4 conditions', size = title_size)
sns.despine()
plt.setp(ax, xticks=[0, 1, 2, 3, 4])
if savefig:
plt.savefig(fig_dir + "/figure7.png", width = w, height = h)
# -
# ## Figure 8
# +
# Create empty figure and plot the individual datapoints
fig, ax = plt.subplots(figsize=(15,9))
for col in df:
ax.plot(df_jitter_2[col], df[col], 'o', alpha=.8, zorder=2, ms=10, mew=1.5)
for idx in df.index:
ax.plot(df_jitter_2.loc[idx,['Before','After']], df.loc[idx,['Before','After']], color = 'gray', linewidth = 2, linestyle = '-',alpha = .2)
for value in df_long_2:
sns.violinplot(x='variable', y='value', data=df_long, hue = 'variable', split = True, inner = 'quartile', cut=1, dodge = True)
sns.boxplot(x='variable', y='value', data=df_long, hue = 'variable', dodge = True, width = 0.2, fliersize = 2)
#Additonal settings
ax.set_xticks(range(len(df.columns)))
ax.set_xticklabels((['Before', 'After']), size= xlab_size)
ax.set_xlim(-1, len(df.columns))
ax.set_ylabel('Value', size = ylab_size)
ax.set_title('Figure 8: individual datapoints with lines, jitter, statistics, box- and violin', size = title_size)
sns.despine()
ax.legend_.remove()
plt.setp(ax.collections, alpha=.1)
if savefig:
plt.savefig(fig_dir + "/figure8.png", width = w, height = h)
# -
# ## General remarks / tips
# - If you want to save your figures in a high-quality manner for e.g., publications, you could save your figure with a `.tif` extension and add `dpi=` as used in the following line of code:
#
# `plt.savefig("/figure.tif", width = w, height = h, dpi = 600)`
# ## That's it! (for now)
#
# You have reached the end of this document.
#
# I hope you'll be able to use this tutorial to create more `open-visualizations` for your research!
#
# If you use this tutorial, please cite it in your work (see DOI above).
#
#
# [open-visualizations](https://github.com/jorvlan/open-visualizations) for repeated measures in `R` and `Python` by [**<NAME>**](https://jordyvanlangen.com)
| Python/tutorial_2/repeated_measures_python_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# 引入所需要的全部包
from sklearn.model_selection import train_test_split # 数据划分的类
from sklearn.linear_model import LinearRegression # 线性回归的类
from sklearn.preprocessing import StandardScaler # 数据标准化
import numpy as np
import matplotlib as mpl
import matplotlib.pyplot as plt
import pandas as pd
from pandas import DataFrame
import time
# -
## 设置字符集,防止中文乱码
mpl.rcParams['font.sans-serif']=[u'simHei']
mpl.rcParams['axes.unicode_minus']=False
# 加载数据
# 日期、时间、有功功率、无功功率、电压、电流、厨房用电功率、洗衣机用电功率、热水器用电功率
path1='datas/household_power_consumption_1000.txt'
df = pd.read_csv(path1, sep=';', low_memory=False)#没有混合类型的时候可以通过low_memory=F调用更多内存,加快效率)
df.head() ## 获取前五行数据查看查看
# 查看格式信息
df.info()
# 异常数据处理(异常数据过滤)
new_df = df.replace('?', np.nan)#替换非法字符为np.nan
datas = new_df.dropna(axis=0, how = 'any') # 只要有一个数据为空,就进行行删除操作
datas.describe().T#观察数据的多种统计指标(只能看数值型的)
# 查看格式信息
df.info()
## 创建一个时间函数格式化字符串
def date_format(dt):
# dt显示是一个series/tuple;dt[0]是date,dt[1]是time
import time
t = time.strptime(' '.join(dt), '%d/%m/%Y %H:%M:%S')
return (t.tm_year, t.tm_mon, t.tm_mday, t.tm_hour, t.tm_min, t.tm_sec)
## 需求:构建时间和功率之间的映射关系,可以认为:特征属性为时间;目标属性为功率值。
# 获取x和y变量, 并将时间转换为数值型连续变量
X = datas.iloc[:,0:2]
X = X.apply(lambda x: pd.Series(date_format(x)), axis=1)
Y = datas['Global_active_power']
X.head(2)
## 对数据集进行测试集合训练集划分
# X:特征矩阵(类型一般是DataFrame)
# Y:特征对应的Label标签(类型一般是Series)
# test_size: 对X/Y进行划分的时候,测试集合的数据占比, 是一个(0,1)之间的float类型的值
# random_state: 数据分割是基于随机器进行分割的,该参数给定随机数种子;给一个值(int类型)的作用就是保证每次分割所产生的数数据集是完全相同的
X_train,X_test,Y_train,Y_test = train_test_split(X, Y, test_size=0.2, random_state=0)
print(X_train.shape)
print(X_test.shape)
print(Y_train.shape)
# 查看训练集上的数据信息(X)
X_train.describe()
## 数据标准化
# StandardScaler:将数据转换为标准差为1的数据集(有一个数据的映射)
# scikit-learn中:如果一个API名字有fit,那么就有模型训练的含义,默认是没法返回值
# scikit-learn中:如果一个API名字中有transform, 那么就表示对数据具有转换的含义操作
# scikit-learn中:如果一个API名字中有predict,那么就表示进行数据预测,会有一个预测结果输出
# scikit-learn中:如果一个API名字中既有fit又有transform的情况下,那就是两者的结合(先做fit,再做transform)
ss = StandardScaler() # 模型对象创建
# ss.fit(X_train) # 模型训练
# X_train = xx.transform(X_train) # 对训练集合数据进行转换
X_train = ss.fit_transform(X_train) # 训练模型并转换训练集
X_test = ss.transform(X_test) ## 直接使用在模型构建数据上进行一个数据标准化操作 (测试集)
pd.DataFrame(X_train).describe()
# +
## 模型训练
lr = LinearRegression(fit_intercept=True) # 模型对象构建
lr.fit(X_train, Y_train) ## 训练模型
## 模型校验
y_predict = lr.predict(X_test) ## 预测结果
print("训练集上R2:",lr.score(X_train, Y_train))
print("测试集上R2:",lr.score(X_test, Y_test))
mse = np.average((y_predict-Y_test)**2)
rmse = np.sqrt(mse)
print("rmse:",rmse)
# -
# 输出模型训练得到的相关参数
print("模型的系数(θ):", end="")
print(lr.coef_)
print("模型的截距:", end='')
print(lr.intercept_)
# +
## 模型保存/持久化
# 在机器学习部署的时候,实际上其中一种方式就是将模型进行输出;另外一种方式就是直接将预测结果输出
# 模型输出一般是将模型输出到磁盘文件
from sklearn.externals import joblib
# 保存模型要求给定的文件所在的文件夹比较存在
joblib.dump(ss, "result/data_ss.model") ## 将标准化模型保存
joblib.dump(lr, "result/data_lr.model") ## 将模型保存
# +
# 加载模型
ss3 = joblib.load("result/data_ss.model") ## 加载模型
lr3 = joblib.load("result/data_lr.model") ## 加载模型
# 使用加载的模型进行预测
data1 = [[2006, 12, 17, 12, 25, 0]]
data1 = ss3.transform(data1)
print(data1)
lr3.predict(data1)
# -
## 预测值和实际值画图比较
t=np.arange(len(X_test))
plt.figure(facecolor='w')#建一个画布,facecolor是背景色
plt.plot(t, Y_test, 'r-', linewidth=2, label='真实值')
plt.plot(t, y_predict, 'g-', linewidth=2, label='预测值')
plt.legend(loc = 'upper left')#显示图例,设置图例的位置
plt.title("线性回归预测时间和功率之间的关系", fontsize=20)
plt.grid(b=True)#加网格
plt.show()
# +
## 功率和电流之间的关系
X = datas.iloc[:,2:4]
Y2 = datas.iloc[:,5]
## 数据分割
X2_train,X2_test,Y2_train,Y2_test = train_test_split(X, Y2, test_size=0.2, random_state=0)
## 模型训练
lr2 = LinearRegression(fit_intercept=False)
lr2.fit(X2_train, Y2_train) ## 训练模型
## 结果预测
Y2_predict = lr2.predict(X2_test)
## 模型评估
print("电流预测R2: ", lr2.score(X2_test,Y2_test))
print("电流参数:", lr2.coef_)
print("模型的截距:", end='')
print(lr2.intercept_)
## 绘制图表
#### 电流关系
t=np.arange(len(X2_test))
plt.figure(facecolor='w')
plt.plot(t, Y2_test, 'r-', linewidth=2, label=u'真实值')
plt.plot(t, Y2_predict, 'g-', linewidth=2, label=u'预测值')
plt.legend(loc = 'lower right')
plt.title(u"线性回归预测功率与电流之间的关系", fontsize=20)
plt.grid(b=True)
plt.show()
# -
| jupyter_book/basic_ml/01_home power usage user linear regression.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
from nldi_xstool.XSGen import XSGen
from nldi_xstool.ancillary import queryDEMsShape, getGageDatum
from nldi_xstool.nldi_xstool import getXSAtPoint, getXSAtEndPts
from shapely.geometry import LineString, Point
import py3dep
from pynhd import NLDI, NHDPlusHR, WaterData
import xarray as xr
from matplotlib import pyplot as plt
import numpy as np
import pandas as pd
import dataretrieval.nwis as nwis
import requests
# import plotly.express as px
import folium
from folium import plugins
import time
from helpers import plotGageLocation, plotGageXS, interpTValues
# -
# # Demonstration of NGHF web services and tools in support of other USGS projects: NGWOS R&D Thresholds project
#
# This demo shows the use of the ndli_xstool, a tool for cross-section extraction along NHDPlus flowlines or given end-points. The nldi_xstool, makes use of 3DEP elevation services for elevation. In this demo, a portion of the Thresholds project excel spreadsheet was written to a datastructure and the information is used to located Threshold project gages, extract a topographic cross-section from user defined endpoints, and plot the cross-section along with it's "thresholds" and plot the data in map view for context.
# ### Create Dictionary of Threshold gage sites where path is location of left-bank and right-bank, both estimated using google earth
# +
gage_path = {'02334480': {'path': [(-84.069592, 34.132959), (-84.070461, 34.131941)],
'name': 'RICHLAND CREEK AT SUWANEE DAM ROAD, NEAR BUFORD,GA'},
'02335350': {'path': [(-84.265284, 33.964690), (-84.264037, 33.965374)],
'name': 'Crooked Creek near Norcross, GA'},
'02207055': {'path': [(-84.059534, 33.824598), (-84.058593, 33.825617)],
'name': 'Jacks Creek at Brannan Road, near Snellville, GA'},
'03321350': {'path': [(-86.888014, 37.618517), (-86.887195, 37.619308)],
'name': 'SOUTH FORK PANTHER CREEK NEAR WHITESVILLE, KY'},
'06811500': {'path': [(-95.812660, 40.392760), (-95.814296, 40.392806)],
'name': 'Little Nemaha River at Auburn, NE'}
}
gage_thresholds = {'02334480': {'Thresholds': {1: {'Type':'Road', 'Value': 17.73, 'Name': 'Street: Suwanee Dam Road',
'lat': 34.132595, 'lon': -84.070117},
2: {'Type':'Top Of Bank', 'Value': 8.85, 'Name': 'Bankfull',
'lat': 34.132561, 'lon': -84.069978}}},
'02335350': {'Thresholds': {1: {'Type':'Top Of Bank', 'Value': 11.37, 'Name': 'Bankfull',
'lat': 33.965156, 'lon': -84.265043},
2: {'Type':'Bridge Deck', 'Value': 19.53, 'Name': 'Bridge Deck Elevation',
'lat': 33.965174, 'lon': -84.264742}}},
'02207055': {'Thresholds': {1: {'Type':'Top Of Bank', 'Value': 2.66, 'Name': 'Bankfull',
'lat': 33.825173, 'lon': -84.058902},
2: {'Type':'Bridge Deck', 'Value': 3.37, 'Name': 'Bridge Deck Elevation',
'lat': 33.82524, 'lon': -84.058999}}},
'03321350': {'Thresholds': {1: {'Type':'Top Of Bank', 'Value': 15.5, 'Name': 'Bankfull',
'lat': 37.61973838, 'lon': -86.8893671}}},
'06811500': {'Thresholds': {1: {'Type':'Bridge Lower Chord', 'Value': 32.5, 'Name': 'Bottom of Bridge Steel',
'lat': 40.392669, 'lon': -95.812861}}}
}
# -
# ## Iterate through dictionary to calculate:
# * Gage Datum (Convert NGDV29 to NAVD88 if necessary)
# * Gage Location from NWIS
# * Gage Comid - may not be necessary
# * 3DEP DEM dictionary of available resolution to provide context to quality of cross-section
# * Cross-section using NHGF projects NLDI_XSTools package (soon to be web-service) returned as Geopandas Dataframe
# * DEM surrounding cross-section to compare measured threshold values with values interpolated off the DEM. This should provide some context to the quality of the cross-section interpolated off the DEM.
#
# NOTE: NWIS package throws errors randomly at times in function getGageDatum
gage_datum_m = []
gage_location = []
gage_comid = []
dem_res = []
dem = []
xs_bbox = []
cross_sections = []
for index, (k,v) in enumerate(gage_path.items()):
gage_datum_m.append(getGageDatum(k, verbose=False))
gage_location.append(NLDI().getfeature_byid('nwissite', f'USGS-{k}'))
gage_comid.append(gage_location[index].comid.values.astype(int)[0])
x = gage_location[index].geometry.x[0]
y = gage_location[index].geometry.y[0]
path = v['path']
cross_sections.append(getXSAtEndPts(path, 101, res=1.0))
lnst = []
for pt in path:
lnst.append(Point(pt[0], pt[1]))
bbox = LineString(lnst).envelope.bounds
dem_res.append(queryDEMsShape(bbox))
# buffer below is a quick hack for now ~ 200m
xs_bbox.append(LineString(lnst).envelope.buffer(.002))
dem.append(py3dep.get_map("DEM",
xs_bbox[index].bounds,
resolution=1,
geo_crs="epsg:4326",
crs="epsg:4326"))
print(k, x, y, gage_comid[index], gage_datum_m[index])
# ## For each gage a plan-view map is provided for context along with the cross-section and thresholds plot
# * Click on Markers to identify
# * Mouse location is indicated in lower right of Map and could be used to adjust position of cross-section end points in gage_thresholds dictionary above
#
# +
index = 0
m = plotGageLocation(index=index,
gage_location=gage_location,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections)
plotGageXS(index=index,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections,
gage_datum=gage_datum_m,
dem = dem,
dem_res=dem_res)
m
# +
index = 1
m = plotGageLocation(index=index,
gage_location=gage_location,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections)
plotGageXS(index=index,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections,
gage_datum=gage_datum_m,
dem = dem,
dem_res=dem_res)
m
# +
index = 2
m = plotGageLocation(index=index,
gage_location=gage_location,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections)
plotGageXS(index=index,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections,
gage_datum=gage_datum_m,
dem = dem,
dem_res=dem_res)
m
# +
index = 3
m = plotGageLocation(index=index,
gage_location=gage_location,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections)
plotGageXS(index=index,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections,
gage_datum=gage_datum_m,
dem = dem,
dem_res=dem_res)
m
# +
index = 4
m = plotGageLocation(index=index,
gage_location=gage_location,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections)
plotGageXS(index=index,
gage_path=gage_path,
gage_thresholds=gage_thresholds,
cross_sections=cross_sections,
gage_datum=gage_datum_m,
dem = dem,
dem_res=dem_res)
m
# -
# ### Check measured thresholds against DEM interpolated values to provide context to the quality of the DEM interpolated cross-section
# + jupyter={"outputs_hidden": true} tags=[]
interpTValues(gage_path=gage_path,
gage_thresholds=gage_thresholds,
gage_datum_m=gage_datum_m,
dem=dem)
# -
| notebooks/Threshold_XSTool_Demo.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creating a Simple Model for SEI Growth
# Before adding a new model, please read the [contribution guidelines](https://github.com/pybamm-team/PyBaMM/blob/master/CONTRIBUTING.md)
# In this notebook, we will run through the steps involved in creating a new model within pybamm. We will then solve and plot the outputs of the model. We have chosen to implement a very simple model of SEI growth. We first give a brief derivation of the model and discuss how to nondimensionalise the model so that we can show the full process of model conception to solution within a single notebook.
# Note: if you run the entire notebook and then try to evaluate the earlier cells, you will likely receive an error. This is because the state of objects is mutated as it is passed through various stages of processing. In this case, we recommend that you restart the Kernel and then evaluate cells in turn through the notebook.
# ## A Simple Model of Solid Electrolyte Interphase (SEI) Growth
# The SEI is a porous layer that forms on the surfaces of negative electrode particles from the products of electrochemical reactions which consume lithium and electrolyte solvents. In the first few cycles of use, a lithium-ion battery loses a large amount of capacity; this is generally attributed to lithium being consumed to produce SEI. However, after a few cycles, the rate of capacity loss slows at a rate often (but not always) reported to scale with the square root of time. SEI growth is therefore often considered to be limited in some way by a diffusion process.
# ### Dimensional Model
# We shall first state our model in dimensional form, but to enter the model in pybamm, we strongly recommend converting models into dimensionless form. The main reason for this is that dimensionless models are typically better conditioned than dimensional models and so several digits of accuracy can be gained. To distinguish between the dimensional and dimensionless models, we shall always employ a superscript $*$ on dimensional variables.
# 
# In our simple SEI model, we consider a one-dimensional SEI which extends from the surface of a planar negative electrode at $x^*=0$ until $x^*=L^*$, where $L^*$ is the thickness of the SEI. Since the SEI is porous, there is some electrolyte within the region $x^*\in[0, L^*]$ and therefore some concentration of solvent, $c^*$. Within the porous SEI, the solvent is transported via a diffusion process according to:
# $$
# \frac{\partial c^*}{\partial t^*} = - \nabla^* \cdot N^*, \quad N^* = - D^*(c^*) \nabla^* c^* \label{dim:eqn:solvent-diffusion}\tag{1}\\
# $$
# where $t^*$ is the time, $N^*$ is the solvent flux, and $D^*(c^*)$ is the effective solvent diffusivity (a function of the solvent concentration).
#
# On the electrode-SEI surface ($x^*=0$) the solvent is consumed by the SEI growth reaction, $R^*$. We assume that diffusion of solvent in the bulk electrolyte ($x^*>L^*$) is fast so that on the SEI-electrolyte surface ($x^*=L^*$) the concentration of solvent is fixed at the value $c^*_{\infty}$. Therefore, the boundary conditions are
# $$
# N^*|_{x^*=0} = - R^*, \quad c^*|_{x^*=L^*} = c^*_{\infty},
# $$
# We also assume that the concentration of solvent within the SEI is initially uniform and equal to the bulk electrolyte solvent concentration, so that the initial condition is
# $$
# c^*|_{t^*=0} = c^*_{\infty}
# $$
#
# Since the SEI is growing, we require an additional equation for the SEI thickness. The thickness of the SEI grows at a rate proportional to the SEI growth reaction $R^*$, where the constant of proportionality is the partial molar volume of the reaction products, $\hat{V}^*$. We also assume that the SEI is initially of thickness $L^*_0$. Therefore, we have
# $$
# \frac{d L^*}{d t^*} = \hat{V}^* R^*, \quad L^*|_{t^*=0} = L^*_0
# $$
#
# Finally, we assume for the sake of simplicity that the SEI growth reaction is irreversible and that the potential difference across the SEI is constant. The reaction is also assumed to be proportional to the concentration of solvent at the electrode-SEI surface ($x^*=0$). Therefore, the reaction flux is given by
# $$
# R^* = k^* c^*|_{x^*=0}
# $$
# where $k^*$ is the reaction rate constant (which is in general dependent upon the potential difference across the SEI).
# ### Non-dimensionalisation
# To convert the model into dimensionless form, we scale the dimensional variables and dimensional functions. For this model, we choose to scale $x^*$ by the current SEI thickness, the current SEI thickness by the initial SEI thickness, solvent concentration with the bulk electrolyte solvent concentration, and the solvent diffusion with the solvent diffusion in the electrolyte. We then use these scalings to infer the scaling for the solvent flux. Therefore, we have
# $$
# x^* = L^* x, \quad L^*= L^*_0 L \quad c^* = c^*_{\infty} c, \quad D^*(c^*) = D^*(c^*_{\infty}) D(c), \quad
# N^* = \frac{D^*(c^*_{\infty}) c^*_{\infty}}{L^*_0}N.
# $$
# We also choose to scale time by the solvent diffusion timescale so that
# $$
# t^* = \frac{(L^*_0)^2}{D^*(c^*_{\infty})}t.
# $$
# Finally, we choose to scale the reaction flux in the same way as the solvent flux so that we have
# $$
# R^* = \frac{D^*(c^*_{\infty}) c^*_{\infty}}{L^*_0} R.
# $$
#
# We note that there are multiple possible choices of scalings. Whilst they will all give the ultimately give the same answer, some choices are better than others depending on the situation under study.
# ### Dimensionless Model
# After substituting in the scalings from the previous section, we obtain the dimensionless form of the model given by:
# Solvent diffusion through SEI:
# \begin{align}
# \frac{\partial c}{\partial t} = \frac{\hat{V} R}{L} x \cdot \nabla c - \frac{1}{L}\nabla \cdot N, \quad N = - \frac{1}{L}D(c) \nabla c, \label{eqn:solvent-diffusion}\tag{1}\\
# N|_{x=0} = - R, \quad c|_{x=1} = 1 \label{bc:solvent-diffusion}\tag{2} \quad
# c|_{t=0} = 1;
# \end{align}
#
# Growth reaction:
# $$
# R = k c|_{x=0}; \label{eqn:reaction}\tag{3}
# $$
#
# SEI thickness:
# $$
# \frac{d L}{d t} = \hat{V} R, \quad L|_{t=0} = 1; \label{eqn:SEI-thickness}\tag{4}
# $$
# where the dimensionless parameters are given by
# $$
# k = \frac{k^* L^*_0}{D^*(c^*_{\infty})}, \quad \hat{V} = \hat{V}^* c^*_{\infty}, \quad
# D(c) = \frac{D^*(c^*)}{D^*(c^*_{\infty})}. \label{parameters}\tag{5}
# $$
# In the above, the additional advective term in the diffusion equation arises due to our choice to scale the spatial coordinate $x^*$ with the time-dependent SEI layer thickness $L^*$.
# ## Entering the Model into PyBaMM
# As always, we begin by importing pybamm and changing our working directory to the root of the pybamm folder.
import pybamm
import numpy as np
import os
os.chdir(pybamm.__path__[0]+'/..')
# A model is defined in six steps:
# 1. Initialise model
# 2. Define parameters and variables
# 3. State governing equations
# 4. State boundary conditions
# 5. State initial conditions
# 6. State output variables
#
# We shall proceed through each step to enter our simple SEI growth model.
# #### 1. Initialise model
# We first initialise the model using the `BaseModel` class. This sets up the required structure for our model.
model = pybamm.BaseModel()
# #### 2. Define parameters and variables
# In our SEI model, we have two dimensionless parameters, $k$ and $\hat{V}$, and one dimensionless function $D(c)$, which are all given in terms of the dimensional parameters, see (5). In pybamm, inputs are dimensional, so we first state all the dimensional parameters. We then define the dimensionless parameters, which are expressed an non-dimensional groupings of dimensional parameters. To define the dimensional parameters, we use the `Parameter` object to create parameter symbols. Parameters which are functions are defined using `FunctionParameter` object and should be defined within a python function as shown.
# +
# dimensional parameters
k_dim = pybamm.Parameter("Reaction rate constant")
L_0_dim = pybamm.Parameter("Initial thickness")
V_hat_dim = pybamm.Parameter("Partial molar volume")
c_inf_dim = pybamm.Parameter("Bulk electrolyte solvent concentration")
def D_dim(cc):
return pybamm.FunctionParameter("Diffusivity", cc)
# dimensionless parameters
k = k_dim * L_0_dim / D_dim(c_inf_dim)
V_hat = V_hat_dim * c_inf_dim
def D(cc):
c_dim = c_inf_dim * cc
return D_dim(c_dim) / D_dim(c_inf_dim)
# -
# We now define the dimensionless variables in our model. Since these are the variables we solve for directly, we do not need to write them in terms of the dimensional variables. We simply use `SpatialVariable` and `Variable` to create the required symbols:
x = pybamm.SpatialVariable("x", domain="SEI layer", coord_sys="cartesian")
c = pybamm.Variable("Solvent concentration", domain="SEI layer")
L = pybamm.Variable("SEI thickness")
# #### 3. State governing equations
# We can now use the symbols we have created for our parameters and variables to write out our governing equations. Note that before we use the reaction flux and solvent flux, we must derive new symbols for them from the defined parameter and variable symbols. Each governing equation must also be stated in the explicit form `d/dt = rhs` since pybamm only stores the right hand side (rhs) and assumes that the left hand side is the time derivative. The governing equations are then simply
# +
# SEI reaction flux
R = k * pybamm.BoundaryValue(c, "left")
# solvent concentration equation
N = - (1 / L) * D(c) * pybamm.grad(c)
dcdt = (V_hat * R) * pybamm.inner(x / L, pybamm.grad(c)) - (1 / L) * pybamm.div(N)
# SEI thickness equation
dLdt = V_hat * R
# -
# Once we have stated the equations, we can add them to the `model.rhs` dictionary. This is a dictionary whose keys are the variables being solved for, and whose values correspond right hand sides of the governing equations for each variable.
model.rhs = {c: dcdt, L: dLdt}
# #### 4. State boundary conditions
# We only have boundary conditions on the solvent concentration equation. We must state where a condition is Neumann (on the gradient) or Dirichlet (on the variable itself).
#
# The boundary condition on the electrode-SEI (x=0) boundary is:
# $$
# N|_{x=0} = - R, \quad N|_{x=0} = - \frac{1}{L} D(c|_{x=0} )\nabla c|_{x=0}
# $$
# which is a Neumann condition. To implement this boundary condition in pybamm, we must first rearrange the equation so that the gradient of the concentration, $\nabla c|_{x=0}$, is the subject. Therefore we have
# $$
# \nabla c|_{x=0} = \frac{L R}{D(c|_{x=0} )}
# $$
# which we enter into pybamm as
# electrode-SEI boundary condition (x=0) (lbc = left boundary condition)
D_left = pybamm.BoundaryValue(D(c), "left") # pybamm requires BoundaryValue(D(c)) and not D(BoundaryValue(c))
grad_c_left = L * R / D_left
# On the SEI-electrolyte boundary (x=1), we have the boundary condition
# $$
# c|_{x=1} = 1
# $$
# which is a Dirichlet condition and is just entered as
c_right = pybamm.Scalar(1)
# We now load these boundary conditions into the `model.boundary_conditions` dictionary in the following way, being careful to state the type of boundary condition:
model.boundary_conditions = {c: {"left": (grad_c_left, "Neumann"), "right": (c_right, "Dirichlet")}}
# #### 5. State initial conditions
# There are two initial conditions in our model:
# $$
# c|_{t=0} = 1, \quad L|_{t=0} = 1
# $$
# which are simply written in pybamm as
c_init = pybamm.Scalar(1)
L_init = pybamm.Scalar(1)
# and then included into the `model.initial_conditions` dictionary:
model.initial_conditions = {c: c_init, L: L_init}
# #### 6. State output variables
# We already have everything required in model for the model to be used and solved, but we have not yet stated what we actually want to output from the model. PyBaMM allows users to output any combination of symbols as an output variable therefore allowing the user the flexibility to output important quantities without further tedious postprocessing steps.
#
# Some useful outputs for this simple model are:
# - the SEI thickness
# - the SEI growth rate
# - the solvent concentration
#
# These are added to the model by adding entries to the `model.variables` dictionary
model.variables = {"SEI thickness": L, "SEI growth rate": dLdt, "Solvent concentration": c}
# We can also output the dimensional versions of these variables by multiplying by the scalings used to non-dimensionalise. By convention, we recommend including the units in the output variables name so that they do not overwrite the dimensionless output variables. To add new entries to the dictionary we used the method `.update()`.
# +
L_dim = L_0_dim * L
dLdt_dim = (D_dim(c_inf_dim) / L_0_dim ) * dLdt
c_dim = c_inf_dim * c
model.variables.update({
"SEI thickness [m]": L_dim,
"SEI growth rate [m/s]": dLdt_dim,
"Solvent concentration [mols/m^3]": c_dim
}
)
# -
# The model is now fully defined and ready to be used. If you plan on reusing the model several times, you can additionally set model defaults which may include: a default geometry to run the model on, a default set of parameter values, a default solver, etc.
# ## Using the Model
# The model will now behave in the same way as any of the inbuilt PyBaMM models. However, to demonstrate that the model works we display the steps involved in solving the model but we will not go into details within this notebook.
# +
# define geometry
geometry = pybamm.Geometry()
geometry.add_domain("SEI layer", {"primary": {x: {"min": pybamm.Scalar(0), "max": pybamm.Scalar(1)}}})
def Diffusivity(cc):
return cc * 10**(-5)
# parameter values (not physically based, for example only!)
param = pybamm.ParameterValues(
{
"Reaction rate constant": 20,
"Initial thickness": 1e-6,
"Partial molar volume": 10,
"Bulk electrolyte solvent concentration": 1,
"Diffusivity": Diffusivity,
}
)
# process model and geometry
param.process_model(model)
param.process_geometry(geometry)
# mesh and discretise
submesh_types = {"SEI layer": pybamm.Uniform1DSubMesh}
var_pts = {x: 100}
mesh = pybamm.Mesh(geometry, submesh_types, var_pts)
spatial_methods = {"SEI layer": pybamm.FiniteVolume()}
disc = pybamm.Discretisation(mesh, spatial_methods)
disc.process_model(model)
# solve
solver = pybamm.ScipySolver()
t = np.linspace(0, 100, 100)
solution = solver.solve(model, t)
# Extract output variables
L_out = solution["SEI thickness"]
c_out = solution["Solvent concentration"]
x = np.linspace(0, 1, 100)
# -
# Using these outputs, we can now plot the SEI thickness as a function of time and also the solvent concentration profile within the SEI. We use a slider to plot the concentration profile at different times.
# +
import matplotlib.pyplot as plt
def plot(t):
f, (ax1, ax2) = plt.subplots(1, 2 ,figsize=(10,5))
ax1.plot(solution.t, L_out(solution.t))
ax1.plot([t], [L_out(t)], 'r.')
plot_c, = ax2.plot(x * L_out(t), c_out(t, x))
ax1.set_ylabel('SEI thickness')
ax1.set_xlabel('t')
ax2.set_ylabel('Solvent concentration')
ax2.set_xlabel('x')
ax2.set_ylim(0, 1.1)
ax2.set_xlim(0, x[-1]*L_out(solution.t[-1]))
plt.show()
import ipywidgets as widgets
widgets.interact(plot, t=widgets.FloatSlider(min=0,max=solution.t[-1],step=0.1,value=0));
# -
# ## Formally adding your model
# The purpose of this notebook has been to go through the steps involved in getting a simple model working within PyBaMM. However, if you plan on reusing your model and want greater flexibility then we recommend that you create a new class for your model. We have set out instructions on how to do this in the "Adding a Model" tutorial in the documentation.
| examples/notebooks/Creating Models/5-a-simple-SEI-model.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] colab_type="text" id="e5O1UdsY202_"
# ##### Copyright 2019 Google LLC.
#
# Licensed under the Apache License, Version 2.0 (the "License");
# + colab={} colab_type="code" id="Zy3bZKW82xP9"
# Licensed under the Apache License, Version 2.0 (the "License");
# you may not use this file except in compliance with the License.
# You may obtain a copy of the License at
#
# https://www.apache.org/licenses/LICENSE-2.0
#
# Unless required by applicable law or agreed to in writing, software
# distributed under the License is distributed on an "AS IS" BASIS,
# WITHOUT WARRANTIES OR CONDITIONS OF ANY KIND, either express or implied.
# See the License for the specific language governing permissions and
# limitations under the License.
# + [markdown] colab_type="text" id="GXhLzrXN27af"
# # Using the Meta-Dataset Data Pipeline
#
# This notebook shows how to use `meta_dataset`’s input pipeline to sample data for the Meta-Dataset benchmark. There are two main ways in which data is sampled:
# 1. **episodic**: Returns N-way classification *episodes*, which contain a *support* (training) set and a *query* (test) set. The number of classes (N) may vary from episode to episode.
# 2. **batch**: Returns batches of images and their corresponding label, sampled from all available classes.
#
# We first import `meta_dataset` and other required packages, and define utility functions for visualization. We’ll make use of `meta_dataset.data.learning_spec` and `meta_dataset.data.pipeline`; their purpose will be made clear later on.
# + cellView="both" colab={} colab_type="code" id="ZyMqBhZIxPQD"
#@title Imports and Utility Functions
from __future__ import absolute_import
from __future__ import division
from __future__ import print_function
import os
from collections import Counter
import gin
import numpy as np
import matplotlib.pyplot as plt
import tensorflow as tf
from meta_dataset.data import config
from meta_dataset.data import dataset_spec as dataset_spec_lib
from meta_dataset.data import learning_spec
from meta_dataset.data import pipeline
def plot_episode(support_images, support_class_ids, query_images,
query_class_ids, size_multiplier=1, max_imgs_per_col=10,
max_imgs_per_row=10):
for name, images, class_ids in zip(('Support', 'Query'),
(support_images, query_images),
(support_class_ids, query_class_ids)):
n_samples_per_class = Counter(class_ids)
n_samples_per_class = {k: min(v, max_imgs_per_col)
for k, v in n_samples_per_class.items()}
id_plot_index_map = {k: i for i, k
in enumerate(n_samples_per_class.keys())}
num_classes = min(max_imgs_per_row, len(n_samples_per_class.keys()))
max_n_sample = max(n_samples_per_class.values())
figwidth = max_n_sample
figheight = num_classes
if name == 'Support':
print('#Classes: %d' % len(n_samples_per_class.keys()))
figsize = (figheight * size_multiplier, figwidth * size_multiplier)
fig, axarr = plt.subplots(
figwidth, figheight, figsize=figsize)
fig.suptitle('%s Set' % name, size='20')
fig.tight_layout(pad=3, w_pad=0.1, h_pad=0.1)
reverse_id_map = {v: k for k, v in id_plot_index_map.items()}
for i, ax in enumerate(axarr.flat):
ax.patch.set_alpha(0)
# Print the class ids, this is needed since, we want to set the x axis
# even there is no picture.
ax.set(xlabel=reverse_id_map[i % figheight], xticks=[], yticks=[])
ax.label_outer()
for image, class_id in zip(images, class_ids):
# First decrement by one to find last spot for the class id.
n_samples_per_class[class_id] -= 1
# If class column is filled or not represented: pass.
if (n_samples_per_class[class_id] < 0 or
id_plot_index_map[class_id] >= max_imgs_per_row):
continue
# If width or height is 1, then axarr is a vector.
if axarr.ndim == 1:
ax = axarr[n_samples_per_class[class_id]
if figheight == 1 else id_plot_index_map[class_id]]
else:
ax = axarr[n_samples_per_class[class_id], id_plot_index_map[class_id]]
ax.imshow(image / 2 + 0.5)
plt.show()
def plot_batch(images, labels, size_multiplier=1):
num_examples = len(labels)
figwidth = np.ceil(np.sqrt(num_examples)).astype('int32')
figheight = num_examples // figwidth
figsize = (figwidth * size_multiplier, (figheight + 1.5) * size_multiplier)
_, axarr = plt.subplots(figwidth, figheight, dpi=300, figsize=figsize)
for i, ax in enumerate(axarr.transpose().ravel()):
# Images are between -1 and 1.
ax.imshow(images[i] / 2 + 0.5)
ax.set(xlabel=labels[i], xticks=[], yticks=[])
plt.show()
# + [markdown] colab_type="text" id="BOn_YZdqPIv5"
# # Primers
# 1. Download your data and process it as explained in [link](https://github.com/google-research/meta-dataset/blob/main/README.md#downloading-and-converting-datasets). Set `BASE_PATH` pointing the processed tf-records (`$RECORDS` in the conversion instructions).
# 2. `meta_dataset` supports many different setting for sampling data. We use [gin-config](https://github.com/google/gin-config) to control default parameters of our functions. You can go to default gin file we are pointing and see the default values.
# 3. You can use `meta_dataset` in **eager** or **graph** mode.
# 4. Let's write a generator that makes the right calls to return data from dataset. `dataset.make_one_shot_iterator()` returns an iterator where each element is an episode.
# 4. SPLIT is used to define which part of the meta-split is going to be used. Different splits have different classes and the details on how they are created can be found in the [paper](https://arxiv.org/abs/1903.03096).
# + colab={} colab_type="code" id="_di9Tczj8joM"
# 1
BASE_PATH = '/path/to/records'
GIN_FILE_PATH = 'meta_dataset/learn/gin/setups/data_config.gin'
# 2
gin.parse_config_file(GIN_FILE_PATH)
# 3
# Comment out to disable eager execution.
tf.enable_eager_execution()
# 4
def iterate_dataset(dataset, n):
if not tf.executing_eagerly():
iterator = dataset.make_one_shot_iterator()
next_element = iterator.get_next()
with tf.Session() as sess:
for idx in range(n):
yield idx, sess.run(next_element)
else:
for idx, episode in enumerate(dataset):
if idx == n:
break
yield idx, episode
# 5
SPLIT = learning_spec.Split.TRAIN
# + [markdown] colab_type="text" id="Pn6ndPMhxs8W"
# # Reading datasets
# In order to sample data, we need to read the dataset_spec files for each dataset. Following snippet reads those files into a list.
# + colab={} colab_type="code" id="Z0uU6WrbxsMa"
ALL_DATASETS = ['aircraft', 'cu_birds', 'dtd', 'fungi', 'ilsvrc_2012',
'omniglot', 'quickdraw', 'vgg_flower']
all_dataset_specs = []
for dataset_name in ALL_DATASETS:
dataset_records_path = os.path.join(BASE_PATH, dataset_name)
dataset_spec = dataset_spec_lib.load_dataset_spec(dataset_records_path)
all_dataset_specs.append(dataset_spec)
# + [markdown] colab_type="text" id="7p448EXYxwbb"
# # (1) Episodic Mode
# `meta_dataset` uses [tf.data.Dataset](https://www.tensorflow.org/api_docs/python/tf/data/Dataset) API and it takes one call to `pipeline.make_multisource_episode_pipeline()`. We loaded or defined most of the variables used during this call above. The remaining parameters are explained below:
#
# - **use_bilevel_ontology_list**: This is a list of booleans indicating whether corresponding dataset in `ALL_DATASETS` should use bilevel ontology. Omniglot is set up with a hierarchy with two level: the alphabet (Latin, Inuktitut...), and the character (with 20 examples per character).
# The flag means that each episode will contain classes from a single alphabet.
# - **use_dag_ontology_list**: This is a list of booleans indicating whether corresponding dataset in `ALL_DATASETS` should use dag_ontology. Same idea for ImageNet, except it uses the hierarchical sampling procedure described in the article.
# - **image_size**: All images from various datasets are down or upsampled to the same size. This is the flag controls the edge size of the square.
# - **shuffle_buffer_size**: Controls the amount of shuffling among examples from any given class.
# + colab={} colab_type="code" id="jPlnBWwkwuGP"
use_bilevel_ontology_list = [False]*len(ALL_DATASETS)
use_dag_ontology_list = [False]*len(ALL_DATASETS)
# Enable ontology aware sampling for Omniglot and ImageNet.
use_bilevel_ontology_list[5] = True
use_dag_ontology_list[4] = True
variable_ways_shots = config.EpisodeDescriptionConfig(
num_query=None, num_support=None, num_ways=None)
dataset_episodic = pipeline.make_multisource_episode_pipeline(
dataset_spec_list=all_dataset_specs,
use_dag_ontology_list=use_dag_ontology_list,
use_bilevel_ontology_list=use_bilevel_ontology_list,
episode_descr_config=variable_ways_shots,
split=SPLIT,
image_size=84,
shuffle_buffer_size=300)
# + [markdown] colab_type="text" id="BN66UXO79Bo2"
# ## Using Dataset
# 1. The episodic dataset consist in a tuple of the form (Episode, data source ID). The data source ID is an integer Tensor containing a value in the range [0, len(all_dataset_specs) - 1]
# signifying which of the datasets of the multisource pipeline the given episode
# came from. Episodes consist of support and query sets and we want to learn to classify images at the query set correctly given the support images. For both support and query set we have `images`, `labels` and `class_ids`. Labels are transformed class_ids offset to zero, so that global class_ids are set to \[0, N\] where N is the number of classes in an episode.
# 3. As one can see the number of images in query set and support set is different. Images are scaled, copied into 84\*84\*3 tensors. Labels are presented in two forms:
# * `*_labels` are relative to the classes selected for the current episode only. They are used as targets for this episode.
# * `*_class_ids` are the original class ids relative to the whole dataset. They are used for visualization and diagnostics.
# 4. It easy to convert tensors of the episode into numpy arrays and use them outside of the Tensorflow framework.
# 5. Classes might have different number of samples in the support set, whereas each class has 10 samples in the query set.
#
# + colab={} colab_type="code" id="lomtjv9rw5WP"
# 1
idx, (episode, source_id) = next(iterate_dataset(dataset_episodic, 1))
print('Got an episode from dataset:', all_dataset_specs[source_id].name)
# 2
for t, name in zip(episode,
['support_images', 'support_labels', 'support_class_ids',
'query_images', 'query_labels', 'query_class_ids']):
print(name, t.shape)
# 3
episode = [a.numpy() for a in episode]
# 4
support_class_ids, query_class_ids = episode[2], episode[5]
print(Counter(support_class_ids))
print(Counter(query_class_ids))
# + [markdown] colab_type="text" id="KxdVUqJiWmTX"
# ## Visualizing Episodes
# Let's visualize the episodes.
#
# - Support and query set for each episode plotted sequentially. Set N_EPISODES to control number of episodes visualized.
# - Each episode is sampled from a single dataset and include N different classes. Each class might have different number of samples in support set, whereas number of images in query set is fixed. We limit number of classes and images per class to 10 in order to create legible plots. Actual episodes might have more classes and samples.
# - Each column represents a distinct class and dataset specific class ids are plotted on the x_axis.
# + colab={} colab_type="code" id="9v2ePLTkoZlE"
# 1
N_EPISODES=2
# 2, 3
for idx, (episode, source_id) in iterate_dataset(dataset_episodic, N_EPISODES):
print('Episode id: %d from source %s' % (idx, all_dataset_specs[source_id].name))
episode = [a.numpy() for a in episode]
plot_episode(support_images=episode[0], support_class_ids=episode[2],
query_images=episode[3], query_class_ids=episode[5])
# + [markdown] colab_type="text" id="pL2AZ5gx3cDS"
# # (2) Batch Mode
# Second mode that `meta_dataset` library provides is the batch mode, where one can sample batches from the list of datasets in a non-episodic manner and use it to train baseline models. There are couple things to note here:
#
# - Each batch is sampled from a different dataset.
# - `ADD_DATASET_OFFSET` controls whether the class_id's returned by the iterator overlaps among different datasets or not. A dataset specific offset is added in order to make returned ids unique.
# - `make_multisource_batch_pipeline()` creates a `tf.data.Dataset` object that returns datasets of the form (Batch, data source ID) where similarly to the
# episodic case, the data source ID is an integer Tensor that identifies which
# dataset the given batch originates from.
# - `shuffle_buffer_size` controls the amount of shuffling done among examples from a given dataset (unlike for the episodic pipeline).
# + colab={} colab_type="code" id="jYY5zd_S6uG6"
BATCH_SIZE = 16
ADD_DATASET_OFFSET = True
# + colab={} colab_type="code" id="BgkTLKKXPh8M"
dataset_batch = pipeline.make_multisource_batch_pipeline(
dataset_spec_list=all_dataset_specs, batch_size=BATCH_SIZE, split=SPLIT,
image_size=84, add_dataset_offset=ADD_DATASET_OFFSET,
shuffle_buffer_size=1000)
for idx, ((images, labels), source_id) in iterate_dataset(dataset_batch, 1):
print(images.shape, labels.shape)
# + colab={} colab_type="code" id="7hGjt6GGonAz"
N_BATCH = 2
for idx, (batch, source_id) in iterate_dataset(dataset_batch, N_BATCH):
print('Batch-%d from source %s' % (idx, all_dataset_specs[source_id].name))
plot_batch(*map(lambda a: a.numpy(), batch), size_multiplier=0.5)
# + [markdown] colab_type="text" id="tu4-jz89xt1f"
# # (3) Fixing Ways and Shots
# 1. `meta_dataset` library provides option to set number of classes/samples per episode. There are 3 main flags you can set.
# - **NUM_WAYS**: Fixes the # classes per episode. We would still get variable number of samples per class in the support set.
# - **NUM_SUPPORT**: Fixes # samples per class in the support set.
# - **NUM_SUPPORT**: Fixes # samples per class in the query set.
# 2. If we want to use fixed `num_ways`, we have to disable ontology based sampling for omniglot and imagenet. We advise using single dataset for using this feature, since using multiple datasets is not supported/tested. In this notebook, we are using Quick, Draw! Dataset.
# 3. We sample episodes and visualize them as we did earlier.
# + colab={} colab_type="code" id="8raM-sad6Igu"
#1
NUM_WAYS = 8
NUM_SUPPORT = 3
NUM_QUERY = 5
fixed_ways_shots = config.EpisodeDescriptionConfig(
num_ways=NUM_WAYS, num_support=NUM_SUPPORT, num_query=NUM_QUERY)
#2
use_bilevel_ontology_list = [False]*len(ALL_DATASETS)
use_dag_ontology_list = [False]*len(ALL_DATASETS)
quickdraw_spec = [all_dataset_specs[6]]
#3
dataset_fixed = pipeline.make_multisource_episode_pipeline(
dataset_spec_list=quickdraw_spec, use_dag_ontology_list=[False],
use_bilevel_ontology_list=use_bilevel_ontology_list, split=SPLIT,
image_size=84, episode_descr_config=fixed_ways_shots)
N_EPISODES = 2
for idx, (episode, source_id) in iterate_dataset(dataset_fixed, N_EPISODES):
print('Episode id: %d from source %s' % (idx, quickdraw_spec[source_id].name))
episode = [a.numpy() for a in episode]
plot_episode(support_images=episode[0], support_class_ids=episode[2],
query_images=episode[3], query_class_ids=episode[5])
# + [markdown] colab_type="text" id="4tKDA6JFxt11"
# # (4) Using Meta-dataset with PyTorch
# As mentioned above it is super easy to consume `meta_dataset` as NumPy arrays. This also enables easy integration into other popular deep learning frameworks like PyTorch. TensorFlow code processes the data and passes it to PyTorch, ready to be consumed. Since the data loader and processing steps do not have any operation on the GPU, TF should not attempt to grab the GPU, and it should be available for PyTorch.
# 1. Let's use an episodic dataset created earlier, `dataset_episodic`, and build on top of it. We will transpose tensor to CHW, which is the common order used by [convolutional layers](https://pytorch.org/docs/stable/nn.html?highlight=conv2d#torch.nn.functional.conv2d) of PyTorch.
# 2. We will use zero-indexed labels, therefore grabbing `e[1]` and `e[4]`. At the end we return a generator that consumes the `tf.Dataset`.
# 3. Using `.cuda()` on PyTorch tensors should distribute them to appropriate devices.
#
# + colab={} colab_type="code" id="2d5w2YW-xt14"
import torch
# 1
to_torch_labels = lambda a: torch.from_numpy(a.numpy()).long()
to_torch_imgs = lambda a: torch.from_numpy(np.transpose(a.numpy(), (0, 3, 1, 2)))
# 2
def data_loader(n_batches):
for i, (e, _) in enumerate(dataset_episodic):
if i == n_batches:
break
yield (to_torch_imgs(e[0]), to_torch_labels(e[1]),
to_torch_imgs(e[3]), to_torch_labels(e[4]))
for i, batch in enumerate(data_loader(n_batches=2)):
#3
data_support, labels_support, data_query, labels_query = [x.cuda() for x in batch]
print(data_support.shape, labels_support.shape, data_query.shape, labels_query.shape)
| .ipynb_checkpoints/Intro_to_Metadataset-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# orphan: true
# ---
# # Running Your Models
#
# - [Setting and Getting Component Variables](set_get.ipynb)
# - [Setup Your Model](setup.ipynb)
# - [Run Your Model](run_model.ipynb)
# - [Run a Driver](run_driver.ipynb)
| openmdao/docs/openmdao_book/features/core_features/running_your_models/main.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Sentiment analysis
# This script uses the cleaned tweets and tags them with a sentiment polarity score (ranging from -1 (negative) to 1 (positive)). After comparing different methods for sentiment analysis, it was found that the [Google Natural Language sentiment analyzer](https://cloud.google.com/natural-language/docs) provided the most accurate result (based on an inspection of the results). Therefore, only this method will be documented here, while the other methods will be covered in Rienje's portfolio.
# +
# Import needed libraries
import pandas as pd
import numpy as np
from deep_translator import GoogleTranslator
from google.cloud import language_v1
# Load cleaned tweets from previous script
df = pd.read_csv('cleaned_sentiment_tweets.csv')
# -
# #### Translating tweets
# Unfortunately, Dutch sentiment analysis is currently not possible due to a [bug](https://issuetracker.google.com/issues/180714982) in the Google Natural Language tool. Therefore, we need to translate the tweets to English prior to analysis. This reduces the accuracy of the sentiment analysis, although the Google API still yields better scores than Dutch sentiment analyses (like [pattern.nl](https://github.com/clips/pattern/wiki/pattern-nl) or analyses based on [classified Dutch tweets](https://github.com/cltl-students/Eva_Zegelaar_Emotion_Classification_Dutch_Political_Tweets)). We translate the tweet using the [deep-translator](https://pypi.org/project/deep-translator/) wrapper for the Google translate API, which actually allows unlimited translating for free.
# The code below takes several hours to run, so we suggest the reader uses the provided data in the next notebook, instead of running it.
# +
# Initiate a list
translatedList = []
# This loop takes a long time (several hours!) to run for all 18k tweets
for tweet in split['text_for_translation']:
translation = GoogleTranslator(source='nl', dest='en').translate(tweet)
translatedList.append(translation)
# Stitch translation to df
df['translation'] = translatedList
# -
# #### The sentiment analysis
# Provided below is the code that sends the tweets to the Google Cloud and returns a sentiment score. This code will probably yield an error if the user does not have a Google Cloud service account. Also, the costs of the sentiment analysis are roughly 1 dollar per 1000 tweets, so running the block below is not recommended. Instead, the tweets with the tagged sentiments will be loaded in the next notebook. The code is an adaptation from [Stackoverflow](https://stackoverflow.com/questions/61319178/how-can-i-send-a-batch-of-strings-to-the-google-cloud-natural-language-api).
#
# +
# Google natural language sentiment analysis
# Costs are roughly 1 dollar per 1000 tweets
# Running the google cloud takes a long time, ~4 hours for 18 000 tweets
# Check instantiation of client
client = language_v1.LanguageServiceClient.from_service_account_json("D:/Users/Rienje/Documents/MGI Wageningen/SmartEnvironmentDataScience/Project/Python/googleNL/sentiment-309511-2a1624b4263f.json")
# Create a function that retrieves sentiment score from Google NL API
def comment_analysis(comment):
# Re-instantiate client
client = language_v1.LanguageServiceClient.from_service_account_json("D:/Users/Rienje/Documents/MGI Wageningen/SmartEnvironmentDataScience/Project/Python/googleNL/sentiment-309511-2a1624b4263f.json")
# Set parameters for analysis
document = {"content":comment,
"type_":language_v1.Document.Type.PLAIN_TEXT,
"language":"en"}
# Sentiment analysis
annotations = client.analyze_sentiment(document=document)
# Append only the sentiment score of the tweet
total_score = annotations.document_sentiment.score
return total_score
# Initiate list
GoogleCloudList = []
# Retrieve sentiment score for al tweets
for tweet in df['translation']:
googlesentiment = comment_analysis(tweet)
GoogleCloudList.append(googlesentiment)
# Add to df and save as csv
df['google_scores'] = GoogleCloudList
df.to_csv('final_sentiment_tweets.csv', header=True, index = False)
# -
# #### Reflection
# There are many modules for sentiment analysis available for Python, although finding an *accurate* sentiment analyzer for Dutch is much more difficult. During the project, an estimation was made of which of four (machine learning, Google Cloud, Pattern and NLTK Vader) was the most 'accurate' for our project. In the end, as described above, the choice was made for the Google Cloud analyzer. This API also has some downsides however.
#
# First, like many of Google's services, it is very black box-y. The user has little insight on how certain tweets are tagged, in contrast with academic classifiers (like [Pattern](https://github.com/clips/pattern) and, to a lesser extent, [NLTK](https://www.nltk.org/api/nltk.sentiment.html). In addition, the texts had to be translated, which inevitably leads to a loss of context, sentence structure and thus, sentiment. The database was too large (or time too limited) to check the translations.
# Therefore, the results of the sentiment analysis should be taken with a grain of salt. In addition to reasons described above, tweet sentiment analysis is quite tricky in itself, as the sentences are short and often informal or sarcastic (especially when concerning complex political tweets).
| 3_Sentiment_Analysis.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# %matplotlib inline
# %config InlineBackend.figure_format = 'retina'
from matplotlib import pyplot as plt
import pandas as pd
# -
churn_data = pd.read_csv('https://raw.githubusercontent.com/'
'treselle-systems/customer_churn_analysis/'
'master/WA_Fn-UseC_-Telco-Customer-Churn.csv')
churn_data.head()
# +
churn_data = churn_data.set_index('customerID')
churn_data = churn_data.drop(['TotalCharges'], axis=1)
# The dataset is naturally hierarchical: some columns only apply to some users. Ex, if you don't have internet
# then the column OnlineBackup isn't applicable, as it's value is "No internet service". We
# are going to map this back to No. We will treat the hierachical nature by stratifying on the
# different services a user may have.
churn_data = churn_data.applymap(lambda x: "No" if str(x).startswith("No ") else x)
churn_data['Churn'] = (churn_data['Churn'] == "Yes")
strata_cols = ['InternetService', 'StreamingMovies', 'StreamingTV', 'PhoneService']
# +
from lifelines import CoxPHFitter
cph = CoxPHFitter().fit(churn_data, 'tenure', 'Churn', strata=strata_cols)
# -
cph
cph.print_summary()
ax = plt.subplots(figsize=(8, 6))
cph.plot(ax=ax[1])
cph.plot_covariate_groups('Contract', values=["Month-to-month", "One year", "Two year"], plot_baseline=False);
| examples/Customer Churn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from functools import partial
from rpy2.ipython import html
html.html_rdataframe=partial(html.html_rdataframe, table_class="docutils")
# # `R` and `pandas` data frames
#
# R `data.frame` and :class:`pandas.DataFrame` objects share a lot of
# conceptual similarities, and :mod:`pandas` chose to use the class name
# `DataFrame` after R objects.
#
# In a nutshell, both are sequences of vectors (or arrays) of consistent
# length or size for the first dimension (the "number of rows").
# if coming from the database world, an other way to look at them is
# column-oriented data tables, or data table API.
#
# rpy2 is providing an interface between Python and R, and a convenience
# conversion layer between :class:`rpy2.robjects.vectors.DataFrame` and
# :class:`pandas.DataFrame` objects, implemented in
# :mod:`rpy2.robjects.pandas2ri`.
# +
import pandas as pd
import rpy2.robjects as ro
from rpy2.robjects.packages import importr
from rpy2.robjects import pandas2ri
from rpy2.robjects.conversion import localconverter
# -
# ## From `pandas` to `R`
#
# Pandas data frame:
# +
pd_df = pd.DataFrame({'int_values': [1,2,3],
'str_values': ['abc', 'def', 'ghi']})
pd_df
# -
# R data frame converted from a `pandas` data frame:
# +
with localconverter(ro.default_converter + pandas2ri.converter):
r_from_pd_df = ro.conversion.py2rpy(pd_df)
r_from_pd_df
# -
# The conversion is automatically happening when calling R functions.
# For example, when calling the R function `base::summary`:
# +
base = importr('base')
with localconverter(ro.default_converter + pandas2ri.converter):
df_summary = base.summary(pd_df)
print(df_summary)
# -
# Note that a `ContextManager` is used to limit the scope of the
# conversion. Without it, rpy2 will not know how to convert a pandas
# data frame:
try:
df_summary = base.summary(pd_df)
except NotImplementedError as nie:
print('NotImplementedError:')
print(nie)
# ## From `R` to `pandas`
#
# Starting from an R data frame this time:
# +
r_df = ro.DataFrame({'int_values': ro.IntVector([1,2,3]),
'str_values': ro.StrVector(['abc', 'def', 'ghi'])})
r_df
# -
# It can be converted to a pandas data frame using the same converter:
# +
with localconverter(ro.default_converter + pandas2ri.converter):
pd_from_r_df = ro.conversion.rpy2py(r_df)
pd_from_r_df
# -
# ## Date and time objects
# +
pd_df = pd.DataFrame({
'Timestamp': pd.date_range('2017-01-01 00:00:00', periods=10, freq='s')
})
pd_df
# +
with localconverter(ro.default_converter + pandas2ri.converter):
r_from_pd_df = ro.conversion.py2rpy(pd_df)
r_from_pd_df
# -
# The timezone used for conversion is the system's default timezone unless `pandas2ri.default_timezone`
# is specified... or unless the time zone is specified in the original time object:
# +
pd_tz_df = pd.DataFrame({
'Timestamp': pd.date_range('2017-01-01 00:00:00', periods=10, freq='s',
tz='UTC')
})
with localconverter(ro.default_converter + pandas2ri.converter):
r_from_pd_tz_df = ro.conversion.py2rpy(pd_tz_df)
r_from_pd_tz_df
| Samples/pandas.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### 1. Problem statement
# - We are tasked by a Fintech firm to analyze mobile app behavior data to identify potential churn customers.
# - The goal is to predict which users are likely to churn, so the firm can focus on re-engaging these users with better products.
# - Below is focusing on modelling.
# ### 2. Importing libraries
import warnings
warnings.filterwarnings("ignore")
import pandas as pd
import numpy as np
import random
import seaborn as sn
import matplotlib.pyplot as plt
from sklearn.model_selection import train_test_split
from sklearn.preprocessing import StandardScaler
# Recursive Feature Elimination
from sklearn.feature_selection import RFE
from sklearn.linear_model import LogisticRegression
from sklearn.model_selection import cross_val_score
from sklearn.metrics import confusion_matrix, accuracy_score, f1_score, precision_score, recall_score
# ### 3. Data process
dataset = pd.read_csv('new_churn_data.csv')
## Data Preparation
user_identifier = dataset['user']
dataset = dataset.drop(columns = ['user'])
# #### 3.1 One-hot encoding
dataset.housing.value_counts()
dataset.groupby('housing')['churn'].nunique().reset_index()
dataset = pd.get_dummies(dataset)
dataset.columns
# #### 3.2 Drop correlated columns (data trap)
dataset = dataset.drop(columns = ['housing_na', 'zodiac_sign_na', 'payment_type_na'])
# #### 3.3 Train test split
X_train, X_test, y_train, y_test = train_test_split(dataset.drop(columns = 'churn'),
dataset['churn'],
test_size = 0.2,
random_state = 0)
# #### 3.4 Data balancing
y_train.value_counts()
# +
pos_index = y_train[y_train.values == 1].index
neg_index = y_train[y_train.values == 0].index
if len(pos_index) > len(neg_index):
higher = pos_index
lower = neg_index
else:
higher = neg_index
lower = pos_index
random.seed(0)
higher = np.random.choice(higher, size=len(lower))
lower = np.asarray(lower)
new_indexes = np.concatenate((lower, higher))
X_train = X_train.loc[new_indexes,]
y_train = y_train[new_indexes]
# -
# #### 3.5 Feature scaling
sc_X = StandardScaler()
X_train2 = pd.DataFrame(sc_X.fit_transform(X_train))
X_test2 = pd.DataFrame(sc_X.transform(X_test))
X_train2.columns = X_train.columns.values
X_test2.columns = X_test.columns.values
X_train2.index = X_train.index.values
X_test2.index = X_test.index.values
X_train = X_train2
X_test = X_test2
X_train.head()
# ### 4. Model create
# #### 4.1 Create model
classifier = LogisticRegression(random_state = 0)
classifier.fit(X_train, y_train)
# Predicting Test Set
y_pred = classifier.predict(X_test)
# Evaluating Results
cm = confusion_matrix(y_test, y_pred)
accuracy_score(y_test, y_pred)
precision_score(y_test, y_pred) # tp / (tp + fp)
recall_score(y_test, y_pred) # tp / (tp + fn)
f1_score(y_test, y_pred)
df_cm = pd.DataFrame(cm, index = (0, 1), columns = (0, 1))
plt.figure(figsize = (6,4))
sn.set(font_scale=1.4)
sn.heatmap(df_cm, annot=True, fmt='g')
print("Test Data Accuracy: %0.4f" % accuracy_score(y_test, y_pred))
# #### 4.2 Cross Validation
accuracies = cross_val_score(estimator = classifier, X = X_train, y = y_train, cv = 10)
print("Accuracy: %0.3f (+/- %0.3f)" % (accuracies.mean(), accuracies.std() * 2))
accuracies.mean()
len(X_train.columns.tolist())
# #### 4.3 Feature importance analysis
# Analyzing Coefficients
pd.concat([pd.DataFrame(X_train.columns, columns = ["features"]),
pd.DataFrame(np.transpose(classifier.coef_), columns = ["coef"])
],axis = 1)
# #### 4.4 Feature Selection
# Model to Test
classifier = LogisticRegression()
# Select Best X Features
rfe = RFE(classifier, 20)
rfe = rfe.fit(X_train, y_train)
# #### summarize the selection of the attributes
print(rfe.support_)
print(rfe.ranking_)
X_train.columns[rfe.support_]
# #### 4.5 Retrain model
classifier = LogisticRegression()
classifier.fit(X_train[X_train.columns[rfe.support_]], y_train)
# Predicting Test Set
y_pred = classifier.predict(X_test[X_train.columns[rfe.support_]])
# Evaluating Results
cm = confusion_matrix(y_test, y_pred)
accuracy_score(y_test, y_pred)
f1_score(y_test, y_pred)
df_cm = pd.DataFrame(cm, index = (1, 0), columns = (1, 0))
plt.figure(figsize = (6,4))
sn.set(font_scale=1.4)
sn.heatmap(df_cm, annot=True, fmt='g')
print("Test Data Accuracy: %0.4f" % accuracy_score(y_test, y_pred))
# #### 4.6 Cross validation (2nd)
# +
# Applying k-Fold Cross Validation
# Fitting Model to the Training Set
accuracies = cross_val_score(estimator = classifier,
X = X_train[X_train.columns[rfe.support_]],
y = y_train, cv = 10)
# -
print("Accuracy: %0.3f (+/- %0.3f)" % (accuracies.mean(), accuracies.std() * 2))
# #### 4.7 Feature importance analysis (2nd)
pd.concat([pd.DataFrame(X_train[X_train.columns[rfe.support_]].columns, columns = ["features"]),
pd.DataFrame(np.transpose(classifier.coef_), columns = ["coef"])
], axis = 1)
y_test
user_identifier
# #### 4.8 Formatting Final Results
final_results = pd.concat([y_test, user_identifier], axis = 1).dropna()
final_results['predicted_churn'] = y_pred
final_results = final_results[['user', 'churn', 'predicted_churn']].reset_index(drop=True)
final_results
| churn_prediction/.ipynb_checkpoints/App_churn_prediction_Logistic_regression_rfe-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Specific Models
# - Feature engineering (PDSH Chapter 5.4)
# - Naive Bayes (PDSH Chapter 5.5)
# - Linear regression (5.6)
# - SVM (5.7)
# - Random forests (5.8)
# - PCA (5.9)
# - Manifold learning (5.10)
# - K-means (5.11)
# - Gaussian mixtures (5.12)
# - Kernel density estimation (5.13)
# - Neural networks (DL, Keras)
| Content/Model/10-SpecificModels.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: TensorFlow 2.3 on Python 3.6 (CUDA 10.1)
# language: python
# name: python3
# ---
# # 케라스 API를 사용한 사용자 정의 모델 만들기 with 텐서플로 2.3+2.4
#
# DLD(Daejeon Learning Day) 2020을 위해 작성된 노트북입니다.
#
# * 깃허브 주소: https://github.com/rickiepark/handson-ml2/blob/master/custom_model_in_keras.ipynb
# * 코랩 주소: https://colab.research.google.com/github/rickiepark/handson-ml2/blob/master/custom_model_in_keras.ipynb
# +
import tensorflow as tf
tf.__version__
# -
# ### MNIST 손글씨 숫자 데이터 적재
# +
(X_train, y_train), (X_test, y_test) = tf.keras.datasets.mnist.load_data()
X_train = X_train.reshape(-1, 784) / 255.
# -
X_train.shape
# ### `Sequential()` 클래스와 함수형 API의 관계
# `Sequential()`:
# 시퀀셜 모델에 10개의 유닛을 가진 완전 연결 층을 추가합니다.
# +
seq_model = tf.keras.Sequential()
seq_model.add(tf.keras.layers.Dense(units=10,
activation='softmax',
input_shape=(784,)))
seq_model.summary()
# -
seq_model.compile(loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
seq_model.fit(X_train, y_train, batch_size=32, epochs=2)
# ### 함수형 API:
#
# 함수형 API를 사용할 때는 `Input()`을 사용해 입력의 크기를 정의해야 합니다. 하지만 `InputLayer` 층이 추가되어 있습니다.
# +
inputs = tf.keras.layers.Input(784)
outputs = tf.keras.layers.Dense(units=10,
activation='softmax')(inputs) # __call()__ 메서드 호출
# dense = tf.keras.layers.Dense(units=10, activation='softmax')
# outputs = dense(inputs)
func_model = tf.keras.Model(inputs, outputs)
func_model.summary()
# -
func_model.compile(loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
func_model.fit(X_train, y_train, batch_size=32, epochs=2)
# `Input`의 정체는 무엇일까요? 이 함수는 `InputLayer` 클래스의 객체를 만들어 그 결과를 반환합니다.
type(tf.keras.layers.Input)
# 사실 신경망의 입력층은 입력 그 자체입니다. `InputLayer` 객체의 입력 노드 출력을 그대로 `Dense` 층에 주입할 수 있습니다. 모든 층은 입력과 출력 노드를 정의합니다.
# +
# inputs = tf.keras.layers.Input(784)
input_layer = tf.keras.layers.InputLayer(784)
inputs = input_layer._inbound_nodes[0].outputs
outputs = tf.keras.layers.Dense(units=10,
activation='softmax')(inputs)
input_layer_model = tf.keras.Model(inputs, outputs)
input_layer_model.summary()
# -
input_layer_model.compile(loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
input_layer_model.fit(X_train, y_train, batch_size=32, epochs=2)
# 함수형 API를 사용한 모델은 `layers` 속성에 `InputLayer` 클래스를 포함합니다.
func_model.layers
# 하지만 시퀀셜 모델은 `layers` 속성에 `InputLayer` 클래스가 보이지 않습니다.
seq_model.layers
# 모델은 감춰진 `_layers` 속성이 또 있습니다. 여기에서 `InputLayer` 클래스를 확인할 수 있습니다.
seq_model._layers
# 또는 `_input_layers` 속성에서도 확인할 수 있습니다.
seq_model._input_layers, func_model._input_layers
seq_model._output_layers, func_model._output_layers
# `Model` 클래스로 만든 `func_model`은 사실 `Functional` 클래스의 객체입니다. `Model` 클래스는 서브클래싱에 사용합니다.
func_model.__class__
# 시퀀셜 모델은 함수형 모델의 특별한 경우입니다. (`Model` --> `Functional` --> `Sequential`)
# ### 사용자 정의 층 만들기
# `tf.layers.Layer` 클래스를 상속하고 `build()` 메서드에서 가중치를 만든다음 `call()` 메서드에서 연산을 구현합니다.
class MyDense(tf.keras.layers.Layer):
def __init__(self, units, activation=None, **kwargs):
# units와 activation 매개변수 외에 나머지 변수를 부모 클래스의 생성자로 전달합니다.
super(MyDense, self).__init__(**kwargs)
self.units = units
# 문자열로 미리 정의된 활성화 함수를 선택합니다. e.g., 'softmax', 'relu'
self.activation = tf.keras.activations.get(activation)
def build(self, input_shape):
# __call__() 메서드를 호출할 때 호출됩니다. 가중치 생성을 지연합니다.
# 가중치와 절편을 생성합니다.
self.kernel = self.add_weight(name='kernel',
shape=[input_shape[-1], self.units],
initializer='glorot_uniform' # 케라스의 기본 초기화
)
self.bias = self.add_weight(name='bias',
shape=[self.units],
initializer='zeros')
def call(self, inputs): # training=None은 training은 배치 정규화나 드롭아웃 같은 경우 사용
# __call__() 메서드를 호출할 때 호출됩니다.
# 실제 연산을 수행합니다. [batch_size, units]
z = tf.matmul(inputs, self.kernel) + self.bias
if self.activation:
return self.activation(z)
return z
# +
inputs = tf.keras.layers.Input(784)
# Layer.__call__() --> MyDense().build() --> Layer.build() --> MyDense().call()
outputs = MyDense(units=10, activation='softmax')(inputs)
my_dense_model = tf.keras.Model(inputs, outputs)
my_dense_model.summary()
# -
my_dense_model.compile(loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
my_dense_model.fit(X_train, y_train, batch_size=32, epochs=2)
# ### 사용자 정의 모델 만들기
# fit(), compile(), predict(), evaluate() 등의 메서드 제공
class MyModel(tf.keras.Model):
def __init__(self):
super(MyModel, self).__init__()
self.output_layer = MyDense(units=10, activation='softmax')
def call(self, inputs):
return self.output_layer(inputs)
# +
my_model = MyModel()
my_model.compile(loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
my_model.fit(X_train, y_train, batch_size=32, epochs=2)
# -
# ### 사용자 정의 훈련
class MyCustomStep(MyModel):
def train_step(self, data):
# fit()에서 전달된 데이터
x, y = data
# 그레이디언트 기록 시작
with tf.GradientTape() as tape:
# 정방향 계산
y_pred = self(x)
# compile() 메서드에서 지정한 손실 계산
loss = self.compiled_loss(y, y_pred)
# 훈련가능한 파라미터에 대한 그레이디언트 계산
gradients = tape.gradient(loss, self.trainable_variables)
# 파라미터 업데이트
self.optimizer.apply_gradients(zip(gradients, self.trainable_variables))
# TF 2.4에서는
# self.optimizer.minimize(loss, self.trainable_variables, tape=tape)
# compile() 메서드에서 지정한 지표 계산
self.compiled_metrics.update_state(y, y_pred)
# 현재까지 지표와 결괏값을 딕셔너리로 반환
return {m.name: m.result() for m in self.metrics}
# +
my_custom_step = MyCustomStep()
my_custom_step.compile(loss='sparse_categorical_crossentropy',
metrics=['accuracy'])
my_custom_step.fit(X_train, y_train, batch_size=32, epochs=2)
# -
| custom_model_in_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Arjun
# language: python
# name: arjun
# ---
# ## GPT Paragraph Similarity using LSTM- head
#
# GPT gives good features for sentence embeddings. These embeddings seem to be separated well between in-domain and out-of-domain topics when measured using cosine similarity.
#
# Paragraph embeddings can be constructed using a linear combination of sentence embeddings. When a naive summing of embeddings was performed, the model failed to construct a reliable paragraph embedding. On tweaking the algorithm to perform summed aggregation of embeddings on groups of sentences such that their combined length was less than the max permissible length of the model, better results were observed. It was noticed however that the last set of sentences seemed to influence the paragraph the most and would skew the results of the paragraph embedding comparison (using cosine similarity metric).
#
# There are a few possible solutions to this problem:
# 1. Use a different metric.
# - Not explored much.
# 2. Divide the paragraph equally into chunks and then feed them into the model before aggregating
# - Improves scores but last sentence bias is not completely negated.
# 3. Use an additional neural network as an aggregator of these sentence embeddings in order to learn paragraph embeddings in a non-linear space. These networks (possibly LSTM based) could be trained on the objective to learn paragraph features from sentence features based on cosine similarity loss.
# - Unidirectional LSTM was prone to bias of last sentence. The bias reduced after shifting to a bidirectional LSTM. The Bi-LSTM was trained by performing cosine similarity between outputs and next/previous inputs for forward/backward cells. Bi-LSTM bi-sequential loss calculation gave the best results.
# 4. Train GPT as a language model in order to remove influence of last sentence on the score.
# - The GPT LM model with an LSTM head is averse to addition of non-domain topics at the end of the paragraph but does not capture context as well as the GPT with Multi Choice Head model, hence was eliminated for consideration of final approach.
#
# %matplotlib inline
# +
import argparse
import os
import csv
import random
import logging
from tqdm import tqdm, trange, tqdm_notebook
from math import ceil
import numpy as np
import torch
import torch.nn as nn
from itertools import combinations, product
from torch.utils.data import (DataLoader, RandomSampler, SequentialSampler,
TensorDataset)
from pytorch_pretrained_bert import (OpenAIGPTDoubleHeadsModel, OpenAIGPTTokenizer,
OpenAIAdam, cached_path, WEIGHTS_NAME, CONFIG_NAME)
from pytorch_pretrained_bert.modeling_openai import OpenAIGPTPreTrainedModel,OpenAIGPTDoubleHeadsModel,OpenAIGPTConfig,OpenAIGPTModel,OpenAIGPTLMHead
from scipy.spatial.distance import cosine, cityblock
logging.basicConfig(format = '%(asctime)s - %(levelname)s - %(name)s - %(message)s',
datefmt = '%m/%d/%Y %H:%M:%S',
level = logging.INFO)
logger = logging.getLogger(__name__)
# +
class OpenAIGPTLMHead_custom(nn.Module):
""" Language Model Head for the transformer """
def __init__(self, model_embeddings_weights, config):
super(OpenAIGPTLMHead_custom, self).__init__()
self.n_embd = config.n_embd
self.vocab_size = config.vocab_size
self.predict_special_tokens = config.predict_special_tokens
embed_shape = model_embeddings_weights.shape
#print("shape check",(model_embeddings_weights[1]))
self.decoder = nn.Linear(embed_shape[1], embed_shape[0], bias=False)
self.set_embeddings_weights(model_embeddings_weights)
def set_embeddings_weights(self, model_embeddings_weights, predict_special_tokens=True):
self.predict_special_tokens = predict_special_tokens
embed_shape = model_embeddings_weights.shape
self.decoder.weight = model_embeddings_weights # Tied weights
def forward(self, hidden_state):
# print('decoder weight')
# print((hidden_state.shape))
lm_logits = self.decoder(hidden_state)
# print(lm_logits.shape)
if not self.predict_special_tokens:
lm_logits = lm_logits[..., :self.vocab_size]
# print("lm_logits.shape: ",lm_logits.shape)
return lm_logits
class OpenAIGPTMultipleChoiceHead_custom(nn.Module):
""" Classifier Head for the transformer """
def __init__(self, config):
super(OpenAIGPTMultipleChoiceHead_custom, self).__init__()
self.n_embd = config.n_embd
self.dropout = nn.Dropout2d(config.resid_pdrop) # To reproduce the noise_shape parameter of TF implementation
self.linear = nn.Linear(config.n_embd, 1)
nn.init.normal_(self.linear.weight, std=0.02)
nn.init.normal_(self.linear.bias, 0)
def forward(self, hidden_states, mc_token_ids):
# Classification logits
# hidden_state (bsz, num_choices, seq_length, hidden_size)
# mc_token_ids (bsz, num_choices)
mc_token_ids = mc_token_ids.unsqueeze(-1).unsqueeze(-1).expand(-1, -1, -1, hidden_states.size(-1))
multiple_choice_h = hidden_states.gather(2, mc_token_ids).squeeze(2)
return multiple_choice_h
class OpenAIGPTDoubleHeadsModel_custom(OpenAIGPTPreTrainedModel):
"""
OpenAI GPT model with a Language Modeling and a Multiple Choice head ("Improving Language Understanding by Generative Pre-Training").
OpenAI GPT use a single embedding matrix to store the word and special embeddings.
Special tokens embeddings are additional tokens that are not pre-trained: [SEP], [CLS]...
Special tokens need to be trained during the fine-tuning if you use them.
The number of special embeddings can be controled using the `set_num_special_tokens(num_special_tokens)` function.
The embeddings are ordered as follow in the token embeddings matrice:
[0, ----------------------
... -> word embeddings
config.vocab_size - 1, ______________________
config.vocab_size,
... -> special embeddings
config.vocab_size + config.n_special - 1] ______________________
where total_tokens_embeddings can be obtained as config.total_tokens_embeddings and is:
total_tokens_embeddings = config.vocab_size + config.n_special
You should use the associate indices to index the embeddings.
Params:
`config`: a OpenAIGPTConfig class instance with the configuration to build a new model
`output_attentions`: If True, also output attentions weights computed by the model at each layer. Default: False
`keep_multihead_output`: If True, saves output of the multi-head attention module with its gradient.
This can be used to compute head importance metrics. Default: False
Inputs:
`input_ids`: a torch.LongTensor of shape [batch_size, num_choices, sequence_length] with the BPE token
indices selected in the range [0, total_tokens_embeddings[
`mc_token_ids`: a torch.LongTensor of shape [batch_size, num_choices] with the index of the token from
which we should take the hidden state to feed the multiple choice classifier (usually last token of the sequence)
`position_ids`: an optional torch.LongTensor with the same shape as input_ids
with the position indices (selected in the range [0, config.n_positions - 1[.
`token_type_ids`: an optional torch.LongTensor with the same shape as input_ids
You can use it to add a third type of embedding to each input token in the sequence
(the previous two being the word and position embeddings).
The input, position and token_type embeddings are summed inside the Transformer before the first
self-attention block.
`lm_labels`: optional language modeling labels: torch.LongTensor of shape [batch_size, num_choices, sequence_length]
with indices selected in [-1, 0, ..., total_tokens_embeddings]. All labels set to -1 are ignored (masked), the loss
is only computed for the labels set in [0, ..., total_tokens_embeddings]
`multiple_choice_labels`: optional multiple choice labels: torch.LongTensor of shape [batch_size]
with indices selected in [0, ..., num_choices].
`head_mask`: an optional torch.Tensor of shape [num_heads] or [num_layers, num_heads] with indices between 0 and 1.
It's a mask to be used to nullify some heads of the transformer. 1.0 => head is fully masked, 0.0 => head is not masked.
Outputs:
if `lm_labels` and `multiple_choice_labels` are not `None`:
Outputs a tuple of losses with the language modeling loss and the multiple choice loss.
else: a tuple with
`lm_logits`: the language modeling logits as a torch.FloatTensor of size [batch_size, num_choices, sequence_length, total_tokens_embeddings]
`multiple_choice_logits`: the multiple choice logits as a torch.FloatTensor of size [batch_size, num_choices]
Example usage:
```python
# Already been converted into BPE token ids
input_ids = torch.LongTensor([[[31, 51, 99], [15, 5, 0]]]) # (bsz, number of choice, seq length)
mc_token_ids = torch.LongTensor([[2], [1]]) # (bsz, number of choice)
config = modeling_openai.OpenAIGPTOpenAIGPTMultipleChoiceHead_customOpenAIGPTMultipleChoiceHead_customConfig()
model = modeling_openai.OpenAIGPTDoubleHeadsModel(config)
lm_logits, multiple_choice_logits = model(input_ids, mc_token_ids)
```
"""
def __init__(self, config, output_attentions=False, keep_multihead_output=False):
super(OpenAIGPTDoubleHeadsModel_custom, self).__init__(config)
self.transformer = OpenAIGPTModel(config, output_attentions=False,
keep_multihead_output=keep_multihead_output)
self.lm_head = OpenAIGPTLMHead_custom(self.transformer.tokens_embed.weight, config)
self.multiple_choice_head = OpenAIGPTMultipleChoiceHead_custom(config)
self.apply(self.init_weights)
def set_num_special_tokens(self, num_special_tokens, predict_special_tokens=True):
""" Update input and output embeddings with new embedding matrice
Make sure we are sharing the embeddings
"""
#self.config.predict_special_tokens = self.transformer.config.predict_special_tokens = predict_special_tokens
self.transformer.set_num_special_tokens(num_special_tokens)
self.lm_head.set_embeddings_weights(self.transformer.tokens_embed.weight, predict_special_tokens=predict_special_tokens)
def forward(self, input_ids, mc_token_ids, lm_labels=None, mc_labels=None, token_type_ids=None,
position_ids=None, head_mask=None):
hidden_states = self.transformer(input_ids, position_ids, token_type_ids, head_mask)
if self.transformer.output_attentions:
all_attentions, hidden_states = hidden_states
hidden_states = hidden_states[-1] #layer #
lm_logits = self.lm_head(hidden_states)
# No input to Multi-Choice head as it gives same output as hidden_states[pos_of_clf_token] during inference
# losses = []
# if lm_labels is not None:
# shift_logits = lm_logits[..., :-1, :].contiguous()
# shift_labels = lm_labels[..., 1:].contiguous()
# loss_fct = nn.CrossEntropyLoss(ignore_index=-1)
# losses.append(loss_fct(shift_logits.view(-1, shift_logits.size(-1)), shift_labels.view(-1)))
return lm_logits, hidden_states #
# -
# +
def accuracy(out, labels):
outputs = np.argmax(out, axis=1)
return np.sum(outputs == labels)
def listRightIndex(alist, value):
return len(alist) - alist[-1::-1].index(value) -1
def pre_process_datasets(encoded_datasets, input_len, cap_length, start_token, delimiter_token, clf_token):
""" Pre-process datasets containing lists of story
To Transformer inputs of shape (n_batch, n_sentence, length) comprising for each batch:
input_ids[batch,n_sentence, :] = [start_token] + story[:cap_length] + [clf_token]
"""
tensor_datasets = []
for dataset in encoded_datasets:
n_batch = ceil(len(dataset[0][0])/cap_length)
input_ids = np.zeros((n_batch, 1, input_len), dtype=np.int64)
mc_token_ids = np.zeros((n_batch, 1), dtype=np.int64)
lm_labels = np.full((n_batch, 1, input_len), fill_value=-1, dtype=np.int64)
mc_labels = np.zeros((n_batch,), dtype=np.int64)
i = 0
init_pos = 0
end_pos = cap_length
for story, cont1, cont2, mc_label in dataset:
if n_batch!=0:
if n_batch==1:
with_cont1 = [start_token] + story[:cap_length] + [clf_token]
input_ids[i, 0, :len(with_cont1)] = with_cont1
mc_token_ids[i, 0] = len(with_cont1) - 1
lm_labels[i, 0, :len(with_cont1)] = with_cont1
mc_labels[i] = mc_label
i+=1
else:
while i!=n_batch and end_pos<len(story):
try:
end_pos = init_pos + listRightIndex(story[init_pos:end_pos],story[-1])
except ValueError:
end_pos = init_pos+story[init_pos:].index(story[-1])
with_cont1 = [start_token] + story[init_pos:end_pos+1] + [clf_token]
input_ids[i, 0, :len(with_cont1)] = with_cont1
mc_token_ids[i, 0] = len(with_cont1) - 1
lm_labels[i, 0, :len(with_cont1)] = with_cont1
mc_labels[i] = mc_label
i+=1
init_pos = end_pos+1
end_pos = min(init_pos+cap_length-1,len(story))
all_inputs = (input_ids, mc_token_ids, lm_labels, mc_labels)
tensor_datasets.append(tuple(torch.tensor(t) for t in all_inputs))
return tensor_datasets
def load_rocstories_dataset(dataset_path):
""" Output a list of tuples(story, 1st continuation, 2nd continuation, label) """
with open(dataset_path, encoding='utf_8') as f:
f = csv.reader(f)
output = []
next(f) # skip the first line
for line in tqdm(f):
output.append(('.'.join(line[0 :4]), line[4], line[5], int(line[-1])))
return output
def tokenize_and_encode(obj):
""" Tokenize and encode a nested object """
if isinstance(obj, str):
return tokenizer.convert_tokens_to_ids(tokenizer.tokenize(obj))
elif isinstance(obj, int):
return obj
return list(tokenize_and_encode(o) for o in obj)
# -
def pre_process_datasets_cos(encoded_datasets, input_len, cap_length,start_token, delimiter_token, clf_token):
""" Pre-process datasets containing lists of stories(paragraphs)
To Transformer inputs of shape (n_batch, n_sentences, length) comprising for each batch, continuation:
input_ids[batch, alternative, :] = [start_token] + story[:cap_length] + [full_stop_id] + [clf_token]
"""
# print("clf_token",clf_token)
tensor_datasets = []
for dataset in encoded_datasets:
#print(dataset)
n_batch = len(dataset)
input_ids = np.zeros((n_batch, 5, input_len), dtype=np.int64)
mc_token_ids = np.zeros((n_batch, 5), dtype=np.int64)
for i, stories in enumerate(dataset):
sents=[]
story = stories[0]
size = len(story)
idx_list = [idx + 1 for idx, val in enumerate(story) if val == 239]
res = [story[i: j] for i, j in zip([0] + idx_list, idx_list + \
([size] if idx_list[-1] != size else []))]
for sent in res:
# print("sent",sent,cap_length)
sents.append([start_token] + sent[:cap_length]+[239] + [clf_token])
for j in range(len(sents)):
input_ids[i, j,:len(sents[j])] = sents[j]
mc_token_ids[i,j] = len(sents[j]) - 1
all_inputs = (input_ids, mc_token_ids)
tensor_datasets.append(tuple(torch.tensor(t) for t in all_inputs))
return tensor_datasets
# +
## Defining constants over here
seed = 42
model_name = 'openai-gpt'
do_train = False
output_dir = '/home/shubham/projects/domain_minds/gpt-experiment/model/'
train_batch_size = 1
device = torch.device("cuda" if torch.cuda.is_available() else "cpu")
n_gpu = torch.cuda.device_count()
logger.info("device: {}, n_gpu {}".format(device, n_gpu))
special_tokens = ['_start_', '_delimiter_', '_classify_']
tokenizer = OpenAIGPTTokenizer.from_pretrained(model_name, special_tokens=special_tokens)
special_tokens_ids = list(tokenizer.convert_tokens_to_ids(token) for token in special_tokens)
model1 = OpenAIGPTDoubleHeadsModel_custom.from_pretrained(output_dir)
tokenizer = OpenAIGPTTokenizer.from_pretrained(output_dir)
model1.to(device)
model1.eval()
tokenizer = OpenAIGPTTokenizer.from_pretrained(output_dir)
logger.info("Ready to encode dataset...")
def feature_extractor(model1,text):
trn_dt = ([text,'','',0],)
datasets = (trn_dt,)
encoded_datasets = tokenize_and_encode(datasets)
# Compute the max input length for the Transformer
# max_length = min(510,ceil(len(encoded_datasets[0][0][0])/ 2)) # For multisentence inputs
max_length = model1.config.n_positions//2 - 2
input_length = len(encoded_datasets[0][0][0])+2 # +2 for start and clf token
input_length = min(input_length, model1.config.n_positions) # Max size of input for the pre-trained model
# Prepare inputs tensors and dataloaders
n_batches = ceil(len(encoded_datasets[0][0][0])/max_length)
tensor_datasets = pre_process_datasets(encoded_datasets, input_length, max_length, *special_tokens_ids)
train_tensor_dataset = tensor_datasets[0]
train_data = TensorDataset(*train_tensor_dataset)
train_dataloader = DataLoader(train_data, batch_size=1)
'''
config = OpenAIGPTConfig.from_json_file('/home/shubham/Project/domain_mind/gpt2_experiment/model/config.json')
model1 = OpenAIGPTMultipleChoiceHead_custom(config)
'''
#eval_loss, eval_accuracy = 0, 0
#nb_eval_steps, nb_eval_examples = 0, 0
final_clf=[]
final_lm=[]
for batch in train_dataloader:
batch = tuple(t.to(device) for t in batch)
input_ids, mc_token_ids, lm_labels, mc_labels = batch
with torch.no_grad():
a, clf_text_feature = model1(input_ids, mc_token_ids, lm_labels, mc_labels)
final_clf.append(clf_text_feature[:,:,-1])
if n_batches>1:
clf_torch = torch.sum(torch.stack(final_clf),0)
return clf_torch
else:
return clf_text_feature[:,:,-1,:]#, lm_text_feature
# -
def load_rocstories_dataset(dataset_path):
""" Output a list of tuples(story, 1st continuation, 2nd continuation, label) """
with open(dataset_path, encoding='utf_8') as f:
f = csv.reader(f)
output = []
next(f) # skip the first line
for line in tqdm(f):
output.append(('.'.join(line[0 :4]), line[4], line[5], int(line[-1])))
return output
# +
train_dataset = '/home/ether/Desktop/gpt_experiments/data/data_para_se_5sent.csv'
import pandas as pd
train_dataset = pd.read_csv(train_dataset,index_col=0)
encoded_datasets = tokenize_and_encode((train_dataset.drop("Num_sentences",axis=1).values,))
max_length = model1.config.n_positions // 2 - 2
input_length = max_length+5
# # Prepare inputs tensors and dataloaders
tensor_datasets = pre_process_datasets_cos(encoded_datasets, input_length, max_length,*special_tokens_ids)
train_tensor_dataset = tensor_datasets[0]
train_data = TensorDataset(*train_tensor_dataset)
train_sampler = RandomSampler(train_data)
train_dataloader = DataLoader(train_data, sampler=train_sampler, batch_size=train_batch_size)
# +
# # Uni/ Bi-LSTM unisequential code
# # for uni, make bidirectional False for self.lstm and set hidden to sizes(1,1,768)
# # for bidirectional unisequential, there is only one loss backprop.
# # make bidirectional True for self.lstm and set hidden to sizes(2,1,768). Uncomment code For Bidirectional.
# class LSTM_Head(nn.Module):
# def __init__(self):
# super(LSTM_Head, self).__init__()
# self.lstm = nn.LSTM(768,768,batch_first=True,bidirectional=False)
# self.linear = nn.Linear(768*2,768)
# def forward(self,input_embeds,mc_token_ids=None,infer=False):
# hidden = (torch.zeros((1,1,768),device=device), \
# torch.zeros((1,1,768),device=device))
# cosloss = nn.CosineSimilarity(dim=-1)
# m = nn.Softmax()
# loss = 0
# hidden_states=[]
# for i in range(len(input_embeds)):
# if not infer:
# # prev_hid,prev_cst = hidden # For Bidirectional
# out, hidden = self.lstm(input_embeds[i][mc_token_ids[i].item()].view(1,1,-1),hidden)
# # hid = torch.sum(torch.stack([hidden[0],prev_hid]),0) # For Bidirectional
# # cst = torch.sum(torch.stack([hidden[1],prev_cst]),0) # For Bidirectional
# # hidden=(hid,cst) # For Bidirectional
# # out = self.linear(out) # For Bidirectional
# if i!=len(input_embeds)-1:
# loss += 1 - cosloss(out,input_embeds[i+1][mc_token_ids[i+1]])
# else:
# # During inference the last output of last lstm cell is considered as paragraph embedding
# out, hidden = self.lstm(input_embeds[i].view(1,1,-1),hidden)
# # out = self.linear(out) # For Bidirectional inference
# if infer:
# return out
# loss = loss/(len(input_embeds)-1)
# return loss
# -
#Bi-LSTM bi-sequential code for truebi files
class LSTM_Head(nn.Module):
def __init__(self):
super(LSTM_Head, self).__init__()
self.lstm = nn.LSTM(768,768,bidirectional=True)
self.linear = nn.Linear(768*2,768)
def forward(self,input_embeds,mc_token_ids=None,infer=False):
hidden = (torch.zeros((2,1,768),device=device), \
torch.zeros((2,1,768),device=device))
# For Cosine Distance
cosloss = nn.CosineSimilarity(dim=-1)
loss = 0
if not infer:
inputs=torch.cat([input_embeds[i][mc_token_ids[i].item()] for i in range(len(input_embeds))]).view(len(input_embeds),1,-1)
out, hidden = self.lstm(inputs,hidden)
lossf=0
lossb=0
outs = out.view(5,2,-1)
for i in range(len(inputs)):
if i!=len(inputs)-1:
# Forward loss claculated as 1-cosloss(current_cell_output,next_cell_input)
lossf += 1-cosloss(outs[i,0],inputs[i+1])
# lossf += cosloss(outs[i,0],inputs[i+1]).acos()/np.pi # Making cosine between (0,1)
if i!=0:
# Backward loss claculated as 1-cosloss(current_cell_output,previous_cell_input)
lossb += 1-cosloss(outs[i,1],inputs[i-1])
# lossb += cosloss(outs[i,1],inputs[i-1]).acos()/np.pi # Making cosine between (0,1)
lossf = lossf/(len(inputs)-1)
lossb = lossb/(len(inputs)-1)
loss = (lossf+lossb)/2
return loss,lossf,lossb
else:
# During inference, output of first lstm_cell(reverse direction) and last lstm_cell(forward direction)
# are concatenated to give the paragraph embedding
out, hidden = self.lstm(input_embeds.view(len(input_embeds),1,-1),hidden)
return hidden[0].view(1,1,-1)
model1.eval()
model = LSTM_Head()
# state_dict = torch.load("../models/lstmheadSGD_bi_mcpos_real_ep2.pt")
# model.load_state_dict(state_dict)
model.to(device)
model.train()
print()
# TRAINING
num_train_epochs = 10
optimizer = torch.optim.SGD(model.parameters(),lr = 1e-5)
for i in tqdm_notebook(range(num_train_epochs)):
tr_loss = 0
nb_tr_steps = 0
tqdm_bar = tqdm_notebook(train_dataloader, desc="Training")
for step, batch in enumerate(tqdm_bar):
batch = tuple(t.to(device) for t in batch)
input_ids, mc_token_ids= batch
with torch.no_grad():
_, sent_feats = model1(input_ids,mc_token_ids)
loss, lossf,lossb= model.forward(sent_feats[0], mc_token_ids[0])
loss.backward()
optimizer.step()
optimizer.zero_grad()
nb_tr_steps += 1
tqdm_bar.desc = "Training losses: {:.2e} {:.2e} {:.2e}".format(loss.item(),lossf.item(),lossb.item())
torch.save(model.state_dict(), "/home/ether/Desktop/gpt_experiments/models/lstmheadSGD_truebi_mcpos_torchcos_ep"+str(i)+".pt")
# ### Testing Ground for LSTM head-based paragraph embeddings
# +
# Collection of paragraphs separated by "\n"
para_docker = '''
Docker is a containerization platform that packages your app and all its dependencies together in the form called a docker container to ensure that your application works seamlessly in any environment. This environment might be a production or staging server. Docker pulls the dependencies needed for your application to run from the cloud and configures them automatically. You don’t need to do any extra work. Cool Right.
Docker communicates natively with the system kernel by passing the middleman on Linux machines and even Windows 10 and Windows Server 2016 and above this means you can run any version of Linux in a container and it will run natively. Not only that Docker uses less disk space to as it is able to reuse files efficiently by using a layered file system. If you have multiple Docker images using the same base image for instance.
Imagine we already have an application running PHP 5.3 on a server and want to deploy a new application which requires PHP 7.2 on that same server. This will cause some version conflict on that server and also might cause some features in the existing application to fail.
In situations like this, we might have to use Docker to sandbox or containerise the new application to run without affecting the old application. This brings us to Docker containers.
Think of a Docker container as above image. There are multiple applications running on the same machine. These applications are put into docker containers and any changes made on these containers does not affect the other container. Each container has different Os running on the same physical machine. Docker helps you to create, deploy and run applications using containers.
A container packages up the code and all its dependencies so the application runs quickly and reliably from one computing environment to another.
A Docker container image is a lightweight, standalone, executable package of software that includes everything needed to run an application: code, runtime, system tools, system libraries and settings.
'''
para_infra = '''
Infrastructure software is a type of enterprise software or program specifically designed to help business organizations perform basic tasks such as workforce support, business transactions and internal services and processes. The most common examples of infrastructure software are database programs, email and other communication software and security applications.
Infrastructure software is used to ensure that people and systems within an organization can connect and do their jobs properly and ensure the efficient execution of business processes, share information, as well as manage touch points with suppliers and customers. This type of software is not necessarily marketing related or used for business transactions such as selling products and services, but is more operations related, ensuring that business applications and processes can keep running effectively.
Infrastructure software can be configured to automatically alert users about best practices and relevant discoveries based on their current activities and job position. Expert systems and knowledge systems fall under this category.
Management of converged infrastructure resources is typically handled by a discrete hardware component that serves a singular purpose. While hyper-converged infrastructure systems are similar in nature to converged infrastructure systems, management of the resources is largely software-defined rather than being handled by one or more hardware components.
Human computation studies need not have extensive or complex software infrastructure. Studies can easily be run through homegrown or customized web applications, together with logging software capable of tracking the details and time of any given interaction. One productive approach for such tools might be to build a database-driven web application capable of storing appropriate demographic background information associated with each participant, along with details of each action and task completed.
You might even add an administrative component capable of managing and enrolling prospective participants. These homegrown applications are generally not terribly difficult to construct, particularly if you have a web-based implementation of the key tasks under consideration, or were planning on building one anyway. For some tasks—particularly those involving collection of fine-grained detail or requiring complex interactions—the freedom associated with constructing your own application may be necessary to get the job done.
The infrastructure provided by Mechanical Turk and similar crowdsourcing platforms provides many advantages over “roll-your-own” designs. As any experienced HCI researcher knows well, the challenges of recruiting, enrolling, and consenting participants can consume substantial amounts of time. Even if you are able to build your own web application to do the trick, you might find that leveraging these platforms—particularly with one of the add-on libraries—might simplify your life considerably. These advantages aside, commercial crowdsourcing tools have potential downsides.
'''
para_mark = '''
I used to find a lot of my illustrators and designers by trawling websites such as Folksy and Etsy.Funnily enough, I always prefered Folksy as an option because we were a UK-based shop, and it made a lot more sense for me to buy from UK designers.I am also a little scared sometimes by the monster that is Etsy! It’s always a minefield having to find out whether someone will ship internationally, let alone navigating tens of thousands of pages of products.
I also find a lot of people on Pinterest, Twitter, Facebook, graduate fairs, local and national exhibitions and design shows so make sure you are linked in with as many of those as you can. Things to be mindful of: The most obvious is email size, but other things to look out for are not putting any images at all in an email. If you are pitching your work, always make sure that you have pictures of this work included — quite a lot of people forget this.
This PDF therefore. It's their signature. Like when did that decision happen? Someone said that was a thing. Yeah. That's a great question. I'm not sure what exactly happened. I would say. My guess is that it happened in the age of the internet, but it's interesting that you bring that up as that is it just because somebody clicked on this PDF and that means it's a signature. I remember back in law school when I studied estate planning and development of wills and trusts and things like that for individuals there.
There is a some sort of Statute the state of Texas where if you just made if that individual just create some sort of marketing. It doesn't necessarily have to be their actual signature or even their name. It could be some sort of marking and that constitutes as a signature or you know authorization if they're granting authorization or whatever. It may be agreement contract if it is a written agreement and.
So I think it's I think the market demanded it because of you know, faxing becoming a thing of the past somewhat. I don't know to fax machine, but I know most Law Offices do have pads machines and courts still use fax machines. And when I hear when I told the court and they don't have use email and like what like what how but it is right and I think it's it's also a matter of investing in other resources in bringing things up to date which hey and in their minds it might be a matter of if it's not broken. We're not going to fix it. You know, it works just fine. Everybody knows how we operate. We'll keep doing it until we can't be morons. Yeah because nobody's objected at this point. So yeah, I don't I don't know when that came about but that's a great question. Yeah. I find it comforting. You don't have a fax machine somehow. I just imagine that.
Ever since their executive shakeup that resulted in Instagram’s original founders being softly pushed out of their positions only to be replaced with Facebook loyalists, Instagram has been toying around with features they’re claiming will make their platform a safer place for their users — from hiding Likes to their recent anti-bullying updates.
While the intention behind these features might be well and good, the changes make the product deviate from the core things that made Instagram spread like wildfire in the first place — specifically by the platform deciding which content you see versus the end user being in control of their in-app experience.Plus, one can’t help to wonder if Facebook is using Instagram company as a buffer, or band-aid, for their own recent mishaps and privacy scandals, which have caused many users to lose faith in the platform.
'''
para_db = '''
SQL is a query language for talking to structured databases. Pretty much all databases that have tables and rows will accept SQL based queries. SQL has many flavors but the fundamentals stay the same. Professionals and amateurs alike use SQL to find, create, update and delete information from their sources of record. It can be used with a ton of different databases like MySQL, Postgres, SQL Server and Oracle. It powers the logic behind popular server-side frameworks like Ruby On Rails and Laravel. If you want to find information associated with a particular account or query what buttons users click in your app there is a good chance SQL can help you out.
Before we hop on the SQL train to Database Town I’d like to acknowledge some alternatives. You can use ORMs to query databases. ORM stands for Object Relational Mapper, which is a fancy way of saying that you can write code in a programming language like PHP or Node.js that translates to SQL queries. Popular ORMs are Active Record for Ruby On Rails, Eloquent for Laravel and Sequelize for Node.js. All of these services allow you to write code that translates to SQL under the hood. SQL is important for building applications with these technologies.
There are many databases that do not use SQL, such as MongoDB and GraphQL. These are newer technologies and not as widely adopted as relational databases. Relational databases have been around a very long time and power the majority of data storage on the internet. To fully appreciate NoSQL technologies and the reasons they came about it’s helpful to know how relational databases and SQL work.
Oracle Corporation provides a range of database cloud services on its Oracle Cloud platform that are designed for different database use cases; from test/dev deployments to small and medium sized workloads to large mission-critical workloads. Oracle Database Cloud Services are available on a choice of general purpose hardware and Exadata engineered systems, in either virtual machines environments or 'bare metal' infrastructure (now known as Oracle Cloud Infrastructure).
Moving away from your database vendor would be like cutting off a foot; self-destructive and painful. More to the point, building a me-too product and entering a full-on competition with the established leaders, is a significantly retrograde step with little tradition of success. Beyond the relational database, there are many new wrinkles that offer attractive niches such as virtual machines, bare metal servers, serverless technologies and micro apps. But I am not seeing a great deal of competition heating up in that space.
Databases are a structured system to put your data in that imposes rules upon that data, and the rules are yours, because the importance of these problems changes based on your needs. Maybe your problem is the size, while someone else has a smaller amount of data where the sensitivity is a high concern.It’s the things you can’t see that are going on in the background; the security, the enforced integrity of the data, the ability to get to it fast and get to it reliably, the robustness; serving lots of people at the same time and even correctly survive crashes and hardware issues without corrupting the data.
In practice it’s very common to have multiple databases. The database that deals with your order and customer information might be completely independent from you database that deals with human resource information. And in many organizations, you don’t just have multiple databases but multiple DBMS. Sometimes it’s because one DBMS is better at something than the other.
'''
para_news = '''
<NAME>, while leaving the NCP, which his father <NAME> has had a long association with, claimed that his shift to the BJP is “keeping the general public’s interest in mind.” This, a senior NCP leader said, was hogwash. “Just because a certain party is winning doesn’t mean they have the public interest in mind. And to claim that someone is leaving a party keeping the public’s interest in mind is a plain lie. These politicians have all proved to be opportunists and don’t care for any party ideology or its legacy,” a senior leader said, requesting anonymity. Several decisions, particularly the one to give 16% reservation to the Maratha caste in the state, has worked in the ruling party’s favour. With the Bombay high court approving the state’s decision, sources in the BJP have indicated that several senior Maratha leaders have been warming up to the party.
When asked if the party accepts its failure in handling the situation, Nirupam said, “I would rather blame the saffron force here. Does the BJP not trust its own cadres to ensure its victory in the state? Why does it need the Congress leadership then?” Bal terms this trend as a “destructive” one. “They are in a destructive mode right now. They want to ensure there is no opposition in the state. They are sure to win the state assembly elections. But before that, the BJP wants to clear off the Congress-NCP from the state. What happened in Karnataka is also playing out in Maharashtra. In fact, it is a trend across the country. Most of these new entrants might not even have any substantial roles to play in the party. But there they are,” Bal points out.
Hindutva bigots also targeted Hindustan Unilever’s Surf Excel ad campaign #RangLayeSang, which featured a young (Hindu) girl helping a young (Muslim) boy in March of this year. Earlier that month they had also targeted a tea brand (Brook Bond) for ‘projecting the Kumbh in the wrong light’ by showing a (presumably Hindu) man deliberately attempting to abandon his father there. The troll brigade aimed to boycott all HUL products, trending the hashtag #BoycottHUL on twitter with pictures of an assortment of products they had bought in the trash. This is also not an India specific trend – across America as well, conservative groups have protested brands taking up a stance against brands that support liberal causes – even when they are as vague as the Gilette ad in January of this year.
The Planetary Society, a non-profit organisation, has been working on the LightSail programme for a decade. The project kicked off in the 1990s, but its first planned prototype, Cosmos 1, was destroyed during a faulty launch on a Russian rocket taking off from a submarine in 2005. The Planetary Society got its the next prototype, LightSail 1, into space in 2015, but technical problems kept it from climbing high enough to be steered by sunlight. The LightSail 2 spacecraft was launched on June 25 and has since been in a low-Earth orbit, according to The Verge. Last week, it deployed four triangular sails – a thin, square swath of mylar about the size of a boxing ring. After launch, engineers on the ground have been remotely adjusting the orientation of the sails to optimise the LightSail 2’s ability to harness solar photons.
Solar sailing isn’t new but the Planetary Society wanted to show that the technique could be used for smaller satellites, which are harder to manoeuvre through space. A majority of the satellites, as senior science reporter <NAME> explained on The Verge, have to rely on thrusters to be mobile. These are “tiny engines that combust chemical propellants to push a vehicle through space.” However, this increases the cost of satellites as well as their launch mass. Smaller satellites like CubeSats cannot accommodate thrusters most of the time, nor can they be closely manoeuvred once they are in space. But with this mission, the Planetary Society has demonstrated that solar sails can guide CubeSats through space. It is set to share the data it receives from this mission to allow other groups to build on this technology. The solar sail technology could reduce the need for expensive, cumbersome rocket propellants and slash the cost of navigating small satellites in space.
Last week’s launch of the Chandrayaan-2 water-finding Moon mission is a significant demonstration of India’s scientific and engineering capacity. It puts India firmly within a select group of countries prowling the solar system for commercial, strategic, and scientific reasons. Pakistanis naturally want to know where they stand in science – of which space exploration is just a small part – and why. What gave India this enormous lead over Pakistan? It is natural that India’s Hindutva government should boast Chandrayaan-2 as its own achievement and claim continuation with imagined glories from Vedic times. But rightfully the credit goes elsewhere. Just imagine if history could be wound back by 70-80 years and Prime Minister Jawaharlal Nehru was replaced by <NAME>.
The atheistic Nehru brought to India an acceptance of European modernity. For this Hindutva hates him even more than it hates India’s Muslims and Christians. Still, his insistence on ‘scientific temper’ – a singularly odd phrase invented while he was still in prison – made India nurture science. Earlier, vigorous reformers like Raja Ram <NAME> (1772-1833) had shown the path. As long as Nehru stood tall no rishi, yogi, or army general could head a science institution Will Pakistan also get a slice of the moon? That depends upon the quality of our scientists and if a culture of science develops. Of course, Pakistan never had a Nehru. A further setback happened in the Zia ul Haq days when Sir Syed Ahmad Khan’s modernism had its remaining flesh eaten off by Allama Iqbal’s shaheen. As if to compensate the loss of appetite for science, buildings for half-a-dozen science institutions were erected along Islamabad’s Constitution Avenue. They could be closed down today and no one would notice. Today’s situation for science – every kind except agriculture and biotechnology – is dire.
'''
para_kuber = '''
Real production apps span multiple containers. Those containers must be deployed across multiple server hosts. Security for containers is multilayered and can be complicated. That's where Kubernetes can help. Kubernetes gives you the orchestration and management capabilities required to deploy containers, at scale, for these workloads. Kubernetes orchestration allows you to build application services that span multiple containers, schedule those containers across a cluster, scale those containers, and manage the health of those containers over time. With Kubernetes you can take real steps towards better IT security.
Of course, this depends on how you’re using containers in your environment. A rudimentary application of Linux containers treats them as efficient, fast virtual machines. Once you scale this to a production environment and multiple applications, it's clear that you need multiple, colocated containers working together to deliver the individual services. This significantly multiplies the number of containers in your environment and as those containers accumulate, the complexity also grows.
Kubernetes fixes a lot of common problems with container proliferation—sorting containers together into a ”pod.” Pods add a layer of abstraction to grouped containers, which helps you schedule workloads and provide necessary services—like networking and storage—to those containers. Other parts of Kubernetes help you load balance across these pods and ensure you have the right number of containers running to support your workloads.
The primary advantage of using Kubernetes in your environment, especially if you are optimizing app dev for the cloud, is that it gives you the platform to schedule and run containers on clusters of physical or virtual machines. More broadly, it helps you fully implement and rely on a container-based infrastructure in production environments. And because Kubernetes is all about automation of operational tasks, you can do many of the same things that other application platforms or management systems let you do, but for your containers.
That’s where Red Hat OpenShift comes in. OpenShift is Kubernetes for the enterprise—and a lot more. OpenShift includes all of the extra pieces of technology that makes Kubernetes powerful and viable for the enterprise, including: registry, networking, telemetry, security, automation, and services. With OpenShift, your developers can make new containerized apps, host them, and deploy them in the cloud with the scalability, control, and orchestration that can turn a good idea into new business quickly and easily.
Kubernetes runs on top of an operating system (Red Hat Enterprise Linux Atomic Host, for example) and interacts with pods of containers running on the nodes. The Kubernetes master takes the commands from an administrator (or DevOps team) and relays those instructions to the subservient nodes. This handoff works with a multitude of services to automatically decide which node is best suited for the task. It then allocates resources and assigns the pods in that node to fulfill the requested work.
The docker technology still does what it's meant to do. When kubernetes schedules a pod to a node, the kubelet on that node will instruct docker to launch the specified containers. The kubelet then continuously collects the status of those containers from docker and aggregates that information in the master. Docker pulls containers onto that node and starts and stops those containers as normal. The difference is that an automated system asks docker to do those things instead of the admin doing so by hand on all nodes for all containers.
'''
# -
text_docker1 = "Docker communicates natively with the system kernel by passing the middleman on Linux machines and even Windows 10 and Windows Server 2016 and above this means you can run any version of Linux in a container and it will run natively. Not only that Docker uses less disk space to as it is able to reuse files efficiently by using a layered file system. If you have multiple Docker images using the same base image for instance Docker only keep a single copy of the files needed and share them with each container. All right. So, how do we use Docker install Docker on your machine and will provide links in the description begin with a Docker file, which can be built into a Docker image which can be run as a Docker container. Okay, let's break that down. The dockerfile is a surprisingly Simple Text document that instructs how the docker image will be built like a blueprint you first select a base image to start with using the from keyword, which you can find a container to use from the docker Hub. Like we mentioned before a bun to an Alpine Linux are popular choices.From there, you can run commands such as downloading installing and running your software of course will link the docks below once our Docker file is complete. We can build it using Docker build followed by the T flag so we can name our image and pass our commands the location of the dockerfile once complete. You can verify your images existence with Docker images. Now, you're built image can run a container of that image or you can push it to the cloud to share with others speaking of sharing with others. If you don't create your own Docker image and you just want to use a premade one in Poland from the docker hub using Docker full and the image names, you may also include a tag if one is available which may specify a version or variant of the software. If you don't specify a tag the latest version will be what statute to run a container pulled down from the docker Hub or build the image and then enter Docker run followed by the image name. There are of course many options available when running your containers such as running it in detached mode, but XD or assigning ports for web services, you can view your running containers with Docker container LS. And as you add more Bill appear here running a single container is fun, but it's annoying to enter all of these.Commands to get a container running and we may want to control several containers as part of a single application such as running an app and a database together something you might want to."
text_docker2 = "Docker is a tool designed to make it easier to create, deploy, and run applications by using containers. Containers allow a developer to package up an application with all of the parts it needs, such as libraries and other dependencies, and ship it all out as one package. By doing so, thanks to the container, the developer can rest assured that the application will run on any other Linux machine regardless of any customized settings that machine might have that could differ from the machine used for writing and testing the code."
text_marketing = "I used to find a lot of my illustrators and designers by trawling websites such as Folksy and Etsy.\
Funnily enough, I always prefered Folksy as an option because we were a UK-based shop, and it made a lot more sense for me to buy from UK designers.\
I am also a little scared sometimes by the monster that is Etsy! It’s always a minefield having to find out whether someone will ship internationally, let alone navigating tens of thousands of pages of products.\
I also find a lot of people on Pinterest, Twitter, Facebook, graduate fairs, local and national exhibitions and design shows so make sure you are linked in with as many of those as you can.\
Things to be mindful of: The most obvious is email size, but other things to look out for are not putting any images at all in an email.\
If you are pitching your work, always make sure that you have pictures of this work included — quite a lot of people forget this."
text_sql = "The uses of SQL include modifying database table and index structures; adding, updating and deleting rows of data; and retrieving subsets of information from within a database for transaction processing and analytics applications. Queries and other SQL operations take the form of commands written as statements -- commonly used SQL statements include select, add, insert, update, delete, create, alter and truncate.SQL became the de facto standard programming language for relational databases after they emerged in the late 1970s and early 1980s. Also known as SQL databases, relational systems comprise a set of tables containing data in rows and columns. Each column in a table corresponds to a category of data -- for example, customer name or address -- while each row contains a data value for the intersecting column."
text_se1 = "If we were to build a solution to collect and querying data related to our customers’ clinical history, probably the Software Architecture will be strongly shaped by lots of politics about how to access data, obfuscation, certificates, tracking, protocols, etc…On the other hand, if we were re-building a system because it’s unmaintainable and technologically obsolete, surely some principles about modularity, testability, new technology stack, etc… will appear.Finally, lightweight Architecture will be needed when working on a new experimental product focused on a new market niche due to the uncertainty of the product itself.Many enterprises have their own framework which implements some of the Architecture’s principles."
text_se2 = "The starting point is a bit of Software Architecture (upfront design) which is retro-feeding with the emergent design of the autonomous teams.Doing so we reach two benefits:Having a reference Architecture which helps us to build our solutionsLet the teams have a degree of innovation that, at the same time, will feed the Architecture and will allow other teams to take advantage of that.When we mean agile and autonomous teams we also refer to multi-skilled teams. Such teams are composed by dev-ops, scrum master, product owner, frontend developer, backend developer, QA, technical leader and so on."
text_nemo = "Parents need to know that even though there are no traditional bad guys in Finding Nemo, there are still some very scary moments, including large creatures with zillions of sharp teeth, the apparent death of a major character, and many tense scenes with characters in peril. And at the very beginning of the movie, Marlin's wife and all but one of their eggs are eaten by a predator -- a scene that could very well upset little kids. Expect a little potty humor amid the movie's messages of teamwork, determination, loyalty, and a father's never-ending love for his son. The issue of Nemo's stunted fin is handled exceptionally well -- matter-of-factly but frankly.Marlin's encounter with the barracuda that decimated his young family drove a permanent stake of fear through his heart. And he transfers his misgivings to his son. Instead of encouraging him to spread his wings—er, flip his fins—he shelters him to a smothering degree. This breeds anger and rebellion in Nemo and creates further unhappiness for Marlin. The film stresses the need to maintain balance in your family life and in the way you introduce your kids to the world. And an extended family of sea turtles provides insight into how steady, loving relationships can flow more smoothly."
# # For testing, use files:
# # 1) /home/ether/Desktop/gpt_experiments/models/lstmheadSGD_truebi_mcpos_torchcos_ep4.pt with Bi-LSTM non-seq
# # 2) /home/ether/Desktop/gpt_experiments/models/lstmheadSGD_uni_mcpos_torchcos_ep4.pt with Unidirectional
# # 3) /home/ether/Desktop/gpt_experiments/models/lstmheadSGD_bi_mcpos_real_ep2.pt with Bi-LSTM seq or non-seq
model = LSTM_Head()
state_dict = torch.load("/home/ether/Desktop/gpt_experiments/models/lstmheadSGD_truebi_mcpos_torchcos_ep4.pt")
model.load_state_dict(state_dict)
model.eval()
model.to(device)
m = nn.Sigmoid()
# for texta, textb in combinations(para_db.strip().split("\n"),2):
for texta, textb in product(para_docker.strip().split("\n"),para_kuber.strip().split("\n")):
with torch.no_grad():
feat1 = [feature_extractor(model1,text.strip()) for text in texta.split(".")[:-1]]
feat2 = [feature_extractor(model1,text.strip()) for text in textb.split(".")[:-1]]
in1 = torch.stack(feat1)
in2 = torch.stack(feat2)
op1 = model(in1.to(device),infer=True)
op2 = model(in2.to(device),infer=True)
print("#"*40,end="\n\n")
# Cosine score of 1 means high similarity
print("With LSTM Cosine score: ", torch.cosine_similarity(op1,op2,dim=-1).detach().cpu().item())
# Cityblock score of 0 means high similarity
print("With LSTM Cityblock score: ", cityblock(m(op1).detach().cpu(),m(op2).detach().cpu()))
print("Without LSTM sum(sent_feat_vecs) Cosine score",torch.cosine_similarity(torch.sum(in1,0),torch.sum(in2,0),dim=-1).detach().cpu().item())
print("Without LSTM sum(sent_feat_vecs) Cityblock score",cityblock(m(torch.sum(in1,0)).detach().cpu(),m(torch.sum(in2,0)).detach().cpu()))
print("*"*40,"<Para1> ",texta,"*"*40,"<Para2> ",textb,sep="\n",end="\n\n")
# lena = len(texta.split("."))-1
# lenb = len(textb.split("."))-1
# print("Lengths",lena,lenb,end="\n\n")
| gpt_paragraph/gpt-paragraph-similarity-LSTM.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.8.5 64-bit (''.venv'': venv)'
# name: python3
# ---
# # Target Outliers
# +
import datetime, time, os
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import plotly.express as px
import plotly.figure_factory as ff
import plotly.graph_objects as go
import pandas as pd
from functools import reduce
from sklearn.metrics import mean_absolute_error, mean_squared_error
# Make numpy printouts easier to read.
np.set_printoptions(precision=3, suppress=True)
import plotly.graph_objects as go
# Create traces
import warnings
warnings.filterwarnings('ignore')
# -
df = pd.read_pickle('../data/pickle/df_merged_5.pickle')
# ## Visualisation
## Check for outliers in 'rebap_eur_mwh' column
'''
Quantiles, specifically the first and third quantiles, which correspond to the 25th and 75th percentiles.
Median, the mid-point in the distribution, which also corresponds to the 50th percentile.
Interquartile range (IQR), the width between the third and first quantiles. Expressed mathematically, we have IQR = Q3—Q1.
Min, minimum value in the dataset excluding outliers, which corresponds to Q1–1.5xIQR.
Max, maximum value in the dataset, excluding outliers, which corresponds to Q3+1.5xIQR.
'''
sns.boxplot(x=df['rebap_eur_mwh'])
plt.show()
# We can observe a high price range from almost -2000 to 4000 €/MWh.
# +
median = np.median(df.rebap_eur_mwh)
upper_quartile = np.percentile(df.rebap_eur_mwh, 75)
lower_quartile = np.percentile(df.rebap_eur_mwh, 25)
iqr = upper_quartile - lower_quartile
lower_outlier = lower_quartile - 1.5 * iqr
upper_outlier = upper_quartile + 1.5 * iqr
#upper_whisker = df[df<=upper_quartile+1.5*iqr].max()
#lower_whisker = df[df>=lower_quartile-1.5*iqr].min()
# -
print(upper_quartile)
print(iqr)
print(lower_outlier)
print(upper_outlier)
count_1 = df["rebap_eur_mwh"][(df["rebap_eur_mwh"]>upper_outlier)].count()
count_2 = df["rebap_eur_mwh"][(df["rebap_eur_mwh"]<lower_outlier)].count()
count_3 = (count_1+count_2)/df["rebap_eur_mwh"].count()
# Check for outliers in 'rz_saldo_mwh' column
sns.boxplot(x=df['rz_saldo_mwh'])
plt.show()
print("This data set contains " + str(count_1) + " prices definded as upper outliers (values > " + str(round(upper_outlier,2)) + " €/MWh) and " + str(count_2) +
" observations lower as " + str(round(lower_outlier,2)) + "€/MWh. This represents " + str(round(count_3,3)*100) + "% of all observations.")
#Create a data frame of outliers data frame.
df_outlier = df[(df["rebap_eur_mwh"]<lower_outlier) | (df["rebap_eur_mwh"]>upper_outlier)]
print("We created a data frame of outliers. It has the lenght of", df_outlier.shape[0]+1, "rows.")
# # Ouliers by time granularity
# We analyse the outlier data frame by categorize it time: months, weekdays, business days and daytime
# ## Outliers by months
df_outlier_month = df_outlier.groupby(df_outlier.index.month).count().reset_index()
print(df_outlier_month["rebap_eur_mwh"].mean())
df_outlier_month
# +
fig = px.bar(df_outlier_month, x='dt_start_utc', y='rebap_eur_mwh', text='rebap_eur_mwh')
fig.update_traces(texttemplate='%{text}', textposition='outside', cliponaxis = False)
fig.update_layout(uniformtext_minsize=8, uniformtext_mode='hide')
fig.update_layout(
title="Outliers by Months",
xaxis_title="Month",
yaxis_title="Outliers per Month",
xaxis = dict(
tickmode = 'array',
tickvals = [1, 2, 3, 4, 5, 6],
ticktext=["January","February", "March", "April", "May", "June"]),
legend_title="",
font=dict(
family="Arial",
size=12,
color="Black"
)
)
fig.show()
# -
# We can see a significant drop of outliers in March.
# ## Weekdays
df_outlier_weekday = df_outlier.groupby(df_outlier.index.weekday).count().reset_index()
print(df_outlier_weekday["rebap_eur_mwh"].mean())
df_outlier_weekday.head()
# +
fig = px.bar(df_outlier_weekday, x='dt_start_utc', y='rebap_eur_mwh', text='rebap_eur_mwh')
fig.update_traces(texttemplate='%{text}', textposition='outside', cliponaxis = False)
fig.update_layout(uniformtext_minsize=8, uniformtext_mode='hide')
fig.update_layout(
title="Outliers on Weekdays",
xaxis_title="Day",
yaxis_title="Outliers per Day",
xaxis = dict(
tickmode = 'array',
tickvals = [0, 1, 2, 3, 4, 5, 6],
ticktext=["Monday","Tueday", "Wednesday", "Thursday", "Friday", "Saturday", "Sunday"]),
legend_title="",
font=dict(
family="Arial",
size=12,
color="Black"
)
)
fig.show()
# -
# On both weekdays, Monday and Tuesday, we observe the max outiers counts. On the following business days the number of outliers drops significantly, while the outliers raise on the weekend again.
# ## Outliers by daytime in hours
df_outlier_hour = df_outlier.groupby(df_outlier.index.hour).count().reset_index()
print(df_outlier_hour["rebap_eur_mwh"].mean())
df_outlier_hour.head()
# +
fig = px.bar(df_outlier_hour, x='dt_start_utc', y='rebap_eur_mwh', text='rebap_eur_mwh')
fig.update_traces(texttemplate='%{text}', textposition='outside', cliponaxis = False)
fig.update_layout(uniformtext_minsize=8, uniformtext_mode='hide')
fig.update_layout(
title="Outliers on Weekdays",
xaxis_title="Day",
yaxis_title="Outliers per Day",
#xaxis = dict(
# tickmode = 'array',
# tickvals = [0, 1, 2, 3, 4, 5, 6],
# ticktext=["Monday","Tueday", "Wednesday", "Thursday", "Friday", "Saturday", "Sunday"]),
legend_title="",
font=dict(
family="Arial",
size=12,
color="Black"
)
)
fig.show()
# -
# We can observe four phases in this distribution. A low phase from 19 - 4 o'clock with a local high at 23 o'clock. A high phase from 5 - 7 o'clock with a local max at 6 o'clock. A constant phase of outliers between 23-29 from 10 to 15 o'clock.
# ## Outliers by daytime on business days
df_bhour_count = df_outlier[df_outlier.index.dayofweek < 5].groupby(df_outlier[df_outlier.index.dayofweek < 5].index.hour).count().reset_index()
print(df_bhour_count["rebap_eur_mwh"].mean())
df_bhour_count.head()
# +
fig = px.bar(df_bhour_count, x='dt_start_utc', y='rebap_eur_mwh', text='rebap_eur_mwh')
fig.update_traces(texttemplate='%{text}', textposition='outside', cliponaxis = False)
fig.update_layout(uniformtext_minsize=8, uniformtext_mode='hide')
fig.update_layout(
title="Outliers on Business Days",
xaxis_title="Hours",
yaxis_title="Outliers per hours",
# xaxis = dict(
# tickmode = 'array',
# tickvals = [0, 4, 8, 12, 16, 20],
# ticktext=["00:00","04:00", "08:00", "12:00", "16:00", "20:00"]),
legend_title="",
font=dict(
family="Arial",
size=12,
color="Black"
)
)
fig.show()
# -
# The business days distribution is quite similar to the weekday distribution, but it has a stronger pronounced right skrewed distribution characteristic.
| 2_1_EDA/Outlier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] id="wIIRUuFu0qpM"
# # Data Augmentation
# + [markdown] id="bCYgudwG0qp9"
# ## Import Necessary Modules
# + id="x_NuqdB00qqA"
import tensorflow as tf
from keras.preprocessing.image import ImageDataGenerator
import cv2
import imutils
import matplotlib.pyplot as plt
from os import listdir
import time
# %matplotlib inline
# + id="xyUm8rl20qqL"
# Nicely formatted time string
def hms_string(sec_elapsed):
h = int(sec_elapsed / (60 * 60))
m = int((sec_elapsed % (60 * 60)) / 60)
s = sec_elapsed % 60
return f"{h}:{m}:{round(s,1)}"
# + id="vtSudxzh0qqO"
def augment_data(file_dir, n_generated_samples, save_to_dir):
"""
Arguments:
file_dir: A string representing the directory where images that we want to augment are found.
n_generated_samples: A string representing the number of generated samples using the given image.
save_to_dir: A string representing the directory in which the generated images will be saved.
"""
#from keras.preprocessing.image import ImageDataGenerator
#from os import listdir
data_gen = ImageDataGenerator(rotation_range=10,
width_shift_range=0.1,
height_shift_range=0.1,
shear_range=0.1,
brightness_range=(0.3, 1.0),
horizontal_flip=True,
vertical_flip=True,
fill_mode='nearest'
)
for filename in listdir(file_dir):
# load the image
image = cv2.imread(file_dir + '\\' + filename)
# reshape the image
image = image.reshape((1,)+image.shape)
# prefix of the names for the generated sampels.
save_prefix = 'aug_' + filename[:-4]
# generate 'n_generated_samples' sample images
i=0
for batch in data_gen.flow(x=image, batch_size=1, save_to_dir=save_to_dir,
save_prefix=save_prefix, save_format='jpg'):
i += 1
if i > n_generated_samples:
break
# + [markdown] id="kTiFrva_0qqT"
# Remember that 61% of the data (155 images) are tumorous. And, 39% of the data (98 images) are non-tumorous.<br>
# So, in order to balance the data we can generate 9 new images for every image that belongs to 'no' class and 6 images for every image that belongs the 'yes' class.<br>
# + id="ziz33EXb0qqX" outputId="ddc17e89-02ea-4eb5-85f3-2fcc31cbdf46"
start_time = time.time()
augmented_data_path = 'augmented data/'
# augment data for the examples with label equal to 'yes' representing tumurous examples
augment_data(file_dir=yes_path, n_generated_samples=6, save_to_dir=augmented_data_path+'yes')
# augment data for the examples with label equal to 'no' representing non-tumurous examples
augment_data(file_dir=no_path, n_generated_samples=9, save_to_dir=augmented_data_path+'no')
end_time = time.time()
execution_time = (end_time - start_time)
print(f"Elapsed time: {hms_string(execution_time)}")
# + [markdown] id="KMhcRKu50qqc"
# Let's see how many tumorous and non-tumorous examples after performing data augmentation:
# + id="wwu2z1l10qql"
def data_summary(main_path):
yes_path = main_path+'yes'
no_path = main_path+'no'
# number of files (images) that are in the the folder named 'yes' that represent tumorous (positive) examples
m_pos = len(listdir(yes_path))
# number of files (images) that are in the the folder named 'no' that represent non-tumorous (negative) examples
m_neg = len(listdir(no_path))
# number of all examples
m = (m_pos+m_neg)
pos_prec = (m_pos* 100.0)/ m
neg_prec = (m_neg* 100.0)/ m
print(f"Number of examples: {m}")
print(f"Percentage of positive examples: {pos_prec}%, number of pos examples: {m_pos}")
print(f"Percentage of negative examples: {neg_prec}%, number of neg examples: {m_neg}")
# + id="EtONxoMy0qqn" outputId="29a6e023-b214-46c3-82cf-76d139c0d730"
data_summary(augmented_data_path)
| Data_Augmentation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:GEOP]
# language: python
# name: conda-env-GEOP-py
# ---
# + run_control={"frozen": false, "read_only": false}
import pandas as pd
import numpy as np
# + run_control={"frozen": false, "read_only": false}
df_ = pd.read_csv("SC-List.csv")
# + run_control={"frozen": false, "read_only": false}
df_state = pd.read_csv("statets.csv", sep="\t", header=None)
# -
df_state.head()
# + run_control={"frozen": false, "read_only": false}
df_state.append({0:"DC", 1:"DC"}, ignore_index=True )
# + run_control={"frozen": false, "read_only": false}
df_['state'] = df_.Company.apply(lambda x: x if isinstance(x, float) else x.replace("University of ", ""))
#for state in df_state:
# + run_control={"frozen": false, "read_only": false}
df_.state.unique()
# + run_control={"frozen": false, "read_only": false}
statesdic = {"Arizona":"Arizona",'Lawrence Livermore National Laboratory': 'California',
'NASA Johnson Space Center': 'California',
'Washington':"Washington", 'Columbia University': 'New York',
'UC Berkeley/LBNL': 'California',
'KIPAC Kavli Institute for Particle Astrophysics and Cosmology': 'California',
'Oxford':"UK", 'Universidad de Antofagasta':"Chile", ' California Davis': 'California',
'Universidad Andres Bello':"Chile",
'Laboratoire de Le Accelerateur Lineaire (LAL)':'France', 'LPNHE':'France',
'INAF-Osservatorio Astronomico di Roma':"Italy",
'Universidad Diego Portales':"Chile", 'LCOGT & UC Santa Barbara':"California",
'Jet Propulsion Lab':"California", 'IPAC Caltech':"California",
'Lawrence Berkeley National Laboratory':"California", 'APC':"France", ' Chicago':"Illinois",
'Laboratory for Atmosphere and Space Physics':"Colorado",
'Colorado at Boulder':"Colorado", 'Pennsylvania State University':"Pennsylvania",
'Yale University':"Connecticut", 'LPNHE/IN2P3':"France", 'Georgia Institute of Technology':"Georgia",
'INAF-Osservatorio Astrofisico di Torino & SNS-Scuola Normale':"Italy",
'Observat\xc3\xb3rio Nacional/BGP-LSST/LIneA':"Chile",
'Queens University Belfast':"Iraland",
'Universidad de La Serena':"Chile", 'Universidad de Atacama':"Chile",
'SLAC Nationatl Acceleratory Laboratory':"California", 'LPSC':"Colorado",
'Universidad Metropolitana de Ciencias de la Educacion (UMCE)':"Chile",
'Pontifilecia Universidad Catolica de Chile':"Chile", ' California Irvine':'California',
'<NAME>':"DC", 'Pontifica Universidad Catolica de Chile':"Chile",
'Stanford Univerisity':'California', 'SUNY-SB':"New York", 'Cornell University':"Illinois",
'Stanford University':'California', 'SLAC National Accelerator Laboratory':'California',
'Michigan State University':"Michigan", ' Texas at Austin':"Texas", ' Michigan':"Michigan",
'INAF':"Italy", ' Cambridge':"UK",
'Harvard Smithsonian Center for Astrophysics':"Massachusetts", ' Pennsylvania':"Pennsylvania",
'Argonne National Laboratory':"Illinois", 'Liverpool John Moores University':"UK",
'New York University':"New York",
'INAF-Osservatorio Astronomico di Trieste':"Italy", 'Laboratoire de l':"France"}
# + run_control={"frozen": false, "read_only": false}
df_['state1'] = df_.state.map(statesdic)
# -
df_[['Email', 'state', 'state1']]
# + run_control={"frozen": false, "read_only": false}
emcountry={"it":"Italy", "fr":"France", "uk":"UK", "au":"Australia", "br":"Brasil", "cl":"Chile",
"nz":"New Zealand","fi":"Finland", "cz":"Czech Republic", "dk":"Damnark","de":"Germany",
"jp":"Japan", "ca":"Canada"}
def statebyEmailEnd(email):
if not isinstance(email, float):
if email.split('.')[-1].strip() in emcountry.keys():
return (emcountry[email.split('.')[-1].strip()])
else: return np.nan
else: return np.nan
# + run_control={"frozen": false, "read_only": false}
df_['state2'] = df_.Email.apply(statebyEmailEnd)
# -
df_[['Email', 'state1', 'state2']]
# + run_control={"frozen": false, "read_only": false}
df_['state3'] = [s[0] if not isinstance(s[0], float) else s[1] for s in zip(df_['state1'], df_['state2'])]
# + run_control={"frozen": false, "read_only": false}
pd.Series([em if isinstance(em, float) else '.'.join(em.split("@")[-1].split('.')[-2:])
for em in df_['Email'][np.array([isinstance(v, float)
for v in df_.state3.values ])].values ]).unique()
# + run_control={"frozen": false, "read_only": false}
df_.state2.unique(), df_.state3.unique()
# + run_control={"frozen": false, "read_only": false}
stateByEmail = {'ucdavis.edu':"California",
'uchicago.edu': "Illinois",
'psu.edu': "Pennsylvania",
'stanford.edu':"California",
'umich.edu': "Michigan",
'harvard.edu': "Massachusetts",
'upenn.edu': "Pennsylvania",
'osu.edu':"Ohio",
'berkeley.edu':"California",
'noao.edu':"Arizona",
'umd.edu':"Maryland",
'lcogt.net':"California",
'princeton.edu':"New Jersey",
'asu.edu':"Arizona",
'nasa.gov':np.nan,
'sarahbridle.net':np.nan,
'ttu.edu':"Texas",
'uwyo.edu':"Wyoming",
'tamu.edu':"Texas",
'caltech.edu':"California",
'illinois.edu':"Illinois",
'jhu.edu':"Maryland",
'uci.edu':"California",
'ucsd.edu':"California",
'lsst.org':"Washington",
'yale.edu':"Connecticut",
'cofc.edu':"South Carolina",
'uw.edu':"Washington",
'cas.cn':np.nan,
'purdue.edu':"Indiana",
'cornell.edu':"Illinois",
'arizona.edu':"Arizona",
'bnl.gov':"California",
'nrao.edu':np.nan,
'ohio.edu':"Ohio",
'wayne.edu':"Michigan",
'lsu.edu':"Louisiana",
'washington.edu':"Washington",
'wwu.edu':"Washington",
'amherst.edu':"Massachusetts",
'columbia.edu':"New York",
'uvi.edu':"Virginia",
'dawsonresearch.com':"California",
'aip.de':"Germany",
'nku.edu':"Kentucky",
'llnl.gov':"California",
'brown.edu':"Rhode Island",
'hawaii.edu':"Hawaii",
'me.com':np.nan,
'ucsb.edu':"California",
'fnal.gov':"Illinois",
'carnegiescience.edu':"California",
'northwestern.edu':"Illinois",
'stsci.edu':"DC",
'navy.mil':"DC",
'utexas.edu':"Texas",
'cmu.edu':"Pennsylvania",
'lanl.gov':"California",
'nd.edu':"Delaware",
'rutgers.edu':"New Jersey",
'ou.edu':"Oklahoma",
'udel.edu':"Delaware",
'ung.si':"Slovenia",
'gatech.edu':"Georgia",
'ucolick.org':"California",
'anl.gov':"Illinois",
'villanova.edu':"Pennsylvania",
'colorado.edu':"Colorado",
'adlerplanetarium.org':"Illinois",
'haverford.edu':"Pennsylvania",
'att.net':np.nan,
'rice.edu':"Texas",
'hws.edu':"Hawaii",
'aavso.org':np.nan,
'gsu.edu':"Georgia",
'umkc.edu':"Missouri",
'bell-labs.com':"California",
'psi.edu':"Pennsylvania",
'uni-heidelberg.de':"Germany",
'utdallas.edu':"Texas",
'utah.edu':"Utah",
'jarvis.net':np.nan,
'h-bar.com':np.nan,
'aob.rs':np.nan,
'virginia.edu':"Virginia",
'oswego.edu':"New York",
'uwm.edu':"Masachussets",
'lbl.gov':"California",
'iac.es':"Spain",
'ac.rs':"Serbia",
'albany.edu':"New York",
'mac.com':np.nan,
'usna.edu':"Maryland",
'yahoo.com':np.nan,
'see.com':np.nan,
'ufl.edu':"Florida",
'icrar.org':"Austrtalia",
'gemini.edu':"Chile",
'nau.edu':"Arizona",
'lowell.edu':"Masachussets",
'pitt.edu':"Pennsylvania",
'gmail.com ':np.nan,
'fit.edu':"Florida",
'longwood.edu':"Virginia",
'lehigh.edu':"Pennsylvania",
'normalesup.org':"France",
'missouristate.edu':"Missouri",
'columbusstate.edu':"Ohio",
'byu.edu':"Utah",
'mpg.de':"Germany",
'usra.edu':"Maryland",
'ucr.edu':"California",
'ucla.edu':"California",
'drexel.edu':"Pennsylvania", 'ucsc.edu':"California",
'jhuapl.edu':"Maryland",
'umn.edu':"Minnesota",
'indiana.edu':"Indiana",
'ucf.edu':"Florida",
'unl.edu':"Nebraska",
'sdsu.edu':"California",
'mtu.edu':"Michigan",
'unt.edu':"Texas",
'ciw.edu':"California",
'apsu.edu':"Tennessee",
'uc.edu':"California",
'gov.pl':"Poland",
'lynchburg.edu':"Virginia",
'ornl.gov':"Tennessee",
'vanderbilt.edu':"Tennessee",
'swri.edu':"Coloradu",
'msu.edu':"Massachusetts",
'lco.global':"California",
'ipmu.jp':"Japan",
'dartmouth.edu':"Hampshire",
'nau.edu ':"Arizona",
'nau.edu':"Arizona",
'sunysb.edu':"New York",
'ubc.ca':"California",
'stvincent.edu':"Pennsylvania",
'uh.edu':"Texas",
'duke.edu':"North Carolina",
'uiuc.edu':"Illinois",
'umass.edu':"Massachusetts",
'bu.edu':"Massachusetts",
'aol.com':np.nan,
'Duke.edu':"North Carolina",
'nyu.edu':"New York"}
# 'case.edu'
def sttatetbyemailend(email):
if isinstance(email, float): return np.nan
else:
em = '.'.join(email.split("@")[-1].split('.')[-2:])
if em in stateByEmail.keys():
return stateByEmail[em]
else: return np.nan
# + run_control={"frozen": false, "read_only": false}
df_['state4'] = df_.Email.apply(sttatetbyemailend)
# + run_control={"frozen": false, "read_only": false}
df_.state4.describe()
# + run_control={"frozen": false, "read_only": false}
df_['state5'] = [s[0] if not isinstance(s[0], float) else s[1] for s in zip(df_['state3'], df_['state4'])]
# -
df_.state5.describe()
df_.describe()
# + run_control={"frozen": false, "read_only": false}
df_.to_csv("mydb.csv")
# + run_control={"frozen": false, "read_only": false}
import geopandas as gpd
# + run_control={"frozen": false, "read_only": false}
import pandas as pd
df_ = pd.read_csv("mydb.csv")
# + run_control={"frozen": false, "read_only": false}
df_.shape, df_.dropna(subset=["state5"]).shape
# + run_control={"frozen": false, "read_only": false}
# %pylab inline
# + run_control={"frozen": false, "read_only": false}
countriesshp = gpd.GeoDataFrame.from_file("TM_WORLD_BORDERS_SIMPL-0.3.shp")
ax = countriesshp.plot()
statetsshp = gpd.GeoDataFrame.from_file("cb_2016_us_state_500k.shp")
statetsshp.plot(ax=ax, color='red')
# + run_control={"frozen": false, "read_only": false}
statetsshp.head()
# + run_control={"frozen": false, "read_only": false}
countriesshp["NAME"][["United" in cn for cn in countriesshp["NAME"].values]]
# -
countriesshp.merge(dfgroup, right_index=True, left_on="NAME")["count"].describe()
statetsshp.NAME
statetsshp.merge(dfgroup, right_index=True, left_on="NAME")["count"].describe()
# + run_control={"frozen": false, "read_only": false}
df_.state5[[v == "UK" for v in df_.state5.values]] = "United Kingdom"
# + run_control={"frozen": false, "read_only": false}
df_.state5[[v == "United Kingdom" for v in df_.state5.values]]
df_.state5[[v == "DC" for v in df_.state5.values]]
# -
df_.state5[[v == "DC" for v in df_.state5.values]] = "Maryland"
# + run_control={"frozen": false, "read_only": false}
dfgroup = df_.drop_duplicates("Email").\
groupby("state5").count()[["Last Name"]].\
rename(columns={"Last Name":"count"})
# + run_control={"frozen": false, "read_only": false}
ax = countriesshp.plot(color="w", edgecolor="k", figsize=(20,20))
countriesshp.merge(dfgroup, right_index=True, left_on="NAME").plot(cmap="BuGn", column="count", ax=ax)
statetsshp.merge(dfgroup, right_index=True, left_on="NAME").plot(cmap="Oranges", column="count", ax=ax)
ax.axis('off')
# + run_control={"frozen": false, "read_only": false}
statetsshp.merge(dfgroup, right_index=True, left_on="NAME")[["NAME",'count']]
# -
pd.options.display.max_rows = 999
# + run_control={"frozen": false, "read_only": false}
countriesshp.merge(dfgroup, right_index=True, left_on="NAME", how="left", indicator=True)
# +
# countriesshp.merge?
# -
| vizs/LSSTSCs/LSSTSCaffiliations/Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Objects in boxes
#
# This tutorial includes everything you need to set up IBM Decision Optimization CPLEX Modeling for Python (DOcplex), build a Mathematical Programming model, and get its solution by solving the model on the cloud with IBM ILOG CPLEX Optimizer.
#
# When you finish this tutorial, you'll have a foundational knowledge of _Prescriptive Analytics_.
#
# >This notebook is part of [Prescriptive Analytics for Python](http://ibmdecisionoptimization.github.io/docplex-doc/)
# >
# >It requires either an [installation of CPLEX Optimizers](http://ibmdecisionoptimization.github.io/docplex-doc/getting_started.html) or it can be run on [IBM Watson Studio Cloud](https://www.ibm.com/cloud/watson-studio/>) (Sign up for a [free IBM Cloud account](https://dataplatform.cloud.ibm.com/registration/stepone?context=wdp&apps=all>)
# and you can start using Watson Studio Cloud right away).
#
#
# Table of contents:
#
# * [Describe the business problem](#Describe-the-business-problem)
# * [How decision optimization (prescriptive analytics) can help](#How--decision-optimization-can-help)
# * [Use decision optimization](#Use-decision-optimization)
# * [Step 1: Import the library](#Step-1:-Import-the-library)
# * [Step 2: Model the data](#Step-2:-Model-the-data)
# * [Step 3: Prepare the data](#Step-3:-Prepare-the-data)
# * [Step 4: Set up the prescriptive model](#Step-4:-Set-up-the-prescriptive-model)
# * [Define the decision variables](#Define-the-decision-variables)
# * [Express the business constraints](#Express-the-business-constraints)
# * [Express the objective](#Express-the-objective)
# * [Solve the model](#Solve-the-model)
# * [Step 5: Investigate the solution and run an example analysis](#Step-5:-Investigate-the-solution-and-then-run-an-example-analysis)
# * [Summary](#Summary)
#
# ****
# ## Describe the business problem
#
# * We wish to put $N$ objects which are scattered in the plane, into a row of $N$ boxes.
#
# * Boxes are aligned from left to right (if $i < i'$, box $i$ is to the left of box $i'$) on the $x$ axis.
# * Box $i$ is located at a point $B_i$ of the $(x,y)$ plane and object $j$ is located at $O_j$.
#
#
# * We want to find an arrangement of objects such that:
# * each box contains exactly one object,
# * each object is stored in one box,
# * the total distance from object $j$ to its storage box is minimal.
#
#
# * First, we solve the problem described, and then we add two new constraints and examine how the cost (and solution) changes.
# * From the first solution, we impose that object #1 is assigned to the box immediately to the left of object #2.
# * Then we impose that object #5 is assigned to a box next to the box of object #6.
# ## How decision optimization can help
#
# * Prescriptive analytics (decision optimization) technology recommends actions that are based on desired outcomes. It takes into account specific scenarios, resources, and knowledge of past and current events. With this insight, your organization can make better decisions and have greater control of business outcomes.
#
# * Prescriptive analytics is the next step on the path to insight-based actions. It creates value through synergy with predictive analytics, which analyzes data to predict future outcomes.
#
# * Prescriptive analytics takes that insight to the next level by suggesting the optimal way to handle that future situation. Organizations that can act fast in dynamic conditions and make superior decisions in uncertain environments gain a strong competitive advantage.
# <br/>
#
# <u>With prescriptive analytics, you can:</u>
#
# * Automate the complex decisions and trade-offs to better manage your limited resources.
# * Take advantage of a future opportunity or mitigate a future risk.
# * Proactively update recommendations based on changing events.
# * Meet operational goals, increase customer loyalty, prevent threats and fraud, and optimize business processes.
# ## Use decision optimization
# ### Step 1: Import the library
#
# Run the following code to import the Decision Optimization CPLEX Modeling library. The *DOcplex* library contains the two modeling packages, Mathematical Programming and Constraint Programming, referred to earlier.
import sys
try:
import docplex.mp
except:
raise Exception('Please install docplex. See https://pypi.org/project/docplex/')
# If *CPLEX* is not installed, install CPLEX Community edition.
try:
import cplex
except:
raise Exception('Please install CPLEX. See https://pypi.org/project/cplex/')
# ### Step 2: Model the data
#
# The input data is the number of objects (and boxes) _N_, and their positions in the (x,y) plane.
#
# ### Step 3: Prepare the data
#
# We use Euclidean distance to compute the distance between an object and its assigned box.
#
#
# +
from math import sqrt
N = 15
box_range = range(1, N+1)
obj_range = range(1, N+1)
import random
o_xmax = N*10
o_ymax = 2*N
box_coords = {b: (10*b, 1) for b in box_range}
obj_coords= {1: (140, 6), 2: (146, 8), 3: (132, 14), 4: (53, 28),
5: (146, 4), 6: (137, 13), 7: (95, 12), 8: (68, 9), 9: (102, 18),
10: (116, 8), 11: (19, 29), 12: (89, 15), 13: (141, 4), 14: (29, 4), 15: (4, 28)}
# the distance matrix from box i to object j
# actually we compute the square of distance to keep integer
# this does not change the essence of the problem
distances = {}
for o in obj_range:
for b in box_range:
dx = obj_coords[o][0]-box_coords[b][0]
dy = obj_coords[o][1]-box_coords[b][1]
d2 = dx*dx + dy*dy
distances[b, o] = d2
# -
# ### Step 4: Set up the prescriptive model
from docplex.mp.environment import Environment
env = Environment()
env.print_information()
# #### Create the DOcplex model
# The model contains all the business constraints and defines the objective.
# +
from docplex.mp.model import Model
mdl = Model("boxes")
# -
# #### Define the decision variables
#
# * For each box $i$ ($i$ in $1..N$) and object $j$ ($j$ in $1..N$), we define a binary variable $X_{i,j}$ equal to $1$ if and only if object $j$ is stored in box $i$.
# decision variables is a 2d-matrix
x = mdl.binary_var_matrix(box_range, obj_range, lambda ij: "x_%d_%d" %(ij[0], ij[1]))
# #### Express the business constraints
#
# * The sum of $X_{i,j}$ over both rows and columns must be equal to $1$, resulting in $2\times N$ constraints.
# +
# one object per box
mdl.add_constraints(mdl.sum(x[i,j] for j in obj_range) == 1
for i in box_range)
# one box for each object
mdl.add_constraints(mdl.sum(x[i,j] for i in box_range) == 1
for j in obj_range)
mdl.print_information()
# -
# #### Express the objective
#
# * The objective is to minimize the total distance between each object and its storage box.
# minimize total displacement
mdl.minimize( mdl.sum(distances[i,j] * x[i,j] for i in box_range for j in obj_range) )
# #### Solve the model
#
# +
mdl.print_information()
assert mdl.solve(), "!!! Solve of the model fails"
# +
mdl.report()
d1 = mdl.objective_value
#mdl.print_solution()
def make_solution_vector(x_vars):
sol = [0]* N
for i in box_range:
for j in obj_range:
if x[i,j].solution_value >= 0.5:
sol[i-1] = j
break
return sol
def make_obj_box_dir(sol_vec):
# sol_vec contains an array of objects in box order at slot b-1 we have obj(b)
return { sol_vec[b]: b+1 for b in range(N)}
sol1 = make_solution_vector(x)
print("* solution: {0!s}".format(sol1))
# -
# #### Additional constraint #1
#
# As an additional constraint, we want to impose that object #1 is stored immediately to the left of object #2.
# As a consequence, object #2 cannot be stored in box #1, so we add:
mdl.add_constraint(x[1,2] == 0)
# Now, we must state that for $k \geq 2$ if $x[k,2] == 1$ then $x[k-1,1] == 1$; this is a logical implication that we express by a relational operator:
mdl.add_constraints(x[k-1,1] >= x[k,2]
for k in range(2,N+1))
mdl.print_information()
# Now let's solve again and check that our new constraint is satisfied, that is, object #1 is immediately left to object #2
ok2 = mdl.solve()
assert ok2, "solve failed"
mdl.report()
d2 = mdl.objective_value
sol2 = make_solution_vector(x)
print(" solution #2 ={0!s}".format(sol2))
# The constraint is indeed satisfied, with a higher objective, as expected.
# #### Additional constraint #2
#
# Now, we want to add a second constraint to state that object #5 is stored in a box that is next to the box of object #6, either to the left or right.
#
# In other words, when $x[k,6]$ is equal to $1$, then one of $x[k-1,5]$ and $x[k+1,5]$ is equal to $1$;
# this is again a logical implication, with an OR in the right side.
#
# We have to handle the case of extremities with care.
# +
# forall k in 2..N-1 then we can use the sum on the right hand side
mdl.add_constraints(x[k,6] <= x[k-1,5] + x[k+1,5]
for k in range(2,N))
# if 6 is in box 1 then 5 must be in 2
mdl.add_constraint(x[1,6] <= x[2,5])
# if 6 is last, then 5 must be before last
mdl.add_constraint(x[N,6] <= x[N-1,5])
# we solve again
ok3 = mdl.solve()
assert ok3, "solve failed"
mdl.report()
d3 = mdl.objective_value
sol3 = make_solution_vector(x)
print(" solution #3 ={0!s}".format(sol3))
# -
# As expected, the constraint is satisfied; objects #5 and #6 are next to each other.
# Predictably, the objective is higher.
#
# ### Step 5: Investigate the solution and then run an example analysis
#
# Present the solution as a vector of object indices, sorted by box indices.
# We use maptplotlib to display the assignment of objects to boxes.
#
# +
import matplotlib.pyplot as plt
from pylab import rcParams
# %matplotlib inline
rcParams['figure.figsize'] = 12, 6
def display_solution(sol):
obj_boxes = make_obj_box_dir(sol)
xs = []
ys = []
for o in obj_range:
b = obj_boxes[o]
box_x = box_coords[b][0]
box_y = box_coords[b][1]
obj_x = obj_coords[o][0]
obj_y = obj_coords[o][1]
plt.text(obj_x, obj_y, str(o), bbox=dict(facecolor='red', alpha=0.5))
plt.plot([obj_x, box_x], [obj_y, box_y])
# -
# The first solution shows no segments crossing, which is to be expected.
display_solution(sol1)
# The second solution, by enforcing that object #1 must be to the left of object #2, introduces crossings.
display_solution(sol2)
display_solution(sol3)
# +
def display(myDict, title):
if True: #env.has_matplotlib:
N = len(myDict)
labels = myDict.keys()
values= myDict.values()
try: # Python 2
ind = xrange(N) # the x locations for the groups
except: # Python 3
ind = range(N)
width = 0.2 # the width of the bars
fig, ax = plt.subplots()
rects1 = ax.bar(ind, values, width, color='g')
ax.set_title(title)
ax.set_xticks([ind[i]+width/2 for i in ind])
ax.set_xticklabels( labels )
#ax.legend( (rects1[0]), (title) )
plt.show()
else:
print("warning: no display")
from collections import OrderedDict
dists = OrderedDict()
dists["d1"]= d1 -8000
dists["d2"] = d2 - 8000
dists["d3"] = d3 - 8000
print(dists)
display(dists, "evolution of distance objective")
# -
# ## Summary
#
# You learned how to set up and use IBM Decision Optimization CPLEX Modeling for Python to formulate a Mathematical Programming model and solve it with CPLEX.
# ## References
# * [CPLEX Modeling for Python documentation](http://ibmdecisionoptimization.github.io/docplex-doc/)
# * [Decision Optimization on Cloud](https://developer.ibm.com/docloud/)
# * Need help with DOcplex or to report a bug? Please go [here](https://stackoverflow.com/questions/tagged/docplex).
# * Contact us at <EMAIL>.
# Copyright © 2017-2019 IBM. IPLA licensed Sample Materials.
| examples/mp/jupyter/boxes.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # The Display Module
#
# The :mod:`diogenes.display` module provides tools for summarizing/exploring data and the performance of trained classifiers.
# ## Exploring data
#
# Display provides a number of tools for examining data before they have been fit to classifiers.
#
# We'll start by pulling and organizing the wine dataset. We read a CSV from The Internet using :func:`diogenes.read.read.open_csv_url`.
# +
# %matplotlib inline
import diogenes
import numpy as np
wine_data = diogenes.read.open_csv_url('http://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-white.csv',
delimiter=';')
# -
# We will then separate labels from features using :func:`diogenes.utils.remove_cols`.
labels = wine_data['quality']
M = diogenes.utils.remove_cols(wine_data, 'quality')
# Finally, we alter labels to make this into a binary classification problem. (At this point, all Diogenes features are available for binary classification, but other kinds of ML have more limited support).
labels = labels < np.average(labels)
# We can look at our summary statistics with :func:`diogenes.display.display.describe_cols`. Like most functions in Diogenes, `describe_cols` produces a Numpy [structured array](http://docs.scipy.org/doc/numpy/user/basics.rec.html).
summary_stats = diogenes.display.describe_cols(M)
print summary_stats.dtype
print summary_stats
# It's a bit confusing to figure out which numbers go to which statistics using default structured array printing, so we provide :func:`diogenes.display.display.pprint_sa` to make it more readable when we print small structured arrays.
diogenes.display.pprint_sa(summary_stats)
# Similarly, we have a number of tools that visualize data. They all return figures, in case the user wants to save them or plot them later.
figure = diogenes.display.plot_correlation_matrix(M)
figure = diogenes.display.plot_correlation_scatter_plot(M)
# There are also a number of tools for exploring the distribution of data in a single column (ie a 1-dimensional Numpy array)
# +
chlorides = M['chlorides']
figure = diogenes.display.plot_box_plot(chlorides)
figure = diogenes.display.plot_kernel_density(chlorides)
figure = diogenes.display.plot_simple_histogram(chlorides)
# -
diogenes.display.pprint_sa(diogenes.display.crosstab(np.round(chlorides, 1), labels))
# ## Examining classifier performance.
#
# First, we will arrange and execute a quick grid_search experiment with :class:`diogenes.grid_search.experiment.Experiment`. This will run Random Forest on our data with a number of different hyper-parameters and a number of different train/test splits. See documentation for grid_search for more detail.
from sklearn.ensemble import RandomForestClassifier
clfs = [{'clf': RandomForestClassifier, 'n_estimators': [10,50],
'max_features': ['sqrt','log2'], 'random_state': [0]}]
exp = diogenes.grid_search.experiment.Experiment(M, labels, clfs=clfs)
_ = exp.run()
# Now, we will extract a single run, which gives us a single fitted classifier and a single set of test data.
run = exp.trials[0].runs[0][0]
fitted_classifier = run.clf
# Sadly, SKLearn doesn't like structured arrays, so we have to convert to the other kind of array
M_test = diogenes.utils.cast_np_sa_to_nd(M[run.test_indices])
labels_test = labels[run.test_indices]
scores = fitted_classifier.predict_proba(M_test)[:,1]
# We can use our fitted classifier and test data to make an ROC curve or a precision-recall curve showing us how well the classifier performs.
roc_fig = diogenes.display.plot_roc(labels_test, scores)
prec_recall_fig = diogenes.display.plot_prec_recall(labels_test, scores)
# For classifiers that offer feature importances, we provide a convenience method to get the top `n` features.
top_features = diogenes.display.get_top_features(fitted_classifier, M=M)
# For random forest classifiers, we also provide a function to examine consecutive occurence of features in decision trees. see :func:`diogenes.display.display.feature_pairs_in_rf` for more detail.
results = diogenes.display.feature_pairs_in_rf(fitted_classifier, n=3)
# ## Making PDF Reports
#
# Finally, diogenes.display provides a simple way to make PDF reports using :class:`diogenes.display.display.Report`.
#
# * Add headings with :meth:`diogenes.display.display.Report.add_heading`
# * Add text blocks with :meth:`diogenes.display.display.Report.add_text`
# * Add tables with :meth:`diogenes.display.display.Report.add_table`
# * Add figures with :meth:`diogenes.display.display.Report.add_fig`
# * Build the report with :meth:`diogenes.display.display.Report.to_pdf`
report = diogenes.display.Report(report_path='display_sample_report.pdf')
report.add_heading('My Great Report About RF', level=1)
report.add_text('I did an experiment with the wine data set '
'(http://archive.ics.uci.edu/ml/machine-learning-databases/wine-quality/winequality-white.csv)')
report.add_heading('Top Features', level=2)
report.add_table(top_features)
report.add_heading('ROC Plot', level=2)
report.add_fig(roc_fig)
full_report_path = report.to_pdf(verbose=False)
# Here's the result:
from IPython.display import HTML
HTML('<iframe src=display_sample_report.pdf width=700 height=350></iframe>')
| doc/notebooks/.ipynb_checkpoints/display-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] pycharm={"name": "#%% md\n"}
# # Broadcast variables
#
# In spark, broadcast variables are read-only shared variables that are cached and available on all nodes in a
# cluster rather than shipping a copy of it with tasks. Instead of sending this data along with every task,
# Spark distributes broadcast variables to the workers using efficient broadcast algorithms to reduce
# communication costs. It means all executor in the same worker can share the same broadcast variable
#
# Spark actions are executed through a set of stages, separated by distributed “shuffle” operations. Spark automatically broadcasts the common data needed by tasks within each stage. The data broadcasted this way is cached in serialized form and deserialized before running each task. This means that explicitly creating broadcast variables is only useful when tasks across multiple stages need the same data or when caching the data in deserialized form is important.
#
# + pycharm={"name": "#%%\n"}
from pyspark.sql import SparkSession
import os
local=True
if local:
spark = SparkSession.builder\
.master("local[4]")\
.appName("BroadcastVariable")\
.config("spark.executor.memory", "2g")\
.getOrCreate()
else:
spark = SparkSession.builder\
.master("k8s://https://kubernetes.default.svc:443")\
.appName("BroadcastVariable")\
.config("spark.kubernetes.container.image", "inseefrlab/jupyter-datascience:master")\
.config("spark.kubernetes.authenticate.driver.serviceAccountName", os.environ['KUBERNETES_SERVICE_ACCOUNT'])\
.config("spark.executor.instances", "4")\
.config("spark.executor.memory","2g")\
.config("spark.kubernetes.namespace", os.environ['KUBERNETES_NAMESPACE'])\
.getOrCreate()
# make the large dataframe show pretty
spark.conf.set("spark.sql.repl.eagerEval.enabled",True)
# + [markdown] pycharm={"name": "#%% md\n"}
# ## Create a broadcast variable
#
# We can create a broadcast variables from any variable, in our case, it's called **states_map**. We use the following command **broadcast_states = spark.sparkContext.broadcast(states_map)** to create a broadcast var called **broadcast_states**. The **broadcast_states** is a wrapper around **states_map**, and its value can be accessed by calling the value method.
#
# Note that broadcast variables are not sent to workers when we created the broadcast var **by calling sc.broadcast()**, the broadcast var will be sent to executors **when they are first used**.
#
# + pycharm={"name": "#%%\n"}
states_map = {"NY": "New York", "CA": "California", "FL": "Florida"}
broadcast_states = spark.sparkContext.broadcast(states_map)
data = [("James", "Smith", "USA", "CA"),
("Michael", "Rose", "USA", "NY"),
("Robert", "Williams", "USA", "CA"),
("Maria", "Jones", "USA", "FL")
]
print("source data: \n {}".format(data))
# + [markdown] pycharm={"name": "#%%\n"}
# ## Use a broadcast variables
#
# After the broadcast variable (i.e. broadcast_states) is created, pay attention to two points:
# 1. Always use the broadcast variable (i.e. broadcast_states) instead of the value (i.e. states_map) in any functions run on the cluster so that **states_map** is shipped to the nodes only once.
# 2. The value **states_map** should not be modified after it is broadcast in order to ensure that all nodes get the same value of the broadcast variable (e.g. if the variable is shipped to a new node later).
#
# +
rdd = spark.sparkContext.parallelize(data)
# get the broadcast state map
states_map = broadcast_states.value
result = rdd.map(lambda x: (x[0], x[1], x[2], states_map[x[3]])).collect()
print("Exp1: after the rdd map on broadcast var ")
print(result)
# -
# Use a broadcast variables in dataframe
# +
columns = ["firstname", "lastname", "country", "state"]
df = spark.createDataFrame(data, schema=columns)
print("Source data frame")
df.show()
# get the broadcast state map
states_map = broadcast_states.value
df1 = df.rdd.map(lambda x: (x[0], x[1], x[2], states_map[x[3]])).toDF(columns)
print("Exp2: after the data frame map on broadcast var ")
df1.show()
# Once the variable is broadcast, we can use it in any dataframe operation which is impossible for local variable
# Because executors does not have access on the local variables defines on spark drivers
local_states_map = ["NY"]
try:
df2 = df.where(df.state.isin(local_states_map))
df2.show()
except:
print("Can't use local variables in dataframe operations")
# isin takes a list, so we need to get the keys of the dict and return it to a list, and I only take the first
# element of the list.
keys = list(states_map.keys())[0:1]
df3 = df.where(df["state"].isin(keys))
df3.show()
# -
| notebooks/pysparkbasics/L01_PySpark_Intro/S04_BroadcastVariable.ipynb |
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .cpp
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: C++14
// language: C++14
// name: xcpp14
// ---
// [](https://lab.mlpack.org/v2/gh/mlpack/examples/master?urlpath=lab%2Ftree%2Fcontact_tracing_clustering_with_dbscan%2F.ipynb)
/**
* @file contact-tracing-dbscan-cpp.ipynb
*
* A simple contact tracing method using DBSCAN.
*
* Once a person is tested positive for the virus,
* it is very important to identify others who may
* have been infected by the diagnosed patients.
* To identify the infected people, a process called
* contact tracing is often used. In this example, we
* applied DBSCAN to perform pseudo location-based
* contact tracing using GPS.
*/
!wget -q https://lab.mlpack.org/data/contact-tracing.csv
// +
#include <mlpack/xeus-cling.hpp>
#include <mlpack/core.hpp>
#include <mlpack/methods/dbscan/dbscan.hpp>
#include <sstream>
// +
// Header files to create and show the plot.
#define WITHOUT_NUMPY 1
#include "matplotlibcpp.h"
#include "xwidgets/ximage.hpp"
#include "../utils/plot3d.hpp"
namespace plt = matplotlibcpp;
// -
using namespace mlpack;
using namespace mlpack::dbscan;
using namespace mlpack::data;
// Load the pseudo location-based dataset for the contact tracing.
// The dataset has 4 columns: timestamp, latitude, longitude, id.
arma::mat input;
DatasetInfo info;
data::Load("contact-tracing.csv", input, info);
// Print the first ten columns of the input data.
std::cout << "timestamp\t"
<< "latitude\t"
<< "longitude\t"
<< "id\t" << std::endl;
std::cout << input.cols(0, 10).t() << std::endl;
// Helper function to generate the data for the 3D plot.
void Data3DPlot(std::stringstream& xData,
std::stringstream& yData,
std::stringstream& time,
std::stringstream& label,
const std::vector<int>& filter)
{
xData.clear();
yData.clear();
time.clear();
label.clear();
for (size_t i = 0; i < info.NumMappings(3); ++i)
{
if (filter.size() != 0 &&
std::find(filter.begin(), filter.end(), i) == filter.end())
continue;
// Get the indices for the current label.
arma::mat dataset = input.cols(arma::find(input.row(3) == (double) i));
// Get the data for the indices.
std::vector<double> t = arma::conv_to<std::vector<double>>::from(dataset.row(0));
std::vector<double> x = arma::conv_to<std::vector<double>>::from(dataset.row(1));
std::vector<double> y = arma::conv_to<std::vector<double>>::from(dataset.row(2));
// Build the strings for the plot.
label << info.UnmapString(i, 3);
for (size_t j = 0; j < t.size(); ++j)
{
xData << x[j] << ";";
yData << y[j] << ";";
// Scale time to make the plot easier to read.
time << t[j] / 1000 << ";";
}
// Prepare for the next row.
xData << "\n";
yData << "\n";
time << "\n";
label << "\n";
}
}
// +
// Plot ids with their latitudes and longitudes across the x-axis and y-axis respectively.
std::stringstream xData, yData, time, label;
std::vector<int> filter;
// Uncomment the lines below to filter for id 0 and 3.
// filter.push_back(0);
// filter.push_back(3);
Data3DPlot(xData, yData, time, label, filter);
Plot3D(xData.str(),
yData.str(),
time.str(),
label.str(),
"x",
"y",
"time",
2, // Mode: 0 = line, 1 = scatter, 2 = line + scatter.
"output.png",
10, // Plot width.
10); // Plot height.
auto im = xw::image_from_file("output.png").finalize();
im
// -
// Plotting all ids can be confusing, so it might be useful to only plot certain ids.
// See the comment above to filter and plot certain ids.
// +
// Generate clusters, and identify the infections by filtering the data in the clusters.
// Radial distance of 6 feet in kilometers.
const double epsilon = 0.0018288;
// Perform Density-Based Spatial Clustering of Applications with Noise
// (DBSCAN).
//
// For more information checkout https://mlpack.org/doc/mlpack-git/doxygen/classmlpack_1_1dbscan_1_1DBSCAN.html
// or uncomment the line below.
// ?DBSCAN<>
DBSCAN<> model(epsilon, 2 /* Minimum number of points for each cluster. */);
// We only use the latitude and longitude attribute.
const arma::mat points = input.submat(
1, 0, input.n_rows - 2 , input.n_cols - 1);
// Perform clustering using DBSCAN, an return the number of clusters.
arma::Row<size_t> assignments;
const size_t numCluster = model.Cluster(points, assignments);
// -
// The model was able to generate 29 clusters, out of which cluster
// 0 to cluster 29 represents data points with neighboring nodes.
std::cout << "Number of clusters: " << numCluster << std::endl;
// +
// Plot cluster with their latitudes and longitudes across the x-axis and y-axis respectively.
plt::figure_size(800, 800);
for (size_t i = 0; i < numCluster; ++i)
{
// Get the indices for the current label.
arma::mat dataset = input.cols(arma::find(assignments == i));
// Get the data for the indices.
std::vector<double> x = arma::conv_to<std::vector<double>>::from(dataset.row(1));
std::vector<double> y = arma::conv_to<std::vector<double>>::from(dataset.row(2));
// Set the label for the legend.
std::map<std::string, std::string> m;
m.insert(std::pair<std::string, std::string>("label", std::to_string(i)));
plt::scatter(x, y, 10, m);
}
plt::xlabel("X");
plt::ylabel("y");
plt::title("ids with their latitudes and longitudes");
plt::legend();
plt::save("./plot.png");
auto im = xw::image_from_file("plot.png").finalize();
im
// -
// Check for people who had been in contact with the infected patient.
void PrintInfected(const std::string& infected /* Infected id e.g. Judy. */,
DatasetInfo& info /* The dataset info object to map between ids and names. */,
const arma::Row<size_t>& assignments /* The generated cluster. */,
const size_t numCluster /* The number of found cluster. */)
{
// Get id from name.
double infectedId = info.MapString<double>(infected, 3);
// Get infected clusters.
arma::Mat<size_t> assignmentsTemp = assignments;
arma::Mat<size_t> cluster = assignmentsTemp.cols(
arma::find(input.row(3) == infectedId));
// Filter out noise cluster.
cluster = cluster.cols(arma::find(cluster <= numCluster));
std::cout << "Infected: " << infected << std::endl;
// Find all names that are in the same infected cluster.
for (size_t c = 0; c < cluster.n_elem; ++c)
{
arma::mat infectedIdsFromCluster = input.cols(
arma::find(assignments == cluster(c)));
if (infectedIdsFromCluster.n_cols <= 0)
std::cout << "No people in the same cluster." << std::endl;
else
std::cout << "Maybe infected others in the cluster: ";
for (size_t n = 0, g = 0; n < infectedIdsFromCluster.n_cols; ++n)
{
size_t id = infectedIdsFromCluster.col(n)(3);
// Skip the name if it's the same as the infected person.
if (info.UnmapString(id, 3) == infected)
continue;
if (g == 0)
std::cout << info.UnmapString(id, 3);
else
std::cout << "," << info.UnmapString(id, 3);
g++;
}
std::cout << std::endl;
}
}
// Check for the people who might be potentially infected from the patient.
PrintInfected("Heidi", info, assignments, numCluster)
// +
// Plot the data for Carol, Frank and Grace, to check the contact over time.
std::vector<int> filterHeidiDavidJudy;
filterHeidiDavidJudy.push_back((int) info.MapString<double>("Heidi", 3));
filterHeidiDavidJudy.push_back((int) info.MapString<double>("David", 3));
filterHeidiDavidJudy.push_back((int) info.MapString<double>("Judy", 3));
std::stringstream xData, yData, time, label;
Data3DPlot(xData, yData, time, label, filterHeidiDavidJudy);
Plot3D(xData.str(),
yData.str(),
time.str(),
label.str(),
"x",
"y",
"time",
2, // Mode: 0 = line, 1 = scatter, 2 = line + scatter.
"contact-heidi-david-judy.png",
10, // Plot width.
10); // Plot height.
auto im = xw::image_from_file("contact-heidi-david-judy.png").finalize();
im
// -
// Check for the people who might be potentially infected from the patient.
PrintInfected("Alice", info, assignments, numCluster)
// +
// Plot the data for Alice and Judy, to check the contact over time.
std::vector<int> filterAliceJudy;
filterAliceJudy.push_back((int) info.MapString<double>("Alice", 3));
filterAliceJudy.push_back((int) info.MapString<double>("Judy", 3));
std::stringstream xData, yData, time, label;
Data3DPlot(xData, yData, time, label, filterAliceJudy);
Plot3D(xData.str(),
yData.str(),
time.str(),
label.str(),
"x",
"y",
"time",
2, // Mode: 0 = line, 1 = scatter, 2 = line + scatter.
"contact-alice-judy.png",
10, // Plot width.
10); // Plot height.
auto im = xw::image_from_file("contact-alice-judy.png").finalize();
im
// -
// Check for the people who might be potentially infected from the patient.
PrintInfected("David", info, assignments, numCluster)
// Check for the people who might be potentially infected from the patient.
PrintInfected("Judy", info, assignments, numCluster)
// Check for the people who might be potentially infected from the patient.
PrintInfected("Carol", info, assignments, numCluster)
| contact_tracing_clustering_with_dbscan/contact-tracing-dbscan-cpp.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3.8.5 64-bit
# metadata:
# interpreter:
# hash: 31f2aee4e71d21fbe5cf8b01ff0e069b9275f58929596ceb00d14d90e3e16cd6
# name: Python 3.8.5 64-bit
# ---
PATH = '../input/cassava-leaf-disease-classification'
import os
import pandas as pd
import matplotlib.pyplot as plt
from sklearn import model_selection
import matplotlib.image as mpimg
# +
train_set = pd.read_csv(f'{PATH}/train.csv')
train_set = pd.read_csv(f'{PATH}/train.csv')
train_set.head(2)
# -
json_mapping = {"0": "Cassava Bacterial Blight (CBB)", "1": "Cassava Brown Streak Disease (CBSD)", "2": "Cassava Green Mottle (CGM)", "3": "Cassava Mosaic Disease (CMD)", "4": "Healthy"}
# +
images_array = [x for x in os.listdir(f'{PATH}/train_images')]
def imshow(position: int):
image_id, category = train_set.iloc[position]
label = json_mapping.get(f"{category}")
image_path = f'{PATH}/train_images/{image_id}'
img = mpimg.imread(image_path)
plt.xlabel(label)
plt.imshow(img)
imshow(2)
# -
# the dataset is skewed towards label 3
train_set.label.value_counts()
# +
kFold = model_selection.StratifiedKFold(n_splits=5)
for f, (t_, v_) in enumerate(kFold.split(X=train_set, y=train_set.label)):
train_set.loc[v_, 'kFold'] = f
train_set.reset_index(drop=True, inplace=True)
# -
# !ls
train_csv = pd.read_csv(f"{PATH}/train.csv")
# adding a column for image location
train_csv['path'] = train_csv['image_id'].map(
lambda x: f"{PATH}/'train_images/{x}")
# shuffling and reset index
print(train_csv.columns)
train_csv.drop('image_id', axis=1,inplace=True)
train_csv = train_csv.sample(frac=1).reset_index(drop=True)
train_csv.head()
print(train_csv.columns)
print(train_csv.iloc[0])
# +
kFold = model_selection.StratifiedKFold(n_splits=5, shuffle=True, random_state=42)
for f, (t_, v_) in enumerate(kFold.split(X=train_csv, y=train_csv.label)):
train_csv.loc[v_, 'kFold'] = f
# -
# +
train_cov = train_csv.groupby('label').apply(
lambda x: x.sample(frac=0.1)).reset_index(drop=True)
df1 = train_cov[train_csv['kFold'] == 4]
df2 = train_cov[train_csv['kFold'] != 4]
df1 = df1.sample(frac=0.1).reset_index(drop=True)
# -
df1.label.value_counts()
# df1.sample(frac=1).reset_index(drop=True)
df1.iloc[0, 1]
# +
from PIL import Image
from PIL import ImageFile
import numpy as np
import torch
image_path = df1.loc[5]['path']
img = "../input/cassava-leaf-disease-classification/train_images/6103.jpg"
image = Image.open(img)
image = np.array(image)
image = torch.tensor(image)
print(2, image)
# -
# !ls "../input/cassava-leaf-disease-classification/'train_images"
# !pip3 install torch
y = train_set['label']
y[4]
import datetime, os
today = str(datetime.date.today())
print(today)
# +
PATHO = f'../output/checkpoints/{today}'
if not os.path.exists(PATHO):
os.mkdir(PATHO)
# -
| notebooks/exploration.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Microblaze Python Libraries
#
# In addition to using the `pynqmb` libraries from C it is also possible to create Python wrappers for the libraries directly. PYNQ provides the `MicroblazeLibrary` class for this purpose.
#
# The `MicroblazeLibrary` class takes a list of libraries as a construction parameter which should be the names of the header files desired without the `.h` file extension. All of the constants and functions will then become members of the instance of the class.
#
# For this example we are going to interact with the Grove ADC device attached to a Pmod-Grove Adapter. We are going to want the `i2c` library for interacting with the device and the `pmod_grove` library to find the pins we want to connect to.
# +
from pynq.overlays.base import BaseOverlay
from pynq.lib import MicroblazeLibrary
base = BaseOverlay('base.bit')
lib = MicroblazeLibrary(base.PMODA, ['i2c', 'pmod_grove'])
# -
# We can now inspect the lib to see all of the functions we can call and constants we have access.
dir(lib)
# Next we need to open our I2C device using the `i2c_open` function. This will return us an `i2c` object we can use for interacting with the bus.
device = lib.i2c_open(lib.PMOD_G4_B, lib.PMOD_G4_A)
# We can check the functions we can call by using `dir` again.
dir(device)
# The Grove ADC responds to address 0x50 and to read from it we need to write the register we want (0 for the result) and then read the two bytes back.
# +
buf = bytearray(2)
buf[0] = 0
device.write(0x50, buf, 1)
device.read(0x50, buf, 2)
((buf[0] & 0x0F) << 8) | buf[1]
# -
# This same approach can be used for all of the other libraries either included with PYNQ or from other sources.
| boards/Pynq-Z1/base/notebooks/microblaze/microblaze_python_libraries.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Supermarket Staple Prices
# ## Imports, Zipcodes, and Staples List
# ### Need to install selenium and webdriver_manager
# #### Currently have:
#
# 1. Giant
# 2. <NAME>
# 3. Safeway
# 4. Aldi
# 5. Target
# 6. Whole Foods Market
# 7. Lidl
# 8. Food Lion
# 9. Amazon
#
# #### No online presence?:
#
# 1. Publix
# 2. Trader Joe's
# 3. Costco
#
# + code_folding=[0]
# imports
# generic imports
import numpy as np
import re
import time
import pandas as pd
import matplotlib.pyplot as plt
# selenium imports
import selenium
from selenium import webdriver
from selenium.webdriver.common.keys import Keys
from webdriver_manager.chrome import ChromeDriverManager
from selenium.webdriver.common.by import By
from selenium.webdriver.support.ui import WebDriverWait
from selenium.webdriver.support import expected_conditions as EC
from selenium.common.exceptions import ElementClickInterceptedException
from selenium.common.exceptions import NoSuchElementException
# + code_folding=[0]
# importing helper functions from .py file!!!
import food_helper
# + code_folding=[0]
# Contains Arlington Zipcodes
zipcodes = np.array([20330, 22201, 22202, 22203, 22204, 22205, 22206, 22207, 22209, 22210, 22211, 22213, 22214,
22215, 22216, 22217, 22219, 22225, 22226, 22227, 22230, 22240, 22241, 22242, 22243, 22244,
22245, 22246, 22350])
# + code_folding=[0]
# reload food_helper file
import importlib
importlib.reload(food_helper)
# -
# ### NOT SURE WHAT IS GOING ON WITH GIANT + HARRIS TEETER??????
# ## GIANT
# + code_folding=[0]
# Giant Baskets
giant_foods = ["Giant Whole Vitamin D Milk", "Giant White Eggs Grade A", "Giant Bagels Plain Pre-Sliced 6 ct", "Giant White Bread",
"Giant Pasta Penne Rigate", "Gala Apple", "Ground Turkey 85%", "Giant Deli Ham Honey Ham Thin Sliced",
"Giant Sweet Cream Butter Sticks 4", "Giant Potatoes Russet", "Navel Orange", "Giant Tomatoes",
"Ground Beef", "Giant 4 Cheese Mexican Blend", "Giant Yogurt", "Giant Cereal Shredded Wheat"]
# Napolitos?? - finish this!
giant_hispanic_foods = ["Avocado", "Giant Beets Whole", "Black Beans", "Giant 4 Cheese Mexican Blend", "Giant Corn Tortillas",
"Green Chiles", "Jalapeno Peppers", "Garbanzo Beans", "Masa",
"Giant Mixed Fruit Large", "Bell Peppers Green", "Pinto Beans", "Giant White Rice Long Grain",
"Pace Chunky Salsa", "Yellow Squash", "Tomatillos", "Tomatoes On The Vine"]
giant_east_af_foods = ["Curry Powder", "Dried Beans", "Premium Dates",
"Garbanzo Beans", "Giant Spaghetti Pasta",
"Giant Tomato Sauce", "Tuna"]
# had to drop fava beana + health bearley...
giant_east_eu_foods = ["Ground Beef", "Giant Beets", "Red Cabbage",
"Giant Lamb", "Giant Sliced Mushrooms", "Giant Pork",
"Giant Potatoes Russet", "Giant Wheat Flour"]
# had to drop health barley
# giant_indig_foods = []
giant_se_asia_foods = ["Baby Corn", "Coconut Milk", "Edamame",
"Eggplant", "Fish Sauce", "Vermicelli",
"Oyster Sauce", "Rice Noodles",
"Jasmine Rice", "Yellow Squash", "Tofu"]
# dropped curry paste, tapioca flour, bamboo shoots
# maybe hae this check for cheap fish sauce...
giant_west_af_foods = ["Black Eye Peas", "Pinto Beans", "Cassava", "Giant Chicken Whole",
"Corn Flour", "Maggi", "Plantain Ripe", "Giant Potatoes Russet",
"Giant Tomato Paste", "Potatoes Yams"]
# + code_folding=[]
# add condition to skip or fill in NA for cultural items not found in some locations!
# + code_folding=[]
# Universal Basket for Giant
giant_df, giant_zips, giant_locs = food_helper.giant(giant_foods, zipcodes)
giant_df.head()
# + code_folding=[]
# Latinx Basket for Giant
giant_hispanic_df = food_helper.giant(giant_hispanic_foods, giant_zips, standard = False, has_zipcodes = True, locs = giant_locs)
giant_hispanic_df.head()
# + code_folding=[0]
# East African Basket for Giant
giant_east_af_df = food_helper.giant(giant_east_af_foods, giant_zips, standard = False, has_zipcodes = True, locs = giant_locs)
giant_east_af_df.head()
# + code_folding=[0]
# Eastern European Basket for Giant
giant_east_eu_df = food_helper.giant(giant_east_eu_foods, giant_zips, standard = False, has_zipcodes = True, locs = giant_locs)
giant_east_eu_df.head()
# + code_folding=[0]
# Southeast Asian Basket for Giant
giant_se_asia_df = food_helper.giant(giant_se_asia_foods, giant_zips, standard = False, has_zipcodes = True, locs = giant_locs)
giant_se_asia_df.head()
# + code_folding=[0]
# West African Basket for Giant
giant_west_af_df = food_helper.giant(giant_west_af_foods, giant_zips, standard = False, has_zipcodes = True, locs = giant_locs)
giant_west_af_df.head()
# + code_folding=[0]
# Giant to csv
# giant_east_af_df.to_csv('Giant_East_Af_Food.csv')
# giant_east_eu_df.to_csv('Giant_East_Eu_Food.csv')
# giant_se_asia_df.to_csv('Giant_SE_Asia_Food.csv')
# giant_west_af_df.to_csv('Giant_West_Af_Food.csv')
# -
# ## <NAME>
# + code_folding=[0]
# Harris Teeter Basket
ht_foods = ["Harris Teeter 3.25% Milk Fat Whole Milk", "Harris Teeter Grade A Large White Eggs", "Thomas Plain Bagels", "Harris Teeter Bread Old Fashioned",
"<NAME> Penne", "Fresh Gala Apple", "Ground Turkey 94%", "<NAME>",
"Harris Teeter Butter Spread", "Russet Potatoes", "Navel Oranges", "Harris Teeter Tomato Sauce",
"Harris Teeter Ground Beef Chuck", "Harris Teeter Shredded Cheese", "Harris Teeter Nonfat Plain Yogurt", "<NAME> Cereal"]
ht_hispanic_foods = ["Hass Avocado", "Fresh Beets", "Harris Teeter Black Beans", "Shredded Mexican Cheese Blend", "Harris Teeter Corn Tortillas",
"Harris Teeter Green Chiles", "Jalapeno Peppers", "Harris Teeter Garbanzo Beans",
"Mixed Fruit", "Bell Peppers Green", "Harris Teeter Pinto Beans", "Harris Teeter Long Grain White Rice",
"Harris Teeter Salsa", "Zucchini Squash", "Tomatillos", "Roma Tomato"]
ht_east_af_foods = ["Pearled Barley", "Curry Powder", "Dried Beans", "Sunsweet Dates",
"Harris Teeter Garbanzo Beans", "Lentil Beans", "Harris Teeter Spaghetti",
"Harris Teeter Tomato Sauce", "Tuna"]
ht_east_eu_foods = ["Pearled Barley", "Harris Teeter Ground Beef Chuck", "Fresh Beets",
"Buckwheat", "Fresh Cabbage Red", "Lamb",
"Harris Teeter Whole White Mushrooms", "Ground Pork Lean", "Russet Potatoes",
"Veal", "Wheat Flour"]
# ht_indig_foods = []
ht_se_asia_foods = ["HT Bamboo Shoots", "Baby Corn", "HT Coconut Milk", "Thai Curry Paste",
"Edamame", "Eggplant", "HT Fish Sauce", "Vermicelli",
"Oyster Sauce", "Rice Flour", "Rice Noodles", "Jasmine Rice Mahatma Thai",
"Fresh Squash", "Tapioca Flour", "Tofu"]
ht_west_af_foods = ["Harris Teeter Dry Blackeye Peas", "Harris Teeter Dry Pinto Beans", "Chicken Whole",
"Fresh Plantain", "Russet Potatoes", "Harris Teeter Tomato Paste"]
# + code_folding=[0]
# Universal Basket for Harris Teeter
ht_df, ht_zips, ht_locs = food_helper.ht(ht_foods, zipcodes)
ht_df.head()
# + code_folding=[0]
# Latinx Basket for Harris Teeter
ht_hispanic_df = food_helper.ht(ht_hispanic_foods, ht_zips, standard = False, has_zipcodes = True, locs = ht_locs)
ht_hispanic_df.head()
# + code_folding=[0]
# East African Basket for Harris Teeter
ht_east_af_df = food_helper.ht(ht_east_af_foods, ht_zips, standard = False, has_zipcodes = True, locs = ht_locs)
ht_east_af_df.head()
# + code_folding=[0]
# Eastern European Basket for Harris Teeter
ht_east_eu_df = food_helper.ht(ht_east_eu_foods, ht_zips, standard = False, has_zipcodes = True, locs = ht_locs)
ht_east_eu_df.head()
# + code_folding=[0]
# Southeast Asian Basket for <NAME>
ht_se_asia_df = food_helper.ht(ht_se_asia_foods, ht_zips, standard = False, has_zipcodes = True, locs = ht_locs)
ht_se_asia_df.head()
# + code_folding=[0]
# West African Basket for <NAME>
ht_west_af_df = food_helper.ht(ht_west_af_foods, ht_zips, standard = False, has_zipcodes = True, locs = ht_locs)
ht_west_af_df.head()
# + code_folding=[0]
# <NAME>eter to csv
# ht_east_af_df.to_csv('HT_East_Af_Food.csv')
# ht_east_eu_df.to_csv('HT_East_Eu_Food.csv')
# ht_se_asia_df.to_csv('HT_SE_Asia_Food.csv')
# ht_west_af_df.to_csv('HT_West_Af_Food.csv')
# -
# ## Safeway
# + code_folding=[0]
# Safeway Basket
sw_foods = ["Value Corner Whole Milk - 1 Gallon", "Large Eggs - 12 Count", "Plain Bagels", "signature Select Bread White",
"Signature Select Pasta 16 oz", "Gala Apple", "Signature Farms Ground Turkey", "Signature Select Ham Cooked 95%",
"Value Corner Butter", "Russet Potatoes 5", "Navel Oranges", "Signature Select Tomato Sauce 8",
"Ground Beef 80% 1.25", "Shredded Cheese 8", "Yogurt Lucerne 32", "Signature Select Cereal Corn Flakes"]
sw_hispanic_foods = ["Hass Avocado", "Beets - 1 Bunch", "Signature Select Black Beans", "Mexican Cheese Blend 8", "Signature Select Corn Tortillas",
"Signature Select Green Chiles", "Jalapeno Peppers", "Signature Select Garbanzo Beans", "Masa",
"Giant Mixed Fruit Large", "Bell Peppers Green", "Signature Select Pinto Beans", "White Rice 5lb",
"Signature Select Salsa 16", "Green Squash", "Tomatillos", "Tomatoes On The Vine"]
# Mixed fruit?
sw_east_af_foods = ["Barley", "Simply Curry Powder", "Dried Beans", "Sunsweet Dates",
"Fava Beans", "Garbanzo Beans", "Lentil Beans", "Signature Spaghetti",
"Signature Tomato Sauce", "Select Tuna Chunk"]
sw_east_eu_foods = ["Barley", "Ground Beef 80% 1.25", "Beets 1 Bunch",
"Red Cabbage", "Lamb Shoulder", "Sliced Mushroom",
"Ground Pork", "Russet Potatoes", "Wheat Flour"]
# had to drop kasha
# sort beets by price
# sw_indig_foods = []
sw_se_asia_foods = ["Light Coconut Milk", "Signature Curry Paste", "Edamame",
"Eggplant", "Fish Sauce", "Oyster Sauce", "Rice Flour", "Rice Noodles",
"Jasmine Rice", "Squash", "Tofu"]
# had to drop bamboo shoots - was getting toothbrush or something?
sw_west_af_foods = ["Black eye Peas", "Dry Pinto Beans", "Chicken Whole", "Corn Flour",
"Plantain", "Russet Potatoes", "Tomato Paste", "Yams"]
# + code_folding=[0]
# Universal Basket for Safeway
sw_df, sw_zips, sw_locs = food_helper.sw(sw_foods, zipcodes)
sw_df.head()
# + code_folding=[0]
# Latinx Basket for Safeway
sw_hispanic_df = food_helper.sw(sw_hispanic_foods, sw_zips, standard = False, has_zipcodes = True, locs = sw_locs)
sw_hispanic_df.head()
# + code_folding=[0]
# East African Basket for Safeway
sw_east_af_df = food_helper.sw(sw_east_af_foods, sw_zips, standard = False, has_zipcodes = True, locs = sw_locs)
sw_east_af_df.head()
# + code_folding=[0]
# Eastern European Basket for Safeway
sw_east_eu_df = food_helper.sw(sw_east_eu_foods, sw_zips, standard = False, has_zipcodes = True, locs = sw_locs)
sw_east_eu_df.head()
# + code_folding=[0]
# Southeast Asian Basket for Safeway
sw_se_asia_df = food_helper.sw(sw_se_asia_foods, sw_zips, standard = False, has_zipcodes = True, locs = sw_locs)
sw_se_asia_df.head()
# + code_folding=[0]
# West African Basket for Safeway
sw_west_af_df = food_helper.sw(sw_west_af_foods, sw_zips, standard = False, has_zipcodes = True, locs = sw_locs)
sw_west_af_df.head()
# + code_folding=[0]
# Safeway to csv
# sw_east_af_df.to_csv('SW_East_Af_Food.csv')
# sw_east_eu_df.to_csv('SW_East_Eu_Food.csv')
# sw_se_asia_df.to_csv('SW_SE_Asia_Food.csv')
# sw_west_af_df.to_csv('SW_West_Af_Food.csv')
# -
# ## Aldi
# + code_folding=[0]
# Aldi Basket
aldi_foods = ["Whole Milk", "Large Eggs A Goldhen Eggs", "L'oven Everything", "White Bread",
"Penne Rigate", "Gala Apple", "Turkey Ground Lean 93", "Deli Sliced Ham",
"Countryside Salted Butter", "Bag Potatoes", "Oranges Bag", "Happy Harvest Tomato Sauce",
"Lean Beef Chub", "Happy Cheese Cubes", "Friendly Farms Nonfat Yogurt Light Plain", "Millville Frosted Shredded"]
aldi_hispanic_foods = ["Large Avocado", "Dakota Black Beans", "Mexican Cheese", "Corn Tortillas",
"Green Chiles", "Jalapeno Peppers", "Garbanzo Beans", "Masa",
"Mixed Fruit", "Bell Peppers Green", "Pinto Beans", "White Rice",
"Casa Medium Salsa", "Yellow Squash", "Tomatillos", "Roma Tomatoes"]
## need to fix the food items in this list.......
# no beets
# no tomatilos???
aldi_east_af_foods = ["Pueblo Pinto Beans", "Garbanzo Beans", "Spaghetti",
"Tomato Sauce", "Canned Tuna"]
aldi_east_eu_foods = ["Lean Beef Chub", "Cabbage", "Lamb", "Sliced Mushrooms",
"Pork", "Bag Potatoes"]
# sort beets by price
# aldi_indig_foods = []
aldi_se_asia_foods = ["Coconut Milk", "Eggplant", "Parboiled Rice",
"Yellow Squash", "Tofu"]
# number 2 option here for coconut milk!
aldi_west_af_foods = ["Dry Pinto Beans", "Chicken Whole", "Corn Flour", "Plantain",
"Bag Potatoes", "Tomato Paste", "Yams"]
# + code_folding=[0]
# Universal Basket for Aldi
aldi_df, aldi_zips, aldi_locs = food_helper.aldi(aldi_foods, zipcodes)
aldi_df.head()
# + code_folding=[0]
# Latinx Basket for Aldi
aldi_hispanic_df = food_helper.aldi(aldi_hispanic_foods, aldi_zips, standard = False, has_zipcodes = True, locs = aldi_locs)
aldi_hispanic_df.head()
# + code_folding=[0]
# East African Basket for Aldi
aldi_east_af_df = food_helper.aldi(aldi_east_af_foods, aldi_zips, standard = False, has_zipcodes = True, locs = aldi_locs)
aldi_east_af_df.head()
# + code_folding=[0]
# Eastern European Basket for Aldi
aldi_east_eu_df = food_helper.aldi(aldi_east_eu_foods, aldi_zips, standard = False, has_zipcodes = True, locs = aldi_locs)
aldi_east_eu_df.head()
# + code_folding=[0]
# Southeast Asian Basket for Aldi
aldi_se_asia_df = food_helper.aldi(aldi_se_asia_foods, aldi_zips, standard = False, has_zipcodes = True, locs = aldi_locs)
aldi_se_asia_df.head()
# + code_folding=[0]
# West African Basket for Aldi
aldi_west_af_df = food_helper.aldi(aldi_west_af_foods, aldi_zips, standard = False, has_zipcodes = True, locs = aldi_locs)
aldi_west_af_df.head()
# + code_folding=[0]
# Aldi to csv
# aldi_east_af_df.to_csv('Aldi_East_Af_Food.csv')
# aldi_east_eu_df.to_csv('Aldi_East_Eu_Food.csv')
# aldi_se_asia_df.to_csv('Aldi_SE_Asia_Food.csv')
# aldi_west_af_df.to_csv('Aldi_West_Af_Food.csv')
# -
# ## Target
# + code_folding=[0]
# Target Basket
target_foods = ["Vitamin D Whole Milk - 1 gal", "Grade A Large Eggs 12", "Plain Bagel 6", "Bread Market Pantry",
"Penne Rigate 16", "Gala Apple Each", "Ground Turkey", "Sliced Ham",
"Butter", "Russet Potatoes", "Navel Orange", "Tomato Sauce 8oz - Good & Gather",
"Ground Beef 1lb 80", "Shredded Cheese", "Yoplait Yogurt 32oz", "Market Pantry Cereal"]
target_hispanic_foods = ["Avocado", "Good and Gather Black Beans", "Mexican Cheese", "Corn Tortillas",
"Green Chiles", "Jalapeno Peppers", "Garbanzo Beans", "Masa",
"Fruit Cocktail Market", "Bell Peppers", "Pinto Beans", "White Rice 5",
"Medium Salsa 24", "Squash", "Good and Gather Salsa Verde", "Roma Tomatoes"]
# SOME HAVE TOMATILLOS, OTHERS TOMATILLO SALSA, AND OTHERS SALSA VERDE...
target_east_af_foods = ["Barley", "Badia Curry Powder", "Dried Beans", "Dried Dates",
"Garbanzo Beans", "Faraon Lentil Beans", "Good And Gather Spaghetti",
"Tomato Sauce 8oz", "Good And Gather Canned Tuna"]
# sort tomato sauce by price
target_east_eu_foods = ["Barley", "Ground Beef 80%", "Good and Gather Sliced Beets", "Cabbage",
"Sliced White Mushrooms", "Russet Potatoes", "Wheat Flour"]
# sort beets by price
# target_indig_foods = []
target_se_asia_foods = ["Baby Corn", "Coconut Milk", "Patak Curry Paste", "Edamame",
"Three Crabs Fish Sauce", "Thai Vermicelli", "Rice Noodles", "Jasmine Rice"]
# number 2 option here for coconut milk!
target_west_af_foods = ["Dry Black Eye Peas", "Dry Pinto Beans", "Corn Flour", "Maggi Cubes Chicken",
"Platano", "Russet Potatoes", "Tomato Paste", "Yams"]
# unsure on chicken...
# make alteration for platano!!1
# + code_folding=[0]
# Universal Basket for Target
target_df, target_zips, target_locs = food_helper.target(target_foods, zipcodes)
target_df.head()
# + code_folding=[0]
# Latinx Basket for Target
target_hispanic_df = food_helper.target(target_hispanic_foods, target_zips, standard = False, has_zipcodes = True, locs = target_locs)
target_hispanic_df.head()
# + code_folding=[0]
# East African Basket for Target
target_east_af_df = food_helper.target(target_east_af_foods, target_zips, standard = False, has_zipcodes = True, locs = target_locs)
target_east_af_df.head()
# + code_folding=[0]
# Eastern European Basket for Target
target_east_eu_df = food_helper.target(target_east_eu_foods, target_zips, standard = False, has_zipcodes = True, locs = target_locs)
target_east_eu_df.head()
# + code_folding=[0]
# Southeast Asian Basket for Target
target_se_asia_df = food_helper.target(target_se_asia_foods, target_zips, standard = False, has_zipcodes = True, locs = target_locs)
target_se_asia_df.head()
# + code_folding=[0]
# West African Basket for Target
target_west_af_df = food_helper.target(target_west_af_foods, target_zips, standard = False, has_zipcodes = True, locs = target_locs)
target_west_af_df.head()
# + code_folding=[0]
# Target to csv
# target_east_af_df.to_csv('Target_East_Af_Food.csv')
# target_east_eu_df.to_csv('Target_East_Eu_Food.csv')
# target_se_asia_df.to_csv('Target_SE_Asia_Food.csv')
# target_west_af_df.to_csv('Target_West_Af_Food.csv')
# -
# ## Whole Foods
# + code_folding=[0]
# Whole Foods Basket
wfm_foods = ["Whole Milk, 1 Gallon", "Large Eggs", "Plain Bagels", "Sandwich Bread Whole Wheat",
"Pasta Penne Rigate", "Fuji Apple", "Ground Turkey Thigh", "Sliced Ham Value Pack",
"Butter, Unsalted 4 Quarters", "Organic Russet Potato", "Navel Orange", "Tomato Sauce",
"Ground Beef 80%", "Shredded Mexican Blend", "Organic Yogurt Whole Milk", "Frosted Wheat Squares"]
wfm_hispanic_foods = ["Large Hass Avocado", "Shelf-Stable Black Beans", "Shreddedd Mexican Blend", "Organic Tortillas",
"Large Green Chiles", "Jalapeno Peppers", "Shelf-Stable Garbanzo Salt",
"Green Bell Pepper", "Shelf-Stable Pinto Salt", "White Rice 80",
"Medium Salsa Batch", "Squash", "Tomatillo", "Roma Tomato"]
# No Masa
# no mixed fruit?
wfm_east_af_foods = ["Barley", "Curry Powder", "Dried Fruit Dates",
"Salt Stable Garbanzo", "Red Lentils", "Pasta Spaghetti",
"Teff", "Tomato Sauce", "Canned Tuna 5oz Everday"]
wfm_east_eu_foods = ["Barley", "Ground Beef 80%", "Red Beet Bunch", "Cabbage",
"Sliced White Mushrooms", "Pork", "Russet Potatoes", "Wheat Flour"]
# wfm_indig_foods = []
wfm_se_asia_foods = ["Bamboo Shoots", "Baby Corn", "Vanilla Coconut Milk", "Thai Curry Paste",
"Fish Sauce", "Vermicelli", "Rice Flour", "Rice Noodles",
"Thai Jasmine Rice", "Yellow Squash", "Tofu"]
wfm_west_af_foods = ["Black Eyed Peas", "Shelf Salt Pinto", "Whole Chicken", "Plantain",
"Russet Potatoes", "Tomato Paste"]
# -
# Probably want to change below: visit
# + code_folding=[0]
# Universal Basket for Whole Foods
wfm_df, wfm_zips, wfm_locs = food_helper.wfm(wfm_foods, zipcodes)
wfm_df.head()
# + code_folding=[0]
# Latinx Basket for Whole Foods
wfm_hispanic_df = food_helper.wfm(wfm_hispanic_foods, wfm_zips, standard = False, has_zipcodes = True, locs = wfm_locs)
wfm_hispanic_df.head()
# + code_folding=[0]
# East African Basket for Whole Foods
wfm_east_af_df = food_helper.wfm(wfm_east_af_foods, wfm_zips, standard = False, has_zipcodes = True, locs = wfm_locs)
wfm_east_af_df.head()
# + code_folding=[0]
# Eastern European Basket for Whole Foods
wfm_east_eu_df = food_helper.wfm(wfm_east_eu_foods, wfm_zips, standard = False, east_euro = True, has_zipcodes = True, locs = wfm_locs)
wfm_east_eu_df.head()
# + code_folding=[0]
# Southeast Asian Basket for Whole Foods
wfm_se_asia_df = food_helper.wfm(wfm_se_asia_foods, wfm_zips, standard = False, has_zipcodes = True, locs = wfm_locs)
wfm_se_asia_df.head()
# + code_folding=[0]
# East African Basket for Whole Foods
wfm_west_af_df = food_helper.wfm(wfm_west_af_foods, wfm_zips, standard = False, west_af = True, has_zipcodes = True, locs = wfm_locs)
wfm_west_af_df.head()
# + code_folding=[0]
# Whole Foods to csv
# wfm_east_af_df.to_csv('WFM_East_Af_Food.csv')
# wfm_east_eu_df.to_csv('WFM_East_Eu_Food.csv')
# wfm_se_asia_df.to_csv('WFM_SE_Asia_Food.csv')
# wfm_west_af_df.to_csv('WFM_West_Af_Food.csv')
# -
# ## Lidl
# + code_folding=[0]
# Lidl Baskets
# lidl
lidl_foods = ["Whole Milk", "12 Large Eggs", "Bagels", "Wheat Bread",
"Penne Pasta Rigate", "Gala Apples", "Ground Turkey", "Sliced Ham",
"Salted Butter", "Russet Potatoes", "Oranges", "Roma Tomatoes",
"80% Ground Beef", "Mexican Cheese", "Yogurt", "Frosted Flakes Cereal"]
# second option with whole milk
lidl_latinx_basket = ["Avocado", "Beets", "Black Beans", "Mexican Cheese",
"Small Corn Tortilla", "Green Bell Peppers, 3 Count",
"Pinto Beans", "Long Grain Enriched White Rice",
"Medium Salsa", "Squash", "Tomatillos", "Roma Tomatoes"]
# coocked beats??
# are we interested in canned or dry beans??
lidl_east_af_basket = ["Pinto Beans", "Dried Dates", "Garbanzo Beans", "Lentils", "Basmati Rice",
"Spaghetti Pasta", "Tomato Sauce", "Tuna"]
lidl_east_eu_basket = ["80% Ground Beef", "Beets", "Green Cabbage", "Ground Lamb", "Lentils",
"Mushrooms", "Pork", "Russet Potatoes", "Wheat Flour"]
lidl_se_asia_basket = ["Goya Coconut Milk", "Edamame", "Eggplant",
"Taste of Thai Rice Noodles", "Jasmine Rice", "Squash"]
lidl_west_af_basket = ["Black Eyed Peas", "Pinto Beans", "Farm Chicken Drumsticks", "Chicken Bouillon Cubes",
"Russet Potatoes", "Tomato Paste", "Red Yams", "Yuca"]
# what part of chicken...
lidl_baskets = [lidl_foods, lidl_latinx_basket, lidl_east_af_basket,
lidl_east_eu_basket, lidl_se_asia_basket, lidl_west_af_basket]
# + code_folding=[0]
# Lidl Drivers for all groups and to csv!
lidl_names = np.array(['Lidl_Universal_Food.csv', 'Lidl_Latinx_Food.csv', 'Lidl_East_Af_Food.csv',
'Lidl_East_Eu_Food.csv', 'Lidl_SE_Asia_Food.csv', 'Lidl_West_Af_Food.csv'])
food_helper.lidl(lidl_names, zipcodes, [], 2, 20, True, lidl_foods, lidl_latinx_basket,
lidl_east_af_basket, lidl_east_eu_basket, lidl_se_asia_basket, lidl_west_af_basket)
# -
# ## Putting DataFrames together
# + code_folding=[0]
# # TO CSV!
# # Universal
# giant_df.to_csv('Giant_Universal_Food.csv')
# ht_df.to_csv('HT_Universal_Food.csv')
# sw_df.to_csv('SW_Universal_Food.csv')
# aldi_df.to_csv('Aldi_Universal_Food.csv')
# target_df.to_csv('Target_Universal_Food.csv')
# wfm_df.to_csv('WFM_Universal_Food.csv')
# # Latinx
# giant_hispanic_df.to_csv('Giant_Latinx_Food.csv')
# ht_hispanic_df.to_csv('HT_Latinx_Food.csv')
# sw_hispanic_df.to_csv('SW_Latinx_Food.csv')
# aldi_hispanic_df.to_csv('Aldi_Latinx_Food.csv')
# target_hispanic_df.to_csv('Target_Latinx_Food.csv')
# wfm_hispanic_df.to_csv('WFM_Latinx_Food.csv')
# # East African
# giant_east_af_df.to_csv('Giant_East_Af_Food.csv')
# ht_east_af_df.to_csv('HT_East_Af_Food.csv')
# sw_east_af_df.to_csv('SW_East_Af_Food.csv')
# aldi_east_af_df.to_csv('Aldi_East_Af_Food.csv')
# target_east_af_df.to_csv('Target_East_Af_Food.csv')
# wfm_east_af_df.to_csv('WFM_East_Af_Food.csv')
# # Eastern European
# giant_east_eu_df.to_csv('Giant_East_Eu_Food.csv')
# ht_east_eu_df.to_csv('HT_East_Eu_Food.csv')
# sw_east_eu_df.to_csv('SW_East_Eu_Food.csv')
# aldi_east_eu_df.to_csv('Aldi_East_Eu_Food.csv')
# target_east_eu_df.to_csv('Target_East_Eu_Food.csv')
# wfm_east_eu_df.to_csv('WFM_East_Eu_Food.csv')
# # Indigenous/Native
# # giant_df.to_csv('Giant_Universal_Food.csv')
# # ht_df.to_csv('HT_Universal_Food.csv')
# # sw_df.to_csv('SW_Universal_Food.csv')
# # aldi_df.to_csv('Aldi_Universal_Food.csv')
# # target_df.to_csv('Target_Universal_Food.csv')
# # wfm_df.to_csv('WFM_Universal_Food.csv')
# # Southeast Asian
# giant_se_asia_df.to_csv('Giant_SE_Asia_Food.csv')
# ht_se_asia_df.to_csv('HT_SE_Asia_Food.csv')
# sw_se_asia_df.to_csv('SW_SE_Asia_Food.csv')
# aldi_se_asia_df.to_csv('Aldi_SE_Asia_Food.csv')
# target_se_asia_df.to_csv('Target_SE_Asia_Food.csv')
# wfm_se_asia_df.to_csv('WFM_SE_Asia_Food.csv')
# # West African
# giant_west_af_df.to_csv('Giant_West_Af_Food.csv')
# ht_west_af_df.to_csv('HT_West_Af_Food.csv')
# sw_west_af_df.to_csv('SW_West_Af_Food.csv')
# aldi_west_af_df.to_csv('Aldi_West_Af_Food.csv')
# target_west_af_df.to_csv('Target_West_Af_Food.csv')
# wfm_west_af_df.to_csv('WFM_West_Af_Food.csv')
# + code_folding=[0]
# # Put data together by basket across stores and into CSVs!
# # Universal
# universal_df = pd.concat([giant_df, ht_df, sw_df, aldi_df, target_df, wfm_df], ignore_index=True)
# display(universal_df.head())
# universal_df.to_csv('Universal_Food.csv')
# # Latinx
# latinx_df = pd.concat([giant_hispanic_df, ht_hispanic_df, sw_hispanic_df, aldi_hispanic_df, target_hispanic_df, wfm_hispanic_df], ignore_index=True)
# display(latinx_df.head())
# latinx_df.to_csv('Latinx_Food.csv')
# -
# ## Load dataframes for inspection
# + code_folding=[]
# LOAD DATAFRAMES
# Universal
giant_df = pd.read_csv('Giant_Universal_Food.csv')
ht_df = pd.read_csv('HT_Universal_Food.csv')
sw_df = pd.read_csv('SW_Universal_Food.csv')
aldi_df = pd.read_csv('Aldi_Universal_Food.csv')
target_df = pd.read_csv('Target_Universal_Food.csv')
wfm_df = pd.read_csv('WFM_Universal_Food.csv')
lidl_df = pd.read_csv('Lidl_Universal_Food.csv', index_col = 0)
# Latinx
giant_hispanic_df = pd.read_csv('Giant_Latinx_Food.csv')
ht_hispanic_df = pd.read_csv('HT_Latinx_Food.csv')
sw_hispanic_df = pd.read_csv('SW_Latinx_Food.csv')
aldi_hispanic_df = pd.read_csv('Aldi_Latinx_Food.csv')
target_hispanic_df = pd.read_csv('Target_Latinx_Food.csv')
wfm_hispanic_df = pd.read_csv('WFM_Latinx_Food.csv')
lidl_hispanic_df = pd.read_csv('Lidl_Latinx_Food.csv', index_col = 0)
# East African
giant_east_af_df = pd.read_csv('Giant_East_Af_Food.csv')
ht_east_af_df = pd.read_csv('HT_East_Af_Food.csv')
sw_east_af_df = pd.read_csv('SW_East_Af_Food.csv')
aldi_east_af_df = pd.read_csv('Aldi_East_Af_Food.csv')
target_east_af_df = pd.read_csv('Target_East_Af_Food.csv')
wfm_east_af_df = pd.read_csv('WFM_East_Af_Food.csv')
lidl_east_af_df = pd.read_csv('Lidl_East_Af_Food.csv', index_col = 0)
# Eastern European
giant_east_eu_df = pd.read_csv('Giant_East_Eu_Food.csv')
ht_east_eu_df = pd.read_csv('HT_East_Eu_Food.csv')
sw_east_eu_df = pd.read_csv('SW_East_Eu_Food.csv')
aldi_east_eu_df = pd.read_csv('Aldi_East_Eu_Food.csv')
target_east_eu_df = pd.read_csv('Target_East_Eu_Food.csv')
wfm_east_eu_df = pd.read_csv('WFM_East_Eu_Food.csv')
lidl_east_eu_df = pd.read_csv('Lidl_East_Eu_Food.csv', index_col = 0)
# Indigenous/Native
# giant_df = pd.read_csv('Giant_Universal_Food.csv')
# ht_df = pd.read_csv('HT_Universal_Food.csv')
# sw_df = pd.read_csv('SW_Universal_Food.csv')
# aldi_df = pd.read_csv('Aldi_Universal_Food.csv')
# target_df = pd.read_csv('Target_Universal_Food.csv')
# wfm_df = pd.read_csv('WFM_Universal_Food.csv')
# Southeast Asian
giant_se_asia_df = pd.read_csv('Giant_SE_Asia_Food.csv')
ht_se_asia_df = pd.read_csv('HT_SE_Asia_Food.csv')
sw_se_asia_df = pd.read_csv('SW_SE_Asia_Food.csv')
aldi_se_asia_df = pd.read_csv('Aldi_SE_Asia_Food.csv')
target_se_asia_df = pd.read_csv('Target_SE_Asia_Food.csv')
wfm_se_asia_df = pd.read_csv('WFM_SE_Asia_Food.csv')
lidl_se_asia_df = pd.read_csv('Lidl_SE_Asia_Food.csv', index_col = 0)
# West African
giant_west_af_df = pd.read_csv('Giant_West_Af_Food.csv')
ht_west_af_df = pd.read_csv('HT_West_Af_Food.csv')
sw_west_af_df = pd.read_csv('SW_West_Af_Food.csv')
aldi_west_af_df = pd.read_csv('Aldi_West_Af_Food.csv')
target_west_af_df = pd.read_csv('Target_West_Af_Food.csv')
wfm_west_af_df = pd.read_csv('WFM_West_Af_Food.csv')
lidl_west_af_df = pd.read_csv('Lidl_West_AF_Food.csv', index_col = 0)
# + code_folding=[]
# COMPILE LARGER DATAFRAMES
# Universal
universal_df = pd.concat([giant_df, ht_df, sw_df, aldi_df,
target_df, wfm_df, lidl_df], ignore_index = True)
universal_df = universal_df.loc[:, ~universal_df.columns.str.contains('^Unnamed')]
display(universal_df.head())
universal_df.to_csv('Universal_Food.csv')
# Latinx
latinx_df = pd.concat([giant_hispanic_df, ht_hispanic_df, sw_hispanic_df, aldi_hispanic_df,
target_hispanic_df, wfm_hispanic_df, lidl_hispanic_df], ignore_index = True)
latinx_df = latinx_df.loc[:, ~latinx_df.columns.str.contains('^Unnamed')]
display(latinx_df.head())
latinx_df.to_csv('Latinx_Food.csv')
# East African
east_af_df = pd.concat([giant_east_af_df, ht_east_af_df, sw_east_af_df, aldi_east_af_df,
target_east_af_df, wfm_east_af_df, lidl_east_af_df], ignore_index = True)
east_af_df = east_af_df.loc[:, ~east_af_df.columns.str.contains('^Unnamed')]
display(east_af_df.head())
east_af_df.to_csv('East_Af_Food.csv')
# Eastern European
east_eu_df = pd.concat([giant_east_eu_df, ht_east_eu_df, sw_east_eu_df, aldi_east_eu_df,
target_east_eu_df, wfm_east_eu_df, lidl_east_eu_df], ignore_index = True)
east_eu_df = east_eu_df.loc[:, ~east_eu_df.columns.str.contains('^Unnamed')]
display(east_eu_df.head())
east_eu_df.to_csv('East_Eu_Food.csv')
# Southeast Asian
se_asia_df = pd.concat([giant_se_asia_df, ht_se_asia_df, sw_se_asia_df, aldi_se_asia_df,
target_se_asia_df, wfm_se_asia_df, lidl_se_asia_df], ignore_index = True)
se_asia_df = se_asia_df.loc[:, ~se_asia_df.columns.str.contains('^Unnamed')]
display(se_asia_df.head())
se_asia_df.to_csv('SE_Asia_Food.csv')
# West African
west_af_df = pd.concat([giant_west_af_df, ht_west_af_df, sw_west_af_df, aldi_west_af_df,
target_west_af_df, wfm_west_af_df, lidl_west_af_df], ignore_index = True)
west_af_df = west_af_df.loc[:, ~west_af_df.columns.str.contains('^Unnamed')]
display(west_af_df.head())
west_af_df.to_csv('West_Af_Food.csv')
# +
universal_df = pd.read_csv('Universal_Food.csv', index_col = 0)
latinx_df = pd.read_csv('Latinx_Food.csv', index_col = 0)
east_af_df = pd.read_csv('East_Af_Food.csv', index_col = 0)
east_eu_df = pd.read_csv('East_Eu_Food.csv', index_col = 0)
se_asia_df = pd.read_csv('SE_Asia_Food.csv', index_col = 0)
west_af_df = pd.read_csv('West_Af_Food.csv', index_col = 0)
universal_df
# -
universal_df[::16]
latinx_df
# + code_folding=[]
# Instacart scrape...
foods = ["Potatoes", "Oranges", "Tomatoes", "Apples", "Ground Turkey", "Chicken Nuggets", "Ground Beef",
"Chicken Eggs", "Shredded Cheese", 'Whole Milk', "Butter", "Yogurt", "Multigrain Bread", "Pasta",
"Bagels", "Cereal"]
ic_driver = webdriver.Chrome(ChromeDriverManager().install())
ic_driver.get("https://www.instacart.com/store/")
ic_driver.maximize_window()
sleep_time = 2
driver_wait = 20
wait = WebDriverWait(ic_driver, driver_wait)
log_in = wait.until(EC.element_to_be_clickable((By.CLASS_NAME, 'css-1cfqs9t')))
log_in.click()
# /html/body/div[4]/div/div/div/div/div[2]/div[1]/form/div[1]/div/div[1]/input
email = wait.until(EC.element_to_be_clickable((By.XPATH, '//div/div/div/div/div[2]/div[1]/form/div[1]/div/div[1]/input')))
email.send_keys('<EMAIL>')
# /html/body/div[4]/div/div/div/div/div[2]/div[1]/form/div[2]/div/div[1]/input
password = wait.until(EC.element_to_be_clickable((By.XPATH, '//div/div/div/div/div[2]/div[1]/form/div[2]/div/div[1]/input')))
password.send_keys('<PASSWORD>')
# /html/body/div[4]/div/div/div/div/div[2]/div[1]/form/div[4]/button
click_log_in = wait.until(EC.element_to_be_clickable((By.XPATH, '//div/div/div/div/div[2]/div[1]/form/div[4]/button')))
click_log_in.click()
go_store = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div[1]/div[2]/div/div[4]/ul/li[1]/a')))
go_store.click()
go_top = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/header/div/div/div[2]/div[4]/div[2]/div/div/div[1]/div[4]/button')))
go_top.click()
info = []
for food in foods:
ic_driver.get('https://www.instacart.com/store/publix/search_v3/{}'.format(food))
element = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/div/div/div/div/div/div[1]/div/div[2]/div/div/div[4]/ul[1]/li[1]/div/div/div/div/a/div[3]')))
info.append(element.text)
# +
# Instacart scrape...
foods = ["Potatoes", "Oranges", "Tomatoes", "Apples", "Ground Turkey", "Chicken Nuggets", "Ground Beef",
"Chicken Eggs", "Shredded Cheese", 'Whole Milk', "Butter", "Yogurt", "Multigrain Bread", "Pasta",
"Bagels", "Cereal"]
ic_driver = webdriver.Chrome(ChromeDriverManager().install())
ic_driver.get("https://www.instacart.com/store/")
ic_driver.maximize_window()
sleep_time = 2
driver_wait = 20
wait = WebDriverWait(ic_driver, driver_wait)
log_in = wait.until(EC.element_to_be_clickable((By.CLASS_NAME, 'css-s55bq')))
log_in.click()#<button class="css-s55bq"><span class="css-utfnc">Log in</span></button>
# /html/body/div[4]/div/div/div/div/div[2]/div[1]/form/div[1]/div/div[1]/input
email = wait.until(EC.element_to_be_clickable((By.XPATH, '//div/div/div/div/div[2]/div[1]/form/div[1]/div/div[1]/input')))
email.send_keys('<EMAIL>')
# /html/body/div[4]/div/div/div/div/div[2]/div[1]/form/div[2]/div/div[1]/input
password = wait.until(EC.element_to_be_clickable((By.XPATH, '//div/div/div/div/div[2]/div[1]/form/div[2]/div/div[1]/input')))
password.send_keys('<PASSWORD>')
# /html/body/div[4]/div/div/div/div/div[2]/div[1]/form/div[4]/button
click_log_in = wait.until(EC.element_to_be_clickable((By.XPATH, '//div/div/div/div/div[2]/div[1]/form/div[4]/button')))
click_log_in.click()
# ERROR HERE
# <a class="css-d20dbd" href="/store/publix/storefront"><span class="css-12thh49"><span class="css-ks8hbe-StoreCompactCard"><span class="css-jewpb8-StoreCompactCard"></span><img src="https://d2d8wwwkmhfcva.cloudfront.net/72x/d2lnr5mha7bycj.cloudfront.net/warehouse/logo/57/29520839-7042-45a0-af82-54f973b4abe5.png" srcset="https://d2d8wwwkmhfcva.cloudfront.net/108x/d2lnr5mha7bycj.cloudfront.net/warehouse/logo/57/29520839-7042-45a0-af82-54f973b4abe5.png 1.5x, https://d2d8wwwkmhfcva.cloudfront.net/144x/d2lnr5mha7bycj.cloudfront.net/warehouse/logo/57/29520839-7042-45a0-af82-54f973b4abe5.png 2x" alt="Publix" width="60" height="60" loading="lazy" role="presentation" class="css-1qsdlnh-StoreCompactCard"></span><span class="css-1im00gz-StoreCompactCard"><span><span class="css-lf0nfd">Publix</span></span><span class="css-2jq6rl">Deli • Groceries • Organic</span><span class="css-e8mvzb"><span class="css-pjl6ah"><span class="css-i5e22u">Pickup 37.9mi</span></span><span class="css-pjl6ah"><span class="css-i5e22u">Accepts EBT</span></span><span class="css-pjl6ah"><span class="css-i5e22u">New</span></span></span></span></span></a>
go_store = wait.until(EC.element_to_be_clickable((By.CLASS_NAME, 'css-d20dbd')))
go_store.click()
# go_top = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/header/div/div/div[2]/div[4]/div[2]/div/div/div[1]/div[4]/button')))
# go_top.click()
info = []
for food in foods:
ic_driver.get('https://www.instacart.com/store/publix/search_v3/{}'.format(food))
element = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/div/div/div/div/div/div[1]/div/div[2]/div/div/div[4]/ul[1]/li[1]/div/div/div/div/a/div[3]')))
info.append(element.text)
# -
# #### Looking to add other stores in the DMV (Virginia specifically for VDH)
# ## Walmart
# + code_folding=[0]
# lists of food items for Walmart
walmart_foods = ["Whole Milk", "Great Value Eggs 12 Count", "Great Value Bagels", "Great Value Bread",
"Great Value Pasta", "Apples", "Ground Turkey", "Great Value Sliced Ham",
"Butter", "Potatoes", "Oranges", "Tomatoes",
"Ground Beef", "Shredded Cheese", "Great Value Yogurt", "Shredded Great Value Cereal 24"]
walmart_latinx_basket = ["Avocado", "Beets", "Great Value Black Beans, 16oz", "Great Value Shredded Cheese",
"Great Value Corn Tortilla", "Great Value Green Chiles", "Great Value Jalapenos",
"Great Value Corn Flour", "Green Bell Pepper", "Great Value Pinto Beans, 32oz",
"Great Value Long Grain White Rice", "Salsa", "Squash", "Tomatillos", "Tomatoes"]
walmart_east_af_basket = ["Barley", "Curry Powder", "Dried Pinto Beans", "Dried Dates",
"Fava Beans", "Great Value Garbanzo Chick Peas", "Lentils",
"Great Value Basmati Rice", "Spaghetti Pasta", "Tomato Sauce", "Great Value Tuna"]
walmart_east_eu_basket = ["Barley", "Beef", "Beets", "Cabbage", "Lentils",
"Sliced Mushrooms", "Pork", "Potatoes", "Enriched Wheat Flour, 5lb"]
walmart_se_asia_basket = ["Bamboo Shoots", "Baby Corn", "Silk Coconut Milk", "Curry Paste", "Eggplant",
"Fish Sauce", "Vermicelli Rice Noodles", "Oyster Sauce", "Rice Flour",
"Brown Rice Noodles", "Great Value Jasmine Rice", "Sardines",
"Squash", "Tofu"]
walmart_west_af_basket = ["Great Value Black Eyed Peas, 16oz", "Brown Beans", "Cassava", "Chicken",
"Great Value Corn Flour", "Knorr Cube Bouillon Chicken 24ct", "Plantains",
"Potatoes", "Tomato Paste"]
# -
# list of csv names
names = np.array(['Walmart_Universal_Food.csv', 'Walmart_Latinx_Food.csv', 'Walmart_East_Af_Food.csv',
'Walmart_East_Eu_Food.csv', 'Walmart_SE_Asia_Food.csv', 'Walmart_West_Af_Food.csv'])
walmart_driver = food_helper.create_walmart_driver()
unique_stores = food_helper.search_zips(walmart_driver, zipcodes)
walmart_all_stores_baskets = food_helper.walmart_driver_func(walmart_driver, unique_stores, 20, walmart_foods, walmart_latinx_basket, walmart_east_af_basket, walmart_east_eu_basket, walmart_se_asia_basket, walmart_west_af_basket)
# + code_folding=[]
## NEED TO FIGURE OUT THE ERROR ABOVE.....
# -
# ## Food Lion
# + code_folding=[0]
# Food Lion Foods
fl_foods = ["Whole Milk", "12 Large Eggs", "Plain Bagels", "Cha Ching Bread",
"Penne Pasta Rigate", "Gala Apples", "Food Lion Ground Turkey", "Food Lion Sliced Ham",
"Butter", "Russet Potatoes Bag", "Navel Oranges Bag", "Tomatoes Vine",
"Food Lion Lean Ground Beef 73%", "Food Lion Shredded Mexican Cheese", "Food Lion Yogurt", "Food Lion Cereal Honey Nut"]
#how to get cheaper whole milk
fl_latinx_basket = ["Hass Avocado", "Beets", "Food Lion Black Beans, 16oz", "Food Lion Shredded Mexican Cheese",
"Banderita Corn Tortilla", "Jalapenos", "Corn Flour",
"Green Bell Pepper", "Pinto Beans", "Food Lion White Rice",
"Food Lion Salsa Medium", "Squash", "Tomatillos", "Tomatoes Vine"]
# are canned beets ok?
# are we interested in canned or dry beans??
fl_east_af_basket = ["Quaker Barley", "Badia Curry Powder", "Pinto Beans",
"Food Lion Garbanzo Beans", "Lentils", "Mahatma Basmati Rice",
"Spaghetti Pasta", "Tomato Sauce", "Great Food Tuna"]
fl_east_eu_basket = ["Quaker Barley", "Food Lion Lean Ground Beef 73%", "Beets", "Buckwheat Grain",
"Cabbage", "Kasha", "Lamb Chuck", "Lentils",
"Sliced Mushrooms", "Pork", "Russet Potatoes Bag", "Wheat Flour"]
fl_se_asia_basket = ["Bamboo Shoots", "Bean Thread", "Geisha Coconut Milk", "Curry Paste Patak", "Eggplant",
"Fish Sauce", "Vermicelli Rice Noodles", "Goya Rice Flour",
"Brown Rice Noodles", "Food Lion Jasmine Rice", "Squash", "Tapioca Flour"]
fl_west_af_basket = ["Black Eyed Peas, 16oz", "Pinto Beans", "La Reyna Yuca Cassava", "Food Lion Chicken",
"Corn Flour", "Food Lion Bouillon", "Plantains",
"Russet Potatoes Bag", "Food Lion Tomato Paste"]
# +
baskets = [fl_foods, fl_latinx_basket, fl_east_af_basket, fl_east_eu_basket,
fl_se_asia_basket, fl_west_af_basket]
names = np.array(['FL_Universal_Food.csv', 'FL_Latinx_Food.csv', 'FL_East_Af_Food.csv',
'FL_East_Eu_Food.csv', 'FL_SE_Asia_Food.csv', 'FL_West_Af_Food.csv'])
food_helper.food_lion(baskets, names, zipcodes)
# +
# ARE PRICES ACTUAL PRICES OR INSTACART PRICES
# +
### Walgreens Prices available in Store only...
# +
### CVS missing lots of options
# +
### Dollar General - prices in store only
# -
# ## Amazon
# + code_folding=[0, 14]
# Amazon Fresh
af_foods = ["Gallon Whole Milk", "12 Eggland Large Eggs", "Fresh Brand Plain Bagels", "Bread",
"Penne Pasta Rigate", "Fresh Brand 3 Gala Apples", "Ground Turkey", "Sliced Ham",
"Amazon Brand Butter", "Potatoes", "Oranges", "Tomatoes Roma",
"Ground Beef", "Mexican Cheese", "Amazon Brand Traditional Non-Fat Yogurt", "Fresh Brand Cereal 18.7"]
#how to get cheaper whole milk
af_latinx_basket = ["Hass Avocado", "Red Beets", "Black Beans", "Mexican Cheese",
"Corn Tortilla Amazon Brand", "Jalapenos", "Corn Flour",
"Green Bell Pepper", "Pinto Beans", "Happy Belly White Rice 5",
"Salsa Medium", "Squash", "Tomatillos", "Tomatoes Roma"]
# are canned beets ok?
# are we interested in canned or dry beans??
af_east_af_basket = ["Happy Belly Barley", "Amazon Brand Curry Powder", "Pinto Beans", "Dried Dates",
"Amazon Brand Garbanzo Beans", "Goya Lentils", "Basmati Rice",
"Amazon Brand Spaghetti Pasta 16", "Teff", "Amazon Brand Tomato Sauce 8", "Fresh Brand Tuna"]
af_east_eu_basket = ["Happy Belly Barley", "Ground Beef", "Red Beets",
"Green Cabbage", "Lamb", "Goya Lentils",
"Fresh Brand White Mushrooms", "Pork", "Potatoes", "King Wheat Flour"]
af_se_asia_basket = ["Bamboo Shoots", "Baby Corn", "Bean Thread",
"Coconut Milk", "Curry Paste", "Amazon Brand Edamame", "Eggplant",
"Fish Sauce", "Rice Flour", "Taste of Thai Rice Noodles",
"Happy Belly Jasmine Rice", "Squash", "Tapioca Flour"]
af_west_af_basket = ["Black Eyed Peas", "Pinto Beans", "Chicken Drumsticks", "Corn Flour",
"Knorr Chicken Bouillon Cubes 24", "Potatoes", "Tomato Paste", "Red Yams"]
# what part of chicken...
# + code_folding=[0]
# run Amazon driver
names = np.array(['AF_Universal_Food.csv', 'AF_Latinx_Food.csv', 'AF_East_Af_Food.csv',
'AF_East_Eu_Food.csv', 'AF_SE_Asia_Food.csv', 'AF_West_Af_Food.csv'])
af_baskets = [af_foods, af_latinx_basket, af_east_af_basket,
af_east_eu_basket, af_se_asia_basket, af_west_af_basket]
food_helper.amazon(zipcodes, names, af_baskets)
# + code_folding=[0]
# load af dataframes
af_df = pd.read_csv('AF_Universal_Food.csv', index_col = 0)
af_hispanic_df = pd.read_csv('AF_Latinx_Food.csv', index_col = 0)
af_east_af_df = pd.read_csv('AF_East_Af_Food.csv', index_col = 0)
af_east_eu_df = pd.read_csv('AF_East_Eu_Food.csv', index_col = 0)
af_se_asia_df = pd.read_csv('AF_SE_Asia_Food.csv', index_col = 0)
af_west_af_df = pd.read_csv('AF_West_AF_Food.csv', index_col = 0)
# +
### BULK RETAILERS?
# -
# ## BJ's
# +
bj_driver = webdriver.Chrome(ChromeDriverManager().install())
bj_driver.get("https://www.bjs.com/search")
bj_driver.maximize_window()
wait = WebDriverWait(bj_driver, driver_wait)
time.sleep(sleep_time)
change_zip = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="header-margin"]/ul/li[4]/app-header-find-aclub-link-molecule/div/div[2]/span[2]')))
change_zip.click()
# +
# change_zip = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="header-margin"]/ul/li[4]/app-header-find-aclub-link-molecule/div/div[2]/span[2]')))
# change_zip.click()wa
# +
#
store_loc = []
for zipcode in zipcodes:
#
enter_zip = wait.until(EC.element_to_be_clickable((By.ID, 'search')))
enter_zip.clear()
enter_zip.send_keys(str(zipcode), '\n')
#
element = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="header-margin"]/ul/li[4]/app-header-find-aclub-link-molecule/div/div[2]/div/app-header-club-modal-details-molecule/div[2]/div[1]/div/div[1]/div[1]/p[1]')))
store_loc.append(element.text)
#
store_loc = np.array(store_loc)
_, idx = np.unique(store_loc, return_index = True)
bj_zipcodes = zipcodes[idx]
# +
bj_foods = ["Whole Milk", "Large Eggs", "Wellsley Bagels", "Bread",
"Penne Pasta Rigate", "Apples", "Turkey", "Ham",
"Butter", "Russet Potatoes", "Navel Oranges", "Tomatoes",
"Ground Beef", "Cheese", "Plain Yogurt", "Cinnamon Cereal"]
# only greek
# turkey is different for 2 stores... potatoes too
bj_latinx_basket = ["Beets", "Black Beans Bag", "Mexican Cheese", "Corn Flour",
"Bell Peppers", "Pinto Beans", "White Rice", "Medium Salsa", "Tomatoes"]
# selection is slightly different for corn flour
bj_east_af_basket = ["Curry Powder", "Pinto Beans", "Chickpeas", "Goya Lentils", "Basmati Rice",
"Spaghetti Pasta", "Tomato Sauce", "Tuna"]
# selection is slightly different for curr powder
# add if to handle this difference
bj_east_eu_basket = ["Ground Beef", "Beets", "Cabbage", "Lamb", "Goya Lentils",
"Mushrooms", "Pork", "Russet Potatoes", "Whole Wheat Flour"]
bj_se_asia_basket = ["Goya Coconut Milk", "Edamame", "Jasmine Rice", "Sardines", "Tofu"]
bj_west_af_basket = ["Pinto Beans", "Chicken", "Corn Flour",
"Bouillon", "Plantains", "Russet Potatoes", "Tomato Paste"]
# what part of chicken...
bj_baskets = [bj_foods, bj_latinx_basket, bj_east_af_basket,
bj_east_eu_basket, bj_se_asia_basket, bj_west_af_basket]
# +
bj_driver = webdriver.Chrome(ChromeDriverManager().install())
bj_driver.get("https://www.bjs.com/search")
bj_driver.maximize_window()
wait = WebDriverWait(bj_driver, driver_wait)
time.sleep(sleep_time)
change_zip = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="header-margin"]/ul/li[4]/app-header-find-aclub-link-molecule/div/div[2]/span[2]')))
change_zip.click()
time.sleep(sleep_time)
#
store_loc = []
all_prices = []
all_info = []
for zipcode in bj_zipcodes:
#
enter_zip = wait.until(EC.element_to_be_clickable((By.ID, 'search')))
enter_zip.clear()
enter_zip.send_keys(str(zipcode), '\n')
time.sleep(sleep_time)
#
element = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="header-margin"]/ul/li[4]/app-header-find-aclub-link-molecule/div/div[2]/div/app-header-club-modal-details-molecule/div[2]')))
store_loc.append(", ".join(element.text.split("Miles")[0].split("\n")[1:-2]))
#
make_my_club = wait.until(EC.element_to_be_clickable((By.CLASS_NAME, 'make-myclub')))
make_my_club.click()
time.sleep(sleep_time) # still didn't click..
store_prices = []
store_info = []
for basket in bj_baskets[:2]:
basket_prices = []
basket_info = []
for food in basket:
#
search = wait.until(EC.element_to_be_clickable((By.ID, 'searchTerm')))
search.clear()
search.send_keys(food, "\n")
time.sleep(2)
#
if food_helper.check_exists_by_xpath(bj_driver, '/html/body/app-root/div/div[2]/div/app-search-main/app-search-result-page-gb/div[1]/app-search-header/div/div[1]/p[2]'):
no_info = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/app-root/div/div[2]/div/app-search-main/app-search-result-page-gb/div[1]/app-search-header/div/div[1]/p[2]')))
#
if (no_info.text == "Here are some related results."):
price = np.nan
info = ""
#
else:
price = np.nan
info = "IDK WHAT HAPPENED"
# too quick and does not line up with the new paths...
elif food_helper.check_exists_by_xpath(bj_driver, '/html/body/app-root/div/div[2]/div/app-search-main/app-search-result-page-gb/div[2]/div[2]/div/app-products-container/div/div/div[1]/app-product-card/div/div'):
element = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/app-root/div/div[2]/div/app-search-main/app-search-result-page-gb/div[2]/div[2]/div/app-products-container/div/div/div[1]/app-product-card/div/div')))
item = element.text.replace("Top Rated\n", "").replace("Best Seller\n", "")
price = (float(item.split("\n$")[-1].split()[0]) + float(item.split("$")[-1].split()[-1])) / 2
info = item.split("\n")[0]
food_info.append(element.text)
#
else:
item = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/app-root/div/div[2]/div/app-pdp-preprocessor/div/app-pdp-layout-template/div/div/div/div[2]/div[1]/div[2]')))
price = float(item.text.split("$")[1].split("\n")[0])
info = item.text.split("\n")[0]
#
basket_prices.append(price)
basket_info.append(info)
#
store_prices.append(basket_prices)
store_info.append(basket_info)
#
change_zip = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="header-margin"]/ul/li[4]/app-header-find-aclub-link-molecule/div/div[2]/span[2]')))
change_zip.click()
all_prices.append(store_prices)
all_info.append(store_info)
#
store_loc = np.array(store_loc)
# +
#
bj_driver = webdriver.Chrome(ChromeDriverManager().install())
bj_driver.get("https://www.bjs.com/search")
bj_driver.maximize_window()
wait = WebDriverWait(bj_driver, driver_wait)
time.sleep(2 * sleep_time)
#
store_loc = []
all_prices = []
all_info = []
for zipcode in bj_zipcodes:
#
change_zip = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="header-margin"]/ul/li[4]/app-header-find-aclub-link-molecule/div/div[2]/span[2]')))
change_zip.click()
time.sleep(2 * sleep_time)
#
enter_zip = wait.until(EC.element_to_be_clickable((By.ID, 'search')))
enter_zip.clear()
enter_zip.send_keys(str(zipcode), '\n')
time.sleep(sleep_time)
#
element = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="header-margin"]/ul/li[4]/app-header-find-aclub-link-molecule/div/div[2]/div/app-header-club-modal-details-molecule/div[2]')))
store_loc.append(", ".join(element.text.split("Miles")[0].split("\n")[1:-2]))
#
make_my_club = wait.until(EC.element_to_be_clickable((By.CLASS_NAME, 'make-myclub')))
make_my_club.click()
time.sleep(sleep_time) # still didn't click..
#
store_prices = []
store_info = []
for basket in bj_baskets[:2]:
#
basket_prices = []
basket_info = []
for food in basket:
#
search = wait.until(EC.element_to_be_clickable((By.ID, 'searchTerm')))
search.clear()
search.send_keys(food, "\n")
time.sleep(2)
#
if food_helper.check_exists_by_xpath(bj_driver, '/html/body/app-root/div/div[2]/div/app-search-main/app-search-result-page-gb/div[1]/app-search-header/div/div[1]/p[2]'):
no_info = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/app-root/div/div[2]/div/app-search-main/app-search-result-page-gb/div[1]/app-search-header/div/div[1]/p[2]')))
#
if (no_info.text == "Here are some related results."):
price = np.nan
info = ""
#
else:
price = np.nan
info = "IDK WHAT HAPPENED"
# too quick and does not line up with the new paths...
elif food_helper.check_exists_by_xpath(bj_driver, '/html/body/app-root/div/div[2]/div/app-search-main/app-search-result-page-gb/div[2]/div[2]/div/app-products-container/div/div/div[1]/app-product-card/div/div'):
element = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/app-root/div/div[2]/div/app-search-main/app-search-result-page-gb/div[2]/div[2]/div/app-products-container/div/div/div[1]/app-product-card/div/div')))
item = element.text.replace("Top Rated\n", "").replace("Best Seller\n", "")
price = (float(item.split("\n$")[-1].split()[0]) + float(item.split("$")[-1].split()[-1])) / 2
info = item.split("\n")[0]
food_info.append(element.text)
#
else:
#
item = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/app-root/div/div[2]/div/app-pdp-preprocessor/div/app-pdp-layout-template/div/div/div/div[2]/div[1]/div[2]')))
info = item.text.split("\n")[0]
#
if "SOLD OUT" in item.text:
price = np.nan
#
else:
price = float(item.text.split("$")[1].split("\n")[0])
#
basket_prices.append(price)
basket_info.append(info)
#
store_prices.append(basket_prices)
store_info.append(basket_info)
#
all_prices.append(store_prices)
all_info.append(store_info)
#
store_loc = np.array(store_loc)
# -
# ## Sam's Club
# + code_folding=[]
# Sam's Club
sam_foods = ["Milk", "Grade A Large White Eggs", "Member's Mark Plain Bagels", "Bread",
"Penne Pasta Rigate", "Gala Apples", "Ground Turkey", "Sliced Ham",
"Butter", "Russet Potatoes", "Large Seedless Oranges", "Tomatoes Roma",
"Ground Beef 80%", "Mexican Cheese", "Nonfat Plain Yogurt", "Cereal Honey"]
#only greek yogurt
sam_latinx_basket = ["Avocado", "Ground Beef 80%", "Member's Mark Black Beans", "Mexican Cheese",
"Corn Tortilla", "Jalapenos", "Masa",
"Green Bell Peppers", "Member's Mark Pinto Beans", "White Rice",
"Salsa Medium", "Squash", "Tomatillos", "Tomatoes Roma"]
# what size for rice?? 25 or 50lbs... // same with salsa medium
sam_east_af_basket = ["Curry Powder", "Member's Mark Pinto Beans", "Chickpeas", "Basmati Rice",
"Spaghetti Pasta", "Tomato Sauce", "Member's Mark Tuna"]
sam_east_eu_basket = ["Ground Beef 80%", "Lamb", "Goya Lentils",
"Mushrooms", "Pork", "Russet Potatoes", "Wheat Flour"]
sam_se_asia_basket = ["Coconut Milk", "Rice Flour", "Taste of Thai Rice Noodles",
"Member's Mark Jasmine Rice", "Sardines", "Squash"]
sam_west_af_basket = ["Member's Mark Pinto Beans", "Chicken Breast", "Masa",
"Bouillon", "Russet Potatoes", "Tomato Paste"]
# what part of chicken...
sam_baskets = [sam_foods, sam_latinx_basket, sam_east_af_basket,
sam_east_eu_basket, sam_se_asia_basket, sam_west_af_basket]
# +
sam_driver = webdriver.Chrome(ChromeDriverManager().install())
sam_driver.get("https://www.samsclub.com/s/")
sam_driver.maximize_window()
# also has reCAPTCHA
# +
names = np.array(['Sam_Universal_Food.csv', 'Sam_Latinx_Food.csv', 'Sam_East_Af_Food.csv',
'Sam_East_Eu_Food.csv', 'Sam_SE_Asia_Food.csv', 'Sam_West_Af_Food.csv'])
sams_club(sam_driver, zipcodes, names, 20, 2, sam_foods, sam_latinx_basket, sam_east_af_basket,
sam_east_eu_basket, sam_se_asia_basket, sam_west_af_basket)
# +
## NEED TO FIGURE OUT LOCATION.. GETTING GENERAL NAME ISN'T TOUGH BUT WHAT ABOUT OTHER STORES...
# location = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div[2]/div[1]/div/div/div[2]/div/nav[2]/div/div[2]/div/div/button')))
# location.click()
# -
universal_df_copy = universal_df.copy()
universal_df_copy["name"] = universal_df_copy["Store"]
universal_df_copy = universal_df_copy.drop(["Item", "Price", "Other_Info", "Store"], axis = 1)
universal_df_copy.head()
# +
address = np.array([re.sub("^\\D*", "", universal_df_copy.Location.iloc[i]).split(",")[0] for i in range(len(universal_df_copy))])
most_info = np.array([re.sub("^\\D*", "", universal_df_copy.Location.iloc[i]) for i in range(len(universal_df_copy))])
zipcode = [int(most_info[i].split(" ")[-1][:5]) for i in range(len(most_info))] # good
state = [most_info[i].split(" ")[-2].strip(",") for i in range(len(most_info))] # includnig washington in DC?
city = [most_info[i].split(",")[-2].strip(" ") for i in range(len(most_info))] # good
# -
universal_df_copy['Address'] = address
universal_df_copy['City'] = city
universal_df_copy['State'] = state
universal_df_copy['zipcode'] = zipcode
universal_df_copy = universal_df_copy.drop(["Location"], axis = 1)
universal_df_copy.head()
universal_df_copy.to_csv("Store_Locations_df.csv")
# +
# # !pip install geopandas
# # !pip install geopy
# -
universal_df_copy = pd.read_csv("Store_Locations_df.csv", index_col = 0)
universal_df_copy_short = universal_df_copy.drop_duplicates().reset_index(drop=True)
universal_df_copy_short["City"] = [universal_df_copy_short["City"][i].replace(" VA", "").replace(" DC", "") for i in range(len(universal_df_copy_short))]
short_address = universal_df_copy_short["Address"]
short_city = universal_df_copy_short["City"]
short_state = universal_df_copy_short["State"]
short_zipcode = universal_df_copy_short["zipcode"]
# +
# errors = 0
# from geopy.geocoders import Nominatim
# locator = Nominatim(user_agent="myGeocoder")
# idxs = []
# lat = []
# long = []
# for i in range(len(short_address)):
# #
# try:
# location = locator.geocode("{}, {}, {} {}".format(short_address[i], short_city[i], short_state[i], short_zipcode[i]))
# print("{}, {}, {} {}".format(short_address[i], short_city[i], short_state[i], short_zipcode[i]))
# print("Latitude = {}, Longitude = {}".format(location.latitude, location.longitude))
# idxs.append(i)
# lat.append(location.latitude)
# long.append(location.longitude)
# #
# except:
# #
# print("Error at {}".format(i))
# errors += 1
# continue
# #
# print("\nThere are {} errors.".format(errors))
# -
universal_df_copy_short_ll = universal_df_copy_short.iloc[np.array(idxs)].copy()
universal_df_copy_short_ll["Latitude"] = lat
universal_df_copy_short_ll["Longitude"] = long
universal_df_copy_short_ll.head()
universal_df_copy_short_ll.to_csv("Store_Locations_df_w_ll.csv")
#trying to get latitude and longitude data for stores
big_location_df = pd.read_csv("export_w_target.csv", index_col = 0)
basic_stores = ["whole foods market", "harris teeter", "aldi", "giant", "safeway", "target"]
basic_stores_location_df = big_location_df[big_location_df['brand'].str.lower().isin(basic_stores)]
narrow_basic_stores_location_df = basic_stores_location_df[['brand', 'lat', 'long']].reset_index(drop = True)
narrow_basic_stores_location_df.to_csv("Narrow_basic_stores_location_df.csv")
# Aldi does not have zipcodes or towns...
# Target info is a bit werid but can be cleaned
# +
# HMart
giant_se_asia_foods = ["Coconut Milk", "Fish Sauce", "Vermicelli", "Oyster Sauce",
"Rice Flour", "Rice Noodles", "Jasmine Rice"]
# start hmart driver
hmart_driver = webdriver.Chrome(ChromeDriverManager().install())
hmart_driver.get("https://www.hmart.com/")
hmart_driver.maximize_window()
wait = WebDriverWait(hmart_driver, 20)
#
all_info = []
random_zips = zipcodes[::5]
for random_zip in random_zips:
#
click_zipcode = wait.until(EC.element_to_be_clickable((By.XPATH, "//header/div[2]/div/div/div[2]/div/div[1]")))
click_zipcode.click()
#
send_zip = wait.until(EC.element_to_be_clickable((By.ID, "zipcode_hfresh")))
send_zip.clear()
send_zip.send_keys(str(random_zip), "\n")
#
info = []
for food in giant_se_asia_foods:
#
search = wait.until(EC.element_to_be_clickable((By.ID, "search")))
search.clear()
search.send_keys(food, "\n")
#
element = wait.until(EC.element_to_be_clickable((By.XPATH, "//ol/li[1]/div/div")))
info.append(element.text.replace("\nAdd to Cart", "").replace("\nSALE", ""))
#
all_info.append(info)
# close hmart driver
hmart_driver.quit()
#
all_info = np.array(all_info)
# -
hmart_prices = np.array([[float(all_info[j][i].split("\n")[-1].split(" ")[0].replace("$", "")) for i in range(len(all_info[j]))] for j in range(len(all_info))])
# ## Kroger's
krogers_driver = webdriver.Chrome(ChromeDriverManager().install())
krogers_driver.get("https://www.k<EMAIL>.<EMAIL>/")
krogers_driver.maximize_window()
# throwing strange errors...
wait = WebDriverWait(krogers_driver, 20)
search = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="SearchBar-input-open"]')))
search.clear()
search.send_keys("Whole Milk", "\n")
# ## Wegmans
# + code_folding=[]
# Wegmans
wegmans_basket = ["Whole Milk", "Large Eggs", "Plain Bagels", "Bread",
"Wegmans Penne", "Gala Apples", "Ground Turkey", "Ham",
"Wegmans Butter", "Wegmans Russet Potatoes", "Oranges", "Tomatoes",
"Ground Beef", "Mexican Cheese", "Wegmans Plain Lowfat Yogurt", "Cereal Honey"]
wegmans_latinx_basket = ["Avocado", "Ground Beef", "Black Beans", "Mexican Cheese",
"Tortillas Corn", "Jalapenos", "Instant Corn Masa",
"Green Bell Peppers", "Pinto Beans", "Long Grain White Rice",
"Wegmans Medium Salsa", "Green Squash", "Tomatillos", "Tomatoes Roma"]
# what size for rice?? 25 or 50lbs... // same with salsa medium
wegmans_east_af_basket = ["Laxmi Curry Powder", "Pinto Beans", "Chickpeas", "Dried Dates",
"Fava Beans", "Goat Cubes", "Goya Lentils", "Wegmans Basmati Rice",
"Spaghetti Pasta", "Tomato Sauce", "Tuna"]
# maybe sort for cheap curry powder
wegmans_east_eu_basket = ["Barley", "Ground Beef", "Beets", "Lamb", "Goya Lentils", "Cabbage", "Kasha",
"Mushrooms", "Pork", "Wegmans Russet Potatoes", "Wheat Flour", "Veal"]
wegmans_se_asia_basket = ["Bamboo Shoots", "Baby Corn", "Bean Threads", "Silk Coconut Milk",
"Curry Paste", "Edamame", "Eggplant", "Fish Sauce",
"Vermicelli", "Oyster Sauce", "Rice Flour", "Thai Rice Noodles",
"Wegmans Jasmine Rice", "Sardines", "Squash", "Tapioca Flour", "Tofu"]
wegmans_west_af_basket = ["Black Eyed Peas", "Cassava", "Chicken Breast", "Instant Corn Masa",
"Goat Cubes", "Bouillon Cubes", "Palm Oil", "Dole Plantains",
"Wegmans Russet Potatoes", "Tomato Paste", "Yams"]
# what part of chicken...
wegmans_baskets = [wegmans_foods, wegmans_latinx_basket, wegmans_east_af_basket,
wegmans_east_eu_basket, wegmans_se_asia_basket, wegmans_west_af_basket]
# -
zip_df = pd.read_csv("csvData.csv")
all_va_zips = zip_df.zip
wegman_cities = ['Alexandria', 'Chantilly', 'Charlottesville', 'Dulles', 'Fairfax',
'Fredericksburg', 'Lake Manassas', 'Leesburg', 'Midlothian', 'Potomac',
'Short Pump', 'Tysons', 'Virginia Beach']
# +
### NEED TO ADD IN OTHER CULTURAL BASKETS!! - GOOD THING IS THAT WE HAVE INFO ON THE ENTIRE STATE!
wegmans_driver = webdriver.Chrome(ChromeDriverManager().install())
wegmans_driver.get("https://shop.wegmans.com/shop/categories")
wegmans_driver.maximize_window()
wait = WebDriverWait(wegmans_driver, 20)
in_store = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/div/div/div/div[2]/div/div/div[2]/span/shopping-context-item[3]/button')))
in_store.click()
all_info = []
for city in wegman_cities:
search = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/div[2]/div/header/div[2]/div[1]/div/div[2]/unata-shopping-selector-nav/div/div/react-context-selector/div/div[3]/button')))
search.click()
enter_zip = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/div/div/div/div[2]/div/div/div/div/div[1]/div[2]/form/div/input')))
enter_zip.clear()
enter_zip.send_keys(city, "\n")
select = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/div/div/div/div[2]/div/div/div/div/div[2]/div[2]/div/div/div/div/div[2]/button')))
select.click()
time.sleep(sleep_time)
info = []
for food in wegmans_basket:
search = wait.until(EC.element_to_be_clickable((By.ID, 'search-nav-input')))
search.clear()
search.send_keys(food, "\n")
time.sleep(sleep_time)
element = wait.until(EC.element_to_be_clickable((By.CLASS_NAME, 'cell-content-wrapper')))
info.append(element.text)
all_info.append(info)
item = np.array([[all_info[j][i].split("\n")[0] for i in range(len(all_info[j]))] for j in range(len(all_info))])
price = np.array([[float(all_info[j][i].split("$")[1].split(" ")[0]) for i in range(len(all_info[j]))] for j in range(len(all_info))])
wegmans_driver.quit()
# -
# Example:
# check if xpath exists, if not return false
def check_exists_by_xpath(driver, xpath):
'''
Description:
Check existence of xpath on page
Inputs:
webdriver: your webdriver
xpath: whatever element we are looking for
Outputs:
returns True if xpath exists, False if not
'''
# try to find element
try:
driver.find_element_by_xpath(xpath)
# throw exception and return false if unable to find
except NoSuchElementException:
return False
return True
# +
### TARGET
def target_locations(zipcodes, sleep_time = 2, driver_wait = 20):
'''
Description:
Identify unique Targets in the Arlington Area
Inputs:
zipcodes:
sleep_time: integer, system sleep time between certain processes, default = 2
driver_wait: integer, wait time for driver - default = 20
Outputs:
target_zipcodes: array of zipcodes shortened to identify unique Targets
unique_target_locs: array of Target locations
'''
# Create Target driver and visit website
target_driver = webdriver.Chrome(ChromeDriverManager().install())
target_driver.maximize_window()
target_driver.get("https://www.target.com/c/grocery/-/N-5xt1a")
# select store option to begin passing in zipcodes
wait = WebDriverWait(target_driver, driver_wait)
store = wait.until(EC.element_to_be_clickable((By.ID, 'storeId-utilityNavBtn')))
store.click()
# loop over zipcodes to collect information on store location
target_info = []
for zipcode in zipcodes:
# click on option to edit zipcode
edit = wait.until(EC.element_to_be_clickable((By.ID, 'zipOrCityState')))
edit.click()
# delete zipcode then send new zipcode (not sure why enter_zip.clear() did not work)
enter_zip = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[9]/div/div/div/div/div[1]/div/div[3]/div[1]/div/input')))
enter_zip.send_keys('\<KEY>')
enter_zip.send_keys(str(zipcode), "\n")
# extract store location information
element = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[9]/div/div/div/div/div[3]')))
target_info.append(element.text)
# close driver
target_driver.quit()
# get unique locations and corresponding zipcodes
unique_target_locs, target_idx = np.unique(target_info, return_index = True)
target_zipcodes = zipcodes[target_idx]
# clean location information
for i, loc in enumerate(unique_target_locs):
unique_target_locs[i] = loc.split("\n")[1]
#
return target_zipcodes, unique_target_locs
def target_driver(target_zipcodes, target_foods, sleep_time = 2, driver_wait = 20, standard = True):
'''
Description:
Extract item information for specific stores for searched food items
Inputs:
target_zipcodes: array of zipcodes to enter to identify closest stores
target_foods: array of food items to search for on website
sleep_time: integer, system sleep time between certain processes, default = 2
driver_wait: integer, wait time for driver - default = 20
standard: Boolean, True if using "Universal" basket of goods
Outputs:
target_items_by_zip: 2D array of food items by store (contains other information)
'''
# start driver and visit Target side
target_driver = webdriver.Chrome(ChromeDriverManager().install())
target_driver.maximize_window()
target_driver.get("https://www.target.com/c/grocery/-/N-5xt1a")
# loop over all unique Targets
target_items_by_zip = []
for k, zipcode in enumerate(target_zipcodes):
# click on option to select a store
wait = WebDriverWait(target_driver, driver_wait)
store = wait.until(EC.element_to_be_clickable((By.ID, 'storeId-utilityNavBtn')))
store.click()
# edit and then add zipcode
edit = wait.until(EC.element_to_be_clickable((By.ID, 'zipOrCityState')))
edit.click()
# erase old zipcode and send new one
enter_zip = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[9]/div/div/div/div/div[1]/div/div[3]/div[1]/div/input')))
enter_zip.send_keys('\<KEY>')
enter_zip.send_keys(str(zipcode), "\n")
# select the top store
go_top_store = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[9]/div/div/div/div/div[3]/div[2]/div[1]/button')))
go_top_store.click()
# loop over staples at store
targ_info = []
for food in target_foods:
# selects next closest beef option for this particular store
if (food == "Ground Beef 1lb 80") and (k == 4):
food = "Ground Beef 1lb 73"
# search for food item
target_driver.get("https://www.target.com/s?searchTerm={}".format(food))
# checks if search item is tomato sauce to sort by price
if (food == "Tomato Sauce 8oz"):
sort = wait.until(EC.element_to_be_clickable((By.XPATH, "/html/body/div[1]/div/div[4]/div[4]/div[2]/div/div[2]/div[3]/div[1]/div[2]/div[2]/div/div[2]/button")))
sort.click()
by_price = wait.until(EC.element_to_be_clickable((By.XPATH, "//li[3]/a/div/div")))
by_price.click()
time.sleep(sleep_time)
# checks if search item is NOT plantains (these are in different section for some reason...)
if (food != "Platano") and (check_exists_by_xpath(target_driver, '//*[@id="mainContainer"]/div[4]/div[2]/div/div[2]/div[3]/div/ul/li[1]/div/div[2]/div')):
# extract slightly cleaned pricing info
# info = wait.until(EC.element_to_be_clickable((By.XPATH, '/html/body/div[1]/div/div[4]/div[4]/div[2]/div/div[2]/div[3]/div/ul/li[1]/div/div[2]/div/div/div')))
info = wait.until(EC.element_to_be_clickable((By.XPATH, '//*[@id="mainContainer"]/div[4]/div[2]/div/div[2]/div[3]/div/ul/li[1]/div/div[2]/div')))
#/html/body/div[1]/div/div[4]/div[4]/div[2]/div/div[2]/div[3]/div/ul/li[1]/div/div[2]/div/div/div
# if food item is plantain - grab info from bottom of webpage
else:
info = wait.until(EC.element_to_be_clickable((By.XPATH, "/html/body/div[1]/div/div[4]/div[4]/div[2]/div/div[1]/div[3]/div[2]/div/ul/li[1]/div/div[2]/div/div/div")))
#
cleaned_info = "".join(re.split('(.\d\d)', info.text)[:-1])
targ_info.append(cleaned_info)
# append list of items to list and close driver
target_items_by_zip.append(targ_info)
target_driver.quit()
# convert from list to array
target_items_by_zip = np.array(target_items_by_zip)
return target_items_by_zip
def target_price_item_other(target_items_by_zip):
'''
Description:
Extracts price, item info, and other information from the 2D array of information passed in
Inputs:
target_items_by_zip: 2D array of food items by store (contains other information)
Outputs:
prices_target: 2D array of prices for each item at each store
cleaned_items_target: 2D array of products at each store
other_info_target: 2D array of additional product information
'''
# clean strings and separate price, items, and other information
cleaned_target = np.array([np.array([target_items_by_zip[j][i] for i in range(len(target_items_by_zip[j]))]) for j in range(len(target_items_by_zip))])
cleaned_items_target = np.array([np.array([cleaned_target[j][i].split("\n")[0] for i in range(len(cleaned_target[j]))]) for j in range(len(cleaned_target))])
# heinous
# prices_target = np.array([np.array([float(cleaned_target[j][i].split("\n")[-3].split(" ")[0].replace("$", "")) if cleaned_target[j][i].split("\n")[-1][:4] == "Free" else float(cleaned_target[j][i].split("\n")[-2].split(" ")[0].replace("$", "")) if cleaned_target[j][i].split("\n")[-1][:3] == "Buy" else float(cleaned_target[j][i].split("\n")[-2].split(" ")[0].replace("$", "")) if cleaned_target[j][i].split("\n")[-1][:3] == "Get" else float(cleaned_target[j][i].split("\n")[-1].split(" ")[0].replace("$", "")) for i in range(len(cleaned_target[j]))]) for j in range(len(cleaned_target))])
prices_target = np.array([np.array([float(re.sub("[^0-9\.]", "", cleaned_target[j][i].split("\n$")[-1].split(" ")[0])) for i in range(len(cleaned_target[j]))]) for j in range(len(cleaned_target))])
other_info_target = np.array([np.array(["" for i in range(len(cleaned_target[j]))]) for j in range(len(cleaned_target))])
return prices_target, cleaned_items_target, other_info_target
def target(foods, zipcodes, standard = True, has_zipcodes = False, locs = []):
'''
Description:
Runs combination of functions to go from a list of zipcodes and food items to final dataframe for Target
Inputs:
foods: array of food items
zipcodes: list of Arlington zipcodes
standard: Boolean, True if using "Universal" basket of goods
Outputs:
df: dataframe with price, info, etc.
'''
# if shortened zipcodes list passed in
if has_zipcodes:
shortened_zipcodes = zipcodes
shortened_locations = locs
# get shortened list of zipcodes if not provided
else:
shortened_zipcodes, shortened_locations = target_locations(zipcodes)
items_by_zip = target_driver(shortened_zipcodes, target_foods = foods, standard = standard)
prices_target, cleaned_items_target, other_info_target = target_price_item_other(items_by_zip)
df = make_df(prices_target, cleaned_items_target, other_info_target, shortened_locations, "Target")
# if we already have zipcodes and locations, return dataframe (else return all 3)
if has_zipcodes:
return df
return df, shortened_zipcodes, shortened_locations
# -
target_zipcodes, unique_target_locs = target_locations(zipcodes)
target_items_by_zip = target_driver(target_zipcodes, target_foods)
prices_target, cleaned_items_target, other_info_target = target_price_item_other(target_items_by_zip)
# +
# hispanic scraping
target_hispanic_foods_big = ["Avocado",
"Beets",
"Good and Gather Black Beans",
"Mexican Cheese",
"Corn Tortillas",
"Crackers",
"Green Chiles",
"Jalapeno Peppers",
"Garbanzo Beans",
"Masa",
"Fruit Cocktail Market",
"Nopalitos",
"Bell Peppers",
"Pinto Beans",
"White Rice 5",
"Medium Salsa 24",
"Squash",
"Tomatillos",
"Good and Gather Salsa Verde",
"Roma Tomatoes"]
target_hispanic_items_by_zip = target_driver(target_zipcodes, target_hispanic_foods_big, standard = False)
# +
# probably need to select in-store pickup??? really not clear here
# -
| data-raw/SupermarketStaplePrices_2.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Programación para *Data Science*
# ============================
#
# Intro101 - 05.1: Conceptos avanzados de Python
# --------------------------------------
#
# En este Notebook encontraréis dos conjuntos de ejercicios: un primer conjunto de ejercicios para practicar y que no puntuan, pero que recomendamos intentar resolver y un segundo conjunto que evaluaremos como actividad.
#
#
# Además, veréis que todas las actividades tienen una etiqueta que indica los recursos necesarios para llevarla a cabo. Hay tres posibles etiquetas:
#
# * <span style="font-family: Courier New; background-color: #82b74b; color: #000000; padding: 3px; ">NM</span> **Sólo materiales**: las herramientas necesarias para realizar la actividad se pueden encontrar en los materiales de la asignatura.
#
# * <span style="font-family: Courier New; background-color: #ffcc5c; color: #000000; padding: 3px; ">EG</span> **Consulta externa guiada**: la actividad puede requerir hacer uso de herramientas que no se encuentran en los materiales de la asignatura, pero el enunciado contiene indicaciones de dónde o cómo encontrar la información adicional necesaria para resolver la actividad.
#
# * <span style="font-family: Courier New; background-color: #f2ae72; color: #000000; padding: 3px; ">EI</span> **Consulta externa independente**: la actividad puede requerir hacer uso de herramientas que no se encuentran en los materiales de la asignatura, y el enunciado puede no incluir la descripción de dónde o cómo encontrar esta información adicional. Será necesario que el estudiante busque esta información utilizando los recursos que se han explicado en la asignatura.
#
# Es importante notar que estas etiquetas no indican el nivel de dificultad del ejercicio, sino únicamente la necesidad de consulta de documentación externa para su resolución. Además, recordad que las **etiquetas son informativas**, pero podréis consultar referencias externas en cualquier momento (aunque no se indique explícitamente) o puede ser que podáis hacer una actividad sin consultar ningún tipo de documentación. Por ejemplo, para resolver una actividad que sólo requiera los materiales de la asignatura, puedéis consultar referencias externas si queréis, ya sea tanto para ayudaros en la resolución como para ampliar el conocimiento!
#
# ---
#
# ## Ejercicios y preguntas teóricas para la actividad
#
# A continuación, encontraréis los **ejercicios y preguntas teóricas que debéis completar en esta actividad** y que forman parte de la evaluación de esta unidad.
# ### Ejercicio 1
#
# Un número primo es aquél que solo es divisible por él mismo y por 1.
#
# a) Escribe un código que compruebe si un número `x = 15` es solo divisible por 1 o por el mismo. Escribe este código usando un iterador (un `for` o un `while`) que barra todos los valores desde `2` a `x-1`. Crea una variable `divisible` que tenga por defecto valor `False` y asigne el valor `True` si a lo largo de la iteración encuentra un número natural divisible. Puedes usar el operador modulo `a % b` para saber si un numero `b` es divisible por `a`.
#
# <span style="font-family: Courier New; background-color: #82b74b; color: #000000; padding: 2px; ">NM</span>
# +
# Respuesta
# -
#
# b) Convierte tu código anterior en una función que compruebe si el número del argumento es primo o no, devolviendo True is es primo y False si no es primo. Comprueba tu función con los valores 492366587, 492366585, 48947 y 2,
#
# <span style="font-family: Courier New; background-color: #82b74b; color: #000000; padding: 2px; ">NM</span>
# +
# Respuesta
# -
#
# c) En el cálculo de la función anterior, una vez se ha encontrado un número que es divisible dentro del rango ya no tiene sentido comprobar el resto de números del rango. Por ejemplo si 10 ya es divisble entre 2, ya no hace falta probar de 3 en adelante pues ya sabemos que el número no es primo.
#
# Modifica la función anterior de la siguiente forma:
# - Una vez se encuentra el divisor, la iteración se interrumpe para no probar el resto de enteros.
# - La función devuelve
# - **Si es primo**: True
# - **Si no es primo**, el primer divisor mayor que 1.
#
# Puedes hacer uso del comando *break* dentro de un bucle para interrumpir este, puedes consultar más información sobre break en la documentación de python [aquí](https://docs.python.org/2/tutorial/controlflow.html).
#
# Comprueba tu función con los valores 492366585, 492366587, 48947 y 2,
#
# <span style="font-family: Courier New; background-color: #82b74b; color: #000000; padding: 2px; ">NM</span>
# +
# Respuesta
# -
# ### Ejercicio 2
#
# La Covid-19 es una enfermedad producida por la infección del virus SARS-CoV-2. La infección es transmisible de persona a persona y su contagiosidad depende de la cantidad del virus en las vías respiratorias. Si cada persona contagiada transmite la enfermedad a $\beta$ contactos en promedio por periodo de tiempo $t$, es posible estimar la evolución del contagio con un modelo matemático sencillo.
#
# Para $t=1$día, las transmisiones en España se han estimado a partir de su histórico de las semanas de Febrero y Marzo del 2020 una $\beta = 0.35$ transmissiones por día por infectado.
#
# Durante un periodo de tiempo (por ejempo un día $d$) la tasa de nuevos contagios se puede estimar como una proporción al número de contagiados del periodo anterior $N$:
#
# $$ \Delta N = N_{1} - N = \beta \cdot N$$ (1)
#
# Por tanto, podemos proyectar el número futuro de afectados como
#
# $$ N_{1} = N + \beta \cdot N = (1+\beta) \cdot N$$ (2)
#
# En dos días:
#
# $$ N_{2} = (1+\beta) \cdot N_{1} = (1+\beta)^2 \cdot N$$ (3)
#
# Y en general en D días tendremos
#
# $$N_{D} = (1+\beta)^D \cdot N$$ (4)
#
# Asumiendo este sencillo modelo:
#
# a) Implementa una función de dos parámetros (N: población infectada inicial, D: número de días), que devuelva el cálculo de afectados para D días siguiendo la ecuación (4). Suponiendo una población afectada de 4250 (población afectada en españa a día 13 de Marzo de 2020), usa la función para calcular la población estimada en 1, 2, 7 y 30 días.
#
# <span style="font-family: Courier New; background-color: #82b74b; color: #000000; padding: 2px; ">NM</span>
# Respuesta
#
# b) Sabiendo que los Servicios de Medicina Intensiva (SMI) disponen de 3363 camas para enfermos graves, y suponiendo que un 10% de los afectados por el covid-19 requerirán de SMI y una supervivencia del 2,5% (Exitus), escribe un código que calcule:
# - El día en curso (Día)
# - El total de afectados por el virus para cada día d (Afectados)
# - El total de ingresados en SMI por el virus para cada día d (Críticos)
# - El total de Exitus por el virus para cada día d (Exitus)
# - Si los servicios de SMI no pueden aceptar los ingresados para cada día $d$ (Estado: indicando Saturación/No Saturación)
#
# Imprime en pantalla la información de cada día durante una simulación de tres semanas, suponiendo que no hay recuperaciones, con una población afectada inicial 4250 y una $\beta = 0.35$ constante.
#
#
# <span style="font-family: Courier New; background-color: #82b74b; color: #000000; padding: 2px; ">NM</span>
# Respuesta
#
# c) Convierte el código anterior en una función que genere un archivo de texto con nombre `output.txt`, siguiendo este formato:
# ```
# Dia, Afectados, Críticos, Exitus, Estado
# 0, 4250, 425, 106, No Saturación
# 1, 5737, 573, 143, No Saturación
# 2, 7745, 774, 193, No Saturación
# ...
# ```
# Con los parámetros de entrada $N$, $D$, $\beta$, camas SMI.
#
# <span style="font-family: Courier New; background-color: #82b74b; color: #000000; padding: 2px; ">NM</span>
# Respuesta
# ### Ejercicio 3
#
# Dado el siguiente diccionario:
d = {"Alex":344334443, "Eva":5533443, "Cristina":443355, "Jonas":33223324}
# Escribid una función que pregunte al usuario que introduzca el nombre de una persona y muestre por pantalla el nombre de la persona y su teléfono.
#
# Tened en cuenta que:
#
# - La función debe controlar que el valor introducido por el usuario es un nombre que existe en el diccionario. En caso contrario, mostrará un mensaje de error ("El nombre introducido no corresponde a ninguna persona") y devolverá el valor False.
# - Debéis tener en cuenta que el nombre de las personas que nos pasan por parámetro puede ser en minúsculas, mayúsculas o una combinación de ambas, y que debemos encontrar el número de teléfono aunque la capitalización de la cadena entrada por el usuario no sea exactamente la misma que hemos almacenado en el diccionario.
# - Suponed que no hay acentos en los nombres.
#
# Nota 1: Para realizar la actividad, tendréis que capturar un texto que entrará el usuario. Consultad la [documentación oficial de la función input](https://docs.python.org/3/library/functions.html#input) para ver cómo hacerlo.
#
# Nota 2: También tendréis que pensar cómo tratar el hecho de que el usuario pueda utilizar mayúsculas y minúsculas en la escritura del nombre en el diccionario. ¡Os animamos a usar un buscador para intentar encontrar alguna alternativa para resolver este subproblema! ¡Recordad citar las referencias que hayáis usado para resolverlo!
#
# <span style="font-family: Courier New; background-color: #ffcc5c; color: #000000; padding: 3px; ">EG</span>
# +
d = {"Alex":344334443, "Eva":5533443, "Cristina":443355, "Jonas":33223324}
# Respuesta
# -
# Referencias consultadas:
#
# *Incluir aquí las referencias*
# ### Ejercicio 4
#
# Python dispone de un **idiom** muy útil conocido como `list comprehension`. Utilizando este **idiom**, proporcionad una expresión que devuelva las listas siguientes.
#
# Nota: Para realizar esta actividad necesitaréis investigar qué son las `list comprehension` y qué sintaxis utilizan. Para ello, se recomienda en primer lugar que utilicéis un buscador para encontrar información genérica sobre esta construcción. Después, os recomendamos que consultéis stackoverflow para ver algunos ejemplos de problemas que se pueden resolver con esta construcción.
#
#
# [stackoverflow](https://stackoverflow.com/) es un sitio de preguntas-y-respuestas muy popular entre programadores. Veréis que para la gran mayoría de las dudas que tengáis, habrá alguien que ya les habrá tenido (y consultado) anteriormente! Así pues, más allá de preguntar vosotros mismos las dudas allí (nosotros ya tenemos el foro del aula para ello!), consultar esta web os permitirá ver qué soluciones proponen otros programadores a estas dudas. A menudo habrá más de una solución a un mismo problema, y podréis valorar cuál es la más adecuada para vuestro problema.
#
# Para ver ejemplos de problemas que son adecuados para resolver con **list comprehensions**, os recomendamos leer las siguientes páginas:
# * https://stackoverflow.com/questions/12555443/squaring-all-elements-in-a-list
# * https://stackoverflow.com/questions/18551458/how-to-frame-two-for-loops-in-list-comprehension-python
# * https://stackoverflow.com/questions/24442091/list-comprehension-with-condition
# * https://stackoverflow.com/questions/41676212/i-want-to-return-only-the-odd-numbers-in-a-list
# * https://stackoverflow.com/questions/4260280/if-else-in-a-list-comprehension
#
# <span style="font-family: Courier New; background-color: #ffcc5c; color: #000000; padding: 3px; ">EG</span>
#
# a) Una lista con los valores $4 x^2$ donde $x$ es cada uno de los números de la lista `list_1`:
# +
list_1 = [1, 2, 3, 4, 5, 6, 7, 8, 9]
# Respuesta
# -
# b) Una lista con los valores $x/(x+1)$ donde $x$ es cada uno de los números de la lista `list_1`:
# +
list_1 = [1, 2, 3, 4, 5, 6, 7, 8, 9]
# Respuesta
# -
# c) Una lista con los valores $4x^2/(4x^2-1)$ donde $x$ es cada uno de los números de la lista `list_1`:
# +
list_1 = [1, 2, 3, 4, 5, 6, 7, 8, 9]
# Respuesta
# -
# ### Ejercicio 5
#
# Las funciones `lambda` son formas de expresar y definir funciones pequeñas sin necesidad de usar el constructor `def funcion():`.
#
# Lee sobre las funciones lambda, por ejemplo [aquí](https://www.w3schools.com/python/python_lambda.asp) o [aquí](https://realpython.com/python-lambda/)
#
# Escribe una función $f$ con argumento $n$, $f(n)$, que **devuelva una función** lambda, que esta a su vez devuelva $n$ copias de una cadena de caracteres en su argumento:
#
# <span style="font-family: Courier New; background-color: #f2ae72; color: #000000; padding: 3px; ">EI</span>
# + jupyter={"source_hidden": true}
# Respuesta
def f(n):
# Definir de la función usando una método lambda
return()
r = f(5)
r("hola") # Donde deberíamos ver 5 copias del literal "Hola "
# -
# ### Ejercicio Opcional
#
# Existe una expresión atribuida a <NAME> (1616) para la estimación del valor de $\pi$, consistente en:
#
# $$
# \frac{\pi}{2} = \prod_{n=1}^{N} (\frac{4n^2}{4n^2 - 1})
# $$
# si $N$ es suficientemente grande $N \to \infty$.
#
# Escribe una función que, dado una aproximación N, calcule una estimación de $\pi$ siguiendo la fórmula de Wallis.
#
#
#
# **Consideraciones:**
#
# - Investigad las funciones map, reduce
# - También podéis usar una list comprehension
# - Las funciones lambda os pueden ser útiles
#
# <span style="font-family: Courier New; background-color: #f2ae72; color: #000000; padding: 3px; ">EI</span>
# + attributes={"classes": ["sourceCode"], "id": ""}
# Respuesta
| 01-intro-101/python/practices/05-python-avanzado/05.2_python_avanzado.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Build a decision tree
# ## Several reminders of this code:
#
# 1. This is a rewriten code references to tutorial from [<NAME>](https://github.com/random-forests), since my original code is not in perfect structure.
# 2. This code includes ID3 and CART algorithm.
# 3. Input data should be pandas DataFrame
# 3. Strongly suggest using hot encoding for categorical data, especially for those categorical features with integers.
# 4. Feel free to contact me if you think there are some problems.
# 5. ***Pruning is not included yet, I will work on it very soon.***
# 6. ***I will add more hyperparameters into this code.***
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import seaborn as sns
import collections
# %matplotlib inline
# +
def is_numeric(value):
# This function is to determine if a value is numerical
return isinstance(value, int) or isinstance(value, float)
def unique_vals(dataset, feature):
"""Find the unique values for a column in a dataset."""
return sorted(set(dataset[feature]))
def class_counts(dataset):
"""
Parameters
------------------
dataset: {DataFrame}, shape = [m_samples, n_features]
Dateset matrix, where 'm_samples' is the number of samples
and 'n_features' is the number of features
Return
------------------
classCounts: dictionary {class: counts}
"""
classCounts = collections.defaultdict(int)
# the number of unique elements and their occurrence
for label in dataset.iloc[:,-1]:
if label not in classCounts:
classCounts[label] = 0
classCounts[label] += 1
return classCounts
# +
class Split:
def __init__(self, dataset, feature, splitPoint):
self.dataset = dataset
self.feature = feature
self.splitPoint = splitPoint
def match(self, comparision_sample):
# This is the function that compare the plit point we choosed to a given value
# If it is numerical, then True if splitPoint >= comparision_sample, False if splitPoint < comparision_sample
# If it is categorical, then True if splitPoint == comparision_sample, False if splitPoint != comparision_sample
val = comparision_sample[self.feature]
if is_numeric(val):
return self.splitPoint <= val
else:
return self.splitPoint == val
def __repr__(self):
# This is just a helper method to print
# the question in a readable format.
header = list(self.dataset.columns)
condition = "=="
if is_numeric(self.splitPoint):
condition = ">="
return "Is %s %s %s?" % (
header[header==self.feature], condition, str(self.splitPoint))
class Leaf:
""" A leaf node
A leaf node holds data including unique values and their counts in a dictionary.
"""
def __init__(self, dataset):
self.prediction = class_counts(dataset)
class Decision_Node:
"""A decision node
A decision node holds the split method and its two child tree.
"""
def __init__(self, split, left_tree, right_tree):
self.split = split
self.left_tree = left_tree
self.right_tree = right_tree
# +
class DecisionTreeClassifier:
def __init__(self, criterion = 'entropy'):
self.criterion = criterion
if self.criterion == 'gini':
self.info_gain = self._gini_gain
elif self.criterion == 'entropy':
self.info_gain = self._entropy_gain
def partition(self, dataset, split):
"""partition the dataset into left and right
For each value a in a feature, compare it to the split point. Partition the dataset into two subsets.
"""
left_set = dataset[split.match(dataset)]
right_set = dataset[split.match(dataset)==False]
return left_set, right_set
def gini_Impurity(self, dataset):
"""
Parameters
------------------
Return
------------------
giniImpurity: float
"""
classCounts = class_counts(dataset)
# calculate gini index
gini = 1.0
for key in classCounts:
# calculate occurrence
prob = float(classCounts[key]/(len(dataset)*1.0))
# calculate entropy
gini -= prob ** 2
return gini
def _gini_gain(self, dataset, left, right):
"""Gini gain
"""
gini = self.gini_Impurity(dataset)
p = float(len(left) / (len(left) +len(right)))
return gini - float(p * self.gini_Impurity(left)) - float((1-p) * self.gini_Impurity(right))
def calcShannonEntropy(self, dataset):
"""
Parameters
------------------
y_labels: {array-like}, shape = [n_samples, 1]
Dateset matrix, where 'n_samples' is the number of samples
and one column of labels
Return
------------------
shannonEntropy: float
"""
m, n = dataset.shape
labelCounts = collections.defaultdict(int)
classCounts = class_counts(dataset)
# calculate shannon entropy
shannonEntropy = 0.0
for key in classCounts:
# calculate occurrence
prob = float(classCounts[key]/m)
if prob == 0:
continue
# calculate entropy
shannonEntropy -= prob * np.log2(prob)
return shannonEntropy
def _entropy_gain(self, dataset, left, right):
"""Entropy gain
"""
entropy = self.calcShannonEntropy(dataset)
p = float(len(left) / (len(left) +len(right)))
return entropy - float(p*self.calcShannonEntropy(left)) - float((1-p)*self.calcShannonEntropy(right))
def find_best_split(self, dataset):
"""
Parameters
------------------
X_data: {array-like} discrete features
preprocessed dataset, could not deal with continuous features,
and categorical feature should be better as binary form
y_labels: {array-like}
Return
------------------
best_gain: the maximum gini index gain
best_split: the feature and split point that get the maximum gini index gain
"""
# Exclude Labels
features = list(dataset.columns[:-1])
best_gain = 0.0
best_split = None
# Loops:
# First for loop: features
# Second for loop: unique values in a feature.
for feature in features:
unique_val = unique_vals(dataset, feature)
for val in unique_val:
split = Split(dataset, feature, val)
left_set, right_set = self.partition(dataset, split)
# Skip this split if it doesn't divide the dataset
if len(left_set) == 0 or len(right_set) == 0:
continue
gain = self.info_gain(dataset, left_set, right_set)
if gain >= best_gain:
best_gain = gain
best_split = split
return best_gain, best_split
def fit(self,dataset):
"""
"""
#Step1: Find the best split feature and point, and create a root.
gain, split = self.find_best_split(dataset)
# If there is no gain, or gain is less than a threshold
# We will not split any more
# And left it as a leaf
if gain == 0:
return Leaf(dataset)
# Partition the dataset into two sub-trees
left_set, right_set = self.partition(dataset, split)
# Recursively build sub-trees, start from left to right
left_tree = self.fit(left_set)
right_tree = self.fit(right_set)
#
return Decision_Node(split, left_tree, right_tree)
def print_tree(self, node, spacing=""):
"""Copy from
World's most elegant tree printing function.
"""
# Base case: we've reached a leaf
if isinstance(node, Leaf):
print (spacing + "Predict", self.print_leaf(node.prediction))
return
# Print the question at this node
print (spacing + str(node.split))
# Call this function recursively on the true branch
print (spacing + '--> True:')
self.print_tree(node.left_tree, spacing + " ")
# Call this function recursively on the false branch
print (spacing + '--> False:')
self.print_tree(node.right_tree, spacing + " ")
def classify(self, dataset, node):
"""See the 'rules of recursion' above."""
# Base case: we've reached a leaf
if isinstance(node, Leaf):
return node.prediction
# Decide whether to follow the true-branch or the false-branch.
# Compare the feature / value stored in the node,
# to the example we're considering.
if node.split.match(dataset):
return self.classify(dataset, node.left_tree)
else:
return self.classify(dataset, node.right_tree)
def print_leaf(self, counts):
"""A nicer way to print the predictions at a leaf."""
total = sum(counts.values()) * 1.0
probs = {}
for lbl in counts.keys():
probs[lbl] = str(int(counts[lbl] / total * 100)) + "%"
return probs
def save_tree(self, inputTree, filename):
import pickle
fw = open(filename, 'wb')
pickle.dump(inputTree, fw)
fw.close()
def read_tree(self, filename):
import pickle
tr = open(filename,'rb')
return pickle.load(tr)
def predict(self, dataset, node):
predictions = []
m,n = dataset.shape
for i in range(m):
leaf = self.classify(dataset.iloc[i],node)
# print('The prediction is: {}'.format(self.print_leaf(leaf)))
predictions.append(self.print_leaf(leaf).keys())
return pd.DataFrame(predictions)
# -
DATA = pd.read_csv('mushrooms.csv')
X = DATA.iloc[:,1:]
y = DATA.iloc[:,0]
# data['labels'] = labels
X.head()
from sklearn.model_selection import train_test_split
X_train, X_test, y_train, y_test = train_test_split( X, y, test_size=0.33, random_state=42)
data = pd.concat([X_train,y_train],axis=1)
data.head()
test = pd.concat([X_test,y_test],axis=1)
dt = DecisionTreeClassifier()
tree = dt.fit(data)
dt.save_tree(tree,'tree')
tree = dt.read_tree('tree')
dt.print_tree(tree)
prediction = dt.predict(test,tree)
prediction
data['class']
from sklearn.metrics import confusion_matrix
confusion_matrix(test['class'],prediction)
# # Too good to be true, only because lack of pruning
| Decision Tree/Decision Tree Classifier.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ## Hello World using Flask
import os
hello_world_script_file = os.path.join(os.path.pardir,'src','models','hello_world_api.py')
# +
# %%writefile $hello_world_script_file
from flask import Flask,request
app = Flask(__name__)
@app.route('/api',methods=['POST'])
def say_hello():
data = request.get_json(force=True)
name = data['name']
return 'hello {0}'.format(name)
if __name__ == '__main__':
app.run(port=3000,debug=True)
# -
import json
import requests
url = 'http://localhost:3000/api'
data = json.dumps({'name' : 'bhavana'}) #creating json object for sending data
r = requests.post(url,data) #response from url
r.text
# ## Maching Learning with API
# #### Building API
import os
machine_learning_api_script_file = os.path.join(os.path.pardir,'src','models','machine_learning_api.py')
# +
# %%writefile $machine_learning_api_script_file
from flask import Flask,request
import pandas as pd
import numpy as np
import json
import pickle
import os
app = Flask(__name__)
model_path = os.path.join(os.path.pardir,os.path.pardir,'models')
model_filepath = os.path.join(model_path,'lr_model.pkl')
scaler_filepath = os.path.join(model_path,'lr_scaler.pkl')
scaler = pickle.load(open(scaler_filepath,'rb'))
model = pickle.load(open(model_filepath,'rb'))
columns = ["Age", "Fare", "FamilySize",
"IsMother", "IsMale", "Deck_A", "Deck_B", "Deck_C", "Deck_D",
"Deck_E", "Deck_F", "Deck_G", "Deck_z", "Pclass_1", "Pclass_2",
"Pclass_3", "Title_Lady", "Title_Master", "Title_Miss", "Title_Mr",
"Title_Mrs", "Title_Officer", "Title_Sir", "Fare_category_very_low",
"Fare_category_low", "Fare_category_high", "Fare_category_very_high", "Embarked_C",
"Embarked_Q", "Embarked_S", "AgeState_Adult", "AgeState_Child"]
@app.route('/api',methods=['POST'])
def make_prediction():
# read json object and convert to json string
data = json.dumps(request.get_json(force=True))
# create dataframe using json string
df = pd.read_json(data)
# extract passengerIds
passenger_ids = df['PassengerId'].ravel()
actuals = df['Survived'].ravel()
# convert data to matrix form
X = df[columns].as_matrix().astype('float')
X_scaled = scaler.transform(X)
predictions = model.predict(X_scaled)
df_response = pd.DataFrame({'PassengerId': passenger_ids, 'Predictions':predictions, 'Actual':actuals})
return df_response.to_json()
if __name__ == '__main__':
app.run(port=3000, debug=True)
# -
# #### Invoking API using requests
import os
import pandas as pd
processed_data_path = os.path.join(os.path.pardir,'data','processed')
train_file_path = os.path.join(processed_data_path,'train.csv')
train_df = pd.read_csv(train_file_path)
survived_passengers = train_df[train_df['Survived'] == 1][:5]
survived_passengers
import requests
def make_api_request(data):
url = 'http://localhost:3000/api'
r = requests.post(url,data)
return r.json()
make_api_request(survived_passengers.to_json())
result = make_api_request(train_df.to_json())
df_result = pd.read_json(json.dumps(result))
df_result.head()
import numpy as np
np.mean(df_result.Actual == df_result.Predictions)
| notebooks/Building-machine-learning-api.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# <a href="https://www.bigdatauniversity.com"><img src = "https://ibm.box.com/shared/static/cw2c7r3o20w9zn8gkecaeyjhgw3xdgbj.png" width = 400, align = "center"></a>
#
# # <center>K-Means Clustering</center>
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Introduction
#
# There are many models for **clustering** out there. In this notebook, we will be presenting the model that is considered the one of the simplest model among them. Despite its simplicity, the **K-means** is vastly used for clustering in many data science applications, especially useful if you need to quickly discover insights from **unlabeled data**. In this notebook, you learn how to use k-Means for customer segmentation.
#
# Some real-world applications of k-means:
# - Customer segmentation
# - Understand what the visitors of a website are trying to accomplish
# - Pattern recognition
# - Machine learning
# - Data compression
#
#
# In this notebook we practice k-means clustering with 2 examples:
# - k-means on a random generated dataset
# - Using k-means for customer segmentation
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Import libraries
# Lets first import the required libraries.
# Also run <b> %matplotlib inline </b> since we will be plotting in this section.
# + button=false new_sheet=false run_control={"read_only": false}
import random
import numpy as np
import matplotlib.pyplot as plt
from sklearn.cluster import KMeans
from sklearn.datasets.samples_generator import make_blobs
# %matplotlib inline
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # k-Means on a randomly generated dataset
# Lets create our own dataset for this lab!
#
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# First we need to set up a random seed. Use <b>numpy's random.seed()</b> function, where the seed will be set to <b>0</b>
# + button=false new_sheet=false run_control={"read_only": false}
np.random.seed(0)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Next we will be making <i> random clusters </i> of points by using the <b> make_blobs </b> class. The <b> make_blobs </b> class can take in many inputs, but we will be using these specific ones. <br> <br>
# <b> <u> Input </u> </b>
# <ul>
# <li> <b>n_samples</b>: The total number of points equally divided among clusters. </li>
# <ul> <li> Value will be: 5000 </li> </ul>
# <li> <b>centers</b>: The number of centers to generate, or the fixed center locations. </li>
# <ul> <li> Value will be: [[4, 4], [-2, -1], [2, -3],[1,1]] </li> </ul>
# <li> <b>cluster_std</b>: The standard deviation of the clusters. </li>
# <ul> <li> Value will be: 0.9 </li> </ul>
# </ul>
# <br>
# <b> <u> Output </u> </b>
# <ul>
# <li> <b>X</b>: Array of shape [n_samples, n_features]. (Feature Matrix)</li>
# <ul> <li> The generated samples. </li> </ul>
# <li> <b>y</b>: Array of shape [n_samples]. (Response Vector)</li>
# <ul> <li> The integer labels for cluster membership of each sample. </li> </ul>
# </ul>
#
# + button=false new_sheet=false run_control={"read_only": false}
X, y = make_blobs(n_samples=5000, centers=[[4,4], [-2, -1], [2, -3], [1, 1]], cluster_std=0.9)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Display the scatter plot of the randomly generated data.
# + button=false new_sheet=false run_control={"read_only": false}
plt.scatter(X[:, 0], X[:, 1], marker='.')
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Setting up K-Means
# Now that we have our random data, let's set up our K-Means Clustering.
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# The KMeans class has many parameters that can be used, but we will be using these three:
# <ul>
# <li> <b>init</b>: Initialization method of the centroids. </li>
# <ul>
# <li> Value will be: "k-means++" </li>
# <li> k-means++: Selects initial cluster centers for k-mean clustering in a smart way to speed up convergence.</li>
# </ul>
# <li> <b>n\_clusters</b>: The number of clusters to form as well as the number of centroids to generate. </li>
# <ul> <li> Value will be: 4 (since we have 4 centers)</li> </ul>
# <li> <b>n\_init</b>: Number of time the k-means algorithm will be run with different centroid seeds. The final results will be the best output of n\_init consecutive runs in terms of inertia. </li>
# <ul> <li> Value will be: 12 </li> </ul>
# </ul>
#
# Initialize KMeans with these parameters, where the output parameter is called <b>k_means</b>.
# + button=false new_sheet=false run_control={"read_only": false}
k_means = KMeans(init = "k-means++", n_clusters = 4, n_init = 12)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Now let's fit the KMeans model with the feature matrix we created above, <b> X </b>
# + button=false new_sheet=false run_control={"read_only": false}
k_means.fit(X)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Now let's grab the labels for each point in the model using KMeans' <b> .labels\_ </b> attribute and save it as <b> k_means_labels </b>
# + button=false new_sheet=false run_control={"read_only": false}
k_means_labels = k_means.labels_
k_means_labels
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# We will also get the coordinates of the cluster centers using KMeans' <b> .cluster_centers_ </b> and save it as <b> k_means_cluster_centers </b>
# + button=false new_sheet=false run_control={"read_only": false}
k_means_cluster_centers = k_means.cluster_centers_
k_means_cluster_centers
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Creating the Visual Plot
# So now that we have the random data generated and the KMeans model initialized, let's plot them and see what it looks like!
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# Please read through the code and comments to understand how to plot the model.
# + button=false new_sheet=false run_control={"read_only": false}
# Initialize the plot with the specified dimensions.
fig = plt.figure(figsize=(6, 4))
# Colors uses a color map, which will produce an array of colors based on
# the number of labels there are. We use set(k_means_labels) to get the
# unique labels.
colors = plt.cm.Spectral(np.linspace(0, 1, len(set(k_means_labels))))
# Create a plot
ax = fig.add_subplot(1, 1, 1)
# For loop that plots the data points and centroids.
# k will range from 0-3, which will match the possible clusters that each
# data point is in.
for k, col in zip(range(len([[4,4], [-2, -1], [2, -3], [1, 1]])), colors):
# Create a list of all data points, where the data poitns that are
# in the cluster (ex. cluster 0) are labeled as true, else they are
# labeled as false.
my_members = (k_means_labels == k)
# Define the centroid, or cluster center.
cluster_center = k_means_cluster_centers[k]
# Plots the datapoints with color col.
ax.plot(X[my_members, 0], X[my_members, 1], 'w', markerfacecolor=col, marker='.')
# Plots the centroids with specified color, but with a darker outline
ax.plot(cluster_center[0], cluster_center[1], 'o', markerfacecolor=col, markeredgecolor='k', markersize=6)
# Title of the plot
ax.set_title('KMeans')
# Remove x-axis ticks
ax.set_xticks(())
# Remove y-axis ticks
ax.set_yticks(())
# Show the plot
plt.show()
# -
# ## Practice
# Try to cluster the above dataset into 3 clusters.
# Notice: do not generate data again, use the same dataset as above.
# +
# write your code here
# -
# Double-click __here__ for the solution.
#
# <!-- Your answer is below:
#
# k_means3 = KMeans(init = "k-means++", n_clusters = 3, n_init = 12)
# k_means3.fit(X)
# fig = plt.figure(figsize=(6, 4))
# colors = plt.cm.Spectral(np.linspace(0, 1, len(set(k_means3.labels_))))
# ax = fig.add_subplot(1, 1, 1)
# for k, col in zip(range(len(k_means3.cluster_centers_)), colors):
# my_members = (k_means3.labels_ == k)
# cluster_center = k_means3.cluster_centers_[k]
# ax.plot(X[my_members, 0], X[my_members, 1], 'w', markerfacecolor=col, marker='.')
# ax.plot(cluster_center[0], cluster_center[1], 'o', markerfacecolor=col, markeredgecolor='k', markersize=6)
# plt.show()
#
#
# -->
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# # Customer Segmentation with K-Means
# Imagine that you have a customer dataset, and you need to apply customer segmentation on this historical data.
# Customer segmentation is the practice of partitioning a customer base into groups of individuals that have similar characteristics. It is a significant strategy as a business can target these specific groups of customers and effectively allocate marketing resources. For example, one group might contain customers who are high-profit and low-risk, that is, more likely to purchase products, or subscribe for a service. A business task is to retaining those customers. Another group might include customers from non-profit organizations. And so on.
#
# Lets download the dataset. To download the data, we will use **`!wget`**. To download the data, we will use `!wget` to download it from IBM Object Storage.
# __Did you know?__ When it comes to Machine Learning, you will likely be working with large datasets. As a business, where can you host your data? IBM is offering a unique opportunity for businesses, with 10 Tb of IBM Cloud Object Storage: [Sign up now for free](http://cocl.us/ML0101EN-IBM-Offer-CC)
# + button=false new_sheet=false run_control={"read_only": false}
# !wget -O Cust_Segmentation.csv https://s3-api.us-geo.objectstorage.softlayer.net/cf-courses-data/CognitiveClass/ML0101ENv3/labs/Cust_Segmentation.csv
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Load Data From CSV File
# Before you can work with the data, you must use the URL to get the Cust_Segmentation.csv.
# + button=false new_sheet=false run_control={"read_only": false}
import pandas as pd
cust_df = pd.read_csv("Cust_Segmentation.csv")
cust_df.head()
# -
# ### Pre-processing
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# As you can see, __Address__ in this dataset is a categorical variable. k-means algorithm isn't directly applicable to categorical variables because Euclidean distance function isn't really meaningful for discrete variables. So, lets drop this feature and run clustering.
# + button=false new_sheet=false run_control={"read_only": false}
df = cust_df.drop('Address', axis=1)
df.head()
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# #### Normalizing over the standard deviation
# Now let's normalize the dataset. But why do we need normalization in the first place? Normalization is a statistical method that helps mathematical-based algorithms to interpret features with different magnitudes and distributions equally. We use __tandardScaler()__ to normalize our dataset.
# + button=false new_sheet=false run_control={"read_only": false}
from sklearn.preprocessing import StandardScaler
X = df.values[:,1:]
X = np.nan_to_num(X)
Clus_dataSet = StandardScaler().fit_transform(X)
Clus_dataSet
# -
# ### Modeling
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# In our example (if we didn't have access to the k-means algorithm), it would be the same as guessing that each customer group would have certain age, income, education, etc, with multiple tests and experiments. However, using the K-means clustering we can do all this process much easier.
#
# Lets apply k-means on our dataset, and take look at cluster labels.
# + button=false new_sheet=false run_control={"read_only": false}
clusterNum = 3
k_means = KMeans(init = "k-means++", n_clusters = clusterNum, n_init = 12)
k_means.fit(X)
labels = k_means.labels_
print(labels)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ### Insights
# We assign the labels to each row in dataframe.
# + button=false new_sheet=false run_control={"read_only": false}
df["Clus_km"] = labels
df.head(5)
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# We can easily check the centroid values by averaging the features in each cluster.
# + button=false new_sheet=false run_control={"read_only": false}
df.groupby('Clus_km').mean()
# -
# Now, lets look at the distribution of customers based on their age and income:
# + button=false new_sheet=false run_control={"read_only": false}
area = np.pi * ( X[:, 1])**2
plt.scatter(X[:, 0], X[:, 3], s=area, c=labels.astype(np.float), alpha=0.5)
plt.xlabel('Age', fontsize=18)
plt.ylabel('Income', fontsize=16)
plt.show()
# +
from mpl_toolkits.mplot3d import Axes3D
fig = plt.figure(1, figsize=(8, 6))
plt.clf()
ax = Axes3D(fig, rect=[0, 0, .95, 1], elev=48, azim=134)
plt.cla()
# plt.ylabel('Age', fontsize=18)
# plt.xlabel('Income', fontsize=16)
# plt.zlabel('Education', fontsize=16)
ax.set_xlabel('Education')
ax.set_ylabel('Age')
ax.set_zlabel('Income')
ax.scatter(X[:, 1], X[:, 0], X[:, 3], c= labels.astype(np.float))
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# k-means will partition your customers into mutually exclusive groups, for example, into 3 clusters. The customers in each cluster are similar to each other demographically.
# Now we can create a profile for each group, considering the common characteristics of each cluster.
# For example, the 3 clusters can be:
#
# - AFFLUENT, EDUCATED AND OLD AGED
# - MIDDLE AGED AND MIDDLE INCOME
# - YOUNG AND LOW INCOME
# + [markdown] button=false new_sheet=false run_control={"read_only": false}
# ## Want to learn more?
#
# IBM SPSS Modeler is a comprehensive analytics platform that has many machine learning algorithms. It has been designed to bring predictive intelligence to decisions made by individuals, by groups, by systems – by your enterprise as a whole. A free trial is available through this course, available here: [SPSS Modeler](http://cocl.us/ML0101EN-SPSSModeler).
#
# Also, you can use Watson Studio to run these notebooks faster with bigger datasets. Watson Studio is IBM's leading cloud solution for data scientists, built by data scientists. With Jupyter notebooks, RStudio, Apache Spark and popular libraries pre-packaged in the cloud, Watson Studio enables data scientists to collaborate on their projects without having to install anything. Join the fast-growing community of Watson Studio users today with a free account at [Watson Studio](https://cocl.us/ML0101EN_DSX)
#
# ### Thanks for completing this lesson!
#
# Notebook created by: <a href = "https://ca.linkedin.com/in/saeedaghabozorgi"><NAME></a>
#
# <hr>
# Copyright © 2018 [Cognitive Class](https://cocl.us/DX0108EN_CC). This notebook and its source code are released under the terms of the [MIT License](https://bigdatauniversity.com/mit-license/).
| ML0101EN-Clus-K-Means-Customer-Seg-py-v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# %matplotlib inline
#
# Cross-validation: some gotchas
# ===============================
#
# Cross-validation is the ubiquitous test of a machine learning model. Yet
# many things can go wrong.
#
#
#
# The uncertainty of measured accuracy
# ------------------------------------
#
# The first thing to have in mind is that the results of a
# cross-validation are noisy estimate of the real prediction accuracy
#
# Let us create a simple artificial data
#
#
from sklearn import datasets, discriminant_analysis
import numpy as np
np.random.seed(0)
data, target = datasets.make_blobs(centers=[(0, 0), (0, 1)])
classifier = discriminant_analysis.LinearDiscriminantAnalysis()
# One cross-validation gives spread out measures
#
#
from sklearn.model_selection import cross_val_score
print(cross_val_score(classifier, data, target))
# What if we try different random shuffles of the data?
#
#
from sklearn import utils
for _ in range(10):
data, target = utils.shuffle(data, target)
print(cross_val_score(classifier, data, target))
# This should not be surprising: if the lassification rate is p, the
# observed distribution of correct classifications on a set of size
# follows a binomial distribution
#
#
from scipy import stats
n = len(data)
distrib = stats.binom(n=n, p=.7)
# We can plot it:
#
#
from matplotlib import pyplot as plt
plt.figure(figsize=(6, 3))
plt.plot(np.linspace(0, 1, n), distrib.pmf(np.arange(0, n)))
# It is wide, because there are not that many samples to mesure the error
# upon: iris is a small dataset
#
# We can look at the interval in which 95% of the observed accuracy lies
# for different sample sizes
#
#
for n in [100, 1000, 10000, 100000]:
distrib = stats.binom(n, .7)
interval = (distrib.isf(.025) - distrib.isf(.975)) / n
print("Size: {0: 7} | interval: {1}%".format(n, 100 * interval))
# At 100 000 samples, 5% of the observed classification accuracy still
# fall more than .5% away of the true rate
#
# **Keep in mind that cross-val is a noisy measure**
#
# Importantly, the variance across folds is not a good measure of this
# error, as the different data folds are not independent. For instance,
# doing many random splits will can reduce the variance arbitrarily, but
# does not provide actually new data points
#
#
# Confounding effects and non independence
# -----------------------------------------
#
#
# Measuring baselines and chance
# -------------------------------
#
# Because of class imbalances, or confounding effects, it is easy to get
# it wrong it terms of what constitutes chances. There are two approaches
# to measure peformances of baselines or chance:
#
# **DummyClassifier** The dummy classifier:
# :class:`sklearn.dummy.DummyClassifier`, with different strategies to
# provide simple baselines
#
#
from sklearn.dummy import DummyClassifier
dummy = DummyClassifier(strategy="stratified")
print(cross_val_score(dummy, data, target))
# **Chance level** To measure actual chance, the most robust approach is
# to use permutations:
# :func:`sklearn.model_selection.permutation_test_score`, which is used
# as cross_val_score
#
#
from sklearn.model_selection import permutation_test_score
score, permuted_scores, p_value = permutation_test_score(classifier, data, target)
print("Classifier score: {0},\np value: {1}\nPermutation scores {2}"
.format(score, p_value, permuted_scores))
| interpreting_ml_tuto/cross_validation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Creole OvineHDSNPList
# the `CREOLE_INIA_UY.zip` dataset is encode in *A/B* format and provide a `OvineHDSNPList.txt` file. Does this file keeps the same information of a manifest file? Is the information consistent for the SNPs in common with *50K* chip?
# +
import re
import csv
import itertools
import collections
from src.features.smarterdb import Dataset, global_connection, VariantSheep
from src.features.utils import sanitize
conn = global_connection()
# -
dataset = Dataset.objects(file="CREOLE_INIA_UY.zip").get()
snplist = dataset.working_dir / "OvineHDSNPList.txt"
# try to detect if the alleles I have in file list are similar to ones I have in database. Print a few samples
with open(snplist) as handle:
reader = csv.reader(handle, delimiter="\t")
header = next(reader)
header = [sanitize(column) for column in header]
print(header)
SnpLine = collections.namedtuple("SnpLine", header)
counter = 0
for record in reader:
# fix elements
record[header.index('snp')] = re.sub(
r'[\[\]]',
"",
record[header.index('snp')])
snpline = SnpLine._make(record)
qs = VariantSheep.objects(name=snpline.name)
if qs.count() > 0:
variant = qs.get()
location = variant.get_location(version="Oar_v3.1", imported_from="manifest")
if snpline.snp != location.illumina or snpline.customer_strand != location.strand or snpline.ilmn_strand != location.illumina_strand:
# if snpline.customer_strand != location.strand:
counter += 1
print(f"{snpline.name} {snpline.chr}:{snpline.position} {location.chrom}:{location.position} {snpline.snp} {location.illumina} {snpline.customer_strand} {location.strand} {snpline.ilmn_strand} {location.illumina_strand}")
if counter > 20:
break
# The `customer_strand` and the `location.strand` are always equal (I suppose since they are the same *OARv3*). However `ilmn_strand` and `location.illumina_strand` can be different. In such case the allele is **reverse complemented**
# ## About coding convenction in general
# Relying on what I see in database, if `illumina_strand` is in *BOT*, I need to reverse complement the allele to have an `illumina_top`. If `strand` is in *BOT*, the `allele` (not the *illumina*) seems to be the `illumina_forward` or `illumina` in reverse order (depending if it comes from *SNPchimp* or *manifest*)
| notebooks/exploratory/0.6.1-bunop-about_creole_snplist.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernel_info:
# name: python3
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ### Note
# * Instructions have been included for each segment. You do not have to follow them exactly, but they are included to help you think through the steps.
# +
# Dependencies and Setup
import pandas as pd
import numpy as np
# File to Load (Remember to Change These)
file_to_load = "Resources/purchase_data.csv"
# Read Purchasing File and store into Pandas data frame
purchase_data = pd.read_csv(file_to_load)
# -
# ## Player Count
# * Display the total number of players
#
# +
Total_Players_in_Game = len(purchase_data["SN"].value_counts())
Total_Players_in_Game = pd.DataFrame(purchase_data)
Total_Players_in_Game
# +
Total_Players_in_Game = len(purchase_data["SN"].value_counts())
Total_Players_in_Game
# -
# ## Purchasing Analysis (Total)
# * Run basic calculations to obtain number of unique items, average price, etc.
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
#
# +
#the number of unique items
number_of_unique_items = len((purchase_data["Item ID"]).unique())
#the average price of said items
average_price = (purchase_data["Price"]).mean()
#the Quantity of Purchases made
number_of_purchases = (purchase_data["Purchase ID"]).count()
#total revenue made in game
total_revenue = (purchase_data["Price"]).sum()
Price_of_item_df = pd.DataFrame({"Number of Unique Items":[number_of_unique_items],
"Average Price":[average_price],
"Number of Purchases": [number_of_purchases],
"Total Revenue": [total_revenue]})
Price_of_item_df.style.format({'Average Price':"${:,.2f}",
'Total Revenue': '${:,.2f}'})
# +
# One third complete
# -
# ## Gender Demographics
# * Percentage and Count of Male Players
#
#
# * Percentage and Count of Female Players
#
#
# * Percentage and Count of Other / Non-Disclosed
#
#
#
# +
#here is the percentages of players by gender
Male_players = purchase_data.loc[purchase_data["Gender"] == "Male"]
Male_count = len(Male_players["SN"].unique())
Male_percent = "{:.2f}%".format(Male_count / Total_Players_in_Game * 100)
Female_players = purchase_data.loc[purchase_data["Gender"] == "Female"]
Female_count = len(Female_players["SN"].unique())
Female_percent = "{:.2f}%".format(Female_count / Total_Players_in_Game * 100)
#below is Data of Genders that does fall into the above categories
other_player_types = purchase_data.loc[purchase_data["Gender"] == "Other / Non-Disclosed"]
other_count = len(other_player_types["SN"].unique())
other_player_types_percent = "{:.2f}%".format(other_count / Total_Players_in_Game * 100)
Gender_Demographics_Table = pd.DataFrame([{
"Gender": "Male", "Total Count": Male_count,
"Percentage of Players": Male_percent},
{"Gender": "Female", "Total Count": Female_count,
"Percentage of Players": Female_percent},
{"Gender": "Other / Non-Disclosed", "Total Count": other_player_types_percent,
"Percentage of Players": other_player_types_percent
}], columns=["Gender", "Total Count", "Percentage of Players"])
Gender_Demographics_Table = Gender_Demographics_Table.set_index("Gender")
Gender_Demographics_Table.index.name = None
Gender_Demographics_Table
# -
#
# ## Purchasing Analysis (Gender)
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. by gender
#
#
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
#Purchase count by those made by Males
Male_Purchase_made_Count = len(Male_players)
#Purchase count by those made by Females
Female_Purchases_Count = len(Female_players)
#purchase from others
OtherKinds_Purchase_Count = len(other_player_types)
#Average Purchase Price as by gender
AvgPrice_male =round((Male_players["Price"].sum())/len(Male_players["Price"]),2)
AvgPrice_Female= round((Female_players["Price"].sum())/len(Female_players["Price"]),2)
AvgPrice_Others= round((other_player_types["Price"].sum())/len(other_player_types["Price"]),2)
#Total Purchases Values
TotalPurchase_M = round(Male_players['Price'].sum(),2)
TotalPurchase_F = round(Female_players['Price'].sum(),2)
TotalPurchase_O = round(other_player_types['Price'].sum(),2)
# Normalised Totals
# male/female/Other
Male_totals = round((TotalPurchase_M/Male_Purchase_made_Count), 2)
Female_totals = round((TotalPurchase_F/Female_Purchases_Count), 2)
Other_totals = round((TotalPurchase_O/OtherKinds_Purchase_Count), 2)
Third_Table_clean = {"Purchase Count":[Male_Purchase_made_Count, Female_Purchases_Count, OtherKinds_Purchase_Count],
"Gender":["Male","Female","Other"],
"Average Purchase Price":[AvgPrice_male, AvgPrice_Female, AvgPrice_Others],
"Total Purchase Value":[TotalPurchase_M, TotalPurchase_F, TotalPurchase_O],
"Normalized Totals":[Male_totals, Female_totals, Other_totals]}
Third_Table_cleaner = pd.DataFrame(Third_Table_clean)
Third_Table_cleaner = Third_Table_cleaner.set_index('Gender')
Third_Table_cleaner= Third_Table_cleaner[['Purchase Count','Average Purchase Price','Total Purchase Value','Normalized Totals']]
Third_Table_cleaner
# -
# ## Age Demographics
# * Establish bins for ages
#
#
# * Categorize the existing players using the age bins. Hint: use pd.cut()
#
#
# * Calculate the numbers and percentages by age group
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: round the percentage column to two decimal points
#
#
# * Display Age Demographics Table
#
# +
# Create the bins in which Data will be held
# Bins are "under 10s", "10-14", "15-19", "20-24", "25-29", "30-34", "35-39", and "40+"
bins_Ages = [0,10,14,19,24,29,34,39,46]
Ages_labels = ["<10","10-14","15-19","20-24","25-29","30-34","35-39","40+"]
Plyr_ages = pd.cut(purchase_data['Age'],bins_Ages,labels= Ages_labels)
Plyr_ages
age_of_Heroes = purchase_data.groupby('Age')
total_count_age = age_of_Heroes["SN"].nunique()
percentage_age = (total_count_age/Total_Players_in_Game) * 100
percentage_age
age_demographics = pd.DataFrame({"Total Count": total_count_age, "Percentage of Players": percentage_age})
age_demographics.style.format({"Percentage of Players":"{: ,.2f}"})
# -
# ## Purchasing Analysis (Age)
# * Bin the purchase_data data frame by age
#
#
# * Run basic calculations to obtain purchase count, avg. purchase price, avg. purchase total per person etc. in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display the summary data frame
# +
bins_Ages2 = [0,10,14,19,24,29,34,39,46]
Ages_labels2 = ["<10","10-14","15-19","20-24","25-29","30-34","35-39","40+"]
Plyr_ages2 = pd.cut(purchase_data['Age'],bins_Ages,labels= Ages_labels)
Plyr_ages2
purchase_count_age = age_of_Heroes["Purchase ID"].count()
avg_purchase_price_age = age_of_Heroes["Price"].mean()
total_purchase_value = age_of_Heroes["Price"].sum()
avg_purchase_per_person_age = total_purchase_value/total_count_age
age_demographics = pd.DataFrame({"Purchase Count": purchase_count_age,
"Average Purchase Price": avg_purchase_price_age,
"Total Purchase Value":total_purchase_value,
"Average Purchase Total per Person": avg_purchase_per_person_age})
age_demographics.index.name = None
age_demographics.style.format({"Average Purchase Price":"${:,.2f}",
"Total Purchase Value":"${:,.2f}",
"Average Purchase Total per Person":"${:,.2f}"})
# -
# ## Top Spenders
# * Run basic calculations to obtain the results in the table below
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the total purchase value column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
theBallers = purchase_data.groupby("SN")
spender_purchase_count = theBallers["Purchase ID"].count()
theAverages = round(theBallers['Price'].mean(),2)
Total = theBallers['Price'].sum()
ScreenName = theBallers["SN"].unique()
TopSpenders = {"SN":ScreenName,"Purchase Count":spender_purchase_count,
"Average Purchase Price":theAverages,"Total Purchase Value":Total}
The_Top_Spender= pd.DataFrame(TopSpenders)
The_Top_Spender= The_Top_Spender.set_index('SN')
The_Top_Spender = The_Top_Spender.sort_values("Total Purchase Value",ascending=False)
The_Top_Spender = The_Top_Spender[['Purchase Count', 'Average Purchase Price', 'Total Purchase Value']]
The_Top_Spender.iloc[:5]
# -
# ## Most Popular Items
# * Retrieve the Item ID, Item Name, and Item Price columns
#
#
# * Group by Item ID and Item Name. Perform calculations to obtain purchase count, average item price, and total purchase value
#
#
# * Create a summary data frame to hold the results
#
#
# * Sort the purchase count column in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the summary data frame
#
#
# +
Total_Items_in_Game = purchase_data["Item ID"].value_counts()
Total_Items_in_Game = pd.DataFrame(purchase_data)
Total_Items_in_Game
items_purchased_count = purchase_data["Price"].count()
popular_GameItems = Total_Items_in_Game.groupby(["Item ID","Item Name"])
items_price = popular_GameItems["Price"].sum()
item_purchase_value = items_price / items_purchased_count
most_popular_items = pd.DataFrame({
"Purchase Count": items_purchased_count,
"Item Price": items_price,
"Total Purchase Value": items_price
})
popular_items_formatted = most_popular_items.sort_values(["Purchase Count"], ascending=False).head()
popular_items_formatted["Item Price"] = popular_items_formatted["Item Price"].astype(float).map("${:,.2f}".format)
popular_items_formatted["Total Purchase Value"] = popular_items_formatted["Total Purchase Value"].astype(float).map("${:,.2f}".format)
popular_items_formatted
# -
# ## Most Profitable Items
# * Sort the above table by total purchase value in descending order
#
#
# * Optional: give the displayed data cleaner formatting
#
#
# * Display a preview of the data frame
#
#
| HeroesOfPymoli_starter.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + tags=[]
# !pip install robin_stocks
import robin_stocks.robinhood as r
import pandas as pd
import numpy as np
import math as m
import matplotlib.pyplot as plt
# -
# Library: [robin_stocks](https://readthedocs.org/projects/robin-stocks/downloads/pdf/latest/)
# ## Authentication and Login
login = r.login('<EMAIL>','<PASSWORD>',store_session=True)
# + [markdown] tags=[]
# ## List of Cryptocurrencies available
# -
crypto_info=r.crypto.get_crypto_currency_pairs()
crypto_list=[]
for i in range(len(crypto_info)):
crypto_list.append(crypto_info[i]['asset_currency']['code'])
# ## Read in Portfolio
df_portfolio=pd.read_csv('portfolio.csv')
df_portfolio.set_index('Parameters',inplace=True)
columns_list=df_portfolio.columns.tolist()
columns_dict={}
for stock in columns_list:
stock_str_rep=stock.replace('-','.')
columns_dict[stock]=stock_str_rep
df_portfolio.rename(columns=columns_dict,inplace=True)
df_portfolio.sort_values(by='weight',axis=1,ascending=False)
port_stocks=df_portfolio.columns.tolist()
df_suggested_equity=df_portfolio.loc['suggested_investment',:]
df_suggested_equity.transpose().plot(kind='bar',figsize=(12,8));
total_port_equity=df_suggested_equity.sum()
print('Total suggested equity value is: {}'.format(total_port_equity))
# ## Print total stocks value
stocks_dict=r.account.build_holdings()
#print('Stocks are: {}'.format(stocks_dict))
tickers=stocks_dict.keys()
ticker_list=list(tickers)
for ticker in ticker_list:
ticker.replace('-','.')
print(ticker_list)
###### Get crypto positions #####
crypto_holdings=r.get_crypto_positions()
print(float(crypto_holdings[3]['quantity']))
price_list=[float(stocks_dict[ticker]['equity']) for ticker in ticker_list]
for i in range(len(crypto_holdings)):
if float(crypto_holdings[i]['quantity'])>0:
crypto_ticker=crypto_holdings[i]['currency']['code'].replace('-USD','')
crypto_qty=float(crypto_holdings[i]['quantity'])
crypto_mark_price=float(r.get_crypto_quote(symbol=crypto_ticker,info='mark_price'))
crypto_price=crypto_qty*crypto_mark_price
price_list.append(crypto_price)
ticker_list.append('{}.USD'.format(crypto_holdings[i]['currency']['code']))
######## Equity of stocks #################
df_allstock_equity=pd.DataFrame(index=['Price'],columns=ticker_list);
df_allstock_equity.loc['Price',:]=price_list;
allstock_equity_cols=df_allstock_equity.columns.tolist()
df_allstock_equity
# ## Stocks common to portfolio and investment
comm_stocks=list(set(port_stocks).intersection(set(ticker_list)))
print('Stocks common to recommended and actual portfolio: ',comm_stocks)
port_stocks=[stock.replace('-','.') for stock in port_stocks]
# ## Stocks not common
# ### Stock in recommended portfolio but not in actual portfolio
not_in_ticker_list=[stock for stock in port_stocks if stock not in ticker_list]
not_in_ticker_list
# ### Stock in actual portfolio but not in recommended portfolio
not_in_port_stocks=[ticker for ticker in ticker_list if ticker not in port_stocks]
not_in_port_stocks
# ## Share Price
share_price={}
share_price_values=[float(stocks_dict[ticker]['price']) for ticker in tickers]
for i in range(len(tickers)):
ticker=ticker_list[i]
share_price[ticker]=share_price_values[i]
share_price=pd.DataFrame(share_price,index=['price'])
share_price
# ## Share Quantity
share_quantity={}
share_quantities=[float(stocks_dict[ticker]['quantity']) for ticker in tickers]
for i in range(len(tickers)):
ticker=ticker_list[i]
share_quantity[ticker]=share_quantities[i]
share_quantity=pd.DataFrame(share_quantity,index=['quantity'])
share_quantity
# ## Total Equity
# +
equity={}
total_equity=0
for i in range(len(ticker_list)):
ticker=ticker_list[i]
equity[ticker]=price_list[i]
total_equity=total_equity+equity[ticker]
print('Total Stocks equity value is: {}'.format(total_equity))
equity_comm={}
total_comm_equity=df_allstock_equity.loc['Price',comm_stocks].sum()
print('Total Common Stocks equity value is: {}'.format(total_comm_equity))
df_equity=pd.DataFrame.from_dict(data=equity,orient='index',columns=['Equity'])
df_plot=df_equity.sort_values(by='Equity',axis=0,ascending=False)
if df_plot.shape[0]>0:
df_plot.plot(kind='bar',figsize=(12,8))
else:
print('no data to plot')
df_equity=df_equity.transpose()
df_equity.sort_values(by='Equity',axis=1,ascending=False)
# + [markdown] jp-MarkdownHeadingCollapsed=true tags=[]
# ## Equity Change
# +
total_equity=np.sum(np.array(price_list))
if len(ticker_list)>0:
equity_change={}
total_equity_change=0
equity_change_percent={}
for i in range(len(ticker_list)):
stock=ticker_list[i]
if stock in list(stocks_dict.keys()):
equity_change[stock]=float(stocks_dict[stock]['equity_change'])
equity_change_percent[stock]=float(stocks_dict[stock]['percent_change'])
else:
equity_change[stock]=df_equity.loc['Equity',stock]-df_suggested_equity[stock]
equity_change_percent[stock]=equity_change[stock]/100
total_equity_change=total_equity_change+equity_change[stock]
equity_change_percent=total_equity_change/total_equity*100
df_equity_change=pd.DataFrame.from_dict(data=equity_change,orient='index',columns=['Equity_Change']).transpose()
df_equity_change_ratio=pd.DataFrame(df_equity_change.
div(total_equity))
df_equity_change_percent=df_equity_change_ratio*100
df_equity_change_ratio.rename(index={'Equity_Change':'Equity_Change_Ratio'},inplace=True)
df_equity_change_percent.rename(index={'Equity_Change':'Equity_Change_Percent'},inplace=True)
print('Total equity change value is: {}'.format(total_equity_change))
print('Total Change Percent is: {}'.format(equity_change_percent))
df_equity_change.sort_values(by='Equity_Change',axis=1,ascending=False).transpose().plot(kind='bar',figsize=(18,6));
print(df_equity_change.sort_values(by='Equity_Change',axis=1,ascending=False))
df_equity_change_percent.sort_values(by='Equity_Change_Percent',axis=1,ascending=False).transpose().plot(kind='bar',figsize=(18,6));
if len(comm_stocks)>0:
equity_comm_change={}
equity_comm_change_percent={}
total_equity_comm_change=0
for i in range(len(comm_stocks)):
comm_stock=comm_stocks[i]
if comm_stock in list(stocks_dict.keys()):
equity_comm_change[comm_stock]=float(stocks_dict[comm_stock]['equity_change'])
equity_comm_change_percent[comm_stock]=float(stocks_dict[comm_stock]['percent_change'])
else:
equity_comm_change[comm_stock]=df_equity.loc['Equity',comm_stock]-df_suggested_equity[comm_stock]
equity_comm_change_percent[comm_stock]=equity_comm_change[comm_stock]/100
total_equity_comm_change=total_equity_comm_change+equity_comm_change[comm_stock]
print('Total Common equity change value is: {}'.format(total_equity_comm_change))
total_equity_comm_change_percent=total_equity_comm_change/total_port_equity*100
print('Total Common Change Percent is: {}'.format(total_equity_comm_change_percent))
df_equity_comm_change=pd.DataFrame.from_dict(data=equity_comm_change,orient='index',columns=['Equity_Comm_Change']).transpose()
df_equity_comm_change_percent=pd.DataFrame.from_dict(data=equity_comm_change_percent,orient='index',columns=['Equity_Comm_Change_Percent']).transpose()
############################ Plots #####################################
# df_equity_comm_change.sort_values(by='Equity_Comm_Change',axis=1,ascending=False).transpose().plot(kind='bar',figsize=(18,6));
# print(df_equity_comm_change.sort_values(by='Equity_Comm_Change',axis=1,ascending=False))
# df_equity_comm_change_percent.sort_values(by='Equity_Comm_Change_Percent',axis=1,ascending=False).transpose().plot(kind='bar',figsize=(18,6));
else:
print('no common stocks')
else:
print('no positions')
# -
# ## Update Stop Loss
# ### Stop Loss Sell Limit Price
# + active=""
# stop_loss_sell_limit_price={}
# if len(comm_stocks)>0:
# for i in range(len(comm_stocks)):
# stock=comm_stocks[i]
# try:
# stop_loss_sell_diff=df_portfolio.loc['price_bound',stock]
# limit_price=float(share_price[stock]-stop_loss_sell_diff)
# if limit_price<0:
# stop_loss_sell_limit_price[stock]=0
# else:
# stop_loss_sell_limit_price[stock]=limit_price
# except:
# print(stock)
# else:
# print('no common stocks')
# stop_loss_sell_limit_price
# -
# ### Cancel Existing Stock Orders
# + active=""
# r.get_all_open_stock_orders()
# + active=""
# r.orders.cancel_all_stock_orders();
# -
# ### Update Stop Loss Sell orders for all stocks
# + tags=[] active=""
# stop_loss_sell_info={}
# if len(comm_stocks)==0:
# print('no common stocks to update stop loss')
# else:
# for i in range(len(comm_stocks)):
# comm_stock=comm_stocks[i]
# comm_stock_modified=comm_stock.replace('-','.')
# sell_quantity=m.floor(share_quantity[comm_stock])
# stop_price=stop_loss_sell_limit_price[comm_stock]
# stop_loss_sell_info[comm_stock]=r.orders.order_sell_stop_loss(symbol=comm_stock_modified,
# quantity=sell_quantity,stopPrice=stop_price,timeInForce='gtc')
# stop_loss_sell_info
# -
# ## Sell and Buy Stocks
# ### Sell all portfolio stocks
# + active=""
# for i in range(len(comm_stocks)):
# comm_stock=comm_stocks[i]
# qty=share_quantity.loc['quantity',comm_stock]
# sell_order=r.orders.order_sell_fractional_by_quantity(symbol=comm_stock,
# quantity=qty,
# timeInForce='gfd')
# if not sell_order['reject_reason']==None:
# print(ticker)
# + [markdown] tags=[]
# ### Sell all stocks
# + tags=[] active=""
# stocks_dict=r.account.build_holdings()
# #print('Stocks are: {}'.format(stocks_dict))
# tickers=stocks_dict.keys()
# ticker_list=list(tickers)
# crypto_ticker_list=[]
# crypto_holdings_temp=crypto_holdings.copy();
# for i in range(len(crypto_holdings_temp)):
# qty=float(crypto_holdings[i]['cost_bases'][0]['direct_quantity'])
# crypto_ticker_list.append(crypto_holdings[i]['currency']['code'])
# if qty>0:
# crypto=crypto_ticker_list[i]
# sell_order=r.orders.order_sell_crypto_by_quantity(symbol=crypto,
# quantity=qty,
# timeInForce='gtc')
# print('Crypto sell orders :',sell_order)
# else:
# print('no crypto to sell')
#
# if len(ticker_list)>0:
# for i in range(len(ticker_list)):
# ticker=ticker_list[i]
# qty=share_quantity.loc['quantity',ticker]
# sell_order=r.orders.order_sell_fractional_by_quantity(symbol=ticker,
# quantity=qty,
# timeInForce='gfd')
# print('stocks_sell_prders :',sell_order)
# else:
# print('no stocks to sell')
# + [markdown] tags=[]
# ### Buy Portfolio Stocks
# + tags=[] active=""
# stocks_dict=r.account.build_holdings()
# #print('Stocks are: {}'.format(stocks_dict))
# tickers=stocks_dict.keys()
# ticker_list=list(tickers)
# ###### Get crypto positions #####
# crypto_holdings=r.get_crypto_positions()
# ### Add crypto position tickers to ticker_list ###
# for i in range(len(crypto_holdings)):
# qty=float(crypto_holdings[i]['cost_bases'][0]['direct_quantity'])
# if qty>0:
# ticker_list.append(crypto_holdings[i]['currency']['code'])
#
# for i in range(len(df_portfolio.columns)):
# ticker_suggested=df_portfolio.columns[i]
# ticker_suggested_modified=ticker_suggested.replace('-','.')
# ticker_suggested_modified=ticker_suggested_modified.replace('.USD','')
# equity=df_portfolio.loc['suggested_investment',ticker_suggested]
# if ticker_suggested_modified not in ticker_list:
# print('{}:{}'.format(ticker_suggested_modified,equity))
# if ticker_suggested_modified in crypto_list:
# buy_order=r.orders.order_buy_crypto_by_price(symbol=ticker_suggested_modified,amountInDollars=equity,
# )
# else:
# buy_order=r.orders.order_buy_fractional_by_price(symbol=ticker_suggested_modified,
# amountInDollars=equity,
# timeInForce='gfd')
# print(buy_order)
# #if not buy_order['reject_reason']==None:
# #print(ticker)
# -
# ####
| View_Stocks/view_robin_stocks_v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# +
import sys
sys.path.insert(0, '/global/common/software/lsst/common/miniconda/py3-4.2.12/lib/python3.6/site-packages')
## Note: if you use Python 2, comment the line above and uncomment the line below
#sys.path.insert(0, '/global/common/cori/contrib/lsst/apps/anaconda/py2-envs/DESCQA/lib/python2.7/site-packages')
# +
import numpy as np
import matplotlib.pyplot as plt
# %matplotlib inline
plt.rcParams['font.size'] = 12
from IPython.display import display
from GCR import GCRQuery
# -
import GCRCatalogs
print(GCRCatalogs.__version__)
# +
# using protoDC2_test here to bypass md5 check.
# using buzzard_test here because full buzzard is very big!
catalogs = ('protoDC2_test', 'buzzard_test')
gc_all = dict(zip(catalogs, (GCRCatalogs.load_catalog(c) for c in catalogs)))
# -
# ## Conditional Luminosity Function Tests
class ConditionalLuminosityFunction(object):
def __init__(self, band='r', magnitude_bins=None, mass_bins=None, z_bins=None, **kwargs):
possible_Mag_fields = ('Mag_true_{}_lsst_z0',
'Mag_true_{}_lsst_z01',
'Mag_true_{}_des_z0',
'Mag_true_{}_des_z01',
'Mag_true_{}_sdss_z0',
'Mag_true_{}_sdss_z01',
)
self.possible_Mag_fields = [f.format(band) for f in possible_Mag_fields]
self.band = band
self.magnitude_bins = magnitude_bins or np.linspace(-25, -18, 29)
self.mass_bins = mass_bins or np.logspace(12, 15, 5)
self.z_bins = z_bins or np.linspace(0, 0.5, 3)
self.n_magnitude_bins = len(self.magnitude_bins) - 1
self.n_mass_bins = len(self.mass_bins) - 1
self.n_z_bins = len(self.z_bins) - 1
self.dmag = self.magnitude_bins[1:] - self.magnitude_bins[:-1]
self.mag_center = (self.magnitude_bins[1:] + self.magnitude_bins[:-1])*0.5
self._other_kwargs = kwargs
def prepare_galaxy_catalog(self, gc):
quantities_needed = {'redshift_true', 'is_central', 'halo_mass'}
if gc.has_quantities(['truth/RHALO', 'truth/R200']):
gc.add_quantity_modifier('r_host', 'truth/RHALO', overwrite=True)
gc.add_quantity_modifier('r_vir', 'truth/R200', overwrite=True)
quantities_needed.add('r_host')
quantities_needed.add('r_vir')
try:
absolute_magnitude_field = gc.first_available(*self.possible_Mag_fields)
except ValueError:
return
quantities_needed.add(absolute_magnitude_field)
if not gc.has_quantities(quantities_needed):
return
return absolute_magnitude_field, quantities_needed
def run_validation_test(self, galaxy_catalog, catalog_name, base_output_dir=None):
prepared = self.prepare_galaxy_catalog(galaxy_catalog)
if prepared is None:
TestResult(skipped=True)
absolute_magnitude_field, quantities_needed = prepared
colnames = [absolute_magnitude_field, 'halo_mass', 'redshift_true']
bins = (self.magnitude_bins, self.mass_bins, self.z_bins)
hist_cen = np.zeros((self.n_magnitude_bins, self.n_mass_bins, self.n_z_bins))
hist_sat = np.zeros_like(hist_cen)
cen_query = GCRQuery('is_central')
sat_query = ~cen_query
if 'r_host' in quantities_needed and 'r_vir' in quantities_needed:
sat_query &= GCRQuery('r_host < r_vir')
for data in galaxy_catalog.get_quantities(quantities_needed, return_iterator=True):
cen_mask = cen_query.mask(data)
sat_mask = sat_query.mask(data)
data = np.stack((data[k] for k in colnames)).T
hist_cen += np.histogramdd(data[cen_mask], bins)[0]
hist_sat += np.histogramdd(data[sat_mask], bins)[0]
del data, cen_mask, sat_mask
halo_counts = hist_cen.sum(axis=0)
clf = dict()
clf['sat'] = hist_sat / halo_counts
clf['cen'] = hist_cen / halo_counts
clf['tot'] = clf['sat'] + clf['cen']
return clf
def make_plot(self, clf, name):
fig, ax = plt.subplots(self.n_mass_bins, self.n_z_bins, sharex=True, sharey=True, figsize=(12,10), dpi=100)
for i in range(self.n_z_bins):
for j in range(self.n_mass_bins):
ax_this = ax[j,i]
for k, ls in zip(('total', 'satellites', 'centrals'), ('-', ':', '--')):
ax_this.semilogy(self.mag_center, clf[k[:3]][:,j,i]/self.dmag, label=k, ls=ls)
ax_this.set_ylim(0.05, 50)
bins = self.mass_bins[j], self.mass_bins[j+1], self.z_bins[i], self.z_bins[i+1]
ax_this.text(-25, 10, '${:.1E}\\leq M <{:.1E}$\n${:g}\\leq z<{:g}$'.format(*bins))
ax_this.legend(loc='lower right', frameon=False, fontsize='medium')
ax = fig.add_subplot(111, frameon=False)
ax.tick_params(labelcolor='none', top='off', bottom='off', left='off', right='off')
ax.grid(False)
ax.set_ylabel(r'$\phi(M_{{{}}}\,|\,M_{{\rm vir}},z)\quad[{{\rm Mag}}^{{-1}}]$'.format(self.band))
ax.set_xlabel(r'$M_{{{}}}\quad[{{\rm Mag}}]$'.format(self.band))
ax.set_title(name)
fig.tight_layout()
display(fig)
plt.close(fig)
clf_test = ConditionalLuminosityFunction()
clf_all = dict()
for label, gc_this in gc_all.items():
clf_all[label] = clf_test.run_validation_test(gc_this, label)
for label, clf_this in clf_all.items():
clf_test.make_plot(clf_this, label)
| examples/CLF Test.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [default]
# language: python
# name: python3
# ---
# # Non-Linear Modulation Amplitude Measurements
#
#
# For NL-SIM we need to have a good way to measure, directly, the NL modulation amplitude. This task is very model dependent. The best way would be to use the fourier transform, but unfortunately the signal to noise and number of points is often too low to make the measurement. For instance if we wanted to nyquist sample a signal with a maximum frequency content of $k_{max}$ we would need a sampling inteval of $1/2 k_{max}$. The period of our pattern is such that we sample $2 \pi$ in $n$ points. So basically our frequency, $f$, is 1. To perfectly Nyquist sample our signal (considering a single harmonic) we need three points across our interval of $[0, 2\pi)$ as that gives the required sampling frequency of $\pi$. Each higher harmonic requires 2 more points. So for 15 points we can nyquist sample up to 7 harmonics. The number of points needed for $H$ harmonics is $2H + 1$
#
# But we still have signal to noise concerns. The other option is to non-linearly fit the signal to a sum of sinusoids, but then you have to fit the phase, amplitude of each sinusoid plus an offset and frequency, so the number of points needed is $2H + 2$ to ensure that the system is not underdetermined.
#
# The final option is to use a variant of LPSVD which would independently fit the amplitude, frequency, phase and offset of each sinusoid. In this case we'd need $4H$ points.
# +
def cosine(x, amp, f, p, o):
'''
Utility function to fit nonlinearly
'''
return amp*np.cos(2*np.pi*f*(x-p))+o
def cosine_sat(x,*params):
o = params[-1]
p = params[-2]
f = params[-3]
amps = params[:len(params)-3]
to_return = zeros_like(x)
for i, amp in enumerate(amps):
#each amp corresponds to the ith harmonic
to_return += cosine(x,amp,f*(i+1),p,0)
return to_return+o
def calc_mod_sat(data, num_harms = 1, nphases = 24,periods = 2):
'''
Need to change this so that it:
- first tries to fit only the amplitude and phase
- if that doesn't work, estimate amp and only fit phase
- then do full fit
'''
#pull internal number of phases
#nphases = self.nphases
#only deal with finite data
#NOTE: could use masked wave here.
finite_args = np.isfinite(data)
data_fixed = data[finite_args]
popt = None
if len(data_fixed) > 4:
#we can't fit data with less than 4 points
#make x-wave
x = np.arange(nphases,dtype=data_fixed.dtype)[finite_args]
#make guesses
#amp of sine wave is sqrt(2) the standard deviation
g_a = np.sqrt(2)*(data_fixed.std())
#offset is mean
g_o = data_fixed.mean()
#frequency is such that `nphases` covers `periods`
g_f = periods/nphases
#guess of phase is from first data point (maybe mean of all?)
g_p = nan
i = 0
while not isfinite(g_p):
g_p = x[i]-np.arccos((data_fixed[i]-g_o)/g_a)/(2*np.pi*g_f)
i+=1
#make guess sequence
if num_harms == 1:
amps = [g_a]
if num_harms == 2:
# https://www.wolframalpha.com/input/?i=expand+1-cos(x)%5E4
amps = [-4/5*g_a, -1/5*g_a]
g_p+=3*pi/2
if num_harms == 3:
# https://www.wolframalpha.com/input/?i=expand+1-cos(x)%5E6
amps = [-15/22*g_a, -6/22*g_a,-1/22*g_a]
g_p+=3*pi/2
if num_harms == 4:
amps = [-56/93*g_a, -28/93*g_a,-8/93*g_a,-1/93*g_a]
g_p+=3*pi/2
pguess = amps + [g_f,g_p,g_o]
try:
popt,pcov = curve_fit(cosine_sat,x,data_fixed,p0=array(pguess))
except RuntimeError as e:
#if fit fails, put nan
print(e)
mod = np.nan
res = nan
except TypeError as e:
print(e)
print(data_fixed)
mod = np.nan
res = nan
else:
if len(popt)==4:
opt_a,opt_f,opt_p,opt_o = popt
opt_a = np.abs(opt_a)
#if any part of the fit is negative, mark as failure
if opt_o - opt_a < 0:
mod = np.nan
else:
#calc mod
mod = 2*opt_a/(opt_o+opt_a)
else:
mod = np.nan
res = (data_fixed-cosine_sat(x,*popt))**2
res = res.sum()
else:
mod = np.nan
res = nan
return popt, res, mod
# +
def cosine(x, amp, f, p, o):
'''
Utility function to fit nonlinearly
'''
return amp*np.cos(2*np.pi*f*(x-p))+o
def cosine_sat(x,*params):
o = params[-1]
p = params[-2]
f = params[-3]
amps = params[:len(params)-3]
to_return = zeros_like(x)
for i, amp in enumerate(amps):
#each amp corresponds to the ith harmonic
to_return += cosine(x,amp,f*(i+1),p,0)
return to_return+o
#Testing the function
x = linspace(0,23,1024)
#popt = array([ -9.23688107e+02, -2.15499865e+01, 1/12, 12*pi, 2.31774918e+03])
popt = array([ -56/128, -28/128, -8/128, -1/128, 1/12, 0, 93/128])
fig, ax = subplots(1,1,figsize=(12,12))
fit = cosine_sat(x,*popt)
ax.plot(x,fit,'r-', label = 'Cosine Sat Func')
amps = popt[:len(popt)-3]
# plot each harmonic seperately
for i, amp in enumerate(amps):
#each amp corresponds to the ith harmonic
o = popt[-1]
p = popt[-2]
f = popt[-3]
mod = 2*abs(amp)/(o+abs(amp))
ax.plot(x,cosine(x,amp,f*(i+1),p,-o*(amp/abs(amps).sum())),'--',label='Mod = {:.2f}\nAmp = {:.2f}'.format(mod, amp))
fit_direct = 1 - (cosine(x, 1/2, f, p , 1/2))**4
ax.plot(x, fit_direct,'k.', label= '$\cos(x)^4$')
ax.legend()
def calc_mod_sat(data, num_harms = 1, nphases = 30,periods = 2):
'''
Need to change this so that it:
- first tries to fit only the amplitude and phase
- if that doesn't work, estimate amp and only fit phase
- then do full fit
'''
#pull internal number of phases
#nphases = self.nphases
#only deal with finite data
#NOTE: could use masked wave here.
finite_args = np.isfinite(data)
data_fixed = data[finite_args]
popt = ones(4)*nan
pguess = ones(4)*nan
res = np.nan
if len(data_fixed) > 3+num_harms:
#we can't fit data if number of parameters exceeds number of points
#make x-wave
x = np.arange(nphases,dtype=data_fixed.dtype)[finite_args]
#make guess sequence
if num_harms == 1:
#make guesses
#amp of sine wave is sqrt(2) the standard deviation
g_a = np.sqrt(2)*(data_fixed.std())
#offset is mean
g_o = data_fixed.mean()
#frequency is such that `nphases` covers `periods`
g_f = periods/nphases
#guess of phase is from median, note that amp is made negative due to the saturation model.
g_p = median((x-np.arccos(-(data_fixed-g_o)/g_a)/(2*np.pi*g_f))[:nphases//periods])
amps = [1]
if num_harms == 2:
amps = [4/5, 1/5]
if num_harms == 3:
amps = [15/22, 6/22,1/22]
if num_harms == 4:
amps = [56/93, 28/93,8/93,1/93]
if num_harms > 1:
#first do the fit for num_harms 1
popt, res, pguess = calc_mod_sat(data, num_harms = 1, nphases = nphases,periods = periods)
g_a, g_f, g_p, g_o = popt
#if the fitted amp is positive that just means we need to adjust the phase
# and make the amp negative
if g_a > 0:
g_p += nphases/periods/2
g_a = -g_a
# for saturated data we expect the peaks to be negative, so shift phase and make amplitudes negative.
pguess = concatenate((array(amps)*g_a, [g_f, g_p, g_o]))
if isfinite(pguess.all()):
try:
popt,pcov = curve_fit(cosine_sat,x,data_fixed,p0=pguess)
except RuntimeError as e:
#if fit fails, put nan
pass
except TypeError as e:
print(e)
print(data_fixed)
else:
res = ((data_fixed-cosine_sat(x, *popt))**2).sum()
return popt, res, pguess
def find_best_num_harms(y, max_harm = 4, verbose =False):
'''
A function to find the best number of harmonics to fit the data
'''
# set up the stopping criterion
amps_valid = False
mods_valid = False
res_old = np.inf
popt_old = np.nan
# loop through having 2 to 4 harmonics, fitting each one
for i in list(range(2, max_harm+1))+[1]:
popt, res, pguess = calc_mod_sat(y, i)
if verbose:
print('pguess', pguess)
print('popt', popt)
# pull the offset
o = popt[-1]
# and the amps
amps = popt[:len(popt)-3]
# check to see if amps are positive
amps_pos = (amps < 0).all()
# check to see that the amplitudes are ordered such that the strength decreasing with increasing harmonics
amps_valid = amps_pos and (amps.argsort() == arange(len(amps))).all()
# check the modulation depths as well.
mods = 2*abs(amps)/(o+abs(amps))
#make sure modulation depths are between 1 and 0, inclusive
mods_valid = np.logical_and(mods <= 1.0, mods >= 0.0).all()
if res > res_old:
# if the residuals have increased, don't update
if verbose:
print('Residuals have increased')
# break
elif mods_valid and (amps_valid or len(popt) == 4):
# if the residuals have decreased and the parameter are valid, update
if verbose:
print('Residuals have decreased and popt valid, updating...')
res_old = res
popt_old = popt
else:
if verbose:
print('Residuals have decreased but popt invalid')
if verbose:
print('{:.3e}'.format(res_old))
if verbose:
print('Final residuals = {:.3e}'.format(res_old))
return popt_old, res_old, pguess
# -
# Probably the best thing to do is to have a function which calculates the saturation directly from a relevant model then you only have to fit:
# - amplitude
# - phase
# - frequency
# - saturation factor
# - offset
#
# For any saturation factor or number of harmonics
#
# The following model functions should be used: power law ($a x^{-b}$), exponential function ($a e^{-b x}$) and saturation ($a/(1 + x/b)$)
| notebooks/NL Modulation Amp Calc.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
from sklearn.linear_model import LinearRegression
import pandas as pd
import numpy as np
dfx = pd.read_csv("Train.csv")
dfxx = pd.read_csv("Test.csv")
# +
X = dfx.values
XT = dfxx.values
X_train = X[:,:-1]
Y_train = X[:,-1]
X_test = XT
# +
# 1 create an object
lr = LinearRegression(normalize=True)
# 2 Training
lr.fit(X_train, Y_train)
# 3 Output Parameters
print(lr.coef_)
print(lr.intercept_)
# -
Y_predict = lr.predict(X_test)
print(Y_predict)
print("Training Score %.4f" %lr.score(X_train, Y_train))
print("Testing score %.4f " %lr.score(X_test,Y_predict))
iD = []
for i in range(X_test.shape[0]):
iD.append(i)
print(iD)
ids = pd.DataFrame(iD)
ids.to_csv('./Pred/prediction3.csv' , index=True)
outcome = pd.DataFrame(Y_predict)
outcome.to_csv('./Pred/prediction3.csv' , index=True)
| Air pollution using Sklearn final.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mazette04/OOP-58002/blob/main/Operations_and_Expressions.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + [markdown] id="CVi6UK2IDx6r"
# Boolean Operators
# + colab={"base_uri": "https://localhost:8080/"} id="6EYcZG80D3Gz" outputId="9dad8b7b-d785-478e-f611-4f4465945341"
a = 7
b =6
print(10>9)
print(10<9)
print(a>b)
# + [markdown] id="DNPZJKWdEstg"
# bool() function
# + colab={"base_uri": "https://localhost:8080/"} id="lyKLgigfEzON" outputId="789469e2-5f7b-4b5e-8e69-f44a03730ad3"
print(bool("Maria"))
print(bool(1))
print(bool(0))
print(bool(None))
print(bool(False))
print(bool(True))
# + [markdown] id="cHSjbDfjFnRp"
# Functions can return a Boolean
# + colab={"base_uri": "https://localhost:8080/"} id="uY6vZFHBFtWZ" outputId="cbf76b28-d3e3-4a61-dee6-77e4757f545d"
def my_Function():
return True
print(my_Function())
# + colab={"base_uri": "https://localhost:8080/"} id="W-pWS7dcGhwC" outputId="ded5f542-0b7f-4d76-933b-518d19edf0b3"
if my_Function():
print("True")
else:
print("False")
# + [markdown] id="ZD9hEDF0KJXQ"
# Application 1
# + colab={"base_uri": "https://localhost:8080/"} id="raVMlOhRLfDu" outputId="4d77ca60-f896-4dff-8495-46b07a9c18aa"
print(a==b)
print(a<b)
# + [markdown] id="bRPGnHh2L8XF"
# Python Operators
# + colab={"base_uri": "https://localhost:8080/"} id="awt80XRpL-2p" outputId="80d857ce-becc-4601-af56-b6d373044e0a"
print(10+5)
print(10-5)
print(10/3)
print(10*5)
print(10%5)
print(10//3) #10/3 = 3.3333
print(10**2)
# + [markdown] id="9QsXZ8S5Ni1-"
# Bitwise operators
# + colab={"base_uri": "https://localhost:8080/"} id="Sa-UI7xtNk_5" outputId="c802bed8-b932-438f-d01e-32562cd729d1"
c = 60 # binary 0011 1100
d = 13 # binary 0000 1101
c&d
print(c|d)
print(c^d)
print(d<<2)
# + [markdown] id="6DVVqwnnPmpX"
# Logical Operators
# + colab={"base_uri": "https://localhost:8080/"} id="pD6vcCWdPot_" outputId="d98eb4d0-f17a-4da8-80ee-79d05c3fd5bd"
h = True
l = False
h and l
h or l
not (h or l)
# + [markdown] id="NtYgw2htQPfw"
# Application 2
# + id="daGDCg9HQSGD"
#Python Assignment Operators
# + colab={"base_uri": "https://localhost:8080/"} id="LSVIB7UtSHbK" outputId="4596d39e-56eb-41a8-b775-19f90786e292"
x = 100
x+=3 # Same as x = x +3, x = 100+3=103
print(x)
# + [markdown] id="V6ONWvbLS44l"
# Identity Operators
# + colab={"base_uri": "https://localhost:8080/"} id="UvP7BTi6S7jy" outputId="9f01f735-12c7-4063-f6b0-700196784c34"
h is l
h is not l
# + [markdown] id="GVVjlhWzTLIw"
# #Control Structure
# + [markdown] id="rnloUuSPTO4H"
# If Statement
# + colab={"base_uri": "https://localhost:8080/"} id="jLmtoCqNTTgV" outputId="5052622d-a5cf-4c45-e4d0-540e1ff0965d"
if a>b:
print("a is greater than b")
# + [markdown] id="joSXySWeUQtR"
# Elif Statement
# + colab={"base_uri": "https://localhost:8080/"} id="EGDKVer0UUMi" outputId="eddfd6e5-2cc4-4567-c45b-7efa2cb98e68"
if a<b:
print("a is less than b")
elif a>b:
print("a is greater than b")
# + [markdown] id="3Gx1gNsoUozm"
# Else Statement
# + colab={"base_uri": "https://localhost:8080/"} id="2tRm_JbbUquT" outputId="74b51357-3002-4342-d98b-a42d8fd91535"
a= 10
b =10
if a>b:
print("a is greater than b")
elif a>b:
print("a is greater than b")
else:
print("a is equal to b")
# + [markdown] id="2BpLjDp7U_Dh"
# Short Hand If Statement
# + colab={"base_uri": "https://localhost:8080/"} id="qTsJOVlFVCl3" outputId="a9f9f888-0b17-4499-f295-1235d3195a19"
if a==b: print("a is equal to b")
# + [markdown] id="gS9fa6rrVtMw"
# Short Hand If...Else Statement
# + colab={"base_uri": "https://localhost:8080/"} id="TaI8UGP4Vw5E" outputId="69b545b3-b20a-43f4-f2a0-40a4c3a6a1ef"
a = 10
b = 9
print("a is greater than b") if a>b else print('b is greater than a')
# + [markdown] id="mDaq7HtfWnbE"
# And
# + colab={"base_uri": "https://localhost:8080/"} id="fV4DLwEzWoTD" outputId="f754bbbf-65eb-4a00-fb27-d72fd8538173"
if a>b and b==b:
print("both conditions are True")
# + [markdown] id="ew4gEVRbXTVF"
# Or
# + colab={"base_uri": "https://localhost:8080/"} id="6-dXGIe4XVkV" outputId="6601ae99-7ec7-4a03-de5c-35c7f5a568b8"
if a<b or b==b:
print("the condition is True")
# + [markdown] id="5PH0qT3SXz6K"
# Nested If
# + colab={"base_uri": "https://localhost:8080/"} id="huXDb6mBX1yM" outputId="b74f33b9-70ad-4dc6-8ff4-23e26864e9e6"
x = int(input())
if x>10:
print("x is above 10")
if x>20:
print("and also above 20")
if x>30:
print("and also above 30")
if x>40:
print("and also above 40")
if x>50:
print("and also above 50")
else:
print("but not above 50")
# + [markdown] id="QG4G7y0zajI2"
# #Loop Statement
# + [markdown] id="IdGNkxVcao5y"
# For Loop
# + colab={"base_uri": "https://localhost:8080/"} id="GslmRjkTaqXc" outputId="4308fc1d-9cfa-45b3-a18b-e87e7f1ee9e7"
week = ['Sunday',"Monday",'Tuesday', "Wednesday","Thursday","Friday","Saturday"]
for x in week:
print(x)
# + [markdown] id="x7hf1DxLbUDB"
# The break statement
# + colab={"base_uri": "https://localhost:8080/"} id="nT7BYBOKbDTJ" outputId="622197cc-cd01-4b72-91cb-7b6c89edf4f1"
#to display Sunday to Wednesday using For loop
for x in week:
print(x)
if x=="Wednesday":
break
# + colab={"base_uri": "https://localhost:8080/"} id="L6XDlUebcq49" outputId="8948a949-5faa-4343-8ac9-71dc41c54f05"
#to display only Wednesday using break statement
for x in week:
if x=="Wednesday":
break
print(x)
# + [markdown] id="SJEioQFLcy6S"
# While Statement
# + colab={"base_uri": "https://localhost:8080/"} id="FTI-C9Osc0bG" outputId="1b35cc9d-757b-484c-e06a-8db7a1b831e0"
i =1
while i<6:
print(i)
i+=1 #same as i = i +1
# + [markdown] id="QxJkggjqdYLU"
# Application 3 - Create a python program that displays no.3 using break statement
# + colab={"base_uri": "https://localhost:8080/"} id="up8Ck7HxdkdF" outputId="767a8039-bc91-4bae-891d-58017984c57d"
i =1
while i<6:
if i==3:
break
i+=1
print(i)
| Operations_and_Expressions.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
#bacteria abundance at Family level
fig, ax = plt.subplots(figsize = (9, 35))
PF = sns.barplot(y='Genus', x='TD_ASDdiff', data=df, ci=None)
# pF = sns.countplot(y='Genus', data=df, hue='Positive',)
plt.title('The bacteria abundance in typically developing and ASD groups')
plt.ylabel('Genus', fontsize=11)
plt.xlabel('Bacteria abundance', fontsize=11)
# plt.legend(title="Group", labels=["ASD", "TD"], loc='lower right')
ax.yaxis.set_tick_params(labelsize='large')
plt.show()
# +
# dfF = (df.groupby('Family')['TD_ASDdiff'].mean().sort_values(ascending=False).to_frame()
# )
# dfF
# -
# At the Family level, family *Bifidobacteriacaea, Verrumoicrobiacaea, Fusobacteriacaea, Neisseriaceae* and *Enterobacteriacaea* were more abundant in ASD group. In TD group, family *Prevotellaceae* and *Bacteroidaceae* were more abundant.
# +
#At phylum level
# sns.barplot(x='TD_ASDdiff', y='Phylum', data =df, ci=None, hue=["b" if y not in ['Bacteroidetes', 'Synergistetes'] else "r" for y in df.Phylum])
# plt.title('The bacteria abundance in TD and ASD group without Error bars')
# plt.ylabel('Bacteria Phylum')
# plt.xlabel('Bacteria abundance')
# plt.legend(title="Group", labels=["ASD"])
# ax.yaxis.set_tick_params(labelsize='large')
# plt.show()
# pt1 = sns.countplot(y='Phylum', data=df, hue='Positive')
# plt.title('Countplot of Bacteria abundance at the Phylum level', fontsize=11)
# plt.xlabel('Abundance', fontsize=11)
# plt.ylabel('Phylum', fontsize=11)
# plt.legend(title="Group", labels=["ASD", "TD"])
# ax.yaxis.set_tick_params(labelsize='large')
# plt.show()
# b1 = sns.barplot(x='Phylum', y='ASDmean', data=df, ci=None, color='lightblue') #order=df.sort_values('Phylum', ascending=False).ASDmean)
# b2= sns.barplot(x='Phylum', y='TDmean', data=df, ci=None, color='salmon', ) #order=df.sort_values('Phylum', ascending=False).TDmean)
# locs, labels = plt.xticks()
# plt.setp(labels, rotation=45, fontsize='large')
# plt.xlabel('Phylum', fontsize=11)
# plt.ylabel('Abundance', fontsize=11)
# top_bar = mpatches.Patch(color="lightblue", label='ASD')
# bottom_bar = mpatches.Patch(color='salmon', label='TD')
# plt.legend(handles=[top_bar, bottom_bar])
# plt.show()
# -
# At the phylum level, Verrucommicrobial, Actinobacteria and Fusobacteria are more abundant in ASD group while Bacteroidetes is more abundant in TD group.
# +
# at class
# dfC = (df.groupby('Class')['TD_ASDdiff'].mean().sort_values(ascending=False).to_frame()
# )
# dfC
# +
#plotting data at class level
# fig, ax = plt.subplots(figsize = (9, 7))
# pC = sns.barplot(y='Class', x='TD_ASDdiff', data=df, ci=None,
# hue=["b" if y not in ['Bacteroidia','Deltaproteobacteria', 'Synergistia', 'Alphaproteobacteria'] else "r" for y in df.Class])
# plt.title('The bacteria abundance in TD and ASD group')
# plt.ylabel('Bacteria Class')
# plt.xlabel('Bacteria abundance')
# plt.legend(title="Group", labels=["TD", "ASD"], loc='lower right')
# ax.yaxis.set_tick_params(labelsize='large')
# plt.show()
#stacked plot
# fig, ax = plt.subplots(figsize = (10, 7))
# b1 = sns.barplot(y='Class', x='ASDmean', data=df, ci=None, color='lightblue')
# b2= sns.barplot(y='Class', x='TDmean', data=df, ci=None, color='salmon')
# locs, labels = plt.xticks()
# plt.setp(labels, rotation=45, fontsize='large')
# plt.xlabel('Abundance', fontsize=11)
# plt.ylabel('Class', fontsize=11)
# top_bar = mpatches.Patch(color="lightblue", label='ASD')
# bottom_bar = mpatches.Patch(color='salmon', label='TD')
# plt.legend(handles=[top_bar, bottom_bar])
# plt.show()
# +
# another way to do a stacked plot at class
# sns.set_color_codes("pastel")
# sns.barplot(y="Class", x="ASDmean", data=df,
# label="ASD", color="b", ci=None)
# sns.set_color_codes("muted")
# sns.barplot(y="Class", x="TDmean", data=df,
# label="TD", color="b", ci=None)
# plt.legend(ncol=2, loc="lower right", frameon=True)
# ax.set(xlim=(0, 240), ylabel="",
# xlabel="Bacteria abundance at phylum level")
# sns.despine(left=True, bottom=True)
# -
# At the class level, Verrucomicrobiae, Actinobacteria, Fusobacteria and Gammaproteobacteria were more abundant in ASD group while Bacteroidia was more abundant in TD group.
# +
# at order
# dfO = (df.groupby('Order')['TD_ASDdiff'].mean().sort_values(ascending=False).to_frame()
# )
# dfO
# +
#bacteria abundance at Order
# fig, ax = plt.subplots(figsize = (8, 7))
# sns.barplot(x='TD_ASDdiff', y='Order', data =df, ci=None)
# #this graph doesn't mactched the dataframe above... and same for all fo the graphs below
# # pO = sns.countplot(y='Order', data=df, hue='Positive')
# plt.title('The bacteria abundance in TD and ASD group')
# plt.ylabel('Order', fontsize=11)
# plt.xlabel('Abundance', fontsize=11)
# # plt.legend(title="Group", labels=["ASD", "TD"], loc='lower right')
# ax.yaxis.set_tick_params(labelsize='large')
# plt.show()
# +
#stacked plot at order
# fig, ax = plt.subplots(figsize = (8, 8))
# b1 = sns.barplot(y='Order', x='ASDmean', data=df, ci=None, color='lightblue')
# b2= sns.barplot(y='Order', x='TDmean', data=df, ci=None, color='salmon')
# locs, labels = plt.xticks()
# plt.setp(labels, rotation=45, fontsize='large')
# plt.xlabel('Abundance', fontsize=11)
# plt.ylabel('Order', fontsize=11)
# top_bar = mpatches.Patch(color="lightblue", label='ASD')
# bottom_bar = mpatches.Patch(color='salmon', label='TD')
# plt.legend(handles=[top_bar, bottom_bar])
# ax.yaxis.set_tick_params(labelsize='large')
# plt.show()
# -
# At the Order level, order Bifidobacteriales(class actinobacteria), Verrucomicrobiales, Fusobacteriales, Neisseriales (class betaproteoabcteria) and Enterobacteriales were more abundant in ASD group, while Bacteroidales was more abundant in TD group.
# +
#plot showing the bacteria abundance in ASD and TD individuals at phylum.
#this graph does work (right size label with wrong color)
# sns.barplot(x='TD_ASDdiff', y='Phylum', data =df, ci=None,palette = ['lightsalmon', 'lightseagreen'])
# plt.title('The bacteria abundance in TD and ASD group')
# plt.ylabel('Bacteria Phylum')
# plt.xlabel('Bacteria abundance')
# ax.yaxis.set_tick_params(labelsize='large')
# plt.show()
# +
#td and asdmean at genus--shows repeated genera on graph (probably belongs to different species)
# fig, ax = plt.subplots(figsize = (9, 25))
# b1 = sns.barplot(y='Genus', x='ASDmean', data=df, ci=None, color='lightsalmon',
# order=df.sort_values('ASDmean', ascending = True).Genus)
# b2= sns.barplot(y='Genus', x='TDmean', data=df, ci=None, color='lightseagreen',
# order=df.sort_values('TDmean', ascending=False).Genus)
# locs, labels = plt.xticks()
# plt.setp(labels, rotation=45, fontsize='large')
# plt.title('The mean bacteria abundance of Typically Deveoping group and ASD group at the genus level', fontsize=13)
# plt.xlabel('Abundance', fontsize=11)
# plt.ylabel('Genus', fontsize=11)
# top_bar = mpatches.Patch(color='lightsalmon', label='ASD')
# bottom_bar = mpatches.Patch(color='lightseagreen', label='TD')
# plt.legend(handles=[top_bar, bottom_bar])
# ax.yaxis.set_tick_params(labelsize='medium')
# ax.xaxis.set_tick_params(labelsize='medium')
# plt.show()
# +
#trying to drop the all the outliers but it didn't work
# df[(np.abs(stats.zscore(df)) < 3).all(axis=1)]
# df[np.abs(df.Data-df.Data.mean()) <= (3*df.Data.std())]
# q = df["col"].quantile(0.99)
# df = df[df.between(df.quantile(.15), df.quantile(.85))]
# def drop_numerical_outliers(df, z_thresh=3):
# # Constrains will contain `True` or `False` depending on if it is a value below the threshold.
# constrains = df.select_dtypes(include=[np.number]) \
# .apply(lambda x: np.abs(stats.zscore(x)) < z_thresh, reduce=False) \
# .all(axis=1)
# # Drop (inplace) values set to be rejected
# df.drop(df.index[~constrains], inplace=True)
# drop_numerical_outliers(df)
# +
#trying to make the neg values in red in pos in green
# def bar_color(df,color1,color2):
# return np.where(df.TD_ASDdiff>0,color1,color2).T
# df.TD_ASDdiff.plot.barh(color=bar_color(df.TD_ASDdiff, 'r', 'g'))
# +
#trying to create a custom palette for the horizontal bar graph
# custom_palette = {}
# for q in df.TD_ASDdiff:
# avr = df[df.TD_ASDdiff == q].Positive
# if avr.where(df['TD_ASDdiff']) == 1:
# custom_palette[q] = 'r'
# else :
# custom_palette[q] = 'y'
# -
| analysis/Tara/milestone2/Junk.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Grand Challenge API Client: interact with grand-challenge.org via python
# ## Chapter: Algorithms
#
# <img src="pipeline.png" />
# GC-API provides a handy functionality to interract with grand-challenge.org via python.
#
# Its purpose and broader functionality is discussed in details in [this](https://grand-challenge.org/blogs/grand-challenge-api-client/) blogpost. In order to use gc-api you will also need to obtain a personal API token. The blog above describe what it is and where to find it.
#
# In this tutorial we will focus on grand-challenge Algorithms and in details go over following steps:
#
# 1. [Uploading input to Algorithms on grand-challenge.org for inference.](#section_1)
# 2. [Downloading inference results from an Algorithm on grand-challenge.org.](#section_2)
# 3. [Uploading multiple item input to an Algorithm on grand-challenge.org.](#section_3)
# 4. [Downloading the results of an algorithm that produces multiple item output.](#section_4)
#
# Remember that you need to request a permission prior to using an algorithm. You do not need to request a permission if you are using your own algorithm.
pip install gcapi --upgrade
#import nessesary libraries
import gcapi
from pathlib import Path
from tqdm import tqdm
import SimpleITK as sitk
import numpy as np
import os
# We use the API token to authentificate to grand-challenge platform via API
#authorise with your personal token
my_personal_GC_API_token = ''
client = gcapi.Client(token=my_personal_GC_API_token)
# ## [1. Uploading input to Algorithms on grand-challenge.org for inference.](#section_1)
# In this section, we will use [Pulmonary Lobe Segmentation](https://grand-challenge.org/algorithms/pulmonary-lobe-segmentation/) by <NAME>. This algorithm performs automatic segmentation of pulmonary lobes of a given chest CT scan. The algorithm uses a contextual two-stage U-Net architecture. We will use example chest CT scans from [coronacases.org](coronacases.org). They are anonimized.
# We will now upload the CT scans of Covid-19 patients to Pulmonary Lobe Segmentation algorithm on grand-challenge using GC-API.
# initialize the algorithm, providing a slug
algorithm_1 = client.algorithms.detail(slug="pulmonary-lobe-segmentation")
# explore, which input the algorithm expects
algorithm_1["inputs"]
# #### Submit the inputs to the algorithm one by one
# Grand-challenge creats a job instance for each set of inputs. To create a job instance use a command:
#
# job = client.run_external_job(algorithm="slug-of-the-algorithm", inputs={ "interface": [ file ] }),
#
# where argument "algorithm" expects a str with a slug of the algorithm you want to use and argument "inputs" expects a dictionary where keys are expected interfaces and the file is str path/url to a particular input file.
#
# Be aware that with this version (0.5.0) the input file path/url needs be placed into a list.
# +
# get the path to the files
files = ["io/case01.mha", "io/case02.mha"]
#timeout
jobs = []
# submit a job for each file in your file list
for file in files:
job = client.run_external_job(
algorithm="pulmonary-lobe-segmentation",
inputs={
"generic-medical-image": [Path(file)]
}
)
jobs.append(job)
# -
# #### Get the statuses of the submitted jobs
# +
jobs = [client.algorithm_jobs.detail(job["pk"]) for job in jobs]
print([job["status"] for job in jobs])
# -
# After all of your jobs have ended up with a status 'Succeeded', you can download the results. You can also use infer the Algorithm on existing Archive on grand-challenge.org (if you have been granted access to it).
# ## [2. Downloading inference results from an Algorithm on grand-challenge.org.](#section_2)
#
# #### Download the results from the algorithm
# loop through input files
for job, input_fname in tqdm(zip(jobs, files)):
# loop through job outputs
for output in job["outputs"]:
# check whether if output exists
if output["image"] is not None:
# get image details
image_details = client(url=output["image"])
print('image_details',image_details)
for file in image_details["files"]:
print('file',file)
# create the output filename
output_file = Path(input_fname.replace(".mha", "_lobes.mha"))
if output_file.suffix != ".mha":
raise ValueError("Output file needs to have .mha extension")
output_file.parent.mkdir(parents=True, exist_ok=True)
with output_file.open("wb") as fp:
# get the impage from url and write it
response = client(url = file["file"],follow_redirects=True).content
fp.write(response)
# ## [3. Uploading multiple item input to an Algorithm on grand-challenge.org.](#section_3)
#
#
# In this section we will take a look, on how to upload multiple item input to an Algorithm on grand-challenge.org. Asw an example we will use Alessa Herrings Algorithm - [Deep Learning-Based Lung Registration](https://grand-challenge.org/algorithms/deep-learning-based-ct-lung-registration/).
#
# This algorithm requires the following inputs:
# 1. fixed image (CT)
# 2. fixed mask (lungs segmentation)
# 3. moving image (CT)
# 4. moving mask (lungs segmentation)
#
# We will use the scans from the previous section as well as the Algorithm output (lung lobes segmentation) in this section. Therefore, we will have to binarize the lobe masks and create lung masks
# #### Binarize the masks obtained from previous example
# provide paths of the lobe segmentations
lobes = [
"io/case01_lobes.mha",
"io/case02_lobes.mha",
]
#loop through the files
for lobe_file in lobes:
#read image with sitk
lobe = sitk.ReadImage(lobe_file)
origin, spacing, direction = lobe.GetOrigin(), lobe.GetSpacing(), lobe.GetDirection()
lobe = sitk.GetArrayFromImage(lobe)
# binarize
lobe[lobe > 1] = 1
lungs = lobe.astype(np.uint8)
lungs = sitk.GetImageFromArray(lungs)
lungs.SetOrigin(origin)
lungs.SetSpacing(spacing)
lungs.SetDirection(direction)
#write the modified image back into file
sitk.WriteImage(lungs, lobe_file.replace("_lobes", "_lungs"), True)
# initialize the algorithm
algorithm_2 = client.algorithms.detail(slug="deep-learning-based-ct-lung-registration")
# #### Inspecting the algorithm object
# You can inspect the algorithm object to understand what kind of inputs it requires
algorithm_2["inputs"]
# #### Submit the inputs to the algorithm
# create a job
registration_job = client.run_external_job(
algorithm="deep-learning-based-ct-lung-registration",
inputs={
"fixed-image": [Path("io/case01.mha")],
"moving-image": [Path("io/case02.mha")],
"fixed-mask": [Path("io/case01_lungs.mha")],
"moving-mask": [Path("io/case02_lungs.mha")],
}
)
# #### Get the status of the job
registration_job = client.algorithm_jobs.detail(registration_job["pk"])
registration_job["status"]
# #### Download the results
#
# After the status of the job is `Succeeded`, you can procceed to downloading the result.
# loop through the outputs
for output in registration_job["outputs"]:
print('output',output)
# get image details
image_details = client(url=output["image"])
output_slug = output["interface"]["slug"]
print("Downloading", output_slug)
for file in image_details["files"]:
output_file = Path(f"{output_slug}.mha")
output_file.parent.mkdir(parents=True, exist_ok=True)
with output_file.open("wb") as fp:
fp.write(client(url=file["file"], follow_redirects=True).content)
# Note that both these algorithms wrote `.mha` files as outputs. For algorithms that require different outputs, you can loop through the outputs of a successful job and search under "interface", which will tell you what kind of outputs you will have to download
# ### [4. Downloading the results of an algorithm that produces multiple item output.](#section_4)
#
# In this section we will focus on how to download results from an algorithm that produces multiple outputs. We will use the algorithm for pulmonary lobes segmentation Covid-19 cases. This algorithm output the segmentation for a particular input as well as a "screenshot" of a middle slice for rapid inspection of algorithm performance.
#
# +
# initialize the algorithm, providing a slug
algorithm_4 = client.algorithms.detail(slug="pulmonary-lobe-segmentation-for-covid-19-ct-scans")
# explore, which input the algorithm expects
algorithm_4["inputs"]
# -
# We will, firstly infer the algorithm on the existing images, exactly the same way we did before.
# +
from gcapi import Client
c = Client(token=my_personal_GC_API_token)
#name of the archive
archive_slug = "coronacases.org"
#save path on your machine
output_archive_dir = 'output_scans'
outputarchivedir_screenshots = 'output_screenshots'
archives = c(url="https://grand-challenge.org/api/v1/archives/")["results"]
corona_archive = None
for archive in archives:
if archive["name"] == archive_slug:
corona_archive = archive
break
if corona_archive is None:
raise Exception("archive not found on GC")
# +
print(corona_archive)
# get information about images in archive from GC API
params = {
'archive': corona_archive['id'],
}
response = c(url="https://grand-challenge.org/api/v1/cases/images/", params=params)
#print(response)
urls=[]
for r in response['results']:
urls.append(r['api_url'])
# -
print(len(urls))
print(urls)
# +
# initialize the archive, providing a slug
jobs = []
# submit a job for each file in your file list
for url in urls[:2]:
print(url)
job = client.run_external_job(
algorithm="pulmonary-lobe-segmentation-for-covid-19-ct-scans",
inputs={
"ct-image": url
}
)
jobs.append(job)
# -
# Lets check the status of the job.
# +
jobs = [client.algorithm_jobs.detail(job["pk"]) for job in jobs]
print([job["status"] for job in jobs])
# -
# If the job status is 'Succeeded' we can procceed to downloading the results. In this part we will go through scenario where we no longer have the details of the particular job that processed algorithm and inputs.
# +
from gcapi import Client
c = Client(token=my_personal_GC_API_token)
# get algorithm providing the slug
algorithm = "pulmonary-lobe-segmentation-for-covid-19-ct-scans"
algorithm_details = c(path="algorithms/", params={"slug": algorithm})
# extract details
algorithm_details = algorithm_details["results"][0]
algorithm_uuid = algorithm_details["pk"]
# Define dictionaries for image uuid mappings
images_mapping = {}
images_mapping_scans = {}
# get the desired archive
archives = c(url="https://grand-challenge.org/api/v1/archives/")["results"]
archive_slug = 'coronacases.org'
target_archive = None
# loop through archives and select the one with a slug that you are looking for ('coronacases.org')
for archive in archives:
if archive["name"] == archive_slug:
target_archive = archive
break
# -
# We have generated a set of outputs to a set of inputs. Now, we need to find out which output corresponds to which input. This can be figured out via unique identifiers of images. Each image in an archive has a unique identifier (uuid). Here we create a **mapping between input image names and uuids**. We collect the uuids to a list.
# get uuids in archive
done = False
iteration = 0
image_uuids = []
# create a mapping between image uuids and input names
while not done:
iteration += 1
if iteration == 1:
# get information about images in archive from GC API
params = {'archive': target_archive['id']}
response = c(url="https://grand-challenge.org/api/v1/cases/images/", params=params)
else:
# get information about images on next page
response = c(url=response["next"])
images = response['results']
for image in images:
# create a mapping for image uuids
uuid = image['pk']
images_mapping[uuid] = image['name'] + "_" + uuid
images_mapping_scans[uuid] = image['name']
image_uuids += [uuid]
if response["next"] is None:
# stop if no next page left
done = True
print('image_uuids:',image_uuids)
print('--------------------------------------------------------------------------------------------------------')
print('images_mapping_scans:',images_mapping_scans)
# Now, we will loop through uuids and collect the algorithm job details corresponding to each unique image identifier.
# get algorithm results for the image
output_image_files = []
screenshot_files = []
counter = 0
# loop through uuid
for image_uuid in image_uuids:
params = {'algorithm_image__algorithm': algorithm_uuid, 'input_image': image_uuid}
# get the jobs details corresponding to a particular uuid and algorithm
results = c.algorithm_jobs.iterate_all(params)
#iterate through the results
for result in results:
counter += 1
print('--------------------------------------------------------------------------------------------------------')
#iterate through the outputs
for output in result['outputs']:
# here we go over different interfaces and write the corresponding output
print('interface:',output["interface"]["slug"])
if output["interface"]["slug"] == "pulmonary-lobes":
image = c(url=output['image'])
for file in image["files"]:
if file['image_type'] == "MHD":
new_file = file['file']#image['files'][0]['file']
output_image_files += [new_file]
dest_path_mha = Path(os.path.join(output_archive_dir, images_mapping_scans[image_uuid]))
with open(dest_path_mha, 'wb') as f1:
response_1 = c(url=new_file, follow_redirects=True)
f1.write(response_1.content)
print(dest_path_mha)
if output["interface"]["slug"] == "pulmonary-lobes-screenshot":
image = c(url=output['image'])
for file in image["files"]:
if file['image_type'] == "TIFF":
new_file = file['file']
screenshot_files += [new_file]
dest_path = os.path.join(outputarchivedir_screenshots, images_mapping[image_uuid] + '.tif')
with open(dest_path, 'wb') as f1:
response_2 = c(url=new_file, follow_redirects=True)
f1.write(response_2.content)
print(dest_path)
| tutorials/gcapi-tutorial-algorithms.ipynb |
// -*- coding: utf-8 -*-
// ---
// jupyter:
// jupytext:
// text_representation:
// extension: .groovy
// format_name: light
// format_version: '1.5'
// jupytext_version: 1.14.4
// kernelspec:
// display_name: Groovy
// language: groovy
// name: groovy
// ---
// # Tablesaw
//
// [Tablesaw](https://tablesaw.tech/) is easy to add to the BeakerX Groovy kernel.
// Tablesaw provides the ability to easily transform, summarize, and filter data, as well as computing descriptive statistics and fundamental machine learning algorithms.
//
// This notebook has some basic demos of how to use Tablesaw, including visualizing the results. This notebook uses the Beaker interactive visualizaiton libraries, but Tablesaw's APIs also work. The notebook covers basic table manipulation, k-means clustering, and linear regression.
// %%classpath add mvn
tech.tablesaw tablesaw-plot 0.11.4
tech.tablesaw tablesaw-smile 0.11.4
tech.tablesaw tablesaw-beakerx 0.11.4
// +
// %import tech.tablesaw.aggregate.*
// %import tech.tablesaw.api.*
// %import tech.tablesaw.api.ml.clustering.*
// %import tech.tablesaw.api.ml.regression.*
// %import tech.tablesaw.columns.*
// display Tablesaw tables with BeakerX table display widget
tech.tablesaw.beakerx.TablesawDisplayer.register()
// -
// 3
tornadoes = Table.read().csv("../../../doc/resources/data/tornadoes_2014.csv")
//cell 3 expected result
Image("../../resources/img/groovy/tablesaw/cell3_case1.png")
// 4
//print dataset structure
tornadoes.structure()
//cell 4 expected result
Image("../../resources/img/groovy/tablesaw/cell4_case1.png")
//get header names
tornadoes.columnNames()
//displays the row and column counts
tornadoes.shape()
// 7
//displays the first n rows
tornadoes.first(10)
//cell 7 expected result
Image("../../resources/img/groovy/tablesaw/cell7_case1.png")
// 8
import static tech.tablesaw.api.QueryHelper.column
tornadoes.structure().selectWhere(column("Column Type").isEqualTo("FLOAT"))
//cell 8 expected result
Image("../../resources/img/groovy/tablesaw/cell8_case1.png")
//summarize the data in each column
tornadoes.summary()
//Mapping operations
def month = tornadoes.dateColumn("Date").month()
tornadoes.addColumn(month);
tornadoes.columnNames()
// 11
//Sorting by column
tornadoes.sortOn("-Fatalities")
//cell 11 expected result
Image("../../resources/img/groovy/tablesaw/cell11_case1.png")
// 12
//Descriptive statistics
tornadoes.column("Fatalities").summary()
//cell 12 expected result
Image("../../resources/img/groovy/tablesaw/cell12_case1.png")
// 13
//Performing totals and sub-totals
def injuriesByScale = tornadoes.median("Injuries").by("Scale")
injuriesByScale.setName("Median injuries by Tornado Scale")
injuriesByScale
//cell 13 expected result
Image("../../resources/img/groovy/tablesaw/cell13_case1.png")
// 14
//Cross Tabs
CrossTab.xCount(tornadoes, tornadoes.categoryColumn("State"), tornadoes.shortColumn("Scale"))
//cell 14 expected result
Image("../../resources/img/groovy/tablesaw/cell14_case1.png")
// ## K-means clustering
//
// K-means is the most common form of “centroid” clustering. Unlike classification, clustering is an unsupervised learning method. The categories are not predetermined. Instead, the goal is to search for natural groupings in the dataset, such that the members of each group are similar to each other and different from the members of the other groups. The K represents the number of groups to find.
//
// We’ll use a well known Scotch Whiskey dataset, which is used to cluster whiskeys according to their taste based on data collected from tasting notes. As always, we start by loading data and printing its structure.
//
// More description is available at https://jtablesaw.wordpress.com/2016/08/08/k-means-clustering-in-java/
// 15
t = Table.read().csv("../../../doc/resources/data/whiskey.csv")
t.structure()
//cell 15 expected result
Image("../../resources/img/groovy/tablesaw/cell15_case1.png")
// +
// 16
model = new Kmeans(
5,
t.nCol(2), t.nCol(3), t.nCol(4), t.nCol(5), t.nCol(6), t.nCol(7),
t.nCol(8), t.nCol(9), t.nCol(10), t.nCol(11), t.nCol(12), t.nCol(13)
);
//print claster formation
model.clustered(t.column("Distillery"));
// -
// cell 16 expected result
Image("../../resources/img/groovy/tablesaw/cell16_case1.png")
// 17
//print centroids for each claster
model.labeledCentroids();
// cell 17 expected result
Image("../../resources/img/groovy/tablesaw/cell17_case1.png")
//gets the distortion for our model
model.distortion()
// +
def n = t.rowCount();
def kValues = new double[n - 2];
def distortions = new double[n - 2];
for (int k = 2; k < n; k++) {
kValues[k - 2] = k;
def kmeans = new Kmeans(k,
t.nCol(2), t.nCol(3), t.nCol(4), t.nCol(5), t.nCol(6), t.nCol(7),
t.nCol(8), t.nCol(9), t.nCol(10), t.nCol(11), t.nCol(12), t.nCol(13)
);
distortions[k - 2] = kmeans.distortion();
}
def linearYPlot = new Plot(title: "K-means clustering demo", xLabel:"K", yLabel: "distortion")
linearYPlot << new Line(x: kValues, y: distortions)
// -
// ## Play (Money)ball with Linear Regression
//
// In baseball, you make the playoffs by winning more games than your rivals. The number of games the rivals win is out of your control so the A’s looked instead at how many wins it took historically to make the playoffs. They decided that 95 wins would give them a strong chance. Here’s how we might check that assumption in Tablesaw.
//
// More description is available at https://jtablesaw.wordpress.com/2016/07/31/play-moneyball-data-science-in-tablesaw/
// +
import static tech.tablesaw.api.QueryHelper.column
baseball = Table.read().csv("../../../doc/resources/data/baseball.csv");
// filter to the data available at the start of the 2002 season
moneyball = baseball.selectWhere(column("year").isLessThan(2002));
wins = moneyball.nCol("W");
year = moneyball.nCol("Year");
playoffs = moneyball.column("Playoffs");
runDifference = moneyball.shortColumn("RS").subtract(moneyball.shortColumn("RA"));
moneyball.addColumn(runDifference);
runDifference.setName("RD");
def Plot = new Plot(title: "RD x Wins", xLabel:"RD", yLabel: "W")
Plot << new Points(x: moneyball.numericColumn("RD").toDoubleArray(), y: moneyball.numericColumn("W").toDoubleArray())
// -
winsModel = LeastSquares.train(wins, runDifference);
def runDiff = new double[1];
runDiff[0] = 135;
def expectedWins = winsModel.predict(runDiff);
runsScored2 = LeastSquares.train(moneyball.nCol("RS"), moneyball.nCol("OBP"), moneyball.nCol("SLG"));
new Histogram(xLabel:"X",
yLabel:"Proportion",
data: Arrays.asList(runsScored2.residuals()),
binCount: 25);
// ## Financial and Economic Data
//
// You can fetch data from [Quandl](https://www.quandl.com/) and load it directly into Tablesaw
// %classpath add mvn com.jimmoores quandl-tablesaw 2.0.0
// %import com.jimmoores.quandl.*
// %import com.jimmoores.quandl.tablesaw.*
// +
// 34
TableSawQuandlSession session = TableSawQuandlSession.create();
Table table = session.getDataSet(DataSetRequest.Builder.of("WIKI/AAPL").build());
// Create a new column containing the year
ShortColumn yearColumn = table.dateColumn("Date").year();
yearColumn.setName("Year");
table.addColumn(yearColumn);
// Create max, min and total volume tables aggregated by year
Table summaryMax = table.groupBy("year").max("Adj. Close");
Table summaryMin = table.groupBy("year").min("Adj. Close");
Table summaryVolume = table.groupBy("year")sum("Volume");
// Create a new table from each of these
summary = Table.create("Summary", summaryMax.column(0), summaryMax.column(1),
summaryMin.column(1), summaryVolume.column(1));
// Add back a DateColumn to the summary...will be used for plotting
DateColumn yearDates = new DateColumn("YearDate");
for(year in summary.column('Year')){
yearDates.append(java.time.LocalDate.of(year,1,1));
}
summary.addColumn(yearDates)
summary
// -
// cell 34 expected result
Image("../../resources/img/groovy/tablesaw/cell34_case1.png")
// +
years = summary.column('YearDate').collect()
plot = new TimePlot(title: 'Price Chart for AAPL', xLabel: 'Time', yLabel: 'Max [Adj. Close]')
plot << new YAxis(label: 'Volume')
plot << new Points(x: years, y: summary.column('Max [Adj. Close]').collect())
plot << new Line(x: years, y: summary.column('Max [Adj. Close]').collect(), color: Color.blue)
plot << new Stems(x: years, y: summary.column('Sum [Volume]').collect(), yAxis: 'Volume')
// -
| test/ipynb/groovy/TablesawTest.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Finding and Cleaning r/loseit Challenge Data
# +
import datetime as dt
import os
import re
import statistics
from pathlib import Path
import matplotlib.pyplot as plt
import numpy as np
import gspread
import lxml
import markdown
import pandas as pd
import praw
import seaborn as sns
from lxml import etree
from oauth2client.service_account import ServiceAccountCredentials
pd.options.mode.chained_assignment = None # default='warn'
# -
# This is a function used to sort some of the lists in human ordered form.
# +
def atoi(text):
return int(text) if text.isdigit() else text
def natural_keys(text):
"""
alist.sort(key=natural_keys) sorts in human order
http://nedbatchelder.com/blog/200712/human_sorting.html
"""
return [atoi(c) for c in re.split("(\d+)", text)]
def month_year(timestamps):
return str(timestamps.date())[:-3]
def get_date(created):
return dt.datetime.fromtimestamp(created)
# -
# Create the Data Directories
if not os.path.exists("./data/raw_data/"):
os.makedirs("./data/raw_data/")
if not os.path.exists("./figures/"):
os.makedirs("./figures/")
if not os.path.exists("./data/cleaned_data/"):
os.makedirs("./data/cleaned_data/")
# loseit_data is the name at the top of the praw.ini file
reddit = praw.Reddit("loseit_data")
loseit_sub = reddit.subreddit("loseit")
challenge_posts = loseit_sub.search("loseit challenge tracker", limit=1000)
topics_dict = {
"title": [],
"score": [],
"id": [],
"url": [],
"comms_num": [],
"created": [],
"body": [],
}
for submission in challenge_posts:
topics_dict["title"].append(submission.title)
topics_dict["score"].append(submission.score)
topics_dict["id"].append(submission.id)
topics_dict["url"].append(submission.url)
topics_dict["comms_num"].append(submission.num_comments)
topics_dict["created"].append(submission.created)
topics_dict["body"].append(submission.selftext)
# +
topics_data = pd.DataFrame(topics_dict)
_timestamp = topics_data["created"].apply(get_date)
topics_data = topics_data.assign(timestamp=_timestamp)
topics_data.to_csv("../data/raw_data/loseit_search_history.csv")
# +
# Now that we have searched through old loseit posts, we need to find the urls.
links = []
for body in topics_dict["body"]:
try:
doc = etree.fromstring(markdown.markdown(re.sub("[\\n]", "", body)))
for link in doc.xpath("//a"):
web_url = link.get("href")
if bool(re.search("spreadsheet", web_url)) and bool(
re.search("oogle", web_url)
):
links.append(web_url)
except etree.XMLSyntaxError:
pass
unique_spreadsheets = list(set(links))
# -
# use creds to create a client to interact with the Google Drive API
names = []
for spreadsheet_link in unique_spreadsheets:
scope = ["https://spreadsheets.google.com/feeds"]
creds = ServiceAccountCredentials.from_json_keyfile_name(
"loseit-sheets-6012c29a1f40.json", scope # this is the google-app.json file
)
gc = gspread.authorize(creds)
sht = gc.open_by_url(spreadsheet_link)
if (
bool(re.search("nter", sht.title)) == False
and bool(re.search("/r/", sht.title)) == False
and bool(re.search("Calculator", sht.title)) == False
):
sheet_name = re.sub(
"_\(responses\)",
"",
re.sub(
",",
"",
re.sub(
"\]",
"",
re.sub(
"\[",
"",
re.sub(
" ",
"_",
re.sub(" ", "_", re.sub("-", "", sht.title.lower())),
),
),
),
),
)
if sheet_name not in names:
print(f"sheet name: {sheet_name}, link: {spreadsheet_link}")
names.append(sheet_name)
try:
data_sheet = sht.worksheet("Tracker")
data_vals = data_sheet.get_all_values()
data_df = pd.DataFrame(data_vals[1:-2], columns=data_vals[0])
data_df.to_csv("../data/raw_data/" + sheet_name + ".csv")
except gspread.WorksheetNotFound:
try:
data_sheet = sht.worksheet("Master Spreadsheet")
data_vals = data_sheet.get_all_values()
data_df = pd.DataFrame(data_vals[1:-2], columns=data_vals[0])
data_df.to_csv("../data/raw_data/" + sheet_name + ".csv")
except gspread.WorksheetNotFound:
print("", end="\t") # sheet_name)
else:
print("", end="\t") # sheet_name)
# The next step is cleaning up some of the column information, and removing the information that is not useful for analysis.
weeks_col = {f"W{x}": f"Week {x}" for x in range(0, 11)}
new_names = {
"W0 (SW)": "Week 0",
"Sex": "Gender",
"Male, Female, Other": "Gender",
"TEAM": "Team",
"Teams": "Team",
"Challenge GW": "Challenge Goal Weight",
"Challenge SW": "Week 0",
"MyFitnessPal Username/Link": "MFP",
}
df_list = []
p = Path("../data/raw_data/")
for idx, challenge in enumerate(p.rglob("*.csv")):
# Challenge Names
challenge_name = re.sub("\d", "", challenge.name[:-4])
# Read in the csv files and change some of the column names
test_df = pd.read_csv(challenge, index_col=0)
test_df.dropna(axis=1, how="all")
test_df.columns = (
test_df.columns.str.strip().str.replace("?", "").str.replace(":", "")
)
test_df.rename(columns=new_names, inplace=True)
# timestamp
if "Timestamp" not in test_df:
test_df["Timestamp"] = (
"October 2018"
if challenge_name == "super_mario_brothers_super_loseit_challenge_tracker"
else "March 2017"
)
test_df.Timestamp = pd.to_datetime(test_df.Timestamp, errors="coerce").apply(
month_year
)
# Age
test_df["Age"] = test_df[
test_df.filter(regex=re.compile("Age", re.IGNORECASE)).columns[0]
]
# Gender
if len(test_df.filter(regex=re.compile("Sex", re.IGNORECASE)).columns):
test_df["Gender"] = test_df[
test_df.filter(regex=re.compile("Sex", re.IGNORECASE)).columns[0]
]
if len(test_df.filter(regex=re.compile("Gender", re.IGNORECASE)).columns):
test_df["Gender"] = test_df[
test_df.filter(regex=re.compile("Gender", re.IGNORECASE)).columns[0]
]
if "Gender" not in test_df:
test_df["Gender"] = "Unknown"
# Ignore KGS
if len(test_df.filter(regex=re.compile("kgs", re.IGNORECASE)).columns):
test_df.drop(
test_df.filter(regex=re.compile("kgs", re.IGNORECASE)).columns[0],
axis=1,
inplace=True,
)
# Keep Just Starting BMI
test_df.drop(
test_df.filter(regex=re.compile("BMI", re.IGNORECASE)).columns[1:],
axis=1,
inplace=True,
)
# Username
test_df.columns = test_df.columns.str.replace(
test_df.filter(like="name").columns[0], "Username"
)
test_df.Username = test_df.Username.astype(str).apply(lambda x: x.lower())
# Weigh-in Data
test_df.rename(columns=weeks_col, inplace=True)
if len(test_df.filter(regex=re.compile("week 0", re.IGNORECASE)).columns):
test_df["Week 0"] = test_df[
test_df.filter(regex=re.compile("week 0", re.IGNORECASE)).columns[0]
]
elif len(test_df.filter(regex=re.compile("sign-up", re.IGNORECASE)).columns):
test_df["Week 0"] = test_df[
test_df.filter(regex=re.compile("sign-up", re.IGNORECASE)).columns[0]
]
elif len(test_df.filter(regex=re.compile("start weight", re.IGNORECASE)).columns):
test_df["Week 0"] = test_df[
test_df.filter(regex=re.compile("start weight", re.IGNORECASE)).columns[0]
]
elif len(test_df.filter(regex=re.compile("Signup weight", re.IGNORECASE)).columns):
test_df["Week 0"] = test_df[
test_df.filter(regex=re.compile("Signup weight", re.IGNORECASE)).columns[0]
]
elif len(
test_df.filter(
regex=re.compile("What is your current weight", re.IGNORECASE)
).columns
):
test_df["Week 0"] = test_df[
test_df.filter(
regex=re.compile("What is your current weight", re.IGNORECASE)
).columns[0]
]
# Height
test_df["Height"] = test_df[
test_df.filter(regex=re.compile("Height", re.IGNORECASE)).columns[0]
]
# Highest Weight
if len(test_df.filter(regex=re.compile("Highest", re.IGNORECASE)).columns):
test_df["Highest Weight"] = test_df[
test_df.filter(regex=re.compile("Highest", re.IGNORECASE)).columns[0]
]
else:
test_df["Highest Weight"] = np.NaN
# Has NSV
test_df["Has NSV"] = (
test_df[test_df.filter(regex=re.compile("NSV", re.IGNORECASE)).columns[0]]
.notnull()
.astype("int")
)
test_df["NSV Text"] = (
test_df[test_df.filter(regex=re.compile("NSV", re.IGNORECASE)).columns[0]]
.astype(str)
.replace("nan", "")
)
# Goal Weight
test_df["Challenge Goal Weight"] = test_df[
test_df.filter(regex=re.compile("Goal Weight", re.IGNORECASE)).columns[0]
]
# Has a food tracker
if len(test_df.filter(regex=re.compile("MyFitnessPal", re.IGNORECASE)).columns):
test_df["MFP"] = (
test_df[
test_df.filter(regex=re.compile("MyFitnessPal", re.IGNORECASE)).columns[
0
]
]
.notnull()
.astype("int")
)
test_df["Has MFP"] = (
test_df[test_df.filter(regex=re.compile("MFP", re.IGNORECASE)).columns[0]]
.notnull()
.astype("int")
)
if len(test_df.filter(regex=re.compile("Loseit", re.IGNORECASE)).columns):
test_df["Has Loseit"] = (
test_df[
test_df.filter(regex=re.compile("Loseit", re.IGNORECASE)).columns[0]
]
.notnull()
.astype("int")
)
else:
test_df["Has Loseit"] = 0
test_df["Has Food Tracker"] = test_df["Has MFP"] + test_df["Has Loseit"]
test_df["Has Food Tracker"] = test_df["Has Food Tracker"].replace(2, 1)
# fitness tracker
if len(test_df.filter(regex=re.compile("Fitbit", re.IGNORECASE)).columns):
test_df["Has Activity Tracker"] = (
test_df[
test_df.filter(regex=re.compile("Fitbit", re.IGNORECASE)).columns[0]
]
.notnull()
.astype("int")
)
elif len(
test_df.filter(regex=re.compile("Fitness tracker", re.IGNORECASE)).columns
):
test_df["Has Activity Tracker"] = (
test_df[
test_df.filter(
regex=re.compile("Fitness Tracker", re.IGNORECASE)
).columns[0]
]
.notnull()
.astype("int")
)
elif len(test_df.filter(regex=re.compile("Garmin", re.IGNORECASE)).columns):
test_df["Has Activity Tracker"] = (
test_df[
test_df.filter(regex=re.compile("Garmin", re.IGNORECASE)).columns[0]
]
.notnull()
.astype("int")
)
elif len(test_df.filter(regex=re.compile("Strava", re.IGNORECASE)).columns):
test_df["Has Activity Tracker"] = (
test_df[
test_df.filter(regex=re.compile("Strava", re.IGNORECASE)).columns[0]
]
.notnull()
.astype("int")
)
# Team and Challenge Names
test_df["Challenge"] = (
challenge_name.replace("_", " ")
.title()
.replace("'", "")
.replace("Tracker", "")
.replace("Master", "")
.replace("Sign Ups", "")
.replace("Spreadsheet", "")
.replace("Loseit", "")
.replace("Challenge", "")
.replace("Edition", "")
.replace(" ", " ")
.strip()
+ " Challenge"
)
test_df["Team"] = test_df["Team"].str.title()
test_df["Team"] = test_df["Team"].str.replace("2Nd", "2nd")
# Starting Weight
test_df["Starting Weight"] = test_df["Week 0"]
# Create the final Data Frame
col_weeks = test_df.filter(regex=re.compile("Week", re.IGNORECASE)).columns.tolist()
col_weeks.sort(key=natural_keys)
col_names = [
"Timestamp",
"Username",
"Team",
"Challenge",
"Age",
"Gender",
"Height",
"Highest Weight",
"Starting Weight",
"Challenge Goal Weight",
"Starting BMI",
"Has NSV",
"Has Food Tracker",
"Has Activity Tracker",
"NSV Text",
]
data_cols = col_names + list(col_weeks)
data_df = test_df[data_cols]
df_list.append((challenge.stem, data_df))
# Now that the data contains only what we are interested in learning, we need to fill in any missing values before we combine all of the challenges together.
# +
big_df_list = []
for data in df_list:
df = data[1].copy()
# Some odties in the data
if data[0] == "spring_time_to_energize_challenge":
df.drop([448, 828], inplace=True)
df.replace({"ERROR": np.NaN}, inplace=True)
if data[0] == "autumn_animal_challenge":
df.drop(971, inplace=True)
df.replace({"#DIV/0!": np.NaN, "old": np.NaN}, inplace=True)
if data[0] == "rebirth_challenge_2017":
df.drop(["Week 7", "Week 8"], axis=1, inplace=True)
df.replace({"20s": 25, "Yes": np.NaN}, inplace=True)
df.Timestamp = statistics.mode(df.Timestamp)
df.dropna(subset=["Username", "Challenge Goal Weight"], axis=0, inplace=True)
df.loc[pd.isnull(df["Gender"]), "Gender"] = "Unknown"
df.loc[~df["Gender"].isin(["Female", "Male", "Unknown"]), "Gender"] = "Other"
df.loc[pd.isnull(df["Highest Weight"]), "Highest Weight"] = df["Week 0"]
df["Timestamp"] = df["Timestamp"].fillna(axis=0, method="ffill", limit=10)
# Now we want to convert the series into the correct types
numberic = [
"Age",
"Height",
"Highest Weight",
"Starting Weight",
"Challenge Goal Weight",
"Starting BMI",
]
df[numberic] = df[numberic].astype(np.float64)
"""
Now we need to work on removing those who dropped out of the challenge.
First, if only one weigh-in was missed we will fill it with the previous weeks
weigh-in. Next, we remove any that are missing the final weigh-in, and lastly,
we fill any of the remaining missing values with the previous weeks data.
"""
weight_cols = df.columns.values[15:].tolist()
df[weight_cols] = df[weight_cols].fillna(axis=1, method="ffill", limit=1)
df.dropna(axis=0, subset=[weight_cols[-1]], inplace=True)
df[weight_cols] = df[weight_cols].fillna(axis=1, method="ffill").astype(np.float64)
new_cols = [
"Final Weight",
"Total Challenge Loss",
"Challenge Percentage Lost",
"Percent of Challenge Goal",
]
df["Challenge Goal Loss"] = df["Starting Weight"].astype(np.float64) - df[
"Challenge Goal Weight"
].astype(np.float64)
df[new_cols[0]] = df[weight_cols[-1]]
df[new_cols[1]] = df[weight_cols[0]] - df[weight_cols[-1]]
df[new_cols[2]] = (df[new_cols[1]] / df[weight_cols[0]]) * 100
df[new_cols[3]] = (
df[new_cols[1]]
/ (
df["Starting Weight"].astype(np.float64)
- df["Challenge Goal Weight"].astype(np.float64)
)
).replace(np.inf, 0).replace(-np.inf, 0) * 100
df[new_cols] = df[new_cols].astype(np.float64)
df = df[df.columns.values[:15].tolist() + ["Challenge Goal Loss"] + new_cols]
# Save the cleaned data and append to the dataframe list
df.to_csv("../data/cleaned_data/cleaned_" + data[0] + ".csv")
big_df_list.append(df)
# -
big_df = pd.concat(big_df_list, ignore_index=True).dropna()
big_df.to_csv("../data/processed_data/cleaned_and_combined_loseit_challenge_data.csv")
# Now that we have the data saved and cleaned, we can move onto [Inspect Challenge Data](02_inspect_loseit_challenge_data.ipynb) to look a little bit deeper into the data to see if there are any outliers and how to possibly deal with them.
| notebooks/01_clean_loseit_challenge_data.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ls
# cd datas
# ls
from sklearn import linear_model
import numpy as np
import pandas as pd
import matplotlib
import matplotlib.pyplot as plt
# %matplotlib inline
train = pd.read_csv('train.csv')
train.head(1)
train["dong"].count()
df = pd.DataFrame(train, columns=['dong'])
df
day_care_center = pd.read_csv('./day_care_center.csv')
day_care_center.head(1)
park = pd.read_csv('./park.csv')
park.head(1)
| real_estate_forecast.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Question Answering with DeepMatcher
#
# Note: you can run **[this notebook live in Google Colab](https://colab.research.google.com/github/anhaidgroup/deepmatcher/blob/master/examples/question_answering.ipynb)**.
#
# DeepMatcher can be easily be used for text matching tasks such Question Answering, Text Entailment, etc. In this tutorial we will see how to use DeepMatcher for Answer Selection, a major sub-task of Question Answering. Specifically, we will look at [WikiQA](https://aclweb.org/anthology/D15-1237), a benchmark dataset for Answer Selection. There are three main steps in this tutorial:
#
# 1. Get data and transform it into DeepMatcher input format
# 2. Setup and train DeepMatcher model
# 3. Evaluate model using QA eval metrics
#
# Before we begin, if you are running this notebook in Colab, you will first need to install necessary packages by running the code below:
try:
import deepmatcher
except:
# !pip install -qqq deepmatcher
# ## Step 1: Get data and transform it into DeepMatcher input format
#
# First let's import relevant packages and download the dataset:
# +
import deepmatcher as dm
import pandas as pd
import os
# !wget -qnc https://download.microsoft.com/download/E/5/F/E5FCFCEE-7005-4814-853D-DAA7C66507E0/WikiQACorpus.zip
# !unzip -qn WikiQACorpus.zip
# -
# Let's see how this dataset looks like:
raw_train = pd.read_csv(os.path.join('WikiQACorpus', 'WikiQA-train.txt'), sep='\t', header=None)
raw_train.head()
# Clearly, it is not in the format `deepmatcher` wants its input data to be in - this file has no column names, no ID column, and its not a CSV file. Let's fix that:
raw_train.columns = ['left_value', 'right_value', 'label']
raw_train.index.name = 'id'
raw_train.head()
# Looks good, now let's save this to disk and transform the validation and test data in the same way:
# +
raw_train.to_csv(os.path.join('WikiQACorpus', 'dm_train.csv'))
raw_files = ['WikiQA-dev.txt', 'WikiQA-test.txt']
csv_files = ['dm_valid.csv', 'dm_test.csv']
for i in range(2):
raw_data = pd.read_csv(os.path.join('WikiQACorpus', raw_files[i]), sep='\t', header=None)
raw_data.columns = ['left_value', 'right_value', 'label']
raw_data.index.name = 'id'
raw_data.to_csv(os.path.join('WikiQACorpus', csv_files[i]))
# -
# ## Step 2: Setup and train DeepMatcher model
#
# Now we are ready to load and process the data for `deepmatcher`:
train, validation, test = dm.data.process(
path='WikiQACorpus',
train='dm_train.csv',
validation='dm_valid.csv',
test='dm_test.csv')
# Next, we create a `deepmatcher` model and train it. Note that since this is a demo, we do not perform hyperparameter tuning - we simply use the default settings for everything except the `pos_neg_ratio` param. This must be set since there are very few "positive matches" (candidates that correctly answer the question) in this dataset. In a real application setting you must tune other model hyperparameters as well to get optimal performance.
model = dm.MatchingModel()
model.run_train(
train,
validation,
epochs=10,
best_save_path='hybrid_model.pth',
pos_neg_ratio=7)
# Now that we have a trained model, we obtain the predictions for the test data. Note that `deepmatcher` computes F1, precision and recall by default but these may not be optimal evaluation metrics for your end task. For instance, in Question Answering, the more relevant metrics are MAP and MRR which we will compute in the next step.
predictions = model.run_prediction(test, output_attributes=True)
# ## Step 3: Evaluate model using QA eval metrics
# Finally, we compute the Mean Average Precision (MAP) and Mean Reciprocal Rank (MRR) using the model's predictions on the test set. Following the approach of the [paper that introduced this dataset](https://aclweb.org/anthology/D15-1237), questions in the test set without answers are ignored when computing these metrics.
# +
MAP, MRR = 0, 0
grouped = predictions.groupby('left_value')
num_questions = 0
for question, answers in grouped:
sorted_answers = answers.sort_values('match_score', ascending=False)
p, ap = 0, 0
top_answer_found = False
for idx, answer in enumerate(sorted_answers.itertuples()):
if answer.label == 1:
if not top_answer_found:
MRR += 1 / (idx + 1)
top_answer_found = True
p += 1
ap += p / (idx + 1)
if p > 0:
ap /= p
num_questions += 1
MAP += ap
MAP /= num_questions
MRR /= num_questions
print('MAP:', MAP)
print('MRR:', MRR)
# -
| examples/question_answering.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda env:gawler] *
# language: python
# name: conda-env-gawler-py
# ---
# ### Where <NAME>?
# In this notebook, we'll be using [H2o's AutoMl](https://docs.h2o.ai/h2o/latest-stable/h2o-docs/automl.html) algorithm to train our first **Binary Classification Model** on the data we prepared in our earlier notebook.
#
# Before we begin, make sure you install h2o in your system. You can refer to the instalation page [here](https://docs.h2o.ai/h2o/latest-stable/h2o-docs/downloading.html#install-in-python).
#
# You can also train your own models on the prepared data using State-Of-The-Art Algorithms such as
# - Tensorflow & Keras
# - Sklearn
# - Pytorch
# - MLBox
# - AutoKeras & Hyperas
# - AutoSkLearn etc.
# +
# Importing the required library - h2o - for data importing & modelling.
import h2o
from h2o.automl import H2OAutoML
# Initializing a local server to train our model upon. Make sure you adjust the Mem_Size as per your system.
# You can always look it up in the documentation to learn more!
h2o.init()
# +
# Importing the file into an H2o dataframe.
df = h2o.import_file('/home/xavian/Downloads/The_Gawler_Challenge/final_files/Model1/merged/Model1_Merged.csv')
# We need to split the data into train and test. Change the ratio as per your preference.
splits = df.split_frame(ratios=[0.8],seed=1)
train = splits[0]
test = splits[1]
# -
train.head()
# ### Specifying x & y
# Now that we have the data loaded, we need to specify `x` & `y` (in other words - training features & lables). A cool feature about H2o's AutoML is that we only need to specify the column names for x & y along with an H2o dataframe (the one that we loaded above) for training.
#
# This saves us a lot of time in converting a dataframe to an array and then reshaping it to fit our model structure. So without any further ado let's proceed with the training!
#
# 
#
#
# Specifying the name of the y column
y = "MINERAL_CL"
x = train.columns
x
# +
# Remove the columns we need to drop from our training frame.
r_rows = ['C1',
'LONGITUDE_',
'LATITUDE_G',
'MINERAL_CL']
for i in range(len(r_rows)):
# print(r_rows[i])
x.remove(r_rows[i])
# -
aml = H2OAutoML(max_runtime_secs=60*20, seed=1)
aml.train(x=x,y=y, training_frame=train)
lb = aml.leaderboard
lb.head()
aml.leaderboard
# Get model ids for all models in the AutoML Leaderboard
model_ids = list(aml.leaderboard['model_id'].as_data_frame().iloc[:,0])
# Get the "All Models" Stacked Ensemble model
se = h2o.get_model([mid for mid in model_ids if "StackedEnsemble_AllModels" in mid][0])
# Get the Stacked Ensemble metalearner model
metalearner = h2o.get_model(se.metalearner()['name'])
h2o.save_model(aml.leader, path="./saved_model")
import matplotlib
# %matplotlib inline
metalearner.std_coef_plot()
y_pred = aml.predict(train)
y_pred = y_pred['predict'].as_data_frame().values.reshape(len(y_pred))
y_true=train['MINERAL_CL'].as_data_frame().values
y_true = y_true.reshape(len(y_true))
y_pred[0] == y_true[0]
# +
a = 0
for i in range(len(y_true)):
if y_pred[i] == y_true[i]:
a+=1
print(a/len(y_true))
# -
pred['predict'].as_data_frame().values
model_path = h2o.save_model(model=aml, path="/saved_model_20_min", force=True)
m.varimp(use_pandas=True)
| models/Model1/Model1_training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Attempting to Correlate Historical Air Quality Index with BOM Weather data
# ## Data Prep and Exploration
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style('whitegrid')
# %matplotlib inline
aqi = pd.read_csv('../data/AQI_Correlate_final_v2.csv')
aqi.head(5)
# ### clean up
print(max(aqi['Quality Min']), min(aqi['Quality Min']))
# n/a in the measurements of precipitation in particular is not good data as we expect precipitation to be important.. drop it
aqi = aqi.dropna()
# quality scores and stn number don't change, product code adds no value
aqi.drop(['Quality Min','Product code','Bureau of Meteorology station number', 'Quality', 'Quality Min', 'Quality Max', 'Period over which rainfall was measured (days)', 'Days of accumulation of minimum temperature', 'Days of accumulation of maximum temperature'], axis=1, inplace=True)
aqi.rename(index=str, columns={"Rainfall amount (millimetres)": "precip",
"Minimum temperature (Degree C)": "temp_min",
"Maximum temperature (Degree C)": "temp_max",
"Randwick_AQI": "aqi",
"Date": "date"
}, inplace=True)
# ### Utility functions for dates and seasons
# +
import dateutil.parser as dparser
# utility functions for frigging around with dates and seasons
def getDateFromDateText(dateText):
return dparser.parse(dateText,dayfirst=True)
def getSeasonFromDate(date):
seasons = {12:'summmer', 1:'summer', 2:'summer', 3: 'autumn', 4: 'autumn', 5: 'autumn', 6: 'winter', 7: 'winter', 8: 'winter', 9: 'spring', 10: 'spring', 11: 'spring'}
return seasons[date.month]
def getSeasonFromDateText(dateText):
return getSeasonFromDate(getDateFromDateText(dateText))
print('I\'m like totally going Paleo so I\'m ready for {}'.format(getSeasonFromDateText('4/02/2016')))
# -
# ### Use the season function and get_dummies to create an indicator field
# apply the season function to the df and append the resultant series to the df as a new field
seasons = aqi['date'].apply(getSeasonFromDateText)
seasons_indicator = pd.get_dummies(seasons, columns=['Season'])
aqi = pd.concat([aqi, seasons_indicator], axis=1)
# ### create a categorical variable from the AQI score
# +
def getAQISeverityFromAQI(_aqi):
if _aqi < 50:return 'vgood';
elif _aqi < 100:return 'good';
elif _aqi < 150: return 'poor';
elif _aqi < 200: return 'danger';
else: return 'extreme';
aqi['aqi_cat'] = aqi['aqi'].apply(getAQISeverityFromAQI)
# -
print('number of samples: {}'.format(len(aqi)))
aqi.head()
# ### Model
# from sklearn import svm
from sklearn.ensemble import RandomForestClassifier
from sklearn.model_selection import train_test_split
# +
from pprint import pprint
def trainAndTestRandForrest(X, y, predictCol):
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.25, random_state=69)
# get a model
clf = RandomForestClassifier(n_jobs=2)
# clf = svm.SVC()
# train the model
clf.fit(X_train, y_train)
# run test set forward through the model and print out some info
predictions = clf.predict(X_test)
getError(predictions, y_test, predictCol)
# random forest is cool because it tells you what was important
importance_list = list(zip(X_train, clf.feature_importances_))
sorted_by_importance = sorted(importance_list, key=lambda tup: tup[1], reverse=True)
pprint(sorted_by_importance)
def getError(predictions, y_test, predictCol):
totalError = 0.0
totalCorrect = 0
for prediction, y in zip(predictions, y_test):
print('prediction: {}, actual: {}'.format(prediction, y))
if prediction ==y:
totalCorrect += 1
print('accuracy: {}%', 100 * totalCorrect / len(y_test))
# -
# ### Train and test the model
X = aqi[['precip', 'temp_min', 'temp_max', 'autumn', 'spring','summer', 'winter']]
y = aqi['aqi_cat']
trainAndTestRandForrest(X, y, 'aqi_cat')
| model/Correllate AQI with Weather.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# %matplotlib inline
import pandas as pd
import cycluster as cy
import os.path as op
import numpy as np
import palettable
from custom_legends import colorLegend
import seaborn as sns
from hclusterplot import *
import matplotlib
import matplotlib.pyplot as plt
import pprint
# +
sns.set_context('paper')
path = "./"
inf = "NICU_PED_Nurses_JC.csv"
dataFilename = op.join(path,inf)
"""A long df has one analyte measurement per row"""
longDf = pd.read_csv(dataFilename)
print(longDf)
# -
longDf['Groups']=longDf['ID'].astype(str)+'_'+longDf['Group']# longDf = longDf.drop(columns= ['ID', 'Influenza.Status', 'Strain', 'Age', 'Sex', 'CMV.Status', 'EBV.Status', 'HSV1_2.Status', 'HHV6.Status', 'VZV.Status'])
longDf
# longDf = longDf.drop(columns= ["IL12p40", "IL10"])
longDf
Df = longDf.pivot_table(index='Groups')
Df.to_excel('Example_2.xlsx')
# tmp.columns[np.isclose(tmp.std(), 0), rtol, atol].tolist()
print(np.isclose(Df.std(), 0))
# +
"""Identify primary day for clustering"""
# df = longDf.set_index(['ptid', 'dpi','cytokine'])['log10_conc'].unstack(['cytokine','dpi'])
# plt.plot([0, 3, 6, 9, 12], df['ifng'].values.T, '-o')
Percent = 20 ## All cytokines with greater(exclusive) percentage of ratio NA : all values will not be considered
rtol = None ## Add tolerance values for threshold variance for cytokine values to be considered
atol = None
"""A wide df has one sample per row (analyte measurements across the columns)"""
def _prepCyDf(tmp, K=3, normed=False, cluster="Cluster", percent= 0, rtol= None, atol= None):
# dayDf = longDf
# tmp = tmp.pivot_table(index='ptid', columns='cytokine', values='log10_conc')
if rtol or atol == None:
noVar = tmp.columns[np.isclose(tmp.std(), 0)].tolist()
else:
noVar = tmp.columns[np.isclose(tmp.std(), 0), rtol, atol].tolist()
naCols = tmp.columns[(tmp.isnull().sum()) / (((tmp.isnull()).sum()) + (tmp.notnull().sum())) > (percent / 100)].tolist() + ["IL12p40", "IL10"]
keepCols = [c for c in tmp.columns if not c in (noVar + naCols)]
# dayDf = dayDf.pivot_table(index='ptid', columns='cytokine', values='log10_conc')[keepCols]
"""By setting normed=True the data our normalized based on correlation with mean analyte concentration"""
tmp = tmp[keepCols]
rcyc = cy.cyclusterClass(studyStr='ADAMTS', sampleStr=cluster, normed=normed, rCyDf=tmp)
rcyc.clusterCytokines(K=K, metric='spearman-signed', minN=0)
rcyc.printModules()
return rcyc
test = _prepCyDf(Df, K=3, normed=False, cluster="All", percent= 10)
# +
"""Now you can use attributes in nserum for plots and testing: cyDf, modDf, dmatDf, etc."""
plt.figure(41, figsize=(15.5, 9.5))
colInds = plotHColCluster(rcyc.cyDf,
method='complete',
metric='pearson-signed',
col_labels=rcyc.labels,
col_dmat=rcyc.dmatDf,
tickSz='large',
vRange=(0,1))
plt.figure(43, figsize = (15.5, 9.5))
colInds = cy.plotting.plotHierClust(1 - rcyc.pwrel,
rcyc.Z,
labels=rcyc.labels,
titleStr='Pairwise reliability (%s)' % rcyc.name,
vRange=(0, 1),
tickSz='large')
plt.figure(901, figsize=(13, 9.7))
cy.plotting.plotModuleEmbedding(rcyc.dmatDf, rcyc.labels, method='kpca', txtSize='large')
colors = palettable.colorbrewer.get_map('Set1', 'qualitative', len(np.unique(rcyc.labels))).mpl_colors
colorLegend(colors, ['%s%1.0f' % (rcyc.sampleStr, i) for i in np.unique(rcyc.labels)], loc='lower left')
# -
# +
import scipy.stats
"""df here should have one column per module and the genotype column"""
ptidDf = longDf[['ptid', 'sample', 'genotype', 'dpi']].drop_duplicates().set_index('ptid')
df = rcyc.modDf.join(ptidDf)
ind = df.genotype == 'WT'
col = 'LUNG1'
# stats.ranksums(df[col].loc[ind], df[col].loc[~ind])
# -
| .ipynb_checkpoints/Cycluster_2-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + colab_type="code" id="mAxBj2nDelo6" colab={}
# #!/usr/bin/env python2
# -*- coding: utf-8 -*-
"""
Created on Sat Mar 23 13:20:15 2019
@author: chin-weihuang
"""
from __future__ import print_function
import numpy as np
import torch
import torch.nn as nn
import torch.optim as optim
import matplotlib.pyplot as plt
import torch.nn.functional as F
import samplers
import jsloss
from jsloss import Discriminator as Discriminator
from jsloss import JSD as JSD
from wgan_gp_code import Critic, WGAN_GPloss
# + colab_type="code" id="0BDQlSMSgK59" outputId="20d61a48-a10c-4a2c-fffc-4b6eda3b1e46" colab={"base_uri": "https://localhost:8080/", "height": 301}
#Question 1.3
output_list = []
epoch_count_js = 500
epoch_count_wgan = 12000
#loop over the values between -1 and 1 for (phi, U(0,1))
for phi in np.linspace(-1,1,21):
jsd = JSD(x_val=phi, minibatch_size=512, epoch_count=epoch_count_js, learning_rate=1e-3, \
input_size=2, hidden_size=10, output_size=1, \
real_sampler = samplers.distribution1, \
fake_sampler = samplers.distribution1 \
)
output_list.append(jsd)
wgan_output_list = []
losses = []
for phi in np.linspace(-1,1,21):
wgan_gp = WGAN_GPloss(x_val=phi, minibatch_size=512, epoch_count=epoch_count_wgan, learning_rate=1e-3, losses = losses,\
input_size=2, hidden_size=10, output_size=1, \
real_sampler = samplers.distribution1, \
fake_sampler = samplers.distribution1 \
)
wgan_output_list.append(wgan_gp)
torch.stack(output_list)
plt.figure()
plt.scatter(np.linspace(-1,1,21), \
torch.stack(output_list).cpu().detach().numpy(), label = "Jensen-Shannon Divergence")
plt.scatter(np.linspace(-1,1,21), \
torch.stack(wgan_output_list).cpu().detach().numpy(), label = "WGAN_GP distance")
plt.xlabel('Values of $\phi \in [-1,1]$ with intervals of 0.1 ' )
plt.ylabel('JS Divergence')
plt.title('Jensen Shanon divergence between (0,U[0,1]) and ($\phi$,U[0,1])')
plt.legend()
plt.show()
# + [markdown] colab_type="text" id="E72UBjVLoxmS"
# The Jensen-Shannon Divergence at $ \phi = 0$ is 0 (overlayed with WGAN_GP distance)
| Generatives_VAEs & GANs/Q1/Q1_3.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# ---
# %load_ext autoreload
# %autoreload 2
# +
# %matplotlib inline
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
# -
# Read information to connect to the database and put it in environment variables
import os
with open('ENVVARS.txt') as f:
for line in f:
parts = line.split('=')
if len(parts) == 2:
os.environ[parts[0]] = parts[1].strip()
db_name = 'ticclat_test'
os.environ['dbname'] = db_name
# +
from ticclat.dbutils import create_ticclat_database
create_ticclat_database(delete_existing=True, dbname=os.environ['dbname'], user=os.environ['user'], passwd=os.environ['password'])
# +
from ticclat.ticclat_schema import Lexicon, Wordform, Anahash
from ticclat.dbutils import get_session, session_scope
Session = get_session(os.environ['user'], os.environ['password'], os.environ['dbname'])
# +
# add two lexicons
from ticclat.dbutils import add_lexicon
name1 = 'l1'
wfs1 = pd.DataFrame()
wfs1['wordform'] = ['wf1', 'wf2', 'wf3']
name2 = 'l2'
wfs2 = pd.DataFrame()
wfs2['wordform'] = ['wf2', 'wf3', 'wf4']
with session_scope(Session) as session:
lex1 = add_lexicon(session, lexicon_name=name1, vocabulary=True, wfs=wfs1)
lex2 = add_lexicon(session, lexicon_name=name2, vocabulary=True, wfs=wfs2)
# +
# add a corpus
from ticclat.tokenize import terms_documents_matrix_counters
from ticclat.sacoreutils import add_corpus_core
name = 'corpus1'
documents = [['wf1', 'wf2'], ['wf2', 'wf3'], ['wf4', 'wf5', 'wf6']]
corpus_matrix, vectorizer = terms_documents_matrix_counters(documents)
print(corpus_matrix.shape)
print(vectorizer.vocabulary_)
metadata = pd.DataFrame()
metadata['title'] = ['doc1', 'doc2', 'doc3']
metadata['pub_year'] = [2018, 2011, 2019]
with session_scope(Session) as session:
add_corpus_core(session, corpus_matrix, vectorizer, name, metadata)
# +
# add another corpus
from ticclat.tokenize import terms_documents_matrix_counters
from ticclat.sacoreutils import add_corpus_core
name = 'corpus2'
documents = [['wf2', 'wf5'], ['wf4', 'wf5', 'wf6']]
corpus_matrix, vectorizer = terms_documents_matrix_counters(documents)
print(corpus_matrix.shape)
print(vectorizer.vocabulary_)
metadata = pd.DataFrame()
metadata['title'] = ['doc4', 'doc5']
metadata['pub_year'] = [2002, 2011]
with session_scope(Session) as session:
add_corpus_core(session, corpus_matrix, vectorizer, name, metadata)
# +
# add another corpus
from ticclat.tokenize import terms_documents_matrix_counters
from ticclat.sacoreutils import add_corpus_core
name = 'corpus3'
documents = [['wf2', 'wf5'], ['wf2', 'wf3', 'wf6'], ['wf2']]
corpus_matrix, vectorizer = terms_documents_matrix_counters(documents)
print(corpus_matrix.shape)
print(vectorizer.vocabulary_)
metadata = pd.DataFrame()
metadata['title'] = ['doc6', 'doc7', 'doc8']
metadata['pub_year'] = [2002, 2011, 2018]
with session_scope(Session) as session:
add_corpus_core(session, corpus_matrix, vectorizer, name, metadata)
# +
from ticclat.ticclat_schema import Lexicon, Wordform, Anahash, Corpus, Document, TextAttestation
with session_scope(Session) as session:
print('number of wordforms:', session.query(Wordform).count())
print('number of lexica:', session.query(Lexicon).count())
print('number of corpora:', session.query(Corpus).count())
print('number of documents:', session.query(Document).count())
print('number of text attestations:', session.query(TextAttestation).count())
# -
| notebooks/create_test_database.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# Import models
import numpy as np
from keras.models import Sequential
from keras.layers import Dense, Activation
from keras.utils.vis_utils import plot_model
# +
# Create model
model = Sequential()
model.add(Dense(32, input_dim = 500))
model.add(Activation(activation="sigmoid"))
model.add(Dense(1))
model.add(Activation(activation="sigmoid"))
# Compile model
model.compile(optimizer='rmsprop',
loss='binary_crossentropy',
metrics=['accuracy'])
# -
# Create dataset
data = np.random.random((1000,500))
labels = np.random.randint(2,size=(1000,1))
# Calculate score
score = model.evaluate(data,labels,verbose=1)
print("Score before training = {}".format(
list(zip(model.metrics_names,score))
))
# Fit model
model.fit(
data,labels, epochs=1000,
batch_size=32,
verbose=1
)
# Score after training
score = model.evaluate(data,labels,verbose=1)
print("Score after training = {}".format(
list(zip(model.metrics_names, score))
))
plot_model(model, to_file='keras_nn_2.png',show_shapes=True)
import matplotlib.pyplot as plt
from cv2 import imread
img = imread('keras_nn_2.png')
plt.imshow(img)
| Keras 2 Layer Neural Network.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import numpy as np
from keras.models import *
from keras import backend as K
from keras.preprocessing.image import ImageDataGenerator
from models.unet import *
from models.unet_se import *
from datahandler import DataHandler
from kfold_data_loader import *
from params import *
import os
import cv2
import skimage.io as io
from tqdm import tqdm
from medpy.io import save
from math import ceil, floor
from matplotlib import pyplot as plt
from sklearn.metrics import f1_score, jaccard_similarity_score
from scipy.ndimage import _ni_support
from scipy.ndimage.morphology import distance_transform_edt, binary_erosion,\
generate_binary_structure
import warnings
warnings.filterwarnings("ignore")
plt.gray()
# -
def destiny_directory(dice_score):
pre = './data/eval/unet_se/'
if dice_score >= 98:
return pre + 'dice_98_100/'
elif dice_score >= 96:
return pre + 'dice_96_98/'
elif dice_score >= 94:
return pre + 'dice_94_96/'
elif dice_score >= 92:
return pre + 'dice_92_94/'
elif dice_score >= 90:
return pre + 'dice_90_92/'
elif dice_score >= 88:
return pre + 'dice_88_90/'
elif dice_score >= 85:
return pre + 'dice_85_88'
elif dice_score >= 80:
return pre + 'dice_80_85/'
elif dice_score >= 70:
return pre + 'dice_70_80/'
elif dice_score >= 60:
return pre + 'dice_60_70/'
else:
return pre + 'dice_less_60'
def getGenerator(images, bs=1):
image_datagen = ImageDataGenerator(rescale=1./255)
image_datagen.fit(images, augment = True)
image_generator = image_datagen.flow(x = images, batch_size=bs,
shuffle = False)
return image_generator
def getDiceScore(ground_truth, prediction):
#convert to boolean values and flatten
ground_truth = np.asarray(ground_truth, dtype=np.bool).flatten()
prediction = np.asarray(prediction, dtype=np.bool).flatten()
return f1_score(ground_truth, prediction)
# +
def hd(result, reference, voxelspacing=None, connectivity=1):
hd1 = __surface_distances(result, reference, voxelspacing, connectivity).max()
hd2 = __surface_distances(reference, result, voxelspacing, connectivity).max()
hd = max(hd1, hd2)
return hd
def hd95(result, reference, voxelspacing=None, connectivity=1):
hd1 = __surface_distances(result, reference, voxelspacing, connectivity)
hd2 = __surface_distances(reference, result, voxelspacing, connectivity)
hd95 = np.percentile(np.hstack((hd1, hd2)), 95)
return hd95
def __surface_distances(result, reference, voxelspacing=None, connectivity=1):
result = np.atleast_1d(result.astype(np.bool))
reference = np.atleast_1d(reference.astype(np.bool))
if voxelspacing is not None:
voxelspacing = _ni_support._normalize_sequence(voxelspacing, result.ndim)
voxelspacing = np.asarray(voxelspacing, dtype=np.float64)
if not voxelspacing.flags.contiguous:
voxelspacing = voxelspacing.copy()
footprint = generate_binary_structure(result.ndim, connectivity)
if 0 == np.count_nonzero(result):
raise RuntimeError('The first supplied array does not contain any binary object.')
if 0 == np.count_nonzero(reference):
raise RuntimeError('The second supplied array does not contain any binary object.')
result_border = result ^ binary_erosion(result, structure=footprint, iterations=1)
reference_border = reference ^ binary_erosion(reference, structure=footprint, iterations=1)
dt = distance_transform_edt(~reference_border, sampling=voxelspacing)
sds = dt[result_border]
return sds
# +
image_files, mask_files = load_data_files('data/kfold_data/')
print(len(image_files))
print(len(mask_files))
skf = getKFolds(image_files, mask_files, n=10)
kfold_indices = []
for train_index, val_index in skf.split(image_files, mask_files):
kfold_indices.append({'train': train_index, 'val': val_index})
# -
def predictMask(model, image):
image_gen = getGenerator(image)
return model.predict_generator(image_gen, steps=len(image))
# +
def prepareForSaving(image):
image = np.squeeze(image)
image = np.moveaxis(image, -1, 0)
return image
def predictAll(model, data, num_data=0):
dice_scores = []
hd_scores = []
hd95_scores = []
for image_file, mask_file in tqdm(data, total=num_data):
fname = image_file[image_file.rindex('/')+1 : image_file.index('.')]
image, hdr = dh.getImageData(image_file)
gt_mask, _ = dh.getImageData(mask_file, is_mask=True)
assert image.shape == gt_mask.shape
if image.shape[1] != 256:
continue
pred_mask = predictMask(model, image)
pred_mask[pred_mask>=0.7] = 1
pred_mask[pred_mask<0.7] = 0
dice_score = getDiceScore(gt_mask, pred_mask)
if dice_score == 0:
continue
dice_scores.append(dice_score)
hd_score = hd(gt_mask, pred_mask)
hd_scores.append(hd_score)
hd95_score = hd95(gt_mask, pred_mask)
hd95_scores.append(hd95_score)
int_dice_score = floor(dice_score * 100)
save_path = destiny_directory(int_dice_score)
pred_mask = prepareForSaving(pred_mask)
image = prepareForSaving(image)
gt_mask = prepareForSaving(gt_mask)
save(pred_mask, os.path.join(save_path, fname + '_' + unet_type + '_'
+ str(int_dice_score) + '.nii'), hdr)
save(image, os.path.join(save_path, fname + '_img.nii'), hdr)
save(gt_mask, os.path.join(save_path, fname + '_mask.nii'), hdr)
return dice_scores, hd_scores, hd95_scores
# +
#Get data and generators
unet_type = 'unet_se'
dh = DataHandler()
all_dice = []
all_hd = []
all_hd95 = []
for i in range(len(kfold_indices)):
exp_name = 'kfold_%s_dice_DA_K%d'%(unet_type, i)
#get parameters
params = getParams(exp_name, unet_type=unet_type)
val_img_files = np.take(image_files, kfold_indices[i]['val'])
val_mask_files = np.take(mask_files, kfold_indices[i]['val'])
if unet_type == 'unet_se':
model = getSEUnet()
else:
model = getUnet()
print('loading weights from %s'%params['checkpoint']['name'])
model.load_weights(params['checkpoint']['name'])
data = zip(val_img_files, val_mask_files)
dice_score, hd_score, hd95_score = predictAll(model, data, num_data=len(val_mask_files))
print('Finished K%d'%i)
all_dice += dice_score
all_hd += hd_score
all_hd95 += hd95_score
print('dice')
for i in range(len(all_dice)):
print(all_dice[i])
print()
print('hd')
for i in range(len(all_hd)):
print(all_hd[i])
print()
print('hd95')
for i in range(len(all_hd95)):
print(all_hd95[i])
print()
print('Final results for %s'%unet_type)
print('dice %f'%np.mean(all_dice))
print('hd %f'%np.mean(all_hd))
print('hd95 %f'%np.mean(all_hd95))
| .ipynb_checkpoints/unet_se_kfold_eval-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Unique Morse Code Words
# International Morse Code defines a standard encoding where each letter is mapped to a series of dots and dashes, as follows: "a" maps to ".-", "b" maps to "-...", "c" maps to "-.-.", and so on.
#
# For convenience, the full table for the 26 letters of the English alphabet is given below:
#
# > [".-","-...","-.-.","-..",".","..-.","--.","....","..",".---","-.-",".-..","--","-.","---",".--.","--.-",".-.","...","-","..-","...-",".--","-..-","-.--","--.."]
#
# Now, given a list of words, each word can be written as a concatenation of the Morse code of each letter. For example, "cba" can be written as "-.-..--...", (which is the concatenation "-.-." + "-..." + ".-"). We'll call such a concatenation, the transformation of a word.
#
# Return the number of different transformations among all words we have.
#
# Example:
# Input: words = ["gin", "zen", "gig", "msg"]
# Output: 2
# Explanation:
# The transformation of each word is:
# "gin" -> "--...-."
# "zen" -> "--...-."
# "gig" -> "--...--."
# "msg" -> "--...--."
#
# There are 2 different transformations, "--...-." and "--...--.".
# Note:
#
# The length of words will be at most 100.
# Each words[i] will have length in range [1, 12].
# words[i] will only consist of lowercase letters.
#
# [Unique Morse Code Words](https://leetcode.com/problems/unique-morse-code-words/)
class Solution:
def uniqueMorseRepresentations(self, words):
"""
:type words: List[str]
:rtype: int
"""
# a->z
codeMap = [".-","-...","-.-.","-..",".","..-.","--.","....","..",".---","-.-",".-..","--","-.","---",".--.","--.-",".-.","...","-","..-","...-",".--","-..-","-.--","--.."]
aAscii = ord("a")
result = set()
for word in words:
code = ""
for char in word:
code += codeMap[ord(char) - aAscii]
result.add(code)
return len(result)
| UniqueMorseCodeWords.ipynb |
# -*- coding: utf-8 -*-
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .jl
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Julia 1.6.3
# language: julia
# name: julia-1.6
# ---
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-217de2f8df81af69", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ## Probabilistic Programming 2: Assignment
#
# In this assignment, we will look at one of the examples in the lecture on the Gaussian distribution. Consider the following factor graph:
#
# 
#
# The variables $x$ and $y$ are Gaussian distributed:
#
# $$\begin{align*}
# x \sim&\ \mathcal{N}(\mu_x,\Sigma_x) \\
# y \sim&\ \mathcal{N}(\mu_y,\Sigma_y) \, .
# \end{align*}$$
#
# Both are two-dimensional. The square node on the left is a vector addition operation, with $\xi_1$ representing the result. The middle square node is a multiplication operation with a clamped matrix $A = \begin{bmatrix} 2 & 0 \\ 0 & 1 \end{bmatrix}$ resulting in $\xi_2$. The right square node is a vector subtraction between $\xi_2$ and a clamped vector $b = \begin{bmatrix} 0 \\ 1 \end{bmatrix}$, producing the variable $z$.
# + nbgrader={"grade": false, "grade_id": "cell-0f4c86237a7e9708", "locked": true, "schema_version": 3, "solution": false, "task": false}
using Pkg
Pkg.activate("workspace/")
Pkg.instantiate();
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-1a26cb45dcb07124", "locked": true, "schema_version": 3, "solution": false, "task": false}
# ### **1) What are the parameters of the message going out of the left square node?**
#
# To answer this question, you'll need to specify a model in ForneyLab. The cell below already has a graph, some clamped parameters, the algorithm compilation and the algorithm execution code. Your job is add the two Gaussian variables $x$, $y$ and the resulting variable $ξ_1$ to the graph. You don't need to assign `:id`'s to these variables, you don't need `placeholder()`s and you don't have to specify a `PosteriorFactorization`.
#
# You can add variables to a graph using the `@RV` macro. Use `@RV var ~ ...` to define a stochastic variable (one that follows a particular distribution) and `@RV var = ...` for a deterministic variable. Options for distributions include `GaussianMeanVariance`, `GaussianMeanPrecision`, `Gamma`, `Wishart`, `Beta`, `Bernoulli`, `Categorical` and `Dirichlet`.
# + nbgrader={"grade": false, "grade_id": "cell-2b06871cebe3fe1d", "locked": false, "schema_version": 3, "solution": true, "task": false}
using ForneyLab
# Start factor graph
g = FactorGraph()
# Set parameters
μ_x = [1.0, 1.0]
Σ_x = [1.0 0.0;
0.0 1.0]
μ_y = [0.0, 0.0]
Σ_y = [2.0 0.0;
0.0 1.0]
### BEGIN SOLUTION
# Add variables
@RV x ~ GaussianMeanVariance(μ_x, Σ_x)
@RV y ~ GaussianMeanVariance(μ_y, Σ_y)
# Vector addition
@RV ξ_1 = x + y
### END SOLUTION
# Compile algorithm
algo = messagePassingAlgorithm(ξ_1)
source_code = algorithmSourceCode(algo)
eval(Meta.parse(source_code))
# Execute inference
messages = Array{Message}(undef, 3)
step!(Dict(), Dict(), messages)
println("Outgoing "*string(messages[3]))
# + nbgrader={"grade": true, "grade_id": "cell-e7c10e3a7fa570f7", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Check your answer
@assert dims(messages[3].dist) == 2
@assert typeof(messages[3]) == Message{GaussianMeanVariance,Multivariate}
### BEGIN HIDDEN TESTS
m = messages[3].dist.params[:m]
@assert sum(abs.(m .- [1.,1.])) < 1e-3
### END HIDDEN TESTS
# + nbgrader={"grade": true, "grade_id": "cell-d2682e2689228314", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Check your answer
@assert dims(messages[3].dist) == 2
@assert typeof(messages[3]) == Message{GaussianMeanVariance,Multivariate}
### BEGIN HIDDEN TESTS
V = messages[3].dist.params[:v]
@assert sum(abs.(V .- [3. 0.;0. 2.])) < 1e-3
### END HIDDEN TESTS
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-69cc7bf0ef19659d", "locked": true, "schema_version": 3, "solution": false, "task": false}
# Suppose $\xi_1$ follows a Gaussian distribution $\xi_1 \sim \mathcal{N}(\mu_{\xi_1}, \Sigma_{\xi_1})$. The middle square node will perform a matrix multiplication $\xi_2 = A \cdot \xi_1$.
#
# ### **2) What are the parameters of the message going out of the middle square node?**
#
# Add $ξ_1$ and $ξ_2$ as variables to the graph.
# + nbgrader={"grade": false, "grade_id": "cell-62055605a3bf0693", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Start factor graph
g = FactorGraph()
# Set parameters
μ_ξ_1 = [1.0, 1.0]
Σ_ξ_1 = [1.0 0.0;
0.0 1.0]
# Transition matrix
A = [2. 0.;
0. 1.]
### BEGIN SOLUTION
# Add variables
@RV ξ_1 ~ GaussianMeanVariance(μ_ξ_1, Σ_ξ_1)
# Vector multiplication
@RV ξ_2 = A*ξ_1
### END SOLUTION
# Compile algorithm
algo = messagePassingAlgorithm(ξ_2)
source_code = algorithmSourceCode(algo)
eval(Meta.parse(source_code))
# Execute inference
messages = Array{Message}(undef, 2)
step!(Dict(), Dict(), messages)
println("Outgoing "*string(messages[2]))
# + nbgrader={"grade": true, "grade_id": "cell-5e91072b62d16dfc", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Check your answer
@assert dims(messages[2].dist) == 2
@assert typeof(messages[2]) == Message{GaussianMeanVariance,Multivariate}
### BEGIN HIDDEN TESTS
m = messages[2].dist.params[:m]
@assert sum(abs.(m .- [2.,1.])) < 1e-3
### END HIDDEN TESTS
# + nbgrader={"grade": true, "grade_id": "cell-9bb8d03309ff5133", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Check your answer
@assert dims(messages[2].dist) == 2
@assert typeof(messages[2]) == Message{GaussianMeanVariance,Multivariate}
### BEGIN HIDDEN TESTS
V = messages[2].dist.params[:v]
@assert sum(abs.(V .- [4. 0.;0. 1.])) < 1e-3
### END HIDDEN TESTS
# + [markdown] nbgrader={"grade": false, "grade_id": "cell-d48b7f00ec1b96d0", "locked": true, "schema_version": 3, "solution": false, "task": false}
# The right square node is a vector subtraction, producing $z = \xi_2 - b$. Suppose the marginal for $\xi_2 \sim \mathcal{N}(\mu_{\xi_2}, \Sigma_{\xi_2})$.
#
# ### **3) What are the parameters for the message going out of the right node?**
#
# Add $\xi_2$ and $z$ as variables to the graph.
# + nbgrader={"grade": false, "grade_id": "cell-e30981f314233cd1", "locked": false, "schema_version": 3, "solution": true, "task": false}
# Start factor graph
g = FactorGraph()
# Set parameters
μ_ξ_2 = [1.0, 1.0]
Σ_ξ_2 = [1.0 0.0;
0.0 1.0]
# Transition matrix
b = [0., 1.]
### BEGIN SOLUTION
# Add variables
@RV ξ_2 ~ GaussianMeanVariance(μ_ξ_2, Σ_ξ_2)
# Vector subtraction
@RV z = ξ_2 - b
### END SOLUTION
# Compile algorithm
algo = messagePassingAlgorithm(z)
source_code = algorithmSourceCode(algo)
eval(Meta.parse(source_code))
# Execute inference
messages = Array{Message}(undef, 2)
step!(Dict(), Dict(), messages)
println("Outgoing "*string(messages[2]))
# + nbgrader={"grade": true, "grade_id": "cell-78542fd40598216b", "locked": true, "points": 1, "schema_version": 3, "solution": false, "task": false}
# Check your answer
@assert dims(messages[2].dist) == 2
@assert typeof(messages[2]) == Message{GaussianMeanVariance,Multivariate}
### BEGIN HIDDEN TESTS
m = messages[2].dist.params[:m]
V = messages[2].dist.params[:v]
@assert sum(abs.(m .- [1., 0.])) < 1e-3
@assert sum(abs.(V .- [1. 0.;0. 1.])) < 1e-3
### END HIDDEN TESTS
| lessons/exercises/probprog/solutions-pp2-exercises.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: conda_python3
# language: python
# name: conda_python3
# ---
# ## Training Notebook
#
# This notebook illustrates training of a simple model to classify digits using the MNIST dataset. This code is used to train the model included with the templates. This is meant to be a starter model to show you how to set up Serverless applications to do inferences. For deeper understanding of how to train a good model for MNIST, we recommend literature from the [MNIST website](http://yann.lecun.com/exdb/mnist/). The dataset is made available under a [Creative Commons Attribution-Share Alike 3.0](https://creativecommons.org/licenses/by-sa/3.0/) license.
# +
# We'll use scikit-learn to load the dataset
# ! pip install -q scikit-learn==0.23.2
# +
# Load the mnist dataset
from sklearn.datasets import fetch_openml
from sklearn.model_selection import train_test_split
X, y = fetch_openml('mnist_784', return_X_y=True)
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=10000)
# -
# ## Tensorflow Model Training
#
# For this example, we will train a simple CNN classifier using Tensorflow to classify the MNIST digits. We will then freeze the model in the `.h5` format. This is same as the starter model file included with the SAM templates.
# ! pip install -q tensorflow==2.4.0
# +
import numpy as np
import tensorflow as tf
print (f'Using TesorFlow version {tf.__version__}')
# Reshape the flat input into a 28x28x1 dim tensor
X_train = X_train.reshape(X_train.shape[0], 28, 28, 1)
X_test = X_test.reshape(X_test.shape[0], 28, 28, 1)
# Convert the output tensors to integers (our data is read as Strings)
y_train = y_train.astype(np.int8)
y_test = y_test.astype(np.int8)
model = tf.keras.Sequential([
# Input layer to match the shape above
tf.keras.layers.Conv2D(32, (3, 3), activation='relu', input_shape=(28, 28, 1)),
tf.keras.layers.MaxPooling2D((2, 2)),
tf.keras.layers.Conv2D(32, (3, 3), activation='relu'),
tf.keras.layers.MaxPooling2D((2, 2)),
tf.keras.layers.Flatten(),
tf.keras.layers.Dense(100, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(100, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(100, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(100, activation='relu'),
tf.keras.layers.BatchNormalization(),
tf.keras.layers.Dense(100, activation='relu'),
tf.keras.layers.BatchNormalization(),
# Output layer for 10 classes
tf.keras.layers.Dense(10)
])
model.compile(optimizer='adam',
loss=tf.keras.losses.SparseCategoricalCrossentropy(from_logits=True),
metrics=['accuracy'])
model.fit(X_train, y_train, epochs=15)
# +
test_loss, test_acc = model.evaluate(X_test, y_test, verbose=2)
print('\nTest accuracy:', test_acc)
# -
# Save model to the disk
model.save('tf_digit_classifier.h5')
| python3.8-image/cookiecutter-ml-apigw-tensorflow/{{cookiecutter.project_name}}/training.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#
# ## First steps:
#
# - Sample some points from a gaussian mixture
# - Fixed sample sizes (one big, one small), 5 different random seeds
# - Create both kde and normal histograms for some binning and bandwidth
# - Value of truth dist bin is area under curve between bin endpoints (just like kde hist!)
# - Make these plots for a range of bandwidths
#
# ### Expected behavior:
#
# - stdev of count estimate across random seeds decreases with more samples
# - for large bandwidth, you will see a bias error, as you're smoothing out the shape of the distribution
#
#
# +
import jax
import jax.numpy as jnp
from jax.random import normal, PRNGKey
rng = PRNGKey(7)
from matplotlib.colors import to_rgb
import matplotlib.pyplot as plt
plt.rc('figure',figsize=[7.3,5],dpi=120,facecolor='w')
from functools import partial
# -
# Let's generate `num_samples` points from a set of normal distributions with slowly increasing means:
lo, hi = -2, 2
grid_points = 500
mu_grid = jnp.linspace(lo, hi, grid_points)
num_samples = 100
points = jnp.tile(
normal(rng, shape = (num_samples,)),
reps = (grid_points,1)
) + mu_grid.reshape(-1,1)
points.shape
# Each index of `points` is a set of `num_samples` samples drawn for a given $\mu$ value. We want to make histograms for these sets of points, and then focus our attention on just one bin.
bins = jnp.linspace(lo-1,hi+1,6)
make_hists = jax.vmap(partial(jnp.histogram, bins = bins))
hists, _ = make_hists(points)
# We can start by inspecting a couple of these histograms to see the behaviour of varying $\mu$ upwards:
# +
centers = bins[:-1] + jnp.diff(bins) / 2.0
width = (bins[-1] - bins[0])/(len(bins) - 1)
fig, axs = plt.subplots(1,3)
# first mu value
axs[0].bar(
centers,
hists[0],
width = width,
label=f'$\mu$={mu_grid[0]}'
)
axs[0].legend()
axs[0].axis('off')
# middle mu value
axs[1].bar(
centers, hists[len(hists)//2],
width = width,
label=f'$\mu$={mu_grid[len(hists)//2]:.2f}',
color = 'C1'
)
axs[1].legend()
axs[1].axis('off')
# last mu value
axs[2].bar(
centers,
hists[-1],
width = width,
label=f'$\mu$={mu_grid[-1]}',
color = 'C2'
)
axs[2].legend()
axs[2].axis('off');
# -
# As one may expect, shifting $\mu$ to the right subsequently skews the resulting histogram.
# Now, let's focus on the behavior of the middle bin by plotting its height across a large range of $\mu$ values:
# +
# cool color scheme
from matplotlib.colors import to_rgb
def fade(c1,c2, num_points):
start = jnp.array(to_rgb(c1))
end = jnp.array(to_rgb(c2))
interp = jax.vmap(partial(jnp.linspace, num=num_points))
return interp(start,end).T
color_scheme = fade('C1', 'C3', num_points=grid_points)
# -
middle = len(bins)//2 - 1
mu_width = mu_grid[1]-mu_grid[0]
plt.bar(mu_grid, hists[:,middle], color=color_scheme, width=mu_width, edgecolor= 'black',linewidth = 0.05,alpha=0.7)
plt.xlabel('$\mu$');
# We can see that this bin goes up then down in value as expected, but it does so in a jagged, unfriendly way, meaning that the gradient of the bin height with respect to $\mu$ is also badly behaved. This gradient is crucial to evaluate if you want to do end-to-end optimization, since histograms are an extremely common component in high-energy physics.
#
# A solution to remedy this jaggedness can be found by changing the way we construct the histogram. In particular, we can perform a kernel density estimate for each set of samples, then discretize the result by partitioning the area under the curve with the same binning as we used to make the histogram.
# +
import jax.scipy as jsc
def kde_hist(events, bins, bandwidth=None, density=False):
edge_hi = bins[1:] # ending bin edges ||<-
edge_lo = bins[:-1] # starting bin edges ->||
# get cumulative counts (area under kde) for each set of bin edges
cdf_up = jsc.stats.norm.cdf(edge_hi.reshape(-1, 1), loc=events, scale=bandwidth)
cdf_dn = jsc.stats.norm.cdf(edge_lo.reshape(-1, 1), loc=events, scale=bandwidth)
# sum kde contributions in each bin
counts = (cdf_up - cdf_dn).sum(axis=1)
if density: # normalize by bin width and counts for total area = 1
db = jnp.array(jnp.diff(bins), float) # bin spacing
return counts / db / counts.sum(axis=0)
return counts
# -
# make hists as before
bins = jnp.linspace(lo-1,hi+1,6)
make_kde_hists = jax.vmap(partial(kde_hist, bins = bins, bandwidth = .5))
kde_hists = make_kde_hists(points)
# +
middle = len(bins)//2 - 1
mu_width = mu_grid[1]-mu_grid[0]
fig, axs = plt.subplots(2,1, sharex=True)
axs[0].bar(
mu_grid,
hists[:,middle],
# fill=False,
color = fade('C1', 'C3', num_points=grid_points),
width = mu_width,
alpha = .7,
label = 'histogram',
edgecolor= 'black',
linewidth = 0.05
)
axs[0].legend()
axs[1].bar(
mu_grid,
kde_hists[:,middle],
color = fade('C0', 'C9', num_points=grid_points),
width = mu_width,
alpha = .7,
label = 'kde',
edgecolor= 'black',
linewidth = 0.05
)
axs[1].legend()
plt.xlabel('$\mu$');
# -
# This envelope is much smoother than that of the original histogram, which follows from the smoothness of the (cumulative) density function defined by the kde, and allows us to get gradients!
# Now that we have a histogram we can differentiate, we need to study its properties (and the gradients themselves!)
#
# Two things to study:
# - Quality of approximation to an actual histogram (and to the true distribution)
# - Stability and validity of gradients
#
# To make this more concrete of a comparison, let's introduce a third plot to the above panel that shows the area under the true distribution:
# +
def true_hist(bins, mu):
edge_hi = bins[1:] # ending bin edges ||<-
edge_lo = bins[:-1] # starting bin edges ->||
# get cumulative counts (area under curve) for each set of bin edges
cdf_up = jsc.stats.norm.cdf(edge_hi.reshape(-1, 1), loc=mu)
cdf_dn = jsc.stats.norm.cdf(edge_lo.reshape(-1, 1), loc=mu)
counts = (cdf_up - cdf_dn).T
return counts
truth = true_hist(bins,mu_grid)
# make hists as before (but normalize)
bins = jnp.linspace(lo-1,hi+1,6)
make_kde_hists = jax.vmap(partial(kde_hist, bins = bins, bandwidth = .5, density=True))
kde_hists = make_kde_hists(points)
make_hists = jax.vmap(partial(jnp.histogram, bins = bins, density = True))
hists, _ = make_hists(points)
# +
middle = len(bins)//2 - 1
mu_width = mu_grid[1]-mu_grid[0]
plt.plot(
mu_grid,
truth[:,middle],
color = 'C6',
alpha = .7,
label = 'true',
)
plt.plot(
mu_grid,
hists[:,middle],
# fill=False,
color = 'C1',
alpha = .7,
label = 'histogram',
)
plt.plot(
mu_grid,
kde_hists[:,middle],
color = 'C9',
alpha = .7,
label = 'kde',
)
plt.legend()
plt.xlabel('$\mu$')
plt.suptitle("bandwidth = 0.5, #samples = 100")
# -
# The hyperparameter that will cause the quality of estimation to vary the most will be the *bandwidth* of the kde, which controls the width of the individual point-wise kernels. Moreover, since the kde is a data-driven estimator, the number of samples will also play a role.
# Let's wrap the above plot construction into a function that we can call.
def make_mu_scan(bandwidth, num_samples, grid_points=500, lo=-2, hi=+2):
mu_grid = jnp.linspace(lo, hi, grid_points)
bins = jnp.linspace(lo-3,hi+3,6)
truth = true_hist(bins,mu_grid)
points = jnp.tile(
normal(rng, shape = (num_samples,)),
reps = (grid_points,1)
) + mu_grid.reshape(-1,1)
make_kde_hists = jax.vmap(
partial(kde_hist, bins = bins, bandwidth = bandwidth, density=True)
)
kde_hists = make_kde_hists(points)
make_hists = jax.vmap(partial(jnp.histogram, bins = bins, density = True))
hists, _ = make_hists(points)
study_bin = len(bins)//2 - 1
return jnp.array([truth[:,study_bin], hists[:,study_bin], kde_hists[:,study_bin]])
# +
# bws = jnp.linspace(0.05,0.8,8)
# lo_samp = jax.vmap(partial(make_mu_scan, num_samples = 100))
# hi_samp = jax.vmap(partial(make_mu_scan, num_samples = 100000))
# lo_hists = lo_samp(bws)
# hi_hists = hi_samp(bws)
# lo_hists.shape
# +
# colors = fade('C3','C9',num_points=8)
# +
# fig, axarr = plt.subplots(2,8, sharex=True, sharey=True)
# up, down = axarr
# for i,hists in enumerate(lo_hists):
# [up[i].plot(mu_grid,hists[j],alpha=.6) for j in [0,1]]
# up[i].plot(mu_grid,hists[2],alpha=.7,color=colors[i])
# up[i].set_title(f'bw={bws[i]:.3f}', color=colors[i])
# up[0].set_ylabel('n=1e2', rotation=0, size='large')
# for i,hists in enumerate(hi_hists):
# [down[i].plot(mu_grid,hists[j],alpha=.6) for j in [0,1]]
# down[i].plot(mu_grid,hists[2],alpha=.7,color=colors[i])
# down[0].set_ylabel('n=1e6', rotation=0, size='large')
# fig.tight_layout();
# -
# Good, but we can do better with the function -- not everything needs to be repeated.
# +
def make_points(num_samples, grid_points=300, lo=-2, hi=+2):
mu_grid = jnp.linspace(lo, hi, grid_points)
rngs = [PRNGKey(i) for i in range(9)]
points = jnp.asarray(
[
jnp.tile(
normal(rng, shape = (num_samples,)),
reps = (grid_points,1)
) + mu_grid.reshape(-1,1) for rng in rngs
]
)
return points, mu_grid
def make_kdes(points, bandwidth, bins):
make_kde_hists = jax.vmap(
partial(kde_hist, bins = bins, bandwidth = bandwidth)
)
return make_kde_hists(points)
def make_mu_scan(bandwidth, num_samples, grid_points=500, lo=-2, hi=+2):
points, mu_grid = make_points(num_samples, grid_points, lo, hi)
bins = jnp.linspace(lo-3,hi+3,6)
truth = true_hist(bins,mu_grid)*num_samples
get_kde_hists = jax.vmap(partial(make_kdes, bins=bins, bandwidth=bandwidth))
kde_hists = get_kde_hists(points)
make_hists = jax.vmap(jax.vmap(partial(jnp.histogram, bins = bins)))
hists, _ = make_hists(points)
study_bin = len(bins)//2 - 1
h = jnp.array([truth[:,study_bin],
hists[:,:,study_bin].mean(axis=0),
kde_hists[:,:,study_bin].mean(axis=0)])
stds = jnp.array([hists[:,:,study_bin].std(axis=0),
kde_hists[:,:,study_bin].std(axis=0)])
return h, stds
# +
bws = jnp.array([0.05,0.5,0.8])
lo_samp = jax.vmap(partial(make_mu_scan, num_samples = 20))
mid_samp = jax.vmap(partial(make_mu_scan, num_samples = 100))
hi_samp = jax.vmap(partial(make_mu_scan, num_samples = 5000))
lo_hists, lo_stds = lo_samp(bws)
mid_hists, mid_stds = mid_samp(bws)
hi_hists, hi_stds = hi_samp(bws)
# +
# colors = fade('C0','C9',num_points=7)
# fig, axarr = plt.subplots(3,7, sharex=True, sharey='row')
# up, mid, down = axarr
# for i,res in enumerate(zip(lo_hists, lo_stds)):
# hists, stds = res
# up[i].plot(mu_grid,hists[0],alpha=.4, color='C2',label="actual")
# up[i].plot(mu_grid,hists[1],alpha=.6, linestyle=':', color='C1',label="histogram")
# up[i].errorbar(mu_grid,hists[2],yerr=stds[1],alpha=.03,color=colors[i])
# up[i].plot(mu_grid,hists[2],alpha=.6,color=colors[i],label="kde histogram")
# up[i].set_title(f'bw={bws[i]:.3f}', color=colors[i])
# #up[0].set_ylabel('n=1e2', rotation=0, size='large')
# for i,res in enumerate(zip(mid_hists, mid_stds)):
# hists, stds = res
# mid[i].plot(mu_grid,hists[0],alpha=.4, color='C2',label="true bin height")
# mid[i].plot(mu_grid,hists[1],alpha=.6, linestyle=':', color='C1',label="histogram")
# mid[i].errorbar(mu_grid,hists[2],yerr=stds[1],alpha=.03,color=colors[i])
# mid[i].plot(mu_grid,hists[2],alpha=.6,color=colors[i],label="kde histogram")
# for i,res in enumerate(zip(hi_hists, hi_stds)):
# hists, stds = res
# down[i].plot(mu_grid,hists[0],alpha=.4, color='C2',label="actual")
# down[i].plot(mu_grid,hists[1],alpha=.6, linestyle=':', color='C1',label="histogram")
# down[i].errorbar(mu_grid,hists[2],yerr=stds[1],alpha=.03,color=colors[i])
# down[i].plot(mu_grid,hists[2],alpha=.6,color=colors[i],label="kde histogram")
# #down[0].set_ylabel('n=1e6', rotation=0, size='large')
# down[3].set_xlabel("$\mu$",size='large')
# mid[0].set_ylabel("frequency",size='large',labelpad=11)
# mid[-1].legend(bbox_to_anchor=(1.1, 1.05), frameon=False)
# fig.tight_layout();
# plt.savefig('samples_vs_bw.png', bbox_inches='tight')
# +
# import matplotlib as mpl
# colors = fade('C0','C9',num_points=7)
# fig, ax = plt.subplots()
# cmap = mpl.colors.ListedColormap(colors)
# norm = mpl.colors.Normalize(vmin=bws[0], vmax=bws[-1])
# mpl.colorbar.ColorbarBase(ax, cmap=cmap, norm=norm)
# +
colors = fade('C0','C9',num_points=7)
fig, axarr = plt.subplots(3,len(bws), sharex=True, sharey='row')
up, mid, down = axarr
for i,res in enumerate(zip(lo_hists, lo_stds)):
hists, stds = res
up[i].plot(mu_grid,hists[0],alpha=.4, color='C3',label="actual", linestyle=':')
up[i].fill_between(mu_grid, hists[1]+stds[0], hists[1]-stds[0], alpha=.2,color='C1',label='histogram variance')
up[i].plot(mu_grid,hists[1],alpha=.4, color='C1',label="histogram")
up[i].fill_between(mu_grid, hists[2]+stds[1], hists[2]-stds[1], alpha=.2,color='C0')
up[i].plot(mu_grid,hists[2],alpha=.6,color='C0',label="kde histogram")
up[i].set_title(f'bw={bws[i]:.2f}', color='C0')
#up[0].set_ylabel('n=1e2', rotation=0, size='large')
for i,res in enumerate(zip(mid_hists, mid_stds)):
hists, stds = res
mid[i].plot(mu_grid,hists[0],alpha=.4, color='C3',label="true bin height", linestyle=':')
mid[i].fill_between(mu_grid, hists[1]+stds[0], hists[1]-stds[0], alpha=.2,color='C1',label='histogram $\pm$ std')
mid[i].plot(mu_grid,hists[1],alpha=.4,color='C1',label="histogram")
mid[i].fill_between(mu_grid, hists[2]+stds[1], hists[2]-stds[1], alpha=.2,color='C0',label='kde histogram $\pm$ std')
mid[i].plot(mu_grid,hists[2],alpha=.6,color='C0',label="kde histogram")
for i,res in enumerate(zip(hi_hists, hi_stds)):
hists, stds = res
down[i].plot(mu_grid,hists[0],alpha=.4, color='C3',label="actual", linestyle=':')
down[i].fill_between(mu_grid, hists[1]+stds[0], hists[1]-stds[0], alpha=.2,color='C1')
down[i].plot(mu_grid,hists[1],alpha=.4, color='C1',label="histogram")
down[i].fill_between(mu_grid, hists[2]+stds[1], hists[2]-stds[1], alpha=.2,color='C0')
down[i].plot(mu_grid,hists[2],alpha=.6,color='C0',label="kde histogram")
#down[0].set_ylabel('n=1e6', rotation=0, size='large')
down[1].set_xlabel("$\mu$",size='large')
mid[0].set_ylabel("frequency",size='large',labelpad=11)
mid[-1].legend(bbox_to_anchor=(1.1, 1.05), frameon=False)
fig.tight_layout();
plt.savefig('samples_vs_bw_nofancy.png', bbox_inches='tight')
# -
# Cool! Now, let's think about gradients.
#
# Since we know analytically that the height of a bin defined by $(a,b)$ for a given $\mu$ value is just
#
# $$bin_{\mathsf{true}}(\mu) = \mathsf{normcdf}(b;\mu) - \mathsf{normcdf}(a;\mu) $$
#
# we can then just diff this wrt $\mu$ by hand!
#
# $$\mathsf{normcdf}(x;\mu) = \frac{1}{2}\left[1+\operatorname{erf}\left(\frac{x-\mu}{\sigma \sqrt{2}}\right)\right]$$
#
# $$\Rightarrow \frac{\partial}{\partial\mu}\mathsf{normcdf}(x;\mu) = \frac{1}{2}\left[1-\left(\frac{2}{\sqrt{2\pi}\sigma} e^{-\frac{(x-\mu)^2}{2\sigma^2}}\right)\right]$$
#
# since $\frac{d}{d z} \operatorname{erf}(z)=\frac{2}{\sqrt{\pi}} e^{-z^{2}}$.
#
# We have $\sigma=1$, making this simpler:
#
# $$\Rightarrow \frac{\partial}{\partial\mu}\mathsf{normcdf}(x;\mu) = \frac{1}{2}\left[1-\left(\frac{2}{\sqrt{2\pi}} e^{-\frac{(x-\mu)^2}{2}}\right)\right]$$
#
# All together:
#
# $$\Rightarrow \frac{\partial}{\partial\mu}bin_{\mathsf{true}}(\mu) = -\frac{1}{\sqrt{2\pi}}\left[\left(e^{-\frac{(b-\mu)^2}{2}}\right) - \left( e^{-\frac{(a-\mu)^2}{2}}\right)\right]$$
#
# The histogram's gradient will be ill-defined, but we can get an estimate of this through finite differences:
#
# $$\mathsf{grad}_{\mathsf{hist}}(bin)(\mu_i) \approx \frac{bin(\mu_{i+1})-bin(\mu_i)}{\mu_{i+1}-\mu_{i}}$$
#
# for a kde, we can just use autodiff.
def true_grad(mu,bins):
b = bins[1:] # ending bin edges ||<-
a = bins[:-1] # starting bin edges ->||
return -(1/((2*jnp.pi)**0.5))*(jnp.exp(-((b-mu)**2)/2) - jnp.exp(-((a-mu)**2)/2))
# +
bins = jnp.linspace(-5,5,6)
mus = jnp.linspace(-2,2,300)
true_grad_many = jax.vmap(partial(true_grad, bins = bins))
grads = true_grad_many(mus)
plt.plot(mus, grads[:,2]);
# -
# Shape looks good!
# +
def gen_points(mu, jrng, nsamples):
points = normal(jrng, shape = (nsamples,))+mu
return points
def bin_height(mu, jrng, bw, nsamples, bins):
points = gen_points(mu, jrng, nsamples)
return kde_hist(points, bins, bandwidth=bw)[2]
def kde_grads(bw, nsamples, lo=-2, hi=+2, grid_size=300):
bins = jnp.linspace(lo-3,hi+3,6)
mu_grid = jnp.linspace(lo,hi,grid_size)
rngs = [PRNGKey(i) for i in range(9)]
grad_fun = jax.grad(bin_height)
grads = []
for i,jrng in enumerate(rngs):
get_grads = jax.vmap(partial(
grad_fun, jrng=jrng, bw=bw, nsamples=nsamples, bins=bins
))
grads.append(get_grads(mu_grid))
return jnp.asarray(grads)
x = kde_grads(0.2,1000).mean(axis=0)
mus = jnp.linspace(-2,2,300)
plt.plot(mus,x)
bins = jnp.linspace(-5,5,6)
true_grad_many = jax.vmap(partial(true_grad, bins = bins))
grads = true_grad_many(mus)*1000
plt.plot(mus, grads[:,2]);
# -
# Okay, looks like the kde grads work as anticipated -- we just need to look at the hist grads now.
# +
def get_hist(mu, jrng, nsamples, bins):
points = gen_points(mu, jrng, nsamples)
hist, _ = jnp.histogram(points, bins)
return hist[2]
def hist_grad_numerical(bin_heights, mu_width):
# in mu plane
lo = bin_heights[:-1]
hi = bin_heights[1:]
bin_width = (bins[1]-bins[0])
grad_left = -(lo-hi)/mu_width
# grad_right = -grad_left
return grad_left
def hist_grads(nsamples, lo=-2, hi=+2, grid_size=300):
bins = jnp.linspace(lo-3,hi+3,6)
mu_grid = jnp.linspace(lo,hi,grid_size)
rngs = [PRNGKey(i) for i in range(9)]
grad_fn = partial(hist_grad_numerical, mu_width=mu_grid[1]-mu_grid[0])
grads = []
for jrng in rngs:
get_heights = jax.vmap(partial(
get_hist, jrng=jrng, nsamples=nsamples, bins=bins
))
grads.append(grad_fn(get_heights(mu_grid)))
return jnp.asarray(grads)
# -
hist_grads(1000).shape
# +
x = kde_grads(0.2,1000).mean(axis=0)
mus = jnp.linspace(-2,2,300)
plt.plot(mus,x, label= 'kde')
bins = jnp.linspace(-5,5,6)
plt.plot(mus[:-1],hist_grads(1000).mean(axis=0), label = 'hist')
true_grad_many = jax.vmap(partial(true_grad, bins = bins))
grads = true_grad_many(mus)*1000
plt.plot(mus, grads[:,2], label='true')
plt.legend();
# -
# Cool! Everything is scaling properly to the number of samples, and we can see the jaggedness of the histogram gradients.
#
# Now let's combine these functions into one, and run that over the same bandwidth and sample numbers as before!~
# +
def both_grads(bw, nsamples, lo=-2, hi=+2, grid_size=300):
bins = jnp.linspace(lo-3,hi+3,6)
mu_grid = jnp.linspace(lo,hi,grid_size)
hist_grad_fun = partial(hist_grad_numerical, mu_width=mu_grid[1]-mu_grid[0])
grad_fun = jax.grad(bin_height)
hist_grads = []
kde_grads = []
rngs = [PRNGKey(i) for i in range(9)]
for jrng in rngs:
get_heights = jax.vmap(partial(
get_hist, jrng=jrng, nsamples=nsamples, bins=bins
))
hist_grads.append(hist_grad_fun(get_heights(mu_grid)))
get_grads = jax.vmap(partial(
grad_fun, jrng=jrng, bw=bw, nsamples=nsamples, bins=bins
))
kde_grads.append(get_grads(mu_grid))
hs = jnp.array(hist_grads)
ks = jnp.array(kde_grads)
h = jnp.array([hs.mean(axis=0),hs.std(axis=0)])
k = jnp.array([ks.mean(axis=0),ks.std(axis=0)])
return h,k
# +
bws = jnp.array([0.05,0.5,0.8])
samps = [20,100,5000]
grid_size = 600
lo_samp = jax.vmap(partial(both_grads, nsamples = samps[0],grid_size=grid_size))
mid_samp = jax.vmap(partial(both_grads, nsamples = samps[1],grid_size=grid_size))
hi_samp = jax.vmap(partial(both_grads, nsamples = samps[2],grid_size=grid_size))
lo_hist, lo_kde = lo_samp(bws)
mid_hist, mid_kde = mid_samp(bws)
hi_hist, hi_kde = hi_samp(bws)
# +
# colors = fade('C0','C9',num_points=7)
mu_grid = jnp.linspace(-2,2,grid_size)
true = [true_grad_many(mu_grid)[:,2]*s for s in samps]
fig, axarr = plt.subplots(3,len(bws), sharex=True, sharey='row')
up, mid, down = axarr
for i,res in enumerate(zip(lo_hist, lo_kde)):
hist_grads, hist_stds = res[0]
kde_grads, kde_stds = res[1]
up[i].plot(mu_grid,true[0],alpha=.4, color='C3',label="actual", linestyle=':')
y = jnp.array(up[i].get_ylim())
# up[i].fill_between(mu_grid[:-1], hist_grads+hist_stds, hist_grads-hist_stds, alpha=.1,color='C1',label='histogram $\pm$ std')
up[i].plot(mu_grid[:-1], hist_grads,alpha=.3, color='C1',label="histogram",linewidth=0.5)
up[i].fill_between(mu_grid, kde_grads+kde_stds, kde_grads-kde_stds, alpha=.2,color='C0',label='kde histogram $\pm$ std')
up[i].plot(mu_grid,kde_grads,alpha=.6,color='C0',label="kde histogram")
up[i].set_title(f'bw={bws[i]:.2f}', color='C0')
up[i].set_ylim(y*1.3)
#up[0].set_ylabel('n=1e2', rotation=0, size='large')
for i,res in enumerate(zip(mid_hist, mid_kde)):
hist_grads, hist_stds = res[0]
kde_grads, kde_stds = res[1]
mid[i].plot(mu_grid,true[1],alpha=.4, color='C3',label="actual", linestyle=':')
y = jnp.array(mid[i].get_ylim())
# mid[i].fill_between(mu_grid[:-1], hist_grads+hist_stds, hist_grads-hist_stds, alpha=.1,color='C1',label='histogram $\pm$ std')
mid[i].plot(mu_grid[:-1], hist_grads,alpha=.3, color='C1',label="histogram",linewidth=0.5)
mid[i].fill_between(mu_grid, kde_grads+kde_stds, kde_grads-kde_stds, alpha=.2,color='C0',label='kde histogram $\pm$ std')
mid[i].plot(mu_grid,kde_grads,alpha=.6,color='C0',label="kde histogram")
mid[i].set_ylim(y*1.3)
for i,res in enumerate(zip(hi_hist, hi_kde)):
hist_grads, hist_stds = res[0]
kde_grads, kde_stds = res[1]
down[i].plot(mu_grid,true[2],alpha=.4, color='C3',label="actual", linestyle=':')
y = jnp.array(down[i].get_ylim())
# down[i].fill_between(mu_grid[:-1], hist_grads+hist_stds, hist_grads-hist_stds, alpha=.1,color='C1',label='histogram variance')
down[i].plot(mu_grid[:-1], hist_grads,alpha=.2, color='C1',label="histogram")
down[i].fill_between(mu_grid, kde_grads+kde_stds, kde_grads-kde_stds, alpha=.2,color='C0')
down[i].plot(mu_grid,kde_grads,alpha=.6,color='C0',label="kde histogram")
down[i].set_ylim(y*1.3)
#down[0].set_ylabel('n=1e6', rotation=0, size='large')
down[1].set_xlabel("$\mu$",size='large')
mid[0].set_ylabel("$\partial\,$frequency / $\partial\mu$",size='large',labelpad=11)
mid[-1].legend(bbox_to_anchor=(1.1, 1.05), frameon=False)
fig.tight_layout();
plt.savefig('samples_vs_bw_nofancy_gradients.png', bbox_inches='tight')
# -
# Suuuuuuper! Let's now look at metrics of quality.
# +
def gen_points(mu, jrng, nsamples):
points = normal(jrng, shape = (nsamples,))+mu
return points
def bin_height(mu, jrng, bw, nsamples, bins):
points = gen_points(mu, jrng, nsamples)
return kde_hist(points, bins, bandwidth=bw)[2]
def kde_grads_mse(bw, nsamples, lo=-2, hi=+2, grid_size=500):
bins = jnp.linspace(lo-3,hi+3,6)
mu_grid = jnp.linspace(lo,hi,grid_size)
rngs = [PRNGKey(i) for i in range(9)]
grad_fun = jax.grad(bin_height)
grads = []
for i,jrng in enumerate(rngs):
get_grads = jax.vmap(partial(
grad_fun, jrng=jrng, bw=bw, nsamples=nsamples, bins=bins
))
grads.append(get_grads(mu_grid))
true_grad_many = jax.vmap(partial(true_grad, bins = bins))
true = true_grad_many(mu_grid)[:,2]*nsamples
mse = jnp.abs((true - jnp.asarray(grads))/true)
return mse.mean(axis=1).mean(axis=0)
bws = jnp.linspace(0.05,1,20)
samps = jnp.linspace(5,5000,20).astype('int')
funcs = [jax.vmap(partial(kde_grads_mse, nsamples=n)) for n in samps]
mses = jnp.array([f(bws) for f in funcs])
# +
X, Y = jnp.meshgrid(bws,samps)
p = plt.contourf(X,Y,mses,levels=50)
c = plt.colorbar(p)
c.set_label('gradient mean relative error',rotation=270, labelpad=15)
mindex = jnp.argmin(mses.ravel())
plt.scatter(X.ravel()[mindex], Y.ravel()[mindex], label = 'minimum error', color='C1')
plt.xlabel('bandwidth')
plt.ylabel('#samples')
plt.legend()
# -
mses[0].shape
# +
# jnp.save('relative_error_bw0.05_1_20_n5_5000_20.npy', mses)
# +
rs = mses[1:-4]
X, Y = jnp.meshgrid(bws,samps)
p = plt.contourf(X[1:-4],Y[1:-4],rs,levels=10)
c = plt.colorbar(p)
c.set_label('gradient mean relative error',rotation=270, labelpad=15)
mindex = jnp.argmin(mses.ravel())
# plt.scatter(X.ravel()[mindex], Y.ravel()[mindex], label = 'minimum error', color='C1')
plt.xlabel('bandwidth')
plt.ylabel('#samples')
plt.legend()
# -
| _notebooks/kde_histograms.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="view-in-github" colab_type="text"
# <a href="https://colab.research.google.com/github/mr7495/image-classification-spatial/blob/main/Sub_ImageNet_ResNet50_Depthw_constraints_224.ipynb" target="_parent"><img src="https://colab.research.google.com/assets/colab-badge.svg" alt="Open In Colab"/></a>
# + id="WMJOLaJwx7Tg"
#This code shows the implementation of our proposed depthwise convolution layer method with constraints for enhancing image classification by extracting spatial data
#Images resolution is 224*224
# + id="t1SzEKNRzFL5" colab={"base_uri": "https://localhost:8080/"} outputId="e116e023-682b-487d-b1e2-71fbdb0a1305"
# !nvidia-smi #show GPU type
# + id="6mw_i4iuzHlC"
import numpy as np
from tensorflow import keras
from tensorflow.keras import layers
import cv2
import zipfile
import shutil
import random
import pandas as pd
import csv
import os
# + id="uP8qFi9uWzAG" colab={"base_uri": "https://localhost:8080/"} outputId="2f58bd9c-14ad-48c6-dce0-e3f3102814b2"
#download Sub-ImageNet dataset from Kaggle (https://www.kaggle.com/mohammadrahimzadeh/imagenet-70classes)
#Get a new download like from kaggle website at the mentioned URL, then replace the link with the "kaggle_linke" input in the next line of code
# !wget -cO - kaggle_link > imagenet_70classes.zip #replace kaggle_link with a new download link from https://www.kaggle.com/mohammadrahimzadeh/imagenet-70classes
# + id="pWNFXlWBWzDg"
archive = zipfile.ZipFile('imagenet_70classes.zip') #Extract Sub-ImageNet Dataset
for file in archive.namelist():
archive.extract(file, 'data')
# + id="79KB2ItAzHqb"
#Set data augmentation techniques
train_datagen = keras.preprocessing.image.ImageDataGenerator(horizontal_flip=True,vertical_flip=True
,zoom_range=0.2,rotation_range=360
,width_shift_range=0.1,height_shift_range=0.1
,channel_shift_range=50
,brightness_range=(0,1.2)
,preprocessing_function=keras.applications.imagenet_utils.preprocess_input)
test_datagen = keras.preprocessing.image.ImageDataGenerator(preprocessing_function=keras.applications.imagenet_utils.preprocess_input)
train_df = pd.read_csv("data/train.csv")
test_df = pd.read_csv("data/test.csv")
# + id="sq8125qva0PJ"
#Replace '\\' with '/' in CSV files
for i in range(len(train_df['filename'])):
name=train_df['filename'][i]
index=name.index('\\')
new_name=name[:index]+'/'+name[index+1:]
train_df['filename'][i]=new_name
for i in range(len(test_df['filename'])):
name=test_df['filename'][i]
index=name.index('\\')
new_name=name[:index]+'/'+name[index+1:]
test_df['filename'][i]=new_name
# + id="L3dqLamH0TBw" colab={"base_uri": "https://localhost:8080/"} outputId="b50fde4d-4223-4f29-cca6-364d992b3ae7"
#Create Data augmentation techniques
batch_size=70
train_generator = train_datagen.flow_from_dataframe(
dataframe=train_df,
directory='data',
x_col="filename",
y_col="class",
target_size=(224, 224),
batch_size=batch_size,
class_mode='categorical',shuffle=True)
validation_generator = test_datagen.flow_from_dataframe(
dataframe=test_df,
directory='data',
x_col="filename",
y_col="class",
target_size=(224, 224),
batch_size=batch_size,
class_mode='categorical',shuffle=True)
# + id="yo9nEsSx7OfK"
name="Sub-ImageNet-ResNet50-Depthw-constraints-224"
# !mkdir "models" #create new folder for saving checkpoints
# !mkdir "reports" #create new folder for saving evaluation reports
keras.backend.clear_session() #clear backend
shape=(224,224,3)
input_tensor=keras.Input(shape=shape)
base_model=keras.applications.ResNet50(input_tensor=input_tensor,weights=None,include_top=False)
depth=keras.layers.DepthwiseConv2D(tuple(base_model.output.shape[1:3]), depthwise_initializer=keras.initializers.RandomNormal(mean=0.0,stddev=0.01),
bias_initializer=keras.initializers.Zeros(),depthwise_constraint=keras.constraints.NonNeg())(base_model.output)
flat=keras.layers.Flatten()(depth)
preds=keras.layers.Dense(70,activation='softmax',
kernel_initializer=keras.initializers.RandomNormal(mean=0.0,stddev=0.01),
bias_initializer=keras.initializers.Zeros(),)(flat)
model=keras.Model(inputs=base_model.input, outputs=preds)
##################################
for layer in model.layers:
layer.trainable = True
model.summary()
filepath="models/%s-{epoch:02d}-{val_accuracy:.4f}.hdf5"%name
checkpoint = keras.callbacks.ModelCheckpoint(filepath, monitor='val_accuracy', save_best_only=False, mode='max') #creating checkpoint to save the best validation accuracy
callbacks_list = [checkpoint]
#Determine adaptive learning rate with an initialization value of 0.045 and decay of 0.94 every two epochs. 31500 is the number of training images
lr_schedule =keras.optimizers.schedules.ExponentialDecay(
initial_learning_rate=0.045,
decay_steps=2*int(31500/batch_size),
decay_rate=0.94,
staircase=True)
optimizer=keras.optimizers.SGD(momentum=0.9,learning_rate=lr_schedule)
model.compile(optimizer=optimizer, loss='categorical_crossentropy',metrics=['accuracy'])
hist=model.fit_generator(train_generator, epochs=148,validation_data=validation_generator,shuffle=True,callbacks=callbacks_list) #start training
with open('reports/{}.csv'.format(name), mode='w',newline='') as csv_file: #write reports
csv_writer = csv.writer(csv_file, delimiter=',', quotechar='"', quoting=csv.QUOTE_MINIMAL)
for key in hist.history:
data=[key]
data.extend(hist.history[key])
csv_writer.writerow(data)
print("Training finished. Reports saved!")
| Sub_ImageNet_ResNet50_Depthw_constraints_224.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import pandas as pan
import numpy as np
import seaborn as sns
import matplotlib.pyplot as plt
import re
import string
tweets =pan.read_csv("covid19_tweets.csv")
tweets.columns = tweets.columns.str.strip().str.lower().str.replace(' ', '_').str.replace('(', '').str.replace(')', '')
tweets.head()
# -
tweets.replace('',np.NaN)
tweets.dropna(inplace=True)
tweets['user_name'].value_counts()
from collections import defaultdict
word_dict = {}
grouped_df = tweets.groupby('user_description').sum().reset_index()
grouped_df['Most used words'] = grouped_df['user_description'].apply(lambda x : word_dict[x])
grouped_df[['user_description', 'Most used words']]
tweets=pan.concat([tweets.user_name,tweets.user_description],axis=1)
tweets.head()
description=tweets['user_description'].replace('([^0-9A-Za-z \t])|(\w+:\/\/\S+)', '', regex=True)
tweets['user_description']=description
tweets.reset_index(inplace=True,drop=True)
tweets.head()
def remove_pattern(user_description, pattern):
r = re.findall(pattern, text)
for i in r:
text = re.sub(i, "", text)
return text
tweets.info()
tweets.head()
tweets['char_count'] = tweets['user_description'].str.len() ## this also includes spaces
tweets[['user_description','char_count']].head()
# +
freq = pan.Series(' '.join(tweets['user_description']).split()).value_counts()[:10]
freq = list(freq.index)
tweets['user_description'] = tweets['user_description'].apply(lambda x: " ".join(x for x in x.split() if x not in freq))
tweets['user_description'].head()
freq = pan.Series(' '.join(tweets['user_description']).split()).value_counts()[-10:]
freq
# -
from nltk.corpus import stopwords
stop = stopwords.words('english')
tweets['user_description'] = tweets['user_description'].apply(lambda x: " ".join(x for x in x.split() if x not in stop))
tweets['user_description'].head()
# +
sub_tweets = tweets[tweets['user_name']=='Time4fisticuffs']
# sample cleaned text and tokens tagged as nouns
sub_tweets['user_description'].sample(2)
# -
def my_tokenizer(text):
return text.split() if text != None else []
tokens = sub_tweets.user_description.map(my_tokenizer).sum()
print(tokens[:100])
# +
from collections import Counter
counter = Counter(tokens)
counter.most_common(20)
# -
tweets.user_description.unique()
# +
freq_df = pan.DataFrame.from_records(counter.most_common(20),
columns=['token', 'count'])
# create bar plot
freq_df.plot(kind='bar', x='token');
# -
tweets['user_descriotion'] = tweets.selftext_lemma\
.map(lambda l: 0 if l==None else len(l.split()))
tweets['user_description'].value_counts()
top_users=tweets.groupby('user_name')['user_description'].count().reset_index()
top_users.columns=['user_name','count']
top_users.sort_values('count',ascending=False,inplace=True)
top_users[0:40].plot(kind='bar',x='user_name',y='count')
plt.xlabel('Users')
plt.ylabel('Tweets')
plt.title('Top 10 tweeters')
plt.show()
tweets.shape
tweets.isnull().sum()
def missing_data(data):
total_count = tweets.isna().count()
total_nulls = tweets.isnull().sum()
percent_nulls = (tweets.isnull().sum()/tweets.isnull().count()*100)
tb = pan.concat([ total_count, total_nulls, percent_nulls], axis=1, keys=[' total_count','Total nulls', 'null Percent'])
types = []
uni_vals = []
for col in tweets:
dtype = str(tweets['user_description'].dtype)
uniques = tweets['user_description'].nunique()
types.append(dtype)
uni_vals.append(uniques)
tb['Types'] = types
tb['Unique values'] = uni_vals
return tb
missing_data(tweets)
politics = tweets['user_description'] == 'wednesday'
politics.head()
# +
def makeList(x):
x = str(x)
x = x.replace('[', '')
x = x.replace(']', '')
x = x.split(',')
return x
tweets_tags = tweets.copy()
tweets_tags['hashes'] = tweets_tags['user_description'].apply(lambda x:makeList(x))
tweets_tags = tweets_tags.explode('hashes')
tweets_tags['user_name'] = tweets_tags['user_name'].str.lower()
tweets_tags['user_name'] = tweets_tags['user_name'].str.replace(" "," ")
tweets_tags['user_name'] = tweets_tags['user_description'].str.replace("'"," ")
tags = tweets_tags['user_name'].value_counts().reset_index()
tags = tags[0:10]
tags.rename(columns = {'index':'Words', 'hashes':'User'}, inplace = True)
tags
# -
fig,ax = plt.subplots(1,1, figsize=(10,10))
sns.set(style="whitegrid")
sns.barplot(x=tags.Words,y=tags.user_name,palette='OrRd_r')
plt.title("Frecuencia de palabras en descripcion creibles",fontsize=15)
plt.xticks(rotation=45,fontsize=12)
def get_countplot(tweets):
"""this function handles the top 10 user specifications"""
user_cols = ['user_name', 'user_description']
for col in user_cols:
tweets[col].value_counts().head(20).plot(kind = 'bar', figsize = (15,5))
plt.show()
get_countplot(tweets)
def clean_text(text):
'''Make text lowercase, remove text in square brackets,remove links,remove punctuation
and remove words containing numbers.'''
text = text.lower()
text = re.sub('\[.*?\]', '', text)
text = re.sub('https?://\S+|www\.\S+', '', text)
text = re.sub('<.*?>+', '', text)
text = re.sub('[%s]' % re.escape(string.punctuation), '', text)
text = re.sub('\n', '', text)
text = re.sub('\w*\d\w*', '', text)
return text
def text_preprocessing(text):
"""
Cleaning and parsing the text.
"""
tokenizer = nltk.tokenize.RegexpTokenizer(r'\w+')
nopunc = clean_text(text)
tokenized_text = tokenizer.tokenize(nopunc)
combined_text = ' '.join(tokenized_text)
return combined_text
tweets['text_clean'] = tweets['user_description'].apply(str).apply(lambda x: text_preprocessing(x))
from sklearn import model_selection
from sklearn.feature_extraction.text import CountVectorizer,TfidfVectorizer
def get_top_n_words(corpus, n=None):
"""
List the top n words in a vocabulary according to occurrence in a text corpus.
"""
vec = CountVectorizer(stop_words = 'english').fit(corpus)
bag_of_words = vec.transform(corpus)
sum_words = bag_of_words.sum(axis=0)
words_freq = [(word, sum_words[0, idx]) for word, idx in vec.vocabulary_.items()]
words_freq =sorted(words_freq, key = lambda x: x[1], reverse=True)
return words_freq[:n]
# +
unigrams = get_top_n_words(tweets['text_clean'],10)
unigram_df = pd.DataFrame(unigrams, columns = ['Text' , 'count']) #Creating df
#Plotting
plt.figure(figsize=(9,10))
sns.barplot(unigram_df["count"],unigram_df["Text"])
plt.title("Top 10 de palabras en descripción")
plt.xlabel("Counts")
plt.ylabel("words")
plt.show()
# -
| Untitled.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
"""
Created on Mon February 22nd, 2022
@author: <NAME>
Description:
This script is designed to read NIfTI files that contain Grade 4 (HGG) tumors from the
local directory, extract the areas where the tumor is present across (x,y,z) by finding
the (min, max) of each axis, then normalize the intensity of the extracted 2D images and
save the slices on a local directory as .png files.
Note: This script was created on a Linux computer and some commands may not work on Windows/MAC OS.
"""
import os.path
import numpy as np
import glob
import nibabel as nib
import pandas as pd
import matplotlib.pyplot as plt
import re
from PIL import Image
from numpy import ndarray
# Folder where the created images will be saved in
out_path = r'/local/data1/elech646/Tumor_grade_classification/Slices'
# Create subfolders
if not os.path.exists(out_path + "/sagittal_grade_classification"):
os.mkdir(out_path + "/sagittal_grade_classification")
if not os.path.exists(out_path + "/frontal_grade_classification"):
os.mkdir(out_path + "/frontal_grade_classification")
if not os.path.exists(out_path + "/trans_grade_classification"):
os.mkdir(out_path + "/trans_grade_classification")
# Add HGG path
sag_path = out_path + "/sagittal_grade_classification" + "/HGG"
fro_path = out_path + "/frontal_grade_classification" + "/HGG"
tra_path = out_path + "/trans_grade_classification" + "/HGG"
if not os.path.exists(sag_path):
os.mkdir(sag_path)
if not os.path.exists(fro_path):
os.mkdir(fro_path)
if not os.path.exists(tra_path):
os.mkdir(tra_path)
filepath = []
# Scanning files + directories
for roots, dirs, files in os.walk("/local/data1/elech646/Tumor_grade_classification/HGG"):
for name in files:
if name.endswith((".nii.gz",".nii")):
filepath.append(roots + os.path.sep + name)
# Creating the images
for i, file in enumerate(filepath):
img = nib.load(file)
img_data = img.get_fdata() # Getting data matrix
patient = file.split("HGG/")[1].split("/")[0] + "/"
if not os.path.exists(sag_path + "/" + patient):
os.mkdir(sag_path + "/" + patient)
if not os.path.exists(fro_path + "/" + patient):
os.mkdir(fro_path + "/" + patient)
if not os.path.exists(tra_path + "/" + patient):
os.mkdir(tra_path + "/" + patient)
# Maximum and minimum slices in which the tumor is present
tr_0 = min(ndarray.nonzero(img_data)[0]) # xmin
tr_1 = max(ndarray.nonzero(img_data)[0]) # xmax
fr_0 = min(ndarray.nonzero(img_data)[1]) # ymin
fr_1 = max(ndarray.nonzero(img_data)[1]) # ymax
sag_0 = min(ndarray.nonzero(img_data)[2]) # zmin
sag_1 = max(ndarray.nonzero(img_data)[2]) # zmax
# Creating the images in the Sagittal Plane (yz)
img_sag = np.rot90(img_data, axes = (1, 2)) # yz plane sagittal
img_sag = np.flip(img_sag, 0) # flip the image left/right since mango was used
for sag in range(sag_0, sag_1 + 1):
perc = int(((sag - sag_0)/(sag_1 - sag_0))*100) # Percentage along the selected slices
tmp = img_sag[sag,:,:]
min_v = img_sag.min()
max_v = img_sag.max()
# Normalize image
tmp_norm = (255*(tmp - min_v) / max_v).astype(np.uint8)
# Add modality
modality = filepath[i].split("_",1)[1].split(".nii", 1)[0].split("_")[-1]
# Name the files
title = sag_path + "/" + patient + patient[:-1] + '_' + modality + str(perc)
# Convert to RGB
im = Image.fromarray(tmp_norm).convert('RGB')
# Create output path
out_path_f = out_path + "/"
# Save images
im.save(os.path.join(out_path_f, title + ".png"))
# Creating the images in the Frontal/Coronal Plane (xz)
img_fr = np.rot90(img_data, axes = (0,2)) # xz plane frontal
for front in range(fr_0, fr_1 + 1):
perc = int(((front - fr_0) /(fr_1 - fr_0))*100) # Percentage along the selected slices
tmp = img_fr[:,front,:]
min_v = img_fr.min()
max_v = img_fr.max()
# Normalize image
tmp_norm = (255*(tmp - min_v) / max_v).astype(np.uint8)
# Add modality
modality = filepath[i].split("_",1)[1].split(".nii", 1)[0].split("_")[-1]
# Name the files
title = fro_path + "/" + patient + patient[:-1] + '_' + modality + str(perc)
# Convert to RGB
im = Image.fromarray(tmp_norm).convert('RGB')
# Create output file path
out_path_f = out_path + "/"
# Save images
im.save(os.path.join(out_path_f, title + ".png"))
# Creating the images in the Transversal/Axial Plane (xy)
img_tr = np.rot90(img_data, 3, axes = (0,1)) # xy plane transversal
x,y,z = img_tr.shape
# Inverting slices upside/down since mango was used
tr_0 = z-max(ndarray.nonzero(img_tr)[0])
tr_1 = z-min(ndarray.nonzero(img_tr)[0])
for transv in range(tr_0, tr_1):
perc = int(((transv - tr_0) / (tr_1 - tr_0))*100) # Percentage along the selected slices
tmp = img_tr[:,:,transv]
min_v = img_tr.min()
max_v = img_tr.max()
# Normalize image
tmp_norm = (255*(tmp - min_v) / max_v).astype(np.uint8)
# Add modality
modality = filepath[i].split("_",1)[1].split(".nii", 1)[0].split("_")[-1]
# Name the files
title = tra_path + "/" + patient + patient[:-1] + '_' + modality + str(perc)
# Convert to RGB
im = Image.fromarray(tmp_norm).convert('RGB')
# Create output file path
out_path_f = out_path + "/"
# Save images
im.save(os.path.join(out_path_f, title + ".png"))
| source/extract_HGG_slices.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import random
import math
# +
MAX_COMPARTMENTS=1000
MAX_PARTICLES=1000
CycleMultiplication=1000
def fc(x):
return math.factorial(x)
def lg(x):
return np.log(x)
# +
N = int(input("Number of Particles N? "))
P = int(input("Number of Compartments p? "))
Ncycle = int(input("Number of Cycles NCycles? "))
if N < 2 or P < 2 or P > MAX_COMPARTMENTS or N > MAX_PARTICLES:
print("Error in input parameters\n")
exit()
NumInComp = np.zeros((Ncycle, P))
Analytical = [0]*P
for l in range(Ncycle):
for i in range(N):
j = random.randint(0, P-1)
NumInComp[l][j] += 1
NumInComp = [x / (N*Ncycle) for x in sum(NumInComp)]
for i in range(1,P):
Analytical[i-1] = np.exp(lg(fc(N)) - lg(fc(i)) - lg(fc(N-i)) - j*lg(P) - (N-i)*lg(P/(P-1)))
# -
#Plotting
fig = plt.figure(figsize=(15,15))
plt.plot(NumInComp,range(1,N+1), label='computation')
plt.plot(trials, re_comput , color='y', linestyle='-', label="theory")
plt.title("Relative error calculated with different n",fontsize=25)
plt.xlabel('Relative error estimated',fontsize=20)
plt.ylabel('Relative error theoritical',fontsize=20)
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('RE_different_n', dpi=300)
plt.plot(range(1,P+1), NumInComp, label='computation')
plt.plot(range(1,P+1), Analytical, label='computation')
# # Boltzmann Distribution
# +
NumberOfEnergyLevels = int(input("NumberOfEnergyLevels ?"))
Temperature = [0.1, 1, 10, 100, 1000, 10000, 100000]
Beta = [1/x for x in Temperature]
tmp = []
fig = plt.figure(figsize=(10,10))
for b in Beta[:]:
for i in range(NumberOfEnergyLevels):
tmp.append(np.exp(-b*i))
Distribution = [x/sum(tmp) for x in tmp]
plt.plot(range(NumberOfEnergyLevels), Distribution, label=f"T={round(1/b)}")
Distribution = []
tmp = []
plt.title("Boltzmann Distribution",fontsize=25)
plt.xlabel('Energy Levels',fontsize=20)
plt.ylabel('Distribution',fontsize=20)
plt.xlim([0, 35])
plt.ylim([0, 0.4])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/boltzman_temp', dpi=300)
# -
temp = pd.read_csv("../Data/temperature.txt")
# +
fig = plt.figure(figsize=(10,10))
for i in temp.columns:
if i =='levels':
continue
plt.plot(temp.levels,temp[i], label=f"T={i}")
plt.title("Boltzmann Distribution",fontsize=25)
plt.xlabel('Energy Levels',fontsize=20)
plt.ylabel('Distribution',fontsize=20)
plt.xlim([0, 35])
plt.ylim([0, 0.4])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/boltzman_temp_2', dpi=300)
# -
# ## Degeneracy
# +
NumberOfEnergyLevels = int(input("NumberOfEnergyLevels ?"))
Temperature = [0.1, 1, 10, 100, 1000, 10000, 100000]
Beta = [1/x for x in Temperature]
tmp = []
fig = plt.figure(figsize=(10,10))
for b in Beta[:]:
for i in range(NumberOfEnergyLevels):
tmp.append((i+1)*np.exp(-b*i))
Distribution = [x/sum(tmp) for x in tmp]
plt.plot(range(NumberOfEnergyLevels), Distribution, label=f"T={round(1/b)}")
Distribution = []
tmp = []
plt.title("Boltzmann Distribution",fontsize=25)
plt.xlabel('Energy Levels',fontsize=20)
plt.ylabel('Distribution',fontsize=20)
plt.xlim([0, 35])
plt.ylim([0, 0.4])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/boltzman_deg', dpi=300)
# -
# ## Linear Rotor
# +
NumberOfEnergyLevels = int(input("NumberOfEnergyLevels ?"))
Temperature = [0.1, 1, 10, 100, 1000, 10000, 100000]
Beta = [1/x for x in Temperature]
tmp = []
fig = plt.figure(figsize=(10,10))
for b in Beta[:]:
for i in range(NumberOfEnergyLevels):
tmp.append((i+1)*np.exp(-b*i))
Distribution = [x/sum(tmp) for x in tmp]
plt.plot(range(NumberOfEnergyLevels), Distribution, label=f"T={round(1/b)}")
Distribution = []
tmp = []
plt.title("Boltzmann Distribution",fontsize=25)
plt.xlabel('Energy Levels',fontsize=20)
plt.ylabel('Distribution',fontsize=20)
plt.xlim([0, 35])
plt.ylim([0, 0.4])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/boltzman_deg', dpi=300)
# +
NumberOfEnergyLevels = int(input("NumberOfEnergyLevels ?"))
Temperature = [0.1, 1, 10, 100, 1000, 10000, 100000]
Beta = [1/x for x in Temperature]
tmp = []
fig = plt.figure(figsize=(10,10))
for b in Beta[:]:
for i in range(NumberOfEnergyLevels):
tmp.append((2*i+1)*np.exp(-b*(0.5*i*(i+1))))
Distribution = [x/sum(tmp) for x in tmp]
plt.plot(range(NumberOfEnergyLevels), Distribution, label=f"T={round(1/b)}")
Distribution = []
tmp = []
plt.title("Boltzmann Distribution",fontsize=25)
plt.xlabel('Energy Levels',fontsize=20)
plt.ylabel('Distribution',fontsize=20)
plt.xlim([0, 35])
plt.ylim([0, 0.4])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/boltzman_rotor', dpi=300)
# -
# ## Comparison with approximate results
Temp=100
q = 2*Beta*1.38064852
q
# # Coupled harmonic oscillators
# +
# %reset
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import random
import math
MAX_NUMBER_OF_OSCILLATORS = 10000
MAX_ENERGY = 100000
CycleMultiplication = 1000
TotalEnergy=0
Beta=0.0
NumberOfCycles = int(input("Number of Cycles? "))
if NumberOfCycles < 3:
print("Number of Cycles must > 3")
Choice = int(input("NVE Ensemble (1) or NPT Ensemble (0) ?"))
if Choice:
TotalEnergy = int(input("Total Energy? "))
TotalOscillators = [4, 8, 10, 15, 25, 40, 50, 100]
#NumberOfOscillators = int(input("Number of Oscillators? "))
else:
Beta = int(input("Beta? "))
NumberOfOscillators = 1
fig = plt.figure(figsize=(10,10))
for NumberOfOscillators in TotalOscillators[:]:
Oscillator = np.zeros(MAX_NUMBER_OF_OSCILLATORS)
Distribution = np.zeros(MAX_ENERGY)
Ninit=NumberOfCycles/2
Sum=0.0
Count=0.0
Normalization=0.0
Utot=0;
if Choice:
i=0
while Utot != TotalEnergy:
if i>=NumberOfOscillators:
i=0
Oscillator[i]+=1
i+=1
Utot+=1
Utot=0
Utot=sum(Oscillator)
print("Initial energy : {}\n".format(Utot))
for i in range(NumberOfCycles):
for j in range(CycleMultiplication):
for k in range(NumberOfOscillators):
if Choice:
OscA = int(random.random()*NumberOfOscillators)
OscB = int(random.random()*NumberOfOscillators)
while OscA == OscB:
OscA = int(random.random()*NumberOfOscillators)
OscB = int(random.random()*NumberOfOscillators)
if random.random() < 0.5:
A = 1
B = -1
else:
A = -1
B = 1
if min(Oscillator[OscA]+A,Oscillator[OscB]+B) >= 0:
Oscillator[OscA]+=A
Oscillator[OscB]+=B
break
else:
if random.random() < 0.5:
A=1
else:
A=-1
if Oscillator[0]+A >= 0 and random.random() < np.exp(-Beta*A):
Oscillator[0]+=A
break
if i > Ninit:
Distribution[int(Oscillator[0])]+=1.0
Normalization+=1.0
Sum+=Oscillator[0]
Count+=1.0
Utot=0
Utot = sum(Oscillator)
print("Final Energy: {}\n".format(Utot))
print("Average Energy First Oscillator 1: {}\n".format(Sum/Count))
distr = [x/Normalization for x in Distribution]
plt.plot(range(MAX_ENERGY), distr, label=f"Oscillators={NumberOfOscillators}")
plt.title("Energy distribution over the harmonic oscillators",fontsize=20)
plt.xlabel('Energy Levels',fontsize=20)
plt.ylabel('Ocupation',fontsize=20)
plt.xlim([0, 40])
plt.ylim([0, 0.5])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/harmonic_dis', dpi=300)
# +
# %reset
import pandas as pd
import numpy as np
import matplotlib.pyplot as plt
import random
import math
MAX_NUMBER_OF_OSCILLATORS = 10000
MAX_ENERGY = 100000
CycleMultiplication = 1000
TotalEnergy=0
Beta=0.0
NumberOfCycles = 1000
TotalChoice = [1,0]
fig = plt.figure(figsize=(10,10))
for Choice in TotalChoice:
if Choice:
TotalEnergy = 100
NumberOfOscillators = 50
else:
Beta = 0.5
NumberOfOscillators = 1
TotalEnergy=0
Oscillator = np.zeros(MAX_NUMBER_OF_OSCILLATORS)
Distribution = np.zeros(MAX_ENERGY)
Ninit=NumberOfCycles/2
Sum=0.0
Count=0.0
Normalization=0.0
Utot=0;
if Choice:
i=0
while Utot != TotalEnergy:
if i>=NumberOfOscillators:
i=0
Oscillator[i]+=1
i+=1
Utot+=1
Utot=0
Utot=sum(Oscillator)
print("Initial energy : {}\n".format(Utot))
for i in range(NumberOfCycles):
for j in range(CycleMultiplication):
for k in range(NumberOfOscillators):
if Choice:
OscA = int(random.random()*NumberOfOscillators)
OscB = int(random.random()*NumberOfOscillators)
while OscA == OscB:
OscA = int(random.random()*NumberOfOscillators)
OscB = int(random.random()*NumberOfOscillators)
if random.random() < 0.5:
A = 1
B = -1
else:
A = -1
B = 1
if min(Oscillator[OscA]+A,Oscillator[OscB]+B) >= 0:
Oscillator[OscA]+=A
Oscillator[OscB]+=B
break
else:
if random.random() < 0.5:
A=1
else:
A=-1
if Oscillator[0]+A >= 0 and random.random() < np.exp(-Beta*A):
Oscillator[0]+=A
break
if i > Ninit:
Distribution[int(Oscillator[0])]+=1.0
Normalization+=1.0
Sum+=Oscillator[0]
Count+=1.0
Utot=0
Utot = sum(Oscillator)
print("Final Energy: {}\n".format(Utot))
print("Average Energy First Oscillator 1: {}\n".format(Sum/Count))
distr = [x/Normalization for x in Distribution]
if Choice:
plt.plot(range(MAX_ENERGY), distr, label=f"NVE (Energy={TotalEnergy})")
else:
plt.plot(range(MAX_ENERGY), distr, label=f"NVT (Beta={Beta})")
plt.title("Energy distribution over the harmonic oscillators for NVE & NVT ensemble",fontsize=20)
plt.xlabel('Energy Levels',fontsize=20)
plt.ylabel('Ocupation',fontsize=20)
plt.xlim([0, 40])
plt.ylim([0, 0.5])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/NVT', dpi=300)
# -
# # Random Walk on a 1D lattice
# +
MAX_LATTICE = 1000
CycleMultiplication = 100
NumberOfCycles = 100
NumberOfJumps = int(input("Number of Jumps/Cycle ? "))
fig = plt.figure(figsize=(10,10))
Distribution = np.zeros(2*NumberOfJumps+1)
Normalization = 0
for i in range(NumberOfCycles):
for j in range(CycleMultiplication):
CurrentPosition=0
for k in range(NumberOfJumps):
if random.random() < 0.5:
CurrentPosition+=1
else:
CurrentPosition-=1
Distribution[CurrentPosition+NumberOfJumps]+=1
Normalization+=1.0
distr = [x/(2*Normalization) for x in Distribution if x != 0]
n_range = np.linspace(-MAX_LATTICE,MAX_LATTICE,MAX_LATTICE*2+1)
n_2_range = np.linspace(-NumberOfJumps,NumberOfJumps,NumberOfJumps+1)
theoritical = 0.5*np.exp(0.5*np.log(2/(NumberOfJumps*np.pi)) - ((n_range**2)/(2*NumberOfJumps)))
plt.plot(n_2_range, distr, linestyle='dashed', label=f"Random walk")
plt.plot(n_range, theoritical, label=f"Analytical")
plt.title("Position distribution with different probability [1D random walk]",fontsize=20)
plt.xlabel('Position relative to origin',fontsize=20)
plt.ylabel('Probability densitiy',fontsize=20)
plt.xlim([-(NumberOfJumps+2), NumberOfJumps+2])
plt.ylim([0, 0.25])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/1D', dpi=300)
# +
MAX_LATTICE = 1000
CycleMultiplication = 100
NumberOfCycles = 100
NumberOfJumps = int(input("Number of Jumps/Cycle ? "))
TotalProbability = [0.5, 0.6, 0.7,0.95]
fig = plt.figure(figsize=(10,10))
rmsd = []
RMSDCurrentPosition = 0
for probability in TotalProbability:
Distribution = np.zeros(2*NumberOfJumps+1)
Normalization = 0
print(probability)
final_distr = []
for i in range(NumberOfCycles):
for j in range(CycleMultiplication):
CurrentPosition=0
for k in range(NumberOfJumps):
if random.random() < probability:
CurrentPosition+=1
else:
CurrentPosition-=1
Distribution[CurrentPosition+NumberOfJumps]+=1
Normalization+=1.0
rmsd.append((CurrentPosition - RMSDCurrentPosition)**2)
RMSDCurrentPosition = CurrentPosition
#distr = [x/(2*Normalization) for x in Distribution if x != 0]
distr = [x/(Normalization) for x in Distribution]
index = []
distrrrr = []
for i in range(len(distr)):
if distr[i] != 0:
index.append(i)
distrrrr.append(distr[i])
range_n3= np.linspace(-len(index)/2,len(index)/2,len(index))
#for i in rang(MAX_LATTICE*2):
# if distr[i+MAX_LATTICE]>0.5:
# final_distr.append(distr)
#n_2_range = np.linspace(-MAX_LATTICE,MAX_LATTICE,MAX_LATTICE*2+1)
plt.plot(range_n3, distrrrr, linestyle='dashed', label=f"Random walk (P={probability})")
plt.title("Position distribution with different probability [1D random walk]",fontsize=20)
plt.xlabel('Position relative to origin',fontsize=20)
plt.ylabel('Probability densitiy',fontsize=20)
plt.xlim([-20, 20])
plt.ylim([0, 0.4])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.savefig('../Images/1D_proba', dpi=300)
# +
rmsd = [x/sum(rmsd) for x in rmsd]
fig = plt.figure(figsize=(10,10))
plt.plot(range(len(rmsd)), rmsd, linestyle='-', label=f"RMSD")
plt.title("Position distribution with different probability [1D random walk]",fontsize=20)
plt.xlabel('Position relative to origin',fontsize=20)
plt.ylabel('Probability densitiy',fontsize=20)
plt.xlim([-20, 20])
plt.ylim([0, 0.4])
plt.legend(loc='best', frameon=True,fontsize=20)
fig.show()
fig.savefig('../Images/1D_proba', dpi=300)
| block1/Notebook/Block_1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
import sys, os
## rewrite it with abs path
sys.path.insert(1, '../base_solver/base_solver_char')
import numpy as np
import torch
import torch.nn as nn
from torch.autograd import Variable
from torch.utils.data import DataLoader, Dataset
from torchvision.utils import save_image
import torchvision.transforms as transforms
import captcha_setting
import my_dataset
from captcha_cnn_model import CNN, Generator
import cv2 as cv
from matplotlib import pyplot as plt
from PIL import Image
import copy
import operator
class CNN(nn.Module):
def __init__(self):
super(CNN, self).__init__()
self.layer1 = nn.Sequential(
nn.Conv2d(1, 32, kernel_size=3, padding=1),
nn.BatchNorm2d(32),
nn.Dropout(0.1), # drop 50% of the neuron
nn.ReLU(),
nn.MaxPool2d(2))
self.layer2 = nn.Sequential(
nn.Conv2d(32, 64, kernel_size=3, padding=1),
nn.BatchNorm2d(64),
nn.Dropout(0.1), # drop 50% of the neuron
nn.ReLU(),
nn.MaxPool2d(2))
self.layer3 = nn.Sequential(
nn.Conv2d(64, 64, kernel_size=3, padding=1),
nn.BatchNorm2d(64),
nn.Dropout(0.1), # drop 50% of the neuron
nn.ReLU(),
nn.MaxPool2d(2))
self.fc = nn.Sequential(
nn.Linear((captcha_setting.IMAGE_WIDTH//8)*(captcha_setting.IMAGE_HEIGHT//8)*64, 1024),
nn.Dropout(0.1), # drop 50% of the neuron
nn.ReLU())
self.rfc = nn.Sequential(
nn.Linear(1024, 256),#captcha_setting.MAX_CAPTCHA*captcha_setting.ALL_CHAR_SET_LEN),
nn.ReLU()
)
self.rfc2 = nn.Sequential(
nn.Linear(256, 10),
)
def forward(self, x):
out = self.layer1(x)
out = self.layer2(out)
out = self.layer3(out)
out = out.view(out.size(0), -1)
out = self.fc(out)
#print(out.shape)
out = self.rfc(out)
out = self.rfc2(out)
#out = out.view(out.size(0), -1)
#print(out.shape)
return out
device = torch.device("cuda:0" if torch.cuda.is_available() else "cpu")
cnn = CNN()
cnn.eval()
# -
# # Simply test the DWGAN, segmentation & solver
# +
# Need to redefine the path here>>>>
img_path = "img_path"
GAN_PATH = '../model/DWGAN_model.pkl'
#<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<<
label = ''
def gaussian_blur(img):
image = np.array(img)
image_blur = cv.GaussianBlur(image,(5,5),0)
new_image = image_blur
return new_image
class testdataset(Dataset):
def __init__(self, folder, transform=None):
self.train_image_file_paths = [os.path.join(folder, image_file) for image_file in os.listdir(folder)]
self.transform = transform
def __len__(self):
return len(self.train_image_file_paths)
def __getitem__(self, idx):
image_root = self.train_image_file_paths[idx]
image_name = image_root.split(os.path.sep)[-1]
image = Image.open(image_root)
image = image.resize((160,60), Image.ANTIALIAS)
label = image_name
image = self.transform(image)
return image, label
transform_1 = transforms.Compose([
# transforms.ColorJitter(),
transforms.Grayscale(),
transforms.ToTensor(),
# transforms.Normalize(mean=[0.485, 0.456, 0.406], std=[0.229, 0.224, 0.225])
])
def get_loader():
dataset = testdataset(img_path, transform=transform_1)
return DataLoader(dataset, batch_size=1, shuffle=False)
dataloader = get_loader()
generator = Generator()
generator.load_state_dict(torch.load(GAN_PATH))
generator.eval()
print("load GAN net.")
label_target = ""
for i, (imgs, label) in enumerate(dataloader):
if(i<32):
continue
print(label)
label_target = label
imgs = torch.tensor(imgs).float()
new_img = generator(imgs)
new_img2 = new_img.data.cpu().numpy()
imgs2 = imgs.data.cpu().numpy()
imgs2 = imgs2[0][0]
imgs2 = imgs2*255
target_img = new_img2[0][0]
target_img = target_img*255
cv.imwrite( "temp.jpg",imgs2)
cv.imwrite( "temp_2.jpg",target_img)
plt.imshow(imgs2)
plt.show()
plt.imshow(target_img)
plt.show()
break
# -
img = cv.imread('temp.jpg')
img_t = cv.imread('temp_2.jpg')
plt.imshow(img)
plt.show()
threshold = 5
n_img = np.zeros((img.shape[0],img.shape[1]))
img_aft = cv.normalize(img, n_img, 0,255,cv.NORM_MINMAX)
plt.imshow(img_aft)
plt.show()
gray = cv.cvtColor(img_aft,cv.COLOR_BGR2GRAY)
ret, thresh = cv.threshold(gray,0,255,cv.THRESH_BINARY_INV+cv.THRESH_OTSU)
plt.imshow(thresh)
plt.show()
def calculate_corner(thresh, nrootdir="cut_image/"):
show_img = cv.imread('temp.jpg')
im2,contours,hierarchy = cv.findContours(thresh,cv.RETR_EXTERNAL,cv.CHAIN_APPROX_SIMPLE)
# print(contours[0])
new_contours = []
cur_contours = []
filter_containor = []
for i in contours:
#print(i)
x, y, w, h = cv.boundingRect(i)
cur_contours.append([x, y, w, h])
contours = sorted(cur_contours, key=operator.itemgetter(0))
for i in range(0,len(contours)):
x = contours[i][0]
y = contours[i][1]
w = contours[i][2]
h = contours[i][3]
newimage=thresh[y:y+h,x:x+w]
nrootdir=("cut_image/")
new_contours.append(contours[i])
t0 = [i[0] for i in new_contours]
t1 = [i[1] for i in new_contours]
t2 = [i[2] for i in new_contours]
t3 = [i[3] for i in new_contours]
x_max = max([new_contours[i][0] + new_contours[i][2] for i in range(len(new_contours))] )+5
x_min = min(t0)
y_max = max([new_contours[i][1] + new_contours[i][3] for i in range(len(new_contours))] )
y_min = min(t1)
x_max = 159
x_min = 0
y_max = 59
y_min = 0
width = (x_max-x_min)//4
for i in range(0,4):
newimage=thresh[y_min:y_max,x_min+i*width:x_min+(i+1)*width]
top, bottom, left, right = [1]*4
newimage = cv.copyMakeBorder(newimage, top, bottom, left, right, cv.BORDER_CONSTANT)
newimage = cv.resize(newimage,(30, 60), interpolation = cv.INTER_CUBIC)
cv.imwrite( "temp.jpg",newimage)
filter_containor.append(Image.open("temp.jpg"))
return filter_containor
# # dataset making for seg
# +
# Rewrite the path for data source and where data will be saved
file_path = ""
save_path = ""
transform = transforms.Compose([
transforms.Grayscale(),
transforms.ToTensor()
])
llist = {'0':0,'1':0,'2':0,'3':0,'4':0,'5':0,'6':0,'7':0,'8':0,'9':0}
test_img_list = os.listdir(file_path)
for img_name in test_img_list:
if('_' in img_name):
label = img_name.split('_')[0].upper()
else:
label = img_name.split('.')[0].upper()
#using GAN
image = Image.open(file_path+img_name)
image = image.resize((160,60), Image.ANTIALIAS)
image = transform(image)
imgs = torch.tensor(image).float()
new_img = generator(imgs)
new_img2 = new_img.data.cpu().numpy()
imgs2 = imgs.data.cpu().numpy()
imgs2 = imgs2[0][0]
imgs2 = imgs2*255
target_img = new_img2[0][0]
target_img = target_img*255
im = Image.fromarray(target_img)
im = im.convert('RGB')
im.save('temp.png')
img = cv.imread('temp.png')
# end using GAN
img = cv.resize(img,(160, 60), interpolation = cv.INTER_CUBIC)
img_aft = cv.normalize(img, n_img, 0,255,cv.NORM_MINMAX)
gray = cv.cvtColor(img_aft,cv.COLOR_BGR2GRAY)
ret, thresh = cv.threshold(gray,0,255,cv.THRESH_BINARY_INV+cv.THRESH_OTSU)
filter_containor = calculate_corner(thresh)
for i in range(0,4):
try:
if(not os.path.exists(save_path+label[i])):
os.mkdir(save_path+label[i])
filter_containor[i].save( save_path+label[i]+'/'+str(llist[label[i]])+'.png')
llist[label[i]] += 1
except:
print(label, i)
break
# -
# # Go Through All Test Data
# +
file_path = "/home/ning_a/Desktop/CAPTCHA/dark_web_captcha/rescator_data/test/"
save_path = "/home/ning_a/Desktop/CAPTCHA/dark_web_captcha/rescator_data/test_char/"
transform = transforms.Compose([
# transforms.ColorJitter(),
transforms.Grayscale(),
# transforms.Lambda(gaussian_blur),
transforms.ToTensor()
])
llist = {'0':0,'1':0,'2':0,'3':0,'4':0,'5':0,'6':0,'7':0,'8':0,'9':0}
test_img_list = os.listdir(file_path)
total = 0
correct = 0
cnn = CNN()
cnn.eval()
cnn.load_state_dict(torch.load('./model_lake/recator_1_char_1.pkl'))
cnn.to(device)
for img_name in test_img_list:
total += 1
if('_' in img_name):
label = img_name.split('_')[0].upper()
else:
label = img_name.split('.')[0].upper()
# #using GAN
# #Uncommon following to enable GAN
# image = Image.open(file_path+img_name)
# image = image.resize((160,60), Image.ANTIALIAS)
# image = transform(image)
# imgs = torch.tensor(image).float()
# new_img = generator(imgs)
# new_img2 = new_img.data.cpu().numpy()
# imgs2 = imgs.data.cpu().numpy()
# imgs2 = imgs2[0][0]
# imgs2 = imgs2*255
# target_img = new_img2[0][0]
# target_img = target_img*255
# im = Image.fromarray(target_img)
# im = im.convert('RGB')
# im.save('temp.png')
# img = cv.imread('temp.png')
# # end using GAN
img = cv.imread(file_path+img_name)
img = cv.resize(img,(160, 60), interpolation = cv.INTER_CUBIC)
img_aft = cv.normalize(img, n_img, 0,255,cv.NORM_MINMAX)
gray = cv.cvtColor(img_aft,cv.COLOR_BGR2GRAY)
ret, thresh = cv.threshold(gray,0,255,cv.THRESH_BINARY_INV+cv.THRESH_OTSU)
filter_containor = calculate_corner(thresh)
p_lable = ''
for i in range(0,4):
image = transform(filter_containor[i]).unsqueeze(0)
image = torch.tensor(image, device=device).float()
image = Variable(image).to(device)
predict_label = cnn(image)
predict_label = predict_label.cpu()
_, predicted = torch.max(predict_label, 1)
p_lable += captcha_setting.ALL_CHAR_SET[predicted]
if(p_lable.upper() == label.upper()):
correct += 1
print(correct)
print(total)
print(float(correct/total))
| Example_and_Record/notebook_example/.ipynb_checkpoints/case_rescator_1-checkpoint.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# name: python3
# ---
# + [markdown] id="q1BH5vnm4W6W"
# ```
# 4. Preprocessing Pipeline
# The partner must:
# describe the data preprocessing pipeline, and
# how this is accomplished via a package/function that is a callable API (that is ultimately accessed by the served, production model).
#
# Evidence must include a description (in the Whitepaper) of how data preprocessing is accomplished, along with the code snippet that accomplishes data preprocessing as a callable API.
# ```
# + [markdown] id="6x1ypzczQCwy"
# # Deploy model using TFX Pipeline
# To deploy the model we will following GCP best practises and use TensorFlow Extended (TFX). A TensorFlow pipeline is a sequence of components taht impoement an ML Pipeline which is specificially designed for scale, deployment, and retraining.
#
# To successfully deploy the model we will need address the 3 phases of the pipeline:
# 1. Ingest & Validate Data
# - ExampleGen
# - StatisticsGen
# - SchemaGen
# - ExampleValidator
# 2. Train & Analyze Model
# - Transform
# - Trainer
# 3. Deploy in Production
# - Pusher
# + [markdown] id="WC9W_S-bONgl"
# ### Install python packages
# We will install required Python packages including TFX and KFP to author ML pipelines and submit jobs to Vertex Pipelines. We will be deploying the TFX pipeline onto the Apache Beam orchestrator.
# + id="iyQtljP-qPHY" outputId="8a87f212-7215-410d-dcd3-d12a2fb5eed3" colab={"base_uri": "https://localhost:8080/"}
# Use the latest version of pip.
# !pip install --upgrade pip
# !pip install --upgrade "tfx[kfp]<2"
# !pip install --upgrade tensorflow_transform
# + [markdown] id="EwT0nov5QO1M"
# ### Restart the Runtime
# You will need to restart the runtime for the libraries to be available in Google Collab. Runtime > Restart Runtime
# + [markdown] id="gckGHdW9iPrq"
# ### Login in to Google for this *notebook*
# + id="kZQA0KrfXCvU"
import sys
if 'google.colab' in sys.modules:
from google.colab import auth
auth.authenticate_user()
# + [markdown] id="3_SveIKxaENu"
# ### Check the package versions.
# + id="Xd-iP9wEaENu" colab={"base_uri": "https://localhost:8080/"} outputId="4582eaeb-e6a1-46a5-ca31-ada3a64359c7"
import tensorflow as tf
print('TensorFlow version: {}'.format(tf.__version__))
from tfx import v1 as tfx
print('TFX version: {}'.format(tfx.__version__))
import kfp
print('KFP version: {}'.format(kfp.__version__))
# + [markdown] id="aDtLdSkvqPHe"
# ### Set up variables
#
# We will set up some variables used to customize the pipelines below. Following
# information is required:
#
# * GCP Project id.
# * GCP Region to run pipelines.
# * Google Cloud Storage Bucket to store pipeline outputs.
# + id="EcUseqJaE2XN"
GOOGLE_CLOUD_PROJECT = 'ml-spec-demo-2-sandbox'
GOOGLE_CLOUD_REGION = 'australia-southeast1'
GCS_BUCKET_NAME = 'black_friday_gcp_bucket'
# + [markdown] id="GAaCPLjgiJrO"
# #### Set `gcloud` to use your project.
# + id="VkWdxe4TXRHk" colab={"base_uri": "https://localhost:8080/"} outputId="bd93c29f-9870-45da-eab2-7180b8b01eca"
{GOOGLE_CLOUD_PROJECT}
# + [markdown] id="l_x_8xoj6k0L"
# ### Set up Global variables for model serving locations
# + id="CPN6UL5CazNy" colab={"base_uri": "https://localhost:8080/"} outputId="75956013-1a23-4e03-9162-99b0709e2dcf"
PIPELINE_NAME = 'black-friday-gcp-vertex-pipelines'
# Path to pipeline artifacts
PIPELINE_ROOT = 'gs://{}/pipeline_root/{}'.format(
GCS_BUCKET_NAME, PIPELINE_NAME)
# Paths for users' Python module
MODULE_ROOT = 'gs://{}/pipeline_module/{}'.format(
GCS_BUCKET_NAME, PIPELINE_NAME)
# Paths to training data
DATA_ROOT = 'gs://{}/data/{}'.format(GCS_BUCKET_NAME, PIPELINE_NAME)
# This is the path where your model will be pushed for serving
SERVING_MODEL_DIR = 'gs://{}/serving_model/{}'.format(
GCS_BUCKET_NAME, PIPELINE_NAME)
# Training data file name
FILE_NAME = 'train.csv'
print('PIPELINE_ROOT: {}'.format(PIPELINE_ROOT))
print('Data root: {}'.format(DATA_ROOT))
# + [markdown] id="11J7XiCq6AFP"
# We need to make our own copy of the dataset. Because TFX ExampleGen reads
# inputs from a directory, we need to create a directory and copy dataset to it
# on GCS.
# + [markdown] id="ASpoNmxKSQjI"
# Take a quick look at the CSV file.
# + id="-eSz28UDSnlG" colab={"base_uri": "https://localhost:8080/"} outputId="509dfedd-6c1e-4017-a2f4-8e5caa9dd254"
# !gsutil cat {DATA_ROOT}/train.csv | head
# + [markdown] id="nH6gizcpSwWV"
# ## Create a pipeline
#
# TFX pipelines are defined using Python APIs. We will define a pipeline which
# consists of three components, CsvExampleGen, Trainer and Pusher. The pipeline
# and model definition is almost the same as
# [Simple TFX Pipeline Tutorial](https://www.tensorflow.org/tfx/tutorials/tfx/penguin_simple).
#
# The only difference is that we don't need to set `metadata_connection_config`
# which is used to locate
# [ML Metadata](https://www.tensorflow.org/tfx/guide/mlmd) database. Because
# Vertex Pipelines uses a managed metadata service, users don't need to care
# of it, and we don't need to specify the parameter.
#
# Before actually define the pipeline, we need to write a model code for the
# Trainer component first.
# + [markdown] id="VyBh_2LO4Ccw"
# # Write Example Component
#
# + [markdown] id="lOjDv93eS5xV"
# ### Write model code.
#
# We will use the same model code as in the
# [Simple TFX Pipeline Tutorial](https://www.tensorflow.org/tfx/tutorials/tfx/penguin_simple).
# + id="aES7Hv5QTDK3"
_trainer_module_file = 'bfs_trainer.py'
_transformer_module_file = 'transformer.py'
_training_pipeline_file = 'training_pipeline.py'
# + id="OB3fD-DSuAJc" colab={"base_uri": "https://localhost:8080/"} outputId="2ef1dc42-31f1-4081-dd9a-0910c8208ccb"
# %%writefile {_transformer_module_file}
from typing import Dict, Text, Any, List
import tensorflow as tf
import tensorflow_transform as tft
FEATURES = [
'Product_ID',
'Gender',
'Age',
'Occupation',
'City_Category',
'Stay_In_Current_City_Years',
'Marital_Status',
'Product_Category_1',
'Product_Category_2',
'Purchase'
]
CATEGORICAL_FEATURE_KEYS = [
'Product_ID',
'Age',
'City_Category',
'Product_Category_1',
'Product_Category_2',
'Stay_In_Current_City_Years',
'Gender'
]
OPTIONAL_NUMERIC_KEY_FEATURES = ['Product_Category_2']
def preprocessing_fn(inputs: Dict[Text, Any], custom_config) -> Dict[Text, Any]:
"""tf.transform's callback function for preprocessing inputs.
Args:
inputs: map from feature keys to raw not-yet-transformed features.
custom_config:
timesteps: The number of timesteps in the look back window
features: Which of the features from the TF.Example to use in the model.
Returns:
Map from string feature key to transformed feature operations.
"""
print('Start preprocessing')
outputs = {}
for key in FEATURES:
key_l = key.lower()
outputs[key_l] = inputs[key]
# Convert optional categories to sparse tensor (fills in blank values basically)
for key in OPTIONAL_NUMERIC_KEY_FEATURES:
key_l = key.lower()
sparse = tf.sparse.SparseTensor(inputs[key].indices, inputs[key].values,
[inputs[key].dense_shape[0], 1])
dense = tf.sparse.to_dense(sp_input=sparse, default_value=0)
# Reshaping from a batch of vectors of size 1 to a batch to scalars.
dense = tf.squeeze(dense, axis=1)
outputs[key_l] = dense
for key in CATEGORICAL_FEATURE_KEYS:
key_l = key.lower()
outputs[key_l] = tft.compute_and_apply_vocabulary(inputs[key])
return outputs
# + id="Gnc67uQNTDfW" colab={"base_uri": "https://localhost:8080/"} outputId="46575526-ccab-4867-b7c5-85b85e49d8b0"
# %%writefile {_trainer_module_file}
from typing import List
from absl import logging
import tensorflow as tf
from tensorflow import keras
from tensorflow_transform.tf_metadata import schema_utils
from tensorflow.keras import layers
from tfx_bsl.tfxio import dataset_options
from tfx import v1 as tfx
from tfx_bsl.public import tfxio
import tensorflow_transform as tft
from tfx.components.example_gen import utils as example_gen_utils
from tensorflow_metadata.proto.v0 import schema_pb2
FEATURES = [
'product_id',
'gender',
'age',
'occupation',
'city_category',
'stay_in_current_city_years',
'marital_status',
'product_category_1',
'product_category_2'
]
LABEL = 'purchase'
# NEW: This function will create a handler function which gets a serialized
# tf.example, preprocess and run an inference with it.
def _get_serve_tf_examples_fn(model, tf_transform_output):
# We must save the tft_layer to the model to ensure its assets are kept and
# tracked.
model.tft_layer_inference = tf_transform_output.transform_features_layer()
@tf.function(input_signature=[
tf.TensorSpec(shape=[None], dtype=tf.string, name='examples')
])
def serve_tf_examples_fn(serialized_tf_examples):
"""Returns the output to be used in the serving signature."""
feature_spec = tf_transform_output.raw_feature_spec()
feature_spec.pop("Purchase")
feature_spec.pop("User_ID")
parsed_features = tf.io.parse_example(serialized_tf_examples, feature_spec)
transformed_features = model.tft_layer_inference(parsed_features)
outputs = model(transformed_features)
return {'outputs': outputs}
return serve_tf_examples_fn
def _make_keras_model() -> tf.keras.Model:
"""Creates a DNN Keras model for classifying penguin data.
Returns:
A Keras Model.
"""
inputs = [keras.layers.Input(shape=(1,), name=f) for f in FEATURES]
d = keras.layers.concatenate(inputs)
d = keras.layers.Dense(128, activation='relu')(d)
d = keras.layers.Dense(256, activation='relu')(d)
d = keras.layers.Dense(128, activation='relu')(d)
outputs = keras.layers.Dense(1)(d)
model = tf.keras.Model(inputs=inputs, outputs=outputs)
model.compile(
optimizer=tf.optimizers.Adam(learning_rate=0.0005),
loss=tf.keras.losses.MeanSquaredError(),
metrics=[keras.metrics.MeanSquaredError()]
)
model.summary(print_fn=logging.info)
return model
# TFX Trainer will call this function.
def run_fn(fn_args: tfx.components.FnArgs):
"""Train the model based on given args.
Args:
fn_args: Holds args used to train the model as name/value pairs.
"""
# This schema is usually either an output of SchemaGen or a manually-curated
# version provided by pipeline author. A schema can also derived from TFT
# graph if a Transform component is used. In the case when either is missing,
# `schema_from_feature_spec` could be used to generate schema from very simple
# feature_spec, but the schema returned would be very primitive.
# get transform component output
tf_transform_output = tft.TFTransformOutput(fn_args.transform_output)
# read input data
train_dataset = fn_args.data_accessor.tf_dataset_factory(
fn_args.train_files,
dataset_options.TensorFlowDatasetOptions(
batch_size=20,
label_key=LABEL
),
tf_transform_output.transformed_metadata.schema,
)
eval_dataset = fn_args.data_accessor.tf_dataset_factory(
fn_args.eval_files,
dataset_options.TensorFlowDatasetOptions(
batch_size=10,
label_key=LABEL
),
tf_transform_output.transformed_metadata.schema,
)
model = _make_keras_model()
# Train model
model.fit(
train_dataset,
steps_per_epoch=fn_args.train_steps,
validation_data=eval_dataset,
validation_steps=fn_args.eval_steps
)
# The layer has to be saved to the model for keras tracking purpases.
model.tft_layer = tf_transform_output.transform_features_layer()
signatures = {
'serving_default': _get_serve_tf_examples_fn(model, tf_transform_output)
}
# The result of the training should be saved in `fn_args.serving_model_dir`
# directory.
model.save(fn_args.serving_model_dir, save_format='tf', signatures=signatures)
# + [markdown] id="-j54Ya94tR1-"
# ### Copy files to bucket
# The transform and trainer module files need to be copied over to the GCP bucket for TFX to read.
# + id="rMMs5wuNYAbc" colab={"base_uri": "https://localhost:8080/"} outputId="d120a9c3-c107-47d2-bf8a-961f62f1502f"
# !gsutil cp {_trainer_module_file} {MODULE_ROOT}/
# !gsutil cp {_transformer_module_file} {MODULE_ROOT}/
# + [markdown] id="q-3jxJZcI7q-"
# ### Create TFX pipeline. This pipeline can then be passed onto an orchestrator, such as KubeFlow, for deployment.
# + id="jPiXvdhXvkGw" colab={"base_uri": "https://localhost:8080/"} outputId="ebb91161-5bef-4fec-c5a8-de4340fe7ed9"
import os
from absl import logging
import tensorflow as tf
from tensorflow import keras
from tensorflow_transform.tf_metadata import schema_utils
from tensorflow.keras import layers
from tfx import v1 as tfx
from tfx_bsl.public import tfxio
from tfx.orchestration.pipeline import Pipeline
from tfx.proto.trainer_pb2 import EvalArgs, TrainArgs
# docs_infra: no_execute
from google.cloud import aiplatform
from google.cloud.aiplatform import pipeline_jobs
def build_pipeline(pipeline_name, pipeline_root, serving_model_dir, data_root, file_name):
print("Running pipeline")
print("Creating example_gen")
# Generate Training Samples from Dataset stored on bucket.
example_gen = tfx.components.CsvExampleGen(
input_base=data_root
)
print("Creating statistics_gen")
# Generate statistics over data for visualization and example validation.
statistics_gen = tfx.components.StatisticsGen(
examples=example_gen.outputs["examples"]
)
print("Creating schema_gen")
# Generates schema based on statistic files
schema_gen = tfx.components.SchemaGen(
statistics=statistics_gen.outputs["statistics"],
#infer_feature_shape=True
)
print("Creating example_validator")
# Performs anomaly dection based on statistics and data schema
example_validator = tfx.components.ExampleValidator(
statistics=statistics_gen.outputs["statistics"],
schema=schema_gen.outputs["schema"]
)
print("Creating transform")
transform = tfx.components.Transform(
examples=example_gen.outputs["examples"],
schema=schema_gen.outputs["schema"],
module_file=os.path.join(MODULE_ROOT, _transformer_module_file)
)
print("Creating trainer")
# Trains the model
trainer = tfx.components.Trainer(
examples=transform.outputs["transformed_examples"],
transform_graph=transform.outputs["transform_graph"],
module_file=os.path.join(MODULE_ROOT, _trainer_module_file),
schema=schema_gen.outputs["schema"],
train_args=TrainArgs(num_steps=150),
eval_args=EvalArgs(num_steps=150)
)
print("Creating pusher")
# Pushes the trained model to vertex
pusher = tfx.components.Pusher(
trainer.outputs['model'],
push_destination=tfx.proto.PushDestination(
filesystem=tfx.proto.PushDestination.Filesystem(
base_directory=serving_model_dir)),
)
print("Creating tfx_pipeline")
tfx_pipeline = Pipeline(
pipeline_name=pipeline_name,
pipeline_root=pipeline_root,
components=[
example_gen,
statistics_gen,
schema_gen,
example_validator,
transform,
trainer,
pusher
],
data_root=data_root,
module_file=os.path.join(MODULE_ROOT, _trainer_module_file),
serving_model_dir=serving_model_dir,
enable_cache=False
)
pipeline_definition_file = pipeline_name + '_pipeline.json'
print("Creating runner")
runner = tfx.orchestration.experimental.KubeflowV2DagRunner(
config=tfx.orchestration.experimental.KubeflowV2DagRunnerConfig(),
output_filename=pipeline_definition_file)
print("Executing runner")
# Following function will write the pipeline definition to PIPELINE_DEFINITION_FILE.
_ = runner.run(tfx_pipeline)
def deploy_to_vertex(pipeline_name, project_name, cloud_region):
aiplatform.init(project=project_name, location=cloud_region)
pipeline_definition_file = pipeline_name + '_pipeline.json'
job = pipeline_jobs.PipelineJob(template_path=pipeline_definition_file,
display_name=pipeline_name)
job.run(sync=False)
build_pipeline(PIPELINE_NAME, PIPELINE_ROOT, SERVING_MODEL_DIR, DATA_ROOT, FILE_NAME)
# + [markdown] id="98QiNea9XtrK"
# # Deploy Pipeline to Vertex AI
# TFX can be deployed to Vertex AI. The build_pipeline function deploys the pipeline to using the Kubeflow V2 orchestrator.
# + id="S9g9yOH5wQfQ" colab={"base_uri": "https://localhost:8080/"} outputId="8e255fa2-9987-4926-e339-56835167160f"
deploy_to_vertex(PIPELINE_NAME, GOOGLE_CLOUD_PROJECT, GOOGLE_CLOUD_REGION)
# + [markdown] id="Z8dyPhVZJj_d"
# ### Model Accuracy
# You can check the model accuracy by reading the logs outputted by trainer module in Vertex AI. The accuracy seems to match the Neural Networks done in the exploration phase coming in with a RME of 80,064,008.
# + [markdown] id="XTEkHdhEvBQo"
# ### Setting Up Vertex AI Endpoints
#
# Once the model is trained it is placed into the Google Bucket under:
#
# ```
# black_friday_gcp_bucket/serving_model/black-friday-gcp-vertex-pipelines/#
# ```
#
# This model will need to be registered in Vertex AI's Model service.
#
# Once this model has been setup, Vertex AI's endpoint service can easily be configured to point to the registered model.
# + [markdown] id="HF8iAToXQPLU"
# ### Inferencing
# Once the endpoint has been set up it can be inferenced using a grpc request.
#
# To inference request's body structure is:
#
# ```
# {"instances":
# [{
# "examples": {
# b64: "<base64 encoded, serialized tensorflow example>"
# }
# }]
# }
# ```
#
#
# This request can be done using the following client code.
# + id="1fOecX2XHsYN"
# GCP Endpoint ID
ENDPOINT = "6519734516804747264"
# + id="zO3vEf2iLlOL"
import tensorflow as tf
# The following functions can be used to convert a value to a type compatible
# with tf.train.Example.
def _bytes_feature(value):
"""Returns a bytes_list from a string / byte."""
if isinstance(value, type(tf.constant(0))):
value = value.numpy() # BytesList won't unpack a string from an EagerTensor.
return tf.train.Feature(bytes_list=tf.train.BytesList(value=[value]))
def _float_feature(value):
"""Returns a float_list from a float / double."""
return tf.train.Feature(float_list=tf.train.FloatList(value=[value]))
def _int64_feature(value):
"""Returns an int64_list from a bool / enum / int / uint."""
return tf.train.Feature(int64_list=tf.train.Int64List(value=[value]))
def serialize_example(product_id, gender, age, occupation, city_category, stay_in_current_city_years, marital_status, product_category_1, product_category_2):
"""
Creates a tf.train.Example message ready to be written to a file.
"""
# Create a dictionary mapping the feature name to the tf.train.Example-compatible
# data type.
feature = {
'Product_ID': _bytes_feature(product_id),
'Gender': _bytes_feature(gender),
'Age': _bytes_feature(age),
'Occupation': _int64_feature(occupation),
'City_Category': _bytes_feature(city_category),
'Stay_In_Current_City_Years': _bytes_feature(stay_in_current_city_years),
'Marital_Status': _int64_feature(marital_status),
'Product_Category_1': _int64_feature(product_category_1),
'Product_Category_2': _int64_feature(product_category_2),
}
# Create a Features message using tf.train.Example.
example_proto = tf.train.Example(features=tf.train.Features(feature=feature))
return example_proto.SerializeToString()
# + id="kQjNcCNuI2uB"
# + colab={"base_uri": "https://localhost:8080/"} id="1WosjtWKLuAr" outputId="f9a0914d-e748-45c6-8471-d9a76ed1819b"
packet = {
"product_id": b'P00069042',
"gender": b'F',
"age": b'0-17',
"occupation": 10,
"city_category": b'A',
"stay_in_current_city_years": b'2',
"marital_status": 0,
"product_category_1": 2,
"product_category_2": 6
}
serialized_example = serialize_example(**packet)
print(f"Serialized_example: {serialized_example}")
# + [markdown] id="2mV2g9wTHMHM"
# Check that the serialized example correctly parses back.
# + colab={"base_uri": "https://localhost:8080/"} id="UCkSMK3r2wjy" outputId="e93a6365-95e5-47c2-9a2e-ec7097409d79"
import tensorflow as tf
example_proto = tf.train.Example.FromString(serialized_example)
example_proto
# + [markdown] id="wTrzJi2DHyLA"
# Encode packet into Base64, build packet and inference.
# + colab={"base_uri": "https://localhost:8080/"} id="CV2UH9G2HWj0" outputId="d56b52f6-ae17-4f52-d9a7-41d01dfd3239"
from google.cloud import aiplatform
import base64
b64_example = base64.b64encode(serialized_example).decode("utf-8")
print(f"Base64 encoded example: {b64_example}")
instances_packet = [{
"examples": {
"b64": b64_example
}
}]
aiplatform.init(project=GOOGLE_CLOUD_PROJECT, location='australia-southeast1')
endpoint = aiplatform.Endpoint(ENDPOINT)
prediction = endpoint.predict(instances=instances_packet)
print(prediction)
# + [markdown] id="mj_xUwkVU5Fl"
# ### Cloud Function
# To get an inference from the model a query needs to be made to the cloud function with the packet structure:
#
# ```
# {
# "User_ID: 1000047 #A Valid User_ID (Int)
# }
# ```
#
# The cloud function will then gather the top 10 Product_IDs the user is most_likely to buy, the User's profile, and Product Information before returning a summation of the Users expected spenditure for the month on these products.
#
# This code can be found in the "Cloud Function" folder under ``` main.py ```.
| Black_Friday_Sales_pipeline_using_TFX.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [conda root]
# language: python
# name: conda-root-py
# ---
# # SMC2017: Exercise set II
#
# ## Setup
# +
import numpy as np
from scipy import stats
import pandas as pd
# %matplotlib inline
import matplotlib.pyplot as plt
import seaborn as sns
sns.set_style()
path = '..\\..\\..\\..\\course_material\\exercise_sheets\\'
# -
# ## II.1 Likelihood estimates for the stochastic volatility model
#
# Consider the stochastic volatility model
# $$
# \begin{align}
# x_t\,|\,x_{t - 1} &\sim \mathcal{N}\left(\phi \cdot x_{t - 1},\,\sigma^2\right) \\
# y_t\,|\,x_t &\sim \mathcal{N}\left(0,\,\beta^2 \exp(x_t)\right) \\
# x_0 &\sim \mathcal{N}\left(0,\,\sigma^2\right)
# \end{align}
# $$
# with parameter vector $\theta = (\phi, \sigma, \beta)$.
# +
data = pd.read_csv(path + 'seOMXlogreturns2012to2014.csv',
header=None, names=['logreturn'])
y = data.logreturn.values
fig, ax = plt.subplots()
ax.plot(y)
# -
# ### a) Likelihood estimation for different values of $\beta$
#
# Consider fixed values for $\phi = 0.98$ and $\sigma = 0.16$. $\beta$ is allowed to vary between 0 and 2.
# +
theta = [0.98, 0.16]
def likelihood_bootstrap_pf(N, y, beta=0.70, resample=True, logweights=True):
# Cumulatively build up log-likelihood
ll = 0.0
# Initialisation
samples = stats.norm.rvs(0, theta[1], N)
weights = 1 / N * np.ones((N,))
weights_normalized = weights
# Determine the number of time steps
T = len(y)
# Loop through all time steps
for t in range(T):
# Resample
if resample:
# Randomly choose ancestors
ancestors = np.random.choice(samples, size=N,
replace=True, p=weights_normalized)
else:
ancestors = samples
# Propagate
samples = stats.norm.rvs(0, 1, N) * theta[1] + theta[0] * ancestors
if logweights:
# Weight
weights = stats.norm.logpdf(y[t], loc=0,
scale=(beta * np.exp(samples / 2)))
# Calculate the max of the weights
max_weights = np.max(weights)
# Subtract the max
weights = weights - max_weights
# Update log-likelihood
ll += max_weights + np.log(np.sum(np.exp(weights))) - np.log(N)
# Normalize weights to be probabilities
weights_normalized = np.exp(weights) / np.sum(np.exp(weights))
else:
# Weight
weights = stats.norm.pdf(y[t], loc=0,
scale=(beta * np.exp(samples / 2)))
# Update log-likelihood
ll += np.log(np.sum(weights)) - np.log(N)
# Normalize weights to be probabilities
weights_normalized = weights / np.sum(weights)
return ll
# -
# Run the bootstrap particle filter to estimate the log-likelihood.
def simulate(N=500, T=500, resample=True):
ll = []
beta_count = len(np.arange(0.5, 2.25, 0.1))
for beta in np.arange(0.5, 2.25, 0.1):
for i in range(10):
ll.append(likelihood_bootstrap_pf(N, y[:T], beta, resample))
ll = np.transpose(np.reshape(ll, (beta_count, 10)))
return ll
# +
fig, ax = plt.subplots(figsize=(10, 5))
ax.boxplot(simulate(500, 500), labels=np.arange(0.5, 2.25, 0.1));
# -
# ### b) Study how $N$ and $T$ affect the variance of the log-likelihood estimate
# +
variances = []
ns = [10, 15, 20, 25, 40, 50, 75, 100, 150, 200]
for N in ns:
lls = []
for i in range(50):
lls.append(likelihood_bootstrap_pf(N, y, beta=0.9))
# Calculate variance
variances.append(np.var(lls))
fig, ax = plt.subplots()
ax.plot(ns, variances, 'o-')
# -
# Variance reduces exponentially with growing $N$.
# +
variances = []
ts = range(10, 501, 35)
for T in ts:
lls = []
for i in range(60):
lls.append(likelihood_bootstrap_pf(200, y[:T], beta=0.9))
# Calculate variance
variances.append(np.var(lls))
fig, ax = plt.subplots()
ax.plot(ts, variances, 'o-')
# -
# Variance increases linearly with growing $T$.
# ### c) Study the influence of resampling on the variance of the estimator
# +
lls = np.zeros((60, 2))
# With resampling
for i in range(60):
lls[i, 0] = likelihood_bootstrap_pf(200, y, beta=0.9)
# Without resampling
for i in range(60):
lls[i, 1] = likelihood_bootstrap_pf(200, y, beta=0.9, resample=False)
fig, ax = plt.subplots()
ax.boxplot(lls, labels=['Resampling', 'No resampling']);
# -
# Without resampling the variance is larger and log-likelihood is generally lower.
# ## II.2 Fully adapted particle filter
# ### b) Implement the FAPF for model (ii) and compare the variance of the estimates of $\mathbb{E}(X_t\,|\,y_{1:t})$ to the estimates obtained by a bootstrap particle filter
#
# The state-space model under consideration is (normal distribution parametrized with $\sigma^2$)
# $$
# \begin{array}{rll}
# x_{t + 1} &= \cos(x_t)^2 + v_t, & v_t \sim N(0, 1) \\
# y_t &= 2 x_t + e_t, & e_t \sim N(0, 0.01)
# \end{array}
# $$
# which leads to the probabilistic model
# $$
# \begin{align}
# p(x_t\,|\,x_{t - 1}) &= N\left(x_t;\,\cos(x_t)^2,\,1\right) \\
# p(y_t\,|\,x_t) &= N\left(y_t;\,2 x_t,\,0.01\right)
# \end{align}
# $$
# This admits the necessary pdfs
# $$
# \begin{align}
# p(y_t\,|\,x_{t - 1}) &= N(y_t;\,2 \cos(x_{t - 1})^2,\,4.01) \\
# p(x_t\,|\,x_{t - 1},\,y_t) &= N\left(x_t;\,\frac{2 y_t + 0.01 \cos(x_{t - 1})^2}{4.01}, \frac{0.01}{4.01}\right)
# \end{align}
# $$
# Simulate a trajectory to use for the particle filters.
# +
T = 100
# Allocate arrays for results
ys = np.zeros((T,))
xs = np.zeros((T + 1,))
# Initial value for state
xs[0] = 0.1
# Walk through all time steps
for t in range(T):
xs[t + 1] = np.power(np.cos(xs[t]), 2) + stats.norm.rvs(0, 1, 1)
ys[t] = 2 * xs[t + 1] + stats.norm.rvs(0, 0.1, 1)
fig, axs = plt.subplots(2, 1, figsize=(10, 10))
axs[0].plot(range(T + 1), xs, 'o-');
axs[1].plot(range(1, T + 1), ys, 'o-r');
# -
def fully_adapted_PF(N, y):
# Save particles
xs = []
# Initialisation
samples = stats.norm.rvs(0, 1, N)
# Save initial data
xs.append(samples)
# Determine length of data
T = len(y)
for t in range(T):
# Calculate resampling weights in case of FAPF
resampling_weights = stats.norm.pdf(
y[t], loc=2*np.power(np.cos(samples), 2), scale=np.sqrt(4.01))
# Normalize the resampling weights
resampling_weights /= np.sum(resampling_weights)
# Resample
ancestors = np.random.choice(samples, size=N, replace=True,
p=resampling_weights)
# Propagate
samples = stats.norm.rvs(0, 1, N) * 0.1 / np.sqrt(4.01) + \
(2 / 4.01) * y[t] + (0.01 / 4.01) * np.power(np.cos(ancestors), 2)
# Save the new samples
xs.append(samples)
return np.array(xs)
# Try to recover the simulated states from the measurements.
xs_filtered = fully_adapted_PF(1000, ys)
# +
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(xs, 'ok')
ax.plot(np.apply_along_axis(np.mean, 1, xs_filtered), 'o-')
ax.legend(['Simulated data', 'FAPF'])
# -
# Holy shit :D
# For comparison, here is the bootstrap particle filter for this model
def bootstrap_PF(N, y):
# Save the history
xs = []
ws = []
# Initialisation
samples = stats.norm.rvs(0, 1, N)
weights = 1 / N * np.ones((N,))
weights_normalized = weights
# Save weights and samples
ws.append(weights_normalized)
xs.append(samples)
# Determine the number of time steps
T = len(y)
# Loop through all time steps
for t in range(T):
# Resample
# Randomly choose ancestors
ancestors = np.random.choice(samples, size=N,
replace=True, p=weights_normalized)
# Propagate
samples = stats.norm.rvs(0, 1, N) + np.power(np.cos(ancestors), 2)
# Save the new x
xs.append(samples)
# Weight
weights = stats.norm.logpdf(y[t], loc=2 * samples, scale=0.1)
# Substract maximum
weights = weights - np.max(weights)
# Normalize weights to be probabilities
weights_normalized = np.exp(weights) / np.sum(np.exp(weights))
# Save the new normalized weights
ws.append(weights_normalized)
return np.array(xs), np.array(ws)
xs_filtered, ws = bootstrap_PF(300, ys)
# +
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(np.apply_along_axis(np.sum, 1, xs_filtered * ws))
ax.plot(xs, '--')
# -
# #### Comparison of variances
# +
M = 50
N = 20
fully_adapted_estimates = np.zeros((M, T + 1))
bootstrap_estimates = np.zeros((M, T + 1))
for k in range(M):
xs_filtered = fully_adapted_PF(N, ys)
fully_adapted_estimates[k, :] = np.apply_along_axis(np.mean, 1, xs_filtered)
xs_filtered, ws = bootstrap_PF(N, ys)
bootstrap_estimates[k, :] = np.apply_along_axis(np.sum, 1, xs_filtered * ws)
fully_adapted_variances = np.apply_along_axis(np.var, 0, fully_adapted_estimates)
bootstrap_variances = np.apply_along_axis(np.var, 0, bootstrap_estimates)
fig, ax = plt.subplots(figsize=(10, 5))
ax.plot(bootstrap_variances);
ax.plot(fully_adapted_variances);
# -
# ## II.3 Likelihood estimator for the APF
#
# This is a theoretical exercise. Look in `exercises_on_paper`.
# ## II.4 Forgetting
#
# Consider the linear state space model (SSM)
# $$
# \begin{array}{rcll}
# X_t & = & 0.7 X_{t - 1} & \\
# Y_t & = & 0.5 X_t + E_t, & \qquad E_t \sim \mathcal{N}(0, 0.1)
# \end{array}
# $$
# with $X_0 \sim \mathcal{N}(0, 1)$.
# Simulate some data from the model. It is not quite clear from the exercise if $Q = 0$ already during data simulation.
# +
# Max. time steps
T = 2000
# Store the simulated measurements
xs_sim = np.zeros((T + 1,))
ys_sim = np.zeros((T,))
# Initial value
xs_sim[0] = stats.norm.rvs()
# Simulate the state and measurement process
for t in range(T):
xs_sim[t + 1] = 0.7 * xs_sim[t] + 0.1 * stats.norm.rvs()
ys_sim[t] = 0.5 * xs_sim[t + 1] + 0.1 * stats.norm.rvs()
fig, axs = plt.subplots(2, 1, figsize=(10, 10))
axs[0].plot(xs_sim);
axs[0].set_title('Simulated states');
axs[0].set_xlabel('Time');
axs[0].set_ylabel('$x_t$');
axs[1].plot(range(1, T + 1), ys_sim, 'r');
axs[1].set_title('Simulated measurements');
axs[1].set_xlabel('Time');
axs[1].set_ylabel('$y_t$');
# -
# Kalman filter, the exact solution to the filtering problem
def kalman_filter(y, A=0.7, C=0.5, Q=0.0, R=0.1, P0=1):
# Determine length of data
T = len(y)
# Filtered means and standard deviations
means_filtered = np.zeros((T + 1,))
covs_filtered = np.zeros((T + 1,))
# Initialize with covariance of prior
covs_filtered[0] = P0
# Kalman recursion
for t in range(T):
# Time update
covs_time_upd = np.power(A, 2) * covs_filtered[t] + Q
# Kalman gain
kalman_gain = C * covs_time_upd / (np.power(C, 2) * covs_time_upd + R)
# Filter updates
means_filtered[t + 1] = A * means_filtered[t] + \
kalman_gain * (y[t] - C * A * means_filtered[t])
covs_filtered[t + 1] = covs_time_upd - kalman_gain * C * covs_time_upd
return means_filtered, covs_filtered
# Bootstrap particle filter for the problem
def bootstrap_PF(y, N=100, A=0.7, C=0.5, Q=0.0, R=0.1, P0=1):
# Length of the data
T = len(y)
# Pre-allocate data storage
xs = np.zeros((N, T + 1))
ws = np.zeros((N, T + 1))
# Initialize
xs[:, 0] = stats.norm.rvs(0, P0, size=N)
ws[:, 0] = 1 / N * np.ones((N,))
for t in range(T):
# Resample
ancestors = np.random.choice(range(N), size=N,
replace=True, p=ws[:, t])
# Propagate
xs[:, t + 1] = A * xs[ancestors, t] + \
np.sqrt(Q) * stats.norm.rvs(size=N)
# Weight
# Use log weights
ws[:, t + 1] = stats.norm.logpdf(y[t], loc=C * xs[:, t + 1],
scale=np.sqrt(R))
# Find maximum and subtract from log weights
ws[:, t + 1] -= np.max(ws[:, t + 1])
# Normalize weights
ws[:, t + 1] = np.exp(ws[:, t + 1]) / np.sum(np.exp(ws[:, t + 1]))
return xs, ws
# Testing both implementations. Bootstrap PF as well as the Kalman filter follow the states rather nicely.
# +
Tmax = 100
N = 50000
means_kf, stddevs_kf = kalman_filter(ys_sim[:Tmax], Q=0.1)
xs, ws = bootstrap_PF(ys_sim[:Tmax], N=N, Q=0.1)
means_bpf = np.sum(xs * ws, axis=0)
fig, ax = plt.subplots()
ax.plot(xs_sim[:Tmax], 'ok')
ax.plot(means_bpf, 'o-')
ax.plot(means_kf, 'x-')
ax.set_xlabel('Time')
ax.set_title("$N = {}$".format(N))
ax.legend(['Simulated state', 'BPF', 'Kalman']);
# -
# If however no noise in the model is assumed, then the state recovery works a lot worse.
# +
Tmax = 100
N = 50000
means_kf, stddevs_kf = kalman_filter(ys_sim[:Tmax], Q=0.0)
xs, ws = bootstrap_PF(ys_sim[:Tmax], N=N, Q=0.0)
means_bpf = np.sum(xs * ws, axis=0)
fig, ax = plt.subplots()
ax.plot(xs_sim[:Tmax], 'ok')
ax.plot(means_bpf, 'o-')
ax.plot(means_kf, 'x-')
ax.set_xlabel('Time')
ax.set_title("$N = {}$".format(N))
ax.legend(['Simulated state', 'BPF', 'Kalman']);
# -
# Looking at the mean-squared-error for the test function $\phi(x_t) = x_t$
# +
M = 100
Tmax = 50
mses = np.zeros((Tmax + 1,))
# Get the exact solution
means_kf, stddevs_kf = kalman_filter(ys_sim[:Tmax], Q=0.1)
# Iterate and repeatedly calculate approximation
for i in range(M):
xs, ws = bootstrap_PF(ys_sim[:Tmax], N=100, Q=0.1)
means_bpf = np.sum(xs * ws, axis=0)
# Add to mean squared errors
mses += np.power(means_bpf - means_kf, 2.0)
# Divide by number of repetitions
mses /= M
fig, ax = plt.subplots()
ax.plot(mses, 'o-')
ax.set_xlabel('Time')
ax.set_ylabel('MSE');
# -
# The errors decrease to zero. Not sure what I am supposed to get in this exercise.
| solutions/code/Python/fheld/exII.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# # Sweep waveguide width
#
# Compute the effective index of different modes for different waveguide widths
#
# we compute mode properties (neff, aeff ...) as a function of the waveguide width
#
# we have to make sure that the simulation region is larger than the waveguide
#
#
# Simulation of mode hybridisation in 220nm thick fully-etched SOI ridge
# waveguides.
#
# Results look the same as those found in [<NAME> and <NAME>, "Mode
# hybridization and conversion in silicon-on-insulator nanowires with angled
# sidewalls," Opt. Express 23, 32452-32464
# (2015)](https://www.osapublishing.org/oe/abstract.cfm?uri=oe-23-25-32452).
# _________________________________
#
# clad_thickness
# width
# <---------->
# ___________ _ _ _ _ _ _
# | |
# _____| |____ |
# wg_heigth
# slab_thickness |
# _______________________ _ _ _ _ __
#
# sub_thickness
# _________________________________
# <------------------------------->
# sub_width
#
# +
import numpy as np
import matplotlib.pyplot as plt
import modes as ms
import opticalmaterialspy as mat
#widths = np.arange(0.3, 2.0, 0.02)
widths = np.arange(0.3, 2.0, 0.2)
wgs = [ms.waveguide(width=width) for width in widths]
wgs[0]
# -
s = ms.sweep_waveguide?
# ## 1550 nm strip waveguides
# Here are some waveguide simulations for C (1550nm) and O (1310) band, where we sweep the waveguide width and compute the effective modes supported by the waveguide.
#
# TE mode (transverse-electrical) means that the light is mainly polarized in the horizontal direction (the main electric field component is Ex) while TM transverse magnetic field modes have the strongest field component (Ey). We can see why for 1550nm a typical waveguide width is 0.5nm, as they only support one TE mode.
# +
# ms.mode_solver_full?
# -
ms.mode_solver_full(width=0.5, plot=True, fields_to_write=('Ex', 'Ey'))
# As waveguides become wider they start supporting more than one single mode. For example, a 2um wide waveguide supports 4 different TE modes (TE0: 1 lobe, TE1: 2 lobes, TE2: 3 lobes, TE3:4 lobes)
ms.mode_solver_full(width=2.0, plot=True, n_modes=4, fields_to_write=('Ex'))
s = ms.sweep_waveguide(wgs, widths, legend=['TE0', 'TM0', 'TE1', 'TM1'], overwrite=False)
# we can create a waveguide compact model that captures the neff variation with width for the fundamental TE mode
s.keys()
n0 = [n[0] for n in s['n_effs']]
plt.plot(widths, n0, '.')
plt.xlabel('width (um)')
plt.ylabel('neff')
p = np.polyfit(widths, n0, 6)
n0f = np.polyval(p, widths)
plt.plot(widths, n0, '.')
plt.plot(widths, n0f, '.')
plt.xlabel('width (um)')
plt.ylabel('neff')
p
# ## 1310 nm strip waveguides
wgs1310 = [ms.waveguide(width=width, wavelength=1.31) for width in widths]
s = ms.sweep_waveguide(wgs1310, widths)
widths = np.arange(0.3, 1.0, 0.1)
overwrite = False
wgs = [ms.waveguide(width=width) for width in widths]
r2 = ms.sweep_waveguide(
wgs, widths, n_modes=2, fraction_mode_list=[1, 2], overwrite=overwrite,
)
# ## Rib waveguide sweep
#
# For 90nm slab
widths = np.arange(0.3, 1.0, 0.1)
overwrite = False
wgs = [ms.waveguide(width=width, slab_thickness=90e-3) for width in widths]
r2 = ms.sweep_waveguide(
wgs, widths, n_modes=3, fraction_mode_list=[1, 2, 3], overwrite=overwrite,
)
| docs/notebooks/20_sweep_width.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: tagifai
# language: python
# name: tagifai
# ---
# + [markdown] id="LPZmAUydQIC9"
# <div align="center">
# <h1><img width="30" src="https://madewithml.com/static/images/rounded_logo.png"> <a href="https://madewithml.com/">Made With ML</a></h1>
# Applied ML · MLOps · Production
# <br>
# Join 30K+ developers in learning how to responsibly <a href="https://madewithml.com/about/">deliver value</a> with ML.
# <br>
# </div>
#
# <br>
#
# <div align="center">
# <a target="_blank" href="https://newsletter.madewithml.com"><img src="https://img.shields.io/badge/Subscribe-30K-brightgreen"></a>
# <a target="_blank" href="https://github.com/GokuMohandas/MadeWithML"><img src="https://img.shields.io/github/stars/GokuMohandas/MadeWithML.svg?style=social&label=Star"></a>
# <a target="_blank" href="https://www.linkedin.com/in/goku"><img src="https://img.shields.io/badge/style--5eba00.svg?label=LinkedIn&logo=linkedin&style=social"></a>
# <a target="_blank" href="https://twitter.com/GokuMohandas"><img src="https://img.shields.io/twitter/follow/GokuMohandas.svg?label=Follow&style=social"></a>
# <br>
# 🔥 Among the <a href="https://github.com/topics/mlops" target="_blank">top MLOps</a> repositories on GitHub
# </div>
#
# <br>
# <hr>
# + [markdown] id="L7--e8qjzvte"
# # Optimize (GPU)
# + [markdown] id="IRyA7luizvtf"
# Use this notebooks to run hyperparameter optimization on Google Colab and utilize it's free GPUs.
# + [markdown] id="rOAOZ5NJzvtl"
# ## Clone repository
# + colab={"base_uri": "https://localhost:8080/"} id="dC_KGdE6zvtl" outputId="8de114ed-97c2-42c5-a41b-b97c8b4bee0f"
# Load repository
# !git clone https://github.com/GokuMohandas/MLOps.git mlops
# + colab={"base_uri": "https://localhost:8080/"} id="aiKzsC9kzvtn" outputId="766de28c-38c4-4097-81ea-4d62f45959c7"
# Files
% cd mlops
# !ls
# + [markdown] id="LnZVQRcZzvtp"
# ## Setup
# + id="lKp6B4M478m_" language="bash"
# !pip install --upgrade pip
# !python -m pip install -e ".[dev]" --no-cache-dir
# + [markdown] id="u90bt1s0SuvM"
# # Download data
# + [markdown] id="4FPUxmU_S7tW"
# We're going to download data directly from GitHub since our blob stores are local. But you can easily load the correct data versions from your cloud blob store using the *.json.dvc pointer files in the [data directory](https://github.com/GokuMohandas/MLOps/tree/main/data).
# + id="O4oQwat9Syf7"
from app import cli
# + colab={"base_uri": "https://localhost:8080/"} id="0PzQcqIuKLkU" outputId="f4b63d56-a8e9-4455-fbe1-f4c315c6bd64"
# Download data
cli.download_data()
# + colab={"base_uri": "https://localhost:8080/"} id="ZsPyGrZYIsmA" outputId="104bc988-7240-4ef5-b81d-e9fdae1cc3cb"
# Check if data downloaded
# !ls data
# -
# # Compute features
# Download data
cli.compute_features()
# Computed features
# !ls data
# + [markdown] id="wzxXb5mjzvts"
# ## Optimize
# + [markdown] id="vn0kS5b8TOL8"
# Now we're going to perform hyperparameter optimization using the objective and parameter distributions defined in the [main script](https://github.com/GokuMohandas/MLOps/blob/main/tagifai/main.py). The best parameters will be written to [config/params.json](https://raw.githubusercontent.com/GokuMohandas/MLOps/main/config/params.json) which will be used to train the best model below.
# + colab={"base_uri": "https://localhost:8080/"} id="WNGWNx_uSvaU" outputId="c7994498-063c-4d3b-d196-84286ce7f0c7"
# Optimize
cli.optimize(num_trials=100)
# + [markdown] id="WBJbUfkTTs5j"
# # Train
# + [markdown] id="jF-9xX7DTweB"
# Once we're identified the best hyperparameters, we're ready to train our best model and save the corresponding artifacts (label encoder, tokenizer, etc.)
# + colab={"base_uri": "https://localhost:8080/"} id="eOT46qHmD1ZR" outputId="df8f66dc-a93d-4cf5-9f6b-66aa7408f6d3"
# Train best model
cli.train_model()
# + [markdown] id="QCNVQICxZU_k"
# # Change metadata
# + [markdown] id="r-bb36cxZWee"
# In order to transfer our trained model and it's artifacts to our local model registry, we should change the metadata to match.
# + id="E0LsygkBZdVb"
from pathlib import Path
from config import config
import yaml
# + id="zuWV3nE-Zdis"
def change_artifact_metadata(fp):
with open(fp) as f:
metadata = yaml.load(f)
for key in ["artifact_location", "artifact_uri"]:
if key in metadata:
metadata[key] = metadata[key].replace(
str(config.MODEL_REGISTRY), model_registry)
with open(fp, "w") as f:
yaml.dump(metadata, f)
# + id="0_qGvxDYZjrQ"
# Change this as necessary
model_registry = "/Users/goku/Documents/madewithml/applied-ml/stores/model"
# + id="er8i_5FOZeTQ"
# Change metadata in all meta.yaml files
experiment_dir = Path(config.MODEL_REGISTRY, "1")
for fp in list(Path(experiment_dir).glob("**/meta.yaml")):
change_artifact_metadata(fp=fp)
# + [markdown] id="w8Wm1xPl0HyF"
# ## Download
# + [markdown] id="uJrowj_lzvtz"
# Download and transfer the trained model's files to your local model registry. If you existing runs, just transfer that run's directory.
# + id="lEkEtbaX0LbU"
from google.colab import files
# + colab={"base_uri": "https://localhost:8080/", "height": 714} id="LJeRbLxh0NxV" outputId="8b1cafed-48fd-4eeb-c04f-538119da8fdf"
# Download
# !zip -r model.zip model
# !zip -r run.zip stores/model/1
files.download("run.zip")
# + id="vbL3YuPSbbIU"
| notebooks/optimize.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# + [markdown] lang="fr"
# <center>
# <img src="https://github.com/Yorko/mlcourse.ai/blob/master/img/ods_stickers.jpg?raw=true" />
#
# # [mlcourse.ai](https://mlcourse.ai) – Open Machine Learning Course
#
# Auteur: [<NAME>](https://yorko.github.io). Traduit et édité par [<NAME>](https://www.linkedin.com/in/christinabutsko/), [<NAME>](https://www.linkedin.com/in/yuanyuanpao/), [<NAME>](https://www.linkedin.com/in/anastasiamanokhina), <NAME>, [<NAME>](https://www.linkedin.com/in/datamove/) et [<NAME>](https://github.com/oussou-dev). Ce matériel est soumis aux termes et conditions de la licence [Creative Commons CC BY-NC-SA 4.0] (https://creativecommons.org/licenses/by-nc-sa/4.0/). L'utilisation gratuite est autorisée à des fins non commerciales.
# + [markdown] lang="fr"
# # <center> Thème 1. Analyse exploratoire de données avec la librairie Pandas
#
# <img src="https://github.com/Yorko/mlcourse.ai/blob/master/img/pandas.jpg?raw=true" width=50% />
# + [markdown] toc=true
# <h1>Sommaire de l'article<span class="tocSkip"></span></h1>
# <div class="toc"><ul class="toc-item"><li><span><a href="#1.-Mise-en-pratique-des-principales-méthodes-de-Pandas" data-toc-modified-id="1.-Mise-en-pratique-des-principales-méthodes-de-Pandas-1">1. Mise en pratique des principales méthodes de Pandas</a></span><ul class="toc-item"><li><span><a href="#Tri" data-toc-modified-id="Tri-1.1">Tri</a></span></li><li><span><a href="#Indexation-et-récupération-de-données" data-toc-modified-id="Indexation-et-récupération-de-données-1.2">Indexation et récupération de données</a></span></li><li><span><a href="#Application-de-fonctions-à-des-cellules,-des-colonnes-et-des-lignes" data-toc-modified-id="Application-de-fonctions-à-des-cellules,-des-colonnes-et-des-lignes-1.3">Application de fonctions à des cellules, des colonnes et des lignes</a></span></li><li><span><a href="#Agrégation-de-données" data-toc-modified-id="Agrégation-de-données-1.4">Agrégation de données</a></span></li><li><span><a href="#Tableaux-récapitulatifs" data-toc-modified-id="Tableaux-récapitulatifs-1.5">Tableaux récapitulatifs</a></span></li><li><span><a href="#Opérations-de-transformations-d'un-DataFrame" data-toc-modified-id="Opérations-de-transformations-d'un-DataFrame-1.6">Opérations de transformations d'un DataFrame</a></span></li></ul></li><li><span><a href="#2.-Prévision-du-churn-(taux-d'attrition)" data-toc-modified-id="2.-Prévision-du-churn-(taux-d'attrition)-2">2. Prévision du churn (taux d'attrition)</a></span></li><li><span><a href="#3.-Mission-pour-s'exercer" data-toc-modified-id="3.-Mission-pour-s'exercer-3">3. Mission pour s'exercer</a></span></li><li><span><a href="#4.-Ressources-utiles" data-toc-modified-id="4.-Ressources-utiles-4">4. Ressources utiles</a></span></li></ul></div>
# + [markdown] lang="fr"
# ## 1. Mise en pratique des principales méthodes de Pandas
# Bien ... Il existe des dizaines de tutoriels intéressants sur la librairie Pandas et l'analyse visuelle des données. Si vous êtes déjà familiarisé avec ces sujets, vous pouvez passer au 3ème article de la série, dans lequel nous abordons le machine learning (apprentissage automatique).
#
# **[Pandas](http://pandas.pydata.org)** est une bibliothèque Python qui fournit des moyens étendus pour l’analyse de données. Les Data scientistes travaillent souvent avec des données stockées dans des formats sous forme de table de données telles que `.csv`,` .tsv` ou `.xlsx`. Pandas est très pratique pour charger, traiter et analyser ces données tabulaires à l’aide de requêtes quasi-similaires aux reques de type SQL. En complément de `Matplotlib` et` Seaborn`, `Pandas` offre un large éventail d'opportunités d'analyse visuelle des données tabulaires.
#
# Les pricipales structures de données dans `Pandas` sont implémentées avec les classes **Series** et **DataFrame**. Le premier est un tableau unidimensionnel indexé d'un type de données fixe. Le second est une structure de données bidimensionnelle - une table - dans laquelle chaque colonne contient des données du même type. Vous pouvez le voir comme un dictionnaire de plusieurs `Series`. Les `DataFrames` sont parfaits pour représenter des données réelles : les lignes correspondent aux instances (exemples, observations, individus etc.) et les colonnes correspondent aux caractéristiques de ces instances (variables).
# +
import numpy as np
import pandas as pd
pd.set_option("display.precision", 2)
# + [markdown] lang="fr"
# Nous allons tester les principales méthodes en analysant un jeu de données ou [dataset](https://bigml.com/user/francisco/gallery/dataset/5163ad540c0b5e5b22000383) sur le taux de désabonnement des clients d'opérateurs téléphoniques. Lisons les données (en utilisant la méthode `read_csv`), et jetons un coup d’œil aux 5 premières lignes en utilisant la méthode` head`:
# +
url = (
"https://raw.githubusercontent.com/Yorko/mlcourse.ai/master/data/telecom_churn.csv"
)
df = pd.read_csv(url)
df.head()
# + [markdown] lang="fr"
# <details>
# <summary>Affichage des DataFrames dans Jupyter notebooks</summary>
# <p>
# Dans les notebooks Jupyter, les DataFrames sont affichés comme ces jolies tables vues ci-dessus alors que `print(df.head())` est moins bien formaté.
# Par défaut, Pandas affiche 20 colonnes et 60 lignes, donc, si votre DataFrame est plus grand, utilisez la fonction `set_option` comme dans l'exemple ci-dessous :
#
# <br>
#
# ```
# pd.set_option('display.max_columns', 100)
# pd.set_option('display.max_rows', 100)
# ```
# </p>
# </details>
#
# Rappelez-vous que chaque ligne correspond à un client, à une **instance** et les colonnes sont les **caractéristiques** de cette instance.
# + [markdown] lang="fr"
# Examinons la dimensionnalité des données, les noms des caractéristiques et les types de caractéristiques.
# -
print(df.shape)
# + [markdown] lang="fr"
# La sortie indique que la table contient 3333 lignes et 20 colonnes.
#
# Essayons maintenant d’afficher les noms de colonnes en utilisant `columns`:
# -
print(df.columns)
# + [markdown] lang="fr"
# Nous pouvons utiliser la méthode `info()` pour générer des informations générales sur le Dataframe :
# -
print(df.info())
# + [markdown] lang="fr"
# `bool`,` int64`, `float64` et` object` sont les types de données de nos caractéristiques. Nous voyons qu'une caractéristique est logique (`bool`), 3 caractéristiques sont de type ` objet`, et 16 caractéristiques sont numériques. Avec cette même méthode, nous pouvons facilement voir s’il manque des valeurs. Ici, il n'y en a pas car chaque colonne contient 3333 observations, le même nombre de lignes que nous avons vu auparavant avec `shape`.
#
# Nous pouvons **changer le type de colonne** avec la méthode `astype`. Appliquons cette méthode à la caractéristique `Churn` pour la convertir en` int64`:
# -
df["Churn"] = df["Churn"].astype("int64")
# + [markdown] lang="fr"
# La méthode `describe` affiche les caractéristiques statistiques de base de chaque caractéristique numérique (type ` int64` et `float64`): nombre de valeurs non manquantes, moyenne, écart-type, amplitude, médiane, (1er : 0,25) et (3ème : 0,75) quartiles.
# -
df.describe()
# + [markdown] lang="fr"
# Afin de voir les statistiques des caractéristiques non numériques, il faut indiquer explicitement ces types de données dans le paramètre `include`.
# -
df.describe(include=["object", "bool"])
# + [markdown] lang="fr"
# Pour les caractéristiques catégorielles (type `objet`) et booléennes (type` bool`), nous pouvons utiliser la méthode `value_counts`. Jetons un coup d'oeil à la distribution de `Churn`:
# -
df["Churn"].value_counts()
# + [markdown] lang="fr"
# 2850 utilisateurs sur 3333 sont *fidèles* clients; leur valeur `Churn` est 0. Pour calculer les pourcentages, passez ` normalize = True` à la fonction `value_counts`.
# -
df["Churn"].value_counts(normalize=True)
# + [markdown] lang="fr"
# ### Tri
#
# Un `DataFrame` peut être trié selon la valeur de l’une des variables (colonnes). Par exemple, nous pouvons trier par *Total day charge*
# (utilisez `ascending = False` pour trier par ordre décroissant):
# -
df.sort_values(by="Total day charge", ascending=False).head()
# + [markdown] lang="fr"
# Nous pouvons également trier sur plusieurs colonnes:
# -
df.sort_values(by=["Churn", "Total day charge"], ascending=[True, False]).head()
# + [markdown] lang="fr"
# ### Indexation et récupération de données
#
# Un `DataFrame` peut être indexé de différentes manières.
#
# Pour obtenir une seule colonne, vous pouvez saisir : `DataFrame['NomDeColonne'] `. Que nous utilisons pour répondre à une question à propos de cette colonne uniquement: **quelle est la proportion d'utilisateurs qui se sont désabonnés dans notre base de données?**
# -
df["Churn"].mean()
# + [markdown] lang="fr"
# 14,5% est en fait assez mauvais pour une entreprise; un tel taux de désabonnement peut entraîner la faillite de l'entreprise.
#
# **L'indexation booléenne** avec une colonne est également très pratique. La syntaxe est `df[P(df['Nom']]]`, où `P` est une condition logique vérifiée pour chaque élément de la colonne ` NomDeColonne`. Le résultat de cette indexation est le `DataFrame` composé uniquement de lignes satisfaisant la condition ` P` de la colonne `NomDeColonne`.
#
# Exemple d'utlisation pour répondre à la question:
#
# **Quelles sont les valeurs moyennes des caractéristiques numériques pour les utilisateurs désabonnés, c'est-à-dire qui ont un Churn égal à 1 ?**
# -
df[df["Churn"] == 1].mean()
# + [markdown] lang="fr"
# **Combien de temps (en moyenne) les utilisateurs désabonnés passent-ils au téléphone pendant la journée?**
# -
df[df["Churn"] == 1]["Total day minutes"].mean()
# + [markdown] lang="fr"
# **Quelle est la durée maximale des appels internationaux parmi les clients fidèles (`Churn == 0`) n'ayant pas de forfait international?**
# -
df[(df["Churn"] == 0) & (df["International plan"] == "No")]["Total intl minutes"].max()
# + [markdown] lang="fr"
# Les DataFrames peuvent être indexés par nom de colonne (étiquette) ou nom de ligne (index) ou par le numéro de série (indice) d'une ligne. La méthode `loc` est utilisée pour **l'indexation par nom**, tandis que` iloc() `est utilisée pour **l'indexation par numéro**.
#
# Dans le premier cas ci-dessous, nous *"récupérons les valeurs des lignes d'index de 0 à 5 (inclus) et des colonnes étiquetées de State à Area code (inclus)"*.
# Dans le second cas, nous *"récupérons les valeurs des cinq premières lignes des trois premières colonnes"* (comme dans un slicing avec Python : la valeur maximale n'est pas incluse).
# -
df.loc[0:5, "State":"Area code"]
df.iloc[0:5, 0:3]
# + [markdown] lang="fr"
# Si nous avons besoin de la première ou de la dernière ligne du dataframe, nous pouvons utiliser la syntaxe : `df[:1]` ou `df[-1:]`:
# -
df[-1:]
# + [markdown] lang="fr"
# ### Application de fonctions à des cellules, des colonnes et des lignes
#
# **Pour appliquer des fonctions à chaque colonne, utilisez `apply ()`:**
# -
df.apply(np.max)
# + [markdown] lang="fr"
# La méthode `apply` peut également être utilisée pour appliquer une fonction à chaque ligne. Pour ce faire, spécifiez `axis = 1`. Les fonctions Lambda sont très pratiques dans de tels scénarios. Par exemple, si nous devons sélectionner tous les _state_ commençant par 'W', nous pouvons le faire comme suit:
# -
df[df["State"].apply(lambda state: state[0] == "W", axis=1)].head()
# + [markdown] lang="fr"
# La méthode `map` peut être utilisée pour **remplacer des valeurs dans une colonne** en transmettant un dictionnaire de la forme` {ancienne_valeur: nouvelle_valeur} `comme argument:
# -
d = {"No": False, "Yes": True}
df["International plan"] = df["International plan"].map(d)
df.head()
# + [markdown] lang="fr"
# Presque la même chose peut être faite avec la méthode `replace`.
#
# <details>
# <summary>Différence dans le traitement des valeurs absentes dans le dictionnaire de mappage</summary>
# <p>
# Il y a une petite différence.
# La méthode `replace` ne fera rien avec des valeurs qui ne se trouvent pas dans le dictionnaire de mappage,
# alors que `map` les changera en `NaN`).
# <br>
#
# ```python
# a_series = pd.Series(['a', 'b', 'c'])
# a_series.replace({'a': 1, 'b': 1}) # 1, 2, c
# a_series.map({'a': 1, 'b': 2}) # 1, 2, NaN
# ```
# </p>
# </details>
# -
df = df.replace({"Voice mail plan": d})
df.head()
# + [markdown] lang="fr"
# ### Agrégation de données
#
# En général, le regroupement des données dans Pandas fonctionne comme suit :
# -
#
# ```python
# df.groupby(by=grouping_columns)[columns_to_show].function()
# ```
# + [markdown] lang="fr"
# 1. Premièrement, la méthode `groupby` divise les colonnes`grouping_columns` par leurs valeurs. Ils deviennent un nouvel index dans le dataframe qui en résulte.
# 2. Ensuite, les colonnes choisies sont sélectionnées (`columns_to_show`). Si `columns_to_show` n'est pas inclus, toutes les clauses non groupby seront incluses.
# 3. Enfin, une ou plusieurs fonctions sont appliquées aux groupes obtenus par colonnes sélectionnées.
#
# Voici un exemple où nous regroupons les données en fonction des valeurs de la variable `Churn` et affichons les statistiques de trois colonnes dans chaque groupe :
# +
columns_to_show = ["Total day minutes", "Total eve minutes", "Total night minutes"]
df.groupby(["Churn"])[columns_to_show].describe(percentiles=[])
# + [markdown] lang="fr"
# Faisons la même chose, mais légèrement différemment, en passant une liste de fonctions à `agg ()`:
# +
columns_to_show = ["Total day minutes", "Total eve minutes", "Total night minutes"]
df.groupby(["Churn"])[columns_to_show].agg([np.mean, np.std, np.min, np.max])
# + [markdown] lang="fr"
# ### Tableaux récapitulatifs
#
# Supposons que nous voulions voir comment les observations de notre ensemble de données sont réparties dans le contexte de deux variables - `Churn` et`International plan`. Pour ce faire, nous pouvons construire un **tableau de contingence** en utilisant la méthode `crosstab`:
# -
pd.crosstab(df["Churn"], df["International plan"])
pd.crosstab(df["Churn"], df["Voice mail plan"], normalize=True)
# + [markdown] lang="fr"
# Nous pouvons constater que la plupart des utilisateurs sont fidèles et n'utilisent pas de services supplémentaires (International Plan/Voice mail).
#
# Cela ressemblera aux **tableaux croisés dynamiques** pour ceux qui connaissent Excel. Et, bien sûr, les tableaux croisés dynamiques sont implémentés dans Pandas: la méthode `pivot_table` prend les paramètres suivants:
#
# *`values` - une liste de variables pour calculer des statistiques,
# * `index` - une liste de variables pour regrouper les données,
# * `aggfunc` - quelles statistiques nous devons calculer pour les groupes, ex. somme, moyenne, maximum, minimum ou autre chose.
#
# Examinons le nombre moyen d'appels de jour, de soir et de nuit par code régional (area code):
# -
df.pivot_table(
["Total day calls", "Total eve calls", "Total night calls"],
["Area code"],
aggfunc="mean",
)
# + [markdown] lang="fr"
# ### Opérations de transformations d'un DataFrame
#
# Comme beaucoup d'autres choses avec Pandas, l'ajout de colonnes à un DataFrame est réalisable de plusieurs manières.
#
# Par exemple, si nous voulons calculer le nombre total d'appels pour tous les utilisateurs, créons la série `total_calls` et collons-la dans le DataFrame:
# -
total_calls = (
df["Total day calls"]
+ df["Total eve calls"]
+ df["Total night calls"]
+ df["Total intl calls"]
)
df.insert(loc=len(df.columns), column="Total calls", value=total_calls)
# le paramètre loc est le nombre de colonnes après lequel l'objet Série doit être inséré
# nous l'initialisons à len(df.columns) pour le coller à la toute fin du dataframe
df.head()
# + [markdown] lang="fr"
# Il est possible d’ajouter une colonne plus facilement sans créer d’instance Series intermédiaire:
# -
df["Total charge"] = (
df["Total day charge"]
+ df["Total eve charge"]
+ df["Total night charge"]
+ df["Total intl charge"]
)
df.head()
# + [markdown] lang="fr"
# Pour supprimer des colonnes ou des lignes, utilisez la méthode `drop`, en passant les index requis et le paramètre` axis` (`1` si vous supprimez des colonnes et rien ou` 0` si vous supprimez des lignes). L'argument `inplace` indique s'il faut modifier le DataFrame d'origine. Avec `inplace = False`, la méthode` drop` ne modifie pas le DataFrame existant et en renvoie un nouveau avec des lignes ou des colonnes supprimées. Avec `inplace = True`, il modifie le DataFrame.
# -
# supprimer les colonnes qui viennent d'être créées
df.drop(["Total charge", "Total calls"], axis=1, inplace=True)
# et voici comment supprimer des lignes
df.drop([1, 2]).head()
# + [markdown] lang="fr"
# ## 2. Prévision du churn (taux d'attrition)
#
# Voyons comment le taux de désabonnement est lié à la caractéristique ou variable *International plan*. Pour ce faire, nous utiliserons un tableau de contingence `` crosstab`` et également une analyse visuelle avec `Seaborn` (l'analyse visuelle sera toutefois traitée plus en détail dans le prochain article).
# -
pd.crosstab(df["Churn"], df["International plan"], margins=True)
# +
# quelques imports pour mettre en place le cadre du graphique
import matplotlib.pyplot as plt
# # !pip install seaborn (pour installer la librairie seaborn via le notebook)
import seaborn as sns
# import de paramètres pour améliorer le rendu visuel
sns.set()
# Les graphiques au format Retina sont plus nets et plus lisibles
# %config InlineBackend.figure_format = 'retina'
# -
sns.countplot(x="International plan", hue="Churn", data=df);
# + [markdown] lang="fr"
# Nous voyons qu'avec *International Plan*, le taux de désabonnement est beaucoup plus élevé, ce qui est une observation intéressante! Peut-être des dépenses importantes et mal contrôlées avec des appels internationaux sont-elles très sujettes aux conflits et suscitent l’insatisfaction des clients de l’opérateur de télécommunications.
#
# Voyons ensuite une autre fonctionnalité importante - *Customer service calls*. Faisons également un tableau de synthèse et une image.
# -
pd.crosstab(df["Churn"], df["Customer service calls"], margins=True)
sns.countplot(x="Customer service calls", hue="Churn", data=df);
# + [markdown] lang="fr"
# Bien que ce ne soit pas évident dans le tableau récapitulatif, il ressort clairement du graphique ci-dessus que le taux de résiliation augmente fortement à partir de 4 appels de service après-vente.
#
# Ajoutons maintenant une variable binaire à notre DataFrame - `Customer service calls > 3` (Appels du service client> 3). Et encore une fois, voyons comment cela se rapporte au désabonnement.
# +
df["Many_service_calls"] = (df["Customer service calls"] > 3).astype("int")
pd.crosstab(df["Many_service_calls"], df["Churn"], margins=True)
# -
sns.countplot(x="Many_service_calls", hue="Churn", data=df);
# + [markdown] lang="fr"
# Construisons une autre table de contingence qui relie *Churn* à la fois à *International plan* et à la variable nouvellement créée *Many_service_calls*.
# -
pd.crosstab(df["Many_service_calls"] & df["International plan"], df["Churn"])
# + [markdown] lang="fr"
# Par conséquent, si un nombre d'appels vers le centre de services est supérieur à 3 et que le *International Plan* est ajouté (et en prédisant Churn=0 sinon), on peut s’attendre à une précision de 85,8%. Ce nombre, 85,8%, que nous avons obtenu grâce à ce raisonnement très simple constitue un bon point de départ (*référence*) pour les autres modèles d’apprentissage automatique que nous allons construire.
#
# Au cours de ce cours, rappelez-vous qu'avant l'avènement de l'apprentissage automatique, le processus d'analyse des données ressemblait à ce que nous venons de réaliser. Récapitulatif :
#
# - La part des clients fidèles dans l'ensemble de données est de 85,5%. Le modèle le plus "simple" qui prédit toujours un "client fidèle" sur de telles données devinera juste dans environ 85,5% des cas. C'est-à-dire que la proportion de réponses correctes (*précision*) des modèles suivants ne devrait pas être inférieure à ce nombre et qu'elle devrait être nettement supérieure;
# - A l’aide d’une simple prévision pouvant être exprimée par la formule suivante: `International plan = True & Customer Service calls > 3 => Churn = 1, else Churn = 0`, on peut s’attendre à un taux de prédiction de 85,8%, qui est juste au-dessus de 85,5%. Ensuite, nous parlerons des arbres de décision et découvrirons comment trouver de telles règles **automatiquement** uniquement sur la base des données d’entrée;
# - Nous avons obtenu ces deux bases sans appliquer l’apprentissage automatique et elles serviront de point de départ pour nos modèles ultérieurs. S'il s'avère qu'avec un effort énorme, nous n'augmentons la précision que de 0,5%, alors nous avons peut-être commis une erreur, et il suffit de nous en tenir à un simple modèle "if-else" avec deux conditions;
# - Avant de former des modèles complexes, il est recommandé de mélanger un peu les données, de tracer des graphiques et de vérifier des hypothèses simples. De plus, dans les applications métier de l'apprentissage automatique, on commence généralement par des solutions simples, pour ensuite expérimenter des solutions plus complexes.
#
# ## 3. Mission pour s'exercer
# Pour vous entraîner avec la librairie Pandas et l’EDA (Analyse Exploratoire de données), vous pouvez remplir [cette mission](https://www.kaggle.com/kashnitsky/a1-demo-pandas-and-uci-adult-dataset) où vous analyserez des données socio-démographiques. Il s'agit juste d'une mission pour vous exerver avec sa [solution](https://www.kaggle.com/kashnitsky/a1-demo-pandas-and-uci-adult-dataset-solution).
#
# ## 4. Ressources utiles
#
# * Le même notebook en mode interactif sur [Kaggle Kernel](https://www.kaggle.com/kashnitsky/topic-1-exploratory-data-analysis-with-pandas)
# * ["Merging DataFrames with pandas"](https://nbviewer.jupyter.org/github/Yorko/mlcourse.ai/blob/master/jupyter_english/tutorials/merging_dataframes_tutorial_max_palko.ipynb) - a tutorial by <NAME> within mlcourse.ai (full list of tutorials is [here](https://mlcourse.ai/tutorials))
# * ["Handle different dataset with dask and trying a little dask ML"](https://nbviewer.jupyter.org/github/Yorko/mlcourse.ai/blob/master/jupyter_english/tutorials/dask_objects_and_little_dask_ml_tutorial_iknyazeva.ipynb) - a tutorial by <NAME> within mlcourse.ai
# * Main course [site](https://mlcourse.ai), [course repo](https://github.com/Yorko/mlcourse.ai), and YouTube [channel](https://www.youtube.com/watch?v=QKTuw4PNOsU&list=PLVlY_7IJCMJeRfZ68eVfEcu-UcN9BbwiX)
# * Official Pandas [documentation](http://pandas.pydata.org/pandas-docs/stable/index.html)
# * Course materials as a [Kaggle Dataset](https://www.kaggle.com/kashnitsky/mlcourse)
# * Medium ["story"](https://medium.com/open-machine-learning-course/open-machine-learning-course-topic-1-exploratory-data-analysis-with-pandas-de57880f1a68) based on this notebook
# * If you read Russian: an [article](https://habrahabr.ru/company/ods/blog/322626/) on Habr.com with ~ the same material. And a [lecture](https://youtu.be/dEFxoyJhm3Y) on YouTube
# * [10 minutes to pandas](http://pandas.pydata.org/pandas-docs/stable/10min.html)
# * [Pandas cheatsheet PDF](https://github.com/pandas-dev/pandas/blob/master/doc/cheatsheet/Pandas_Cheat_Sheet.pdf)
# * GitHub repos: [Pandas exercises](https://github.com/guipsamora/pandas_exercises/) and ["Effective Pandas"](https://github.com/TomAugspurger/effective-pandas)
# * [scipy-lectures.org](http://www.scipy-lectures.org/index.html) — tutorials on pandas, numpy, matplotlib and scikit-learn
| jupyter_french/topic01_pandas_data_analysis/topic1_pandas_data_analysis-fr_def.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # MLP applied to quantum metrology
# Here I will show how we used MLP to infer a probability distribution of the laser detunning
"Code imports"
import tensorflow as tf
from tensorflow import keras
from tensorflow.keras.callbacks import EarlyStopping
from tensorflow.keras.callbacks import ModelCheckpoint
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from numpy import loadtxt
from numpy import savetxt
from scipy.sparse.linalg import expm, expm_multiply
from scipy import sparse
# +
"Now we import and preprocess the registers"
"We import the registers"
n_Delta=100
Delta_vec=np.linspace(1,5,n_Delta)
Delta_class=list(range(n_Delta))
diccionario_clases=dict(zip(Delta_vec,Delta_class))
out_name = "C:/Users/Manuel/Desktop/universidad/beca_gefes_2021/proyecto/archivos_clicks/n_Delta_100_n_clicks_100/clicks_deltas_Omega_3.csv"
Deltas_and_clicks=loadtxt(out_name,delimiter=",")
y_full=Deltas_and_clicks[:,0]
y_full_class=np.vectorize(diccionario_clases.get)(y_full)
X_full=Deltas_and_clicks[:,1:]
"We are only interested in the time difference between photons"
X_full[:,1:]=X_full[:,1:]-X_full[:,:-1]
ratio=0.8
limit=int(ratio*len(y_full))
y_train,y_valid=y_full_class[:limit],y_full_class[limit:]
"We reescale the photon times so that the inputs are between 0 and 1 as it is in that range where we have"
"the most expresivity of the activation functions"
maxX=np.amax(X_full)
X_train,X_valid=X_full[:limit,:]/maxX,X_full[limit:,:]/maxX
"Lastly we shuffle the training data"
indices = tf.range(start=0, limit=tf.shape(X_train)[0], dtype=tf.int32)
shuffled_indices = tf.random.shuffle(indices)
X_train = tf.gather(X_train, shuffled_indices)
y_train = tf.gather(y_train, shuffled_indices)
# +
"We train our first neural network, with early stopping so that we retrive the best weights"
es = EarlyStopping(monitor='val_loss',
mode='min', verbose=1, patience=10, min_delta=0.01,
restore_best_weights=True)
model=keras.models.Sequential([
keras.layers.Dense(300,activation="relu"),
keras.layers.Dense(100,activation="relu"),
keras.layers.Dense(100,activation="relu"),
keras.layers.Dense(n_Delta,activation="softmax")
# El activador es softmax para que salga una distribucion normalizada
])
model.compile(loss="sparse_categorical_crossentropy",
optimizer="adam",metrics=["accuracy"])
model.fit( X_train, y_train, epochs = 50,
validation_data = (X_valid, y_valid),
callbacks = [es])
# -
# The Early Stopping callback saves us from the overfitting problem. We use sparse categorical crossentropy
# instead of categorical crossentropy because of the representation of delta's used. We could just do one hot encoding
# and apply categorical crossentropy but we chose not.
#
# Now I will plot a few probability distributions and the delta used to make the simulations. Please note that since this
# is a stochastic process the delta simulated doesn't have to coincide with the maximum of the probability function, but
# should be reasonably close.
# +
y_pred = model.predict(X_valid)
for i in [0,25,50,99]:
plt.figure()
plt.plot(Delta_vec, y_pred[i, :], "b-")
plt.xlabel("$\delta$ (normalised units)" , fontsize = 12)
plt.ylabel("Prob" , fontsize = 12)
plt.title("Probability density", fontsize = 14)
plt.vlines(Delta_vec[i], 0, np.amax(y_pred[i, :]))
plt.legend(["MLP probability density", "Simulated $\delta$"], fontsize = 12)
# -
# Now I will introduce a function really useful to evaluate the model performance.
# In this function we make the neural network predict the whole X_valid dataset, then we aggregate the
# probability registers though multiplication and we take the network prediction and the confidence interval
# (which in this case is the full width at half maximum of the distribution) for every value of delta.
#
# We then put it in a plot.
def interval_plot(model, X_valid, Delta_vec, title: str):
# Aquí ponemos el código del benchmark
y_pred=model.predict(X_valid)
n_blocks=int(np.floor(y_pred.shape[0]/100))
# numero de bloques, siendo un bloque las predicciones para todo el dominio
# de deltas
y_pred_product=y_pred[:100,:]**(1/n_blocks) # Los productos
for i in range(n_blocks-1):
y_pred_product=y_pred_product*y_pred[(i+1)*100:(i+1)*100+100,:]**(1/n_blocks)
for ii in range(y_pred_product.shape[0]):
y_pred_product[ii,:]=y_pred_product[ii,:]/(np.sum(y_pred_product[ii,:]))
# Ahora sacamos las lineas de la predicción y del intervalo de confianza
Delta_upper=np.zeros(100)
Delta_mid=np.zeros(100)
Delta_lower=np.zeros(100)
for i in range(100):
arr=y_pred_product[i,:]
max_h=np.amax(arr)
pos_arr=np.where(arr>max_h/2)
Delta_lower[i]=Delta_vec[pos_arr[0][0]]# Tengo que poner un [0] delante para indicar que quiero sacar un elemento de un
Delta_upper[i]=Delta_vec[pos_arr[0][-1]] # array. Personalmente, npi del sentido
for ii in range(100):
Delta_mid[ii]=np.sum(y_pred_product[ii,:]*Delta_vec[:])
plt.figure()
plt.plot(Delta_vec,Delta_vec,"k--")
plt.plot(Delta_vec,Delta_mid,"b-")
plt.fill_between(Delta_vec,Delta_upper,Delta_lower,alpha=0.3,color="red")
plt.legend(["Simulated $\delta$","Network prediction","Confidence interval"],
fontsize = 12)
plt.xlabel("$\delta$",fontsize = 12)
plt.ylabel("Prediction", fontsize = 12)
plt.title(title, fontsize = 14)
plt.show
interval_plot(model, X_valid, Delta_vec, title = "Model performance")
"In case you only want the lines for your own plots"
def interval_plot_lines(model, X_valid, Delta_vec):
# Aquí ponemos el código del benchmark
y_pred=model.predict(X_valid)
n_blocks=int(np.floor(y_pred.shape[0]/100))
# numero de bloques, siendo un bloque las predicciones para todo el dominio
# de deltas
y_pred_product=y_pred[:100,:]**(1/n_blocks) # Los productos
for i in range(n_blocks-1):
y_pred_product=y_pred_product*y_pred[(i+1)*100:(i+1)*100+100,:]**(1/n_blocks)
for ii in range(y_pred_product.shape[0]):
y_pred_product[ii,:]=y_pred_product[ii,:]/(np.sum(y_pred_product[ii,:]))
# Ahora sacamos las lineas de la predicción y del intervalo de confianza
Delta_upper=np.zeros(100)
Delta_mid=np.zeros(100)
Delta_lower=np.zeros(100)
for i in range(100):
arr=y_pred_product[i,:]
max_h=np.amax(arr)
pos_arr=np.where(arr>max_h/2)
Delta_lower[i]=Delta_vec[pos_arr[0][0]]# Tengo que poner un [0] delante para indicar que quiero sacar un elemento de un
Delta_upper[i]=Delta_vec[pos_arr[0][-1]] # array. Personalmente, npi del sentido
for ii in range(100):
Delta_mid[ii]=np.sum(y_pred_product[ii,:]*Delta_vec[:])
return Delta_upper, Delta_mid, Delta_lower
| Basic MLP quantum metrology.ipynb |
/ -*- coding: utf-8 -*-
/ ---
/ jupyter:
/ jupytext:
/ text_representation:
/ extension: .q
/ format_name: light
/ format_version: '1.5'
/ jupytext_version: 1.14.4
/ kernelspec:
/ display_name: SQL
/ language: sql
/ name: SQL
/ ---
/ + [markdown] azdata_cell_guid="e01663cc-427c-457f-84db-b16d0fca3a90"
/ # Query CSV files
/
/ Serverless Synapse SQL pool enables you to read CSV files from Azure storage (DataLake or blob storage).
/
/ ## Read csv file
/
/ The easiest way to see to the content of your `CSV` file is to provide file URL to `OPENROWSET` function and specify format `CSV`. If the file is publicly available or if your Azure AD identity can access this file, you should be able to see the content of the file using the query like the one shown in the following example:
/ + azdata_cell_guid="dbc4f12e-388c-49fa-9d85-0fbea3b19d1b"
select top 10 *
from openrowset(
bulk 'https://pandemicdatalake.blob.core.windows.net/public/curated/covid-19/ecdc_cases/latest/ecdc_cases.csv',
format = 'csv',
parser_version = '2.0',
firstrow = 2 ) as rows
/ + [markdown] azdata_cell_guid="a373fa76-bfdf-4bb6-8098-73c9ef436eb8"
/ ## Data source usage
/
/ Previous example uses full path to the file. As an alternative, you can create an external data source with the location that points to the root folder of the storage:
/ + azdata_cell_guid="48b6ee55-09ec-47df-bea5-707dc2f42aa8"
create external data source covid
with (
location = 'https://pandemicdatalake.blob.core.windows.net/public/curated/covid-19/ecdc_cases'
);
/ + [markdown] azdata_cell_guid="c4145b77-8663-4e59-914b-721955a02635"
/ Once you create a data source, you can use that data source and the relative path to the file in `OPENROWSET` function:
/ + azdata_cell_guid="f3da158c-c168-45b0-8e38-7ee2d430420f"
select
top 10 *
from
openrowset(
bulk 'latest/ecdc_cases.csv',
data_source = 'covid',
format = 'csv',
parser_version ='2.0',
firstrow = 2
) as rows
/ + [markdown] azdata_cell_guid="745b2c81-01eb-4bf5-9cad-47a03dcff194"
/ ## Explicitly specify schema
/
/ `OPENROWSET` enables you to explicitly specify what columns you want to read from the file using `WITH` clause:
/ + azdata_cell_guid="e7bacd03-45d4-4b0b-b1d0-9522e1a54436"
select
top 10 *
from
openrowset(
bulk 'https://pandemicdatalake.blob.core.windows.net/public/curated/covid-19/ecdc_cases/latest/ecdc_cases.csv',
format = 'csv',
parser_version ='2.0',
firstrow = 2
) with (
date_rep date 1,
cases int 5,
geo_id varchar(6) 8
) as rows
/ + [markdown] azdata_cell_guid="4397f453-4b20-4083-ae0e-4966d789993f"
/ The numbers after a data type in the `WITH` clause represent column index in the CSV file.
| Notebooks/TSQL/Jupiter/content/quickstarts/csv.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# ---
#
# _You are currently looking at **version 1.1** of this notebook. To download notebooks and datafiles, as well as get help on Jupyter notebooks in the Coursera platform, visit the [Jupyter Notebook FAQ](https://www.coursera.org/learn/python-text-mining/resources/d9pwm) course resource._
#
# ---
# # Assignment 1
#
# In this assignment, you'll be working with messy medical data and using regex to extract relevant infromation from the data.
#
# Each line of the `dates.txt` file corresponds to a medical note. Each note has a date that needs to be extracted, but each date is encoded in one of many formats.
#
# The goal of this assignment is to correctly identify all of the different date variants encoded in this dataset and to properly normalize and sort the dates.
#
# Here is a list of some of the variants you might encounter in this dataset:
# * 04/20/2009; 04/20/09; 4/20/09; 4/3/09
# * Mar-20-2009; Mar 20, 2009; March 20, 2009; Mar. 20, 2009; Mar 20 2009;
# * 20 Mar 2009; 20 March 2009; 20 Mar. 2009; 20 March, 2009
# * Mar 20th, 2009; Mar 21st, 2009; Mar 22nd, 2009
# * Feb 2009; Sep 2009; Oct 2010
# * 6/2008; 12/2009
# * 2009; 2010
#
# Once you have extracted these date patterns from the text, the next step is to sort them in ascending chronological order accoring to the following rules:
# * Assume all dates in xx/xx/xx format are mm/dd/yy
# * Assume all dates where year is encoded in only two digits are years from the 1900's (e.g. 1/5/89 is January 5th, 1989)
# * If the day is missing (e.g. 9/2009), assume it is the first day of the month (e.g. September 1, 2009).
# * If the month is missing (e.g. 2010), assume it is the first of January of that year (e.g. January 1, 2010).
# * Watch out for potential typos as this is a raw, real-life derived dataset.
#
# With these rules in mind, find the correct date in each note and return a pandas Series in chronological order of the original Series' indices.
#
# For example if the original series was this:
#
# 0 1999
# 1 2010
# 2 1978
# 3 2015
# 4 1985
#
# Your function should return this:
#
# 0 2
# 1 4
# 2 0
# 3 1
# 4 3
#
# Your score will be calculated using [Kendall's tau](https://en.wikipedia.org/wiki/Kendall_rank_correlation_coefficient), a correlation measure for ordinal data.
#
# *This function should return a Series of length 500 and dtype int.*
# +
import pandas as pd
import re
doc = []
with open('dates.txt') as file:
for line in file:
doc.append(line)
df = pd.Series(doc)
df.head(10)
# +
def date_sorter(df):
## establish dataframe
df_t = pd.DataFrame(df)
df_t = df_t.rename(columns={0:'Note'})
# Regex to extract date infor and generate a date df
#df_date = df_t['Note'].iloc[0:125].str.extractall('(?P<month>\d{1,4})\/(?P<day>\d{1,4})\/(?P<year>\d{2,4})')
df_date = df_t['Note'].iloc[0:125].str.extractall('(?P<month>\d{1,4})[/-](?P<day>\d{1,4})[/-](?P<year>\d{2,4})')
df_date2 = df_t['Note'].iloc[125:194].str.extractall('(?P<day>\d{2}) (?P<month>(?:Jan|Feb|Mar|Apr|May|Jun|Aug|Sep|Oct|Nov|Dec)[a-z]*) (?P<year>\d{4})')
df_date3 = df_t['Note'].iloc[125:228].str.extractall('(?P<month>(?:Jan|Feb|Mar|Apr|May|Jun|Jul|Aug|Sep|Oct|Nov|Dec)[a-z,.]*) (?P<day>\d{2})[, ]* (?P<year>\d{4})')
#only month & year
df_date4 = df_t['Note'].iloc[228:343].str.extractall('(?P<month>(?:Jan|Feb|Mar|Apr|May|Jun|Jul|Aug|Sep|Oct|Nov|Dec)[a-z,]*) (?P<year>\d{0,4})')
df_date5 = df_t['Note'].iloc[343:455].str.extractall('(?P<month>\d{1,2})(?P<slash>[\/])(?P<year>\d{2,4})')
df_date5=df_date5.drop('slash', axis='columns')
#only year
df_date6 = df_t['Note'].iloc[455:].str.extractall('(?P<year>\d{4})')
# concat dataframes and manually correct errors
df_trans = pd.concat([df_date, df_date2, df_date3, df_date4,df_date5, df_date6])
df_trans.drop((72, 1), axis='rows', inplace=True)
df_trans.drop((254, 0), axis='rows', inplace=True)
df_trans.drop((289, 1), axis='rows', inplace=True)
df_trans.drop((297, 0), axis='rows', inplace=True)
#df_trans.reset_index(inplace=True)
#df_trans = df_trans.drop('match', axis='columns')
# manually clean data, replace NaN w/ 1's
df_trans = df_trans.fillna(1)
#correct month names
df_trans['month'] = df_trans['month'].str.strip()
df_trans['month'] = df_trans['month'].apply(lambda x: 1 if (x =='Jan' or x=='January'or x=='January,' or x=='Janaury') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 2 if (x=='Feb' or x=='February' or x=='February,') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 3 if (x=='Mar' or x=='March' or x=='Mar.' or x=='Mar,' or x=='March,') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 4 if (x=='Apr' or x=='April' or x=='Apr,') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 5 if (x=='May' or x=='May,') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 6 if (x=='Jun' or x=='June' or x=='June,') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 7 if (x=='Jul' or x=='July' or x=='July,') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 8 if (x=='Aug' or x=='August') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 9 if (x=='Sep' or x=='September' or x=='Sep.' or x=='September.') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 10 if(x=='Oct' or x=='October' or x=='October.') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 11 if(x=='Nov' or x=='November') else x)
df_trans['month'] = df_trans['month'].apply(lambda x: 12 if(x=='Dec' or x=='December' or x=='Decemeber') else x)
df_trans = df_trans.fillna(1)
# convert to numeric
df_trans['year']=pd.to_numeric(df_trans['year'], errors='raise', downcast='integer')
df_trans['day']=pd.to_numeric(df_trans['day'], errors='raise', downcast='integer')
df_trans['month']=pd.to_numeric(df_trans['month'], errors='raise', downcast='integer')
#convert year
df_trans['year'] = df_trans['year'].apply(lambda x: 1900 + x if(x <100 and x>30) else x)
#sort
df_trans.sort_values(by=['year', 'month', 'day'], inplace = True)
df_trans.reset_index(inplace=True)
df_s = df_trans['level_0']
# '''
# https://www.coursera.org/learn/python-text-mining/discussions/weeks/1/threads/9xo_YiTyEeuZ9Q5xsF_ZCQ
# https://www.coursera.org/learn/python-text-mining/discussions/weeks/1/threads/9xo_YiTyEeuZ9Q5xsF_ZCQ
# https://www.coursera.org/learn/python-text-mining/discussions/weeks/1/threads/AoLX8rSbEeiqnRI0WnAb-A
# '''
return df_s # Your answer here
a = date_sorter(df)
a
# -
| Python/Applied_Text_Mining_Assignment+1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# + [markdown] id="zHZRulFLg3RQ"
# #### Importing dependencies
# ---
# + id="DsGHwcezg3RQ" pycharm={"name": "#%%\n"}
import pandas as pd
import numpy as np
import os
os.environ['TF_CPP_MIN_LOG_LEVEL'] = '3'
import tensorflow as tf
from sklearn.preprocessing import MinMaxScaler, StandardScaler
from sklearn.metrics import accuracy_score
from sklearn.model_selection import train_test_split
from sklearn.model_selection import KFold
# -
os.chdir(os.getcwd().replace('Notebooks', ''))
os.getcwd()
from GitMarco.tf.utils import limit_memory
limit_memory(2000)
# + [markdown] id="k7pk2MjRg3RR"
# #### Loading data (csv format)
# ---
# + id="trnAsZedg3RR" pycharm={"name": "#%%\n"}
# Reading the train and test data
import os
df = pd.read_excel('tabular/df.xlsx')
# + [markdown] id="B5PVbJ1dg3RS"
# #### Check data type
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="ozun0gXpg3RS" outputId="3ac86e1d-db46-4a3e-d139-246bfa2759b1" pycharm={"name": "#%%\n"}
df.info()
# + [markdown] id="N9ihvqZdg3RT"
# #### Drop desired columns from the dataset
# ---
# + id="_TFrLZ9Ag3RT" pycharm={"name": "#%%\n"}
rem_cols = None
# + id="fw6tOtsvg3RU" pycharm={"name": "#%%\n"}
if rem_cols is not None:
df.drop(rem_cols, axis=1, inplace=True)
df.head()
# + [markdown] id="QLSTw5tug3RU"
# #### Cleaning
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="DeoLSEB0g3RU" outputId="6d3069d7-2984-4fb3-e9bb-24190038f4bb" pycharm={"name": "#%%\n"}
print(df.shape)
# df = df[df.CNT_CHILDREN != 12]
print(df.shape)
# + [markdown] id="NR9WEIceg3RV"
# #### Removing NaN values
# ---
#
# + colab={"base_uri": "https://localhost:8080/"} id="6evkhF2yg3RV" outputId="55e437bf-0db9-42fe-a6f5-fd7f0a6722b6" pycharm={"name": "#%%\n"}
print('Before', df.shape)
df.dropna(axis=0, how='any', thresh=None, subset=None, inplace=True)
print('After', df.shape)
# + [markdown] id="C2R_HrK3g3RV"
# #### Fixing data
# ---
# + id="5fFrvHBUg3RV" pycharm={"name": "#%%\n"}
# df.DAYS_EMPLOYED[df.DAYS_EMPLOYED == 365243] = 1
# df.head()
# + [markdown] id="-JMdag5Wg3RW"
# #### Performing Factorization
# ---
# We transform the desired columns into factorized classes
# + id="lBiW8ubLg3RW" pycharm={"name": "#%%\n"}
cols = None
if cols is not None:
df[cols] = df[cols].apply(lambda x: pd.factorize(x)[0])
df.head()
# + [markdown] id="Mayvi5vGg3RW"
# #### Checking again data dtype
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="ZER68AIeg3RX" outputId="8d702474-7a16-4203-c6d8-cb4675799e17" pycharm={"name": "#%%\n"}
df.info()
# + [markdown] id="3gCNDarFg3RX"
# #### Checking stats
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="CnuMzo37g3RX" outputId="ee9e2fb2-87c6-4493-f653-5b6ec84a05bd" pycharm={"name": "#%%\n"}
df.describe().T
# + [markdown] id="5YvvBACYg3RX"
# #### Ditribution
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 281} id="7suVLbWhg3RX" outputId="71845f2a-1f7a-4207-86d3-386738583d43" pycharm={"name": "#%%\n"}
df.hist(df.columns[-4])
pass
# + [markdown] id="_gz9uvysg3RX"
# #### Preparing data for training, validation and test
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 317} id="8TZvHf4Bg3RX" outputId="acba2508-1e85-4c87-9bb2-6fb2608c5e9f" pycharm={"name": "#%%\n"}
input_data = df.copy()
input_data.head()
# + [markdown] id="S1cpM8bSg3RX"
# #### Normalizing data
# ---
# + colab={"base_uri": "https://localhost:8080/", "height": 424} id="WCe9UkoOg3RX" outputId="c655f358-83ad-41e4-e566-766da36442f4" pycharm={"name": "#%%\n"}
scaler = StandardScaler()
scaler.fit(input_data)
normed_data = pd.DataFrame(scaler.transform(input_data), columns=input_data.columns)
normed_data.describe().T
# + [markdown] id="QjayZ6USg3RY"
# #### Splitting data into training, validation and testing
# ---
# + id="JVRwa8gxg3RY" pycharm={"name": "#%%\n"}
n_labels = 1
labels = normed_data.drop(columns=df.columns[:-n_labels])
normed_data = normed_data.drop(columns=df.columns[-n_labels:])
train_data, test_data, train_labels, test_labels = train_test_split(
normed_data, labels, test_size=0.2, shuffle=True)
# + [markdown] id="U8IFX7Dhg3RZ"
# #### Creating a parametric sequential model
# ---
# + id="vxxiJuAmg3RZ" pycharm={"name": "#%%\n"}
from GitMarco.tf.metrics import r_squared
from GitMarco.graphics.matplotlib import validation_plot
def create_model(dropout_rate: float = 0.0,
neurons: int = 32,
activation: str = 'relu',
n_layers: int = 2,
learning_rate: float = 0.001,
optimizer = tf.keras.optimizers.Adam,
nesterov: bool = True,
momentum: float = .9,
) -> tf.keras.Model:
model = tf.keras.models.Sequential()
model.add(tf.keras.layers.Dense(neurons,
activation=activation,
input_shape=(normed_data.shape[1], )))
model.add(tf.keras.layers.Dropout(dropout_rate))
for i in range(1, n_layers):
model.add(tf.keras.layers.Dense(neurons, activation=activation))
model.add(tf.keras.layers.Dropout(dropout_rate))
model.add(tf.keras.layers.Dense(train_labels.shape[1]))
if isinstance(optimizer, tf.keras.optimizers.SGD):
model.compile(optimizer=optimizer(learning_rate=learning_rate, nesterov=nesterov, momentum=momentum),
loss=tf.keras.losses.mean_squared_error,
# metrics=[r_squared]
)
else:
model.compile(optimizer=optimizer(learning_rate=learning_rate),
loss=tf.keras.losses.mean_squared_error,
# metrics=[r_squared]
)
model.summary()
return model
# + [markdown] id="ulRLM-mBg3RZ"
# #### Wrap keras model with sklearn
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="NSNVYxmFg3RZ" outputId="af563806-d836-4cf3-9c8a-8cf9197fea8f" pycharm={"name": "#%%\n"}
from keras.wrappers.scikit_learn import KerasRegressor
model = KerasRegressor(build_fn=create_model, verbose=0)
# + [markdown] id="SVbEC0ENg3RZ"
# #### Performing cross-validation and hyper-parameters optimization
# ---
# + id="Fgaf_Wpfg3RZ" pycharm={"name": "#%%\n"}
# We specify here the parameters of the grid search, in the form of lists
learning_rate = [0.01,]
dropout_rate = [0.2]
batch_size = [100]
epochs = [2000, ]
neurons = [256,]
activation = ['relu',]
n_layers = [2,]
nesterov = [True,]
momentum = [.9, ]
optimizer = [
tf.keras.optimizers.Adam,
# tf.keras.optimizers.SGD,
# tf.keras.optimizers.RMSprop,
]
# Make a dictionary of the grid search parameters
param_grid = dict(learning_rate=learning_rate,
dropout_rate=dropout_rate,
batch_size=batch_size,
epochs=epochs,
neurons=neurons,
activation=activation,
n_layers=n_layers,
optimizer=optimizer,
nesterov=nesterov,
momentum=momentum)
# + colab={"base_uri": "https://localhost:8080/"} id="3TK5PWVXg3RZ" outputId="e57612aa-66f9-4ec1-93f7-ce9ee8000fba" pycharm={"name": "#%%\n"} tags=[]
from sklearn.model_selection import GridSearchCV
from sklearn.metrics import r2_score
seed = 22 # Set random state
jobs = 1 # Set the number of parallel processes (-1 => all available cores)
n_folds = 5 # Number of cross-validation folds
# Build and fit the GridSearchCV
grid = GridSearchCV(estimator=model,
param_grid=param_grid,
cv=KFold(random_state=seed,
n_splits=n_folds,
shuffle=True
),
scoring='neg_mean_squared_error',
error_score='raise',
verbose=0,
n_jobs=jobs)
grid_results = grid.fit(train_data, train_labels, verbose=0)
# + [markdown] id="SEzWq_5Gg3RZ"
# #### Explore Results
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="-UwHHO6yg3RZ" outputId="090ef7a2-e18d-4bd5-b680-c6043c917140" pycharm={"name": "#%%\n"}
# Summarize the results in a readable format
print("Best: {0}, using {1} \n".format(grid_results.best_score_, grid_results.best_params_))
# + colab={"base_uri": "https://localhost:8080/"} id="VZ81bfOzg3RZ" outputId="ea7e4509-0214-4ced-beb4-3e98118a1ef5" pycharm={"name": "#%%\n"}
means = grid_results.cv_results_['mean_test_score']
stds = grid_results.cv_results_['std_test_score']
params = grid_results.cv_results_['params']
for mean, stdev, param in zip(means, stds, params):
print('{0} ({1}) with: {2}\n'.format(mean, stdev, param))
# + colab={"base_uri": "https://localhost:8080/"} id="Jry2qBKsg3RZ" outputId="87b9d88a-bcf2-4723-e4a9-4c3fa636aa36" pycharm={"name": "#%%\n"}
vars(grid_results).keys()
# + [markdown] id="90I_Of5cg3RZ"
# It is possible to post-process the results of the analysis in various ways in order to create detailed reports describing the performance of the classifier as its hyperparameters change @TODO
# + [markdown] id="G278Wfo8g3RZ"
# #### Extracting the best model
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="OvGDO_Jtg3Ra" outputId="3974dded-65ab-4d91-85ac-b080a8ccc661" pycharm={"name": "#%%\n"}
vars(grid_results.best_estimator_).keys()
# + id="R6TEq4irg3Ra" pycharm={"name": "#%%\n"}
final_model = grid_results.best_estimator_.model
# + [markdown] id="9RaxzM8-g3Ra"
# #### Evaluating the best model on the test set
# ---
# + colab={"base_uri": "https://localhost:8080/"} id="Ldt39A45g3Ra" outputId="5f0bb981-d71a-408f-9c32-45dbd9ebc05b" pycharm={"name": "#%%\n"}
train_scores = final_model.evaluate(train_data, train_labels)
print('Train loss: {0}'.format(train_scores))
# + colab={"base_uri": "https://localhost:8080/"} id="odlie9xM05ih" outputId="d01a89d8-69e7-4881-f9c1-1f5bdab2efab"
scores = []
for i in range(train_labels.shape[1]):
R2_ = r2_score(train_labels.to_numpy()[:, i], final_model.predict(train_data)[:, i])
print(R2_)
scores.append(R2_)
print('\n')
np.mean(scores)
# + colab={"base_uri": "https://localhost:8080/"} id="fhJdYUAtg3Ra" outputId="3ca15a41-65ad-4d9f-a5e8-b09611df0b5c" pycharm={"name": "#%%\n"}
test_scores = final_model.evaluate(test_data, test_labels)
print('Test loss: {0}'.format(test_scores))
# + colab={"base_uri": "https://localhost:8080/"} id="NFGIHcf81Eu6" outputId="e891c15a-227b-49bc-8646-bcbfeee7ac62"
r2_score(test_labels, final_model.predict(test_data))
# + [markdown] id="D9ePtAvcg3Ra"
# #### Saving training data, test data and best model
# ---
# + id="clwcQi4Rg3Ra" pycharm={"name": "#%%\n"}
results_path = 'results_tabular'
# + id="5OGk1g6-g3Ra" pycharm={"name": "#%%\n"}
if os.path.exists(results_path):
os.system('rm -r {0}'.format(results_path))
os.mkdir(results_path)
else:
os.mkdir(results_path)
# + id="m4S_YhYrg3Ra" pycharm={"name": "#%%\n"}
np_to_csv = lambda x, y: np.savetxt(f"{y}.csv", x, delimiter=",")
# + id="XO2zUOwwg3Ra" pycharm={"name": "#%%\n"}
train_data.to_csv(os.path.join(results_path, 'train_data.csv'))
test_data.to_csv(os.path.join(results_path, 'test_data.csv'))
np_to_csv(train_labels, os.path.join(results_path, 'train_labels'))
np_to_csv(test_labels, os.path.join(results_path, 'test_labels'))
# + colab={"base_uri": "https://localhost:8080/"} id="RFeNiMSTg3Ra" outputId="365a0338-9356-4745-93f4-422884ff671b" pycharm={"name": "#%%\n"}
final_model.save(os.path.join(results_path, 'best_model'))
# + colab={"base_uri": "https://localhost:8080/"} id="Q__6dSre0AQX" outputId="b6d90aa5-91ef-4f18-a865-ad34ae9579eb"
# !zip -r results_tabular/best_model.zip results_tabular/best_model
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="KxrwtCcD13Jc" outputId="e1314efe-41fa-428b-fb48-7a096b223129"
import matplotlib.pyplot as plt
plt.style.use('seaborn-darkgrid')
for j in range(train_labels.shape[1]):
plot = validation_plot(train_labels.iloc[:, j].to_numpy(), final_model.predict(train_data)[:, j], show=True, title='Training', marker_color='red')
# + colab={"base_uri": "https://localhost:8080/", "height": 1000} id="N-dDXwBZ1-iD" outputId="1cfc3bb4-c114-44a2-e802-15b6c50169a8"
for k in range(test_labels.shape[1]):
plot = validation_plot(test_labels.iloc[:, k].to_numpy(), final_model.predict(test_data)[:, k], show=True, title='Test', marker_color='red')
# + id="qLtqaprZ218M"
| GitMarco/notebooks/standard_regression_keras.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Exercise 1: Extending the Pan-Tompkins Algorithm
#
# The Pan-Tompkins algorithm in the previous video is a basic version of the algorithm. In this exercise we will add features to the decision rules to improve its performance.
#
# ## Imports
# +
import numpy as np
ts = np.arange(0, 5, 1/100)
sinusoid = 3 * np.sin(2 * np.pi * 1 * ts + np.pi) + 10
# +
import glob
import os
import numpy as np
import pandas as pd
import scipy as sp
import scipy.signal
np.warnings.filterwarnings('ignore')
# -
# ## Performance Evaluation Helpers
#
# First, we need to build a function that tells us the performance of our QRS estimates. We will optimize for precision and recall. These two functions should help us do that.
def Evaluate(reference_peak_indices, estimate_peak_indices, tolerance_samples=40):
"""Evaluates algorithm performance for a single dataset.
It is not expected that reference and estimate peak indices overlap exactly.
Instead say a QRS estimate is correct if it is within <tolerance_samples> of
a reference estimate.
Args:
reference_peak_indices: (np.array) ground-truth array of QRS complex locations
estiamte_peak_indices: (np.array) array of QRS complex estimates
tolerance_samples: (number) How close a QRS estimate needs to be to a reference
location to be correct.
Returns:
n_correct: (number) The number of QRS complexes that were correctly detected
n_missed: (number) The number of QRS complexes that the algorithm failed
to detect
n_extra: (number) The number of spurious QRS complexes detected by the
algorithm
"""
# Keep track of the number of QRS peaks that were found correctly
n_correct = 0
# ... that were missed
n_missed = 0
# ... and that are spurious
n_extra = 0
# Loop counters
i, j = 0, 0
while (i < len(reference_peak_indices)) and (j < len(estimate_peak_indices)):
# Iterate through the arrays of QRS peaks, counting the number of peaks
# that are correct, missed, and extra.
ref = reference_peak_indices[i]
est = estimate_peak_indices[j]
if abs(ref - est) < tolerance_samples:
# If the reference peak and the estimate peak are within <tolerance_samples>,
# then we mark this beat correctly detected and move on to the next one.
n_correct += 1
i += 1
j += 1
continue
if ref < est:
# Else, if they are farther apart and the reference is before the estimate,
# then the detector missed a beat and we advance the reference array.
n_missed += 1
i += 1
continue
# Else, the estimate is before the reference. This means we found an extra beat
# in the estimate array. We advance the estimate array to check the next beat.
j += 1
n_extra += 1
# Don't forget to count the number of missed or extra peaks at the end of the array.
n_missed += len(reference_peak_indices[i:])
n_extra += len(estimate_peak_indices[j:])
return n_correct, n_missed, n_extra
# Now we need a function that can compute precision and recall for us.
def PrecisionRecall(n_correct, n_missed, n_extra):
# TODO: Compute precision and recall from the input arguments.
precision = None
recall = None
return precision, recall
# ## Pan-Tompkins Algorithm
#
# We will start with the same algorithm that you saw in the last video. This starter code differs only in that we do not *LocalizeMaxima* on the output peaks. This is because for this dataset the QRS complexes could be pointing up or down and if we try to find the maxima when the QRS complex is pointing downward we will hurt our algorithm performance. Instead we will be happy with the approximate QRS locations that our algorithm detects.
#
# The current version of the algorithm has a precision and recall of 0.89 and 0.74. Verify this by running the next cell. Your task is to improve the performance of the algorithm by adding the following features.
#
# ### Refractory Period Blanking
# Recall from the physiology lesson that the QRS complex is a result of ventricular depolarization, and that cellular depolarization happens when ions travel across the cell membrane. There is a physiological constraint on how soon consecutive depolarization can occur. This constraint is 200 ms. Read more about it [here](https://en.wikipedia.org/wiki/Refractory_period_(physiology)#Cardiac_refractory_period). We can take advantage of this phenomenon in our algorithm by removing detections that occur within 200ms of another one. Preserve the larger detection.
#
# ### Adaptive Thresholding
# The QRS complex height can change over time as contact with the electrodes changes or shifts. Instead of using a fixed threshold, we should use one that changes over time. Make the detection threshold 70% of the average peak height for the last 8 peaks.
#
# ### T-Wave Discrimination
# One error mode is to detect T-waves as QRS complexes. We can avoid picking T-waves by doing the following:
# * Find peaks that follow a previous one by 360ms or less
# * Compute the maximum absolute slope within 60ms of each peak. Eg `np.max(np.abs(np.diff(ecg[peak - 60ms: peak + 60ms])))`
# * If the slope of the second peak is less than half of the slope of the first peak, discard the second peak as a T-wave
# Read another description of this technique [here](https://en.wikipedia.org/wiki/Pan%E2%80%93Tompkins_algorithm#T_wave_discrimination)
#
# After implementing these three techniques you should see a significant increase in precision and recall. I ended up with 0.95 and 0.87. See if you can beat that!
# +
def BandpassFilter(signal, fs=300):
"""Bandpass filter the signal between 5 and 15 Hz."""
b, a = sp.signal.butter(3, (5, 15), btype='bandpass', fs=fs)
return sp.signal.filtfilt(b, a, signal)
def MovingSum(signal, fs=300):
"""Moving sum operation with window size of 150ms."""
n_samples = int(round(fs * 0.150))
return pd.Series(signal).rolling(n_samples, center=True).sum().values
def FindPeaks(signal, order=10):
"""A simple peak detection algorithm."""
msk = (signal[order:-order] > signal[:-order * 2]) & (signal[order:-order] > signal[order * 2:])
for o in range(1, order):
msk &= (signal[order:-order] > signal[o: -order * 2 + o])
msk &= (signal[order:-order] > signal[order * 2 - o: -o])
return msk.nonzero()[0] + order
def ThresholdPeaks(filtered_signal, peaks):
"""Threshold detected peaks to select the QRS complexes."""
thresh = np.mean(filtered_signal[peaks])
return peaks[filtered_signal[peaks] > thresh]
def AdaptiveThresholdPeaks(filtered_signal, peaks):
# TODO: Implement adaptive thresholding
pass
def RefractoryPeriodBlanking(filtered_signal, peaks, fs, refractory_period_ms=200):
# TODO: Implement refractory period blanking
pass
def TWaveDiscrimination(signal, peaks, fs, twave_window_ms=360, slope_window_ms=60):
# TODO: Implement t-wave discrimination
pass
def PanTompkinsPeaks(signal, fs):
"""Pan-Tompkins QRS complex detection algorithm."""
filtered_signal = MovingSum(
np.square(
np.diff(
BandpassFilter(signal, fs))), fs)
peaks = FindPeaks(filtered_signal)
#peaks = RefractoryPeriodBlanking(filtered_signal, peaks, fs) # TODO: Uncomment this line
peaks = ThresholdPeaks(filtered_signal, peaks) # TODO: Remove this line
#peaks = AdaptiveThresholdPeaks(filtered_signal, peaks) # TODO: Uncomment this line
#peaks = TWaveDiscrimination(signal, peaks, fs) # TODO: Uncomment this line
return peaks
# -
# ## Load Data and Evaluate Performance
#
# As we add features to the algorithm we can continue to evaluate it and see the change in performance. Use the code below to compute an overall precision and recall for QRS detection. You must first implement the `PrecisionRecall` function above.
# +
# This dataset is sampled at 300 Hz.
fs = 300
files = glob.glob('../../data/cinc/*.npz')
# Keep track of the total number of correct, missed, and extra detections.
total_correct, total_missed, total_extra = 0, 0, 0
for i, fl in enumerate(files):
# For each file, load the data...
with np.load(fl) as npz:
ecg = npz['ecg']
reference_peak_indices = npz['qrs']
# Compute our QRS location estimates...
estimate_peak_indices = PanTompkinsPeaks(ecg, fs)
# Compare our estimates against the reference...
n_correct, n_missed, n_extra = Evaluate(reference_peak_indices, estimate_peak_indices)
# And add them to our running totals.
total_correct += n_correct
total_missed += n_missed
total_extra += n_extra
print('\r{}/{} files processed...'.format(i+1, len(files)), end='')
print('') # print a newline
# Compute and report the overall performance.
precision, recall = PrecisionRecall(total_correct, total_missed, total_extra)
print('Total performance:\n\tPrecision = {:0.2f}\n\tRecall = {:0.2f}'.format(precision, recall))
| AI-for-Healthcare/wearable-data/lesson 4/1_pan_tompkins_algorithm.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Advanced Tutorial 5: Scheduler
# In this tutorial, we will talk about:
# * [Scheduler](#ta05scheduler)
# * [Concept](#ta05concept)
# * [EpochScheduler](#ta05epoch)
# * [RepeatScheduler](#ta05repeat)
# * [Things You Can Schedule](#ta05things)
# * [Datasets](#ta05dataset)
# * [Batch Size](#ta05batch)
# * [NumpyOps](#ta05numpy)
# * [Optimizers](#ta05optimizer)
# * [TensorOps](#ta05tensor)
# * [Traces](#ta05trace)
# * [Related Apphub Examples](#ta05apphub)
# <a id='ta05scheduler'></a>
# ## Scheduler
# <a id='ta05concept'></a>
# ### Concept
# Deep learning training is getting more complicated every year. One major aspect of this complexity is time-dependent training. For example:
#
# * Using different datasets for different training epochs.
# * Applying different preprocessing for different epochs.
# * Training different networks on different epochs.
# * ...
#
# The list goes on and on. In order to provide an easy way for users to accomplish time-dependent training, we provide the `Scheduler` class which can help you schedule any part of the training.
#
# Please note that the basic time unit that `Scheduler` can handle is `epochs`. If users want arbitrary scheduling cycles, the simplest way is to customize the length of one epoch in `Estimator` using max_train_steps_per_epoch.
# <a id='ta05epoch'></a>
# ### EpochScheduler
# The most straightforward way to schedule things is through an epoch-value mapping. For example, If users want to schedule the batch size in the following way:
#
# * epoch 1 - batchsize 16
# * epoch 2 - batchsize 32
# * epoch 3 - batchsize 32
# * epoch 4 - batchsize 64
# * epoch 5 - batchsize 64
#
# You can do the following:
from fastestimator.schedule import EpochScheduler
batch_size = EpochScheduler(epoch_dict={1:16, 2:32, 4:64})
for epoch in range(1, 6):
print("At epoch {}, batch size is {}".format(epoch, batch_size.get_current_value(epoch)))
# <a id='ta05repeat'></a>
# ### RepeatScheduler
# If your schedule follows a repeating pattern, then you don't want to specify that for all epochs. `RepeatScheduler` is here to help you. Let's say we want the batch size on odd epochs to be 32, and on even epochs to be 64:
# +
from fastestimator.schedule import RepeatScheduler
batch_size = RepeatScheduler(repeat_list=[32, 64])
for epoch in range(1, 6):
print("At epoch {}, batch size is {}".format(epoch, batch_size.get_current_value(epoch)))
# -
# <a id='ta05things'></a>
# ## Things You Can Schedule:
# <a id='ta05dataset'></a>
# ### Datasets
# Scheduling training or evaluation datasets is very common in deep learning. For example, in curriculum learning people will train on an easy dataset first and then gradually move on to harder datasets. For illustration purposes, let's use two different instances of the same MNIST dataset:
# +
from fastestimator.dataset.data import mnist, cifair10
from fastestimator.schedule import EpochScheduler
train_data1, eval_data = mnist.load_data()
train_data2, _ = mnist.load_data()
train_data = EpochScheduler(epoch_dict={1:train_data1, 3: train_data2})
# -
# <a id='ta05batch'></a>
# ### Batch Size
# We can also schedule the batch size on different epochs, which may help resolve GPU resource constraints.
batch_size = RepeatScheduler(repeat_list=[32,64])
# <a id='ta05numpy'></a>
# ### NumpyOps
# Preprocessing operators can also be scheduled. For illustration purpose, we will apply a `Rotation` for the first two epochs and then not apply it for the third epoch:
# +
from fastestimator.op.numpyop.univariate import ExpandDims, Minmax
from fastestimator.op.numpyop.multivariate import Rotate
import fastestimator as fe
rotate_op = EpochScheduler(epoch_dict={1:Rotate(image_in="x", image_out="x",limit=30), 3:None})
pipeline = fe.Pipeline(train_data=train_data,
eval_data=eval_data,
batch_size=batch_size,
ops=[ExpandDims(inputs="x", outputs="x"), rotate_op, Minmax(inputs="x", outputs="x")])
# -
# <a id='ta05optimizer'></a>
# ### Optimizers
# For fast convergence, some people like to use different optimizers at different training phases. In our example, we will use `adam` for the first epoch and `sgd` for the second epoch.
# +
from fastestimator.architecture.tensorflow import LeNet
model_1 = fe.build(model_fn=LeNet, optimizer_fn=EpochScheduler(epoch_dict={1:"adam", 2: "sgd"}), model_name="m1")
# -
# <a id='ta05tensor'></a>
# ### TensorOps
# We can schedule `TensorOps` just like `NumpyOps`. Let's define another model `model_2` such that:
# * epoch 1-2: train `model_1`
# * epoch 3: train `model_2`
# +
from fastestimator.op.tensorop.model import ModelOp, UpdateOp
from fastestimator.op.tensorop.loss import CrossEntropy
model_2 = fe.build(model_fn=LeNet, optimizer_fn="adam", model_name="m2")
model_map = {1: ModelOp(model=model_1, inputs="x", outputs="y_pred"),
3: ModelOp(model=model_2, inputs="x", outputs="y_pred")}
update_map = {1: UpdateOp(model=model_1, loss_name="ce"), 3: UpdateOp(model=model_2, loss_name="ce")}
network = fe.Network(ops=[EpochScheduler(model_map),
CrossEntropy(inputs=("y_pred", "y"), outputs="ce"),
EpochScheduler(update_map)])
# -
# <a id='ta05trace'></a>
# ### Traces
# `Traces` can also be scheduled. For example, we will save `model_1` at the end of second epoch and save `model_3` at the end of third epoch:
# +
from fastestimator.trace.io import ModelSaver
import tempfile
save_folder = tempfile.mkdtemp()
#Disable model saving by setting None on 3rd epoch:
modelsaver1 = EpochScheduler({2:ModelSaver(model=model_1,save_dir=save_folder), 3:None})
modelsaver2 = EpochScheduler({3:ModelSaver(model=model_2,save_dir=save_folder)})
traces=[modelsaver1, modelsaver2]
# -
# ## Let the training begin
# Nothing special in here, create the estimator then start the training:
estimator = fe.Estimator(pipeline=pipeline, network=network, traces=traces, epochs=3, log_steps=300)
estimator.fit()
# <a id='ta05apphub'></a>
# ## Apphub Examples
# You can find some practical examples of the concepts described here in the following FastEstimator Apphubs:
#
# * [PGGAN](../../apphub/image_generation/pggan/pggan.ipynb)
| tutorial/advanced/t05_scheduler.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
# Setup
from hcipy import *
import numpy as np
from matplotlib import pyplot as plt
N = 256
D = 9.96
sps = int(40 * N/128)
pupsep = 65/39.3
D_grid = 3.6e-3
aperture = circular_aperture(D)
pupil_grid = make_pupil_grid(N, D)
wf = Wavefront(aperture(pupil_grid))
aberrated = wf.copy()
amplitude = 0.3
spatial_frequency = 5
aberrated.electric_field *= np.exp(1j * amplitude * np.sin(2*np.pi * pupil_grid.x / D * spatial_frequency))
# +
# Making an aberration basis. This is in O(N^2) so doubling it quadruples the runtime.
aberration_mode_basis = []
try:
for i in range(N):
for j in range(N):
wf = Wavefront(aperture(pupil_grid))
wf.electric_field.shape = (N, N)
l = wf.electric_field.tolist()
if np.real(l[i][j]) > 0:
l[i][j] = 0
wf.electric_field = Field(np.asarray(l).ravel(), wf.grid)
aberration_mode_basis.append(wf)
basis_size = len(aberration_mode_basis)
print(basis_size)
except KeyboardInterrupt:
print(len(aberration_mode_basis))
# -
# Propagating to pyramid. (Longest step, about 8-9 minutes, and probably also O(N^2) so 36 minutes for N=256)
keck_pyramid = PyramidWavefrontSensorOptics(pupil_grid, pupil_separation=pupsep, num_pupil_pixels=sps)
pyramid_output_basis = [keck_pyramid.forward(x) for x in aberration_mode_basis]
# +
def get_sub_images(intensity):
pyramid_grid = make_pupil_grid(N, D_grid)
images = Field(np.asarray(intensity).ravel(), pyramid_grid)
pysize = int(np.sqrt(images.size))
images.shape = (pysize, pysize)
sub_images = [images[pysize-sps-1:pysize-1, pysize-sps-1:pysize-1],
images[pysize-sps-1:pysize-1, 0:sps],
images[0:sps, 0:sps],
images[0:sps, pysize-sps-1:pysize-1]]
subimage_grid = make_pupil_grid(sps, D_grid * sps / N)
for count, img in enumerate(sub_images):
img = img.ravel()
img.grid = subimage_grid
sub_images[count] = img
return sub_images
sub_images_basis = [get_sub_images(x.intensity) for x in pyramid_output_basis]
# +
def estimate(EstimatorObject, images_list):
# Restored the numbering convention to what it was originally because this version of sub_images handles it.
I_a = images_list[0]
I_b = images_list[1]
I_c = images_list[2]
I_d = images_list[3]
norm = I_a + I_b + I_c + I_d
I_x = (I_a + I_b - I_c - I_d) / norm
I_y = (I_a - I_b - I_c + I_d) / norm
return [I_x.ravel(), I_y.ravel()]
keck_pyramid_estimator = PyramidWavefrontSensorEstimator(aperture, make_pupil_grid(sps*2, D_grid*sps*2/N))
estimated_basis = [estimate(keck_pyramid_estimator, x) for x in sub_images_basis]
# -
# All the above steps, to generate flat wavefront slopes
flat_mode = Wavefront(aperture(pupil_grid))
flat_pyramid_output = keck_pyramid.forward(flat_mode)
flat_x, flat_y = estimate(keck_pyramid_estimator, get_sub_images(flat_pyramid_output.intensity))
estimated_basis = np.asarray([[x - flat_x, y - flat_y] for x, y in estimated_basis])
# In regular Python, write to file here, and read in a separate file. Here, continuing as usual.
M_inv = field_inverse_tikhonov(Field(estimated_basis, make_pupil_grid(sps, D)), 1e-15)
M_inv = M_inv.copy()
M_inv.shape = (basis_size, 2*sps*sps)
# Quickly do all the above without explaining anything
get_pyramid_output = lambda wf: np.asarray(estimate(keck_pyramid_estimator, get_sub_images(keck_pyramid.forward(wf).intensity))).ravel() #should return a (2*sps*sps,) size NumPy Array
# +
# make
aberrated_images = get_pyramid_output(aberrated)
flat_images = get_pyramid_output(Wavefront(aperture(pupil_grid)))
aberrated_res = aberrated_images - flat_images
reconstructed = M_inv.dot(aberrated_res).tolist()
project_onto = Wavefront(aperture(pupil_grid)).electric_field
project_onto.shape = (N, N)
project_onto = project_onto.tolist()
count, i, j = 0, 0, 0
while count < basis_size:
if np.real(project_onto[i][j]) > 0:
project_onto[i][j] = reconstructed[count]
count += 1
j += 1
if j == N - 1:
j = 0
i += 1
# -
imshow_field(aberrated.phase - Wavefront(aperture(pupil_grid)).intensity, pupil_grid)
plt.colorbar()
plt.show()
imshow_field(np.asarray(project_onto).ravel() * aperture(pupil_grid), pupil_grid)
plt.colorbar()
plt.show()
imshow_field(np.asarray(project_onto).ravel() * aperture(pupil_grid), pupil_grid, vmin=-0.05, vmax=0.05)
plt.colorbar()
plt.show()
| pwfs-keck/t/Wavefront Reconstruction v1.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# <script async src="https://www.googletagmanager.com/gtag/js?id=UA-59152712-8"></script>
# <script>
# window.dataLayer = window.dataLayer || [];
# function gtag(){dataLayer.push(arguments);}
# gtag('js', new Date());
#
# gtag('config', 'UA-59152712-8');
# </script>
#
# # $H_{\rm Orb, NS}$, up to and including third post-Newtonian order
#
# ## This notebook constructs the orbital, nonspinning Hamiltonian up to third post-Newtonian order, as summarized in [Buonanno, Chen, and Damour (2006)](https://arxiv.org/abs/gr-qc/0508067) (see references therein for sources)
#
# **Notebook Status:** <font color='green'><b> Validated </b></font>
#
# **Validation Notes:** All expressions in this notebook were transcribed twice by hand on separate occasions, and expressions were corrected as needed to ensure consistency with published PN expressions. In addition, this tutorial notebook has been confirmed to be self-consistent with its corresponding NRPy+ module, as documented [below](#code_validation). **Additional validation tests may have been performed, but are as yet, undocumented.**
#
#
# ## Author: <NAME>
#
# ### This notebook exists as the following Python module:
# 1. [PN_Hamiltonian_NS.py](../../edit/NRPyPN/PN_Hamiltonian_NS.py)
#
# ### This notebook & corresponding Python module depend on the following NRPy+/NRPyPN Python modules:
# 1. [indexedexp.py](../../edit/indexedexp.py): [**documentation+tutorial**](../Tutorial-Indexed_Expressions.ipynb)
# 1. [NRPyPN_shortcuts.py](../../edit/NRPyPN/NRPyPN_shortcuts.py): [**documentation**](NRPyPN_shortcuts.ipynb)
# <a id='toc'></a>
#
# # Table of Contents
# $$\label{toc}$$
#
# 1. Part 1: [$H_{\rm Orb,\ NS,\ Newt}+H_{\rm Orb,\ NS,\ 1PN}+H_{\rm Orb,\ NS,\ 2PN}$](#uptotwopn): Contributions up to and including second post-Newtonian order
# 1. Part 2: [$H_{\rm Orb,\ NS,\ 3PN}$](#threepn): Third post-Newtonian contribution
# 1. Part 3: [Validation against second transcription and corresponding Python module](#code_validation)
# 1. Part 4: [LaTeX PDF output](#latex_pdf_output): $\LaTeX$ PDF Output
# <a id='uptotwopn'></a>
#
# # Part 1: $H_{\rm Orb,\ NS,\ Newt}+H_{\rm Orb,\ NS,\ 1PN}+H_{\rm Orb,\ NS,\ 2PN}$ \[Back to [top](#toc)\]
# $$\label{uptotwopn}$$
#
# As detailed in [Buonanno, Chen, and Damour (2006)](https://arxiv.org/abs/gr-qc/0508067) (henceforth BCD2006),
# this model assumes two point masses of mass $m_1$ and $m_2$ with corresponding momentum vectors $\mathbf{P}_1$ and $\mathbf{P}_2$, and displacement vectors $\mathbf{X}_1$ and $\mathbf{X}_2$ with respect to the center of mass.
#
# Following [BCD2006](https://arxiv.org/abs/gr-qc/0508067), we define the following quantities
#
# \begin{align}
# \mu &= m_1 m_2 / (m_1+m_2)\\
# \eta &= m_1 m_2 / (m_1+m_2)^2\\
# \mathbf{p} &= \mathbf{P}_1/\mu = -\mathbf{P}_2/\mu\\
# \mathbf{q} &= (\mathbf{X}_1-\mathbf{X}_2)/M\\
# q &= |\mathbf{q}|\\
# \mathbf{n} &= \frac{\mathbf{q}}{q}
# \end{align}
#
# Then the Hamiltonian up to and including second PN order is given by (to reduce possibility of copying error, these equations are taken directly from Eqs 2.2-4 of the LaTeX source code of [BCD2006](https://arxiv.org/abs/gr-qc/0508067), and only mildly formatted to (1) improve presentation in Jupyter notebooks and (2) to ensure some degree of consistency in notation across different terms in other Hamiltonian notebooks):
#
# \begin{align}
# H_{\rm Newt}\left({\bf q},{\bf p}\right) &= \mu \left[\frac{{\bf p}^2}{2} -
# \frac{1}{q}\right]\,, \\
# H_{\rm 1PN}\left({\bf q},{\bf p}\right) &= \mu\left[\frac{1}{8}(3\eta-1)({\bf p}^2)^2
# - \frac{1}{2}\left[(3+\eta){\bf p}^2+\eta({\bf n}\cdot{\bf p})^2\right]\frac{1}{q} + \frac{1}{2q^2}\right]\,,\\
# H_{\rm 2PN}\left({\bf q},{\bf p}\right)
# &= \mu\left[\frac{1}{16}\left(1-5\eta+5\eta^2\right)({\bf p}^2)^3
# + \frac{1}{8} \left[
# \left(5-20\eta-3\eta^2\right)({\bf p}^2)^2-2\eta^2({\bf n}\cdot{\bf p})^2{\bf p}^2-3\eta^2({\bf n}\cdot{\bf p})^4 \right]\frac{1}{q}\right.
# \\
# &\quad\quad\quad \left.+ \frac{1}{2} \left[(5+8\eta){\bf p}^2+3\eta({\bf n}\cdot{\bf p})^2\right]\frac{1}{q^2}
# - \frac{1}{4}(1+3\eta)\frac{1}{q^3}\right]\,,
# \end{align}
# +
# Step 0: Add NRPy's directory to the path
# https://stackoverflow.com/questions/16780014/import-file-from-parent-directory
import sympy as sp # SymPy: The Python computer algebra package upon which NRPy+ depends
import indexedexpNRPyPN as ixp # NRPy+: Symbolic indexed expression (e.g., tensors, vectors, etc.) support
from NRPyPN_shortcuts import div,dot # NRPyPN: shortcuts for e.g., vector operations
def f_H_Newt__H_NS_1PN__H_NS_2PN(m1,m2, PU, nU, q):
mu = m1*m2 / (m1+m2)
eta = m1*m2 / (m1+m2)**2
pU = ixp.zerorank1()
for i in range(3):
pU[i] = PU[i]/mu
global H_Newt, H_NS_1PN, H_NS_2PN
H_Newt = mu*(+div(1,2)*dot(pU,pU) - 1/q)
H_NS_1PN = mu*(+div(1,8)*(3*eta-1)*dot(pU,pU)**2
-div(1,2)*((3+eta)*dot(pU,pU) + eta*dot(nU,pU)**2)/q
+div(1,2)/q**2)
H_NS_2PN = mu*(+div(1,16)*(1 - 5*eta + 5*eta**2)*dot(pU,pU)**3
+div(1,8)*(+(5 - 20*eta - 3*eta**2)*dot(pU,pU)**2
-2*eta**2*dot(nU,pU)**2*dot(pU,pU)
-3*eta**2*dot(nU,pU)**4)/q
+div(1,2)*((5+8*eta)*dot(pU,pU) + 3*eta*dot(nU,pU)**2)/q**2
-div(1,4)*(1+3*eta)/q**3)
# -
# Second version. This one was mostly a search+replace version
# of the original TeX'ed up equations in the paper.
# Used for validation purposes only.
def f_H_Newt__H_NS_1PN__H_NS_2PNv2(m1,m2, PU, nU, q):
mu = m1*m2/(m1+m2)
eta = m1*m2/(m1+m2)**2
pU = ixp.zerorank1()
for i in range(3):
pU[i] = PU[i]/mu
p_dot_p = dot(pU,pU)
n_dot_p = dot(nU,pU)
# H_{\rm Newt} = \frac{p^i p^i}{2} - \frac{1}{q}
global H_Newtv2, H_NS_1PNv2, H_NS_2PNv2
H_Newtv2 = mu*(div(1,2)*p_dot_p - 1/q)
H_NS_1PNv2 = mu*(div(1,8)*(3*eta-1)*p_dot_p**2 - \
div(1,2)*((3+eta)*p_dot_p + eta*n_dot_p**2)/q + 1/(2*q**2))
H_NS_2PNv2 = mu*(div(1,16)*(1 - 5*eta + 5*eta**2)*p_dot_p**3 +
div(1,8)*((5 - 20*eta - 3*eta**2)*p_dot_p**2
- 2*eta**2*n_dot_p**2*p_dot_p - 3*eta**2*n_dot_p**4)/q +
div(1,2)*((5 + 8*eta)*p_dot_p + 3*eta*n_dot_p**2)/q**2 -
div(1,4)*(1 + 3*eta)/q**3)
# <a id='threepn'></a>
#
# # Part 2: $H_{\rm Orb,\ NS,\ 3PN}$ Third post-Newtonian contribution \[Back to [top](#toc)\]
# $$\label{threepn}$$
#
# To reduce possibility of copying error, equations are taken directly from the LaTeX source code of Eqs 2.2-4 in [BCD2006](https://arxiv.org/abs/gr-qc/0508067), and only mildly formatted to (1) improve presentation in Jupyter notebooks and (2) to ensure some degree of consistency in notation across different terms in other Hamiltonian notebooks:
#
# \begin{align}
# H_{\rm 3PN}\left({\bf q},{\bf p}\right)
# &= \mu\left\{\frac{1}{128}\left(-5+35\eta-70\eta^2+35\eta^3\right)({\bf p}^2)^4\right.
# \\
# &\quad\quad + \frac{1}{16}\left[
# \left(-7+42\eta-53\eta^2-5\eta^3\right)({\bf p}^2)^3
# + (2-3\eta)\eta^2({\bf n}\cdot{\bf p})^2({\bf p}^2)^2
# + 3(1-\eta)\eta^2({\bf n}\cdot{\bf p})^4{\bf p}^2 - 5\eta^3({\bf n}\cdot{\bf p})^6
# \right]\frac{1}{q}
# \\
# &\quad\quad +\left[ \frac{1}{16}\left(-27+136\eta+109\eta^2\right)({\bf p}^2)^2
# + \frac{1}{16}(17+30\eta)\eta({\bf n}\cdot{\bf p})^2{\bf p}^2 + \frac{1}{12}(5+43\eta)\eta({\bf n}\cdot{\bf p})^4
# \right]\frac{1}{q^2} \\
# &\quad\quad +\left\{ \left[ -\frac{25}{8} + \left(\frac{1}{64}\pi^2-\frac{335}{48}\right)\eta
# - \frac{23}{8}\eta^2 \right]{\bf p}^2
# + \left(-\frac{85}{16}-\frac{3}{64}\pi^2-\frac{7}{4}\eta\right)\eta({\bf n}\cdot{\bf p})^2
# \right\}\frac{1}{q^3}
# \\
# &\quad\quad\left. + \left[ \frac{1}{8} + \left(\frac{109}{12}-\frac{21}{32}\pi^2\right)\eta
# \right]\frac{1}{q^4}\right\}\,,
# \end{align}
def f_H_NS_3PN(m1,m2, PU, nU, q):
mu = m1*m2 / (m1+m2)
eta = m1*m2 / (m1+m2)**2
pU = ixp.zerorank1()
for i in range(3):
pU[i] = PU[i]/mu
global H_NS_3PN
H_NS_3PN = mu*(+div(1,128)*(-5 + 35*eta - 70*eta**2 + 35*eta**3)*dot(pU,pU)**4
+div(1, 16)*(+(-7 + 42*eta - 53*eta**2 - 5*eta**3)*dot(pU,pU)**3
+(2-3*eta)*eta**2*dot(nU,pU)**2*dot(pU,pU)**2
+3*(1-eta)*eta**2*dot(nU,pU)**4*dot(pU,pU) - 5*eta**3*dot(nU,pU)**6)/q
+(+div(1,16)*(-27 + 136*eta + 109*eta**2)*dot(pU,pU)**2
+div(1,16)*(+17 + 30*eta)*eta*dot(nU,pU)**2*dot(pU,pU)
+div(1,12)*(+ 5 + 43*eta)*eta*dot(nU,pU)**4)/q**2
+(+(-div(25, 8) + (div(1,64)*sp.pi**2 - div(335,48))*eta - div(23,8)*eta**2)*dot(pU,pU)
+(-div(85,16) - div(3,64)*sp.pi**2 - div(7,4)*eta)*eta*dot(nU,pU)**2)/q**3
+(+div(1,8)+(div(109,12) - div(21,32)*sp.pi**2)*eta)/q**4)
# Second version. This one was mostly a search+replace version
# of the original TeX'ed up equations in the paper.
# Used for validation purposes only.
def f_H_NS_3PNv2(m1,m2, pU, nU, q):
mu = m1*m2/(m1+m2)
eta = m1*m2/(m1+m2)**2
PU = ixp.zerorank1()
for i in range(3):
PU[i] = pU[i]/mu
P_dot_P = dot(PU,PU)
n_dot_P = dot(nU,PU)
global H_NS_3PNv2
# The following is simply by-hand search/replaced from the above LaTeX to minimize error
H_NS_3PNv2 = \
mu*( div(1,128)*(-5+35*eta-70*eta**2+35*eta**3)*P_dot_P**4 +
div(1,16)* ( (-7+42*eta-53*eta**2-5*eta**3)*P_dot_P**3
+(2-3*eta)*eta**2*n_dot_P**2*P_dot_P**2 +
+3*(1-eta)*eta**2*n_dot_P**4*P_dot_P - 5*eta**3*n_dot_P**6 )/(q) +
( div(1,16)*(-27+136*eta+109*eta**2)*P_dot_P**2
+ div(1,16)*(17+30*eta)*eta*n_dot_P**2*P_dot_P + div(1,12)*(5+43*eta)*eta*n_dot_P**4)/(q**2) +
( ( -div(25,8) + (div(1,64)*sp.pi**2-div(335,48))*eta
- div(23,8)*eta**2 )*P_dot_P
+ (-div(85,16)-div(3,64)*sp.pi**2-div(7,4)*eta)*eta*n_dot_P**2)/(q**3) +
( div(1,8) + (div(109,12)-div(21,32)*sp.pi**2)*eta)/(q**4) )
# <a id='code_validation'></a>
#
# # Part 3: Validation against second transcription and corresponding Python module \[Back to [top](#toc)\]
# $$\label{code_validation}$$
#
# As a code validation check, we verify agreement between
# * the SymPy expressions transcribed from the cited published work on two separate occasions, and
# * the SymPy expressions generated in this notebook, and the corresponding Python module.
# +
from NRPyPN_shortcuts import m1,m2,pU,nU,q # NRPyPN: import needed input variables.
f_H_Newt__H_NS_1PN__H_NS_2PN(m1,m2, pU, nU, q)
f_H_NS_3PN(m1,m2, pU, nU, q)
def error(varname):
print("ERROR: When comparing Python module & notebook, "+varname+" was found not to match.")
sys.exit(1)
# Validation against second transcription of the expressions.
f_H_Newt__H_NS_1PN__H_NS_2PNv2(m1,m2, pU, nU, q)
f_H_NS_3PNv2(m1,m2, pU, nU, q)
if sp.simplify(H_Newt - H_Newtv2) != 0: error("H_Newtv2")
if sp.simplify(H_NS_1PN - H_NS_1PNv2) != 0: error("H_NS_1PNv2")
if sp.simplify(H_NS_2PN - H_NS_2PNv2) != 0: error("H_NS_2PNv2")
if sp.simplify(H_NS_3PN - H_NS_3PNv2) != 0: error("H_NS_3PNv2")
# Validation against corresponding Python module
import PN_Hamiltonian_NS as HNS
HNS.f_H_Newt__H_NS_1PN__H_NS_2PN(m1,m2, pU, nU, q)
HNS.f_H_NS_3PN(m1,m2, pU, nU, q)
if sp.simplify(H_Newt - HNS.H_Newt) != 0: error("H_Newt")
if sp.simplify(H_NS_1PN - HNS.H_NS_1PN) != 0: error("H_NS_1PN")
if sp.simplify(H_NS_2PN - HNS.H_NS_2PN) != 0: error("H_NS_2PN")
if sp.simplify(H_NS_3PN - HNS.H_NS_3PN) != 0: error("H_NS_3PN")
print("ALL TESTS PASS")
# -
# <a id='latex_pdf_output'></a>
#
# # Step 4: Output this notebook to $\LaTeX$-formatted PDF file \[Back to [top](#toc)\]
# $$\label{latex_pdf_output}$$
#
# The following code cell converts this Jupyter notebook into a proper, clickable $\LaTeX$-formatted PDF file. After the cell is successfully run, the generated PDF may be found in the root NRPy+ tutorial directory, with filename
# [PN-Hamiltonian-Nonspinning.pdf](PN-Hamiltonian-Nonspinning.pdf) (Note that clicking on this link may not work; you may need to open the PDF file through another means.)
import os,sys # Standard Python modules for multiplatform OS-level functions
import cmdline_helperNRPyPN as cmd # NRPy+: Multi-platform Python command-line interface
cmd.output_Jupyter_notebook_to_LaTeXed_PDF("PN-Hamiltonian-Nonspinning",location_of_template_file=os.path.join(".."))
| NRPyPN/PN-Hamiltonian-Nonspinning.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# +
###### #Following instructions in DataCamp course:
#https://campus.datacamp.com/courses/generalized-linear-models-in-python/modeling-binary-data?ex=15
#https://towardsdatascience.com/a-quick-guide-on-descriptive-statistics-using-pandas-and-seaborn-2aadc7395f32
#https://github.com/VIS-SIG/Wonderful-Wednesdays/tree/master/data/2020/2020-12-09
#Import libraries
import statsmodels.api as sm #Array based model
from statsmodels.formula.api import glm
import numpy as np
import matplotlib.pyplot as plt
#from lifelines import KaplanMeierFitter
from sklearn.ensemble import RandomForestClassifier
# -
import seaborn as sns
import pandas as pd
import os
print(os.getcwd()) #/Users/zahraSari
# +
cwd=os.chdir('/Users/zahraSari/Desktop/')
#Change directory
Files = os.listdir(cwd)
# +
#Data from Github :
#https://github.com/VIS-SIG/Wonderful-Wednesdays/blob/master/data/2020/2020-12-09/Reexcision.csv
data= pd.read_csv('Book.csv')
print(data)
#age
#tumorsize
#histology (hist; 0: others, 1: Invasive-duct./ductal-lob.)
#Multifocality (mult.foc; 0: no, 1: yes)
#Accomp. in situ (acc.in.situ; 0: others, 1: DCIS and LCIS)
#Lymphovascular invasion (lymph.inv; 0: no, 1: yes)
#Estrogen-receptor (estr.rec; 0: no, 1: yes)
#Progesterone-receptor (prog.rec; 0: no, 1: yes)
# +
print(data.columns.tolist()) #View the column names
# -
data.info() # Tumor size has 9 unknown variables
data.mean()
total_rows=len(data.axes[0])
total_cols=len(data.axes[1])
print("Number of Rows: "+str(total_rows))
print("Number of Columns: "+str(total_cols))
# +
data['tumorsize']= pd.to_numeric(data['tumorsize'])
# -
data=data.dropna() #Removing the 9 unknown variables from Data Frame
data.mean()
# +
# Plot the age variable
sns.distplot(data['age'])
# plt.axvline(np.median(data['age']),color='b', linestyle='--')
plt.axvline(np.mean(data['age']),color='b', linestyle='-')
#Display the plot
plt.title('Age Distrubtion, Mean = 56.74')
plt.show()
data.age.mean()
# +
# Plot first variable
sns.distplot(data['tumorsize'])
# plt.axvline(np.median(data['tumorsize']),color='b', linestyle='--')
plt.axvline(np.mean(data['tumorsize']),color='b', linestyle='-')
# Display the
plt.title('Tumor Size Distrubtion, Mean = 23.14')
plt.show()
data.tumorsize.mean()
# +
#Interpretation: People with re-exision
g = sns.FacetGrid(data, col='RE')
g.map(plt.hist, 'tumorsize', bins=20)
# Adjust title and axis labels directly
g.set_axis_labels(x_var="Tumor Size", y_var="Tumor Size Distribution by RE")
# Clearly ones with RE=1 have smaller tumorsize
# +
g = sns.FacetGrid(data, col='RE')
g.map(plt.hist, 'age', bins=20)
# Adjust title and axis labels directly
g.set_axis_labels(x_var="Age", y_var="Age Distribution by RE")
#RE=1 has younger age distribution
# -
# # Pivoting Features
data['hist'].value_counts().sort_values()
# +
sns.countplot(x='hist',data=data,palette='hls')
plt.show()
#
# -
sns.countplot(x='RE',data=data,palette='hls')
plt.title('Re-excision Frequency')
plt.show()
# +
# Summary Chart Re-excision
ax = ((100 *data["RE"].value_counts() / len(data))).plot.bar(rot=0)
ax.set( ylabel="%", title="Re-excision Percentage")
plt.show()
# Summary Chart hist
ax = ((100 *data["hist"].value_counts() / len(data))).plot.bar(rot=0)
ax.set( ylabel="%", title="Histology Percentage")
plt.show()
# Summary Chart multfoc
ax = ((100 *data["multfoc"].value_counts() / len(data))).plot.bar(rot=0)
ax.set( ylabel="%", title="Multfoc Percentage")
plt.show()
# Summary Chart accinsitu
ax = ((100 *data["accinsitu"].value_counts() / len(data))).plot.bar(rot=0)
ax.set( ylabel="%", title="Accomp. in situ Percentage")
plt.show()
# +
data.groupby(['RE']).mean() #Mean of variables for RE of 0 or 1
# +
#Interpretation: Age distribution for patinets who had Re-excision is lower than those with
# %matplotlib inline
sns.set(style="whitegrid")
plt.figure(figsize=(10,8))
plt.title('Boxplot of Age by Re-excision (0 vs. 1)')
ax = sns.boxplot( x='RE',y='age', data=data, orient="v")
plt.xlabel('Re-excision')
plt.ylabel('Age')
# +
# %matplotlib inline
sns.set(style="whitegrid")
plt.figure(figsize=(10,8))
ax = sns.boxplot(y='tumorsize' , x='RE', data=data, orient="v")
plt.title('Boxplot of Tumor Size by Re-excision (0 vs. 1)')
plt.xlabel('Re-excision')
plt.ylabel('Tumor Size')
#The box plot shows you how a feature's values spread out for each class.
#It's a compact representation of the distribution, showing the extreme high value,
#upper quartile, median, lower quartile and extreme low value.
# -
# # Correlating categorical features
# +
grid = sns.FacetGrid(data, row='accinsitu', size=2.2, aspect=1.6)
grid.map(sns.pointplot, 'hist', 'RE' , 'lymphinv', palette='deep')
grid.add_legend()
# -
print(data.corr)
# # Correlation
# +
####Correlation Plot#######
corr=data.corr()
# Generate a mask for the upper triangle
mask = np.zeros_like(corr, dtype=np.bool)
mask[np.triu_indices_from(mask)] = True
# Set up the matplotlib figure
f, ax = plt.subplots(figsize=(15, 13))
# Generate a custom diverging colormap
cmap = sns.diverging_palette(220, 10, as_cmap=True)
# Draw the heatmap with the mask and correct aspect ratio
sns.heatmap(corr, mask=mask, cmap=cmap, vmax=.3, center=0,
square=True, linewidths=.5, cbar_kws={"shrink": .5})
plt.title('Correlation Plot')
# -
# # Statistical Model - Logistic Model
# +
#########Statistical Model############
#Fit logistic regression model
#Logistic regression is an improved version of linear regression.
# +
model = sm.GLM.from_formula("RE ~ hist + age + tumorsize + hist + multfoc + accinsitu + lymphinv + estrrec + progrec ", family = sm.families.Binomial(), data=data)
result = model.fit()
result.summary()
#Based on p-value being less than 0.05,
#Significant variables are: hist, tumorsize, accinsitu, lymphinv
#Age is very close to 0.05 so speculuative whether it is significant,
#Similarly for Intercept, p-value is very close to 0.05 but doesn't pass
#Coef for hist: (Thinking of linear regression formula Y = AX + B)
#where A= -1.2014 , B is 0 since it is non-significant.
#If a person’s hist is 1 unit more s/he will have a 0.052 (coefficient with age in the table above) unit more
#chance of having heart disease based on the p-value in the table.
#Generally, positive coefficients indicate that the event becomes more likely as the predictor increases.
#Negative coefficients indicate that the event becomes less likely as the predictor increases.
# +
#Removing non-significant variables and re-fitting the model
#Age seems to be significant now
model = sm.GLM.from_formula("RE ~ hist + age + tumorsize + accinsitu + lymphinv -1 ", family = sm.families.Binomial(), data=data)
result = model.fit()
result.summary()
#coeffiecnt -1.2849 for hist shows increase of odds
#for ones with hist=1 than ones with hist=0
#According to this fitted model, older people are more
#likely to have Reexicision than younger people. The
#log odds for heart disease increases by 0.0545 units for each year.
#If a person is 10 years older his or her chance of having RE
#increases by 0.0545 * 10 = 0.545 units.
# +
data[['RE','hist', 'age' , 'tumorsize' , 'accinsitu' , 'lymphinv']].corr()
# -
# # Visualization of the Fitted Model
# +
#https://towardsdatascience.com/logistic-regression-model-fitting-and-finding-the-correlation-p-value-z-score-confidence-8330fb86db19
#With help from this site
# -
from statsmodels.sandbox.predict_functional import predict_functional
values = {"hist": 1, "tumorsize": 23, "accinsitu":0 , "lymphinv" :0 }
pr, cb, fv = predict_functional(result, "age", values=values, ci_method="simultaneous")
# +
ax = sns.lineplot(fv, pr, lw=4)
ax.fill_between(fv, cb[:, 0], cb[:, 1], color='grey', alpha=0.4)
ax.set_xlabel("Age")
ax.set_ylabel("Re-excision")
ax.set_title('Fitted Model: Log-odds of Re-excision by Age for patients with hist=1, tumorsize=23,accinsitu=0 ,lymphinv=0 ')
#This plot of fitted log-odds visualizes the effect of age on reexcision for
#hist=0, tumorsize=23, accinsitu=0 and lumphinv=0 by the glm fitted model
#Slight negative correlation of age and RE are visible in this plot
#For the specific described variables
# -
# +
from statsmodels.sandbox.predict_functional import predict_functional
values = {"hist": 1, "age": 57, "accinsitu":0 , "lymphinv" :0 }
pr, cb, fv = predict_functional(result, "tumorsize", values=values, ci_method="simultaneous")
ax = sns.lineplot(fv, pr, lw=4)
ax.fill_between(fv, cb[:, 0], cb[:, 1], color='grey', alpha=0.4)
ax.set_xlabel("Tumor Size")
ax.set_ylabel("Re-excision")
ax.set_title('Fitted Model: Log-odds of Re-excision by Tumorsize for patients with hist=1,age=57,accinsitu=0,lymphinv=0')
#This plot of fitted log-odds visualizes the effect of tumorsize on reexcision for
#hist=0, age=45, accinsitu=0 and lumphinv=0 by the glm fitted model
#Clear Positive correlation of tumorsize and RE are visible in this plot
# -
# +
import seaborn as sns
#Plot the relationship between two variables in a DataFrame and
#add overlay with the logistic fit
sns.regplot(x = 'tumorsize', y = 'RE',
y_jitter = 0.03,
data = data,
logistic = True,
ci = 95)
plt.title('Fitted Plot for Re-excision by Tumorsize')
plt.xlabel('Tumor Size')
plt.ylabel('Re-excision')
# Display the plot
plt.show()
#Interpretation: the lower tumor sizes are associated with value 0 for Reexcision,
#higher values of tumorsize.
#Tumor sizes of over 55 are associated with value of 1 for Reexicision.
#The confidence interval gets wider as the value of the predictor increases. The
#wide interval is partly due to the small amount of data for larger tumor size.
# +
#Plot the relationship between two variables in a DataFrame and add overlay with the logistic fit
sns.regplot(x = 'age', y = 'RE',
y_jitter = 0.03,
data = data,
logistic = True,
ci = 95)
plt.title('Fitted Plot for Re-excision by Age')
plt.xlabel('Age')
plt.ylabel('Re-excision')
# Display the plot
plt.show()
#Interpretation: the lower values age is associated with value 0 for Reexcision, higher values of tumorsize
#eg. tumor size of over 55 are associated with value of 1 for Reexicision.
#The confidence interval gets wider as the value of the predictor increases. The
#wide interval is partly due to the small amount of data for larger lower and higher ages.
# +
# Examine the first 5 computed predictions
print(data[['RE', 'hist', 'multfoc', 'accinsitu', 'lymphinv', 'estrrec', 'progrec']].head())
# +
# Define the cutoff
cutoff = 0.5
# Compute class predictions: y_prediction
y_prediction = np.where(prediction > cutoff, 1, 0)
# -
# # Machine Learning
# +
#Random Forest
# +
from sklearn.model_selection import train_test_split
# +
y=data['RE']
# +
train_df = data.drop(['RE'], axis=1)
# +
import numpy as np
from sklearn.model_selection import train_test_split
X, y = np.arange(10).reshape((5, 2)), range(5)
X
list(y)
# -
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.33, random_state=42)
X_train
y_train
X_test
y_test
# +
#print("Before", train_df.shape, test_df.shape, combine[0].shape, combine[1].shape)
#train_df = train_df.drop(['Ticket', 'Cabin'], axis=1)
#test_df = test_df.drop(['Ticket', 'Cabin'], axis=1)
#combine = [train_df, test_df]
#"After", train_df.shape, test_df.shape, combine[0].shape, combine[1].shape
# +
train_df = data.drop(['RE'], axis=1)
# +
df = pd.DataFrame(np.random.randn(100, 2))
msk = np.random.rand(len(df)) < 0.8
train = df[msk]
test = df[~msk]
len(test)
len(train)
# +
#####################DATACAMP#################
# Import train_test_split function
#from sklearn.model_selection import train_test_split
#X=data[['sepal length', 'sepal width', 'petal length', 'petal width']] # Features
#y=data['species'] # Labels
# Split dataset into training set and test set
#X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3) # 70% training and 30% test
# Import train_test_split function
from sklearn.model_selection import train_test_split
X=data[['age' , 'tumorsize' , 'hist', 'multfoc' , 'accinsitu', 'lymphinv', 'estrrec' , 'progrec']] # Features
y=data['RE'] # Labels
# Split dataset into training set and test set
X_train, X_test, y_train, y_test = train_test_split(X, y, test_size=0.3) # 70% training and 30% test
# -
# +
#Import Random Forest Model
from sklearn.ensemble import RandomForestClassifier
#Create a Gaussian Classifier
clf=RandomForestClassifier(n_estimators=100)
#Train the model using the training sets y_pred=clf.predict(X_test)
clf.fit(X_train,y_train)
y_pred=clf.predict(X_test)
# +
#Import scikit-learn metrics module for accuracy calculation
from sklearn import metrics
# Model Accuracy, how often is the classifier correct?
print("Accuracy:",metrics.accuracy_score(y_test, y_pred))
# +
RandomForestClassifier(bootstrap=True, class_weight=None, criterion='gini',
max_depth=None, max_features='auto', max_leaf_nodes=None,
min_impurity_decrease=0.0, min_impurity_split=None,
min_samples_leaf=1, min_samples_split=2,
min_weight_fraction_leaf=0.0, n_estimators=100, n_jobs=1,
oob_score=False, random_state=None, verbose=0,
warm_start=False)
# -
| posts/2021-01-11-wonderful-wednesdays-january-2021/code/Residual Tumor - Zara Sari.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python [Root]
# language: python
# name: Python [Root]
# ---
# # Machine Learning with Iris flower data set & Python's sckit-learn
#
#
# The [Iris flower dataset](https://en.wikipedia.org/wiki/Iris_flower_data_set) is composed of 4 variables (aka features) & 1 label (type of flower).
#
# ## 1. Importing dataset
from sklearn.datasets import load_iris
iris = load_iris()
# let's check the names of the feature & target (ie; label)
print(iris.feature_names)
print(iris.target_names)
# We can see that we have 4 features & 3 target labels:
len(iris.feature_names), len(iris.target_names)
print(iris.data[0])
# If we look at the target values we see that there are 3 digits 0, 1, & 2:
print(iris.target)
# We know that there are three labels but which is which? Checking the dataset at Wikipedia tells us that the first entry is a 'I. setosa' (hence label 0). Similarly, 1 represents 'I. versicolor' and 2 represents 'I. virginica'.
# ## 2. Classified training
# We now perform the usual practice of splitting the dataset into two parts, a training dataset usually 25-30%, and a testing dataset usually 65-70% of the original dataset. We'll go wih 30% and 70%.
import numpy as np
from sklearn import tree
# The dataset as you can see on Wikipedia is composed of 50 observations per label. We extract this from the dataset and assign it to `nl`:
nl = len(iris.target[iris.target == 0]) # 50
# +
n = int(nl * 0.3); # number of observations to use for test data (30% of 50)
# indices of observations to use for test dataset
test_indx = list( range(0,n) ) + list( range(50,50+n) ) + list( range(100,100+n) )
# -
# We now delete the observations corresponding to these indices from each of the target and data columns and assign the result to the training datasets.
# training data
train_target = np.delete(iris.target, test_indx)
train_data = np.delete(iris.data, test_indx, axis = 0)
# To form the testing data we need the obervations corresponding to those indice:
# testing data
test_target = iris.target[test_indx]
test_data = iris.data[test_indx]
# Next we train the classifier:
clf = tree.DecisionTreeClassifier() # clf for short
clf.fit(train_data, train_target)
# ## 3. Label prediction of test data
# We now feed the test data (predictors) into the decision tree and get back the (predicted) test labels:
clf.predict(test_data)
# If we look at the original test labels we find they are the same:
test_target
# An easier way to check is with a conditional statment which shows us that the predicted and original labels are the same:
clf.predict(test_data) == test_target
# ## 4. Visualizing the decision tree
# Decision trees are very useful in making decisions and best of all is that they can be visualized showing us how the classifier works.
# This requires `pydot` and `Graphviz` to be installed. With Anaconda under Ubuntu:
# +
# in terminal:
# conda install pydot
# sudo apt-get install graphviz
# +
# %matplotlib inline
# code borrowed scikit learn website
from sklearn.externals.six import StringIO
import pydot
dot_data = StringIO()
tree.export_graphviz(clf, out_file=dot_data)
graph = pydot.dot_parser.parse_dot_data(dot_data.getvalue())
# graph.write_pdf("iris.pdf")
from IPython.display import Image
dot_data = StringIO()
tree.export_graphviz(clf, out_file=dot_data,
feature_names=iris.feature_names,
class_names=iris.target_names,
filled=True, rounded=True,
special_characters=True)
# graph = pydot.graph_from_dot_data(dot_data.getvalue())
# graph = pydot.dot_parser.parse_dot_data(dot_data.getvalue())
# Image(graph.create_png())
graph = pydot.graph_from_dot_data(dot_data.getvalue())
Image(graph.)
# +
from sklearn import tree
X = [[0, 0], [1, 1]]
Y = [0, 1]
clf = tree.DecisionTreeClassifier()
clf = clf.fit(X, Y)
# After being fitted, the model can then be used to predict the class of samples:
clf.predict([[2., 2.]])
# Alternatively, the probability of each class can be predicted, which is the fraction of training samples of the same class in a leaf:
clf.predict_proba([[2., 2.]])
# DecisionTreeClassifier is capable of both binary (where the labels are [-1, 1]) classification and multiclass (where the labels are [0, ..., K-1]) classification.
# Using the Iris dataset, we can construct a tree as follows:
from sklearn.datasets import load_iris
from sklearn import tree
iris = load_iris()
clf = tree.DecisionTreeClassifier()
clf = clf.fit(iris.data, iris.target)
# Once trained, we can export the tree in Graphviz format using the export_graphviz exporter. Below is an example export of a tree trained on the entire iris dataset:
with open("iris.dot", 'w') as f:
f = tree.export_graphviz(clf, out_file=f)
# Then we can use Graphviz’s dot tool to create a PDF file (or any other supported file type): dot -Tpdf iris.dot -o iris.pdf.
import os
os.unlink('iris.dot')
# # Alternatively, if we have Python module pydotplus installed, we can generate a PDF file (or any other supported file type) directly in Python:
# import pydotplus
# dot_data = tree.export_graphviz(clf, out_file=None)
# graph = pydotplus.graph_from_dot_data(dot_data)
# graph.write_pdf("iris.pdf")
# # The export_graphviz exporter also supports a variety of aesthetic options, including coloring nodes by their class (or value for regression) and using explicit variable and class names if desired. IPython notebooks can also render these plots inline using the Image() function:
# from IPython.display import Image
# dot_data = tree.export_graphviz(clf, out_file=None,
# feature_names=iris.feature_names,
# class_names=iris.target_names,
# filled=True, rounded=True,
# special_characters=True)
# graph = pydotplus.graph_from_dot_data(dot_data)
# Image(graph.create_png())
# -
# +
# code borrowed scikit learn website
from sklearn.externals.six import StringIO
import pydot
dot_data = StringIO()
tree.export_graphviz(clf, out_file=dot_data)
graph = pydot.graph_from_dot_data(dot_data.getvalue())
graph.write_pdf("iris.pdf")
from IPython.display import Image
dot_data = StringIO()
tree.export_graphviz(clf, out_file=dot_data,
feature_names=iris.feature_names,
class_names=iris.target_names,
filled=True, rounded=True,
special_characters=True)
graph = pydot.graph_from_dot_data(dot_data.getvalue())
Image(graph.create_png())
# -
# ## How does a decision tree work?
# In this case, it starts by checking if the petal width is less than or equal to 0.8.If it is the tree predicts it belongs to the `setosa` class. Otherwise, it asks if the petal length is less than or equal to 4.75. If it is it classifies it as a `versicolor`, otherwise a `virginica`. .
# This is summarized in the first three levels of the above decision tree.
# ### Examples
# * For the first prediction we see that it was assigned as a setosa (label 0) since the petal width is 0.2 < 0.8.
print(iris.feature_names, iris.target_names)
print(test_data[0], test_target[0])
# * For the 20th prediction it was assigned as a `versicolor` since the petal width is 1.3 > 0.8 (hence not a `setosa`) and the petal length is 4.5 < 4.75.
print(iris.feature_names, iris.target_names)
print(test_data[20], test_target[20])
# * For the 40th prediction it was assigned as a `virginica` since the petal width is 2.0 > 0.8 (hence not a `setosa`) and the petal length is 5.1 > 4.75. At this point it could be either a veriscolor or a virginica. The decision tree keeps going through these branches until it reaches a decision and hence a prediction. I leave the rest to you for this case.
print(iris.feature_names, iris.target_names)
print(test_data[40], test_target[40])
# ## Notes
# * Feature selection (and engineering) is vital in decision trees and machine learning in general.
# ## Resources
# * https://en.wikipedia.org/wiki/Decision_tree_learning
# * https://en.wikipedia.org/wiki/Predictive_analytics
| notebooks/Decision trees with Iris dataset with Scikit learn.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
#hide
from fastai.basics import *
# # The fastai book
#
# > How to get started with the fastai book.
# This information is for readers of the Early Access version of [Deep Learning for Coders with fastai and PyTorch](https://www.amazon.com/Deep-Learning-Coders-fastai-PyTorch/dp/1492045527) and of the fastai [draft notebooks](https://github.com/fastai/fastbook).
#
# Note that these require you to use fastai v2, which is currently in pre-release. During pre-release, this module is called `fastai`. The draft notebooks contain the correct imports for `fastai`, but the PDF book does not - it uses `fastai`. Therefore, you should change `fastai` to `fastai` in all import statements.
# ## Install
# Clone the [draft notebooks repo](https://github.com/fastai/fastbook), and then from that directory:
#
# pip install -r requirements.txt
#
# It is best to first install fastai v1 to ensure you have all the correct dependencies. See [the docs](https://docs.fast.ai/) for details.
# ## How to use
# To get started, run Jupyter Notebook (or use one of the online Jupyter platforms listed at https://course.fast.ai) and click on the *app_jupyter.ipynb* notebook.
#
# We'll be adding more information to this site as the official book release in July 2020 gets closer.
| nbs/index.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
import matplotlib.pyplot as plt
import numpy as np
import pandas as pd
lb=["I01","I02","I03","I04","I05","I06","I07","I08","I09","I10","I11","I12","I13","I14","I15","I16","I17","I18"]
# plt.figure(figsize=(12,7))
# plt.boxplot([t1.IO1,t1.IO2,t1.IO3,t1.IO4,t1.IO5,t1.IO6,t1.IO7,t1.IO8,t1.IO9,t1.I10,t1.I11,t1.I12,t1.I13,t1.I14,t1.I15,t1.I16,t1.I17,t1.I18],0,"gD",labels=lb,)
# plt.xlabel("Indices")
# plt.ylabel("T1",rotation=0)
# plt.savefig("t1GBMJumpSqueeze",fExpansionormat="png")
# +
x=pd.ExcelFile('/home/sharan/Desktop/Transport/IISER/ClusterOutput/Squeeze/GBM/t1FinalSqueeze.xlsx')
t1=x.parse(0)
y=pd.ExcelFile('/home/sharan/Desktop/Transport/IISER/ClusterOutput/Squeeze/Stat.xlsx')
page=y.parse(0)
t1star=np.array(page.Mean)
# t2star=np.append(t2star,t2star[len(t2star)-1])
plt.figure(figsize=(12,7))
plt.boxplot([t1.IO1,t1.IO2,t1.IO3,t1.IO4,t1.IO5,t1.IO6,t1.IO7,t1.IO8,t1.IO9,t1.I10,t1.I11,t1.I12,t1.I13,t1.I14,t1.I15,t1.I16,t1.I17,t1.I18],0," ",labels=lb)
plt.scatter(x=range(1,19),y=t1star,marker="D",c="g")
# print(len(t2star))
# print(len(lb))
plt.xlabel("Indices")
plt.ylabel("T1",rotation=0)
plt.savefig("t1GBMJumpSqueeze",format="png")
plt.show()
# +
x=pd.ExcelFile('/home/sharan/Desktop/Transport/IISER/ClusterOutput/Squeeze/GBM/t2FinalSqueeze.xlsx')
t2=x.parse(0)
# y=pd.ExcelFile('/home/thegodfather/Desktop/IISER/Codes&Data2/Squeeze/GBM/MainSqueezeMax.xlsx')
# page=y.parse(0)
t2star=np.array(page.SD)
# t2star=np.append(t2star,t2star[len(t2star)-1])
plt.figure(figsize=(12,7))
plt.boxplot([t2.IO1,t2.IO2,t2.IO3,t2.IO4,t2.IO5,t2.IO6,t2.IO7,t2.IO8,t2.IO9,t2.I10,t2.I11,t2.I12,t2.I13,t2.I14,t2.I15,t2.I16,t2.I17,t2.I18],0," ",labels=lb)
plt.scatter(x=range(1,19),y=t2star,marker="D",c="g")
# print(len(t2star))
# print(len(lb))
plt.xlabel("Indices")
plt.ylabel("T2",rotation=0)
plt.savefig("t2GBMJumpSqueeze",format="png")
plt.show()
# +
x=pd.ExcelFile('/home/sharan/Desktop/Transport/IISER/ClusterOutput/Squeeze/GBM/t3FinalSqueeze.xlsx')
t3=x.parse(0)
# y=pd.ExcelFile('/home/thegodfather/Desktop/IISER/Codes&Data2/Squeeze/GBM/MainSqueezeMax.xlsx')
# page=y.parse(0)
t3star=np.array(page.Skew)
plt.figure(figsize=(12,7))
plt.boxplot([t3.IO1,t3.IO2,t3.IO3,t3.IO4,t3.IO5,t3.IO6,t3.IO7,t3.IO8,t3.IO9,t3.I10,t3.I11,t3.I12,t3.I13,t3.I14,t3.I15,t3.I16,t3.I17,t3.I18],0," ",labels=lb)
plt.scatter(x=range(1,19),y=t3star,marker="D",c="g")
plt.xlabel("Indices")
plt.ylabel("T3",rotation=0)
# plt.ylabe(rotation=90)
plt.savefig("t3GBMJumpSqueeze",format="png")
plt.show()
plt.close()
# +
x=pd.ExcelFile('/home/sharan/Desktop/Transport/IISER/ClusterOutput/Squeeze/GBM/t4FinalSqueeze.xlsx')
t4=x.parse(0)
# y=pd.ExcelFile('/home/thegodfather/Desktop/IISER/Codes&Data2/Squeeze/GBM/MainSqueezeMax.xlsx')
# page=y.parse(0)
t4star=np.array(page.Kurtosis)
plt.figure(figsize=(12,7))
plt.boxplot([t4.IO1,t4.IO2,t4.IO3,t4.IO4,t4.IO5,t4.IO6,t4.IO7,t4.IO8,t4.IO9,t4.I10,t4.I11,t4.I12,t4.I13,t4.I14,t4.I15,t4.I16,t4.I17,t4.I18],0," ",labels=lb)
plt.scatter(x=range(1,19),y=t4star,marker="D",c="g")
plt.xlabel("Indices")
plt.ylabel("T4",rotation=0)
# plt.ylabe(rotation=90)
plt.savefig("t4GBMJumpSqueeze",format="png")
plt.show()
| GBMPlots.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3
# language: python
# name: python3
# ---
# # Hiding Input Cells
#
# In many cases, what matters in a notebook is the result. For a cleaner presentation of the generated data, you might want to hide the code producing the output.
#
# This little script allows to do just that.
# + language="javascript"
# document.code_shown = true;
# document.code_toggle = function(element, when_shown, when_hidden) {
# console.log(element);
# var text;
# if (document.code_shown){
# $('div.input').hide('500');
# text = when_hidden;
# } else {
# $('div.input').show('500');
# text = when_shown;
# }
# element.innerHTML = text;
# document.code_shown = !document.code_shown;
# }
# -
# Notice that the function `code_toggle` is created as a member of `document`, to make it visible to other cells.
#
# All you need now is a dynamic component that can call this code in order to Show or Hide the input blocks. Any place in the document and any dynamic widget will do; for this example, we can use a simple button.
# + language="html"
# <button id="toggleButton"
# onclick="document.code_toggle(this, 'Hide', 'Show')"
# >Hide</button>
# -
# ## Hiding code by default at startup
#
# The following code snippet will invoke the function when the page is first loaded (just add it to any `%%javascript` code cell in the notebook, or `<script>` block in an `%%html` code cell).
#
# ```javascscript
# $( document ).ready(function(){
# document.code_shown=false;
# $('div.input').hide()
# });
# ```
| HideCode.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 3 (ipykernel)
# language: python
# name: python3
# ---
# +
from atomphys import Atom
import numpy as np
import matplotlib.pyplot as plt
from math import pi as π
# This example uses pint, it has to be installed
try:
import pint
except ImportError:
print('This example makes extensive use of the pint module. If it is not installed several code blocks will error')
# -
# fetch the NIST transition data for Mg+
Mg = Atom('Mg+')
units = Mg.units
c = units.c
ε_0 = units.ε_0
Mg('S1/2')
# calculate the static polarizability for the ground state
Mg('S1/2').α().to('h Hz/(V/cm)^2')
# calculate the dynamic polarizability for the ground state at 1064 nm
α0 = Mg('S1/2').α(λ=1064 * units.nm)
print((α0/(2*c*ε_0)).to('h Hz/(W/cm^2)'))
# +
# make a figure of the AC stark shift
λ = np.linspace(100,1200, 10000) * units.nm
plt.plot(λ, ( (1/(2*c*ε_0)) * Mg('S1/2').α(λ=λ) ).to('h Hz/(W/cm^2)'), label=Mg('S1/2').term)
plt.plot(λ, ( (1/(2*c*ε_0)) * Mg('P1/2').α(λ=λ) ).to('h Hz/(W/cm^2)'), label=Mg('P1/2').term)
plt.legend()
plt.xlabel('wavelength ({:})'.format(λ.units))
plt.ylabel('AC Stark Shift h Hz/(W/cm^2))')
plt.ylim(-20,20);
# -
Rb = Atom('Rb')
# +
λ = np.linspace(500, 2000, 100000) * units.nm
I0 = units('2 W/(π*(10 um)^2)')
plt.plot(λ, ( -I0/(2*c*ε_0) * Rb('S1/2').α(λ=λ) ).to('h MHz'), label=Rb('S1/2').term)
plt.plot(λ, ( -I0/(2*c*ε_0) * Rb('P1/2').α(λ=λ) ).to('h MHz'), label=Rb('P1/2').term)
plt.ylim(-200, 200)
# -
Rb('S1/2').to('P1/2')
Rb('P1/2').transitions
| examples/Example Calculation.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: Python 2
# language: python
# name: python2
# ---
# # Problem Set 104a
#
# Complete the problems described below.
#
# Submit your work via email. A special email address has been setup for this purpose, and you can attach whatever files are appropriate to any mail sent to the address below. So I can tell which files below to whom, please **title all submitted files as**:
#
# LASTNAME firstname.py
#
# Submit this file to the class file submission email:
#
# <EMAIL>
#
# For one or more of the problems described below, a drawing or series of drawings may be required. These may be completed digitally (in Illustrator or similar) or by hand (in pencil or similar, then scanned). Please submit a single **multi-page** PDF (one page per problem) to the problem set submission email address.
from decodes.core import *
from decodes.io.jupyter_out import JupyterOut
out = JupyterOut.two_pi( grid_color = Color(0.95,0.95,0.95) )
from math import *
# ## Ontology of Collections
# ### 104a.01
# As discussed in Chapter 1.04, a variety of Python collections will be discussed in this class. Five types are briefly described around page 113 of the text. Following the samples found there, write code that constructs each of the following collections:
#
# * A ***String*** assigned to the variable `str_a` and set equal to your first and last name.
# * A ***List*** assigned to the variable `lst_a` and populated with three Points of your choosing.
# * A ***Tuple*** assigned to the variable `tup_a` and populated with a Point and an Integer number of your choosing.
# * A ***Set*** assigned to the variable `set_a` and populated with six Integer numbers of your choosing.
#
#
str_a = "<NAME>"
lst_a = [Point(),Point(),Point()]
tup_a = (Point(),10)
set_a = set([1,2,3,4,5,6])
# ## Local Structures of Control
# ### Mechanisms of Choice
# ### 104a.02
# The variable `porridge_temperature` represents the temperature in farenheit of a delicious bowl of oatmeal that a bear we know will only eat when within ten degrees of his desired temperature of 120.0 degrees. Write an `if-elif-else` block to use the print statements provided properly.
#
# +
porridge_temperature = random.uniform(70.0, 180.0)
if porridge_temperature > 130:
print('{:.2f} degrees?!! the porridge is too hot'.format(porridge_temperature))
elif porridge_temperature < 110:
print('{:.2f} degrees?!! the porridge is too cold'.format(porridge_temperature))
else:
print('THIS BEAR GONNA EAT')
# -
# ### Iteration by Condition
# ### 104a.03
# The variable `pt_a` is a Point located at some distance away from the world origin along the y-axis. Using a `while` loop, move this Point toward the origin one unit at a time until it is at or below the x-axis. Print the coordinates of `pt_a` at every step.
#
#
# +
pt_a = Point(0,random.uniform(30.0, 50.0))
while pt_a.y > 0:
pt_a.y = pt_a.y - 1
print(pt_a)
print(pt_a)
# -
# ### Exceptions
# ### 104a.04
# The given code is fragile. Modify it to include a `try-except` structure.
#
# Two Vecs are defined, `vec_b` and `vec_a` which is randomly assigned to one of the unit vectors (x,y, or z). The cross product of the two are taken, assigned to `vec_cross`, and then the angle between `vec_b` and `vec_cross` is calculated. Because the cross product of parallel vectors is the zero vector, this angle calculation might fail ungracefully and raise a ***ZeroDivisionError***. Modify the following code as to catch this error, and assign the variable `angle` to the string "NOT A NUMBER" in cases of failure.
#
#
# +
# vec_a is randomly assigned to either the unit x,y, or z-axis.
vec_a = random.choice( [Vec.ux(), Vec.uy(), Vec.uz()] )
vec_b = Vec(0,4)
#### START YOUR WORK HERE ####
# this code is fragile. wrap it carefully in a try-except structure
try:
vec_cross = vec_b.cross(vec_a)
angle = math.degrees( vec_b.angle(vec_cross) )
except:
angle = "NOT A NUMBER"
#### END YOUR WORK HERE ####
print("the angle is {}".format( angle ))
# -
# ## Sequence Types in Python
# ### Basic Features of Sequences
# ### 104a.05
# Two sequences, `looksay_seq` and `pascal_seq`, and two integers, `int_a` and `int_b` are given. Using these objects, complete the following tasks using some of the common operators and methods related to sequence types.
#
# * `int_c` is defined as an integer describing the number of objects contained in `pascal_seq`.
# * `bool_a` is a boolean that tells us if `int_a` may be found in `pascal_seq`.
# * `int_d` is the index of `int_a` in `pascal_seq`.
# * `int_e` is the number of times we may find `int_a` in `pascal_seq`.
# * `lookpas_seq` is the concatenation of `looksay_seq` and `pascal_seq` (in that order).
# * `rep_seq` is the repetition of two integers, the first item in `pascal_seq` and the last item in `looksay_seq`, three times.
#
#
# +
looksay_seq = [1, 11, 21, 1211, 111221, 312211]
pascal_seq = [1, 8, 28, 56, 70, 56, 28, 8, 1]
int_a = 56
int_b = 3
int_c = len(pascal_seq)
bool_a = int_a in pascal_seq
int_d = pascal_seq.index(int_a)
int_e = pascal_seq.count(int_a)
lookpas_seq = looksay_seq + pascal_seq
rep_seq = [pascal_seq[0],looksay_seq[-1]] * 3
print(int_b)
print(bol_a)
print(int_c)
print(int_d)
print(lookpas_seq)
print(rep_seq)
# -
# ### Slicing
# ### 104a.06
# Anagram time!
#
# Using only slices of `str_b` and the concatenation operator, define `str_c` as a String that reads `"they see"`
#
#
# +
str_b = "the eyes"
str_c = str_b[:3] + str_b[5] + str_b[3] + str_b[-1] + str_b[2] + str_b[2]
print(str_c)
# -
# ### 104a.07
# The Integer `cnt` is given, which may be *any positive **even** number*.
#
# Two Lists of Points, `pts_a` and `pts_b` are given, each of which contains `cnt` number of Points. Slice the Lists `pts_a` and `pts_b` such that then given code produces a ***shoelace-like*** 'zig-zag' pattern of Segments.
#
#
# +
cnt = 8
pts_a = [Point(-1,y) for y in Interval(-3,3).divide(cnt,True) ]
pts_b = [Point(1,y) for y in Interval(-3,3).divide(cnt,True) ]
out.put(pts_a)
out.put(pts_b)
#### START YOUR WORK HERE ####
pts_a1 = pts_a[::2]
pts_b1 = pts_b[1::2]
#pts_a2 = pts_a[1::2]
#pts_b2 = pts_b[::2]
#### END YOUR WORK HERE ####
for n in range(int(cnt/2)):
seg_zig1 = Segment(pts_a1[n],pts_b1[n])
seg_zag1 = Segment(pts_b1[n],pts_a1[n+1])
out.put([seg_zig1,seg_zag1])
#seg_zig2 = Segment(pts_b2[n],pts_a2[n])
#seg_zag2 = Segment(pts_a2[n],pts_b2[n+1])
#out.put([seg_zig2,seg_zag2])
out.draw()
out.clear()
# -
# ## Tuples
# ### 104a.08
# Here we demonstrate the calculation of the weighted centroid of a set of Points.
#
# Two Lists of length `cnt` are given: a List of Points at random locations `pts_c`, and a List of numeric weights `weights` that range from 0.0 to 1.0. Code is given to calculate the weighted centroid of these Points and to plot the result to the canvas.
#
# Complete this code by writing a loop that iterates over corresponding objects in the `pts_c` and `weights` Lists. At each cycle, calcuate the **sum** of **the x-coordinate of the Point** and the **cooresponding weight value** to the `x_sum` variable. Do the same for the y-coordinate of the Point and the `y_sum` variable.
#
#
# +
cnt = 5
pts_c = [Point.random(Interval(-3.5,3.5)) for n in range(cnt)]
weights = [random.random() for n in range(cnt)]
w_sum = sum(weights)
x_sum, y_sum = 0,0
#### START YOUR WORK HERE ####
for pt, weight in zip(pts_c,weights):
x_sum += pt.x * weight
y_sum += pt.y * weight
#### END YOUR WORK HERE ####
pt_cent = Point( x_sum/w_sum, y_sum/w_sum )
pt_cent.set_color(1,0,0)
for n in range(cnt):
pt = pts_c[n]
pt.set_weight(Interval(2,10).eval( weights[n] ) )
out.put(pt)
out.put(pt_cent)
out.draw()
out.clear()
# -
# ## Strings
# ### 104a.09
# In CNC machining, G-Code is a common file format for defining the movements of numerically-controlled machines. At its most basic, the format is simple: a series of desired positions is described along with an indication of how to move from one place to the next (along with other encoded commands that we won't discuss here). Each line of a G-code file might begin with a descriptor of the desired movement, for example G01 indicates linear interpolation, followed by the x-y coordinates of the desired location.
#
# Using the code provided below, ***complete the defining of the G-Code file String `g_code`***. To this end, two corresponding Lists of x- and y- values (converted to Strings) are given, `x_coords` and `y_coords` that describe the intended location of the CNC actuator. Also given is a String `fmt_str` that should be used along with the `format()` method in the construction of individual lines of G-Code.
#
# If you're curious, you might note that the `\n` code creates a new line in text formatting.
#
#
# +
cnt = 10
x_coords = ['{:.3f}'.format(2.0*cos(t)) for t in Interval.twopi()/cnt]
y_coords = ['{:.3f}'.format(3.0*sin(t)) for t in Interval.twopi()/cnt]
print(x_coords)
print(y_coords)
fmt_str = "G01 {0},{1}\n"
g_code = "OPEN PROG 1000 CLEAR\n"
#### START YOUR WORK HERE ####
for x,y in zip(x_coords, y_coords):
g_code += fmt_str.format(x,y)
#### END YOUR WORK HERE ####
g_code += "CLOSE"
print(g_code)
# -
# ## Sorting
# ### 104a.10
# The decorate-sort-undecorate pattern of code is exceedingly useful in computational geometry for design applications. Use this pattern to solve the following problem.
#
# A number of Points that fall around the model origin are given as `pts_unsorted`. Sort these Points by their angle relative to the world x-axis and assign them to the `pts_sorted` variable. Use the `decorated_tups` variable in your process. A routine is given for visualizing the order of a List of Points by color, and may be applied to the Points as given or as resulting.
# +
# the following construction is termed a 'list comprehension'.
# We'll know more about this soon. For now we can just accept
# that a collection of Points at random location is defined.
pts_unsorted = [Point.random(Interval(-3,3),True) for n in range(100)]
pts_sorted = []
#### START YOUR WORK HERE ####
decorated_tups = []
for pt in pts_unsorted:
# each tuple pairs a numeric angle with a related Point
decorated_tups.append( (Vec(pt).angle(Vec.ux()), pt) )
#the tuples are sorted by their first value
decorated_tups.sort()
# the list of tuples is 'unpacked' to a regular list of points
for tup in decorated_tups:
pts_sorted.append(tup[1])
# change this assignment to pts_sorted to see the results of your sort
pts_to_visualize = pts_sorted
#### END YOUR WORK HERE ####
# what follows is a routine for visualizing the order of a List of Points.
# it's okay if you don't understand all this.
ca,cb = Color(1,0,0), Color(0,1,1)
for n, pt in enumerate(pts_to_visualize): pt.set_color( Color.interpolate( ca,cb,float(n)/len(pts_to_visualize) ) )
out.put(pts_to_visualize)
out.draw()
out.clear()
# -
| problem sets/Problem Set 104a - KEY.ipynb |
# ---
# jupyter:
# jupytext:
# text_representation:
# extension: .py
# format_name: light
# format_version: '1.5'
# jupytext_version: 1.14.4
# kernelspec:
# display_name: 'Python 3.7.3 64-bit (''base'': conda)'
# language: python
# name: python37364bitbaseconda210f926cb548430eaeeaaca39b8496cc
# ---
# # Rearranging the system matrices from (<NAME>, 1999)
#
# [1] <NAME>., & <NAME>. (1999). Analytical characterization of the unique properties of planetary gear free vibration. Journal of Vibration and Acoustics, Transactions of the ASME, 121(3), 316–321. http://doi.org/10.1115/1.2893982 \
# [2] <NAME>., & <NAME>. (2012). Vibration Properties of High-Speed Planetary Gears With Gyroscopic Effects. Journal of Vibration and Acoustics, 134(6). http://doi.org/10.1115/1.4006646
# +
from sympy import *
init_printing()
def symb(x,y):
return symbols('{0}_{1}'.format(x,y), type = float)
| notes/.ipynb_checkpoints/Untitled-checkpoint.ipynb |